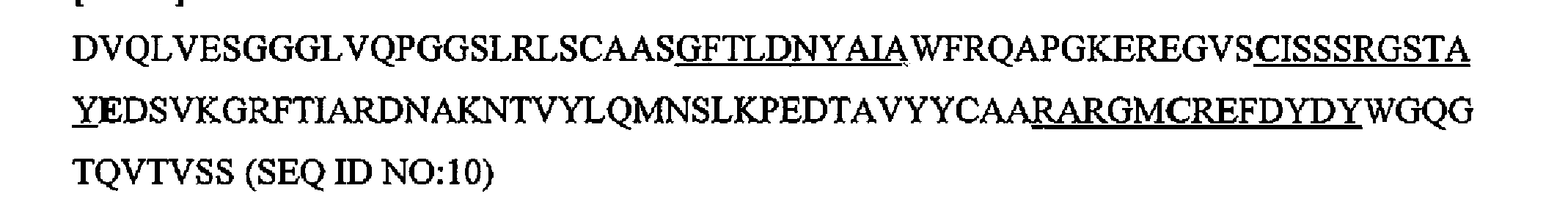WO2024243292A2 - Novel complement system inhibiting antibodies - Google Patents
Novel complement system inhibiting antibodies Download PDFInfo
- Publication number
- WO2024243292A2 WO2024243292A2 PCT/US2024/030542 US2024030542W WO2024243292A2 WO 2024243292 A2 WO2024243292 A2 WO 2024243292A2 US 2024030542 W US2024030542 W US 2024030542W WO 2024243292 A2 WO2024243292 A2 WO 2024243292A2
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- seq
- amino acid
- raav
- acid sequence
- sequence
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Pending
Links
Classifications
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K16/00—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies
- C07K16/18—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/20—Immunoglobulins specific features characterized by taxonomic origin
- C07K2317/22—Immunoglobulins specific features characterized by taxonomic origin from camelids, e.g. camel, llama or dromedary
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/20—Immunoglobulins specific features characterized by taxonomic origin
- C07K2317/24—Immunoglobulins specific features characterized by taxonomic origin containing regions, domains or residues from different species, e.g. chimeric, humanized or veneered
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/30—Immunoglobulins specific features characterized by aspects of specificity or valency
- C07K2317/33—Crossreactivity, e.g. for species or epitope, or lack of said crossreactivity
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/30—Immunoglobulins specific features characterized by aspects of specificity or valency
- C07K2317/34—Identification of a linear epitope shorter than 20 amino acid residues or of a conformational epitope defined by amino acid residues
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/50—Immunoglobulins specific features characterized by immunoglobulin fragments
- C07K2317/56—Immunoglobulins specific features characterized by immunoglobulin fragments variable (Fv) region, i.e. VH and/or VL
- C07K2317/569—Single domain, e.g. dAb, sdAb, VHH, VNAR or nanobody®
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/70—Immunoglobulins specific features characterized by effect upon binding to a cell or to an antigen
- C07K2317/76—Antagonist effect on antigen, e.g. neutralization or inhibition of binding
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/90—Immunoglobulins specific features characterized by (pharmaco)kinetic aspects or by stability of the immunoglobulin
- C07K2317/92—Affinity (KD), association rate (Ka), dissociation rate (Kd) or EC50 value
Definitions
- a computer readable XML file entitled “090400-5023-WO Sequence Listing” created on May 21, 2024, with a file size of about 249,282 bytes contains the sequence listing for this application and is hereby incorporated by reference in its entirety.
- a number of human diseases are caused by complement dysregulation, resulting in complement-mediated autologous tissue injury.
- the complement dysregulation may arise from mutations, either somatic or germline, in complement regulator or regulator- related genes such that these regulators no longer function normally.
- mAbs monoclonal antibodies
- peptides or other small molecules that bind and block specific alternative pathway or terminal pathway complement components.
- a clinically validated example is Eculizumab, a humanized mAb against complement C5 which has been approved for the treatment of paroxysmal nocturnal hemoglobinuria (PNH) and atypical hemolytic uremic syndrome (aHUS).
- PNH paroxysmal nocturnal hemoglobinuria
- aHUS atypical hemolytic uremic syndrome
- Other approaches that have been described include mAbs against factor B (fB), factor D (fD), or properdin (fP), and a cyclic peptide that binds and inhibits C3.
- fB factor B
- fD factor D
- fP properdin
- cyclic peptide that binds and inhibits C3.
- IV intravenous
- recombinant regulatory proteins such as soluble DAF, CR1, CRIg and proteins comprising minimal domains of fluid phase regulator fH (N-terminal short consensus repeat [SCR] 1-5 and C-terminal SCR 19-20) or fusion proteins between fH and CR2 (TT30) have been tested.
- fluid phase regulator fH N-terminal short consensus repeat [SCR] 1-5 and C-terminal SCR 19-20
- TT30 fusion proteins between fH and CR2
- polypeptides comprising antibodies, preferably comprising a single variable domain on a heavy chain (VHH or nanobody) capable of modulating complement activity by specifically binding to human complement factor C3, C3a, C3b, C5, C5a and/or C5b.
- VHH heavy chain
- the polypeptide comprises a VH or VHH domain of, or derived from, a camelid heavy chain antibody.
- the VH or VHH domain comprises one or more complementarity determining regions (CDRs) as set forth in Tables 1 and 2.
- the VH or VHH domain comprises the CDRS (CDR1, CDR2 and CDR3) of one or more antibodies selected from the group consisting of SEQ ID Nos: 1-14 and SEQ ID Nos:57-65.
- said CDR1, CDR2 and CDR3 are separated by frame regions FR1, FR2, FR3, and FR4 of the VH or VHH domain.
- the VH or VHH domain comprises or consists of any one of the amino acid sequences set forth as SEQ ID Nos: 1-14 or any one of the amino acid sequences set forth as SEQ ID Nos: 57-65.
- the VH or VHH domain is humanized and comprises or consists of any one of amino acid sequences set forth as SEQ ID Nos:94-136 or any one of the amino acid sequences set forth as SEQ ID Nos: 137-163.
- the VH or VHH domain comprises a tag, such as a Strep or his6 tag.
- the VH or VHH domain is fused to another polypeptide.
- a polynucleotide (DNA or RNA) is provided, wherein said polynucleotide comprises a nucleotide sequence that encodes a polypeptide comprising one or more CDRs of a VH or VHH domain as herein described, or encodes a polypeptide comprising a VH or VHH domain as herein described.
- an expression vector comprising said polynucleotide is provided, wherein said nucleotide sequence is operably linked to an expression control sequence (e.g., a promoter).
- a pharmaceutical composition comprising a polypeptide or expression vector as described herein and a pharmaceutically acceptable carrier is provided.
- recombinant AAV (rAAV) virions comprising a variant AAV capsid sequence encapsulating a heterologous nucleic acid comprising a nucleotide sequence encoding an anti-03 and/or anti-C5 antibody as herein described.
- the nucleotide sequence encoding the antibody is operably linked to an expression control sequence.
- the variant AAV capsid protein of the rAAV comprises a capsid protein comprising a peptide insertion of from about 7 amino acids to about 20 amino acids (a “heterologous peptide” or “peptide insertion”) in the GH-loop of the capsid protein, preferably in a surface-exposed region of the GH-loop, relative to a corresponding parental AAV capsid protein, wherein the peptide insertion comprises the amino acid sequence ISDQTKH (SEQ ID NO: 168).
- the peptide insertion comprises one to three spacer amino acids (Y1-Y3) at the amino and/or carboxy terminal of the amino acid sequence ISDQTKH (SEQ ID NO: 168), wherein each of Y1-Y3 is independently selected from Ala, Leu, Gly, Ser, Thr, and Pro.
- the peptide insertion comprises, consists essentially of or consists of the amino acid sequence LAISDQTKHA (SEQ ID NO: 169).
- the peptide is inserted following any of the amino acids in positions 584-591 in VP1 of AAV2 or a corresponding position in another AAV serotype (i.e., the insertion site is
- the capsid protein further comprises one or more amino acid substitutions relative to VP1 capsid of AAV2 or one or more corresponding substitutions in another AAV serotype, preferably wherein the capsid protein further comprises a P34A amino acid substitution relative to VP1 capsid of AAV2 or the corresponding substitution in another AAV serotype.
- a method for delivering a heterologous nucleic acid comprising a nucleotide sequence encoding an antibody as herein described to a mammalian subject comprising administering to the mammal an effective amount of an rAAV as herein described or a pharmaceutical composition comprising same, preferably wherein the rAAV or pharmaceutical composition is administered by intravitreal injection.
- the heterologous nucleic acid is delivered to a retinal cell of the subject, e.g., a photoreceptor cell (e.g., rods; cones), a retinal ganglion cell (RGC), a glial cell (e.g., a Muller glial cell, a microglial cell), a bipolar cell, an amacrine cell, a horizontal cell, and/or a retinal pigmented epithelium (RPE) cell of the subject.
- a retinal cell of the subject e.g., a photoreceptor cell (e.g., rods; cones), a retinal ganglion cell (RGC), a glial cell (e.g., a Muller glial cell, a microglial cell), a bipolar cell, an amacrine cell, a horizontal cell, and/or a retinal pigmented epithelium (RPE) cell of the subject.
- a photoreceptor cell e.
- detectable plasma levels of the antibody are present in a subject for at least a week, at least two weeks, at least three weeks, at least a month, at least two months, or at least 6 months following administration of the rAAV to the subject.
- the rAAV is administered to a subject by intravitreal administration.
- a pharmaceutical composition comprising an rAAV as described herein and a pharmaceutically acceptable excipient.
- a method for treating a complement related disorder by delivering to the subject an rAAV as herein described or a pharmaceutical composition comprising the rAAV.
- Complement related disorders include, without limitation, membranoproliferative glomerulonephritis, atypical hemolytic uremic syndrome (aHUS), age related macular degeneration (AMD), geographic atrophy secondary to AMD microangiopathic haemolytic anemia, thrombocytopenia, acute renal failure, paroxysmal
- a method for treating dry age-related macular degeneration (AMD) e.g., late-stage dry AMD
- AMD dry age-related macular degeneration
- a pharmaceutical composition comprising the rAAV.
- the rAAV or pharmaceutical composition is administered to a subject for the treatment of geographic atrophy secondary to AMD.
- the rAAV or pharmaceutical composition is administered to the subject by intravitreal injection.
- Figures 1 A-C Induced pluripotent stem cell derived retinal pigmented epithelial cells (iPSC-RPE) were transduced with C3-N10 rAAV transgene (encoding anti-C3 antibody of SEQ ID NO: 14) at three multiplicities of infection, (MOI) 5,000, 10,000 and 20,000. A dose response was observed for nanobody (VHH) expression, (Fig. 1A), and MAC formation (Fig. IB).
- VHH nanobody
- Fig. IB MAC formation
- MAC complement membrane attack complex
- Figures 2A-C Complement Inhibition by Wild Type Lead Nanobodies in Alternative, Lectin & Classical Complement Pathways. All three wild type nanobodies’ complement inhibition activity were compared using Wieslab’ s alternative, MBL/lectin, and classical complement pathway assay kits. As shown in Figure 2A (alternative), Figure 2B (lectin) and Figure 2C (classical), while C5S2 has similarly strong inhibitory potency against all three pathways, C3S3 only exhibits activity against the alternative pathway. C3N10 appears to have stronger inhibitory affect towards the alternative pathway, milder potency against the lectin and classical pathways.
- Figures 4A-B IC50 potency of C3N10 and C3N10.3 against classical ( Figure 4A) and lectin ( Figure 4B) pathways.
- Figure 4A IC50 potency of C3N10 and C3N10.3 against classical ( Figure 4A) and lectin ( Figure 4B) pathways.
- Figure 4B IC50 potency of C3N10 and C3N10.3 against classical ( Figure 4A) and lectin ( Figure 4B) pathways.
- Figure 5 illustrates Cross Reactivity of Nanobody Wild Type Leads with Mouse Serum.
- Murine cross-inhibitory activity of the wild type nanobodies was examined using Hycult’s 03 Mouse ELISA Kit and the mouse-anti -mouse C3 antibody BB5.1 as control. None of the anti-human C3 or C5 nanobodies, C3N10, C3S3, nor C5S2 exhibited inhibitory activity against the mouse serum mediated alternative complement pathway.
- Figures 6A-B IC50 of wild type and humanized nanobodies against alternative complement pathway in cynomolgus serum. Both wild type and their humanized counterparts were serial diluted and their cynomolgus serum mediated alternative complement pathway inhibitory activity was calculated using the respective Wieslab assay.
- C3N10 WT and C3N10.3 Figure 6A
- C3S3 WT and C3S3.2 Figure 6B
- C5S2 and C5S2.2 did not exhibit any inhibitory activity in this NHP serum mediated assay (data not shown).
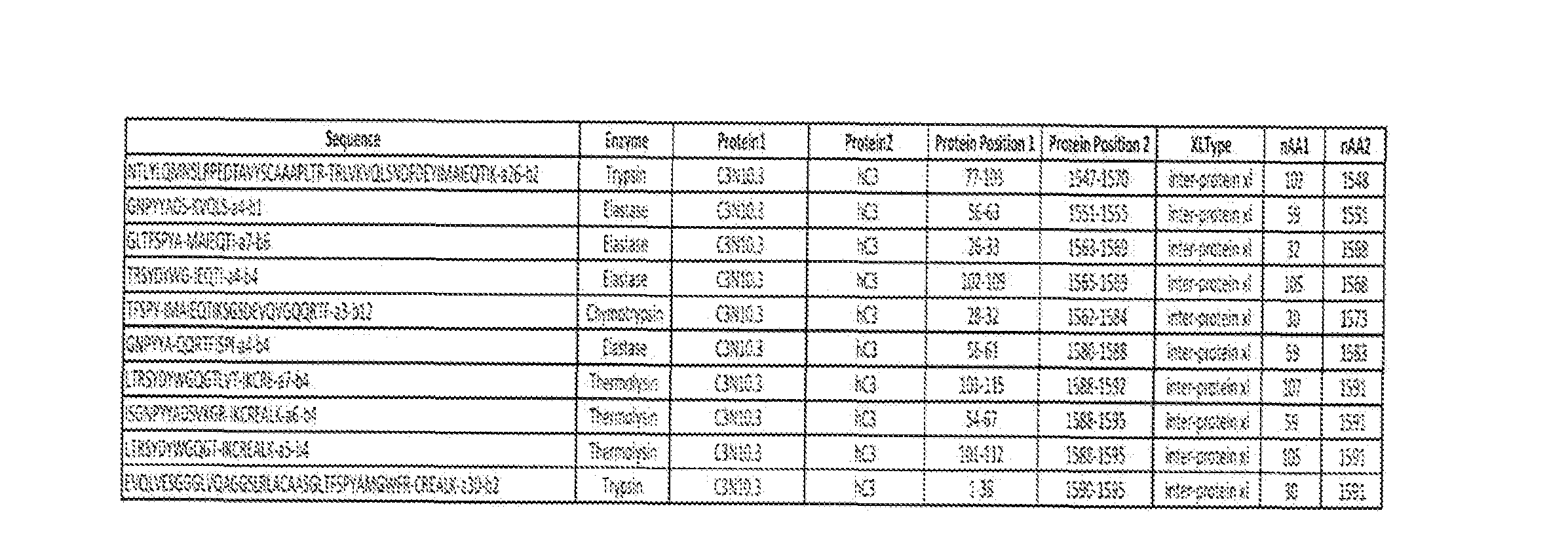
- Humanized 03 S3.2 was allowed to associate human 03 protein and cross-linking mass spectrometry (CovalX) was performed to identify potential binding peptide epitopes.
- An epitope between amino acids 110-130 was identified (amino acids 114-124) and another epitope between amino acids 490-510 (amino acids 497-502 and 502-505) of human C3 protein was identified.
- Figure 10 illustrates potential binding domains of C3N1O.3 and C3S3.2 to human 03 protein. Based upon the cross-linking mass spectrometry data, the binding regions of C3N10.3 and C3S3.2 to human C3 protein are shown, wherein the former associates within the alphachain’s C345C, while the latter interacts across both MG1 and MG5 domains of the beta-chain, respectively.
- FIG 11 shows Eculizumab and C5S2.2 cross block binding to human C5: Probe anti-human Fc Sandwich.
- the Gator/BLI machine was used to examine whether C5S2.2 cross blocks eculizumab, a clinically approved anti-human C5 targeting hIgG4 antibody.
- Anti-human Fc probes were pre-bound by eculizumab, then washed. Then, either human C5-alone or increasing concentrations of human C5 pre-bound with C5S2.2 were allowed to associate with the antibody-loaded probes. Binding kinetics was only observed with free C5 protein indicating that the pre-bound C5S2.2 VHH was blocking eculizumab’s C5 binding epitope. Time (sec) is on the X-axis and Shift (nm) is on the Y-axis.
- Figure 12 shows Eculizumab and C5S2.2 cross block binding to human 05: Probe anti-StepII sandwich.
- the Gator/BLI machine was used to examine whether C5S2.2 cross blocks eculizumab, a clinically approved anti-human C5 targeting h!gG4 antibody.
- Anti-Strep II probes were pre-bound with Strep-tagged anti-human C5 C5S2.2 VHH. After washing, the nanobody loaded probes were dipped into human C5 containing binding buffer and association kinetics was observed. When this pre-bound complex was then dipped into eculizumab containing solution, no interaction was seen, implicating that the latter’s binding epitope was blocked. Time (sec) is on the X-axis and Shift (nm) is on the Y-axis.
- Figures 13A-C show C5S2.2 binding to human and cynomolgus C5 mutants.
- Eculizumab is refractory to two naturally occurring variants: the R885H human polymorphism and the W917S human vs cynomolgus divergence.
- Human C5 R885H, human C5 W917S, and cynomolgus C5 S917W mutant proteins were recombinantly produced and C5S2.2’s affinity to them was compared to eculizumab ( Figures 14A-C) using the Gator.
- FIG. 14A-C shows Eculizumab binding to human and cynomolgus C5 mutants.
- Eculizumab is refractory to two naturally occurring variants: the R885H human polymorphism and the W917S human vs cynomolgus divergence.
- Human C5 R885H, human C5 W917S, and cynomolgus C5 S917W mutant proteins were recombinantly produced and eculizumab's affinity to them was reflected using the Gator
- Normal human donor serum was tested for ADA against the listed nanobodies and controls using meso-scale discovery immunoassays. Samples were diluted 1 : 100 and applied to plate wells passively coated with the respective antibodies. Bound human antibodies were detected using Sulfo-TAG goat-anti-human IgG and measured in a Meso Sector S 600.
- Figures 16A-C show functional assessment of VHH expressed following plasmid transfection by breakdown products of complement cascade associated proteins.
- Figure 16A C3 VHH inhibitors, but not C5 VHH inhibitors, prevent C3 cleavage as detected by absence of C3 a breakdown product.
- Figure 16B C3 VHH inhibitors and C5 VHH inhibitors prevent 05 cleavage as detected by absence of C5a breakdown product.
- Figure 16C 03 VHH inhibitors, but not C5 VHH inhibitors, prevent Bb cleavage as detected by absence of Factor B breakdown products, v, Version; h, humanized; kDa, kilodalton; NT, non-transfected.
- Figure 17 is a Western blot demonstrating a single product for each VHH variant examined following transduction with AAV carrying VHH.
- NT non-transduced
- kDa kilodalton
- MOI 20,000 33 ⁇ g protein loaded per lane; seven days post transduction.
- Figures 18A-C show expression of secreted VHH (C3N10 and C3N10 humanized, Figure ISA; C3S3 and humanized C3S3, Figure 18B; C5S2 and humanized C5S2, Figure 18C), following AAV transduction, as measured by ELISA and illustrating a dose response observed for each VHH (MOIs of 1,000, 5,000 and 20,000).
- NT non-transduced
- MOI multiplicity of infection
- n 3 wells/MOI.
- Figure 19 shows function of secreted VHH (C3N10, C3N10 h, C353, C353 h, C5S2, C5S2 h) as measured by complement inhibition represented as a percent. Each VHH exhibited some level of complement inhibition.
- NT non-transduced
- h humanized
- MOI multiplicity of infection
- error bars ⁇ standard deviation
- n 3 wells/MOI.
- an “antibody” is used herein in the broadest sense to refer to a polypeptide or protein with an immunoglobulin-like domain capable of recognizing and binding an antigen and includes full-size antibodies, individual chains thereof and all portions, domains or fragments thereof (including but not limited to, antigen binding domains or fragments such as VHH domains or VH/VL domains, respectively).
- the antigen binding site of an antibody preferably comprises at least one complementarity determining region (CDR).
- CDR complementarity determining region
- the present disclosure relates primarily to single variable domain on a heavy chain (VHH) antibodies.
- the terms “single domain antibody”, “single variable domain antibody”, “VHH antibody”, and “nanobody” have the same meaning referring to a variable region of a heavy chain of an antibody, and construct a single domain antibody consisting of only one heavy chain variable region.
- the antigen binding site of a single variable domain is formed by no more than three CDRs.
- the antibodies with a natural deficiency of the light chain and the heavy chain constant region 1 (CHI) are first obtained, the variable regions of the heavy chain of the antibody are therefore cloned to construct a single domain antibody (VHH) consisting of only one heavy chain variable region.
- the VHH antibody is preferably derived from llama.
- an “antigen” is a molecule comprising at least one epitope.
- the antigen may for example be a polypeptide, nucleic acid, polysaccharide, protein, lipoprotein or glycoprotein.
- a “complementarity determining region” or “CDR” is a hypervariable region of the antigen-binding region of an antibody.
- the CDRs are interspersed between regions that are more conserved, termed framework regions (FRs).
- FRs framework regions
- the antigen-binding region of an antibody may thus comprise one or more CDRs and FRs, usually in each variable domain three CDRs and four FRs arranged from amino-terminus to carboxy-terminus in the following order: FR1 , CDR1 , FR2, CDR2, FR3, CDR3, FR4.
- An “epitope” is a determinant capable of specific binding to an antibody.
- Epitopes may for example be comprised within polypeptides or proteins.
- Epitopes may be continuous or discontinuous, wherein a discontinuous epitope is a conformational epitope on an antigen which is formed from at least two separate regions in the primary sequence of the protein, nucleic acid or polysaccharide.
- affinity refers to the strength of binding between an antibody and its antigen.
- the affinity of an antibody can be defined in terms of the dissociation constant, KD, which is an equilibrium constant that measures the propensity of a molecular complex to separate (dissociate) reversibly into the molecules forming the complex.
- KD is defined as the ratio kotr / kon , where kotr and kon are the rate constants for association and dissociation of the molecular complex.
- affinity is determined by calculating the dissociation constant KD based on IC50 values. Thus, the affinity is measured as an apparent affinity.
- isolated designates a biological material (cell, nucleic acid or protein) that has been removed from its original environment (the environment in which it is naturally present). For example, a polynucleotide present in the natural state in a plant or an animal is not isolated, however the same polynucleotide separated from the adjacent nucleic acids in which it is naturally present, is considered “isolated.”
- a "coding region” or “coding sequence” is a portion of polynucleotide which consists of codons translatable into amino acids.
- a “stop codon” (TAG, TGA, or TAA) is typically not translated into an amino acid, it can be considered to be part of a coding region, but any flanking sequences, for example promoters, ribosome binding sites, transcriptional terminators, introns, and the like, are not part of a coding region.
- the boundaries of a coding region are typically determined by a start codon at the 5' terminus, encoding the
- Two or more coding regions can be present in a single polynucleotide construct, e.g., on a single vector, or in separate polynucleotide constructs, e.g., on separate (different) vectors. It follows, then that a single vector can contain just a single coding region, or comprise two or more coding regions.
- regulatory region refers to nucleotide sequences located upstream (5' non-coding sequences), within, or downstream (3' non-coding sequences) of a coding region, and which influence the transcription, RNA processing, stability, or translation of the associated coding region. Regulatory regions can include promoters, translation leader sequences, introns, polyadenylation recognition sequences, RNA processing sites, effector binding sites and stem-loop structures. If a coding region is intended for expression in a eukaryotic cell, a polyadenylation signal and transcription termination sequence will usually be located 3' to the coding sequence.
- nucleic acid is interchangeable with “polynucleotide” or “nucleic acid molecule” and a polymer of nucleotides is intended.
- a polynucleotide which encodes a gene product can include a promoter and/or other transcription or translation control elements operably associated with one or more coding regions.
- a coding region for a gene product e.g., a polypeptide
- a coding region and a promoter are "operably associated" if induction of promoter function results in the transcription of mRNA encoding the gene product encoded by the coding region, and if the nature of the linkage between the promoter and the coding region does not interfere with the ability of the promoter to direct the expression of the gene product or interfere with the ability of the DNA template to be transcribed.
- Other transcription control elements besides a promoter, for example enhancers, operators, repressors, and transcription termination signals, can also be operably associated with a coding region to direct gene product expression.
- Transcriptional control sequences refer to DNA regulatory sequences, such as promoters, enhancers, terminators, and the like, that provide for the expression of a coding
- transcription control regions are known to those skilled in the art. These include, without limitation, transcription control regions which function in vertebrate cells, such as, but not limited to, promoter and enhancer segments from cytomegaloviruses (the immediate early promoter, in conjunction with intron-A), simian virus 40 (the early promoter), and retroviruses (such as Rous sarcoma virus). Other transcription control regions include those derived from vertebrate genes such as actin, heat shock protein, bovine growth hormone and rabbit beta-globin, as well as other sequences capable of controlling gene expression in eukaryotic cells. Additional suitable transcription control regions include tissuespecific promoters and enhancers as well as lymphokine-inducible promoters (e.g., promoters inducible by interferons or interleukins).
- tissuespecific promoters and enhancers as well as lymphokine-inducible promoters (e.g., promoters inducible by interferons or interleukins).
- translation control elements include, but are not limited to ribosome binding sites, translation initiation and termination codons, and elements derived from picomaviruses (particularly an internal ribosome entry site, or IRES, also referred to as a CITE sequence).
- RNA messenger RNA
- tRNA transfer RNA
- shRNA small hairpin RNA
- siRNA small interfering RNA
- expression produces a "gene product.”
- a gene product can be either a nucleic acid, e.g., a messenger RNA produced by transcription of a gene, or a polypeptide which is translated from a transcript.
- Gene products described herein further include nucleic acids with post transcriptional modifications, e.g., polyadenylation or splicing, or polypeptides with post translational modifications, e.g., methylation, glycosylation, the addition of lipids, association with other protein subunits, or proteolytic cleavage.
- post transcriptional modifications e.g., polyadenylation or splicing
- polypeptides with post translational modifications e.g., methylation, glycosylation, the addition of lipids, association with other protein subunits, or proteolytic cleavage.
- Promoter and “promoter sequence” are used interchangeably and refer to a DNA sequence capable of controlling the expression of a coding sequence or functional RNA.
- a coding sequence is located 3' to a promoter sequence. Promoters can be derived in their entirety from a native gene, or be composed of different elements derived from different
- promoters found in nature, or even comprise synthetic DNA segments. It is understood by those skilled in the art that different promoters can direct the expression of a gene in different tissues or cell types, or at different stages of development, or in response to different environmental or physiological conditions. Promoters that cause a gene to be expressed in most cell types at most times are commonly referred to as “constitutive promoters.” Promoters that cause a gene to be expressed in a specific cell type are commonly referred to as “cell-specific promoters” or “tissuespecific promoters.” Promoters that cause a gene to be expressed at a specific stage of development or cell differentiation are commonly referred to as “developmentally-specific promoters” or “cell differentiation-specific promoters.” Promoters that are induced and cause a gene to be expressed following exposure or treatment of the cell with an agent, biological molecule, chemical, ligand, light, or the like that induces the promoter are commonly referred to as “inducible promoters” or “regulatable promoters.” It is further recognized that since in
- Plasmid refers to an extra-chromosomal element often carrying a gene that is not part of the central metabolism of the cell, and usually in the form of circular doublestranded DNA molecules.
- Such elements can be autonomously replicating sequences, genome integrating sequences, phage or nucleotide sequences, linear, circular, or supercoiled, of a single- or double-stranded DNA or RNA, derived from any source, in which a number of nucleotide sequences have been joined or recombined into a unique construction which is capable of introducing a promoter fragment and DNA sequence for a selected gene product along with appropriate 3' untranslated sequence into a cell.
- a polynucleotide or polypeptide has a certain percent "sequence identity" to another polynucleotide or polypeptide, meaning that, when aligned, that percentage of bases or amino acids are the same when comparing the two sequences.
- sequence identity is related to sequence homology. Homology comparisons may be conducted by eye, or more usually, with the aid of readily available sequence comparison programs. Sequence similarity or sequence homology can be determined in a number of different manners. Commercially available computer programs may calculate percent (%) homology between two or more sequences and may also calculate the sequence identity shared by two or more amino acids
- Sequence homologies may be generated by any of a number of computer programs known in the art, including BLAST, available over the world wide web at ncbi.nlm.nih.gov/BLAST/. Another alignment algorithm is PASTA, available in the Genetics Computing Group (GCG) package, from Madison, Wis., USA. Other techniques for alignment are described in Methods in Enzymology, vol. 266: Computer Methods for Macromolecular Sequence Analysis (1996), ed. Doolittle, Academic Press, Inc. Of particular interest are alignment programs that permit gaps in the sequence. The Smith- Waterman is one type of algorithm that permits gaps in sequence alignments. See Meth. Mol. Biol. 70: 173-187 (1997). Also, the GAP program using the Needleman and Wunsch alignment method can be utilized to align sequences. See J. Mol. Biol. 48: 443-453 (1970).
- % homology may be calculated over contiguous sequences, i.e., one sequence is aligned with the other sequence and each amino acid or nucleotide in one sequence is directly compared with the corresponding amino acid or nucleotide in the other sequence, one residue at a time. This is called an “ungapped” alignment. Typically, such ungapped alignments are performed only over a relatively short number of residues.
- High gap penalties may, of course, produce optimized alignments with fewer gaps. Most alignment programs allow the gap penalties to be modified. However, it is preferred to use the default values when using such software for sequence comparisons. For example, when using the GCG Wisconsin Bestfit package the default gap penalty for amino acid sequences is -12 for a gap and -4 for each extension.
- BLAST and FASTA are available for offline and online searching (see Ausubel et al., 1999, Short Protocols in Molecular Biology, pages 7-58 to 7-60). However, for some applications, it is preferred to use the GCG Bestfit program.
- a new tool, called BLAST 2 Sequences is also available for comparing protein and nucleotide sequences (see FEMS Microbiol Lett. 1999 174(2): 247-50; FEMS Microbiol Lett. 1999 177(1): 187-8 and the website of the National Center for Biotechnology information at the website of the National Institutes for Health).
- the final % homology may be measured in terms of identity, the alignment process itself is typically not based on an all-or-nothing pair comparison. Instead, a scaled similarity score matrix is generally used that assigns scores to each pair-wise comparison based on chemical similarity or evolutionary distance.
- An example of such a matrix commonly used is the BLOSUM62 matrix — the default matrix for the BLAST suite of programs. GCG Wisconsin programs generally use either the public default values or a custom symbol comparison table, if supplied (see user manual for further details). For some applications, it is preferred to use the public default values for the GCG package, or in the case of other software, the default matrix, such as BLOSUM62.
- percentage homologies may be calculated using the multiple alignment feature in DNASISTM (Hitachi Software), based on an algorithm, analogous to CLUSTAL
- sequences may also have deletions, insertions or substitutions of amino acid residues which produce a silent change and result in a functionally equivalent substance.
- Deliberate amino acid substitutions may be made on the basis of similarity in amino acid properties (such as polarity, charge, solubility, hydrophobicity, hydrophilicity, and/or the amphipathic nature of the residues) and it is therefore useful to group amino acids together in functional groups.
- Amino acids may be grouped together based on the properties of their side chains alone. However, it is more useful to include mutation data as well.
- the sets of amino acids thus derived are likely to be conserved for structural reasons. These sets may be described in the form of a Venn diagram (Livingstone C. D. and Barton G.
- Embodiments of the invention include sequences (both polynucleotide or polypeptide) which may comprise homologous substitution (substitution and replacement are both used herein to mean the interchange of an existing amino acid residue or nucleotide, with an alternative residue or nucleotide) that may occur i.e., like-for-like substitution in the case of amino acids such as basic for basic, acidic for acidic, polar for polar, etc.
- Non-homologous substitution may also occur i.e., from one class of residue to another or alternatively involving the inclusion of unnatural amino acids such as ornithine (hereinafter referred to as Z), diaminobutyric acid
- B 16 ornithine (hereinafter referred to as B), norleucine ornithine (hereinafter referred to as 0), pyriylalanine, thienylalanine, naphthylalanine and phenylglycine.
- Variant amino acid sequences may include suitable spacer groups that may be inserted between any two amino acid residues of the sequence including alkyl groups such as methyl, ethyl or propyl groups in addition to amino acid spacers such as glycine or P-alanine residues.
- alkyl groups such as methyl, ethyl or propyl groups
- amino acid spacers such as glycine or P-alanine residues.
- a further form of variation which involves the presence of one or more amino acid residues in peptoid form, may be well understood by those skilled in the art.
- the peptoid form is used to refer to variant amino acid residues wherein the a-carbon substituent group is on the residue's nitrogen atom rather than the a-carbon.
- amino acid substitution and its synonyms described above are intended to encompass modification of an amino acid sequence by replacement of an amino acid with another, substituting, amino acid.
- the substitution may be a conservative substitution. It may also be a non-conservative substitution.
- conservative in referring to two amino acids, is intended to mean that the amino acids share a common property recognized by one of skill in the art. For example, amino acids having hydrophobic nonacidic side chains, amino acids having hydrophobic acidic side chains, amino acids having hydrophilic nonacidic side chains, amino acids having hydrophilic acidic side chains, and amino acids having hydrophilic basic side chains.
- Common properties may also be amino acids having hydrophobic side chains, amino acids having aliphatic hydrophobic side chains, amino acids having aromatic hydrophobic side chains, amino acids with polar neutral side chains, amino acids with electrically charged side chains, amino acids with electrically charged acidic side chains, and amino acids with electrically charged basic side chains.
- Both naturally occurring and non-naturally occurring amino acids are known in the art and may be used as substituting amino acids in embodiments.
- Methods for replacing an amino acid are well known to the skilled in the art and include, but are not limited to, mutations of the nucleotide sequence encoding the amino acid sequence. Reference to "one or more" herein is intended to encompass the individual embodiments of, for example, 1, 2, 3, 4, 5, 6, or more.
- treatment, 11 refers to obtaining a desired pharmacologic and/or physiologic effect.
- the effect may be prophylactic in terms of completely or partially preventing a disease or symptom thereof and/or may be therapeutic in terms of a partial or complete cure for a disease and/or adverse effect attributable to the disease.
- Treatment covers any treatment of a disease in a mammal, particularly in a human, and includes: (a) preventing the disease (and/or symptoms caused by the disease) from occurring in a subject which may be predisposed to the disease or at risk of acquiring the disease but has not yet been diagnosed as having it; (b) inhibiting the disease (and/or symptoms caused by the disease), i.e., arresting its development; and (c) relieving the disease (and/or symptoms caused by the disease), i.e., causing regression of the disease (and/or symptoms caused by the disease), i.e., ameliorating the disease and/or one or more symptoms of the disease.
- treating complement factor H disorders may encompass alleviating, reducing, and/or ameliorating symptoms, and/or preventing the development of additional symptoms associated with complement factor H disorder, which can manifest as several different phenotypes, including asymptomatic, recurrent bacterial infections, and renal failure. This is typically characterized by decreased serum levels of factor H, complement component C3, and a decrease in other terminal complement components, indicating activation of the alternative complement pathway. Serum levels of complement component C5 may also be decreased. This disorder is associated with a number of renal diseases with variable clinical presentation and progression, including C3 glomerulopathy and atypical hemolytic uremic syndrome.
- treating complement associated disorders includes alleviating, reducing, and/or ameliorating symptoms, both of the complement factor H disorders identified above, but also other disorders associated with uncontrolled alternative pathway complement regulation.
- Complement-mediated disorders may encompass symptoms associated with complement dysregulation which can manifest as several different phenotypes, including asymptomatic, recurrent bacterial infections, and various tissue injuries including but not limited to renal diseases. Unless otherwise specified, both homozygous subjects and heterozygous subjects are encompassed within this definition. Complement dysregulation is typically caused by loss of function mutations in, or auto-antibodies against, complement regulatory proteins including but not limited to fH, factor I (fl) and membrane cofactor protein (MCP) or by gain of function mutations in other complement proteins including but not limited to C3, C5 and factor B (fB).
- complement regulatory proteins including but not limited to fH, factor I (fl) and membrane cofactor protein (MCP)
- MCP membrane cofactor protein
- Complement dysregulation is typically, though not always, characterized by decreased serum levels of factor H, complement component C3, fB and a decrease in other terminal complement components, indicating activation of the alternative and/or the terminal complement pathway.
- Complement- mediated pathologies that can be treated by the present invention of composition and method include but are not limited to the following diseases with variable clinical presentation and progression: C3 glomerulopathy (formally called membranoproliferative glomerulonephritis type II or MPGNII), of which there are two known forms - dense deposit disease (DDD) and C3 glomerulonephritis (C3GN); thrombotic microangiopathy (TMA) including but not limited to atypical hemolytic uremic syndrome (aHUS), Shiga-like toxin-producing E.
- C3 glomerulopathy (formally called membranoproliferative glomerulonephritis type II or MPGNII), of which there are two known forms - dense deposit disease (DDD) and C3
- thrombotic thrombocytopenia purpura TTP
- retinal degenerative eye disease including age related macular degeneration (AMD), RPE degeneration, chorioretinal degeneration, photoreceptor degeneration, paroxysmal noctural hemoglobinuria (PNH), ischemia reperfusion injury of all organs and settings, rheumatoid arthritis, hemodialysis, diabetic nephropathy, diabetic vasculopathy, asthma, systemic lupus erythematosus (SLE), ischemic stroke, abdominal aortic aneurysm (AAA), antineutrophil cytoplasmic antibody (ANCA) mediated vasculitis (ANCA vasculitis), ANCA- mediated hemorrhagic lung injury and disease, ANCA glomerulonephritis, graft versus host disease (GvHD), acute or delay graft rejection in organ transplantation, Crohn's disease, p
- NMO neuromyelitis optica
- BP Bullous pemphigoid
- AD Alzheimer's disease
- bacterial infections caused by recruitment of bacterial pathogens (e.g., Aspergillus spp.; Borrelia burgdorferi; B. duttonii; B. recurrentis; Candida albicans; Fr and sella tularensis; Haemophilus influenzae; Neisseria meningitidis;
- the terms "individual,” “host,” “subject,” and “patient” are used interchangeably herein, and refer to a mammal, including, but not limited to, primates (e.g., humans; non- human primates, including simians); mammalian sport animals (e.g., horses); mammalian farm animals (e.g., sheep, goats, etc.); mammalian pets (dogs, cats, etc.); and rodents (e.g., mice, rats, etc.).
- primates e.g., humans; non- human primates, including simians
- mammalian sport animals e.g., horses
- mammalian farm animals e.g., sheep, goats, etc.
- mammalian pets dogs, cats, etc.
- rodents e.g., mice, rats, etc.
- an effective amount is an amount sufficient to effect beneficial or desired clinical results.
- An effective amount can be administered in one or more administrations.
- an effective amount of a compound e.g., an infectious rAAV virion
- an effective amount of an infectious rAAV virion is an amount that is sufficient to palliate, ameliorate, stabilize, reverse, prevent, slow or delay the progression of (and/or symptoms associated with) a particular disease state (e.g., a disorder associated with complement dysfunction).
- an effective amount of an infectious rAAV virion is an amount of the infectious rAAV virion that is able to effectively deliver a heterologous nucleic acid to a target cell (or target cells) of the individual.
- Effective amounts may be determined preclinically by, e.g., detecting in the cell or tissue the gene product (RNA, protein) that is encoded by the heterologous nucleic acid sequence using techniques that are well understood in the art, e.g. RT-PCR, western blotting, ELISA, fluorescence or other reporter readouts, and the like. Effective amounts may be determined clinically by, e.g. detecting a change in the onset or progression of disease using methods known in the art, e.g. 6-minute walk test, left ventricular ejection fraction, hand-held dynamometry, Vignos Scale and the like as described herein and as known in the art.
- Novel anti-C3 and anti-C5 antibodies and nucleic acids encoding same are described herein.
- rAAV virions comprising a nucleic acid encoding the anti-C3 and/or antibodies are also described. These rAAV virions are characterized by durable and robust expression of anti-C3
- Delivery of these rAAV virions to subjects in need thereof may be achieved via a number of routes, preferably by intravitreal administration. Also provided are methods of using these rAAV virions in regimens for treating complement factor C3- and C5-associated disorders, particularly dry AMD and geographic atrophy secondary to AMD.
- Naturally occurring human antibodies are heterotetramers.
- the antibodies provided herein in one aspect comprise an antigen binding site in a single polypeptide.
- the antibodies are therefore herein referred to as “single variable domain” or “VHH” antibodies.
- Single variable domain antibodies are also known as nanobodies.
- the single variable domain antibodies disclosed herein may, though, in certain embodiments, be bispecific or multispecific single variable domain antibodies, in which single variable domain antibodies are coupled.
- a single variable domain antibody is an antibody fragment consisting of a single monomeric variable antibody domain. Like a whole antibody, it is able to bind selectively to a specific antigen.
- Single variable domain antibodies typically have molecular weights in the range of 12-15 kDa, i.e. much lower than common antibodies, ranging typically from 150 to 160 kDa.
- Single variable domain antibodies are also smaller than Fab fragments (-50 kDa) of heterotetrameric antibodies comprising one light chain and half a heavy chain.
- Single domain antibodies can derive from antibodies found in nature, for example in camelids (VHH) and cartilaginous fishes (VNAR).
- VHH camelids
- VNAR cartilaginous fishes
- New or Nurse Shark Antigen Receptor (NAR) protein exists as a dimer of two heavy chains with no associated light chains. Each chain is composed of one variable (V) and five constant domains. The NAR proteins thus constitute a single immunoglobulin variable-like domain.
- Single heavy- chain antibodies are also found in camelids, such as such as dromedaries, camels, llamas and alpacas, where the heavy chain has lost one of its constant domains and underwent modifications in the variable domain, both of which are structural elements necessary for the binding of light chains.
- single variable domain antibodies can also be engineered by recombinant methods.
- One approach is to split the dimeric variable domains from common immunoglobulin G (IgG) from humans or mice into monomers.
- Single domains which are derived from light chains, also bind specifically to target epitopes.
- the single variable domain antibody may be derived from any suitable organism.
- Single domain camelid antibodies are equal to regular antibodies in terms of specificity. Single domain antibodies are easily isolated, for example by using phage panning procedures. The smaller size and single domain architecture make these antibodies easier to express as proteins in bacterial cells for large scale production, making them ideal for commercial exploitation.
- the antibodies of the present invention are therefore single variable domain antibodies, preferably derived from camelid antibodies, preferably llama antibodies, including functional homologs, fragments thereof and fusion macromolecules containing VHH covalently linked to glycan, nucleic acid, protein, or chemical groups not being a macromolecule.
- an antibody provided herein is an antibody fragment.
- the antibody fragment is a Fab, Fab', Fab'-SH, or F(ab')2 fragment, in particular a Fab fragment.
- Papain digestion of intact antibodies produces two identical antigen-binding fragments, called “Fab” fragments containing each the heavy- and light-chain variable domains (VH and VL, respectively) and also the constant domain of the light chain (CL) and the first constant domain of the heavy chain (CHI).
- Fab fragment thus refers to an antibody fragment comprising a light chain comprising a VL domain and a CL domain, and a heavy chain fragment comprising a VH domain and a CHI domain.
- Fab' fragments differ from Fab fragments by the addition of residues at the carboxy terminus of the CHI domain including one or more cysteines from the antibody hinge region.
- Fab'-SH are Fab' fragments in which the cysteine residue(s) of the constant domains bear a free thiol group.
- Pepsin treatment yields an F(ab')2 fragment that has two antigen-binding sites (two Fab fragments) and a part of the Fc region.
- Antibody fragments can be made by various techniques, including but not limited to proteolytic digestion of an intact antibody as well as recombinant production by recombinant host cells (e.g., E. coli, CHO).
- recombinant host cells e.g., E. coli, CHO.
- the antibody provided herein is a Fab fragment.
- the VH domain of the antibody provided herein comprises a human VH3 framework.
- the VL domain of the antibody provided herein comprises a human Vkappal framework.
- the CL domain of the antibody provided herein is of kappa isotype.
- the CHI domain of the antibody provided herein is of human IgGl isotype.
- the antibody provided herein is a Fab fragment comprising a CL domain of kappa isotype and a CHI domain of human IgGl isotype.
- an antibody provided herein is a multispecific antibody.
- Multispecific antibodies are monoclonal antibodies that have binding specificities for at least two different sites, i.e., different epitopes on different antigens or different epitopes on the same antigen. In certain aspects, the multispecific antibody has three or more binding specificities.
- Multispecific antibodies with three or more binding specificities comprising antibodies provided herein may be provided in an asymmetric form with a domain crossover in one or more binding arms of the same antigen specificity, i.e. by exchanging the VH/VL domains (see e.g., WO 2009/080252 and WO 2015/150447), the CH1/CL domains (see e.g., WO 2009/080253) or the complete Fab arms (see e.g., WO 2009/080251, WO 2016/016299, also see Schaefer et al, PNAS, 108 (2011) 1187-1191, and Klein at al cache MAbs 8 (2016) 1010-20).
- Various further molecular formats for multispecific antibodies are known in the art and are included herein (see e.g., Spiess et al., Mol Immunol 67 (2015) 95-106).
- amino acid sequence variants of the antibodies provided herein are contemplated. For example, it may be desirable to alter the binding affinity and/or other biological properties of the antibody.
- Amino acid sequence variants of an antibody may be prepared by introducing appropriate modifications into the nucleotide sequence encoding the antibody, or by peptide synthesis. Such modifications include, for example, deletions from,
- antibody variants having one or more amino acid substitutions are provided.
- Sites of interest for substitutional mutagenesis include the CDRs and FRs.
- amino acid side chain classes amino acids side chain classes.
- Amino acid substitutions may be introduced into an antibody of interest and the products screened for a desired activity, e.g., retained/improved antigen binding, decreased immunogenicity, or improved ADCC or CDC.
- Amino acids may be grouped according to common side-chain properties:
- Non-conservative substitutions will entail exchanging a member of one of these classes for a member of another class.
- substitutional variant involves substituting one or more CDR residues of a parent antibody (e.g., a humanized or human antibody).
- a parent antibody e.g., a humanized or human antibody.
- the resulting variant(s) selected for further study will have modifications (e.g., improvements) in certain biological properties (e.g., increased affinity, reduced immunogenicity) relative to the parent antibody and/or will have substantially retained certain biological properties of the parent antibody.
- An exemplary substitutional variant is an affinity matured antibody, which may be conveniently generated, e.g., using phage display-based affinity maturation techniques such as those described herein. Briefly, one or more CDR residues are mutated and the variant antibodies displayed on phage and screened for a particular biological activity (e.g., binding affinity).
- substitutions, insertions, or deletions may occur within one or more CDRs so long as such alterations do not substantially reduce the ability of the antibody to bind antigen.
- conservative alterations e.g., conservative substitutions as provided herein
- that do not substantially reduce binding affinity may be made in the CDRs.
- 25 alterations may, for example, be outside of antigen contacting residues in the CDRs.
- each CDR either is unaltered, or contains no more than one, two or three amino acid substitutions.
- a useful method for identification of residues or regions of an antibody that may be targeted for mutagenesis is called “alanine scanning mutagenesis” as described by Cunningham and Wells (1989) Science, 244:1081-1085.
- a residue or group of target residues e.g., charged residues such as arg, asp, his, lys, and glu
- a neutral or negatively charged amino acid e.g., alanine or polyalanine
- a crystal structure of an antigen-antibody complex may be used to identify contact points between the antibody and antigen. Such contact residues and neighboring residues may be targeted or eliminated as candidates for substitution. Variants may be screened to determine whether they contain the desired properties.
- Amino acid sequence insertions include amino- and/or carboxyl-terminal fusions ranging in length from one residue to polypeptides containing a hundred or more residues, as well as intrasequence insertions of single or multiple amino acid residues.
- terminal insertions include an antibody with an N-terminal methionyl residue.
- Other insertional variants of the antibody molecule include the fusion to the N- or C-terminus of the antibody to an enzyme (e.g., for ADEPT (antibody directed enzyme prodrug therapy)) or a polypeptide which increases the serum half-life of the antibody.
- an antibody provided herein is altered to increase or decrease the extent to which the antibody is glycosylated.
- Addition or deletion of glycosylation sites to an antibody may be conveniently accomplished by altering the amino acid sequence such that one or more glycosylation sites is created or removed.
- the oligosaccharide attached thereto may be altered.
- Native antibodies produced by mammalian cells typically comprise a branched,
- the oligosaccharide may include various carbohydrates, e.g., mannose, N-acetyl glucosamine (GlcNAc), galactose, and sialic acid, as well as a fucose attached to a GlcNAc in the “stem” of the biantennary oligosaccharide structure.
- modifications of the oligosaccharide in an antibody of the invention may be made in order to create antibody variants with certain improved properties.
- antibody variants having a non-fucosylated oligosaccharide, i.e. an oligosaccharide structure that lacks fucose attached (directly or indirectly) to an Fc region.
- a non-fucosylated oligosaccharide also referred to as “afucosylated” oligosaccharide
- Such non-fucosylated oligosaccharide particularly is an N-linked oligosaccharide which lacks a fucose residue attached to the first GlcNAc in the stem of the biantennary oligosaccharide structure.
- antibody variants having an increased proportion of non-fucosylated oligosaccharides in the Fc region as compared to a native or parent antibody.
- the proportion of non-fucosylated oligosaccharides may be at least about 20%, at least about 40%, at least about 60%, at least about 80%, or even about 100% (i.e. no fucosylated oligosaccharides are present).
- the percentage of non-fucosylated oligosaccharides is the (average) amount of oligosaccharides lacking fucose residues, relative to the sum of all oligosaccharides attached to Asn 297 (e. g.
- Asn297 refers to the asparagine residue located at about position 297 in the Fc region (EU numbering of Fc region residues); however, Asn297 may also be located about ⁇ 3 amino acids upstream or downstream of position 297, i.e., between positions 294 and 300, due to minor sequence variations in antibodies.
- Such antibodies having an increased proportion of non-fucosylated oligosaccharides in the Fc region may have improved FcyRIIIa receptor binding and/or improved effector function, in particular improved ADCC function. See, e.g., US 2003/0157108; US 2004/0093621.
- Examples of cell lines capable of producing antibodies with reduced fucosylation include Lee 13 CHO cells deficient in protein fucosylation (Ripka et al. Arch. Biochem. Biophys. 249:533-545 (1986); US 2003/0157108; and WO 2004/056312, especially at Example 11), and knockout cell lines, such as alpha- 1,6-fucosyltransferase gene, FUT8, knockout CHO cells (see, e.g., Yamane-Ohnuki et al. Biotech. Bioeng. 87:614-622 (2004); Kanda, Y. et al., Biotechnol.
- antibody variants are provided with bisected oligosaccharides, e.g., in which a biantennary oligosaccharide attached to the Fc region of the antibody is bisected by GlcNAc.
- Such antibody variants may have reduced fucosylation and/or improved ADCC function as described above. Examples of such antibody variants are described, e.g., in Umana et al., Nat Biotechnol 17, 176-180 (1999); Ferrara et al., Biotechn Bioeng 93, 851-861 (2006); WO 99/54342; WO 2004/065540, WO 2003/011878.
- Antibody variants with at least one galactose residue in the oligosaccharide attached to the Fc region are also provided. Such antibody variants may have improved CDC function. Such antibody variants are described, e.g., in WO 1997/30087; WO 1998/58964; and WO 1999/22764.
- one or more amino acid modifications may be introduced into the Fc region of an antibody provided herein, thereby generating an Fc region variant.
- the Fc region variant may comprise a human Fc region sequence (e.g., a human IgGl, IgG2, IgG3 or IgG4 Fc region) comprising an amino acid modification (e.g., a substitution) at one or more amino acid positions.
- the invention contemplates an antibody variant that possesses some but not all effector functions, which make it a desirable candidate for applications in which the half life of the antibody in vivo is important yet certain effector functions (such as complementdependent cytotoxicity (CDC) and antibody-dependent cell-mediated cytotoxicity (ADCC)) are unnecessary or deleterious.
- In vitro and/or in vivo cytotoxicity assays can be conducted to confirm the reduction/depletion of CDC and/or ADCC activities.
- Fc receptor (FcR) binding assays can be conducted to ensure that the antibody lacks FcyR binding (hence likely lacking ADCC activity), but retains FcRn binding ability.
- the primary cells for mediating ADCC, NK cells express FcyRIII only, whereas monocytes express FcyRI, FcyRII and FcyRIII.
- non-radioactive assays methods may be employed (see, for example, ACTITM non-radioactive cytotoxicity assay for flow cytometry (CellTechnology, Inc. Mountain View, CA; and CytoTox 96 ⁇ non-radioactive cytotoxicity assay (Promega, Madison, WI).
- Useful effector cells for such assays include peripheral blood mononuclear cells (PBMC) and Natural Killer (NK) cells.
- PBMC peripheral blood mononuclear cells
- NK Natural Killer
- ADCC activity of the molecule of interest may be assessed in vivo, e.g., in a animal model such as that disclosed in Clynes et al. Proc. Nat'l Acad. Sci.
- Clq binding assays may also be carried out to confirm that the antibody is unable to bind Clq and hence lacks CDC activity. See, e.g., Clq and C3c binding ELISA in WO 2006/029879 and WO 2005/100402.
- a CDC assay may be performed (see, for example, Gazzano- Santoro et al., J. Immunol. Methods 202:163 (1996); Cragg, M. S. et al., Blood 101:1045-1052 (2003); and Cragg, M. S. and M. J. Glennie, Blood 103:2738-2743 (2004)).
- FcRn binding and in vivo clearance/half life determinations can also be performed using methods known in the art (see, e.g., Petkova, S. B. et al., Inti Immunol. 18(12): 1759-1769 (2006); WO 2013/120929 Al).
- Antibodies with reduced effector function include those with substitution of one or more of Fc region residues 238, 265, 269, 270, 297, 327 and 329 (U.S. Pat. No. 6,737,056).
- Fc mutants include Fc mutants with substitutions at two or more of amino acid positions 265, 269, 270, 297 and 327, including the so-called “DANA” Fc mutant with substitution of residues 265 and 297 to alanine (U.S. Pat. No. 7,332,581).
- an antibody variant comprises an Fc region with one or more amino acid substitutions which improve ADCC, e.g., substitutions at positions 298, 333, and/or 334 of the Fc region (EU numbering of residues).
- an antibody variant comprises an Fc region with one or more amino acid substitutions which diminish FcyR binding, e.g., substitutions at positions 234 and 235 of the Fc region (EU numbering of residues).
- the substitutions are L234A and L235A (LALA).
- the antibody variant further comprises D265A and/or P329G in an Fc region derived from a human IgGlFc region.
- the substitutions are L234A, L235A and P329G (LALA-PG) in an Fc region derived from a human IgGl Fc region. (See, e.g., WO 2012/130831).
- the substitutions are L234A, L235A and D265A (LALA- DA) in an Fc region derived from a human IgGI Fc region.
- alterations are made in the Fc region that result in altered (i.e., either improved or diminished) Cl q binding and/or Complement Dependent Cytotoxicity (CDC), e.g., as described in U.S. Pat. No. 6,194,551, WO 99/51642, and Idusogie et al. J. Immunol. 164: 4178-4184 (2000).
- CDC Complement Dependent Cytotoxicity
- Such Fc variants include those with substitutions at one or more of Fc region residues: 238, 252, 254, 256, 265, 272, 286, 303, 305, 307, 311, 312, 317, 340, 356, 360, 362, 376, 378, 380, 382, 413, 424 or 434, e.g., substitution of Fc region residue 434 (See, e.g., U.S. Pat. No. 7,371,826; Dall'Acqua, W. F., et al. J. Biol. Chem. 281 (2006) 23514-23524).
- Fc region residues critical to the mouse Fc-mouse FcRn interaction have been identified by site-directed mutagenesis (see e.g. Dall'Acqua, W. F., et al. J. Immunol 169 (2002) 5171-5180).
- Residues 1253, H310, H433, N434, and H435 are involved in the interaction (Medesan, C., et al., Eur. J. Immunol. 26 (1996) 2533; Firan, M., et al., Int. Immunol. 13 (2001) 993; Kim, J. K., et al., Eur. J. Immunol. 24 (1994) 542).
- an antibody variant comprises an Fc region with one or more amino acid substitutions, which reduce FcRn binding, e.g., substitutions at positions 253, and/or 310, and/or 435 of the Fc-region (EU numbering of residues).
- the antibody variant comprises an Fc region with the amino acid substitutions at positions 253, 310 and 435.
- the substitutions are 1253 A, H310A and H435A in an Fc region derived from a human IgGl Fc-region. See, e.g., Grevys, A., et al., J. Immunol. 194 (2015) 5497-5508.
- an antibody variant comprises an Fc region with one or more amino acid substitutions, which reduce FcRn binding, e.g., substitutions at positions 310, and/or 433, and/or 436 of the Fc region (EU numbering of residues).
- the antibody variant comprises an Fc region with the amino acid substitutions at positions 310, 433 and 436.
- the substitutions are H310A, H433A and Y436A in an Fc region derived from a human IgGl Fc-region. (See, e.g., WO 2014/177460 Al).
- an antibody variant comprises an Fc region with one or more amino acid substitutions which increase FcRn binding, e.g., substitutions at positions 252, and/or 254, and/or 256 of the Fc region (EU numbering of residues).
- the antibody variant comprises an Fc region with amino acid substitutions at positions 252, 254, and 256.
- the substitutions are M252Y, S254T and T256E in an Fc region derived from a human IgGl Fc-region. See also Duncan & Winter, Nature 322:738-40 (1988); U.S. Pat. Nos.
- the C-terminus of the heavy chain of the antibody as reported herein can be a complete C-terminus ending with the amino acid residues PGK.
- the C-terminus of the heavy chain can be a shortened C-terminus in which one or two of the C terminal amino acid residues
- an antibody comprising a heavy chain including a C-terminal CH3 domain as specified herein comprises the C-terminal glycine-lysine dipeptide (G446 and K447, EU index numbering of amino acid positions).
- an antibody comprising a heavy chain including a C- terminal CH3 domain, as specified herein comprises a C-terminal glycine residue (G446, EU index numbering of amino acid positions).
- cysteine engineered antibodies e.g., THIOMABTM antibodies
- the substituted residues occur at accessible sites of the antibody.
- reactive thiol groups are thereby positioned at accessible sites of the antibody and may be used to conjugate the antibody to other moieties, such as drug moieties or linker-drug moieties, to create an immunoconjugate, as described further herein.
- Cysteine engineered antibodies may be generated as described, e.g., in U.S. Pat. Nos. 7,521,541, 8,30,930, 7,855,275, 9,000,130, or WO 2016040856.
- the invention also provides immunoconjugates comprising an antibody provided herein conjugated (chemically bonded) to one or more agents; in one embodiment such as cytotoxic agents, chemotherapeutic agents, drugs, growth inhibitory agents, toxins (e.g., protein toxins, enzymatically active toxins of bacterial, fungal, plant, or animal origin, or fragments thereof), or radioactive isotopes.
- agents such as cytotoxic agents, chemotherapeutic agents, drugs, growth inhibitory agents, toxins (e.g., protein toxins, enzymatically active toxins of bacterial, fungal, plant, or animal origin, or fragments thereof), or radioactive isotopes.
- the invention provides immunoconjugates comprising an antibody provided herein conjugated to a polymer.
- polymer used herein includes chemical polymers and protein polymers.
- the immunoconjugate comprises the antibody provided herein conjugated to an extended recombinant polypeptide (XTEN).
- the immunoconjugate comprises an XTEN (a) comprising
- GGSPAGSCTSP GASASCAPSTG, TAEAAGCGTAEAA, and GPEPTCPAPSG.
- (b) being 36 to 3000 L-amino acid residues in length, and/or (c) wherein the sum of glycine (G), alanine (A), serine (S), threonine (T), glutamate (E) and proline (P) residues constitutes more than 90% of the total amino acid residues of the XTEN.
- Antibodies may be produced using recombinant methods and compositions, e.g., as described in U.S. Pat. No. 4,816,567. For these methods one or more isolated nucleic acid(s) encoding an antibody are provided.
- nucleic acids encoding an antibody of the invention are provided. Such nucleic acids may be readily isolated and sequenced using conventional procedures (e.g., by using oligonucleotide probes that are capable of binding specifically to genes encoding the heavy and light chains of the antibody) or produced by recombinant methods or obtained by chemical synthesis.
- Suitable host cells for cloning or expression of antibody-encoding vectors include prokaryotic or eukaryotic cells described herein.
- antibodies may be produced in bacteria, in particular when glycosylation and Fc effector function are not needed.
- For expression of antibody fragments and polypeptides in bacteria see, e.g., U.S. Pat. Nos. 5,648,237, 5,789,199, and 5,840,523. (See also Charlton, K. A., In: Methods in Molecular Biology, Vol.
- the antibody may be isolated from the bacterial cell paste in a soluble fraction and can be further purified.
- the host cell is an E. coli cell.
- Vertebrate cells may also be used as hosts.
- mammalian cell lines that are adapted to grow in suspension may be useful.
- useful mammalian host cell lines are monkey kidney CV1 line transformed by SV40 (COS-7); human embryonic kidney line (293 or 293T cells as described, e.g., in Graham, F. L. et al., J. Gen Virol. 36 (1977) 59-74); baby hamster kidney cells (BHK); mouse sertoli cells (TM4 cells as described, e.g., in Mather, J. P., Biol. Reprod. 23 (1980) 243-252); monkey kidney cells (CV1); African green monkey kidney
- VERO-76 human cervical carcinoma cells
- HELA human cervical carcinoma cells
- MDCK canine kidney cells
- BBL 3 A canine kidney cells
- W138 human liver cells
- Hep G2 human liver cells
- MMT 060562 mouse mammary tumor
- TRI cells as described, e.g., in Mather, J. P. et al., Annals N.Y. Acad. Sci. 383 (1982) 44-68); MRC 5 cells; and FS4 cells.
- Other useful mammalian host cell lines include Chinese hamster ovary (CHO) cells, including DHFR- CHO cells (Urlaub, G. et al., Proc. Natl. Acad. Sci.
- myeloma cell lines such as Y0, NS0 and Sp2/0.
- myeloma cell lines such as Y0, NS0 and Sp2/0.
- the host cell is eukaryotic, e.g., a Chinese Hamster Ovary (CHO) cell or lymphoid cell (e.g., Y0, NS0, Sp20 cell).
- the host cell is a CHO cell. Production of antibodies of the invention in CHO cells may improve syringeability of the antibody.
- an anti-C3 antibody as herein described is capable of specifically binding to an epitope of human complement factor C3 and/or proteolytic derivatives C3a and/or C3b.
- an anti-C3 antibody is capable of specifically binding to an epitope in any region of the polypeptide sequence identified by GenBank Accession No. NP 000055.2, the entire contents of which are incorporated herein by reference.
- the antibodies described herein comprise one or more CDRs.
- the CDRs may identify the specificity of the antibody and accordingly it is preferred that the antigen binding site comprises at least 2, and more preferably comprises 3 CDRs.
- the antibodies provided herein are preferably derived from natural antibodies, such as camelid antibodies.
- the antibody is selected from the following sequences or sequences at least 75% identical thereto (CDR sequences are underlined):
- the antibody comprises a CDR having at least one, at least two, or at least three amino acid differences relative to a CDR as set forth in Table 1.
- the antibody comprises a CDR1 having the following sequence: G - F/L - T/S - F/L - S/G/D - P/V/Y/N - Y - A/D - M/I - G/S/A (wherein the first amino acid is G, the second amino acid is F or L, etc.) and/or comprises a CDR2 having the following sequence: G/S/C - 1 -
- the antibody comprises a CDR1, CDR2 and/or CDR3 selected from the group consisting of the CDR sequences set forth at Table 1.
- the antibody comprises a CDR1 selected from SEQ ID Nos: 15, 18, 21, 24, 27, 30, 33, 36, 39, 42, 45, 48, 51, and 54, or comprises a CDR1 comprising a sequence at least 75% identical to any one of SEQ ID Nos:15, 18, 21, 24, 27, 30, 33, 36, 39, 42, 45, 48, 51, and 54; and/or comprises a CDR2 selected from SEQ ID Nos:16, 19, 22, 25, 28, 31, 34, 37, 40, 43, 46, 49, 52, and 55 or comprises a CDR2 comprising a sequence at least 75% identical to any one of SEQ ID Nos: 16, 19, 22, 25, 28, 31, 34, 37, 40, 43, 46, 49, 52, and 55; and/or comprises a CDR3 selected from SEQ ID Nos: 17, 20, 23, 26, 29, 32, 35, 38, 41, 44, 47, 50, 53, and 56 or comprises a CDR1 comprising a sequence at least 75% identical to any one of SEQ ID Nos:
- the antibody comprises a CDR1, CDR2, and CDR3 selected from: SEQ ID Nos: 15-17, 18-20, 21-23, 24-26, 27-29, 30-32, 33-35, 36-38, 39-41, 42-44, 45-47, 48-50, 51-53 and 54-56 or comprises a CDR1, CDR2 and CDR3 at least 75% identical to SEQ ID Nos: 15-17, 18- 20, 21-23, 24-26, 27-29, 30-32, 33-35, 36-38, 39-41, 42-44, 45-47, 48-50, 51-53 and 54-56.
- the antibody comprises a CDR1, CDR2 and CDR3 of SEQ ID Nos: 54-56, respectively, or comprises CDRs at least 75% (e.g., at least 77%, at least 78%, at least 79%, at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97% or at least 98%) identical thereto.
- CDR1, CDR2 and CDR3 of SEQ ID Nos: 54-56 respectively, or comprises CDRs at least 75% (e.g., at least 77%, at least 78%, at least 79%, at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%
- amino acid sequence of human C3 is provided below:
- an anti-C3 antibody as herein described (e.g. C3N1O.3) binds to an epitope within the 03 alpha-chain at the C345C domain. In some aspects, an anti-C3 antibody as herein described binds to an epitope within amino acids 1540-1600 of human C3 or an amino acid sequence at least 80% identical thereto.
- an anti-03 antibody as herein described binds to an epitope comprising amino acids 1548-1561 of human C3 or an amino acid sequence at least 80% identical thereto and/or binds to an epitope comprising amino acids 1568- 40 1573 of human C3 or an amino acid sequence at least 80% identical thereto and/or binds to an epitope comprising amino acids 1583-1591 of human C3 or an amino acid sequence at least 80% identical thereto.
- an anti-C3 antibody as herein described (e.g. C3S3.2) binds to at least one discontinuous epitope within the C3 beta-chain at the MG1 and MGS domains. In some aspects, an anti-C3 antibody as herein described binds to at least one discontinuous epitope within amino acids 110-130 of human C3 and within amino acids 490-501 of human C3. In some aspects, an anti-C3 antibody as herein described binds to an epitope comprising amino acids 114-124 of human C3 or an amino acid sequence at least 80% identical thereto and/or binds to an epitope comprising amino acids 497-505 of human C3 or an amino acid sequence at least 80% identical thereto.
- the anti-C3 antibody comprises a CDR1, CDR2 and/or CDR3 of the anti-C3 antibody of any one of SEQ ID Nos: 2, 14, 98 and 135 or comprises a CDR1, CDR2 and/or CDR3 at least 70% identical thereto (or comprising four or fewer amino acid substitutions relative to CDR1, CDR2, CDR3 of any one of SEQ ID Nos: 2, 14, 98 and 135).
- an anti-C5 antibody e.g., nanobody
- an anti-C5 antibody is capable of specifically binding to an epitope of human complement factor C5 and/or proteolytic derivatives C5a and/or C5b.
- an anti-C5 antibody is capable of specifically binding to an epitope in any region of the polypeptide sequence identified by GenBank Accession No. NP_001304092.1, the entire contents of which are incorporated herein by reference.
- the antibodies described herein comprise one or more CDRs.
- the CDRs may identify the specificity of the antibody and accordingly it is preferred that the antigen binding site comprises at least 2, and more preferably comprises 3 CDRs.
- the antibodies provided herein are preferably derived from natural antibodies, such as camelid antibodies.
- the antibody is selected from the following sequences or sequences at least 75% identical thereto (CDR sequences are underlined):
- the antibody comprises a CDR having at least one, at least two, or at least three amino acid differences relative to a CDR as set forth in Table 2.
- the antibody comprises a CDR1 having the following sequence: G - Y/N/S/T - 1 -
- the antibody comprises a CDR1 having the following sequence: G - Y/N/S/T - 1 - F - Y/H/F/A/P/D - D/Q/A/Y/S/E - D/S/T/E/Y/W - D - M - G and/or comprises a CDR2 having the following sequence - A/T - 1 - D - V/Y/G/D - G - A - S/N - T - Y.
- the antibody comprises a CDR1, CDR2 and/or CDR3 selected from the group consisting of the CDR sequences set forth at Table 2.
- the antibody comprises a CDR1 selected from SEQ ID Nos: 66, 69, 72, 75, 78, 81, 84, 87 and 90 or comprises a CDR1 comprising a sequence at least 75% identical to any one of SEQ ID Nos: 66, 69, 72, 75, 78, 81, 84, 87 and 90; and/or comprises a CDR2 selected from SEQ ID Nos:67, 70, 73, 76, 79, 82, 85, 88 and 91 or comprises a CDR2 comprising a sequence at least 75% identical to any one of SEQ ID Nos: 67, 70, 73, 76, 79, 82, 85, 88 and 91; and/or comprises a CDR3 selected from SEQ ID Nos: 68, 71, 74, 77, 80, 83, 86, 89 and 92 or comprises a CDRI comprising a sequence at least 75% identical to any one of SEQ ID Nos: 66,
- the antibody comprises a CDRI, CDR2, and CDR3 selected from: SEQ ID Nos:66-68, 69-71, 72-74, 75-77, 78-80, 81-83, 84-86, 87-89 and 90-92 or comprises a CDRI, CDR2 and CDR3 at least 75% identical to SEQ ID Nos: 66-68, 69-71, 72-74, 75-77, 78-80, 81-83, 84-86, 87-89 and 90-92.
- amino acid sequence of human C5 is provided below:
- an anti-C5 antibody as herein described (e.g., C5S2.2) competitively inhibits eculizumab from binding to human C5.
- the anti-C5 antibody comprises a CDR1, CDR2 and/or CDR3 of the anti-C5 antibody of any one of SEQ ID Nos: 58 and 141 or comprises a CDR1, CDR2 and/or CDR3 at least 70% identical thereto (or comprising four or fewer amino acid substitutions relative to CDR1, CDR2, CDR3 of any one of SEQ ID Nos: 58 and 141).
- the antibodies provided herein also include functional variants thereof.
- the term “functional variant” is meant to include those variants, which retain some or essentially all the ability of an antibody to selectively binding its antigen or ligand, such as any of the ligands
- Functional variants include any variant, which is at least 75% identical to antibodies provided herein, such as at least 80, 81 , 82, 83, 84, 85, 86, 87, 88, 89, such as 90, 91 , 92, 93, 94, 95, 96, such as 97, such as 98, such as 99, such as at least 99.5% identical to an antibody provided herein, such as any of those identified by SEQ ID NOs: 1-14 and 57-65.
- the antibody comprises the amino acid sequence set forth as SEQ ID NO: 14 or an amino acid sequence at least 75% identical thereto.
- Functional variants include any variant antibody comprising one or more CDR(s), which is at least 75% identical to a CDR of an antibody provided herein, such as at least 80, 81 , 82, 83, 84, 85, 86, 87, 88, 89, such as 90, 91 , 92, 93, 94, 95, 96, such as 97, such as 98, such as 99, such as at least 99.5% identical to said CDR.
- the CDRs of the respective antibodies identified by SEQ ID NOs: 1-14 and 57-65 are indicated above as SEQ ID Nos: 15-56 and 66- 92, respectively.
- antibodies comprise one or more regions having at least 75%, such as at least 80, 81 , 82, 83, 84, 85, 86, 87, 88, 89, such as 90, 91 , 92, 93, 94, 95, 96, such as 97, such as 98, such as 99, such as at least 99.5% identity to one or more regions identified by any one of SEQ ID Nos: 15-56 and 66-92.
- Amino acid substitutions include conservative amino acid substitutions, which refer to substitution of one amino acid with another amino acid residue having a side chain with similar properties.
- a group of amino acids having aliphatic side chains is glycine, alanine, valine, leucine, and isoleucine
- a group of amino acids having aliphatic-hydroxyl side chains is serine and threonine
- a group of amino acids having amide-containing side chains is asparagine and glutamine
- a group of amino acids having aromatic side chains is phenylalanine, tyrosine, and tryptophan
- a group of amino acids having basic side chains is lysine, arginine, and histidine
- a group of amino acids having sulfur-containing side chains is cysteine and methionine.
- Preferred conservative amino acids substitution groups are: valine-leucine-isoleucine, phenylalanine-tyrosine, lysine-arginine, alanine-valine, and asparagine-glutamine.
- one amino acid may be substituted for another within the groups of amino acids indicated herein below: i) Amino acids having polar side chains (Asp, Glu, Lys, Arg, His, Asn, Gin, Ser, Thr, Tyr, and Cys,) ii) Amino acids having non-polar side chains (Gly, Ala, Vai, Leu, lie, Phe, Trp, Pro, and Met)
- Amino acids having aliphatic side chains (Gly, Ala Vai, Leu, lie) iv) Amino acids having cyclic side chains (Phe, Tyr, Tip, His, Pro) v) Amino acids having aromatic side chains (Phe, Tyr, Trp) vi) Amino acids having acidic side chains (Asp, Glu) vii) Amino acids having basic side chains (Lys, Arg, His) viii) Amino acids having amide side chains (Asn, Gin) ix) Amino acids having hydroxy side chains (Ser, Thr) x) Amino acids having sulphur-containing side chains (Cys, Met), xi) Neutral, weakly hydrophobic amino acids (Pro, Ala, Gly, Ser, Thr) xii) Hydrophilic, acidic amino acids (Gin, Asn, Glu, Asp), and xiii) Hydrophobic amino acids (Leu, lie, Vai)
- Functional variants of antibodies may in one preferred embodiment be a fragment of an antibody, preferably an antigen-binding fragment or a variable region.
- antibody fragments useful with the present invention include fragments of VHH and VNAR.
- the antibodies may thus be obtained by immunization of any suitable organism, in particular camelids, sharks or the like.
- the antibodies could also be generated from a synthetic library with randomized or designed CDRs.
- an antibody capable of specifically binding to an epitope of human complement factor C3, C3a and/or C3b provided.
- an antibody capable of specifically binding to an epitope of human complement factor C5, C5a and/or C5b provided.
- the antibody is coupled to a histidine tag, such as a his6 tag at the N- or C-terminus of the polypeptide.
- the antibody is coupled to a non-native Strep or his6 tag (e.g., WSHPQFEKHHHHHH (SEQ ID NO:93) at the C -terminal.
- the antibody is coupled to an Fc-fragmcnt at the N-terminal.
- N-terminal additions are particularly preferred, and in a preferred embodiment, the antibody provided herein comprises an additional N- terminal region.
- the additional N-terminal region can be selected from any relevant additional moieties, depending on the contemplated application of the antibody and the desired functionalities to the final antibody product.
- Albumin may be added for increasing circulation time and protect the product from degradation.
- Other antigen binding fragments, antibodies or fragments thereof may be added for introducing a second affinity /binding specificity to the antibody product.
- an anti-03 antibody as herein described forms a first domain in a fusion protein.
- the fusion protein is a heterodimeric fusion protein comprising a first domain comprising a polypeptide comprising an antibody as herein described and a second domain comprising an immunoglobulin Fc domain.
- the first domain and second domain are joined by a linker.
- the fusion protein is a heterodimeric fusion protein comprising the amino acid sequence of any one of SEQ ID Nos: 178- 180 or an amino acid sequence at least 70% identical thereto.
- the fusion protein is a homodimeric fusion protein comprising a first domain comprising a polypeptide comprising an antibody as herein described and a second domain identical to the first domain.
- the first domain and second domain are joined by a linker, e.g., a poly-Glycine-Serine (G4S) linker, preferably a (G4S)2 linker.
- the fusion protein is a homodimeric fusion protein comprising the amino acid sequence of SEQ ID NO: 181 or 182 or an amino acid sequence at least 70% identical there.
- a bispecific/multispecific antibody e.g., a single peptide chain comprising two antigen-binding regions, which may be separated by a linker sequence, wherein one of the two antigen-binding regions comprises a VHH domain as herein described.
- a VHH domain as herein described is coupled to another VHH domain yielding a bispecific antibody, which consists of or comprises (i) a VHH domain as herein described and (ii) a VHH domain capable of specifically binding to an epitope of a second target.
- the antibodies disclosed herein may in preferred embodiments comprise modifications, which improve the function of the antibody.
- modifications which improve the function of the antibody.
- an antibody provided herein may be a humanized antibody.
- the humanized antibody is a humanized form of the llama nanobodies described above and is selected from among the following amino acid sequences or amino acid sequences at least 75% identical thereto (the bolded amino acids are the humanized form amino acids that have been changed from the corresponding llama nanobody amino acid):
- VSS (SEQ ID NO:97)
- VSS (SEQ ID NO:98)
- VSS (SEQ ID NO:99)
- TLVTVSS (SEQ ID NO: 100)
- TLVTVSS (SEQ ID NO: 101)
- TLVTVSS (SEQ ID NO: 102)
- TLVTVSS (SEQ ID NO: 103)
- VSS (SEQ ID NO: 106)
- VSS (SEQ ID NO: 107)
- VOL VESGGGLVQAGGSLRLSCAASGYIFVYSTMGWYROAPGKEREF VASIN YGSITYY
- VSS (SEQ ID NO: 118)
- VSS (SEQ ID NO: 119)
- VSS (SEQ ID NO: 120)
- TLVTVSS (SEQ ID NO: 124)
- TLVTVSS (SEQ ID NO: 126)
- VSS (SEQ ID NO:129)
- VTVSS (SEQ ID NO: 130)
- VSS (SEQ ID NO: 132)
- VSS (SEQ ID NO: 133)
- VSS (SEQ ID NO: 134)
- VTVSS (SEQ ID NO: 138)
- VTVSS (SEQ ID NO: 139)
- VSS (SEQ ID NO: 141)
- VTVSS (SEQ ID NO:143)
- VSS (SEQ ID NO: 147)
- VSS (SEQ ID NO: 148)
- VSS (SEQ ID NO: 149)
- TLVTVSS (SEQ ID NO: 153)
- VTVSS (SEQ ID NO: 157)
- VTVSS (SEQ ID NO: 158)
- the antibody comprises the sequence of any one of SEQ ID Nos:94-163 or comprises an amino acid sequence at least 75%, such as at least 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or at least 99.5% identical thereto and preferably comprises the bolded amino acids.
- nucleic acid comprising a nucleotide sequence encoding an antibody as herein described is provided.
- the nucleic acid encoding the antibody as herein described forms part of an expression vector in which the nucleotide sequence encoding the antibody is operably linked to a suitable promoter.
- the RNA and/or cDNA coding sequences are designed for optimal expression in human cells.
- Codon-optimized coding regions can be designed by various different methods. This optimization may be performed using methods which are available online, published methods, or a company which provides codon optimizing services.
- One codon optimizing method is described, e.g., in WO 2015/012924 A2, which is incorporated by reference herein. Briefly, the nucleic acid sequence encoding the product is modified with synonymous codon sequences. Suitably, the entire length of the open reading frame (ORF) for the product is modified. However, in some embodiments, only a fragment of the ORF may be altered. By using one of these methods, one can apply the frequencies to any given polypeptide sequence, and produce a nucleic acid fragment of a codon- optimized coding region which encodes the polypeptide.
- a pharmaceutical composition comprising a pharmaceutically acceptable carrier and a nucleic acid comprising a nucleotide sequence encoding an antibody as herein described, said nucleotide sequence operably linked to a promoter.
- a pharmaceutical composition comprising an antibody as herein described and a pharmaceutically acceptable carrier.
- novel rAAV virions described herein comprise (i) a capsid as herein described and (ii) a heterologous nucleic acid encoding an anti-C3 and/or anti-C5 antibody as herein described, preferably wherein the nucleotide sequence encoding the anti-C3 and/or anti-C5 antibody is operably linked to an expression control sequence.
- the heterologous nucleic acid includes an AAV genome with the rep and cap genes deleted and/or replaced by the antibody sequence and its associated expression control sequences.
- the antibody sequence is typically inserted adjacent to one or two (i.e., is flanked by) AAV TRs or TR elements adequate for viral replication (Xiao et al., 1997, J.
- Virol. 71(2): 941-948 in place of the nucleic acid encoding viral rep and cap proteins.
- Other regulatory sequences suitable for use in facilitating tissue-specific expression of the antibody y gene sequence in the target cell may also be included.
- the anti-C3 antibody encoded by the heterologous nucleic acid of the rAAV is able to prevent binding of C3 to the C3 convertases of the alternative pathway and the classical pathway which in turn inhibits C3 cleavage.
- the rAAV virion comprises a heterologous nucleic acid comprising (a) an AAV2 terminal repeat (b) a transcription control sequence (c) a nucleotide sequence encoding an anti-C3 antibody as herein described (d) a polyadenylation sequence and (e) an AAV2 terminal repeat.
- the rAAV virion comprises a heterologous nucleic acid comprising (a) an AAV2 terminal repeat (b) a transcription control sequence (c) a nucleotide sequence encoding an anti-C5 antibody as herein described (d) a polyadenylation sequence and (e) an AAV2 terminal repeat.
- the inverted terminal repeats (ITR(s)) selected for use in the rAAV virion are preferably AAV sequences, with serotypes 1, 2, 3, 4, 5 and 6 being preferred.
- the ITRs may be synthetic sequences that function as AAV inverted terminal repeats, such as the "double-D sequence" as described in U.S. Pat. No. 5,478,745 to Samulski et al., the entire disclosure of
- the TRs are from the same parvovirus, e.g., both ITR sequences are from AAV2.
- the heterologous nucleic encapsulated by the rAAV virion comprises a 5' ITR with the following sequence:
- the heterologous nucleic acid encapsulated by the rAAV virion comprises a 3' ITR with the following sequence:
- the antibody gene encoded by the rAAV is preferably operatively linked to at least one transcription control sequence, preferably a transcription control sequence that is heterologous to the nucleic acid.
- the transcription control sequence comprises a cell- or tissue-specific promoter that results in cell-specific expression of the nucleic acid e.g. in photoreceptor cells, such as human rod photoreceptor-specific human G-protein coupled receptor rhodopsin kinase 1 (hGRK) promoter or a human interphotoreceptor retinoid-binding protein (IRBP) promoter.
- the transcription control sequence comprises a constitutive promoter that results in similar expression level of the nucleic acid in many cell types.
- Suitable constitutive promoters include CAG promoter comprising (C) cytomegalovirus (CMV) immediate-early enhancer element, (A) first exon and the first intron of chicken beta-actin gene and (G) splice acceptor of the rabbit beta-globin gene (see Miyazaki et al. (1989) Gene 79(2): 269-277), cytomegalovirus promoter (CMV) (Stinski et al, (1985) Journal of Virology 55(2): 431-441), human elongation factor la promoter (EFla) (Kim et al.
- CMV cytomegalovirus
- A first exon and the first intron of chicken beta-actin gene
- G splice acceptor of the rabbit beta-globin gene
- the antibody gene encoded by the rAAV is operably linked to a CAG promoter.
- the CAG promoter comprises the
- sequence of SEQ ID NO:166 or comprises a sequence at least 95%, at least 96%, at least 97%, at least 98% or at least 99% identical thereto:
- the heterologous nucleic acid encapsulated by the rAAV virion comprises a SV40 polyadenylation sequence the following sequence:
- the variant AAV capsid of the rAAV - encapsulating the heterologous nucleic acid encoding an antibody or functional variant thereof - comprises a variant AAV capsid protein comprising an insertion of from about 7 to about 20 amino acids (a “heterologous peptide” or “peptide insertion”) in the GH-loop of a parental AAV capsid protein, wherein the peptide comprises the amino acid sequence ISDQTKH (SEQ ID NO: 168).
- the variant capsid protein when present in an AAV virion, confers increased infectivity of a retinal cell compared to the infectivity of a retinal cell by an AAV virion comprising the corresponding parental capsid protein.
- the GH loop or loop IV, of the AAV capsid protein it is meant the solvent- accessible portion referred to in the art as the GH loop, or loop IV, of AAV capsid protein.
- the insertion site can be within about amino acids 570-611 of AAV2 VP1.
- the peptide insertion has from 1 to 3 spacer amino acids (Y i- Y3) at the amino and/or carboxyl terminus of the amino acid sequence ISDQTKH (SEQ ID NO: 168).
- spacer amino acids include, without limitation, leucine (L), alanine (A), glycine (G), serine (S), threonine (T), and proline (P).
- a peptide insertion comprises 2 spacer amino acids at the N-terminus and 2 spacer amino acids at the C- terminus.
- a peptide insertion comprises 2 spacer amino acids at the N- terminus and 1 spacer amino acids at the C -terminus.
- the peptide insertion comprises or consists of the amino acid sequence LAISDQTKHA (SEQ ID NO: 169).
- the variant AAV capsid protein comprises a peptide insertion comprising the amino acid sequence ISDQTKH (SEQ ID NO: 168) and further comprises one or more amino acid substitutions relative to a corresponding parental AAV capsid protein.
- amino acid substitutions may be found at e.g., col. 26, lines 40-65 of U.S. Patent No. 11,576,983, the entire contents of which are incorporated herein by reference.
- the variant AAV capsid protein comprises a peptide insertion comprising the amino acid sequence ISDQTKH (SEQ ID NO: 168) and further
- 64 comprises a P34A amino acid substitution relative to VP1 capsid of AAV2 or the corresponding substitution in another AAV serotype.
- the variant capsid protein may comprise one or more features disclosed in U.S. Patent No. 11,576,983, in particular, one or more features disclosed at column 26, line 66 to column 29, line 50 of U.S. Patent No. 11,576,983.
- the variant capsid protein comprises the following amino acid sequence or comprises an amino acid sequence at least 80%, at least 90%, least 95%, at least 98%, or at least 99% identical to the following amino acid sequence:
- the variant AAV capsid protein of SEQ ID NO: 170 contains the following modifications relative to native AAV2 capsid: (i) a proline (P) to alanine (A) mutation at amino acid position 34, which is located inside the assembled capsid (VP1 protein only), and (ii) an insertion of 10 amino acids (leucine-alanine-isoleucine-serine-aspartic acid-glutamine-threonine- lysine-histidine-alanine/LAISDQTKHA (SEQ ID NO: 169)) at amino acid position 588, which is present in VP1, VP2, and VP3.
- the capsid comprises a variant capsid protein comprising a sequence at least 90%, at least 95%, at least 98%, at least 99% identical to SEQ ID NO: 170 and comprising a P34A substitution and an LAISDQTKHA (SEQ ID NO: 169) peptide insertion at amino acid position 588.
- packaging cells which are encompassed by “host cells,” which may be cultured to produce packaged viral vectors of the invention.
- the packaging cells of the invention generally include cells with heterologous (1) viral vector function(s), (2)
- the vectors can be made by several methods known to skilled artisans (see, e.g., WO 2013/063379). A preferred method is described in Grieger, et al. 2015, Molecular Therapy 24(2):287-297, the contents of which are incorporated by reference herein for all purposes. Briefly, efficient transfection of HEK293 cells is used as a starting point, wherein an adherent HEK293 cell line from a qualified clinical master cell bank is used to grow in animal component-free suspension conditions in shaker flasks and WAVE bioreactors that allow for rapid and scalable rAAV production.
- the suspension HEK293 cell line generates greater than 10 5 vector genome containing particles (vg)/cell or greater than 10 14 vg/L of cell culture when harvested 48 hours post-transfection.
- triple transfection refers to the fact that the packaging cell is transfected with three plasmids: one plasmid encodes the AAV rep and cap genes, another plasmid encodes various helper functions (e.g., adenovirus or HSV proteins such as Ela, Elb, E2a, E4, and VA RNA, and another plasmid encodes the transgene and its various control elements (e.g., modified GLA gene and GAG promoter).
- helper functions e.g., adenovirus or HSV proteins such as Ela, Elb, E2a, E4, and VA RNA
- transgene and its various control elements e.g., modified GLA gene and GAG promoter
- This scalable manufacturing technology has been utilized to manufacture GMP Phase I clinical AAV vectors for retinal neovascularization (AAV2), Hemophilia B (scAAV8), Giant Axonal Neuropathy (scAAV9) and Retinitis Pigmentosa (AAV2), which have been administered into patients.
- AAV2 retinal neovascularization
- scAAV8 Hemophilia B
- scAAV9 Giant Axonal Neuropathy
- AAV2 Retinitis Pigmentosa
- a minimum of a 5-fold increase in overall vector production by implementing a perfusion method that entails harvesting rAAV from the culture media at numerous time-points post-transfection.
- the packaging cells include viral vector functions, along with packaging and vector functions.
- the viral vector functions typically include a portion of a parvovirus genome, such as an AAV genome, with rep and cap deleted and replaced by the modified GLA sequence and its associated expression control sequences.
- the viral vector functions include sufficient expression control sequences to result in replication of the viral vector for packaging.
- the viral vector includes a portion of a parvovirus genome, such as an AAV genome with rep and cap deleted and replaced by the transgene and its associated expression control sequences.
- the transgene is typically flanked by two AAV TRs, in place of the deleted viral rep and cap ORFs.
- transgene is typically a nucleic acid sequence that can be expressed to produce a therapeutic polypeptide or a marker polypeptide.
- the terminal repeats (TR(s)) (resolvable and non-resolvable) selected for use in the viral vectors are preferably AAV sequences, with serotypes 1, 2, 3, 4, 5 and 6 being preferred.
- Resolvable AAV TRs need not have a wild-type TR sequence (e.g., a wild-type sequence may be altered by insertion, deletion, truncation or missense mutations), as long as the TR mediates the desired functions, e.g., virus packaging, integration, and/or provirus rescue, and the like.
- the TRs may be synthetic sequences that function as AAV inverted terminal repeats, such as the "double-D sequence" as described in U.S. Pat. No. 5,478,745 to Samulski et at, the entire disclosure of which is incorporated in its entirety herein by reference.
- the TRs are from the same parvovirus, e.g., both TR sequences are from AAV2.
- the packaging functions include variant capsid components as described above.
- the packaged viral vector includes an anti-C3 antibody transgene and expression control sequences flanked by TR elements, referred to herein as the "transgene” or “transgene expression cassette,” sufficient to result in packaging of the vector DNA and subsequent expression of the gene sequence in the transduced cell.
- the viral vector functions may, for example, be supplied to the cell as a component of a plasmid or an amplicon.
- the viral vector functions may exist extrachromosomally within the cell line and/or may be integrated into the cell's chromosomal DNA.
- any method of introducing the nucleotide sequence carrying the viral vector functions into a cellular host for replication and packaging may be employed, including but not limited to, electroporation, calcium phosphate precipitation, microinj ection, cationic or anionic liposomes, and liposomes in combination with a nuclear localization signal.
- the viral vector functions are provided by transfection using a virus vector; standard methods for producing viral infection may be used.
- the packaging functions include genes for viral vector replication and packaging.
- the packaging functions may include, as needed, functions necessary for viral gene expression, viral vector replication, rescue of the viral vector from the integrated state, viral gene expression, and packaging of the viral vector into a viral particle.
- the packaging functions may be supplied together or separately to the packaging cell using a genetic construct such as a plasmid or an amplicon, a Baculovirus, or HS V helper construct.
- the packaging functions may exist extrachromosomally within the packaging cell, but are preferably integrated into the cell's chromosomal DNA. Examples include genes encoding AAV Rep and Cap proteins.
- the helper functions include helper virus elements needed for establishing active infection of the packaging cell, which is required to initiate packaging of the viral vector. Examples include functions derived from adenovirus, baculovirus and/or herpes virus sufficient to result in packaging of the viral vector.
- adenovirus helper functions will typically include adenovirus components Ela, Elb, E2a, E4, and VA RNA.
- the packaging functions may be supplied by infection of the packaging cell with the required virus.
- the packaging functions may be supplied together or separately to the packaging cell using a genetic construct such as a plasmid or an amplicon. See, e.g., pXR helper plasmids as described in Rabinowitz et al., 2002, J.
- the packaging functions may exist extrachromosomally within the packaging cell, but are preferably integrated into the cell's chromosomal DNA (e.g., El or E3 in HEK 293 cells).
- helper virus functions may be employed.
- the packaging cells are insect cells
- baculovirus may serve as a helper virus.
- Herpes virus may also be used as a helper virus in AAV packaging methods.
- Hybrid herpes viruses encoding the AAV Rep protein(s) may advantageously facilitate for more scalable AAV vector production schemes.
- any method of introducing the nucleotide sequence carrying the helper functions into a cellular host for replication and packaging may be employed, including but not limited to, electroporation, calcium phosphate precipitation, microinjection, cationic or anionic liposomes, and liposomes in combination with a nuclear localization signal.
- the helper functions are provided by transfection using a virus vector or infection using a helper virus; standard methods for producing viral infection may be used.
- any suitable permissive or packaging cell known in the art may be employed in the production of the packaged viral vector.
- Mammalian cells or insect cells are preferred.
- Examples of cells useful for the production of packaging cells in the practice of the invention include, for example, human cell lines, such as VERO, WI38, MRC5, A549, HEK 293 cells (which express functional adenoviral El under the control of a constitutive promoter), B-50 or any other HeLa cells, HepG2, Saos-2, HuH7, and HT1080 cell lines.
- the packaging cell is capable of growing in suspension culture, more preferably, the cell is capable of growing in serum-free culture.
- the packaging cell is a HEK293 that grows in suspension in serum free medium.
- the packaging cell is the HEK293 cell described in U.S. Pat. No. 9,441,206 and deposited as ATCC No. PT A 13274. Numerous rAAV packaging cell lines are known in the art, including, but not limited to, those disclosed in WO 2002/46359.
- the packaging cell is cultured in the form of a cell stack (e.g. 10-layer cell stack seeded with HEK293 cells).
- Cell lines for use as packaging cells include insect cell lines. Any insect cell which allows for replication of AAV and which can be maintained in culture can be used in accordance with the present invention. Examples include Spodoptera frugiperda, such as the Sf9 or S£21 cell lines, Drosophila spp. cell lines, or mosquito cell lines, e.g., Aedes albopictus derived cell lines. A preferred cell line is the Spodoptera frugiperda Sf9 cell line.
- the following references are incorporated herein for their teachings concerning use of insect cells for expression of heterologous polypeptides, methods of introducing nucleic acids into such cells, and methods of maintaining such cells in culture: Methods in Molecular Biology, ed.
- Virus capsids according to the invention can be produced using any method known in the art, e.g., by expression from a baculovirus (Brown et al., (1994) Virology 198:477-488).
- the virus vectors of the invention can be produced in insect cells using baculovirus vectors to deliver the rep/cap genes and rAAV template as described, for example, by Urabe et al., 2002, Human Gene Therapy 13:1935-1943.
- the present invention provides for a method of rAAV production in insect cells wherein a baculovirus packaging system or vectors may be constructed to carry the AAV Rep and Cap coding region by engineering these genes into the polyhedrin coding region of a baculovirus vector and producing viral recombinants by transfection into a host cell.
- a baculovirus packaging system or vectors may be constructed to carry the AAV Rep and Cap coding region by engineering these genes into the polyhedrin coding region of a baculovirus vector and producing viral recombinants by transfection into a host cell.
- the AAV DNA vector product is a self- complementary AAV like molecule without using mutation to the AAV ITR. This appears to be a by-product of inefficient AAV rep nicking in insect cells which results in a self-complementary DNA molecule by virtue of lack of functional Rep enzyme activity.
- the host cell is a baculovirus-infected cell or has introduced therein additional nucleic acid encoding baculovirus helper functions or includes these baculovirus helper functions therein.
- These baculovirus viruses can express the AAV components and subsequently facilitate the production of the capsids.
- the packaging cells generally include one or more viral vector functions along with helper functions and packaging functions sufficient to result in replication and packaging of the viral vector. These various functions may be supplied together or separately to the packaging cell using a genetic construct such as a plasmid or an amplicon, and they may exist extrachromosomally within the cell line or integrated into the cell's chromosomes.
- the cells may be supplied with any one or more of the stated functions already incorporated, e.g., a cell line with one or more vector functions incorporated extrachromosomally or integrated into the cell's chromosomal DNA, a cell line with one or more packaging functions incorporated extrachromosomally or integrated into the cell's chromosomal DNA, or a cell line with helper functions incorporated extrachromosomally or integrated into the cell's chromosomal DNA.
- a cell line with one or more vector functions incorporated extrachromosomally or integrated into the cell's chromosomal DNA e.g., a cell line with one or more vector functions incorporated extrachromosomally or integrated into the cell's chromosomal DNA, a cell line with one or more packaging functions incorporated extrachromosomally or integrated into the cell's chromosomal DNA, or a cell line with helper functions incorporated extrachromosomally or integrated into the cell's chromosomal DNA.
- methods for delivering heterologous nucleotide sequences encoding anti-C3 and/or anti-C5 antibodies to the retina are provided utilizing an rAAV as herein described.
- the rAAV may be employed to deliver a nucleotide sequence encoding the antibody to a retinal cell in vitro, e.g., to produce an antibody polypeptide or nucleic acid in vitro for ex vivo gene therapy.
- the rAAV are additionally useful in a method of delivering a nucleotide sequence to a subject in need thereof, e.g., to express the antibody in a subject in need thereof, such as human with dry AMD or a human with geographic atrophy. In this manner, the antibody may thus be produced in vivo in the subject to restore complement regulation.
- a method of delivering a nucleic acid encoding an anti-C3 and/or anti-C5 antibody or antigen-binding fragment thereof to a retinal cell comprising contacting the retinal cell with an rAAV virion as herein described.
- a method of delivering a nucleic acid encoding an anti-C3 and/or anti-C5 antibody or antigen-binding variant thereof to a retinal cell in a mammalian subject comprising administering an effective amount of the rAAV virion as herein described or a pharmaceutical formulation comprising same to a mammalian subject.
- the rAAV may be administered to retina of a subject by any suitable route.
- the rAAV is administered to the subject intraocularly, preferably by subretinal, suprachoroidal, and/or intravitreal injection.
- rAAV is administered to a subject via intravitreal injection, more preferably by a single intravitreal injection.
- a method for the treatment of a geographic atrophy in a subject in need of such treatment comprising administering to the subject a recombinant adeno-associated virus (rAAV) virion comprising: (a) a variant AAV capsid protein comprising a heterologous peptide insertion with a length of 7 to 20 amino acids covalently inserted in the GH-loop of the AAV capsid protein, wherein the peptide insertion comprises the amino acid sequence ISDQTKH (SEQ ID NO: 168) and (ii) a heterologous nucleic acid comprising a nucleotide sequence encoding an anti-C3 and/or anti-C5 antibody or antigenbinding variant thereof as herein described, said nucleotide sequence operably linked to a promoter, or administering to the subject a pharmaceutical composition comprising said rAAV virion and a pharmaceutically acceptable carrier, preferably wherein said rAAV or said pharmaceutical
- the variant AAV capsid protein comprises an amino acid sequence at least 80%, at least 85%, at least 90%, at least 95%, at least 98%, at least 99% identical to, or is
- amino acid sequence set forth as SEQ ID NO: 170 and comprises a P34A substitution and an LAISDQTKHA (SEQ ID NO: 169) peptide insertion at amino acid position 588.
- the heterologous nucleic acid encapsulated by the variant AAV capsid protein comprises a nucleotide sequence that encodes an antibody comprising one or more CDRs comprising the amino acid sequence according to Table 1 or Table 2 or comprising an amino acid sequence at least 75% identical thereto.
- the antibody comprises at least one, more preferably at least two, most preferably three CDRs, each comprising an amino acid sequence selected from those set forth as SEQ ID Nos: 54-56 or comprising an amino acid sequence at least 75% identical to an amino acid sequence selected from those set forth as SEQ ID Nos: 54-56.
- the nucleotide sequence encodes an antibody comprising an amino acid sequence selected from those set forth as SEQ ID Nos: 14 and 133-136 or comprising an amino acid sequence at least 75% identical (e.g., at least 80%, at 90%, at least 95%, at least 98%, or at least 99%) identical to an amino acid sequence selected from those set forth as SEQ ID Nos: 14 and 133-136.
- the nucleotide sequence encoding the nanobody comprises the nucleotide sequence set forth as any one of SEQ ID Nos: 133-136.
- the nucleotide sequence encodes an antibody comprising an amino acid sequence selected from those set forth as SEQ ID Nos: 137-163 or comprising an amino acid sequence at least 75% identical (e.g., at least 80%, at 90%, at least 95%, at least 98%, or at least 99% identical) to an amino acid sequence selected from those set forth as SEQ ID Nos: 137-163.
- the rAAV is administered by periocular, intravitreal, suprachoroidal and/or subretinal injection, preferably by intravitreal injection, to a subject with dry AMD and/or geographic atrophy at a dose of from about 1x10 8 vector genomes (vg)/eye to about 1x10 13 vg/eye, from abou1tx 10 9 vg/eye to about 1x 10 12 vg/eye, from about 1x 10 9 vg/eye to about 1x10 11 vg/eye or from about 6x10 9 vg/eye to about 6x10 10 vg/eye.
- vg vector genomes
- a method for treating dry AMD and/or geographic atrophy in a subject in need thereof comprising administering to the subject by periocular, intravitreal, suprachoroidal and/or subretinal injection, an effective amount of an rAAV virion
- 73 comprising (i) a capsid comprising a capsid protein comprising the amino acid sequence set forth as SEQ ID NO: 170 and (ii) a heterologous nucleic acid comprising from 5' to 3': (a) an AAV2 terminal repeat (b) a CAG or CMV promoter (c) a nucleotide sequence encoding an amino acid sequence as set forth in any of SEQ ID Nos: 14 and 133-136 (d) a polyadenylation sequence and (e) an AAV2 terminal repeat.
- a method for treating dry AMD and/or geographic atrophy in a subject in need thereof comprising administering to the subject by periocular, intravitreal, suprachoroidal and/or subretinal injection, a pharmaceutical composition comprising a pharmaceutically acceptable carrier and an rAAV virion comprising (i) a capsid protein comprising the amino acid sequence set forth as SEQ ID NO: 170 and (ii) a heterologous nucleic acid comprising from 5' to 3': (a) an AAV2 terminal repeat (b) a CAG or CMV promoter (c) a nucleotide sequence encoding an amino acid sequence as set forth in any of SEQ ID Nos: 14 and 133-136 (d) a polyadenylation sequence and (e) an AAV2 terminal repeat.
- a pharmaceutical composition comprising a pharmaceutically acceptable carrier and an rAAV virion comprising (i) a capsid protein comprising the amino acid sequence set forth as SEQ ID NO: 170 and (ii) a
- the pharmaceutical composition comprises between about 1x10 8 vg to about 1x10 13 vg, between about 1x10 9 vg to about 1x10 12 vg, between about 1x10 9 vg to about 1x10 11 vg or between about 6x10 9 vg to about 6x10 10 vg.
- a method for treating dry AMD and/or geographic atrophy in a subject in need thereof comprising administering to the subject by periocular, intravitreal, suprachoroidal and/or subretinal injection, an effective amount of an rAAV virion comprising (i) a capsid comprising a capsid protein comprising the amino acid sequence set forth as SEQ ID NO: 170 and (ii) a heterologous nucleic acid comprising from 5' to 3': (a) an AAV2 terminal repeat (b) a CAG or CMV promoter (c) a nucleotide sequence encoding an amino acid sequence as set forth in any of SEQ ID Nos:137-163 (d) a polyadenylation sequence and (e) an AAV2 terminal repeat.
- an rAAV virion comprising (i) a capsid comprising a capsid protein comprising the amino acid sequence set forth as SEQ ID NO: 170 and (ii) a heterologous nucleic acid comprising from 5
- a method for treating dry AMD and/or geographic atrophy in a subject in need thereof comprising administering to the subject by periocular, intravitreal, suprachoroidal and/or subretinal injection, a pharmaceutical composition comprising a pharmaceutically acceptable carrier and an rAAV virion comprising (i) a capsid protein comprising the amino acid sequence set forth as SEQ ID NO: 170 and (ii) a heterologous nucleic
- the pharmaceutical composition comprises between about1x 10 8 vg to about 1 x 10 13 vg, between about 1 x 10 9 vg to about 1 x 10 12 vg, between about 1 x 10 9 vg to about 1 x 10 n vg or between about 6x 10 9 vg to about 6x 10 10 vg.
- compositions comprising an rAAV as described herein are provided.
- the pharmaceutical composition comprises about 1 x 10 8 to about 1 x 10 14 vector particles or vector genomes, about 1 x 10 s to about 1 x 10 13 vector particles or vector genomes, about 1 x 10 9 to about 1 x 10 12 vector particles or vector genomes, or about 1 x 10 9 , about 2 x 10 9 , 3x 10 9 , about 4 x 10 9 , about 5 x 10 9 , about 6 x 10 9 , about 7 x IO 9 , about 8 x 10 9 , about 9 x 10 9 , about 1 x 10 10 , about 2 x 10 10 , about 3 x 10 10 , about 4 x 10 10 , about 5 x 10 10 , about 6 x 10 10 , about 7 x 10 10 , about 8 x 10 10 , about 9 x 10 10 , about 1 x 10 11 , about 2 x 10 11 , about 3
- the pharmaceutical composition comprises about 1 x 10 9 to about 1 x 10 11 vg, and preferably comprises about 6x10 9 vg to about 6x10 10 vg.
- the pharmaceutical composition is administered to a human with dry AMD and/or geographic atrophy via intravitreal injection.
- MAC retinal pigment epithelium
- Nanobodies against human complement factor C3 (SEQ ID Nos: I -14) were isolated through phage display panning against human C3 protein and tested for their ability to specifically bind to C3 and inhibit complement.
- Recombinant AAV comprising a capsid of SEQ ID NO: 107 was generated encapsulating a heterologous nucleic acid encoding a representative anti-C3 nanobody and assessed for complement inhibition.
- Recombinant protein was produced in HEK293 cells by NeoClone and GeneScript for each nanobody identified.
- HEK293T cells were grown on tissue culture plastic at 5% CO2 in normoxic conditions in DMEM, 10% fetal bovine serum (FBS), and with 1% Penicillin/Streptomycin.
- iPSC-RPE were grown on Matrigel coated plates and matured for 30 days at 5% CO2 in normoxic conditions in XVI VO- 10 media.
- iPSC-RPE were transduced ⁇ 30 days after seeding with a multiplicity of infection (MOI, vg/cell) of 5,000, 10,000 or 20,000. Samples were collected 7 days post infection for analysis.
- MOI multiplicity of infection
- the Wieslab® Complement system Alternative pathway kit (Eagle Biosciences Inc. (Cat. No. COMPL AP 330) is an enzyme immunoassay for the qualitative determination of functional alternative complement pathway in human serum and was executed according to
- the Wieslab Complement assay combines the specific activation of the pathway with the use of labeled antibodies specific for a neoepitope of the termina complement complex, C5b-9, produced as the result of complement activation (the amount of C5b-9 generated is proportional to the functional activity of the complement system).
- C5b-9 a neoepitope of the termina complement complex
- the amount of C5b-9 generated is proportional to the functional activity of the complement system.
- normal human or cynomolgus serum (Complement Tech) was diluted 1:20, along with 3.75 ug of purified nanobodies. Control and test serum was added at 100 uL/well to the provided plates. After incubating at 37C for 1 hour, plates were washed 3 times with washing solution. Prepared conjugate alkaline phosphatase-labelled antibodies to C5b-9 was added and incubated for 30 minutes at room temperature. After washing, substrate solution was added to each well and absorbance read at 405
- the plate Post incubation, the plate was washed 3 times with the wash buffer provided in the kit and incubated with the conjugate for 30 min at room temperature. Following incubation, the plate was washed 3 times with the wash buffer provided.
- One hundred micro-liter of the substrate solution provided with the kit was added to the microtiter plate and absorbance was read at 405 nm every 5 min intervals for 60 minutes. For data analysis the OD 405nm at 60 min was plotted as a function of concentration for the dose response curve.
- iPSC-RPE were exposed to either 1% human serum or 1% human serum plus 5 ⁇ g/ml Zymosan to induce the alternative complement pathway. Twenty-four hours post stimulation, cells were fixed with 4% paraformaldehyde and then permeabilized with 0.01% Triton X-100.
- rAAV with the capsid amino acid sequence listed in SEQ ID NO: 107 was produced via PEI-mediated triple transfection into HEK293 cells.
- Cells were cultured in DMEM supplemented with FBS and maintained at 37 °C in a 5% CO2 environment.
- Antigen protein was pre-coated in standard ELISA plates at 500 ng/well then blocked with 1% BSA. Nanobodies were diluted to 10 ug/mL and 150 uL were added to each well. After incubation and washing, remaining bound nanobodies were detected with HRP conjugated monoclonal rabbit anti-camelid VHH. Control wells with no nanobodies was used as background and this reading was subtracted from test wells.
- Biolayer Interferometry via the Gator or Octet machines were used to determine the binding affinities of nanobodies to their respective antigens.
- the anti-His probes were first equilibrated in buffer (PBST+5%BSA) at 37C and at 1000 ipm in a 96 well plate. After equilibration, the 6XHis tagged nanobodies (load sample) were immobilized onto the sensors. Once the probes reached 60% capacity, the loaded sensors were washed in buffer to remove any non-specifically binding. The nanobody loaded probes are then immersed in a serial dilution of untagged target antigen protein (analyte sample) for the association step to obtain “Kon”.
- PBST+5%BSA buffer
- 6XHis tagged nanobodies load sample
- the loaded sensors were washed in buffer to remove any non-specifically binding.
- the nanobody loaded probes are then immersed in a serial dilution of untagged target antigen protein (analyte sample) for
- Koff dissociation rate constant (1/s).
- KD the equilibrium dissociation rate constant which is the direct measure of binding affinity of the binding pair.
- iPSC-RPE Induced pluripotent stem ceil derived retinal pigmented epithelial cells
- rAAV capsid amino acid sequence listed in SEQ ID NO: 107 carrying a C3-N10 transgene (SEQ ID NO: 14) at three multiplicities of infection, (MOI) 5,000, 10,000 and 20,000.
- MOI multiplicities of infection
- VHH nanobody protein levels were analyzed in supernatants via ELISA and a dose response was observed in expression (Figure la).
- Nanobodies against human complement factor C5 were isolated through phage display panning against human C protein and tested for their ability to specifically bind to C5 and inhibit complement.
- the Wieslab® Complement system Alternative pathway kit (Eagle Biosciences Inc. (Cat. No. COMPL AP 330) is an enzyme immunoassay for the qualitative determination of functional alternative complement pathway in human serum and was executed according to manufacturer’s instructions.
- the Wieslab Complement assay combines the specific activation of the pathway with the use of labeled antibodies specific for a neoepitope of the termina complement complex, C5b-9, produced as the result of complement activation (the amount of C5b-9 generated is proportional to the functional activity of the complement system).
- Antigen protein was pre-coated in standard ELISA plates at 500 ng/well then blocked with 1% BSA. Nanobodies were diluted to 10 ug/mL and 150 uL were added to each well. After incubation and washing, remaining bound nanobodies were detected with HRP conjugated monoclonal rabbit anti-camelid VHH. Control wells with no nanobodies was used as background and this reading was subtracted from test wells.
- Biolayer Interferometry via the Gator or Octet machines were used to determine the binding affinities of nanobodies to their respective antigens.
- the anti-His probes were first equilibrated in buffer (PBST+5%BSA) at 37C and at 1000 rpm in a 96 well plate. After equilibration, the 6XHis tagged nanobodies (load sample) were immobilized onto the sensors. Once the probes reached 60% capacity, the loaded sensors were washed in buffer to remove any
- the nanobody loaded probes are then immersed in a serial dilution of untagged target antigen protein (analyte sample) for the association step to obtain “Kon”.
- the sensor is then transferred to a blank buffer to obtain off rate “Koff dissociation rate constant (1/s).
- Koff dissociation rate constant (1/s).
- the ratio of Koff/Kon is the “KD” is the equilibrium dissociation rate constant which is the direct measure of binding affinity of the binding pair.
- both the wild type and mutant complement sequences were cloned with a C-terminal TEV cleavage site and 6XHis tag into pcDNA plasmid with a mouse IgG secretory sequence and transiently transfected into Expi293 cells.
- the cells were harvested, spin at 250Xg for 30 min and the supernatant collected.
- the recombinant protein was purified by single step purification by using Immobilized metal affinity chromatography (IMAC) affinity chromatography.
- IMAC Immobilized metal affinity chromatography
- the clarified supernatant was passed through a 0.2-micron filter and passed through a 1 ml His excel trap column preequilibrated with equilibration buffer (50mM Tris, 150mM NaCl pH 8.0) at 1 mL/min flow rate. Following capture step, the column was washed with a wash buffer (10mM Imidazole, 50mM Tris, 150nM NaCl pH 8) and eluted with his tag elution buffer (250nM Imidazole, 50mM Tris, 150mM NaCl pH 8). The his-elution was subjected to TEV protease for 6XHis tag cleavage.
- equilibration buffer 50mM Tris, 150mM NaCl pH 8.0
- the 6XHis tag is removed by passing the eluate through IMAC resin.
- the flowthrough from IMAC column is passed size exclusion chromatography to buffer exchange into PBS (137mM NaCl, 2.7 mM KC1, 4.3mM NaH2PO4, 1.4Mm KH2PO4) and passed through 0.4 nm filter and stored at -20C in aliquots.
- the column was washed with a wash buffer (10mM Imidazole, 50mM Tris, 150nM NaCl pH 8) and eluted with his tag elution buffer (250nM Imidazole, 50mM Tris, 150mM NaCl pH 8).
- the his-elution subjected to buffer exchange into PBS and passed through 0.4 nm filter and stored at -20C in aliquots.
- the VHH cDNAs were cloned into pCDNA3 with a C-terminal human IgG2 hinge-human IgG4 Fc to express the nanobodies as Fc dimers in Expi293 cells with a mouse IgG signal peptide for secretion in the supernatants.
- the cells were harvested and centrifuged at 250 xg for 30 min to pellet the cells and supernatant and pass through a 0.45 micron membrane.
- the recombinant VHH-Fc dimer was passed through Mabselect sure column at pH 7 in IxPBS buffer to capture the VHH- Fc fusion.
- Post loading the column is washed with wash buffer and eluted using a low pH buffer, 0.1M Citrate buffer, 150mM NaCl pH 3.
- the eluate was neutralized immediately with 1 M Tris pH 8 followed by buffer exchanging into IXPBS pH 7 and aliquoted.
- the Wieslab alternative complement pathway assay system was employed for semiquantitative estimation of the inhibitory activity of the nanobodies.
- the Wieslab Alternative pathway kit is an enzyme immunoassay for the qualitative determination of functional alternative complement pathway in human serum.
- the assay combines the principles of hemolytic assay for complement activation with the use of labeled antibodies specific for neoantigen produced because of complement activation.
- the assay plate is coated with specific activators of alternate pathway activators and the diluent solution provided contains specific blocker to ensure activation of only alternate pathway.
- normal human serum was diluted 1 :20 in the diluent
- recombinantly purified nanobody was spiked in the wells in duplicates along with control and test serum and mixed on a plate shaker for 5 min at 400 rpm. Following mixing the contents of the 96 well plate is transferred into the assay plate using a multichannel pipette. After incubating at 37C for 1 hour, plates were washed 3 times with washing solution and conjugate alkaline phosphatase-labelled antibodies to C5b-9 was added and incubated for 30 minutes at room temperature. After washing, substrate solution was added to the wells and incubated at room temperature for 45 minutes. The amount of complement activation correlates with the color intensity and is measured in terms of absorbance/ optical density at 405nm.
- IC50 MAC half-maximal inhibitory concentration membrane associated complex formation
- Mouse C3/ C5 Elisa was employed to check the cross functional activity of the C3 and C5 nanobodies to mouse complement proteins.
- the assay provides a micro well plate coated with LPS for specific activation of the alternative pathway.
- the mouse serum was diluted 1 : 20 in the dilution buffer provided and transferred to 96 well plate and the serial dilutions of the nanobodies were spiked in the serum in triplicates along with a positive control mouse BB5.1 antibody.
- the 96 well plate was agitated for 5 min on plate shaker at 400 rpm and the contents of the plate transferred to the micro well plate coated with LPS and incubated at 37C for 60 min.
- the anti-strep II antibody coated probes were used for binding affinity determination by Octet/ Gator.
- the anti-strep II antibody probe was immersed in buffer (PBST+5%BSA) for equilibration at 37C and at 1000 rpm in a 96 well plate. Following equilibration, the probes were immersed in well containing VHH-strep-tagged VHH for loading. Once the probes are about 60% loaded, the anti-strep II antibody probes were immersed in a buffer well for a washing step to remove any non-specifically bound VHH to the probes.
- buffer PBST+5%BSA
- the probes are then immersed in a serial dilution of either recombinantly purified human or cynomolgus C3 or C5 wild type or the C5 single and double mutant proteins (untagged) for association step to obtain “Kon” which is the association rate constant (1/s).
- a dissociation step is performed in buffer to obtain off rate “Koff dissociation rate constant (1/s).
- KD the equilibrium dissociation rate constant which is the direct measure of binding affinity of the binding pair.
- Gator BLI platform was used to investigate binding epitope(s) of C5S2.2 nanobody. Like eculizumab, C5S2.2 nanobody did not show cross binding to cynomolgus C5 protein nor have inhibitory alternative complement activity in cynomolgus serum. Hence binding epitope was interrogated to find out whether eculizumab and C5S2.2 cross-block each other to human C5.
- a sandwich approach was used on BLI where anti-human Fc-binding probes were used to immobilize eculizumab for 1-2 min. After washing the pre-loaded probes were then dipped into untagged C5 alone or C5 preincubated with C5S2 in the recommended binding buffer for 5-6 min to allow for association of the either the C5 or C5S2-C5 complex.
- a reverse sandwich was performed by immobilizing dilution series of C5S2 nanobody on the anti-strep- II probes. After washing, this pre-loaded probe was immersed into C5 containing binding solution for an to check for letting the C5 protein associate with immobilized nanobody for 2-3 min. Once the C5 protein is associated the probe holding the VHH-C5 complex was immersed in the well containing eculizumab for 5-6 minutes for association.
- hC3 was cross linked with DDS for 180 min and cleaved by five different proteases (trypsin, Chymotrypsin Elastase, Asp-N and Thermolysin) and measured on for maximum sequence coverage. Post cleavage both the control and crosslinked samples are subjected to Liquid chromatography for peptide separation followed by mass analysis on Q- Exactive Plus mass spectrometer. Resulting peptides were analyzed by using CovalX proprietary software and peptide fingerprint map for both the cross linked and control sample was generated.
- proteases trypsin, Chymotrypsin Elastase, Asp-N and Thermolysin
- Miniprep cultures were grown from the resulting colonies following cloning and transformation, in Terrific Broth, supplemented with Kanamycin Sulphate at 100 mg/mL and grown at 30°C overnight with shaking. DNA was isolated with QIAprep Spin Miniprep Kit following manufacturer’s protocol and sequenced using Primordium Lab’s sequencing technology to identify positive clones of sCFH and CFH.
- HEK293T cells were grown in DMEM media with 1% Penicillin/Streptomycin and 10% fetal bovine serum. Cells were passaged twice a week and not used beyond passage 20.
- iPSC-RPE were grown on Matrigel coated plates and matured for 30 days at 5% CO2 in normoxic conditions in XVIVO-10 media supplemented with 10 pM RHO/ROCK pathway inhibitor Y-27632.
- Recombinant rAAVs (comprising a capsid of SEQ ID NO:170), for in vitro studies were produced via PEI -mediated triple transfection into HEK293 cells.
- Cells were cultured in DMEM supplemented with FBS and maintained at 37 °C in a 5% CO2 environment.
- HEK293T cells were seeded in 12-well plates at 2x 10 5 cells/well and transfected with various plasmid versions of C3N10, C3S3 and C5S2. Supernatants were collected and tested for transgene expression by various assays.
- iPSC-RPE were transduced ⁇ 30 days after seeding with a multiplicity of infection (MOI, vg/cell) of 1,000, 5,000 or 20,000. Supernatant samples were collected 7 days post infection for analysis.
- MOI multiplicity of infection
- Anti-VHH capture antibody was diluted 1 :500 in PBS and incubated in a MaxiSorp microtiter plate overnight at 4 °C. The plate was then washed 5x with PBST and incubated in a blocking solution of 2% BSA in PBS at room temperature for 2 hours. Plate washing was likewise performed between each following step unless otherwise noted. Supernatants from transduced cells were diluted in 1% BSA in PBS; dilutions were uniform across all samples of each VHH tested. Diluted samples were applied to the microtiter plate and incubated at room temperature for 2 hours.
- the plate was next incubated with HRP-conjugated anti-VHH detection antibody diluted 1 : 10,000 in 1% BSA in PBST for 1 hour at room temperature. Following incubation with detection antibody, 100 ⁇ l TMB substrate was applied to the plate and color development was monitored. 100 ⁇ l 450 nM TMB stop solution was applied to the plate without a wash step 5-15 minutes after addition of TMB substrate. The optical density (OD) was measured with a Cytation 3 photometer at 450 run within 15 minutes of applying the stop solution.
- Protein concentrations of cell supernatants were quantified using the Pierce Micro BCA Protein assay according to the manufacturer’s instructions. Appropriate volume of supernatant to achieve 33 ⁇ g of total protein was then diluted in PBS to 15 ⁇ l and mixed with 5.78 ⁇ l 4x LDS buffer and 2.31 ⁇ l 10x Reducing agent and incubated at 90 °C for 10 minutes. Samples were loaded on a 15-well Bolt 12% Bis-Tris Plus polyacrylamide gel or Novex 10 to 20% Tricine polyaciylamide gels and run in 1 x MOPS buffer at 160 volts for 45 minutes.
- the Wieslab® Complement system Alternative pathway kit is an enzyme immunoassay for the qualitative determination of functional alternative complement pathway in human serum and was executed according to manufacturer’s instructions.
- Normal human serum was diluted to 2% in the provided dilution buffer and 200 ⁇ l of the 2% serum was transferred to a microtiter plate. 20 ⁇ l supernatant samples from transduced RPE cultures were applied to the plate and mixed. Following mixing, the diluted serum with added supernatant samples was transferred in duplicate to the assay plate, 100 ⁇ l per well, and incubated for 60 min at 37°C. Post incubation, the plate was washed 3 times with the wash buffer provided in the kit and incubated with the conjugate for 30 min at room temperature.
- 92 cynomolgus C3 and C5 complement protein targets were calculated via biolayer interferometry (BLI) using the Gator. Not only do neither 03 nor C5 targeted wild type nanobodies cross-bind to the other antigen, but none of them also associate with other members of the alpha-2 -macroglobulin family: human C4b, alpha-2-macroglobulin, and alpha-2 - macroglobuline-like-1 protein (data not shown).
- the innate immune complement system is initiated by three separate pathways: the classical, lectin, and alternative pathways.
- the C3 protein acts as the point of convergence of each pathway’s activation, while C5 protein cleavage initiates the formation of the final component of complement activation: the membrane attack complex.
- the wild type nanobodies functional potential were evaluated in classical, alternative, and classical pathway assay kits in human serum available from Wieslab. As shown in Figure 2, all three nanobodies have inhibitory activity against the alternative pathway. However, while C5S2 has strong inhibitory activity in all three pathways, C3S3 only has activity against the alternative pathway. C3N10 shows most inhibitory activity against the alternative pathway and moderate potency against both the classical and lectin pathways.
- C3N 10.3, C3 S3.2, and C5 S2.2 were engineered and expressed as homo-dimeric fusion proteins (aka, C3N10.3-Fc, C3S3.2-Fc and C5S2.2-Fc) with the human G2 hinge-G4 Fc domain to emulate native immunoglobulin bivalent structure and function.
- Their respective binding rate to target and alternative pathway inhibition potency was calculated using biolayer interferometry and the Wieslab assays and compared to the original monomers in Table 8 and 9, respectively.
- Avidity association was observed with C3S3.2-Fc and C5S2.2-Fc while synergistic alternative pathway inhibitory activity (greater than two-fold) appears to be seen with C5S2-Fc ( Figure 7).
- cross-linking mass spectrometry (XL-MS, CovalX) was used to identify C3N10.3 and C3S3.2 binding epitope. Both humanized nanobodies and purified C3 human protein were provided.
- Tables of cross-linked peptides between the nanobody and antigen along with illustrated aminoacid interactions are shown in Table 10 and Figure 8 for C3N10.3 and Table 11 and Figure 9 for
- GVDYVYKTRLVKVQLSNDFDEYIMAIEQTIKSGSDEVQVGQQRTFISPIKCREALKLEEK K (SEQ ID NO:171)), while C3S3.2 appears to associate with two non-linear epitopes in the beta-chain across the MG1 (110-130: ATFGTQVVEKVVLVSLQSGYL (SEQ ID NO: 172) and MG5 (490-510: YTYLIMNKGRLLKAGRQVREP (SEQ ID NO:173) domains, Figure 10.
- eculizumab like C5S2, binds to human C5 protein, inhibits all three complement pathways in human serum, and neither binds to cynomolgus C5 nor inhibits complement pathways in cynomolgus serum (data not shown).
- the Gator was used to investigate whether eculizumab and C5S2 also cross-block each other to human C5.
- C5S2 in the recommended binding buffer.
- C5 protein alone bound rapidly to the eculizumab preloaded tips as expected while the C5 preincubated with C5S2 did not, indicating the latter prebound nanobody was blocking the former full-size antibody’s epitope.
- Eculizumab is refractory to two naturally occurring variants: the R885H human polymorphism and the W917S human vs cynomolgus divergence (Brachet G et.al. 2016; Nishimura J et.al. 2014).
- Human C5 R885H, human C5 W917S, and cynomolgus C5 S917W mutant proteins were recombinantly produced mid C5S2.2’s and eculizumab’s affinity to them was compared using the Gator.
- Figures 13A-C show, while C5S2.2 had high affinity for the wild-type human C5 protein, it lost all binding activity to human C5 R885H and W917S mutants. On the other hand, while it was unable to bind to wild type cynomolgus C5, C5S2.2 had high affinity to the mutant cynomolgus C5 S917W. This pattern of interaction to wild-type vs mutant was reflected by eculizumab, as expected, in Figures 14A-C, indicating that C5S2.2 and eculizumab have similar binding epitopes on human 05.
- vascular endothelial growth factor A 98 vascular endothelial growth factor A while Cablivi (caplacizumab) is a tri-alanine-linked homodimeric nanobody against the Al domain of von Willebrand factor.
- VHH containing plasmids were assessed for function via western blot assays.
- C3 gets broken down into C3a and C3b.
- C3 breakdown triggers activation of C5 degradation resulting in C5a and C5b.
- a feedback loop of C3 activation can be initiated by this activation and result in Factor B breakdown.
- C3 and C5 VHH inhibitors can be assessed for function to stop these degradation products, which can be visualized by western blot.
- VHH within the AAV supportive plasmid backbone were manufactured into AAV.
- Lead VHH AAV were characterized by expression and activity following transduction at three MOI in iPSC-RPE.
- Figure 17 depicts a Western blot of the supernatant collected post transduction, showing a single product at the expected size of 15kDa for each VHH examined ( Figure 17).
- VHH expression in the supernatant was also determined by ELISA ( Figure 18). Each VHH exhibited a dose response as indicated by an increase in OD at 450nm with increasing dose.
Landscapes
- Chemical & Material Sciences (AREA)
- Organic Chemistry (AREA)
- Health & Medical Sciences (AREA)
- Medicinal Chemistry (AREA)
- Biochemistry (AREA)
- Biophysics (AREA)
- General Health & Medical Sciences (AREA)
- Genetics & Genomics (AREA)
- Life Sciences & Earth Sciences (AREA)
- Molecular Biology (AREA)
- Proteomics, Peptides & Aminoacids (AREA)
- Immunology (AREA)
- Medicines Containing Material From Animals Or Micro-Organisms (AREA)
- Medicines That Contain Protein Lipid Enzymes And Other Medicines (AREA)
- Micro-Organisms Or Cultivation Processes Thereof (AREA)
- Peptides Or Proteins (AREA)
Abstract
Description
Claims
Priority Applications (3)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| AU2024274562A AU2024274562A1 (en) | 2023-05-23 | 2024-05-22 | Novel complement system inhibiting antibodies |
| IL324504A IL324504A (en) | 2023-05-23 | 2025-11-06 | Complement system inhibiting antibodies |
| MX2025013869A MX2025013869A (en) | 2023-05-23 | 2025-11-20 | Novel complement system inhibiting antibodies |
Applications Claiming Priority (4)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| US202363503745P | 2023-05-23 | 2023-05-23 | |
| US202363503768P | 2023-05-23 | 2023-05-23 | |
| US63/503,745 | 2023-05-23 | ||
| US63/503,768 | 2023-05-23 |
Publications (2)
| Publication Number | Publication Date |
|---|---|
| WO2024243292A2 true WO2024243292A2 (en) | 2024-11-28 |
| WO2024243292A3 WO2024243292A3 (en) | 2025-04-17 |
Family
ID=93590362
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/US2024/030542 Pending WO2024243292A2 (en) | 2023-05-23 | 2024-05-22 | Novel complement system inhibiting antibodies |
Country Status (4)
| Country | Link |
|---|---|
| AU (1) | AU2024274562A1 (en) |
| IL (1) | IL324504A (en) |
| MX (1) | MX2025013869A (en) |
| WO (1) | WO2024243292A2 (en) |
Cited By (1)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN119409815A (en) * | 2025-01-07 | 2025-02-11 | 天辰生物医药(苏州)有限公司 | Anti-human complement C5 VHH antibody and its application |
Family Cites Families (3)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| EP2474632B1 (en) * | 2002-12-20 | 2015-08-12 | Celera Corporation | Genetic polymorphisms associated with myocardial infarction, methods of detection and uses thereof |
| GB2583560A (en) * | 2018-12-11 | 2020-11-04 | Admirx Inc | Fusion protein constructs for complement associated disease |
| US20230416942A1 (en) * | 2020-08-21 | 2023-12-28 | Yale University | Nanobody Compositions and Methods of Use of the Same |
-
2024
- 2024-05-22 WO PCT/US2024/030542 patent/WO2024243292A2/en active Pending
- 2024-05-22 AU AU2024274562A patent/AU2024274562A1/en active Pending
-
2025
- 2025-11-06 IL IL324504A patent/IL324504A/en unknown
- 2025-11-20 MX MX2025013869A patent/MX2025013869A/en unknown
Cited By (1)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN119409815A (en) * | 2025-01-07 | 2025-02-11 | 天辰生物医药(苏州)有限公司 | Anti-human complement C5 VHH antibody and its application |
Also Published As
| Publication number | Publication date |
|---|---|
| IL324504A (en) | 2026-01-01 |
| MX2025013869A (en) | 2026-01-07 |
| AU2024274562A1 (en) | 2025-11-27 |
| WO2024243292A3 (en) | 2025-04-17 |
| AU2024274562A9 (en) | 2025-12-11 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| CN112074605B (en) | Adeno-associated virus (AAV) delivery of anti-FAM19A5 antibodies | |
| KR20190141223A (en) | Antibody Production Methods to Minimize Disulfide Bond Reduction | |
| KR20140019385A (en) | Bi- and monospecific, asymmetric antibodies and methods of generating the same | |
| JP2025098167A (en) | Methods for treating antibody-mediated disorders with fcrn antagonists | |
| US12168696B2 (en) | Antibodies to mucin-16 (MUC16), encoding polynucleotides and methods of treating MUC16-positive cancer | |
| KR20230025672A (en) | Antibodies that bind to CD3 and CD19 | |
| WO2022156653A1 (en) | Expression vector for anti-sars-cov-2 neutralizing antibodies | |
| WO2024243292A2 (en) | Novel complement system inhibiting antibodies | |
| WO2020260315A1 (en) | Deimmunized antibodies binding to alpha-4 integrin and uses thereof | |
| US20250376526A1 (en) | Fc VARIANT WITH ENHANCED AFFINITY TO Fc RECEPTORS AND IMPROVED THERMAL STABILITY | |
| CN114364695B (en) | Antibodies specifically recognizing C5A and their applications | |
| EP4399226A1 (en) | Antibodies binding to tetanus toxin and uses thereof | |
| KR20260016923A (en) | Novel complement system inhibitory antibodies | |
| TW202241943A (en) | Tau-specific antibody gene therapy compositions, methods and uses thereof | |
| JP2021508472A (en) | Anti-VEGF antibody and method of use | |
| CN117203230A (en) | Antibodies that specifically recognize C5A and their applications | |
| JP7781174B2 (en) | Monoclonal antibodies against CLDN18.2 and Fc-engineered versions thereof | |
| US20240034772A1 (en) | Antibodies binding to f-protein of metapneumovirus and uses thereof | |
| CN121311247A (en) | Antibodies that specifically bind MASP3 and multispecific antibodies that specifically bind MASP3 and MASP2 | |
| CN116848133A (en) | Antibodies that bind metapneumovirus F protein and their uses | |
| CN119101165A (en) | Antibodies that specifically bind to MASP3 and multispecific antibodies that specifically bind to MASP3 and MASP2 | |
| CN112262154A (en) | Novel IGG5-like recombinant immunoglobulin encoded by human heavy chain pseudo-γ gene | |
| CN117377685A (en) | Fc variants with enhanced affinity for Fc receptors and improved thermal stability |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| WWE | Wipo information: entry into national phase |
Ref document number: 324504 Country of ref document: IL |
|
| WWE | Wipo information: entry into national phase |
Ref document number: AU2024274562 Country of ref document: AU Ref document number: 827185 Country of ref document: NZ |
|
| WWE | Wipo information: entry into national phase |
Ref document number: MX/A/2025/013869 Country of ref document: MX |
|
| WWE | Wipo information: entry into national phase |
Ref document number: 202547114914 Country of ref document: IN |
|
| ENP | Entry into the national phase |
Ref document number: 2024274562 Country of ref document: AU Date of ref document: 20240522 Kind code of ref document: A |
|
| WWP | Wipo information: published in national office |
Ref document number: 827185 Country of ref document: NZ |
|
| WWE | Wipo information: entry into national phase |
Ref document number: 1020257040281 Country of ref document: KR |
|
| REG | Reference to national code |
Ref country code: BR Ref legal event code: B01A Ref document number: 112025025559 Country of ref document: BR |
|
| WWP | Wipo information: published in national office |
Ref document number: 202547114914 Country of ref document: IN |
|
| WWE | Wipo information: entry into national phase |
Ref document number: 2024811828 Country of ref document: EP Ref document number: 2025137194 Country of ref document: RU |
|
| NENP | Non-entry into the national phase |
Ref country code: DE |
|
| WWP | Wipo information: published in national office |
Ref document number: 324504 Country of ref document: IL |
|
| WWP | Wipo information: published in national office |
Ref document number: MX/A/2025/013869 Country of ref document: MX |
|
| 121 | Ep: the epo has been informed by wipo that ep was designated in this application |
Ref document number: 24811828 Country of ref document: EP Kind code of ref document: A2 |
|
| WWP | Wipo information: published in national office |
Ref document number: 2025137194 Country of ref document: RU |