WO2024256580A1 - Concurrent sequencing with spatially separated rings - Google Patents
Concurrent sequencing with spatially separated rings Download PDFInfo
- Publication number
- WO2024256580A1 WO2024256580A1 PCT/EP2024/066446 EP2024066446W WO2024256580A1 WO 2024256580 A1 WO2024256580 A1 WO 2024256580A1 EP 2024066446 W EP2024066446 W EP 2024066446W WO 2024256580 A1 WO2024256580 A1 WO 2024256580A1
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- immobilised
- proportion
- primers
- solid support
- immobilised primers
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Pending
Links
Classifications
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q1/00—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions
- C12Q1/68—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions involving nucleic acids
- C12Q1/6806—Preparing nucleic acids for analysis, e.g. for polymerase chain reaction [PCR] assay
Definitions
- the invention relates to solid supports and methods for use in nucleic acid sequencing, in particular solid supports and methods for use in concurrent sequencing.
- next-generation sequencing technologies
- a nucleic acid cluster is created on a flow cell by amplifying an original template nucleic acid strand. Sequencing cycles may be performed as complementary strands of the template nucleic acids are being synthesized, i.e., using sequencing-by-synthesis (SBS) processes.
- SBS sequencing-by-synthesis
- deoxyribonucleic acid analogs conjugated to fluorescent labels are hybridized to the template nucleic acids, and excitation light sources are used to excite the fluorescent labels on the deoxyribonucleic acid analogs.
- Detectors capture fluorescent emissions from the fluorescent labels and identify the deoxyribonucleic acid analogs.
- the sequence of the template nucleic acids may be determined by repeatedly performing such sequencing cycles.
- NGS allows for the sequencing of a number of different template nucleic acids simultaneously, which has significantly reduced the cost of sequencing in the last twenty years.
- a solid support comprising: a plurality of first immobilised primers, and a plurality of second immobilised primers, wherein a first proportion of the first immobilised primers are configured to be cleavable under first cleavage conditions and are unblocked, wherein a second proportion of the first immobilised primers are configured to be non-cleavable under first cleavage conditions and are blocked, wherein a third proportion of the second immobilised primers are configured to be cleavable under second cleavage conditions and are blocked; and wherein a fourth proportion of the second immobilised primers are configured to be non-cleavable under second cleavage conditions and are unblocked.
- a ratio between unblocked immobilised primers and blocked immobilised primers is between 20:80 to 80:20.
- the ratio between unblocked immobilised primers and blocked immobilised primers is between 50:50 to 75:25.
- the ratio between unblocked immobilised primers and blocked immobilised primers is between 60:40 to 70:30.
- the ratio between unblocked immobilised primers and blocked immobilised primers is about 2: 1.
- the first immobilised primers of the second proportion that are blocked each comprise a blocking group at a 3 ’ end of the first immobilised primer.
- the blocking group is selected from the group consisting of: a hairpin loop, a deoxynucleotide, a deoxyribonucleotide, a hydrogen atom instead of a 3 ’-OH group, a phosphate group, a phosphorothioate group, a propyl spacer, a modification blocking the 3 ’-hydroxyl group, or an inverted nucleobase.
- the blocking group is a phosphate group or a hairpin loop.
- the blocking group is a phosphate group.
- the second immobilised primers of the third proportion that are blocked each comprise a blocking group at a 3 ’ end of the second immobilised primer.
- the blocking group is selected from the group consisting of: a hairpin loop, a deoxynucleotide, a deoxyribonucleotide, a hydrogen atom instead of a 3 ’-OH group, a phosphate group, a phosphorothioate group, a propyl spacer, a modification blocking the 3 ’-hydroxyl group, or an inverted nucleobase.
- the blocking group is a phosphate group or a hairpin loop. In one aspect, the blocking group is a phosphate group.
- the first immobilised primers of the first proportion are configured to be cleavable by a thermal trigger, a light trigger, and/or a chemical/biochemical trigger.
- the first immobilised primers of the first proportion are configured to be cleavable by a metal catalyst.
- the first immobilised primers of the first proportion are configured to be cleavable by a transition metal catalyst.
- the first immobilised primers of the first proportion are configured to be cleavable by a palladium-based or a nickel-based catalyst.
- the first immobilised primers of the first proportion are configured to be cleavable by a palladium-based catalyst.
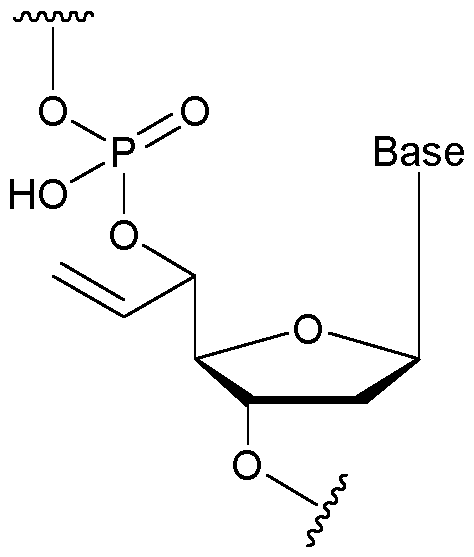
- each first immobilised primer of the first proportion comprises a nucleotide that comprises an allyl group.
- the allyl group is attached to a sugar group of the nucleotide.
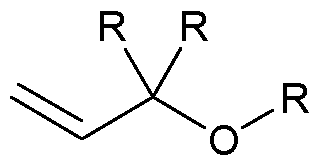
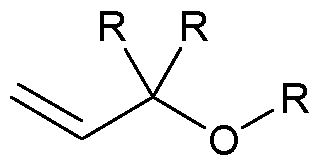
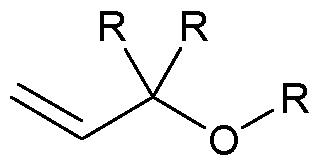
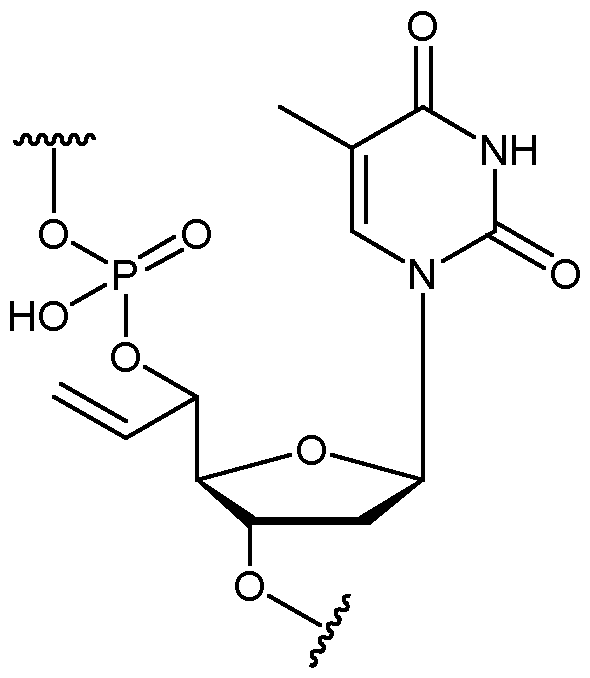
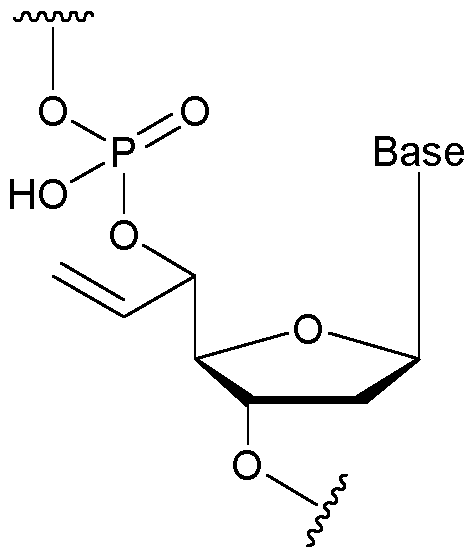
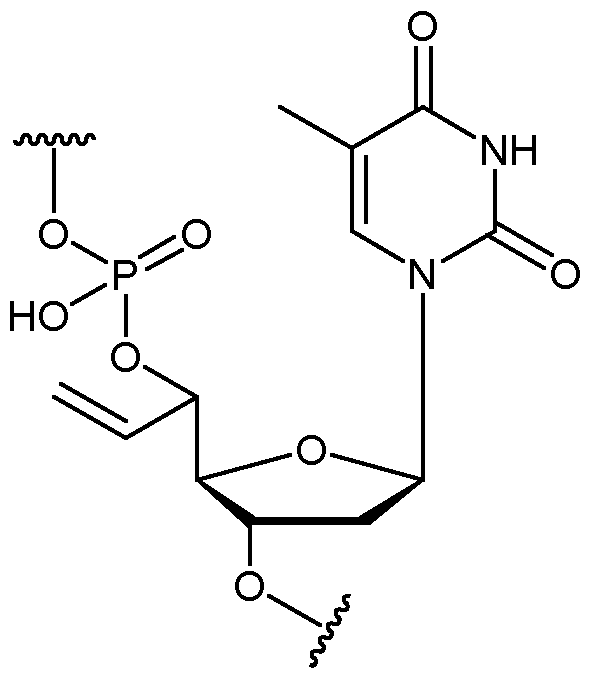
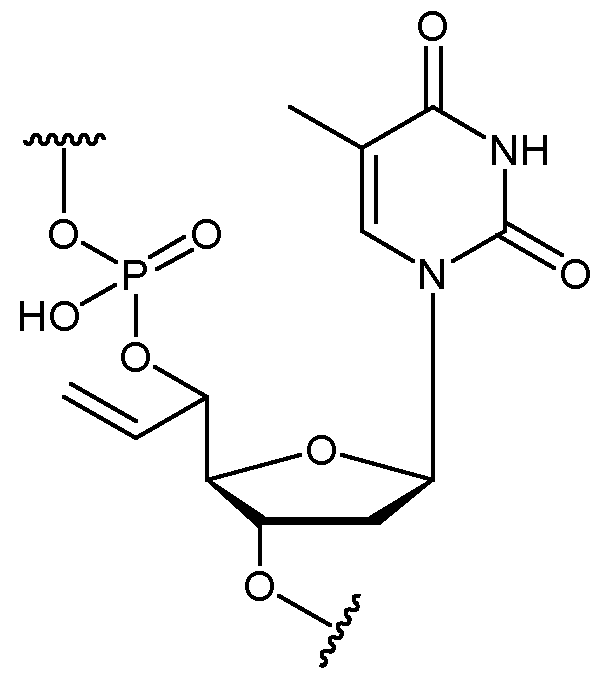
- the allyl group has a structure according to the following: wherein R represents an attachment point to hydrogen (in which case, R is hydrogen) or the remainder of the sugar group.
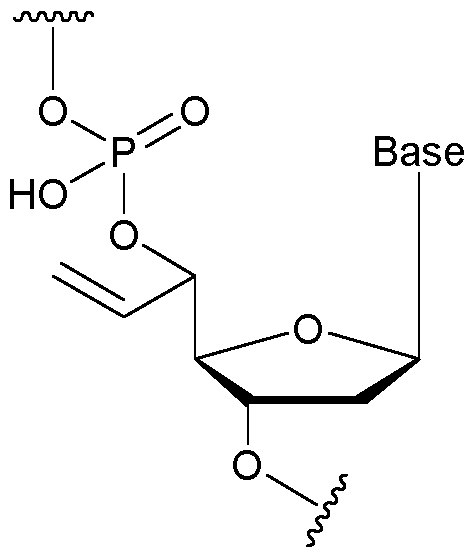
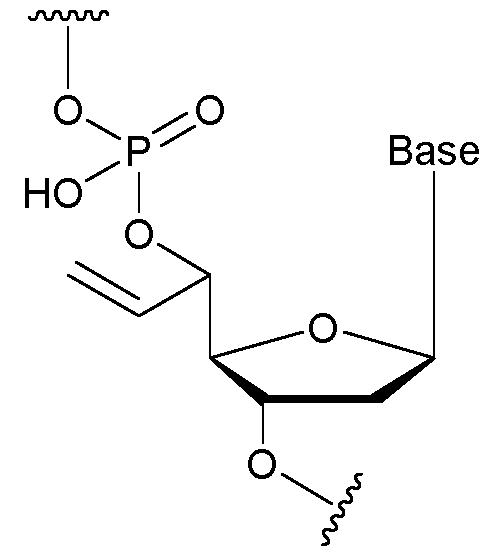
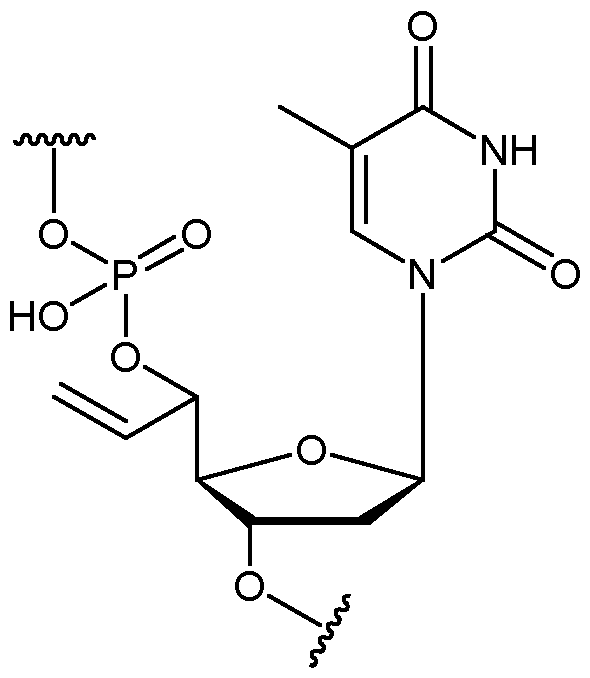
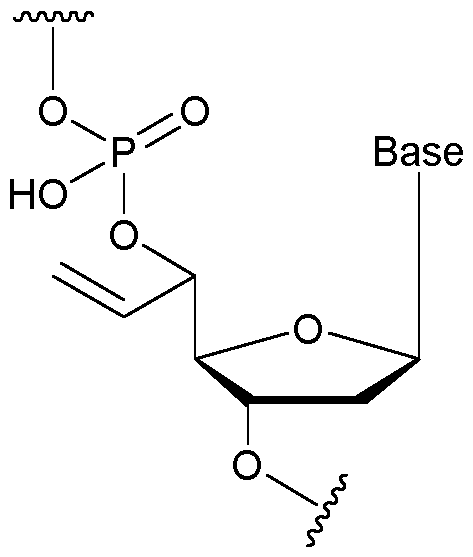
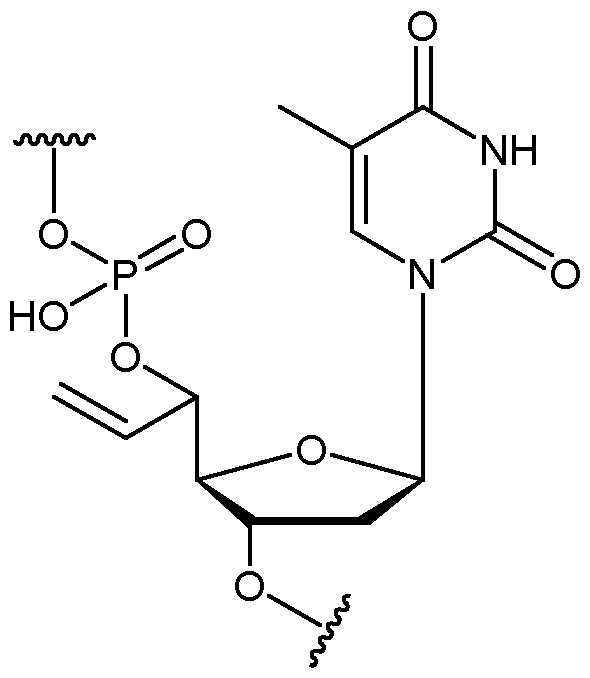
- the nucleotide has a structure according to the following: wherein represents an attachment point of the nucleotide to the rest of the first immobilised primer of the first proportion, and wherein “base” represents adenine, guanine, cytosine, thymine or uracil, or a derivative thereof.
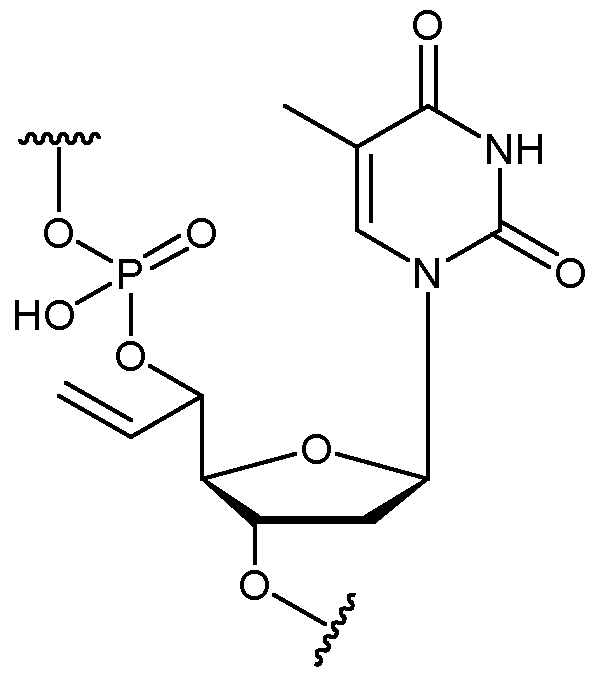
- the nucleotide has a structure according to the following: wherein represents an attachment point of the nucleotide to the rest of the first immobilised primer of the first proportion.
- the first immobilised primers of the first proportion are configured to be cleavable by a glycosylase.
- each first immobilised primer of the first proportion comprises a nucleobase which is not selected from guanine, cytosine, adenine or thymine when the first immobilised primer is a DNA sequence; or wherein each first immobilised primer of the first proportion comprises a nucleobase which is not selected from guanine, cytosine, adenine or uracil when the first immobilised primer is an RNA sequence.
- each first immobilised primer of the first proportion comprises oxoguanine (e.g. 8- oxoguanine) or uracil when the first immobilised primer is a DNA sequence, or wherein each first immobilised primer of the first proportion comprises oxoguanine (e.g. 8-oxoguanine) when the first immobilised primer is an RNA sequence.
- oxoguanine e.g. 8- oxoguanine
- uracil when the first immobilised primer is a DNA sequence
- each first immobilised primer of the first proportion comprises oxoguanine (e.g. 8-oxoguanine) when the first immobilised primer is an RNA sequence.
- the second immobilised primers of the third proportion are configured to be cleavable by a metal catalyst. In one aspect, the second immobilised primers of the third proportion are configured to be cleavable by a transition metal catalyst.
- the second immobilised primers of the third proportion are configured to be cleavable by a palladium-based or a nickel-based catalyst.
- the second immobilised primers of the third proportion are configured to be cleavable by a palladium-based catalyst.
- each second immobilised primer of the third proportion comprises a nucleotide that comprises an allyl group.
- the allyl group is attached to a sugar group of the nucleotide.
- the allyl group has a structure according to the following: wherein R represents an attachment point to hydrogen (in which case, R is hydrogen) or the remainder of the sugar group.
- the nucleotide has a structure according to the following: wherein represents an attachment point of the nucleotide to the rest of the second immobilised primer of the third proportion, and wherein “base” represents adenine, guanine, cytosine, thymine or uracil, or a derivative thereof.
- the nucleotide has a structure according to the following: wherein represents an attachment point of the nucleotide to the rest of the second immobilised primer of the third proportion.
- the second immobilised primers of the third proportion are configured to be cleavable by a glycosylase.
- each second immobilised primer of the third proportion comprises a nucleobase which is not selected from guanine, cytosine, adenine or thymine when the second immobilised primer is a DNA sequence; or wherein each second immobilised primer of the third proportion comprises a nucleobase which is not selected from guanine, cytosine, adenine or uracil when the second immobilised primer is an RNA sequence.
- each second immobilised primer of the third proportion comprises oxoguanine (e.g. 8-oxoguanine) or uracil when the second immobilised primer is a DNA sequence, or wherein each second immobilised primer of the third proportion comprises oxoguanine (e.g. 8-oxoguanine) when the second immobilised primer is an RNA sequence.
- oxoguanine e.g. 8-oxoguanine
- uracil when the second immobilised primer is a DNA sequence
- each second immobilised primer of the third proportion comprises oxoguanine (e.g. 8-oxoguanine) when the second immobilised primer is an RNA sequence.
- first cleavage conditions and the second cleavage conditions are the same or are different.
- the first cleavage conditions and the second cleavage conditions are the same.
- the first proportion of first immobilised primers that are cleavable are further configured to be linearisable under first linearisation conditions.
- the third proportion of second immobilised primers that are cleavable are further configured to be linearisable under second linearisation conditions.
- the first linearisation conditions and the second linearisation conditions are the same or are different.
- the first linearisation conditions and the second linearisation conditions are the same.
- each first immobilised primer comprises a sequence as defined in SEQ ID NO. 1 or 5, or a variant or fragment thereof; and each second immobilised primer comprises a sequence as defined in SEQ ID NO. 2, or a variant or fragment thereof; or wherein each first immobilised primer comprises a sequence as defined in SEQ ID NO. 2, or a variant or fragment thereof; and each second immobilised primer comprises a sequence as defined in SEQ ID NO. 1 or 5, or a variant or fragment thereof.
- a solid support comprising: a plurality of first immobilised primers, and a plurality of second immobilised primers, wherein the plurality of first immobilised primers are located in a first region, wherein the plurality of second immobilised primers are located in a second region surrounding the first region.
- the second region is contiguous.
- the second region is an annular region.
- a concentration of first immobilised primers in the first region is greater than a concentration of second immobilised primers in the second region.
- a ratio between the concentration of first immobilised primers in the first region and the concentration of second immobilised primers in the second region is between 1.25: 1 to 5: 1.
- the ratio between the concentration of first immobilised primers in the first region and the concentration of second immobilised primers in the second region is between 1.5 : 1 to 3 : 1.
- the ratio between the concentration of first immobilised primers in the first region and the concentration of second immobilised primers in the second region is about 2: 1. In one aspect, a concentration of first immobilised primers in the first region is less than a concentration of second immobilised primers in the second region.
- a ratio between the concentration of first immobilised primers in the first region and the concentration of second immobilised primers in the second region is between 1: 1.25 to 1:5.
- the ratio between the concentration of first immobilised primers in the first region and the concentration of second immobilised primers in the second region is between 1 : 1.5 to 1:3.
- the ratio between the concentration of first immobilised primers in the first region and the concentration of second immobilised primers in the second region is about 1:2.
- a concentration of first immobilised primers in the first region is equal or substantially equal to a concentration of second immobilised primers in the second region.
- the solid support is a flow cell.
- kits comprising a solid support as described herein.
- a process of manufacturing a solid support comprising:
- steps (b) immobilising a plurality of second precursor primers onto the solid support to form a plurality of second immobilised primers, wherein a third proportion of the second precursor primers are configured to be cleavable under second cleavage conditions and are blocked, and wherein a fourth proportion of the second precursor primers are configured to be non-cleavable under second cleavage conditions and are unblocked.
- steps (a) and (b) are conducted sequentially or simultaneously.
- step (b) is conducted after step (a).
- step (a) is conducted after step (b).
- steps (a) and (b) are conducted simultaneously.
- immobilisation comprises forming covalent linkages between the solid support and each of the plurality of first precursor primers, and between the solid support and each of the plurality of second precursor primers.
- forming covalent linkages involves using a click reaction.
- forming covalent linkages involves forming a 1,2, 3 -triazole linkage.
- a method of preparing polynucleotide sequences for identification comprising: providing a solid support as described herein, and synthesising a plurality of template sequences that extend from the second immobilised primers in the fourth proportion and a plurality of template complement sequences that extend from the first immobilised primers in the first proportion.
- the step of synthesising the plurality of template sequences extending from the second immobilised primers in the fourth proportion and the plurality of template complement sequences extending from the first immobilised primers in the first proportion involves conducting amplification.
- amplification is bridge amplification.
- amplification is conducted over 20 to 40 cycles.
- amplification is conducted over 25 to 35 cycles.
- the method further comprises a step of removing second immobilised primers in the fourth proportion that are not yet extended, and removing first immobilised primers in the first proportion that are not yet extended.
- the step of removing second immobilised primers in the fourth proportion that are not yet extended and removing first immobilised primers in the first proportion that are not yet extended is conducted using a 3 ’-5’ exonuclease (e.g. Exol).
- a 3 ’-5’ exonuclease e.g. Exol
- the method further comprises a step of unblocking the second proportion of first immobilised primers and unblocking the third proportion of second immobilised primers.
- the method further comprises a step of synthesising a plurality of template sequences that extend from the second immobilised primers in the third proportion and a plurality of template complement sequences that extend from the first immobilised primers in the second proportion.
- the step of synthesising the plurality of template sequences extending from the second immobilised primers in the third proportion and the plurality of template complement sequences extending from the first immobilised primers in the second proportion involves conducting amplification.
- amplification is bridge amplification.
- amplification is conducted over 5 to 25 cycles.
- amplification is conducted over 10 to 20 cycles.
- the number of amplification cycles used for synthesising the plurality of template sequences extending from the second immobilised primers in the third proportion and the plurality of template complement sequences extending from the first immobilised primers in the second proportion is less than the number of amplification cycles used for synthesising the plurality of template sequences extending from the second immobilised primers in the fourth proportion and the plurality of template complement sequences extending from the first immobilised primers in the first proportion.
- the number of amplification cycles used for synthesising the plurality of template sequences extending from the second immobilised primers in the third proportion and the plurality of template complement sequences extending from the first immobilised primers in the second proportion is more than the number of amplification cycles used for synthesising the plurality of template sequences extending from the second immobilised primers in the fourth proportion and the plurality of template complement sequences extending from the first immobilised primers in the first proportion.
- the number of amplification cycles used for synthesising the plurality of template sequences extending from the second immobilised primers in the third proportion and the plurality of template complement sequences extending from the first immobilised primers in the second proportion is the same as the number of amplification cycles used for synthesising the plurality of template sequences extending from the second immobilised primers in the fourth proportion and the plurality of template complement sequences extending from the first immobilised primers in the first proportion.
- the method further comprises a step of removing second immobilised primers in the third proportion that are not yet extended, and removing first immobilised primers in the second proportion that are not yet extended.
- the step of removing second immobilised primers in the third proportion that are not yet extended and removing first immobilised primers in the second proportion that are not yet extended is conducted using a 3 ’-5’ exonuclease (e.g. Exol).
- a 3 ’-5’ exonuclease e.g. Exol
- the method further comprises a step of exposing the solid support to first cleavage conditions and/or second cleavage conditions.
- the first cleavage conditions and/or second cleavage conditions comprise exposure to a thermal trigger, a light trigger, and/or a chemical/biochemical trigger.
- the solid support is exposed to a metal catalyst.
- the solid support is exposed to a transition metal catalyst.
- the solid support is exposed to a palladium-based or a nickel-based catalyst.
- the solid support is exposed to a palladium-based catalyst.
- the solid support is exposed to a glycosylase.
- the method further comprises a step of linearising the plurality of template sequences extending from the second immobilised primers of the fourth proportion and linearising the plurality of template complement sequences extending from the first immobilised primers of the second proportion.
- the method further comprises treating the linearised template sequences and the linearised template complement sequences with a single-stranded binding protein.
- a method of sequencing polynucleotide sequences comprising: preparing polynucleotide sequences for identification using a method as described herein; and concurrently sequencing nucleobases in the template sequences extending from the second immobilised primers of the fourth proportion and the template complement sequences extending from the first immobilised primers of the second proportion.
- the step of concurrently sequencing nucleobases comprises performing sequencing- by-synthesis or sequencing-by-ligation.
- the step of concurrently sequencing nucleobases comprises treatment with a strand displacement polymerase (e.g. phi29).
- a strand displacement polymerase e.g. phi29
- the method further comprises a step of conducting paired-end reads.
- the step of concurrently sequencing nucleobases comprises:
- first intensity data comprising a combined intensity of a first signal component obtained based upon a respective first nucleobase at a first portion of the template sequence and a second signal component obtained based upon a respective second nucleobase at a second portion of the template complement sequence, wherein the first and second signal components are obtained simultaneously;
- selecting the classification based on the first and second intensity data comprises selecting the classification based on the combined intensity of the first and second signal components and the combined intensity of the third and fourth signal components.
- the plurality of classifications comprises sixteen classifications, each classification representing one of sixteen unique combinations of first and second nucleobases.
- the first signal component, second signal component, third signal component and fourth signal component are generated based on light emissions associated with the respective nucleobase.
- the light emissions are detected by a sensor, wherein the sensor is configured to provide a single output based upon the first and second signals.
- the senor comprises a single sensing element.
- the method further comprises repeating steps (a) to (d) for each of a plurality of base calling cycles.
- the step of concurrently sequencing nucleobases comprises:
- first intensity data comprising a combined intensity of a first signal component obtained based upon a respective first nucleobase at a first portion of the template sequence and a second signal component obtained based upon a respective second nucleobase at the second portion of the template complement sequence, wherein the first and second signal components are obtained simultaneously;
- each classification of the plurality of classifications represents one or more possible combinations of respective first and second nucleobases, and wherein at least one classification of the plurality of classifications represents more than one possible combination of respective first and second nucleobases;
- selecting the classification based on the first and second intensity data comprises selecting the classification based on the combined intensity of the first and second signal components and the combined intensity of the third and fourth signal components.
- an intensity of the first signal component is substantially the same as an intensity of the second signal component and an intensity of the third signal component is substantially the same as an intensity of the fourth signal component.
- the plurality of classifications consists of a predetermined number of classifications.
- the plurality of classifications comprises: one or more classifications representing matching first and second nucleobases; and one or more classifications representing mismatching first and second nucleobases
- determining sequence information of the first portion and second portion comprises: in response to selecting a classification representing matching first and second nucleobases, determining a match between the first and second nucleobases; or in response to selecting a classification representing mismatching first and second nucleobases, determining a mismatch between the first and second nucleobases.
- determining sequence information of the first portion and the second portion comprises, in response to selecting a classification representing a match between the first and second nucleobases, base calling the first and second nucleobases. In one aspect, determining sequence information of the first portion and the second portion comprises, based on the selected classification, determining that the second portion is modified relative to the first portion at a location associated with the first and second nucleobases.
- the first signal component, second signal component, third signal component and fourth signal component are generated based on light emissions associated with the respective nucleobase.
- the light emissions are detected by a sensor, wherein the sensor is configured to provide a single output based upon the first and second signals.
- the senor comprises a single sensing element.
- the method further comprises repeating steps (a) to (d) for each of a plurality of base calling cycles.
- the first portion is at least 25 base pairs and the second portion is at least 25 base pairs.
- kits comprising instructions for preparing polynucleotide sequences for identification as described herein; and/or sequencing polynucleotide sequences as described herein.
- a data processing device comprising means for carrying out a method as described herein.
- the data processing device is a polynucleotide sequencer.
- a computer program product comprising instructions which, when the program is executed by a processor, cause the processor to carry out a method as described herein.
- a computer-readable storage medium comprising instructions which, when executed by a processor, cause the processor to carry out a method as described herein.
- a computer-readable data carrier having stored thereon a computer program product as described herein.
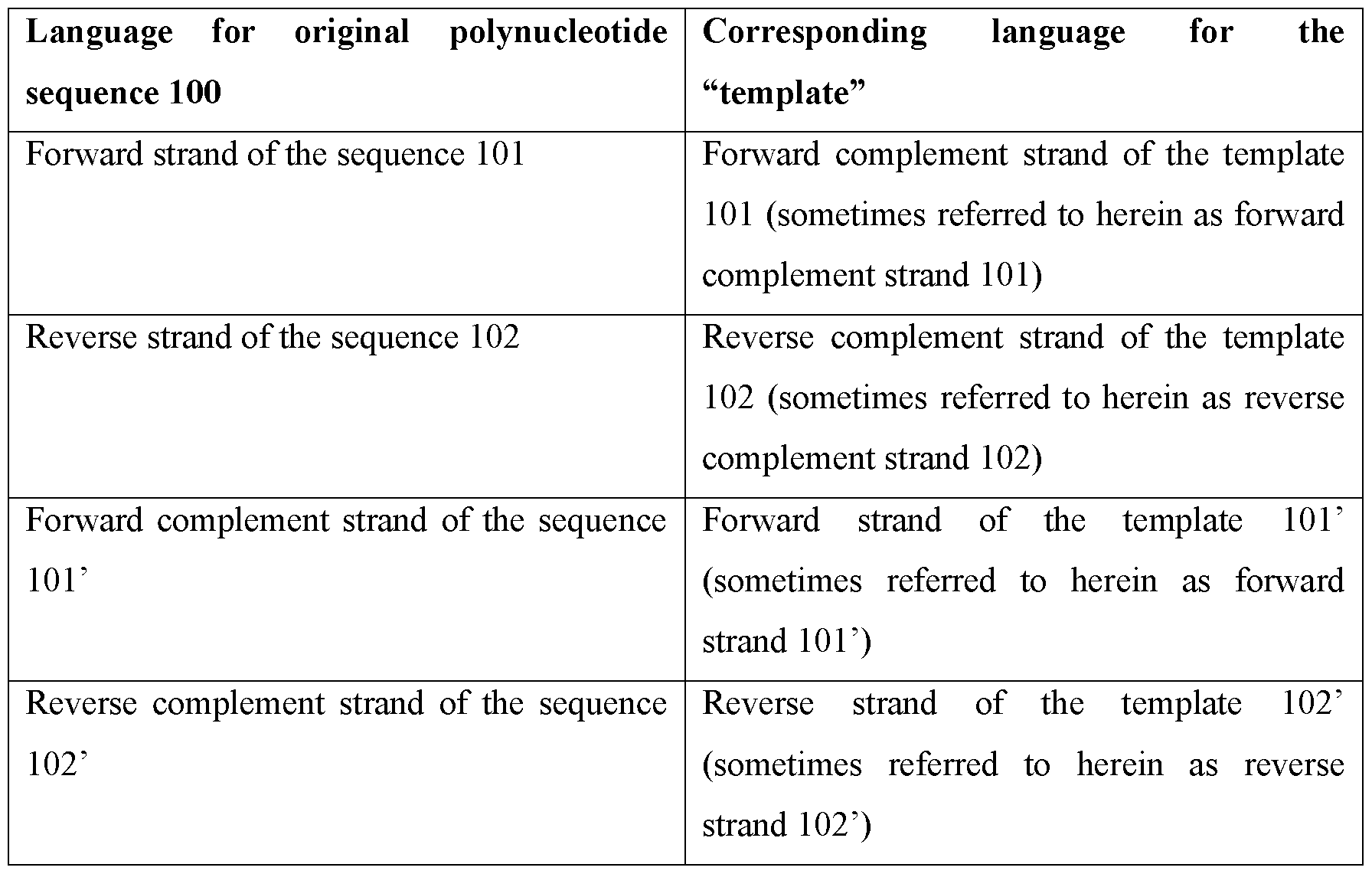
- Figure 1 shows a forward strand, reverse strand, forward complement strand, and reverse complement strand of a polynucleotide molecule.
- Figure 2 shows an example of a polynucleotide sequence (or insert) with 5’ and 3’ adaptor sequences.
- Figure 3 shows a typical polynucleotide with 5’ and 3’ adaptor sequences.
- Figure 4 shows an example of PCR stitching process.
- two sequences - a strand of a human library and a strand of a phiX library are joined together to create a single polynucleotide strand comprising both a first insert (comprising the strand of the human sequence) and a second insert (comprising the strand of the phiX sequence), as well as terminal and internal adaptor sequences.
- Figure 5 shows the preparation of a polynucleotide sequence using a loop fork method.
- Figure 6 shows a typical solid support with bound oligonucleotides.
- Figure 7 shows the stages of bridge amplification and the generation of an amplified cluster comprising (Panel A) a library strand hybridising to an immobilised primer; (Panel B) generation of a template strand from the library strand; (Panel C) dehybridisation and washing away the library strand; (Panel D) hybridisation of the template strand to another immobilised primer; (Panel E) generation of a template complement strand from the template strand via bridge amplification; (Panel F) dehybridisation of the sequence bridge; (Panel G) hybridisation of the template strand and template complement strand to immobilised primers; and (Panel H) subsequent bridge amplification to provide a plurality of template and template complement strands.
- Figure 8 is a set of charts showing the detection of nucleobases using 4-channel, 2-channel and 1 -channel chemistry.
- Figure 9 is a plot showing two-dimensional graphical representations of sixteen distributions of signals generated by polynucleotide sequences according to one embodiment.
- Figure 10 is a flow diagram showing a method for base calling according to one embodiment.
- Figure 11 is a plot showing graphical representations of nine distributions of signals generated by polynucleotide sequences according to one embodiment.
- Figure 12 is a plot showing graphical representations of nine distributions of signals generated by polynucleotide sequences according to one embodiment, highlighting distributions that may be associated with library preparation errors.
- Figure 13 is a flow diagram showing a method for determining sequence information according to one embodiment.
- Figure 14 shows a process for preparing solid supports having central first regions and respective surrounding second regions.
- the first step involves grafting a flow cell with linearisable P5 lawn primers (i.e. a type of cleavable and unblocked first immobilised primer of the first proportion as described herein), non-linearisable P7 lawn primers (i.e. a type of non-cleavable and unblocked second immobilised primer of the fourth proportion as described herein), blocked non-linearisable P5 lawn primers (i.e. a type of non-cleavable and blocked first immobilised primer of the second proportion as described herein), and blocked linearisable P7 lawn primers (i.e.
- linearisable P5 lawn primers i.e. a type of cleavable and unblocked first immobilised primer of the first proportion as described herein
- non-linearisable P7 lawn primers i.e. a type of non-cleavable and unblocked second immobilised
- the second step involves annealing a library strand (not shown) to a particular site, and generating template and template complement strands on the flow cell using amplification. As amplification cycles are conducted, template and template complement strands are generated in a radial direction (with the initial annealing point of the library strand as its centre), thus forming a “central” region.
- the third step involves removal of unused linearisable P5 lawn primers and unused non-linearisable P7 lawn primers using Exol.
- the fourth step involves unblocking of non-linearisable P5 lawn primers and linearisable P7 lawn primers.
- the fifth step involves generating further template and template complement strands on the flow cell using amplification.
- template and template complement strands are again generated in a radial direction (i.e. towards the right on the figure for step 5), thus forming a surrounding ringed region around the “central” region. Since the “central” region is already saturated with template and template complement strands, further strands are generally formed in the surrounding region rather than within the “central” region.
- the central region is predominantly formed of template and template complement strands extending from linearisable P5 lawn primers and non-linearisable P7 lawn primers
- the surrounding ringed region is predominantly formed of template and template complement strands extending from non-linearisable P5 lawn primers and linearisable P7 lawn primers.
- the hairpin loops can be substituted with other blocking groups such as phosphate groups (see Examples).
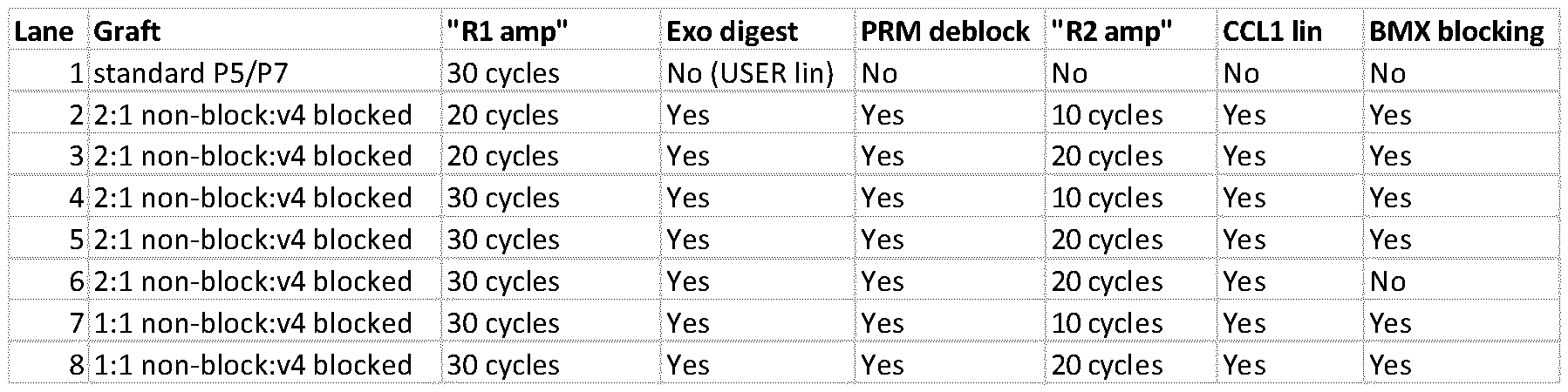
- Figure 15 shows images from solid supports having central first regions and respective surrounding second regions. Different ratios of unblocked to blocked lawn primers were used (e.g. 2: 1 unblocked vs. blocked lawn primers, or 1: 1 unblocked vs. blocked lawn primers).
- the number of amplification cycles is indicated by the notation [m+n], where “m” indicates the number of amplification cycles used for generating the central first region, whereas “n” indicates the number of amplification cycles used for generating the surrounding second regions.
- 30 amplification cycles for the central first region gave good intensity centres, whilst the brightness of the ringed regions could be tuned (e.g. using 10 cycles for less intense rings, to 20 cycles for more intense rings).
- variant refers to a variant polypeptide sequence or part of the polypeptide sequence that retains desired function of the full non-variant sequence.
- a desired function of the immobilised primer retains the ability to bind (i.e. hybridise) to a target sequence.
- a “variant” has at least 25%, 26%, 27%, 28%, 29%, 30%,
- sequence identity of a variant can be determined using any number of sequence alignment programs known in the art.
- fragment refers to a functionally active series of consecutive nucleic acids from a longer nucleic acid sequence.
- the fragment may be at least 99%, at least 95%, at least 90%, at least 80%, at least 70%, at least 60%, at least 50%, at least 40% or at least 30% the length of the longer nucleic acid sequence.
- a fragment as used herein also retains the ability to bind (i.e. hybridise) to a target sequence.
- Sequencing generally comprises four fundamental steps: 1) library preparation to form a plurality of target polynucleotides for identification; 2) cluster generation to form an array of amplified template polynucleotides; 3) sequencing the cluster array of amplified template polynucleotides; and 4) data analysis to identify characteristics of the target polynucleotides from the amplified template polynucleotide sequences. These steps are described in greater detail below. Library strands and template terminology
- the polynucleotide sequence 100 comprises a forward strand of the sequence 101 and a reverse strand of the sequence 102.
- replication of the polynucleotide sequence 100 provides a double-stranded polynucleotide sequence 100a that comprises a forward strand of the sequence 101 and a forward complement strand of the sequence 101’, and a double-stranded polynucleotide sequence 100b that comprises a reverse strand of the sequence 102 and a reverse complement strand of the sequence 102’.
- the term “template” may be used to describe a complementary version of the double-stranded polynucleotide sequence 100.
- the “template” comprises a forward complement strand of the sequence 101’ and a reverse complement strand of the sequence 102’.
- a sequencing process e.g. a sequencing-by-synthesis or a sequencing-by-ligation process
- a sequencing process e.g. a sequencing-by-synthesis or a sequencing-by-ligation process
- the two strands in the template may also be referred to as a forward strand of the template 101 ’ and a reverse strand of the template 102’.
- the complement of the forward strand of the template 101 ’ is termed the forward complement strand of the template 101
- the complement of the reverse strand of the template 102’ is termed the reverse complement strand of the template 102.
- forward strand, reverse strand, forward complement strand, and reverse complement strand are used herein without qualifying whether they are with respect to the original polynucleotide sequence 100 or with respect to the “template”, these terms may be interpreted as referring to the “template”.
- Library preparation is the first step in any high-throughput sequencing platform. These libraries allow templates to be generated via complementary base pairing that can subsequently be clustered and amplified. During library preparation, nucleic acid sequences, for example genomic DNA sample, or cDNA or RNA sample, is converted into a sequencing library, which can then be sequenced.
- the first step in library preparation is random fragmentation of the DNA sample. Sample DNA is first fragmented and the fragments of a specific size (typically 200-500 bp, but can be larger) are ligated, sub-cloned or “inserted” inbetween two oligo adaptors (adaptor sequences). The original sample DNA fragments are referred to as “inserts”.
- the target polynucleotides may advantageously also be size-fractionated prior to modification with the adaptor sequences.
- the templates to be generated typically include separate polynucleotide sequences, in particular a first polynucleotide sequence comprising a first portion and a second polynucleotide sequence comprising a second portion.
- Generating these templates from particular libraries may be performed according to methods known to persons of skill in the art. However, some example approaches of preparing libraries suitable for generation of such templates are described below.
- the library may be prepared by ligating adaptor sequences to doublestranded polynucleotide sequences, each comprising a forward strand of the sequence and a reverse strand of the sequence, as described in more detail in e.g. WO 07/052006, which is incorporated herein by reference.
- “tagmentation” can be used to attach the sample DNA to the adaptors, as described in more detail in e.g. WO 10/048605, US 2012/0301925, US 2013/0143774 and WO 2016/189331, each of which are incorporated herein by reference.
- tagmentation double-stranded DNA is simultaneously fragmented and tagged with adaptor sequences and PCR primer binding sites.
- the combined reaction eliminates the need for a separate mechanical shearing step during library preparation.
- These procedures may be used, for example, for preparing templates including a first polynucleotide sequence comprising a first portion and a second polynucleotide sequence comprising a second portion, wherein the first portion is a forward strand of the template, and the second portion is a forward complement strand of the template - i.e. a copy of the forward strand (or alternatively, wherein the first portion is a reverse strand of the template, and the second portion is a reverse complement strand of the template).
- library preparation may comprise ligating a first primer-binding sequence 301’ (e.g. P5’, such as SEQ ID NO. 3) and a second terminal sequencing primer binding site 304 (e.g. SBS3’, for example, SEQ ID NO. 8) to a 3’-end of a forward strand of a sequence 101. See Figure 2.
- the library preparation may be arranged such that the second terminal sequencing primer binding site 304 is attached (e.g. directly attached) to the 3 ’-end of the forward strand of the sequence 101, and such that the first primer-binding sequence 301’ is attached (e.g. directly attached) to the 3 ’-end of the second terminal sequencing primer binding site 304.
- the library preparation may further comprise ligating a complement of first terminal sequencing primer binding site 303’ (e.g. SBS12, such as SEQ ID NO. 9) (also referred to herein as a first terminal sequencing primer binding site complement 303’) and a complement of a second primerbinding sequence 302 (also referred to herein as a second primer-binding complement sequence 302) (e.g. P7, such as SEQ ID NO. 2) to a 5 ’-end of the forward strand of the sequence 101.
- the library preparation may be arranged such that first terminal sequencing primer binding site complement 303’ is attached (e.g. directly attached) to the 5 ’-end of the forward strand of the sequence 101, and such that second primer-binding complement sequence 302 is attached (e.g. directly attached) to the 5 ’-end of first terminal sequencing primer binding site complement 303’.
- one strand of a polynucleotide within a polynucleotide library may comprise, in a 5 ’ to 3 ’ direction, a second primer-binding complement sequence 302 (e.g. P7), a first terminal sequencing primer binding site complement 303’ (e.g. SBS12), a forward strand of the sequence 101, a second terminal sequencing primer binding site 304 (e.g. SBS3’), and a first primer-binding sequence 301’ (e.g. P5’) ( Figure 2 - bottom strand).
- a second primer-binding complement sequence 302 e.g. P7
- a first terminal sequencing primer binding site complement 303’ e.g. SBS12
- a forward strand of the sequence 101 e.g. SBS3’
- a second terminal sequencing primer binding site 304 e.g. SBS3’
- a first primer-binding sequence 301’ e.g. P5’
- the strand may further comprise one or more index sequences.
- a first index sequence (e.g. i7) may be provided between the second primer-binding complement sequence 302 (e.g. P7) and the first terminal sequencing primer binding site complement 303’ (e.g. SB SI 2).
- a second index complement sequence (e.g. i5 ’) may be provided between the second terminal sequencing primer binding site 304 (e.g. SBS3’) and the first primer-binding sequence 301’ (e.g. P5’).
- one strand of a polynucleotide within a polynucleotide library may comprise, in a 5’ to 3’ direction, a second primer-binding complement sequence 302 (e.g. P7), a first index sequence (e.g. i7), a first terminal sequencing primer binding site complement 303’ (e.g. SBS12), a forward strand of the sequence 101, a second terminal sequencing primer binding site 304 (e.g. SBS3’), a second index complement sequence (e.g. i5’), and a first primer-binding sequence 301’ (e.g. P5’).
- a typical polynucleotide is shown in Figure 3 (bottom strand).
- the library preparation may also comprise ligating a second primer-binding sequence 302’ (e.g. P7’) and a first terminal sequencing primer binding site 303 (e.g. SBS12’) to a 3’-end of a reverse strand of a sequence 102.
- the library preparation may be arranged such that first terminal sequencing primer binding site 303 is attached (e.g. directly attached) to the 3 ’-end of the reverse strand of the sequence 102, and such that the second primer-binding sequence 302’ is attached (e.g. directly attached) to the 3 ’-end of first terminal sequencing primer binding site 303.
- the library preparation may further comprise ligating a complement of a second terminal sequencing primer binding site 304’ (e.g. SBS3) (also referred to herein as a second terminal sequencing primer binding site complement 304’) and a complement of a first primer-binding sequence 301 (also referred to herein as a first primer-binding complement sequence 301) (e.g. P5) to a 5 ’-end of the reverse strand of the sequence 102.
- the library preparation may be arranged such that the second terminal sequencing primer binding site complement 304’ is attached (e.g. directly attached) to the 5 ’-end of the reverse strand of the sequence 102, and such that the first primer-binding complement sequence 301 is attached (e.g. directly attached) to the 5 ’-end of the second terminal sequencing primer binding site complement 304’.
- another strand of a polynucleotide within a polynucleotide library may comprise, in a 5 ’ to 3’ direction, a first primer-binding complement sequence 301 (e.g. P5), a second terminal sequencing primer binding site complement 304’ (e.g. SBS3), a reverse strand of the sequence 102, a first terminal sequencing primer binding site 303 (e.g. SBS12’), and a second primerbinding sequence 302’ (e.g. P7’) ( Figure 2 - top strand).
- a first primer-binding complement sequence 301 e.g. P5
- a second terminal sequencing primer binding site complement 304 e.g. SBS3
- a reverse strand of the sequence 102 e.g. SBS12’
- a first terminal sequencing primer binding site 303 e.g. SBS12’
- a second primerbinding sequence 302’ e.g. P7’
- the another strand may further comprise one or more index sequences.
- a second index sequence (e.g. i5) may be provided between the first primerbinding complement sequence 301 (e.g. P5) and the second terminal sequencing primer binding site complement 304’ (e.g. SBS3).
- a first index complement sequence (e.g. i7’) may be provided between the first terminal sequencing primer binding site 303 (e.g. SBS12’) and the second primer-binding sequence 302’ (e.g. P7’).
- another strand of a polynucleotide within a polynucleotide library may comprise, in a 5’ to 3’ direction, a first primer-binding complement sequence 301 (e.g. P5), a second index sequence (e.g. i5), a second terminal sequencing primer binding site complement 304’ (e.g. SBS3), a reverse strand of the sequence 102, a first terminal sequencing primer binding site 303 (e.g. SBS12’), a first index complement sequence (e.g. i7’), and a second primer-binding sequence 302’ (e.g. P7’).
- a typical polynucleotide is shown in Figure 3 (top strand).
- the library may be prepared using PCR stitching methods, such as (splicing by) overlap extension PCR (also known as OE-PCR or SOE-PCR), as described in more detail in e.g. Higuchi et al. (Nucleic Acids Res., 1988, vol. 16, pp. 7351-7367), which is incorporated herein by reference.
- PCR stitching methods such as (splicing by) overlap extension PCR (also known as OE-PCR or SOE-PCR), as described in more detail in e.g. Higuchi et al. (Nucleic Acids Res., 1988, vol. 16, pp. 7351-7367), which is incorporated herein by reference.
- a representative process for conducting PCR stitching for a human and PhiX library is shown in Figure 4.
- the term “genetically unrelated” refers to portions which are not related in the sense of being any two of the group consisting of: forward strands, reverse strands, forward complement strands, and reverse complement strands.
- the “genetically unrelated” sequences could be different fragment sequences which are derived from the same source, but are different fragments from that source (e.g. from the same fragmented library preparation process). This includes sequences that can be overlapping in sequence (but not identical in sequence).
- further processes may be used to generate templates including a first polynucleotide sequence comprising a first portion and a second polynucleotide sequence comprising a second portion, wherein the first portion and the second portion are genetically unrelated.
- the library may be prepared using a loop fork method, which is described below. This procedure may be used, for example, for preparing templates including a first polynucleotide sequence comprising a first portion and a second polynucleotide sequence comprising a second portion, wherein the first portion is a forward strand of the template, and the second portion is a reverse complement strand of the template (or alternatively, wherein the first portion is a reverse strand of the template, and the second portion is a forward complement strand of the template).
- Such libraries may also be referred to as self-tandem inserts.
- a representative process for conducting a loop fork method is shown in Figure 5.
- adaptors may be ligated to a first end of the sequence (e.g. using processes as described in more detail in e.g. WO 07/052006, or “tagmentation” methods as described above).
- a second end of the sequence (different from the first end) may be ligated to a loop, which connects the forward strand of the sequence and the reverse strand of the sequence, thus generating a loop fork ligated polynucleotide sequence.
- templates including a first polynucleotide sequence comprising a first portion and a second polynucleotide sequence comprising a second portion, wherein the first portion is a forward strand of the template, and the second portion is a reverse complement strand of the template (or alternatively, wherein the first portion is a reverse strand of the template, and the second portion is a forward complement strand of the template).
- a double-stranded nucleic acid will typically be formed from two complementary polynucleotide strands comprised of deoxyribonucleotides or ribonucleotides joined by phosphodiester bonds, but may additionally include one or more ribonucleotides and/or non-nucleotide chemical moieties and/or non-naturally occurring nucleotides and/or non-naturally occurring backbone linkages.
- the double-stranded nucleic acid may include non-nucleotide chemical moieties, e.g. linkers or spacers, at the 5' end of one or both strands.
- the double-stranded nucleic acid may include methylated nucleotides, uracil bases, phosphorothioate groups, peptide conjugates etc.
- Such non-DNA or non-natural modifications may be included in order to confer some desirable property to the nucleic acid, for example to enable covalent, non-covalent or metal-coordination attachment to a solid support, or to act as spacers to position the site of cleavage an optimal distance from the solid support.
- a single stranded nucleic acid consists of one such polynucleotide strand.
- a polynucleotide strand is only partially hybridised to a complementary strand - for example, a long polynucleotide strand hybridised to a short nucleotide primer - it may still be referred to herein as a single stranded nucleic acid.
- a sequence comprising at least a primer-binding sequence (a primer-binding sequence and a sequencing primer binding site, in another aspect, a combination of a primer-binding sequence, an index sequence and a sequencing primer binding site) may be referred to herein as an adaptor sequence, and an insert is flanked by a 5’ adaptor sequence and a 3’ adaptor sequence.
- the primerbinding sequence may also comprise a sequencing primer for the index read.
- an “adaptor” refers to a sequence that comprises a short sequence-specific oligonucleotide that is ligated to the 5' and 3' ends of each DNA (or RNA) fragment in a sequencing library as part of library preparation.
- the adaptor sequence may further comprise nonpeptide linkers.
- the P5’ and P7’ primer-binding sequences are complementary to short primer sequences (or lawn primers) present on the surface of a flow cell. Binding of P5’ and P7’ to their complements (P5 and P7) on - for example - the surface of the flow cell, permits nucleic acid amplification. As used herein denotes the complementary strand.
- the primer-binding sequences in the adaptor which permit hybridisation to amplification primers will typically be around 20-40 nucleotides in length, although the invention is not limited to sequences of this length.
- the precise identity of the amplification primers (e.g. lawn primers), and hence the cognate sequences in the adaptors, are generally not material to the invention, as long as the primer-binding sequences are able to interact with the amplification primers in order to direct PCR amplification.
- sequence of the amplification primers may be specific for a particular target nucleic acid that it is desired to amplify, but in other embodiments these sequences may be "universal" primer sequences which enable amplification of any target nucleic acid of known or unknown sequence which has been modified to enable amplification with the universal primers.
- the criteria for design of PCR primers are generally well known to those of ordinary skill in the art.
- the index sequences are unique short DNA (or RNA) sequences that are added to each DNA (or RNA) fragment during library preparation.
- the unique sequences allow many libraries to be pooled together and sequenced simultaneously. Sequencing reads from pooled libraries are identified and sorted computationally, based on their barcodes, before final data analysis. Library multiplexing is also a useful technique when working with small genomes or targeting genomic regions of interest. Multiplexing with barcodes can exponentially increase the number of samples analysed in a single run, without drastically increasing run cost or run time. Examples of tag sequences are found in WO05/068656, whose contents are incorporated herein by reference in their entirety.
- the tag can be read at the end of the first read, or equally at the end of the second read, for example using a sequencing primer complementary to the strand marked P7.
- the invention is not limited by the number of reads per cluster, for example two reads per cluster: three or more reads per cluster are obtainable simply by dehybridising a first extended sequencing primer, and rehybridising a second primer before or after a cluster repopulation/strand resynthesis step. Methods of preparing suitable samples for indexing are described in, for example WO 2008/093098, which is incorporated herein by reference. Single or dual indexing may also be used. With single indexing, up to 48 unique 6-base indexes can be used to generate up to 48 uniquely tagged libraries.
- up to 24 unique 8-base Index 1 sequences and up to 16 unique 8-base Index 2 sequences can be used in combination to generate up to 384 uniquely tagged libraries. Pairs of indexes can also be used such that every i5 index and every i7 index are used only one time. With these unique dual indexes, it is possible to identify and filter indexed hopped reads, providing even higher confidence in multiplexed samples.
- the sequencing primer binding sites are sequencing and/or index primer binding sites and indicate the starting point of the sequencing read.
- a sequencing primer anneals (i.e. hybridises) to at least a portion of the sequencing primer binding site on the template strand.
- the polymerase enzyme binds to this site and incorporates complementary nucleotides base by base into the growing opposite strand.
- a double stranded nucleic acid library is formed, typically, the library has previously been subjected to denaturing conditions to provide single stranded nucleic acids. Suitable denaturing conditions will be apparent to the skilled reader with reference to standard molecular biology protocols (Sambrook et al., 2001, Molecular Cloning, A Laboratory Manual, 4th Ed, Cold Spring Harbor Laboratory Press, Cold Spring Harbor Laboratory Press, NY; Current Protocols, eds Ausubel et al). In one embodiment, chemical denaturation may be used.
- a single-stranded library may be contacted in free solution onto a solid support comprising surface capture moieties (for example P5 and P7 lawn primers).
- a solid support 200 such as a flowcell.
- seeding and clustering can be conducted off- flowcell using other types of solid support.
- the solid support 200 may comprise a substrate 204. See Figure 6.
- the substrate 204 comprises at least one well 203 (e.g. a nanowell), and typically comprises a plurality of wells 203 (e.g. a plurality of nanowells).
- the solid support comprises a plurality of first immobilised primers and a plurality of second immobilised primers.
- each well 203 may comprise a plurality of first immobilised primers 201.
- each well 203 may comprise a plurality of second immobilised primers 202.
- each well 203 may comprise a plurality of first immobilised primers 201 and a plurality of second immobilised primers 202.
- the first immobilised primer 201 may be attached via a 5 ’-end of its polynucleotide chain to the solid support 200.
- the extension may be in a direction away from the solid support 200.
- the second immobilised primer 202 may be attached via a 5 ’-end of its polynucleotide chain to the solid support 200.
- the extension may be in a direction away from the solid support 200.
- the first immobilised primer 201 may be different to the second immobilised primer 202 and/or a complement of the second immobilised primer 202.
- the second immobilised primer 202 may be different to the first immobilised primer 201 and/or a complement of the first immobilised primer 201.
- the (or each of the) first immobilised primer(s) 201 may comprise a sequence as defined in SEQ ID NO. 1 or 5, or a variant or fragment thereof.
- the (or each of the) second immobilised primer(s) 202 may comprise a sequence as defined in SEQ ID NO. 2, or a variant or fragment thereof. Whilst first immobilised primer(s) 201 are shown here to correspond to P5 and second immobilised primer(s) 202 are shown here to correspond to P7, the definitions of these may be swapped - in other words, first immobilised primer(s) 201 may correspond instead to P7, and second immobilised primer(s) 202 may correspond to P5.
- the solid support may be contacted with the template to be amplified under conditions which permit hybridisation (or annealing - such terms may be used interchangeably) between the template and the immobilised primers.
- the template is usually added in free solution under suitable hybridisation conditions, which will be apparent to the skilled reader.
- hybridisation conditions are, for example, 5xSSC at 40°C.
- other temperatures may be used during hybridisation, for example about 50°C to about 75°C, about 55°C to about 70°C, or about 60°C to about 65°C. Solid-phase amplification can then proceed.
- the first step of the amplification is a primer extension step in which nucleotides are added to the 3' end of the immobilised primer using the template to produce a fully extended complementary strand.
- the template is then typically washed off the solid support.
- the complementary strand will include at its 3' end a primer-binding sequence (i.e. either P5’ or P7’) which is capable of bridging to the second primer molecule immobilised on the solid support and binding.
- Further rounds of amplification leads to the formation of clusters or colonies of template molecules bound to the solid support. This is called clustering.
- amplification may be isothermal amplification using a strand displacement polymerase; or may be exclusion amplification as described in WO 2013/188582. Further information on amplification can be found in WO 02/06456 and WO 07/107710, the contents of which are incorporated herein in their entirety by reference.
- a cluster of template molecules comprising copies of a template strand and copies of the complement of the template strand.
- each first polynucleotide sequence may be attached (via the 5 ’-end of the first polynucleotide sequence) to a first immobilised primer, and wherein each second polynucleotide sequence is attached (via the 5 ’-end of the second polynucleotide sequence) to a second immobilised primer.
- Each first polynucleotide sequence may comprise a second adaptor sequence, wherein the second adaptor sequence comprises a portion, which is substantially complementary to the second immobilised primer (or is substantially complementary to the second immobilised primer).
- the second adaptor sequence may be at a 3 ’-end of the first polynucleotide sequence.
- Each second polynucleotide sequence may comprise a first adaptor sequence, wherein the first adaptor sequence comprises a portion, which is substantially complementary to the first immobilised primer (or is substantially complementary to the first immobilised primer).
- the first adaptor sequence may be at a 3 ’-end of the second polynucleotide sequence.
- a solution comprising a polynucleotide library prepared by ligating adaptor sequences to double-stranded polynucleotide sequences as described above may be flown across a flowcell.
- a particular polynucleotide strand from the polynucleotide library to be sequenced comprising, in a 5’ to 3’ direction, a second primer-binding complement sequence 302 (e.g. P7), a first terminal binding site complement 303’ (e.g. SBS12), a forward strand of the sequence 101, a second terminal sequencing primer binding site 304 (e.g. SBS3’) and a first primer-binding sequence 301’ (e.g. P5’), may anneal (via the first primer-binding sequence 301’) to the first immobilised primer 201 (e.g. P5 lawn primer) located within a particular well 203 ( Figure 7A).
- a second primer-binding complement sequence 302 e.g. P7
- a first terminal binding site complement 303’ e.g. SBS12
- a forward strand of the sequence 101 e.g. SBS3’
- a first primer-binding sequence 301’ e.g.
- the polynucleotide library may comprise other polynucleotide strands with different forward strands of the sequence 101.
- Such other polynucleotide strands may anneal to corresponding first immobilised primers 201 (e.g. P5 lawn primers) in different wells 203, thus enabling parallel processing of the various different strands within the polynucleotide library.
- first immobilised primers 201 e.g. P5 lawn primers
- a new polynucleotide strand may then be synthesised, extending from the first immobilised primer 201 (e.g. P5 lawn primer) in a direction away from the substrate 204.
- this generates a template strand comprising, in a 5 ’ to 3 ’ direction, the first immobilised primer 201 (e.g. P5 lawn primer) which is attached to the solid support 200, a second terminal sequencing primer binding site complement 304’ (e.g. SBS3), a forward strand of the template 101’ (which represents a type of “first portion”), a first terminal sequencing primer binding site 303 (which represents atype of “first sequencing primer binding site”) (e.g. SBS12’), and a second primer-binding sequence 302’ (e.g. P7’) ( Figure 7B).
- Such a process may utilise an appropriate polymerase, such as a DNA or RNA polymerase.
- the polynucleotides in the library comprise index sequences, then corresponding index sequences are also produced in the template.
- the polynucleotide strand from the polynucleotide library may then be dehybridised and washed away, leaving a template strand attached to the first immobilised primer 201 (e.g. P5 lawn primer) ( Figure 7C).
- first immobilised primer 201 e.g. P5 lawn primer
- the second primer-binding sequence 302’ (e.g. P7’) on the template strand may then anneal to a second immobilised primer 202 (e.g. P7 lawn primer) located within the well 203. This forms a “bridge” ( Figure 7D).
- a second immobilised primer 202 e.g. P7 lawn primer
- a new polynucleotide strand may then be synthesised by bridge amplification, extending from the second immobilised primer 202 (e.g. P7 lawn primer) (initially) in a direction away from the substrate 204.
- the second immobilised primer 202 e.g. P7 lawn primer
- a first terminal sequencing primer binding site complement 303’ e.g. SBS12
- a forward complement strand of the template 101 which represents a type of “second portion”
- a second terminal sequencing primer binding site 304 which represents a type of “second sequencing primer binding site”
- a first primer-binding sequence 301’ e.g. P5’
- Figure 7E may utilise a suitable polymerase, such as a DNA or RNA polymerase.
- the strand attached to the second immobilised primer 202 may then be dehybridised from the strand attached to the first immobilised primer 201 (e.g. P5 lawn primer) ( Figure 7F).
- a subsequent bridge amplification cycle can then lead to amplification of the strand attached to the first immobilised primer 201 (e.g. P5 lawn primer) and the strand attached to the second immobilised primer 202 (e.g. P7 lawn primer).
- the second primer-binding sequence 302’ e.g. P7’
- the first primer-binding sequence 301’ e.g. P5’
- the second immobilised primer 202 e.g. P7 lawn primer
- the first primer-binding sequence 301’ on the template strand attached to the second immobilised primer 202 (e.g. P7 lawn primer) may then anneal to another first immobilised primer 201 (e.g. P5 lawn primer) located within the well 203 ( Figure 7G).
- Completion of bridge amplification and dehybridisation may then provide an amplified (duoclonal) cluster, thus providing a plurality of first polynucleotide sequences comprising the forward strand of the template 101’ (i.e. “first portions”), and a plurality of second polynucleotide sequences comprising the forward complement strand of the template 101 (i.e. “second portions”) (Figure 7H).
- further bridge amplification cycles may be conducted to increase the number of first polynucleotide sequences and second polynucleotide sequences within the well 203.
- the “first portion” corresponds with the forward strand of the template 101’
- the “second portion” corresponds with the forward complement strand of the template 101.
- a portion at or close to the loop may be cleaved (e.g. by nicking).
- the loop may comprise a cleavage site (e.g. a restriction recognition site, a cleavable linker, a modified nucleotide, or the like).
- a portion at or close to the overlap region may comprise a cleavage site (e.g. a restriction recognition site, a cleavable linker, a modified nucleotide, or the like).
- a cleavage site e.g. a restriction recognition site, a cleavable linker, a modified nucleotide, or the like.
- first portions and second portions may be prepared for templates including a first polynucleotide sequence comprising a first portion and a second polynucleotide sequence comprising a second portion, and as such the forward strand of the template 101’ and the forward complement strand of the template 101 may be substituted as appropriate.
- the template provides information (e.g. identification of the genetic sequence, identification of epigenetic modifications) on the original target polynucleotide sequence.
- a sequencing process e.g. a sequencing-by-synthesis or sequencing-by- ligation process
- sequencing may be carried out using any suitable "sequencing-by-synthesis" technique, wherein nucleotides are added successively in cycles to the free 3' hydroxyl group, resulting in synthesis of a polynucleotide chain in the 5' to 3' direction.
- the nature of the nucleotide added may be determined after each addition.
- One particular sequencing method relies on the use of modified nucleotides that can act as reversible chain terminators. Such reversible chain terminators comprise removable 3' blocking groups.
- the modified nucleotides may carry a label to facilitate their detection.
- a label may be configured to emit a signal, such as an electromagnetic signal, or a (visible) light signal.
- the label is a fluorescent label (e.g. a dye).
- a fluorescent label e.g. a dye
- the label may be configured to emit an electromagnetic signal, or a (visible) light signal.
- One method for detecting the fluorescently labelled nucleotides comprises using laser light of a wavelength specific for the labelled nucleotides, or the use of other suitable sources of illumination.
- the fluorescence from the label on an incorporated nucleotide may be detected by a CCD camera or other suitable detection means. Suitable detection means are described in PCT/US2007/007991, the contents of which are incorporated herein by reference in their entirety.
- the detectable label need not be a fluorescent label. Any label can be used which allows the detection of the incorporation of the nucleotide into the DNA sequence.
- Each cycle may involve simultaneous delivery of four different nucleotide types to the array of template molecules. Alternatively, different nucleotide types can be added sequentially and an image of the array of template molecules can be obtained between each addition step.
- each nucleotide type may have a (spectrally) distinct label.
- four channels may be used to detect four nucleobases (also known as 4-channel chemistry) ( Figure 8 - left).
- a first nucleotide type e.g. A
- a second nucleotide type e.g. G
- a second label e.g. configured to emit a second wavelength, such as blue light
- a third nucleotide type e.g. T
- a third label e.g.
- a fourth nucleotide type may include a fourth label (e.g. configured to emit a fourth wavelength, such as yellow light).
- Four images can then be obtained, each using a detection channel that is selective for one of the four different labels.
- the first nucleotide type e.g. A
- the second nucleotide type e.g. G
- the second channel e.g. configured to detect the second wavelength, such as blue light
- the third nucleotide type e.g. T
- a third channel e.g.
- the fourth nucleotide type (e.g. C) may be detected in a fourth channel (e.g. configured to detect the fourth wavelength, such as yellow light).
- a fourth channel e.g. configured to detect the fourth wavelength, such as yellow light.
- detection of each nucleotide type may be conducted using fewer than four different labels.
- sequencing-by-synthesis may be performed using methods and systems described in US 2013/0079232, which is incorporated herein by reference.
- two channels may be used to detect four nucleobases (also known as 2-channel chemistry) ( Figure 8 - middle).
- a first nucleotide type e.g. A
- a second label e.g. configured to emit a second wavelength, such as red light
- a second nucleotide type e.g. G
- a third nucleotide type e.g. T
- the first label e.g.
- the first nucleotide type (e.g. A) may be detected in both a first channel (e.g. configured to detect the first wavelength, such as red light) and a second channel (e.g. configured to detect the second wavelength, such as green light), the second nucleotide type (e.g.
- the third nucleotide type (e.g. T) may be detected in the first channel (e.g. configured to detect the first wavelength, such as red light) and may not be detected in the second channel
- the fourth nucleotide type (e.g. C) may not be detected in the first channel and may be detected in the second channel (e.g. configured to detect the second wavelength, such as green light).
- one channel may be used to detect four nucleobases (also known as 1- channel chemistry) ( Figure 8 - right).
- a first nucleotide type e.g. A
- a second nucleotide type e.g. G
- a third nucleotide type e.g. T
- a non-cleavable label e.g. configured to emit the wavelength, such as green light
- a fourth nucleotide type e.g. C
- a label-accepting site which does not include the label.
- a first image can then be obtained, and a subsequent treatment carried out to cleave the label attached to the first nucleotide type, and to attach the label to the label-accepting site on the fourth nucleotide type.
- a second image may then be obtained.
- the first nucleotide type e.g. A
- the second nucleotide type e.g. G
- the third nucleotide type e.g. T
- the channel e.g.
- the fourth nucleotide type (e.g. C) may not be detected in the channel in the first image and may be detected in the channel in the second image (e.g. configured to detect the wavelength, such as green light).
- the sequencing process comprises a first sequencing read and second sequencing read.
- the first sequencing read and the second sequencing read may be conducted concurrently. In other words, the first sequencing read and the second sequencing read may be conducted at the same time.
- the first sequencing read may comprise the binding of a first sequencing primer (also known as a read 1 sequencing primer) to the first sequencing primer binding site (e.g. first terminal sequencing primer binding site 303 in templates including a first polynucleotide sequence comprising a first portion and a second polynucleotide sequence comprising a second portion).
- the second sequencing read may comprise the binding of a second sequencing primer (also known as a read 2 sequencing primer) to the second sequencing primer binding site (e.g. second terminal sequencing primer binding site 304 in templates including a first polynucleotide sequence comprising a first portion and a second polynucleotide sequence comprising a second portion).
- first portion e.g. forward strand of the template 101’ in templates including a first polynucleotide sequence comprising a first portion and a second polynucleotide sequence comprising a second portion; or other types of first portion if different library preparations are used, such as by PCR stitching or loop fork methods
- second portion e.g. forward complement strand of the template 101 in templates including a first polynucleotide sequence comprising a first portion and a second polynucleotide sequence comprising a second portion; or other types of first portion if different library preparations are used, such as by PCR stitching or loop fork methods.
- sequencing by ligation for example as described in US 6,306,597 or WO 06/084132, the contents of which are incorporated herein by reference.
- Figure 9 is a scatter plot showing an example of sixteen distributions of signals generated by polynucleotide sequences disclosed herein.
- the two-dimensional scatter plot of Figure 9 shows sixteen distributions (or bins) of intensity values from the combination of a brighter signal (i.e. a first signal as described herein) and a dimmer signal (i.e. a second signal as described herein); the two signals may be co-localized and may not be optically resolved as described above.
- the intensity values shown in Figure 9 may be up to a scale or normalisation factor; the units of the intensity values may be arbitrary or relative (i.e., representing the ratio of the actual intensity to a reference intensity).
- the sum of the brighter signal generated by the first portions and the dimmer signal generated by the second portions results in a combined signal.
- the combined signal may be captured by a first optical channel and a second optical channel.
- the brighter signal may be A, T, C or G
- the dimmer signal may be A, T, C or G
- the computer system can map the combined signal generated into one of the sixteen bins, and thus determine the added nucleobase at the first portion and the added nucleobase at the second portion, respectively.
- the computer processor base calls both the added nucleobase at the first portion and the added nucleobase at the second portion as C.
- the processor base calls the added nucleobase at the first portion as C and the added nucleobase at the second portion as T.
- the processor base calls the added nucleobase at the first portion as C and the added nucleobase at the second portion as G.
- the processor base calls the added nucleobase at the first portion as C and the added nucleobase at the second portion as A.
- the processor base calls the added nucleobase at the first portion as T and the added nucleobase at the second portion as C.
- the processor base calls both the added nucleobase at the first portion and the added nucleobase at the second portion as T.
- the processor base calls the added nucleobase at the first portion as T and the added nucleobase at the second portion as G.
- the processor base calls the added nucleobase at the first portion as T and the added nucleobase at the second portion as A.
- the processor base calls the added nucleobase at the first portion as G and the added nucleobase at the second portion as C.
- the processor base calls the added nucleobase at the first portion as G and the added nucleobase at the second portion as T.
- the processor base calls both the added nucleobase at the first portion and the added nucleobase at the second portion as G.
- the processor base calls the added nucleobase at the first portion as G and the added nucleobase at the second portion as A.
- the processor base calls the added nucleobase at the first portion as A and the added nucleobase at the second portion as C.
- the processor base calls the added nucleobase at the first portion as A and the added nucleobase at the second portion as T.
- the processor base calls the added nucleobase at the first portion as A and the added nucleobase at the second portion as G.
- the processor base calls both the added nucleobase at the first portion and the added nucleobase at the second portion as A.
- T is configured to emit a signal in both the IMAGE 1 channel and the IMAGE 2 channel
- A is configured to emit a signal in the IMAGE 1 channel only
- C is configured to emit a signal in the IMAGE 2 channel only
- G does not emit a signal in either channel.
- A may be configured to emit a signal in both the IMAGE 1 channel and the IMAGE 2 channel
- T may be configured to emit a signal in the IMAGE 1 channel only
- C may be configured to emit a signal in the IMAGE 2 channel only
- G may be configured to not emit a signal in either channel.
- Figure 10 is a flow diagram showing a method 1700 of base calling according to the present disclosure.
- the described method allows for simultaneous sequencing of two (or more) portions (e.g. the first portion and the second portion) in a single sequencing run from a single combined signal obtained from the first portion and the second portion, thus requiring less sequencing reagent consumption and faster generation of data from both the first portion and the second portion.
- the simplified method may reduce the number of workflow steps while producing the same yield as compared to existing next-generation sequencing methods. Thus, the simplified method may result in reduced sequencing runtime.
- the disclosed method 1700 may start from block 1701. The method may then move to block 1710.
- intensity data is obtained.
- the intensity data includes first intensity data and second intensity data.
- the first intensity data comprises a combined intensity of a first signal component obtained based upon a respective first nucleobase of the first portion and a second signal component obtained based upon a respective second nucleobase of the second portion.
- the second intensity data comprises a combined intensity of a third signal component obtained based upon the respective first nucleobase of the first portion and a fourth signal component obtained based upon the respective second nucleobase of the second portion.
- the first portion is capable of generating a first signal comprising a first signal component and a third signal component.
- the second portion is capable of generating a second signal comprising a second signal component and a fourth signal component.
- the first portion and the second portion may be arranged on the solid support such that signals from the first portion and the second portion are detected by a single sensing portion and/or may comprise a single cluster such that first signals and second signals from each of the respective first portions and second portions cannot be spatially resolved.
- obtaining the intensity data comprises selecting intensity data that corresponds to two (or more) different portions (e.g. the first portion and the second portion).
- intensity data is selected based upon a chastity score.
- a chastity score may be calculated as the ratio of the brightest base intensity divided by the sum of the brightest and second brightest base intensities.
- the desired chastity score may be different depending upon the expected intensity ratio of the light emissions associated with the different portions. As described above, it may be desired to produce clusters comprising the first portion and the second portion, which give rise to signals in a ratio of 2: 1.
- high-quality data corresponding to two portions with an intensity ratio of 2: 1 may have a chastity score of around 0.8 to 0.9.
- the method may proceed to block 1720.
- one of a plurality of classifications is selected based on the intensity data.
- Each classification represents a possible combination of respective first and second nucleobases.
- the plurality of classifications comprises sixteen classifications as shown in Figure 9, each representing a unique combination of first and second nucleobases. Where there are two portions, there are sixteen possible combinations of first and second nucleobases.
- Selecting the classification based on the first and second intensity data comprises selecting the classification based on the combined intensity of the first and second signal components and the combined intensity of the third and fourth signal components.
- the method may then proceed to block 1730, where the respective first and second nucleobases are base called based on the classification selected in block 1720.
- the signals generated during a cycle of a sequencing are indicative of the identity of the nucleobase(s) added during sequencing (e.g. using sequencing-by-synthesis). It will be appreciated that there is a direct correspondence between the identity of the nucleobases that are incorporated and the identity of the complementary base at the corresponding position of the template sequence bound to the solid support. Therefore, any references herein to the base calling of respective nucleobases at the two portions encompasses the base calling of nucleobases hybridised to the template sequences and, alternatively or additionally, the identification of the corresponding nucleobases of the template sequences.
- the method may then end at block 1740.
- nucleobases at any given position there are sixteen possible combinations of nucleobases at any given position (i.e., an A in the first portion and an A in the second portion, an A in the first portion and a T in the second portion, and so on).
- the light emissions associated with each target sequence during the relevant base calling cycle will be characteristic of the same nucleobase.
- the two portions behave as a single portion, and the identity of the bases at that position are uniquely callable.
- the signals associated with each portion in the relevant base calling cycle will be characteristic of different nucleobases.
- the first signal coming from the first portion have substantially the same intensity as the second signal coming from the second portion.
- the two signals may also be co-localised, and may not be spatially and/or optically resolved. Therefore, when different nucleobases are present at corresponding positions of the two portions, the identity of the nucleobases cannot be uniquely called from the combined signal alone. However, useful sequencing information can still be determined from these signals.
- the scatter plot of Figure 11 shows nine distributions (or bins) of intensity values from the combination of two co-localised signals of substantially equal intensity.
- the intensity values shown in Figure 11 may be up to a scale or normalisation factor; the units of the intensity values may be arbitrary or relative (i.e., representing the ratio of the actual intensity to a reference intensity).
- the sum of the first signal generated from the first portion and the second signal generated from the second portion results in a combined signal.
- the combined signal may be captured by a first optical channel and a second optical channel.
- the computer system can map the combined signal generated into one of the nine bins, and thus determine sequence information relating to the added nucleobase at the first portion and the added nucleobase at the second portion. Bins are selected based upon the combined intensity of the signals originating from each target sequence during the base calling cycle.
- bin 1803 may be selected following the detection of a high-intensity (or “on/on”) signal in the first channel and a high-intensity signal in the second channel.
- Bin 1806 may be selected following the detection of a high-intensity signal in the first channel and an intermediate-intensity (“on/off’ or “off/on”) signal in the second channel.
- Bin 1809 may be selected following the detection of a high-intensity signal in the first channel and a low-intensity or zero-intensity (“off/off’) signal in the second channel.
- Bin 1802 may be selected following the detection of an intermediate-intensity signal in the first channel and a high-intensity signal in the second channel.
- Bin 1805 may be selected following the detection of an intermediate-intensity signal in the first channel and an intermediate-intensity signal in the second channel.
- Bin 1808 may be selected following the detection of an intermediateintensity signal in the first channel and a low-intensity or zero-intensity signal in the second channel.
- Bin 1801 may be selected following the detection of a low-intensity signal in the first channel and a high-intensity signal in the second channel.
- Bin 1804 may be selected following the detection of a low-intensity or zero-intensity signal in the first channel and an intermediateintensity signal in the second channel.
- Bin 1807 may be selected following the detection of a low- intensity or zero-intensity signal in the first channel and a low-intensity signal in the second channel.
- the computer processor may detect a match between the first portion and the second portion at the sensed position.
- the computer processor may base call the respective nucleobases. For example, when the combined signal is mapped to bin 1801 for a base calling cycle, the computer processor base calls both the added nucleobase at the first portion and the added nucleobase at the second portion as T.
- the processor base calls both the added nucleobase at the first portion and the added nucleobase at the second portion as A.
- the processor base calls both the added nucleobase at the first portion and the added nucleobase at the second portion as G.
- the processor base calls both the added nucleobase at the first portion and the added nucleobase at the second portion as C.
- bins 1802, 1804, 1806, and 1808 each represent two possible combinations of first and second nucleobases.
- Bin 1805 meanwhile, represents four possible combinations.
- mapping the combined signal to an ambiguous bin may still allow for sequencing information to be determined.
- bins 1802, 1804, 1805, 1806, and 1808 represent mismatches between respective nucleobases of the two portions sensed during the cycle. Therefore, in response to mapping the combined signal to a bin representing a mismatch, the computer processor may detect a mismatch between the first portion and the second portion at the sensed position.
- A is configured to emit a signal in both the first channel and the second channel
- C is configured to emit a signal in the first channel only
- T is configured to emit a signal in the second channel only
- G does not emit a signal in either channel.
- different permutations of nucleobases can be used to achieve the same effect by performing dye swaps.
- A may be configured to emit a signal in both the first channel and the second channel
- T may be configured to emit a signal in the first channel only
- C may be configured to emit a signal in the second channel only
- G may be configured to not emit a signal in either channel.
- the number of classifications which may be selected based upon the combined signal intensities may be predetermined, for example based on the number of portions expected to be present in the nucleic acid cluster. Whilst Figure 11 shows a set of nine possible classifications, the number of classifications may be greater or smaller.
- the mapping of the combined signal to each of the different bins can provide additional information about the first portion and the second portion, or about sequences from which the first portion and the second portion were derived. For example, given the nucleic acid material input and the processing methods used to generate the nucleic acid clusters, the first portion and the second portion may be expected to be identical at a given position. In this case, the mapping of the combined signal to a bin representing a mismatch may be indicative of an error introduced during library preparation.
- Errors may arise during NGS library preparation, for example due to PCR artifacts or DNA damage.
- the error rate is determined by the library preparation method used, for example the number of cycles of PCR amplification carried out, and a typical error rate may be of the order of 0.1%. This can limit the sensitivity of diagnostic assays based on the sequencing method, and may obscure true variants.
- the present methods allow for the identification of library preparation errors from fewer sequencing reads. In the absence of any library preparation/sequencing errors, the signals produced by sequencing the two portions (e.g. using sequencing-by-synthesis) will match.
- the combined signal may therefore be mapped to one of the four “comer” clouds shown in Figure 11 and Figure 12, and the identity of the nucleobase at the corresponding position of the original library polynucleotide can be determined. Should the identity of the nucleobase at that position suggest a rare, or even unknown, variant, it can be determined with a high level of confidence that the base call represents a true variant, as opposed to a library preparation error. If, on the other hand, the combined signal is mapped to any of the other clouds, this indicates that the sequences of the first portion and the second portion do not match, and that an error has occurred in library preparation. Therefore, in response to mapping the combined signal to a classification representing a mismatch between the two nucleobases, a library preparation error may be identified.
- Figure 13 is a flow diagram showing a method 1900 of determining sequence information according to the present disclosure.
- the described method allows for the determination of sequence information from two (or more) portions (e.g. the first portion and the second portion) in a single sequencing run from a single combined signal obtained from the first portion and the second portion.
- the disclosed method 1900 may start from block 1901. The method may then move to block 1910.
- intensity data is obtained.
- the intensity data includes first intensity data and second intensity data.
- the first intensity data comprises a combined intensity of a first signal component obtained based upon a respective first nucleobase of the first portion and a second signal component obtained based upon a respective second nucleobase of the second portion.
- the second intensity data comprises a combined intensity of a third signal component obtained based upon the respective first nucleobase of the first portion and a fourth signal component obtained based upon the respective second nucleobase of the second portion.
- the first portion is capable of generating a first signal comprising a first signal component and a third signal component.
- the second portion is capable of generating a second signal comprising a second signal component and a fourth signal component.
- the first portion and the second portion may be arranged on the solid support such that signals from the first portion and the second portion are detected by a single sensing portion and/or may comprise a single cluster such that first signals and second signals from each of the respective first portions and second portions cannot be spatially resolved.
- obtaining the intensity data comprises selecting intensity data, for example based upon a chastity score.
- a chastity score may be calculated as the ratio of the brightest base intensity divided by the sum of the brightest and second brightest base intensities.
- high- quality data corresponding to two portions with a substantially equal intensity ratio may have a chastity score of around 0.8 to 0.9, for example 0.89-0.9.
- the method may proceed to block 1920.
- one of a plurality of classifications is selected based on the intensity data.
- Each classification represents one or more possible combinations of respective first and second nucleobases, and at least one classification of the plurality of classifications represents more than one possible combination of respective first and second nucleobases.
- the plurality of classifications comprises nine classifications as shown in Figure 11. Selecting the classification based on the first and second intensity data comprises selecting the classification based on the combined intensity of the first and second signal components and the combined intensity of the third and fourth signal components.
- the method may then proceed to block 1930, where sequence information of the respective first and second nucleobases is determined based on the classification selected in block 1920.
- the signals generated during a cycle of a sequencing are indicative of the identity of the nucleobase(s) added during sequencing (e.g. using sequencing-by-synthesis). For example, it may be determined that there is a match or a mismatch between the respective first and second nucleobases. Where it is determined that there is a match between the first and second respective nucleobases, the nucleobases may be base called. Whether there is a match or a mismatch, additional or alternative information may be obtained, as described above.
- any references herein to the base calling of respective nucleobases at the two portions encompasses the base calling of nucleobases hybridised to the template sequences and, alternatively or additionally, the identification of the corresponding nucleobases of the template sequences.
- the method may then end at block 1940.
- Solid supports enabling ringed complementary reads and methods of preparing polynucleotide sequences for identification
- the present invention is directed to solid supports that enable ringed complementary reads and methods of preparing polynucleotide sequences for identification using the solid support.
- the solid support comprises first immobilised primers and second immobilised primers, wherein some of the first immobilised primers are cleavable and unblocked, some of the first immobilised primers are non-cleavable and blocked, some of the second immobilised primers are cleavable and blocked, and some of the second immobilised primers are non-cleavable and unblocked.
- first immobilised primers and second immobilised primers allows the generation of solid supports that have a central read portion (also referred to herein as a “first region”), and a read portion surrounding the central read portion (also referred to herein as a “second region”).
- Such solid supports are tuneable as the number of amplification cycles used for generating the first region and the second region can be carefully controlled.
- a lower number of amplification cycles may be used for generating the surrounding second region, compared to the number of amplification cycles used for generating the central first region.
- a higher number of amplification cycles may be used for generating the surrounding second region, compared to the number of amplification cycles used for generating the central first region.
- this generates a difference in intensity of signal that can be emitted by the central first region compared to the surrounding second region.
- Such a difference in intensity of signal can be decoded by data analysis methods as described herein, and as such polynucleotide sequences located in the central first region and polynucleotide sequences located in the surrounding second region can be concurrently sequenced.
- the same number of amplification cycles may be used for generating the surrounding second region and the central first region.
- information regarding polynucleotide sequences located in the central first region and polynucleotide sequences located in the surrounding second region can be determined after concurrent sequencing.
- a solid support comprising: a plurality of first immobilised primers, and a plurality of second immobilised primers, wherein a first proportion of the first immobilised primers are configured to be cleavable under first cleavage conditions and are unblocked, wherein a second proportion of the first immobilised primers are configured to be non-cleavable under first cleavage conditions and are blocked, wherein a third proportion of the second immobilised primers are configured to be cleavable under second cleavage conditions and are blocked; and wherein a fourth proportion of the second immobilised primers are configured to be non-cleavable under second cleavage conditions and are unblocked.
- the second immobilised primer is different in sequence to the first immobilised primer.
- the total population of the plurality of first immobilised primers may consist of the first proportion of the first immobilised primers and the second proportion of the first immobilised primers.
- the first immobilised primers may either be configured to be cleavable under first cleavage conditions and are unblocked (i.e. members of the first proportion), or if not, then may be configured to be non-cleavable under first cleavage conditions and are blocked (i.e. members of the second proportion).
- the total population of the plurality of second immobilised primers may consist of the third proportion of the second immobilised primers and the fourth proportion of the second immobilised primers.
- the second immobilised primers may either be configured to be cleavable under second cleavage conditions and are blocked (i.e. members of the third proportion), or if not, then may be configured to be non-cleavable under second cleavage conditions and are unblocked (i.e. members of the fourth proportion).
- a ratio between unblocked immobilised primers and blocked immobilised primers may be between 20:80 to 80:20; in a further embodiment, between 50:50 to 75:25; in an even further embodiment, between 60:40 to 70:30; and in a yet even further embodiment, about 2: 1.
- Such a ratio may advantageously provide a central first region that is able to output a signal that is intense enough to read during sequencing.
- the first proportion of first immobilised primers relative to the total population of first immobilised primers may be between 0.2 to 0.8; in a further embodiment, between 0.5 to 0.75; in an even further embodiment, between 0.6 to 0.7; in a yet even further embodiment, about %.
- the second proportion of first immobilised primers relative to the total population of first immobilised primers may be between 0.8 to 0.2; in a further embodiment, between 0.5 to 0.25; in an even further embodiment, between 0.4 to 0.3; in a yet even further embodiment, about 'A.
- the first proportion of first immobilised primers relative to the total population of first immobilised primers may be between 0.2 to 0.8, 0.5 to 0.75 (in a further embodiment), 0.6 to 0.7 (in an even further embodiment), or about % (in a yet even further embodiment), whilst the respective second proportion of first immobilised primers relative to the total population of first immobilised primers may be between 0.8 to 0.2, 0.5 to 0.25 (in the further embodiment), 0.4 to 0.3 (in the even further embodiment), or about V (in the yet even further embodiment) (wherein the first proportion of first immobilised primers and the second proportion of first immobilised primers sums to 1).
- the fourth proportion of second immobilised primers relative to the total population of second immobilised primers may be between 0.2 to 0.8; in a further embodiment, between 0.5 to 0.75; in an even further embodiment, between 0.6 to 0.7; in a yet even further embodiment, about %.
- the third proportion of second immobilised primers relative to the total population of second immobilised primers may be between 0.8 to 0.2; in a further embodiment, between 0.5 to 0.25; in an even further embodiment, between 0.4 to 0.3; in a yet even further embodiment, about 'A.
- the fourth proportion of second immobilised primers relative to the total population of second immobilised primers may be between 0.2 to 0.8, 0.5 to 0.75 (in a further embodiment), 0.6 to 0.7 (in an even further embodiment), about % (in a yet even further embodiment), whilst the respective third proportion of second immobilised primers relative to the total population of second immobilised primers may be between 0.8 to 0.2, 0.5 to 0.25 (in the further embodiment), 0.4 to 0.3 (in the even further embodiment), or about (in the yet even further embodiment) (wherein the fourth proportion of second immobilised primers and the third proportion of second immobilised primers sums to 1).
- the total population of the first immobilised primers may be equal to the total population of the second immobilised primers.
- the first proportion of first immobilised primers may be equal to the fourth proportion of second immobilised primers (i.e. non-cleavable, unblocked second immobilised primers).
- the second proportion of first immobilised primers may be equal to the third proportion of second immobilised primers (i.e. cleavable, blocked second immobilised primers).
- the first proportion of first immobilised primers may be equal to the fourth proportion of second immobilised primers (i.e. non-cleavable, unblocked second immobilised primers), and the second proportion of first immobilised primers (i.e. non-cleavable, blocked first immobilised primers) may be equal to the third proportion of second immobilised primers (i.e. cleavable, blocked second immobilised primers).
- blocking groups include a hairpin loop (e.g. a polynucleotide attached to the 3 ’-end, comprising in a 5’ to 3’ direction, a cleavable site such as a nucleotide comprising uracil, a loop portion, and a complement portion, wherein the complement portion is substantially complementary to all or a portion of the immobilised primer), a deoxynucleotide, a deoxyribonucleotide, a hydrogen atom instead of a 3 ’-OH group, a phosphate group, a phosphorothioate group, a propyl spacer (e.g.
- a modification blocking the 3 ’-hydroxyl group e.g. hydroxyl protecting groups, such as silyl ether groups (e.g. trimethylsilyl, triethylsilyl, triisopropylsilyl, t-butyl(dimethyl)silyl, t- butyl(diphenyl)silyl), ether groups (e.g. benzyl, allyl, t-butyl, methoxymethyl (MOM), 2- methoxyethoxymethyl (MEM), tetrahydropyranyl), or acyl groups (e.g. acetyl, benzoyl)), or an inverted nucleobase.
- the blocking group may be any modification that prevents extension (i.e. elongation) of the primer by a polymerase.
- the first immobilised primers of the second proportion that are blocked may each comprise a blocking group at a 3 ’ end of the first immobilised primer.
- the blocking group may be selected from the group consisting of: a hairpin loop (e.g.
- a polynucleotide attached to the 3 ’-end comprising in a 5’ to 3’ direction, a cleavable site such as a nucleotide comprising uracil, a loop portion, and a complement portion, wherein the complement portion is substantially complementary to all or a portion of the immobilised primer), a deoxynucleotide, a deoxyribonucleotide, a hydrogen atom instead of a 3 ’-OH group, a phosphate group, a phosphorothioate group, a propyl spacer (e.g.
- a modification blocking the 3 ’-hydroxyl group e.g. hydroxyl protecting groups, such as silyl ether groups (e.g. trimethylsilyl, triethylsilyl, triisopropylsilyl, t-butyl(dimethyl)silyl, t- butyl(diphenyl)silyl), ether groups (e.g. benzyl, allyl, t-butyl, methoxymethyl (MOM), 2- methoxyethoxymethyl (MEM), tetrahydropyranyl), or acyl groups (e.g.
- hydroxyl protecting groups such as silyl ether groups (e.g. trimethylsilyl, triethylsilyl, triisopropylsilyl, t-butyl(dimethyl)silyl, t- butyl(diphenyl)silyl), ether groups (e.g. benzyl, allyl, t-buty
- the blocking group may be a phosphate group or a hairpin loop (e.g. a polynucleotide attached to the 3 ’-end, comprising in a 5’ to 3’ direction, a cleavable site such as a nucleotide comprising uracil, a loop portion, and a complement portion, wherein the complement portion is substantially complementary to all or a portion of the immobilised primer).
- the blocking group may be a phosphate group.
- the second immobilised primers of the third proportion that are blocked may each comprise a blocking group at a 3’ end of the second immobilised primer.
- the blocking group may be selected from the group consisting of: a hairpin loop (e.g. a polynucleotide attached to the 3 ’-end, comprising in a 5’ to 3’ direction, a cleavable site such as a nucleotide comprising uracil, a loop portion, and a complement portion, wherein the complement portion is substantially complementary to all or a portion of the immobilised primer), a deoxynucleotide, a deoxyribonucleotide, a hydrogen atom instead of a 3 ’-OH group, a phosphate group, a phosphorothioate group, a propyl spacer (e.g.
- a modification blocking the 3 ’-hydroxyl group e.g. hydroxyl protecting groups, such as silyl ether groups (e.g. trimethylsilyl, triethylsilyl, triisopropylsilyl, t-butyl(dimethyl)silyl, t- butyl(diphenyl)silyl), ether groups (e.g. benzyl, allyl, t-butyl, methoxymethyl (MOM), 2- methoxyethoxymethyl (MEM), tetrahydropyranyl), or acyl groups (e.g.
- hydroxyl protecting groups such as silyl ether groups (e.g. trimethylsilyl, triethylsilyl, triisopropylsilyl, t-butyl(dimethyl)silyl, t- butyl(diphenyl)silyl), ether groups (e.g. benzyl, allyl, t-buty
- the blocking group may be a phosphate group or a hairpin loop (e.g. a polynucleotide attached to the 3 ’-end, comprising in a 5’ to 3’ direction, a cleavable site such as a nucleotide comprising uracil, a loop portion, and a complement portion, wherein the complement portion is substantially complementary to all or a portion of the immobilised primer).
- the blocking group may be a phosphate group.
- the location at which the first immobilised primer of the first proportion is configured to be cleavable under first cleavage conditions may also be referred to as a first cleavage site.
- the first immobilised primer of the first proportion comprises a first cleavage site.
- the first cleavage site may comprise a cleavable covalent bond.
- sequencing to occur starting from the nick location (e.g. in conjunction with a strand displacement polymerase).
- the first cleavage site when the first cleavage site is nicked, this allows linearisation to occur since one side of a “bridge” formed during bridge amplification can be detached from a solid support.
- the first cleavage site may also be referred to as a first linearisation site (or that the first immobilised primer of the first proportion is configured to be linearisable under first linearisation conditions).
- the location at which the second immobilised primer of the third proportion is configured to be cleavable under second cleavage conditions may also be referred to as a second cleavage site.
- the second immobilised primer of the third proportion comprises a second cleavage site.
- the second cleavage site may comprise a cleavable covalent bond.
- sequencing to occur starting from the nick location (e.g. in conjunction with a strand displacement polymerase).
- the second cleavage site when the second cleavage site is nicked, this allows linearisation to occur since one side of a “bridge” formed during bridge amplification can be detached from a solid support.
- the second cleavage site may also be referred to as a second linearisation site (or that the second immobilised primer of the third proportion is configured to be linearisable under second linearisation conditions).
- first cleavage conditions refers to reaction conditions that cause cleavage within the first immobilised primer of the first proportion (i.e. at the first cleavage site).
- the first cleavage conditions may involve exposure to a thermal trigger (e.g. by heating), a light trigger (e.g. by exposure to ultraviolet light), and/or a chemical/biochemical trigger (e.g. an enzyme, a metal catalyst including transition metal catalysts such as a palladium-based or a nickel-based catalyst, periodate).
- a thermal trigger e.g. by heating
- a light trigger e.g. by exposure to ultraviolet light
- a chemical/biochemical trigger e.g. an enzyme, a metal catalyst including transition metal catalysts such as a palladium-based or a nickel-based catalyst, periodate.
- the “first cleavage conditions” may allow linearisation to occur, and may be referred to as “first linearisation conditions”.
- the first immobilised primers of the first proportion may be configured to be cleavable by a thermal trigger (e.g. by heating), a light trigger (e.g. by exposure to ultraviolet light), and/or a chemical/biochemical trigger (e.g. an enzyme, a metal catalyst including transition metal catalysts such as a palladium-based or a nickel-based catalyst, periodate).
- a thermal trigger e.g. by heating
- a light trigger e.g. by exposure to ultraviolet light
- a chemical/biochemical trigger e.g. an enzyme, a metal catalyst including transition metal catalysts such as a palladium-based or a nickel-based catalyst, periodate.
- the first immobilised primers of the first proportion may be configured to be cleavable by a metal catalyst.
- the first cleavage conditions involve exposure to a metal catalyst.
- the metal catalyst is a transition metal catalyst.
- the metal catalyst is a palladium-based or a nickel-based catalyst.
- the metal catalyst is a palladium-based catalyst (e.g. a Pd(0) catalyst).
- Non-limiting examples of suitable palladium-based catalysts are described in WO 2019/222264 Al (which is incorporated herein by reference), such as Pd(THP)2, Pd(THP)s, Pd(THP)4, Pd(THM)4 (where THP is tris(hydroxypropyl)phosphine and where THM is tris(hydroxymethyl)phosphine).
- Such palladium-based catalysts may be generated in situ from corresponding Pd(0) or Pd(II) complexes.
- each first immobilised primer of the first proportion may comprise a nucleotide that comprises an allyl group, for example an allyl group attached to a sugar group (e.g. deoxyribose or ribose) of the nucleotide.
- the allyl group may have a structure according to the following: wherein R represents an attachment point to hydrogen (in which case, R is hydrogen) or the remainder of the sugar group (e.g. deoxyribose or ribose).
- the nucleotide may have a structure according to the following: wherein represents an attachment point of the nucleotide to the rest of the first immobilised primer of the first proportion, and wherein “base” represents adenine, guanine, cytosine, thymine or uracil, or a derivative thereof.
- the nucleotide may have a structure according to the following: wherein represents an attachment point of the nucleotide to the rest of the first immobilised primer of the first proportion.
- the first immobilised primers of the first proportion may be configured to be cleavable by a glycosylase.
- the first cleavage conditions involve exposure to a glycosylase.
- the first proportion of first immobilised primers may be configured to be cleavable by a glycosylase that recognises any nitrogenous base (e.g.
- the first proportion of first immobilised primers may be configured to be cleavable by a glycosylase that recognises any nitrogenous base (e.g. purine or pyrimidine) which is not selected from guanine (G), cytosine (C), adenine (A) and uracil (U) when the first immobilised primer is an RNA sequence.
- the glycosylase may recognise an unnatural nucleobase (i.e. one which is not usually present in a typical DNA sequence an RNA sequence).
- unnatural nucleobases may include oxoguanine (e.g. 8 -oxoguanine), hypoxanthine, xanthine, methylguanines (e.g. O 6 -methylguanine, N 7 - methylguanine), methyladenines (e.g. 3 -methyladenine, N 6 -methyladenine), modified cytosines including methylcytosines (e.g. 5-methylcytosine, 5-hydroxymethylcytosine, 5-formylcytosine, 5 -carboxylcytosine), dihydrouracil, inosine, and uracil (if the first immobilised primer is a DNA sequence).
- oxoguanine e.g. 8 -oxoguanine
- hypoxanthine e.g. O 6 -methylguanine, N 7 - methylguanine
- methyladenines e.g. 3 -methyladenine, N 6 -
- the first immobilised primers of the first proportion may be configured to be cleavable by a uracil glycosylase (when the first immobilised primer is a DNA sequence) or an oxoguanine glycosylase (e.g. 8-oxoguanine glycosylase).
- each first immobilised primer of the first proportion may comprise a nucleobase which is not selected from guanine, cytosine, adenine or thymine when the first immobilised primer is a DNA sequence; or wherein each first immobilised primer of the first proportion may comprise a nucleobase which is not selected from guanine, cytosine, adenine or uracil when the first immobilised primer is an RNA sequence.
- each first immobilised primer of the first proportion may comprise an unnatural nucleobase (i.e. one which is not usually present in a typical DNA sequence an RNA sequence).
- examples of unnatural nucleobases may include oxoguanine (e.g.
- 8-oxoguanine hypoxanthine, xanthine, methylguanines (e.g. O 6 -methylguanine, N 7 -methylguanine), methyladenines (e.g. 3- methyladenine, N 6 -methyladenine), modified cytosines including methylcytosines (e.g. 5- methylcytosine, 5-hydroxymethylcytosine, 5-formylcytosine, 5 -carboxylcytosine), dihydrouracil, inosine, and uracil (if the first immobilised primer is a DNA sequence).
- methylcytosines e.g. 5- methylcytosine, 5-hydroxymethylcytosine, 5-formylcytosine, 5 -carboxylcytosine
- dihydrouracil inosine
- uracil if the first immobilised primer is a DNA sequence.
- each first immobilised primer of the first proportion may comprise oxoguanine (e.g. 8-oxoguanine) or uracil when the first immobilised primer is a DNA sequence, or wherein each first immobilised primer of the first proportion may comprise oxoguanine (e.g. 8-oxoguanine) when the first immobilised primer is an RNA sequence.
- oxoguanine e.g. 8-oxoguanine
- uracil when the first immobilised primer is a DNA sequence
- each first immobilised primer of the first proportion may comprise oxoguanine (e.g. 8-oxoguanine) when the first immobilised primer is an RNA sequence.
- the term “second cleavage conditions” refers to reaction conditions that cause cleavage within the second immobilised primer of the third proportion (i.e. at the second cleavage site).
- the second cleavage conditions may involve exposure to athermal trigger (e.g. by heating), a light trigger (e.g. by exposure to ultraviolet light), and/or a chemical/biochemical trigger (e.g. an enzyme, a metal catalyst including transition metal catalysts such as a palladium-based or a nickel-based catalyst, periodate).
- the “second cleavage conditions” may allow linearisation to occur, and may be referred to as “second linearisation conditions”.
- the second immobilised primers of the third proportion may be configured to be cleavable by a thermal trigger (e.g. by heating), a light trigger (e.g. by exposure to ultraviolet light), and/or a chemical/biochemical trigger (e.g. an enzyme, a metal catalyst including transition metal catalysts such as a palladium-based or a nickel-based catalyst, periodate).
- a thermal trigger e.g. by heating
- a light trigger e.g. by exposure to ultraviolet light
- a chemical/biochemical trigger e.g. an enzyme, a metal catalyst including transition metal catalysts such as a palladium-based or a nickel-based catalyst, periodate.
- the second immobilised primers of the third proportion may be configured to be cleavable by a metal catalyst.
- the first cleavage conditions involve exposure to a metal catalyst.
- the metal catalyst is a transition metal catalyst.
- the metal catalyst is a palladium-based or a nickel-based catalyst.
- the metal catalyst is a palladium-based catalyst (e.g. a Pd(0) catalyst).
- Non-limiting examples of suitable palladium-based catalysts are described in WO 2019/222264 Al (which is incorporated herein by reference), such as Pd(THP)2, Pd(THP)s, Pd(THP)4, Pd(THM)4 (where THP is tris(hydroxypropyl)phosphine and where THM is tris(hydroxymethyl)phosphine).
- Such palladium-based catalysts may be generated in situ from corresponding Pd(0) or Pd(II) complexes.
- each second immobilised primer of the third proportion may comprise a nucleotide that comprises an allyl group, for example an allyl group attached to a sugar group (e.g. deoxyribose or ribose) of the nucleotide.
- the allyl group may have a structure according to the following: wherein R represents an attachment point to hydrogen (in which case, R is hydrogen) or the remainder of the sugar group (e.g. deoxyribose or ribose).
- the nucleotide may have a structure according to the following: wherein represents an attachment point of the nucleotide to the rest of the second immobilised primer of the third proportion, and wherein “base” represents adenine, guanine, cytosine, thymine or uracil, or a derivative thereof.
- the nucleotide may have a structure according to the following: wherein represents an attachment point of the nucleotide to the rest of the second immobilised primer of the third proportion.
- the second immobilised primers of the third proportion may be configured to be cleavable by a glycosylase.
- the second cleavage conditions involve exposure to a glycosylase.
- the third proportion of second immobilised primers may be configured to be cleavable by a glycosylase that recognises any nitrogenous base (e.g.
- the third proportion of second immobilised primers may be configured to be cleavable by a glycosylase that recognises any nitrogenous base (e.g. purine or pyrimidine) which is not selected from guanine (G), cytosine (C), adenine (A) and uracil (U) when the second immobilised primer is an RNA sequence.
- the glycosylase may recognise an unnatural nucleobase (i.e. one which is not usually present in a typical DNA sequence an RNA sequence).
- unnatural nucleobases may include oxoguanine (e.g. 8 -oxoguanine), hypoxanthine, xanthine, methylguanines (e.g. O 6 - methylguanine, N 7 -methylguanine), methyladenines (e.g. 3 -methyladenine, N 6 -methyladenine), modified cytosines including methylcytosines (e.g. 5 -methylcytosine, 5-hydroxymethylcytosine, 5 -formylcytosine, 5 -carboxylcytosine), dihydrouracil, inosine, and uracil (if the second immobilised primer is a DNA sequence).
- oxoguanine e.g. 8 -oxoguanine
- hypoxanthine e.g. O 6 - methylguanine, N 7 -methylguanine
- methyladenines e.g. 3 -methyladenine,
- the second immobilised primers of the third proportion may be configured to be cleavable by a uracil glycosylase (when the second immobilised primer is a DNA sequence) or an oxoguanine glycosylase (e.g. 8-oxoguanine glycosylase).
- each second immobilised primer of the third proportion may comprise a nucleobase which is not selected from guanine, cytosine, adenine or thymine when the second immobilised primer is a DNA sequence, or wherein each second immobilised primer of the third proportion may comprise a nucleobase which is not selected from guanine, cytosine, adenine or uracil when the second immobilised primer is an RNA sequence.
- each second immobilised primer of the third proportion may comprise an unnatural nucleobase (i.e. one which is not usually present in a typical DNA sequence an RNA sequence).
- examples of unnatural nucleobases may include oxoguanine (e.g.
- methylguanines e.g. O 6 -methylguanine, N 7 -methylguanine
- methyladenines e.g. 3- methyladenine, N 6 -methyladenine
- modified cytosines including methylcytosines (e.g. 5- methylcytosine, 5-hydroxymethylcytosine, 5-formylcytosine, 5 -carboxylcytosine), dihydrouracil, inosine, and uracil (if the second immobilised primer is a DNA sequence).
- each second immobilised primer of the third proportion may comprise oxoguanine (e.g. 8-oxoguanine) or uracil when the second immobilised primer is a DNA sequence, or wherein each second immobilised primer of the third proportion may comprise oxoguanine (e.g. 8-oxoguanine) when the second immobilised primer is an RNA sequence.
- oxoguanine e.g. 8-oxoguanine
- uracil when the second immobilised primer is a DNA sequence
- each second immobilised primer of the third proportion may comprise oxoguanine (e.g. 8-oxoguanine) when the second immobilised primer is an RNA sequence.
- each first immobilised primer of the first proportion and each second immobilised primer of the third proportion may comprise a nucleotide that comprises an allyl group, for example an allyl group attached to a sugar group (e.g. deoxyribose or ribose) of the nucleotide.
- the allyl group may have a structure according to the following: wherein R represents an attachment point to hydrogen (in which case, R is hydrogen) or the remainder of the sugar group (e.g. deoxyribose or ribose).
- the nucleotide may have a structure according to the following: wherein represents an attachment point of the nucleotide to the rest of the first immobilised primer of the first proportion or the rest of the second immobilised primer of the third proportion, and wherein “base” represents adenine, guanine, cytosine, thymine or uracil, or a derivative thereof.
- the nucleotide may have a structure according to the following: wherein represents an attachment point of the nucleotide to the rest of the first immobilised primer of the first proportion or the rest of the second immobilised primer of the third proportion.
- the term “glycosylase” may refer to an enzyme which catalyses the removal of a nitrogenous base from one of the nucleotides in a (poly)nucleotide chain by breaking a N- glycosidic bond, resulting in the formation of an apurinic/apyrimidinic site (AP site).
- the glycosylase may recognise any nitrogenous base (e.g. purine or pyrimidine) which is not selected from guanine (G), cytosine (C), adenine (A) and thymine (T); for RNA chains, the glycosylase may recognise any nitrogenous base (e.g.
- purine or pyrimidine which is not selected from guanine (G), cytosine (C), adenine (A) and uracil (U).
- G guanine
- C cytosine
- A adenine
- U uracil
- typical nitrogenous bases recognised by glycosylases include oxoguanine (e.g. 8-oxoguanine), uracil and alkylpurines.
- Glycosylases may be monofunctional, such that they only possess glycosylase activity (i.e. breaking of the N-glycosidic bond) - cleavage of a phosphodiester bond in the sugar-phosphate backbone may then occur in an uncatalysed manner by elimination.
- Other glycosylases may be bifimctional, such that they also possess AP lyase activity by catalysing the phosphodiester bond of the (poly)nucleotide chain.
- the glycosylase is bifunctional (i.e. possesses both glycosylase and AP lyase activity).
- the first cleavage conditions and the second cleavage conditions may be the same or may be different. In some embodiments, the first cleavage conditions and the second cleavage conditions may be the same. This allows the cleavage within the first immobilised primers and the second immobilised primers to occur using a single exposure, which reduces the number of steps required for preparing first polynucleotide sequences and second polynucleotide sequences for concurrent sequencing. In alternative embodiments, the first cleavage conditions and the second cleavage conditions may be different. This allows control of which of the first immobilised primer and/or the second immobilised primer become cleaved, should this be necessary during the preparation processes.
- the first proportion of first immobilised primers that are cleavable may be further configured to be linearisable under first linearisation conditions.
- the third proportion of second immobilised primers that are cleavable may be further configured to be linearisable under second linearisation conditions.
- first proportion of first immobilised primers that are cleavable may be further configured to be linearisable under first linearisation conditions; and the third proportion of second immobilised primers that are cleavable may be further configured to be linearisable under second linearisation conditions.
- first linearisation conditions and the second linearisation conditions may be the same or may be different. In some embodiments, the first linearisation conditions and the second linearisation conditions may be the same. This allows the linearisation within the first immobilised primers and the second immobilised primers to occur using a single exposure, which reduces the number of steps required for preparing first polynucleotide sequences and second polynucleotide sequences for concurrent sequencing. In alternative embodiments, the first linearisation conditions and the second linearisation conditions may be different. This allows control of which of the first immobilised primer and/or the second immobilised primer become linearised, should this be necessary during the preparation processes.
- each first immobilised primer may comprise a sequence as defined in SEQ ID NO. 1 or 5, or a variant or fragment thereof; and each second immobilised primer may comprise a sequence as defined in SEQ ID NO. 2, or a variant or fragment thereof; or wherein each first immobilised primer may comprise a sequence as defined in SEQ ID NO. 2, or a variant or fragment thereof; and each second immobilised primer may comprise a sequence as defined in SEQ ID NO. 1 or 5, or a variant or fragment thereof.
- the solid support described above comprising the first proportion of first immobilised primers, the second proportion of first immobilised primers, the third proportion of second immobilised primers and the fourth proportion of second immobilised primers may be considered to be a precursor solid support that, once processed according to the methods described herein, forms a solid support that comprises a central first region and a surrounding second region.
- another aspect of the present invention is directed to a solid support, comprising: a plurality of first immobilised primers, and a plurality of second immobilised primers, wherein the plurality of first immobilised primers are located in a first region, wherein the plurality of second immobilised primers are located in a second region surrounding the first region.
- the second immobilised primer is different in sequence to the first immobilised primer.
- the second region may be contiguous.
- the second region may be an annular region.
- the second region may be an annular region surrounding a circular first region.
- the solid support may comprise a plurality of first regions with a respective plurality of second regions surrounding the first regions. Each first region and second region may be considered to be a “well” on the solid support.
- the plurality of first immobilised primers in the first region may be extended to form a plurality of template sequences.
- the plurality of second immobilised primers in the second region may be extended to form a plurality of template complement sequences.
- the plurality of first immobilised primers in the first region may be extended to form a plurality of template sequences, and the plurality of second immobilised primers in the second region may be extended to form a plurality of template complement sequences.
- the template sequences and the template complement sequences may have been removed from the solid support. Accordingly, in some embodiments, the plurality of first immobilised primers in the first region may not be extended. In some embodiments, the plurality of second immobilised primers in the second region may not be extended. In some embodiments, the plurality of first immobilised primers in the first region may not be extended, and the plurality of second immobilised primers in the second region may not be extended.
- the concentration of first immobilised primers in the first region may be the same or different compared to the concentration of second immobilised primers in the second region.
- concentrations are different, this generates a difference in intensity of signal that can be emitted by the central first region compared to the surrounding second region, which allows polynucleotide sequences located in the central first region and the surrounding second region to be concurrently sequenced (in particular, to determine sequence information, mismatched base pairs, or other information in the polynucleotide sequences including epigenetic modifications).
- concentrations are the same, information can still be obtained using concurrent sequencing (e.g. mismatched base pairs, or other information in the polynucleotide sequences including epigenetic modifications).
- a concentration of first immobilised primers in the first region may be greater than a concentration of second immobilised primers in the second region.
- a ratio between the concentration of first immobilised primers in the first region and the concentration of second immobilised primers in the second region may be between 1.25: 1 to 5 : 1 ; in an even further embodiment, between 1.5 : 1 to 3 : 1 ; in a yet even further embodiment, about 2: 1.
- a concentration of first immobilised primers in the first region may be less than a concentration of second immobilised primers in the second region.
- a ratio between the concentration of first immobilised primers in the first region and the concentration of second immobilised primers in the second region may be between 1: 1.25 to 1:5; in an even further embodiment, between 1: 1.5 to 1:3; in a yet even further embodiment, about 1:2.
- a concentration of first immobilised primers in the first region may be equal or substantially equal to a concentration of second immobilised primers in the second region.
- solid supports as described herein are useful in nucleic acid sequencing, particularly concurrent sequencing.
- a solid support as described herein in nucleic acid sequencing there is provided a process of manufacturing a solid support, comprising:
- the second precursor primer is different in sequence to the first precursor primer.
- the solid support to be manufactured may be a solid support as described herein (in particular, a precursor solid support comprising first proportions, second proportions, third proportions and fourth proportions as described herein). Accordingly, aspects relating to the solid support and other characteristics of the solid support (such as the first immobilised primers and the second immobilised primers) as described herein apply equally to processes as described herein for manufacturing the solid support.
- first precursor primer refers to a state of the first immobilised primers of the solid support before they are immobilised to the solid support. As such, the first precursor primers may be provided as “free” primers in solution. After immobilisation, the “first precursor primers” are then referred to as “first immobilised primers”.
- second precursor primer refers to a state of the second immobilised primers of the solid support before they are immobilised to the solid support. As such, the second precursor primers may be provided as “free” primers in solution. After immobilisation, the “second precursor primers” are then referred to as “second immobilised primers”.
- Steps (a) and (b) may be conducted sequentially or simultaneously.
- step (b) may be conducted after step (a).
- step (a) may be conducted after step (b).
- steps (a) and (b) may be conducted simultaneously.
- the immobilisation method is not particularly limited provided that the first immobilised primers and the second immobilised primers remain on the solid support during amplification, clustering and sequencing.
- immobilisation may comprise forming covalent linkages between the solid support and each of the plurality of first precursor primers, and between the solid support and each of the plurality of second precursor primers.
- the forming covalent linkages involves using a click reaction (e.g. metal-catalysed azide-alkyne cycloaddition reactions, such as copper-catalysed azide-alkyne cycloaddition reactions and strain-promoted azide-alkyne cycloadditions).
- forming covalent linkages may involve forming a 1,2,3-triazole linkage.
- the solid support prior to immobilisation may include azide moieties (e.g. PAZAM), whilst the first precursor primers and the second precursor primers may each comprise alkyne moieties (e.g. terminal alkynes, cycloalkynes).
- a click reaction between the azide moieties on the solid support and the alkyne moieties on the first precursor primers and the second precursor primers allows a 1,2,3-triazole linkage to be formed.
- azide moieties and alkyne moieties can also be swapped, for example by including alkyne moieties on the solid support prior to immobilisation, and including azide moieties on each of the first precursor primers and the second precursor primers.
- solid supports as described herein may be useful in methods of preparing polynucleotide sequences for identification.
- a method of preparing polynucleotide sequences for identification comprising: providing a solid support as described herein, and synthesising a plurality of template sequences that extend from the second immobilised primers in the fourth proportion and a plurality of template complement sequences that extend from the first immobilised primers in the first proportion.
- identification is meant here obtaining genetic information from the polynucleotide strands. This may include identification of the genetic sequence of the polynucleotide strands (i.e. sequencing). Furthermore, this may instead, or additionally, include identification of mismatched base pairs. In addition, this may instead, or additionally, include identification of any epigenetic modifications, for example methylation. Accordingly, “identification” may mean identification of the genetic sequence of the polynucleotide strands, mismatched base pairs, and/or identification of any epigenetic modifications.
- the present invention can be applied to (separate) polynucleotide strands where a first strand comprises a first portion to be identified and a second strand comprises a second portion to be identified.
- first portions also referred to as “template sequences” herein
- second portions also referred to as “template complement sequences” herein
- the first portions and second portions may be different polynucleotide sequences. That is, the sequences may be genetically unrelated and/or derived from different sources.
- first portions and second portions may be genetically related.
- the (separate) polynucleotide strands may comprise a first strand that comprises a first portion that may comprise (or be) the forward strand of a polynucleotide sequence (e.g. forward strand of a template), and a second strand that comprises a second portion that may comprise (or be) the reverse strand of the polynucleotide sequence (e.g. reverse strand of the template) or the forward complement strand of the polynucleotide sequence (e.g. forward complement strand of the template).
- the (separate) polynucleotide strands may comprise a first strand that comprises a first portion that may comprise (or be) the reverse strand of a polynucleotide sequence (e.g. reverse strand of a template), and a second strand that comprises a second portion that may comprise (or be) the forward strand of the polynucleotide sequence (e.g. forward strand of the template) or the reverse complement strand of the polynucleotide sequence (e.g. reverse complement strand of the template).
- the (separate) polynucleotide strands may comprise a first strand that comprises a first portion that may comprise (or be) the forward strand of a polynucleotide sequence (e.g. forward strand of a template), and a second strand that comprises a second portion that may comprise (or be) the reverse complement strand of the polynucleotide sequence (e.g. reverse complement strand of the template) (in effect, a reverse complement strand may be considered a “copy” of the forward strand).
- a reverse complement strand may be considered a “copy” of the forward strand.
- the (separate) polynucleotide strands may comprise a first strand that comprises a first portion that may comprise (or be) the reverse strand of a polynucleotide sequence (e.g. reverse strand of a template), and a second strand that comprises a second portion that may comprise (or be) the forward complement strand of the polynucleotide sequence (e.g. forward complement strand of the template) (in effect, a forward complement strand may be considered a “copy” of the reverse strand).
- the first portion may be derived from a forward strand of a target polynucleotide to be sequenced, and the second portion may be derived from a reverse complement strand of the target polynucleotide to be sequenced; or the first portion may be derived from a reverse strand of a target polynucleotide to be sequenced, and the second portion may be derived from a forward complement strand of the target polynucleotide to be sequenced.
- concurrent sequencing of both the forward and reverse complement strands (or the reverse and forward complement strands) allows mismatched base pairs and/or epigenetic modification to be detected.
- the first portion may be referred to herein as read 1 (Rl).
- the second portion may be referred to herein as read 2 (R2).
- the first portion is at least 25 or at least 50 base pairs and the second portion is at least 25 base pairs or at least 50 base pairs.
- the polynucleotide strands may form or be part of a cluster on the solid support.
- the cluster may refer to polynucleotide strands located within a central first region and its associated surrounding second region.
- cluster may refer to a clonal group of template polynucleotides (e.g. DNA or RNA) bound within a single well of a solid support (e.g. flow cell).
- a cluster may refer to the population of polynucleotide molecules within a well that are then sequenced.
- a “cluster” may contain a sufficient number of copies of template polynucleotides such that the cluster is able to output a signal (e.g. a light signal) that allows sequencing reads to be performed on the cluster.
- a “cluster” may comprise, for example, about 500 to about 2000 copies, about 600 to about 1800 copies, about 700 to about 1600 copies, about 800 to about 1400 copies, about 900 to about 1200 copies, or about 1000 copies of template polynucleotides.
- a cluster may be formed by bridge amplification, as described above.
- the cluster formed may be a duoclonal cluster.
- duoclonal cluster is meant that the population of polynucleotide sequences that are then sequenced (as the next step) are substantially of two types - e.g. a first sequence and a second sequence.
- a “duoclonal” cluster may refer to the population of single first sequences and single second sequences within a well that are then sequenced.
- a “duoclonal” cluster may contain a sufficient number of copies of a single first sequence and copies of a single second sequence such that the cluster is able to output a signal (e.g. a light signal) that allows sequencing reads to be performed on the “monoclonal” cluster.
- a “duoclonal” cluster may comprise, for example, about 500 to about 2000 combined copies, about 600 to about 1800 combined copies, about 700 to about 1600 combined copies, about 800 to about 1400 combined copies, about 900 to about 1200 combined copies, or about 1000 combined copies of single first sequences and single second sequences.
- the copies of single first sequences and single second sequences together may comprise at least about 50%, at least about 60%, at least about 70%, even at least about 80%, at least about 90%, or about 95%, 98%, 99% or 100% of all polynucleotides within a single well of the flow cell, and thus providing a substantially duoclonal “cluster”.
- a first signal that is capable of being produced by the first portion and a second signal that is capable of being produced by the second portion may be optically unresolved.
- the step of synthesising the plurality of template sequences extending from the second immobilised primers in the fourth proportion and the plurality of template complement sequences extending from the first immobilised primers in the first proportion may involve conducting amplification; in a further embodiment, bridge amplification.
- amplification (of the plurality of template sequences extending from the second immobilised primers in the fourth proportion and the plurality of template complement sequences extending from the first immobilised primers in the first proportion) may be conducted over 20 to 40 cycles; in a further embodiment, 25 to 35 cycles.
- the method may further comprise a step of removing second immobilised primers in the fourth proportion that are not yet extended, and removing first immobilised primers in the first proportion that are not yet extended.
- the step of removing second immobilised primers in the fourth proportion that are not yet extended, and removing first immobilised primers in the first proportion that are not yet extended may be conducted after (e.g. immediately after) the step of synthesising the plurality of template sequences that extend from the second immobilised primers in the fourth proportion and the plurality of template complement sequences that extend from the first immobilised primers in the first proportion.
- the step of removing second immobilised primers in the fourth proportion that are not yet extended and removing first immobilised primers in the first proportion that are not yet extended may be conducted using a 3 ’-5’ exonuclease (e.g. Exol).
- a 3 ’-5’ exonuclease e.g. Exol
- the method may further comprise a step of unblocking the second proportion of first immobilised primers and unblocking the third proportion of second immobilised primers (hereinafter referred to as “unblocking conditions”).
- the step of unblocking the second proportion of first immobilised primers and unblocking the third proportion of second immobilised primers may be conducted after (e.g. immediately after) the step of removing second immobilised primers in the fourth proportion that are not yet extended, and removing first immobilised primers in the first proportion that are not yet extended.
- the step of unblocking the second proportion of first immobilised primers and unblocking the third proportion of second immobilised primers may involve removing phosphate groups or hairpin loops from the second proportion of first immobilised primers and removing phosphate groups or hairpin loops from the third proportion of second immobilised primers.
- the step of unblocking the second proportion of first immobilised primers and unblocking the third proportion of second immobilised primers may involve removing phosphate groups from the second proportion of first immobilised primers and removing phosphate groups from the third proportion of second immobilised primers.
- the unblocking conditions may be different to those of the first cleavage conditions and/or the second cleavage conditions.
- the step of unblocking the second proportion of first immobilised primers and unblocking the third proportion of second immobilised primers may be conducted selectively over cleaving the first proportion of first immobilised primers and/or cleaving the third proportion of second immobilised primers.
- the method may further comprise a step of synthesising a plurality of template sequences that extend from the second immobilised primers in the third proportion and a plurality of template complement sequences that extend from the first immobilised primers in the second proportion.
- the step of synthesising the plurality of template sequences that extend from the second immobilised primers in the third proportion and the plurality of template complement sequences that extend from the first immobilised primers in the second proportion may be conducted after (e.g. immediately after) the step of unblocking the second proportion of first immobilised primers and unblocking the third proportion of second immobilised primers.
- the step of synthesising the plurality of template sequences extending from the second immobilised primers in the third proportion and the plurality of template complement sequences extending from the first immobilised primers in the second proportion may involve conducting amplification; in a further embodiment, bridge amplification.
- amplification (of the plurality of template sequences extending from the second immobilised primers in the third proportion and the plurality of template complement sequences extending from the first immobilised primers in the second proportion) may be conducted over 5 to 25 cycles; in a further embodiment, 10 to 20 cycles.
- the number of amplification cycles used for generating the central first region and the surrounding second region can be tuned.
- the number of amplification cycles used for synthesising the plurality of template sequences extending from the second immobilised primers in the third proportion and the plurality of template complement sequences extending from the first immobilised primers in the second proportion may be less than the number of amplification cycles used for synthesising the plurality of template sequences extending from the second immobilised primers in the fourth proportion and the plurality of template complement sequences extending from the first immobilised primers in the first proportion.
- the number of amplification cycles used for synthesising the plurality of template sequences extending from the second immobilised primers in the third proportion and the plurality of template complement sequences extending from the first immobilised primers in the second proportion may be more than the number of amplification cycles used for synthesising the plurality of template sequences extending from the second immobilised primers in the fourth proportion and the plurality of template complement sequences extending from the first immobilised primers in the first proportion.
- the number of amplification cycles used for synthesising the plurality of template sequences extending from the second immobilised primers in the third proportion and the plurality of template complement sequences extending from the first immobilised primers in the second proportion may be the same as the number of amplification cycles used for synthesising the plurality of template sequences extending from the second immobilised primers in the fourth proportion and the plurality of template complement sequences extending from the first immobilised primers in the first proportion.
- the method may further comprise a step of removing second immobilised primers in the third proportion that are not yet extended, and removing first immobilised primers in the second proportion that are not yet extended.
- the step of removing second immobilised primers in the third proportion that are not yet extended, and removing first immobilised primers in the second proportion that are not yet extended may be conducted after (e.g. immediately after) the step of synthesising the plurality of template sequences that extend from the second immobilised primers in the third proportion and the plurality of template complement sequences that extend from the first immobilised primers in the second proportion.
- the step of removing second immobilised primers in the third proportion that are not yet extended and removing first immobilised primers in the second proportion that are not yet extended may be conducted using a 3 ’-5’ exonuclease (e.g. Exol).
- the method may further comprise a step of exposing the solid support to first cleavage conditions and/or second cleavage conditions after the step of synthesising the plurality of template sequences that extend from the second immobilised primers in the third proportion and the plurality of template complement sequences that extend from the first immobilised primers in the second proportion.
- Suitable first cleavage conditions and second cleavage conditions are described above.
- the first cleavage conditions and/or second cleavage conditions may comprise exposure to a thermal trigger (e.g. by heating), a light trigger (e.g. by exposure to ultraviolet light), and/or a chemical/biochemical trigger (e.g. an enzyme, a metal catalyst including transition metal catalysts such as a palladium-based or a nickel-based catalyst, periodate).
- the solid support may be exposed to the first cleavage conditions and then subsequently exposed to the second cleavage conditions, or may be exposed to the second cleavage conditions and then subsequently exposed to the first cleavage conditions (i.e. sequentially). Alternatively, the solid support may be exposed to the first cleavage conditions and the second cleavage conditions at the same time (i.e. simultaneously).
- the solid support may be exposed to a metal catalyst.
- the first cleavage conditions and/or second cleavage conditions may involve exposure to a metal catalyst. Suitable metal catalysts are described above.
- the metal catalyst is a transition metal catalyst.
- the metal catalyst is a palladium-based or a nickel-based catalyst.
- the metal catalyst is a palladiumbased catalyst (e.g. a Pd(0) catalyst).
- the solid support may be exposed to a glycosylase.
- the first cleavage conditions and/or second cleavage conditions may involve exposure to a glycosylase. Suitable glycosylases are described above.
- the solid support is exposed to a uracil glycosylase and/or an oxoguanine glycosylase (e.g. 8-oxoguanine glycosylase).
- the method may further comprise a step of linearising the plurality of template sequences extending from the second immobilised primers of the fourth proportion and linearising the plurality of template complement sequences extending from the first immobilised primers of the second proportion.
- the method may further comprise treating the linearised template sequences and the linearised template complement sequences with a single-stranded binding protein.
- Also described herein is a method of sequencing polynucleotide sequences, comprising preparing polynucleotide sequences for identification using a method as described herein; and concurrently sequencing nucleobases in the template sequences extending from the second immobilised primers of the fourth proportion and the template complement sequences extending from the first immobilised primers of the second proportion.
- sequencing is performed by a sequencing-by-synthesis or sequencing-by- ligation process.
- the step of concurrently sequencing nucleobases may comprise treatment with a strand displacement polymerase (e.g. phi29).
- a strand displacement polymerase e.g. phi29
- the method may further comprise a step of conducting paired-end reads.
- the data may be analysed using 16 QAM as mentioned herein. For example, this may be utilised when a concentration of first immobilised primers in the first region is greater than a concentration of second immobilised primers in the second region on certain types of solid support, or when a concentration of first immobilised primers in the first region is less than a concentration of second immobilised primers in the second region on certain types of solid support.
- the step of concurrently sequencing nucleobases may comprise: (a) obtaining first intensity data comprising a combined intensity of a first signal component obtained based upon a respective first nucleobase at a first portion of the template sequence and a second signal component obtained based upon a respective second nucleobase at a second portion of the template complement sequence, wherein the first and second signal components are obtained simultaneously;
- each classification represents a possible combination of respective first and second nucleobases
- selecting the classification based on the first and second intensity data may comprise selecting the classification based on the combined intensity of the first and second signal components and the combined intensity of the third and fourth signal components.
- the plurality of classifications may comprise sixteen classifications, each classification representing one of sixteen unique combinations of first and second nucleobases.
- the first signal component, second signal component, third signal component and fourth signal component may be generated based on light emissions associated with the respective nucleobase.
- the light emissions may be detected by a sensor, wherein the sensor is configured to provide a single output based upon the first and second signals.
- the senor may comprise a single sensing element.
- the method may further comprise repeating steps (a) to (d) for each of a plurality of base calling cycles.
- the data may be analysed using 9 QAM as mentioned herein. For example, this may be utilised when a concentration of first immobilised primers in the first region is equal or substantially equal to a concentration of second immobilised primers in the second region on certain types of solid support.
- the step of concurrently sequencing nucleobases may comprise:
- first intensity data comprising a combined intensity of a first signal component obtained based upon a respective first nucleobase at a first portion of the template sequence and a second signal component obtained based upon a respective second nucleobase at a second portion of the template complement sequence, wherein the first and second signal components are obtained simultaneously;
- each classification of the plurality of classifications represents one or more possible combinations of respective first and second nucleobases, and wherein at least one classification of the plurality of classifications represents more than one possible combination of respective first and second nucleobases;
- selecting the classification based on the first and second intensity data may comprise selecting the classification based on the combined intensity of the first and second signal components and the combined intensity of the third and fourth signal components.
- an intensity of the first signal component when based on a nucleobase of the same identity, may be substantially the same as an intensity of the second signal component and an intensity of the third signal component is substantially the same as an intensity of the fourth signal component.
- the plurality of classifications may consist of a predetermined number of classifications.
- the plurality of classifications may comprise: one or more classifications representing matching first and second nucleobases; and one or more classifications representing mismatching first and second nucleobases, and wherein determining sequence information of the first portion and second portion comprises: in response to selecting a classification representing matching first and second nucleobases, determining a match between the first and second nucleobases; or in response to selecting a classification representing mismatching first and second nucleobases, determining a mismatch between the first and second nucleobases.
- determining sequence information of the first portion and the second portion may comprise, in response to selecting a classification representing a match between the first and second nucleobases, base calling the first and second nucleobases.
- determining sequence information of the first portion and the second portion may comprise, based on the selected classification, determining that the second portion is modified relative to the first portion at a location associated with the first and second nucleobases.
- the first signal component, second signal component, third signal component and fourth signal component may be generated based on light emissions associated with the respective nucleobase.
- the light emissions may be detected by a sensor, wherein the sensor is configured to provide a single output based upon the first and second signals.
- the senor may comprise a single sensing element.
- the method may further comprise repeating steps (a) to (d) for each of a plurality of base calling cycles.
- Methods as described herein may be performed by a user physically.
- a user may themselves conduct the methods of preparing polynucleotide sequences for identification as described herein, and as such the methods as described herein may not need to be computer- implemented.
- a kit comprising a solid support as described herein.
- kits comprising instructions for preparing polynucleotide sequences for identification according to the methods described herein and/or sequencing polynucleotide sequences according to the methods described herein.
- methods as described herein may be performed by a computer.
- a computer may contain instructions to conduct the methods of preparing polynucleotide sequences for identification as described herein, and as such the methods as described herein may be computer-implemented.
- a data processing device comprising means for carrying out the methods as described herein.
- the data processing device may be a polynucleotide sequencer.
- the data processing device may comprise reagents used for methods as described herein.
- the data processing device may comprise a solid support as described herein, such as a flow cell.
- a computer program product comprising instructions which, when the program is executed by a processor, cause the processor to carry out the methods as described herein.
- a computer-readable storage medium comprising instructions which, when executed by a processor, cause the processor to carry out the methods as described herein.
- a computer-readable data carrier having stored thereon the computer program product as described herein.
- a data carrier signal carrying the computer program product as described herein.
- the various illustrative imaging or data processing techniques described in connection with the embodiments disclosed herein can be implemented as electronic hardware, computer software, or combinations of both.
- various illustrative components, blocks, modules, and steps have been described above generally in terms of their functionality. Whether such functionality is implemented as hardware or software depends upon the particular application and design constraints imposed on the overall system.
- the described functionality can be implemented in varying ways for each particular application, but such implementation decisions should not be interpreted as causing a departure from the scope of the disclosure.
- DSP digital signal processor
- ASIC application specific integrated circuit
- FPGA field programmable gate array
- a processor can be a microprocessor, but in the alternative, the processor can be a controller, microcontroller, or state machine, combinations of the same, or the like.
- a processor can also be implemented as a combination of computing devices, e.g., a combination of a DSP and a microprocessor, a plurality of microprocessors, one or more microprocessors in conjunction with a DSP core, or any other such configuration.
- a processor can also be implemented as a combination of computing devices, e.g., a combination of a DSP and a microprocessor, a plurality of microprocessors, one or more microprocessors in conjunction with a DSP core, or any other such configuration.
- systems described herein may be implemented using a discrete memory chip, a portion of memory in a microprocessor, flash, EPROM, or other types of memory.
- a software module can reside in RAM memory, flash memory, ROM memory, EPROM memory, EEPROM memory, registers, hard disk, a removable disk, a CD-ROM, or any other form of computer-readable storage medium known in the art.
- An exemplary storage medium can be coupled to the processor such that the processor can read information from, and write information to, the storage medium. In the alternative, the storage medium can be integral to the processor.
- the processor and the storage medium can reside in an ASIC.
- a software module can comprise computer-executable instructions which cause a hardware processor to execute the computer-executable instructions. Computer-executable instructions may be stored in a (transitory or non-transitory) computer readable storage medium (e.g., memory, storage system, etc.) storing code, or computer readable instructions.
- Disjunctive language such as the phrase “at least one of X, Y or Z,” unless specifically stated otherwise, is otherwise understood with the context as used in general to present that an item, term, etc., may be either X, Y or Z, or any combination thereof (e.g., X, Y and/or Z). Thus, such disjunctive language is not generally intended to, and should not, imply that certain embodiments require at least one of X, at least one of Y or at least one of Z to each be present.
- the terms “about” or “approximate” and the like are synonymous and are used to indicate that the value modified by the term has an understood range associated with it, where the range can be ⁇ 20%, ⁇ 15%, ⁇ 10%, ⁇ 5%, or ⁇ 1%.
- the term “substantially” is used to indicate that a result (e.g., measurement value) is close to a targeted value, where close can mean, for example, the result is within 80% of the value, within 90% of the value, within 95% of the value, or within 99% of the value.
- the term “partially” is used to indicate that an effect is only in part or to a limited extent.
- a device configured to or “a device to” are intended to include one or more recited devices.
- Such one or more recited devices can also be collectively configured to carry out the stated recitations.
- a processor to carry out recitations A, B and C can include a first processor configured to carry out recitation A working in conjunction with a second processor configured to carry out recitations B and C.
- X denotes alkene-dT linkage, which can be linearised with 6mM palladium in THP cleavage mix: wherein represents an attachment point of the nucleotide to the rest of the oligo.
- a 2 1 ratio used oligos at 1.5pM each for the more concentrated pair, 0.75pM each for the less concentrated pair.
- a 1 1 ratio used all 4 oligos at 0.75pM in the grafting mix.
- Flowcell lanes were seeded with 5pM of a TruSeq Nano E. coli library and then bridge amplified for either 20 or 30 cycles of initial amplification to establish the cluster R1 “centres”. Differing numbers of amplification cycles were achieved by programming the cBot script to pump the different cycles of amplification from different positions of the cBot reagent plate. For instance, cycles 1 to 20 may pump the reagents LDR (formamide), LPM (water) and AMX (Bst amplification mix) from positions 5, 3 and 4 on the reagent plate, then cycles 21 to 30 would pump those reagents from positions 9, 2 and 6. For lanes which were kept at the 20 cycles amplification stage, those later reagent positions were filled with HT2 wash buffer.
- LDR formamide
- LPM water
- AMX Bst amplification mix
- lanes 2-5, 7 and 8 were treated with BMX (Blocking mix, a mixture of terminal transferase, a sequencing polymerase and ddNTPs) for 30mins at 37 °C and then 15mins at 60 °C.
- Lanes 1 and 6 had HT2 wash buffer during this step to be a “no blocking” test. This treatment blocks any free 3 ’OH ends in the lanes.
- Example 3 Sequencing using solid support with ringed region and central region
- a solid support comprising: a plurality of first immobilised primers, and a plurality of second immobilised primers, wherein a first proportion of the first immobilised primers are configured to be cleavable under first cleavage conditions and are unblocked, wherein a second proportion of the first immobilised primers are configured to be non-cleavable under first cleavage conditions and are blocked, wherein a third proportion of the second immobilised primers are configured to be cleavable under second cleavage conditions and are blocked; and wherein a fourth proportion of the second immobilised primers are configured to be non-cleavable under second cleavage conditions and are unblocked.
- Clause 6. A solid support according to any one of clauses 1 to 5, wherein the first immobilised primers of the second proportion that are blocked each comprise a blocking group at a 3 ’ end of the first immobilised primer.
- the blocking group is selected from the group consisting of: a hairpin loop, a deoxynucleotide, a deoxyribonucleotide, a hydrogen atom instead of a 3 ’-OH group, a phosphate group, a phosphorothioate group, a propyl spacer, a modification blocking the 3 ’-hydroxyl group, or an inverted nucleobase.
- Clause 8 A solid support according to clause 7, wherein the blocking group is a phosphate group or a hairpin loop.
- Clause 10 A solid support according to any one of clauses 1 to 9, wherein the second immobilised primers of the third proportion that are blocked each comprise a blocking group at a 3 ’ end of the second immobilised primer.
- a solid support according to clause 10, wherein the blocking group is selected from the group consisting of: a hairpin loop, a deoxynucleotide, a deoxyribonucleotide, a hydrogen atom instead of a 3 ’-OH group, a phosphate group, a phosphorothioate group, a propyl spacer, a modification blocking the 3 ’-hydroxyl group, or an inverted nucleobase.
- Clause 14 A solid support according to any one of clauses 1 to 13, wherein the first immobilised primers of the first proportion are configured to be cleavable by a thermal trigger, a light trigger, and/or a chemical/biochemical trigger.
- Clause 16 A solid support according to clause 15, wherein the first immobilised primers of the first proportion are configured to be cleavable by a transition metal catalyst.
- Clause 17 A solid support according to clause 16, wherein the first immobilised primers of the first proportion are configured to be cleavable by a palladium-based or a nickel-based catalyst.
- Clause 18 A solid support according to clause 17, wherein the first immobilised primers of the first proportion are configured to be cleavable by a palladium-based catalyst.
- each first immobilised primer of the first proportion comprises a nucleotide that comprises an allyl group.
- Clause 20 A solid support according to clause 19, wherein the allyl group is attached to a sugar group of the nucleotide.
- nucleotide has a structure according to the following: se wherein represents an attachment point of the nucleotide to the rest of the first immobilised primer of the first proportion, and wherein “base” represents adenine, guanine, cytosine, thymine or uracil, or a derivative thereof.
- base represents adenine, guanine, cytosine, thymine or uracil, or a derivative thereof.
- Clause 24 A solid support according to clause 14, wherein the first immobilised primers of the first proportion are configured to be cleavable by a glycosylase.
- each first immobilised primer of the first proportion comprises a nucleobase which is not selected from guanine, cytosine, adenine or thymine when the first immobilised primer is a DNA sequence; or wherein each first immobilised primer of the first proportion comprises a nucleobase which is not selected from guanine, cytosine, adenine or uracil when the first immobilised primer is an RNA sequence.
- each first immobilised primer of the first proportion comprises oxoguanine (e.g. 8-oxoguanine) or uracil when the first immobilised primer is a DNA sequence, or wherein each first immobilised primer of the first proportion comprises oxoguanine (e.g. 8-oxoguanine) when the first immobilised primer is an RNA sequence.
- oxoguanine e.g. 8-oxoguanine
- uracil when the first immobilised primer is a DNA sequence
- each first immobilised primer of the first proportion comprises oxoguanine (e.g. 8-oxoguanine) when the first immobilised primer is an RNA sequence.
- Clause 27 A solid support according to any one of clauses 1 to 26, wherein the second immobilised primers of the third proportion are configured to be cleavable by a thermal trigger, a light trigger, or a chemical/biochemical trigger.
- Clause 28 A solid support according to clause 27, wherein the second immobilised primers of the third proportion are configured to be cleavable by a metal catalyst.
- Clause 29 A solid support according to clause 28, wherein the second immobilised primers of the third proportion are configured to be cleavable by a transition metal catalyst.
- Clause 30 A solid support according to clause 29, wherein the second immobilised primers of the third proportion are configured to be cleavable by a palladium-based or a nickel-based catalyst.
- Clause 31 A solid support according to clause 30, wherein the second immobilised primers of the third proportion are configured to be cleavable by a palladium-based catalyst.
- each second immobilised primer of the third proportion comprises a nucleotide that comprises an allyl group.
- Clause 35 A solid support according to clause 34, wherein the nucleotide has a structure according to the following: wherein represents an attachment point of the nucleotide to the rest of the second immobilised primer of the third proportion, and wherein “base” represents adenine, guanine, cytosine, thymine or uracil, or a derivative thereof.
- base represents adenine, guanine, cytosine, thymine or uracil, or a derivative thereof.
- Clause 37 A solid support according to clause 27, wherein the second immobilised primers of the third proportion are configured to be cleavable by a glycosylase.
- each second immobilised primer of the third proportion comprises a nucleobase which is not selected from guanine, cytosine, adenine or thymine when the second immobilised primer is a DNA sequence; or wherein each second immobilised primer of the third proportion comprises a nucleobase which is not selected from guanine, cytosine, adenine or uracil when the second immobilised primer is an RNA sequence.
- each second immobilised primer of the third proportion comprises oxoguanine (e.g. 8-oxoguanine) or uracil when the second immobilised primer is a DNA sequence, or wherein each second immobilised primer of the third proportion comprises oxoguanine (e.g. 8-oxoguanine) when the second immobilised primer is an RNA sequence.
- oxoguanine e.g. 8-oxoguanine
- uracil when the second immobilised primer is a DNA sequence
- each second immobilised primer of the third proportion comprises oxoguanine (e.g. 8-oxoguanine) when the second immobilised primer is an RNA sequence.
- Clause 40 A solid support according to any one of clauses 1 to 39, wherein the first cleavage conditions and the second cleavage conditions are the same or are different.
- Clause 41 A solid support according to clause 40, wherein the first cleavage conditions and the second cleavage conditions are the same.
- Clause 42 A solid support according to any one of clauses 1 to 41, wherein the first proportion of first immobilised primers that are cleavable are further configured to be linearisable under first linearisation conditions.
- Clause 43 A solid support according to any one of clauses 1 to 42, wherein the third proportion of second immobilised primers that are cleavable are further configured to be linearisable under second linearisation conditions.
- Clause 44 A solid support according to clause 43 as dependent on clause 42, wherein the first linearisation conditions and the second linearisation conditions are the same or are different.
- Clause 45 A solid support according to clause 44, wherein the first linearisation conditions and the second linearisation conditions are the same.
- each first immobilised primer comprises a sequence as defined in SEQ ID NO. 1 or 5, or a variant or fragment thereof; and each second immobilised primer comprises a sequence as defined in SEQ ID NO. 2, or a variant or fragment thereof; or wherein each first immobilised primer comprises a sequence as defined in SEQ ID NO. 2, or a variant or fragment thereof; and each second immobilised primer comprises a sequence as defined in SEQ ID NO. 1 or 5, or a variant or fragment thereof.
- a solid support comprising: a plurality of first immobilised primers, and a plurality of second immobilised primers, wherein the plurality of first immobilised primers are located in a first region, wherein the plurality of second immobilised primers are located in a second region surrounding the first region.
- Clause 48 A solid support according to clause 47, wherein the second region is contiguous.
- Clause 50 A solid support according to any one of clauses 47 to 49, wherein a concentration of first immobilised primers in the first region is greater than a concentration of second immobilised primers in the second region.
- Clause 51 A solid support according to clause 50, wherein a ratio between the concentration of first immobilised primers in the first region and the concentration of second immobilised primers in the second region is between 1.25: 1 to 5: 1.
- Clause 52 A solid support according to clause 51, wherein the ratio between the concentration of first immobilised primers in the first region and the concentration of second immobilised primers in the second region is between 1.5 : 1 to 3 : 1.
- Clause 53 A solid support according to clause 52, wherein the ratio between the concentration of first immobilised primers in the first region and the concentration of second immobilised primers in the second region is about 2: 1.
- Clause 54 A solid support according to any one of clauses 47 to 49, wherein a concentration of first immobilised primers in the first region is less than a concentration of second immobilised primers in the second region.
- Clause 55 A solid support according to clause 54, wherein a ratio between the concentration of first immobilised primers in the first region and the concentration of second immobilised primers in the second region is between 1: 1.25 to 1:5.
- Clause 56 A solid support according to clause 55, wherein the ratio between the concentration of first immobilised primers in the first region and the concentration of second immobilised primers in the second region is between 1: 1.5 to 1:3.
- Clause 57 A solid support according to clause 56, wherein the ratio between the concentration of first immobilised primers in the first region and the concentration of second immobilised primers in the second region is about 1:2.
- Clause 58 A solid support according to any one of clauses 47 to 49, wherein a concentration of first immobilised primers in the first region is equal or substantially equal to a concentration of second immobilised primers in the second region.
- Clause 59 A solid support according to any one of clauses 1 to 58, wherein the solid support is a flow cell.
- Clause 60 A kit comprising a solid support according to any one of clauses 1 to 59.
- Clause 61 Use of a solid support according to any one of clauses 1 to 59 in nucleic acid sequencing.
- a process of manufacturing a solid support comprising:
- Clause 63 A process according to clause 62, wherein steps (a) and (b) are conducted sequentially or simultaneously.
- Clause 64 A process according to clause 63, wherein step (b) is conducted after step (a).
- Clause 65 A process according to clause 63, wherein step (a) is conducted after step (b).
- Clause 66 A process according to clause 63, wherein steps (a) and (b) are conducted simultaneously.
- Clause 67 A process according to any one of clauses 62 to 66, wherein immobilisation comprises forming covalent linkages between the solid support and each of the plurality of first precursor primers, and between the solid support and each of the plurality of second precursor primers.
- Clause 68 A process according to clause 67, wherein forming covalent linkages involves using a click reaction.
- Clause 69 A process according to clause 67 or clause 68, wherein forming covalent linkages involves forming a 1,2, 3 -triazole linkage.
- Clause 70 A method of preparing polynucleotide sequences for identification, comprising: providing a solid support according to any one of clauses 1 to 46, and synthesising a plurality of template sequences that extend from the second immobilised primers in the fourth proportion and a plurality of template complement sequences that extend from the first immobilised primers in the first proportion.
- Clause 71 A method according to clause 70, wherein the step of synthesising the plurality of template sequences extending from the second immobilised primers in the fourth proportion and the plurality of template complement sequences extending from the first immobilised primers in the first proportion involves conducting amplification.
- Clause 72 A method according to clause 71, wherein amplification is bridge amplification.
- Clause 73 A method according to clause 71 or clause 72, wherein amplification is conducted over 20 to 40 cycles.
- Clause 74 A method according to clause 73, wherein amplification is conducted over 25 to 35 cycles.
- Clause 75 A method according to any one of clauses 70 to 74, further comprising a step of removing second immobilised primers in the fourth proportion that are not yet extended, and removing first immobilised primers in the first proportion that are not yet extended.
- Clause 76 A method according to clause 75, wherein the step of removing second immobilised primers in the fourth proportion that are not yet extended and removing first immobilised primers in the first proportion that are not yet extended is conducted using a 3 ’-5’ exonuclease (e.g. Exol).
- a 3 ’-5’ exonuclease e.g. Exol
- Clause 77 A method according to clause 75 or clause 76, further comprising a step of unblocking the second proportion of first immobilised primers and unblocking the third proportion of second immobilised primers.
- Clause 78 A method according to clause 77, further comprising a step of synthesising a plurality of template sequences that extend from the second immobilised primers in the third proportion and a plurality of template complement sequences that extend from the first immobilised primers in the second proportion.
- Clause 79 A method according to clause 78, wherein the step of synthesising the plurality of template sequences extending from the second immobilised primers in the third proportion and the plurality of template complement sequences extending from the first immobilised primers in the second proportion involves conducting amplification.
- Clause 80 A method according to clause 79, wherein amplification is bridge amplification.
- Clause 81 A method according to clause 79 or clause 80, wherein amplification is conducted over 5 to 25 cycles.
- Clause 82 A method according to clause 81, wherein amplification is conducted over 10 to 20 cycles.
- Clause 83 A method according to clause 79 or clause 80, wherein the number of amplification cycles used for synthesising the plurality of template sequences extending from the second immobilised primers in the third proportion and the plurality of template complement sequences extending from the first immobilised primers in the second proportion is less than the number of amplification cycles used for synthesising the plurality of template sequences extending from the second immobilised primers in the fourth proportion and the plurality of template complement sequences extending from the first immobilised primers in the first proportion.
- Clause 84 A method according to clause 79 or clause 80, wherein the number of amplification cycles used for synthesising the plurality of template sequences extending from the second immobilised primers in the third proportion and the plurality of template complement sequences extending from the first immobilised primers in the second proportion is more than the number of amplification cycles used for synthesising the plurality of template sequences extending from the second immobilised primers in the fourth proportion and the plurality of template complement sequences extending from the first immobilised primers in the first proportion.
- Clause 85 A method according to clause 79 or clause 80, wherein the number of amplification cycles used for synthesising the plurality of template sequences extending from the second immobilised primers in the third proportion and the plurality of template complement sequences extending from the first immobilised primers in the second proportion is the same as the number of amplification cycles used for synthesising the plurality of template sequences extending from the second immobilised primers in the fourth proportion and the plurality of template complement sequences extending from the first immobilised primers in the first proportion.
- Clause 86 A method according to any one of clauses 78 to 85, further comprising a step of removing second immobilised primers in the third proportion that are not yet extended, and removing first immobilised primers in the second proportion that are not yet extended.
- Clause 87 A method according to clause 86, wherein the step of removing second immobilised primers in the third proportion that are not yet extended and removing first immobilised primers in the second proportion that are not yet extended is conducted using a 3 ’-5’ exonuclease (e.g. Exol).
- a 3 ’-5’ exonuclease e.g. Exol
- Clause 88 A method according to any one of clauses 78 to 87, further comprising a step of exposing the solid support to first cleavage conditions and/or second cleavage conditions.
- Clause 89 A method according to clause 88, wherein the first cleavage conditions and/or second cleavage conditions comprise exposure to a thermal trigger, a light trigger, and/or a chemical/biochemical trigger.
- Clause 90 A method according to clause 89, wherein the solid support is exposed to a metal catalyst.
- Clause 91 A method according to clause 90, wherein the solid support is exposed to a transition metal catalyst.
- Clause 92 A solid support according to clause 91, wherein the solid support is exposed to a palladium-based or a nickel-based catalyst.
- Clause 93 A solid support according to clause 92, wherein the solid support is exposed to a palladium-based catalyst.
- Clause 94 A method according to clause 89, wherein the solid support is exposed to a glycosylase.
- Clause 95 A method according to any one of clauses 78 to 94, wherein the method further comprises a step of linearising the plurality of template sequences extending from the second immobilised primers of the fourth proportion and linearising the plurality of template complement sequences extending from the first immobilised primers of the second proportion.
- Clause 96 A method according to clause 95, wherein the method further comprises treating the linearised template sequences and the linearised template complement sequences with a singlestranded binding protein.
- Clause 97 A method of sequencing polynucleotide sequences, comprising: preparing polynucleotide sequences for identification using a method according to any one of clauses 70 to 96; and concurrently sequencing nucleobases in the template sequences extending from the second immobilised primers of the fourth proportion and the template complement sequences extending from the first immobilised primers of the second proportion.
- Clause 98 A method according to clause 97, wherein the step of concurrently sequencing nucleobases comprises performing sequencing-by-synthesis or sequencing-by-ligation.
- Clause 99 A method according to clause 97 or clause 98, wherein the step of concurrently sequencing nucleobases comprises treatment with a strand displacement polymerase (e.g. phi29).
- a strand displacement polymerase e.g. phi29
- Clause 100 A method according to any one of clauses 97 to 99, wherein the method further comprises a step of conducting paired-end reads.
- a kit comprising instructions for preparing polynucleotide sequences for identification according to any one of clauses 70 to 96; and/or sequencing polynucleotide sequences according to any one of clauses 97 to 100.
- Clause 102 A data processing device comprising means for carrying out a method according to any one of clauses 70 to 100.
- Clause 103 A data processing device according to clause 102, wherein the data processing device is a polynucleotide sequencer.
- Clause 104 A computer program product comprising instructions which, when the program is executed by a processor, cause the processor to carry out a method according to any one of clauses 70 to 100.
- Clause 105 A computer-readable storage medium comprising instructions which, when executed by a processor, cause the processor to carry out a method according to any one of clauses 70 to 100.
- Clause 106. A computer-readable data carrier having stored thereon a computer program product according to clause 104.
- Clause 107 A data carrier signal carrying a computer program product according to clause 104.
- SEQ ID NO. 2 P7 sequence
- SEQ ID NO. 4 P7’ sequence (complementary to P7)
- SEQ ID NO. 7 SBS3
- SEQ ID NO. 9 SBS12
- X denotes alkene-dT linkage: wherein represents an attachment point of the nucleotide to the rest of the oligo.
- ‘X” denotes alkene-dT linkage: wherein represents an attachment point of the nucleotide to the rest of the oligo.
- alkyne -T TTTTTCAAG C AG AAG AC GG CAT AC GAG AT
Landscapes
- Chemical & Material Sciences (AREA)
- Organic Chemistry (AREA)
- Life Sciences & Earth Sciences (AREA)
- Analytical Chemistry (AREA)
- Zoology (AREA)
- Wood Science & Technology (AREA)
- Proteomics, Peptides & Aminoacids (AREA)
- Health & Medical Sciences (AREA)
- Engineering & Computer Science (AREA)
- Biophysics (AREA)
- Immunology (AREA)
- Microbiology (AREA)
- Molecular Biology (AREA)
- Biotechnology (AREA)
- Physics & Mathematics (AREA)
- Chemical Kinetics & Catalysis (AREA)
- Biochemistry (AREA)
- Bioinformatics & Cheminformatics (AREA)
- General Engineering & Computer Science (AREA)
- General Health & Medical Sciences (AREA)
- Genetics & Genomics (AREA)
- Measuring Or Testing Involving Enzymes Or Micro-Organisms (AREA)
Abstract
The invention relates to solid supports and methods for use in nucleic acid sequencing, in particular solid supports and methods for use in concurrent sequencing.
Description
CONCURRENT SEQUENCING WITH SPATIAEEY SEPARATED RINGS
Field of the Invention
The invention relates to solid supports and methods for use in nucleic acid sequencing, in particular solid supports and methods for use in concurrent sequencing.
Background of the Invention
In some types of next-generation sequencing (NGS) technologies, a nucleic acid cluster is created on a flow cell by amplifying an original template nucleic acid strand. Sequencing cycles may be performed as complementary strands of the template nucleic acids are being synthesized, i.e., using sequencing-by-synthesis (SBS) processes.
In each sequencing cycle, deoxyribonucleic acid analogs conjugated to fluorescent labels are hybridized to the template nucleic acids, and excitation light sources are used to excite the fluorescent labels on the deoxyribonucleic acid analogs. Detectors capture fluorescent emissions from the fluorescent labels and identify the deoxyribonucleic acid analogs. As a result, the sequence of the template nucleic acids may be determined by repeatedly performing such sequencing cycles.
NGS allows for the sequencing of a number of different template nucleic acids simultaneously, which has significantly reduced the cost of sequencing in the last twenty years. However, there remains a desire for further improvements in sequencing throughput and speed.
Summary of the Invention
According to an aspect of the present invention, there is provided a solid support, comprising: a plurality of first immobilised primers, and a plurality of second immobilised primers, wherein a first proportion of the first immobilised primers are configured to be cleavable under first cleavage conditions and are unblocked, wherein a second proportion of the first immobilised primers are configured to be non-cleavable under first cleavage conditions and are blocked, wherein a third proportion of the second immobilised primers are configured to be cleavable under second cleavage conditions and are blocked; and
wherein a fourth proportion of the second immobilised primers are configured to be non-cleavable under second cleavage conditions and are unblocked.
In one aspect, a ratio between unblocked immobilised primers and blocked immobilised primers is between 20:80 to 80:20.
In one aspect, the ratio between unblocked immobilised primers and blocked immobilised primers is between 50:50 to 75:25.
In one aspect, the ratio between unblocked immobilised primers and blocked immobilised primers is between 60:40 to 70:30.
In one aspect, the ratio between unblocked immobilised primers and blocked immobilised primers is about 2: 1.
In one aspect, the first immobilised primers of the second proportion that are blocked each comprise a blocking group at a 3 ’ end of the first immobilised primer.
In one aspect, the blocking group is selected from the group consisting of: a hairpin loop, a deoxynucleotide, a deoxyribonucleotide, a hydrogen atom instead of a 3 ’-OH group, a phosphate group, a phosphorothioate group, a propyl spacer, a modification blocking the 3 ’-hydroxyl group, or an inverted nucleobase.
In one aspect, the blocking group is a phosphate group or a hairpin loop.
In one aspect, the blocking group is a phosphate group.
In one aspect, the second immobilised primers of the third proportion that are blocked each comprise a blocking group at a 3 ’ end of the second immobilised primer.
In one aspect, the blocking group is selected from the group consisting of: a hairpin loop, a deoxynucleotide, a deoxyribonucleotide, a hydrogen atom instead of a 3 ’-OH group, a phosphate group, a phosphorothioate group, a propyl spacer, a modification blocking the 3 ’-hydroxyl group, or an inverted nucleobase.
In one aspect, the blocking group is a phosphate group or a hairpin loop.
In one aspect, the blocking group is a phosphate group.
In one aspect, the first immobilised primers of the first proportion are configured to be cleavable by a thermal trigger, a light trigger, and/or a chemical/biochemical trigger.
In one aspect, the first immobilised primers of the first proportion are configured to be cleavable by a metal catalyst.
In one aspect, the first immobilised primers of the first proportion are configured to be cleavable by a transition metal catalyst.
In one aspect, the first immobilised primers of the first proportion are configured to be cleavable by a palladium-based or a nickel-based catalyst.
In one aspect, the first immobilised primers of the first proportion are configured to be cleavable by a palladium-based catalyst.
In one aspect, each first immobilised primer of the first proportion comprises a nucleotide that comprises an allyl group.
In one aspect, the allyl group is attached to a sugar group of the nucleotide.
In one aspect, the allyl group has a structure according to the following:
wherein R represents an attachment point to hydrogen (in which case, R is hydrogen) or the remainder of the sugar group.
In one aspect, the nucleotide has a structure according to the following:
wherein represents an attachment point of the nucleotide to the rest of the first immobilised primer of the first proportion, and wherein “base” represents adenine, guanine, cytosine, thymine or uracil, or a derivative thereof.
In one aspect, the nucleotide has a structure according to the following:
wherein represents an attachment point of the nucleotide to the rest of the first immobilised primer of the first proportion.
In one aspect, the first immobilised primers of the first proportion are configured to be cleavable by a glycosylase.
In one aspect, each first immobilised primer of the first proportion comprises a nucleobase which is not selected from guanine, cytosine, adenine or thymine when the first immobilised primer is a DNA sequence; or wherein each first immobilised primer of the first proportion comprises a nucleobase which is not selected from guanine, cytosine, adenine or uracil when the first immobilised primer is an RNA sequence.
In one aspect, each first immobilised primer of the first proportion comprises oxoguanine (e.g. 8- oxoguanine) or uracil when the first immobilised primer is a DNA sequence, or wherein each first immobilised primer of the first proportion comprises oxoguanine (e.g. 8-oxoguanine) when the first immobilised primer is an RNA sequence.
In one aspect, the second immobilised primers of the third proportion are configured to be cleavable by a thermal trigger, a light trigger, or a chemical/biochemical trigger.
In one aspect, the second immobilised primers of the third proportion are configured to be cleavable by a metal catalyst.
In one aspect, the second immobilised primers of the third proportion are configured to be cleavable by a transition metal catalyst.
In one aspect, the second immobilised primers of the third proportion are configured to be cleavable by a palladium-based or a nickel-based catalyst.
In one aspect, the second immobilised primers of the third proportion are configured to be cleavable by a palladium-based catalyst.
In one aspect, each second immobilised primer of the third proportion comprises a nucleotide that comprises an allyl group.
In one aspect, the allyl group is attached to a sugar group of the nucleotide.
In one aspect, the allyl group has a structure according to the following:
wherein R represents an attachment point to hydrogen (in which case, R is hydrogen) or the remainder of the sugar group.
In one aspect, the nucleotide has a structure according to the following:
wherein represents an attachment point of the nucleotide to the rest of the second immobilised primer of the third proportion, and wherein “base” represents adenine, guanine, cytosine, thymine or uracil, or a derivative thereof.
In one aspect, the nucleotide has a structure according to the following:
wherein represents an attachment point of the nucleotide to the rest of the second immobilised primer of the third proportion.
In one aspect, the second immobilised primers of the third proportion are configured to be cleavable by a glycosylase.
In one aspect, each second immobilised primer of the third proportion comprises a nucleobase which is not selected from guanine, cytosine, adenine or thymine when the second immobilised primer is a DNA sequence; or wherein each second immobilised primer of the third proportion comprises a nucleobase which is not selected from guanine, cytosine, adenine or uracil when the second immobilised primer is an RNA sequence.
In one aspect, each second immobilised primer of the third proportion comprises oxoguanine (e.g. 8-oxoguanine) or uracil when the second immobilised primer is a DNA sequence, or wherein each second immobilised primer of the third proportion comprises oxoguanine (e.g. 8-oxoguanine) when the second immobilised primer is an RNA sequence.
In one aspect, the first cleavage conditions and the second cleavage conditions are the same or are different.
In one aspect, the first cleavage conditions and the second cleavage conditions are the same.
In one aspect, the first proportion of first immobilised primers that are cleavable are further configured to be linearisable under first linearisation conditions.
In one aspect, the third proportion of second immobilised primers that are cleavable are further configured to be linearisable under second linearisation conditions.
In one aspect, the first linearisation conditions and the second linearisation conditions are the same or are different.
In one aspect, the first linearisation conditions and the second linearisation conditions are the same.
In one aspect, each first immobilised primer comprises a sequence as defined in SEQ ID NO. 1 or 5, or a variant or fragment thereof; and each second immobilised primer comprises a sequence as defined in SEQ ID NO. 2, or a variant or fragment thereof; or wherein each first immobilised primer comprises a sequence as defined in SEQ ID NO. 2, or a variant or fragment thereof; and each second immobilised primer comprises a sequence as defined in SEQ ID NO. 1 or 5, or a variant or fragment thereof.
According to another aspect of the present invention, there is provided a solid support, comprising: a plurality of first immobilised primers, and a plurality of second immobilised primers, wherein the plurality of first immobilised primers are located in a first region, wherein the plurality of second immobilised primers are located in a second region surrounding the first region.
In one aspect, the second region is contiguous.
In one aspect, the second region is an annular region.
In one aspect, a concentration of first immobilised primers in the first region is greater than a concentration of second immobilised primers in the second region.
In one aspect, a ratio between the concentration of first immobilised primers in the first region and the concentration of second immobilised primers in the second region is between 1.25: 1 to 5: 1.
In one aspect, the ratio between the concentration of first immobilised primers in the first region and the concentration of second immobilised primers in the second region is between 1.5 : 1 to 3 : 1.
In one aspect, the ratio between the concentration of first immobilised primers in the first region and the concentration of second immobilised primers in the second region is about 2: 1.
In one aspect, a concentration of first immobilised primers in the first region is less than a concentration of second immobilised primers in the second region.
In one aspect, a ratio between the concentration of first immobilised primers in the first region and the concentration of second immobilised primers in the second region is between 1: 1.25 to 1:5.
In one aspect, the ratio between the concentration of first immobilised primers in the first region and the concentration of second immobilised primers in the second region is between 1 : 1.5 to 1:3.
In one aspect, the ratio between the concentration of first immobilised primers in the first region and the concentration of second immobilised primers in the second region is about 1:2.
In one aspect, a concentration of first immobilised primers in the first region is equal or substantially equal to a concentration of second immobilised primers in the second region.
In one aspect, the solid support is a flow cell.
According to another aspect of the present invention, there is provided a kit comprising a solid support as described herein.
According to another aspect of the present invention, there is provided a use of a solid support as described herein in nucleic acid sequencing.
According to another aspect of the present invention, there is provided a process of manufacturing a solid support, comprising:
(a) immobilising a plurality of first precursor primers onto a solid support to form a plurality of first immobilised primers, wherein a first proportion of the first precursor primers are configured to be cleavable under first cleavage conditions and are unblocked, and wherein a second proportion of the first precursor primers are configured to be non-cleavable under first cleavage conditions and are blocked; and
(b) immobilising a plurality of second precursor primers onto the solid support to form a plurality of second immobilised primers, wherein a third proportion of the second precursor primers are configured to be cleavable under second cleavage conditions and are blocked, and wherein a fourth proportion of the second precursor primers are configured to be non-cleavable under second cleavage conditions and are unblocked.
In one aspect, steps (a) and (b) are conducted sequentially or simultaneously.
In one aspect, step (b) is conducted after step (a).
In one aspect, step (a) is conducted after step (b).
In one aspect, steps (a) and (b) are conducted simultaneously.
In one aspect, immobilisation comprises forming covalent linkages between the solid support and each of the plurality of first precursor primers, and between the solid support and each of the plurality of second precursor primers.
In one aspect, forming covalent linkages involves using a click reaction.
In one aspect, forming covalent linkages involves forming a 1,2, 3 -triazole linkage.
According to another aspect of the present invention, there is provided a method of preparing polynucleotide sequences for identification, comprising: providing a solid support as described herein, and synthesising a plurality of template sequences that extend from the second immobilised primers in the fourth proportion and a plurality of template complement sequences that extend from the first immobilised primers in the first proportion.
In one aspect, the step of synthesising the plurality of template sequences extending from the second immobilised primers in the fourth proportion and the plurality of template complement sequences extending from the first immobilised primers in the first proportion involves conducting amplification.
In one aspect, amplification is bridge amplification.
In one aspect, amplification is conducted over 20 to 40 cycles.
In one aspect, amplification is conducted over 25 to 35 cycles.
In one aspect, the method further comprises a step of removing second immobilised primers in the fourth proportion that are not yet extended, and removing first immobilised primers in the first proportion that are not yet extended.
In one aspect, the step of removing second immobilised primers in the fourth proportion that are not yet extended and removing first immobilised primers in the first proportion that are not yet extended is conducted using a 3 ’-5’ exonuclease (e.g. Exol).
In one aspect, the method further comprises a step of unblocking the second proportion of first immobilised primers and unblocking the third proportion of second immobilised primers.
In one aspect, the method further comprises a step of synthesising a plurality of template sequences that extend from the second immobilised primers in the third proportion and a plurality of template complement sequences that extend from the first immobilised primers in the second proportion.
In one aspect, the step of synthesising the plurality of template sequences extending from the second immobilised primers in the third proportion and the plurality of template complement sequences extending from the first immobilised primers in the second proportion involves conducting amplification.
In one aspect, amplification is bridge amplification.
In one aspect, amplification is conducted over 5 to 25 cycles.
In one aspect, amplification is conducted over 10 to 20 cycles.
In one aspect, the number of amplification cycles used for synthesising the plurality of template sequences extending from the second immobilised primers in the third proportion and the plurality of template complement sequences extending from the first immobilised primers in the second proportion is less than the number of amplification cycles used for synthesising the plurality of template sequences extending from the second immobilised primers in the fourth proportion and the plurality of template complement sequences extending from the first immobilised primers in the first proportion.
In one aspect, the number of amplification cycles used for synthesising the plurality of template sequences extending from the second immobilised primers in the third proportion and the plurality
of template complement sequences extending from the first immobilised primers in the second proportion is more than the number of amplification cycles used for synthesising the plurality of template sequences extending from the second immobilised primers in the fourth proportion and the plurality of template complement sequences extending from the first immobilised primers in the first proportion.
In one aspect, the number of amplification cycles used for synthesising the plurality of template sequences extending from the second immobilised primers in the third proportion and the plurality of template complement sequences extending from the first immobilised primers in the second proportion is the same as the number of amplification cycles used for synthesising the plurality of template sequences extending from the second immobilised primers in the fourth proportion and the plurality of template complement sequences extending from the first immobilised primers in the first proportion.
In one aspect, the method further comprises a step of removing second immobilised primers in the third proportion that are not yet extended, and removing first immobilised primers in the second proportion that are not yet extended.
In one aspect, the step of removing second immobilised primers in the third proportion that are not yet extended and removing first immobilised primers in the second proportion that are not yet extended is conducted using a 3 ’-5’ exonuclease (e.g. Exol).
In one aspect, the method further comprises a step of exposing the solid support to first cleavage conditions and/or second cleavage conditions.
In one aspect, the first cleavage conditions and/or second cleavage conditions comprise exposure to a thermal trigger, a light trigger, and/or a chemical/biochemical trigger.
In one aspect, the solid support is exposed to a metal catalyst.
In one aspect, the solid support is exposed to a transition metal catalyst.
In one aspect, the solid support is exposed to a palladium-based or a nickel-based catalyst.
In one aspect, the solid support is exposed to a palladium-based catalyst.
In one aspect, the solid support is exposed to a glycosylase.
In one aspect, the method further comprises a step of linearising the plurality of template sequences extending from the second immobilised primers of the fourth proportion and linearising the plurality of template complement sequences extending from the first immobilised primers of the second proportion.
In one aspect, the method further comprises treating the linearised template sequences and the linearised template complement sequences with a single-stranded binding protein.
According to another aspect of the present invention, there is provided a method of sequencing polynucleotide sequences, comprising: preparing polynucleotide sequences for identification using a method as described herein; and concurrently sequencing nucleobases in the template sequences extending from the second immobilised primers of the fourth proportion and the template complement sequences extending from the first immobilised primers of the second proportion.
In one aspect, the step of concurrently sequencing nucleobases comprises performing sequencing- by-synthesis or sequencing-by-ligation.
In one aspect, the step of concurrently sequencing nucleobases comprises treatment with a strand displacement polymerase (e.g. phi29).
In one aspect, the method further comprises a step of conducting paired-end reads.
In one aspect, the step of concurrently sequencing nucleobases comprises:
(a) obtaining first intensity data comprising a combined intensity of a first signal component obtained based upon a respective first nucleobase at a first portion of the template sequence and a second signal component obtained based upon a respective second nucleobase at a second portion of the template complement sequence, wherein the first and second signal components are obtained simultaneously;
(b) obtaining second intensity data comprising a combined intensity of a third signal component obtained based upon the respective first nucleobase at the first portion and a fourth signal component obtained based upon the respective second nucleobase at the second portion, wherein the third and fourth signal components are obtained simultaneously;
(c) selecting one of a plurality of classifications based on the first and the second intensity data, wherein each classification represents a possible combination of respective first and second nucleobases; and
(d) based on the selected classification, base calling the respective first and second nucleobases.
In one aspect, selecting the classification based on the first and second intensity data comprises selecting the classification based on the combined intensity of the first and second signal components and the combined intensity of the third and fourth signal components.
In one aspect, the plurality of classifications comprises sixteen classifications, each classification representing one of sixteen unique combinations of first and second nucleobases.
In one aspect, the first signal component, second signal component, third signal component and fourth signal component are generated based on light emissions associated with the respective nucleobase.
In one aspect, the light emissions are detected by a sensor, wherein the sensor is configured to provide a single output based upon the first and second signals.
In one aspect, the sensor comprises a single sensing element.
In one aspect, the method further comprises repeating steps (a) to (d) for each of a plurality of base calling cycles.
In one aspect, the step of concurrently sequencing nucleobases comprises:
(a) obtaining first intensity data comprising a combined intensity of a first signal component obtained based upon a respective first nucleobase at a first portion of the template sequence and a second signal component obtained based upon a respective second nucleobase at the second portion of the template complement sequence, wherein the first and second signal components are obtained simultaneously;
(b) obtaining second intensity data comprising a combined intensity of a third signal component obtained based upon the respective first nucleobase at the first portion and a fourth signal component obtained based upon the respective second nucleobase at the second portion, wherein the third and fourth signal components are obtained simultaneously;
(c) selecting one of a plurality of classifications based on the first and the second intensity data, wherein each classification of the plurality of classifications represents one or more
possible combinations of respective first and second nucleobases, and wherein at least one classification of the plurality of classifications represents more than one possible combination of respective first and second nucleobases; and
(d) based on the selected classification, determining sequence information from the first portion and the second portion.
In one aspect, selecting the classification based on the first and second intensity data comprises selecting the classification based on the combined intensity of the first and second signal components and the combined intensity of the third and fourth signal components.
In one aspect, when based on a nucleobase of the same identity, an intensity of the first signal component is substantially the same as an intensity of the second signal component and an intensity of the third signal component is substantially the same as an intensity of the fourth signal component.
In one aspect, the plurality of classifications consists of a predetermined number of classifications.
In one aspect, the plurality of classifications comprises: one or more classifications representing matching first and second nucleobases; and one or more classifications representing mismatching first and second nucleobases, and wherein determining sequence information of the first portion and second portion comprises: in response to selecting a classification representing matching first and second nucleobases, determining a match between the first and second nucleobases; or in response to selecting a classification representing mismatching first and second nucleobases, determining a mismatch between the first and second nucleobases.
In one aspect, determining sequence information of the first portion and the second portion comprises, in response to selecting a classification representing a match between the first and second nucleobases, base calling the first and second nucleobases.
In one aspect, determining sequence information of the first portion and the second portion comprises, based on the selected classification, determining that the second portion is modified relative to the first portion at a location associated with the first and second nucleobases.
In one aspect, the first signal component, second signal component, third signal component and fourth signal component are generated based on light emissions associated with the respective nucleobase.
In one aspect, the light emissions are detected by a sensor, wherein the sensor is configured to provide a single output based upon the first and second signals.
In one aspect, the sensor comprises a single sensing element.
In one aspect, the method further comprises repeating steps (a) to (d) for each of a plurality of base calling cycles.
In one aspect, the first portion is at least 25 base pairs and the second portion is at least 25 base pairs.
According to another aspect of the present invention, there is provided a kit comprising instructions for preparing polynucleotide sequences for identification as described herein; and/or sequencing polynucleotide sequences as described herein.
According to another aspect of the present invention, there is provided a data processing device comprising means for carrying out a method as described herein.
In one aspect, the data processing device is a polynucleotide sequencer.
According to another aspect of the present invention, there is provided a computer program product comprising instructions which, when the program is executed by a processor, cause the processor to carry out a method as described herein.
According to another aspect of the present invention, there is provided a computer-readable storage medium comprising instructions which, when executed by a processor, cause the processor to carry out a method as described herein.
According to another aspect of the present invention, there is provided a computer-readable data carrier having stored thereon a computer program product as described herein.
According to another aspect of the present invention, there is provided a data carrier signal carrying a computer program product as described herein.
Brief Description of the Drawings
Features of examples of the present disclosure will become apparent by reference to the following detailed description and drawings, in which like reference numerals correspond to similar, though perhaps not identical, components. For the sake of brevity, reference numerals or features having a previously described function may or may not be described in connection with other drawings in which they appear.
Figure 1 shows a forward strand, reverse strand, forward complement strand, and reverse complement strand of a polynucleotide molecule.
Figure 2 shows an example of a polynucleotide sequence (or insert) with 5’ and 3’ adaptor sequences.
Figure 3 shows a typical polynucleotide with 5’ and 3’ adaptor sequences.
Figure 4 shows an example of PCR stitching process. Here, two sequences - a strand of a human library and a strand of a phiX library are joined together to create a single polynucleotide strand comprising both a first insert (comprising the strand of the human sequence) and a second insert (comprising the strand of the phiX sequence), as well as terminal and internal adaptor sequences.
Figure 5 shows the preparation of a polynucleotide sequence using a loop fork method.
Figure 6 shows a typical solid support with bound oligonucleotides.
Figure 7 shows the stages of bridge amplification and the generation of an amplified cluster comprising (Panel A) a library strand hybridising to an immobilised primer; (Panel B) generation of a template strand from the library strand; (Panel C) dehybridisation and washing away the library strand; (Panel D) hybridisation of the template strand to another immobilised primer; (Panel E) generation of a template complement strand from the template strand via bridge amplification; (Panel F) dehybridisation of the sequence bridge; (Panel G) hybridisation of the
template strand and template complement strand to immobilised primers; and (Panel H) subsequent bridge amplification to provide a plurality of template and template complement strands.
Figure 8 is a set of charts showing the detection of nucleobases using 4-channel, 2-channel and 1 -channel chemistry.
Figure 9 is a plot showing two-dimensional graphical representations of sixteen distributions of signals generated by polynucleotide sequences according to one embodiment.
Figure 10 is a flow diagram showing a method for base calling according to one embodiment.
Figure 11 is a plot showing graphical representations of nine distributions of signals generated by polynucleotide sequences according to one embodiment.
Figure 12 is a plot showing graphical representations of nine distributions of signals generated by polynucleotide sequences according to one embodiment, highlighting distributions that may be associated with library preparation errors.
Figure 13 is a flow diagram showing a method for determining sequence information according to one embodiment.
Figure 14 shows a process for preparing solid supports having central first regions and respective surrounding second regions. The first step involves grafting a flow cell with linearisable P5 lawn primers (i.e. a type of cleavable and unblocked first immobilised primer of the first proportion as described herein), non-linearisable P7 lawn primers (i.e. a type of non-cleavable and unblocked second immobilised primer of the fourth proportion as described herein), blocked non-linearisable P5 lawn primers (i.e. a type of non-cleavable and blocked first immobilised primer of the second proportion as described herein), and blocked linearisable P7 lawn primers (i.e. a type of cleavable and blocked second immobilised primer of the third proportion as described herein). The second step involves annealing a library strand (not shown) to a particular site, and generating template and template complement strands on the flow cell using amplification. As amplification cycles are conducted, template and template complement strands are generated in a radial direction (with the initial annealing point of the library strand as its centre), thus forming a “central” region. The third step involves removal of unused linearisable P5 lawn primers and unused non-linearisable P7 lawn primers using Exol. The fourth step involves unblocking of non-linearisable P5 lawn primers and linearisable P7 lawn primers. The fifth step involves generating further template and
template complement strands on the flow cell using amplification. As amplification cycles are conducted, template and template complement strands are again generated in a radial direction (i.e. towards the right on the figure for step 5), thus forming a surrounding ringed region around the “central” region. Since the “central” region is already saturated with template and template complement strands, further strands are generally formed in the surrounding region rather than within the “central” region. Thus, the central region is predominantly formed of template and template complement strands extending from linearisable P5 lawn primers and non-linearisable P7 lawn primers, whereas the surrounding ringed region is predominantly formed of template and template complement strands extending from non-linearisable P5 lawn primers and linearisable P7 lawn primers. Whilst the method is exemplified in Figure 14 with hairpin loops to act as blocking groups for the purpose of illustration, the hairpin loops can be substituted with other blocking groups such as phosphate groups (see Examples).
Figure 15 shows images from solid supports having central first regions and respective surrounding second regions. Different ratios of unblocked to blocked lawn primers were used (e.g. 2: 1 unblocked vs. blocked lawn primers, or 1: 1 unblocked vs. blocked lawn primers). The number of amplification cycles is indicated by the notation [m+n], where “m” indicates the number of amplification cycles used for generating the central first region, whereas “n” indicates the number of amplification cycles used for generating the surrounding second regions. 30 amplification cycles for the central first region gave good intensity centres, whilst the brightness of the ringed regions could be tuned (e.g. using 10 cycles for less intense rings, to 20 cycles for more intense rings). Whilst a blocking mix (BMX) is used in most cases to block any free 3 ’-OH ends on the flow cells before introduction of the sequencing primers, the similarity between the result of the “2: 1, 30+20” case (where BMX was used) compared to the result of the “2: 1, 30+20, no BMX” case (where no BMX was used) seems to show that the blocking is not strictly necessary.
Detailed Description
All patents, patent applications, and other publications referred to herein, including all sequences disclosed within these references, are expressly incorporated herein by reference, to the same extent as if each individual publication, patent or patent application was specifically and individually indicated to be incorporated by reference. All documents cited are, in relevant part, incorporated herein by reference in their entireties for the purposes indicated by the context of their citation herein. However, the citation of any document is not to be construed as an admission that it is prior art with respect to the present disclosure.
Embodiments can be used in sequencing, in particular concurrent sequencing reactions. Methodologies applicable to the present invention have been described in WO 08/041002, WO 07/052006, WO 98/44151, WO 00/18957, WO 02/06456, WO 07/107710, WO 05/068656, US 13/661,524 and US 2012/0316086, the contents of which are herein incorporated by reference. Further information can be found in US 20060024681, US 20060292611, WO 06/110855, WO 06/135342, WO 03/074734, WO 07/010252, WO 07/091077, WO 00/179553, WO 98/44152 and WO 2022/087150, the contents of which are herein incorporated by reference.
As used herein, the term “variant” refers to a variant polypeptide sequence or part of the polypeptide sequence that retains desired function of the full non-variant sequence. For example, a desired function of the immobilised primer retains the ability to bind (i.e. hybridise) to a target sequence.
As used in any aspect described herein, a “variant” has at least 25%, 26%, 27%, 28%, 29%, 30%,
31%, 32%, 33%, 34%, 35%, 36%, 37%, 38%, 39%, 40%, 41%, 42%, 43%, 44%, 45%, 46%, 47%.
48%, 49%, 50%, 51%, 52%, 53%, 54%, 55%, 56%, 57%, 58%, 59%, 60%, 61%, 62%, 63%, 64%.
65%, 66%, 67%, 68%, 69%, 70%, 71%, 72%, 73%, 74%, 75%, 76%, 77%, 78%, 79%, 80%, 81 %.
82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or at least 99% overall sequence identity to the non- variant nucleic acid sequence. The sequence identity of a variant can be determined using any number of sequence alignment programs known in the art. As an example, Emboss Stretcher from the EMBL-EBI may be used as found on the Internet at: ebi.ac.uk/Tools/psa/emboss_stretcher/ (using default parameters: pair output format, Matrix = BLOSUM62, Gap open = 1, Gap extend = 1 for proteins; pair output format, Matrix = DNAfull, Gap open = 16, Gap extend = 4 for nucleotides).
As used herein, the term “fragment” refers to a functionally active series of consecutive nucleic acids from a longer nucleic acid sequence. The fragment may be at least 99%, at least 95%, at least 90%, at least 80%, at least 70%, at least 60%, at least 50%, at least 40% or at least 30% the length of the longer nucleic acid sequence. In one embodiment, a fragment as used herein also retains the ability to bind (i.e. hybridise) to a target sequence.
Sequencing generally comprises four fundamental steps: 1) library preparation to form a plurality of target polynucleotides for identification; 2) cluster generation to form an array of amplified template polynucleotides; 3) sequencing the cluster array of amplified template polynucleotides; and 4) data analysis to identify characteristics of the target polynucleotides from the amplified template polynucleotide sequences. These steps are described in greater detail below.
Library strands and template terminology
As shown in Figure 1, for a given double-stranded polynucleotide sequence 100 to be identified, the polynucleotide sequence 100 comprises a forward strand of the sequence 101 and a reverse strand of the sequence 102.
When the polynucleotide sequence 100 is replicated (e.g. using a DNA/RNA polymerase), complementary versions of the forward strand 101 of the sequence 100 and the reverse strand 102 of the sequence 100 are generated. Thus, replication of the polynucleotide sequence 100 provides a double-stranded polynucleotide sequence 100a that comprises a forward strand of the sequence 101 and a forward complement strand of the sequence 101’, and a double-stranded polynucleotide sequence 100b that comprises a reverse strand of the sequence 102 and a reverse complement strand of the sequence 102’.
The term “template” may be used to describe a complementary version of the double-stranded polynucleotide sequence 100. As such, the “template” comprises a forward complement strand of the sequence 101’ and a reverse complement strand of the sequence 102’. Thus, by using the forward complement strand of the sequence 101’ as a template for complementary base pairing, a sequencing process (e.g. a sequencing-by-synthesis or a sequencing-by-ligation process) reproduces information that was present in the original forward strand of the sequence 101. Similarly, by using the reverse complement strand of the sequence 102’ as a template for complementary base pairing, a sequencing process (e.g. a sequencing-by-synthesis or a sequencing-by-ligation process) reproduces information that was present in the original reverse strand of the sequence 102.
The two strands in the template may also be referred to as a forward strand of the template 101 ’ and a reverse strand of the template 102’. The complement of the forward strand of the template 101 ’ is termed the forward complement strand of the template 101, whilst the complement of the reverse strand of the template 102’ is termed the reverse complement strand of the template 102.
Generally, where forward strand, reverse strand, forward complement strand, and reverse complement strand are used herein without qualifying whether they are with respect to the original polynucleotide sequence 100 or with respect to the “template”, these terms may be interpreted as referring to the “template”.
Library preparation
Library preparation is the first step in any high-throughput sequencing platform. These libraries allow templates to be generated via complementary base pairing that can subsequently be clustered and amplified. During library preparation, nucleic acid sequences, for example genomic DNA sample, or cDNA or RNA sample, is converted into a sequencing library, which can then be sequenced. By way of example with a DNA sample, the first step in library preparation is random fragmentation of the DNA sample. Sample DNA is first fragmented and the fragments of a specific size (typically 200-500 bp, but can be larger) are ligated, sub-cloned or “inserted” inbetween two oligo adaptors (adaptor sequences). The original sample DNA fragments are referred to as “inserts”. The target polynucleotides may advantageously also be size-fractionated prior to modification with the adaptor sequences.
As described herein, the templates to be generated typically include separate polynucleotide sequences, in particular a first polynucleotide sequence comprising a first portion and a second polynucleotide sequence comprising a second portion. Generating these templates from particular libraries may be performed according to methods known to persons of skill in the art. However, some example approaches of preparing libraries suitable for generation of such templates are described below.
In some embodiments, the library may be prepared by ligating adaptor sequences to doublestranded polynucleotide sequences, each comprising a forward strand of the sequence and a
reverse strand of the sequence, as described in more detail in e.g. WO 07/052006, which is incorporated herein by reference. In some cases, “tagmentation” can be used to attach the sample DNA to the adaptors, as described in more detail in e.g. WO 10/048605, US 2012/0301925, US 2013/0143774 and WO 2016/189331, each of which are incorporated herein by reference. In tagmentation, double-stranded DNA is simultaneously fragmented and tagged with adaptor sequences and PCR primer binding sites. The combined reaction eliminates the need for a separate mechanical shearing step during library preparation. These procedures may be used, for example, for preparing templates including a first polynucleotide sequence comprising a first portion and a second polynucleotide sequence comprising a second portion, wherein the first portion is a forward strand of the template, and the second portion is a forward complement strand of the template - i.e. a copy of the forward strand (or alternatively, wherein the first portion is a reverse strand of the template, and the second portion is a reverse complement strand of the template).
Where features herein are described in relation to the “forward” strand, it should be considered that these features could equally be applied to the “reverse strand”.
Where libraries are prepared by ligating adaptor sequences to double-stranded polynucleotide sequences as described above, library preparation may comprise ligating a first primer-binding sequence 301’ (e.g. P5’, such as SEQ ID NO. 3) and a second terminal sequencing primer binding site 304 (e.g. SBS3’, for example, SEQ ID NO. 8) to a 3’-end of a forward strand of a sequence 101. See Figure 2. The library preparation may be arranged such that the second terminal sequencing primer binding site 304 is attached (e.g. directly attached) to the 3 ’-end of the forward strand of the sequence 101, and such that the first primer-binding sequence 301’ is attached (e.g. directly attached) to the 3 ’-end of the second terminal sequencing primer binding site 304.
The library preparation may further comprise ligating a complement of first terminal sequencing primer binding site 303’ (e.g. SBS12, such as SEQ ID NO. 9) (also referred to herein as a first terminal sequencing primer binding site complement 303’) and a complement of a second primerbinding sequence 302 (also referred to herein as a second primer-binding complement sequence 302) (e.g. P7, such as SEQ ID NO. 2) to a 5 ’-end of the forward strand of the sequence 101. The library preparation may be arranged such that first terminal sequencing primer binding site complement 303’ is attached (e.g. directly attached) to the 5 ’-end of the forward strand of the sequence 101, and such that second primer-binding complement sequence 302 is attached (e.g. directly attached) to the 5 ’-end of first terminal sequencing primer binding site complement 303’.
Thus, one strand of a polynucleotide within a polynucleotide library may comprise, in a 5 ’ to 3 ’ direction, a second primer-binding complement sequence 302 (e.g. P7), a first terminal
sequencing primer binding site complement 303’ (e.g. SBS12), a forward strand of the sequence 101, a second terminal sequencing primer binding site 304 (e.g. SBS3’), and a first primer-binding sequence 301’ (e.g. P5’) (Figure 2 - bottom strand).
Although not shown in Figure 2, the strand may further comprise one or more index sequences. As such, a first index sequence (e.g. i7) may be provided between the second primer-binding complement sequence 302 (e.g. P7) and the first terminal sequencing primer binding site complement 303’ (e.g. SB SI 2). Separately, or in addition, a second index complement sequence (e.g. i5 ’) may be provided between the second terminal sequencing primer binding site 304 (e.g. SBS3’) and the first primer-binding sequence 301’ (e.g. P5’). Thus, in some embodiments, one strand of a polynucleotide within a polynucleotide library may comprise, in a 5’ to 3’ direction, a second primer-binding complement sequence 302 (e.g. P7), a first index sequence (e.g. i7), a first terminal sequencing primer binding site complement 303’ (e.g. SBS12), a forward strand of the sequence 101, a second terminal sequencing primer binding site 304 (e.g. SBS3’), a second index complement sequence (e.g. i5’), and a first primer-binding sequence 301’ (e.g. P5’). A typical polynucleotide is shown in Figure 3 (bottom strand).
When a double-stranded sequence 100 is used, the library preparation may also comprise ligating a second primer-binding sequence 302’ (e.g. P7’) and a first terminal sequencing primer binding site 303 (e.g. SBS12’) to a 3’-end of a reverse strand of a sequence 102. The library preparation may be arranged such that first terminal sequencing primer binding site 303 is attached (e.g. directly attached) to the 3 ’-end of the reverse strand of the sequence 102, and such that the second primer-binding sequence 302’ is attached (e.g. directly attached) to the 3 ’-end of first terminal sequencing primer binding site 303.
The library preparation may further comprise ligating a complement of a second terminal sequencing primer binding site 304’ (e.g. SBS3) (also referred to herein as a second terminal sequencing primer binding site complement 304’) and a complement of a first primer-binding sequence 301 (also referred to herein as a first primer-binding complement sequence 301) (e.g. P5) to a 5 ’-end of the reverse strand of the sequence 102. The library preparation may be arranged such that the second terminal sequencing primer binding site complement 304’ is attached (e.g. directly attached) to the 5 ’-end of the reverse strand of the sequence 102, and such that the first primer-binding complement sequence 301 is attached (e.g. directly attached) to the 5 ’-end of the second terminal sequencing primer binding site complement 304’.
Thus, another strand of a polynucleotide within a polynucleotide library may comprise, in a 5 ’ to 3’ direction, a first primer-binding complement sequence 301 (e.g. P5), a second terminal
sequencing primer binding site complement 304’ (e.g. SBS3), a reverse strand of the sequence 102, a first terminal sequencing primer binding site 303 (e.g. SBS12’), and a second primerbinding sequence 302’ (e.g. P7’) (Figure 2 - top strand).
Although not shown in Figure 2, the another strand may further comprise one or more index sequences. As such, a second index sequence (e.g. i5) may be provided between the first primerbinding complement sequence 301 (e.g. P5) and the second terminal sequencing primer binding site complement 304’ (e.g. SBS3). Separately, or in addition, a first index complement sequence (e.g. i7’) may be provided between the first terminal sequencing primer binding site 303 (e.g. SBS12’) and the second primer-binding sequence 302’ (e.g. P7’). Thus, in some embodiments, another strand of a polynucleotide within a polynucleotide library may comprise, in a 5’ to 3’ direction, a first primer-binding complement sequence 301 (e.g. P5), a second index sequence (e.g. i5), a second terminal sequencing primer binding site complement 304’ (e.g. SBS3), a reverse strand of the sequence 102, a first terminal sequencing primer binding site 303 (e.g. SBS12’), a first index complement sequence (e.g. i7’), and a second primer-binding sequence 302’ (e.g. P7’). A typical polynucleotide is shown in Figure 3 (top strand).
In some embodiments, the library may be prepared using PCR stitching methods, such as (splicing by) overlap extension PCR (also known as OE-PCR or SOE-PCR), as described in more detail in e.g. Higuchi et al. (Nucleic Acids Res., 1988, vol. 16, pp. 7351-7367), which is incorporated herein by reference. A representative process for conducting PCR stitching for a human and PhiX library is shown in Figure 4.
As used herein, the term “genetically unrelated” refers to portions which are not related in the sense of being any two of the group consisting of: forward strands, reverse strands, forward complement strands, and reverse complement strands. However, the “genetically unrelated” sequences could be different fragment sequences which are derived from the same source, but are different fragments from that source (e.g. from the same fragmented library preparation process). This includes sequences that can be overlapping in sequence (but not identical in sequence).
As will be described later, during clustering and amplification, further processes may be used to generate templates including a first polynucleotide sequence comprising a first portion and a second polynucleotide sequence comprising a second portion, wherein the first portion and the second portion are genetically unrelated.
In some embodiments, the library may be prepared using a loop fork method, which is described below. This procedure may be used, for example, for preparing templates including a first
polynucleotide sequence comprising a first portion and a second polynucleotide sequence comprising a second portion, wherein the first portion is a forward strand of the template, and the second portion is a reverse complement strand of the template (or alternatively, wherein the first portion is a reverse strand of the template, and the second portion is a forward complement strand of the template). Such libraries may also be referred to as self-tandem inserts. A representative process for conducting a loop fork method is shown in Figure 5.
Starting from a double-stranded polynucleotide sequence comprising a forward strand of the sequence and a reverse strand of the sequence, adaptors may be ligated to a first end of the sequence (e.g. using processes as described in more detail in e.g. WO 07/052006, or “tagmentation” methods as described above). A second end of the sequence (different from the first end) may be ligated to a loop, which connects the forward strand of the sequence and the reverse strand of the sequence, thus generating a loop fork ligated polynucleotide sequence. Conducting PCR on the loop fork ligated polynucleotide sequence produces a new doublestranded polynucleotide sequence, one strand comprising the forward strand of the sequence and the reverse strand of the sequence, and the other strand comprising a forward complement strand of the sequence and a reverse complement strand of the sequence. The library is now ready for seeding, clustering and amplification.
As will be described later, during clustering and amplification, further processes may be used to generate templates including a first polynucleotide sequence comprising a first portion and a second polynucleotide sequence comprising a second portion, wherein the first portion is a forward strand of the template, and the second portion is a reverse complement strand of the template (or alternatively, wherein the first portion is a reverse strand of the template, and the second portion is a forward complement strand of the template).
As will be understood by the skilled person, a double-stranded nucleic acid will typically be formed from two complementary polynucleotide strands comprised of deoxyribonucleotides or ribonucleotides joined by phosphodiester bonds, but may additionally include one or more ribonucleotides and/or non-nucleotide chemical moieties and/or non-naturally occurring nucleotides and/or non-naturally occurring backbone linkages. In particular, the double-stranded nucleic acid may include non-nucleotide chemical moieties, e.g. linkers or spacers, at the 5' end of one or both strands. By way of non-limiting example, the double-stranded nucleic acid may include methylated nucleotides, uracil bases, phosphorothioate groups, peptide conjugates etc. Such non-DNA or non-natural modifications may be included in order to confer some desirable property to the nucleic acid, for example to enable covalent, non-covalent or metal-coordination attachment to a solid support, or to act as spacers to position the site of cleavage an optimal
distance from the solid support. A single stranded nucleic acid consists of one such polynucleotide strand. Where a polynucleotide strand is only partially hybridised to a complementary strand - for example, a long polynucleotide strand hybridised to a short nucleotide primer - it may still be referred to herein as a single stranded nucleic acid.
A sequence comprising at least a primer-binding sequence (a primer-binding sequence and a sequencing primer binding site, in another aspect, a combination of a primer-binding sequence, an index sequence and a sequencing primer binding site) may be referred to herein as an adaptor sequence, and an insert is flanked by a 5’ adaptor sequence and a 3’ adaptor sequence. The primerbinding sequence may also comprise a sequencing primer for the index read.
As used herein, an “adaptor” refers to a sequence that comprises a short sequence-specific oligonucleotide that is ligated to the 5' and 3' ends of each DNA (or RNA) fragment in a sequencing library as part of library preparation. The adaptor sequence may further comprise nonpeptide linkers.
In a further embodiment, the P5’ and P7’ primer-binding sequences are complementary to short primer sequences (or lawn primers) present on the surface of a flow cell. Binding of P5’ and P7’ to their complements (P5 and P7) on - for example - the surface of the flow cell, permits nucleic acid amplification. As used herein denotes the complementary strand.
The primer-binding sequences in the adaptor which permit hybridisation to amplification primers (e.g. lawn primers) will typically be around 20-40 nucleotides in length, although the invention is not limited to sequences of this length. The precise identity of the amplification primers (e.g. lawn primers), and hence the cognate sequences in the adaptors, are generally not material to the invention, as long as the primer-binding sequences are able to interact with the amplification primers in order to direct PCR amplification. The sequence of the amplification primers may be specific for a particular target nucleic acid that it is desired to amplify, but in other embodiments these sequences may be "universal" primer sequences which enable amplification of any target nucleic acid of known or unknown sequence which has been modified to enable amplification with the universal primers. The criteria for design of PCR primers are generally well known to those of ordinary skill in the art.
The index sequences (also known as a barcode or tag sequence) are unique short DNA (or RNA) sequences that are added to each DNA (or RNA) fragment during library preparation. The unique sequences allow many libraries to be pooled together and sequenced simultaneously. Sequencing reads from pooled libraries are identified and sorted computationally, based on their barcodes,
before final data analysis. Library multiplexing is also a useful technique when working with small genomes or targeting genomic regions of interest. Multiplexing with barcodes can exponentially increase the number of samples analysed in a single run, without drastically increasing run cost or run time. Examples of tag sequences are found in WO05/068656, whose contents are incorporated herein by reference in their entirety. The tag can be read at the end of the first read, or equally at the end of the second read, for example using a sequencing primer complementary to the strand marked P7. The invention is not limited by the number of reads per cluster, for example two reads per cluster: three or more reads per cluster are obtainable simply by dehybridising a first extended sequencing primer, and rehybridising a second primer before or after a cluster repopulation/strand resynthesis step. Methods of preparing suitable samples for indexing are described in, for example WO 2008/093098, which is incorporated herein by reference. Single or dual indexing may also be used. With single indexing, up to 48 unique 6-base indexes can be used to generate up to 48 uniquely tagged libraries. With dual indexing, up to 24 unique 8-base Index 1 sequences and up to 16 unique 8-base Index 2 sequences can be used in combination to generate up to 384 uniquely tagged libraries. Pairs of indexes can also be used such that every i5 index and every i7 index are used only one time. With these unique dual indexes, it is possible to identify and filter indexed hopped reads, providing even higher confidence in multiplexed samples.
The sequencing primer binding sites are sequencing and/or index primer binding sites and indicate the starting point of the sequencing read. During the sequencing process, a sequencing primer anneals (i.e. hybridises) to at least a portion of the sequencing primer binding site on the template strand. The polymerase enzyme binds to this site and incorporates complementary nucleotides base by base into the growing opposite strand.
Cluster generation and amplification
Once a double stranded nucleic acid library is formed, typically, the library has previously been subjected to denaturing conditions to provide single stranded nucleic acids. Suitable denaturing conditions will be apparent to the skilled reader with reference to standard molecular biology protocols (Sambrook et al., 2001, Molecular Cloning, A Laboratory Manual, 4th Ed, Cold Spring Harbor Laboratory Press, Cold Spring Harbor Laboratory Press, NY; Current Protocols, eds Ausubel et al). In one embodiment, chemical denaturation may be used.
Pollowing denaturation, a single-stranded library may be contacted in free solution onto a solid support comprising surface capture moieties (for example P5 and P7 lawn primers).
Thus, embodiments of the present invention may be performed on a solid support 200, such as a flowcell. However, in alternative embodiments, seeding and clustering can be conducted off- flowcell using other types of solid support.
The solid support 200 may comprise a substrate 204. See Figure 6. The substrate 204 comprises at least one well 203 (e.g. a nanowell), and typically comprises a plurality of wells 203 (e.g. a plurality of nanowells).
In one embodiment, the solid support comprises a plurality of first immobilised primers and a plurality of second immobilised primers.
Thus, each well 203 may comprise a plurality of first immobilised primers 201. In addition, each well 203 may comprise a plurality of second immobilised primers 202. Thus, each well 203 may comprise a plurality of first immobilised primers 201 and a plurality of second immobilised primers 202.
The first immobilised primer 201 may be attached via a 5 ’-end of its polynucleotide chain to the solid support 200. When extension occurs from first immobilised primer 201, the extension may be in a direction away from the solid support 200.
The second immobilised primer 202 may be attached via a 5 ’-end of its polynucleotide chain to the solid support 200. When extension occurs from second immobilised primer 202, the extension may be in a direction away from the solid support 200.
The first immobilised primer 201 may be different to the second immobilised primer 202 and/or a complement of the second immobilised primer 202. The second immobilised primer 202 may be different to the first immobilised primer 201 and/or a complement of the first immobilised primer 201.
The (or each of the) first immobilised primer(s) 201 may comprise a sequence as defined in SEQ ID NO. 1 or 5, or a variant or fragment thereof. The (or each of the) second immobilised primer(s) 202 may comprise a sequence as defined in SEQ ID NO. 2, or a variant or fragment thereof. Whilst first immobilised primer(s) 201 are shown here to correspond to P5 and second immobilised primer(s) 202 are shown here to correspond to P7, the definitions of these may be swapped - in other words, first immobilised primer(s) 201 may correspond instead to P7, and second immobilised primer(s) 202 may correspond to P5.
By way of brief example, following attachment of the P5 and P7 primers to the solid support, the solid support may be contacted with the template to be amplified under conditions which permit hybridisation (or annealing - such terms may be used interchangeably) between the template and the immobilised primers. The template is usually added in free solution under suitable hybridisation conditions, which will be apparent to the skilled reader. Typically, hybridisation conditions are, for example, 5xSSC at 40°C. However, other temperatures may be used during hybridisation, for example about 50°C to about 75°C, about 55°C to about 70°C, or about 60°C to about 65°C. Solid-phase amplification can then proceed. The first step of the amplification is a primer extension step in which nucleotides are added to the 3' end of the immobilised primer using the template to produce a fully extended complementary strand. The template is then typically washed off the solid support. The complementary strand will include at its 3' end a primer-binding sequence (i.e. either P5’ or P7’) which is capable of bridging to the second primer molecule immobilised on the solid support and binding. Further rounds of amplification (analogous to a standard PCR reaction) leads to the formation of clusters or colonies of template molecules bound to the solid support. This is called clustering.
Thus, solid-phase amplification by either a method analogous to that of WO 98/44151 or that of WO 00/18957 (the contents of which are incorporated herein in their entirety by reference) will result in production of a clustered array comprised of colonies of "bridged" amplification products. This process is known as bridge amplification. Both strands of the amplification products will be immobilised on the solid support at or near the 5' end, this attachment being derived from the original attachment of the amplification primers. Typically, the amplification products within each colony will be derived from amplification of a single template molecule. Other amplification procedures may be used, and will be known to the skilled person. For example, amplification may be isothermal amplification using a strand displacement polymerase; or may be exclusion amplification as described in WO 2013/188582. Further information on amplification can be found in WO 02/06456 and WO 07/107710, the contents of which are incorporated herein in their entirety by reference.
Through such approaches, a cluster of template molecules is formed, comprising copies of a template strand and copies of the complement of the template strand.
The steps of cluster generation and amplification for templates including a first polynucleotide sequence comprising a first portion and a second polynucleotide sequence comprising a second portion, are illustrated below and in Figure 7.
In cases where (separate) polynucleotide strands are used, each first polynucleotide sequence may be attached (via the 5 ’-end of the first polynucleotide sequence) to a first immobilised primer, and wherein each second polynucleotide sequence is attached (via the 5 ’-end of the second polynucleotide sequence) to a second immobilised primer. Each first polynucleotide sequence may comprise a second adaptor sequence, wherein the second adaptor sequence comprises a portion, which is substantially complementary to the second immobilised primer (or is substantially complementary to the second immobilised primer). The second adaptor sequence may be at a 3 ’-end of the first polynucleotide sequence. Each second polynucleotide sequence may comprise a first adaptor sequence, wherein the first adaptor sequence comprises a portion, which is substantially complementary to the first immobilised primer (or is substantially complementary to the first immobilised primer). The first adaptor sequence may be at a 3 ’-end of the second polynucleotide sequence.
In an embodiment, a solution comprising a polynucleotide library prepared by ligating adaptor sequences to double-stranded polynucleotide sequences as described above may be flown across a flowcell.
A particular polynucleotide strand from the polynucleotide library to be sequenced comprising, in a 5’ to 3’ direction, a second primer-binding complement sequence 302 (e.g. P7), a first terminal binding site complement 303’ (e.g. SBS12), a forward strand of the sequence 101, a second terminal sequencing primer binding site 304 (e.g. SBS3’) and a first primer-binding sequence 301’ (e.g. P5’), may anneal (via the first primer-binding sequence 301’) to the first immobilised primer 201 (e.g. P5 lawn primer) located within a particular well 203 (Figure 7A).
The polynucleotide library may comprise other polynucleotide strands with different forward strands of the sequence 101. Such other polynucleotide strands may anneal to corresponding first immobilised primers 201 (e.g. P5 lawn primers) in different wells 203, thus enabling parallel processing of the various different strands within the polynucleotide library.
A new polynucleotide strand may then be synthesised, extending from the first immobilised primer 201 (e.g. P5 lawn primer) in a direction away from the substrate 204. By using complementary base-pairing, this generates a template strand comprising, in a 5 ’ to 3 ’ direction, the first immobilised primer 201 (e.g. P5 lawn primer) which is attached to the solid support 200, a second terminal sequencing primer binding site complement 304’ (e.g. SBS3), a forward strand of the template 101’ (which represents a type of “first portion”), a first terminal sequencing primer binding site 303 (which represents atype of “first sequencing primer binding site”) (e.g. SBS12’),
and a second primer-binding sequence 302’ (e.g. P7’) (Figure 7B). Such a process may utilise an appropriate polymerase, such as a DNA or RNA polymerase.
If the polynucleotides in the library comprise index sequences, then corresponding index sequences are also produced in the template.
The polynucleotide strand from the polynucleotide library may then be dehybridised and washed away, leaving a template strand attached to the first immobilised primer 201 (e.g. P5 lawn primer) (Figure 7C).
The second primer-binding sequence 302’ (e.g. P7’) on the template strand may then anneal to a second immobilised primer 202 (e.g. P7 lawn primer) located within the well 203. This forms a “bridge” (Figure 7D).
A new polynucleotide strand may then be synthesised by bridge amplification, extending from the second immobilised primer 202 (e.g. P7 lawn primer) (initially) in a direction away from the substrate 204. By using complementary base-pairing, this generates a template strand comprising, in a 5’ to 3’ direction, the second immobilised primer 202 (e.g. P7 lawn primer) which is attached to the solid support 200, a first terminal sequencing primer binding site complement 303’ (e.g. SBS12), a forward complement strand of the template 101 (which represents a type of “second portion”), a second terminal sequencing primer binding site 304 (which represents a type of “second sequencing primer binding site”) (e.g. SBS3’), and a first primer-binding sequence 301’ (e.g. P5’) (Figure 7E). Again, such a process may utilise a suitable polymerase, such as a DNA or RNA polymerase.
The strand attached to the second immobilised primer 202 (e.g. P7 lawn primer) may then be dehybridised from the strand attached to the first immobilised primer 201 (e.g. P5 lawn primer) (Figure 7F).
A subsequent bridge amplification cycle can then lead to amplification of the strand attached to the first immobilised primer 201 (e.g. P5 lawn primer) and the strand attached to the second immobilised primer 202 (e.g. P7 lawn primer). Similar to Figure 7D, the second primer-binding sequence 302’ (e.g. P7’) on the template strand attached to the first immobilised primer 201 (e.g. P5 lawn primer) may then anneal to another second immobilised primer 202 (e.g. P7 lawn primer) located within the well 203. In a similar fashion, the first primer-binding sequence 301’ (e.g. P5’) on the template strand attached to the second immobilised primer 202 (e.g. P7 lawn primer) may
then anneal to another first immobilised primer 201 (e.g. P5 lawn primer) located within the well 203 (Figure 7G).
Completion of bridge amplification and dehybridisation may then provide an amplified (duoclonal) cluster, thus providing a plurality of first polynucleotide sequences comprising the forward strand of the template 101’ (i.e. “first portions”), and a plurality of second polynucleotide sequences comprising the forward complement strand of the template 101 (i.e. “second portions”) (Figure 7H).
If desired, further bridge amplification cycles may be conducted to increase the number of first polynucleotide sequences and second polynucleotide sequences within the well 203.
In this particular example, the “first portion” corresponds with the forward strand of the template 101’, and the “second portion” corresponds with the forward complement strand of the template 101.
However, other set-ups may be obtained by changing the library used. For example, by using a loop fork method to prepare a library, a portion at or close to the loop (or the loop complement) may be cleaved (e.g. by nicking). In these cases, the loop may comprise a cleavage site (e.g. a restriction recognition site, a cleavable linker, a modified nucleotide, or the like). By conducting cleavage at the loop, it is possible to produce a well 203, where the “first portion” corresponds with a forward strand of the template, and the “second portion” corresponds with a reverse complement strand of the template. In addition, by using a PCR stitching method to prepare a library, a portion at or close to the overlap region may comprise a cleavage site (e.g. a restriction recognition site, a cleavable linker, a modified nucleotide, or the like). By conducting cleavage at the overlap region, it is possible to produce a well 203, where the “first portion” corresponds with a first insert sequence, and the “second portion” corresponds with a second insert sequence that is genetically unrelated to the first insert sequence. As such, different types of strands for the “first portions” and “second portions” may be prepared for templates including a first polynucleotide sequence comprising a first portion and a second polynucleotide sequence comprising a second portion, and as such the forward strand of the template 101’ and the forward complement strand of the template 101 may be substituted as appropriate.
Sequencing
As described herein, the template provides information (e.g. identification of the genetic sequence, identification of epigenetic modifications) on the original target polynucleotide
sequence. For example, a sequencing process (e.g. a sequencing-by-synthesis or sequencing-by- ligation process) may reproduce information that was present in the original target polynucleotide sequence, by using complementary base pairing.
In one embodiment, sequencing may be carried out using any suitable "sequencing-by-synthesis" technique, wherein nucleotides are added successively in cycles to the free 3' hydroxyl group, resulting in synthesis of a polynucleotide chain in the 5' to 3' direction. The nature of the nucleotide added may be determined after each addition. One particular sequencing method relies on the use of modified nucleotides that can act as reversible chain terminators. Such reversible chain terminators comprise removable 3' blocking groups. Once such a modified nucleotide has been incorporated into the growing polynucleotide chain complementary to the region of the template being sequenced there is no free 3 '-OH group available to direct further sequence extension and therefore the polymerase cannot add further nucleotides. Once the nature of the base incorporated into the growing chain has been determined, the 3' block may be removed to allow addition of the next successive nucleotide. By ordering the products derived using these modified nucleotides it is possible to deduce the DNA sequence of the DNA template. Such reactions can be done in a single experiment if each of the modified nucleotides has attached thereto a different label, known to correspond to the particular base, to facilitate discrimination between the bases added at each incorporation step. Suitable labels are described in PCT application PCT/GB2007/001770, the contents of which are incorporated herein by reference in their entirety. Alternatively, a separate reaction may be carried out containing each of the modified nucleotides added individually.
The modified nucleotides may carry a label to facilitate their detection. Such a label may be configured to emit a signal, such as an electromagnetic signal, or a (visible) light signal.
In a particular embodiment, the label is a fluorescent label (e.g. a dye). Thus, such a label may be configured to emit an electromagnetic signal, or a (visible) light signal. One method for detecting the fluorescently labelled nucleotides comprises using laser light of a wavelength specific for the labelled nucleotides, or the use of other suitable sources of illumination. The fluorescence from the label on an incorporated nucleotide may be detected by a CCD camera or other suitable detection means. Suitable detection means are described in PCT/US2007/007991, the contents of which are incorporated herein by reference in their entirety.
However, the detectable label need not be a fluorescent label. Any label can be used which allows the detection of the incorporation of the nucleotide into the DNA sequence.
Each cycle may involve simultaneous delivery of four different nucleotide types to the array of template molecules. Alternatively, different nucleotide types can be added sequentially and an image of the array of template molecules can be obtained between each addition step.
In some embodiments, each nucleotide type may have a (spectrally) distinct label. In other words, four channels may be used to detect four nucleobases (also known as 4-channel chemistry) (Figure 8 - left). For example, a first nucleotide type (e.g. A) may include a first label (e.g. configured to emit a first wavelength, such as red light), a second nucleotide type (e.g. G) may include a second label (e.g. configured to emit a second wavelength, such as blue light), a third nucleotide type (e.g. T) may include a third label (e.g. configured to emit a third wavelength, such as green light), and a fourth nucleotide type (e.g. C) may include a fourth label (e.g. configured to emit a fourth wavelength, such as yellow light). Four images can then be obtained, each using a detection channel that is selective for one of the four different labels. For example, the first nucleotide type (e.g. A) may be detected in a first channel (e.g. configured to detect the first wavelength, such as red light), the second nucleotide type (e.g. G) may be detected in a second channel (e.g. configured to detect the second wavelength, such as blue light), the third nucleotide type (e.g. T) may be detected in a third channel (e.g. configured to detect the third wavelength, such as green light), and the fourth nucleotide type (e.g. C) may be detected in a fourth channel (e.g. configured to detect the fourth wavelength, such as yellow light). Although specific pairings of bases to signal types (e.g. wavelengths) are described above, different signal types (e.g. wavelengths) and/or permutations may also be used.
In some embodiments, detection of each nucleotide type may be conducted using fewer than four different labels. For example, sequencing-by-synthesis may be performed using methods and systems described in US 2013/0079232, which is incorporated herein by reference.
Thus, in some embodiments, two channels may be used to detect four nucleobases (also known as 2-channel chemistry) (Figure 8 - middle). For example, a first nucleotide type (e.g. A) may include a first label (e.g. configured to emit a first wavelength, such as green light) and a second label (e.g. configured to emit a second wavelength, such as red light), a second nucleotide type (e.g. G) may not include the first label and may not include the second label, a third nucleotide type (e.g. T) may include the first label (e.g. configured to emit the first wavelength, such as green light) and may not include the second label, and a fourth nucleotide type (e.g. C) may not include the first label and may include the second label (e.g. configured to emit the second wavelength, such as red light). Two images can then be obtained, using detection channels for the first label and the second label. For example, the first nucleotide type (e.g. A) may be detected in both a first channel (e.g. configured to detect the first wavelength, such as red light) and a second channel
(e.g. configured to detect the second wavelength, such as green light), the second nucleotide type (e.g. G) may not be detected in the first channel and may not be detected in the second channel, the third nucleotide type (e.g. T) may be detected in the first channel (e.g. configured to detect the first wavelength, such as red light) and may not be detected in the second channel, and the fourth nucleotide type (e.g. C) may not be detected in the first channel and may be detected in the second channel (e.g. configured to detect the second wavelength, such as green light). Although specific pairings of bases to signal types (e.g. wavelengths) and/or combinations of channels are described above, different signal types (e.g. wavelengths) and/or permutations may also be used.
In some embodiments, one channel may be used to detect four nucleobases (also known as 1- channel chemistry) (Figure 8 - right). For example, a first nucleotide type (e.g. A) may include a cleavable label (e.g. configured to emit a wavelength, such as green light), a second nucleotide type (e.g. G) may not include a label, a third nucleotide type (e.g. T) may include a non-cleavable label (e.g. configured to emit the wavelength, such as green light), and a fourth nucleotide type (e.g. C) may include a label-accepting site which does not include the label. A first image can then be obtained, and a subsequent treatment carried out to cleave the label attached to the first nucleotide type, and to attach the label to the label-accepting site on the fourth nucleotide type. A second image may then be obtained. For example, the first nucleotide type (e.g. A) may be detected in a channel (e.g. configured to detect the wavelength, such as green light) in the first image and not detected in the channel in the second image, the second nucleotide type (e.g. G) may not be detected in the channel in the first image and may not be detected in the channel in the second image, the third nucleotide type (e.g. T) may be detected in the channel (e.g. configured to detect the wavelength, such as green light) in the first image and may be detected in the channel (e.g. configured to detect the wavelength, such as green light) in the second image, and the fourth nucleotide type (e.g. C) may not be detected in the channel in the first image and may be detected in the channel in the second image (e.g. configured to detect the wavelength, such as green light). Although specific pairings of bases to signal types (e.g. wavelengths) and/or combinations of images are described above, different signal types (e.g. wavelengths), images and/or permutations may also be used.
In one embodiment, the sequencing process comprises a first sequencing read and second sequencing read. The first sequencing read and the second sequencing read may be conducted concurrently. In other words, the first sequencing read and the second sequencing read may be conducted at the same time.
The first sequencing read may comprise the binding of a first sequencing primer (also known as a read 1 sequencing primer) to the first sequencing primer binding site (e.g. first terminal
sequencing primer binding site 303 in templates including a first polynucleotide sequence comprising a first portion and a second polynucleotide sequence comprising a second portion). The second sequencing read may comprise the binding of a second sequencing primer (also known as a read 2 sequencing primer) to the second sequencing primer binding site (e.g. second terminal sequencing primer binding site 304 in templates including a first polynucleotide sequence comprising a first portion and a second polynucleotide sequence comprising a second portion).
This leads to sequencing of the first portion (e.g. forward strand of the template 101’ in templates including a first polynucleotide sequence comprising a first portion and a second polynucleotide sequence comprising a second portion; or other types of first portion if different library preparations are used, such as by PCR stitching or loop fork methods) and the second portion (e.g. forward complement strand of the template 101 in templates including a first polynucleotide sequence comprising a first portion and a second polynucleotide sequence comprising a second portion; or other types of first portion if different library preparations are used, such as by PCR stitching or loop fork methods).
Alternative methods of sequencing may include sequencing by ligation, for example as described in US 6,306,597 or WO 06/084132, the contents of which are incorporated herein by reference.
Data analysis using 16 QaM
Figure 9 is a scatter plot showing an example of sixteen distributions of signals generated by polynucleotide sequences disclosed herein.
The two-dimensional scatter plot of Figure 9 shows sixteen distributions (or bins) of intensity values from the combination of a brighter signal (i.e. a first signal as described herein) and a dimmer signal (i.e. a second signal as described herein); the two signals may be co-localized and may not be optically resolved as described above. The intensity values shown in Figure 9 may be up to a scale or normalisation factor; the units of the intensity values may be arbitrary or relative (i.e., representing the ratio of the actual intensity to a reference intensity). The sum of the brighter signal generated by the first portions and the dimmer signal generated by the second portions results in a combined signal. The combined signal may be captured by a first optical channel and a second optical channel. Since the brighter signal may be A, T, C or G, and the dimmer signal may be A, T, C or G, there are sixteen possibilities for the combined signal, corresponding to sixteen distinguishable patterns when optically captured. That is, each of the sixteen possibilities corresponds to a bin shown in Figure 9. The computer system can map the combined signal
generated into one of the sixteen bins, and thus determine the added nucleobase at the first portion and the added nucleobase at the second portion, respectively.
For example, when the combined signal is mapped to bin 1612 for a base calling cycle, the computer processor base calls both the added nucleobase at the first portion and the added nucleobase at the second portion as C. When the combined signal is mapped to bin 1614 for the base calling cycle, the processor base calls the added nucleobase at the first portion as C and the added nucleobase at the second portion as T. When the combined signal is mapped to bin 1616 for the base calling cycle, the processor base calls the added nucleobase at the first portion as C and the added nucleobase at the second portion as G. When the combined signal is mapped to bin 1618 for the base calling cycle, the processor base calls the added nucleobase at the first portion as C and the added nucleobase at the second portion as A.
When the combined signal is mapped to bin 1622 for the base calling cycle, the processor base calls the added nucleobase at the first portion as T and the added nucleobase at the second portion as C. When the combined signal is mapped to bin 1624 for the base calling cycle, the processor base calls both the added nucleobase at the first portion and the added nucleobase at the second portion as T. When the combined signal is mapped to bin 1626 for the base calling cycle, the processor base calls the added nucleobase at the first portion as T and the added nucleobase at the second portion as G. When the combined signal is mapped to bin 1628 for the base calling cycle, the processor base calls the added nucleobase at the first portion as T and the added nucleobase at the second portion as A.
When the combined signal is mapped to bin 1632 for the base calling cycle, the processor base calls the added nucleobase at the first portion as G and the added nucleobase at the second portion as C. When the combined signal is mapped to bin 1634 for the base calling cycle, the processor base calls the added nucleobase at the first portion as G and the added nucleobase at the second portion as T. When the combined signal is mapped to bin 1636 for the base calling cycle, the processor base calls both the added nucleobase at the first portion and the added nucleobase at the second portion as G. When the combined signal is mapped to bin 1638 for the base calling cycle, the processor base calls the added nucleobase at the first portion as G and the added nucleobase at the second portion as A.
When the combined signal is mapped to bin 1642 for the base calling cycle, the processor base calls the added nucleobase at the first portion as A and the added nucleobase at the second portion as C. When the combined signal is mapped to bin 1644 for the base calling cycle, the processor base calls the added nucleobase at the first portion as A and the added nucleobase at the second
portion as T. When the combined signal is mapped to bin 1646 for the base calling cycle, the processor base calls the added nucleobase at the first portion as A and the added nucleobase at the second portion as G. When the combined signal is mapped to bin 1648 for the base calling cycle, the processor base calls both the added nucleobase at the first portion and the added nucleobase at the second portion as A.
In this particular example, T is configured to emit a signal in both the IMAGE 1 channel and the IMAGE 2 channel, A is configured to emit a signal in the IMAGE 1 channel only, C is configured to emit a signal in the IMAGE 2 channel only, and G does not emit a signal in either channel. However, different permutations of nucleobases can be used to achieve the same effect by performing dye swaps. For example, A may be configured to emit a signal in both the IMAGE 1 channel and the IMAGE 2 channel, T may be configured to emit a signal in the IMAGE 1 channel only, C may be configured to emit a signal in the IMAGE 2 channel only, and G may be configured to not emit a signal in either channel.
Further details regarding performing base-calling based on a scatter plot having sixteen bins may be found in U.S. Patent Application Publication No. 2019/0212294, the disclosure of which is incorporated herein by reference.
Figure 10 is a flow diagram showing a method 1700 of base calling according to the present disclosure. The described method allows for simultaneous sequencing of two (or more) portions (e.g. the first portion and the second portion) in a single sequencing run from a single combined signal obtained from the first portion and the second portion, thus requiring less sequencing reagent consumption and faster generation of data from both the first portion and the second portion. Further, the simplified method may reduce the number of workflow steps while producing the same yield as compared to existing next-generation sequencing methods. Thus, the simplified method may result in reduced sequencing runtime.
As shown in Figure 10, the disclosed method 1700 may start from block 1701. The method may then move to block 1710.
At block 1710, intensity data is obtained. The intensity data includes first intensity data and second intensity data. The first intensity data comprises a combined intensity of a first signal component obtained based upon a respective first nucleobase of the first portion and a second signal component obtained based upon a respective second nucleobase of the second portion. Similarly, the second intensity data comprises a combined intensity of a third signal component
obtained based upon the respective first nucleobase of the first portion and a fourth signal component obtained based upon the respective second nucleobase of the second portion.
As such, the first portion is capable of generating a first signal comprising a first signal component and a third signal component. The second portion is capable of generating a second signal comprising a second signal component and a fourth signal component.
As described above, the first portion and the second portion may be arranged on the solid support such that signals from the first portion and the second portion are detected by a single sensing portion and/or may comprise a single cluster such that first signals and second signals from each of the respective first portions and second portions cannot be spatially resolved.
In one example, obtaining the intensity data comprises selecting intensity data that corresponds to two (or more) different portions (e.g. the first portion and the second portion). In one example, intensity data is selected based upon a chastity score. A chastity score may be calculated as the ratio of the brightest base intensity divided by the sum of the brightest and second brightest base intensities. The desired chastity score may be different depending upon the expected intensity ratio of the light emissions associated with the different portions. As described above, it may be desired to produce clusters comprising the first portion and the second portion, which give rise to signals in a ratio of 2: 1. In one example, high-quality data corresponding to two portions with an intensity ratio of 2: 1 may have a chastity score of around 0.8 to 0.9.
After the intensity data has been obtained, the method may proceed to block 1720. In this step, one of a plurality of classifications is selected based on the intensity data. Each classification represents a possible combination of respective first and second nucleobases. In one example, the plurality of classifications comprises sixteen classifications as shown in Figure 9, each representing a unique combination of first and second nucleobases. Where there are two portions, there are sixteen possible combinations of first and second nucleobases. Selecting the classification based on the first and second intensity data comprises selecting the classification based on the combined intensity of the first and second signal components and the combined intensity of the third and fourth signal components.
The method may then proceed to block 1730, where the respective first and second nucleobases are base called based on the classification selected in block 1720. The signals generated during a cycle of a sequencing are indicative of the identity of the nucleobase(s) added during sequencing (e.g. using sequencing-by-synthesis). It will be appreciated that there is a direct correspondence between the identity of the nucleobases that are incorporated and the identity of the
complementary base at the corresponding position of the template sequence bound to the solid support. Therefore, any references herein to the base calling of respective nucleobases at the two portions encompasses the base calling of nucleobases hybridised to the template sequences and, alternatively or additionally, the identification of the corresponding nucleobases of the template sequences. The method may then end at block 1740.
Data analysis using 9 QaM
For two portions of polynucleotide sequences (e.g. a first portion and a second portion as described herein), there are sixteen possible combinations of nucleobases at any given position (i.e., an A in the first portion and an A in the second portion, an A in the first portion and a T in the second portion, and so on). When the same nucleobase is present at a given position in both portions, the light emissions associated with each target sequence during the relevant base calling cycle will be characteristic of the same nucleobase. In effect, the two portions behave as a single portion, and the identity of the bases at that position are uniquely callable.
However, when a nucleobase of the first portion is different from a nucleobase at a corresponding position of the second portion, the signals associated with each portion in the relevant base calling cycle will be characteristic of different nucleobases. In one embodiment, the first signal coming from the first portion have substantially the same intensity as the second signal coming from the second portion. The two signals may also be co-localised, and may not be spatially and/or optically resolved. Therefore, when different nucleobases are present at corresponding positions of the two portions, the identity of the nucleobases cannot be uniquely called from the combined signal alone. However, useful sequencing information can still be determined from these signals.
The scatter plot of Figure 11 shows nine distributions (or bins) of intensity values from the combination of two co-localised signals of substantially equal intensity.
The intensity values shown in Figure 11 may be up to a scale or normalisation factor; the units of the intensity values may be arbitrary or relative (i.e., representing the ratio of the actual intensity to a reference intensity). The sum of the first signal generated from the first portion and the second signal generated from the second portion results in a combined signal. The combined signal may be captured by a first optical channel and a second optical channel. The computer system can map the combined signal generated into one of the nine bins, and thus determine sequence information relating to the added nucleobase at the first portion and the added nucleobase at the second portion.
Bins are selected based upon the combined intensity of the signals originating from each target sequence during the base calling cycle. For example, bin 1803 may be selected following the detection of a high-intensity (or “on/on”) signal in the first channel and a high-intensity signal in the second channel. Bin 1806 may be selected following the detection of a high-intensity signal in the first channel and an intermediate-intensity (“on/off’ or “off/on”) signal in the second channel. Bin 1809 may be selected following the detection of a high-intensity signal in the first channel and a low-intensity or zero-intensity (“off/off’) signal in the second channel. Bin 1802 may be selected following the detection of an intermediate-intensity signal in the first channel and a high-intensity signal in the second channel. Bin 1805 may be selected following the detection of an intermediate-intensity signal in the first channel and an intermediate-intensity signal in the second channel. Bin 1808 may be selected following the detection of an intermediateintensity signal in the first channel and a low-intensity or zero-intensity signal in the second channel. Bin 1801 may be selected following the detection of a low-intensity signal in the first channel and a high-intensity signal in the second channel. Bin 1804 may be selected following the detection of a low-intensity or zero-intensity signal in the first channel and an intermediateintensity signal in the second channel. Bin 1807 may be selected following the detection of a low- intensity or zero-intensity signal in the first channel and a low-intensity signal in the second channel.
Four of the nine bins represent matches between respective nucleobases of the two portions sensed during the cycle (bins 1801, 1803, 1807, and 1809). In response to mapping the combined signal to a bin representing a match, the computer processor may detect a match between the first portion and the second portion at the sensed position. In response to mapping the combined signal to a bin representing a match, the computer processor may base call the respective nucleobases. For example, when the combined signal is mapped to bin 1801 for a base calling cycle, the computer processor base calls both the added nucleobase at the first portion and the added nucleobase at the second portion as T. When the combined signal is mapped to bin 1803 for the base calling cycle, the processor base calls both the added nucleobase at the first portion and the added nucleobase at the second portion as A. When the combined signal is mapped to bin 1807 for the base calling cycle, the processor base calls both the added nucleobase at the first portion and the added nucleobase at the second portion as G. When the combined signal is mapped to bin 1809 for the base calling cycle, the processor base calls both the added nucleobase at the first portion and the added nucleobase at the second portion as C.
The remaining five bins are “ambiguous”. That is to say that these bins each represent more than one possible combination of first and second nucleobases. Bins 1802, 1804, 1806, and 1808 each represent two possible combinations of first and second nucleobases. Bin 1805, meanwhile,
represents four possible combinations. Nevertheless, mapping the combined signal to an ambiguous bin may still allow for sequencing information to be determined. For example, bins 1802, 1804, 1805, 1806, and 1808 represent mismatches between respective nucleobases of the two portions sensed during the cycle. Therefore, in response to mapping the combined signal to a bin representing a mismatch, the computer processor may detect a mismatch between the first portion and the second portion at the sensed position.
In this particular example, A is configured to emit a signal in both the first channel and the second channel, C is configured to emit a signal in the first channel only, T is configured to emit a signal in the second channel only, and G does not emit a signal in either channel. However, different permutations of nucleobases can be used to achieve the same effect by performing dye swaps. For example, A may be configured to emit a signal in both the first channel and the second channel, T may be configured to emit a signal in the first channel only, C may be configured to emit a signal in the second channel only, and G may be configured to not emit a signal in either channel.
The number of classifications which may be selected based upon the combined signal intensities may be predetermined, for example based on the number of portions expected to be present in the nucleic acid cluster. Whilst Figure 11 shows a set of nine possible classifications, the number of classifications may be greater or smaller.
In addition to identifying matches and mismatches, the mapping of the combined signal to each of the different bins (e.g. in combination with additional knowledge, such as the library preparation methods used) can provide additional information about the first portion and the second portion, or about sequences from which the first portion and the second portion were derived. For example, given the nucleic acid material input and the processing methods used to generate the nucleic acid clusters, the first portion and the second portion may be expected to be identical at a given position. In this case, the mapping of the combined signal to a bin representing a mismatch may be indicative of an error introduced during library preparation.
Errors may arise during NGS library preparation, for example due to PCR artifacts or DNA damage. The error rate is determined by the library preparation method used, for example the number of cycles of PCR amplification carried out, and a typical error rate may be of the order of 0.1%. This can limit the sensitivity of diagnostic assays based on the sequencing method, and may obscure true variants. In some embodiments, the present methods allow for the identification of library preparation errors from fewer sequencing reads.
In the absence of any library preparation/sequencing errors, the signals produced by sequencing the two portions (e.g. using sequencing-by-synthesis) will match. The combined signal may therefore be mapped to one of the four “comer” clouds shown in Figure 11 and Figure 12, and the identity of the nucleobase at the corresponding position of the original library polynucleotide can be determined. Should the identity of the nucleobase at that position suggest a rare, or even unknown, variant, it can be determined with a high level of confidence that the base call represents a true variant, as opposed to a library preparation error. If, on the other hand, the combined signal is mapped to any of the other clouds, this indicates that the sequences of the first portion and the second portion do not match, and that an error has occurred in library preparation. Therefore, in response to mapping the combined signal to a classification representing a mismatch between the two nucleobases, a library preparation error may be identified.
Figure 13 is a flow diagram showing a method 1900 of determining sequence information according to the present disclosure. The described method allows for the determination of sequence information from two (or more) portions (e.g. the first portion and the second portion) in a single sequencing run from a single combined signal obtained from the first portion and the second portion.
As shown in Figure 13, the disclosed method 1900 may start from block 1901. The method may then move to block 1910.
At block 1910, intensity data is obtained. The intensity data includes first intensity data and second intensity data. The first intensity data comprises a combined intensity of a first signal component obtained based upon a respective first nucleobase of the first portion and a second signal component obtained based upon a respective second nucleobase of the second portion. Similarly, the second intensity data comprises a combined intensity of a third signal component obtained based upon the respective first nucleobase of the first portion and a fourth signal component obtained based upon the respective second nucleobase of the second portion.
As such, the first portion is capable of generating a first signal comprising a first signal component and a third signal component. The second portion is capable of generating a second signal comprising a second signal component and a fourth signal component.
As described above, the first portion and the second portion may be arranged on the solid support such that signals from the first portion and the second portion are detected by a single sensing portion and/or may comprise a single cluster such that first signals and second signals from each of the respective first portions and second portions cannot be spatially resolved.
In one example, obtaining the intensity data comprises selecting intensity data, for example based upon a chastity score. A chastity score may be calculated as the ratio of the brightest base intensity divided by the sum of the brightest and second brightest base intensities. In one example, high- quality data corresponding to two portions with a substantially equal intensity ratio may have a chastity score of around 0.8 to 0.9, for example 0.89-0.9.
After the intensity data has been obtained, the method may proceed to block 1920. In this step, one of a plurality of classifications is selected based on the intensity data. Each classification represents one or more possible combinations of respective first and second nucleobases, and at least one classification of the plurality of classifications represents more than one possible combination of respective first and second nucleobases. In one example, the plurality of classifications comprises nine classifications as shown in Figure 11. Selecting the classification based on the first and second intensity data comprises selecting the classification based on the combined intensity of the first and second signal components and the combined intensity of the third and fourth signal components.
The method may then proceed to block 1930, where sequence information of the respective first and second nucleobases is determined based on the classification selected in block 1920. The signals generated during a cycle of a sequencing are indicative of the identity of the nucleobase(s) added during sequencing (e.g. using sequencing-by-synthesis). For example, it may be determined that there is a match or a mismatch between the respective first and second nucleobases. Where it is determined that there is a match between the first and second respective nucleobases, the nucleobases may be base called. Whether there is a match or a mismatch, additional or alternative information may be obtained, as described above. It will be appreciated that there is a direct correspondence between the identity of the nucleobases that are incorporated and the identity of the complementary base at the corresponding position of the template sequence bound to the solid support. Therefore, any references herein to the base calling of respective nucleobases at the two portions encompasses the base calling of nucleobases hybridised to the template sequences and, alternatively or additionally, the identification of the corresponding nucleobases of the template sequences. The method may then end at block 1940.
Solid supports enabling ringed complementary reads and methods of preparing polynucleotide sequences for identification
The present invention is directed to solid supports that enable ringed complementary reads and methods of preparing polynucleotide sequences for identification using the solid support. The
solid support comprises first immobilised primers and second immobilised primers, wherein some of the first immobilised primers are cleavable and unblocked, some of the first immobilised primers are non-cleavable and blocked, some of the second immobilised primers are cleavable and blocked, and some of the second immobilised primers are non-cleavable and unblocked. Advantageously, this configuration of first immobilised primers and second immobilised primers allows the generation of solid supports that have a central read portion (also referred to herein as a “first region”), and a read portion surrounding the central read portion (also referred to herein as a “second region”). Such solid supports are tuneable as the number of amplification cycles used for generating the first region and the second region can be carefully controlled.
For example, in some cases, a lower number of amplification cycles may be used for generating the surrounding second region, compared to the number of amplification cycles used for generating the central first region. In other cases, a higher number of amplification cycles may be used for generating the surrounding second region, compared to the number of amplification cycles used for generating the central first region. In both these cases, this generates a difference in intensity of signal that can be emitted by the central first region compared to the surrounding second region. Such a difference in intensity of signal can be decoded by data analysis methods as described herein, and as such polynucleotide sequences located in the central first region and polynucleotide sequences located in the surrounding second region can be concurrently sequenced.
In other cases, the same number of amplification cycles may be used for generating the surrounding second region and the central first region. Again, information regarding polynucleotide sequences located in the central first region and polynucleotide sequences located in the surrounding second region can be determined after concurrent sequencing.
Accordingly, we describe a solid support, comprising: a plurality of first immobilised primers, and a plurality of second immobilised primers, wherein a first proportion of the first immobilised primers are configured to be cleavable under first cleavage conditions and are unblocked, wherein a second proportion of the first immobilised primers are configured to be non-cleavable under first cleavage conditions and are blocked, wherein a third proportion of the second immobilised primers are configured to be cleavable under second cleavage conditions and are blocked; and wherein a fourth proportion of the second immobilised primers are configured to be non-cleavable under second cleavage conditions and are unblocked.
In one embodiment, the second immobilised primer is different in sequence to the first immobilised primer.
The total population of the plurality of first immobilised primers may consist of the first proportion of the first immobilised primers and the second proportion of the first immobilised primers. In other words, the first immobilised primers may either be configured to be cleavable under first cleavage conditions and are unblocked (i.e. members of the first proportion), or if not, then may be configured to be non-cleavable under first cleavage conditions and are blocked (i.e. members of the second proportion).
The total population of the plurality of second immobilised primers may consist of the third proportion of the second immobilised primers and the fourth proportion of the second immobilised primers. In other words, the second immobilised primers may either be configured to be cleavable under second cleavage conditions and are blocked (i.e. members of the third proportion), or if not, then may be configured to be non-cleavable under second cleavage conditions and are unblocked (i.e. members of the fourth proportion).
In one embodiment, a ratio between unblocked immobilised primers and blocked immobilised primers may be between 20:80 to 80:20; in a further embodiment, between 50:50 to 75:25; in an even further embodiment, between 60:40 to 70:30; and in a yet even further embodiment, about 2: 1.
Such a ratio may advantageously provide a central first region that is able to output a signal that is intense enough to read during sequencing.
In one embodiment, the first proportion of first immobilised primers relative to the total population of first immobilised primers may be between 0.2 to 0.8; in a further embodiment, between 0.5 to 0.75; in an even further embodiment, between 0.6 to 0.7; in a yet even further embodiment, about %.
In one embodiment, the second proportion of first immobilised primers relative to the total population of first immobilised primers may be between 0.8 to 0.2; in a further embodiment, between 0.5 to 0.25; in an even further embodiment, between 0.4 to 0.3; in a yet even further embodiment, about 'A. In particular, the first proportion of first immobilised primers relative to the total population of first immobilised primers may be between 0.2 to 0.8, 0.5 to 0.75 (in a further embodiment), 0.6 to 0.7 (in an even further embodiment), or about % (in a yet even further
embodiment), whilst the respective second proportion of first immobilised primers relative to the total population of first immobilised primers may be between 0.8 to 0.2, 0.5 to 0.25 (in the further embodiment), 0.4 to 0.3 (in the even further embodiment), or about V (in the yet even further embodiment) (wherein the first proportion of first immobilised primers and the second proportion of first immobilised primers sums to 1).
In one embodiment, the fourth proportion of second immobilised primers relative to the total population of second immobilised primers may be between 0.2 to 0.8; in a further embodiment, between 0.5 to 0.75; in an even further embodiment, between 0.6 to 0.7; in a yet even further embodiment, about %.
In one embodiment, the third proportion of second immobilised primers relative to the total population of second immobilised primers may be between 0.8 to 0.2; in a further embodiment, between 0.5 to 0.25; in an even further embodiment, between 0.4 to 0.3; in a yet even further embodiment, about 'A. In particular, the fourth proportion of second immobilised primers relative to the total population of second immobilised primers may be between 0.2 to 0.8, 0.5 to 0.75 (in a further embodiment), 0.6 to 0.7 (in an even further embodiment), about % (in a yet even further embodiment), whilst the respective third proportion of second immobilised primers relative to the total population of second immobilised primers may be between 0.8 to 0.2, 0.5 to 0.25 (in the further embodiment), 0.4 to 0.3 (in the even further embodiment), or about (in the yet even further embodiment) (wherein the fourth proportion of second immobilised primers and the third proportion of second immobilised primers sums to 1).
In one embodiment, the total population of the first immobilised primers may be equal to the total population of the second immobilised primers.
In one embodiment, the first proportion of first immobilised primers (i.e. cleavable, unblocked first immobilised primers) may be equal to the fourth proportion of second immobilised primers (i.e. non-cleavable, unblocked second immobilised primers).
In one embodiment, the second proportion of first immobilised primers (i.e. non-cleavable, blocked first immobilised primers) may be equal to the third proportion of second immobilised primers (i.e. cleavable, blocked second immobilised primers).
In one embodiment, the first proportion of first immobilised primers (i.e. cleavable, unblocked first immobilised primers) may be equal to the fourth proportion of second immobilised primers (i.e. non-cleavable, unblocked second immobilised primers), and the second proportion of first
immobilised primers (i.e. non-cleavable, blocked first immobilised primers) may be equal to the third proportion of second immobilised primers (i.e. cleavable, blocked second immobilised primers).
By “blocked” is meant that the immobilised primer comprises a blocking group at a 3’ end of the immobilised primer. Suitable blocking groups include a hairpin loop (e.g. a polynucleotide attached to the 3 ’-end, comprising in a 5’ to 3’ direction, a cleavable site such as a nucleotide comprising uracil, a loop portion, and a complement portion, wherein the complement portion is substantially complementary to all or a portion of the immobilised primer), a deoxynucleotide, a deoxyribonucleotide, a hydrogen atom instead of a 3 ’-OH group, a phosphate group, a phosphorothioate group, a propyl spacer (e.g. -O-(CH2)3-OH instead of a 3 ’-OH group), a modification blocking the 3 ’-hydroxyl group (e.g. hydroxyl protecting groups, such as silyl ether groups (e.g. trimethylsilyl, triethylsilyl, triisopropylsilyl, t-butyl(dimethyl)silyl, t- butyl(diphenyl)silyl), ether groups (e.g. benzyl, allyl, t-butyl, methoxymethyl (MOM), 2- methoxyethoxymethyl (MEM), tetrahydropyranyl), or acyl groups (e.g. acetyl, benzoyl)), or an inverted nucleobase. However, the blocking group may be any modification that prevents extension (i.e. elongation) of the primer by a polymerase.
As such, the first immobilised primers of the second proportion that are blocked may each comprise a blocking group at a 3 ’ end of the first immobilised primer. In particular, the blocking group may be selected from the group consisting of: a hairpin loop (e.g. a polynucleotide attached to the 3 ’-end, comprising in a 5’ to 3’ direction, a cleavable site such as a nucleotide comprising uracil, a loop portion, and a complement portion, wherein the complement portion is substantially complementary to all or a portion of the immobilised primer), a deoxynucleotide, a deoxyribonucleotide, a hydrogen atom instead of a 3 ’-OH group, a phosphate group, a phosphorothioate group, a propyl spacer (e.g. -O-(CH2)3-OH instead of a 3 ’-OH group), a modification blocking the 3 ’-hydroxyl group (e.g. hydroxyl protecting groups, such as silyl ether groups (e.g. trimethylsilyl, triethylsilyl, triisopropylsilyl, t-butyl(dimethyl)silyl, t- butyl(diphenyl)silyl), ether groups (e.g. benzyl, allyl, t-butyl, methoxymethyl (MOM), 2- methoxyethoxymethyl (MEM), tetrahydropyranyl), or acyl groups (e.g. acetyl, benzoyl)), or an inverted nucleobase. In a further embodiment, the blocking group may be a phosphate group or a hairpin loop (e.g. a polynucleotide attached to the 3 ’-end, comprising in a 5’ to 3’ direction, a cleavable site such as a nucleotide comprising uracil, a loop portion, and a complement portion, wherein the complement portion is substantially complementary to all or a portion of the immobilised primer). In an even further embodiment, the blocking group may be a phosphate group.
Similarly, the second immobilised primers of the third proportion that are blocked may each comprise a blocking group at a 3’ end of the second immobilised primer. In particular, the blocking group may be selected from the group consisting of: a hairpin loop (e.g. a polynucleotide attached to the 3 ’-end, comprising in a 5’ to 3’ direction, a cleavable site such as a nucleotide comprising uracil, a loop portion, and a complement portion, wherein the complement portion is substantially complementary to all or a portion of the immobilised primer), a deoxynucleotide, a deoxyribonucleotide, a hydrogen atom instead of a 3 ’-OH group, a phosphate group, a phosphorothioate group, a propyl spacer (e.g. -O-(CH2)3-OH instead of a 3 ’-OH group), a modification blocking the 3 ’-hydroxyl group (e.g. hydroxyl protecting groups, such as silyl ether groups (e.g. trimethylsilyl, triethylsilyl, triisopropylsilyl, t-butyl(dimethyl)silyl, t- butyl(diphenyl)silyl), ether groups (e.g. benzyl, allyl, t-butyl, methoxymethyl (MOM), 2- methoxyethoxymethyl (MEM), tetrahydropyranyl), or acyl groups (e.g. acetyl, benzoyl)), or an inverted nucleobase. In a further embodiment, the blocking group may be a phosphate group or a hairpin loop (e.g. a polynucleotide attached to the 3 ’-end, comprising in a 5’ to 3’ direction, a cleavable site such as a nucleotide comprising uracil, a loop portion, and a complement portion, wherein the complement portion is substantially complementary to all or a portion of the immobilised primer). In an even further embodiment, the blocking group may be a phosphate group.
The location at which the first immobilised primer of the first proportion is configured to be cleavable under first cleavage conditions may also be referred to as a first cleavage site. As such, the first immobilised primer of the first proportion comprises a first cleavage site. The first cleavage site may comprise a cleavable covalent bond. In some cases, when the first cleavage site is nicked, this allows sequencing to occur starting from the nick location (e.g. in conjunction with a strand displacement polymerase). In some cases, when the first cleavage site is nicked, this allows linearisation to occur since one side of a “bridge” formed during bridge amplification can be detached from a solid support. In such linearisation cases, the first cleavage site may also be referred to as a first linearisation site (or that the first immobilised primer of the first proportion is configured to be linearisable under first linearisation conditions).
The location at which the second immobilised primer of the third proportion is configured to be cleavable under second cleavage conditions may also be referred to as a second cleavage site. As such, the second immobilised primer of the third proportion comprises a second cleavage site. The second cleavage site may comprise a cleavable covalent bond. In some cases, when the second cleavage site is nicked, this allows sequencing to occur starting from the nick location (e.g. in conjunction with a strand displacement polymerase). In some cases, when the second cleavage site is nicked, this allows linearisation to occur since one side of a “bridge” formed
during bridge amplification can be detached from a solid support. In such linearisation cases, the second cleavage site may also be referred to as a second linearisation site (or that the second immobilised primer of the third proportion is configured to be linearisable under second linearisation conditions).
As used herein, the term “first cleavage conditions” refers to reaction conditions that cause cleavage within the first immobilised primer of the first proportion (i.e. at the first cleavage site). The first cleavage conditions may involve exposure to a thermal trigger (e.g. by heating), a light trigger (e.g. by exposure to ultraviolet light), and/or a chemical/biochemical trigger (e.g. an enzyme, a metal catalyst including transition metal catalysts such as a palladium-based or a nickel-based catalyst, periodate). In some cases, the “first cleavage conditions” may allow linearisation to occur, and may be referred to as “first linearisation conditions”.
Accordingly, in an embodiment, the first immobilised primers of the first proportion may be configured to be cleavable by a thermal trigger (e.g. by heating), a light trigger (e.g. by exposure to ultraviolet light), and/or a chemical/biochemical trigger (e.g. an enzyme, a metal catalyst including transition metal catalysts such as a palladium-based or a nickel-based catalyst, periodate).
In one embodiment, the first immobilised primers of the first proportion may be configured to be cleavable by a metal catalyst. In other words, the first cleavage conditions involve exposure to a metal catalyst. In a further embodiment, the metal catalyst is a transition metal catalyst. In an even further embodiment, the metal catalyst is a palladium-based or a nickel-based catalyst. In a yet even further embodiment, the metal catalyst is a palladium-based catalyst (e.g. a Pd(0) catalyst). Non-limiting examples of suitable palladium-based catalysts are described in WO 2019/222264 Al (which is incorporated herein by reference), such as Pd(THP)2, Pd(THP)s, Pd(THP)4, Pd(THM)4 (where THP is tris(hydroxypropyl)phosphine and where THM is tris(hydroxymethyl)phosphine). Such palladium-based catalysts may be generated in situ from corresponding Pd(0) or Pd(II) complexes.
In one embodiment, each first immobilised primer of the first proportion may comprise a nucleotide that comprises an allyl group, for example an allyl group attached to a sugar group (e.g. deoxyribose or ribose) of the nucleotide. In a further embodiment, the allyl group may have a structure according to the following:
wherein R represents an attachment point to hydrogen (in which case, R is hydrogen) or the remainder of the sugar group (e.g. deoxyribose or ribose). In an even further embodiment, the nucleotide may have a structure according to the following:
wherein represents an attachment point of the nucleotide to the rest of the first immobilised primer of the first proportion, and wherein “base” represents adenine, guanine, cytosine, thymine or uracil, or a derivative thereof. In a yet even further embodiment, the nucleotide may have a structure according to the following:
wherein represents an attachment point of the nucleotide to the rest of the first immobilised primer of the first proportion.
In one embodiment, the first immobilised primers of the first proportion may be configured to be cleavable by a glycosylase. In other words, the first cleavage conditions involve exposure to a glycosylase. In a further embodiment, the first proportion of first immobilised primers may be configured to be cleavable by a glycosylase that recognises any nitrogenous base (e.g. purine or pyrimidine) which is not selected from guanine (G), cytosine (C), adenine (A) and thymine (T) when the first immobilised primer is a DNA sequence; or the first proportion of first immobilised primers may be configured to be cleavable by a glycosylase that recognises any nitrogenous base (e.g. purine or pyrimidine) which is not selected from guanine (G), cytosine (C), adenine (A) and uracil (U) when the first immobilised primer is an RNA sequence. In other words, the glycosylase may recognise an unnatural nucleobase (i.e. one which is not usually present in a typical DNA
sequence an RNA sequence). Examples of unnatural nucleobases may include oxoguanine (e.g. 8 -oxoguanine), hypoxanthine, xanthine, methylguanines (e.g. O6-methylguanine, N7- methylguanine), methyladenines (e.g. 3 -methyladenine, N6-methyladenine), modified cytosines including methylcytosines (e.g. 5-methylcytosine, 5-hydroxymethylcytosine, 5-formylcytosine, 5 -carboxylcytosine), dihydrouracil, inosine, and uracil (if the first immobilised primer is a DNA sequence).
In an even further embodiment, the first immobilised primers of the first proportion may be configured to be cleavable by a uracil glycosylase (when the first immobilised primer is a DNA sequence) or an oxoguanine glycosylase (e.g. 8-oxoguanine glycosylase).
In one embodiment, each first immobilised primer of the first proportion may comprise a nucleobase which is not selected from guanine, cytosine, adenine or thymine when the first immobilised primer is a DNA sequence; or wherein each first immobilised primer of the first proportion may comprise a nucleobase which is not selected from guanine, cytosine, adenine or uracil when the first immobilised primer is an RNA sequence. In other words, each first immobilised primer of the first proportion may comprise an unnatural nucleobase (i.e. one which is not usually present in a typical DNA sequence an RNA sequence). As mentioned above, examples of unnatural nucleobases may include oxoguanine (e.g. 8-oxoguanine), hypoxanthine, xanthine, methylguanines (e.g. O6-methylguanine, N7-methylguanine), methyladenines (e.g. 3- methyladenine, N6-methyladenine), modified cytosines including methylcytosines (e.g. 5- methylcytosine, 5-hydroxymethylcytosine, 5-formylcytosine, 5 -carboxylcytosine), dihydrouracil, inosine, and uracil (if the first immobilised primer is a DNA sequence).
In a further embodiment, each first immobilised primer of the first proportion may comprise oxoguanine (e.g. 8-oxoguanine) or uracil when the first immobilised primer is a DNA sequence, or wherein each first immobilised primer of the first proportion may comprise oxoguanine (e.g. 8-oxoguanine) when the first immobilised primer is an RNA sequence.
As used herein, the term “second cleavage conditions” refers to reaction conditions that cause cleavage within the second immobilised primer of the third proportion (i.e. at the second cleavage site). The second cleavage conditions may involve exposure to athermal trigger (e.g. by heating), a light trigger (e.g. by exposure to ultraviolet light), and/or a chemical/biochemical trigger (e.g. an enzyme, a metal catalyst including transition metal catalysts such as a palladium-based or a nickel-based catalyst, periodate). In some cases, the “second cleavage conditions” may allow linearisation to occur, and may be referred to as “second linearisation conditions”.
Accordingly, in an embodiment, the second immobilised primers of the third proportion may be configured to be cleavable by a thermal trigger (e.g. by heating), a light trigger (e.g. by exposure to ultraviolet light), and/or a chemical/biochemical trigger (e.g. an enzyme, a metal catalyst including transition metal catalysts such as a palladium-based or a nickel-based catalyst, periodate).
In one embodiment, the second immobilised primers of the third proportion may be configured to be cleavable by a metal catalyst. In other words, the first cleavage conditions involve exposure to a metal catalyst. In a further embodiment, the metal catalyst is a transition metal catalyst. In an even further embodiment, the metal catalyst is a palladium-based or a nickel-based catalyst. In a yet even further embodiment, the metal catalyst is a palladium-based catalyst (e.g. a Pd(0) catalyst). Non-limiting examples of suitable palladium-based catalysts are described in WO 2019/222264 Al (which is incorporated herein by reference), such as Pd(THP)2, Pd(THP)s, Pd(THP)4, Pd(THM)4 (where THP is tris(hydroxypropyl)phosphine and where THM is tris(hydroxymethyl)phosphine). Such palladium-based catalysts may be generated in situ from corresponding Pd(0) or Pd(II) complexes.
In one embodiment, each second immobilised primer of the third proportion may comprise a nucleotide that comprises an allyl group, for example an allyl group attached to a sugar group (e.g. deoxyribose or ribose) of the nucleotide. In a further embodiment, the allyl group may have a structure according to the following:
wherein R represents an attachment point to hydrogen (in which case, R is hydrogen) or the remainder of the sugar group (e.g. deoxyribose or ribose). In an even further embodiment, the nucleotide may have a structure according to the following:
wherein represents an attachment point of the nucleotide to the rest of the second immobilised primer of the third proportion, and wherein “base” represents adenine, guanine,
cytosine, thymine or uracil, or a derivative thereof. In a yet even further embodiment, the nucleotide may have a structure according to the following:
wherein represents an attachment point of the nucleotide to the rest of the second immobilised primer of the third proportion.
In one embodiment, the second immobilised primers of the third proportion may be configured to be cleavable by a glycosylase. In other words, the second cleavage conditions involve exposure to a glycosylase. In a further embodiment, the third proportion of second immobilised primers may be configured to be cleavable by a glycosylase that recognises any nitrogenous base (e.g. purine or pyrimidine) which is not selected from guanine (G), cytosine (C), adenine (A) and thymine (T) when the second immobilised primer is a DNA sequence; or the third proportion of second immobilised primers may be configured to be cleavable by a glycosylase that recognises any nitrogenous base (e.g. purine or pyrimidine) which is not selected from guanine (G), cytosine (C), adenine (A) and uracil (U) when the second immobilised primer is an RNA sequence. In other words, the glycosylase may recognise an unnatural nucleobase (i.e. one which is not usually present in a typical DNA sequence an RNA sequence). Examples of unnatural nucleobases may include oxoguanine (e.g. 8 -oxoguanine), hypoxanthine, xanthine, methylguanines (e.g. O6- methylguanine, N7-methylguanine), methyladenines (e.g. 3 -methyladenine, N6-methyladenine), modified cytosines including methylcytosines (e.g. 5 -methylcytosine, 5-hydroxymethylcytosine, 5 -formylcytosine, 5 -carboxylcytosine), dihydrouracil, inosine, and uracil (if the second immobilised primer is a DNA sequence).
In an even further embodiment, the second immobilised primers of the third proportion may be configured to be cleavable by a uracil glycosylase (when the second immobilised primer is a DNA sequence) or an oxoguanine glycosylase (e.g. 8-oxoguanine glycosylase).
In one embodiment, each second immobilised primer of the third proportion may comprise a nucleobase which is not selected from guanine, cytosine, adenine or thymine when the second
immobilised primer is a DNA sequence, or wherein each second immobilised primer of the third proportion may comprise a nucleobase which is not selected from guanine, cytosine, adenine or uracil when the second immobilised primer is an RNA sequence. In other words, each second immobilised primer of the third proportion may comprise an unnatural nucleobase (i.e. one which is not usually present in a typical DNA sequence an RNA sequence). As mentioned above, examples of unnatural nucleobases may include oxoguanine (e.g. 8 -oxoguanine), hypoxanthine, xanthine, methylguanines (e.g. O6-methylguanine, N7-methylguanine), methyladenines (e.g. 3- methyladenine, N6-methyladenine), modified cytosines including methylcytosines (e.g. 5- methylcytosine, 5-hydroxymethylcytosine, 5-formylcytosine, 5 -carboxylcytosine), dihydrouracil, inosine, and uracil (if the second immobilised primer is a DNA sequence).
In a further embodiment, each second immobilised primer of the third proportion may comprise oxoguanine (e.g. 8-oxoguanine) or uracil when the second immobilised primer is a DNA sequence, or wherein each second immobilised primer of the third proportion may comprise oxoguanine (e.g. 8-oxoguanine) when the second immobilised primer is an RNA sequence.
In one embodiment, each first immobilised primer of the first proportion and each second immobilised primer of the third proportion may comprise a nucleotide that comprises an allyl group, for example an allyl group attached to a sugar group (e.g. deoxyribose or ribose) of the nucleotide. In a further embodiment, the allyl group may have a structure according to the following:
wherein R represents an attachment point to hydrogen (in which case, R is hydrogen) or the remainder of the sugar group (e.g. deoxyribose or ribose). In an even further embodiment, the nucleotide may have a structure according to the following:
wherein represents an attachment point of the nucleotide to the rest of the first immobilised primer of the first proportion or the rest of the second immobilised primer of the third proportion, and wherein “base” represents adenine, guanine, cytosine, thymine or uracil, or a derivative
thereof. In a yet even further embodiment, the nucleotide may have a structure according to the following:
wherein represents an attachment point of the nucleotide to the rest of the first immobilised primer of the first proportion or the rest of the second immobilised primer of the third proportion.
As used herein, the term “glycosylase” may refer to an enzyme which catalyses the removal of a nitrogenous base from one of the nucleotides in a (poly)nucleotide chain by breaking a N- glycosidic bond, resulting in the formation of an apurinic/apyrimidinic site (AP site). For DNA chains, the glycosylase may recognise any nitrogenous base (e.g. purine or pyrimidine) which is not selected from guanine (G), cytosine (C), adenine (A) and thymine (T); for RNA chains, the glycosylase may recognise any nitrogenous base (e.g. purine or pyrimidine) which is not selected from guanine (G), cytosine (C), adenine (A) and uracil (U). Examples of typical nitrogenous bases recognised by glycosylases include oxoguanine (e.g. 8-oxoguanine), uracil and alkylpurines.
Glycosylases may be monofunctional, such that they only possess glycosylase activity (i.e. breaking of the N-glycosidic bond) - cleavage of a phosphodiester bond in the sugar-phosphate backbone may then occur in an uncatalysed manner by elimination. Other glycosylases may be bifimctional, such that they also possess AP lyase activity by catalysing the phosphodiester bond of the (poly)nucleotide chain. In one embodiment, the glycosylase is bifunctional (i.e. possesses both glycosylase and AP lyase activity).
In one embodiment, the first cleavage conditions and the second cleavage conditions may be the same or may be different. In some embodiments, the first cleavage conditions and the second cleavage conditions may be the same. This allows the cleavage within the first immobilised primers and the second immobilised primers to occur using a single exposure, which reduces the number of steps required for preparing first polynucleotide sequences and second polynucleotide sequences for concurrent sequencing. In alternative embodiments, the first cleavage conditions and the second cleavage conditions may be different. This allows control of which of the first
immobilised primer and/or the second immobilised primer become cleaved, should this be necessary during the preparation processes.
In one embodiment, the first proportion of first immobilised primers that are cleavable may be further configured to be linearisable under first linearisation conditions.
In one embodiment, the third proportion of second immobilised primers that are cleavable may be further configured to be linearisable under second linearisation conditions.
In one embodiment, the first proportion of first immobilised primers that are cleavable may be further configured to be linearisable under first linearisation conditions; and the third proportion of second immobilised primers that are cleavable may be further configured to be linearisable under second linearisation conditions.
In one embodiment, the first linearisation conditions and the second linearisation conditions may be the same or may be different. In some embodiments, the first linearisation conditions and the second linearisation conditions may be the same. This allows the linearisation within the first immobilised primers and the second immobilised primers to occur using a single exposure, which reduces the number of steps required for preparing first polynucleotide sequences and second polynucleotide sequences for concurrent sequencing. In alternative embodiments, the first linearisation conditions and the second linearisation conditions may be different. This allows control of which of the first immobilised primer and/or the second immobilised primer become linearised, should this be necessary during the preparation processes.
In one embodiment, each first immobilised primer may comprise a sequence as defined in SEQ ID NO. 1 or 5, or a variant or fragment thereof; and each second immobilised primer may comprise a sequence as defined in SEQ ID NO. 2, or a variant or fragment thereof; or wherein each first immobilised primer may comprise a sequence as defined in SEQ ID NO. 2, or a variant or fragment thereof; and each second immobilised primer may comprise a sequence as defined in SEQ ID NO. 1 or 5, or a variant or fragment thereof.
The solid support described above comprising the first proportion of first immobilised primers, the second proportion of first immobilised primers, the third proportion of second immobilised primers and the fourth proportion of second immobilised primers may be considered to be a precursor solid support that, once processed according to the methods described herein, forms a solid support that comprises a central first region and a surrounding second region.
Accordingly, another aspect of the present invention is directed to a solid support, comprising: a plurality of first immobilised primers, and a plurality of second immobilised primers, wherein the plurality of first immobilised primers are located in a first region, wherein the plurality of second immobilised primers are located in a second region surrounding the first region.
In one embodiment, the second immobilised primer is different in sequence to the first immobilised primer.
In one embodiment, the second region may be contiguous.
In one embodiment, the second region may be an annular region. In particular, the second region may be an annular region surrounding a circular first region.
In one embodiment, the solid support may comprise a plurality of first regions with a respective plurality of second regions surrounding the first regions. Each first region and second region may be considered to be a “well” on the solid support.
In some embodiments, the plurality of first immobilised primers in the first region may be extended to form a plurality of template sequences. In some embodiments, the plurality of second immobilised primers in the second region may be extended to form a plurality of template complement sequences. In some embodiments, the plurality of first immobilised primers in the first region may be extended to form a plurality of template sequences, and the plurality of second immobilised primers in the second region may be extended to form a plurality of template complement sequences.
In other cases, the template sequences and the template complement sequences may have been removed from the solid support. Accordingly, in some embodiments, the plurality of first immobilised primers in the first region may not be extended. In some embodiments, the plurality of second immobilised primers in the second region may not be extended. In some embodiments, the plurality of first immobilised primers in the first region may not be extended, and the plurality of second immobilised primers in the second region may not be extended.
Depending on the number of amplification cycles used to generate the first region and the second region, the concentration of first immobilised primers in the first region may be the same or different compared to the concentration of second immobilised primers in the second region.
When the concentrations are different, this generates a difference in intensity of signal that can be emitted by the central first region compared to the surrounding second region, which allows polynucleotide sequences located in the central first region and the surrounding second region to be concurrently sequenced (in particular, to determine sequence information, mismatched base pairs, or other information in the polynucleotide sequences including epigenetic modifications). When the concentrations are the same, information can still be obtained using concurrent sequencing (e.g. mismatched base pairs, or other information in the polynucleotide sequences including epigenetic modifications).
In an embodiment, a concentration of first immobilised primers in the first region may be greater than a concentration of second immobilised primers in the second region. In a further embodiment, a ratio between the concentration of first immobilised primers in the first region and the concentration of second immobilised primers in the second region may be between 1.25: 1 to 5 : 1 ; in an even further embodiment, between 1.5 : 1 to 3 : 1 ; in a yet even further embodiment, about 2: 1.
In an alternative embodiment, a concentration of first immobilised primers in the first region may be less than a concentration of second immobilised primers in the second region. In a further embodiment, a ratio between the concentration of first immobilised primers in the first region and the concentration of second immobilised primers in the second region may be between 1: 1.25 to 1:5; in an even further embodiment, between 1: 1.5 to 1:3; in a yet even further embodiment, about 1:2.
In yet another embodiment, a concentration of first immobilised primers in the first region may be equal or substantially equal to a concentration of second immobilised primers in the second region.
As mentioned above, solid supports as described herein (whether it is a precursor solid support comprising first proportions, second proportions, third proportions and fourth proportions as described herein; or whether it is a solid support comprising a central first region and a surrounding second region as described herein) are useful in nucleic acid sequencing, particularly concurrent sequencing.
Accordingly, in another aspect of the present invention, there is provided a use of a solid support as described herein in nucleic acid sequencing.
In another aspect of the present invention, there is provided a process of manufacturing a solid support, comprising:
(a) immobilising a plurality of first precursor primers onto a solid support to form a plurality of first immobilised primers, wherein a first proportion of the first precursor primers are configured to be cleavable under first cleavage conditions and are unblocked, and wherein a second proportion of the first precursor primers are configured to be non-cleavable under first cleavage conditions and are blocked; and
(b) immobilising a plurality of second precursor primers onto the solid support to form a plurality of second immobilised primers, wherein a third proportion of the second precursor primers are configured to be cleavable under second cleavage conditions and are blocked, and wherein a fourth proportion of the second precursor primers are configured to be non-cleavable under second cleavage conditions and are unblocked.
In one embodiment, the second precursor primer is different in sequence to the first precursor primer.
The solid support to be manufactured may be a solid support as described herein (in particular, a precursor solid support comprising first proportions, second proportions, third proportions and fourth proportions as described herein). Accordingly, aspects relating to the solid support and other characteristics of the solid support (such as the first immobilised primers and the second immobilised primers) as described herein apply equally to processes as described herein for manufacturing the solid support.
The term “first precursor primer” refers to a state of the first immobilised primers of the solid support before they are immobilised to the solid support. As such, the first precursor primers may be provided as “free” primers in solution. After immobilisation, the “first precursor primers” are then referred to as “first immobilised primers”.
The term “second precursor primer” refers to a state of the second immobilised primers of the solid support before they are immobilised to the solid support. As such, the second precursor primers may be provided as “free” primers in solution. After immobilisation, the “second precursor primers” are then referred to as “second immobilised primers”.
Steps (a) and (b) may be conducted sequentially or simultaneously.
For example, in one embodiment, where steps (a) and (b) are conducted sequentially, step (b) may be conducted after step (a). Alternatively, step (a) may be conducted after step (b).
In one embodiment, steps (a) and (b) may be conducted simultaneously.
The immobilisation method is not particularly limited provided that the first immobilised primers and the second immobilised primers remain on the solid support during amplification, clustering and sequencing.
In one embodiment, immobilisation may comprise forming covalent linkages between the solid support and each of the plurality of first precursor primers, and between the solid support and each of the plurality of second precursor primers. In a further embodiment, the forming covalent linkages involves using a click reaction (e.g. metal-catalysed azide-alkyne cycloaddition reactions, such as copper-catalysed azide-alkyne cycloaddition reactions and strain-promoted azide-alkyne cycloadditions).
In particular, forming covalent linkages may involve forming a 1,2,3-triazole linkage. The solid support prior to immobilisation may include azide moieties (e.g. PAZAM), whilst the first precursor primers and the second precursor primers may each comprise alkyne moieties (e.g. terminal alkynes, cycloalkynes). A click reaction between the azide moieties on the solid support and the alkyne moieties on the first precursor primers and the second precursor primers allows a 1,2,3-triazole linkage to be formed. The configuration of azide moieties and alkyne moieties can also be swapped, for example by including alkyne moieties on the solid support prior to immobilisation, and including azide moieties on each of the first precursor primers and the second precursor primers.
The solid supports as described herein (in particular, a precursor solid support comprising first proportions, second proportions, third proportions and fourth proportions as described herein) may be useful in methods of preparing polynucleotide sequences for identification.
Accordingly, in another aspect of the present invention, there is provided a method of preparing polynucleotide sequences for identification, comprising: providing a solid support as described herein, and synthesising a plurality of template sequences that extend from the second immobilised primers in the fourth proportion and a plurality of template complement sequences that extend from the first immobilised primers in the first proportion.
Typical steps that are used in the method of preparing polynucleotide sequences for identification are shown in Figure 14.
By “identification” is meant here obtaining genetic information from the polynucleotide strands. This may include identification of the genetic sequence of the polynucleotide strands (i.e. sequencing). Furthermore, this may instead, or additionally, include identification of mismatched base pairs. In addition, this may instead, or additionally, include identification of any epigenetic modifications, for example methylation. Accordingly, “identification” may mean identification of the genetic sequence of the polynucleotide strands, mismatched base pairs, and/or identification of any epigenetic modifications.
The present invention can be applied to (separate) polynucleotide strands where a first strand comprises a first portion to be identified and a second strand comprises a second portion to be identified. Thus, the polynucleotide strands comprising first portions (also referred to as “template sequences” herein) may be located in the central first region, and the polynucleotide strands comprising second portions (also referred to as “template complement sequences” herein) may be located in the surrounding second region.
The first portions and second portions may be different polynucleotide sequences. That is, the sequences may be genetically unrelated and/or derived from different sources.
Alternatively, the first portions and second portions may be genetically related.
For example, the (separate) polynucleotide strands may comprise a first strand that comprises a first portion that may comprise (or be) the forward strand of a polynucleotide sequence (e.g. forward strand of a template), and a second strand that comprises a second portion that may comprise (or be) the reverse strand of the polynucleotide sequence (e.g. reverse strand of the template) or the forward complement strand of the polynucleotide sequence (e.g. forward complement strand of the template). As a further alternative, the (separate) polynucleotide strands may comprise a first strand that comprises a first portion that may comprise (or be) the reverse strand of a polynucleotide sequence (e.g. reverse strand of a template), and a second strand that comprises a second portion that may comprise (or be) the forward strand of the polynucleotide sequence (e.g. forward strand of the template) or the reverse complement strand of the polynucleotide sequence (e.g. reverse complement strand of the template).
Alternatively, the (separate) polynucleotide strands may comprise a first strand that comprises a first portion that may comprise (or be) the forward strand of a polynucleotide sequence (e.g. forward strand of a template), and a second strand that comprises a second portion that may comprise (or be) the reverse complement strand of the polynucleotide sequence (e.g. reverse
complement strand of the template) (in effect, a reverse complement strand may be considered a “copy” of the forward strand). As a further alternative, the (separate) polynucleotide strands may comprise a first strand that comprises a first portion that may comprise (or be) the reverse strand of a polynucleotide sequence (e.g. reverse strand of a template), and a second strand that comprises a second portion that may comprise (or be) the forward complement strand of the polynucleotide sequence (e.g. forward complement strand of the template) (in effect, a forward complement strand may be considered a “copy” of the reverse strand). In some embodiments, the first portion may be derived from a forward strand of a target polynucleotide to be sequenced, and the second portion may be derived from a reverse complement strand of the target polynucleotide to be sequenced; or the first portion may be derived from a reverse strand of a target polynucleotide to be sequenced, and the second portion may be derived from a forward complement strand of the target polynucleotide to be sequenced. In these particular embodiments, concurrent sequencing of both the forward and reverse complement strands (or the reverse and forward complement strands) allows mismatched base pairs and/or epigenetic modification to be detected.
The first portion may be referred to herein as read 1 (Rl). The second portion may be referred to herein as read 2 (R2).
In one embodiment, the first portion is at least 25 or at least 50 base pairs and the second portion is at least 25 base pairs or at least 50 base pairs.
The polynucleotide strands may form or be part of a cluster on the solid support. In particular, the cluster may refer to polynucleotide strands located within a central first region and its associated surrounding second region.
As used herein, the term “cluster” may refer to a clonal group of template polynucleotides (e.g. DNA or RNA) bound within a single well of a solid support (e.g. flow cell). As such, a cluster may refer to the population of polynucleotide molecules within a well that are then sequenced. A “cluster” may contain a sufficient number of copies of template polynucleotides such that the cluster is able to output a signal (e.g. a light signal) that allows sequencing reads to be performed on the cluster. A “cluster” may comprise, for example, about 500 to about 2000 copies, about 600 to about 1800 copies, about 700 to about 1600 copies, about 800 to about 1400 copies, about 900 to about 1200 copies, or about 1000 copies of template polynucleotides.
A cluster may be formed by bridge amplification, as described above.
The cluster formed may be a duoclonal cluster.
By “duoclonal” cluster is meant that the population of polynucleotide sequences that are then sequenced (as the next step) are substantially of two types - e.g. a first sequence and a second sequence. As such, a “duoclonal” cluster may refer to the population of single first sequences and single second sequences within a well that are then sequenced. A “duoclonal” cluster may contain a sufficient number of copies of a single first sequence and copies of a single second sequence such that the cluster is able to output a signal (e.g. a light signal) that allows sequencing reads to be performed on the “monoclonal” cluster. A “duoclonal” cluster may comprise, for example, about 500 to about 2000 combined copies, about 600 to about 1800 combined copies, about 700 to about 1600 combined copies, about 800 to about 1400 combined copies, about 900 to about 1200 combined copies, or about 1000 combined copies of single first sequences and single second sequences. The copies of single first sequences and single second sequences together may comprise at least about 50%, at least about 60%, at least about 70%, even at least about 80%, at least about 90%, or about 95%, 98%, 99% or 100% of all polynucleotides within a single well of the flow cell, and thus providing a substantially duoclonal “cluster”.
A first signal that is capable of being produced by the first portion and a second signal that is capable of being produced by the second portion may be optically unresolved.
In one embodiment, the step of synthesising the plurality of template sequences extending from the second immobilised primers in the fourth proportion and the plurality of template complement sequences extending from the first immobilised primers in the first proportion may involve conducting amplification; in a further embodiment, bridge amplification.
In one embodiment, amplification (of the plurality of template sequences extending from the second immobilised primers in the fourth proportion and the plurality of template complement sequences extending from the first immobilised primers in the first proportion) may be conducted over 20 to 40 cycles; in a further embodiment, 25 to 35 cycles.
In one embodiment, the method may further comprise a step of removing second immobilised primers in the fourth proportion that are not yet extended, and removing first immobilised primers in the first proportion that are not yet extended. The step of removing second immobilised primers in the fourth proportion that are not yet extended, and removing first immobilised primers in the first proportion that are not yet extended, may be conducted after (e.g. immediately after) the step of synthesising the plurality of template sequences that extend from the second immobilised primers in the fourth proportion and the plurality of template complement sequences that extend
from the first immobilised primers in the first proportion. In a further embodiment, the step of removing second immobilised primers in the fourth proportion that are not yet extended and removing first immobilised primers in the first proportion that are not yet extended may be conducted using a 3 ’-5’ exonuclease (e.g. Exol).
In one embodiment, the method may further comprise a step of unblocking the second proportion of first immobilised primers and unblocking the third proportion of second immobilised primers (hereinafter referred to as “unblocking conditions”). The step of unblocking the second proportion of first immobilised primers and unblocking the third proportion of second immobilised primers may be conducted after (e.g. immediately after) the step of removing second immobilised primers in the fourth proportion that are not yet extended, and removing first immobilised primers in the first proportion that are not yet extended. In particular, the step of unblocking the second proportion of first immobilised primers and unblocking the third proportion of second immobilised primers may involve removing phosphate groups or hairpin loops from the second proportion of first immobilised primers and removing phosphate groups or hairpin loops from the third proportion of second immobilised primers. In a further embodiment, the step of unblocking the second proportion of first immobilised primers and unblocking the third proportion of second immobilised primers may involve removing phosphate groups from the second proportion of first immobilised primers and removing phosphate groups from the third proportion of second immobilised primers.
In one embodiment, the unblocking conditions may be different to those of the first cleavage conditions and/or the second cleavage conditions. Thus, the step of unblocking the second proportion of first immobilised primers and unblocking the third proportion of second immobilised primers may be conducted selectively over cleaving the first proportion of first immobilised primers and/or cleaving the third proportion of second immobilised primers.
In one embodiment, the method may further comprise a step of synthesising a plurality of template sequences that extend from the second immobilised primers in the third proportion and a plurality of template complement sequences that extend from the first immobilised primers in the second proportion. The step of synthesising the plurality of template sequences that extend from the second immobilised primers in the third proportion and the plurality of template complement sequences that extend from the first immobilised primers in the second proportion may be conducted after (e.g. immediately after) the step of unblocking the second proportion of first immobilised primers and unblocking the third proportion of second immobilised primers.
In one embodiment, the step of synthesising the plurality of template sequences extending from the second immobilised primers in the third proportion and the plurality of template complement sequences extending from the first immobilised primers in the second proportion may involve conducting amplification; in a further embodiment, bridge amplification.
In one embodiment, amplification (of the plurality of template sequences extending from the second immobilised primers in the third proportion and the plurality of template complement sequences extending from the first immobilised primers in the second proportion) may be conducted over 5 to 25 cycles; in a further embodiment, 10 to 20 cycles.
As mentioned above, the number of amplification cycles used for generating the central first region and the surrounding second region can be tuned.
Accordingly, in an embodiment, the number of amplification cycles used for synthesising the plurality of template sequences extending from the second immobilised primers in the third proportion and the plurality of template complement sequences extending from the first immobilised primers in the second proportion may be less than the number of amplification cycles used for synthesising the plurality of template sequences extending from the second immobilised primers in the fourth proportion and the plurality of template complement sequences extending from the first immobilised primers in the first proportion.
In an alternative embodiment, the number of amplification cycles used for synthesising the plurality of template sequences extending from the second immobilised primers in the third proportion and the plurality of template complement sequences extending from the first immobilised primers in the second proportion may be more than the number of amplification cycles used for synthesising the plurality of template sequences extending from the second immobilised primers in the fourth proportion and the plurality of template complement sequences extending from the first immobilised primers in the first proportion.
In yet another embodiment, the number of amplification cycles used for synthesising the plurality of template sequences extending from the second immobilised primers in the third proportion and the plurality of template complement sequences extending from the first immobilised primers in the second proportion may be the same as the number of amplification cycles used for synthesising the plurality of template sequences extending from the second immobilised primers in the fourth proportion and the plurality of template complement sequences extending from the first immobilised primers in the first proportion.
In one embodiment, the method may further comprise a step of removing second immobilised primers in the third proportion that are not yet extended, and removing first immobilised primers in the second proportion that are not yet extended. The step of removing second immobilised primers in the third proportion that are not yet extended, and removing first immobilised primers in the second proportion that are not yet extended may be conducted after (e.g. immediately after) the step of synthesising the plurality of template sequences that extend from the second immobilised primers in the third proportion and the plurality of template complement sequences that extend from the first immobilised primers in the second proportion. In a further embodiment, the step of removing second immobilised primers in the third proportion that are not yet extended and removing first immobilised primers in the second proportion that are not yet extended may be conducted using a 3 ’-5’ exonuclease (e.g. Exol).
In one embodiment, the method may further comprise a step of exposing the solid support to first cleavage conditions and/or second cleavage conditions after the step of synthesising the plurality of template sequences that extend from the second immobilised primers in the third proportion and the plurality of template complement sequences that extend from the first immobilised primers in the second proportion. Suitable first cleavage conditions and second cleavage conditions are described above. In a further embodiment, the first cleavage conditions and/or second cleavage conditions may comprise exposure to a thermal trigger (e.g. by heating), a light trigger (e.g. by exposure to ultraviolet light), and/or a chemical/biochemical trigger (e.g. an enzyme, a metal catalyst including transition metal catalysts such as a palladium-based or a nickel-based catalyst, periodate).
The solid support may be exposed to the first cleavage conditions and then subsequently exposed to the second cleavage conditions, or may be exposed to the second cleavage conditions and then subsequently exposed to the first cleavage conditions (i.e. sequentially). Alternatively, the solid support may be exposed to the first cleavage conditions and the second cleavage conditions at the same time (i.e. simultaneously).
In one embodiment, the solid support may be exposed to a metal catalyst. In other words, the first cleavage conditions and/or second cleavage conditions may involve exposure to a metal catalyst. Suitable metal catalysts are described above. In a further embodiment, the metal catalyst is a transition metal catalyst. In an even further embodiment, the metal catalyst is a palladium-based or a nickel-based catalyst. In a yet even further embodiment, the metal catalyst is a palladiumbased catalyst (e.g. a Pd(0) catalyst).
In one embodiment, the solid support may be exposed to a glycosylase. In other words, the first cleavage conditions and/or second cleavage conditions may involve exposure to a glycosylase. Suitable glycosylases are described above. In a further embodiment, the solid support is exposed to a uracil glycosylase and/or an oxoguanine glycosylase (e.g. 8-oxoguanine glycosylase).
In one embodiment, the method may further comprise a step of linearising the plurality of template sequences extending from the second immobilised primers of the fourth proportion and linearising the plurality of template complement sequences extending from the first immobilised primers of the second proportion. In a further embodiment, the method may further comprise treating the linearised template sequences and the linearised template complement sequences with a single-stranded binding protein.
Methods of sequencing
Also described herein is a method of sequencing polynucleotide sequences, comprising preparing polynucleotide sequences for identification using a method as described herein; and concurrently sequencing nucleobases in the template sequences extending from the second immobilised primers of the fourth proportion and the template complement sequences extending from the first immobilised primers of the second proportion.
In one embodiment, sequencing is performed by a sequencing-by-synthesis or sequencing-by- ligation process.
In one embodiment, the step of concurrently sequencing nucleobases may comprise treatment with a strand displacement polymerase (e.g. phi29).
In one embodiment, the method may further comprise a step of conducting paired-end reads.
In some embodiments, the data may be analysed using 16 QAM as mentioned herein. For example, this may be utilised when a concentration of first immobilised primers in the first region is greater than a concentration of second immobilised primers in the second region on certain types of solid support, or when a concentration of first immobilised primers in the first region is less than a concentration of second immobilised primers in the second region on certain types of solid support.
Accordingly, the step of concurrently sequencing nucleobases may comprise:
(a) obtaining first intensity data comprising a combined intensity of a first signal component obtained based upon a respective first nucleobase at a first portion of the template sequence and a second signal component obtained based upon a respective second nucleobase at a second portion of the template complement sequence, wherein the first and second signal components are obtained simultaneously;
(b) obtaining second intensity data comprising a combined intensity of a third signal component obtained based upon the respective first nucleobase at the first portion and a fourth signal component obtained based upon the respective second nucleobase at the second portion, wherein the third and fourth signal components are obtained simultaneously;
(c) selecting one of a plurality of classifications based on the first and the second intensity data, wherein each classification represents a possible combination of respective first and second nucleobases; and
(d) based on the selected classification, base calling the respective first and second nucleobases.
In one embodiment, selecting the classification based on the first and second intensity data may comprise selecting the classification based on the combined intensity of the first and second signal components and the combined intensity of the third and fourth signal components.
In one embodiment, the plurality of classifications may comprise sixteen classifications, each classification representing one of sixteen unique combinations of first and second nucleobases.
In one embodiment, the first signal component, second signal component, third signal component and fourth signal component may be generated based on light emissions associated with the respective nucleobase.
In one example, the light emissions may be detected by a sensor, wherein the sensor is configured to provide a single output based upon the first and second signals.
In one embodiment, the sensor may comprise a single sensing element.
In one embodiment, the method may further comprise repeating steps (a) to (d) for each of a plurality of base calling cycles.
In some embodiments, the data may be analysed using 9 QAM as mentioned herein. For example, this may be utilised when a concentration of first immobilised primers in the first region is equal
or substantially equal to a concentration of second immobilised primers in the second region on certain types of solid support.
Accordingly, the step of concurrently sequencing nucleobases may comprise:
(a) obtaining first intensity data comprising a combined intensity of a first signal component obtained based upon a respective first nucleobase at a first portion of the template sequence and a second signal component obtained based upon a respective second nucleobase at a second portion of the template complement sequence, wherein the first and second signal components are obtained simultaneously;
(b) obtaining second intensity data comprising a combined intensity of a third signal component obtained based upon the respective first nucleobase at the first portion and a fourth signal component obtained based upon the respective second nucleobase at the second portion, wherein the third and fourth signal components are obtained simultaneously;
(c) selecting one of a plurality of classifications based on the first and the second intensity data, wherein each classification of the plurality of classifications represents one or more possible combinations of respective first and second nucleobases, and wherein at least one classification of the plurality of classifications represents more than one possible combination of respective first and second nucleobases; and
(d) based on the selected classification, determining sequence information from the first portion and the second portion.
In one embodiment, selecting the classification based on the first and second intensity data may comprise selecting the classification based on the combined intensity of the first and second signal components and the combined intensity of the third and fourth signal components.
In one aspect, when based on a nucleobase of the same identity, an intensity of the first signal component may be substantially the same as an intensity of the second signal component and an intensity of the third signal component is substantially the same as an intensity of the fourth signal component.
In one embodiment, the plurality of classifications may consist of a predetermined number of classifications.
In one embodiment, the plurality of classifications may comprise: one or more classifications representing matching first and second nucleobases; and
one or more classifications representing mismatching first and second nucleobases, and wherein determining sequence information of the first portion and second portion comprises: in response to selecting a classification representing matching first and second nucleobases, determining a match between the first and second nucleobases; or in response to selecting a classification representing mismatching first and second nucleobases, determining a mismatch between the first and second nucleobases.
In one embodiment, determining sequence information of the first portion and the second portion may comprise, in response to selecting a classification representing a match between the first and second nucleobases, base calling the first and second nucleobases.
In another embodiment, determining sequence information of the first portion and the second portion may comprise, based on the selected classification, determining that the second portion is modified relative to the first portion at a location associated with the first and second nucleobases.
In one example, the first signal component, second signal component, third signal component and fourth signal component may be generated based on light emissions associated with the respective nucleobase.
In one aspect, the light emissions may be detected by a sensor, wherein the sensor is configured to provide a single output based upon the first and second signals.
In one embodiment, the sensor may comprise a single sensing element.
In one embodiment, the method may further comprise repeating steps (a) to (d) for each of a plurality of base calling cycles.
Kits
Methods as described herein may be performed by a user physically. In other words, a user may themselves conduct the methods of preparing polynucleotide sequences for identification as described herein, and as such the methods as described herein may not need to be computer- implemented.
In another aspect of the invention, there is provided a kit comprising a solid support as described herein.
In another aspect of the invention, there is provided a kit comprising instructions for preparing polynucleotide sequences for identification according to the methods described herein and/or sequencing polynucleotide sequences according to the methods described herein.
Computer programs and products
In other embodiments, methods as described herein may be performed by a computer. In other words, a computer may contain instructions to conduct the methods of preparing polynucleotide sequences for identification as described herein, and as such the methods as described herein may be computer-implemented.
Accordingly, in another aspect of the invention, there is provided a data processing device comprising means for carrying out the methods as described herein.
The data processing device may be a polynucleotide sequencer.
The data processing device may comprise reagents used for methods as described herein.
The data processing device may comprise a solid support as described herein, such as a flow cell.
In another aspect of the invention, there is provided a computer program product comprising instructions which, when the program is executed by a processor, cause the processor to carry out the methods as described herein.
In another aspect of the invention, there is provided a computer-readable storage medium comprising instructions which, when executed by a processor, cause the processor to carry out the methods as described herein.
In another aspect of the invention, there is provided a computer-readable data carrier having stored thereon the computer program product as described herein.
In another aspect of the invention, there is provided a data carrier signal carrying the computer program product as described herein.
The various illustrative imaging or data processing techniques described in connection with the embodiments disclosed herein can be implemented as electronic hardware, computer software, or combinations of both. To illustrate this interchangeability of hardware and software, various illustrative components, blocks, modules, and steps have been described above generally in terms of their functionality. Whether such functionality is implemented as hardware or software depends upon the particular application and design constraints imposed on the overall system. The described functionality can be implemented in varying ways for each particular application, but such implementation decisions should not be interpreted as causing a departure from the scope of the disclosure.
The various illustrative detection systems described in connection with the embodiments disclosed herein can be implemented or performed by a machine, such as a processor configured with specific instructions, a digital signal processor (DSP), an application specific integrated circuit (ASIC), a field programmable gate array (FPGA) or other programmable logic device, discrete gate or transistor logic, discrete hardware components, or any combination thereof designed to perform the functions described herein. A processor can be a microprocessor, but in the alternative, the processor can be a controller, microcontroller, or state machine, combinations of the same, or the like. A processor can also be implemented as a combination of computing devices, e.g., a combination of a DSP and a microprocessor, a plurality of microprocessors, one or more microprocessors in conjunction with a DSP core, or any other such configuration. For example, systems described herein may be implemented using a discrete memory chip, a portion of memory in a microprocessor, flash, EPROM, or other types of memory.
The elements of a method, process, or algorithm described in connection with the embodiments disclosed herein can be embodied directly in hardware, in a software module executed by a processor, or in a combination of the two. A software module can reside in RAM memory, flash memory, ROM memory, EPROM memory, EEPROM memory, registers, hard disk, a removable disk, a CD-ROM, or any other form of computer-readable storage medium known in the art. An exemplary storage medium can be coupled to the processor such that the processor can read information from, and write information to, the storage medium. In the alternative, the storage medium can be integral to the processor. The processor and the storage medium can reside in an ASIC. A software module can comprise computer-executable instructions which cause a hardware processor to execute the computer-executable instructions.
Computer-executable instructions may be stored in a (transitory or non-transitory) computer readable storage medium (e.g., memory, storage system, etc.) storing code, or computer readable instructions.
Additional Notes
The embodiments described herein are exemplary. Modifications, rearrangements, substitute processes, etc. may be made to these embodiments and still be encompassed within the teachings set forth herein. One or more of the steps, processes, or methods described herein may be carried out by one or more processing and/or digital devices, suitably programmed.
Conditional language used herein, such as, among others, “can,” “might,” “may,” “e.g.,” and the like, unless specifically stated otherwise, or otherwise understood within the context as used, is generally intended to convey that certain embodiments include, while other embodiments do not include, certain features, elements and/or states. Thus, such conditional language is not generally intended to imply that features, elements and/or states are in any way required for one or more embodiments or that one or more embodiments necessarily include logic for deciding, with or without author input or prompting, whether these features, elements and/or states are included or are to be performed in any particular embodiment. The terms “comprising,” “including,” “having,” “involving,” and the like are synonymous and are used inclusively, in an open-ended fashion, and do not exclude additional elements, features, acts, operations, and so forth. Also, the term “or” is used in its inclusive sense (and not in its exclusive sense) so that when used, for example, to connect a list of elements, the term “or” means one, some, or all of the elements in the list. The term “comprising” may be considered to encompass “consisting”.
Disjunctive language such as the phrase “at least one of X, Y or Z,” unless specifically stated otherwise, is otherwise understood with the context as used in general to present that an item, term, etc., may be either X, Y or Z, or any combination thereof (e.g., X, Y and/or Z). Thus, such disjunctive language is not generally intended to, and should not, imply that certain embodiments require at least one of X, at least one of Y or at least one of Z to each be present.
The terms “about” or “approximate” and the like are synonymous and are used to indicate that the value modified by the term has an understood range associated with it, where the range can be ±20%, ±15%, ±10%, ±5%, or ±1%. The term “substantially” is used to indicate that a result (e.g., measurement value) is close to a targeted value, where close can mean, for example, the result is within 80% of the value, within 90% of the value, within 95% of the value, or within
99% of the value. The term “partially” is used to indicate that an effect is only in part or to a limited extent.
Unless otherwise explicitly stated, articles such as “a” or “an” should generally be interpreted to include one or more described items. Accordingly, phrases such as “a device configured to” or “a device to” are intended to include one or more recited devices. Such one or more recited devices can also be collectively configured to carry out the stated recitations. For example, “a processor to carry out recitations A, B and C” can include a first processor configured to carry out recitation A working in conjunction with a second processor configured to carry out recitations B and C.
While the above detailed description has shown, described, and pointed out novel features as applied to illustrative embodiments, it will be understood that various omissions, substitutions, and changes in the form and details of the devices or algorithms illustrated can be made without departing from the spirit of the disclosure. As will be recognized, certain embodiments described herein can be embodied within a form that does not provide all of the features and benefits set forth herein, as some features can be used or practiced separately from others. All changes which come within the meaning and range of equivalency of the claims are to be embraced within their scope.
It should be appreciated that all combinations of the foregoing concepts (provided such concepts are not mutually inconsistent) are contemplated as being part of the inventive subject matter disclosed herein. In particular, all combinations of claimed subject matter appearing at the end of this disclosure are contemplated as being part of the inventive subject matter disclosed herein.
The present invention will now be described by way of the following non-limiting examples.
Examples
Example 1 : Preparation of (precursor) solid support
Oligos: v4 P5-nonlin block
5’alkyne-TTTTTTAATGATACGGCGACCACCGATCTA*C*A*C-phosphate (SEQ ID NO. 11) v4 P7-lin block
5’alkyne-TTTTTTXCAAGCAGAAGACGGCATACGA*G*A*T-phosphate (SEQ ID NO. 12)
P5-lin non-block
5’alkyne-TTTTTTXAATGATACGGCGACCACCGATCTACAC (SEQ ID NO. 13)
P7-nonlin non-block
5’alkyne-TTTTTTCAAGCAGAAGACGGCATACGAGAT (SEQ ID NO. 14)
“*” denotes a thiophosphate linkage between last 3 nucleotides to protect the oligos from the Exo digestion step later on.
“X” denotes alkene-dT linkage, which can be linearised with 6mM palladium in THP cleavage mix:
wherein represents an attachment point of the nucleotide to the rest of the oligo.
An SFA coated, non-grafted HiSeq2000 flowcell was grafted with mixtures of oligos as detailed in the following table:
A 2: 1 ratio used oligos at 1.5pM each for the more concentrated pair, 0.75pM each for the less concentrated pair. A 1: 1 ratio used all 4 oligos at 0.75pM in the grafting mix.
1.5ml of carbonate buffer based grafting mix was made up by combining:
705 pl water
750 l IM carbonate buffer (pHl 0)
14pl PMDETA (Sigma 369497)
15 pl CuSCft (Sigma C2284)
15.4pl of freshly dissolved 500mg/ml sodium ascorbate in water
175 pl of the above buffer was spiked with the appropriate primers as detailed above and used in the grafting reaction on an Illumina cBot. The grafting mix was pumped onto the flowcell surface and incubated at 65 °C for 30mins to allow the click chemistry between the 5’ alkynes on the oligos and the free azides. After the grafting step, the flowcell lanes were washed with HT1 buffer and then oligo grafting checked via a TET-QC assay (hybridisation of TET labelled complements to the P5/P7 oligo sequences, Typhoon instrument scanning to assess the levels of TET signal in each lane). TET-QC oligos were removed by 0.1N NaOH dehyb before the flowcell was used to make clusters.
Example 2: Preparation of solid support with ringed region and central region
1. Flowcell lanes were seeded with 5pM of a TruSeq Nano E. coli library and then bridge amplified for either 20 or 30 cycles of initial amplification to establish the cluster R1 “centres”. Differing numbers of amplification cycles were achieved by programming the cBot script to pump the different cycles of amplification from different positions of the
cBot reagent plate. For instance, cycles 1 to 20 may pump the reagents LDR (formamide), LPM (water) and AMX (Bst amplification mix) from positions 5, 3 and 4 on the reagent plate, then cycles 21 to 30 would pump those reagents from positions 9, 2 and 6. For lanes which were kept at the 20 cycles amplification stage, those later reagent positions were filled with HT2 wash buffer.
2. After the initial amplification was complete, the flowcell lanes were treated with exonuclease I (NEB) for 30 mins at 37 °C, to remove excess “non-block” oligos. During this step, the standard control P5/P7 clusters in lane 1 were linearised by using USER mix instead of the Exol mix.
3. The “v4 blocked” oligo pair in lanes 2-8 were then unblocked by treatment of the flowcell lanes with T4 kinase containing reagent (PRM, “Patterned Resynthesis Mix”, Illumina) for 30 mins at 37 °C.
4. A 2nd bridge amplification was then done for either 10 or 20 cycles, again varying cycle number in a similar way to step 1. This 2nd amplification would create the cluster R2 “rings” around the R1 “centres”.
5. After the 2nd amplification was complete, all cluster DNA in lanes 2 to 8 were linearised by using the “CCL1” reagent (Cluster Chemical Linearisation 1) - a mix of 6mM palladium in standard SBS cleavage reagent (THP). This was incubated for 10 mins at 60 °C to enable linearisation at the alkene-dT residues in the “P5-lin non-block” and “v4 P7- lin block” oligos.
6. After linearisation, lanes 2-5, 7 and 8 were treated with BMX (Blocking mix, a mixture of terminal transferase, a sequencing polymerase and ddNTPs) for 30mins at 37 °C and then 15mins at 60 °C. Lanes 1 and 6 had HT2 wash buffer during this step to be a “no blocking” test. This treatment blocks any free 3 ’OH ends in the lanes.
Example 3: Sequencing using solid support with ringed region and central region
1. After blocking, the flowcell lanes were then denatured and hybed with a mixture of R1 and R2 sequencing primers.
2. The flowcell was then put onto a standard HiSeq2000 for sequencing. Images in Figure 15 are close-ups of images from this run at 1st cycle.
Full details of the various steps and amplification cycle numbers done for each lane are in the table below:
Embodiments are set out in the following clauses:
Clause 1. A solid support, comprising: a plurality of first immobilised primers, and a plurality of second immobilised primers, wherein a first proportion of the first immobilised primers are configured to be cleavable under first cleavage conditions and are unblocked, wherein a second proportion of the first immobilised primers are configured to be non-cleavable under first cleavage conditions and are blocked, wherein a third proportion of the second immobilised primers are configured to be cleavable under second cleavage conditions and are blocked; and wherein a fourth proportion of the second immobilised primers are configured to be non-cleavable under second cleavage conditions and are unblocked.
Clause 2. A solid support according to clause 1, wherein a ratio between unblocked immobilised primers and blocked immobilised primers is between 20:80 to 80:20.
Clause 3. A solid support according to clause 2, wherein the ratio between unblocked immobilised primers and blocked immobilised primers is between 50:50 to 75:25.
Clause 4. A solid support according to clause 3, wherein the ratio between unblocked immobilised primers and blocked immobilised primers is between 60:40 to 70:30.
Clause 5. A solid support according to clause 4, wherein the ratio between unblocked immobilised primers and blocked immobilised primers is about 2: 1.
Clause 6. A solid support according to any one of clauses 1 to 5, wherein the first immobilised primers of the second proportion that are blocked each comprise a blocking group at a 3 ’ end of the first immobilised primer.
Clause 7. A solid support according to clause 6, wherein the blocking group is selected from the group consisting of: a hairpin loop, a deoxynucleotide, a deoxyribonucleotide, a hydrogen atom instead of a 3 ’-OH group, a phosphate group, a phosphorothioate group, a propyl spacer, a modification blocking the 3 ’-hydroxyl group, or an inverted nucleobase.
Clause 8. A solid support according to clause 7, wherein the blocking group is a phosphate group or a hairpin loop.
Clause 9. A solid support according to clause 8, wherein the blocking group is a phosphate group.
Clause 10. A solid support according to any one of clauses 1 to 9, wherein the second immobilised primers of the third proportion that are blocked each comprise a blocking group at a 3 ’ end of the second immobilised primer.
Clause 11. A solid support according to clause 10, wherein the blocking group is selected from the group consisting of: a hairpin loop, a deoxynucleotide, a deoxyribonucleotide, a hydrogen atom instead of a 3 ’-OH group, a phosphate group, a phosphorothioate group, a propyl spacer, a modification blocking the 3 ’-hydroxyl group, or an inverted nucleobase.
Clause 12. A solid support according to clause 11, wherein the blocking group is a phosphate group or a hairpin loop.
Clause 13. A solid support according to clause 12, wherein the blocking group is a phosphate group.
Clause 14. A solid support according to any one of clauses 1 to 13, wherein the first immobilised primers of the first proportion are configured to be cleavable by a thermal trigger, a light trigger, and/or a chemical/biochemical trigger.
Clause 15. A solid support according to clause 14, wherein the first immobilised primers of the first proportion are configured to be cleavable by a metal catalyst.
Clause 16. A solid support according to clause 15, wherein the first immobilised primers of the first proportion are configured to be cleavable by a transition metal catalyst.
Clause 17. A solid support according to clause 16, wherein the first immobilised primers of the first proportion are configured to be cleavable by a palladium-based or a nickel-based catalyst.
Clause 18. A solid support according to clause 17, wherein the first immobilised primers of the first proportion are configured to be cleavable by a palladium-based catalyst.
Clause 19. A solid support according to any one of clauses 1 to 18, wherein each first immobilised primer of the first proportion comprises a nucleotide that comprises an allyl group.
Clause 20. A solid support according to clause 19, wherein the allyl group is attached to a sugar group of the nucleotide.
Clause 21. A solid support according to clause 20, wherein the allyl group has a structure according to the following:
wherein R represents an attachment point to hydrogen or the remainder of the sugar group.
Clause 22. A solid support according to clause 21, wherein the nucleotide has a structure according to the following: se
wherein represents an attachment point of the nucleotide to the rest of the first immobilised primer of the first proportion, and wherein “base” represents adenine, guanine, cytosine, thymine or uracil, or a derivative thereof.
Clause 23. A solid support according to clause 22, wherein the nucleotide has a structure according to the following:
wherein represents an attachment point of the nucleotide to the rest of the first immobilised primer of the first proportion.
Clause 24. A solid support according to clause 14, wherein the first immobilised primers of the first proportion are configured to be cleavable by a glycosylase.
Clause 25. A solid support according to any one of clauses 1 to 24, wherein each first immobilised primer of the first proportion comprises a nucleobase which is not selected from guanine, cytosine, adenine or thymine when the first immobilised primer is a DNA sequence; or wherein each first immobilised primer of the first proportion comprises a nucleobase which is not selected from guanine, cytosine, adenine or uracil when the first immobilised primer is an RNA sequence.
Clause 26. A solid support according to clause 25, wherein each first immobilised primer of the first proportion comprises oxoguanine (e.g. 8-oxoguanine) or uracil when the first immobilised primer is a DNA sequence, or wherein each first immobilised primer of the first proportion comprises oxoguanine (e.g. 8-oxoguanine) when the first immobilised primer is an RNA sequence.
Clause 27. A solid support according to any one of clauses 1 to 26, wherein the second immobilised primers of the third proportion are configured to be cleavable by a thermal trigger, a light trigger, or a chemical/biochemical trigger.
Clause 28. A solid support according to clause 27, wherein the second immobilised primers of the third proportion are configured to be cleavable by a metal catalyst.
Clause 29. A solid support according to clause 28, wherein the second immobilised primers of the third proportion are configured to be cleavable by a transition metal catalyst.
Clause 30. A solid support according to clause 29, wherein the second immobilised primers of the third proportion are configured to be cleavable by a palladium-based or a nickel-based catalyst.
Clause 31. A solid support according to clause 30, wherein the second immobilised primers of the third proportion are configured to be cleavable by a palladium-based catalyst.
Clause 32. A solid support according to any one of clauses 1 to 31, wherein each second immobilised primer of the third proportion comprises a nucleotide that comprises an allyl group.
Clause 33. A solid support according to clause 32, wherein the allyl group is attached to a sugar group of the nucleotide.
Clause 34. A solid support according to clause 33, wherein the allyl group has a structure according to the following:
wherein R represents an attachment point to hydrogen or the remainder of the sugar group.
Clause 35. A solid support according to clause 34, wherein the nucleotide has a structure according to the following:
wherein represents an attachment point of the nucleotide to the rest of the second immobilised primer of the third proportion, and wherein “base” represents adenine, guanine, cytosine, thymine or uracil, or a derivative thereof.
Clause 36. A solid support according to clause 35, wherein the nucleotide has a structure according to the following:
wherein represents an attachment point of the nucleotide to the rest of the second immobilised primer of the third proportion.
Clause 37. A solid support according to clause 27, wherein the second immobilised primers of the third proportion are configured to be cleavable by a glycosylase.
Clause 38. A solid support according to any one of clauses 1 to 37, wherein each second immobilised primer of the third proportion comprises a nucleobase which is not selected from guanine, cytosine, adenine or thymine when the second immobilised primer is a DNA sequence; or wherein each second immobilised primer of the third proportion comprises a nucleobase which is not selected from guanine, cytosine, adenine or uracil when the second immobilised primer is an RNA sequence.
Clause 39. A solid support according to clause 38, wherein each second immobilised primer of the third proportion comprises oxoguanine (e.g. 8-oxoguanine) or uracil when the second immobilised primer is a DNA sequence, or wherein each second immobilised primer of the third proportion comprises oxoguanine (e.g. 8-oxoguanine) when the second immobilised primer is an RNA sequence.
Clause 40. A solid support according to any one of clauses 1 to 39, wherein the first cleavage conditions and the second cleavage conditions are the same or are different.
Clause 41. A solid support according to clause 40, wherein the first cleavage conditions and the second cleavage conditions are the same.
Clause 42. A solid support according to any one of clauses 1 to 41, wherein the first proportion of first immobilised primers that are cleavable are further configured to be linearisable under first linearisation conditions.
Clause 43. A solid support according to any one of clauses 1 to 42, wherein the third proportion of second immobilised primers that are cleavable are further configured to be linearisable under second linearisation conditions.
Clause 44. A solid support according to clause 43 as dependent on clause 42, wherein the first linearisation conditions and the second linearisation conditions are the same or are different.
Clause 45. A solid support according to clause 44, wherein the first linearisation conditions and the second linearisation conditions are the same.
Clause 46. A solid support according to any one of clauses 1 to 45, wherein each first immobilised primer comprises a sequence as defined in SEQ ID NO. 1 or 5, or a variant or fragment thereof; and each second immobilised primer comprises a sequence as defined in SEQ ID NO. 2, or a variant or fragment thereof; or wherein each first immobilised primer comprises a sequence as defined in SEQ ID NO. 2, or a variant or fragment thereof; and each second immobilised primer comprises a sequence as defined in SEQ ID NO. 1 or 5, or a variant or fragment thereof.
Clause 47. A solid support, comprising: a plurality of first immobilised primers, and a plurality of second immobilised primers, wherein the plurality of first immobilised primers are located in a first region, wherein the plurality of second immobilised primers are located in a second region surrounding the first region.
Clause 48. A solid support according to clause 47, wherein the second region is contiguous.
Clause 49. A solid support according to clause 47 or clause 48, wherein the second region is an annular region.
Clause 50. A solid support according to any one of clauses 47 to 49, wherein a concentration of first immobilised primers in the first region is greater than a concentration of second immobilised primers in the second region.
Clause 51. A solid support according to clause 50, wherein a ratio between the concentration of first immobilised primers in the first region and the concentration of second immobilised primers in the second region is between 1.25: 1 to 5: 1.
Clause 52. A solid support according to clause 51, wherein the ratio between the concentration of first immobilised primers in the first region and the concentration of second immobilised primers in the second region is between 1.5 : 1 to 3 : 1.
Clause 53. A solid support according to clause 52, wherein the ratio between the concentration of first immobilised primers in the first region and the concentration of second immobilised primers in the second region is about 2: 1.
Clause 54. A solid support according to any one of clauses 47 to 49, wherein a concentration of first immobilised primers in the first region is less than a concentration of second immobilised primers in the second region.
Clause 55. A solid support according to clause 54, wherein a ratio between the concentration of first immobilised primers in the first region and the concentration of second immobilised primers in the second region is between 1: 1.25 to 1:5.
Clause 56. A solid support according to clause 55, wherein the ratio between the concentration of first immobilised primers in the first region and the concentration of second immobilised primers in the second region is between 1: 1.5 to 1:3.
Clause 57. A solid support according to clause 56, wherein the ratio between the concentration of first immobilised primers in the first region and the concentration of second immobilised primers in the second region is about 1:2.
Clause 58. A solid support according to any one of clauses 47 to 49, wherein a concentration of first immobilised primers in the first region is equal or substantially equal to a concentration of second immobilised primers in the second region.
Clause 59. A solid support according to any one of clauses 1 to 58, wherein the solid support is a flow cell.
Clause 60. A kit comprising a solid support according to any one of clauses 1 to 59.
Clause 61. Use of a solid support according to any one of clauses 1 to 59 in nucleic acid sequencing.
Clause 62. A process of manufacturing a solid support, comprising:
(a) immobilising a plurality of first precursor primers onto a solid support to form a plurality of first immobilised primers, wherein a first proportion of the first precursor primers are configured to be cleavable under first cleavage conditions and are unblocked, and wherein a second proportion of the first precursor primers are configured to be non- cleavable under first cleavage conditions and are blocked; and
(b) immobilising a plurality of second precursor primers onto the solid support to form a plurality of second immobilised primers, wherein a third proportion of the second precursor primers are configured to be cleavable under second cleavage conditions and are blocked, and wherein a fourth proportion of the second precursor primers are configured to be non-cleavable under second cleavage conditions and are unblocked.
Clause 63. A process according to clause 62, wherein steps (a) and (b) are conducted sequentially or simultaneously.
Clause 64. A process according to clause 63, wherein step (b) is conducted after step (a).
Clause 65. A process according to clause 63, wherein step (a) is conducted after step (b).
Clause 66. A process according to clause 63, wherein steps (a) and (b) are conducted simultaneously.
Clause 67. A process according to any one of clauses 62 to 66, wherein immobilisation comprises forming covalent linkages between the solid support and each of the plurality of first precursor primers, and between the solid support and each of the plurality of second precursor primers.
Clause 68. A process according to clause 67, wherein forming covalent linkages involves using a click reaction.
Clause 69. A process according to clause 67 or clause 68, wherein forming covalent linkages involves forming a 1,2, 3 -triazole linkage.
Clause 70. A method of preparing polynucleotide sequences for identification, comprising: providing a solid support according to any one of clauses 1 to 46, and
synthesising a plurality of template sequences that extend from the second immobilised primers in the fourth proportion and a plurality of template complement sequences that extend from the first immobilised primers in the first proportion.
Clause 71. A method according to clause 70, wherein the step of synthesising the plurality of template sequences extending from the second immobilised primers in the fourth proportion and the plurality of template complement sequences extending from the first immobilised primers in the first proportion involves conducting amplification.
Clause 72. A method according to clause 71, wherein amplification is bridge amplification.
Clause 73. A method according to clause 71 or clause 72, wherein amplification is conducted over 20 to 40 cycles.
Clause 74. A method according to clause 73, wherein amplification is conducted over 25 to 35 cycles.
Clause 75. A method according to any one of clauses 70 to 74, further comprising a step of removing second immobilised primers in the fourth proportion that are not yet extended, and removing first immobilised primers in the first proportion that are not yet extended.
Clause 76. A method according to clause 75, wherein the step of removing second immobilised primers in the fourth proportion that are not yet extended and removing first immobilised primers in the first proportion that are not yet extended is conducted using a 3 ’-5’ exonuclease (e.g. Exol).
Clause 77. A method according to clause 75 or clause 76, further comprising a step of unblocking the second proportion of first immobilised primers and unblocking the third proportion of second immobilised primers.
Clause 78. A method according to clause 77, further comprising a step of synthesising a plurality of template sequences that extend from the second immobilised primers in the third proportion and a plurality of template complement sequences that extend from the first immobilised primers in the second proportion.
Clause 79. A method according to clause 78, wherein the step of synthesising the plurality of template sequences extending from the second immobilised primers in the third proportion and
the plurality of template complement sequences extending from the first immobilised primers in the second proportion involves conducting amplification.
Clause 80. A method according to clause 79, wherein amplification is bridge amplification.
Clause 81. A method according to clause 79 or clause 80, wherein amplification is conducted over 5 to 25 cycles.
Clause 82. A method according to clause 81, wherein amplification is conducted over 10 to 20 cycles.
Clause 83. A method according to clause 79 or clause 80, wherein the number of amplification cycles used for synthesising the plurality of template sequences extending from the second immobilised primers in the third proportion and the plurality of template complement sequences extending from the first immobilised primers in the second proportion is less than the number of amplification cycles used for synthesising the plurality of template sequences extending from the second immobilised primers in the fourth proportion and the plurality of template complement sequences extending from the first immobilised primers in the first proportion.
Clause 84. A method according to clause 79 or clause 80, wherein the number of amplification cycles used for synthesising the plurality of template sequences extending from the second immobilised primers in the third proportion and the plurality of template complement sequences extending from the first immobilised primers in the second proportion is more than the number of amplification cycles used for synthesising the plurality of template sequences extending from the second immobilised primers in the fourth proportion and the plurality of template complement sequences extending from the first immobilised primers in the first proportion.
Clause 85. A method according to clause 79 or clause 80, wherein the number of amplification cycles used for synthesising the plurality of template sequences extending from the second immobilised primers in the third proportion and the plurality of template complement sequences extending from the first immobilised primers in the second proportion is the same as the number of amplification cycles used for synthesising the plurality of template sequences extending from the second immobilised primers in the fourth proportion and the plurality of template complement sequences extending from the first immobilised primers in the first proportion.
Clause 86. A method according to any one of clauses 78 to 85, further comprising a step of removing second immobilised primers in the third proportion that are not yet extended, and removing first immobilised primers in the second proportion that are not yet extended.
Clause 87. A method according to clause 86, wherein the step of removing second immobilised primers in the third proportion that are not yet extended and removing first immobilised primers in the second proportion that are not yet extended is conducted using a 3 ’-5’ exonuclease (e.g. Exol).
Clause 88. A method according to any one of clauses 78 to 87, further comprising a step of exposing the solid support to first cleavage conditions and/or second cleavage conditions.
Clause 89. A method according to clause 88, wherein the first cleavage conditions and/or second cleavage conditions comprise exposure to a thermal trigger, a light trigger, and/or a chemical/biochemical trigger.
Clause 90. A method according to clause 89, wherein the solid support is exposed to a metal catalyst.
Clause 91. A method according to clause 90, wherein the solid support is exposed to a transition metal catalyst.
Clause 92. A solid support according to clause 91, wherein the solid support is exposed to a palladium-based or a nickel-based catalyst.
Clause 93. A solid support according to clause 92, wherein the solid support is exposed to a palladium-based catalyst.
Clause 94. A method according to clause 89, wherein the solid support is exposed to a glycosylase.
Clause 95. A method according to any one of clauses 78 to 94, wherein the method further comprises a step of linearising the plurality of template sequences extending from the second immobilised primers of the fourth proportion and linearising the plurality of template complement sequences extending from the first immobilised primers of the second proportion.
Clause 96. A method according to clause 95, wherein the method further comprises treating the linearised template sequences and the linearised template complement sequences with a singlestranded binding protein.
Clause 97. A method of sequencing polynucleotide sequences, comprising: preparing polynucleotide sequences for identification using a method according to any one of clauses 70 to 96; and concurrently sequencing nucleobases in the template sequences extending from the second immobilised primers of the fourth proportion and the template complement sequences extending from the first immobilised primers of the second proportion.
Clause 98. A method according to clause 97, wherein the step of concurrently sequencing nucleobases comprises performing sequencing-by-synthesis or sequencing-by-ligation.
Clause 99. A method according to clause 97 or clause 98, wherein the step of concurrently sequencing nucleobases comprises treatment with a strand displacement polymerase (e.g. phi29).
Clause 100. A method according to any one of clauses 97 to 99, wherein the method further comprises a step of conducting paired-end reads.
Clause 101. A kit comprising instructions for preparing polynucleotide sequences for identification according to any one of clauses 70 to 96; and/or sequencing polynucleotide sequences according to any one of clauses 97 to 100.
Clause 102. A data processing device comprising means for carrying out a method according to any one of clauses 70 to 100.
Clause 103. A data processing device according to clause 102, wherein the data processing device is a polynucleotide sequencer.
Clause 104. A computer program product comprising instructions which, when the program is executed by a processor, cause the processor to carry out a method according to any one of clauses 70 to 100.
Clause 105. A computer-readable storage medium comprising instructions which, when executed by a processor, cause the processor to carry out a method according to any one of clauses 70 to 100.
Clause 106. A computer-readable data carrier having stored thereon a computer program product according to clause 104. Clause 107. A data carrier signal carrying a computer program product according to clause 104.
SEQUENCE LISTING
SEQ ID NO. 1 : P5 sequence
AATGATACGGCGACCACCGAGATCTACAC
SEQ ID NO. 2 : P7 sequence
CAAGCAGAAGACGGCATACGAGAT
SEQ ID NO. 3 : P5’ sequence (complementary to P5)
GTGTAGATCTCGGTGGTCGCCGTATCATT
SEQ ID NO. 4 : P7’ sequence (complementary to P7)
ATCTCGTATGCCGTCTTCTGCTTG
SEQ ID NO. 5 : Alternative P5 sequence
AATGATACGGCGACCGA
SEQ ID NO. 6 : Alternative P5 ’ sequence (complementary to alternative P5 sequence)
TCGGTCGCCGTATCATT
SEQ ID NO. 7: SBS3
ACACTCTTTCCCTACACGACGCTCTTCCGATCT
SEQ ID NO. 8: SBS3’
AGATCGGAAGAGCGTCGTGTAGGGAAAGAGTGT
SEQ ID NO. 9: SBS12
GTGACTGGAGTTCAGACGTGTGCTCTTCCGATCT
SEQ ID NO. 10: SBS12’
AGAT C GG AAGAGC AC AC GT CT GAAC T C C AGT C AC
SEQ ID NO. 11: v4 P5-nonlin block
5’alkyne-TTTTTTAATGATACGGCGACCACCGATCTA*C*A*C-phosphate
“*” denotes a thiophosphate linkage.
SEQ ID NO. 12: v4 P7-lin block
5’alkyne-TTTTTTXCAAGCAGAAGACGGCATACGA*G*A*T-phosphate
“*” denotes a thiophosphate linkage.
X” denotes alkene-dT linkage:
wherein represents an attachment point of the nucleotide to the rest of the oligo.
SEQ ID NO. 13: P5-lin non-block
5’alkyne-TTTTTTXAATGATACGGCGACCACCGATCTACAC
‘X” denotes alkene-dT linkage:
wherein represents an attachment point of the nucleotide to the rest of the oligo.
SEQ ID NO. 14: P7-nonlin non-block
5 ’ alkyne -T TTTTTCAAG C AG AAG AC GG CAT AC GAG AT
Claims
1. A solid support, comprising: a plurality of first immobilised primers, and a plurality of second immobilised primers, wherein a first proportion of the first immobilised primers are configured to be cleavable under first cleavage conditions and are unblocked, wherein a second proportion of the first immobilised primers are configured to be non-cleavable under first cleavage conditions and are blocked, wherein a third proportion of the second immobilised primers are configured to be cleavable under second cleavage conditions and are blocked; and wherein a fourth proportion of the second immobilised primers are configured to be non-cleavable under second cleavage conditions and are unblocked.
2. A solid support according to claim 1, wherein a ratio between unblocked immobilised primers and blocked immobilised primers is between 20:80 to 80:20; optionally between 50:50 to 75:25, between 60:40 to 70:30, or about 2: 1.
3. A solid support according to claim 1 or claim 2, wherein the first immobilised primers of the second proportion that are blocked each comprise a blocking group at a 3 ’ end of the first immobilised primer.
4. A solid support according to any one of claims 1 to 3, wherein the second immobilised primers of the third proportion that are blocked each comprise a blocking group at a 3 ’ end of the second immobilised primer.
5. A solid support according to any one of claims 1 to 4, wherein the first immobilised primers of the first proportion are configured to be cleavable by a thermal trigger, a light trigger, and/or a chemical/biochemical trigger.
6. A solid support according to claim 5, wherein the first immobilised primers of the first proportion are configured to be cleavable by a metal catalyst, or a glycosylase.
7. A solid support according to any one of claims 1 to 6, wherein each first immobilised primer of the first proportion comprises a nucleobase which is not selected from guanine, cytosine, adenine or thymine when the first immobilised primer is a DNA sequence; or wherein each first immobilised primer of the first proportion comprises a nucleobase
which is not selected from guanine, cytosine, adenine or uracil when the first immobilised primer is an RNA sequence; optionally wherein each first immobilised primer of the first proportion comprises oxoguanine (e.g. 8-oxoguanine) or uracil when the first immobilised primer is a DNA sequence, or wherein each first immobilised primer of the first proportion comprises oxoguanine (e.g. 8-oxoguanine) when the first immobilised primer is an RNA sequence.
8. A solid support according to any one of claims 1 to 7, wherein the second immobilised primers of the third proportion are configured to be cleavable by a thermal trigger, a light trigger, or a chemical/biochemical trigger.
9. A solid support according to claim 8, wherein the second immobilised primers of the third proportion are configured to be cleavable by a metal catalyst, or a glycosylase.
10. A solid support according to any one of claims 1 to 9, wherein each second immobilised primer of the third proportion comprises a nucleobase which is not selected from guanine, cytosine, adenine or thymine when the second immobilised primer is a DNA sequence; or wherein each second immobilised primer of the third proportion comprises a nucleobase which is not selected from guanine, cytosine, adenine or uracil when the second immobilised primer is an RNA sequence; optionally wherein each second immobilised primer of the third proportion comprises oxoguanine (e.g. 8-oxoguanine) or uracil when the second immobilised primer is a DNA sequence, or wherein each second immobilised primer of the third proportion comprises oxoguanine (e.g. 8-oxoguanine) when the second immobilised primer is an RNA sequence.
11. A solid support according to any one of claims 1 to 10, wherein the first cleavage conditions and the second cleavage conditions are the same or are different; optionally wherein the first cleavage conditions and the second cleavage conditions are the same.
12. A solid support according to any one of claims 1 to 11, wherein the first proportion of first immobilised primers that are cleavable are further configured to be linearisable under first linearisation conditions.
13. A solid support according to any one of claims 1 to 12, wherein the third proportion of second immobilised primers that are cleavable are further configured to be linearisable under second linearisation conditions.
14. A solid support according to claim 13 as dependent on claim 12, wherein the first linearisation conditions and the second linearisation conditions are the same or are different; optionally wherein the first linearisation conditions and the second linearisation conditions are the same.
15. A solid support, comprising: a plurality of first immobilised primers, and a plurality of second immobilised primers, wherein the plurality of first immobilised primers are located in a first region, wherein the plurality of second immobilised primers are located in a second region surrounding the first region.
16. A solid support according to claim 15, wherein the second region is contiguous.
17. A solid support according to claim 15 or claim 16, wherein the second region is an annular region.
18. A solid support according to any one of claims 15 to 17, wherein a concentration of first immobilised primers in the first region is greater than a concentration of second immobilised primers in the second region.
19. A solid support according to claim 18, wherein a ratio between the concentration of first immobilised primers in the first region and the concentration of second immobilised primers in the second region is between 1.25 : 1 to 5 : 1 ; optionally between 1.5 : 1 to 3 : 1 , or about 2: 1.
20. A solid support according to any one of claims 15 to 17, wherein a concentration of first immobilised primers in the first region is less than a concentration of second immobilised primers in the second region.
21. A solid support according to claim 20, wherein a ratio between the concentration of first immobilised primers in the first region and the concentration of second immobilised primers in the second region is between 1: 1.25 to 1:5; optionally between 1: 1.5 to 1:3, or about 1:2.
22. A solid support according to any one of claims 15 to 17, wherein a concentration of first immobilised primers in the first region is equal or substantially equal to a concentration of second immobilised primers in the second region.
23. A solid support according to any one of claims 1 to 22, wherein the solid support is a flow cell.
24. A kit comprising a solid support according to any one of claims 1 to 23.
25. Use of a solid support according to any one of claims 1 to 23 in nucleic acid sequencing.
26. A process of manufacturing a solid support, comprising:
(a) immobilising a plurality of first precursor primers onto a solid support to form a plurality of first immobilised primers, wherein a first proportion of the first precursor primers are configured to be cleavable under first cleavage conditions and are unblocked, and wherein a second proportion of the first precursor primers are configured to be non- cleavable under first cleavage conditions and are blocked; and
(b) immobilising a plurality of second precursor primers onto the solid support to form a plurality of second immobilised primers, wherein a third proportion of the second precursor primers are configured to be cleavable under second cleavage conditions and are blocked, and wherein a fourth proportion of the second precursor primers are configured to be non-cleavable under second cleavage conditions and are unblocked.
27. A process according to claim 26, wherein steps (a) and (b) are conducted sequentially or simultaneously.
28. A process according to claim 27, wherein step (b) is conducted after step (a), wherein step (a) is conducted after step (b), or wherein steps (a) and (b) are conducted simultaneously.
29. A method of preparing polynucleotide sequences for identification, comprising: providing a solid support according to any one of claims 1 to 14, and synthesising a plurality of template sequences that extend from the second immobilised primers in the fourth proportion and a plurality of template complement sequences that extend from the first immobilised primers in the first proportion.
30. A method according to claim 29, wherein the step of synthesising the plurality of template sequences extending from the second immobilised primers in the fourth proportion and the plurality of template complement sequences extending from the first immobilised primers in the first proportion involves conducting amplification.
31. A method according to claim 30, wherein amplification is bridge amplification.
32. A method according to claim 30 or claim 31, wherein amplification is conducted over 20 to 40 cycles; optionally 25 to 35 cycles.
33. A method according to any one of claims 29 to 32, further comprising a step of removing second immobilised primers in the fourth proportion that are not yet extended, and removing first immobilised primers in the first proportion that are not yet extended.
34. A method according to claim 33, wherein the step of removing second immobilised primers in the fourth proportion that are not yet extended and removing first immobilised primers in the first proportion that are not yet extended is conducted using a 3 ’-5’ exonuclease (e.g. Exol).
35. A method according to claim 33 or claim 34, further comprising a step of unblocking the second proportion of first immobilised primers and unblocking the third proportion of second immobilised primers.
36. A method according to claim 35, further comprising a step of synthesising a plurality of template sequences that extend from the second immobilised primers in the third proportion and a plurality of template complement sequences that extend from the first immobilised primers in the second proportion.
37. A method according to claim 36, wherein the step of synthesising the plurality of template sequences extending from the second immobilised primers in the third proportion and the plurality of template complement sequences extending from the first immobilised primers in the second proportion involves conducting amplification.
38. A method according to claim 37, wherein amplification is bridge amplification.
39. A method according to claim 37 or claim 38, wherein amplification is conducted over 5 to 25 cycles; optionally 10 to 20 cycles.
40. A method according to claim 37 or claim 38, wherein:
(i) the number of amplification cycles used for synthesising the plurality of template sequences extending from the second immobilised primers in the third proportion and the plurality of template complement sequences extending from the first immobilised primers in the second proportion is less than the number of amplification cycles used for synthesising the plurality of template sequences extending from the second immobilised primers in the fourth proportion and the plurality of template complement sequences extending from the first immobilised primers in the first proportion; or
(ii) the number of amplification cycles used for synthesising the plurality of template sequences extending from the second immobilised primers in the third proportion and the plurality of template complement sequences extending from the first immobilised primers in the second proportion is more than the number of amplification cycles used for synthesising the plurality of template sequences extending from the second immobilised primers in the fourth proportion and the plurality of template complement sequences extending from the first immobilised primers in the first proportion; or
(iii) the number of amplification cycles used for synthesising the plurality of template sequences extending from the second immobilised primers in the third proportion and the plurality of template complement sequences extending from the first immobilised primers in the second proportion is the same as the number of amplification cycles used for synthesising the plurality of template sequences extending from the second immobilised primers in the fourth proportion and the plurality of template complement sequences extending from the first immobilised primers in the first proportion.
41. A method according to any one of claims 36 to 40, further comprising a step of removing second immobilised primers in the third proportion that are not yet extended, and removing first immobilised primers in the second proportion that are not yet extended; optionally wherein the step of removing second immobilised primers in the third proportion that are not yet extended and removing first immobilised primers in the second proportion that are not yet extended is conducted using a 3 ’-5’ exonuclease (e.g. Exol).
42. A method according to any one of claims 36 to 41, further comprising a step of exposing the solid support to first cleavage conditions and/or second cleavage conditions.
43. A method according to claim 42, wherein the first cleavage conditions and/or second cleavage conditions comprise exposure to a thermal trigger, a light trigger, and/or a chemical/biochemical trigger.
44. A method according to claim 43, wherein the solid support is exposed to a metal catalyst, or a glycosylase.
45. A method according to any one of claims 36 to 44, wherein the method further comprises a step of linearising the plurality of template sequences extending from the second immobilised primers of the fourth proportion and linearising the plurality of template complement sequences extending from the first immobilised primers of the second proportion.
46. A method according to claim 45, wherein the method further comprises treating the linearised template sequences and the linearised template complement sequences with a single-stranded binding protein.
47. A method of sequencing polynucleotide sequences, comprising: preparing polynucleotide sequences for identification using a method according to any one of claims 29 to 46; and concurrently sequencing nucleobases in the template sequences extending from the second immobilised primers of the fourth proportion and the template complement sequences extending from the first immobilised primers of the second proportion.
48. A kit comprising instructions for preparing polynucleotide sequences for identification according to any one of claims 29 to 46; and/or sequencing polynucleotide sequences according to claim 47.
49. A data processing device comprising means for carrying out a method according to any one of claims 29 to 47; optionally wherein the data processing device is a polynucleotide sequencer.
50. A computer program product comprising instructions which, when the program is executed by a processor, cause the processor to carry out a method according to any one of claims 29 to 47.
51. A computer-readable storage medium comprising instructions which, when executed by a processor, cause the processor to carry out a method according to any one of claims 29 to 47.
52. A computer-readable data carrier having stored thereon a computer program product according to claim 50.
53. A data carrier signal carrying a computer program product according to claim 50.
Applications Claiming Priority (2)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| US202363508070P | 2023-06-14 | 2023-06-14 | |
| US63/508,070 | 2023-06-14 |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| WO2024256580A1 true WO2024256580A1 (en) | 2024-12-19 |
Family
ID=91616878
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/EP2024/066446 Pending WO2024256580A1 (en) | 2023-06-14 | 2024-06-13 | Concurrent sequencing with spatially separated rings |
Country Status (1)
| Country | Link |
|---|---|
| WO (1) | WO2024256580A1 (en) |
Citations (31)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO1998044152A1 (en) | 1997-04-01 | 1998-10-08 | Glaxo Group Limited | Method of nucleic acid sequencing |
| WO1998044151A1 (en) | 1997-04-01 | 1998-10-08 | Glaxo Group Limited | Method of nucleic acid amplification |
| WO2000018957A1 (en) | 1998-09-30 | 2000-04-06 | Applied Research Systems Ars Holding N.V. | Methods of nucleic acid amplification and sequencing |
| US6306597B1 (en) | 1995-04-17 | 2001-10-23 | Lynx Therapeutics, Inc. | DNA sequencing by parallel oligonucleotide extensions |
| WO2002006456A1 (en) | 2000-07-13 | 2002-01-24 | Invitrogen Corporation | Methods and compositions for rapid protein and peptide extraction and isolation using a lysis matrix |
| WO2003074734A2 (en) | 2002-03-05 | 2003-09-12 | Solexa Ltd. | Methods for detecting genome-wide sequence variations associated with a phenotype |
| WO2005068656A1 (en) | 2004-01-12 | 2005-07-28 | Solexa Limited | Nucleic acid characterisation |
| US20060024681A1 (en) | 2003-10-31 | 2006-02-02 | Agencourt Bioscience Corporation | Methods for producing a paired tag from a nucleic acid sequence and methods of use thereof |
| WO2006084132A2 (en) | 2005-02-01 | 2006-08-10 | Agencourt Bioscience Corp. | Reagents, methods, and libraries for bead-based squencing |
| WO2006110855A2 (en) | 2005-04-12 | 2006-10-19 | 454 Life Sciences Corporation | Methods for determining sequence variants using ultra-deep sequencing |
| WO2006135342A1 (en) | 2005-06-14 | 2006-12-21 | Agency For Science, Technology And Research | Method of processing and/or genome mapping of ditag sequences |
| US20060292611A1 (en) | 2005-06-06 | 2006-12-28 | Jan Berka | Paired end sequencing |
| WO2007010252A1 (en) | 2005-07-20 | 2007-01-25 | Solexa Limited | Method for sequencing a polynucleotide template |
| WO2007052006A1 (en) | 2005-11-01 | 2007-05-10 | Solexa Limited | Method of preparing libraries of template polynucleotides |
| WO2007091077A1 (en) | 2006-02-08 | 2007-08-16 | Solexa Limited | Method for sequencing a polynucleotide template |
| WO2007107710A1 (en) | 2006-03-17 | 2007-09-27 | Solexa Limited | Isothermal methods for creating clonal single molecule arrays |
| WO2008041002A2 (en) | 2006-10-06 | 2008-04-10 | Illumina Cambridge Limited | Method for sequencing a polynucleotide template |
| WO2008093098A2 (en) | 2007-02-02 | 2008-08-07 | Illumina Cambridge Limited | Methods for indexing samples and sequencing multiple nucleotide templates |
| WO2010048605A1 (en) | 2008-10-24 | 2010-04-29 | Epicentre Technologies Corporation | Transposon end compositions and methods for modifying nucleic acids |
| US20120301925A1 (en) | 2011-05-23 | 2012-11-29 | Alexander S Belyaev | Methods and compositions for dna fragmentation and tagging by transposases |
| US20120316086A1 (en) | 2011-06-09 | 2012-12-13 | Illumina, Inc. | Patterned flow-cells useful for nucleic acid analysis |
| US20130079232A1 (en) | 2011-09-23 | 2013-03-28 | Illumina, Inc. | Methods and compositions for nucleic acid sequencing |
| US20130143774A1 (en) | 2011-12-05 | 2013-06-06 | The Regents Of The University Of California | Methods and compositions for generating polynucleic acid fragments |
| WO2013188582A1 (en) | 2012-06-15 | 2013-12-19 | Illumina, Inc. | Kinetic exclusion amplification of nucleic acid libraries |
| US9347092B2 (en) * | 2009-02-25 | 2016-05-24 | Roche Molecular System, Inc. | Solid support for high-throughput nucleic acid analysis |
| WO2016189331A1 (en) | 2015-05-28 | 2016-12-01 | Illumina Cambridge Limited | Surface-based tagmentation |
| US9670529B2 (en) * | 2012-02-28 | 2017-06-06 | Population Genetics Technologies Ltd. | Method for attaching a counter sequence to a nucleic acid sample |
| US20180037950A1 (en) * | 2014-11-11 | 2018-02-08 | Illumina Cambridge Limited | Methods and arrays for producing and sequencing monoclonal clusters of nucleic acid |
| US20190212294A1 (en) | 2018-01-08 | 2019-07-11 | Illumina, Inc. | High-Throughput Sequencing with Semiconductor-Based Detection |
| WO2019222264A1 (en) | 2018-05-15 | 2019-11-21 | Illumina, Inc. | Compositions and methods for chemical cleavage and deprotection of surface-bound oligonucleotides |
| WO2022087150A2 (en) | 2020-10-21 | 2022-04-28 | Illumina, Inc. | Sequencing templates comprising multiple inserts and compositions and methods for improving sequencing throughput |
-
2024
- 2024-06-13 WO PCT/EP2024/066446 patent/WO2024256580A1/en active Pending
Patent Citations (31)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US6306597B1 (en) | 1995-04-17 | 2001-10-23 | Lynx Therapeutics, Inc. | DNA sequencing by parallel oligonucleotide extensions |
| WO1998044151A1 (en) | 1997-04-01 | 1998-10-08 | Glaxo Group Limited | Method of nucleic acid amplification |
| WO1998044152A1 (en) | 1997-04-01 | 1998-10-08 | Glaxo Group Limited | Method of nucleic acid sequencing |
| WO2000018957A1 (en) | 1998-09-30 | 2000-04-06 | Applied Research Systems Ars Holding N.V. | Methods of nucleic acid amplification and sequencing |
| WO2002006456A1 (en) | 2000-07-13 | 2002-01-24 | Invitrogen Corporation | Methods and compositions for rapid protein and peptide extraction and isolation using a lysis matrix |
| WO2003074734A2 (en) | 2002-03-05 | 2003-09-12 | Solexa Ltd. | Methods for detecting genome-wide sequence variations associated with a phenotype |
| US20060024681A1 (en) | 2003-10-31 | 2006-02-02 | Agencourt Bioscience Corporation | Methods for producing a paired tag from a nucleic acid sequence and methods of use thereof |
| WO2005068656A1 (en) | 2004-01-12 | 2005-07-28 | Solexa Limited | Nucleic acid characterisation |
| WO2006084132A2 (en) | 2005-02-01 | 2006-08-10 | Agencourt Bioscience Corp. | Reagents, methods, and libraries for bead-based squencing |
| WO2006110855A2 (en) | 2005-04-12 | 2006-10-19 | 454 Life Sciences Corporation | Methods for determining sequence variants using ultra-deep sequencing |
| US20060292611A1 (en) | 2005-06-06 | 2006-12-28 | Jan Berka | Paired end sequencing |
| WO2006135342A1 (en) | 2005-06-14 | 2006-12-21 | Agency For Science, Technology And Research | Method of processing and/or genome mapping of ditag sequences |
| WO2007010252A1 (en) | 2005-07-20 | 2007-01-25 | Solexa Limited | Method for sequencing a polynucleotide template |
| WO2007052006A1 (en) | 2005-11-01 | 2007-05-10 | Solexa Limited | Method of preparing libraries of template polynucleotides |
| WO2007091077A1 (en) | 2006-02-08 | 2007-08-16 | Solexa Limited | Method for sequencing a polynucleotide template |
| WO2007107710A1 (en) | 2006-03-17 | 2007-09-27 | Solexa Limited | Isothermal methods for creating clonal single molecule arrays |
| WO2008041002A2 (en) | 2006-10-06 | 2008-04-10 | Illumina Cambridge Limited | Method for sequencing a polynucleotide template |
| WO2008093098A2 (en) | 2007-02-02 | 2008-08-07 | Illumina Cambridge Limited | Methods for indexing samples and sequencing multiple nucleotide templates |
| WO2010048605A1 (en) | 2008-10-24 | 2010-04-29 | Epicentre Technologies Corporation | Transposon end compositions and methods for modifying nucleic acids |
| US9347092B2 (en) * | 2009-02-25 | 2016-05-24 | Roche Molecular System, Inc. | Solid support for high-throughput nucleic acid analysis |
| US20120301925A1 (en) | 2011-05-23 | 2012-11-29 | Alexander S Belyaev | Methods and compositions for dna fragmentation and tagging by transposases |
| US20120316086A1 (en) | 2011-06-09 | 2012-12-13 | Illumina, Inc. | Patterned flow-cells useful for nucleic acid analysis |
| US20130079232A1 (en) | 2011-09-23 | 2013-03-28 | Illumina, Inc. | Methods and compositions for nucleic acid sequencing |
| US20130143774A1 (en) | 2011-12-05 | 2013-06-06 | The Regents Of The University Of California | Methods and compositions for generating polynucleic acid fragments |
| US9670529B2 (en) * | 2012-02-28 | 2017-06-06 | Population Genetics Technologies Ltd. | Method for attaching a counter sequence to a nucleic acid sample |
| WO2013188582A1 (en) | 2012-06-15 | 2013-12-19 | Illumina, Inc. | Kinetic exclusion amplification of nucleic acid libraries |
| US20180037950A1 (en) * | 2014-11-11 | 2018-02-08 | Illumina Cambridge Limited | Methods and arrays for producing and sequencing monoclonal clusters of nucleic acid |
| WO2016189331A1 (en) | 2015-05-28 | 2016-12-01 | Illumina Cambridge Limited | Surface-based tagmentation |
| US20190212294A1 (en) | 2018-01-08 | 2019-07-11 | Illumina, Inc. | High-Throughput Sequencing with Semiconductor-Based Detection |
| WO2019222264A1 (en) | 2018-05-15 | 2019-11-21 | Illumina, Inc. | Compositions and methods for chemical cleavage and deprotection of surface-bound oligonucleotides |
| WO2022087150A2 (en) | 2020-10-21 | 2022-04-28 | Illumina, Inc. | Sequencing templates comprising multiple inserts and compositions and methods for improving sequencing throughput |
Non-Patent Citations (2)
| Title |
|---|
| HIGUCHI ET AL., NUCLEIC ACIDS RES., vol. 16, 1988, pages 7351 - 7367 |
| SAMBROOK ET AL.: "Molecular Cloning, A Laboratory Manual", 2001, COLD SPRING HARBOR LABORATORY PRESS |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| US20230098456A1 (en) | Methods for sequencing a polynucleotide template | |
| CN109415761B (en) | Hybrid chain reaction method for in situ molecular detection | |
| CA2810931C (en) | Direct capture, amplification and sequencing of target dna using immobilized primers | |
| US20100041561A1 (en) | Preparation of Nucleic Acid Templates for Solid Phase Amplification | |
| US20090226975A1 (en) | Constant cluster seeding | |
| CA3183217A1 (en) | Compositions and methods for in situ single cell analysis using enzymatic nucleic acid extension | |
| WO2022247555A1 (en) | Sequencing method | |
| AU2023233839A1 (en) | Concurrent sequencing of forward and reverse complement strands on concatenated polynucleotides for methylation detection | |
| CN102639714A (en) | Detection method of target nucleic acid | |
| CN115521977A (en) | Spatial sequencing Using MICTAG | |
| KR20240069835A (en) | Improved method and kit for the generation of dna libraries for massively parallel sequencing | |
| EP4499873A1 (en) | Paired-end re-synthesis using blocked p5 primers | |
| US20250043275A1 (en) | Methods of preparing loop fork libraries | |
| DK2456892T3 (en) | Procedure for sequencing of a polynukleotidskabelon | |
| CN114207229A (en) | Flexible and high-throughput sequencing of target genomic regions | |
| WO2024256581A1 (en) | Determination of modified cytosines | |
| WO2024256580A1 (en) | Concurrent sequencing with spatially separated rings | |
| WO2025062001A1 (en) | Optimised nucleic acid sequencing | |
| WO2025062002A1 (en) | Concurrent sequencing using nick translation | |
| EP4590854A1 (en) | Deformable polymers comprising immobilised primers |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| 121 | Ep: the epo has been informed by wipo that ep was designated in this application |
Ref document number: 24734826 Country of ref document: EP Kind code of ref document: A1 |
|
| WWE | Wipo information: entry into national phase |
Ref document number: 2024734826 Country of ref document: EP |