WO2024246083A1 - Bispecific antibodies targeting bcma and cd28 - Google Patents
Bispecific antibodies targeting bcma and cd28 Download PDFInfo
- Publication number
- WO2024246083A1 WO2024246083A1 PCT/EP2024/064688 EP2024064688W WO2024246083A1 WO 2024246083 A1 WO2024246083 A1 WO 2024246083A1 EP 2024064688 W EP2024064688 W EP 2024064688W WO 2024246083 A1 WO2024246083 A1 WO 2024246083A1
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- seq
- antibody
- amino acid
- bcma
- acid sequence
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Pending
Links
Classifications
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K16/00—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies
- C07K16/18—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans
- C07K16/28—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans against receptors, cell surface antigens or cell surface determinants
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K16/00—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies
- C07K16/18—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans
- C07K16/28—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans against receptors, cell surface antigens or cell surface determinants
- C07K16/2803—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans against receptors, cell surface antigens or cell surface determinants against the immunoglobulin superfamily
- C07K16/2809—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans against receptors, cell surface antigens or cell surface determinants against the immunoglobulin superfamily against the T-cell receptor (TcR)-CD3 complex
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K16/00—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies
- C07K16/18—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans
- C07K16/28—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans against receptors, cell surface antigens or cell surface determinants
- C07K16/2803—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans against receptors, cell surface antigens or cell surface determinants against the immunoglobulin superfamily
- C07K16/2818—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans against receptors, cell surface antigens or cell surface determinants against the immunoglobulin superfamily against CD28 or CD152
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K16/00—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies
- C07K16/18—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans
- C07K16/28—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans against receptors, cell surface antigens or cell surface determinants
- C07K16/2878—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans against receptors, cell surface antigens or cell surface determinants against the NGF-receptor/TNF-receptor superfamily, e.g. CD27, CD30, CD40, CD95
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K16/00—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies
- C07K16/18—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans
- C07K16/28—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans against receptors, cell surface antigens or cell surface determinants
- C07K16/30—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans against receptors, cell surface antigens or cell surface determinants from tumour cells
- C07K16/3061—Blood cells
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K39/00—Medicinal preparations containing antigens or antibodies
- A61K2039/505—Medicinal preparations containing antigens or antibodies comprising antibodies
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K39/00—Medicinal preparations containing antigens or antibodies
- A61K2039/505—Medicinal preparations containing antigens or antibodies comprising antibodies
- A61K2039/507—Comprising a combination of two or more separate antibodies
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K39/00—Medicinal preparations containing antigens or antibodies
- A61K2039/545—Medicinal preparations containing antigens or antibodies characterised by the dose, timing or administration schedule
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/20—Immunoglobulins specific features characterized by taxonomic origin
- C07K2317/24—Immunoglobulins specific features characterized by taxonomic origin containing regions, domains or residues from different species, e.g. chimeric, humanized or veneered
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/30—Immunoglobulins specific features characterized by aspects of specificity or valency
- C07K2317/31—Immunoglobulins specific features characterized by aspects of specificity or valency multispecific
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/30—Immunoglobulins specific features characterized by aspects of specificity or valency
- C07K2317/33—Crossreactivity, e.g. for species or epitope, or lack of said crossreactivity
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/30—Immunoglobulins specific features characterized by aspects of specificity or valency
- C07K2317/35—Valency
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/50—Immunoglobulins specific features characterized by immunoglobulin fragments
- C07K2317/51—Complete heavy chain or Fd fragment, i.e. VH + CH1
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/50—Immunoglobulins specific features characterized by immunoglobulin fragments
- C07K2317/515—Complete light chain, i.e. VL + CL
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/50—Immunoglobulins specific features characterized by immunoglobulin fragments
- C07K2317/52—Constant or Fc region; Isotype
- C07K2317/524—CH2 domain
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/50—Immunoglobulins specific features characterized by immunoglobulin fragments
- C07K2317/52—Constant or Fc region; Isotype
- C07K2317/526—CH3 domain
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/50—Immunoglobulins specific features characterized by immunoglobulin fragments
- C07K2317/56—Immunoglobulins specific features characterized by immunoglobulin fragments variable (Fv) region, i.e. VH and/or VL
- C07K2317/565—Complementarity determining region [CDR]
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/50—Immunoglobulins specific features characterized by immunoglobulin fragments
- C07K2317/56—Immunoglobulins specific features characterized by immunoglobulin fragments variable (Fv) region, i.e. VH and/or VL
- C07K2317/567—Framework region [FR]
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/60—Immunoglobulins specific features characterized by non-natural combinations of immunoglobulin fragments
- C07K2317/66—Immunoglobulins specific features characterized by non-natural combinations of immunoglobulin fragments comprising a swap of domains, e.g. CH3-CH2, VH-CL or VL-CH1
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/70—Immunoglobulins specific features characterized by effect upon binding to a cell or to an antigen
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/70—Immunoglobulins specific features characterized by effect upon binding to a cell or to an antigen
- C07K2317/71—Decreased effector function due to an Fc-modification
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/70—Immunoglobulins specific features characterized by effect upon binding to a cell or to an antigen
- C07K2317/73—Inducing cell death, e.g. apoptosis, necrosis or inhibition of cell proliferation
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/70—Immunoglobulins specific features characterized by effect upon binding to a cell or to an antigen
- C07K2317/75—Agonist effect on antigen
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/90—Immunoglobulins specific features characterized by (pharmaco)kinetic aspects or by stability of the immunoglobulin
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/90—Immunoglobulins specific features characterized by (pharmaco)kinetic aspects or by stability of the immunoglobulin
- C07K2317/92—Affinity (KD), association rate (Ka), dissociation rate (Kd) or EC50 value
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/90—Immunoglobulins specific features characterized by (pharmaco)kinetic aspects or by stability of the immunoglobulin
- C07K2317/94—Stability, e.g. half-life, pH, temperature or enzyme-resistance
Definitions
- the present invention relates to new humanized BCMA antibodies and bispecific antibodies that specifically bind to BCMA and CD28, methods for their production, pharmaceutical compositions containing these antibodies, and methods of using the same.
- Cancer immunotherapy is becoming an increasingly effective therapy option that can result in dramatic and durable responses in cancer types such as melanoma, non-small cell lung cancer and renal cell carcinoma. This is mostly driven by the success of several immune checkpoint inhibitors including anti-PD-1 (e.g. Keytruda, Merck; Opdivo, BMS), anti-CTLA-4 (e.g. Yervoy, BMS) and anti-PD-Ll (e.g. Tecentriq, Roche).
- anti-PD-1 e.g. Keytruda, Merck; Opdivo, BMS
- anti-CTLA-4 e.g. Yervoy, BMS
- anti-PD-Ll e.g. Tecentriq, Roche.
- multiple myeloma is one of the most common hematological malignancies with remaining high unmet medical need.
- Multiple myeloma also known as plasma cell myeloma, is characterized by terminally differentiated plasma cells that secrete non-functional monoclonal immunoglobulins. As the cancerous plasma cells accumulate in the bone marrow, they interfere with the production of normal blood cells, leading to various symptoms and complications. Common symptoms of multiple myeloma include bone pain, especially in the back or ribs, fatigue, weakness, frequent infections, weight loss, excessive thirst, and increased urination.
- Treatment options for multiple myeloma depend on various factors, including the stage of the disease or the patient's overall health.
- the immunomodulatory drugs such as lenalidomide and pomalidomide (ImiDs), and proteasome inhibitors such as carfilzomib or bortezomib may remain the backbone of 1st line therapy for multiple myeloma (Moreau et al, The Lancet Oncology 2021, 22(3), el05- el 18).
- these drugs do not target specifically the diseased tumor cells e.g. diseased plasma cells (PC). Efforts have been made towards selectively depleting the plasma cells in multiple myeloma.
- PC diseased plasma cells
- BCMA B cell maturation antigen
- TNFRSF17 tumor necrosis factor receptor superfamily 17
- CAR immunomodulatory drug
- CD28 is the founding member of a subfamily of costimulatory molecules characterized by paired V-set immunoglobulin superfamily (IgSF) domains attached to single transmembrane domains and cytoplasmic domains that contain critical signaling motifs (Carreno and Collins, Annu Rev Immunol. 2002, 20, 29-53). Other members of the subfamily include ICOS, CTLA-4, PD1, PD1H, TIGIT, and BTLA (Chen and Flies, Nat Rev Immunol. 2013, 13(4), 227-42). CD28 expression is restricted to T cells and prevalent on all naive and a majority of antigen- experienced subsets, including those that express PD-1 or CTLA-4.
- IgSF immunoglobulin superfamily
- CD28 and CTLA-4 are highly homologous and compete for binding to the same B7 molecules CD80 and CD86, which are expressed on dendritic cells, B cells, macrophages, and tumor cells (Linsley et al., Proc Natl Acad Sci USA. 1990, 87(13), 5031-5).
- the higher affinity of CTLA-4 for the B7 family of ligands allows CTLA-4 to outcompete CD28 for ligand binding and suppress effector T cells responses (Engelhardt et al., J Immunol 2006, 177, 1052-1061).
- PD-1 was shown to inhibit CD28 signaling by in part dephosphorylating the cytoplasmic domain of CD28 (Hui et al., Science 2017, 355, 1428-1433).
- CD28 ligands also promotes the expression of inducible costimulatory receptors such as OX-40, ICOS, and 4- IBB (Acuto and Michel, Nat Rev Immunol 2003, 3, 939-951).
- CD28 Upon ligation of CD28, a disulfide-linked homodimer, the membrane proximal YMNM motif and the distal PYAP motif have been shown to complex with several kinases and adaptor proteins (Boomer and Green, Cold Spring Harb Perspect Biol 2010, 2, a002436). These motifs are important for the induction of IL2 transcription, which is mediated by the CD28-dependent activation of NF AT, AP-1, and NFKB family transcription factors (Fraser et al., Science 1991, 251, 313-316). However, additional poorly characterized sites for phosphorylation and ubiquitination are found within the cytoplasmic domain of CD28.
- CD28-initiated pathways have critical roles in promoting the proliferation and effector function of conventional T cells.
- CD28 ligation also promotes the anti-inflammatory function of regulatory T cells.
- CD28 co-stimulates T cells by in part augmenting signals from the T cell receptor, but was also shown to mediate unique signaling events (Acuto and Michel, 2003; Boomer and Green, 2010).
- Signals specifically triggered by CD28 control many important aspects of T cell function, including phosphorylation and other post-translational modifications of downstream proteins (e.g., PI3K mediated phosphorylation), transcriptional changes (eg. Bcl-xL expression), epigenetic changes (e.g.
- CD28 -deficient mice have reduced responses to infectious pathogens, allograft antigens, graft-versus-host disease, contact hypersensitivity and asthma (Acuto and Michel, 2003). Lack of CD28-mediated co-stimulation results in reduced T cell proliferation in vitro and in vivo, in severe inhibition of germinal -centre formation and immunoglobulin isotype-class switching, reduced T helper (Th)-cell differentiation and the expression of Th2-type cytokines. CD4-dependent cytotoxic CD8+ T-cell responses are also affected.
- CD28-deficient naive T cells showed a reduced proliferative response particularly at lower antigen concentrations.
- a growing body of literature supports the idea that engaging CD28 on T cells has anti-tumor potential.
- Recent evidence demonstrates that the anti-cancer effects of PD-L1/PD-1 and CTLA-4 checkpoint inhibitors depend on CD28 (Kamphorst et al., Science 2017, 355, 1423-1427).
- Clinical studies investigating the therapeutic effects of CTLA-4 and PD-1 blockade have shown exceptionally promising results in patients with advanced melanoma and other cancers.
- infusion of genetically engineered T cells expressing artificial chimeric T cell receptors comprising an extracellular antigen recognition domain fused to the intracellular TCR signaling domains (CD3z) and intracellular co-stimulatory domains (CD28 and/or 4- IBB domains) has shown high rates and durability of response in B cell cancers and other cancers.
- CD28 agonistic antibodies can be divided into two categories: (i) CD28 superagonistic antibodies and (ii) CD28 conventional agonistic antibodies. Normally, for the activation of naive T cells both engagement of the T cell antigen receptor (TCR, signal 1) and costimulatory signaling by CD28 (signal 2) is required.
- CD28 Superagonists CD28SA are CD28-specific monoclonal antibodies, which are able to autonomously activate T cells without overt T cell receptor engagement (Hiinig, Nat Rev Immunol 2012, 12, 317-318). In rodents, CD28SA activates conventional and regulatory T cells. CD28SA antibodies are therapeutically effective in multiple models of autoimmunity, inflammation and transplantation.
- Theralizumab (TGN1412 or TAB08) has been re-evaluated in an open-label, multicenter dose escalation study in RA patients and patients with metastatic or unresectable advanced solid malignancies.
- CD28 conventional agonistic antibodies such as clone 9.3, mimic CD28 natural ligands and are only able to enhance T cell activation in presence of a T cell receptor signal (signal 1).
- the binding epitope of the antibody has a major impact on whether the agonistic antibody is a superagonist or a conventional agonist (Beyersdorf et al., Ann. Rheum. Dis. 2005, 64, iv91-iv95).
- the superagonistic TGN1412 binds to a lateral motif of CD28, while the conventional agonistic molecule 9.3 binds close to the ligand binding epitope.
- superagonistic and conventional agonistic antibodies differ in their ability to form linear complexes of CD28 molecules on the surface of T cells.
- TGN1412 is able to efficiently form linear arrays of CD28, which presumably leads to aggregated signaling components which are sufficient to surpass the threshold for T cell activation.
- the conventional agonist 9.3 leads to complexes which are not linear in structure.
- An attempt to convert conventional agonistic binders based on the 9.3 clone has been previously published (Otz et al., Leukemia 2009, 23(1), 71-77) using a recombinant bi-specific single-chain antibody directed to a melanoma-associated proteoglycan and CD28.
- the reported bispecific single chain antibody was reported to exert “supra-agonistic” activity despite the use of a conventional CD28 agonistic binder 9.3, based in the intrinsic tendency of bispecific single chain antibodies to form multimeric constructs.
- T cell bispecific antibodies i.e. T cell bispecific antibodies (TCBs)
- T cell bispecific antibodies i.e. T cell bispecific antibodies
- CD28 is expressed at baseline on T cells in various tumor indications and activation of CD28 signaling enhances T cell receptor signals
- the combination of a TCB molecule with a tumor-targeted CD28 molecule can act synergistically to induce strong and long-lasting anti-tumor responses.
- WO 2020/127618 Al describes tumor-targeted agonistic CD28 antigen binding molecules.
- Various tumor targets are described therein.
- CD28 agonism in Multiple Myeloma may exert different biological functions on immune, respective MM plasma cells. While co-activation of T-cells via CD28 is expected to drive antitumor responses, CD28 agonism on MM cells mediates pro-survival signaling via regulation of PI3K/Akt, FoxO3a, and Bimm which in turn is described to induce chemotherapeutic resistance in multiple myeloma (Murray et al, Blood 2014, 123(24), 3770-3779). Over-expression of CD28 on newly diagnosed Multiple Myeloma plasma cells is described to correlate with worse clinical outcome. However, CD28 activation inhibits myeloma cell proliferation (Bahlis et al., Blood 2007, 109(11), 5002-5010).
- Agonizing CD28 in presence of a strong immune cell mediated response, such as a T-cell bispecific activation of T-cells, can further boost efficient anti-tumor responses.
- a strong immune cell mediated response such as a T-cell bispecific activation of T-cells
- We herein provide bispecific agonistic CD28 antigen binding molecules that specifically bind to BCMA. Enhancing a T cell response with a CD28 bispecific antibody targeting BCMA on myeloma cells may be an option to improve the treatment of Multiple Myeloma and there is a need to provide BCMA-targeted anti-CD28 antibodies with advantageous properties.
- the present invention describes new BCMA-targeted bispecific agonistic CD28 antigen binding molecules which achieve a tumor-dependent T cell activation and tumor cell killing without the necessity to form multimers.
- the bispecific CD28 antigen binding molecules of the present invention are characterized by monovalent binding to CD28 and in that they comprise a certain antigen binding domain as defined herein capable of specific binding to BCMA. Furthermore, they possess an Fc domain composed of a first and a second subunit capable of stable association comprising one or more amino acid substitution that reduces the binding affinity of the antigen binding molecule to an Fc receptor and/or effector function. Fc receptor- mediated cross-linking is thereby abrogated and tumor-specific activation is achieved by crosslinking through binding of the second antigen binding domain capable of specific binding to BCMA.
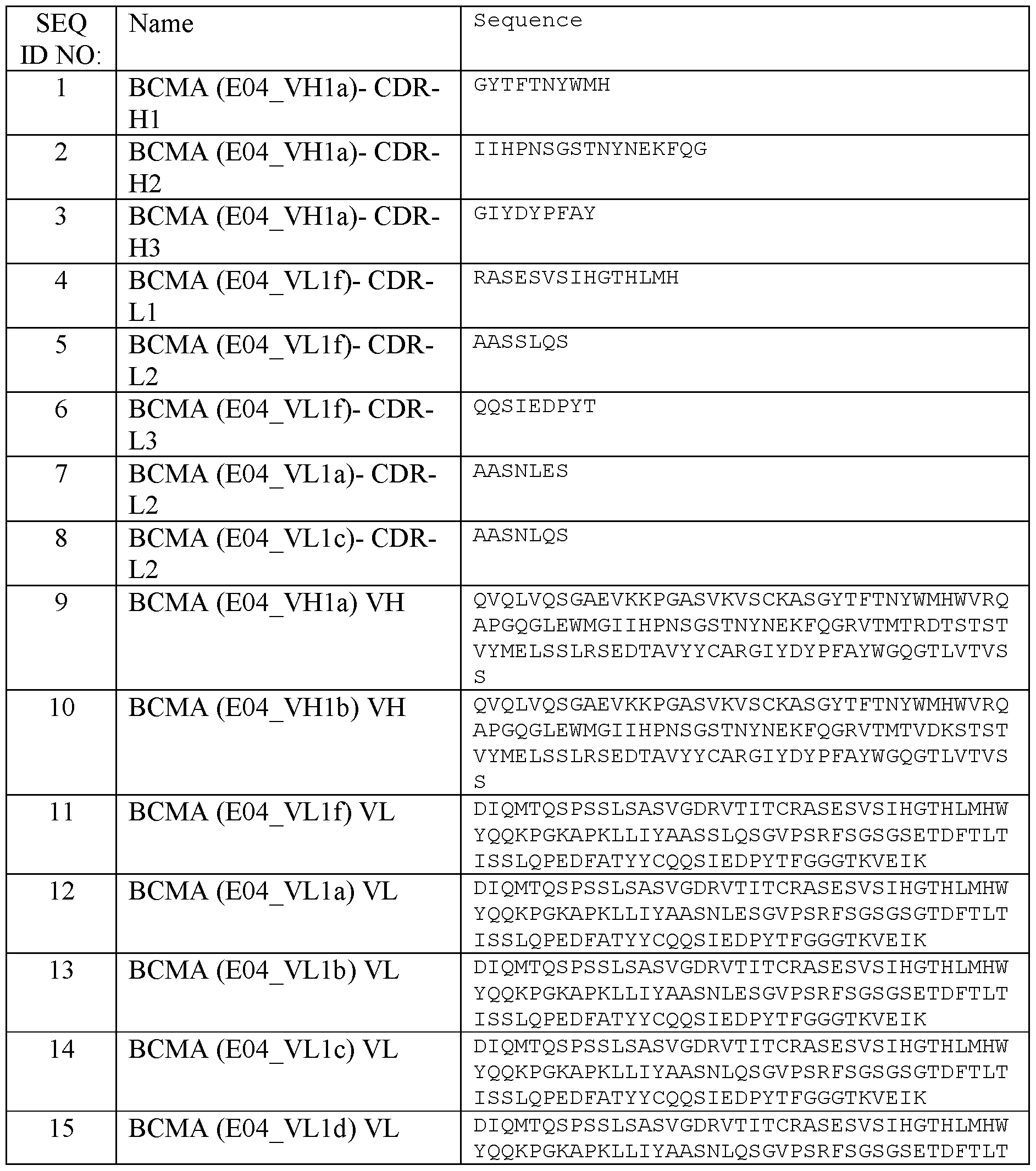
- an antibody that specifically binds to B cell maturation agent (BCMA), wherein the antibody comprises a first antigen binding domain comprising
- VH BCMA heavy chain variable region
- VH BCMA heavy chain variable region comprising a heavy chain complementary determining region CDR-H1 of SEQ ID NO: 1 (GYTFTNYWMH), a CDR-H2 of SEQ ID NO: 2 (IH4PNSGSTNYNEKFQG), and a CDR-H3 of SEQ ID NO: 3 (GIYDYPFAY), and
- VLBCMA light chain variable region
- VL comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 4 (RASES VSIHGTHLMH), a CDR-L2 of SEQ ID NO: 5 (AASSLQS) and a CDR-L3 of SEQ ID NO: 6 (QQSIEDPYT); or
- VL comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 4 (RASES VSIHGTHLMH), a CDR-L2 of SEQ ID NO: 7 (AASNLES) and a CDR-L3 of SEQ ID NO: 6 (QQSIEDPYT), or
- VL comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 4 (RASES VSIHGTHLMH), a CDR-L2 of SEQ ID NO: 8 (AASNLQS) and a CDR-L3 of SEQ ID NO: 6 (QQSIEDPYT).
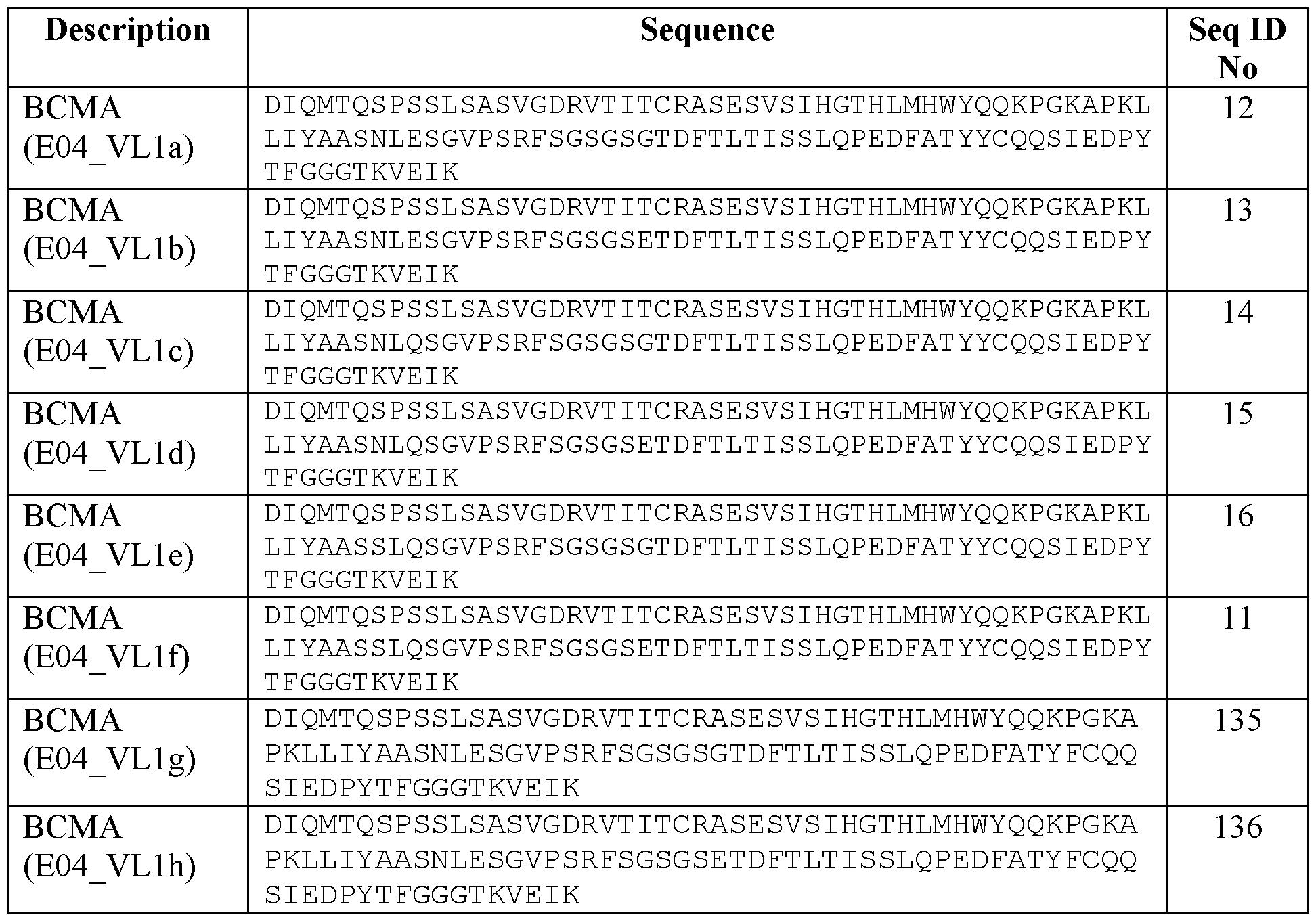
- an antibody that specifically binds to BCMA comprising an amino acid sequence selected from the group consisting of SEQ ID NO: 9 (VHla) and SEQ ID NO: 10 (VHlb), and/or the VL BCMA comprises an amino acid sequence selected from the group consisting of SEQ ID NO: 11 (VLlf) , SEQ ID NO: 12 (VLla), SEQ ID NO: 13 (VLlb), SEQ ID NO: 14 (VLlc), SEQ ID NO: 15 (VLld), and SEQ ID NO: 16 (VLle).
- antibody that specifically binds to BCMA, wherein the antibody comprises a first antigen binding domain comprising
- VH BCMA comprising an amino acid sequence of SEQ ID NO:9
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 11, or
- VH BCMA comprising an amino acid sequence of SEQ ID NOV
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 12.
- the first antigen binding domain is a Fab molecule.
- the antibody that specifically binds to BCMA comprises an Fc domain composed of a first and a second subunit.
- the antibody that specifically binds to BCMA comprises a second antigen binding domain that specifically binds to a second antigen, i.e. is a bispecific antibody.
- the second antigen binding domain that specifically binds to a second antigen is a Fab molecule wherein the variable domains VL and VH or the constant domains CL and CHI, particularly the variable domains VL and VH, of the Fab light chain and the Fab heavy chain are replaced by each other.
- an antibody that specifically binds to BCMA comprising a first antigen binding domain that specifically binds to BCMA, wherein the first antigen binding domain is a Fab molecule wherein in the constant domain CL the amino acid at position 123 (numbering according to Kabat EU index) is substituted by an amino acid selected from lysine (K), arginine (R) or histidine (H) and the amino acid at position 124 (numbering according to Kabat EU index) is substituted independently by lysine (K), arginine (R) or histidine (H), and wherein in the constant domain CHI the amino acid at position 147 (numbering according to Kabat EU index) is substituted independently by glutamic acid (E) or aspartic acid (D) and the amino acid at position 213 (numbering according to Kabat EU index) is substituted independently by glutamic acid (E), or aspartic acid (D) (numbering according to Kabat EU index).
- the antibody that specifically binds to BCMA comprises an Fc domain, wherein the Fc domain is an IgG, particularly an IgGl Fc domain.
- the Fc domain is a human Fc domain.
- the Fc domain comprises a modification promoting the association of the first and the second subunit of the Fc domain. In one aspect, the Fc domain comprises knobs into hole modifications. In one aspect, the first subunit of the Fc domain comprises the amino acid substitutions S354C and T366W (EU numbering) and the second subunit of the Fc domain comprises the amino acid substitutions Y349C, T366S and Y407V (numbering according to Kabat EU index).
- the Fc domain comprises one or more amino acid substitution that reduces binding to an Fc receptor and/or effector function.
- the the Fc domain is of human IgGl subclass and comprises the amino acid mutations L234A, L235A and P329G (numbering according to Kabat EU index).
- antibodies that specifically bind to BCMA and that comprise a second antigen binding domain that specifically binds to CD28.
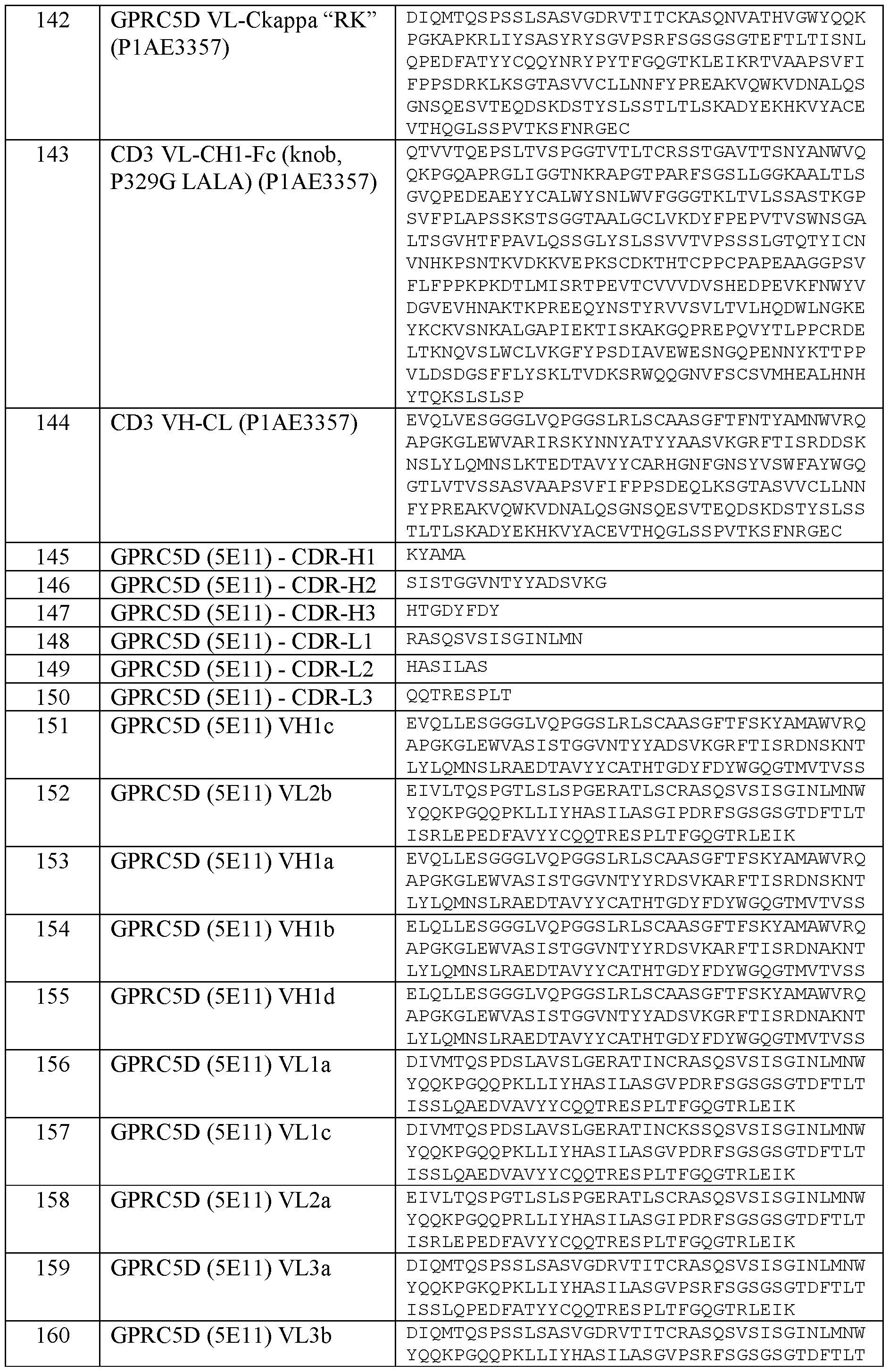
- the second antigen binding domain that specifically binds to CD28 comprises a heavy chain variable region (VHCD28) comprising a heavy chain complementary determining region CDR-H1 of SEQ ID NO: 17, a CDR-H2 of SEQ ID NO: 18, and a CDR-H3 of SEQ ID NO: 19, and a light chain variable region (VLCD28) comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 20, a CDR-L2 of SEQ ID NO: 21 and a CDR-L3 of SEQ ID NO: 22.
- VHCD28 heavy chain variable region comprising a heavy chain complementary determining region CDR-H1 of SEQ ID NO: 17, a CDR-H2 of SEQ ID NO: 18, and a CDR-H3 of SEQ ID NO: 19, and a light chain variable region (VLCD28) comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 20, a CDR-L2 of SEQ ID NO: 21 and
- the second antigen binding domain that specifically binds to CD28 comprises a heavy chain variable region (VHCD28) comprising an amino acid sequence of SEQ ID NO:23, and a light chain variable region (VLCD28) comprising an amino acid sequence of SEQ ID NO:24 (v8).
- VHCD28 heavy chain variable region
- VLCD28 light chain variable region
- the antibody as described herein comprises a first antigen binding domain comprising a VH BCMA comprising an amino acid sequence of SEQ ID NO:9 and VL BCMA comprising an amino acid sequence of SEQ ID NO: 11 and a second antigen binding domain comprising VH CD28 comprising an amino acid sequence of SEQ ID NO:23 and VL CD28 comprising an amino acid sequence of SEQ ID NO:24.
- the antibody comprises a first light chain comprising the amino acid sequence of SEQ ID NO:25, a first heavy chain comprising the amino acid sequence of SEQ ID NO:26, a second heavy chain comprising the amino acid sequence of SEQ ID NO:27 and a second light chain comprising the amino acid sequence of SEQ ID NO:28.
- VH BCMA heavy chain variable region
- VH comprising a heavy chain complementary determining region CDR-H1 of SEQ ID NO: 29 (GFTFSNAWMD), a CDR-H2 of SEQ ID NO: 30 (QITAKSNNYATYYADSVKG), and a CDR-H3 of SEQ ID NO: 31 (DGYH), and
- VH comprising a heavy chain complementary determining region CDR-H1 of SEQ ID NO: 29 (GFTFSNAWMD), a CDR-H2 of SEQ ID NO: 32 (QITAKSNNYATYYAAPVKG), and a CDR-H3 of SEQ ID NO: 31 (DGYH), and
- VLBCMA light chain variable region
- the antibody comprises (A) a first antigen binding domain that specifically binds to BCMA, wherein the VH BCMA comprises an amino acid sequence selected from the group consisting of SEQ ID NO: 36 (VH2a) and SEQ ID NO: 38 (VHlb), and/or the V L BCMA comprises an amino acid sequence selected from the group consisting of SEQ ID NO: 37 (VL2a) and SEQ ID NO:39 (VLla).
- an antibody that specifically binds to BCMA and CD28, wherein the antibody comprises a first antigen binding domain comprising
- VH BCMA comprising an amino acid sequence of SEQ ID NO:36 and a VLBCMA comprising an amino acid sequence of SEQ ID NO:37, or
- VH BCMA comprising an amino acid sequence of SEQ ID NO:38 and a VLBCMA comprising an amino acid sequence of SEQ ID NO:39.
- an antibody that specifically binds to BCMA and CD28, wherein the first antigen binding domain that binds to BCMA is a Fab molecule.
- the antibody that specifically binds to BCMA and CD28 comprises an Fc domain composed of a first and a second subunit.
- the second antigen binding domain that specifically binds to CD28 is a Fab molecule.
- the second antigen binding domain that specifically binds to CD28 is a Fab molecule wherein the variable domains VL and VH or the constant domains CL and CHI, particularly the variable domains VL and VH, of the Fab light chain and the Fab heavy chain are replaced by each other.
- an antibody that specifically binds to BCMA and CD28, wherein the first antigen binding domain that binds to BCMA is a Fab molecule wherein in the constant domain CL the amino acid at position 123 (numbering according to Kabat EU index) is substituted by an amino acid selected from lysine (K), arginine (R) or histidine (H) and the amino acid at position 124 (numbering according to Kabat EU index) is substituted independently by lysine (K), arginine (R) or histidine (H), and wherein in the constant domain CHI the amino acid at position 147 (numbering according to Kabat EU index) is substituted independently by glutamic acid (E) or aspartic acid (D) and the amino acid at position 213 (numbering according to Kabat EU index) is substituted independently by glutamic acid (E), or aspartic acid (D) (numbering according to Kabat EU index).
- the antibody that specifically binds to BCMA and CD28 comprises an Fc domain, wherein the Fc domain is an IgG, particularly an IgGl Fc domain.
- the Fc domain is a human Fc domain.
- the Fc domain comprises a modification promoting the association of the first and the second subunit of the Fc domain. In one aspect, the Fc domain comprises knobs into hole modifications. In one aspect, the first subunit of the Fc domain comprises the amino acid substitutions S354C and T366W (EU numbering) and the second subunit of the Fc domain comprises the amino acid substitutions Y349C, T366S and Y407V (numbering according to Kabat EU index).
- the Fc domain comprises one or more amino acid substitution that reduces binding to an Fc receptor and/or effector function.
- the the Fc domain is of human IgGl subclass and comprises the amino acid mutations L234A, L235A and P329G (numbering according to Kabat EU index).
- the second antigen binding domain that specifically binds to CD28 comprises a heavy chain variable region (VHCD28) comprising a heavy chain complementary determining region CDR-H1 of SEQ ID NO: 17, a CDR-H2 of SEQ ID NO: 18, and a CDR-H3 of SEQ ID NO: 19, and a light chain variable region (VLCD28) comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 20, a CDR-L2 of SEQ ID NO: 21 and a CDR-L3 of SEQ ID NO: 22.
- VHCD28 heavy chain variable region comprising a heavy chain complementary determining region CDR-H1 of SEQ ID NO: 17, a CDR-H2 of SEQ ID NO: 18, and a CDR-H3 of SEQ ID NO: 19, and a light chain variable region (VLCD28) comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 20, a CDR-L2 of SEQ ID NO: 21 and
- the second antigen binding domain that specifically binds to CD28 comprises a heavy chain variable region (VHCD28) comprising an amino acid sequence of SEQ ID NO:23, and a light chain variable region (VLCD28) comprising an amino acid sequence of SEQ ID NO:24.
- VHCD28 heavy chain variable region
- VLCD28 light chain variable region
- an antibody that specifically binds to BCMA and CD28, wherein the antibody comprises a first antigen binding domain comprising a VH BCMA comprising an amino acid sequence of SEQ ID NO:36 and VLBCMA comprising an amino acid sequence of SEQ ID NO:37 and a second antigen binding domain comprising VH CD28 comprising an amino acid sequence of SEQ ID NO:23 and VL CD28 comprising an amino acid sequence of SEQ ID NO:24.
- the antibody that specifically binds to BCMA and CD28 comprises a first light chain comprising the amino acid sequence of SEQ ID NO:40, a first heavy chain comprising the amino acid sequence of SEQ ID NO:41, a second heavy chain comprising the amino acid sequence of SEQ ID NO:27 and a second light chain comprising the amino acid sequence of SEQ ID NO:28.
- the invention further provides a vector, particularly an expression vector, comprising the isolated polynucleotide of the invention and a host cell comprising the isolated nucleic acid or the expression vector of the invention.
- the host cell is an eukaryotic cell, particularly a mammalian cell.
- a method of producing an antibody that specifically binds to BCMA or a bispecific BCMA antibody as described herein before comprising the steps of a) culturing the host cell as described above under conditions suitable for the expression of the antibody, and optionally b) recovering the an antibody that specifically binds to BCMA or the bispecific BCMA antibody.
- the invention also encompasses the antibody or bispecific antibody as produced by the method of the invention.
- composition comprising an antibody that specifically binds to BCMA or a bispecific BCMA antibody as described herein before and at least one pharmaceutically acceptable excipient.
- the pharmaceutical composition comprises an additional therapeutic agent.
- the antibody that specifically binds to BCMA or the bispecific BCMA antibody as described herein before or the pharmaceutical composition, for use in enhancing (a) T cell activation or (b) T cell effector functions.
- the antibody that specifically binds to BCMA or the bispecific BCMA antibody as described herein before or the pharmaceutical composition for use in the treatment of a disease.
- the disease is cancer, in particular multiple myeloma (MM).
- the antibody that specifically binds to BCMA or the bispecific BCMA antibody as described herein before for use in the treatment of cancer wherein the use is for administration in combination with a chemotherapeutic agent, radiation therapy and/ or other agents for use in cancer immunotherapy.
- the antibody that specifically binds to BCMA or the bispecific BCMA antibody as described herein before is for use in the treatment of cancer, wherein the use is for administration in combination with a T-cell activating anti-CD3 bispecific antibody.
- the T-cell activating anti-CD3 bispecific antibody is an anti- GPRC5D/anti-CD3 antibody.
- the invention provides a method of inhibiting the growth of tumor cells in an individual comprising administering to the individual an effective amount of the antibody that specifically binds to BCMA or the bispecific BCMA antibody as described herein before, or the pharmaceutical composition of the invention, to inhibit the growth of the tumor cells.
- the invention provides a method of treating or delaying cancer in an individual comprising administering to the individual an effective amount of the antibody that specifically binds to BCMA or the bispecific BCMA antibody as described herein before, or the pharmaceutical composition of the invention.
- the antibody that specifically binds to BCMA or the bispecific BCMA antibody as described herein before for the manufacture of a medicament for the treatment of a disease in an individual in need thereof, in particular for the manufacture of a medicament for the treatment of cancer, as well as a method of treating a disease in an individual, comprising administering to said individual a therapeutically effective amount of a composition comprising the antibody that specifically binds to BCMA or the bispecific BCMA antibody of the invention in a pharmaceutically acceptable form.
- the disease is cancer.
- the individual is a mammal, particularly a human.
- FIG. 1A shows a schematic illustration of the CD28 agonistic antibody variants as monovalent hu IgGl PGLALA isotype (“Fc silent”).
- Fig. IB shows a bispecific BCMA-CD28 antigen binding molecule in 1+1 format, wherein in the Fab molecule comprising the CD28 antigen binding domain the VH and VL domains are exchanged with each other (VH/VL crossfab) and wherein in the Fab molecule comprising the BCMA antigen binding domain certain amino acids in the CHI and CL domain are exchanged (charged variants) to allow better pairing with the light chain.
- Fig. 1A shows a schematic illustration of the CD28 agonistic antibody variants as monovalent hu IgGl PGLALA isotype (“Fc silent”).
- Fig. IB shows a bispecific BCMA-CD28 antigen binding molecule in 1+1 format, wherein in the Fab molecule comprising the CD28 antigen binding domain the VH and VL domains are exchanged with each other
- 1C shows a bispecific BCMA-CD28 antigen binding molecule in 1+1 format, wherein in the Fab molecule comprising the BCMA antigen binding domain the VH and VL domains are exchanged with each other (VH/VL crossfab) and wherein in the Fab molecule comprising the CD28 antigen binding domain certain amino acids in the CHI and CL domain are exchanged (charged variants) to allow better pairing with the light chain.
- Figures 2A and 2B show the binding of various BCMA-CD28 bispecific antigen binding molecules to CD28-expressing CHO cells (CHO-kl-huCD28 cells). All BCMA-CD28 bispecific antigen binding molecules were able to bind to human CD28 on CHO-kl-huCD28 cells in a concentration dependent manner, assessed by flow cytometry. However, binding to human CD28 does not reach saturation due to low affinity binders and is comparable among the CD28v8 bispecific antibodies (Fig. 2A) and CD28vl5 bispecific antibodies (Fig. 2B), respectively.
- Figures 3A and 3B show the binding of various BCMA-CD28 bispecific antigen binding molecules to BCMA-expressing CHO cells (CHO-huBCMA cells). All BCMA-CD28 bispecific antigen binding molecules were able to bind to human BCMA in a concentration dependent manner, assessed by flow cytometry.
- the molecules with the BCMA antibody PR described in WO 2020/127618 Al have lower ECso values compared to the molecules with the new BCMA antibodies as described herein, but maximal binding (Emax) is comparable among all molecules tested.
- Figures 3C to 3H show binding of two BCMA-CD28 bispecific antigen binding molecules Pl AG7191 and Pl AG7207 as well as the BCMA-targeted CD3 T cell engager Alnuctamab, to CHO cells expressing human BCMA with indicated point mutations. Shown is the binding in case no BCMA is present (Fig. 3C) and to human wt BCMA (Fig. 3D) as well as to BCMA variants human BCMA P33S (Fig. 3E), human BCMA P343del (Fig. 3F), human BCMA R27P (Fig. 3G) and human BCMA S3 Odel (Fig. 3H).
- BCMA-targeted CD3 T cell engager Teclistamab does not bind to two mutated BCMA variants, namely R27P (Fig. 3G) and S30del (Fig. 3H) and Elranatamab only weakly binds to BCMA with the R27P mutation at high concentrations. Neither of the tested bispecific molecules bind to BCMA-negative CHOkl cells (3C).
- Figures 4A to 4F show the dose-dependent activation of IL2 signaling in Jurkat IL2 reporter cells. Titrated amounts of (200.0-0.5 nM) of BCMA-CD28 vl5 BsAbs (Fig. 4A to 4C) and BCMA-CD28 v8 BsAbs (Fig. 4D to 4F) were added together with three different concentrations (20, 200, and 2000 pM) of GPRC5D-TCB to the mixture of the target (NCI-H929) and effector (Jurkat IL2 reporter) cells.
- Figures 5A to 5D provide a summary of the data obtained from Jurkat NFkB reporter assay.
- Fig. 5A shows the ECso values and Fig. 5B compares the efficacy of various BCMA- CD28 vl5 BsAbs and the non-targeted CD28 vl5 control
- Fig. 5C shows the ECso values and Fig. 5D compares the efficacy of the various BCMA-CD28 v8 BsAbs and the non-targeted CD28 control.
- the data was calculated from three independent experiments performed in triplicate. ECso data of Fig. 5 A and Fig. 5C were added together from experiments with different concentrations of GPRC5D-TCB, and the same was applied for efficacy data. All data are shown as mean ⁇ s.d.
- FIGS. 6A and 6B show that the dose-dependent activation of CD8 + T cells through BCMA-CD28 vl5 bispecific antibodies (BsAbs) occurs only in presence of a first signal (TCB) and BCMA expression.
- the BCMA-expressing MM cell line NCI-H929 (Fig. 6A) was cocultured with healthy donor PBMCs (ratio of 1 : 1).
- NCI-H929 BCMAko cells knock-out cells that do not express BCMA
- Fig. 6B Co-cultures were treated with GPRC5D-TCB (providing 1st signal) alone or in combination with BCMA- CD28 BsAbs or an untargeted CD28 (neg.
- FIGS. 8A and 8B show that the dose-dependent activation of CD4 + T cells through BCMA-CD28 v8 bispecific antibodies (BsAbs) occurs only in presence of a first signal (TCB) and BCMA expression.
- the BCMA-expressing MM cell line NCI-H929 (Fig. 8A) was cocultured with healthy donor PBMCs (ratio of 1 : 1).
- NCI-H929 BCMAko cells knock-out cells that do not express BCMA
- Fig. 8B Co-cultures were treated with GPRC5D-TCB (providing 1st signal) alone or in combination with BCMA- CD28 BsAbs or an untargeted CD28 (neg.
- FIGS 9A and 9B show that the dose-dependent proliferation of CD4 + T cells through BCMA-CD28 v8 bispecific antibodies (BsAbs). Proliferation occurs only in presence of a first signal (TCB) and BCMA expression.
- the BCMA-expressing MM cell line NCI-H929 (Fig. 9A) was co-cultured with healthy donor PBMCs (ratio of 1 : 1).
- NCI-H929 BCMAko cells knock-out cells that do not express BCMA
- Fig. 9B Co-cultures were treated with GPRC5D-TCB (providing 1st signal) alone or in combination with BCMA- CD28 BsAbs or an untargeted CD28 (neg.
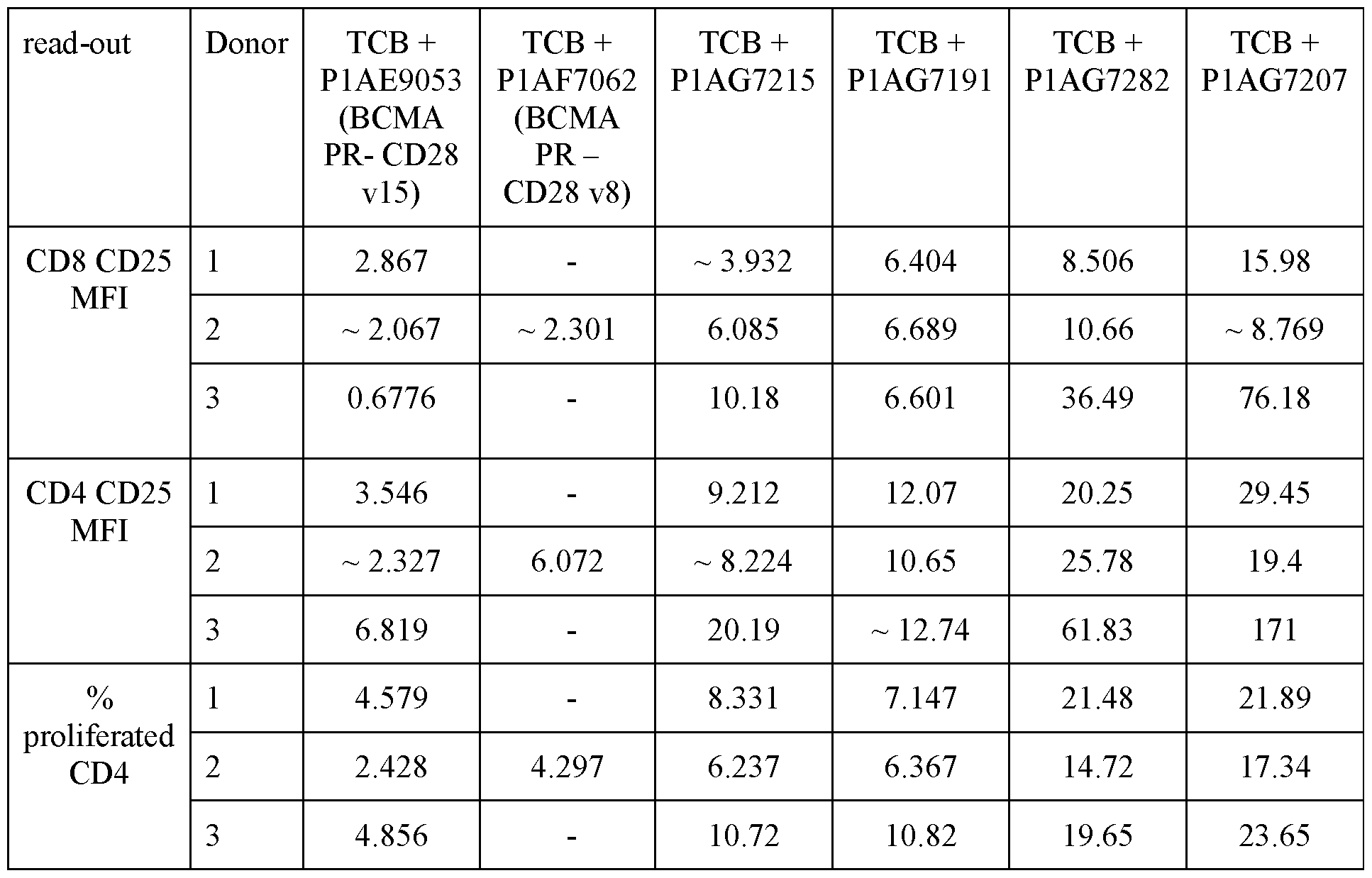
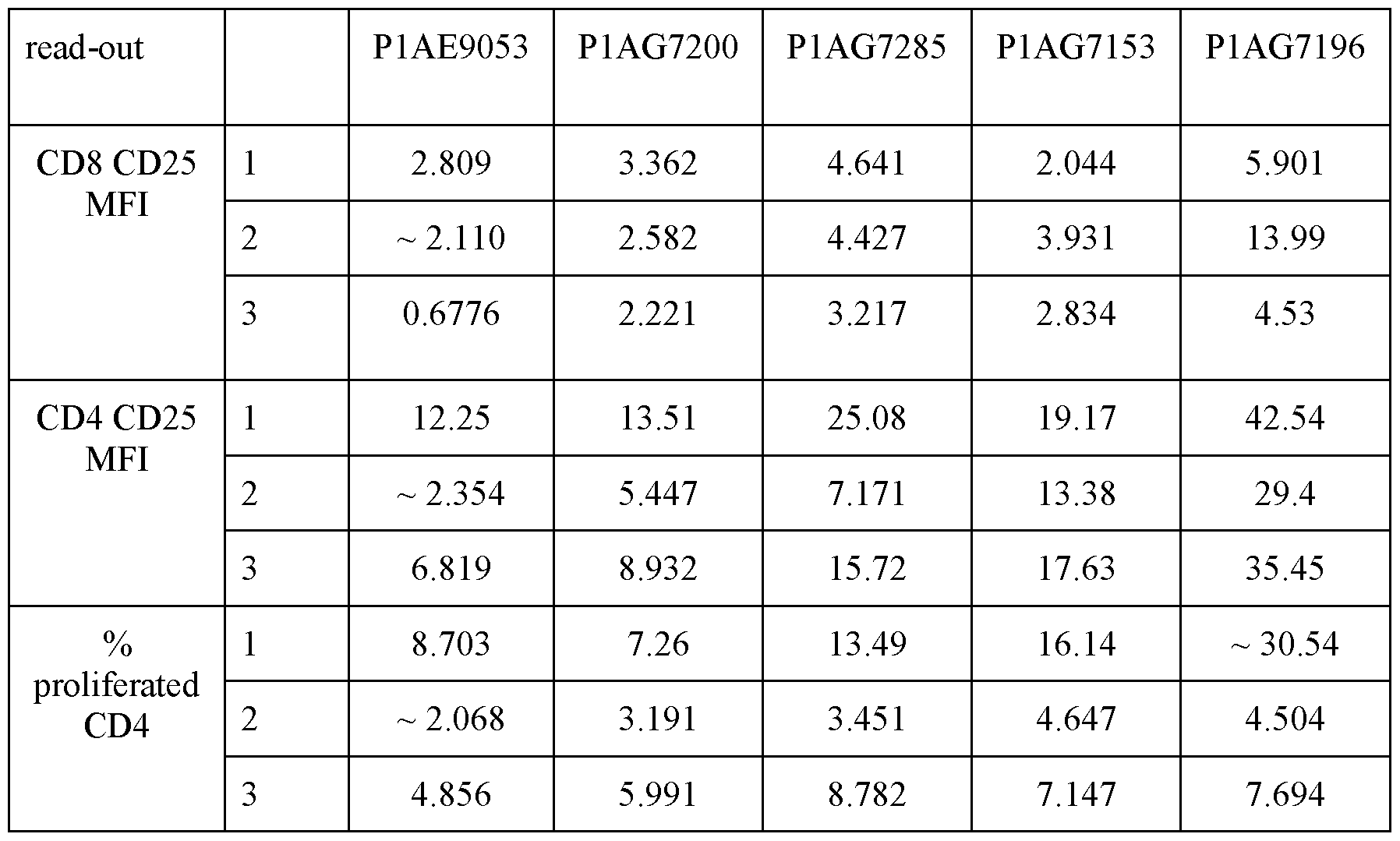
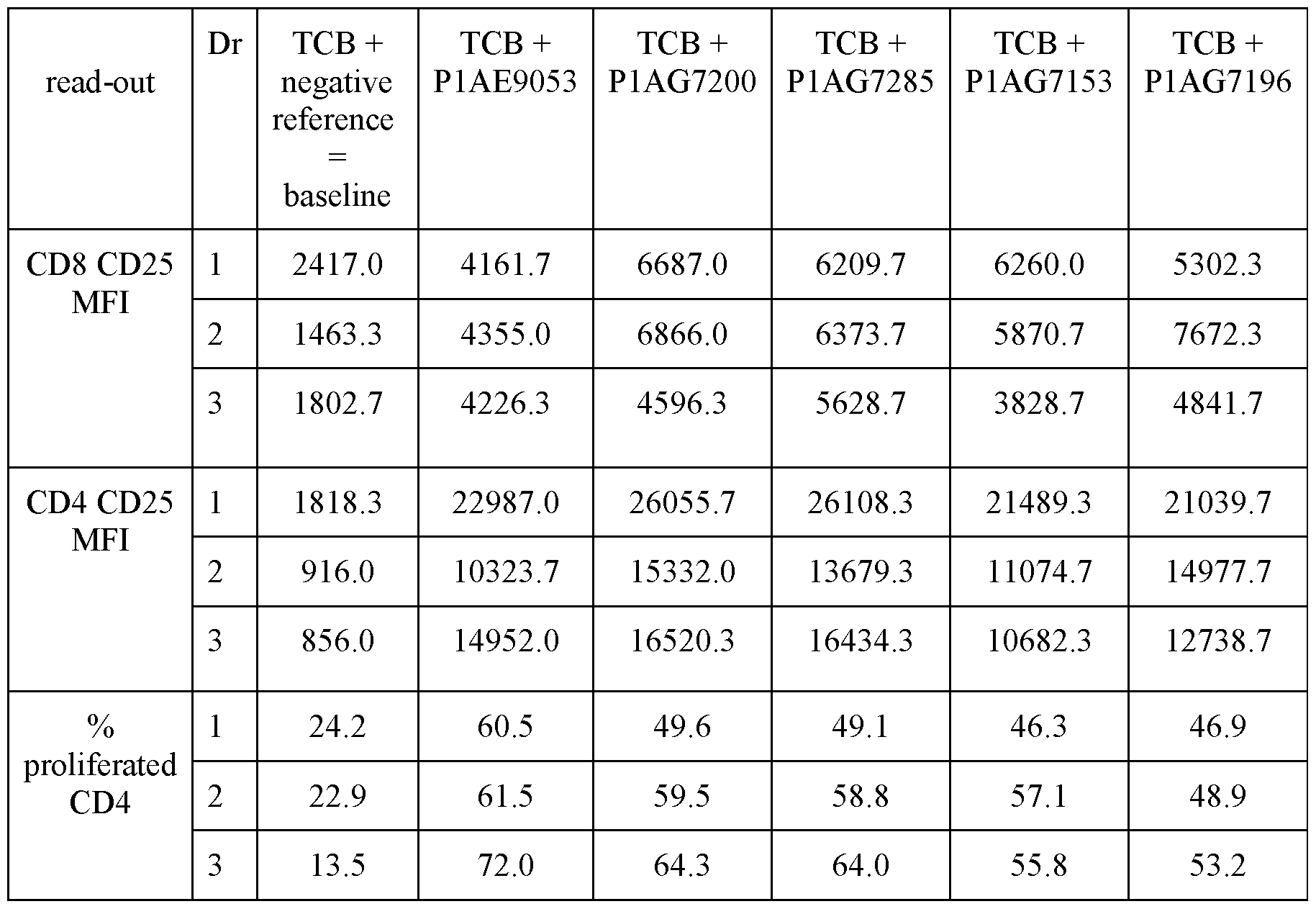
- Figure 10 shows results of an exemplary ex vivo test, using primary MM patients’ bone marrow samples as described in Example 4.3. T-cell activation was determined by flow cytometry, assessing the upregulation of CD25 on CD8 + T cells upon incubation with 1 nM GPRC5D-TCB in absence or presence of 200 nM of the indicated BCMA-CD28 bispecific antibody or untargeted negative reference molecule for 96 hours. All data shown refer to single tubes measurement per condition.
- Figure 11 shows results of an exemplary ex vivo test, using primary MM patients’ bone marrow samples.
- TCL Tumor Cell Lysis
- Figures 12A to 12D show the results of an ex vivo experiment, using primary MM patients’ bone marrow samples. T-cell activation, respective degranulation was determined by flow cytometry, assessing the upregulation of CD25 (see Fig. 12A and 12B) respective CD107A (see Fig. 12C and 12D) on CD4 + (see Fig. 12A and 12C) or CD8 + (see Fig. 12B and 12D) T cells upon incubation with 10 nM GPRC5D-TCB in absence or presence of 200 nM of the indicated BCMA-CD28 bispecific antibodies or reference molecules for 96 hours.
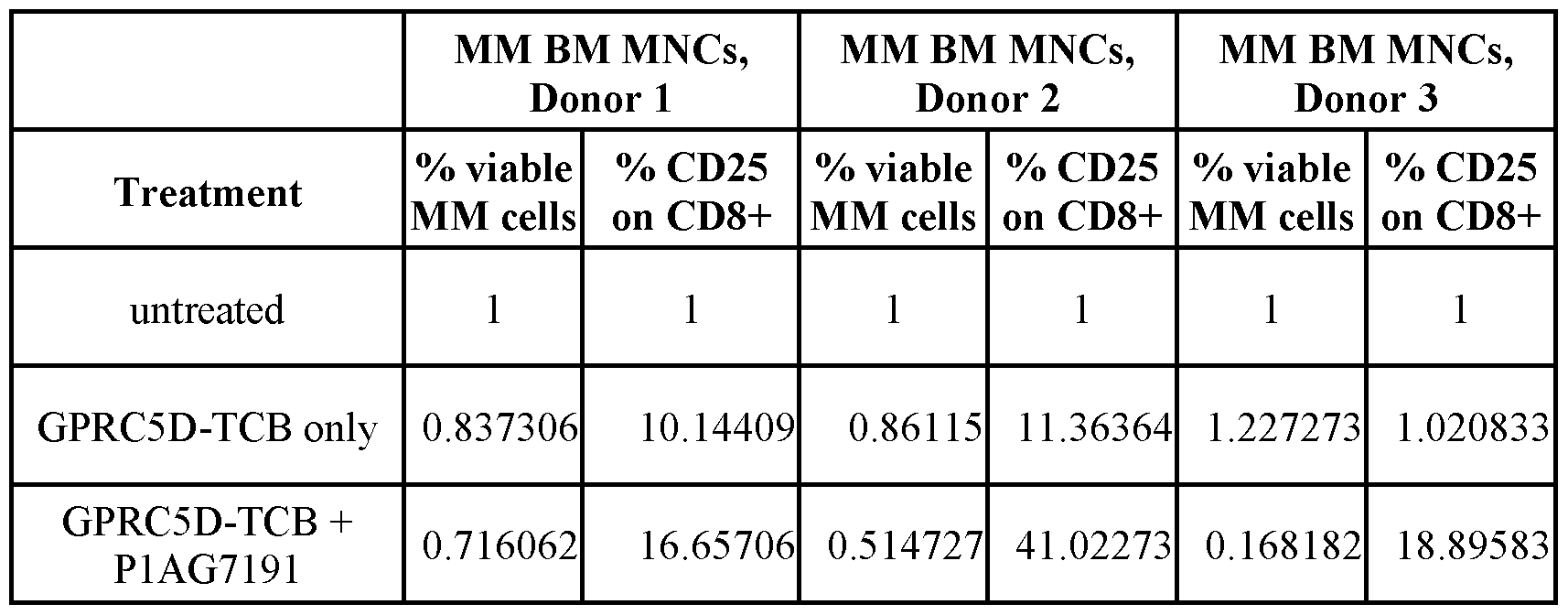
- Figures 13A and 13B show the results of an ex vivo test, using BM MNCs of primary MM patients’ bone marrow samples. Tumor Cell lysis respective T-cell activation was determined by flow cytometry, assessing the percentage of viable MM PCs (Fig. 13 A) or the upregulation of CD25 on CD8+ T cells (Fig. 13B) upon incubation with 0.01 nM GPRC5D-TCB in absence or presence of 800 nM of the indicated BCMA-CD28 bispecific antibodies or reference molecules for 96 hours.
- Figures 14A to 14F depict the results from an in vivo experiment testing the efficacy of GPRC5D x CD3 monotherapy and its combination with BCMA-CD28 bispecific antibodies (lOmg/kg P1AG7215 and 10 mg/kg P1AG7282) in the NCI-H929 tumor model.
- Figures 14A to 14 D show the tumor growth inhibition in single animals and
- Figure 14E shows the tumor volume as Median (+/- IQR) per treatment group. Animals with a terminal tumor load below the size at treatment start were defined as responders.
- Tumor volume at termination is shown for GPRC5D x CD3 monotherapy group and both combinations as Median (+/- IQR) is shown in Fig. 14F.
- Figures 15A to 15L show the results from an in vivo experiment testing the efficacy of GPRC5D x CD3 monotherapy and its combination with BCMA-CD28 var8 bispecific antibody P1AG7215 (20 mg/kg, 10 mg/kg, 5 mg/kg), BCMA-CD28 varl5 bispecific antibody P1AG7282 (20 mg/kg, 10 mg/kg, 2 mg/kg) and BCMA(PR)-CD28 bispecific antibodies P1AE9053 (10 mg/kg) and P1AF7062 (10 mg/kg) in the NCI-H929 tumor model. Tumor growth inhibition is shown in single animals ( Figures 15A to 15 J) and as Median (+/- IQR) per treatment group (Fig. 15K).
- Tumor volume at termination (study day 40) is shown for GPRC5D x CD3 monotherapy and combination groups as Median (+/- IQR) (C). in the NCI-H929 tumor model.
- Figures 15A to 15J show the tumor growth inhibition in single animals and Figure 15K shows the tumor volume as Median (+/- IQR) per treatment group. Animals with a terminal tumor load below the size at treatment start were defined as responders.
- Tumor volume at termination (study day 41) is shown for GPRC5D x CD3 monotherapy group and both combinations as Median (+/- IQR) is shown in Fig. 15L.
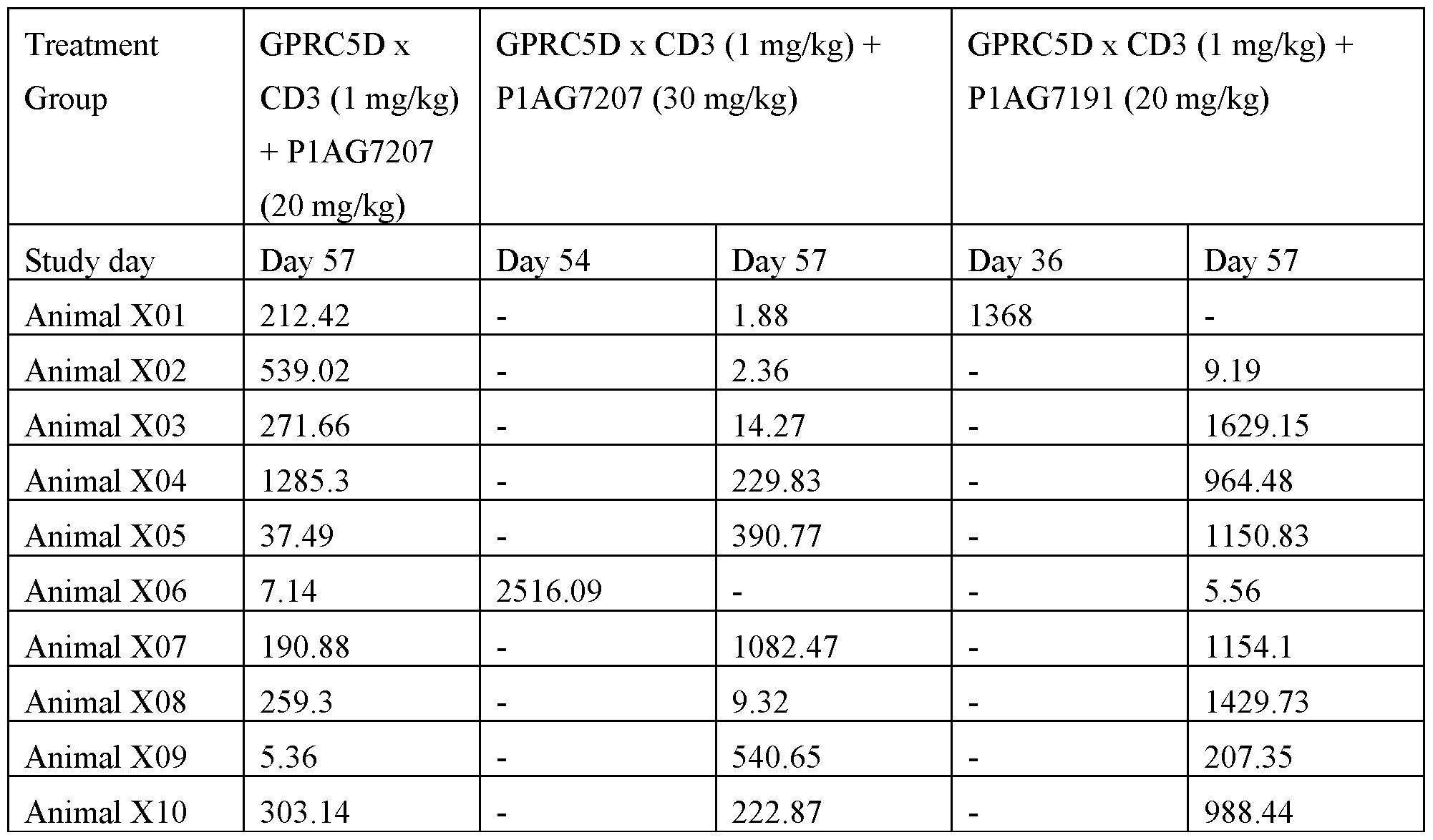
- Figures 16A to 16G show the results of an in vivo experiment testing the efficacy of GPRC5D x CD3 monotherapy (1 mg/kg) and combination with BCMA-CD28 (20 and 30 mg/kg) Pl AG7207 (20 and 30 mg/kg) and Pl AG7191 (20 mg/kg), respectively in the NCI-H929 tumor model.
- Figures 16A to 16E show the tumor growth inhibition in single animals.
- Figure 16F shows the tumor volume as Median (+/- IQR) per treatment group. Animals with a terminal tumor load below the size at treatment start were defined as responders. Treatment with the combination of GPRC5D x CD3 and 20 mg/kg Pl AG7207 led to 4 responders and with 30 mg/kg Pl AG7207 to 5 responders.
- FIGS 17A to 17G show the results of an in vivo experiment testing the efficacy of GPRC5D x CD3 monotherapy and in combination with BCMA-CD28 bispecific antibodies Pl AG7191 (40, 10 and 1 mg/kg) and P1AE9053 (10 mg/kg) in the NCI-H929 tumor model.
- Tumor growth inhibition is shown as Median (+/- IQR) per treatment group (Fig. 17A) and in single animals (Fig. 17B to 17G). Animals with a terminal tumor load below the size at treatment start were defined as responders.
- the two highest doses of Pl AG7191 (40 and 10 mg/kg) delay the time to tumor relapse compared to monotherapy and inhibit the tumor regrowth.
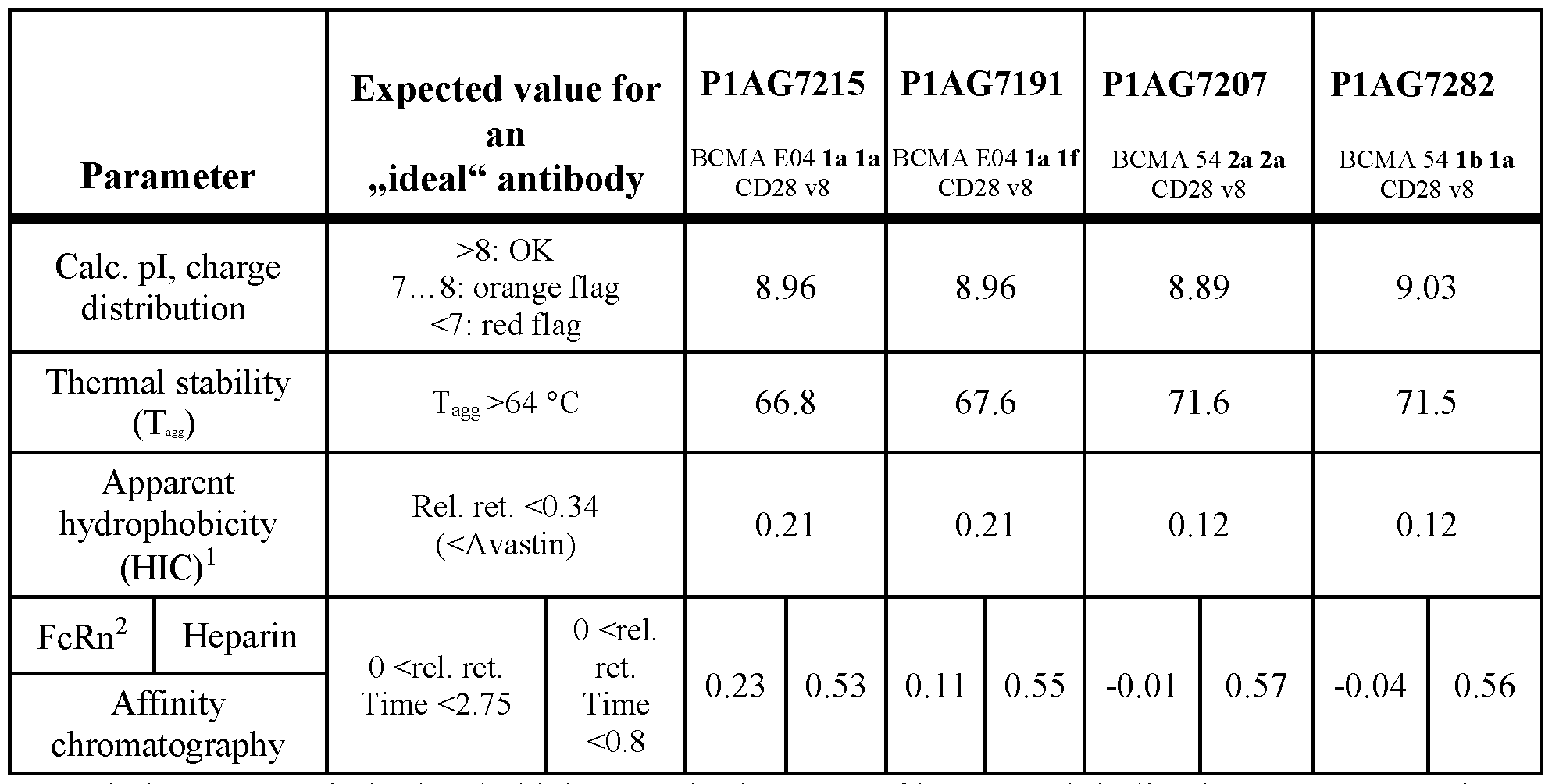
- Figure 18 shows a comparison of the plasma concentration-time profiles of bispecific antibodies P1AG7282, P1AG7215, P1AG7191 and P1AG7207 as measured in HuFcRN transgenic mice (PK study).
- Figure 19 shows a comparative heatMAPPS representation of the results observed in the MAPPs assay (Example 6.3.1) for the bispecific antibodies P1AG7282, P1AG7215, P1AG7191 and Pl AG7207. Clusters found in each domain are highlighted.
- antigen binding molecule refers in its broadest sense to a molecule that specifically binds an antigenic determinant.
- antigen binding molecules are antibodies, multispecific antibodies (e.g., bispecific antibodies), antibody fragments and scaffold antigen binding proteins.
- the term “antigen binding domain that binds to a tumor-associated antigen” or "moiety capable of specific binding to a tumor-associated antigen” refers to a polypeptide molecule that specifically binds to the tumor-associated antigen BCMA.
- the antigen binding domain is able to activate signaling through BCMA.
- the antigen binding domain is able to direct the entity to which it is attached (e.g. the CD28 antibody) to a BCMA-expressing cell, for example to a specific type of tumor cell.
- Antigen binding domains capable of specific binding to BCMA include antibodies and fragments thereof as further defined herein.
- antigen binding domains capable of specific binding to a tumor-associated antigen may include scaffold antigen binding proteins as further defined herein, e.g. binding domains which are based on designed repeat proteins or designed repeat domains (see e.g. WO 2002/020565).
- an antigen binding domain refers to the part of the molecule that comprises the area which specifically binds to and is complementary to part or all of an antigen.
- An antigen binding domain capable of specific antigen binding may be provided, for example, by one or more antibody variable domains (also called antibody variable regions).
- an antigen binding domain capable of specific antigen binding comprises an antibody light chain variable region (VL) and an antibody heavy chain variable region (VH).
- VL antibody light chain variable region
- VH antibody heavy chain variable region
- the "antigen binding domain capable of specific binding to a tumor-associated antigen can also be a Fab fragment or a crossFab fragment.
- the terms “first”, “second” or “third” with respect to antigen binding domains etc. are used for convenience of distinguishing when there is more than one of each type of moiety. Use of these terms is not intended to confer a specific order or orientation of the moiety unless explicitly so stated.
- antibody herein is used in the broadest sense and encompasses various antibody structures, including but not limited to monoclonal antibodies, polyclonal antibodies, monospecific and multispecific antibodies (e.g., bispecific antibodies), and antibody fragments so long as they exhibit the desired antigen-binding activity.
- the term “monoclonal antibody” as used herein refers to an antibody obtained from a population of substantially homogeneous antibodies, i.e., the individual antibodies comprising the population are identical and/or bind the same epitope, except for possible variant antibodies, e.g. containing naturally occurring mutations or arising during production of a monoclonal antibody preparation, such variants generally being present in minor amounts.
- polyclonal antibody preparations typically include different antibodies directed against different determinants (epitopes)
- each monoclonal antibody of a monoclonal antibody preparation is directed against a single determinant on an antigen.
- bispecific antibody denotes an antibody that has one or more binding sites each of which bind to the same epitope of the same antigen.
- the term “bispecific” means that the antigen binding molecule is able to specifically bind to at least two distinct antigenic determinants. Typically, a bispecific antigen binding molecule comprises two antigen binding sites, each of which is specific for a different antigenic determinant. However, a bispecific antigen binding molecule may also comprise additional antigen binding sites which bind to further antigenic determinants. In certain aspects, the bispecific antigen binding molecule is capable of simultaneously binding two antigenic determinants, particularly two antigenic determinants expressed on two distinct cells or on the same cell.
- the term “bispecific” in accordance with the present invention thus may also include a trispecific molecule, e.g. a bispecific molecule comprising a CD28 antibody and two antigen binding domains directed to two different target cell antigens.
- valent as used within the current application denotes the presence of a specified number of binding sites specific for one distinct antigenic determinant in an antigen binding molecule that are specific for one distinct antigenic determinant.
- bivalent tetravalent
- hexavalent denote the presence of two binding sites, four binding sites, and six binding sites specific for a certain antigenic determinant, respectively, in an antigen binding molecule.
- the bispecific antigen binding molecules according to the invention can be monovalent for a certain antigenic determinant, meaning that they have only one binding site for said antigenic determinant or they can be bivalent or tetravalent for a certain antigenic determinant, meaning that they have two binding sites or four binding sites, respectively, for said antigenic determinant.
- full length antibody “intact antibody”, and “whole antibody” are used herein interchangeably to refer to an antibody having a structure substantially similar to a native antibody structure.
- Native antibodies refer to naturally occurring immunoglobulin molecules with varying structures.
- native IgG-class antibodies are heterotetrameric glycoproteins of about 150,000 daltons, composed of two light chains and two heavy chains that are disulfide-bonded. From N- to C-terminus, each heavy chain has a variable region (VH), also called a variable heavy domain or a heavy chain variable domain, followed by three constant domains (CHI, CH2, and CH3), also called a heavy chain constant region.
- each light chain has a variable region (VL), also called a variable light domain or a light chain variable domain, followed by a light chain constant domain (CL), also called a light chain constant region.
- the heavy chain of an antibody may be assigned to one of five types, called a (IgA), 5 (IgD), a (IgE), y (IgG), or p (IgM), some of which may be further divided into subtypes, e.g. yl (IgGl), y2 (IgG2), y3 (IgG3), y4 (IgG4), al (IgAl) and a2 (IgA2).
- the light chain of an antibody may be assigned to one of two types, called kappa (K) and lambda (X), based on the amino acid sequence of its constant domain.
- antibody fragment refers to a molecule other than an intact antibody that comprises a portion of an intact antibody that binds the antigen to which the intact antibody binds.
- antibody fragments include but are not limited to Fv, Fab, Fab', Fab’-SH, F(ab')2; diabodies, triabodies, tetrabodies, crossFab fragments; linear antibodies; single-chain antibody molecules (e.g. scFv); and single domain antibodies.
- scFv single domain antibodies.
- Diabodies are antibody fragments with two antigen-binding sites that may be bivalent or bispecific, see, for example, EP 404,097; WO 1993/01161; Hudson et al., Nat Med 9, 129-134 (2003); and Hollinger et al., Proc Natl Acad Sci USA 90, 6444-6448 (1993). Triabodies and tetrabodies are also described in Hudson et al., Nat Med 9, 129-134 (2003).
- Single-domain antibodies are antibody fragments comprising all or a portion of the heavy chain variable domain or all or a portion of the light chain variable domain of an antibody.
- a single-domain antibody is a human single-domain antibody (Domantis, Inc., Waltham, MA; see e.g. U.S. Patent No. 6,248,516 Bl).
- Antibody fragments can be made by various techniques, including but not limited to proteolytic digestion of an intact antibody as well as production by recombinant host cells (e.g. E. coli or phage), as described herein.
- Fab fragments containing each the heavy- and light-chain variable domains and also the constant domain of the light chain and the first constant domain (CHI) of the heavy chain.
- Fab fragment or Fab molecule refers to an antibody fragment comprising a light chain fragment comprising a variable light chain (VL) domain and a constant domain of a light chain (CL), and a variable heavy chain (VH) domain and a first constant domain (CHI) of a heavy chain.
- Fab’ fragments differ from Fab fragments by the addition of a few residues at the carboxy terminus of the heavy chain CHI domain including one or more cysteins from the antibody hinge region.
- Fab’-SH are Fab’ fragments in which the cysteine residue(s) of the constant domains bear a free thiol group. Pepsin treatment yields an F(ab')2 fragment that has two antigen-combining sites (two Fab fragments) and a part of the Fc region.
- a “conventional Fab fragment” is comprised of a VL-CL light chain and a VH-CH1 heavy chain.
- crossFab fragment or “xFab fragment” or “crossover Fab fragment” refers to a Fab fragment, wherein either the variable regions or the constant regions of the heavy and light chain are exchanged.
- Two different chain compositions of a crossover Fab molecule are possible and comprised in the bispecific antibodies of the invention: On the one hand, the variable regions of the Fab heavy and light chain are exchanged, i.e. the crossover Fab molecule comprises a peptide chain composed of the light chain variable (VL) domain and the heavy chain constant domain (CHI), and a peptide chain composed of the heavy chain variable domain (VH) and the light chain constant domain (CL).
- VL light chain variable
- CHI heavy chain constant domain
- VH heavy chain variable domain
- CL light chain constant domain
- This crossover Fab molecule is also referred to as CrossFab (VLVH).
- the crossover Fab molecule comprises a peptide chain composed of the heavy chain variable domain (VH) and the light chain constant domain (CL), and a peptide chain composed of the light chain variable domain (VL) and the heavy chain constant domain (CHI).
- VH heavy chain variable domain
- CL light chain constant domain
- CHI heavy chain constant domain
- a “single chain Fab fragment” or “scFab” is a polypeptide consisting of an antibody heavy chain variable domain (VH), an antibody constant domain 1 (CHI), an antibody light chain variable domain (VL), an antibody light chain constant domain (CL) and a linker, wherein said antibody domains and said linker have one of the following orders in N-terminal to C-terminal direction: a) VH-CH1 -linker- VL-CL, b) VL-CL-linker-VH-CHl, c) VH-CL-linker-VL-CHl or d) VL-CH1 -linker- VH-CL; and wherein said linker is a polypeptide of at least 30 amino acids, preferably between 32 and 50 amino acids.
- Said single chain Fab fragments are stabilized via the natural disulfide bond between the CL domain and the CHI domain.
- these single chain Fab molecules might be further stabilized by generation of interchain disulfide bonds via insertion of cysteine residues (e.g. position 44 in the variable heavy chain and position 100 in the variable light chain according to Kabat numbering).
- a “crossover single chain Fab fragment” or “x-scFab” is a is a polypeptide consisting of an antibody heavy chain variable domain (VH), an antibody constant domain 1 (CHI), an antibody light chain variable domain (VL), an antibody light chain constant domain (CL) and a linker, wherein said antibody domains and said linker have one of the following orders in N- terminal to C-terminal direction: a) VH-CL-linker-VL-CHl and b) VL-CH1 -linker- VH-CL; wherein VH and VL form together an antigen-binding site which binds specifically to an antigen and wherein said linker is a polypeptide of at least 30 amino acids.
- these x-scFab molecules might be further stabilized by generation of interchain disulfide bonds via insertion of cysteine residues (e.g. position 44 in the variable heavy chain and position 100 in the variable light chain according to Kabat numbering).
- a “single-chain variable fragment (scFv)” is a fusion protein of the variable regions of the heavy (VH) and light chains (VL) of an antibody, connected with a short linker peptide of ten to about 25 amino acids.
- the linker is usually rich in glycine for flexibility, as well as serine or threonine for solubility, and can either connect the N-terminus of the VH with the C-terminus of the VL, or vice versa. This protein retains the specificity of the original antibody, despite removal of the constant regions and the introduction of the linker.
- scFv antibodies are, e.g. described in Houston, J.S., Methods in Enzymol. 203 (1991) 46-96).
- antibody fragments comprise single chain polypeptides having the characteristics of a VH domain, namely being able to assemble together with a VL domain, or of a VL domain, namely being able to assemble together with a VH domain to a functional antigen binding site and thereby providing the antigen binding property of full length antibodies.
- fibronectin and designed ankyrin repeat proteins have been used as alternative scaffolds for antigenbinding domains, see, e.g., Gebauer and Skerra, Engineered protein scaffolds as next-generation antibody therapeutics. Curr Opin Chem Biol 13:245-255 (2009) and Stumpp et al., Darpins: A new generation of protein therapeutics. Drug Discovery Today 13: 695-701 (2008).
- a scaffold antigen binding protein is selected from the group consisting of CTLA-4 (Evibody), Lipocalins (Anticalin), a Protein A-derived molecule such as Z-domain of Protein A (Affibody), an A-domain (Avimer/Maxibody), a serum transferrin (/ra//.s-body); a designed ankyrin repeat protein (DARPin), a variable domain of antibody light chain or heavy chain (single-domain antibody, sdAb), a variable domain of antibody heavy chain (nanobody, aVH), VNAR fragments, a fibronectin (AdNectin), a C-type lectin domain (Tetranectin); a variable domain of a new antigen receptor beta-lactamase (VNAR fragments), a human gammacrystallin or ubiquitin (Affilin molecules); a kunitz type domain of human protease inhibitors, microbodies such as the group consisting of CTLA
- CTLA-4 Cytotoxic T Lymphocyte-associated Antigen 4
- CTLA-4 is a CD28-family receptor expressed on mainly CD4 + T-cells. Its extracellular domain has a variable domain- like Ig fold. Loops corresponding to CDRs of antibodies can be substituted with heterologous sequence to confer different binding properties.
- CTLA-4 molecules engineered to have different binding specificities are also known as Evibodies (e.g. US7166697B1). Evibodies are around the same size as the isolated variable region of an antibody (e.g. a domain antibody). For further details, see Journal of Immunological Methods 248 (1-2), 31-45 (2001).
- Lipocalins are a family of extracellular proteins which transport small hydrophobic molecules such as steroids, bilins, retinoids and lipids. They have a rigid beta-sheet secondary structure with a number of loops at the open end of the conical structure which can be engineered to bind to different target antigens. Anticalins are between 160-180 amino acids in size, and are derived from lipocalins. For further details, see Biochim Biophys Acta 1482: 337-350 (2000), US7250297B1 and US20070224633.
- An affibody is a scaffold derived from Protein A of Staphylococcus aureus which can be engineered to bind to antigen.
- the domain consists of a three-helical bundle of approximately 58 amino acids. Libraries have been generated by randomization of surface residues. For further details, see Protein Eng. Des. Sei. 2004, 17, 455-462 and EP 1641818A1. Avimers are multidomain proteins derived from the A-domain scaffold family. The native domains of approximately 35 amino acids adopt a defined disulfide bonded structure. Diversity is generated by shuffling of the natural variation exhibited by the family of A-domains. For further details, see Nature Biotechnology 23(12), 1556 - 1561 (2005) and Expert Opinion on Investigational Drugs 16(6), 909-917 (June 2007). A transferrin is a monomeric serum transport glycoprotein.
- Transferrins can be engineered to bind different target antigens by insertion of peptide sequences in a permissive surface loop.
- engineered transferrin scaffolds include the Transbody.
- Designed Ankyrin Repeat Proteins are derived from Ankyrin which is a family of proteins that mediate attachment of integral membrane proteins to the cytoskeleton.
- a single ankyrin repeat is a 33 residue motif consisting of two alpha-helices and a beta-turn. They can be engineered to bind different target antigens by randomizing residues in the first alpha-helix and a beta-turn of each repeat.
- a singledomain antibody is an antibody fragment consisting of a single monomeric variable antibody domain.
- the first single domains were derived from the variable domain of the antibody heavy chain from camelids (nanobodies or VHH fragments).
- the term single-domain antibody includes an autonomous human heavy chain variable domain (aVH) or VNAR fragments derived from sharks.
- Fibronectin is a scaffold which can be engineered to bind to antigen.
- Adnectins consists of a backbone of the natural amino acid sequence of the 10th domain of the 15 repeating units of human fibronectin type III (FN3). Three loops at one end of the betasandwich can be engineered to enable an Adnectin to specifically recognize a therapeutic target of interest. For further details, see Protein Eng. Des. Sei. 18, 435- 444 (2005), US20080139791, W02005056764 and US6818418B1.
- Peptide aptamers are combinatorial recognition molecules that consist of a constant scaffold protein, typically thioredoxin (TrxA) which contains a constrained variable peptide loop inserted at the active site. For further details, see Expert Opin. Biol. Ther. 5, 783-797 (2005).
- Microbodies are derived from naturally occurring microproteins of 25-50 amino acids in length which contain 3-4 cysteine bridges - examples of microproteins include KalataBI and conotoxin and knottins.
- the microproteins have a loop which can beengineered to include upto 25 amino acids without affecting the overall fold of the microprotein. For further details of engineered knottin domains, see W02008098796.
- an “antibody that binds to the same epitope” as a reference molecule refers to an antibody that blocks binding of the reference molecule to its antigen in a competition assay by 50% or more, and conversely, the reference molecule blocks binding of the antigen binding molecule to its antigen in a competition assay by 50% or more.
- an antigen binding domain refers to the part of an antigen binding molecule that comprises the area which specifically binds to and is complementary to part or all of an antigen. Where an antigen is large, an antigen binding molecule may only bind to a particular part of the antigen, which part is termed an epitope.
- An antigen binding domain may be provided by, for example, one or more variable domains (also called variable regions).
- an antigen binding domain comprises an antibody light chain variable domain (VL) and an antibody heavy chain variable domain (VH).
- VL antibody light chain variable domain
- VH antibody heavy chain variable domain
- Useful antigenic determinants can be found, for example, on the surfaces of tumor cells, on the surfaces of virus-infected cells, on the surfaces of other diseased cells, on the surface of immune cells, free in blood serum, and/or in the extracellular matrix (ECM).
- ECM extracellular matrix
- the proteins useful as antigens herein can be any native form the proteins from any vertebrate source, including mammals such as primates (e.g. humans) and rodents (e.g. mice and rats), unless otherwise indicated.
- the antigen is a human protein.
- the term encompasses the “full-length”, unprocessed protein as well as any form of the protein that results from processing in the cell.
- the term also encompasses naturally occurring variants of the protein, e.g. splice variants or allelic variants.
- ELISA enzyme-linked immunosorbent assay
- SPR Surface Plasmon Resonance
- the extent of binding of an antigen binding molecule to an unrelated protein is less than about 10% of the binding of the antigen binding molecule to the antigen as measured, e.g. by SPR.
- an molecule that binds to the antigen has a dissociation constant (Kd) of ⁇ 1 pM, ⁇ 100 nM, ⁇ 10 nM, ⁇ 1 nM, ⁇ 0.1 nM, ⁇ 0.01 nM, or ⁇ 0.001 nM (e.g. 10' 8 M or less, e.g. from 10' 8 M to 10' 13 M, e.g. from 10' 9 M to 10' 13 M).
- Binding affinity refers to the strength of the sum total of non-covalent interactions between a single binding site of a molecule (e.g. an antibody) and its binding partner (e.g. an antigen). Unless indicated otherwise, as used herein, “binding affinity” refers to intrinsic binding affinity which reflects a 1 : 1 interaction between members of a binding pair (e.g. antibody and antigen).
- the affinity of a molecule X for its partner Y can generally be represented by the dissociation constant (Kd), which is the ratio of dissociation and association rate constants (koff and kon, respectively).
- Kd dissociation constant
- equivalent affinities may comprise different rate constants, as long as the ratio of the rate constants remains the same. Affinity can be measured by common methods known in the art, including those described herein. A particular method for measuring affinity is Surface Plasmon Resonance (SPR).
- an “activating T cell antigen” as used herein refers to an antigenic determinant expressed on the surface of a T lymphocyte, particularly a cytotoxic T lymphocyte, which is capable of inducing T cell activation upon interaction with an antibody. Specifically, interaction of an antibody with an activating T cell antigen may induce T cell activation by triggering the signaling cascade of the T cell receptor complex.
- the activating T cell antigen is CD3, particularly the epsilon subunit of CD3 (see UniProt no. P07766 (version 189), NCBI RefSeq no. NP 000724.1, SEQ ID NO: 167 for the human sequence; or UniProt no. Q95LI5 (version 49), NCBI GenBank no. BAB71849.1, SEQ ID NO: 168 for the cynomolgus [Macaca fascicularis] sequence).
- T cell activation refers to one or more cellular response of a T lymphocyte, particularly a cytotoxic T lymphocyte, selected from proliferation, differentiation, cytokine secretion, cytotoxic effector molecule release, cytotoxic activity, and expression of activation markers. Suitable assays to measure T cell activation are known in the art and described herein.
- T cell effector functions refers to the activities of T cells that play a key role in the adaptive immune system. T cells are responsible for initiating and coordinating the body’s immune response against foreign invaders, such as viruses or bacteria as well as tumor cells. Effector functions refer to the various activities carried out by T cells to eliminate these offenders, which include releasing cytokines, stimulating other cells, and directly attacking and eliminating infected cells.
- TAA tumor-associated antigen
- a target cell for example a cell in a tumor such as a cancer cell, a cell of the tumor stroma, a malignant B lymphocyte or a melanoma cell.
- the target cell antigen is an antigen on the surface of a tumor cell.
- TAA is BCMA.
- BCMA refers to B cell maturation antigen, also termed tumor necrosis factor receptor superfamily member 17 (TNFRS17) or CD269, and is a type III transmembrane protein without a signal-peptide and containing cysteine-rich extracellular domains.
- Ligands for BCMA include B cell activating factor (BAFF) and a proliferation-inducing ligand (APRIL), of which APRIL has a higher affinity for BCMA.
- BAFF B cell activating factor
- APRIL proliferation-inducing ligand
- BCMA is preferentially expressed by mature B lymphocytes, with minimal expression in hematopoietic stem cells or non-hematopoietic tissue, and is essential for the survival of long-lived bone marrow plasma cells.
- BCMA soluble BCMA
- BAFF transmembrane activator and calcium modulator and cyclophilin ligand interactor
- BCMA as used herein refers to any BCMA protein from any vertebrate source, including mammals such as primates (e.g. humans) non-human primates (e.g. cynomolgus monkeys) and rodents (e.g. mice and rats), unless otherwise indicated.
- the amino acid sequence of human BCMA is shown in UniProt (www.uniprot.org) accession no. Q02223 (SEQ ID NO: 99).
- CD28 Cluster of differentiation 28, Tp44 refers to any CD28 protein from any vertebrate source, including mammals such as primates (e.g. humans) non-human primates (e.g. cynomolgus monkeys) and rodents (e.g. mice and rats), unless otherwise indicated.
- CD28 is expressed on T cells and provides co-stimulatory signals required for T cell activation and survival. T cell stimulation through CD28 in addition to the T-cell receptor (TCR) can provide a potent signal for the production of various interleukins.
- CD28 is the receptor for CD80 (B7.1) and CD86 (B7.2) proteins and is the only B7 receptor constitutively expressed on naive T cells.
- the amino acid sequence of human CD28 is shown in UniProt (www.uniprot.org) accession no. Pl 0747 (SEQ ID NO: 100).
- an “agonistic antibody” refers to an antibody that comprises an agonistic function against a given receptor.
- an agonist ligand factor
- the tertiary structure of the receptor protein changes, and the receptor is activated (when the receptor is a membrane protein, a cell growth signal or such is usually transducted).
- the receptor is a dimerforming type, an agonistic antibody can dimerize the receptor at an appropriate distance and angle, thus acting similarly to a ligand.
- An appropriate anti-receptor antibody can mimic dimerization of receptors performed by ligands, and thus can become an agonistic antibody.
- a “CD28 agonistic antibody” or “CD28 conventional agonistic antibody” is an antibody that mimics CD28 natural ligands (CD80 or CD86) in their role to enhance T cell activation in presence of a T cell receptor signal (“signal 2”).
- a T cell needs two signals to become fully activated.
- signal 1 arises from the interaction of T cell receptor (TCR) molecules with peptide/major histocompatibility complex (MHC) complexes on antigen presenting cells (APCs) and “signal 2” is provided by engagement of a costimulatory receptor, e.g. CD28.
- a CD28 agonistic antibody is able to costimulate T cells (signal 2).
- CD28 agonistic antibody is not capable of fully activating T cells without additional stimulation of the TCR.
- CD28 superagonistic antibodies There is however a subclass of CD28 specific antigen binding molecules, the so-called CD28 superagonistic antibodies.
- a “CD28 superagonistic antibody” is a CD28 antibody which is capable of fully activating T cells without additional stimulation of the TCR.
- a CD28 superagonistic anitbody is capable to induce T cell proliferation and cytokine secretion without prior T cell activation (signal 1).
- variable domain refers to the domain of an antibody heavy or light chain that is involved in binding the antigen binding molecule to antigen.
- the variable domains of the heavy chain and light chain (VH and VL, respectively) of a native antibody generally have similar structures, with each domain comprising four conserved framework regions (FRs) and three hypervariable regions (HVRs). See, e.g., Kindt et al., Kuby Immunology, 6th ed., W.H. Freeman and Co., page 91 (2007).
- a single VH or VL domain may be sufficient to confer antigen-binding specificity.
- hypervariable region refers to each of the regions of an antigen binding variable domain which are hypervariable in sequence and which determine antigen binding specificity, for example “complementarity determining regions” (“CDRs”).
- CDRs complementarity determining regions
- antigen binding domains comprise six CDRs: three in the VH (CDR-H1, CDR-H2, CDR-H3), and three in the VL (CDR-L1, CDR-L2, CDR-L3).
- Exemplary CDRs herein include:
- the CDRs are determined according to Kabat et al., supra.
- the CDR designations can also be determined according to Chothia, supra, McCallum, supra, or any other scientifically accepted nomenclature.
- Kabat et al. also defined a numbering system for variable region sequences that is applicable to any antibody.
- One of ordinary skill in the art can unambiguously assign this system of "Kabat numbering" to any variable region sequence, without reliance on any experimental data beyond the sequence itself.
- Kabat numbering refers to the numbering system set forth by Kabat et al., U.S. Dept, of Health and Human Services, "Sequence of Proteins of Immunological Interest" (1983). Unless otherwise specified, references to the numbering of specific amino acid residue positions in an antibody variable region are according to the Kabat numbering system.
- affinity matured in the context of antigen binding molecules (e.g., antibodies) refers to an antigen binding molecule that is derived from a reference antigen binding molecule, e.g., by mutation, binds to the same antigen, preferably binds to the same epitope, as the reference antibody; and has a higher affinity for the antigen than that of the reference antigen binding molecule.
- Affinity maturation generally involves modification of one or more amino acid residues in one or more CDRs of the antigen binding molecule.
- the affinity matured antigen binding molecule binds to the same epitope as the initial reference antigen binding molecule.
- FR Framework
- the FR of a variable domain generally consists of four FR domains: FR1, FR2, FR3, and FR4. Accordingly, the HVR and FR sequences generally appear in the following sequence in VH (or VL): FR1-H1(L1)-FR2-H2(L2)-FR3-H3(L3)-FR4.
- An “acceptor human framework” for the purposes herein is a framework comprising the amino acid sequence of a light chain variable domain (VL) framework or a heavy chain variable domain (VH) framework derived from a human immunoglobulin framework or a human consensus framework, as defined below.
- acceptor human framework “derived from” a human immunoglobulin framework or a human consensus framework may comprise the same amino acid sequence thereof, or it may contain amino acid sequence changes. In some embodiments, the number of amino acid changes are 10 or less, 9 or less, 8 or less, 7 or less, 6 or less, 5 or less, 4 or less, 3 or less, or 2 or less. In some embodiments, the VL acceptor human framework is identical in sequence to the VL human immunoglobulin framework sequence or human consensus framework sequence.
- chimeric antibody refers to an antibody in which a portion of the heavy and/or light chain is derived from a particular source or species, while the remainder of the heavy and/or light chain is derived from a different source or species.
- the “class” of an antibody refers to the type of constant domain or constant region possessed by its heavy chain.
- the heavy chain constant domains that correspond to the different classes of immunoglobulins are called a, 5, 8, y, and p respectively.
- a “humanized” antibody refers to a chimeric antibody comprising amino acid residues from non-human HVRs and amino acid residues from human FRs.
- a humanized antibody will comprise substantially all of at least one, and typically two, variable domains, in which all or substantially all of the HVRs (e.g., CDRs) correspond to those of a non- human antibody, and all or substantially all of the FRs correspond to those of a human antibody.
- a humanized antibody optionally may comprise at least a portion of an antibody constant region derived from a human antibody.
- a “humanized form” of an antibody, e.g., a non-human antibody refers to an antibody that has undergone humanization.
- Other forms of "humanized antibodies" encompassed by the present invention are those in which the constant region has been additionally modified or changed from that of the original antibody to generate the properties according to the invention, especially in regard to Clq binding and/or Fc receptor (FcR) binding.
- a “human” antibody is one which possesses an amino acid sequence which corresponds to that of an antibody produced by a human or a human cell or derived from a non-human source that utilizes human antibody repertoires or other human antibody-encoding sequences.
- This definition of a human antibody specifically excludes a humanized antibody comprising non- human antigen-binding residues.
- a “human” or “humanized” antibody comprises a constant region of human origin, particularly of the IgG isotype, more particularly of the IgGl isotype, comprising a human CHI, CH2, CH3 and/or CL domain.
- the term "CL domain” denotes the constant part of an antibody light chain polypeptide. Exemplary sequences of human constant domains are given in SEQ ID Nos: 101 and 102 (human kappa and lambda CL domains, respectively).
- CHI domain denotes the part of an antibody heavy chain polypeptide that extends approximately from EU position 118 to EU position 215 (EU numbering system according to Kabat).
- a CHI domain has the amino acid sequence of ASTKGPSVFP LAPSSKSTSG GTAALGCLVK DYFPEPVTVS WNSGALTSGV HTFPAVLQSS GLYSLSSVVT VPSSSLGTQT YICNVNHKPS NTKVDKKV (SEQ ID NO: 103).
- a segment having the amino acid sequence of EPKSC (SEQ ID NO: 104) is following to link the CHI domain to the hinge region.
- hinge region denotes the part of an antibody heavy chain polypeptide that joins in a wild-type antibody heavy chain the CHI domain and the CH2 domain, e. g. from about position 216 to about position 230 according to the EU number system of Kabat, or from about position 226 to about position 230 according to the EU number system of Kabat.
- the hinge regions of other IgG subclasses can be determined by aligning with the hinge-region cysteine residues of the IgGl subclass sequence.
- the hinge region is normally a dimeric molecule consisting of two polypeptides with identical amino acid sequence.
- the hinge region generally comprises up to 25 amino acid residues and is flexible allowing the associated target binding sites to move independently.
- the hinge region can be subdivided into three domains: the upper, the middle, and the lower hinge domain (see e.g. Roux, et al., J. Immunol. 161 (1998) 4083).
- the hinge region has the amino acid sequence DKTHTCPXCP (SEQ ID NO: 105), wherein X is either S or P. In one aspect, the hinge region has the amino acid sequence HTCPXCP (SEQ ID NO: 106), wherein X is either S or P. In one aspect, the hinge region has the amino acid sequence CPXCP (SEQ ID NO: 107), wherein X is either S or P.
- Fc domain or “Fc region” herein is used to define a C-terminal region of an antibody heavy chain that contains at least a portion of the constant region.
- the term includes native sequence Fc regions and variant Fc regions.
- Fc domain may only refer to an IgG CH2 and an IgG CH3 domain.
- the “CH2 domain” of a human IgG Fc region usually extends from an amino acid residue at about EU position 231 to an amino acid residue at about EU position 340 (EU numbering system according to Kabat).
- a CH2 domain has the amino acid sequence of APELLGGPSV FLFPPKPKDT LMISRTPEVT CVWDVSHEDP EVKFNWYVDG VEVHNAKTKP REEQESTYRW SVLTVLHQDW LNGKEYKCKV SNKALPAPIE KTISKAK (SEQ ID NO: 108).
- the CH2 domain is unique in that it is not closely paired with another domain. Rather, two N-linked branched carbohydrate chains are interposed between the two CH2 domains of an intact native Fc-region.
- carbohydrate may provide a substitute for the domaindomain pairing and help stabilize the CH2 domain.
- a carbohydrate chain is attached to the CH2 domain.
- the CH2 domain herein may be a native sequence CH2 domain or variant CH2 domain.
- the “CH3 domain” comprises the stretch of residues C-terminal to a CH2 domain in an Fc region denotes the part of an antibody heavy chain polypeptide that extends approximately from EU position 341 to EU position 446 (EU numbering system according to Kabat).
- the CH3 domain has the amino acid sequence of GQPREPQVYT LPPSRDELTK NQVSLTCLVK GFYPSDIAVE WESNGQPENN YKTTPPVLDS DGSFFLYSKL TVDKSRWQQG NVFSCSVMHE ALHNHYTQKS LSLSPGK (SEQ ID NO: 109).
- the CH3 region herein may be a native sequence CH3 domain or a variant CH3 domain (e.g.
- CH3 domain with an introduced “protuberance” (“knob”) in one chain thereof and a corresponding introduced “cavity” (“hole”) in the other chain thereof; see US Patent No. 5,821,333, expressly incorporated herein by reference).
- Such variant CH3 domains may be used to promote heterodimerization of two non-identical antibody heavy chains as herein described.
- a human IgG heavy chain Fc region extends from Cys226, or from Pro230, to the carboxyl -terminus of the heavy chain.
- the C-terminal lysine (Lys447) of the Fc region may or may not be present.
- EU numbering system also called the EU index, as described in Kabat et al., Sequences of Proteins of Immunological Interest, 5th Ed. Public Health Service, National Institutes of Health, Bethesda, MD, 1991.
- the “knob-into-hole” technology is described e.g. in US 5,731,168; US 7,695,936; Ridgway et al., Prot Eng 9, 617-621 (1996) and Carter, J Immunol Meth 248, 7-15 (2001).
- the method involves introducing a protuberance (“knob”) at the interface of a first polypeptide and a corresponding cavity (“hole”) in the interface of a second polypeptide, such that the protuberance can be positioned in the cavity so as to promote heterodimer formation and hinder homodimer formation.
- Protuberances are constructed by replacing small amino acid side chains from the interface of the first polypeptide with larger side chains (e.g. tyrosine or tryptophan).
- Compensatory cavities of identical or similar size to the protuberances are created in the interface of the second polypeptide by replacing large amino acid side chains with smaller ones (e.g. alanine or threonine).
- the protuberance and cavity can be made by altering the nucleic acid encoding the polypeptides, e.g. by site-specific mutagenesis, or by peptide synthesis.
- a knob modification comprises the amino acid substitution T366W in one of the two subunits of the Fc domain
- the hole modification comprises the amino acid substitutions T366S, L368A and Y407V in the other one of the two subunits of the Fc domain.
- the subunit of the Fc domain comprising the knob modification additionally comprises the amino acid substitution S354C
- the subunit of the Fc domain comprising the hole modification additionally comprises the amino acid substitution Y349C.
- a "region equivalent to the Fc region of an immunoglobulin" is intended to include naturally occurring allelic variants of the Fc region of an immunoglobulin as well as variants having alterations which produce substitutions, additions, or deletions but which do not decrease substantially the ability of the immunoglobulin to mediate effector functions (such as antibodydependent cellular cytotoxicity).
- one or more amino acids can be deleted from the N-terminus or C-terminus of the Fc region of an immunoglobulin without substantial loss of biological function.
- Such variants can be selected according to general rules known in the art so as to have minimal effect on activity (see, e.g., Bowie, J. U. et al., Science 247: 1306-10 (1990)).
- wild-type Fc domain denotes an amino acid sequence identical to the amino acid sequence of an Fc domain found in nature.
- Wild-type human Fc domains include a native human IgGl Fc-region (non- A and A allotypes), native human IgG2 Fc-region, native human IgG3 Fc-region, and native human IgG4 Fc-region as well as naturally occurring variants thereof.
- Wild-type Fc-regions are denoted in SEQ ID NO: 110 (IgGl, Caucasian allotype), SEQ ID NO: 111 (IgGl, afroamerican allotype), SEQ ID NO: 112 (IgG2), SEQ ID NO: 113 (IgG3) and SEQ ID NO: 114 (IgG4).
- variant (human) Fc domain denotes an amino acid sequence which differs from that of a “wild-type” (human) Fc domain amino acid sequence by virtue of at least one “amino acid mutation”.
- the variant Fc-region has at least one amino acid mutation compared to a native Fc-region, e.g. from about one to about ten amino acid mutations, and in one aspect from about one to about five amino acid mutations in a native Fc-region.
- the (variant) Fc-region has at least about 95 % homology with a wild-type Fc-region.
- a specific variant Fc domain disclosed herein is the human IgGl heavy chain constant region with mutations L234A, L235A and P329G comprising the amino acid sequence of SEQ ID NO: 115.
- effector functions refers to those biological activities attributable to the Fc region of an antibody, which vary with the antibody isotype.
- antibody effector functions include: Clq binding and complement dependent cytotoxicity (CDC), Fc receptor binding, antibody-dependent cell-mediated cytotoxicity (ADCC), antibody-dependent cellular phagocytosis (ADCP), cytokine secretion, immune complex -mediated antigen uptake by antigen presenting cells, down regulation of cell surface receptors (e.g. B cell receptor), and B cell activation.
- Fc receptor binding dependent effector functions can be mediated by the interaction of the Fc-region of an antibody with Fc receptors (FcRs), which are specialized cell surface receptors on hematopoietic cells.
- Fc receptors belong to the immunoglobulin superfamily, and have been shown to mediate both the removal of antibody-coated pathogens by phagocytosis of immune complexes, and the lysis of erythrocytes and various other cellular targets (e.g. tumor cells) coated with the corresponding antibody, via antibody dependent cell mediated cytotoxicity (ADCC) (see e.g. Van de Winkel, J.G. and Anderson, C.L., J. Leukoc. Biol. 49 (1991) 511-524).
- ADCC antibody dependent cell mediated cytotoxicity
- FcRs are defined by their specificity for immunoglobulin isotypes: Fc receptors for IgG antibodies are referred to as FcyR. Fc receptor binding is described e.g. in Ravetch, J.V. and Kinet, J.P., Annu. Rev. Immunol. 9 (1991) 457-492; Capel, P.J., et al., Immunomethods 4 (1994) 25-34; de Haas, M., et al., J. Lab. Clin. Med. 126 (1995) 330-341; and Gessner, J.E., et al., Ann. Hematol. 76 (1998) 231-248.
- FcyR Fc-region of IgG antibodies
- FcyRI binds monomeric IgG with high affinity and is expressed on macrophages, monocytes, neutrophils and eosinophils.
- Modification in the Fc-region IgG at least at one of the amino acid residues E233-G236, P238, D265, N297, A327 and P329 (numbering according to EU index of Kabat) reduce binding to FcyRI.
- FcyRIIA is found on many cells involved in killing (e.g. macrophages, monocytes, neutrophils) and seems able to activate the killing process.
- FcyRIIB seems to play a role in inhibitory processes and is found on B cells, macrophages and on mast cells and eosinophils. On B-cells it seems to function to suppress further immunoglobulin production and isotype switching to, for example, the IgE class.
- FcyRIIB acts to inhibit phagocytosis as mediated through FcyRIIA.
- the B-form may help to suppress activation of these cells through IgE binding to its separate receptor.
- Reduced binding for FcyRIIA is found e.g. for antibodies comprising an IgG Fc-region with mutations at least at one of the amino acid residues E233- G236, P238, D265, N297, A327, P329, D270, Q295, A327, R292, and K414 (numbering according to EU index of Kabat).
- FcyRIII (CD 16) binds IgG with medium to low affinity and exists as two types.
- FcyRIIIA is found on NK cells, macrophages, eosinophils and some monocytes and T cells and mediates ADCC.
- FcyRIIIB is highly expressed on neutrophils. Reduced binding to FcyRIIIA is found e.g.
- antibodies comprising an IgG Fc-region with mutation at least at one of the amino acid residues E233-G236, P238, D265, N297, A327, P329, D270, Q295, A327, S239, E269, E293, Y296, V303, A327, K338 and D376 (numbering according to EU index of Kabat).
- ADCC antibody-dependent cellular cytotoxicity
- the term “reduced ADCC” is defined as either a reduction in the number of target cells that are lysed in a given time, at a given concentration of antibody in the medium surrounding the target cells, by the mechanism of ADCC defined above, and/or an increase in the concentration of antibody in the medium surrounding the target cells, required to achieve the lysis of a given number of target cells in a given time, by the mechanism of ADCC.
- the reduction in ADCC is relative to the ADCC mediated by the same antibody produced by the same type of host cells, using the same standard production, purification, formulation and storage methods (which are known to those skilled in the art), but that has not been engineered.
- the reduction in ADCC mediated by an antibody comprising in its Fc domain an amino acid substitution that reduces ADCC is relative to the ADCC mediated by the same antibody without this amino acid substitution in the Fc domain.
- Suitable assays to measure ADCC are well known in the art (see e.g. PCT publication no. WO 2006/082515 or PCT publication no. WO 2012/130831).
- the capacity of the antibody to induce the initial steps mediating ADCC is investigated by measuring their binding to Fey receptors expressing cells, such as cells, recombinantly expressing FcyRI and/or FcyRIIA or NK cells (expressing essentially FcyRIIIA). In particular, binding to FcyR on NK cells is measured.
- an “activating Fc receptor” is an Fc receptor that following engagement by an Fc region of an antibody elicits signaling events that stimulate the receptor-bearing cell to perform effector functions.
- Activating Fc receptors include FcyRIIIa (CD16a), FcyRI (CD64), FcyRIIa (CD32), and FcaRI (CD89).
- a particular activating Fc receptor is human FcyRIIIa (SEQ ID NO: 116, UniProt accession no. P08637, version 141).
- Ectodomain is the domain of a membrane protein that extends into the extracellular space (i.e. the space outside the target cell). Ectodomains are usually the parts of proteins that initiate contact with surfaces, which leads to signal transduction.
- peptide linker refers to a peptide comprising one or more amino acids, typically about 2 to 20 amino acids.
- Peptide linkers are known in the art or are described herein.
- Suitable, non-immunogenic linker peptides are, for example, (G4S) n , (SG4)n or G4(SG4) n peptide linkers, wherein “n” is generally a number between 1 and 5, typically between 2 and 4, in particular 2, i.e.
- GGGGS GGGGSGGGG (SEQ ID NO: 198), GGGGSGGGGS (SEQ ID NO: 118), SGGGGSGGGG (SEQ ID NO: 119) and GGGGS GGGGSGGGG (SEQ ID NO: 120), but also include the sequences GSPGSSSSGS (SEQ ID NO: 121), (G4S) 3 (SEQ ID NO: 122), (G4S) 4 (SEQ ID NO: 123), GSGSGSGS (SEQ ID NO: 124), GSGSGNGS (SEQ ID NO: 125), GGSGSGSG (SEQ ID NO: 126), GGSGSG (SEQ ID NO: 127), GGSG (SEQ ID NO: 128), GGSGNGSG (SEQ ID NO:129), GGNGSGSG (SEQ ID NO: 130) and GGNGSG (SEQ ID NO:131).
- Peptide linkers of particular interest are (G4S) (SEQ ID NO: 117), GGGGSGGGG (SEQ ID NO: 198), (G 4 S) 2 or GGGGSGGGGS (SEQ ID NO: 118), (G4S) 3 (SEQ ID NO: 122) and (G4S) 4 (SEQ ID NO: 123).
- amino acid denotes the group of naturally occurring carboxy a-amino acids comprising alanine (three letter code: ala, one letter code: A), arginine (arg, R), asparagine (asn, N), aspartic acid (asp, D), cysteine (cys, C), glutamine (gin, Q), glutamic acid (glu, E), glycine (gly, G), histidine (his, H), isoleucine (ile, I), leucine (leu, L), lysine (lys, K), methionine (met, M), phenylalanine (phe, F), proline (pro, P), serine (ser, S), threonine (thr, T), tryptophan (trp, W), tyrosine (tyr, Y), and valine (val, V).
- alanine three letter code: ala, one letter code: A
- arginine arg, R
- fused or “connected” is meant that the components (e.g. a polypeptide and another polypeptide) are linked by peptide bonds, either directly or via one or more peptide linkers.
- Percent (%) amino acid sequence identity with respect to a reference polypeptide (protein) sequence is defined as the percentage of amino acid residues in a candidate sequence that are identical with the amino acid residues in the reference polypeptide sequence, after aligning the sequences and introducing gaps, if necessary, to achieve the maximum percent sequence identity, and not considering any conservative substitutions as part of the sequence identity. Alignment for purposes of determining percent amino acid sequence identity can be achieved in various ways that are within the skill in the art, for instance, using publicly available computer software such as BLAST, BLAST-2, ALIGN. SAWI or Megalign (DNASTAR) software. Those skilled in the art can determine appropriate parameters for aligning sequences, including any algorithms needed to achieve maximal alignment over the full length of the sequences being compared.
- % amino acid sequence identity values are generated using the sequence comparison computer program ALIGN-2.
- the ALIGN- 2 sequence comparison computer program was authored by Genentech, Inc., and the source code has been filed with user documentation in the U.S. Copyright Office, Washington D.C., 20559, where it is registered under U.S. Copyright Registration No. TXU510087.
- the ALIGN-2 program is publicly available from Genentech, Inc., South San Francisco, California, or may be compiled from the source code.
- the ALIGN-2 program should be compiled for use on a UNIX operating system, including digital UNIX V4.0D. All sequence comparison parameters are set by the ALIGN-2 program and do not vary.
- the % amino acid sequence identity of a given amino acid sequence A to, with, or against a given amino acid sequence B is calculated as follows: 100 times the fraction X/Y, where X is the number of amino acid residues scored as identical matches by the sequence alignment program ALIGN-2 in that program’s alignment of A and B, and where Y is the total number of amino acid residues in B.
- amino acid sequence variants of the BCMA antibodies or bispecific BCMA antibodies provided herein are contemplated. For example, it may be desirable to improve the binding affinity and/or other biological properties of the BCMA antibodies or bispecific BCMA antibodies.
- Amino acid sequence variants of the BCMA antibodies or bispecific BCMA antibodies may be prepared by introducing appropriate modifications into the nucleotide sequence encoding the molecules, or by peptide synthesis. Such modifications include, for example, deletions from, and/or insertions into and/or substitutions of residues within the amino acid sequences of the antibody. Any combination of deletion, insertion, and substitution can be made to arrive at the final construct, provided that the final construct possesses the desired characteristics, e.g., antigen-binding.
- Sites of interest for substitutional mutagenesis include the CDRs and Framework (FRs). Conservative substitutions are provided in Table B under the heading “Preferred Substitutions” and further described below in reference to amino acid side chain classes (1) to (6). Amino acid substitutions may be introduced into the molecule of interest and the products screened for a desired activity, e.g., retained/improved antigen binding, decreased immunogenicity, or improved ADCC or CDC.
- Amino acids may be grouped according to common side-chain properties:
- Non-conservative substitutions will entail exchanging a member of one of these classes for another class.
- amino acid sequence variants includes substantial variants wherein there are amino acid substitutions in one or more hypervariable region residues of a parent antigen binding molecule (e.g. a humanized or human antibody).
- a parent antigen binding molecule e.g. a humanized or human antibody.
- the resulting variant(s) selected for further study will have modifications (e.g., improvements) in certain biological properties (e.g., increased affinity, reduced immunogenicity) relative to the parent antigen binding molecule and/or will have substantially retained certain biological properties of the parent antigen binding molecule.
- An exemplary substitutional variant is an affinity matured antibody, which may be conveniently generated, e.g., using phage display-based affinity maturation techniques such as those described herein.
- one or more CDR residues are mutated and the variant antigen binding molecules displayed on phage and screened for a particular biological activity (e.g. binding affinity).
- substitutions, insertions, or deletions may occur within one or more CDRs so long as such alterations do not substantially reduce the ability of the antigen binding molecule to bind antigen.
- conservative alterations e.g., conservative substitutions as provided herein
- a useful method for identification of residues or regions of an antibody that may be targeted for mutagenesis is called "alanine scanning mutagenesis" as described by Cunningham and Wells (1989) Science, 244: 1081-1085.
- a residue or group of target residues e.g., charged residues such as Arg, Asp, His, Lys, and Glu
- a neutral or negatively charged amino acid e.g., alanine or polyalanine
- Further substitutions may be introduced at the amino acid locations demonstrating functional sensitivity to the initial substitutions.
- a crystal structure of an antigen-antigen binding molecule complex to identify contact points between the antibody and antigen. Such contact residues and neighboring residues may be targeted or eliminated as candidates for substitution.
- Variants may be screened to determine whether they contain the desired properties.
- Amino acid sequence insertions include amino- and/or carboxyl-terminal fusions ranging in length from one residue to polypeptides containing a hundred or more residues, as well as intrasequence insertions of single or multiple amino acid residues.
- Examples of insertions include BCMA antibodies or bispecific BCMA antibodies with a fusion to the N- or C-terminus to a polypeptide which increases the serum half-life of the CD28 antigen binding molecules.
- the BCMA antibodies or bispecific BCMA antibodies provided herein are altered to increase or decrease the extent to which the antibody is glycosylated.
- Glycosylation variants of the molecules may be conveniently obtained by altering the amino acid sequence such that one or more glycosylation sites is created or removed.
- the agonistic ICOS- binding molecule comprises an Fc domain
- the carbohydrate attached thereto may be altered.
- Native antibodies produced by mammalian cells typically comprise a branched, biantennary oligosaccharide that is generally attached by an N-linkage to Asn297 of the CH2 domain of the Fc region. See, e.g., Wright et al. TIBTECH 15:26-32 (1997).
- the oligosaccharide may include various carbohydrates, e.g., mannose, N-acetyl glucosamine (GlcNAc), galactose, and sialic acid, as well as a fucose attached to a GlcNAc in the “stem” of the biantennary oligosaccharide structure.
- modifications of the oligosaccharide in BCMA antibodies or bispecific BCMA antibodies may be made in order to create variants with certain improved properties.
- variants of BCMA antibodies or bispecific BCMA antibodies are provided having a carbohydrate structure that lacks fucose attached (directly or indirectly) to an Fc region.
- Such fucosylation variants may have improved ADCC function, see e.g. US Patent Publication Nos. US 2003/0157108 (Presta, L.) or US 2004/0093621 (Kyowa Hakko Kogyo Co., Ltd).
- Further variants of the BCMA antibodies or bispecific BCMA antibodies include those with bisected oligosaccharides, e.g., in which a biantennary oligosaccharide attached to the Fc region is bisected by GlcNAc.
- Such variants may have reduced fucosylation and/or improved ADCC function., see for example WO 2003/011878 (Jean-Mairet et al.); US Patent No.
- cysteine engineered variants of the CD28 antigen binding molecules of the invention e.g., “thioMAbs,” in which one or more residues of the molecule are substituted with cysteine residues.
- the substituted residues occur at accessible sites of the molecule.
- reactive thiol groups are thereby positioned at accessible sites of the antibody and may be used to conjugate the antibody to other moieties, such as drug moieties or linker-drug moieties, to create an immunoconjugate.
- any one or more of the following residues may be substituted with cysteine: V205 (Kabat numbering) of the light chain; Al 18 (EU numbering) of the heavy chain; and S400 (EU numbering) of the heavy chain Fc region.
- Cysteine engineered antigen binding molecules may be generated as described, e.g., in U.S. Patent No. 7,521,541.
- the BCMA antibodies or bispecific BCMA antibodies provided herein may be further modified to contain additional non-proteinaceous moieties that are known in the art and readily available.
- the moieties suitable for derivatization of the antibody include but are not limited to water soluble polymers.
- Non-limiting examples of water soluble polymers include, but are not limited to, polyethylene glycol (PEG), copolymers of ethylene glycol/propylene glycol, carboxymethylcellulose, dextran, polyvinyl alcohol, polyvinyl pyrrolidone, poly-1, 3- dioxolane, poly-1, 3, 6-trioxane, ethylene/maleic anhydride copolymer, polyaminoacids (either homopolymers or random copolymers), and dextran or poly(n-vinyl pyrrolidone)polyethylene glycol, propropylene glycol homopolymers, prolypropylene oxide/ethylene oxide co-polymers, polyoxyethylated polyols (e.g., glycerol), polyvinyl alcohol, and mixtures thereof.
- PEG polyethylene glycol
- copolymers of ethylene glycol/propylene glycol carboxymethylcellulose
- dextran polyvinyl alcohol
- Polyethylene glycol propionaldehyde may have advantages in manufacturing due to its stability in water.
- the polymer may be of any molecular weight, and may be branched or unbranched.
- the number of polymers attached to the antibody may vary, and if more than one polymer is attached, they can be the same or different molecules. In general, the number and/or type of polymers used for derivatization can be determined based on considerations including, but not limited to, the particular properties or functions of the antibody to be improved, whether the bispecific antibody derivative will be used in a therapy under defined conditions, etc.
- conjugates of an antibody and non-proteinaceous moiety that may be selectively heated by exposure to radiation are provided.
- the non-proteinaceous moiety is a carbon nanotube (Kam, N.W. et al., Proc. Natl. Acad. Sci. USA 102 (2005) 11600-11605).
- the radiation may be of any wavelength, and includes, but is not limited to, wavelengths that do not harm ordinary cells, but which heat the non-proteinaceous moiety to a temperature at which cells proximal to the antibody-non-proteinaceous moiety are killed.
- immunoconjugates of the BCMA antibodies or bispecific BCMA antibodies provided herein may be obtained.
- An “immunoconjugate” is an antibody conjugated to one or more heterologous molecule(s), for instance a small molecule agent.
- polynucleotide refers to an isolated nucleic acid molecule or construct, e.g. messenger RNA (mRNA), virally-derived RNA, or plasmid DNA (pDNA).
- a polynucleotide may comprise a conventional phosphodiester bond or a non-conventional bond (e.g. an amide bond, such as found in peptide nucleic acids (PNA).
- PNA peptide nucleic acids
- nucleic acid molecule refers to any one or more nucleic acid segments, e.g. DNA or RNA fragments, present in a polynucleotide. Each nucleotide is composed of a base, specifically a purine- or pyrimidine base (i.e.
- nucleic acid molecule is described by the sequence of bases, whereby said bases represent the primary structure (linear structure) of a nucleic acid molecule.
- sequence of bases is typically represented from 5’ to 3’.
- nucleic acid molecule encompasses deoxyribonucleic acid (DNA) including e.g., complementary DNA (cDNA) and genomic DNA, ribonucleic acid (RNA), in particular messenger RNA (mRNA), synthetic forms of DNA or RNA, and mixed polymers comprising two or more of these molecules.
- DNA deoxyribonucleic acid
- cDNA complementary DNA
- RNA ribonucleic acid
- mRNA messenger RNA
- the nucleic acid molecule may be linear or circular.
- nucleic acid molecule includes both, sense and antisense strands, as well as single stranded and double stranded forms.
- the herein described nucleic acid molecule can contain naturally occurring or non-naturally occurring nucleotides.
- nucleic acid molecules also encompass DNA and RNA molecules which are suitable as a vector for direct expression of an antibody of the invention in vitro and/or in vivo, e.g., in a host or patient.
- DNA e.g., cDNA
- RNA e.g., mRNA
- mRNA can be chemically modified to enhance the stability of the RNA vector and/or expression of the encoded molecule so that mRNA can be injected into a subject to generate the antibody in vivo (see e.g., Stadler et al. (2017) Nature Medicine 23:815-817, or EP 2 101 823 Bl).
- isolated nucleic acid molecule or polynucleotide is meant a nucleic acid molecule, DNA or RNA, which has been removed from its native environment.
- a recombinant polynucleotide encoding a polypeptide contained in a vector is considered isolated for the purposes of the present invention.
- Further examples of an isolated polynucleotide include recombinant polynucleotides maintained in heterologous host cells or purified (partially or substantially) polynucleotides in solution.
- An isolated polynucleotide includes a polynucleotide molecule contained in cells that ordinarily contain the polynucleotide molecule, but the polynucleotide molecule is present extrachromosomally or at a chromosomal location that is different from its natural chromosomal location.
- Isolated RNA molecules include in vivo or in vitro RNA transcripts of the present invention, as well as positive and negative strand forms, and double-stranded forms. Isolated polynucleotides or nucleic acids according to the present invention further include such molecules produced synthetically.
- a polynucleotide or a nucleic acid may be or may include a regulatory element such as a promoter, ribosome binding site, or a transcription terminator.
- nucleic acid or polynucleotide having a nucleotide sequence at least, for example, 95% "identical" to a reference nucleotide sequence of the present invention it is intended that the nucleotide sequence of the polynucleotide is identical to the reference sequence except that the polynucleotide sequence may include up to five point mutations per each 100 nucleotides of the reference nucleotide sequence.
- a polynucleotide having a nucleotide sequence at least 95% identical to a reference nucleotide sequence up to 5% of the nucleotides in the reference sequence may be deleted or substituted with another nucleotide, or a number of nucleotides up to 5% of the total nucleotides in the reference sequence may be inserted into the reference sequence.
- These alterations of the reference sequence may occur at the 5’ or 3’ terminal positions of the reference nucleotide sequence or anywhere between those terminal positions, interspersed either individually among residues in the reference sequence or in one or more contiguous groups within the reference sequence.
- any particular polynucleotide sequence is at least 80%, 85%, 90%, 95%, 96%, 97%, 98% or 99% identical to a nucleotide sequence of the present invention can be determined conventionally using known computer programs, such as the ones discussed above for polypeptides (e.g. ALIGN-2).
- expression cassette refers to a polynucleotide generated recombinantly or synthetically, with a series of specified nucleic acid elements that permit transcription of a particular nucleic acid in a target cell.
- the recombinant expression cassette can be incorporated into a plasmid, chromosome, mitochondrial DNA, plastid DNA, virus, or nucleic acid fragment.
- the recombinant expression cassette portion of an expression vector includes, among other sequences, a nucleic acid sequence to be transcribed and a promoter.
- the expression cassette of the invention comprises polynucleotide sequences that encode bispecific antigen binding molecules of the invention or fragments thereof.
- vector or "expression vector” is synonymous with "expression construct” and refers to a DNA molecule that is used to introduce and direct the expression of a specific gene to which it is operably associated in a target cell.
- the term includes the vector as a self-replicating nucleic acid structure as well as the vector incorporated into the genome of a host cell into which it has been introduced.
- the expression vector of the present invention comprises an expression cassette. Expression vectors allow transcription of large amounts of stable mRNA. Once the expression vector is inside the target cell, the ribonucleic acid molecule or protein that is encoded by the gene is produced by the cellular transcription and/or translation machinery.
- the expression vector of the invention comprises an expression cassette that comprises polynucleotide sequences that encode bispecific antigen binding molecules of the invention or fragments thereof.
- host cell refers to cells into which exogenous nucleic acid has been introduced, including the progeny of such cells.
- Host cells include “transformants” and “transformed cells,” which include the primary transformed cell and progeny derived therefrom without regard to the number of passages. Progeny may not be completely identical in nucleic acid content to a parent cell, but may contain mutations. Mutant progeny that have the same function or biological activity as screened or selected for in the originally transformed cell are included herein.
- a host cell is any type of cellular system that can be used to generate the bispecific antigen binding molecules of the present invention.
- Host cells include cultured cells, e.g.
- mammalian cultured cells such as CHO cells, BHK cells, NSO cells, SP2/0 cells, YO myeloma cells, P3X63 mouse myeloma cells, PER cells, PER.C6 cells or hybridoma cells, yeast cells, insect cells, and plant cells, to name only a few, but also cells comprised within a transgenic animal, transgenic plant or cultured plant or animal tissue.
- an “effective amount” of an agent refers to the amount that is necessary to result in a physiological change in the cell or tissue to which it is administered.
- a “therapeutically effective amount” of an agent refers to an amount effective, at dosages and for periods of time necessary, to achieve the desired therapeutic or prophylactic result.
- a therapeutically effective amount of an agent for example eliminates, decreases, delays, minimizes or prevents adverse effects of a disease.
- mammals include, but are not limited to, domesticated animals (e.g. cows, sheep, cats, dogs, and horses), primates (e.g. humans and nonhuman primates such as monkeys), rabbits, and rodents (e.g. mice and rats). Particularly, the individual or subject is a human.
- domesticated animals e.g. cows, sheep, cats, dogs, and horses
- primates e.g. humans and nonhuman primates such as monkeys
- rabbits e.g. mice and rats
- rodents e.g. mice and rats
- composition refers to a preparation which is in such form as to permit the biological activity of an active ingredient contained therein to be effective, and which contains no additional components which are unacceptably toxic to a subject to which the formulation would be administered.
- a “pharmaceutically acceptable excipient” refers to an ingredient in a pharmaceutical composition, other than an active ingredient, which is nontoxic to a subject.
- a pharmaceutically acceptable excipient includes, but is not limited to, a buffer, a stabilizer, or a preservative.
- package insert is used to refer to instructions customarily included in commercial packages of therapeutic products, that contain information about the indications, usage, dosage, administration, combination therapy, contraindications and/or warnings concerning the use of such therapeutic products.
- treatment refers to clinical intervention in an attempt to alter the natural course of the individual being treated, and can be performed either for prophylaxis or during the course of clinical pathology. Desirable effects of treatment include, but are not limited to, preventing occurrence or recurrence of disease, alleviation of symptoms, diminishment of any direct or indirect pathological consequences of the disease, preventing metastasis, decreasing the rate of disease progression, amelioration or palliation of the disease state, and remission or improved prognosis.
- the molecules of the invention are used to delay development of a disease or to slow the progression of a disease.
- the term “therapy” refers to any protocol, method and/or agent that can be used in the prevention, management, treatment and/or amelioration of a disease (or symptom related thereto) or cancer.
- the terms “therapies” and “therapy” refer to a biological therapy, supportive therapy, and/or other therapies useful in the prevention, management, treatment and/or amelioration of a disease or cancer known to one of skill in the art such as a physician.
- combination treatment or “co-administration” as noted herein encompasses combined administration (where two or more therapeutic agents are included in the same or separate formulations), and separate administration, in which case, administration of an antibody as reported herein can occur prior to, simultaneously, and/or following, administration of the additional therapeutic agent or agents, preferably an antibody or antibodies.
- cancer refers to or describes the physiological condition in mammals that is typically characterized by unregulated cell growth/proliferation.
- cancer refers to proliferative diseases, such as carcinoma, lymphomas (e.g., Hodgkin’s and nonHodgkin’s lymphoma), blastoma, sarcoma, and leukemia.
- cancer includes lymphocytic leukemias, lung cancer, non-small cell lung (NSCL) cancer, bronchi oloalviolar cell lung cancer, bone cancer, pancreatic cancer, skin cancer, cancer of the head or neck, cutaneous or intraocular melanoma, uterine cancer, ovarian cancer, rectal cancer, cancer of the anal region, stomach cancer, gastric cancer, colon cancer, breast cancer, uterine cancer, carcinoma of the fallopian tubes, carcinoma of the endometrium, carcinoma of the cervix, carcinoma of the vagina, carcinoma of the vulva, Hodgkin's Disease, cancer of the esophagus, cancer of the small intestine, cancer of the endocrine system, cancer of the thyroid gland, cancer of the parathyroid gland, cancer of the adrenal gland, sarcoma of soft tissue, cancer of the urethra, cancer of the penis, prostate cancer, cancer of the bladder, cancer of the kidney or ureter, renal cell carcinoma, carcinoma of the renal
- the cancer is a solid tumor.
- the cancer is a haematological cancer, particularly leukemia, most particularly acute lymphoblastic leukemia (ALL) or acute myelogenous leukemia (AML).
- ALL acute lymphoblastic leukemia
- AML acute myelogenous leukemia
- the term cancer refers to any canceer in which BCMA is expressed. More preferably, the cancer is multiple myeloma (MM).
- novel antibodies and/or antibody fragments that specifically bind to B cell maturation antigen (BCMA).
- novel antibodies and/or antibody fragments that specifically bind to the extracellular domain of human BCMA comprising the amino acid sequence of SEQ ID NO: 132.
- SEQ ID NO: 132 the amino acid sequence of SEQ ID NO: 132.
- these antibodies specifically bind to human BCMA.
- These antibodies are able to bind to human BCMA and to cynomolgus BCMA.
- the new antibodies are also able to bind to mutated human BCMA variants hu BCMA R27P (SEQ ID NO:210), Hu BCMA_S30del (SEQ ID NO:211), hu BCMA P33S (SEQ ID NO:212) and hu BCMA_P34del (SEQ ID NO:213). They are therefore not affected by point mutations in the ECD of BCMA and do not lose the therapeutic activity as has been observed with other BCMA-targeted molecules such as Elranatamab and Teclistamab ((Lee et al., Nature Medicine 2023, 29, 2295-2306).
- the new antibodies are further characterized in that they are producable in high amounts and with high titers, that they show high thermal stability (as measured by the aggregation temperature T agg ), or in that they possess a high degree of humanness and may therefore be less immunogenic in the human body.
- the percentage of humanness of the VH and VL sequences as compared to the human germline sequences can be determined by the methods described in Abhinandan, K. R. and Martin, Andrew C. R. 2007, J. Mol. Biol. 2007, 369, 852-862.
- VH BCMA heavy chain variable region
- VH BCMA heavy chain variable region comprising a heavy chain complementary determining region CDR-H1 of SEQ ID NO: 1 (GYTFTNYWMH), a CDR-H2 of SEQ ID NO: 2 (IIHPNSGSTNYNEKFQG), and a CDR-H3 of SEQ ID NO: 3 (GIYDYPFAY), and
- VLBCMA light chain variable region
- VL comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 4 (RASES VSIHGTHLMH), a CDR-L2 of SEQ ID NO: 5 (AASSLQS) and a CDR-L3 of SEQ ID NO: 6 (QQSIEDPYT); or
- VL comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 4 (RASES VSIHGTHLMH), a CDR-L2 of SEQ ID NO: 7 (AASNLES) and a CDR-L3 of SEQ ID NO: 6 (QQSIEDPYT), or
- VL comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 4 (RASES VSIHGTHLMH), a CDR-L2 of SEQ ID NO: 8 (AASNLQS) and a CDR-L3 of SEQ ID NO: 6 (QQSIEDPYT).
- a BCMA antibody (or an antigen binding domain that specifically binds to BCMA), wherein said antibody comprises a VH BCMA comprising an amino acid sequence selected from the group consisting of SEQ ID NO: 9 (VHla) and SEQ ID NO: 10 (VHlb), and/or a VLBCMA comprising an amino acid sequence selected from the group consisting of SEQ ID NO: 11 (VLlf) , SEQ ID NO: 12 (VLla), SEQ ID NO: 13 (VLlb), SEQ ID NO: 14 (VLlc), SEQ ID NO: 15 (VLld), and SEQ ID NO: 16 (VLle).
- a BCMA antibody (or an antigen binding domain that specifically binds to BCMA), wherein said antibody (or antigen binding domain) comprises
- VH BCMA comprising an amino acid sequence of SEQ ID NO:9
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 11, or
- VH BCMA comprising an amino acid sequence of SEQ ID NOV and VLBCMA comprising an amino acid sequence of SEQ ID NO: 12;
- VH BCMA comprising an amino acid sequence of SEQ ID NOV
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 13
- VH BCMA comprising an amino acid sequence of SEQ ID NOV and VLBCMA comprising an amino acid sequence of SEQ ID NO: 14;
- VH BCMA comprising an amino acid sequence of SEQ ID NOV
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 15, or
- VH BCMA comprising an amino acid sequence of SEQ ID NOV
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 16
- VH BCMA comprising an amino acid sequence of SEQ ID NOV
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 135, or
- VH BCMA comprising an amino acid sequence of SEQ ID NOV and VLBCMA comprising an amino acid sequence of SEQ ID NO: 136;
- VH BCMA comprising an amino acid sequence of SEQ ID NO: 10
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 11
- VH BCMA comprising an amino acid sequence of SEQ ID NO: 10 and VLBCMA comprising an amino acid sequence of SEQ ID NO: 12;
- VH BCMA comprising an amino acid sequence of SEQ ID NO: 10
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 13, or
- VH BCMA comprising an amino acid sequence of SEQ ID NO: 10
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 14
- VH BCMA comprising an amino acid sequence of SEQ ID NO: 10
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 15, or
- VH BCMA comprising an amino acid sequence of SEQ ID NO: 10 and VLBCMA comprising an amino acid sequence of SEQ ID NO: 16;
- VH BCMA comprising an amino acid sequence of SEQ ID NO: 10
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 135, or
- VH BCMA comprising an amino acid sequence of SEQ ID NO: 10
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 136.
- the BCMA antibody (or the antigen binding domain that specifically binds to BCMA) comprises
- VH BCMA comprising an amino acid sequence of SEQ ID NOV and VLBCMA comprising an amino acid sequence of SEQ ID NO: 11, or
- VH BCMA comprising an amino acid sequence of SEQ ID NO:9 and VLBCMA comprising an amino acid sequence of SEQ ID NO: 12; or
- VH BCMA comprising an amino acid sequence of SEQ ID NOV
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 13
- VH BCMA comprising an amino acid sequence of SEQ ID NOV and VLBCMA comprising an amino acid sequence of SEQ ID NO: 14;
- VH BCMA comprising an amino acid sequence of SEQ ID NOV
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 15, or
- VH BCMA comprising an amino acid sequence of SEQ ID NOV
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 16
- VH BCMA comprising an amino acid sequence of SEQ ID NOV
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 135, or
- VH BCMA comprising an amino acid sequence of SEQ ID NOV
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 136.
- the BCMA antibody (or the antigen binding domain that specifically binds to BCMA) comprises
- VH BCMA comprising an amino acid sequence of SEQ ID NO: 10
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 11
- VH BCMA comprising an amino acid sequence of SEQ ID NO: 10 and VLBCMA comprising an amino acid sequence of SEQ ID NO: 12;
- VH BCMA comprising an amino acid sequence of SEQ ID NO: 10
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 13, or
- VH BCMA comprising an amino acid sequence of SEQ ID NO: 10
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 14
- VH BCMA comprising an amino acid sequence of SEQ ID NO: 10
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 15, or
- VH BCMA comprising an amino acid sequence of SEQ ID NO: 10 and VLBCMA comprising an amino acid sequence of SEQ ID NO: 16;
- VH BCMA comprising an amino acid sequence of SEQ ID NO: 10
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 135, or
- VH BCMA comprising an amino acid sequence of SEQ ID NO: 10
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 136.
- BCMA antibody or an antigen binding domain that specifically binds to BCMA, wherein said antibody (or antigen binding domain) comprises
- VH BCMA comprising an amino acid sequence of SEQ ID NOV
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 11
- the BCMA antibody (or the antigen binding domain that specifically binds to BCMA) comprises a VH BCMA comprising an amino acid sequence of SEQ ID NO:9 and a VL BCMA comprising an amino acid sequence of SEQ ID NO: 11.
- VH BCMA heavy chain variable region
- VH comprising a heavy chain complementary determining region CDR-H1 of SEQ ID NO: 29 (GFTFSNAWMD), a CDR-H2 of SEQ ID NO: 30 (QITAKSNNYATYYADSVKG), and a CDR-H3 of SEQ ID NO: 31 (DGYH), and
- VH comprising a heavy chain complementary determining region CDR-H1 of SEQ ID NO: 29 (GFTFSNAWMD), a CDR-H2 of SEQ ID NO: 32 (QITAKSNNYATYYAAPVKG), and a CDR-H3 of SEQ ID NO: 31 (DGYH), and
- VLBCMA light chain variable region
- RASEDIRNGLA light chain complementary determining region CDR-L1 of SEQ ID NO: 33
- NANSLHT NANSLHT
- EDTSKYPYT a light chain variable region comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 33 (RASEDIRNGLA), a CDR-L2 of SEQ ID NO: 34 (NANSLHT) and a CDR-L3 of SEQ ID NO: 35 (EDTSKYPYT).
- VLBCMA light chain variable region comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 33 (RASEDIRNGLA), a CDR-L2 of SEQ ID NO: 34 (NANSLHT) and a CDR-L3 of SEQ ID NO: 35 (EDTSKYPYT).
- a BCMA antibody (or an antigen binding domain that specifically binds to BCMA), wherein said antibody (or antigen binding domain) comprises a heavy chain variable region (VHBCMA) comprising a CDR-H1 of SEQ ID NO: 29 (GFTFSNAWMD), a CDR-H2 of SEQ ID NO: 30 (QITAKSNNYATYYADSVKG), and a CDR-H3 of SEQ ID NO: 31 (DGYH), and a light chain variable region (VLBCMA) comprising a CDR-L1 of SEQ ID NO: 33 (RASEDIRNGLA), a CDR-L2 of SEQ ID NO: 34 (NANSLHT) and a CDR-L3 of SEQ ID NO: 35 (EDTSKYPYT).
- VHBCMA heavy chain variable region
- RASEDIRNGLA CDR-L1 of SEQ ID NO: 33
- NANSLHT NANSLHT
- CDR-L3 of SEQ ID NO: 35 EDTSKY
- a BCMA antibody (or an antigen binding domain that specifically binds to BCMA), wherein said antibody (or antigen binding domain) comprises a heavy chain variable region (VHBCMA) comprising a CDR-H1 of SEQ ID NO: 29 (GFTFSNAWMD), a CDR-H2 of SEQ ID NO: 32 (QITAKSNNYATYYAAPVKG), and a CDR-H3 of SEQ ID NO: 31 (DGYH), and a light chain variable region (VLBCMA) comprising a CDR-L1 of SEQ ID NO: 33 (RASEDIRNGLA), a CDR-L2 of SEQ ID NO: 34 (NANSLHT) and a CDR-L3 of SEQ ID NO: 35 (EDTSKYPYT).
- VHBCMA heavy chain variable region
- RASEDIRNGLA a CDR-L2 of SEQ ID NO: 34
- NANSLHT NANSLHT
- a BCMA antibody (or an antigen binding domain that specifically binds to BCMA), wherein said antibody (or antigen binding domain) comprises a VH BCMA comprising an amino acid sequence selected from the group consisting of SEQ ID NO: 36 (VH2a), SEQ ID NO: 38 (VHlb), SEQ ID NO:48 (VHla), SEQ ID NO:49 (VHla_Y292D), SEQ ID NO:50 (VHlc hu CDR2) and SEQ ID NO:51 (VHla_W197Y), and/or the V L BCMA comprising an amino acid sequence selected from the group consisting of SEQ ID NO: 37 (VL2a), SEQ ID NO:39 (VLla), SEQ ID NO:52 (VLla_L2_GL), SEQ ID NO:53 (VL2a_L2_GL), SEQ ID NO:54 (VLla_Ll_pGL), SEQ ID NO:55 (VLla_N651A), SEQ ID
- a BCMA antibody (or an antigen binding domain that specifically binds to BCMA), wherein said antibody (or antigen binding domain) comprises a VH BCMA comprising an amino acid sequence selected from the group consisting of SEQ ID NO: 36 (VH2a) and SEQ ID NO: 38 (VHlb), and/or the VL BCMA comprising an amino acid sequence selected from the group consisting of SEQ ID NO: 37 (VL2a) and SEQ ID NO:39 (VLla).
- BCMA antibody or an antigen binding domain that specifically binds to BCMA, wherein said antibody (or antigen binding domain) comprises
- VH BCMA comprising an amino acid sequence of SEQ ID NO:36 and a VLBCMA comprising an amino acid sequence of SEQ ID NO:37, or
- VH BCMA comprising an amino acid sequence of SEQ ID NO:38 and a VLBCMA comprising an amino acid sequence of SEQ ID NO:39;
- VH BCMA comprising an amino acid sequence of SEQ ID NO:48 and VLBCMA comprising an amino acid sequence of SEQ ID NO:52, or
- VH BCMA comprising an amino acid sequence of SEQ ID NO:49 and VLBCMA comprising an amino acid sequence of SEQ ID NO:39;
- VH BCMA comprising an amino acid sequence of SEQ ID NO:49 and VLBCMA comprising an amino acid sequence of SEQ ID NO:52, or
- VHBCMA comprising an amino acid sequence of SEQ ID NO:49 and VLBCMA comprising an amino acid sequence of SEQ ID NO:58;
- VH BCMA comprising an amino acid sequence of SEQ ID NO:49 and VLBCMA comprising an amino acid sequence of SEQ ID NO:37, or
- VH BCMA comprising an amino acid sequence of SEQ ID NO:38 and VLBCMA comprising an amino acid sequence of SEQ ID NO:54;
- VH BCMA comprising an amino acid sequence of SEQ ID NO:38 and VLBCMA comprising an amino acid sequence of SEQ ID NO:55, or
- VH BCMA comprising an amino acid sequence of SEQ ID NO:38 and VLBCMA comprising an amino acid sequence of SEQ ID NO:56;
- VH BCMA comprising an amino acid sequence of SEQ ID NO:38 and VLBCMA comprising an amino acid sequence of SEQ ID NO:58, or
- VH BCMA comprising an amino acid sequence of SEQ ID NO:38 and VLBCMA comprising an amino acid sequence of SEQ ID NO:37;
- VH BCMA comprising an amino acid sequence of SEQ ID NO:36 and VLBCMA comprising an amino acid sequence of SEQ ID NO:39, or
- VH BCMA comprising an amino acid sequence of SEQ ID NO:36 and VLBCMA comprising an amino acid sequence of SEQ ID NO:54; or (o) a VH BCMA comprising an amino acid sequence of SEQ ID NO:36 and VLBCMA comprising an amino acid sequence of SEQ ID NO:58.
- BCMA antibody or an antigen binding domain that specifically binds to BCMA, wherein said antibody (or antigen binding domain) comprises
- VH BCMA comprising an amino acid sequence of SEQ ID NO:36 and a VLBCMA comprising an amino acid sequence of SEQ ID NO:37, or
- VH BCMA comprising an amino acid sequence of SEQ ID NO:38 and a VLBCMA comprising an amino acid sequence of SEQ ID NO:39.
- the BCMA antibody comprises a VH BCMA comprising an amino acid sequence of SEQ ID NO:36 and a VLBCMA comprising an amino acid sequence of SEQ ID NO:37.
- the antibody that specifically binds to BCMA is a full-length antibody, in particular of human IgGl subclass. In one particular aspect, it comprises the amino acid mutations L234A, L235A and P329G (numbering according to Kabat EU index).
- Bispecific CD28 agonistic antibodies comprising the new BCMA antibodies
- the patent application also provides new BCMA antibodies that comprise a second antigen binding domain that specifically binds to second antigen, in particular to CD28.
- These bispecific CD28 agonistic antibodies possess advantageous properties such as excellent producibility, stability, binding affinity, biological activity, targeting efficiency, reduced toxicity, an extended dosage range that can be given to a patient and thereby a possibly an enhanced efficacy can be observed.
- the new bispecific CD28 agonistic antibodies comprise an Fc domain composed of a first and a second subunit capable of stable association comprising one or more amino acid substitution that reduces the binding affinity of the antigen binding molecule to an Fc receptor and/or effector function (Fc silent) and thus unspecific cross-linking via Fc receptors is avoided.
- BCMA antigen binding domains capable of specific binding to BCMA which causes cross-linking at the tumor site.
- the inventors have found that based on their binding properties the BCMA antigen binding domains as described herein have advantageous properties that makes them more usable in the bispecific format.
- the bispecific agonistic CD28 antigen binding molecule comprising these BCMA antigen binding domains possess an improved functionality and ability to increase T cell activation, particularly in the presence of T-cell activating anti-CD3 bispecific antibodies. Thus, an enhanced tumor-specific T cell activation is achieved.
- a bispecific antibody that specifically binds to B cell maturation agent (BCMA) and CD28, wherein the antibody comprises
- a first antigen binding domain comprising (i) a heavy chain variable region (VH BCMA) comprising a heavy chain complementary determining region CDR-H1 of SEQ ID NO: 1 (GYTFTNYWMH), a CDR-H2 of SEQ ID NO: 2 (IH4PNSGSTNYNEKFQG), and a CDR-H3 of SEQ ID NO: 3 (GIYDYPFAY), and
- VLBCMA light chain variable region
- VL comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 4 (RASES VSIHGTHLMH), a CDR-L2 of SEQ ID NO: 5 (AASSLQS) and a CDR-L3 of SEQ ID NO: 6 (QQSIEDPYT); or
- VL comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 4 (RASES VSIHGTHLMH), a CDR-L2 of SEQ ID NO: 7 (AASNLES) and a CDR-L3 of SEQ ID NO: 6 (QQSIEDPYT), or
- VL comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 4 (RASES VSIHGTHLMH), a CDR-L2 of SEQ ID NO: 8 (AASNLQS) and a CDR-L3 of SEQ ID NO: 6 (QQSIEDPYT), and
- an antibody that specifically binds to BCMA and CD28, wherein said antibody comprises a VH BCMA comprising an amino acid sequence selected from the group consisting of SEQ ID NO: 9 (VHla) and SEQ ID NO: 10 (VHlb), and/or a VLBCMA comprising an amino acid sequence selected from the group consisting of SEQ ID NO: 11 (VLlf) , SEQ ID NO: 12 (VLla), SEQ ID NO: 13 (VLlb), SEQ ID NO: 14 (VLlc), SEQ ID NO: 15 (VLld), and SEQ ID NO: 16 (VLle).
- VH BCMA comprising an amino acid sequence selected from the group consisting of SEQ ID NO: 9 (VHla) and SEQ ID NO: 10 (VHlb)
- VLBCMA comprising an amino acid sequence selected from the group consisting of SEQ ID NO: 11 (VLlf) , SEQ ID NO: 12 (VLla), SEQ ID NO: 13 (VLl
- an antibody that specifically binds to BCMA and CD28 wherein said antibody comprises
- VH BCMA comprising an amino acid sequence of SEQ ID NOV
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 11, or
- VH BCMA comprising an amino acid sequence of SEQ ID NOV and VLBCMA comprising an amino acid sequence of SEQ ID NO: 12;
- VH BCMA comprising an amino acid sequence of SEQ ID NOV
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 13
- VH BCMA comprising an amino acid sequence of SEQ ID NOV and VLBCMA comprising an amino acid sequence of SEQ ID NO: 14;
- VH BCMA comprising an amino acid sequence of SEQ ID NOV
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 15, or
- VHBCMA comprising an amino acid sequence of SEQ ID NOV
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 16
- VH BCMA comprising an amino acid sequence of SEQ ID NOV
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 135, or
- VH BCMA comprising an amino acid sequence of SEQ ID NOV and VLBCMA comprising an amino acid sequence of SEQ ID NO: 136; or (i) a VH BCMA comprising an amino acid sequence of SEQ ID NO: 10 and VLBCMA comprising an amino acid sequence of SEQ ID NO: 11, or
- VH BCMA comprising an amino acid sequence of SEQ ID NO: 10 and VLBCMA comprising an amino acid sequence of SEQ ID NO: 12;
- VH BCMA comprising an amino acid sequence of SEQ ID NO: 10
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 13, or
- VH BCMA comprising an amino acid sequence of SEQ ID NO: 10
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 14
- VH BCMA comprising an amino acid sequence of SEQ ID NO: 10
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 15, or
- VH BCMA comprising an amino acid sequence of SEQ ID NO: 10 and VLBCMA comprising an amino acid sequence of SEQ ID NO: 16;
- VH BCMA comprising an amino acid sequence of SEQ ID NO: 10
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 135, or
- VH BCMA comprising an amino acid sequence of SEQ ID NO: 10
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 136.
- the antibody that specifically binds to BCMA and CD28 comprises
- VH BCMA comprising an amino acid sequence of SEQ ID NOV
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 11, or
- VH BCMA comprising an amino acid sequence of SEQ ID NOV and VLBCMA comprising an amino acid sequence of SEQ ID NO: 12;
- VH BCMA comprising an amino acid sequence of SEQ ID NOV
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 13
- VH BCMA comprising an amino acid sequence of SEQ ID NOV and VLBCMA comprising an amino acid sequence of SEQ ID NO: 14;
- VH BCMA comprising an amino acid sequence of SEQ ID NOV
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 15, or
- VH BCMA comprising an amino acid sequence of SEQ ID NOV
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 16
- VH BCMA comprising an amino acid sequence of SEQ ID NOV
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 135, or
- VH BCMA comprising an amino acid sequence of SEQ ID NOV
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 136.
- the antibody that specifically binds to BCMA and CD28 comprises
- VH BCMA comprising an amino acid sequence of SEQ ID NO: 10
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 11
- VH BCMA comprising an amino acid sequence of SEQ ID NO: 10 and VLBCMA comprising an amino acid sequence of SEQ ID NO: 12; or (k) a VH BCMA comprising an amino acid sequence of SEQ ID NO: 10 and VLBCMA comprising an amino acid sequence of SEQ ID NO: 13, or
- VH BCMA comprising an amino acid sequence of SEQ ID NO: 10
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 14
- VH BCMA comprising an amino acid sequence of SEQ ID NO: 10
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 15, or
- VH BCMA comprising an amino acid sequence of SEQ ID NO: 10 and VLBCMA comprising an amino acid sequence of SEQ ID NO: 16;
- VH BCMA comprising an amino acid sequence of SEQ ID NO: 10
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 135, or
- VH BCMA comprising an amino acid sequence of SEQ ID NO: 10
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 136.
- an antibody that specifically binds to BCMA and CD28 wherein said antibody comprises
- VH BCMA comprising an amino acid sequence of SEQ ID NOV
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 11
- VH BCMA comprising an amino acid sequence of SEQ ID NOV
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 12.
- the antibody that specifically binds to BCMA and CD28 comprises a VH BCMA comprising an amino acid sequence of SEQ ID NOV and a VL BCMA comprising an amino acid sequence of SEQ ID NO: 11.
- the first antigen binding domain that specifically binds to BCMA can be a Fab molecule.
- the first antigen binding domain that specifically binds to BCMA can be a cross-Fab molecule, i.e a Fab molecule wherein the variable domains VL and VH or the constant domains CL and CHI, particularly the variable domains VL and VH, of the Fab light chain and the Fab heavy chain are replaced by each other.
- the first antigen binding domain that specifically binds to BCMA is a Fab molecule wherein in the constant domain CL the amino acid at position 123 (numbering according to Kabat EU index) is substituted by an amino acid selected from lysine (K), arginine (R) or histidine (H) and the amino acid at position 124 (numbering according to Kabat EU index) is substituted independently by lysine (K), arginine (R) or histidine (H), and wherein in the constant domain CHI the amino acid at position 147 (numbering according to Kabat EU index) is substituted independently by glutamic acid (E) or aspartic acid (D) and the amino acid at position 213 (numbering according to Kabat EU index) is substituted independently by glutamic acid (E), or aspartic acid (D) (numbering according to Kabat EU index).
- the second antigen binding domain that specifically binds to CD28 can be a Fab molecule.
- the second antigen binding domain that specifically binds to CD28 is a cross-Fab molecule, i.e a Fab molecule wherein the variable domains VL and VH or the constant domains CL and CHI, particularly the variable domains VL and VH, of the Fab light chain and the Fab heavy chain are replaced by each other.
- the second antigen binding domain that specifically binds to CD28 comprises a heavy chain variable region (VHCD28) comprising a heavy chain complementary determining region CDR-H1 of SEQ ID NO: 17, a CDR-H2 of SEQ ID NO: 18, and a CDR-H3 of SEQ ID NO: 19, and a light chain variable region (VLCD28) comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 20, a CDR-L2 of SEQ ID NO: 21 and a CDR-L3 of SEQ ID NO: 22.
- VHCD28 heavy chain variable region comprising a heavy chain complementary determining region CDR-H1 of SEQ ID NO: 17, a CDR-H2 of SEQ ID NO: 18, and a CDR-H3 of SEQ ID NO: 19, and a light chain variable region (VLCD28) comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 20, a CDR-L2 of SEQ ID NO: 21 and
- the second antigen binding domain that specifically binds to CD28 comprises a heavy chain variable region (VHCD28) comprising an amino acid sequence of SEQ ID NO:23, and a light chain variable region (VLCD28) comprising an amino acid sequence of SEQ ID NO:24 (v8).
- VHCD28 heavy chain variable region
- VLCD28 light chain variable region
- an antibody that specifically binds to BCMA and CD28, wherein said antibody comprises a first antigen binding domain comprising a VH BCMA comprising an amino acid sequence of SEQ ID NOV and VLBCMA comprising an amino acid sequence of SEQ ID NO: 11 and a second antigen binding domain comprising VH CD28 comprising an amino acid sequence of SEQ ID NO:23 and VL CD28 comprising an amino acid sequence of SEQ ID NO:24.
- the antibody that specifically binds to BCMA and CD28 comprises a polypeptide that is at least 95%, 96%, 97%, 98%, or 99% identical to the sequence of SEQ ID NO: 25, a polypeptide that is at least 95%, 96%, 97%, 98%, or 99% identical to the sequence of SEQ ID NO: 26, a polypeptide that is at least 95%, 96%, 97%, 98%, or 99% identical to the sequence of SEQ ID NO: 27, and a polypeptide that is at least 95%, 96%, 97%, 98%, or 99% identical to the sequence of SEQ ID NO: 28.
- an antibody that specifically binds to BCMA and CD28, wherein said antibody comprises a first light chain comprising the amino acid sequence of SEQ ID NO:25, a first heavy chain comprising the amino acid sequence of SEQ ID NO:26, a second heavy chain comprising the amino acid sequence of SEQ ID NO:27 and a second light chain comprising the amino acid sequence of SEQ ID NO:28.
- an antibody that specifically binds to BCMA and CD28, wherein said antibody comprises a first antigen binding domain comprising a VH BCMA comprising an amino acid sequence of SEQ ID NOV and VLBCMA comprising an amino acid sequence of SEQ ID NO: 12 and a second antigen binding domain comprising VH CD28 comprising an amino acid sequence of SEQ ID NO:23 and VL CD28 comprising an amino acid sequence of SEQ ID NO:24.
- the antibody that specifically binds to BCMA and CD28 comprises a polypeptide that is at least 95%, 96%, 97%, 98%, or 99% identical to the sequence of SEQ ID NO: 92, a polypeptide that is at least 95%, 96%, 97%, 98%, or 99% identical to the sequence of SEQ ID NO: 26, a polypeptide that is at least 95%, 96%, 97%, 98%, or 99% identical to the sequence of SEQ ID NO: 27, and a polypeptide that is at least 95%, 96%, 97%, 98%, or 99% identical to the sequence of SEQ ID NO: 28.
- an antibody that specifically binds to BCMA and CD28, wherein said antibody comprises a first light chain comprising the amino acid sequence of SEQ ID NO:92, a first heavy chain comprising the amino acid sequence of SEQ ID NO:26, a second heavy chain comprising the amino acid sequence of SEQ ID NO:27 and a second light chain comprising the amino acid sequence of SEQ ID NO:28.
- an antibody that specifically binds to B cell maturation agent (BCMA) and CD28, wherein the antibody comprises
- VH BCMA heavy chain variable region
- VH comprising a heavy chain complementary determining region CDR-H1 of SEQ ID NO: 29 (GFTFSNAWMD), a CDR-H2 of SEQ ID NO: 30 (QITAKSNNYATYYADSVKG), and a CDR-H3 of SEQ ID NO: 31 (DGYH), and
- VH comprising a heavy chain complementary determining region CDR-H1 of SEQ ID NO: 29 (GFTFSNAWMD), a CDR-H2 of SEQ ID NO: 32 (QITAKSNNYATYYAAPVKG), and a CDR-H3 of SEQ ID NO: 31 (DGYH), and
- VLBCMA light chain variable region
- an antibody that specifically binds to BCMA and CD28, wherein said antibody comprises a heavy chain variable region (VH BCMA) comprising a CDR-H1 of SEQ ID NO: 29 (GFTFSNAWMD), a CDR-H2 of SEQ ID NO: 30 (QITAKSNNYATYYADSVKG), and a CDR-H3 of SEQ ID NO: 31 (DGYH), and a light chain variable region (VL BCMA) comprising a CDR-L1 of SEQ ID NO: 33 (RASEDIRNGLA), a CDR-L2 of SEQ ID NO: 34 (NANSLHT) and a CDR-L3 of SEQ ID NO: 35 (EDTSKYPYT).
- VH BCMA heavy chain variable region
- VL BCMA light chain variable region comprising a CDR-L1 of SEQ ID NO: 33
- RANSLHT NANSLHT
- EDTSKYPYT CDR-L3 of SEQ ID NO: 35
- an antibody that specifically binds to BCMA and CD28, wherein said antibody comprises a heavy chain variable region (VH BCMA) comprising a CDR-H1 of SEQ ID NO: 29 (GFTFSNAWMD), a CDR-H2 of SEQ ID NO: 32 (QITAKSNNYATYYAAPVKG), and a CDR-H3 of SEQ ID NO: 31 (DGYH), and a light chain variable region (VL BCMA) comprising a CDR-L1 of SEQ ID NO: 33 (RASEDIRNGLA), a CDR-L2 of SEQ ID NO: 34 (NANSLHT) and a CDR-L3 of SEQ ID NO: 35 (EDTSKYPYT).
- VH BCMA heavy chain variable region
- VL BCMA light chain variable region comprising a CDR-L1 of SEQ ID NO: 33
- RANSLHT NANSLHT
- EDTSKYPYT CDR-L3 of SEQ ID NO: 35
- an antibody that specifically binds to BCMA and CD28 wherein said antibody comprises a VH BCMA comprising an amino acid sequence selected from the group consisting of SEQ ID NO: 36 (VH2a), SEQ ID NO: 38 (VHlb), SEQ ID NO:48 (VHla), SEQ ID NO:49 (VHla_Y292D), SEQ ID NO:50 (VHlc hu CDR2) and SEQ ID NO:51 (VHla_W197Y), and/or the VL BCMA comprising an amino acid sequence selected from the group consisting of SEQ ID NO: 37 (VL2a), SEQ ID NO:39 (VLla), SEQ ID NO:52 (VLla_L2_GL), SEQ ID NO:53 (VL2a_L2_GL), SEQ ID NO:54 (VLla_Ll_pGL), SEQ ID NO 55 (VLla_N651A), SEQ ID NO:56 (VLla_N651A_N69
- an antibody that specifically binds to BCMA and CD28, wherein said antibody comprises a VH BCMA comprising an amino acid sequence selected from the group consisting of SEQ ID NO: 36 (VH2a) and SEQ ID NO: 38 (VHlb), and/or the V L BCMA comprising an amino acid sequence selected from the group consisting of SEQ ID NO: 37 (VL2a) and SEQ ID NO:39 (VLla).
- an antibody that specifically binds to BCMA and CD28 wherein said antibody comprises
- VH BCMA comprising an amino acid sequence of SEQ ID NO:36 and a VL BCMA comprising an amino acid sequence of SEQ ID NO:37, or
- VH BCMA comprising an amino acid sequence of SEQ ID NO:38 and a VL BCMA comprising an amino acid sequence of SEQ ID NO:39;
- VH BCMA comprising an amino acid sequence of SEQ ID NO:48 and VLBCMA comprising an amino acid sequence of SEQ ID NO:52, or
- VH BCMA comprising an amino acid sequence of SEQ ID NO:49 and VLBCMA comprising an amino acid sequence of SEQ ID NO:39;
- VH BCMA comprising an amino acid sequence of SEQ ID NO:49 and VLBCMA comprising an amino acid sequence of SEQ ID NO:52, or
- VH BCMA comprising an amino acid sequence of SEQ ID NO:49 and VLBCMA comprising an amino acid sequence of SEQ ID NO:58;
- VH BCMA comprising an amino acid sequence of SEQ ID NO:49 and VLBCMA comprising an amino acid sequence of SEQ ID NO:37, or
- VH BCMA comprising an amino acid sequence of SEQ ID NO:38 and VLBCMA comprising an amino acid sequence of SEQ ID NO:54;
- VH BCMA comprising an amino acid sequence of SEQ ID NO:38 and VLBCMA comprising an amino acid sequence of SEQ ID NO:55, or
- VH BCMA comprising an amino acid sequence of SEQ ID NO:38 and VLBCMA comprising an amino acid sequence of SEQ ID NO:56;
- VH BCMA comprising an amino acid sequence of SEQ ID NO:38 and VLBCMA comprising an amino acid sequence of SEQ ID NO:58, or
- VH BCMA comprising an amino acid sequence of SEQ ID NO:38 and VLBCMA comprising an amino acid sequence of SEQ ID NO:37; or
- VH BCMA comprising an amino acid sequence of SEQ ID NO:36 and VLBCMA comprising an amino acid sequence of SEQ ID NO:39, or
- VH BCMA comprising an amino acid sequence of SEQ ID NO:36 and VLBCMA comprising an amino acid sequence of SEQ ID NO:54;
- VH BCMA comprising an amino acid sequence of SEQ ID NO:36 and VLBCMA comprising an amino acid sequence of SEQ ID NO:58.
- an antibody that specifically binds to BCMA and CD28 wherein said antibody comprises
- VH BCMA comprising an amino acid sequence of SEQ ID NO:36 and a VLBCMA comprising an amino acid sequence of SEQ ID NO:37, or
- VH BCMA comprising an amino acid sequence of SEQ ID NO:38 and a VLBCMA comprising an amino acid sequence of SEQ ID NO:39.
- the antibody that specifically binds to BCMA and CD28 comprises a VH BCMA comprising an amino acid sequence of SEQ ID NO:36 and a VLBCMA comprising an amino acid sequence of SEQ ID NO:37.
- the first antigen binding domain that specifically binds to BCMA can be a Fab molecule.
- the first antigen binding domain that specifically binds to BCMA can be a cross-Fab molecule, i.e a Fab molecule wherein the variable domains VL and VH or the constant domains CL and CHI, particularly the variable domains VL and VH, of the Fab light chain and the Fab heavy chain are replaced by each other.
- the first antigen binding domain that specifically binds to BCMA is a Fab molecule wherein in the constant domain CL the amino acid at position 123 (numbering according to Kabat EU index) is substituted by an amino acid selected from lysine (K), arginine (R) or histidine (H) and the amino acid at position 124 (numbering according to Kabat EU index) is substituted independently by lysine (K), arginine (R) or histidine (H), and wherein in the constant domain CHI the amino acid at position 147 (numbering according to Kabat EU index) is substituted independently by glutamic acid (E) or aspartic acid (D) and the amino acid at position 213 (numbering according to Kabat EU index) is substituted independently by glutamic acid (E), or aspartic acid (D) (numbering according to Kabat EU index).
- the second antigen binding domain that specifically binds to CD28 can be a Fab molecule.
- the second antigen binding domain that specifically binds to CD28 is a cross-Fab molecule, i.e a Fab molecule wherein the variable domains VL and VH or the constant domains CL and CHI, particularly the variable domains VL and VH, of the Fab light chain and the Fab heavy chain are replaced by each other.
- the second antigen binding domain that specifically binds to CD28 comprises a heavy chain variable region (VHCD28) comprising a heavy chain complementary determining region CDR-H1 of SEQ ID NO: 17, a CDR-H2 of SEQ ID NO: 18, and a CDR-H3 of SEQ ID NO: 19, and a light chain variable region (VLCD28) comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 20, a CDR-L2 of SEQ ID NO: 21 and a CDR-L3 of SEQ ID NO: 22.
- VHCD28 heavy chain variable region comprising a heavy chain complementary determining region CDR-H1 of SEQ ID NO: 17, a CDR-H2 of SEQ ID NO: 18, and a CDR-H3 of SEQ ID NO: 19, and a light chain variable region (VLCD28) comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 20, a CDR-L2 of SEQ ID NO: 21 and
- the second antigen binding domain that specifically binds to CD28 comprises a heavy chain variable region (VHCD28) comprising an amino acid sequence of SEQ ID NO:23, and a light chain variable region (VLCD28) comprising an amino acid sequence of SEQ ID NO:24 (v8).
- VHCD28 heavy chain variable region
- VLCD28 light chain variable region
- an antibody that specifically binds to BCMA and CD28, wherein said antibody comprises a first antigen binding domain comprising a VH BCMA comprising an amino acid sequence of SEQ ID NO:36 and VLBCMA comprising an amino acid sequence of SEQ ID NO:37 and a second antigen binding domain comprising VH CD28 comprising an amino acid sequence of SEQ ID NO:23 and VL CD28 comprising an amino acid sequence of SEQ ID NO:24.
- the antibody that specifically binds to BCMA and CD28 comprises a polypeptide that is at least 95%, 96%, 97%, 98%, or 99% identical to the sequence of SEQ ID NO: 40, a polypeptide that is at least 95%, 96%, 97%, 98%, or 99% identical to the sequence of SEQ ID NO: 41, a polypeptide that is at least 95%, 96%, 97%, 98%, or 99% identical to the sequence of SEQ ID NO: 27, and a polypeptide that is at least 95%, 96%, 97%, 98%, or 99% identical to the sequence of SEQ ID NO: 28.
- an antibody that specifically binds to BCMA and CD28, wherein said antibody comprises a first light chain comprising the amino acid sequence of SEQ ID NO:40, a first heavy chain comprising the amino acid sequence of SEQ ID NO:41, a second heavy chain comprising the amino acid sequence of SEQ ID NO:27 and a second light chain comprising the amino acid sequence of SEQ ID NO:28.
- an antibody that specifically binds to BCMA and CD28, wherein said antibody comprises a first antigen binding domain comprising a VH BCMA comprising an amino acid sequence of SEQ ID NO:38 and VLBCMA comprising an amino acid sequence of SEQ ID NO:39 and a second antigen binding domain comprising VH CD28 comprising an amino acid sequence of SEQ ID NO:23 and VL CD28 comprising an amino acid sequence of SEQ ID NO:24.
- the antibody that specifically binds to BCMA and CD28 comprises a polypeptide that is at least 95%, 96%, 97%, 98%, or 99% identical to the sequence of SEQ ID NO: 94, a polypeptide that is at least 95%, 96%, 97%, 98%, or 99% identical to the sequence of SEQ ID NO: 93, a polypeptide that is at least 95%, 96%, 97%, 98%, or 99% identical to the sequence of SEQ ID NO: 27, and a polypeptide that is at least 95%, 96%, 97%, 98%, or 99% identical to the sequence of SEQ ID NO: 28.
- an antibody that specifically binds to BCMA and CD28, wherein said antibody comprises a first light chain comprising the amino acid sequence of SEQ ID NO:94, a first heavy chain comprising the amino acid sequence of SEQ ID NO:93, a second heavy chain comprising the amino acid sequence of SEQ ID NO:27 and a second light chain comprising the amino acid sequence of SEQ ID NO:28.
- the antibody that specifically binds to BCMA and CD28 comprises a Fc domain.
- the Fc domain is an IgG, particularly an IgGl Fc domain.
- the Fc domain is a human Fc domain, particularly a human IgGl Fc domain.
- the Fc domain comprises one or more amino acid substitution that reduces binding to an Fc receptor and/or effector function. In one particular aspect, it comprises the amino acid mutations L234A, L235A and P329G (numbering according to Kabat EU index).
- the Fc domain of the antibody that specifically binds to BCMA and CD28 consists of a pair of polypeptide chains comprising heavy chain domains of an immunoglobulin molecule.
- the Fc domain of an immunoglobulin G (IgG) molecule is a dimer, each subunit of which comprises the CH2 and CH3 IgG heavy chain constant domains.
- the two subunits of the Fc domain are capable of stable association with each other.
- the Fc domain confers favorable pharmacokinetic properties to the antigen binding molecules of the invention, including a long serum half-life which contributes to good accumulation in the target tissue and a favorable tissue-blood distribution ratio. On the other side, it may, however, lead to undesirable targeting of the bispecific antibodies of the invention to cells expressing Fc receptors rather than to the preferred antigen-bearing cells.
- the Fc domain of the antibody that specifically binds to BCMA and CD28 exhibits reduced binding affinity to an Fc receptor and/or reduced effector function, as compared to a native IgGl Fc domain.
- the Fc domain does not substantially bind to an Fc receptor and/or does not induce effector function.
- the Fc receptor is an Fey receptor.
- the Fc receptor is a human Fc receptor.
- the Fc receptor is an activating human Fey receptor, more specifically human FcyRIIIa, FcyRI or FcyRIIa, most specifically human FcyRIIIa.
- the Fc domain does not induce effector function.
- the reduced effector function can include, but is not limited to, one or more of the following: reduced complement dependent cytotoxicity (CDC), reduced antibody-dependent cell-mediated cytotoxicity (ADCC), reduced antibody-dependent cellular phagocytosis (ADCP), reduced cytokine secretion, reduced immune complex-mediated antigen uptake by antigen- presenting cells, reduced binding to NK cells, reduced binding to macrophages, reduced binding to monocytes, reduced binding to polymorphonuclear cells, reduced direct signaling inducing apoptosis, reduced dendritic cell maturation, or reduced T cell priming.
- CDC complement dependent cytotoxicity
- ADCC reduced antibody-dependent cell-mediated cytotoxicity
- ADCP reduced antibody-dependent cellular phagocytosis
- cytokine secretion reduced immune complex-mediated antigen uptake by antigen- presenting cells
- reduced binding to NK cells reduced binding to macrophages
- monocytes reduced binding to monocytes
- polymorphonuclear cells reduced direct signaling induc
- one or more amino acid modifications may be introduced into the Fc region of an antibody provided herein, thereby generating an Fc region variant.
- the Fc region variant may comprise a human Fc region sequence (e.g., a human IgGl, IgG2, IgG3 or IgG4 Fc region) comprising an amino acid modification (e.g. a substitution) at one or more amino acid positions.
- an antibody that specifically binds to BCMA and CD28 wherein the Fc region comprises one or more amino acid substitution that reduces binding to an Fc receptor, in particular towards Fey receptor.
- the Fc region comprises one or more amino acid substitution and wherein the ADCC induced by the antibody is reduced to 0-20% of the ADCC induced by an antibody comprising the wild-type human IgGl Fc region.
- the Fc domain of the antibody described herein comprises one or more amino acid mutation that reduces the binding affinity of the Fc domain to an Fc receptor and/or effector function.
- the same one or more amino acid mutation is present in each of the two subunits of the Fc domain.
- the Fc domain comprises an amino acid substitution at a position of E233, L234, L235, N297, P331 and P329 (EU numbering).
- the Fc domain comprises amino acid substitutions at positions 234 and 235 (EU numbering) and/or 329 (EU numbering) of the IgG heavy chains.
- an antibody that specifically binds to BCMA and CD28 which comprises an Fc domain with the amino acid substitutions L234A, L235A and P329G (“P329G LALA”, EU numbering) in the IgG heavy chains.
- the amino acid substitutions L234A and L235A refer to the so-called LALA mutation.
- the “P329G LALA” combination of amino acid substitutions almost completely abolishes Fey receptor binding of a human IgGl Fc domain and is described in International Patent Appl. Publ. No. WO 2012/130831 Al which also describes methods of preparing such mutant Fc domains and methods for determining its properties such as Fc receptor binding or effector functions.
- Fc domains with reduced Fc receptor binding and/or effector function also include those with substitution of one or more of Fc domain residues 238, 265, 269, 270, 297, 327 and 329 (U.S. Patent No. 6,737,056).
- Such Fc mutants include Fc mutants with substitutions at two or more of amino acid positions 265, 269, 270, 297 and 327, including the so-called “DANA” Fc mutant with substitution of residues 265 and 297 to alanine (US Patent No. 7,332,581).
- the Fc domain is an IgG4 Fc domain.
- IgG4 antibodies exhibit reduced binding affinity to Fc receptors and reduced effector functions as compared to IgGl antibodies.
- the Fc domain is an IgG4 Fc domain comprising an amino acid substitution at position S228 (Kabat numbering), particularly the amino acid substitution S228P.
- the Fc domain is an IgG4 Fc domain comprising amino acid substitutions L235E and S228P and P329G (EU numbering).
- IgG4 Fc domain mutants and their Fey receptor binding properties are also described in WO 2012/130831.
- Mutant Fc domains can be prepared by amino acid deletion, substitution, insertion or modification using genetic or chemical methods well known in the art. Genetic methods may include site-specific mutagenesis of the encoding DNA sequence, PCR, gene synthesis, and the like. The correct nucleotide changes can be verified for example by sequencing.
- Binding to Fc receptors can be easily determined e.g. by ELISA, or by Surface Plasmon Resonance (SPR) using standard instrumentation such as a BIAcore instrument (GE Healthcare), and Fc receptors such as may be obtained by recombinant expression.
- binding affinity of Fc domains or cell activating antibodies comprising an Fc domain for Fc receptors may be evaluated using cell lines known to express particular Fc receptors, such as human NK cells expressing Fcyllla receptor.
- Effector function of an Fc domain, or antigen binding molecules of the invention comprising an Fc domain can be measured by methods known in the art.
- a suitable assay for measuring ADCC is described herein.
- Other examples of in vitro assays to assess ADCC activity of a molecule of interest are described in U.S. Patent No. 5,500,362; Hellstrom et al. Proc Natl Acad Sci USA 83, 7059-7063 (1986) and Hellstrom et al., Proc Natl Acad Sci USA 82, 1499- 1502 (1985); U.S. Patent No. 5,821,337; Bruggemann et al., J Exp Med 166, 1351-1361 (1987).
- non-radioactive assays methods may be employed (see, for example, ACTE M nonradioactive cytotoxicity assay for flow cytometry (CellTechnology, Inc. Mountain View, CA); and CytoTox 96® non-radioactive cytotoxicity assay (Promega, Madison, WI)).
- Useful effector cells for such assays include peripheral blood mononuclear cells (PBMC) and Natural Killer (NK) cells.
- PBMC peripheral blood mononuclear cells
- NK Natural Killer
- ADCC activity of the molecule of interest may be assessed in vivo, e.g. in an animal model such as that disclosed in Clynes et al., Proc Natl Acad Sci USA 95, 652-656 (1998).
- binding of the Fc domain to a complement component, specifically to Clq is reduced. Accordingly, in some aspects wherein the Fc domain is engineered to have reduced effector function, said reduced effector function includes reduced CDC.
- Clq binding assays may be carried out to determine whether the bispecific antibodies of the invention are able to bind Clq and hence has CDC activity. See e.g., Clq and C3c binding ELISA in WO 2006/029879 and WO 2005/100402.
- a CDC assay may be performed (see, for example, Gazzano-Santoro et al., J Immunol Methods 202, 163 (1996); Cragg et al., Blood 101, 1045-1052 (2003); and Cragg and Glennie, Blood 103, 2738-2743 (2004)).
- the Fc domain exhibiting reduced binding affinity to an Fc receptor and/or reduced effector function, as compared to a native IgGl Fc domain is a human IgGl Fc domain comprising the amino acid substitutions L234A, L235A and optionally P329G, or a human IgG4 Fc domain comprising the amino acid substitutions S228P, L235E and optionally P329G (numberings according to Kabat EU index). More particularly, it is a human IgGl Fc domain comprising the amino acid substitutions L234A, L235A and P329G (numbering according to Kabat EU index).
- the antibody that specifically binds to BCMA and CD28 comprises different antigenbinding sites, fused to one or the other of the two subunits of the Fc domain, thus the two subunits of the Fc domain may be comprised in two non-identical polypeptide chains. Recombinant co-expression of these polypeptides and subsequent dimerization leads to several possible combinations of the two polypeptides. To improve the yield and purity of the bispecific antigen binding molecules of the invention in recombinant production, it will thus be advantageous to introduce in the Fc domain of the antibody described herein a modification promoting the association of the desired polypeptides.
- an antibody that specifically binds to BCMA and CD28 comprising a Fc domain composed of a first and a second subunit capable of stable association, wherein the Fc domain comprises a modification promoting the association of the first and second subunit of the Fc domain.
- the site of most extensive protein-protein interaction between the two subunits of a human IgG Fc domain is in the CH3 domain of the Fc domain.
- said modification is in the CH3 domain of the Fc domain.
- said modification is a so-called “knob-into-hole” modification, comprising a “knob” modification in one of the two subunits of the Fc domain and a “hole” modification in the other one of the two subunits of the Fc domain.
- an antibody that specifically binds to BCMA and CD28 wherein the antibody comprises a Fc domain composed of a first and a second subunit capable of stable association comprising one or more amino acid substitution that reduces the binding affinity of the antigen binding molecule to an Fc receptor and/or effector function, wherein the first subunit of the Fc domain comprises knobs and the second subunit of the Fc domain comprises holes according to the knobs into holes method.
- the first subunit of the Fc domain comprises the amino acid substitutions S354C and T366W (EU numbering) and the second subunit of the Fc domain comprises the amino acid substitutions Y349C, T366S and Y407V (numbering according to Kabat EU index).
- the method involves introducing a protuberance (“knob”) at the interface of a first polypeptide and a corresponding cavity (“hole”) in the interface of a second polypeptide, such that the protuberance can be positioned in the cavity so as to promote heterodimer formation and hinder homodimer formation.
- Protuberances are constructed by replacing small amino acid side chains from the interface of the first polypeptide with larger side chains (e.g. tyrosine or tryptophan).
- Compensatory cavities of identical or similar size to the protuberances are created in the interface of the second polypeptide by replacing large amino acid side chains with smaller ones (e.g. alanine or threonine).
- an amino acid residue is replaced with an amino acid residue having a larger side chain volume, thereby generating a protuberance within the CH3 domain of the first subunit which is positionable in a cavity within the CH3 domain of the second subunit, and in the CH3 domain of the second subunit of the Fc domain an amino acid residue is replaced with an amino acid residue having a smaller side chain volume, thereby generating a cavity within the CH3 domain of the second subunit within which the protuberance within the CH3 domain of the first subunit is positionable.
- the protuberance and cavity can be made by altering the nucleic acid encoding the polypeptides, e.g. by site-specific mutagenesis, or by peptide synthesis.
- the threonine residue at position 366 is replaced with a tryptophan residue (T366W)
- T366W tryptophan residue
- Y407V valine residue
- the threonine residue at position 366 is replaced with a serine residue (T366S) and the leucine residue at position 368 is replaced with an alanine residue (L368A).
- the serine residue at position 354 is replaced with a cysteine residue (S354C)
- the tyrosine residue at position 349 is replaced by a cysteine residue (Y349C).
- the first subunit of the Fc domain comprises the amino acid substitutions S354C and T366W (EU numbering) and the second subunit of the Fc domain comprises the amino acid substitutions Y349C, T366S and Y407V (numbering according to Kabat EU index).
- a modification promoting association of the first and the second subunit of the Fc domain comprises a modification mediating electrostatic steering effects, e.g. as described in PCT publication WO 2009/089004.
- this method involves replacement of one or more amino acid residues at the interface of the two Fc domain subunits by charged amino acid residues so that homodimer formation becomes electrostatically unfavorable but heterodimerization electrostatically favorable.
- the C-terminus of the heavy chain of the antibody as reported herein can be a complete C- terminus ending with the amino acid residues PGK.
- the C-terminus of the heavy chain can be a shortened C-terminus in which one or two of the C terminal amino acid residues have been removed.
- the C-terminus of the heavy chain is a shortened C-terminus ending P.
- the C-terminus of the heavy chain is a shortened C-terminus ending PG.
- a CD28 antigen binding molecule comprising a heavy chain including a C-terminal CH3 domain as specified herein comprises the C -terminal glycine-lysine dipeptide (G446 and K447, numbering according to Kabat EU index).
- a CD28 antigen binding molecule comprising a heavy chain including a C-terminal CH3 domain comprises a C-terminal glycine residue (G446, numbering according to Kabat EU index).
- an antibody that specifically binds to BCMA and CD28 characterized by monovalent binding to BCMA and CD28 comprising (a) a first antigen binding domain capable of specific binding to BCMA, (b) a second antigen binding domain capable of specific binding to CD28, and (c) a Fc domain composed of a first and a second subunit capable of stable association comprising one or more amino acid substitution that reduces the binding affinity of the antigen binding molecule to an Fc receptor and/or effector function, wherein the second antigen binding domain capable of specific binding to CD28 is a Fab fragment and in the Fab fragment either the variable domains VH and VL or the constant domains CHI and CL are exchanged according to the Crossmab technology.
- an antibody that specifically binds to BCMA and CD28 characterized by monovalent binding to BCMA and CD28 comprising (a) a first antigen binding domain capable of specific binding to BCMA, (b) a second antigen binding domain capable of specific binding to CD28, and (c) a Fc domain composed of a first and a second subunit capable of stable association comprising one or more amino acid substitution that reduces the binding affinity of the antigen binding molecule to an Fc receptor and/or effector function, wherein the first antigen binding domain capable of specific binding to BCMA is a Fab fragment and in the Fab fragment either the variable domains VH and VL or the constant domains CHI and CL are exchanged according to the Crossmab technology.
- Multispecific antibodies with a domain replacement/exchange in one binding arm are described in detail in W02009/080252 and Schaefer, W. et al, PNAS, 108 (2011) 11187-1191. They clearly reduce the byproducts caused by the mismatch of a light chain against a first antigen with the wrong heavy chain against the second antigen (compared to approaches without such domain exchange).
- the invention relates to an antibody that specifically binds to BCMA and CD28 comprising (a) a first antigen binding domain capable of specific binding to BCMA, (b) a second antigen binding domain capable of specific binding to CD28, and (c) a Fc domain composed of a first and a second subunit capable of stable association comprising one or more amino acid substitution that reduces the binding affinity of the antigen binding molecule to an Fc receptor and/or effector function, wherein in the Fab fragment capable of specific binding to CD28 the variable domains VL and VH are replaced by each other so that the VH domain is part of the light chain and the VL domain is part of the heavy chain.
- the invention relates to an antibody that specifically binds to BCMA and CD28 comprising (a) a first antigen binding domain capable of specific binding to BCMA, (b) a second antigen binding domain capable of specific binding to CD28, and (c) a Fc domain composed of a first and a second subunit capable of stable association comprising one or more amino acid substitution that reduces the binding affinity of the antigen binding molecule to an Fc receptor and/or effector function, wherein in the Fab fragment capable of specific binding to BCMA the variable domains VL and VH are replaced by each other so that the VH domain is part of the light chain and the VL domain is part of the heavy chain.
- the antibody that specifically binds to BCMA and CD28 comprises (a) a first antigen binding domain capable of specific binding to BCMA, (b) a second antigen binding domains capable of specific binding to a CD28, and (c) a Fc domain composed of a first and a second subunit capable of stable association comprising one or more amino acid substitution that reduces the binding affinity of the antigen binding molecule to an Fc receptor and/or effector function, can contain different charged amino acid substitutions (so-called “charged residues”). These modifications are introduced in the crossed or non-crossed CHI and CL domains.
- the invention relates to an antibody that specifically binds to BCMA and CD28, wherein in one of CL domains the amino acid at position 123 (EU numbering) has been replaced by arginine (R) and the amino acid at position 124 (EU numbering) has been substituted by lysine (K) and wherein in one of the CHI domains the amino acids at position 147 (EU numbering) and at position 213 (EU numbering) have been substituted by glutamic acid (E).
- the invention further provides isolated polynucleotides encoding a BCMA antibody or an antibody that specifically binds to BCMA and CD28 as described herein.
- the one or more isolated polynucleotides encoding the BCMA antibody or antibody that specifically binds to BCMA and CD28 may be expressed as a single polynucleotide that encodes the entire antigen binding molecule or as multiple (e.g., two or more) polynucleotides that are co-expressed.
- Polypeptides encoded by polynucleotides that are co-expressed may associate through, e.g., disulfide bonds or other means to form a functional antigen binding molecule.
- the light chain portion of an immunoglobulin may be encoded by a separate polynucleotide from the heavy chain portion of the immunoglobulin. When co-expressed, the heavy chain polypeptides will associate with the light chain polypeptides to form the immunoglobulin.
- the isolated polynucleotide encodes the entire antibody that specifically binds to BCMA and CD28 as described herein. In other aspects, the isolated polynucleotide encodes a polypeptide comprised in the antibody that specifically binds to BCMA and CD28 as described herein.
- the polynucleotide or nucleic acid is DNA.
- a polynucleotide of the present invention is RNA, for example, in the form of messenger RNA (mRNA). RNA of the present invention may be single stranded or double stranded.
- BCMA antibodies or antibodies that specifically bind to BCMA and CD28 as described herein may be obtained, for example, by solid-state peptide synthesis (e.g. Merrifield solid phase synthesis) or recombinant production.
- solid-state peptide synthesis e.g. Merrifield solid phase synthesis
- polynucleotide encoding the antibody that specifically binds to BCMA and CD28 or polypeptide fragments thereof, e.g., as described above is isolated and inserted into one or more vectors for further cloning and/or expression in a host cell.
- Such polynucleotide may be readily isolated and sequenced using conventional procedures.
- a vector, preferably an expression vector, comprising one or more of the polynucleotides as described herein is provided.
- the expression vector can be part of a plasmid, virus, or may be a nucleic acid fragment.
- the expression vector includes an expression cassette into which the polynucleotide encoding the antibody or polypeptide fragments thereof (i.e. the coding region) is cloned in operable association with a promoter and/or other transcription or translation control elements.
- a "coding region" is a portion of nucleic acid which consists of codons translated into amino acids.
- a "stop codon" (TAG, TGA, or TAA) is not translated into an amino acid, it may be considered to be part of a coding region, if present, but any flanking sequences, for example promoters, ribosome binding sites, transcriptional terminators, introns, 5' and 3' untranslated regions, and the like, are not part of a coding region.
- Two or more coding regions can be present in a single polynucleotide construct, e.g. on a single vector, or in separate polynucleotide constructs, e.g. on separate (different) vectors.
- any vector may contain a single coding region, or may comprise two or more coding regions, e.g.
- a vector of the present invention may encode one or more polypeptides, which are post- or co-translationally separated into the final proteins via proteolytic cleavage.
- a vector, polynucleotide, or nucleic acid of the invention may encode heterologous coding regions, either fused or unfused to a polynucleotide encoding the antibody of the invention or polypeptide fragments thereof, or variants or derivatives thereof.
- Heterologous coding regions include without limitation specialized elements or motifs, such as a secretory signal peptide or a heterologous functional domain. An operable association is when a coding region for a gene product, e.g.
- a polypeptide is associated with one or more regulatory sequences in such a way as to place expression of the gene product under the influence or control of the regulatory sequence(s).
- Two DNA fragments (such as a polypeptide coding region and a promoter associated therewith) are "operably associated” if induction of promoter function results in the transcription of mRNA encoding the desired gene product and if the nature of the linkage between the two DNA fragments does not interfere with the ability of the expression regulatory sequences to direct the expression of the gene product or interfere with the ability of the DNA template to be transcribed.
- a promoter region would be operably associated with a nucleic acid encoding a polypeptide if the promoter was capable of effecting transcription of that nucleic acid.
- the promoter may be a cell-specific promoter that directs substantial transcription of the DNA only in predetermined cells.
- Other transcription control elements besides a promoter, for example enhancers, operators, repressors, and transcription termination signals, can be operably associated with the polynucleotide to direct cell-specific transcription.
- transcription control regions which function in vertebrate cells, such as, but not limited to, promoter and enhancer segments from cytomegaloviruses (e.g. the immediate early promoter, in conjunction with intron-A), simian virus 40 (e.g. the early promoter), and retroviruses (such as, e.g. Rous sarcoma virus).
- transcription control regions include those derived from vertebrate genes such as actin, heat shock protein, bovine growth hormone and rabbit a-globin, as well as other sequences capable of controlling gene expression in eukaryotic cells.
- tissue-specific promoters and enhancers as well as inducible promoters (e.g. promoters inducible tetracyclins).
- inducible promoters e.g. promoters inducible tetracyclins
- translation control elements include, but are not limited to ribosome binding sites, translation initiation and termination codons, and elements derived from viral systems (particularly an internal ribosome entry site, or IRES, also referred to as a CITE sequence).
- the expression cassette may also include other features such as an origin of replication, and/or chromosome integration elements such as retroviral long terminal repeats (LTRs), or adeno-associated viral (AAV) inverted terminal repeats (ITRs).
- LTRs retroviral long terminal repeats
- AAV adeno-associated viral inverted terminal repeats
- Polynucleotide and nucleic acid coding regions as described herein may be associated with additional coding regions which encode secretory or signal peptides, which direct the secretion of a polypeptide encoded by a polynucleotide of the present invention.
- DNA encoding a signal sequence may be placed upstream of the nucleic acid encoding an antibody as described herein or polypeptide fragments thereof.
- proteins secreted by mammalian cells have a signal peptide or secretory leader sequence which is cleaved from the mature protein once export of the growing protein chain across the rough endoplasmic reticulum has been initiated.
- polypeptides secreted by vertebrate cells generally have a signal peptide fused to the N-terminus of the polypeptide, which is cleaved from the translated polypeptide to produce a secreted or "mature" form of the polypeptide.
- the native signal peptide e.g. an immunoglobulin heavy chain or light chain signal peptide is used, or a functional derivative of that sequence that retains the ability to direct the secretion of the polypeptide that is operably associated with it.
- a heterologous mammalian signal peptide, or a functional derivative thereof may be used.
- the wild-type leader sequence may be substituted with the leader sequence of human tissue plasminogen activator (TP A) or mouse P-glucuronidase.
- DNA encoding a short protein sequence that could be used to facilitate later purification (e.g. a histidine tag) or assist in labeling the antibody as described herein may be included within or at the ends of the polynucleotide encoding the antibody as described herein or polypeptide fragments thereof.
- a host cell comprising one or more polynucleotides as described herein.
- a host cell comprising one or more vectors as described herein.
- the polynucleotides and vectors may incorporate any of the features, singly or in combination, described herein in relation to polynucleotides and vectors, respectively.
- a host cell comprises (e.g. has been transformed or transfected with) a vector comprising a polynucleotide that encodes (part of) an antibody as disclosed herein.
- the term "host cell” refers to any kind of cellular system which can be engineered to generate the fusion proteins of the invention or fragments thereof.
- Host cells suitable for replicating and for supporting expression of antigen binding molecules are well known in the art. Such cells may be transfected or transduced as appropriate with the particular expression vector and large quantities of vector containing cells can be grown for seeding large scale fermenters to obtain sufficient quantities of the antibody for clinical applications.
- Suitable host cells include prokaryotic microorganisms, such as E. coli, or various eukaryotic cells, such as Chinese hamster ovary cells (CHO), insect cells, or the like.
- polypeptides may be produced in bacteria in particular when glycosylation is not needed. After expression, the polypeptide may be isolated from the bacterial cell paste in a soluble fraction and can be further purified.
- eukaryotic microbes such as filamentous fungi or yeast are suitable cloning or expression hosts for polypeptide-encoding vectors, including fungi and yeast strains whose glycosylation pathways have been “humanized”, resulting in the production of a polypeptide with a partially or fully human glycosylation pattern. See Gerngross, Nat Biotech 22, 1409-1414 (2004), and Li et al., Nat Biotech 24, 210-215 (2006).
- Suitable host cells for the expression of (glycosylated) polypeptides are also derived from multicellular organisms (invertebrates and vertebrates).
- invertebrate cells include plant and insect cells. Numerous baculoviral strains have been identified which may be used in conjunction with insect cells, particularly for transfection of Spodoptera frugiperda cells. Plant cell cultures can also be utilized as hosts. See e.g. US Patent Nos. 5,959,177, 6,040,498, 6,420,548, 7,125,978, and 6,417,429 (describing PLANTIBODIESTM technology for producing antibodies in transgenic plants). Vertebrate cells may also be used as hosts.
- mammalian cell lines that are adapted to grow in suspension may be useful.
- useful mammalian host cell lines are monkey kidney CV1 line transformed by SV40 (COS-7); human embryonic kidney line (293 or 293T cells as described, e.g., in Graham et al., J Gen Virol 36, 59 (1977)), baby hamster kidney cells (BHK), mouse sertoli cells (TM4 cells as described, e.g., in Mather, Biol Reprod 23, 243-251 (1980)), monkey kidney cells (CV1), African green monkey kidney cells (VERO-76), human cervical carcinoma cells (HELA), canine kidney cells (MDCK), buffalo rat liver cells (BRL 3 A), human lung cells (W138), human liver cells (Hep G2), mouse mammary tumor cells (MMT 060562), TRI cells (as described, e.g., in Mather et al., Annals N.Y.
- MRC 5 cells MRC 5 cells
- FS4 cells Other useful mammalian host cell lines include Chinese hamster ovary (CHO) cells, including dhfr- CHO cells (Urlaub et al., Proc Natl Acad Sci USA 77, 4216 (1980)); and myeloma cell lines such as YO, NS0, P3X63 and Sp2/0.
- CHO Chinese hamster ovary
- dhfr- CHO cells Urlaub et al., Proc Natl Acad Sci USA 77, 4216 (1980)
- myeloma cell lines such as YO, NS0, P3X63 and Sp2/0.
- Host cells include cultured cells, e.g., mammalian cultured cells, yeast cells, insect cells, bacterial cells and plant cells, to name only a few, but also cells comprised within a transgenic animal, transgenic plant or cultured plant or animal tissue.
- the host cell is a eukaryotic cell, preferably a mammalian cell, such as a Chinese Hamster Ovary (CHO) cell, a human embryonic kidney (HEK) cell or a lymphoid cell (e.g., Y0, NSO, Sp20 cell). Standard technologies are known in the art to express foreign genes in these systems.
- Cells expressing a polypeptide comprising either the heavy or the light chain of an immunoglobulin may be engineered so as to also express the other of the immunoglobulin chains such that the expressed product is an immunoglobulin that has both a heavy and a light chain.
- a method of producing an antibody that specifically binds to BCMA and CD28 or polypeptide fragments thereof comprises culturing a host cell comprising polynucleotides encoding the antibody or polypeptide fragments thereof, as provided herein, under conditions suitable for expression of the antibody or polypeptide fragments thereof, and recovering the antibody as described herein or polypeptide fragments thereof from the host cell (or host cell culture medium).
- the antigen binding domain capable of specific binding to BCMA e.g. a Fab fragment
- the antigen binding domain capable of specific binding to BCMA comprises at least an immunoglobulin variable region capable of binding to an antigen.
- Variable regions can form part of and be derived from naturally or non-naturally occurring antibodies and fragments thereof.
- Methods to produce polyclonal antibodies and monoclonal antibodies are well known in the art (see e.g. Harlow and Lane, "Antibodies, a laboratory manual", Cold Spring Harbor Laboratory, 1988).
- Non-naturally occurring antibodies can be constructed using solid phase-peptide synthesis, can be produced recombinantly (e.g. as described in U.S. patent No. 4,186,567) or can be obtained, for example, by screening combinatorial libraries comprising variable heavy chains and variable light chains (see e.g. U.S. Patent. No. 5,969,108 to McCafferty).
- Non-limiting immunoglobulins useful can be of murine, primate, or human origin. If the antibody is intended for human use, a chimeric form of immunoglobulin may be used wherein the constant regions of the immunoglobulin are from a human.
- a humanized or fully human form of the immunoglobulin can also be prepared in accordance with methods well known in the art (see e. g. U.S. Patent No. 5,565,332 to Winter). Humanization may be achieved by various methods including, but not limited to (a) grafting the non-human (e.g., donor antibody) CDRs onto human (e.g.
- recipient antibody framework and constant regions with or without retention of critical framework residues (e.g. those that are important for retaining good antigen binding affinity or antibody functions), (b) grafting only the non-human specificity-determining regions (SDRs or a-CDRs; the residues critical for the antibody-antigen interaction) onto human framework and constant regions, or (c) transplanting the entire non-human variable domains, but "cloaking" them with a human-like section by replacement of surface residues.
- critical framework residues e.g. those that are important for retaining good antigen binding affinity or antibody functions
- Particular immunoglobulins according to the invention are human immunoglobulins.
- Human antibodies and human variable regions can be produced using various techniques known in the art. Human antibodies are described generally in van Dijk and van de Winkel, Curr Opin Pharmacol 5, 368-74 (2001) and Lonberg, Curr Opin Immunol 20, 450-459 (2008). Human variable regions can form part of and be derived from human monoclonal antibodies made by the hybridoma method (see e.g. Monoclonal Antibody Production Techniques and Applications, pp. 51-63 (Marcel Dekker, Inc., New York, 1987)).
- Human antibodies and human variable regions may also be prepared by administering an immunogen to a transgenic animal that has been modified to produce intact human antibodies or intact antibodies with human variable regions in response to antigenic challenge (see e.g. Lonberg, Nat Biotech 23, 1117-1125 (2005). Human antibodies and human variable regions may also be generated by isolating Fv clone variable region sequences selected from human-derived phage display libraries (see e.g., Hoogenboom et al.
- Phage typically display antibody fragments, either as single-chain Fv (scFv) fragments or as Fab fragments.
- the antigen binding domains comprised in the antibodies described herein are engineered to have enhanced binding affinity according to, for example, the methods disclosed in PCT publication WO 2012/020006 (see Examples relating to affinity maturation) or U.S. Pat. Appl. Publ. No. 2004/0132066.
- the ability of the antibodies of the invention to bind to a specific antigenic determinant can be measured either through an enzyme-linked immunosorbent assay (ELISA) or other techniques familiar to one of skill in the art, e.g. surface plasmon resonance technique (Liljeblad, et al., Glyco J 17, 323-329 (2000)), and traditional binding assays (Heeley, Endocr Res 28, 217-229 (2002)).
- ELISA enzyme-linked immunosorbent assay
- Competition assays may be used to identify an antigen binding molecule that competes with a reference antibody for binding to a particular antigen.
- a competing antigen binding molecule binds to the same epitope (e.g. a linear or a conformational epitope) that is bound by the reference antigen binding molecule.
- epitope e.g. a linear or a conformational epitope
- Detailed exemplary methods for mapping an epitope to which an antigen binding molecule binds are provided in Morris (1996) “Epitope Mapping Protocols”, in Methods in Molecular Biology vol. 66 (Humana Press, Totowa, NJ).
- immobilized antigen is incubated in a solution comprising a first labeled antigen binding molecule that binds to the antigen and a second unlabeled antigen binding molecule that is being tested for its ability to compete with the first antigen binding molecule for binding to the antigen.
- the second antigen binding molecule may be present in a hybridoma supernatant.
- immobilized antigen is incubated in a solution comprising the first labeled antigen binding molecule but not the second unlabeled antigen binding molecule. After incubation under conditions permissive for binding of the first antibody to the antigen, excess unbound antibody is removed, and the amount of label associated with immobilized antigen is measured.
- Bispecific antibodies prepared as described herein may be purified by art-known techniques such as high performance liquid chromatography, ion exchange chromatography, gel electrophoresis, affinity chromatography, size exclusion chromatography, and the like.
- the actual conditions used to purify a particular protein will depend, in part, on factors such as net charge, hydrophobicity, hydrophilicity etc., and will be apparent to those having skill in the art.
- affinity chromatography purification an antibody, ligand, receptor or antigen can be used to which the antigen binding molecule binds.
- a matrix with protein A or protein G may be used.
- Sequential Protein A or G affinity chromatography and size exclusion chromatography can be used to isolate an antigen binding molecule essentially as described in the Examples.
- the purity of the antibody or fragments thereof can be determined by any of a variety of well-known analytical methods including gel electrophoresis, high pressure liquid chromatography, and the like.
- the bispecific antibody expressed as described in the Examples was shown to be intact and properly assembled as demonstrated by reducing and non-reducing SDS-PAGE.
- BCMA antibody or the antibody that specifically binds to BCMA and CD28 provided herein may be identified, screened for, or characterized for its physical/chemical properties and/or biological activities by various assays known in the art. 1. Affinity assays
- the affinity of the antigen binding molecule provided herein for the corresponding target can be determined in accordance with the methods set forth in the Examples by surface plasmon resonance (SPR), using standard instrumentation such as a Proteon instrument (Bio-rad), and receptors or target proteins such as may be obtained by recombinant expression.
- the affinity of the antigen binding molecule for the target cell antigen can also be determined by surface plasmon resonance (SPR), using standard instrumentation such as a Proteon instrument (Biorad), and receptors or target proteins such as may be obtained by recombinant expression.
- KD is measured by surface plasmon resonance using a Proteon ® machine (Bio-Rad) at 25 °C.
- Binding of the antibody or bispecific antibody provided herein to the corresponding receptor expressing cells may be evaluated using cell lines expressing the particular receptor or target antigen, for example by flow cytometry (FACS).
- FACS flow cytometry
- CHO cells expressing human CD28 parental cell line CHO-kl ATCC #CCL-61, modified to stably overexpress human CD28 are used in the binding assay.
- cancer cell lines expressing BCMA were used to demonstrate the binding of the bispecific antigen binding molecules to the target cell antigen.
- assays are provided for identifying the biuological activity of the antibody that specifically binds to BCMA and CD28.
- Biological activity may include, e.g. T cell proliferation and cytokine secretion as measured with the methods as described in Example 4.
- Antigen binding molecules having such biological activity in vivo and/or in vitro are also provided.
- the invention provides pharmaceutical compositions comprising any of the antibodies that specifically bind to BCMA and CD28 provided herein, e.g., for use in any of the below therapeutic methods.
- a pharmaceutical composition comprises an antibody that specifically binds to BCMA and CD28 provided herein and at least one pharmaceutically acceptable excipient.
- a pharmaceutical composition comprises an antibody that specifically binds to BCMA and CD28 provided herein and at least one additional therapeutic agent, e.g., as described below.
- Pharmaceutical compositions as disclosed herein comprise a therapeutically effective amount of one or more antigen binding molecules dissolved or dispersed in a pharmaceutically acceptable excipient.
- compositions that are generally non-toxic to recipients at the dosages and concentrations employed, i.e. do not produce an adverse, allergic or other untoward reaction when administered to an animal, such as, for example, a human, as appropriate.
- the preparation of a pharmaceutical composition that contains at least one antibody that specifically binds to BCMA and CD28 and optionally an additional active ingredient will be known to those of skill in the art in light of the present disclosure, as exemplified by Remington's Pharmaceutical Sciences, 18th Ed. Mack Printing Company, 1990, incorporated herein by reference.
- the compositions are lyophilized formulations or aqueous solutions.
- pharmaceutically acceptable excipient includes any and all solvents, buffers, dispersion media, coatings, surfactants, antioxidants, preservatives (e.g. antibacterial agents, antifungal agents), isotonic agents, salts, stabilizers and combinations thereof, as would be known to one of ordinary skill in the art.
- Parenteral compositions include those designed for administration by injection, e.g. subcutaneous, intradermal, intralesional, intravenous, intraarterial intramuscular, intrathecal or intraperitoneal injection.
- the antibodies or bispecific antibodies may be formulated in aqueous solutions, preferably in physiologically compatible buffers such as Hanks' solution, Ringer's solution, or physiological saline buffer.
- the solution may contain formulation agents such as suspending, stabilizing and/or dispersing agents.
- the antibody that specifically binds to BCMA and CD28 may be in powder form for constitution with a suitable vehicle, e.g., sterile pyrogen-free water, before use.
- Sterile injectable solutions are prepared by incorporating the fusion proteins of the invention in the required amount in the appropriate solvent with various of the other ingredients enumerated below, as required. Sterility may be readily accomplished, e.g., by filtration through sterile filtration membranes. Generally, dispersions are prepared by incorporating the various sterilized active ingredients into a sterile vehicle which contains the basic dispersion medium and/or the other ingredients. In the case of sterile powders for the preparation of sterile injectable solutions, suspensions or emulsion, the preferred methods of preparation are vacuum-drying or freeze-drying techniques which yield a powder of the active ingredient plus any additional desired ingredient from a previously sterile- filtered liquid medium thereof.
- the liquid medium should be suitably buffered if necessary and the liquid diluent first rendered isotonic prior to injection with sufficient saline or glucose.
- the composition must be stable under the conditions of manufacture and storage, and preserved against the contaminating action of microorganisms, such as bacteria and fungi. It will be appreciated that endotoxin contamination should be kept minimally at a safe level, for example, less than 0.5 ng/mg protein.
- Suitable pharmaceutically acceptable excipients include, but are not limited to: buffers such as phosphate, citrate, and other organic acids; antioxidants including ascorbic acid and methionine; preservatives (such as octadecyldimethylbenzyl ammonium chloride; hexamethonium chloride; benzalkonium chloride; benzethonium chloride; phenol, butyl or benzyl alcohol; alkyl parabens such as methyl or propyl paraben; catechol; resorcinol; cyclohexanol; 3-pentanol; and m-cresol); low molecular weight (less than about 10 residues) polypeptides; proteins, such as serum albumin, gelatin, or immunoglobulins; hydrophilic polymers such as polyvinylpyrrolidone; amino acids such as glycine, glutamine, asparagine, histidine, arginine, or lysine; monos
- Aqueous injection suspensions may contain compounds which increase the viscosity of the suspension, such as sodium carboxymethyl cellulose, sorbitol, dextran, or the like.
- the suspension may also contain suitable stabilizers or agents which increase the solubility of the compounds to allow for the preparation of highly concentrated solutions.
- suspensions of the active compounds may be prepared as appropriate oily injection suspensions.
- Suitable lipophilic solvents or vehicles include fatty oils such as sesame oil, or synthetic fatty acid esters, such as ethyl cleats or triglycerides, or liposomes.
- Active ingredients may be entrapped in microcapsules prepared, for example, by coacervation techniques or by interfacial polymerization, for example, hydroxymethylcellulose or gelatin-microcapsules and poly-(methylmethacylate) microcapsules, respectively, in colloidal drug delivery systems (for example, liposomes, albumin microspheres, microemulsions, nanoparticles and nanocapsules) or in macroemulsions.
- colloidal drug delivery systems for example, liposomes, albumin microspheres, microemulsions, nanoparticles and nanocapsules
- Sustained-release preparations may be prepared.
- sustained-release preparations include semipermeable matrices of solid hydrophobic polymers containing the polypeptide, which matrices are in the form of shaped articles, e.g. films, or microcapsules.
- prolonged absorption of an injectable composition can be brought about by the use in the compositions of agents delaying absorption, such as, for example, aluminum monostearate, gelatin or combinations thereof.
- Exemplary pharmaceutically acceptable excipients herein further include insterstitial drug dispersion agents such as soluble neutral -active hyaluronidase glycoproteins (sHASEGP), for example, human soluble PH-20 hyaluronidase glycoproteins, such as rHuPH20 (HYLENEX®, Baxter International, Inc.).
- sHASEGP soluble neutral -active hyaluronidase glycoproteins
- rHuPH20 HYLENEX®, Baxter International, Inc.
- Certain exemplary sHASEGPs and methods of use, including rHuPH20 are described in US Patent Publication Nos. 2005/0260186 and 2006/0104968.
- a sHASEGP is combined with one or more additional glycosaminoglycanases such as chondroitinases.
- additional glycosaminoglycanases such as chondroitinases.
- Exemplary lyophilized antibody formulations are described in US Patent No
- Aqueous antibody formulations include those described in US Patent No. 6,171,586 and W02006/044908, the latter formulations including a histidine-acetate buffer.
- the antibody that specifically binds to BCMA and CD28 may also be formulated as a depot preparation. Such long acting formulations may be administered by implantation (for example subcutaneously or intramuscularly) or by intramuscular injection.
- the antibody that specifically binds to BCMA and CD28 may be formulated with suitable polymeric or hydrophobic materials (for example as an emulsion in an acceptable oil) or ion exchange resins, or as sparingly soluble derivatives, for example, as a sparingly soluble salt.
- compositions comprising the antibody that specifically binds to BCMA and CD28 may be manufactured by means of conventional mixing, dissolving, emulsifying, encapsulating, entrapping or lyophilizing processes. Pharmaceutical compositions may be formulated in conventional manner using one or more physiologically acceptable carriers, diluents, excipients or auxiliaries which facilitate processing of the proteins into preparations that can be used pharmaceutically. Proper formulation is dependent upon the route of administration chosen.
- the antibody that specifically binds to BCMA and CD28 may be formulated into a composition in a free acid or base, neutral or salt form.
- Pharmaceutically acceptable salts are salts that substantially retain the biological activity of the free acid or base. These include the acid addition salts, e.g.
- salts formed with the free carboxyl groups can also be derived from inorganic bases such as for example, sodium, potassium, ammonium, calcium or ferric hydroxides; or such organic bases as isopropylamine, trimethylamine, histidine or procaine. Pharmaceutical salts tend to be more soluble in aqueous and other protic solvents than are the corresponding free base forms.
- composition herein may also contain more than one active ingredient as necessary for the particular indication being treated, preferably those with complementary activities that do not adversely affect each other.
- active ingredients are suitably present in combination in amounts that are effective for the purpose intended.
- the formulations to be used for in vivo administration are generally sterile. Sterility may be readily accomplished, e.g., by filtration through sterile filtration membranes.
- any of the antibodies that specifically bind to BCMA and CD28 provided herein may be used in therapeutic methods, either alone or in combination.
- an antibody that specifically binds to BCMA and CD28 for use as a medicament is provided.
- an antibody that specifically binds to BCMA and CD28 for use in treating cancer is provided.
- an antibody that specifically binds to BCMA and CD28 for use in treating hematologic malignancies is provided.
- the term “hematologic malignancies” includes diseases selected from the group consisting of multiple myeloma (MM), chronic lymphocytic leukemia, acute B-lymphoblastic leukemia, non-Hodgkin lymphoma (NHL), and Hodgkin lymphoma, but also acute myeloid leukemia and acute lymphoblastic leukemia.
- an antibody that specifically binds to BCMA and CD28 for use in treating multiple myeloma (MM) is provided.
- an antibody that specifically binds to BCMA and CD28 for use in a method of treatment is provided.
- an antibody that specifically binds to BCMA and CD28 for use in a method of treating an individual having cancer comprising administering to the individual an effective amount of the antibody that specifically binds to BCMA and CD28.
- the method further comprises administering to the individual an effective amount of at least one additional therapeutic agent.
- the antibody that specifically binds to BCMA and CD28 as described herein is for use in inhibiting the growth of BCMA-expressing cancer cells.
- an antibody that specifically binds to BCMA and CD28 for use in a method of treatment is provided.
- an antibody that specifically binds to BCMA and CD28 for use in a method of treating an individual having cancer comprising administering to the individual an effective amount of the antibody that specifically binds to BCMA and CD28.
- an antibody that specifically binds to BCMA and CD28 for use in a method of treating an individual having BCMA-expressing cancer, in particular hematologic malignancies selected from the group consisting of multiple myeloma (MM), chronic lymphocytic leukemia, acute B-lymphoblastic leukemia, non-Hodgkin lymphoma (NHL), Hodgkin lymphoma, acute myeloid leukemia and acute lymphoblastic leukemia, comprising administering to the individual an effective amount of the antibody that specifically binds to BCMA and CD28.
- the method further comprises administering to the individual an effective amount of at least one additional therapeutic agent.
- the medicament is for treatment of cancer, particularly BCMA-expressing cancer.
- the medicament is for use in a method of treating cancer comprising administering to an individual having cancer an effective amount of the medicament.
- the method further comprises administering to the individual an effective amount of at least one additional therapeutic agent, e.g., as described below.
- the medicament is for treatment of BCMA-expressing cancer.
- the medicament is for use in a method of treating cancer, in particular BCMA-expressing cancer, comprising administering to an individual having cancer an effective amount of the medicament.
- a method for treating a cancer in particular BCMA-expressing cancer.
- the method comprises administering to an individual having cancer an effective amount of an antibody that specifically binds to BCMA and CD28.
- the method further comprises administering to the individual an effective amount of at least one additional therapeutic agent, as described below.
- An “individual” according to any of the above aspects may be a human.
- a pharmaceutical formulation comprising any of the antibodies that specifically bind to BCMA and CD28 as reported herein, e.g., for use in any of the above therapeutic methods.
- a pharmaceutical formulation comprises an antibody that specifically binds to BCMA and CD28 as reported herein and a pharmaceutically acceptable carrier.
- a pharmaceutical formulation comprises an antibody that specifically binds to BCMA and CD28 as reported herein and at least one additional therapeutic agent.
- an antibody that specifically binds to BCMA and CD28 as reported herein can be used either alone or in combination with other agents in a therapy.
- an antibody that specifically binds to BCMA and CD28 as reported herein may be co-administered with at least one additional therapeutic agent.
- an antibody that specifically binds to BCMA and CD28 as described herein for use in cancer immunotherapy is provided.
- an antibody that specifically binds to BCMA and CD28 for use in a method of cancer immunotherapy is provided.
- An “individual” according to any of the above aspects is preferably a human.
- combination therapies noted above encompass combined administration (where two or more therapeutic agents are included in the same or separate formulations), and separate administration, in which case, administration of the antibody as reported herein can occur prior to, simultaneously, and/or following, administration of the additional therapeutic agent or agents.
- administration of the antibody that specifically binds to BCMA and CD28 and administration of an additional therapeutic agent occur within about one month, or within about one, two or three weeks, or within about one, two, three, four, five, or six days, of each other.
- An antigen binding molecule as reported herein can be administered by any suitable means, including parenteral, intrapulmonary, and intranasal, and, if desired for local treatment, intralesional administration.
- Parenteral infusions include intramuscular, intravenous, intraarterial, intraperitoneal, or subcutaneous administration. Dosing can be by any suitable route, e.g. by injections, such as intravenous or subcutaneous injections, depending in part on whether the administration is brief or chronic.
- Various dosing schedules including but not limited to single or multiple administrations over various time-points, bolus administration, and pulse infusion are contemplated herein.
- Antibodies that specifically bind to BCMA and CD28 as described herein would be formulated, dosed, and administered in a fashion consistent with good medical practice. Factors for consideration in this context include the particular disorder being treated, the particular mammal being treated, the clinical condition of the individual patient, the cause of the disorder, the site of delivery of the agent, the method of administration, the scheduling of administration, and other factors known to medical practitioners.
- the antibody that specifically binds to BCMA and CD28 needs not be, but is optionally formulated with one or more agents currently used to prevent or treat the disorder in question. The effective amount of such other agents depends on the amount of antibody present in the formulation, the type of disorder or treatment, and other factors discussed above. These are generally used in the same dosages and with administration routes as described herein, or about from 1 to 99% of the dosages described herein, or in any dosage and by any route that is empirically/clinically determined to be appropriate.
- an antibody that specifically binds to BCMA and CD28 as described herein when used alone or in combination with one or more other additional therapeutic agents, will depend on the type of disease to be treated, the type of antibody, the severity and course of the disease, whether the antibody is administered for preventive or therapeutic purposes, previous therapy, the patient's clinical history and response to the antibody, and the discretion of the attending physician.
- the antibody that specifically binds to BCMA and CD28 is suitably administered to the patient at one time or over a series of treatments. Depending on the type and severity of the disease, about 1 pg/kg to 15 mg/kg (e.g.
- 0.5 mg/kg - 10 mg/kg) of bispecific agonistic CD28 antigen binding molecule can be an initial candidate dosage for administration to the patient, whether, for example, by one or more separate administrations, or by continuous infusion.
- One typical daily dosage might range from about 1 pg/kg to 100 mg/kg or more, depending on the factors mentioned above.
- the treatment would generally be sustained until a desired suppression of disease symptoms occurs.
- One exemplary dosage of the antibody would be in the range from about 0.05 mg/kg to about 10 mg/kg.
- one or more doses of about 0.5 mg/kg, 2.0 mg/kg, 4.0 mg/kg or 10 mg/kg (or any combination thereof) may be administered to the patient.
- Such doses may be administered intermittently, e.g. every week or every three weeks (e.g. such that the patient receives from about two to about twenty, or e.g. about six doses of the antibody).
- An initial higher loading dose, followed by one or more lower doses may be administered.
- other dosage regimens may be useful. The progress of this therapy is easily monitored by conventional techniques and assays.
- the antibody that specifically binds to BCMA and CD28 may be administered in combination with one or more other agents in therapy.
- an antigen binding molecule as described herein may be co-administered with at least one additional therapeutic agent.
- therapeutic agent encompasses any agent that can be administered for treating a symptom or disease in an individual in need of such treatment.
- additional therapeutic agent may comprise any active ingredients suitable for the particular indication being treated, preferably those with complementary activities that do not adversely affect each other.
- an additional therapeutic agent is another anti-cancer agent, for example a microtubule disruptor, an antimetabolite, a topoisomerase inhibitor, a DNA intercalator, an alkylating agent, a hormonal therapy, a kinase inhibitor, a receptor antagonist, an activator of tumor cell apoptosis, or an anti angiogenic agent.
- an additional therapeutic agent is an immunomodulatory agent, a cytostatic agent, an inhibitor of cell adhesion, a cytotoxic or cytostatic agent, an activator of cell apoptosis, or an agent that increases the sensitivity of cells to apoptotic inducers.
- antibodies that specifically bind to BCMA and CD28 as disclosed herein or pharmaceutical compositions comprising them for use in the treatment of cancer wherein the bispecific antibody is administered in combination with a chemotherapeutic agent, radiation and/ or other agents for use in cancer immunotherapy.
- Such other agents are suitably present in combination in amounts that are effective for the purpose intended.
- the effective amount of such other agents depends on the amount of fusion protein used, the type of disorder or treatment, and other factors discussed above.
- the bispecific antigen binding molecule or antibody of the invention are generally used in the same dosages and with administration routes as described herein, or about from 1 to 99% of the dosages described herein, or in any dosage and by any route that is empirically/clinically determined to be appropriate.
- combination therapies noted above encompass combined administration (where two or more therapeutic agents are included in the same or separate compositions), and separate administration, in which case, administration of the bispecific antigen binding molecule or antibody of the invention can occur prior to, simultaneously, and/or following, administration of the additional therapeutic agent and/or adjuvant.
- immunomodulator refers to any substance including a monoclonal antibody that effects the immune system.
- the molecules of the inventions can be considered immunomodulators.
- Immunomodulators can be used as anti -neoplastic agents for the treatment of cancer.
- immunomodulators include, but are not limited to anti- CTLA4 antibodies (e.g. ipilimumab), anti-PDl antibodies (e.g.
- combination therapies noted above encompass combined administration (where two or more therapeutic agents are included in the same or separate compositions), and separate administration, in which case, administration of the bispecific antigen binding molecule can occur prior to, simultaneously, and/or following, administration of the additional therapeutic agent and/or adjuvant.
- the antibodies that specifically bind to BCMA and CD28 may be administered in combination with T-cell activating anti-CD3 bispecific antibodies.
- the T-cell activating anti-CD3 bispecific antibodies are specific for a tumor-associated antigen, for example GPRC5D, CD38, FcRH5 or BCMA.
- the T-cell activating anti-CD3 bispecific antibody is an anti-GPRC5D/anti-CD3 bispecific antibody.
- the anti-GPRC5D/anti-CD3 bispecific antibody comprises at least one antigen binding domain capable of specific binding to GPRC5D which comprises a heavy chain variable region (VHGPRC5D) comprising (i) CDR-H1 comprising the amino acid sequence of SEQ ID NO: 145, (ii) CDR-H2 comprising the amino acid sequence of SEQ ID NO: 146, and (iii) CDR-H3 comprising the amino acid sequence of SEQ ID NO: 147, and a light chain variable region (VLGPRC5D) comprising (iv) CDR-L1 comprising the amino acid sequence of SEQ ID NO: 148, (v) CDR-L2 comprising the amino acid sequence of SEQ ID NO: 149, and (vi) CDR-L3 comprising the amino acid sequence of SEQ ID NO: 150.
- VHGPRC5D heavy chain variable region
- VLGPRC5D light chain variable region
- the anti-GPRC5D/anti- CD3 bispecific antibody comprises at least one antigen binding domain capable of specific binding to GPRC5D which comprises a heavy chain variable region (VHGPRC5D) comprising an amino acid sequence selected from the group consisting of SEQ ID NO: 151, SEQ ID NO: 153, SEQ ID NO: 154 and SEQ ID NO: 155 and a light chain variable region (VLGPRC5D) comprising an amino acid sequence selected from the group consisting of SEQ ID NO: 152, SEQ ID NO: 156, SEQ ID NO: 157, SEQ ID NO: 158, SEQ ID NO: 159 and SEQ ID NO:160.
- VHGPRC5D heavy chain variable region
- VLGPRC5D light chain variable region
- the anti-GPRC5D/anti-CD3 bispecific antibody comprises at least one antigen binding domain capable of specific binding to GPRC5D which comprises a heavy chain variable region (VHGPRC5D) comprising an amino acid sequence that is at least about 95%, 96%, 97%, 98%, 99% or 100% identical to the amino acid sequence of SEQ ID NO: 151, and a light chain variable region (VLGPRC5D) comprising an amino acid sequence that is at least about 95%, 96%, 97%, 98%, 99% or 100% identical to the amino acid sequence of SEQ ID NO: 152.
- VHGPRC5D heavy chain variable region
- VLGPRC5D light chain variable region
- the antigen binding domain capable of specific binding to GPRC5D comprises a heavy chain variable region (VHGPRC5D) comprising the amino acid sequence of SEQ ID NO: 151 and a light chain variable region (VLGPRC5D) comprising the amino acid sequence of SEQ ID NO: 152.
- VHGPRC5D heavy chain variable region
- VLGPRC5D light chain variable region
- the anti-GPRC5D/anti-CD3 bispecific antibody comprises at least one antigen binding domain capable of specific binding to GPRC5D which comprises a heavy chain variable region (VHGPRC5D) comprising (i) CDR-H1 comprising the amino acid sequence of SEQ ID NO: 161, (ii) CDR-H2 comprising the amino acid sequence of SEQ ID NO: 162, and (iii) CDR-H3 comprising the amino acid sequence of SEQ ID NO: 163, and a light chain variable region (VLGPRC5D) comprising (iv) CDR-L1 comprising the amino acid sequence of SEQ ID NO: 164, (v) CDR-L2 comprising the amino acid sequence of SEQ ID NO: 165, and (vi) CDR-L3 comprising the amino acid sequence of SEQ ID NO: 166.
- VHGPRC5D heavy chain variable region
- VLGPRC5D light chain variable region
- the anti-GPRC5D/anti- CD3 bispecific antibody comprises at least one antigen binding domain capable of specific binding to GPRC5D which comprises a heavy chain variable region (VHGPRC5D) comprising an amino acid sequence selected from the group consisting of SEQ ID NO: 167, SEQ ID NO: 168, SEQ ID NO: 169, SEQ ID NO: 170, SEQ ID NO: 171 and SEQ ID NO: 172 and a light chain variable region (VLGPRC5D) comprising an amino acid sequence selected from the group consisting of SEQ ID NO: 173, SEQ ID NO: 174, SEQ ID NO: 175, SEQ ID NO: 176 and SEQ ID NO: 177.
- the anti-GPRC5D/anti-CD3 bispecific antibody comprises at least one antigen binding domain capable of specific binding to GPRC5D which comprises
- VHGPRC5D heavy chain variable region
- VLGPRC5D light chain variable region
- VHGPRC5D heavy chain variable region
- VLGPRC5D light chain variable region
- VHGPRC5D a heavy chain variable region comprising the amino acid sequence of SEQ ID NO: 151 and a light chain variable region (VLGPRC5D) comprising the amino acid sequence of SEQ ID NO: 158, or
- VHGPRC5D heavy chain variable region
- VLGPRC5D light chain variable region
- VHGPRC5D heavy chain variable region
- VLGPRC5D light chain variable region
- the anti-GPRC5D/anti-CD3 bispecific antibody comprises an antigen binding domain capable of specific binding to CD3 which comprises a heavy chain variable region (VHCD3) comprising (i) CDR-H1 comprising the amino acid sequence of SEQ ID NO: 178, (ii) CDR-H2 comprising the amino acid sequence of SEQ ID NO: 179, and (iii) CORED comprising the amino acid sequence of SEQ ID NO: 180, and a light chain variable region (VLCD3) comprising (iv) CDR-L1 comprising the amino acid sequence of SEQ ID NO: 181, (v) CDR-L2 comprising the amino acid sequence of SEQ ID NO: 182, and (vi) CDR-L3 comprising the amino acid sequence of SEQ ID NO: 183.
- VHCD3 heavy chain variable region
- the anti-GPRC5D/anti-CD3 bispecific antibody comprises an antigen binding domain capable of specific binding to CD3 which comprises a heavy chain variable region (VHCD3) comprising an amino acid sequence of SEQ ID NO: 184 and a light chain variable region (VLCD3) comprising an amino acid sequence of SEQ ID NO:185.
- VHCD3 heavy chain variable region
- VLCD3 light chain variable region
- the anti-GPRC5D/anti-CD3 bispecific antibody comprises an antigen binding domain capable of specific binding to CD3 which comprises a heavy chain variable region (VHCD3) comprising (i) CDR-H1 comprising the amino acid sequence of SEQ ID NO: 186, (ii) CDR-H2 comprising the amino acid sequence of SEQ ID NO: 187, and (iii) CORED comprising the amino acid sequence of SEQ ID NO: 188, and a light chain variable region (VLCD3) comprising (iv) CDR-L1 comprising the amino acid sequence of SEQ ID NO: 189, (v) CDR-L2 comprising the amino acid sequence of SEQ ID NO: 190, and (vi) CDR-L3 comprising the amino acid sequence of SEQ ID NO: 191.
- VHCD3 heavy chain variable region
- CDR-H1 comprising the amino acid sequence of SEQ ID NO: 186
- CDR-H2 comprising the amino acid sequence of SEQ ID NO: 187
- CORED compris
- the anti-GPRC5D/anti-CD3 bispecific antibody comprises an antigen binding domain capable of specific binding to CD3 which comprises a heavy chain variable region (VHCD3) comprising an amino acid sequence of SEQ ID NO: 192 and a light chain variable region (VLCD3) comprising an amino acid sequence of SEQ ID NO:193.
- VHCD3 heavy chain variable region
- VLCD3 light chain variable region
- the anti-GPRC5D/anti-CD3 bispecific antibody comprises an antigen binding domain capable of specific binding to CD3 which comprises a heavy chain variable region (VHCD3) comprising an amino acid sequence of SEQ ID NO: 184 and a light chain variable region (VLCD3) comprising an amino acid sequence of SEQ ID NO: 185, and two antigen binding domains capable of specifically binding to GPRC5D, wherein the antigen binding domains capable of specifically binding to GPRC5D each comprise a heavy chain variable region (VHGPRC5D) comprising the amino acid sequence of SEQ ID NO: 151 and a light chain variable region (VLGPRC5D) comprising the amino acid sequence of SEQ ID NO: 152.
- VHCD3 heavy chain variable region
- VLCD3 light chain variable region
- the anti-GPRC5D/anti-CD3 bispecific antibody comprises a polypeptide that is at least 95%, 96%, 97%, 98%, or 99% identical to the sequence of SEQ ID NO: 137, two polypeptides that are at least 95%, 96%, 97%, 98%, or 99% identical to the sequence of SEQ ID NO: 138, a polypeptide that is at least 95%, 96%, 97%, 98%, or 99% identical to the sequence of SEQ ID NO: 139, and a polypeptide that is at least 95%, 96%, 97%, 98%, or 99% identical to the sequence of SEQ ID NO: 140.
- the bispecific antibody comprises a polypeptide sequence of SEQ ID NO: 137, two polypeptide sequences of SEQ ID NO: 138, a polypeptide sequence of SEQ ID NO: 139 and a polypeptide sequence of SEQ ID NO: 140 (GPRC5D CD3 TCB).
- the anti- GPRC5D/anti-CD3 bispecific antibody is forimtamig.
- the anti-GPRC5D/anti-CD3 bispecific antibody comprises a polypeptide that is at least 95%, 96%, 97%, 98%, or 99% identical to the sequence of SEQ ID NO: 141, a polypeptide that is at least 95%, 96%, 97%, 98%, or 99% identical to the sequence of SEQ ID NO: 142, a polypeptide that is at least 95%, 96%, 97%, 98%, or 99% identical to the sequence of SEQ ID NO: 143, and a polypeptide that is at least 95%, 96%, 97%, 98%, or 99% identical to the sequence of SEQ ID NO: 144.
- the bispecific antibody comprises a polypeptide sequence of SEQ ID NO: 141, a polypeptide sequence of SEQ ID NO: 142, a polypeptide sequence of SEQ ID NO: 143 and a polypeptide sequence of SEQ ID NO: 144 (GPRC5D CD3 1+1 bispecific antibody).
- the T-cell activating anti-CD3 bispecific antibody is an anti -BCM A/anti - CD3 bispecific antibody.
- the anti-BCMA/anti-CD3 bispecific antibody comprises an amino acid sequence of SEQ ID NO: 198, two amino acid sequences of SEQ ID NO: 199, an amino acid sequence of SEQ ID NO: 200 and amino acid sequence of SEQ ID NO: 201.
- the anti-BCMA/anti-CD3 bispecific antibody comprises an amino acid sequence of SEQ ID NO: 202, an amino acid sequence of SEQ ID NO: 203, an amino acid sequence of SEQ ID NO: 204 and amino acid sequence of SEQ ID NO: 205.
- the anti- BCMA/anti-CD3 bispecific antibody comprises an amino acid sequence of SEQ ID NO: 206, an amino acid sequence of SEQ ID NO: 207, an amino acid sequence of SEQ ID NO: 208 and amino acid sequence of SEQ ID NO: 209.
- the anti-BCMA/anti-CD3 bispecific antibody is selected from the group consisting of Alnuctamab, Elranatamab and Teclistamab.
- the T-cell activating anti-CD3 bispecific antibody is an anti- FcRH5/anti-CD3 bispecific antibody.
- Anti-FcRH5/anti-CD3 bispecific antibodies are for instance described in WO 2016/205520.
- Such combination therapies noted above encompass combined administration (where two or more therapeutic agents are included in the same or separate formulations), and separate administration, in which case, administration of the therapeutic agent can occur prior to, simultaneously, and/or following, administration of an additional therapeutic agent or agents.
- administration of the therapeutic agent and administration of an additional therapeutic agent occur within about one month, or within about one, two or three weeks, or within about one, two, three, four, five, or six days, of each other.
- an article of manufacture containing materials useful for the treatment, prevention and/or diagnosis of the disorders described above comprises a container and a label or package insert on or associated with the container.
- Suitable containers include, for example, bottles, vials, syringes, IV solution bags, etc.
- the containers may be formed from a variety of materials such as glass or plastic.
- the container holds a composition which is by itself or combined with another composition effective for treating, preventing and/or diagnosing the condition and may have a sterile access port (for example the container may be an intravenous solution bag or a vial having a stopper that is pierceable by a hypodermic injection needle).
- At least one active agent in the composition is an antibody that specifically binds to BCMA and CD28 as disclosed herein.
- the label or package insert indicates that the composition is used for treating the condition of choice.
- the article of manufacture may comprise (a) a first container with a composition contained therein, wherein the composition comprises an antibody that specifically binds to BCMA and CD28; and (b) a second container with a composition contained therein, wherein the composition comprises a further cytotoxic or otherwise therapeutic agent.
- the article of manufacture in this embodiment of the invention may further comprise a package insert indicating that the compositions can be used to treat a particular condition.
- the article of manufacture may further comprise a second (or third) container comprising a pharmaceutically-acceptable buffer, such as bacteriostatic water for injection (BWFI), phosphate-buffered saline, Ringer's solution and dextrose solution. It may further include other materials desirable from a commercial and user standpoint, including other buffers, diluents, filters, needles, and syringes.
- BWFI bacteriostatic water for injection
- phosphate-buffered saline such as bacteriostatic water for injection (BWFI), phosphate-buffered saline, Ringer's solution and dextrose solution.
- BWFI bacteriostatic water for injection
- phosphate-buffered saline such as bacteriostatic water for injection (BWFI), phosphate-buffered saline, Ringer's solution and dextrose solution.
- BWFI bacteriostatic water for injection
- Ringer's solution such as phosphate
- An antibody that specifically binds to B cell maturation agent (BCMA), wherein the antibody comprises a first antigen binding domain comprising
- VH BCMA heavy chain variable region
- VH BCMA heavy chain variable region comprising a heavy chain complementary determining region CDR-H1 of SEQ ID NO: 1 (GYTFTNYWMH), a CDR-H2 of SEQ ID NO: 2 (IIHPNSGSTNYNEKFQG), and a CDR-H3 of SEQ ID NO: 3 (GIYDYPFAY), and
- VLBCMA light chain variable region
- VL comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 4 (RASES VSIHGTHLMH), a CDR-L2 of SEQ ID NO: 5 (AASSLQS) and a CDR-L3 of SEQ ID NO: 6 (QQSIEDPYT); or
- VL comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 4 (RASES VSIHGTHLMH), a CDR-L2 of SEQ ID NO: 7 (AASNLES) and a CDR-L3 of SEQ ID NO: 6 (QQSIEDPYT), or
- VL comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 4 (RASES VSIHGTHLMH), a CDR-L2 of SEQ ID NO: 8 (AASNLQS) and a CDR-L3 of SEQ ID NO: 6 (QQSIEDPYT).
- VH BCMA comprises an amino acid sequence selected from the group consisting of SEQ ID NO: 9 (VHla) and SEQ ID NO: 10 (VHlb)
- VL BCMA comprises an amino acid sequence selected from the group consisting of SEQ ID NO: 11 (VLlf) , SEQ ID NO: 12 (VLla), SEQ ID NO: 13 (VLlb), SEQ ID NO: 14 (VLlc), SEQ ID NO: 15 (VLld), and SEQ ID NO: 16 (VLle).
- VH BCMA comprising an amino acid sequence of SEQ ID NOV
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 11, or
- VH BCMA comprising an amino acid sequence of SEQ ID NOV
- VLBCMA comprising an amino acid sequence of SEQ ID NO: 12.
- the antibody of any one of paras 1 to 4 comprising an Fc domain composed of a first and a second subunit.
- the second antigen binding domain that specifically binds to a second antigen is a Fab molecule wherein the variable domains VL and VH or the constant domains CL and CHI, particularly the variable domains VL and VH, of the Fab light chain and the Fab heavy chain are replaced by each other.
- the first antigen binding domain is a Fab molecule wherein in the constant domain CL the amino acid at position 123 (numbering according to Kabat EU index) is substituted by an amino acid selected from lysine (K), arginine (R) or histidine (H) and the amino acid at position 124 (numbering according to Kabat EU index) is substituted independently by lysine (K), arginine (R) or histidine (H), and wherein in the constant domain CHI the amino acid at position 147 (numbering according to Kabat EU index) is substituted independently by glutamic acid (E) or aspartic acid (D) and the amino acid at position 213 (numbering according to Kabat EU index) is substituted independently by glutamic acid (E), or aspartic acid (D) (numbering according to Kabat EU index).
- the antibody of para 15, wherein the second antigen binding domain that specifically binds to CD28 comprises a heavy chain variable region (VHCD28) comprising a heavy chain complementary determining region CDR-H1 of SEQ ID NO: 17, a CDR-H2 of SEQ ID NO: 18, and a CDR-H3 of SEQ ID NO: 19, and a light chain variable region (VLCD28) comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 20, a CDR-L2 of SEQ ID NO: 21 and a CDR-L3 of SEQ ID NO: 22. 17.
- VHCD28 heavy chain variable region
- VLCD28 light chain variable region
- the antibody of paras 15 or 16, wherein the second antigen binding domain that specifically binds to CD28 comprises a heavy chain variable region (VHCD28) comprising an amino acid sequence of SEQ ID NO:23, and a light chain variable region (VLCD28) comprising an amino acid sequence of SEQ ID NO:24 (v8).
- VHCD28 heavy chain variable region
- VLCD28 light chain variable region
- the antibody of any one of paras 1 to 18, comprising a first light chain comprising the amino acid sequence of SEQ ID NO:25, a first heavy chain comprising the amino acid sequence of SEQ ID NO:26, a second heavy chain comprising the amino acid sequence of SEQ ID NO:27 and a second light chain comprising the amino acid sequence of SEQ ID NO:28.
- VH BCMA heavy chain variable region
- VH comprising a heavy chain complementary determining region CDR-H1 of SEQ ID NO: 29 (GFTFSNAWMD), a CDR-H2 of SEQ ID NO: 30 (QITAKSNNYATYYADSVKG), and a CDR-H3 of SEQ ID NO: 31 (DGYH), and
- VH comprising a heavy chain complementary determining region CDR-H1 of SEQ ID NO: 29 (GFTFSNAWMD), a CDR-H2 of SEQ ID NO: 32 (QITAKSNNYATYYAAPVKG), and a CDR-H3 of SEQ ID NO: 31 (DGYH), and
- VLBCMA light chain variable region
- VH BCMA comprises an amino acid sequence selected from the group consisting of SEQ ID NO: 36 (VH2a) and SEQ ID NO: 38 (VHlb), and/or the VL BCMA comprises an amino acid sequence selected from the group consisting of SEQ ID NO: 37 (VL2a) and SEQ ID NO:39 (VLla).
- VH BCMA comprising an amino acid sequence of SEQ ID NO:36 and a VL BCMA comprising an amino acid sequence of SEQ ID NO:37
- VH BCMA comprising an amino acid sequence of SEQ ID NO:38 and a VLBCMA comprising an amino acid sequence of SEQ ID NO:39.
- the second antigen binding domain that specifically binds to a second antigen is a Fab molecule wherein the variable domains VL and VH or the constant domains CL and CHI, particularly the variable domains VL and VH, of the Fab light chain and the Fab heavy chain are replaced by each other.
- the first antigen binding domain is a Fab molecule wherein in the constant domain CL the amino acid at position 123 (numbering according to Kabat EU index) is substituted by an amino acid selected from lysine (K), arginine (R) or histidine (H) and the amino acid at position 124 (numbering according to Kabat EU index) is substituted independently by lysine (K), arginine (R) or histidine (H), and wherein in the constant domain CHI the amino acid at position 147 (numbering according to Kabat EU index) is substituted independently by glutamic acid (E) or aspartic acid (D) and the amino acid at position 213 (numbering according to Kabat EU index) is substituted independently by glutamic acid (E), or aspartic acid (D) (numbering according to Kabat EU index).
- the antibody of any one of paras 24 to 32, wherein the Fc domain is of human IgGl subclass and comprises the amino acid mutations L234A, L235A and P329G (numbering according to Kabat EU index).
- the antibody of any one of paras 20 to 33, wherein the second antigen binding domain that specifically binds to CD28 comprises a heavy chain variable region (VHCD28) comprising a heavy chain complementary determining region CDR-H1 of SEQ ID NO: 17, a CDR-H2 of SEQ ID NO: 18, and a CDR-H3 of SEQ ID NO: 19, and a light chain variable region (VLCD28) comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 20, a CDR-L2 of SEQ ID NO: 21 and a CDR-L3 of SEQ ID NO: 22.
- VHCD28 heavy chain variable region
- VLCD28 light chain variable region
- the antibody of para 34, wherein the second antigen binding domain that specifically binds to CD28 comprises a heavy chain variable region (VHCD28) comprising an amino acid sequence of SEQ ID NO:23, and a light chain variable region (VLCD28) comprising an amino acid sequence of SEQ ID NO: 24.
- VHCD28 heavy chain variable region
- VLCD28 light chain variable region
- the antibody of any one of paras 20 to 36 comprising a first light chain comprising the amino acid sequence of SEQ ID NO:40, a first heavy chain comprising the amino acid sequence of SEQ ID NO:41, a second heavy chain comprising the amino acid sequence of SEQ ID NO:27 and a second light chain comprising the amino acid sequence of SEQ ID NO:28.
- a host cell comprising the polynucleotide(s) of para 38 or the vector(s) of para 39.
- a method of producing an antibody that specifically binds to BCMA comprising the steps of a) culturing the host cell of para 40 under conditions suitable for the expression of the antibody and optionally b) recovering the antibody.
- a pharmaceutical composition comprising the antibody of any one of paras 1 to 37 or 42 and at least one pharmaceutically acceptable excipient.
- T-cell activating anti-CD3 bispecific antibody is an anti-GPRC5D/anti-CD3 antibody.
- a method of treating a disease, particularly cancer, in an individual comprising administering to said individual an effective amount of the antibody of paras 1 to 37 or 42, or the pharmaceutical composition of para 43.
- DNA sequencing DNA sequences were determined by double strand sequencing.
- Desired gene segments were either generated by PCR using appropriate templates or were synthesized at Geneart AG (Regensburg, Germany) or Genscript (New Jersey, USA) from synthetic oligonucleotides and PCR products by automated gene synthesis.
- the gene segments flanked by singular restriction endonuclease cleavage sites were cloned into standard cloning / sequencing vectors.
- the plasmid DNA was purified from transformed bacteria and concentration determined by UV spectroscopy.
- the DNA sequence of the subcloned gene fragments was confirmed by DNA sequencing.
- Gene segments were designed with suitable restriction sites to allow subcloning into the respective expression vectors. All constructs were designed with a 5 ’-end DNA sequence coding for a leader peptide which targets proteins for secretion in eukaryotic cells.
- IgG and bispecific antibodies The DNA sequences encoding the variable heavy and light chain regions of the BCMA antibodies (and, where applicable, the CD28 antibodies) were cloned into mammalian expression vectors using conventional cloning techniques. The antibodies described herein were produced using shaking flasks with FedBatch mode. The recombinant production was performed by transient transfection of Expi293TM Cells in a defined, serum-free medium. For transfection ExpiFectamineTM 293 Transfection Kit was used (Gibco). Cell culture supernatants were harvested 7-12 days after transfection.
- Quantification of protein titer The protein titer of supernatant samples was determined by affinity chromatography using a POROS A 20 pm column, 2.1 x 30 mm (Life Technologies, Carlsbad, CA, USA) on a High Performance Liquid Chromatography system (Ultimate 3000 HPLC system, Thermo Scientific, Waltham, MA, USA). The supernatant was loaded onto the column equilibrated with 0.2 M Na2HPO4, pH 7.4, followed by elution with 0.1 M citric acid, 0.2 M NaCl, pH 2.5. Titers were quantified by measuring absorption at 280 nm, and subsequently calculating the protein concentration by comparing the elution peak area (under the curve) of the analyte with a reference standard curve.
- quantification of Fc containing constructs in supernatants was performed by Protein A - HPLC on an Agilent HPLC System with UV detector. Supernatants are injected on POROS A 20 pm (Applied Biosystems), washed with 10 mM Tris, 50 mM Glycine, 100 mM NaCl, pH 8.0 and eluted in the same buffer at pH 2.0. Titers were quantified by measuring absorption at 280 nm, and subsequently calculating the protein concentration by comparing the elution peak area (under the curve) of the analyte with a reference standard curve.
- Proteins were purified from cell culture supernatants referring to standard protocols.
- Fc containing proteins were purified from cell culture supernatants by Protein A-affinity chromatography (equilibration buffer: 20 mM sodium citrate, 20 mM sodium phosphate, pH 7.5 or PBS; elution buffer: 20 mM, 25 mM or 50 mM sodium citrate, pH 3.0). Elution was achieved at pH 3.0 followed by immediate pH neutralization of the sample.
- the protein was concentrated by centrifugation (Millipore Amicon® ULTRA-15, #UFC903096), and aggregated protein was separated from monomeric protein by size exclusion chromatography in 20 mM histidine, 140 mM sodium chloride, pH 6.0.
- the concentrations of purified proteins were determined by measuring the absorption at 280 nm using the mass extinction coefficient calculated on the basis of the amino acid sequence according to Pace, et al., Protein Science, 1995, 4, 2411-1423. Purity and molecular weight of the proteins were analyzed by CE-SDS in the presence and absence of a reducing agent using a LabChipGXII or LabChip GX Touch (Perkin Elmer).
- VH/VL CrossMabs VH/VL CrossMabs
- ESI-MS electrospray ionization mass spectrometry
- the VH/VL CrossMabs were deglycosylated with N-Glycosidase F in a phosphate or Tris buffer at 37°C for up to 17 h at a protein concentration of 1 mg/ml.
- the plasmin or limited LysC (Roche) digestions were performed with 100 pg deglycosylated VH/VL CrossMabs in a Tris buffer pH 8 at room temperature for 120 hours and at 37°C for 40 min, respectively. Prior to mass spectrometry the samples were desalted via HPLC on a Sephadex G25 column (GE Healthcare). The total mass was determined via ESI-MS on a maXis 4G UHR-QTOF MS system (Bruker Daltonik) equipped with a TriVersa NanoMate source (Advion).
- Binding of the generated antibodies to the respective antigens is investigated by surface plasmon resonance using a BIACORE instrument (GE Healthcare Biosciences AB, Uppsala, Sweden). Briefly, for affinity measurements Goat- Anti -Human IgG, JTR 109-005-098 antibodies are immobilized on a CM5 chip via amine coupling for presentation of the antibodies against the respective antigen. Binding is measured in HBS buffer (HBS-P (10 mM HEPES, 150 mM NaCl, 0.005% Tween 20, ph 7.4), 25°C (or alternatively at 37°C).
- HBS buffer HBS-P (10 mM HEPES, 150 mM NaCl, 0.005% Tween 20, ph 7.4
- Antigen (R&D Systems or in house purified) was added in various concentrations in solution. Association was measured by an antigen injection of 80 seconds to 3 minutes; dissociation was measured by washing the chip surface with HBS buffer for 3 - 10 minutes and a KD value was estimated using a 1 : 1 Langmuir binding model. Negative control data (e.g. buffer curves) are subtracted from sample curves for correction of system intrinsic baseline drift and for noise signal reduction. The respective Biacore Evaluation Software is used for analysis of sensorgrams and for calculation of affinity data.
- Anti-BCMA antibody E04 is disclosed in WO 2012/163805 and has the VH domain of SEQ ID NO:42 and the VL domain of SEQ ID NO:43. Optimized variants thereof were created as described in the following.
- a suitable human acceptor framework during the humanization, a combination of two methodologies was used. On the one hand, a classical approach was taken by querying a BLASTp database of human V- and J-region sequences for the murine input sequences (cropped to the variable part).
- Selective criteria for the choice of human acceptor framework were sequence homology, same or similar CDR lengths, and the estimated frequency of the human germline, but also the conservation of certain amino acids at the VH-VL domain interface.
- the CDRs of the murine input sequences were grafted onto the human acceptor framework regions.
- Each amino acid difference between these initial CDR grafts and the parental antibody was rated for possible impact on the structural integrity of the respective variable region, and “back mutations” towards the parental sequence were introduced whenever deemed appropriate.
- the structural assessment was based on Fv region homology models of both the parental antibody and the humanization variants, created with an in-house antibody structure homology modeling protocol implemented using the BIO VIA Discovery Studio Environment, version 17R2.
- “forward mutations” were included, i.e., amino acid exchanges that change the original amino acid occurring at a given CDR position of the parental binder to the amino acid found at the equivalent position of the human acceptor germline. The aim is to increase the overall human character of the humanization variants (beyond the framework regions) to further reduce the immunogenicity risk.
- the acceptor framework was chosen as described in Table 1 below:
- Post-CDR3 framework regions were adapted from human IGHJ germline IGHJ4*01 (YFDYWGQGTLVTVSS, SEQ ID NO: 194) and human IGKJ germline IGKJ4*01 (LTFGGGTKVEIK, SEQ ID NO: 195).
- the part relevant for the acceptor framework is indicated as underlined.
- forward mutations with f e.g., bS49A refers to a back mutation (human germline amino acid to parental antibody amino acid) from serine to alanine at position 49. All residue indices are given in Kabat numbering. 1.1.3 T-cell Epitope Prediction
- the NetMHCIIpan 4.0 predictor (Reynisson B et. al: NetMHCpan-4.1 and NetMHCIIpan-4.0: improved predictions of MHC antigen presentation by concurrent motif deconvolution and integration of MS MHC eluted ligand data, Nucl. Acids Res., 48(W1): W449-W454 (2020)) was employed.
- the thresholds for strong and weak binding 15mer peptides were set to a percentile rank of 1 and 5, respectively. Binding 15mer peptides with a percentile rank above 5 were not considered. Likewise, all binding 15mer peptides with a 9mer core peptide occurring in 10 or more human V-region germlines were not considered.
- the germline sequences were obtained from the IMGT database (Giudicelli, V. et al.: IMGT/LIGM-DB, the IMGT® comprehensive database of immunoglobulin and T cell receptor nucleotide sequences. Nucl. Acids Res., 34(S1):D781-D784 (2006)). As many of the predicted 15mer binders are sharing the same 9mer core peptide, Table 3 below also details the number of unique 9mer cores present in the respective sequence and predicted to bind in the percentile rank range ⁇ 5.
- the resulting VH domains of the humanized BCMA antibodies can be found in Table 4 below and the resulting VL domains of the humanized BCMA antibodies are listed in Table 5 below.
- the humanized amino acid sequences for heavy and light chain variable domains of E04 humanized variants were fused to a one-armed human IgGl backbone/human CH1-Hinge-CH2- CH3 with an effector silent Fc domain (P329G; L234A, L235A) to abrogate binding to Fey receptors according to the method described in WO 2012/130831 Al and into a light chain.
- an effector silent Fc domain P329G; L234A, L235A
- Anti-BCMA antibody 54 is disclosed in WO 2013/072415 and has the VH domain of SEQ ID NO:44 and the VL domain of SEQ ID NO:45.
- BCMA-54 is a humanized antibody with 84.8% identity to the most similar human HV germline (IGHV3-15*01) and 83.2% identity to the most similar human KV germline (IGKV1-6*O1).
- the variable region of BCMA-54 is based on framework regions of human origin, there are several positions in these regions which are not corresponding to the human germline amino acid. Examples for this include the positions VH-16 (Ala), VH-44 (Arg), VH-84 (Lys), VL-22 (Ala), VL-83 (Glu), and VL-95 (He).
- VH-16 Al
- VH-44 Arg
- VH-84 Lys
- VL-22 Al
- VL-83 Glu
- VL-95 He
- suitable human acceptor frameworks were identified by querying a BLASTp database of human V- and J-region sequences for the original BCMA-54 sequences. Selective criteria for the choice of human acceptor framework were sequence homology, same or similar CDR lengths, and the estimated frequency of the human germline, but also the conservation of certain amino acids at the VH-VL domain interface. Following the germline identification step, the CDRs of the BCMA-54 input sequences were grafted onto the human acceptor framework regions. Each amino acid difference between these initial CDR grafts and the parental antibodies was rated for possible impact on the structural integrity of the respective variable region, and “back mutations” towards the parental sequence were introduced whenever deemed appropriate.
- Acceptor framework Post-CDR3 framework regions were adapted from human IGHJ germline IGHJ 1*01 (AEYFOHWGQGTLVTVSS, SEQ ID NO: 196) and human IGKJ germline IGKJ2*01 (YTF GQGTK LE IK, SEQ ID NO: 197). The part relevant for the acceptor framework is indicated as underlined.
- Back mutations are prefixed with b, forward mutations with f, e.g., bS49A refers to a back mutation (human germline amino acid to parental antibody amino acid) from serine to alanine at position 49. All residue indices are given in Kabat numbering..
- the NetMHCIIpan 4.0 predictor (Reynisson B et. al: NetMHCpan-4.1 and NetMHCIIpan-4.0: improved predictions of MHC antigen presentation by concurrent motif deconvolution and integration of MS MHC eluted ligand data, Nucl. Acids Res., 48(W1): W449-W454 (2020)) was employed.
- the thresholds for strong and weak binding 15mer peptides were set to a percentile rank of 1 and 5, respectively. Binding 15mer peptides with a percentile rank above 5 were not considered. Likewise, all binding 15mer peptides with a 9mer core peptide occurring in 10 or more human V-region germlines were not considered.
- the germline sequences were obtained from the IMGT database (Giudicelli, V. et al.: IMGT/LIGM-DB, the IMGT® comprehensive database of immunoglobulin and T cell receptor nucleotide sequences. Nucl. Acids Res., 34(S1):D781-D784 (2006)). As many of the predicted 15mer binders are sharing the same 9mer core peptide, Table 8 below also details the number of unique 9mer cores present in the respective sequence and predicted to bind in the percentile rank range ⁇ 5.
- VH domains of the humanized BCMA antibodies can be found in Table 9 below and the resulting VL domains of the humanized BCMA antibodies are listed in Table 10 below.
- Table 9 Amino acid sequences of the VH domains of humanized BCMA antibodies
- the humanized amino acid sequences for heavy and light chain variable domains of E04 humanized variants were fused to a one-armed human IgGl backbone/human CH1-Hinge-CH2- CH3 with an effector silent Fc domain (P329G; L234A, L235A) to abrogate binding to Fey receptors according to the method described in WO 2012/130831 Al and containing a knob mutation according to Knobs-into-Holes technology and into a light chain.
- a human Fc containing an effector silent Fc domain was used for the right assembly of the one-armed IgGl .
- the amino acid sequences were backtranslated into DNA and the resulting cDNA were synthesized (GeneArt or Twist Biosciences) and then cloned into heavy chain expression vectors as fusion proteins with human IgGl backbones expression vectors as fusion proteins to human C-kappa.
- Light chain (LC) and heavy chain (HC) plasmids were then co-transfected into HEK293 cells and purified after 7 days from supernatants by standard methods for antibody purification.
- the protein concentration of purified constructs was determined by measuring the optical density (OD) at 280 nm, using the mass extinction coefficient calculated on the basis of the amino acid sequence according to Pace, et al., Protein Science, 1995, 4, 2411-1423. Purity and molecular weight of the proteins were analyzed by CE-SDS in the presence and absence of a reducing agent using a LabChipGXII (Perkin Elmer).
- the binding kinetics of the monovalent humanized BCMA antibody variants to human BCMA and to cynomolgus BCMA were investigated by surface plasmon resonance (SPR) using a BIACORE T200 instrument (GE Healthcare). All experiments were performed at 25° C using HBS-P Buffer (10 mM HEPES, 150 mM NaCl pH 7.4, 0.05% surfactant P20) as running buffer and as dilution buffer.
- Anti-BCMA antibodies were captured on the surface for 30 s at a flow rate of 5 pl/min leading to a capturing response of 50-200 RU.
- a dilution series of the antigen human BCMA Fc homodimer (R&D Systems) or cynomolgus BCMA Fc homodimer (R&D Systems), respectively
- the dissociation phase was monitored for 300-600 sec by washing with running buffer.
- the surface was regenerated by injecting 5 mM NaOH (freshly prepared) for 2 x 30 s.
- the KD values for the monovalent constructs including the 29 preferred BCMA variants are shown in Table 12 below.
- Table 12 Production, Purification and Binding Properties expressed monovalent anti- BCMA variants Based on the results of the in-silico assessment of the occurrence of potential T-cell epitopes in the humanized sequences with the NetMHCIIpan 4.0 predictor (see Example 1.2.3), the antibodies P1AG5080, P1AG5072, P1AG5028, P1AG5031, P1AG5063 and P1AG5036 (4 BCMA 54 variants and 2 BCMA E04 variants) were selected as the molecules with the lowest potential T-cell epitopes. In combination with their binding behavior, Pl AG5072 and Pl AG5031 were choosen as BCMA 54 variants and Pl AG5063 and Pl AG5036 were selected as the BCMA E04 variant antibodies to be included in the bispecific antibodies.
- the sequences of the respective variable domains were used and sub-cloned in frame with the respective constant regions which are preinserted in the respective recipient mammalian expression vector.
- Pro329Gly, Leu234Ala and Leu235Ala mutations (PG-LALA) have been introduced in the constant region of the human IgGl heavy chains to abrogate binding to Fc gamma receptors according to the method described in International Patent Appl. Publ. No. WO 2012/130831.
- Fc fragments contained either the “knob” (S354C/T366W mutations, numbering according to Kabat EU index) or “hole” mutations (Y349C/T366S/L368A/Y407V mutations according to Kabat EU index) to avoid mispairing of the heavy chains.
- “knob” S354C/T366W mutations, numbering according to Kabat EU index
- “hole” mutations Y349C/T366S/L368A/Y407V mutations according to Kabat EU index
- exchange of VH/VL or CHl/Ckappa domains was introduced in one binding moiety (CrossFab technology).
- charges were introduced into the CHI and Ckappa domains as described in International Patent Appl. Publ. No. WO 2015/150447.
- CD28 v.8 has a VH of SEQ ID NO:23 and a VL of SEQ ID NO:24.
- CD28 v.15 has a VH of SEQ ID NO: 90 and a VL of SEQ ID NO:91.
- FIG. 1C Schematic illustrations of the bispecific antibody structures are shown in Figures IB or 1C.
- Table 13 summarizes specific anti-BCMA/anti-CD28 bispecific antibodies made, their identifiers and the sequences of the heavy chains (HC1 knob and HC2 hole) and light chains (LC1 and LC2).
- the DNA sequences encoding the variable heavy and light chain regions of the BCMA and CD28 antigen binding domains were cloned into mammalian expression vectors using conventional cloning techniques.
- the bispecific antibodies described herein were produced using shaking flasks with FedBatch mode.
- the recombinant production was performed by transient transfection of Expi293TM Cells in a defined, serum-free medium. For transfection ExpiFectamineTM 293 Transfection Kit was used (Gibco). Cell culture supernatants were harvested 7-12 days after transfection.
- Quantification of protein titer The protein titer of supernatant samples was determined by affinity chromatography using a POROS A 20 pm column, 2.1 x 30 mm (Life Technologies, Carlsbad, CA, USA) on a High Performance Liquid Chromatography system (Ultimate 3000 HPLC system, Thermo Scientific, Waltham, MA, USA). The supernatant was loaded onto the column equilibrated with 0.2 M Na2HPO4, pH 7.4, followed by elution with 0.1 M citric acid, 0.2 M NaCl, pH 2.5. Titers were quantified by measuring absorption at 280 nm, and subsequently calculating the protein concentration by comparing the elution peak area (under the curve) of the analyte with a reference standard curve.
- Proteins were purified from cell culture supernatants referring to standard protocols.
- Fc containing proteins were purified from cell culture supernatants by Protein A-affinity chromatography (equilibration buffer: 20 mM sodium citrate, 20 mM sodium phosphate, pH 7.5 or PBS; elution buffer: 20 mM, 25 mM or 50 mM sodium citrate, pH 3.0). Elution was achieved at pH 3.0 followed by immediate pH neutralization of the sample.
- the protein was concentrated by centrifugation (Millipore Amicon® ULTRA- 15, #UFC903096), and aggregated protein was separated from monomeric protein by size exclusion chromatography in 20 mM histidine, 140 mM sodium chloride, pH 6.0.
- the concentrations of purified proteins were determined by measuring the absorption at 280 nm using the mass extinction coefficient calculated on the basis of the amino acid sequence according to Pace, et al., Protein Science, 1995, 4, 2411-1423. Purity and molecular weight of the proteins were analyzed by CE-SDS in the presence and absence of a reducing agent using a LabChipGXII or LabChip GX Touch (Perkin Elmer).
- Determination of the aggregate content was performed by HPLC chromatography at 25°C using analytical size-exclusion column (TSKgel G3000 SW XL or UP-SW3000, Tosoh Bioscience) equilibrated in running buffer (200 mM KH2PO4, 250 mM KC1 pH 6.2, 0.02% NaNs).
- Table 14 Summary of the production and purification of bispecific CD28 antigen binding molecules
- BCMA-CD28 bsAbs had favoarable pl values in the range of >8 (see Table 15 below).
- Thermal stability of the BCMA-CD28 bsAbs prepared was monitored by Dynamic Light Scattering (DLS) and by monitoring of temperature dependent intrinsic protein fluorescence by applying a temperature ramp using an Optim 2 instrument (Avacta Analytical, UK).
- the apparent hydrophobicity of the bispecific antibodies was assessed by hydrophobic interaction chromatography (HIC) as relative retention time compared to hydrophobicity standards (90 % of immunoglobulins in IVIG preparation have a relative retention time ⁇ 0.35).
- HIC hydrophobic interaction chromatography
- 20 pg of sample was injected onto a HIC-Ether-5PW (Tosoh) column equilibrated with 25 mM Na-phosphate, 1.5 M ammonium sulfate, pH 7.0. Elution was performed with a linear gradient from 0 to 100% buffer B (25 mM Na-phosphate, pH 7.0) within 60 minutes. Retention times were then compared to protein standards with known hydrophobicity (e.g. Avastin).
- FcRn affinity chromatography FcRn was expressed, purified and biotinylated as described (Schlothauer et al., MAbs 2013, 5(4), 576-86).
- the prepared receptor was added to streptavidin-sepharose (GE Healthcare).
- streptavidin-sepharose GE Healthcare
- the resulting FcRn-sepharose matrix was packed in a column housing.
- the column was equilibrated with 20 mM 2-(N-morpholine)- ethanesulfonic acid (MES) and 140 mM NaCl, pH 5.5 (eluent A) at a 0.5 ml/min flow rate.
- MES 2-(N-morpholine)- ethanesulfonic acid
- Heparin affinity was determined by injecting 30-50 pg of sample onto a TSKgel Heparin- 5PW (Tosoh) column equilibrated with 50 mM Tris, pH 7.4. Elution was performed with a linear gradient from 0 to 100% buffer B (50 mM Tris, IM NaCl, pH 7.4 mM) within 37 minutes. Retention times were compared to protein standards with known affinities. The expected values for the retention times and the measured retention times for particular BCMA-CD28 bsAbs are shown in Table 15.
- BCMA-CD28 bispecific antibodies with a concentration of 2 pg/ml were injected for 30 s at a flow rate of 5 pl/min, and dissociation was monitored for 120 s.
- the surface was regenerated by injecting 10 mM glycine buffer, pH 1.5, for 60 s. Bulk refractive index differences were corrected by subtracting blank injections and by subtracting the response obtained from a blank control flow cell. For evaluation, the binding response 5 seconds after injection end was taken.
- the BCMA and CD28 binding was divided by the anti-hu IgG response (the signal (RU) obtained upon capture of the BCMA-CD28 bsAbs on the immobilized anti-hu IgG antibody). The relative binding activity was calculated by referencing each temperature stressed sample to the corresponding, non-stressed sample. As shown in Table 16, all BCMA-CD28 bsAbs showed a stabile binding upon stress to BCMA and CD28.
- Alnuctamab is a BCMA x CD3 bispecific antibody in a 2+1 format, based on an IgGl Fc with L234A/L235A/P329G (EU numbering) mutations.
- the sequences are identified in the International Non-Proprietary Nomenclature List (Recommended INN: List 85; WHO Drug Information, Vol. 35, No. 1, 2021).
- Alnuctamab comprises an amino acid sequence of SEQ ID NO: 198, two amino acid sequences of SEQ ID NO: 199, an amino acid sequence of SEQ ID NO: 200 and amino acid sequence of SEQ ID NO: 201 (2+1 format).
- Elranatamab is a BCMA x CD3 bispecific antibody in a 1+1 format. The sequences are identified in the International Non-Proprietary Nomenclature List (Recommended INN: List 87, WHO Drug Information, Vol. 36, No. 1, 2022). Elranatamab comprises an amino acid sequence of SEQ ID NO: 202, an amino acid sequence of SEQ ID NO: 203, an amino acid sequence of SEQ ID NO: 204 and amino acid sequence of SEQ ID NO: 205.
- Alnuctamab P1AF0105
- the DNA sequences encoding the variable heavy and light chain regions of the respective binding domains were cloned into mammalian expression vectors using conventional cloning techniques.
- Antibodies were generated by transient transfection of Expi293F cells. Cells were seeded in Expi293 media (Gibco, #1435101) at a density of 2.5 x 10 6 /mL.
- Expression plasmids and ExpiFectamine (Gibco, ExpiFectamine transfection kit, #13385544) were separately mixed in OptiMEM (Gibco, #11520386).
- Elranatamab (P1AH5054) was produced and purified by Proteros according to their standard methods and protocols.
- the quantification of Fc containing constructs in supernatants was performed by Protein A - HPLC on an Agilent HPLC System with UV detector. Supernatants are injected on POROS 20 A (Applied Biosystems), washed with 10 mM Tris, 50 mM Glycine, 100 mM NaCl, pH 8.0 and eluted in the same buffer at pH 2.0. Titers were quantified by measuring absorption at 280 nm, and subsequently calculating the protein concentration by comparing the elution peak area (under the curve) of the analyte with a reference standard curve.
- Proteins were purified from filtered cell culture supernatants referring to standard protocols.
- Fc containing proteins were purified from cell culture supernatants by Protein A-affinity chromatography (equilibration buffer: 20 mM sodium citrate, 20 mM sodium phosphate, pH 7.5; elution buffer: 20 mM sodium citrate, pH 3.0). Elution was achieved at pH 3.0 followed by immediate pH neutralization of the sample.
- the protein was concentrated by centrifugation using Millipore Amicon® ULTRA- 15 (Merck, #UFC903096), and aggregated protein was separated from monomeric protein by size exclusion chromatography in 20 mM histidine, 140 mM sodium chloride, pH 6.0.
- the concentrations of purified proteins were determined by measuring the absorption at 280 nm using the mass extinction coefficient calculated on the basis of the amino acid sequence according to Pace, et al., Protein Science, 1995, 4, 2411-1423. Purity and molecular weight of the proteins were analyzed by CE-SDS in the presence and absence of a reducing agent using a LabChipGXII or LabChip GX Touch (Perkin Elmer) (Perkin Elmer).
- Table 16A Summary of the production and purification of bispecific BCMA x CD3 antigen binding molecules
- Teclistamab is a BCMA x CD3 bispecific antibody with an IgG4-F234A/L235A/S228P (EU numbering) Fc. The sequences are identified in the International Non-Proprietary Nomenclature List (Recommended INN: List 82, WHO Drug Information, Vol. 33, No. 3, 2019). It was obtained from a Supplier (FarmaMondo, Lot Nr. AT1334P1). Teclistamab comprises an amino acid sequence of SEQ ID NO: 206, an amino acid sequence of SEQ ID NO: 207, an amino acid sequence of SEQ ID NO: 208 and amino acid sequence of SEQ ID NO: 209.
- CHO-K1 cell lines expressing BCMA extracellular domain non -truncating mutants were generated as follows: Full-length cDNA encoding human BCMA (UniProt: Q02223) and its corresponding mutants (R27P, S30del, P33S and P34del) were subcloned into lentiviral transfer vectors controlled by the CMV promoter.
- Lentiviral particles were prepared by transiently cotransfecting HEK 293-derived viral production cells (Gibco, #A35347) with transfer plasmid and lentiviral packaging mix (pRSV-Rev, pCgpV, and pCMV-VSV-G) using the LV-MAX Transfection Kit (Gibco, # A35346) according to the manufacturer's protocol. Viral supernatants were harvested 48 h after transfection, filtered through a 0.45-pm low protein binding filter and stored at -80°C until use.
- CHO-K1 (ATCC CRL-9618) cells were seeded per well in a 24-well plate. The following day, the culture medium was replaced with 300 pL of purified lentiviral supernatant and 100 pL of fresh culture medium DMEM/F-12 (Gibco, #11320033) supplemented with 10% fetal bovine serum (Gibco, #16140063) and 1% GlutaMAX Supplement (Gibco; #31331-028). To facilitate viral transduction, the infectious medium was further supplemented with 2.5 pL of TransDuxTM reagent and 100 pL of MAX Enhancer (SBI; # LV860A-1). The cells were then incubated for 24 hours at 37 °C. After the incubation period, the viral medium was discarded and the cells were subsequently maintained in fresh culture medium.
- DMEM/F-12 Gibco, #11320033
- fetal bovine serum Gibco, #16140063
- GlutaMAX Supplement Gibco; #3
- the culture medium was supplemented with 6 pg/mL puromycin (Invivogen; #ant-pr-l).
- the cells exhibiting human BCMA surface expression were isolated by BD FACSAria III cell sorter (BD Biosciences) and subsequently cultured to generate stable clones. Following a 4-week stability test, the surface expression and its stability were verified by flow cytometry analysis using mouse PE-conjugated anti-human BCMA (BioLegend, #357503).
- the stable CHO transfectants (parental cell line CHO-kl ATCC #CCL-61) were cultured in DMEM/F-12 (Gibco, #10565018) supplemented with 10 fetal bovine serum (Gibco, #16140063 or Sigma-Aldrich in case of F4135) and 1% GlutaMAX Supplement (Gibco; #31331-028), including 6 pg/ml puromycin (Invivogen; #ant-pr-l).
- Adherent CHO cells were detached using Cell Dissociation Buffer (Gibco, #13151014) or trypsin (Gibco by ThermoFisher Scientific, TrypLETMExpress Enzyme #2605-010), counted and checked for viability. All subsequent steps were performed at 4°C.
- CHO-huCD28 cells were resuspended in FACS buffer (PBS, 2% Fetal Bovine Serum; 1% 0.5 M EDTA pH 8; 0.25% NaNs Sodium azide) at 1 Mio cells per ml.
- FACS buffer PBS, 2% Fetal Bovine Serum; 1% 0.5 M EDTA pH 8; 0.25% NaNs Sodium azide
- 0.1 Mio cells were plated per well of a round-bottom 96-well-plate and washed with 150 pl FACS buffer per well and supernatants were discarded.
- Cells were stained in a total volume of 50 pl per well and increasing concentrations of the indicated BCMA-CD28 bispecific molecules (0.48 pM - 2000 nM) for 60 minutes at 4°C.
- CHO-huBCMA cells were resuspended in FACS buffer (PBS, 2% Fetal Bovine Serum; 1% 0.5 M EDTA pH 8; 0.25% NaNs Sodium azide) at 1 Mio cells per ml.
- FACS buffer PBS, 2% Fetal Bovine Serum; 1% 0.5 M EDTA pH 8; 0.25% NaNs Sodium azide
- 0.1 Mio cells were plated per well of a round-bottom 96-well-plate and washed with 150 pl FACS buffer per well and supernatants were discarded.
- Cells were stained in a total volume of 50 pl per well and increasing concentrations of the indicated BCMA-CD28 bispecific molecules (0.48 pM - 2000 nM) for 30 minutes at 4°C. Afterwards, the cells were centrifuged and washed with 150 pl FACS buffer twice.
- the bispecific antibodies tested comprised CD28 antibody CD28v8 or CD28vl5, respectively.
- Fig. 2A and 3A molecules comprising antibody CD28v8
- Fig. 2B and 3B molecules comprising antibody CD28vl5
- Fig. 3 A and 3B show that all bispecific CD28 molecules are able to bind both, human CD28 (Fig. 2A and 2B), as well as human BCMA (Fig. 3 A and 3B) in a concentration-dependent manner. Binding to human CD28 does not reach saturation due to low affinity binders and is comparable among the CD28v8 bispecific antibodies (Fig. 2A) and CD28vl5 bispecific antibodies (Fig. 2B), respectively.
- the bispecific antibodies tested comprised the CD28 antibody CD28v8 and the BCMA binder BCMA (54 2a2a) or BCMA (E04_lalf). Since one of the relapse mechanisms in MM can be non-truncating, missense mutations or in-frame deletions in the extracellular domain of BCMA, the above mentioned molecules were compared head-to-head to well described BCMA-targeted CD3 T cell engagers Teclistamab, Alnuctamab and Elranatamab (Lee et al., Nature Medicine 2023, 29, 2295-2306).
- CHO transfectants were resuspended in PBS (Gibco by ThermoFisher Scientific, #20012050) and counted. Live-dead staining of target cells was performed by incubation of cells with 1 : 1000 diluted Zombie Aqua Viability dye (BioLegend #423102) in PBS at 1.5 x 10 6 cells per ml for 10 minutes in the dark, followed by a washing step with PBS and centrifugation at 400 x g at 4°C for 4 minutes.
- PBS Gibco by ThermoFisher Scientific, #20012050
- Live-dead staining of target cells was performed by incubation of cells with 1 : 1000 diluted Zombie Aqua Viability dye (BioLegend #423102) in PBS at 1.5 x 10 6 cells per ml for 10 minutes in the dark, followed by a washing step with PBS and centrifugation at 400 x g at 4°C for 4 minutes.
- CHO-K1 cells were adjusted to 1.25 x 10 6 cells per ml in PBS and 40 pl were seeded per well of a 384-U bottom well plate (ThermoFisher Scientific #264573). Plates were centrifuged once more at 400 x g at 4°C for 4 minutes and 20 pl supernatant was removed.
- Cells were stained in a total volume of 40 pl per well by adding 20 pl of 2x concentrated dilution of the indicated BCMA-CD28 antigen binding molecules (final concentrations ranged from 0.008 - 125 nM). Cells were then incubated for 30 minutes at 4°C in the dark. Afterwards, the cells were centrifuged and washed with 40 pl FACS buffer three times.
- BCMA-targeted CD28v8 bispecific molecules were able to bind to the wt and to all mutated forms of human BCMA in a concentration-dependent manner.
- Elranatamab displayed a significant reduction of binding to the point mutation R27P, which translated into a weak binding signal at the highest concentrations only.
- Teclistamab was not able to bind to the point mutation R27P and the S30del.
- Efficient binding to BCMA is a crucial requisite for the therapeutic activity of BCMA-targeted bispecific molecules, suggesting an advantage for molecules who are not affected by point mutations in the ECD of BCMA.
- the difference in the maximal values for the wt versus the mutated BCMA transfectants is due to different expression levels of the human BCMA wt and variants on the transfectants and is consistent for all molecules tested.
- Table 17A High-level summary of binding properties of the indicated BCMA-targeting molecules to either human wildtype (wt) BCMA or the indicated point mutations in the extracellular domain of human BCMA
- BCMA-CD28 bispecific antibodies were assessed in a Jurkat IL2 reporter assay, where the molecules simultaneously bind to human CD28 on Jurkat IL-2 cells and BCMA expressed on the indicated MM cell lines.
- a CD3/TCR-mediated first activation signal the crosslinking of the BCMA-CD28 bispecific antibodies to Jurkat and MM cells induces IL-2 release from activated Jurkat cells which in turn drives the expression of an IL-2 promoter driven luciferase gene, which can be quantified by determination of luciferase, using a conventional microplate reader.
- BCMA-CD28 bispecific antibodies were evaluated in a PBMC co-culture assay in presence of BCMA-expressing MM cell lines and a fixed concentration of an anti-GPRC5D/anti-CD3 bispecific antibody (GPRC5D TCB, P1AE6625) providing the simultaneous 1st signal for T-cell activation. Read-outs were upregulation of surface activation markers on T cells as well as T cell proliferation, as indicated. 3.
- Jurkat IL2 reporter assays were used to address how different BCMA-CD28 bispecific antibodies can co-stimulate T cells after activation by GPRC5D TCB (P1AE6625). The assay has been performed as described in the following:
- NCI-H929 cells were cultivated in RPMI1640 (Gibco) supplemented with 10% FCS (PAN-biotech), 2 mM L-glutamine (Sigma-Aldrich), 1 mM sodium pyruvate (Thermofisher), and 50pM 2-Mercaptoethanol (Thermofisher). Cells were cultured twice per week to maintain a density between 0.5 - 2.0 xlO 6 cells/ml.
- Jurkat IL2 reporter and Jurkat NF- KB/4-1BB reporter cell lines were grown in RPMI1640 medium supplemented with 10% FCS, 25 mM HEPES (Thermofisher), 2 mM L-glutamine, 0.1 mM non-essential amino acid (Thermofisher), and 1 mM sodium pyruvate.
- the medium was supplemented with 200 pg/ml of Hygromycin B (Roche) for Jurkat IL2 reporter cell line, and with 400 pg/ml of Hygromycin B and 600pg/mL of Geneticin (Sigma-Aldrich) for Jurkat NF-KB/4-1BB reporter cell line.
- Jurkat cells were cultured twice per week to maintain a density between 0.1 - 0.5 xlO 6 cells/ml.
- the target (NCI-H929) and effector (Jurkat IL2 reporter) cells were harvested and resuspended in assay medium (Jurkat cell medium without antibiotics) to obtain a cell density of 6 xlO 6 cells/ml for target cells and 3 xlO 6 cells/ml for effector cells. Then cells were mixed at a ratio E:T 1 :2, and 20 pl of effector-target cells mixture were plated per well of a white-walled 384-flat bottom well plate (FalconTM 384-Well White Flat-Bottom Tissue Culture Treated Microplate). Next, 10 pl of titrated amount (200.0-0.05 nM) of BCMA-CD28 bispecific antibody (BsAb) were added to the plate in triplicates.
- BsAb BCMA-CD28 bispecific antibody
- GPRC5D-TCB 10 pl of fixed concentration (2000, 200, or 20 pM) GPRC5D-TCB was added to the plate, reaching the final volume of 40 pl per well.
- three separate control conditions were prepared that contained target and effector cells alone in order to indicate effector cell-induced IL2 signaling without stimulus.
- Target and effector cells with BCMA-CD28 BsAb and without GPRC5D-TCB were added together to address the unspecific activation of effector cells without a first signal.
- Last control, target, and effector cells with GPRC5D-TCB (2000, 200, or 20 pM) and without BCMA-CD28 BsAb were prepared to indicate the baseline of IL2 signaling in the effector cells signal.
- Assay medium was used to reach the final volume (40 pl) for the control wells. Assay plates were centrifuged for 1 min at 350 g and incubated for 24 hrs at 37°C in a humidified CO2 incubator. Assay plates were incubated for 5 min at room temperature (RT) before adding 20 pl of the ONE-Glo solution (Promega). Further, plates were centrifuged for 1 min at 350g and incubated for 10 min at room temperature in the dark to achieve full lysis of the cells. Luminescence was measured (read: 1 s per well) by using the Tecan SparklOM.
- the target NCLH929 cell line expressing high levels of BCMA was mixed at the ratio 1 :2 (E:T) with the effector Jurkat IL 2 reporter cell line that expresses luciferase in an IL2 promoter-dependent fashion. Then, titrated (200.0-0.04 nM) BCMA-CD28 BsAbs together with (2000, 200, or 20 pM GPRC5D-TCB were added and luminescence was measured 24 hrs later. Results showed that all tested BCMA-CD28 BsAbs including CD28 antibody v8 or vl5 induced co-stimulation of T cells in a dose-dependent manner (Fig. 4A-4F).
- Table 18 EC50 values (nM) and Emax (%) for T cell activation as measured in the Jurkat
- Figures 5A to 5D show a summary of the data obtained from Jurkat NFkB reporter assay.
- Fig. 5 A compares the ECso values and Fig. 5B the efficacy of the various BCMA-CD28(vl5) BsAbs
- Fig. 5C the ECso values and Fig. 5D the efficacy of the various BCMA-CD28(v8) BsAbs compared to the untargeted CD28 controls Pl AG0760 and P1AF8794, respectively, and were calculated from three independent experiments performed in triplicate.
- Target Cells NCI-H929 & NCI-H929 BCMAko, as indicated respectively.
- NCI-H929 (ATCC® CRL-9068TM) is a human multiple myeloma cell line that expresses BCMA.
- BCMA knockout variant of the NCI-H929 cell line (generated using CRISPR/Cas9 technology) was tested as well.
- the cells were cultivated in RPMI 1640 (GibcoTM 31870074) supplemented with 10% FCS (GibcoTM 16140-071), 10 mM HEPES (GibcoTM 15630056), 2 mM GlutaMAX-I (GibcoTM 35050-038), 1 mM sodium pyruvate (GibcoTM 11360039), and 50pM 2-Mercapto- ethanol (GibcoTM 31350010). Cells were passaged 2-3 times per week by adding fresh medium to maintain a density between 0.5xl0 6 /ml and 2.5xl0 6 /ml. Cells were incubated at 37 °C with 5% CO2.
- the target cells were harvested, counted and re-suspended in assay medium (RPMI 1640 w/ HEPES, w/GlutaMax GibcoTM 31870074 plus 10% FCS) to obtain a cell density of 1.2 xlO 6 cells/ml.
- assay medium RPMI 1640 w/ HEPES, w/GlutaMax GibcoTM 31870074 plus 10% FCS
- PBMCs Frozen human peripheral blood mononuclear cells (PBMC) from whole human blood in CPD were received from Cambridge bioscience. PBMCs were stored in nitrogen vapor phase and thawed on the day of the assay. Cells were counted and then labeled with Cell Proliferation Dye eFluorTM 450 (65-0842-90, eBioscienceTM ). Briefly, PBMCs were washed once with DPBS (Gibco), supernatant was discarded and cells resuspended to 2 Mio cells per well using DPBS.
- DPBS Gibco
- 50 pl (0.06xl0 6 ) NCI-H929 or NCI-H929 BCMAko cells and 50 pl (0.06xl0 6 PBMCs) were plated per well in 96-well round bottom plates (TPP) resulting in a 1 : 1 ratio.
- 50 pl of titrated amount (0.12 - 500 nM) of BCMA-CD28 bispecific antibody (BsAb) were added to the plate in triplicates.
- 50 pl of fixed concentration (0.32 pM) GPRC5D-TCB (P1AE6625) was added to the plate, reaching the final volume of 200 pl per well.
- three separate control conditions were prepared.
- Target and effector cells with BCMA-CD28 BsAbs and without GPRC5D-TCB were added to address potential activation of effector cells by CD28 bispecific molecules without a first signal.
- target, and effector cells were treated with GPRC5D-TCB alone to indicate the baseline.
- a control with target and effector cells was prepared in which no antibody was added (untreated).
- Assay medium was used to reach the final volume (200 pl) for the control wells. Assay plates were centrifuged for 1 min at 350 g and incubated for 4 days at 37°C in a humidified CO2 incubator.
- the cells were stained to evaluate T cell activation and proliferation.
- the cells were first washed once with 200 pL PBS and then stained with LIVE/DEADTM Fixable Near-IR Dead Cell Stain (1 :500) (ThermoFisher Scientific Catalog No. L34976), FITC antihuman CD4 (clone RPA-T4), BV711 anti -human CD8 (clone RPA-T8), APC anti -human CD25 (clone BC96), PerCP-Cy5.5 anti-human CD 137 (clone 4B4-1), all from BioLegend in PBS for 30 min at 4°C.
- Flow cytometry acquisition was performed on a custom-designed BD Biosciences Fortessa and analyzed using FlowJo software (Tree Star, Ashland, OR) and GraphPad Prism software.
- Tables 19 to 22 summarize the ECso values, as well the maximum from the data shown in Figures 6A to 9A. ECso values were calculated using GraphPadPrism6.
- Table 20 ECso values (nM): CD28 v!5 bispecific antibodies versus BCMA (PR)-CD28 v!5 bispecific antibody
- the therapeutic efficacy of the bispecific BCMA-CD28 antibodies was tested in combination therapy with GPRC5D-TCB in heparinized primary MM patient bone marrow aspirate (BMA) samples obtained within the first 48 hours after extraction. Cell amounts, their viability and phenotype, as well as percent of diseased MM plasma cells at baseline were analyzed by flow cytometry. Next, at least 0.1 million MM plasma cells (MM PCs) were seeded in 24- or 48-well plates. A fixed concentration of a GPRC5D-CD3 bispecific molecule (GPRC5D-TCB, 1 or 10 nM as indicated) was tested alone or in combination with increasing doses of the different BCMA-CD28 bispecific antibodies.
- BMA heparinized primary MM patient bone marrow aspirate
- TCB TCB isotype, respectively an untargeted CD28 molecule was added.
- Sterile PBS was added up to 1 mL.
- Cells were centrifuged at 540 g for 5 min and erythrocytes were lysed for 15 minutes according to standard methods.
- Cells were centrifuged at 800 g for 10 minutes, washed with sterile PBS once and incubated at room temperature for 20 minutes with maleimide to stain dead versus living cells.
- Cells were washed with 12 ml of PBS, including 0.09% of NaNs and 0.5 % of BSA to remove unbound maleimide.
- cell pellets were resuspended in 500 pL PBS and filtered through 5 ml polystyrene round-bottom tubes with cell-strainer cap to discard cell clots and remove potential membrane aggregates. Cells were centrifuged once more at 540 g for 5 minutes. Finally, cells were stained with the indicated antibodies against different surface markers for 20 min at RT protected from light (Table 23 and Table 24), washed as described above and resuspended in sterile PBS. The total sample was acquired by flow.
- Table 23 Antibody combination used for the analysis of pre-treatment sample
- Table 24 Antibody combination used to determine MM-PC lysis and effector cell activation
- the therapeutic efficacy of the bispecific BCMA-CD28 antibodies was tested in combination therapy with GPRC5D-TCB using frozen and thawed primary MM bone marrow mononuclear cells (BMMNCs) as described below.
- BMMNCs primary MM bone marrow mononuclear cells
- samples were thawed and resuspended in StemSpan SFEMII (StemCell) medium, including 20 % human serum, 55 uM P-mercaptoethanol (Gibco cat# 11528926) and 100 U/ml penicillin and 100 pg/ml streptomycin (Gibco, lOOx stock).
- 3 Mio cells were plated per well of a 12-well-plate, including 100 ng/mLof recombinant human IL-6 and stored in the incubator at 37°C, 5 % CO2 for 24 hours.
- Samples were washed twice with sterile PBS and stained as follows: First, a live/dead staining was performed for 20 minutes at room temperature in the dark, using a NIR dye (ThermoFisher). After one washing step, human TruStain Fcx Blocking was added in FACS buffer (PBS with 2% Fetal Bovine Serum, 1% 0.5 M EDTA pH 8, 0.25% NaNs Sodium azide) and cells were incubated for another 10 minutes at room temperature (RT) in the dark prior to addition of the respective antibody mix (see Table 25).
- FACS buffer PBS with 2% Fetal Bovine Serum, 1% 0.5 M EDTA pH 8, 0.25% NaNs Sodium azide
- the FACS surface antibodies were purchased from BD Biosciences, Miltenyi or LuBiosciences and used according to the manufacturers’ recommendations.
- the detection antibody for human GPRC5D was generated by Roche.
- Results: Fig. 10 and Table 26 demonstrate that all BCMA-CD28 bispecific antibodies were able to significantly boost activation of CD8+ T cells (CD25 upregulation) on top of the GPRC5D-targeted CD3 engager (GPRC5D-TCB) mediated activation.
- the strongest additional activation at 200 nM was induced by BCMA-CD28 bsAb P1AG7215, followed by BCMA- CD28 Pl AG7191 and BCMA-CD28 Pl AG7207.
- No activation on top of the TCB was induced in presence of 200 nM of the untargeted CD28, which demonstrates that the CD28 molecules need to be crosslinked via the tumor antigen-targeting moiety to be active.
- Table 26 Up-regulation of the activation marker CD25 on CD8+ T cells upon incubation of primary MM BM aspirate samples with the indicated molecules for 96 hours, as assessed by flow cytometry
- T cell bispecific (TCB) isotype control T cell bispecific (TCB) isotype control
- FIGS 12A to 12D and Table 28 depict the BCMA-target dependent upregulation of T cell activation (CD25 on either CD4+ or CD8+ T cells), as well as the increase of the degranulation marker CD 107 on CD4+ and CD8+ T cells on top of the TCB-mediated activation/degranulation in presence of the indicated BCMA-CD28 bispecific antibodies but not in presence of the untargeted negative reference.
- the BCMA-CD28 bispecific antibody P1AG7215 induced a similar effect on activation of T cells as BCMA (PR)-CD28 v8 and BCMA (PR)-CD28 vl5, whereas BCMA-CD28 P1AG7282 induces slightly more moderate effects on top of the TCB-mediated ones.
- BCMA-CD28 bispecific antibodies which solely differ in their CD28 variant: CD28v8 (7062) or CD28vl5 (9053).
- Levels of CD107 on CD4+ T cells indicate strongest effects on top of TCB induced by BCMA (PR)-CD28 v8 and BCMA-CD28 P1AG7215, followed by BCMA-CD28 Pl AG7282.
- BCMA-CD28 Pl AG7282 On CD8+ T cells, only BCMA-CD28 Pl AG7282 reveals increased levels of CD107 on top of the one induced by TCB. Differences observed between the effect on T cell activation (CD25) and degranulation (CD 107) might be based on different kinetics of those markers.
- CD25 the effect on T cell activation
- CD 107 degranulation
- Table 28 Up-regulation of the activation marker CD25 or the degranulation marker CD107 on CD4+ or CD8+ T cells upon incubation of primary MM BM aspirate samples with the indicated molecules for 96 hours, as assessed by flow cytometry Another series of experiments was conducted using BM MNCs from primary MM patients after thawing. As indicated in Figures 13A and 13B and Table 29, BCMA-CD28 Pl AG7191 induced significant lysis of malignant MM plasma cells as well as upregulation of the frequency of CD25-positive CD8+ T cells on top of GPRC5D TCB in all three samples assessed.
- Table 29 Increased lysis of primary malignant MM plasma cells, respectively upregulation of the activation marker CD25 on CD8+ T cells upon incubation of primary MM BM MNCs with the indicated molecules for 96 hours, as assessed by flow cytometry
- Human NCI-H929 cells obtained from Roche Nutley were cultured in RPMI1640 high glucose medium containing 10% FCS, 2 mM L-Glutamine, 10 mM HEPES and 1 mM Sodiumpyruvate (37 °C at 5 % CO2).
- 50 microliters cell suspension 2.5 xlO 6 cells
- Mouse model Humanized NSG mice were provided by Jackson Laboratories, Sacramento USA.
- mice were irradiated (140cGy) and injected with CD34 + cord blood cells (9xl0 4 cells) from healthy donors (hematopoietic stem cells; HSC). After arrival animals were maintained for one week to get accustomed to new environment and for observation. Mice were maintained under specific pathogen-free conditions with daily cycles of 12 h light /12 h darkness according to committed guidelines (GV-Solas; Felasa; TierschG). Continuous health monitoring was carried out on a regular basis. Experimental study protocol was reviewed and approved by the local government (ROB-55.2-2532. Vet_03-20-170).
- Combinations of P1AG7215 and P1AG7200 were found to act synergistically with GPRC5D x CD3 and improve tumor growth inhibition in the NCI-H929 model (Fig. 15E to 15 J).
- the in vivo efficacy of the assessed new BCMA-CD28 bispecific antibodies was in a similar range as compared to the BCMA-CD28 bispecific antibodies comprising prior art BCMA antibody PR.
- the treatment groups of P1AG7215 (20 mg/kg, see Fig. 15E) and P1AG7200 (20 mg/kg, see Fig. 15H) showed lowest tumor volumes at termination.
- Table 30 Tumor volume (mm 3 ) at termination
- Table 31 P-values of combination groups compared to GPRC5D x CD3
- Pl AG7191 was tested in different doses (40, 10 and 1 mg/kg) in combination with GPRC5D x CD3 (0.05 mg/kg Pl AE6625) and compared to the GPRC5D x CD3 monotherapy and combination with the molecule P1AE9053 comprising BCMA clone PR (10 mg/kg).
- P1AG7191 and P1AE9053 were injected 24 hours after GPRC5D x CD3.
- the GPRC5D x CD3 monotherapy induces strong tumor growth inhibition in the NCI-H929 model (Fig. 17A). However, tumors start to relapse after the 4th treatment cycle (Table 31).
- the two highest doses of Pl AG7191 (40 and 10 mg/kg) delay the time to tumor relapse and inhibit the tumor regrowth.
- 6 and 8 responders were left in the P1AG7191 40 mg/kg and 10 mg/kg combination groups, respectively.
- the molecule P1AE9053 shows lower combination effects compared to Pl AG7191 as only 5 responders were left at termination and more tumors escape at earlier time points (Table 32).
- mice carrying the human FcRn instead of the mouse FcRn are considered more predictive for the clearance in human than in wild-type mice (C57/B16).
- Serum samples were analyzed using an immunoassay with anti-human FCpan (CH2) or kCH capture and detection reagents. As shown in Figure 18 and Table 33, all tested BCMA-CD28 bispecific antibodies have a similar plasma concentration-time profiles and clearance values ranging between 4 to 7 mL/kg/day in these mouse strains.
- CH2 anti-human FCpan
- kCH capture and detection reagents As shown in Figure 18 and Table 33, all tested BCMA-CD28 bispecific antibodies have a similar plasma concentration-time profiles and clearance values ranging between 4 to 7 mL/kg/day in these mouse strains.
- the Cell Microarray Technology (Charles River Laboratories) was used to screen for potential off-target binding interactions.
- Bispecific IgGl antibodies were screened for binding against fixed HEK293 cells expressing 6019 individual full-length human plasma membrane proteins and cell surface-tethered human secreted proteins, as well as a further 397 human heterodimers. Fluorescent images of the spotted cells were analyzed using ImageQuant software (GE healthcare, Version 8.2).
- Table 34 reports the results of the screen as strong, medium, weak or no interaction after visual inspection of the spots intensity.
- MAPPs Major histocompatibility complex-II (MHC-II)-Associated Peptide Proteomics (MAPPs) is a mass spectrometry-based approach to identify and relatively quantitate naturally processed and presented MHC-II-associated peptides that can potentially activate T cells and contribute to the immunogenicity of a drug.
- MHC-II Major histocompatibility complex-II
- MAPPs is a mass spectrometry-based approach to identify and relatively quantitate naturally processed and presented MHC-II-associated peptides that can potentially activate T cells and contribute to the immunogenicity of a drug.
- PBMCs Peripheral blood mononuclear cells
- Ficoll-Paque PLUS GE Healthcare Europe GmbH, Glattbrugg, Switzerland
- Monocytes were isolated by positive immunoselection using anti-CD14-coated microbeads and a magnetic separator (MACS, Miltenyi Biotech, Bergisch Gladbach, Germany).
- CD14 + cells were then cultured at a concentration of 0.3 x 10 6 cells/mL in 100mm ultra-low attachment culture dishes (Coming Inc., Corning, NY, USA) in serum-free Cellgro medium containing 1% GlutaMAX, 1% penicillin/streptomycin.
- Monocytes were differentiated into immature DCs with 50 ng/mL of GM-CSF and 5 ng/mL of IL-4 for 5 days at 37 °C with 5% CO2 before being challenged with a test protein at 50 pg/mL in the presence of 1 pg/mL of lipopolysaccharide (LPS) from Salmonella abortus equi (Sigma-Aldrich Chemie GmbH, Buchs, Switzerland) for 24 h.
- LPS lipopolysaccharide
- Mature DCs were harvested, washed with phosphate-buffered saline (PBS) and the cell pellets were frozen at -80 °C prior to subsequent immunoprecipitation.
- PBS phosphate-buffered saline
- HLA-DR-Presented Peptides Cell pellets were lysed in 20 mM Tris-buffer solution pH 7.8 containing 1% (v/v) Triton X-100 and protease inhibitors (Roche Diagnostics GmbH, Mannheim, Germany) for 1 h at 4 °C on a ThermoMixer at 1100 rpm.
- the HLA-DR immune complexes were isolated by immunoprecipitation using biotin-conjugated anti-human HLA-DR antibodies (clone L243, RayBiotech). Lysates were incubated with the antibody on a rotator overnight at 4°C.
- MHC-II peptide preparations obtained from about 3 x 10 6 human monocyte-derived DCs (moDCs) per sample were separated on a nanocapillary liquid chromatography system (UltiMate 3000 RSLC, Thermo Scientific, CA, USA) using self-packed fused-silica C18 reversed phase column (75 pm i.d. x 170 mm, ReproSil-Pur C18-AQ, 3 pm, Dr.
- DataMAPPs is a Roche-created data analysis tool to help visualising otherwise complex mass spectrometry-derived data in the form of a heat map.
- the program is publicly available at https://www.R-project.org/ and can be executed with a standard R installation. Please see reference (Steiner et al. 2020) for an extensive description of the package.
- DataMAPPs is invoked after processing the LC-MS/MS data with the PEAKS software and exporting results in a suitable tabular format.
- the dataMAPPs processing pipeline consists of (1) data import and consistency check: sample annotation and peptide quantification (PEAKS output) files are read into memory and checked for consistency.
- Cutoffs can be modified by the user after inspecting a collection of QC plots, which allows for a suitable, data set-specific processing while being able to rerun the procedure in a reproducible way; (3) data normalisation: peptide abundance (based on peak area) is normalised based on an adjusted version of the GRSN (global rank-invariant set normalisation) procedure. (4) replicate aggregation: technical replicates (i.e. samples run for the same donor, same treatment, same dose) can be averaged to retain information on all peptides that were detected in at least one of the replicates; (5) peptide mapping: antibody-associated peptides are mapped on the tested biologic’s protein amino acid sequences.
- peptides with neighbouring amino acid positions are binned to distinct epitope clusters (hotspots), whose abundance reflects the summed intensities of each of the constitutive MHC-II peptides;
- data export and visualisation the dataMAPPs workflow contains standard functionalities to generate several heatmap plots (epitope cluster or individual peptide level, per antibody or a global experiment summary) for a quick visualisation and comparison of the results
- P1AG7207 revealed three binding clusters. Among them, two clusters (Cl and C2) contained non-germline residues with frequencies of 2/12 and 5/12, respectively. In total 5/12 donors (41.6%) presented a potential immunogenic T cell epitope that could contribute to the immunogenicity of the compound.
- Pl AG7282 revealed three binding clusters. Among them, one cluster (Cl) contained non-germline residues with a frequency of 1/12 donors. In total 1/12 donors (8.3%) presented a potential immunogenic T cell epitope that could contribute to the immunogenicity of the compound..
- T cell activation is an important part of the immune response to therapeutic proteins and is usually required for the development of anti -drug antibodies in the clinic.
- the DC-T cell assay was used to assess the capacity of four BCMA-CD28 bispecific antibodies: Pl AG7191, P1AG215, P1AG7207, P1AG7282, to induce CD4 + T cells upon presentation of T cell epitopes by APCs.
- KLH Keyhole limpet haemocyanin
- Avastin® bevacizumab
- Monocytes were isolated from frozen PBMC samples by magnetic bead selection and differentiated into immature DC (iDC) using GM-CSF and IL-4. iDC were then harvested, washed and loaded with each individual test protein/peptide for 4 hours at 37°C. A DC maturation cocktail containing TNFa and IL-ip was then added for a further 40-42 hours to activate/mature the DC (mDC).
- the expression of key DC surface markers (CD11c, CD14, CD40, CD80, CD83, CD86, CD209 and HLA-DR) at both the immature and mature stage were assessed by flow cytometry to ensure the DC were activated prior to T cell interaction.
- the mDCs were then co-cultured with autologous CD4 + T cells (isolated by magnetic bead selection) for 6 days at 37°C, 5% CO2 in a humidified atmosphere.
- autologous monocytes were isolated from PBMC using magnetic bead selection and loaded with the selected protein/peptide that were initially used to load the DC.
- the monocytes were added to anti-IFNy pre-coated FluoroSpot plates (Mabtech) along with the corresponding DC:CD4 co-culture in quadruplicate.
- the FluoroSpot plates were incubated for 40-42 hours at 37°C, 5% CO2 in a humidified atmosphere. After incubation the FluoroSpot plates were developed using an in-house procedure and the spotforming cells (SFC) per well assessed for each cytokine in each test condition.
- SFC spotforming cells
- Data analysis Data management and statistical analysis has been performed in the R programming language (https://www.R-project.org/, v. 3.6.1). Data are transformed to a log2 scale and a Generalized linear model (GLM) is applied to quantify SI (fold change and 95% CI). Adjustments are applied to the dataset (exponential type of heteroscedasticity adjusted, gaussian noise injection at low end of SFU scale, linear regression and extrapolation of each SI to a blank value of 0) and QC plot are generated (DC differentiation markers, reproducibility on compound and donor level, relative stimulation of donors).
- GLM Generalized linear model
- SI Stimulation Index
- the DC used in this assay were of high quality and expressed high levels of the T cell co-stimulatory molecules required for the activation of T cells. Based on the readouts of the DC-T cell assay, all four BCMA-CD28 bispecific antibodies P1AG7191, P1AG215, P1AG7207 and Pl AG7282 were associated with a low risk of sequence-related immunogenicity for CD4 + T cell response (below the threshold of 10% as described in Siegel et al, Pharmaceutics 2022, 14(12), 2672).
- BCMA-CD28 bispecific antibodies may cause the formation of anti-drug antibodies (AD As), which may negatively impact the safety of the therapeutic result (e.g., allergic reactions, immune complex -mediated diseases).
- AD As anti-drug antibodies
- the risk for the four BCMA-CD28 bispecific antibodies P1AF7191, P1AF215, P1AF7207 and P1AF7282 to induce an unwanted immune response and the consequences for such a response were assessed through a combination of non-clinical assays (MAPPs and DC-T cell assay) as descibed herein before.
- MAPPs and DC-T cell assay non-clinical assays
- Cytokine analysis Determination of cytokine concentrations was performed on frozen plasma samples diluted 1 :5. Pre-tests revealed that levels of cytokine did not differ between fresh and thawed samples. Analyte concentrations were determined by ELISA using the Human Cytokine chemiluminescent assay kit (Aushon Ciraplex, Cat. No 101-269- 1-AB) with the SignaturePLUSTM imaging system and the Cirasoft analysis software. Results are expressed as pg/ml.
- ULOQ concentrations were assigned the concentration of the highest standard as follows: IFN-y, 500 pg/mL ; IL-2, 200 pg/mL ; IL-6, 2000 pg/mL ; IL-8, 4000 pg/mL ; TNF-a, 1000 pg/ml. Sample values below LLOQ concentrations were assigned LLOQ concentrations : IFN-y, 0.24 pg/mL ; IL-2, 98 fg/mL ; IL-6, 0.98 pg/mL ; IL-8, 1.95 pg/mL ; TNF-a, 0.49 pg/mL.
- BCMA-CD28 bispecific antibodies P1AG7215 and P1AG7062 did not trigger any cytokine release that was above the level of that mediated by Erbitux®, the molecule P1AE9053 (BCMA (PR)- CD28 vl5) triggered a release of IL-6 and IL-8 in 50 % of the donors.
Landscapes
- Health & Medical Sciences (AREA)
- Chemical & Material Sciences (AREA)
- Immunology (AREA)
- Organic Chemistry (AREA)
- Life Sciences & Earth Sciences (AREA)
- General Health & Medical Sciences (AREA)
- Biochemistry (AREA)
- Biophysics (AREA)
- Genetics & Genomics (AREA)
- Medicinal Chemistry (AREA)
- Molecular Biology (AREA)
- Proteomics, Peptides & Aminoacids (AREA)
- Cell Biology (AREA)
- Hematology (AREA)
- Peptides Or Proteins (AREA)
- Preparation Of Compounds By Using Micro-Organisms (AREA)
Abstract
The present invention relates to new humanized BCMA antibodies and bispecific antibodies that specifically bind to BCMA and CD28, methods for their production, pharmaceutical compositions containing these antibodies, and methods of using the same.
Description
Bispecific antibodies targeting BCMA and CD28
FIELD OF THE INVENTION
The present invention relates to new humanized BCMA antibodies and bispecific antibodies that specifically bind to BCMA and CD28, methods for their production, pharmaceutical compositions containing these antibodies, and methods of using the same.
BACKGROUND
Cancer immunotherapy is becoming an increasingly effective therapy option that can result in dramatic and durable responses in cancer types such as melanoma, non-small cell lung cancer and renal cell carcinoma. This is mostly driven by the success of several immune checkpoint inhibitors including anti-PD-1 (e.g. Keytruda, Merck; Opdivo, BMS), anti-CTLA-4 (e.g. Yervoy, BMS) and anti-PD-Ll (e.g. Tecentriq, Roche). These agents are likely to serve as standard of care for many cancer types, or as the backbone of combination therapies, however, only a fraction of patients (<25%) benefits from such therapies. Furthermore, various cancers (prostate cancer, colorectal cancer, pancreatic cancer, sarcomas, non-triple negative breast cancer etc.) present primary resistance to these immunomodulators. A number of reports indicate that the absence of pre-existing anti-tumor T cells contributes to the absence or poor response of some patients. In summary, despite impressive anti -cancer effects of existing immunotherapies, there is a clear medical need for addressing a large cancer patient population and for developing therapies that aim to induce and enhance novel tumor-specific T cell responses.
Affecting -75,000 new patients every year in the EU and US, multiple myeloma (MM) is one of the most common hematological malignancies with remaining high unmet medical need. Multiple myeloma, also known as plasma cell myeloma, is characterized by terminally differentiated plasma cells that secrete non-functional monoclonal immunoglobulins. As the cancerous plasma cells accumulate in the bone marrow, they interfere with the production of normal blood cells, leading to various symptoms and complications. Common symptoms of multiple myeloma include bone pain, especially in the back or ribs, fatigue, weakness, frequent infections, weight loss, excessive thirst, and increased urination. Treatment options for multiple myeloma depend on various factors, including the stage of the disease or the patient's overall health. In the short-term, the immunomodulatory drugs such as lenalidomide and pomalidomide (ImiDs), and proteasome inhibitors such as carfilzomib or bortezomib may remain the backbone
of 1st line therapy for multiple myeloma (Moreau et al, The Lancet Oncology 2021, 22(3), el05- el 18). However, these drugs do not target specifically the diseased tumor cells e.g. diseased plasma cells (PC). Efforts have been made towards selectively depleting the plasma cells in multiple myeloma. The lack of surface proteins that specifically mark plasma cells has hampered the development of antibodies or cellular therapies for multiple myeloma. So far, there are few cases of successful biologies, including daratumumab (anti-CD38) and elotuzumab (anti-CD319), with the caveat that both antigens are also expressed on other normal tissues including hematopoietic lineages and immune effector cells, which may limit their long-term clinical use.
B cell maturation antigen (BCMA), a transmembrane glycoprotein in the tumor necrosis factor receptor superfamily 17 (TNFRSF17), is expressed at significantly higher levels in all patient MM cells but not on other normal tissues except normal plasma cells. BCMA-chimeric antigen receptor (CAR) T-cells have already shown significant clinical activities in patients with relapsed/refractory MM (RRMM) who have undergone at least three prior treatments, including with a proteasome inhibitor and an immunomodulatory drug (IMiD). Additional modalities, including anti-BCMA antibody-drug conjugate also has achieved significant clinical responses in patients who failed at least three prior lines of therapy, including an anti-CD38 antibody, a proteasome inhibitor, and an immunomodulatory drug (Cho et al, Front Immunolog. 2018, 9, 1821). However, there is still a need for better treatment options of Multiple Myeloma.
CD28 is the founding member of a subfamily of costimulatory molecules characterized by paired V-set immunoglobulin superfamily (IgSF) domains attached to single transmembrane domains and cytoplasmic domains that contain critical signaling motifs (Carreno and Collins, Annu Rev Immunol. 2002, 20, 29-53). Other members of the subfamily include ICOS, CTLA-4, PD1, PD1H, TIGIT, and BTLA (Chen and Flies, Nat Rev Immunol. 2013, 13(4), 227-42). CD28 expression is restricted to T cells and prevalent on all naive and a majority of antigen- experienced subsets, including those that express PD-1 or CTLA-4. CD28 and CTLA-4 are highly homologous and compete for binding to the same B7 molecules CD80 and CD86, which are expressed on dendritic cells, B cells, macrophages, and tumor cells (Linsley et al., Proc Natl Acad Sci USA. 1990, 87(13), 5031-5). The higher affinity of CTLA-4 for the B7 family of ligands allows CTLA-4 to outcompete CD28 for ligand binding and suppress effector T cells responses (Engelhardt et al., J Immunol 2006, 177, 1052-1061). In contrast, PD-1 was shown to inhibit CD28 signaling by in part dephosphorylating the cytoplasmic domain of CD28 (Hui et al., Science 2017, 355, 1428-1433). Ligation of CD28 by CD80 or CD86 on the surface of professional antigen-presenting cells is strictly required for functional de novo priming of naive T cells, subsequent clonal expansion, cytokine production, target cell lysis, and formation of long-lived memory. Binding of CD28 ligands also promotes the expression of inducible costimulatory receptors such as OX-40, ICOS, and 4- IBB (Acuto and Michel, Nat Rev Immunol 2003, 3, 939-951). Upon ligation of CD28, a disulfide-linked homodimer, the membrane
proximal YMNM motif and the distal PYAP motif have been shown to complex with several kinases and adaptor proteins (Boomer and Green, Cold Spring Harb Perspect Biol 2010, 2, a002436). These motifs are important for the induction of IL2 transcription, which is mediated by the CD28-dependent activation of NF AT, AP-1, and NFKB family transcription factors (Fraser et al., Science 1991, 251, 313-316). However, additional poorly characterized sites for phosphorylation and ubiquitination are found within the cytoplasmic domain of CD28. As reviewed by (Esensten et al., Immunity 2016, 44, 973-988), CD28-initiated pathways have critical roles in promoting the proliferation and effector function of conventional T cells. CD28 ligation also promotes the anti-inflammatory function of regulatory T cells. CD28 co-stimulates T cells by in part augmenting signals from the T cell receptor, but was also shown to mediate unique signaling events (Acuto and Michel, 2003; Boomer and Green, 2010). Signals specifically triggered by CD28 control many important aspects of T cell function, including phosphorylation and other post-translational modifications of downstream proteins (e.g., PI3K mediated phosphorylation), transcriptional changes (eg. Bcl-xL expression), epigenetic changes (e.g. IL-2 promoter), cytoskeletal remodeling (e.g. orientation of the microtubule-organizing center) and changes in the glycolytic rate (e.g. glycolytic flux). CD28 -deficient mice have reduced responses to infectious pathogens, allograft antigens, graft-versus-host disease, contact hypersensitivity and asthma (Acuto and Michel, 2003). Lack of CD28-mediated co-stimulation results in reduced T cell proliferation in vitro and in vivo, in severe inhibition of germinal -centre formation and immunoglobulin isotype-class switching, reduced T helper (Th)-cell differentiation and the expression of Th2-type cytokines. CD4-dependent cytotoxic CD8+ T-cell responses are also affected. Importantly, CD28-deficient naive T cells showed a reduced proliferative response particularly at lower antigen concentrations. A growing body of literature supports the idea that engaging CD28 on T cells has anti-tumor potential. Recent evidence demonstrates that the anti-cancer effects of PD-L1/PD-1 and CTLA-4 checkpoint inhibitors depend on CD28 (Kamphorst et al., Science 2017, 355, 1423-1427). Clinical studies investigating the therapeutic effects of CTLA-4 and PD-1 blockade have shown exceptionally promising results in patients with advanced melanoma and other cancers. In addition, infusion of genetically engineered T cells expressing artificial chimeric T cell receptors comprising an extracellular antigen recognition domain fused to the intracellular TCR signaling domains (CD3z) and intracellular co-stimulatory domains (CD28 and/or 4- IBB domains) has shown high rates and durability of response in B cell cancers and other cancers.
CD28 agonistic antibodies can be divided into two categories: (i) CD28 superagonistic antibodies and (ii) CD28 conventional agonistic antibodies. Normally, for the activation of naive T cells both engagement of the T cell antigen receptor (TCR, signal 1) and costimulatory signaling by CD28 (signal 2) is required. CD28 Superagonists (CD28SA) are CD28-specific monoclonal antibodies, which are able to autonomously activate T cells without overt T cell
receptor engagement (Hiinig, Nat Rev Immunol 2012, 12, 317-318). In rodents, CD28SA activates conventional and regulatory T cells. CD28SA antibodies are therapeutically effective in multiple models of autoimmunity, inflammation and transplantation. However, a phase I study of the human CD28SA antibody TGN1412 resulted in a life-threatening cytokine storm in 2006. Follow-up studies have suggested that the toxicity was caused by dosing errors due to differences in the CD28 responsiveness of human T cells and T cells of preclinical animal models. Theralizumab (TGN1412 or TAB08) has been re-evaluated in an open-label, multicenter dose escalation study in RA patients and patients with metastatic or unresectable advanced solid malignancies. CD28 conventional agonistic antibodies, such as clone 9.3, mimic CD28 natural ligands and are only able to enhance T cell activation in presence of a T cell receptor signal (signal 1). Published insights indicate that the binding epitope of the antibody has a major impact on whether the agonistic antibody is a superagonist or a conventional agonist (Beyersdorf et al., Ann. Rheum. Dis. 2005, 64, iv91-iv95). The superagonistic TGN1412 binds to a lateral motif of CD28, while the conventional agonistic molecule 9.3 binds close to the ligand binding epitope. As a consequence of the different binding epitopes, superagonistic and conventional agonistic antibodies differ in their ability to form linear complexes of CD28 molecules on the surface of T cells. Precisely, TGN1412 is able to efficiently form linear arrays of CD28, which presumably leads to aggregated signaling components which are sufficient to surpass the threshold for T cell activation. The conventional agonist 9.3, on the other hand, leads to complexes which are not linear in structure. An attempt to convert conventional agonistic binders based on the 9.3 clone has been previously published (Otz et al., Leukemia 2009, 23(1), 71-77) using a recombinant bi-specific single-chain antibody directed to a melanoma-associated proteoglycan and CD28. The reported bispecific single chain antibody was reported to exert “supra-agonistic” activity despite the use of a conventional CD28 agonistic binder 9.3, based in the intrinsic tendency of bispecific single chain antibodies to form multimeric constructs.
It has been found that a better T cell activation is achieved when limiting amounts of anti- CD3 bispecific antibodies, i.e. T cell bispecific antibodies (TCBs), are combined with agonistic anti-CD28 molecules. Given, that CD28 is expressed at baseline on T cells in various tumor indications and activation of CD28 signaling enhances T cell receptor signals, the combination of a TCB molecule with a tumor-targeted CD28 molecule can act synergistically to induce strong and long-lasting anti-tumor responses. WO 2020/127618 Al describes tumor-targeted agonistic CD28 antigen binding molecules. Various tumor targets are described therein.
CD28 agonism in Multiple Myeloma may exert different biological functions on immune, respective MM plasma cells. While co-activation of T-cells via CD28 is expected to drive antitumor responses, CD28 agonism on MM cells mediates pro-survival signaling via regulation of PI3K/Akt, FoxO3a, and Bimm which in turn is described to induce chemotherapeutic resistance in multiple myeloma (Murray et al, Blood 2014, 123(24), 3770-3779). Over-expression of CD28
on newly diagnosed Multiple Myeloma plasma cells is described to correlate with worse clinical outcome. However, CD28 activation inhibits myeloma cell proliferation (Bahlis et al., Blood 2007, 109(11), 5002-5010).
Agonizing CD28 in presence of a strong immune cell mediated response, such as a T-cell bispecific activation of T-cells, can further boost efficient anti-tumor responses. We herein provide bispecific agonistic CD28 antigen binding molecules that specifically bind to BCMA. Enhancing a T cell response with a CD28 bispecific antibody targeting BCMA on myeloma cells may be an option to improve the treatment of Multiple Myeloma and there is a need to provide BCMA-targeted anti-CD28 antibodies with advantageous properties.
SUMMARY
The present invention describes new BCMA-targeted bispecific agonistic CD28 antigen binding molecules which achieve a tumor-dependent T cell activation and tumor cell killing without the necessity to form multimers. The bispecific CD28 antigen binding molecules of the present invention are characterized by monovalent binding to CD28 and in that they comprise a certain antigen binding domain as defined herein capable of specific binding to BCMA. Furthermore, they possess an Fc domain composed of a first and a second subunit capable of stable association comprising one or more amino acid substitution that reduces the binding affinity of the antigen binding molecule to an Fc receptor and/or effector function. Fc receptor- mediated cross-linking is thereby abrogated and tumor-specific activation is achieved by crosslinking through binding of the second antigen binding domain capable of specific binding to BCMA.
Thus, provided herein is an antibody that specifically binds to B cell maturation agent (BCMA), wherein the antibody comprises a first antigen binding domain comprising
(i) a heavy chain variable region (VH BCMA) comprising a heavy chain complementary determining region CDR-H1 of SEQ ID NO: 1 (GYTFTNYWMH), a CDR-H2 of SEQ ID NO: 2 (IH4PNSGSTNYNEKFQG), and a CDR-H3 of SEQ ID NO: 3 (GIYDYPFAY), and
(ii) a light chain variable region (VLBCMA) selected from the group consisting of
(a) a VL comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 4 (RASES VSIHGTHLMH), a CDR-L2 of SEQ ID NO: 5 (AASSLQS) and a CDR-L3 of SEQ ID NO: 6 (QQSIEDPYT); or
(b) a VL comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 4 (RASES VSIHGTHLMH), a CDR-L2 of SEQ ID NO: 7 (AASNLES) and a CDR-L3 of SEQ ID NO: 6 (QQSIEDPYT), or
(c) a VL comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 4
(RASES VSIHGTHLMH), a CDR-L2 of SEQ ID NO: 8 (AASNLQS) and a CDR-L3 of SEQ ID NO: 6 (QQSIEDPYT).
In one aspect, provided is an antibody that specifically binds to BCMA, wherein the antibody comprises an amino acid sequence selected from the group consisting of SEQ ID NO: 9 (VHla) and SEQ ID NO: 10 (VHlb), and/or the VL BCMA comprises an amino acid sequence selected from the group consisting of SEQ ID NO: 11 (VLlf) , SEQ ID NO: 12 (VLla), SEQ ID NO: 13 (VLlb), SEQ ID NO: 14 (VLlc), SEQ ID NO: 15 (VLld), and SEQ ID NO: 16 (VLle).
In one aspect, provided is antibody that specifically binds to BCMA, wherein the antibody comprises a first antigen binding domain comprising
(a) a VH BCMA comprising an amino acid sequence of SEQ ID NO:9 and VLBCMA comprising an amino acid sequence of SEQ ID NO: 11, or
(b) a VH BCMA comprising an amino acid sequence of SEQ ID NOV and VLBCMA comprising an amino acid sequence of SEQ ID NO: 12.
In one particular aspect, the first antigen binding domain is a Fab molecule.
In one aspect, the antibody that specifically binds to BCMA comprises an Fc domain composed of a first and a second subunit.
In one aspect, the antibody that specifically binds to BCMA comprises a second antigen binding domain that specifically binds to a second antigen, i.e. is a bispecific antibody.
In one aspect, the second antigen binding domain that specifically binds to a second antigen is a Fab molecule wherein the variable domains VL and VH or the constant domains CL and CHI, particularly the variable domains VL and VH, of the Fab light chain and the Fab heavy chain are replaced by each other.
In one aspect, provided is an antibody that specifically binds to BCMA comprising a first antigen binding domain that specifically binds to BCMA, wherein the first antigen binding domain is a Fab molecule wherein in the constant domain CL the amino acid at position 123 (numbering according to Kabat EU index) is substituted by an amino acid selected from lysine (K), arginine (R) or histidine (H) and the amino acid at position 124 (numbering according to Kabat EU index) is substituted independently by lysine (K), arginine (R) or histidine (H), and wherein in the constant domain CHI the amino acid at position 147 (numbering according to Kabat EU index) is substituted independently by glutamic acid (E) or aspartic acid (D) and the amino acid at position 213 (numbering according to Kabat EU index) is substituted independently by glutamic acid (E), or aspartic acid (D) (numbering according to Kabat EU index).
In all of these aspects, the antibody that specifically binds to BCMA comprises an Fc domain, wherein the Fc domain is an IgG, particularly an IgGl Fc domain. In one aspect, the the Fc domain is a human Fc domain.
In one aspect, the Fc domain comprises a modification promoting the association of the first and the second subunit of the Fc domain. In one aspect, the Fc domain comprises knobs into
hole modifications. In one aspect, the first subunit of the Fc domain comprises the amino acid substitutions S354C and T366W (EU numbering) and the second subunit of the Fc domain comprises the amino acid substitutions Y349C, T366S and Y407V (numbering according to Kabat EU index).
In one further aspect, the the Fc domain comprises one or more amino acid substitution that reduces binding to an Fc receptor and/or effector function. In one particular aspect, the the Fc domain is of human IgGl subclass and comprises the amino acid mutations L234A, L235A and P329G (numbering according to Kabat EU index).
Provided herein are further antibodies that specifically bind to BCMA and that comprise a second antigen binding domain that specifically binds to CD28.
In one aspect, the second antigen binding domain that specifically binds to CD28 comprises a heavy chain variable region (VHCD28) comprising a heavy chain complementary determining region CDR-H1 of SEQ ID NO: 17, a CDR-H2 of SEQ ID NO: 18, and a CDR-H3 of SEQ ID NO: 19, and a light chain variable region (VLCD28) comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 20, a CDR-L2 of SEQ ID NO: 21 and a CDR-L3 of SEQ ID NO: 22.
In one aspect, the second antigen binding domain that specifically binds to CD28 comprises a heavy chain variable region (VHCD28) comprising an amino acid sequence of SEQ ID NO:23, and a light chain variable region (VLCD28) comprising an amino acid sequence of SEQ ID NO:24 (v8).
In one particular aspect, the antibody as described herein comprises a first antigen binding domain comprising a VH BCMA comprising an amino acid sequence of SEQ ID NO:9 and VL BCMA comprising an amino acid sequence of SEQ ID NO: 11 and a second antigen binding domain comprising VH CD28 comprising an amino acid sequence of SEQ ID NO:23 and VL CD28 comprising an amino acid sequence of SEQ ID NO:24.
In one aspect, the antibody comprises a first light chain comprising the amino acid sequence of SEQ ID NO:25, a first heavy chain comprising the amino acid sequence of SEQ ID NO:26, a second heavy chain comprising the amino acid sequence of SEQ ID NO:27 and a second light chain comprising the amino acid sequence of SEQ ID NO:28.
Provided herein is also an antibody that specifically binds to B cell maturation agent (BCMA) and CD28, wherein the antibody comprises
(A) a first antigen binding domain comprising
(i) a heavy chain variable region (VH BCMA) selected from the group consisting of
(a) a VH comprising a heavy chain complementary determining region CDR-H1 of SEQ ID NO: 29 (GFTFSNAWMD), a CDR-H2 of SEQ ID NO: 30 (QITAKSNNYATYYADSVKG), and a CDR-H3 of SEQ ID NO: 31 (DGYH), and
(b) a VH comprising a heavy chain complementary determining region CDR-H1 of SEQ ID NO:
29 (GFTFSNAWMD), a CDR-H2 of SEQ ID NO: 32 (QITAKSNNYATYYAAPVKG), and a CDR-H3 of SEQ ID NO: 31 (DGYH), and
(ii) a light chain variable region (VLBCMA) comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 33 (RASEDIRNGLA), a CDR-L2 of SEQ ID NO: 34 (NANSLHT) and a CDR-L3 of SEQ ID NO: 35 (EDTSKYPYT), and
(B) a second antigen binding domain that specifically binds to CD28.
In one aspect, the antibody comprises (A) a first antigen binding domain that specifically binds to BCMA, wherein the VH BCMA comprises an amino acid sequence selected from the group consisting of SEQ ID NO: 36 (VH2a) and SEQ ID NO: 38 (VHlb), and/or the VL BCMA comprises an amino acid sequence selected from the group consisting of SEQ ID NO: 37 (VL2a) and SEQ ID NO:39 (VLla).
In one aspect, provided is an antibody that specifically binds to BCMA and CD28, wherein the antibody comprises a first antigen binding domain comprising
(a) a VH BCMA comprising an amino acid sequence of SEQ ID NO:36 and a VLBCMA comprising an amino acid sequence of SEQ ID NO:37, or
(b) a VH BCMA comprising an amino acid sequence of SEQ ID NO:38 and a VLBCMA comprising an amino acid sequence of SEQ ID NO:39.
In one aspect, provided is an antibody that specifically binds to BCMA and CD28, wherein the first antigen binding domain that binds to BCMA is a Fab molecule.
In one aspect, the antibody that specifically binds to BCMA and CD28 comprises an Fc domain composed of a first and a second subunit.
In one aspect, the second antigen binding domain that specifically binds to CD28 is a Fab molecule. In one aspect, the second antigen binding domain that specifically binds to CD28 is a Fab molecule wherein the variable domains VL and VH or the constant domains CL and CHI, particularly the variable domains VL and VH, of the Fab light chain and the Fab heavy chain are replaced by each other.
In one aspect, provided is an antibody that specifically binds to BCMA and CD28, wherein the first antigen binding domain that binds to BCMA is a Fab molecule wherein in the constant domain CL the amino acid at position 123 (numbering according to Kabat EU index) is substituted by an amino acid selected from lysine (K), arginine (R) or histidine (H) and the amino acid at position 124 (numbering according to Kabat EU index) is substituted independently by lysine (K), arginine (R) or histidine (H), and wherein in the constant domain CHI the amino acid at position 147 (numbering according to Kabat EU index) is substituted independently by glutamic acid (E) or aspartic acid (D) and the amino acid at position 213 (numbering according to Kabat EU index) is substituted independently by glutamic acid (E), or aspartic acid (D) (numbering according to Kabat EU index).
In all of these aspects, the antibody that specifically binds to BCMA and CD28 comprises an Fc domain, wherein the Fc domain is an IgG, particularly an IgGl Fc domain. In one aspect, the the Fc domain is a human Fc domain.
In one aspect, the Fc domain comprises a modification promoting the association of the first and the second subunit of the Fc domain. In one aspect, the Fc domain comprises knobs into hole modifications. In one aspect, the first subunit of the Fc domain comprises the amino acid substitutions S354C and T366W (EU numbering) and the second subunit of the Fc domain comprises the amino acid substitutions Y349C, T366S and Y407V (numbering according to Kabat EU index).
In one further aspect, the the Fc domain comprises one or more amino acid substitution that reduces binding to an Fc receptor and/or effector function. In one particular aspect, the the Fc domain is of human IgGl subclass and comprises the amino acid mutations L234A, L235A and P329G (numbering according to Kabat EU index).
In all of these aspects, the second antigen binding domain that specifically binds to CD28 comprises a heavy chain variable region (VHCD28) comprising a heavy chain complementary determining region CDR-H1 of SEQ ID NO: 17, a CDR-H2 of SEQ ID NO: 18, and a CDR-H3 of SEQ ID NO: 19, and a light chain variable region (VLCD28) comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 20, a CDR-L2 of SEQ ID NO: 21 and a CDR-L3 of SEQ ID NO: 22.
In one particular aspect, the second antigen binding domain that specifically binds to CD28 comprises a heavy chain variable region (VHCD28) comprising an amino acid sequence of SEQ ID NO:23, and a light chain variable region (VLCD28) comprising an amino acid sequence of SEQ ID NO:24.
In one aspect, provided is an antibody that specifically binds to BCMA and CD28, wherein the antibody comprises a first antigen binding domain comprising a VH BCMA comprising an amino acid sequence of SEQ ID NO:36 and VLBCMA comprising an amino acid sequence of SEQ ID NO:37 and a second antigen binding domain comprising VH CD28 comprising an amino acid sequence of SEQ ID NO:23 and VL CD28 comprising an amino acid sequence of SEQ ID NO:24.
In one aspect, the antibody that specifically binds to BCMA and CD28 comprises a first light chain comprising the amino acid sequence of SEQ ID NO:40, a first heavy chain comprising the amino acid sequence of SEQ ID NO:41, a second heavy chain comprising the amino acid sequence of SEQ ID NO:27 and a second light chain comprising the amino acid sequence of SEQ ID NO:28.
According to another aspect of the invention, there is provided one or more isolated polynucleotide encoding the antibody as described herein before. The invention further provides a vector, particularly an expression vector, comprising the isolated polynucleotide of the invention and a host cell comprising the isolated nucleic acid or the expression vector of the
invention. In some aspects, the host cell is an eukaryotic cell, particularly a mammalian cell. In another aspect, provided is a method of producing an antibody that specifically binds to BCMA or a bispecific BCMA antibody as described herein before, comprising the steps of a) culturing the host cell as described above under conditions suitable for the expression of the antibody, and optionally b) recovering the an antibody that specifically binds to BCMA or the bispecific BCMA antibody. The invention also encompasses the antibody or bispecific antibody as produced by the method of the invention.
Further provided is a pharmaceutical composition comprising an antibody that specifically binds to BCMA or a bispecific BCMA antibody as described herein before and at least one pharmaceutically acceptable excipient. In one aspect, the pharmaceutical composition comprises an additional therapeutic agent.
Also encompassed is the antibody that specifically binds to BCMA or the bispecific BCMA antibody as described herein before, or the pharmaceutical composition comprising the bispecific BCMA antibody, for use as a medicament.
In one aspect, provided is the antibody that specifically binds to BCMA or the bispecific BCMA antibody as described herein before or the pharmaceutical composition, for use in enhancing (a) T cell activation or (b) T cell effector functions.
In one aspect, provided is the antibody that specifically binds to BCMA or the bispecific BCMA antibody as described herein before or the pharmaceutical composition, for use in the treatment of a disease. In one aspect the disease is cancer, in particular multiple myeloma (MM).
In one specific aspect, provided is the antibody that specifically binds to BCMA or the bispecific BCMA antibody as described herein before or the pharmaceutical composition, for use in the treatment of cancer, in particular multiple myeloma. In another specific aspect, provided is the antibody that specifically binds to BCMA or the bispecific BCMA antibody as described herein before for use in the treatment of cancer, wherein the use is for administration in combination with a chemotherapeutic agent, radiation therapy and/ or other agents for use in cancer immunotherapy. In one aspect, the antibody that specifically binds to BCMA or the bispecific BCMA antibody as described herein before is for use in the treatment of cancer, wherein the use is for administration in combination with a T-cell activating anti-CD3 bispecific antibody. In one parrticular aspect, the T-cell activating anti-CD3 bispecific antibody is an anti- GPRC5D/anti-CD3 antibody.
In a further aspect, the invention provides a method of inhibiting the growth of tumor cells in an individual comprising administering to the individual an effective amount of the antibody that specifically binds to BCMA or the bispecific BCMA antibody as described herein before, or the pharmaceutical composition of the invention, to inhibit the growth of the tumor cells. In another aspect, the invention provides a method of treating or delaying cancer in an individual comprising administering to the individual an effective amount of the antibody that specifically
binds to BCMA or the bispecific BCMA antibody as described herein before, or the pharmaceutical composition of the invention.
Also provided is the use of the antibody that specifically binds to BCMA or the bispecific BCMA antibody as described herein before for the manufacture of a medicament for the treatment of a disease in an individual in need thereof, in particular for the manufacture of a medicament for the treatment of cancer, as well as a method of treating a disease in an individual, comprising administering to said individual a therapeutically effective amount of a composition comprising the antibody that specifically binds to BCMA or the bispecific BCMA antibody of the invention in a pharmaceutically acceptable form. In a specific aspect, the disease is cancer. In any of the above aspects the individual is a mammal, particularly a human.
BRIEF DESCRIPTION OF THE DRAWINGS
In Figures 1A to 1C schematic illustrations of exemplary molecules as described herein are shown. Fig. 1A shows a schematic illustration of the CD28 agonistic antibody variants as monovalent hu IgGl PGLALA isotype (“Fc silent”). Fig. IB shows a bispecific BCMA-CD28 antigen binding molecule in 1+1 format, wherein in the Fab molecule comprising the CD28 antigen binding domain the VH and VL domains are exchanged with each other (VH/VL crossfab) and wherein in the Fab molecule comprising the BCMA antigen binding domain certain amino acids in the CHI and CL domain are exchanged (charged variants) to allow better pairing with the light chain. Fig. 1C shows a bispecific BCMA-CD28 antigen binding molecule in 1+1 format, wherein in the Fab molecule comprising the BCMA antigen binding domain the VH and VL domains are exchanged with each other (VH/VL crossfab) and wherein in the Fab molecule comprising the CD28 antigen binding domain certain amino acids in the CHI and CL domain are exchanged (charged variants) to allow better pairing with the light chain.
Figures 2A and 2B show the binding of various BCMA-CD28 bispecific antigen binding molecules to CD28-expressing CHO cells (CHO-kl-huCD28 cells). All BCMA-CD28 bispecific antigen binding molecules were able to bind to human CD28 on CHO-kl-huCD28 cells in a concentration dependent manner, assessed by flow cytometry. However, binding to human CD28 does not reach saturation due to low affinity binders and is comparable among the CD28v8 bispecific antibodies (Fig. 2A) and CD28vl5 bispecific antibodies (Fig. 2B), respectively.
Figures 3A and 3B show the binding of various BCMA-CD28 bispecific antigen binding molecules to BCMA-expressing CHO cells (CHO-huBCMA cells). All BCMA-CD28 bispecific antigen binding molecules were able to bind to human BCMA in a concentration dependent manner, assessed by flow cytometry. The molecules with the BCMA antibody PR described in WO 2020/127618 Al have lower ECso values compared to the molecules with the new BCMA
antibodies as described herein, but maximal binding (Emax) is comparable among all molecules tested.
Figures 3C to 3H show binding of two BCMA-CD28 bispecific antigen binding molecules Pl AG7191 and Pl AG7207 as well as the BCMA-targeted CD3 T cell engager Alnuctamab, to CHO cells expressing human BCMA with indicated point mutations. Shown is the binding in case no BCMA is present (Fig. 3C) and to human wt BCMA (Fig. 3D) as well as to BCMA variants human BCMA P33S (Fig. 3E), human BCMA P343del (Fig. 3F), human BCMA R27P (Fig. 3G) and human BCMA S3 Odel (Fig. 3H). Another BCMA-targeted CD3 T cell engager Teclistamab does not bind to two mutated BCMA variants, namely R27P (Fig. 3G) and S30del (Fig. 3H) and Elranatamab only weakly binds to BCMA with the R27P mutation at high concentrations. Neither of the tested bispecific molecules bind to BCMA-negative CHOkl cells (3C).
Figures 4A to 4F show the dose-dependent activation of IL2 signaling in Jurkat IL2 reporter cells. Titrated amounts of (200.0-0.5 nM) of BCMA-CD28 vl5 BsAbs (Fig. 4A to 4C) and BCMA-CD28 v8 BsAbs (Fig. 4D to 4F) were added together with three different concentrations (20, 200, and 2000 pM) of GPRC5D-TCB to the mixture of the target (NCI-H929) and effector (Jurkat IL2 reporter) cells.
Figures 5A to 5D provide a summary of the data obtained from Jurkat NFkB reporter assay. Fig. 5A shows the ECso values and Fig. 5B compares the efficacy of various BCMA- CD28 vl5 BsAbs and the non-targeted CD28 vl5 control, and Fig. 5C shows the ECso values and Fig. 5D compares the efficacy of the various BCMA-CD28 v8 BsAbs and the non-targeted CD28 control. The data was calculated from three independent experiments performed in triplicate. ECso data of Fig. 5 A and Fig. 5C were added together from experiments with different concentrations of GPRC5D-TCB, and the same was applied for efficacy data. All data are shown as mean ± s.d.
Figures 6A and 6B show that the dose-dependent activation of CD8+ T cells through BCMA-CD28 vl5 bispecific antibodies (BsAbs) occurs only in presence of a first signal (TCB) and BCMA expression. The BCMA-expressing MM cell line NCI-H929 (Fig. 6A) was cocultured with healthy donor PBMCs (ratio of 1 : 1). Alternatively, NCI-H929 BCMAko cells (knock-out cells that do not express BCMA) were used as target cells (Fig. 6B). Co-cultures were treated with GPRC5D-TCB (providing 1st signal) alone or in combination with BCMA- CD28 BsAbs or an untargeted CD28 (neg. ref.) which were titrated from 500 nM to 0.03 nM (1 :4 dilution steps) and incubated for 4 days. Controls were left untreated or were treated with the BCMA-CD28 BsAbs alone (3 highest concentrations). Data are shown as mean of triplicates ± s.d. of one donor (representative for 3 donors tested; BCMA ko N=l).
Figures 7A and 7B show that the dose-dependent activation of CD8+ T cells through BCMA-CD28 v8 bispecific antibodies (BsAbs). Activation occurs only in presence of a first signal (TCB) and BCMA expression. The BCMA-expressing MM cell line NCI-H929 (Fig. 7A) was co-cultured with healthy donor PBMCs (ratio of 1: 1). Alternatively, NCI-H929 BCMAko cells (knock-out cells that do not express BCMA) were used as target cells (Fig. 7B). Co-cultures were treated with GPRC5D-TCB (providing 1st signal) alone or in combination with BCMA- CD28 BsAbs or an untargeted CD28 (neg. ref.) which were titrated from 500 nM to 0.12 nM (1 :4 dilution steps) and incubated for 4 days. Controls were left untreated or were treated with the BCMA-CD28 BsAbs alone. Data are shown as mean of triplicates ± s.d. of one donor (representative for 3 donors tested; BCMA ko N=l).
Figures 8A and 8B show that the dose-dependent activation of CD4+ T cells through BCMA-CD28 v8 bispecific antibodies (BsAbs) occurs only in presence of a first signal (TCB) and BCMA expression. The BCMA-expressing MM cell line NCI-H929 (Fig. 8A) was cocultured with healthy donor PBMCs (ratio of 1 : 1). Alternatively, NCI-H929 BCMAko cells (knock-out cells that do not express BCMA) were used as target cells (Fig. 8B). Co-cultures were treated with GPRC5D-TCB (providing 1st signal) alone or in combination with BCMA- CD28 BsAbs or an untargeted CD28 (neg. ref.) which were titrated from 500 nM to 0.12 nM (1 :4 dilution steps) and incubated for 4 days. Controls were left untreated or were treated with the BCMA-CD28 BsAbs alone (3 highest concentrations). Data are shown as mean of triplicates ± s.d. of one donor (representative for 3 donors tested; BCMA ko N=l).
Figures 9A and 9B show that the dose-dependent proliferation of CD4+ T cells through BCMA-CD28 v8 bispecific antibodies (BsAbs). Proliferation occurs only in presence of a first signal (TCB) and BCMA expression. The BCMA-expressing MM cell line NCI-H929 (Fig. 9A) was co-cultured with healthy donor PBMCs (ratio of 1 : 1). Alternatively, NCI-H929 BCMAko cells (knock-out cells that do not express BCMA) were used as target cells (Fig. 9B). Co-cultures were treated with GPRC5D-TCB (providing 1st signal) alone or in combination with BCMA- CD28 BsAbs or an untargeted CD28 (neg. ref.) which were titrated from 500 nM to 0.12 nM (1 :4 dilution steps) and incubated for 4 days. Controls were left untreated or were treated with the BCMA-CD28 BsAbs alone. Data are shown as mean of triplicates ± s.d. of one donor (representative for 3 donors tested; BCMA ko N=l).
Figure 10 shows results of an exemplary ex vivo test, using primary MM patients’ bone marrow samples as described in Example 4.3. T-cell activation was determined by flow cytometry, assessing the upregulation of CD25 on CD8+ T cells upon incubation with 1 nM GPRC5D-TCB in absence or presence of 200 nM of the indicated BCMA-CD28 bispecific antibody or untargeted negative reference molecule for 96 hours. All data shown refer to single tubes measurement per condition.
Figure 11 shows results of an exemplary ex vivo test, using primary MM patients’ bone marrow samples. Tumor Cell Lysis (TCL) was determined by flow cytometry, defining maleimide positive cells with low FSC as dead cells, upon incubation with 1 nM GPRC5D-TCB in absence or presence of 400 nM of the indicated BCMA-CD28 bispecific antibody or untargeted negative reference molecule for 96 hours. All data shown refer to single tube measurement per condition.
Figures 12A to 12D show the results of an ex vivo experiment, using primary MM patients’ bone marrow samples. T-cell activation, respective degranulation was determined by flow cytometry, assessing the upregulation of CD25 (see Fig. 12A and 12B) respective CD107A (see Fig. 12C and 12D) on CD4+ (see Fig. 12A and 12C) or CD8+ (see Fig. 12B and 12D) T cells upon incubation with 10 nM GPRC5D-TCB in absence or presence of 200 nM of the indicated BCMA-CD28 bispecific antibodies or reference molecules for 96 hours.
Figures 13A and 13B show the results of an ex vivo test, using BM MNCs of primary MM patients’ bone marrow samples. Tumor Cell lysis respective T-cell activation was determined by flow cytometry, assessing the percentage of viable MM PCs (Fig. 13 A) or the upregulation of CD25 on CD8+ T cells (Fig. 13B) upon incubation with 0.01 nM GPRC5D-TCB in absence or presence of 800 nM of the indicated BCMA-CD28 bispecific antibodies or reference molecules for 96 hours.
Figures 14A to 14F depict the results from an in vivo experiment testing the efficacy of GPRC5D x CD3 monotherapy and its combination with BCMA-CD28 bispecific antibodies (lOmg/kg P1AG7215 and 10 mg/kg P1AG7282) in the NCI-H929 tumor model. Figures 14A to 14 D show the tumor growth inhibition in single animals and Figure 14E shows the tumor volume as Median (+/- IQR) per treatment group. Animals with a terminal tumor load below the size at treatment start were defined as responders. Tumor volume at termination (study day 41) is shown for GPRC5D x CD3 monotherapy group and both combinations as Median (+/- IQR) is shown in Fig. 14F.
Figures 15A to 15L show the results from an in vivo experiment testing the efficacy of GPRC5D x CD3 monotherapy and its combination with BCMA-CD28 var8 bispecific antibody P1AG7215 (20 mg/kg, 10 mg/kg, 5 mg/kg), BCMA-CD28 varl5 bispecific antibody P1AG7282 (20 mg/kg, 10 mg/kg, 2 mg/kg) and BCMA(PR)-CD28 bispecific antibodies P1AE9053 (10 mg/kg) and P1AF7062 (10 mg/kg) in the NCI-H929 tumor model. Tumor growth inhibition is shown in single animals (Figures 15A to 15 J) and as Median (+/- IQR) per treatment group (Fig. 15K). Animals with a terminal tumor load below the size at treatment start were defined as responders. Significant differences shown as asterisk. Tumor volume at termination (study day 40) is shown for GPRC5D x CD3 monotherapy and combination groups as Median (+/- IQR) (C).
in the NCI-H929 tumor model. Figures 15A to 15J show the tumor growth inhibition in single animals and Figure 15K shows the tumor volume as Median (+/- IQR) per treatment group. Animals with a terminal tumor load below the size at treatment start were defined as responders. Tumor volume at termination (study day 41) is shown for GPRC5D x CD3 monotherapy group and both combinations as Median (+/- IQR) is shown in Fig. 15L.
Figures 16A to 16G show the results of an in vivo experiment testing the efficacy of GPRC5D x CD3 monotherapy (1 mg/kg) and combination with BCMA-CD28 (20 and 30 mg/kg) Pl AG7207 (20 and 30 mg/kg) and Pl AG7191 (20 mg/kg), respectively in the NCI-H929 tumor model. Figures 16A to 16E show the tumor growth inhibition in single animals. Figure 16F shows the tumor volume as Median (+/- IQR) per treatment group. Animals with a terminal tumor load below the size at treatment start were defined as responders. Treatment with the combination of GPRC5D x CD3 and 20 mg/kg Pl AG7207 led to 4 responders and with 30 mg/kg Pl AG7207 to 5 responders. 2 responders were observed in the group treated with GPRC5D x CD3 and 20 mg/kg P1AG7191. In Fig. 16G tumor weight at termination (study day 57) is shown for the GPRC5D x CD3 monotherapy group and the combinations as Median +/- IQR.
Figures 17A to 17G show the results of an in vivo experiment testing the efficacy of GPRC5D x CD3 monotherapy and in combination with BCMA-CD28 bispecific antibodies Pl AG7191 (40, 10 and 1 mg/kg) and P1AE9053 (10 mg/kg) in the NCI-H929 tumor model. Tumor growth inhibition is shown as Median (+/- IQR) per treatment group (Fig. 17A) and in single animals (Fig. 17B to 17G). Animals with a terminal tumor load below the size at treatment start were defined as responders. The two highest doses of Pl AG7191 (40 and 10 mg/kg) delay the time to tumor relapse compared to monotherapy and inhibit the tumor regrowth.
Figure 18 shows a comparison of the plasma concentration-time profiles of bispecific antibodies P1AG7282, P1AG7215, P1AG7191 and P1AG7207 as measured in HuFcRN transgenic mice (PK study).
Figure 19 shows a comparative heatMAPPS representation of the results observed in the MAPPs assay (Example 6.3.1) for the bispecific antibodies P1AG7282, P1AG7215, P1AG7191 and Pl AG7207. Clusters found in each domain are highlighted.
DETAILED DESCRIPTION OF THE INVENTION
Definitions
Unless defined otherwise, technical and scientific terms used herein have the same meaning as generally used in the art to which this invention belongs. For purposes of interpreting
this specification, the following definitions will apply and whenever appropriate, terms used in the singular will also include the plural and vice versa.
As used herein, the term "antigen binding molecule" refers in its broadest sense to a molecule that specifically binds an antigenic determinant. Examples of antigen binding molecules are antibodies, multispecific antibodies (e.g., bispecific antibodies), antibody fragments and scaffold antigen binding proteins.
As used herein, the term “antigen binding domain that binds to a tumor-associated antigen” or "moiety capable of specific binding to a tumor-associated antigen" refers to a polypeptide molecule that specifically binds to the tumor-associated antigen BCMA. In one aspect, the antigen binding domain is able to activate signaling through BCMA. In a particular aspect, the antigen binding domain is able to direct the entity to which it is attached (e.g. the CD28 antibody) to a BCMA-expressing cell, for example to a specific type of tumor cell. Antigen binding domains capable of specific binding to BCMA include antibodies and fragments thereof as further defined herein. In addition, antigen binding domains capable of specific binding to a tumor-associated antigen may include scaffold antigen binding proteins as further defined herein, e.g. binding domains which are based on designed repeat proteins or designed repeat domains (see e.g. WO 2002/020565).
In relation to an antigen binding molecule, i.e. an antibody or fragment thereof, the term "antigen binding domain" refers to the part of the molecule that comprises the area which specifically binds to and is complementary to part or all of an antigen. An antigen binding domain capable of specific antigen binding may be provided, for example, by one or more antibody variable domains (also called antibody variable regions). Particularly, an antigen binding domain capable of specific antigen binding comprises an antibody light chain variable region (VL) and an antibody heavy chain variable region (VH). In another aspect, the "antigen binding domain capable of specific binding to a tumor-associated antigen " can also be a Fab fragment or a crossFab fragment. As used herein, the terms “first”, “second” or “third” with respect to antigen binding domains etc., are used for convenience of distinguishing when there is more than one of each type of moiety. Use of these terms is not intended to confer a specific order or orientation of the moiety unless explicitly so stated.
The term "antibody" herein is used in the broadest sense and encompasses various antibody structures, including but not limited to monoclonal antibodies, polyclonal antibodies, monospecific and multispecific antibodies (e.g., bispecific antibodies), and antibody fragments so long as they exhibit the desired antigen-binding activity.
The term “monoclonal antibody” as used herein refers to an antibody obtained from a population of substantially homogeneous antibodies, i.e., the individual antibodies comprising the population are identical and/or bind the same epitope, except for possible variant antibodies, e.g. containing naturally occurring mutations or arising during production of a monoclonal antibody preparation, such variants generally being present in minor amounts. In contrast to
polyclonal antibody preparations, which typically include different antibodies directed against different determinants (epitopes), each monoclonal antibody of a monoclonal antibody preparation is directed against a single determinant on an antigen.
The term “monospecific” antibody as used herein denotes an antibody that has one or more binding sites each of which bind to the same epitope of the same antigen. The term “bispecific” means that the antigen binding molecule is able to specifically bind to at least two distinct antigenic determinants. Typically, a bispecific antigen binding molecule comprises two antigen binding sites, each of which is specific for a different antigenic determinant. However, a bispecific antigen binding molecule may also comprise additional antigen binding sites which bind to further antigenic determinants. In certain aspects, the bispecific antigen binding molecule is capable of simultaneously binding two antigenic determinants, particularly two antigenic determinants expressed on two distinct cells or on the same cell. The term “bispecific” in accordance with the present invention thus may also include a trispecific molecule, e.g. a bispecific molecule comprising a CD28 antibody and two antigen binding domains directed to two different target cell antigens.
The term “valent” as used within the current application denotes the presence of a specified number of binding sites specific for one distinct antigenic determinant in an antigen binding molecule that are specific for one distinct antigenic determinant. As such, the terms “bivalent”, “tetravalent”, and “hexavalent” denote the presence of two binding sites, four binding sites, and six binding sites specific for a certain antigenic determinant, respectively, in an antigen binding molecule. In particular aspects of the invention, the bispecific antigen binding molecules according to the invention can be monovalent for a certain antigenic determinant, meaning that they have only one binding site for said antigenic determinant or they can be bivalent or tetravalent for a certain antigenic determinant, meaning that they have two binding sites or four binding sites, respectively, for said antigenic determinant.
The terms “full length antibody”, “intact antibody”, and “whole antibody” are used herein interchangeably to refer to an antibody having a structure substantially similar to a native antibody structure. “Native antibodies” refer to naturally occurring immunoglobulin molecules with varying structures. For example, native IgG-class antibodies are heterotetrameric glycoproteins of about 150,000 daltons, composed of two light chains and two heavy chains that are disulfide-bonded. From N- to C-terminus, each heavy chain has a variable region (VH), also called a variable heavy domain or a heavy chain variable domain, followed by three constant domains (CHI, CH2, and CH3), also called a heavy chain constant region. Similarly, from N- to C-terminus, each light chain has a variable region (VL), also called a variable light domain or a light chain variable domain, followed by a light chain constant domain (CL), also called a light chain constant region. The heavy chain of an antibody may be assigned to one of five types, called a (IgA), 5 (IgD), a (IgE), y (IgG), or p (IgM), some of which may be further divided into subtypes, e.g. yl (IgGl), y2 (IgG2), y3 (IgG3), y4 (IgG4), al (IgAl) and a2 (IgA2). The light
chain of an antibody may be assigned to one of two types, called kappa (K) and lambda (X), based on the amino acid sequence of its constant domain.
An "antibody fragment" refers to a molecule other than an intact antibody that comprises a portion of an intact antibody that binds the antigen to which the intact antibody binds. Examples of antibody fragments include but are not limited to Fv, Fab, Fab', Fab’-SH, F(ab')2; diabodies, triabodies, tetrabodies, crossFab fragments; linear antibodies; single-chain antibody molecules (e.g. scFv); and single domain antibodies. For a review of certain antibody fragments, see Hudson et al., Nat Med 9, 129-134 (2003). For a review of scFv fragments, see e.g. Pliickthun, in The Pharmacology of Monoclonal Antibodies, vol. 113, Rosenburg and Moore eds., Springer-Verlag, New York, pp. 269-315 (1994); see also WO 93/16185; and U.S. Patent Nos. 5,571,894 and 5,587,458. For discussion of Fab and F(ab')2 fragments comprising salvage receptor binding epitope residues and having increased in vivo half-life, see U.S. Patent No. 5,869,046. Diabodies are antibody fragments with two antigen-binding sites that may be bivalent or bispecific, see, for example, EP 404,097; WO 1993/01161; Hudson et al., Nat Med 9, 129-134 (2003); and Hollinger et al., Proc Natl Acad Sci USA 90, 6444-6448 (1993). Triabodies and tetrabodies are also described in Hudson et al., Nat Med 9, 129-134 (2003). Single-domain antibodies are antibody fragments comprising all or a portion of the heavy chain variable domain or all or a portion of the light chain variable domain of an antibody. In certain embodiments, a single-domain antibody is a human single-domain antibody (Domantis, Inc., Waltham, MA; see e.g. U.S. Patent No. 6,248,516 Bl). Antibody fragments can be made by various techniques, including but not limited to proteolytic digestion of an intact antibody as well as production by recombinant host cells (e.g. E. coli or phage), as described herein.
Papain digestion of intact antibodies produces two identical antigen-binding fragments, called “Fab” fragments containing each the heavy- and light-chain variable domains and also the constant domain of the light chain and the first constant domain (CHI) of the heavy chain. As used herein, Thus, the term “Fab fragment” or “Fab molecule” refers to an antibody fragment comprising a light chain fragment comprising a variable light chain (VL) domain and a constant domain of a light chain (CL), and a variable heavy chain (VH) domain and a first constant domain (CHI) of a heavy chain. Fab’ fragments differ from Fab fragments by the addition of a few residues at the carboxy terminus of the heavy chain CHI domain including one or more cysteins from the antibody hinge region. Fab’-SH are Fab’ fragments in which the cysteine residue(s) of the constant domains bear a free thiol group. Pepsin treatment yields an F(ab')2 fragment that has two antigen-combining sites (two Fab fragments) and a part of the Fc region. A “conventional Fab fragment” is comprised of a VL-CL light chain and a VH-CH1 heavy chain.
The term “crossFab fragment” or “xFab fragment” or “crossover Fab fragment” refers to a Fab fragment, wherein either the variable regions or the constant regions of the heavy and light chain are exchanged. Two different chain compositions of a crossover Fab molecule are possible and comprised in the bispecific antibodies of the invention: On the one hand, the variable regions
of the Fab heavy and light chain are exchanged, i.e. the crossover Fab molecule comprises a peptide chain composed of the light chain variable (VL) domain and the heavy chain constant domain (CHI), and a peptide chain composed of the heavy chain variable domain (VH) and the light chain constant domain (CL). This crossover Fab molecule is also referred to as CrossFab (VLVH). On the other hand, when the constant regions of the Fab heavy and light chain are exchanged, the crossover Fab molecule comprises a peptide chain composed of the heavy chain variable domain (VH) and the light chain constant domain (CL), and a peptide chain composed of the light chain variable domain (VL) and the heavy chain constant domain (CHI). This crossover Fab molecule is also referred to as CrossFab (CLCHI).
A “single chain Fab fragment” or “scFab” is a polypeptide consisting of an antibody heavy chain variable domain (VH), an antibody constant domain 1 (CHI), an antibody light chain variable domain (VL), an antibody light chain constant domain (CL) and a linker, wherein said antibody domains and said linker have one of the following orders in N-terminal to C-terminal direction: a) VH-CH1 -linker- VL-CL, b) VL-CL-linker-VH-CHl, c) VH-CL-linker-VL-CHl or d) VL-CH1 -linker- VH-CL; and wherein said linker is a polypeptide of at least 30 amino acids, preferably between 32 and 50 amino acids. Said single chain Fab fragments are stabilized via the natural disulfide bond between the CL domain and the CHI domain. In addition, these single chain Fab molecules might be further stabilized by generation of interchain disulfide bonds via insertion of cysteine residues (e.g. position 44 in the variable heavy chain and position 100 in the variable light chain according to Kabat numbering).
A “crossover single chain Fab fragment” or “x-scFab” is a is a polypeptide consisting of an antibody heavy chain variable domain (VH), an antibody constant domain 1 (CHI), an antibody light chain variable domain (VL), an antibody light chain constant domain (CL) and a linker, wherein said antibody domains and said linker have one of the following orders in N- terminal to C-terminal direction: a) VH-CL-linker-VL-CHl and b) VL-CH1 -linker- VH-CL; wherein VH and VL form together an antigen-binding site which binds specifically to an antigen and wherein said linker is a polypeptide of at least 30 amino acids. In addition, these x-scFab molecules might be further stabilized by generation of interchain disulfide bonds via insertion of cysteine residues (e.g. position 44 in the variable heavy chain and position 100 in the variable light chain according to Kabat numbering).
A “single-chain variable fragment (scFv)” is a fusion protein of the variable regions of the heavy (VH) and light chains (VL) of an antibody, connected with a short linker peptide of ten to about 25 amino acids. The linker is usually rich in glycine for flexibility, as well as serine or threonine for solubility, and can either connect the N-terminus of the VH with the C-terminus of the VL, or vice versa. This protein retains the specificity of the original antibody, despite removal of the constant regions and the introduction of the linker. scFv antibodies are, e.g. described in Houston, J.S., Methods in Enzymol. 203 (1991) 46-96). In addition, antibody fragments comprise single chain polypeptides having the characteristics of a VH domain, namely being
able to assemble together with a VL domain, or of a VL domain, namely being able to assemble together with a VH domain to a functional antigen binding site and thereby providing the antigen binding property of full length antibodies.
“Scaffold antigen binding proteins” are known in the art, for example, fibronectin and designed ankyrin repeat proteins (DARPins) have been used as alternative scaffolds for antigenbinding domains, see, e.g., Gebauer and Skerra, Engineered protein scaffolds as next-generation antibody therapeutics. Curr Opin Chem Biol 13:245-255 (2009) and Stumpp et al., Darpins: A new generation of protein therapeutics. Drug Discovery Today 13: 695-701 (2008). In one aspect of the invention, a scaffold antigen binding protein is selected from the group consisting of CTLA-4 (Evibody), Lipocalins (Anticalin), a Protein A-derived molecule such as Z-domain of Protein A (Affibody), an A-domain (Avimer/Maxibody), a serum transferrin (/ra//.s-body); a designed ankyrin repeat protein (DARPin), a variable domain of antibody light chain or heavy chain (single-domain antibody, sdAb), a variable domain of antibody heavy chain (nanobody, aVH), VNAR fragments, a fibronectin (AdNectin), a C-type lectin domain (Tetranectin); a variable domain of a new antigen receptor beta-lactamase (VNAR fragments), a human gammacrystallin or ubiquitin (Affilin molecules); a kunitz type domain of human protease inhibitors, microbodies such as the proteins from the knottin family, peptide aptamers and fibronectin (adnectin). CTLA-4 (Cytotoxic T Lymphocyte-associated Antigen 4) is a CD28-family receptor expressed on mainly CD4+ T-cells. Its extracellular domain has a variable domain- like Ig fold. Loops corresponding to CDRs of antibodies can be substituted with heterologous sequence to confer different binding properties. CTLA-4 molecules engineered to have different binding specificities are also known as Evibodies (e.g. US7166697B1). Evibodies are around the same size as the isolated variable region of an antibody (e.g. a domain antibody). For further details, see Journal of Immunological Methods 248 (1-2), 31-45 (2001). Lipocalins are a family of extracellular proteins which transport small hydrophobic molecules such as steroids, bilins, retinoids and lipids. They have a rigid beta-sheet secondary structure with a number of loops at the open end of the conical structure which can be engineered to bind to different target antigens. Anticalins are between 160-180 amino acids in size, and are derived from lipocalins. For further details, see Biochim Biophys Acta 1482: 337-350 (2000), US7250297B1 and US20070224633. An affibody is a scaffold derived from Protein A of Staphylococcus aureus which can be engineered to bind to antigen. The domain consists of a three-helical bundle of approximately 58 amino acids. Libraries have been generated by randomization of surface residues. For further details, see Protein Eng. Des. Sei. 2004, 17, 455-462 and EP 1641818A1. Avimers are multidomain proteins derived from the A-domain scaffold family. The native domains of approximately 35 amino acids adopt a defined disulfide bonded structure. Diversity is generated by shuffling of the natural variation exhibited by the family of A-domains. For further details, see Nature Biotechnology 23(12), 1556 - 1561 (2005) and Expert Opinion on Investigational Drugs 16(6), 909-917 (June 2007). A transferrin is a monomeric serum transport glycoprotein.
Transferrins can be engineered to bind different target antigens by insertion of peptide sequences in a permissive surface loop. Examples of engineered transferrin scaffolds include the Transbody. For further details, see J. Biol. Chem 274, 24066-24073 (1999). Designed Ankyrin Repeat Proteins (DARPins) are derived from Ankyrin which is a family of proteins that mediate attachment of integral membrane proteins to the cytoskeleton. A single ankyrin repeat is a 33 residue motif consisting of two alpha-helices and a beta-turn. They can be engineered to bind different target antigens by randomizing residues in the first alpha-helix and a beta-turn of each repeat. Their binding interface can be increased by increasing the number of modules (a method of affinity maturation). For further details, see J. Mol. Biol. 332, 489-503 (2003), PNAS 100(4), 1700-1705 (2003) and J. Mol. Biol. 369, 1015-1028 (2007) and US20040132028A1. A singledomain antibody is an antibody fragment consisting of a single monomeric variable antibody domain. The first single domains were derived from the variable domain of the antibody heavy chain from camelids (nanobodies or VHH fragments). Furthermore, the term single-domain antibody includes an autonomous human heavy chain variable domain (aVH) or VNAR fragments derived from sharks. Fibronectin is a scaffold which can be engineered to bind to antigen. Adnectins consists of a backbone of the natural amino acid sequence of the 10th domain of the 15 repeating units of human fibronectin type III (FN3). Three loops at one end of the betasandwich can be engineered to enable an Adnectin to specifically recognize a therapeutic target of interest. For further details, see Protein Eng. Des. Sei. 18, 435- 444 (2005), US20080139791, W02005056764 and US6818418B1. Peptide aptamers are combinatorial recognition molecules that consist of a constant scaffold protein, typically thioredoxin (TrxA) which contains a constrained variable peptide loop inserted at the active site. For further details, see Expert Opin. Biol. Ther. 5, 783-797 (2005). Microbodies are derived from naturally occurring microproteins of 25-50 amino acids in length which contain 3-4 cysteine bridges - examples of microproteins include KalataBI and conotoxin and knottins. The microproteins have a loop which can beengineered to include upto 25 amino acids without affecting the overall fold of the microprotein. For further details of engineered knottin domains, see W02008098796.
An “antibody that binds to the same epitope” as a reference molecule refers to an antibody that blocks binding of the reference molecule to its antigen in a competition assay by 50% or more, and conversely, the reference molecule blocks binding of the antigen binding molecule to its antigen in a competition assay by 50% or more.
The term "antigen binding domain" refers to the part of an antigen binding molecule that comprises the area which specifically binds to and is complementary to part or all of an antigen. Where an antigen is large, an antigen binding molecule may only bind to a particular part of the antigen, which part is termed an epitope. An antigen binding domain may be provided by, for example, one or more variable domains (also called variable regions). Preferably, an antigen binding domain comprises an antibody light chain variable domain (VL) and an antibody heavy chain variable domain (VH).
As used herein, the term "antigenic determinant" is synonymous with "antigen" and "epitope," and refers to a site (e.g. a contiguous stretch of amino acids or a conformational configuration made up of different regions of non-contiguous amino acids) on a polypeptide macromolecule to which an antigen binding moiety binds, forming an antigen binding moiety- antigen complex. Useful antigenic determinants can be found, for example, on the surfaces of tumor cells, on the surfaces of virus-infected cells, on the surfaces of other diseased cells, on the surface of immune cells, free in blood serum, and/or in the extracellular matrix (ECM). The proteins useful as antigens herein can be any native form the proteins from any vertebrate source, including mammals such as primates (e.g. humans) and rodents (e.g. mice and rats), unless otherwise indicated. In a particular embodiment the antigen is a human protein. Where reference is made to a specific protein herein, the term encompasses the “full-length”, unprocessed protein as well as any form of the protein that results from processing in the cell. The term also encompasses naturally occurring variants of the protein, e.g. splice variants or allelic variants.
By "specific binding" is meant that the binding is selective for the antigen and can be discriminated from unwanted or non-specific interactions. The ability of an antigen binding molecule to bind to a specific antigen can be measured either through an enzyme-linked immunosorbent assay (ELISA) or other techniques familiar to one of skill in the art, e.g. Surface Plasmon Resonance (SPR) technique (analyzed on a BIAcore instrument) (Liljeblad et al., Glyco J 17, 323-329 (2000)), and traditional binding assays (Heeley, Endocr Res 28, 217-229 (2002)). In one embodiment, the extent of binding of an antigen binding molecule to an unrelated protein is less than about 10% of the binding of the antigen binding molecule to the antigen as measured, e.g. by SPR. In certain embodiments, an molecule that binds to the antigen has a dissociation constant (Kd) of < 1 pM, < 100 nM, < 10 nM, < 1 nM, < 0.1 nM, < 0.01 nM, or < 0.001 nM (e.g. 10'8 M or less, e.g. from 10'8 M to 10'13 M, e.g. from 10'9 M to 10'13 M).
“Affinity” or “binding affinity” refers to the strength of the sum total of non-covalent interactions between a single binding site of a molecule (e.g. an antibody) and its binding partner (e.g. an antigen). Unless indicated otherwise, as used herein, “binding affinity” refers to intrinsic binding affinity which reflects a 1 : 1 interaction between members of a binding pair (e.g. antibody and antigen). The affinity of a molecule X for its partner Y can generally be represented by the dissociation constant (Kd), which is the ratio of dissociation and association rate constants (koff and kon, respectively). Thus, equivalent affinities may comprise different rate constants, as long as the ratio of the rate constants remains the same. Affinity can be measured by common methods known in the art, including those described herein. A particular method for measuring affinity is Surface Plasmon Resonance (SPR).
An “activating T cell antigen” as used herein refers to an antigenic determinant expressed on the surface of a T lymphocyte, particularly a cytotoxic T lymphocyte, which is capable of inducing T cell activation upon interaction with an antibody. Specifically, interaction of an antibody with an activating T cell antigen may induce T cell activation by triggering the
signaling cascade of the T cell receptor complex. In a particular embodiment the activating T cell antigen is CD3, particularly the epsilon subunit of CD3 (see UniProt no. P07766 (version 189), NCBI RefSeq no. NP 000724.1, SEQ ID NO: 167 for the human sequence; or UniProt no. Q95LI5 (version 49), NCBI GenBank no. BAB71849.1, SEQ ID NO: 168 for the cynomolgus [Macaca fascicularis] sequence).
“T cell activation” as used herein refers to one or more cellular response of a T lymphocyte, particularly a cytotoxic T lymphocyte, selected from proliferation, differentiation, cytokine secretion, cytotoxic effector molecule release, cytotoxic activity, and expression of activation markers. Suitable assays to measure T cell activation are known in the art and described herein.
The term “T cell effector functions” refers to the activities of T cells that play a key role in the adaptive immune system. T cells are responsible for initiating and coordinating the body’s immune response against foreign invaders, such as viruses or bacteria as well as tumor cells. Effector functions refer to the various activities carried out by T cells to eliminate these offenders, which include releasing cytokines, stimulating other cells, and directly attacking and eliminating infected cells.
A “tumor-associated antigen” or TAA as used herein refers to an antigenic determinant presented on the surface of a target cell, for example a cell in a tumor such as a cancer cell, a cell of the tumor stroma, a malignant B lymphocyte or a melanoma cell. In certain aspects, the target cell antigen is an antigen on the surface of a tumor cell. In one particular aspect, TAA is BCMA.
The term “BCMA” refers to B cell maturation antigen, also termed tumor necrosis factor receptor superfamily member 17 (TNFRS17) or CD269, and is a type III transmembrane protein without a signal-peptide and containing cysteine-rich extracellular domains. Ligands for BCMA include B cell activating factor (BAFF) and a proliferation-inducing ligand (APRIL), of which APRIL has a higher affinity for BCMA. BCMA is preferentially expressed by mature B lymphocytes, with minimal expression in hematopoietic stem cells or non-hematopoietic tissue, and is essential for the survival of long-lived bone marrow plasma cells. Membrane-bound BCMA can undergo gamma-secretase mediated shedding from the cell surface, leading to circulation of soluble BCMA (sBCMA) and reduced activation of surface BCMA by APRIL and BAFF. BCMA is overexpressed at significantly higher levels in all patient MM cells but not on other normal tissues except normal plasma cells. BCMA, along with two related TNFR superfamily B-cell activation factor receptor (BAFF-R) and transmembrane activator and calcium modulator and cyclophilin ligand interactor (TACI), critically regulate B cell proliferation and survival, as well as maturation and differentiation into plasma cells. These three functionally related receptors support long-term survival of B cells at different stages of development by binding to BAFF and/or APRIL, their cognate ligands. BCMA as used herein refers to any BCMA protein from any vertebrate source, including mammals such as primates (e.g. humans) non-human primates (e.g. cynomolgus monkeys) and rodents (e.g. mice and rats),
unless otherwise indicated. The amino acid sequence of human BCMA is shown in UniProt (www.uniprot.org) accession no. Q02223 (SEQ ID NO: 99).
The term “CD28” (Cluster of differentiation 28, Tp44) refers to any CD28 protein from any vertebrate source, including mammals such as primates (e.g. humans) non-human primates (e.g. cynomolgus monkeys) and rodents (e.g. mice and rats), unless otherwise indicated. CD28 is expressed on T cells and provides co-stimulatory signals required for T cell activation and survival. T cell stimulation through CD28 in addition to the T-cell receptor (TCR) can provide a potent signal for the production of various interleukins. CD28 is the receptor for CD80 (B7.1) and CD86 (B7.2) proteins and is the only B7 receptor constitutively expressed on naive T cells. The amino acid sequence of human CD28 is shown in UniProt (www.uniprot.org) accession no. Pl 0747 (SEQ ID NO: 100).
An “agonistic antibody” refers to an antibody that comprises an agonistic function against a given receptor. In general, when an agonist ligand (factor) binds to a receptor, the tertiary structure of the receptor protein changes, and the receptor is activated (when the receptor is a membrane protein, a cell growth signal or such is usually transducted). If the receptor is a dimerforming type, an agonistic antibody can dimerize the receptor at an appropriate distance and angle, thus acting similarly to a ligand. An appropriate anti-receptor antibody can mimic dimerization of receptors performed by ligands, and thus can become an agonistic antibody.
A “CD28 agonistic antibody” or “CD28 conventional agonistic antibody” is an antibody that mimics CD28 natural ligands (CD80 or CD86) in their role to enhance T cell activation in presence of a T cell receptor signal (“signal 2”). A T cell needs two signals to become fully activated. Under physiological conditions “signal 1” arises from the interaction of T cell receptor (TCR) molecules with peptide/major histocompatibility complex (MHC) complexes on antigen presenting cells (APCs) and “signal 2” is provided by engagement of a costimulatory receptor, e.g. CD28. A CD28 agonistic antibody is able to costimulate T cells (signal 2). It is also able to induce T cell proliferation and cytokine secretion in combination with a molecule with specificity for the TCR complex, however the CD28 agonistic antibody is not capable of fully activating T cells without additional stimulation of the TCR. There is however a subclass of CD28 specific antigen binding molecules, the so-called CD28 superagonistic antibodies. A “CD28 superagonistic antibody” is a CD28 antibody which is capable of fully activating T cells without additional stimulation of the TCR. A CD28 superagonistic anitbody is capable to induce T cell proliferation and cytokine secretion without prior T cell activation (signal 1).
The term “variable domain” or “variable region” refers to the domain of an antibody heavy or light chain that is involved in binding the antigen binding molecule to antigen. The variable domains of the heavy chain and light chain (VH and VL, respectively) of a native antibody generally have similar structures, with each domain comprising four conserved framework regions (FRs) and three hypervariable regions (HVRs). See, e.g., Kindt et al., Kuby
Immunology, 6th ed., W.H. Freeman and Co., page 91 (2007). A single VH or VL domain may be sufficient to confer antigen-binding specificity.
The term “hypervariable region” or “HVR” as used herein refers to each of the regions of an antigen binding variable domain which are hypervariable in sequence and which determine antigen binding specificity, for example “complementarity determining regions” (“CDRs”). Generally, antigen binding domains comprise six CDRs: three in the VH (CDR-H1, CDR-H2, CDR-H3), and three in the VL (CDR-L1, CDR-L2, CDR-L3). Exemplary CDRs herein include:
(a) hypervariable loops occurring at amino acid residues 26-32 (LI), 50-52 (L2), 91-96 (L3), 26-32 (Hl), 53-55 (H2), and 96-101 (H3) (Chothia and Lesk, J. Mol. Biol. 196:901-917 (1987));
(b) CDRs occurring at amino acid residues 24-34 (LI), 50-56 (L2), 89-97 (L3), 31 -35b (Hl), 50-65 (H2), and 95-102 (H3) (Kabat et al., Sequences of Proteins of Immunological Interest, 5th Ed. Public Health Service, National Institutes of Health, Bethesda, MD (1991)); and
(c) antigen contacts occurring at amino acid residues 27c-36 (LI), 46-55 (L2), 89-96 (L3), 30-35b (Hl), 47-58 (H2), and 93-101 (H3) (MacCallum et al. J. Mol. Biol. 262: 732-745 (1996)).
Unless otherwise indicated, the CDRs are determined according to Kabat et al., supra. One of skill in the art will understand that the CDR designations can also be determined according to Chothia, supra, McCallum, supra, or any other scientifically accepted nomenclature. Kabat et al. also defined a numbering system for variable region sequences that is applicable to any antibody. One of ordinary skill in the art can unambiguously assign this system of "Kabat numbering" to any variable region sequence, without reliance on any experimental data beyond the sequence itself. As used herein, "Kabat numbering" refers to the numbering system set forth by Kabat et al., U.S. Dept, of Health and Human Services, "Sequence of Proteins of Immunological Interest" (1983). Unless otherwise specified, references to the numbering of specific amino acid residue positions in an antibody variable region are according to the Kabat numbering system.
As used herein, the term “affinity matured” in the context of antigen binding molecules (e.g., antibodies) refers to an antigen binding molecule that is derived from a reference antigen binding molecule, e.g., by mutation, binds to the same antigen, preferably binds to the same epitope, as the reference antibody; and has a higher affinity for the antigen than that of the reference antigen binding molecule. Affinity maturation generally involves modification of one or more amino acid residues in one or more CDRs of the antigen binding molecule. Typically, the affinity matured antigen binding molecule binds to the same epitope as the initial reference antigen binding molecule.
"Framework" or "FR" refers to variable domain residues other than hypervariable region (HVR) residues. The FR of a variable domain generally consists of four FR domains: FR1, FR2, FR3, and FR4. Accordingly, the HVR and FR sequences generally appear in the following sequence in VH (or VL): FR1-H1(L1)-FR2-H2(L2)-FR3-H3(L3)-FR4.
An “acceptor human framework” for the purposes herein is a framework comprising the amino acid sequence of a light chain variable domain (VL) framework or a heavy chain variable domain (VH) framework derived from a human immunoglobulin framework or a human consensus framework, as defined below. An acceptor human framework “derived from” a human immunoglobulin framework or a human consensus framework may comprise the same amino acid sequence thereof, or it may contain amino acid sequence changes. In some embodiments, the number of amino acid changes are 10 or less, 9 or less, 8 or less, 7 or less, 6 or less, 5 or less, 4 or less, 3 or less, or 2 or less. In some embodiments, the VL acceptor human framework is identical in sequence to the VL human immunoglobulin framework sequence or human consensus framework sequence.
The term "chimeric" antibody refers to an antibody in which a portion of the heavy and/or light chain is derived from a particular source or species, while the remainder of the heavy and/or light chain is derived from a different source or species.
The “class” of an antibody refers to the type of constant domain or constant region possessed by its heavy chain. There are five major classes of antibodies: IgA, IgD, IgE, IgG, and IgM, and several of these may be further divided into subclasses (isotypes), e.g. IgGi, IgG?, IgG?, IgG4, IgAi, and IgA?. The heavy chain constant domains that correspond to the different classes of immunoglobulins are called a, 5, 8, y, and p respectively..
A “humanized” antibody refers to a chimeric antibody comprising amino acid residues from non-human HVRs and amino acid residues from human FRs. In certain embodiments, a humanized antibody will comprise substantially all of at least one, and typically two, variable domains, in which all or substantially all of the HVRs (e.g., CDRs) correspond to those of a non- human antibody, and all or substantially all of the FRs correspond to those of a human antibody. A humanized antibody optionally may comprise at least a portion of an antibody constant region derived from a human antibody. A “humanized form” of an antibody, e.g., a non-human antibody, refers to an antibody that has undergone humanization. Other forms of "humanized antibodies" encompassed by the present invention are those in which the constant region has been additionally modified or changed from that of the original antibody to generate the properties according to the invention, especially in regard to Clq binding and/or Fc receptor (FcR) binding.
A “human” antibody is one which possesses an amino acid sequence which corresponds to that of an antibody produced by a human or a human cell or derived from a non-human source that utilizes human antibody repertoires or other human antibody-encoding sequences. This definition of a human antibody specifically excludes a humanized antibody comprising non- human antigen-binding residues. Particularly, a “human” or “humanized” antibody comprises a constant region of human origin, particularly of the IgG isotype, more particularly of the IgGl isotype, comprising a human CHI, CH2, CH3 and/or CL domain.
The term "CL domain" denotes the constant part of an antibody light chain polypeptide. Exemplary sequences of human constant domains are given in SEQ ID Nos: 101 and 102 (human kappa and lambda CL domains, respectively).
The term "CHI domain" denotes the part of an antibody heavy chain polypeptide that extends approximately from EU position 118 to EU position 215 (EU numbering system according to Kabat). In one aspect, a CHI domain has the amino acid sequence of ASTKGPSVFP LAPSSKSTSG GTAALGCLVK DYFPEPVTVS WNSGALTSGV HTFPAVLQSS GLYSLSSVVT VPSSSLGTQT YICNVNHKPS NTKVDKKV (SEQ ID NO: 103). Usually, a segment having the amino acid sequence of EPKSC (SEQ ID NO: 104) is following to link the CHI domain to the hinge region.
The term "hinge region" denotes the part of an antibody heavy chain polypeptide that joins in a wild-type antibody heavy chain the CHI domain and the CH2 domain, e. g. from about position 216 to about position 230 according to the EU number system of Kabat, or from about position 226 to about position 230 according to the EU number system of Kabat. The hinge regions of other IgG subclasses can be determined by aligning with the hinge-region cysteine residues of the IgGl subclass sequence. The hinge region is normally a dimeric molecule consisting of two polypeptides with identical amino acid sequence. The hinge region generally comprises up to 25 amino acid residues and is flexible allowing the associated target binding sites to move independently. The hinge region can be subdivided into three domains: the upper, the middle, and the lower hinge domain (see e.g. Roux, et al., J. Immunol. 161 (1998) 4083).
In one aspect, the hinge region has the amino acid sequence DKTHTCPXCP (SEQ ID NO: 105), wherein X is either S or P. In one aspect, the hinge region has the amino acid sequence HTCPXCP (SEQ ID NO: 106), wherein X is either S or P. In one aspect, the hinge region has the amino acid sequence CPXCP (SEQ ID NO: 107), wherein X is either S or P.
The term “Fc domain” or “Fc region” herein is used to define a C-terminal region of an antibody heavy chain that contains at least a portion of the constant region. The term includes native sequence Fc regions and variant Fc regions. In the context of a molecule already defined by a Fab fragment (including the CHI domain), the term “Fc domain” may only refer to an IgG CH2 and an IgG CH3 domain.
The “CH2 domain” of a human IgG Fc region usually extends from an amino acid residue at about EU position 231 to an amino acid residue at about EU position 340 (EU numbering system according to Kabat). In one aspect, a CH2 domain has the amino acid sequence of APELLGGPSV FLFPPKPKDT LMISRTPEVT CVWDVSHEDP EVKFNWYVDG VEVHNAKTKP REEQESTYRW SVLTVLHQDW LNGKEYKCKV SNKALPAPIE KTISKAK (SEQ ID NO: 108). The CH2 domain is unique in that it is not closely paired with another domain. Rather, two N-linked branched carbohydrate chains are interposed between the two CH2 domains of an intact native Fc-region. It has been speculated that the carbohydrate may provide a substitute for the domaindomain pairing and help stabilize the CH2 domain. Burton, Mol. Immunol. 22 (1985) 161-206.
In one embodiment, a carbohydrate chain is attached to the CH2 domain. The CH2 domain herein may be a native sequence CH2 domain or variant CH2 domain.
The “CH3 domain” comprises the stretch of residues C-terminal to a CH2 domain in an Fc region denotes the part of an antibody heavy chain polypeptide that extends approximately from EU position 341 to EU position 446 (EU numbering system according to Kabat). In one aspect, the CH3 domain has the amino acid sequence of GQPREPQVYT LPPSRDELTK NQVSLTCLVK GFYPSDIAVE WESNGQPENN YKTTPPVLDS DGSFFLYSKL TVDKSRWQQG NVFSCSVMHE ALHNHYTQKS LSLSPGK (SEQ ID NO: 109). The CH3 region herein may be a native sequence CH3 domain or a variant CH3 domain (e.g. a CH3 domain with an introduced “protuberance” (“knob”) in one chain thereof and a corresponding introduced “cavity” (“hole”) in the other chain thereof; see US Patent No. 5,821,333, expressly incorporated herein by reference). Such variant CH3 domains may be used to promote heterodimerization of two non-identical antibody heavy chains as herein described. In one embodiment, a human IgG heavy chain Fc region extends from Cys226, or from Pro230, to the carboxyl -terminus of the heavy chain. However, the C-terminal lysine (Lys447) of the Fc region may or may not be present. Unless otherwise specified herein, numbering of amino acid residues in the Fc region or constant region is according to the EU numbering system, also called the EU index, as described in Kabat et al., Sequences of Proteins of Immunological Interest, 5th Ed. Public Health Service, National Institutes of Health, Bethesda, MD, 1991.
The “knob-into-hole” technology is described e.g. in US 5,731,168; US 7,695,936; Ridgway et al., Prot Eng 9, 617-621 (1996) and Carter, J Immunol Meth 248, 7-15 (2001). Generally, the method involves introducing a protuberance (“knob”) at the interface of a first polypeptide and a corresponding cavity (“hole”) in the interface of a second polypeptide, such that the protuberance can be positioned in the cavity so as to promote heterodimer formation and hinder homodimer formation. Protuberances are constructed by replacing small amino acid side chains from the interface of the first polypeptide with larger side chains (e.g. tyrosine or tryptophan). Compensatory cavities of identical or similar size to the protuberances are created in the interface of the second polypeptide by replacing large amino acid side chains with smaller ones (e.g. alanine or threonine). The protuberance and cavity can be made by altering the nucleic acid encoding the polypeptides, e.g. by site-specific mutagenesis, or by peptide synthesis. In a specific embodiment a knob modification comprises the amino acid substitution T366W in one of the two subunits of the Fc domain, and the hole modification comprises the amino acid substitutions T366S, L368A and Y407V in the other one of the two subunits of the Fc domain. In a further specific embodiment, the subunit of the Fc domain comprising the knob modification additionally comprises the amino acid substitution S354C, and the subunit of the Fc domain comprising the hole modification additionally comprises the amino acid substitution Y349C. Introduction of these two cysteine residues results in the formation of a disulfide bridge between
the two subunits of the Fc region, thus further stabilizing the dimer (Carter, J Immunol Methods 248, 7-15 (2001)).
A "region equivalent to the Fc region of an immunoglobulin" is intended to include naturally occurring allelic variants of the Fc region of an immunoglobulin as well as variants having alterations which produce substitutions, additions, or deletions but which do not decrease substantially the ability of the immunoglobulin to mediate effector functions (such as antibodydependent cellular cytotoxicity). For example, one or more amino acids can be deleted from the N-terminus or C-terminus of the Fc region of an immunoglobulin without substantial loss of biological function. Such variants can be selected according to general rules known in the art so as to have minimal effect on activity (see, e.g., Bowie, J. U. et al., Science 247: 1306-10 (1990)).
The term “wild-type Fc domain” denotes an amino acid sequence identical to the amino acid sequence of an Fc domain found in nature. Wild-type human Fc domains include a native human IgGl Fc-region (non- A and A allotypes), native human IgG2 Fc-region, native human IgG3 Fc-region, and native human IgG4 Fc-region as well as naturally occurring variants thereof. Wild-type Fc-regions are denoted in SEQ ID NO: 110 (IgGl, Caucasian allotype), SEQ ID NO: 111 (IgGl, afroamerican allotype), SEQ ID NO: 112 (IgG2), SEQ ID NO: 113 (IgG3) and SEQ ID NO: 114 (IgG4).
The term “variant (human) Fc domain” denotes an amino acid sequence which differs from that of a “wild-type” (human) Fc domain amino acid sequence by virtue of at least one “amino acid mutation”. In one aspect, the variant Fc-region has at least one amino acid mutation compared to a native Fc-region, e.g. from about one to about ten amino acid mutations, and in one aspect from about one to about five amino acid mutations in a native Fc-region. In one aspect, the (variant) Fc-region has at least about 95 % homology with a wild-type Fc-region. A specific variant Fc domain disclosed herein is the human IgGl heavy chain constant region with mutations L234A, L235A and P329G comprising the amino acid sequence of SEQ ID NO: 115.
The term “effector functions” refers to those biological activities attributable to the Fc region of an antibody, which vary with the antibody isotype. Examples of antibody effector functions include: Clq binding and complement dependent cytotoxicity (CDC), Fc receptor binding, antibody-dependent cell-mediated cytotoxicity (ADCC), antibody-dependent cellular phagocytosis (ADCP), cytokine secretion, immune complex -mediated antigen uptake by antigen presenting cells, down regulation of cell surface receptors (e.g. B cell receptor), and B cell activation.
Fc receptor binding dependent effector functions can be mediated by the interaction of the Fc-region of an antibody with Fc receptors (FcRs), which are specialized cell surface receptors on hematopoietic cells. Fc receptors belong to the immunoglobulin superfamily, and have been shown to mediate both the removal of antibody-coated pathogens by phagocytosis of immune complexes, and the lysis of erythrocytes and various other cellular targets (e.g. tumor cells) coated with the corresponding antibody, via antibody dependent cell mediated cytotoxicity
(ADCC) (see e.g. Van de Winkel, J.G. and Anderson, C.L., J. Leukoc. Biol. 49 (1991) 511-524). FcRs are defined by their specificity for immunoglobulin isotypes: Fc receptors for IgG antibodies are referred to as FcyR. Fc receptor binding is described e.g. in Ravetch, J.V. and Kinet, J.P., Annu. Rev. Immunol. 9 (1991) 457-492; Capel, P.J., et al., Immunomethods 4 (1994) 25-34; de Haas, M., et al., J. Lab. Clin. Med. 126 (1995) 330-341; and Gessner, J.E., et al., Ann. Hematol. 76 (1998) 231-248.
Cross-linking of receptors for the Fc-region of IgG antibodies (FcyR) triggers a wide variety of effector functions including phagocytosis, antibody-dependent cellular cytotoxicity, and release of inflammatory mediators, as well as immune complex clearance and regulation of antibody production. In humans, three classes of FcyR have been characterized, which are:
- FcyRI (CD64) binds monomeric IgG with high affinity and is expressed on macrophages, monocytes, neutrophils and eosinophils. Modification in the Fc-region IgG at least at one of the amino acid residues E233-G236, P238, D265, N297, A327 and P329 (numbering according to EU index of Kabat) reduce binding to FcyRI. IgG2 residues at positions 233-236, substituted into IgGl and IgG4, reduced binding to FcyRI by 103-fold and eliminated the human monocyte response to antibody-sensitized red blood cells (Armour, K.L., et al., Eur. J. Immunol. 29 (1999) 2613-2624).
-FcyRII (CD32) binds complexed IgG with medium to low affinity and is widely expressed. This receptor can be divided into two sub-types, FcyRIIA and FcyRIIB. FcyRIIA is found on many cells involved in killing (e.g. macrophages, monocytes, neutrophils) and seems able to activate the killing process. FcyRIIB seems to play a role in inhibitory processes and is found on B cells, macrophages and on mast cells and eosinophils. On B-cells it seems to function to suppress further immunoglobulin production and isotype switching to, for example, the IgE class. On macrophages, FcyRIIB acts to inhibit phagocytosis as mediated through FcyRIIA. On eosinophils and mast cells the B-form may help to suppress activation of these cells through IgE binding to its separate receptor. Reduced binding for FcyRIIA is found e.g. for antibodies comprising an IgG Fc-region with mutations at least at one of the amino acid residues E233- G236, P238, D265, N297, A327, P329, D270, Q295, A327, R292, and K414 (numbering according to EU index of Kabat).
- FcyRIII (CD 16) binds IgG with medium to low affinity and exists as two types. FcyRIIIA is found on NK cells, macrophages, eosinophils and some monocytes and T cells and mediates ADCC. FcyRIIIB is highly expressed on neutrophils. Reduced binding to FcyRIIIA is found e.g. for antibodies comprising an IgG Fc-region with mutation at least at one of the amino acid residues E233-G236, P238, D265, N297, A327, P329, D270, Q295, A327, S239, E269, E293, Y296, V303, A327, K338 and D376 (numbering according to EU index of Kabat).
Mapping of the binding sites on human IgGl for Fc receptors, the above mentioned mutation sites and methods for measuring binding to FcyRI and FcyRIIA are described in Shields, R.L., et al. J. Biol. Chem. 276 (2001) 6591-6604.
The term “ADCC” or “antibody-dependent cellular cytotoxicity” is an immune mechanism leading to lysis of antibody-coated target cells by immune effector cells. The target cells are cells to which antibodies or derivatives thereof comprising an Fc region specifically bind, generally via the protein part that is N-terminal to the Fc region. As used herein, the term “reduced ADCC” is defined as either a reduction in the number of target cells that are lysed in a given time, at a given concentration of antibody in the medium surrounding the target cells, by the mechanism of ADCC defined above, and/or an increase in the concentration of antibody in the medium surrounding the target cells, required to achieve the lysis of a given number of target cells in a given time, by the mechanism of ADCC. The reduction in ADCC is relative to the ADCC mediated by the same antibody produced by the same type of host cells, using the same standard production, purification, formulation and storage methods (which are known to those skilled in the art), but that has not been engineered. For example, the reduction in ADCC mediated by an antibody comprising in its Fc domain an amino acid substitution that reduces ADCC, is relative to the ADCC mediated by the same antibody without this amino acid substitution in the Fc domain. Suitable assays to measure ADCC are well known in the art (see e.g. PCT publication no. WO 2006/082515 or PCT publication no. WO 2012/130831). For example, the capacity of the antibody to induce the initial steps mediating ADCC is investigated by measuring their binding to Fey receptors expressing cells, such as cells, recombinantly expressing FcyRI and/or FcyRIIA or NK cells (expressing essentially FcyRIIIA). In particular, binding to FcyR on NK cells is measured.
An “activating Fc receptor” is an Fc receptor that following engagement by an Fc region of an antibody elicits signaling events that stimulate the receptor-bearing cell to perform effector functions. Activating Fc receptors include FcyRIIIa (CD16a), FcyRI (CD64), FcyRIIa (CD32), and FcaRI (CD89). A particular activating Fc receptor is human FcyRIIIa (SEQ ID NO: 116, UniProt accession no. P08637, version 141).
An “ectodomain” is the domain of a membrane protein that extends into the extracellular space (i.e. the space outside the target cell). Ectodomains are usually the parts of proteins that initiate contact with surfaces, which leads to signal transduction.
The term “peptide linker” refers to a peptide comprising one or more amino acids, typically about 2 to 20 amino acids. Peptide linkers are known in the art or are described herein. Suitable, non-immunogenic linker peptides are, for example, (G4S)n, (SG4)n or G4(SG4)n peptide linkers, wherein “n” is generally a number between 1 and 5, typically between 2 and 4, in particular 2, i.e. the peptides selected from the group consisting of GGGGS (SEQ ID NO: 117), GGGGSGGGG (SEQ ID NO: 198), GGGGSGGGGS (SEQ ID NO: 118), SGGGGSGGGG (SEQ ID NO: 119) and GGGGS GGGGSGGGG (SEQ ID NO: 120), but also include the sequences GSPGSSSSGS (SEQ ID NO: 121), (G4S)3 (SEQ ID NO: 122), (G4S)4 (SEQ ID NO: 123), GSGSGSGS (SEQ ID NO: 124), GSGSGNGS (SEQ ID NO: 125), GGSGSGSG (SEQ ID NO: 126), GGSGSG (SEQ ID NO: 127), GGSG (SEQ ID NO: 128), GGSGNGSG (SEQ ID
NO:129), GGNGSGSG (SEQ ID NO: 130) and GGNGSG (SEQ ID NO:131). Peptide linkers of particular interest are (G4S) (SEQ ID NO: 117), GGGGSGGGG (SEQ ID NO: 198), (G4S)2 or GGGGSGGGGS (SEQ ID NO: 118), (G4S)3 (SEQ ID NO: 122) and (G4S)4 (SEQ ID NO: 123).
The term ’’amino acid” as used within this application denotes the group of naturally occurring carboxy a-amino acids comprising alanine (three letter code: ala, one letter code: A), arginine (arg, R), asparagine (asn, N), aspartic acid (asp, D), cysteine (cys, C), glutamine (gin, Q), glutamic acid (glu, E), glycine (gly, G), histidine (his, H), isoleucine (ile, I), leucine (leu, L), lysine (lys, K), methionine (met, M), phenylalanine (phe, F), proline (pro, P), serine (ser, S), threonine (thr, T), tryptophan (trp, W), tyrosine (tyr, Y), and valine (val, V).
By “fused” or “connected” is meant that the components (e.g. a polypeptide and another polypeptide) are linked by peptide bonds, either directly or via one or more peptide linkers.
“Percent (%) amino acid sequence identity" with respect to a reference polypeptide (protein) sequence is defined as the percentage of amino acid residues in a candidate sequence that are identical with the amino acid residues in the reference polypeptide sequence, after aligning the sequences and introducing gaps, if necessary, to achieve the maximum percent sequence identity, and not considering any conservative substitutions as part of the sequence identity. Alignment for purposes of determining percent amino acid sequence identity can be achieved in various ways that are within the skill in the art, for instance, using publicly available computer software such as BLAST, BLAST-2, ALIGN. SAWI or Megalign (DNASTAR) software. Those skilled in the art can determine appropriate parameters for aligning sequences, including any algorithms needed to achieve maximal alignment over the full length of the sequences being compared. For purposes herein, however, % amino acid sequence identity values are generated using the sequence comparison computer program ALIGN-2. The ALIGN- 2 sequence comparison computer program was authored by Genentech, Inc., and the source code has been filed with user documentation in the U.S. Copyright Office, Washington D.C., 20559, where it is registered under U.S. Copyright Registration No. TXU510087. The ALIGN-2 program is publicly available from Genentech, Inc., South San Francisco, California, or may be compiled from the source code. The ALIGN-2 program should be compiled for use on a UNIX operating system, including digital UNIX V4.0D. All sequence comparison parameters are set by the ALIGN-2 program and do not vary. In situations where ALIGN-2 is employed for amino acid sequence comparisons, the % amino acid sequence identity of a given amino acid sequence A to, with, or against a given amino acid sequence B (which can alternatively be phrased as a given amino acid sequence A that has or comprises a certain % amino acid sequence identity to, with, or against a given amino acid sequence B) is calculated as follows: 100 times the fraction X/Y, where X is the number of amino acid residues scored as identical matches by the sequence alignment program ALIGN-2 in that program’s alignment of A and B, and where Y is the total number of amino acid residues in B. It will be appreciated that where the length of amino acid sequence A is not equal to the length of amino acid sequence B, the % amino acid sequence
identity of A to B will not equal the % amino acid sequence identity of B to A. Unless specifically stated otherwise, all % amino acid sequence identity values used herein are obtained as described in the immediately preceding paragraph using the ALIGN-2 computer program.
In certain embodiments, amino acid sequence variants of the BCMA antibodies or bispecific BCMA antibodies provided herein are contemplated. For example, it may be desirable to improve the binding affinity and/or other biological properties of the BCMA antibodies or bispecific BCMA antibodies. Amino acid sequence variants of the BCMA antibodies or bispecific BCMA antibodies may be prepared by introducing appropriate modifications into the nucleotide sequence encoding the molecules, or by peptide synthesis. Such modifications include, for example, deletions from, and/or insertions into and/or substitutions of residues within the amino acid sequences of the antibody. Any combination of deletion, insertion, and substitution can be made to arrive at the final construct, provided that the final construct possesses the desired characteristics, e.g., antigen-binding. Sites of interest for substitutional mutagenesis include the CDRs and Framework (FRs). Conservative substitutions are provided in Table B under the heading “Preferred Substitutions” and further described below in reference to amino acid side chain classes (1) to (6). Amino acid substitutions may be introduced into the molecule of interest and the products screened for a desired activity, e.g., retained/improved antigen binding, decreased immunogenicity, or improved ADCC or CDC.
Amino acids may be grouped according to common side-chain properties:
(1) hydrophobic: Norleucine, Met, Ala, Vai, Leu, He;
(2) neutral hydrophilic: Cys, Ser, Thr, Asn, Gin;
(3) acidic: Asp, Glu;
(4) basic: His, Lys, Arg;
(5) residues that influence chain orientation: Gly, Pro;
(6) aromatic: Trp, Tyr, Phe.
Non-conservative substitutions will entail exchanging a member of one of these classes for another class.
The term “amino acid sequence variants” includes substantial variants wherein there are amino acid substitutions in one or more hypervariable region residues of a parent antigen binding molecule (e.g. a humanized or human antibody). Generally, the resulting variant(s) selected for further study will have modifications (e.g., improvements) in certain biological properties (e.g., increased affinity, reduced immunogenicity) relative to the parent antigen binding molecule and/or will have substantially retained certain biological properties of the parent antigen binding molecule. An exemplary substitutional variant is an affinity matured antibody, which may be conveniently generated, e.g., using phage display-based affinity maturation techniques such as those described herein. Briefly, one or more CDR residues are mutated and the variant antigen binding molecules displayed on phage and screened for a particular biological activity (e.g. binding affinity). In certain embodiments, substitutions, insertions, or deletions may occur within one or more CDRs so long as such alterations do not substantially reduce the ability of the antigen binding molecule to bind antigen. For example, conservative alterations (e.g., conservative substitutions as provided herein) that do not substantially reduce binding affinity may be made in CDRs. A useful method for identification of residues or regions of an antibody that may be targeted for mutagenesis is called "alanine scanning mutagenesis" as described by Cunningham and Wells (1989) Science, 244: 1081-1085. In this method, a residue or group of target residues (e.g., charged residues such as Arg, Asp, His, Lys, and Glu) are identified and
replaced by a neutral or negatively charged amino acid (e.g., alanine or polyalanine) to determine whether the interaction of the antibody with antigen is affected. Further substitutions may be introduced at the amino acid locations demonstrating functional sensitivity to the initial substitutions. Alternatively, or additionally, a crystal structure of an antigen-antigen binding molecule complex to identify contact points between the antibody and antigen. Such contact residues and neighboring residues may be targeted or eliminated as candidates for substitution. Variants may be screened to determine whether they contain the desired properties.
Amino acid sequence insertions include amino- and/or carboxyl-terminal fusions ranging in length from one residue to polypeptides containing a hundred or more residues, as well as intrasequence insertions of single or multiple amino acid residues. Examples of insertions include BCMA antibodies or bispecific BCMA antibodies with a fusion to the N- or C-terminus to a polypeptide which increases the serum half-life of the CD28 antigen binding molecules.
In certain aspects, the BCMA antibodies or bispecific BCMA antibodies provided herein are altered to increase or decrease the extent to which the antibody is glycosylated. Glycosylation variants of the molecules may be conveniently obtained by altering the amino acid sequence such that one or more glycosylation sites is created or removed. Where the agonistic ICOS- binding molecule comprises an Fc domain, the carbohydrate attached thereto may be altered. Native antibodies produced by mammalian cells typically comprise a branched, biantennary oligosaccharide that is generally attached by an N-linkage to Asn297 of the CH2 domain of the Fc region. See, e.g., Wright et al. TIBTECH 15:26-32 (1997). The oligosaccharide may include various carbohydrates, e.g., mannose, N-acetyl glucosamine (GlcNAc), galactose, and sialic acid, as well as a fucose attached to a GlcNAc in the “stem” of the biantennary oligosaccharide structure. In some embodiments, modifications of the oligosaccharide in BCMA antibodies or bispecific BCMA antibodies may be made in order to create variants with certain improved properties. In one aspect, variants of BCMA antibodies or bispecific BCMA antibodies are provided having a carbohydrate structure that lacks fucose attached (directly or indirectly) to an Fc region. Such fucosylation variants may have improved ADCC function, see e.g. US Patent Publication Nos. US 2003/0157108 (Presta, L.) or US 2004/0093621 (Kyowa Hakko Kogyo Co., Ltd). Further variants of the BCMA antibodies or bispecific BCMA antibodies include those with bisected oligosaccharides, e.g., in which a biantennary oligosaccharide attached to the Fc region is bisected by GlcNAc. Such variants may have reduced fucosylation and/or improved ADCC function., see for example WO 2003/011878 (Jean-Mairet et al.); US Patent No. 6,602,684 (Umana et al.); and US 2005/0123546 (Umana et al ). Variants with at least one galactose residue in the oligosaccharide attached to the Fc region are also provided. Such antibody variants may have improved CDC function and are described, e.g., in WO 1997/30087 (Patel et al.); WO 1998/58964 (Raju, S.); and WO 1999/22764 (Raju, S.).
In certain embodiments, it may be desirable to create cysteine engineered variants of the CD28 antigen binding molecules of the invention, e.g., “thioMAbs,” in which one or more
residues of the molecule are substituted with cysteine residues. In particular embodiments, the substituted residues occur at accessible sites of the molecule. By substituting those residues with cysteine, reactive thiol groups are thereby positioned at accessible sites of the antibody and may be used to conjugate the antibody to other moieties, such as drug moieties or linker-drug moieties, to create an immunoconjugate. In certain embodiments, any one or more of the following residues may be substituted with cysteine: V205 (Kabat numbering) of the light chain; Al 18 (EU numbering) of the heavy chain; and S400 (EU numbering) of the heavy chain Fc region. Cysteine engineered antigen binding molecules may be generated as described, e.g., in U.S. Patent No. 7,521,541.
In certain aspects, the BCMA antibodies or bispecific BCMA antibodies provided herein may be further modified to contain additional non-proteinaceous moieties that are known in the art and readily available. The moieties suitable for derivatization of the antibody include but are not limited to water soluble polymers. Non-limiting examples of water soluble polymers include, but are not limited to, polyethylene glycol (PEG), copolymers of ethylene glycol/propylene glycol, carboxymethylcellulose, dextran, polyvinyl alcohol, polyvinyl pyrrolidone, poly-1, 3- dioxolane, poly-1, 3, 6-trioxane, ethylene/maleic anhydride copolymer, polyaminoacids (either homopolymers or random copolymers), and dextran or poly(n-vinyl pyrrolidone)polyethylene glycol, propropylene glycol homopolymers, prolypropylene oxide/ethylene oxide co-polymers, polyoxyethylated polyols (e.g., glycerol), polyvinyl alcohol, and mixtures thereof. Polyethylene glycol propionaldehyde may have advantages in manufacturing due to its stability in water. The polymer may be of any molecular weight, and may be branched or unbranched. The number of polymers attached to the antibody may vary, and if more than one polymer is attached, they can be the same or different molecules. In general, the number and/or type of polymers used for derivatization can be determined based on considerations including, but not limited to, the particular properties or functions of the antibody to be improved, whether the bispecific antibody derivative will be used in a therapy under defined conditions, etc. In another aspect, conjugates of an antibody and non-proteinaceous moiety that may be selectively heated by exposure to radiation are provided. In one embodiment, the non-proteinaceous moiety is a carbon nanotube (Kam, N.W. et al., Proc. Natl. Acad. Sci. USA 102 (2005) 11600-11605). The radiation may be of any wavelength, and includes, but is not limited to, wavelengths that do not harm ordinary cells, but which heat the non-proteinaceous moiety to a temperature at which cells proximal to the antibody-non-proteinaceous moiety are killed. In another aspect, immunoconjugates of the BCMA antibodies or bispecific BCMA antibodies provided herein may be obtained. An “immunoconjugate” is an antibody conjugated to one or more heterologous molecule(s), for instance a small molecule agent.
The term "polynucleotide" refers to an isolated nucleic acid molecule or construct, e.g. messenger RNA (mRNA), virally-derived RNA, or plasmid DNA (pDNA). A polynucleotide may comprise a conventional phosphodiester bond or a non-conventional bond (e.g. an amide
bond, such as found in peptide nucleic acids (PNA). The term "nucleic acid molecule" refers to any one or more nucleic acid segments, e.g. DNA or RNA fragments, present in a polynucleotide. Each nucleotide is composed of a base, specifically a purine- or pyrimidine base (i.e. cytosine (C), guanine (G), adenine (A), thymine (T) or uracil (U)), a sugar (i.e. deoxyribose or ribose), and a phosphate group. Often, the nucleic acid molecule is described by the sequence of bases, whereby said bases represent the primary structure (linear structure) of a nucleic acid molecule. The sequence of bases is typically represented from 5’ to 3’. Herein, the term nucleic acid molecule encompasses deoxyribonucleic acid (DNA) including e.g., complementary DNA (cDNA) and genomic DNA, ribonucleic acid (RNA), in particular messenger RNA (mRNA), synthetic forms of DNA or RNA, and mixed polymers comprising two or more of these molecules. The nucleic acid molecule may be linear or circular. In addition, the term nucleic acid molecule includes both, sense and antisense strands, as well as single stranded and double stranded forms. Moreover, the herein described nucleic acid molecule can contain naturally occurring or non-naturally occurring nucleotides. Examples of non-naturally occurring nucleotides include modified nucleotide bases with derivatized sugars or phosphate backbone linkages or chemically modified residues. Nucleic acid molecules also encompass DNA and RNA molecules which are suitable as a vector for direct expression of an antibody of the invention in vitro and/or in vivo, e.g., in a host or patient. Such DNA (e.g., cDNA) or RNA (e.g., mRNA) vectors, can be unmodified or modified. For example, mRNA can be chemically modified to enhance the stability of the RNA vector and/or expression of the encoded molecule so that mRNA can be injected into a subject to generate the antibody in vivo (see e.g., Stadler et al. (2017) Nature Medicine 23:815-817, or EP 2 101 823 Bl).
By "isolated" nucleic acid molecule or polynucleotide is meant a nucleic acid molecule, DNA or RNA, which has been removed from its native environment. For example, a recombinant polynucleotide encoding a polypeptide contained in a vector is considered isolated for the purposes of the present invention. Further examples of an isolated polynucleotide include recombinant polynucleotides maintained in heterologous host cells or purified (partially or substantially) polynucleotides in solution. An isolated polynucleotide includes a polynucleotide molecule contained in cells that ordinarily contain the polynucleotide molecule, but the polynucleotide molecule is present extrachromosomally or at a chromosomal location that is different from its natural chromosomal location. Isolated RNA molecules include in vivo or in vitro RNA transcripts of the present invention, as well as positive and negative strand forms, and double-stranded forms. Isolated polynucleotides or nucleic acids according to the present invention further include such molecules produced synthetically. In addition, a polynucleotide or a nucleic acid may be or may include a regulatory element such as a promoter, ribosome binding site, or a transcription terminator.
By a nucleic acid or polynucleotide having a nucleotide sequence at least, for example, 95% "identical" to a reference nucleotide sequence of the present invention, it is intended that the
nucleotide sequence of the polynucleotide is identical to the reference sequence except that the polynucleotide sequence may include up to five point mutations per each 100 nucleotides of the reference nucleotide sequence. In other words, to obtain a polynucleotide having a nucleotide sequence at least 95% identical to a reference nucleotide sequence, up to 5% of the nucleotides in the reference sequence may be deleted or substituted with another nucleotide, or a number of nucleotides up to 5% of the total nucleotides in the reference sequence may be inserted into the reference sequence. These alterations of the reference sequence may occur at the 5’ or 3’ terminal positions of the reference nucleotide sequence or anywhere between those terminal positions, interspersed either individually among residues in the reference sequence or in one or more contiguous groups within the reference sequence. As a practical matter, whether any particular polynucleotide sequence is at least 80%, 85%, 90%, 95%, 96%, 97%, 98% or 99% identical to a nucleotide sequence of the present invention can be determined conventionally using known computer programs, such as the ones discussed above for polypeptides (e.g. ALIGN-2).
The term "expression cassette" refers to a polynucleotide generated recombinantly or synthetically, with a series of specified nucleic acid elements that permit transcription of a particular nucleic acid in a target cell. The recombinant expression cassette can be incorporated into a plasmid, chromosome, mitochondrial DNA, plastid DNA, virus, or nucleic acid fragment. Typically, the recombinant expression cassette portion of an expression vector includes, among other sequences, a nucleic acid sequence to be transcribed and a promoter. In certain embodiments, the expression cassette of the invention comprises polynucleotide sequences that encode bispecific antigen binding molecules of the invention or fragments thereof.
The term “vector” or "expression vector" is synonymous with "expression construct" and refers to a DNA molecule that is used to introduce and direct the expression of a specific gene to which it is operably associated in a target cell. The term includes the vector as a self-replicating nucleic acid structure as well as the vector incorporated into the genome of a host cell into which it has been introduced. The expression vector of the present invention comprises an expression cassette. Expression vectors allow transcription of large amounts of stable mRNA. Once the expression vector is inside the target cell, the ribonucleic acid molecule or protein that is encoded by the gene is produced by the cellular transcription and/or translation machinery. In one embodiment, the expression vector of the invention comprises an expression cassette that comprises polynucleotide sequences that encode bispecific antigen binding molecules of the invention or fragments thereof.
The terms "host cell", "host cell line," and "host cell culture" are used interchangeably and refer to cells into which exogenous nucleic acid has been introduced, including the progeny of such cells. Host cells include "transformants" and "transformed cells," which include the primary transformed cell and progeny derived therefrom without regard to the number of passages. Progeny may not be completely identical in nucleic acid content to a parent cell, but may contain
mutations. Mutant progeny that have the same function or biological activity as screened or selected for in the originally transformed cell are included herein. A host cell is any type of cellular system that can be used to generate the bispecific antigen binding molecules of the present invention. Host cells include cultured cells, e.g. mammalian cultured cells, such as CHO cells, BHK cells, NSO cells, SP2/0 cells, YO myeloma cells, P3X63 mouse myeloma cells, PER cells, PER.C6 cells or hybridoma cells, yeast cells, insect cells, and plant cells, to name only a few, but also cells comprised within a transgenic animal, transgenic plant or cultured plant or animal tissue.
An "effective amount" of an agent refers to the amount that is necessary to result in a physiological change in the cell or tissue to which it is administered.
A "therapeutically effective amount" of an agent, e.g. a pharmaceutical composition, refers to an amount effective, at dosages and for periods of time necessary, to achieve the desired therapeutic or prophylactic result. A therapeutically effective amount of an agent for example eliminates, decreases, delays, minimizes or prevents adverse effects of a disease.
An “individual” or “subject” is a mammal. Mammals include, but are not limited to, domesticated animals (e.g. cows, sheep, cats, dogs, and horses), primates (e.g. humans and nonhuman primates such as monkeys), rabbits, and rodents (e.g. mice and rats). Particularly, the individual or subject is a human.
The term "pharmaceutical composition" refers to a preparation which is in such form as to permit the biological activity of an active ingredient contained therein to be effective, and which contains no additional components which are unacceptably toxic to a subject to which the formulation would be administered.
A “pharmaceutically acceptable excipient” refers to an ingredient in a pharmaceutical composition, other than an active ingredient, which is nontoxic to a subject. A pharmaceutically acceptable excipient includes, but is not limited to, a buffer, a stabilizer, or a preservative.
The term “package insert” is used to refer to instructions customarily included in commercial packages of therapeutic products, that contain information about the indications, usage, dosage, administration, combination therapy, contraindications and/or warnings concerning the use of such therapeutic products.
As used herein, “treatment” (and grammatical variations thereof such as “treat” or “treating”) refers to clinical intervention in an attempt to alter the natural course of the individual being treated, and can be performed either for prophylaxis or during the course of clinical pathology. Desirable effects of treatment include, but are not limited to, preventing occurrence or recurrence of disease, alleviation of symptoms, diminishment of any direct or indirect pathological consequences of the disease, preventing metastasis, decreasing the rate of disease progression, amelioration or palliation of the disease state, and remission or improved prognosis. In some embodiments, the molecules of the invention are used to delay development of a disease or to slow the progression of a disease.
The term “therapy” refers to any protocol, method and/or agent that can be used in the prevention, management, treatment and/or amelioration of a disease (or symptom related thereto) or cancer. In certain aspects, the terms “therapies” and “therapy” refer to a biological therapy, supportive therapy, and/or other therapies useful in the prevention, management, treatment and/or amelioration of a disease or cancer known to one of skill in the art such as a physician.
The term "combination treatment" or “co-administration” as noted herein encompasses combined administration (where two or more therapeutic agents are included in the same or separate formulations), and separate administration, in which case, administration of an antibody as reported herein can occur prior to, simultaneously, and/or following, administration of the additional therapeutic agent or agents, preferably an antibody or antibodies.
The term “cancer” refers to or describes the physiological condition in mammals that is typically characterized by unregulated cell growth/proliferation. Thus, the term cancer as used herein refers to proliferative diseases, such as carcinoma, lymphomas (e.g., Hodgkin’s and nonHodgkin’s lymphoma), blastoma, sarcoma, and leukemia. In particular, the term cancer includes lymphocytic leukemias, lung cancer, non-small cell lung (NSCL) cancer, bronchi oloalviolar cell lung cancer, bone cancer, pancreatic cancer, skin cancer, cancer of the head or neck, cutaneous or intraocular melanoma, uterine cancer, ovarian cancer, rectal cancer, cancer of the anal region, stomach cancer, gastric cancer, colon cancer, breast cancer, uterine cancer, carcinoma of the fallopian tubes, carcinoma of the endometrium, carcinoma of the cervix, carcinoma of the vagina, carcinoma of the vulva, Hodgkin's Disease, cancer of the esophagus, cancer of the small intestine, cancer of the endocrine system, cancer of the thyroid gland, cancer of the parathyroid gland, cancer of the adrenal gland, sarcoma of soft tissue, cancer of the urethra, cancer of the penis, prostate cancer, cancer of the bladder, cancer of the kidney or ureter, renal cell carcinoma, carcinoma of the renal pelvis, mesothelioma, hepatocellular cancer, biliary cancer, neoplasms of the central nervous system (CNS), spinal axis tumors, brain stem glioma, glioblastoma multiforme, astrocytomas, schwanomas, ependymonas, medulloblastomas, meningiomas, squamous cell carcinomas, pituitary adenoma and Ewings sarcoma, including refractory versions of any of the above cancers, or a combination of one or more of the above cancers. In one aspect, the cancer is a solid tumor. In another aspect, the cancer is a haematological cancer, particularly leukemia, most particularly acute lymphoblastic leukemia (ALL) or acute myelogenous leukemia (AML). In a preferred aspect, the term cancer refers to any canceer in which BCMA is expressed. More preferably, the cancer is multiple myeloma (MM).
Exemplary new BCMA antibodies
Provided herein are novel antibodies and/or antibody fragments that specifically bind to B cell maturation antigen (BCMA). Provided are novel antibodies and/or antibody fragments that specifically bind to the extracellular domain of human BCMA comprising the amino acid sequence of SEQ ID NO: 132. Thus, these antibodies specifically bind to human BCMA.
These antibodies are able to bind to human BCMA and to cynomolgus BCMA.
They bind to the human BCMA extracellular domain (ECD of the amino acid sequence of SEQ ID NO: 132) with an KD value of less than 5 nM as measured by surface plasmon resonance (SPR) (see Example 1.3).
As is demonstrated in Example 3, the new antibodies are also able to bind to mutated human BCMA variants hu BCMA R27P (SEQ ID NO:210), Hu BCMA_S30del (SEQ ID NO:211), hu BCMA P33S (SEQ ID NO:212) and hu BCMA_P34del (SEQ ID NO:213). They are therefore not affected by point mutations in the ECD of BCMA and do not lose the therapeutic activity as has been observed with other BCMA-targeted molecules such as Elranatamab and Teclistamab ((Lee et al., Nature Medicine 2023, 29, 2295-2306).
The new antibodies are further characterized in that they are producable in high amounts and with high titers, that they show high thermal stability (as measured by the aggregation temperature Tagg), or in that they possess a high degree of humanness and may therefore be less immunogenic in the human body. The percentage of humanness of the VH and VL sequences as compared to the human germline sequences can be determined by the methods described in Abhinandan, K. R. and Martin, Andrew C. R. 2007, J. Mol. Biol. 2007, 369, 852-862.
In one aspect, provided herein is an a antibody that specifically binds to B cell maturation agent (BCMA), wherein the antibody comprises
(i) a heavy chain variable region (VH BCMA) comprising a heavy chain complementary determining region CDR-H1 of SEQ ID NO: 1 (GYTFTNYWMH), a CDR-H2 of SEQ ID NO: 2 (IIHPNSGSTNYNEKFQG), and a CDR-H3 of SEQ ID NO: 3 (GIYDYPFAY), and
(ii) a light chain variable region (VLBCMA) selected from the group consisting of
(a) a VL comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 4 (RASES VSIHGTHLMH), a CDR-L2 of SEQ ID NO: 5 (AASSLQS) and a CDR-L3 of SEQ ID NO: 6 (QQSIEDPYT); or
(b) a VL comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 4 (RASES VSIHGTHLMH), a CDR-L2 of SEQ ID NO: 7 (AASNLES) and a CDR-L3 of SEQ ID NO: 6 (QQSIEDPYT), or
(c) a VL comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 4 (RASES VSIHGTHLMH), a CDR-L2 of SEQ ID NO: 8 (AASNLQS) and a CDR-L3 of SEQ ID NO: 6 (QQSIEDPYT).
In one aspect, provided is a BCMA antibody (or an antigen binding domain that specifically binds to BCMA), wherein said antibody comprises a VH BCMA comprising an amino acid sequence selected from the group consisting of SEQ ID NO: 9 (VHla) and SEQ ID NO: 10 (VHlb), and/or a VLBCMA comprising an amino acid sequence selected from the group consisting of SEQ ID NO: 11 (VLlf) , SEQ ID NO: 12 (VLla), SEQ ID NO: 13 (VLlb), SEQ ID NO: 14 (VLlc), SEQ ID NO: 15 (VLld), and SEQ ID NO: 16 (VLle).
In one aspect, provided is a BCMA antibody (or an antigen binding domain that specifically binds to BCMA), wherein said antibody (or antigen binding domain) comprises
(a) a VH BCMA comprising an amino acid sequence of SEQ ID NO:9 and VLBCMA comprising an amino acid sequence of SEQ ID NO: 11, or
(b) a VH BCMA comprising an amino acid sequence of SEQ ID NOV and VLBCMA comprising an amino acid sequence of SEQ ID NO: 12; or
(c) a VH BCMA comprising an amino acid sequence of SEQ ID NOV and VLBCMA comprising an amino acid sequence of SEQ ID NO: 13, or
(d) a VH BCMA comprising an amino acid sequence of SEQ ID NOV and VLBCMA comprising an amino acid sequence of SEQ ID NO: 14; or
(e) a VH BCMA comprising an amino acid sequence of SEQ ID NOV and VLBCMA comprising an amino acid sequence of SEQ ID NO: 15, or
(f) a VH BCMA comprising an amino acid sequence of SEQ ID NOV and VLBCMA comprising an amino acid sequence of SEQ ID NO: 16; or
(g) a VH BCMA comprising an amino acid sequence of SEQ ID NOV and VLBCMA comprising an amino acid sequence of SEQ ID NO: 135, or
(h) a VH BCMA comprising an amino acid sequence of SEQ ID NOV and VLBCMA comprising an amino acid sequence of SEQ ID NO: 136; or
(i) a VH BCMA comprising an amino acid sequence of SEQ ID NO: 10 and VLBCMA comprising an amino acid sequence of SEQ ID NO: 11, or
(j) a VH BCMA comprising an amino acid sequence of SEQ ID NO: 10 and VLBCMA comprising an amino acid sequence of SEQ ID NO: 12; or
(k) a VH BCMA comprising an amino acid sequence of SEQ ID NO: 10 and VLBCMA comprising an amino acid sequence of SEQ ID NO: 13, or
(l) a VH BCMA comprising an amino acid sequence of SEQ ID NO: 10 and VLBCMA comprising an amino acid sequence of SEQ ID NO: 14; or
(m) a VH BCMA comprising an amino acid sequence of SEQ ID NO: 10 and VLBCMA comprising an amino acid sequence of SEQ ID NO: 15, or
(n) a VH BCMA comprising an amino acid sequence of SEQ ID NO: 10 and VLBCMA comprising an amino acid sequence of SEQ ID NO: 16; or
(o) a VH BCMA comprising an amino acid sequence of SEQ ID NO: 10 and VLBCMA comprising an amino acid sequence of SEQ ID NO: 135, or
(p) a VH BCMA comprising an amino acid sequence of SEQ ID NO: 10 and VLBCMA comprising an amino acid sequence of SEQ ID NO: 136.
In one aspect, the BCMA antibody (or the antigen binding domain that specifically binds to BCMA) comprises
(a) a VH BCMA comprising an amino acid sequence of SEQ ID NOV and VLBCMA comprising an amino acid sequence of SEQ ID NO: 11, or
(b) a VH BCMA comprising an amino acid sequence of SEQ ID NO:9 and VLBCMA comprising an amino acid sequence of SEQ ID NO: 12; or
(c) a VH BCMA comprising an amino acid sequence of SEQ ID NOV and VLBCMA comprising an amino acid sequence of SEQ ID NO: 13, or
(d) a VH BCMA comprising an amino acid sequence of SEQ ID NOV and VLBCMA comprising an amino acid sequence of SEQ ID NO: 14; or
(e) a VH BCMA comprising an amino acid sequence of SEQ ID NOV and VLBCMA comprising an amino acid sequence of SEQ ID NO: 15, or
(f) a VH BCMA comprising an amino acid sequence of SEQ ID NOV and VLBCMA comprising an amino acid sequence of SEQ ID NO: 16; or
(g) a VH BCMA comprising an amino acid sequence of SEQ ID NOV and VLBCMA comprising an amino acid sequence of SEQ ID NO: 135, or
(h) a VH BCMA comprising an amino acid sequence of SEQ ID NOV and VLBCMA comprising an amino acid sequence of SEQ ID NO: 136.
In one aspect, the BCMA antibody (or the antigen binding domain that specifically binds to BCMA) comprises
(i) a VH BCMA comprising an amino acid sequence of SEQ ID NO: 10 and VLBCMA comprising an amino acid sequence of SEQ ID NO: 11, or
(j) a VH BCMA comprising an amino acid sequence of SEQ ID NO: 10 and VLBCMA comprising an amino acid sequence of SEQ ID NO: 12; or
(k) a VH BCMA comprising an amino acid sequence of SEQ ID NO: 10 and VLBCMA comprising an amino acid sequence of SEQ ID NO: 13, or
(l) a VH BCMA comprising an amino acid sequence of SEQ ID NO: 10 and VLBCMA comprising an amino acid sequence of SEQ ID NO: 14; or
(m) a VH BCMA comprising an amino acid sequence of SEQ ID NO: 10 and VLBCMA comprising an amino acid sequence of SEQ ID NO: 15, or
(n) a VH BCMA comprising an amino acid sequence of SEQ ID NO: 10 and VLBCMA comprising an amino acid sequence of SEQ ID NO: 16; or
(o) a VH BCMA comprising an amino acid sequence of SEQ ID NO: 10 and VLBCMA comprising an amino acid sequence of SEQ ID NO: 135, or
(p) a VH BCMA comprising an amino acid sequence of SEQ ID NO: 10 and VLBCMA comprising an amino acid sequence of SEQ ID NO: 136.
In one particular aspect, provided is a BCMA antibody (or an antigen binding domain that specifically binds to BCMA), wherein said antibody (or antigen binding domain) comprises
(a) a VH BCMA comprising an amino acid sequence of SEQ ID NOV and a VLBCMA comprising an amino acid sequence of SEQ ID NO: 11, or
(b) a VH BCMA comprising an amino acid sequence of SEQ ID NOV and a VLBCMA comprising an amino acid sequence of SEQ ID NO: 12.
More particularly, the BCMA antibody (or the antigen binding domain that specifically binds to BCMA) comprises a VH BCMA comprising an amino acid sequence of SEQ ID NO:9 and a VL BCMA comprising an amino acid sequence of SEQ ID NO: 11.
In another aspect, provided herein is an a antibody that specifically binds to B cell maturation agent (BCMA), wherein the antibody comprises
(i) a heavy chain variable region (VH BCMA) selected from the group consisting of
(a) a VH comprising a heavy chain complementary determining region CDR-H1 of SEQ ID NO: 29 (GFTFSNAWMD), a CDR-H2 of SEQ ID NO: 30 (QITAKSNNYATYYADSVKG), and a CDR-H3 of SEQ ID NO: 31 (DGYH), and
(b) a VH comprising a heavy chain complementary determining region CDR-H1 of SEQ ID NO: 29 (GFTFSNAWMD), a CDR-H2 of SEQ ID NO: 32 (QITAKSNNYATYYAAPVKG), and a CDR-H3 of SEQ ID NO: 31 (DGYH), and
(ii) a light chain variable region (VLBCMA) comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 33 (RASEDIRNGLA), a CDR-L2 of SEQ ID NO: 34 (NANSLHT) and a CDR-L3 of SEQ ID NO: 35 (EDTSKYPYT).
In one aspect, provided is a BCMA antibody (or an antigen binding domain that specifically binds to BCMA), wherein said antibody (or antigen binding domain) comprises a heavy chain variable region (VHBCMA) comprising a CDR-H1 of SEQ ID NO: 29 (GFTFSNAWMD), a CDR-H2 of SEQ ID NO: 30 (QITAKSNNYATYYADSVKG), and a CDR-H3 of SEQ ID NO: 31 (DGYH), and a light chain variable region (VLBCMA) comprising a CDR-L1 of SEQ ID NO: 33 (RASEDIRNGLA), a CDR-L2 of SEQ ID NO: 34 (NANSLHT) and a CDR-L3 of SEQ ID NO: 35 (EDTSKYPYT).
In one aspect, provided is a BCMA antibody (or an antigen binding domain that specifically binds to BCMA), wherein said antibody (or antigen binding domain) comprises a heavy chain variable region (VHBCMA) comprising a CDR-H1 of SEQ ID NO: 29 (GFTFSNAWMD), a CDR-H2 of SEQ ID NO: 32 (QITAKSNNYATYYAAPVKG), and a CDR-H3 of SEQ ID NO: 31 (DGYH), and a light chain variable region (VLBCMA) comprising a CDR-L1 of SEQ ID NO: 33 (RASEDIRNGLA), a CDR-L2 of SEQ ID NO: 34 (NANSLHT) and a CDR-L3 of SEQ ID NO: 35 (EDTSKYPYT).
In one aspect, provided is a BCMA antibody (or an antigen binding domain that specifically binds to BCMA), wherein said antibody (or antigen binding domain) comprises a VH BCMA comprising an amino acid sequence selected from the group consisting of SEQ ID NO: 36 (VH2a), SEQ ID NO: 38 (VHlb), SEQ ID NO:48 (VHla), SEQ ID NO:49 (VHla_Y292D), SEQ ID NO:50 (VHlc hu CDR2) and SEQ ID NO:51 (VHla_W197Y), and/or the VL BCMA comprising an amino acid sequence selected from the group consisting of SEQ ID NO: 37 (VL2a), SEQ ID NO:39 (VLla), SEQ ID NO:52 (VLla_L2_GL), SEQ ID NO:53 (VL2a_L2_GL), SEQ ID NO:54 (VLla_Ll_pGL), SEQ ID NO:55 (VLla_N651A), SEQ ID
NO:56 (VLla_N651A_N695S), SEQ ID NO:57 (VLla_H698Q), SEQ ID NO:58 (VLla_T699S) and SEQ ID NO:59 (VLla_H698Q_T699S).
In one aspect, provided is a BCMA antibody (or an antigen binding domain that specifically binds to BCMA), wherein said antibody (or antigen binding domain) comprises a VH BCMA comprising an amino acid sequence selected from the group consisting of SEQ ID NO: 36 (VH2a) and SEQ ID NO: 38 (VHlb), and/or the VL BCMA comprising an amino acid sequence selected from the group consisting of SEQ ID NO: 37 (VL2a) and SEQ ID NO:39 (VLla).
In one aspect, provided is a BCMA antibody (or an antigen binding domain that specifically binds to BCMA), wherein said antibody (or antigen binding domain) comprises
(a) a VH BCMA comprising an amino acid sequence of SEQ ID NO:36 and a VLBCMA comprising an amino acid sequence of SEQ ID NO:37, or
(b) a VH BCMA comprising an amino acid sequence of SEQ ID NO:38 and a VLBCMA comprising an amino acid sequence of SEQ ID NO:39; or
(c) a VH BCMA comprising an amino acid sequence of SEQ ID NO:48 and VLBCMA comprising an amino acid sequence of SEQ ID NO:52, or
(d) a VH BCMA comprising an amino acid sequence of SEQ ID NO:49 and VLBCMA comprising an amino acid sequence of SEQ ID NO:39; or
(e) a VH BCMA comprising an amino acid sequence of SEQ ID NO:49 and VLBCMA comprising an amino acid sequence of SEQ ID NO:52, or
(f) a VHBCMA comprising an amino acid sequence of SEQ ID NO:49 and VLBCMA comprising an amino acid sequence of SEQ ID NO:58; or
(g) a VH BCMA comprising an amino acid sequence of SEQ ID NO:49 and VLBCMA comprising an amino acid sequence of SEQ ID NO:37, or
(h) a VH BCMA comprising an amino acid sequence of SEQ ID NO:38 and VLBCMA comprising an amino acid sequence of SEQ ID NO:54; or
(i) a VH BCMA comprising an amino acid sequence of SEQ ID NO:38 and VLBCMA comprising an amino acid sequence of SEQ ID NO:55, or
(j) a VH BCMA comprising an amino acid sequence of SEQ ID NO:38 and VLBCMA comprising an amino acid sequence of SEQ ID NO:56; or
(k) a VH BCMA comprising an amino acid sequence of SEQ ID NO:38 and VLBCMA comprising an amino acid sequence of SEQ ID NO:58, or
(l) a VH BCMA comprising an amino acid sequence of SEQ ID NO:38 and VLBCMA comprising an amino acid sequence of SEQ ID NO:37; or
(m) a VH BCMA comprising an amino acid sequence of SEQ ID NO:36 and VLBCMA comprising an amino acid sequence of SEQ ID NO:39, or
(n) a VH BCMA comprising an amino acid sequence of SEQ ID NO:36 and VLBCMA comprising an amino acid sequence of SEQ ID NO:54; or
(o) a VH BCMA comprising an amino acid sequence of SEQ ID NO:36 and VLBCMA comprising an amino acid sequence of SEQ ID NO:58.
In one particular aspect, provided is a BCMA antibody (or an antigen binding domain that specifically binds to BCMA), wherein said antibody (or antigen binding domain) comprises
(a) a VH BCMA comprising an amino acid sequence of SEQ ID NO:36 and a VLBCMA comprising an amino acid sequence of SEQ ID NO:37, or
(b) a VH BCMA comprising an amino acid sequence of SEQ ID NO:38 and a VLBCMA comprising an amino acid sequence of SEQ ID NO:39.
More particularly, the BCMA antibody comprises a VH BCMA comprising an amino acid sequence of SEQ ID NO:36 and a VLBCMA comprising an amino acid sequence of SEQ ID NO:37.
In one aspect, the antibody that specifically binds to BCMA is a full-length antibody, in particular of human IgGl subclass. In one particular aspect, it comprises the amino acid mutations L234A, L235A and P329G (numbering according to Kabat EU index).
Bispecific CD28 agonistic antibodies comprising the new BCMA antibodies
The patent application also provides new BCMA antibodies that comprise a second antigen binding domain that specifically binds to second antigen, in particular to CD28. These bispecific CD28 agonistic antibodies possess advantageous properties such as excellent producibility, stability, binding affinity, biological activity, targeting efficiency, reduced toxicity, an extended dosage range that can be given to a patient and thereby a possibly an enhanced efficacy can be observed. The new bispecific CD28 agonistic antibodies comprise an Fc domain composed of a first and a second subunit capable of stable association comprising one or more amino acid substitution that reduces the binding affinity of the antigen binding molecule to an Fc receptor and/or effector function (Fc silent) and thus unspecific cross-linking via Fc receptors is avoided. Instead, they comprise specific antigen binding domain capable of specific binding to BCMA which causes cross-linking at the tumor site. Surprisingly, the inventors have found that based on their binding properties the BCMA antigen binding domains as described herein have advantageous properties that makes them more usable in the bispecific format. Furthermore, it has been found that the bispecific agonistic CD28 antigen binding molecule comprising these BCMA antigen binding domains possess an improved functionality and ability to increase T cell activation, particularly in the presence of T-cell activating anti-CD3 bispecific antibodies. Thus, an enhanced tumor-specific T cell activation is achieved.
In one aspect, provided herein is a bispecific antibody that specifically binds to B cell maturation agent (BCMA) and CD28, wherein the antibody comprises
(A) a first antigen binding domain comprising
(i) a heavy chain variable region (VH BCMA) comprising a heavy chain complementary determining region CDR-H1 of SEQ ID NO: 1 (GYTFTNYWMH), a CDR-H2 of SEQ ID NO: 2 (IH4PNSGSTNYNEKFQG), and a CDR-H3 of SEQ ID NO: 3 (GIYDYPFAY), and
(ii) a light chain variable region (VLBCMA) selected from the group consisting of
(a) a VL comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 4 (RASES VSIHGTHLMH), a CDR-L2 of SEQ ID NO: 5 (AASSLQS) and a CDR-L3 of SEQ ID NO: 6 (QQSIEDPYT); or
(b) a VL comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 4 (RASES VSIHGTHLMH), a CDR-L2 of SEQ ID NO: 7 (AASNLES) and a CDR-L3 of SEQ ID NO: 6 (QQSIEDPYT), or
(c) a VL comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 4 (RASES VSIHGTHLMH), a CDR-L2 of SEQ ID NO: 8 (AASNLQS) and a CDR-L3 of SEQ ID NO: 6 (QQSIEDPYT), and
(B) a second antigen binding domain that specifically binds to CD28.
In one aspect, provided is an antibody that specifically binds to BCMA and CD28, wherein said antibody comprises a VH BCMA comprising an amino acid sequence selected from the group consisting of SEQ ID NO: 9 (VHla) and SEQ ID NO: 10 (VHlb), and/or a VLBCMA comprising an amino acid sequence selected from the group consisting of SEQ ID NO: 11 (VLlf) , SEQ ID NO: 12 (VLla), SEQ ID NO: 13 (VLlb), SEQ ID NO: 14 (VLlc), SEQ ID NO: 15 (VLld), and SEQ ID NO: 16 (VLle).
In one aspect, provided is an antibody that specifically binds to BCMA and CD28, wherein said antibody comprises
(a) a VH BCMA comprising an amino acid sequence of SEQ ID NOV and VLBCMA comprising an amino acid sequence of SEQ ID NO: 11, or
(b) a VH BCMA comprising an amino acid sequence of SEQ ID NOV and VLBCMA comprising an amino acid sequence of SEQ ID NO: 12; or
(c) a VH BCMA comprising an amino acid sequence of SEQ ID NOV and VLBCMA comprising an amino acid sequence of SEQ ID NO: 13, or
(d) a VH BCMA comprising an amino acid sequence of SEQ ID NOV and VLBCMA comprising an amino acid sequence of SEQ ID NO: 14; or
(e) a VH BCMA comprising an amino acid sequence of SEQ ID NOV and VLBCMA comprising an amino acid sequence of SEQ ID NO: 15, or
(f) a VHBCMA comprising an amino acid sequence of SEQ ID NOV and VLBCMA comprising an amino acid sequence of SEQ ID NO: 16; or
(g) a VH BCMA comprising an amino acid sequence of SEQ ID NOV and VLBCMA comprising an amino acid sequence of SEQ ID NO: 135, or
(h) a VH BCMA comprising an amino acid sequence of SEQ ID NOV and VLBCMA comprising an amino acid sequence of SEQ ID NO: 136; or
(i) a VH BCMA comprising an amino acid sequence of SEQ ID NO: 10 and VLBCMA comprising an amino acid sequence of SEQ ID NO: 11, or
(j) a VH BCMA comprising an amino acid sequence of SEQ ID NO: 10 and VLBCMA comprising an amino acid sequence of SEQ ID NO: 12; or
(k) a VH BCMA comprising an amino acid sequence of SEQ ID NO: 10 and VLBCMA comprising an amino acid sequence of SEQ ID NO: 13, or
(l) a VH BCMA comprising an amino acid sequence of SEQ ID NO: 10 and VLBCMA comprising an amino acid sequence of SEQ ID NO: 14; or
(m) a VH BCMA comprising an amino acid sequence of SEQ ID NO: 10 and VLBCMA comprising an amino acid sequence of SEQ ID NO: 15, or
(n) a VH BCMA comprising an amino acid sequence of SEQ ID NO: 10 and VLBCMA comprising an amino acid sequence of SEQ ID NO: 16; or
(o) a VH BCMA comprising an amino acid sequence of SEQ ID NO: 10 and VLBCMA comprising an amino acid sequence of SEQ ID NO: 135, or
(p) a VH BCMA comprising an amino acid sequence of SEQ ID NO: 10 and VLBCMA comprising an amino acid sequence of SEQ ID NO: 136.
In one aspect, the antibody that specifically binds to BCMA and CD28 comprises
(a) a VH BCMA comprising an amino acid sequence of SEQ ID NOV and VLBCMA comprising an amino acid sequence of SEQ ID NO: 11, or
(b) a VH BCMA comprising an amino acid sequence of SEQ ID NOV and VLBCMA comprising an amino acid sequence of SEQ ID NO: 12; or
(c) a VH BCMA comprising an amino acid sequence of SEQ ID NOV and VLBCMA comprising an amino acid sequence of SEQ ID NO: 13, or
(d) a VH BCMA comprising an amino acid sequence of SEQ ID NOV and VLBCMA comprising an amino acid sequence of SEQ ID NO: 14; or
(e) a VH BCMA comprising an amino acid sequence of SEQ ID NOV and VLBCMA comprising an amino acid sequence of SEQ ID NO: 15, or
(f) a VH BCMA comprising an amino acid sequence of SEQ ID NOV and VLBCMA comprising an amino acid sequence of SEQ ID NO: 16; or
(g) a VH BCMA comprising an amino acid sequence of SEQ ID NOV and VLBCMA comprising an amino acid sequence of SEQ ID NO: 135, or
(h) a VH BCMA comprising an amino acid sequence of SEQ ID NOV and VLBCMA comprising an amino acid sequence of SEQ ID NO: 136.
In one aspect, the antibody that specifically binds to BCMA and CD28 comprises
(i) a VH BCMA comprising an amino acid sequence of SEQ ID NO: 10 and VLBCMA comprising an amino acid sequence of SEQ ID NO: 11, or
(j) a VH BCMA comprising an amino acid sequence of SEQ ID NO: 10 and VLBCMA comprising an amino acid sequence of SEQ ID NO: 12; or
(k) a VH BCMA comprising an amino acid sequence of SEQ ID NO: 10 and VLBCMA comprising an amino acid sequence of SEQ ID NO: 13, or
(l) a VH BCMA comprising an amino acid sequence of SEQ ID NO: 10 and VLBCMA comprising an amino acid sequence of SEQ ID NO: 14; or
(m) a VH BCMA comprising an amino acid sequence of SEQ ID NO: 10 and VLBCMA comprising an amino acid sequence of SEQ ID NO: 15, or
(n) a VH BCMA comprising an amino acid sequence of SEQ ID NO: 10 and VLBCMA comprising an amino acid sequence of SEQ ID NO: 16; or
(o) a VH BCMA comprising an amino acid sequence of SEQ ID NO: 10 and VLBCMA comprising an amino acid sequence of SEQ ID NO: 135, or
(p) a VH BCMA comprising an amino acid sequence of SEQ ID NO: 10 and VLBCMA comprising an amino acid sequence of SEQ ID NO: 136.
In one particular aspect, provided is an antibody that specifically binds to BCMA and CD28, wherein said antibody comprises
(a) a VH BCMA comprising an amino acid sequence of SEQ ID NOV and a VLBCMA comprising an amino acid sequence of SEQ ID NO: 11, or
(b) a VH BCMA comprising an amino acid sequence of SEQ ID NOV and a VLBCMA comprising an amino acid sequence of SEQ ID NO: 12.
More particularly, the antibody that specifically binds to BCMA and CD28 comprises a VH BCMA comprising an amino acid sequence of SEQ ID NOV and a VL BCMA comprising an amino acid sequence of SEQ ID NO: 11.
In all these aspects as mentioned herein before, the first antigen binding domain that specifically binds to BCMA can be a Fab molecule. In some aspects, the first antigen binding domain that specifically binds to BCMA can be a cross-Fab molecule, i.e a Fab molecule wherein the variable domains VL and VH or the constant domains CL and CHI, particularly the variable domains VL and VH, of the Fab light chain and the Fab heavy chain are replaced by each other.
In a particular aspect, the first antigen binding domain that specifically binds to BCMA is a Fab molecule wherein in the constant domain CL the amino acid at position 123 (numbering according to Kabat EU index) is substituted by an amino acid selected from lysine (K), arginine (R) or histidine (H) and the amino acid at position 124 (numbering according to Kabat EU index) is substituted independently by lysine (K), arginine (R) or histidine (H), and wherein in the constant domain CHI the amino acid at position 147 (numbering according to Kabat EU index) is substituted independently by glutamic acid (E) or aspartic acid (D) and the amino acid at position 213 (numbering according to Kabat EU index) is substituted independently by glutamic acid (E), or aspartic acid (D) (numbering according to Kabat EU index).
In any of the aspects as mentioned herein before, the second antigen binding domain that specifically binds to CD28 can be a Fab molecule. In a particular aspect, the second antigen binding domain that specifically binds to CD28 is a cross-Fab molecule, i.e a Fab molecule wherein the variable domains VL and VH or the constant domains CL and CHI, particularly the variable domains VL and VH, of the Fab light chain and the Fab heavy chain are replaced by each other.
In one aspect, the second antigen binding domain that specifically binds to CD28 comprises a heavy chain variable region (VHCD28) comprising a heavy chain complementary determining region CDR-H1 of SEQ ID NO: 17, a CDR-H2 of SEQ ID NO: 18, and a CDR-H3 of SEQ ID NO: 19, and a light chain variable region (VLCD28) comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 20, a CDR-L2 of SEQ ID NO: 21 and a CDR-L3 of SEQ ID NO: 22. In one aspect, the second antigen binding domain that specifically binds to CD28 comprises a heavy chain variable region (VHCD28) comprising an amino acid sequence of SEQ ID NO:23, and a light chain variable region (VLCD28) comprising an amino acid sequence of SEQ ID NO:24 (v8).
In one particular aspect, provided is an antibody that specifically binds to BCMA and CD28, wherein said antibody comprises a first antigen binding domain comprising a VH BCMA comprising an amino acid sequence of SEQ ID NOV and VLBCMA comprising an amino acid sequence of SEQ ID NO: 11 and a second antigen binding domain comprising VH CD28 comprising an amino acid sequence of SEQ ID NO:23 and VL CD28 comprising an amino acid sequence of SEQ ID NO:24.
In one aspect, the antibody that specifically binds to BCMA and CD28 comprises a polypeptide that is at least 95%, 96%, 97%, 98%, or 99% identical to the sequence of SEQ ID NO: 25, a polypeptide that is at least 95%, 96%, 97%, 98%, or 99% identical to the sequence of SEQ ID NO: 26, a polypeptide that is at least 95%, 96%, 97%, 98%, or 99% identical to the sequence of SEQ ID NO: 27, and a polypeptide that is at least 95%, 96%, 97%, 98%, or 99% identical to the sequence of SEQ ID NO: 28. In one particular aspect, provided is an antibody that specifically binds to BCMA and CD28, wherein said antibody comprises a first light chain comprising the amino acid sequence of SEQ ID NO:25, a first heavy chain comprising the amino acid sequence of SEQ ID NO:26, a second heavy chain comprising the amino acid sequence of SEQ ID NO:27 and a second light chain comprising the amino acid sequence of SEQ ID NO:28.
In another aspect, provided is an antibody that specifically binds to BCMA and CD28, wherein said antibody comprises a first antigen binding domain comprising a VH BCMA comprising an amino acid sequence of SEQ ID NOV and VLBCMA comprising an amino acid sequence of SEQ ID NO: 12 and a second antigen binding domain comprising VH CD28 comprising an amino acid sequence of SEQ ID NO:23 and VL CD28 comprising an amino acid sequence of SEQ ID NO:24.
In one aspect, the antibody that specifically binds to BCMA and CD28 comprises a polypeptide that is at least 95%, 96%, 97%, 98%, or 99% identical to the sequence of SEQ ID NO: 92, a polypeptide that is at least 95%, 96%, 97%, 98%, or 99% identical to the sequence of SEQ ID NO: 26, a polypeptide that is at least 95%, 96%, 97%, 98%, or 99% identical to the sequence of SEQ ID NO: 27, and a polypeptide that is at least 95%, 96%, 97%, 98%, or 99% identical to the sequence of SEQ ID NO: 28. In one aspect, provided is an antibody that specifically binds to BCMA and CD28, wherein said antibody comprises a first light chain comprising the amino acid sequence of SEQ ID NO:92, a first heavy chain comprising the amino acid sequence of SEQ ID NO:26, a second heavy chain comprising the amino acid sequence of SEQ ID NO:27 and a second light chain comprising the amino acid sequence of SEQ ID NO:28.
In a further aspect, provided is an antibody that specifically binds to B cell maturation agent (BCMA) and CD28, wherein the antibody comprises
(A) a first antigen binding domain comprising
(i) a heavy chain variable region (VH BCMA) selected from the group consisting of
(a) a VH comprising a heavy chain complementary determining region CDR-H1 of SEQ ID NO: 29 (GFTFSNAWMD), a CDR-H2 of SEQ ID NO: 30 (QITAKSNNYATYYADSVKG), and a CDR-H3 of SEQ ID NO: 31 (DGYH), and
(b) a VH comprising a heavy chain complementary determining region CDR-H1 of SEQ ID NO: 29 (GFTFSNAWMD), a CDR-H2 of SEQ ID NO: 32 (QITAKSNNYATYYAAPVKG), and a CDR-H3 of SEQ ID NO: 31 (DGYH), and
(ii) a light chain variable region (VLBCMA) comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 33 (RASEDIRNGLA), a CDR-L2 of SEQ ID NO: 34 (NANSLHT) and a CDR-L3 of SEQ ID NO: 35 (EDTSKYPYT), and
(B) a second antigen binding domain that specifically binds to CD28.
In one aspect, provided is an antibody that specifically binds to BCMA and CD28, wherein said antibody comprises a heavy chain variable region (VH BCMA) comprising a CDR-H1 of SEQ ID NO: 29 (GFTFSNAWMD), a CDR-H2 of SEQ ID NO: 30 (QITAKSNNYATYYADSVKG), and a CDR-H3 of SEQ ID NO: 31 (DGYH), and a light chain variable region (VL BCMA) comprising a CDR-L1 of SEQ ID NO: 33 (RASEDIRNGLA), a CDR-L2 of SEQ ID NO: 34 (NANSLHT) and a CDR-L3 of SEQ ID NO: 35 (EDTSKYPYT).
In one aspect, provided is an antibody that specifically binds to BCMA and CD28, wherein said antibody comprises a heavy chain variable region (VH BCMA) comprising a CDR-H1 of SEQ ID NO: 29 (GFTFSNAWMD), a CDR-H2 of SEQ ID NO: 32 (QITAKSNNYATYYAAPVKG), and a CDR-H3 of SEQ ID NO: 31 (DGYH), and a light chain variable region (VL BCMA) comprising a CDR-L1 of SEQ ID NO: 33 (RASEDIRNGLA), a CDR-L2 of SEQ ID NO: 34 (NANSLHT) and a CDR-L3 of SEQ ID NO: 35 (EDTSKYPYT).
In one aspect, provided is an antibody that specifically binds to BCMA and CD28, wherein said antibody comprises a VH BCMA comprising an amino acid sequence selected from the group consisting of SEQ ID NO: 36 (VH2a), SEQ ID NO: 38 (VHlb), SEQ ID NO:48 (VHla), SEQ ID NO:49 (VHla_Y292D), SEQ ID NO:50 (VHlc hu CDR2) and SEQ ID NO:51 (VHla_W197Y), and/or the VL BCMA comprising an amino acid sequence selected from the group consisting of SEQ ID NO: 37 (VL2a), SEQ ID NO:39 (VLla), SEQ ID NO:52 (VLla_L2_GL), SEQ ID NO:53 (VL2a_L2_GL), SEQ ID NO:54 (VLla_Ll_pGL), SEQ ID NO 55 (VLla_N651A), SEQ ID NO:56 (VLla_N651A_N695S), SEQ ID NO:57 (VLla_H698Q), SEQ ID NO:58 (VLla_T699S) and SEQ ID NO:59 (VLla_H698Q_T699S).
In one aspect, provided is an antibody that specifically binds to BCMA and CD28, wherein said antibody comprises a VH BCMA comprising an amino acid sequence selected from the group consisting of SEQ ID NO: 36 (VH2a) and SEQ ID NO: 38 (VHlb), and/or the VL BCMA comprising an amino acid sequence selected from the group consisting of SEQ ID NO: 37 (VL2a) and SEQ ID NO:39 (VLla).
In one aspect, provided is an antibody that specifically binds to BCMA and CD28, wherein said antibody comprises
(a) a VH BCMA comprising an amino acid sequence of SEQ ID NO:36 and a VL BCMA comprising an amino acid sequence of SEQ ID NO:37, or
(b) a VH BCMA comprising an amino acid sequence of SEQ ID NO:38 and a VL BCMA comprising an amino acid sequence of SEQ ID NO:39; or
(c) a VH BCMA comprising an amino acid sequence of SEQ ID NO:48 and VLBCMA comprising an amino acid sequence of SEQ ID NO:52, or
(d) a VH BCMA comprising an amino acid sequence of SEQ ID NO:49 and VLBCMA comprising an amino acid sequence of SEQ ID NO:39; or
(e) a VH BCMA comprising an amino acid sequence of SEQ ID NO:49 and VLBCMA comprising an amino acid sequence of SEQ ID NO:52, or
(f) a VH BCMA comprising an amino acid sequence of SEQ ID NO:49 and VLBCMA comprising an amino acid sequence of SEQ ID NO:58; or
(g) a VH BCMA comprising an amino acid sequence of SEQ ID NO:49 and VLBCMA comprising an amino acid sequence of SEQ ID NO:37, or
(h) a VH BCMA comprising an amino acid sequence of SEQ ID NO:38 and VLBCMA comprising an amino acid sequence of SEQ ID NO:54; or
(i) a VH BCMA comprising an amino acid sequence of SEQ ID NO:38 and VLBCMA comprising an amino acid sequence of SEQ ID NO:55, or
(j) a VH BCMA comprising an amino acid sequence of SEQ ID NO:38 and VLBCMA comprising an amino acid sequence of SEQ ID NO:56; or
(k) a VH BCMA comprising an amino acid sequence of SEQ ID NO:38 and VLBCMA comprising an amino acid sequence of SEQ ID NO:58, or
(l) a VH BCMA comprising an amino acid sequence of SEQ ID NO:38 and VLBCMA comprising an amino acid sequence of SEQ ID NO:37; or
(m) a VH BCMA comprising an amino acid sequence of SEQ ID NO:36 and VLBCMA comprising an amino acid sequence of SEQ ID NO:39, or
(n) a VH BCMA comprising an amino acid sequence of SEQ ID NO:36 and VLBCMA comprising an amino acid sequence of SEQ ID NO:54; or
(o) a VH BCMA comprising an amino acid sequence of SEQ ID NO:36 and VLBCMA comprising an amino acid sequence of SEQ ID NO:58.
In one particular aspect, provided is an antibody that specifically binds to BCMA and CD28, wherein said antibody comprises
(a) a VH BCMA comprising an amino acid sequence of SEQ ID NO:36 and a VLBCMA comprising an amino acid sequence of SEQ ID NO:37, or
(b) a VH BCMA comprising an amino acid sequence of SEQ ID NO:38 and a VLBCMA comprising an amino acid sequence of SEQ ID NO:39.
More particularly, the antibody that specifically binds to BCMA and CD28 comprises a VH BCMA comprising an amino acid sequence of SEQ ID NO:36 and a VLBCMA comprising an amino acid sequence of SEQ ID NO:37.
In all these aspects as mentioned herein before, the first antigen binding domain that specifically binds to BCMA can be a Fab molecule. In some aspects, the first antigen binding domain that specifically binds to BCMA can be a cross-Fab molecule, i.e a Fab molecule wherein the variable domains VL and VH or the constant domains CL and CHI, particularly the variable domains VL and VH, of the Fab light chain and the Fab heavy chain are replaced by each other.
In a particular aspect, the first antigen binding domain that specifically binds to BCMA is a Fab molecule wherein in the constant domain CL the amino acid at position 123 (numbering according to Kabat EU index) is substituted by an amino acid selected from lysine (K), arginine (R) or histidine (H) and the amino acid at position 124 (numbering according to Kabat EU index) is substituted independently by lysine (K), arginine (R) or histidine (H), and wherein in the constant domain CHI the amino acid at position 147 (numbering according to Kabat EU index) is substituted independently by glutamic acid (E) or aspartic acid (D) and the amino acid at position 213 (numbering according to Kabat EU index) is substituted independently by glutamic acid (E), or aspartic acid (D) (numbering according to Kabat EU index).
In any of the aspects as mentioned herein before, the second antigen binding domain that specifically binds to CD28 can be a Fab molecule. In a particular aspect, the second antigen binding domain that specifically binds to CD28 is a cross-Fab molecule, i.e a Fab molecule wherein the variable domains VL and VH or the constant domains CL and CHI, particularly the
variable domains VL and VH, of the Fab light chain and the Fab heavy chain are replaced by each other.
In one aspect, the second antigen binding domain that specifically binds to CD28 comprises a heavy chain variable region (VHCD28) comprising a heavy chain complementary determining region CDR-H1 of SEQ ID NO: 17, a CDR-H2 of SEQ ID NO: 18, and a CDR-H3 of SEQ ID NO: 19, and a light chain variable region (VLCD28) comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 20, a CDR-L2 of SEQ ID NO: 21 and a CDR-L3 of SEQ ID NO: 22. In one aspect, the second antigen binding domain that specifically binds to CD28 comprises a heavy chain variable region (VHCD28) comprising an amino acid sequence of SEQ ID NO:23, and a light chain variable region (VLCD28) comprising an amino acid sequence of SEQ ID NO:24 (v8).
In one particular aspect, provided is an antibody that specifically binds to BCMA and CD28, wherein said antibody comprises a first antigen binding domain comprising a VH BCMA comprising an amino acid sequence of SEQ ID NO:36 and VLBCMA comprising an amino acid sequence of SEQ ID NO:37 and a second antigen binding domain comprising VH CD28 comprising an amino acid sequence of SEQ ID NO:23 and VL CD28 comprising an amino acid sequence of SEQ ID NO:24.
In one aspect, the antibody that specifically binds to BCMA and CD28 comprises a polypeptide that is at least 95%, 96%, 97%, 98%, or 99% identical to the sequence of SEQ ID NO: 40, a polypeptide that is at least 95%, 96%, 97%, 98%, or 99% identical to the sequence of SEQ ID NO: 41, a polypeptide that is at least 95%, 96%, 97%, 98%, or 99% identical to the sequence of SEQ ID NO: 27, and a polypeptide that is at least 95%, 96%, 97%, 98%, or 99% identical to the sequence of SEQ ID NO: 28. In one particular aspect, provided is an antibody that specifically binds to BCMA and CD28, wherein said antibody comprises a first light chain comprising the amino acid sequence of SEQ ID NO:40, a first heavy chain comprising the amino acid sequence of SEQ ID NO:41, a second heavy chain comprising the amino acid sequence of SEQ ID NO:27 and a second light chain comprising the amino acid sequence of SEQ ID NO:28.
In another aspect, provided is an antibody that specifically binds to BCMA and CD28, wherein said antibody comprises a first antigen binding domain comprising a VH BCMA comprising an amino acid sequence of SEQ ID NO:38 and VLBCMA comprising an amino acid sequence of SEQ ID NO:39 and a second antigen binding domain comprising VH CD28 comprising an amino acid sequence of SEQ ID NO:23 and VL CD28 comprising an amino acid sequence of SEQ ID NO:24.
In one aspect, the antibody that specifically binds to BCMA and CD28 comprises a polypeptide that is at least 95%, 96%, 97%, 98%, or 99% identical to the sequence of SEQ ID NO: 94, a polypeptide that is at least 95%, 96%, 97%, 98%, or 99% identical to the sequence of SEQ ID NO: 93, a polypeptide that is at least 95%, 96%, 97%, 98%, or 99% identical to the sequence of SEQ ID NO: 27, and a polypeptide that is at least 95%, 96%, 97%, 98%, or 99%
identical to the sequence of SEQ ID NO: 28. In one particular aspect, provided is an antibody that specifically binds to BCMA and CD28, wherein said antibody comprises a first light chain comprising the amino acid sequence of SEQ ID NO:94, a first heavy chain comprising the amino acid sequence of SEQ ID NO:93, a second heavy chain comprising the amino acid sequence of SEQ ID NO:27 and a second light chain comprising the amino acid sequence of SEQ ID NO:28.
In any of the aspects mentioned above, the antibody that specifically binds to BCMA and CD28, comprises a Fc domain. In one aspect, the Fc domain is an IgG, particularly an IgGl Fc domain. In one particular aspect, the Fc domain is a human Fc domain, particularly a human IgGl Fc domain. In any of the aspects described above, the Fc domain comprises one or more amino acid substitution that reduces binding to an Fc receptor and/or effector function. In one particular aspect, it comprises the amino acid mutations L234A, L235A and P329G (numbering according to Kabat EU index).
Fc domain modifications reducing Fc receptor binding and/or effector function
The Fc domain of the antibody that specifically binds to BCMA and CD28 consists of a pair of polypeptide chains comprising heavy chain domains of an immunoglobulin molecule. For example, the Fc domain of an immunoglobulin G (IgG) molecule is a dimer, each subunit of which comprises the CH2 and CH3 IgG heavy chain constant domains. The two subunits of the Fc domain are capable of stable association with each other. The Fc domain confers favorable pharmacokinetic properties to the antigen binding molecules of the invention, including a long serum half-life which contributes to good accumulation in the target tissue and a favorable tissue-blood distribution ratio. On the other side, it may, however, lead to undesirable targeting of the bispecific antibodies of the invention to cells expressing Fc receptors rather than to the preferred antigen-bearing cells.
Accordingly, the Fc domain of the antibody that specifically binds to BCMA and CD28 exhibits reduced binding affinity to an Fc receptor and/or reduced effector function, as compared to a native IgGl Fc domain. In one aspect, the Fc domain does not substantially bind to an Fc receptor and/or does not induce effector function. In a particular aspect, the Fc receptor is an Fey receptor. In one aspect, the Fc receptor is a human Fc receptor. In a specific aspect, the Fc receptor is an activating human Fey receptor, more specifically human FcyRIIIa, FcyRI or FcyRIIa, most specifically human FcyRIIIa. In one aspect, the Fc domain does not induce effector function. The reduced effector function can include, but is not limited to, one or more of the following: reduced complement dependent cytotoxicity (CDC), reduced antibody-dependent cell-mediated cytotoxicity (ADCC), reduced antibody-dependent cellular phagocytosis (ADCP), reduced cytokine secretion, reduced immune complex-mediated antigen uptake by antigen- presenting cells, reduced binding to NK cells, reduced binding to macrophages, reduced binding
to monocytes, reduced binding to polymorphonuclear cells, reduced direct signaling inducing apoptosis, reduced dendritic cell maturation, or reduced T cell priming.
In certain aspects, one or more amino acid modifications may be introduced into the Fc region of an antibody provided herein, thereby generating an Fc region variant. The Fc region variant may comprise a human Fc region sequence (e.g., a human IgGl, IgG2, IgG3 or IgG4 Fc region) comprising an amino acid modification (e.g. a substitution) at one or more amino acid positions.
In one particular aspect, provided is an antibody that specifically binds to BCMA and CD28, wherein the Fc region comprises one or more amino acid substitution that reduces binding to an Fc receptor, in particular towards Fey receptor. In one aspect, provided is an antibody that specifically binds to BCMA and CD28, wherein the Fc region comprises one or more amino acid substitution and wherein the ADCC induced by the antibody is reduced to 0-20% of the ADCC induced by an antibody comprising the wild-type human IgGl Fc region.
In one aspect, the Fc domain of the antibody described herein comprises one or more amino acid mutation that reduces the binding affinity of the Fc domain to an Fc receptor and/or effector function. Typically, the same one or more amino acid mutation is present in each of the two subunits of the Fc domain. In particular, the Fc domain comprises an amino acid substitution at a position of E233, L234, L235, N297, P331 and P329 (EU numbering). In particular, the Fc domain comprises amino acid substitutions at positions 234 and 235 (EU numbering) and/or 329 (EU numbering) of the IgG heavy chains. More particularly, provided is an antibody that specifically binds to BCMA and CD28 which comprises an Fc domain with the amino acid substitutions L234A, L235A and P329G (“P329G LALA”, EU numbering) in the IgG heavy chains. The amino acid substitutions L234A and L235A refer to the so-called LALA mutation. The “P329G LALA” combination of amino acid substitutions almost completely abolishes Fey receptor binding of a human IgGl Fc domain and is described in International Patent Appl. Publ. No. WO 2012/130831 Al which also describes methods of preparing such mutant Fc domains and methods for determining its properties such as Fc receptor binding or effector functions.
Fc domains with reduced Fc receptor binding and/or effector function also include those with substitution of one or more of Fc domain residues 238, 265, 269, 270, 297, 327 and 329 (U.S. Patent No. 6,737,056). Such Fc mutants include Fc mutants with substitutions at two or more of amino acid positions 265, 269, 270, 297 and 327, including the so-called “DANA” Fc mutant with substitution of residues 265 and 297 to alanine (US Patent No. 7,332,581).
In another aspect, the Fc domain is an IgG4 Fc domain. IgG4 antibodies exhibit reduced binding affinity to Fc receptors and reduced effector functions as compared to IgGl antibodies. In a more specific aspect, the Fc domain is an IgG4 Fc domain comprising an amino acid
substitution at position S228 (Kabat numbering), particularly the amino acid substitution S228P. In a more specific aspect, the Fc domain is an IgG4 Fc domain comprising amino acid substitutions L235E and S228P and P329G (EU numbering). Such IgG4 Fc domain mutants and their Fey receptor binding properties are also described in WO 2012/130831.
Mutant Fc domains can be prepared by amino acid deletion, substitution, insertion or modification using genetic or chemical methods well known in the art. Genetic methods may include site-specific mutagenesis of the encoding DNA sequence, PCR, gene synthesis, and the like. The correct nucleotide changes can be verified for example by sequencing.
Binding to Fc receptors can be easily determined e.g. by ELISA, or by Surface Plasmon Resonance (SPR) using standard instrumentation such as a BIAcore instrument (GE Healthcare), and Fc receptors such as may be obtained by recombinant expression. Alternatively, binding affinity of Fc domains or cell activating antibodies comprising an Fc domain for Fc receptors may be evaluated using cell lines known to express particular Fc receptors, such as human NK cells expressing Fcyllla receptor.
Effector function of an Fc domain, or antigen binding molecules of the invention comprising an Fc domain, can be measured by methods known in the art. A suitable assay for measuring ADCC is described herein. Other examples of in vitro assays to assess ADCC activity of a molecule of interest are described in U.S. Patent No. 5,500,362; Hellstrom et al. Proc Natl Acad Sci USA 83, 7059-7063 (1986) and Hellstrom et al., Proc Natl Acad Sci USA 82, 1499- 1502 (1985); U.S. Patent No. 5,821,337; Bruggemann et al., J Exp Med 166, 1351-1361 (1987). Alternatively, non-radioactive assays methods may be employed (see, for example, ACTEM nonradioactive cytotoxicity assay for flow cytometry (CellTechnology, Inc. Mountain View, CA); and CytoTox 96® non-radioactive cytotoxicity assay (Promega, Madison, WI)). Useful effector cells for such assays include peripheral blood mononuclear cells (PBMC) and Natural Killer (NK) cells. Alternatively, or additionally, ADCC activity of the molecule of interest may be assessed in vivo, e.g. in an animal model such as that disclosed in Clynes et al., Proc Natl Acad Sci USA 95, 652-656 (1998).
In some aspects, binding of the Fc domain to a complement component, specifically to Clq, is reduced. Accordingly, in some aspects wherein the Fc domain is engineered to have reduced effector function, said reduced effector function includes reduced CDC. Clq binding assays may be carried out to determine whether the bispecific antibodies of the invention are able to bind Clq and hence has CDC activity. See e.g., Clq and C3c binding ELISA in WO 2006/029879 and WO 2005/100402. To assess complement activation, a CDC assay may be performed (see, for example, Gazzano-Santoro et al., J Immunol Methods 202, 163 (1996); Cragg et al., Blood 101, 1045-1052 (2003); and Cragg and Glennie, Blood 103, 2738-2743 (2004)).
In one particular aspect, the Fc domain exhibiting reduced binding affinity to an Fc receptor and/or reduced effector function, as compared to a native IgGl Fc domain, is a human IgGl Fc domain comprising the amino acid substitutions L234A, L235A and optionally P329G, or a human IgG4 Fc domain comprising the amino acid substitutions S228P, L235E and optionally P329G (numberings according to Kabat EU index). More particularly, it is a human IgGl Fc domain comprising the amino acid substitutions L234A, L235A and P329G (numbering according to Kabat EU index).
Fc domain modifications promoting heterodimerization
The antibody that specifically binds to BCMA and CD28 comprises different antigenbinding sites, fused to one or the other of the two subunits of the Fc domain, thus the two subunits of the Fc domain may be comprised in two non-identical polypeptide chains. Recombinant co-expression of these polypeptides and subsequent dimerization leads to several possible combinations of the two polypeptides. To improve the yield and purity of the bispecific antigen binding molecules of the invention in recombinant production, it will thus be advantageous to introduce in the Fc domain of the antibody described herein a modification promoting the association of the desired polypeptides.
Accordingly, in particular aspects, provided is an antibody that specifically binds to BCMA and CD28 comprising a Fc domain composed of a first and a second subunit capable of stable association, wherein the Fc domain comprises a modification promoting the association of the first and second subunit of the Fc domain. The site of most extensive protein-protein interaction between the two subunits of a human IgG Fc domain is in the CH3 domain of the Fc domain. Thus, in one aspect said modification is in the CH3 domain of the Fc domain.
In a specific aspect said modification is a so-called “knob-into-hole” modification, comprising a “knob” modification in one of the two subunits of the Fc domain and a “hole” modification in the other one of the two subunits of the Fc domain. Thus, provided is an antibody that specifically binds to BCMA and CD28, wherein the antibody comprises a Fc domain composed of a first and a second subunit capable of stable association comprising one or more amino acid substitution that reduces the binding affinity of the antigen binding molecule to an Fc receptor and/or effector function, wherein the first subunit of the Fc domain comprises knobs and the second subunit of the Fc domain comprises holes according to the knobs into holes method. In a particular aspect, the first subunit of the Fc domain comprises the amino acid substitutions S354C and T366W (EU numbering) and the second subunit of the Fc domain comprises the amino acid substitutions Y349C, T366S and Y407V (numbering according to Kabat EU index).
The knob-into-hole technology is described e.g. in US 5,731,168; US 7,695,936; Ridgway et al., Prot Eng 9, 617-621 (1996) and Carter, J Immunol Meth 248, 7-15 (2001). Generally, the
method involves introducing a protuberance (“knob”) at the interface of a first polypeptide and a corresponding cavity (“hole”) in the interface of a second polypeptide, such that the protuberance can be positioned in the cavity so as to promote heterodimer formation and hinder homodimer formation. Protuberances are constructed by replacing small amino acid side chains from the interface of the first polypeptide with larger side chains (e.g. tyrosine or tryptophan). Compensatory cavities of identical or similar size to the protuberances are created in the interface of the second polypeptide by replacing large amino acid side chains with smaller ones (e.g. alanine or threonine).
Accordingly, in one aspect, in the CH3 domain of the first subunit of the Fc domain of the antibody that specifically binds to BCMA and CD28 an amino acid residue is replaced with an amino acid residue having a larger side chain volume, thereby generating a protuberance within the CH3 domain of the first subunit which is positionable in a cavity within the CH3 domain of the second subunit, and in the CH3 domain of the second subunit of the Fc domain an amino acid residue is replaced with an amino acid residue having a smaller side chain volume, thereby generating a cavity within the CH3 domain of the second subunit within which the protuberance within the CH3 domain of the first subunit is positionable. The protuberance and cavity can be made by altering the nucleic acid encoding the polypeptides, e.g. by site-specific mutagenesis, or by peptide synthesis. In a specific aspect, in the CH3 domain of the first subunit of the Fc domain the threonine residue at position 366 is replaced with a tryptophan residue (T366W), and in the CH3 domain of the second subunit of the Fc domain the tyrosine residue at position 407 is replaced with a valine residue (Y407V). In one aspect, in the second subunit of the Fc domain additionally the threonine residue at position 366 is replaced with a serine residue (T366S) and the leucine residue at position 368 is replaced with an alanine residue (L368A).
In yet a further aspect, in the first subunit of the Fc domain additionally the serine residue at position 354 is replaced with a cysteine residue (S354C), and in the second subunit of the Fc domain additionally the tyrosine residue at position 349 is replaced by a cysteine residue (Y349C). Introduction of these two cysteine residues results in formation of a disulfide bridge between the two subunits of the Fc domain, further stabilizing the dimer (Carter (2001), J Immunol Methods 248, 7-15). In a particular aspect, the first subunit of the Fc domain comprises the amino acid substitutions S354C and T366W (EU numbering) and the second subunit of the Fc domain comprises the amino acid substitutions Y349C, T366S and Y407V (numbering according to Kabat EU index).
In an alternative aspect, a modification promoting association of the first and the second subunit of the Fc domain comprises a modification mediating electrostatic steering effects, e.g. as described in PCT publication WO 2009/089004. Generally, this method involves replacement of one or more amino acid residues at the interface of the two Fc domain subunits by charged
amino acid residues so that homodimer formation becomes electrostatically unfavorable but heterodimerization electrostatically favorable.
The C-terminus of the heavy chain of the antibody as reported herein can be a complete C- terminus ending with the amino acid residues PGK. The C-terminus of the heavy chain can be a shortened C-terminus in which one or two of the C terminal amino acid residues have been removed. In one preferred aspect, the C-terminus of the heavy chain is a shortened C-terminus ending P. In one preferred aspect, the C-terminus of the heavy chain is a shortened C-terminus ending PG. In one aspect of all aspects as reported herein, a CD28 antigen binding molecule comprising a heavy chain including a C-terminal CH3 domain as specified herein, comprises the C -terminal glycine-lysine dipeptide (G446 and K447, numbering according to Kabat EU index). In one aspect of all aspects as reported herein, a CD28 antigen binding molecule comprising a heavy chain including a C-terminal CH3 domain, as specified herein, comprises a C-terminal glycine residue (G446, numbering according to Kabat EU index).
Modifications in the Fab domains
In one aspect, provided is an antibody that specifically binds to BCMA and CD28 characterized by monovalent binding to BCMA and CD28 comprising (a) a first antigen binding domain capable of specific binding to BCMA, (b) a second antigen binding domain capable of specific binding to CD28, and (c) a Fc domain composed of a first and a second subunit capable of stable association comprising one or more amino acid substitution that reduces the binding affinity of the antigen binding molecule to an Fc receptor and/or effector function, wherein the second antigen binding domain capable of specific binding to CD28 is a Fab fragment and in the Fab fragment either the variable domains VH and VL or the constant domains CHI and CL are exchanged according to the Crossmab technology.
In another aspect, provided is an antibody that specifically binds to BCMA and CD28 characterized by monovalent binding to BCMA and CD28 comprising (a) a first antigen binding domain capable of specific binding to BCMA, (b) a second antigen binding domain capable of specific binding to CD28, and (c) a Fc domain composed of a first and a second subunit capable of stable association comprising one or more amino acid substitution that reduces the binding affinity of the antigen binding molecule to an Fc receptor and/or effector function, wherein the first antigen binding domain capable of specific binding to BCMA is a Fab fragment and in the Fab fragment either the variable domains VH and VL or the constant domains CHI and CL are exchanged according to the Crossmab technology.
Multispecific antibodies with a domain replacement/exchange in one binding arm (CrossMabVH-VL or CrossMabCH-CL) are described in detail in W02009/080252 and Schaefer, W. et al, PNAS, 108 (2011) 11187-1191. They clearly reduce the byproducts caused
by the mismatch of a light chain against a first antigen with the wrong heavy chain against the second antigen (compared to approaches without such domain exchange).
In one aspect, the invention relates to an antibody that specifically binds to BCMA and CD28 comprising (a) a first antigen binding domain capable of specific binding to BCMA, (b) a second antigen binding domain capable of specific binding to CD28, and (c) a Fc domain composed of a first and a second subunit capable of stable association comprising one or more amino acid substitution that reduces the binding affinity of the antigen binding molecule to an Fc receptor and/or effector function, wherein in the Fab fragment capable of specific binding to CD28 the variable domains VL and VH are replaced by each other so that the VH domain is part of the light chain and the VL domain is part of the heavy chain. In another aspect, the invention relates to an antibody that specifically binds to BCMA and CD28 comprising (a) a first antigen binding domain capable of specific binding to BCMA, (b) a second antigen binding domain capable of specific binding to CD28, and (c) a Fc domain composed of a first and a second subunit capable of stable association comprising one or more amino acid substitution that reduces the binding affinity of the antigen binding molecule to an Fc receptor and/or effector function, wherein in the Fab fragment capable of specific binding to BCMA the variable domains VL and VH are replaced by each other so that the VH domain is part of the light chain and the VL domain is part of the heavy chain.
In another aspect, and to further improve correct pairing, the antibody that specifically binds to BCMA and CD28 (characterized by monovalent binding to CD28) comprises (a) a first antigen binding domain capable of specific binding to BCMA, (b) a second antigen binding domains capable of specific binding to a CD28, and (c) a Fc domain composed of a first and a second subunit capable of stable association comprising one or more amino acid substitution that reduces the binding affinity of the antigen binding molecule to an Fc receptor and/or effector function, can contain different charged amino acid substitutions (so-called “charged residues”). These modifications are introduced in the crossed or non-crossed CHI and CL domains. In a particular aspect, the invention relates to an antibody that specifically binds to BCMA and CD28, wherein in one of CL domains the amino acid at position 123 (EU numbering) has been replaced by arginine (R) and the amino acid at position 124 (EU numbering) has been substituted by lysine (K) and wherein in one of the CHI domains the amino acids at position 147 (EU numbering) and at position 213 (EU numbering) have been substituted by glutamic acid (E). In one particular aspect, in the CL domain of the Fab fragment capable of specific binding to CD28 the amino acid at position 123 (EU numbering) has been replaced by arginine (R) and the amino acid at position 124 (EU numbering) has been substituted by lysine (K) and in the CHI domain of the Fab fragment capable of specific binding to CD28 the amino acids at position 147 (EU numbering) and at position 213 (EU numbering) have been substituted by glutamic acid (E).
Polynucleotides
The invention further provides isolated polynucleotides encoding a BCMA antibody or an antibody that specifically binds to BCMA and CD28 as described herein. The one or more isolated polynucleotides encoding the BCMA antibody or antibody that specifically binds to BCMA and CD28 may be expressed as a single polynucleotide that encodes the entire antigen binding molecule or as multiple (e.g., two or more) polynucleotides that are co-expressed. Polypeptides encoded by polynucleotides that are co-expressed may associate through, e.g., disulfide bonds or other means to form a functional antigen binding molecule. For example, the light chain portion of an immunoglobulin may be encoded by a separate polynucleotide from the heavy chain portion of the immunoglobulin. When co-expressed, the heavy chain polypeptides will associate with the light chain polypeptides to form the immunoglobulin. In some aspects, the isolated polynucleotide encodes the entire antibody that specifically binds to BCMA and CD28 as described herein. In other aspects, the isolated polynucleotide encodes a polypeptide comprised in the antibody that specifically binds to BCMA and CD28 as described herein. In certain aspects the polynucleotide or nucleic acid is DNA. In other aspects, a polynucleotide of the present invention is RNA, for example, in the form of messenger RNA (mRNA). RNA of the present invention may be single stranded or double stranded.
Recombinant Methods
BCMA antibodies or antibodies that specifically bind to BCMA and CD28 as described herein may be obtained, for example, by solid-state peptide synthesis (e.g. Merrifield solid phase synthesis) or recombinant production. For recombinant production one or more polynucleotide encoding the antibody that specifically binds to BCMA and CD28 or polypeptide fragments thereof, e.g., as described above, is isolated and inserted into one or more vectors for further cloning and/or expression in a host cell. Such polynucleotide may be readily isolated and sequenced using conventional procedures. In one aspect, a vector, preferably an expression vector, comprising one or more of the polynucleotides as described herein is provided. Methods which are well known to those skilled in the art can be used to construct expression vectors containing the coding sequence of the antibody (fragment) along with appropriate transcriptional/translational control signals. These methods include in vitro recombinant DNA techniques, synthetic techniques and in vivo recombination/genetic recombination. See, for example, the techniques described in Maniatis et al., MOLECULAR CLONING: A LABORATORY MANUAL, Cold Spring Harbor Laboratory, N.Y. (1989); and Ausubel et al., CURRENT PROTOCOLS IN MOLECULAR BIOLOGY, Greene Publishing Associates and Wiley Interscience, N.Y. (1989). The expression vector can be part of a plasmid, virus, or may be a nucleic acid fragment. The expression vector includes an expression cassette into which the polynucleotide encoding the antibody or polypeptide fragments thereof (i.e. the coding region) is
cloned in operable association with a promoter and/or other transcription or translation control elements. As used herein, a "coding region" is a portion of nucleic acid which consists of codons translated into amino acids. Although a "stop codon" (TAG, TGA, or TAA) is not translated into an amino acid, it may be considered to be part of a coding region, if present, but any flanking sequences, for example promoters, ribosome binding sites, transcriptional terminators, introns, 5' and 3' untranslated regions, and the like, are not part of a coding region. Two or more coding regions can be present in a single polynucleotide construct, e.g. on a single vector, or in separate polynucleotide constructs, e.g. on separate (different) vectors. Furthermore, any vector may contain a single coding region, or may comprise two or more coding regions, e.g. a vector of the present invention may encode one or more polypeptides, which are post- or co-translationally separated into the final proteins via proteolytic cleavage. In addition, a vector, polynucleotide, or nucleic acid of the invention may encode heterologous coding regions, either fused or unfused to a polynucleotide encoding the antibody of the invention or polypeptide fragments thereof, or variants or derivatives thereof. Heterologous coding regions include without limitation specialized elements or motifs, such as a secretory signal peptide or a heterologous functional domain. An operable association is when a coding region for a gene product, e.g. a polypeptide, is associated with one or more regulatory sequences in such a way as to place expression of the gene product under the influence or control of the regulatory sequence(s). Two DNA fragments (such as a polypeptide coding region and a promoter associated therewith) are "operably associated" if induction of promoter function results in the transcription of mRNA encoding the desired gene product and if the nature of the linkage between the two DNA fragments does not interfere with the ability of the expression regulatory sequences to direct the expression of the gene product or interfere with the ability of the DNA template to be transcribed. Thus, a promoter region would be operably associated with a nucleic acid encoding a polypeptide if the promoter was capable of effecting transcription of that nucleic acid. The promoter may be a cell- specific promoter that directs substantial transcription of the DNA only in predetermined cells. Other transcription control elements, besides a promoter, for example enhancers, operators, repressors, and transcription termination signals, can be operably associated with the polynucleotide to direct cell-specific transcription.
Suitable promoters and other transcription control regions are disclosed herein. A variety of transcription control regions are known to those skilled in the art. These include, without limitation, transcription control regions, which function in vertebrate cells, such as, but not limited to, promoter and enhancer segments from cytomegaloviruses (e.g. the immediate early promoter, in conjunction with intron-A), simian virus 40 (e.g. the early promoter), and retroviruses (such as, e.g. Rous sarcoma virus). Other transcription control regions include those derived from vertebrate genes such as actin, heat shock protein, bovine growth hormone and rabbit a-globin, as well as other sequences capable of controlling gene expression in eukaryotic
cells. Additional suitable transcription control regions include tissue-specific promoters and enhancers as well as inducible promoters (e.g. promoters inducible tetracyclins). Similarly, a variety of translation control elements are known to those of ordinary skill in the art. These include, but are not limited to ribosome binding sites, translation initiation and termination codons, and elements derived from viral systems (particularly an internal ribosome entry site, or IRES, also referred to as a CITE sequence). The expression cassette may also include other features such as an origin of replication, and/or chromosome integration elements such as retroviral long terminal repeats (LTRs), or adeno-associated viral (AAV) inverted terminal repeats (ITRs).
Polynucleotide and nucleic acid coding regions as described herein may be associated with additional coding regions which encode secretory or signal peptides, which direct the secretion of a polypeptide encoded by a polynucleotide of the present invention. For example, if secretion of the antibody or polypeptide fragments thereof is desired, DNA encoding a signal sequence may be placed upstream of the nucleic acid encoding an antibody as described herein or polypeptide fragments thereof. According to the signal hypothesis, proteins secreted by mammalian cells have a signal peptide or secretory leader sequence which is cleaved from the mature protein once export of the growing protein chain across the rough endoplasmic reticulum has been initiated. Those of ordinary skill in the art are aware that polypeptides secreted by vertebrate cells generally have a signal peptide fused to the N-terminus of the polypeptide, which is cleaved from the translated polypeptide to produce a secreted or "mature" form of the polypeptide. In certain aspects, the native signal peptide, e.g. an immunoglobulin heavy chain or light chain signal peptide is used, or a functional derivative of that sequence that retains the ability to direct the secretion of the polypeptide that is operably associated with it. Alternatively, a heterologous mammalian signal peptide, or a functional derivative thereof, may be used. For example, the wild-type leader sequence may be substituted with the leader sequence of human tissue plasminogen activator (TP A) or mouse P-glucuronidase. DNA encoding a short protein sequence that could be used to facilitate later purification (e.g. a histidine tag) or assist in labeling the antibody as described herein may be included within or at the ends of the polynucleotide encoding the antibody as described herein or polypeptide fragments thereof.
In a further aspect, a host cell comprising one or more polynucleotides as described herein is provided. In certain aspects, a host cell comprising one or more vectors as described herein is provided. The polynucleotides and vectors may incorporate any of the features, singly or in combination, described herein in relation to polynucleotides and vectors, respectively. In one aspect, a host cell comprises (e.g. has been transformed or transfected with) a vector comprising a polynucleotide that encodes (part of) an antibody as disclosed herein. As used herein, the term "host cell" refers to any kind of cellular system which can be engineered to generate the fusion proteins of the invention or fragments thereof. Host cells suitable for replicating and for
supporting expression of antigen binding molecules are well known in the art. Such cells may be transfected or transduced as appropriate with the particular expression vector and large quantities of vector containing cells can be grown for seeding large scale fermenters to obtain sufficient quantities of the antibody for clinical applications. Suitable host cells include prokaryotic microorganisms, such as E. coli, or various eukaryotic cells, such as Chinese hamster ovary cells (CHO), insect cells, or the like. For example, polypeptides may be produced in bacteria in particular when glycosylation is not needed. After expression, the polypeptide may be isolated from the bacterial cell paste in a soluble fraction and can be further purified. In addition to prokaryotes, eukaryotic microbes such as filamentous fungi or yeast are suitable cloning or expression hosts for polypeptide-encoding vectors, including fungi and yeast strains whose glycosylation pathways have been “humanized”, resulting in the production of a polypeptide with a partially or fully human glycosylation pattern. See Gerngross, Nat Biotech 22, 1409-1414 (2004), and Li et al., Nat Biotech 24, 210-215 (2006).
Suitable host cells for the expression of (glycosylated) polypeptides are also derived from multicellular organisms (invertebrates and vertebrates). Examples of invertebrate cells include plant and insect cells. Numerous baculoviral strains have been identified which may be used in conjunction with insect cells, particularly for transfection of Spodoptera frugiperda cells. Plant cell cultures can also be utilized as hosts. See e.g. US Patent Nos. 5,959,177, 6,040,498, 6,420,548, 7,125,978, and 6,417,429 (describing PLANTIBODIES™ technology for producing antibodies in transgenic plants). Vertebrate cells may also be used as hosts. For example, mammalian cell lines that are adapted to grow in suspension may be useful. Other examples of useful mammalian host cell lines are monkey kidney CV1 line transformed by SV40 (COS-7); human embryonic kidney line (293 or 293T cells as described, e.g., in Graham et al., J Gen Virol 36, 59 (1977)), baby hamster kidney cells (BHK), mouse sertoli cells (TM4 cells as described, e.g., in Mather, Biol Reprod 23, 243-251 (1980)), monkey kidney cells (CV1), African green monkey kidney cells (VERO-76), human cervical carcinoma cells (HELA), canine kidney cells (MDCK), buffalo rat liver cells (BRL 3 A), human lung cells (W138), human liver cells (Hep G2), mouse mammary tumor cells (MMT 060562), TRI cells (as described, e.g., in Mather et al., Annals N.Y. Acad Sci 383, 44-68 (1982)), MRC 5 cells, and FS4 cells. Other useful mammalian host cell lines include Chinese hamster ovary (CHO) cells, including dhfr- CHO cells (Urlaub et al., Proc Natl Acad Sci USA 77, 4216 (1980)); and myeloma cell lines such as YO, NS0, P3X63 and Sp2/0. For a review of certain mammalian host cell lines suitable for protein production, see, e.g., Yazaki and Wu, Methods in Molecular Biology, Vol. 248 (B.K.C. Lo, ed., Humana Press, Totowa, NJ), pp. 255-268 (2003). Host cells include cultured cells, e.g., mammalian cultured cells, yeast cells, insect cells, bacterial cells and plant cells, to name only a few, but also cells comprised within a transgenic animal, transgenic plant or cultured plant or animal tissue. In one embodiment, the host cell is a eukaryotic cell, preferably a mammalian cell, such as a Chinese
Hamster Ovary (CHO) cell, a human embryonic kidney (HEK) cell or a lymphoid cell (e.g., Y0, NSO, Sp20 cell). Standard technologies are known in the art to express foreign genes in these systems. Cells expressing a polypeptide comprising either the heavy or the light chain of an immunoglobulin, may be engineered so as to also express the other of the immunoglobulin chains such that the expressed product is an immunoglobulin that has both a heavy and a light chain.
In one aspect, a method of producing an antibody that specifically binds to BCMA and CD28 or polypeptide fragments thereof is provided, wherein the method comprises culturing a host cell comprising polynucleotides encoding the antibody or polypeptide fragments thereof, as provided herein, under conditions suitable for expression of the antibody or polypeptide fragments thereof, and recovering the antibody as described herein or polypeptide fragments thereof from the host cell (or host cell culture medium).
In certain aspects the antigen binding domain capable of specific binding to BCMA (e.g. a Fab fragment) forming part of the antibody comprises at least an immunoglobulin variable region capable of binding to an antigen. Variable regions can form part of and be derived from naturally or non-naturally occurring antibodies and fragments thereof. Methods to produce polyclonal antibodies and monoclonal antibodies are well known in the art (see e.g. Harlow and Lane, "Antibodies, a laboratory manual", Cold Spring Harbor Laboratory, 1988). Non-naturally occurring antibodies can be constructed using solid phase-peptide synthesis, can be produced recombinantly (e.g. as described in U.S. patent No. 4,186,567) or can be obtained, for example, by screening combinatorial libraries comprising variable heavy chains and variable light chains (see e.g. U.S. Patent. No. 5,969,108 to McCafferty).
Any animal species of immunoglobulin can be used for the methods described herein. Non-limiting immunoglobulins useful can be of murine, primate, or human origin. If the antibody is intended for human use, a chimeric form of immunoglobulin may be used wherein the constant regions of the immunoglobulin are from a human. A humanized or fully human form of the immunoglobulin can also be prepared in accordance with methods well known in the art (see e. g. U.S. Patent No. 5,565,332 to Winter). Humanization may be achieved by various methods including, but not limited to (a) grafting the non-human (e.g., donor antibody) CDRs onto human (e.g. recipient antibody) framework and constant regions with or without retention of critical framework residues (e.g. those that are important for retaining good antigen binding affinity or antibody functions), (b) grafting only the non-human specificity-determining regions (SDRs or a-CDRs; the residues critical for the antibody-antigen interaction) onto human framework and constant regions, or (c) transplanting the entire non-human variable domains, but "cloaking" them with a human-like section by replacement of surface residues. Humanized antibodies and methods of making them are reviewed, e.g., in Almagro and Fransson, Front
Biosci 13, 1619-1633 (2008), and are further described, e.g., in Riechmann et al., Nature 332, 323-329 (1988); Queen et al., Proc Natl Acad Sci USA 86, 10029-10033 (1989); US Patent Nos. 5,821,337, 7,527,791, 6,982,321, and 7,087,409; Jones et al., Nature 321, 522-525 (1986); Morrison et al., Proc Natl Acad Sci 81, 6851-6855 (1984); Morrison and Oi, Adv Immunol 44, 65-92 (1988); Verhoeyen et al., Science 239, 1534-1536 (1988); Padlan, Molec Immun 31(3), 169-217 (1994); Kashmiri et al., Methods 36, 25-34 (2005) (describing SDR (a-CDR) grafting); Padlan, Mol Immunol 28, 489-498 (1991) (describing “resurfacing”); Dall’Acqua et al., Methods 36, 43-60 (2005) (describing “FR shuffling”); and Osbourn et al., Methods 36, 61-68 (2005) and Klimka et al., Br J Cancer 83, 252-260 (2000) (describing the “guided selection” approach to FR shuffling). Particular immunoglobulins according to the invention are human immunoglobulins. Human antibodies and human variable regions can be produced using various techniques known in the art. Human antibodies are described generally in van Dijk and van de Winkel, Curr Opin Pharmacol 5, 368-74 (2001) and Lonberg, Curr Opin Immunol 20, 450-459 (2008). Human variable regions can form part of and be derived from human monoclonal antibodies made by the hybridoma method (see e.g. Monoclonal Antibody Production Techniques and Applications, pp. 51-63 (Marcel Dekker, Inc., New York, 1987)). Human antibodies and human variable regions may also be prepared by administering an immunogen to a transgenic animal that has been modified to produce intact human antibodies or intact antibodies with human variable regions in response to antigenic challenge (see e.g. Lonberg, Nat Biotech 23, 1117-1125 (2005). Human antibodies and human variable regions may also be generated by isolating Fv clone variable region sequences selected from human-derived phage display libraries (see e.g., Hoogenboom et al. in Methods in Molecular Biology 178, 1-37 (O’Brien et al., ed., Human Press, Totowa, NJ, 2001); and McCafferty et al., Nature 348, 552-554; Clackson et al., Nature 352, 624-628 (1991)). Phage typically display antibody fragments, either as single-chain Fv (scFv) fragments or as Fab fragments.
In certain aspects, the antigen binding domains comprised in the antibodies described herein are engineered to have enhanced binding affinity according to, for example, the methods disclosed in PCT publication WO 2012/020006 (see Examples relating to affinity maturation) or U.S. Pat. Appl. Publ. No. 2004/0132066. The ability of the antibodies of the invention to bind to a specific antigenic determinant can be measured either through an enzyme-linked immunosorbent assay (ELISA) or other techniques familiar to one of skill in the art, e.g. surface plasmon resonance technique (Liljeblad, et al., Glyco J 17, 323-329 (2000)), and traditional binding assays (Heeley, Endocr Res 28, 217-229 (2002)). Competition assays may be used to identify an antigen binding molecule that competes with a reference antibody for binding to a particular antigen. In certain aspects, such a competing antigen binding molecule binds to the same epitope (e.g. a linear or a conformational epitope) that is bound by the reference antigen binding molecule. Detailed exemplary methods for mapping an epitope to which an antigen
binding molecule binds are provided in Morris (1996) “Epitope Mapping Protocols”, in Methods in Molecular Biology vol. 66 (Humana Press, Totowa, NJ). In an exemplary competition assay, immobilized antigen is incubated in a solution comprising a first labeled antigen binding molecule that binds to the antigen and a second unlabeled antigen binding molecule that is being tested for its ability to compete with the first antigen binding molecule for binding to the antigen. The second antigen binding molecule may be present in a hybridoma supernatant. As a control, immobilized antigen is incubated in a solution comprising the first labeled antigen binding molecule but not the second unlabeled antigen binding molecule. After incubation under conditions permissive for binding of the first antibody to the antigen, excess unbound antibody is removed, and the amount of label associated with immobilized antigen is measured. If the amount of label associated with immobilized antigen is substantially reduced in the test sample relative to the control sample, then that indicates that the second antigen binding molecule is competing with the first antigen binding molecule for binding to the antigen. See Harlow and Lane (1988) Antibodies: A Laboratory Manual ch.14 (Cold Spring Harbor Laboratory, Cold Spring Harbor, NY).
Bispecific antibodies prepared as described herein may be purified by art-known techniques such as high performance liquid chromatography, ion exchange chromatography, gel electrophoresis, affinity chromatography, size exclusion chromatography, and the like. The actual conditions used to purify a particular protein will depend, in part, on factors such as net charge, hydrophobicity, hydrophilicity etc., and will be apparent to those having skill in the art. For affinity chromatography purification an antibody, ligand, receptor or antigen can be used to which the antigen binding molecule binds. For example, for affinity chromatography purification of antigen binding molecules of the invention, a matrix with protein A or protein G may be used. Sequential Protein A or G affinity chromatography and size exclusion chromatography can be used to isolate an antigen binding molecule essentially as described in the Examples. The purity of the antibody or fragments thereof can be determined by any of a variety of well-known analytical methods including gel electrophoresis, high pressure liquid chromatography, and the like. For example, the bispecific antibody expressed as described in the Examples was shown to be intact and properly assembled as demonstrated by reducing and non-reducing SDS-PAGE.
Assays
The BCMA antibody or the antibody that specifically binds to BCMA and CD28 provided herein may be identified, screened for, or characterized for its physical/chemical properties and/or biological activities by various assays known in the art.
1. Affinity assays
The affinity of the antigen binding molecule provided herein for the corresponding target can be determined in accordance with the methods set forth in the Examples by surface plasmon resonance (SPR), using standard instrumentation such as a Proteon instrument (Bio-rad), and receptors or target proteins such as may be obtained by recombinant expression. The affinity of the antigen binding molecule for the target cell antigen can also be determined by surface plasmon resonance (SPR), using standard instrumentation such as a Proteon instrument (Biorad), and receptors or target proteins such as may be obtained by recombinant expression. According to one aspect, KD is measured by surface plasmon resonance using a Proteon ® machine (Bio-Rad) at 25 °C.
2. Binding assays and other assays
Binding of the antibody or bispecific antibody provided herein to the corresponding receptor expressing cells may be evaluated using cell lines expressing the particular receptor or target antigen, for example by flow cytometry (FACS). In one aspect, CHO cells expressing human CD28 (parental cell line CHO-kl ATCC #CCL-61, modified to stably overexpress human CD28) are used in the binding assay.
In a further aspect, cancer cell lines expressing BCMA were used to demonstrate the binding of the bispecific antigen binding molecules to the target cell antigen.
3. Activity assays
In one aspect, assays are provided for identifying the biuological activity of the antibody that specifically binds to BCMA and CD28. Biological activity may include, e.g. T cell proliferation and cytokine secretion as measured with the methods as described in Example 4. Antigen binding molecules having such biological activity in vivo and/or in vitro are also provided.
Pharmaceutical Compositions, Formulations and Routes of Administration
In a further aspect, the invention provides pharmaceutical compositions comprising any of the antibodies that specifically bind to BCMA and CD28 provided herein, e.g., for use in any of the below therapeutic methods. In one aspect, a pharmaceutical composition comprises an antibody that specifically binds to BCMA and CD28 provided herein and at least one pharmaceutically acceptable excipient. In another aspect, a pharmaceutical composition comprises an antibody that specifically binds to BCMA and CD28 provided herein and at least one additional therapeutic agent, e.g., as described below.
Pharmaceutical compositions as disclosed herein comprise a therapeutically effective amount of one or more antigen binding molecules dissolved or dispersed in a pharmaceutically acceptable excipient. The phrases "pharmaceutical or pharmacologically acceptable" refers to molecular entities and compositions that are generally non-toxic to recipients at the dosages and concentrations employed, i.e. do not produce an adverse, allergic or other untoward reaction when administered to an animal, such as, for example, a human, as appropriate. The preparation of a pharmaceutical composition that contains at least one antibody that specifically binds to BCMA and CD28 and optionally an additional active ingredient will be known to those of skill in the art in light of the present disclosure, as exemplified by Remington's Pharmaceutical Sciences, 18th Ed. Mack Printing Company, 1990, incorporated herein by reference. In particular, the compositions are lyophilized formulations or aqueous solutions. As used herein, "pharmaceutically acceptable excipient" includes any and all solvents, buffers, dispersion media, coatings, surfactants, antioxidants, preservatives (e.g. antibacterial agents, antifungal agents), isotonic agents, salts, stabilizers and combinations thereof, as would be known to one of ordinary skill in the art.
Parenteral compositions include those designed for administration by injection, e.g. subcutaneous, intradermal, intralesional, intravenous, intraarterial intramuscular, intrathecal or intraperitoneal injection. For injection, the antibodies or bispecific antibodies may be formulated in aqueous solutions, preferably in physiologically compatible buffers such as Hanks' solution, Ringer's solution, or physiological saline buffer. The solution may contain formulation agents such as suspending, stabilizing and/or dispersing agents. Alternatively, the antibody that specifically binds to BCMA and CD28 may be in powder form for constitution with a suitable vehicle, e.g., sterile pyrogen-free water, before use. Sterile injectable solutions are prepared by incorporating the fusion proteins of the invention in the required amount in the appropriate solvent with various of the other ingredients enumerated below, as required. Sterility may be readily accomplished, e.g., by filtration through sterile filtration membranes. Generally, dispersions are prepared by incorporating the various sterilized active ingredients into a sterile vehicle which contains the basic dispersion medium and/or the other ingredients. In the case of sterile powders for the preparation of sterile injectable solutions, suspensions or emulsion, the preferred methods of preparation are vacuum-drying or freeze-drying techniques which yield a powder of the active ingredient plus any additional desired ingredient from a previously sterile- filtered liquid medium thereof. The liquid medium should be suitably buffered if necessary and the liquid diluent first rendered isotonic prior to injection with sufficient saline or glucose. The composition must be stable under the conditions of manufacture and storage, and preserved against the contaminating action of microorganisms, such as bacteria and fungi. It will be appreciated that endotoxin contamination should be kept minimally at a safe level, for example, less than 0.5 ng/mg protein. Suitable pharmaceutically acceptable excipients include, but are not
limited to: buffers such as phosphate, citrate, and other organic acids; antioxidants including ascorbic acid and methionine; preservatives (such as octadecyldimethylbenzyl ammonium chloride; hexamethonium chloride; benzalkonium chloride; benzethonium chloride; phenol, butyl or benzyl alcohol; alkyl parabens such as methyl or propyl paraben; catechol; resorcinol; cyclohexanol; 3-pentanol; and m-cresol); low molecular weight (less than about 10 residues) polypeptides; proteins, such as serum albumin, gelatin, or immunoglobulins; hydrophilic polymers such as polyvinylpyrrolidone; amino acids such as glycine, glutamine, asparagine, histidine, arginine, or lysine; monosaccharides, disaccharides, and other carbohydrates including glucose, mannose, or dextrins; chelating agents such as EDTA; sugars such as sucrose, mannitol, trehalose or sorbitol; salt-forming counter-ions such as sodium; metal complexes (e.g. Zn-protein complexes); and/or non-ionic surfactants such as polyethylene glycol (PEG). Aqueous injection suspensions may contain compounds which increase the viscosity of the suspension, such as sodium carboxymethyl cellulose, sorbitol, dextran, or the like. Optionally, the suspension may also contain suitable stabilizers or agents which increase the solubility of the compounds to allow for the preparation of highly concentrated solutions. Additionally, suspensions of the active compounds may be prepared as appropriate oily injection suspensions. Suitable lipophilic solvents or vehicles include fatty oils such as sesame oil, or synthetic fatty acid esters, such as ethyl cleats or triglycerides, or liposomes.
Active ingredients may be entrapped in microcapsules prepared, for example, by coacervation techniques or by interfacial polymerization, for example, hydroxymethylcellulose or gelatin-microcapsules and poly-(methylmethacylate) microcapsules, respectively, in colloidal drug delivery systems (for example, liposomes, albumin microspheres, microemulsions, nanoparticles and nanocapsules) or in macroemulsions. Such techniques are disclosed in Remington's Pharmaceutical Sciences (18th Ed. Mack Printing Company, 1990). Sustained-release preparations may be prepared. Suitable examples of sustained-release preparations include semipermeable matrices of solid hydrophobic polymers containing the polypeptide, which matrices are in the form of shaped articles, e.g. films, or microcapsules. In particular embodiments, prolonged absorption of an injectable composition can be brought about by the use in the compositions of agents delaying absorption, such as, for example, aluminum monostearate, gelatin or combinations thereof. Exemplary pharmaceutically acceptable excipients herein further include insterstitial drug dispersion agents such as soluble neutral -active hyaluronidase glycoproteins (sHASEGP), for example, human soluble PH-20 hyaluronidase glycoproteins, such as rHuPH20 (HYLENEX®, Baxter International, Inc.). Certain exemplary sHASEGPs and methods of use, including rHuPH20, are described in US Patent Publication Nos. 2005/0260186 and 2006/0104968. In one aspect, a sHASEGP is combined with one or more additional glycosaminoglycanases such as chondroitinases. Exemplary lyophilized antibody formulations are described in US Patent No. 6,267,958. Aqueous antibody formulations include
those described in US Patent No. 6,171,586 and W02006/044908, the latter formulations including a histidine-acetate buffer. In addition to the compositions described previously, the antibody that specifically binds to BCMA and CD28 may also be formulated as a depot preparation. Such long acting formulations may be administered by implantation (for example subcutaneously or intramuscularly) or by intramuscular injection. Thus, for example, the antibody that specifically binds to BCMA and CD28 may be formulated with suitable polymeric or hydrophobic materials (for example as an emulsion in an acceptable oil) or ion exchange resins, or as sparingly soluble derivatives, for example, as a sparingly soluble salt.
Pharmaceutical compositions comprising the antibody that specifically binds to BCMA and CD28 may be manufactured by means of conventional mixing, dissolving, emulsifying, encapsulating, entrapping or lyophilizing processes. Pharmaceutical compositions may be formulated in conventional manner using one or more physiologically acceptable carriers, diluents, excipients or auxiliaries which facilitate processing of the proteins into preparations that can be used pharmaceutically. Proper formulation is dependent upon the route of administration chosen. The antibody that specifically binds to BCMA and CD28 may be formulated into a composition in a free acid or base, neutral or salt form. Pharmaceutically acceptable salts are salts that substantially retain the biological activity of the free acid or base. These include the acid addition salts, e.g. those formed with the free amino groups of a proteinaceous composition, or which are formed with inorganic acids such as for example, hydrochloric or phosphoric acids, or such organic acids as acetic, oxalic, tartaric or mandelic acid. Salts formed with the free carboxyl groups can also be derived from inorganic bases such as for example, sodium, potassium, ammonium, calcium or ferric hydroxides; or such organic bases as isopropylamine, trimethylamine, histidine or procaine. Pharmaceutical salts tend to be more soluble in aqueous and other protic solvents than are the corresponding free base forms. The composition herein may also contain more than one active ingredient as necessary for the particular indication being treated, preferably those with complementary activities that do not adversely affect each other. Such active ingredients are suitably present in combination in amounts that are effective for the purpose intended. The formulations to be used for in vivo administration are generally sterile. Sterility may be readily accomplished, e.g., by filtration through sterile filtration membranes.
Therapeutic methods and compositions
Any of the antibodies that specifically bind to BCMA and CD28 provided herein may be used in therapeutic methods, either alone or in combination.
In one aspect, an antibody that specifically binds to BCMA and CD28 for use as a medicament is provided. In further aspects, an antibody that specifically binds to BCMA and CD28 for use in treating cancer is provided. In one particular aspect, an antibody that specifically binds to BCMA and CD28 for use in treating hematologic malignancies is provided. The term
“hematologic malignancies” includes diseases selected from the group consisting of multiple myeloma (MM), chronic lymphocytic leukemia, acute B-lymphoblastic leukemia, non-Hodgkin lymphoma (NHL), and Hodgkin lymphoma, but also acute myeloid leukemia and acute lymphoblastic leukemia. In one particular aspect, an antibody that specifically binds to BCMA and CD28 for use in treating multiple myeloma (MM) is provided.
In certain aspects, an antibody that specifically binds to BCMA and CD28 for use in a method of treatment is provided. In certain aspects, herein is provided an antibody that specifically binds to BCMA and CD28 for use in a method of treating an individual having cancer comprising administering to the individual an effective amount of the antibody that specifically binds to BCMA and CD28. In one such aspect, the method further comprises administering to the individual an effective amount of at least one additional therapeutic agent.
In one aspect, the antibody that specifically binds to BCMA and CD28 as described herein is for use in inhibiting the growth of BCMA-expressing cancer cells.
In certain aspects, an antibody that specifically binds to BCMA and CD28 for use in a method of treatment is provided. In certain aspects, herein is provided an antibody that specifically binds to BCMA and CD28 for use in a method of treating an individual having cancer comprising administering to the individual an effective amount of the antibody that specifically binds to BCMA and CD28. In another aspect, provided is an antibody that specifically binds to BCMA and CD28 for use in a method of treating an individual having BCMA-expressing cancer, in particular hematologic malignancies selected from the group consisting of multiple myeloma (MM), chronic lymphocytic leukemia, acute B-lymphoblastic leukemia, non-Hodgkin lymphoma (NHL), Hodgkin lymphoma, acute myeloid leukemia and acute lymphoblastic leukemia, comprising administering to the individual an effective amount of the antibody that specifically binds to BCMA and CD28. In one such aspect, the method further comprises administering to the individual an effective amount of at least one additional therapeutic agent.
In a further aspect, herein is provided for the use of an antibody that specifically binds to BCMA and CD28 as described herein in the manufacture or preparation of a medicament. In one embodiment, the medicament is for treatment of cancer, particularly BCMA-expressing cancer. In a further aspect, the medicament is for use in a method of treating cancer comprising administering to an individual having cancer an effective amount of the medicament. In one such aspect, the method further comprises administering to the individual an effective amount of at least one additional therapeutic agent, e.g., as described below. In another aspect, the medicament is for treatment of BCMA-expressing cancer. In a further aspect, the medicament is for use in a method of treating cancer, in particular BCMA-expressing cancer, comprising administering to an individual having cancer an effective amount of the medicament. In a further aspect, herein is provided a method for treating a cancer, in particular BCMA-expressing cancer. In one aspect, the method comprises administering to an individual having cancer an effective
amount of an antibody that specifically binds to BCMA and CD28. In one such aspect, the method further comprises administering to the individual an effective amount of at least one additional therapeutic agent, as described below. An “individual” according to any of the above aspects may be a human.
In a further aspect, herein are provided pharmaceutical formulations comprising any of the antibodies that specifically bind to BCMA and CD28 as reported herein, e.g., for use in any of the above therapeutic methods. In one aspect, a pharmaceutical formulation comprises an antibody that specifically binds to BCMA and CD28 as reported herein and a pharmaceutically acceptable carrier. In another aspect, a pharmaceutical formulation comprises an antibody that specifically binds to BCMA and CD28 as reported herein and at least one additional therapeutic agent.
An antibody that specifically binds to BCMA and CD28 as reported herein can be used either alone or in combination with other agents in a therapy. For instance, an antibody that specifically binds to BCMA and CD28 as reported herein may be co-administered with at least one additional therapeutic agent. Thus, an antibody that specifically binds to BCMA and CD28 as described herein for use in cancer immunotherapy is provided. In certain aspects, an antibody that specifically binds to BCMA and CD28 for use in a method of cancer immunotherapy is provided. An “individual” according to any of the above aspects is preferably a human.
Such combination therapies noted above encompass combined administration (where two or more therapeutic agents are included in the same or separate formulations), and separate administration, in which case, administration of the antibody as reported herein can occur prior to, simultaneously, and/or following, administration of the additional therapeutic agent or agents. In one aspect, administration of the antibody that specifically binds to BCMA and CD28 and administration of an additional therapeutic agent occur within about one month, or within about one, two or three weeks, or within about one, two, three, four, five, or six days, of each other.
An antigen binding molecule as reported herein (and any additional therapeutic agent) can be administered by any suitable means, including parenteral, intrapulmonary, and intranasal, and, if desired for local treatment, intralesional administration. Parenteral infusions include intramuscular, intravenous, intraarterial, intraperitoneal, or subcutaneous administration. Dosing can be by any suitable route, e.g. by injections, such as intravenous or subcutaneous injections, depending in part on whether the administration is brief or chronic. Various dosing schedules including but not limited to single or multiple administrations over various time-points, bolus administration, and pulse infusion are contemplated herein.
Antibodies that specifically bind to BCMA and CD28 as described herein would be formulated, dosed, and administered in a fashion consistent with good medical practice. Factors for consideration in this context include the particular disorder being treated, the particular mammal being treated, the clinical condition of the individual patient, the cause of the disorder, the site of delivery of the agent, the method of administration, the scheduling of administration,
and other factors known to medical practitioners. The antibody that specifically binds to BCMA and CD28 needs not be, but is optionally formulated with one or more agents currently used to prevent or treat the disorder in question. The effective amount of such other agents depends on the amount of antibody present in the formulation, the type of disorder or treatment, and other factors discussed above. These are generally used in the same dosages and with administration routes as described herein, or about from 1 to 99% of the dosages described herein, or in any dosage and by any route that is empirically/clinically determined to be appropriate.
For the prevention or treatment of disease, the appropriate dosage of an antibody that specifically binds to BCMA and CD28 as described herein (when used alone or in combination with one or more other additional therapeutic agents) will depend on the type of disease to be treated, the type of antibody, the severity and course of the disease, whether the antibody is administered for preventive or therapeutic purposes, previous therapy, the patient's clinical history and response to the antibody, and the discretion of the attending physician. The antibody that specifically binds to BCMA and CD28 is suitably administered to the patient at one time or over a series of treatments. Depending on the type and severity of the disease, about 1 pg/kg to 15 mg/kg (e.g. 0.5 mg/kg - 10 mg/kg) of bispecific agonistic CD28 antigen binding molecule can be an initial candidate dosage for administration to the patient, whether, for example, by one or more separate administrations, or by continuous infusion. One typical daily dosage might range from about 1 pg/kg to 100 mg/kg or more, depending on the factors mentioned above. For repeated administrations over several days or longer, depending on the condition, the treatment would generally be sustained until a desired suppression of disease symptoms occurs. One exemplary dosage of the antibody would be in the range from about 0.05 mg/kg to about 10 mg/kg. Thus, one or more doses of about 0.5 mg/kg, 2.0 mg/kg, 4.0 mg/kg or 10 mg/kg (or any combination thereof) may be administered to the patient. Such doses may be administered intermittently, e.g. every week or every three weeks (e.g. such that the patient receives from about two to about twenty, or e.g. about six doses of the antibody). An initial higher loading dose, followed by one or more lower doses may be administered. However, other dosage regimens may be useful. The progress of this therapy is easily monitored by conventional techniques and assays.
Other agents and treatments
As described before, the antibody that specifically binds to BCMA and CD28 may be administered in combination with one or more other agents in therapy. For instance, an antigen binding molecule as described herein may be co-administered with at least one additional therapeutic agent. The term “therapeutic agent” encompasses any agent that can be administered for treating a symptom or disease in an individual in need of such treatment. Such additional therapeutic agent may comprise any active ingredients suitable for the particular indication being treated, preferably those with complementary activities that do not adversely affect each other. In
certain aspects, an additional therapeutic agent is another anti-cancer agent, for example a microtubule disruptor, an antimetabolite, a topoisomerase inhibitor, a DNA intercalator, an alkylating agent, a hormonal therapy, a kinase inhibitor, a receptor antagonist, an activator of tumor cell apoptosis, or an anti angiogenic agent. In certain aspects, an additional therapeutic agent is an immunomodulatory agent, a cytostatic agent, an inhibitor of cell adhesion, a cytotoxic or cytostatic agent, an activator of cell apoptosis, or an agent that increases the sensitivity of cells to apoptotic inducers.
Thus, provided are antibodies that specifically bind to BCMA and CD28 as disclosed herein or pharmaceutical compositions comprising them for use in the treatment of cancer, wherein the bispecific antibody is administered in combination with a chemotherapeutic agent, radiation and/ or other agents for use in cancer immunotherapy.
Such other agents are suitably present in combination in amounts that are effective for the purpose intended. The effective amount of such other agents depends on the amount of fusion protein used, the type of disorder or treatment, and other factors discussed above. The bispecific antigen binding molecule or antibody of the invention are generally used in the same dosages and with administration routes as described herein, or about from 1 to 99% of the dosages described herein, or in any dosage and by any route that is empirically/clinically determined to be appropriate. Such combination therapies noted above encompass combined administration (where two or more therapeutic agents are included in the same or separate compositions), and separate administration, in which case, administration of the bispecific antigen binding molecule or antibody of the invention can occur prior to, simultaneously, and/or following, administration of the additional therapeutic agent and/or adjuvant.
In a further aspect, provided is an antibody that specifically binds to BCMA and CD28 as described herein for use in the treatment of cancer, in particular BCMA-expressing cancer, wherein the bispecific antigen binding molecule is administered in combination with another immunomodulator. The term “immunomodulator” refers to any substance including a monoclonal antibody that effects the immune system. The molecules of the inventions can be considered immunomodulators. Immunomodulators can be used as anti -neoplastic agents for the treatment of cancer. In one aspect, immunomodulators include, but are not limited to anti- CTLA4 antibodies (e.g. ipilimumab), anti-PDl antibodies (e.g. nivolumab or pembrolizumab), PD-L1 antibodies (e.g. atezolizumab, avelumab or durvalumab), OX-40 antibodies, 4-1BB antibodies and GITR antibodies. Such combination therapies noted above encompass combined administration (where two or more therapeutic agents are included in the same or separate compositions), and separate administration, in which case, administration of the bispecific antigen binding molecule can occur prior to, simultaneously, and/or following, administration of the additional therapeutic agent and/or adjuvant.
-n-
Combination with T cell bispecific antibodies
In one aspect, the antibodies that specifically bind to BCMA and CD28 may be administered in combination with T-cell activating anti-CD3 bispecific antibodies. The T-cell activating anti-CD3 bispecific antibodies are specific for a tumor-associated antigen, for example GPRC5D, CD38, FcRH5 or BCMA. In one aspect, the T-cell activating anti-CD3 bispecific antibody is an anti-GPRC5D/anti-CD3 bispecific antibody.
In one aspect, the anti-GPRC5D/anti-CD3 bispecific antibody comprises at least one antigen binding domain capable of specific binding to GPRC5D which comprises a heavy chain variable region (VHGPRC5D) comprising (i) CDR-H1 comprising the amino acid sequence of SEQ ID NO: 145, (ii) CDR-H2 comprising the amino acid sequence of SEQ ID NO: 146, and (iii) CDR-H3 comprising the amino acid sequence of SEQ ID NO: 147, and a light chain variable region (VLGPRC5D) comprising (iv) CDR-L1 comprising the amino acid sequence of SEQ ID NO: 148, (v) CDR-L2 comprising the amino acid sequence of SEQ ID NO: 149, and (vi) CDR-L3 comprising the amino acid sequence of SEQ ID NO: 150. In one aspect, the anti-GPRC5D/anti- CD3 bispecific antibody comprises at least one antigen binding domain capable of specific binding to GPRC5D which comprises a heavy chain variable region (VHGPRC5D) comprising an amino acid sequence selected from the group consisting of SEQ ID NO: 151, SEQ ID NO: 153, SEQ ID NO: 154 and SEQ ID NO: 155 and a light chain variable region (VLGPRC5D) comprising an amino acid sequence selected from the group consisting of SEQ ID NO: 152, SEQ ID NO: 156, SEQ ID NO: 157, SEQ ID NO: 158, SEQ ID NO: 159 and SEQ ID NO:160.
In one aspect, the anti-GPRC5D/anti-CD3 bispecific antibody comprises at least one antigen binding domain capable of specific binding to GPRC5D which comprises a heavy chain variable region (VHGPRC5D) comprising an amino acid sequence that is at least about 95%, 96%, 97%, 98%, 99% or 100% identical to the amino acid sequence of SEQ ID NO: 151, and a light chain variable region (VLGPRC5D) comprising an amino acid sequence that is at least about 95%, 96%, 97%, 98%, 99% or 100% identical to the amino acid sequence of SEQ ID NO: 152. Particularly, the antigen binding domain capable of specific binding to GPRC5D comprises a heavy chain variable region (VHGPRC5D) comprising the amino acid sequence of SEQ ID NO: 151 and a light chain variable region (VLGPRC5D) comprising the amino acid sequence of SEQ ID NO: 152.
In another aspect, the anti-GPRC5D/anti-CD3 bispecific antibody comprises at least one antigen binding domain capable of specific binding to GPRC5D which comprises a heavy chain variable region (VHGPRC5D) comprising (i) CDR-H1 comprising the amino acid sequence of SEQ ID NO: 161, (ii) CDR-H2 comprising the amino acid sequence of SEQ ID NO: 162, and (iii) CDR-H3 comprising the amino acid sequence of SEQ ID NO: 163, and a light chain variable region (VLGPRC5D) comprising (iv) CDR-L1 comprising the amino acid sequence of SEQ ID NO: 164, (v) CDR-L2 comprising the amino acid sequence of SEQ ID NO: 165, and (vi) CDR-L3
comprising the amino acid sequence of SEQ ID NO: 166. In one aspect, the anti-GPRC5D/anti- CD3 bispecific antibody comprises at least one antigen binding domain capable of specific binding to GPRC5D which comprises a heavy chain variable region (VHGPRC5D) comprising an amino acid sequence selected from the group consisting of SEQ ID NO: 167, SEQ ID NO: 168, SEQ ID NO: 169, SEQ ID NO: 170, SEQ ID NO: 171 and SEQ ID NO: 172 and a light chain variable region (VLGPRC5D) comprising an amino acid sequence selected from the group consisting of SEQ ID NO: 173, SEQ ID NO: 174, SEQ ID NO: 175, SEQ ID NO: 176 and SEQ ID NO: 177. In another aspect, the anti-GPRC5D/anti-CD3 bispecific antibody comprises at least one antigen binding domain capable of specific binding to GPRC5D which comprises
(a) a heavy chain variable region (VHGPRC5D) comprising the amino acid sequence of SEQ ID NO: 151 and a light chain variable region (VLGPRC5D) comprising the amino acid sequence of SEQ ID NO: 152, or
(b) a heavy chain variable region (VHGPRC5D) comprising the amino acid sequence of SEQ ID NO: 155 and a light chain variable region (VLGPRC5D) comprising the amino acid sequence of SEQ ID NO: 158, or
(c) a heavy chain variable region (VHGPRC5D) comprising the amino acid sequence of SEQ ID NO: 151 and a light chain variable region (VLGPRC5D) comprising the amino acid sequence of SEQ ID NO: 158, or
(d) a heavy chain variable region (VHGPRC5D) comprising the amino acid sequence of SEQ ID NO: 167 and a light chain variable region (VLGPRC5D) comprising the amino acid sequence of SEQ ID NO: 175, or
(e) a heavy chain variable region (VHGPRC5D) comprising the amino acid sequence of SEQ ID NO: 169 and a light chain variable region (VLGPRC5D) comprising the amino acid sequence of SEQ ID NO: 174.
In one aspect, the anti-GPRC5D/anti-CD3 bispecific antibody comprises an antigen binding domain capable of specific binding to CD3 which comprises a heavy chain variable region (VHCD3) comprising (i) CDR-H1 comprising the amino acid sequence of SEQ ID NO: 178, (ii) CDR-H2 comprising the amino acid sequence of SEQ ID NO: 179, and (iii) CORED comprising the amino acid sequence of SEQ ID NO: 180, and a light chain variable region (VLCD3) comprising (iv) CDR-L1 comprising the amino acid sequence of SEQ ID NO: 181, (v) CDR-L2 comprising the amino acid sequence of SEQ ID NO: 182, and (vi) CDR-L3 comprising the amino acid sequence of SEQ ID NO: 183. In one aspect, the anti-GPRC5D/anti-CD3 bispecific antibody comprises an antigen binding domain capable of specific binding to CD3 which comprises a heavy chain variable region (VHCD3) comprising an amino acid sequence of SEQ ID NO: 184 and a light chain variable region (VLCD3) comprising an amino acid sequence of SEQ ID NO:185.
In another aspect, the anti-GPRC5D/anti-CD3 bispecific antibody comprises an antigen binding domain capable of specific binding to CD3 which comprises a heavy chain variable
region (VHCD3) comprising (i) CDR-H1 comprising the amino acid sequence of SEQ ID NO: 186, (ii) CDR-H2 comprising the amino acid sequence of SEQ ID NO: 187, and (iii) CORED comprising the amino acid sequence of SEQ ID NO: 188, and a light chain variable region (VLCD3) comprising (iv) CDR-L1 comprising the amino acid sequence of SEQ ID NO: 189, (v) CDR-L2 comprising the amino acid sequence of SEQ ID NO: 190, and (vi) CDR-L3 comprising the amino acid sequence of SEQ ID NO: 191. In one aspect, the anti-GPRC5D/anti-CD3 bispecific antibody comprises an antigen binding domain capable of specific binding to CD3 which comprises a heavy chain variable region (VHCD3) comprising an amino acid sequence of SEQ ID NO: 192 and a light chain variable region (VLCD3) comprising an amino acid sequence of SEQ ID NO:193.
In one aspect, the anti-GPRC5D/anti-CD3 bispecific antibody comprises an antigen binding domain capable of specific binding to CD3 which comprises a heavy chain variable region (VHCD3) comprising an amino acid sequence of SEQ ID NO: 184 and a light chain variable region (VLCD3) comprising an amino acid sequence of SEQ ID NO: 185, and two antigen binding domains capable of specifically binding to GPRC5D, wherein the antigen binding domains capable of specifically binding to GPRC5D each comprise a heavy chain variable region (VHGPRC5D) comprising the amino acid sequence of SEQ ID NO: 151 and a light chain variable region (VLGPRC5D) comprising the amino acid sequence of SEQ ID NO: 152. In one particular aspect, the anti-GPRC5D/anti-CD3 bispecific antibody comprises a polypeptide that is at least 95%, 96%, 97%, 98%, or 99% identical to the sequence of SEQ ID NO: 137, two polypeptides that are at least 95%, 96%, 97%, 98%, or 99% identical to the sequence of SEQ ID NO: 138, a polypeptide that is at least 95%, 96%, 97%, 98%, or 99% identical to the sequence of SEQ ID NO: 139, and a polypeptide that is at least 95%, 96%, 97%, 98%, or 99% identical to the sequence of SEQ ID NO: 140. In a further particular aspect, the bispecific antibody comprises a polypeptide sequence of SEQ ID NO: 137, two polypeptide sequences of SEQ ID NO: 138, a polypeptide sequence of SEQ ID NO: 139 and a polypeptide sequence of SEQ ID NO: 140 (GPRC5D CD3 TCB). In one particular aspect, the anti- GPRC5D/anti-CD3 bispecific antibody is forimtamig.
In another aspect, the anti-GPRC5D/anti-CD3 bispecific antibody comprises a polypeptide that is at least 95%, 96%, 97%, 98%, or 99% identical to the sequence of SEQ ID NO: 141, a polypeptide that is at least 95%, 96%, 97%, 98%, or 99% identical to the sequence of SEQ ID NO: 142, a polypeptide that is at least 95%, 96%, 97%, 98%, or 99% identical to the sequence of SEQ ID NO: 143, and a polypeptide that is at least 95%, 96%, 97%, 98%, or 99% identical to the sequence of SEQ ID NO: 144. In one aspect, the bispecific antibody comprises a polypeptide sequence of SEQ ID NO: 141, a polypeptide sequence of SEQ ID NO: 142, a polypeptide sequence of SEQ ID NO: 143 and a polypeptide sequence of SEQ ID NO: 144 (GPRC5D CD3 1+1 bispecific antibody).
In another aspect, the T-cell activating anti-CD3 bispecific antibody is an anti -BCM A/anti - CD3 bispecific antibody. In one aspect, the anti-BCMA/anti-CD3 bispecific antibody comprises an amino acid sequence of SEQ ID NO: 198, two amino acid sequences of SEQ ID NO: 199, an amino acid sequence of SEQ ID NO: 200 and amino acid sequence of SEQ ID NO: 201. In another aspect, the anti-BCMA/anti-CD3 bispecific antibody comprises an amino acid sequence of SEQ ID NO: 202, an amino acid sequence of SEQ ID NO: 203, an amino acid sequence of SEQ ID NO: 204 and amino acid sequence of SEQ ID NO: 205. In yet another aspect, the anti- BCMA/anti-CD3 bispecific antibody comprises an amino acid sequence of SEQ ID NO: 206, an amino acid sequence of SEQ ID NO: 207, an amino acid sequence of SEQ ID NO: 208 and amino acid sequence of SEQ ID NO: 209. In one aspect, the anti-BCMA/anti-CD3 bispecific antibody is selected from the group consisting of Alnuctamab, Elranatamab and Teclistamab.
In a further aspect, the T-cell activating anti-CD3 bispecific antibody is an anti- FcRH5/anti-CD3 bispecific antibody. Anti-FcRH5/anti-CD3 bispecific antibodies are for instance described in WO 2016/205520.
Such combination therapies noted above encompass combined administration (where two or more therapeutic agents are included in the same or separate formulations), and separate administration, in which case, administration of the therapeutic agent can occur prior to, simultaneously, and/or following, administration of an additional therapeutic agent or agents. In one embodiment, administration of the therapeutic agent and administration of an additional therapeutic agent occur within about one month, or within about one, two or three weeks, or within about one, two, three, four, five, or six days, of each other.
Articles of Manufacture
In another aspect of the invention, an article of manufacture containing materials useful for the treatment, prevention and/or diagnosis of the disorders described above is provided. The article of manufacture comprises a container and a label or package insert on or associated with the container. Suitable containers include, for example, bottles, vials, syringes, IV solution bags, etc. The containers may be formed from a variety of materials such as glass or plastic. The container holds a composition which is by itself or combined with another composition effective for treating, preventing and/or diagnosing the condition and may have a sterile access port (for example the container may be an intravenous solution bag or a vial having a stopper that is pierceable by a hypodermic injection needle). At least one active agent in the composition is an antibody that specifically binds to BCMA and CD28 as disclosed herein. The label or package insert indicates that the composition is used for treating the condition of choice. Moreover, the article of manufacture may comprise (a) a first container with a composition contained therein, wherein the composition comprises an antibody that specifically binds to BCMA and CD28; and
(b) a second container with a composition contained therein, wherein the composition comprises a further cytotoxic or otherwise therapeutic agent. The article of manufacture in this embodiment of the invention may further comprise a package insert indicating that the compositions can be used to treat a particular condition. Alternatively, or additionally, the article of manufacture may further comprise a second (or third) container comprising a pharmaceutically-acceptable buffer, such as bacteriostatic water for injection (BWFI), phosphate-buffered saline, Ringer's solution and dextrose solution. It may further include other materials desirable from a commercial and user standpoint, including other buffers, diluents, filters, needles, and syringes.
General information regarding the nucleotide sequences of human immunoglobulins light and heavy chains is given in: Kabat, E.A., et al., Sequences of Proteins of Immunological Interest, 5th ed., Public Health Service, National Institutes of Health, Bethesda, MD (1991). Amino acids of antibody chains are numbered and referred to according to the numbering systems according to Kabat (Kabat, E.A., et al., Sequences of Proteins of Immunological Interest, 5th ed., Public Health Service, National Institutes of Health, Bethesda, MD (1991)) as defined above.
The following numbered paragraphs (paras) describe aspects of the present invention:
1. An antibody that specifically binds to B cell maturation agent (BCMA), wherein the antibody comprises a first antigen binding domain comprising
(i) a heavy chain variable region (VH BCMA) comprising a heavy chain complementary determining region CDR-H1 of SEQ ID NO: 1 (GYTFTNYWMH), a CDR-H2 of SEQ ID NO: 2 (IIHPNSGSTNYNEKFQG), and a CDR-H3 of SEQ ID NO: 3 (GIYDYPFAY), and
(ii) a light chain variable region (VLBCMA) selected from the group consisting of
(a) a VL comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 4 (RASES VSIHGTHLMH), a CDR-L2 of SEQ ID NO: 5 (AASSLQS) and a CDR-L3 of SEQ ID NO: 6 (QQSIEDPYT); or
(b) a VL comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 4 (RASES VSIHGTHLMH), a CDR-L2 of SEQ ID NO: 7 (AASNLES) and a CDR-L3 of SEQ ID NO: 6 (QQSIEDPYT), or
(c) a VL comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 4 (RASES VSIHGTHLMH), a CDR-L2 of SEQ ID NO: 8 (AASNLQS) and a CDR-L3 of SEQ ID NO: 6 (QQSIEDPYT).
2. The antibody of para 1, wherein the VH BCMA comprises an amino acid sequence selected from the group consisting of SEQ ID NO: 9 (VHla) and SEQ ID NO: 10 (VHlb), and/or the VL BCMA comprises an amino acid sequence selected from the group consisting of SEQ ID NO: 11 (VLlf) , SEQ ID NO: 12 (VLla), SEQ ID NO: 13 (VLlb), SEQ ID NO: 14 (VLlc), SEQ ID NO: 15 (VLld), and SEQ ID NO: 16 (VLle).
3. An antibody that specifically binds to BCMA, wherein the antibody comprises a first antigen binding domain comprising
(a) a VH BCMA comprising an amino acid sequence of SEQ ID NOV and VLBCMA comprising an amino acid sequence of SEQ ID NO: 11, or
(b) a VH BCMA comprising an amino acid sequence of SEQ ID NOV and VLBCMA comprising an amino acid sequence of SEQ ID NO: 12.
4. The antibody of any one of paras 1 to 3, wherein the first antigen binding domain is a Fab molecule.
5. The antibody of any one of paras 1 to 4, comprising an Fc domain composed of a first and a second subunit.
6. The antibody of any one of paras 1 to 5, comprising a second antigen binding domain that specifically binds to a second antigen.
7. The antibody of any one of paras 1 to 6, wherein the second antigen binding domain that specifically binds to a second antigen is a Fab molecule wherein the variable domains VL and
VH or the constant domains CL and CHI, particularly the variable domains VL and VH, of the Fab light chain and the Fab heavy chain are replaced by each other.
8. The antibody of any one of paras 1 to 7, wherein the first antigen binding domain is a Fab molecule wherein in the constant domain CL the amino acid at position 123 (numbering according to Kabat EU index) is substituted by an amino acid selected from lysine (K), arginine (R) or histidine (H) and the amino acid at position 124 (numbering according to Kabat EU index) is substituted independently by lysine (K), arginine (R) or histidine (H), and wherein in the constant domain CHI the amino acid at position 147 (numbering according to Kabat EU index) is substituted independently by glutamic acid (E) or aspartic acid (D) and the amino acid at position 213 (numbering according to Kabat EU index) is substituted independently by glutamic acid (E), or aspartic acid (D) (numbering according to Kabat EU index).
9. The antibody of any one of paras 5 to 8, wherein the Fc domain is an IgG, particularly an IgGl Fc domain.
10. The antibody of any one of paras 5 to 9, wherein the Fc domain is a human Fc domain.
11. The antibody of any one of paras 5 to 10, wherein the Fc comprises a modification promoting the association of the first and the second subunit of the Fc domain.
12. The antibody of para 11, wherein the first subunit of the Fc domain comprises the amino acid substitutions S354C and T366W (EU numbering) and the second subunit of the Fc domain comprises the amino acid substitutions Y349C, T366S and Y407V (numbering according to Kabat EU index).
13. The antibody of any one of paras 5 to 12, wherein the Fc domain comprises one or more amino acid substitution that reduces binding to an Fc receptor and/or effector function.
14. The antibody of any one of paras 5 to 13, wherein the Fc domain is of human IgGl subclass and comprises the amino acid mutations L234A, L235A and P329G (numbering according to Kabat EU index).
15. The antibody of any one of paras 6 to 14, wherein the second antigen is CD28.
16. The antibody of para 15, wherein the second antigen binding domain that specifically binds to CD28 comprises a heavy chain variable region (VHCD28) comprising a heavy chain complementary determining region CDR-H1 of SEQ ID NO: 17, a CDR-H2 of SEQ ID NO: 18, and a CDR-H3 of SEQ ID NO: 19, and a light chain variable region (VLCD28) comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 20, a CDR-L2 of SEQ ID NO: 21 and a CDR-L3 of SEQ ID NO: 22.
17. The antibody of paras 15 or 16, wherein the second antigen binding domain that specifically binds to CD28 comprises a heavy chain variable region (VHCD28) comprising an amino acid sequence of SEQ ID NO:23, and a light chain variable region (VLCD28) comprising an amino acid sequence of SEQ ID NO:24 (v8).
18. The antibody of any one of paras 1 to 17, wherein the antibody comprises a first antigen binding domain comprising a VH BCMA comprising an amino acid sequence of SEQ ID NO:9 and VL BCMA comprising an amino acid sequence of SEQ ID NO: 11 and a second antigen binding domain comprising VH CD28 comprising an amino acid sequence of SEQ ID NO:23 and VL CD28 comprising an amino acid sequence of SEQ ID NO:24.
19. The antibody of any one of paras 1 to 18, comprising a first light chain comprising the amino acid sequence of SEQ ID NO:25, a first heavy chain comprising the amino acid sequence of SEQ ID NO:26, a second heavy chain comprising the amino acid sequence of SEQ ID NO:27 and a second light chain comprising the amino acid sequence of SEQ ID NO:28.
20. An antibody that specifically binds to B cell maturation agent (BCMA) and CD28, wherein the antibody comprises
(A) a first antigen binding domain comprising
(i) a heavy chain variable region (VH BCMA) selected from the group consisting of
(a) a VH comprising a heavy chain complementary determining region CDR-H1 of SEQ ID NO: 29 (GFTFSNAWMD), a CDR-H2 of SEQ ID NO: 30 (QITAKSNNYATYYADSVKG), and a CDR-H3 of SEQ ID NO: 31 (DGYH), and
(b) a VH comprising a heavy chain complementary determining region CDR-H1 of SEQ ID NO: 29 (GFTFSNAWMD), a CDR-H2 of SEQ ID NO: 32 (QITAKSNNYATYYAAPVKG), and a CDR-H3 of SEQ ID NO: 31 (DGYH), and
(ii) a light chain variable region (VLBCMA) comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 33 (RASEDIRNGLA), a CDR-L2 of SEQ ID NO: 34 (NANSLHT) and a CDR-L3 of SEQ ID NO: 35 (EDTSKYPYT), and
(B) a second antigen binding domain that specifically binds to CD28.
21. The antibody of para 20, wherein the VH BCMA comprises an amino acid sequence selected from the group consisting of SEQ ID NO: 36 (VH2a) and SEQ ID NO: 38 (VHlb), and/or the VL BCMA comprises an amino acid sequence selected from the group consisting of SEQ ID NO: 37 (VL2a) and SEQ ID NO:39 (VLla).
22. An antibody that specifically binds to BCMA and CD28, wherein the antibody comprises a first antigen binding domain comprising
(a) a VH BCMA comprising an amino acid sequence of SEQ ID NO:36 and a VL BCMA comprising an amino acid sequence of SEQ ID NO:37, or
(b) a VH BCMA comprising an amino acid sequence of SEQ ID NO:38 and a VLBCMA comprising an amino acid sequence of SEQ ID NO:39.
23. The antibody of any one of paras 20 to 22, wherein the first antigen binding domain is a Fab molecule.
24. The antibody of any one of paras 20 to 23, comprising an Fc domain composed of a first and a second subunit.
25. The antibody of any one of paras 20 to 24, comprising a second antigen binding domain that specifically binds to a second antigen.
26. The antibody of any one of paras 20 to 25, wherein the second antigen binding domain that specifically binds to a second antigen is a Fab molecule wherein the variable domains VL and VH or the constant domains CL and CHI, particularly the variable domains VL and VH, of the Fab light chain and the Fab heavy chain are replaced by each other.
27. The antibody of any one of paras 20 to 26, wherein the first antigen binding domain is a Fab molecule wherein in the constant domain CL the amino acid at position 123 (numbering according to Kabat EU index) is substituted by an amino acid selected from lysine (K), arginine (R) or histidine (H) and the amino acid at position 124 (numbering according to Kabat EU index) is substituted independently by lysine (K), arginine (R) or histidine (H), and wherein in the constant domain CHI the amino acid at position 147 (numbering according to Kabat EU index) is substituted independently by glutamic acid (E) or aspartic acid (D) and the amino acid at position 213 (numbering according to Kabat EU index) is substituted independently by glutamic acid (E), or aspartic acid (D) (numbering according to Kabat EU index).
28. The antibody of any one of paras 20 to 27, wherein the Fc domain is an IgG, particularly an IgGl Fc domain.
29. The antibody of any one of paras 24 to 28, wherein the Fc domain is a human Fc domain.
30. The antibody of any one of paras 24 to 29, wherein the Fc comprises a modification promoting the association of the first and the second subunit of the Fc domain.
31. The antibody of para 30, wherein the first subunit of the Fc domain comprises the amino acid substitutions S354C and T366W (EU numbering) and the second subunit of the Fc domain comprises the amino acid substitutions Y349C, T366S and Y407V (numbering according to Kabat EU index).
32. The antibody of any one of paras 24 to 31, wherein the Fc domain comprises one or more amino acid substitution that reduces binding to an Fc receptor and/or effector function.
33. The antibody of any one of paras 24 to 32, wherein the Fc domain is of human IgGl subclass and comprises the amino acid mutations L234A, L235A and P329G (numbering according to Kabat EU index).
34. The antibody of any one of paras 20 to 33, wherein the second antigen binding domain that specifically binds to CD28 comprises a heavy chain variable region (VHCD28) comprising a heavy chain complementary determining region CDR-H1 of SEQ ID NO: 17, a CDR-H2 of SEQ ID NO: 18, and a CDR-H3 of SEQ ID NO: 19, and a light chain variable region (VLCD28) comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 20, a CDR-L2 of SEQ ID NO: 21 and a CDR-L3 of SEQ ID NO: 22.
35. The antibody of para 34, wherein the second antigen binding domain that specifically binds to CD28 comprises a heavy chain variable region (VHCD28) comprising an amino acid sequence of SEQ ID NO:23, and a light chain variable region (VLCD28) comprising an amino acid sequence of SEQ ID NO: 24.
36. The antibody of any one of paras 20 to 35, wherein the antibody comprises a first antigen binding domain comprising a VH BCMA comprising an amino acid sequence of SEQ ID NO:36 and VL BCMA comprising an amino acid sequence of SEQ ID NO:37 and a second antigen binding domain comprising VH CD28 comprising an amino acid sequence of SEQ ID NO:23 and VL CD28 comprising an amino acid sequence of SEQ ID NO:24.
37. The antibody of any one of paras 20 to 36, comprising a first light chain comprising the amino acid sequence of SEQ ID NO:40, a first heavy chain comprising the amino acid sequence of SEQ ID NO:41, a second heavy chain comprising the amino acid sequence of SEQ ID NO:27 and a second light chain comprising the amino acid sequence of SEQ ID NO:28.
38. One or more isolated polynucleotide encoding the antibody of any one of paras 1 to 37.
39. One or more vector, particularly expression vector, comprising the polynucleotide(s) of para 38.
40. A host cell comprising the polynucleotide(s) of para 38 or the vector(s) of para 39.
41. A method of producing an antibody that specifically binds to BCMA, comprising the steps of a) culturing the host cell of para 40 under conditions suitable for the expression of the antibody and optionally b) recovering the antibody.
42. An antibody that specifically binds to BCMA produced by the method of para 41.
43. A pharmaceutical composition comprising the antibody of any one of paras 1 to 37 or 42 and at least one pharmaceutically acceptable excipient.
44. The antibody of any one of paras 1 to 37 or 42, or the pharmaceutical composition of claim 43, for use as a medicament.
45. The antibody of any one of paras 1 to 37 or 42, or the pharmaceutical composition of claim 43, for use in enhancing (a) T cell activation or (b) T cell effector functions.
46. The antibody of any one of paras 1 to 37 or 42, or the pharmaceutical composition of claim 43, for use in the treatment of a disease.
47. The antibody or the pharmaceutical composition for use of para 46, wherein the disease is cancer, in particular multiple myeloma.
48. The antibody of any one of paras 1 to 37 or 42, or the pharmaceutical composition of claim 43, for use in the treatment of cancer, wherein the use is for administration in combination with a chemotherapeutic agent, radiation therapy and/ or other agents for use in cancer immunotherapy.
49. The antibody of any one of paras 1 to 37 or 42, or the pharmaceutical composition of claim 43, for use in the treatment of cancer, wherein the use is for administration in combination with a T-cell activating anti-CD3 bispecific antibody.
50. The antibody or the pharmaceutical composition for use of para 49, wherein the T-cell activating anti-CD3 bispecific antibody is an anti-GPRC5D/anti-CD3 antibody.
52. Use of the antibody of any one of paras 1 to 37 or 42, or the pharmaceutical composition of para 43, in the manufacture of a medicament for the treatment of a disease, particularly for the treatment of cancer.
53. A method of treating a disease, particularly cancer, in an individual, comprising administering to said individual an effective amount of the antibody of paras 1 to 37 or 42, or the pharmaceutical composition of para 43.
54. The method of para 53, further comprising administration in combination with a chemotherapeutic agent, radiation therapy and/ or other agents for use in cancer immunotherapy, particularly in combination with a T-cell activating anti-CD3 bispecific antibody.
***
EXAMPLES
The following are examples of methods and compositions of the invention. It is understood that various other aspects may be practiced, given the general description provided above.
Recombinant DNA techniques: Standard methods were used to manipulate DNA as described in Sambrook et al., Molecular cloning: A laboratory manual; Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, 1989. The molecular biological reagents were used according to the manufacturer's instructions. General information regarding the nucleotide sequences of human immunoglobulin light and heavy chains is given in: Kabat, E.A. et al., (1991) Sequences of Proteins of Immunological Interest, Fifth Ed., NIH Publication No 91-3242.
DNA sequencing: DNA sequences were determined by double strand sequencing.
Gene synthesis: Desired gene segments, where required, were either generated by PCR using appropriate templates or were synthesized at Geneart AG (Regensburg, Germany) or Genscript (New Jersey, USA) from synthetic oligonucleotides and PCR products by automated gene synthesis. The gene segments flanked by singular restriction endonuclease cleavage sites were cloned into standard cloning / sequencing vectors. The plasmid DNA was purified from transformed bacteria and concentration determined by UV spectroscopy. The DNA sequence of the subcloned gene fragments was confirmed by DNA sequencing. Gene segments were designed with suitable restriction sites to allow subcloning into the respective expression vectors. All constructs were designed with a 5 ’-end DNA sequence coding for a leader peptide which targets proteins for secretion in eukaryotic cells.
Production of IgG and bispecific antibodies: The DNA sequences encoding the variable heavy and light chain regions of the BCMA antibodies (and, where applicable, the CD28 antibodies) were cloned into mammalian expression vectors using conventional cloning techniques. The antibodies described herein were produced using shaking flasks with FedBatch mode. The recombinant production was performed by transient transfection of Expi293™ Cells in a defined, serum-free medium. For transfection ExpiFectamine™ 293 Transfection Kit was used (Gibco). Cell culture supernatants were harvested 7-12 days after transfection.
Quantification of protein titer: The protein titer of supernatant samples was determined by affinity chromatography using a POROS A 20 pm column, 2.1 x 30 mm (Life Technologies, Carlsbad, CA, USA) on a High Performance Liquid Chromatography system (Ultimate 3000 HPLC system, Thermo Scientific, Waltham, MA, USA). The supernatant was loaded onto the column equilibrated with 0.2 M Na2HPO4, pH 7.4, followed by elution with 0.1 M citric acid, 0.2
M NaCl, pH 2.5. Titers were quantified by measuring absorption at 280 nm, and subsequently calculating the protein concentration by comparing the elution peak area (under the curve) of the analyte with a reference standard curve.
Alternatively, quantification of Fc containing constructs in supernatants was performed by Protein A - HPLC on an Agilent HPLC System with UV detector. Supernatants are injected on POROS A 20 pm (Applied Biosystems), washed with 10 mM Tris, 50 mM Glycine, 100 mM NaCl, pH 8.0 and eluted in the same buffer at pH 2.0. Titers were quantified by measuring absorption at 280 nm, and subsequently calculating the protein concentration by comparing the elution peak area (under the curve) of the analyte with a reference standard curve.
Purification of IgG and bispecific antibodies: Proteins were purified from cell culture supernatants referring to standard protocols. In brief, Fc containing proteins were purified from cell culture supernatants by Protein A-affinity chromatography (equilibration buffer: 20 mM sodium citrate, 20 mM sodium phosphate, pH 7.5 or PBS; elution buffer: 20 mM, 25 mM or 50 mM sodium citrate, pH 3.0). Elution was achieved at pH 3.0 followed by immediate pH neutralization of the sample. The protein was concentrated by centrifugation (Millipore Amicon® ULTRA-15, #UFC903096), and aggregated protein was separated from monomeric protein by size exclusion chromatography in 20 mM histidine, 140 mM sodium chloride, pH 6.0.
Analytics of IgG and bispecific antibodies: The concentrations of purified proteins were determined by measuring the absorption at 280 nm using the mass extinction coefficient calculated on the basis of the amino acid sequence according to Pace, et al., Protein Science, 1995, 4, 2411-1423. Purity and molecular weight of the proteins were analyzed by CE-SDS in the presence and absence of a reducing agent using a LabChipGXII or LabChip GX Touch (Perkin Elmer). Determination of the aggregate content was performed by HPLC chromatography at 25°C using analytical size-exclusion column (TSKgel G3000 SW XL or UP- SW3000, Tosoh Bioscience) equilibrated in running buffer (200 mM KH2PO4, 250 mM KC1 pH 6.2, 0.02% NaNs). The final quality was good for all molecules with varying monomer content from around 70% to almost 100% monomer content and >90% purity on CE-SDS. In conclusion, all IgGs and bispecific antibodies were produced in good quality.
Mass spectrometry: This section describes the characterization of the multispecific antibodies with VH/VL exchange (VH/VL CrossMabs) with emphasis on their correct assembly. The expected primary structures were analyzed by electrospray ionization mass spectrometry (ESI-MS) of the deglycosylated intact CrossMabs and deglycosylated/plasmin digested or alternatively deglycosylated/limited LysC digested CrossMabs. The VH/VL CrossMabs were deglycosylated with N-Glycosidase F in a phosphate or Tris buffer at 37°C for up to 17 h at a protein concentration of 1 mg/ml. The plasmin or limited LysC (Roche) digestions were
performed with 100 pg deglycosylated VH/VL CrossMabs in a Tris buffer pH 8 at room temperature for 120 hours and at 37°C for 40 min, respectively. Prior to mass spectrometry the samples were desalted via HPLC on a Sephadex G25 column (GE Healthcare). The total mass was determined via ESI-MS on a maXis 4G UHR-QTOF MS system (Bruker Daltonik) equipped with a TriVersa NanoMate source (Advion).
Determination of binding and binding affinity of multispecific antibodies to the respective antigens using surface resonance (SPR) (BIACORE) Binding of the generated antibodies to the respective antigens is investigated by surface plasmon resonance using a BIACORE instrument (GE Healthcare Biosciences AB, Uppsala, Sweden). Briefly, for affinity measurements Goat- Anti -Human IgG, JTR 109-005-098 antibodies are immobilized on a CM5 chip via amine coupling for presentation of the antibodies against the respective antigen. Binding is measured in HBS buffer (HBS-P (10 mM HEPES, 150 mM NaCl, 0.005% Tween 20, ph 7.4), 25°C (or alternatively at 37°C). Antigen (R&D Systems or in house purified) was added in various concentrations in solution. Association was measured by an antigen injection of 80 seconds to 3 minutes; dissociation was measured by washing the chip surface with HBS buffer for 3 - 10 minutes and a KD value was estimated using a 1 : 1 Langmuir binding model. Negative control data (e.g. buffer curves) are subtracted from sample curves for correction of system intrinsic baseline drift and for noise signal reduction. The respective Biacore Evaluation Software is used for analysis of sensorgrams and for calculation of affinity data.
Example 1
Generation and Production of optimal anti-BCMA antibodies
1.1 Generation of humanized variants of anti-BCMA antibody E04
1.1.1 Methodology
Anti-BCMA antibody E04 is disclosed in WO 2012/163805 and has the VH domain of SEQ ID NO:42 and the VL domain of SEQ ID NO:43. Optimized variants thereof were created as described in the following. For the identification of a suitable human acceptor framework during the humanization, a combination of two methodologies was used. On the one hand, a classical approach was taken by querying a BLASTp database of human V- and J-region sequences for the murine input sequences (cropped to the variable part). Selective criteria for the choice of human acceptor framework were sequence homology, same or similar CDR lengths, and the estimated frequency of the human germline, but also the conservation of certain amino acids at the VH-VL domain interface. Following the germline identification step, the CDRs of the murine input sequences were grafted onto the human acceptor framework regions. Each amino acid difference between these initial CDR grafts and the parental antibody was rated for
possible impact on the structural integrity of the respective variable region, and “back mutations” towards the parental sequence were introduced whenever deemed appropriate. The structural assessment was based on Fv region homology models of both the parental antibody and the humanization variants, created with an in-house antibody structure homology modeling protocol implemented using the BIO VIA Discovery Studio Environment, version 17R2. In some humanization variants, “forward mutations” were included, i.e., amino acid exchanges that change the original amino acid occurring at a given CDR position of the parental binder to the amino acid found at the equivalent position of the human acceptor germline. The aim is to increase the overall human character of the humanization variants (beyond the framework regions) to further reduce the immunogenicity risk.
On the other hand, an in silico tool developed in-house was used to predict the VH-VL domain orientation of the paired VH and VL humanization variants (WO 2016/062734). The results were compared to the predicted VH-VL domain orientation of the parental binder to select for framework combinations which are close in geometry to the original antibody. The rational is to detect possible amino acid exchanges in the VH-VL interface region that might lead to disruptive changes in the pairing of the two domains that in turn might have detrimental effects on the binding properties.
1.1.2 Choice of acceptor framework and adaptations thereof
The acceptor framework was chosen as described in Table 1 below:
Post-CDR3 framework regions were adapted from human IGHJ germline IGHJ4*01 (YFDYWGQGTLVTVSS, SEQ ID NO: 194) and human IGKJ germline IGKJ4*01 (LTFGGGTKVEIK, SEQ ID NO: 195). The part relevant for the acceptor framework is indicated as underlined.
Based on structural considerations, back mutations from the human acceptor framework to the amino acid in the parental clone were introduced at certain positions of the E04 humanization variants. Furthermore, some positions were identified as promising candidates for forward mutations, where the amino acid in a CDR of the parental binder is substituted by the amino acid found in the human acceptor germline. The changes are detailed in Table 2 below.
Table 2: List of Variants
Note: Back mutations are prefixed with b, forward mutations with f, e.g., bS49A refers to a back mutation (human germline amino acid to parental antibody amino acid) from serine to alanine at position 49. All residue indices are given in Kabat numbering. 1.1.3 T-cell Epitope Prediction
To assess the occurrence of potential T-cell epitopes in the humanized sequences, the NetMHCIIpan 4.0 predictor (Reynisson B et. al: NetMHCpan-4.1 and NetMHCIIpan-4.0: improved predictions of MHC antigen presentation by concurrent motif deconvolution and integration of MS MHC eluted ligand data, Nucl. Acids Res., 48(W1): W449-W454 (2020)) was employed. The predictions were made for the following human MHC class II alleles: DRBl*01:01, DRBl*03:01, DRBl*04:01, DRBl*07:01, DRBl*08:01, DRBl*09:01, DRBl*l l:01, DRBl*13:01, and DRBl*15:01.
The thresholds for strong and weak binding 15mer peptides were set to a percentile rank of 1 and 5, respectively. Binding 15mer peptides with a percentile rank above 5 were not considered. Likewise, all binding 15mer peptides with a 9mer core peptide occurring in 10 or more human V-region germlines were not considered. The germline sequences were obtained from the IMGT database (Giudicelli, V. et al.: IMGT/LIGM-DB, the IMGT® comprehensive
database of immunoglobulin and T cell receptor nucleotide sequences. Nucl. Acids Res., 34(S1):D781-D784 (2006)). As many of the predicted 15mer binders are sharing the same 9mer core peptide, Table 3 below also details the number of unique 9mer cores present in the respective sequence and predicted to bind in the percentile rank range < 5. Table 3: T cell epitopes
1.1.4 VH and VL domains of the resulting humanized BCMA antibodies
The resulting VH domains of the humanized BCMA antibodies can be found in Table 4 below and the resulting VL domains of the humanized BCMA antibodies are listed in Table 5 below.
Table 4: Amino acid sequences of the VH domains of humanized BCMA antibodies
The humanized amino acid sequences for heavy and light chain variable domains of E04 humanized variants were fused to a one-armed human IgGl backbone/human CH1-Hinge-CH2- CH3 with an effector silent Fc domain (P329G; L234A, L235A) to abrogate binding to Fey receptors according to the method described in WO 2012/130831 Al and into a light chain. For the right assembly of the one-armed IgGl a human Fc containing an effector silent Fc domain was used. The amino acid sequences were backtranslated in to DNA and the resulting cDNA were synthesized (GeneArt or Twist Biosciences) and then cloned into heavy chain expression vectors as fusion proteins with human IgGl backbones expression vectors as fusion proteins to human C-kappa. Light chain (LC) and heavy chain (HC) plasmids were then co-transfected into HEK293 cells and purified after 7 days from supernatants by standard methods for antibody purification.
1.2 Generation of humanized variants of anti-BCMA antibody 54
1.2.1 Methodology
Anti-BCMA antibody 54 is disclosed in WO 2013/072415 and has the VH domain of SEQ ID NO:44 and the VL domain of SEQ ID NO:45. BCMA-54 is a humanized antibody with 84.8% identity to the most similar human HV germline (IGHV3-15*01) and 83.2% identity to the most similar human KV germline (IGKV1-6*O1). Even though the variable region of BCMA-54 is based on framework regions of human origin, there are several positions in these regions which are not corresponding to the human germline amino acid. Examples for this include the positions VH-16 (Ala), VH-44 (Arg), VH-84 (Lys), VL-22 (Ala), VL-83 (Glu), and VL-95 (He). Furthermore, there exists the possibility to humanize parts of the CDRs to decrease the immunogenic potential of this binder and reduce the number of potential T-cell epitopes.
To realize this potential for optimization, suitable human acceptor frameworks were identified by querying a BLASTp database of human V- and J-region sequences for the original BCMA-54 sequences. Selective criteria for the choice of human acceptor framework were sequence homology, same or similar CDR lengths, and the estimated frequency of the human germline, but also the conservation of certain amino acids at the VH-VL domain interface. Following the germline identification step, the CDRs of the BCMA-54 input sequences were grafted onto the human acceptor framework regions. Each amino acid difference between these initial CDR grafts and the parental antibodies was rated for possible impact on the structural integrity of the respective variable region, and “back mutations” towards the parental sequence were introduced whenever deemed appropriate. The structural assessment was based on Fv region homology models of both BCMA-54 and the optimization candidate variants, created with an in-house antibody structure homology modeling protocol implemented using the BIO VIA Discovery Studio Environment, version 17R2. In some humanization variants, “forward mutations” were included, i.e., amino acid exchanges that change the original amino acid occurring at a given CDR position of the parental binder to the amino acid found at the equivalent position of the human acceptor germline.
An in silico tool developed in-house was used to predict the VH-VL domain orientation of the paired VH and VL humanization variants (WO 2016/062734). The results were compared to the predicted VH-VL domain orientation of the parental binder to select for framework combinations which are close in geometry to the original antibody. The rationale is to detect possible amino acid exchanges in the VH-VL interface region that might lead to disruptive changes in the pairing of the two domains that in turn might have detrimental effects on the binding properties..
1.2.2 Choice of acceptor framework and adaptations thereof
The following acceptor frameworks were chosen as described in Table 6 below:
Table 6: Acceptor framework
Post-CDR3 framework regions were adapted from human IGHJ germline IGHJ 1*01 (AEYFOHWGQGTLVTVSS, SEQ ID NO: 196) and human IGKJ germline IGKJ2*01 (YTF GQGTK LE IK, SEQ ID NO: 197). The part relevant for the acceptor framework is indicated as underlined.
Based on structural considerations, back mutations from the human acceptor framework to the amino acid in the original BCMA-54 sequence were introduced at certain positions of the optimization variants. Furthermore, some positions were identified as promising candidates for forward mutations, wherein the amino acid in a CDR of the parental binder is substituted by the amino acid found in the human acceptor germline. On top of the basis variants (VHla, VHlb, VH2a, VLla and VL2a) additional sequence variants were defined which typically introduce additional forward mutations (“germlining”), either of individual positions or stretches of the respective basis sequence. One variant (VHla_W197Y) aimed at ameliorating a predicted hydrophobic surface patch, thereby potentially improving the biophysical properties of the VH region. The changes are detailed in Table 7 below.
Table 7: List of Variants
Note: Back mutations are prefixed with b, forward mutations with f, e.g., bS49A refers to a back mutation (human germline amino acid to parental antibody amino acid) from serine to alanine at position 49. All residue indices are given in Kabat numbering..
1.2.3 T-cell Epitope Prediction
To assess the occurrence of potential T-cell epitopes in the humanized sequences, the NetMHCIIpan 4.0 predictor (Reynisson B et. al: NetMHCpan-4.1 and NetMHCIIpan-4.0: improved predictions of MHC antigen presentation by concurrent motif deconvolution and integration of MS MHC eluted ligand data, Nucl. Acids Res., 48(W1): W449-W454 (2020)) was employed. The predictions were made for the following human MHC class II alleles: DRBl*01:01, DRBl*03:01, DRBl*04:01, DRBl*07:01, DRBl*08:01, DRBl*09:01, DRBl*l l:01, DRBl*13:01, and DRBl*15:01.
The thresholds for strong and weak binding 15mer peptides were set to a percentile rank of 1 and 5, respectively. Binding 15mer peptides with a percentile rank above 5 were not considered. Likewise, all binding 15mer peptides with a 9mer core peptide occurring in 10 or more human V-region germlines were not considered. The germline sequences were obtained from the IMGT database (Giudicelli, V. et al.: IMGT/LIGM-DB, the IMGT® comprehensive database of immunoglobulin and T cell receptor nucleotide sequences. Nucl. Acids Res., 34(S1):D781-D784 (2006)). As many of the predicted 15mer binders are sharing the same 9mer core peptide, Table 8 below also details the number of unique 9mer cores present in the respective sequence and predicted to bind in the percentile rank range < 5.
1.2.4 VH and VL domains of the resulting humanized BCMA antibodies
The resulting VH domains of the humanized BCMA antibodies can be found in Table 9 below and the resulting VL domains of the humanized BCMA antibodies are listed in Table 10 below.
Table 9: Amino acid sequences of the VH domains of humanized BCMA antibodies
The humanized amino acid sequences for heavy and light chain variable domains of E04 humanized variants were fused to a one-armed human IgGl backbone/human CH1-Hinge-CH2- CH3 with an effector silent Fc domain (P329G; L234A, L235A) to abrogate binding to Fey receptors according to the method described in WO 2012/130831 Al and containing a knob mutation according to Knobs-into-Holes technology and into a light chain. For the right assembly of the one-armed IgGl a human Fc containing an effector silent Fc domain was used. The amino acid sequences were backtranslated into DNA and the resulting cDNA were synthesized (GeneArt or Twist Biosciences) and then cloned into heavy chain expression vectors as fusion proteins with human IgGl backbones expression vectors as fusion proteins to human C-kappa. Light chain (LC) and heavy chain (HC) plasmids were then co-transfected into HEK293 cells and purified after 7 days from supernatants by standard methods for antibody purification.
1.3 Characterization of the humanized anti-BCMA variants
In order to characterize the humanized anti-BCMA antibody variants, all clones were expressed as monovalent one-armed IgG-like constructs (Figure 1A). This format was chosen in order to characterize the binding to BCMA in a 1 : 1 model.
With the intention of selecting two preferred humanized variants of anti-BCMA antibody E04 and two preferred humanized variants of anti-BCMA antibody 54, 76 variants were produced. From these 76 variants, the 29 variants (15 BCMA 54 variants and 14 BCMA E04 variants) with the highest affinity to huBCMA were pre-selected and further characterized by measuring affinity to cyBCMA.
The corresponding VH/VL pairs, the construct ID (TaPIR ID) and the corresponding SEQ ID NO:s of the 29 variants and the parental antibodies are listed in Table 11 below.
The protein concentration of purified constructs was determined by measuring the optical density (OD) at 280 nm, using the mass extinction coefficient calculated on the basis of the amino acid sequence according to Pace, et al., Protein Science, 1995, 4, 2411-1423. Purity and molecular weight of the proteins were analyzed by CE-SDS in the presence and absence of a reducing agent using a LabChipGXII (Perkin Elmer). Determination of the aggregate content was performed by HPLC chromatography at 25°C using analytical size-exclusion column (TSKgel G3000 SW XL or UP-SW3000) equilibrated in running buffer (25 mM K2HPO4, 125 mM NaCl, 200mM L- Arginine Monohydrocloride, pH 6.7 or 200 mM KH2PO4, 250 mM KC1 pH 6.2 respectively). A summary of the product amount and purification parameters of all monovalent anti-BCMA variants is given in Table 12.
The binding kinetics of the monovalent humanized BCMA antibody variants to human BCMA and to cynomolgus BCMA were investigated by surface plasmon resonance (SPR) using a BIACORE T200 instrument (GE Healthcare). All experiments were performed at 25° C using HBS-P Buffer (10 mM HEPES, 150 mM NaCl pH 7.4, 0.05% surfactant P20) as running buffer and as dilution buffer. An anti-Fc IgG capturing antibody, specific for the PGLALA mutant Fc region, was immobilized on a series S Sensor Chip CM5 (Cytiva) by using standard amine coupling chemistry, resulting in a surface density of approximately 15000 resonance units (RU). Anti-BCMA antibodies were captured on the surface for 30 s at a flow rate of 5 pl/min leading to a capturing response of 50-200 RU. A dilution series of the antigen (human BCMA Fc homodimer (R&D Systems) or cynomolgus BCMA Fc homodimer (R&D Systems), respectively) was injected for 120 s at 30 pl/min with concentrations from 3 up to 300 nM onto the surface (association phase). The dissociation phase was monitored for 300-600 sec by washing with running buffer. The surface was regenerated by injecting 5 mM NaOH (freshly prepared) for 2 x 30 s. Bulk refractive index differences were corrected by subtracting blank injections and by substracting the response obtained from the reference flow cell without captured antibody. The derived curves were fitted to a 1 : 1 Langmuir binding model using the BIAevaluation software.
The KD values for the monovalent constructs including the 29 preferred BCMA variants are shown in Table 12 below.
Table 12: Production, Purification and Binding Properties expressed monovalent anti- BCMA variants
Based on the results of the in-silico assessment of the occurrence of potential T-cell epitopes in the humanized sequences with the NetMHCIIpan 4.0 predictor (see Example 1.2.3), the antibodies P1AG5080, P1AG5072, P1AG5028, P1AG5031, P1AG5063 and P1AG5036 (4 BCMA 54 variants and 2 BCMA E04 variants) were selected as the molecules with the lowest potential T-cell epitopes. In combination with their binding behavior, Pl AG5072 and Pl AG5031 were choosen as BCMA 54 variants and Pl AG5063 and Pl AG5036 were selected as the BCMA E04 variant antibodies to be included in the bispecific antibodies.
Example 2
Generation and Production of bispecific antigen binding molecules targeting BCMA
2.1 Cloning of bispecific antigen binding molecules targeting BCMA and CD28
For the generation of the expression plasmids, the sequences of the respective variable domains were used and sub-cloned in frame with the respective constant regions which are preinserted in the respective recipient mammalian expression vector. In the Fc domain, Pro329Gly, Leu234Ala and Leu235Ala mutations (PG-LALA) have been introduced in the constant region of the human IgGl heavy chains to abrogate binding to Fc gamma receptors according to the method described in International Patent Appl. Publ. No. WO 2012/130831. For the generation of bispecific antibodies, Fc fragments contained either the “knob” (S354C/T366W mutations, numbering according to Kabat EU index) or “hole” mutations (Y349C/T366S/L368A/Y407V mutations according to Kabat EU index) to avoid mispairing of the heavy chains. In order to avoid mispairing of light chains in the bispecific antigen binding molecules, exchange of VH/VL or CHl/Ckappa domains was introduced in one binding moiety (CrossFab technology). In another binding moiety, charges were introduced into the CHI and Ckappa domains as described in International Patent Appl. Publ. No. WO 2015/150447.
The generation and preparation of the anti-CD28 antibodies CD28 v.8 and CD28 v.15 as used herein is described in International Patent Application Publication No. WO 2020/127618 Al. CD28 v.8 has a VH of SEQ ID NO:23 and a VL of SEQ ID NO:24. CD28 v.15 has a VH of SEQ ID NO: 90 and a VL of SEQ ID NO:91.
Schematic illustrations of the bispecific antibody structures are shown in Figures IB or 1C. Table 13 summarizes specific anti-BCMA/anti-CD28 bispecific antibodies made, their identifiers and the sequences of the heavy chains (HC1 knob and HC2 hole) and light chains (LC1 and LC2).
2.2 Production and purification of bispecific antigen binding molecules targeting BCMA and CD28
The DNA sequences encoding the variable heavy and light chain regions of the BCMA and CD28 antigen binding domains were cloned into mammalian expression vectors using
conventional cloning techniques. The bispecific antibodies described herein were produced using shaking flasks with FedBatch mode. The recombinant production was performed by transient transfection of Expi293™ Cells in a defined, serum-free medium. For transfection ExpiFectamine™ 293 Transfection Kit was used (Gibco). Cell culture supernatants were harvested 7-12 days after transfection.
Quantification of protein titer: The protein titer of supernatant samples was determined by affinity chromatography using a POROS A 20 pm column, 2.1 x 30 mm (Life Technologies, Carlsbad, CA, USA) on a High Performance Liquid Chromatography system (Ultimate 3000 HPLC system, Thermo Scientific, Waltham, MA, USA). The supernatant was loaded onto the column equilibrated with 0.2 M Na2HPO4, pH 7.4, followed by elution with 0.1 M citric acid, 0.2 M NaCl, pH 2.5. Titers were quantified by measuring absorption at 280 nm, and subsequently calculating the protein concentration by comparing the elution peak area (under the curve) of the analyte with a reference standard curve.
Purification of bispecific antibodies: Proteins were purified from cell culture supernatants referring to standard protocols. In brief, Fc containing proteins were purified from cell culture supernatants by Protein A-affinity chromatography (equilibration buffer: 20 mM sodium citrate, 20 mM sodium phosphate, pH 7.5 or PBS; elution buffer: 20 mM, 25 mM or 50 mM sodium citrate, pH 3.0). Elution was achieved at pH 3.0 followed by immediate pH neutralization of the sample. The protein was concentrated by centrifugation (Millipore Amicon® ULTRA- 15, #UFC903096), and aggregated protein was separated from monomeric protein by size exclusion chromatography in 20 mM histidine, 140 mM sodium chloride, pH 6.0.
Analytics of bispecific antibodies: The concentrations of purified proteins were determined by measuring the absorption at 280 nm using the mass extinction coefficient calculated on the basis of the amino acid sequence according to Pace, et al., Protein Science, 1995, 4, 2411-1423. Purity and molecular weight of the proteins were analyzed by CE-SDS in the presence and absence of a reducing agent using a LabChipGXII or LabChip GX Touch (Perkin Elmer). Determination of the aggregate content was performed by HPLC chromatography at 25°C using analytical size-exclusion column (TSKgel G3000 SW XL or UP-SW3000, Tosoh Bioscience) equilibrated in running buffer (200 mM KH2PO4, 250 mM KC1 pH 6.2, 0.02% NaNs).
A summary of the purification parameters of selected molecules is given in Table 14.
Table 14: Summary of the production and purification of bispecific CD28 antigen binding molecules
2.3 Biophysical and biochemical characterization of bispecific antibodies targeting BCMA and CD28
In order to predict the developability of the bispecific antibodies, its biophysical and biochemical properties were assessed by calculation methods and assays.
A sequence-based prediction of the net charge of the protein as a function of pH by using standard software (e.g. the EMBOSS tool), and thus it isoelectric point (calc, pl) helped to estimate whether the bispecific antibody is suitable for binding and non-binding to cation and anion exchange chromatography media, and thus suitable for the general purification methods during production. All produced BCMA-CD28 bsAbs had favoarable pl values in the range of >8 (see Table 15 below).
Thermal stability of the BCMA-CD28 bsAbs prepared was monitored by Dynamic Light Scattering (DLS) and by monitoring of temperature dependent intrinsic protein fluorescence by applying a temperature ramp using an Optim 2 instrument (Avacta Analytical, UK). 10 pg of filtered protein sample with a protein concentration of 1 mg/ml was applied in duplicate to the Optim 2 instrument. The temperature was ramped from 25°C to 85°C at 0.1°C/min, with the ratio of fluorescence intensity at 350 nm/330 nm and scattering intensity at 266 nm being collected. The results are shown in Table 15. The aggregation temperature (Tagg) of all the tested BCMA-CD28 bsAbs produced is favorable.
The apparent hydrophobicity of the bispecific antibodies was assessed by hydrophobic interaction chromatography (HIC) as relative retention time compared to hydrophobicity standards (90 % of immunoglobulins in IVIG preparation have a relative retention time <0.35). In detail, 20 pg of sample was injected onto a HIC-Ether-5PW (Tosoh) column equilibrated with 25 mM Na-phosphate, 1.5 M ammonium sulfate, pH 7.0. Elution was performed with a linear gradient from 0 to 100% buffer B (25 mM Na-phosphate, pH 7.0) within 60 minutes. Retention times were then compared to protein standards with known hydrophobicity (e.g. Avastin).
For the FcRn affinity chromatography, FcRn was expressed, purified and biotinylated as described (Schlothauer et al., MAbs 2013, 5(4), 576-86). For coupling, the prepared receptor was added to streptavidin-sepharose (GE Healthcare). The resulting FcRn-sepharose matrix was packed in a column housing. The column was equilibrated with 20 mM 2-(N-morpholine)- ethanesulfonic acid (MES) and 140 mM NaCl, pH 5.5 (eluent A) at a 0.5 ml/min flow rate. 30 pg of antibody samples were diluted at a volume ratio of 1 : 1 with eluent A and applied to the FcRn column. The column was washed with 5 column volumes of eluent A followed by elution with a linear gradient from 20 to 100% 20 mM Tris/HCl and 140 mM NaCl, pH 8.8 (eluent B) in 35 column volumes. The analysis was performed with a column oven at 25 °C. The elution profile was monitored by continuous measurement of the absorbance at 280 nm. Retention times were compared to protein standards with known affinities.
Heparin affinity was determined by injecting 30-50 pg of sample onto a TSKgel Heparin- 5PW (Tosoh) column equilibrated with 50 mM Tris, pH 7.4. Elution was performed with a linear gradient from 0 to 100% buffer B (50 mM Tris, IM NaCl, pH 7.4 mM) within 37 minutes. Retention times were compared to protein standards with known affinities. The expected values for the retention times and the measured retention times for particular BCMA-CD28 bsAbs are shown in Table 15.
1 - Relative to generic hydrophobicity standards. 90 % of immunoglobulins in IVIG preparation have a relative retention time <0.35.
2 - Relative to generic FcRn affinity standard. Normal IgGs have a relative retention time between 0 and 2.75.
A characterization of binding potency by surface plasmon resonance (SPR) after stress of was carried out with particular BCMA-CD28 bsAbs. The results are shown in Table 16. The reduction in binding potency caused by incubation of the molecules for 14 days at 37°C, pH 7.4 and at 40°C, pH 6 was quantified by surface plasmon resonance using a Biacore T200 instrument (GE Healthcare). Samples stored at -80°C and pH 6 were used as reference. The reference samples and the samples stressed at 40°C were in 20 mM Histidine buffer, 140 mM NaCl, pH 6.0, and the samples stressed at 37°C in PBS buffer, pH 7.4, all at a concentration of 1.0 mg/ml. After the stress period (14 days) samples in PBS buffer were dialyzed back to 20 mM Histidine buffer, 140 mM NaCl, pH 6.0 for further analysis.
All SPR experiments were performed using a BIACORE instrument (GE Healthcare Biosciences AB, Uppsala, Sweden) at 25°C with HBS-P+ buffer (10 mM HEPES, 150 mM NaCl, pH 7.4, 0.05% Surfactant P20) as running and dilution buffer. Antigen (R&D Systems or in house purified) was added in various concentrations in solution. Biotinylated human BCMA and CD28, as well as biotinylated anti-hu IgG (Capture Select, Thermo Scientific, #7103262100) were immobilized on a Series S Sensor Chip SA (GE Healthcare, #29104992), resulting in surface densities of at least 1000 resonance units (RU). BCMA-CD28 bispecific antibodies with a concentration of 2 pg/ml were injected for 30 s at a flow rate of 5 pl/min, and dissociation was monitored for 120 s. The surface was regenerated by injecting 10 mM glycine buffer, pH 1.5, for 60 s. Bulk refractive index differences were corrected by subtracting blank injections and by subtracting the response obtained from a blank control flow cell. For evaluation, the binding response 5 seconds after injection end was taken.
To normalize the binding signal, the BCMA and CD28 binding was divided by the anti-hu IgG response (the signal (RU) obtained upon capture of the BCMA-CD28 bsAbs on the immobilized anti-hu IgG antibody). The relative binding activity was calculated by referencing each temperature stressed sample to the corresponding, non-stressed sample. As shown in Table 16, all BCMA-CD28 bsAbs showed a stabile binding upon stress to BCMA and CD28.
2.4 Production and purification of bispecific antigen binding molecules targeting BCMA and CD3
For comparison, the well-described BCMA-targeted CD3 T cell engagers Alnuctamab and Elranatamab were prepared. Alnuctamab is a BCMA x CD3 bispecific antibody in a 2+1 format, based on an IgGl Fc with L234A/L235A/P329G (EU numbering) mutations. The sequences are
identified in the International Non-Proprietary Nomenclature List (Recommended INN: List 85; WHO Drug Information, Vol. 35, No. 1, 2021). Alnuctamab comprises an amino acid sequence of SEQ ID NO: 198, two amino acid sequences of SEQ ID NO: 199, an amino acid sequence of SEQ ID NO: 200 and amino acid sequence of SEQ ID NO: 201 (2+1 format).
Elranatamab is a BCMA x CD3 bispecific antibody in a 1+1 format. The sequences are identified in the International Non-Proprietary Nomenclature List (Recommended INN: List 87, WHO Drug Information, Vol. 36, No. 1, 2022). Elranatamab comprises an amino acid sequence of SEQ ID NO: 202, an amino acid sequence of SEQ ID NO: 203, an amino acid sequence of SEQ ID NO: 204 and amino acid sequence of SEQ ID NO: 205.
For the production of Alnuctamab (P1AF0105), the DNA sequences encoding the variable heavy and light chain regions of the respective binding domains were cloned into mammalian expression vectors using conventional cloning techniques. Antibodies were generated by transient transfection of Expi293F cells. Cells were seeded in Expi293 media (Gibco, #1435101) at a density of 2.5 x 106/mL. Expression plasmids and ExpiFectamine (Gibco, ExpiFectamine transfection kit, #13385544) were separately mixed in OptiMEM (Gibco, #11520386). After 5 minutes both solutions were combined, mixed by pipetting and incubated for 15-20 minutes at room temperature. Cells were added to the plasmid/ExpiFectamine solution and incubated for 24 hours at 37°C in a shaking incubator with a 5% CO2 atmosphere. One day post transfection, supplements (Enhancer 1+2, ExpiFectamine transfection kit) were added. Cell supernatants were harvested after 4-5 days by centrifugation and subsequent filtration (0.2 pm filter), and proteins were purified from the harvested supernatant by standard methods as indicated below.
Elranatamab (P1AH5054) was produced and purified by Proteros according to their standard methods and protocols.
The quantification of Fc containing constructs in supernatants was performed by Protein A - HPLC on an Agilent HPLC System with UV detector. Supernatants are injected on POROS 20 A (Applied Biosystems), washed with 10 mM Tris, 50 mM Glycine, 100 mM NaCl, pH 8.0 and eluted in the same buffer at pH 2.0. Titers were quantified by measuring absorption at 280 nm, and subsequently calculating the protein concentration by comparing the elution peak area (under the curve) of the analyte with a reference standard curve.
Proteins were purified from filtered cell culture supernatants referring to standard protocols. In brief, Fc containing proteins were purified from cell culture supernatants by Protein A-affinity chromatography (equilibration buffer: 20 mM sodium citrate, 20 mM sodium phosphate, pH 7.5; elution buffer: 20 mM sodium citrate, pH 3.0). Elution was achieved at pH 3.0 followed by immediate pH neutralization of the sample. The protein was concentrated by centrifugation using Millipore Amicon® ULTRA- 15 (Merck, #UFC903096), and aggregated protein was separated
from monomeric protein by size exclusion chromatography in 20 mM histidine, 140 mM sodium chloride, pH 6.0. The concentrations of purified proteins were determined by measuring the absorption at 280 nm using the mass extinction coefficient calculated on the basis of the amino acid sequence according to Pace, et al., Protein Science, 1995, 4, 2411-1423. Purity and molecular weight of the proteins were analyzed by CE-SDS in the presence and absence of a reducing agent using a LabChipGXII or LabChip GX Touch (Perkin Elmer) (Perkin Elmer). Determination of the aggregate content was performed by HPLC chromatography at 25°C using analytical size-exclusion column (TSKgel G3000 SW XL or UP-SW3000) equilibrated in running buffer (200 mM KH2PO4, 250 mM KC1 pH 6.2, 0.02% NaNs). A summary of the purification parameters is given in Table 16A.
Table 16A: Summary of the production and purification of bispecific BCMA x CD3 antigen binding molecules
Teclistamab is a BCMA x CD3 bispecific antibody with an IgG4-F234A/L235A/S228P (EU numbering) Fc. The sequences are identified in the International Non-Proprietary Nomenclature List (Recommended INN: List 82, WHO Drug Information, Vol. 33, No. 3, 2019). It was obtained from a Supplier (FarmaMondo, Lot Nr. AT1334P1). Teclistamab comprises an amino acid sequence of SEQ ID NO: 206, an amino acid sequence of SEQ ID NO: 207, an amino acid sequence of SEQ ID NO: 208 and amino acid sequence of SEQ ID NO: 209.
Example 3
Binding of bispecific CD28 agonistic antigen binding molecules targeting BCMA to cells expressing CD28 or BCMA
To measure the binding to BCMA or CD28 we performed FACS-based binding assay on CHO transfectants, that were transduced to stably overexpress either human BCMA or human CD28.
CHO-K1 cell lines expressing BCMA extracellular domain non -truncating mutants were generated as follows: Full-length cDNA encoding human BCMA (UniProt: Q02223) and its corresponding mutants (R27P, S30del, P33S and P34del) were subcloned into lentiviral transfer vectors controlled by the CMV promoter. Lentiviral particles were prepared by transiently cotransfecting HEK 293-derived viral production cells (Gibco, #A35347) with transfer plasmid and lentiviral packaging mix (pRSV-Rev, pCgpV, and pCMV-VSV-G) using the LV-MAX Transfection Kit (Gibco, # A35346) according to the manufacturer's protocol. Viral supernatants were harvested 48 h after transfection, filtered through a 0.45-pm low protein binding filter and stored at -80°C until use.
One day prior to transduction, 5 x 104 CHO-K1 (ATCC CRL-9618) cells were seeded per well in a 24-well plate. The following day, the culture medium was replaced with 300 pL of purified lentiviral supernatant and 100 pL of fresh culture medium DMEM/F-12 (Gibco, #11320033) supplemented with 10% fetal bovine serum (Gibco, #16140063) and 1% GlutaMAX Supplement (Gibco; #31331-028). To facilitate viral transduction, the infectious medium was further supplemented with 2.5 pL of TransDux™ reagent and 100 pL of MAX Enhancer (SBI; # LV860A-1). The cells were then incubated for 24 hours at 37 °C. After the incubation period, the viral medium was discarded and the cells were subsequently maintained in fresh culture medium.
Three days post transduction, the culture medium was supplemented with 6 pg/mL puromycin (Invivogen; #ant-pr-l). After initial selection, the cells exhibiting human BCMA surface expression were isolated by BD FACSAria III cell sorter (BD Biosciences) and subsequently cultured to generate stable clones. Following a 4-week stability test, the surface expression and its stability were verified by flow cytometry analysis using mouse PE-conjugated anti-human BCMA (BioLegend, #357503).
The stable CHO transfectants (parental cell line CHO-kl ATCC #CCL-61) were cultured in DMEM/F-12 (Gibco, #10565018) supplemented with 10 fetal bovine serum (Gibco, #16140063 or Sigma-Aldrich in case of F4135) and 1% GlutaMAX Supplement (Gibco; #31331-028), including 6 pg/ml puromycin (Invivogen; #ant-pr-l). Adherent CHO cells were detached using Cell Dissociation Buffer (Gibco, #13151014) or trypsin (Gibco by ThermoFisher Scientific, TrypLE™Express Enzyme #2605-010), counted and checked for viability. All subsequent steps were performed at 4°C.
CD28 binding protocol
To assess binding to CD28, CHO-huCD28 cells were resuspended in FACS buffer (PBS, 2% Fetal Bovine Serum; 1% 0.5 M EDTA pH 8; 0.25% NaNs Sodium azide) at 1 Mio cells per ml. 0.1 Mio cells were plated per well of a round-bottom 96-well-plate and washed with 150 pl FACS buffer per well and supernatants were discarded. Cells were stained in a total volume of
50 pl per well and increasing concentrations of the indicated BCMA-CD28 bispecific molecules (0.48 pM - 2000 nM) for 60 minutes at 4°C. Afterwards, due to low affinity of the CD28 antigen binding domains - the cells were not washed but immediately fixed by adding 150 pl of 4% PF A (in PBS) on top of the antibody solution and incubating the plates at 4°C for 15 min. Next, plates were centrifuged (5 min, 450xg) and washed with 150 pl FACS buffer twice. The plates were analyzed on a BD Fortessa flow cytometer, equipped with the software FACS Diva. Binding curves and ECso values were obtained using GraphPadPrism6. The data as measured for various BCMA-CD28 bsAbs and controls are shown in Fig. 2A and 2B.
BCMA binding protocol
For the assessment of the binding to BCMA, CHO-huBCMA cells were resuspended in FACS buffer (PBS, 2% Fetal Bovine Serum; 1% 0.5 M EDTA pH 8; 0.25% NaNs Sodium azide) at 1 Mio cells per ml. 0.1 Mio cells were plated per well of a round-bottom 96-well-plate and washed with 150 pl FACS buffer per well and supernatants were discarded. Cells were stained in a total volume of 50 pl per well and increasing concentrations of the indicated BCMA-CD28 bispecific molecules (0.48 pM - 2000 nM) for 30 minutes at 4°C. Afterwards, the cells were centrifuged and washed with 150 pl FACS buffer twice.
A total of 25 pl per well of pre-diluted secondary antibody (PE-AffiniPure F(ab')2 Fragment Goat Anti-Human IgG, Fey Fragment Specific; Jackson Immunoresearch, 109-116- 170, 1 :50 diluted in FACS buffer) was added and plates were incubated for 30 minutes at 4°C. Cells were washed twice and then fixed by adding 50 pl of 1% PFA (in PBS) and an incubation of 20 min at RT. Cells were washed once with 150 pl FACS buffer and analyzed on a BD Fortessa flow cytometer, equipped with the software FACS Diva. Binding curves and ECso values were obtained using GraphPadPrism6. The data as measured for various BCMA-CD28 bsAbs and controls (for instance “2nd antibody only refers to samples containing only the flurophore labeled secondary antibody) are shown in Fig. 3A and 3B.
The bispecific antibodies tested comprised CD28 antibody CD28v8 or CD28vl5, respectively. Fig. 2A and 3A (molecules comprising antibody CD28v8) and Fig. 2B and 3B (molecules comprising antibody CD28vl5) show that all bispecific CD28 molecules are able to bind both, human CD28 (Fig. 2A and 2B), as well as human BCMA (Fig. 3 A and 3B) in a concentration-dependent manner. Binding to human CD28 does not reach saturation due to low affinity binders and is comparable among the CD28v8 bispecific antibodies (Fig. 2A) and CD28vl5 bispecific antibodies (Fig. 2B), respectively. When it comes to BCMA binding, the molecules with the BCMA antibody PR described in WO 2020/127618 Al have lower ECso values compared to the molecules with the new BCMA antibodies as described herein, but maximal binding (Emax) is comparable among all molecules tested. The ECso values are listed in Table 17 below.
Table 17: EC50 values (nM) and Emax for binding of the indicated bispecific BCMA-CD28 bsAbs to BCMA, expressed on cells
In a second assay, the bispecific antibodies tested comprised the CD28 antibody CD28v8 and the BCMA binder BCMA (54 2a2a) or BCMA (E04_lalf). Since one of the relapse mechanisms in MM can be non-truncating, missense mutations or in-frame deletions in the extracellular domain of BCMA, the above mentioned molecules were compared head-to-head to well described BCMA-targeted CD3 T cell engagers Teclistamab, Alnuctamab and Elranatamab (Lee et al., Nature Medicine 2023, 29, 2295-2306). For the assessment of the binding to BCMA variants with the indicated point mutations in the extracellular domain of hu BCMA, CHO transfectants were resuspended in PBS (Gibco by ThermoFisher Scientific, #20012050) and counted. Live-dead staining of target cells was performed by incubation of cells with 1 : 1000 diluted Zombie Aqua Viability dye (BioLegend #423102) in PBS at 1.5 x 106 cells per ml for 10 minutes in the dark, followed by a washing step with PBS and centrifugation at 400 x g at 4°C for 4 minutes. CHO-K1 cells were adjusted to 1.25 x 106 cells per ml in PBS and 40 pl were seeded per well of a 384-U bottom well plate
(ThermoFisher Scientific #264573). Plates were centrifuged once more at 400 x g at 4°C for 4 minutes and 20 pl supernatant was removed.
Cells were stained in a total volume of 40 pl per well by adding 20 pl of 2x concentrated dilution of the indicated BCMA-CD28 antigen binding molecules (final concentrations ranged from 0.008 - 125 nM). Cells were then incubated for 30 minutes at 4°C in the dark. Afterwards, the cells were centrifuged and washed with 40 pl FACS buffer three times.
A total of 20 pl per well of pre-diluted secondary antibody (R-Phycoerythrin AffiniPure Fab Fragment Goat Anti-Human IgG, Fey fragment specific; Jackson Immunoresearch, 109-117- 008, 1 : 100 diluted in PBS) was added to plates already containing 20 ul (final dilution 1 :200). Cells were incubated for 30 minutes at 4°C and then washed twice with 40 pl PBS per well and then fixed by adding 40 pl of 1% PFA (in PBS) overnight at 4°C. Cells were washed twice with 40 pl FACS buffer and analyzed on a BD FAC Symphony™ A5 Cell Analyzer. Binding curves and ECso values were obtained using GraphPadPrism6. The data as measured for various BCMA-CD28 antigen binding molecules and controls (for instance “2nd antibody only” refers to samples containing only the fluorophore labeled secondary antibody) are shown in Fig. 3C to 3H. The binding properties of the various BCMA-targeting molecules are summarized in Table 17A.
Both BCMA-targeted CD28v8 bispecific molecules, as well as “Alnuctamab” were able to bind to the wt and to all mutated forms of human BCMA in a concentration-dependent manner. In contrast, Elranatamab displayed a significant reduction of binding to the point mutation R27P, which translated into a weak binding signal at the highest concentrations only. Teclistamab was not able to bind to the point mutation R27P and the S30del. Efficient binding to BCMA is a crucial requisite for the therapeutic activity of BCMA-targeted bispecific molecules, suggesting an advantage for molecules who are not affected by point mutations in the ECD of BCMA. The difference in the maximal values for the wt versus the mutated BCMA transfectants is due to different expression levels of the human BCMA wt and variants on the transfectants and is consistent for all molecules tested.
Table 17A: High-level summary of binding properties of the indicated BCMA-targeting molecules to either human wildtype (wt) BCMA or the indicated point mutations in the extracellular domain of human BCMA
“Yes” means binding to the BCMA variant.
Example 4
In vitro functional characterization of bispecific CD28 agonistic antigen binding molecules targeting BCMA Several cell-based in vitro and ex vivo assays were performed to evaluate the capability of bispecific BCMA-CD28 molecules to boost T-cell bispecific-mediated activation. (Jurkat and) T-cell activation and proliferation as determined by flow cytometry were main read-outs.
1. The activity of BCMA-CD28 bispecific antibodies was assessed in a Jurkat IL2 reporter assay, where the molecules simultaneously bind to human CD28 on Jurkat IL-2 cells and BCMA expressed on the indicated MM cell lines. In presence of a CD3/TCR-mediated first activation signal, the crosslinking of the BCMA-CD28 bispecific antibodies to Jurkat and MM cells induces IL-2 release from activated Jurkat cells which in turn drives the expression of an IL-2 promoter driven luciferase gene, which can be quantified by determination of luciferase, using a conventional microplate reader. 2. In another read-out, the functionality of BCMA-CD28 bispecific antibodies was evaluated in a PBMC co-culture assay in presence of BCMA-expressing MM cell lines and a fixed concentration of an anti-GPRC5D/anti-CD3 bispecific antibody (GPRC5D TCB, P1AE6625) providing the simultaneous 1st signal for T-cell activation. Read-outs were upregulation of surface activation markers on T cells as well as T cell proliferation, as indicated.
3. A similar set-up was used to assess the functional activity of BCMA-CD28 bispecific antibodies in presence of a GPRC5D TCB in an ex vivo setting, using unprocessed MM patients bone marrow aspirate samples containing autologous T effector cells and BCMA- as well as GPRC5D-expressing MM target cells.
4.1 T cell activation - Jurkat IL2 reporter in vitro assay
Jurkat IL2 reporter assays were used to address how different BCMA-CD28 bispecific antibodies can co-stimulate T cells after activation by GPRC5D TCB (P1AE6625). The assay has been performed as described in the following:
Cell cultures: NCI-H929 cells were cultivated in RPMI1640 (Gibco) supplemented with 10% FCS (PAN-biotech), 2 mM L-glutamine (Sigma-Aldrich), 1 mM sodium pyruvate (Thermofisher), and 50pM 2-Mercaptoethanol (Thermofisher). Cells were cultured twice per week to maintain a density between 0.5 - 2.0 xlO6 cells/ml. Jurkat IL2 reporter and Jurkat NF- KB/4-1BB reporter cell lines were grown in RPMI1640 medium supplemented with 10% FCS, 25 mM HEPES (Thermofisher), 2 mM L-glutamine, 0.1 mM non-essential amino acid (Thermofisher), and 1 mM sodium pyruvate. In addition, the medium was supplemented with 200 pg/ml of Hygromycin B (Roche) for Jurkat IL2 reporter cell line, and with 400 pg/ml of Hygromycin B and 600pg/mL of Geneticin (Sigma-Aldrich) for Jurkat NF-KB/4-1BB reporter cell line. Jurkat cells were cultured twice per week to maintain a density between 0.1 - 0.5 xlO6 cells/ml.
The target (NCI-H929) and effector (Jurkat IL2 reporter) cells were harvested and resuspended in assay medium (Jurkat cell medium without antibiotics) to obtain a cell density of 6 xlO6 cells/ml for target cells and 3 xlO6 cells/ml for effector cells. Then cells were mixed at a ratio E:T 1 :2, and 20 pl of effector-target cells mixture were plated per well of a white-walled 384-flat bottom well plate (Falcon™ 384-Well White Flat-Bottom Tissue Culture Treated Microplate). Next, 10 pl of titrated amount (200.0-0.05 nM) of BCMA-CD28 bispecific antibody (BsAb) were added to the plate in triplicates. Further, 10 pl of fixed concentration (2000, 200, or 20 pM) GPRC5D-TCB was added to the plate, reaching the final volume of 40 pl per well. In addition, three separate control conditions were prepared that contained target and effector cells alone in order to indicate effector cell-induced IL2 signaling without stimulus. Target and effector cells with BCMA-CD28 BsAb and without GPRC5D-TCB were added together to address the unspecific activation of effector cells without a first signal. Last control, target, and effector cells with GPRC5D-TCB (2000, 200, or 20 pM) and without BCMA-CD28 BsAb were prepared to indicate the baseline of IL2 signaling in the effector cells signal. Assay medium was used to reach the final volume (40 pl) for the control wells. Assay plates were centrifuged for 1 min at 350 g and incubated for 24 hrs at 37°C in a humidified CO2 incubator. Assay plates were
incubated for 5 min at room temperature (RT) before adding 20 pl of the ONE-Glo solution (Promega). Further, plates were centrifuged for 1 min at 350g and incubated for 10 min at room temperature in the dark to achieve full lysis of the cells. Luminescence was measured (read: 1 s per well) by using the Tecan SparklOM. Briefly, the target NCLH929 cell line expressing high levels of BCMA was mixed at the ratio 1 :2 (E:T) with the effector Jurkat IL 2 reporter cell line that expresses luciferase in an IL2 promoter-dependent fashion. Then, titrated (200.0-0.04 nM) BCMA-CD28 BsAbs together with (2000, 200, or 20 pM GPRC5D-TCB were added and luminescence was measured 24 hrs later. Results showed that all tested BCMA-CD28 BsAbs including CD28 antibody v8 or vl5 induced co-stimulation of T cells in a dose-dependent manner (Fig. 4A-4F). In general two variants P1AG7200 and P1AG7282 for CD28 vl5 and two variants, P1AG7215 and P1AG7282, for CD28 v8 outperformed other variants and co-stimulated T cells equally well as the prior art molecule P1AE9053 (BCMA(PR)-CD28 vl5) and P1AF7062 (BCMA(PR)-CD28 v8) (Fig.4A- 4C). Data were normalized against the prior art molecule P1AE9053 in Fig.4A-4C and against P1AF7062 in Fig. 4D-4F, and the GPRC5D-TCB induced signal was substrated. Obtained data are shown as mean ± s.d. of one independent experiment performed in triplicates out of three independent experiments in Table 18 below.
Table 18: EC50 values (nM) and Emax (%) for T cell activation as measured in the Jurkat
Figures 5A to 5D show a summary of the data obtained from Jurkat NFkB reporter assay. Fig. 5 A compares the ECso values and Fig. 5B the efficacy of the various BCMA-CD28(vl5) BsAbs, and Fig. 5C the ECso values and Fig. 5D the efficacy of the various BCMA-CD28(v8) BsAbs compared to the untargeted CD28 controls Pl AG0760 and P1AF8794, respectively, and were calculated from three independent experiments performed in triplicate. ECso data of Fig. 5 A and Fig. 5C were added together from experiments with different concentrations of GPRC5D- TCB (20, 200 or 2000 pM), and the same was applied for efficacy data. All data are shown as mean ± s.d. Data were analyzed by ANOVA and Tukey’s multiple comparison correction. Asterisks indicate p< 0.05 (*), p<0.001 (***) and p<0.0001 (***).
In summary, a strong boosting of GPRC5D-TCB mediated T cells (Jurkat IL2 reporter cells) activation with all BCMA-CD28 bsAbs was observed. No significant differences in efficacy (Emax) and ECso values were observed. Overall, the efficiency (Emax) and EC50 values were in a similar range, with the exception of Pl AG7207 being significantly less potent than P1AF7062.
4.2 PBMC co-culture assay in presence of BCMA-expressing MM cell lines
To assess the ability of BCMA-CD28 bsAbs to function in a PBMC co-culture assay in presence of BCMA-expressing cells, the following method was carried out.
Target Cells: NCI-H929 & NCI-H929 BCMAko, as indicated respectively.
NCI-H929 (ATCC® CRL-9068™) is a human multiple myeloma cell line that expresses BCMA. In order to assess potential target-independent activity of the BCMA-CD28 BsAbs, a BCMA knockout variant of the NCI-H929 cell line (generated using CRISPR/Cas9 technology) was tested as well. The cells were cultivated in RPMI 1640 (Gibco™ 31870074) supplemented with 10% FCS (Gibco™ 16140-071), 10 mM HEPES (Gibco™ 15630056), 2 mM GlutaMAX-I (Gibco™ 35050-038), 1 mM sodium pyruvate (Gibco™ 11360039), and 50pM 2-Mercapto- ethanol (Gibco™ 31350010). Cells were passaged 2-3 times per week by adding fresh medium to maintain a density between 0.5xl06/ml and 2.5xl06/ml. Cells were incubated at 37 °C with 5% CO2. At the day of the assay the target cells were harvested, counted and re-suspended in assay medium (RPMI 1640 w/ HEPES, w/GlutaMax Gibco™ 31870074 plus 10% FCS) to obtain a cell density of 1.2 xlO6 cells/ml.
Effector cells: PBMCs
Frozen human peripheral blood mononuclear cells (PBMC) from whole human blood in CPD were received from Cambridge bioscience. PBMCs were stored in nitrogen vapor phase and thawed on the day of the assay. Cells were counted and then labeled with Cell Proliferation Dye eFluor™ 450 (65-0842-90, eBioscience™ ). Briefly, PBMCs were washed once with DPBS (Gibco), supernatant was discarded and cells resuspended to 2 Mio cells per well using DPBS. 10 ml of the cell suspension was added to 50ml -falcon tubes and then 10 ml of a 10 pM concentrated Cell Proliferation Dye eFluor™ 450 was added while carefully vortexing (final dye concentration was 5 pM). After an incubation of 10 min at 37°C (water bath) 30 ml of cold assay medium was added. Cells were centrifuged, counted and re-suspended in assay medium to obtain a cell density of 1.2 xlO6 PBMCs/ml.
Preparation of Co-culture
50 pl (0.06xl06) NCI-H929 or NCI-H929 BCMAko cells and 50 pl (0.06xl06 PBMCs) were plated per well in 96-well round bottom plates (TPP) resulting in a 1 : 1 ratio. Next, 50 pl of titrated amount (0.12 - 500 nM) of BCMA-CD28 bispecific antibody (BsAb) were added to the plate in triplicates. Further, 50 pl of fixed concentration (0.32 pM) GPRC5D-TCB (P1AE6625) was added to the plate, reaching the final volume of 200 pl per well. In addition, three separate control conditions were prepared. Target and effector cells with BCMA-CD28 BsAbs and without GPRC5D-TCB were added to address potential activation of effector cells by CD28 bispecific molecules without a first signal. In another control, target, and effector cells were treated with GPRC5D-TCB alone to indicate the baseline. Last, a control with target and effector cells was prepared in which no antibody was added (untreated). Assay medium was used to reach the final volume (200 pl) for the control wells. Assay plates were centrifuged for 1 min at 350 g and incubated for 4 days at 37°C in a humidified CO2 incubator.
FACS staining and read-out
After incubation, the cells were stained to evaluate T cell activation and proliferation. The cells were first washed once with 200 pL PBS and then stained with LIVE/DEAD™ Fixable Near-IR Dead Cell Stain (1 :500) (ThermoFisher Scientific Catalog No. L34976), FITC antihuman CD4 (clone RPA-T4), BV711 anti -human CD8 (clone RPA-T8), APC anti -human CD25 (clone BC96), PerCP-Cy5.5 anti-human CD 137 (clone 4B4-1), all from BioLegend in PBS for 30 min at 4°C. Flow cytometry acquisition was performed on a custom-designed BD Biosciences Fortessa and analyzed using FlowJo software (Tree Star, Ashland, OR) and GraphPad Prism software.
Results
As illustrated in Figures 6A and 6B, 7A and 7B, 8A and 8B and 9A and 9B, all BCMA- CD28 bispecific antibodies were able to significantly boost activation of CD8+ and CD4+ T cells (CD25 upregulation) as well as the proliferation of CD4+ T cells in the presence of BCMA- expressing target cells and a first signal provided by a low concentration of GPRC5D-TCB (Fig. 6A, 7A, 8A and 9A). None of the BCMA-CD28 molecules was inducing T cell activation in the absence of a first signal/TCB at the concentrations tested (500 nM and lower). Furthermore, there was no boosting of TCB-mediated activation of T cells in the absence of BCMA- expressing targets at concentrations up to 125 nM (Fig. 6B, 7B, 8B and 9B). Only the molecule BCMA (PR)-CD28 vl5 led to boosting of TCB-mediated T cell activation and proliferation in the absence of target expression, thus being able of target-independent activation as demonstrated in Fig. 8B and 9B.
Tables 19 to 22 summarize the ECso values, as well the maximum from the data shown in Figures 6A to 9A. ECso values were calculated using GraphPadPrism6.
Table 19: ECso values (nM): CD28 v8 bispecific antibodies versus BCMA (PR)-CD28 bispecific antibodies
Table 20: ECso values (nM): CD28 v!5 bispecific antibodies versus BCMA (PR)-CD28 v!5 bispecific antibody
Table 21: Maximal values: CD28 v8 bispecific antibodies versus BCMA (PR1-CD28 bispecific antibodies
D = Donor
Table 22: Maximal values: CD28 v!5 bispecific antibodies versus BCMA (PR)-CD28 v!5 bispecific antibody
D = Donor 4.3 Ex vivo functional characterization of bispecific CD28 agonistic antigen binding molecules targeting EpCAM
The therapeutic efficacy of the bispecific BCMA-CD28 antibodies was tested in combination therapy with GPRC5D-TCB in heparinized primary MM patient bone marrow aspirate (BMA) samples obtained within the first 48 hours after extraction. Cell amounts, their viability and phenotype, as well as percent of diseased MM plasma cells at baseline were analyzed by flow cytometry. Next, at least 0.1 million MM plasma cells (MM PCs) were seeded in 24- or 48-well plates. A fixed concentration of a GPRC5D-CD3 bispecific molecule (GPRC5D-TCB, 1 or 10 nM as indicated) was tested alone or in combination with increasing doses of the different BCMA-CD28 bispecific antibodies. As negative references, a TCB isotype, respectively an untargeted CD28 molecule was added. After 96 hours cells were transferred into 15 ml falcon tubes. Sterile PBS was added up to 1 mL. Cells were centrifuged at 540 g for 5 min and erythrocytes were lysed for 15 minutes according to standard methods. Cells were centrifuged at 800 g for 10 minutes, washed with sterile PBS once and incubated at room temperature for 20 minutes with maleimide to stain dead versus living cells. Cells were washed
with 12 ml of PBS, including 0.09% of NaNs and 0.5 % of BSA to remove unbound maleimide. Thereafter, cell pellets were resuspended in 500 pL PBS and filtered through 5 ml polystyrene round-bottom tubes with cell-strainer cap to discard cell clots and remove potential membrane aggregates. Cells were centrifuged once more at 540 g for 5 minutes. Finally, cells were stained with the indicated antibodies against different surface markers for 20 min at RT protected from light (Table 23 and Table 24), washed as described above and resuspended in sterile PBS. The total sample was acquired by flow.
Table 23: Antibody combination used for the analysis of pre-treatment sample
Table 24: Antibody combination used to determine MM-PC lysis and effector cell activation
In another experiment, the therapeutic efficacy of the bispecific BCMA-CD28 antibodies was tested in combination therapy with GPRC5D-TCB using frozen and thawed primary MM bone marrow mononuclear cells (BMMNCs) as described below.
On day one, samples were thawed and resuspended in StemSpan SFEMII (StemCell) medium, including 20 % human serum, 55 uM P-mercaptoethanol (Gibco cat# 11528926) and 100 U/ml penicillin and 100 pg/ml streptomycin (Gibco, lOOx stock). 3 Mio cells were plated per well of a 12-well-plate, including 100 ng/mLof recombinant human IL-6 and stored in the incubator at 37°C, 5 % CO2 for 24 hours.
Thereafter, samples were harvested and 0.1 Mio cells were used for baseline characterization. The remaining cells were washed twice with sterile PBS and stained with a NIR live/dead stain prior being sorted for viability using the FACS BD Aria sorter. Living cells were resuspended in medium and 0.1 Mio cells were plated per well of a 96-well-plate. Cells were incubated for 96 hours in the absence or presence of the indicated antibodies in a total volume of 150 uL per well at 37 °C, 5 % CO2. Samples were washed twice with sterile PBS and stained as follows: First, a live/dead staining was performed for 20 minutes at room temperature in the dark, using a NIR dye (ThermoFisher). After one washing step, human TruStain Fcx Blocking was added in FACS buffer (PBS with 2% Fetal Bovine Serum, 1% 0.5 M EDTA pH 8, 0.25% NaNs Sodium azide) and cells were incubated for another 10 minutes at room temperature (RT) in the dark prior to addition of the respective antibody mix (see Table 25).
Surface staining was performed for 20 minutes at RT in the dark. Cells were washed with FACS buffer once and fixed in PBS, containing 2 % paraformaldehyde (PF A) for 10 minutes at RT in the dark. Samples were washed with FACS buffer, resuspended in FACS buffer, and analyzed by flow cytometry after addition of counting beads to determine the absolute number of cells.
Table 25: Antibody combination used to do the baseline characterization and effector cell activation
The FACS surface antibodies were purchased from BD Biosciences, Miltenyi or LuBiosciences and used according to the manufacturers’ recommendations. The detection antibody for human GPRC5D was generated by Roche. Results: Fig. 10 and Table 26 demonstrate that all BCMA-CD28 bispecific antibodies were able to significantly boost activation of CD8+ T cells (CD25 upregulation) on top of the GPRC5D-targeted CD3 engager (GPRC5D-TCB) mediated activation. The strongest additional activation at 200 nM was induced by BCMA-CD28 bsAb P1AG7215, followed by BCMA- CD28 Pl AG7191 and BCMA-CD28 Pl AG7207. No activation on top of the TCB was induced in presence of 200 nM of the untargeted CD28, which demonstrates that the CD28 molecules need to be crosslinked via the tumor antigen-targeting moiety to be active.
Table 26: Up-regulation of the activation marker CD25 on CD8+ T cells upon incubation of primary MM BM aspirate samples with the indicated molecules for 96 hours, as assessed by flow cytometry
Samples were normalized to wells treated with the T cell bispecific (TCB) isotype control (TCB isotype=0).
This could be demonstrated as well with another primary MM patient BM aspirate sample, as depicted in Fig. 11 and Table 27. Here, boosting of TCB-mediated lysis of primary malignant MM plasma cells could be observed in presence of 400 nM of the BCMA-targeted molecule Pl AG7191 but not in presence of the untargeted CD28. In presence of 400 nM BCMA (PR)- CD28 v8 only a minor add-on effect on top of the TCB-mediated lysis of primary MM plasma cells could be observed, indicating that the ex vivo efficacy of the BCMA-CD28 bispecific antibody 7191 is stronger compared to the one of BCMA (PR)-CD28 v8. Table 27: Induction of MM plasma cell lysis (MM PC) upon incubation of primary MM BM aspirate samples with the indicated molecules for 96 hours, as assessed by flow cytometry
Samples were normalized to weels with untreated BM aspirate samples (untreated = 0).
Figures 12A to 12D and Table 28 depict the BCMA-target dependent upregulation of T cell activation (CD25 on either CD4+ or CD8+ T cells), as well as the increase of the
degranulation marker CD 107 on CD4+ and CD8+ T cells on top of the TCB-mediated activation/degranulation in presence of the indicated BCMA-CD28 bispecific antibodies but not in presence of the untargeted negative reference. At 200 nM, the BCMA-CD28 bispecific antibody P1AG7215 induced a similar effect on activation of T cells as BCMA (PR)-CD28 v8 and BCMA (PR)-CD28 vl5, whereas BCMA-CD28 P1AG7282 induces slightly more moderate effects on top of the TCB-mediated ones. Furthermore, the experiment reveals similar effects induced by 200 nM BCMA-CD28 bispecific antibodies, which solely differ in their CD28 variant: CD28v8 (7062) or CD28vl5 (9053). Levels of CD107 on CD4+ T cells indicate strongest effects on top of TCB induced by BCMA (PR)-CD28 v8 and BCMA-CD28 P1AG7215, followed by BCMA-CD28 Pl AG7282. On CD8+ T cells, only BCMA-CD28 Pl AG7282 reveals increased levels of CD107 on top of the one induced by TCB. Differences observed between the effect on T cell activation (CD25) and degranulation (CD 107) might be based on different kinetics of those markers. Overall, the data shows that all assessed BCMA-CD28 bispecific antibodies were able to further boost T cell activation respectively degranulation on top of a TCB providing the first signal to T cells.
Table 28: Up-regulation of the activation marker CD25 or the degranulation marker CD107 on CD4+ or CD8+ T cells upon incubation of primary MM BM aspirate samples with the indicated molecules for 96 hours, as assessed by flow cytometry
Another series of experiments was conducted using BM MNCs from primary MM patients after thawing. As indicated in Figures 13A and 13B and Table 29, BCMA-CD28 Pl AG7191 induced significant lysis of malignant MM plasma cells as well as upregulation of the frequency of CD25-positive CD8+ T cells on top of GPRC5D TCB in all three samples assessed. The maximal effect induced by the TCB, as well as by the combination of GPRC5D TCB and BCMA-CD28 bispecific antibody differed between the MM patient samples, suggesting that the immune cell composition, the effector to target cell ratio and T cell fitness may impact the overall results.
Table 29: Increased lysis of primary malignant MM plasma cells, respectively upregulation of the activation marker CD25 on CD8+ T cells upon incubation of primary MM BM MNCs with the indicated molecules for 96 hours, as assessed by flow cytometry
Samples were normalized to wells with untreated BM MNCs (untreated=l).
Example 5
In vivo functional characterization of bispecific CD28 agonistic antigen binding molecules targeting BCMA
The efficacy studies described herein were aimed to understand the potency of the BCMA- CD28 bispecific antibodies in combination with an anti-GPRC5D/anti-CD3 bispecific antibody (GPRC5D x CD3).
Cell culture: Human NCI-H929 cells (obtained from Roche Nutley) were cultured in RPMI1640 high glucose medium containing 10% FCS, 2 mM L-Glutamine, 10 mM HEPES and 1 mM Sodiumpyruvate (37 °C at 5 % CO2). In order to generate tumor bearing mice, 50 microliters cell suspension (2.5 xlO6 cells) were co-injected with 50 pl Matrigel subcutaneously in the right flank of anaesthetized, humanized NSG mice.
Mouse model: Humanized NSG mice were provided by Jackson Laboratories, Sacramento USA. For engraftment, animals were irradiated (140cGy) and injected with CD34+ cord blood cells (9xl04 cells) from healthy donors (hematopoietic stem cells; HSC). After arrival animals were maintained for one week to get accustomed to new environment and for observation. Mice were maintained under specific pathogen-free conditions with daily cycles of 12 h light /12 h darkness according to committed guidelines (GV-Solas; Felasa; TierschG). Continuous health monitoring was carried out on a regular basis. Experimental study protocol was reviewed and approved by the local government (ROB-55.2-2532. Vet_03-20-170).
Randomization and treatment: When subcutaneous tumors reached a tumor volume of 200 mm3, humanized mice were randomized into different treatment groups based on tumor volume and body weight (n=10/group). All antibodies and vehicle (Histidin buffer) were administered intravenously (i.v.) once weekly. Costimulator molecules were injected 48 hours (Fig. 14A to 14F) or 24 hours (Fig. 15A to 15L, Fig. 16A to 16G) after GPRC5D x CD3 treatment.
Monitoring of animals: Animals were controlled daily for clinical symptoms and detection of adverse effects. Body weight and tumor growth (Caliper measurement) was monitored twice weekly. Animals were sacrificed according to the termination criteria or at the end of the experiment.
Statistics: Graphs were generated using GraphPad prism Software. A time-to-event analysis was performed using the internal tool DOPSa (based on the software R). The critical tumor volume was chosen as the maximum observed baseline tumor volume plus the standard deviation of the baseline tumor volume (rounded to the nearest multiple of ten). Study groups were compared using the Log-Rank test and p-values have been corrected for multiple testing using the Bonferroni-Holm method. Significant changes compared to the control group (GPRC5D x CD3) with p<0.05 were depicted with asterisks (* p<0.05, ** p<0.01, *** p<0.001).
Combination of GPRC5D x CD3 with BCMA-CD28
In order to compare GPRC5D x CD3 monotherapy to combination with BCMA-CD28 bispecific antibody, NCI-H929 tumors were established in humanized NSG mice and efficacy was evaluated in vivo. The results are shown in Figures 14A to 14F. In summary, GPRC5D x CD3 monotherapy (1 mg/kg P1AE3357) induced a moderate and heterogeneous tumor growth inhibition (Fig 14B). Although there was no statistical significant difference observed in tumor growth inhibition, combination of GPRC5D x CD3 with P1AG7215 and especially P1AG7282 (10 mg/kg; 48 hours after GPRC5D x CD3) seemed to result in a more homogeneous and overall stronger response than the monotherapy (Fig. 14C and 14D).
In a second in vivo study different doses of BCMA-CD28 bispecific antibodies were tested for their costimulatory efficacy (Pl AG7215: 20 mg/kg, 10 mg/kg, 5 mg/kg and Pl AG7200: 20 mg/kg, 10 mg/kg, 2 mg/kg; 48 hours after GPRC5D x CD3 (P1AE3357) and compared to BCMA (PR)-CD28 bispecific antibodies (P1AE9053: 10 mg/kg and P1AF7062 10 mg/kg; 48 hours after GPRC5D x CD3) in combination with GPRC5D x CD3 therapy. The results are shown in Figures 15A to 15L. Combinations of P1AG7215 and P1AG7200 were found to act synergistically with GPRC5D x CD3 and improve tumor growth inhibition in the NCI-H929 model (Fig. 15E to 15 J). The in vivo efficacy of the assessed new BCMA-CD28 bispecific antibodies was in a similar range as compared to the BCMA-CD28 bispecific antibodies comprising prior art BCMA antibody PR. The treatment groups of P1AG7215 (20 mg/kg, see Fig. 15E) and P1AG7200 (20 mg/kg, see Fig. 15H) showed lowest tumor volumes at termination.
In a third in vivo study, high doses of BCMA-CD28 costimulators Pl AG7207 and P1AG7191 were tested (20 and 30 mg/kg P1AG7207 and 20 mg/kg P1AG7191; 24 hours scheduling after GPRC5D x CD3 P1AE3357) with a number of 10 animals per treatment group. Both molecules induced significant tumor growth inhibition compared to GPRC5D x CD3 monotherapy (Fig 16A to 16G, Table 31). The higher dose of Pl AG7207 induced a larger number of responders compared to the 20 mg/kg combinations (n=5 compared to n=4 or n=2, respectively) and overall smaller tumor volumes at termination (Table 30).
Table 30: Tumor volume (mm3) at termination
Table 31: P-values of combination groups compared to GPRC5D x CD3
In a fourth in vivo study, Pl AG7191 was tested in different doses (40, 10 and 1 mg/kg) in combination with GPRC5D x CD3 (0.05 mg/kg Pl AE6625) and compared to the GPRC5D x CD3 monotherapy and combination with the molecule P1AE9053 comprising BCMA clone PR (10 mg/kg). P1AG7191 and P1AE9053 were injected 24 hours after GPRC5D x CD3. The GPRC5D x CD3 monotherapy induces strong tumor growth inhibition in the NCI-H929 model (Fig. 17A). However, tumors start to relapse after the 4th treatment cycle (Table 31). The two highest doses of Pl AG7191 (40 and 10 mg/kg) delay the time to tumor relapse and inhibit the tumor regrowth. At termination of the experiment (day 64), 6 and 8 responders were left in the P1AG7191 40 mg/kg and 10 mg/kg combination groups, respectively. In the GPRC5D x CD3 monotherapy group and in the combination with the 1 mg/kg Pl AG7191 group only 2 responders were left. The molecule P1AE9053 shows lower combination effects compared to Pl AG7191 as only 5 responders were left at termination and more tumors escape at earlier time points (Table 32).
Taken together, the results obtained from these in vivo studies provide preclinical evidence that the therapeutic response to GPRC5D x CD3 can be boosted by BCMA-CD28 bispecific antibodies.
Example 6
Pharmacological Properties of bispecific antigen binding molecules targeting BCMA and CD28
6.1 PK properties in huFcRn transgenic mice
Transgenic mice carrying the human FcRn instead of the mouse FcRn are considered more predictive for the clearance in human than in wild-type mice (C57/B16). Female huFcRn Tg32 homozygous SCID mice (n = 3) received a single IV bolus dose of Pl AG7207 at 5 mg/kg. Female huFcRn Tg32 homozygous BL6 mice (n = 3) received a single IV bolus dose of P1AG7282, P1AG7191, or P1AG7215, respectively, at 5 mg/kg. Blood samples were taken at 0.17, 7, 24, 48, 72, 168, 336, 408, and 504 hours after receiving the dose and processed to serum prior to bioanalysis. Serum samples were analyzed using an immunoassay with anti-human FCpan (CH2) or kCH capture and detection reagents. As shown in Figure 18 and Table 33, all tested BCMA-CD28 bispecific antibodies have a similar plasma concentration-time profiles and clearance values ranging between 4 to 7 mL/kg/day in these mouse strains.
The Cell Microarray Technology (Charles River Laboratories) was used to screen for potential off-target binding interactions. Bispecific IgGl antibodies were screened for binding against fixed HEK293 cells expressing 6019 individual full-length human plasma membrane proteins and cell surface-tethered human secreted proteins, as well as a further 397 human heterodimers. Fluorescent images of the spotted cells were analyzed using ImageQuant software (GE healthcare, Version 8.2).
Table 34 reports the results of the screen as strong, medium, weak or no interaction after visual inspection of the spots intensity.
As a conclusion, all antibodies showed a strong interaction with their primary targets, i.e. BCMA and CD28. P1AE9053 showed an additional interaction with REN (renin). Pl AG7191 and P1AG7215 showed very weak or medium interactions with LDLR (low density lipoprotein receptor). No other interactions were detected for P1AG7215 and P1AG7282, indicating a high specificity of these molecules for their primary targets.
6.3 Immunogenicity Testing
6.3.1 MAPPs Assay
The MAPPs assay was used to identify potential T cell epitopes from four BCMA-CD28 bispecific antibodies: P1AG7191, P1AG7215, P1AG7207 and P1AG7282 (Figure 19). Major histocompatibility complex-II (MHC-II)-Associated Peptide Proteomics (MAPPs) is a mass spectrometry-based approach to identify and relatively quantitate naturally processed and presented MHC-II-associated peptides that can potentially activate T cells and contribute to the immunogenicity of a drug.
Generation of Human Monocyte-Derived DCs and the Assay Procedure: Peripheral blood mononuclear cells (PBMCs) were isolated from huffy coats of healthy donors by gradient density centrifugation using Ficoll-Paque PLUS (GE Healthcare Europe GmbH, Glattbrugg, Switzerland). Monocytes were isolated by positive immunoselection using anti-CD14-coated microbeads and a magnetic separator (MACS, Miltenyi Biotech, Bergisch Gladbach, Germany). CD14+ cells were then cultured at a concentration of 0.3 x 106 cells/mL in 100mm ultra-low attachment culture dishes (Coming Inc., Corning, NY, USA) in serum-free Cellgro medium containing 1% GlutaMAX, 1% penicillin/streptomycin. Monocytes were differentiated into immature DCs with 50 ng/mL of GM-CSF and 5 ng/mL of IL-4 for 5 days at 37 °C with 5% CO2 before being challenged with a test protein at 50 pg/mL in the presence of 1 pg/mL of lipopolysaccharide (LPS) from Salmonella abortus equi (Sigma-Aldrich Chemie GmbH, Buchs, Switzerland) for 24 h. Mature DCs were harvested, washed with phosphate-buffered saline (PBS) and the cell pellets were frozen at -80 °C prior to subsequent immunoprecipitation.
Isolation of HLA-DR-Presented Peptides: Cell pellets were lysed in 20 mM Tris-buffer solution pH 7.8 containing 1% (v/v) Triton X-100 and protease inhibitors (Roche Diagnostics GmbH, Mannheim, Germany) for 1 h at 4 °C on a ThermoMixer at 1100 rpm. The HLA-DR immune complexes were isolated by immunoprecipitation using biotin-conjugated anti-human HLA-DR antibodies (clone L243, RayBiotech). Lysates were incubated with the antibody on a rotator overnight at 4°C. Samples were washed five times with a buffer containing 20 mM N-(2- Hydroxyethyl) piperazine-A’-2-ethane sulfonic acid-NaOH (pH 7.9), 150 mM KC1, 1 mM MgCh, 0.2 mM CaCh, 0.2 mM ethylenediaminetetraacetate, 10% (v/v) glycerol, and 0.1% (v/v) Digitonin and five times with purified water. MHC-II peptides were eluted twice from HLA-DR molecules by adding 18 pL of 0.1% trifluoroacetic acid. The eluates were collected and kept at 4°C prior to mass spectrometric analysis.
Data acquisition: Data acquisition and data processing were performed essentially as described in Steiner et al.. J. Proteome Res. 2020, 19, 3792-3806. MHC-II peptide preparations obtained from about 3 x 106 human monocyte-derived DCs (moDCs) per sample were separated on a nanocapillary liquid chromatography system (UltiMate 3000 RSLC, Thermo Scientific, CA, USA) using self-packed fused-silica C18 reversed phase column (75 pm i.d. x 170 mm, ReproSil-Pur C18-AQ, 3 pm, Dr. Maisch GmbH) connected to a Q-Exactive HFX Orbitrap mass spectrometer (Thermo Scientific) via electrospray ionisation (LC-ESI-MS/MS). Samples (15 pL volume dissolved in 0.5% (v/v) formic acid in 2% (v/v) acetonitrile/water) were loaded for 2-3 min at 8 pL/min onto an Acclaim PepMap C18 trap column (100 pm i.d. x 20 mm, Thermo Scientific) using a Vented Tee design. Peptides were then eluted at a flow rate of 250 nL/min using a nonlinear 39 min gradient of 2-45% B, followed by an 11 min column wash, and reequilibration for 10 min [buffer A: 0.1% (v/v) formic acid in 2% (v/v) acetonitrile/water; buffer B: 0.1% (v/v) formic acid in acetonitrile], MHC-II peptides were analysed by tandem MS using
standard operating parameters. Survey scans (scanning range m/z 400-1650) were recorded in the Orbitrap mass analyzer at a resolution of 60,000, with the lock mass option enabled. Data- dependent MS/MS spectra of the 18 most abundant ions from the survey scan were recorded in the Orbitrap cell at a resolution of 15,000. Target ions selected for MS/MS were dynamically excluded for 7 s. Peptides were identified using the most updated PEAKS Studio version available at the time (version X Pro, Bioinformatics Solutions Inc., ON, Canada). The raw MS data were searched against the human protein database UniProtKB (http ://www.uniprot. org, release 2015 10, approx. 88,500 TrEMBL and SwissProt entries containing the amino acid sequences of the test therapeutic proteins) with a mass tolerance of ±10 ppm for precursor ions and ±0.025 Da for fragment ions. Met-sulfoxide, Asn/Gln deamidation, and N-terminal pyroglutamyl ati on were considered as differential modifications. Data were searched without enzyme specificity, and peptide results were reported at 1% specFDR cutoff. All LC-MS/MS runs of a given donor were processed batchwise, and the areas under the curve of identified features (2 min retention time shift tolerance; features reporting option: all) were exported in a tab-delimited table without further normalisation.
Statistical analysis: DataMAPPs is a Roche-created data analysis tool to help visualising otherwise complex mass spectrometry-derived data in the form of a heat map. The program is publicly available at https://www.R-project.org/ and can be executed with a standard R installation. Please see reference (Steiner et al. 2020) for an extensive description of the package. DataMAPPs is invoked after processing the LC-MS/MS data with the PEAKS software and exporting results in a suitable tabular format. In short, the dataMAPPs processing pipeline consists of (1) data import and consistency check: sample annotation and peptide quantification (PEAKS output) files are read into memory and checked for consistency. Only peptides of length 10-30 amino acids are considered for analysis and mapped against the sequences of the studied biologies. Further filtering is performed to retain only the antibody-related signals that are specific to the studied molecules in a given LC-MS/MS analysis. A final step eliminates duplicate entries and sums signals from peptides identified with different charge states Peak area is transformed to the log2 scale prior to processing; (2) data QC: the pipeline flags samples with a low number of identified peptides (cutoff typically set to half of the median peptide count for the respective donor) or low similarity (median Pearson correlation with all other samples from the same donor <0.8). Cutoffs can be modified by the user after inspecting a collection of QC plots, which allows for a suitable, data set-specific processing while being able to rerun the procedure in a reproducible way; (3) data normalisation: peptide abundance (based on peak area) is normalised based on an adjusted version of the GRSN (global rank-invariant set normalisation) procedure. (4) replicate aggregation: technical replicates (i.e. samples run for the same donor, same treatment, same dose) can be averaged to retain information on all peptides that were detected in at least one of the replicates; (5) peptide mapping: antibody-associated peptides are
mapped on the tested biologic’s protein amino acid sequences. In addition, peptides with neighbouring amino acid positions are binned to distinct epitope clusters (hotspots), whose abundance reflects the summed intensities of each of the constitutive MHC-II peptides; (6) data export and visualisation: the dataMAPPs workflow contains standard functionalities to generate several heatmap plots (epitope cluster or individual peptide level, per antibody or a global experiment summary) for a quick visualisation and comparison of the results
Results: The analysis of BCMA-CD28 bispecific antibody P1AG7191 revealed three binding clusters. Among them, one cluster (C5) contained non-germline residues. Though the frequency for this cluster appears low (1/12 donors). In total 1/12 donors (8.3%) presented a potential immunogenic T cell epitope that could contribute to the immunogenicity of the compound.
The analysis of Pl AG7215 revealed one binding cluster which did not contain any nongermline residues. As a consequence, none of the donors presented a potential immunogenic T cell epitope that could contribute to the immunogenicity of the compound.
The analysis of P1AG7207 revealed three binding clusters. Among them, two clusters (Cl and C2) contained non-germline residues with frequencies of 2/12 and 5/12, respectively. In total 5/12 donors (41.6%) presented a potential immunogenic T cell epitope that could contribute to the immunogenicity of the compound.
The analysis of Pl AG7282 revealed three binding clusters. Among them, one cluster (Cl) contained non-germline residues with a frequency of 1/12 donors. In total 1/12 donors (8.3%) presented a potential immunogenic T cell epitope that could contribute to the immunogenicity of the compound..
6.3.2 DC:T cell assay (Epibase® DC:CD4 re-stimulation assay developed by Lonza)
T cell activation is an important part of the immune response to therapeutic proteins and is usually required for the development of anti -drug antibodies in the clinic. The DC-T cell assay was used to assess the capacity of four BCMA-CD28 bispecific antibodies: Pl AG7191, P1AG215, P1AG7207, P1AG7282, to induce CD4+ T cells upon presentation of T cell epitopes by APCs.
Materials: Donors were recruited at Phase I clinical trial units in the UK. All samples were collected under an ethical protocol approved by a local REC (research ethics committee) and written informed consent was obtained from each donor prior to sample donation. All samples were stored according to the terms of Lonza’s HTA (Human Tissue Authority) license for the use of samples in research. PBMCs from healthy donors were prepared from whole blood within
six hours of blood withdrawal. Cells were cryopreserved in vapour phase nitrogen until use in the assays. The quality and functionality of each PBMC preparation was analyzed by seven days of activation with positive controls such as KLH to assess naive T cell responses.
Keyhole limpet haemocyanin (KLH) was used as a technical control, reconstituted and stored at -80°C in single use aliquots according to the manufacturer’s recommendations under sterile conditions. Additionally, bevacizumab (Avastin®) was included as a positive benchmark protein. All samples were tested at a final concentration of 0.3 pM for the DC stimulation stage and for the APC re-stimulation stage.
Methods: Monocytes were isolated from frozen PBMC samples by magnetic bead selection and differentiated into immature DC (iDC) using GM-CSF and IL-4. iDC were then harvested, washed and loaded with each individual test protein/peptide for 4 hours at 37°C. A DC maturation cocktail containing TNFa and IL-ip was then added for a further 40-42 hours to activate/mature the DC (mDC). The expression of key DC surface markers (CD11c, CD14, CD40, CD80, CD83, CD86, CD209 and HLA-DR) at both the immature and mature stage were assessed by flow cytometry to ensure the DC were activated prior to T cell interaction. The mDCs were then co-cultured with autologous CD4+ T cells (isolated by magnetic bead selection) for 6 days at 37°C, 5% CO2 in a humidified atmosphere. On day 6, autologous monocytes were isolated from PBMC using magnetic bead selection and loaded with the selected protein/peptide that were initially used to load the DC. After incubation at 37°C, 5% CO2 in a humidified atmosphere for 4 hours, the monocytes were added to anti-IFNy pre-coated FluoroSpot plates (Mabtech) along with the corresponding DC:CD4 co-culture in quadruplicate. The FluoroSpot plates were incubated for 40-42 hours at 37°C, 5% CO2 in a humidified atmosphere. After incubation the FluoroSpot plates were developed using an in-house procedure and the spotforming cells (SFC) per well assessed for each cytokine in each test condition.
Surface marker QC checks were performed on the monocyte derived DC at both the immature and mature stage to determine any possible influence of the test product on the DC differentiation and allows for the assessment of the quality of the DCs before subsequent coculture with CD4+ T cells. Surface markers are assessed by flow cytometry using fluorescently labelled antibodies and the Guava® easyCyte™ 8HT flow cytometer.
Data analysis: Data management and statistical analysis has been performed in the R programming language (https://www.R-project.org/, v. 3.6.1). Data are transformed to a log2 scale and a Generalized linear model (GLM) is applied to quantify SI (fold change and 95% CI). Adjustments are applied to the dataset (exponential type of heteroscedasticity adjusted, gaussian noise injection at low end of SFU scale, linear regression and extrapolation of each SI to a blank
value of 0) and QC plot are generated (DC differentiation markers, reproducibility on compound and donor level, relative stimulation of donors).
To help for the immunogenicity risk assessment the Stimulation Index (SI) is calculated for each test condition in each donor (ratio between SFU/well and the matched blank). A positive donor response is counted if at least 2-fold SI change is established with p<0.05 (non-adjusted p- value from GLM). The number of positive donor responses to a treatment within the 30 healthy donor cohort gives the response rate relative to this treatment.
Results: The DC used in this assay were of high quality and expressed high levels of the T cell co-stimulatory molecules required for the activation of T cells. Based on the readouts of the DC-T cell assay, all four BCMA-CD28 bispecific antibodies P1AG7191, P1AG215, P1AG7207 and Pl AG7282 were associated with a low risk of sequence-related immunogenicity for CD4+ T cell response (below the threshold of 10% as described in Siegel et al, Pharmaceutics 2022, 14(12), 2672).
6.3.3 Integrated Risk for Immunogenicity
Administration of therapeutic antibodies may cause the formation of anti-drug antibodies (AD As), which may negatively impact the safety of the therapeutic result (e.g., allergic reactions, immune complex -mediated diseases). The risk for the four BCMA-CD28 bispecific antibodies P1AF7191, P1AF215, P1AF7207 and P1AF7282 to induce an unwanted immune response and the consequences for such a response were assessed through a combination of non-clinical assays (MAPPs and DC-T cell assay) as descibed herein before. Overall, the BCMA-CD28 bispecific antibodies can be ranked as follows, from the lowest risk of immunogenicity to the highest risk: P1AG215, P1AG7191, P1AG7282 and PlAG7207.
6.4 Evaluation for the risk of cytokine release in an in vitro human Whole blood assay
In order to evaluate potential safety risks relating to cytokine release upon first dosing in humans, in vitro non-GLP testing for cytokine secretion mediated by BCMA-CD28 bispecific antibodies was conducted with fresh undiluted human whole blood samples from healthy donors. Erbitux®, an anti-EGFR IgGl mAb, was used as a negative comparator and Lemtrada®, a humanized anti-CD52 IgGl, known to induce first infusion reactions (IRRs) in greater than 90% of recipients, was used as a positive comparator.
Venous blood from 6 healthy donors was collected in vacutainer tubes with lithium heparin as anticoagulant (Roche Infirmary Services, Switzerland and Roche) and kept at room temperature until initiation of the assay, which was within 3 hrs of blood withdrawal. 195 pl of whole blood was added to the test items (5 pL volume, in triplicate) in round bottom 96 well
plates followed by incubation for 24 hrs at 37°C with 5% CO2. Cells and plasma were separated by centrifugation at 1800 g for 5 min and plasma samples (70 pL volume) were stored at -80°C until analysis of cytokine content.
All compounds were tested at concentrations of 0.1, 1, 10 and 100 pg/mL. In addition, PBS (Dulbecco's phosphate-buffered saline, Gibco No.14190, technical negative control) and LPS at a concentration of 0.1 pg/ml (lipopolysaccharide derived from Salmonella abortus, Sigma, Product. No. L5886, technical positive control) were also included in the assay.
Cytokine analysis: Determination of cytokine concentrations was performed on frozen plasma samples diluted 1 :5. Pre-tests revealed that levels of cytokine did not differ between fresh and thawed samples. Analyte concentrations were determined by ELISA using the Human Cytokine chemiluminescent assay kit (Aushon Ciraplex, Cat. No 101-269- 1-AB) with the SignaturePLUS™ imaging system and the Cirasoft analysis software. Results are expressed as pg/ml. Values above ULOQ concentrations were assigned the concentration of the highest standard as follows: IFN-y, 500 pg/mL ; IL-2, 200 pg/mL ; IL-6, 2000 pg/mL ; IL-8, 4000 pg/mL ; TNF-a, 1000 pg/ml. Sample values below LLOQ concentrations were assigned LLOQ concentrations : IFN-y, 0.24 pg/mL ; IL-2, 98 fg/mL ; IL-6, 0.98 pg/mL ; IL-8, 1.95 pg/mL ; TNF-a, 0.49 pg/mL.
Statistical analysis was performed with JMP 12.1.10 software (SAS Institute AG) using a custom-made script (Sabine Wilson, PS BiOmics/Pathology) for data import, descriptive analysis, replicate and sample outlier removal, cut-point determination and visualization of results (frequency and ratio graphs). PBS data were used to determine the population distribution, the technical background, decision limit (Lc) for each cytokine and identification of statistical outliers. The Lc represents the cytokine signal for which only a 5% chance is expected that an actually negative (PBS like) sample exceeds this value. The Lc values were determined based on PBS samples from healthy donors. In other words, the chance of a true blank to exhibit a signal > Lc is 5%. In order to compare the test items to the clinically well-characterized control Erbitux®, a cut-off was calculated for each concentration and cytokine based on the distribution of Erbitux®-induced values. This was done by a 95% quantile approach of the measured Erbitux® values. Thus, there is a 5% chance that values from ‘negative’ samples exceed these cutoffs.
Whereas the BCMA-CD28 bispecific antibodies P1AG7215 and P1AG7062 did not trigger any cytokine release that was above the level of that mediated by Erbitux®, the molecule P1AE9053 (BCMA (PR)- CD28 vl5) triggered a release of IL-6 and IL-8 in 50 % of the donors.
***
Claims
1. An antibody that specifically binds to B cell maturation agent (BCMA), wherein the antibody comprises a first antigen binding domain comprising
(i) a heavy chain variable region (VH BCMA) comprising a heavy chain complementary determining region CDR-H1 of SEQ ID NO: 1 (GYTFTNYWMH), a CDR-H2 of SEQ ID NO: 2 (IIHPNSGSTNYNEKFQG), and a CDR-H3 of SEQ ID NO: 3 (GIYDYPFAY), and
(ii) a light chain variable region (VLBCMA) selected from the group consisting of
(a) a VL comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 4 (RASES VSIHGTHLMH), a CDR-L2 of SEQ ID NO: 5 (AASSLQS) and a CDR-L3 of SEQ ID NO: 6 (QQSIEDPYT); or
(b) a VL comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 4 (RASES VSIHGTHLMH), a CDR-L2 of SEQ ID NO: 7 (AASNLES) and a CDR-L3 of SEQ ID NO: 6 (QQSIEDPYT), or
(c) a VL comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 4 (RASES VSIHGTHLMH), a CDR-L2 of SEQ ID NO: 8 (AASNLQS) and a CDR-L3 of SEQ ID NO: 6 (QQSIEDPYT).
2. The antibody of claim 1, wherein the VH BCMA comprises an amino acid sequence selected from the group consisting of SEQ ID NO: 9 (VHla) and SEQ ID NO: 10 (VHlb), and/or the VL BCMA comprises an amino acid sequence selected from the group consisting of SEQ ID NO: 11 (VLlf) , SEQ ID NO: 12 (VLla), SEQ ID NO: 13 (VLlb), SEQ ID NO: 14 (VLlc), SEQ ID NO: 15 (VLld), and SEQ ID NO: 16 (VLle).
3. An antibody that specifically binds to BCMA, wherein the antibody comprises a first antigen binding domain comprising
(a) a VH BCMA comprising an amino acid sequence of SEQ ID NOV and VLBCMA comprising an amino acid sequence of SEQ ID NO: 11, or
(b) a VH BCMA comprising an amino acid sequence of SEQ ID NOV and VLBCMA comprising an amino acid sequence of SEQ ID NO: 12.
4. The antibody of any one of claims 1 to 3, wherein the first antigen binding domain is a Fab molecule.
5. The antibody of any one of claims 1 to 4, comprising an Fc domain composed of a first and a second subunit.
6. The antibody of any one of claims 1 to 5, comprising a second antigen binding domain that specifically binds to a second antigen.
7. The antibody of any one of claims 1 to 6, wherein the second antigen binding domain that specifically binds to a second antigen is a Fab molecule wherein the variable domains VL and VH or the constant domains CL and CHI, particularly the variable domains VL and VH, of the Fab light chain and the Fab heavy chain are replaced by each other.
8. The antibody of any one of claims 6 or 7, wherein the second antigen is CD28.
9. The antibody of claim 8, wherein the second antigen binding domain that specifically binds to CD28 comprises a heavy chain variable region (VHCD28) comprising a heavy chain complementary determining region CDR-H1 of SEQ ID NO: 17, a CDR-H2 of SEQ ID NO: 18, and a CDR-H3 of SEQ ID NO: 19, and a light chain variable region (VLCD28) comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 20, a CDR-L2 of SEQ ID NO: 21 and a CDR-L3 of SEQ ID NO: 22.
10. The antibody of claims 8 or 9, wherein the second antigen binding domain that specifically binds to CD28 comprises a heavy chain variable region (VHCD28) comprising an amino acid sequence of SEQ ID NO:23, and a light chain variable region (VLCD28) comprising an amino acid sequence of SEQ ID NO:24 (v8).
11. The antibody of any one of claims 1 to 10, wherein the antibody comprises a first antigen binding domain comprising a VH BCMA comprising an amino acid sequence of SEQ ID NOV and VL BCMA comprising an amino acid sequence of SEQ ID NO: 11 and a second antigen binding domain comprising VH CD28 comprising an amino acid sequence of SEQ ID NO:23 and VL CD28 comprising an amino acid sequence of SEQ ID NO:24.
12. The antibody of any one of claims 1 to 11, comprising a first light chain comprising the amino acid sequence of SEQ ID NO:25, a first heavy chain comprising the amino acid sequence of SEQ ID NO:26, a second heavy chain comprising the amino acid sequence of SEQ ID NO:27 and a second light chain comprising the amino acid sequence of SEQ ID NO:28.
13. An antibody that specifically binds to B cell maturation agent (BCMA) and CD28, wherein the antibody comprises
(A) a first antigen binding domain comprising
(i) a heavy chain variable region (VH BCMA) selected from the group consisting of
(a) a VH comprising a heavy chain complementary determining region CDR-H1 of SEQ ID NO: 29 (GFTFSNAWMD), a CDR-H2 of SEQ ID NO: 30 (QITAKSNNYATYYADSVKG), and a CDR-H3 of SEQ ID NO: 31 (DGYH), and
(b) a VH comprising a heavy chain complementary determining region CDR-H1 of SEQ ID NO: 29 (GFTFSNAWMD), a CDR-H2 of SEQ ID NO: 32 (QITAKSNNYATYYAAPVKG), and a CDR-H3 of SEQ ID NO: 31 (DGYH), and
(ii) a light chain variable region (VLBCMA) comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 33 (RASEDIRNGLA), a CDR-L2 of SEQ ID NO: 34 (NANSLHT) and a CDR-L3 of SEQ ID NO: 35 (EDTSKYPYT), and
(B) a second antigen binding domain that specifically binds to CD28.
14. The antibody of claim 13, wherein the VH BCMA comprises an amino acid sequence selected from the group consisting of SEQ ID NO: 36 (VH2a) and SEQ ID NO: 38 (VHlb), and/or the VL BCMA comprises an amino acid sequence selected from the group consisting of SEQ ID NO: 37 (VL2a) and SEQ ID NO:39 (VLla).
15. An antibody that specifically binds to BCMA and CD28, wherein the antibody comprises a first antigen binding domain comprising
(a) a VH BCMA comprising an amino acid sequence of SEQ ID NO:36 and a VLBCMA comprising an amino acid sequence of SEQ ID NO:37, or
(b) a VH BCMA comprising an amino acid sequence of SEQ ID NO:38 and a VLBCMA comprising an amino acid sequence of SEQ ID NO:39.
16. The antibody of any one of claims 13 to 15, wherein the first antigen binding domain is a Fab molecule.
17. The antibody of any one of claims 13 to 16, wherein the second antigen binding domain that specifically binds to a second antigen is a Fab molecule wherein the variable domains VL and VH or the constant domains CL and CHI, particularly the variable domains VL and VH, of the Fab light chain and the Fab heavy chain are replaced by each other.
18. The antibody of any one of claims 13 to 17, wherein the second antigen binding domain that specifically binds to CD28 comprises a heavy chain variable region (VHCD28) comprising a heavy chain complementary determining region CDR-H1 of SEQ ID NO: 17, a CDR-H2 of SEQ ID NO: 18, and a CDR-H3 of SEQ ID NO: 19, and a light chain variable region (VLCD28) comprising a light chain complementary determining region CDR-L1 of SEQ ID NO: 20, a CDR-L2 of SEQ ID NO: 21 and a CDR-L3 of SEQ ID NO: 22.
19. The antibody of claim 18, wherein the second antigen binding domain that specifically binds to CD28 comprises a heavy chain variable region (VHCD28) comprising an amino acid sequence of SEQ ID NO:23, and a light chain variable region (VLCD28) comprising an amino acid sequence of SEQ ID NO: 24.
20. The antibody of any one of claims 13 to 19, wherein the antibody comprises a first antigen binding domain comprising a VH BCMA comprising an amino acid sequence of SEQ ID NO:36 and VLBCMA comprising an amino acid sequence of SEQ ID NO:37 and a second
antigen binding domain comprising VH CD28 comprising an amino acid sequence of SEQ ID NO:23 and VL CD28 comprising an amino acid sequence of SEQ ID NO:24.
21. The antibody of any one of claims 13 to 20, comprising a first light chain comprising the amino acid sequence of SEQ ID NO:40, a first heavy chain comprising the amino acid sequence of SEQ ID NO:41, a second heavy chain comprising the amino acid sequence of SEQ ID NO:27 and a second light chain comprising the amino acid sequence of SEQ ID NO:28.
22. One or more isolated polynucleotide encoding the antibody of any one of claims 1 to 21.
23. One or more vector, particularly expression vector, comprising the polynucleotide(s) of claim 22.
24. A host cell comprising the polynucleotide(s) of claim 22 or the vector(s) of claim 23.
25. A method of producing an antibody that specifically binds to BCMA, comprising the steps of a) culturing the host cell of claim 24 under conditions suitable for the expression of the antibody and optionally b) recovering the antibody.
26. A pharmaceutical composition comprising the antibody of any one of claims 1 to 21 and at least one pharmaceutically acceptable excipient.
27. The antibody of any one of claims 1 to 21, or the pharmaceutical composition of claim 26, for use as a medicament.
28. The antibody of any one of claims 1 to 21, or the pharmaceutical composition of claim 26, for use in enhancing (a) T cell activation or (b) T cell effector functions.
29. The antibody of any one of claims 1 to 21, or the pharmaceutical composition of claim 26, for use in the treatment of cancer, in particular multiple myeloma.
30. The antibody of any one of claims 1 to 21, or the pharmaceutical composition of claim 26, for use according to claim 29, wherein the use is for administration in combination with a chemotherapeutic agent, radiation therapy and/ or other agents for use in cancer immunotherapy.
31. The antibody of any one of claims 1 to 21, or the pharmaceutical composition of claim 26, for use according to claim 29, wherein the use is for administration in combination with a T- cell activating anti-CD3 bispecific antibody.
32. The antibody or the pharmaceutical composition for use of claim 31, wherein the T-cell activating anti-CD3 bispecific antibody is an anti-GPRC5D/anti-CD3 antibody.
33. Use of the antibody of any one of claims 1 to 21, or the pharmaceutical composition of claim 26, in the manufacture of a medicament for the treatment of a disease, particularly for the treatment of cancer.
34. A method of treating a disease, particularly cancer, in an individual, comprising administering to said individual an effective amount of the antibody of claims 1 to 21, or the pharmaceutical composition of claim 26.
35. The method of claim 34, further comprising administration in combination with a chemotherapeutic agent, radiation therapy and/ or other agents for use in cancer immunotherapy, particularly in combination with a T-cell activating anti-CD3 bispecific antibody. ***
Priority Applications (1)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| CN202480036148.2A CN121241068A (en) | 2023-06-01 | 2024-05-29 | Bispecific antibodies targeting BCMA and CD28 |
Applications Claiming Priority (4)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| EP23176608.0 | 2023-06-01 | ||
| EP23176608 | 2023-06-01 | ||
| EP23189801 | 2023-08-04 | ||
| EP23189801.6 | 2023-08-04 |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| WO2024246083A1 true WO2024246083A1 (en) | 2024-12-05 |
Family
ID=91375677
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/EP2024/064688 Pending WO2024246083A1 (en) | 2023-06-01 | 2024-05-29 | Bispecific antibodies targeting bcma and cd28 |
Country Status (3)
| Country | Link |
|---|---|
| CN (1) | CN121241068A (en) |
| TW (1) | TW202504918A (en) |
| WO (1) | WO2024246083A1 (en) |
Citations (59)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US4186567A (en) | 1977-04-18 | 1980-02-05 | Hitachi Metals, Ltd. | Ornament utilizing rare earth-cobalt magnet |
| EP0404097A2 (en) | 1989-06-22 | 1990-12-27 | BEHRINGWERKE Aktiengesellschaft | Bispecific and oligospecific, mono- and oligovalent receptors, production and applications thereof |
| WO1993001161A1 (en) | 1991-07-11 | 1993-01-21 | Pfizer Limited | Process for preparing sertraline intermediates |
| WO1993016185A2 (en) | 1992-02-06 | 1993-08-19 | Creative Biomolecules, Inc. | Biosynthetic binding protein for cancer marker |
| US5500362A (en) | 1987-01-08 | 1996-03-19 | Xoma Corporation | Chimeric antibody with specificity to human B cell surface antigen |
| US5565332A (en) | 1991-09-23 | 1996-10-15 | Medical Research Council | Production of chimeric antibodies - a combinatorial approach |
| US5571894A (en) | 1991-02-05 | 1996-11-05 | Ciba-Geigy Corporation | Recombinant antibodies specific for a growth factor receptor |
| US5587458A (en) | 1991-10-07 | 1996-12-24 | Aronex Pharmaceuticals, Inc. | Anti-erbB-2 antibodies, combinations thereof, and therapeutic and diagnostic uses thereof |
| WO1997030087A1 (en) | 1996-02-16 | 1997-08-21 | Glaxo Group Limited | Preparation of glycosylated antibodies |
| US5731168A (en) | 1995-03-01 | 1998-03-24 | Genentech, Inc. | Method for making heteromultimeric polypeptides |
| US5821337A (en) | 1991-06-14 | 1998-10-13 | Genentech, Inc. | Immunoglobulin variants |
| WO1998058964A1 (en) | 1997-06-24 | 1998-12-30 | Genentech, Inc. | Methods and compositions for galactosylated glycoproteins |
| US5869046A (en) | 1995-04-14 | 1999-02-09 | Genentech, Inc. | Altered polypeptides with increased half-life |
| WO1999022764A1 (en) | 1997-10-31 | 1999-05-14 | Genentech, Inc. | Methods and compositions comprising glycoprotein glycoforms |
| US5959177A (en) | 1989-10-27 | 1999-09-28 | The Scripps Research Institute | Transgenic plants expressing assembled secretory antibodies |
| US5969108A (en) | 1990-07-10 | 1999-10-19 | Medical Research Council | Methods for producing members of specific binding pairs |
| US6040498A (en) | 1998-08-11 | 2000-03-21 | North Caroline State University | Genetically engineered duckweed |
| US6171586B1 (en) | 1997-06-13 | 2001-01-09 | Genentech, Inc. | Antibody formulation |
| US6248516B1 (en) | 1988-11-11 | 2001-06-19 | Medical Research Council | Single domain ligands, receptors comprising said ligands methods for their production, and use of said ligands and receptors |
| US6267958B1 (en) | 1995-07-27 | 2001-07-31 | Genentech, Inc. | Protein formulation |
| WO2002020565A2 (en) | 2000-09-08 | 2002-03-14 | Universität Zürich | Collections of repeat proteins comprising repeat modules |
| US6420548B1 (en) | 1999-10-04 | 2002-07-16 | Medicago Inc. | Method for regulating transcription of foreign genes |
| WO2003011878A2 (en) | 2001-08-03 | 2003-02-13 | Glycart Biotechnology Ag | Antibody glycosylation variants having increased antibody-dependent cellular cytotoxicity |
| US6602684B1 (en) | 1998-04-20 | 2003-08-05 | Glycart Biotechnology Ag | Glycosylation engineering of antibodies for improving antibody-dependent cellular cytotoxicity |
| US20030157108A1 (en) | 2001-10-25 | 2003-08-21 | Genentech, Inc. | Glycoprotein compositions |
| US20040093621A1 (en) | 2001-12-25 | 2004-05-13 | Kyowa Hakko Kogyo Co., Ltd | Antibody composition which specifically binds to CD20 |
| US6737056B1 (en) | 1999-01-15 | 2004-05-18 | Genentech, Inc. | Polypeptide variants with altered effector function |
| US20040132066A1 (en) | 2002-02-14 | 2004-07-08 | Kalobios, Inc. | Methods for affinity maturation |
| US6818418B1 (en) | 1998-12-10 | 2004-11-16 | Compound Therapeutics, Inc. | Protein scaffolds for antibody mimics and other binding proteins |
| US20050123546A1 (en) | 2003-11-05 | 2005-06-09 | Glycart Biotechnology Ag | Antigen binding molecules with increased Fc receptor binding affinity and effector function |
| WO2005056764A2 (en) | 2003-12-05 | 2005-06-23 | Compound Therapeutics, Inc. | Inhibitors of type 2 vascular endothelial growth factor receptors |
| WO2005100402A1 (en) | 2004-04-13 | 2005-10-27 | F.Hoffmann-La Roche Ag | Anti-p-selectin antibodies |
| US20050260186A1 (en) | 2003-03-05 | 2005-11-24 | Halozyme, Inc. | Soluble glycosaminoglycanases and methods of preparing and using soluble glycosaminoglycanases |
| US6982321B2 (en) | 1986-03-27 | 2006-01-03 | Medical Research Council | Altered antibodies |
| WO2006029879A2 (en) | 2004-09-17 | 2006-03-23 | F.Hoffmann-La Roche Ag | Anti-ox40l antibodies |
| EP1641818A1 (en) | 2003-07-04 | 2006-04-05 | Affibody AB | Polypeptides having binding affinity for her2 |
| WO2006044908A2 (en) | 2004-10-20 | 2006-04-27 | Genentech, Inc. | Antibody formulation in histidine-acetate buffer |
| US20060104968A1 (en) | 2003-03-05 | 2006-05-18 | Halozyme, Inc. | Soluble glycosaminoglycanases and methods of preparing and using soluble glycosaminogly ycanases |
| US7087409B2 (en) | 1997-12-05 | 2006-08-08 | The Scripps Research Institute | Humanization of murine antibody |
| WO2006082515A2 (en) | 2005-02-07 | 2006-08-10 | Glycart Biotechnology Ag | Antigen binding molecules that bind egfr, vectors encoding same, and uses thereof |
| US7125978B1 (en) | 1999-10-04 | 2006-10-24 | Medicago Inc. | Promoter for regulating expression of foreign genes |
| US7166697B1 (en) | 1998-03-06 | 2007-01-23 | Diatech Pty. Ltd. | V-like domain binding molecules |
| US7250297B1 (en) | 1997-09-26 | 2007-07-31 | Pieris Ag | Anticalins |
| US20070224633A1 (en) | 2003-08-25 | 2007-09-27 | Pieris Ag | Muteins of Tear Lipocalin |
| US20080139791A1 (en) | 1998-12-10 | 2008-06-12 | Adnexus Therapeutics, Inc. | Pharmaceutically acceptable Fn3 Polypeptides for human treatments |
| WO2008098796A1 (en) | 2007-02-16 | 2008-08-21 | Nascacell Technologies Ag | Polypeptide comprising a knottin protein moiety |
| US7521541B2 (en) | 2004-09-23 | 2009-04-21 | Genetech Inc. | Cysteine engineered antibodies and conjugates |
| US7527791B2 (en) | 2004-03-31 | 2009-05-05 | Genentech, Inc. | Humanized anti-TGF-beta antibodies |
| WO2009080252A1 (en) | 2007-12-21 | 2009-07-02 | F. Hoffmann-La Roche Ag | Bivalent, bispecific antibodies |
| WO2009089004A1 (en) | 2008-01-07 | 2009-07-16 | Amgen Inc. | Method for making antibody fc-heterodimeric molecules using electrostatic steering effects |
| WO2012020006A2 (en) | 2010-08-13 | 2012-02-16 | Roche Glycart Ag | Anti-fap antibodies and methods of use |
| WO2012130831A1 (en) | 2011-03-29 | 2012-10-04 | Roche Glycart Ag | Antibody fc variants |
| WO2012163805A1 (en) | 2011-05-27 | 2012-12-06 | Glaxo Group Limited | Bcma (cd269/tnfrsf17) -binding proteins |
| WO2013072415A1 (en) | 2011-11-15 | 2013-05-23 | Amgen Research (Munich) Gmbh | Binding molecules for bcma and cd3 |
| WO2015150447A1 (en) | 2014-04-02 | 2015-10-08 | F. Hoffmann-La Roche Ag | Multispecific antibodies |
| WO2016062734A1 (en) | 2014-10-24 | 2016-04-28 | F. Hoffmann-La Roche Ag | Vh-vl-interdomain angle based antibody humanization |
| EP2101823B1 (en) | 2007-01-09 | 2016-11-23 | CureVac AG | Rna-coded antibody |
| WO2016205520A1 (en) | 2015-06-16 | 2016-12-22 | Genentech, Inc. | Humanized and affinity matured antibodies to fcrh5 and methods of use |
| WO2020127618A1 (en) | 2018-12-21 | 2020-06-25 | F. Hoffmann-La Roche Ag | Tumor-targeted agonistic cd28 antigen binding molecules |
-
2024
- 2024-05-29 WO PCT/EP2024/064688 patent/WO2024246083A1/en active Pending
- 2024-05-29 TW TW113119794A patent/TW202504918A/en unknown
- 2024-05-29 CN CN202480036148.2A patent/CN121241068A/en active Pending
Patent Citations (64)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US4186567A (en) | 1977-04-18 | 1980-02-05 | Hitachi Metals, Ltd. | Ornament utilizing rare earth-cobalt magnet |
| US6982321B2 (en) | 1986-03-27 | 2006-01-03 | Medical Research Council | Altered antibodies |
| US5500362A (en) | 1987-01-08 | 1996-03-19 | Xoma Corporation | Chimeric antibody with specificity to human B cell surface antigen |
| US6248516B1 (en) | 1988-11-11 | 2001-06-19 | Medical Research Council | Single domain ligands, receptors comprising said ligands methods for their production, and use of said ligands and receptors |
| EP0404097A2 (en) | 1989-06-22 | 1990-12-27 | BEHRINGWERKE Aktiengesellschaft | Bispecific and oligospecific, mono- and oligovalent receptors, production and applications thereof |
| US6417429B1 (en) | 1989-10-27 | 2002-07-09 | The Scripps Research Institute | Transgenic plants expressing assembled secretory antibodies |
| US5959177A (en) | 1989-10-27 | 1999-09-28 | The Scripps Research Institute | Transgenic plants expressing assembled secretory antibodies |
| US5969108A (en) | 1990-07-10 | 1999-10-19 | Medical Research Council | Methods for producing members of specific binding pairs |
| US5571894A (en) | 1991-02-05 | 1996-11-05 | Ciba-Geigy Corporation | Recombinant antibodies specific for a growth factor receptor |
| US5821337A (en) | 1991-06-14 | 1998-10-13 | Genentech, Inc. | Immunoglobulin variants |
| WO1993001161A1 (en) | 1991-07-11 | 1993-01-21 | Pfizer Limited | Process for preparing sertraline intermediates |
| US5565332A (en) | 1991-09-23 | 1996-10-15 | Medical Research Council | Production of chimeric antibodies - a combinatorial approach |
| US5587458A (en) | 1991-10-07 | 1996-12-24 | Aronex Pharmaceuticals, Inc. | Anti-erbB-2 antibodies, combinations thereof, and therapeutic and diagnostic uses thereof |
| WO1993016185A2 (en) | 1992-02-06 | 1993-08-19 | Creative Biomolecules, Inc. | Biosynthetic binding protein for cancer marker |
| US7695936B2 (en) | 1995-03-01 | 2010-04-13 | Genentech, Inc. | Knobs and holes heteromeric polypeptides |
| US5731168A (en) | 1995-03-01 | 1998-03-24 | Genentech, Inc. | Method for making heteromultimeric polypeptides |
| US5821333A (en) | 1995-03-01 | 1998-10-13 | Genetech, Inc. | Method for making heteromultimeric polypeptides |
| US5869046A (en) | 1995-04-14 | 1999-02-09 | Genentech, Inc. | Altered polypeptides with increased half-life |
| US6267958B1 (en) | 1995-07-27 | 2001-07-31 | Genentech, Inc. | Protein formulation |
| WO1997030087A1 (en) | 1996-02-16 | 1997-08-21 | Glaxo Group Limited | Preparation of glycosylated antibodies |
| US6171586B1 (en) | 1997-06-13 | 2001-01-09 | Genentech, Inc. | Antibody formulation |
| WO1998058964A1 (en) | 1997-06-24 | 1998-12-30 | Genentech, Inc. | Methods and compositions for galactosylated glycoproteins |
| US7250297B1 (en) | 1997-09-26 | 2007-07-31 | Pieris Ag | Anticalins |
| WO1999022764A1 (en) | 1997-10-31 | 1999-05-14 | Genentech, Inc. | Methods and compositions comprising glycoprotein glycoforms |
| US7087409B2 (en) | 1997-12-05 | 2006-08-08 | The Scripps Research Institute | Humanization of murine antibody |
| US7166697B1 (en) | 1998-03-06 | 2007-01-23 | Diatech Pty. Ltd. | V-like domain binding molecules |
| US6602684B1 (en) | 1998-04-20 | 2003-08-05 | Glycart Biotechnology Ag | Glycosylation engineering of antibodies for improving antibody-dependent cellular cytotoxicity |
| US6040498A (en) | 1998-08-11 | 2000-03-21 | North Caroline State University | Genetically engineered duckweed |
| US6818418B1 (en) | 1998-12-10 | 2004-11-16 | Compound Therapeutics, Inc. | Protein scaffolds for antibody mimics and other binding proteins |
| US20080139791A1 (en) | 1998-12-10 | 2008-06-12 | Adnexus Therapeutics, Inc. | Pharmaceutically acceptable Fn3 Polypeptides for human treatments |
| US7332581B2 (en) | 1999-01-15 | 2008-02-19 | Genentech, Inc. | Polypeptide variants with altered effector function |
| US6737056B1 (en) | 1999-01-15 | 2004-05-18 | Genentech, Inc. | Polypeptide variants with altered effector function |
| US6420548B1 (en) | 1999-10-04 | 2002-07-16 | Medicago Inc. | Method for regulating transcription of foreign genes |
| US7125978B1 (en) | 1999-10-04 | 2006-10-24 | Medicago Inc. | Promoter for regulating expression of foreign genes |
| US20040132028A1 (en) | 2000-09-08 | 2004-07-08 | Stumpp Michael Tobias | Collection of repeat proteins comprising repeat modules |
| WO2002020565A2 (en) | 2000-09-08 | 2002-03-14 | Universität Zürich | Collections of repeat proteins comprising repeat modules |
| WO2003011878A2 (en) | 2001-08-03 | 2003-02-13 | Glycart Biotechnology Ag | Antibody glycosylation variants having increased antibody-dependent cellular cytotoxicity |
| US20030157108A1 (en) | 2001-10-25 | 2003-08-21 | Genentech, Inc. | Glycoprotein compositions |
| US20040093621A1 (en) | 2001-12-25 | 2004-05-13 | Kyowa Hakko Kogyo Co., Ltd | Antibody composition which specifically binds to CD20 |
| US20040132066A1 (en) | 2002-02-14 | 2004-07-08 | Kalobios, Inc. | Methods for affinity maturation |
| US20060104968A1 (en) | 2003-03-05 | 2006-05-18 | Halozyme, Inc. | Soluble glycosaminoglycanases and methods of preparing and using soluble glycosaminogly ycanases |
| US20050260186A1 (en) | 2003-03-05 | 2005-11-24 | Halozyme, Inc. | Soluble glycosaminoglycanases and methods of preparing and using soluble glycosaminoglycanases |
| EP1641818A1 (en) | 2003-07-04 | 2006-04-05 | Affibody AB | Polypeptides having binding affinity for her2 |
| US20070224633A1 (en) | 2003-08-25 | 2007-09-27 | Pieris Ag | Muteins of Tear Lipocalin |
| US20050123546A1 (en) | 2003-11-05 | 2005-06-09 | Glycart Biotechnology Ag | Antigen binding molecules with increased Fc receptor binding affinity and effector function |
| WO2005056764A2 (en) | 2003-12-05 | 2005-06-23 | Compound Therapeutics, Inc. | Inhibitors of type 2 vascular endothelial growth factor receptors |
| US7527791B2 (en) | 2004-03-31 | 2009-05-05 | Genentech, Inc. | Humanized anti-TGF-beta antibodies |
| WO2005100402A1 (en) | 2004-04-13 | 2005-10-27 | F.Hoffmann-La Roche Ag | Anti-p-selectin antibodies |
| WO2006029879A2 (en) | 2004-09-17 | 2006-03-23 | F.Hoffmann-La Roche Ag | Anti-ox40l antibodies |
| US7521541B2 (en) | 2004-09-23 | 2009-04-21 | Genetech Inc. | Cysteine engineered antibodies and conjugates |
| WO2006044908A2 (en) | 2004-10-20 | 2006-04-27 | Genentech, Inc. | Antibody formulation in histidine-acetate buffer |
| WO2006082515A2 (en) | 2005-02-07 | 2006-08-10 | Glycart Biotechnology Ag | Antigen binding molecules that bind egfr, vectors encoding same, and uses thereof |
| EP2101823B1 (en) | 2007-01-09 | 2016-11-23 | CureVac AG | Rna-coded antibody |
| WO2008098796A1 (en) | 2007-02-16 | 2008-08-21 | Nascacell Technologies Ag | Polypeptide comprising a knottin protein moiety |
| WO2009080252A1 (en) | 2007-12-21 | 2009-07-02 | F. Hoffmann-La Roche Ag | Bivalent, bispecific antibodies |
| WO2009089004A1 (en) | 2008-01-07 | 2009-07-16 | Amgen Inc. | Method for making antibody fc-heterodimeric molecules using electrostatic steering effects |
| WO2012020006A2 (en) | 2010-08-13 | 2012-02-16 | Roche Glycart Ag | Anti-fap antibodies and methods of use |
| WO2012130831A1 (en) | 2011-03-29 | 2012-10-04 | Roche Glycart Ag | Antibody fc variants |
| WO2012163805A1 (en) | 2011-05-27 | 2012-12-06 | Glaxo Group Limited | Bcma (cd269/tnfrsf17) -binding proteins |
| WO2013072415A1 (en) | 2011-11-15 | 2013-05-23 | Amgen Research (Munich) Gmbh | Binding molecules for bcma and cd3 |
| WO2015150447A1 (en) | 2014-04-02 | 2015-10-08 | F. Hoffmann-La Roche Ag | Multispecific antibodies |
| WO2016062734A1 (en) | 2014-10-24 | 2016-04-28 | F. Hoffmann-La Roche Ag | Vh-vl-interdomain angle based antibody humanization |
| WO2016205520A1 (en) | 2015-06-16 | 2016-12-22 | Genentech, Inc. | Humanized and affinity matured antibodies to fcrh5 and methods of use |
| WO2020127618A1 (en) | 2018-12-21 | 2020-06-25 | F. Hoffmann-La Roche Ag | Tumor-targeted agonistic cd28 antigen binding molecules |
Non-Patent Citations (107)
| Title |
|---|
| "Monoclonal Antibody Production Techniques and Applications", 1987, MARCEL DEKKER, INC, pages: 51 - 63 |
| "NCBI GenBank", Database accession no. BAB71849.1 |
| "NCBI", Database accession no. NP_000724.1 |
| "Remington's Pharmaceutical Sciences", 1990, MACK PRINTING COMPANY |
| "UniProt accession", Database accession no. P08637 |
| "UniProt", Database accession no. P10747 |
| ABHINANDAN, K. R.MARTIN, ANDREW C. R, J. MOL. BIOL., vol. 369, 2007, pages 1015 - 1028 |
| ACUTOMICHEL, NAT REV IMMUNOL, vol. 3, 2003, pages 939 - 951 |
| ALEXANDER BUJOTZEK ET AL: "VH-VL orientation prediction for antibody humanization candidate selection: A case study", MABS, vol. 8, no. 2, 4 December 2015 (2015-12-04), US, pages 288 - 305, XP055416051, ISSN: 1942-0862, DOI: 10.1080/19420862.2015.1117720 * |
| ALMAGROFRANSSON, FRONT BIOSCI, vol. 13, 2008, pages 1619 - 1633 |
| ARMOUR, K.L. ET AL., EUR. J. IMMUNOL., vol. 29, 1999, pages 2613 - 2624 |
| BAHLIS ET AL., BLOOD, vol. 109, no. 11, 2007, pages 5002 - 5010 |
| BEYERSDORF ET AL., ANN. RHEUM. DIS, vol. 64, 2005, pages iv91 - iv95 |
| BIOCHIM BIOPHYS ACTA, vol. 1482, 2000, pages 337 - 350 |
| BOWIE, J. U ET AL., SCIENCE, vol. 247, 1990, pages 1306 - 10 |
| BRUGGEMANN ET AL., J EXP MED, vol. 166, 1987, pages 1351 - 1361 |
| BURTON, MOL. IMMUNOL., vol. 22, 1985, pages 161 - 206 |
| CAPEL, P.J. ET AL., IMMUNOMETHODS, vol. 113, 1994, pages 269 - 315 |
| CARRENOCOLLINS, ANNU REV IMMUNOL, vol. 20, 2002, pages 29 - 53 |
| CARTER, J IMMUNOL METH, vol. 248, 2001, pages 7 - 15 |
| CARTER, J IMMUNOL METHODS, vol. 248, 2001, pages 7 - 15 |
| CHENFLIES, NAT REV IMMUNOL., vol. 13, no. 4, 2013, pages 227 - 42 |
| CHO ET AL., FRONT IMMUNOLOG, vol. 9, 2018, pages 1821 |
| CHOTHIALESK, J. MOL. BIOL., vol. 196, 1987, pages 901 - 917 |
| CLYNES ET AL., PROC NATL ACAD SCI USA, vol. 95, 1998, pages 652 - 656 |
| CRAGG ET AL., BLOOD, vol. 101, 2003, pages 1045 - 1052 |
| CRAGGGLENNIE, BLOOD, vol. 103, 2004, pages 2738 - 2743 |
| CUNNINGHAMWELLS, SCIENCE, vol. 244, 1989, pages 1081 - 1085 |
| DALL' ACQUA ET AL., METHODS, vol. 36, 2005, pages 61 - 68 |
| DE HAAS, M. ET AL., J. LAB. CLIN. MED., vol. 126, 1995, pages 330 - 341 |
| ENGELHARDT ET AL., J IMMUNOL, vol. 177, 2006, pages 1052 - 1061 |
| ESENSTEN ET AL., IMMUNITY, vol. 44, 2016, pages 973 - 988 |
| EXPERT OPIN. BIOL. THER., vol. 5, 2005, pages 783 - 797 |
| EXPERT OPINION ON INVESTIGATIONAL DRUGS, vol. 16, no. 6, June 2007 (2007-06-01), pages 909 - 917 |
| FRASER ET AL., SCIENCE, vol. 251, 1991, pages 313 - 316 |
| GAZZANO-SANTORO ET AL., J IMMUNOL METHODS, vol. 202, 1996, pages 163 |
| GEBAUERSKERRA: "Engineered protein scaffolds as next-generation antibody therapeutics", CURR OPIN CHEM BIOL, vol. 13, 2009, pages 245 - 255 |
| GERNGROSS, NAT BIOTECH, vol. 22, 2004, pages 1409 - 1414 |
| GESSNER, J.E. ET AL., ANN. HEMATOL., vol. 76, 1998, pages 231 - 248 |
| GIUDICELLI, V. ET AL.: "IMGT/LIGM-DB, the IMGT® comprehensive database of immunoglobulin and T cell receptor nucleotide sequences", NUCL. ACIDS RES, vol. 34, no. S1, 2006, pages 781 - 784 |
| GIUDICELLI, V. ET AL.: "IMGT/LIGM-DB, the IMGT® comprehensive database of immunoglobulin and T cell receptor nucleotide sequences.", NUCL. ACIDS RES., vol. 34, no. 1, 2006, pages D781 - D784, XP002532807, DOI: 10.1093/NAR/GKJ088 |
| GRAHAM ET AL., J GEN VIROL, vol. 36, 1977, pages 59 |
| HARLOWLANE: "Antibodies: A Laboratory Manual", 1988, COLD SPRING HARBOR LABORATORY |
| HEELEY, ENDOCR RES, vol. 28, 2002, pages 217 - 229 |
| HELLSTROM ET AL., PROC NATL ACAD SCI USA, vol. 82, 1985, pages 1499 - 1502 |
| HELLSTROM ET AL., PROC NATL ACAD SCI USA, vol. 83, 1986, pages 7059 - 7063 |
| HOLLINGER ET AL., PROC NATL ACAD SCI USA, vol. 90, 1993, pages 6444 - 6448 |
| HOOGENBOOM ET AL.: "Methods in Molecular Biology", vol. 178, 1996, HUMANA PRESS, article "Epitope Mapping Protocols", pages: 255 - 268 |
| HOUSTON, J.S, METHODS IN ENZYMOL., vol. 203, 1991, pages 46 - 96 |
| HUDSON ET AL., NAT MED, vol. 9, 2003, pages 129 - 134 |
| HUNIG, NAT REV IMMUNOL, vol. 12, 2012, pages 317 - 318 |
| J. BIOL. CHEM, vol. 274, 1999, pages 24066 - 24073 |
| J. MOL. BIOL., vol. 332, 2003, pages 489 - 503 |
| JAMES R. APGAR ET AL: "Beyond CDR-grafting: Structure-guided humanization of framework and CDR regions of an anti-myostatin antibody", MABS, vol. 8, no. 7, 13 September 2016 (2016-09-13), US, pages 1302 - 1318, XP055416065, ISSN: 1942-0862, DOI: 10.1080/19420862.2016.1215786 * |
| JONES ET AL., NATURE, vol. 321, 1986, pages 522 - 525 |
| JOURNAL OF IMMUNOLOGICAL METHODS, vol. 248, no. 1-2, 2001, pages 31 - 45 |
| KABAT ET AL.: "U.S. Dept. of Health and Human Services", SEQUENCE OF PROTEINS OF IMMUNOLOGICAL INTEREST, 1983 |
| KABAT, E.A. ET AL.: "Sequences of Proteins of Immunological Interest", 1991, PUBLIC HEALTH SERVICE, NATIONAL INSTITUTES OF HEALTH |
| KAM, N.W. ET AL., PROC. NATL. ACAD. SCI. USA, vol. 102, 2005, pages 11600 - 11605 |
| KAMPHORST ET AL., SCIENCE, vol. 355, 2017, pages 1423 - 1427 |
| KINDT ET AL.: "Kuby Immunology", 2007, W.H. FREEMAN AND CO., pages: 91 |
| KLIMKA ET AL., BR J CANCER, vol. 83, 2000, pages 252 - 260 |
| LEE ET AL., NATURE MEDICINE, vol. 29, 2023, pages 2295 - 2306 |
| LI ET AL., NAT BIOTECH, vol. 24, 2006, pages 210 - 215 |
| LILJEBLAD ET AL., GLYCO J, vol. 17, 2000, pages 323 - 329 |
| LINSLEY ET AL., PROC NATL ACAD SCI USA, vol. 87, no. 13, 1990, pages 5031 - 5 |
| LONBERG, CURR OPIN IMMUNOL, vol. 20, 2008, pages 450 - 459 |
| LONBERG, NAT BIOTECH, vol. 23, 2005, pages 1117 - 1125 |
| MACCALLUM ET AL., J. MOL. BIOL., vol. 262, 1996, pages 732 - 745 |
| MATHER ET AL., ANNALS N.Y. ACAD SCI, vol. 383, 1982, pages 44 - 68 |
| MATHER, BIOL REPROD, vol. 23, 1980, pages 243 - 251 |
| MCCAFFERTY ET AL., NATURE, vol. 352, 1991, pages 624 - 628 |
| MOREAU ET AL., THE LANCET ONCOLOGY, vol. 22, no. 3, 2021, pages e105 - e118 |
| MORRISON ET AL., PROC NATL ACAD SCI, vol. 81, 1984, pages 6851 - 6855 |
| MORRISONOI, ADV IMMUNOL, vol. 44, 1988, pages 65 - 92 |
| MURRAY ET AL., BLOOD, vol. 123, no. 24, 2014, pages 3770 - 3779 |
| NATURE BIOTECHNOLOGY, vol. 23, no. 12, 2005, pages 1556 - 1561 |
| OTZ ET AL., LEUKEMIA, vol. 23, no. 1, 2009, pages 71 - 77 |
| PACE ET AL., PROTEIN SCIENCE, vol. 4, 1995, pages 2411 - 1423 |
| PADLAN, MOL IMMUNOL, vol. 28, 1991, pages 489 - 498 |
| PADLAN, MOLEC IMMUN, vol. 31, no. 3, 1994, pages 169 - 217 |
| PNAS, vol. 100, no. 4, 2003, pages 1700 - 1705 |
| PROTEIN ENG. DES. SEL., vol. 17, 2004, pages 455 - 462 |
| PROTEIN ENG. DES. SEL., vol. 18, 2005, pages 435 - 444 |
| QUEEN ET AL., PROC NATL ACAD SCI USA, vol. 86, 1989, pages 10029 - 10033 |
| RAVETCH, J.V.KINET, J.P., ANNU. REV. IMMUNOL, vol. 9, 1991, pages 457 - 492 |
| REYNISSON B: "NetMHCpan-4.1 and NetMHCIIpan-4.0: improved predictions of MHC antigen presentation by concurrent motif deconvolution and integration of MS MHC eluted ligand data", NUCL. ACIDS RES, vol. 48, no. 1, 2020, pages W449 - W454 |
| RIDGWAY ET AL., PROT ENG, vol. 9, 1996, pages 617 - 621 |
| RIECHMANN ET AL., NATURE, vol. 332, 1988, pages 323 - 329 |
| ROUX ET AL., J. IMMUNOL., vol. 161, 1998, pages 4083 |
| SAMBROOK ET AL.: "Molecular cloning: A laboratory manual", 1989, COLD SPRING HARBOR LABORATORY PRESS |
| SCHAEFER, W. ET AL., PNAS, vol. 108, 2011, pages 11187 - 1191 |
| SCHLOTHAUER ET AL., MABS, vol. 5, no. 4, 2013, pages 576 - 86 |
| SHIELDS, R.L. ET AL., J. BIOL. CHEM., vol. 276, 2001, pages 6591 - 6604 |
| SIEGEL ET AL., PHARMACEUTICS, vol. 14, no. 12, 2022, pages 2672 |
| STADLER ET AL., NATURE MEDICINE, vol. 23, 2017, pages 815 - 817 |
| STEINER ET AL., J. PROTEOME RES, vol. 19, 2020, pages 3792 - 3806 |
| STUMPP ET AL.: "Darpins: A new generation of protein therapeutics", DRUG DISCOVERY TODAY, vol. 13, 2008, pages 695 - 701, XP023440383, DOI: 10.1016/j.drudis.2008.04.013 |
| TAN P ET AL: ""SUPERHUMANIZED" ANTIBODIES: REDUCTION OF IMMUNOGENIC POTENTIAL BY COMPLEMENTARITY-DETERMINING REGION GRAFTING WITH HUMAN GERMLINE SEQUENCES: APPLICATION TO AN ANTI-CD28", THE JOURNAL OF IMMUNOLOGY, WILLIAMS & WILKINS CO, US, vol. 169, no. 2, 15 July 2002 (2002-07-15), pages 1119 - 1125, XP008056579, ISSN: 0022-1767 * |
| URLAUB ET AL., PROC NATL ACAD SCI USA, vol. 77, 1980, pages 4216 |
| VAN DE WINKEL, J.G.ANDERSON, C.L, J. LEUKOC. BIOL., vol. 49, 1991, pages 511 - 524 |
| VAN DIJKVAN DE WINKEL, CURR OPIN PHARMACOL, vol. 5, 2001, pages 368 - 74 |
| VERHOEYEN ET AL., SCIENCE, vol. 239, 1988, pages 1534 - 1536 |
| WHO DRUG INFORMATION, vol. 33, no. 3, 2019 |
| WHO DRUG INFORMATION, vol. 35, no. 1, 2021 |
| WHO DRUG INFORMATION, vol. 36, no. 1, 2022 |
| WRIGHT ET AL., TIBTECH, vol. 15, 1997, pages 26 - 32 |
Also Published As
| Publication number | Publication date |
|---|---|
| TW202504918A (en) | 2025-02-01 |
| CN121241068A (en) | 2025-12-30 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| US20230357397A1 (en) | Combination therapy with targeted 4-1bb (cd137) agonists | |
| US20230416365A1 (en) | Tumor-targeted agonistic cd28 antigen binding molecules | |
| US11718680B2 (en) | Combination therapy of anti-CD20/anti-CD3 bispecific antibodies and 4-1BB (CD137) agonists | |
| US12240911B2 (en) | Bispecific antibodies specific for OX40 | |
| US20200223925A1 (en) | Tumor-targeted superagonistic cd28 antigen binding molecules | |
| US20230416366A1 (en) | Anti-cd3/anti-cd28 bispecific antigen binding molecules | |
| EP3999536A1 (en) | Antibodies binding to nkg2d | |
| EP3703821A2 (en) | Bispecific 2+1 contorsbodies | |
| EP3359568A1 (en) | Bispecific antibodies with tetravalency for a costimulatory tnf receptor | |
| EP3765489A1 (en) | Therapeutic combination of 4-1 bb agonists with anti-cd20 antibodies | |
| EP4168448A1 (en) | Agonistic cd28 antigen binding molecules targeting her2 | |
| WO2024246083A1 (en) | Bispecific antibodies targeting bcma and cd28 | |
| WO2020260326A1 (en) | Novel icos antibodies and tumor-targeted antigen binding molecules comprising them | |
| WO2024246086A1 (en) | Immunostimulatory antigen binding molecules that specifically bind to bcma | |
| CA3123493C (en) | Tumor-targeted agonistic cd28 antigen binding molecules |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| 121 | Ep: the epo has been informed by wipo that ep was designated in this application |
Ref document number: 24730637 Country of ref document: EP Kind code of ref document: A1 |
|
| WWE | Wipo information: entry into national phase |
Ref document number: 2024730637 Country of ref document: EP |
|
| NENP | Non-entry into the national phase |
Ref country code: DE |