WO2025019675A1 - Prime editing system and uses thereof - Google Patents
Prime editing system and uses thereof Download PDFInfo
- Publication number
- WO2025019675A1 WO2025019675A1 PCT/US2024/038551 US2024038551W WO2025019675A1 WO 2025019675 A1 WO2025019675 A1 WO 2025019675A1 US 2024038551 W US2024038551 W US 2024038551W WO 2025019675 A1 WO2025019675 A1 WO 2025019675A1
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- editing
- pegrna
- sequence
- seq
- cells
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Pending
Links
Classifications
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/87—Introduction of foreign genetic material using processes not otherwise provided for, e.g. co-transformation
- C12N15/90—Stable introduction of foreign DNA into chromosome
- C12N15/902—Stable introduction of foreign DNA into chromosome using homologous recombination
- C12N15/907—Stable introduction of foreign DNA into chromosome using homologous recombination in mammalian cells
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/11—DNA or RNA fragments; Modified forms thereof; Non-coding nucleic acids having a biological activity
- C12N15/113—Non-coding nucleic acids modulating the expression of genes, e.g. antisense oligonucleotides; Antisense DNA or RNA; Triplex- forming oligonucleotides; Catalytic nucleic acids, e.g. ribozymes; Nucleic acids used in co-suppression or gene silencing
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/11—DNA or RNA fragments; Modified forms thereof; Non-coding nucleic acids having a biological activity
- C12N15/52—Genes encoding for enzymes or proenzymes
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/11—DNA or RNA fragments; Modified forms thereof; Non-coding nucleic acids having a biological activity
- C12N15/62—DNA sequences coding for fusion proteins
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N9/00—Enzymes; Proenzymes; Compositions thereof; Processes for preparing, activating, inhibiting, separating or purifying enzymes
- C12N9/10—Transferases (2.)
- C12N9/12—Transferases (2.) transferring phosphorus containing groups, e.g. kinases (2.7)
- C12N9/1241—Nucleotidyltransferases (2.7.7)
- C12N9/1276—RNA-directed DNA polymerase (2.7.7.49), i.e. reverse transcriptase or telomerase
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N9/00—Enzymes; Proenzymes; Compositions thereof; Processes for preparing, activating, inhibiting, separating or purifying enzymes
- C12N9/14—Hydrolases (3)
- C12N9/16—Hydrolases (3) acting on ester bonds (3.1)
- C12N9/22—Ribonucleases [RNase]; Deoxyribonucleases [DNase]
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K38/00—Medicinal preparations containing peptides
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N2310/00—Structure or type of the nucleic acid
- C12N2310/10—Type of nucleic acid
- C12N2310/14—Type of nucleic acid interfering nucleic acids [NA]
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N2310/00—Structure or type of the nucleic acid
- C12N2310/30—Chemical structure
- C12N2310/31—Chemical structure of the backbone
- C12N2310/315—Phosphorothioates
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N2310/00—Structure or type of the nucleic acid
- C12N2310/30—Chemical structure
- C12N2310/32—Chemical structure of the sugar
- C12N2310/321—2'-O-R Modification
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N2740/00—Reverse transcribing RNA viruses
- C12N2740/00011—Details
- C12N2740/10011—Retroviridae
- C12N2740/16011—Human Immunodeficiency Virus, HIV
- C12N2740/16041—Use of virus, viral particle or viral elements as a vector
- C12N2740/16043—Use of virus, viral particle or viral elements as a vector viral genome or elements thereof as genetic vector
Definitions
- the present disclosure relates to compositions and systems for genome editing and uses thereof.
- the present disclosure shows modifying synthetic pegRNAs to allow SSB protein binding and overexpression of small RNA binding exonuclease protection factor La (SSB) protein can improve prime editing.
- SSB small RNA binding exonuclease protection factor La
- a system comprising a Cas9 nickase; a reverse transcriptase; a small RNA binding exonuclease protection factor La (SSB) protein; and a prime editing guide RNA (pegRNA).
- SSB small RNA binding exonuclease protection factor La
- pegRNA prime editing guide RNA
- the pegRNA comprises a 3 ’-polyuridine domain.
- the 3 ’-polyuridine domain comprises at least one chemically modified uridine (for example, at least two, three, four, or five chemically modified uridines).
- the 3 ’-polyuridine domain comprises at least one unmodified uridine (for example, at least two, three, four, or five unmodified uridines).
- the 3’- polyuridine domain comprises at least one chemically modified uridine (for example, at least two, three, four, or five chemically modified uridines) and at least one unmodified uridine (for example, at least one, two, three, four, or five unmodified uridines).
- the at least one unmodified uridine locates at the 3’ end of the pegRNA.
- the chemical modification is 2’ -O-m ethylation and/or replacement of a phosphodiester bond to a phosphorothioate bond.
- the 3 ’-polyuridine domain comprises at least one (for example, at least two, three, four, or five) uridine with unmodified 2'-hydroxyl (OH) group.
- the at least one uridine with unmodified 2'- OH group locates at the 3’ end of the pegRNA.
- the SSB protein of the system of any preceding aspect comprises a sequence at least 80% identical to SEQ ID NO: 34 or a fragment thereof.
- the SSB protein comprises a La motif and/or an RNA recognition motif (RRM) (e.g., amino acid residues 1-194 or 2-194 of SEQ ID NO: 34).
- RRM RNA recognition motif
- the SSB protein comprises a sequence at least 80% identical to SEQ ID NO: 33 or SEQ ID NO: 35 or a fragment thereof.
- the Cas9 nickase comprises a sequence at least 80% identical to SEQ ID NO: 26 or 27, or a fragment thereof.
- the SSB protein is operatively linked to the Cas9 nickase and the reverse transcriptase.
- the system of any preceding aspect comprises a recombinant polypeptide, wherein the recombinant polypeptide comprises a sequence at least 80% identical to any of SEQ ID NOs: 1-12 or a fragment thereof.
- a system comprising a first polynucleotide encoding a Cas9 nickase; a second polynucleotide encoding a reverse transcriptase; and a third polynucleotide encoding a small RNA binding exonuclease protection factor La (SSB) protein; and a prime editing guide RNA (pegRNA).
- SSB small RNA binding exonuclease protection factor La
- pegRNA prime editing guide RNA
- the third polynucleotide encodes a La motif and/or an RNA recognition motif (RRM) of the SSB protein.
- the third polynucleotide comprises a sequence at least 80% identical to SEQ ID NO: 50 or a fragment thereof.
- the third polynucleotide comprises a sequence at least 80% identical to SEQ ID NO: 49 or 51 or a fragment thereof.
- the first, second, and third polynucleotides are operatively linked thereby forming one recombinant polynucleotide.
- the recombinant polynucleotide comprises a sequence at least 80% identical to any of SEQ ID NOs: 13-24.
- the first, second, and third polynucleotides are located on a same or different vectors.
- composition comprising the system, polypeptides, or polynucleotides of any preceding aspect.
- Also disclosed herein is a method of treating a genetic disorder in a subject in need, comprising administering to the subject a therapeutically effective amount of the system, polypeptides, or polynucleotides of any preceding aspect or a pharmaceutical composition of any preceding aspect. Also disclosed herein is a method for altering expression of a gene product in a cell, comprising introducing into the cell an effective amount of the system, polypeptides, or polynucleotides of any preceding aspect.
- FIGS. 1A-1H show Genome-scale CRISPRi screens identify La as a determinant of prime editing.
- FIG. 1 A shows schematic of prime editing.
- FIG. IB shows a schematic of FACS reporter that expresses GFP upon installation of +7 GGto CA substitution prime edit (mCherry marker constitutively expressed).
- FIG. 1C shows flow cytometry analysis of GFP expression in K562 CRISPRi cells with integrated FACS reporter with and without prime editing (+7 GG to CA, PE3 with a +50 complementary strand nick) and with and without transduction of an MSH2-targeting sgRNA.
- FIG. ID shows the gene-level phenotypes from genome-scale CRISPRi screen performed in FACS reporter cells with +7 GGto CA edit using PE3 approach.
- Phenotypes represent enrichment of normalized sgRNA counts in GFP+ over GFP- populations after prime editing. Genes identified as hits using CRISPhieRmix (FDR ⁇ 0.01) and pseudogene controls generated from randomly selected non-targeting sgRNAs are indicated.
- FIG. IE shows the quantification of CRISPRi-mediated La depletion. RT-qPCR data collected from K562 CRISPRi cells with integrated MCS reporter. Data are normalized to ACTB and presented relative to a non-targeting sgRNA.
- FIG. 1G show a comparison of prime editing outcomes using a pegRNA (left) or an epegRNA (right, tevopreQi) with the PE2 approach (plasmid delivery) at stably integrated FACS reporter in K562 CRISPRi cells after transduction of a La-targeting or non-targeting sgRNA.
- FIG. 1H shows a comparison of prime editing outcomes with the indicated edit and approach (plasmid delivery) at integrated MCS reporter with and without depletion of La in K562 CRISPRi cells.
- FIGS. 2A-2H show that La promotes prime editing across edit types and genomic loci in multiple cell lines. FIG.
- FIG. 2A shows western blot analysis of K562 cells constitutively expressing PEmax (K562 PEmax cell line) and derived clones with genetic disruption of La (La-kol through La-ko5). Whole-cell lysates were sequentially immunoblotted for La, GAPDH, and prime editor protein (PEmax) with corresponding antibodies. Asterisks denote La knockout cell lines used in this study.
- FIG. 2B shows percentages of intended prime editing and indels at multiple genomic loci in K562 PEmax and La-ko4 cells. pegRNAs and epegRNAs (evopreQi) were delivered as plasmids without or with MLHldn (PE2 or PE4, respectively).
- FIG. 2c shows percentages of intended prime editing and indels at the endogenous DNMT1 locus in K562 PEmax and La-ko4/5 cells with or without ectopic expression of La.
- Expression plasmids for La or an mRFP control were delivered alongside plasmids encoding pegRNA or epegRNA (evopreQi) specifying a +5 G to T edit.
- FIG. 2D shows quantification of siRNA-mediated La depletion. RT-qPCR data were collected from HEK293T cells at specified time points during prime editing.
- FIG. 2E shows fold changes in indicated editing outcomes across ten PE3 edits (substitutions, insertions, and deletions) at five genomic loci in HEK293T cells with or without La depletion by siRNAs. Editing components delivered by plasmid transfection. Editing percentages presented in FIG. 8F.
- FIG. 2F shows effects of La depletion on DSB repair. Schematic of the MCS reporter (top), with distances between predicted SaCas9 cut site and sequences required for GFP expression indicated.
- FIG. 2G shows fold changes in SaPE2, PE4 approach-, SaCas9-, SaBE4- and SaABE8e-induced editing outcomes in La-ko4 relative to parental controls (intended edits only).
- the same pegRNA expression plasmid was used for the editing systems at each of the four genomic loci. Editing percentages presented in FIGS. 9C-9F.
- FIGS. 9C-9F show fold changes in SaCas9-, SaBE4-, and SaABE8e-induced editing outcomes in La-ko4 relative to parental controls (intended edits only).
- the same sgRNA expression plasmid was used for the editing systems at each of the four genomic loci. Editing percentages presented in FIGS. 9C-9F.
- Indel frequency for each sample included adjacent to corresponding intending editing efficiency in FIGS. 2B-2C.
- FIGS. 3A-3J show that La functionally interacts with the 3' ends of polyuridylated pegRNAs.
- FIG. 3 A shows schematics of La domain architecture (top) and five La mutants used in FIG. 3B (bottom).
- FIG. 3B shows percentages of intended prime editing and indels at the endogenous DNMT1 locus with or without ectopic expression of La or La mutants in a.
- Expression plasmids (La, mutants, or mRFP control) were delivered to K562 PEmax and La- ko4 cells alongside plasmids encoding one pegRNA (+5 G to T edit).
- FIG. 1 shows schematics of La domain architecture (top) and five La mutants used in FIG. 3B (bottom).
- FIG. 3B shows percentages of intended prime editing and indels at the endogenous DNMT1 locus with or without ectopic expression of La or La mutants in a.
- Expression plasmids (La, mutants, or
- FIG. 3C shows the chemical structure of RNA with phosphorothioate bonds (*) and 2'-O-methylation modifications (m).
- FIG. 3D shows percentages of intended prime editing and indels at the endogenous DNMT1 locus in K562 PEmax and La-ko4 cells using 100 pmole of synthetic pegRNAs with indicated 3' end sequences and chemical modifications.
- FIG. 3E shows fold changes in average intended prime editing achieved at four genomic loci in La-ko4 cells relative to parental controls using 100 pmole synthetic pegRNAs with indicated no-polyU, blocked, or La-accessible end configurations. Editing percentages presented in FIG. 10E.
- FIG. 3F shows a model of La interaction with 3' ends of polyuridylated pegRNAs promotes prime editing.
- FIG. 3G shows a schematic of pegRNA specifying RUNX1 +5 G to T with the minimum sequence defining each class of small RNA-seq fragments highlighted (c/.s-active, orange; /ra//.s-active, purple). The edit-encoding nucleotide and cryptic terminators are also indicated (white base and green asterisks, respectively).
- FIGS. 3H-3I show coverage plots of small RNA-seq fragments aligned to pegRNA (h) or epegRNA (i) encoding RUNX1 +5 G to T.
- Alignment categories are indicated (human small RNA, gray; cv.s-active, orange; trans-active, purple; premature termination, green) and genes with adjusted /z-values ⁇ 0.05 are highlighted in light gray (calculated by DESeq2 using the Wald test).
- Nucleotide position 0 denotes the 5' end of the RNA, and positions of the edit-encoding nucleotide (vertical solid line) and the start of PBS (vertical dashed line) are indicated. Shaded areas represent sgRNA sequence, Pol III terminator for pegRNA, and linker plus evopreQi and Pol III terminator for epegRNA.
- FIGS. 4A-4I show fusion of La RNA-binding N-terminal domain to PEmax improves prime editing.
- FIG. 4A shows architectures of prime editors. Medium gray NLS, bpNLS SV40 ; Dark gray NLS, NLS c ‘ Myc ; A, 32 amino acid linker; B, 34 amino acid linker; C, SGGS linker; La, full length La or the N-terminal domain of La (La 1-194 ); MMLV-RT, human codon- optimized MMLV-RT.
- FIG. 4A shows architectures of prime editors.
- FIG. 4B shows percentages of intended prime edits and indels produced with the indicated editors (from a) and pegRNA or epegRNA (evopreQi) at the endogenous DNMT1 and VEGFA loci in HEK293T and U2OS cells, respectively. Editing components (PE2) delivered by plasmid transfection.
- FIG. 4C shows percentages of intended prime editing and indels at the endogenous DNMT1 and VEGFA loci in HEK293T, HeLa, and U2OS cells. Editing components delivered by plasmid transfection.
- FIG. 1 shows percentages of intended prime edits and indels produced with the indicated editors (from a) and pegRNA or epegRNA (evopreQi) at the endogenous DNMT1 and VEGFA loci in HEK293T and U2OS cells, respectively. Editing components (PE2) delivered by plasmid transfection.
- FIG. 4C shows percentages of intended prime editing and ind
- FIG. 4D shows percentages of intended prime editing and indels at eight endogenous loci in U2OS cells using pegRNAs or epegRNAs (mpknot: HEK3, tevopreQi: HEK4, evopreQi: all other loci). Editing components delivered by plasmid transfection. pegRNA data also presented in FIG. 15B.
- FIG. 4E shows percentages of intended prime editing and indels at the endogenous HEK3 locus in HEK293T cells. Editing components delivered by plasmid transfection.
- FIG. 4F shows fold changes in intended prime editing (left) and ratios of intended editing to indel frequency (right) for each indicated condition. Editing percentages presented in FIG. 4D.
- FIG. 4G shows a schematic of interactions between La N-terminal domain and RNA with 3 '-UUUOH. Four residues were mutated in PE7 mutant to disrupt 3' polyU binding (Q20, Y23, Y24 and F35; indicated in red).
- FIG. 4H shows a schematic of PE7 mutant harboring four mutations in La 1-194 to disrupt 3’ polyU binding (Q20A, Y23A, Y24F and F35A; indicated by red lines).
- FIG. 41 shows percentages of intended prime edits and indels produced with PEmax, PE7 or PE7 mutant at the endogenous RUNX1 and VEGFA loci in U2OS cells. Editing components delivered by plasmid transfection.
- FIGS. 4C-4E, and 41 Indel frequency for each sample included adjacent to corresponding intending editing efficiency in FIGS. 4C-4E, and 41.
- One-tailed unpaired Student’s /-test (FIGS. 4B-4E, and 41). *P ⁇ 0.05.
- FIGS. 5A-5H show that PE7 enhances prime editing at disease-related targets and primary human cells.
- FIG. 5A-5H show that PE7 enhances prime editing at disease-related targets and primary human cells.
- FIG. 5A Percentages of intended prime editing and indels at six endogenous loci in U2OS cells using pegRNAs and epegRNAs (tevopreQi). Editing components delivered by plasmid transfection. pegRNA data also presented in FIG. 5C.
- FIG. 5B Fold changes in intended prime editing for each indicated condition. Editing percentages presented in FIG. 5A.
- FIG. 5C Summary plot of intended prime edit and indel frequencies produced with indicated editor and prime editing approaches at genomic loci. Data for PE2 and PE4 from six loci indicated in a plus HBG1/2. Data for PE3 and PE5 from a subset of those targets (HBB, PRNP, IL2RB, CXCR4).
- FIG. 5D Percentages of intended prime editing and indels at four genomic loci in K562 cells using indicated editor and synthetic pegRNAs with no-polyU, blocked or La-accessible end configurations.
- FIG. 5E Fold changes in average intended prime editing in K562 cells using PE7 mRNA relative to PEmax mRNA for synthetic pegRNAs with each indicated end configuration. Editing percentages presented in FIG. 5D.
- FIG. 5F Percentages of intended prime editing and indels in primary human T cells using PEmax or PE7 mRNA and synthetic pegRNAs with indicated end configurations.
- FIG. 5G Fold changes in intended prime editing in primary human T cells using PE7 mRNA and synthetic pegRNAs with La-accessible end configuration relative to intended editing with PEmax mRNA and the same pegRNAs at eight genomic loci.
- Data in FIG. 5G indicate ratios of values for individual edits and donors (4 different T cell donors) and horizontal bar in FIG. 5G indicate median. Fold changes included for select comparisons in FIGS. 5A, 5D and 5F.
- FIGS. 6A-6I show characterization of prime editing reporters before and during genome-scale CRISPRi screens.
- FIG. 6A shows a schematic of isolating prime edited cells with intended edit using the FACS reporter.
- the reporter expresses GFP upon installation of select prime edits, thus enabling separation of cells into mostly edited or mostly unedited populations using flow cytometry.
- the complete FACS reporter is depicted in FIG. IB.
- FIG. 6B shows a schematic of isolating prime edited cells with intended edit using the MCS reporter.
- the reporter expresses a synthetic cell surface marker (IgK-hlgGl-Fc-PDGFRP) upon installation of select prime edits, thus enabling facile separation of cells into mostly edited or mostly unedited populations using magnetic Protein G beads.
- IgK-hlgGl-Fc-PDGFRP synthetic cell surface marker
- FIG. 6C shows three prime edits capable of ‘switching on’ the FACS and MCS reporters (depicted with the former).
- FIG. 6D shows flow cytometry analysis of GFP expression from the FACS reporter after prime editing with each of the edits depicted in FIG. 6C.
- FIG. 6E shows percentages of prime editing outcomes in GFP+ or GFP- cell populations sorted by flow cytometry after editing with each of the edits depicted in FIG. 6C. Outcomes quantified by sequencing the FACS reporter target site.
- FIG. 6F shows percentages of prime editing outcomes in bead-bound or unbound cell populations isolated by magnetic separation after editing with each of the edits depicted in c.
- FIG. 6G shows flow cytometry analysis of GFP expression in the FACS reporter cells (ie., K562 CRISPRi cells with stably integrated FACS reporter) after transduction with genome-scale CRISPRi library (hCRISPRi-v2) and prime editing by plasmid transfection (+7 GG to CA, PE3). Data from repeat measurements of each replicate of the genome-scale screen.
- FIG. 6H shows percentages of prime editing outcomes observed in GFP+ or GFP- cell populations for each replicate of the genome-scale FACS screen. Outcomes quantified by sequencing the FACS reporter target site.
- FIG. 6G shows flow cytometry analysis of GFP expression in the FACS reporter cells (ie., K562 CRISPRi cells with stably integrated FACS reporter) after transduction with genome-scale CRISPRi library (hCRISPRi-v2) and prime editing by plasmid transfection (+7 GG to CA, PE3). Data from repeat measurements of each replicate of the genome-scale screen.
- FIG. 6H shows
- FIG. 61 shows sequences and frequencies of alleles observed at the FACS reporter target site in cell populations sorted for replicate 1 of the genome-scale FACS screen.
- FIGS. 7A-7I show results of genome-scale CRISPRi screens performed with FACS and MCS reporters.
- FIG. 7A Pearson correlations of read counts per sgRNA between each pair of samples isolated from genome-scale FACS screen.
- FIG. 7B sgRNA-level phenotypes from each replicate of the genome-scale FACS screen.
- FIG. 7C Gene-level phenotypes (average of replicates) and per gene FDRs from the genome-scale FACS screen, as determined by CRISPhieRmix analysis.
- FIG. 7D Pearson correlations of read counts per sgRNA between each pair of samples isolated from the genome-scale MCS screen performed with the PE3 approach.
- FIG. 7A Pearson correlations of read counts per sgRNA between each pair of samples isolated from genome-scale FACS screen.
- FIG. 7F Gene-level phenotypes (average of replicates) from genome- scale FACS and MCS screens performed with the PE3 approach.
- FIGS. 7G-7I Gene-level phenotypes from each replicate of MCS reporter screens performed with the PE3 (FIG. 7G), PE4 (FIG. 7H) and PE5 (FIG. 71) approaches.
- FIGS. 7B-7E sgRNAs targeting genes identified as hits (FDR ⁇ 0.01) using CRISPhieRmix are highlighted in red.
- FIGS. 7C, 7F- 7G genes identified as hits (FDR ⁇ 0.01) in the indicated screen using CRISPhieRmix are highlighted in red.
- FIGS. 8A-8F show validation of La phenotypes with various genetic perturbation modalities.
- FIG. 8A shows a schematic of workflow used to engineer K562 clonal cell lines with PEmax expressed constitutively from the AAVS1 safe-harbor locus (K562 PEmax cell line).
- FIG. 8B shows sequences and frequencies of alleles observed at the La locus in the La- knockout clones used in this study (La-ko3 through La-ko5).
- FIG. 8C shows full images of western blot presented in FIG. 2 A.
- FIG. 8D shows cumulative population doublings of La-ko4 and La-ko5 cells compared to K562 PEmax parental cells.
- FIG. 8A shows a schematic of workflow used to engineer K562 clonal cell lines with PEmax expressed constitutively from the AAVS1 safe-harbor locus (K562 PEmax cell line).
- FIG. 8B shows sequences and frequencies of alleles observed at the La locus
- FIG. 8E shows flow cytometry analysis of GFP expressed from the PEmax construct at the AAVS1 locus in K562 PEmax parental, La-ko3, La-ko4 and La-ko5 cells prior to transfection in FIG. 2C.
- FIG. 8F shows percentages of intended prime editing and indels across ten edits with pegRNAs (top) or epegRNAs (bottom, mpknot: HEK3, evopreQi : all other loci) at five genomic loci in HEK293T cells with and without depletion of La by siRNAs.
- Prime editing components PE3 were delivered as expression plasmids. Percentages in FIG.
- FIGS. 9A-9F show that La has a stronger impact on prime editing than other editing modalities.
- FIG. 9A shows flow cytometry analysis of GFP expression from a stably integrated MCS reporter in K562 CRISPRi cells after transduction of indicated sgRNAs and editing with SaCas9 nuclease. Editing components (SaCas9, +7 GG to CA pegRNA) delivered by plasmid transfection.
- FIG. 9B shows quantification of SaCas9-induced indels at stably integrated MCS reporter described in FIG. 9 A.
- FIGS. 9A-9F show that La has a stronger impact on prime editing than other editing modalities.
- FIG. 9A shows flow cytometry analysis of GFP expression from a stably integrated MCS reporter in K562 CRISPRi cells after transduction of indicated sgRNAs and editing with SaCas9 nuclease. Editing components (SaCas9, +7 GG to CA peg
- FIGS. 9C-9F show percentages of intended editing achieved in K562 PEmax parental and La-knockout cells using SaPE2 with PE4 approach, SaCas9, SaBE4, and SaABE8e across four genomic loci, HEK3 (FIG. 9C), EMX1 (FIG. 9D), FANCF (FIG. 9E) and HBB (FIG. 9F).
- the same pegRNA or sgRNA expression plasmid was used for the editing systems at each target, with select combinations excluded (ie., SaPE2 with PE4 approach with any sgRNA and SaBE4 at EMX1).
- FIGS. 10A-10E show prime editing with synthetic pegRNAs designed to block or allow La binding reveals functional interaction between La and polyuridylated 3' ends.
- FIG. 10A shows chemical structures of ribonucleotides linked by a phosphorothioate bond (left) or with substitution of ribose 2'-OH for 2'-O-methyl groups (2'-0Me) (right).
- FIG. 10B shows percentages of intended prime editing and indels at the endogenous DNMT1 locus in K562 PEmax parental cells using synthetic pegRNA with indicated 3' end configuration. Input was titrated from 0 to 500 pmole at 100 pmole increments.
- FIGS. 10A shows chemical structures of ribonucleotides linked by a phosphorothioate bond (left) or with substitution of ribose 2'-OH for 2'-O-methyl groups (2'-0Me) (right).
- FIG. 10B shows percentages of intended prime editing and
- FIG. 10C-10D shows percentages of intended prime editing and indels at the endogenous HEK3 (c, PE3) and DNMT1 (b, PE2) loci in K562 PEmax parental, La-ko4, and La-ko5 (where indicated) cells using 100 pmole of synthetic pegRNAs with specified 3' end sequences and chemical modifications.
- FIG. 10E shows percentages of intended prime editing and indels at endogenous DNMT1, CXCR4, VEGFA and RUNX1 loci in K562 PEmax parental and La-ko4 cells using 100 pmole of synthetic pegRNAs with indicated 3' end configurations.
- FIGS. 11 A-l ID show details of small RNA-seq experiment performed with two sets of (e)pegRNAs.
- FIG. 11A shows composition of small RNA-seq libraries from K562 PEmax parental or La-ko4 cells. Data from samples collected one and two days after transfection of eleven (e)pegRNAs in two sets.
- FIG. 1 IB shows fold changes in normalized counts of indicated biotypes in La-ko4 cells relative to parental controls, from samples collected one and two days after transfection of eleven (e)pegRNAs in two sets. Counts were calculated per replicate independently for each set of (e)pegRNAs as the sum of properly aligned fragments classified as each biotype and normalized by total RNA counts.
- FIG. 11A shows composition of small RNA-seq libraries from K562 PEmax parental or La-ko4 cells. Data from samples collected one and two days after transfection of eleven (e)pegRNAs in two sets.
- FIG. 1 IB shows fold changes in normalized
- Data are from samples collected one day after pegRNA plasmid transfection and normalized by counts of fragments from total human small RNA (top) or those within the corresponding bins (bottom). Plotted data represent coverages of indicated bins (c/.s-active, /ra/z.s-active or inactive) in specified cell lines.
- FIGS. 12A-12C show additional coverage plots of (e)pegRNAs from small RNA-seq experiment performed with two sets of (e)pegRNAs.
- FIGS. 12A-12C Coverage plots of small RNA-seq fragments aligned to pegRNA (left) or epegRNA (right) encoding EMX1 +5 G to T (FIG. 12 A), HEK3 +1 T to A (FIG. 12B) or DNMT1 +5 G to T (FIG. 12C).
- Data are from samples collected one day after (e)pegRNA plasmid transfection and normalized by counts of fragments from total human small RNA (top) or those within the corresponding bins (bottom).
- Nucleotide position 0 denotes the 5' end of the RNA, and positions of the edit-encoding nucleotide (vertical solid line) and the start of PBS (vertical dashed line) are indicated.
- Shaded areas represent sgRNA sequence, Pol III terminator for pegRNAs, and linker plus evopreQi/mpknot and Pol III terminator for epegRNAs.
- FIGS. 13A-13E show details of small RNA-seq experiment performed with nontargeting pegRNA and epegRNA, each specifying a +6 G to C edit in the Mus musculus DNMT1 gene.
- FIG. 13A shows the composition of small RNA-seq libraries from K562 PEmax parental or La-ko4 cells. Data from samples collected one and two days after transfection of plasmids encoding a pegRNA or an epegRNA specifying mouse DNMT1 +6 Gto C.
- FIG. 13A shows the composition of small RNA-seq libraries from K562 PEmax parental or La-ko4 cells. Data from samples collected one and two days after transfection of plasmids encoding a pegRNA or an epegRNA specifying mouse DNMT1 +6 Gto C.
- FIGS. 13C-13D show coverage plots of small RNA-seq fragments aligned to the pegRNA (left) or the epegRNA (right) specifying mouse DNMT1 +6 G to C edit. Data are from cells without the (e)pegRNA target collected one (FIG.
- FIGS. 14A-14I show PE7 has no or negligible effects on cell viability, cell growth, and mRNA abundance compared to PEmax and PE7 mutant.
- FIG. 14A shows percentages of intended prime editing and indels at the endogenous HEK3 and PRNP loci in K562 cells with PEmax, PE7 or PE7 mutant. Editing components delivered by plasmid transfection. Cells from this experiment were used for analyses in FIGS. 14B-14I.
- FIG. 14B shows percentages of viable K562 cells quantified by flow cytometry one, two and three days after transfection of PEmax, PE7 or PE7 mutant editor plasmid and pegRNA plasmid specifying either HEK3 +1 T to A or PRNP +6 G to T.
- FIG. 14A shows percentages of intended prime editing and indels at the endogenous HEK3 and PRNP loci in K562 cells with PEmax, PE7 or PE7 mutant. Editing components delivered by plasmid transfection. Cells from
- FIGS. 14G-14I shows venn diagrams of differentially expressed genes (p ⁇ 0.05) in K562 cells edited at two different loci across three comparisons: PE7 relative to PEmax (FIG. 14g), PE7 relative to PE7 mutant (FIG. 14H), and PEmax relative to PE7 mutant (FIG. 141). Indel frequency for each sample included adjacent to corresponding intending editing efficiency in a.
- FIGS. 15A-15G show PE7 improves prime editing with different approaches and delivery strategies without substantially increasing off-target effect.
- FIG. 15A-15G show PE7 improves prime editing with different approaches and delivery strategies without substantially increasing off-target effect.
- FIG. 15A shows percentages of editing outcomes produced by PEmax or PE7 with the PE2 approach at on- and off-target sites using pegRNAs targeting the EMX1 (left), FANCF (middle left), HEK3 (middle right), and HEK4 (right) loci in U2OS cells.
- On-target editing data also presented in FIG. 15B and FIG. 4D.
- FIG. 15B shows a summary plot of intended prime edit and indel frequencies observed at genomic loci with indicated editor and prime editing approaches.
- Data for PE2 and PE4 from eight loci indicated in FIG. 4D.
- Data for PE3 and PE5 from a subset of those targets (RNF2, HEK3, DNMT1 and VEGFA).
- FIG. 15C shows percentages of intended prime editing and indels at endogenous HEK3 (top) and DNMT1 (bottom) loci after lentiviral transduction of pegRNAs or (e)pegRNAs (tevopreQi) and transfection of PEmax or PE7 editor encoded on mRNA or plasmid in HeLa (left) and U2OS (right) cells. (e)pegRNAs use a modified sgRNA scaffold.
- FIG. 15D shows percentages of intended prime editing and indels at endogenous DNMT1 (left) and HEK3 (right) loci after lentiviral transduction of editing components in K562 cells. Two different editor expression constructs (as indicated) were tested.
- FIG. 15E shows percentages of intended prime editing and indels at three genomic loci in U2OS cells using indicated editor mRNA and synthetic pegRNAs with no-polyU, blocked or La-accessible end configurations.
- FIG. 15F shows fold changes in average intended prime editing in U2OS cells using PE7 mRNA relative to PEmax mRNA for synthetic pegRNAs with each indicated end configuration. Editing percentages presented in FIG. 15E.
- FIG. 15G shows percentages of intended prime editing and indels at five genomic loci in primary human T cells using PEmax or PE7 mRNA and synthetic pegRNAs with La-accessible end configuration.
- administering to a subject includes any route of introducing or delivering to a subject an agent. Administration can be carried out by any suitable route, including oral, topical, intravenous, subcutaneous, transcutaneous, transdermal, intramuscular, intra-joint, parenteral, intra-arteriole, intradermal, intraventricular, intracranial, intraperitoneal, intralesional, intranasal, rectal, vaginal, by inhalation, via an implanted reservoir, or via a transdermal patch, and the like. Administration includes self-administration and the administration by another.
- beneficial agent and “active agent” are used interchangeably herein to refer to a chemical compound or composition that has a beneficial biological effect.
- beneficial biological effects include both therapeutic effects, i.e., treatment of a disorder or other undesirable physiological condition, and prophylactic effects, i.e., prevention of a disorder or other undesirable physiological condition.
- the terms also encompass pharmaceutically acceptable, pharmacologically active derivatives of beneficial agents specifically mentioned herein, including, but not limited to, salts, esters, amides, prodrugs, active metabolites, isomers, fragments, analogs, and the like.
- “Complementary” or “substantially complementary” refers to the hybridization or base pairing or the formation of a duplex between nucleotides or nucleic acids, such as, for instance, between the two strands of a double stranded DNA molecule or between an oligonucleotide primer and a primer binding site on a single stranded nucleic acid.
- Complementary nucleotides are, generally, A and T/U, or C and G.
- Cas9 recognizes a short motif in the CRISPR repeat sequences (the PAM or protospacer adjacent motif) to help distinguish self versus non-self.
- Cas9 nuclease sequences and structures are well known to those of skill in the art (see, e.g., “Complete genome sequence of an Ml strain of Streptococcus pyogenes.” Ferretti et al., J. J., McShan W. M., Ajdic D. J., Savic D. J., Savic G., Lyon K., Primeaux C., Sezate S., Suvorov A. N., Kenton S., Lai H. S., Lin S. P., Qian Y., Jia H.
- an “effective amount” of an agent can also refer to an amount covering both therapeutically effective amounts and prophylactically effective amounts.
- An “effective amount” of an agent necessary to achieve a therapeutic effect may vary according to factors such as the age, sex, and weight of the subject. Dosage regimens can be adjusted to provide the optimum therapeutic response. For example, several divided doses may be administered daily or the dose may be proportionally reduced as indicated by the exigencies of the therapeutic situation.
- Encoding refers to the inherent property of specific sequences of nucleotides in a polynucleotide, such as a gene, a cDNA, or an mRNA, to serve as templates for synthesis of other polymers and macromolecules in biological processes having either a defined sequence of nucleotides (i.e., rRNA, tRNA and mRNA) or a defined sequence of amino acids and the biological properties resulting therefrom, Thus, a gene encodes a protein if transcription and translation of mRNA occurs.
- a polynucleotide such as a gene, a cDNA, or an mRNA
- an expression cassette refers to a nucleic acid construct, which when introduced into a host cell, results in transcription and/or translation of a RNA or polypeptide, respectively.
- an expression cassette comprising a promoter operably linked to a second nucleic acid (e.g. polynucleotide) may include a promoter that is heterologous to the second nucleic acid (e.g.
- an expression cassette comprising a terminator (or termination sequence) operably linked to a second nucleic acid may include a terminator that is heterologous to the second nucleic acid (e.g., polynucleotide) as the result of human manipulation.
- the expression cassette comprises a promoter operably linked to a second nucleic acid (e.g., polynucleotide) and a terminator operably linked to the second nucleic acid (e.g., polynucleotide) as the result of human manipulation.
- the expression cassette comprises an endogenous promoter.
- the expression cassette comprises an endogenous terminator.
- the expression cassette comprises a synthetic (or non-natural) promoter.
- the expression cassette comprises a synthetic (or non-natural) terminator.
- fragments can include insertions, deletions, substitutions, or other selected modifications of particular regions or specific amino acids residues, provided the activity of the fragment is not significantly altered or impaired compared to the nonmodified peptide or protein. These modifications can provide for some additional property, such as to remove or add amino acids capable of disulfide bonding, to increase its bio-longevity, to alter its secretory characteristics, etc.
- gene refers to the coding sequence or control sequence, or fragments thereof.
- a gene may include any combination of coding sequence and control sequence, or fragments thereof.
- a “gene” as referred to herein may be all or part of a native gene.
- a polynucleotide sequence as referred to herein may be used interchangeably with the term “gene”, or may include any coding sequence, non-coding sequence or control sequence, fragments thereof, and combinations thereof.
- the term “gene” or “gene sequence” includes, for example, control sequences upstream of the coding sequence.
- genetically engineered cell or “genetically modified cell” as used herein refers to a cell modified by means of genetic engineering.
- engineered or “modified” thereof may refer to one or more changes of nucleic acids, such as nucleic acids within the genome of an organism.
- engineered or “modified” may refer to a change, addition and/or deletion of a gene.
- Engineered cells or modified cells can also refer to cells that contain added, deleted, and/or changed genes.
- nucleic acids or polypeptide sequences refer to two or more sequences or subsequences that are the same or have a specified percentage of amino acid residues or nucleotides that are the same (i.e., about 60% identity, preferably 61%, 62%, 63%, 64%, 65%, 66%, 67%, 68%, 69%, 70%, 71%, 72%, 73%, 74%, 75%, 76%, 77%, 78%, 79%, 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or higher identity over a specified region when compared and aligned for maximum correspondence over a comparison window or designated region) as measured using a BLAST or BLAST 2.0 sequence comparison algorithms with default parameters described below, or by manual alignment and visual inspection (see,
- sequences are then said to be “substantially identical.”
- This definition also refers to, or may be applied to, the compliment of a test sequence.
- the definition also includes sequences that have deletions and/or additions, as well as those that have substitutions.
- the preferred algorithms can account for gaps and the like.
- identity exists over a region that is at least about 10 amino acids or 20 nucleotides in length, or more preferably over a region that is 10-50 amino acids or 20-50 nucleotides in length.
- percent (%) amino acid sequence identity is defined as the percentage of amino acids in a candidate sequence that are identical to the amino acids in a reference sequence, after aligning the sequences and introducing gaps, if necessary, to achieve the maximum percent sequence identity.
- Alignment for purposes of determining percent sequence identity can be achieved in various ways that are within the skill in the art, for instance, using publicly available computer software such as BLAST, BLAST-2, ALIGN, ALIGN-2 or Megalign (DNASTAR) software. Appropriate parameters for measuring alignment, including any algorithms needed to achieve maximal alignment over the full-length of the sequences being compared can be determined by known methods.
- sequence comparisons typically one sequence acts as a reference sequence, to which test sequences are compared.
- test and reference sequences are entered into a computer, subsequence coordinates are designated, if necessary, and sequence algorithm program parameters are designated.
- sequence algorithm program parameters Preferably, default program parameters can be used, or alternative parameters can be designated.
- sequence comparison algorithm then calculates the percent sequence identities for the test sequences relative to the reference sequence, based on the program parameters.
- HSPs high scoring sequence pairs
- T is referred to as the neighborhood word score threshold (Altschul et al. (1990) J. Mol. Biol. 215:403-410). These initial neighborhood word hits act as seeds for initiating searches to find longer HSPs containing them. The word hits are extended in both directions along each sequence for as far as the cumulative alignment score can be increased. Cumulative scores are calculated using, for nucleotide sequences, the parameters M (reward score for a pair of matching residues; always >0) and N (penalty score for mismatching residues; always ⁇ 0). For amino acid sequences, a scoring matrix is used to calculate the cumulative score.
- Extension of the word hits in each direction are halted when: the cumulative alignment score falls off by the quantity X from its maximum achieved value; the cumulative score goes to zero or below, due to the accumulation of one or more negative-scoring residue alignments; or the end of either sequence is reached.
- the BLAST algorithm parameters W, T, and X determine the sensitivity and speed of the alignment.
- the BLAST algorithm also performs a statistical analysis of the similarity between two sequences (see, e.g., Karlin and Altschul (1993) Proc. Natl. Acad. Set. USA 90:5873-5787).
- One measure of similarity provided by the BLAST algorithm is the smallest sum probability (P(N)), which provides an indication of the probability by which a match between two nucleotide or amino acid sequences would occur by chance.
- P(N) the smallest sum probability
- a nucleic acid is considered similar to a reference sequence if the smallest sum probability in a comparison of the test nucleic acid to the reference nucleic acid is less than about 0.2, more preferably less than about 0.01.
- “increased” or “increase” as used herein generally means an increase by a statically significant amount; for the avoidance of any doubt, “increased” means an increase of at least 10% as compared to a reference level, for example an increase of at least about 20%, or at least about 30%, or at least about 40%, or at least about 50%, or at least about 60%, or at least about 70%, or at least about 80%, or at least about 90% or up to and including a 100% increase or any increase between 10-100% as compared to a reference level, or at least about a 2-fold, or at least about a 3-fold, or at least about a 4-fold, or at least about a 5-fold or at least about a 10-fold increase, or any increase between 2-fold and 10-fold or greater as compared to a reference level so long as the increase is statistically significant.
- the terms “may,” “optionally,” and “may optionally” are used interchangeably and are meant to include cases in which the condition occurs as well as cases in which the condition does not occur.
- the statement that a formulation “may include an excipient” is meant to include cases in which the formulation includes an excipient as well as cases in which the formulation does not include an excipient.
- Nucleic acid is “operably linked” when it is placed into a functional relationship with another nucleic acid sequence or amino acid sequence.
- DNA for a presequence or secretory leader is operably linked to DNA for a polypeptide if it is expressed as a preprotein that participates in the secretion of the polypeptide;
- a promoter or enhancer is operably linked to a coding sequence if it affects the transcription of the sequence;
- a ribosome binding site is operably linked to a coding sequence if it is positioned so as to facilitate translation.
- “operably linked” means that the DNA sequences being linked are near each other, and, in the case of a secretory leader, contiguous and in reading phase.
- operably linked nucleic acids do not have to be contiguous. Linking is accomplished by ligation at convenient restriction sites. If such sites do not exist, the synthetic oligonucleotide adaptors or linkers are used in accordance with conventional practice.
- a promoter is operably linked with a coding sequence when it is capable of affecting (e.g., modulating relative to the absence of the promoter) the expression of a protein from that coding sequence (i.e., the coding sequence is under the transcriptional control of the promoter).
- promoter refers to a region or sequence determinants located upstream or downstream from the start of transcription and which are involved in recognition and binding of RNA polymerase and other proteins to initiate transcription. Promoters need not be of bacterial origin, for example, promoters derived from viruses or from other organisms can be used in the compositions, systems, or methods described herein.
- regulatory element is intended to include promoters, enhancers, internal ribosomal entry sites (IRES), and other expression control elements (e.g., transcription termination signals, such as polyadenylation signals and poly-U sequences).
- Regulatory elements include those that direct constitutive expression of a nucleotide sequence in many types of host cell and those that direct expression of the nucleotide sequence only in certain host cells (e.g., tissue-specific regulatory sequences).
- tissue-specific promoter may direct expression primarily in a desired tissue of interest, such as muscle, neuron, bone, skin, blood, specific organs, or particular cell types.
- Regulatory elements may also direct expression in a temporal-dependent manner, such as in a cell-cycle dependent or developmental stage-dependent manner, which may or may not also be tissue or cell-type specific.
- a vector comprises one or more pol III promoter (e.g., 1, 2, 3, 4, 5, or more pol I promoters), one or more pol II promoters (e.g., 1, 2, 3, 4, 5, or more pol II promoters), one or more pol I promoters (e.g., 1, 2, 3, 4, 5, or more pol I promoters), or combinations thereof.
- pol III promoters include, but are not limited to, U6 and Hl promoters.
- pol II promoters include, but are not limited to, the retroviral Rous sarcoma virus (RSV) LTR promoter (optionally with the RSV enhancer), the cytomegalovirus (CMV) promoter (optionally with the CMV enhancer) [see, e.g., Boshart et al, Cell, 41 :521-530 (1985)], the SV40 promoter, the dihydrofolate reductase promoter, the P-actin promoter, the phosphoglycerol kinase (PGK) promoter, and the EFla promoter.
- RSV Rous sarcoma virus
- CMV cytomegalovirus
- PGK phosphoglycerol kinase
- enhancer elements such as WPRE; CMV enhancers; the R-U5' segment in LTR of HTLV-I (Mol. Cell. Biol., Vol. 8(1), p. 466-472, 1988); SV40 enhancer; and the intron sequence between exons 2 and 3 of rabbit P-globin (Proc. Natl. Acad. Sci. USA., Vol. 78(3), p. 1527-31, 1981).
- WPRE WPRE
- CMV enhancers the R-U5' segment in LTR of HTLV-I
- SV40 enhancer SV40 enhancer
- the intron sequence between exons 2 and 3 of rabbit P-globin Proc. Natl. Acad. Sci. USA., Vol. 78(3), p. 1527-31, 1981.
- recombinant refers to a human manipulated nucleic acid (e.g., polynucleotide) or a copy or complement of a human manipulated nucleic acid (e.g., polynucleotide), or if in reference to a protein (i.e, a “recombinant protein”), a protein encoded by a recombinant nucleic acid (e.g., polynucleotide).
- a recombinant expression cassette comprising a promoter operably linked to a second nucleic acid (e.g., polynucleotide) may include a promoter that is heterologous to the second nucleic acid (e.g.
- a recombinant expression cassette may comprise nucleic acids (e.g., polynucleotides) combined in such a way that the nucleic acids (e.g., polynucleotides) are extremely unlikely to be found in nature.
- “Pharmaceutically acceptable” component can refer to a component that is not biologically or otherwise undesirable, i.e., the component may be incorporated into a pharmaceutical formulation of the invention and administered to a subject as described herein without causing significant undesirable biological effects or interacting in a deleterious manner with any of the other components of the formulation in which it is contained.
- the term When used in reference to administration to a human, the term generally implies the component has met the required standards of toxicological and manufacturing testing or that it is included on the Inactive Ingredient Guide prepared by the U.S. Food and Drug Administration.
- nucleic acid as used herein means a polymer composed of nucleotides, e.g., deoxyribonucleotides or ribonucleotides.
- nucleobase refers to the part of a nucleotide that bears the Watson/Crick base-pairing functionality.
- the most common naturally-occurring nucleobases, adenine (A), guanine (G), uracil (U), cytosine (C), and thymine (T) bear the hydrogen-bonding functionality that binds one nucleic acid strand to another in a sequence specific manner.
- ribonucleic acid and “RNA” as used herein mean a polymer composed of ribonucleotides.
- deoxyribonucleic acid and “DNA” as used herein mean a polymer composed of deoxyribonucleotides.
- polynucleotide refers to a single or double stranded polymer composed of nucleotide monomers.
- the term “prime editing system” involves a Cas9 nickase (for example, a Cas9 H840A nickase or a Cas9 R221K N394K H840A nickase) and a reverse transcriptase, in combination with a guide RNA (herein referred as “prime editing guide RNA” or “pegRNA”).
- the pegRNA is a sgRNA with a primer binding site (PBS) and a DNA synthesis template appended to the 3’ end containing the desired nucleic acid sequence.
- PBS primer binding site
- the primer binding site allows the 3’ end of a nicked DNA strand to hybridize to the pegRNA, while the RT template serves as a template for the synthesis of edited genetic information.
- the pegRNA encodes the new sequence and allows DNA synthesis to introduce the desired mutations.
- the prime editing systems and pegRNAs are those described in U.S. Patent Nol, 1447,770, which is incorporated herein by reference in its entirety.
- the pegRNA comprises an engineered pegRNA (epegRNA).
- a pegRNA can be longer than standard sgRNAs commonly used for CRISPR gene editing.
- the pegRNA disclosed herein can be at least 60 nt, 70 nt, 80 nt, 90 nt, 100 nt, 120 nt, 140 nt, 160 nt, 180 nt, 200 nt, 250 nt, 300 nt, 350 nt, 400 nt, or 500 nt in length.
- guide RNA refers to the polynucleotide sequence comprising the guide sequence, the tracrRNAand the crRNA.
- guide sequence refers to the about 20 bp sequence within the guide RNA that specifies the target site and may be used interchangeably with the term “guide” or “spacer”.
- a “crRNA” is a bacterial RNA that confers target specificity and requires tracrRNA to bind to Cas9.
- a “tracrRNA” is a bacterial RNA that links the crRNA to the Cas9 nuclease and typically can bind any crRNA.
- the sequence specificity of a Cas DNA-binding protein is determined by gRNAs, which have nucleotide base-pairing complementarity to target DNA sequences.
- the pegRNA comprises a 3 ’-polyuridine domain (which comprises, for example, at least 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 25, 30, 35, 40, 45, 50, 60, 70, 80, 90, or 100 uridines).
- the 3 ’-polyuridine domain can locate at the 3’- end of the pegRNA or near the 3’ - end of the pegRNA.
- the 3 ’-polyuridine domain comprises at least one chemically modified uridine (for example, at least one, two, three, four, or five chemically modified uridines).
- the 3’- polyuridine domain comprises at least one unmodified uridine (for example, at least one, two, three, four, or five unmodified uridines).
- the 3 ’-polyuridine domain comprises at least one chemically modified uridine (for example, at least one, two, three, four, or five chemically modified uridines) and at least one unmodified uridine (for example, at least one, two, three, four, or five unmodified uridines).
- the at least one unmodified uridine locates at the 3’ end of the pegRNA.
- the pegRNA disclosed herein can have two unmodified uridines locating at its 3’ end, downstream of at least one, two, three, four, or five chemically modified uridines.
- the 3 ’-polyuridine domain comprises the sequence UU*mU*mU*mUU.
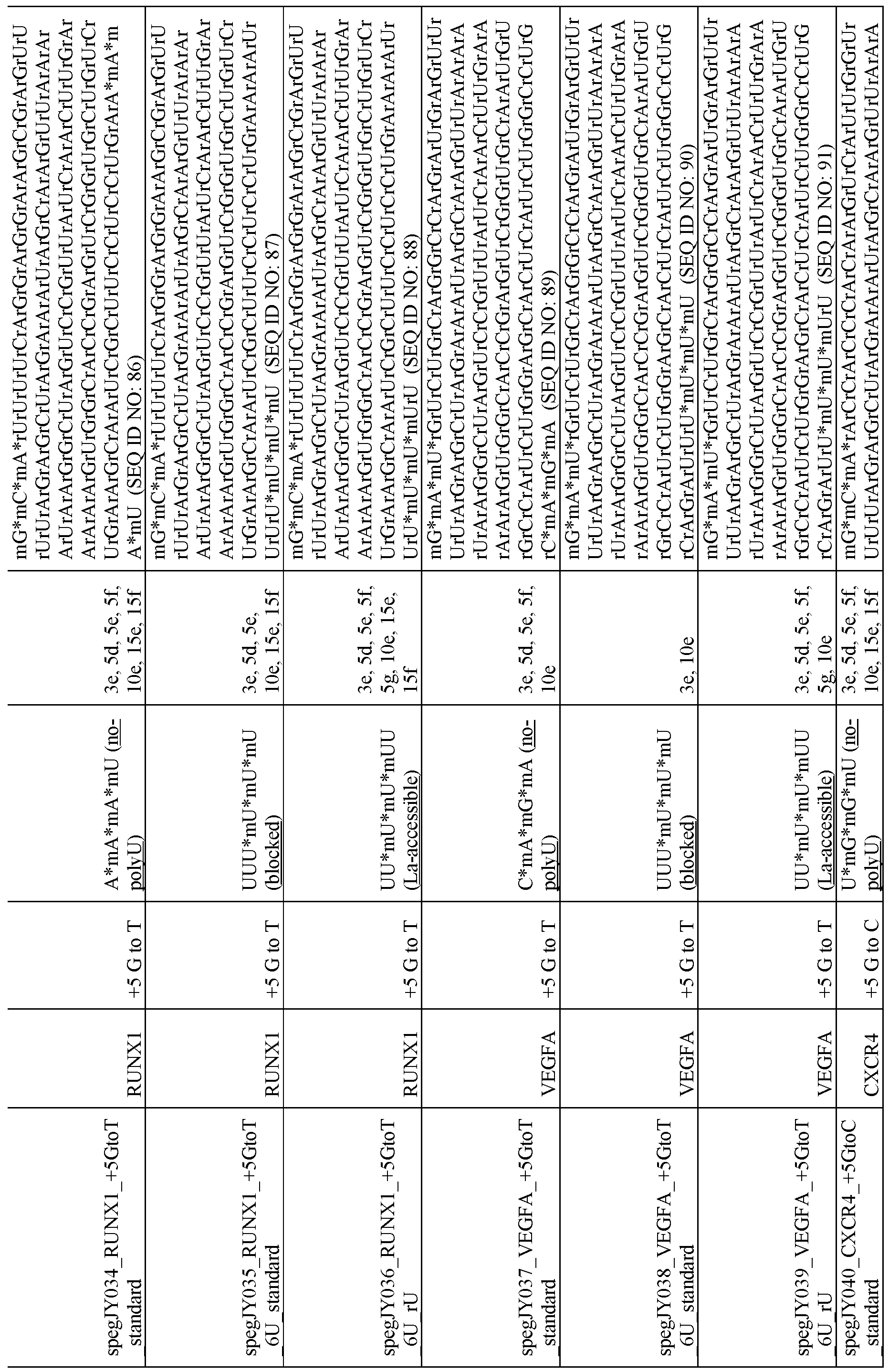
- the 3’- polyuridine domain comprises a 3’ sequence fragment selected from the sequences in Table 3.
- the 3 ’-polyuridine domain comprises a 3’ sequence fragment selected from SEQ ID NOs: 63-108.
- the chemical modification is 2’ -O-m ethylation and/or replacement of a phosphodiester bond to a phosphorothioate bond.
- the 3 ’-polyuridine domain comprises at least one (for example, at least two, three, four, or five) uridine with unmodified 2'-hydroxyl (OH) group.
- the at least one uridine with unmodified 2'- OH group locates at the 3’ end of the pegRNA.
- the SSB protein comprises a La motif and/or an RNA recognition motif (RRM) (e.g., amino acid residues 1-194 or 2-194 of SEQ ID NO: 34).
- RRM RNA recognition motif
- the SSB protein comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 33 or a fragment thereof.
- the SSB protein comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 34 or a fragment thereof.
- the SSB protein comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical SEQ ID NO: 35 or a fragment thereof. In some embodiments, the SSB protein comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 36 or a fragment thereof. In some embodiments, the Cas9 nickase of the system disclosed herein comprises a sequence at least 80% identical to SEQ ID NO: 26, or a fragment thereof. In some embodiments, the Cas9 nickase of the system disclosed herein comprises a sequence at least 80% identical to SEQ ID NO: 27, or a fragment thereof.
- the reverse transcriptase of the system disclosed herein comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 31 or a fragment thereof. In some embodiments, the reverse transcriptase of the system disclosed herein comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 32 or a fragment thereof.
- the Cas9 nickase is operatively linked to the reverse transcriptase (e.g., directly or through a linker).
- the SSB protein is operatively linked to the Cas9 nickase and the reverse transcriptase (e.g., directly or through one or more linkers).
- the system disclosed herein comprises a recombinant polypeptide that comprises the reverse transcriptase, the Cas9 nickase, and the SSB protein disclosed herein.
- the recombinant polypeptide can further comprise one or more linkers and/or one or more nuclear localization sequences (NLS) (e.g., aNLS of SV40 or c-Myc).
- the linker comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 28, 29, 30, or 40, or a fragment thereof.
- the NLS comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 25, 37, 38, 39, or 40 or a fragment thereof.
- the recombinant polypeptide comprising the reverse transcriptase, the Cas9 nickase, and the SSB protein comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to any of SEQ ID NOs: 1-12 or a fragment thereof. In some embodiments, the recombinant polypeptide comprising the reverse transcriptase, the Cas9 nickase, and the SSB protein comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 5 or a fragment thereof.
- the recombinant polypeptide comprising the reverse transcriptase, the Cas9 nickase, and the SSB protein comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 1 or a fragment thereof. In some embodiments, the recombinant polypeptide comprising the reverse transcriptase, the Cas9 nickase, and the SSB protein comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 2 or a fragment thereof.
- the recombinant polypeptide comprising the reverse transcriptase, the Cas9 nickase, and the SSB protein comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 3 or a fragment thereof. In some embodiments, the recombinant polypeptide comprising the reverse transcriptase, the Cas9 nickase, and the SSB protein comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 4 or a fragment thereof.
- the recombinant polypeptide comprising the reverse transcriptase, the Cas9 nickase, and the SSB protein comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 5 or a fragment thereof. In some embodiments, the recombinant polypeptide comprising the reverse transcriptase, the Cas9 nickase, and the SSB protein comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 6 or a fragment thereof.
- the recombinant polypeptide comprising the reverse transcriptase, the Cas9 nickase, and the SSB protein comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 7 or a fragment thereof. In some embodiments, the recombinant polypeptide comprising the reverse transcriptase, the Cas9 nickase, and the SSB protein comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 8 or a fragment thereof.
- the recombinant polypeptide comprising the reverse transcriptase, the Cas9 nickase, and the SSB protein comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 9 or a fragment thereof. In some embodiments, the recombinant polypeptide comprising the reverse transcriptase, the Cas9 nickase, and the SSB protein comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 10 or a fragment thereof.
- the recombinant polypeptide comprising the reverse transcriptase, the Cas9 nickase, and the SSB protein comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 11 or a fragment thereof. In some embodiments, the recombinant polypeptide comprising the reverse transcriptase, the Cas9 nickase, and the SSB protein comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 12 or a fragment thereof.
- a system comprising a Cas9 nickase; a reverse transcriptase; and a small RNA binding exonuclease protection factor La (SSB) protein.
- the Cas9 nickase, reverse transcriptase, and the SSB protein can be on a same or different polypeptide.
- the Cas9 nickase, reverse transcriptase, and the SSB protein can be on a same or different pharmaceutically acceptable carriers.
- a system comprising a first polynucleotide encoding a Cas9 nickase; a second polynucleotide encoding a reverse transcriptase; and a third polynucleotide encoding a small RNA binding exonuclease protection factor La (SSB) protein; and a prime editing guide RNA (pegRNA).
- SSB small RNA binding exonuclease protection factor La
- pegRNA prime editing guide RNA
- the third polynucleotide encodes a La motif and/or an RNA recognition motif (RRM) of the SSB protein.
- the third polynucleotide comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 50 or a fragment thereof.
- the third polynucleotide comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 49 or a fragment thereof.
- the first, second, and third polynucleotides are operatively linked thereby forming one recombinant polynucleotide.
- the system disclosed herein comprises a recombinant polynucleotide encoding a recombinant polypeptide that comprises the reverse transcriptase, the Cas9 nickase, and the SSB protein disclosed herein.
- the recombinant polypeptide can further comprise one or more linkers and/or one or more nuclear localization sequences (NLS) (e.g., a NLS of SV40 or c-Myc).
- NLS nuclear localization sequences
- the polynucleotide encoding the linker comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 44, 45, 46, or 56, or a fragment thereof.
- the polynucleotide encoding the NLS comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 41, 53, 54, 55, 56 or a fragment thereof.
- the recombinant nucleotide comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to any of SEQ ID NOs: 13-24 or a fragment thereof. In some embodiments, the recombinant nucleotide comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 17 or a fragment thereof.
- compositions disclosed herein may be in solution, suspension (for example, incorporated into microparticles (such as exosomes) or liposomes). These may be targeted to a particular cell type via antibodies, receptors, or receptor ligands.
- the compositions disclosed herein may be in exosomes.
- exosome refers to a cell-derived membranous vesicle. They refer to extracellular vesicles, which are generally of between 30 and 200 nm in size, for example in the range of 50-100 nm in size.
- the exosomes can be engineered to express one or more ligands or molecules for cell-targeting delivery.
- compositions may potentially be administered as a pharmaceutically acceptable acid- or base- addition salt, formed by reaction with inorganic acids such as hydrochloric acid, hydrobromic acid, perchloric acid, nitric acid, thiocyanic acid, sulfuric acid, and phosphoric acid, and organic acids such as formic acid, acetic acid, propionic acid, glycolic acid, lactic acid, pyruvic acid, oxalic acid, malonic acid, succinic acid, maleic acid, and fumaric acid, or by reaction with an inorganic base such as sodium hydroxide, ammonium hydroxide, potassium hydroxide, and organic bases such as mono-, di-, trialkyl and aryl amines and substituted ethanolamines.
- inorganic acids such as hydrochloric acid, hydrobromic acid, perchloric acid, nitric acid, thiocyanic acid, sulfuric acid, and phosphoric acid
- organic acids such as formic acid, acetic acid, propionic acid, glyco
- compositions required will vary from subject to subject, depending on the species, age, weight and general condition of the subject, the severity of the allergic disorder being treated, the particular nucleic acid or vector used, its mode of administration and the like. Thus, it is not possible to specify an exact amount for every composition. However, an appropriate amount can be determined by one of ordinary skill in the art using only routine experimentation given the teachings herein.
- dosing frequency for the composition disclosed herein includes, but is not limited to, no more than once every 30 years, every 25 years, every 20 years, every 15 years, every 10 years, every 5 years, every 4 years, every 3 years, every 2 years, every 12 months, or every 6 months.
- the interval between each administration is less than about 4 months, less than about 3 months, less than about 2 months, less than about a month, less than about 3 weeks, less than about 2 weeks, or less than less than about a week, such as less than about any of 6, 5, 4, 3, 2, or 1 day.
- the dosing frequency for the composition includes, but is not limited to, at least once a day, twice a day, or three times a day.
- the interval between each administration is less than about 48 hours, 36 hours, 24 hours, 22 hours, 20 hours, 18 hours, 16 hours, 14 hours, 12 hours, 10 hours, 9 hours, 8 hours, or 7 hours.
- the interval between each administration is less than about 24 hours, 22 hours, 20 hours, 18 hours, 16 hours, 14 hours, 12 hours, 10 hours, 9 hours, 8 hours, 7 hours, or 6 hours. In some embodiments, the interval between each administration is constant. For example, the administration can be carried out daily, every two days, every three days, every four days, every five days, or weekly. Administration can also be continuous and adjusted to maintaining a level of the compound within any desired and specified range. It should be understood and herein contemplated that the compositions disclosed herein can be used in combination with a pain reliever and, in some examples, reduce the dosing frequency of the pain reliever.
- the therapeutically effective amount typically will vary from about 0.001 mg/kg to about 1000 mg/kg, from about 0.01 mg/kg to about 750 mg/kg, from about 100 mg/kg to about 500 mg/kg, from about 1 mg/kg to about 250 mg/kg, from about 10 mg/kg to about 150 mg/kg in one or more dose administrations daily, for one or several days (depending of course of the mode of administration and the factors discussed above).
- Other suitable dose ranges include 1 mg to 10,000 mg per day, 100 mg to 10,000 mg per day, 500 mg to 10,000 mg per day, and 500 mg to 1,000 mg per day.
- the amount is less than 10,000 mg per day with a range of 750 mg to 9,000 mg per day.
- the doses of the compositions disclosed herein for gene editing in a cell or a subject is less (e.g., about 2-fold less, about 3-fold less, about 4-fold less, about 5-fold less, about 6-fold less, about 7-fold less, about 8-fold less, about 9-fold less, about 10- fold less, about 15-fold less, about 20-fold less, about 30-fold less, about 40-fold less, about 50-fold less, about 100-fold less, or about 1000-fold less) than the doses commonly known in the art for gene editing.
- Parenteral administration of the composition is generally characterized by injection.
- Injectables can be prepared in conventional forms, either as liquid solutions or suspensions, solid forms suitable for solution of suspension in liquid prior to injection, or as emulsions.
- a more recently revised approach for parenteral administration involves use of a slow release or sustained release system such that a constant dosage is maintained. See, e.g., U.S. Patent No. 3,610,795, which is incorporated by reference herein.
- Example 1 Genome-scale CRISPRi screens identify La (SSB) as a strong mediator of prime editing
- Prime editing allows precise modification of genomes.
- scalable prime editing reporters were developed and performed genome-scale CRISPR-interference screens. From these screens, a single factor emerged as the strongest mediator of prime editing: the small RNA-binding exonuclease protection factor La (SSB).
- La small RNA-binding exonuclease protection factor
- La binds polyuridine tracts at the 3' ends of RNA polymerase III transcripts and protects those transcripts from cellular exonucleases. Accordingly, functional interaction were observed between La and the 3' ends of polyuridylated prime editing guide RNAs. Guided by these insights, a strategy was developed to improve prime editing, namely fusing the RNA-binding, N-terminal domain of La to the prime editor protein PEmax. Application of the editor dramatically increased prime editing efficiencies. The results provide key insights into how prime editing components interact with the cellular environment and suggest general strategies for stabilizing exogenous small RNAs therein.
- Prime editing minimally consists of an engineered Cas9 protein (Cas9 H840A nickase fused to a reverse transcriptase) and a prime editing guide RNA (pegRNA) that specifies both the DNA target and the intended edit (FIG. 1 A).
- the editor protein binds its cognate guide RNA and, directed by the spacer sequence, finds a complementary DNA target. Once bound to the target, the editing complex nicks the displaced DNA strand and releases a 3' DNA end. This end can then hybridize to the 3' extension of the pegRNA and prime reverse transcription of the pegRNA-encoded edit, which is ultimately incorporated into the genome or removed by DNA mismatch repair (MMR).
- MMR DNA mismatch repair
- Prime editing Several features that impact prime editing have already been reported, including the expression, stability, localization and activity of editing components, as well as chromatin context of target loci. Additionally, demonstrating that mechanistic understanding can reveal avenues for technological improvement, Previous results showed that small prime edits can be installed with higher efficiency and precision when MMR is suppressed or evaded. Studies of prime editing to date, however, have been limited in focus, with inquiry restricted to optimization of editing components or examination of inferred cellular determinants (e.g., DNA repair). By interrogating prime editing with unbiased, genome-scale CRISPR-interference (CRISPRi) screens, an unanticipated mediator of prime editing was identified: the small RNA-binding protein La (SSB). Subsequent characterization of this factor, then showed how exploiting an interaction between La and pegRNAs can dramatically enhance prime editing.
- CRISPRi genome-scale CRISPR-interference
- Prime editing reporter system in which installation of an intended edit ‘switches on’ a reporter gene was developed (FIG. IB). By design, this system expresses a single bicistronic mRNA but, due to lack of a properly positioned start codon, produces only a constitutive marker protein (driven by an internal ribosome entry site) until an upstream, inframe ATG is edited into a defined target site to induce expression of a different reporter gene.
- the system was designed for use with an orthogonal Staphylococcus aureus Cas9 (SaCas9)-based prime editor (SaPE2).
- SaPE2 protospacers in the target site were included: one for ATG installation and another at which a +50 complementary strand nick can be introduced.
- Such nicks have been shown to enhance prime editing, and their inclusion, by use of additional single guide RNAs (sgRNAs), constitutes the PE3 approach.
- sgRNAs single guide RNAs
- Two versions of the reporter system were built: one that uses the fluorescent protein EGFP to report on editing and another that uses a synthetic cell surface protein (IgK-hlgGl-Fc- PDGFRP) (FIGS. 6A-6B).
- the gene products encoded by each of these reporters were chosen to allow efficient isolation of successfully edited, marker-positive cells: GFP through fluorescence-activated cell sorting (FACS reporter) and the surface protein via magnetic cell separation with protein G beads (MCS reporter).
- FACS reporter fluorescence-activated cell sorting
- MCS reporter magnetic cell separation with protein G beads
- Each of these reporters were transduced into K562 cells constitutively expressing CRISPRi machinery (K562 CRISPRi cells) and, to validate their performance, edited the resulting cells with substitution or insertion edits designed to install one or more start codons (FIG. 6C).
- the FACS reporter After editing, the FACS reporter produced a clear population of GFP+ cells (FIG. 1C). Two observations also demonstrated that that the percentage of those marker-positive cells faithfully reports intended prime editing efficiency: (1) perturbation of MSH2, an MMR gene known to suppress small substitution edits, increased GFP+ percentage (FIG. 1C) and (2) PE3-based editing, which is more efficient than PE2, showed higher GFP+ percentage (FIG. 6D). Additionally, confirming reporter accuracy, quantification of editing outcomes from GFP+ and GFP- populations of FACS reporter cells separated by flow cytometry, and from MCS reporter cells that either bound protein G beads or did not, revealed enrichment of intended edits in GFP+ and bead-bound cells, respectively (FIGS. 6E-6F).
- FACS reporter cells were transduced with the hCRISPRi-v2 library (18,905 targeted genes, 5 sgRNAs per gene), introduced prime editing components by plasmid transfection (SaPE2, +7 GG to CA pegRNA, +50 nicking sgRNA), and separated resulting GFP+/- populations.
- Flow cytometry analysis prior to sorting confirmed reasonable editing efficiencies (FIG. 6G) and sequencing of the target site showed expected enrichment in sorted populations (FIGS. 6H-6I).
- Example 2 La promotes prime editing across pegRNA and editor designs, programmed edits, endogenous targets, and cell types
- La a ubiquitously expressed eukaryotic protein, is involved in diverse aspects of RNA metabolism, but one of its most characterized roles is binding polyuridine (polyU) tracts at the 3' ends of nascent RNA polymerase III (Pol III) transcripts and protecting them from exonucleases.
- polyU polyuridine
- Pol III nascent RNA polymerase III
- the La phenotypes observed therefrom may represent an interaction between La and the pegRNA used for screening.
- pegRNAs and epegRNAs were tested.
- the latter contain structured motifs at their 3' ends and have been shown to enhance prime editing, with improvements loosely attributed to pegRNA stabilization.
- This difference fits a model wherein La promotes editing by interacting with the 3' ends of (e) pegRNAs but has a stronger effect on pegRNAs, which may be less stable or more accessible to La due to less structured 3’ ends.
- Example 3 The effect of La on prime editing does not extend to other editing modalities
- Prime editing relies on pegRNA 3' extensions, which encode intended edits, but other editing modalities such as nuclease-mediated gene disruption and base editing do not. This difference was a prompt to examine the effects of La on other genome editing approaches.
- SaCas9 was used to induce DNA double-strand breaks (DSBs) in the MCS prime editing reporter using the +7 GG to CA pegRNA, which targets SaCas9 to a locus near a transduced GFP marker gene but not directly within sequences required for expression (FIG. 2F). Because Cas9-induced DSBs often generate large deletions, such breaks can disrupt nearby reporter genes, even when those genes are distant from the target site and especially when targeting transduced lentiviral constructs.
- Example 4 La promotes prime editing by interacting with the 3' ends of polyuridylated pegRNAs
- La has been tenuously implicated in Pol Ill-mediated transcription, with phosphorylation of a single residue (S336) potentially involved in transcriptional modulation via Pol III recycling.
- S336 phosphorylation of a single residue
- La mutants were examined.
- La is a 408-residue protein consisting of a highly conserved La motif, two RNA recognition motifs (RRMs), and a flexible region with a nuclear localization signal (NLS) at the C-terminus (FIG. 3 A).
- the N-terminal domain of La (La 1-194 ), which contains the La motif and RRM1, is necessary and sufficient for high-affinity binding to 3' polyU, while regulation of Pol III recycling has been attributed to the phosphorylation status of Ser366 (S366). It was reasoned that if La promotes prime editing through transcription, truncation of the C-terminal domain or mutation of S366 could abolish or alter its impact, but if La promotes prime editing by binding to the 3' ends of pegRNAs, La 1-194 alone should be sufficient for that activity.
- Example 7 PE7 enhances prime editing of therapeutic-relevant targets and cell types
- 2.1E8 MCS reporter cells were transduced with hCRISPRi-v2 viruses at a 0.16 MOI (15% infection) for the screen conditions and were selected by 3 pg mL' 1 puromycin 48 hours after transduction. 7 days after transduction, 1E8 fully selected cells were nucleofected for each replicate of each edit with SE Cell Line 4D- Nucleofector X Kit L (Lonza V4XC-1024) and pulse code FF120, according to the manufacturer’s protocol. Each nucleofection consists of 1E7 cells and varying amounts of plasmids encoding prime editing components.
- PE3 7500 ng pCMV-SaPE2, 2500 ng +7 GG to CA pegRNA plasmid, 833 ng +50 nicking sgRNA plasmid (PE3) were used per nucleofection.
- PE4 and PE5 6000 ng pCMV-SaPE2, 3000 ng pEFla- hMLHldn (Addgene #174823), 2000 ng +7 GG to CA pegRNA plasmid and 667 ng +50 nicking sgRNA plasmid (PE5) were used. 4 days post nucleofection, cells from each replicate and condition were magnetically separated into bound and unbound fractions as previously described.
- the lysis buffer consisted of 10 mM Tris pH 8.0 (Gibco AM9855G), 0.05% SDS (Invitrogen 15553027), 25 pg mL 1 proteinase K (Invitrogen AM2546) and Nuclease-Free Water (AM9939).
- the genomic DNA extract was incubated at 37 °C for 90 minutes and transferred into PCR strips (USA Scientific 1402-4700) for 80°C inactivation of proteinase K for 30 minutes in Bio-Rad T100 Thermal Cycler.
- 1E6 cells were nucleofected with specified amounts of plasmids or synthetic guide RNAs using SE Cell Line 4D-Nucleofector X Kit S (Lonza V4XC-1032) and program FF-120, according to the manufacturer’s protocol.
- 900 ng pCMV-SaPE2 300 ng pegRNA plasmid, 100 ng nicking sgRNA plasmid (PE3/5) and 450 ng pEFla-hMLHldn (PE4/5) were nucleofected.
- 500 ng pegRNA plasmid was nucleofected.
- 500 ng pegRNA plasmid and 1000 ng plasmid encoding La, La mutants or mRFP control were nucleofected.
- 800 ng pX600 (Addgene #61592) and 400 ng +7 GG to CA pegRNA plasmid were nucleofected.
- Synthetic pegRNAs and nicking sgRNAs with specified sequences and modifications were ordered as Custom Alt-R gRNA from Integrated DNA Technologies (Table 3). According to an incremental titration of a DNMT1 +5 G to T synthetic pegRNA with standard chemical modifications in K562 PEmax parental cells, intended editing efficiencies were already saturated at 100 pmole input (FIG. 10B). Therefore, 100 pmole synthetic pegRNA and 50 pmole nicking sgRNA (PE3) were used for nucleofection unless otherwise specified.
- 1E6-2E6 cells were harvested in 1.5 mL tubes (Eppendorf 0030123611), washed with 1 mL DPBS (Gibco 14190144) and resuspended in 100 pL freshly prepared lysis buffer described above.
- the genomic DNA extract was incubated at 37 °C for 120 minutes and transferred into PCR strips (USA Scientific 1402-4700) for 80°C inactivation of proteinase K for 40 minutes in Bio-Rad T100 Thermal Cycler.
- 1E6 K562 and 1E5 U2OS cells were nucleofected with 1 pg editor mRNA and 50 pmole synthetic pegRNA using SE Cell Line 4D-Nucleofector X Kit S (Lonza V4XC-1032) with program FF-120 and DN-100, respectively, according to the manufacturer’s protocols. After nucleofection, cells were cultured for 72 hours and harvested for genomic DNA extract.
- cells were transduced with lentiviruses expressing (e)pegRNAs (20-40% infection) and were fully selected by 3 pg mL' 1 puromycin.
- Stably transduced HeLa and U2OS cells were nucleofected with 750 ng editor plasmid or 1 pg editor mRNA using SE Cell Line 4D-Nucleofector X Kit S (Lonza V4XC-1032) with program CN- 114 and DN-100, respectively, according to the manufacturer’s protocols. After nucleofection, cells were cultured for 72 hours and harvested for genomic DNA extract.
- K562 cells were transduced with lentiviruses expressing PEmax or PE7 (with IRES2-driven EGFP or EGFP-T2A-NeoR as selectable marker).
- the transduced populations (EGFP+, 20- 30%) were isolated by BD FACSAria Fusion Flow Cytometer 9 days post transduction, further transduced with lentiviruses expressing (e)pegRNAs (approximately 50% infection), fully selected by 3 pg mL' 1 puromycin and harvested 11 days after second transduction for genomic DNA extract.
- Genomic DNA sequences containing target sites were amplified through two rounds of PCR reactions (PCR1 and 2). In PCR1, genomic regions of interest were amplified with primers containing forward and reverse adapters for Illumina sequencing (Integrated DNA Technologies).
- Each PCR1 reaction consisted of 1 pL genomic DNA extract, 0.1 pL of each 100 pM forward and reverse primer (0.5 pM final concentration), 10 pL Phusion U Green Multiplex PCR Master Mix (Thermo Scientific F564L) and 8.8 pL Nuclease-Free Water (AM9939) and was performed with the following cycling conditions: 98 °C for 2 min, 28 cycles of [98 °C for 10 s, 61 °C for 20 s, and 72 °C for 30 s], followed by 72 °C for 2 min.
- PCR1 amplification was confirmed by 1% agarose (Goldbio A-201-100) gel electrophoresis before proceeding to PCR2 to uniquely index each sample with both forward and reverse Illumina barcoding primers.
- Each 14 pL PCR2 reaction consisted of 1 pL unpurified PCR 1 reaction, 0.5 pM of each forward and reverse Illumina barcoding primer, 7 pL Phusion U Green Multiplex PCR Master Mix (Thermo Scientific F564L) and Nuclease-Free Water (AM9939) and was performed with the following cycling conditions: 98 °C for 2 min, 9 cycles of [98 °C for 10 s, 61 °C for 20 s, and 72 °C for 30 s], followed by 72 °C for 2 min.
- the gel purified PCR2 products were quantified by Qubit IX dsDNA High Sensitivity kit (Invitrogen Q33231) and a high sensitivity DNA chip (Agilent Technologies 5067-4626) on an Agilent 2100 Bioanalyzer and sequenced with the MiSeq Reagent Micro Kit v2 300 cycles (Illumina MS- 103 -1002) or Nano Kit v2 300 cycles (Illumina MS-103-1001) with 300 cycles for R1 read, 8 cycles i7 index read and 8 cycles i5 index read.
- Sequencing reads were demultiplexed through HTSEQ (Princeton University High Throughput Sequencing Database) or bcl2fastq2 (Illumina) and sequencing adapters were trimmed using Cutadapt with the parameter “-a AGATCGGAAGAGCACACGTCTGAACTCCAGTCAC” (SEQ ID NO: 61).
- amplicon sequencing reads were aligned to corresponding reference sequences with CRISPResso2 in HDR batch mode using the intended editing outcome as the expected allele (-e) with the parameters “-q 30” and “- discard indel reads TRUE”.
- the CRISPResso2 quantification window was centered at the pegRNA nick (“-wc -3”) and the window size (“-w”) was set to 10 + the distance between nicks generated by the pegRNA and the nicking sgRNA.
- the same parameters were used for PE2, PE3, PE4 and PE5 conditions.
- the frequency of intended editing without indels was calculated as: (number of non-discarded HDR-aligned reads)/(number of reads that aligned all amplicons).
- the frequency of intended editing with indels was calculated as: (number of discarded HDR-aligned reads)/(number of reads that aligned all amplicons).
- the frequency of total intended editing (with or without indels) was calculated as (number of HDR-aligned reads)/(number of reads that aligned all amplicons).
- the frequency of total indels was calculated as: (number of discarded reads)/( number of reads that aligned all amplicons).
- the frequency of indels without intended editing was calculated as (number of discarded reference- aligned reads)/(number of reads that aligned all amplicons).
- the intended prime editing efficiencies referred to frequencies of intended editing without indels and the indel efficiencies referred to frequencies of total indels in this study unless otherwise specified.
- Each off-target amplicon sequence was compared to the 3' DNA flap sequence encoded by the pegRNA extension starting from the nucleotide 3 ' of Cas9 nick to the downstream until reaching the first nucleotide on the off-target amplicon that is different from the 3' DNA flap. Any reads with this nucleotide converted to that on the 3 ' DNA flap were considered off-target reads and the number of such reads can be found in the output file “Nucleotide frequency summary around sgRNA”. Off-target editing efficiencies were calculated as (number of off-target reads + number of indel-containing reads)/(number of reads that aligned all amplicons).
- CRISPResso2 was run in standard batch mode with the parameters “-q 30” and “-discard indel reads TRUE”.
- the intended editing efficiency referred to the frequency of indels which was calculated as (number of discarded reference- aligned reads)/ (number of reads that aligned all amplicons). Base editing outcomes were quantified by CRISPResso2 as previously described.
- Each 20 pL qPCR consists of 2 pL cDNA, 0.3 pM of each forward and reverse primer, 10 pL S YBR Green PCR Master Mix (Applied Biosystems 4309155) and Nuclease-Free Water (AM9939) and were performed in technical triplicate on a ViiA 7 Real-Time PCR System (Applied Biosystems) with the following cycling conditions: 50 °C for 2 min, 95 °C for 10 min, and 40 cycles of [95 °C for 15 s, 60 °C for 1 min].
- Relative La expression levels were calculated using the 2' AACT method with ACTB (a housekeeping gene) as the internal control in comparison to a non-targeting sgRNA or a non-targeting control siRNA pool.
- La knock-out K562 PEmax cells 122 pmole Alt-R S .p. Cas9 Nuclease V3 (Integrated DNA Technologies 1081058) and 200 pmole Alt-R CRISPR-Cas9 sgRNA targeting La (Integrated DNA Technologies Hs.Cas9.SSB. l.AA) were complexed for 20 minutes at room temperature and were nucleofected into 5E5 K562 PEmax parental cells using the SE Cell Line 4D-Nucleofector X Kit (Lonza V4XC-1032) and program FF-120, according to the manufacturer’s protocol.
- cells were sorted by BD FACSAria Fusion Flow Cytometer into 96-well plates at 1 cell per well with 150 pL conditioned culture medium. Single cells were grown and expanded for 2-3 weeks into clonal lines. Clones with high EGFP+ cell% according to AttueNXT flow cytometry analysis were selected for further characterization by targeted sequencing at genomic La locus and CRISPResso2 analysis.
- Antibodies were diluted in 5% Blotto (5% nonfat dry milk in TBST) and incubated with the membrane for 1 hour at room temperature.
- the following primary antibodies were used: anti-La mouse monoclonal antibody (1 :5000; Abeam ab75927); anti-GAPDH rabbit monoclonal antibody (1 :5000; Abeam abl81602); Guide-it Cas9 rabbit Polyclonal Antibody (1 : 1000; Takara 632607).
- the following secondary antibodies were used: HRP-conjugated sheep anti-mouse polyclonal antibody (1 :2000; VWR 95017-332) and HRP-conjugated donkey anti-rabbit polyclonal antibody (1 :2000; VWR 95017-556).
- the membrane was washed with TBST and immersed into Lumi-LightPLUS Western Blotting Substrate (Sigma 12015196001) for 3 minutes in dark prior to exposure with Azure Biosystems 600.
- the Restore Western Blot Stripping Buffer (Thermo Scientific 21059) was applied to strip the membrane before reprobing.
- Small RNA sequencing Small RNA sequencing.
- the small RNA sequencing (small RNA-seq) with targeting (e)pegRNAs was performed in triplicate and for each replicate, 5E6 K562 PEmax parental or La-ko4 cells were nucleofected with 2500 ng either one of the two (e)pegRNA plasmid sets (Set 1 and 2) using the SE Cell Line 4D-Nucleofector X Kit L (Lonza V4XC-1024) and pulse code FF120, according to the manufacturer’s protocol.
- Set 1 consists of plasmids encoding FANCF +5 G to T pegRNA, HEK3 +1 T to A pegRNA, DNMT1 +5 G to T pegRNA, RUNX1 +5 G to T epegRNA (evopreQi), VEGFA +5 G to T pegRNA and EMX1 +5 G to T epegRNA (mpknot).
- Set 2 consists of plasmids encoding RNF2 +1 C to A pegRNA, HEK3 +1 T to A epegRNA (mpknot), DNMT1 +5 G to T epegRNA (evopreQi), RUNX1 +5 G to T pegRNA, VEGFA +5 G to T pegRNA and EMX1 +5 G to T pegRNA.
- the VEGFA +5 G to T pegRNA plasmid was shared by both sets and served as the internal control for potential cross-set normalization.
- the FANCF +5 G to T pegRNA plasmid and the RNF2 +1 C to A pegRNA were specific to set 1 and 2 respectively.
- each set has the pegRNA plasmid while the other has the epegRNA plasmid encoding the same prime edit.
- Each set only had one evopreQi epegRNA plasmid and one mpknot epegRNA plasmid.
- the sets were formulated so that each (e)pegRNA transcript from cells nucleofected with one set can be aligned uniquely to the corresponding (e)pegRNA in that set, based on the observation in preliminary experiments that few fragments were solely mapped to the sgRNA scaffold shared by different (e)pegRNAs.
- RNA-seq with non-targeting mDNMTl +6 G to C pegRNA and epegRNA were performed in quadruplicate and for each replicate, 5E6 K562 PEmax parental or La-ko4 cells were nucleofected with 5000 ng (e)pegRNA plasmid using the SE Cell Line 4D-Nucleofector X Kit L (Lonza V4XC-1024) and pulse code FF120, according to the manufacturer’s protocol.
- a small RNA library was constructed with 1 pg total RNA as the input using NEBNext Multiplex Small RNA Library Prep Set for Illumina (Set 1) (New England Biolabs E7300S) and NEBNext Multiplex Oligos for Illumina Index Primers Set 3 (New England Biolabs E7710S) and Set 4 (New England Biolabs E7730S) according to the manufacturer’s protocol.
- K562 PEmax parental and La-ko4 cells were transduced with lentiviruses harboring the mDNMTl target.
- 1E6 each transduced cells were nucleofected with 500 or 1000 ng pegRNA or epegRNA plasmid using the SE Cell Line 4D-Nucleofector X Kit (Lonza V4XC-1032) and program FF-120, according to the manufacturer’s protocol. 14 the amount of Cells from each nucleofection were harvested 1, 2, 3 and 4 days after nucleofection and the editing outcomes were quantified by high-throughput DNA sequencing and CRISPResso2 analysis.
- the trimmed reads were then aligned to the sequence(s) of (e)pegRNA(s) the sample was nucleofected with, using Bowtie2 with default alignment options. Reads that did not align to the (e)pegRNA references were then aligned to the human genome (GRCh38 primary assembly from Ensembl release 107) using Bowtie2 with default alignment parameters. Downstream analysis of the alignments used only reads mapped in a proper pair, ensuring both ends of the sequenced fragment were properly mapped.
- Quantifications of human small RNA including assigning fragments to human transcripts, genes, and biotypes as well as counting, were performed on properly paired alignments using a custom Python script available in the GitHub repository. To distinguish between overlapping annotations, each aligned fragment was assigned to the annotation that most closely matched the start and end point of the fragment.
- the (e)pegRNA(s) were quantified for each sample by assigning each properly aligned fragment into one of three bins defined in the main text (c/.s-active, /ra/z.s-active and inactive) using Rsamtools and plyranges.
- Differential expression was calculated using DESeq2 version 1.38.3 with a design consisting of two covariates: (e)pegRNA plasmid set nucleofected (set 1 or 2) and cell line (K562 PEmax or La-ko4). Default parameters were used to estimate library size factors, genewise dispersion, and fitting of the negative binomial GLM to determine log2 fold change values. Log fold change shrinkage was performed using the apeglm algorithm. The default two-sided Wald test was used to determine the p values and the Bonferroni Holm method was used for multiple test correction. Coverage plots were generated using ggplot2 on data organized using the readr, dplyr, tidyr, and stringr packages.
- RNA-seq and data analysis. Each condition of RNA-seq was performed in quadruplicate and for each replicate, 1E6 K562 cells were nucleofected with 750 ng PEmax or PE7 editor plasmid and 250 ng pegRNA plasmid encoding HEK3 +1 T to A or PRNP + 6 G to T using SE Cell Line 4D-Nucleofector X Kit S (Lonza V4XC-1032) with program FF-120, according to the manufacturer’s protocols. Nucleofected cells were cultured in 6-well plates with 2.5 mL medium per well.
- Sequencing libraries were pooled, quantified by Qubit IX dsDNA High Sensitivity kit (Invitrogen Q33231) and a high sensitivity DNA chip (Agilent Technologies 5067-4626) on an Agilent 2100 Bioanalyzer and sequenced with the NovaSeq 6000 SP Reagent kit vl.5 100 cycles (Illumina 20028401) with 112 cycles for R1 read, 10 cycles index read.
- Sequencing reads were demultiplexed through HTSEQ (Princeton University High Throughput Sequencing Database). Alignment, quantification, and differential expression were performed using a Snakemake workflow and R scripts available on GitHub github.com/Princeton-LSI-ResearchComputing/PE-mRNA-seq-diffexp. The reads were aligned to the GRCh38 genome from Ensembl release 100 using STAR with default alignment parameters. Quantification was performed by STAR during alignment. Differential expression between editors was performed separately for each pegRNA. The standard DESeq2 procedure was performed to determine the differential expression between each editor within the set of samples for each pegRNA. Fold changes for lowly expressed genes were shrunken using the adaptive shrinkage estimator from the ashr package. Figures were generated using R packages ggplot2 and ggpubr.
- T cell isolation culture and prime editing.
- Human peripheral blood Leukopaks enriched for PBMCs were sourced from STEMCELL Technologies (catalog # 200-0092). No preference was given with regard to sex, gender, ethnicity or race.
- T cells were isolated with the Easy Sep Human T cell isolation kit (STEMCELL Technologies 100-0695) according to manufacturer's instructions. Immediately after isolation, T cells were used directly for in vitro experiments.
- T cells were cultured in complete X-VIVO 15 consisting of X- VI VO 15 (Lonza Bioscience 04-418Q) supplemented with 5% FBS (R&D systems), 4mM N-acetyl- cysteine (RPI A10040) and 55 pM 2-mercaptoethanol (GIbco 21985023).
- Pan CD3+ T cells were thawed and activated with anti-CD3/anti-CD28 dynabeads (Gibco 40203D) at a 1 : 1 bead:cell ratio in presence of 500 IU mL’ 1 IL-2.
- T cells were magnetically de-beaded and taken up in P3 buffer with supplement (Lonza Bioscience V4SP- 3096) at 37.5E6 cells mL' 1 .
- 1.5 pg PEmax or PE7 mRNA mixed with 50 pmole synthetic pegRNA (IDT) was added per 20pL cells, not exceeding 25 pL total volume per reaction.
- Cells were subsequently electroporated on a Lonza 4D Nucleofector using program DS-137.
- PCR was performed with 25 uL of eluted genomic DNA per sample in an 100 pL PCR reaction with KAPA HiFi HotStart ReadyMix (Roche 9420398001) with the following cycling conditions: 95 °C for 3 min, 28 cycles of [98 °C for 20 s, 63 °C for 15 s, and 72 °C for 60 s], followed by 72 °C for 2 min.
- PCR products were purified by SPRI selection (Beckman Coulter B23317) and 2 pL eluted product was used for 8 cycles of additional PCR with KAPA HiFi HotStart ReadyMix to add Illumina sequencing adapters and indices.
- the final PCR products were purified by SPRI selection, quantified with Qubit IX dsDNA High Sensitivity (HS) assay kit (Invitrogen Q33230), equimolarly pooled, and sequenced with the MiSeq Reagent Kit v2 300 cycles (Illumina MS-102-2002) with 300 cycles for R1 read, 8 cycles i7 index read and 8 cycles i5 index read. Sequencing data were demultiplexed using BaseSpace and analyzed by CRISPResso2.
- mRNA in vitro transcription template plasmids for HPSC experiments were constructed by cloning PEmax and PE7 into a previously described vector. mRNA was generated using HiScribe T7 High Yield RNA Synthesis Kit (New England Biolabs E2040S) and BbsI linearized plasmids as templates with UTP fully replaced by NkMethylpseudouridine-S '-triphosphate (TriLink Biotechnologies N-1081) and co-transcriptional capping by CleanCap AG (TriLink Biotechnologies N-7113).
- mRNA was purified using the Monarch RNA Cleanup kit (500 pg) (NEB T2050S), eluted in IDTE pH 7.5 (Integrated DNA Technologies 11-05-01-15) and quantified using Qubit RNA High Sensitivity (HS) Assay Kit (Invitrogen Q32852).
- Synthetic pegRNAs were ordered as Custom Alt-R gRNA from Integrated DNA Technologies and resuspended at 200 pM in IDTE pH 7.5.
- Cryopreserved human CD34 + HSPCs from mobilized peripheral blood of deidentified healthy donors were obtained from the Fred Hutchinson Cancer Research Center (Seattle, Washington).
- CD34 + HSPCs were cultured with X-Vivo-15 media supplemented with 100 ng mL' 1 human Stem Cell Growth Factor (SCF), 100 ng mL' 1 human thrombopoietin (TPO), and 100 ng mL' 1 recombinant human FMS-like Tyrosine Kinase 3 Ligand (Flt3-L).
- CD34 + HSPCs were thawed and cultured for 24 hours in the presence of cytokines prior to nucleofection.
- 2.5E5 CD34 + HSPCs were electroporated using the P3 Primary Cell X kit S (Lonza Bioscience V4SP-3096) according to manufacturer’s recommendations with 2000 ng PEmax or PE7 mRNA and 200 pmole synthetic pegRNA using pulse code DS-130. Genomic DNA was harvested 3 days post nucleofection with QuickExtract DNA Extraction Solution (LGC Biosearch Technologies QE09050) following manufacturer’s recommendations. Prime editing outcomes were quantified by high-throughput DNA sequencing and CRISPResso2 analysis as described earlier.
Landscapes
- Health & Medical Sciences (AREA)
- Life Sciences & Earth Sciences (AREA)
- Genetics & Genomics (AREA)
- Engineering & Computer Science (AREA)
- Chemical & Material Sciences (AREA)
- Biomedical Technology (AREA)
- Bioinformatics & Cheminformatics (AREA)
- Organic Chemistry (AREA)
- Zoology (AREA)
- Wood Science & Technology (AREA)
- Molecular Biology (AREA)
- General Engineering & Computer Science (AREA)
- Biotechnology (AREA)
- Microbiology (AREA)
- Biochemistry (AREA)
- General Health & Medical Sciences (AREA)
- Plant Pathology (AREA)
- Biophysics (AREA)
- Physics & Mathematics (AREA)
- Medicinal Chemistry (AREA)
- Cell Biology (AREA)
- Mycology (AREA)
- Micro-Organisms Or Cultivation Processes Thereof (AREA)
Abstract
The present disclosure relates to prime editing systems, compositions, and methods for altering target gene sequences and/or expression of target gene sequences.
Description
PRIME EDITING SYSTEM AND USES THEREOF
CROSS REFERENCE TO RELATED APPLICATIONS
This application claims priority to, and the benefit of, U.S. Provisional Patent Application No. 63/527,444, filed July 18, 2023, and U.S. Provisional Patent Application No. 63/611,931, filed December 19, 2023, each of which is incorporated by reference herein in its entirety.
STATEMENT REGARDING FEDERALLY SPONSORED RESEARCH
This invention was made with government support under Grant Nos. GM138167, HG009490, and CA072720 awarded by the National Institutes of Health. The government has certain rights in the invention.
REFERENCE TO SEQUENCE LISTING
The sequence listing submitted on July 18, 2024, as an .XML file entitled “11676- 002W01_ST26” created on July 12, 2024, and having a file size of 447,258 bytes is hereby incorporated by reference pursuant to 37 C.F.R. § 1.52(e)(5).
FIELD
The present disclosure relates to compositions and systems for genome editing and uses thereof.
BACKGROUND
Efforts to repurpose CRISPR-Cas systems have produced a suite of genome editing tools, including programmable nucleases, base editors, and prime editors. Use of these tools in research has expanded the understanding of genomes, gene function, and many biological processes; additionally, as an emerging class of clinical agents, these tools hold promise for addressing a host of unmet medical needs. Application of genome editing technologies and development of new approaches, however, have outpaced the knowledge of how existing approaches work. A compelling example is prime editing. Prime editing uses reverse transcription to install different types of edits (z.e., substitutions, small insertions, and deletions) into genomes with minimal unwanted mutational byproducts. Although a relatively new technology, the remarkable versatility and precision of prime editing have already motivated
many efforts to build enhanced prime editing systems, with intense focus on improving editing efficiency, which remains low and highly variable. While such efforts have been relatively successful, because each has to date considered only prime editing components or inferred determinants, much remains unknown about how the approach works and how interactions with the cellular environment promote or suppress prime editing processes. While many companies are working on using prime editing for therapeutic purposes, stability of pegRNA remains a problem. What is needed are new prime editing systems for enhancing pegRNA stability.
SUMMARY
The present disclosure shows modifying synthetic pegRNAs to allow SSB protein binding and overexpression of small RNA binding exonuclease protection factor La (SSB) protein can improve prime editing.
Accordingly, in some aspects, disclosed herein is a system comprising a Cas9 nickase; a reverse transcriptase; a small RNA binding exonuclease protection factor La (SSB) protein; and a prime editing guide RNA (pegRNA).
In some embodiments, the pegRNA comprises a 3 ’-polyuridine domain. In some embodiments, the 3 ’-polyuridine domain comprises at least one chemically modified uridine (for example, at least two, three, four, or five chemically modified uridines). In some embodiments, the 3 ’-polyuridine domain comprises at least one unmodified uridine (for example, at least two, three, four, or five unmodified uridines). In some embodiments, the 3’- polyuridine domain comprises at least one chemically modified uridine (for example, at least two, three, four, or five chemically modified uridines) and at least one unmodified uridine (for example, at least one, two, three, four, or five unmodified uridines). In some embodiments, the at least one unmodified uridine locates at the 3’ end of the pegRNA.
In some embodiments, the chemical modification is 2’ -O-m ethylation and/or replacement of a phosphodiester bond to a phosphorothioate bond.
In some embodiments, the 3 ’-polyuridine domain comprises at least one (for example, at least two, three, four, or five) uridine with unmodified 2'-hydroxyl (OH) group. In some embodiments, the at least one uridine with unmodified 2'- OH group locates at the 3’ end of the pegRNA.
In some embodiments, the SSB protein of the system of any preceding aspect comprises a sequence at least 80% identical to SEQ ID NO: 34 or a fragment thereof. In some embodiments, the SSB protein comprises a La motif and/or an RNA recognition motif (RRM) (e.g., amino acid residues 1-194 or 2-194 of SEQ ID NO: 34). In some embodiments, the SSB protein comprises a sequence at least 80% identical to SEQ ID NO: 33 or SEQ ID NO: 35 or a fragment thereof.
In some embodiments, the Cas9 nickase comprises a sequence at least 80% identical to SEQ ID NO: 26 or 27, or a fragment thereof.
In some embodiments, the SSB protein is operatively linked to the Cas9 nickase and the reverse transcriptase. Accordingly, in some embodiments, the system of any preceding aspect comprises a recombinant polypeptide, wherein the recombinant polypeptide comprises a sequence at least 80% identical to any of SEQ ID NOs: 1-12 or a fragment thereof.
Also disclosed herein is a system comprising a first polynucleotide encoding a Cas9 nickase; a second polynucleotide encoding a reverse transcriptase; and a third polynucleotide encoding a small RNA binding exonuclease protection factor La (SSB) protein; and a prime editing guide RNA (pegRNA).
In some embodiments, the third polynucleotide encodes a La motif and/or an RNA recognition motif (RRM) of the SSB protein. In some embodiments, the third polynucleotide comprises a sequence at least 80% identical to SEQ ID NO: 50 or a fragment thereof. In some embodiments, the third polynucleotide comprises a sequence at least 80% identical to SEQ ID NO: 49 or 51 or a fragment thereof.
In some embodiments, the first, second, and third polynucleotides are operatively linked thereby forming one recombinant polynucleotide. In some embodiments, the recombinant polynucleotide comprises a sequence at least 80% identical to any of SEQ ID NOs: 13-24. In some embodiments, the first, second, and third polynucleotides are located on a same or different vectors.
Also disclosed herein is a pharmaceutical composition comprising the system, polypeptides, or polynucleotides of any preceding aspect.
Also disclosed herein is a method of treating a genetic disorder in a subject in need, comprising administering to the subject a therapeutically effective amount of the system, polypeptides, or polynucleotides of any preceding aspect or a pharmaceutical composition of any preceding aspect.
Also disclosed herein is a method for altering expression of a gene product in a cell, comprising introducing into the cell an effective amount of the system, polypeptides, or polynucleotides of any preceding aspect.
BRIEF DESCRIPTION OF THE DRAWINGS
The accompanying figures, which are incorporated in and constitute a part of this specification, illustrate several aspects described below.
FIGS. 1A-1H show Genome-scale CRISPRi screens identify La as a determinant of prime editing. FIG. 1 A shows schematic of prime editing. FIG. IB shows a schematic of FACS reporter that expresses GFP upon installation of +7 GGto CA substitution prime edit (mCherry marker constitutively expressed). FIG. 1C shows flow cytometry analysis of GFP expression in K562 CRISPRi cells with integrated FACS reporter with and without prime editing (+7 GG to CA, PE3 with a +50 complementary strand nick) and with and without transduction of an MSH2-targeting sgRNA. FIG. ID shows the gene-level phenotypes from genome-scale CRISPRi screen performed in FACS reporter cells with +7 GGto CA edit using PE3 approach. Phenotypes represent enrichment of normalized sgRNA counts in GFP+ over GFP- populations after prime editing. Genes identified as hits using CRISPhieRmix (FDR < 0.01) and pseudogene controls generated from randomly selected non-targeting sgRNAs are indicated. FIG. IE shows the quantification of CRISPRi-mediated La depletion. RT-qPCR data collected from K562 CRISPRi cells with integrated MCS reporter. Data are normalized to ACTB and presented relative to a non-targeting sgRNA. FIG. IF and FIG. 1G show a comparison of prime editing outcomes using a pegRNA (left) or an epegRNA (right, tevopreQi) with the PE2 approach (plasmid delivery) at stably integrated FACS reporter in K562 CRISPRi cells after transduction of a La-targeting or non-targeting sgRNA. Intended editing quantified by flow cytometry (FIG. IF) or sequencing (FIG. 1G). Quantification of indels with and without the intended edit included directly adjacent to intended editing efficiency for each sample in FIG. 1G. FIG. 1H shows a comparison of prime editing outcomes with the indicated edit and approach (plasmid delivery) at integrated MCS reporter with and without depletion of La in K562 CRISPRi cells. Percentages of intended prime editing without indels (top), intended prime editing with indels (middle), and indels without the intended edit (bottom) plotted separately. Data and error bars in FIGS. 1F-1H indicate means and standard deviations (n=4, 4 and 3, respectively). Horizontal bars in FIG. IE indicate geometric means (n=3). Fold changes included for select comparisons in FIGS. 1F-1H. One-tailed unpaired Student’s /-test (FIGS. 1F-1H). * < 0.05.
FIGS. 2A-2H show that La promotes prime editing across edit types and genomic loci in multiple cell lines. FIG. 2A shows western blot analysis of K562 cells constitutively expressing PEmax (K562 PEmax cell line) and derived clones with genetic disruption of La (La-kol through La-ko5). Whole-cell lysates were sequentially immunoblotted for La, GAPDH, and prime editor protein (PEmax) with corresponding antibodies. Asterisks denote La knockout cell lines used in this study. FIG. 2B shows percentages of intended prime editing and indels at multiple genomic loci in K562 PEmax and La-ko4 cells. pegRNAs and epegRNAs (evopreQi) were delivered as plasmids without or with MLHldn (PE2 or PE4, respectively). PE4 was used for RNF2 and FANCF loci to ensure reasonable baseline editing in K562 PEmax cells. FIG. 2c shows percentages of intended prime editing and indels at the endogenous DNMT1 locus in K562 PEmax and La-ko4/5 cells with or without ectopic expression of La. Expression plasmids for La or an mRFP control were delivered alongside plasmids encoding pegRNA or epegRNA (evopreQi) specifying a +5 G to T edit. FIG. 2D shows quantification of siRNA-mediated La depletion. RT-qPCR data were collected from HEK293T cells at specified time points during prime editing. Data were normalized to ACTB and presented relative to a non-targeting siRNA pool. FIG. 2E shows fold changes in indicated editing outcomes across ten PE3 edits (substitutions, insertions, and deletions) at five genomic loci in HEK293T cells with or without La depletion by siRNAs. Editing components delivered by plasmid transfection. Editing percentages presented in FIG. 8F. FIG. 2F shows effects of La depletion on DSB repair. Schematic of the MCS reporter (top), with distances between predicted SaCas9 cut site and sequences required for GFP expression indicated. Flow cytometry analysis of MCS reporter cells with and without CRISPRi-mediated La depletion after DSB induction by SaCas9 and pegRNA encoding +7 GG to CA edit. Reporter also schematized in FIG. 6B. FIG. 2G shows fold changes in SaPE2, PE4 approach-, SaCas9-, SaBE4- and SaABE8e-induced editing outcomes in La-ko4 relative to parental controls (intended edits only). The same pegRNA expression plasmid was used for the editing systems at each of the four genomic loci. Editing percentages presented in FIGS. 9C-9F. FIG. 2H shows fold changes in SaCas9-, SaBE4-, and SaABE8e-induced editing outcomes in La-ko4 relative to parental controls (intended edits only). The same sgRNA expression plasmid was used for the editing systems at each of the four genomic loci. Editing percentages presented in FIGS. 9C-9F. Indel frequency for each sample included adjacent to corresponding intending editing efficiency in FIGS. 2B-2C. Data and error bars indicate means and standard deviations in b (n=4) and c (n=3). Horizontal bars in d indicate geometric means (n=3). Horizontal bars in e indicate medians of fold changes for individual replicates (n=4). Fold changes included for
select comparisons in FIGS. 2B-2C. One-tailed unpaired Student’s /-test (FIGS. 2B-2C). *P < 0.05.
FIGS. 3A-3J show that La functionally interacts with the 3' ends of polyuridylated pegRNAs. FIG. 3 A shows schematics of La domain architecture (top) and five La mutants used in FIG. 3B (bottom). FIG. 3B shows percentages of intended prime editing and indels at the endogenous DNMT1 locus with or without ectopic expression of La or La mutants in a. Expression plasmids (La, mutants, or mRFP control) were delivered to K562 PEmax and La- ko4 cells alongside plasmids encoding one pegRNA (+5 G to T edit). FIG. 3C shows the chemical structure of RNA with phosphorothioate bonds (*) and 2'-O-methylation modifications (m). FIG. 3D shows percentages of intended prime editing and indels at the endogenous DNMT1 locus in K562 PEmax and La-ko4 cells using 100 pmole of synthetic pegRNAs with indicated 3' end sequences and chemical modifications. FIG. 3E shows fold changes in average intended prime editing achieved at four genomic loci in La-ko4 cells relative to parental controls using 100 pmole synthetic pegRNAs with indicated no-polyU, blocked, or La-accessible end configurations. Editing percentages presented in FIG. 10E. FIG. 3F shows a model of La interaction with 3' ends of polyuridylated pegRNAs promotes prime editing. FIG. 3G shows a schematic of pegRNA specifying RUNX1 +5 G to T with the minimum sequence defining each class of small RNA-seq fragments highlighted (c/.s-active, orange; /ra//.s-active, purple). The edit-encoding nucleotide and cryptic terminators are also indicated (white base and green asterisks, respectively). FIGS. 3H-3I show coverage plots of small RNA-seq fragments aligned to pegRNA (h) or epegRNA (i) encoding RUNX1 +5 G to T. Data are from samples collected one day after (e)pegRNA plasmid transfection and normalized by counts of fragments from total human small RNA (top) or those within the corresponding bins (bottom). FIG. 3J shows a plot (MA) of small RNA-seq data displaying mean normalized expression versus log2-fold change in expression in La-ko4 relative to parental controls (n=3). Data are from samples collected one day after transfection of plasmids encoding seven pegRNAs and four epegRNAs. Alignment categories are indicated (human small RNA, gray; cv.s-active, orange; trans-active, purple; premature termination, green) and genes with adjusted /z-values < 0.05 are highlighted in light gray (calculated by DESeq2 using the Wald test). Indel frequency for each sample included adjacent to corresponding intending editing efficiency in FIGS. 3B-3D. Data and error bars in FIGS. 3B-3D indicate means and standard deviations (n=3). Fold changes included for select comparisons in d. One-tailed unpaired Student’s /-test (d). *P < 0.05, ns, not significant. Vertical bars in e indicate medians of ratios of means (n=3). In h and i, plotted data represent coverages of indicated bins (cis-
active, /ra/z.s-active or inactive) in specified cell lines (n=3). Nucleotide position 0 denotes the 5' end of the RNA, and positions of the edit-encoding nucleotide (vertical solid line) and the start of PBS (vertical dashed line) are indicated. Shaded areas represent sgRNA sequence, Pol III terminator for pegRNA, and linker plus evopreQi and Pol III terminator for epegRNA.
FIGS. 4A-4I show fusion of La RNA-binding N-terminal domain to PEmax improves prime editing. FIG. 4A shows architectures of prime editors. Medium gray NLS, bpNLSSV40; Dark gray NLS, NLSc‘Myc; A, 32 amino acid linker; B, 34 amino acid linker; C, SGGS linker; La, full length La or the N-terminal domain of La (La1-194); MMLV-RT, human codon- optimized MMLV-RT. FIG. 4B shows percentages of intended prime edits and indels produced with the indicated editors (from a) and pegRNA or epegRNA (evopreQi) at the endogenous DNMT1 and VEGFA loci in HEK293T and U2OS cells, respectively. Editing components (PE2) delivered by plasmid transfection. FIG. 4C shows percentages of intended prime editing and indels at the endogenous DNMT1 and VEGFA loci in HEK293T, HeLa, and U2OS cells. Editing components delivered by plasmid transfection. FIG. 4D shows percentages of intended prime editing and indels at eight endogenous loci in U2OS cells using pegRNAs or epegRNAs (mpknot: HEK3, tevopreQi: HEK4, evopreQi: all other loci). Editing components delivered by plasmid transfection. pegRNA data also presented in FIG. 15B. FIG. 4E shows percentages of intended prime editing and indels at the endogenous HEK3 locus in HEK293T cells. Editing components delivered by plasmid transfection. FIG. 4F shows fold changes in intended prime editing (left) and ratios of intended editing to indel frequency (right) for each indicated condition. Editing percentages presented in FIG. 4D. FIG. 4G shows a schematic of interactions between La N-terminal domain and RNA with 3 '-UUUOH. Four residues were mutated in PE7 mutant to disrupt 3' polyU binding (Q20, Y23, Y24 and F35; indicated in red). FIG. 4H shows a schematic of PE7 mutant harboring four mutations in La1-194 to disrupt 3’ polyU binding (Q20A, Y23A, Y24F and F35A; indicated by red lines). FIG. 41 shows percentages of intended prime edits and indels produced with PEmax, PE7 or PE7 mutant at the endogenous RUNX1 and VEGFA loci in U2OS cells. Editing components delivered by plasmid transfection. Indel frequency for each sample included adjacent to corresponding intending editing efficiency in FIGS. 4C-4E, and 41. Data in 4B indicate values of individual replicates (n = 9 for PEmax and 6 for PE7 with DNMT1 edit, and n=3 for others). Data and error bars in FIGS. 4C-4E and 41 indicate means and standard deviations (n=3). Horizontal bars in FIG. 4F indicate medians of ratios of means (n=3). Fold changes included for select comparisons in FIGS. 4B-4E, and 41. One-tailed unpaired Student’s /-test (FIGS. 4B-4E, and 41). *P < 0.05.
FIGS. 5A-5H show that PE7 enhances prime editing at disease-related targets and primary human cells. FIG. 5A, Percentages of intended prime editing and indels at six endogenous loci in U2OS cells using pegRNAs and epegRNAs (tevopreQi). Editing components delivered by plasmid transfection. pegRNA data also presented in FIG. 5C. FIG. 5B, Fold changes in intended prime editing for each indicated condition. Editing percentages presented in FIG. 5A. FIG. 5C, Summary plot of intended prime edit and indel frequencies produced with indicated editor and prime editing approaches at genomic loci. Data for PE2 and PE4 from six loci indicated in a plus HBG1/2. Data for PE3 and PE5 from a subset of those targets (HBB, PRNP, IL2RB, CXCR4). FIG. 5D, Percentages of intended prime editing and indels at four genomic loci in K562 cells using indicated editor and synthetic pegRNAs with no-polyU, blocked or La-accessible end configurations. FIG. 5E, Fold changes in average intended prime editing in K562 cells using PE7 mRNA relative to PEmax mRNA for synthetic pegRNAs with each indicated end configuration. Editing percentages presented in FIG. 5D. FIG. 5F, Percentages of intended prime editing and indels in primary human T cells using PEmax or PE7 mRNA and synthetic pegRNAs with indicated end configurations. FIG. 5G, Fold changes in intended prime editing in primary human T cells using PE7 mRNA and synthetic pegRNAs with La-accessible end configuration relative to intended editing with PEmax mRNA and the same pegRNAs at eight genomic loci. FIG. 5H, Percentages of intended prime editing and indels at endogenous HBB locus in primary HPSCs using PEmax or PE7 mRNA and synthetic pegRNAs with blocked or La-accessible end configuration. Indel frequency for each sample included adjacent to corresponding intending editing efficiency in FIGS. 5 A, 5C-5D, 5F and 5H. Data and error bars indicate means and standard deviations (n=3) in FIGS. 5A, 5D and 5H (3 different HPSC donors). Horizontal bars in FIG. 5C indicate medians of fold changes for individual replicates (n=3) with 99% confidence interval. Horizontal and vertical bars in FIGS. 5B-5E indicate medians of ratios of means (n=3). Data and error bars in FIG. 5F indicate means and standard deviations (n=6 and 2 different T cell donors for synthetic pegRNAs with La-accessible and blocked end configuration, respectively). Data in FIG. 5G indicate ratios of values for individual edits and donors (4 different T cell donors) and horizontal bar in FIG. 5G indicate median. Fold changes included for select comparisons in FIGS. 5A, 5D and 5F. One-tailed unpaired Student’s Ltest (d). *P < 0.05.
FIGS. 6A-6I show characterization of prime editing reporters before and during genome-scale CRISPRi screens. FIG. 6A shows a schematic of isolating prime edited cells with intended edit using the FACS reporter. The reporter expresses GFP upon installation of select prime edits, thus enabling separation of cells into mostly edited or mostly unedited
populations using flow cytometry. The complete FACS reporter is depicted in FIG. IB. FIG. 6B shows a schematic of isolating prime edited cells with intended edit using the MCS reporter. The reporter expresses a synthetic cell surface marker (IgK-hlgGl-Fc-PDGFRP) upon installation of select prime edits, thus enabling facile separation of cells into mostly edited or mostly unedited populations using magnetic Protein G beads. The complete MCS reporter is depicted in FIG. 2F. FIG. 6C shows three prime edits capable of ‘switching on’ the FACS and MCS reporters (depicted with the former). FIG. 6D shows flow cytometry analysis of GFP expression from the FACS reporter after prime editing with each of the edits depicted in FIG. 6C. FIG. 6E shows percentages of prime editing outcomes in GFP+ or GFP- cell populations sorted by flow cytometry after editing with each of the edits depicted in FIG. 6C. Outcomes quantified by sequencing the FACS reporter target site. FIG. 6F shows percentages of prime editing outcomes in bead-bound or unbound cell populations isolated by magnetic separation after editing with each of the edits depicted in c. Outcomes quantified by sequencing the MCS reporter target site. FIG. 6G shows flow cytometry analysis of GFP expression in the FACS reporter cells (ie., K562 CRISPRi cells with stably integrated FACS reporter) after transduction with genome-scale CRISPRi library (hCRISPRi-v2) and prime editing by plasmid transfection (+7 GG to CA, PE3). Data from repeat measurements of each replicate of the genome-scale screen. FIG. 6H shows percentages of prime editing outcomes observed in GFP+ or GFP- cell populations for each replicate of the genome-scale FACS screen. Outcomes quantified by sequencing the FACS reporter target site. FIG. 61 shows sequences and frequencies of alleles observed at the FACS reporter target site in cell populations sorted for replicate 1 of the genome-scale FACS screen. In FIGS. 6D-6F, prime editing components were delivered into K562 CRISPRi cells as expression plasmids after stable transduction of the FACS or MCS reporter. Data indicates means in FIG. 6E (n=l) and FIG. 6F (n=3).
FIGS. 7A-7I show results of genome-scale CRISPRi screens performed with FACS and MCS reporters. FIG. 7A, Pearson correlations of read counts per sgRNA between each pair of samples isolated from genome-scale FACS screen. FIG. 7B, sgRNA-level phenotypes from each replicate of the genome-scale FACS screen. FIG. 7C, Gene-level phenotypes (average of replicates) and per gene FDRs from the genome-scale FACS screen, as determined by CRISPhieRmix analysis. FIG. 7D, Pearson correlations of read counts per sgRNA between each pair of samples isolated from the genome-scale MCS screen performed with the PE3 approach. FIG. 7E, sgRNA-level phenotypes from each replicate of the genome-scale MCS screen performed with the PE3 approach. Compare to FIG. 7B for screen-to- screen differences in technical variability. FIG. 7F, Gene-level phenotypes (average of replicates) from genome-
scale FACS and MCS screens performed with the PE3 approach. FIGS. 7G-7I, Gene-level phenotypes from each replicate of MCS reporter screens performed with the PE3 (FIG. 7G), PE4 (FIG. 7H) and PE5 (FIG. 71) approaches. In FIGS. 7B-7E, sgRNAs targeting genes identified as hits (FDR < 0.01) using CRISPhieRmix are highlighted in red. In FIGS. 7C, 7F- 7G, genes identified as hits (FDR < 0.01) in the indicated screen using CRISPhieRmix are highlighted in red.
FIGS. 8A-8F show validation of La phenotypes with various genetic perturbation modalities. FIG. 8A shows a schematic of workflow used to engineer K562 clonal cell lines with PEmax expressed constitutively from the AAVS1 safe-harbor locus (K562 PEmax cell line). FIG. 8B shows sequences and frequencies of alleles observed at the La locus in the La- knockout clones used in this study (La-ko3 through La-ko5). FIG. 8C shows full images of western blot presented in FIG. 2 A. FIG. 8D shows cumulative population doublings of La-ko4 and La-ko5 cells compared to K562 PEmax parental cells. FIG. 8E shows flow cytometry analysis of GFP expressed from the PEmax construct at the AAVS1 locus in K562 PEmax parental, La-ko3, La-ko4 and La-ko5 cells prior to transfection in FIG. 2C. FIG. 8F shows percentages of intended prime editing and indels across ten edits with pegRNAs (top) or epegRNAs (bottom, mpknot: HEK3, evopreQi : all other loci) at five genomic loci in HEK293T cells with and without depletion of La by siRNAs. Prime editing components (PE3) were delivered as expression plasmids. Percentages in FIG. 8D indicate means and standard deviations (n=7) of daily cell doublings relative to K562 PEmax parental cells across 8-day time course. Indel frequency for each sample included adjacent to corresponding intending editing efficiency in FIG. 8F. Data and error bars in FIG. 8F indicate means and standard deviations (n=4). One-tailed unpaired Student’s Ltest (f). *P < 0.05, ns, not significant.
FIGS. 9A-9F show that La has a stronger impact on prime editing than other editing modalities. FIG. 9A shows flow cytometry analysis of GFP expression from a stably integrated MCS reporter in K562 CRISPRi cells after transduction of indicated sgRNAs and editing with SaCas9 nuclease. Editing components (SaCas9, +7 GG to CA pegRNA) delivered by plasmid transfection. FIG. 9B shows quantification of SaCas9-induced indels at stably integrated MCS reporter described in FIG. 9 A. FIGS. 9C-9F show percentages of intended editing achieved in K562 PEmax parental and La-knockout cells using SaPE2 with PE4 approach, SaCas9, SaBE4, and SaABE8e across four genomic loci, HEK3 (FIG. 9C), EMX1 (FIG. 9D), FANCF (FIG. 9E) and HBB (FIG. 9F). The same pegRNA or sgRNA expression plasmid was used for the editing systems at each target, with select combinations excluded (ie., SaPE2 with PE4 approach with any sgRNA and SaBE4 at EMX1). Data and error bars in FIGS. 9A-9F indicate means and
standard deviations (n=3). Two-tailed unpaired Student’s Z-test (FIGS. 9A-9B). ns, not significant.
FIGS. 10A-10E show prime editing with synthetic pegRNAs designed to block or allow La binding reveals functional interaction between La and polyuridylated 3' ends. FIG. 10A shows chemical structures of ribonucleotides linked by a phosphorothioate bond (left) or with substitution of ribose 2'-OH for 2'-O-methyl groups (2'-0Me) (right). FIG. 10B shows percentages of intended prime editing and indels at the endogenous DNMT1 locus in K562 PEmax parental cells using synthetic pegRNA with indicated 3' end configuration. Input was titrated from 0 to 500 pmole at 100 pmole increments. FIGS. 10C-10D shows percentages of intended prime editing and indels at the endogenous HEK3 (c, PE3) and DNMT1 (b, PE2) loci in K562 PEmax parental, La-ko4, and La-ko5 (where indicated) cells using 100 pmole of synthetic pegRNAs with specified 3' end sequences and chemical modifications. FIG. 10E shows percentages of intended prime editing and indels at endogenous DNMT1, CXCR4, VEGFA and RUNX1 loci in K562 PEmax parental and La-ko4 cells using 100 pmole of synthetic pegRNAs with indicated 3' end configurations. Indel frequency for each sample included adjacent to corresponding intending editing efficiency in FIGS. 10C-10E. Data and error bars in FIGS. 10B-10E indicate means and standard deviations (n=3). Fold changes included for select comparisons in FIGS. 10C-10E. One-tailed unpaired Student’s Z-test (c-e). *P < 0.05, ns, not significant.
FIGS. 11 A-l ID show details of small RNA-seq experiment performed with two sets of (e)pegRNAs. FIG. 11A shows composition of small RNA-seq libraries from K562 PEmax parental or La-ko4 cells. Data from samples collected one and two days after transfection of eleven (e)pegRNAs in two sets. FIG. 1 IB shows fold changes in normalized counts of indicated biotypes in La-ko4 cells relative to parental controls, from samples collected one and two days after transfection of eleven (e)pegRNAs in two sets. Counts were calculated per replicate independently for each set of (e)pegRNAs as the sum of properly aligned fragments classified as each biotype and normalized by total RNA counts. FIG. 11C shows a plot (MA) of small RNA-seq data displaying mean normalized expression versus log2-fold change in expression in La-ko4 cells relative to parental controls (n=3). Data are from samples collected two days after transfection of plasmids encoding seven pegRNAs and four epegRNAs. Alignment categories are indicated (human small RNA, gray; cv.s-active, orange; Z/z///.s-active, purple; premature termination, green) and genes with adjusted - values < 0.05 are highlighted in light gray (calculated by DESeq2 using the Wald test). FIG. 11D shows coverage plots of small RNA-seq fragments aligned to pegRNAs encoding VEGFA +5 G to T (left, n=6), FANCF +5
G to T (middle, n=3) or RNF2 +1 C to A (right, n=3). Data are from samples collected one day after pegRNA plasmid transfection and normalized by counts of fragments from total human small RNA (top) or those within the corresponding bins (bottom). Plotted data represent coverages of indicated bins (c/.s-active, /ra/z.s-active or inactive) in specified cell lines. Nucleotide position 0 denotes the 5' end of the RNA, and positions of the edit-encoding nucleotide (vertical solid line) and the start of PBS (vertical dashed line) are indicated. Shaded areas represent sgRNA sequence and Pol III terminator. Data indicate means in a (n=3). Horizontal bars in FIG. 1 IB indicate medians.
FIGS. 12A-12C show additional coverage plots of (e)pegRNAs from small RNA-seq experiment performed with two sets of (e)pegRNAs. FIGS. 12A-12C, Coverage plots of small RNA-seq fragments aligned to pegRNA (left) or epegRNA (right) encoding EMX1 +5 G to T (FIG. 12 A), HEK3 +1 T to A (FIG. 12B) or DNMT1 +5 G to T (FIG. 12C). Data are from samples collected one day after (e)pegRNA plasmid transfection and normalized by counts of fragments from total human small RNA (top) or those within the corresponding bins (bottom). Plotted data represent coverages of indicated bins (cv.s-active, /ra/z.s-active or inactive) in specified cell lines (n=3). Nucleotide position 0 denotes the 5' end of the RNA, and positions of the edit-encoding nucleotide (vertical solid line) and the start of PBS (vertical dashed line) are indicated. Shaded areas represent sgRNA sequence, Pol III terminator for pegRNAs, and linker plus evopreQi/mpknot and Pol III terminator for epegRNAs.
FIGS. 13A-13E show details of small RNA-seq experiment performed with nontargeting pegRNA and epegRNA, each specifying a +6 G to C edit in the Mus musculus DNMT1 gene. FIG. 13A shows the composition of small RNA-seq libraries from K562 PEmax parental or La-ko4 cells. Data from samples collected one and two days after transfection of plasmids encoding a pegRNA or an epegRNA specifying mouse DNMT1 +6 Gto C. FIG. 13B shows fold changes in normalized counts of indicated biotypes in La-ko4 cells relative to parental controls, from samples collected one and two days after transfection of plasmids encoding a pegRNA or an epegRNA specifying mouse DNMT1 +6 G to C. Counts were calculated per replicate independently for the pegRNA and the epegRNA as the sum of properly aligned fragments classified as each biotype and normalized by total RNA counts. FIGS. 13C-13D show coverage plots of small RNA-seq fragments aligned to the pegRNA (left) or the epegRNA (right) specifying mouse DNMT1 +6 G to C edit. Data are from cells without the (e)pegRNA target collected one (FIG. 13C) and two (FIG. 13D) days after (e)pegRNA plasmid transfection and normalized by counts of fragments from total human small RNA (top) or those within the corresponding bins (bottom). Plotted data represent coverages of indicated
bins (c/.s-active, /ra/z.s-active or inactive) in specified cell lines (n=4). Nucleotide position 0 denotes the 5' end of the RNA, and positions of the edit-encoding nucleotide (vertical solid line) and the start of PBS (vertical dashed line) are indicated. Shaded areas represent sgRNA sequence, Pol III terminator for pegRNA, and tevopreQi plus Pol III terminator for epegRNA. FIG. 13E show percentages of intended prime editing and indels in K562 PEmax parental and La-ko4 cells transduced with lentiviruses harboring mouse DNMT1 target and transfected with a plasmid encoding a pegRNA or epegRNA specifying mouse DNMT1 +6 G to C. Data are from samples collected on indicated days. Data indicates means in a (n=4). Horizontal bars in FIG 13B indicate medians. Data and error bars in FIG. 13E indicate means and standard deviations (n=3).
FIGS. 14A-14I show PE7 has no or negligible effects on cell viability, cell growth, and mRNA abundance compared to PEmax and PE7 mutant. FIG. 14A shows percentages of intended prime editing and indels at the endogenous HEK3 and PRNP loci in K562 cells with PEmax, PE7 or PE7 mutant. Editing components delivered by plasmid transfection. Cells from this experiment were used for analyses in FIGS. 14B-14I. FIG. 14B shows percentages of viable K562 cells quantified by flow cytometry one, two and three days after transfection of PEmax, PE7 or PE7 mutant editor plasmid and pegRNA plasmid specifying either HEK3 +1 T to A or PRNP +6 G to T. FIG. 14C shows cumulative population doublings of K562 cells two and three days after transfection of PEmax, PE7 or PE7 mutant editor plasmid and pegRNA plasmid specifying either HEK3 +1 T to A or PRNP +6 G to T. FIGS. 14D-14F shows a plot (MA) of RNA-seq data (n=4) displaying mean normalized gene expression versus log2-fold change in gene expression from K562 cells edited with PE7 relative to PEmax (FIG. 14D), PE7 relative to PE7 mutant (FIG. 14E), and PEmax relative to PE7 mutant (FIG. 14F). Analyses were performed with cells edited using two different pegRNAs, one specifying HEK3 +1 T to A (top) and one specifying PRNP +6 G to T (bottom). Upregulated and downregulated genes with adjusted /i-values < 0.05 (calculated by DESeq2 using the Wald test) are highlighted in red and blue respectively. FIGS. 14G-14I shows venn diagrams of differentially expressed genes (p < 0.05) in K562 cells edited at two different loci across three comparisons: PE7 relative to PEmax (FIG. 14g), PE7 relative to PE7 mutant (FIG. 14H), and PEmax relative to PE7 mutant (FIG. 141). Indel frequency for each sample included adjacent to corresponding intending editing efficiency in a. Data and error bars in FIG. 14A indicate means and standard deviations (n=4). Horizontal bars in FIGS. 14B-14C indicate means (n=4). Fold changes included for select comparisons in FIG. 14 A. One-tailed unpaired Student’s /-test (FIG. 14 A) and one-way ANOVA (FIG. 14C). *P < 0.05, ns, not significant.
FIGS. 15A-15G show PE7 improves prime editing with different approaches and delivery strategies without substantially increasing off-target effect. FIG. 15A shows percentages of editing outcomes produced by PEmax or PE7 with the PE2 approach at on- and off-target sites using pegRNAs targeting the EMX1 (left), FANCF (middle left), HEK3 (middle right), and HEK4 (right) loci in U2OS cells. On-target editing data also presented in FIG. 15B and FIG. 4D. FIG. 15B shows a summary plot of intended prime edit and indel frequencies observed at genomic loci with indicated editor and prime editing approaches. Data for PE2 and PE4 from eight loci indicated in FIG. 4D. Data for PE3 and PE5 from a subset of those targets (RNF2, HEK3, DNMT1 and VEGFA). FIG. 15C shows percentages of intended prime editing and indels at endogenous HEK3 (top) and DNMT1 (bottom) loci after lentiviral transduction of pegRNAs or (e)pegRNAs (tevopreQi) and transfection of PEmax or PE7 editor encoded on mRNA or plasmid in HeLa (left) and U2OS (right) cells. (e)pegRNAs use a modified sgRNA scaffold. FIG. 15D shows percentages of intended prime editing and indels at endogenous DNMT1 (left) and HEK3 (right) loci after lentiviral transduction of editing components in K562 cells. Two different editor expression constructs (as indicated) were tested. (e)pegRNAs use a modified sgRNA scaffold and epegRNAs use tevopreQi. FIG. 15E shows percentages of intended prime editing and indels at three genomic loci in U2OS cells using indicated editor mRNA and synthetic pegRNAs with no-polyU, blocked or La-accessible end configurations. FIG. 15F shows fold changes in average intended prime editing in U2OS cells using PE7 mRNA relative to PEmax mRNA for synthetic pegRNAs with each indicated end configuration. Editing percentages presented in FIG. 15E. FIG. 15G shows percentages of intended prime editing and indels at five genomic loci in primary human T cells using PEmax or PE7 mRNA and synthetic pegRNAs with La-accessible end configuration. Indel frequency for each sample included adjacent to corresponding intending editing efficiency in FIGS. 15B, 15E and 15G. Data and error bars indicate means and standard deviations in FIGS. 15A (n=3), 15E (n=3) and 15G (n=6 different T cell donors). Horizontal bars in FIG. 15B indicate medians of fold changes for individual replicates (n=3) with 99% confidence interval. Data in FIGS. 15C-15D indicate values of individual replicates (n=3). Vertical bars in FIG. 15F indicate medians of ratios of means (n=3). Fold changes included for select comparisons in FIGS. 15A and 15E. Two-tailed (FIG. 15 A) and one-tailed (FIG. 15E) Ltest. *P < 0.05, ns, not significant.
DETAILED DESCRIPTION
Reference will now be made in detail to the embodiments of the invention, examples of which are illustrated in the drawings and the examples. This invention may, however, be
embodied in many different forms and should not be construed as limited to the embodiments set forth herein.
Unless defined otherwise, all technical and scientific terms used herein have the same meaning as commonly understood to one of ordinary skill in the art to which this disclosure belongs. The term “comprising” and variations thereof as used herein is used synonymously with the term “including” and variations thereof and are open, non-limiting terms. Although the terms “comprising” and “including” have been used herein to describe various embodiments, the terms “consisting essentially of’ and “consisting of’ can be used in place of “comprising” and “including” to provide for more specific embodiments and are also disclosed. As used in this disclosure and in the appended claims, the singular forms “a”, “an”, “the”, include plural referents unless the context clearly dictates otherwise.
The following definitions are provided for the full understanding of terms used in this specification.
Terminology
The term “about” as used herein when referring to a measurable value such as an amount, a percentage, and the like, is meant to encompass variations of ±20%, ±10%, ±5%, or ±1% from the measurable value.
“Administration” to a subject includes any route of introducing or delivering to a subject an agent. Administration can be carried out by any suitable route, including oral, topical, intravenous, subcutaneous, transcutaneous, transdermal, intramuscular, intra-joint, parenteral, intra-arteriole, intradermal, intraventricular, intracranial, intraperitoneal, intralesional, intranasal, rectal, vaginal, by inhalation, via an implanted reservoir, or via a transdermal patch, and the like. Administration includes self-administration and the administration by another.
As used here, the terms “beneficial agent” and “active agent” are used interchangeably herein to refer to a chemical compound or composition that has a beneficial biological effect. Beneficial biological effects include both therapeutic effects, i.e., treatment of a disorder or other undesirable physiological condition, and prophylactic effects, i.e., prevention of a disorder or other undesirable physiological condition. The terms also encompass pharmaceutically acceptable, pharmacologically active derivatives of beneficial agents specifically mentioned herein, including, but not limited to, salts, esters, amides, prodrugs, active metabolites, isomers, fragments, analogs, and the like. When the terms “beneficial agent” or “active agent” are used, then, or when a particular agent is specifically identified, it is to be understood that the term includes the agent per se as well as pharmaceutically acceptable,
pharmacologically active salts, esters, amides, prodrugs, conjugates, active metabolites, isomers, fragments, analogs, etc.
The phrases "concurrent administration", "administration in combination", "simultaneous administration" or "administered simultaneously" as used herein, means that the compounds are administered at the same point in time or immediately following one another.
The phrase “codon optimized” as it refers to genes or coding regions of nucleic acid molecules for the transformation of various hosts, refers to the alteration of codons in the gene or coding regions of nucleic acid molecules to reflect the typical codon usage of a selected organism without altering the polypeptide encoded by the DNA. Such optimization includes replacing at least one, or more than one, or a significant number, of codons with one or more codons that are more frequently used in the genes of that selected organism.
“Complementary” or “substantially complementary” refers to the hybridization or base pairing or the formation of a duplex between nucleotides or nucleic acids, such as, for instance, between the two strands of a double stranded DNA molecule or between an oligonucleotide primer and a primer binding site on a single stranded nucleic acid. Complementary nucleotides are, generally, A and T/U, or C and G. Two single-stranded RNA or DNA molecules are said to be substantially complementary when the nucleotides of one strand, optimally aligned and compared and with appropriate nucleotide insertions or deletions, pair with at least about 80% of the nucleotides of the other strand, usually at least about 90% to 95%, and more preferably from about 98 to 100%. Alternatively, substantial complementarity exists when an RNA or DNA strand will hybridize under selective hybridization conditions to its complement. Typically, selective hybridization will occur when there is at least about 65% complementary over a stretch of at least 14 to 25 nucleotides, at least about 75%, or at least about 90% complementary. See Kanehisa (1984) Nucl. Acids Res. 12:203.
The term “Cas9” or “Cas9 nuclease” refers to an RNA-guided nuclease comprising a Cas9 protein, or a fragment thereof (e.g., a protein comprising an active, inactive, or partially active DNA cleavage domain of Cas9, and/or the gRNA binding domain of Cas9). A Cas9 nuclease is also referred to sometimes as a casnl nuclease or a CRISPR (clustered regularly interspaced short palindromic repeat)-associated nuclease. CRISPR is an adaptive immune system that provides protection against mobile genetic elements (viruses, transposable elements and conjugative plasmids). CRISPR clusters contain spacers, sequences complementary to antecedent mobile elements, and target invading nucleic acids. CRISPR clusters are transcribed and processed into CRISPR RNA (crRNA). In type II CRISPR systems correct processing of pre-crRNA requires a trans-encoded small RNA (tracrRNA), endogenous
ribonuclease 3 (rnc) and a Cas9 protein. The tracrRNA serves as a guide for ribonuclease 3- aided processing of pre-crRNA. Subsequently, Cas9/crRNA/tracrRNA endonucleolytically cleaves linear or circular dsDNA target complementary to the spacer. The target strand not complementary to crRNA is first cut endonucleolytically, then trimmed 3 ’-5’ exonucleolytically. In nature, DNA-binding and cleavage typically requires protein and both RNAs. However, single guide RNAs (“sgRNA”, or simply “gNRA”) can be engineered so as to incorporate aspects of both the crRNA and tracrRNA into a single RNA species. See, e.g., Jinek M., Chylinski K., Fonfara I., Hauer M., Doudna J. A., Charpentier E. Science 337:816- 821(2012), the entire contents of which is hereby incorporated by reference. Cas9 recognizes a short motif in the CRISPR repeat sequences (the PAM or protospacer adjacent motif) to help distinguish self versus non-self. Cas9 nuclease sequences and structures are well known to those of skill in the art (see, e.g., “Complete genome sequence of an Ml strain of Streptococcus pyogenes.” Ferretti et al., J. J., McShan W. M., Ajdic D. J., Savic D. J., Savic G., Lyon K., Primeaux C., Sezate S., Suvorov A. N., Kenton S., Lai H. S., Lin S. P., Qian Y., Jia H. G., Najar F. Z., Ren Q., Zhu H., Song L., White J., Yuan X., Clifton S. W., Roe B. A., McLaughlin R. E., Proc. Natl. Acad. Sci. U.S.A. 98:4658-4663(2001); “CRISPR RNA maturation by transencoded small RNA and host factor RNase III.” Deltcheva E., Chylinski K., Sharma C. M., Gonzales K., Chao Y., Pirzada Z. A., Eckert M. R., Vogel J., Charpentier E., Nature 471 :602- 607(2011); and “A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity.” Jinek M., Chylinski K., Fonfara I., Hauer M., Doudna J. A., Charpentier E. Science 337:816-821(2012), the entire contents of each of which are incorporated herein by reference). Cas9 orthologs have been described in various species, including, but not limited to, S. pyogenes and S. thermophilus. Additional suitable Cas9 nucleases and sequences will be apparent to those of skill in the art based on this disclosure, and such Cas9 nucleases and sequences include Cas9 sequences from the organisms and loci disclosed in Chylinski, Rhun, and Charpentier, “The tracrRNA and Cas9 families of type II CRISPR-Cas immunity systems” (2013) RNA Biology 10:5, 726-737; the entire contents of which are incorporated herein by reference.
The term “nickase” as used herein, refers to a nuclease that cleaves only a single DNA strand, either due to its natural function or because it has been engineered to cleave only a single DNA strand. Jinek et al., “A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity” Science 337(6096):816-821 (2012) and Cong et al. Multiplex genome engineering using CRISPR/Cas systems” Science 339(6121):819-823 (2013).
“Effective amount” of an agent refers to a sufficient amount of an agent to provide a desired effect. The amount of agent that is “effective” will vary from subject to subject, depending on many factors such as the age and general condition of the subject, the particular agent or agents, and the like. Thus, it is not always possible to specify a quantified “effective amount.” However, an appropriate “effective amount” in any subject case may be determined by one of ordinary skill in the art using routine experimentation. Also, as used herein, and unless specifically stated otherwise, an “effective amount” of an agent can also refer to an amount covering both therapeutically effective amounts and prophylactically effective amounts. An “effective amount” of an agent necessary to achieve a therapeutic effect may vary according to factors such as the age, sex, and weight of the subject. Dosage regimens can be adjusted to provide the optimum therapeutic response. For example, several divided doses may be administered daily or the dose may be proportionally reduced as indicated by the exigencies of the therapeutic situation.
"Encoding" refers to the inherent property of specific sequences of nucleotides in a polynucleotide, such as a gene, a cDNA, or an mRNA, to serve as templates for synthesis of other polymers and macromolecules in biological processes having either a defined sequence of nucleotides (i.e., rRNA, tRNA and mRNA) or a defined sequence of amino acids and the biological properties resulting therefrom, Thus, a gene encodes a protein if transcription and translation of mRNA occurs.
The term “expression cassette” or “vector” refers to a nucleic acid construct, which when introduced into a host cell, results in transcription and/or translation of a RNA or polypeptide, respectively. In embodiments, an expression cassette comprising a promoter operably linked to a second nucleic acid (e.g. polynucleotide) may include a promoter that is heterologous to the second nucleic acid (e.g. polynucleotide) as the result of human manipulation (e.g., by methods described in Sambrook et al., Molecular Cloning — A Laboratory Manual, Cold Spring Harbor Laboratory, Cold Spring Harbor, N.Y., (1989) or Current Protocols in Molecular Biology Volumes 1-3, John Wiley & Sons, Inc. (1994-1998)). In some embodiments, an expression cassette comprising a terminator (or termination sequence) operably linked to a second nucleic acid (e.g. polynucleotide) may include a terminator that is heterologous to the second nucleic acid (e.g., polynucleotide) as the result of human manipulation. In some embodiments, the expression cassette comprises a promoter operably linked to a second nucleic acid (e.g., polynucleotide) and a terminator operably linked to the second nucleic acid (e.g., polynucleotide) as the result of human manipulation. In some embodiments, the expression cassette comprises an endogenous promoter. In some
embodiments, the expression cassette comprises an endogenous terminator. In some embodiments, the expression cassette comprises a synthetic (or non-natural) promoter. In some embodiments, the expression cassette comprises a synthetic (or non-natural) terminator.
The “fragments,” whether attached to other sequences or not, can include insertions, deletions, substitutions, or other selected modifications of particular regions or specific amino acids residues, provided the activity of the fragment is not significantly altered or impaired compared to the nonmodified peptide or protein. These modifications can provide for some additional property, such as to remove or add amino acids capable of disulfide bonding, to increase its bio-longevity, to alter its secretory characteristics, etc.
The term "gene" or "gene sequence" refers to the coding sequence or control sequence, or fragments thereof. A gene may include any combination of coding sequence and control sequence, or fragments thereof. Thus, a "gene" as referred to herein may be all or part of a native gene. A polynucleotide sequence as referred to herein may be used interchangeably with the term "gene”, or may include any coding sequence, non-coding sequence or control sequence, fragments thereof, and combinations thereof. The term "gene" or "gene sequence" includes, for example, control sequences upstream of the coding sequence.
The term “genetically engineered cell” or “genetically modified cell” as used herein refers to a cell modified by means of genetic engineering. The term as used herein “engineered” or “modified” thereof may refer to one or more changes of nucleic acids, such as nucleic acids within the genome of an organism. The term “engineered” or “modified” may refer to a change, addition and/or deletion of a gene. Engineered cells or modified cells can also refer to cells that contain added, deleted, and/or changed genes.
The terms “identical” or percent “identity,” in the context of two or more nucleic acids or polypeptide sequences, refer to two or more sequences or subsequences that are the same or have a specified percentage of amino acid residues or nucleotides that are the same (i.e., about 60% identity, preferably 61%, 62%, 63%, 64%, 65%, 66%, 67%, 68%, 69%, 70%, 71%, 72%, 73%, 74%, 75%, 76%, 77%, 78%, 79%, 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or higher identity over a specified region when compared and aligned for maximum correspondence over a comparison window or designated region) as measured using a BLAST or BLAST 2.0 sequence comparison algorithms with default parameters described below, or by manual alignment and visual inspection (see, e.g., NCBI web site or the like). Such sequences are then said to be “substantially identical.” This definition also refers to, or may be applied to, the compliment of a test sequence. The definition also includes sequences that have deletions and/or additions,
as well as those that have substitutions. As described below, the preferred algorithms can account for gaps and the like. Preferably, identity exists over a region that is at least about 10 amino acids or 20 nucleotides in length, or more preferably over a region that is 10-50 amino acids or 20-50 nucleotides in length. As used herein, percent (%) amino acid sequence identity is defined as the percentage of amino acids in a candidate sequence that are identical to the amino acids in a reference sequence, after aligning the sequences and introducing gaps, if necessary, to achieve the maximum percent sequence identity. Alignment for purposes of determining percent sequence identity can be achieved in various ways that are within the skill in the art, for instance, using publicly available computer software such as BLAST, BLAST-2, ALIGN, ALIGN-2 or Megalign (DNASTAR) software. Appropriate parameters for measuring alignment, including any algorithms needed to achieve maximal alignment over the full-length of the sequences being compared can be determined by known methods.
For sequence comparisons, typically one sequence acts as a reference sequence, to which test sequences are compared. When using a sequence comparison algorithm, test and reference sequences are entered into a computer, subsequence coordinates are designated, if necessary, and sequence algorithm program parameters are designated. Preferably, default program parameters can be used, or alternative parameters can be designated. The sequence comparison algorithm then calculates the percent sequence identities for the test sequences relative to the reference sequence, based on the program parameters.
One example of an algorithm that is suitable for determining percent sequence identity and sequence similarity are the BLAST and BLAST 2.0 algorithms, which are described in Altschul et al. (1977) Nuc. Acids Res. 25:3389-3402, and Altschul et al. (1990) J. Mol. Biol. 215:403-410, respectively. Software for performing BLAST analyses is publicly available through the National Center for Biotechnology Information (ncbi.nlm.nih.gov/). This algorithm involves first identifying high scoring sequence pairs (HSPs) by identifying short words of length W in the query sequence, which either match or satisfy some positive-valued threshold score T when aligned with a word of the same length in a database sequence. T is referred to as the neighborhood word score threshold (Altschul et al. (1990) J. Mol. Biol. 215:403-410). These initial neighborhood word hits act as seeds for initiating searches to find longer HSPs containing them. The word hits are extended in both directions along each sequence for as far as the cumulative alignment score can be increased. Cumulative scores are calculated using, for nucleotide sequences, the parameters M (reward score for a pair of matching residues; always >0) and N (penalty score for mismatching residues; always <0). For amino acid sequences, a scoring matrix is used to calculate the cumulative score. Extension of
the word hits in each direction are halted when: the cumulative alignment score falls off by the quantity X from its maximum achieved value; the cumulative score goes to zero or below, due to the accumulation of one or more negative-scoring residue alignments; or the end of either sequence is reached. The BLAST algorithm parameters W, T, and X determine the sensitivity and speed of the alignment. The BLASTN program (for nucleotide sequences) uses as defaults a wordlength (W) of 11, an expectation (E) or 10, M=5, N=-4 and a comparison of both strands. For amino acid sequences, the BLASTP program uses as defaults a wordlength of 3, and expectation (E) of 10, and the BLOSUM62 scoring matrix (see Henikoff and Henikoff (1989) Proc. Natl. Acad. Set. USA 89: 10915) alignments (B) of 50, expectation (E) of 10, M=5, N=-4, and a comparison of both strands.
The BLAST algorithm also performs a statistical analysis of the similarity between two sequences (see, e.g., Karlin and Altschul (1993) Proc. Natl. Acad. Set. USA 90:5873-5787). One measure of similarity provided by the BLAST algorithm is the smallest sum probability (P(N)), which provides an indication of the probability by which a match between two nucleotide or amino acid sequences would occur by chance. For example, a nucleic acid is considered similar to a reference sequence if the smallest sum probability in a comparison of the test nucleic acid to the reference nucleic acid is less than about 0.2, more preferably less than about 0.01.
The term “increased” or “increase” as used herein generally means an increase by a statically significant amount; for the avoidance of any doubt, “increased” means an increase of at least 10% as compared to a reference level, for example an increase of at least about 20%, or at least about 30%, or at least about 40%, or at least about 50%, or at least about 60%, or at least about 70%, or at least about 80%, or at least about 90% or up to and including a 100% increase or any increase between 10-100% as compared to a reference level, or at least about a 2-fold, or at least about a 3-fold, or at least about a 4-fold, or at least about a 5-fold or at least about a 10-fold increase, or any increase between 2-fold and 10-fold or greater as compared to a reference level so long as the increase is statistically significant.
As used herein, the terms “may,” “optionally,” and “may optionally” are used interchangeably and are meant to include cases in which the condition occurs as well as cases in which the condition does not occur. Thus, for example, the statement that a formulation “may include an excipient” is meant to include cases in which the formulation includes an excipient as well as cases in which the formulation does not include an excipient.
Nucleic acid is “operably linked” when it is placed into a functional relationship with another nucleic acid sequence or amino acid sequence. For example, DNA for a presequence
or secretory leader is operably linked to DNA for a polypeptide if it is expressed as a preprotein that participates in the secretion of the polypeptide; a promoter or enhancer is operably linked to a coding sequence if it affects the transcription of the sequence; or a ribosome binding site is operably linked to a coding sequence if it is positioned so as to facilitate translation. Generally, “operably linked” means that the DNA sequences being linked are near each other, and, in the case of a secretory leader, contiguous and in reading phase. However, operably linked nucleic acids (e.g., enhancers and coding sequences) do not have to be contiguous. Linking is accomplished by ligation at convenient restriction sites. If such sites do not exist, the synthetic oligonucleotide adaptors or linkers are used in accordance with conventional practice. In embodiments, a promoter is operably linked with a coding sequence when it is capable of affecting (e.g., modulating relative to the absence of the promoter) the expression of a protein from that coding sequence (i.e., the coding sequence is under the transcriptional control of the promoter).
The term “promoter” or “regulatory element” refers to a region or sequence determinants located upstream or downstream from the start of transcription and which are involved in recognition and binding of RNA polymerase and other proteins to initiate transcription. Promoters need not be of bacterial origin, for example, promoters derived from viruses or from other organisms can be used in the compositions, systems, or methods described herein. The term “regulatory element” is intended to include promoters, enhancers, internal ribosomal entry sites (IRES), and other expression control elements (e.g., transcription termination signals, such as polyadenylation signals and poly-U sequences). Such regulatory elements are described, for example, in Goeddel, Gene Expression Technology: Methods in Enzymology, Academic Press, San Diego, Calif. (1990). Regulatory elements include those that direct constitutive expression of a nucleotide sequence in many types of host cell and those that direct expression of the nucleotide sequence only in certain host cells (e.g., tissue-specific regulatory sequences). A tissue-specific promoter may direct expression primarily in a desired tissue of interest, such as muscle, neuron, bone, skin, blood, specific organs, or particular cell types. Regulatory elements may also direct expression in a temporal-dependent manner, such as in a cell-cycle dependent or developmental stage-dependent manner, which may or may not also be tissue or cell-type specific. In some embodiments, a vector comprises one or more pol III promoter (e.g., 1, 2, 3, 4, 5, or more pol I promoters), one or more pol II promoters (e.g., 1, 2, 3, 4, 5, or more pol II promoters), one or more pol I promoters (e.g., 1, 2, 3, 4, 5, or more pol I promoters), or combinations thereof. Examples of pol III promoters include, but are not limited to, U6 and Hl promoters. Examples of pol II promoters include, but are not limited to,
the retroviral Rous sarcoma virus (RSV) LTR promoter (optionally with the RSV enhancer), the cytomegalovirus (CMV) promoter (optionally with the CMV enhancer) [see, e.g., Boshart et al, Cell, 41 :521-530 (1985)], the SV40 promoter, the dihydrofolate reductase promoter, the P-actin promoter, the phosphoglycerol kinase (PGK) promoter, and the EFla promoter. Also encompassed by the term “regulatory element” are enhancer elements, such as WPRE; CMV enhancers; the R-U5' segment in LTR of HTLV-I (Mol. Cell. Biol., Vol. 8(1), p. 466-472, 1988); SV40 enhancer; and the intron sequence between exons 2 and 3 of rabbit P-globin (Proc. Natl. Acad. Sci. USA., Vol. 78(3), p. 1527-31, 1981). It will be appreciated by those skilled in the art that the design of the expression vector can depend on such factors as the choice of the host cell to be transformed, the level of expression desired, etc.
The term “recombinant” refers to a human manipulated nucleic acid (e.g., polynucleotide) or a copy or complement of a human manipulated nucleic acid (e.g., polynucleotide), or if in reference to a protein (i.e, a “recombinant protein”), a protein encoded by a recombinant nucleic acid (e.g., polynucleotide). In embodiments, a recombinant expression cassette comprising a promoter operably linked to a second nucleic acid (e.g., polynucleotide) may include a promoter that is heterologous to the second nucleic acid (e.g. polynucleotide) as the result of human manipulation (e.g., by methods described in Sambrook et al., Molecular Cloning — A Laboratory Manual, Cold Spring Harbor Laboratory, Cold Spring Harbor, N.Y., (1989) or Current Protocols in Molecular Biology Volumes 1-3, John Wiley & Sons, Inc. (1994-1998)). In another example, a recombinant expression cassette may comprise nucleic acids (e.g., polynucleotides) combined in such a way that the nucleic acids (e.g., polynucleotides) are extremely unlikely to be found in nature. For instance, human manipulated restriction sites or plasmid vector sequences may flank or separate the promoter from the second nucleic acid (e.g., polynucleotide). One of skill will recognize that nucleic acids (e.g., polynucleotides) can be manipulated in many ways and are not limited to the examples above.
The term “reduced”, “reduce”, “reduction”, or “decrease” as used herein generally means a decrease by a statistically significant amount. However, for avoidance of doubt, “reduced” means a decrease by at least 10% as compared to a reference level, for example a decrease by at least about 20%, or at least about 30%, or at least about 40%, or at least about 50%, or at least about 60%, or at least about 70%, or at least about 80%, or at least about 90% or up to and including a 100% decrease (i.e. absent level as compared to a reference sample), or any decrease between 10-100% as compared to a reference level so long as the decrease is statistically significant.
As used throughout, by a "subject" (or a “host”) is meant an individual. Thus, the "subject" can include, for example, domesticated animals, such as cats, dogs, etc., livestock (e.g., cattle, horses, pigs, sheep, goats, etc.), laboratory animals (e.g., mouse, rabbit, rat, guinea pig, etc.) mammals, non-human mammals, primates, non-human primates, rodents, birds, reptiles, amphibians, fish, and any other animal. The subject can be a mammal such as a primate or a human. Administration of the therapeutic agents can be carried out at dosages and for periods of time effective for treatment of a subject.
As used herein, a “target”, “target molecule”, or “target cell” refers to a biomolecule or a cell that can be the focus of a therapeutic drug strategy, diagnostic assay, or a combination thereof, sometimes referred to as a theranostic. Therefore, a target can include, without limitation, many organic molecules that can be produced by a living organism or synthesized, for example, a protein or portion thereof, a peptide, a polysaccharide, an oligosaccharide, a sugar, a glycoprotein, a lipid, a phospholipid, a polynucleotide or portion thereof, an oligonucleotide, an aptamer, a nucleotide, a nucleoside, DNA, RNA, a DNA/RNA chimera, an antibody or fragment thereof, a receptor or a fragment thereof, a receptor ligand, a nucleic acidprotein fusion, a hapten, a nucleic acid, a virus or a portion thereof, an enzyme, a co-factor, a cytokine, a chemokine, as well as small molecules (e.g., a chemical compound), for example, primary metabolites, secondary metabolites, and other biological or chemical molecules that are capable of activating, inhibiting, or modulating a biochemical pathway or process, and/or any other affinity agent, among others.
“Therapeutically effective amount” or “therapeutically effective dose” of a composition refers to an amount that is effective to achieve a desired therapeutic result. In some embodiments, a desired therapeutic result is reduction or clearance of a pathogen. Therapeutically effective amounts of a given therapeutic agent will typically vary with respect to factors such as the type and severity of the disorder or disease being treated and the age, gender, and weight of the subject. The term can also refer to an amount of a therapeutic agent, or a rate of delivery of a therapeutic agent (e.g., amount over time), effective to facilitate a desired therapeutic effect. The precise desired therapeutic effect will vary according to the condition to be treated, the tolerance of the subject, the agent and/or agent formulation to be administered (e.g., the potency of the therapeutic agent, the concentration of agent in the formulation, and the like), and a variety of other factors that are appreciated by those of ordinary skill in the art. In some instances, a desired biological or medical response is achieved following administration of multiple dosages of the composition to the subject over a period of days, weeks, or years.
As used herein, the terms “treating” or “treatment” of a subject includes the administration of a drug to a subject with the purpose of curing, healing, alleviating, relieving, altering, remedying, ameliorating, improving, stabilizing or affecting a disease or disorder, or a symptom of a disease or disorder. The terms “treating” and “treatment” can also refer to reduction in severity and/or frequency of symptoms, elimination of symptoms and/or underlying cause, and improvement or remediation of damage.
As used herein, the term “preventing” a disease, a disorder, or unwanted physiological event in a subject refers to the prevention of a disease, a disorder, or unwanted physiological event or prevention of a symptom of a disease, a disorder, or unwanted physiological event
"Pharmaceutically acceptable" component can refer to a component that is not biologically or otherwise undesirable, i.e., the component may be incorporated into a pharmaceutical formulation of the invention and administered to a subject as described herein without causing significant undesirable biological effects or interacting in a deleterious manner with any of the other components of the formulation in which it is contained. When used in reference to administration to a human, the term generally implies the component has met the required standards of toxicological and manufacturing testing or that it is included on the Inactive Ingredient Guide prepared by the U.S. Food and Drug Administration.
"Pharmaceutically acceptable carrier" (sometimes referred to as a “carrier”) means a carrier or excipient that is useful in preparing a pharmaceutical or therapeutic composition that is generally safe and non-toxic, and includes a carrier that is acceptable for veterinary and/or human pharmaceutical or therapeutic use. The terms "carrier" or "pharmaceutically acceptable carrier" can include, but are not limited to, phosphate buffered saline solution, water, emulsions (such as an oil/water or water/oil emulsion) and/or various types of wetting agents. As used herein, the term "carrier" encompasses, but is not limited to, any excipient, diluent, filler, salt, buffer, stabilizer, solubilizer, lipid, stabilizer, or other material well known in the art for use in pharmaceutical formulations and as described further herein.
“Therapeutic agent” refers to any composition that has a beneficial biological effect. Beneficial biological effects include both therapeutic effects, e.g., treatment of a disorder or other undesirable physiological condition, and prophylactic effects, e.g., prevention of a disorder or other undesirable physiological condition. The terms also encompass pharmaceutically acceptable, pharmacologically active derivatives of beneficial agents specifically mentioned herein, including, but not limited to, cells, salts, esters, amides, proagents, active metabolites, isomers, fragments, analogs, and the like. When the term “therapeutic agent” is used, or when a particular agent is specifically identified, it is to be
understood that the term includes the agent per se as well as pharmaceutically acceptable, pharmacologically active salts, esters, amides, proagents, conjugates, active metabolites, isomers, fragments, analogs, etc.
The term “polypeptide” refers to a compound made up of a single chain of D- or L- amino acids or a mixture of D- and L-amino acids joined by peptide bonds.
The term “nucleic acid” as used herein means a polymer composed of nucleotides, e.g., deoxyribonucleotides or ribonucleotides.
The term "nucleobase" refers to the part of a nucleotide that bears the Watson/Crick base-pairing functionality. The most common naturally-occurring nucleobases, adenine (A), guanine (G), uracil (U), cytosine (C), and thymine (T) bear the hydrogen-bonding functionality that binds one nucleic acid strand to another in a sequence specific manner.
The terms “ribonucleic acid” and “RNA” as used herein mean a polymer composed of ribonucleotides.
The terms “deoxyribonucleic acid” and “DNA” as used herein mean a polymer composed of deoxyribonucleotides.
The term “polynucleotide” refers to a single or double stranded polymer composed of nucleotide monomers.
Disclosed herein are the components to be used to prepare the disclosed compositions as to be used in the methods disclosed herein. These and other materials are disclosed herein, and it is understood that when combinations, subsets, interactions, groups, etc. of these materials are disclosed that while specific reference of each various individual and collective combinations and permutation of these compounds may not be explicitly disclosed, each is specifically contemplated and described herein. If a class of molecules A, B, and C are disclosed as well as a class of molecules D, E, and F and an example of a combination molecule, A-D is disclosed, then even if each is not individually recited each is individually and collectively contemplated meaning combinations, A-E, A-F, B-D, B-E, B-F, C-D, C-E, and C- F are considered disclosed. Likewise, any subset or combination of these is also disclosed. Thus, for example, the sub-group of A-E, B-F, and C-E would be considered disclosed. This concept applies to all aspects of this application including, but not limited to, steps in methods of making and using the disclosed compositions. Thus, if there are a variety of additional steps that can be performed it is understood that each of these additional steps can be performed with any specific embodiment or combination of embodiments of the disclosed methods.
Gene-Editing Systems
In some aspects, disclosed herein is a system comprising a Cas9 nickase; a reverse transcriptase; a small RNA binding exonuclease protection factor La (SSB) protein; and a prime editing guide RNA (pegRNA).
The term “prime editing system” involves a Cas9 nickase (for example, a Cas9 H840A nickase or a Cas9 R221K N394K H840A nickase) and a reverse transcriptase, in combination with a guide RNA (herein referred as “prime editing guide RNA” or “pegRNA”). The pegRNA is a sgRNA with a primer binding site (PBS) and a DNA synthesis template appended to the 3’ end containing the desired nucleic acid sequence. During genome editing, the primer binding site allows the 3’ end of a nicked DNA strand to hybridize to the pegRNA, while the RT template serves as a template for the synthesis of edited genetic information. The pegRNA encodes the new sequence and allows DNA synthesis to introduce the desired mutations. In some examples, the prime editing systems and pegRNAs are those described in U.S. Patent Nol, 1447,770, which is incorporated herein by reference in its entirety. In some embodiments, the pegRNA comprises an engineered pegRNA (epegRNA).
A pegRNA can be longer than standard sgRNAs commonly used for CRISPR gene editing. In some embodiments, the pegRNA disclosed herein can be at least 60 nt, 70 nt, 80 nt, 90 nt, 100 nt, 120 nt, 140 nt, 160 nt, 180 nt, 200 nt, 250 nt, 300 nt, 350 nt, 400 nt, or 500 nt in length.
The terms “guide RNA”, “single guide RNA”, or “sgRNA” are used interchangeably and refer to the polynucleotide sequence comprising the guide sequence, the tracrRNAand the crRNA. The term “guide sequence” refers to the about 20 bp sequence within the guide RNA that specifies the target site and may be used interchangeably with the term “guide” or “spacer”. A “crRNA” is a bacterial RNA that confers target specificity and requires tracrRNA to bind to Cas9. A “tracrRNA” is a bacterial RNA that links the crRNA to the Cas9 nuclease and typically can bind any crRNA. The sequence specificity of a Cas DNA-binding protein is determined by gRNAs, which have nucleotide base-pairing complementarity to target DNA sequences.
In some embodiments, the pegRNA comprises a 3 ’-polyuridine domain (which comprises, for example, at least 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 25, 30, 35, 40, 45, 50, 60, 70, 80, 90, or 100 uridines). The 3 ’-polyuridine domain can locate at the 3’- end of the pegRNA or near the 3’ - end of the pegRNA. In some embodiments, the 3 ’-polyuridine domain comprises at least one chemically modified uridine (for example, at
least one, two, three, four, or five chemically modified uridines). In some embodiments, the 3’- polyuridine domain comprises at least one unmodified uridine (for example, at least one, two, three, four, or five unmodified uridines). In some embodiments, the 3 ’-polyuridine domain comprises at least one chemically modified uridine (for example, at least one, two, three, four, or five chemically modified uridines) and at least one unmodified uridine (for example, at least one, two, three, four, or five unmodified uridines). In some embodiments, the at least one unmodified uridine locates at the 3’ end of the pegRNA. For example, the pegRNA disclosed herein can have two unmodified uridines locating at its 3’ end, downstream of at least one, two, three, four, or five chemically modified uridines. In some embodiments, the 3 ’-polyuridine domain comprises the sequence UU*mU*mU*mUU. In some embodiments, the 3’- polyuridine domain comprises a 3’ sequence fragment selected from the sequences in Table 3. In some embodiments, the 3 ’-polyuridine domain comprises a 3’ sequence fragment selected from SEQ ID NOs: 63-108.
In some embodiments, the chemical modification is 2’ -O-m ethylation and/or replacement of a phosphodiester bond to a phosphorothioate bond.
When a synthetic pegRNA has a 3’-polyU and as long as the 2’-OH of the last uridine is not modified, SSB can interact with synthetic pegRNA. For example, *mUrU described herein is the modification pattern whose last uridine is not modified, whereas *mU*rU has its last uridine technically modified due to the replacement of phosphodiester bond to phosphorothioate bond. Both modification patterns allow synthetic pegRNAs to interact with SSB. Accordingly, in some embodiments, the 3 ’-polyuridine domain comprises at least one (for example, at least two, three, four, or five) uridine with unmodified 2'-hydroxyl (OH) group. In some embodiments, the at least one uridine with unmodified 2'- OH group locates at the 3’ end of the pegRNA.
In some embodiments, the SSB protein comprises a La motif and/or an RNA recognition motif (RRM) (e.g., amino acid residues 1-194 or 2-194 of SEQ ID NO: 34). In some embodiments, the SSB protein comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 33 or a fragment thereof. In some embodiments, the SSB protein comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 34 or a fragment thereof. In some embodiments, the SSB protein comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical SEQ ID NO: 35 or a fragment thereof. In some embodiments, the SSB protein comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 36 or a fragment thereof.
In some embodiments, the Cas9 nickase of the system disclosed herein comprises a sequence at least 80% identical to SEQ ID NO: 26, or a fragment thereof. In some embodiments, the Cas9 nickase of the system disclosed herein comprises a sequence at least 80% identical to SEQ ID NO: 27, or a fragment thereof.
In some embodiments, the reverse transcriptase of the system disclosed herein comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 31 or a fragment thereof. In some embodiments, the reverse transcriptase of the system disclosed herein comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 32 or a fragment thereof.
In some embodiments, the Cas9 nickase is operatively linked to the reverse transcriptase (e.g., directly or through a linker). In some embodiments, the SSB protein is operatively linked to the Cas9 nickase and the reverse transcriptase (e.g., directly or through one or more linkers). Accordingly, in some embodiments, the system disclosed herein comprises a recombinant polypeptide that comprises the reverse transcriptase, the Cas9 nickase, and the SSB protein disclosed herein. The recombinant polypeptide can further comprise one or more linkers and/or one or more nuclear localization sequences (NLS) (e.g., aNLS of SV40 or c-Myc). In some embodiments, the linker comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 28, 29, 30, or 40, or a fragment thereof. In some embodiments, the NLS comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 25, 37, 38, 39, or 40 or a fragment thereof.
In some embodiments, the recombinant polypeptide comprising the reverse transcriptase, the Cas9 nickase, and the SSB protein comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to any of SEQ ID NOs: 1-12 or a fragment thereof. In some embodiments, the recombinant polypeptide comprising the reverse transcriptase, the Cas9 nickase, and the SSB protein comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 5 or a fragment thereof.
In some embodiments, the recombinant polypeptide comprising the reverse transcriptase, the Cas9 nickase, and the SSB protein comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 1 or a fragment thereof. In some embodiments, the recombinant polypeptide comprising the reverse transcriptase, the Cas9 nickase, and the SSB protein comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 2 or a fragment thereof. In some embodiments, the recombinant polypeptide comprising the reverse transcriptase, the Cas9
nickase, and the SSB protein comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 3 or a fragment thereof. In some embodiments, the recombinant polypeptide comprising the reverse transcriptase, the Cas9 nickase, and the SSB protein comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 4 or a fragment thereof. In some embodiments, the recombinant polypeptide comprising the reverse transcriptase, the Cas9 nickase, and the SSB protein comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 5 or a fragment thereof. In some embodiments, the recombinant polypeptide comprising the reverse transcriptase, the Cas9 nickase, and the SSB protein comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 6 or a fragment thereof. In some embodiments, the recombinant polypeptide comprising the reverse transcriptase, the Cas9 nickase, and the SSB protein comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 7 or a fragment thereof. In some embodiments, the recombinant polypeptide comprising the reverse transcriptase, the Cas9 nickase, and the SSB protein comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 8 or a fragment thereof. In some embodiments, the recombinant polypeptide comprising the reverse transcriptase, the Cas9 nickase, and the SSB protein comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 9 or a fragment thereof. In some embodiments, the recombinant polypeptide comprising the reverse transcriptase, the Cas9 nickase, and the SSB protein comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 10 or a fragment thereof. In some embodiments, the recombinant polypeptide comprising the reverse transcriptase, the Cas9 nickase, and the SSB protein comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 11 or a fragment thereof. In some embodiments, the recombinant polypeptide comprising the reverse transcriptase, the Cas9 nickase, and the SSB protein comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 12 or a fragment thereof.
Also disclosed herein is a system comprising a Cas9 nickase; a reverse transcriptase; and a small RNA binding exonuclease protection factor La (SSB) protein.
The Cas9 nickase, reverse transcriptase, and the SSB protein can be on a same or different polypeptide. In some embodiments, the Cas9 nickase, reverse transcriptase, and the SSB protein can be on a same or different pharmaceutically acceptable carriers.
Also disclosed herein is a system comprising a first polynucleotide encoding a Cas9 nickase; a second polynucleotide encoding a reverse transcriptase; and a third polynucleotide encoding a small RNA binding exonuclease protection factor La (SSB) protein; and a prime editing guide RNA (pegRNA).
In some embodiments, the third polynucleotide encodes a La motif and/or an RNA recognition motif (RRM) of the SSB protein. In some embodiments, the third polynucleotide comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 50 or a fragment thereof. In some embodiments, the third polynucleotide comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 49 or a fragment thereof.
In some embodiments, the first, second, and third polynucleotides are operatively linked thereby forming one recombinant polynucleotide. Accordingly, in some embodiments, the system disclosed herein comprises a recombinant polynucleotide encoding a recombinant polypeptide that comprises the reverse transcriptase, the Cas9 nickase, and the SSB protein disclosed herein. The recombinant polypeptide can further comprise one or more linkers and/or one or more nuclear localization sequences (NLS) (e.g., a NLS of SV40 or c-Myc). In some embodiments, the polynucleotide encoding the linker comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 44, 45, 46, or 56, or a fragment thereof. In some embodiments, the polynucleotide encoding the NLS comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 41, 53, 54, 55, 56 or a fragment thereof.
In some embodiments, the recombinant nucleotide comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to any of SEQ ID NOs: 13-24 or a fragment thereof. In some embodiments, the recombinant nucleotide comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 17 or a fragment thereof.
Also disclosed herein is a system comprising a first polynucleotide encoding a Cas9 nickase; a second polynucleotide encoding a reverse transcriptase; and
a third polynucleotide encoding a small RNA binding exonuclease protection factor La (SSB) protein.
In some embodiments, the first, second, and third polynucleotides disclosed herein are located on a same or different pharmaceutically acceptable carriers or vectors.
It is herein contemplated that the polypeptides, polynucleotides, or systems disclosed herein can be applied as paired prime editing for precise genomic deletions. In some examples, the polypeptides, polynucleotides, or systems disclosed herein can be used for the PRIME-Del, which induces a deletion using a pair of prime editing sgRNAs (pegRNAs) that target opposite DNA strands, programming not only the sites that are nicked but also the outcome of the repair. The PRIME-Del technology is known in the art. See, Choi, J., Chen, W., Suiter, C.C. et al. Precise genomic deletions using paired prime editing. Nat Biotechnol 40, 218-226 (2022)., incorporated by reference herein in its entirety. In some examples, the polypeptides, polynucleotides, or systems disclosed herein can be used for twin prime editing (or twinPE), a DSB-independent method that uses a prime editor protein and two prime editing guide RNAs (pegRNAs) for the programmable replacement or excision of DNA sequences at endogenous human genomic sites. Twin prime editing can be used for programmable deletion, replacement, integration and inversion of large DNA sequences (for example, enabling targeted integration of gene-sized DNA plasmids (>5,000 bp) and targeted sequence inversions of 40 kb in human cells). The twin prime editing technology is known in the art. See, Anzalone, A.V., Gao, X.D., Podracky, C.J. et al. Programmable deletion, replacement, integration and inversion of large DNA sequences with twin prime editing. Nat Biotechnol 40, 731-740 (2022)., incorporated by reference herein in its entirety. In some examples, the polypeptides, polynucleotides, or systems disclosed herein can be used for programmable addition via site-specific targeting elements (PASTE), which uses a CRISPR-Cas9 nickase fused to both a reverse transcriptase and serine integrase for targeted genomic recruitment and integration of desired payloads. PASTE enables integration of large sequences (e.g., as large as ~36 kilobases) at multiple genomic loci into cells (e.g., human cell lines, primary T cells, hematopoietic stem and progenitor cells (HSPCs), and non-dividing primary human hepatocytes). The PASTE technology is known in the art. See Yarnall, M.T.N., loannidi, E.I., Schmitt-Ulms, C. et al. Drag-and-drop genome insertion of large sequences without double-strand DNA cleavage using CRISPR-directed integrases. Nat Biotechnol 41, 500-512 (2023).)., which is incorporated by reference herein in its entirety.
Also disclosed herein is an engineered cell with the polypeptides, polynucleotides, or systems disclosed herein. The compositions and methods disclosed herein can be used for
making cells or cell lines with desired sequence changes, e.g., mutations, large insertions and large deletions). In some embodiments, the engineered cell is a stem cell (e.g., an induced pluripotent stem cell, hematopoietic stem cell, or a hematopoietic stem and progenitor cell (HSPC)), or an immune cell (e.g., a CAR-T cell, a primary T cell).
Also disclosed herein is an animal model with the polypeptides, polynucleotides, systems, or cells disclosed herein. In some embodiments, the animal model is genetically engineered.
Also disclosed herein is a pharmaceutical composition comprising the polypeptides, polynucleotides, systems, or engineered cells disclosed herein.
In certain embodiments, the disclosure contemplates pharmaceutical compositions comprising the polypeptides, the polynucleotides, the vectors, or the systems disclosed herein, or optionally other pharmaceutical agent, or pharmaceutically acceptable salts thereof, and a pharmaceutically acceptable excipient. In certain embodiments, this disclosure contemplates the production of a medicament comprising the polypeptides, polynucleotides, vectors, or systems disclosed herein, or agents disclosed herein and uses for methods disclosed herein.
Methods of Treatment
Also disclosed herein is a method of preventing and/or treating a genetic disorder in a subject in need, comprising administering to the subject a therapeutically effective amount of the polypeptides, polynucleotides, system, engineered cells, or the pharmaceutical composition disclosed herein.
Also disclosed herein is a method for altering expression of a gene product in a cell, comprising introducing into the cell an effective amount of the polypeptides, polynucleotides, or system, or the pharmaceutical composition disclosed herein.
In some embodiments, the system comprises a Cas9 nickase; a reverse transcriptase; a small RNA binding exonuclease protection factor La (SSB) protein; and a prime editing guide RNA (pegRNA).
In some embodiments, the system comprises a Cas9 nickase; a reverse transcriptase; and a small RNA binding exonuclease protection factor La (SSB) protein.
The Cas9 nickase, reverse transcriptase, SSB protein, and pegRNA can be administered together or separately to the subject or cell.
In some embodiments, the pegRNA comprises a 3 ’-polyuridine domain (which comprises, for example, at least 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 25, 30, 35, 40, 45, 50, 60, 70, 80, 90, or 100 uridines). The 3 ’-polyuridine domain can locate at the 3’- end of the pegRNA or near the 3’ - end of the pegRNA. In some embodiments, the 3 ’-polyuridine domain comprises at least one chemically modified uridine (for example, at least one, two, three, four, or five chemically modified uridines). In some embodiments, the 3’- polyuridine domain comprises at least one unmodified uridine (for example, at least one, two, three, four, or five unmodified uridines). In some embodiments, the 3 ’-polyuridine domain comprises at least one chemically modified uridine (for example, at least one, two, three, four, or five chemically modified uridines) and at least one unmodified uridine (for example, at least one, two, three, four, or five unmodified uridines). In some embodiments, the at least one unmodified uridine locates at the 3’ end of the pegRNA. For example, the pegRNA disclosed herein can have two unmodified uridines locating at its 3’ end, downstream of at least one, two, three, four, or five chemically modified uridines. In some embodiments, the uridines are chemically modified with 2’-O-methylation.
In some embodiments, the chemical modification is 2’ -O-m ethylation and/or replacement of a phosphodiester bond to a phosphorothioate bond. In some embodiments, the 3 ’-polyuridine domain comprises at least one (for example, at least two, three, four, or five) uridine with unmodified 2'-hydroxyl (OH) group. In some embodiments, the at least one uridine with unmodified 2'- OH group locates at the 3’ end of the pegRNA.
In some embodiments, the SSB protein comprises a La motif and/or an RNA recognition motif (RRM) (e.g., amino acid residues 1-194 or 2-194 of SEQ ID NO: 34). In some embodiments, the SSB protein comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 33 or a fragment thereof. In some embodiments, the SSB protein comprises a sequence at least 80% (e.g., at least 80%, 85%,
90%, 95%, 98%, or 99%) identical to SEQ ID NO: 34 or a fragment thereof. In some embodiments, the SSB protein comprises a sequence at least 80% (e.g., at least 80%, 85%,
90%, 95%, 98%, or 99%) identical SEQ ID NO: 35 or a fragment thereof. In some embodiments, the SSB protein comprises a sequence at least 80% (e.g., at least 80%, 85%,
90%, 95%, 98%, or 99%) identical to SEQ ID NO: 36 or a fragment thereof.
In some embodiments, the Cas9 nickase of the system disclosed herein comprises a sequence at least 80% identical to SEQ ID NO: 26, or a fragment thereof. In some embodiments,
the Cas9 nickase of the system disclosed herein comprises a sequence at least 80% identical to SEQ ID NO: 27, or a fragment thereof.
In some embodiments, the reverse transcriptase of the system disclosed herein comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 31 or a fragment thereof. In some embodiments, the reverse transcriptase of the system disclosed herein comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 32 or a fragment thereof.
In some embodiments, the Cas9 nickase is operatively linked to the reverse transcriptase (e.g., through a linker). In some embodiments, the SSB protein is operatively linked to the Cas9 nickase and the reverse transcriptase. Accordingly, in some embodiments, the system disclosed herein comprises a recombinant polypeptide that comprises the reverse transcriptase, the Cas9 nickase, and the SSB protein disclosed herein. The recombinant polypeptide can further comprise one or more linkers and/or one or more nuclear localization sequences (NLS) (e.g., a NLS of SV40 or c-Myc). In some embodiments, the linker comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 28, 29, 30, or 40, or fragment thereof. In some embodiments, the NLS comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 25, 37, 38, 39, or 40. In some embodiments, the recombinant polypeptide comprises the reverse transcriptase, the Cas9 nickase, and the SSB protein comprises a sequence at least 80% identical to any of SEQ ID NOs: 1-12 or a fragment thereof.
In some embodiments, the system comprises a first polynucleotide encoding a Cas9 nickase; a second polynucleotide encoding a reverse transcriptase; and a third polynucleotide encoding a small RNA binding exonuclease protection factor La (SSB) protein; and a prime editing guide RNA (pegRNA).
In some embodiments, the system comprises a first polynucleotide encoding a Cas9 nickase; a second polynucleotide encoding a reverse transcriptase; and a third polynucleotide encoding a small RNA binding exonuclease protection factor La (SSB) protein
The first, second, and third polynucleotides and the pegRNA can be administered together or separately to the subject or cell.
In some embodiments, the first polynucleotide comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 42 or a fragment thereof. In some embodiments, the first polynucleotide comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 43 or a fragment thereof.
In some embodiments, the second polynucleotide comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 47 or a fragment thereof. In some embodiments, the second polynucleotide comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 48 or a fragment thereof.
In some embodiments, the third polynucleotide comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 49 or a fragment thereof. In some embodiments, the third polynucleotide comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 50 or a fragment thereof. In some embodiments, the third polynucleotide comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 51 or a fragment thereof. In some embodiments, the third polynucleotide comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 52 or a fragment thereof. In some embodiments, the third polynucleotide encodes a La motif and/or an RNA recognition motif (RRM) of the SSB protein.
In some embodiments, the first, second, and third polynucleotides are operatively linked thereby forming one recombinant polynucleotide.
In some embodiments, the recombinant polynucleotide comprises the reverse transcriptase, the Cas9 nickase, and the SSB protein and comprises a sequence at least 80% identical to any of SEQ ID NOs: 13-24 or a fragment thereof.
In some embodiments, the SSB protein is operatively linked to the Cas9 nickase and the reverse transcriptase. Accordingly, in some embodiments, the system disclosed herein comprises a recombinant polynucleotide encoding a recombinant polypeptide that comprises the reverse transcriptase, the Cas9 nickase, and the SSB protein disclosed herein. The recombinant polypeptide can further comprise one or more linkers and/or one or more nuclear localization sequences (NLS) (e.g., a NLS of SV40 or c-Myc). In some embodiments, the polynucleotide encoding the linker comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 44, 45, 46, or 56, or fragment thereof. In some embodiments, the polynucleotide encoding the NLS comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 41, 53, 54, 55, or
56. In some embodiments, the recombinant nucleotide comprises a sequence at least 80% identical to any of SEQ ID NOs: 13-24 or a fragment thereof.
In some embodiments, the recombinant nucleotide comprises a sequence at least 80% identical to SEQ ID NO: 13 or a fragment thereof. In some embodiments, the recombinant nucleotide comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 14 or a fragment thereof. In some embodiments, the recombinant nucleotide comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 15 or a fragment thereof. In some embodiments, the recombinant nucleotide comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 16 or a fragment thereof. In some embodiments, the recombinant nucleotide comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 17 or a fragment thereof. In some embodiments, the recombinant nucleotide comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 18 or a fragment thereof. In some embodiments, the recombinant nucleotide comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 19 or a fragment thereof. In some embodiments, the recombinant nucleotide comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 20 or a fragment thereof. In some embodiments, the recombinant nucleotide comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 21 or a fragment thereof. In some embodiments, the recombinant nucleotide comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 22 or a fragment thereof. In some embodiments, the recombinant nucleotide comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 23 or a fragment thereof. In some embodiments, the recombinant nucleotide comprises a sequence at least 80% (e.g., at least 80%, 85%, 90%, 95%, 98%, or 99%) identical to SEQ ID NO: 24 or a fragment thereof.
The nucleic acids that are delivered to cells typically contain expression controlling systems. For example, the inserted genes in viral and retroviral systems usually contain promoters, and/or enhancers to help control the expression of the desired gene product. A promoter is generally a sequence or sequences of DNA that function when in a relatively fixed location in regard to the transcription start site. A promoter contains core elements required for basic interaction of RNA polymerase and transcription factors, and may contain upstream elements and response elements.
Preferred promoters controlling transcription from vectors in mammalian host cells may be obtained from various sources, for example, the genomes of viruses such as: polyoma, Simian Virus 40 (SV40), adenovirus, retroviruses, hepatitis-B virus and most preferably cytomegalovirus, or from heterologous mammalian promoters, e.g., beta actin promoter. The early and late promoters of the SV40 virus are conveniently obtained as an SV40 restriction fragment which also contains the SV40 viral origin of replication (Fiers et al., Nature, 273: 113 (1978)). The immediate early promoter of the human cytomegalovirus is conveniently obtained as a Hindlll E restriction fragment (Greenway, P.J. et al., Gene 18: 355-360 (1982)). Of course, promoters from the host cell or related species also are useful herein.
Enhancer generally refers to a sequence of DNA that functions at no fixed distance from the transcription start site and can be either 5' (Laimins, L. et al., Proc. Natl. Acad. Sci. 78: 993 (1981)) or 3' (Lusky, M.L., et al., Mol. Cell Bio. 3: 1108 (1983)) to the transcription unit. Furthermore, enhancers can be within an intron (Banerji, J.L. et al., Cell 33: 729 (1983)) as well as within the coding sequence itself (Osborne, T.F., et al., Mol. Cell Bio. 4: 1293 (1984)). They are usually between 10 and 300 bp in length, and they function in cis. Enhancers function to increase transcription from nearby promoters. Enhancers also often contain response elements that mediate the regulation of transcription. Promoters can also contain response elements that mediate the regulation of transcription. Enhancers often determine the regulation of expression of a gene. While many enhancer sequences are now known from mammalian genes (globin, elastase, albumin, -fetoprotein and insulin), typically one will use an enhancer from a eukaryotic cell virus for general expression. Preferred examples are the SV40 enhancer on the late side of the replication origin (bp 100-270), the cytomegalovirus early promoter enhancer, the polyoma enhancer on the late side of the replication origin, and adenovirus enhancers.
In certain embodiments, the promoter and/or enhancer region can act as a constitutive promoter and/or enhancer to maximize expression of the region of the transcription unit to be transcribed. In certain constructs the promoter and/or enhancer region be active in all eukaryotic cell types, even if it is only expressed in a particular type of cell at a particular time. A preferred promoter of this type is the CMV promoter (650 bases). Other preferred promoters are SV40 promoters, cytomegalovirus (full length promoter), and retroviral vector LTR.
The compositions disclosed herein may be in solution, suspension (for example, incorporated into microparticles (such as exosomes) or liposomes). These may be targeted to a particular cell type via antibodies, receptors, or receptor ligands. The compositions disclosed herein may be in exosomes. The term “exosome”, as used herein, refers to a cell-derived
membranous vesicle. They refer to extracellular vesicles, which are generally of between 30 and 200 nm in size, for example in the range of 50-100 nm in size. The exosomes can be engineered to express one or more ligands or molecules for cell-targeting delivery.
Drug load or loading capacity refers to the amount of the composition that can be present in the exosome can be from about 0.1 % to about 60 % of its exosome weight. For example, the amount of the composition present in the exosome can be from about 0.1%, about 0.2%, about 0.3%, about 0.4%, about 0.5%, about 0.6%, about 0.7%, about 0.8%, about 0.9%, about 1%, about 2%, about 2.5%, about 3%, about 3.5%, about 4%, about 4.5%, about 5%, about 5.5%, about 6%, about 6.5%, about 7%, about 7.5%, about 8%, about 8.5%, about 9%, about 9.5%, about 10%, about 10.5%, about 11%, about 11.5%, about 12%, about 12.5%, about 13%, about 13.5%, about 14%, about 15%, about 16%, about 17%, about 18%, about 19%, about 20%, about 22%, about 24%, about 26%, about 28%, about 30%, about 32%, about 34%, about 36%, about 38%, about 40%, about 55%, or about 60% of its exosome weight.
Suitable carriers and their formulations are described in Remington: The Science and Practice of Pharmacy (19th ed.) ed. A.R. Gennaro, Mack Publishing Company, Easton, PA 1995. Typically, an appropriate amount of a pharmaceutically-acceptable salt is used in the formulation to render the formulation isotonic. Examples of the pharmaceutically-acceptable carrier include, but are not limited to, saline, Ringer's solution and dextrose solution. The pH of the solution is preferably from about 5 to about 8, and more preferably from about 7 to about 7.5. Further carriers include sustained release preparations such as semipermeable matrices of solid hydrophobic polymers containing the antibody, which matrices are in the form of shaped articles, e.g., films, liposomes or microparticles. It will be apparent to those persons skilled in the art that certain carriers may be more preferable depending upon, for instance, the route of administration and concentration of composition being administered.
Pharmaceutical carriers are known to those skilled in the art. These most typically would be standard carriers for administration of drugs to humans, including solutions such as sterile water, saline, and buffered solutions at physiological pH. The compositions can be administered intramuscularly or subcutaneously. Other compounds will be administered according to standard procedures used by those skilled in the art.
Pharmaceutical compositions may include carriers, thickeners, diluents, buffers, preservatives, surface active agents and the like in addition to the molecule of choice.
Preparations for parenteral administration include sterile aqueous or non-aqueous solutions, suspensions, and emulsions. Examples of non-aqueous solvents are propylene glycol, polyethylene glycol, vegetable oils such as olive oil, and injectable organic esters such as ethyl
oleate. Aqueous carriers include water, alcoholic/aqueous solutions, emulsions or suspensions, including saline and buffered media. Parenteral vehicles include sodium chloride solution, Ringer's dextrose, dextrose and sodium chloride, lactated Ringer's, or fixed oils. Intravenous vehicles include fluid and nutrient replenishers, electrolyte replenishers (such as those based on Ringer's dextrose), and the like. Preservatives and other additives may also be present such as, for example, antimicrobials, anti-oxidants, chelating agents, and inert gases and the like.
Formulations for topical administration may include ointments, lotions, creams, gels, drops, suppositories, sprays, liquids and powders. Conventional pharmaceutical carriers, aqueous, powder or oily bases, thickeners and the like may be necessary or desirable.
Compositions for oral administration include powders or granules, suspensions or solutions in water or non-aqueous media, capsules, sachets, or tablets. Thickeners, flavorings, diluents, emulsifiers, dispersing aids or binders may be desirable.
Some of the compositions may potentially be administered as a pharmaceutically acceptable acid- or base- addition salt, formed by reaction with inorganic acids such as hydrochloric acid, hydrobromic acid, perchloric acid, nitric acid, thiocyanic acid, sulfuric acid, and phosphoric acid, and organic acids such as formic acid, acetic acid, propionic acid, glycolic acid, lactic acid, pyruvic acid, oxalic acid, malonic acid, succinic acid, maleic acid, and fumaric acid, or by reaction with an inorganic base such as sodium hydroxide, ammonium hydroxide, potassium hydroxide, and organic bases such as mono-, di-, trialkyl and aryl amines and substituted ethanolamines.
The exact amount of the compositions required will vary from subject to subject, depending on the species, age, weight and general condition of the subject, the severity of the allergic disorder being treated, the particular nucleic acid or vector used, its mode of administration and the like. Thus, it is not possible to specify an exact amount for every composition. However, an appropriate amount can be determined by one of ordinary skill in the art using only routine experimentation given the teachings herein.
In some embodiments, the genetic disorder is Sickle cell anemia. Mutations in the HBB gene on chromosome 11 can cause Sickle cell anemia. Accordingly, in some embodiments, the compositions disclosed herein target a mutated HBB gene. In some embodiments, the genetic disorders are those described in U.S. Patent No. 8,697,359, which is incorporated herein by reference in its entirety.
In some embodiments, dosing frequency for the composition disclosed herein, includes, but is not limited to, no more than once every 30 years, every 25 years, every 20 years, every
15 years, every 10 years, every 5 years, every 4 years, every 3 years, every 2 years, every 12 months, or every 6 months.
Dosing frequency for the composition disclosed herein, includes, but is not limited to, at least once every 30 years, every 25 years, every 20 years, every 15 years, every 10 years, every 5 years, every 4 years, every 3 years, every 2 years, every 12 months, once every 11 months, once every 10 months, once every 9 months, once every 8 months, once every 7 months, once every 6 months, once every 5 months, once every 4 months, once every 3 months, once every two months, once every month; or at least once every three weeks, once every two weeks, once a week, twice a week, three times a week, four times a week, five times a week, six times a week, or daily. In some embodiments, the interval between each administration is less than about 4 months, less than about 3 months, less than about 2 months, less than about a month, less than about 3 weeks, less than about 2 weeks, or less than less than about a week, such as less than about any of 6, 5, 4, 3, 2, or 1 day. In some embodiments, the dosing frequency for the composition includes, but is not limited to, at least once a day, twice a day, or three times a day. In some embodiments, the interval between each administration is less than about 48 hours, 36 hours, 24 hours, 22 hours, 20 hours, 18 hours, 16 hours, 14 hours, 12 hours, 10 hours, 9 hours, 8 hours, or 7 hours. In some embodiments, the interval between each administration is less than about 24 hours, 22 hours, 20 hours, 18 hours, 16 hours, 14 hours, 12 hours, 10 hours, 9 hours, 8 hours, 7 hours, or 6 hours. In some embodiments, the interval between each administration is constant. For example, the administration can be carried out daily, every two days, every three days, every four days, every five days, or weekly. Administration can also be continuous and adjusted to maintaining a level of the compound within any desired and specified range. It should be understood and herein contemplated that the compositions disclosed herein can be used in combination with a pain reliever and, in some examples, reduce the dosing frequency of the pain reliever.
In some embodiments, the therapeutically effective amount typically will vary from about 0.001 mg/kg to about 1000 mg/kg, from about 0.01 mg/kg to about 750 mg/kg, from about 100 mg/kg to about 500 mg/kg, from about 1 mg/kg to about 250 mg/kg, from about 10 mg/kg to about 150 mg/kg in one or more dose administrations daily, for one or several days (depending of course of the mode of administration and the factors discussed above). Other suitable dose ranges include 1 mg to 10,000 mg per day, 100 mg to 10,000 mg per day, 500 mg to 10,000 mg per day, and 500 mg to 1,000 mg per day. In some embodiments, the amount is less than 10,000 mg per day with a range of 750 mg to 9,000 mg per day.
In some embodiments, the doses of the compositions disclosed herein for gene editing in a cell or a subject is less (e.g., about 2-fold less, about 3-fold less, about 4-fold less, about 5-fold less, about 6-fold less, about 7-fold less, about 8-fold less, about 9-fold less, about 10- fold less, about 15-fold less, about 20-fold less, about 30-fold less, about 40-fold less, about 50-fold less, about 100-fold less, or about 1000-fold less) than the doses commonly known in the art for gene editing.
Parenteral administration of the composition, if used, is generally characterized by injection. Injectables can be prepared in conventional forms, either as liquid solutions or suspensions, solid forms suitable for solution of suspension in liquid prior to injection, or as emulsions. A more recently revised approach for parenteral administration involves use of a slow release or sustained release system such that a constant dosage is maintained. See, e.g., U.S. Patent No. 3,610,795, which is incorporated by reference herein.
EXAMPLES
The following examples are set forth below to illustrate the compositions, cells, methods, and results according to the disclosed subject matter. These examples are not intended to be inclusive of all aspects of the subject matter disclosed herein, but rather to illustrate representative methods and results. These examples are not intended to exclude equivalents and variations of the present invention which are apparent to one skilled in the art.
Example 1: Genome-scale CRISPRi screens identify La (SSB) as a strong mediator of prime editing
Prime editing allows precise modification of genomes. To identify cellular determinants of prime editing, scalable prime editing reporters were developed and performed genome-scale CRISPR-interference screens. From these screens, a single factor emerged as the strongest mediator of prime editing: the small RNA-binding exonuclease protection factor La (SSB). Further investigation revealed that La promotes prime editing across approaches (PE2, PE3, PE4, PE5), edit types (substitutions, insertions, deletions), guide RNA designs (pegRNAs, epegRNAs), endogenous loci, and cell types but has no consistent effect on genome editing approaches that rely on standard, unextended guide RNAs. La binds polyuridine tracts at the 3' ends of RNA polymerase III transcripts and protects those transcripts from cellular exonucleases. Accordingly, functional interaction were observed between La and the 3' ends of polyuridylated prime editing guide RNAs. Guided by these insights, a strategy was developed to improve prime editing, namely fusing the RNA-binding, N-terminal domain of
La to the prime editor protein PEmax. Application of the editor dramatically increased prime editing efficiencies. The results provide key insights into how prime editing components interact with the cellular environment and suggest general strategies for stabilizing exogenous small RNAs therein.
Prime editing minimally consists of an engineered Cas9 protein (Cas9 H840A nickase fused to a reverse transcriptase) and a prime editing guide RNA (pegRNA) that specifies both the DNA target and the intended edit (FIG. 1 A). As with other CRISPR-Cas systems, the editor protein binds its cognate guide RNA and, directed by the spacer sequence, finds a complementary DNA target. Once bound to the target, the editing complex nicks the displaced DNA strand and releases a 3' DNA end. This end can then hybridize to the 3' extension of the pegRNA and prime reverse transcription of the pegRNA-encoded edit, which is ultimately incorporated into the genome or removed by DNA mismatch repair (MMR). Several features that impact prime editing have already been reported, including the expression, stability, localization and activity of editing components, as well as chromatin context of target loci. Additionally, demonstrating that mechanistic understanding can reveal avenues for technological improvement, Previous results showed that small prime edits can be installed with higher efficiency and precision when MMR is suppressed or evaded. Studies of prime editing to date, however, have been limited in focus, with inquiry restricted to optimization of editing components or examination of inferred cellular determinants (e.g., DNA repair). By interrogating prime editing with unbiased, genome-scale CRISPR-interference (CRISPRi) screens, an unanticipated mediator of prime editing was identified: the small RNA-binding protein La (SSB). Subsequent characterization of this factor, then showed how exploiting an interaction between La and pegRNAs can dramatically enhance prime editing.
Genetic screens have been performed previously to study prime editing, but previous efforts have been limited to genes associated with DNA repair processes. To elucidate cellular determinants of prime editing with less bias, it was sought to perform genome-scale genetic screens — which have yet to be realized for this or any other CRISPR-based technology. For screening, a prime editing reporter system in which installation of an intended edit ‘switches on’ a reporter gene was developed (FIG. IB). By design, this system expresses a single bicistronic mRNA but, due to lack of a properly positioned start codon, produces only a constitutive marker protein (driven by an internal ribosome entry site) until an upstream, inframe ATG is edited into a defined target site to induce expression of a different reporter gene. To allow the reporter system to be paired with CRISPRi, which relies on Streptococcus pyogenes Cas9 (SpCas9), the system was designed for use with an orthogonal Staphylococcus
aureus Cas9 (SaCas9)-based prime editor (SaPE2). Specifically, two SaPE2 protospacers in the target site were included: one for ATG installation and another at which a +50 complementary strand nick can be introduced. Such nicks have been shown to enhance prime editing, and their inclusion, by use of additional single guide RNAs (sgRNAs), constitutes the PE3 approach. Prime editing without a complementary strand nick, on the other hand, is called the PE2 approach.
Two versions of the reporter system were built: one that uses the fluorescent protein EGFP to report on editing and another that uses a synthetic cell surface protein (IgK-hlgGl-Fc- PDGFRP) (FIGS. 6A-6B). The gene products encoded by each of these reporters were chosen to allow efficient isolation of successfully edited, marker-positive cells: GFP through fluorescence-activated cell sorting (FACS reporter) and the surface protein via magnetic cell separation with protein G beads (MCS reporter). Each of these reporters were transduced into K562 cells constitutively expressing CRISPRi machinery (K562 CRISPRi cells) and, to validate their performance, edited the resulting cells with substitution or insertion edits designed to install one or more start codons (FIG. 6C). After editing, the FACS reporter produced a clear population of GFP+ cells (FIG. 1C). Two observations also demonstrated that that the percentage of those marker-positive cells faithfully reports intended prime editing efficiency: (1) perturbation of MSH2, an MMR gene known to suppress small substitution edits, increased GFP+ percentage (FIG. 1C) and (2) PE3-based editing, which is more efficient than PE2, showed higher GFP+ percentage (FIG. 6D). Additionally, confirming reporter accuracy, quantification of editing outcomes from GFP+ and GFP- populations of FACS reporter cells separated by flow cytometry, and from MCS reporter cells that either bound protein G beads or did not, revealed enrichment of intended edits in GFP+ and bead-bound cells, respectively (FIGS. 6E-6F).
Given these results, the study proceeded to genome-scale screening. Briefly, FACS reporter cells were transduced with the hCRISPRi-v2 library (18,905 targeted genes, 5 sgRNAs per gene), introduced prime editing components by plasmid transfection (SaPE2, +7 GG to CA pegRNA, +50 nicking sgRNA), and separated resulting GFP+/- populations. Flow cytometry analysis prior to sorting confirmed reasonable editing efficiencies (FIG. 6G) and sequencing of the target site showed expected enrichment in sorted populations (FIGS. 6H-6I). To identify genes that, when depleted, strongly alter prime editing efficiency, the relative enrichment or depletion of each CRISPRi sgRNA across GFP+/- populations was determined by sequencing (FIGS. 7A-7B) and then calculated gene-level phenotypes. From this analysis, identified were 35 negative regulators and 1 positive regulator, La (FDR < 0.01, CRISPhieRmix pipeline) (FIG.
ID, FIG. 7C). Due to the relative ease of cell separation with the MCS reporter, MCS-based, genome-scale screens, were performed specifically using PE3 and two enhanced systems of prime editing called PE4 and PE5, which are PE2 and PE3, respectively, but with inclusion of a dominant negative MMR protein (MLHldn). Data from these screens were noisier, with higher technical variability (Methods), but the results confirmed regulators from the first screen, including MMR genes (MSH2, MSH6, MLH1, PMS2) and ones with unknown roles (CASP8AP2, P0LR1D) (FIGS. 7D-71). Additionally, across all screens a single gene displayed the strongest negative phenotypes: La (SSB) (FIG. ID, FIGS. 7B-7C, 7G-7I).
Example 2: La promotes prime editing across pegRNA and editor designs, programmed edits, endogenous targets, and cell types
La, a ubiquitously expressed eukaryotic protein, is involved in diverse aspects of RNA metabolism, but one of its most characterized roles is binding polyuridine (polyU) tracts at the 3' ends of nascent RNA polymerase III (Pol III) transcripts and protecting them from exonucleases. Because the genome-scale CRISPRi screens relied on Pol Ill-transcribed pegRNAs, the La phenotypes observed therefrom may represent an interaction between La and the pegRNA used for screening. However, before evaluating this possibility, it was sought to validate the effect of La on prime editing. Using the reporter system and two La-targeting CRISPRi sgRNAs, each of which depleted La mRNA by >89% (FIG. IE), it was confirmed that cells with low La expression were compromised for prime editing (FIGS. 1F-1H), specifically observing strong defects with PE2, PE3, PE4, and PE5 approaches using the SaPE2 editor with two ATG-specifying pegRNAs (+7 GG to CA, +1 21-bp His-tag insertion) and comparitively weaker defects when editing with one ‘engineered’ pegRNA design called epegRNA (discussed below). Additionally, by subcategorizing unwanted outcomes (‘indels’) from experiments with the MCS repoprter into two bins according to the presence or absence of co-occurring intended edits, it was found that La promotes intended edits with and without accompanying indels but not outcomes with indels alone (FIG. 1H). This observation implies that La does not modulate DNA target recognition or target site nicking but rather incorporation of the edit. Strong phenotypes with the 21-bp insertion edit, which due to its length should not be impacted by MMR, and with approaches that involve MMR suppression (PE4 and PE5) also indicated that the role of La in prime editing is orthogonal to MMR.
Next tested was the impact of La on prime editing at several endogenous loci using an optimized SpCas9-based prime editor, PEmax. For these experiments, a K562 cell line, was engineered, that constitutively expresses PEmax from the AAVS1 safe-harbor locus (K562
PEmax cells) and derived La-knockout clones (La-kol - La-ko5) (FIG. 2A, FIGS. 8A-8C). Consistent with previous validation results, PE2- and PE4-based intended editing efficiencies were significantly lower in La-knockout cells than parental cells, when evaluated with both pegRNAs and epegRNAs (FIGS. 2B-2C). Demonstrating that these defects were due to loss of La, but not unrelated differences arising from clonal isolation, ectopic expression of La rescued intended editing with DNMT1 +5 G to T (FIG. 2C), and no obvious relationship was observed between editing efficiencies and several measured features of the La-knockout lines (z.e., cell growth and PEmax expression) (FIGS. 8C-8E). Next investigates was if the role of La in prime editing is cell- or edit-type specific by evaluating PE3 in HEK293T cells transfected with La-targeting or non-targeting siRNAs (FIGS. 2D-2E, FIG. 8F). Amplicon sequencing of five genomic loci, each targeted with a substitution and an insertion or deletion edit, revealed decreased intended editing efficiencies in La-depleted cells, with a median reduction of 41% across loci and edits. Phenotypes from this experiment were generally weaker than those observed with La-knockout cells, but it was found that La expression rebounded from RNAi-mediated depletion during editing (FIG. 2D), which, together with the observation that ectopic La expression increased editing in parental cells (2.6- and 1.7- fold with a pegRNA and an epegRNA, respectively) (FIG. 2C), suggests a gene dosage effect.
Throughout these experiments, both pegRNAs and epegRNAs were tested. The latter contain structured motifs at their 3' ends and have been shown to enhance prime editing, with improvements loosely attributed to pegRNA stabilization. It was found that La promoted editing with both pegRNAs and epegRNAs, but phenotypes were consistently stronger with pegRNAs (FIGS. 1F-1G, FIGS. 2B-2C, 2E, FIG. 8F). This difference fits a model wherein La promotes editing by interacting with the 3' ends of (e) pegRNAs but has a stronger effect on pegRNAs, which may be less stable or more accessible to La due to less structured 3’ ends.
Example 3: The effect of La on prime editing does not extend to other editing modalities
Prime editing relies on pegRNA 3' extensions, which encode intended edits, but other editing modalities such as nuclease-mediated gene disruption and base editing do not. This difference was a prompt to examine the effects of La on other genome editing approaches. As a preliminary test, SaCas9 was used to induce DNA double-strand breaks (DSBs) in the MCS prime editing reporter using the +7 GG to CA pegRNA, which targets SaCas9 to a locus near a transduced GFP marker gene but not directly within sequences required for expression (FIG. 2F). Because Cas9-induced DSBs often generate large deletions, such breaks can disrupt nearby reporter genes, even when those genes are distant from the target site and especially
when targeting transduced lentiviral constructs. After editing with SaCas9, an observed loss of GFP in the MCS reporter cells, indicating large deletions, but neither GFP loss nor the frequencies of small, DSB-induced indels at the target site were significantly altered upon La depletion (FIG. 2F, FIGS. 9A-9B), suggesting that La had no effect on either type of outcome. Next investigated was if La impacts editing with SaCas9-induced DSBs and two base editing systems at endogenous loci, four genomic targets were selected where a single pegRNA was able to elicit editing with SaCas9, SaBE4-Gram, SaABE8e, and SaPE2 with the PE4 approach, plasmids encoding each of those pegRNAs or an sgRNA with the same spacer (together with other editing components) were transfected into La-knockout cells and parental controls. Targeted sequencing revealed that loss of La had by far the strongest and most consistent effect on prime editing and only moderate, inconsistent effects on other approaches, which were further dampened when editing with sgRNAs (FIGS. 2G-2H, FIGS. 4C-4F). Given these results, it was concluded that La has a specific effect on prime editing.
Example 4: La promotes prime editing by interacting with the 3' ends of polyuridylated pegRNAs
In addition to binding polyU tracts at the 3' ends of Pol III transcripts and protecting them from exonucleases, La has been tenuously implicated in Pol Ill-mediated transcription, with phosphorylation of a single residue (S336) potentially involved in transcriptional modulation via Pol III recycling. To explore the possibility that La affects prime editing through transcriptional effects (z.e., pegRNA expression), various La mutants were examined. La is a 408-residue protein consisting of a highly conserved La motif, two RNA recognition motifs (RRMs), and a flexible region with a nuclear localization signal (NLS) at the C-terminus (FIG. 3 A). The N-terminal domain of La (La1-194), which contains the La motif and RRM1, is necessary and sufficient for high-affinity binding to 3' polyU, while regulation of Pol III recycling has been attributed to the phosphorylation status of Ser366 (S366). It was reasoned that if La promotes prime editing through transcription, truncation of the C-terminal domain or mutation of S366 could abolish or alter its impact, but if La promotes prime editing by binding to the 3' ends of pegRNAs, La1-194 alone should be sufficient for that activity. Evaluating DNMT1 +5 G to T editing in cells transfected with La, two S366 mutants (S366D/G), or La1- 194 fused to an NLS in different configurations revealed that each mutant (five total) rescued PE2 editing in La-knockout cells to levels higher than those observed in parental cells without ectopic La (or mutant) expression (FIG. 3B). Additionally, each La1-194 construct was sufficient for rescue, although S366 mutants and full-length La promoted prime editing to even higher
levels. These results indicate that La promotes prime editing primarily through the N-terminal domain, with contribution from the C-terminus, but little to no contribution from S366.
Having established the importance of La1'194 for prime editing, which binds polyU at the 3' ends of RNA oligomers with nanomolar affinity in vitro, next explored was the hypothesis that La promotes prime editing through interaction with the 3' ends of polyuridylated pegRNAs. To test this idea, several sets of synthetic pegRNAs with or without polyU tails and different patterns of 3' chemical modifications, including 2'-O-methylation (2'- OMe) and phosphorothioate linkages (*) were designed (FIG. 3C). Two considerations guided design of these pegRNAs: (1) Previous biochemical characterization of La1-194-RNA binding has shown that replacing the ribose 2'-hydroxyl group (2'-OH) of the most terminal uridine on an RNA substrate with 2'-0Me disrupts La binding (>30-fold reduction), presumably by creating a steric block. Thus, while addition of a polyU should promote interaction with La, inclusion of terminal 2'-0Me should disrupt the interaction (FIG. 3C). (2) Chemical modifications also confer resistance to RNA exonucleases and are thus typically included in synthetic guide RNAs to boost editing efficiencies59,60. Editing with the first two sets of synthetic pegRNAs confirmed this effect, specifically demonstrating that pegRNAs with 3' chemical modifications produced generally higher editing efficiencies than those without such modifications (FIGS. 10A-10D).
Editing with the first two sets of synthetic pegRNAs also revealed a clear trend among pegRNAs with 3' chemical modifications, those with modifications near their 3' ends but upstream of unmodified polyU tails (‘La-accessible’) were strongly compromised for intended editing in the absence of La, while those with a terminal 2'-0Me and with or without a polyU tail (‘blocked’ and ‘no-polyU’, respectively) were minimally or not impacted by La loss (FIGS. 10C-10D). These results establish an association between the capability of pegRNAs to bind La and their reliance on La for editing. Testing pegRNAs with additional end configurations then confirmed that La strongly impacts editing efficiency when the last 2'-OH of an appended polyU is kept unmodified, suggesting that even one ‘normal’ uridine at the 3' terminus is sufficient for La engagement (FIG. 3D). To evaluate the generality of these observations, four genomic loci were next edited with pegRNAs terminating in a La-accessible end (UU*mU*mU*mUU), a blocked end (UUU*mU*mU*mU), or a no-polyU end (N*mN*mN*mN). Results from this experiment confirmed that loss of La strongly reduces editing efficiencies with the La-accessible end but has less or no effect with the other two end configurations (FIG. 3E, FIG. 10E). Altogether, these results show that endogenous La promotes prime editing when pegRNA ends are expected to bind La1-194 and independent of
pegRNA transcription (FIG. 3F). Moreover, because several La-accessible pegRNAs produced modestly higher intended editing efficiencies compared their blocked or no-polyU conterparts in the presence of La (FIGS. 10C-10E), these data provide motivation to further explore interaction between La and pegRNA 3' ends as a means to improve prime editing.
Example 5: La protects the 3' ends of pegRNAs
Recent studies have shown that 3' ends of pegRNAs are actively degraded within cells and that truncated pegRNAs, resulting from such degradation can interfere with prime editing30. To explore the possibility that interaction with La impacts this process, small RNA sequencing (small RNA-seq) of (e)pegRNAs expressed in the K562 PEmax parental and La-knockout (La- ko4) cells was performed. Two sets of plasmids encoding eleven (e)pegRNAs were trasnfected, each targeting one of seven genomic loci and, after one and two days, prepared and sequenced small RNA libraries from isolated total RNA. By aligning paired-end reads to reference (e)pegRNA sequences and GRCh38, resulting RNA fragments were designed and categorized, with each fragment representing an RNA molecule whose sequence was either determined from an overlapping read pair or inferred from alignment. The majority of these fragments were categorized as snoRNA, retained intron, IncRNA, rRNA or miRNA (FIGS. 11A-11B), but (e)pegRNA fragments were also readily identifiable, with 97.4 ± 0.7% uniquely mapped to one of the eleven (e)pegRNAs.
Cursory evaluation revealed that uniquely mapped (e)pegRNA fragments had distinct features, prompting further categorization into three mutually exclusive bins (FIG. 3G). Fragments with at least 15 nt of spacer sequence and a defined region of the sgRNA scaffold (z.e., the repeat: anti -repeat duplex and stem loop 1) were categorized as ‘c/.s-active’ (orange). These fragments represent pegRNAs with the minimally required sequence for editor protein binding and efficient target engagement and are thus necessary but not sufficient for prime editing in cis. Remaining fragments with the edit-encoding nucleotide and at least the first 5' nucleotide of the primer binding site (PBS) were defined as 7ra//.s-active’ (purple). Given recent reports that untethered templates can enable efficient prime editing, it was reasoned that this sequence represents a minimal requirement for templating prime editing in trans. Finally, remaining fragments were called ‘inactive’, including those ending at cryptic terminator sequences (green asterisks), which are unlikely to impact editing.
For each (e)pegRNA and bin, the number of fragments (coverage), were counted at each nucleotide position and normalized those counts in two ways: either to the number of fragments within the corresponding bin (relative coverage) or to the number of fragments from
the same sample assigned to human RNA (absolute coverage). Examining coverages across (e)pegRNA first demonstrated that, independent of the presence or absence of La, c/.s-active fragments contained the entirety of their spacer and scaffold sequences (including sequences outside of the 58-nt region that defined the category) but as early as one day post transfection, lacked a large portion of their 3' extensions (FIGS. 3H-3I, FIG. 11D, FIG. 12A-12C). While this observation is similar to published results demonstrating loss of 3' extensions by Northern blotting, the magnitude of the effect was surprisingly strong for many of the (e)pegRNAs. Moreover, steep, reproducible decreases were observed in coverages at particular nucleotides for both pegRNAs and epegRNAs, suggesting stepwise or heterogeneous end processing. These observations support the idea that sequences bound by the Cas9 nickase are protected, while exposed 3' extensions are highly unstable.
Comparing the relative coverages of the cv.s-active (e)pegRNA fragments (normalized per bin) from parental cells to those from La-ko4 cells next showed that loss of La preferentially reduced coverages within the 3' extensions of both pegRNAs and epegRNAs, suggesting that 3' extensions are more actively degraded without La (FIGS. 3H-3I, FIGS. 1 ID, FIG. 12). The most striking of these effects were observed for (e)pegRNAs with appreciable coverage of their 3' extensions in parental cell and thus a reasonable baseline for comparison (RUNX1 +5 G to T and VEGFA +5 G to T), but differences were also apparent for those (e)pegRNAs with minimal extension coverages in parental cells. Evaluating the absolute coverages of (e)pegRNA fragments (normalized per sample), as well as performing expression analyses with all fragments, then demonstrated that absolute levels of (e)pegRNA fragments were also affected by loss of La, with evident reduction of cv.s-active, /ra/z.s-active and premature termination fragments in the La-ko4 cells (FIGS. 3H-3J, FIGS. 11C-11D, FIG. 12). These observations show that loss of La destabilizes (e)pegRNAs and, consistent with an interaction between La1-194 and (e)pegRNA polyU tails, renders their 3' ends particularly susceptible to degradation. Dramatic loss of 3' extensions in both the presence and absense of La and variability in truncation patterns, though, complicate functional interpreation of these phenotypes (Discussion).
By design, prime editing generates RNA-DNA hybrids during editing. Cellular RNases (RNase H1/H2) or the intrinsic RNase H activity of the MMLV reverse transcriptase (MMLV- RT) in PEmax may therefore be responsible for some of the dramatic 3 ' end degradation that was observed. To evaluate the effect of La on (e)pegRNAs without such effects, a small RNA- seq was performed on a pegRNA and epegRNA pair (Mus DNMT1 +6 G to C) with no obvious genomic target in human cells (FIGS. 13A-13D). In stark constrast to actively editing
(e)pegRNAs, a majority of cz.s-active fragments from the non-targeting pair retained 3' extensions in K562 PEmax cells one day post transfection, and truncation patterns were nearly bimodal (z.e., fragments were mostly intact or had extensions fully truncated). Preferential loss of extension sequences from cz.s-active fragments was neverthless again observed in La-ko4 cells. These results thus also support an interaction between La and the 3' ends of (e)pegRNA, but because destabilization phenotypes for the pegRNA and epegRNA were of vastly different magnitudes while both of them demonstrated strong dependence on La for editing when an exogenous target was provided (FIG. 13E), it was reasoned that destabilization may be exacerbated by the presence of the target or that stabilization of (e)pegRNAs is only part of La’s role in prime editing.
Example 6: Fusing N-terminal domain of La to PEmax dramatically improves prime editing efficiency
Given evidence that La promotes prime editing primarily through La1-194, it was next asked if tethering that domain to the prime editor protein could offer improvement. Remarkably, fusing full-length La or La1-194 to PEmax in multiple positions (z.e., N-terminus, C-terminus, or between Cas9 nickase and MMLV-RT) significantly improved editing efficiencies when evaluated with the PE2 approach using pegRNA and/or epegRNA in two different cell lines (FIG. 4A-4B). Among constructs with full length La, highest median intended editing was achieved with an internal fusion (PE-Lmax-2), and among La1-194 fusion constructs, a C- terminal fusion (PEmax-C) was the most efficient. Subsequent characterization of the latter, which was named PE7, revealed dramatic improvement over PEmax across eight genomic loci in three different cell lines (293T, HeLa, U2OS) and three distinct edit types (single nucleotide substitutions, two insertions, a 15-bp deletion), with the biggest improvements observed in MMR proficient HeLa and U2OS cells (FIGS. 4C-4E). In particular, PE7 improved intended editing efficiencies in U2OS cells by 21.2- and 5.5-fold (median) compared to PEmax when using the PE2 approach with pegRNAs and epegRNAs (respectively), thus increasing absolute levels of PE2 editing from extremely low to potentially actionable (FIGS. 4D-4F). Consistent with previous studies, epegRNAs increased baseline editing with PEmax, but interestingly, these modified reagents offered no additional improvement over pegRNAs when using PE7, with no signficant observed at four genomic loci (RUNX1, FANCF, HEK4, EMX1, P < 0.05, two-tailed unpaired Student’s /-test) and significant decrease in intended editing with epegRNAs at four other sites (RNF2, DNMT1, HEK3, VEGFA, P < 0.05, one-tailed unpaired Student’s /-test), suggesting that, if the underlying mechanisms by which epegRNA and PE7
improve prime editing are similar (e.g., reagent stabilization), the latter has a saturating or combinatorically negative effect. Additionally, despite moderately increasing indel frequencies, PE7 did not increase those outcomes proportionally with the intended edits and unwanted outcomes at the target sites remained very low.
To confirm that the effect of PE7 on prime editing was due to the RNA-binding activity of the fused La1'194, a PE7 mutant with four mutations previously shown to disrupt interaction between La1'194 and polyuridylated RNA oligomers was generated (FIGS. 4G-4H). Supporting the model that La promotes prime editing by binding pegRNA 3' ends (FIG. 3F), these mutations completely abolished improvements from fusing La1-194 to PEmax, when evaluated with four edits in two cell lines (U2OS and K562) (FIG. 41, FIG. 14A).
It was next asked if PE7 causes deleterious effects on cell growth or increases off-target editing. Expression of PE7 in K562 cells produced negligible changes to cell viability and caused no significant difference in the number of population doublings observed during editing, relative to PEmax and PE7 mutant (FIGS. 14A-14C). Moreover, gene expression analysis of cells transfected with PEmax, PE7, or PE7 mutant together with PRNP- or HEK3 -targeting pegRNAs produced minimal changes to the cellular transcriptome, with no genes demonstrating >2 -fold up or downregulation in any comparisons, and only four genes similarly and significantly changed in more than one comparison (FIGS. 14D-14I). To evaluate off- target editing, several of the most common Cas9 off-target sites associated with four of the prime edited loci (EMX1, FANCF, HEK3, HEK4) were sequenced. These genomic loci have been used previously to demonstrate the specificity of prime editing. Accordingly, editing with PEmax in U2OS cells produced few changes at these sites (median 0.053%), and compared to PEmax, PE7 increased off-targeting significantly at only two of the thirteen, suggesting PE7 generally boosts prime editing activity but few off-target sites are susceptible to that effect.
Example 7: PE7 enhances prime editing of therapeutic-relevant targets and cell types
To evaluate the potential utility of PE7, next, editing at additional genomic targets was evaluated, including ones associated with sickle cell anemia (HBB), prion disease (PRNP), familial hypercholesterolemia (PSCK9), adoptive T cell transfer therapy (IL2RB), HIV infection (CXCR4), and CDKL5 deficiency disorder (CDKL5) (FIGS. 5A-5B). Editing with the PE2 approach at these loci showed similar improvement over PEmax as the panel of eight test edits with both pegRNAs and epegRNAs (median 23.8- and 7.9-fold for pegRNAs and epegRNAs, respectively) (FIGS. 4D, 4F, FIGS. 5A-5B) although one edit (PNRP +6 G to T) for which use of an epegRNA with PE7 strongly outperformed a matched pegRNA provided
an outlier to previous observations and indicated that, depending on sequence or target site, select epegRNAs may synergize with PE7. It was then asked if editing efficiency could be further increased by pairing PE7 with the more efficient PE3, PE4, and PE5 prime editing approaches. Across seven disease-relevant edits (or a subset thereof), PE7 produced median 10.4-, 7.7-, and 5.7-fold improvement over PEmax in intended editing with each of those approaches, respectively (median 10.0-, 3.8-, and 8.9-fold increases in indels) (Fig. 5C). Across all or a subset of the previous eight test edits, PE7 also showed improvement (median 6.9-, 6.5-, and 3.3-fold increase in intended editing, median 6.8-, 7.8-, and 7.6-fold increases in indels) (Fig. 4D, Fig. 10B). Remarkably, when paired with the most advanced system (PE5), PE7 achieved 50.2% median intended editing across 8 edits. These results show that PE7 supports substantially higher prime editing efficiency across approaches and genomic loci. Moreover, results with PE4 and PE5 confirm the earlier observations that the effect of La on prime editing is not redundant with MMR.
Next evaluation was of the performance of PE7 with different strategies of editor and (e)pegRNA delivery. Across tests with the PE2 approach, PE7 outperformed PEmax, including when editors were delivered by plasmids or in vitro transcribed mRNA to cells stably expressing pegRNAs or epegRNAs and when both editors and (e)pegRNAs were delivered by lentiviral transduction (FIGS. 15C-15D). The latter demonstrates robustness of PE7 without high-copy delivery. Additionally, mRNA-expressed PE7 yielded higher frequencies of intended editing than PEmax when combined with La-accessible pegRNAs (UU*mU*mU*mUU). By contrast, PE7 and PEmax were similar in performance when paired with no-polyU pegRNA designs (N*mN*mN*mN), further supporting the conclusion that La binding to the 3' ends of pegRNAs promotes prime editing. Contrary to expectations from experiments in La-knockout cells (FIG. 3E), however, PE7 also increased intended editing efficiencies over PEmax with La-blocked pegRNAs, possibly due to enhanced interaction with La-blocked pegRNAs when those elements are in proximity, as in the effector complex or at the site of editing. Relevant to this idea, the binding pocket for 3' polyU within La1-194 does not involve the canonical RNA-binding surface of the constituent RRM1, leaving open the possibility that multiple, typically low-affinity interactions contribute to such an effect.
Finally, it was confirmed that PE7 improves prime editing in primary cells. Consistent with results in K562 and U2OS cells, editing human primary T cells with mRNA expressed PE7 and La-accessible pegRNAs yielded higher frequencies of intended editing than other pairings of mRNA expressed editors and synthetic pegRNAs, overall demonstrating 2.1-, 3.2- and 5.2-fold improvement over standard reagents (z.e., PEmax with no-polyU pegRNAs) at
three different sites (FIG. 5F). When paired with La-accessible pegRNAs, PE7 also achieved a reasonable 20.0% median intended editing efficiency across eight targets with PE2 approach, which represents a median 2.3-fold improvement over PEmax (FIGS. 5F-5G, FIG. 15G). Similarly, editing the HBB locus in primary human haematopoietic stem cells (HPSCs) showed clear advantage for PE7 with La-accessible pegRNAs, although maximal efficiency remained relatively low (FIG. 5H). These data show proof-of-principle for leveraging the RNA-binding activity of La to optimize prime editing in primary cells.
Genome-scale genetic screens were performed aimed at identifying cellular determinants of prime editing and identified La, a small RNA-binding protein, as a strong promoting factor. It was then shown that endogenous La promotes prime editing through its N- terminal domain (La1-194), which is both necessary and sufficient for high-affinity binding to the 3' polyU of RNA molecules. Guided by previous structural and biochemical studies, synthetic pegRNAs, were designed, with features that in principle allow La binding to their 3' ends and show that these pegRNAs functionally interact with La. Motivated by these results, full-length La protein or La1-194 was fused to the PEmax prime editor and found that such fusions improved prime editing efficiency. Further characterization of one such fusion (PE7) demonstrated strong enhancement in efficiency of intended prime editing across cell and edit types, genomic loci, and delivery methods, with infrequent effects on off-target editing and little to no effect on cell growth or gene expression. In particular, with the simplest prime editing approach (PE2), PE7 increased intended editing levels from extremely low by PEmax to potentially actionable and, with more advanced PE5 approach, achieved an impressive 50.2% median efficiency.
The previous results also show that endogenous La can stabilize (e)pegRNAs, which completements an emerging understanding that poor stability of reverse transcription templates (z.e., pegRNA 3' extensions or untethered linear templates) limits prime editing efficiency. However, two observations from the small RNA-seq analyses leave unanswered the question of how La promotes prime editing: (1) Actively editing (e)pegRNA were extensively truncated with or without La, resulting in a mixture of fragments with difficult-to-interpret effects on prime editing (z.e., potentially supportive, neutral, or inhibitory) and/or dramatic loss of 3' extensions (z.e., RNF2 +1 C to A, DNMT1 +5 G to T). These features confound functional interpretation of (e)pegRNA destabilization phenotypes therefrom. (2) Examining (e)pegRNA stability in the absence of their genomic targets, where truncation of cz.s-active fragments was less dramatic in the presence of La, revealed that the stability of one pegRNA (Mus DNMT1 +6 Gto C) was minimally affected by loss of La, demonstrating that (e)pegRNAs do not always
depend strongly on La for stability. Therefore, although it was observed that La promotes (e)pegRNA stability, it cannot be concluded that this effect alone explains the role of La in promoting prime editing. Indeed, La may promote multiple steps in prime editing, including nuclear retention of (e)pegRNAs or effector complex formation. Alternatively, La may impact (e)pegRNA stability in multiple ways, for example offering variable protection from cellular exonucleases independent of editing activity and safeguarding (e)pegRNAs from a targetdependent mode of degradation (possibly by RNase H). Independent of the exact mechanism, insights provided herein will enable future studies of (e)pegRNA stability.
Previous efforts to mitigate poor pegRNA stability include adding structured RNA motifs to the 3' ends of pegRNAs, as in epegRNAs, and circularizing untethered templates. Suggesting that the role of La in prime editing is at least partially redundant with epegRNAs, it was found that, compared to pegRNAs, epegRNAs buffer La-associated editing phenotypes. However, when editing with PE7, epegRNAs provided no additional benefit except in a minority of cases, suggesting that, if PE7 and epegRNAs improve editing through the same mechanism, the former often has a saturating effect. From an application standpoint, this offers practical benefit, because while modifying reverse transcription templates can improve editing, such approaches complicate reagent procurement, especially with RNA delivery, due to requiring chemical synthesis of significantly longer RNA molecules or circularization. Nevertheless, for select applications, defining when and how epegRNAs offer additional benefit over pegRNAs with PE7 may also prove beneficial.
More subtly, the study highlights how end modification strategies developed to protect synthetic sgRNAs from RNA exonucleases have been somewhat haphazardly applied to pegRNAs. Unlike sgRNAs, which are almost entirely protected by bound Cas9 proteins, pegRNAs rely on exposed 3' extensions. Therefore, it cannot expect chemical modification strategies developed for sgRNAs to be optimal or even sufficient for synthetic pegRNAs. Additionally, although chemically synthesized guide RNAs can be produced without a 3 ' polyU, terminal uridines are often included with unclear functional consequences, and studies deploying synthetic pegRNAs have arbitrarily included or excluded them. Combined with commercially recommended chemical modifications for sgRNAs, these polyU tails result in pegRNAs that should block (3'-mU*mU*mU from Synthego) or allow (3'-mU*mU*mU*U from Agilent and IDT) La binding, which could have effects on editing even without using PE7, as demonstrated in several of the experiments (e.g., FIG. 5H).
In summary, through the identification and characterization of La as a cellular determinant of prime editing, this study expands understanding of the cellular processes that
directly impact prime editing and demonstrates methods for improving prime editing efficiencies. Discovery of this effector, as well as the results, tools and insights provided herein, will also no doubt aid additional efforts to understand mechanisms of prime editing in exquisite detail.
Example 8: Methods
General methods. CRISPRi sgRNAs were cloned into pU6-sgRNA EFlAlpha-puro- T2A-BFP (Addgene #60955) as described in weissman.wi.mit.edu/resources/sgRNACloningProtocol.pdf. Plasmids for transfection expressing pegRNAs, epegRNAs and non-CRISPRi sgRNAs were cloned by Gibson Assembly of gene fragments without adapters from Twist Bioscience and pU6-pegRNA-GG- acceptor plasmid (Addgene #132777) digested by Ndel or BsaAI/BsaI-HFv2 (New England Biolabs R0111S, R0531S, R3733S). Plasmids for transduction expressing pegRNAs and epegRNAs were cloned by Gibson Assembly of gBlock from Integrated DNA Technologies and pU6-sgRNA EFlAlpha-puro-T2A-BFP digested by BstXI and Xhol (New England Biolabs R0113S and R0146S). The FACS and MCS reporter plasmids were cloned by Gibson Assembly with pALD-lentiEGFP-A (Aldevron) as the backbone, IRES2 from pLenti- DsRed lRES EGFP (Addgene #92194) and the synthetic surface marker from pJT039 (Addgene # 161927). The AAVS1 PEmax knock-in plasmid was generated by restriction cloning with a backbone modified from pAAVSl-Nst-MCS (Addgene # 80487), PEmax editor from pCMV-PEmax (Addgene #174820) and IRES2 from pLenti-DsRed lRES EGFP. Plasmids of PEmax fused to La or La N-terminal domain were generated by restriction cloning using pCMV-PEmax as the backbone (linker A, SGGSx2-XTEN16-SGGSx2; linker B, SGGSx2-bpNLSSV40 -SGGSx2; linker C, SGGS). pCMV-PE7-P2A-hMLHldn was cloned by Gibson Assembly with pCMV-PE7 as the backbone and insert fragment PCR amplified from pCMV-PEmax-P2A-hMLHldn (Addgene #174828). pCMV-PE7-mutant (Q20A/Y23A/Y24F/F35A) was cloned by Gibson Assembly with pCMV-PE7 as backbone and mutation-containing gene fragment without adapters from Twist Bioscience. pT7-PE7 for IVT was cloned by Gibson Assembly with pT7-PEmax for IVT (#178113) as the backbone and insert fragment PCR amplified from pCMV-PE7. Lentiviral transfer plasmids expressing PEmax or PE7 with IRES2-driven EGFP or EGFP-T2A-NeoR as selectable marker were cloned by Gibson Assembly with pU6-sgRNA EFlAlpha-puro-T2A-BFP as the backbone, UCOE and SFFV promoter from pMHOOOl (Addgene #85969), IRES2 from pLenti- DsRed_IRES_EGFP and T2A-NeoR from pAAVSl-Nst-MCS. FACS/MCS reporter plasmids,
AAVS1 PEmax knock-in plasmid, pCMV-PE7, pCMV-PE7-P2A-hMLHldn and pT7-PE7 for IVT will be available on Addgene. DNA amplification for molecular cloning was performed with Platinum SuperFi II PCR Master Mix (Invitrogen 12368010). Plasmids were extracted with NucleoSpin Plasmid, Mini kit (Macherey-Nagel 740588.250), ZymoPURE II Plasmid Midiprep Kit (Zymo Research D4201) or EndoFree Plasmid Maxi Kit (Qiagen 12362). Primers were ordered from Integrated DNA Technologies.
In vitro transcription of prime editor mRNA. Prime editor mRNA was in vitro transcribed as previously described. Plasmids with PEmax or PE7 coding sequence flanked by an inactivated T7 promoter, a 5' untranslated region (UTR) and Kozak sequence in the upstream as well as a 3' UTR in the downstream were purchased from Addgene (pT7-PEmax for IVT) or cloned as described above (pT7-PE7 for IVT). In vitro transcription templates were generated by PCR to correct T7 promoter and install a 119-nt poly(A) tail downstream of the 3' UTR. PCR products were purified by DNA Clean & Concentrator-5 (Zymo Research D4003) and SPRI selection (Beckman Coulter B23317) for cell line and T cell experiments respectively and stored at -20 °C until further use. mRNA was generated using Hi Scribe T7 mRNA Kit with CleanCap Reagent AG (New England BioLabs E2080S) for cell line experiments and HiScribe T7 High Yield RNA Synthesis Kit (New England Biolabs E2040S) in the presence of RNase Inhibitor (New England Biolabs M0314L) and Yeast Inorganic Pyrophosphatase (New England Biolabs M2403L) for T cell experiments. All mRNA was produced with UTP fully replaced with N^Methylpseudouridine-S'-triphosphate (TriLink Biotechnologies N-1081) and co-transcriptional capped by CleanCap AG (TriLink Biotechnologies N-7113). Transcribed mRNA was precipitated by 2.5 M lithium chloride (Invitrogen AM9480), resuspended in Nuclease-Free Water (Invitrogen AM9939), quantified by a NanoDrop One UV-Vis spectrophotometer (Thermo Scientific), normalized to 1 pg pL-1 and stored at -80 °C. mRNA for T cell experiments was additionally quantified by Agilent 4200 TapeStation.
General mammalian cell culture conditions. Lenti-X 293 T was purchased from Takara (632180). K562 (CCL-243), HeLa (CCL-2) and U2OS (HTB-96) were purchased from ATCC. K562 CRISPRi cell line constitutively expressing dCas9-BFP-KRAB (pHR-SFFV- dCas9-BFP-KRAB, Addgene #46911) was a gift from Jonathan Weissman. Lenti-X 293T, HeLa and U2OS were cultured and passaged in Dulbecco’s modified Eagle’s medium (DMEM) (Corning 10-013-CV), DMEM (Corning 10-013-CV) and McCoy's 5A (Modified) Medium (16600082) supplemented with 10% (v/v) fetal bovine serum (Corning 35-010-CV) and lx Penicillin-Streptomycin (Pen-Strep) (Corning 30-002-CI). For lipofection and nucleofection, lx Pen-Strep was not supplemented. K562 and K562 CRISPRi were cultured and passaged in
RPMI medium 1640 (Gibco 22400089) supplemented with 10% (v/v) fetal bovine serum (Corning 35-010-CV) and lx penicillin-streptomycin-glutamine (Gibco 10378016). For nucleofection, lx penicillin-streptomycin-glutamine was replaced by lx L-Glutamine at 292 pg mL'1 final concentration (Coming 25-005-CI). All cell types were incubated, maintained, and cultured at 37 °C with 5% CO2. Cell lines were authenticated by their respective suppliers or short tandem repeat profiling and tested negative for mycoplasma.
Lentivirus packaging and transduction. To package lentiviruses, Lenti-X 293T were seeded at 9 x 105 cells/well in 6-well plates (Greiner Bio-One 657165) and were transfected at 70% confluency. For transfection, 6 pL TransIT-LTl (Mims MIR 2300) was mixed and incubated with 250 pL Opti-MEM I Reduced Serum Medium (Gibco 31985070) at room temperature for 15 minutes, then mixed with 100 ng pALD-Rev-A (Aldevron), 100 ng pALD- GagPol-A (Aldevron), 200 ng pALD-VSV-G-A (Aldevron) and 1500 ng transfer plasmids at room temperature for another 15 minutes, and was added dropwise to Lenti-X 293T followed by gentle swirling for proper mixing. 10 hours after transfection, ViralBoost reagent (ALSTEM VB100) was added at lx final concentration. 48 hours after transfection, the virus-containing supernatant was collected, filtered through a 0.45-pm cellulose acetate filter (VWR 76479- 040), and stored at -80 °C. Lentiviruses for CRISPRi screens were similarly packaged with hCRISPRi-v2 library (Addgene #83969) as transfer plasmids in 145 mm plates (Greiner Bio- One 639160). For transduction of K562, cells were resuspended in fresh culture medium supplemented with 8 pg mL'1 polybrene (Santa Cruz Biotechnology sc-134220), mixed with lentivirus-containing supernatant, and were centrifuged at 1000 g at room temperature for 2 hours. For transduction of U2OS and HeLa, cell culture was supplemented with 8 pg mL'1 polybrene and lentivirus-containing supernatant. The percentages of transduced (fluorescent protein marker positive) cells were determined by AttueNXT flow cytometry 72 hours after transduction. To generate stably transduced cell lines, cells were selected by 3 pg mL'1 puromycin 48 hours after transduction until >95% of live cells were marker positive.
Construction of FACS reporter cell line and FACS-based genome-scale CRISPRi screen. To construct the FACS reporter cell line, K562 CRISPRi was transduced with FACS reporter lentiviruses at a 0.17 MOI (15.3% infection). The transduced (mCherry+) population was isolated by BD FACSAria Fusion Flow Cytometer and expanded as the FACS reporter cell line. For the FACS-based genome-scale CRISPRi screen, two replicates were independently performed a day apart. For each replicate, 2.4E8 FACS reporter cells were transduced with hCRISPRi-v2 viruses at a 0.29 MOI (25% infection) and were selected by 3 pg mL'1 puromycin 48 hours after transduction. 7 days after transduction, 3.2E8 fully selected
cells were nucleofected with SE Cell Line 4D-Nucleofector X Kit L (Lonza V4XC-1024) and pulse code FF120, according to the manufacturer’s protocol. Each nucleofection consists of 1E7 cells, 7500 ng pCMV-SaPE2 (Addgene #174817), 2500 ng +7 GGto CA pegRNA plasmid and 833 ng +50 nicking sgRNA plasmid. 3 days post nucleofection, 1.5E8 cells were sorted by BD FACSAria Fusion Flow Cytometer. More specifically, cells were first gated on mCherry+ and BFP+, of which EGFP+ and EGFP- populations were collected. Genomic DNA (gDNA) was extracted from both populations with NucleoSpin Blood XL Maxi kit (Macherey-Nagel 740950.50). The entirety of gDNA from both populations was used for PCR amplification of integrated hCRISPRi-v2 sgRNAs. Each 100 pL PCR reaction was performed with 10 pg of gDNA, 1 pM of forward primer (P5 and i7) that anneals in mouse U6 promoter, 1 pM of reverse primer (P7) that anneals to the sgRNA constant region, and 50 pL of NEBNext Ultra II Q5 Master Mix (New England BioLabs M0544X) with the following cycling conditions: 98°C for 30 s, 23 cycles of [98°C for 10 s, 65°C for 75 s], followed by 65°C for 5 min. The PCR product was purified using SPRIselect (Beckman Coulter B23318) with a double size selection (0.65x right side and 1.35x left side), quantified by Qubit IX dsDNA High Sensitivity kit (Invitrogen Q33231) and a high sensitivity DNA chip (Agilent Technologies 5067-4626) on an Agilent 2100 Bioanalyzer, and sequenced with the NovaSeq 6000 SP Reagent kit vl.5 100 cycles (Illumina 20028401) with 50 cycles for R1 read and 8 cycles i7 index read.
Construction of MCS reporter cell line and MCS-based genome-scale CRISPRi screen. To construct the MCS reporter cell line, K562 CRISPRi was transduced with MCS reporter lentiviruses at a 0.09 MOI (8.5 % infection). The transduced (EGFP+) population was isolated by BD FACSAria Fusion Flow Cytometer and expanded as the MCS reporter cell line. MCS-based genome-scale CRISPRi screens with +7 GGto CAPE3+50, PE4 and PE5+50 edits were performed in parallel with two replicates each. 2.1E8 MCS reporter cells were transduced with hCRISPRi-v2 viruses at a 0.16 MOI (15% infection) for the screen conditions and were selected by 3 pg mL'1 puromycin 48 hours after transduction. 7 days after transduction, 1E8 fully selected cells were nucleofected for each replicate of each edit with SE Cell Line 4D- Nucleofector X Kit L (Lonza V4XC-1024) and pulse code FF120, according to the manufacturer’s protocol. Each nucleofection consists of 1E7 cells and varying amounts of plasmids encoding prime editing components. More specifically, for PE2 and PE3, 7500 ng pCMV-SaPE2, 2500 ng +7 GG to CA pegRNA plasmid, 833 ng +50 nicking sgRNA plasmid (PE3) were used per nucleofection. For PE4 and PE5, 6000 ng pCMV-SaPE2, 3000 ng pEFla- hMLHldn (Addgene #174823), 2000 ng +7 GG to CA pegRNA plasmid and 667 ng +50 nicking sgRNA plasmid (PE5) were used. 4 days post nucleofection, cells from each replicate
and condition were magnetically separated into bound and unbound fractions as previously described. The gDNA extraction, PCR, NGS library QC and sequencing were performed as described in the section above. It was noted that the MCS reporter cannot remove dead cells, debris, or doublets from bead-bound or unbound fraction. Therefore, it was less efficient in cell separation than the FACS reporter.
Analysis of genome-scale CRISPRi screen. Sequencing reads were aligned to the human CRISPRi_v2 library (5 sgRNA/gene) using custom Python scripts as previously described (scripts available at github.com/mhorlbeck/ScreenProcessing). sgRNA-level phenotypes were calculated as the log2 enrichment of normalized read counts (sgRNA counts normalized to total counts from the sample and relative to the median of non-targeting controls) within populations of marker-positive cells (GFP+ or bound) compared to marker-negative cells (GFP- or unbound). Prior to calculation, a read count minimum of 50 was imposed for each sgRNA within each sample. Gene-level phenotypes were then calculated for each annotated transcription start site by averaging the phenotypes of the strongest 3 sgRNAs by absolute value. Negative control pseudogenes were generated by random sampling, assigning 5 non-targeting sgRNAs to each pseudogene. sgRNA-level phenotypes were used as input to the CRISPhieRmix pipeline under default parameters with p = 2 to formally evaluate the effect each gene has on prime editing efficiency.
Tissue culture transfection protocols and genomic DNA extraction. For La knockdown in Lenti-X 293 T by siRNA reverse transfection, 120 pmole ON-TARGETplus Human La siRNA (Horizon LQ-006877-01-0005) or ON-TARGETplus Non-targeting Control Pool (Horizon D-001810-10-05) were mixed thoroughly with 500 pL Opti-MEM I Reduced Serum Medium (Gibco 31985070) and 4 pL Lipofectamine RNAiMAX Transfection Reagent (Invitrogen 13778150) in each well of 6-well plates (Greiner Bio-One 657165), incubated at room temperature for 15 minutes before 4E5 Lenti-X 293T in 2.5 mL Pen- Strep-free medium were added. The reverse transfected cells were used for RT-qPCR or downstream prime editing experiments as described in corresponding Methods sections.
For prime editing in Lenti-X 293T by plasmid transfection, 18,000 cells were seeded in 100 pL Pen- Strep-free medium per well in 96-well plates (Nunc 167008). 18 hours after seeding, a 10 pL mixture of 200 ng pCMV-PE2 (Addgene #132775), 66 ng pegRNA, 22 ng nicking sgRNA, 0.5 pL Lipofectamine 2000 Transfection Reagent (Invitrogen 11668027) and Opti-MEM I Reduced Serum Medium (Gibco 31985070) was incubated at room temperature for 15 minutes and added to each well. 72 hours after transfection, culture medium was removed, cells were washed with DPBS (Gibco 14190144) and genomic DNA was extracted
by adding 40 pL freshly prepared lysis buffer into each well. The lysis buffer consisted of 10 mM Tris pH 8.0 (Gibco AM9855G), 0.05% SDS (Invitrogen 15553027), 25 pg mL 1 proteinase K (Invitrogen AM2546) and Nuclease-Free Water (AM9939). The genomic DNA extract was incubated at 37 °C for 90 minutes and transferred into PCR strips (USA Scientific 1402-4700) for 80°C inactivation of proteinase K for 30 minutes in Bio-Rad T100 Thermal Cycler.
For prime editing in Lenti-X 293T, HeLa and U2OS cells by plasmid nucleofection, 750 ng prime editor plasmid and 250 ng pegRNA plasmid were nucleofected. For each sample, 2E5 LentiX-293T cells, 1E5 HeLa cells or 1E5 U2OS cells were nucleofected using SF (Lonza V4XC-2032), SE (Lonza V4XC-1032) and SE Cell Line 4D-Nucleofector X Kit S with program CM-130, CN-114 and DN-100, respectively, according to the manufacturer’s protocols. After nucleofection, cells were cultured in 24-well plates and 72 hours post nucleofection, culture medium was removed, cells were washed with DPBS (Gibco 14190144) and genomic DNA was extracted by adding 110 pL freshly prepared lysis buffer described above into each well. The genomic DNA extract was incubated at 37 °C for 90 minutes and transferred into PCR strips (USA Scientific 1402-4700) for 80°C inactivation of proteinase K for 40 minutes in Bio-Rad T100 Thermal Cycler.
For nucleofections in K562 (except CRISPRi screens, AAVS1 knock-in, La knockout and small RNA sequencing), 1E6 cells were nucleofected with specified amounts of plasmids or synthetic guide RNAs using SE Cell Line 4D-Nucleofector X Kit S (Lonza V4XC-1032) and program FF-120, according to the manufacturer’s protocol. For testing FACS- and MCS- reporter and validation of La phenotype in reporter lines, 900 ng pCMV-SaPE2, 300 ng pegRNA plasmid, 100 ng nicking sgRNA plasmid (PE3/5) and 450 ng pEFla-hMLHldn (PE4/5) were nucleofected. For validation of La phenotype in K562 PEmax and La-knockout clones, 500 ng pegRNA plasmid was nucleofected. For rescue experiments, 500 ng pegRNA plasmid and 1000 ng plasmid encoding La, La mutants or mRFP control were nucleofected. For SaCas9 cutting in MCS reporter cells, 800 ng pX600 (Addgene #61592) and 400 ng +7 GG to CA pegRNA plasmid were nucleofected. For SaPE4 editing in K562 PEmax parental and La-ko4 cells, 800 ng pCMV-SaPE2, 400 ng pegRNA plasmid and 400 ng pEFla- hMLHldn were nucleofected. For SaCas9, SaBE4 and SaABE8e editing in K562 PEmax parental and La-ko4 cells, 400 ng pegRNA or sgRNA plasmid together with 800 ng pX600, SaBE4-Gram (Addgene #100809) or SaABE8e (Addgene #138500) were nucleofected. Synthetic pegRNAs and nicking sgRNAs with specified sequences and modifications were ordered as Custom Alt-R gRNA from Integrated DNA Technologies (Table 3). According to
an incremental titration of a DNMT1 +5 G to T synthetic pegRNA with standard chemical modifications in K562 PEmax parental cells, intended editing efficiencies were already saturated at 100 pmole input (FIG. 10B). Therefore, 100 pmole synthetic pegRNA and 50 pmole nicking sgRNA (PE3) were used for nucleofection unless otherwise specified. 72 hours post nucleofection, 1E6-2E6 cells were harvested in 1.5 mL tubes (Eppendorf 0030123611), washed with 1 mL DPBS (Gibco 14190144) and resuspended in 100 pL freshly prepared lysis buffer described above. The genomic DNA extract was incubated at 37 °C for 120 minutes and transferred into PCR strips (USA Scientific 1402-4700) for 80°C inactivation of proteinase K for 40 minutes in Bio-Rad T100 Thermal Cycler.
For prime editing in K562 and U2OS cells using editor mRNA and synthetic pegRNA, 1E6 K562 and 1E5 U2OS cells were nucleofected with 1 pg editor mRNA and 50 pmole synthetic pegRNA using SE Cell Line 4D-Nucleofector X Kit S (Lonza V4XC-1032) with program FF-120 and DN-100, respectively, according to the manufacturer’s protocols. After nucleofection, cells were cultured for 72 hours and harvested for genomic DNA extract.
For prime editing in HeLa and U2OS cells by lentiviral delivery of (e)pegRNAs and nucleofection of editor plasmids or mRNA, cells were transduced with lentiviruses expressing (e)pegRNAs (20-40% infection) and were fully selected by 3 pg mL'1 puromycin. Stably transduced HeLa and U2OS cells were nucleofected with 750 ng editor plasmid or 1 pg editor mRNA using SE Cell Line 4D-Nucleofector X Kit S (Lonza V4XC-1032) with program CN- 114 and DN-100, respectively, according to the manufacturer’s protocols. After nucleofection, cells were cultured for 72 hours and harvested for genomic DNA extract.
For prime editing in K562 cells by lentiviral delivery of prime editors and (e)pegRNAs, K562 cells were transduced with lentiviruses expressing PEmax or PE7 (with IRES2-driven EGFP or EGFP-T2A-NeoR as selectable marker). The transduced populations (EGFP+, 20- 30%) were isolated by BD FACSAria Fusion Flow Cytometer 9 days post transduction, further transduced with lentiviruses expressing (e)pegRNAs (approximately 50% infection), fully selected by 3 pg mL'1 puromycin and harvested 11 days after second transduction for genomic DNA extract.
High-throughput DNA sequencing of genomic DNA samples. Genomic DNA sequences containing target sites were amplified through two rounds of PCR reactions (PCR1 and 2). In PCR1, genomic regions of interest were amplified with primers containing forward and reverse adapters for Illumina sequencing (Integrated DNA Technologies). Each PCR1 reaction consisted of 1 pL genomic DNA extract, 0.1 pL of each 100 pM forward and reverse primer (0.5 pM final concentration), 10 pL Phusion U Green Multiplex PCR Master Mix
(Thermo Scientific F564L) and 8.8 pL Nuclease-Free Water (AM9939) and was performed with the following cycling conditions: 98 °C for 2 min, 28 cycles of [98 °C for 10 s, 61 °C for 20 s, and 72 °C for 30 s], followed by 72 °C for 2 min. Successful PCR1 amplification was confirmed by 1% agarose (Goldbio A-201-100) gel electrophoresis before proceeding to PCR2 to uniquely index each sample with both forward and reverse Illumina barcoding primers. Each 14 pL PCR2 reaction consisted of 1 pL unpurified PCR 1 reaction, 0.5 pM of each forward and reverse Illumina barcoding primer, 7 pL Phusion U Green Multiplex PCR Master Mix (Thermo Scientific F564L) and Nuclease-Free Water (AM9939) and was performed with the following cycling conditions: 98 °C for 2 min, 9 cycles of [98 °C for 10 s, 61 °C for 20 s, and 72 °C for 30 s], followed by 72 °C for 2 min. Successful PCR2 amplification was confirmed by 1% agarose gel electrophoresis before being pooled by common amplicons. 30 pL pooled PCR2 reactions of each common amplicon were purified by 1% agarose gel electrophoresis with a manual size selection of 200 to 600 bp according to a 100 bp DNA ladder (Goldbio D001-500), extracted with Zymoclean Gel DNA Recovery Kit (Zymo Research D4001) and eluted in 30 pL Buffer EB (Qiagen 19086). The gel purified PCR2 products were quantified by Qubit IX dsDNA High Sensitivity kit (Invitrogen Q33231) and a high sensitivity DNA chip (Agilent Technologies 5067-4626) on an Agilent 2100 Bioanalyzer and sequenced with the MiSeq Reagent Micro Kit v2 300 cycles (Illumina MS- 103 -1002) or Nano Kit v2 300 cycles (Illumina MS-103-1001) with 300 cycles for R1 read, 8 cycles i7 index read and 8 cycles i5 index read. Sequencing reads were demultiplexed through HTSEQ (Princeton University High Throughput Sequencing Database) or bcl2fastq2 (Illumina) and sequencing adapters were trimmed using Cutadapt with the parameter “-a AGATCGGAAGAGCACACGTCTGAACTCCAGTCAC” (SEQ ID NO: 61).
To quantify prime editing outcomes, amplicon sequencing reads were aligned to corresponding reference sequences with CRISPResso2 in HDR batch mode using the intended editing outcome as the expected allele (-e) with the parameters “-q 30” and “- discard indel reads TRUE”. For each amplicon, the CRISPResso2 quantification window was centered at the pegRNA nick (“-wc -3”) and the window size (“-w”) was set to 10 + the distance between nicks generated by the pegRNA and the nicking sgRNA. The same parameters were used for PE2, PE3, PE4 and PE5 conditions. The frequency of intended editing without indels was calculated as: (number of non-discarded HDR-aligned reads)/(number of reads that aligned all amplicons). The frequency of intended editing with indels was calculated as: (number of discarded HDR-aligned reads)/(number of reads that aligned all amplicons). The frequency of total intended editing (with or without indels) was calculated as (number of HDR-aligned
reads)/(number of reads that aligned all amplicons). The frequency of total indels was calculated as: (number of discarded reads)/( number of reads that aligned all amplicons). The frequency of indels without intended editing was calculated as (number of discarded reference- aligned reads)/(number of reads that aligned all amplicons). The intended prime editing efficiencies referred to frequencies of intended editing without indels and the indel efficiencies referred to frequencies of total indels in this study unless otherwise specified.
To quantify off-target prime editing, two to four most common Cas9 off-target sites experimentally determined for each on-target locus were amplified from genomic DNA extract of U2OS cells nucleofected with plasmids encoding PEmax or PE7 and pegRNAs targeting HEK3, HEK4, FANCF and EMX1 loci in FIG. 4D. Off-target editing was quantified as previous described with minor modifications. Specifically, reads were aligned to corresponding off-target reference sequences using CRISPResso2 in standard batch mode with parameters “- q 30”, “-w 10” and “-discard indel reads TRUE”. Each off-target amplicon sequence was compared to the 3' DNA flap sequence encoded by the pegRNA extension starting from the nucleotide 3 ' of Cas9 nick to the downstream until reaching the first nucleotide on the off-target amplicon that is different from the 3' DNA flap. Any reads with this nucleotide converted to that on the 3 ' DNA flap were considered off-target reads and the number of such reads can be found in the output file “Nucleotide frequency summary around sgRNA”. Off-target editing efficiencies were calculated as (number of off-target reads + number of indel-containing reads)/(number of reads that aligned all amplicons).
To quantify Cas9 cutting outcomes, CRISPResso2 was run in standard batch mode with the parameters “-q 30” and “-discard indel reads TRUE”. The intended editing efficiency referred to the frequency of indels which was calculated as (number of discarded reference- aligned reads)/ (number of reads that aligned all amplicons). Base editing outcomes were quantified by CRISPResso2 as previously described.
RT-qPCR. To quantify knockdown efficiencies of La-targeting CRISPRi sgRNAs in MCS reporter cells or La siRNA in Lenti-X 293T, total RNA was extracted using Quick-RNA Miniprep Kit (Zymo Research R1054) with DNase I treatment and 1 pg total RNA was converted to cDNA with SuperScript IV First-Strand Synthesis System (Invitrogen 18091050), according to the manufacturer’s protocol. Each 20 pL qPCR consists of 2 pL cDNA, 0.3 pM of each forward and reverse primer, 10 pL S YBR Green PCR Master Mix (Applied Biosystems 4309155) and Nuclease-Free Water (AM9939) and were performed in technical triplicate on a ViiA 7 Real-Time PCR System (Applied Biosystems) with the following cycling conditions: 50 °C for 2 min, 95 °C for 10 min, and 40 cycles of [95 °C for 15 s, 60 °C for 1 min]. Relative
La expression levels were calculated using the 2'AACT method with ACTB (a housekeeping gene) as the internal control in comparison to a non-targeting sgRNA or a non-targeting control siRNA pool.
Generation of K562 clones with PEmax knock-in at AAVS1. 91.5 pmole Alt-R S.p. Cas9 Nuclease V3 (Integrated DNA Technologies 1081058) and 150 pmole Custom Alt-R gRNA targeting AAVS1 (Integrated DNA Technologies) were complexed for 20 minutes at room temperature and were nucleofected together with 2000 ng AAVS1 PEmax knock-in plasmid as the HDR template into 7.5E5 K562 cells using the SE Cell Line 4D-Nucleofector X Kit (Lonza V4XC-1032) and program FF-120, according to the manufacturer’s protocol. 4 days after nucleofection, cells were selected with 400 pg mL’1 Geneticin (Gibco 10131027) for 2 weeks before sorted by BD FACSAria Fusion Flow Cytometer into 96-well plates at 1 cell per well with 150 pL conditioned culture medium. Single cells were grown and expanded for 2-3 weeks into clonal lines, from which the one with the highest and most homogenous EGFP expression by AttueNXT flow cytometry analysis was selected as the K562 PEmax parental cell line.
Generation of La knock-out K562 PEmax cells. 122 pmole Alt-R S .p. Cas9 Nuclease V3 (Integrated DNA Technologies 1081058) and 200 pmole Alt-R CRISPR-Cas9 sgRNA targeting La (Integrated DNA Technologies Hs.Cas9.SSB. l.AA) were complexed for 20 minutes at room temperature and were nucleofected into 5E5 K562 PEmax parental cells using the SE Cell Line 4D-Nucleofector X Kit (Lonza V4XC-1032) and program FF-120, according to the manufacturer’s protocol. 5 days post nucleofection, cells were sorted by BD FACSAria Fusion Flow Cytometer into 96-well plates at 1 cell per well with 150 pL conditioned culture medium. Single cells were grown and expanded for 2-3 weeks into clonal lines. Clones with high EGFP+ cell% according to AttueNXT flow cytometry analysis were selected for further characterization by targeted sequencing at genomic La locus and CRISPResso2 analysis.
Western blotting. Cells were washed with DPBS (Gibco 14190144), lysed in 2X western lysis buffer, boiled for 5 minutes at 95°C, and stored at -80°C prior to use. For SDS- PAGE, samples were reheated at 95°C for 5 minutes, well-mixed, loaded to the 10% gel, and run for 1.5 hours at 150 V. The Precision Plus Protein Dual Color Standards (Bio-Rad 161- 0374) was loaded as the marker. The proteins were transferred into a nitrocellulose membrane (VWR 10120-060) with the Trans-Blot SD semi-dry transfer cell (Bio-Rad). Antibodies were diluted in 5% Blotto (5% nonfat dry milk in TBST) and incubated with the membrane for 1 hour at room temperature. The following primary antibodies were used: anti-La mouse monoclonal antibody (1 :5000; Abeam ab75927); anti-GAPDH rabbit monoclonal antibody
(1 :5000; Abeam abl81602); Guide-it Cas9 rabbit Polyclonal Antibody (1 : 1000; Takara 632607). The following secondary antibodies were used: HRP-conjugated sheep anti-mouse polyclonal antibody (1 :2000; VWR 95017-332) and HRP-conjugated donkey anti-rabbit polyclonal antibody (1 :2000; VWR 95017-556). After incubating with secondary antibodies, the membrane was washed with TBST and immersed into Lumi-LightPLUS Western Blotting Substrate (Sigma 12015196001) for 3 minutes in dark prior to exposure with Azure Biosystems 600. The Restore Western Blot Stripping Buffer (Thermo Scientific 21059) was applied to strip the membrane before reprobing.
Cell growth assay. To quantify the effect of La knockout on cell growth, K562 PEmax parental cells, La-ko4 and La-ko5 cells were monitored using AttueNXT flow cytometry with three individual replicates per cell line and each replicate in a 100 mm cell culture dish (Greiner Bio-One 664160). On each day, live cell density (average of three repeat measurements) of each replicate and each cell line was quantified by flow cytometry, diluted to approximately 5E5 mL’1, and quantified again immediately and 24 hours after dilution. The relative cell doubling was calculated as the ratio of live cell density measured 24 hours after dilution to that measured immediately after dilution in log2 scale.
Small RNA sequencing. The small RNA sequencing (small RNA-seq) with targeting (e)pegRNAs was performed in triplicate and for each replicate, 5E6 K562 PEmax parental or La-ko4 cells were nucleofected with 2500 ng either one of the two (e)pegRNA plasmid sets (Set 1 and 2) using the SE Cell Line 4D-Nucleofector X Kit L (Lonza V4XC-1024) and pulse code FF120, according to the manufacturer’s protocol. Set 1 consists of plasmids encoding FANCF +5 G to T pegRNA, HEK3 +1 T to A pegRNA, DNMT1 +5 G to T pegRNA, RUNX1 +5 G to T epegRNA (evopreQi), VEGFA +5 G to T pegRNA and EMX1 +5 G to T epegRNA (mpknot). Set 2 consists of plasmids encoding RNF2 +1 C to A pegRNA, HEK3 +1 T to A epegRNA (mpknot), DNMT1 +5 G to T epegRNA (evopreQi), RUNX1 +5 G to T pegRNA, VEGFA +5 G to T pegRNA and EMX1 +5 G to T pegRNA. The VEGFA +5 G to T pegRNA plasmid was shared by both sets and served as the internal control for potential cross-set normalization. The FANCF +5 G to T pegRNA plasmid and the RNF2 +1 C to A pegRNA were specific to set 1 and 2 respectively. For HEK3, DNMT1, RUNX1, and EMX1 genomic loci, one set has the pegRNA plasmid while the other has the epegRNA plasmid encoding the same prime edit. Each set only had one evopreQi epegRNA plasmid and one mpknot epegRNA plasmid. The sets were formulated so that each (e)pegRNA transcript from cells nucleofected with one set can be aligned uniquely to the corresponding (e)pegRNA in that set, based on the
observation in preliminary experiments that few fragments were solely mapped to the sgRNA scaffold shared by different (e)pegRNAs.
The small RNA-seq with non-targeting mDNMTl +6 G to C pegRNA and epegRNA (tevopreQi) were performed in quadruplicate and for each replicate, 5E6 K562 PEmax parental or La-ko4 cells were nucleofected with 5000 ng (e)pegRNA plasmid using the SE Cell Line 4D-Nucleofector X Kit L (Lonza V4XC-1024) and pulse code FF120, according to the manufacturer’s protocol.
In both experiments, half of the cells from each nucleofection were harvested 24 or 48 hours after nucleofection and total RNA was extracted using mirVana miRNA Isolation Kit with phenol (Invitrogen AMI 560) and was quantified using a NanoDrop One UV-Vis spectrophotometer (Thermo Scientific). For each sample, a small RNA library was constructed with 1 pg total RNA as the input using NEBNext Multiplex Small RNA Library Prep Set for Illumina (Set 1) (New England Biolabs E7300S) and NEBNext Multiplex Oligos for Illumina Index Primers Set 3 (New England Biolabs E7710S) and Set 4 (New England Biolabs E7730S) according to the manufacturer’s protocol. Equivolume libraries of all samples were pooled, purified using SPRIselect (Beckman Coulter B23318) with a double size selection (0.5x right side and 1.35x left side), quantified by Qubit IX dsDNA High Sensitivity kit (Invitrogen Q33231) and a high sensitivity DNA chip (Agilent Technologies 5067-4626) on an Agilent 2100 Bioanalyzer, and was sequenced with the NovaSeq 6000 SP Reagent kit vl.5 100 cycles (Illumina 20028401) with 40 cycles for R1 read, 8 cycles i7 index read and 90 cycles for R2 read.
To validate La phenotype with non-targeting mus DNMT1 (mDNMTl) +6 G to C (e)pegRNAs, K562 PEmax parental and La-ko4 cells were transduced with lentiviruses harboring the mDNMTl target. 1E6 each transduced cells were nucleofected with 500 or 1000 ng pegRNA or epegRNA plasmid using the SE Cell Line 4D-Nucleofector X Kit (Lonza V4XC-1032) and program FF-120, according to the manufacturer’s protocol. 14 the amount of Cells from each nucleofection were harvested 1, 2, 3 and 4 days after nucleofection and the editing outcomes were quantified by high-throughput DNA sequencing and CRISPResso2 analysis.
Small RNA sequencing data analysis. Sequencing reads were demultiplexed through HTSEQ (Princeton University High Throughput Sequencing Database). The reads were trimmed, aligned, and processed using a Snakemake workflow and R scripts available at github.com/Princeton-LSI-ResearchComputing/PE-small-RNA-seq-analysis. Adapters were trimmed using Cutadapt -a AGATCGGAAGAGCACACGTCTGAACTCCAGTCAC -
AGATCGTCGGACTGTAGAACTCTGAACGTGTAGATCTCGGTGGTCGCCGTATCAT T (SEQ ID NO: 62). The trimmed reads were then aligned to the sequence(s) of (e)pegRNA(s) the sample was nucleofected with, using Bowtie2 with default alignment options. Reads that did not align to the (e)pegRNA references were then aligned to the human genome (GRCh38 primary assembly from Ensembl release 107) using Bowtie2 with default alignment parameters. Downstream analysis of the alignments used only reads mapped in a proper pair, ensuring both ends of the sequenced fragment were properly mapped.
Quantifications of human small RNA, including assigning fragments to human transcripts, genes, and biotypes as well as counting, were performed on properly paired alignments using a custom Python script available in the GitHub repository. To distinguish between overlapping annotations, each aligned fragment was assigned to the annotation that most closely matched the start and end point of the fragment. The (e)pegRNA(s) were quantified for each sample by assigning each properly aligned fragment into one of three bins defined in the main text (c/.s-active, /ra/z.s-active and inactive) using Rsamtools and plyranges. Differential expression was calculated using DESeq2 version 1.38.3 with a design consisting of two covariates: (e)pegRNA plasmid set nucleofected (set 1 or 2) and cell line (K562 PEmax or La-ko4). Default parameters were used to estimate library size factors, genewise dispersion, and fitting of the negative binomial GLM to determine log2 fold change values. Log fold change shrinkage was performed using the apeglm algorithm. The default two-sided Wald test was used to determine the p values and the Bonferroni Holm method was used for multiple test correction. Coverage plots were generated using ggplot2 on data organized using the readr, dplyr, tidyr, and stringr packages.
RNA-seq and data analysis. Each condition of RNA-seq was performed in quadruplicate and for each replicate, 1E6 K562 cells were nucleofected with 750 ng PEmax or PE7 editor plasmid and 250 ng pegRNA plasmid encoding HEK3 +1 T to A or PRNP + 6 G to T using SE Cell Line 4D-Nucleofector X Kit S (Lonza V4XC-1032) with program FF-120, according to the manufacturer’s protocols. Nucleofected cells were cultured in 6-well plates with 2.5 mL medium per well. 24, 48 and 72 hours after nucleofection, 150 pL cell culture from each replicate and condition was analyzed by AttueNXT flow cytometry to quantify cell viability and live cell density. 72 hours after nucleofection, 1 mL cell culture from each replicate and condition was harvested for genomic DNA extract to quantify prime editing outcomes at HEK3 or PRNP locus. The remaining ImL cell culture was pelleted and washed with DPBS (Gibco 14190144) for total RNA extraction using RNeasy Plus Mini Kit (Qiagen 74134) with on column DNase I treatment. Total RNA was quantified by a NanoDrop One
UV-Vis spectrophotometer (Thermo Scientific) and RNA 6000 Pico chips (Agilent Technologies 5067-1513) on an Agilent 2100 Bioanalyzer. 3' mRNA SMART-seq libraries were prepared using total RNA as input on an Apollo NGS library prep system (Takara 640078) following the manufacturer’s protocol. Sequencing libraries were pooled, quantified by Qubit IX dsDNA High Sensitivity kit (Invitrogen Q33231) and a high sensitivity DNA chip (Agilent Technologies 5067-4626) on an Agilent 2100 Bioanalyzer and sequenced with the NovaSeq 6000 SP Reagent kit vl.5 100 cycles (Illumina 20028401) with 112 cycles for R1 read, 10 cycles index read.
Sequencing reads were demultiplexed through HTSEQ (Princeton University High Throughput Sequencing Database). Alignment, quantification, and differential expression were performed using a Snakemake workflow and R scripts available on GitHub github.com/Princeton-LSI-ResearchComputing/PE-mRNA-seq-diffexp. The reads were aligned to the GRCh38 genome from Ensembl release 100 using STAR with default alignment parameters. Quantification was performed by STAR during alignment. Differential expression between editors was performed separately for each pegRNA. The standard DESeq2 procedure was performed to determine the differential expression between each editor within the set of samples for each pegRNA. Fold changes for lowly expressed genes were shrunken using the adaptive shrinkage estimator from the ashr package. Figures were generated using R packages ggplot2 and ggpubr.
T cell isolation, culture and prime editing. Human peripheral blood Leukopaks enriched for PBMCs were sourced from STEMCELL Technologies (catalog # 200-0092). No preference was given with regard to sex, gender, ethnicity or race. T cells were isolated with the Easy Sep Human T cell isolation kit (STEMCELL Technologies 100-0695) according to manufacturer's instructions. Immediately after isolation, T cells were used directly for in vitro experiments. All T cells were cultured in complete X-VIVO 15 consisting of X- VI VO 15 (Lonza Bioscience 04-418Q) supplemented with 5% FBS (R&D systems), 4mM N-acetyl- cysteine (RPI A10040) and 55 pM 2-mercaptoethanol (GIbco 21985023). Pan CD3+ T cells were thawed and activated with anti-CD3/anti-CD28 dynabeads (Gibco 40203D) at a 1 : 1 bead:cell ratio in presence of 500 IU mL’1 IL-2. Two days after stimulation, T cells were magnetically de-beaded and taken up in P3 buffer with supplement (Lonza Bioscience V4SP- 3096) at 37.5E6 cells mL'1. 1.5 pg PEmax or PE7 mRNA mixed with 50 pmole synthetic pegRNA (IDT) was added per 20pL cells, not exceeding 25 pL total volume per reaction. Cells were subsequently electroporated on a Lonza 4D Nucleofector using program DS-137. Immediately after electroporation, 80 pL warm complete X-VIVO 15 was added to each
electroporation well and cells were incubated for 30 minutes in a 5% CO2 incubator at 37 °C followed by distribution of each electroporation reaction into 3 wells of a 96 well round bottom plate. Each well was brought to 200 pL complete XVIV015 and 200 IU mL'1 IL-2. Cells were subcultured and expanded by addition of fresh media and IL-2 every 2 to 3 days. Four days after electroporation, approximately 5E5 cells were spun down at 500 g for 5 minutes, and genomic DNA was extracted using the DNeasy Blood & Tissue Kit (Qiagen 69506) per manufacturer's instructions with an elution volume of 100 pL. To assess editing efficiency, PCR was performed with 25 uL of eluted genomic DNA per sample in an 100 pL PCR reaction with KAPA HiFi HotStart ReadyMix (Roche 9420398001) with the following cycling conditions: 95 °C for 3 min, 28 cycles of [98 °C for 20 s, 63 °C for 15 s, and 72 °C for 60 s], followed by 72 °C for 2 min. PCR products were purified by SPRI selection (Beckman Coulter B23317) and 2 pL eluted product was used for 8 cycles of additional PCR with KAPA HiFi HotStart ReadyMix to add Illumina sequencing adapters and indices. The final PCR products were purified by SPRI selection, quantified with Qubit IX dsDNA High Sensitivity (HS) assay kit (Invitrogen Q33230), equimolarly pooled, and sequenced with the MiSeq Reagent Kit v2 300 cycles (Illumina MS-102-2002) with 300 cycles for R1 read, 8 cycles i7 index read and 8 cycles i5 index read. Sequencing data were demultiplexed using BaseSpace and analyzed by CRISPResso2.
HPSC isolation, culture and prime editing. mRNA in vitro transcription template plasmids for HPSC experiments were constructed by cloning PEmax and PE7 into a previously described vector. mRNA was generated using HiScribe T7 High Yield RNA Synthesis Kit (New England Biolabs E2040S) and BbsI linearized plasmids as templates with UTP fully replaced by NkMethylpseudouridine-S '-triphosphate (TriLink Biotechnologies N-1081) and co-transcriptional capping by CleanCap AG (TriLink Biotechnologies N-7113). Following IVT, mRNA was purified using the Monarch RNA Cleanup kit (500 pg) (NEB T2050S), eluted in IDTE pH 7.5 (Integrated DNA Technologies 11-05-01-15) and quantified using Qubit RNA High Sensitivity (HS) Assay Kit (Invitrogen Q32852). Synthetic pegRNAs were ordered as Custom Alt-R gRNA from Integrated DNA Technologies and resuspended at 200 pM in IDTE pH 7.5. Cryopreserved human CD34+ HSPCs from mobilized peripheral blood of deidentified healthy donors were obtained from the Fred Hutchinson Cancer Research Center (Seattle, Washington). CD34+ HSPCs were cultured with X-Vivo-15 media supplemented with 100 ng mL'1 human Stem Cell Growth Factor (SCF), 100 ng mL'1 human thrombopoietin (TPO), and 100 ng mL'1 recombinant human FMS-like Tyrosine Kinase 3 Ligand (Flt3-L). CD34+ HSPCs were thawed and cultured for 24 hours in the presence of cytokines prior to nucleofection.
2.5E5 CD34+ HSPCs were electroporated using the P3 Primary Cell X kit S (Lonza Bioscience V4SP-3096) according to manufacturer’s recommendations with 2000 ng PEmax or PE7 mRNA and 200 pmole synthetic pegRNA using pulse code DS-130. Genomic DNA was harvested 3 days post nucleofection with QuickExtract DNA Extraction Solution (LGC Biosearch Technologies QE09050) following manufacturer’s recommendations. Prime editing outcomes were quantified by high-throughput DNA sequencing and CRISPResso2 analysis as described earlier.
Unless defined otherwise, all technical and scientific terms used herein have the same meanings as commonly understood by one of skill in the art to which the disclosed invention belongs. Publications cited herein and the materials for which they are cited are specifically incorporated by reference.
Those skilled in the art will appreciate that numerous changes and modifications can be made to the preferred embodiments of the invention and that such changes and modifications can be made without departing from the spirit of the invention. It is, therefore, intended that the appended claims cover all such equivalent variations as fall within the true spirit and scope of the invention.
SEQUENCES
SEQ ID NO: 1
MKRTADGSEFESPKKKRKVDKKYSIGLDIGTNSVGWAVITDEYKVPSKKFKVLGNTDRHSIKKNLIGA
LLFDSGETAEATRLKRTARRRYTRRKNRICYLQEIFSNEMAKVDDSFFHRLEESFLVEEDKKHERHPIFG
NIVDEVAYHEKYPTIYHLRKKLVDSTDKADLRLIYLALAHMIKFRGHFLIEGDLNPDNSDVDKLFIQLV
QTYNQLFEENPINASGVDAKAILSARLSKSRRLENLIAQLPGEKKNGLFGNLIALSLGLTPNFKSNFDLA
EDAKLQLSKDTYDDDLDNLLAQIGDQYADLFLAAKNLSDAILLSDILRVNTEITKAPLSASMIKRYDEH
HQDLTLLKALVRQQLPEKYKEIFFDQSKNGYAGYIDGGASQEEFYKFIKPILEKMDGTEELLVKLNRED
LLRKQRTFDNGSIPHQIHLGELHAILRRQEDFYPFLKDNREKIEKILTFRIPYYVGPLARGNSRFAWMTR
KSEETITPWNFEEWDKGASAQSFIERMTNFDKNLPNEKVLPKHSLLYEYFTVYNELTKVKYVTEGMR
KPAFLSGEQKKAIVDLLFKTNRKVTVKQLKEDYFKKIECFDSVEISGVEDRFNASLGTYHDLLKIIKDKD
FLDNEENEDILEDIVLTLTLFEDREMIEERLKTYAHLFDDKVMKQLKRRRYTGWGRLSRKLINGIRDKQ
SGKTILDFLKSDGFANRNFMQLIHDDSLTFKEDIQKAQVSGQGDSLHEHIANLAGSPAIKKGILQTVKV
VDELVKVMGRHKPENIVIEMARENQTTQKGQKNSRERMKRIEEGIKELGSQILKEHPVENTQLQNEKL
YLYYLQNGRDMYVDQELDINRLSDYDVDAIVPQSFLKDDSIDNKVLTRSDKNRGKSDNVPSEEWKK
MKNYWRQLLNAKLITQRKFDNLTKAERGGLSELDKAGFIKRQLVETRQITKHVAQILDSRMNTKYDE
NDKLIREVKVITLKSKLVSDFRKDFQFYKVREINNYHHAHDAYLNAWGTALIKKYPKLESEFVYGDY
KVYDVRKMIAKSEQEIGKATAKYFFYSNIMNFFKTEITLANGEIRKRPLIETNGETGEIVWDKGRDFAT
VRKVLSMPQVNIVKKTEVQTGGFSKESILPKRNSDKLIARKKDWDPKKYGGFDSPTVAYSVLWAKVE
KGKSKKLKSVKELLGITIMERSSFEKNPIDFLEAKGYKEVKKDLIIKLPKYSLFELENGRKRMLASAGEL
QKGNELALPSKYVNFLYLASHYEKLKGSPEDNEQKQLFVEQHKHYLDEIIEQISEFSKRVILADANLDK
VLSAYNKHRDKPIREQAENIIHLFTLTNLGAPAAFKYFDTTIDRKRYTSTKEVLDATLIHQSITGLYETRI
DLSQLGGDSGGSSGGSSGSETPGTSESATPESSGGSSGGSSTLNIEDEYRLHETSKEPDVSLGSTWLSDFP
QAWAETGGMGLAVRQAPLIIPLKATSTPVSIKQYPMSQEARLGIKPHIQRLLDQGILVPCQSPWNTPLLP
VKKPGTNDYRPVQDLREVNKRVEDIHPTVPNPYNLLSGLPPSHQWYTVLDLKDAFFCLRLHPTSQPLF
AFEWRDPEMGISGQLTWTRLPQGFKNSPTLFNEALHRDLADFRIQHPDLILLQYVDDLLLAATSELDCQ
QGTRALLQTLGNLGYRASAKKAQICQKQVKYLGYLLKEGQRWLTEARKETVMGQPTPKTPRQLREFL
GKAGFCRLFIPGFAEMAAPLYPLTKPGTLFNWGPDQQKAYQEIKQALLTAPALGLPDLTKPFELFVDEK
QGYAKGVLTQKLGPWRRPVAYLSKKLDPVAAGWPPCLRMVAAIAVLTKDAGKLTMGQPLVILAPHA
VEALVKQPPDRWLSNARMTHYQALLLDTDRVQFGPWALNPATLLPLPEEGLQHNCLDILAEAHGTRP
DLTDQPLPDADHTWYTDGSSLLQEGQRKAGAAVTTETEVIWAKALPAGTSAQRAELIALTQALKMAE
GKKLNVYTDSRYAFATAHIHGEIYRRRGWLTSEGKEIKNKDEILALLKALFLPKRLSIIHCPGHQKGHSA
EARGNRMADQAARKAAITETPDTSTLLIENSSPSGGSSGGSSGSETPGTSESATPESSGGSSGGSMAENG
DNEKMAALEAKICHQIEYYFGDFNLPRDKFLKEQIKLDEGWVPLEIMIKFNRLNRLTTDFNVIVEALSK
SKAELMEISEDKTKIRRSPSKPLPEVTDEYKNDVKNRSVYIKGFPTDATLDDIKEWLEDKGQVLNIQMR
RTLHKAFKGSIFWFDSIESAKKFVETPGQKYKETDLLILFKDDYFAKKNESGGSKRTADGSEFEPKKK
RKV*
SEQ ID NO: 2
MKRTADGSEFESPKKKRKVAENGDNEKMAALEAKICHQIEYYFGDFNLPRDKFLKEQIKLDEGWVPLE IMIKFNRLNRLTTDFNVIVEALSKSKAELMEISEDKTKIRRSPSKPLPEVTDEYKNDVKNRSVYIKGFPTD ATLDDIKEWLEDKGQVLNIQMRRTLHKAFKGSIFWFDSIESAKKFVETPGQKYKETDLLILFKDDYFA KKNESGGSSGGSSGSETPGTSESATPESSGGSSGGSDKKYSIGLDIGTNSVGWAVITDEYKVPSKKFKVL GNTDRHSIKKNLIGALLFDSGETAEATRLKRTARRRYTRRKNRICYLQEIFSNEMAKVDDSFFHRLEESF LVEEDKKHERHPIFGNIVDEVAYHEKYPTIYHLRKKLVDSTDKADLRLIYLALAHMIKFRGHFLIEGDL
NPDNSDVDKLFIQLVQTYNQLFEENPINASGVDAKAILSARLSKSRRLENLIAQLPGEKKNGLFGNLIAL SLGLTPNFKSNFDLAEDAKLQLSKDTYDDDLDNLLAQIGDQYADLFLAAKNLSDAILLSDILRVNTEIT KAPLSASMIKRYDEHHQDLTLLKALVRQQLPEKYKEIFFDQSKNGYAGYIDGGASQEEFYKFIKPILEK MDGTEELLVKLNREDLLRKQRTFDNGSIPHQIHLGELHAILRRQEDFYPFLKDNREKIEKILTFRIPYYV GPLARGNSRFAWMTRKSEETITPWNFEEVVDKGASAQSFIERMTNFDKNLPNEKVLPKHSLLYEYFTV YNELTKVKYVTEGMRKPAFLSGEQKKAIVDLLFKTNRKVTVKQLKEDYFKKIECFDSVEISGVEDRFN
ASLGTYHDLLKIIKDKDFLDNEENEDILEDIVLTLTLFEDREMIEERLKTYAHLFDDKVMKQLKRRRYT GWGRLSRKLINGIRDKQSGKTILDFLKSDGFANRNFMQLIHDDSLTFKEDIQKAQVSGQGDSLHEHIAN LAGSPAIKKGILQTVKWDELVKVMGRHKPENIVIEMARENQTTQKGQKNSRERMKRIEEGIKELGSQI LKEHPVENTQLQNEKLYLYYLQNGRDMYVDQELDINRLSDYDVDAIVPQSFLKDDSIDNKVLTRSDKN RGKSDNVPSEEWKKMKNYWRQLLNAKLITQRKFDNLTKAERGGLSELDKAGFIKRQLVETRQITKH VAQILDSRMNTKYDENDKLIREVKVITLKSKLVSDFRKDFQFYKVREINNYHHAHDAYLNAWGTALI
KKYPKLESEFVYGDYKVYDVRKMIAKSEQEIGKATAKYFFYSNIMNFFKTEITLANGEIRKRPLIETNGE
TGEIVWDKGRDFATVRKVLSMPQVNIVKKTEVQTGGFSKESILPKRNSDKLIARKKDWDPKKYGGFDS
PTVAYSVLWAKVEKGKSKKLKSVKELLGITIMERSSFEKNPIDFLEAKGYKEVKKDLIIKLPKYSLFEL
ENGRKRMLASAGELQKGNELALPSKYVNFLYLASHYEKLKGSPEDNEQKQLFVEQHKHYLDEIIEQISE
FSKRVILADANLDKVLSAYNKHRDKPIREQAENIIHLFTLTNLGAPAAFKYFDTTIDRKRYTSTKEVLDA
TLIHQSITGLYETRIDLSQLGGDSGGSSGGSSGSETPGTSESATPESSGGSSGGSSTLNIEDEYRLHETSKE
PDVSLGSTWLSDFPQAWAETGGMGLAVRQAPLIIPLKATSTPVSIKQYPMSQEARLGIKPHIQRLLDQGI
LVPCQSPWNTPLLPVKKPGTNDYRPVQDLREVNKRVEDIHPTVPNPYNLLSGLPPSHQWYTVLDLKDA
FFCLRLHPTSQPLFAFEWRDPEMGISGQLTWTRLPQGFKNSPTLFNEALHRDLADFRIQHPDLILLQYVD
DLLLAATSELDCQQGTRALLQTLGNLGYRASAKKAQICQKQVKYLGYLLKEGQRWLTEARKETVMG
QPTPKTPRQLREFLGKAGFCRLFIPGFAEMAAPLYPLTKPGTLFNWGPDQQKAYQEIKQALLTAPALGL
PDLTKPFELFVDEKQGYAKGVLTQKLGPWRRPVAYLSKKLDPVAAGWPPCLRMVAAIAVLTKDAGK
LTMGQPLVILAPHAVEALVKQPPDRWLSNARMTHYQALLLDTDRVQFGPVVALNPATLLPLPEEGLQ
HNCLDILAEAHGTRPDLTDQPLPDADHTWYTDGSSLLQEGQRKAGAAVTTETEVIWAKALPAGTSAQ
RAELIALTQALKMAEGKKLNVYTDSRYAFATAHIHGEIYRRRGWLTSEGKEIKNKDEILALLKALFLPK
RLSHHCPGHQKGHSAEARGNRMADQAARKAAITETPDTSTLLIENSSPSGGSKRTADGSEFEPKKKRK
V*
SEQ ID NO: 3
MKRTADGSEFESPKKKRKVSGGSAENGDNEKMAALEAKICHQIEYYFGDFNLPRDKFLKEQIKLDEG
WVPLEIMIKFNRLNRLTTDFNVIVEALSKSKAELMEISEDKTKIRRSPSKPLPEVTDEYKNDVKNRSVYI
KGFPTDATLDDIKEWLEDKGQVLNIQMRRTLHKAFKGSIFWFDSIESAKKFVETPGQKYKETDLLILF
KDDYFAKKNESGGSSGGSSGSETPGTSESATPESSGGSSGGSDKKYSIGLDIGTNSVGWAVITDEYKVPS
KKFKVLGNTDRHSIKKNLIGALLFDSGETAEATRLKRTARRRYTRRKNRICYLQEIFSNEMAKVDDSFF
HRLEESFLVEEDKKHERHPIFGNIVDEVAYHEKYPTIYHLRKKLVDSTDKADLRLIYLALAHMIKFRGH
FLIEGDLNPDNSDVDKLFIQLVQTYNQLFEENPINASGVDAKAILSARLSKSRRLENLIAQLPGEKKNGL
FGNLIALSLGLTPNFKSNFDLAEDAKLQLSKDTYDDDLDNLLAQIGDQYADLFLAAKNLSDAILLSDIL
RVNTEITKAPLSASMIKRYDEHHQDLTLLKALVRQQLPEKYKEIFFDQSKNGYAGYIDGGASQEEFYKF
IKPILEKMDGTEELLVKLNREDLLRKQRTFDNGSIPHQIHLGELHAILRRQEDFYPFLKDNREKIEKILTF
RIPYYVGPLARGNSRFAWMTRKSEETITPWNFEEWDKGASAQSFIERMTNFDKNLPNEKVLPKHSLL
YEYFTVYNELTKVKYVTEGMRKPAFLSGEQKKAIVDLLFKTNRKVTVKQLKEDYFKKIECFDSVEISG
VEDRFNASLGTYHDLLKIIKDKDFLDNEENEDILEDIVLTLTLFEDREMIEERLKTYAHLFDDKVMKQL
KRRRYTGWGRLSRKLINGIRDKQSGKTILDFLKSDGFANRNFMQLIHDDSLTFKEDIQKAQVSGQGDSL
HEHIANLAGSPAIKKGILQTVKWDELVKVMGRHKPENIVIEMARENQTTQKGQKNSRERMKRIEEGIK
ELGSQILKEHPVENTQLQNEKLYLYYLQNGRDMYVDQELDINRLSDYDVDAIVPQSFLKDDSIDNKVL
TRSDKNRGKSDNVPSEEWKKMKNYWRQLLNAKLITQRKFDNLTKAERGGLSELDKAGFIKRQLVET
RQITKHVAQILDSRMNTKYDENDKLIREVKVITLKSKLVSDFRKDFQFYKVREINNYHHAHDAYLNAV
VGTALIKKYPKLESEFVYGDYKVYDVRKMIAKSEQEIGKATAKYFFYSNIMNFFKTEITLANGEIRKRP
LIETNGETGEIVWDKGRDFATVRKVLSMPQVNIVKKTEVQTGGFSKESILPKRNSDKLIARKKDWDPK
KYGGFDSPTVAYSVLVVAKVEKGKSKKLKSVKELLGITIMERSSFEKNPIDFLEAKGYKEVKKDLIIKLP
KYSLFELENGRKRMLASAGELQKGNELALPSKYVNFLYLASHYEKLKGSPEDNEQKQLFVEQHKHYL
DEIIEQISEFSKRVILADANLDKVLSAYNKHRDKPIREQAENIIHLFTLTNLGAPAAFKYFDTTIDRKRYT
STKEVLDATLIHQSITGLYETRIDLSQLGGDSGGSSGGSSGSETPGTSESATPESSGGSSGGSSTLNIEDEY
RLHETSKEPDVSLGSTWLSDFPQAWAETGGMGLAVRQAPLIIPLKATSTPVSIKQYPMSQEARLGIKPHI
QRLLDQGILVPCQSPWNTPLLPVKKPGTNDYRPVQDLREVNKRVEDIHPTVPNPYNLLSGLPPSHQWY
TVLDLKDAFFCLRLHPTSQPLFAFEWRDPEMGISGQLTWTRLPQGFKNSPTLFNEALHRDLADFRIQHP
DLILLQYVDDLLLAATSELDCQQGTRALLQTLGNLGYRASAKKAQICQKQVKYLGYLLKEGQRWLTE
ARKETVMGQPTPKTPRQLREFLGKAGFCRLFIPGFAEMAAPLYPLTKPGTLFNWGPDQQKAYQEIKQA
LLTAPALGLPDLTKPFELFVDEKQGYAKGVLTQKLGPWRRPVAYLSKKLDPVAAGWPPCLRMVAAIA
VLTKDAGKLTMGQPLVILAPHAVEALVKQPPDRWLSNARMTHYQALLLDTDRVQFGPWALNPATLL
PLPEEGLQHNCLDILAEAHGTRPDLTDQPLPDADHTWYTDGSSLLQEGQRKAGAAVTTETEVIWAKAL
PAGTSAQRAELIALTQALKMAEGKKLNVYTDSRYAFATAHIHGEIYRRRGWLTSEGKEIKNKDEILALL
KALFLPKRLSIIHCPGHQKGHSAEARGNRMADQAARKAAITETPDTSTLLIENSSPSGGSKRTADGSEFE PKKKRKV*
SEQ ID NO: 4
MKRTADGSEFESPKKKRKVDKKYSIGLDIGTNSVGWAVITDEYKVPSKKFKVLGNTDRHSIKKNLIGA
LLFDSGETAEATRLKRTARRRYTRRKNRICYLQEIFSNEMAKVDDSFFHRLEESFLVEEDKKHERHPIFG
NIVDEVAYHEKYPTIYHLRKKLVDSTDKADLRLIYLALAHMIKFRGHFLIEGDLNPDNSDVDKLFIQLV QTYNQLFEENPINASGVDAKAILSARLSKSRRLENLIAQLPGEKKNGLFGNLIALSLGLTPNFKSNFDLA EDAKLQLSKDTYDDDLDNLLAQIGDQYADLFLAAKNLSDAILLSDILRVNTEITKAPLSASMIKRYDEH
HQDLTLLKALVRQQLPEKYKEIFFDQSKNGYAGYIDGGASQEEFYKFIKPILEKMDGTEELLVKLNRED LLRKQRTFDNGSIPHQIHLGELHAILRRQEDFYPFLKDNREKIEKILTFRIPYYVGPLARGNSRFAWMTR KSEETITPWNFEEWDKGASAQSFIERMTNFDKNLPNEKVLPKHSLLYEYFTVYNELTKVKYVTEGMR KPAFLSGEQKKAIVDLLFKTNRKVTVKQLKEDYFKKIECFDSVEISGVEDRFNASLGTYHDLLKIIKDKD FLDNEENEDILEDIVLTLTLFEDREMIEERLKTYAHLFDDKVMKQLKRRRYTGWGRLSRKLINGIRDKQ SGKTILDFLKSDGFANRNFMQLIHDDSLTFKEDIQKAQVSGQGDSLHEHIANLAGSPAIKKGILQTVKV VDELVKVMGRHKPENIVIEMARENQTTQKGQKNSRERMKRIEEGIKELGSQILKEHPVENTQLQNEKL YLYYLQNGRDMYVDQELDINRLSDYDVDAIVPQSFLKDDSIDNKVLTRSDKNRGKSDNVPSEEWKK MKNYWRQLLNAKLITQRKFDNLTKAERGGLSELDKAGFIKRQLVETRQITKHVAQILDSRMNTKYDE NDKLIREVKVITLKSKLVSDFRKDFQFYKVREINNYHHAHDAYLNAWGTALIKKYPKLESEFVYGDY KVYDVRKMIAKSEQEIGKATAKYFFYSNIMNFFKTEITLANGEIRKRPLIETNGETGEIVWDKGRDFAT VRKVLSMPQVNIVKKTEVQTGGFSKESILPKRNSDKLIARKKDWDPKKYGGFDSPTVAYSVLWAKVE KGKSKKLKSVKELLGITIMERSSFEKNPIDFLEAKGYKEVKKDLIIKLPKYSLFELENGRKRMLASAGEL QKGNELALPSKYVNFLYLASHYEKLKGSPEDNEQKQLFVEQHKHYLDEIIEQISEFSKRVILADANLDK VLSAYNKHRDKPIREQAENIIHLFTLTNLGAPAAFKYFDTTIDRKRYTSTKEVLDATLIHQSITGLYETRI DLSQLGGDSGGSSGGSSGSETPGTSESATPESSGGSSGGSMAENGDNEKMAALEAKICHQIEYYFGDFN LPRDKFLKEQIKLDEGWVPLEIMIKFNRLNRLTTDFNVIVEALSKSKAELMEISEDKTKIRRSPSKPLPEV TDEYKNDVKNRSVYIKGFPTDATLDDIKEWLEDKGQVLNIQMRRTLHKAFKGSIFWFDSIESAKKFVE TPGQKYKETDLLILFKDDYFAKKNESGGSSGGSSGSETPGTSESATPESSGGSSGGSSTLNIEDEYRLHET SKEPDVSLGSTWLSDFPQAWAETGGMGLAVRQAPLIIPLKATSTPVSIKQYPMSQEARLGIKPHIQRLLD QGILVPCQSPWNTPLLPVKKPGTNDYRPVQDLREVNKRVEDIHPTVPNPYNLLSGLPPSHQWYTVLDL
KDAFFCLRLHPTSQPLFAFEWRDPEMGISGQLTWTRLPQGFKNSPTLFNEALHRDLADFRIQHPDLILLQ YVDDLLLAATSELDCQQGTRALLQTLGNLGYRASAKKAQICQKQVKYLGYLLKEGQRWLTEARKET VMGQPTPKTPRQLREFLGKAGFCRLFIPGFAEMAAPLYPLTKPGTLFNWGPDQQKAYQEIKQ ALLTAP ALGLPDLTKPFELFVDEKQGYAKGVLTQKLGPWRRPVAYLSKKLDPVAAGWPPCLRMVAAIAVLTKD AGKLTMGQPLVILAPHAVEALVKQPPDRWLSNARMTHYQALLLDTDRVQFGPWALNPATLLPLPEE GLQHNCLDILAEAHGTRPDLTDQPLPDADHTWYTDGSSLLQEGQRKAGAAVTTETEVIWAKALPAGT SAQRAELIALTQALKMAEGKKLNVYTDSRYAFATAHIHGEIYRRRGWLTSEGKEIKNKDEILALLKALF LPKRLSIIHCPGHQKGHSAEARGNRMADQAARKAAITETPDTSTLLIENSSPSGGSKRTADGSEFEPKKK RKV*
SEQ ID NO: 5
MKRTADGSEFESPKKKRKVDKKYSIGLDIGTNSVGWAVITDEYKVPSKKFKVLGNTDRHSIKKNLIGA LLFDSGETAEATRLKRTARRRYTRRKNRICYLQEIFSNEMAKVDDSFFHRLEESFLVEEDKKHERHPIFG NIVDEVAYHEKYPTIYHLRKKLVDSTDKADLRLIYLALAHMIKFRGHFLIEGDLNPDNSDVDKLFIQLV QTYNQLFEENPINASGVDAKAILSARLSKSRKLENLIAQLPGEKKNGLFGNLIALSLGLTPNFKSNFDLA EDAKLQLSKDTYDDDLDNLLAQIGDQYADLFLAAKNLSDAILLSDILRVNTEITKAPLSASMIKRYDEH HQDLTLLKALVRQQLPEKYKEIFFDQSKNGYAGYIDGGASQEEFYKFIKPILEKMDGTEELLVKLKRED LLRKQRTFDNGSIPHQIHLGELHAILRRQEDFYPFLKDNREKIEKILTFRIPYYVGPLARGNSRFAWMTR KSEETITPWNFEEWDKGASAQSFIERMTNFDKNLPNEKVLPKHSLLYEYFTVYNELTKVKYVTEGMR KPAFLSGEQKKAIVDLLFKTNRKVTVKQLKEDYFKKIECFDSVEISGVEDRFNASLGTYHDLLKIIKDKD FLDNEENEDILEDIVLTLTLFEDREMIEERLKTYAHLFDDKVMKQLKRRRYTGWGRLSRKLINGIRDKQ SGKTILDFLKSDGFANRNFMQLIHDDSLTFKEDIQKAQVSGQGDSLHEHIANLAGSPAIKKGILQTVKV VDELVKVMGRHKPENIVIEMARENQTTQKGQKNSRERMKRIEEGIKELGSQILKEHPVENTQLQNEKL YLYYLQNGRDMYVDQELDINRLSDYDVDAIVPQSFLKDDSIDNKVLTRSDKNRGKSDNVPSEEWKK MKNYWRQLLNAKLITQRKFDNLTKAERGGLSELDKAGFIKRQLVETRQITKHVAQILDSRMNTKYDE NDKLIREVKVITLKSKLVSDFRKDFQFYKVREINNYHHAHDAYLNAWGTALIKKYPKLESEFVYGDY KVYDVRKMIAKSEQEIGKATAKYFFYSNIMNFFKTEITLANGEIRKRPLIETNGETGEIVWDKGRDFAT
VRKVLSMPQVNIVKKTEVQTGGFSKESILPKRNSDKLIARKKDWDPKKYGGFDSPTVAYSVLWAKVE KGKSKKLKSVKELLGITIMERSSFEKNPIDFLEAKGYKEVKKDLIIKLPKYSLFELENGRKRMLASAGEL QKGNELALPSKYVNFLYLASHYEKLKGSPEDNEQKQLFVEQHKHYLDEIIEQISEFSKRVILADANLDK VLSAYNKHRDKPIREQAENIIHLFTLTNLGAPAAFKYFDTTIDRKRYTSTKEVLDATLIHQSITGLYETRI DLSQLGGDSGGSSGGSKRTADGSEFESPKKKRKVSGGSSGGSTLNIEDEYRLHETSKEPDVSLGSTWLS DFPQAWAETGGMGLAVRQAPLIIPLKATSTPVSIKQYPMSQEARLGIKPHIQRLLDQGILVPCQSPWNTP LLPVKKPGTNDYRPVQDLREVNKRVEDIHPTVPNPYNLLSGLPPSHQWYTVLDLKDAFFCLRLHPTSQP LFAFEWRDPEMGISGQLTWTRLPQGFKNSPTLFNEALHRDLADFRIQHPDLILLQYVDDLLLAATSELD CQQGTRALLQTLGNLGYRASAKKAQICQKQVKYLGYLLKEGQRWLTEARKETVMGQPTPKTPRQLR EFLGKAGFCRLFIPGFAEMAAPLYPLTKPGTLFNWGPDQQKAYQEIKQALLTAPALGLPDLTKPFELFV DEKQGYAKGVLTQKLGPWRRPVAYLSKKLDPVAAGWPPCLRMVAAIAVLTKDAGKLTMGQPLVILA PHAVEALVKQPPDRWLSNARMTHYQALLLDTDRVQFGPWALNPATLLPLPEEGLQHNCLDILAEAH
GTRPDLTDQPLPDADHTWYTDGSSLLQEGQRKAGAAVTTETEVIWAKALPAGTSAQRAELIALTQALK MAEGKKLNVYTDSRYAFATAHIHGEIYRRRGWLTSEGKEIKNKDEILALLKALFLPKRLSIIHCPGHQK GHSAEARGNRMADQAARKAAITETPDTSTLLIENSSPSGGSSGGSSGSETPGTSESATPESSGGSSGGSM AENGDNEKMAALEAKICHQIEYYFGDFNLPRDKFLKEQIKLDEGWVPLEIMIKFNRLNRLTTDFNVIVE ALSKSKAELMEISEDKTKIRRSPSKPLPEVTDEYKNDVKNRSVYIKGFPTDATLDDIKEWLEDKGQVLNI
QMRRTLHKAFKGSIFWFDSIESAKKFVETPGQKYKETDLLILFKDDYFAKKNESGGSKRTADGSEFES PKKKRKVGSGPAAKRVKLD*
SEQ ID NO: 6
MKRTADGSEFESPKKKRKVSGGSAENGDNEKMAALEAKICHQIEYYFGDFNLPRDKFLKEQIKLDEG WVPLEIMIKFNRLNRLTTDFNVIVEALSKSKAELMEISEDKTKIRRSPSKPLPEVTDEYKNDVKNRSVYI KGFPTDATLDDIKEWLEDKGQVLNIQMRRTLHKAFKGSIFWFDSIESAKKFVETPGQKYKETDLLILF KDDYFAKKNESGGSSGGSSGSETPGTSESATPESSGGSSGGSDKKYSIGLDIGTNSVGWAVITDEYKVPS KKFKVLGNTDRHSIKKNLIGALLFDSGETAEATRLKRTARRRYTRRKNRICYLQEIFSNEMAKVDDSFF
HRLEESFLVEEDKKHERHPIFGNIVDEVAYHEKYPTIYHLRKKLVDSTDKADLRLIYLALAHMIKFRGH FLIEGDLNPDNSDVDKLFIQLVQTYNQLFEENPINASGVDAKAILSARLSKSRKLENLIAQLPGEKKNGL FGNLIALSLGLTPNFKSNFDLAEDAKLQLSKDTYDDDLDNLLAQIGDQYADLFLAAKNLSDAILLSDIL RVNTEITKAPLSASMIKRYDEHHQDLTLLKALVRQQLPEKYKEIFFDQSKNGYAGYIDGGASQEEFYKF IKPILEKMDGTEELLVKLKREDLLRKQRTFDNGSIPHQIHLGELHAILRRQEDFYPFLKDNREKIEKILTF
RIPYYVGPLARGNSRFAWMTRKSEETITPWNFEEWDKGASAQSFIERMTNFDKNLPNEKVLPKHSLL YEYFTVYNELTKVKYVTEGMRKPAFLSGEQKKAIVDLLFKTNRKVTVKQLKEDYFKKIECFDSVEISG VEDRFNASLGTYHDLLKIIKDKDFLDNEENEDILEDIVLTLTLFEDREMIEERLKTYAHLFDDKVMKQL KRRRYTGWGRLSRKLINGIRDKQSGKTILDFLKSDGFANRNFMQLIHDDSLTFKEDIQKAQVSGQGDSL HEHIANLAGSPAIKKGILQTVKWDELVKVMGRHKPENIVIEMARENQTTQKGQKNSRERMKRIEEGIK
ELGSQILKEHPVENTQLQNEKLYLYYLQNGRDMYVDQELDINRLSDYDVDAIVPQSFLKDDSIDNKVL TRSDKNRGKSDNVPSEEWKKMKNYWRQLLNAKLITQRKFDNLTKAERGGLSELDKAGFIKRQLVET RQITKHVAQILDSRMNTKYDENDKLIREVKVITLKSKLVSDFRKDFQFYKVREINNYHHAHDAYLNAV VGTALIKKYPKLESEFVYGDYKVYDVRKMIAKSEQEIGKATAKYFFYSNIMNFFKTEITLANGEIRKRP LIETNGETGEIVWDKGRDFATVRKVLSMPQVNIVKKTEVQTGGFSKESILPKRNSDKLIARKKDWDPK
KYGGFDSPTVAYSVLVVAKVEKGKSKKLKSVKELLGITIMERSSFEKNPIDFLEAKGYKEVKKDLIIKLP KYSLFELENGRKRMLASAGELQKGNELALPSKYVNFLYLASHYEKLKGSPEDNEQKQLFVEQHKHYL DEIIEQISEFSKRVILADANLDKVLSAYNKHRDKPIREQAENIIHLFTLTNLGAPAAFKYFDTTIDRKRYT STKEVLDATLIHQSITGLYETRIDLSQLGGDSGGSSGGSKRTADGSEFESPKKKRKVSGGSSGGSTLNIE DEYRLHETSKEPDVSLGSTWLSDFPQAWAETGGMGLAVRQAPLIIPLKATSTPVSIKQYPMSQEARLGI
KPHIQRLLDQGILVPCQSPWNTPLLPVKKPGTNDYRPVQDLREVNKRVEDIHPTVPNPYNLLSGLPPSH QWYTVLDLKDAFFCLRLHPTSQPLFAFEWRDPEMGISGQLTWTRLPQGFKNSPTLFNEALHRDLADFRI QHPDLILLQYVDDLLLAATSELDCQQGTRALLQTLGNLGYRASAKKAQICQKQVKYLGYLLKEGQRW LTEARKETVMGQPTPKTPRQLREFLGKAGFCRLFIPGFAEMAAPLYPLTKPGTLFNWGPDQQKAYQEI KQALLTAPALGLPDLTKPFELFVDEKQGYAKGVLTQKLGPWRRPVAYLSKKLDPVAAGWPPCLRMV
AAIAVLTKDAGKLTMGQPLVILAPHAVEALVKQPPDRWLSNARMTHYQALLLDTDRVQFGPWALNP ATLLPLPEEGLQHNCLDILAEAHGTRPDLTDQPLPDADHTWYTDGSSLLQEGQRKAGAAVTTETEVIW AKALPAGTSAQRAELIALTQALKMAEGKKLNVYTDSRYAFATAHIHGEIYRRRGWLTSEGKEIKNKDE ILALLKALFLPKRLSIIHCPGHQKGHSAEARGNRMADQAARKAAITETPDTSTLLIENSSPSGGSKRTAD GSEFESPKKKRKVGSGPAAKRVKLD*
SEQ ID NO: 7
MKRTADGSEFESPKKKRKVDKKYSIGLDIGTNSVGWAVITDEYKVPSKKFKVLGNTDRHSIKKNLIGA LLFDSGETAEATRLKRTARRRYTRRKNRICYLQEIFSNEMAKVDDSFFHRLEESFLVEEDKKHERHPIFG NIVDEVAYHEKYPTIYHLRKKLVDSTDKADLRLIYLALAHMIKFRGHFLIEGDLNPDNSDVDKLFIQLV QTYNQLFEENPINASGVDAKAILSARLSKSRKLENLIAQLPGEKKNGLFGNLIALSLGLTPNFKSNFDLA EDAKLQLSKDTYDDDLDNLLAQIGDQYADLFLAAKNLSDAILLSDILRVNTEITKAPLSASMIKRYDEH
HQDLTLLKALVRQQLPEKYKEIFFDQSKNGYAGYIDGGASQEEFYKFIKPILEKMDGTEELLVKLKRED LLRKQRTFDNGSIPHQIHLGELHAILRRQEDFYPFLKDNREKIEKILTFRIPYYVGPLARGNSRFAWMTR KSEETITPWNFEEWDKGASAQSFIERMTNFDKNLPNEKVLPKHSLLYEYFTVYNELTKVKYVTEGMR KPAFLSGEQKKAIVDLLFKTNRKVTVKQLKEDYFKKIECFDSVEISGVEDRFNASLGTYHDLLKIIKDKD FLDNEENEDILEDIVLTLTLFEDREMIEERLKTYAHLFDDKVMKQLKRRRYTGWGRLSRKLINGIRDKQ
SGKTILDFLKSDGFANRNFMQLIHDDSLTFKEDIQKAQVSGQGDSLHEHIANLAGSPAIKKGILQTVKV VDELVKVMGRHKPENIVIEMARENQTTQKGQKNSRERMKRIEEGIKELGSQILKEHPVENTQLQNEKL YLYYLQNGRDMYVDQELDINRLSDYDVDAIVPQSFLKDDSIDNKVLTRSDKNRGKSDNVPSEEWKK MKNYWRQLLNAKLITQRKFDNLTKAERGGLSELDKAGFIKRQLVETRQITKHVAQILDSRMNTKYDE
NDKLIREVKVITLKSKLVSDFRKDFQFYKVREINNYHHAHDAYLNAVVGTALIKKYPKLESEFVYGDY KVYDVRKMIAKSEQEIGKATAKYFFYSNIMNFFKTEITLANGEIRKRPLIETNGETGEIVWDKGRDFAT VRKVLSMPQVNIVKKTEVQTGGFSKESILPKRNSDKLIARKKDWDPKKYGGFDSPTVAYSVLWAKVE KGKSKKLKSVKELLGITIMERSSFEKNPIDFLEAKGYKEVKKDLIIKLPKYSLFELENGRKRMLASAGEL QKGNELALPSKYVNFLYLASHYEKLKGSPEDNEQKQLFVEQHKHYLDEIIEQISEFSKRVILADANLDK VLSAYNKHRDKPIREQAENIIHLFTLTNLGAPAAFKYFDTTIDRKRYTSTKEVLDATLIHQSITGLYETRI DLSQLGGDSGGSSGGSSGSETPGTSESATPESSGGSSGGSMAENGDNEKMAALEAKICHQIEYYFGDFN LPRDKFLKEQIKLDEGWVPLEIMIKFNRLNRLTTDFNVIVEALSKSKAELMEISEDKTKIRRSPSKPLPEV TDEYKNDVKNRSVYIKGFPTDATLDDIKEWLEDKGQVLNIQMRRTLHKAFKGSIFWFDSIESAKKFVE TPGQKYKETDLLILFKDDYFAKKNESGGSSGGSKRTADGSEFESPKKKRKVSGGSSGGSTLNIEDEYRL HETSKEPDVSLGSTWLSDFPQAWAETGGMGLAVRQAPLIIPLKATSTPVSIKQYPMSQEARLGIKPHIQR LLDQGILVPCQSPWNTPLLPVKKPGTNDYRPVQDLREVNKRVEDIHPTVPNPYNLLSGLPPSHQWYTV LDLKDAFFCLRLHPTSQPLFAFEWRDPEMGISGQLTWTRLPQGFKNSPTLFNEALHRDLADFRIQHPDLI LLQYVDDLLLAATSELDCQQGTRALLQTLGNLGYRASAKKAQICQKQVKYLGYLLKEGQRWLTEAR KETVMGQPTPKTPRQLREFLGKAGFCRLFIPGFAEMAAPLYPLTKPGTLFNWGPDQQKAYQEIKQALL TAPALGLPDLTKPFELFVDEKQGYAKGVLTQKLGPWRRPVAYLSKKLDPVAAGWPPCLRMVAAIAVL TKDAGKLTMGQPLVILAPHAVEALVKQPPDRWLSNARMTHYQALLLDTDRVQFGPWALNPATLLPL PEEGLQHNCLDILAEAHGTRPDLTDQPLPDADHTWYTDGSSLLQEGQRKAGAAVTTETEVIWAKALPA GTSAQRAELIALTQALKMAEGKKLNVYTDSRYAFATAHIHGEIYRRRGWLTSEGKEIKNKDEILALLK ALFLPKRLSIIHCPGHQKGHSAEARGNRMADQAARKAAITETPDTSTLLIENSSPSGGSKRTADGSEFES PKKKRKVGSGPAAKRVKLD*
SEQ ID NO: 8
MKRTADGSEFESPKKKRKVDKKYSIGLDIGTNSVGWAVITDEYKVPSKKFKVLGNTDRHSIKKNLIGA LLFDSGETAEATRLKRTARRRYTRRKNRICYLQEIFSNEMAKVDDSFFHRLEESFLVEEDKKHERHPIFG NIVDEVAYHEKYPTIYHLRKKLVDSTDKADLRLIYLALAHMIKFRGHFLIEGDLNPDNSDVDKLFIQLV QTYNQLFEENPINASGVDAKAILSARLSKSRKLENLIAQLPGEKKNGLFGNLIALSLGLTPNFKSNFDLA EDAKLQLSKDTYDDDLDNLLAQIGDQYADLFLAAKNLSDAILLSDILRVNTEITKAPLSASMIKRYDEH HQDLTLLKALVRQQLPEKYKEIFFDQSKNGYAGYIDGGASQEEFYKFIKPILEKMDGTEELLVKLKRED LLRKQRTFDNGSIPHQIHLGELHAILRRQEDFYPFLKDNREKIEKILTFRIPYYVGPLARGNSRFAWMTR KSEETITPWNFEEWDKGASAQSFIERMTNFDKNLPNEKVLPKHSLLYEYFTVYNELTKVKYVTEGMR KPAFLSGEQKKAIVDLLFKTNRKVTVKQLKEDYFKKIECFDSVEISGVEDRFNASLGTYHDLLKIIKDKD FLDNEENEDILEDIVLTLTLFEDREMIEERLKTYAHLFDDKVMKQLKRRRYTGWGRLSRKLINGIRDKQ SGKTILDFLKSDGFANRNFMQLIHDDSLTFKEDIQKAQVSGQGDSLHEHIANLAGSPAIKKGILQTVKV VDELVKVMGRHKPENIVIEMARENQTTQKGQKNSRERMKRIEEGIKELGSQILKEHPVENTQLQNEKL YLYYLQNGRDMYVDQELDINRLSDYDVDAIVPQSFLKDDSIDNKVLTRSDKNRGKSDNVPSEEWKK MKNYWRQLLNAKLITQRKFDNLTKAERGGLSELDKAGFIKRQLVETRQITKHVAQILDSRMNTKYDE NDKLIREVKVITLKSKLVSDFRKDFQFYKVREINNYHHAHDAYLNAWGTALIKKYPKLESEFVYGDY
KVYDVRKMIAKSEQEIGKATAKYFFYSNIMNFFKTEITLANGEIRKRPLIETNGETGEIVWDKGRDFAT VRKVLSMPQVNIVKKTEVQTGGFSKESILPKRNSDKLIARKKDWDPKKYGGFDSPTVAYSVLWAKVE KGKSKKLKSVKELLGITIMERSSFEKNPIDFLEAKGYKEVKKDLIIKLPKYSLFELENGRKRMLASAGEL QKGNELALPSKYVNFLYLASHYEKLKGSPEDNEQKQLFVEQHKHYLDEIIEQISEFSKRVILADANLDK VLSAYNKHRDKPIREQAENIIHLFTLTNLGAPAAFKYFDTTIDRKRYTSTKEVLDATLIHQSITGLYETRI DLSQLGGDSGGSSGGSKRTADGSEFESPKKKRKVSGGSSGGSMAENGDNEKMAALEAKICHQIEYYFG DFNLPRDKFLKEQIKLDEGWVPLEIMIKFNRLNRLTTDFNVIVEALSKSKAELMEISEDKTKIRRSPSKPL PEVTDEYKNDVKNRSVYIKGFPTDATLDDIKEWLEDKGQVLNIQMRRTLHKAFKGSIFWFDSIESAKK FVETPGQKYKETDLLILFKDDYFAKKNESGGSSGGSSGSETPGTSESATPESSGGSSGGSTLNIEDEYRL HETSKEPDVSLGSTWLSDFPQAWAETGGMGLAVRQAPLIIPLKATSTPVSIKQYPMSQEARLGIKPHIQR LLDQGILVPCQSPWNTPLLPVKKPGTNDYRPVQDLREVNKRVEDIHPTVPNPYNLLSGLPPSHQWYTV LDLKDAFFCLRLHPTSQPLFAFEWRDPEMGISGQLTWTRLPQGFKNSPTLFNEALHRDLADFRIQHPDLI LLQYVDDLLLAATSELDCQQGTRALLQTLGNLGYRASAKKAQICQKQVKYLGYLLKEGQRWLTEAR KETVMGQPTPKTPRQLREFLGKAGFCRLFIPGFAEMAAPLYPLTKPGTLFNWGPDQQKAYQEIKQALL TAPALGLPDLTKPFELFVDEKQGYAKGVLTQKLGPWRRPVAYLSKKLDPVAAGWPPCLRMVAAIAVL TKDAGKLTMGQPLVILAPHAVEALVKQPPDRWLSNARMTHYQALLLDTDRVQFGPWALNPATLLPL PEEGLQHNCLDILAEAHGTRPDLTDQPLPDADHTWYTDGSSLLQEGQRKAGAAVTTETEVIWAKALPA GTSAQRAELIALTQALKMAEGKKLNVYTDSRYAFATAHIHGEIYRRRGWLTSEGKEIKNKDEILALLK ALFLPKRLSIIHCPGHQKGHSAEARGNRMADQAARKAAITETPDTSTLLIENSSPSGGSKRTADGSEFES PKKKRKVGSGPAAKRVKLD*
SEQ ID NO: 9
MKRTADGSEFESPKKKRKVDKKYSIGLDIGTNSVGWAVITDEYKVPSKKFKVLGNTDRHSIKKNLIGA LLFDSGETAEATRLKRTARRRYTRRKNRICYLQEIFSNEMAKVDDSFFHRLEESFLVEEDKKHERHPIFG NIVDEVAYHEKYPTIYHLRKKLVDSTDKADLRLIYLALAHMIKFRGHFLIEGDLNPDNSDVDKLFIQLV QTYNQLFEENPINASGVDAKAILSARLSKSRKLENLIAQLPGEKKNGLFGNLIALSLGLTPNFKSNFDLA EDAKLQLSKDTYDDDLDNLLAQIGDQYADLFLAAKNLSDAILLSDILRVNTEITKAPLSASMIKRYDEH HQDLTLLKALVRQQLPEKYKEIFFDQSKNGYAGYIDGGASQEEFYKFIKPILEKMDGTEELLVKLKRED LLRKQRTFDNGSIPHQIHLGELHAILRRQEDFYPFLKDNREKIEKILTFRIPYYVGPLARGNSRFAWMTR KSEETITPWNFEEWDKGASAQSFIERMTNFDKNLPNEKVLPKHSLLYEYFTVYNELTKVKYVTEGMR
KPAFLSGEQKKAIVDLLFKTNRKVTVKQLKEDYFKKIECFDSVEISGVEDRFNASLGTYHDLLKIIKDKD FLDNEENEDILEDIVLTLTLFEDREMIEERLKTYAHLFDDKVMKQLKRRRYTGWGRLSRKLINGIRDKQ SGKTILDFLKSDGFANRNFMQLIHDDSLTFKEDIQKAQVSGQGDSLHEHIANLAGSPAIKKGILQTVKV VDELVKVMGRHKPENIVIEMARENQTTQKGQKNSRERMKRIEEGIKELGSQILKEHPVENTQLQNEKL YLYYLQNGRDMYVDQELDINRLSDYDVDAIVPQSFLKDDSIDNKVLTRSDKNRGKSDNVPSEEWKK MKNYWRQLLNAKLITQRKFDNLTKAERGGLSELDKAGFIKRQLVETRQITKHVAQILDSRMNTKYDE NDKLIREVKVITLKSKLVSDFRKDFQFYKVREINNYHHAHDAYLNAWGTALIKKYPKLESEFVYGDY KVYDVRKMIAKSEQEIGKATAKYFFYSNIMNFFKTEITLANGEIRKRPLIETNGETGEIVWDKGRDFAT
VRKVLSMPQVNIVKKTEVQTGGFSKESILPKRNSDKLIARKKDWDPKKYGGFDSPTVAYSVLWAKVE KGKSKKLKSVKELLGITIMERSSFEKNPIDFLEAKGYKEVKKDLIIKLPKYSLFELENGRKRMLASAGEL QKGNELALPSKYVNFLYLASHYEKLKGSPEDNEQKQLFVEQHKHYLDEIIEQISEFSKRVILADANLDK VLSAYNKHRDKPIREQAENIIHLFTLTNLGAPAAFKYFDTTIDRKRYTSTKEVLDATLIHQSITGLYETRI DLSQLGGDSGGSSGGSKRTADGSEFESPKKKRKVSGGSSGGSTLNIEDEYRLHETSKEPDVSLGSTWLS DFPQAWAETGGMGLAVRQAPLIIPLKATSTPVSIKQYPMSQEARLGIKPHIQRLLDQGILVPCQSPWNTP LLPVKKPGTNDYRPVQDLREVNKRVEDIHPTVPNPYNLLSGLPPSHQWYTVLDLKDAFFCLRLHPTSQP LFAFEWRDPEMGISGQLTWTRLPQGFKNSPTLFNEALHRDLADFRIQHPDLILLQYVDDLLLAATSELD
CQQGTRALLQTLGNLGYRASAKKAQICQKQVKYLGYLLKEGQRWLTEARKETVMGQPTPKTPRQLR EFLGKAGFCRLFIPGFAEMAAPLYPLTKPGTLFNWGPDQQKAYQEIKQALLTAPALGLPDLTKPFELFV DEKQGYAKGVLTQKLGPWRRPVAYLSKKLDPVAAGWPPCLRMVAAIAVLTKDAGKLTMGQPLVILA PHAVEALVKQPPDRWLSNARMTHYQALLLDTDRVQFGPWALNPATLLPLPEEGLQHNCLDILAEAH GTRPDLTDQPLPDADHTWYTDGSSLLQEGQRKAGAAVTTETEVIWAKALPAGTSAQRAELIALTQALK MAEGKKLNVYTDSRYAFATAHIHGEIYRRRGWLTSEGKEIKNKDEILALLKALFLPKRLSIIHCPGHQK GHSAEARGNRMADQAARKAAITETPDTSTLLIENSSPSGGSSGGSSGSETPGTSESATPESSGGSSGGSM AENGDNEKMAALEAKICHQIEYYFGDFNLPRDKFLKEQIKLDEGWVPLEIMIKFNRLNRLTTDFNVIVE
ALSKSKAELMEISEDKTKIRRSPSKPLPEVTDEYKNDVKNRSVYIKGFPTDATLDDIKEWLEDKGQVLNI QMRRTLHKAFKGSIFWFDSIESAKKFVETPGQKYKETDLLILFKDDYFAKKNEERKQNKVEAKLRAK QEQEAKQKLEEDAEMKSLEEKIGCLLKFSGDLDDQTCREDLHILFSNHGEIKWIDFVRGAKEGIILFKEK AKEALGKAKDANNGNLQLRNKEVTWEVLEGEVEKEALKKIIEDQQESLNKWKSKGRRFKGKGKGNK AAQPGSGKGKVQFQGKKTKFASDDEHDEHDENGATGPVKRAREETDKEEPASKQQKTENGAGDQSG GSKRTADGSEFESPKKKRKVGSGPAAKRVKLD*
SEQ ID NO: 10
MKRTADGSEFESPKKKRKVSGGSAENGDNEKMAALEAKICHQIEYYFGDFNLPRDKFLKEQIKLDEG WVPLEIMIKFNRLNRLTTDFNVIVEALSKSKAELMEISEDKTKIRRSPSKPLPEVTDEYKNDVKNRSVYI
KGFPTDATLDDIKEWLEDKGQVLNIQMRRTLHKAFKGSIFWFDSIESAKKFVETPGQKYKETDLLILF KDDYFAKKNEERKQNKVEAKLRAKQEQEAKQKLEEDAEMKSLEEKIGCLLKFSGDLDDQTCREDLHI LFSNHGEIKWIDFVRGAKEGIILFKEKAKEALGKAKDANNGNLQLRNKEVTWEVLEGEVEKEALKKIIE DQQESLNKWKSKGRRFKGKGKGNKAAQPGSGKGKVQFQGKKTKFASDDEHDEHDENGATGPVKRA REETDKEEPASKQQKTENGAGDQSGGSSGGSSGSETPGTSESATPESSGGSSGGSDKKYSIGLDIGTNSV GWAVITDEYKVPSKKFKVLGNTDRHSIKKNLIGALLFDSGETAEATRLKRTARRRYTRRKNRICYLQEI FSNEMAKVDDSFFHRLEESFLVEEDKKHERHPIFGNIVDEVAYHEKYPTIYHLRKKLVDSTDKADLRLI YLALAHMIKFRGHFLIEGDLNPDNSDVDKLFIQLVQTYNQLFEENPINASGVDAKAILSARLSKSRKLEN
LIAQLPGEKKNGLFGNLIALSLGLTPNFKSNFDLAEDAKLQLSKDTYDDDLDNLLAQIGDQYADLFLAA KNLSDAILLSDILRVNTEITKAPLSASMIKRYDEHHQDLTLLKALVRQQLPEKYKEIFFDQSKNGYAGYI DGGASQEEFYKFIKPILEKMDGTEELLVKLKREDLLRKQRTFDNGSIPHQIHLGELHAILRRQEDFYPFL KDNREKIEKILTFRIPYYVGPLARGNSRFAWMTRKSEETITPWNFEEWDKGASAQSFIERMTNFDKNL PNEKVLPKHSLLYEYFTVYNELTKVKYVTEGMRKPAFLSGEQKKAIVDLLFKTNRKVTVKQLKEDYF KKIECFDSVEISGVEDRFNASLGTYHDLLKIIKDKDFLDNEENEDILEDIVLTLTLFEDREMIEERLKTYA HLFDDKVMKQLKRRRYTGWGRLSRKLINGIRDKQSGKTILDFLKSDGFANRNFMQLIHDDSLTFKEDI QKAQVSGQGDSLHEHIANLAGSPAIKKGILQTVKVVDELVKVMGRHKPENIVIEMARENQTTQKGQK
NSRERMKRIEEGIKELGSQILKEHPVENTQLQNEKLYLYYLQNGRDMYVDQELDINRLSDYDVDAIVP
QSFLKDDSIDNKVLTRSDKNRGKSDNVPSEEWKKMKNYWRQLLNAKLITQRKFDNLTKAERGGLSE
LDKAGFIKRQLVETRQITKHVAQILDSRMNTKYDENDKLIREVKVITLKSKLVSDFRKDFQFYKVREIN
NYHHAHDAYLNAVVGTALIKKYPKLESEFVYGDYKVYDVRKMIAKSEQEIGKATAKYFFYSNIMNFF
KTEITLANGEIRKRPLIETNGETGEIVWDKGRDFATVRKVLSMPQVNIVKKTEVQTGGFSKESILPKRNS
DKLIARKKDWDPKKYGGFDSPTVAYSVLWAKVEKGKSKKLKSVKELLGITIMERSSFEKNPIDFLEAK
GYKEVKKDLIIKLPKYSLFELENGRKRMLASAGELQKGNELALPSKYVNFLYLASHYEKLKGSPEDNE
QKQLFVEQHKHYLDEIIEQISEFSKRVILADANLDKVLSAYNKHRDKPIREQAENIIHLFTLTNLGAPAAF
KYFDTTIDRKRYTSTKEVLDATLIHQSITGLYETRIDLSQLGGDSGGSSGGSKRTADGSEFESPKKKRKV
SGGSSGGSTLNIEDEYRLHETSKEPDVSLGSTWLSDFPQAWAETGGMGLAVRQAPLIIPLKATSTPVSIK
QYPMSQEARLGIKPHIQRLLDQGILVPCQSPWNTPLLPVKKPGTNDYRPVQDLREVNKRVEDIHPTVPN
PYNLLSGLPPSHQWYTVLDLKDAFFCLRLHPTSQPLFAFEWRDPEMGISGQLTWTRLPQGFKNSPTLFN
EALHRDLADFRIQHPDLILLQYVDDLLLAATSELDCQQGTRALLQTLGNLGYRASAKKAQICQKQVKY
LGYLLKEGQRWLTEARKETVMGQPTPKTPRQLREFLGKAGFCRLFIPGFAEMAAPLYPLTKPGTLFNW
GPDQQKAYQEIKQALLTAPALGLPDLTKPFELFVDEKQGYAKGVLTQKLGPWRRPVAYLSKKLDPVA
AGWPPCLRMVAAIAVLTKDAGKLTMGQPLVILAPHAVEALVKQPPDRWLSNARMTHYQALLLDTDR
VQFGPWALNPATLLPLPEEGLQHNCLDILAEAHGTRPDLTDQPLPDADHTWYTDGSSLLQEGQRKAG
AAVTTETEVIWAKALPAGTSAQRAELIALTQALKMAEGKKLNVYTDSRYAFATAHIHGEIYRRRGWLT
SEGKEIKNKDEILALLKALFLPKRLSIIHCPGHQKGHSAEARGNRMADQAARKAAITETPDTSTLLIENS
SPSGGSKRTADGSEFESPKKKRKVGSGPAAKRVKLD*
SEQ ID NO: 11
MKRTADGSEFESPKKKRKVDKKYSIGLDIGTNSVGWAVITDEYKVPSKKFKVLGNTDRHSIKKNLIGA
LLFDSGETAEATRLKRTARRRYTRRKNRICYLQEIFSNEMAKVDDSFFHRLEESFLVEEDKKHERHPIFG
NIVDEVAYHEKYPTIYHLRKKLVDSTDKADLRLIYLALAHMIKFRGHFLIEGDLNPDNSDVDKLFIQLV
QTYNQLFEENPINASGVDAKAILSARLSKSRKLENLIAQLPGEKKNGLFGNLIALSLGLTPNFKSNFDLA
EDAKLQLSKDTYDDDLDNLLAQIGDQYADLFLAAKNLSDAILLSDILRVNTEITKAPLSASMIKRYDEH
HQDLTLLKALVRQQLPEKYKEIFFDQSKNGYAGYIDGGASQEEFYKFIKPILEKMDGTEELLVKLKRED
LLRKQRTFDNGSIPHQIHLGELHAILRRQEDFYPFLKDNREKIEKILTFRIPYYVGPLARGNSRFAWMTR
KSEETITPWNFEEWDKGASAQSFIERMTNFDKNLPNEKVLPKHSLLYEYFTVYNELTKVKYVTEGMR
KPAFLSGEQKKAIVDLLFKTNRKVTVKQLKEDYFKKIECFDSVEISGVEDRFNASLGTYHDLLKIIKDKD
FLDNEENEDILEDIVLTLTLFEDREMIEERLKTYAHLFDDKVMKQLKRRRYTGWGRLSRKLINGIRDKQ
SGKTILDFLKSDGFANRNFMQLIHDDSLTFKEDIQKAQVSGQGDSLHEHIANLAGSPAIKKGILQTVKV
VDELVKVMGRHKPENIVIEMARENQTTQKGQKNSRERMKRIEEGIKELGSQILKEHPVENTQLQNEKL
YLYYLQNGRDMYVDQELDINRLSDYDVDAIVPQSFLKDDSIDNKVLTRSDKNRGKSDNVPSEEWKK
MKNYWRQLLNAKLITQRKFDNLTKAERGGLSELDKAGFIKRQLVETRQITKHVAQILDSRMNTKYDE
NDKLIREVKVITLKSKLVSDFRKDFQFYKVREINNYHHAHDAYLNAWGTALIKKYPKLESEFVYGDY
KVYDVRKMIAKSEQEIGKATAKYFFYSNIMNFFKTEITLANGEIRKRPLIETNGETGEIVWDKGRDFAT
VRKVLSMPQVNIVKKTEVQTGGFSKESILPKRNSDKLIARKKDWDPKKYGGFDSPTVAYSVLWAKVE
KGKSKKLKSVKELLGITIMERSSFEKNPIDFLEAKGYKEVKKDLIIKLPKYSLFELENGRKRMLASAGEL
QKGNELALPSKYVNFLYLASHYEKLKGSPEDNEQKQLFVEQHKHYLDEIIEQISEFSKRVILADANLDK
VLSAYNKHRDKPIREQAENIIHLFTLTNLGAPAAFKYFDTTIDRKRYTSTKEVLDATLIHQSITGLYETRI
DLSQLGGDSGGSSGGSSGSETPGTSESATPESSGGSSGGSMAENGDNEKMAALEAKICHQIEYYFGDFN
LPRDKFLKEQIKLDEGWVPLEIMIKFNRLNRLTTDFNVIVEALSKSKAELMEISEDKTKIRRSPSKPLPEV
TDEYKNDVKNRSVYIKGFPTDATLDDIKEWLEDKGQVLNIQMRRTLHKAFKGSIFWFDSIESAKKFVE
TPGQKYKETDLLILFKDDYFAKKNEERKQNKVEAKLRAKQEQEAKQKLEEDAEMKSLEEKIGCLLKFS
GDLDDQTCREDLHILFSNHGEIKWIDFVRGAKEGIILFKEKAKEALGKAKDANNGNLQLRNKEVTWEV
LEGEVEKEALKKIIEDQQESLNKWKSKGRRFKGKGKGNKAAQPGSGKGKVQFQGKKTKFASDDEHDE
HDENGATGPVKRAREETDKEEPASKQQKTENGAGDQSGGSSGGSKRTADGSEFESPKKKRKVSGGSS
GGSTLNIEDEYRLHETSKEPDVSLGSTWLSDFPQAWAETGGMGLAVRQAPLIIPLKATSTPVSIKQYPM
SQEARLGIKPHIQRLLDQGILVPCQSPWNTPLLPVKKPGTNDYRPVQDLREVNKRVEDIHPTVPNPYNL
LSGLPPSHQWYTVLDLKDAFFCLRLHPTSQPLFAFEWRDPEMGISGQLTWTRLPQGFKNSPTLFNEALH
RDLADFRIQHPDLILLQYVDDLLLAATSELDCQQGTRALLQTLGNLGYRASAKKAQICQKQVKYLGYL
LKEGQRWLTEARKETVMGQPTPKTPRQLREFLGKAGFCRLFIPGFAEMAAPLYPLTKPGTLFNWGPDQ
QKAYQEIKQALLTAPALGLPDLTKPFELFVDEKQGYAKGVLTQKLGPWRRPVAYLSKKLDPVAAGWP
PCLRMVAAIAVLTKDAGKLTMGQPLVILAPHAVEALVKQPPDRWLSNARMTHYQALLLDTDRVQFGP
WALNPATLLPLPEEGLQHNCLDILAEAHGTRPDLTDQPLPDADHTWYTDGSSLLQEGQRKAGAAVTT
ETEVIWAKALPAGTSAQRAELIALTQALKMAEGKKLNVYTDSRYAFATAHIHGEIYRRRGWLTSEGKE
IKNKDEILALLKALFLPKRLSIIHCPGHQKGHSAEARGNRMADQAARKAAITETPDTSTLLIENSSPSGGS
KRTADGSEFESPKKKRKVGSGPAAKRVKLD*
SEQ ID NO: 12
MKRTADGSEFESPKKKRKVDKKYSIGLDIGTNSVGWAVITDEYKVPSKKFKVLGNTDRHSIKKNLIGA LLFDSGETAEATRLKRTARRRYTRRKNRICYLQEIFSNEMAKVDDSFFHRLEESFLVEEDKKHERHPIFG NIVDEVAYHEKYPTIYHLRKKLVDSTDKADLRLIYLALAHMIKFRGHFLIEGDLNPDNSDVDKLFIQLV QTYNQLFEENPINASGVDAKAILSARLSKSRKLENLIAQLPGEKKNGLFGNLIALSLGLTPNFKSNFDLA EDAKLQLSKDTYDDDLDNLLAQIGDQYADLFLAAKNLSDAILLSDILRVNTEITKAPLSASMIKRYDEH HQDLTLLKALVRQQLPEKYKEIFFDQSKNGYAGYIDGGASQEEFYKFIKPILEKMDGTEELLVKLKRED LLRKQRTFDNGSIPHQIHLGELHAILRRQEDFYPFLKDNREKIEKILTFRIPYYVGPLARGNSRFAWMTR KSEETITPWNFEEWDKGASAQSFIERMTNFDKNLPNEKVLPKHSLLYEYFTVYNELTKVKYVTEGMR KPAFLSGEQKKAIVDLLFKTNRKVTVKQLKEDYFKKIECFDSVEISGVEDRFNASLGTYHDLLKIIKDKD FLDNEENEDILEDIVLTLTLFEDREMIEERLKTYAHLFDDKVMKQLKRRRYTGWGRLSRKLINGIRDKQ SGKTILDFLKSDGFANRNFMQLIHDDSLTFKEDIQKAQVSGQGDSLHEHIANLAGSPAIKKGILQTVKV VDELVKVMGRHKPENIVIEMARENQTTQKGQKNSRERMKRIEEGIKELGSQILKEHPVENTQLQNEKL YLYYLQNGRDMYVDQELDINRLSDYDVDAIVPQSFLKDDSIDNKVLTRSDKNRGKSDNVPSEEWKK MKNYWRQLLNAKLITQRKFDNLTKAERGGLSELDKAGFIKRQLVETRQITKHVAQILDSRMNTKYDE NDKLIREVKVITLKSKLVSDFRKDFQFYKVREINNYHHAHDAYLNAWGTALIKKYPKLESEFVYGDY KVYDVRKMIAKSEQEIGKATAKYFFYSNIMNFFKTEITLANGEIRKRPLIETNGETGEIVWDKGRDFAT VRKVLSMPQVNIVKKTEVQTGGFSKESILPKRNSDKLIARKKDWDPKKYGGFDSPTVAYSVLWAKVE KGKSKKLKSVKELLGITIMERSSFEKNPIDFLEAKGYKEVKKDLIIKLPKYSLFELENGRKRMLASAGEL
QKGNELALPSKYVNFLYLASHYEKLKGSPEDNEQKQLFVEQHKHYLDEIIEQISEFSKRVILADANLDK VLSAYNKHRDKPIREQAENIIHLFTLTNLGAPAAFKYFDTTIDRKRYTSTKEVLDATLIHQSITGLYETRI DLSQLGGDSGGSSGGSKRTADGSEFESPKKKRKVSGGSSGGSMAENGDNEKMAALEAKICHQIEYYFG DFNLPRDKFLKEQIKLDEGWVPLEIMIKFNRLNRLTTDFNVIVEALSKSKAELMEISEDKTKIRRSPSKPL PEVTDEYKNDVKNRSVYIKGFPTDATLDDIKEWLEDKGQVLNIQMRRTLHKAFKGSIFWFDSIESAKK FVETPGQKYKETDLLILFKDDYFAKKNEERKQNKVEAKLRAKQEQEAKQKLEEDAEMKSLEEKIGCLL KFSGDLDDQTCREDLHILFSNHGEIKWIDFVRGAKEGIILFKEKAKEALGKAKDANNGNLQLRNKEVT WEVLEGEVEKEALKKIIEDQQESLNKWKSKGRRFKGKGKGNKAAQPGSGKGKVQFQGKKTKFASDD EHDEHDENGATGPVKRAREETDKEEPASKQQKTENGAGDQSGGSSGGSSGSETPGTSESATPESSGGSS GGSTLNIEDEYRLHETSKEPDVSLGSTWLSDFPQAWAETGGMGLAVRQAPLIIPLKATSTPVSIKQYPM SQEARLGIKPHIQRLLDQGILVPCQSPWNTPLLPVKKPGTNDYRPVQDLREVNKRVEDIHPTVPNPYNL LSGLPPSHQWYTVLDLKDAFFCLRLHPTSQPLFAFEWRDPEMGISGQLTWTRLPQGFKNSPTLFNEALH RDLADFRIQHPDLILLQYVDDLLLAATSELDCQQGTRALLQTLGNLGYRASAKKAQICQKQVKYLGYL LKEGQRWLTEARKETVMGQPTPKTPRQLREFLGKAGFCRLFIPGFAEMAAPLYPLTKPGTLFNWGPDQ QKAYQEIKQALLTAPALGLPDLTKPFELFVDEKQGYAKGVLTQKLGPWRRPVAYLSKKLDPVAAGWP PCLRMVAAIAVLTKDAGKLTMGQPLVILAPHAVEALVKQPPDRWLSNARMTHYQALLLDTDRVQFGP WALNPATLLPLPEEGLQHNCLDILAEAHGTRPDLTDQPLPDADHTWYTDGSSLLQEGQRKAGAAVTT ETEVIWAKALPAGTSAQRAELIALTQALKMAEGKKLNVYTDSRYAFATAHIHGEIYRRRGWLTSEGKE
IKNKDEILALLKALFLPKRLSIIHCPGHQKGHSAEARGNRMADQAARKAAITETPDTSTLLIENSSPSGGS KRTADGSEFESPKKKRKVGSGPAAKRVKLD*
SEQ ID NO: 13
ATGAAACGGACAGCCGACGGAAGCGAGTTCGAGTCACCAAAGAAGAAGCGGAAAGTCGACAAGA AGTACAGCATCGGCCTGGACATCGGCACCAACTCTGTGGGCTGGGCCGTGATCACCGACGAGTAC AAGGTGCCCAGCAAGAAATTCAAGGTGCTGGGCAACACCGACCGGCACAGCATCAAGAAGAACC TGATCGGAGCCCTGCTGTTCGACAGCGGCGAAACAGCCGAGGCCACCCGGCTGAAGAGAACCGCC AGAAGAAGATACACCAGACGGAAGAACCGGATCTGCTATCTGCAAGAGATCTTCAGCAACGAGAT GGCCAAGGTGGACGACAGCTTCTTCCACAGACTGGAAGAGTCCTTCCTGGTGGAAGAGGATAAGA AGCACGAGCGGCACCCCATCTTCGGCAACATCGTGGACGAGGTGGCCTACCACGAGAAGTACCCC ACCATCTACCACCTGAGAAAGAAACTGGTGGACAGCACCGACAAGGCCGACCTGCGGCTGATCTA TCTGGCCCTGGCCCACATGATCAAGTTCCGGGGCCACTTCCTGATCGAGGGCGACCTGAACCCCGA CAACAGCGACGTGGACAAGCTGTTCATCCAGCTGGTGCAGACCTACAACCAGCTGTTCGAGGAAA ACCCCATCAACGCCAGCGGCGTGGACGCCAAGGCCATCCTGTCTGCCAGACTGAGCAAGAGCAGA CGGCTGGAAAATCTGATCGCCCAGCTGCCCGGCGAGAAGAAGAATGGCCTGTTCGGAAACCTGAT TGCCCTGAGCCTGGGCCTGACCCCCAACTTCAAGAGCAACTTCGACCTGGCCGAGGATGCCAAACT GCAGCTGAGCAAGGACACCTACGACGACGACCTGGACAACCTGCTGGCCCAGATCGGCGACCAGT ACGCCGACCTGTTTCTGGCCGCCAAGAACCTGTCCGACGCCATCCTGCTGAGCGACATCCTGAGAG TGAACACCGAGATCACCAAGGCCCCCCTGAGCGCCTCTATGATCAAGAGATACGACGAGCACCAC CAGGACCTGACCCTGCTGAAAGCTCTCGTGCGGCAGCAGCTGCCTGAGAAGTACAAAGAGATTTT CTTCGACCAGAGCAAGAACGGCTACGCCGGCTACATTGACGGCGGAGCCAGCCAGGAAGAGTTCT ACAAGTTCATCAAGCCCATCCTGGAAAAGATGGACGGCACCGAGGAACTGCTCGTGAAGCTGAAC
AGAGAGGACCTGCTGCGGAAGCAGCGGACCTTCGACAACGGCAGCATCCCCCACCAGATCCACCT
GGGAGAGCTGCACGCCATTCTGCGGCGGCAGGAAGATTTTTACCCATTCCTGAAGGACAACCGGG
AAAAGATCGAGAAGATCCTGACCTTCCGCATCCCCTACTACGTGGGCCCTCTGGCCAGGGGAAAC
AGCAGATTCGCCTGGATGACCAGAAAGAGCGAGGAAACCATCACCCCCTGGAACTTCGAGGAAGT
GGTGGACAAGGGCGCTTCCGCCCAGAGCTTCATCGAGCGGATGACCAACTTCGATAAGAACCTGC
CCAACGAGAAGGTGCTGCCCAAGCACAGCCTGCTGTACGAGTACTTCACCGTGTATAACGAGCTG
ACCAAAGTGAAATACGTGACCGAGGGAATGAGAAAGCCCGCCTTCCTGAGCGGCGAGCAGAAAA
AGGCCATCGTGGACCTGCTGTTCAAGACCAACCGGAAAGTGACCGTGAAGCAGCTGAAAGAGGAC
TACTTCAAGAAAATCGAGTGCTTCGACTCCGTGGAAATCTCCGGCGTGGAAGATCGGTTCAACGCC
TCCCTGGGCACATACCACGATCTGCTGAAAATTATCAAGGACAAGGACTTCCTGGACAATGAGGA
AAACGAGGACATTCTGGAAGATATCGTGCTGACCCTGACACTGTTTGAGGACAGAGAGATGATCG
AGGAACGGCTGAAAACCTATGCCCACCTGTTCGACGACAAAGTGATGAAGCAGCTGAAGCGGCGG
AGATACACCGGCTGGGGCAGGCTGAGCCGGAAGCTGATCAACGGCATCCGGGACAAGCAGTCCG
GCAAGACAATCCTGGATTTCCTGAAGTCCGACGGCTTCGCCAACAGAAACTTCATGCAGCTGATCC
ACGACGACAGCCTGACCTTTAAAGAGGACATCCAGAAAGCCCAGGTGTCCGGCCAGGGCGATAGC
CTGCACGAGCACATTGCCAATCTGGCCGGCAGCCCCGCCATTAAGAAGGGCATCCTGCAGACAGT
GAAGGTGGTGGACGAGCTCGTGAAAGTGATGGGCCGGCACAAGCCCGAGAACATCGTGATCGAA
ATGGCCAGAGAGAACCAGACCACCCAGAAGGGACAGAAGAACAGCCGCGAGAGAATGAAGCGG
ATCGAAGAGGGCATCAAAGAGCTGGGCAGCCAGATCCTGAAAGAACACCCCGTGGAAAACACCC
AGCTGCAGAACGAGAAGCTGTACCTGTACTACCTGCAGAATGGGCGGGATATGTACGTGGACCAG
GAACTGGACATCAACCGGCTGTCCGACTACGATGTGGACGCTATCGTGCCTCAGAGCTTTCTGAAG
GACGACTCCATCGACAACAAGGTGCTGACCAGAAGCGACAAGAACCGGGGCAAGAGCGACAACG
TGCCCTCCGAAGAGGTCGTGAAGAAGATGAAGAACTACTGGCGGCAGCTGCTGAACGCCAAGCTG
ATTACCCAGAGAAAGTTCGACAATCTGACCAAGGCCGAGAGAGGCGGCCTGAGCGAACTGGATAA
GGCCGGCTTCATCAAGAGACAGCTGGTGGAAACCCGGCAGATCACAAAGCACGTGGCACAGATCC
TGGACTCCCGGATGAACACTAAGTACGACGAGAATGACAAGCTGATCCGGGAAGTGAAAGTGATC
ACCCTGAAGTCCAAGCTGGTGTCCGATTTCCGGAAGGATTTCCAGTTTTACAAAGTGCGCGAGATC
AACAACTACCACCACGCCCACGACGCCTACCTGAACGCCGTCGTGGGAACCGCCCTGATCAAAAA
GTACCCTAAGCTGGAAAGCGAGTTCGTGTACGGCGACTACAAGGTGTACGACGTGCGGAAGATGA
TCGCCAAGAGCGAGCAGGAAATCGGCAAGGCTACCGCCAAGTACTTCTTCTACAGCAACATCATG
AACTTTTTCAAGACCGAGATTACCCTGGCCAACGGCGAGATCCGGAAGCGGCCTCTGATCGAGAC
AAACGGCGAAACCGGGGAGATCGTGTGGGATAAGGGCCGGGATTTTGCCACCGTGCGGAAAGTGC
TGAGCATGCCCCAAGTGAATATCGTGAAAAAGACCGAGGTGCAGACAGGCGGCTTCAGCAAAGA
GTCTATCCTGCCCAAGAGGAACAGCGATAAGCTGATCGCCAGAAAGAAGGACTGGGACCCTAAGA
AGTACGGCGGCTTCGACAGCCCCACCGTGGCCTATTCTGTGCTGGTGGTGGCCAAAGTGGAAAAG
GGCAAGTCCAAGAAACTGAAGAGTGTGAAAGAGCTGCTGGGGATCACCATCATGGAAAGAAGCA
GCTTCGAGAAGAATCCCATCGACTTTCTGGAAGCCAAGGGCTACAAAGAAGTGAAAAAGGACCTG
ATCATCAAGCTGCCTAAGTACTCCCTGTTCGAGCTGGAAAACGGCCGGAAGAGAATGCTGGCCTCT
GCCGGCGAACTGCAGAAGGGAAACGAACTGGCCCTGCCCTCCAAATATGTGAACTTCCTGTACCT
GGCCAGCCACTATGAGAAGCTGAAGGGCTCCCCCGAGGATAATGAGCAGAAACAGCTGTTTGTGG
AACAGCACAAGCACTACCTGGACGAGATCATCGAGCAGATCAGCGAGTTCTCCAAGAGAGTGATC
CTGGCCGACGCTAATCTGGACAAAGTGCTGTCCGCCTACAACAAGCACCGGGATAAGCCCATCAG
AGAGCAGGCCGAGAATATCATCCACCTGTTTACCCTGACCAATCTGGGAGCCCCTGCCGCCTTCAA
GTACTTTGACACCACCATCGACCGGAAGAGGTACACCAGCACCAAAGAGGTGCTGGACGCCACCC
TGATCCACCAGAGCATCACCGGCCTGTACGAGACACGGATCGACCTGTCTCAGCTGGGAGGTGAC
TCTGGAGGATCTAGCGGAGGATCCTCTGGCAGCGAGACACCAGGAACAAGCGAGTCAGCAACACC
AGAGAGCAGTGGCGGCAGCAGCGGCGGCAGCAGCACCCTAAATATAGAAGATGAGTATCGGCTA
CATGAGACCTCAAAAGAGCCAGATGTTTCTCTAGGGTCCACATGGCTGTCTGATTTTCCTCAGGCC
TGGGCGGAAACCGGGGGCATGGGACTGGCAGTTCGCCAAGCTCCTCTGATCATACCTCTGAAAGC
AACCTCTACCCCCGTGTCCATAAAACAATACCCCATGTCACAAGAAGCCAGACTGGGGATCAAGC
CCCACATACAGAGACTGTTGGACCAGGGAATACTGGTACCCTGCCAGTCCCCCTGGAACACGCCC
CTGCTACCCGTTAAGAAACCAGGGACTAATGATTATAGGCCTGTCCAGGATCTGAGAGAAGTCAA
CAAGCGGGTGGAAGACATCCACCCCACCGTGCCCAACCCTTACAACCTCTTGAGCGGGCTCCCACC
GTCCCACCAGTGGTACACTGTGCTTGATTTAAAGGATGCCTTTTTCTGCCTGAGACTCCACCCCACC
AGTCAGCCTCTCTTCGCCTTTGAGTGGAGAGATCCAGAGATGGGAATCTCAGGACAATTGACCTGG
ACCAGACTCCCACAGGGTTTCAAAAACAGTCCCACCCTGTTTAATGAGGCACTGCACAGAGACCT
AGCAGACTTCCGGATCCAGCACCCAGACTTGATCCTGCTACAGTACGTGGATGACTTACTGCTGGC
CGCCACTTCTGAGCTAGACTGCCAACAAGGTACTCGGGCCCTGTTACAAACCCTAGGGAACCTCGG
GTATCGGGCCTCGGCCAAGAAAGCCCAAATTTGCCAGAAACAGGTCAAGTATCTGGGGTATCTTCT
AAAAGAGGGTCAGAGATGGCTGACTGAGGCCAGAAAAGAGACTGTGATGGGGCAGCCTACTCCG
AAGACCCCTCGACAACTAAGGGAGTTCCTAGGGAAGGCAGGCTTCTGTCGCCTCTTCATCCCTGGG TTTGCAGAAATGGCAGCCCCCCTGTACCCTCTCACCAAACCGGGGACTCTGTTTAATTGGGGCCCA GACCAACAAAAGGCCTATCAAGAAATCAAGCAAGCTCTTCTAACTGCCCCAGCCCTGGGGTTGCC
AGATTTGACTAAGCCCTTTGAACTCTTTGTCGACGAGAAGCAGGGCTACGCCAAAGGTGTCCTAAC GCAAAAACTGGGACCTTGGCGTCGGCCGGTGGCCTACCTGTCCAAAAAGCTAGACCCAGTAGCAG CTGGGTGGCCCCCTTGCCTACGGATGGTAGCAGCCATTGCCGTACTGACAAAGGATGCAGGCAAG
CTAACCATGGGACAGCCACTAGTCATTCTGGCCCCCCATGCAGTAGAGGCACTAGTCAAACAACC
CCCCGACCGCTGGCTTTCCAACGCCCGGATGACTCACTATCAGGCCTTGCTTTTGGACACGGACCG GGTCCAGTTCGGACCGGTGGTAGCCCTGAACCCGGCTACGCTGCTCCCACTGCCTGAGGAAGGGCT GCAACACAACTGCCTTGATATCCTGGCCGAAGCCCACGGAACCCGACCCGACCTAACGGACCAGC
CGCTCCCAGACGCCGACCACACCTGGTACACGGATGGAAGCAGTCTCTTACAAGAGGGACAGCGT AAGGCGGGAGCTGCGGTGACCACCGAGACCGAGGTAATCTGGGCTAAAGCCCTGCCAGCCGGGAC ATCCGCTCAGCGGGCTGAACTGATAGCACTCACCCAGGCCCTAAAGATGGCAGAAGGTAAGAAGC
TAAATGTTTATACTGATAGCCGTTATGCTTTTGCTACTGCCCATATCCATGGAGAAATATACAGAA GGCGTGGGTGGCTCACATCAGAAGGCAAAGAGATCAAAAATAAAGACGAGATCTTGGCCCTACTA AAAGCCCTCTTTCTGCCCAAAAGACTTAGCATAATCCATTGTCCAGGACATCAAAAGGGACACAG
CGCCGAGGCTAGAGGCAACCGGATGGCTGACCAAGCGGCCCGAAAGGCAGCCATCACAGAGACT CCAGACACCTCTACCCTCCTCATAGAAAATTCATCACCCTCCGGAGGATCTAGCGGAGGCTCCTCT GGCTCTGAGACACCTGGCACAAGCGAGAGCGCAACACCTGAAAGCAGCGGGGGCAGCAGCGGGG
GGTCAATGGCTGAAAATGGTGATAATGAAAAGATGGCTGCCCTGGAGGCCAAAATCTGTCATCAA ATTGAGTATTATTTTGGCGACTTCAATTTGCCACGGGACAAGTTTCTAAAGGAACAGATAAAACTG GATGAAGGCTGGGTACCTTTGGAGATAATGATAAAATTCAACAGGTTGAACCGTCTAACAACAGA
CTTTAATGTAATTGTGGAAGCATTGAGCAAATCCAAGGCAGAACTCATGGAAATCAGTGAAGATA AAACTAAAATCAGAAGGTCTCCAAGCAAACCCCTACCTGAAGTGACTGATGAGTATAAAAATGAT GTAAAAAACAGATCTGTTTATATTAAAGGCTTCCCAACTGATGCAACTCTTGATGACATAAAAGAA
TGGTTAGAAGATAAAGGTCAAGTACTAAATATTCAGATGAGAAGAACATTGCATAAAGCATTTAA GGGATCAATTTTTGTTGTGTTTGATAGCATTGAATCTGCTAAGAAATTTGTAGAGACCCCTGGCCA GAAGTACAAAGAAACAGACCTGCTAATACTTTTCAAGGACGATTACTTTGCCAAAAAAAATGAAT
CTGGCGGCTCAAAAAGAACCGCCGACGGCAGCGAATTCGAGCCCAAGAAGAAGAGGAAAGTCTA A
SEQ ID NO: 14
ATGAAACGGACAGCCGACGGAAGCGAGTTCGAGTCACCAAAGAAGAAGCGGAAAGTCGCTGAAA ATGGTGATAATGAAAAGATGGCTGCCCTGGAGGCCAAAATCTGTCATCAAATTGAGTATTATTTTG GCGACTTCAATTTGCCACGGGACAAGTTTCTAAAGGAACAGATAAAACTGGATGAAGGCTGGGTA
CCTTTGGAGATAATGATAAAATTCAACAGGTTGAACCGTCTAACAACAGACTTTAATGTAATTGTG GAAGCATTGAGCAAATCCAAGGCAGAACTCATGGAAATCAGTGAAGATAAAACTAAAATCAGAA GGTCTCCAAGCAAACCCCTACCTGAAGTGACTGATGAGTATAAAAATGATGTAAAAAACAGATCT
GTTTATATTAAAGGCTTCCCAACTGATGCAACTCTTGATGACATAAAAGAATGGTTAGAAGATAAA GGTCAAGTACTAAATATTCAGATGAGAAGAACATTGCATAAAGCATTTAAGGGATCAATTTTTGTT GTGTTTGATAGCATTGAATCTGCTAAGAAATTTGTAGAGACCCCTGGCCAGAAGTACAAAGAAAC
AGACCTGCTAATACTTTTCAAGGACGATTACTTTGCCAAAAAAAATGAATCCGGAGGATCTAGCG GAGGCTCCTCTGGCTCTGAGACACCTGGCACAAGCGAGAGCGCAACACCTGAAAGCAGCGGGGGC AGCAGCGGGGGGTCAGACAAGAAGTACAGCATCGGCCTGGACATCGGCACCAACTCTGTGGGCTG
GGCCGTGATCACCGACGAGTACAAGGTGCCCAGCAAGAAATTCAAGGTGCTGGGCAACACCGACC GGCACAGCATCAAGAAGAACCTGATCGGAGCCCTGCTGTTCGACAGCGGCGAAACAGCCGAGGCC ACCCGGCTGAAGAGAACCGCCAGAAGAAGATACACCAGACGGAAGAACCGGATCTGCTATCTGC
AAGAGATCTTCAGCAACGAGATGGCCAAGGTGGACGACAGCTTCTTCCACAGACTGGAAGAGTCC TTCCTGGTGGAAGAGGATAAGAAGCACGAGCGGCACCCCATCTTCGGCAACATCGTGGACGAGGT GGCCTACCACGAGAAGTACCCCACCATCTACCACCTGAGAAAGAAACTGGTGGACAGCACCGACA
AGGCCGACCTGCGGCTGATCTATCTGGCCCTGGCCCACATGATCAAGTTCCGGGGCCACTTCCTGA TCGAGGGCGACCTGAACCCCGACAACAGCGACGTGGACAAGCTGTTCATCCAGCTGGTGCAGACC TACAACCAGCTGTTCGAGGAAAACCCCATCAACGCCAGCGGCGTGGACGCCAAGGCCATCCTGTC
TGCCAGACTGAGCAAGAGCAGACGGCTGGAAAATCTGATCGCCCAGCTGCCCGGCGAGAAGAAG AATGGCCTGTTCGGAAACCTGATTGCCCTGAGCCTGGGCCTGACCCCCAACTTCAAGAGCAACTTC GACCTGGCCGAGGATGCCAAACTGCAGCTGAGCAAGGACACCTACGACGACGACCTGGACAACCT
GCTGGCCCAGATCGGCGACCAGTACGCCGACCTGTTTCTGGCCGCCAAGAACCTGTCCGACGCCAT CCTGCTGAGCGACATCCTGAGAGTGAACACCGAGATCACCAAGGCCCCCCTGAGCGCCTCTATGA TCAAGAGATACGACGAGCACCACCAGGACCTGACCCTGCTGAAAGCTCTCGTGCGGCAGCAGCTG
CCTGAGAAGTACAAAGAGATTTTCTTCGACCAGAGCAAGAACGGCTACGCCGGCTACATTGACGG
CGGAGCCAGCCAGGAAGAGTTCTACAAGTTCATCAAGCCCATCCTGGAAAAGATGGACGGCACCG
AGGAACTGCTCGTGAAGCTGAACAGAGAGGACCTGCTGCGGAAGCAGCGGACCTTCGACAACGGC
AGCATCCCCCACCAGATCCACCTGGGAGAGCTGCACGCCATTCTGCGGCGGCAGGAAGATTTTTAC
CCATTCCTGAAGGACAACCGGGAAAAGATCGAGAAGATCCTGACCTTCCGCATCCCCTACTACGT
GGGCCCTCTGGCCAGGGGAAACAGCAGATTCGCCTGGATGACCAGAAAGAGCGAGGAAACCATC
ACCCCCTGGAACTTCGAGGAAGTGGTGGACAAGGGCGCTTCCGCCCAGAGCTTCATCGAGCGGAT
GACCAACTTCGATAAGAACCTGCCCAACGAGAAGGTGCTGCCCAAGCACAGCCTGCTGTACGAGT
ACTTCACCGTGTATAACGAGCTGACCAAAGTGAAATACGTGACCGAGGGAATGAGAAAGCCCGCC
TTCCTGAGCGGCGAGCAGAAAAAGGCCATCGTGGACCTGCTGTTCAAGACCAACCGGAAAGTGAC
CGTGAAGCAGCTGAAAGAGGACTACTTCAAGAAAATCGAGTGCTTCGACTCCGTGGAAATCTCCG
GCGTGGAAGATCGGTTCAACGCCTCCCTGGGCACATACCACGATCTGCTGAAAATTATCAAGGAC
AAGGACTTCCTGGACAATGAGGAAAACGAGGACATTCTGGAAGATATCGTGCTGACCCTGACACT
GTTTGAGGACAGAGAGATGATCGAGGAACGGCTGAAAACCTATGCCCACCTGTTCGACGACAAAG
TGATGAAGCAGCTGAAGCGGCGGAGATACACCGGCTGGGGCAGGCTGAGCCGGAAGCTGATCAA
CGGCATCCGGGACAAGCAGTCCGGCAAGACAATCCTGGATTTCCTGAAGTCCGACGGCTTCGCCA
ACAGAAACTTCATGCAGCTGATCCACGACGACAGCCTGACCTTTAAAGAGGACATCCAGAAAGCC
CAGGTGTCCGGCCAGGGCGATAGCCTGCACGAGCACATTGCCAATCTGGCCGGCAGCCCCGCCAT
TAAGAAGGGCATCCTGCAGACAGTGAAGGTGGTGGACGAGCTCGTGAAAGTGATGGGCCGGCAC
AAGCCCGAGAACATCGTGATCGAAATGGCCAGAGAGAACCAGACCACCCAGAAGGGACAGAAGA
ACAGCCGCGAGAGAATGAAGCGGATCGAAGAGGGCATCAAAGAGCTGGGCAGCCAGATCCTGAA
AGAACACCCCGTGGAAAACACCCAGCTGCAGAACGAGAAGCTGTACCTGTACTACCTGCAGAATG
GGCGGGATATGTACGTGGACCAGGAACTGGACATCAACCGGCTGTCCGACTACGATGTGGACGCT
ATCGTGCCTCAGAGCTTTCTGAAGGACGACTCCATCGACAACAAGGTGCTGACCAGAAGCGACAA
GAACCGGGGCAAGAGCGACAACGTGCCCTCCGAAGAGGTCGTGAAGAAGATGAAGAACTACTGG
CGGCAGCTGCTGAACGCCAAGCTGATTACCCAGAGAAAGTTCGACAATCTGACCAAGGCCGAGAG
AGGCGGCCTGAGCGAACTGGATAAGGCCGGCTTCATCAAGAGACAGCTGGTGGAAACCCGGCAG
ATCACAAAGCACGTGGCACAGATCCTGGACTCCCGGATGAACACTAAGTACGACGAGAATGACAA
GCTGATCCGGGAAGTGAAAGTGATCACCCTGAAGTCCAAGCTGGTGTCCGATTTCCGGAAGGATTT
CCAGTTTTACAAAGTGCGCGAGATCAACAACTACCACCACGCCCACGACGCCTACCTGAACGCCG
TCGTGGGAACCGCCCTGATCAAAAAGTACCCTAAGCTGGAAAGCGAGTTCGTGTACGGCGACTAC
AAGGTGTACGACGTGCGGAAGATGATCGCCAAGAGCGAGCAGGAAATCGGCAAGGCTACCGCCA
AGTACTTCTTCTACAGCAACATCATGAACTTTTTCAAGACCGAGATTACCCTGGCCAACGGCGAGA
TCCGGAAGCGGCCTCTGATCGAGACAAACGGCGAAACCGGGGAGATCGTGTGGGATAAGGGCCG
GGATTTTGCCACCGTGCGGAAAGTGCTGAGCATGCCCCAAGTGAATATCGTGAAAAAGACCGAGG
TGCAGACAGGCGGCTTCAGCAAAGAGTCTATCCTGCCCAAGAGGAACAGCGATAAGCTGATCGCC
AGAAAGAAGGACTGGGACCCTAAGAAGTACGGCGGCTTCGACAGCCCCACCGTGGCCTATTCTGT
GCTGGTGGTGGCCAAAGTGGAAAAGGGCAAGTCCAAGAAACTGAAGAGTGTGAAAGAGCTGCTG
GGGATCACCATCATGGAAAGAAGCAGCTTCGAGAAGAATCCCATCGACTTTCTGGAAGCCAAGGG
CTACAAAGAAGTGAAAAAGGACCTGATCATCAAGCTGCCTAAGTACTCCCTGTTCGAGCTGGAAA
ACGGCCGGAAGAGAATGCTGGCCTCTGCCGGCGAACTGCAGAAGGGAAACGAACTGGCCCTGCCC
TCCAAATATGTGAACTTCCTGTACCTGGCCAGCCACTATGAGAAGCTGAAGGGCTCCCCCGAGGAT
AATGAGCAGAAACAGCTGTTTGTGGAACAGCACAAGCACTACCTGGACGAGATCATCGAGCAGAT
CAGCGAGTTCTCCAAGAGAGTGATCCTGGCCGACGCTAATCTGGACAAAGTGCTGTCCGCCTACA
ACAAGCACCGGGATAAGCCCATCAGAGAGCAGGCCGAGAATATCATCCACCTGTTTACCCTGACC
AATCTGGGAGCCCCTGCCGCCTTCAAGTACTTTGACACCACCATCGACCGGAAGAGGTACACCAG
CACCAAAGAGGTGCTGGACGCCACCCTGATCCACCAGAGCATCACCGGCCTGTACGAGACACGGA
TCGACCTGTCTCAGCTGGGAGGTGACTCTGGAGGATCTAGCGGAGGATCCTCTGGCAGCGAGACA
CCAGGAACAAGCGAGTCAGCAACACCAGAGAGCAGTGGCGGCAGCAGCGGCGGCAGCAGCACCC
TAAATATAGAAGATGAGTATCGGCTACATGAGACCTCAAAAGAGCCAGATGTTTCTCTAGGGTCC
ACATGGCTGTCTGATTTTCCTCAGGCCTGGGCGGAAACCGGGGGCATGGGACTGGCAGTTCGCCA
AGCTCCTCTGATCATACCTCTGAAAGCAACCTCTACCCCCGTGTCCATAAAACAATACCCCATGTC
ACAAGAAGCCAGACTGGGGATCAAGCCCCACATACAGAGACTGTTGGACCAGGGAATACTGGTAC
CCTGCCAGTCCCCCTGGAACACGCCCCTGCTACCCGTTAAGAAACCAGGGACTAATGATTATAGGC
CTGTCCAGGATCTGAGAGAAGTCAACAAGCGGGTGGAAGACATCCACCCCACCGTGCCCAACCCT
TACAACCTCTTGAGCGGGCTCCCACCGTCCCACCAGTGGTACACTGTGCTTGATTTAAAGGATGCC
TTTTTCTGCCTGAGACTCCACCCCACCAGTCAGCCTCTCTTCGCCTTTGAGTGGAGAGATCCAGAG
ATGGGAATCTCAGGACAATTGACCTGGACCAGACTCCCACAGGGTTTCAAAAACAGTCCCACCCT
GTTTAATGAGGCACTGCACAGAGACCTAGCAGACTTCCGGATCCAGCACCCAGACTTGATCCTGCT
ACAGTACGTGGATGACTTACTGCTGGCCGCCACTTCTGAGCTAGACTGCCAACAAGGTACTCGGGC
CCTGTTACAAACCCTAGGGAACCTCGGGTATCGGGCCTCGGCCAAGAAAGCCCAAATTTGCCAGA
AACAGGTCAAGTATCTGGGGTATCTTCTAAAAGAGGGTCAGAGATGGCTGACTGAGGCCAGAAAA GAGACTGTGATGGGGCAGCCTACTCCGAAGACCCCTCGACAACTAAGGGAGTTCCTAGGGAAGGC AGGCTTCTGTCGCCTCTTCATCCCTGGGTTTGCAGAAATGGCAGCCCCCCTGTACCCTCTCACCAAA
CCGGGGACTCTGTTTAATTGGGGCCCAGACCAACAAAAGGCCTATCAAGAAATCAAGCAAGCTCT TCTAACTGCCCCAGCCCTGGGGTTGCCAGATTTGACTAAGCCCTTTGAACTCTTTGTCGACGAGAA GCAGGGCTACGCCAAAGGTGTCCTAACGCAAAAACTGGGACCTTGGCGTCGGCCGGTGGCCTACC
TGTCCAAAAAGCTAGACCCAGTAGCAGCTGGGTGGCCCCCTTGCCTACGGATGGTAGCAGCCATT GCCGTACTGACAAAGGATGCAGGCAAGCTAACCATGGGACAGCCACTAGTCATTCTGGCCCCCCA TGCAGTAGAGGCACTAGTCAAACAACCCCCCGACCGCTGGCTTTCCAACGCCCGGATGACTCACTA
TCAGGCCTTGCTTTTGGACACGGACCGGGTCCAGTTCGGACCGGTGGTAGCCCTGAACCCGGCTAC GCTGCTCCCACTGCCTGAGGAAGGGCTGCAACACAACTGCCTTGATATCCTGGCCGAAGCCCACG GAACCCGACCCGACCTAACGGACCAGCCGCTCCCAGACGCCGACCACACCTGGTACACGGATGGA
AGCAGTCTCTTACAAGAGGGACAGCGTAAGGCGGGAGCTGCGGTGACCACCGAGACCGAGGTAAT CTGGGCTAAAGCCCTGCCAGCCGGGACATCCGCTCAGCGGGCTGAACTGATAGCACTCACCCAGG CCCTAAAGATGGCAGAAGGTAAGAAGCTAAATGTTTATACTGATAGCCGTTATGCTTTTGCTACTG
CCCATATCCATGGAGAAATATACAGAAGGCGTGGGTGGCTCACATCAGAAGGCAAAGAGATCAAA AATAAAGACGAGATCTTGGCCCTACTAAAAGCCCTCTTTCTGCCCAAAAGACTTAGCATAATCCAT TGTCCAGGACATCAAAAGGGACACAGCGCCGAGGCTAGAGGCAACCGGATGGCTGACCAAGCGG
CCCGAAAGGCAGCCATCACAGAGACTCCAGACACCTCTACCCTCCTCATAGAAAATTCATCACCCT CTGGCGGCTCAAAAAGAACCGCCGACGGCAGCGAATTCGAGCCCAAGAAGAAGAGGAAAGTCTA A
SEQ ID NO: 15
ATGAAACGGACAGCCGACGGAAGCGAGTTCGAGTCACCAAAGAAGAAGCGGAAAGTCTCTGGCG GCTCAGCTGAAAATGGTGATAATGAAAAGATGGCTGCCCTGGAGGCCAAAATCTGTCATCAAATT GAGTATTATTTTGGCGACTTCAATTTGCCACGGGACAAGTTTCTAAAGGAACAGATAAAACTGGAT
GAAGGCTGGGTACCTTTGGAGATAATGATAAAATTCAACAGGTTGAACCGTCTAACAACAGACTT TAATGTAATTGTGGAAGCATTGAGCAAATCCAAGGCAGAACTCATGGAAATCAGTGAAGATAAAA CTAAAATCAGAAGGTCTCCAAGCAAACCCCTACCTGAAGTGACTGATGAGTATAAAAATGATGTA
AAAAACAGATCTGTTTATATTAAAGGCTTCCCAACTGATGCAACTCTTGATGACATAAAAGAATGG TTAGAAGATAAAGGTCAAGTACTAAATATTCAGATGAGAAGAACATTGCATAAAGCATTTAAGGG ATCAATTTTTGTTGTGTTTGATAGCATTGAATCTGCTAAGAAATTTGTAGAGACCCCTGGCCAGAA
GTACAAAGAAACAGACCTGCTAATACTTTTCAAGGACGATTACTTTGCCAAAAAAAATGAATCCG GAGGATCTAGCGGAGGCTCCTCTGGCTCTGAGACACCTGGCACAAGCGAGAGCGCAACACCTGAA AGCAGCGGGGGCAGCAGCGGGGGGTCAGACAAGAAGTACAGCATCGGCCTGGACATCGGCACCA
ACTCTGTGGGCTGGGCCGTGATCACCGACGAGTACAAGGTGCCCAGCAAGAAATTCAAGGTGCTG GGCAACACCGACCGGCACAGCATCAAGAAGAACCTGATCGGAGCCCTGCTGTTCGACAGCGGCGA AACAGCCGAGGCCACCCGGCTGAAGAGAACCGCCAGAAGAAGATACACCAGACGGAAGAACCGG
ATCTGCTATCTGCAAGAGATCTTCAGCAACGAGATGGCCAAGGTGGACGACAGCTTCTTCCACAG ACTGGAAGAGTCCTTCCTGGTGGAAGAGGATAAGAAGCACGAGCGGCACCCCATCTTCGGCAACA TCGTGGACGAGGTGGCCTACCACGAGAAGTACCCCACCATCTACCACCTGAGAAAGAAACTGGTG
GACAGCACCGACAAGGCCGACCTGCGGCTGATCTATCTGGCCCTGGCCCACATGATCAAGTTCCG
GGGCCACTTCCTGATCGAGGGCGACCTGAACCCCGACAACAGCGACGTGGACAAGCTGTTCATCC AGCTGGTGCAGACCTACAACCAGCTGTTCGAGGAAAACCCCATCAACGCCAGCGGCGTGGACGCC AAGGCCATCCTGTCTGCCAGACTGAGCAAGAGCAGACGGCTGGAAAATCTGATCGCCCAGCTGCC
CGGCGAGAAGAAGAATGGCCTGTTCGGAAACCTGATTGCCCTGAGCCTGGGCCTGACCCCCAACT TCAAGAGCAACTTCGACCTGGCCGAGGATGCCAAACTGCAGCTGAGCAAGGACACCTACGACGAC GACCTGGACAACCTGCTGGCCCAGATCGGCGACCAGTACGCCGACCTGTTTCTGGCCGCCAAGAA
CCTGTCCGACGCCATCCTGCTGAGCGACATCCTGAGAGTGAACACCGAGATCACCAAGGCCCCCCT GAGCGCCTCTATGATCAAGAGATACGACGAGCACCACCAGGACCTGACCCTGCTGAAAGCTCTCG TGCGGCAGCAGCTGCCTGAGAAGTACAAAGAGATTTTCTTCGACCAGAGCAAGAACGGCTACGCC
GGCTACATTGACGGCGGAGCCAGCCAGGAAGAGTTCTACAAGTTCATCAAGCCCATCCTGGAAAA GATGGACGGCACCGAGGAACTGCTCGTGAAGCTGAACAGAGAGGACCTGCTGCGGAAGCAGCGG ACCTTCGACAACGGCAGCATCCCCCACCAGATCCACCTGGGAGAGCTGCACGCCATTCTGCGGCG
GCAGGAAGATTTTTACCCATTCCTGAAGGACAACCGGGAAAAGATCGAGAAGATCCTGACCTTCC GCATCCCCTACTACGTGGGCCCTCTGGCCAGGGGAAACAGCAGATTCGCCTGGATGACCAGAAAG AGCGAGGAAACCATCACCCCCTGGAACTTCGAGGAAGTGGTGGACAAGGGCGCTTCCGCCCAGAG
CTTCATCGAGCGGATGACCAACTTCGATAAGAACCTGCCCAACGAGAAGGTGCTGCCCAAGCACA GCCTGCTGTACGAGTACTTCACCGTGTATAACGAGCTGACCAAAGTGAAATACGTGACCGAGGGA ATGAGAAAGCCCGCCTTCCTGAGCGGCGAGCAGAAAAAGGCCATCGTGGACCTGCTGTTCAAGAC
CAACCGGAAAGTGACCGTGAAGCAGCTGAAAGAGGACTACTTCAAGAAAATCGAGTGCTTCGACT
CCGTGGAAATCTCCGGCGTGGAAGATCGGTTCAACGCCTCCCTGGGCACATACCACGATCTGCTGA
AAATTATCAAGGACAAGGACTTCCTGGACAATGAGGAAAACGAGGACATTCTGGAAGATATCGTG
CTGACCCTGACACTGTTTGAGGACAGAGAGATGATCGAGGAACGGCTGAAAACCTATGCCCACCT
GTTCGACGACAAAGTGATGAAGCAGCTGAAGCGGCGGAGATACACCGGCTGGGGCAGGCTGAGC
CGGAAGCTGATCAACGGCATCCGGGACAAGCAGTCCGGCAAGACAATCCTGGATTTCCTGAAGTC
CGACGGCTTCGCCAACAGAAACTTCATGCAGCTGATCCACGACGACAGCCTGACCTTTAAAGAGG
ACATCCAGAAAGCCCAGGTGTCCGGCCAGGGCGATAGCCTGCACGAGCACATTGCCAATCTGGCC
GGCAGCCCCGCCATTAAGAAGGGCATCCTGCAGACAGTGAAGGTGGTGGACGAGCTCGTGAAAGT
GATGGGCCGGCACAAGCCCGAGAACATCGTGATCGAAATGGCCAGAGAGAACCAGACCACCCAG
AAGGGACAGAAGAACAGCCGCGAGAGAATGAAGCGGATCGAAGAGGGCATCAAAGAGCTGGGC
AGCCAGATCCTGAAAGAACACCCCGTGGAAAACACCCAGCTGCAGAACGAGAAGCTGTACCTGTA
CTACCTGCAGAATGGGCGGGATATGTACGTGGACCAGGAACTGGACATCAACCGGCTGTCCGACT
ACGATGTGGACGCTATCGTGCCTCAGAGCTTTCTGAAGGACGACTCCATCGACAACAAGGTGCTG
ACCAGAAGCGACAAGAACCGGGGCAAGAGCGACAACGTGCCCTCCGAAGAGGTCGTGAAGAAGA
TGAAGAACTACTGGCGGCAGCTGCTGAACGCCAAGCTGATTACCCAGAGAAAGTTCGACAATCTG
ACCAAGGCCGAGAGAGGCGGCCTGAGCGAACTGGATAAGGCCGGCTTCATCAAGAGACAGCTGG
TGGAAACCCGGCAGATCACAAAGCACGTGGCACAGATCCTGGACTCCCGGATGAACACTAAGTAC
GACGAGAATGACAAGCTGATCCGGGAAGTGAAAGTGATCACCCTGAAGTCCAAGCTGGTGTCCGA
TTTCCGGAAGGATTTCCAGTTTTACAAAGTGCGCGAGATCAACAACTACCACCACGCCCACGACGC
CTACCTGAACGCCGTCGTGGGAACCGCCCTGATCAAAAAGTACCCTAAGCTGGAAAGCGAGTTCG
TGTACGGCGACTACAAGGTGTACGACGTGCGGAAGATGATCGCCAAGAGCGAGCAGGAAATCGG
CAAGGCTACCGCCAAGTACTTCTTCTACAGCAACATCATGAACTTTTTCAAGACCGAGATTACCCT
GGCCAACGGCGAGATCCGGAAGCGGCCTCTGATCGAGACAAACGGCGAAACCGGGGAGATCGTG
TGGGATAAGGGCCGGGATTTTGCCACCGTGCGGAAAGTGCTGAGCATGCCCCAAGTGAATATCGT
GAAAAAGACCGAGGTGCAGACAGGCGGCTTCAGCAAAGAGTCTATCCTGCCCAAGAGGAACAGC
GATAAGCTGATCGCCAGAAAGAAGGACTGGGACCCTAAGAAGTACGGCGGCTTCGACAGCCCCAC
CGTGGCCTATTCTGTGCTGGTGGTGGCCAAAGTGGAAAAGGGCAAGTCCAAGAAACTGAAGAGTG
TGAAAGAGCTGCTGGGGATCACCATCATGGAAAGAAGCAGCTTCGAGAAGAATCCCATCGACTTT
CTGGAAGCCAAGGGCTACAAAGAAGTGAAAAAGGACCTGATCATCAAGCTGCCTAAGTACTCCCT
GTTCGAGCTGGAAAACGGCCGGAAGAGAATGCTGGCCTCTGCCGGCGAACTGCAGAAGGGAAAC
GAACTGGCCCTGCCCTCCAAATATGTGAACTTCCTGTACCTGGCCAGCCACTATGAGAAGCTGAAG
GGCTCCCCCGAGGATAATGAGCAGAAACAGCTGTTTGTGGAACAGCACAAGCACTACCTGGACGA
GATCATCGAGCAGATCAGCGAGTTCTCCAAGAGAGTGATCCTGGCCGACGCTAATCTGGACAAAG
TGCTGTCCGCCTACAACAAGCACCGGGATAAGCCCATCAGAGAGCAGGCCGAGAATATCATCCAC
CTGTTTACCCTGACCAATCTGGGAGCCCCTGCCGCCTTCAAGTACTTTGACACCACCATCGACCGG
AAGAGGTACACCAGCACCAAAGAGGTGCTGGACGCCACCCTGATCCACCAGAGCATCACCGGCCT
GTACGAGACACGGATCGACCTGTCTCAGCTGGGAGGTGACTCTGGAGGATCTAGCGGAGGATCCT
CTGGCAGCGAGACACCAGGAACAAGCGAGTCAGCAACACCAGAGAGCAGTGGCGGCAGCAGCGG
CGGCAGCAGCACCCTAAATATAGAAGATGAGTATCGGCTACATGAGACCTCAAAAGAGCCAGATG
TTTCTCTAGGGTCCACATGGCTGTCTGATTTTCCTCAGGCCTGGGCGGAAACCGGGGGCATGGGAC
TGGCAGTTCGCCAAGCTCCTCTGATCATACCTCTGAAAGCAACCTCTACCCCCGTGTCCATAAAAC
AATACCCCATGTCACAAGAAGCCAGACTGGGGATCAAGCCCCACATACAGAGACTGTTGGACCAG
GGAATACTGGTACCCTGCCAGTCCCCCTGGAACACGCCCCTGCTACCCGTTAAGAAACCAGGGACT
AATGATTATAGGCCTGTCCAGGATCTGAGAGAAGTCAACAAGCGGGTGGAAGACATCCACCCCAC
CGTGCCCAACCCTTACAACCTCTTGAGCGGGCTCCCACCGTCCCACCAGTGGTACACTGTGCTTGA
TTTAAAGGATGCCTTTTTCTGCCTGAGACTCCACCCCACCAGTCAGCCTCTCTTCGCCTTTGAGTGG
AGAGATCCAGAGATGGGAATCTCAGGACAATTGACCTGGACCAGACTCCCACAGGGTTTCAAAAA
CAGTCCCACCCTGTTTAATGAGGCACTGCACAGAGACCTAGCAGACTTCCGGATCCAGCACCCAG
ACTTGATCCTGCTACAGTACGTGGATGACTTACTGCTGGCCGCCACTTCTGAGCTAGACTGCCAAC
AAGGTACTCGGGCCCTGTTACAAACCCTAGGGAACCTCGGGTATCGGGCCTCGGCCAAGAAAGCC
CAAATTTGCCAGAAACAGGTCAAGTATCTGGGGTATCTTCTAAAAGAGGGTCAGAGATGGCTGAC
TGAGGCCAGAAAAGAGACTGTGATGGGGCAGCCTACTCCGAAGACCCCTCGACAACTAAGGGAGT
TCCTAGGGAAGGCAGGCTTCTGTCGCCTCTTCATCCCTGGGTTTGCAGAAATGGCAGCCCCCCTGT
ACCCTCTCACCAAACCGGGGACTCTGTTTAATTGGGGCCCAGACCAACAAAAGGCCTATCAAGAA
ATCAAGCAAGCTCTTCTAACTGCCCCAGCCCTGGGGTTGCCAGATTTGACTAAGCCCTTTGAACTC
TTTGTCGACGAGAAGCAGGGCTACGCCAAAGGTGTCCTAACGCAAAAACTGGGACCTTGGCGTCG
GCCGGTGGCCTACCTGTCCAAAAAGCTAGACCCAGTAGCAGCTGGGTGGCCCCCTTGCCTACGGAT
GGTAGCAGCCATTGCCGTACTGACAAAGGATGCAGGCAAGCTAACCATGGGACAGCCACTAGTCA
TTCTGGCCCCCCATGCAGTAGAGGCACTAGTCAAACAACCCCCCGACCGCTGGCTTTCCAACGCCC
GGATGACTCACTATCAGGCCTTGCTTTTGGACACGGACCGGGTCCAGTTCGGACCGGTGGTAGCCC
TGAACCCGGCTACGCTGCTCCCACTGCCTGAGGAAGGGCTGCAACACAACTGCCTTGATATCCTGG
CCGAAGCCCACGGAACCCGACCCGACCTAACGGACCAGCCGCTCCCAGACGCCGACCACACCTGG
TACACGGATGGAAGCAGTCTCTTACAAGAGGGACAGCGTAAGGCGGGAGCTGCGGTGACCACCGA
GACCGAGGTAATCTGGGCTAAAGCCCTGCCAGCCGGGACATCCGCTCAGCGGGCTGAACTGATAG
CACTCACCCAGGCCCTAAAGATGGCAGAAGGTAAGAAGCTAAATGTTTATACTGATAGCCGTTAT
GCTTTTGCTACTGCCCATATCCATGGAGAAATATACAGAAGGCGTGGGTGGCTCACATCAGAAGG
CAAAGAGATCAAAAATAAAGACGAGATCTTGGCCCTACTAAAAGCCCTCTTTCTGCCCAAAAGAC
TTAGCATAATCCATTGTCCAGGACATCAAAAGGGACACAGCGCCGAGGCTAGAGGCAACCGGATG
GCTGACCAAGCGGCCCGAAAGGCAGCCATCACAGAGACTCCAGACACCTCTACCCTCCTCATAGA
AAATTCATCACCCTCTGGCGGCTCAAAAAGAACCGCCGACGGCAGCGAATTCGAGCCCAAGAAGA AGAGGAAAGTCTAA
SEQ ID NO: 16
ATGAAACGGACAGCCGACGGAAGCGAGTTCGAGTCACCAAAGAAGAAGCGGAAAGTCGACAAGA
AGTACAGCATCGGCCTGGACATCGGCACCAACTCTGTGGGCTGGGCCGTGATCACCGACGAGTAC
AAGGTGCCCAGCAAGAAATTCAAGGTGCTGGGCAACACCGACCGGCACAGCATCAAGAAGAACC
TGATCGGAGCCCTGCTGTTCGACAGCGGCGAAACAGCCGAGGCCACCCGGCTGAAGAGAACCGCC
AGAAGAAGATACACCAGACGGAAGAACCGGATCTGCTATCTGCAAGAGATCTTCAGCAACGAGAT
GGCCAAGGTGGACGACAGCTTCTTCCACAGACTGGAAGAGTCCTTCCTGGTGGAAGAGGATAAGA
AGCACGAGCGGCACCCCATCTTCGGCAACATCGTGGACGAGGTGGCCTACCACGAGAAGTACCCC
ACCATCTACCACCTGAGAAAGAAACTGGTGGACAGCACCGACAAGGCCGACCTGCGGCTGATCTA
TCTGGCCCTGGCCCACATGATCAAGTTCCGGGGCCACTTCCTGATCGAGGGCGACCTGAACCCCGA
CAACAGCGACGTGGACAAGCTGTTCATCCAGCTGGTGCAGACCTACAACCAGCTGTTCGAGGAAA
ACCCCATCAACGCCAGCGGCGTGGACGCCAAGGCCATCCTGTCTGCCAGACTGAGCAAGAGCAGA
CGGCTGGAAAATCTGATCGCCCAGCTGCCCGGCGAGAAGAAGAATGGCCTGTTCGGAAACCTGAT
TGCCCTGAGCCTGGGCCTGACCCCCAACTTCAAGAGCAACTTCGACCTGGCCGAGGATGCCAAACT
GCAGCTGAGCAAGGACACCTACGACGACGACCTGGACAACCTGCTGGCCCAGATCGGCGACCAGT
ACGCCGACCTGTTTCTGGCCGCCAAGAACCTGTCCGACGCCATCCTGCTGAGCGACATCCTGAGAG
TGAACACCGAGATCACCAAGGCCCCCCTGAGCGCCTCTATGATCAAGAGATACGACGAGCACCAC
CAGGACCTGACCCTGCTGAAAGCTCTCGTGCGGCAGCAGCTGCCTGAGAAGTACAAAGAGATTTT
CTTCGACCAGAGCAAGAACGGCTACGCCGGCTACATTGACGGCGGAGCCAGCCAGGAAGAGTTCT
ACAAGTTCATCAAGCCCATCCTGGAAAAGATGGACGGCACCGAGGAACTGCTCGTGAAGCTGAAC
AGAGAGGACCTGCTGCGGAAGCAGCGGACCTTCGACAACGGCAGCATCCCCCACCAGATCCACCT
GGGAGAGCTGCACGCCATTCTGCGGCGGCAGGAAGATTTTTACCCATTCCTGAAGGACAACCGGG
AAAAGATCGAGAAGATCCTGACCTTCCGCATCCCCTACTACGTGGGCCCTCTGGCCAGGGGAAAC
AGCAGATTCGCCTGGATGACCAGAAAGAGCGAGGAAACCATCACCCCCTGGAACTTCGAGGAAGT
GGTGGACAAGGGCGCTTCCGCCCAGAGCTTCATCGAGCGGATGACCAACTTCGATAAGAACCTGC
CCAACGAGAAGGTGCTGCCCAAGCACAGCCTGCTGTACGAGTACTTCACCGTGTATAACGAGCTG
ACCAAAGTGAAATACGTGACCGAGGGAATGAGAAAGCCCGCCTTCCTGAGCGGCGAGCAGAAAA
AGGCCATCGTGGACCTGCTGTTCAAGACCAACCGGAAAGTGACCGTGAAGCAGCTGAAAGAGGAC
TACTTCAAGAAAATCGAGTGCTTCGACTCCGTGGAAATCTCCGGCGTGGAAGATCGGTTCAACGCC
TCCCTGGGCACATACCACGATCTGCTGAAAATTATCAAGGACAAGGACTTCCTGGACAATGAGGA
AAACGAGGACATTCTGGAAGATATCGTGCTGACCCTGACACTGTTTGAGGACAGAGAGATGATCG
AGGAACGGCTGAAAACCTATGCCCACCTGTTCGACGACAAAGTGATGAAGCAGCTGAAGCGGCGG
AGATACACCGGCTGGGGCAGGCTGAGCCGGAAGCTGATCAACGGCATCCGGGACAAGCAGTCCG
GCAAGACAATCCTGGATTTCCTGAAGTCCGACGGCTTCGCCAACAGAAACTTCATGCAGCTGATCC
ACGACGACAGCCTGACCTTTAAAGAGGACATCCAGAAAGCCCAGGTGTCCGGCCAGGGCGATAGC
CTGCACGAGCACATTGCCAATCTGGCCGGCAGCCCCGCCATTAAGAAGGGCATCCTGCAGACAGT
GAAGGTGGTGGACGAGCTCGTGAAAGTGATGGGCCGGCACAAGCCCGAGAACATCGTGATCGAA
ATGGCCAGAGAGAACCAGACCACCCAGAAGGGACAGAAGAACAGCCGCGAGAGAATGAAGCGG
ATCGAAGAGGGCATCAAAGAGCTGGGCAGCCAGATCCTGAAAGAACACCCCGTGGAAAACACCC
AGCTGCAGAACGAGAAGCTGTACCTGTACTACCTGCAGAATGGGCGGGATATGTACGTGGACCAG
GAACTGGACATCAACCGGCTGTCCGACTACGATGTGGACGCTATCGTGCCTCAGAGCTTTCTGAAG
GACGACTCCATCGACAACAAGGTGCTGACCAGAAGCGACAAGAACCGGGGCAAGAGCGACAACG
TGCCCTCCGAAGAGGTCGTGAAGAAGATGAAGAACTACTGGCGGCAGCTGCTGAACGCCAAGCTG
ATTACCCAGAGAAAGTTCGACAATCTGACCAAGGCCGAGAGAGGCGGCCTGAGCGAACTGGATAA
GGCCGGCTTCATCAAGAGACAGCTGGTGGAAACCCGGCAGATCACAAAGCACGTGGCACAGATCC
TGGACTCCCGGATGAACACTAAGTACGACGAGAATGACAAGCTGATCCGGGAAGTGAAAGTGATC
ACCCTGAAGTCCAAGCTGGTGTCCGATTTCCGGAAGGATTTCCAGTTTTACAAAGTGCGCGAGATC
AACAACTACCACCACGCCCACGACGCCTACCTGAACGCCGTCGTGGGAACCGCCCTGATCAAAAA
GTACCCTAAGCTGGAAAGCGAGTTCGTGTACGGCGACTACAAGGTGTACGACGTGCGGAAGATGA
TCGCCAAGAGCGAGCAGGAAATCGGCAAGGCTACCGCCAAGTACTTCTTCTACAGCAACATCATG
AACTTTTTCAAGACCGAGATTACCCTGGCCAACGGCGAGATCCGGAAGCGGCCTCTGATCGAGAC
AAACGGCGAAACCGGGGAGATCGTGTGGGATAAGGGCCGGGATTTTGCCACCGTGCGGAAAGTGC
TGAGCATGCCCCAAGTGAATATCGTGAAAAAGACCGAGGTGCAGACAGGCGGCTTCAGCAAAGA
GTCTATCCTGCCCAAGAGGAACAGCGATAAGCTGATCGCCAGAAAGAAGGACTGGGACCCTAAGA
AGTACGGCGGCTTCGACAGCCCCACCGTGGCCTATTCTGTGCTGGTGGTGGCCAAAGTGGAAAAG
GGCAAGTCCAAGAAACTGAAGAGTGTGAAAGAGCTGCTGGGGATCACCATCATGGAAAGAAGCA
GCTTCGAGAAGAATCCCATCGACTTTCTGGAAGCCAAGGGCTACAAAGAAGTGAAAAAGGACCTG
ATCATCAAGCTGCCTAAGTACTCCCTGTTCGAGCTGGAAAACGGCCGGAAGAGAATGCTGGCCTCT
GCCGGCGAACTGCAGAAGGGAAACGAACTGGCCCTGCCCTCCAAATATGTGAACTTCCTGTACCT
GGCCAGCCACTATGAGAAGCTGAAGGGCTCCCCCGAGGATAATGAGCAGAAACAGCTGTTTGTGG
AACAGCACAAGCACTACCTGGACGAGATCATCGAGCAGATCAGCGAGTTCTCCAAGAGAGTGATC
CTGGCCGACGCTAATCTGGACAAAGTGCTGTCCGCCTACAACAAGCACCGGGATAAGCCCATCAG
AGAGCAGGCCGAGAATATCATCCACCTGTTTACCCTGACCAATCTGGGAGCCCCTGCCGCCTTCAA
GTACTTTGACACCACCATCGACCGGAAGAGGTACACCAGCACCAAAGAGGTGCTGGACGCCACCC
TGATCCACCAGAGCATCACCGGCCTGTACGAGACACGGATCGACCTGTCTCAGCTGGGAGGTGAC
TCCGGAGGATCTAGCGGAGGCTCCTCTGGCTCTGAGACACCTGGCACAAGCGAGAGCGCAACACC
TGAAAGCAGCGGGGGCAGCAGCGGGGGGTCAATGGCTGAAAATGGTGATAATGAAAAGATGGCT
GCCCTGGAGGCCAAAATCTGTCATCAAATTGAGTATTATTTTGGCGACTTCAATTTGCCACGGGAC
AAGTTTCTAAAGGAACAGATAAAACTGGATGAAGGCTGGGTACCTTTGGAGATAATGATAAAATT
CAACAGGTTGAACCGTCTAACAACAGACTTTAATGTAATTGTGGAAGCATTGAGCAAATCCAAGG
CAGAACTCATGGAAATCAGTGAAGATAAAACTAAAATCAGAAGGTCTCCAAGCAAACCCCTACCT
GAAGTGACTGATGAGTATAAAAATGATGTAAAAAACAGATCTGTTTATATTAAAGGCTTCCCAACT
GATGCAACTCTTGATGACATAAAAGAATGGTTAGAAGATAAAGGTCAAGTACTAAATATTCAGAT
GAGAAGAACATTGCATAAAGCATTTAAGGGATCAATTTTTGTTGTGTTTGATAGCATTGAATCTGC
TAAGAAATTTGTAGAGACCCCTGGCCAGAAGTACAAAGAAACAGACCTGCTAATACTTTTCAAGG
ACGATTACTTTGCCAAAAAAAATGAATCTGGAGGATCTAGCGGAGGATCCTCTGGCAGCGAGACA
CCAGGAACAAGCGAGTCAGCAACACCAGAGAGCAGTGGCGGCAGCAGCGGCGGCAGCAGCACCC
TAAATATAGAAGATGAGTATCGGCTACATGAGACCTCAAAAGAGCCAGATGTTTCTCTAGGGTCC
ACATGGCTGTCTGATTTTCCTCAGGCCTGGGCGGAAACCGGGGGCATGGGACTGGCAGTTCGCCA
AGCTCCTCTGATCATACCTCTGAAAGCAACCTCTACCCCCGTGTCCATAAAACAATACCCCATGTC
ACAAGAAGCCAGACTGGGGATCAAGCCCCACATACAGAGACTGTTGGACCAGGGAATACTGGTAC
CCTGCCAGTCCCCCTGGAACACGCCCCTGCTACCCGTTAAGAAACCAGGGACTAATGATTATAGGC
CTGTCCAGGATCTGAGAGAAGTCAACAAGCGGGTGGAAGACATCCACCCCACCGTGCCCAACCCT
TACAACCTCTTGAGCGGGCTCCCACCGTCCCACCAGTGGTACACTGTGCTTGATTTAAAGGATGCC
TTTTTCTGCCTGAGACTCCACCCCACCAGTCAGCCTCTCTTCGCCTTTGAGTGGAGAGATCCAGAG
ATGGGAATCTCAGGACAATTGACCTGGACCAGACTCCCACAGGGTTTCAAAAACAGTCCCACCCT
GTTTAATGAGGCACTGCACAGAGACCTAGCAGACTTCCGGATCCAGCACCCAGACTTGATCCTGCT
ACAGTACGTGGATGACTTACTGCTGGCCGCCACTTCTGAGCTAGACTGCCAACAAGGTACTCGGGC
CCTGTTACAAACCCTAGGGAACCTCGGGTATCGGGCCTCGGCCAAGAAAGCCCAAATTTGCCAGA
AACAGGTCAAGTATCTGGGGTATCTTCTAAAAGAGGGTCAGAGATGGCTGACTGAGGCCAGAAAA
GAGACTGTGATGGGGCAGCCTACTCCGAAGACCCCTCGACAACTAAGGGAGTTCCTAGGGAAGGC
AGGCTTCTGTCGCCTCTTCATCCCTGGGTTTGCAGAAATGGCAGCCCCCCTGTACCCTCTCACCAAA
CCGGGGACTCTGTTTAATTGGGGCCCAGACCAACAAAAGGCCTATCAAGAAATCAAGCAAGCTCT
TCTAACTGCCCCAGCCCTGGGGTTGCCAGATTTGACTAAGCCCTTTGAACTCTTTGTCGACGAGAA
GCAGGGCTACGCCAAAGGTGTCCTAACGCAAAAACTGGGACCTTGGCGTCGGCCGGTGGCCTACC
TGTCCAAAAAGCTAGACCCAGTAGCAGCTGGGTGGCCCCCTTGCCTACGGATGGTAGCAGCCATT
GCCGTACTGACAAAGGATGCAGGCAAGCTAACCATGGGACAGCCACTAGTCATTCTGGCCCCCCA
TGCAGTAGAGGCACTAGTCAAACAACCCCCCGACCGCTGGCTTTCCAACGCCCGGATGACTCACTA
TCAGGCCTTGCTTTTGGACACGGACCGGGTCCAGTTCGGACCGGTGGTAGCCCTGAACCCGGCTAC
GCTGCTCCCACTGCCTGAGGAAGGGCTGCAACACAACTGCCTTGATATCCTGGCCGAAGCCCACG
GAACCCGACCCGACCTAACGGACCAGCCGCTCCCAGACGCCGACCACACCTGGTACACGGATGGA
AGCAGTCTCTTACAAGAGGGACAGCGTAAGGCGGGAGCTGCGGTGACCACCGAGACCGAGGTAAT
CTGGGCTAAAGCCCTGCCAGCCGGGACATCCGCTCAGCGGGCTGAACTGATAGCACTCACCCAGG
CCCTAAAGATGGCAGAAGGTAAGAAGCTAAATGTTTATACTGATAGCCGTTATGCTTTTGCTACTG
CCCATATCCATGGAGAAATATACAGAAGGCGTGGGTGGCTCACATCAGAAGGCAAAGAGATCAAA
AATAAAGACGAGATCTTGGCCCTACTAAAAGCCCTCTTTCTGCCCAAAAGACTTAGCATAATCCAT
TGTCCAGGACATCAAAAGGGACACAGCGCCGAGGCTAGAGGCAACCGGATGGCTGACCAAGCGG
CCCGAAAGGCAGCCATCACAGAGACTCCAGACACCTCTACCCTCCTCATAGAAAATTCATCACCCT
CTGGCGGCTCAAAAAGAACCGCCGACGGCAGCGAATTCGAGCCCAAGAAGAAGAGGAAAGTCTA A
SEQ ID NO: 17
ATGAAACGGACAGCCGACGGAAGCGAGTTCGAGTCACCAAAGAAGAAGCGGAAAGTCGACAAGA
AGTACAGCATCGGCCTGGACATCGGCACCAACTCTGTGGGCTGGGCCGTGATCACCGACGAGTAC
AAGGTGCCCAGCAAGAAATTCAAGGTGCTGGGCAACACCGACCGGCACAGCATCAAGAAGAACC
TGATCGGAGCCCTGCTGTTCGACAGCGGCGAAACAGCCGAGGCCACCCGGCTGAAGAGAACCGCC
AGAAGAAGATACACCAGACGGAAGAACCGGATCTGCTATCTGCAAGAGATCTTCAGCAACGAGAT
GGCCAAGGTGGACGACAGCTTCTTCCACAGACTGGAAGAGTCCTTCCTGGTGGAAGAGGATAAGA
AGCACGAGCGGCACCCCATCTTCGGCAACATCGTGGACGAGGTGGCCTACCACGAGAAGTACCCC
ACCATCTACCACCTGAGAAAGAAACTGGTGGACAGCACCGACAAGGCCGACCTGCGGCTGATCTA
TCTGGCCCTGGCCCACATGATCAAGTTCCGGGGCCACTTCCTGATCGAGGGCGACCTGAACCCCGA
CAACAGCGACGTGGACAAGCTGTTCATCCAGCTGGTGCAGACCTACAACCAGCTGTTCGAGGAAA
ACCCCATCAACGCCAGCGGCGTGGACGCCAAGGCCATCCTGTCTGCCAGACTGAGCAAGAGCAGA
AAGCTGGAAAATCTGATCGCCCAGCTGCCCGGCGAGAAGAAGAATGGCCTGTTCGGAAACCTGAT
TGCCCTGAGCCTGGGCCTGACCCCCAACTTCAAGAGCAACTTCGACCTGGCCGAGGATGCCAAACT
GCAGCTGAGCAAGGACACCTACGACGACGACCTGGACAACCTGCTGGCCCAGATCGGCGACCAGT
ACGCCGACCTGTTTCTGGCCGCCAAGAACCTGTCCGACGCCATCCTGCTGAGCGACATCCTGAGAG
TGAACACCGAGATCACCAAGGCCCCCCTGAGCGCCTCTATGATCAAGAGATACGACGAGCACCAC
CAGGACCTGACCCTGCTGAAAGCTCTCGTGCGGCAGCAGCTGCCTGAGAAGTACAAAGAGATTTT
CTTCGACCAGAGCAAGAACGGCTACGCCGGCTACATTGACGGCGGAGCCAGCCAGGAAGAGTTCT
ACAAGTTCATCAAGCCCATCCTGGAAAAGATGGACGGCACCGAGGAACTGCTCGTGAAGCTGAAG
AGAGAGGACCTGCTGCGGAAGCAGCGGACCTTCGACAACGGCAGCATCCCCCACCAGATCCACCT
GGGAGAGCTGCACGCCATTCTGCGGCGGCAGGAAGATTTTTACCCATTCCTGAAGGACAACCGGG
AAAAGATCGAGAAGATCCTGACCTTCCGCATCCCCTACTACGTGGGCCCTCTGGCCAGGGGAAAC
AGCAGATTCGCCTGGATGACCAGAAAGAGCGAGGAAACCATCACCCCCTGGAACTTCGAGGAAGT
GGTGGACAAGGGCGCTTCCGCCCAGAGCTTCATCGAGCGGATGACCAACTTCGATAAGAACCTGC
CCAACGAGAAGGTGCTGCCCAAGCACAGCCTGCTGTACGAGTACTTCACCGTGTATAACGAGCTG
ACCAAAGTGAAATACGTGACCGAGGGAATGAGAAAGCCCGCCTTCCTGAGCGGCGAGCAGAAAA
AGGCCATCGTGGACCTGCTGTTCAAGACCAACCGGAAAGTGACCGTGAAGCAGCTGAAAGAGGAC
TACTTCAAGAAAATCGAGTGCTTCGACTCCGTGGAAATCTCCGGCGTGGAAGATCGGTTCAACGCC
TCCCTGGGCACATACCACGATCTGCTGAAAATTATCAAGGACAAGGACTTCCTGGACAATGAGGA
AAACGAGGACATTCTGGAAGATATCGTGCTGACCCTGACACTGTTTGAGGACAGAGAGATGATCG
AGGAACGGCTGAAAACCTATGCCCACCTGTTCGACGACAAAGTGATGAAGCAGCTGAAGCGGCGG
AGATACACCGGCTGGGGCAGGCTGAGCCGGAAGCTGATCAACGGCATCCGGGACAAGCAGTCCG
GCAAGACAATCCTGGATTTCCTGAAGTCCGACGGCTTCGCCAACAGAAACTTCATGCAGCTGATCC
ACGACGACAGCCTGACCTTTAAAGAGGACATCCAGAAAGCCCAGGTGTCCGGCCAGGGCGATAGC
CTGCACGAGCACATTGCCAATCTGGCCGGCAGCCCCGCCATTAAGAAGGGCATCCTGCAGACAGT
GAAGGTGGTGGACGAGCTCGTGAAAGTGATGGGCCGGCACAAGCCCGAGAACATCGTGATCGAA
ATGGCCAGAGAGAACCAGACCACCCAGAAGGGACAGAAGAACAGCCGCGAGAGAATGAAGCGG
ATCGAAGAGGGCATCAAAGAGCTGGGCAGCCAGATCCTGAAAGAACACCCCGTGGAAAACACCC
AGCTGCAGAACGAGAAGCTGTACCTGTACTACCTGCAGAATGGGCGGGATATGTACGTGGACCAG
GAACTGGACATCAACCGGCTGTCCGACTACGATGTGGACGCTATCGTGCCTCAGAGCTTTCTGAAG
GACGACTCCATCGACAACAAGGTGCTGACCAGAAGCGACAAGAACCGGGGCAAGAGCGACAACG
TGCCCTCCGAAGAGGTCGTGAAGAAGATGAAGAACTACTGGCGGCAGCTGCTGAACGCCAAGCTG
ATTACCCAGAGAAAGTTCGACAATCTGACCAAGGCCGAGAGAGGCGGCCTGAGCGAACTGGATAA
GGCCGGCTTCATCAAGAGACAGCTGGTGGAAACCCGGCAGATCACAAAGCACGTGGCACAGATCC
TGGACTCCCGGATGAACACTAAGTACGACGAGAATGACAAGCTGATCCGGGAAGTGAAAGTGATC
ACCCTGAAGTCCAAGCTGGTGTCCGATTTCCGGAAGGATTTCCAGTTTTACAAAGTGCGCGAGATC
AACAACTACCACCACGCCCACGACGCCTACCTGAACGCCGTCGTGGGAACCGCCCTGATCAAAAA
GTACCCTAAGCTGGAAAGCGAGTTCGTGTACGGCGACTACAAGGTGTACGACGTGCGGAAGATGA
TCGCCAAGAGCGAGCAGGAAATCGGCAAGGCTACCGCCAAGTACTTCTTCTACAGCAACATCATG
AACTTTTTCAAGACCGAGATTACCCTGGCCAACGGCGAGATCCGGAAGCGGCCTCTGATCGAGAC
AAACGGCGAAACCGGGGAGATCGTGTGGGATAAGGGCCGGGATTTTGCCACCGTGCGGAAAGTGC
TGAGCATGCCCCAAGTGAATATCGTGAAAAAGACCGAGGTGCAGACAGGCGGCTTCAGCAAAGA
GTCTATCCTGCCCAAGAGGAACAGCGATAAGCTGATCGCCAGAAAGAAGGACTGGGACCCTAAGA
AGTACGGCGGCTTCGACAGCCCCACCGTGGCCTATTCTGTGCTGGTGGTGGCCAAAGTGGAAAAG
GGCAAGTCCAAGAAACTGAAGAGTGTGAAAGAGCTGCTGGGGATCACCATCATGGAAAGAAGCA
GCTTCGAGAAGAATCCCATCGACTTTCTGGAAGCCAAGGGCTACAAAGAAGTGAAAAAGGACCTG
ATCATCAAGCTGCCTAAGTACTCCCTGTTCGAGCTGGAAAACGGCCGGAAGAGAATGCTGGCCTCT
GCCGGCGAACTGCAGAAGGGAAACGAACTGGCCCTGCCCTCCAAATATGTGAACTTCCTGTACCT
GGCCAGCCACTATGAGAAGCTGAAGGGCTCCCCCGAGGATAATGAGCAGAAACAGCTGTTTGTGG
AACAGCACAAGCACTACCTGGACGAGATCATCGAGCAGATCAGCGAGTTCTCCAAGAGAGTGATC
CTGGCCGACGCTAATCTGGACAAAGTGCTGTCCGCCTACAACAAGCACCGGGATAAGCCCATCAG
AGAGCAGGCCGAGAATATCATCCACCTGTTTACCCTGACCAATCTGGGAGCCCCTGCCGCCTTCAA
GTACTTTGACACCACCATCGACCGGAAGAGGTACACCAGCACCAAAGAGGTGCTGGACGCCACCC
TGATCCACCAGAGCATCACCGGCCTGTACGAGACACGGATCGACCTGTCTCAGCTGGGAGGTGAC
TCCGGCGGAAGCTCTGGTGGCAGCAAGCGGACCGCCGACGGCTCTGAATTCGAGAGCCCTAAGAA
GAAAAGAAAGGTGAGCGGAGGCTCTAGCGGCGGAAGCACCCTGAACATTGAAGACGAGTATAGA
CTGCATGAAACAAGCAAGGAACCCGACGTGTCCCTGGGCTCCACCTGGCTGTCCGACTTTCCCCAG
GCCTGGGCCGAGACAGGAGGAATGGGCCTGGCCGTGCGGCAGGCACCCCTGATCATCCCTCTGAA
GGCCACCTCTACACCCGTGAGCATCAAGCAGTACCCTATGTCTCAGGAGGCCAGACTGGGCATCA
AGCCTCACATCCAGAGGCTGCTGGACCAGGGCATCCTGGTGCCATGCCAGAGCCCCTGGAACACA
CCACTGCTGCCCGTGAAGAAGCCAGGCACCAATGACTATAGACCCGTGCAGGATCTGAGAGAGGT
GAACAAGAGGGTGGAGGATATCCACCCCACCGTGCCCAACCCTTACAATCTGCTGTCCGGCCTGCC
CCCTTCTCACCAGTGGTATACAGTGCTGGACCTGAAGGATGCCTTCTTTTGTCTGAGACTGCACCCT
ACCAGCCAGCCACTGTTCGCCTTTGAGTGGAGGGACCCTGAGATGGGCATCTCTGGCCAGCTGACC
TGGACACGCCTGCCTCAGGGCTTCAAGAATAGCCCAACACTGTTTAACGAGGCCCTGCACCGCGA
CCTGGCAGATTTCCGGATCCAGCACCCAGATCTGATCCTGCTGCAGTACGTGGACGATCTGCTGCT
GGCCGCCACCAGCGAGCTGGATTGCCAGCAGGGAACACGCGCCCTGCTGCAGACCCTGGGAAACC
TGGGATATAGGGCATCCGCCAAGAAGGCCCAGATCTGTCAGAAGCAGGTGAAGTACCTGGGCTAT
CTGCTGAAGGAGGGCCAGAGATGGCTGACAGAGGCCAGGAAGGAGACAGTGATGGGCCAGCCAA
CACCCAAGACCCCAAGACAGCTGAGGGAGTTCCTGGGCAAAGCAGGATTTTGCAGGCTGTTCATC
CCAGGATTCGCAGAGATGGCAGCACCTCTGTACCCACTGACCAAGCCGGGCACCCTGTTTAATTGG
GGCCCTGACCAGCAGAAGGCCTATCAGGAGATCAAGCAGGCCCTGCTGACAGCACCAGCCCTGGG
CCTGCCAGACCTGACCAAGCCTTTCGAGCTGTTTGTGGATGAGAAGCAGGGCTACGCCAAGGGCG
TGCTGACCCAGAAGCTGGGACCATGGAGACGGCCCGTGGCCTATCTGTCCAAGAAGCTGGACCCA
GTGGCAGCAGGATGGCCACCATGCCTGAGGATGGTGGCAGCAATCGCCGTGCTGACAAAGGATGC
CGGCAAGCTGACCATGGGACAGCCACTGGTCATCCTGGCACCACACGCAGTGGAGGCCCTGGTGA
AGCAGCCTCCAGATCGCTGGCTGTCTAACGCCCGGATGACACACTACCAGGCCCTGCTGCTGGACA
CCGATCGCGTGCAGTTTGGCCCTGTGGTGGCCCTGAATCCAGCCACCCTGCTGCCTCTGCCAGAGG
AGGGCCTGCAGCACAACTGTCTGGACATCCTGGCAGAGGCACACGGAACAAGGCCAGACCTGACC
GATCAGCCCCTGCCTGACGCCGATCACACATGGTATACCGATGGAAGCTCCCTGCTGCAGGAGGG
CCAGAGGAAGGCAGGAGCAGCAGTGACCACAGAGACAGAAGTGATCTGGGCCAAGGCCCTGCCA
GCAGGCACATCCGCCCAGCGGGCCGAGCTGATCGCCCTGACCCAGGCCCTGAAGATGGCCGAGGG
CAAGAAGCTGAACGTGTACACAGACTCCAGATATGCCTTCGCCACCGCACACATCCACGGAGAGA
TCTACAGGCGCCGGGGCTGGCTGACCTCTGAGGGCAAGGAGATCAAGAACAAGGATGAGATCCTG
GCCCTGCTGAAGGCCCTGTTTCTGCCCAAGCGGCTGAGCATCATCCACTGTCCTGGACACCAGAAG
GGACACTCCGCCGAGGCAAGGGGCAATCGGATGGCCGACCAGGCCGCCAGAAAGGCTGCTATTAC
TGAAACTCCCGACACTTCCACTCTGCTGATTGAAAACTCCTCCCCTTCCGGAGGATCTAGCGGAGG
CTCCTCTGGCTCTGAGACACCTGGCACAAGCGAGAGCGCAACACCTGAAAGCAGCGGGGGCAGCA
GCGGGGGGTCAATGGCTGAAAATGGTGATAATGAAAAGATGGCTGCCCTGGAGGCCAAAATCTGT
CATCAAATTGAGTATTATTTTGGCGACTTCAATTTGCCACGGGACAAGTTTCTAAAGGAACAGATA
AAACTGGATGAAGGCTGGGTACCTTTGGAGATAATGATAAAATTCAACAGGTTGAACCGTCTAAC
AACAGACTTTAATGTAATTGTGGAAGCATTGAGCAAATCCAAGGCAGAACTCATGGAAATCAGTG
AAGATAAAACTAAAATCAGAAGGTCTCCAAGCAAACCCCTACCTGAAGTGACTGATGAGTATAAA
AATGATGTAAAAAACAGATCTGTTTATATTAAAGGCTTCCCAACTGATGCAACTCTTGATGACATA
AAAGAATGGTTAGAAGATAAAGGTCAAGTACTAAATATTCAGATGAGAAGAACATTGCATAAAGC
ATTTAAGGGATCAATTTTTGTTGTGTTTGATAGCATTGAATCTGCTAAGAAATTTGTAGAGACCCCT
GGCCAGAAGTACAAAGAAACAGACCTGCTAATACTTTTCAAGGACGATTACTTTGCCAAAAAAAA TGAATCTGGCGGCTCAAAAAGAACCGCCGACGGCAGCGAATTCGAGTCTCCCAAGAAGAAGAGG AAAGTCGGCTCTGGCCCTGCCGCTAAGAGAGTGAAGCTGGACTAA
SEQ ID NO: 18
ATGAAACGGACAGCCGACGGAAGCGAGTTCGAGTCACCAAAGAAGAAGCGGAAAGTCTCTGGCG
GCTCAGCTGAAAATGGTGATAATGAAAAGATGGCTGCCCTGGAGGCCAAAATCTGTCATCAAATT
GAGTATTATTTTGGCGACTTCAATTTGCCACGGGACAAGTTTCTAAAGGAACAGATAAAACTGGAT
GAAGGCTGGGTACCTTTGGAGATAATGATAAAATTCAACAGGTTGAACCGTCTAACAACAGACTT
TAATGTAATTGTGGAAGCATTGAGCAAATCCAAGGCAGAACTCATGGAAATCAGTGAAGATAAAA
CTAAAATCAGAAGGTCTCCAAGCAAACCCCTACCTGAAGTGACTGATGAGTATAAAAATGATGTA
AAAAACAGATCTGTTTATATTAAAGGCTTCCCAACTGATGCAACTCTTGATGACATAAAAGAATGG
TTAGAAGATAAAGGTCAAGTACTAAATATTCAGATGAGAAGAACATTGCATAAAGCATTTAAGGG
ATCAATTTTTGTTGTGTTTGATAGCATTGAATCTGCTAAGAAATTTGTAGAGACCCCTGGCCAGAA
GTACAAAGAAACAGACCTGCTAATACTTTTCAAGGACGATTACTTTGCCAAAAAAAATGAATCCG
GAGGATCTAGCGGAGGCTCCTCTGGCTCTGAGACACCTGGCACAAGCGAGAGCGCAACACCTGAA
AGCAGCGGGGGCAGCAGCGGGGGGTCAGACAAGAAGTACAGCATCGGCCTGGACATCGGCACCA
ACTCTGTGGGCTGGGCCGTGATCACCGACGAGTACAAGGTGCCCAGCAAGAAATTCAAGGTGCTG
GGCAACACCGACCGGCACAGCATCAAGAAGAACCTGATCGGAGCCCTGCTGTTCGACAGCGGCGA
AACAGCCGAGGCCACCCGGCTGAAGAGAACCGCCAGAAGAAGATACACCAGACGGAAGAACCGG
ATCTGCTATCTGCAAGAGATCTTCAGCAACGAGATGGCCAAGGTGGACGACAGCTTCTTCCACAG
ACTGGAAGAGTCCTTCCTGGTGGAAGAGGATAAGAAGCACGAGCGGCACCCCATCTTCGGCAACA
TCGTGGACGAGGTGGCCTACCACGAGAAGTACCCCACCATCTACCACCTGAGAAAGAAACTGGTG
GACAGCACCGACAAGGCCGACCTGCGGCTGATCTATCTGGCCCTGGCCCACATGATCAAGTTCCG
GGGCCACTTCCTGATCGAGGGCGACCTGAACCCCGACAACAGCGACGTGGACAAGCTGTTCATCC
AGCTGGTGCAGACCTACAACCAGCTGTTCGAGGAAAACCCCATCAACGCCAGCGGCGTGGACGCC
AAGGCCATCCTGTCTGCCAGACTGAGCAAGAGCAGAAAGCTGGAAAATCTGATCGCCCAGCTGCC
CGGCGAGAAGAAGAATGGCCTGTTCGGAAACCTGATTGCCCTGAGCCTGGGCCTGACCCCCAACT
TCAAGAGCAACTTCGACCTGGCCGAGGATGCCAAACTGCAGCTGAGCAAGGACACCTACGACGAC
GACCTGGACAACCTGCTGGCCCAGATCGGCGACCAGTACGCCGACCTGTTTCTGGCCGCCAAGAA
CCTGTCCGACGCCATCCTGCTGAGCGACATCCTGAGAGTGAACACCGAGATCACCAAGGCCCCCCT
GAGCGCCTCTATGATCAAGAGATACGACGAGCACCACCAGGACCTGACCCTGCTGAAAGCTCTCG
TGCGGCAGCAGCTGCCTGAGAAGTACAAAGAGATTTTCTTCGACCAGAGCAAGAACGGCTACGCC
GGCTACATTGACGGCGGAGCCAGCCAGGAAGAGTTCTACAAGTTCATCAAGCCCATCCTGGAAAA
GATGGACGGCACCGAGGAACTGCTCGTGAAGCTGAAGAGAGAGGACCTGCTGCGGAAGCAGCGG
ACCTTCGACAACGGCAGCATCCCCCACCAGATCCACCTGGGAGAGCTGCACGCCATTCTGCGGCG
GCAGGAAGATTTTTACCCATTCCTGAAGGACAACCGGGAAAAGATCGAGAAGATCCTGACCTTCC
GCATCCCCTACTACGTGGGCCCTCTGGCCAGGGGAAACAGCAGATTCGCCTGGATGACCAGAAAG
AGCGAGGAAACCATCACCCCCTGGAACTTCGAGGAAGTGGTGGACAAGGGCGCTTCCGCCCAGAG
CTTCATCGAGCGGATGACCAACTTCGATAAGAACCTGCCCAACGAGAAGGTGCTGCCCAAGCACA
GCCTGCTGTACGAGTACTTCACCGTGTATAACGAGCTGACCAAAGTGAAATACGTGACCGAGGGA
ATGAGAAAGCCCGCCTTCCTGAGCGGCGAGCAGAAAAAGGCCATCGTGGACCTGCTGTTCAAGAC
CAACCGGAAAGTGACCGTGAAGCAGCTGAAAGAGGACTACTTCAAGAAAATCGAGTGCTTCGACT
CCGTGGAAATCTCCGGCGTGGAAGATCGGTTCAACGCCTCCCTGGGCACATACCACGATCTGCTGA
AAATTATCAAGGACAAGGACTTCCTGGACAATGAGGAAAACGAGGACATTCTGGAAGATATCGTG
CTGACCCTGACACTGTTTGAGGACAGAGAGATGATCGAGGAACGGCTGAAAACCTATGCCCACCT
GTTCGACGACAAAGTGATGAAGCAGCTGAAGCGGCGGAGATACACCGGCTGGGGCAGGCTGAGC
CGGAAGCTGATCAACGGCATCCGGGACAAGCAGTCCGGCAAGACAATCCTGGATTTCCTGAAGTC
CGACGGCTTCGCCAACAGAAACTTCATGCAGCTGATCCACGACGACAGCCTGACCTTTAAAGAGG
ACATCCAGAAAGCCCAGGTGTCCGGCCAGGGCGATAGCCTGCACGAGCACATTGCCAATCTGGCC
GGCAGCCCCGCCATTAAGAAGGGCATCCTGCAGACAGTGAAGGTGGTGGACGAGCTCGTGAAAGT
GATGGGCCGGCACAAGCCCGAGAACATCGTGATCGAAATGGCCAGAGAGAACCAGACCACCCAG
AAGGGACAGAAGAACAGCCGCGAGAGAATGAAGCGGATCGAAGAGGGCATCAAAGAGCTGGGC
AGCCAGATCCTGAAAGAACACCCCGTGGAAAACACCCAGCTGCAGAACGAGAAGCTGTACCTGTA
CTACCTGCAGAATGGGCGGGATATGTACGTGGACCAGGAACTGGACATCAACCGGCTGTCCGACT
ACGATGTGGACGCTATCGTGCCTCAGAGCTTTCTGAAGGACGACTCCATCGACAACAAGGTGCTG
ACCAGAAGCGACAAGAACCGGGGCAAGAGCGACAACGTGCCCTCCGAAGAGGTCGTGAAGAAGA
TGAAGAACTACTGGCGGCAGCTGCTGAACGCCAAGCTGATTACCCAGAGAAAGTTCGACAATCTG
ACCAAGGCCGAGAGAGGCGGCCTGAGCGAACTGGATAAGGCCGGCTTCATCAAGAGACAGCTGG
TGGAAACCCGGCAGATCACAAAGCACGTGGCACAGATCCTGGACTCCCGGATGAACACTAAGTAC
GACGAGAATGACAAGCTGATCCGGGAAGTGAAAGTGATCACCCTGAAGTCCAAGCTGGTGTCCGA
TTTCCGGAAGGATTTCCAGTTTTACAAAGTGCGCGAGATCAACAACTACCACCACGCCCACGACGC
CTACCTGAACGCCGTCGTGGGAACCGCCCTGATCAAAAAGTACCCTAAGCTGGAAAGCGAGTTCG
TGTACGGCGACTACAAGGTGTACGACGTGCGGAAGATGATCGCCAAGAGCGAGCAGGAAATCGG
CAAGGCTACCGCCAAGTACTTCTTCTACAGCAACATCATGAACTTTTTCAAGACCGAGATTACCCT
GGCCAACGGCGAGATCCGGAAGCGGCCTCTGATCGAGACAAACGGCGAAACCGGGGAGATCGTG
TGGGATAAGGGCCGGGATTTTGCCACCGTGCGGAAAGTGCTGAGCATGCCCCAAGTGAATATCGT
GAAAAAGACCGAGGTGCAGACAGGCGGCTTCAGCAAAGAGTCTATCCTGCCCAAGAGGAACAGC
GATAAGCTGATCGCCAGAAAGAAGGACTGGGACCCTAAGAAGTACGGCGGCTTCGACAGCCCCAC
CGTGGCCTATTCTGTGCTGGTGGTGGCCAAAGTGGAAAAGGGCAAGTCCAAGAAACTGAAGAGTG
TGAAAGAGCTGCTGGGGATCACCATCATGGAAAGAAGCAGCTTCGAGAAGAATCCCATCGACTTT
CTGGAAGCCAAGGGCTACAAAGAAGTGAAAAAGGACCTGATCATCAAGCTGCCTAAGTACTCCCT
GTTCGAGCTGGAAAACGGCCGGAAGAGAATGCTGGCCTCTGCCGGCGAACTGCAGAAGGGAAAC
GAACTGGCCCTGCCCTCCAAATATGTGAACTTCCTGTACCTGGCCAGCCACTATGAGAAGCTGAAG
GGCTCCCCCGAGGATAATGAGCAGAAACAGCTGTTTGTGGAACAGCACAAGCACTACCTGGACGA
GATCATCGAGCAGATCAGCGAGTTCTCCAAGAGAGTGATCCTGGCCGACGCTAATCTGGACAAAG
TGCTGTCCGCCTACAACAAGCACCGGGATAAGCCCATCAGAGAGCAGGCCGAGAATATCATCCAC
CTGTTTACCCTGACCAATCTGGGAGCCCCTGCCGCCTTCAAGTACTTTGACACCACCATCGACCGG
AAGAGGTACACCAGCACCAAAGAGGTGCTGGACGCCACCCTGATCCACCAGAGCATCACCGGCCT
GTACGAGACACGGATCGACCTGTCTCAGCTGGGAGGTGACTCCGGCGGAAGCTCTGGTGGCAGCA
AGCGGACCGCCGACGGCTCTGAATTCGAGAGCCCTAAGAAGAAAAGAAAGGTGAGCGGAGGCTC
TAGCGGCGGAAGCACCCTGAACATTGAAGACGAGTATAGACTGCATGAAACAAGCAAGGAACCC
GACGTGTCCCTGGGCTCCACCTGGCTGTCCGACTTTCCCCAGGCCTGGGCCGAGACAGGAGGAATG
GGCCTGGCCGTGCGGCAGGCACCCCTGATCATCCCTCTGAAGGCCACCTCTACACCCGTGAGCATC
AAGCAGTACCCTATGTCTCAGGAGGCCAGACTGGGCATCAAGCCTCACATCCAGAGGCTGCTGGA
CCAGGGCATCCTGGTGCCATGCCAGAGCCCCTGGAACACACCACTGCTGCCCGTGAAGAAGCCAG
GCACCAATGACTATAGACCCGTGCAGGATCTGAGAGAGGTGAACAAGAGGGTGGAGGATATCCAC
CCCACCGTGCCCAACCCTTACAATCTGCTGTCCGGCCTGCCCCCTTCTCACCAGTGGTATACAGTGC
TGGACCTGAAGGATGCCTTCTTTTGTCTGAGACTGCACCCTACCAGCCAGCCACTGTTCGCCTTTGA
GTGGAGGGACCCTGAGATGGGCATCTCTGGCCAGCTGACCTGGACACGCCTGCCTCAGGGCTTCA
AGAATAGCCCAACACTGTTTAACGAGGCCCTGCACCGCGACCTGGCAGATTTCCGGATCCAGCAC
CCAGATCTGATCCTGCTGCAGTACGTGGACGATCTGCTGCTGGCCGCCACCAGCGAGCTGGATTGC
CAGCAGGGAACACGCGCCCTGCTGCAGACCCTGGGAAACCTGGGATATAGGGCATCCGCCAAGAA
GGCCCAGATCTGTCAGAAGCAGGTGAAGTACCTGGGCTATCTGCTGAAGGAGGGCCAGAGATGGC
TGACAGAGGCCAGGAAGGAGACAGTGATGGGCCAGCCAACACCCAAGACCCCAAGACAGCTGAG
GGAGTTCCTGGGCAAAGCAGGATTTTGCAGGCTGTTCATCCCAGGATTCGCAGAGATGGCAGCAC
CTCTGTACCCACTGACCAAGCCGGGCACCCTGTTTAATTGGGGCCCTGACCAGCAGAAGGCCTATC
AGGAGATCAAGCAGGCCCTGCTGACAGCACCAGCCCTGGGCCTGCCAGACCTGACCAAGCCTTTC
GAGCTGTTTGTGGATGAGAAGCAGGGCTACGCCAAGGGCGTGCTGACCCAGAAGCTGGGACCATG
GAGACGGCCCGTGGCCTATCTGTCCAAGAAGCTGGACCCAGTGGCAGCAGGATGGCCACCATGCC
TGAGGATGGTGGCAGCAATCGCCGTGCTGACAAAGGATGCCGGCAAGCTGACCATGGGACAGCCA
CTGGTCATCCTGGCACCACACGCAGTGGAGGCCCTGGTGAAGCAGCCTCCAGATCGCTGGCTGTCT
AACGCCCGGATGACACACTACCAGGCCCTGCTGCTGGACACCGATCGCGTGCAGTTTGGCCCTGTG
GTGGCCCTGAATCCAGCCACCCTGCTGCCTCTGCCAGAGGAGGGCCTGCAGCACAACTGTCTGGAC
ATCCTGGCAGAGGCACACGGAACAAGGCCAGACCTGACCGATCAGCCCCTGCCTGACGCCGATCA
CACATGGTATACCGATGGAAGCTCCCTGCTGCAGGAGGGCCAGAGGAAGGCAGGAGCAGCAGTG
ACCACAGAGACAGAAGTGATCTGGGCCAAGGCCCTGCCAGCAGGCACATCCGCCCAGCGGGCCGA
GCTGATCGCCCTGACCCAGGCCCTGAAGATGGCCGAGGGCAAGAAGCTGAACGTGTACACAGACT
CCAGATATGCCTTCGCCACCGCACACATCCACGGAGAGATCTACAGGCGCCGGGGCTGGCTGACC
TCTGAGGGCAAGGAGATCAAGAACAAGGATGAGATCCTGGCCCTGCTGAAGGCCCTGTTTCTGCC
CAAGCGGCTGAGCATCATCCACTGTCCTGGACACCAGAAGGGACACTCCGCCGAGGCAAGGGGCA
ATCGGATGGCCGACCAGGCCGCCAGAAAGGCTGCTATTACTGAAACTCCCGACACTTCCACTCTGC
TGATTGAAAACTCCTCCCCTTCTGGCGGCTCAAAAAGAACCGCCGACGGCAGCGAATTCGAGTCTC
CCAAGAAGAAGAGGAAAGTCGGCTCTGGCCCTGCCGCTAAGAGAGTGAAGCTGGACTAA
SEQ ID NO: 19
ATGAAACGGACAGCCGACGGAAGCGAGTTCGAGTCACCAAAGAAGAAGCGGAAAGTCGACAAGA
AGTACAGCATCGGCCTGGACATCGGCACCAACTCTGTGGGCTGGGCCGTGATCACCGACGAGTAC
AAGGTGCCCAGCAAGAAATTCAAGGTGCTGGGCAACACCGACCGGCACAGCATCAAGAAGAACC
TGATCGGAGCCCTGCTGTTCGACAGCGGCGAAACAGCCGAGGCCACCCGGCTGAAGAGAACCGCC
AGAAGAAGATACACCAGACGGAAGAACCGGATCTGCTATCTGCAAGAGATCTTCAGCAACGAGAT
GGCCAAGGTGGACGACAGCTTCTTCCACAGACTGGAAGAGTCCTTCCTGGTGGAAGAGGATAAGA
AGCACGAGCGGCACCCCATCTTCGGCAACATCGTGGACGAGGTGGCCTACCACGAGAAGTACCCC
ACCATCTACCACCTGAGAAAGAAACTGGTGGACAGCACCGACAAGGCCGACCTGCGGCTGATCTA
TCTGGCCCTGGCCCACATGATCAAGTTCCGGGGCCACTTCCTGATCGAGGGCGACCTGAACCCCGA
CAACAGCGACGTGGACAAGCTGTTCATCCAGCTGGTGCAGACCTACAACCAGCTGTTCGAGGAAA
ACCCCATCAACGCCAGCGGCGTGGACGCCAAGGCCATCCTGTCTGCCAGACTGAGCAAGAGCAGA
AAGCTGGAAAATCTGATCGCCCAGCTGCCCGGCGAGAAGAAGAATGGCCTGTTCGGAAACCTGAT
TGCCCTGAGCCTGGGCCTGACCCCCAACTTCAAGAGCAACTTCGACCTGGCCGAGGATGCCAAACT
GCAGCTGAGCAAGGACACCTACGACGACGACCTGGACAACCTGCTGGCCCAGATCGGCGACCAGT
ACGCCGACCTGTTTCTGGCCGCCAAGAACCTGTCCGACGCCATCCTGCTGAGCGACATCCTGAGAG
TGAACACCGAGATCACCAAGGCCCCCCTGAGCGCCTCTATGATCAAGAGATACGACGAGCACCAC
CAGGACCTGACCCTGCTGAAAGCTCTCGTGCGGCAGCAGCTGCCTGAGAAGTACAAAGAGATTTT
CTTCGACCAGAGCAAGAACGGCTACGCCGGCTACATTGACGGCGGAGCCAGCCAGGAAGAGTTCT
ACAAGTTCATCAAGCCCATCCTGGAAAAGATGGACGGCACCGAGGAACTGCTCGTGAAGCTGAAG
AGAGAGGACCTGCTGCGGAAGCAGCGGACCTTCGACAACGGCAGCATCCCCCACCAGATCCACCT
GGGAGAGCTGCACGCCATTCTGCGGCGGCAGGAAGATTTTTACCCATTCCTGAAGGACAACCGGG
AAAAGATCGAGAAGATCCTGACCTTCCGCATCCCCTACTACGTGGGCCCTCTGGCCAGGGGAAAC
AGCAGATTCGCCTGGATGACCAGAAAGAGCGAGGAAACCATCACCCCCTGGAACTTCGAGGAAGT
GGTGGACAAGGGCGCTTCCGCCCAGAGCTTCATCGAGCGGATGACCAACTTCGATAAGAACCTGC
CCAACGAGAAGGTGCTGCCCAAGCACAGCCTGCTGTACGAGTACTTCACCGTGTATAACGAGCTG
ACCAAAGTGAAATACGTGACCGAGGGAATGAGAAAGCCCGCCTTCCTGAGCGGCGAGCAGAAAA
AGGCCATCGTGGACCTGCTGTTCAAGACCAACCGGAAAGTGACCGTGAAGCAGCTGAAAGAGGAC
TACTTCAAGAAAATCGAGTGCTTCGACTCCGTGGAAATCTCCGGCGTGGAAGATCGGTTCAACGCC
TCCCTGGGCACATACCACGATCTGCTGAAAATTATCAAGGACAAGGACTTCCTGGACAATGAGGA
AAACGAGGACATTCTGGAAGATATCGTGCTGACCCTGACACTGTTTGAGGACAGAGAGATGATCG
AGGAACGGCTGAAAACCTATGCCCACCTGTTCGACGACAAAGTGATGAAGCAGCTGAAGCGGCGG
AGATACACCGGCTGGGGCAGGCTGAGCCGGAAGCTGATCAACGGCATCCGGGACAAGCAGTCCG
GCAAGACAATCCTGGATTTCCTGAAGTCCGACGGCTTCGCCAACAGAAACTTCATGCAGCTGATCC
ACGACGACAGCCTGACCTTTAAAGAGGACATCCAGAAAGCCCAGGTGTCCGGCCAGGGCGATAGC
CTGCACGAGCACATTGCCAATCTGGCCGGCAGCCCCGCCATTAAGAAGGGCATCCTGCAGACAGT
GAAGGTGGTGGACGAGCTCGTGAAAGTGATGGGCCGGCACAAGCCCGAGAACATCGTGATCGAA
ATGGCCAGAGAGAACCAGACCACCCAGAAGGGACAGAAGAACAGCCGCGAGAGAATGAAGCGG
ATCGAAGAGGGCATCAAAGAGCTGGGCAGCCAGATCCTGAAAGAACACCCCGTGGAAAACACCC
AGCTGCAGAACGAGAAGCTGTACCTGTACTACCTGCAGAATGGGCGGGATATGTACGTGGACCAG
GAACTGGACATCAACCGGCTGTCCGACTACGATGTGGACGCTATCGTGCCTCAGAGCTTTCTGAAG
GACGACTCCATCGACAACAAGGTGCTGACCAGAAGCGACAAGAACCGGGGCAAGAGCGACAACG
TGCCCTCCGAAGAGGTCGTGAAGAAGATGAAGAACTACTGGCGGCAGCTGCTGAACGCCAAGCTG
ATTACCCAGAGAAAGTTCGACAATCTGACCAAGGCCGAGAGAGGCGGCCTGAGCGAACTGGATAA
GGCCGGCTTCATCAAGAGACAGCTGGTGGAAACCCGGCAGATCACAAAGCACGTGGCACAGATCC
TGGACTCCCGGATGAACACTAAGTACGACGAGAATGACAAGCTGATCCGGGAAGTGAAAGTGATC
ACCCTGAAGTCCAAGCTGGTGTCCGATTTCCGGAAGGATTTCCAGTTTTACAAAGTGCGCGAGATC
AACAACTACCACCACGCCCACGACGCCTACCTGAACGCCGTCGTGGGAACCGCCCTGATCAAAAA
GTACCCTAAGCTGGAAAGCGAGTTCGTGTACGGCGACTACAAGGTGTACGACGTGCGGAAGATGA
TCGCCAAGAGCGAGCAGGAAATCGGCAAGGCTACCGCCAAGTACTTCTTCTACAGCAACATCATG
AACTTTTTCAAGACCGAGATTACCCTGGCCAACGGCGAGATCCGGAAGCGGCCTCTGATCGAGAC
AAACGGCGAAACCGGGGAGATCGTGTGGGATAAGGGCCGGGATTTTGCCACCGTGCGGAAAGTGC
TGAGCATGCCCCAAGTGAATATCGTGAAAAAGACCGAGGTGCAGACAGGCGGCTTCAGCAAAGA
GTCTATCCTGCCCAAGAGGAACAGCGATAAGCTGATCGCCAGAAAGAAGGACTGGGACCCTAAGA
AGTACGGCGGCTTCGACAGCCCCACCGTGGCCTATTCTGTGCTGGTGGTGGCCAAAGTGGAAAAG
GGCAAGTCCAAGAAACTGAAGAGTGTGAAAGAGCTGCTGGGGATCACCATCATGGAAAGAAGCA
GCTTCGAGAAGAATCCCATCGACTTTCTGGAAGCCAAGGGCTACAAAGAAGTGAAAAAGGACCTG
ATCATCAAGCTGCCTAAGTACTCCCTGTTCGAGCTGGAAAACGGCCGGAAGAGAATGCTGGCCTCT
GCCGGCGAACTGCAGAAGGGAAACGAACTGGCCCTGCCCTCCAAATATGTGAACTTCCTGTACCT
GGCCAGCCACTATGAGAAGCTGAAGGGCTCCCCCGAGGATAATGAGCAGAAACAGCTGTTTGTGG
AACAGCACAAGCACTACCTGGACGAGATCATCGAGCAGATCAGCGAGTTCTCCAAGAGAGTGATC
CTGGCCGACGCTAATCTGGACAAAGTGCTGTCCGCCTACAACAAGCACCGGGATAAGCCCATCAG
AGAGCAGGCCGAGAATATCATCCACCTGTTTACCCTGACCAATCTGGGAGCCCCTGCCGCCTTCAA
GTACTTTGACACCACCATCGACCGGAAGAGGTACACCAGCACCAAAGAGGTGCTGGACGCCACCC
TGATCCACCAGAGCATCACCGGCCTGTACGAGACACGGATCGACCTGTCTCAGCTGGGAGGTGAC
TCCGGAGGATCTAGCGGAGGCTCCTCTGGCTCTGAGACACCTGGCACAAGCGAGAGCGCAACACC
TGAAAGCAGCGGGGGCAGCAGCGGGGGGTCAATGGCTGAAAATGGTGATAATGAAAAGATGGCT
GCCCTGGAGGCCAAAATCTGTCATCAAATTGAGTATTATTTTGGCGACTTCAATTTGCCACGGGAC
AAGTTTCTAAAGGAACAGATAAAACTGGATGAAGGCTGGGTACCTTTGGAGATAATGATAAAATT
CAACAGGTTGAACCGTCTAACAACAGACTTTAATGTAATTGTGGAAGCATTGAGCAAATCCAAGG
CAGAACTCATGGAAATCAGTGAAGATAAAACTAAAATCAGAAGGTCTCCAAGCAAACCCCTACCT
GAAGTGACTGATGAGTATAAAAATGATGTAAAAAACAGATCTGTTTATATTAAAGGCTTCCCAACT
GATGCAACTCTTGATGACATAAAAGAATGGTTAGAAGATAAAGGTCAAGTACTAAATATTCAGAT
GAGAAGAACATTGCATAAAGCATTTAAGGGATCAATTTTTGTTGTGTTTGATAGCATTGAATCTGC
TAAGAAATTTGTAGAGACCCCTGGCCAGAAGTACAAAGAAACAGACCTGCTAATACTTTTCAAGG
ACGATTACTTTGCCAAAAAAAATGAATCCGGCGGAAGCTCTGGTGGCAGCAAGCGGACCGCCGAC
GGCTCTGAATTCGAGAGCCCTAAGAAGAAAAGAAAGGTGAGCGGAGGCTCTAGCGGCGGAAGCA
CCCTGAACATTGAAGACGAGTATAGACTGCATGAAACAAGCAAGGAACCCGACGTGTCCCTGGGC
TCCACCTGGCTGTCCGACTTTCCCCAGGCCTGGGCCGAGACAGGAGGAATGGGCCTGGCCGTGCG
GCAGGCACCCCTGATCATCCCTCTGAAGGCCACCTCTACACCCGTGAGCATCAAGCAGTACCCTAT
GTCTCAGGAGGCCAGACTGGGCATCAAGCCTCACATCCAGAGGCTGCTGGACCAGGGCATCCTGG
TGCCATGCCAGAGCCCCTGGAACACACCACTGCTGCCCGTGAAGAAGCCAGGCACCAATGACTAT
AGACCCGTGCAGGATCTGAGAGAGGTGAACAAGAGGGTGGAGGATATCCACCCCACCGTGCCCAA
CCCTTACAATCTGCTGTCCGGCCTGCCCCCTTCTCACCAGTGGTATACAGTGCTGGACCTGAAGGA
TGCCTTCTTTTGTCTGAGACTGCACCCTACCAGCCAGCCACTGTTCGCCTTTGAGTGGAGGGACCCT
GAGATGGGCATCTCTGGCCAGCTGACCTGGACACGCCTGCCTCAGGGCTTCAAGAATAGCCCAAC
ACTGTTTAACGAGGCCCTGCACCGCGACCTGGCAGATTTCCGGATCCAGCACCCAGATCTGATCCT
GCTGCAGTACGTGGACGATCTGCTGCTGGCCGCCACCAGCGAGCTGGATTGCCAGCAGGGAACAC
GCGCCCTGCTGCAGACCCTGGGAAACCTGGGATATAGGGCATCCGCCAAGAAGGCCCAGATCTGT
CAGAAGCAGGTGAAGTACCTGGGCTATCTGCTGAAGGAGGGCCAGAGATGGCTGACAGAGGCCA
GGAAGGAGACAGTGATGGGCCAGCCAACACCCAAGACCCCAAGACAGCTGAGGGAGTTCCTGGG
CAAAGCAGGATTTTGCAGGCTGTTCATCCCAGGATTCGCAGAGATGGCAGCACCTCTGTACCCACT
GACCAAGCCGGGCACCCTGTTTAATTGGGGCCCTGACCAGCAGAAGGCCTATCAGGAGATCAAGC
AGGCCCTGCTGACAGCACCAGCCCTGGGCCTGCCAGACCTGACCAAGCCTTTCGAGCTGTTTGTGG
ATGAGAAGCAGGGCTACGCCAAGGGCGTGCTGACCCAGAAGCTGGGACCATGGAGACGGCCCGT
GGCCTATCTGTCCAAGAAGCTGGACCCAGTGGCAGCAGGATGGCCACCATGCCTGAGGATGGTGG
CAGCAATCGCCGTGCTGACAAAGGATGCCGGCAAGCTGACCATGGGACAGCCACTGGTCATCCTG
GCACCACACGCAGTGGAGGCCCTGGTGAAGCAGCCTCCAGATCGCTGGCTGTCTAACGCCCGGAT
GACACACTACCAGGCCCTGCTGCTGGACACCGATCGCGTGCAGTTTGGCCCTGTGGTGGCCCTGAA
TCCAGCCACCCTGCTGCCTCTGCCAGAGGAGGGCCTGCAGCACAACTGTCTGGACATCCTGGCAGA
GGCACACGGAACAAGGCCAGACCTGACCGATCAGCCCCTGCCTGACGCCGATCACACATGGTATA
CCGATGGAAGCTCCCTGCTGCAGGAGGGCCAGAGGAAGGCAGGAGCAGCAGTGACCACAGAGAC
AGAAGTGATCTGGGCCAAGGCCCTGCCAGCAGGCACATCCGCCCAGCGGGCCGAGCTGATCGCCC
TGACCCAGGCCCTGAAGATGGCCGAGGGCAAGAAGCTGAACGTGTACACAGACTCCAGATATGCC
TTCGCCACCGCACACATCCACGGAGAGATCTACAGGCGCCGGGGCTGGCTGACCTCTGAGGGCAA
GGAGATCAAGAACAAGGATGAGATCCTGGCCCTGCTGAAGGCCCTGTTTCTGCCCAAGCGGCTGA
GCATCATCCACTGTCCTGGACACCAGAAGGGACACTCCGCCGAGGCAAGGGGCAATCGGATGGCC
GACCAGGCCGCCAGAAAGGCTGCTATTACTGAAACTCCCGACACTTCCACTCTGCTGATTGAAAAC
TCCTCCCCTTCTGGCGGCTCAAAAAGAACCGCCGACGGCAGCGAATTCGAGTCTCCCAAGAAGAA
GAGGAAAGTCGGCTCTGGCCCTGCCGCTAAGAGAGTGAAGCTGGACTAA
SEQ ID NO: 20
ATGAAACGGACAGCCGACGGAAGCGAGTTCGAGTCACCAAAGAAGAAGCGGAAAGTCGACAAGA
AGTACAGCATCGGCCTGGACATCGGCACCAACTCTGTGGGCTGGGCCGTGATCACCGACGAGTAC
AAGGTGCCCAGCAAGAAATTCAAGGTGCTGGGCAACACCGACCGGCACAGCATCAAGAAGAACC
TGATCGGAGCCCTGCTGTTCGACAGCGGCGAAACAGCCGAGGCCACCCGGCTGAAGAGAACCGCC
AGAAGAAGATACACCAGACGGAAGAACCGGATCTGCTATCTGCAAGAGATCTTCAGCAACGAGAT
GGCCAAGGTGGACGACAGCTTCTTCCACAGACTGGAAGAGTCCTTCCTGGTGGAAGAGGATAAGA
AGCACGAGCGGCACCCCATCTTCGGCAACATCGTGGACGAGGTGGCCTACCACGAGAAGTACCCC
ACCATCTACCACCTGAGAAAGAAACTGGTGGACAGCACCGACAAGGCCGACCTGCGGCTGATCTA
TCTGGCCCTGGCCCACATGATCAAGTTCCGGGGCCACTTCCTGATCGAGGGCGACCTGAACCCCGA
CAACAGCGACGTGGACAAGCTGTTCATCCAGCTGGTGCAGACCTACAACCAGCTGTTCGAGGAAA
ACCCCATCAACGCCAGCGGCGTGGACGCCAAGGCCATCCTGTCTGCCAGACTGAGCAAGAGCAGA
AAGCTGGAAAATCTGATCGCCCAGCTGCCCGGCGAGAAGAAGAATGGCCTGTTCGGAAACCTGAT
TGCCCTGAGCCTGGGCCTGACCCCCAACTTCAAGAGCAACTTCGACCTGGCCGAGGATGCCAAACT
GCAGCTGAGCAAGGACACCTACGACGACGACCTGGACAACCTGCTGGCCCAGATCGGCGACCAGT
ACGCCGACCTGTTTCTGGCCGCCAAGAACCTGTCCGACGCCATCCTGCTGAGCGACATCCTGAGAG
TGAACACCGAGATCACCAAGGCCCCCCTGAGCGCCTCTATGATCAAGAGATACGACGAGCACCAC
CAGGACCTGACCCTGCTGAAAGCTCTCGTGCGGCAGCAGCTGCCTGAGAAGTACAAAGAGATTTT
CTTCGACCAGAGCAAGAACGGCTACGCCGGCTACATTGACGGCGGAGCCAGCCAGGAAGAGTTCT
ACAAGTTCATCAAGCCCATCCTGGAAAAGATGGACGGCACCGAGGAACTGCTCGTGAAGCTGAAG
AGAGAGGACCTGCTGCGGAAGCAGCGGACCTTCGACAACGGCAGCATCCCCCACCAGATCCACCT
GGGAGAGCTGCACGCCATTCTGCGGCGGCAGGAAGATTTTTACCCATTCCTGAAGGACAACCGGG
AAAAGATCGAGAAGATCCTGACCTTCCGCATCCCCTACTACGTGGGCCCTCTGGCCAGGGGAAAC
AGCAGATTCGCCTGGATGACCAGAAAGAGCGAGGAAACCATCACCCCCTGGAACTTCGAGGAAGT
GGTGGACAAGGGCGCTTCCGCCCAGAGCTTCATCGAGCGGATGACCAACTTCGATAAGAACCTGC
CCAACGAGAAGGTGCTGCCCAAGCACAGCCTGCTGTACGAGTACTTCACCGTGTATAACGAGCTG
ACCAAAGTGAAATACGTGACCGAGGGAATGAGAAAGCCCGCCTTCCTGAGCGGCGAGCAGAAAA
AGGCCATCGTGGACCTGCTGTTCAAGACCAACCGGAAAGTGACCGTGAAGCAGCTGAAAGAGGAC
TACTTCAAGAAAATCGAGTGCTTCGACTCCGTGGAAATCTCCGGCGTGGAAGATCGGTTCAACGCC
TCCCTGGGCACATACCACGATCTGCTGAAAATTATCAAGGACAAGGACTTCCTGGACAATGAGGA
AAACGAGGACATTCTGGAAGATATCGTGCTGACCCTGACACTGTTTGAGGACAGAGAGATGATCG
AGGAACGGCTGAAAACCTATGCCCACCTGTTCGACGACAAAGTGATGAAGCAGCTGAAGCGGCGG
AGATACACCGGCTGGGGCAGGCTGAGCCGGAAGCTGATCAACGGCATCCGGGACAAGCAGTCCG
GCAAGACAATCCTGGATTTCCTGAAGTCCGACGGCTTCGCCAACAGAAACTTCATGCAGCTGATCC
ACGACGACAGCCTGACCTTTAAAGAGGACATCCAGAAAGCCCAGGTGTCCGGCCAGGGCGATAGC
CTGCACGAGCACATTGCCAATCTGGCCGGCAGCCCCGCCATTAAGAAGGGCATCCTGCAGACAGT
GAAGGTGGTGGACGAGCTCGTGAAAGTGATGGGCCGGCACAAGCCCGAGAACATCGTGATCGAA
ATGGCCAGAGAGAACCAGACCACCCAGAAGGGACAGAAGAACAGCCGCGAGAGAATGAAGCGG
ATCGAAGAGGGCATCAAAGAGCTGGGCAGCCAGATCCTGAAAGAACACCCCGTGGAAAACACCC
AGCTGCAGAACGAGAAGCTGTACCTGTACTACCTGCAGAATGGGCGGGATATGTACGTGGACCAG
GAACTGGACATCAACCGGCTGTCCGACTACGATGTGGACGCTATCGTGCCTCAGAGCTTTCTGAAG
GACGACTCCATCGACAACAAGGTGCTGACCAGAAGCGACAAGAACCGGGGCAAGAGCGACAACG
TGCCCTCCGAAGAGGTCGTGAAGAAGATGAAGAACTACTGGCGGCAGCTGCTGAACGCCAAGCTG
ATTACCCAGAGAAAGTTCGACAATCTGACCAAGGCCGAGAGAGGCGGCCTGAGCGAACTGGATAA
GGCCGGCTTCATCAAGAGACAGCTGGTGGAAACCCGGCAGATCACAAAGCACGTGGCACAGATCC
TGGACTCCCGGATGAACACTAAGTACGACGAGAATGACAAGCTGATCCGGGAAGTGAAAGTGATC
ACCCTGAAGTCCAAGCTGGTGTCCGATTTCCGGAAGGATTTCCAGTTTTACAAAGTGCGCGAGATC
AACAACTACCACCACGCCCACGACGCCTACCTGAACGCCGTCGTGGGAACCGCCCTGATCAAAAA
GTACCCTAAGCTGGAAAGCGAGTTCGTGTACGGCGACTACAAGGTGTACGACGTGCGGAAGATGA
TCGCCAAGAGCGAGCAGGAAATCGGCAAGGCTACCGCCAAGTACTTCTTCTACAGCAACATCATG
AACTTTTTCAAGACCGAGATTACCCTGGCCAACGGCGAGATCCGGAAGCGGCCTCTGATCGAGAC
AAACGGCGAAACCGGGGAGATCGTGTGGGATAAGGGCCGGGATTTTGCCACCGTGCGGAAAGTGC
TGAGCATGCCCCAAGTGAATATCGTGAAAAAGACCGAGGTGCAGACAGGCGGCTTCAGCAAAGA
GTCTATCCTGCCCAAGAGGAACAGCGATAAGCTGATCGCCAGAAAGAAGGACTGGGACCCTAAGA
AGTACGGCGGCTTCGACAGCCCCACCGTGGCCTATTCTGTGCTGGTGGTGGCCAAAGTGGAAAAG
GGCAAGTCCAAGAAACTGAAGAGTGTGAAAGAGCTGCTGGGGATCACCATCATGGAAAGAAGCA
GCTTCGAGAAGAATCCCATCGACTTTCTGGAAGCCAAGGGCTACAAAGAAGTGAAAAAGGACCTG
ATCATCAAGCTGCCTAAGTACTCCCTGTTCGAGCTGGAAAACGGCCGGAAGAGAATGCTGGCCTCT
GCCGGCGAACTGCAGAAGGGAAACGAACTGGCCCTGCCCTCCAAATATGTGAACTTCCTGTACCT
GGCCAGCCACTATGAGAAGCTGAAGGGCTCCCCCGAGGATAATGAGCAGAAACAGCTGTTTGTGG
AACAGCACAAGCACTACCTGGACGAGATCATCGAGCAGATCAGCGAGTTCTCCAAGAGAGTGATC
CTGGCCGACGCTAATCTGGACAAAGTGCTGTCCGCCTACAACAAGCACCGGGATAAGCCCATCAG
AGAGCAGGCCGAGAATATCATCCACCTGTTTACCCTGACCAATCTGGGAGCCCCTGCCGCCTTCAA
GTACTTTGACACCACCATCGACCGGAAGAGGTACACCAGCACCAAAGAGGTGCTGGACGCCACCC
TGATCCACCAGAGCATCACCGGCCTGTACGAGACACGGATCGACCTGTCTCAGCTGGGAGGTGAC
TCCGGCGGAAGCTCTGGTGGCAGCAAGCGGACCGCCGACGGCTCTGAATTCGAGAGCCCTAAGAA
GAAAAGAAAGGTGAGCGGAGGCTCTAGCGGCGGAAGCATGGCTGAAAATGGTGATAATGAAAAG
ATGGCTGCCCTGGAGGCCAAAATCTGTCATCAAATTGAGTATTATTTTGGCGACTTCAATTTGCCA
CGGGACAAGTTTCTAAAGGAACAGATAAAACTGGATGAAGGCTGGGTACCTTTGGAGATAATGAT
AAAATTCAACAGGTTGAACCGTCTAACAACAGACTTTAATGTAATTGTGGAAGCATTGAGCAAAT
CCAAGGCAGAACTCATGGAAATCAGTGAAGATAAAACTAAAATCAGAAGGTCTCCAAGCAAACCC
CTACCTGAAGTGACTGATGAGTATAAAAATGATGTAAAAAACAGATCTGTTTATATTAAAGGCTTC
CCAACTGATGCAACTCTTGATGACATAAAAGAATGGTTAGAAGATAAAGGTCAAGTACTAAATAT
TCAGATGAGAAGAACATTGCATAAAGCATTTAAGGGATCAATTTTTGTTGTGTTTGATAGCATTGA
ATCTGCTAAGAAATTTGTAGAGACCCCTGGCCAGAAGTACAAAGAAACAGACCTGCTAATACTTTT
CAAGGACGATTACTTTGCCAAAAAAAATGAATCCGGAGGATCTAGCGGAGGCTCCTCTGGCTCTG
AGACACCTGGCACAAGCGAGAGCGCAACACCTGAAAGCAGCGGGGGCAGCAGCGGGGGGTCAAC
CCTGAACATTGAAGACGAGTATAGACTGCATGAAACAAGCAAGGAACCCGACGTGTCCCTGGGCT
CCACCTGGCTGTCCGACTTTCCCCAGGCCTGGGCCGAGACAGGAGGAATGGGCCTGGCCGTGCGG
CAGGCACCCCTGATCATCCCTCTGAAGGCCACCTCTACACCCGTGAGCATCAAGCAGTACCCTATG
TCTCAGGAGGCCAGACTGGGCATCAAGCCTCACATCCAGAGGCTGCTGGACCAGGGCATCCTGGT
GCCATGCCAGAGCCCCTGGAACACACCACTGCTGCCCGTGAAGAAGCCAGGCACCAATGACTATA
GACCCGTGCAGGATCTGAGAGAGGTGAACAAGAGGGTGGAGGATATCCACCCCACCGTGCCCAAC
CCTTACAATCTGCTGTCCGGCCTGCCCCCTTCTCACCAGTGGTATACAGTGCTGGACCTGAAGGAT
GCCTTCTTTTGTCTGAGACTGCACCCTACCAGCCAGCCACTGTTCGCCTTTGAGTGGAGGGACCCT
GAGATGGGCATCTCTGGCCAGCTGACCTGGACACGCCTGCCTCAGGGCTTCAAGAATAGCCCAAC
ACTGTTTAACGAGGCCCTGCACCGCGACCTGGCAGATTTCCGGATCCAGCACCCAGATCTGATCCT
GCTGCAGTACGTGGACGATCTGCTGCTGGCCGCCACCAGCGAGCTGGATTGCCAGCAGGGAACAC
GCGCCCTGCTGCAGACCCTGGGAAACCTGGGATATAGGGCATCCGCCAAGAAGGCCCAGATCTGT
CAGAAGCAGGTGAAGTACCTGGGCTATCTGCTGAAGGAGGGCCAGAGATGGCTGACAGAGGCCA
GGAAGGAGACAGTGATGGGCCAGCCAACACCCAAGACCCCAAGACAGCTGAGGGAGTTCCTGGG
CAAAGCAGGATTTTGCAGGCTGTTCATCCCAGGATTCGCAGAGATGGCAGCACCTCTGTACCCACT
GACCAAGCCGGGCACCCTGTTTAATTGGGGCCCTGACCAGCAGAAGGCCTATCAGGAGATCAAGC
AGGCCCTGCTGACAGCACCAGCCCTGGGCCTGCCAGACCTGACCAAGCCTTTCGAGCTGTTTGTGG
ATGAGAAGCAGGGCTACGCCAAGGGCGTGCTGACCCAGAAGCTGGGACCATGGAGACGGCCCGT
GGCCTATCTGTCCAAGAAGCTGGACCCAGTGGCAGCAGGATGGCCACCATGCCTGAGGATGGTGG
CAGCAATCGCCGTGCTGACAAAGGATGCCGGCAAGCTGACCATGGGACAGCCACTGGTCATCCTG
GCACCACACGCAGTGGAGGCCCTGGTGAAGCAGCCTCCAGATCGCTGGCTGTCTAACGCCCGGAT
GACACACTACCAGGCCCTGCTGCTGGACACCGATCGCGTGCAGTTTGGCCCTGTGGTGGCCCTGAA
TCCAGCCACCCTGCTGCCTCTGCCAGAGGAGGGCCTGCAGCACAACTGTCTGGACATCCTGGCAGA
GGCACACGGAACAAGGCCAGACCTGACCGATCAGCCCCTGCCTGACGCCGATCACACATGGTATA
CCGATGGAAGCTCCCTGCTGCAGGAGGGCCAGAGGAAGGCAGGAGCAGCAGTGACCACAGAGAC
AGAAGTGATCTGGGCCAAGGCCCTGCCAGCAGGCACATCCGCCCAGCGGGCCGAGCTGATCGCCC
TGACCCAGGCCCTGAAGATGGCCGAGGGCAAGAAGCTGAACGTGTACACAGACTCCAGATATGCC
TTCGCCACCGCACACATCCACGGAGAGATCTACAGGCGCCGGGGCTGGCTGACCTCTGAGGGCAA
GGAGATCAAGAACAAGGATGAGATCCTGGCCCTGCTGAAGGCCCTGTTTCTGCCCAAGCGGCTGA
GCATCATCCACTGTCCTGGACACCAGAAGGGACACTCCGCCGAGGCAAGGGGCAATCGGATGGCC
GACCAGGCCGCCAGAAAGGCTGCTATTACTGAAACTCCCGACACTTCCACTCTGCTGATTGAAAAC
TCCTCCCCTTCTGGCGGCTCAAAAAGAACCGCCGACGGCAGCGAATTCGAGTCTCCCAAGAAGAA
GAGGAAAGTCGGCTCTGGCCCTGCCGCTAAGAGAGTGAAGCTGGACTAA
SEQ ID NO: 21
ATGAAACGGACAGCCGACGGAAGCGAGTTCGAGTCACCAAAGAAGAAGCGGAAAGTCGACAAGA
AGTACAGCATCGGCCTGGACATCGGCACCAACTCTGTGGGCTGGGCCGTGATCACCGACGAGTAC
AAGGTGCCCAGCAAGAAATTCAAGGTGCTGGGCAACACCGACCGGCACAGCATCAAGAAGAACC
TGATCGGAGCCCTGCTGTTCGACAGCGGCGAAACAGCCGAGGCCACCCGGCTGAAGAGAACCGCC
AGAAGAAGATACACCAGACGGAAGAACCGGATCTGCTATCTGCAAGAGATCTTCAGCAACGAGAT
GGCCAAGGTGGACGACAGCTTCTTCCACAGACTGGAAGAGTCCTTCCTGGTGGAAGAGGATAAGA
AGCACGAGCGGCACCCCATCTTCGGCAACATCGTGGACGAGGTGGCCTACCACGAGAAGTACCCC
ACCATCTACCACCTGAGAAAGAAACTGGTGGACAGCACCGACAAGGCCGACCTGCGGCTGATCTA
TCTGGCCCTGGCCCACATGATCAAGTTCCGGGGCCACTTCCTGATCGAGGGCGACCTGAACCCCGA
CAACAGCGACGTGGACAAGCTGTTCATCCAGCTGGTGCAGACCTACAACCAGCTGTTCGAGGAAA
ACCCCATCAACGCCAGCGGCGTGGACGCCAAGGCCATCCTGTCTGCCAGACTGAGCAAGAGCAGA
AAGCTGGAAAATCTGATCGCCCAGCTGCCCGGCGAGAAGAAGAATGGCCTGTTCGGAAACCTGAT
TGCCCTGAGCCTGGGCCTGACCCCCAACTTCAAGAGCAACTTCGACCTGGCCGAGGATGCCAAACT
GCAGCTGAGCAAGGACACCTACGACGACGACCTGGACAACCTGCTGGCCCAGATCGGCGACCAGT
ACGCCGACCTGTTTCTGGCCGCCAAGAACCTGTCCGACGCCATCCTGCTGAGCGACATCCTGAGAG
TGAACACCGAGATCACCAAGGCCCCCCTGAGCGCCTCTATGATCAAGAGATACGACGAGCACCAC
CAGGACCTGACCCTGCTGAAAGCTCTCGTGCGGCAGCAGCTGCCTGAGAAGTACAAAGAGATTTT
CTTCGACCAGAGCAAGAACGGCTACGCCGGCTACATTGACGGCGGAGCCAGCCAGGAAGAGTTCT
ACAAGTTCATCAAGCCCATCCTGGAAAAGATGGACGGCACCGAGGAACTGCTCGTGAAGCTGAAG
AGAGAGGACCTGCTGCGGAAGCAGCGGACCTTCGACAACGGCAGCATCCCCCACCAGATCCACCT
GGGAGAGCTGCACGCCATTCTGCGGCGGCAGGAAGATTTTTACCCATTCCTGAAGGACAACCGGG
AAAAGATCGAGAAGATCCTGACCTTCCGCATCCCCTACTACGTGGGCCCTCTGGCCAGGGGAAAC
AGCAGATTCGCCTGGATGACCAGAAAGAGCGAGGAAACCATCACCCCCTGGAACTTCGAGGAAGT
GGTGGACAAGGGCGCTTCCGCCCAGAGCTTCATCGAGCGGATGACCAACTTCGATAAGAACCTGC
CCAACGAGAAGGTGCTGCCCAAGCACAGCCTGCTGTACGAGTACTTCACCGTGTATAACGAGCTG
ACCAAAGTGAAATACGTGACCGAGGGAATGAGAAAGCCCGCCTTCCTGAGCGGCGAGCAGAAAA
AGGCCATCGTGGACCTGCTGTTCAAGACCAACCGGAAAGTGACCGTGAAGCAGCTGAAAGAGGAC
TACTTCAAGAAAATCGAGTGCTTCGACTCCGTGGAAATCTCCGGCGTGGAAGATCGGTTCAACGCC
TCCCTGGGCACATACCACGATCTGCTGAAAATTATCAAGGACAAGGACTTCCTGGACAATGAGGA
AAACGAGGACATTCTGGAAGATATCGTGCTGACCCTGACACTGTTTGAGGACAGAGAGATGATCG
AGGAACGGCTGAAAACCTATGCCCACCTGTTCGACGACAAAGTGATGAAGCAGCTGAAGCGGCGG
AGATACACCGGCTGGGGCAGGCTGAGCCGGAAGCTGATCAACGGCATCCGGGACAAGCAGTCCG
GCAAGACAATCCTGGATTTCCTGAAGTCCGACGGCTTCGCCAACAGAAACTTCATGCAGCTGATCC
ACGACGACAGCCTGACCTTTAAAGAGGACATCCAGAAAGCCCAGGTGTCCGGCCAGGGCGATAGC
CTGCACGAGCACATTGCCAATCTGGCCGGCAGCCCCGCCATTAAGAAGGGCATCCTGCAGACAGT
GAAGGTGGTGGACGAGCTCGTGAAAGTGATGGGCCGGCACAAGCCCGAGAACATCGTGATCGAA
ATGGCCAGAGAGAACCAGACCACCCAGAAGGGACAGAAGAACAGCCGCGAGAGAATGAAGCGG
ATCGAAGAGGGCATCAAAGAGCTGGGCAGCCAGATCCTGAAAGAACACCCCGTGGAAAACACCC
AGCTGCAGAACGAGAAGCTGTACCTGTACTACCTGCAGAATGGGCGGGATATGTACGTGGACCAG
GAACTGGACATCAACCGGCTGTCCGACTACGATGTGGACGCTATCGTGCCTCAGAGCTTTCTGAAG
GACGACTCCATCGACAACAAGGTGCTGACCAGAAGCGACAAGAACCGGGGCAAGAGCGACAACG
TGCCCTCCGAAGAGGTCGTGAAGAAGATGAAGAACTACTGGCGGCAGCTGCTGAACGCCAAGCTG
ATTACCCAGAGAAAGTTCGACAATCTGACCAAGGCCGAGAGAGGCGGCCTGAGCGAACTGGATAA
GGCCGGCTTCATCAAGAGACAGCTGGTGGAAACCCGGCAGATCACAAAGCACGTGGCACAGATCC
TGGACTCCCGGATGAACACTAAGTACGACGAGAATGACAAGCTGATCCGGGAAGTGAAAGTGATC
ACCCTGAAGTCCAAGCTGGTGTCCGATTTCCGGAAGGATTTCCAGTTTTACAAAGTGCGCGAGATC
AACAACTACCACCACGCCCACGACGCCTACCTGAACGCCGTCGTGGGAACCGCCCTGATCAAAAA
GTACCCTAAGCTGGAAAGCGAGTTCGTGTACGGCGACTACAAGGTGTACGACGTGCGGAAGATGA
TCGCCAAGAGCGAGCAGGAAATCGGCAAGGCTACCGCCAAGTACTTCTTCTACAGCAACATCATG
AACTTTTTCAAGACCGAGATTACCCTGGCCAACGGCGAGATCCGGAAGCGGCCTCTGATCGAGAC
AAACGGCGAAACCGGGGAGATCGTGTGGGATAAGGGCCGGGATTTTGCCACCGTGCGGAAAGTGC
TGAGCATGCCCCAAGTGAATATCGTGAAAAAGACCGAGGTGCAGACAGGCGGCTTCAGCAAAGA
GTCTATCCTGCCCAAGAGGAACAGCGATAAGCTGATCGCCAGAAAGAAGGACTGGGACCCTAAGA
AGTACGGCGGCTTCGACAGCCCCACCGTGGCCTATTCTGTGCTGGTGGTGGCCAAAGTGGAAAAG
GGCAAGTCCAAGAAACTGAAGAGTGTGAAAGAGCTGCTGGGGATCACCATCATGGAAAGAAGCA
GCTTCGAGAAGAATCCCATCGACTTTCTGGAAGCCAAGGGCTACAAAGAAGTGAAAAAGGACCTG
ATCATCAAGCTGCCTAAGTACTCCCTGTTCGAGCTGGAAAACGGCCGGAAGAGAATGCTGGCCTCT
GCCGGCGAACTGCAGAAGGGAAACGAACTGGCCCTGCCCTCCAAATATGTGAACTTCCTGTACCT
GGCCAGCCACTATGAGAAGCTGAAGGGCTCCCCCGAGGATAATGAGCAGAAACAGCTGTTTGTGG
AACAGCACAAGCACTACCTGGACGAGATCATCGAGCAGATCAGCGAGTTCTCCAAGAGAGTGATC
CTGGCCGACGCTAATCTGGACAAAGTGCTGTCCGCCTACAACAAGCACCGGGATAAGCCCATCAG
AGAGCAGGCCGAGAATATCATCCACCTGTTTACCCTGACCAATCTGGGAGCCCCTGCCGCCTTCAA
GTACTTTGACACCACCATCGACCGGAAGAGGTACACCAGCACCAAAGAGGTGCTGGACGCCACCC
TGATCCACCAGAGCATCACCGGCCTGTACGAGACACGGATCGACCTGTCTCAGCTGGGAGGTGAC
TCCGGCGGAAGCTCTGGTGGCAGCAAGCGGACCGCCGACGGCTCTGAATTCGAGAGCCCTAAGAA
GAAAAGAAAGGTGAGCGGAGGCTCTAGCGGCGGAAGCACCCTGAACATTGAAGACGAGTATAGA
CTGCATGAAACAAGCAAGGAACCCGACGTGTCCCTGGGCTCCACCTGGCTGTCCGACTTTCCCCAG
GCCTGGGCCGAGACAGGAGGAATGGGCCTGGCCGTGCGGCAGGCACCCCTGATCATCCCTCTGAA
GGCCACCTCTACACCCGTGAGCATCAAGCAGTACCCTATGTCTCAGGAGGCCAGACTGGGCATCA
AGCCTCACATCCAGAGGCTGCTGGACCAGGGCATCCTGGTGCCATGCCAGAGCCCCTGGAACACA
CCACTGCTGCCCGTGAAGAAGCCAGGCACCAATGACTATAGACCCGTGCAGGATCTGAGAGAGGT
GAACAAGAGGGTGGAGGATATCCACCCCACCGTGCCCAACCCTTACAATCTGCTGTCCGGCCTGCC
CCCTTCTCACCAGTGGTATACAGTGCTGGACCTGAAGGATGCCTTCTTTTGTCTGAGACTGCACCCT
ACCAGCCAGCCACTGTTCGCCTTTGAGTGGAGGGACCCTGAGATGGGCATCTCTGGCCAGCTGACC
TGGACACGCCTGCCTCAGGGCTTCAAGAATAGCCCAACACTGTTTAACGAGGCCCTGCACCGCGA
CCTGGCAGATTTCCGGATCCAGCACCCAGATCTGATCCTGCTGCAGTACGTGGACGATCTGCTGCT
GGCCGCCACCAGCGAGCTGGATTGCCAGCAGGGAACACGCGCCCTGCTGCAGACCCTGGGAAACC
TGGGATATAGGGCATCCGCCAAGAAGGCCCAGATCTGTCAGAAGCAGGTGAAGTACCTGGGCTAT
CTGCTGAAGGAGGGCCAGAGATGGCTGACAGAGGCCAGGAAGGAGACAGTGATGGGCCAGCCAA
CACCCAAGACCCCAAGACAGCTGAGGGAGTTCCTGGGCAAAGCAGGATTTTGCAGGCTGTTCATC
CCAGGATTCGCAGAGATGGCAGCACCTCTGTACCCACTGACCAAGCCGGGCACCCTGTTTAATTGG
GGCCCTGACCAGCAGAAGGCCTATCAGGAGATCAAGCAGGCCCTGCTGACAGCACCAGCCCTGGG
CCTGCCAGACCTGACCAAGCCTTTCGAGCTGTTTGTGGATGAGAAGCAGGGCTACGCCAAGGGCG
TGCTGACCCAGAAGCTGGGACCATGGAGACGGCCCGTGGCCTATCTGTCCAAGAAGCTGGACCCA
GTGGCAGCAGGATGGCCACCATGCCTGAGGATGGTGGCAGCAATCGCCGTGCTGACAAAGGATGC
CGGCAAGCTGACCATGGGACAGCCACTGGTCATCCTGGCACCACACGCAGTGGAGGCCCTGGTGA
AGCAGCCTCCAGATCGCTGGCTGTCTAACGCCCGGATGACACACTACCAGGCCCTGCTGCTGGACA
CCGATCGCGTGCAGTTTGGCCCTGTGGTGGCCCTGAATCCAGCCACCCTGCTGCCTCTGCCAGAGG
AGGGCCTGCAGCACAACTGTCTGGACATCCTGGCAGAGGCACACGGAACAAGGCCAGACCTGACC
GATCAGCCCCTGCCTGACGCCGATCACACATGGTATACCGATGGAAGCTCCCTGCTGCAGGAGGG
CCAGAGGAAGGCAGGAGCAGCAGTGACCACAGAGACAGAAGTGATCTGGGCCAAGGCCCTGCCA
GCAGGCACATCCGCCCAGCGGGCCGAGCTGATCGCCCTGACCCAGGCCCTGAAGATGGCCGAGGG
CAAGAAGCTGAACGTGTACACAGACTCCAGATATGCCTTCGCCACCGCACACATCCACGGAGAGA
TCTACAGGCGCCGGGGCTGGCTGACCTCTGAGGGCAAGGAGATCAAGAACAAGGATGAGATCCTG
GCCCTGCTGAAGGCCCTGTTTCTGCCCAAGCGGCTGAGCATCATCCACTGTCCTGGACACCAGAAG
GGACACTCCGCCGAGGCAAGGGGCAATCGGATGGCCGACCAGGCCGCCAGAAAGGCTGCTATTAC
TGAAACTCCCGACACTTCCACTCTGCTGATTGAAAACTCCTCCCCTTCCGGAGGATCTAGCGGAGG
CTCCTCTGGCTCTGAGACACCTGGCACAAGCGAGAGCGCAACACCTGAAAGCAGCGGGGGCAGCA
GCGGGGGGTCAATGGCTGAAAATGGTGATAATGAAAAGATGGCTGCCCTGGAGGCCAAAATCTGT
CATCAAATTGAGTATTATTTTGGCGACTTCAATTTGCCACGGGACAAGTTTCTAAAGGAACAGATA
AAACTGGATGAAGGCTGGGTACCTTTGGAGATAATGATAAAATTCAACAGGTTGAACCGTCTAAC
AACAGACTTTAATGTAATTGTGGAAGCATTGAGCAAATCCAAGGCAGAACTCATGGAAATCAGTG
AAGATAAAACTAAAATCAGAAGGTCTCCAAGCAAACCCCTACCTGAAGTGACTGATGAGTATAAA
AATGATGTAAAAAACAGATCTGTTTATATTAAAGGCTTCCCAACTGATGCAACTCTTGATGACATA
AAAGAATGGTTAGAAGATAAAGGTCAAGTACTAAATATTCAGATGAGAAGAACATTGCATAAAGC
ATTTAAGGGATCAATTTTTGTTGTGTTTGATAGCATTGAATCTGCTAAGAAATTTGTAGAGACCCCT
GGCCAGAAGTACAAAGAAACAGACCTGCTAATACTTTTCAAGGACGATTACTTTGCCAAAAAAAA
TGAAGAAAGAAAACAAAATAAAGTGGAAGCTAAATTAAGAGCTAAACAGGAGCAAGAAGCAAAA
CAAAAGTTAGAAGAAGATGCTGAAATGAAATCTCTAGAAGAAAAGATTGGATGCTTGCTGAAATT
TTCGGGTGATTTAGATGATCAGACCTGTAGAGAAGATTTACACATACTTTTCTCAAATCATGGTGA
AATAAAATGGATAGACTTCGTCAGAGGAGCAAAAGAGGGGATAATTCTATTTAAAGAAAAAGCCA
AGGAAGCATTGGGTAAAGCCAAAGATGCAAATAATGGTAACCTACAATTAAGGAACAAAGAAGT
GACTTGGGAAGTACTAGAAGGAGAGGTGGAAAAAGAAGCACTGAAGAAAATAATAGAAGACCAA
CAAGAATCCCTAAACAAATGGAAGTCAAAAGGTCGTAGATTTAAAGGAAAAGGAAAGGGTAATA
AAGCTGCCCAGCCTGGGTCTGGTAAAGGAAAAGTACAGTTTCAGGGCAAGAAAACGAAATTTGCT
AGTGATGATGAACATGATGAACATGATGAAAATGGTGCAACTGGACCTGTGAAAAGAGCAAGAG
AAGAAACAGACAAAGAAGAACCTGCATCCAAACAACAGAAAACAGAAAATGGTGCTGGAGACCA
GTCTGGCGGCTCAAAAAGAACCGCCGACGGCAGCGAATTCGAGTCTCCCAAGAAGAAGAGGAAA
GTCGGCTCTGGCCCTGCCGCTAAGAGAGTGAAGCTGGACTAA
SEQ ID NO: 22
ATGAAACGGACAGCCGACGGAAGCGAGTTCGAGTCACCAAAGAAGAAGCGGAAAGTCTCTGGCG
GCTCAGCTGAAAATGGTGATAATGAAAAGATGGCTGCCCTGGAGGCCAAAATCTGTCATCAAATT
GAGTATTATTTTGGCGACTTCAATTTGCCACGGGACAAGTTTCTAAAGGAACAGATAAAACTGGAT
GAAGGCTGGGTACCTTTGGAGATAATGATAAAATTCAACAGGTTGAACCGTCTAACAACAGACTT
TAATGTAATTGTGGAAGCATTGAGCAAATCCAAGGCAGAACTCATGGAAATCAGTGAAGATAAAA
CTAAAATCAGAAGGTCTCCAAGCAAACCCCTACCTGAAGTGACTGATGAGTATAAAAATGATGTA
AAAAACAGATCTGTTTATATTAAAGGCTTCCCAACTGATGCAACTCTTGATGACATAAAAGAATGG
TTAGAAGATAAAGGTCAAGTACTAAATATTCAGATGAGAAGAACATTGCATAAAGCATTTAAGGG
ATCAATTTTTGTTGTGTTTGATAGCATTGAATCTGCTAAGAAATTTGTAGAGACCCCTGGCCAGAA
GTACAAAGAAACAGACCTGCTAATACTTTTCAAGGACGATTACTTTGCCAAAAAAAATGAAGAAA
GAAAACAAAATAAAGTGGAAGCTAAATTAAGAGCTAAACAGGAGCAAGAAGCAAAACAAAAGTT
AGAAGAAGATGCTGAAATGAAATCTCTAGAAGAAAAGATTGGATGCTTGCTGAAATTTTCGGGTG
ATTTAGATGATCAGACCTGTAGAGAAGATTTACACATACTTTTCTCAAATCATGGTGAAATAAAAT
GGATAGACTTCGTCAGAGGAGCAAAAGAGGGGATAATTCTATTTAAAGAAAAAGCCAAGGAAGC
ATTGGGTAAAGCCAAAGATGCAAATAATGGTAACCTACAATTAAGGAACAAAGAAGTGACTTGGG
AAGTACTAGAAGGAGAGGTGGAAAAAGAAGCACTGAAGAAAATAATAGAAGACCAACAAGAATC
CCTAAACAAATGGAAGTCAAAAGGTCGTAGATTTAAAGGAAAAGGAAAGGGTAATAAAGCTGCC
CAGCCTGGGTCTGGTAAAGGAAAAGTACAGTTTCAGGGCAAGAAAACGAAATTTGCTAGTGATGA
TGAACATGATGAACATGATGAAAATGGTGCAACTGGACCTGTGAAAAGAGCAAGAGAAGAAACA
GACAAAGAAGAACCTGCATCCAAACAACAGAAAACAGAAAATGGTGCTGGAGACCAGTCCGGAG
GATCTAGCGGAGGCTCCTCTGGCTCTGAGACACCTGGCACAAGCGAGAGCGCAACACCTGAAAGC
AGCGGGGGCAGCAGCGGGGGGTCAGACAAGAAGTACAGCATCGGCCTGGACATCGGCACCAACT
CTGTGGGCTGGGCCGTGATCACCGACGAGTACAAGGTGCCCAGCAAGAAATTCAAGGTGCTGGGC
AACACCGACCGGCACAGCATCAAGAAGAACCTGATCGGAGCCCTGCTGTTCGACAGCGGCGAAAC
AGCCGAGGCCACCCGGCTGAAGAGAACCGCCAGAAGAAGATACACCAGACGGAAGAACCGGATC
TGCTATCTGCAAGAGATCTTCAGCAACGAGATGGCCAAGGTGGACGACAGCTTCTTCCACAGACT
GGAAGAGTCCTTCCTGGTGGAAGAGGATAAGAAGCACGAGCGGCACCCCATCTTCGGCAACATCG
TGGACGAGGTGGCCTACCACGAGAAGTACCCCACCATCTACCACCTGAGAAAGAAACTGGTGGAC
AGCACCGACAAGGCCGACCTGCGGCTGATCTATCTGGCCCTGGCCCACATGATCAAGTTCCGGGG
CCACTTCCTGATCGAGGGCGACCTGAACCCCGACAACAGCGACGTGGACAAGCTGTTCATCCAGC
TGGTGCAGACCTACAACCAGCTGTTCGAGGAAAACCCCATCAACGCCAGCGGCGTGGACGCCAAG
GCCATCCTGTCTGCCAGACTGAGCAAGAGCAGAAAGCTGGAAAATCTGATCGCCCAGCTGCCCGG
CGAGAAGAAGAATGGCCTGTTCGGAAACCTGATTGCCCTGAGCCTGGGCCTGACCCCCAACTTCA
AGAGCAACTTCGACCTGGCCGAGGATGCCAAACTGCAGCTGAGCAAGGACACCTACGACGACGAC
CTGGACAACCTGCTGGCCCAGATCGGCGACCAGTACGCCGACCTGTTTCTGGCCGCCAAGAACCTG
TCCGACGCCATCCTGCTGAGCGACATCCTGAGAGTGAACACCGAGATCACCAAGGCCCCCCTGAG
CGCCTCTATGATCAAGAGATACGACGAGCACCACCAGGACCTGACCCTGCTGAAAGCTCTCGTGC
GGCAGCAGCTGCCTGAGAAGTACAAAGAGATTTTCTTCGACCAGAGCAAGAACGGCTACGCCGGC
TACATTGACGGCGGAGCCAGCCAGGAAGAGTTCTACAAGTTCATCAAGCCCATCCTGGAAAAGAT
GGACGGCACCGAGGAACTGCTCGTGAAGCTGAAGAGAGAGGACCTGCTGCGGAAGCAGCGGACC
TTCGACAACGGCAGCATCCCCCACCAGATCCACCTGGGAGAGCTGCACGCCATTCTGCGGCGGCA
GGAAGATTTTTACCCATTCCTGAAGGACAACCGGGAAAAGATCGAGAAGATCCTGACCTTCCGCA
TCCCCTACTACGTGGGCCCTCTGGCCAGGGGAAACAGCAGATTCGCCTGGATGACCAGAAAGAGC
GAGGAAACCATCACCCCCTGGAACTTCGAGGAAGTGGTGGACAAGGGCGCTTCCGCCCAGAGCTT
CATCGAGCGGATGACCAACTTCGATAAGAACCTGCCCAACGAGAAGGTGCTGCCCAAGCACAGCC
TGCTGTACGAGTACTTCACCGTGTATAACGAGCTGACCAAAGTGAAATACGTGACCGAGGGAATG
AGAAAGCCCGCCTTCCTGAGCGGCGAGCAGAAAAAGGCCATCGTGGACCTGCTGTTCAAGACCAA
CCGGAAAGTGACCGTGAAGCAGCTGAAAGAGGACTACTTCAAGAAAATCGAGTGCTTCGACTCCG
TGGAAATCTCCGGCGTGGAAGATCGGTTCAACGCCTCCCTGGGCACATACCACGATCTGCTGAAA
ATTATCAAGGACAAGGACTTCCTGGACAATGAGGAAAACGAGGACATTCTGGAAGATATCGTGCT
GACCCTGACACTGTTTGAGGACAGAGAGATGATCGAGGAACGGCTGAAAACCTATGCCCACCTGT
TCGACGACAAAGTGATGAAGCAGCTGAAGCGGCGGAGATACACCGGCTGGGGCAGGCTGAGCCG
GAAGCTGATCAACGGCATCCGGGACAAGCAGTCCGGCAAGACAATCCTGGATTTCCTGAAGTCCG
ACGGCTTCGCCAACAGAAACTTCATGCAGCTGATCCACGACGACAGCCTGACCTTTAAAGAGGAC
ATCCAGAAAGCCCAGGTGTCCGGCCAGGGCGATAGCCTGCACGAGCACATTGCCAATCTGGCCGG
CAGCCCCGCCATTAAGAAGGGCATCCTGCAGACAGTGAAGGTGGTGGACGAGCTCGTGAAAGTGA
TGGGCCGGCACAAGCCCGAGAACATCGTGATCGAAATGGCCAGAGAGAACCAGACCACCCAGAA
GGGACAGAAGAACAGCCGCGAGAGAATGAAGCGGATCGAAGAGGGCATCAAAGAGCTGGGCAGC
CAGATCCTGAAAGAACACCCCGTGGAAAACACCCAGCTGCAGAACGAGAAGCTGTACCTGTACTA
CCTGCAGAATGGGCGGGATATGTACGTGGACCAGGAACTGGACATCAACCGGCTGTCCGACTACG
ATGTGGACGCTATCGTGCCTCAGAGCTTTCTGAAGGACGACTCCATCGACAACAAGGTGCTGACCA
GAAGCGACAAGAACCGGGGCAAGAGCGACAACGTGCCCTCCGAAGAGGTCGTGAAGAAGATGAA
GAACTACTGGCGGCAGCTGCTGAACGCCAAGCTGATTACCCAGAGAAAGTTCGACAATCTGACCA
AGGCCGAGAGAGGCGGCCTGAGCGAACTGGATAAGGCCGGCTTCATCAAGAGACAGCTGGTGGA
AACCCGGCAGATCACAAAGCACGTGGCACAGATCCTGGACTCCCGGATGAACACTAAGTACGACG
AGAATGACAAGCTGATCCGGGAAGTGAAAGTGATCACCCTGAAGTCCAAGCTGGTGTCCGATTTC
CGGAAGGATTTCCAGTTTTACAAAGTGCGCGAGATCAACAACTACCACCACGCCCACGACGCCTA
CCTGAACGCCGTCGTGGGAACCGCCCTGATCAAAAAGTACCCTAAGCTGGAAAGCGAGTTCGTGT
ACGGCGACTACAAGGTGTACGACGTGCGGAAGATGATCGCCAAGAGCGAGCAGGAAATCGGCAA
GGCTACCGCCAAGTACTTCTTCTACAGCAACATCATGAACTTTTTCAAGACCGAGATTACCCTGGC
CAACGGCGAGATCCGGAAGCGGCCTCTGATCGAGACAAACGGCGAAACCGGGGAGATCGTGTGG
GATAAGGGCCGGGATTTTGCCACCGTGCGGAAAGTGCTGAGCATGCCCCAAGTGAATATCGTGAA
AAAGACCGAGGTGCAGACAGGCGGCTTCAGCAAAGAGTCTATCCTGCCCAAGAGGAACAGCGAT
AAGCTGATCGCCAGAAAGAAGGACTGGGACCCTAAGAAGTACGGCGGCTTCGACAGCCCCACCGT
GGCCTATTCTGTGCTGGTGGTGGCCAAAGTGGAAAAGGGCAAGTCCAAGAAACTGAAGAGTGTGA
AAGAGCTGCTGGGGATCACCATCATGGAAAGAAGCAGCTTCGAGAAGAATCCCATCGACTTTCTG
GAAGCCAAGGGCTACAAAGAAGTGAAAAAGGACCTGATCATCAAGCTGCCTAAGTACTCCCTGTT
CGAGCTGGAAAACGGCCGGAAGAGAATGCTGGCCTCTGCCGGCGAACTGCAGAAGGGAAACGAA
CTGGCCCTGCCCTCCAAATATGTGAACTTCCTGTACCTGGCCAGCCACTATGAGAAGCTGAAGGGC
TCCCCCGAGGATAATGAGCAGAAACAGCTGTTTGTGGAACAGCACAAGCACTACCTGGACGAGAT
CATCGAGCAGATCAGCGAGTTCTCCAAGAGAGTGATCCTGGCCGACGCTAATCTGGACAAAGTGC
TGTCCGCCTACAACAAGCACCGGGATAAGCCCATCAGAGAGCAGGCCGAGAATATCATCCACCTG
TTTACCCTGACCAATCTGGGAGCCCCTGCCGCCTTCAAGTACTTTGACACCACCATCGACCGGAAG
AGGTACACCAGCACCAAAGAGGTGCTGGACGCCACCCTGATCCACCAGAGCATCACCGGCCTGTA
CGAGACACGGATCGACCTGTCTCAGCTGGGAGGTGACTCCGGCGGAAGCTCTGGTGGCAGCAAGC
GGACCGCCGACGGCTCTGAATTCGAGAGCCCTAAGAAGAAAAGAAAGGTGAGCGGAGGCTCTAG
CGGCGGAAGCACCCTGAACATTGAAGACGAGTATAGACTGCATGAAACAAGCAAGGAACCCGAC
GTGTCCCTGGGCTCCACCTGGCTGTCCGACTTTCCCCAGGCCTGGGCCGAGACAGGAGGAATGGGC
CTGGCCGTGCGGCAGGCACCCCTGATCATCCCTCTGAAGGCCACCTCTACACCCGTGAGCATCAAG
CAGTACCCTATGTCTCAGGAGGCCAGACTGGGCATCAAGCCTCACATCCAGAGGCTGCTGGACCA
GGGCATCCTGGTGCCATGCCAGAGCCCCTGGAACACACCACTGCTGCCCGTGAAGAAGCCAGGCA
CCAATGACTATAGACCCGTGCAGGATCTGAGAGAGGTGAACAAGAGGGTGGAGGATATCCACCCC
ACCGTGCCCAACCCTTACAATCTGCTGTCCGGCCTGCCCCCTTCTCACCAGTGGTATACAGTGCTG
GACCTGAAGGATGCCTTCTTTTGTCTGAGACTGCACCCTACCAGCCAGCCACTGTTCGCCTTTGAGT
GGAGGGACCCTGAGATGGGCATCTCTGGCCAGCTGACCTGGACACGCCTGCCTCAGGGCTTCAAG
AATAGCCCAACACTGTTTAACGAGGCCCTGCACCGCGACCTGGCAGATTTCCGGATCCAGCACCCA
GATCTGATCCTGCTGCAGTACGTGGACGATCTGCTGCTGGCCGCCACCAGCGAGCTGGATTGCCAG
CAGGGAACACGCGCCCTGCTGCAGACCCTGGGAAACCTGGGATATAGGGCATCCGCCAAGAAGGC
CCAGATCTGTCAGAAGCAGGTGAAGTACCTGGGCTATCTGCTGAAGGAGGGCCAGAGATGGCTGA
CAGAGGCCAGGAAGGAGACAGTGATGGGCCAGCCAACACCCAAGACCCCAAGACAGCTGAGGGA
GTTCCTGGGCAAAGCAGGATTTTGCAGGCTGTTCATCCCAGGATTCGCAGAGATGGCAGCACCTCT
GTACCCACTGACCAAGCCGGGCACCCTGTTTAATTGGGGCCCTGACCAGCAGAAGGCCTATCAGG
AGATCAAGCAGGCCCTGCTGACAGCACCAGCCCTGGGCCTGCCAGACCTGACCAAGCCTTTCGAG
CTGTTTGTGGATGAGAAGCAGGGCTACGCCAAGGGCGTGCTGACCCAGAAGCTGGGACCATGGAG
ACGGCCCGTGGCCTATCTGTCCAAGAAGCTGGACCCAGTGGCAGCAGGATGGCCACCATGCCTGA
GGATGGTGGCAGCAATCGCCGTGCTGACAAAGGATGCCGGCAAGCTGACCATGGGACAGCCACTG
GTCATCCTGGCACCACACGCAGTGGAGGCCCTGGTGAAGCAGCCTCCAGATCGCTGGCTGTCTAAC
GCCCGGATGACACACTACCAGGCCCTGCTGCTGGACACCGATCGCGTGCAGTTTGGCCCTGTGGTG
GCCCTGAATCCAGCCACCCTGCTGCCTCTGCCAGAGGAGGGCCTGCAGCACAACTGTCTGGACATC
CTGGCAGAGGCACACGGAACAAGGCCAGACCTGACCGATCAGCCCCTGCCTGACGCCGATCACAC
ATGGTATACCGATGGAAGCTCCCTGCTGCAGGAGGGCCAGAGGAAGGCAGGAGCAGCAGTGACC
ACAGAGACAGAAGTGATCTGGGCCAAGGCCCTGCCAGCAGGCACATCCGCCCAGCGGGCCGAGCT
GATCGCCCTGACCCAGGCCCTGAAGATGGCCGAGGGCAAGAAGCTGAACGTGTACACAGACTCCA
GATATGCCTTCGCCACCGCACACATCCACGGAGAGATCTACAGGCGCCGGGGCTGGCTGACCTCT
GAGGGCAAGGAGATCAAGAACAAGGATGAGATCCTGGCCCTGCTGAAGGCCCTGTTTCTGCCCAA
GCGGCTGAGCATCATCCACTGTCCTGGACACCAGAAGGGACACTCCGCCGAGGCAAGGGGCAATC
GGATGGCCGACCAGGCCGCCAGAAAGGCTGCTATTACTGAAACTCCCGACACTTCCACTCTGCTGA
TTGAAAACTCCTCCCCTTCTGGCGGCTCAAAAAGAACCGCCGACGGCAGCGAATTCGAGTCTCCCA
AGAAGAAGAGGAAAGTCGGCTCTGGCCCTGCCGCTAAGAGAGTGAAGCTGGACTAA
SEQ ID NO: 23
ATGAAACGGACAGCCGACGGAAGCGAGTTCGAGTCACCAAAGAAGAAGCGGAAAGTCGACAAGA
AGTACAGCATCGGCCTGGACATCGGCACCAACTCTGTGGGCTGGGCCGTGATCACCGACGAGTAC
AAGGTGCCCAGCAAGAAATTCAAGGTGCTGGGCAACACCGACCGGCACAGCATCAAGAAGAACC
TGATCGGAGCCCTGCTGTTCGACAGCGGCGAAACAGCCGAGGCCACCCGGCTGAAGAGAACCGCC
AGAAGAAGATACACCAGACGGAAGAACCGGATCTGCTATCTGCAAGAGATCTTCAGCAACGAGAT
GGCCAAGGTGGACGACAGCTTCTTCCACAGACTGGAAGAGTCCTTCCTGGTGGAAGAGGATAAGA
AGCACGAGCGGCACCCCATCTTCGGCAACATCGTGGACGAGGTGGCCTACCACGAGAAGTACCCC
ACCATCTACCACCTGAGAAAGAAACTGGTGGACAGCACCGACAAGGCCGACCTGCGGCTGATCTA
TCTGGCCCTGGCCCACATGATCAAGTTCCGGGGCCACTTCCTGATCGAGGGCGACCTGAACCCCGA
CAACAGCGACGTGGACAAGCTGTTCATCCAGCTGGTGCAGACCTACAACCAGCTGTTCGAGGAAA
ACCCCATCAACGCCAGCGGCGTGGACGCCAAGGCCATCCTGTCTGCCAGACTGAGCAAGAGCAGA
AAGCTGGAAAATCTGATCGCCCAGCTGCCCGGCGAGAAGAAGAATGGCCTGTTCGGAAACCTGAT
TGCCCTGAGCCTGGGCCTGACCCCCAACTTCAAGAGCAACTTCGACCTGGCCGAGGATGCCAAACT
GCAGCTGAGCAAGGACACCTACGACGACGACCTGGACAACCTGCTGGCCCAGATCGGCGACCAGT
ACGCCGACCTGTTTCTGGCCGCCAAGAACCTGTCCGACGCCATCCTGCTGAGCGACATCCTGAGAG
TGAACACCGAGATCACCAAGGCCCCCCTGAGCGCCTCTATGATCAAGAGATACGACGAGCACCAC
CAGGACCTGACCCTGCTGAAAGCTCTCGTGCGGCAGCAGCTGCCTGAGAAGTACAAAGAGATTTT
CTTCGACCAGAGCAAGAACGGCTACGCCGGCTACATTGACGGCGGAGCCAGCCAGGAAGAGTTCT
ACAAGTTCATCAAGCCCATCCTGGAAAAGATGGACGGCACCGAGGAACTGCTCGTGAAGCTGAAG
AGAGAGGACCTGCTGCGGAAGCAGCGGACCTTCGACAACGGCAGCATCCCCCACCAGATCCACCT
GGGAGAGCTGCACGCCATTCTGCGGCGGCAGGAAGATTTTTACCCATTCCTGAAGGACAACCGGG
AAAAGATCGAGAAGATCCTGACCTTCCGCATCCCCTACTACGTGGGCCCTCTGGCCAGGGGAAAC
AGCAGATTCGCCTGGATGACCAGAAAGAGCGAGGAAACCATCACCCCCTGGAACTTCGAGGAAGT
GGTGGACAAGGGCGCTTCCGCCCAGAGCTTCATCGAGCGGATGACCAACTTCGATAAGAACCTGC
CCAACGAGAAGGTGCTGCCCAAGCACAGCCTGCTGTACGAGTACTTCACCGTGTATAACGAGCTG
ACCAAAGTGAAATACGTGACCGAGGGAATGAGAAAGCCCGCCTTCCTGAGCGGCGAGCAGAAAA
AGGCCATCGTGGACCTGCTGTTCAAGACCAACCGGAAAGTGACCGTGAAGCAGCTGAAAGAGGAC
TACTTCAAGAAAATCGAGTGCTTCGACTCCGTGGAAATCTCCGGCGTGGAAGATCGGTTCAACGCC
TCCCTGGGCACATACCACGATCTGCTGAAAATTATCAAGGACAAGGACTTCCTGGACAATGAGGA
AAACGAGGACATTCTGGAAGATATCGTGCTGACCCTGACACTGTTTGAGGACAGAGAGATGATCG
AGGAACGGCTGAAAACCTATGCCCACCTGTTCGACGACAAAGTGATGAAGCAGCTGAAGCGGCGG
AGATACACCGGCTGGGGCAGGCTGAGCCGGAAGCTGATCAACGGCATCCGGGACAAGCAGTCCG
GCAAGACAATCCTGGATTTCCTGAAGTCCGACGGCTTCGCCAACAGAAACTTCATGCAGCTGATCC
ACGACGACAGCCTGACCTTTAAAGAGGACATCCAGAAAGCCCAGGTGTCCGGCCAGGGCGATAGC
CTGCACGAGCACATTGCCAATCTGGCCGGCAGCCCCGCCATTAAGAAGGGCATCCTGCAGACAGT
GAAGGTGGTGGACGAGCTCGTGAAAGTGATGGGCCGGCACAAGCCCGAGAACATCGTGATCGAA
ATGGCCAGAGAGAACCAGACCACCCAGAAGGGACAGAAGAACAGCCGCGAGAGAATGAAGCGG
ATCGAAGAGGGCATCAAAGAGCTGGGCAGCCAGATCCTGAAAGAACACCCCGTGGAAAACACCC
AGCTGCAGAACGAGAAGCTGTACCTGTACTACCTGCAGAATGGGCGGGATATGTACGTGGACCAG
GAACTGGACATCAACCGGCTGTCCGACTACGATGTGGACGCTATCGTGCCTCAGAGCTTTCTGAAG
GACGACTCCATCGACAACAAGGTGCTGACCAGAAGCGACAAGAACCGGGGCAAGAGCGACAACG
TGCCCTCCGAAGAGGTCGTGAAGAAGATGAAGAACTACTGGCGGCAGCTGCTGAACGCCAAGCTG
ATTACCCAGAGAAAGTTCGACAATCTGACCAAGGCCGAGAGAGGCGGCCTGAGCGAACTGGATAA
GGCCGGCTTCATCAAGAGACAGCTGGTGGAAACCCGGCAGATCACAAAGCACGTGGCACAGATCC
TGGACTCCCGGATGAACACTAAGTACGACGAGAATGACAAGCTGATCCGGGAAGTGAAAGTGATC
ACCCTGAAGTCCAAGCTGGTGTCCGATTTCCGGAAGGATTTCCAGTTTTACAAAGTGCGCGAGATC
AACAACTACCACCACGCCCACGACGCCTACCTGAACGCCGTCGTGGGAACCGCCCTGATCAAAAA
GTACCCTAAGCTGGAAAGCGAGTTCGTGTACGGCGACTACAAGGTGTACGACGTGCGGAAGATGA
TCGCCAAGAGCGAGCAGGAAATCGGCAAGGCTACCGCCAAGTACTTCTTCTACAGCAACATCATG
AACTTTTTCAAGACCGAGATTACCCTGGCCAACGGCGAGATCCGGAAGCGGCCTCTGATCGAGAC
AAACGGCGAAACCGGGGAGATCGTGTGGGATAAGGGCCGGGATTTTGCCACCGTGCGGAAAGTGC
TGAGCATGCCCCAAGTGAATATCGTGAAAAAGACCGAGGTGCAGACAGGCGGCTTCAGCAAAGA
GTCTATCCTGCCCAAGAGGAACAGCGATAAGCTGATCGCCAGAAAGAAGGACTGGGACCCTAAGA
AGTACGGCGGCTTCGACAGCCCCACCGTGGCCTATTCTGTGCTGGTGGTGGCCAAAGTGGAAAAG
GGCAAGTCCAAGAAACTGAAGAGTGTGAAAGAGCTGCTGGGGATCACCATCATGGAAAGAAGCA
GCTTCGAGAAGAATCCCATCGACTTTCTGGAAGCCAAGGGCTACAAAGAAGTGAAAAAGGACCTG
ATCATCAAGCTGCCTAAGTACTCCCTGTTCGAGCTGGAAAACGGCCGGAAGAGAATGCTGGCCTCT
GCCGGCGAACTGCAGAAGGGAAACGAACTGGCCCTGCCCTCCAAATATGTGAACTTCCTGTACCT
GGCCAGCCACTATGAGAAGCTGAAGGGCTCCCCCGAGGATAATGAGCAGAAACAGCTGTTTGTGG
AACAGCACAAGCACTACCTGGACGAGATCATCGAGCAGATCAGCGAGTTCTCCAAGAGAGTGATC
CTGGCCGACGCTAATCTGGACAAAGTGCTGTCCGCCTACAACAAGCACCGGGATAAGCCCATCAG
AGAGCAGGCCGAGAATATCATCCACCTGTTTACCCTGACCAATCTGGGAGCCCCTGCCGCCTTCAA
GTACTTTGACACCACCATCGACCGGAAGAGGTACACCAGCACCAAAGAGGTGCTGGACGCCACCC
TGATCCACCAGAGCATCACCGGCCTGTACGAGACACGGATCGACCTGTCTCAGCTGGGAGGTGAC
TCCGGAGGATCTAGCGGAGGCTCCTCTGGCTCTGAGACACCTGGCACAAGCGAGAGCGCAACACC
TGAAAGCAGCGGGGGCAGCAGCGGGGGGTCAATGGCTGAAAATGGTGATAATGAAAAGATGGCT
GCCCTGGAGGCCAAAATCTGTCATCAAATTGAGTATTATTTTGGCGACTTCAATTTGCCACGGGAC
AAGTTTCTAAAGGAACAGATAAAACTGGATGAAGGCTGGGTACCTTTGGAGATAATGATAAAATT
CAACAGGTTGAACCGTCTAACAACAGACTTTAATGTAATTGTGGAAGCATTGAGCAAATCCAAGG
CAGAACTCATGGAAATCAGTGAAGATAAAACTAAAATCAGAAGGTCTCCAAGCAAACCCCTACCT
GAAGTGACTGATGAGTATAAAAATGATGTAAAAAACAGATCTGTTTATATTAAAGGCTTCCCAACT
GATGCAACTCTTGATGACATAAAAGAATGGTTAGAAGATAAAGGTCAAGTACTAAATATTCAGAT
GAGAAGAACATTGCATAAAGCATTTAAGGGATCAATTTTTGTTGTGTTTGATAGCATTGAATCTGC
TAAGAAATTTGTAGAGACCCCTGGCCAGAAGTACAAAGAAACAGACCTGCTAATACTTTTCAAGG
ACGATTACTTTGCCAAAAAAAATGAAGAAAGAAAACAAAATAAAGTGGAAGCTAAATTAAGAGC
TAAACAGGAGCAAGAAGCAAAACAAAAGTTAGAAGAAGATGCTGAAATGAAATCTCTAGAAGAA
AAGATTGGATGCTTGCTGAAATTTTCGGGTGATTTAGATGATCAGACCTGTAGAGAAGATTTACAC
ATACTTTTCTCAAATCATGGTGAAATAAAATGGATAGACTTCGTCAGAGGAGCAAAAGAGGGGAT
AATTCTATTTAAAGAAAAAGCCAAGGAAGCATTGGGTAAAGCCAAAGATGCAAATAATGGTAACC
TACAATTAAGGAACAAAGAAGTGACTTGGGAAGTACTAGAAGGAGAGGTGGAAAAAGAAGCACT
GAAGAAAATAATAGAAGACCAACAAGAATCCCTAAACAAATGGAAGTCAAAAGGTCGTAGATTT
AAAGGAAAAGGAAAGGGTAATAAAGCTGCCCAGCCTGGGTCTGGTAAAGGAAAAGTACAGTTTC
AGGGCAAGAAAACGAAATTTGCTAGTGATGATGAACATGATGAACATGATGAAAATGGTGCAACT
GGACCTGTGAAAAGAGCAAGAGAAGAAACAGACAAAGAAGAACCTGCATCCAAACAACAGAAAA
CAGAAAATGGTGCTGGAGACCAGTCCGGCGGAAGCTCTGGTGGCAGCAAGCGGACCGCCGACGG
CTCTGAATTCGAGAGCCCTAAGAAGAAAAGAAAGGTGAGCGGAGGCTCTAGCGGCGGAAGCACC
CTGAACATTGAAGACGAGTATAGACTGCATGAAACAAGCAAGGAACCCGACGTGTCCCTGGGCTC
CACCTGGCTGTCCGACTTTCCCCAGGCCTGGGCCGAGACAGGAGGAATGGGCCTGGCCGTGCGGC
AGGCACCCCTGATCATCCCTCTGAAGGCCACCTCTACACCCGTGAGCATCAAGCAGTACCCTATGT
CTCAGGAGGCCAGACTGGGCATCAAGCCTCACATCCAGAGGCTGCTGGACCAGGGCATCCTGGTG
CCATGCCAGAGCCCCTGGAACACACCACTGCTGCCCGTGAAGAAGCCAGGCACCAATGACTATAG
ACCCGTGCAGGATCTGAGAGAGGTGAACAAGAGGGTGGAGGATATCCACCCCACCGTGCCCAACC
CTTACAATCTGCTGTCCGGCCTGCCCCCTTCTCACCAGTGGTATACAGTGCTGGACCTGAAGGATG
CCTTCTTTTGTCTGAGACTGCACCCTACCAGCCAGCCACTGTTCGCCTTTGAGTGGAGGGACCCTG
AGATGGGCATCTCTGGCCAGCTGACCTGGACACGCCTGCCTCAGGGCTTCAAGAATAGCCCAACA
CTGTTTAACGAGGCCCTGCACCGCGACCTGGCAGATTTCCGGATCCAGCACCCAGATCTGATCCTG
CTGCAGTACGTGGACGATCTGCTGCTGGCCGCCACCAGCGAGCTGGATTGCCAGCAGGGAACACG
CGCCCTGCTGCAGACCCTGGGAAACCTGGGATATAGGGCATCCGCCAAGAAGGCCCAGATCTGTC
AGAAGCAGGTGAAGTACCTGGGCTATCTGCTGAAGGAGGGCCAGAGATGGCTGACAGAGGCCAG
GAAGGAGACAGTGATGGGCCAGCCAACACCCAAGACCCCAAGACAGCTGAGGGAGTTCCTGGGC
AAAGCAGGATTTTGCAGGCTGTTCATCCCAGGATTCGCAGAGATGGCAGCACCTCTGTACCCACTG
ACCAAGCCGGGCACCCTGTTTAATTGGGGCCCTGACCAGCAGAAGGCCTATCAGGAGATCAAGCA
GGCCCTGCTGACAGCACCAGCCCTGGGCCTGCCAGACCTGACCAAGCCTTTCGAGCTGTTTGTGGA
TGAGAAGCAGGGCTACGCCAAGGGCGTGCTGACCCAGAAGCTGGGACCATGGAGACGGCCCGTG
GCCTATCTGTCCAAGAAGCTGGACCCAGTGGCAGCAGGATGGCCACCATGCCTGAGGATGGTGGC
AGCAATCGCCGTGCTGACAAAGGATGCCGGCAAGCTGACCATGGGACAGCCACTGGTCATCCTGG
CACCACACGCAGTGGAGGCCCTGGTGAAGCAGCCTCCAGATCGCTGGCTGTCTAACGCCCGGATG
ACACACTACCAGGCCCTGCTGCTGGACACCGATCGCGTGCAGTTTGGCCCTGTGGTGGCCCTGAAT
CCAGCCACCCTGCTGCCTCTGCCAGAGGAGGGCCTGCAGCACAACTGTCTGGACATCCTGGCAGA
GGCACACGGAACAAGGCCAGACCTGACCGATCAGCCCCTGCCTGACGCCGATCACACATGGTATA
CCGATGGAAGCTCCCTGCTGCAGGAGGGCCAGAGGAAGGCAGGAGCAGCAGTGACCACAGAGAC
AGAAGTGATCTGGGCCAAGGCCCTGCCAGCAGGCACATCCGCCCAGCGGGCCGAGCTGATCGCCC
TGACCCAGGCCCTGAAGATGGCCGAGGGCAAGAAGCTGAACGTGTACACAGACTCCAGATATGCC
TTCGCCACCGCACACATCCACGGAGAGATCTACAGGCGCCGGGGCTGGCTGACCTCTGAGGGCAA
GGAGATCAAGAACAAGGATGAGATCCTGGCCCTGCTGAAGGCCCTGTTTCTGCCCAAGCGGCTGA
GCATCATCCACTGTCCTGGACACCAGAAGGGACACTCCGCCGAGGCAAGGGGCAATCGGATGGCC
GACCAGGCCGCCAGAAAGGCTGCTATTACTGAAACTCCCGACACTTCCACTCTGCTGATTGAAAAC
TCCTCCCCTTCTGGCGGCTCAAAAAGAACCGCCGACGGCAGCGAATTCGAGTCTCCCAAGAAGAA
GAGGAAAGTCGGCTCTGGCCCTGCCGCTAAGAGAGTGAAGCTGGACTAA
SEQ ID NO: 24
ATGAAACGGACAGCCGACGGAAGCGAGTTCGAGTCACCAAAGAAGAAGCGGAAAGTCGACAAGA
AGTACAGCATCGGCCTGGACATCGGCACCAACTCTGTGGGCTGGGCCGTGATCACCGACGAGTAC
AAGGTGCCCAGCAAGAAATTCAAGGTGCTGGGCAACACCGACCGGCACAGCATCAAGAAGAACC
TGATCGGAGCCCTGCTGTTCGACAGCGGCGAAACAGCCGAGGCCACCCGGCTGAAGAGAACCGCC
AGAAGAAGATACACCAGACGGAAGAACCGGATCTGCTATCTGCAAGAGATCTTCAGCAACGAGAT
GGCCAAGGTGGACGACAGCTTCTTCCACAGACTGGAAGAGTCCTTCCTGGTGGAAGAGGATAAGA
AGCACGAGCGGCACCCCATCTTCGGCAACATCGTGGACGAGGTGGCCTACCACGAGAAGTACCCC
ACCATCTACCACCTGAGAAAGAAACTGGTGGACAGCACCGACAAGGCCGACCTGCGGCTGATCTA
TCTGGCCCTGGCCCACATGATCAAGTTCCGGGGCCACTTCCTGATCGAGGGCGACCTGAACCCCGA
CAACAGCGACGTGGACAAGCTGTTCATCCAGCTGGTGCAGACCTACAACCAGCTGTTCGAGGAAA
ACCCCATCAACGCCAGCGGCGTGGACGCCAAGGCCATCCTGTCTGCCAGACTGAGCAAGAGCAGA
AAGCTGGAAAATCTGATCGCCCAGCTGCCCGGCGAGAAGAAGAATGGCCTGTTCGGAAACCTGAT
TGCCCTGAGCCTGGGCCTGACCCCCAACTTCAAGAGCAACTTCGACCTGGCCGAGGATGCCAAACT
GCAGCTGAGCAAGGACACCTACGACGACGACCTGGACAACCTGCTGGCCCAGATCGGCGACCAGT
ACGCCGACCTGTTTCTGGCCGCCAAGAACCTGTCCGACGCCATCCTGCTGAGCGACATCCTGAGAG
TGAACACCGAGATCACCAAGGCCCCCCTGAGCGCCTCTATGATCAAGAGATACGACGAGCACCAC
CAGGACCTGACCCTGCTGAAAGCTCTCGTGCGGCAGCAGCTGCCTGAGAAGTACAAAGAGATTTT
CTTCGACCAGAGCAAGAACGGCTACGCCGGCTACATTGACGGCGGAGCCAGCCAGGAAGAGTTCT
ACAAGTTCATCAAGCCCATCCTGGAAAAGATGGACGGCACCGAGGAACTGCTCGTGAAGCTGAAG
AGAGAGGACCTGCTGCGGAAGCAGCGGACCTTCGACAACGGCAGCATCCCCCACCAGATCCACCT
GGGAGAGCTGCACGCCATTCTGCGGCGGCAGGAAGATTTTTACCCATTCCTGAAGGACAACCGGG
AAAAGATCGAGAAGATCCTGACCTTCCGCATCCCCTACTACGTGGGCCCTCTGGCCAGGGGAAAC
AGCAGATTCGCCTGGATGACCAGAAAGAGCGAGGAAACCATCACCCCCTGGAACTTCGAGGAAGT
GGTGGACAAGGGCGCTTCCGCCCAGAGCTTCATCGAGCGGATGACCAACTTCGATAAGAACCTGC
CCAACGAGAAGGTGCTGCCCAAGCACAGCCTGCTGTACGAGTACTTCACCGTGTATAACGAGCTG
ACCAAAGTGAAATACGTGACCGAGGGAATGAGAAAGCCCGCCTTCCTGAGCGGCGAGCAGAAAA
AGGCCATCGTGGACCTGCTGTTCAAGACCAACCGGAAAGTGACCGTGAAGCAGCTGAAAGAGGAC
TACTTCAAGAAAATCGAGTGCTTCGACTCCGTGGAAATCTCCGGCGTGGAAGATCGGTTCAACGCC
TCCCTGGGCACATACCACGATCTGCTGAAAATTATCAAGGACAAGGACTTCCTGGACAATGAGGA
AAACGAGGACATTCTGGAAGATATCGTGCTGACCCTGACACTGTTTGAGGACAGAGAGATGATCG
AGGAACGGCTGAAAACCTATGCCCACCTGTTCGACGACAAAGTGATGAAGCAGCTGAAGCGGCGG
AGATACACCGGCTGGGGCAGGCTGAGCCGGAAGCTGATCAACGGCATCCGGGACAAGCAGTCCG
GCAAGACAATCCTGGATTTCCTGAAGTCCGACGGCTTCGCCAACAGAAACTTCATGCAGCTGATCC
ACGACGACAGCCTGACCTTTAAAGAGGACATCCAGAAAGCCCAGGTGTCCGGCCAGGGCGATAGC
CTGCACGAGCACATTGCCAATCTGGCCGGCAGCCCCGCCATTAAGAAGGGCATCCTGCAGACAGT
GAAGGTGGTGGACGAGCTCGTGAAAGTGATGGGCCGGCACAAGCCCGAGAACATCGTGATCGAA
ATGGCCAGAGAGAACCAGACCACCCAGAAGGGACAGAAGAACAGCCGCGAGAGAATGAAGCGG
ATCGAAGAGGGCATCAAAGAGCTGGGCAGCCAGATCCTGAAAGAACACCCCGTGGAAAACACCC
AGCTGCAGAACGAGAAGCTGTACCTGTACTACCTGCAGAATGGGCGGGATATGTACGTGGACCAG
GAACTGGACATCAACCGGCTGTCCGACTACGATGTGGACGCTATCGTGCCTCAGAGCTTTCTGAAG
GACGACTCCATCGACAACAAGGTGCTGACCAGAAGCGACAAGAACCGGGGCAAGAGCGACAACG
TGCCCTCCGAAGAGGTCGTGAAGAAGATGAAGAACTACTGGCGGCAGCTGCTGAACGCCAAGCTG
ATTACCCAGAGAAAGTTCGACAATCTGACCAAGGCCGAGAGAGGCGGCCTGAGCGAACTGGATAA
GGCCGGCTTCATCAAGAGACAGCTGGTGGAAACCCGGCAGATCACAAAGCACGTGGCACAGATCC
TGGACTCCCGGATGAACACTAAGTACGACGAGAATGACAAGCTGATCCGGGAAGTGAAAGTGATC
ACCCTGAAGTCCAAGCTGGTGTCCGATTTCCGGAAGGATTTCCAGTTTTACAAAGTGCGCGAGATC
AACAACTACCACCACGCCCACGACGCCTACCTGAACGCCGTCGTGGGAACCGCCCTGATCAAAAA
GTACCCTAAGCTGGAAAGCGAGTTCGTGTACGGCGACTACAAGGTGTACGACGTGCGGAAGATGA
TCGCCAAGAGCGAGCAGGAAATCGGCAAGGCTACCGCCAAGTACTTCTTCTACAGCAACATCATG
AACTTTTTCAAGACCGAGATTACCCTGGCCAACGGCGAGATCCGGAAGCGGCCTCTGATCGAGAC
AAACGGCGAAACCGGGGAGATCGTGTGGGATAAGGGCCGGGATTTTGCCACCGTGCGGAAAGTGC
TGAGCATGCCCCAAGTGAATATCGTGAAAAAGACCGAGGTGCAGACAGGCGGCTTCAGCAAAGA
GTCTATCCTGCCCAAGAGGAACAGCGATAAGCTGATCGCCAGAAAGAAGGACTGGGACCCTAAGA
AGTACGGCGGCTTCGACAGCCCCACCGTGGCCTATTCTGTGCTGGTGGTGGCCAAAGTGGAAAAG
GGCAAGTCCAAGAAACTGAAGAGTGTGAAAGAGCTGCTGGGGATCACCATCATGGAAAGAAGCA
GCTTCGAGAAGAATCCCATCGACTTTCTGGAAGCCAAGGGCTACAAAGAAGTGAAAAAGGACCTG
ATCATCAAGCTGCCTAAGTACTCCCTGTTCGAGCTGGAAAACGGCCGGAAGAGAATGCTGGCCTCT
GCCGGCGAACTGCAGAAGGGAAACGAACTGGCCCTGCCCTCCAAATATGTGAACTTCCTGTACCT
GGCCAGCCACTATGAGAAGCTGAAGGGCTCCCCCGAGGATAATGAGCAGAAACAGCTGTTTGTGG
AACAGCACAAGCACTACCTGGACGAGATCATCGAGCAGATCAGCGAGTTCTCCAAGAGAGTGATC
CTGGCCGACGCTAATCTGGACAAAGTGCTGTCCGCCTACAACAAGCACCGGGATAAGCCCATCAG
AGAGCAGGCCGAGAATATCATCCACCTGTTTACCCTGACCAATCTGGGAGCCCCTGCCGCCTTCAA
GTACTTTGACACCACCATCGACCGGAAGAGGTACACCAGCACCAAAGAGGTGCTGGACGCCACCC
TGATCCACCAGAGCATCACCGGCCTGTACGAGACACGGATCGACCTGTCTCAGCTGGGAGGTGAC
TCCGGCGGAAGCTCTGGTGGCAGCAAGCGGACCGCCGACGGCTCTGAATTCGAGAGCCCTAAGAA
GAAAAGAAAGGTGAGCGGAGGCTCTAGCGGCGGAAGCATGGCTGAAAATGGTGATAATGAAAAG
ATGGCTGCCCTGGAGGCCAAAATCTGTCATCAAATTGAGTATTATTTTGGCGACTTCAATTTGCCA
CGGGACAAGTTTCTAAAGGAACAGATAAAACTGGATGAAGGCTGGGTACCTTTGGAGATAATGAT
AAAATTCAACAGGTTGAACCGTCTAACAACAGACTTTAATGTAATTGTGGAAGCATTGAGCAAAT
CCAAGGCAGAACTCATGGAAATCAGTGAAGATAAAACTAAAATCAGAAGGTCTCCAAGCAAACCC
CTACCTGAAGTGACTGATGAGTATAAAAATGATGTAAAAAACAGATCTGTTTATATTAAAGGCTTC
CCAACTGATGCAACTCTTGATGACATAAAAGAATGGTTAGAAGATAAAGGTCAAGTACTAAATAT
TCAGATGAGAAGAACATTGCATAAAGCATTTAAGGGATCAATTTTTGTTGTGTTTGATAGCATTGA
ATCTGCTAAGAAATTTGTAGAGACCCCTGGCCAGAAGTACAAAGAAACAGACCTGCTAATACTTTT
CAAGGACGATTACTTTGCCAAAAAAAATGAAGAAAGAAAACAAAATAAAGTGGAAGCTAAATTA
AGAGCTAAACAGGAGCAAGAAGCAAAACAAAAGTTAGAAGAAGATGCTGAAATGAAATCTCTAG
AAGAAAAGATTGGATGCTTGCTGAAATTTTCGGGTGATTTAGATGATCAGACCTGTAGAGAAGATT
TACACATACTTTTCTCAAATCATGGTGAAATAAAATGGATAGACTTCGTCAGAGGAGCAAAAGAG
GGGATAATTCTATTTAAAGAAAAAGCCAAGGAAGCATTGGGTAAAGCCAAAGATGCAAATAATGG
TAACCTACAATTAAGGAACAAAGAAGTGACTTGGGAAGTACTAGAAGGAGAGGTGGAAAAAGAA
GCACTGAAGAAAATAATAGAAGACCAACAAGAATCCCTAAACAAATGGAAGTCAAAAGGTCGTA
GATTTAAAGGAAAAGGAAAGGGTAATAAAGCTGCCCAGCCTGGGTCTGGTAAAGGAAAAGTACA
GTTTCAGGGCAAGAAAACGAAATTTGCTAGTGATGATGAACATGATGAACATGATGAAAATGGTG
CAACTGGACCTGTGAAAAGAGCAAGAGAAGAAACAGACAAAGAAGAACCTGCATCCAAACAACA
GAAAACAGAAAATGGTGCTGGAGACCAGTCCGGAGGATCTAGCGGAGGCTCCTCTGGCTCTGAGA
CACCTGGCACAAGCGAGAGCGCAACACCTGAAAGCAGCGGGGGCAGCAGCGGGGGGTCAACCCT
GAACATTGAAGACGAGTATAGACTGCATGAAACAAGCAAGGAACCCGACGTGTCCCTGGGCTCCA
CCTGGCTGTCCGACTTTCCCCAGGCCTGGGCCGAGACAGGAGGAATGGGCCTGGCCGTGCGGCAG
GCACCCCTGATCATCCCTCTGAAGGCCACCTCTACACCCGTGAGCATCAAGCAGTACCCTATGTCT
CAGGAGGCCAGACTGGGCATCAAGCCTCACATCCAGAGGCTGCTGGACCAGGGCATCCTGGTGCC
ATGCCAGAGCCCCTGGAACACACCACTGCTGCCCGTGAAGAAGCCAGGCACCAATGACTATAGAC
CCGTGCAGGATCTGAGAGAGGTGAACAAGAGGGTGGAGGATATCCACCCCACCGTGCCCAACCCT
TACAATCTGCTGTCCGGCCTGCCCCCTTCTCACCAGTGGTATACAGTGCTGGACCTGAAGGATGCC
TTCTTTTGTCTGAGACTGCACCCTACCAGCCAGCCACTGTTCGCCTTTGAGTGGAGGGACCCTGAG
ATGGGCATCTCTGGCCAGCTGACCTGGACACGCCTGCCTCAGGGCTTCAAGAATAGCCCAACACTG
TTTAACGAGGCCCTGCACCGCGACCTGGCAGATTTCCGGATCCAGCACCCAGATCTGATCCTGCTG
CAGTACGTGGACGATCTGCTGCTGGCCGCCACCAGCGAGCTGGATTGCCAGCAGGGAACACGCGC
CCTGCTGCAGACCCTGGGAAACCTGGGATATAGGGCATCCGCCAAGAAGGCCCAGATCTGTCAGA
AGCAGGTGAAGTACCTGGGCTATCTGCTGAAGGAGGGCCAGAGATGGCTGACAGAGGCCAGGAA
GGAGACAGTGATGGGCCAGCCAACACCCAAGACCCCAAGACAGCTGAGGGAGTTCCTGGGCAAA
GCAGGATTTTGCAGGCTGTTCATCCCAGGATTCGCAGAGATGGCAGCACCTCTGTACCCACTGACC
AAGCCGGGCACCCTGTTTAATTGGGGCCCTGACCAGCAGAAGGCCTATCAGGAGATCAAGCAGGC
CCTGCTGACAGCACCAGCCCTGGGCCTGCCAGACCTGACCAAGCCTTTCGAGCTGTTTGTGGATGA
GAAGCAGGGCTACGCCAAGGGCGTGCTGACCCAGAAGCTGGGACCATGGAGACGGCCCGTGGCCT
ATCTGTCCAAGAAGCTGGACCCAGTGGCAGCAGGATGGCCACCATGCCTGAGGATGGTGGCAGCA
ATCGCCGTGCTGACAAAGGATGCCGGCAAGCTGACCATGGGACAGCCACTGGTCATCCTGGCACC
ACACGCAGTGGAGGCCCTGGTGAAGCAGCCTCCAGATCGCTGGCTGTCTAACGCCCGGATGACAC
ACTACCAGGCCCTGCTGCTGGACACCGATCGCGTGCAGTTTGGCCCTGTGGTGGCCCTGAATCCAG
CCACCCTGCTGCCTCTGCCAGAGGAGGGCCTGCAGCACAACTGTCTGGACATCCTGGCAGAGGCA
CACGGAACAAGGCCAGACCTGACCGATCAGCCCCTGCCTGACGCCGATCACACATGGTATACCGA
TGGAAGCTCCCTGCTGCAGGAGGGCCAGAGGAAGGCAGGAGCAGCAGTGACCACAGAGACAGAA
GTGATCTGGGCCAAGGCCCTGCCAGCAGGCACATCCGCCCAGCGGGCCGAGCTGATCGCCCTGAC
CCAGGCCCTGAAGATGGCCGAGGGCAAGAAGCTGAACGTGTACACAGACTCCAGATATGCCTTCG
CCACCGCACACATCCACGGAGAGATCTACAGGCGCCGGGGCTGGCTGACCTCTGAGGGCAAGGAG
ATCAAGAACAAGGATGAGATCCTGGCCCTGCTGAAGGCCCTGTTTCTGCCCAAGCGGCTGAGCAT
CATCCACTGTCCTGGACACCAGAAGGGACACTCCGCCGAGGCAAGGGGCAATCGGATGGCCGACC
AGGCCGCCAGAAAGGCTGCTATTACTGAAACTCCCGACACTTCCACTCTGCTGATTGAAAACTCCT
CCCCTTCTGGCGGCTCAAAAAGAACCGCCGACGGCAGCGAATTCGAGTCTCCCAAGAAGAAGAGG
AAAGTCGGCTCTGGCCCTGCCGCTAAGAGAGTGAAGCTGGACTAA
SEQ ID NO: 25 (bpNLSSV40) MKRTADGSEFESPKKKRKV
SEQ ID NO: 26 (Cas9 H840A nickase)
DKKYSIGLDIGTNSVGWAVITDEYKVPSKKFKVLGNTDRHSIKKNLIGALLFDSGETAEATRLKRTARR
RYTRRKNRICYLQEIFSNEMAKVDDSFFHRLEESFLVEEDKKHERHPIFGNIVDEVAYHEKYPTIYHLRK
KLVDSTDKADLRLIYLALAHMIKFRGHFLIEGDLNPDNSDVDKLFIQLVQTYNQLFEENPINASGVDAK
AILSARLSKSRRLENLIAQLPGEKKNGLFGNLIALSLGLTPNFKSNFDLAEDAKLQLSKDTYDDDLDNLL
AQIGDQYADLFLAAKNLSDAILLSDILRVNTEITKAPLSASMIKRYDEHHQDLTLLKALVRQQLPEKYK
EIFFDQSKNGYAGYIDGGASQEEFYKFIKPILEKMDGTEELLVKLNREDLLRKQRTFDNGSIPHQIHLGE
LHAILRRQEDFYPFLKDNREKIEKILTFRIPYYVGPLARGNSRFAWMTRKSEETITPWNFEEWDKGASA
QSFIERMTNFDKNLPNEKVLPKHSLLYEYFTVYNELTKVKYVTEGMRKPAFLSGEQKKAIVDLLFKTN
RKVTVKQLKEDYFKKIECFDSVEISGVEDRFNASLGTYHDLLKIIKDKDFLDNEENEDILEDIVLTLTLFE
DREMIEERLKTYAHLFDDKVMKQLKRRRYTGWGRLSRKLINGIRDKQSGKTILDFLKSDGFANRNFM
QLIHDDSLTFKEDIQKAQVSGQGDSLHEHIANLAGSPAIKKGILQTVKWDELVKVMGRHKPENIVIEM
ARENQTTQKGQKNSRERMKRIEEGIKELGSQILKEHPVENTQLQNEKLYLYYLQNGRDMYVDQELDIN
RLSDYDVDAIVPQSFLKDDSIDNKVLTRSDKNRGKSDNVPSEEWKKMKNYWRQLLNAKLITQRKFD
NLTKAERGGLSELDKAGFIKRQLVETRQITKHVAQILDSRMNTKYDENDKLIREVKVITLKSKLVSDFR
KDFQFYKVREINNYHHAHDAYLNAWGTALIKKYPKLESEFVYGDYKVYDVRKMIAKSEQEIGKATA
KYFFYSNIMNFFKTEITLANGEIRKRPLIETNGETGEIVWDKGRDFATVRKVLSMPQVNIVKKTEVQTG
GFSKESILPKRNSDKLIARKKDWDPKKYGGFDSPTVAYSVLWAKVEKGKSKKLKSVKELLGITIMERS
SFEKNPIDFLEAKGYKEVKKDLIIKLPKYSLFELENGRKRMLASAGELQKGNELALPSKYVNFLYLASH
YEKLKGSPEDNEQKQLFVEQHKHYLDEIIEQISEFSKRVILADANLDKVLSAYNKHRDKPIREQAENIIH
LFTLTNLGAPAAFKYFDTTIDRKRYTSTKEVLDATLIHQSITGLYETRIDLSQLGGD
SEQ ID NO: 27 (Cas9 R221K N394K H840A nickase)
DKKYSIGLDIGTNSVGWAVITDEYKVPSKKFKVLGNTDRHSIKKNLIGALLFDSGETAEATRLKRTARR
RYTRRKNRICYLQEIFSNEMAKVDDSFFHRLEESFLVEEDKKHERHPIFGNIVDEVAYHEKYPTIYHLRK
KLVDSTDKADLRLIYLALAHMIKFRGHFLIEGDLNPDNSDVDKLFIQLVQTYNQLFEENPINASGVDAK
AILSARLSKSRKLENLIAQLPGEKKNGLFGNLIALSLGLTPNFKSNFDLAEDAKLQLSKDTYDDDLDNLL
AQIGDQYADLFLAAKNLSDAILLSDILRVNTEITKAPLSASMIKRYDEHHQDLTLLKALVRQQLPEKYK
EIFFDQSKNGYAGYIDGGASQEEFYKFIKPILEKMDGTEELLVKLKREDLLRKQRTFDNGSIPHQIHLGE
LHAILRRQEDFYPFLKDNREKIEKILTFRIPYYVGPLARGNSRFAWMTRKSEETITPWNFEEWDKGASA
QSFIERMTNFDKNLPNEKVLPKHSLLYEYFTVYNELTKVKYVTEGMRKPAFLSGEQKKAIVDLLFKTN
RKVTVKQLKEDYFKKIECFDSVEISGVEDRFNASLGTYHDLLKIIKDKDFLDNEENEDILEDIVLTLTLFE
DREMIEERLKTYAHLFDDKVMKQLKRRRYTGWGRLSRKLINGIRDKQSGKTILDFLKSDGFANRNFM
QLIHDDSLTFKEDIQKAQVSGQGDSLHEHIANLAGSPAIKKGILQTVKWDELVKVMGRHKPENIVIEM
ARENQTTQKGQKNSRERMKRIEEGIKELGSQILKEHPVENTQLQNEKLYLYYLQNGRDMYVDQELDIN
RLSDYDVDAIVPQSFLKDDSIDNKVLTRSDKNRGKSDNVPSEEWKKMKNYWRQLLNAKLITQRKFD
NLTKAERGGLSELDKAGFIKRQLVETRQITKHVAQILDSRMNTKYDENDKLIREVKVITLKSKLVSDFR
KDFQFYKVREINNYHHAHDAYLNAWGTALIKKYPKLESEFVYGDYKVYDVRKMIAKSEQEIGKATA
KYFFYSNIMNFFKTEITLANGEIRKRPLIETNGETGEIVWDKGRDFATVRKVLSMPQVNIVKKTEVQTG
GFSKESILPKRNSDKLIARKKDWDPKKYGGFDSPTVAYSVLWAKVEKGKSKKLKSVKELLGITIMERS
SFEKNPIDFLEAKGYKEVKKDLIIKLPKYSLFELENGRKRMLASAGELQKGNELALPSKYVNFLYLASH
YEKLKGSPEDNEQKQLFVEQHKHYLDEIIEQISEFSKRVILADANLDKVLSAYNKHRDKPIREQAENIIH
LFTLTNLGAPAAFKYFDTTIDRKRYTSTKEVLDATLIHQSITGLYETRIDLSQLGGD
SEQ ID NO: 28, linker A* (SGGS2x-XTEN16-SGGS2x)
SGGSSGGSSGSETPGTSESATPESSGGSSGGSS
SEQ ID NO: 29, linker A (SGGS2x-XTEN16-SGGS2x)
SGGSSGGSSGSETPGTSESATPESSGGSSGGS
SEQ ID NO: 30, SGGS linker
SGGS
SEQ ID NO: 31, MMLV-RT
TLNIEDEYRLHETSKEPDVSLGSTWLSDFPQAWAETGGMGLAVRQAPLIIPLKATSTPVSIKQYPMSQE
ARLGIKPHIQRLLDQGILVPCQSPWNTPLLPVKKPGTNDYRPVQDLREVNKRVEDIHPTVPNPYNLLSG
LPPSHQWYTVLDLKDAFFCLRLHPTSQPLFAFEWRDPEMGISGQLTWTRLPQGFKNSPTLFNEALHRDL
ADFRIQHPDLILLQYVDDLLLAATSELDCQQGTRALLQTLGNLGYRASAKKAQICQKQVKYLGYLLKE
GQRWLTEARKETVMGQPTPKTPRQLREFLGKAGFCRLFIPGFAEMAAPLYPLTKPGTLFNWGPDQQKA
YQEIKQALLTAPALGLPDLTKPFELFVDEKQGYAKGVLTQKLGPWRRPVAYLSKKLDPVAAGWPPCL
RMVAAIAVLTKDAGKLTMGQPLVILAPHAVEALVKQPPDRWLSNARMTHYQALLLDTDRVQFGPVV
ALNPATLLPLPEEGLQHNCLDILAEAHGTRPDLTDQPLPDADHTWYTDGSSLLQEGQRKAGAAVTTET
EVIWAKALPAGTSAQRAELIALTQALKMAEGKKLNVYTDSRYAFATAHIHGEIYRRRGWLTSEGKEIK
NKDEILALLKALFLPKRLSIIHCPGHQKGHSAEARGNRMADQAARKAAITETPDTSTLLIENSSP
SEQ ID NO: 32, human codon optimized MMLV-RT
TLNIEDEYRLHETSKEPDVSLGSTWLSDFPQAWAETGGMGLAVRQAPLIIPLKATSTPVSIKQYPMSQE
ARLGIKPHIQRLLDQGILVPCQSPWNTPLLPVKKPGTNDYRPVQDLREVNKRVEDIHPTVPNPYNLLSG
LPPSHQWYTVLDLKDAFFCLRLHPTSQPLFAFEWRDPEMGISGQLTWTRLPQGFKNSPTLFNEALHRDL
ADFRIQHPDLILLQYVDDLLLAATSELDCQQGTRALLQTLGNLGYRASAKKAQICQKQVKYLGYLLKE
GQRWLTEARKETVMGQPTPKTPRQLREFLGKAGFCRLFIPGFAEMAAPLYPLTKPGTLFNWGPDQQKA
YQEIKQALLTAPALGLPDLTKPFELFVDEKQGYAKGVLTQKLGPWRRPVAYLSKKLDPVAAGWPPCL
RMVAAIAVLTKDAGKLTMGQPLVILAPHAVEALVKQPPDRWLSNARMTHYQALLLDTDRVQFGPVV
ALNPATLLPLPEEGLQHNCLDILAEAHGTRPDLTDQPLPDADHTWYTDGSSLLQEGQRKAGAAVTTET
EVIWAKALPAGTSAQRAELIALTQALKMAEGKKLNVYTDSRYAFATAHIHGEIYRRRGWLTSEGKEIK
NKDEILALLKALFLPKRLSIIHCPGHQKGHSAEARGNRMADQAARKAAITETPDTSTLLIENSSP
SEQ ID NO: 33 (SSB1 194)
MAENGDNEKMAALEAKICHQIEYYFGDFNLPRDKFLKEQIKLDEGWVPLEIMIKFNRLNRLTTDFNVIV
EALSKSKAELMEISEDKTKIRRSPSKPLPEVTDEYKNDVKNRSVYIKGFPTDATLDDIKEWLEDKGQVL
NIQMRRTLHKAFKGSIFWFDSIESAKKFVETPGQKYKETDLLILFKDDYFAKKNE
SEQ ID NO: 34 (SSB)
MAENGDNEKMAALEAKICHQIEYYFGDFNLPRDKFLKEQIKLDEGWVPLEIMIKFNRLNRLTTDFNVIV
EALSKSKAELMEISEDKTKIRRSPSKPLPEVTDEYKNDVKNRSVYIKGFPTDATLDDIKEWLEDKGQVL
NIQMRRTLHKAFKGSIFWFDSIESAKKFVETPGQKYKETDLLILFKDDYFAKKNEERKQNKVEAKLRA
KQEQEAKQKLEEDAEMKSLEEKIGCLLKFSGDLDDQTCREDLHILFSNHGEIKWIDFVRGAKEGIILFKE KAKEALGKAKDANNGNLQLRNKEVTWEVLEGEVEKEALKKIIEDQQESLNKWKSKGRRFKGKGKGN KAAQPGSGKGKVQFQGKKTKFASDDEHDEHDENGATGPVKRAREETDKEEPASKQQKTENGAGDQ
SEQ ID NO: 35 (SSB2-194)
AENGDNEKMAALEAKICHQIEYYFGDFNLPRDKFLKEQIKLDEGWVPLEIMIKFNRLNRLTTDFNVIVE ALSKSKAELMEISEDKTKIRRSPSKPLPEVTDEYKNDVKNRSVYIKGFPTDATLDDIKEWLEDKGQVLNI QMRRTLHKAFKGSIFWFDSIESAKKFVETPGQKYKETDLLILFKDDYFAKKNE
SEQ ID NO: 36 (SSB2-408)
AENGDNEKMAALEAKICHQIEYYFGDFNLPRDKFLKEQIKLDEGWVPLEIMIKFNRLNRLTTDFNVIVE ALSKSKAELMEISEDKTKIRRSPSKPLPEVTDEYKNDVKNRSVYIKGFPTDATLDDIKEWLEDKGQVLNI QMRRTLHKAFKGSIFWFDSIESAKKFVETPGQKYKETDLLILFKDDYFAKKNEERKQNKVEAKLRAK QEQEAKQKLEEDAEMKSLEEKIGCLLKFSGDLDDQTCREDLHILFSNHGEIKWIDFVRGAKEGIILFKEK AKEALGKAKDANNGNLQLRNKEVTWEVLEGEVEKEALKKIIEDQQESLNKWKSKGRRFKGKGKGNK AAQPGSGKGKVQFQGKKTKFASDDEHDEHDENGATGPVKRAREETDKEEPASKQQKTENGAGDQ
SEQ ID NO: 37 (bpNLSSV4°)
KRTADGSEFEPKKKRKV
SEQ ID NO: 38 (bpNLSSV40)
KRTADGSEFESPKKKRKV
SEQ ID NO: 39 (NLSc Myc)
GSGPAAKRVKLD
SEQ ID NO: 40 linker B (SGGSx2-NLSSV40-SGGSx2)
SGGSSGGSKRTADGSEFESPKKKRKVSGGSSGGS
SEQ ID NO: 41 (bpNLSSV40), DNA
ATGAAACGGACAGCCGACGGAAGCGAGTTCGAGTCACCAAAGAAGAAGCGGAAAGTC
SEQ ID NO: 42 (Cas9 H840A nickase), DNA
GACAAGAAGTACAGCATCGGCCTGGACATCGGCACCAACTCTGTGGGCTGGGCCGTGATCACCGA
CGAGTACAAGGTGCCCAGCAAGAAATTCAAGGTGCTGGGCAACACCGACCGGCACAGCATCAAG
AAGAACCTGATCGGAGCCCTGCTGTTCGACAGCGGCGAAACAGCCGAGGCCACCCGGCTGAAGAG
AACCGCCAGAAGAAGATACACCAGACGGAAGAACCGGATCTGCTATCTGCAAGAGATCTTCAGCA
ACGAGATGGCCAAGGTGGACGACAGCTTCTTCCACAGACTGGAAGAGTCCTTCCTGGTGGAAGAG
GATAAGAAGCACGAGCGGCACCCCATCTTCGGCAACATCGTGGACGAGGTGGCCTACCACGAGAA
GTACCCCACCATCTACCACCTGAGAAAGAAACTGGTGGACAGCACCGACAAGGCCGACCTGCGGC
TGATCTATCTGGCCCTGGCCCACATGATCAAGTTCCGGGGCCACTTCCTGATCGAGGGCGACCTGA
ACCCCGACAACAGCGACGTGGACAAGCTGTTCATCCAGCTGGTGCAGACCTACAACCAGCTGTTC
GAGGAAAACCCCATCAACGCCAGCGGCGTGGACGCCAAGGCCATCCTGTCTGCCAGACTGAGCAA
GAGCAGACGGCTGGAAAATCTGATCGCCCAGCTGCCCGGCGAGAAGAAGAATGGCCTGTTCGGAA
ACCTGATTGCCCTGAGCCTGGGCCTGACCCCCAACTTCAAGAGCAACTTCGACCTGGCCGAGGATG
CCAAACTGCAGCTGAGCAAGGACACCTACGACGACGACCTGGACAACCTGCTGGCCCAGATCGGC
GACCAGTACGCCGACCTGTTTCTGGCCGCCAAGAACCTGTCCGACGCCATCCTGCTGAGCGACATC
CTGAGAGTGAACACCGAGATCACCAAGGCCCCCCTGAGCGCCTCTATGATCAAGAGATACGACGA
GCACCACCAGGACCTGACCCTGCTGAAAGCTCTCGTGCGGCAGCAGCTGCCTGAGAAGTACAAAG
AGATTTTCTTCGACCAGAGCAAGAACGGCTACGCCGGCTACATTGACGGCGGAGCCAGCCAGGAA
GAGTTCTACAAGTTCATCAAGCCCATCCTGGAAAAGATGGACGGCACCGAGGAACTGCTCGTGAA
GCTGAACAGAGAGGACCTGCTGCGGAAGCAGCGGACCTTCGACAACGGCAGCATCCCCCACCAGA
TCCACCTGGGAGAGCTGCACGCCATTCTGCGGCGGCAGGAAGATTTTTACCCATTCCTGAAGGACA
ACCGGGAAAAGATCGAGAAGATCCTGACCTTCCGCATCCCCTACTACGTGGGCCCTCTGGCCAGG
GGAAACAGCAGATTCGCCTGGATGACCAGAAAGAGCGAGGAAACCATCACCCCCTGGAACTTCGA
GGAAGTGGTGGACAAGGGCGCTTCCGCCCAGAGCTTCATCGAGCGGATGACCAACTTCGATAAGA
ACCTGCCCAACGAGAAGGTGCTGCCCAAGCACAGCCTGCTGTACGAGTACTTCACCGTGTATAAC
GAGCTGACCAAAGTGAAATACGTGACCGAGGGAATGAGAAAGCCCGCCTTCCTGAGCGGCGAGC
AGAAAAAGGCCATCGTGGACCTGCTGTTCAAGACCAACCGGAAAGTGACCGTGAAGCAGCTGAAA
GAGGACTACTTCAAGAAAATCGAGTGCTTCGACTCCGTGGAAATCTCCGGCGTGGAAGATCGGTT
CAACGCCTCCCTGGGCACATACCACGATCTGCTGAAAATTATCAAGGACAAGGACTTCCTGGACA
ATGAGGAAAACGAGGACATTCTGGAAGATATCGTGCTGACCCTGACACTGTTTGAGGACAGAGAG
ATGATCGAGGAACGGCTGAAAACCTATGCCCACCTGTTCGACGACAAAGTGATGAAGCAGCTGAA
GCGGCGGAGATACACCGGCTGGGGCAGGCTGAGCCGGAAGCTGATCAACGGCATCCGGGACAAG
CAGTCCGGCAAGACAATCCTGGATTTCCTGAAGTCCGACGGCTTCGCCAACAGAAACTTCATGCAG
CTGATCCACGACGACAGCCTGACCTTTAAAGAGGACATCCAGAAAGCCCAGGTGTCCGGCCAGGG
CGATAGCCTGCACGAGCACATTGCCAATCTGGCCGGCAGCCCCGCCATTAAGAAGGGCATCCTGC
AGACAGTGAAGGTGGTGGACGAGCTCGTGAAAGTGATGGGCCGGCACAAGCCCGAGAACATCGT
GATCGAAATGGCCAGAGAGAACCAGACCACCCAGAAGGGACAGAAGAACAGCCGCGAGAGAATG
AAGCGGATCGAAGAGGGCATCAAAGAGCTGGGCAGCCAGATCCTGAAAGAACACCCCGTGGAAA
ACACCCAGCTGCAGAACGAGAAGCTGTACCTGTACTACCTGCAGAATGGGCGGGATATGTACGTG
GACCAGGAACTGGACATCAACCGGCTGTCCGACTACGATGTGGACGCTATCGTGCCTCAGAGCTTT
CTGAAGGACGACTCCATCGACAACAAGGTGCTGACCAGAAGCGACAAGAACCGGGGCAAGAGCG
ACAACGTGCCCTCCGAAGAGGTCGTGAAGAAGATGAAGAACTACTGGCGGCAGCTGCTGAACGCC
AAGCTGATTACCCAGAGAAAGTTCGACAATCTGACCAAGGCCGAGAGAGGCGGCCTGAGCGAACT
GGATAAGGCCGGCTTCATCAAGAGACAGCTGGTGGAAACCCGGCAGATCACAAAGCACGTGGCAC
AGATCCTGGACTCCCGGATGAACACTAAGTACGACGAGAATGACAAGCTGATCCGGGAAGTGAAA
GTGATCACCCTGAAGTCCAAGCTGGTGTCCGATTTCCGGAAGGATTTCCAGTTTTACAAAGTGCGC
GAGATCAACAACTACCACCACGCCCACGACGCCTACCTGAACGCCGTCGTGGGAACCGCCCTGAT
CAAAAAGTACCCTAAGCTGGAAAGCGAGTTCGTGTACGGCGACTACAAGGTGTACGACGTGCGGA
AGATGATCGCCAAGAGCGAGCAGGAAATCGGCAAGGCTACCGCCAAGTACTTCTTCTACAGCAAC
ATCATGAACTTTTTCAAGACCGAGATTACCCTGGCCAACGGCGAGATCCGGAAGCGGCCTCTGATC
GAGACAAACGGCGAAACCGGGGAGATCGTGTGGGATAAGGGCCGGGATTTTGCCACCGTGCGGA
AAGTGCTGAGCATGCCCCAAGTGAATATCGTGAAAAAGACCGAGGTGCAGACAGGCGGCTTCAGC
AAAGAGTCTATCCTGCCCAAGAGGAACAGCGATAAGCTGATCGCCAGAAAGAAGGACTGGGACC
CTAAGAAGTACGGCGGCTTCGACAGCCCCACCGTGGCCTATTCTGTGCTGGTGGTGGCCAAAGTGG
AAAAGGGCAAGTCCAAGAAACTGAAGAGTGTGAAAGAGCTGCTGGGGATCACCATCATGGAAAG
AAGCAGCTTCGAGAAGAATCCCATCGACTTTCTGGAAGCCAAGGGCTACAAAGAAGTGAAAAAGG
ACCTGATCATCAAGCTGCCTAAGTACTCCCTGTTCGAGCTGGAAAACGGCCGGAAGAGAATGCTG
GCCTCTGCCGGCGAACTGCAGAAGGGAAACGAACTGGCCCTGCCCTCCAAATATGTGAACTTCCT
GTACCTGGCCAGCCACTATGAGAAGCTGAAGGGCTCCCCCGAGGATAATGAGCAGAAACAGCTGT
TTGTGGAACAGCACAAGCACTACCTGGACGAGATCATCGAGCAGATCAGCGAGTTCTCCAAGAGA
GTGATCCTGGCCGACGCTAATCTGGACAAAGTGCTGTCCGCCTACAACAAGCACCGGGATAAGCC
CATCAGAGAGCAGGCCGAGAATATCATCCACCTGTTTACCCTGACCAATCTGGGAGCCCCTGCCGC
CTTCAAGTACTTTGACACCACCATCGACCGGAAGAGGTACACCAGCACCAAAGAGGTGCTGGACG
CCACCCTGATCCACCAGAGCATCACCGGCCTGTACGAGACACGGATCGACCTGTCTCAGCTGGGA
GGTGAC
SEQ ID NO: 43 (Cas9 R221K N394K H840A nickase), DNA
GACAAGAAGTACAGCATCGGCCTGGACATCGGCACCAACTCTGTGGGCTGGGCCGTGATCACCGA
CGAGTACAAGGTGCCCAGCAAGAAATTCAAGGTGCTGGGCAACACCGACCGGCACAGCATCAAG
AAGAACCTGATCGGAGCCCTGCTGTTCGACAGCGGCGAAACAGCCGAGGCCACCCGGCTGAAGAG
AACCGCCAGAAGAAGATACACCAGACGGAAGAACCGGATCTGCTATCTGCAAGAGATCTTCAGCA
ACGAGATGGCCAAGGTGGACGACAGCTTCTTCCACAGACTGGAAGAGTCCTTCCTGGTGGAAGAG
GATAAGAAGCACGAGCGGCACCCCATCTTCGGCAACATCGTGGACGAGGTGGCCTACCACGAGAA
GTACCCCACCATCTACCACCTGAGAAAGAAACTGGTGGACAGCACCGACAAGGCCGACCTGCGGC
TGATCTATCTGGCCCTGGCCCACATGATCAAGTTCCGGGGCCACTTCCTGATCGAGGGCGACCTGA
ACCCCGACAACAGCGACGTGGACAAGCTGTTCATCCAGCTGGTGCAGACCTACAACCAGCTGTTC
GAGGAAAACCCCATCAACGCCAGCGGCGTGGACGCCAAGGCCATCCTGTCTGCCAGACTGAGCAA
GAGCAGAAAGCTGGAAAATCTGATCGCCCAGCTGCCCGGCGAGAAGAAGAATGGCCTGTTCGGAA
ACCTGATTGCCCTGAGCCTGGGCCTGACCCCCAACTTCAAGAGCAACTTCGACCTGGCCGAGGATG
CCAAACTGCAGCTGAGCAAGGACACCTACGACGACGACCTGGACAACCTGCTGGCCCAGATCGGC
GACCAGTACGCCGACCTGTTTCTGGCCGCCAAGAACCTGTCCGACGCCATCCTGCTGAGCGACATC
CTGAGAGTGAACACCGAGATCACCAAGGCCCCCCTGAGCGCCTCTATGATCAAGAGATACGACGA
GCACCACCAGGACCTGACCCTGCTGAAAGCTCTCGTGCGGCAGCAGCTGCCTGAGAAGTACAAAG
AGATTTTCTTCGACCAGAGCAAGAACGGCTACGCCGGCTACATTGACGGCGGAGCCAGCCAGGAA
GAGTTCTACAAGTTCATCAAGCCCATCCTGGAAAAGATGGACGGCACCGAGGAACTGCTCGTGAA
GCTGAAGAGAGAGGACCTGCTGCGGAAGCAGCGGACCTTCGACAACGGCAGCATCCCCCACCAGA
TCCACCTGGGAGAGCTGCACGCCATTCTGCGGCGGCAGGAAGATTTTTACCCATTCCTGAAGGACA
ACCGGGAAAAGATCGAGAAGATCCTGACCTTCCGCATCCCCTACTACGTGGGCCCTCTGGCCAGG
GGAAACAGCAGATTCGCCTGGATGACCAGAAAGAGCGAGGAAACCATCACCCCCTGGAACTTCGA
GGAAGTGGTGGACAAGGGCGCTTCCGCCCAGAGCTTCATCGAGCGGATGACCAACTTCGATAAGA
ACCTGCCCAACGAGAAGGTGCTGCCCAAGCACAGCCTGCTGTACGAGTACTTCACCGTGTATAAC
GAGCTGACCAAAGTGAAATACGTGACCGAGGGAATGAGAAAGCCCGCCTTCCTGAGCGGCGAGC
AGAAAAAGGCCATCGTGGACCTGCTGTTCAAGACCAACCGGAAAGTGACCGTGAAGCAGCTGAAA
GAGGACTACTTCAAGAAAATCGAGTGCTTCGACTCCGTGGAAATCTCCGGCGTGGAAGATCGGTT
CAACGCCTCCCTGGGCACATACCACGATCTGCTGAAAATTATCAAGGACAAGGACTTCCTGGACA
ATGAGGAAAACGAGGACATTCTGGAAGATATCGTGCTGACCCTGACACTGTTTGAGGACAGAGAG
ATGATCGAGGAACGGCTGAAAACCTATGCCCACCTGTTCGACGACAAAGTGATGAAGCAGCTGAA
GCGGCGGAGATACACCGGCTGGGGCAGGCTGAGCCGGAAGCTGATCAACGGCATCCGGGACAAG
CAGTCCGGCAAGACAATCCTGGATTTCCTGAAGTCCGACGGCTTCGCCAACAGAAACTTCATGCAG
CTGATCCACGACGACAGCCTGACCTTTAAAGAGGACATCCAGAAAGCCCAGGTGTCCGGCCAGGG
CGATAGCCTGCACGAGCACATTGCCAATCTGGCCGGCAGCCCCGCCATTAAGAAGGGCATCCTGC
AGACAGTGAAGGTGGTGGACGAGCTCGTGAAAGTGATGGGCCGGCACAAGCCCGAGAACATCGT
GATCGAAATGGCCAGAGAGAACCAGACCACCCAGAAGGGACAGAAGAACAGCCGCGAGAGAATG
AAGCGGATCGAAGAGGGCATCAAAGAGCTGGGCAGCCAGATCCTGAAAGAACACCCCGTGGAAA
ACACCCAGCTGCAGAACGAGAAGCTGTACCTGTACTACCTGCAGAATGGGCGGGATATGTACGTG
GACCAGGAACTGGACATCAACCGGCTGTCCGACTACGATGTGGACGCTATCGTGCCTCAGAGCTTT
CTGAAGGACGACTCCATCGACAACAAGGTGCTGACCAGAAGCGACAAGAACCGGGGCAAGAGCG
ACAACGTGCCCTCCGAAGAGGTCGTGAAGAAGATGAAGAACTACTGGCGGCAGCTGCTGAACGCC
AAGCTGATTACCCAGAGAAAGTTCGACAATCTGACCAAGGCCGAGAGAGGCGGCCTGAGCGAACT
GGATAAGGCCGGCTTCATCAAGAGACAGCTGGTGGAAACCCGGCAGATCACAAAGCACGTGGCAC
AGATCCTGGACTCCCGGATGAACACTAAGTACGACGAGAATGACAAGCTGATCCGGGAAGTGAAA
GTGATCACCCTGAAGTCCAAGCTGGTGTCCGATTTCCGGAAGGATTTCCAGTTTTACAAAGTGCGC
GAGATCAACAACTACCACCACGCCCACGACGCCTACCTGAACGCCGTCGTGGGAACCGCCCTGAT
CAAAAAGTACCCTAAGCTGGAAAGCGAGTTCGTGTACGGCGACTACAAGGTGTACGACGTGCGGA
AGATGATCGCCAAGAGCGAGCAGGAAATCGGCAAGGCTACCGCCAAGTACTTCTTCTACAGCAAC
ATCATGAACTTTTTCAAGACCGAGATTACCCTGGCCAACGGCGAGATCCGGAAGCGGCCTCTGATC
GAGACAAACGGCGAAACCGGGGAGATCGTGTGGGATAAGGGCCGGGATTTTGCCACCGTGCGGA
AAGTGCTGAGCATGCCCCAAGTGAATATCGTGAAAAAGACCGAGGTGCAGACAGGCGGCTTCAGC
AAAGAGTCTATCCTGCCCAAGAGGAACAGCGATAAGCTGATCGCCAGAAAGAAGGACTGGGACC
CTAAGAAGTACGGCGGCTTCGACAGCCCCACCGTGGCCTATTCTGTGCTGGTGGTGGCCAAAGTGG
AAAAGGGCAAGTCCAAGAAACTGAAGAGTGTGAAAGAGCTGCTGGGGATCACCATCATGGAAAG
AAGCAGCTTCGAGAAGAATCCCATCGACTTTCTGGAAGCCAAGGGCTACAAAGAAGTGAAAAAGG
ACCTGATCATCAAGCTGCCTAAGTACTCCCTGTTCGAGCTGGAAAACGGCCGGAAGAGAATGCTG
GCCTCTGCCGGCGAACTGCAGAAGGGAAACGAACTGGCCCTGCCCTCCAAATATGTGAACTTCCT
GTACCTGGCCAGCCACTATGAGAAGCTGAAGGGCTCCCCCGAGGATAATGAGCAGAAACAGCTGT
TTGTGGAACAGCACAAGCACTACCTGGACGAGATCATCGAGCAGATCAGCGAGTTCTCCAAGAGA
GTGATCCTGGCCGACGCTAATCTGGACAAAGTGCTGTCCGCCTACAACAAGCACCGGGATAAGCC
CATCAGAGAGCAGGCCGAGAATATCATCCACCTGTTTACCCTGACCAATCTGGGAGCCCCTGCCGC
CTTCAAGTACTTTGACACCACCATCGACCGGAAGAGGTACACCAGCACCAAAGAGGTGCTGGACG
CCACCCTGATCCACCAGAGCATCACCGGCCTGTACGAGACACGGATCGACCTGTCTCAGCTGGGA
GGTGAC
SEQ ID NO: 44, linker A* (SGGS2x-XTEN16-SGGS2x), DNA
TCTGGAGGATCTAGCGGAGGATCCTCTGGCAGCGAGACACCAGGAACAAGCGAGTCAGCAACACC
AGAGAGCAGTGGCGGCAGCAGCGGCGGCAGCAGC
SEQ ID NO: 45, linker A (SGGS2x-XTEN16-SGGS2x), DNA
TCCGGAGGATCTAGCGGAGGCTCCTCTGGCTCTGAGACACCTGGCACAAGCGAGAGCGCAACACC
TGAAAGCAGCGGGGGCAGCAGCGGGGGGTCA
SEQ ID NO: 46, SGGS linker, DNA
TCTGGCGGCTCA
SEQ ID NO: 47, MMLV-RT, DNA
ACCCTAAATATAGAAGATGAGTATCGGCTACATGAGACCTCAAAAGAGCCAGATGTTTCTCTAGG
GTCCACATGGCTGTCTGATTTTCCTCAGGCCTGGGCGGAAACCGGGGGCATGGGACTGGCAGTTCG
CCAAGCTCCTCTGATCATACCTCTGAAAGCAACCTCTACCCCCGTGTCCATAAAACAATACCCCAT
GTCACAAGAAGCCAGACTGGGGATCAAGCCCCACATACAGAGACTGTTGGACCAGGGAATACTGG
TACCCTGCCAGTCCCCCTGGAACACGCCCCTGCTACCCGTTAAGAAACCAGGGACTAATGATTATA
GGCCTGTCCAGGATCTGAGAGAAGTCAACAAGCGGGTGGAAGACATCCACCCCACCGTGCCCAAC
CCTTACAACCTCTTGAGCGGGCTCCCACCGTCCCACCAGTGGTACACTGTGCTTGATTTAAAGGAT
GCCTTTTTCTGCCTGAGACTCCACCCCACCAGTCAGCCTCTCTTCGCCTTTGAGTGGAGAGATCCAG
AGATGGGAATCTCAGGACAATTGACCTGGACCAGACTCCCACAGGGTTTCAAAAACAGTCCCACC
CTGTTTAATGAGGCACTGCACAGAGACCTAGCAGACTTCCGGATCCAGCACCCAGACTTGATCCTG
CTACAGTACGTGGATGACTTACTGCTGGCCGCCACTTCTGAGCTAGACTGCCAACAAGGTACTCGG
GCCCTGTTACAAACCCTAGGGAACCTCGGGTATCGGGCCTCGGCCAAGAAAGCCCAAATTTGCCA
GAAACAGGTCAAGTATCTGGGGTATCTTCTAAAAGAGGGTCAGAGATGGCTGACTGAGGCCAGAA
AAGAGACTGTGATGGGGCAGCCTACTCCGAAGACCCCTCGACAACTAAGGGAGTTCCTAGGGAAG
GCAGGCTTCTGTCGCCTCTTCATCCCTGGGTTTGCAGAAATGGCAGCCCCCCTGTACCCTCTCACCA
AACCGGGGACTCTGTTTAATTGGGGCCCAGACCAACAAAAGGCCTATCAAGAAATCAAGCAAGCT
CTTCTAACTGCCCCAGCCCTGGGGTTGCCAGATTTGACTAAGCCCTTTGAACTCTTTGTCGACGAG
AAGCAGGGCTACGCCAAAGGTGTCCTAACGCAAAAACTGGGACCTTGGCGTCGGCCGGTGGCCTA
CCTGTCCAAAAAGCTAGACCCAGTAGCAGCTGGGTGGCCCCCTTGCCTACGGATGGTAGCAGCCA
TTGCCGTACTGACAAAGGATGCAGGCAAGCTAACCATGGGACAGCCACTAGTCATTCTGGCCCCC
CATGCAGTAGAGGCACTAGTCAAACAACCCCCCGACCGCTGGCTTTCCAACGCCCGGATGACTCA
CTATCAGGCCTTGCTTTTGGACACGGACCGGGTCCAGTTCGGACCGGTGGTAGCCCTGAACCCGGC
TACGCTGCTCCCACTGCCTGAGGAAGGGCTGCAACACAACTGCCTTGATATCCTGGCCGAAGCCCA
CGGAACCCGACCCGACCTAACGGACCAGCCGCTCCCAGACGCCGACCACACCTGGTACACGGATG
GAAGCAGTCTCTTACAAGAGGGACAGCGTAAGGCGGGAGCTGCGGTGACCACCGAGACCGAGGT
AATCTGGGCTAAAGCCCTGCCAGCCGGGACATCCGCTCAGCGGGCTGAACTGATAGCACTCACCC
AGGCCCTAAAGATGGCAGAAGGTAAGAAGCTAAATGTTTATACTGATAGCCGTTATGCTTTTGCTA
CTGCCCATATCCATGGAGAAATATACAGAAGGCGTGGGTGGCTCACATCAGAAGGCAAAGAGATC
AAAAATAAAGACGAGATCTTGGCCCTACTAAAAGCCCTCTTTCTGCCCAAAAGACTTAGCATAATC
CATTGTCCAGGACATCAAAAGGGACACAGCGCCGAGGCTAGAGGCAACCGGATGGCTGACCAAG
CGGCCCGAAAGGCAGCCATCACAGAGACTCCAGACACCTCTACCCTCCTCATAGAAAATTCATCA
CCC
SEQ ID NO: 48, human codon optimized MMLV-RT, DNA
ACCCTGAACATTGAAGACGAGTATAGACTGCATGAAACAAGCAAGGAACCCGACGTGTCCCTGGG
CTCCACCTGGCTGTCCGACTTTCCCCAGGCCTGGGCCGAGACAGGAGGAATGGGCCTGGCCGTGC
GGCAGGCACCCCTGATCATCCCTCTGAAGGCCACCTCTACACCCGTGAGCATCAAGCAGTACCCTA
TGTCTCAGGAGGCCAGACTGGGCATCAAGCCTCACATCCAGAGGCTGCTGGACCAGGGCATCCTG
GTGCCATGCCAGAGCCCCTGGAACACACCACTGCTGCCCGTGAAGAAGCCAGGCACCAATGACTA
TAGACCCGTGCAGGATCTGAGAGAGGTGAACAAGAGGGTGGAGGATATCCACCCCACCGTGCCCA
ACCCTTACAATCTGCTGTCCGGCCTGCCCCCTTCTCACCAGTGGTATACAGTGCTGGACCTGAAGG
ATGCCTTCTTTTGTCTGAGACTGCACCCTACCAGCCAGCCACTGTTCGCCTTTGAGTGGAGGGACC
CTGAGATGGGCATCTCTGGCCAGCTGACCTGGACACGCCTGCCTCAGGGCTTCAAGAATAGCCCA
ACACTGTTTAACGAGGCCCTGCACCGCGACCTGGCAGATTTCCGGATCCAGCACCCAGATCTGATC
CTGCTGCAGTACGTGGACGATCTGCTGCTGGCCGCCACCAGCGAGCTGGATTGCCAGCAGGGAAC
ACGCGCCCTGCTGCAGACCCTGGGAAACCTGGGATATAGGGCATCCGCCAAGAAGGCCCAGATCT
GTCAGAAGCAGGTGAAGTACCTGGGCTATCTGCTGAAGGAGGGCCAGAGATGGCTGACAGAGGCC
AGGAAGGAGACAGTGATGGGCCAGCCAACACCCAAGACCCCAAGACAGCTGAGGGAGTTCCTGG
GCAAAGCAGGATTTTGCAGGCTGTTCATCCCAGGATTCGCAGAGATGGCAGCACCTCTGTACCCAC
TGACCAAGCCGGGCACCCTGTTTAATTGGGGCCCTGACCAGCAGAAGGCCTATCAGGAGATCAAG
CAGGCCCTGCTGACAGCACCAGCCCTGGGCCTGCCAGACCTGACCAAGCCTTTCGAGCTGTTTGTG
GATGAGAAGCAGGGCTACGCCAAGGGCGTGCTGACCCAGAAGCTGGGACCATGGAGACGGCCCG
TGGCCTATCTGTCCAAGAAGCTGGACCCAGTGGCAGCAGGATGGCCACCATGCCTGAGGATGGTG
GCAGCAATCGCCGTGCTGACAAAGGATGCCGGCAAGCTGACCATGGGACAGCCACTGGTCATCCT
GGCACCACACGCAGTGGAGGCCCTGGTGAAGCAGCCTCCAGATCGCTGGCTGTCTAACGCCCGGA
TGACACACTACCAGGCCCTGCTGCTGGACACCGATCGCGTGCAGTTTGGCCCTGTGGTGGCCCTGA
ATCCAGCCACCCTGCTGCCTCTGCCAGAGGAGGGCCTGCAGCACAACTGTCTGGACATCCTGGCAG
AGGCACACGGAACAAGGCCAGACCTGACCGATCAGCCCCTGCCTGACGCCGATCACACATGGTAT
ACCGATGGAAGCTCCCTGCTGCAGGAGGGCCAGAGGAAGGCAGGAGCAGCAGTGACCACAGAGA
CAGAAGTGATCTGGGCCAAGGCCCTGCCAGCAGGCACATCCGCCCAGCGGGCCGAGCTGATCGCC
CTGACCCAGGCCCTGAAGATGGCCGAGGGCAAGAAGCTGAACGTGTACACAGACTCCAGATATGC
CTTCGCCACCGCACACATCCACGGAGAGATCTACAGGCGCCGGGGCTGGCTGACCTCTGAGGGCA
AGGAGATCAAGAACAAGGATGAGATCCTGGCCCTGCTGAAGGCCCTGTTTCTGCCCAAGCGGCTG
AGCATCATCCACTGTCCTGGACACCAGAAGGGACACTCCGCCGAGGCAAGGGGCAATCGGATGGC CGACCAGGCCGCCAGAAAGGCTGCTATTACTGAAACTCCCGACACTTCCACTCTGCTGATTGAAAA
CTCCTCCCCT
SEQ ID NO: 49 (SSB1494), DNA
ATGGCTGAAAATGGTGATAATGAAAAGATGGCTGCCCTGGAGGCCAAAATCTGTCATCAAATTGA
GTATTATTTTGGCGACTTCAATTTGCCACGGGACAAGTTTCTAAAGGAACAGATAAAACTGGATGA
AGGCTGGGTACCTTTGGAGATAATGATAAAATTCAACAGGTTGAACCGTCTAACAACAGACTTTA
ATGTAATTGTGGAAGCATTGAGCAAATCCAAGGCAGAACTCATGGAAATCAGTGAAGATAAAACT
AAAATCAGAAGGTCTCCAAGCAAACCCCTACCTGAAGTGACTGATGAGTATAAAAATGATGTAAA
AAACAGATCTGTTTATATTAAAGGCTTCCCAACTGATGCAACTCTTGATGACATAAAAGAATGGTT
AGAAGATAAAGGTCAAGTACTAAATATTCAGATGAGAAGAACATTGCATAAAGCATTTAAGGGAT CAATTTTTGTTGTGTTTGATAGCATTGAATCTGCTAAGAAATTTGTAGAGACCCCTGGCCAGAAGT
ACAAAGAAACAGACCTGCTAATACTTTTCAAGGACGATTACTTTGCCAAAAAAAATGAA
SEQ ID NO: 50 (SSB), DNA
ATGGCTGAAAATGGTGATAATGAAAAGATGGCTGCCCTGGAGGCCAAAATCTGTCATCAAATTGA
GTATTATTTTGGCGACTTCAATTTGCCACGGGACAAGTTTCTAAAGGAACAGATAAAACTGGATGA
AGGCTGGGTACCTTTGGAGATAATGATAAAATTCAACAGGTTGAACCGTCTAACAACAGACTTTA
ATGTAATTGTGGAAGCATTGAGCAAATCCAAGGCAGAACTCATGGAAATCAGTGAAGATAAAACT
AAAATCAGAAGGTCTCCAAGCAAACCCCTACCTGAAGTGACTGATGAGTATAAAAATGATGTAAA
AAACAGATCTGTTTATATTAAAGGCTTCCCAACTGATGCAACTCTTGATGACATAAAAGAATGGTT
AGAAGATAAAGGTCAAGTACTAAATATTCAGATGAGAAGAACATTGCATAAAGCATTTAAGGGAT
CAATTTTTGTTGTGTTTGATAGCATTGAATCTGCTAAGAAATTTGTAGAGACCCCTGGCCAGAAGT
ACAAAGAAACAGACCTGCTAATACTTTTCAAGGACGATTACTTTGCCAAAAAAAATGAAGAAAGA
AAACAAAATAAAGTGGAAGCTAAATTAAGAGCTAAACAGGAGCAAGAAGCAAAACAAAAGTTAG
AAGAAGATGCTGAAATGAAATCTCTAGAAGAAAAGATTGGATGCTTGCTGAAATTTTCGGGTGAT
TTAGATGATCAGACCTGTAGAGAAGATTTACACATACTTTTCTCAAATCATGGTGAAATAAAATGG
ATAGACTTCGTCAGAGGAGCAAAAGAGGGGATAATTCTATTTAAAGAAAAAGCCAAGGAAGCATT
GGGTAAAGCCAAAGATGCAAATAATGGTAACCTACAATTAAGGAACAAAGAAGTGACTTGGGAA
GTACTAGAAGGAGAGGTGGAAAAAGAAGCACTGAAGAAAATAATAGAAGACCAACAAGAATCCC
TAAACAAATGGAAGTCAAAAGGTCGTAGATTTAAAGGAAAAGGAAAGGGTAATAAAGCTGCCCA
GCCTGGGTCTGGTAAAGGAAAAGTACAGTTTCAGGGCAAGAAAACGAAATTTGCTAGTGATGATG
AACATGATGAACATGATGAAAATGGTGCAACTGGACCTGTGAAAAGAGCAAGAGAAGAAACAGA CAAAGAAGAACCTGCATCCAAACAACAGAAAACAGAAAATGGTGCTGGAGACCAG
SEQ ID NO: 51 (SSB2 194), DNA
GCTGAAAATGGTGATAATGAAAAGATGGCTGCCCTGGAGGCCAAAATCTGTCATCAAATTGAGTA
TTATTTTGGCGACTTCAATTTGCCACGGGACAAGTTTCTAAAGGAACAGATAAAACTGGATGAAGG
CTGGGTACCTTTGGAGATAATGATAAAATTCAACAGGTTGAACCGTCTAACAACAGACTTTAATGT
AATTGTGGAAGCATTGAGCAAATCCAAGGCAGAACTCATGGAAATCAGTGAAGATAAAACTAAAA
TCAGAAGGTCTCCAAGCAAACCCCTACCTGAAGTGACTGATGAGTATAAAAATGATGTAAAAAAC
AGATCTGTTTATATTAAAGGCTTCCCAACTGATGCAACTCTTGATGACATAAAAGAATGGTTAGAA
GATAAAGGTCAAGTACTAAATATTCAGATGAGAAGAACATTGCATAAAGCATTTAAGGGATCAAT
TTTTGTTGTGTTTGATAGCATTGAATCTGCTAAGAAATTTGTAGAGACCCCTGGCCAGAAGTACAA
AGAAACAGACCTGCTAATACTTTTCAAGGACGATTACTTTGCCAAAAAAAATGAA
SEQ ID NO: 52 (SSB2408), DNA
GCTGAAAATGGTGATAATGAAAAGATGGCTGCCCTGGAGGCCAAAATCTGTCATCAAATTGAGTA
TTATTTTGGCGACTTCAATTTGCCACGGGACAAGTTTCTAAAGGAACAGATAAAACTGGATGAAGG
CTGGGTACCTTTGGAGATAATGATAAAATTCAACAGGTTGAACCGTCTAACAACAGACTTTAATGT
AATTGTGGAAGCATTGAGCAAATCCAAGGCAGAACTCATGGAAATCAGTGAAGATAAAACTAAAA
TCAGAAGGTCTCCAAGCAAACCCCTACCTGAAGTGACTGATGAGTATAAAAATGATGTAAAAAAC
AGATCTGTTTATATTAAAGGCTTCCCAACTGATGCAACTCTTGATGACATAAAAGAATGGTTAGAA
GATAAAGGTCAAGTACTAAATATTCAGATGAGAAGAACATTGCATAAAGCATTTAAGGGATCAAT
TTTTGTTGTGTTTGATAGCATTGAATCTGCTAAGAAATTTGTAGAGACCCCTGGCCAGAAGTACAA
AGAAACAGACCTGCTAATACTTTTCAAGGACGATTACTTTGCCAAAAAAAATGAAGAAAGAAAAC
AAAATAAAGTGGAAGCTAAATTAAGAGCTAAACAGGAGCAAGAAGCAAAACAAAAGTTAGAAGA
AGATGCTGAAATGAAATCTCTAGAAGAAAAGATTGGATGCTTGCTGAAATTTTCGGGTGATTTAGA
TGATCAGACCTGTAGAGAAGATTTACACATACTTTTCTCAAATCATGGTGAAATAAAATGGATAGA
CTTCGTCAGAGGAGCAAAAGAGGGGATAATTCTATTTAAAGAAAAAGCCAAGGAAGCATTGGGTA
AAGCCAAAGATGCAAATAATGGTAACCTACAATTAAGGAACAAAGAAGTGACTTGGGAAGTACTA
GAAGGAGAGGTGGAAAAAGAAGCACTGAAGAAAATAATAGAAGACCAACAAGAATCCCTAAACA
AATGGAAGTCAAAAGGTCGTAGATTTAAAGGAAAAGGAAAGGGTAATAAAGCTGCCCAGCCTGG
GTCTGGTAAAGGAAAAGTACAGTTTCAGGGCAAGAAAACGAAATTTGCTAGTGATGATGAACATG
ATGAACATGATGAAAATGGTGCAACTGGACCTGTGAAAAGAGCAAGAGAAGAAACAGACAAAGA AGAACCTGCATCCAAACAACAGAAAACAGAAAATGGTGCTGGAGACCAG
SEQ ID NO: 53 (bpNLSSV40), DNA
AAAAGAACCGCCGACGGCAGCGAATTCGAGCCCAAGAAGAAGAGGAAAGTC
SEQ ID NO: 54 (bpNLSSV40), DNA
AAAAGAACCGCCGACGGCAGCGAATTCGAGTCTCCCAAGAAGAAGAGGAAAGTC
SEQ ID NO: 55 (NLSc K'c). DNA
GGCTCTGGCCCTGCCGCTAAGAGAGTGAAGCTGGAC
SEQ ID NO: 56 linker B (SGGSx2-NLSSV40-SGGSx2), DNA
TCCGGCGGAAGCTCTGGTGGCAGCAAGCGGACCGCCGACGGCTCTGAATTCGAGAGCCCTAAGAA GAAAAGAAAGGTGAGCGGAGGCTCTAGCGGCGGAAGC
SEQ ID NO: 57 PEmax-La1 194(Q20A_Y23A_Y24F_F35A)(PE7 mutant)
MKRTADGSEFESPKKKRKVDKKYSIGLDIGTNSVGWAVITDEYKVPSKKFKVLGNTDRHSIKKNLIGA
LLFDSGETAEATRLKRTARRRYTRRKNRICYLQEIFSNEMAKVDDSFFHRLEESFLVEEDKKHERHPIFG
NIVDEVAYHEKYPTIYHLRKKLVDSTDKADLRLIYLALAHMIKFRGHFLIEGDLNPDNSDVDKLFIQLV
QTYNQLFEENPINASGVDAKAILSARLSKSRKLENLIAQLPGEKKNGLFGNLIALSLGLTPNFKSNFDLA
EDAKLQLSKDTYDDDLDNLLAQIGDQYADLFLAAKNLSDAILLSDILRVNTEITKAPLSASMIKRYDEH
HQDLTLLKALVRQQLPEKYKEIFFDQSKNGYAGYIDGGASQEEFYKFIKPILEKMDGTEELLVKLKRED
LLRKQRTFDNGSIPHQIHLGELHAILRRQEDFYPFLKDNREKIEKILTFRIPYYVGPLARGNSRFAWMTR
KSEETITPWNFEEWDKGASAQSFIERMTNFDKNLPNEKVLPKHSLLYEYFTVYNELTKVKYVTEGMR
KPAFLSGEQKKAIVDLLFKTNRKVTVKQLKEDYFKKIECFDSVEISGVEDRFNASLGTYHDLLKIIKDKD
FLDNEENEDILEDIVLTLTLFEDREMIEERLKTYAHLFDDKVMKQLKRRRYTGWGRLSRKLINGIRDKQ
SGKTILDFLKSDGFANRNFMQLIHDDSLTFKEDIQKAQVSGQGDSLHEHIANLAGSPAIKKGILQTVKV
VDELVKVMGRHKPENIVIEMARENQTTQKGQKNSRERMKRIEEGIKELGSQILKEHPVENTQLQNEKL
YLYYLQNGRDMYVDQELDINRLSDYDVDAIVPQSFLKDDSIDNKVLTRSDKNRGKSDNVPSEEWKK
MKNYWRQLLNAKLITQRKFDNLTKAERGGLSELDKAGFIKRQLVETRQITKHVAQILDSRMNTKYDE
NDKLIREVKVITLKSKLVSDFRKDFQFYKVREINNYHHAHDAYLNAWGTALIKKYPKLESEFVYGDY
KVYDVRKMIAKSEQEIGKATAKYFFYSNIMNFFKTEITLANGEIRKRPLIETNGETGEIVWDKGRDFAT
VRKVLSMPQVNIVKKTEVQTGGFSKESILPKRNSDKLIARKKDWDPKKYGGFDSPTVAYSVLWAKVE
KGKSKKLKSVKELLGITIMERSSFEKNPIDFLEAKGYKEVKKDLIIKLPKYSLFELENGRKRMLASAGEL
QKGNELALPSKYVNFLYLASHYEKLKGSPEDNEQKQLFVEQHKHYLDEIIEQISEFSKRVILADANLDK
VLSAYNKHRDKPIREQAENIIHLFTLTNLGAPAAFKYFDTTIDRKRYTSTKEVLDATLIHQSITGLYETRI
DLSQLGGDSGGSSGGSKRTADGSEFESPKKKRKVSGGSSGGSTLNIEDEYRLHETSKEPDVSLGSTWLS
DFPQAWAETGGMGLAVRQAPLIIPLKATSTPVSIKQYPMSQEARLGIKPHIQRLLDQGILVPCQSPWNTP
LLPVKKPGTNDYRPVQDLREVNKRVEDIHPTVPNPYNLLSGLPPSHQWYTVLDLKDAFFCLRLHPTSQP
LFAFEWRDPEMGISGQLTWTRLPQGFKNSPTLFNEALHRDLADFRIQHPDLILLQYVDDLLLAATSELD
CQQGTRALLQTLGNLGYRASAKKAQICQKQVKYLGYLLKEGQRWLTEARKETVMGQPTPKTPRQLR
EFLGKAGFCRLFIPGFAEMAAPLYPLTKPGTLFNWGPDQQKAYQEIKQALLTAPALGLPDLTKPFELFV
DEKQGYAKGVLTQKLGPWRRPVAYLSKKLDPVAAGWPPCLRMVAAIAVLTKDAGKLTMGQPLVILA
PHAVEALVKQPPDRWLSNARMTHYQALLLDTDRVQFGPWALNPATLLPLPEEGLQHNCLDILAEAH
GTRPDLTDQPLPDADHTWYTDGSSLLQEGQRKAGAAVTTETEVIWAKALPAGTSAQRAELIALTQALK
MAEGKKLNVYTDSRYAFATAHIHGEIYRRRGWLTSEGKEIKNKDEILALLKALFLPKRLSIIHCPGHQK
GHSAEARGNRMADQAARKAAITETPDTSTLLIENSSPSGGSSGGSSGSETPGTSESATPESSGGSSGGSM
AENGDNEKMAALEAKICHAIEAFFGDFNLPRDKALKEQIKLDEGWVPLEIMIKFNRLNRLTTDFNVIVE
ALSKSKAELMEISEDKTKIRRSPSKPLPEVTDEYKNDVKNRSVYIKGFPTDATLDDIKEWLEDKGQVLNI
QMRRTLHKAFKGSIFWFDSIESAKKFVETPGQKYKETDLLILFKDDYFAKKNESGGSKRTADGSEFES PKKKRKVGSGPAAKRVKLD*
SEQ ID NO: 58 PEmax-La1 194(Q20A_Y23A_Y24F_F35A)(PE7 mutant), DNA
ATGAAACGGACAGCCGACGGAAGCGAGTTCGAGTCACCAAAGAAGAAGCGGAAAGTCGACAAGA
AGTACAGCATCGGCCTGGACATCGGCACCAACTCTGTGGGCTGGGCCGTGATCACCGACGAGTAC
AAGGTGCCCAGCAAGAAATTCAAGGTGCTGGGCAACACCGACCGGCACAGCATCAAGAAGAACC
TGATCGGAGCCCTGCTGTTCGACAGCGGCGAAACAGCCGAGGCCACCCGGCTGAAGAGAACCGCC
AGAAGAAGATACACCAGACGGAAGAACCGGATCTGCTATCTGCAAGAGATCTTCAGCAACGAGAT
GGCCAAGGTGGACGACAGCTTCTTCCACAGACTGGAAGAGTCCTTCCTGGTGGAAGAGGATAAGA
AGCACGAGCGGCACCCCATCTTCGGCAACATCGTGGACGAGGTGGCCTACCACGAGAAGTACCCC
ACCATCTACCACCTGAGAAAGAAACTGGTGGACAGCACCGACAAGGCCGACCTGCGGCTGATCTA
TCTGGCCCTGGCCCACATGATCAAGTTCCGGGGCCACTTCCTGATCGAGGGCGACCTGAACCCCGA
CAACAGCGACGTGGACAAGCTGTTCATCCAGCTGGTGCAGACCTACAACCAGCTGTTCGAGGAAA
ACCCCATCAACGCCAGCGGCGTGGACGCCAAGGCCATCCTGTCTGCCAGACTGAGCAAGAGCAGA
AAGCTGGAAAATCTGATCGCCCAGCTGCCCGGCGAGAAGAAGAATGGCCTGTTCGGAAACCTGAT
TGCCCTGAGCCTGGGCCTGACCCCCAACTTCAAGAGCAACTTCGACCTGGCCGAGGATGCCAAACT
GCAGCTGAGCAAGGACACCTACGACGACGACCTGGACAACCTGCTGGCCCAGATCGGCGACCAGT
ACGCCGACCTGTTTCTGGCCGCCAAGAACCTGTCCGACGCCATCCTGCTGAGCGACATCCTGAGAG
TGAACACCGAGATCACCAAGGCCCCCCTGAGCGCCTCTATGATCAAGAGATACGACGAGCACCAC
CAGGACCTGACCCTGCTGAAAGCTCTCGTGCGGCAGCAGCTGCCTGAGAAGTACAAAGAGATTTT
CTTCGACCAGAGCAAGAACGGCTACGCCGGCTACATTGACGGCGGAGCCAGCCAGGAAGAGTTCT
ACAAGTTCATCAAGCCCATCCTGGAAAAGATGGACGGCACCGAGGAACTGCTCGTGAAGCTGAAG
AGAGAGGACCTGCTGCGGAAGCAGCGGACCTTCGACAACGGCAGCATCCCCCACCAGATCCACCT
GGGAGAGCTGCACGCCATTCTGCGGCGGCAGGAAGATTTTTACCCATTCCTGAAGGACAACCGGG
AAAAGATCGAGAAGATCCTGACCTTCCGCATCCCCTACTACGTGGGCCCTCTGGCCAGGGGAAAC
AGCAGATTCGCCTGGATGACCAGAAAGAGCGAGGAAACCATCACCCCCTGGAACTTCGAGGAAGT
GGTGGACAAGGGCGCTTCCGCCCAGAGCTTCATCGAGCGGATGACCAACTTCGATAAGAACCTGC
CCAACGAGAAGGTGCTGCCCAAGCACAGCCTGCTGTACGAGTACTTCACCGTGTATAACGAGCTG
ACCAAAGTGAAATACGTGACCGAGGGAATGAGAAAGCCCGCCTTCCTGAGCGGCGAGCAGAAAA
AGGCCATCGTGGACCTGCTGTTCAAGACCAACCGGAAAGTGACCGTGAAGCAGCTGAAAGAGGAC
TACTTCAAGAAAATCGAGTGCTTCGACTCCGTGGAAATCTCCGGCGTGGAAGATCGGTTCAACGCC
TCCCTGGGCACATACCACGATCTGCTGAAAATTATCAAGGACAAGGACTTCCTGGACAATGAGGA
AAACGAGGACATTCTGGAAGATATCGTGCTGACCCTGACACTGTTTGAGGACAGAGAGATGATCG
AGGAACGGCTGAAAACCTATGCCCACCTGTTCGACGACAAAGTGATGAAGCAGCTGAAGCGGCGG
AGATACACCGGCTGGGGCAGGCTGAGCCGGAAGCTGATCAACGGCATCCGGGACAAGCAGTCCG
GCAAGACAATCCTGGATTTCCTGAAGTCCGACGGCTTCGCCAACAGAAACTTCATGCAGCTGATCC
ACGACGACAGCCTGACCTTTAAAGAGGACATCCAGAAAGCCCAGGTGTCCGGCCAGGGCGATAGC
CTGCACGAGCACATTGCCAATCTGGCCGGCAGCCCCGCCATTAAGAAGGGCATCCTGCAGACAGT
GAAGGTGGTGGACGAGCTCGTGAAAGTGATGGGCCGGCACAAGCCCGAGAACATCGTGATCGAA
ATGGCCAGAGAGAACCAGACCACCCAGAAGGGACAGAAGAACAGCCGCGAGAGAATGAAGCGG
ATCGAAGAGGGCATCAAAGAGCTGGGCAGCCAGATCCTGAAAGAACACCCCGTGGAAAACACCC
AGCTGCAGAACGAGAAGCTGTACCTGTACTACCTGCAGAATGGGCGGGATATGTACGTGGACCAG
GAACTGGACATCAACCGGCTGTCCGACTACGATGTGGACGCTATCGTGCCTCAGAGCTTTCTGAAG
GACGACTCCATCGACAACAAGGTGCTGACCAGAAGCGACAAGAACCGGGGCAAGAGCGACAACG
TGCCCTCCGAAGAGGTCGTGAAGAAGATGAAGAACTACTGGCGGCAGCTGCTGAACGCCAAGCTG
ATTACCCAGAGAAAGTTCGACAATCTGACCAAGGCCGAGAGAGGCGGCCTGAGCGAACTGGATAA
GGCCGGCTTCATCAAGAGACAGCTGGTGGAAACCCGGCAGATCACAAAGCACGTGGCACAGATCC
TGGACTCCCGGATGAACACTAAGTACGACGAGAATGACAAGCTGATCCGGGAAGTGAAAGTGATC
ACCCTGAAGTCCAAGCTGGTGTCCGATTTCCGGAAGGATTTCCAGTTTTACAAAGTGCGCGAGATC
AACAACTACCACCACGCCCACGACGCCTACCTGAACGCCGTCGTGGGAACCGCCCTGATCAAAAA
GTACCCTAAGCTGGAAAGCGAGTTCGTGTACGGCGACTACAAGGTGTACGACGTGCGGAAGATGA
TCGCCAAGAGCGAGCAGGAAATCGGCAAGGCTACCGCCAAGTACTTCTTCTACAGCAACATCATG
AACTTTTTCAAGACCGAGATTACCCTGGCCAACGGCGAGATCCGGAAGCGGCCTCTGATCGAGAC
AAACGGCGAAACCGGGGAGATCGTGTGGGATAAGGGCCGGGATTTTGCCACCGTGCGGAAAGTGC
TGAGCATGCCCCAAGTGAATATCGTGAAAAAGACCGAGGTGCAGACAGGCGGCTTCAGCAAAGA
GTCTATCCTGCCCAAGAGGAACAGCGATAAGCTGATCGCCAGAAAGAAGGACTGGGACCCTAAGA
AGTACGGCGGCTTCGACAGCCCCACCGTGGCCTATTCTGTGCTGGTGGTGGCCAAAGTGGAAAAG
GGCAAGTCCAAGAAACTGAAGAGTGTGAAAGAGCTGCTGGGGATCACCATCATGGAAAGAAGCA
GCTTCGAGAAGAATCCCATCGACTTTCTGGAAGCCAAGGGCTACAAAGAAGTGAAAAAGGACCTG
ATCATCAAGCTGCCTAAGTACTCCCTGTTCGAGCTGGAAAACGGCCGGAAGAGAATGCTGGCCTCT
GCCGGCGAACTGCAGAAGGGAAACGAACTGGCCCTGCCCTCCAAATATGTGAACTTCCTGTACCT
GGCCAGCCACTATGAGAAGCTGAAGGGCTCCCCCGAGGATAATGAGCAGAAACAGCTGTTTGTGG
AACAGCACAAGCACTACCTGGACGAGATCATCGAGCAGATCAGCGAGTTCTCCAAGAGAGTGATC
CTGGCCGACGCTAATCTGGACAAAGTGCTGTCCGCCTACAACAAGCACCGGGATAAGCCCATCAG
AGAGCAGGCCGAGAATATCATCCACCTGTTTACCCTGACCAATCTGGGAGCCCCTGCCGCCTTCAA
GTACTTTGACACCACCATCGACCGGAAGAGGTACACCAGCACCAAAGAGGTGCTGGACGCCACCC
TGATCCACCAGAGCATCACCGGCCTGTACGAGACACGGATCGACCTGTCTCAGCTGGGAGGTGAC
TCCGGCGGAAGCTCTGGTGGCAGCAAGCGGACCGCCGACGGCTCTGAATTCGAGAGCCCTAAGAA
GAAAAGAAAGGTGAGCGGAGGCTCTAGCGGCGGAAGCACCCTGAACATTGAAGACGAGTATAGA
CTGCATGAAACAAGCAAGGAACCCGACGTGTCCCTGGGCTCCACCTGGCTGTCCGACTTTCCCCAG
GCCTGGGCCGAGACAGGAGGAATGGGCCTGGCCGTGCGGCAGGCACCCCTGATCATCCCTCTGAA
GGCCACCTCTACACCCGTGAGCATCAAGCAGTACCCTATGTCTCAGGAGGCCAGACTGGGCATCA
AGCCTCACATCCAGAGGCTGCTGGACCAGGGCATCCTGGTGCCATGCCAGAGCCCCTGGAACACA
CCACTGCTGCCCGTGAAGAAGCCAGGCACCAATGACTATAGACCCGTGCAGGATCTGAGAGAGGT
GAACAAGAGGGTGGAGGATATCCACCCCACCGTGCCCAACCCTTACAATCTGCTGTCCGGCCTGCC
CCCTTCTCACCAGTGGTATACAGTGCTGGACCTGAAGGATGCCTTCTTTTGTCTGAGACTGCACCCT
ACCAGCCAGCCACTGTTCGCCTTTGAGTGGAGGGACCCTGAGATGGGCATCTCTGGCCAGCTGACC
TGGACACGCCTGCCTCAGGGCTTCAAGAATAGCCCAACACTGTTTAACGAGGCCCTGCACCGCGA
CCTGGCAGATTTCCGGATCCAGCACCCAGATCTGATCCTGCTGCAGTACGTGGACGATCTGCTGCT
GGCCGCCACCAGCGAGCTGGATTGCCAGCAGGGAACACGCGCCCTGCTGCAGACCCTGGGAAACC
TGGGATATAGGGCATCCGCCAAGAAGGCCCAGATCTGTCAGAAGCAGGTGAAGTACCTGGGCTAT
CTGCTGAAGGAGGGCCAGAGATGGCTGACAGAGGCCAGGAAGGAGACAGTGATGGGCCAGCCAA
CACCCAAGACCCCAAGACAGCTGAGGGAGTTCCTGGGCAAAGCAGGATTTTGCAGGCTGTTCATC
CCAGGATTCGCAGAGATGGCAGCACCTCTGTACCCACTGACCAAGCCGGGCACCCTGTTTAATTGG
GGCCCTGACCAGCAGAAGGCCTATCAGGAGATCAAGCAGGCCCTGCTGACAGCACCAGCCCTGGG
CCTGCCAGACCTGACCAAGCCTTTCGAGCTGTTTGTGGATGAGAAGCAGGGCTACGCCAAGGGCG
TGCTGACCCAGAAGCTGGGACCATGGAGACGGCCCGTGGCCTATCTGTCCAAGAAGCTGGACCCA
GTGGCAGCAGGATGGCCACCATGCCTGAGGATGGTGGCAGCAATCGCCGTGCTGACAAAGGATGC
CGGCAAGCTGACCATGGGACAGCCACTGGTCATCCTGGCACCACACGCAGTGGAGGCCCTGGTGA
AGCAGCCTCCAGATCGCTGGCTGTCTAACGCCCGGATGACACACTACCAGGCCCTGCTGCTGGACA
CCGATCGCGTGCAGTTTGGCCCTGTGGTGGCCCTGAATCCAGCCACCCTGCTGCCTCTGCCAGAGG
AGGGCCTGCAGCACAACTGTCTGGACATCCTGGCAGAGGCACACGGAACAAGGCCAGACCTGACC
GATCAGCCCCTGCCTGACGCCGATCACACATGGTATACCGATGGAAGCTCCCTGCTGCAGGAGGG
CCAGAGGAAGGCAGGAGCAGCAGTGACCACAGAGACAGAAGTGATCTGGGCCAAGGCCCTGCCA
GCAGGCACATCCGCCCAGCGGGCCGAGCTGATCGCCCTGACCCAGGCCCTGAAGATGGCCGAGGG
CAAGAAGCTGAACGTGTACACAGACTCCAGATATGCCTTCGCCACCGCACACATCCACGGAGAGA
TCTACAGGCGCCGGGGCTGGCTGACCTCTGAGGGCAAGGAGATCAAGAACAAGGATGAGATCCTG
GCCCTGCTGAAGGCCCTGTTTCTGCCCAAGCGGCTGAGCATCATCCACTGTCCTGGACACCAGAAG
GGACACTCCGCCGAGGCAAGGGGCAATCGGATGGCCGACCAGGCCGCCAGAAAGGCTGCTATTAC
TGAAACTCCCGACACTTCCACTCTGCTGATTGAAAACTCCTCCCCTTCCGGAGGATCTAGCGGAGG
CTCCTCTGGCTCTGAGACACCTGGCACAAGCGAGAGCGCAACACCTGAAAGCAGCGGGGGCAGCA
GCGGGGGGTCAATGGCTGAAAATGGTGATAATGAAAAGATGGCTGCCCTGGAGGCCAAAATCTGT
CATGCCATTGAGGCCTTCTTTGGCGACTTCAATTTGCCACGGGACAAGGCCCTAAAGGAACAGATA
AAACTGGATGAAGGCTGGGTACCTTTGGAGATAATGATAAAATTCAACAGGTTGAACCGTCTAAC
AACAGACTTTAATGTAATTGTGGAAGCATTGAGCAAATCCAAGGCAGAACTCATGGAAATCAGTG
AAGATAAAACTAAAATCAGAAGGTCTCCAAGCAAACCCCTACCTGAAGTGACTGATGAGTATAAA
AATGATGTAAAAAACAGATCTGTTTATATTAAAGGCTTCCCAACTGATGCAACTCTTGATGACATA
AAAGAATGGTTAGAAGATAAAGGTCAAGTACTAAATATTCAGATGAGAAGAACATTGCATAAAGC
ATTTAAGGGATCAATTTTTGTTGTGTTTGATAGCATTGAATCTGCTAAGAAATTTGTAGAGACCCCT GGCCAGAAGTACAAAGAAACAGACCTGCTAATACTTTTCAAGGACGATTACTTTGCCAAAAAAAA
TGAATCTGGCGGCTCAAAAAGAACCGCCGACGGCAGCGAATTCGAGTCTCCCAAGAAGAAGAGG
AAAGTCGGCTCTGGCCCTGCCGCTAAGAGAGTGAAGCTGGACTAA
SEQ ID NO: 59 SSB1-194 (Q20A Y23A Y24F F35A)
MAENGDNEKMAALEAKICHAIEAFFGDFNLPRDKALKEQIKLDEGWVPLEIMIKFNRLNRLTTDFNVIV EALSKSKAELMEISEDKTKIRRSPSKPLPEVTDEYKNDVKNRSVYIKGFPTDATLDDIKEWLEDKGQVL
NIQMRRTLHKAFKGSIFWFDSIESAKKFVETPGQKYKETDLLILFKDDYFAKKNE
SEQ ID NO: 60 SSB1 194 (Q20A Y23A Y24F F35A), DNA
ATGGCTGAAAATGGTGATAATGAAAAGATGGCTGCCCTGGAGGCCAAAATCTGTCATgccATTGAG gccttcTTTGGCGACTTCAATTTGCCACGGGACAAGgccCTAAAGGAACAGATAAAACTGGATGAAGG
CTGGGTACCTTTGGAGATAATGATAAAATTCAACAGGTTGAACCGTCTAACAACAGACTTTAATGT
AATTGTGGAAGCATTGAGCAAATCCAAGGCAGAACTCATGGAAATCAGTGAAGATAAAACTAAAA
TCAGAAGGTCTCCAAGCAAACCCCTACCTGAAGTGACTGATGAGTATAAAAATGATGTAAAAAAC
AGATCTGTTTATATTAAAGGCTTCCCAACTGATGCAACTCTTGATGACATAAAAGAATGGTTAGAA GATAAAGGTCAAGTACTAAATATTCAGATGAGAAGAACATTGCATAAAGCATTTAAGGGATCAAT
TTTTGTTGTGTTTGATAGCATTGAATCTGCTAAGAAATTTGTAGAGACCCCTGGCCAGAAGTACAA
AGAAACAGACCTGCTAATACTTTTCAAGGACGATTACTTTGCCAAAAAAAATGAA
Table 3 Legend m = 2’-O-methylation (or 2’-O-methyl modification)
* = phosphorothioate bond r = RNA base (or ribonucleotide)
Claims
1. A system comprising: a Cas9 nickase; a reverse transcriptase; a small RNA binding exonuclease protection factor La (SSB) protein; and a prime editing guide RNA (pegRNA).
2. The system of claim 1, wherein the pegRNA comprises a 3 ’-polyuridine domain.
3. The system of claim 2, wherein the 3 ’-polyuridine domain comprises at least one chemical modification.
4. The system of claim 3, wherein the at least one chemical modification is 2’-O- methylation.
5. The system of claim 3 or 4, wherein the at least one chemical modification is replacement of a phosphodiester bond to a phosphorothioate bond.
6. The system of claim 2 or 3, wherein the 3 ’-polyuridine domain comprises at least one uridine with unmodified 2'-hydroxyl (OH) group.
7. The system of claim 6, wherein the at least one uridine with unmodified 2'- OH group locates at a 3’ end of the pegRNA.
8. The system of any one of claims 1-7, wherein the SSB protein comprises a La motif and/or an RNA recognition motif (RRM).
9. The system of any one of claims 1-7, wherein the SSB protein comprises a sequence at least 80% identical to SEQ ID NO: 34 or a fragment thereof.
10. The system of claim 9, wherein the SSB protein comprises amino acid residues 1-194 or 2-194 of SEQ ID NO: 34.
11. The system of claim 9, wherein the SSB protein comprises a sequence at least 80% identical to SEQ ID NO: 33 or SEQ ID NO: 35 or a fragment thereof.
12. The system of any one of claims 1-11, wherein the Cas9 nickase comprises a sequence at least 80% identical to SEQ ID NO: 26 or 27, or a fragment thereof.
13. The system of any one of claims 1-12, wherein the SSB protein is operatively linked to the Cas9 nickase and the reverse transcriptase.
14. The system of claim 13, comprising a recombinant polypeptide that comprises the SSB protein, Cas9 nickase, and the reverse transcriptase, wherein the recombinant polypeptide comprises a sequence at least 80% identical to any of SEQ ID NOs: 1-12 or a fragment thereof.
15. A system comprising: a first polynucleotide encoding a Cas9 nickase; a second polynucleotide encoding a reverse transcriptase; and a third polynucleotide encoding a small RNA binding exonuclease protection factor La (SSB) protein; and a prime editing guide RNA (pegRNA).
16. The system of claim 15, wherein the pegRNA comprises a 3 ’-polyuridine domain.
17. The system of claim 16, wherein the 3 ’-polyuridine domain comprises at least one chemical modification.
18. The system of claim 17, wherein the at least one chemical modification is 2’-O- methylation .
19. The system of claim 17 or 18, wherein the at least one chemical modification is replacement of a phosphodiester bond to a phosphorothioate bond.
20. The system of claim 16 or 17, wherein the 3 ’-polyuridine domain comprises at least one uridine with unmodified 2'-hydroxyl (OH) group.
21. The system of claim 20, wherein the at least one uridine with unmodified 2'-hydroxyl (OH) group locates at a 3’ end of the pegRNA.
22. The system of any one of claims 15-21, wherein the third polynucleotide encodes a La motif and/or an RNA recognition motif (RRM) of the SSB protein.
23. The system of any one of claims 15-22, wherein the third polynucleotide comprises a sequence at least 80% identical to SEQ ID NO: 49, 50, 51, or 52, or a fragment thereof.
24. The system of any one of claims 15-23, wherein the Cas9 nickase comprises a sequence at least 80% identical to SEQ ID NO: 42 or 43, or a fragment thereof.
25. The system of any one of claims 15-24, wherein the first, second, and third polynucleotides are operatively linked.
26. The system of any one of claims 15-25, comprising a sequence at least 80% identical to any of SEQ ID NOs: 13-24.
27. The system of any one of claims 15-26, wherein the first, second, and third polynucleotides are located on the same or different vectors.
28. A pharmaceutical composition comprising the system of any one of claims 1-27.
29. A method of treating a genetic disorder in a subject in need, comprising administering to the subject a therapeutically effective amount of the system of any one of claims 1- 27 or the pharmaceutical composition of claim 28.
30. A method for altering expression of a gene product in a cell, comprising introducing into the cell an effective amount of the system of any one of claims 1-27.
31. The method of claim 30, wherein the cell is a mammalian cell.
32. The method of claim 31, wherein the mammalian cell is in a human.
Applications Claiming Priority (4)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| US202363527444P | 2023-07-18 | 2023-07-18 | |
| US63/527,444 | 2023-07-18 | ||
| US202363611931P | 2023-12-19 | 2023-12-19 | |
| US63/611,931 | 2023-12-19 |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| WO2025019675A1 true WO2025019675A1 (en) | 2025-01-23 |
Family
ID=94282680
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/US2024/038551 Pending WO2025019675A1 (en) | 2023-07-18 | 2024-07-18 | Prime editing system and uses thereof |
Country Status (1)
| Country | Link |
|---|---|
| WO (1) | WO2025019675A1 (en) |
Citations (3)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO2023039586A1 (en) * | 2021-09-10 | 2023-03-16 | Agilent Technologies, Inc. | Guide rnas with chemical modification for prime editing |
| WO2023070110A2 (en) * | 2021-10-21 | 2023-04-27 | Prime Medicine, Inc. | Genome editing compositions and methods for treatment of retinitis pigmentosa |
| WO2023070062A2 (en) * | 2021-10-21 | 2023-04-27 | Prime Medicine, Inc. | Genome editing compositions and methods for treatment of usher syndrome type 3 |
-
2024
- 2024-07-18 WO PCT/US2024/038551 patent/WO2025019675A1/en active Pending
Patent Citations (3)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO2023039586A1 (en) * | 2021-09-10 | 2023-03-16 | Agilent Technologies, Inc. | Guide rnas with chemical modification for prime editing |
| WO2023070110A2 (en) * | 2021-10-21 | 2023-04-27 | Prime Medicine, Inc. | Genome editing compositions and methods for treatment of retinitis pigmentosa |
| WO2023070062A2 (en) * | 2021-10-21 | 2023-04-27 | Prime Medicine, Inc. | Genome editing compositions and methods for treatment of usher syndrome type 3 |
Non-Patent Citations (2)
| Title |
|---|
| ALFANO CATERINA, SANFELICE DOMENICO, BABON JEFF, KELLY GEOFF, JACKS AMANDA, CURRY STEPHEN, CONTE MARIA R: "Structural analysis of cooperative RNA binding by the La motif and central RRM domain of human La protein", NATURE STRUCTURAL & MOLECULAR BIOLOGY, NATURE PUBLISHING GROUP US, NEW YORK, vol. 11, no. 4, 1 April 2004 (2004-04-01), New York , pages 323 - 329, XP093268078, ISSN: 1545-9993, DOI: 10.1038/nsmb747 * |
| YAN ET AL.: "Improving prime editing with an endogenous small RNA-binding protein", NATURE, vol. 628, 3 April 2024 (2024-04-03), pages 639 - 647, XP038062744, DOI: 10.1038/s41586-024-07259-6 * |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| Chen et al. | Enhanced prime editing systems by manipulating cellular determinants of editing outcomes | |
| US20210310022A1 (en) | Massively parallel combinatorial genetics for crispr | |
| JP2024503437A (en) | Prime editing factor variants, constructs, and methods to improve prime editing efficiency and accuracy | |
| JP2023159185A (en) | Compositions and methods for treating hemoglobinopathies | |
| Voit et al. | Nuclease-mediated gene editing by homologous recombination of the human globin locus | |
| CA3026110A1 (en) | Novel crispr enzymes and systems | |
| EP3872177B1 (en) | Compositions and methods for enhancing homologous recombination | |
| WO2017070429A1 (en) | Methods involving editing polynucleotides that encode t cell receptor | |
| WO2016123071A1 (en) | Methods of identifying essential protein domains | |
| Zhao et al. | Ligation-assisted homologous recombination enables precise genome editing by deploying both MMEJ and HDR | |
| JP2024545144A (en) | OMNI-103 CRISPR nuclease-RNA complex | |
| WO2024003810A1 (en) | Guide rna with chemical modifications | |
| Iyer et al. | Efficient homology-directed repair with circular ssDNA donors | |
| EP4288088A2 (en) | Lymphocyte activation gene 3 (lag3) compositions and methods for immunotherapy | |
| WO2025019675A1 (en) | Prime editing system and uses thereof | |
| Cattle et al. | An enhanced Eco1 retron editor enables precision genome engineering in human cells without double-strand breaks | |
| Yan | Mapping the Cellular Determinants of Genome Editing | |
| Ponnienselvan | Addressing Bottlenecks of Prime Editing Through Improved pegRNA Designs and Rationally Engineered Prime Editor Variants | |
| WO2025076306A1 (en) | Prime editors having improved prime editing efficiency | |
| HK40080440A (en) | Compositions and methods for the treatment of hemoglobinopathies | |
| HK40058696A (en) | Compositions and methods for enhancing homologous recombination | |
| JP2026505389A (en) | Engineered OMNI-50 nuclease variants | |
| EP4392060A1 (en) | Programmed cell death protein 1 (pd1) compositions and methods for cell-based therapy | |
| HK40116323A (en) | Omni-103 crispr nuclease-rna complexes | |
| HK40111655A (en) | Novel omni 115, 124, 127, 144-149, 159, 218, 237, 248, 251-253 and 259 crispr nucleases |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| 121 | Ep: the epo has been informed by wipo that ep was designated in this application |
Ref document number: 24843960 Country of ref document: EP Kind code of ref document: A1 |