WO2025017151A1 - Stable poly(a)-encoding messenger rna templates - Google Patents
Stable poly(a)-encoding messenger rna templates Download PDFInfo
- Publication number
- WO2025017151A1 WO2025017151A1 PCT/EP2024/070452 EP2024070452W WO2025017151A1 WO 2025017151 A1 WO2025017151 A1 WO 2025017151A1 EP 2024070452 W EP2024070452 W EP 2024070452W WO 2025017151 A1 WO2025017151 A1 WO 2025017151A1
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- template
- poly
- tail
- mrna
- adenosines
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Pending
Links
Classifications
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/63—Introduction of foreign genetic material using vectors; Vectors; Use of hosts therefor; Regulation of expression
- C12N15/67—General methods for enhancing the expression
- C12N15/68—Stabilisation of the vector
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/63—Introduction of foreign genetic material using vectors; Vectors; Use of hosts therefor; Regulation of expression
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/63—Introduction of foreign genetic material using vectors; Vectors; Use of hosts therefor; Regulation of expression
- C12N15/67—General methods for enhancing the expression
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12P—FERMENTATION OR ENZYME-USING PROCESSES TO SYNTHESISE A DESIRED CHEMICAL COMPOUND OR COMPOSITION OR TO SEPARATE OPTICAL ISOMERS FROM A RACEMIC MIXTURE
- C12P19/00—Preparation of compounds containing saccharide radicals
- C12P19/26—Preparation of nitrogen-containing carbohydrates
- C12P19/28—N-glycosides
- C12P19/30—Nucleotides
- C12P19/34—Polynucleotides, e.g. nucleic acids, oligoribonucleotides
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N2830/00—Vector systems having a special element relevant for transcription
- C12N2830/50—Vector systems having a special element relevant for transcription regulating RNA stability, not being an intron, e.g. poly A signal
Definitions
- the present invention relates to templates comprising a nucleic acid sequence encoding a segmented poly(A) tail for the in vitro transcription (IVT) of messenger RNA (mRNA).
- the templates of the invention are stably maintained in bacterial cells during propagation and are non-inferior to templates comprising a nucleic acid sequence encoding a nucleic acid sequence without poly(A) tail or a standard non-segmented poly(A) tail when used for IVT.
- mRNA messenger RNA
- mRNA therapy can be used to restore normal levels of an endogenous protein or provide an exogenous therapeutic protein (e.g., a vaccine antigen or antibody) without permanently altering the genome sequence or entering the nucleus of the cell.
- mRNA therapy takes advantage of the cell’s own protein production and processing machinery to express a therapeutic peptide, polypeptide, or protein, is flexible to tailored dosing and formulation, and is broadly applicable to any disease or condition that is treatable through the provision of an exogenous protein.
- the process of manufacturing mRNA for use in therapy typically involves the in vitro transcription (IVT) of mRNA from a DNA template using a phage-derived DNA-dependent RNA polymerase.
- the DNA template is typically prepared by standard molecular biology techniques. Specifically, a nucleic acid sequence encoding a peptide, polypeptide or protein of interest is cloned into a multi-copy plasmid which is then propagated in Escherichia coli (E. colt) and purified for use as a template in an IVT reaction.
- poly(A) tail With the exception of replication-dependent histone transcripts, naturally occurring mRNAs include a poly(A) tail at their 3’ end.
- the poly(A) tail commonly consists of a homopolymeric segment of adenosines. In mammals, poly(A) tails have been reported to consist of approximately 250 nucleotides. In vivo, the poly(A) tail is typically enzymatically added to mRNAs. This process is known as polyadenylation and occurs during mRNA maturation. It involves cleavage at the 3’ end followed by the enzymatic addition of adenine bases by poly(A) polymerase. The poly(A) tail facilitates the transport of mRNA from the nucleus to the cytoplasm. It is considered important for stability of the mRNA in vivo and the efficient translation of the mRNA-encoded protein (Sachs, Curr Opin Cell Biol. 1990; 2(6): 1092-1098).
- templates comprising segmented poly(A) tails that have high stability during propagation in bacterial cells (e.g., with a stability of the segmented poly(A) tail above 90% after two passages in E. coli (assessed by Sanger sequencing) while also maintaining translation efficiency of the resulting mRNA in target cells in vivo.
- the present invention is based on the discovery that high stability of templates for the in vitro transcription (IVT) of mRNA can be achieved during propagation in bacterial cells when the nucleic acid sequence encoding the poly(A) tail comprises three or more segments of 10-60 adenosines, wherein at least two of said segments consist of 40 adenosines or less and each segment is separated from the next segment by a spacer, wherein at least two of the spacers consists of two nucleotides other than adenosine.
- the inventors found that, when using such templates, 100% stability of a plasmid comprising the template could be achieved during propagation of the template in bacterial cells such as E. coli.
- the inventors also found that the mRNA resulting from such templates are non-inferior in terms of translation efficiency in comparison to mRNAs comprising standard non-segmented poly(A) tails. Surprisingly, the inventors also observed that the segmented poly(A) tails of the mRNAs resulting from IVT of templates of the invention were consistently of the desired length, whereas the length of non-segmented poly(A) tails of correspondingly produced mRNAs varied significantly. Thus, the templates of the invention can usefully be employed in the large-scale manufacture of mRNA for therapeutic purposes to provide batches of mRNAs with segmented poly(A) tails of a consistent length.
- the invention relates to a template A.g, a DNA template such as a plasmid) for in vitro transcription of messenger RNA (mRNA), wherein the template comprises in the 5’ to 3’ direction a first nucleic acid sequence encoding a peptide, polypeptide or protein and a second nucleic acid sequence encoding a poly(A) tail comprising three or more segments of 10-60 adenosines, wherein at least two of said segments consist of 40 adenosines or less, wherein each segment is separated from the next segment by a spacer, and wherein at least two of the spacers independently consists of two nucleotides other than adenosine.
- further spacers independently consist of one or two nucleotides other than adenosine.
- the poly(A) tail comprises three segments.
- the poly(A) tail further comprises a fourth segment of less than 10 adenosines.
- the fourth segment is at the 3’ end of the poly(A) tail.
- the fourth segment consists of less than 5 adenosines (e.g., 3 adenosines).
- each of the three or more segments consists of 25- 40 adenosines. In other embodiments, the at least two of the three or more segments consist of 20 adenosines or less (e.g., 18 or 19 adenosines). In one embodiment, the template comprises one segment of 50-60 adenosines. In another embodiment, the template comprises at least 5 segments (e.g., 6 segments) of 20 adenosines or less. In some embodiments, one or more of the segments comprise(s) different numbers of adenosines. In some embodiments, one or more of the segments comprise(s) the same number of adenosines.
- each of the segments comprises a different number of adenosines. In some embodiments, each of the segments comprises the same number of adenosines. In some embodiments, the number of adenosines per segment decreases in a 5’ to 3’ direction.
- the non-adenosine nucleotides of each spacer independently are guanosine or cytidine. In one embodiment, the non-adenosine nucleotides of each spacer are guanosine. In another embodiment, the non-adenosine nucleotides of each spacer are cytidine. In some embodiments, the at least two of the spacers independently consist of two guanosines or two cytidines. In one embodiment, the at least two spacers consist of two guanosines. In another embodiment, the at least two spacers consist of two cytidines. In some embodiments, at least one of the further spacers consists of one guanosine or one cytidine.
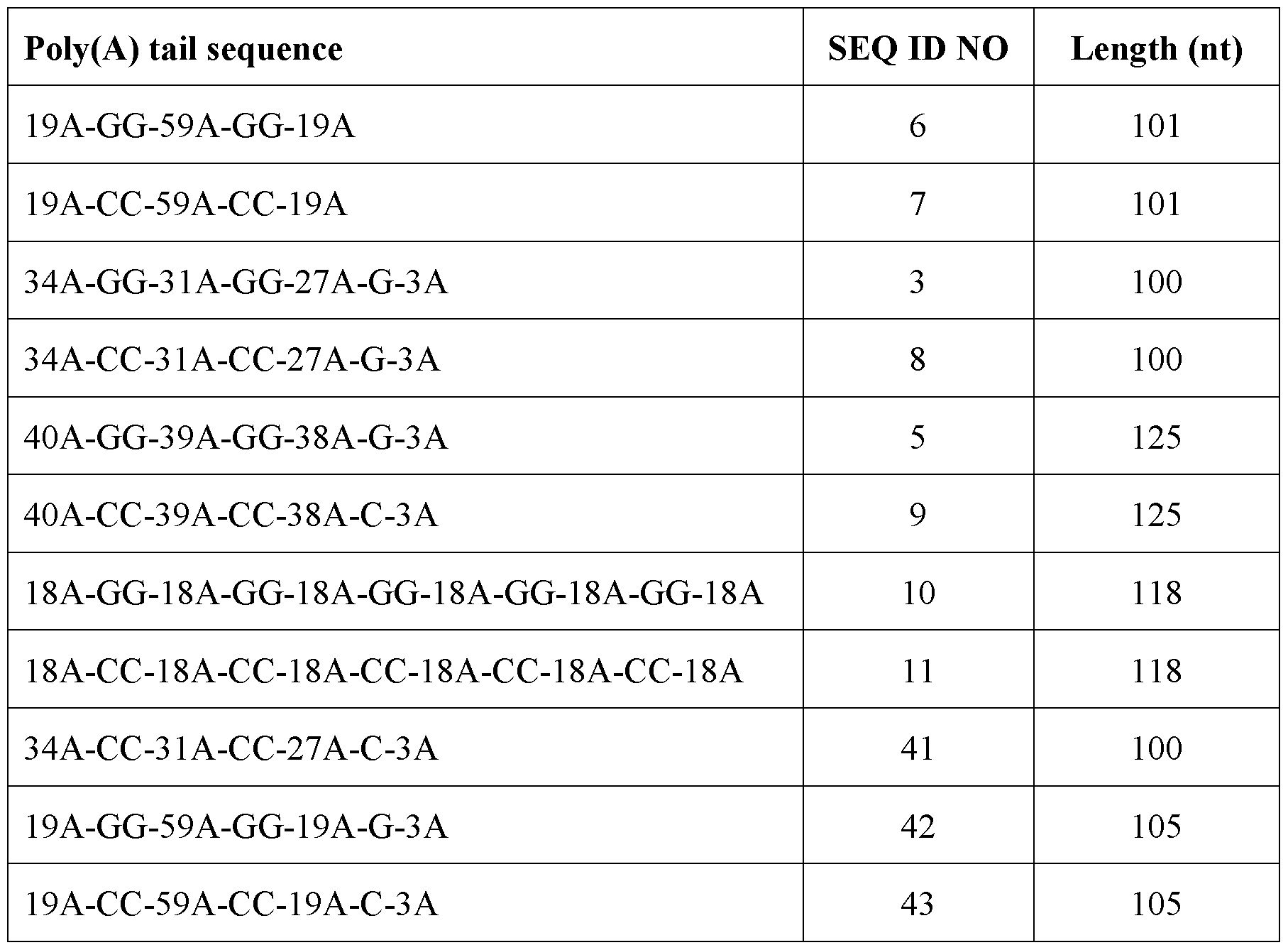
- the poly(A) tail sequence comprises or consists of the nucleic acid sequence 34A-GG-31A-GG-27A-G-3A (SEQ ID NO: 3). In some embodiments, the poly(A) tail sequence comprises or consists of the nucleic acid sequence 40A-GG-39A-GG- 25A-T-12A-G-3A (SEQ ID NO: 4). In some embodiments, the poly(A) tail sequence comprises or consists of the nucleic acid sequence 40A-GG-39A-GG-38A-G-3A (SEQ ID NO: 5).
- the poly(A) tail sequence comprises or consists of the nucleic acid sequence 34A-GG-31A-GG-27A-G-3A (SEQ ID NO: 3). [18] In some embodiments, the poly(A) tail is composed of 150 nucleotides or less. In particular embodiments, the poly(A) tail is composed of 80-150 nucleotides, e.g., 100-125 nucleotides.
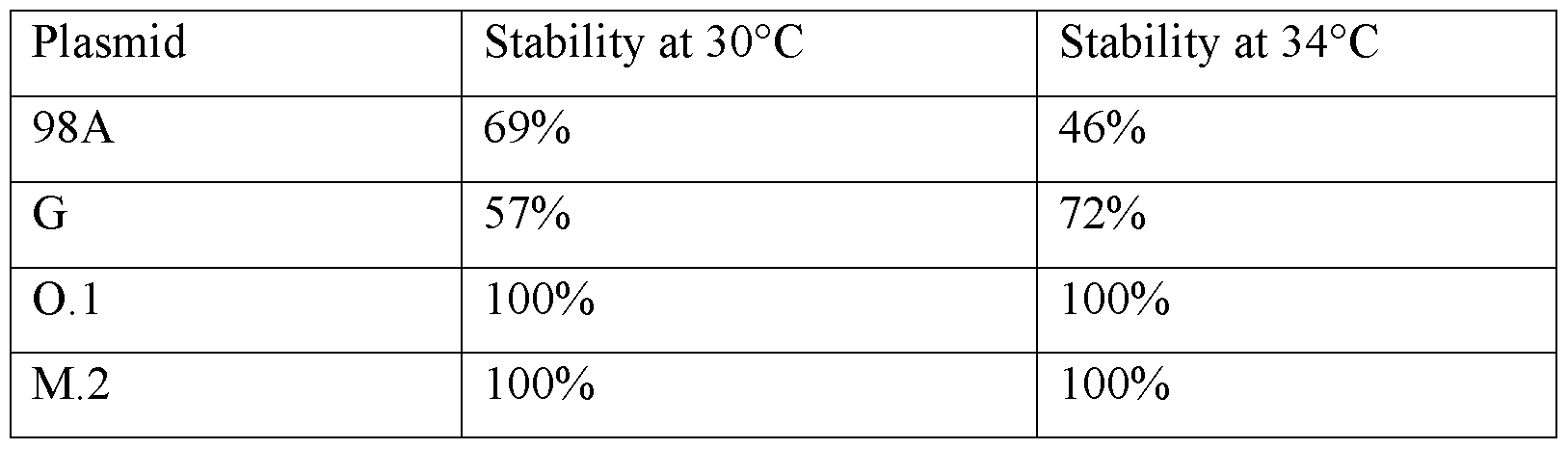
- the template is capable of being stably propagated in a bacterial cell such as E. coli at 30°C and/or 34°C.
- a segmented poly(A) tail in accordance with the invention has a stability of 90% or greater, e.g., 92% or greater, 94% or greater, 95% or greater, 96% or greater, 97% or greater, 98% or greater, or 99% or greater during propagation in a bacterial cell such as E. coli.
- the template is capable of being propagated in E. coli cells at 30°C and/or 34°C without any loss of the poly(A) tail.
- the template is capable of being propagated in E.
- the template is capable of being propagated in E. coli cells at 30°C or 34°C without loss.
- the template is a ColEl plasmid.
- the ColEl plasmid comprises a Cer sequence.
- the first nucleic acid sequence comprises a 5’ untranslated sequence (5’ UTR) and/or a 3’ untranslated sequence (3’ UTR).
- the template comprises a polymerase promoter operably linked to the first and second nucleic acid sequences.
- the promoter is an SP6, T7 or KP34 phage polymerase promoter.
- the invention also relates to a method of producing mRNA comprising incubating a template in accordance with the present invention in the presence of an RNA polymerase under conditions suitable for in vitro transcription (IVT).
- IVTT in vitro transcription
- the invention provides a method of making an mRNA molecule, the method comprising (a) providing a template in accordance with the present invention, and (b) contacting the template with an RNA polymerase and a reaction mixture under conditions suitable for in vitro transcription of the first and second nucleic acid sequences.
- the method further comprises adding a 5’ cap to the resulting mRNA molecule.
- the method comprises a 5 ’cap analogue in the IVT reaction mixture.
- the invention also relates to a method of improving stability of a homopolymeric sequence comprising at least 80 adenosines during propagation in a bacterial cells (e.g., E.
- the method comprises modifying the homopolymeric sequence to comprise three or more segments of 10-60 adenosines, wherein at least two of said segments consist of 40 adenosines or less, wherein each segment is separated from the next segment by a spacer, wherein at least two of the spacers independently consists of two nucleotides other than adenosine.
- the homopolymeric sequence is comprised in a vector suitable for propagation in a bacterial cell.
- the method further comprises transforming a bacterial cell with the vector comprising the modified homopolymeric sequence.
- the invention further relates to a population of mRNAs obtainable by a method of making or producing an mRNA molecule described herein.
- at least 80% of the mRNAs in a population of mRNAs obtained in accordance with the present invention comprise a poly(A) tail of the desired length.
- the invention also relates to an mRNA molecule with a 3’ poly(A) tail comprising at three or more segments of 10-60 adenosines, wherein at least two of said segments consist of 40 adenosines or less, wherein each segment is separated from the next segment by a spacer, and wherein at least two of the spacers independently consists of two nucleotides other than adenosine.
- further spacers independently consist of one or two nucleotides other than adenosine.
- the poly(A) tail comprises three segments.
- the poly(A) tail further comprises a fourth segment of less than 10 adenosines.
- the fourth segment is at the 3’ end of the poly(A) tail.
- the fourth segment consists of less than 5 adenosines (e.g., 3 adenosines).
- the mRNA comprises three segments, each consisting of 25-40 adenosines. In another embodiment, at least two of the three or more segments consist of 20 adenosines or less (e.g., 18 or 19 adenosines). In one embodiment, the poly(A) tail comprises at least one segment of 50-60 adenosines. In another embodiment, the poly(A) tail comprises at least 5 segments (e.g., 6 segments) of 20 adenosines or less. [31] In some embodiments, one or more of the segments comprise(s) different numbers of adenosines. In some embodiments, one or more of the segments comprise(s) the same number of adenosines.
- each of the segments comprises a different number of adenosines. In some embodiments, each of the segments comprises the same number of adenosines. In some embodiments, the number of adenosines per segment decreases in a 5’ to 3’ direction.
- the non-adenosine nucleotides of each spacer in an mRNA molecule of the invention independently are guanosine or cytidine. In one embodiment, the non-adenosine nucleotides of each spacer are guanosine. In another embodiment, the non- adenosine nucleotides of each spacer are cytidine. In some embodiments, the at least two spacers independently consist of two guanosines or two cytidines. In one embodiment, the at least two spacers consist of two guanosines. In another embodiment, the at least two spacers consist of two cytidines. In some embodiments, at least one of the further spacers consists of one guanosine or one cytidine.
- the poly(A) tail in an mRNA molecule of the invention is composed of 150 nucleotides or less.
- the poly(A) tail is composed of 80-150 nucleotides, e.g., 100-125 nucleotides.
- the invention also relates to a pharmaceutical composition
- a pharmaceutical composition comprising an mRNA molecule in accordance with the invention and a pharmaceutically acceptable carrier.
- the mRNA molecule comprised in the pharmaceutical composition is encapsulated in a lipid nanoparticle.
- the composition is an immunogenic composition (e.g., a vaccine).
- the immunogenic composition is used for immunizing a subject.
- the immunogenic composition prevents a virus infection in the subject, decreases the subject’s likelihood of contracting a virus infection, or reduces the subject’s likelihood of getting seriously ill from a virus infection.
- the immunogenic composition raises a protective immune response in the subject.
- the subject is a human.
- the human is 6 months of age or older, less than 18 years of age, at least 6 months of age and less than 18 years of age, at least 18 years of age and less than 65 years of age, at least 6 months of age and less than 5 years of age, at least 5 years of age and less than 65 years of age, at least 60 years of age, or at least 65 years of age.
- immunization is performed intramuscularly, intradermally, subcutaneously, intravenously, intranasally, by inhalation, or intraperitoneally.
- FIG. 1 schematically outlines an assay that can be used to determine plasmid stability in a bacterial cell such as Escherichia coli (E. coif).
- a glycerol stock of a bacterial clone comprising a plasmid of interest is thawed and used to inoculate a preculture.
- the preculture is incubated for 8 hours at 30°C.
- This preculture is then used to inoculate a liquid culture Pl which comprises a growth medium and an antibiotic to maintain the plasmid in the bacterial cell (e.g., 25 mL Luria-Bertani [LB] broth + kanamycin [Kana] at 25 pg/ml as indicated in the figure).
- Pl comprises a growth medium and an antibiotic to maintain the plasmid in the bacterial cell
- a suitable solid culture medium e.g., LB medium + kanamycin at 25 pg/mL.
- 16 to 24 colonies are isolated from each plate and cultured in 1.2 mL of liquid culture medium (e.g., LB broth supplemented with kanamycin at 25 pg/mL).
- liquid culture medium e.g., LB broth supplemented with kanamycin at 25 pg/mL.
- plasmid DNA is purified from the resulting 16 to 24 cultures, e.g., by miniprep plasmid purification.
- Figure 2 shows the results of a plasmid stability assay performed as illustrated in Figure 1.
- the stability of plasmid templates encoding a segmented poly(A) tail or nonsegmented poly(A) tail is compared.
- Each filled black circle depicts the length of the poly(A) tail measured by sequencing of a template plasmid.
- Each plasmid was obtained from a colony isolated from a P2 culture as shown in Figure 1.
- Figure 3 illustrates the impact of segmented and non-segmented poly (A) tails on intracellular stability and translation efficiency of mRNA transcripts encoding a eGFP degron.
- Figure 3A shows the mean fluorescence intensity (MFI) per cell for eGFP degron expression at different time-points post-transfection of various mRNA transcripts as indicated. The MFI values were corrected for MFI background.
- Figure 3B shows the translation efficiency and intracellular stability of the tested mRNA transcripts calculated based on eGFP expression data. As can be seen from this figure, most of the values clustered in the lower left corner of the plot. Only mRNA transcripts comprising the segmented 0.1 poly(A) tail sequence stood out by having both exceptional intracellular stability and very high translation efficiency.
- Figure 4 illustrates the reactogenicity of mRNAs comprising a segmented or nonsegmented poly(A) tail as determined in THPl-DualTM cells. These cells comprise an NF-KP inducible reporter gene encoding secreted alkaline phosphatase (SEAP) and an interferonsensitive response element (ISRE)-inducible reporter gene encoding Lucia luciferase.
- SEAP secreted alkaline phosphatase
- ISRE interferonsensitive response element
- Figure 4B shows the optical density of secreted alkaline phosphatase (SEAP) within supernatants obtained from cells electroporated with HAH3 mRNA transcripts comprising segmented or non-segmented poly (A) tails as indicated. Optical density was measured at 620 nm. Bars in Figures 4A and 4B represent mean value ⁇ SEM. The supernatant from cells electroporated with mRNA transcripts encoding eGFP was included as a positive control. Reactogenicity of the tested mRNA transcripts was minimal. The presence of segmented poly(A) tails on mRNA transcripts did not increase reactogenicity relative to non-segmented poly(A) tails. DEFINITIONS
- a ribonucleotide is understood to represent one or more ribonucleotides.
- the terms “a” (or “an”), “one or more”, and “at least one” can be used interchangeably herein.
- the term "and/or" as used in a phrase such as “A, B, and/or C” is intended to encompass each of the following aspects: A, B, and C; A, B, or C; A or C; A or B; B or C; A and C; A and B; B and C; A (alone); B (alone); and C (alone).
- the term “about” refers to an interval of accuracy that a person skilled in the art will understand to still ensure the technical effect of the feature in question.
- the term indicates a deviation from the indicated numerical value of ⁇ 10%. In some embodiments, the deviation is ⁇ 5% of the indicated numerical value. In certain embodiments, the deviation is ⁇ 1% of the indicated numerical value.
- mRNA refers to a polyribonucleotide that encodes at least one peptide, polypeptide, or protein.
- mRNA as used herein encompasses both modified and unmodified RNA.
- mRNA may contain one or more coding and non-coding regions (e.g., a 5’ untranslated region and a 3’ untranslated region).
- mRNA can be purified from natural sources, produced using recombinant expression systems and optionally purified, in vitro transcribed, or chemically synthesized.
- the invention particularly relates to the manufacturing of mRNA by in vitro transcription (IVT).
- mRNA can comprise nucleotide analogues such as analogues having chemically modified bases or sugars, backbone modifications, etc.
- An mRNA sequence is presented in the 5’ to 3’ direction unless otherwise indicated.
- a typical mRNA comprises a 5’ cap, a 5’ untranslated region (5’ UTR), a protein-coding region, and a 3’ untranslated region (3’ UTR).
- a poly(A) tail is typically added to or included in in vitro transcribed mRNAs to mimic naturally occurring mRNAs. The presence of a poly(A) tail at the 3’ end of an mRNA molecule prevents degradation inside cells, thereby enhancing its half-life in vivo.
- nucleic acid sequence encoding a polypeptide refers to any nucleic acid sequence that encodes an amino acid sequence which, upon expression, forms a peptide, polypeptide, or protein.
- a nucleic acid sequence may further comprise a 5’- untranslated region (5’ UTR) and 3 ’-untranslated region (3 ’-UTR).
- poly(A) refers to a nucleic acid comprising one or more homopolymeric segments of adenosines typically located at the 3’ end of an mRNA molecule.
- the homopolymeric segments of adenosines may be separated from each other by spacers comprising one or more nucleotides other than adenosine.
- Poly(A) tails comprising one or more spacers are referred to herein as “segmented poly(A) tails” to distinguish them from poly(A) tails that consist entirely of a single homopolymeric region of adenosines (referred to herein as “non-segmented poly (A) tails”).
- reference template and “reference plasmid” refers to a DNA template or plasmid that serves as a comparator to assess, e.g., stability during propagation, and differs from a template or plasmid of the invention only by a single sequence element, typically the nucleic acid sequence encoding the poly(A) tail.
- the term “spacer” refers to one or more nucleotides other than adenosine within the poly(A) tail. In some embodiments, the spacer consists of either one or two consecutive cytidine(s) or guanosine(s).
- sequence-optimized is used to describe a nucleotide sequence that is modified relative to a naturally occurring or wild-type nucleotide sequence. In the case of a sequence-optimized mRNA, such modifications may include, e.g., codon optimization and/or the use of 5’ UTRs and 3’ UTRs which are not normally associated with the naturally occurring or wild-type nucleic acid.
- cognitivation optimization and “codon-optimized” refer to modifications of the codon composition of a naturally occurring or wild-type nucleic acid encoding a peptide, polypeptide or protein that do not alter its amino acid sequence, thereby improving protein expression of said nucleic acid.
- “codon optimization” may also refer to the process by which one or more optimized nucleotide sequences are arrived at by removing, with filters, less than optimal nucleotide sequences from a list of nucleotide sequences, such as filtering by guanine-cytosine (GC) content, codon adaptation index (CAI), presence of destabilizing nucleic acid sequences or motifs, and/or presence of pause sites and/or terminator signals.
- GC guanine-cytosine
- CAI codon adaptation index
- the term “template” refers to a molecule (typically a DNA molecule such as a plasmid) comprising a nucleic acid sequence encoding an mRNA transcript to be synthesized by in vitro transcription (IVT).
- the template is used as template for IVT in order to produce a mRNA transcript encoded by the template.
- the template comprises all elements necessary for IVT, particularly a promoter element for binding of an RNA polymerase, which is operably linked to the nucleic acid sequence encoding a desired mRNA transcript.
- the template may be a linear or a circular DNA molecule that is linearized before IVT.
- the term “batch” refers to a quantity or amount of mRNA synthesized at one time, e.g., produced according to a single manufacturing order during the same cycle of manufacture.
- a batch may refer to a population of mRNAs synthesized in one reaction that occurs via a single aliquot of enzyme and/or a single aliquot of template for continuous synthesis under one set of conditions.
- the term “batch” does not refer to populations of mRNAs synthesized at different times and/or in different reactions that are combined to achieve a desired amount.
- the term “expression” of a nucleic acid sequence refers to translation of an mRNA into a peptide, polypeptide, or protein in vitro or in vivo.
- the term “therapeutic” refers to any pharmaceutical, drug, or composition that can be used to treat or prevent a disease, illness, condition, or disorder of bodily function.
- the term “substantially” refers to the qualitative condition of exhibiting total or near-total extent or degree of a characteristic or property of interest.
- One of ordinary skill in the biological arts will understand that biological and chemical phenomena rarely, if ever, go to completion and/or proceed to completeness or achieve or avoid an absolute result.
- the term “substantially” is therefore used herein to capture the potential lack of completeness inherent in many biological and chemical phenomena.
- zzz vitro refers to events that occur in an artificial environment, e.g., in a test tube or reaction vessel, or in a cell in cell culture, etc., rather than within a multi-cellular organism.
- zzz vzvo refers to events that occur within a multi-cellular organism, such as a human or a non-human animal.
- isolated refers to a substance and/or entity that has been (1) separated from at least some of the components with which it was associated when initially produced (whether in nature and/or in an experimental setting), and/or (2) produced, prepared, and/or manufactured by the hand of man.
- the term “immunogenic composition” refers to a composition that generates an immune response that may or may not be a protective immune response or protective immunity.
- the term “immune response” refers to a response of a cell of the immune system, such as a B cell, T cell, dendritic cell, macrophage or polymorphonucleocyte, to a stimulus such as an antigen, immunogen, or vaccine.
- An immune response can include any cell of the body involved in a host defense response, including for example, an epithelial cell that secretes an interferon or a cytokine.
- An immune response includes, but is not limited to, an innate and/or adaptive immune response.
- lymphocytes such as B or T cells
- An antibody response or humoral response is an immune response in which antibodies are produced.
- a “cellular immune response” is one mediated by T cells and/or other white blood cells.
- a “protective immune response” or “protective immunity” refers to an immune response that protects a subject from infection (prevents infection or prevents the development of disease associated with infection) or reduces the symptoms of infection (for instance, an infection by an influenza virus). Vaccines may elicit both prophylactic (preventative) and therapeutic responses.
- the present invention relates to templates for in vitro transcription (IVT) of messenger RNA (mRNA).
- Such templates typically may comprise in 5’ to 3’ direction a first nucleic acid sequence encoding a polypeptide and a second nucleic acid encoding a poly(A) tail.
- mRNAs used in therapy commonly include a poly(A) tail at their 3’ end to prevent rapid degradation of the mRNAs inside cell in vivo and to improve expression of the encoded peptide, polypeptide, or protein. It is often advantageous to use co-transcriptional tailing, e.g., because it avoids the costs associated with a separate enzymatic tailing step, and in theory reduces variability of the poly(A) tail length and hence reproducibility.
- the inventors have surprisingly found that templates were maintained faithfully during propagation in bacterial cells when the poly(A) tail-encoding nucleic acid sequence comprised three segments of 10-60 adenosines, of which at least two consisted of 40 adenosines or less, wherein each segment was separated from the next segment by a spacer, of which at least two independently consisted of two nucleotides other than adenosine.
- the inventors found that mRNAs obtained by in vitro transcription (IVT) of such templates were translated as efficiently as mRNA with a standard non-segmented poly(A) tail. Unexpectedly, the resulting population of mRNAs consistently had poly(A) tails of the desired lengths.
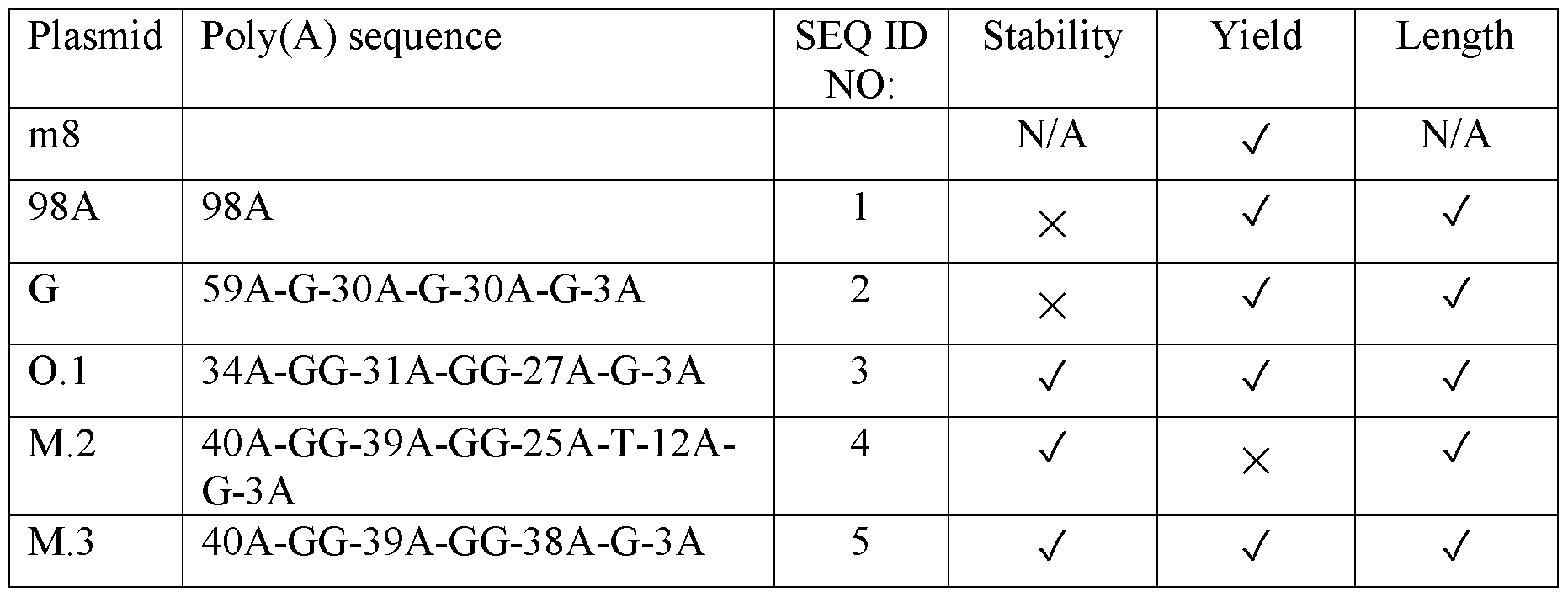
- templates encoding a poly(A) tail sequence comprising the nucleic acid sequence as set forth in SEQ ID NO: 3-5 were stably maintained during propagation in bacterial cells, produced acceptable yields of mRNA transcripts with poly(A) tails having the expected length and were minimally reactogenic. Higher mRNA transcript yields were achieved with templates encoding a poly(A) tail sequence comprising the nucleic acid sequence of SEQ ID NO: 3 or SEQ ID NO: 5.
- mRNA transcripts comprising the segmented poly(A) tail sequence of SEQ ID NO: 3 were found to have higher intracellular stability and translation efficiency.
- the invention relates to a template for in vitro transcription (IVT) of messenger RNA (mRNA), wherein the template comprises in 5’ to 3’ direction a first nucleic acid sequence encoding a polypeptide and a second nucleic acid sequence encoding a poly(A) tail comprising three or more segments of 10-60 adenosines, wherein at least two of said segments consist of 40 adenosines or less, wherein each segment is separated from the next segment by a spacer, and at least two of the spacers independently consist of two nucleotides other than adenosine.
- IVTT in vitro transcription
- further spacers independently consist of one or two nucleotides other than adenosine.
- the poly(A) tail comprises three segments. [71] In some embodiments, the poly(A) tail further comprises a fourth segment of less than 10 adenosines. In some embodiments, the fourth segment is at the 3’ end of the poly(A) tail. In some embodiments, the fourth segment consists of less than 5 adenosines (e.g., 3 adenosines).
- each of the three segments consists of 25-40 adenosines. In other embodiments, the at least two of the three segments consist of 20 adenosines or less (e.g., 18 or 19 adenosines). In one embodiment, the template comprises one segment of 50- 60 adenosines. In another embodiment, the template comprises at least 5 segments (e.g., 6 segments) of 20 adenosines or less.
- the number of adenosines in the three segments is about the same or the same.
- the first, second and third segments in 5’ to 3’ direction each consist of about 40 adenosines (e.g., 40, 39 and 38, respectively).
- the number of adenosines in each segment is different.
- the number of adenosines in at least one of the three (or four) segments is different to the number of adenosines in the other two (or three) segments.
- the first segment in 5’ to 3’ direction comprises more than 30 adenosines, e.g., 40 adenosines or more (e.g., 34 to 59), whereas the second, third and optional fourth segment comprise fewer adenosines (e.g., 30 adenosines or less).
- the first and third segments consist of 20 adenosines or less (e.g., 19, 18, 17, 16 or 15), whereas the second segment consists of 60 adenosines or less (e.g., 59, 58, 56, or 55).
- At least two spacers in a segmented poly(A) tail in accordance with the invention may consist of two nucleotides other than adenosine. Further spacers may independently consist of one or two nucleotides other than adenosine.
- the nucleotide other than adenosine is guanosine.
- the nucleotide other than adenosine is cytidine.
- the use of cytidine(s) in the spacers may advantageously be employed to boost expression of the polypeptide encoded by an mRNA comprising a poly(A) tail of the invention.
- the same type of nucleotide either guanosine or cytidine
- some spacers consist of cytidine(s) and some spacers consists of guanosine(s).
- further spacers in the poly(A) tail consist of a single nucleotide (e.g., a guanosine or a cytidine).
- each spacer in the poly(A) tail consists of two nucleotides (e.g., two guanosines or two cytidines).
- the poly(A) tail comprises at least two spacers consisting of two nucleotides (e.g., a guanosine or a cytidine) and at least one spacer consisting of a single nucleotide (e.g., a guanosine or a cytidine).
- the at least two spacers consist of two cytidines, and the at least one spacer of a single cytidine.
- the at least two spacers consist of two guanosines, and the at least one spacer of one guanosine.
- the poly(A) tail comprises at least one spacer consisting of two cytidines and at least one spacer consisting of two guanosines. In some embodiments, the poly(A) tail comprises at least one further spacer consisting of a single cytidine or a single guanosine.
- a segmented poly(A) tail in accordance with the invention consists of four segments, wherein each of the first three segments in the 5’ to 3’ direction consist of 25-40 adenosines.
- the fourth segment comprises less than 10 (e.g., less than 5) adenosines and is separated from the third segment by a spacer consisting of a single nucleotide other than adenosine.
- the same type of non-adenosine nucleotide is used in each spacer.
- a poly(A) tail comprising 60 and 150 nucleotides (e.g., 90-120 nucleotides) is sufficient to maintain mRNA stability in vivo.
- the length of the poly(A) sequence of the template is less than 150 nucleotides e.g., less than 145 nucleotides, less than 140 nucleotides, less than 135 nucleotides, less than 130 nucleotides, less than 125 nucleotides, less than 120 nucleotides, less than 115 nucleotides, less than 110 nucleotides, less than 105 nucleotides, less than 100 nucleotides, less than 99 nucleotides, less than 98 nucleotides, less than 95 nucleotides, less than 90 nucleotides, less than 85 nucleotides or 80 nucleotides.
- the average length of a poly(A) tail of an in vitro transcribed mRNA prepared in accordance with the invention is 80-150 nucleotides e.g., 90-125 nucleotides or 100-120 nucleotides. Size and arrangement of segments
- the poly(A) tail comprises or consists of a nucleic acid of formula I, wherein A is a segment consisting of adenosines and S is a spacer consisting of cytidine(s) and/or guanosine(s):
- nl, n2 and n3 are the number of adenosines in the respective segments, wherein each of nl, n2 and n3 is independently selected from 19 to 59, ml and m2 are the number of nucleotides in the respective spacers, wherein each of ml and m2 is 2.
- each of nl, n2 and n3 of formula I is independently selected from 35 to 40. In some embodiments, each of nl, n2 and n3 of formula I is independently selected from 25-35. In some embodiments, nl is greater than 30, and n2 and n3 are less than 40. In some embodiment, each of nl and n3 of formula I is less than 20 (e.g., 15-19), and n2 is more than 40 (e.g., 50-60).
- nl is 34, n2 is 31, n3 is 27, S is independently selected from guanosine and cytosine.
- formula 1 is followed by a single nucleotide spacer (e.g., G or C) and 3 adenosines, e.g., GAAA.
- each of the spacers consist of guanosines. In other embodiments, each of the spacers consist of cytidines.
- the poly(A) tail comprises or consists of a nucleic acid of formula II, wherein A is a segment consisting of adenosines and S is a spacer consisting of cytidine(s) and/or guanosine(s):
- nl, n2, n3 and n4 are the number of adenosines in the respective segments, nl is 34 to 59, n2 and n3 are independently selected from 27 to 39, n4 is 3, ml, m2 and m3 are the number of nucleotides in the respective spacers, wherein each of ml and m2 is 2 and m3 is 1 or 2.
- each of nl, n2 and n3 of formula II is independently selected from 35 to 40. In some embodiments, each of nl, n2 and n3 of formula II is independently selected from 25-35. In some embodiments, nl is greater than 30, and n2 and n3 are less than 40.
- each of the spacers consists of guanosine. In other embodiments, each of the spacers consists of cytidine.
- m3 of formula II is 1 and the spacer consists of a guanosine. In other embodiments, m3 is 1 and the spacer consists of a cytidine.
- nl is 34, n2 is 31, n3 is 27, n4 is 3, S is independently selected from guanosine and cytosine, ml and m2 are both 2, and m3 is 1.
- each of the spacers consist of guanosines. In other embodiments, each of the spacers consist of cytidines.
- the poly(A) tail comprises or consists of a nucleic acid of formula III, wherein A is a segment consisting of adenosines and S is a spacer consisting of cytidine(s) and/or guanosine(s):
- n is the number of adenosines in the respective segments, wherein n is independently selected from 15 to 18, m is the number of nucleotides in the respective spacers, wherein m is 2, and r is the number of repeats, wherein r is selected from 5 to 7.
- each instance of n in formula III is 15. In some embodiments, each instance of n in formula III is 18. [96] In some embodiments, r is 6. In some embodiments r is 7.
- each spacer consists of guanosines. In other embodiments, each spacer consists of cytidines.
- the poly(A) tail sequence comprises or consists of the nucleic acid sequence 34A-GG-31A-GG-27A-G-3A (SEQ ID NO: 3). In some embodiments, the poly(A) tail sequence comprises or consists of the nucleic acid sequence 40A-GG-39A-GG- 25A-T-12A-G-3A (SEQ ID NO: 4). In some embodiments, the poly(A) tail sequence comprises or consists of the nucleic acid sequence 40A-GG-39A-GG-38A-G-3A (SEQ ID NO: 5).
- mRNA transcripts comprising a poly(A) tail with the nucleic acid sequence set forth in SEQ ID NO: 3 were found to have exceptionally high intracellular stability and translation efficiency. This combination of high intracellular stability and translation efficiency is especially useful for in vivo therapeutic uses.
- the poly(A) tail sequence comprises or consists of the nucleic acid sequence 34A-GG-31A-GG- 27A-G-3A (SEQ ID NO: 3).
- Templates of the invention can easily be propagated in bacterial cells to produce large quantities for use in the in vitro transcription (IVT) of messenger RNA (mRNA).
- the template is a DNA template.
- a template of the invention is a plasmid.
- Plasmid backbones suitable for use with the invention are well-known to the skilled person and include the backbones of medium and high copy number plasmids routinely employed in the cloning and expression of gene sequences. These include pUC and pBR series of plasmids, including pUC19 and pBR322.
- a plasmid for use in the invention can be propagated in a bacterial cell such as E. coli at 100 copies or more without jeopardizing long-term plasmid maintenance and stability.
- the plasmid may comprise additional elements (e.g., ori and/or cer) that contribute to its stability during propagation in a bacterial cell.
- a template in accordance with the invention such as a plasmid comprises an origin of replication (ori).
- the ori can control replication and copy number of the template during propagation.
- the ori is chosen for combability with a host species such as a bacterial cell (e.g., E. coli).
- the ori is from pUC19, pMBl or pCoIEl.
- a vector backbone comprising an ori having the nucleotide sequence of SEQ ID NO: 12 is particularly advantageous for use with the invention:
- this ori comprises a stabilizing mutation, rendering plasmids containing this ori especially suitable for propagation in E. coli.
- a template such as a plasmid, includes a ColEl resolution (cer) sequence
- cer is a target site for multimer resolution in bacterial cells. Including a cer sequence prevents plasmid loss during propagation in bacterial cells such as E. coli.
- a template for use with the invention typically comprises one or more restriction site(s).
- the inclusion of one or more restriction sites may be useful for a variety of reasons.
- templates that are easily propagated in bacterial cells are typically circular, whereas in vitro transcription (IVT) is typically performed with linear templates to prevent transcriptional readthrough.
- a template of the invention may comprise a restriction site to linearize the template with a suitable restriction enzyme prior to IVT.
- the same template backbone is typically used for the production of various mRNAs, each of which encodes a different peptide, polypeptide or protein.
- the template backbone may include additional restriction sites to insert/replace the nucleic acid sequence encoding the peptide, polypeptide or protein.
- the restriction sites are typically located within the template in such a manner that the inserted nucleic acid sequence encoding the peptide, polypeptide or protein is flanked by a 5’ untranslated region (5’ UTR) and a 3’ untranslated region (3’ UTR) to yield an mRNA coding sequence that is operably linked to an RNA polymerase promoter.
- the template backbone may include one or more restriction sites to insert/replace a nucleic acid sequence encoding a poly(A) tail of the invention.
- the restriction site is positioned such that the nucleic acid sequence encoding the poly(A) tail is immediately 3’ to the 3’ UTR in the template.
- Suitable restriction sites include those that are cut by a type II restriction enzyme, such as a type IIP or type IIS restriction enzyme.
- the restriction enzyme is Xhol, EcoRI, BciVI, Spel, Xbal, Ndel, AflII, SacI, Kpnl, Smal, BamHI, Sall, Sbfl, PstI, BspQI (or its isoschizomers SapI), or Hindlll.
- a restriction site for a restriction enzyme such as Clal is used to insert a segmented poly(A) of the invention into a template such as a plasmid.
- the restriction site is transcribed into the mRNA molecule. The presence of the restriction site in the template or the mRNA molecule may be used for analytical purposes (e.g., to assess stability of the poly (A) sequence or the presence of the poly (A) tail).
- a template according to the invention may comprise a selection marker gene.
- a selection marker gene allows for the identification of cells that contain the template (e.g., during cloning).
- the template typically comprises a positive selection marker gene.
- a positive selection marker gene can encode a gene product (e.g., a protein or enzyme) that provides resistance to an antibiotic, enabling the selection of bacterial colonies carrying the template. Moreover, the presence of an antibiotic in the growth medium during propagation can aid in the maintenance of the template. Commonly used positive selection marker genes include kanamycin resistance (aph) and ampicillin resistance (bla) genes.
- a template of the invention e.g., a plasmid
- the templates of the invention typically include a suitable promoter sequence operably linked to the nucleic acid sequence encoding the peptide, polypeptide, or protein.
- a DNA-dependent RNA polymerase initiates transcription by contacting the promoter sequence.
- DNA-dependent RNA polymerases commonly used for IVT include, e.g., SP6 RNA polymerase, T7 RNA polymerase, T3 RNA polymerase and Klebsiella phage KP34 RNA polymerase.
- the promoter sequence is an SP6 RNA polymerase promoter sequence.
- the promoter sequence is a T7 RNA polymerase promoter sequence.
- the promoter sequence is a T3 RNA polymerase promoter sequence.
- the promoter sequence is a KP34 RNA polymerase promoter sequence.
- An SP6 promoter sequence may comprise 5' ATTTAGGTGACACTATAG-3' (SEQ ID NO: 13), or a variant thereof that is optimized for recognition and/or binding of SP6 RNA polymerase.
- Such variants may have the nucleic acid sequence 5'-ATTTAGGTGACACTATAGAAGNG-3' (SEQ ID NO: 14), where N is A, C, T or G. Exemplary variants are shown in Table 2.
- the SP6 promoter sequence is upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 34A-GG-31A-GG-27A-G- 3A (SEQ ID NO: 3). In some embodiments, the SP6 promoter sequence is upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 40A-GG-39A- GG-25A-T-12A-G-3A (SEQ ID NO: 4). In some embodiments, the SP6 promoter sequence is upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 40A-GG-39A-GG-38A-G-3 A (SEQ ID NO: 5). In particular embodiments, the SP6 promoter sequence is upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 34A-GG-31A-GG-27A-G-3A (SEQ ID NO: 3).
- the SP6 promoter comprises a sequence selected from the group consisting of SEQ ID NOs: 15-23 and is upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 34A-GG-31A-GG-27A-G-3A (SEQ ID NO: 3). In some embodiments, the SP6 promoter comprises a sequence selected from the group consisting of SEQ ID NOs: 15-23 and is upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 40A-GG-39A-GG-25A-T-12A-G-3A (SEQ ID NO: 4).
- the SP6 promoter comprises a sequence selected from the group consisting of SEQ ID NOs: 15-23 and is upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 40A-GG-39A-GG-38A-G-3A (SEQ ID NO: 5).
- the SP6 promoter comprises a sequence selected from the group consisting of SEQ ID NOs: 15-23 and is upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 34A-GG-31A-GG-27A-G-3A (SEQ ID NO: 3).
- T7 RNA polymerase Any promoter sequence that can be recognized by an T7 RNA polymerase may be used in the present invention.
- a suitable T7 promoter sequence comprises 5'- TAATACGACTCACTATAG-3' (SEQ ID NO: 24).
- the T7 promoter sequence is upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 34A-GG-31A-GG-27A-G- 3 A (SEQ ID NO: 3). In some embodiments, the T7 promoter sequence is upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 40A-GG-39A- GG-25A-T-12A-G-3A (SEQ ID NO: 4). In some embodiments, the T7 promoter sequence is upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 40A-GG-39A-GG-38A-G-3A (SEQ ID NO: 5). In particular embodiments, the T7 promoter sequence is upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 34A-GG-31A-GG-27A-G-3A (SEQ ID NO: 3).
- the T7 promoter sequence comprises SEQ ID NO: 24 and is upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 34A-GG-31A-GG-27A-G-3A (SEQ ID NO: 3). In some embodiments, the T7 promoter sequence comprises SEQ ID NO: 24 and is upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 40A-GG-39A-GG-25A-T-12A-G-3A (SEQ ID NO: 4).
- the T7 promoter sequence comprises SEQ ID NO: 24 and is upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 40A-GG-39A-GG-38A-G-3A (SEQ ID NO: 5).
- the T7 promoter sequence comprises SEQ ID NO: 24 and is upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 34A-GG-31A-GG-27A-G- 3A (SEQ ID NO: 3).
- a suitable Klebsiella phage KP34 core promoter sequence may comprise the nucleic acid sequence 5'-TAATGTTACAGGAGTA- 3' (SEQ ID NO: 25) or 5'-ATGTTACAGGAGTA-3' (SEQ ID NO: 26), or variants thereof
- Exemplary variants include TAATGTTACAGGAGTAGGnANiN 2 N3N 4 W (SEQ ID NO: 27), wherein NI- 4 is any one of A, C, G or T, wherein W is A or T, and wherein n is number of guanosines.
- N1N2N3N4 is CAGA.
- Exemplary variants are shown in Table 3.
- the KP34 promoter sequence is upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 34A-GG-31A-GG-27A-G- 3A (SEQ ID NO: 3). In some embodiments, the KP34 promoter sequence is upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 40A-GG-39A- GG-25A-T-12A-G-3A (SEQ ID NO: 4). In some embodiments, the KP34 promoter sequence is upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 40A-GG-39A-GG-38A-G-3A (SEQ ID NO: 5).
- the KP34 promoter sequence is upstream of a poly(A) tail sequence comprises or consists of the nucleic acid sequence 34A-GG-31A-GG-27A-G-3A (SEQ ID NO: 3).
- the KP34 promoter comprises a sequence selected from the group consisting of SEQ ID NOs: 28-33 and is upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 34A-GG-31A-GG-27A-G-3A (SEQ ID NO: 3).
- the KP34 promoter comprises a sequence selected from the group consisting of SEQ ID NOs: 28-33 and is upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 40A-GG-39A-GG-25A-T-12A-G-3A (SEQ ID NO: 4). In some embodiments, the KP34 promoter comprises a sequence selected from the group consisting of SEQ ID NOs: 28-33 and is upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 40A-GG-39A-GG-38A-G-3A (SEQ ID NO: 5).
- the KP34 promoter comprises a sequence selected from the group consisting of SEQ ID NOs: 28-33 and is upstream of a poly(A) tail sequence comprises or consists of the nucleic acid sequence 34A-GG-31A-GG-27A-G-3A (SEQ ID NO: 3 mRNA
- the templates of the invention are used for in vitro transcription (IVT) of messenger RNA (mRNA).
- the templates comprise in 5’ to 3’ direction a first nucleic acid sequence encoding a polypeptide and a second nucleic acid sequence encoding a poly(A) tail of the invention.
- the nucleic acid sequence encoding a polypeptide is optimized to facilitate more efficient transcription. For example, it may be optimized to remove cis- regulatory elements (e.g., termination signals, and protein binding sites), artificial recombination sites, chi sites, polymerase slippage sites, and/or other elements that could interfere with transcription. In some embodiments, the nucleic acid sequence encoding a polypeptide is optimized to reduce GC content or CpG dinucleotide content.
- the nucleic acid sequence encoding a polypeptide is optimized to remove cryptic splice sites, repetitive sequences, RNA instability motifs, and other elements relevant to mRNA processing and/or stability. In some embodiments, the nucleic acid sequence encoding a polypeptide is optimized to stabilize the mRNA secondary structure or stable free energy of the mRNA.
- An mRNA comprises a nucleotide sequence encoding a peptide, polypeptide or protein. It also typically comprises a 5’ untranslated sequence (5’ UTR) and a 3’ untranslated sequence (3’ UTR).
- the nucleotide sequences of a 5’ UTR or 3’ UTR encoded by a template of the invention is typically different from the 5’ UTR present in a naturally occurring mRNA encoding the peptide, polypeptide or protein in question.
- the 5’ UTR may include one or more elements that affect the mRNA’s stability or translation, e.g., an iron responsive element.
- the 5’ UTR may be about 50 to about 500 nucleotides in length.
- a 3’ UTR may include a binding site for proteins that affect an mRNA’s stability or location in a cell.
- a 3’ UTR may comprise one or more binding sites for a miRNA.
- a 3 ’ UTR may be about 50 to about 500 nucleotides in length.
- a 3’ UTR includes one or more poly(A)denylation signal(s).
- a poly(A)denylation signal typically comprises a hexamer e.g., AAUAAA.
- the 5’ and/or 3’ UTR sequences can be derived from mRNA which are stable (e.g., globin, actin, GAPDH, tubulin, histone, or citric acid cycle enzymes) to increase the stability of the mRNA.
- a 5’ UTR sequence may include a partial sequence of a CMV immediate-early 1 (IE1) gene or immediate-early gene 9 (IE9), or a fragment thereof, to improve nuclease resistance and/or half-life of the mRNA.
- IE1 CMV immediate-early 1
- IE9 immediate-early gene 9
- hGH human growth hormone
- An exemplary 5’ UTR may include a sequence derived from a CMV immediate-early 1 (IE1) gene (U.S. Publication Nos. 2014/0206753 and 2015/0157565, each of which is incorporated herein by reference), or the sequences provided in Example 1 of U.S. Publication No. 2016/0151409, incorporated herein by reference.
- an exemplary 5’ UTR may include a sequence derived from an immediate-early 9 (IE9) gene of the Towne strain of CMV.
- the 5’ UTR may be derived from the 5’ UTR of a TOP gene.
- TOP genes are typically characterized by the presence of a 5 ’-terminal oligopyrimidine (TOP) tract. Furthermore, most TOP genes are characterized by growth-associated translational regulation. However, TOP genes with a tissue specific translational regulation are also known.
- the 5’ UTR derived from the 5’ UTR of a TOP gene lacks the 5’ TOP motif (the oligopyrimidine tract) (e.g., U.S. Publication Nos. 2017/0029847, 2016/0304883, 2016/0235864, and 2016/0166710, each of which is incorporated herein by reference).
- the 5’ UTR is derived from a ribosomal protein Large 32 (L32) gene (U.S. Publication No. 2017/0029847, supra).
- the 5’ UTR is derived from the 5’ UTR of a hydroxysteroid (17-b) dehydrogenase 4 gene (HSD17B4) (U.S. Publication No. 2016/0166710, supra).
- the 5’ UTR is derived from the 5’ UTR of an ATP5A1 gene (U.S. Publication No. 2016/0166710, supra).
- an internal ribosome entry site (IRES) is used instead of a 5’ UTR.
- a 5’ UTR suitable for use with the invention may comprise one of the nucleic acid sequences shown in Table 4.
- the nucleic acid sequence comprised in the 5’ UTR is selected from any one of SEQ ID NOs: 34-39 and is located upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 34A-GG-31A-GG-27A-G-3A (SEQ ID NO: 3).
- the nucleic acid sequence comprised in the 5’ UTR is selected from any one of SEQ ID NOs: 34-39 and is located upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 40A-GG-39A-GG-25A-T-12A-G-3A (SEQ ID NO: 4).
- the nucleic acid sequence comprised in the 5’ UTR is selected from any one of SEQ ID NOs: 34-39 and is located upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 40A-GG-39A-GG-38A-G- 3A (SEQ ID NO: 5).
- the nucleic acid sequence comprised in the 5’ UTR is selected from any one of SEQ ID NOs: 34-39 and is located upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 34A-GG-31A-GG-27A-G- 3A (SEQ ID NO: 3).
- a 3’ UTR suitable for use with the invention may comprise the following nucleotide sequence:
- the nucleic acid sequence comprised in the 3’ UTR comprises SEQ ID NO: 40 and is located upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 34A-GG-31A-GG-27A-G-3A (SEQ ID NO: 3).
- the nucleic acid sequence comprised in the 3’ UTR comprises SEQ ID NOs: 40 and is located upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 40A-GG-39A-GG-25A-T-12A-G-3A (SEQ ID NO: 4).
- the nucleic acid sequence comprised in the 3’ UTR comprises SEQ ID NOs: 40 and is located upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 40A-GG-39A-GG-38A-G-3A (SEQ ID NO: 5).
- the nucleic acid sequence comprised in the 3’ UTR comprises SEQ ID NOs: 40 and is located upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 34A-GG-31A-GG-27A-G-3A (SEQ ID NO: 3).
- the nucleotide sequence encoding a peptide, polypeptide or protein may be optimized regarding codon usage bias, codon adaptability, internal chi sites, ribosomal binding sites (e.g., IRES), premature poly(A) sites, Shine-Dalgarno (SD) sequences, and/or other elements relevant to translation.
- the nucleotide sequence encoding a peptide, polypeptide or protein is optimized regarding codon context, codon-anticodon interaction, translational pause sites, and/or other elements relevant to protein folding.
- bacterial cells used to propagate the template are E. coli cells.
- the E. coli strain is K12-derived or JM109-derived, e.g., DH5a, DHIOb, Stbl2, Stbl3, NEBStableTM, or EnduraTM.
- the inventors found that high stability of templates for use in the in vitro transcription (IVT) of mRNA can be achieved during propagation in bacterial cells when the nucleic acid sequence encoding the poly (A) tail comprises three or more segments of 10-60 adenosines, wherein at least two of said segments consist of 40 adenosines or less, and wherein each segment is separated from the next segment by a spacer, wherein at least two of the spacers independently consists of two nucleotides other than adenosine.
- such templates were 100% stable during propagation in bacterial cells (specifically E. coli .
- Stability can be assessed by culturing a bacterial clone to provide a starter culture for a first time-period (e.g., 8 hours).
- the first time-period is typically selected in such a manner that the bacterial cells are in an exponential growth phase.
- the starter culture is then used to inoculate a first liquid culture.
- a second time-period of incubation e.g., 20 hours
- an aliquot of this first liquid culture is taken to inoculate a second liquid culture.
- the second liquid culture is incubated for a third time-period (e.g., 20 hours).
- the second and third timeperiods are typically selected in such a manner that the bacterial cells are entering a stationary phase.
- Each of the cultures comprise an appropriate growth medium (e.g., LB broth) and are incubated at a temperature suitable for growth (e.g., 30°C, 34°C, or 37°C).
- the growth medium may be supplemented with an antibiotic to maintain the template in the bacterial cell.
- an aliquot is taken from the first and second liquid cultures and plated on a suitable solid culture medium (e.g., after a 10-fold serial dilution). After a suitable incubation period (e.g., 16-24 hours), 10-24 individual bacterial colonies are isolated from each plate and cultured. After a suitable period of incubation (e.g., overnight), the template is isolated from each of the cultures.
- the purified template is then analyzed by sequencing or another suitable method, e.g., restriction fragment length polymorphism.
- the resulting data is compared to data obtained with the same analytical method for a reference template (typically the template isolated from the bacterial clone that was used to inoculate the starter culture).
- Figure 1 schematically outlines an assay that can be used to determine plasmid stability in abacterial cell such as A. coli.
- a bacteria cell such as A. coli.
- an 8-hour incubation period corresponds to 32 generations
- a 20-hour incubation period corresponds to 80 generations.
- the template (e.g., a DNA plasmid) is capable of being propagated in A. coli cells at 34°C with less than 30% loss, e.g., less than 10%, less than 5% or less than 1% loss. In some embodiments, the template is capable of being propagated in A. coli cells over at least about 70 generations with less than 30% loss, e.g., less than 20% loss, less than 15% loss, or less than 10% loss. For templates of the invention, loss is more typically less than 5%, e.g., less than 1%. In some embodiments, the template is capable of being propagated in a bacterial cell (e.g, A.
- a template of the invention is capable of being propagated in a bacterial cell (e.g., A. coli) for about 100-200 generations (e.g., about 150-200 generations) without loss or mutation to the nucleic acid sequence encoding the poly(A) tail.
- the template is capable of being propagated without loss or mutation to the nucleic acid sequence encoding the poly(A) tail in A. coli cells at 30°C or 34°C.
- nucleic acid encoding a segmented poly(A) tail confers comparable expression of the encoded protein compared to a nucleic acid encoding a non-segmented poly(A) tail.
- Quantitative methods for determining protein expression are known in the art, e.g., Western blotting, flow cytometry, enzyme-linked immunosorbent assay (ELISA) and mass spectrometry.
- IVTT In Vitro Transcription
- IVT is typically performed with a reaction mixture comprising a DNA template, a pool of ribonucleotide triphosphates, a buffering reagent (that may include DTT), and one or more salts (e.g., MgCh and NaCl).
- a typical IVT reaction buffer may also include spermidine. The exact conditions will vary according to the specific application.
- the IVT process may include a cap analogue which is included in the mRNA molecules co-transcriptionally.
- a cap analogue is included in the IVT reaction mixture.
- the cap analogue can be incorporated as the first “base” in a nascent RNA strand.
- the cap analogue may be Cap 0, Cap 1, Cap 2, m 6 Am, or a chemical cap analogue.
- the following chemical cap analogues may be used to generate the 5 ’-guanosine cap structure according to the manufacturer’s instructions: 3’-O-Me-m7G(5’)ppp(5’)G (the ARCA cap); G(5’)ppp(5’)A; G(5’)ppp(5’)G; m7G(5’)ppp(5’)A; m7G(5’)ppp(5’)G; m7G(5')ppp(5')(2'OMeA)-pG; m7G(5’)ppp(5’)(2’OmeA)pU; m7G(5’)ppp(5’)(2’OmeG)pG (New England BioLabs, Ipswich, MA; TriLink Biotechnologies).
- the in vitro transcribed mRNA may comprise a 5’ cap with the following structure: Modified RNA
- modified mRNA transcripts are synthesized with one or more modifications (z.e., as modified mRNA).
- modified mRNA comprises a modified ribonucleotide, such as ribonucleotide analogue (e.g., adenosine analogue, guanosine analogue, cytidine analogue, and/or uridine analogue).
- ribonucleotide analogue e.g., adenosine analogue, guanosine analogue, cytidine analogue, and/or uridine analogue.
- the presence of a modified ribonucleotide may render the mRNA more stable and/or less immunogenic than a control mRNA with the same sequence but containing only naturally occurring ribonucleotides.
- the modified ribonucleotide is selected from pseudouridine, N1 -methylpseudouridine, 2-thiouridine, 4’ -thiouridine, 5-methylcytidine, 2-thio-l -m ethyl- 1- deaza-pseudouridine, 2-thio-l-methyl-pseudouridine, 2-thio-5-aza-uridine, 2-thio- dihydropseudouridine, 2-thio-dihydrouridine, 2-thio-pseudouridine, 4-methoxy-2-thio- pseudouridine, 4-methoxy-pseudouridine, 4-thio-l-methyl-pseudouridine, 4-thio- pseudouridine, 5-aza-uridine, dihydropseudouridine, 5-methyluridine, 5-methoxyuridine, and 2’-O-methyl uridine.
- the modified ribonucleotide typically takes the place of a naturally occurring nucleotide.
- the in vitro transcribed mRNA of the invention comprises both unmodified and modified ribonucleotides.
- Such in vitro transcribed mRNA can be prepared by including a modified ribonucleotide in the IVT reaction mixture, typically in place of a naturally occurring ribonucleotide (e.g., N1 -methylpseudouridine in place of uridine).
- the in vitro transcribed mRNA may be RNA wherein 25% of uracil residues are 2-thio-uracil and 25% of cytosine residues are 5-methylcytosine.
- Teachings for the use of such modified RNA are disclosed in US Patent Publication US 2012/0195936 and international publication WO 2011/012316, both of which are hereby incorporated by reference in their entirety.
- the length of an IVT reaction may depend on the length of the mRNA transcript.
- the mRNA transcript comprises at least 500 ribonucleotides. In some embodiments, the mRNA transcript comprises about 500 to about 3,000 ribonucleotides. In some embodiments, the mRNA transcript comprises about 700 to about 3,000 ribonucleotides. In some embodiments, the mRNA transcript comprises about 800 to about 2,000 ribonucleotides. In some embodiments, the mRNA transcript comprises about 1,000 to about 2,000 ribonucleotides. In some embodiments, the mRNA transcript comprises less than 3,000 ribonucleotides.
- the period over which IVT may take place to synthesize mRNA can vary widely. In some embodiments, IVT takes place over a period of about thirty minutes to about six hours. In some embodiments, IVT takes place over a period of about sixty to about ninety minutes.
- IVT can be terminated by removing the DNA template, e.g., through the addition of DNase I and a suitable buffer.
- the polymerase reaction can be quenched by addition of DNase I and a DNase I buffer (100 mM Tris-HCl, 5 mM MgCh and 25 mM CaCh, pH 7.6 at lOx) to facilitate digestion of the double-stranded DNA template in preparation for purification.
- DNase I buffer 100 mM Tris-HCl, 5 mM MgCh and 25 mM CaCh, pH 7.6 at lOx
- the mRNA is synthesized in batches. In some embodiments, the present invention relates to the large-scale manufacture of mRNA.
- a batch comprises at least 1 g of in vitro transcribed mRNA (e.g., 5 g, 10 g, or 25 g). In other embodiments, a batch comprises at least 50 g of in vitro transcribed mRNA (e.g., 75 g, 100 g, 150 g, 200 g, or 250 g).
- a method according to the invention synthesizes at least 500 g, 750 g, 1 kg, 5 kg, 10 kg, 50 kg, 100 kg, 1000 kg, or more of mRNA in a single batch. In some embodiments, 10 kg mRNA or more is synthesized in a single batch. In some embodiments, between 10 kg and 100 kg of mRNA is synthesized in a single batch.
- Exemplary IVT conditions
- a suitable reaction mixture comprises a double-stranded DNA template of the invention, an RNA polymerase, RNase inhibitor, pyrophosphatase, NTPs, 10 mM DTT and a reaction buffer (29.6 mM Tris-HCl, 3.8 mM MgCh, 8.9 mM KC1, and pH 7.5).
- this reaction mixture is incubated at 37°C for the length of time needed to complete IVT of the mRNA transcript encoded by the DNA template.
- a reaction mixture includes each NTP at a concentration ranging from 1-10 mM, a DNA template at a concentration ranging from 0.01-0.5 mg/mL, and an RNA polymerase at a concentration ranging from 0.01-0.1 mg/mL.
- a 5’ cap structure can be added enzymatically after in vitro transcription (IVT) has been completed.
- IVT in vitro transcription
- at least 90% of in vitro transcribed mRNA subjected to separate enzymatic capping can comprise Capl structures.
- a 7-m ethylguanosine cap (also referred to as “m7G” or “Cap 0”), comprises a guanosine that is linked through a 5 ’-5 ’-triphosphate bond to the first transcribed nucleotide.
- a 5’ cap is typically added as follows: first, an RNA terminal phosphatase removes one of the terminal phosphate groups from the 5’ nucleotide, leaving two terminal phosphates; guanosine triphosphate (GTP) is then added to the terminal phosphates via a guanylyl transferase, producing a 5’5’5 triphosphate linkage; and the 7- nitrogen of guanine is then methylated by a methyltransferase.
- GTP guanosine triphosphate
- Examples of cap structures include, but are not limited to, m7G(5’)ppp, (5’(A,G(5’)ppp(5’)A, and G(5’)ppp(5’)G. Additional cap structures are described in U.S. Publication Nos. US 2016/0032356 and US 2018/0125989, which are incorporated herein by reference.
- a vaccinia virus capping enzyme may be used to generate the Cap 0 structure: m7G(5’)ppp(5’)G.
- a Cap 1 structure may be generated using both vaccinia virus capping enzyme and a 2’-0 methyl-transferase to generate: m7G(5’)ppp(5’)G-2’-O-m ethyl.
- a Cap 2 structure may be generated from the Cap 1 structure followed by the 2’-O-methylation of the 5 ’-antepenultimate nucleotide using a 2’-0 methyltransferase.
- a Cap 3 structure may be generated from the Cap 2 structure followed by the 2’-O-methylation of the 5’- preantepenultimate nucleotide using a 2’-0 methyltransferase.
- a method of producing mRNA that comprises incubating a template of the invention in the presence of an RNA polymerase under conditions suitable for in vitro transcription (IVT) may further comprise a separate step of capping the resulting in vitro transcribed mRNA.
- the capping step may involve adding a capping enzyme (guanylyltransferase) and a guanosine triphosphate (GTP).
- a suitable capping enzyme may be derived from a Vaccinia virus (Vaccine virus guanylyltransferase).
- the capping step also comprises adding a guanine methyltransferase and a 2 -O-m ethyltransferase. Capping may be performed separately, e.g., after IVT.
- the mRNA transcripts obtained from an IVT reaction and/or a separate capping step may be purified.
- Various methods may be used to purify mRNA.
- the mRNA transcripts are purified by precipitation and centrifugation.
- the mRNA is purified by filtration using, e.g., Normal Flow Filtration (NFF) or Tangential Flow Filtration (TFF).
- Suitable purification methods include those described in published U.S. Application Nos. US 2016/0040154, US 2015/0376220, US 2018/0251755, US 2018/0251754, US 2020/0095571, US 2021/0388338, and US 2021/0002635, and in International Patent Publication No. WO 2022/072836, all of which are incorporated by reference herein.
- a template encoding a segmented poly(A) tail of the invention yields mRNA transcripts with poly(A) tails having a single length distribution around the expected value
- a reference template encoding a standard non-segmented poly(A) tail can yield mRNA transcripts with poly(A) tails having a bimodal length distribution.
- the methods of the invention can be used to obtain a population of mRNA molecules with poly(A) tails having a single length distribution around a desired value.
- At least 80% of the mRNAs in a population comprise a poly(A) tail of the desired length e.g., at least 85%, at least 90%, or at least 95%. In some embodiments, at least 80% (e.g., at least 85%, 90% or 95%) of the mRNAs in the population are within 10 nucleotides or less (e.g., 5 nucleotides or less, or 3 nucleotides or less) of the desired length value.
- Methods for determining the length of mRNA molecules are well- known and include capillary gel electrophoresis. A suitable assay for measuring poly(A) tail length using a minor-groove binding dye and one or more ribonucleases is described in WO 2022/232499A1.
- the poly(A) tail is composed of 150 nucleotides or less e.g., 140 nucleotides or less, 130 nucleotides or less, 120 nucleotides or less, 110 nucleotides or less, 100 nucleotides or less, 90 nucleotides or less or 80 nucleotides or less.
- the poly(A) tail is composed of 80-150 nucleotides e.g., 100-125 nucleotides, 125-150 nucleotides, or 80-100 nucleotides.
- the desired tail length is at least 100 nucleotides (e.g., 100- 200 nucleotides).
- the desired tail length may be a particular value (e.g., 100 or 125 nucleotides), and the average length of the tailed mRNA may be within about 10% or about 5% of that value.
- At least 80% of the mRNA in a population is tailed and has an average tail length within 10% of the desired length.
- At least 80% (e.g., at least 85%, at least 90%, or at least 95%) of the mRNAs in a population comprise a Capl structure at the 5’ end.
- Suitable methods for determining the presence of a Capl structure are known in the art and include mass spectroscopy (see, e.g., WO 2014/152659A1).
- compositions comprising mRNA molecules of the invention may include one or more pharmaceutically acceptable excipients.
- pharmaceutically acceptable excipients e.g., one or more of a buffering such as Tris, a salt such as NaCl, or a sugar, e.g., a disaccharide such as sucrose or trehalose
- compositions comprising mRNA molecules of the invention may include one or more pharmaceutically acceptable excipients.
- Such excipients may be selected from a buffering agent (e.g., Tris), a salt (e.g., NaCl), and a sugar (e.g., a disaccharide such as sucrose or trehalose).
- a buffering agent e.g., Tris
- a salt e.g., NaCl
- a sugar e.g., a disaccharide such as sucrose or trehalose.
- compositions comprising an mRNA molecule in accordance with the invention may further comprise one or more agents that protect the mRNA molecule from rapid degradation and/or enhance delivery of the mRNA to target cells in vivo.
- agents include lipid nanoparticles and other carrier molecules (e.g., complex-forming carriers such as protamine).
- the mRNA molecule may be encapsulated in a lipid nanoparticle.
- Suitable lipid nanoparticles comprise a cationic lipid, one or more non-cationic lipids (e.g., DOPE and cholesterol) and a PEG-modified lipid (e.g., DMG-PEG2000).
- compositions disclosed herein can be formulated for administration in any way known in the art of drug delivery, for example, orally, parenterally, intravenously, intramuscularly, subcutaneously, intradermally, transdermally, intrathecally, submucosally, sublingually, rectally, vaginally, etc.
- the composition is formulated for sublingual administration, intramuscular administration, intradermal administration, subcutaneous administration, intravenous administration, intranasal administration, administration by inhalation, or intraperitoneal administration.
- the composition is formulated for sublingual administration.
- the composition e.g., an immunogenic composition such as a vaccine
- intramuscular injection e.g., an immunogenic composition such as a vaccine
- compositions disclosed herein such as pharmaceutical compositions may be packaged in a container, such as a prefilled syringe, a vial, or an autoinjector.
- a composition e.g., an immunogenic composition such as a vaccine
- the composition is packaged in a vial.
- the composition is in an autoinjector.
- the composition of the disclosure are packaged cartridges for patient-friendly autoinjector and infusion pump devices.
- a composition disclosed herein is an immunogenic composition or a vaccine.
- the polypeptide encoded by an mRNA molecule in accordance with the invention is an antigenic protein for use in an immunogenic composition or a vaccine, such as an antigenic protein from an influenza virus, a coronavirus, a RSV, a parainfluenza virus, a human immunodeficiency virus (HIV), a herpesvirus, a human papilloma virus, a rotavirus virus, a norovirus, a varicella zoster virus, a hepatitis virus, a paramyxovirus, a monkey pox virus, a parvovirus, an Ebola virus, a dengue virus, a hantavirus, a Zika virus, a west Nile virus, a poliovirus, or a rabies virus
- Immunogenic compositions are administered to a subject to prevent an infectious disease (e.g., virus infection, such as influenza, coronavirus, or a respiratory syncytial virus (RSV) infection) in the subject, to decrease the subject’s likelihood of contracting an infectious disease (e.g., virus infection), or to reduce the subject’s likelihood of getting serious ill from an infectious disease (e.g., virus infection, such as an influenza virus, coronavirus, or RSV infection).
- infectious disease e.g., virus infection, such as influenza, coronavirus, or a respiratory syncytial virus (RSV) infection
- RSV respiratory syncytial virus
- composition disclosed herein may be used for the manufacture of an immunogenic composition (e.g., a vaccine) for use in immunizing a subject against an infectious disease (e.g., virus infection, such as an influenza virus, coronavirus, or RSV infection).
- immunization comprises administering to a subject in need thereof an immunologically effective amount of any of the vaccines described herein.
- infectious disease e.g., virus infection, such as an influenza virus, coronavirus, or RSV infection.
- immunization comprises administering to a subject in need thereof an immunologically effective amount of any of the vaccines described herein.
- the term “immunologically effective amount” or “therapeutically effective amount” means an amount sufficient to immunize a subject.
- the disclosure provides a method of immunizing a subject comprising administering to the subject in need thereof an immunologically effective amount of any of the vaccines described herein.
- immunize or “immunizing” means to induce in a subject a protective immune response against an infectious disease (e.g., viral infection, such as influenza, coronavirus, or RSV infection).
- an infectious disease e.g., viral infection, such as influenza, coronavirus, or RSV infection
- a method of reducing one or more symptoms of an infectious disease e.g., viral infection, such as influenza, coronavirus, or RSV infection
- a prophylactically effective amount of any of an immunogenic composition e.g., a vaccine
- the methods and uses of the immunogenic compositions and vaccines described herein include administration of a single dose to a subject (z.e., no booster dose).
- the methods and uses of the vaccines described herein include prime-boost vaccination strategies.
- Prime-boost vaccination comprises administering a priming immunogenic composition (e.g., a priming vaccine) and then, after a period of time has passed, administering to the subject a boosting immunogenic composition (e.g., a boosting vaccine).
- the immune response is “primed” upon administration of the priming immunogenic composition (e.g., the priming vaccine) and is “boosted” upon administration of the boosting immunogenic composition (e.g., the boosting vaccine).
- the immunogenic compositions and vaccines described herein are administered to subjects, wherein the subjects can be any member of the animal kingdom.
- the subject is a non-human animal.
- the nonhuman subject is an avian (e.g., a chicken or a bird), a reptile, an amphibian, a fish, an insect, and/or a worm.
- the non-human subject is a mammal (e.g., a ferret, a rodent, a mouse, a rat, a rabbit, a monkey, a dog, a cat, a sheep, cattle, a primate, and/or a Pig)-
- the immunogenic compositions and vaccines described herein are administered to a human subject.
- a human subject is 6 months of age or older, 6 months through 35 months of age, at least two years of age, at least 3 years of age, 36 months through 8 years of age, 9 years of age or older, at least 6 months of age and less than 5 years of age, at least 6 months of age and less than 18 years of age, or at least 3 years of age and less than 18 years of age.
- the human subject is an infant (less than 36 months).
- the human subject is a child or adolescent (less than 18 years of age).
- the human subject is a child of at least 6 months of age and less than 5 years of age. In some embodiments, the human subject is at least 5 years of age and less than 60 years of age. In some embodiments, the human subject is at least 5 years of age and less than 65 years of age. In some embodiments, the human subject is elderly (at least 60 years of age or at least 65 years of age). In some embodiments, the human subject is a non-elderly adult (at least 18 years of age and less than 65 years of age or at least 18 years of age and less than 60 years of age).
- a template for in vitro transcription of messenger RNA wherein the template comprises in 5’ to 3’ direction a first nucleic acid sequence encoding a polypeptide and a second nucleic acid sequence encoding a poly(A) tail comprising three or more segments of 10-60 adenosines, wherein at least two of said segments consist of 40 adenosines or less, wherein each segment is separated from the next segment by a spacer, wherein at least two of the spacers independently consists of two nucleic acids other than adenosine.
- mRNA messenger RNA
- poly(A) tail further comprises a fourth segment of less than 10 adenosines.
- each of the three segments consists of 25-40 adenosines.
- poly(A) tail is composed of 150 nucleotides or less.
- poly(A) tail is composed of 80-150 nucleosides, e.g., 100-125 nucleosides.
- the template is a DNA template, e.g., a plasmid.
- template comprises a polymerase promoter operably linked to the first and second nucleic acid sequences.
- a method of producing mRNA comprising incubating the template of any one of the preceding embodiments in the presence of an RNA polymerase under conditions suitable for in vitro transcription.
- a method of improving stability of a homopolymeric sequence comprising at least 80 adenosines during propagation in a bacterial cells comprises modifying the homopolymeric sequence to comprise three or more segments of 10-60 adenosines, wherein at least two of said segments consist of 40 adenosines or less, wherein each segment is separated from the next segment by a spacer, wherein at least two of the spacers independently consists of two nucleic acids other than adenosine.
- a population of mRNAs obtainable by the method of embodiment 31.
- An mRNA molecule with a 3’ poly(A) tail comprising three or more segments of 10- 60 adenosines, wherein at least two of said segments consist of 40 adenosines or less, wherein each segment is separated from the next segment by a spacer, wherein at least two of the spacers independently consists of two nucleic acids other than adenosine.
- mRNA molecule of any one of embodiments 37-42 comprising three segments, each consisting of 25-40 adenosines.
- the mRNA molecule of embodiment 44, wherein the poly(A) tail comprises at least 5 segments (e.g., 6 segments) of 20 adenosines or less.
- the mRNA molecule of embodiment 50 wherein the at least two spacers consist of two guanosines. 52. The mRNA molecule of embodiment 50, wherein the at least two spacers consist of two cytidines.
- mRNA molecule of any one of embodiments 37-54, wherein the poly(A) tail is composed of 80-150 nucleosides, e.g., 100-125 nucleosides.
- a pharmaceutical composition comprising the mRNA molecule of any one of embodiments 37-55 and a pharmaceutically acceptable carrier.
- composition of embodiment 56 or 57, wherein the pharmaceutical composition is an immunogenic composition (e.g., a vaccine).
- the immunogenic composition according to embodiment 58 for use in immunizing a subject.
- the immunogenic composition for use according to embodiment 62 wherein the human is 6 months of age or older, less than 18 years of age, at least 6 months of age and less than 18 years of age, at least 18 years of age and less than 65 years of age, at least 6 months of age and less than 5 years of age, at least 5 years of age and less than 65 years of age, at least 60 years of age, or at least 65 years of age.
- Example 1 Generating templates with high stability during propagation in bacterial cells
- a template plasmid was prepared comprising a nucleic acid sequence encoding an mRNA.
- This nucleic acid sequence included the following elements: a 5’ untranslated region (5’ UTR), a protein coding sequence, and a 3’ untranslated region (3’ UTR).
- the 3’ UTR was followed by a multiple cloning site (MCS) for the insertion of poly(A) tail constructs.
- the template plasmid also included an RNA polymerase promoter (specific for SP6 RNA polymerase) operably linked to the nucleic acid sequence encoding the mRNA.
- the template plasmid also included a Cer sequence to increase plasmid stability and an antibiotic resistance gene (providing resistance to kanamycin) for the selection of transformants and plasmid maintenance.
- the template plasmid without a poly tail insert is identified as “m8” in subsequent examples.
- the MCS was used for the insertion of four different nucleic acid sequences encoding a poly(A) tail.
- the poly(A) tail sequence encoded by each nucleic acid sequence is shown in Table 5.
- Escherichia coli (E. coll) cells were transformed with the resulting template plasmids (98A, G, 0.1 and M.2). Successful transformants were selected on kanamycin-containing solid medium.
- the template plasmid was isolated and sequenced. Transformants containing a plasmid with the desired poly(A) tail sequences were expanded in liquid medium to prepare a glycerol stock.
- Plasmid stability experiment A schematic representation of the plasmid stability experiment is shown in Figure 1. The results are summarized in Table 6. Table 6: Plasmid stability
- plasmid stability was higher for the tested poly(A) tail sequences that comprised three or more segments of 10-60 adenosines separated by at least two spacers of two nucleotides others than adenosine.
- the poly(A) tail was maintained intact in 100% of the tested colonies.
- each spacer consisted of a single guanosine (plasmid G) had higher stability than the plasmid with the non-segmented poly(A) tail (plasmid 98A) which was included as a control. 72% of the tested colonies contained an intact version of plasmid G, whereas only 46% of the tested colonies contained an intact version of plasmid 98A.
- This example illustrates that the templates encoding a poly(A) tail sequence comprising at least three segments of 10-60 adenosines separated by at least two spacers of two nucleotides others than adenosine have very high stability independent of the incubation temperature of the bacterial cells during propagation.
- Example 1 of WO 2021/168052 In vitro transcribed mRNA was prepared as described in Example 1 of WO 2021/168052, which is incorporated herein by reference. Briefly, the template plasmids isolated in Example 1 were linearized by digestion with a restriction enzyme. For each gram of mRNA transcribed, a reaction containing a linearized template plasmid, an RNA polymerase (SP6), RNase inhibitor, pyrophosphatase, NTPs, DTT, and a buffering reagent was prepared with RNase-free water.
- the reaction buffer (pH 7.5) comprised 29.6 mM Tris- HC1, 3.8 mM MgCh, 8.9 mM KC1.
- a control plasmid m8 lacking a nucleic acid sequence encoding a poly(A) tail and plasmids 98A, G, O.l and M.2 as described in Example 1 were linearized by restriction enzyme digestion and used as templates for IVT reactions as described in Example 2. For each template, IVT reactions were performed in triplicate. The mRNA yield was assessed by UV spectrometry. The length of the mRNA transcripts was assessed by capillary gel electrophoresis.
- plasmid M.2 contains a single thymidine as spacer after the third segment of 25 adenosines.
- the stability of plasmid M.2 was comparable to the stability of plasmid O.l .
- An alternative plasmid construct was created in which the single thymidine was replaced with adenosine to generate plasmid M.3 with the following poly(A) tail sequence: 40A-GG-39A-GG-38A-G-3A (SEQ ID NO: 5)
- plasmid M.3 was found to have the same stability profile as plasmid M.2.
- mRNA yield of about 150 pg or greater were achieved with plasmids M.3 and O.l. As a comparison, the mRNA yield was about 200 pg with a template plasmid including a standard poly(A) tail (98 A) or no poly tail (m8). For plasmid M.2, the mRNA yield was 100 pg.
- tails of two representative IVT reactions using plasmids 98A and 0.1 as templates were analyzed).
- poly(A) tails of mRNA transcripts obtained from plasmid 98A, encoding a standard non-segmented poly(A) tail were longer than expected and had a bimodal distribution. In contrast, only a single distribution of poly(A) tail length was observed in the population of mRNA transcripts obtained from plasmid 0.1 and the vast majority of poly(A) tails had a length within 5 nucleotides of the expected tail length of 100 nucleotides.
- a template encoding a segmented poly(A) tail can advantageously yield mRNA transcripts with poly(A) tails having a single length distribution around the desired value.
- mRNA transcripts obtained from IVT reactions using plasmids m8, 98A, G, 0.1, M.2 and M.3 were used for the transfection of cultured cells.
- the mRNA transcripts obtained from plasmid m8 were enzymatically tailed with a 120 nucleotide-long poly(A) tail prior to transfection. Expression was assessed both by Western blot and flow cytometry using an antibody specific for the influenza virus A HA-H3 protein. Western blot analysis on cell lysates obtained 24 hours after transfection confirmed that all transfected cells expressed the HA-H3 protein.
- HA surface expression was determined on intact cells 24 and 48 hours after transfection. At both time points, cells transfected with mRNA transcripts comprising a segmented poly(A) tail (G, 0.1, M.2 and M.3) expressed the HA protein on their surface at levels comparable to that observed on cells transfected with mRNA transcripts comprising a standard non-segmented tail (98A). Similarly, at both time points, HA expression on cells transfected with mRNA transcripts obtained from plasmid G, 0.1, M.2 and M.3 was noninferior to that on cells transfected with mRNA transcripts comprising a standard nonsegmented that was added enzymatically (m8).
- This example demonstrates that a template encoding a segmented poly(A) tail as described herein has increased plasmid stability compared to a template encoding a nonsegmented poly(A) tail as well as a template encoding a differently configured segmented poly(A) tail that forms part of the state of the art.
- Plasmid stability was determined as described in Example 1. A 50 pl aliquot was taken of culture P2 (see Figure 1) at the end of the incubation period and plated. The incubation temperature was 30°C. A template encoding a non-segmented poly(A) tail was used as a control (112A).
- templates comprising the segmented poly(A) tail sequences A18GG, A18CC, 0.1, O-C, A40GG, S-GG, and S-CC were stable, having poly(A) tail sequences of the expected length (see Figure 2).
- Control templates comprising BioNTech’ s segmented poly(A) tails (BioNTech 40 and BioNTech 30) were likewise stable.
- This example illustrates that templates encoding a poly(A) tail sequence comprising at least three segments of 10-60 adenosines separated by at least two spacers of two nucleotides other than adenosine are stable during propagation in bacterial cells. Stability was improved in comparison to a template comprising non-segmented poly(A) tail and a template encoding a differently configured segmented poly(A) tails.
- Example 7 mRNA with segmented poly (A) tail yields improved translation efficiency
- mRNA transcripts were prepared from templates by IVT as described in Example 2. Templates comprised a nucleic acid sequence encoding the eGFP degron and a segmented test poly(A) tail sequence (S-GG, S-CC, A18GG, M-C or A40GG; see Table 8). In addition, templates comprising a non-segmented control poly(A) tail sequence (112A, 100A (SEQ ID NO: 48), 80A (SEQ ID NO: 49)) were used to prepare control mRNA. To benchmark the performance of the test mRNA transcripts, additional mRNA transcripts were prepared from templates comprising the Trepotech and BioNTech segmented poly(A) tail sequences shown in Table 8. Similar mRNA yields were achieved from each of these templates as found in Example 3.
- a transfection mix was prepared by diluting Lipofectamine MessengerMAX in OptiMEM followed by the addition of 5 pl of 100 ng/pl mRNA.
- Kinetic monitoring of protein expression was performed using the Operetta CLS Imaging system and Harmony software. Data points were collected at 60-minute intervals for 25 hours and then at 120- minute intervals for 26 hours. Data acquisition was performed on 16 fields/well using the following parameters: eGFP (Time 200 ms; Power 100%; Height -9 pm; Excitation 460-490 nm; Emission 500-550 nm) and Hoechst (Time 200 ms; Power 100%; Height -9 pm; Excitation 355-385 nm; Emission 430-500 nm).
- the eGFP degron expression data were used to calculate translation efficiency and intracellular stability values to assess the overall effect of the poly(A) tail sequence.
- the translation efficiency value was calculated by dividing the MFI maximum of a given mRNA transcript sample by the MFI maximum of the mRNA transcript reference sample comprising the non-segmented 112A tail sequence.
- the intracellular stability index value was calculated by subtracting the hours post transfection for the MFI maximum of the mRNA transcript reference sample from the hours post transfection for the MFI maximum of a given mRNA transcript sample. The values were then plotted as shown in Figure 3B.
- mRNA transcripts comprising a segmented poly(A) tail sequence comprising at least three segments of 10-60 adenosines separated by at least two spacers of two nucleotides other than adenosine have low reactogenicity.
- Exogenous single-stranded and double-stranded RNA can induce an inflammatory response via Toll-like receptors (TLRs) 7/8 and 3, respectively.
- TLR signaling can trigger the expression of NF -KP/IRF3 -induced genes.
- THPl-DualTM cells Invivogen were used to determine in vitro reactogenicity of mRNA transcripts comprising segmented poly(A) tail sequences.
- a reporter gene encoding secreted alkaline phosphatase (SEAP) is operably linked to NF-KP inducible promoter, and a reporter gene encoding secreted luciferase (Lucia) is operably linked to an Interferon-sensitive response element (ISRE).
- SEAP secreted alkaline phosphatase
- Lucia reporter gene encoding secreted luciferase
- ISRE Interferon-sensitive response element
- Non-differentiated THPl-DualTM cells were electroporated with HAH3-encoding mRNA prepared as described in Example 2.
- the HAH3-encoding mRNA comprised poly(A) tail sequences A18GG, A18CC, O-C, M-C and S-CC, as described in Table 8.
- H3HA- encoding mRNA comprising a segmented poly(A) tail comprising single-base spacers as shown in Table 9 below were also assessed for reactogenicity.
- a non-segmented control (112A) was also included as a reference.
- Negative controls included a cell culture supernatant sample of untreated cells (depicted as “None”) and a cell culture supernatant sample of mock- treated cells (no mRNA; depicted as “None + EP”) .
- Plasmids encoding the poly(A) tail sequences listed in Table 8 were used as templates for IVT reactions as described in Example 2. Stability of the mRNA transcripts was evaluated at -80°C, 4°C and 20°C over a period of 14 days and at 37°C over a period of 8 days. mRNA integrity was determined using capillary electrophoresis profiling. No statistically significant difference in mRNA stability was observed at these temperatures with all tested mRNA transcripts.
- This example illustrates that the presence of a segmented poly(A) tail sequence comprising at least three segments of 10-60 adenosines separated by at least two spacers of two nucleotides other than adenosine does not negatively affect mRNA thermostability.
Landscapes
- Health & Medical Sciences (AREA)
- Genetics & Genomics (AREA)
- Life Sciences & Earth Sciences (AREA)
- Engineering & Computer Science (AREA)
- Organic Chemistry (AREA)
- Chemical & Material Sciences (AREA)
- Zoology (AREA)
- Wood Science & Technology (AREA)
- Bioinformatics & Cheminformatics (AREA)
- Biotechnology (AREA)
- General Engineering & Computer Science (AREA)
- Biomedical Technology (AREA)
- Molecular Biology (AREA)
- Microbiology (AREA)
- Biochemistry (AREA)
- General Health & Medical Sciences (AREA)
- Plant Pathology (AREA)
- Biophysics (AREA)
- Physics & Mathematics (AREA)
- Chemical Kinetics & Catalysis (AREA)
- General Chemical & Material Sciences (AREA)
- Pharmaceuticals Containing Other Organic And Inorganic Compounds (AREA)
Abstract
The present disclosure relates to a template for in vitro transcription (IVT) of messenger RNA (mRNA), wherein the template comprises in 5' to 3' direction a first nucleic acid sequence encoding the peptide, polypeptide or protein and a second nucleic acid sequence encoding a poly(A) tail comprising three or more segments of 10-60 adenosines, wherein at least two of said segments consist of 40 adenosines or less, wherein each segment is separated from the next segment by a spacer, wherein at least two of the spacers independently consists of two nucleotides other than adenosine. The invention also relates to methods of producing mRNA that employ a template in accordance with the invention, as well as mRNA molecules obtainable by such methods and pharmaceutical compositions comprising them.
Description
STABLE POLY(A)-ENCODING MESSENGER RNA TEMPLATES
CROSS-REFERENCE TO RELATED APPLICATIONS
[1] This application claims priority to European Application No. 23306234.8 filed 18 July 2023, the contents of which is incorporated herein by reference in its entirety.
SEQUENCE LISTING
[2] The present specification makes reference to a Sequence Listing, submitted electronically as an .xml file name “2024-07- 16_PAT23027-WO-PCT_Sequence_Li sting” on 18 July 2024. The .xml file was generated on 16 July 2024 and is 50 KB in size. The entire contents of the sequence listing are herein incorporated by reference.
FIELD OF THE INVENTION
[3] The present invention relates to templates comprising a nucleic acid sequence encoding a segmented poly(A) tail for the in vitro transcription (IVT) of messenger RNA (mRNA). The templates of the invention are stably maintained in bacterial cells during propagation and are non-inferior to templates comprising a nucleic acid sequence encoding a nucleic acid sequence without poly(A) tail or a standard non-segmented poly(A) tail when used for IVT.
BACKGROUND OF THE INVENTION
[4] Messenger RNA (mRNA) is becoming increasingly important as a therapeutic agent. mRNA therapy can be used to restore normal levels of an endogenous protein or provide an exogenous therapeutic protein (e.g., a vaccine antigen or antibody) without permanently altering the genome sequence or entering the nucleus of the cell. mRNA therapy takes advantage of the cell’s own protein production and processing machinery to express a therapeutic peptide, polypeptide, or protein, is flexible to tailored dosing and formulation, and is broadly applicable to any disease or condition that is treatable through the provision of an exogenous protein.
[5] The process of manufacturing mRNA for use in therapy typically involves the in vitro transcription (IVT) of mRNA from a DNA template using a phage-derived DNA-dependent RNA polymerase. The DNA template is typically prepared by standard molecular biology techniques. Specifically, a nucleic acid sequence encoding a peptide, polypeptide or protein
of interest is cloned into a multi-copy plasmid which is then propagated in Escherichia coli (E. colt) and purified for use as a template in an IVT reaction.
[6] With the exception of replication-dependent histone transcripts, naturally occurring mRNAs include a poly(A) tail at their 3’ end. The poly(A) tail commonly consists of a homopolymeric segment of adenosines. In mammals, poly(A) tails have been reported to consist of approximately 250 nucleotides. In vivo, the poly(A) tail is typically enzymatically added to mRNAs. This process is known as polyadenylation and occurs during mRNA maturation. It involves cleavage at the 3’ end followed by the enzymatic addition of adenine bases by poly(A) polymerase. The poly(A) tail facilitates the transport of mRNA from the nucleus to the cytoplasm. It is considered important for stability of the mRNA in vivo and the efficient translation of the mRNA-encoded protein (Sachs, Curr Opin Cell Biol. 1990; 2(6): 1092-1098).
[7] It is often convenient for the DNA template to encode a poly(A) tail. However, the presence of a homopolymeric sequence encoding the poly(A) tail in a DNA template can be problematic for its propagation in bacterial cells. Homopolymeric sequences have a tendency to recombine during plasmid replication (see, e.g., Preiss et al, RNA 1998; 4(11): 1321- 1331). Especially at large scale, the plasmid encoded homopolymeric poly(A) tail tends to recombine and lose adenosines. This can be detrimental for the stability of the mRNA produced from the template. This leads to variability between the mRNAs produced, which may result in differences in their action and may require re-cloning of the template, which is costly.
[8] Previous attempts have been made to address this problem and various solutions have been proposed. These include the insertion of spacer nucleotides other than adenosine between poly(A) segments in the nucleic acid sequence that encodes the 3’ tail of the mRNA. For example, WO 2016/005324 and WO 2016/091391 demonstrate that segmentation of a poly(A) tail with a spacer comprising nucleotides other than adenosine can increase plasmid stability in E. coli by avoiding recombination in the polyA tail region. However, the inclusion of longer spacers (10 nucleotides or more) in the 3’ poly(A) tail can negatively affect the translation efficiency of the mRNA. While templates encoding a poly(A) tail with shorter spacers (6 nucleotides) do not appear to suffer from the same disadvantages concerning translation efficiency, they are often less efficient in preventing recombination between the homopolymeric segment of the 3’ tail. Shortening of the spacer to a single nucleotide seems
to reduce the occurrence of recombination events only in some instances (cf. WO 2020/074642 and Trepotec et al., RNA 2019; 25(4):507-518).
[9] Accordingly, a need exists for templates comprising segmented poly(A) tails that have high stability during propagation in bacterial cells (e.g., with a stability of the segmented poly(A) tail above 90% after two passages in E. coli (assessed by Sanger sequencing) while also maintaining translation efficiency of the resulting mRNA in target cells in vivo.
SUMMARY OF THE INVENTION
[10] The present invention is based on the discovery that high stability of templates for the in vitro transcription (IVT) of mRNA can be achieved during propagation in bacterial cells when the nucleic acid sequence encoding the poly(A) tail comprises three or more segments of 10-60 adenosines, wherein at least two of said segments consist of 40 adenosines or less and each segment is separated from the next segment by a spacer, wherein at least two of the spacers consists of two nucleotides other than adenosine. The inventors found that, when using such templates, 100% stability of a plasmid comprising the template could be achieved during propagation of the template in bacterial cells such as E. coli. The inventors also found that the mRNA resulting from such templates are non-inferior in terms of translation efficiency in comparison to mRNAs comprising standard non-segmented poly(A) tails. Surprisingly, the inventors also observed that the segmented poly(A) tails of the mRNAs resulting from IVT of templates of the invention were consistently of the desired length, whereas the length of non-segmented poly(A) tails of correspondingly produced mRNAs varied significantly. Thus, the templates of the invention can usefully be employed in the large-scale manufacture of mRNA for therapeutic purposes to provide batches of mRNAs with segmented poly(A) tails of a consistent length.
[11] Accordingly, the invention relates to a template A.g, a DNA template such as a plasmid) for in vitro transcription of messenger RNA (mRNA), wherein the template comprises in the 5’ to 3’ direction a first nucleic acid sequence encoding a peptide, polypeptide or protein and a second nucleic acid sequence encoding a poly(A) tail comprising three or more segments of 10-60 adenosines, wherein at least two of said segments consist of 40 adenosines or less, wherein each segment is separated from the next segment by a spacer, and wherein at least two of the spacers independently consists of two nucleotides other than adenosine.
[12] In some embodiments, further spacers independently consist of one or two nucleotides other than adenosine.
[13] In some embodiments, the poly(A) tail comprises three segments.
[14] In some embodiments, the poly(A) tail further comprises a fourth segment of less than 10 adenosines. In some embodiments, the fourth segment is at the 3’ end of the poly(A) tail. In some embodiments, the fourth segment consists of less than 5 adenosines (e.g., 3 adenosines).
[15] In some embodiments, each of the three or more segments consists of 25- 40 adenosines. In other embodiments, the at least two of the three or more segments consist of 20 adenosines or less (e.g., 18 or 19 adenosines). In one embodiment, the template comprises one segment of 50-60 adenosines. In another embodiment, the template comprises at least 5 segments (e.g., 6 segments) of 20 adenosines or less. In some embodiments, one or more of the segments comprise(s) different numbers of adenosines. In some embodiments, one or more of the segments comprise(s) the same number of adenosines. In some embodiments, each of the segments comprises a different number of adenosines. In some embodiments, each of the segments comprises the same number of adenosines. In some embodiments, the number of adenosines per segment decreases in a 5’ to 3’ direction.
[16] In some embodiments, the non-adenosine nucleotides of each spacer independently are guanosine or cytidine. In one embodiment, the non-adenosine nucleotides of each spacer are guanosine. In another embodiment, the non-adenosine nucleotides of each spacer are cytidine. In some embodiments, the at least two of the spacers independently consist of two guanosines or two cytidines. In one embodiment, the at least two spacers consist of two guanosines. In another embodiment, the at least two spacers consist of two cytidines. In some embodiments, at least one of the further spacers consists of one guanosine or one cytidine.
[17] In some embodiments, the poly(A) tail sequence comprises or consists of the nucleic acid sequence 34A-GG-31A-GG-27A-G-3A (SEQ ID NO: 3). In some embodiments, the poly(A) tail sequence comprises or consists of the nucleic acid sequence 40A-GG-39A-GG- 25A-T-12A-G-3A (SEQ ID NO: 4). In some embodiments, the poly(A) tail sequence comprises or consists of the nucleic acid sequence 40A-GG-39A-GG-38A-G-3A (SEQ ID NO: 5). In particular embodiments, the poly(A) tail sequence comprises or consists of the nucleic acid sequence 34A-GG-31A-GG-27A-G-3A (SEQ ID NO: 3).
[18] In some embodiments, the poly(A) tail is composed of 150 nucleotides or less. In particular embodiments, the poly(A) tail is composed of 80-150 nucleotides, e.g., 100-125 nucleotides.
[19] In some embodiments, the template is capable of being stably propagated in a bacterial cell such as E. coli at 30°C and/or 34°C. In some embodiments, a segmented poly(A) tail in accordance with the invention has a stability of 90% or greater, e.g., 92% or greater, 94% or greater, 95% or greater, 96% or greater, 97% or greater, 98% or greater, or 99% or greater during propagation in a bacterial cell such as E. coli. In some embodiments, the template is capable of being propagated in E. coli cells at 30°C and/or 34°C without any loss of the poly(A) tail. In some embodiments, the template is capable of being propagated in E. coli cells at 34°C with less than 30% loss, e.g., less than 10% loss, less than 5% loss, or less than 1% loss. In some embodiments, the template is capable of being propagated in E. coli cells at 30°C or 34°C without loss.
[20] In some embodiments, the template is a ColEl plasmid. In some embodiments, the ColEl plasmid comprises a Cer sequence.
[21] In some embodiments, the first nucleic acid sequence comprises a 5’ untranslated sequence (5’ UTR) and/or a 3’ untranslated sequence (3’ UTR).
[22] In some embodiments, the template comprises a polymerase promoter operably linked to the first and second nucleic acid sequences. In some embodiments, the promoter is an SP6, T7 or KP34 phage polymerase promoter.
[23] The invention also relates to a method of producing mRNA comprising incubating a template in accordance with the present invention in the presence of an RNA polymerase under conditions suitable for in vitro transcription (IVT).
[24] In some embodiments, the invention provides a method of making an mRNA molecule, the method comprising (a) providing a template in accordance with the present invention, and (b) contacting the template with an RNA polymerase and a reaction mixture under conditions suitable for in vitro transcription of the first and second nucleic acid sequences. In some embodiments, the method further comprises adding a 5’ cap to the resulting mRNA molecule. In some embodiments, the method comprises a 5 ’cap analogue in the IVT reaction mixture.
[25] The invention also relates to a method of improving stability of a homopolymeric sequence comprising at least 80 adenosines during propagation in a bacterial cells (e.g., E. coli), wherein the method comprises modifying the homopolymeric sequence to comprise three or more segments of 10-60 adenosines, wherein at least two of said segments consist of 40 adenosines or less, wherein each segment is separated from the next segment by a spacer, wherein at least two of the spacers independently consists of two nucleotides other than adenosine. In some embodiments, the homopolymeric sequence is comprised in a vector suitable for propagation in a bacterial cell. In some embodiments, the method further comprises transforming a bacterial cell with the vector comprising the modified homopolymeric sequence.
[26] The invention further relates to a population of mRNAs obtainable by a method of making or producing an mRNA molecule described herein. In some embodiments, at least 80% of the mRNAs in a population of mRNAs obtained in accordance with the present invention comprise a poly(A) tail of the desired length.
[27] The invention also relates to an mRNA molecule with a 3’ poly(A) tail comprising at three or more segments of 10-60 adenosines, wherein at least two of said segments consist of 40 adenosines or less, wherein each segment is separated from the next segment by a spacer, and wherein at least two of the spacers independently consists of two nucleotides other than adenosine.
[28] In some embodiments, further spacers independently consist of one or two nucleotides other than adenosine. In some embodiments, the poly(A) tail comprises three segments.
[29] In some embodiments, the poly(A) tail further comprises a fourth segment of less than 10 adenosines. In some embodiments, the fourth segment is at the 3’ end of the poly(A) tail. In some embodiments, the fourth segment consists of less than 5 adenosines (e.g., 3 adenosines).
[30] In one embodiment, the mRNA comprises three segments, each consisting of 25-40 adenosines. In another embodiment, at least two of the three or more segments consist of 20 adenosines or less (e.g., 18 or 19 adenosines). In one embodiment, the poly(A) tail comprises at least one segment of 50-60 adenosines. In another embodiment, the poly(A) tail comprises at least 5 segments (e.g., 6 segments) of 20 adenosines or less.
[31] In some embodiments, one or more of the segments comprise(s) different numbers of adenosines. In some embodiments, one or more of the segments comprise(s) the same number of adenosines. In some embodiments, each of the segments comprises a different number of adenosines. In some embodiments, each of the segments comprises the same number of adenosines. In some embodiments, the number of adenosines per segment decreases in a 5’ to 3’ direction.
[32] In some embodiments, the non-adenosine nucleotides of each spacer in an mRNA molecule of the invention independently are guanosine or cytidine. In one embodiment, the non-adenosine nucleotides of each spacer are guanosine. In another embodiment, the non- adenosine nucleotides of each spacer are cytidine. In some embodiments, the at least two spacers independently consist of two guanosines or two cytidines. In one embodiment, the at least two spacers consist of two guanosines. In another embodiment, the at least two spacers consist of two cytidines. In some embodiments, at least one of the further spacers consists of one guanosine or one cytidine.
[33] In some embodiments, the poly(A) tail in an mRNA molecule of the invention is composed of 150 nucleotides or less. In particular embodiments, the poly(A) tail is composed of 80-150 nucleotides, e.g., 100-125 nucleotides.
[34] The invention also relates to a pharmaceutical composition comprising an mRNA molecule in accordance with the invention and a pharmaceutically acceptable carrier. In some embodiments, the mRNA molecule comprised in the pharmaceutical composition is encapsulated in a lipid nanoparticle. In some embodiments, the composition is an immunogenic composition (e.g., a vaccine). In some embodiments, the immunogenic composition is used for immunizing a subject. In some embodiments, the immunogenic composition prevents a virus infection in the subject, decreases the subject’s likelihood of contracting a virus infection, or reduces the subject’s likelihood of getting seriously ill from a virus infection. In some embodiments, the immunogenic composition raises a protective immune response in the subject. In some embodiments, the subject is a human. In some embodiments, the human is 6 months of age or older, less than 18 years of age, at least 6 months of age and less than 18 years of age, at least 18 years of age and less than 65 years of age, at least 6 months of age and less than 5 years of age, at least 5 years of age and less than 65 years of age, at least 60 years of age, or at least 65 years of age. In some embodiments,
immunization is performed intramuscularly, intradermally, subcutaneously, intravenously, intranasally, by inhalation, or intraperitoneally.
[35] Other features, objects, and advantages of the present invention are apparent in the detailed description, drawings and embodiments that follow. It should be understood, however, that the detailed description, the drawings, and the embodiments, while indicating embodiments of the present invention, are given by way of illustration only, not limitation. Various changes and modifications will become apparent to those skilled in the art.
BRIEF DESCRIPTION OF THE DRAWINGS
[36] Embodiments of the invention will be described, by way of example, with reference to the following drawing.
[37] Figure 1 schematically outlines an assay that can be used to determine plasmid stability in a bacterial cell such as Escherichia coli (E. coif). A glycerol stock of a bacterial clone comprising a plasmid of interest is thawed and used to inoculate a preculture. The preculture is incubated for 8 hours at 30°C. This preculture is then used to inoculate a liquid culture Pl which comprises a growth medium and an antibiotic to maintain the plasmid in the bacterial cell (e.g., 25 mL Luria-Bertani [LB] broth + kanamycin [Kana] at 25 pg/ml as indicated in the figure). After 18 to 20 hours of incubation, an aliquot of this culture (Pl) is taken to inoculate liquid culture P2, which has the same composition as culture Pl. The optical density of the inoculum at 600 nm can be adjusted to 0.005 to 0.02, as indicated in the figure. Culture P2 is incubated for 18 to 20 hours under the same conditions as culture Pl. A suitable incubation temperature is 30°C or 34°C. At the end of the respective 18-20- hour incubation periods, a 50 pL aliquot is taken from each of culture Pl and culture P2 and used for an 8-fold serial dilution series. 100 pL of the last dilution is plated on a suitable solid culture medium (e.g., LB medium + kanamycin at 25 pg/mL). After a suitable incubation period (e.g., 16-24 hours), 16 to 24 colonies are isolated from each plate and cultured in 1.2 mL of liquid culture medium (e.g., LB broth supplemented with kanamycin at 25 pg/mL). After a suitable period of incubation (e.g., overnight), plasmid DNA is purified from the resulting 16 to 24 cultures, e.g., by miniprep plasmid purification. The purified plasmid DNA is then analyzed, e.g., by Sanger sequencing, to determine the presence of the intact nucleic acid sequence of the plasmid or a particular portion of interest (e.g., the nucleic acid sequence encoding the poly (A) tail).
[38] Figure 2 shows the results of a plasmid stability assay performed as illustrated in Figure 1. The stability of plasmid templates encoding a segmented poly(A) tail or nonsegmented poly(A) tail is compared. Each filled black circle depicts the length of the poly(A) tail measured by sequencing of a template plasmid. Each plasmid was obtained from a colony isolated from a P2 culture as shown in Figure 1.
[39] Figure 3 illustrates the impact of segmented and non-segmented poly (A) tails on intracellular stability and translation efficiency of mRNA transcripts encoding a eGFP degron. Figure 3A shows the mean fluorescence intensity (MFI) per cell for eGFP degron expression at different time-points post-transfection of various mRNA transcripts as indicated. The MFI values were corrected for MFI background. Figure 3B shows the translation efficiency and intracellular stability of the tested mRNA transcripts calculated based on eGFP expression data. As can be seen from this figure, most of the values clustered in the lower left corner of the plot. Only mRNA transcripts comprising the segmented 0.1 poly(A) tail sequence stood out by having both exceptional intracellular stability and very high translation efficiency.
[40] Figure 4 illustrates the reactogenicity of mRNAs comprising a segmented or nonsegmented poly(A) tail as determined in THPl-Dual™ cells. These cells comprise an NF-KP inducible reporter gene encoding secreted alkaline phosphatase (SEAP) and an interferonsensitive response element (ISRE)-inducible reporter gene encoding Lucia luciferase. Figure 4A shows the relative luminescence intensity of Lucia luciferase within supernatants obtained from cells electroporated with HAH3 mRNA transcripts comprising segmented or non-segmented poly(A) tails as indicated. Figure 4B shows the optical density of secreted alkaline phosphatase (SEAP) within supernatants obtained from cells electroporated with HAH3 mRNA transcripts comprising segmented or non-segmented poly (A) tails as indicated. Optical density was measured at 620 nm. Bars in Figures 4A and 4B represent mean value ± SEM. The supernatant from cells electroporated with mRNA transcripts encoding eGFP was included as a positive control. Reactogenicity of the tested mRNA transcripts was minimal. The presence of segmented poly(A) tails on mRNA transcripts did not increase reactogenicity relative to non-segmented poly(A) tails.
DEFINITIONS
[41] In order for the present invention to be more readily understood, certain terms are first defined below. Additional definitions for the following terms and other terms are set forth throughout the specification.
[42] As used in this specification and the appended claims, the singular forms “a”, “an”, and “the” include plural referents unless the context clearly dictates otherwise. For example, “a ribonucleotide” is understood to represent one or more ribonucleotides. As such, the terms “a” (or “an”), “one or more”, and “at least one” can be used interchangeably herein.
[43] Unless specifically stated or obvious from context, as used herein, the term “or” is understood to be inclusive and covers both “or” and “and”. Furthermore, “and/or” where used herein is to be taken as specific disclosure of each of the two specified features or components with or without the other. Thus, the term “and/or” as used in a phrase such as “A and/or B” herein is intended to include “A and B”, “A or B”, “A” (alone), and “B” (alone). Likewise, the term "and/or" as used in a phrase such as “A, B, and/or C” is intended to encompass each of the following aspects: A, B, and C; A, B, or C; A or C; A or B; B or C; A and C; A and B; B and C; A (alone); B (alone); and C (alone).
[44] Throughout this specification and embodiments, the words “have” and “comprise”, or variations such as “has”, “having”, “comprises”, or “comprising” will be understood to imply the inclusion of a stated integer or group of integers but not the exclusion of any other integer or group of integers. It is further understood that wherever embodiments are described herein with the language “comprising” or “having” of grammatical equivalents thereof, otherwise analogous embodiments described in terms of “consisting of’ and/or “consisting essentially of’ are also provided.
[45] As used herein, the term “about” refers to an interval of accuracy that a person skilled in the art will understand to still ensure the technical effect of the feature in question. The term indicates a deviation from the indicated numerical value of ±10%. In some embodiments, the deviation is ±5% of the indicated numerical value. In certain embodiments, the deviation is ±1% of the indicated numerical value.
[46] As used herein, the term “mRNA” refers to a polyribonucleotide that encodes at least one peptide, polypeptide, or protein. mRNA as used herein encompasses both modified and unmodified RNA. mRNA may contain one or more coding and non-coding regions (e.g., a
5’ untranslated region and a 3’ untranslated region). mRNA can be purified from natural sources, produced using recombinant expression systems and optionally purified, in vitro transcribed, or chemically synthesized. The invention particularly relates to the manufacturing of mRNA by in vitro transcription (IVT). Where appropriate, e.g., in the case of chemically synthesized molecules, mRNA can comprise nucleotide analogues such as analogues having chemically modified bases or sugars, backbone modifications, etc. An mRNA sequence is presented in the 5’ to 3’ direction unless otherwise indicated. A typical mRNA comprises a 5’ cap, a 5’ untranslated region (5’ UTR), a protein-coding region, and a 3’ untranslated region (3’ UTR). A poly(A) tail is typically added to or included in in vitro transcribed mRNAs to mimic naturally occurring mRNAs. The presence of a poly(A) tail at the 3’ end of an mRNA molecule prevents degradation inside cells, thereby enhancing its half-life in vivo.
[47] The terms “peptide”, “polypeptide” or “protein” are used interchangeably herein. For example, the expression “a nucleic acid sequence encoding a polypeptide” refers to any nucleic acid sequence that encodes an amino acid sequence which, upon expression, forms a peptide, polypeptide, or protein. Such a nucleic acid sequence may further comprise a 5’- untranslated region (5’ UTR) and 3 ’-untranslated region (3 ’-UTR).
[48] As used herein, the term “poly(A)” refers to a nucleic acid comprising one or more homopolymeric segments of adenosines typically located at the 3’ end of an mRNA molecule. The homopolymeric segments of adenosines may be separated from each other by spacers comprising one or more nucleotides other than adenosine. Poly(A) tails comprising one or more spacers are referred to herein as “segmented poly(A) tails” to distinguish them from poly(A) tails that consist entirely of a single homopolymeric region of adenosines (referred to herein as “non-segmented poly (A) tails”).
[49] As used herein, the terms “reference template” and “reference plasmid” refers to a DNA template or plasmid that serves as a comparator to assess, e.g., stability during propagation, and differs from a template or plasmid of the invention only by a single sequence element, typically the nucleic acid sequence encoding the poly(A) tail.
[50] As used herein, the term “spacer” refers to one or more nucleotides other than adenosine within the poly(A) tail. In some embodiments, the spacer consists of either one or two consecutive cytidine(s) or guanosine(s).
[51] As used herein, the term “sequence-optimized” is used to describe a nucleotide sequence that is modified relative to a naturally occurring or wild-type nucleotide sequence. In the case of a sequence-optimized mRNA, such modifications may include, e.g., codon optimization and/or the use of 5’ UTRs and 3’ UTRs which are not normally associated with the naturally occurring or wild-type nucleic acid.
[52] As used herein, the terms “codon optimization” and “codon-optimized” refer to modifications of the codon composition of a naturally occurring or wild-type nucleic acid encoding a peptide, polypeptide or protein that do not alter its amino acid sequence, thereby improving protein expression of said nucleic acid. In the context of the present invention, “codon optimization” may also refer to the process by which one or more optimized nucleotide sequences are arrived at by removing, with filters, less than optimal nucleotide sequences from a list of nucleotide sequences, such as filtering by guanine-cytosine (GC) content, codon adaptation index (CAI), presence of destabilizing nucleic acid sequences or motifs, and/or presence of pause sites and/or terminator signals.
[53] As used herein, the term “template” refers to a molecule (typically a DNA molecule such as a plasmid) comprising a nucleic acid sequence encoding an mRNA transcript to be synthesized by in vitro transcription (IVT). The template is used as template for IVT in order to produce a mRNA transcript encoded by the template. The template comprises all elements necessary for IVT, particularly a promoter element for binding of an RNA polymerase, which is operably linked to the nucleic acid sequence encoding a desired mRNA transcript. The template may be a linear or a circular DNA molecule that is linearized before IVT.
[54] As used herein, the term “batch” refers to a quantity or amount of mRNA synthesized at one time, e.g., produced according to a single manufacturing order during the same cycle of manufacture. A batch may refer to a population of mRNAs synthesized in one reaction that occurs via a single aliquot of enzyme and/or a single aliquot of template for continuous synthesis under one set of conditions. The term “batch” does not refer to populations of mRNAs synthesized at different times and/or in different reactions that are combined to achieve a desired amount.
[55] As used herein, the term “expression” of a nucleic acid sequence refers to translation of an mRNA into a peptide, polypeptide, or protein in vitro or in vivo.
[56] As used herein, the term “therapeutic” refers to any pharmaceutical, drug, or composition that can be used to treat or prevent a disease, illness, condition, or disorder of bodily function.
[57] As used herein, the term “substantially” refers to the qualitative condition of exhibiting total or near-total extent or degree of a characteristic or property of interest. One of ordinary skill in the biological arts will understand that biological and chemical phenomena rarely, if ever, go to completion and/or proceed to completeness or achieve or avoid an absolute result. The term “substantially” is therefore used herein to capture the potential lack of completeness inherent in many biological and chemical phenomena.
[58] As used herein, the term “zzz vitro" refers to events that occur in an artificial environment, e.g., in a test tube or reaction vessel, or in a cell in cell culture, etc., rather than within a multi-cellular organism.
[59] As used herein, the term “zzz vzvo” refers to events that occur within a multi-cellular organism, such as a human or a non-human animal.
[60] As used herein, the term “isolated” refers to a substance and/or entity that has been (1) separated from at least some of the components with which it was associated when initially produced (whether in nature and/or in an experimental setting), and/or (2) produced, prepared, and/or manufactured by the hand of man.
[61] As used herein, the term “immunogenic composition” refers to a composition that generates an immune response that may or may not be a protective immune response or protective immunity. The term “immune response” refers to a response of a cell of the immune system, such as a B cell, T cell, dendritic cell, macrophage or polymorphonucleocyte, to a stimulus such as an antigen, immunogen, or vaccine. An immune response can include any cell of the body involved in a host defense response, including for example, an epithelial cell that secretes an interferon or a cytokine. An immune response includes, but is not limited to, an innate and/or adaptive immune response. Methods of measuring immune responses are well known in the art and include, for example, measuring proliferation and/or activity of lymphocytes (such as B or T cells), secretion of cytokines or chemokines, inflammation, antibody production and the like. An antibody response or humoral response is an immune response in which antibodies are produced. A “cellular immune response” is one mediated by T cells and/or other white blood cells.
[62] As used herein, the term “vaccine” refers to a composition that generates a protective immune response or a protective immunity in a subject. A “protective immune response” or “protective immunity” refers to an immune response that protects a subject from infection (prevents infection or prevents the development of disease associated with infection) or reduces the symptoms of infection (for instance, an infection by an influenza virus). Vaccines may elicit both prophylactic (preventative) and therapeutic responses.
[63] Unless otherwise defined herein, technical and scientific terms used herein have the same meaning as commonly understood by one of ordinary skill in the art to which this application belongs and as commonly used in the art to which this application belongs. Exemplary methods and materials are described below, although methods and materials similar or equivalent to those described herein can also be used in the practice or testing of the present disclosure. In case of conflict, the present specification, including definitions, will control.
[64] Generally, nomenclature used in connection with, and techniques of, cell and tissue culture, molecular biology, virology, immunology, microbiology, genetics, analytical chemistry, synthetic organic chemistry, medicinal and pharmaceutical chemistry, and protein and nucleic acid chemistry and hybridization described herein are those well-known and commonly used in the art. Enzymatic reactions and purification techniques are performed according to manufacturer’s specifications, as commonly accomplished in the art or as described herein. Further, unless otherwise required by context, singular terms shall include pluralities and plural terms shall include the singular.
[65] All publications and other reference materials referenced herein are hereby incorporated by reference in their entirety. Although a number of documents are cited herein, this citation does not constitute an admission that any of these documents forms part of the common general knowledge in the art.
DETAILED DESCRIPTION OF THE INVENTION
[66] The present invention relates to templates for in vitro transcription (IVT) of messenger RNA (mRNA). Such templates typically may comprise in 5’ to 3’ direction a first nucleic acid sequence encoding a polypeptide and a second nucleic acid encoding a poly(A) tail. mRNAs used in therapy commonly include a poly(A) tail at their 3’ end to prevent rapid degradation of the mRNAs inside cell in vivo and to improve expression of the encoded
peptide, polypeptide, or protein. It is often advantageous to use co-transcriptional tailing, e.g., because it avoids the costs associated with a separate enzymatic tailing step, and in theory reduces variability of the poly(A) tail length and hence reproducibility.
[67] The inventors have surprisingly found that templates were maintained faithfully during propagation in bacterial cells when the poly(A) tail-encoding nucleic acid sequence comprised three segments of 10-60 adenosines, of which at least two consisted of 40 adenosines or less, wherein each segment was separated from the next segment by a spacer, of which at least two independently consisted of two nucleotides other than adenosine. Moreover, the inventors found that mRNAs obtained by in vitro transcription (IVT) of such templates were translated as efficiently as mRNA with a standard non-segmented poly(A) tail. Unexpectedly, the resulting population of mRNAs consistently had poly(A) tails of the desired lengths.
[68] The inventors found that templates encoding a poly(A) tail sequence comprising the nucleic acid sequence as set forth in SEQ ID NO: 3-5 were stably maintained during propagation in bacterial cells, produced acceptable yields of mRNA transcripts with poly(A) tails having the expected length and were minimally reactogenic. Higher mRNA transcript yields were achieved with templates encoding a poly(A) tail sequence comprising the nucleic acid sequence of SEQ ID NO: 3 or SEQ ID NO: 5. Surprisingly, mRNA transcripts comprising the segmented poly(A) tail sequence of SEQ ID NO: 3 were found to have higher intracellular stability and translation efficiency.
Poly(A) tail
[69] Accordingly, the invention relates to a template for in vitro transcription (IVT) of messenger RNA (mRNA), wherein the template comprises in 5’ to 3’ direction a first nucleic acid sequence encoding a polypeptide and a second nucleic acid sequence encoding a poly(A) tail comprising three or more segments of 10-60 adenosines, wherein at least two of said segments consist of 40 adenosines or less, wherein each segment is separated from the next segment by a spacer, and at least two of the spacers independently consist of two nucleotides other than adenosine.
[70] In some embodiments, further spacers independently consist of one or two nucleotides other than adenosine. In some embodiments, the poly(A) tail comprises three segments.
[71] In some embodiments, the poly(A) tail further comprises a fourth segment of less than 10 adenosines. In some embodiments, the fourth segment is at the 3’ end of the poly(A) tail. In some embodiments, the fourth segment consists of less than 5 adenosines (e.g., 3 adenosines).
[72] In some embodiments, each of the three segments consists of 25-40 adenosines. In other embodiments, the at least two of the three segments consist of 20 adenosines or less (e.g., 18 or 19 adenosines). In one embodiment, the template comprises one segment of 50- 60 adenosines. In another embodiment, the template comprises at least 5 segments (e.g., 6 segments) of 20 adenosines or less.
[73] In some embodiments, the number of adenosines in the three segments is about the same or the same. For example, in some embodiments, the first, second and third segments in 5’ to 3’ direction each consist of about 40 adenosines (e.g., 40, 39 and 38, respectively).
[74] In other embodiments, the number of adenosines in each segment is different. In some embodiments, the number of adenosines in at least one of the three (or four) segments is different to the number of adenosines in the other two (or three) segments. For example, in some embodiments, the first segment in 5’ to 3’ direction comprises more than 30 adenosines, e.g., 40 adenosines or more (e.g., 34 to 59), whereas the second, third and optional fourth segment comprise fewer adenosines (e.g., 30 adenosines or less). In other embodiments, the first and third segments consist of 20 adenosines or less (e.g., 19, 18, 17, 16 or 15), whereas the second segment consists of 60 adenosines or less (e.g., 59, 58, 56, or 55).
Spacer
[75] At least two spacers in a segmented poly(A) tail in accordance with the invention may consist of two nucleotides other than adenosine. Further spacers may independently consist of one or two nucleotides other than adenosine.
[76] In some embodiments, the nucleotide other than adenosine is guanosine. In some embodiments, the nucleotide other than adenosine is cytidine. The use of cytidine(s) in the spacers may advantageously be employed to boost expression of the polypeptide encoded by an mRNA comprising a poly(A) tail of the invention. Typically, the same type of nucleotide (either guanosine or cytidine) is used in each spacer of the poly(A) tail. In some embodiments, some spacers consist of cytidine(s) and some spacers consists of guanosine(s).
[77] In some embodiments, further spacers in the poly(A) tail consist of a single nucleotide (e.g., a guanosine or a cytidine). In some embodiments, each spacer in the poly(A) tail consists of two nucleotides (e.g., two guanosines or two cytidines).
[78] In particular embodiments, the poly(A) tail comprises at least two spacers consisting of two nucleotides (e.g., a guanosine or a cytidine) and at least one spacer consisting of a single nucleotide (e.g., a guanosine or a cytidine). In some embodiments, the at least two spacers consist of two cytidines, and the at least one spacer of a single cytidine. In some embodiments, the at least two spacers consist of two guanosines, and the at least one spacer of one guanosine.
[79] In some embodiments, the poly(A) tail comprises at least one spacer consisting of two cytidines and at least one spacer consisting of two guanosines. In some embodiments, the poly(A) tail comprises at least one further spacer consisting of a single cytidine or a single guanosine.
[80] In some embodiments, a segmented poly(A) tail in accordance with the invention consists of four segments, wherein each of the first three segments in the 5’ to 3’ direction consist of 25-40 adenosines. In some embodiments, the fourth segment comprises less than 10 (e.g., less than 5) adenosines and is separated from the third segment by a spacer consisting of a single nucleotide other than adenosine. In some embodiments, the same type of non-adenosine nucleotide (either guanosine or cytidine) is used in each spacer.
Length
[81] Typically, a poly(A) tail comprising 60 and 150 nucleotides (e.g., 90-120 nucleotides) is sufficient to maintain mRNA stability in vivo. In some embodiments, the length of the poly(A) sequence of the template is less than 150 nucleotides e.g., less than 145 nucleotides, less than 140 nucleotides, less than 135 nucleotides, less than 130 nucleotides, less than 125 nucleotides, less than 120 nucleotides, less than 115 nucleotides, less than 110 nucleotides, less than 105 nucleotides, less than 100 nucleotides, less than 99 nucleotides, less than 98 nucleotides, less than 95 nucleotides, less than 90 nucleotides, less than 85 nucleotides or 80 nucleotides.
[82] In some embodiments, the average length of a poly(A) tail of an in vitro transcribed mRNA prepared in accordance with the invention is 80-150 nucleotides e.g., 90-125 nucleotides or 100-120 nucleotides.
Size and arrangement of segments
[83] The inventors have found that particular sizes and arrangements of at least three segments comprising of 10-60 adenosines are particularly effective in enhancing the stability of templates during propagation in bacterial cells.
[84] In some embodiments, the poly(A) tail comprises or consists of a nucleic acid of formula I, wherein A is a segment consisting of adenosines and S is a spacer consisting of cytidine(s) and/or guanosine(s):
Ani - Smi - An2 - Sm2 - An3 (formula I) wherein: nl, n2 and n3 are the number of adenosines in the respective segments, wherein each of nl, n2 and n3 is independently selected from 19 to 59, ml and m2 are the number of nucleotides in the respective spacers, wherein each of ml and m2 is 2.
[85] In some embodiments, each of nl, n2 and n3 of formula I is independently selected from 35 to 40. In some embodiments, each of nl, n2 and n3 of formula I is independently selected from 25-35. In some embodiments, nl is greater than 30, and n2 and n3 are less than 40. In some embodiment, each of nl and n3 of formula I is less than 20 (e.g., 15-19), and n2 is more than 40 (e.g., 50-60).
[86] In some embodiments, nl is 34, n2 is 31, n3 is 27, S is independently selected from guanosine and cytosine. In some embodiments, formula 1 is followed by a single nucleotide spacer (e.g., G or C) and 3 adenosines, e.g., GAAA.
[87] In some embodiments, each of the spacers consist of guanosines. In other embodiments, each of the spacers consist of cytidines.
[88] In some embodiments, the poly(A) tail comprises or consists of a nucleic acid of formula II, wherein A is a segment consisting of adenosines and S is a spacer consisting of cytidine(s) and/or guanosine(s):
Anl - Sml - An2 “ Sm2 “ An3 “ Sm3 “ An4 (formula II) wherein: nl, n2, n3 and n4 are the number of adenosines in the respective segments,
nl is 34 to 59, n2 and n3 are independently selected from 27 to 39, n4 is 3, ml, m2 and m3 are the number of nucleotides in the respective spacers, wherein each of ml and m2 is 2 and m3 is 1 or 2.
[89] In some embodiments, each of nl, n2 and n3 of formula II is independently selected from 35 to 40. In some embodiments, each of nl, n2 and n3 of formula II is independently selected from 25-35. In some embodiments, nl is greater than 30, and n2 and n3 are less than 40.
[90] In some embodiments, each of the spacers consists of guanosine. In other embodiments, each of the spacers consists of cytidine.
[91] In some embodiments, m3 of formula II is 1 and the spacer consists of a guanosine. In other embodiments, m3 is 1 and the spacer consists of a cytidine.
[92] In some embodiments, nl is 34, n2 is 31, n3 is 27, n4 is 3, S is independently selected from guanosine and cytosine, ml and m2 are both 2, and m3 is 1.
[93] In some embodiments, each of the spacers consist of guanosines. In other embodiments, each of the spacers consist of cytidines.
[94] In some embodiments, the poly(A) tail comprises or consists of a nucleic acid of formula III, wherein A is a segment consisting of adenosines and S is a spacer consisting of cytidine(s) and/or guanosine(s):
(An - Sm)r An (formula III) wherein: n is the number of adenosines in the respective segments, wherein n is independently selected from 15 to 18, m is the number of nucleotides in the respective spacers, wherein m is 2, and r is the number of repeats, wherein r is selected from 5 to 7.
[95] In some embodiments, each instance of n in formula III is 15. In some embodiments, each instance of n in formula III is 18.
[96] In some embodiments, r is 6. In some embodiments r is 7.
[97] In some embodiments, each spacer consists of guanosines. In other embodiments, each spacer consists of cytidines.
Exemplary poly(A) tail sequences
[98] Exemplary poly(A) tail sequences of the present invention are shown in Table 1.
[99] In some embodiments, the poly(A) tail sequence comprises or consists of the nucleic acid sequence 34A-GG-31A-GG-27A-G-3A (SEQ ID NO: 3). In some embodiments, the poly(A) tail sequence comprises or consists of the nucleic acid sequence 40A-GG-39A-GG- 25A-T-12A-G-3A (SEQ ID NO: 4). In some embodiments, the poly(A) tail sequence comprises or consists of the nucleic acid sequence 40A-GG-39A-GG-38A-G-3A (SEQ ID NO: 5).
[100] mRNA transcripts comprising a poly(A) tail with the nucleic acid sequence set forth in SEQ ID NO: 3 were found to have exceptionally high intracellular stability and translation efficiency. This combination of high intracellular stability and translation efficiency is especially useful for in vivo therapeutic uses. Accordingly, in particular embodiments, the
poly(A) tail sequence comprises or consists of the nucleic acid sequence 34A-GG-31A-GG- 27A-G-3A (SEQ ID NO: 3).
Template
[101] Templates of the invention can easily be propagated in bacterial cells to produce large quantities for use in the in vitro transcription (IVT) of messenger RNA (mRNA). In some embodiments, the template is a DNA template. In a typical embodiment, a template of the invention is a plasmid. Plasmid backbones suitable for use with the invention are well-known to the skilled person and include the backbones of medium and high copy number plasmids routinely employed in the cloning and expression of gene sequences. These include pUC and pBR series of plasmids, including pUC19 and pBR322.
[102] Typically, a plasmid for use in the invention can be propagated in a bacterial cell such as E. coli at 100 copies or more without jeopardizing long-term plasmid maintenance and stability. The plasmid may comprise additional elements (e.g., ori and/or cer) that contribute to its stability during propagation in a bacterial cell.
Origin of replication
[103] In a typical embodiment, a template in accordance with the invention such as a plasmid comprises an origin of replication (ori). The ori can control replication and copy number of the template during propagation. The ori is chosen for combability with a host species such as a bacterial cell (e.g., E. coli). In some embodiments, the ori is from pUC19, pMBl or pCoIEl.
[104] The inventors have found that a vector backbone comprising an ori having the nucleotide sequence of SEQ ID NO: 12 is particularly advantageous for use with the invention:
CCCGTAGAAAAGATCAAAAGATCTTCTTGAGATCCTTTTTTTCTGCGCGTAATC TGCTACTTGCAAACAAAAAAACCACCGCTACCAGCGGTGGTTTGTTTGCCGAAT CAAGAGCTACCAACTCTTTTTCCGAAGGTAACTGGCTTTAGCAGAGCGCAGATA CCAAATACTGTCCTTCTAGTGTAGCCGTAGTTAGGCCACCACTTCAAGAACTCT GTAGCACCGCCTACATACCTCGCTCTGCTAATCCTGTTACCAGTGGCTGCTGCC AGTGGCGATAAGTCGTGTCTTACCGGGTTGGACTCAAGACGATAGTTACCGGA TAAGGCGCAGCGGTCGGGCTGAACGGGGGGTTCGTGCACACAGCCCAGCTTGG AGCGAACGACCTACACCGAACTGAGATACCTACAGCGTGAGCTATGAGAAAGC
GCCACGCTTCCCGAAGGGAGAAAGGCGGACAGGTATCCGGTAAGCGGCAGGG TCGGAACAGGAGAGCGCACGAGGGAGCTTCCAGGGGGAAACGCCTGGTATCTT TATAGTCCTGTCGGGTTTCGCCACCTCTGACTTGAGCGTCGATTTTTGTGATGCT CGTCAGGGGGGCGGAGCCTATGGAAAAACGCCAGCAACGCGGCCTTTTTACGG TTCCTGGCCTTTTGCTGGCCTTTTGCTCACATTG (SEQ ID NO: 12).
[105] Without wishing to be bound by any particular theory, the inventors believe that this ori comprises a stabilizing mutation, rendering plasmids containing this ori especially suitable for propagation in E. coli.
ColEl resolution sequence
[106] In some embodiments, a template such as a plasmid, includes a ColEl resolution (cer) sequence, cer is a target site for multimer resolution in bacterial cells. Including a cer sequence prevents plasmid loss during propagation in bacterial cells such as E. coli.
[107] During replication, daughter plasmids are segregated by recombining across duplicated cer sequences in a plasmid dimer, yielding two monomers. The insertion of a cer sequence allows for the stable inheritance of ColEl and related plasmids. This occurs via the prevention of a “dimer catastrophe” caused by the runaway accumulation of multimers, cer is a target site for multimer resolution in bacterial cells (Peubez etal., Microb Cell Fact 2010; 9:65).
Restriction sites
[108] A template for use with the invention, e.g., a DNA template, typically comprises one or more restriction site(s). The inclusion of one or more restriction sites may be useful for a variety of reasons. For example, templates that are easily propagated in bacterial cells are typically circular, whereas in vitro transcription (IVT) is typically performed with linear templates to prevent transcriptional readthrough. Accordingly, in some embodiments, a template of the invention may comprise a restriction site to linearize the template with a suitable restriction enzyme prior to IVT.
[109] Moreover, the same template backbone is typically used for the production of various mRNAs, each of which encodes a different peptide, polypeptide or protein. Accordingly, the template backbone may include additional restriction sites to insert/replace the nucleic acid sequence encoding the peptide, polypeptide or protein. The restriction sites are typically located within the template in such a manner that the inserted nucleic acid sequence encoding
the peptide, polypeptide or protein is flanked by a 5’ untranslated region (5’ UTR) and a 3’ untranslated region (3’ UTR) to yield an mRNA coding sequence that is operably linked to an RNA polymerase promoter. Similarly, the template backbone may include one or more restriction sites to insert/replace a nucleic acid sequence encoding a poly(A) tail of the invention. The restriction site is positioned such that the nucleic acid sequence encoding the poly(A) tail is immediately 3’ to the 3’ UTR in the template.
[HO] Suitable restriction sites include those that are cut by a type II restriction enzyme, such as a type IIP or type IIS restriction enzyme. In some embodiments, the restriction enzyme is Xhol, EcoRI, BciVI, Spel, Xbal, Ndel, AflII, SacI, Kpnl, Smal, BamHI, Sall, Sbfl, PstI, BspQI (or its isoschizomers SapI), or Hindlll. In some embodiments, a restriction site for a restriction enzyme such as Clal is used to insert a segmented poly(A) of the invention into a template such as a plasmid. In some embodiments, the restriction site is transcribed into the mRNA molecule. The presence of the restriction site in the template or the mRNA molecule may be used for analytical purposes (e.g., to assess stability of the poly (A) sequence or the presence of the poly (A) tail).
Selection marker
[Hl] A template according to the invention, e.g., a DNA template such as a plasmid, may comprise a selection marker gene. A selection marker gene allows for the identification of cells that contain the template (e.g., during cloning). The template typically comprises a positive selection marker gene. A positive selection marker gene can encode a gene product (e.g., a protein or enzyme) that provides resistance to an antibiotic, enabling the selection of bacterial colonies carrying the template. Moreover, the presence of an antibiotic in the growth medium during propagation can aid in the maintenance of the template. Commonly used positive selection marker genes include kanamycin resistance (aph) and ampicillin resistance (bla) genes. In a particular embodiment, a template of the invention (e.g., a plasmid) comprises a kanamycin resistance gene.
Promoter
[112] The templates of the invention, e.g., a DNA template, typically include a suitable promoter sequence operably linked to the nucleic acid sequence encoding the peptide, polypeptide, or protein. During in vitro transcription (IVT), a DNA-dependent RNA polymerase initiates transcription by contacting the promoter sequence. DNA-dependent
RNA polymerases commonly used for IVT include, e.g., SP6 RNA polymerase, T7 RNA polymerase, T3 RNA polymerase and Klebsiella phage KP34 RNA polymerase.
[113] Accordingly, in some embodiments, the promoter sequence is an SP6 RNA polymerase promoter sequence. In other embodiments, the promoter sequence is a T7 RNA polymerase promoter sequence. In other embodiments, the promoter sequence is a T3 RNA polymerase promoter sequence. In other embodiments, the promoter sequence is a KP34 RNA polymerase promoter sequence.
[114] Any promoter sequence that can be recognized by an SP6 RNA polymerase may be used in the present invention. An SP6 promoter sequence may comprise 5' ATTTAGGTGACACTATAG-3' (SEQ ID NO: 13), or a variant thereof that is optimized for recognition and/or binding of SP6 RNA polymerase. Such variants may have the nucleic acid sequence 5'-ATTTAGGTGACACTATAGAAGNG-3' (SEQ ID NO: 14), where N is A, C, T or G. Exemplary variants are shown in Table 2.
[115] In some embodiments, the SP6 promoter sequence is upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 34A-GG-31A-GG-27A-G- 3A (SEQ ID NO: 3). In some embodiments, the SP6 promoter sequence is upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 40A-GG-39A- GG-25A-T-12A-G-3A (SEQ ID NO: 4). In some embodiments, the SP6 promoter sequence is upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 40A-GG-39A-GG-38A-G-3 A (SEQ ID NO: 5). In particular embodiments, the SP6 promoter
sequence is upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 34A-GG-31A-GG-27A-G-3A (SEQ ID NO: 3).
[116] In some embodiments, the SP6 promoter comprises a sequence selected from the group consisting of SEQ ID NOs: 15-23 and is upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 34A-GG-31A-GG-27A-G-3A (SEQ ID NO: 3). In some embodiments, the SP6 promoter comprises a sequence selected from the group consisting of SEQ ID NOs: 15-23 and is upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 40A-GG-39A-GG-25A-T-12A-G-3A (SEQ ID NO: 4). In some embodiments, the SP6 promoter comprises a sequence selected from the group consisting of SEQ ID NOs: 15-23 and is upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 40A-GG-39A-GG-38A-G-3A (SEQ ID NO: 5). In particular embodiments, the SP6 promoter comprises a sequence selected from the group consisting of SEQ ID NOs: 15-23 and is upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 34A-GG-31A-GG-27A-G-3A (SEQ ID NO: 3).
[117] Any promoter sequence that can be recognized by an T7 RNA polymerase may be used in the present invention. A suitable T7 promoter sequence comprises 5'- TAATACGACTCACTATAG-3' (SEQ ID NO: 24).
[118] In some embodiments, the T7 promoter sequence is upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 34A-GG-31A-GG-27A-G- 3 A (SEQ ID NO: 3). In some embodiments, the T7 promoter sequence is upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 40A-GG-39A- GG-25A-T-12A-G-3A (SEQ ID NO: 4). In some embodiments, the T7 promoter sequence is upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 40A-GG-39A-GG-38A-G-3A (SEQ ID NO: 5). In particular embodiments, the T7 promoter sequence is upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 34A-GG-31A-GG-27A-G-3A (SEQ ID NO: 3).
[119] In some embodiments, the T7 promoter sequence comprises SEQ ID NO: 24 and is upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 34A-GG-31A-GG-27A-G-3A (SEQ ID NO: 3). In some embodiments, the T7 promoter sequence comprises SEQ ID NO: 24 and is upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 40A-GG-39A-GG-25A-T-12A-G-3A
(SEQ ID NO: 4). In some embodiments, the T7 promoter sequence comprises SEQ ID NO: 24 and is upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 40A-GG-39A-GG-38A-G-3A (SEQ ID NO: 5). In particular embodiments, the T7 promoter sequence comprises SEQ ID NO: 24 and is upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 34A-GG-31A-GG-27A-G- 3A (SEQ ID NO: 3).
[120] Any promoter sequence that can be recognized by a Klebsiella phage KP34 RNA polymerase may be used in the present invention. A suitable Klebsiella phage KP34 core promoter sequence may comprise the nucleic acid sequence 5'-TAATGTTACAGGAGTA- 3' (SEQ ID NO: 25) or 5'-ATGTTACAGGAGTA-3' (SEQ ID NO: 26), or variants thereof Exemplary variants include TAATGTTACAGGAGTAGGnANiN2N3N4W (SEQ ID NO: 27), wherein NI-4 is any one of A, C, G or T, wherein W is A or T, and wherein n is number of guanosines. In particular embodiments, N1N2N3N4 is CAGA. Exemplary variants are shown in Table 3.
[121] In some embodiments, the KP34 promoter sequence is upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 34A-GG-31A-GG-27A-G- 3A (SEQ ID NO: 3). In some embodiments, the KP34 promoter sequence is upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 40A-GG-39A- GG-25A-T-12A-G-3A (SEQ ID NO: 4). In some embodiments, the KP34 promoter sequence is upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 40A-GG-39A-GG-38A-G-3A (SEQ ID NO: 5). In particular embodiments, the KP34 promoter sequence is upstream of a poly(A) tail sequence comprises or consists of the nucleic acid sequence 34A-GG-31A-GG-27A-G-3A (SEQ ID NO: 3).
[122] In some embodiments, the KP34 promoter comprises a sequence selected from the group consisting of SEQ ID NOs: 28-33 and is upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 34A-GG-31A-GG-27A-G-3A (SEQ ID NO: 3). In some embodiments, the KP34 promoter comprises a sequence selected from the group consisting of SEQ ID NOs: 28-33 and is upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 40A-GG-39A-GG-25A-T-12A-G-3A (SEQ ID NO: 4). In some embodiments, the KP34 promoter comprises a sequence selected from the group consisting of SEQ ID NOs: 28-33 and is upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 40A-GG-39A-GG-38A-G-3A (SEQ ID NO: 5). In particular embodiments, the KP34 promoter comprises a sequence selected from the group consisting of SEQ ID NOs: 28-33 and is upstream of a poly(A) tail sequence comprises or consists of the nucleic acid sequence 34A-GG-31A-GG-27A-G-3A (SEQ ID NO: 3 mRNA
[123] The templates of the invention, e.g. , DNA templates, are used for in vitro transcription (IVT) of messenger RNA (mRNA). As such, the templates comprise in 5’ to 3’ direction a first nucleic acid sequence encoding a polypeptide and a second nucleic acid sequence encoding a poly(A) tail of the invention.
[124] In some embodiments, the nucleic acid sequence encoding a polypeptide is optimized to facilitate more efficient transcription. For example, it may be optimized to remove cis- regulatory elements (e.g., termination signals, and protein binding sites), artificial recombination sites, chi sites, polymerase slippage sites, and/or other elements that could interfere with transcription. In some embodiments, the nucleic acid sequence encoding a polypeptide is optimized to reduce GC content or CpG dinucleotide content.
[125] In some embodiments, the nucleic acid sequence encoding a polypeptide is optimized to remove cryptic splice sites, repetitive sequences, RNA instability motifs, and other elements relevant to mRNA processing and/or stability. In some embodiments, the nucleic acid sequence encoding a polypeptide is optimized to stabilize the mRNA secondary structure or stable free energy of the mRNA.
[126] An mRNA comprises a nucleotide sequence encoding a peptide, polypeptide or protein. It also typically comprises a 5’ untranslated sequence (5’ UTR) and a 3’ untranslated sequence (3’ UTR).
5 ’ and 3 ’ untranslated regions
[127] The nucleotide sequences of a 5’ UTR or 3’ UTR encoded by a template of the invention, e.g., a DNA template, or present in an mRNA transcribed from it is typically different from the 5’ UTR present in a naturally occurring mRNA encoding the peptide, polypeptide or protein in question. For example, the 5’ UTR may include one or more elements that affect the mRNA’s stability or translation, e.g., an iron responsive element. The 5’ UTR may be about 50 to about 500 nucleotides in length.
[128] A 3’ UTR may include a binding site for proteins that affect an mRNA’s stability or location in a cell. Alternatively, or in addition, a 3’ UTR may comprise one or more binding sites for a miRNA. In some embodiments, a 3 ’ UTR may be about 50 to about 500 nucleotides in length.
[129] In some embodiments, a 3’ UTR includes one or more poly(A)denylation signal(s). A poly(A)denylation signal typically comprises a hexamer e.g., AAUAAA.
[130] For example, suitable 5’ and 3’ UTRs are described in WO 2012/075040, which is incorporated herein by reference.
[131] In certain embodiments, the 5’ and/or 3’ UTR sequences can be derived from mRNA which are stable (e.g., globin, actin, GAPDH, tubulin, histone, or citric acid cycle enzymes) to increase the stability of the mRNA. For example, a 5’ UTR sequence may include a partial sequence of a CMV immediate-early 1 (IE1) gene or immediate-early gene 9 (IE9), or a fragment thereof, to improve nuclease resistance and/or half-life of the mRNA. Also contemplated is the inclusion of a sequence encoding human growth hormone (hGH), or a fragment thereof, to the 3’ end or untranslated region of the mRNA.
[132] An exemplary 5’ UTR may include a sequence derived from a CMV immediate-early 1 (IE1) gene (U.S. Publication Nos. 2014/0206753 and 2015/0157565, each of which is incorporated herein by reference), or the sequences provided in Example 1 of U.S. Publication No. 2016/0151409, incorporated herein by reference. Alternatively, an exemplary 5’ UTR may include a sequence derived from an immediate-early 9 (IE9) gene of the Towne strain of CMV.
[133] In various embodiments, the 5’ UTR may be derived from the 5’ UTR of a TOP gene. TOP genes are typically characterized by the presence of a 5 ’-terminal oligopyrimidine (TOP) tract. Furthermore, most TOP genes are characterized by growth-associated translational regulation. However, TOP genes with a tissue specific translational regulation are also known. In certain embodiments, the 5’ UTR derived from the 5’ UTR of a TOP gene lacks the 5’ TOP motif (the oligopyrimidine tract) (e.g., U.S. Publication Nos. 2017/0029847, 2016/0304883, 2016/0235864, and 2016/0166710, each of which is incorporated herein by reference).
[134] In certain embodiments, the 5’ UTR is derived from a ribosomal protein Large 32 (L32) gene (U.S. Publication No. 2017/0029847, supra).
[135] In certain embodiments, the 5’ UTR is derived from the 5’ UTR of a hydroxysteroid (17-b) dehydrogenase 4 gene (HSD17B4) (U.S. Publication No. 2016/0166710, supra).
[136] In certain embodiments, the 5’ UTR is derived from the 5’ UTR of an ATP5A1 gene (U.S. Publication No. 2016/0166710, supra).
[137] In some embodiments, an internal ribosome entry site (IRES) is used instead of a 5’ UTR.
[138] A 5’ UTR suitable for use with the invention may comprise one of the nucleic acid sequences shown in Table 4.
[139] In some embodiments, the nucleic acid sequence comprised in the 5’ UTR is selected from any one of SEQ ID NOs: 34-39 and is located upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 34A-GG-31A-GG-27A-G-3A (SEQ ID NO: 3). In some embodiments, the nucleic acid sequence comprised in the 5’ UTR is selected
from any one of SEQ ID NOs: 34-39 and is located upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 40A-GG-39A-GG-25A-T-12A-G-3A (SEQ ID NO: 4). In some embodiments, the nucleic acid sequence comprised in the 5’ UTR is selected from any one of SEQ ID NOs: 34-39 and is located upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 40A-GG-39A-GG-38A-G- 3A (SEQ ID NO: 5).
[140] In particular embodiments, the nucleic acid sequence comprised in the 5’ UTR is selected from any one of SEQ ID NOs: 34-39 and is located upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 34A-GG-31A-GG-27A-G- 3A (SEQ ID NO: 3).
[141] A 3’ UTR suitable for use with the invention may comprise the following nucleotide sequence:
CGGGTGGCATCCCTGTGACCCCTCCCCAGTGCCTCTCCTGGCCCTGGAAGTTGC CACTCCAGTGCCCACCAGCCTTGTCCTAATAAAATTAAGTTGCATC (SEQ ID NO: 40).
[142] In some embodiments, the nucleic acid sequence comprised in the 3’ UTR comprises SEQ ID NO: 40 and is located upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 34A-GG-31A-GG-27A-G-3A (SEQ ID NO: 3). In some embodiments, the nucleic acid sequence comprised in the 3’ UTR comprises SEQ ID NOs: 40 and is located upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 40A-GG-39A-GG-25A-T-12A-G-3A (SEQ ID NO: 4). In some embodiments, the nucleic acid sequence comprised in the 3’ UTR comprises SEQ ID NOs: 40 and is located upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 40A-GG-39A-GG-38A-G-3A (SEQ ID NO: 5).
[143] In particular embodiments, the nucleic acid sequence comprised in the 3’ UTR comprises SEQ ID NOs: 40 and is located upstream of a poly(A) tail sequence that comprises or consists of the nucleic acid sequence 34A-GG-31A-GG-27A-G-3A (SEQ ID NO: 3).
Coding sequence
[144] The nucleotide sequence encoding a peptide, polypeptide or protein may be optimized regarding codon usage bias, codon adaptability, internal chi sites, ribosomal binding sites (e.g., IRES), premature poly(A) sites, Shine-Dalgarno (SD) sequences, and/or other elements
relevant to translation. In some embodiments, the nucleotide sequence encoding a peptide, polypeptide or protein is optimized regarding codon context, codon-anticodon interaction, translational pause sites, and/or other elements relevant to protein folding.
[145] A suitable sequence optimization method is described in International Patent Publication No. WO 2021/226461 Al, which is incorporated herein by reference.
Template propagation
[146] Various cells can be used for the high-yield production of templates of the invention. Bacterial cells that stably maintain high copy number plasmids are particularly suitable for use with the invention. In some embodiments, the bacterial cells used to propagate the template are E. coli cells. In particular embodiments, the E. coli strain is K12-derived or JM109-derived, e.g., DH5a, DHIOb, Stbl2, Stbl3, NEBStable™, or Endura™.
Assessing template stability
[147] The inventors found that high stability of templates for use in the in vitro transcription (IVT) of mRNA can be achieved during propagation in bacterial cells when the nucleic acid sequence encoding the poly (A) tail comprises three or more segments of 10-60 adenosines, wherein at least two of said segments consist of 40 adenosines or less, and wherein each segment is separated from the next segment by a spacer, wherein at least two of the spacers independently consists of two nucleotides other than adenosine. As demonstrated herein in Example 1, such templates were 100% stable during propagation in bacterial cells (specifically E. coli .
[148] Stability can be assessed by culturing a bacterial clone to provide a starter culture for a first time-period (e.g., 8 hours). The first time-period is typically selected in such a manner that the bacterial cells are in an exponential growth phase. The starter culture is then used to inoculate a first liquid culture. After a second time-period of incubation (e.g., 20 hours), an aliquot of this first liquid culture is taken to inoculate a second liquid culture. The second liquid culture is incubated for a third time-period (e.g., 20 hours). The second and third timeperiods are typically selected in such a manner that the bacterial cells are entering a stationary phase. Each of the cultures comprise an appropriate growth medium (e.g., LB broth) and are incubated at a temperature suitable for growth (e.g., 30°C, 34°C, or 37°C). The growth medium may be supplemented with an antibiotic to maintain the template in the bacterial cell.
[149] At the end of the respective second and third time-periods, an aliquot is taken from the first and second liquid cultures and plated on a suitable solid culture medium (e.g., after a 10-fold serial dilution). After a suitable incubation period (e.g., 16-24 hours), 10-24 individual bacterial colonies are isolated from each plate and cultured. After a suitable period of incubation (e.g., overnight), the template is isolated from each of the cultures. The purified template is then analyzed by sequencing or another suitable method, e.g., restriction fragment length polymorphism. The resulting data is compared to data obtained with the same analytical method for a reference template (typically the template isolated from the bacterial clone that was used to inoculate the starter culture).
[150] Figure 1 schematically outlines an assay that can be used to determine plasmid stability in abacterial cell such as A. coli. For A. coli, an 8-hour incubation period corresponds to 32 generations, and a 20-hour incubation period corresponds to 80 generations.
[151] In some embodiments, the template (e.g., a DNA plasmid) is capable of being propagated in A. coli cells at 34°C with less than 30% loss, e.g., less than 10%, less than 5% or less than 1% loss. In some embodiments, the template is capable of being propagated in A. coli cells over at least about 70 generations with less than 30% loss, e.g., less than 20% loss, less than 15% loss, or less than 10% loss. For templates of the invention, loss is more typically less than 5%, e.g., less than 1%. In some embodiments, the template is capable of being propagated in a bacterial cell (e.g, A. coli) for at least about 100 generations (e.g, at least about 110, 120, 130, 140, 150, 160, 170, 180, 190, or 200 generations) without loss or mutation to the nucleic acid sequence encoding the poly(A) tail. In particular embodiments, a template of the invention is capable of being propagated in a bacterial cell (e.g., A. coli) for about 100-200 generations (e.g., about 150-200 generations) without loss or mutation to the nucleic acid sequence encoding the poly(A) tail. In some embodiments, the template is capable of being propagated without loss or mutation to the nucleic acid sequence encoding the poly(A) tail in A. coli cells at 30°C or 34°C.
[152] The inventors also found that a nucleic acid encoding a segmented poly(A) tail confers comparable expression of the encoded protein compared to a nucleic acid encoding a non-segmented poly(A) tail. Quantitative methods for determining protein expression are known in the art, e.g., Western blotting, flow cytometry, enzyme-linked immunosorbent assay (ELISA) and mass spectrometry.
In Vitro Transcription (IVT)
[153] Various methods for synthesizing mRNA via IVT are described in US Patent Publication No. US 2018/0258423 and International Patent Publication No. WO 2021/168052A1, which are incorporated herein by reference, and can be used to practice the present invention. Briefly, IVT is typically performed with a reaction mixture comprising a DNA template, a pool of ribonucleotide triphosphates, a buffering reagent (that may include DTT), and one or more salts (e.g., MgCh and NaCl). A typical IVT reaction buffer may also include spermidine. The exact conditions will vary according to the specific application.
Cap analog
[154] In vitro transcribed mRNAs with a methylated 5’ cap structure can be translated very efficiently in vivo. For the large-scale production of mRNA, it can be more convenient and cost-effective to perform capping co-transcriptionally. Accordingly, the IVT process may include a cap analogue which is included in the mRNA molecules co-transcriptionally.
[155] During co-transcriptional capping, a cap analogue is included in the IVT reaction mixture. The cap analogue can be incorporated as the first “base” in a nascent RNA strand. The cap analogue may be Cap 0, Cap 1, Cap 2, m6Am, or a chemical cap analogue. For example, the following chemical cap analogues may be used to generate the 5 ’-guanosine cap structure according to the manufacturer’s instructions: 3’-O-Me-m7G(5’)ppp(5’)G (the ARCA cap); G(5’)ppp(5’)A; G(5’)ppp(5’)G; m7G(5’)ppp(5’)A; m7G(5’)ppp(5’)G; m7G(5')ppp(5')(2'OMeA)-pG; m7G(5’)ppp(5’)(2’OmeA)pU; m7G(5’)ppp(5’)(2’OmeG)pG (New England BioLabs, Ipswich, MA; TriLink Biotechnologies).
[156] In a specific embodiment, the in vitro transcribed mRNA may comprise a 5’ cap with the following structure:
Modified RNA
[157] In some embodiments, mRNA transcripts are synthesized with one or more modifications (z.e., as modified mRNA). In a particular embodiment, modified mRNA comprises a modified ribonucleotide, such as ribonucleotide analogue (e.g., adenosine analogue, guanosine analogue, cytidine analogue, and/or uridine analogue). The presence of a modified ribonucleotide may render the mRNA more stable and/or less immunogenic than a control mRNA with the same sequence but containing only naturally occurring ribonucleotides.
[158] In some embodiments, the modified ribonucleotide is selected from pseudouridine, N1 -methylpseudouridine, 2-thiouridine, 4’ -thiouridine, 5-methylcytidine, 2-thio-l -m ethyl- 1- deaza-pseudouridine, 2-thio-l-methyl-pseudouridine, 2-thio-5-aza-uridine, 2-thio- dihydropseudouridine, 2-thio-dihydrouridine, 2-thio-pseudouridine, 4-methoxy-2-thio- pseudouridine, 4-methoxy-pseudouridine, 4-thio-l-methyl-pseudouridine, 4-thio- pseudouridine, 5-aza-uridine, dihydropseudouridine, 5-methyluridine, 5-methoxyuridine, and 2’-O-methyl uridine.
[159] The modified ribonucleotide typically takes the place of a naturally occurring nucleotide. Accordingly, in one aspect, the in vitro transcribed mRNA of the invention comprises both unmodified and modified ribonucleotides. Such in vitro transcribed mRNA can be prepared by including a modified ribonucleotide in the IVT reaction mixture, typically in place of a naturally occurring ribonucleotide (e.g., N1 -methylpseudouridine in place of uridine). This results in in vitro transcribed mRNA in which 100% of the naturally occurring ribonucleotide is replaced by a corresponding modified ribonucleotide (e.g., 100% of the uridines are replaced with Nl-methyl-pseudouri dine).
[160] In some embodiments, the in vitro transcribed mRNA may be RNA wherein 25% of uracil residues are 2-thio-uracil and 25% of cytosine residues are 5-methylcytosine. Teachings for the use of such modified RNA are disclosed in US Patent Publication US 2012/0195936 and international publication WO 2011/012316, both of which are hereby incorporated by reference in their entirety.
Transcript length
[161] The length of an IVT reaction may depend on the length of the mRNA transcript. In a typical embodiment, the mRNA transcript comprises at least 500 ribonucleotides. In some
embodiments, the mRNA transcript comprises about 500 to about 3,000 ribonucleotides. In some embodiments, the mRNA transcript comprises about 700 to about 3,000 ribonucleotides. In some embodiments, the mRNA transcript comprises about 800 to about 2,000 ribonucleotides. In some embodiments, the mRNA transcript comprises about 1,000 to about 2,000 ribonucleotides. In some embodiments, the mRNA transcript comprises less than 3,000 ribonucleotides.
[162] Accordingly, the period over which IVT may take place to synthesize mRNA can vary widely. In some embodiments, IVT takes place over a period of about thirty minutes to about six hours. In some embodiments, IVT takes place over a period of about sixty to about ninety minutes.
Termination
[163] IVT can be terminated by removing the DNA template, e.g., through the addition of DNase I and a suitable buffer. For example, the polymerase reaction can be quenched by addition of DNase I and a DNase I buffer (100 mM Tris-HCl, 5 mM MgCh and 25 mM CaCh, pH 7.6 at lOx) to facilitate digestion of the double-stranded DNA template in preparation for purification.
Large-scale synthesis
[164] In some embodiments, the mRNA is synthesized in batches. In some embodiments, the present invention relates to the large-scale manufacture of mRNA.
[165] In some embodiments, about 1 g to about 100 kg of mRNA (e.g., 100 g to 10 kg, or 250 g to 5 kg) is synthesized in a single batch. In some embodiments, a batch comprises at least 1 g of in vitro transcribed mRNA (e.g., 5 g, 10 g, or 25 g). In other embodiments, a batch comprises at least 50 g of in vitro transcribed mRNA (e.g., 75 g, 100 g, 150 g, 200 g, or 250 g).
[166] In some embodiments, a method according to the invention synthesizes at least 500 g, 750 g, 1 kg, 5 kg, 10 kg, 50 kg, 100 kg, 1000 kg, or more of mRNA in a single batch. In some embodiments, 10 kg mRNA or more is synthesized in a single batch. In some embodiments, between 10 kg and 100 kg of mRNA is synthesized in a single batch.
Exemplary IVT conditions
[167] In some embodiments, a suitable reaction mixture comprises a double-stranded DNA template of the invention, an RNA polymerase, RNase inhibitor, pyrophosphatase, NTPs, 10 mM DTT and a reaction buffer (29.6 mM Tris-HCl, 3.8 mM MgCh, 8.9 mM KC1, and pH 7.5). In some embodiments, this reaction mixture is incubated at 37°C for the length of time needed to complete IVT of the mRNA transcript encoded by the DNA template.
[168] In some embodiments, a reaction mixture includes each NTP at a concentration ranging from 1-10 mM, a DNA template at a concentration ranging from 0.01-0.5 mg/mL, and an RNA polymerase at a concentration ranging from 0.01-0.1 mg/mL.
Post-synthesis processing
Enzymatic capping
[169] A 5’ cap structure can be added enzymatically after in vitro transcription (IVT) has been completed. Advantageously, at least 90% of in vitro transcribed mRNA subjected to separate enzymatic capping can comprise Capl structures.
[170] Several types of 5’ caps are known. A 7-m ethylguanosine cap (also referred to as “m7G” or “Cap 0”), comprises a guanosine that is linked through a 5 ’-5 ’-triphosphate bond to the first transcribed nucleotide. A 5’ cap is typically added as follows: first, an RNA terminal phosphatase removes one of the terminal phosphate groups from the 5’ nucleotide, leaving two terminal phosphates; guanosine triphosphate (GTP) is then added to the terminal phosphates via a guanylyl transferase, producing a 5’5’5 triphosphate linkage; and the 7- nitrogen of guanine is then methylated by a methyltransferase. Examples of cap structures include, but are not limited to, m7G(5’)ppp, (5’(A,G(5’)ppp(5’)A, and G(5’)ppp(5’)G. Additional cap structures are described in U.S. Publication Nos. US 2016/0032356 and US 2018/0125989, which are incorporated herein by reference.
[171] A vaccinia virus capping enzyme may be used to generate the Cap 0 structure: m7G(5’)ppp(5’)G. A Cap 1 structure may be generated using both vaccinia virus capping enzyme and a 2’-0 methyl-transferase to generate: m7G(5’)ppp(5’)G-2’-O-m ethyl. A Cap 2 structure may be generated from the Cap 1 structure followed by the 2’-O-methylation of the 5 ’-antepenultimate nucleotide using a 2’-0 methyltransferase. A Cap 3 structure may be generated from the Cap 2 structure followed by the 2’-O-methylation of the 5’- preantepenultimate nucleotide using a 2’-0 methyltransferase.
[172] Accordingly, a method of producing mRNA that comprises incubating a template of the invention in the presence of an RNA polymerase under conditions suitable for in vitro transcription (IVT) may further comprise a separate step of capping the resulting in vitro transcribed mRNA. The capping step may involve adding a capping enzyme (guanylyltransferase) and a guanosine triphosphate (GTP). A suitable capping enzyme may be derived from a Vaccinia virus (Vaccine virus guanylyltransferase). Typically, the capping step also comprises adding a guanine methyltransferase and a 2 -O-m ethyltransferase. Capping may be performed separately, e.g., after IVT.
Purification
[173] The mRNA transcripts obtained from an IVT reaction and/or a separate capping step may be purified. Various methods may be used to purify mRNA. In some embodiments, the mRNA transcripts are purified by precipitation and centrifugation. In some embodiments, the mRNA is purified by filtration using, e.g., Normal Flow Filtration (NFF) or Tangential Flow Filtration (TFF).
[174] Suitable purification methods include those described in published U.S. Application Nos. US 2016/0040154, US 2015/0376220, US 2018/0251755, US 2018/0251754, US 2020/0095571, US 2021/0388338, and US 2021/0002635, and in International Patent Publication No. WO 2022/072836, all of which are incorporated by reference herein.
Compositions
[175] As shown herein, the inventors found that a template encoding a segmented poly(A) tail of the invention yields mRNA transcripts with poly(A) tails having a single length distribution around the expected value, whereas a reference template encoding a standard non-segmented poly(A) tail can yield mRNA transcripts with poly(A) tails having a bimodal length distribution. Accordingly, the methods of the invention can be used to obtain a population of mRNA molecules with poly(A) tails having a single length distribution around a desired value. In some embodiments, at least 80% of the mRNAs in a population comprise a poly(A) tail of the desired length e.g., at least 85%, at least 90%, or at least 95%. In some embodiments, at least 80% (e.g., at least 85%, 90% or 95%) of the mRNAs in the population are within 10 nucleotides or less (e.g., 5 nucleotides or less, or 3 nucleotides or less) of the desired length value. Methods for determining the length of mRNA molecules are well- known and include capillary gel electrophoresis. A suitable assay for measuring poly(A) tail
length using a minor-groove binding dye and one or more ribonucleases is described in WO 2022/232499A1.
[176] In some embodiments, the poly(A) tail is composed of 150 nucleotides or less e.g., 140 nucleotides or less, 130 nucleotides or less, 120 nucleotides or less, 110 nucleotides or less, 100 nucleotides or less, 90 nucleotides or less or 80 nucleotides or less. In some embodiments, the poly(A) tail is composed of 80-150 nucleotides e.g., 100-125 nucleotides, 125-150 nucleotides, or 80-100 nucleotides.
[177] The inventors observed that the average tail length was within about 5%-l 0% of the desired value when using a template of the invention to obtain a population of mRNA molecules. In one embodiment, the desired tail length is at least 100 nucleotides (e.g., 100- 200 nucleotides). For example, the desired tail length may be a particular value (e.g., 100 or 125 nucleotides), and the average length of the tailed mRNA may be within about 10% or about 5% of that value.
[178] In some embodiments, at least 80% of the mRNA in a population is tailed and has an average tail length within 10% of the desired length. In some embodiments, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, or at least 99% of the mRNA in a population s tailed and has an average tail length within 5%-10% of the desired length.
[179] In some embodiments, at least 80% (e.g., at least 85%, at least 90%, or at least 95%) of the mRNAs in a population comprise a Capl structure at the 5’ end. Suitable methods for determining the presence of a Capl structure are known in the art and include mass spectroscopy (see, e.g., WO 2014/152659A1).
[180] An mRNA molecule obtained by a method of the invention may be formulated as a pharmaceutical composition. Formulations may include the addition of pharmaceutically acceptable excipients (e.g., one or more of a buffering such as Tris, a salt such as NaCl, or a sugar, e.g., a disaccharide such as sucrose or trehalose) to a population of mRNA molecules (e.g., after purification). Accordingly, in some embodiments, compositions comprising mRNA molecules of the invention may include one or more pharmaceutically acceptable excipients. Such excipients may be selected from a buffering agent (e.g., Tris), a salt (e.g., NaCl), and a sugar (e.g., a disaccharide such as sucrose or trehalose).
Lipid nanoparticles
[181] Compositions comprising an mRNA molecule in accordance with the invention (including pharmaceutical compositions) may further comprise one or more agents that protect the mRNA molecule from rapid degradation and/or enhance delivery of the mRNA to target cells in vivo. Suitable agents include lipid nanoparticles and other carrier molecules (e.g., complex-forming carriers such as protamine). For example, the mRNA molecule may be encapsulated in a lipid nanoparticle. Suitable lipid nanoparticles comprise a cationic lipid, one or more non-cationic lipids (e.g., DOPE and cholesterol) and a PEG-modified lipid (e.g., DMG-PEG2000).
Administration
[182] The compositions disclosed herein such as pharmaceutical compositions can be formulated for administration in any way known in the art of drug delivery, for example, orally, parenterally, intravenously, intramuscularly, subcutaneously, intradermally, transdermally, intrathecally, submucosally, sublingually, rectally, vaginally, etc. In some embodiments, the composition is formulated for sublingual administration, intramuscular administration, intradermal administration, subcutaneous administration, intravenous administration, intranasal administration, administration by inhalation, or intraperitoneal administration. In some embodiments, the composition is formulated for sublingual administration. In some embodiments, the composition (e.g., an immunogenic composition such as a vaccine) is formulated for intramuscular injection.
[183] The compositions disclosed herein such as pharmaceutical compositions may be packaged in a container, such as a prefilled syringe, a vial, or an autoinjector. In some embodiments, a composition (e.g., an immunogenic composition such as a vaccine) is packaged in a prefilled syringe. In some embodiments, the composition is packaged in a vial. In some embodiments, the composition is in an autoinjector. In other embodiments, the composition of the disclosure are packaged cartridges for patient-friendly autoinjector and infusion pump devices.
Immunogenic compositions and vaccines
[184] In some embodiments, a composition disclosed herein is an immunogenic composition or a vaccine. In some embodiments, the polypeptide encoded by an mRNA molecule in accordance with the invention is an antigenic protein for use in an immunogenic
composition or a vaccine, such as an antigenic protein from an influenza virus, a coronavirus, a RSV, a parainfluenza virus, a human immunodeficiency virus (HIV), a herpesvirus, a human papilloma virus, a rotavirus virus, a norovirus, a varicella zoster virus, a hepatitis virus, a paramyxovirus, a monkey pox virus, a parvovirus, an Ebola virus, a dengue virus, a hantavirus, a Zika virus, a west Nile virus, a poliovirus, or a rabies virus
[185] Immunogenic compositions (e.g., vaccines) are administered to a subject to prevent an infectious disease (e.g., virus infection, such as influenza, coronavirus, or a respiratory syncytial virus (RSV) infection) in the subject, to decrease the subject’s likelihood of contracting an infectious disease (e.g., virus infection), or to reduce the subject’s likelihood of getting serious ill from an infectious disease (e.g., virus infection, such as an influenza virus, coronavirus, or RSV infection).
[186] The composition disclosed herein may be used for the manufacture of an immunogenic composition (e.g., a vaccine) for use in immunizing a subject against an infectious disease (e.g., virus infection, such as an influenza virus, coronavirus, or RSV infection). In some embodiments, immunization comprises administering to a subject in need thereof an immunologically effective amount of any of the vaccines described herein. As used herein, the term “immunologically effective amount” or “therapeutically effective amount” means an amount sufficient to immunize a subject.
[187] Accordingly, in some embodiments, the disclosure provides a method of immunizing a subject comprising administering to the subject in need thereof an immunologically effective amount of any of the vaccines described herein. As used herein, “immunize” or “immunizing” means to induce in a subject a protective immune response against an infectious disease (e.g., viral infection, such as influenza, coronavirus, or RSV infection). Also provided is a method of reducing one or more symptoms of an infectious disease (e.g., viral infection, such as influenza, coronavirus, or RSV infection) comprising administering to a subject in need thereof a prophylactically effective amount of any of an immunogenic composition (e.g., a vaccine) described herein.
[188] The methods and uses of the immunogenic compositions and vaccines described herein include administration of a single dose to a subject (z.e., no booster dose). In some embodiments, the methods and uses of the vaccines described herein include prime-boost vaccination strategies. Prime-boost vaccination comprises administering a priming immunogenic composition (e.g., a priming vaccine) and then, after a period of time has
passed, administering to the subject a boosting immunogenic composition (e.g., a boosting vaccine). The immune response is “primed” upon administration of the priming immunogenic composition (e.g., the priming vaccine) and is “boosted” upon administration of the boosting immunogenic composition (e.g., the boosting vaccine).
[189] In some embodiments, the immunogenic compositions and vaccines described herein are administered to subjects, wherein the subjects can be any member of the animal kingdom. In some embodiments, the subject is a non-human animal. In some embodiments, the nonhuman subject is an avian (e.g., a chicken or a bird), a reptile, an amphibian, a fish, an insect, and/or a worm. In some embodiments, the non-human subject is a mammal (e.g., a ferret, a rodent, a mouse, a rat, a rabbit, a monkey, a dog, a cat, a sheep, cattle, a primate, and/or a Pig)-
[190] In some embodiments, the immunogenic compositions and vaccines described herein are administered to a human subject. In some embodiments, a human subject is 6 months of age or older, 6 months through 35 months of age, at least two years of age, at least 3 years of age, 36 months through 8 years of age, 9 years of age or older, at least 6 months of age and less than 5 years of age, at least 6 months of age and less than 18 years of age, or at least 3 years of age and less than 18 years of age. In some embodiments, the human subject is an infant (less than 36 months). In some embodiments, the human subject is a child or adolescent (less than 18 years of age). In some embodiments, the human subject is a child of at least 6 months of age and less than 5 years of age. In some embodiments, the human subject is at least 5 years of age and less than 60 years of age. In some embodiments, the human subject is at least 5 years of age and less than 65 years of age. In some embodiments, the human subject is elderly (at least 60 years of age or at least 65 years of age). In some embodiments, the human subject is a non-elderly adult (at least 18 years of age and less than 65 years of age or at least 18 years of age and less than 60 years of age).
Further embodiments
[191] The invention is described further by reference to the following numbered embodiments:
1. A template for in vitro transcription of messenger RNA (mRNA), wherein the template comprises in 5’ to 3’ direction a first nucleic acid sequence encoding a polypeptide and a second nucleic acid sequence encoding a poly(A) tail comprising three or more
segments of 10-60 adenosines, wherein at least two of said segments consist of 40 adenosines or less, wherein each segment is separated from the next segment by a spacer, wherein at least two of the spacers independently consists of two nucleic acids other than adenosine.
2. The template of embodiment 1, wherein further spacers independently consist of one or two nucleic acids other than adenosine.
3. The template of embodiment 1 or 2, wherein the poly(A) tail comprises three segments.
4. The template of any one of embodiments 1-3, wherein the poly(A) tail further comprises a fourth segment of less than 10 adenosines.
5. The template of embodiment 4, wherein the fourth segment is at the 3’ end of the poly (A) tail.
6. The template of embodiment 4 or 5, wherein the fourth segment consists of less than 5 adenosines (e.g., 3 adenosines).
7. The template of any one of the preceding embodiments, wherein each of the three segments consists of 25-40 adenosines.
8. The template of any one of embodiments 1-6, wherein the at least two of the three or more segments consist of 20 adenosines or less (e.g., 18 or 19 adenosines).
9. The template of embodiment 8, wherein the template comprises one segment of 50- 60 adenosines.
10. The template of embodiment 8, wherein the template comprises at least 5 segments (e.g., 6 segments) of 20 adenosines or less.
11. The template of any one of the preceding embodiments, wherein the one or two nucleic acids of each spacer independently is/are guanosine or cytidine.
12. The template of embodiment 11, wherein the one or two nucleic acids of each spacer is/are guanosine.
13. The template of embodiment 11, wherein the one or two nucleic acids of each spacer is/are cytidine.
14. The template of any one of the preceding embodiments, wherein the at least two of the spacers independently consist of two guanosines or two cytidines.
15. The template of embodiment 14, wherein the at least two spacers consist of two guanosines.
16. The template of embodiment 14, wherein the at least two spacers consist of two cytidines.
17. The template of any one of the preceding embodiments, wherein at least one of the further spacers consists of one guanosine or one cytidine.
18. The template of any one of the preceding embodiments, wherein the poly(A) tail is composed of 150 nucleotides or less.
19. The template of any one of the preceding embodiments, wherein the poly(A) tail is composed of 80-150 nucleosides, e.g., 100-125 nucleosides.
20. The template of any one of the preceding embodiments, wherein the template is a DNA template, e.g., a plasmid.
21. The template of any one of the preceding embodiments, wherein the template is capable of being stably propagated in E. coli cells at 34°C, optionally wherein the second nucleic acid sequence encoding the poly(A) tail has a stability of 90% or greater, e.g., 95% or greater, 98% or greater, or 99% or greater.
22. The template of any one of the preceding embodiments, wherein the template is capable of being propagated in A. coli cells at 30°C or 34°C without loss of the second nucleic acid sequence encoding the poly(A) tail.
23. The template of any one of the preceding embodiments, wherein the template is capable of being propagated in E. coli cells at 34°C with less than 30% loss of the second nucleic acid encoding the poly(A) tail, e.g., less than 10% loss, less than 5% loss, or less than 1% loss.
24. The template of any one of the preceding embodiments, wherein the template is capable of being propagated in A. coli cells at 30°C or 34°C without loss.
25. The template of any of the preceding embodiments, wherein the template is a ColEl plasmid.
26. The template of embodiment 25, wherein the ColEl plasmid comprises a Cer sequence.
27. The template of any one of the preceding embodiments, wherein the template further comprises a nucleic acid sequence encoding an antibiotic resistance gene.
28. The template of any one of the preceding embodiments, wherein the first nucleic acid sequence comprises a 5’ untranslated sequence (5’ UTR) and/or a 3’ untranslated sequence (3’ UTR).
29. The template of any one of the preceding embodiments, wherein template comprises a polymerase promoter operably linked to the first and second nucleic acid sequences.
30. The template of embodiment 29, wherein the promoter is an SP6, T7 or KP34 phage polymerase promoter.
31. A method of producing mRNA comprising incubating the template of any one of the preceding embodiments in the presence of an RNA polymerase under conditions suitable for in vitro transcription.
32. A method of improving stability of a homopolymeric sequence comprising at least 80 adenosines during propagation in a bacterial cells, wherein the method comprises modifying the homopolymeric sequence to comprise three or more segments of 10-60 adenosines, wherein at least two of said segments consist of 40 adenosines or less, wherein each segment is separated from the next segment by a spacer, wherein at least two of the spacers independently consists of two nucleic acids other than adenosine.
33. The method of embodiment 32, wherein the homopolymeric sequence is comprised in a vector suitable for propagation in a bacterial cell.
34. The method of embodiment 32 or 33, wherein the method further comprises transforming a bacterial cell with the vector comprising the modified homopolymeric sequence.
35. A population of mRNAs obtainable by the method of embodiment 31.
36. The population of mRNAs of embodiment 35, wherein at least 80% of the mRNAs comprise a poly(A) tail of the desired length.
37. An mRNA molecule with a 3’ poly(A) tail comprising three or more segments of 10- 60 adenosines, wherein at least two of said segments consist of 40 adenosines or less, wherein each segment is separated from the next segment by a spacer, wherein at least two of the spacers independently consists of two nucleic acids other than adenosine.
38. The RNA molecule of embodiment 37, wherein further spacers independently consist of one or two nucleic acids other than adenosine.
39. The mRNA molecule of embodiment 37 or 38, wherein the poly(A) tail comprises three segments.
40. The mRNA molecule of any one of embodiments 37-39, wherein the poly(A) tail further comprises a fourth segment of less than 10 adenosines.
41. The mRNA molecule of embodiment 40, wherein the fourth segment is at the 3’ end of the poly (A) tail.
42. The mRNA molecule of embodiment 40 or 41, wherein the fourth segment consists of less than 5 adenosines (e.g., 3 adenosines).
43. The mRNA molecule of any one of embodiments 37-42, comprising three segments, each consisting of 25-40 adenosines.
44. The mRNA molecule of any one of embodiments 37-42, wherein at least two of the three or more segments consist of 20 adenosines or less (e.g., 18 or 19 adenosines).
45. The mRNA molecule of embodiment 44, wherein the poly(A) tail comprises at least one segment of 50-60 adenosines.
46. The mRNA molecule of embodiment 44, wherein the poly(A) tail comprises at least 5 segments (e.g., 6 segments) of 20 adenosines or less.
47. The mRNA molecule of any one of embodiments 37-46, wherein the one or two nucleic acids of each spacer independently is/are guanosine or cytidine.
48. The mRNA molecule of embodiment 47, wherein the one or two nucleic acids of each spacer is/are guanosine.
49. The mRNA molecule of embodiment 47, wherein the one or two nucleic acids of each spacer is/are cytidine.
50. The mRNA molecule of any one of embodiments 37-49, wherein the at least two of the spacers independently consist of two guanosines or two cytidines.
51. The mRNA molecule of embodiment 50, wherein the at least two spacers consist of two guanosines.
52. The mRNA molecule of embodiment 50, wherein the at least two spacers consist of two cytidines.
53. The mRNA molecule of any one of embodiments 37-52, wherein at least one of the further spacers consists of one guanosine or one cytidine.
54. The mRNA molecule of any one of embodiments 37-53, wherein the poly(A) tail is composed of 150 nucleotides or less.
55. The mRNA molecule of any one of embodiments 37-54, wherein the poly(A) tail is composed of 80-150 nucleosides, e.g., 100-125 nucleosides.
56. A pharmaceutical composition comprising the mRNA molecule of any one of embodiments 37-55 and a pharmaceutically acceptable carrier.
57. The pharmaceutical composition of embodiment 56, wherein the mRNA molecule is encapsulated in a lipid nanoparticle.
58. The pharmaceutical composition of embodiment 56 or 57, wherein the pharmaceutical composition is an immunogenic composition (e.g., a vaccine).
59. The immunogenic composition according to embodiment 58 for use in immunizing a subject.
60. The immunogenic composition for use according to embodiment 59, wherein immunization prevents a virus infection in the subject, decreases the subject’s likelihood of contracting a virus infection, or reduces the subject’s likelihood of getting seriously ill from a virus infection.
61. The immunogenic composition for use according to embodiment 60, wherein immunization raises a protective immune response in the subject.
62. The immunogenic composition for use according to any one of embodiments 59-61, wherein the subject is a human.
63. The immunogenic composition for use according to embodiment 62, wherein the human is 6 months of age or older, less than 18 years of age, at least 6 months of age and less than 18 years of age, at least 18 years of age and less than 65 years of age, at least 6 months of age and less than 5 years of age, at least 5 years of age and less than 65 years of age, at least 60 years of age, or at least 65 years of age.
64. The immunogenic composition for use according to any one of embodiments 59-63, wherein immunization is performed intramuscularly, intradermally, subcutaneously, intravenously, intranasally, by inhalation, or intraperitoneally.
Equivalents
[192] Although methods and materials similar or equivalent to those described herein can be used in the practice or testing of the present invention, suitable methods and materials are described below.
EXAMPLES
[193] The following examples are included for illustrative purposes only and are not intended to limit the scope of the invention.
Example 1. Generating templates with high stability during propagation in bacterial cells
[194] This example illustrates the generation of templates that have high stability during propagation in bacterial cells.
[195] A template plasmid was prepared comprising a nucleic acid sequence encoding an mRNA. This nucleic acid sequence included the following elements: a 5’ untranslated region (5’ UTR), a protein coding sequence, and a 3’ untranslated region (3’ UTR). The 3’ UTR was followed by a multiple cloning site (MCS) for the insertion of poly(A) tail constructs. The template plasmid also included an RNA polymerase promoter (specific for SP6 RNA polymerase) operably linked to the nucleic acid sequence encoding the mRNA. In addition, the template plasmid also included a Cer sequence to increase plasmid stability and an antibiotic resistance gene (providing resistance to kanamycin) for the selection of transformants and plasmid maintenance. The template plasmid without a poly tail insert is identified as “m8” in subsequent examples.
[196] The protein coding sequence in the template plasmid encoded hemagglutinin of the H3 subtype (HA-H3) derived from influenza virus A (H3N2). This protein was chosen because it is expressed on the surface of target cells and its expression therefore can be detected easily using flow cytometry (see Example 5).
[197] The MCS was used for the insertion of four different nucleic acid sequences encoding a poly(A) tail. The poly(A) tail sequence encoded by each nucleic acid sequence is shown in Table 5.
[198] Escherichia coli (E. coll) cells were transformed with the resulting template plasmids (98A, G, 0.1 and M.2). Successful transformants were selected on kanamycin-containing solid medium. The template plasmid was isolated and sequenced. Transformants containing a plasmid with the desired poly(A) tail sequences were expanded in liquid medium to prepare a glycerol stock.
[199] To test plasmid stability, a frozen vial of glycerol stock was thawed to prepare a preculture in Luria-Bertani (LB) broth. The preculture was incubated for 8 hours at 30°C and then used to inoculate 25 ml of LB broth. After 20 hours of incubation, an aliquot of this culture (Pl) was taken to inoculate a second 25 ml LB broth culture (P2). The inoculum had an optical density of 0.005 at 600 nm and was also incubated for 20 hours under the same conditions as the Pl culture. At each step, plasmid maintenance was supported by the presence of 25 pg/ml of kanamycin in the growth medium. Duplicate experiments were performed. The only variable was the incubation temperature. It was either 30°C or 34°C. A 50 pl aliquot was taken of each culture Pl and P2 at the end of the incubation period and plated.
[200] 16 colonies were isolated from each plate and cultured in 1.2 ml of LB broth. Plasmid DNA was isolated (purified) from the resulting cultures by miniprep and subjected to Sanger sequencing to determine the presence of the intact nucleic acid sequence encoding the poly(A) tail. A schematic representation of the plasmid stability experiment is shown in Figure 1. The results are summarized in Table 6.
Table 6: Plasmid stability
[201] At an incubation temperature of 34°C, plasmid stability was higher for the tested poly(A) tail sequences that comprised three or more segments of 10-60 adenosines separated by at least two spacers of two nucleotides others than adenosine. The two plasmids in which at least two of the spacers consisted of two guanosines (plasmids 0.1 and M.2) had the highest stability. The poly(A) tail was maintained intact in 100% of the tested colonies. At 34°C, the plasmid in which each spacer consisted of a single guanosine (plasmid G) had higher stability than the plasmid with the non-segmented poly(A) tail (plasmid 98A) which was included as a control. 72% of the tested colonies contained an intact version of plasmid G, whereas only 46% of the tested colonies contained an intact version of plasmid 98A.
[202] At an incubation period of 30°C, the poly(A) tail sequence remained intact in 100% of the tested colonies comprising plasmids O. l and M.2. In contrast, only 69% and 57% of the tested colonies comprising plasmids 98A and G, respectively, had intact poly(A) tail inserts.
[203] This example illustrates that the templates encoding a poly(A) tail sequence comprising at least three segments of 10-60 adenosines separated by at least two spacers of two nucleotides others than adenosine have very high stability independent of the incubation temperature of the bacterial cells during propagation.
Example 2. Preparing in vitro transcribed mRNA
[204] In vitro transcribed mRNA was prepared as described in Example 1 of WO 2021/168052, which is incorporated herein by reference. Briefly, the template plasmids isolated in Example 1 were linearized by digestion with a restriction enzyme. For each gram of mRNA transcribed, a reaction containing a linearized template plasmid, an RNA polymerase (SP6), RNase inhibitor, pyrophosphatase, NTPs, DTT, and a buffering reagent
was prepared with RNase-free water. The reaction buffer (pH 7.5) comprised 29.6 mM Tris- HC1, 3.8 mM MgCh, 8.9 mM KC1. The reaction mixtures were incubated at 37°C for 60 to 90 min. In vitro transcribed mRNA prepared with ATP, UTP, CTP, and GTP is referred to herein as unmodified (abbreviated as “unmod”). Where indicated, UDP was replaced with N1 -methylpseudouridine to prepare in vitro transcribed mRNA comprising a modified ribonucleotide (abbreviated as “mod”). DNase I was added to stop the reaction, and the reaction mixture was incubated for 15 more minutes at 37°C. The resulting in vitro transcribed mRNA was purified.
Example 3. mRNA yield and transcript length
[205] This example demonstrates that the mRNA transcript length and yield from a template encoding a segmented poly(A) tail can be comparable to the yield from a template encoding a standard non-segmented poly A tail.
[206] A control plasmid m8 lacking a nucleic acid sequence encoding a poly(A) tail and plasmids 98A, G, O.l and M.2 as described in Example 1 were linearized by restriction enzyme digestion and used as templates for IVT reactions as described in Example 2. For each template, IVT reactions were performed in triplicate. The mRNA yield was assessed by UV spectrometry. The length of the mRNA transcripts was assessed by capillary gel electrophoresis.
[207] As shown in Table 5, plasmid M.2 contains a single thymidine as spacer after the third segment of 25 adenosines. The stability of plasmid M.2 was comparable to the stability of plasmid O.l . An alternative plasmid construct was created in which the single thymidine was replaced with adenosine to generate plasmid M.3 with the following poly(A) tail sequence: 40A-GG-39A-GG-38A-G-3A (SEQ ID NO: 5)
[208] In a separate set of experiments, plasmid M.3 was found to have the same stability profile as plasmid M.2.
[209] An acceptable mRNA yield of about 150 pg or greater were achieved with plasmids M.3 and O.l. As a comparison, the mRNA yield was about 200 pg with a template plasmid including a standard poly(A) tail (98 A) or no poly tail (m8). For plasmid M.2, the mRNA yield was 100 pg.
[210] Capillary gel electrophoresis confirmed that the length of the mRNA transcripts obtained from the IVT reaction was close to the expected size, although generally shorter.
This was consistent across the tested constructs. The transcript length was comparable to the length of transcripts obtained from plasmid 98A (encoding a standard non-segmented poly(A) tail), in particular taking into account that the segmented poly(A) tail encoded by plasmids G, 0.1, M.2 and M.3 was longer than that of plasmid 98A. In conclusion, the presence of a nucleic acid sequence encoding a segmented poly(A) tail in template plasmids G, 0.1, M.2 and M.3 did not negatively impact mRNA transcription.
[211] The results of the experiments described in this example are summarized in Table 7.
[212] This example demonstrates that the presence of a nucleic acid sequence encoding a segmented poly(A) tail does not negatively affect in vitro transcription (IVT) as assessed by mRNA transcript length and yield.
Example 4. poly(A) tail length
[213] This example demonstrates that a template encoding a segmented poly(A) tail yields mRNA transcripts with poly(A) tails having a single length distribution around the expected value.
[214] To confirm that the presence of a nucleic acid sequence encoding a segmented poly(A) tail did not result in a truncation of the poly(A) tail in the resulting mRNA transcripts, tails of two representative IVT reactions using plasmids 98A and 0.1 as templates were analyzed).
[215] The poly(A) tails of mRNA transcripts obtained from plasmid 98A, encoding a standard non-segmented poly(A) tail, were longer than expected and had a bimodal distribution. In contrast, only a single distribution of poly(A) tail length was observed in the
population of mRNA transcripts obtained from plasmid 0.1 and the vast majority of poly(A) tails had a length within 5 nucleotides of the expected tail length of 100 nucleotides.
[216] This example shows that, in contrast to a template encoding a standard nonsegmented poly(A) tail, a template encoding a segmented poly(A) tail can advantageously yield mRNA transcripts with poly(A) tails having a single length distribution around the desired value.
Example 5. Expression of the mRNA-encoded protein
[217] This example demonstrates that the presence of a segmented poly(A) tail on mRNA transcripts does not negatively affect expression of the mRNA-encoded protein.
[218] mRNA transcripts obtained from IVT reactions using plasmids m8, 98A, G, 0.1, M.2 and M.3 (see Example 3) were used for the transfection of cultured cells. The mRNA transcripts obtained from plasmid m8 were enzymatically tailed with a 120 nucleotide-long poly(A) tail prior to transfection. Expression was assessed both by Western blot and flow cytometry using an antibody specific for the influenza virus A HA-H3 protein. Western blot analysis on cell lysates obtained 24 hours after transfection confirmed that all transfected cells expressed the HA-H3 protein.
[219] HA surface expression was determined on intact cells 24 and 48 hours after transfection. At both time points, cells transfected with mRNA transcripts comprising a segmented poly(A) tail (G, 0.1, M.2 and M.3) expressed the HA protein on their surface at levels comparable to that observed on cells transfected with mRNA transcripts comprising a standard non-segmented tail (98A). Similarly, at both time points, HA expression on cells transfected with mRNA transcripts obtained from plasmid G, 0.1, M.2 and M.3 was noninferior to that on cells transfected with mRNA transcripts comprising a standard nonsegmented that was added enzymatically (m8).
[220] These results confirm that the presence of a segmented poly(A) tail on mRNA transcripts does not negatively affect expression of the mRNA-encoded protein.
Example 6. Stability comparison
[221] This example demonstrates that a template encoding a segmented poly(A) tail as described herein has increased plasmid stability compared to a template encoding a nonsegmented poly(A) tail as well as a template encoding a differently configured segmented poly(A) tail that forms part of the state of the art.
[222] Plasmid stability was determined as described in Example 1. A 50 pl aliquot was taken of culture P2 (see Figure 1) at the end of the incubation period and plated. The incubation temperature was 30°C. A template encoding a non-segmented poly(A) tail was used as a control (112A). Templates encoding differently configured segmented poly(A) tails as disclosed in W02020/074642 (Trepotec) and W02016/005324 (BioNTech 40 and BioNTech 30) were also included for comparison. Table 8 provides the tested poly(A) tail sequences.
[223] As shown in Figure 2, the template encoding a non-segmented poly(A) tail (112A) had much reduced stability as indicated in the variation of the tail length in the different bacterial colonies isolated from the P2 culture. A similar observation was also made for the control template comprising the segmented Trepotec poly(A) tail sequence.
[224] In contrast, templates comprising the segmented poly(A) tail sequences A18GG, A18CC, 0.1, O-C, A40GG, S-GG, and S-CC were stable, having poly(A) tail sequences of the expected length (see Figure 2). Control templates comprising BioNTech’ s segmented poly(A) tails (BioNTech 40 and BioNTech 30) were likewise stable.
[225] This example illustrates that templates encoding a poly(A) tail sequence comprising at least three segments of 10-60 adenosines separated by at least two spacers of two nucleotides other than adenosine are stable during propagation in bacterial cells. Stability was improved in comparison to a template comprising non-segmented poly(A) tail and a template encoding a differently configured segmented poly(A) tails.
Example 7. mRNA with segmented poly (A) tail yields improved translation efficiency
[226] This example demonstrates that an mRNA comprising a segmented poly(A) tail is translated more efficiently as result of increased mRNA stability.
[227] To assess whether the presence of a segmented poly(A) tail affects the production of the mRNA-encoded protein, expression of mRNA transcripts encoding an eGFP degron was analyzed. Briefly, HeLa cells were plated in a 96-well plate at a seeding density of 6000 cells/well. Following overnight incubation, Hoechst stain was added to the cells at a final concentration of 0.09 pM for 30 minutes prior to the addition of transfection reagents.
[228] mRNA transcripts were prepared from templates by IVT as described in Example 2. Templates comprised a nucleic acid sequence encoding the eGFP degron and a segmented test poly(A) tail sequence (S-GG, S-CC, A18GG, M-C or A40GG; see Table 8). In addition, templates comprising a non-segmented control poly(A) tail sequence (112A, 100A (SEQ ID NO: 48), 80A (SEQ ID NO: 49)) were used to prepare control mRNA. To benchmark the performance of the test mRNA transcripts, additional mRNA transcripts were prepared from templates comprising the Trepotech and BioNTech segmented poly(A) tail sequences shown in Table 8. Similar mRNA yields were achieved from each of these templates as found in Example 3.
[229] A transfection mix was prepared by diluting Lipofectamine MessengerMAX in OptiMEM followed by the addition of 5 pl of 100 ng/pl mRNA. Kinetic monitoring of protein expression was performed using the Operetta CLS Imaging system and Harmony software. Data points were collected at 60-minute intervals for 25 hours and then at 120- minute intervals for 26 hours. Data acquisition was performed on 16 fields/well using the following parameters: eGFP (Time 200 ms; Power 100%; Height -9 pm; Excitation 460-490 nm; Emission 500-550 nm) and Hoechst (Time 200 ms; Power 100%; Height -9 pm; Excitation 355-385 nm; Emission 430-500 nm). Manual thresholding was used for positive cell gating and at least 800 cells were sampled per well.
[230] As shown in Figure 3 A, surprisingly high expression was observed for eGFP degron- encoding mRNA transcripts with the segmented 0.1 poly(A) tail sequence. Expression of the eGFP degron, as assessed by median fluorescence intensity (MFI) corrected for background MFI, was statistically significantly higher for mRNA transcripts comprising the 0.1 poly(A) tail sequence in comparison to mRNA transcripts with the non-segmented 112A tail at both 6 hours and 24 hours post transfection. None of the other test mRNA transcript showed a statistically significant increase in eGFP degron expression relative to the 112A mRNA transcript control. A similar observation was also made in comparison to templates encoding prior art segmented poly(A) tail sequences (data not shown). Only mRNA transcripts with the BioNTech30 poly(A) tail sequences showed statistically significantly higher expression at 6 hours but not 24 hours post transfection.
[231] The eGFP degron expression data were used to calculate translation efficiency and intracellular stability values to assess the overall effect of the poly(A) tail sequence. The translation efficiency value was calculated by dividing the MFI maximum of a given mRNA transcript sample by the MFI maximum of the mRNA transcript reference sample comprising the non-segmented 112A tail sequence. The intracellular stability index value was calculated by subtracting the hours post transfection for the MFI maximum of the mRNA transcript reference sample from the hours post transfection for the MFI maximum of a given mRNA transcript sample. The values were then plotted as shown in Figure 3B. The BioNTech30, BioNTech40, and Trepotec poly(A) tail sequences are referred to as “BT30”, “BT40” and “T) in Figure 3B. As can be seen from this figure, most of the value clustered in the lower left corner of the plot. Only mRNA transcripts comprising the segmented 0.1 poly(A) tail sequence stood out. They had both exceptional intracellular stability and - likely as a consequence - very high translation efficiency.
[232] Overall, the data in this example confirm that the presence of a segmented poly(A) tail on mRNA transcripts does not negatively affect expression of the mRNA-encoded protein, consistent with the observations made in Example 6. Surprisingly, the data also indicate that the segmented 0.1 poly(A) tail sequence can provide mRNA transcripts with higher in vivo stability mRNA and consequently higher translation efficiency.
Example 8. mRNA reactogenicity
[233] This example demonstrates that mRNA transcripts comprising a segmented poly(A) tail sequence comprising at least three segments of 10-60 adenosines separated by at least two spacers of two nucleotides other than adenosine have low reactogenicity.
[234] Exogenous single-stranded and double-stranded RNA can induce an inflammatory response via Toll-like receptors (TLRs) 7/8 and 3, respectively. In particular, TLR signaling can trigger the expression of NF -KP/IRF3 -induced genes. THPl-Dual™ cells (Invivogen) were used to determine in vitro reactogenicity of mRNA transcripts comprising segmented poly(A) tail sequences. These cells are derived from the human THP-1 monocyte cell line by stable integration of two inducible reporter constructs: A reporter gene encoding secreted alkaline phosphatase (SEAP) is operably linked to NF-KP inducible promoter, and a reporter gene encoding secreted luciferase (Lucia) is operably linked to an Interferon-sensitive response element (ISRE).
[235] Non-differentiated THPl-Dual™ cells were electroporated with HAH3-encoding mRNA prepared as described in Example 2. The HAH3-encoding mRNA comprised poly(A) tail sequences A18GG, A18CC, O-C, M-C and S-CC, as described in Table 8. H3HA- encoding mRNA comprising a segmented poly(A) tail comprising single-base spacers as shown in Table 9 below were also assessed for reactogenicity. A non-segmented control (112A) was also included as a reference. Negative controls included a cell culture supernatant sample of untreated cells (depicted as “None”) and a cell culture supernatant sample of mock- treated cells (no mRNA; depicted as “None + EP”) .
[236] Reactogenicity was assessed by monitoring SEAP activity (Figure 4A) and Lucia activity (Figure 4B) using an Ensight Multimode Plate Reader (PE) following the addition of a SEAP detection reagent (Quanti-Blue) and a luciferase detection reagent (Quanti-Luc). Trilink’s eGFP mRNA was included was used as a positive control.
[237] As can be seen from Figures 4A and 4B, all tested mRNA transcripts resulted in minimal induction of reporter gene expression. These results confirm that the presence of the tested segmented poly(A) tails on mRNA transcripts does not increase reactogenicity.
Example 9. Thermostability
[238] This example demonstrates that the presence of a segmented poly(A) tail does not modify the thermostability of mRNA transcripts.
[239] Plasmids encoding the poly(A) tail sequences listed in Table 8 were used as templates for IVT reactions as described in Example 2. Stability of the mRNA transcripts was evaluated at -80°C, 4°C and 20°C over a period of 14 days and at 37°C over a period of 8 days. mRNA integrity was determined using capillary electrophoresis profiling. No statistically significant difference in mRNA stability was observed at these temperatures with all tested mRNA transcripts.
[240] This example illustrates that the presence of a segmented poly(A) tail sequence comprising at least three segments of 10-60 adenosines separated by at least two spacers of two nucleotides other than adenosine does not negatively affect mRNA thermostability.
Claims
1. A template for in vitro transcription of messenger RNA (mRNA), wherein the template comprises in 5’ to 3’ direction a first nucleic acid sequence encoding a polypeptide and a second nucleic acid sequence encoding a poly(A) tail comprising three or more segments of 10-60 adenosines, wherein at least two of said segments consist of 40 adenosines or less, wherein each segment is separated from the next segment by a spacer, wherein at least two of the spacers independently consists of two nucleotides other than adenosine.
2. The template of claim 1, wherein further spacers independently consist of one or two nucleotides other than adenosine.
3. The template of claim 1 or 2, wherein the poly(A) tail comprises three segments.
4. The template of any one of claims 1-3, wherein the poly(A) tail further comprises a fourth segment of less than 10 adenosines.
5. The template of claim 4, wherein the fourth segment is at the 3’ end of the poly(A) tail.
6. The template of claim 4 or 5, wherein the fourth segment consists of less than 5 adenosines (e.g., 3 adenosines).
7. The template of any one of the preceding claims, wherein each of the three segments consists of 25-40 adenosines.
8. The template of any one of the preceding claims, wherein the one or two nucleotides of each spacer independently is/are guanosine or cytidine.
9. The template of claim 8, wherein the one or two nucleotides of each spacer is/are guanosine.
10. The template of claim 8, wherein the one or two nucleotides of each spacer is/are cytidine.
11. The template of any one of the preceding claims, wherein the at least two of the spacers independently consist of two guanosines or two cytidines.
12. The template of claim 11, wherein the at least two spacers consist of two guanosines.
13. The template of claim 11, wherein the at least two spacers consist of two cytidines.
14. The template of any one of the preceding claims, wherein at least one of the further spacers consists of one guanosine or one cytidine.
15. The template of any one of the preceding claims, wherein the poly(A) tail is composed of 150 nucleotides or less.
16. The template of any one of the preceding claims, wherein the poly(A) tail is composed of 80-150 nucleotides, e.g., 100-125 nucleotides.
17. The template of any one of the preceding claims, wherein the template is a DNA template, e.g., a plasmid.
18. The template of any one of the preceding claims, wherein the template is capable of being stably propagated in E. coli cells at 34°C, optionally wherein the second nucleic acid sequence encoding the poly(A) tail has a stability of 90% or greater, e.g, 95% or greater, 98% or greater, or 99% or greater.
19. The template of any one of the preceding claims, wherein the template is capable of being propagated in E. coli cells at 30°C or 34°C without loss of the second nucleic acid sequence encoding the poly (A) tail.
20. The template of any one of the preceding claims, wherein the template is capable of being propagated in E. coli cells at 34°C with less than 30% loss of the second nucleic acid encoding the poly(A) tail, e.g., less than 10% loss, less than 5% loss, or less than 1% loss.
21. The template of any one of the preceding claims, wherein the template is capable of being propagated in A. coli cells at 30°C or 34°C without loss.
22. The template of any of the preceding claims, wherein the template is a ColEl plasmid.
23. The template of claim 22, wherein the ColEl plasmid comprises a Cer sequence.
24. The template of any one of the preceding claims, wherein the template further comprises a nucleic acid sequence encoding an antibiotic resistance gene.
25. The template of any one of the preceding claims, wherein the first nucleic acid sequence comprises a 5’ untranslated sequence (5’ UTR) and/or a 3’ untranslated sequence (3’ UTR).
26. The template of any one of the preceding claims, wherein template comprises a polymerase promoter operably linked to the first and second nucleic acid sequences.
27. The template of claim 26, wherein the promoter is an SP6, T7 or KP34 phage polymerase promoter.
28. A method of producing mRNA comprising incubating the template of any one of the preceding claims in the presence of an RNA polymerase under conditions suitable for in vitro transcription.
29. A method of improving stability of a homopolymeric sequence comprising at least 80 adenosines during propagation in a bacterial cells, wherein the method comprises modifying the homopolymeric sequence to comprise three or more segments of 10-60 adenosines, wherein at least two of said segments consist of 40 adenosines or less, wherein each segment is separated from the next segment by a spacer, wherein at least two of the spacers independently consists of two nucleotides other than adenosine.
30. The method of claim 29, wherein the homopolymeric sequence is comprised in a vector suitable for propagation in a bacterial cell.
31. The method of claim 29 or 30, wherein the method further comprises transforming a bacterial cell with the vector comprising the modified homopolymeric sequence.
32. A population of mRNAs obtainable by the method of claim 28.
33. The population of mRNAs of claim 32, wherein at least 80% of the mRNAs comprise a poly(A) tail of the desired length.
34. An mRNA molecule with a 3’ poly(A) tail comprising three or more segments of 10- 60 adenosines, wherein at least two of said segments consist of 40 adenosines or less, wherein each segment is separated from the next segment by a spacer, wherein at least two of the spacers independently consists of two nucleotides other than adenosine.
35. The RNA molecule of embodiment 34, wherein further spacers independently consist of one or two nucleotides other than adenosine.
36. The mRNA molecule of claim 34 or 35, wherein the poly(A) tail comprises three segments.
37. The mRNA molecule of any one of claims 34-36, wherein the poly(A) tail further comprises a fourth segment of less than 10 adenosines.
38. The mRNA molecule of claim 37, wherein the fourth segment is at the 3’ end of the poly (A) tail.
39. The mRNA molecule of claim 37 or 38, wherein the fourth segment consists of less than 5 adenosines (e.g., 3 adenosines).
40. The mRNA molecule of any one of claims 34-39, comprising three segments, each consisting of 25-40 adenosines.
41. The mRNA molecule of any one of claims 34-40, wherein the one or two nucleotides of each spacer independently is/are guanosine or cytidine.
42. The mRNA molecule of claim 41, wherein the one or two nucleotides of each spacer is/are guanosine.
43. The mRNA molecule of claim 41, wherein the one or two nucleotides of each spacer is/are cytidine.
44. The mRNA molecule of any one of claims 34-43, wherein the at least two of the spacers independently consist of two guanosines or two cytidines.
45. The mRNA molecule of claim 44, wherein the at least two spacers consist of two guanosines.
46. The mRNA molecule of claim 44, wherein the at least two spacers consist of two cytidines.
47. The mRNA molecule of any one of claims 34-46, wherein at least one of the further spacers consists of one guanosine or one cytidine.
48. The mRNA molecule of any one of claims 34-47, wherein the poly(A) tail is composed of 150 nucleotides or less.
49. The mRNA molecule of any one of claims 34-48, wherein the poly(A) tail is composed of 80-150 nucleotides, e.g., 100-125 nucleotides.
50. A pharmaceutical composition comprising the mRNA molecule of any one of claims 34-49 and a pharmaceutically acceptable carrier.
51. The pharmaceutical composition of claim 50, wherein the mRNA molecule is encapsulated in a lipid nanoparticle.
52. The pharmaceutical composition of claim 50 or 51, wherein the pharmaceutical composition is an immunogenic composition (e.g., a vaccine).
53. The immunogenic composition according to claim 52 for use in immunizing a subject.
54. The immunogenic composition for use according to claim 53, wherein immunization prevents a virus infection in the subject, decreases the subject’s likelihood of contracting a virus infection, or reduces the subject’s likelihood of getting seriously ill from a virus infection.
55. The immunogenic composition for use according to claim 54, wherein immunization raises a protective immune response in the subject.
56. The immunogenic composition for use according to any one of claims 53-55, wherein the subject is a human.
57. The immunogenic composition for use according to claim 56, wherein the human is 6 months of age or older, less than 18 years of age, at least 6 months of age and less than 18 years of age, at least 18 years of age and less than 65 years of age, at least 6 months of age and less than 5 years of age, at least 5 years of age and less than 65 years of age, at least 60 years of age, or at least 65 years of age.
58. The immunogenic composition for use according to any one of claims 53-57, wherein immunization is performed intramuscularly, intradermally, subcutaneously, intravenously, intranasally, by inhalation, or intraperitoneally.
Applications Claiming Priority (2)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| EP23306234 | 2023-07-18 | ||
| EP23306234.8 | 2023-07-18 |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| WO2025017151A1 true WO2025017151A1 (en) | 2025-01-23 |
Family
ID=88018230
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/EP2024/070452 Pending WO2025017151A1 (en) | 2023-07-18 | 2024-07-18 | Stable poly(a)-encoding messenger rna templates |
Country Status (1)
| Country | Link |
|---|---|
| WO (1) | WO2025017151A1 (en) |
Citations (28)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO2004011618A2 (en) * | 2002-07-29 | 2004-02-05 | Hmgene, Inc. | Methods of identifying adipocyte specific genes, the genes identified, and their uses |
| WO2005012875A2 (en) * | 2003-07-29 | 2005-02-10 | Bristol-Myers Squibb Company | Biomarkers of cyclin-dependent kinase modulation |
| WO2011012316A2 (en) | 2009-07-31 | 2011-02-03 | Ludwig-Maximilians-Universität | Rna with a combination of unmodified and modified nucleotides for protein expression |
| WO2012075040A2 (en) | 2010-11-30 | 2012-06-07 | Shire Human Genetic Therapies, Inc. | mRNA FOR USE IN TREATMENT OF HUMAN GENETIC DISEASES |
| US20140206753A1 (en) | 2011-06-08 | 2014-07-24 | Shire Human Genetic Therapies, Inc. | Lipid nanoparticle compositions and methods for mrna delivery |
| WO2014152659A1 (en) | 2013-03-14 | 2014-09-25 | Shire Human Genetic Therapies, Inc. | Quantitative assessment for cap efficiency of messenger rna |
| US20150157565A1 (en) | 2012-06-08 | 2015-06-11 | Shire Human Genetic Therapies, Inc. | Pulmonary delivery of mrna to non-lung target cells |
| US20150376220A1 (en) | 2014-04-25 | 2015-12-31 | Shire Human Genetic Therapies, Inc. | Methods for purification of messenger rna |
| WO2016005324A1 (en) | 2014-07-11 | 2016-01-14 | Biontech Rna Pharmaceuticals Gmbh | Stabilization of poly(a) sequence encoding dna sequences |
| US20160040154A1 (en) | 2013-03-14 | 2016-02-11 | Shire Human Genetic Therapies, Inc. | Methods for purification of messenger rna |
| US20160151409A1 (en) | 2013-03-15 | 2016-06-02 | Shire Human Genetic Therapies, Inc. | Synergistic enhancement of the delivery of nucleic acids via blended formulations |
| WO2016091391A1 (en) | 2014-12-12 | 2016-06-16 | Curevac Ag | Artificial nucleic acid molecules for improved protein expression |
| US20160166710A1 (en) | 2013-08-21 | 2016-06-16 | Curevac Ag | Method for increasing expression of rna-encoded proteins |
| US20160235864A1 (en) | 2013-11-01 | 2016-08-18 | Curevac Ag | Modified rna with decreased immunostimulatory properties |
| US20160304883A1 (en) | 2013-12-30 | 2016-10-20 | Curevac Ag | Artificial nucleic acid molecules |
| US20170029847A1 (en) | 2013-12-30 | 2017-02-02 | Curevac Ag | Artificial nucleic acid molecules |
| US20180125989A1 (en) | 2016-11-10 | 2018-05-10 | Translate Bio, Inc. | Ice-based lipid nanoparticle formulation for delivery of mrna |
| US20180251754A1 (en) | 2017-02-27 | 2018-09-06 | Translate Bio, Inc. | Methods for purification of messenger rna |
| US20180251755A1 (en) | 2017-02-27 | 2018-09-06 | Translate Bio, Inc. | Methods For Purification of Messenger RNA |
| US20180258423A1 (en) | 2017-02-27 | 2018-09-13 | Translate Bio, Inc. | Large scale synthesis of messenger rna |
| US20200095571A1 (en) | 2018-08-24 | 2020-03-26 | Translate Bio, Inc. | Methods for purification of messenger rna |
| WO2020074642A1 (en) | 2018-10-11 | 2020-04-16 | Ethris Gmbh | Plasmid containing a sequence encoding an mrna with a segmented poly(a) tail |
| US20210002635A1 (en) | 2019-05-15 | 2021-01-07 | Translate Bio, Inc. | Methods for purification of messenger rna |
| WO2021168052A1 (en) | 2020-02-18 | 2021-08-26 | Translate Bio, Inc. | Improved processes for in vitro transcription of messenger rna |
| WO2021226461A1 (en) | 2020-05-07 | 2021-11-11 | Translate Bio, Inc. | Generation of optimized nucleotide sequences |
| US20210388338A1 (en) | 2018-11-08 | 2021-12-16 | Translate Bio, Inc. | Methods and Compositions for Messenger RNA Purification |
| WO2022072836A2 (en) | 2020-10-01 | 2022-04-07 | Translate Bio, Inc. | Methods for purification of messenger rna |
| WO2022232499A1 (en) | 2021-04-29 | 2022-11-03 | Translate Bio, Inc. | Methods for measuring poly a tail length |
-
2024
- 2024-07-18 WO PCT/EP2024/070452 patent/WO2025017151A1/en active Pending
Patent Citations (30)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO2004011618A2 (en) * | 2002-07-29 | 2004-02-05 | Hmgene, Inc. | Methods of identifying adipocyte specific genes, the genes identified, and their uses |
| WO2005012875A2 (en) * | 2003-07-29 | 2005-02-10 | Bristol-Myers Squibb Company | Biomarkers of cyclin-dependent kinase modulation |
| WO2011012316A2 (en) | 2009-07-31 | 2011-02-03 | Ludwig-Maximilians-Universität | Rna with a combination of unmodified and modified nucleotides for protein expression |
| US20120195936A1 (en) | 2009-07-31 | 2012-08-02 | Ethris Gmbh | Rna with a combination of unmodified and modified nucleotides for protein expression |
| WO2012075040A2 (en) | 2010-11-30 | 2012-06-07 | Shire Human Genetic Therapies, Inc. | mRNA FOR USE IN TREATMENT OF HUMAN GENETIC DISEASES |
| US20140206753A1 (en) | 2011-06-08 | 2014-07-24 | Shire Human Genetic Therapies, Inc. | Lipid nanoparticle compositions and methods for mrna delivery |
| US20150157565A1 (en) | 2012-06-08 | 2015-06-11 | Shire Human Genetic Therapies, Inc. | Pulmonary delivery of mrna to non-lung target cells |
| US20160040154A1 (en) | 2013-03-14 | 2016-02-11 | Shire Human Genetic Therapies, Inc. | Methods for purification of messenger rna |
| WO2014152659A1 (en) | 2013-03-14 | 2014-09-25 | Shire Human Genetic Therapies, Inc. | Quantitative assessment for cap efficiency of messenger rna |
| US20160032356A1 (en) | 2013-03-14 | 2016-02-04 | Shire Human Genetic Therapies, Inc. | Quantitative assessment for cap efficiency of messenger rna |
| US20160151409A1 (en) | 2013-03-15 | 2016-06-02 | Shire Human Genetic Therapies, Inc. | Synergistic enhancement of the delivery of nucleic acids via blended formulations |
| US20160166710A1 (en) | 2013-08-21 | 2016-06-16 | Curevac Ag | Method for increasing expression of rna-encoded proteins |
| US20160235864A1 (en) | 2013-11-01 | 2016-08-18 | Curevac Ag | Modified rna with decreased immunostimulatory properties |
| US20160304883A1 (en) | 2013-12-30 | 2016-10-20 | Curevac Ag | Artificial nucleic acid molecules |
| US20170029847A1 (en) | 2013-12-30 | 2017-02-02 | Curevac Ag | Artificial nucleic acid molecules |
| US20150376220A1 (en) | 2014-04-25 | 2015-12-31 | Shire Human Genetic Therapies, Inc. | Methods for purification of messenger rna |
| WO2016005324A1 (en) | 2014-07-11 | 2016-01-14 | Biontech Rna Pharmaceuticals Gmbh | Stabilization of poly(a) sequence encoding dna sequences |
| WO2016091391A1 (en) | 2014-12-12 | 2016-06-16 | Curevac Ag | Artificial nucleic acid molecules for improved protein expression |
| US20180125989A1 (en) | 2016-11-10 | 2018-05-10 | Translate Bio, Inc. | Ice-based lipid nanoparticle formulation for delivery of mrna |
| US20180251755A1 (en) | 2017-02-27 | 2018-09-06 | Translate Bio, Inc. | Methods For Purification of Messenger RNA |
| US20180251754A1 (en) | 2017-02-27 | 2018-09-06 | Translate Bio, Inc. | Methods for purification of messenger rna |
| US20180258423A1 (en) | 2017-02-27 | 2018-09-13 | Translate Bio, Inc. | Large scale synthesis of messenger rna |
| US20200095571A1 (en) | 2018-08-24 | 2020-03-26 | Translate Bio, Inc. | Methods for purification of messenger rna |
| WO2020074642A1 (en) | 2018-10-11 | 2020-04-16 | Ethris Gmbh | Plasmid containing a sequence encoding an mrna with a segmented poly(a) tail |
| US20210388338A1 (en) | 2018-11-08 | 2021-12-16 | Translate Bio, Inc. | Methods and Compositions for Messenger RNA Purification |
| US20210002635A1 (en) | 2019-05-15 | 2021-01-07 | Translate Bio, Inc. | Methods for purification of messenger rna |
| WO2021168052A1 (en) | 2020-02-18 | 2021-08-26 | Translate Bio, Inc. | Improved processes for in vitro transcription of messenger rna |
| WO2021226461A1 (en) | 2020-05-07 | 2021-11-11 | Translate Bio, Inc. | Generation of optimized nucleotide sequences |
| WO2022072836A2 (en) | 2020-10-01 | 2022-04-07 | Translate Bio, Inc. | Methods for purification of messenger rna |
| WO2022232499A1 (en) | 2021-04-29 | 2022-11-03 | Translate Bio, Inc. | Methods for measuring poly a tail length |
Non-Patent Citations (6)
| Title |
|---|
| JIA JINPING ET AL: "Annotation and expression profile analysis of 2073 full-length cDNAs from stress-induced maize ( Zea mays L.) seedlings", THE PLANT JOURNAL, vol. 48, no. 5, 31 October 2006 (2006-10-31), GB, pages 710 - 727, XP093203787, ISSN: 0960-7412, DOI: 10.1111/j.1365-313X.2006.02905.x * |
| KO CHEMYONG ET AL: "Xenopus hoxc8 during early development", BIOCHEMICAL AND BIOPHYSICAL RESEARCH COMMUNICATIONS, vol. 300, no. 1, 1 January 2003 (2003-01-01), Amsterdam NL, pages 9 - 15, XP093203674, ISSN: 0006-291X, DOI: 10.1016/S0006-291X(02)02699-2 * |
| PEUBEZ ET AL., MICROB CELL FACT, vol. 9, 2010, pages 65 |
| PREISS ET AL., RNA, vol. 4, no. 11, 1998, pages 1321 - 1331 |
| SACHS, CURR OPIN CELL BIOL, vol. 2, no. 6, 1990, pages 1092 - 1098 |
| TREPOTEC ET AL., RNA, vol. 25, no. 4, 2019, pages 507 - 518 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| US12480117B2 (en) | Compositions comprising modified circular polyribonucleotides and uses thereof | |
| AU2018385757B2 (en) | Compositions comprising circular polyribonucleotides and uses thereof | |
| KR102186497B1 (en) | Artificial nucleic acid molecules | |
| JP5735927B2 (en) | Re-engineering the primary structure of mRNA to enhance protein production | |
| JP2025090772A (en) | Cyclic polyribonucleotide and pharmaceutical composition thereof | |
| AU2005232565B2 (en) | MicroRNA and methods for inhibiting same | |
| CA3164192A1 (en) | Modified guide rnas for gene editing | |
| WO2023222114A1 (en) | Methods of making circular rna | |
| CN115916993A (en) | In vitro production and purification of therapeutic mRNA | |
| JP2024504981A (en) | Novel engineered and chimeric nucleases | |
| CN111979273A (en) | Method for preparing humanized ACE2 mouse model | |
| WO2005117557A2 (en) | Expression system | |
| WO2023227124A1 (en) | Skeleton for constructing mrna in-vitro transcription template | |
| WO2025102994A1 (en) | Rna in-vitro cyclization and rolling circle translation method based on group i intron, and use | |
| CN118679248A (en) | Synthesis production of circular DNA vectors | |
| EP4269585A1 (en) | Guide rna for editing polyadenylation signal sequence of target rna | |
| WO2024145248A1 (en) | Compositions and methods for generating circular rna | |
| WO2025017151A1 (en) | Stable poly(a)-encoding messenger rna templates | |
| EP2814964B1 (en) | Nucleic acid comprising or coding for a histone stem-loop and a poly(a) sequence or a polyadenylation signal for increasing the expression of an encoded allergenic antigen or an autoimmune self-antigen | |
| KR102129584B1 (en) | Insertion method of target genes in linearized vectors | |
| JP2015180203A (en) | REENGINEERING mRNA PRIMARY STRUCTURE FOR ENHANCED PROTEIN PRODUCTION | |
| KR102918706B1 (en) | Modified RNA for preparing mRNA vaccines and therapeutics | |
| US20250101480A1 (en) | Compositions and methods for preparing capped mrna | |
| WO2025076037A1 (en) | Circular rna vectors comprising modified nucleotides |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| 121 | Ep: the epo has been informed by wipo that ep was designated in this application |
Ref document number: 24746687 Country of ref document: EP Kind code of ref document: A1 |