WO2024229398A1 - Artificial amalgavirus satellite rnas - Google Patents
Artificial amalgavirus satellite rnas Download PDFInfo
- Publication number
- WO2024229398A1 WO2024229398A1 PCT/US2024/027769 US2024027769W WO2024229398A1 WO 2024229398 A1 WO2024229398 A1 WO 2024229398A1 US 2024027769 W US2024027769 W US 2024027769W WO 2024229398 A1 WO2024229398 A1 WO 2024229398A1
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- rna
- virus
- hrv
- sequence
- dna
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Pending
Links
Classifications
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/63—Introduction of foreign genetic material using vectors; Vectors; Use of hosts therefor; Regulation of expression
- C12N15/79—Vectors or expression systems specially adapted for eukaryotic hosts
- C12N15/82—Vectors or expression systems specially adapted for eukaryotic hosts for plant cells, e.g. plant artificial chromosomes (PACs)
- C12N15/8201—Methods for introducing genetic material into plant cells, e.g. DNA, RNA, stable or transient incorporation, tissue culture methods adapted for transformation
- C12N15/8202—Methods for introducing genetic material into plant cells, e.g. DNA, RNA, stable or transient incorporation, tissue culture methods adapted for transformation by biological means, e.g. cell mediated or natural vector
- C12N15/8203—Virus mediated transformation
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/63—Introduction of foreign genetic material using vectors; Vectors; Use of hosts therefor; Regulation of expression
- C12N15/79—Vectors or expression systems specially adapted for eukaryotic hosts
- C12N15/82—Vectors or expression systems specially adapted for eukaryotic hosts for plant cells, e.g. plant artificial chromosomes (PACs)
- C12N15/8241—Phenotypically and genetically modified plants via recombinant DNA technology
- C12N15/8261—Phenotypically and genetically modified plants via recombinant DNA technology with agronomic (input) traits, e.g. crop yield
Definitions
- Amalgaviruses are double-stranded (ds) RNA viruses having a monopartite genome which encodes an RNA dependent RNA polymerase (RdRP) and a putative capsid protein (Krupovic et al., Biol Direct.2015 Mar 29;10:12. doi: 10.1186/s13062-015-0047-8).
- RNA molecules comprising from 5’ terminus to 3’ terminus: (a) a 5’ RNA replication element recognized by an amalgavirus RNA-dependent RNA polymerase (RdRP); (b) a cargo RNA molecule (cargo RNA sequence); and (c) a 3’ RNA replication element recognized by the RdRP; wherein the 5’ RNA replication element, the cargo RNA molecule, and the 3’ RNA replication element are operably linked, and wherein the cargo RNA molecule is heterologous to the 5’ RNA replication element and the 3’ RNA replication element.
- RdRP amalgavirus RNA-dependent RNA polymerase
- the 5’ RNA replication element and the 3’ RNA replication element are obtained from the same amalgavirus genome or from amalgavirus genomes having at least 85%, 90%, 95%, 98%, or 99% sequence identity to one another and which are optionally related;
- the 5’ RNA replication element, the 3’ RNA replication element, and the RdRP are obtained from the same amalgavirus genome or from amalgavirus genomes having at least 85%, 90%, 95%, 98%, or 99% sequence identity to one another and which are optionally related; or
- the 5’ RNA replication element, the 3’ RNA replication element, and/or the RdRP coding region are obtained from different amalgavirus genomes, and the members of each respective set of the 5’ RNA replication elements, 3’ RNA replication elements, and/or RdRP coding regions have at least 85%, 90%, 95%, 98%, or 99% sequence identity to one another.
- the RNA molecule comprises: at least one heterologous RNA virus (HRV) amplicon in sense or antisense orientation to the first 5’ RNA replication element comprising: I. (i) a heterologous RNA virus (HRV) 5’ replication region (HRV 5’RR); (ii) the cargo RNA molecule; and Agent Ref: P14357WO00 - 2 - (iii) the heterologous RNA virus (HRV) 3’ RNA replication region (HRV 3’RR); wherein the HRV 5’ RR and HRV 3’ RR HRV are recognized by a heterologous RNA virus RNA-dependent RNA polymerase (hrvRdRP); and wherein the HRV 5’RR, cargo RNA molecule, and HRV 3’RR are operably linked; or II.
- HRV heterologous RNA virus
- heterologous RNA virus (HRV) subgenomic promoter operably linked to the cargo RNA molecule; wherein the subgenomic promoter is recognized by a heterologous RNA virus RNA-dependent RNA polymerase (hrvRdRP).
- HRV heterologous RNA virus RNA-dependent RNA polymerase
- Agricultural formulations as well as bacterial, fungal, plant, insect, and invertebrate animal cells comprising the herein disclosed recombinant RNAs are also provided.
- RNA molecules comprising the herein disclosed recombinant RNA molecules; and a cell containing the recombinant RNA molecule and an RdRP protein that recognizes the 5’ and 3’ RNA replication elements of the recombinant RNA molecule.
- the cell further comprises one or more of: (i) a viral capsid protein (CP); (ii) an RNA-binding protein (RBP) that binds to the RNA molecule, optionally wherein the RBP binds to an RNA effecter; (iii) an RNA cleavage agent that cleaves the RNA molecule; (iv) a second RNA-dependent RNA polymerase (2 nd RdRP) protein that recognizes an HRV 5’ or 3’ replication region and/or a subgenomic promoter in the RNA molecule; (v) a viral movement protein (MP); (vi) a heterologous RNA virus (HRV); or (vii) an hrvRdRP, optionally wherein the hrvRdRP recognizes the HRV 5’ or 3’ replication region and/or the subgenomic promoter.
- CP a viral capsid protein
- RBP RNA-binding protein
- RBP RNA-binding protein
- methods of establishing a synthetic amalgavirus satellite RNA in a plant cell comprising: providing to a plant cell any of the herein disclosed recombinant RNA molecules, wherein the plant cell comprises an RdRP protein that recognizes the 5’ RNA replication element and 3’ RNA replication element, wherein the RNA molecule optionally comprises an encapsidation recognition element (ERE) and is or can be encapsidated by a capsid protein, whereby the RdRP protein catalyzes synthesis of the synthetic amalgavirus satellite RNA from the recombinant RNA molecule.
- ERP encapsidation recognition element
- Also provided are methods of obtaining a phenotypic change in a plant or plant cell comprising: providing to a plant or plant cell any of the herein disclosed recombinant RNA molecules, wherein the cargo RNA molecule comprises RNA that effects a phenotypic change in the plant or plant cell in comparison to a plant or plant cell lacking the recombinant RNA, wherein the plant or plant cell comprises an RdRP protein that recognizes the 5’ RNA replication element and 3’ RNA replication element and catalyzes synthesis of a synthetic amalgavirus RNA from the recombinant RNA molecule, and wherein and the cargo RNA molecule effects the phenotypic change.
- the methods further comprise providing an hrvRdRP to the plant which recognizes the HRV 5’ or 3’ replication region and/or the subgenomic promoter in the synthetic amalgavirus satellite RNA, optionally wherein the hrvRdRP is provided by introducing a recombinant DNA or RNA encoding the hrvRdRP into the plant or a part thereof.
- Agent Ref P14357WO00 - 3 - [0010] Also provided are methods of manufacturing a synthetic amalgavirus satellite particle, comprising combining any of the herein disclosed recombinant RNA molecules with a viral capsid protein, wherein the recombinant RNA molecule comprises an encapsidation recognition element (ERE), and wherein the ERE provides for encapsidation of the RNA by the viral capsid protein.
- ERE encapsidation recognition element
- plant propagules comprising any of the herein disclosed recombinant RNA molecules and an amalgavirus RdRP, optionally wherein the plant propagule further comprises a heterologous RNA virus RdRP which recognizes an HRV 5’ or 3’ replication region and/or a subgenomic promoter in the synthetic amalgavirus satellite RNA.
- the plant propagule further comprises a heterologous RNA virus RdRP which recognizes an HRV 5’ or 3’ replication region and/or a subgenomic promoter in the synthetic amalgavirus satellite RNA
- the heterologous RNA virus RdRP is an Alphaflexivirus, Betaflexivirus, Bromovirus, Celavirus, Closterovirus, Comovirus, Potexvirus, Potyvirus, Tobamovirus, Tombusvirus, Tospoviridae, Trivirinae, Tymovirus, Varicosavirus, or Secoviridae RdRP.
- plants comprising any of the herein disclosed recombinant RNA molecules and an amalgavirus RdRP, optionally wherein the plant propagule further comprises a heterologous RNA virus RdRP which recognizes an HRV 5’ or 3’ replication region and/or a subgenomic promoter in the synthetic amalgavirus satellite RNA.
- Amalgavirus satellite systems that are self-replicating when introduced into a plant or plant cell, comprising: (1) any of the herein disclosed recombinant amalgavirus satellite RNAs (e.g., recombinant RNA molecules); and (2) an exogenous amalgavirus that is capable of replication in the plant or plant cells and that encodes the amalgavirus RdRP that recognizes the 5’ and 3’ replicase recognition sequences in the recombinant amalgavirus satellite RNA, optionally wherein the amalgavirus satellite system further comprises a heterologous RNA virus RdRP which recognizes an HRV 5’ or 3’ replication region and/or a subgenomic promoter in the synthetic amalgavirus satellite RNA.
- any of the herein disclosed recombinant amalgavirus satellite RNAs e.g., recombinant RNA molecules
- an exogenous amalgavirus that is capable of replication in the plant or plant cells and that encodes the am
- Figure 1 shows a non-limiting embodiment of a structure of an amalgavirus satellite construct.
- the 5’ RNA replication element is labelled “5’ RRE” and the 3’ RNA replication element is labeled “3’ RRE.”
- Figure 2 shows non-limiting embodiments of an amalgavirus satellite construct containing a heterologous RNA virus (HRV) amplicon comprising: (i) a heterologous RNA virus (HRV) 5’ replication region (HRV 5’RR); (ii) the cargo RNA molecule; and (iii) the heterologous RNA virus (HRV) 3’ RNA replication region (HRV 3’RR).
- HRV heterologous RNA virus
- cleavable sequence is located: (i) 5’ to the 5’ RNA replication element or 3’ to the 3’ RNA replication element; and/or (ii) between the 3’ end of the 5’ RNA replication element and the HRV amplicon and/or between the HRV amplicon and the 5’ end of the 3’ RNA replication element.
- the cleavable sequence is optionally a self-cleaving ribozyme, a self-cleaving inducible ribozyme, or an Agent Ref: P14357WO00 - 4 - siRNA or miRNA recognition site.
- a subgenomic promoter and/or an IRES is/are operably linked to the cargo RNA.
- Figure 3 shows non-limiting embodiments of an amalgavirus satellite construct containing a heterologous RNA virus (HRV) amplicon comprising a heterologous RNA virus (HRV) subgenomic promoter (HRV sgp) which is recognized by a heterologous RNA virus RNA-dependent RNA polymerase (hrvRdRP) and which is operably linked to the cargo RNA molecule.
- HRV sgp and cargo RNA are in sense and antisense orientation relative to the amalgavirus 5’ RRE are shown.
- Figure 4 depicts an amalgavirus satellite construct comprising heterologous RNA virus (HRV) subgenomic promoters (HRV sgp) with: (i) one HRV sgp operably linked to a cargo RNA; and (ii) one HRV sgp operably linked to RNA encoding an hrvRdRP which recognizes both of the HRV sgp (i.e., can drive expression of the operably linked hrvRdRP and cargo RNA).
- HRV sgp heterologous RNA virus
- HRV sgp heterologous RNA virus subgenomic promoters
- an IRES is operably linked to the cargo RNA and/or an IRES is operably linked to the RNA encoding the hrvRdRP.
- FIG. 5 depicts non-limiting embodiments of an amalgavirus satellite construct containing a heterologous RNA virus (HRV) amplicon comprising a heterologous RNA virus (HRV) subgenomic promoter (HRV sgp) which is recognized by a heterologous RNA virus RNA-dependent RNA polymerase (hrvRdRP) and which is operably linked to the cargo RNA molecule flanked by HRV 5’RR and HRV 3’ RR.
- HRV sgp and cargo RNA are in sense or antisense orientation relative to the amalgavirus 5’ RRE are shown.
- the HRV 5’ RR and 3’ RR which flank the cargo RNA provide for hrvRdRP-mediated replication of an RNA comprising from 5’ to 3’ the HRV 5’ RR, cargo RNA, and HRV 3’ RR.
- the HRV 5’ RR and 3’ RR are flanked by ribozymes.
- Figure 6 depicts a commensal satellite with a cargo RNA molecule including an HRV (“HRV1”, e.g., tobacco mosaic virus, TMV) amplicon designed to be amplified by the HRV (“HRV1”, e.g., TMV) RdRP binding to either the replication regions or to the subgenomic promoter, where the commensal satellite is an amalgavirus satellite.
- HRV1 tobacco mosaic virus
- RdRP RNA encoding the HRV RdRP
- solid squares which can further amplify the HRV amplicon, as well as RNA encoding another cargo (solid circles). In the absence of the commensal virus, no amplification of the commensal satellite occurs.
- Figure 7 depicts a commensal satellite with a cargo RNA molecule including an HRV (HRV1, e.g., tobacco mosaic virus, TMV) amplicon designed to be amplified by the HRV RdRP (“HRV1 RdRP”, solid squares) binding to the HRV1 replication regions, where the commensal satellite is an amalgavirus satellite.
- HRV tobacco mosaic virus
- the resulting transcripts include RNA encoding the HRV (HRV1) RdRP which can further amplify the HRV amplicon.
- the HRV1 amplicon includes sequence for a HRV2 amplicon (indicated in italicized text), encoding a coding and/or noncoding cargo (solid circles) and designed to be amplified in the presence of a second acute viral RdRP (“HRV2 RdRP”, hexagonal symbol), which can be provided, e.g., by introduction of a second acute virus (“HRV2”, e.g., cowpea mosaic virus, CPMV) into the plant. In the absence of the commensal virus, no amplification of the commensal satellite occurs.
- HRV2 RdRP hexagonal symbol
- Figure 8 depicts a commensal satellite with a cargo RNA molecule including an HRV (“HRV1”, e.g., tobacco mosaic virus, TMV) amplicon designed to be amplified by the HRV (“HRV1”, e.g., TMV) RdRP binding to either of two subgenomic promoters, where the commensal satellite is an amalgavirus satellite.
- HRV1 tobacco mosaic virus
- TMV tobacco mosaic virus
- the resulting transcripts include RNA encoding the HRV RdRP (“HRV1 RdRP”, solid squares), which can further amplify the HRV amplicon, as well as RNA encoding a noncoding RNAi cargo, the sense and antisense strands of which are formed during the amplification process to yield a double-stranded RNA molecule (dsRNA) for silencing of a target gene.
- HRV1 RdRP solid squares
- RNA encoding a noncoding RNAi cargo the sense and antisense strands of which are formed during the amplification process to yield a double-stranded RNA molecule (dsRNA) for silencing of a target gene.
- dsRNA double-stranded RNA molecule
- the term “and/or” as used in a phrase such as “A and/or B” herein is intended to include “A and B,” “A or B,” “A” (alone), and “B” (alone).
- the term “and/or” as used in a phrase such as “A, B, and/or C” is intended to encompass each of the following embodiments: A, B, and C; A, B, or C; A or C; A or B; B or C; A and C; A and B; B and C; A (alone); B (alone); and C (alone).
- F 1 ,” “F 2 ,” and the like refer to plants or seed obtained from a parent plant which has been selfed or that has been crossed to another plant.
- heterologous when used to describe a first element in reference to a second element means that the first element and second element do not exist in nature disposed as described.
- a heterologous nucleic acid molecule or sequence is a nucleic acid molecule or sequence that (a) is not native to a cell in which it is expressed, (b) is linked or fused to a nucleic acid molecule or sequence with which it is not linked to or fused to in nature, or with which it is not linked to or fused to in nature in the same way, (c) has been altered or mutated by the hand of man relative to its native state, or (d) has altered expression as compared to its native expression levels under similar conditions.
- a “heterologous promoter” is used to drive transcription of a sequence that is not one that is natively transcribed by that promoter (e.g., a eukaryote promoter used to drive transcription of a DNA molecule encoding an amalgavirus RNA sequence); thus, a “heterologous promoter” sequence can be included in an expression construct by a recombinant nucleic acid technique.
- a recombinant polynucleotide such as those provided by this disclosure includes genetic sequences of two or more different amalgaviruses, which genetic sequences are “heterologous” in that they would not naturally occur together.
- heterologous refers to a molecule or to a discrete part of a molecule; for example, referring to a cargo RNA molecule (e.g., a nucleic acid such as a protein-encoding RNA, an ssRNA, a regulatory RNA, an interfering RNA, or a guide RNA), which can be part of a larger molecule, or referring to a structure (e.g., structures including a promoter (e.g., for a DNA dependent RNA polymerase) or subgenomic promoter (e.g., for an RNA-dependent RNA polymerase), an RNA effecter, RNA cleavage agent recognition site, or a polynucleotide comprising or Agent Ref: P14357WO00 - 6 - encoding an expression-enhancing element, encapsidation recognition element (ERE), selectable or scoreable marker, DNA aptamer, RNA aptamer; a transcription factor binding
- IRES internal ribosome entry site
- An IRES element is generally between 100-800 nucleotides.
- IRES encephalomyocarditis virus IRES
- ECMV encephalomyocarditis virus
- maize hsp101 IRES 5’UTR crucifer infecting tobamovirus crTMV CR-CP 148 IRES
- tobacco etch virus (TEV) IRES 5’UTR hibiscus chlorotic ringspot virus (HCRSV) IRES.
- HCRSV hibiscus chlorotic ringspot virus
- an IRES sequence is derived from non-plant eukaryotic virus sequences that include but are not limited to: acute bee paralysis virus (ABPV), classical swine fever virus (CSFV), coxsackievirus B3 virus (CVB3), encephalomyocarditis virus (ECMV), enterovirus 71 (E71), hepatitis A virus (HAV), human rhinovirus (HRV2), human rhinovirus (HRV2), human lymphotropic virus (HTLV), polyoma virus (PV), and Zea mays (ZmHSP101).
- a virus A virus
- HRV2 human rhinovirus
- HRV2 human rhinovirus
- HTLV human lymphotropic virus
- PV polyoma virus
- ZmHSP101 Zea mays
- the phrase “operably linked” refers to a juxtaposition wherein the components so described are in a relationship permitting them to function in their intended manner.
- a promoter is operably linked to a coding sequence if the promoter provides for transcription or expression of the coding sequence.
- percent identity refers to percent (%) sequence identity with respect to a reference polynucleotide or polypeptide sequence following alignment by standard techniques. Alignment for purposes of determining percent nucleic acid or amino acid sequence identity can be achieved in various ways that are within the capabilities of one of skill in the art, for example, using publicly available computer software such as BLAST, BLAST-2, PSI-BLAST, or Megalign software.
- the software is MUSCLE (Edgar, Nucleic Acids Res., 32(5): 1792-1797, 2004).
- MUSCLE Nucleic Acids Res., 32(5): 1792-1797, 2004.
- percent sequence identity values are generated using the sequence comparison computer program BLAST (Altschul et al. (1990) J. Mol. Biol., 215:403-410).
- the percent sequence identity of a given nucleic acid or amino acid sequence, A, to, with, or against a given nucleic acid or amino acid sequence, B, (which can alternatively be phrased as a given Agent Ref: P14357WO00 - 7 - nucleic acid or amino acid sequence, A that has a certain percent sequence identity to, with, or against a given nucleic acid or amino acid sequence, B) is calculated as follows: 100 multiplied by (the fraction X/Y), where X is the number of nucleotides or amino acids scored as identical matches by a sequence alignment program (e.g., BLAST) in that program’s alignment of A and B, and where Y is the total number of nucleotides or amino acids in B.
- a sequence alignment program e.g., BLAST
- the term “plant” includes a whole plant and any descendant, cell, tissue, or part of a plant.
- plant parts include any part(s) of a plant, including, for example and without limitation: seed (including mature seed and immature seed); a plant cutting; a plant cell; a plant cell culture; or a plant organ (e.g., pollen, mature or immature embryos, flowers, fruits, shoots, leaves, roots, stems, and explants).
- a plant tissue or plant organ is or includes a seed, protoplast, callus, or any other group of plant cells that is organized into a structural or functional unit.
- a plant cell or tissue culture is capable of regenerating a plant having the physiological and morphological characteristics of the plant from which the cell or tissue was obtained, and of regenerating a plant having substantially the same genotype as the plant.
- Regenerable cells in a plant cell or tissue culture can include embryos, protoplasts, meristematic cells, callus, pollen, leaves, anthers, roots, root tips, flowers, and/or stalks.
- some plant cells are not capable of being regenerated to produce plants and are referred to herein as “non-regenerable” plant cells.
- the term “transcriptome” refers to the sum total of all RNA molecules expressed in a cell.
- RNA molecules include mRNAs, tRNAs, ribosomal RNAs, miRNAs, viral RNAs (both genomic and sub-genomic), and long non-coding RNAs.
- RNA molecules include mRNAs, tRNAs, ribosomal RNAs, miRNAs, viral RNAs (both genomic and sub-genomic), and long non-coding RNAs.
- any of the preceding definitions is inconsistent with definitions provided in any patent or non-patent reference incorporated herein by reference, any patent or non-patent reference cited herein, or in any patent or non-patent reference found elsewhere, it is understood that the preceding definition will be used herein.
- nucleic acid sequences described herein are given, when read from left to right, in the 5’ to 3’ direction. Nucleic acid sequences may be provided as DNA or as RNA, as specified.
- nucleic acid sequences can encode the same polypeptide sequence, and such modified nucleic acid sequences (e.g., for the purposes of codon optimization for a given species) are within the scope of the present disclosure.
- modified nucleic acid sequences e.g., for the purposes of codon optimization for a given species
- recombinant polynucleotides e.g., recombinant DNA, recombinant RNA, recombinant ssRNAs, recombinant dsRNAs, recombinant vectors, etc.
- AV amalgavirus
- RdRP amalgavirus RNA-dependent RNA polymerase
- This disclosure is further related to methods of making and using such recombinant polynucleotides, for example, by employing such recombinant polynucleotides to express a heterologous cargo sequence in a plant and optionally thereby modifying expression of an endogenous target sequence and/or genotype or phenotype Agent Ref: P14357WO00 - 8 - of the plant.
- the amalgavirus is a commensal amalgavirus, that is, an amalgavirus that is endemic or native to a given eukaryote host (such as a host plant) without causing apparent negative effects on the host (i.e., is considered non-pathogenic), is often present at a persistent but low population (i.e., low viral titer), and is often vertically transmitted to succeeding generations of the host.
- this disclosure is related to a recombinant DNA molecule that includes a promoter that is functional in a cell and is operably linked to a DNA sequence encoding an RNA molecule.
- the RNA molecule includes, in 5’ to 3’ order: (a) a 5’ RNA replication element that is capable of being recognized by an amalgavirus RNA-dependent RNA polymerase (RdRP); (b) a cargo RNA sequence; and (c) a 3’ RNA replication element that is capable of being recognized by the amalgavirus RdRP.
- Figure 1 shows an embodiment of a generalized structure of a DNA polynucleotide encoding an amalgavirus satellite, where in certain embodiments the 5’ RNA replication element corresponds to the 5’ untranslated region (UTR) of an amalgavirus and where the 3’ RNA replication element corresponds to the 3’ untranslated region (UTR) of an amalgavirus.
- the 5’ RNA replication element and/or the 3’ RNA replication element include nucleotides that extend into the predicted coding sequence or open reading frame of the amalgavirus.
- Recombinant DNA molecules provided herein can include a promoter that is functional in a cell (e.g., a bacterial cell, a plant cell, a fungal cell, or an animal cell) and is operably linked to a DNA sequence encoding an RNA molecule (e.g.
- RNA replication element a 5’ RNA replication element, a cargo RNA sequence; and a 3’ RNA replication element; a ribozyme, an intron, or a RNA encoding a protein (e.g., a capsid, movement, RdRP, or an RdRP protein that recognizes an HRV 5’ or 3’ replication region and/or a subgenomic promoter).
- a protein e.g., a capsid, movement, RdRP, or an RdRP protein that recognizes an HRV 5’ or 3’ replication region and/or a subgenomic promoter.
- a promoter functional in a plant cell provides for systemic gene expression, or alternatively for cell-, tissue-, or organ-specific gene expression, or expression that is inducible by external signals or agents (for example, light-, pathogen-, wound-, stress-, or hormone-inducible elements, or chemical inducers) or elements that are capable of cell-cycle regulated gene transcription; such elements may be located in the 5’ or 3’ regions of the native gene or engineered into a polynucleotide.
- Promoters include those from viruses, bacteria, fungi, animals, and plants.
- RNA polymerase e.g., RNA pol I, pol II, or pol III
- RNA polymerase e.g., RNA pol I, pol II, or pol III
- Embodiments of promoters include those from cauliflower mosaic virus (e.g., p35S), bacteriophage (e.g., pT7), and plants (e.g., pATUBQ10).
- the promoter is operably linked to nucleotide sequences encoding multiple guide RNAs, wherein the sequences encoding guide RNAs are separated by a cleavage site such as a nucleotide sequence encoding a microRNA recognition/cleavage site or a self-cleaving ribozyme (see, e.g., Ferré-D’Amaré and Scott (2010) Cold Spring Harbor Perspectives Biol., 2:a003574).
- the promoter is a pol II promoter operably linked to a nucleotide sequence encoding the RNA.
- the promoter operably linked to one or more polynucleotides encoding elements of a genome-editing system is a constitutive promoter that Agent Ref: P14357WO00 - 9 - drives DNA expression in plant cells.
- the promoter drives DNA expression in the nucleus or in an organelle such as a chloroplast or mitochondrion.
- constitutive promoters active in plant cells include a CaMV 35S promoter as disclosed in U.S. Pat. Nos.5,858,742 and 5,322,938, a rice actin promoter as disclosed in U.S. Pat. No.5,641,876, a maize chloroplast aldolase promoter as disclosed in U.S. Pat.
- the promoter operably linked to one or more polynucleotides encoding elements of a genome-editing system is a promoter from figwort mosaic virus (FMV), a RUBISCO promoter, or a pyruvate phosphate dikinase (PDK) promoter, which is active in the chloroplasts of mesophyll cells.
- FMV figwort mosaic virus
- RUBISCO RUBISCO promoter
- PDK pyruvate phosphate dikinase
- the promoter is heterologous to the cell it is functional in and/or to the other elements to which the promoter is operably linked.
- Embodiments of recombinant polynucleotides provided herein comprise or encode RNA molecules containing 5’ and 3’ RNA replication elements recognized by an amalgavirus RNA-dependent RNA polymerase (RdRP).
- RdRP amalgavirus RNA-dependent RNA polymerase
- recognition by an amalgavirus RdRP is identified in an in vitro RdRP assay (e.g., an assay adapted from Horiuchi et al. Plant Cell Physiol.42(2):197-203, 2001).
- recognition by an amalgavirus RdRP is identified by an in vivo RdRP assay wherein an RNA comprising 5’ and 3’ RNA replication elements is introduced into a cell comprising the RdRP, and replication of the RNA is assayed (e.g., by an RT-PCR assay or an assay for a reporter gene encoded by a cargo RNA located in the RNA comprising 5’ and 3’ RNA replication elements).
- cells comprising the RdRP are engineered by introducing a gene or RNA molecule encoding the RdRP into the cell.
- the cell comprising the RdRP is a cell which contains an amalgavirus which expresses the RdRP; in such embodiments the amalgavirus can be one that is native to or is known to naturally occur in the cell, or it can be a non-native amalgavirus.
- a recombinant virus of any suitable viral family is engineered to express the amalgavirus RdRP.
- the recombinant polynucleotides comprise a 5’ RNA replication element and a 3’ RNA replication element obtained from the same amalgavirus genome or from amalgavirus genomes having at least 85%, 90%, 95%, 98%, or 99% sequence identity to one another and which are optionally related.
- the amalgavirus genomes having at least 85%, 90%, 95%, 98%, or 99% sequence identity to one another are taxonomically related, e.g., genomes classified as belonging to the same genus, family, and/or order.
- the recombinant polynucleotides comprise a 5’ RNA replication element, a 3’ RNA replication element, and an RdRP are obtained from the same amalgavirus genome or from amalgavirus genomes having at least 85%, 90%, 95%, 98%, or 99% sequence identity to one another and which are optionally taxonomically related, e.g., genomes classified as belonging to the same genus, family, and/or order.
- Non-limiting examples of a 5’ RNA replication element and a 3’ RNA replication element from the same amalgavirus capsid protein genome Agent Ref: P14357WO00 - 10 - include those set forth in each row of Table 18.
- the RdRPs set forth in Table 18 also recognize the corresponding 5’ RNA replication element and a 3’ RNA replication element from the amalgavirus capsid protein genome corresponding to the same amalgavirus (i.e., an amalgavirus having the 5’ RRE and 3’ RRE of Table 18 obtained from the amalgavirus genome encoding the RdRP).
- the recombinant polynucleotides comprise a 5’ RNA replication element, a 3’ RNA replication element, and/or an RdRP coding region are obtained from two amalgavirus genomes wherein the members of each pair of the 5’ RNA replication elements, 3’ RNA replication elements, and RdRP coding regions of the two amalgavirus genomes have at least 85%, 90%, 95%, 98%, or 99% sequence identity to one another.
- the recombinant polynucleotides comprise a 5’ RNA replication element and a 3’ RNA replication element obtained from distinct amalgavirus genomes.
- the recombinant polynucleotides e.g., recombinant DNAs or recombinant RNAs
- the distinct amalgavirus genomes will have less than 85%, 80%, 75%, or 70% sequence identity to one another.
- the distinct amalgavirus genomes will have 50%, 60%, or 65% to any one of 70%, 75%, 80%, or 84% sequence identity to one another.
- the combination of 5’ RNA replication elements, 3’ RNA replication elements, and RdRP set forth in a single row of Table 18, or variants thereof having at least 85%, 90%, 95%, 98%, or 99% sequence identity to the 5’ RNA replication element, 3’ RNA replication elements, and RdRP, or variants thereof wherein the secondary structures of the RNA replication elements are conserved, are used together in an expression system, plant cell, plant propagule, plant, or method provided herein.
- the 5’ RNA replication elements and 3’ RNA replication elements in a given row of Table 18 or variants thereof are operably linked to a cargo RNA and replicated by the corresponding RdRP or variant thereof in the row.
- the combination of 5’ RNA replication elements, 3’ RNA replication elements, and RdRP set forth in any one row of Table 18 or aforementioned or otherwise disclosed variants thereof are used in a dicot plant cell-based expression system, dicot plant cell, dicot plant propagule, dicot plant, or related dicot plant- based method provided herein.
- the aforementioned dicot is a member of the genus Arachis, Brassica, Capsicum, Cicer, Cucumis, Cucurbita, Gossypium, Lens, Nicotiana, Solanum, Phaseolus, Pisum, Vicia, Vigna, or Glycine.
- the 5’ RNA replication elements, 3’ RNA replication elements, and RdRP used in the dicot are obtained and/or derived from viral sequences which were originally isolated from a dicot host (e.g., as set forth in Table 17).
- the combination of 5’ RNA replication elements, 3’ RNA replication elements, and RdRP set forth in any one row of Table 18 or aforementioned or otherwise disclosed variants thereof are used in a monocot plant cell-based expression system, monocot plant cell, monocot plant propagule, monocot plant, or monocot dicot plant-based method provided herein.
- the aforementioned monocot Agent Ref: P14357WO00 - 11 - is a member of the genus Allium, Avena, Festuca, Hordeum, Lilium, Lolium, Miscanthus, Oryza, Saccharum, Secale, Setaria, Sorghum, x Triticosecale, Triticum, or Zea.
- the 5’ RNA replication elements, 3’ RNA replication elements, and RdRP used in the monocot are obtained and/or derived from viral sequences which were originally isolated from a monocot host (e.g., as set forth in Table 17).
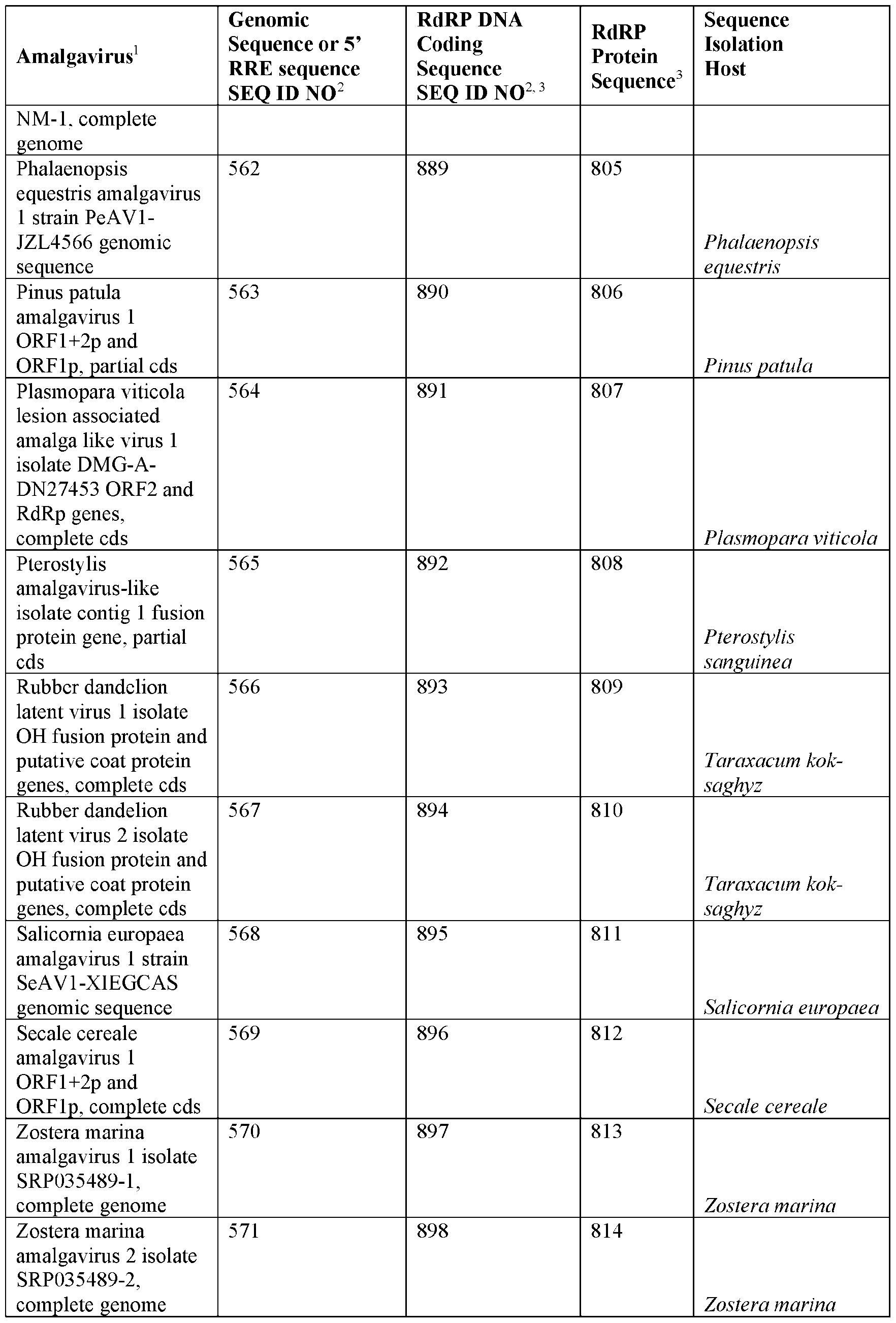
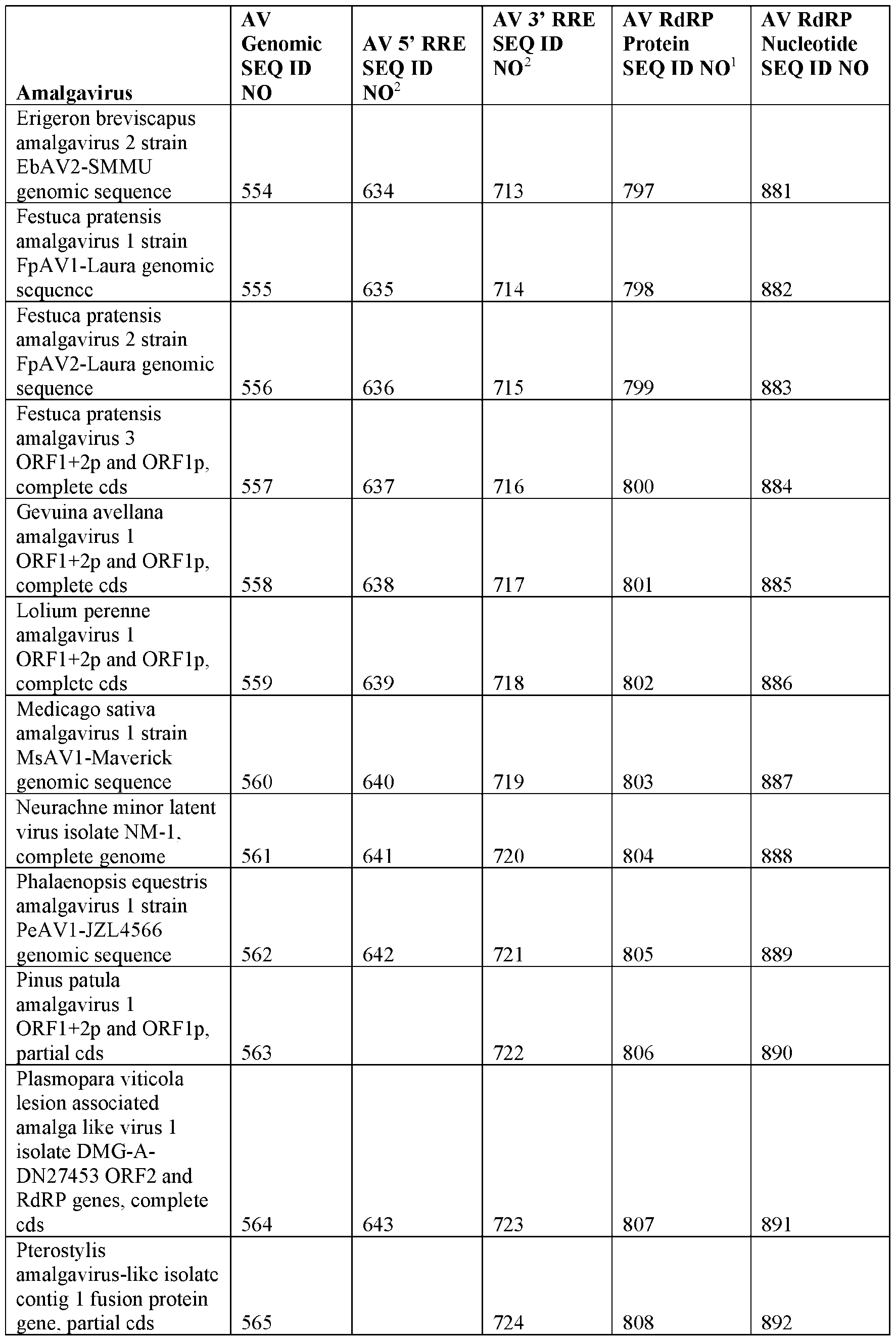
- Examples of DNA molecules which encode RNA molecules comprising or containing 5’ and 3’ RNA replication elements recognized by an amalgavirus RdRP are set forth in Table 1.
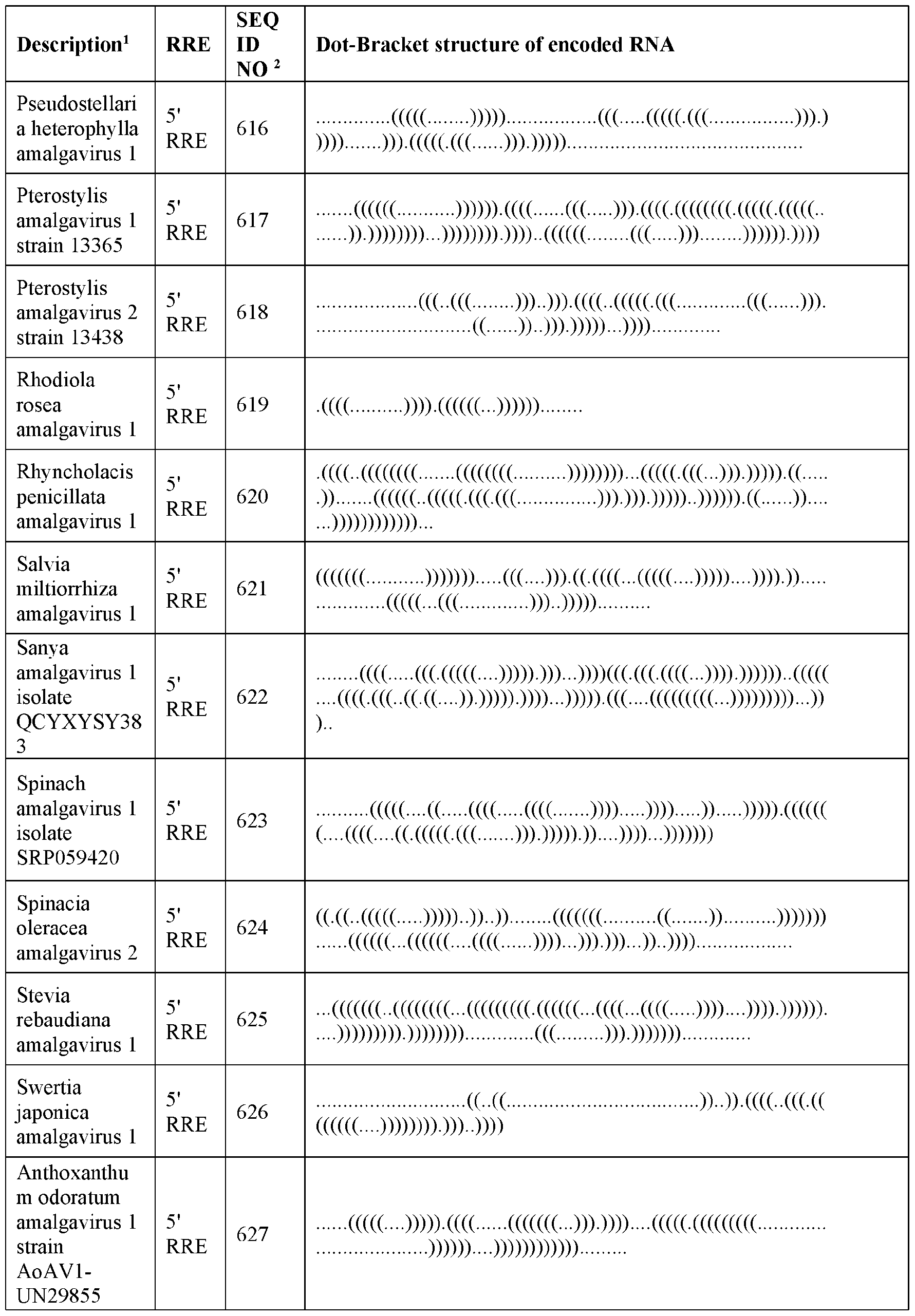
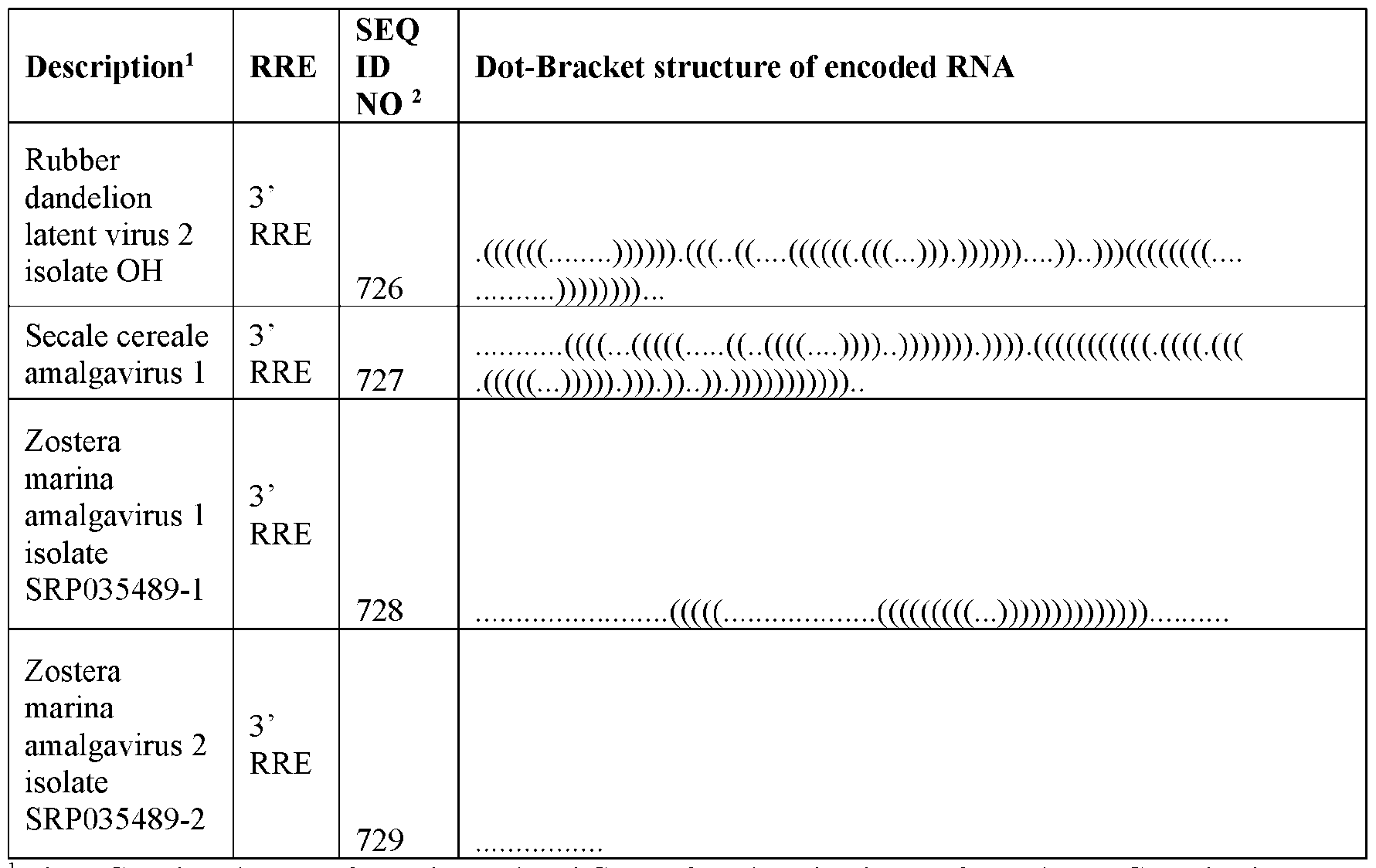
- DNA molecules which encode RNAs comprising or containing 5’ RNA replication elements recognized by an amalgavirus RdRP include SEQ ID NOs: 467, 469, 471 and 572 to 649 (AV 5’ RRE) and SEQ ID NOs: 468, 470, 472, and 650 to 729 (AV 3’ RRE).
- RNA replication elements (RRE) are shown in Table 1 by way of dot bracket notation.
- the dot bracket notation provided in Table 1 was generated using RNA Fold software for predicting RNA secondary structure based on minimum free energy predictions of base pair probabilities.
- a dot ‘.’ signifies an unpaired base and a bracket ‘(‘ or ‘)’ represents a paired base.
- Dot bracket notation is further described in Mattei et al., Nucleic Acids Research, 42(10): 6146-6157, 2014; Ramlan and Zauner In: International Workshop on Computing With Biomolecules, E.
- Such structural features can range in size from 20, 30, or 40 to about 500 nucleotides (nt). These structural features are useful for designing engineered polynucleotide sequences that function as amalgavirus RNA replication elements and/or for constructing variants of the sequences set forth in SEQ ID NO: 467 to 472 that function as 5’ and 3’ RNA replication elements (RRE).
- RRE RNA replication elements
- one of more residues in the RNA secondary structure set forth in Table 1 or in equivalent RNAs are substituted with distinct nucleotides which maintain the RNA secondary structure (e.g., presence or absence of base pairing).
- the RNA secondary structure set forth in Table 1 or in equivalent RNAs the RNA secondary structure is maintained by making substitutions in the nucleotide sequence that result in no changes in the position of base-paired nucleotides or non-base-paired nucleotides.
- RNA secondary structure set forth in Table 1 or in equivalent RNAs is maintained by substituting nucleotides in the secondary structure which are not base paired with nucleotides which will not base pair, and/or by substituting nucleotides in the secondary structure which are base paired with nucleotides which will base pair.
- maintaining the RNA secondary structure need not be absolute (e.g., the structure is partially maintained).
- a dsRNA structure is partially maintained when one, two, three or more nucleotides, particularly at the 5’ end and/or 3’ end of a hairpin-forming structure are substituted with nucleotides which do not base pair and thus reduce the total length of dsRNA in the structure.
- an unpaired RNA structure is partially maintained when one, two, three or more Agent Ref: P14357WO00 - 12 - nucleotides, particularly at the 5’ end and/or 3’ end of a loop structure are substituted with nucleotides which base pair and thus reduce the total length of ssRNA in the loop structure.
- Embodiments of amalgavirus satellite RNAs include those where the 5’ RNA replication element includes one or more of these 5’ structural features and/or wherein the 3’ RNA replication element includes one or more of these 3’ structural features.
- the 5’ RNA replication elements comprise an RNA encoded by a DNA having at least 70%, 75%, 80%, 85%, 90%, 95%, 98%, 99%, or 100% sequence identity to SEQ ID NO: 467, 469, 471, or 572 to 649, optionally wherein the encoded RNA maintains or partially maintains a corresponding structural feature set forth in Table 1.
- the 3’ RNA replication elements comprise an RNA encoded by a DNA having at least 70%, 75%, 80%, 85%, 90%, 95%, 98%, 99%, or 100% sequence identity to SEQ ID NO: 468, 470, 472, or 650 to 729, wherein the encoded RNA optionally maintains or partially maintains a corresponding structural feature set forth in Table 1.
- Recombinant polynucleotides e.g., recombinant DNA, recombinant RNA, recombinant ssRNAs, recombinant dsRNAs, recombinant vectors, etc.
- Recombinant polynucleotides e.g., recombinant DNA, recombinant RNA, recombinant ssRNAs, recombinant dsRNAs, recombinant vectors, etc.
- Embodiments of additional RNA elements include RNAs encoding an amalgavirus RdRP.
- DNA sequences encoding an amalgavirus RdRP include the corresponding sequences of AV RdRP genomes set forth in Table 1 under descriptors which refer to the National Center for Biotechnology Information database accession number for entries in the world wide web internet database “ncbi[dot]nlm[dot]nih.gov/nuccore”.
- DNA sequences encoding amalgavirus RdRP also include the sequence of the DNA encoding the AV RdRP disclosed in SEQ ID NO: 473, 474, or 475 as well as DNA sequences having at least 85%, 90%, 95%, 98%, or 99% identity thereto.
- DNA sequences encoding amalgavirus RdRP and amalgavirus RdRP protein sequences also include the sequences set forth in Table 17 as well as the DNA and protein sequences having at least 85%, 90%, 95%, 98%, or 99% identity thereto.
- Embodiments of additional RNA elements include RNAs encoding a viral movement protein (MP).
- the cargo RNA comprises an RNA encoding a viral MP.
- the viral movement protein is believed to bind to the RNA and to assist its movement (and thus the movement of the cargo RNA) throughout the plant, e.g., via the plasmodesmata.
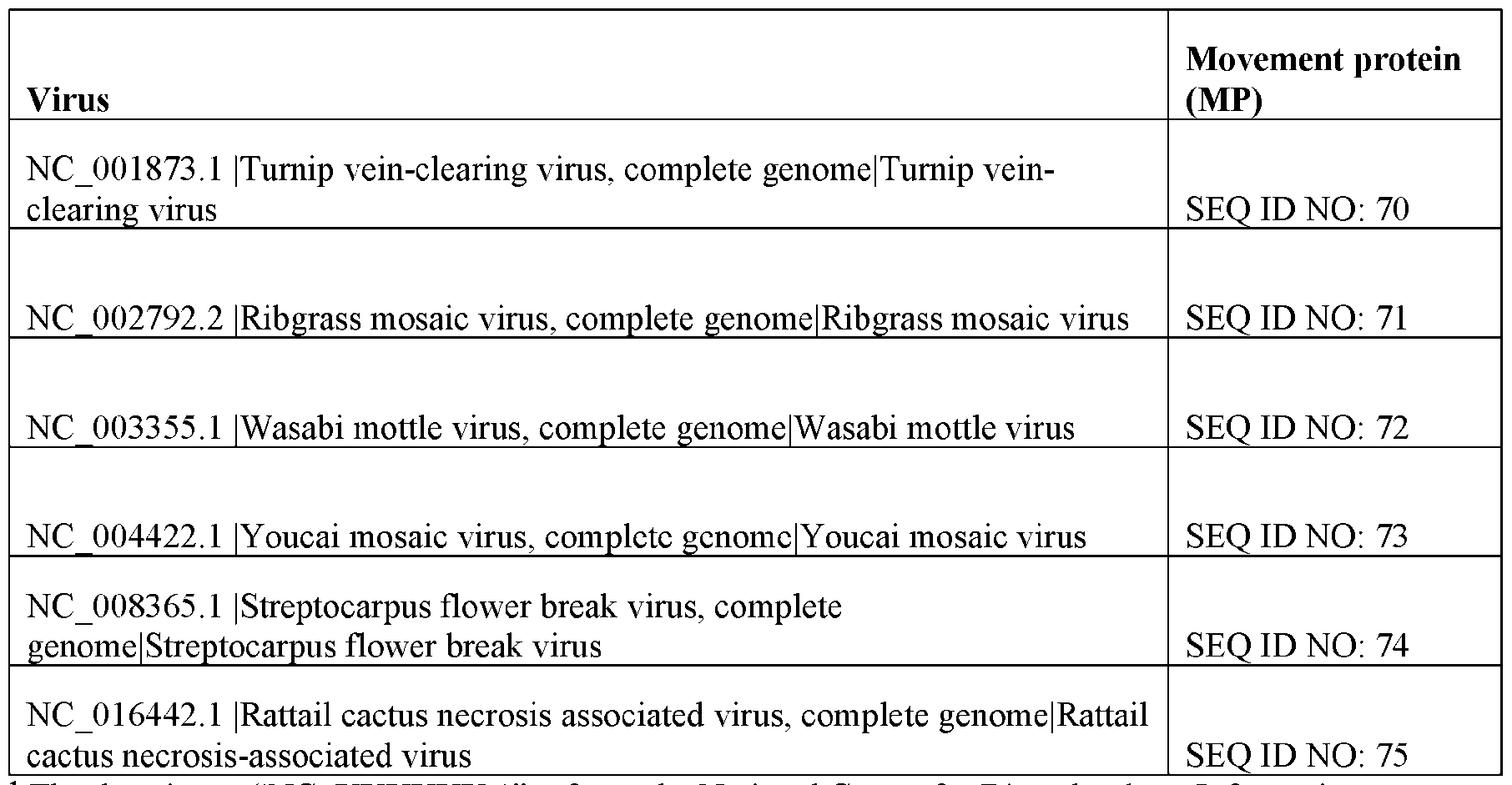
- Viral MPs include movement proteins identified from tobacco mosaic virus (TMV), cowpea mosaic virus, potato leafroll virus, tomato spotted wilt virus, and tomato mosaic virus. MPs from a variety of viruses are described in Table 3.
- TLS tRNA-like sequences
- TLS can trigger mobility of otherwise nonmobile RNAs, assisting to increase systemic delivery of the RNA molecule.
- TLS includes tRNAs and tRNA-like sequences identified from other genetic elements, e.g., mRNAs.
- An isoleucine tRNA encoded by SEQ ID NO: 466 is an example of a useful tRNA-like Agent Ref: P14357WO00 - 13 - sequence.
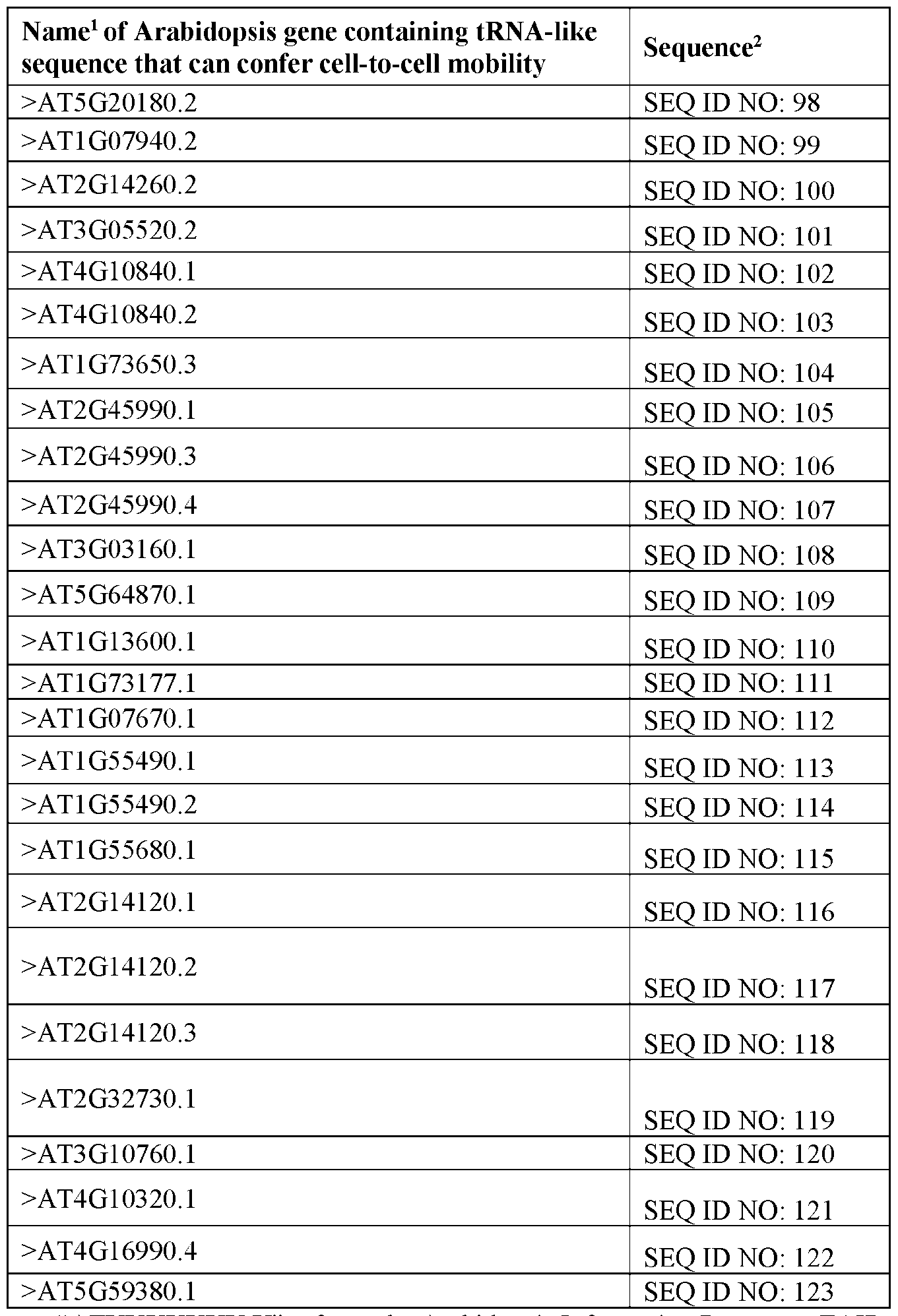
- Other mobile RNAs including TLS identified in Arabidopsis which are useful for building polynucleotides are described in Table 4.
- mobile mRNA sequences were downloaded from the PLAMOM database for Arabidopsis.
- the tRNA “seed alignment” from the RFAM database was downloaded in stockholm format (multiple sequence alignment + secondary structure).
- a covariance model was created with INFERNAL for the tRNA stockholm alignment.
- PLAMOM mRNA sequences were scanned for significant similarity to tRNAs based on primary and secondary structure features. mRNA sequences with significant hits (E-val ⁇ 1) were then saved to a fasta file.
- a tRNA-like sequence includes a tRNA-like sequence from an Arabidopsis Flowering Time T (FT) mRNA.
- the RNA molecule includes at least one RNA encoding a viral MP, a tRNA-like sequence from an Arabidopsis FT mRNA, and an encapsidation recognition element (ERE) comprising TMV-OAS.
- the RNA molecule includes a tRNA-like sequence encoded by a DNA sequence selected from the group consisting of SEQ ID NOs: 76-123, and 466. In some embodiments, the RNA molecule includes a modified tRNA-like sequence that has at least 90%, 95%, 98%, or 99% sequence identity to a scaffold tRNA-like sequence encoded by a DNA sequence selected from the group consisting of SEQ ID NOs: 76-123, and 466 and that maintains the secondary structure of the scaffold tRNA-like sequence.
- Embodiments of additional RNA elements include RNAs encoding a viral capsid protein (CP).
- capsid proteins are also sometimes referred to as coat proteins, with both capsid and coat proteins being referred to as “CP.”
- the capsid protein is heterologous to the amalgavirus.
- the cargo RNA comprises an RNA encoding a viral CP.
- CP can be provided, e.g., by co-expression of a recombinant construct encoding the CP or by native expression by a virus endogenous to or introduced into a plant cell. Encapsidation of an RNA molecule by the CP is achieved provided it contains an encapsidation recognition element (ERE), e.g., an origin of assembly sequence (OAS).
- ERP encapsidation recognition element
- Table 2 describes several OAS and CP sequences from a variety of viruses useful in engineering constructs which provide for RNA encapsidation.
- the OAS is positioned near the 3’ end of a construct, e.g., within the 3’ region of a cargo RNA or 3’ to a cargo RNA.
- the OAS is found 5’ to the 3’ RNA replication elements (e.g., the 3’ RNA replication elements set forth in Table 1).
- a TMV-OAS positioned at the 3’ end of the RNA molecule is recognized by the TMV capsid protein, leading to assembly of a TMV virion around the RNA.
- Embodiments wherein the recombinant RNAs are complexed with RNA binding proteins (RBPs) are also provided herein.
- RBPs include RNA recognition motifs (RRMs) such as: (i) Lys/Arg-Gly-Phe/Tyr-Gly/Ala-Phe/Tyr-Val/Ile/Leu-X-Phe/Tyr, where X can be any amino acid (SEQ ID NO: 464); (ii) Ile/Val/Leu-Phe/Tyr-Ile/Val/Leu-X-Asn-Leu, where X can be any amino acid (SEQ ID NO: 465).
- RRMs RNA recognition motifs
- RNA elements include at least one ribozyme.
- Ribozymes include self-cleaving ribozyme, a ligand-responsive ribozyme (aptazyme), a trans-cleaving ribozyme designed to cleave a target sequence (e.g., a trans-cleaving hammerhead ribozyme designed to cleave the pepper phytoene desaturase (PDS) sequence (the RNA encoded by SEQ ID NO: 421), a hepatitis delta virus (HDV) ribozyme (the RNA encoded by SEQ ID NO: 423), or a hammerhead ribozyme (the RNA encoded by SEQ ID NO: 420).
- PDS pepper phytoene desaturase
- HDV hepatitis delta virus
- multiple ribozymes are included in a polynucleotide.
- Useful ribozymes include Twister, Hammerhead, Hairpin, and other ribozymes.
- Non- limiting examples of useful ribozymes include those provided in Table 14.
- such a ribozyme e.g., a self-cleaving ribozyme
- such a ribozyme e.g., a self-cleaving ribozyme
- a ribozyme is located 5’ to the HRV 5’ RNA replication region and/or 3’ to the HRV 3’ RNA replication region in a recombinant RNA comprising an imbedded heterologous RNA virus (HRV) amplicon.
- HRV heterologous RNA virus
- intronic sequences are placed in a 5’UTR downstream of a promoter (e.g., a promoter active in plant cells) used to drive expression of a recombinant RNA.
- a promoter e.g., a promoter active in plant cells
- intronic sequences are placed 5’ to a 5’ RNA replication element, in a cargo RNA, or 3’ to a 3’ RNA replication element.
- Embodiments of recombinant polynucleotides and additional RNA elements include subgenomic promoters recognized by an RNA-dependent RNA polymerase (RdRP) and/or RNA molecules encoding an RNA-dependent RNA polymerase (RdRP).
- RdRP RNA-dependent RNA polymerase
- RdRP RNA-dependent RNA polymerase
- Examples of such subgenomic promoters and RdRP include a Brome Mosaic Virus subgenomic promoter and RdRP (Siegal et al.1998, doi: 10.1073/pnas.95.20.11613), barley yellow dwarf virus (BYDV) sgRNA1, sgRNA2, and sgRNA3 subgenomic promoters and RdRP (Koev and Miller; J Virol.2000 Jul;74(13):5988-96.
- Brome Mosaic Virus subgenomic promoter and RdRP (Siegal et al.1998, doi: 10.1073/pnas.95.20.11613), barley yellow dwarf virus (BYDV) sgRNA1, sgRNA2, and sgRNA3 subgenomic promoters and RdRP (Koev and Miller; J Virol.2000 Jul;74(13):5988-96.
- such subgenomic promoters are placed either 5’ and/or 3’ to an RNA molecule comprising a 5’ RNA replication element, a cargo RNA, and a 3’ RNA replication element to permit production of either or both + and – strands of the RNA molecule when the RdRP is provided.
- such subgenomic promoters are operably linked to a cargo RNA molecule and/or to any additional RNA element to permit production of the corresponding cargo and/or additional RNA when the RdRP is provided.
- the subgenomic promoters are operably linked to a cargo RNA comprising an HRV-inhibitory RNA or to a cargo RNA that encodes a protein which inhibits infection, movement, transmission, and/or replication of the HRV.
- the subgenomic promoters are operably linked to a cargo RNA comprising an RNA having at least 20 Agent Ref: P14357WO00 - 15 - contiguous nucleotides having an identical or complementary sequence to a segment of equivalent length of the genomic RNA of the HRV.
- the subgenomic promoters are operably linked to a cargo RNA comprising an RNA having at least 20 contiguous nucleotides having an identical or complementary sequence to a segment of equivalent length of the genomic RNA of the HRV which does not encode the hrvRdRP.
- Embodiments of other optional elements in the recombinant polynucleotides provided herein include: a) a discrete expression cassette including a second promoter operably linked to a DNA sequence to be transcribed, and optionally a terminator element (see, e.g., a NOS or CaMV35S terminator); (b) an expression-enhancing element (e.g., a DNA encoding an expression-enhancing intronic sequence); (c) a DNA or RNA sequence encoding a marker (e.g., a selectable marker such as DNA or RNA encoding an antibiotic resistance or herbicide resistance sequence; DNA encoding a scorable marker or detectable label (e.g., a beta-glucuronidase, fluorescent protein, luciferase, etc.); (d) a DNA aptamer; (e) a DNA or RNA sequence encoding an RNA aptamer; (f) T-DNA left and right border DNA sequences; (g) a
- recombinant polynucleotides comprising a cargo RNA molecule or comprising DNA encoding a cargo RNA molecule.
- the recombinant polynucleotide includes a single cargo RNA molecule.
- the recombinant polynucleotide includes at least two cargo RNA molecules, e.g., 2, 3, 4, 5, 6, 7, 8, 9, 10, or more than 10 cargo RNA molecules; in embodiments, the at least two cargo RNA molecules are the same (e.g., multiple copies of a non-coding RNA sequence or multiple copies of a RNA sequence encoding a polypeptide) or are different (e.g., two or more different non-coding RNA sequences, or two or more different coding RNA sequences, or combinations of non-coding and coding cargo RNA sequences).
- a cargo RNA molecule is up to about 3.5 kilobases (kb) in length.
- Cargo RNA molecules can range in length from any one of about 20 nucleotides (nt), 100nt, 200nt, 300nt, 400nt, 500nt, 600nt, 700nt, 800nt, or 900nt to any one of about 1kb, 2kb, 3kb, or 3.2 kb in length.
- Other lengths of the cargo RNA molecule are less than or equal to 100 nucleotides (nt) can range in length from any one of about 20nt, 30nt, or 40nt to any one of about 50nt, 60nt, 70nt, 80nt, 90nt, or 100nt in length.
- Recombinant RNAs comprising a cargo RNA of up to about 3.5kb, 4kb, 5kb, 6kb, 7kb, 8kb, 9kb, 10kb, 11kb, 12kb, 13kb, or 14kb in length can in certain embodiments be encapsidated by a Agent Ref: P14357WO00 - 16 - heterologous viral capsid protein set forth in Table 2.
- recombinant RNAs comprising a cargo RNA of up to about 14kb and encapsidated by a heterologous viral capsid protein can comprise an OAS element set forth in Table 2 and be encapsidated by a corresponding capsid protein set forth in Table 2.
- the cargo RNA molecule is greater than 14kb, for example, 15kb, 16kb, 17kb, 18kb, 19kb, or even 20kb.
- the cargo RNA molecule includes: (a) at least one coding sequence, (b) at least one non-coding sequence, or (c) both at least one coding sequence and at least one non-coding sequence.
- Such cargo RNA molecules include combinations of coding/non- coding sequence; multiple non-coding/coding sequences; as well as aptamers, ribozymes, and other elements as is described herein.
- the cargo RNA molecule includes (a) a coding sequence to be expressed in a plant, and (b) at least one non-coding sequence that modifies expression or translation of the coding sequence, such as a recognition and cleavage sequence for an siRNA or miRNA that is endogenously expressed in the plant (see, e.g., US Patent Nos.8,334,430, 9,139,838, 9,976,152, 10,793,869, 10,876,126) and can bind to and cleave an RNA transcript containing the recognition and cleavage sequence; in such embodiments, it is possible to achieve spatially or temporally or developmentally specific expression of the coding sequence in the plant.
- a coding or non-coding cargo RNA can be optimized for expression in plants by methods which include using codons which occur more frequently in plant genes and/or by eliminating polyadenylation sites in the cargo RNA (e.g., as described in at least US Pat. Nos.5380831, 5689052, and 7741118, which are each incorporated herein by reference in their entireties).
- the cargo RNA molecule includes at least one coding sequence (e.g., a translatable sequence).
- the coding sequence is accordingly a protein or a polypeptide such as those described in this disclosure’s working examples.
- a cargo RNA comprises a selectable marker RNA encoding an antibiotic resistance or herbicide resistance polypeptide sequence or a scorable marker RNA encoding a scorable marker protein (e.g., a beta- glucuronidase, fluorescent protein, luciferase, etc.).
- selectable marker/selection agent combinations include glyphosate-resistant EPSPS enzymes and/or glyphosate oxidases/glyphosate, a bialaphos resistance (bar) or phosphinothricin acyl transferase (pat) enzyme/glufosinate, or a neomycin phosphotransferase (npt)/neomycin or kanamycin.
- scorable markers include ⁇ -glucuronidase (GUS), luciferase, and fluorescent proteins such as green fluorescent protein (GFP), yellow fluorescent protein (YFP), and cyan fluorescent protein (CFP).
- GUS ⁇ -glucuronidase
- GFP green fluorescent protein
- YFP yellow fluorescent protein
- CFP cyan fluorescent protein
- the cargo RNA sequence encodes at least one protein or polypeptide that provides a desirable trait in a plant in which the protein or polypeptide is expressed.
- polypeptides useful in agricultural applications include, for example, bacteriocins, lysins, antimicrobial peptides, nodule C-rich peptides, and bacteriocyte regulatory peptides.
- Such polypeptides can be used to alter the level, activity, or metabolism of target microorganisms for increasing the fitness of beneficial insects (such as honeybees and silkworms) or for decreasing the fitness of pest invertebrates (such as aphids, caterpillars, beetle larvae, and mites).
- beneficial insects such as honeybees and silkworms
- pest invertebrates such as aphids, caterpillars, beetle larvae, and mites.
- Embodiments of agriculturally useful polypeptides include peptide toxins, such as those Agent Ref: P14357WO00 - 17 - naturally produced by entomopathogenic bacteria (e.g., Bacillus thuringiensis, Photorhabdus luminescens, Serratia entomophila, or Xenorhabdus nematophila), as is known in the art.
- Embodiments of agriculturally useful polypeptides include polypeptides (including small peptides such as cyclodipeptides or diketopiperazines) for controlling agriculturally important pests or pathogens, e.g., antimicrobial polypeptides or antifungal polypeptides for controlling diseases in plants, or pesticidal polypeptides (e.g., insecticidal polypeptides and/or nematicidal polypeptides) for controlling invertebrate pests such as insects or nematodes.
- polypeptides including small peptides such as cyclodipeptides or diketopiperazines
- antimicrobial polypeptides or antifungal polypeptides for controlling diseases in plants
- pesticidal polypeptides e.g., insecticidal polypeptides and/or nematicidal polypeptides
- invertebrate pests such as insects or nematodes.
- Embodiments of antimicrobial polypeptides include cathelicidins, cecropins, beta-defensins, amphibian antimicrobial peptides (e.g., aurein-like peptides, esculentin, gaegurin, brevinin, rugosin, ranatuerin, ranacyclin, uperin, ocellatin, grahamin, nigrocin, dermoseptin, temporin, bombinin, maximin), enterocins, ponicerins, megourins, apidaecins, abaecins, attacin, bacteriocins and lantibiotics, dermcidin, formaecin, halocidins, lactocin, tachystatins, and some insecticidal toxins produced by spiders and scorpions.
- amphibian antimicrobial peptides e.g., aurein-like peptides, esculentin
- Embodiments of agriculturally useful polypeptides include antibodies, nanobodies, and fragments thereof, e.g., antibody or nanobody fragments that retain at least some (e.g., at least 10%) of the specific binding activity of the intact antibody or nanobody.
- Embodiments of agriculturally useful polypeptides include transcription factors, e.g., plant transcription factors; see., e.g., the “AtTFDB” database listing the transcription factor families identified in the model plant Arabidopsis thaliana), publicly available at agris- knowledgebase[dot]org/AtTFDB/.
- Embodiments of agriculturally useful polypeptides include nucleases, for example, exonucleases or endonucleases (e.g., Cas nucleases such as Cas9 or Cas12a).
- Embodiments of agriculturally useful polypeptides further include cell-penetrating peptides, enzymes (e.g., amylases, cellulases, peptidases, lipases, chitinases), peptide pheromones (for example, yeast or fungal mating pheromones, invertebrate reproductive and larval signaling pheromones, see, e.g., Altstein (2004) Peptides, 25:1373–1376).
- enzymes e.g., amylases, cellulases, peptidases, lipases, chitinases
- peptide pheromones for example, yeast or fungal mating pheromones, invertebrate reproductive and
- Embodiments of agriculturally useful polypeptides confer a beneficial agronomic trait, e.g., herbicide tolerance, insect control, modified yield, increased fungal or oomycte disease resistance, increased virus resistance, increased nematode resistance, increased bacterial disease resistance, plant growth and development, modified starch production, modified oils production, high oil production, modified fatty acid content, high protein production, fruit ripening, enhanced animal and human nutrition, production of biopolymers, environmental stress resistance, pharmaceutical peptides (e.g., hormones, enzymes, transcription factors, antigens, antibodies, or antibody fragments) and secretable peptides, improved processing traits, improved digestibility (e.g., reduced levels of toxins or reduced levels of compounds with “anti-nutritive” qualities such as lignins, lectins, and phytates), enzyme production, flavor, nitrogen fixation, hybrid seed production, fiber production, and biofuel production.
- beneficial agronomic trait e.g., herbicide tolerance, insect control, modified
- Non-limiting examples of agriculturally useful polypeptides include polypeptides that confer herbicide resistance (US Patent Nos.6,803,501; 6,448,476; 6,248,876; 6,225,114; 6,107,549; 5,866,775; 5,804,425; 5,633,435; and 5,463,175), increased yield (US Patent Nos.
- the cargo RNA encodes one or more small signaling peptides (SSPs), also called peptide hormones, which are an attractive option for use as cargoes in RNA commensal satellites due to their small size (5-75 amino acids) and potency.
- SSPs result from processing longer precursor polypeptides (derived from ORF regions).
- SSPs originate from a wider range of sources including intergenic/intronic regions, long non-coding RNAs, pri-miRNAs, and 5′ and 3′ UTRs of mRNAs.
- Non-limiting examples of SSPs include miPEP172c, miPEP171d, BomiPEP397a, AtmiPEP397a, BvmiPEP164b, and AtmiPEP164b peptides set forth in Table 13.
- the RNA molecule further includes an internal ribosome entry site (IRES) located 5’ and immediately adjacent to the at least one coding sequence.
- IRS internal ribosome entry site
- the cargo RNA molecule includes multiple coding sequences, and the RNA molecule further includes an IRES located 5’ and immediately adjacent to each of the coding sequences (e.g., open translational reading frames encoding a protein of interest.
- IRES sequences include those depicted in Table 5.
- the cargo RNA molecule includes a non-coding sequence such as those described in this disclosure’s working examples.
- non-coding sequences include a hairpin RNA (hpRNA); an RNA that forms multiple stem-loops; an RNA pseudoknot; an RNA sequence that forms at least partially double-stranded RNA; a small interfering RNA (siRNA) or siRNA precursor; a microRNA (miRNA) or miRNA precursor; a ribozyme; a ligand-responsive ribozyme (aptazyme); an RNA aptamer; or a long noncoding RNA (lncRNA).
- hpRNA hairpin RNA
- siRNA small interfering RNA
- miRNA microRNA
- aptazyme a ligand-responsive ribozyme
- RNA aptamer or a long noncoding RNA (lncRNA).
- the cargo RNA includes a selectable or scorable RNA marker, such as an RNA aptamer or a regulatory RNA, such as an siRNA or siRNA precursor (see, e.g., US Patent No.8,404,927, 8,455,716, 9,777,288, 10,378,012), a miRNA or a miRNA precursor (see, e.g., US Patent Nos.8,410,334, 8,395,023, 9,708,620), a trans-acting siRNA or trans-acting siRNA precursor (see, e.g., US Patent Nos.8,030,473, 8,476,422, 8,816,061, 9,018,002), a phased sRNA or phased sRNA precursor (see, e.g., US Patent No.8,404,928), an siRNA or miRNA decoy (see, e.g., US Patent Nos.8,946,511, 9,873,888), an siRNA or miRNA cleavage blocker (see, e
- RNA aptamers include those that exhibit fluorescence upon binding a molecule.
- the fluorescent RNA aptamer can be the Broccoli RNA aptamer.
- RNA aptamers that can be used include, but are not limited to, Spinach, Spinach2, Carrot, Radish, Corn, Red Broccoli, Orange Broccoli, and Broccoli Fluorets.
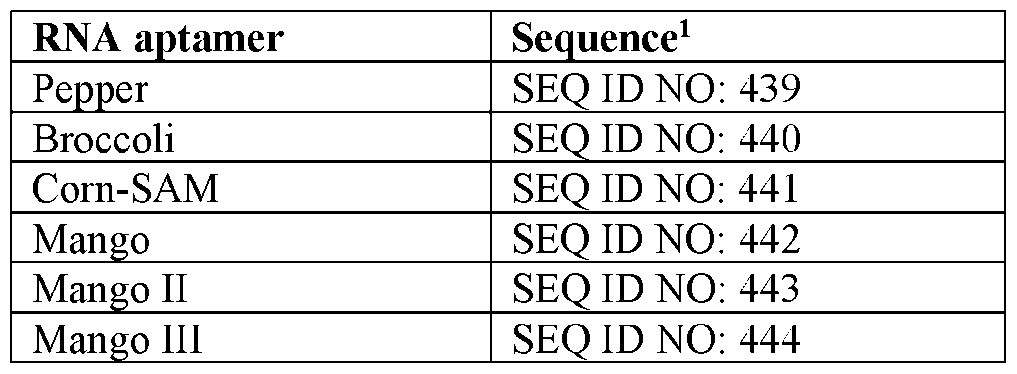
- Other useful RNA aptamers that can be used include those provided in Table 15.
- Suitable regulatory RNAs can be used to down-regulate (i.e., silence) the expression of a marker gene.
- PDS phytoene desaturase
- silencing of the gene yields a photobleached phenotype is widely used as a marker gene because silencing of the gene yields a photobleached phenotype.
- Antiviral cargo RNAs and in particular antiviral cargo RNAs directed against viral pathogens are provided herein.
- the antiviral cargo RNAs comprise a heterologous RNA Virus (HRV)-inhibitory RNA or encode an HRV-inhibitory protein, wherein the HRV-inhibitory RNA or protein inhibits infection, movement, transmission, and/or replication of the HRV.
- HRV heterologous RNA Virus
- Target viral pathogens include an Alphaflexivirus, Betaflexivirus, Bromovirus, Celavirus, a Closterovirus, a Comovirus, a Potexvirus, Potyvirus, Tobamovirus, Tombusvirus, Tospoviridae, Trivirinae, Tymovirus, Varicosavirus, or Secoviridae.
- the target viral pathogen is Cucumber Mosaic Virus, Brome mosaic virus, Citrus tristeza virus, Beet yellows virus, Cowpea mosaic virus, Potato virus X; Pepper mottle virus, Bean yellow mosaic virus, Barley stripe mosaic virus, Wheat stripe mosaic virus, Rice yellow mottle virus, Maize dwarf mosaic virus, zucchini yellow mosaic virus, watermelon mosaic virus, sugarcane mosaic virus, Tobacco mosaic virus, Tomato mosaic virus, Tomato brown rugose fruit virus, Turnip vein-clearing virus, Pepper mild mottle virus, Turnip crinkle virus, Tomato bushy stunt virus, Tomato spotted wilt virus, watermelon bud necrosis virus, Turnip yellow mosaic virus, Spinach latent Agent Ref: P14357WO00 - 20 - virus, Olive latent virus 2, Citrus yellow vein clearing virus, Potato latent virus, Apple stem grooving virus, Citrus leaf blotch virus, Apple latent spherical virus, Soybean latent spherical virus
- the targeted viral pathogen is a heterologous RNA virus disclosed in Table 7.
- antiviral inhibitory RNAs (RNAi sequences) used as cargo RNAs are obtained for a chosen target gene of a viral pathogen using siRNA/miRNA prediction tools (see, e.g., on the world wide web internet site “zhaolab[dot]org/pssRNAit/).
- siRNA/miRNA prediction tools see, e.g., on the world wide web internet site “zhaolab[dot]org/pssRNAit/).
- Other examples of non-coding RNA sequences having antiviral activity e.g., dsRNA molecules which produce miRNA or siRNA
- examples include those disclosed in US Patent No.8,455,716, which is incorporated herein by reference in its entirety.
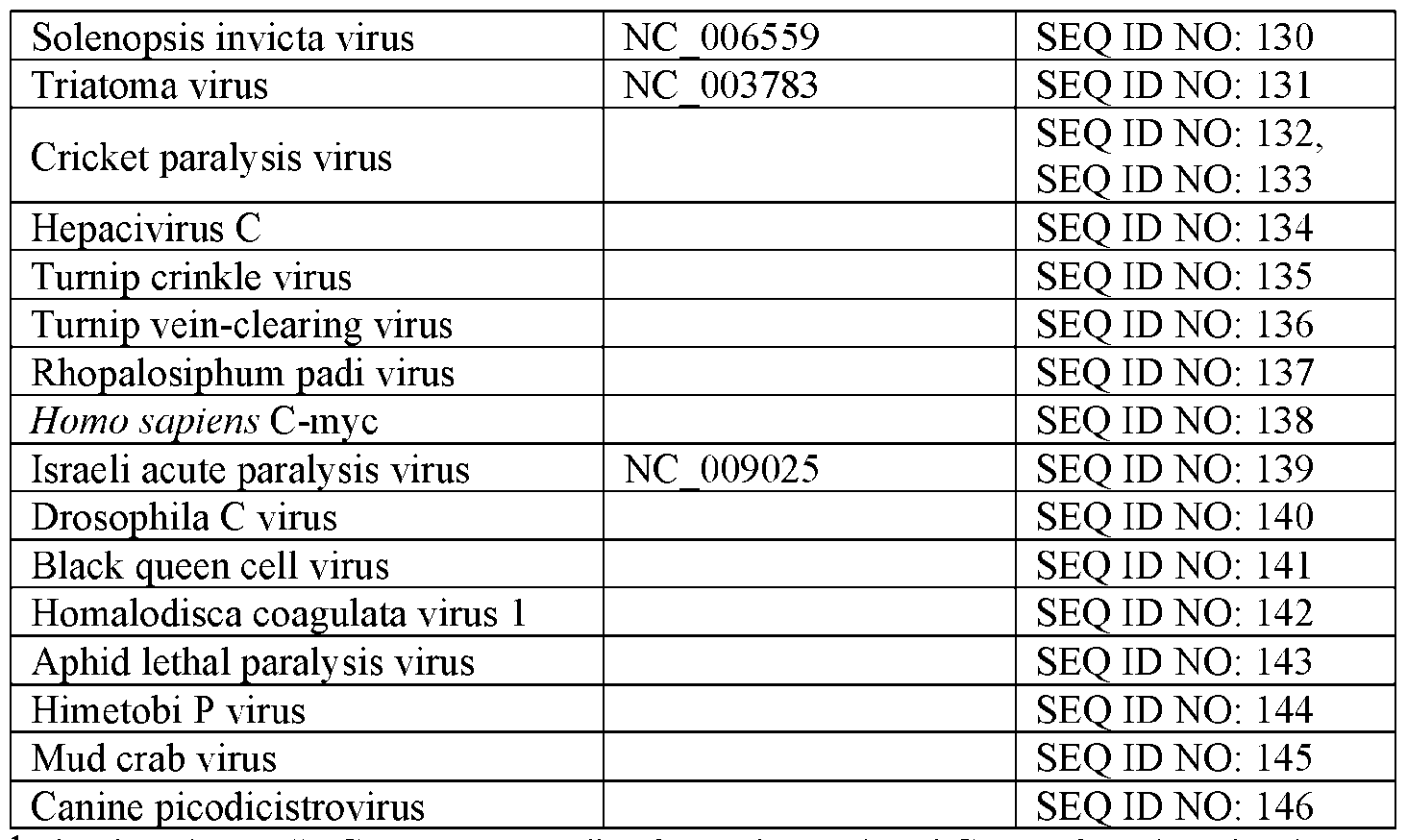
- Non-limiting examples of viral targets for antiviral cargo RNA molecules include the viral genes and genomes provided in Table 7 as well as other variants of those viral sequences.
- cargo RNAs encoding antiviral proteins are provided.
- Non-limiting examples of antiviral proteins include the N protein (Whitham, S. et al. Cell 78, 1101–1115 (1994)) and endogenous plant viral resistance proteins provided in Table 8.
- Antifungal cargo RNAs, and in particular antifungal cargo RNAs directed against plant fungal pathogens, are provided herein.
- Target fungal pathogens include Botrytis, Fusarium, Magnaporthe, Phytophthora, Rhizoctonia, Sclerotinia, and Verticillium sp.
- the antifungal cargo RNA comprises a non-coding RNA sequence having antifungal activity (e.g., dsRNA molecules which produce miRNA or siRNA) and in particular a dsRNA directed against a fungal pathogen target gene.
- such antifungal cargo RNAs comprising dsRNA-mediated control of fungal pathogens are modeled after those described in Qiao et al., 2021, doi: 10.1111/pbi.13589; Duanis-Assaf, et al., 2022, DOI: 10.1111/pbi.13708; Yang et al., 2022, doi: 10.3389/fmicb.2021.660976; Sundaresha et al., doi: 10.20944/preprints202102.0280.v1; and Gaffar et al., 2019, doi: 10.3389/fmicb.2019.01662.
- Non-limiting examples of antifungal cargo RNAi targets are provided in Table 10.
- antifungal inhibitory RNAs (RNAi sequences) used as cargo RNAs are obtained for a chosen target gene of a fungal pathogen (e.g., a fungal pathogen gene set forth in Table 10) using siRNA/miRNA prediction tools (see, e.g., on the world wide web internet site “zhaolab[dot]org/pssRNAit/).
- antifungal cargo RNAs encode antifungal proteins.
- Useful antifungal proteins include nodule-specific cysteine-rich antimicrobial peptides (Vellivelli et al., 2020, doi: 10.1073/pnas.2003526117), defensins (Asano et al., 2013, doi: 10.1371/journal.ppat.1003581), the conidial germination-inhibiting antifungal peptides disclosed in International Patent Application publication WO2023/004435, including their homodimers, heterodimers, and fusions with signal or cell- penetrating peptides (e.g., the sequences provided in Tables 4 and 5 of WO2023/004435), which is incorporated herein by reference in its entirety; the various antifungal antimicrobial peptides disclosed in De Cesare et al.
- Insecticidal cargo RNAs and in particular insecticidal or insect inhibitory cargo RNAs directed against insects are provided herein.
- Target insects include sucking insects (e.g., heteropteran and homopteran insects including aphids, whiteflies, and plant bugs), caterpillars (e.g., lepidopteran insects including fall army, black cutworm, corn earworm, soybean looper, and velvetbean caterpillar), beetles (e.g., coleopteran insects including Colorado Potato Beetle and corn rootworms), and flies (e.g., dipteran insects including Ceratitis capitata).
- Insecticidal or insect inhibitory cargo RNAs provided herein can be directed against insects at various stages of their development (e.g., embryonic, larval, pupal, or adult stages).
- the insecticidal or insect inhibitory cargo RNA comprises a non-coding RNA sequence having insecticidal or insect inhibitory activity (e.g., dsRNA molecules which produce miRNA or siRNA) and in particular a dsRNA directed against an insect target gene.
- insecticidal cargo RNAs comprising dsRNA-mediated control of insects comprise or are modeled after those described in US Patent Nos.11,091,770 and 11,186,837, which are each incorporated herein by reference in their entireties.
- Non-limiting examples of insecticidal or insect- inhibitory cargo RNAi targets are provided in Table 9.
- Non-limiting examples of insecticidal cargo RNAi targets include insect Actin, SNF7, Tyrosine hydroxylase, C002, Hunchback, V-ATPase subunit A, COPI coatomer beta prime subunit, ribosomal protein L19, and ubiquitin C genes.
- insecticidal or insect inhibitory RNAs (RNAi sequences) used as cargo are obtained for a chosen target gene of an insect (e.g., an insect gene set forth in Table 9 or US Patent Nos.11,091,770 and 11,186,837) using siRNA/miRNA prediction tools (see, e.g., on the world wide web internet site “zhaolab[dot]org/pssRNAit/).
- insecticidal cargo RNAs encode insecticidal proteins.
- Useful insecticidal proteins encoded by insecticidal cargo RNAs include native and modified Bacillus thuringiensis Cry, vegetative insecticidal proteins (VIP), and Cyt proteins (Palma et al.2014, doi: 10.3390/toxins6123296; US Patent No.11,267,849, incorporated herein by reference in its entirety) as well as insecticidal or insect-inhibitory proteins provided in Table 9.
- Cargo RNAs can also encode “resistance” or “R” genes which confer resistance to certain arthropods, bacteria such as Pseudomonas sp., Xanthomonas sp., and Erwinia sp., and fungal pathogens including Cochliobolus, Blumeria, Fusarium, Melampsora, and Magnaporthe sp.
- R genes encoded by cargo RNAs include those provided in Table 12.
- the cargo RNA molecule comprises a CRISPR guide RNA, e.g., a crRNA, gRNA, or sgRNA.
- CRISPR-associated endonucleases such as Cas9, Cas12, and Cas13 endonucleases are used as genome editing tools in different plants; see, e.g., Wolter et al. (2019) BMC Plant Biol., 19:176-183); Aman et al. (2016) Genome Biol., 19:1-10.
- CRISPR/Cas9 requires a two- component crRNA:tracrRNA “guide RNA” (“gRNA”) that contains a targeting sequence (the “CRISPR RNA” or “crRNA” sequence) and a Cas9 nuclease-recruiting sequence (tracrRNA).
- gRNA guide RNA
- sgRNA single guide RNA
- an engineered Agent Ref: P14357WO00 - 22 - (synthetic) single RNA molecule that mimics a naturally occurring crRNA-tracrRNA complex and contains both a tracrRNA (for binding the nuclease) and at least one crRNA (to guide the nuclease to the sequence targeted for editing); see, for example, Cong et al. (2013) Science, 339:819-823; Xing et al. (2014) BMC Plant Biol., 14:327-340.
- Chemically modified sgRNAs have been demonstrated to be effective in genome editing; see, for example, Hendel et al. (2015) Nature Biotechnol., 985-991.
- Commercial manufacturers of CRISPR nucleases and guide RNAs provide algorithms for designing guide RNA sequences; see, e.g., guide design tools provided by Integrated DNA Technologies at www[dot]idtdna[dot]com/pages/products/crispr-genome-editing/alt-r-crispr-cas9-system.
- RNAs are imbedded within a heterologous RNA virus (HRV) amplicon comprising; (i) an HRV 5’ replication region (HRV 5’ RR); (ii) the cargo RNA molecule; and (iii) the heterologous RNA virus (HRV) 3’ RNA replication region (HRV 3’RR), wherein (i), (ii), and (iii) are operably linked.
- HRV heterologous RNA virus
- an illustrative example of an amalgavirus satellite construct with such an HRV amplicon is shown in Figure 2.
- Amplification of such an HRV amplicon in plants comprising an amalgavirus satellite construct, the amalgavirus, and an HRV RdRP is illustrated in Figure 6.
- Examples of HRV 5’ replication regions (5’RR), 3’ replication regions (3’RR), and corresponding HRV RNA- dependent RNA Polymerases (RdRP) that recognize such replication regions are set forth in Table 7.
- an internal ribosome entry site IRES; e.g. an IRES in Table 5
- IRES internal ribosome entry site
- one or more self-cleaving or inducible ribozymes are operably linked to the 5’ end of the HRV 5’ RR and to the 3’ end of the HRV 3’ RR.
- the HRV amplicon further comprises a subgenomic promoter which is operably linked to the cargo RNA molecule.
- subgenomic promoters include a subgenomic promoter of the HRV and/or a Brome Mosaic Virus subgenomic promoter (Siegal et al.1998, doi: 10.1073/pnas.95.20.11613), barley yellow dwarf virus (BYDV) sgRNA1, sgRNA2, and sgRNA3 subgenomic promoters (Koev and Miller; J Virol.2000 Jul;74(13):5988-96, doi: 10.1128/jvi.74.13.5988-5996.2000), and an Alternanthera mosaic virus (AltMV-MU) sgp1, sgp2, or sgp3 subgenomic promoter (Putlyaev et al., Biochemistry (Mosc).;80(8):1039-46, DOI: 10.1134/S000629791508009X).
- HRV amplicons can be in the sense or antisense orientation with respect to the amalgavirus 5’ RNA replication element.
- HRV amplicon When the HRV amplicon is oriented in the sense orientation relative to the amalgavirus 5’ RNA replication element, the HRV 5’ RR and 3’ RR are present in the recombinant RNA molecule in the sense orientation, as in the corresponding sequences found in the plus (+) strand of the HRV genomic RNA.
- the HRV 5’ RR and 3’ RR are present in the recombinant RNA Agent Ref: P14357WO00 - 23 - molecule in antisense orientation, as in the corresponding sequences found in the negative (-) strand of the HRV genomic RNA.
- RNA Agent Ref P14357WO00 - 23 - molecule in antisense orientation, as in the corresponding sequences found in the negative (-) strand of the HRV genomic RNA.
- hrvRdRP RNA-dependent RNA polymerase that recognizes the HRV 5’ RR and 3’RR
- the HRV amplicon undergoes amplification (e.g., hrvRdRP-mediated replication).
- Such hrvRdRP can be provided by sources that include: (i) infection by the HRV; (ii) introduction by vector-mediated delivery of a polynucleotide encoding the hrvRdRP (e.g., Agrobacterium-mediated delivery or viral vector mediated delivery); or (iii) introduction of a nucleic acid encoding the hrvRdRP.
- sources that include: (i) infection by the HRV; (ii) introduction by vector-mediated delivery of a polynucleotide encoding the hrvRdRP (e.g., Agrobacterium-mediated delivery or viral vector mediated delivery); or (iii) introduction of a nucleic acid encoding the hrvRdRP.
- HRV1 RdRP and “HRV2 RdRP”
- Amplification e.g., an increase in copy number of the HRV amplicon
- the recombinant nucleotides provided herein comprise amalgavirus 5’ and 3’ RNA replication elements flanking a heterologous RNA virus (HRV) subgenomic promoter operably linked to the cargo RNA molecule; wherein the subgenomic promoter is recognized by a heterologous RNA virus RNA-dependent RNA polymerase (hrvRdRP).
- HRV heterologous RNA virus
- hrvRdRP heterologous RNA virus RNA-dependent RNA polymerase
- FIG. 8 Another illustrative example where subgenomic promoters drive expression of an HRV RdRP and a dsRNA cargo in plants comprising an amalgavirus satellite construct is shown in Figure 8.
- an internal ribosome entry site IRES; e.g. an IRES in Table 5
- IRES internal ribosome entry site
- Embodiments of subgenomic promoters include a subgenomic promoter of the HRV and/or a Brome Mosaic Virus subgenomic promoter (Siegal et al.1998, doi: 10.1073/pnas.95.20.11613), barley yellow dwarf virus (BYDV) sgRNA1, sgRNA2, and sgRNA3 subgenomic promoters (Koev and Miller; J Virol.2000 Jul;74(13):5988-96.
- a subgenomic promoter of the HRV and/or a Brome Mosaic Virus subgenomic promoter (Siegal et al.1998, doi: 10.1073/pnas.95.20.11613), barley yellow dwarf virus (BYDV) sgRNA1, sgRNA2, and sgRNA3 subgenomic promoters (Koev and Miller; J Virol.2000 Jul;74(13):5988-96.
- BYDV barley yellow dwarf
- RNAs from subgenomic promoters provides for additional copies of the cargo RNA and an enhancement of desirable phenotypes conferred by the cargo RNA (e.g., increased antiviral, antifungal, or insecticidal activity in comparison to control plants lacking the additional expressed cargo RNA or lacking the cargo RNA).
- the subgenomic promoters and operably linked cargo RNAs are oriented in the sense orientation relative to the amalgavirus 5’ RNA replication element, the subgenomic promoters and operably linked cargo RNAs are present in the recombinant RNA molecule as a sense strand where the subgenomic promoter is recognized by the hrvRdRP to produce the desired cargo RNA.
- the Agent Ref P14357WO00 - 24 - subgenomic promoters and operably linked cargo RNAs are oriented in the sense orientation relative to the amalgavirus 5’ RNA replication element in the recombinant RNA molecule (positive strand)
- the subgenomic promoter can be recognized by the hrvRdRP to produce the desired cargo RNA.
- the HRV amplicon is oriented in the antisense orientation relative to the amalgavirus 5’ RNA replication element in the recombinant RNA molecule, the subgenomic promoter cannot be recognized by the hrvRdRP to produce the desired cargo RNA.
- the negative strand of the recombinant RNA molecule produced by the amalgavirus RdRP would contain the subgenomic promoters and operably linked cargo RNA in a sense orientation where the subgenomic promoter can be recognized by the hrvRdRP to produce the desired cargo RNA.
- the HRV amplicons further comprise a HRV 5’ RR and 3’ RR which flank the cargo RNA and provide for hrvRdRP-mediated replication of an RNA comprising from 5’ to 3’ the HRV 5’ RR, cargo RNA, and HRV 3’ RR (e.g., as illustrated in the non-limiting example of Figure 5).
- an RNA encoding the cargo molecule can be produced (e.g., via hrvRdRP-mediated synthesis of the cargo RNA from the subgenomic promoter).
- Such hrvRdRP can be provided by sources that include: (i) infection by the HRV; (ii) introduction by vector-mediated delivery (e.g., Agrobacterium-mediated delivery or viral vector mediated delivery); (iii) introduction of a nucleic acid encoding the hrvRdRP; or (iv) inclusion of a cargo RNA in the recombinant nucleotides comprising amalgavirus 5’ and 3’ RNA replication elements.
- vector-mediated delivery e.g., Agrobacterium-mediated delivery or viral vector mediated delivery
- nucleic acid encoding the hrvRdRP e.g., Agrobacterium-mediated delivery or viral vector mediated delivery
- inclusion of a cargo RNA in the recombinant nucleotides comprising amalgavirus 5’ and 3’ RNA replication elements.
- the subgenomic promoter and operably linked cargo RNA are present in the recombinant RNA molecule as the antisense strand, and the cargo RNA encodes both an hrvRdRP and a second coding or non-coding RNA where both the hrvRdRP and a second coding or non-coding RNA are operably linked to a subgenomic promoter recognized by the hrvRdRP.
- an IRES is operably linked to the RNA encoding the hrvRdRP.
- RNA replication elements Production of the negative strand of the recombinant nucleotides comprising amalgavirus 5’ and 3’ RNA replication elements results in an RNA where the subgenomic promoters recognized by the hrvRdRP can drive expression of the HRV RdRP and a second coding or non-coding RNA.
- An illustrative example of an amalgavirus satellite construct with subgenomic promoters in antisense orientation relative to the 5’ RRE and driving expression of both an hrvRdRP that recognizes the subgenomic promoters and a second cargo RNA is shown in Figure 4.
- an RNA molecule including at least one HRV amplicon is amplified directly by the hrvRdRP (e.g., without initial or further amplification by the commensal viral RdRP).
- the HRV amplicon includes, in 5’ to 3’ order, (i) a heterologous RNA virus (HRV) 5’ replication region (HRV 5’RR); (ii) a cargo RNA molecule; and (iii) a heterologous RNA virus (HRV) 3’ RNA replication region (HRV 3’RR); wherein the HRV 5’ RR and HRV 3’ RR HRV are recognized by a heterologous RNA virus RNA-dependent RNA polymerase (hrvRdRP); and wherein the HRV 5’RR, cargo RNA molecule, and HRV 3’RR are operably linked.
- the HRV amplicon Agent Ref: P14357WO00 - 25 - includes, in 5’ to 3’ order, a subgenomic promoter that is operably linked to (v) a cargo RNA molecule, wherein the subgenomic promoter is recognized by the hrvRdRP.
- HRV amplicons can be provided in either an isolated form or in a composition.
- the RNA molecule including the HRV amplicon can be provided to a plant, for example, by transcription in the plant from a recombinant DNA molecule encoding the RNA molecule that is transiently expressed in the plant or that is stably integrated into the plant’s genome, or by delivery to the plant of an exogenous RNA molecule including the HRV amplicon, for example, by contacting a surface of the plant with the exogenous RNA molecule including the HRV amplicon, or by introducing the exogenous RNA molecule including the HRV amplicon into the plant’s vascular system (e.g., by injection, infusion, petiole uptake, root uptake).
- a plant for example, by transcription in the plant from a recombinant DNA molecule encoding the RNA molecule that is transiently expressed in the plant or that is stably integrated into the plant’s genome, or by delivery to the plant of an exogenous RNA molecule including the HRV amplicon, for example, by
- RNA molecules are delivered by injection into the vascular system of plants, such as perennial or woody plants.
- Examples of devices and methods for delivery via injection into plant vascular systems include those described in US Patent No.11844318; examples of delivery via injection into olive trees are described in PCT published application WO 2023/161802 and Grandi et al. (2023) Front. Plant Sci., 14:2023; doi.org/10.3389/fpls.2023.1180632; and examples of delivery via injection into citrus trees are described in PCT published application WO 2023/240208 and Al-Rimawi et al. (2019) Antibiotics, 8, 196; doi:10.3390/antibiotics8040196.
- the cargo RNA molecule includes at least one antiviral RNA (e.g., an antiviral inhibitory RNA or an RNA encoding an antiviral polypeptide) that provides the plant with resistance to at least one viral pathogen (which in some instances can be the heterologous RNA virus itself).
- antiviral RNA e.g., an antiviral inhibitory RNA or an RNA encoding an antiviral polypeptide
- Such embodiments are useful as antiviral treatments for plants, to prevent or decrease the severity of infection of a plant by a viral pathogen.
- plants including solanaceous plants that comprise amalgavirus satellite RNAs containing HRV amplicons disclosed herein can exhibit control of HRV that include Tobacco mosaic virus, Cucumber mosaic virus, Pepino mosaic virus, Potato virus X, Potato virus Y, Tobacco etch virus, Potato leafroll virus, Tomato apical stunt viroid, Tomato aspermy virus, Tomato black ring virus, Tomato blistering mosaic tymovirus, Tomato bushy stunt virus, Tomato chlorosis virus, Tomato chlorotic dwarf viroid, Tomato chlorotic spot orthotospovirus, Tomato fruit blotch virus, Tomato infectious chlorosis virus, Tomato mild mottle virus, Tomato mosaic virus, Tomato brown rugose fruit virus, Tomato necrotic streak virus, Tomato necrotic stunt virus, Tomato planta macho viroid, Tomato spotted wilt orthotospovirus, Tomato torrado virus, Tomato yellow ring orthotospovirus, Tomato zonate
- plants such as tomato plants that comprise Southern Tomato Virus (STV) amalgavirus RdRPs and satellite RNAs with 5’RRE and 3’RRE provided in Table 17 or variants thereof and containing an HRV amplicon can exhibit control of HRV that include Tobacco mosaic virus, Cucumber mosaic virus, Pepino mosaic virus, Potato virus X, Potato virus Y, Tobacco etch virus, Agent Ref: P14357WO00 - 26 - Potato leafroll virus, Tomato apical stunt viroid, Tomato aspermy virus, Tomato black ring virus, Tomato blistering mosaic tymovirus, Tomato bushy stunt virus, Tomato chlorosis virus, Tomato chlorotic dwarf viroid, Tomato chlorotic spot orthotospovirus, Tomato fruit blotch virus, Tomato infectious chlorosis virus, Tomato mild mottle virus, Tomato mosaic virus, Tomato brown rugose fruit virus, Tomato necrotic streak virus, Tomato necrotic stunt virus, Tomato planta mach
- the HRV amplicons or elements comprise sequences provided in Table 7 or variants thereof.
- RNA molecules that contain HRV amplification sequences (such as the HRV amplicons described herein which contain either (1) a pair of HRV 5’ and 3’ RNA replication regions, or (2) a subgenomic promoter that is recognized by the hrvRdRP) potentially also serve as a “sponge” or “decoy” that reduces the corresponding hrvRdRP’s efficiency in recognizing and amplifying the HRV viral genome itself, thus potentially decreasing a pathogenic HRV’s deleterious effects on an infected plant.
- RNA polynucleotides comprising at least one cleavable sequence are provided.
- the at least one cleavable sequence is located: (i) 5’ to the 5’ RNA replication element or 3’ to the 3’ RNA replication element; and/or (ii) between the 3’ end of the 5’ RNA replication element and the HRV amplicon and/or between the HRV amplicon and the 5’ end of the 3’ RNA replication element.
- the cleavable sequence is a self-cleaving ribozyme (e.g., a hammerhead ribozyme; Tang and Breaker. Proc Natl Acad Sci USA.2000 May 23;97(11):5784-9.
- a cargo RNA molecule that is integrated into a polynucleotide includes at least one CRISPR guide RNA; release of the guide RNA is mediated, e.g., by flanking DR sequences, ribozyme sequences, or other self-cleaving or trans-cleaving RNAs, or by cleavage by an endogenous ribonuclease.
- the corresponding Cas nuclease can be provided by separate or concurrent delivery, e.g., by co-delivery with a vector or polynucleotide, or by transient or stable expression of the corresponding Cas nuclease in the cell to which the polynucleotide is delivered.
- guide sequence designs are constrained by the requirement that the DNA target sequence (to which the crRNA is designed to be complementary) must be adjacent to a proto-spacer adjacent motif (“PAM”) sequence that is recognized by the specific Cas nuclease to be employed.
- PAM proto-spacer adjacent motif
- Cas nucleases recognize specific PAM sequences and there is a diversity of nucleases and corresponding PAM sequences; see, e.g., Smakov et al. (2017) Nature Reviews Microbiol., doi:10.1038/nrmicro.2016.184.
- Cas9 nucleases cleave dsDNA, require a GC-rich PAM sequence located 3’ to the DNA target sequence to be targeted by the crRNA component of the guide RNA, and cleave leaving blunt ends.
- Cas12a nucleases cleave dsDNA require a T-rich PAM sequence located 5’ to the DNA target sequence to be targeted by the crRNA component of the guide RNA, Agent Ref: P14357WO00 - 27 - and cleave leaving staggered ends with a 5’ overhang.
- Cas13 nucleases cleave single-stranded RNAs and do not require a PAM sequence; instead, Cas13 nuclease are guided to their targets by a single crRNA with a direct repeat (“DR”).
- the crRNA component of a guide RNA is generally designed to have a length of between 17 – 24 nucleotides (frequently 19, 20, or 21 nucleotides) and exact complementarity (i.
- Non-limiting examples of effective guide RNA design are found, e.g., in US Patent Application Publications US 2019/0032131, 2015/0082478, and 2019/0352655, which are each incorporated by reference in their entirety herein.
- CRISPR “arrays” can be designed to include one or multiple guide RNA sequences corresponding to one or more desired target DNA sequence(s); see, for example, Cong et al. (2013) Science, 339:819-823; Ran et al. (2013) Nature Protocols, 8:2281-2308.
- the 5’ RNA replication element includes a 5’ UTR element of an amalgavirus genome (e.g., either a CP or RdRP AV genome including an AV 5’ UTR set forth in Table 1).
- the 5’ RNA replication element further includes a genomic sequence of the amalgavirus that is natively located 3’ to and optionally adjacent or immediately adjacent to the 5’ UTR sequence.
- the 3’ RNA replication element includes a 3’ UTR sequence of an amalgavirus genome (e.g., either a CP or RdRP AV genome including an AV 3’ UTR set forth in Table 1).
- the 3’ RNA replication element further includes a genomic sequence of the amalgavirus that is natively located 5’ to and optionally adjacent or immediately adjacent to the 3’ UTR sequence.
- the RNA molecule further includes at least one RNA molecule encoding a viral MP.
- the at least one RNA molecule encoding an MP is located (a) before the cargo RNA molecule, (b) after the cargo RNA molecule, or (c) both before and after the cargo RNA molecule.
- the at least one RNA sequence encoding an MP includes at least two RNA sequences encoding different MPs or a single RNA sequence encoding multiple copies of MPs.
- the recombinant DNA molecule further includes a discrete expression cassette including a second promoter that is functional in the cell and is operably linked to a DNA sequence encoding at least one viral movement protein, and optionally a terminator element.
- the RNA molecule further includes an encapsidation recognition element (ERE), where the ERE is located close to or adjacent to the 3’ RNA replication element, and optionally wherein the 3’ RNA replication element includes a 3’ UTR sequence of the amalgavirus.
- ERE includes a viral OAS such as a tobacco mosaic virus OAS (TMV-OAS) or an OAS set forth in Table 2.
- the RNA molecule further includes at least one tRNA-like sequence (TLS), and wherein the at least one tRNA-like sequence includes a tRNA-like sequence from an Arabidopsis FT mRNA (e.g. a TLS in an Arabidopsis FT mRNA of Table 4).
- the RNA molecule includes a tRNA-like sequence encoded by a DNA sequence selected from the group consisting of SEQ ID NOs: 76-123, and 466.
- the RNA molecule includes a modified tRNA-like sequence that has at least 90% sequence identity to a scaffold tRNA-like sequence encoded by a DNA sequence selected from the group consisting of SEQ ID NOs: 76-123, and 466and that maintains the secondary structure of the scaffold tRNA-like sequence.
- the RNA molecule further includes at least one RNA encoding a viral MP, a tRNA-like sequence from an Arabidopsis FT mRNA, and an encapsidation recognition sequence including TMV-OAS.
- the cargo RNA molecule is up to about 3.5kb or more in length.
- the cargo RNA molecule includes: (a) at least one coding sequence, (b) at least one non- coding sequence, or (c) both at least one coding sequence and at least one non-coding sequence.
- the cargo RNA molecule includes at least one coding sequence, and wherein the RNA molecule further includes an internal ribosome entry site (IRES) located 5’ and immediately adjacent to the at least one coding sequence.
- the cargo RNA molecule includes multiple coding sequences, and wherein the RNA molecule further includes an IRES located 5’ and immediately adjacent to each of the coding sequences.
- the cargo RNA molecule includes at least one non-coding sequence, and wherein the at least one non-coding sequence is selected from the group consisting of a hairpin RNA (hpRNA); an RNA that forms multiple stem-loops; an RNA pseudoknot; an RNA sequence that forms at least partially double-stranded RNA; a small interfering RNA (siRNA) or siRNA precursor; a microRNA (miRNA) or miRNA precursor; a ribozyme; a ligand-responsive ribozyme (aptazyme); an RNA aptamer; and a long noncoding RNA (lncRNA).
- hpRNA hairpin RNA
- RNA pseudoknot an RNA sequence that forms at least partially double-stranded RNA
- siRNA small interfering RNA
- miRNA microRNA
- miRNA miRNA
- a ribozyme a ligand-responsive ribozyme
- RNA aptamer a long noncoding RNA (l
- a DNA sequence encoding at least one ribozyme is provided. In embodiments, the at least one ribozyme is located 5’ to the 5’ RNA replication element or 3’ to the 3’ RNA replication element. In embodiments, a DNA sequence encoding at least one ligand-responsive ribozyme (aptazyme) is provided. In embodiments, the at least one ligand-responsive ribozyme is located 5’ to the 5’ RNA replication element or 3’ to the 3’ RNA replication element.
- RNA molecules comprising the aforementioned or otherwise disclosed 5’ RNA replication elements, a cargo RNA molecule(s), and 3’ RNA replication elements, as well as additional aforementioned or otherwise disclosed elements are also provided herein.
- the recombinant RNA molecules are produced by a recombinant DNA molecule provided herein.
- the recombinant RNA molecules are produced by an in vivo or in vitro (e.g., cell free) RNA replication process through the action of a RdRP acting on: (i) 5’ and 3’ RNA replication elements; and/or (ii) a subgenomic promoter.
- RNA molecule comprising, in 5’ to 3’ order: (i) a 5’ RNA replication element; (ii) a cargo RNA molecule; and (iii) a 3’ RNA replication element; and, optionally, further comprising at least one additional RNA or other element.
- Embodiments of additional elements include at least one RNA encoding a viral MP, at least one tRNA-like sequence, an OAS, an RdRP protein that recognizes the 5’ and 3’ RNA replication elements, a subgenomic promoter, and/or an RdRP that recognizes an HRV 5’ or 3’ replication region and/or a subgenomic promoter.
- the RdRP protein is provided by an amalgavirus, e.g. an amalgavirus that is endogenous to a cell in which expression is desired.
- cell-based expression system can include (a) a recombinant DNA molecule including a heterologous promoter that is functional in a cell and is operably linked to a DNA sequence encoding an RNA molecule comprising, in 5’ to 3’ order: (i) a 5’ RNA replication element; (ii) a cargo RNA molecule; and (iii) a 3’ RNA replication element; and, optionally, further comprising at least one additional RNA or other element.
- Embodiments of additional elements include at least one RNA encoding a viral MP, at least one tRNA-like sequence, an OAS, an RdRP protein that recognizes the 5’ and 3’ RNA replication elements, a subgenomic promoter, and/or an RdRP that recognizes the subgenomic promoter.
- cell-based expression system can include (a) a recombinant RNA molecule comprising, in 5’ to 3’ order: (i) a 5’ RNA replication element; (ii) a cargo RNA molecule; and (iii) a 3’ RNA replication element; and, optionally, further comprising at least one additional RNA or other element.
- Embodiments of additional RNA elements include at least one RNA encoding a viral MP, at least one tRNA-like sequence, an OAS, an RdRP protein that recognizes the 5’ and 3’ RNA replication elements, a subgenomic promoter, and/or an RdRP that recognizes an HRV 5’ or 3’ replication region and/or a subgenomic promoter.
- the RdRP protein is provided by an amalgavirus.
- an RdRP that recognizes an HRV 5’ or 3’ replication region and/or a subgenomic promoter is provided by a heterologous RNA virus (HRV) or by another nucleic acid introduced into the cell (e.g., by a vector or other recombinant nucleic acid).
- HRV heterologous RNA virus
- the 5’ RNA replication element and the 3’ RNA replication element are obtained from the same amalgavirus and/or from the same amalgavirus genome (e.g., both obtained from the same AV capsid genome or both obtained from the same AV RdRP genome) or from amalgavirus genomes having at least 85%, 90%, 95%, 98%, or 99% sequence identity to one another and which are optionally related.
- a cell used in the expression system is a bacterial cell, a plant cell, a fungal cell, or an animal cell (e.g., an insect cell).
- a cell used in the expression system endogenously contains an amalgavirus having a genome that encodes an RdRP that recognizes the 5’ and 3’ RNA replication elements.
- the expression system further includes a viral capsid protein that is recognized by the encapsidation recognition element and encapsidates the RNA molecule.
- the viral capsid protein is: (a) expressed by the recombinant DNA molecule in the cell (e.g., where the recombinant DNA molecule further includes a discrete expression Agent Ref: P14357WO00 - 30 - cassette comprising a second promoter operably linked to a DNA sequence encoding the viral capsid protein, and optionally a terminator element), (b) co-expressed by a second recombinant DNA molecule in the cell; (c) provided exogenously to the cell; or (d) expressed by a virus in the cell.
- the RdRP protein is heterologous to the cell. In embodiments, the RdRP protein is provided exogenously to the cell.
- the RdRP protein that recognizes the 5’ and 3’ RNA replication elements is endogenously expressed in the plant cell by the amalgavirus (e.g., where the amalgavirus occurs naturally in the plant cell).
- the amalgavirus is native to or endemic to the plant cell.
- the amalgavirus that is endemic to the plant cell is non-pathogenic.
- the amalgavirus that is endemic to the plant cell is non-pathogenic and commensal.
- the amalgavirus is an exogenously introduced amalgavirus (i.e., not endemic or native to the host, but artificially introduced).
- an amalgavirus natively found in one plant species, variety, or germplasm can be introduced, with or without a corresponding recombinant amalgavirus satellite RNA, into a different plant species, variety, or germplasm.
- a complete self- replicating amalgavirus satellite system is introduced into a plant or plant cells, wherein the self- replicating amalgavirus satellite system includes: (1) a recombinant amalgavirus satellite RNA comprising, from 5’ terminus to 3’ terminus: (a) a 5’ RNA replication element recognized by an amalgavirus RNA-dependent RNA polymerase (RdRP); (b) a cargo RNA molecule; and (c) a 3’ RNA replication element recognized by the RdRP; wherein the 5’ RNA replication element, the cargo RNA molecule, and the 3’ RNA replication element are operably linked, and wherein the promoter and the cargo RNA molecule are heterologous to the 5’ RNA replication element and the 3
- the amalgavirus RdRP comprises a protein having at least 85%, 90%, 95%, 97%, or 99% sequence identity to SEQ ID NO: 476, 477, 478, 730 to 813, or 814.
- the recombinant DNA molecule or recombinant RNA molecule further comprises at least one RNA encoding a viral MP, a tRNA-like sequence from an Arabidopsis FT mRNA, and an encapsidation recognition sequence including a TMV-OAS.
- Cells comprising any of the aforementioned or otherwise disclosed recombinant polynucleotides are provided herein.
- Cells comprising the recombinant polynucleotides include prokaryotic (e.g., a bacterium, such as a bacterium capable of transforming a eukaryotic cell) or eukaryotic (e.g., a plant cell, fungal cell, or animal cell such as an insect cell) cells.
- the cells are bacterial cells capable of transforming a plant cell (e.g., an Agrobacterium sp., a Sinorhizobium sp., a Mesorhizobium sp., Bradyrhizobium sp., Rhizobium sp., or an Ensifer sp. cell.
- Bacterial cells capable of transforming a plant cell suitable for use with the recombinant polynucleotides provided herein include Agrobacterium sp., a Sinorhizobium sp., a Mesorhizobium sp., Bradyrhizobium sp., Rhizobium sp., or an Ensifer sp.
- Vectors suitable for maintenance, propagation, and/or expression of the recombinant polynucleotides in the aforementioned prokaryotic or eukaryotic cells are also provided herein.
- Such vectors can comprise any of the aforementioned or otherwise disclosed recombinant polynucleotides, recombinant DNA molecules, and recombinant RNA molecules as well as those polynucleotides molecules described in the Examples.
- the bacterium that mediates the plant transformation is an Agrobacterium sp., a Sinorhizobium sp., a Mesorhizobium sp., Bradyrhizobium sp., Rhizobium sp., or an Ensifer sp.
- the vector includes T-DNAs flanking the recombinant DNA molecule encoding the recombinant RNA molecule (e.g., as described in US patent application publications US20170369898 and US20180312854, each incorporated herein by reference in their entireties).
- the vector is contained within a plant cell or within a bacterial cell (Agrobacterium sp., a Sinorhizobium sp., a Mesorhizobium sp., Bradyrhizobium sp., Rhizobium sp., or an Ensifer sp. cell).
- a bacterial cell Agrobacterium sp., a Sinorhizobium sp., a Mesorhizobium sp., Bradyrhizobium sp., Rhizobium sp., or an Ensifer sp. cell.
- Viral particles comprising any of the aforementioned or otherwise disclosed recombinant RNA molecules are also provided.
- the recombinant RNA is introduced into a host or production plant by using Agrobacterium-mediated transformation with a polynucleotide comprising (5’ to 3’): (i) a promoter which is operably linked to a viral MP coding sequence and a TLS element, flanked by amalgavirus 5’ and 3’ RNA replication elements; (ii) a promoter which is operably linked to a cargo RNA molecule and a TLS element, flanked by amalgavirus 5’ and 3’ RNA replication elements; (iii) a promoter operably linked to an RdRP coding sequence; and (iv) a promoter operably linked to a CP encoding sequence.
- a polynucleotide comprising (5’ to 3’): (i) a promoter which is operably linked to a viral MP coding sequence and a TLS element, flanked by amalgavirus 5’ and 3’ RNA replication elements; (ii) a promote
- Heterologous promoters independently drive expression of the capsid protein and the cargo as depicted.
- the RNA expressed from the polynucleotide includes an OAS.
- a host plant is transformed for production of the synthetic amalgavirus satellite RNA and satellite particles comprising the encapsidated RNA.
- the expressed and encapsidated amalgavirus satellite RNA is subsequently isolated from leaf material or other tissue of the host plant, purified (and, if desired, formulated) for high pressure spraying onto plants that endogenously contain the corresponding amalgavirus or a recombinant (e.g., stably transformed or transiently expressed in the plant) source of the AV RdRP for subsequent expression and replication of the amalgavirus satellite RNA and satellite particles comprising the same in encapsidated form.
- spraying with the encapsidated satellite particles with certain cargo RNA molecules can be used to modify the plant as desired.
- plants without a systemic amalgavirus which provides the AV RdRP can further comprise a recombinant DNA or RNA molecule which encodes and provides the RdRP, e.g., stably integrated into the plant’s genome or transiently expressed in the plant.
- a Agent Ref: P14357WO00 - 32 - plant that transgenically or transiently expresses an amalgavirus RdRP is also useful for evaluating recombinant amalgavirus satellites in planta.
- Target plants and plant cells used as hosts for synthetic amalgavirus satellite RNAs include both monocot and dicot plants and plant cells which can support amalgavirus replication.
- the amalgavirus is endogenous to (endemic to or natively found in) the plant or plant cell.
- the amalgavirus is introduced to and becomes established in the plant or plant cell.
- Embodiments include row crop plants, fruit-producing plants and trees, vegetables, trees, and ornamental plants including ornamental flowers, shrubs, trees, groundcovers, and turf grasses.
- the host plants and plant cells for synthetic amalgavirus satellite RNAs include row crop plants, fruit-producing plants and trees, vegetables, trees, and ornamental plants including ornamental flowers, shrubs, trees, groundcovers, and turf grasses.
- the host plants and plant cells for synthetic amalgavirus satellite RNAs include commercially important cultivated crops, trees, and plants, including: alfalfa (Medicago sativa), almonds (Prunus dulcis), apples (Malus x domestica), apricots (Prunus armeniaca, P. brigantine, P. mandshurica, P. mume, P.
- sibirica asparagus (Asparagus officinalis), bananas (Musa spp.), barley (Hordeum vulgare), beans (Phaseolus spp.), blueberries and cranberries (Vaccinium spp.), cacao (Theobroma cacao), canola and rapeseed or oilseed rape, (Brassica napus), Polish canola (Brassica rapa), and related cruciferous vegetables including broccoli, kale, cabbage, and turnips (Brassica carinata, B. juncea, B. oleracea, B. napus, B. nigra, and B.
- Coffea arabica, Coffea canephora, and Coffea liberica cotton (Gossypium hirsutum L.), cowpea (Vigna unguiculata and other Vigna spp.), fava beans (Vicia faba), cucumber (Cucumis sativus), currants and gooseberries (Ribes spp.), date (Phoenix dactylifera), duckweeds (family Lemnoideae), eggplant or aubergine (Solanum melongena), eucalyptus (Eucalyptus spp.), flax (Linum usitatissumum L.), geraniums (Pelargonium spp.), grapefruit (Citrus x paradisi), grapes (Vitus spp.) including wine grapes (Vitus vinifera and hybrids thereof), guava (Psidium guajava), hops (Humulus lupulus), hemp and cannabis
- the host plant or plant cells for synthetic amalgavirus satellite RNAs is a dicot plant or plant cell selected from the genera Arachis, Brassica, Capsicum, Cicer, Cucumis, Cucurbita, Gossypium, Lens, Nicotiana, Solanum, Phaseolus, Pisum, Vicia, Vigna, or Glycine.
- the host plant or plant cells for synthetic amalgavirus satellite RNAs is a monocot plant or plant cell selected from the genera Allium, Avena, Festuca, Hordeum, Lilium, Lolium, Miscanthus, Oryza, Saccharum, Secale, Setaria, Sorghum, x Triticosecale, Triticum, or Zea.
- monocot target plants and plant cells used as hosts for synthetic amalgavirus satellite RNAs include oats (Avena sativa), barley (Hordeum vulgare), rice (Oryza sativa, Oryza glaberrima, Oryza rufipogen), rye (Secale cereale), wheat (Triticum aestivum), sorghum (Sorghum bicolor), and maize (Zea mays) plants and plant cells.
- the recombinant RNA molecule or a formulation thereof is provided by contacting the plant or plant cell with the recombinant RNA molecule or formulation thereof.
- the recombinant RNA molecule is provided by expressing in the plant or plant cell a DNA molecule that encodes the recombinant RNA molecule or a formulation.
- the recombinant RNA molecule is provided by contacting the plant or plant cell with cells (such as bacterial cells) which comprise a DNA molecule that encodes the recombinant RNA molecule and are capable of transforming the plant or plant cell.
- the recombinant RNA molecule is provided by contacting the plant or plant cell with a satellite particle comprising an encapsidated recombinant RNA molecule or a formulation thereof.
- the 5’ RNA replication element has a nucleotide sequence obtained or derived from an amalgavirus genomic sequence; and/or the 3’ RNA replication element has a nucleotide sequence obtained or derived from an amalgavirus genomic sequence.
- the 5’ and/or 3’ RNA replication element can be obtained from the corresponding amalgavirus genomic sequence by synthesizing or cloning a copy of the corresponding amalgavirus genomic sequence.
- the 5’ and/or 3’ RNA replication element can be Agent Ref: P14357WO00 - 34 - derived from the corresponding amalgavirus genomic sequence by synthesizing a copy of a modified amalgavirus genomic sequence or sequences.
- Such modifications of amalgavirus genomic sequences present in a derived sequence include: (i) substitutions of nucleotides which maintain the RNA secondary structure; (ii) substitution of nucleotides based on a consensus obtained by alignment of 5’ or 3’ RNA replication elements; (iii) insertions, deletions, and/or substitution of nucleotides to facilitate assembly and/or operable linkage to other elements in the satellite RNA which include cargo RNA molecules, tRNA-like elements, encapsidation recognition element (ERE), RNA encoding a viral movement protein (MP), IRES elements, an HRV 5’RR, HRV 3’RR, and/or HRV subgenomic promoter; or (iv) any combination of (i), (ii), or (iii).
- the plant cell includes the amalgavirus, and the RdRP protein is provided to the plant cell by the amalgavirus.
- the amalgavirus is endemic to the plant cell.
- the amalgavirus that is endemic to the plant cell is non- pathogenic and/or commensal to the plant cell.
- the amalgavirus is exogenously provided to the plant cell.
- the RdRP protein is exogenously provided to the plant cell.
- the recombinant RNA molecule is produced in a fermentation system.
- the recombinant RNA molecule is provided to the plant cell by transcribing in the plant cell a recombinant DNA construct including a promoter functional in the plant cell and operably linked to a DNA sequence encoding the recombinant RNA molecule.
- the recombinant RNA molecule further includes an encapsidation recognition element (ERE), and the plant cell further includes a viral capsid protein (CP) capable of encapsidating the synthetic amalgavirus satellite RNA.
- EEE encapsidation recognition element
- CP viral capsid protein
- the viral capsid protein is exogenously provided to the plant cell.
- the recombinant DNA construct further includes a DNA sequence encoding a viral capsid protein.
- the recombinant DNA construct further includes a second promoter functional in the plant cell and operably linked to the DNA sequence encoding the viral capsid protein.
- the viral capsid protein is expressed in the plant cell and encapsidates the synthetic amalgavirus satellite RNA.
- the plant cell includes the amalgavirus, and the amalgavirus provides to the plant cell: (a) the RdRP protein, (b) the viral capsid protein, or (c) both the RdRP protein and the viral capsid protein.
- the methods can further comprise a first step of providing a population of plants comprising the plant cells comprising: (i) the amalgavirus which provides the RdRP; or (ii) recombinant polynucleotide molecule that encodes the RdRP; and then providing the recombinant RNA molecule to the plants comprising the plant cells.
- the methods can further comprise the step of determining if the plant cell comprises an amalgavirus which can provide the RdRP.
- the plant cell comprises the amalgavirus which can provide the RdRP, the amalgavirus, the RdRP protein, and/or the recombinant polynucleotide encoding the RdRP is optionally not exogenously provided to the plant cell.
- the RdRP protein or the recombinant polynucleotide encoding the RdRP is exogenously provided to the plant cell, or a Agent Ref: P14357WO00 - 35 - combination of the amalgavirus, RdRP protein, or polynucleotide encoding the RdRP is exogenously provided to the plant cell.
- a complete self-replicating amalgavirus satellite system is introduced into a plant or plant cells, wherein the self-replicating amalgavirus satellite system includes: (1) a recombinant amalgavirus satellite RNA comprising, from 5’ terminus to 3’ terminus: (a) a 5’ RNA replication element recognized by an amalgavirus RNA-dependent RNA polymerase (RdRP); (b) a cargo RNA molecule; and (c) a 3’ RNA replication element recognized by the RdRP; wherein the 5’ RNA replication element, the cargo RNA molecule, and the 3’ RNA replication element are operably linked, and wherein the promoter and the cargo RNA molecule are heterologous to the 5’ RNA replication element and the 3’ RNA replication element; and (2) an exogenous amalgavirus (e.g., an amalgavirus that is not endemic or native to the plant or plant cells) that is capable of replication in the plant or plant cells and that encode
- the amalgavirus RdRP comprises a protein having at least 85%, 90%, 95%, 97%, or 99% sequence identity to SEQ ID NO: 476, 477, 478, 730 to 813, or 814.
- the presence or absence of an amalgavirus in a target plant can be determined by an RNA detection assay (e.g., an RT-PCR assay) using nucleic acid probes and/or primers which can detect any part of an amalgavirus genome including a 5’ RNA replication element, a CP and/or RdRP coding region, and/or a 3’ RNA replication element.
- Such probes and primers include those which detect any of the 5’ or 3’ RNA replication elements set forth in Table 1 or having significant sequence identity thereto (e.g., at least about 80%, 85%, 90%, 95%, 98%, or 99% of a length of at least about 18, 20, 30, 40 or 50 nt).
- the presence or absence of an amalgavirus in a target plant can be determined by a protein detection assay (e.g., an immunoassay) directed to an AV CP or RdRP (e.g., a CP or RdRP encoded by or homologous to a CP or RdRP encoded by an AV genome disclosed in Table 1).
- Target plants and plant cells used in the methods include all aforementioned target plants and plant cell hosts for synthetic amalgavirus satellite RNAs (e.g., recombinant RNAs).
- the recombinant RNA that effects: (i) a phenotypic change in the plant or plant cell; (ii) increases a plant’s resistance to a pest or pathogen; or (iii) increases a plant’s resistance to stress can include an RNA for modulating a target gene’s expression relative to the target gene’s expression in a control plant or plant cell not provided with the recombinant RNA molecule, and the phenotypic change, increased resistance to the pest or pathogen, or increased resistance to stress is a result of the modulation.
- the modulation is (a) an increase of the target gene’s expression; or (b) a decrease of the target gene’s expression.
- expression of the target gene is increased by up to about 1%, 2%, 3%, %, 5%, 6%, 7%, 8%, 9%, 10%, 11%, 12%, 13%, 14%, 15%, 16%, 17%, 18%, 19%, 20%, 21%, 22%, 23%, 24%, 25%, 26%, 27%, 28%, 29%, 30%, 31%, 32%, 33%, 34%, 35%, 36%, 37%, 38%, 39%, 40%, 41%, 42%, 43%, 44%, 45%, 46%, 47%, 48%, 49%, 50%, 51%, 52%, 53%, 54%, 55%, 56%, 57%, 58%, 59%, 60%, 61%, 62%, 63%, 64%, 65%, 66%, 67%, 68%, 69%, 70%, 71%
- expression of the target gene is increased by up to about 2-, 3-, 4-, 5-, 6-, 8-, 9-, 10-fold, or more relative to a reference level (e.g., a level found in a control plant or plant cell lacking the recombinant RNA molecule).
- a reference level e.g., a level found in a control plant or plant cell lacking the recombinant RNA molecule.
- expression of the target gene is decreased by at least about 1%, 2%, 3%, 4%, 5%, 6%, 7%, 8%, 9%, 10%, 11%, 12%, 13%, 14%, 15%, 16%, 17%, 18%, 19%, 20%, 21%, 22%, 23%, 24%, 25%, 26%, 27%, 28%, 29%, 30%, 31%, 32%, 33%, 34%, 35%, 36%, 37%, 38%, 39%, 40%, 41%, 42%, 43%, 44%, 45%, 46%, 47%, 48%, 49%, 50%, 51%, 52%, 53%, 54%, 55%, 56%, 57%, 58%, 59%, 60%, 61%, 62%, 63%, 64%, 65%, 66%, 67%, 68%, 69%, 70%, 71%, 72%, 73%, 74%, 75%, 76%, 77%, 78%, 79%, 80%, 81%, 82%, 83%
- RNAs for modifying the genome include gRNAs recognized by CAS nucleases, RNAs encoding TALENs or artificial zinc finger proteins (aZFN).
- RNAs for modifying the epigenome include RNAs which provide RNA directed DNA methylation such as in promoter regions of target genes (Matzke and Mosher (2014); doi: 10.1038/nrg3683).
- Embodiments of an RNA for modifying the transcriptome include one or more RNAs that comprise any of: a hairpin RNA (hpRNA); an RNA that forms multiple stem-loops; an RNA pseudoknot; an RNA molecule that forms at least partially double-stranded RNA; a small interfering RNA (siRNA) or siRNA precursor; a microRNA (miRNA) or miRNA precursor; a phased siRNA or phased siRNA precursor (see, e.g., US Patent No.8,404,928); a ribozyme; a ligand-responsive ribozyme (aptazyme); an RNA aptamer; or a long noncoding RNA (lncRNA).
- hpRNA hairpin RNA
- the cargo RNA molecule can comprise an RNA that effects a phenotypic change in the plant or plant cell in comparison to a plant or plant cell lacking the recombinant RNA.
- phenotypes that are changed include developmental rate, growth rate, size, yield (e.g., intrinsic yield), vigor, photosynthetic capability, flavor, starch production, protein content, carbohydrate content, oil content, fatty acid content, lipid content, digestibility, biomass, shoot length, root length, root architecture, seed set, seed weight, seed quality (e.g., nutritional content), germination, fruit set, rate of fruit ripening, production of biopolymers, production of fibers, production of biofuels, production of pharmaceutical peptides (e.g., hormones, enzymes, transcription factors, antigens, antibodies, or antibody fragments), production of secretable peptides, enzyme production, improved processing traits, or amount of harvestable produce.
- developmental rate e.g., growth rate, size, yield (e.g., intrinsic yield), vigor, photosynthetic capability, flavor, starch production, protein content, carbohydrate content, oil content, fatty acid content, lipid content, digestibility, biomass, shoot length, root length
- phenotypes that are changed include taste, appearance, or shelf-life of a product harvested from the plant.
- phenotypes that are changed include flower size, flower color, flower patterning, flower morphology including presence or absence of stamens, flower number, flower longevity, flower fragrance, leaf size, leaf color, leaf patterning, leaf morphology, plant height, or plant architecture.
- the recombinant RNA can comprise an RNA that inhibits expression of a gene of the pest or pathogen and/or inhibits replication of the genome of the pest or pathogen.
- the pest or pathogen is selected from the group comprising: a bacterium, a virus other than an amalgavirus, a fungus, an oomycete, and an invertebrate (e.g., an arthropod or a nematode).
- Target viruses other than an amalgavirus include; (i) positive-strand RNA viruses in the Bromoviridae, Closteroviridae, Luteoviridae, or Potyviridae family; (ii) negative strand RNA viruses in the Bunyaviridae and Rhabdoviridae family; (iii) dsDNA viruses in the family Caulimoviridae; and (iv) ssDNA viruses in the family Geminiviridae.
- Target arthropods pests include coleopteran and lepidopteran insects.
- Target fungal pathogens include Magnaporthe spp., Botrytis spp., Puccinia spp.; Fusarium spp., Blumeria spp., Mycosphaerella spp., Colletotrichum spp., Ustilago spp., Melampsora spp., Phakopsora spp., Phytophthora spp., and Rhizoctonia spp.
- the cargo RNA molecule effects an increase in the plant’s resistance to a pest or pathogen, relative to that in a plant not provided with the recombinant RNA molecule.
- the recombinant RNA can comprise an RNA that targets a plant gene which provides such resistance.
- the RNA that effects an increase in the plant’s resistance to stress in the plant or plant cell includes an RNA for modulating the target gene’s expression relative to the target gene’s expression in a control plant or plant cell not provided with the recombinant RNA molecule, and wherein the increase stress resistance is a result of the modulation.
- the modulation is (a) an increase of the target gene’s expression; or (b) a decrease of the target gene’s expression.
- the RNA that effects an increase in the plant’s resistance to stress in the plant or plant cell comprises a messenger RNA encoding a protein which confers the stress resistance.
- the messenger RNA includes an RNA sequence absent in the transcriptome of the plant or plant cell lacking the recombinant RNA.
- the stress includes at least one abiotic stress selected from the group including: nutrient stress, light stress, drought stress, heat stress, and cold stress.
- the stress includes at least one biotic stress selected from the group including: crowding, shading, and allelopathy (e.g., resulting from allelopathic chemicals including a juglone produced by walnut trees).
- the cargo RNA can encode the exogenous polypeptide.
- the polypeptide is isolated (e.g., separated from at least one other cellular components such as a carbohydrate, a lipid, or another protein) or polypeptide is purified.
- manufacture can occur in either a cell-based system or a cell-free system.
- Cell-based methods of manufacturing a synthetic amalgavirus satellite particle can comprise: (a) providing to a plant cell the recombinant RNA molecule, wherein the recombinant RNA molecule comprises an encapsidation recognition element (ERE), and wherein the plant cell comprises an RdRP protein that recognizes the 5’ RNA replication element and 3’ RNA replication element catalyzes synthesis of a synthetic amalgavirus satellite RNA from the recombinant RNA molecule and a viral capsid protein, wherein the ERE provides for encapsidation of the RNA by the viral capsid protein; and optionally isolating the synthetic amalgavirus satellite particle from the plant cell, a plant comprising the plant cell, or from media in which the plant cell or the plant had been grown.
- ERP encapsidation recognition element
- Cell-free methods of manufacturing a synthetic amalgavirus satellite particle include methods where the recombinant RNA molecule is combined with a viral capsid protein in a vessel, wherein the recombinant RNA molecule comprises an ERE, and wherein the ERE provides for encapsidation of the RNA by the viral capsid protein in the vessel; optionally wherein the method further comprises isolating the synthetic amalgavirus satellite particle from uncombined RNA and/or viral capsid protein in the vessel.
- the synthetic satellite particle is isolated (e.g., separated from at least one other cellular components such as an organelle, a membrane, a carbohydrate, a lipid, or another protein) or is purified.
- the methods can further comprise formulating the synthetic amalgavirus satellite particle.
- Synthetic amalgavirus satellite particles comprising the recombinant RNA, including those made by the aforementioned methods are also provided.
- Methods of providing any of the aforementioned synthetic amalgavirus satellite particles to a plant including contacting (e.g., spraying, dusting, injecting, soaking, etc.) the plant with the synthetic amalgavirus satellite particle or a formulation thereof are also provided.
- the recombinant polynucleotides, cells comprising the same, and synthetic amalgavirus satellite particles described herein can be formulated either in pure form (e.g., the composition contains only the recombinant polynucleotide) or together with one or more additional formulation components to facilitate application or delivery of the compositions.
- the additional formulation component includes, e.g., a carrier (i.e., a component that has an active role in delivering the active agent (e.g., recombinant polynucleotide); for example, a carrier can encapsulate, covalently or non-covalently modify, or otherwise associate with the active agent in a manner that improves delivery of the active agent) or an excipient (e.g., a delivery vehicle, adjuvant, diluent, surfactant, stabilizer, or tonicity agent).
- the composition is formulated for delivery to a plant.
- the disclosure provides a formulation comprising any of the compositions described herein.
- the formulation is a liquid, a gel, or a powder.
- the formulation is configured to be sprayed on plants, to be injected into plants or otherwise introduced into the vascular system of a plant, to be rubbed on leaves, to be soaked into plants, Agent Ref: P14357WO00 - 39 - to be coated onto plants, or be coated on seeds, or to be delivered through root uptake (e.g., in a hydroponic system or via soil).
- the composition can be formulated into emulsifiable concentrates, suspension concentrates, directly sprayable or dilutable solutions, coatable pastes, diluted emulsions, spray powders, soluble powders, dispersible powders, wettable powders, dusts, granules, encapsulations in polymeric substances, microcapsules, foams, aerosols, carbon dioxide gas preparations, or tablets.
- the composition is a liquid.
- the composition is a solid.
- the composition is an aerosol, such as in a pressurized aerosol can.
- the recombinant polynucleotide makes up about 0.1% to about 100% of the composition, such as any one of about 0.01% to about 100%, about 1% to about 99.9%, about 0.1% to about 10%, about 1% to about 25%, about 10% to about 50%, about 50% to about 99%, or about 0.1% to about 90% of active ingredients (e.g., recombinant polynucleotides).
- the composition includes at least any of 0.1%, 0.5%, 1%, 5%, 10%, 20%, 30%, 40%, 50%, 60%, 70%, 80%, 90%, 95%, or more active ingredients (e.g., recombinant polynucleotides).
- the concentrated agents are preferred as commercial products, the final user normally uses diluted agents, which have a substantially lower concentration of active ingredient.
- the composition is formulated for topical delivery to a plant.
- the topical delivery is spraying, leaf rubbing (e.g., with or without an abrasive), soaking, coating (e.g., coating using micro-particulates or nano-particulates), or delivery through root uptake (e.g., delivery in a hydroponic system or by a root drench).
- the composition further comprises a carrier and/or an excipient.
- the composition does not comprise a carrier or excipient, e.g., comprises a naked polynucleotide (e.g., a naked RNA).
- a naked polynucleotide e.g., a naked RNA
- the recombinant polynucleotide is delivered at a concentration of at least 0.1 grams per acre, e.g., at least 0.1, 1, 2, 5, 10, 15, 20, 25, 30, 35, 40, 45, or 50 grams per acre. In some embodiments, less than 120 liters per acre is delivered, e.g., less than 110, 100, 90, 80, 70, 60, 50, 40, 30, 20, 15, 10, 5, or 2 liters per acre or less than 1 liter per acre.
- the formulation comprises a carrier.
- the formulation is an emulsion or a reverse emulsion, a liquid, or a gel.
- the formulation includes a carrier that serves as a physical support (e.g., solid or semi-solid surfaces or matrices, powders, or particles or nanoparticles).
- the active agent is encapsulated or enclosed in or attached to or complexed with a carrier including a liposome, vesicle, micelle, or other fluid compartment.
- the active agent is encapsulated or enclosed in or attached to or complexed with a carrier including a naturally occurring or synthetic, branched or linear polymer (e.g., pectin, agarose, chitin, chitosan, DEAE-dextran, polyvinylpyrrolidone (“PVP”), or polyethylenimine (“PEI”).
- a carrier including a naturally occurring or synthetic, branched or linear polymer (e.g., pectin, agarose, chitin, chitosan, DEAE-dextran, polyvinylpyrrolidone (“PVP”), or polyethylenimine (“PEI”).
- PVP polyvinylpyrrolidone
- PEI polyethylenimine
- the carrier includes cations or a cationic charge, such as cationic liposomes or cationic polymers such as Agent Ref: P14357WO00 - 40 - polyamines (e.g., sper
- the carrier includes a polypeptide such as an enzyme, (e.g., cellulase, pectolyase, maceroenzyme, pectinase), a cell penetrating or pore- forming peptide (e.g., poly-lysine, poly-arginine, or polyhomoarginine peptides).
- an enzyme e.g., cellulase, pectolyase, maceroenzyme, pectinase
- a cell penetrating or pore- forming peptide e.g., poly-lysine, poly-arginine, or polyhomoarginine peptides.
- Non-limiting examples of carriers include cationic liposomes and polymer nanoparticles such as those reviewed by Zhang et al. (2007) J. Controlled Release, 123:1 - 10, and the cross-linked multilamellar liposomes described in US Patent Application Publication 2014/035
- the carrier includes a nanomaterial, such as carbon or silica nanoparticles, carbon nanotubes, carbon nanofibers, or carbon quantum dots.
- a nanomaterial such as carbon or silica nanoparticles, carbon nanotubes, carbon nanofibers, or carbon quantum dots.
- Non-limiting examples of carriers include particles or nanoparticles (e.g., particles or nanoparticles made of materials such as carbon, silicon, silicon carbide, gold, tungsten, polymers, or ceramics) in various size ranges and shapes, magnetic particles or nanoparticles (e.g., silenceMag Magnetotransfection TM agent, OZ Biosciences, San Diego, CA), abrasive or scarifying agents, needles or microneedles, matrices, and grids.
- particles or nanoparticles e.g., particles or nanoparticles made of materials such as carbon, silicon, silicon carbide, gold, tungsten, polymers, or ceramics
- magnetic particles or nanoparticles e.g., silenceMag Magneto
- particulates and nanoparticulates are useful in delivery of the polynucleotide composition or the nuclease or both.
- Useful particulates and nanoparticles include those made of metals (e.g., gold, silver, tungsten, iron, cerium), ceramics (e.g., aluminum oxide, silicon carbide, silicon nitride, tungsten carbide), polymers (e.g., polystyrene, polydiacetylene, and poly(3,4- ethylenedioxythiophene) hydrate), semiconductors (e.g., quantum dots), silicon (e.g., silicon carbide), carbon (e.g., graphite, graphene, graphene oxide, or carbon nanosheets, nanocomplexes, or nanotubes), and composites (e.g., polyvinylcarbazole/graphene, polystyrene/graphene, platinum/graphene, palladium/graphene nanocomposites
- such particulates and nanoparticulates are further covalently or non-covalently functionalized, or further include modifiers or cross-linked materials such as polymers (e.g., linear or branched polyethylenimine, poly-lysine), polynucleotides (e.g., DNA or RNA), polysaccharides, lipids, polyglycols (e.g., polyethylene glycol, thiolated polyethylene glycol), polypeptides or proteins, and detectable labels (e.g., a fluorophore, an antigen, an antibody, or a quantum dot).
- polymers e.g., linear or branched polyethylenimine, poly-lysine
- polynucleotides e.g., DNA or RNA
- polysaccharides e.g., DNA or RNA
- lipids lipids
- polyglycols e.g., polyethylene glycol, thiolated polyethylene glycol
- Embodiments of compositions including particulates include those formulated, e.g., as liquids, colloids, dispersions, suspensions, aerosols, gels, and solids.
- Embodiments include nanoparticles affixed to a surface or support, e.g., an array of carbon nanotubes vertically aligned on a silicon or copper wafer substrate.
- Embodiments include polynucleotide compositions including particulates (e.g., gold or tungsten or magnetic particles) delivered by a Biolistic-type technique or with magnetic force.
- the size of the particles used in Biolistics is generally in the “microparticle” range, for example, gold microcarriers in the 0.6, 1.0, and 1.6 micrometer size ranges (see, e.g., instruction manual for the Helios® Gene Gun System, Bio-Rad, Hercules, CA; Randolph-Anderson et al.
- nanoparticles which are generally in the nanometer (nm) size range or less than 1 micrometer, e.g., with a diameter of less than about 1 nm, less than about 3 nm, less than about 5 nm, less than about 10 nm, less than about 20 nm, less than about 40 nm, less than about 60 nm, less than about 80 nm, and less than about 100 nm.
- nanoparticles commercially available (all from Sigma-Aldrich Corp., St.
- Louis, MO include gold nanoparticles with diameters of 5, 10, or 15 nm; silver nanoparticles with particle sizes of 10, 20, 40, 60, or 100 nm; palladium “nanopowder” of less than 25 nm particle size; single-, double-, and multi-walled carbon nanotubes, e.g., with diameters of 0.7 - 1.1, 1.3 - 2.3, 0.7 - 0.9, or 0.7 - 1.3 nm, or with nano tube bundle dimensions of 2 - 10 nm by 1- 5 micrometers, 6 - 9 nm by 5 micrometers, 7 - 15 nm by 0.5 - 10 micrometers, 7 - 12 nm by 0.5 - 10 micrometers, 110 - 170 nm by 5 - 9 micrometers, 6 - 13 nm by 2.5 - 20 micrometers.
- Embodiments include polynucleotide compositions including materials such as gold, silicon, cerium, or carbon, e.g., gold or gold-coated nanoparticles, silicon carbide whiskers, carborundum, porous silica nanoparticles, gelatin/silica nanoparticles, nanoceria or cerium oxide nanoparticles (CNPs), carbon nanotubes (CNTs) such as single-, double-, or multi-walled carbon nanotubes and their chemically functionalized versions (e.g., carbon nanotubes functionalized with amide, amino, carboxylic acid, sulfonic acid, or polyethylene glycol moieties), and graphene or graphene oxide or graphene complexes; see, for example, Wong et al.
- materials such as gold, silicon, cerium, or carbon, e.g., gold or gold-coated nanoparticles, silicon carbide whiskers, carborundum, porous silica nanoparticles, gelatin/silica nanoparticle
- the composition includes an excipient, e.g., a delivery vehicle, adjuvant, diluent, surfactant, stabilizer, or tonicity agent or a combination thereof.
- the excipient is a crop oil concentrate, a vegetable oil concentrate, a modified vegetable oil, a nitrogen source, a deposition (drift control) and/or retention agent (with or without ammonium sulfate and/or defoamer), a compatibility agent, a buffering agent and/or acidifier, a water conditioning agent, a basic blend, a spreader-sticker and/or extender, an adjuvant plus foliar fertilizer, an antifoam agent, a foam marker, a scent, or a tank cleaner and/or neutralizer.
- the excipient is an adjuvant described in the Compendium of Herbicide Adjuvants (Young et al. (2016).
- delivery vehicles and diluents include, but are not limited to, lactose, dextrose, sucrose, sorbitol, mannitol, starches, gum acacia, calcium phosphate, alginates, tragacanth, gelatin, calcium silicate, microcrystalline cellulose, polyvinylpyrrolidone, cellulose, water, saline solution, syrup, methylcellulose, methyl- and propylhydroxybenzoates, talc, magnesium stearate, and mineral oil.
- delivery vehicles include, but are not limited to, solid or liquid excipient materials, solvents, stabilizers, slow-release excipients, colorings, and surface-active substances (surfactants).
- the excipient e.g., delivery vehicle
- the stabilizing vehicle includes, e.g., an epoxidized vegetable oil, an antifoaming agent, e.g. silicone oil, a preservative, a viscosity regulator, a binding agent, or a tackifier.
- the stabilizing vehicle is a buffer suitable for the recombinant polynucleotide.
- the composition is microencapsulated in a polymer bead delivery vehicle.
- the stabilizing vehicle protects the recombinant polynucleotide against UV and/or acidic conditions.
- the delivery vehicle contains a pH buffer.
- the composition is formulated to have a pH in the range of about 4.5 to about 9.0, including for example pH ranges of about any one of 5.0 to about 8.0, about 6.5 to about 7.5, or about 6.5 to about 7.0.
- the composition provided herein includes an adjuvant.
- Adjuvants are agents that do not possess the polynucleotide activity, but impart beneficial properties to a formulation. For example, adjuvants are either pre-mixed in the formulation or added to a spray tank to improve mixing or application or to enhance performance.
- Adjuvants can be used to customize the formulation to specific needs and compensate for local conditions. Adjuvants can be designed to perform specific functions, including wetting, spreading, sticking, reducing evaporation, reducing volatilization, buffering, emulsifying, dispersing, reducing spray drift, and reducing foaming. No single adjuvant can perform all these functions, but compatible adjuvants often can be combined to perform multiple functions simultaneously.
- adjuvants included in the formulation are binders, dispersants and stabilizers, specifically, for example, casein, gelatin, polysaccharides (e.g., starch, gum arabic, cellulose derivatives, alginic acid, etc.), lignin derivatives, bentonite, sugars, synthetic water-soluble polymers (e.g., polyvinyl alcohol, polyvinylpyrrolidone, polyacrylic acid, etc.), PAP (acidic isopropyl phosphate), BHT (2,6-di-t-butyl-4-methylphenol), BHA (a mixture of 2-t-butyl-4-methoxyphenol and 3- t-butyl-4-methoxyphenol), vegetable oils, mineral oils, fatty acids and fatty acid esters.
- binders specifically, for example, casein, gelatin, polysaccharides (e.g., starch, gum arabic, cellulose derivatives, alginic acid, etc.), lignin derivatives, benton
- the compositions provided herein are in a liquid formulation.
- Liquid formulations are generally mixed with water, but in some instances are used with crop oil, diesel fuel, kerosene, or other light oil as an excipient.
- the amount of active ingredient e.g., recombinant polynucleotides
- an emulsifiable concentrate formulation contains a liquid active ingredient, one or more petroleum-based solvents, and an agent that allows the formulation to be mixed with water to form an emulsion.
- Such concentrates can be used in agricultural, ornamental and turf, forestry, Agent Ref: P14357WO00 - 43 - structural, food processing, livestock, and public health pest formulations.
- these are adaptable to application equipment from small portable sprayers to hydraulic sprayers, low-volume ground sprayers, mist blowers, and low-volume aircraft sprayers.
- Some active ingredients readily dissolve in a liquid excipient. When mixed with an excipient, they form a solution that does not settle out or separate, e.g., a homogenous solution.
- formulations of these types include an active ingredient, a carrier and/or an excipient, and one or more other ingredients. Solutions can be used in any type of sprayer, indoors and outdoors.
- the composition is formulated as an invert emulsion.
- An invert emulsion is a water-soluble active ingredient dispersed in an oil excipient.
- Invert emulsions require an emulsifier that allows the active ingredient to be mixed with a large volume of petroleum-based excipient, usually fuel oil.
- Invert emulsions aid in reducing drift. With other formulations, some spray drift results when water droplets begin to evaporate before reaching target surfaces; as a result, the droplets become very small and lightweight. Because oil evaporates more slowly than water, invert emulsion droplets shrink less, and more active ingredient reaches the target. Oil further helps to reduce runoff and improve rain resistance.
- a flowable or liquid formulation combines many of the characteristics of emulsifiable concentrates and wettable powders. Manufacturers use these formulations when the active ingredient is a solid that does not dissolve in either water or oil. The active ingredient, impregnated on a substance such as clay, is ground to a very fine powder. The powder is then suspended in a small amount of liquid. The resulting liquid product is quite thick.
- Flowables and liquids share many of the features of emulsifiable concentrates, and they have similar disadvantages. They require moderate agitation to keep them in suspension and leave visible residues, similar to those of wettable powders.
- Flowables/liquids are easy to handle and apply. Because they are liquids, they are subject to spilling and splashing. They contain solid particles, so they contribute to abrasive wear of nozzles and pumps. Flowable and liquid suspensions settle out in their containers. Because flowable and liquid formulations tend to settle, packaging in containers of five gallons or less makes remixing easier.
- Aerosol formulations contain one or more active ingredients and a solvent. Most aerosols contain a low percentage of active ingredients.
- Ready to use aerosol formulations are usually small, self-contained units that release the formulation when the nozzle valve is triggered. The formulation is driven through a fine opening by an inert gas under pressure, creating fine droplets. These products are used in greenhouses, in small areas Agent Ref: P14357WO00 - 44 - inside buildings, or in localized outdoor areas. Commercial models, which hold five to 5 pounds of active ingredient, are usually refillable.
- Smoke or fog aerosol formulations are not under pressure.
- the composition comprises a liquid excipient.
- a liquid excipient includes, for example, aromatic or aliphatic hydrocarbons (e.g., xylene, toluene, alkylnaphthalene, phenylxylylethane, kerosene, gas oil, hexane, cyclohexane, etc.), halogenated hydrocarbons (e.g., chlorobenzene, dichloromethane, dichloroethane, trichloroethane, etc.), alcohols (e.g., methanol, ethanol, isopropyl alcohol, butanol, hexanol, benzyl alcohol, ethylene glycol, etc.), ethers (e.g., diethyl ether, ethylene glycol
- the composition comprises a gaseous excipient.
- Gaseous excipients include, for example, butane gas, floron gas, liquefied petroleum gas (LPG), dimethyl ether, and carbon dioxide gas.
- LPG liquefied petroleum gas
- the compositions are provided as a dry formulation.
- Dry formulations can be divided into two types: ready-to-use and concentrates that must be mixed with water to be applied as a spray. Most dust formulations are ready to use and contain a low percentage of active ingredients (less than about 10 percent by weight), plus a very fine, dry inert excipient (e.g., talc, chalk, clay, nut hulls, or volcanic ash). The size of individual dust particles varies. A few dust formulations are concentrates and contain a high percentage of active ingredients. In some embodiments, these are mixed with dry inert excipients before applying. In some embodiments, dusts are used dry and can easily drift to non-target sites. [0120] In some instances, the composition is formulated as a powder.
- the composition is formulated as a wettable powder.
- Wettable powders are dry, finely ground formulations that look like dusts. They usually must be mixed with water for application as a spray. A few products, however, can be applied either as a dust or as a wettable powder—the choice is left to the applicator. Wettable powders have about 1 to about 95 percent active ingredient by weight; in some cases, more than about 50 percent. The particles do not dissolve in water. They settle out quickly unless constantly agitated to keep them suspended. They can be used for most pest problems and in most types of spray equipment where agitation is possible. Wettable powders have excellent residual activity.
- the composition is formulated as a soluble powder.
- Soluble powder formulations look like wettable powders. However, when mixed with water, soluble powders dissolve readily and form a true solution. After they are mixed thoroughly, no additional agitation is necessary.
- the amount of active ingredient in soluble powders ranges from about 15 to about 95 percent by weight; in some cases, more than about 50 percent. Soluble powders have all the advantages of wettable powders and none of the disadvantages, except the inhalation hazard during mixing.
- the composition is formulated as a water-dispersible granule.
- Water- dispersible granules also known as dry flowables, are like wettable powders, except instead of being dust-like, they are formulated as small, easily measured granules.
- Water-dispersible granules must be mixed with water to be applied. Once in water, the granules break apart into fine particles similar to wettable powders. The formulation requires constant agitation to keep it suspended in water. The percentage of active ingredient is high, often as much as 90 percent by weight. Water-dispersible granules share many of the same advantages and disadvantages of wettable powders, except they are more easily measured and mixed.
- the composition comprises a solid excipient.
- Solid excipients include finely-divided powder or granules of clay (e.g.
- kaolin clay diatomaceous earth, bentonite, Fubasami clay, acid clay, etc.
- synthetic hydrated silicon oxide talc, ceramics, other inorganic minerals (e.g., sericite, quartz, sulfur, activated carbon, calcium carbonate, hydrated silica, etc.), a substance which can be sublimated and is in the solid form at room temperature (e.g., 2,4,6-triisopropyl-1,3,5-trioxane, naphthalene, p-dichlorobenzene, camphor, adamantan, etc.); wool; silk; cotton; hemp; pulp; synthetic resins (e.g., polyethylene resins such as low-density polyethylene, straight low-density polyethylene and high-density polyethylene; ethylene-vinyl ester copolymers such as ethylene-vinyl acetate copolymers; ethylene-methacrylic acid ester copolymers such as ethylene-methyl methacrylate copo
- the composition is provided in a microencapsulated formulation (e.g., a nanocapsule). Microencapsulated formulations are mixed with water and sprayed in the same manner as other sprayable formulations. After spraying, the encapsulation shell or coating breaks down and slowly releases the active ingredient. [0125] In some instances, the composition is provided in a liposome. In some instances, the composition is provided in a vesicle. [0126] In some instances, a composition provided herein includes a surfactant. Surfactants, also called wetting agents and spreaders, physically alter the surface tension of a spray droplet.
- surfactants also called wetting agents and spreaders
- Surfactants For a formulation to perform its function properly, a spray droplet must be able to wet the foliage and spread out evenly over a leaf. Surfactants enlarge the area of formulation coverage, thereby increasing exposure to the active agent. Surfactants are particularly important when applying a formulation to waxy or hairy leaves. Without proper wetting and spreading, spray droplets often run off or fail to cover leaf surfaces adequately. Too much surfactant, however, can cause excessive runoff and reduce efficacy. [0127] Surfactants can be classified as anionic, cationic, or nonionic. Formulation activity in the presence of a nonionic surfactant can be quite different from activity in the presence of a cationic or anionic surfactant.
- anionic surfactants are most effective when used with contact pesticides (pesticides that control a pest by direct contact rather than being absorbed systemically).
- Cationic surfactants are not typically used as stand-alone surfactants because they usually are phytotoxic.
- Nonionic surfactants often used with systemic pesticides, help sprays to penetrate plant cuticles.
- Nonionic surfactants are compatible with most pesticides, and most EPA-registered pesticides that require a surfactant recommend a nonionic type.
- Adjuvants include, but are not limited to, stickers, extenders, plant penetrants, compatibility agents, buffers or pH modifiers, drift control additives, defoaming agents, and thickeners.
- surfactants included in the compositions described herein are alkyl sulfate ester salts, alkyl sulfonates, alkyl aryl sulfonates, alkyl aryl ethers and polyoxyethylenated products thereof, polyethylene glycol ethers, polyvalent alcohol esters and sugar alcohol derivatives.
- the surfactant is a nonionic surfactant, a surfactant plus nitrogen source, an organo- silicone surfactant, or a high surfactant oil concentrate.
- the recombinant polynucleotide can, in embodiments, be in a mixture with other active compounds, such as pesticidal agents (e.g., insecticides, sterilants, acaricides, nematicides, molluscicides, or fungicides, attractants, growth-regulating substances, or herbicides).
- pesticidal agents e.g., insecticides, sterilants, acaricides, nematicides, molluscicides, or fungicides, attractants, growth-regulating substances, or herbicides.
- pesticide refers to any substance or mixture of substances intended for preventing, destroying, repelling, or mitigating any pest.
- a pesticide can be a chemical substance or biological agent used against pests including insects, mollusks, pathogens, weeds, nematodes, and microbes that compete with humans for food, destroy property, spread disease, or are a nuisance.
- the term “pesticidal agent” further encompasses other bioactive molecules such as antibiotics, antivirals pesticides, antifungals, antihelminthics, nutrients, pollen, sucrose, and/or agents that stun or slow insect movement.
- this disclosure is related to a method of producing a modified plant propagule that comprises at least one plant cell comprising a recombinant RNA molecule.
- the method includes the steps of: isolating a plant propagule comprising at least one plant cell comprising a recombinant RNA molecule and an amalgavirus RNA-dependent RNA polymerase (RdRP) from a mixed population of plant cells comprising both plant cells comprising the recombinant RNA molecule and plant cells lacking the recombinant RNA molecule, wherein the recombinant RNA molecule comprises, in 5’ to 3’ order, a 5’ RNA replication element that is capable of being recognized by the amalgavirus RdRP; a cargo RNA sequence; and a 3’ RNA replication element that is capable of being recognized by the amalgavirus RdRP.
- RdRP amalgavirus RNA-dependent RNA polymerase
- the isolated plant propagule comprising at least one plant cell comprising a recombinant RNA molecule will be free or substantially free of plant cells lacking the recombinant RNA.
- Such isolated plant propagules which are substantially free of plant cells lacking the recombinant RNA can in certain embodiments comprise plant propagules where at least 75%, 80%, 85%, 90%, 95%, 98%, or 99% of the plant cells in the plant propagule contain the recombinant DNA molecule.
- the mixed population of plant cells comprise a population of protoplasts or a population of cells in callus, an explant, a plant part, or whole plant.
- the mixed population of plant cells can comprise a population of plant cells where less than 70%, 60%, 50%, 40%, 30%, 20%, 10%, 5%, or 1% of the plant cells in the population contain the recombinant RNA molecule.
- the mixed population of plant cells comprise plant cells comprising the amalgavirus RdRP and plant cells lacking the amalgavirus RdRP.
- the plant cells lacking the amalgavirus RdRP will also lack the recombinant RNA.
- the mixed population of plant cells comprise plant cells comprising the amalgavirus RdRP.
- the plant cells comprising the amalgavirus RdRP can further comprise the recombinant RNA.
- the cargo RNA sequence comprises RNA that encodes a selectable or scorable marker.
- the mixed population of plant cells is screened or selected for the presence of the plant cell comprising the recombinant RNA molecule prior to isolating the plant propagule. In such screens, the mixed population of cells or a portion thereof are subjected to an assay for a screenable marker for the presence of the recombinant RNA molecule (e.g., an RNA sequence diagnostic for presence of the recombinant RNA molecule or a polypeptide encoded by the recombinant RNA molecule) and separated from plant cells lacking the recombinant RNA molecule.
- an assay for a screenable marker for the presence of the recombinant RNA molecule e.g., an RNA sequence diagnostic for presence of the recombinant RNA molecule or a polypeptide encoded by the recombinant RNA molecule
- the isolation comprises Agent Ref: P14357WO00 - 48 - selecting for the plant cell comprising the recombinant RNA molecule prior to isolating the plant propagule.
- Selections in instances where the recombinant RNA encodes a selectable marker can comprise exposing the mixed population of plant cells to a selection agent (e.g., an herbicide or antibiotic) and isolating plant cells which survive exposure to the selection agent.
- selectable marker/selection agent combinations include glyphosate-resistant EPSPS enzymes and/or glyphosate oxidases/glyphosate, a bialaphos resistance (bar) or phosphinothricin acyl transferase (pat) enzyme/glufosinate, or a neomycin phosphotransferase (npt)/neomycin or kanamycin.
- the selectable or scorable marker is an RNA aptamer (e.g., a Broccoli aptamer) or a regulatory RNA (e.g., an siRNA, siRNA precursor, miRNA, or miRNA precursor, or a phased siRNA or phased siRNA precursor that downregulates expression of an endogenous gene in the plant, resulting in a detectable phenotype, e.g., bleaching caused by downregulation of a pigment-producing gene).
- the mixed population is located within a plant or a plant part.
- the plant or plant part is screened or selected for presence of the recombinant RNA molecule prior to isolating the plant propagule.
- the plant or plant part is screened or selected for systemic presence of the recombinant RNA molecule prior to isolating the plant propagule.
- the plant cells, plant, or plant part in the mixed population or that are isolated lack DNA that encodes the recombinant RNA molecule.
- the plant propagule comprising the recombinant RNA molecule is isolated by detecting the RNA molecule in one or more plant cells comprising the recombinant RNA molecule and separating the one or more plant cells comprising the recombinant RNA molecule from the plant cells lacking the recombinant DNA molecule.
- the plant propagule is a mosaic comprising both plant cells comprising the recombinant RNA molecule and plant cells lacking the recombinant RNA molecule.
- the modified plant propagule is a mosaic comprising both plant cells comprising the amalgavirus RdRP and plant cells lacking the amalgavirus RdRP.
- at least 99%, 98%, 95%, 90%, 85%, 80%, 70%, 60%, 50%, 40%, 30%, 20%, 10%, 5%, or 1% of the plant cells in the mosaic can comprise the recombinant RNA molecule.
- the plant propagule lacks DNA that encodes the recombinant RNA molecule.
- the modified plant propagule comprises the cell comprising the recombinant RNA molecule, or a seed, seedling, ovule, embryo, pollen, root, stem, leaf, shoot, tuber, rhizome, stolon, bulb, explant, or callus comprising the cell comprising the recombinant RNA molecule.
- Plant propagules made by any of the aforementioned methods and/or incorporating any of the aforementioned features are also provided herein.
- any of the aforementioned methods can further comprise multiplying the cell, seed, seedling, ovule, embryo, pollen, root, stem, leaf, shoot, tuber, rhizome, stolon, bulb, explant, or callus to obtain progeny, wherein the progeny comprise the recombinant RNA molecule.
- the multiplying of the cells consists of culturing a plurality of explants obtained from Agent Ref: P14357WO00 - 49 - the cell, seed, seedling, ovule, embryo, pollen, root, stem, leaf, shoot, tuber, rhizome, stolon, bulb, explant, or callus.
- the isolated propagule comprises the cell and the aforementioned methods can further comprise regenerating a plant, seedling, ovule, embryo, pollen, root, stem, leaf, shoot, tuber, rhizome, stolon, bulb, explant, or callus comprising the recombinant RNA from the cell.
- the isolated propagule comprises callus and the aforementioned methods can further comprise regenerating a plant, seedling, ovule, embryo, pollen, root, stem, leaf, shoot, tuber, rhizome, stolon, bulb, or explant comprising the recombinant RNA from said callus.
- a plant is regenerated and the aforementioned methods can further comprise recovering F1 seed or F1 progeny or clonal progeny comprising the recombinant RNA from the plant.
- this disclosure is related to a method of providing a synthetic amalgavirus satellite RNA to a plant or plant part by grafting one plant part to another plant part.
- the methods can comprise grafting a scion onto a rootstock comprising any of the aforementioned or otherwise disclosed recombinant DNA molecules and/or recombinant RNA molecules (e.g., a recombinant RNA comprising, in 5’ to 3’ order, a 5’ RNA replication element that is capable of being recognized by an amalgavirus RNA-dependent RNA polymerase (RdRP); a cargo RNA sequence; and a 3’ RNA replication element that is capable of being recognized by the amalgavirus RdRP), wherein at least one cell of the rootstock and/or the scion comprises the amalgavirus RdRP.
- a recombinant RNA comprising, in 5’ to 3’ order, a 5’ RNA replication element that is capable of being recognized by an amalgavirus RNA-dependent RNA polymerase (RdRP); a cargo RNA sequence; and a 3’ RNA replication element that is capable of being recognized by the amalgavirus
- the scion can comprise a plant shoot, an apical or other meristem, a leaf attached to a petiole, or other plant part and the rootstock can comprise roots and aerial portions of the plant including the main stem, secondary stems, leaves, and/or reproductive structures of the plant,
- DNA that encodes the recombinant RNA molecule is absent in the scion and/or the rootstock.
- the scion lacks the recombinant RNA molecule prior to grafting.
- the rootstock comprises the amalgavirus RdRP.
- the amalgavirus RdRP is provided by an amalgavirus endemic to the rootstock (e.g., an amalgavirus which is non-pathogenic and/or commensal).
- the amalgavirus RdRP is exogenously provided to the rootstock (e.g., via a DNA expression cassette which is integrated into the chromosomal or plastidic DNA of the rootstock or via a recombinant viral vector comprising DNA or RNA encoding the RdRP).
- the scion comprises the amalgavirus RdRP.
- the RdRP is provided by an amalgavirus endemic to the scion (e.g., an amalgavirus which is non-pathogenic and/or commensal).
- the RdRP is exogenously provided to the scion (e.g., via a DNA expression cassette which is integrated into the chromosomal or plastidic DNA of the scion or via a recombinant viral vector comprising DNA or RNA encoding the RdRP).
- the amalgavirus RdRP comprises a protein having at least 85%, 90%, 95%, 97%, or 99% sequence identity to SEQ ID NO: 476, 477, 478, 730 to 813, or 814.
- the rootstock and/or the scion comprises a heterologous viral coat protein which can encapsidate the recombinant RNA molecule. In some embodiments, the rootstock and/or the scion comprises the recombinant RNA molecule encapsidated by a heterologous viral coat protein.
- this disclosure is related to a method of producing a grafted plant comprising a recombinant RNA molecule comprising, in 5’ to 3’ order, a 5’ RNA replication element that is capable of being recognized by an amalgavirus RNA-dependent RNA polymerase (RdRP); a cargo RNA sequence; and a 3’ RNA replication element that is capable of being recognized by an the amalgavirus RdRP.
- RdRP amalgavirus RNA-dependent RNA polymerase
- the recombinant RNA molecule is provided by contacting the scion, the rootstock, or both the scion and the rootstock with a composition comprising the recombinant RNA molecule prior to grafting the scion onto the rootstock to produce the grafted plant.
- at least one cell of the rootstock and/or the scion comprises an amalgavirus RdRP prior to contacting the scion, the rootstock, or both the scion and the rootstock with the composition.
- the rootstock comprises the amalgavirus RdRP.
- the amalgavirus RdRP is provided by an amalgavirus endemic to the rootstock (e.g., an amalgavirus which is non- pathogenic and/or commensal).
- the amalgavirus RdRP is exogenously provided to the rootstock (e.g., via a DNA expression cassette which is integrated into the chromosomal or plastidic DNA of the rootstock or via a recombinant viral vector comprising DNA or RNA encoding the RdRP).
- the scion comprises the amalgavirus RdRP.
- the RdRP is provided by an amalgavirus endemic to the scion (e.g., an amalgavirus which is non-pathogenic and/or commensal).
- the RdRP is exogenously provided to the scion (e.g., via a DNA expression cassette which is integrated into the chromosomal or plastidic DNA of the scion or via a recombinant viral vector comprising DNA or RNA encoding the RdRP).
- the amalgavirus RdRP comprises a protein having at least 85%, 90%, 95%, 97%, or 99% sequence identity to SEQ ID NO: 476, 477, 478, 730 to 813, or 814.
- DNA that encodes the recombinant RNA molecule is absent in the scion, the rootstock, and/or the grafted plant.
- the composition can be provided to the scion, the rootstock, or both the scion and the rootstock according to any of the formulations disclosed herein.
- the formulation is a liquid, a gel, or a powder.
- the formulation is configured to be sprayed on to the scion, the rootstock, or both the scion and the rootstock; to be injected into the scion, the rootstock, or both the scion and the rootstock; to be soaked into the scion, the rootstock, or both the scion and the rootstock; or to be coated onto the scion, the rootstock, or both the scion and the rootstock.
- the contacting comprises dipping the scion, the rootstock, or both the scion and the rootstock into the composition prior to grafting.
- this disclosure is related to a method for producing a plant that transmits any of the aforementioned or otherwise disclosed recombinant RNA molecules provided herein to progeny plants or seed.
- the methods include the steps of: isolating an F 1 progeny plant or seed comprising at least one cell comprising an amalgavirus RNA-dependent RNA polymerase (RdRP) and the recombinant RNA molecule comprising, in 5’ to 3’ order, a 5’ RNA replication element that is capable of being recognized by the amalgavirus RdRP); a cargo RNA sequence; and a 3’ RNA replication element that is capable of being recognized by the amalgavirus RdRP from a population of F 1 plants or seed obtained from at least one parent plant comprising the recombinant RNA molecule.
- RdRP amalgavirus RNA-dependent RNA polymerase
- the parent plant or a part thereof comprising the plant cell is screened or selected for presence of the recombinant RNA molecule prior to isolating the F 1 progeny plant or seed.
- the parent plant or one or more parts thereof are screened for systemic presence of the recombinant RNA molecule prior to isolating the F 1 progeny plants.
- floral tissue e.g., whole flowers or buds, sepal, calyx, or petal
- male reproductive tissue e.g., stamen, anther, or pollen
- female reproductive tissue e.g., whole fruit, ovary, pericarp, ovule, seed coat, endosperm, or embryo
- F 1 seeds are obtained from a parent plant selected for presence of the recombinant RNA molecule in pericarp tissue.
- the F 1 progeny plant or seed comprising the cell is isolated by screening the population of F 1 plants or seed obtained from a parent plant for the presence of the recombinant RNA molecule and propagating the F 1 progeny plant or seed comprising the recombinant RNA molecule.
- the progeny plants or seed thereof are subjected to an assay for a screenable marker for the presence of the recombinant RNA (e.g., an RNA sequence diagnostic for presence of the recombinant RNA molecule or a polypeptide encoded by the recombinant RNA molecule) and separated from progeny plants and seed lacking the recombinant RNA progeny plants and seed lacking the recombinant RNA.
- a screenable marker for the presence of the recombinant RNA e.g., an RNA sequence diagnostic for presence of the recombinant RNA molecule or a polypeptide encoded by the recombinant RNA molecule
- Such screening assays can be non-destructive assays wherein a portion of the progeny seed or plant is removed and assayed without loss of the viability or ability to propagate the seed or plant tissue comprising the recombinant RNA.
- an F 1 seed of the parent plant is non-destructively screened for presence of the recombinant RNA molecule.
- the F 1 seed of the parent plant is non-destructively screened by assaying maternally derived or endosperm tissue of the seed for the presence of the recombinant RNA molecule.
- the cargo RNA sequence comprises RNA that encodes a selectable or scorable marker.
- the recombinant RNA molecule encodes a selectable marker and the F 1 progeny plant or seed comprising the recombinant RNA molecule is isolated by selecting the F 1 progeny plant or seed comprising the recombinant RNA molecule for presence of the selectable marker.
- Examples of such selections in instances where the recombinant RNA encodes a selectable marker can comprise exposing the progeny seeds or plants to a selection agent (e.g., an herbicide or antibiotic) and isolating progeny seeds or plants which survive exposure to the selection agent.
- a selectable marker e.g., a protein which confers resistance to a selection agent such as an herbicide or antibiotic
- selectable marker/selection agent combinations include glyphosate-resistant EPSPS enzymes and/or glyphosate oxidases/glyphosate, a bialaphos resistance (bar) or phosphinothricin acyl transferase (pat) enzymes/glufosinate, and neomycin phosphotransferase (npt)/neomycin or kanamycin.
- the selectable or scorable marker is an RNA aptamer or a regulatory RNA.
- the F 1 progeny plant or seed lacks DNA that encodes the recombinant RNA molecule.
- the parent plant lacks DNA that encodes the recombinant RNA Agent Ref: P14357WO00 - 52 - molecule.
- the selected F 1 progeny plant transmits the recombinant RNA molecule to at least F 2 progeny.
- the F 1 progeny plant or seed population is obtained from a parent plant used as a pollen recipient.
- the F 1 progeny plant or seed population is obtained from a parent plant used as a pollen donor.
- the F 1 progeny plant or seed population is obtained by selfing the parent plant.
- the F 1 progeny plant or seed population is obtained from the sexual crossing of two parent plants.
- the parent plant that comprises the recombinant RNA molecule is the female parent plant. In other embodiments, the parent plant that comprises the recombinant RNA molecule is the male parent plant, and the recombinant RNA molecule is transmitted in pollen of the male parent plant. [0137] In certain embodiments, the methods can further comprise introducing the recombinant RNA molecule or a polynucleotide encoding the recombinant RNA molecule into a plant cell and obtaining the parent plant comprising the recombinant RNA molecule from the plant cell.
- the recombinant RNA molecule further comprises at least one additional element selected from the group consisting of: (a) at least one RNA encoding a viral movement protein (MP); (b) at least one tRNA-like sequence; and c) an origin-of-assembly sequence (OAS).
- a parent and/or plant comprises a heterologous viral coat protein which can encapsidate the recombinant RNA molecule.
- the parent and/or progeny plant comprises the recombinant RNA molecule encapsidated by a heterologous viral coat protein.
- this disclosure is related to a method of barcoding a plant, plant cell, progeny thereof, or part thereof.
- the methods comprise providing to the plant or plant cell any of the aforementioned or otherwise disclosed recombinant RNA molecules provided herein, wherein the cargo RNA of the recombinant RNA molecule comprises a barcode RNA molecule, and wherein the plant or plant cell comprises an amalgavirus RdRP.
- the barcode RNA molecule comprises a sequence that uniquely identifies the plant, plant cell, progeny thereof, or part thereof.
- the barcode RNA can be a randomly generated sequence.
- the barcode RNA molecule comprises a sequence that is not present in the genome and/or transcriptome of a wild-type plant of the same species, a pathogen thereof, or a symbiont thereof. In some embodiments, the barcode RNA molecule comprises a forward primer binding site and a reverse primer binding site for detection of the barcode RNA molecule. In embodiments, the barcode RNA molecule comprises a non- protein coding sequence. In some embodiments, the barcode RNA sequence is up to about 3.2 kb in length. In some embodiments, the barcode RNA has a length of 10 to 5000 nucleotides, 20 to 1000 nucleotides, or 50 to 500 nucleotides.
- the barcode RNA molecule has a length of 10, 20, 30, 40, 50, 60, 70, 80, 90, 100, 110, 120, 130, 140, 150, 160, 170, 180, 190, 200, 250, 300, 350, 400, 450, 500, 550, 600, 650, 700, 750, 800, 850, 900, 950, 1000, 1500, 2000, 2500, 3000, 3500, 4000, 4500, or 5000 nucleotides.
- the plant transmits the recombinant RNA molecule comprising the barcode RNA to progeny.
- the plant, plant cell, progeny thereof, or part thereof lacks DNA that encodes the recombinant RNA molecule.
- the methods Agent Ref: P14357WO00 - 53 - can further comprise isolating an F 1 progeny plant or seed comprising at least one cell comprising the amalgavirus RdRP and the recombinant RNA molecule.
- the F 1 progeny plant or seed is obtained from the plant used as a pollen recipient.
- the F 1 progeny plant or seed is obtained from the plant used as a pollen donor.
- the F 1 progeny plant or seed is obtained by selfing the parent plant.
- the methods can further comprise propagating the plant or plant cell to obtain a plant part or a plant propagule comprising the barcode RNA molecule.
- this disclosure is related to a method of identifying a barcoded plant, plant part, or plant cell.
- the methods comprise screening for the presence of a barcode RNA molecule in the plant, plant part, or plant cell, wherein the plant, plant part, or plant cell comprises any of the aforementioned or otherwise disclosed recombinant RNA molecules provided herein, and wherein the cargo RNA of the recombinant RNA molecule comprises the barcode RNA molecule.
- the methods comprise obtaining a nucleic acid sample from the plant, plant part, or plant cell; and detecting the presence of the barcode RNA molecule in the sample.
- Assays for detection of a barcode RNA include RNA detection assays (e.g., an RT-PCR assay) using nucleic acid probes and/or primers which can detect the barcode RNA and/or sequencing of the barcode RNA.
- Such screening assays can be non-destructive assays wherein a portion of the seed or plant is removed and assayed without loss of the viability or ability to propagate the seed or plant tissue comprising the barcode RNA.
- Methods for non-destructive assays of seed or other plant tissue which can be adapted for such screens include but are not limited to those disclosed in US patent applications US20220221377 and US20210259176, both incorporated herein by reference in their entireties.
- a seed of the plant is non-destructively screened for presence of the barcode RNA molecule.
- a seedling, ovule, embryo, pollen, root, stem, leaf, shoot, tuber, rhizome, stolon, bulb, explant, or callus is screened for the presence of the barcode RNA molecule.
- the methods disclosed herein are not processes for modifying the germ line or genetic identity of human beings.
- the methods disclosed herein are not processes for modifying the genetic identity of animals which are likely to cause them suffering without any substantial medical benefit to man or animal, and are also not drawn to animals resulting from such processes.
- the methods disclosed herein are not methods for treatment of the human or animal body by surgery or therapy.
- the cells disclosed herein are not human embryos.
- the cells disclosed herein are not the human body or its parts, at the various stages of its formation and development.
- the plant cells, plant propagules e.g., a seed, seedling, ovule, embryo, pollen, root, stem, leaf, shoot, tuber, rhizome, stolon, bulb, explant, or callus
- plants provided herein are not produced by an exclusively biological process.
- the methods for producing plant cells, plant propagules e.g., a seed, seedling, ovule, embryo, pollen, root, stem, leaf, shoot, tuber, rhizome, stolon, bulb, explant, or callus
- plants provided herein are not exclusively biological processes.
- Various embodiments of the compositions, systems, and methods described herein are set forth in the following set of numbered embodiments. [0142] 1.
- a recombinant RNA molecule comprising from 5’ terminus to 3’ terminus: (a) a 5’ RNA replication element recognized by an amalgavirus RNA-dependent RNA polymerase (RdRP); (b) a cargo RNA molecule; and (c) a 3’ RNA replication element recognized by the RdRP; wherein the 5’ RNA replication element, the cargo RNA molecule, and the 3’ RNA replication element are operably linked and wherein the cargo RNA molecule is heterologous to the 5’ RNA replication element and the 3’ RNA replication element, optionally wherein: (i) the 5’ RNA replication element and the 3’ RNA replication element are obtained from the same amalgavirus genome or from amalgavirus genomes having at least 85%, 90%, 95%, 98%, or 99% sequence identity to one another and which are optionally related; (ii) the 5’ RNA replication element, the 3’ RNA replication element, and the RdRP are obtained from the same amalgavirus genome or
- RNA replication element comprises at least one RNA secondary structure provided in Table 1 or adopted by an RNA molecule encoded by SEQ ID NO: 467, 469, 471, or 572 to 649; and/or (b) the 3’ RNA replication element comprises at least one RNA secondary structure provided in Table 1 or adopted by an RNA molecule encoded by SEQ ID NO: 468, 470, 472, or 650 to 729.
- the 5’ RNA replication element comprises at least one RNA secondary structure provided in Table 1 or adopted by an RNA molecule encoded by SEQ ID NO: 467, 469, 471, or 572 to 649; and/or (b) the 3’ RNA replication element comprises at least one RNA secondary structure provided in Table 1 or adopted by an RNA molecule encoded by SEQ ID NO: 468, 470, 472, or 650 to 729.
- RNA replication element comprises an RNA molecule encoded by SEQ ID NO: 467, 469, 471, or 572 to 649; a variant thereof encoded by a DNA molecule having at least 70%, 75%, 80%, 85%, 90%, 95%, 98%, or 99% sequence identity to SEQ ID NO: 467, 469, 471, or 572 to 649; or a variant thereof wherein one or more nucleotides in the RNA secondary structure are substituted with distinct nucleotides which maintain the RNA secondary structure; and/or (b) the 3’ RNA replication element comprises an RNA molecule encoded by SEQ ID NO: 468, 470, 472, or 650 to 729, or; a variant thereof encoded by a DNA molecule having at least 70%, 75%, 80%, 85%, 90%, 95%, 98%, or 99% sequence identity to SEQ ID NO: 468, 470, 472, or 650 to
- Agent Ref P14357WO00 - 55 - [0145] 4.
- the 5’ RNA replication element comprises at least a segment of the 5’ untranslated region (UTR) of the amalgavirus genome or a variant thereof having one or more nucleotide substitutions, insertions, and/or deletions, wherein the variant is recognized by the RdRP, optionally wherein the 5’ RNA replication element further comprises a genomic sequence of the amalgavirus that is natively located 3’ to and adjacent to the 5’ UTR sequence; and/or (b) the 3’ RNA replication element comprises at least a segment of the 3’ UTR of the amalgavirus genome or a variant thereof having one or more nucleotide substitutions, insertions, and/or deletions, wherein the variant is recognized by the RdRP, optionally wherein the 3’ RNA replication element further comprises a genomic sequence of the amalgavirus that is natively located 5’ to and adjacent to the 3’ UTR sequence, and optional
- RNA molecule of any one of embodiments 1 to 5, wherein the RNA molecule further comprises at least one of: (i) a tRNA-like element, optionally wherein the tRNA-like element provides for intercellular movement of the RNA and optionally wherein the intercellular movement is mediated by a viral movement protein (MP); (ii) an encapsidation recognition element (ERE), optionally wherein the ERE provides for encapsidation of the RNA by a heterologous viral capsid protein; (iii) an RNA effecter; and/or (iv) RNA encoding a viral movement protein (MP), optionally wherein an internal ribosome entry site (IRES) is operably linked to the RNA encoding the MP.
- MP viral movement protein
- ERE encapsidation recognition element
- RNA-like element comprises a tRNA-like molecule from an Arabidopsis FT mRNA or is a tRNA-like sequence encoded by a DNA sequence selected from the group consisting of SEQ ID NOs: 76-123, and 466 or is a modified tRNA-like sequence that has at least 90% sequence identity to a scaffold tRNA-like sequence encoded by a DNA sequence selected from the group consisting of SEQ ID NOs: 76-123, and 466 and that maintains the secondary structure of the scaffold tRNA-like sequence, and/or wherein the ERE is a tobacco mosaic virus (TMV) OAS.
- TMV tobacco mosaic virus
- the recombinant RNA molecule of embodiment 1, wherein the cargo RNA molecule is up to about 3.2kb in length.
- 9 The recombinant RNA molecule of embodiment 1, wherein the cargo RNA molecule comprises: (a) at least one coding sequence, optionally wherein the coding sequence encodes a selectable or scoreable marker; (b) at least one non-coding sequence; or (c) both at least one coding sequence and at least one non-coding sequence.
- Agent Ref P14357WO00 - 56 - [0151] 10.
- RNA molecule of embodiment 1 wherein the cargo RNA molecule comprises at least one coding sequence, and wherein the RNA molecule further comprises an internal ribosome entry site (IRES) which is operably linked to at least one coding sequence, optionally wherein the operably linked IRES is located 5’ and immediately adjacent to the coding sequence.
- IRES internal ribosome entry site
- hpRNA hairpin RNA
- RNA hairpin RNA
- siRNA small interfering RNA
- miRNA microRNA
- miRNA miRNA
- a ribozyme a ligand-responsive ribozyme
- RNA aptamer or a long noncoding RNA (
- RNA binding proteins comprise an RNA recognition motif.
- RNA molecule comprises: at least one heterologous RNA virus (HRV) amplicon in sense or antisense orientation to the first 5’ RNA replication element comprising: I. (i) a heterologous RNA virus (HRV) 5’ replication region (HRV 5’RR); (ii) the cargo RNA molecule; and (iii) the heterologous RNA virus (HRV) 3’ RNA replication region (HRV 3’RR); wherein the HRV 5’ RR and HRV 3’ RR HRV are recognized by a heterologous RNA virus RNA-dependent RNA polymerase (hrvRdRP); and wherein the HRV 5’RR, cargo RNA molecule, and HRV 3’RR are operably linked; or II.
- HRV heterologous RNA virus
- heterologous RNA virus (HRV) subgenomic promoter operably linked to the cargo RNA molecule; wherein the subgenomic promoter is recognized by a heterologous RNA virus RNA-dependent RNA polymerase (hrvRdRP).
- RNA molecule comprises from 5’ terminus to 3’ terminus: (a) the 5’ RNA replication element; Agent Ref: P14357WO00 - 57 - (b) the HRV amplicon in antisense orientation to the first 5’ RNA replication element; optionally wherein the HRV amplicon further comprises: (i) an RNA molecule encoding an HRV RNA-dependent RNA polymerase (hrvRdRP) which is operably linked to the HRV 5’RR and HRV 3’RR, wherein the RNA molecule encoding the HRV RNA- dependent RNA polymerase (hrvRdRP) is optionally operably linked to a subgenomic promoter recognized by the hrvRdRP; or (ii) an RNA molecule encoding an HRV RNA-dependent RNA polymerase (hrvRdRP) which is operably linked to linked to a subgenomic promoter recognized by the hrvRdRP; or (ii) an RNA
- a virus selected from the group consisting of an Alphaflexivirus, Betaflexivirus, Bromovirus, a Closterovirus, a Comovirus, a Potexvirus, Potyvirus, Tobamovirus, Tombusvirus, Tospoviridae, Trivirinae, Tymovirus, and Secoviridae.
- the Bromovirus is a Cucumber Mosaic Virus, Spinach latent virus, Olive latent virus, or Brome mosaic virus
- the Closterovirus is a Citrus tristeza virus or Beet yellows virus
- the Comovirus is Cowpea mosaic virus, Apple latent spherical virus, or Soybean latent spherical virus
- the Potexvirus is Potato virus X or Citrus yellow vein clearing virus
- the Potyvirus is a Pepper mottle virus, Bean yellow mosaic virus, Barley stripe mosaic virus, Wheat stripe mosaic virus, Rice yellow mottle virus, Maize dwarf mosaic virus, zucchini yellow mosaic virus, watermelon mosaic virus, or sugarcane mosaic virus
- the Tobamovirus is a Tobacco mosaic virus, Tomato mosaic virus, Tomato brown rugose fruit virus, Turnip vein-clearing virus, or Pepper mild mottle virus
- the Tombus is a Tobacco mosaic virus, Tomato mosaic virus, Tomato brown rugose fruit virus, Turnip vein-clearing
- the HRV 5’ RR comprises at least one RNA secondary structure adopted by an RNA encoded by SEQ ID NO: 161 to 186 or 909 to 918, or comprises the RNA sequence encoded by SEQ ID NO: 161 to 186 or 909 to 918, or comprises a contiguous fragment of at least 80%, 85%, 90%, or 95% of the full sequence of the RNA encoded by SEQ ID NO: 161 to 186 or 909 to 918; and/or (b) the HRV 3’ RR comprises at least one RNA secondary structure adopted by an RNA encoded by SEQ ID NO: 187 to 211 or 919 to 928, or comprises the RNA sequence of the RNA encoded Agent Ref: P14357WO00 - 58 - by SEQ ID NO: 187 to 211 or 919 to 928, or comprises a contiguous fragment of at least 80%, 85%, 90%, or 95% of
- the HRV 3’ RR comprises an RNA molecule containing at least a segment of the 3’ untranslated region (UTR) of the HRV genome, or a variant thereof having one or more nucleotide substitutions, insertions, and/or deletions, wherein the HRV 3’ RR or the variant is recognized by the hrvRdRP, optionally wherein the RNA comprising the HRV 3’ RR further comprises a genomic sequence of the HRV that is natively located 5’ to and adjacent to the 3’ UTR sequence; and/or (b) the HRV 5’ RR comprises an RNA molecule containing at least a segment of the 3’ untranslated region (UTR) of the HRV genome, or a variant thereof having one or more nucleotide substitutions, insertions, and/or deletions, wherein the HRV 5’ RR or the variant is recognized by the hrvRdRP, optionally
- RNA comprises an HRV-inhibitory RNA or encodes an HRV-inhibitory protein, wherein the HRV- inhibitory RNA or HRV-inhibitory protein inhibits infection, movement, transmission, and/or replication of the HRV.
- the cargo RNA comprises an RNA having at least 20 contiguous nucleotides having an identical or complementary sequence to a segment of equivalent length of the genomic RNA of the HRV.
- RNA comprises an RNA having at least 20 contiguous nucleotides having an identical or complementary sequence to a segment of equivalent length of the genomic RNA of the HRV which does not encode the hrvRdRP.
- RNA encoded by the Cucumber Mosaic Virus and the HRV 5’RR comprises an RNA encoded by the Cucumber Mosaic Virus 5’ RR DNA sequence in Table 7, an RNA encoded by a DNA having at least 85%, 90%, 95%, or 99% sequence identity to the Cucumber Mosaic Virus 5’ RR DNA sequence in Table 7,
- the HRV 3’ RR comprises an RNA encoded by the Cucumber Mosaic Virus 3’ RR DNA sequence in Table 7, or an RNA encoded by a DNA having at least 85%, 90%, 95%, or 99% sequence identity to the Cucumber Mosaic Virus 3’ RR DNA sequence in Table 7;
- the HRV is a Brome mosaic virus and the HRV 5’RR comprises an RNA encoded by the Brome mosaic virus 5’ RR DNA sequence in Table 7, an RNA encoded
- the cleavable sequence is optionally a self-cleaving ribozyme, a self-cleaving inducible ribozyme, or an siRNA or miRNA recognition site.
- [0170] 29 An agricultural formulation comprising the recombinant RNA molecule of any one of embodiments 1 to 28.
- 30 The agricultural formulation of embodiment 29, wherein the recombinant RNA molecule is complexed with one or more RNA binding proteins or encapsidated by a viral capsid protein, optionally wherein the RNA comprises an ERE which provides for encapsidation of the RNA by a heterologous capsid protein.
- 31. The agricultural formulation of embodiment 30, wherein the RNA binding proteins comprise an RNA recognition motif.
- 32 The agricultural formulation of embodiment 30 or 31, wherein the viral capsid protein is heterologous to the amalgavirus.
- 33 The agricultural formulation of embodiment 30 or 31, wherein the viral capsid protein is heterologous to the amalgavirus.
- a cell comprising the recombinant RNA molecule of any one of embodiments 1 to 28, wherein the cell is a bacterial cell, a fungal cell, a plant cell, an insect cell, or an invertebrate animal cell.
- the cell is a plant cell and DNA which encodes the recombinant RNA molecule is absent from the cell.
- 36 The cell of embodiment 34, wherein the cell is a plant cell and DNA which encodes the recombinant RNA molecule is absent from the cell.
- the cell of embodiment 34 wherein the cell comprises a recombinant DNA molecule which encodes the recombinant RNA molecule.
- An expression system comprising: (a) an RNA molecule comprising the recombinant RNA molecule of any one of embodiments 1 to 28; and (b) a cell containing the recombinant RNA molecule and an RdRP protein that recognizes the 5’ and 3’ RNA replication elements of the recombinant RNA molecule.
- RNA molecule comprises an operably linked encapsidation recognition element (ERE) recognized by a viral capsid protein, and wherein the cell contains the viral capsid protein.
- EEE operably linked encapsidation recognition element
- 39 The expression system of embodiment 37, wherein the RNA molecule is encapsidated by the viral capsid protein and/or wherein the viral capsid protein comprises the viral capsid protein of any one of SEQ ID NOs: 26 to 50.
- 40 The expression system of embodiment 37, 38, or 39, further comprising the reverse complement of the recombinant RNA molecule. [0182] 41.
- the cell further comprises: (i) a viral capsid protein (CP), (ii) an RNA-binding protein (RBP) that can bind to the RNA molecule, optionally wherein the RBP binds to an RNA effecter; (iii) an RNA cleavage agent that cleaves the RNA molecule; (iv) a second RNA-dependent RNA polymerase (RdRP) protein that recognizes an HRV 5’ or 3’ replication region and/or a subgenomic promoter in the RNA molecule (2 nd RdRP); (v) a viral movement protein (MP); (v) a heterologous RNA virus (HRV); or (vi) an hr
- CP viral capsid protein
- RBP RNA-binding protein
- the expression system of embodiment 46 wherein the plant cell contains an amalgavirus which expresses the RdRP protein that recognizes the 5’ RNA replication element and the 3’ RNA replication element and/or wherein the plant cell contains an HRV that expresses the 2 nd RdRP or hrvRdRP protein.
- 48 The expression system of embodiment 47, wherein the amalgavirus occurs naturally in the plant cell.
- 49 A method of providing a synthetic amalgavirus satellite RNA to a plant, comprising contacting the plant with the recombinant RNA molecule of any one of embodiments 1 to 28. [0191] 50.
- contacting comprises spraying, dusting, injecting, or soaking the plant or a part thereof with the recombinant RNA molecule or the formulation.
- 51. The method of embodiment 49 or 50, further comprising providing an hrvRdRP to the plant which recognizes the HRV 5’ or 3’ replication region and/or the subgenomic promoter, optionally wherein the hrvRdRP is provided by introducing a recombinant DNA or RNA encoding the hrvRdRP into the plant or a part thereof.
- a method of establishing a synthetic amalgavirus satellite RNA in a plant cell comprising: providing to a plant cell the recombinant RNA molecule of any one of embodiments 1 to 28; wherein the plant cell comprises an RdRP protein that recognizes the 5’ RNA replication element and 3’ RNA replication element, wherein the RNA molecule is optionally comprises an ERE and is encapsidated by a capsid protein, whereby the RdRP protein catalyzes synthesis of the synthetic amalgavirus satellite RNA from the recombinant RNA molecule. [0194] 53.
- the method of embodiment 52 wherein the plant cell comprises an amalgavirus and wherein the RdRP protein is provided to the plant cell by the amalgavirus.
- 54 The method of embodiment 52 or 53, wherein the amalgavirus is endemic to the plant cell, optionally wherein the amalgavirus which is endemic to the plant cell is non-pathogenic and/or commensal.
- 55 The method of embodiment 52, 53, or 54, wherein the wherein the recombinant RNA molecule comprises an operably linked encapsidation recognition element (ERE) recognized by a viral capsid protein, optionally wherein capsid protein comprises a viral capsid protein of SEQ ID NO: 26 to 50.
- ERP operably linked encapsidation recognition element
- a method of obtaining a phenotypic change in a plant or plant cell comprising: providing to a plant or plant cell a recombinant RNA molecule of any one of embodiments 1 to 28, wherein the cargo RNA molecule comprises RNA that effects a phenotypic change in the plant or plant cell in comparison to a plant or plant cell lacking the recombinant RNA, wherein the plant or plant cell comprises an RdRP protein that recognizes the 5’ RNA replication element and 3’ RNA replication element and catalyzes synthesis of a synthetic amalgavirus RNA from the recombinant RNA molecule, and wherein the cargo RNA molecule effects the phenotypic change. [0199] 58.
- RNA that effects a phenotypic change in the plant or plant cell comprises at least one RNA selected from an siRNA or siRNA precursor, a miRNA or miRNA precursor, and a phased siRNA or phased siRNA precursor.
- RNA that effects a phenotypic change in the plant or plant cell comprises a messenger RNA.
- the messenger RNA comprises an RNA molecule absent in the genome of the plant or plant cell.
- RNA that effects a phenotypic change in the plant or plant cell comprises an RNA for modifying the genome of the plant or plant cell.
- RNA that effects a phenotypic change in the plant or plant cell comprises an RNA for modifying the transcriptome and/or epigenome of the plant or plant cell, optionally wherein the RNA for modifying the epigenome targets an endogenous plant gene for RNA-induced transcriptional silencing.
- RNA for modifying the epigenome targets an endogenous plant gene for RNA-induced transcriptional silencing.
- phenotypic change comprises an increase in the plant’s resistance to a pest or pathogen, optionally wherein the pest or pathogen is selected from the group comprising a bacterium, a virus other than an amalgavirus, a fungus, an oomycete, and an invertebrate.
- the pest or pathogen is selected from the group comprising a bacterium, a virus other than an amalgavirus, a fungus, an oomycete, and an invertebrate.
- the pathogen is a heterologous RNA virus (HRV), optionally wherein the HRV is a virus selected from the group consisting of an Alphaflexivirus, Betaflexivirus, Bromovirus, a Closterovirus, a Comovirus, a Potexvirus, Potyvirus, Tobamovirus, Tombusvirus, Tospoviridae, Trivirinae, Tymovirus, and Secoviridae.
- HRV heterologous RNA virus
- the Bromovirus is a Cucumber Mosaic Virus, Spinach latent virus, Olive latent virus, or Brome mosaic virus;
- the Closterovirus is a Citrus tristeza virus or Beet yellows virus;
- the Comovirus is Cowpea mosaic virus, Apple latent spherical virus, or Soybean latent spherical virus;
- the Potexvirus is Potato virus X or Citrus yellow vein clearing virus;
- the Potyvirus is a Pepper mottle virus, Bean yellow mosaic virus, Barley stripe mosaic virus, Wheat stripe mosaic virus, Rice yellow mottle virus, Maize dwarf mosaic virus, zucchini yellow mosaic virus, watermelon mosaic virus, or sugarcane mosaic virus;
- the Tobamovirus is a Tobacco mosaic virus, Tomato mosaic virus, Tomato brown rugose fruit virus, Turnip vein-clearing virus, or Pepper mild mottle virus;
- the Tombusvirus is a Turn
- phenotypic change comprises an increase in the plant’s resistance to stress, optionally wherein the stress comprises at least one abiotic stress comprising nutrient stress, light stress, water stress, heat stress, and/or cold stress, or optionally wherein the stress comprises at least one biotic stress comprising crowding, shading, or allelopathy.
- the stress comprises at least one abiotic stress comprising nutrient stress, light stress, water stress, heat stress, and/or cold stress, or optionally wherein the stress comprises at least one biotic stress comprising crowding, shading, or allelopathy.
- the recombinant RNA molecule is provided to the plant of plant cell in the form of an RNA, an encapsidated RNA, or a formulation thereof.
- RNA comprises a synthetic amalgavirus satellite particle.
- the providing comprises contacting the plant or plant cell with the RNA, encapsidated RNA, or formulation thereof, optionally wherein contacting comprises spraying, dusting, injecting, or soaking the plant or plant cell with the RNA, encapsidated RNA, or formulation thereof.
- the recombinant RNA further comprises its reverse complementary RNA molecule.
- any one of embodiments 57 to 70 further comprising providing an hrvRdRP to the plant which recognizes the HRV 5’ or 3’ replication region and/or the subgenomic promoter in the synthetic amalgavirus satellite RNA, optionally wherein the hrvRdRP is provided by introducing a recombinant DNA or RNA encoding the hrvRdRP into the plant or a part thereof. [0213] 72.
- a method of manufacturing a synthetic amalgavirus satellite particle comprising combining the recombinant RNA molecule of any one of embodiments 1 to 28 with a viral capsid protein, wherein the recombinant RNA molecule comprises an encapsidation recognition element (ERE), and wherein the ERE provides for encapsidation of the RNA by the viral capsid protein. .
- ERE encapsidation recognition element
- the combining comprises (a) providing to a plant cell the recombinant RNA molecule, wherein the recombinant RNA molecule comprises an encapsidation recognition element (ERE), and wherein the plant cell comprises an RdRP protein that recognizes the 5’ RNA replication element and 3’ RNA replication element catalyzes synthesis of a synthetic amalgavirus satellite RNA from the recombinant RNA molecule and a viral capsid protein, wherein the ERE provides for encapsidation of the RNA by the viral capsid protein; and optionally Agent Ref: P14357WO00 - 71 - (b) isolating the synthetic amalgavirus satellite particle from the plant cell, a plant comprising the plant cell, or from media in which the plant cell or the plant had been grown.
- ERP encapsidation recognition element
- [0216] 75 The method of embodiment 72, 73, or 74, further comprising the step of formulating the synthetic amalgavirus satellite particle wherein the formulating comprises combining the synthetic amalgavirus satellite particle with a carrier, an excipient, and/or an adjuvant.
- 76 A plant propagule comprising the recombinant RNA molecule of any one of embodiments 1 to 28 and an amalgavirus RdRP, optionally wherein the plant propagule further comprises a heterologous RNA virus RdRP which recognizes an HRV 5’ or 3’ replication region and/or the subgenomic promoter in the synthetic amalgavirus satellite RNA.
- 77 A plant propagule comprising the recombinant RNA molecule of any one of embodiments 1 to 28 and an amalgavirus RdRP, optionally wherein the plant propagule further comprises a heterologous RNA virus RdRP which recognizes an HRV 5’ or 3’ replication region and/or the subgenomic promoter
- the plant propagule of embodiment 76 wherein the plant propagule is a seed, a seedling, root, stem, leaf, shoot, tuber, rhizome, stolon, bulb, explant, embryo, or callus.
- 78. The plant propagule of embodiment 76 or 77, wherein the plant propagule is a mosaic comprising both plant cells comprising the recombinant RNA molecule and plant cells lacking the recombinant RNA molecule.
- 79. The plant propagule of embodiment 76, 77,or 78, wherein the plant propagule lacks DNA encoding the recombinant RNA molecule.
- a heterologous RNA virus RdRP which recognizes an HRV 5’ or 3’ replication region and/or the subgenomic promoter in the synthetic amalgavirus satellite RNA
- the heterologous RNA virus RdRP is an Alphaflexivirus, Betaflexi
- a plant comprising the recombinant RNA molecule of any one of embodiments 1 to 28 and an amalgavirus RdRP, optionally wherein the plant propagule further comprises a heterologous RNA virus RdRP which recognizes an HRV 5’ or 3’ replication region and/or the subgenomic promoter in the synthetic amalgavirus satellite RNA.
- a heterologous RNA virus RdRP which recognizes an HRV 5’ or 3’ replication region and/or the subgenomic promoter in the synthetic amalgavirus satellite RNA.
- 85. The plant of any one of embodiments 81 to 84, wherein the plant comprises an amalgavirus, and wherein the amalgavirus RdRP is provided to the plant cell by the amalgavirus, optionally wherein the amalgavirus RdRP comprises a protein having at least 85%, 90%, 95%, 97%, or 99% sequence identity to SEQ ID NO: 476, 477, 478, 730 to 813, or 814. [0227] 86.
- the plant of any one of embodiments 81 to 86, wherein the amalgavirus RdRP, the 5’ RNA replication element, and/or the 3’ RNA replication element are derived from an amalgavirus comprising one or both of the amalgavirus RdRP, 5’ RNA replication element, and/or 3’ RNA replication elements. [0229] 88.
- RNA virus RdRP which recognizes an HRV 5’ or 3’ replication region and/or the subgenomic promoter in the synthetic amalgavirus satellite RNA
- the heterologous RNA virus RdRP is an Alphaflexivirus, Betaflexivirus, Bromovirus, Celavirus, a Closterovirus, a Comovirus, a Potexvirus, Potyvirus, Tobamovirus, Tombusvirus, Tospoviridae, Trivirinae, Tymovirus, Varicosavirus, or Secoviridae RdRP or an RdRP set forth in Table 7. [0233] 92.
- An amalgavirus satellite system that is self-replicating when introduced into a plant or plant cell, comprising: (a) a recombinant amalgavirus satellite RNA of any one of embodiments 1 to 28;and (b) an exogenous amalgavirus that is capable of replication in the plant or plant cells and that encodes the amalgavirus RdRP that recognizes the 5’ and 3’ replicase recognition sequences in the recombinant amalgavirus satellite RNA, optionally wherein the amalgavirus satellite system further comprises a heterologous RNA virus RdRP which recognizes an HRV 5’ or 3’ replication region and/or the subgenomic promoter in the synthetic amalgavirus satellite RNA. [0234] 93.
- the self-replicating amalgavirus satellite system of embodiment 92 wherein the exogenous amalgavirus is endemic or native to a different species, variety, or germplasm of plant.
- the amalgavirus satellite system further comprises a heterologous RNA virus (HRV) RdRP which recognizes an HRV 5’ or 3’ replication region and/or the subgenomic promoter in the synthetic amalgavirus satellite RNA, and the heterologous RNA virus RdRP is optionally an Alphaflexivirus, Betaflexivirus, Bromovirus, Celavirus, a Closterovirus, a Comovirus, a Potexvirus, Potyvirus, Tobamovirus, Tombusvirus, Tospoviridae, Trivirinae, Tymovirus, Varicosavirus, or Secoviridae RdRP or an RdRP set forth in Table 7.
- HRV heterologous RNA virus
- Agent Ref P14357WO00 - 73 - [0236]
- a recombinant DNA molecule comprising a first promoter which is operably linked to DNA encoding the RNA molecule of any one of embodiments 1-28.
- 96. A cell comprising the recombinant DNA molecule of embodiment 95, wherein the cell is a bacterial cell, a fungal cell, a plant cell, an insect cell, or an invertebrate animal cell.
- 97 A vector for bacterially mediated plant transformation, comprising the recombinant DNA molecule of embodiment 95.
- the vector of embodiment 97 wherein the bacterium that mediates the plant transformation is an Agrobacterium sp., a Sinorhizobium sp., a Mesorhizobium sp., a Bradyrhizobium sp., Rhizobium sp., or an Ensifer sp. and the vector is adapted for transformation with the bacterium.
- the vector of embodiment 97 or 98, wherein the bacterium that mediates the plant transformation is an Agrobacterium sp., and wherein the vector further comprises T-DNAs flanking the DNA molecule encoding the recombinant RNA molecule. [0241] 100.
- An expression system comprising: (a) the recombinant DNA molecule of embodiment 95; and(b) a cell containing the recombinant DNA molecule and an RdRP protein that recognizes the 5’ and 3’ RNA replication elements encoded by the DNA molecule.
- the recombinant DNA molecule further comprises at least one additional element comprising: (i) DNA encoding at least one RNA encoding a viral movement protein (MP); (ii) DNA encoding at least one tRNA-like molecule; (iii) DNA encoding an encapsidation recognition element (ERE); (iv) DNA encoding an RNA comprising, from 5’ to 3’ and operably linked, a 5’ RNA replication element recognized by an amalgavirus RNA-dependent RNA polymerase (RdRP), the RNA of (i) and optionally an operably linked RNA of (ii) and/or (iii), and a 3’ RNA replication element; (v) DNA encoding an RNA promoter; (vi) DNA encoding an RNA- dependent RNA polymerase (RdRP) that recognizes an HRV 5’ or 3’ replication region and/or a subgenomic promoter; and/or (vii) a
- 103 The expression system of embodiments 101 or 102, wherein the cell is a bacterial cell, a plant cell, a fungal cell, an insect cell, an invertebrate animal cell, or a vertebrate animal cell, optionally wherein the bacterial cell is an Agrobacterium sp., a Sinorhizobium sp., a Mesorhizobium sp., Bradyrhizobium sp., Rhizobium sp., or an Ensifer sp. cell. [0245] 104.
- RNA-binding protein that can bind to the RNA molecule encoded by the DNA molecule, optionally wherein the RBP binds to an RNA effecter;
- RBP RNA-binding protein
- an RNA cleavage agent that cleaves the RNA molecule; and/or
- an RNA promoter dependent RNA polymerase (RdRP) protein that recognizes an HRV 5’ or 3’ replication region and/or a subgenomic promoter recognizes an RNA promoter in the RNA molecule.
- Agent Ref P14357WO00 - 74 - [0246] 105.
- invention 104 or 105 wherein: (i) the capsid protein, viral movement protein (MP), RdRP protein, and/or the RdRP protein that recognizes an HRV 5’ or 3’ replication region and/or a subgenomic promoter is heterologous to the cell and/or (ii) wherein the RdRP protein or a polynucleotide encoding the RdRP protein is provided exogenously to the cell.
- MP viral movement protein
- RdRP protein and/or the RdRP protein that recognizes an HRV 5’ or 3’ replication region and/or a subgenomic promoter is heterologous to the cell and/or (ii) wherein the RdRP protein or a polynucleotide encoding the RdRP protein is provided exogenously to the cell.
- 107 The expression system of any one of embodiments 101 to 106, wherein the cell is a plant cell.
- 108 The expression system of any one of embodiments 101 to 106, wherein the cell is a
- the expression system of 107 wherein the plant cell contains an amalgavirus which expresses the RdRP protein that recognizes the 5’ RNA replication element and the 3’ RNA replication element.
- 109 The expression system of embodiment 108, wherein the amalgavirus occurs naturally in the plant cell.
- 110 The expression system of any one of embodiments 101 to 109, wherein the recombinant DNA molecule further comprises at least one RNA encoding a viral MP, a tRNA-like molecule from an Arabidopsis FT mRNA, and an encapsidation recognition element comprising a TMV-OAS.
- An agricultural formulation comprising the expression system of any one of embodiments 101 to 110.
- a method of producing an exogenous polypeptide in a plant or plant cell comprising: providing a plant or plant cell comprising the recombinant RNA molecule of any one of embodiments 1 to 28 or the recombinant DNA molecule of embodiment 95, wherein the cargo RNA molecule encoded by the RNA or DNA molecule comprises a translatable messenger RNA encoding the exogenous polypeptide, wherein the plant or plant cell comprises an RdRP protein that recognizes the 5’ RNA replication element and the 3’ RNA replication element of the recombinant RNA and that catalyzes synthesis of a synthetic amalgavirus satellite RNA from the recombinant RNA molecule, and wherein the exogenous polypeptide is translated from the translatable messenger; optionally wherein the plant or plant cell further comprises a heterologous RNA virus (HRV) RNA promoter dependent RNA polymerase (hrvRdRP) protein that recognizes an HRV 5’ or 3’ replication region and/or a sub
- Agent Ref P14357WO00 - 75 - [0257] 116.
- the method of embodiment 115 further comprising a first step of providing a population of plants comprising the plant cells comprising the amalgavirus which provides the RdRP protein; and then providing the recombinant RNA or DNA molecule, the cell, the vector. the recombinant RNA or DNA molecule, or formulation thereof to the plants comprising the plant cells and optionally further comprising providing the hrvRdRP to the plants comprising the plant cells.
- 121 The method of any one of embodiments 115 to 120, further comprising the step of determining if the plant cell comprises an amalgavirus which can provide the RdRP.
- 122 The method of any one of embodiments 115 to 120, further comprising the step of determining if the plant cell comprises an amalgavirus which can provide the RdRP.
- RdRP amalgavirus RNA- dependent RNA polymerase
- the method of embodiment 122 or 123, wherein the mixed population of plant cells comprise plant cells comprising the amalgavirus RdRP and plant cells lacking the amalgavirus RdRP.
- 125 The method of embodiment 122 or 123, wherein the mixed population of plant cells comprise plant cells comprising the amalgavirus RdRP.
- 126 The method of any one of embodiments 122 to 125, wherein the mixed population of plant cells is screened or selected for the presence of the plant cell comprising the recombinant RNA molecule prior to isolating the plant propagule.
- 127 The method of any one of embodiments 122 to 125, wherein the mixed population of plant cells is screened or selected for the presence of the plant cell comprising the recombinant RNA molecule prior to isolating the plant propagule.
- the method of any one of embodiments 122 to 126, wherein the isolation comprises selecting for the plant cell comprising the recombinant RNA molecule prior to isolating the plant propagule. [0269] 128. The method of any one of embodiments 122 to 127, wherein the mixed population is located within a plant or a plant part. Agent Ref: P14357WO00 - 76 - [0270] 129. The method of any one of embodiments 122 to 128, wherein the plant or plant part is screened or selected for presence of the recombinant RNA molecule prior to isolating the plant propagule. [0271] 130.
- any one of embodiments 122 to 132 wherein the plant propagule comprising the recombinant RNA molecule is isolated by detecting the RNA molecule in one or more plant cells comprising the recombinant RNA molecule and separating the one or more plant cells comprising the recombinant RNA molecule from the plant cells lacking the recombinant DNA molecule.
- the plant propagule is a mosaic comprising both plant cells comprising the recombinant RNA molecule and plant cells lacking the recombinant RNA molecule.
- any one of embodiments 122 to 140 wherein the isolated propagule comprises callus and the method further comprises regenerating a plant, seedling, root, stem, leaf, shoot, tuber, rhizome, stolon, bulb, or explant comprising the recombinant RNA from said callus.
- 143 The method of embodiment 141 or 142, wherein a plant is regenerated and wherein the method further comprises recovering F 1 seed or F 1 progeny comprising the recombinant RNA from the plant.
- 144 The method of any one of embodiments 122 to 140, wherein the isolated propagule comprises callus and the method further comprises regenerating a plant, seedling, root, stem, leaf, shoot, tuber, rhizome, stolon, bulb, or explant comprising the recombinant RNA from said callus.
- a method of providing a synthetic amalgavirus satellite RNA to a plant comprising: grafting a scion onto a rootstock comprising recombinant RNA molecule of any one of embodiments 1 to 28, wherein at least one cell of the rootstock and/or the scion comprises the amalgavirus RdRP.
- a scion onto a rootstock comprising recombinant RNA molecule of any one of embodiments 1 to 28, wherein at least one cell of the rootstock and/or the scion comprises the amalgavirus RdRP.
- a method for producing a plant that transmits a recombinant RNA molecule to progeny plants or seed comprising isolating an F 1 progeny plant or seed comprising at least one cell comprising an amalgavirus RNA-dependent RNA polymerase (RdRP) and the recombinant RNA molecule of any one of embodiments 1 to 28 from a population of F 1 plants or seed obtained from a parent plant comprising the recombinant RNA molecule.
- RdRP amalgavirus RNA-dependent RNA polymerase
- the method of embodiment 151, wherein the F 1 progeny plant or seed comprising the cell is isolated by screening the population of F 1 plants or seed obtained from a parent plant for the presence of the recombinant RNA molecule and propagating the F 1 progeny plant or seed comprising the recombinant RNA molecule.
- the recombinant RNA molecule encodes a selectable marker and the F 1 progeny plant or seed comprising the recombinant RNA molecule is isolated by selecting the F 1 progeny plant or seed comprising the recombinant RNA molecule for presence of the selectable marker.
- the method of any one of embodiments 151 to 156, wherein the F 1 progeny plant or seed population is obtained from a parent plant used as a pollen donor.
- 159. The method of any one of embodiments 151 to 156, wherein the F 1 progeny plant or seed population is obtained by selfing the parent plant.
- 160. The method of any one of embodiments 151 to 159, wherein the parent plant or a part thereof comprising the plant cell is screened or selected for presence of the recombinant RNA molecule prior to isolating the F 1 progeny plant or seed. [0302] 161.
- any one of embodiments 151 to 160 wherein the parent plant or one or more parts thereof are screened for systemic presence of the recombinant RNA molecule prior to isolating the F 1 progeny plants, optionally wherein the part comprises floral tissue or male or female reproductive tissue.
- the part comprises floral tissue or male or female reproductive tissue.
- 162 The method of any one of embodiments 151 to 161, wherein the pericarp of the parent plant is screened or selected for presence of the recombinant RNA molecule.
- F 1 seeds are obtained from a parent plant selected for presence of the recombinant RNA molecule in pericarp tissue.
- any one of embodiments 151 to 165 further comprising introducing the recombinant RNA molecule or a polynucleotide encoding the recombinant RNA molecule into a plant cell and obtaining the parent plant comprising the recombinant RNA molecule from the plant cell.
- 167 The method of any one of embodiments 151 to 166, wherein the propagule, plant, plant part, scion, and/or rootstock comprises a heterologous viral coat protein which can encapsidate the recombinant RNA molecule and/or comprises the recombinant RNA molecule encapsidated by a heterologous viral coat protein.
- the propagule, plant, plant part, scion, and/or rootstock comprises a heterologous viral coat protein which can encapsidate the recombinant RNA molecule and/or comprises the recombinant RNA molecule encapsidated by a heterologous viral coat protein.
- a method of barcoding a plant, plant cell, progeny thereof, or part thereof comprising providing to the plant or plant cell the recombinant RNA molecule of any one of embodiments 1 to 28, Agent Ref: P14357WO00 - 79 - wherein the cargo RNA of the recombinant RNA molecule comprises a barcode RNA molecule, and wherein the plant or plant cell comprises an amalgavirus RNA-dependent RNA polymerase (RdRP).
- RdRP amalgavirus RNA-dependent RNA polymerase
- the barcode RNA molecule comprises a sequence that is not present in the genome and/or transcriptome of a wild-type plant of the same species, a pathogen thereof, or a symbiont thereof. [0312] 171. The method of any one of embodiments 168 to 170, wherein the barcode RNA molecule comprises a random sequence. [0313] 172. The method of any one of embodiments 168 to 171, wherein the barcode RNA molecule comprises a forward primer binding site and a reverse primer binding site that is not present in the genome and/or transcriptome of a wild-type plant of the same species, a pathogen thereof, or a symbiont thereof. [0314] 173.
- any one of embodiments 168 to 172, wherein the barcode RNA molecule is up to about 3.2 kb in length. [0315] 174.
- any one of embodiments 168 to 176 further comprising isolating an F 1 progeny plant or seed comprising at least one cell comprising the amalgavirus RdRP and the recombinant RNA molecule.
- 179. The method of embodiment 178, wherein the F 1 progeny plant or seed is obtained from the plant used as a pollen recipient.
- 180. The method of embodiment 178, wherein the F 1 progeny plant or seed is obtained from the plant used as a pollen donor.
- 181 The method of embodiment 178, wherein the F 1 progeny plant or seed is obtained by selfing the plant.
- Agent Ref P14357WO00 - 80 - [0323] 183.
- a method of identifying a barcoded plant, plant part, or plant cell the method comprising screening for the presence of a barcode RNA molecule in the plant, plant part, or plant cell, wherein the plant, plant part, or plant cell comprises the recombinant RNA molecule of any one of embodiments 1 to 28, wherein the cargo RNA of the recombinant RNA molecule comprises the barcode RNA molecule. Summary of Tables Table 1.
- RNA replication element DNA Coding Sequences SEQ Description 1 RRE ID Dot-Bracket structure of encoded RNA 2 Agent Ref: P14357WO00 - 81 - SEQ Description 1 RRE ID Dot-Bracket structure of encoded RNA NO 2 Agent Ref: P14357WO00 - 82 - SEQ Description 1 RRE ID Dot-Bracket structure of encoded RNA NO 2 Agent Ref: P14357WO00 - 83 - SEQ Description 1 RRE ID Dot-Bracket structure of encoded RNA NO 2 Agent Ref: P14357WO00 - 84 - SEQ Description 1 RRE ID Dot-Bracket structure of encoded RNA NO 2 Agent Ref: P14357WO00 - 85 - SEQ Description 1 RRE ID Dot-Bracket structure of encoded RNA NO 2 Agent Ref: P14357WO00 - 85 - SEQ Description 1 RRE ID Dot-Bracket structure of encoded RNA NO 2 Agent Ref: P14357WO00
- IRES sequences IRES source NCBI Accession ID 1 Sequence 2 Agent Ref: P14357WO00 - 99 - Solenopsis invicta virus NC_006559 SEQ ID NO: 130 Triatoma virus NC_003783 SEQ ID NO: 131 S EQ ID NO: 132 y Information database “ncbi[dot]nlm[dot]nih.gov/nuccore.” 2 RNA equivalents of the DNA sequences are also contemplated and can be obtained from the DNA sequences provided. Table 6.
- Insecticidal or Insect Inhibitory and Nematocidal or Nematode Inhibitory Cargo Molecules NT AA Examples Se uence Se uence of Tar et Agent Ref: P14357WO00 - 104 - NT AA Examples Sequence Sequence of Target NCBI Accession (SEQ ID (SEQ ID Cargo Crops , Agent Ref: P14357WO00 - 105 - NT AA Examples Sequence Sequence of Target NCBI Accession (SEQ ID (SEQ ID Cargo Crops Agent Ref: P14357WO00 - 106 - NT AA Examples Sequence Sequence of Target NCBI Accession (SEQ ID (SEQ ID Cargo Crops , , , , er for entries in the world wide web internet database ncbi[dot]nlm[dot]nih.gov/nuccore.
- RNAi Targets Cargo (RNAi NCBI Accession Examples of target tar et ene) ID 1 Se uence Tar et athoen cro Agent Ref: P14357WO00 - 107 - Cargo (RNAi NCBI Accession Examples of target target gene) ID 1 Sequence Target pathogen crop SEQ ID Agent Ref: P14357WO00 - 108 - Cargo (RNAi NCBI Accession Examples of target target gene) ID 1 Sequence Target pathogen crop Polygalacturonase SEQ ID tomato wheat er o e es e wo w e we e e aaase c o o .gov uccoe. Table 11.
- Antifungal cargo protein Examples NCBI Accession NT AA Target of target r Agent Ref: P14357WO00 - 109 - Examples NCBI Accession NT AA Target of target Cargo ID 1 Sequence Sequence pathogen crop ell y rn ell ell rn Agent Ref: P14357WO00 - 110 - Examples NCBI Accession NT AA Target of target Cargo ID 1 Sequence Sequence pathogen crop rn s s s s s s, s, s, s, s, Agent Ref: P14357WO00 - 111 - Examples NCBI Accession NT AA Target of target Cargo ID 1 Sequence Sequence pathogen crop ber Table 12.
- Agent Ref P14357WO00 - 112 - NCBI or TAIR Gene Source Effector Accession NT AA ce , ce ce ce
- Bioactive Plant peptides Peptide name Sequence Function SEQ ID NO: Increase nodulation in soybean (improved nitrogen fixation, Ribozymes Ribozyme NCBI Accession ID Rfam ID Sequence 1 KV7674721 Agent Ref: P14357WO00 - 116 - Ribozyme NCBI Accession ID Rfam ID Sequence 1 RF03160; URS0000D69167 4558 sequences provided.
- Amalgavirus RdRP sequences Genomic RdRP DNA R Sequence Sequ dRP 1 ence or 5’ Coding Isolation m Agent Ref: P14357WO00 - 118 - Genomic RdRP DNA Sequenc uence or 5 RdR e Seq P Amalgavirus 1 ’ Coding Pro Isolation RRE sequence Sequence tein Host a .
- Agent Ref P14357WO00 - 119 - Genomic RdRP DNA Sequence Sequence or 5’
- RdRP Amalgavirus 1 Coding Prot Isolation RRE sequence Sequence ein Host a a a is Agent Ref: P14357WO00 - 120 - Genomic RdRP DNA R Sequence Sequence or 5’
- the RdRP coding sequence of SEQ ID NO: 473 and the RdRP Protein sequence of SEQ ID NO: 476 are from NCBI database entry NC_011591.1 (Southern tomato virus, isolate Mexico-1) Table 18.
- RRE AV RdRP AV RdRP Genomic AV 5’ RRE SEQ ID Protein Nucleotide 2 1 O Agent Ref: P14357WO00 - 125 - AV AV 3’
- RRE AV RdRP AV RdRP Genomic AV 5’ RRE SEQ ID Protein Nucleotide SEQ ID SEQ ID
- An antiviral amalgavirus satellite with an imbedded HRV amplicon and RNAi cargo is generally provided in 5’ to 3’ orientation as follows: (i) 5’ RNA replication element from the amalgavirus; (ii) a heterologous RNA virus (HRV) 5’ replication region (HRV 5’RR); (iii) an antiviral RNA molecule which induces an RNAi response; (iv) the heterologous RNA virus (HRV) 3’ RNA replication region (HRV 3’RR); and (v) and the 3’ RNA replication element from the amalgavirus.
- COMSAT amalgavirus satellite that carries an antiviral inhibitory RNA (RNAi) cargo is generally provided in 5’ to 3’ orientation as follows: (i) 5’ RNA replication element from the amalgavirus; (ii) a heterologous RNA virus (HRV) 5’ replication region (HRV 5’RR); (iii) an antiviral RNA molecule which induces an
- a COMSAT comprising amalgavirus 5’ and 3’ RNA replication elements flanking a tobacco mosaic virus (TMV) amplicon containing a cargo RNA containing a Pepper mild mottle virus (PMMoV) RNAi inducing sequence (e.g., a PMMoV sequence which can form a dsRNA) is provided as SEQ ID NO: 482.
- TMV tobacco mosaic virus
- PMMoV Pepper mild mottle virus
- This satellite includes TMV 5’ and 3’ replication region sequences (HRV 5’ and 3’ RR sequences) to promote secondary amplification by TMV, which is a pathogenic tobamovirus.
- the elements of the satellite are set forth in Table 19.
- Similar antiviral amalgavirus satellites are designed for secondary amplification by tomato mosaic virus or tomato brown rugose fruit virus (which are also Agent Ref: P14357WO00 - 131 - pathogenic tobamoviruses), and include an N protein (SEQ ID NO:254), L4 protein (SEQ ID NO:256), or a combination of N and L4 protein as the antiviral cargo sequence(s), and replacing the HRV 5’ and 3’ RR sequences of TMV with the HRV 5’ and 3’ RR sequences of either tomato mosaic virus or tomato brown rugose fruit virus as provided in Table 7, to promote secondary amplification by tomato mosaic virus or tomato brown rugose fruit virus, respectively.
- N protein SEQ ID NO:254
- L4 protein SEQ ID NO:256
- a combination of N and L4 protein as the antiviral cargo sequence(s)
- replacing the HRV 5’ and 3’ RR sequences of TMV with the HRV 5’ and 3’ RR sequences of either tomato mosaic
- Solanaceous plants containing the amalgavirus (or that otherwise are provided with the appropriate amalgavirus RdRP, e.g., through transient or transgenic expression) and that are provided with these antiviral amalgavirus satellites are expected to exhibit resistance to tobacco mosaic virus, tomato mosaic virus, or tomato brown rugose fruit virus, respectively.
- Table 19 Genetic element Nucleotide position in SEQ ID NO: 482 amalgavirus 5’ RNA replication element 1-128 p .
- An amalgavirus satellite (COMSAT) with an imbedded HRV amplicon that carries an antiviral protein cargo is generally provided in 5’ to 3’ orientation as follows: (i) 5’ RNA replication element from the amalgavirus; (ii) a heterologous RNA virus (HRV) 5’ replication region (HRV 5’RR); (iii) an antiviral protein cargo; (iv) the heterologous RNA virus (HRV) 3’ RNA replication region (HRV 3’RR); and (v) the 3’ RNA replication element from the amalgavirus.
- COMSAT amalgavirus satellite
- a COMSAT comprising amalgavirus 5’ and 3’ RNA replication elements flanking a TMV amplicon containing an antiviral cargo protein containing the coding sequence of the N gene of tobacco is provided as SEQ ID NO: 481.
- the elements of the satellite are set forth in Table 20.
- An amalgavirus satellite (COMSAT) that carries an antiviral RNAi cargo is generally provided in 5’ to 3’ orientation as follows: (i) 5’ RNA replication element from the amalgavirus; (ii) an antiviral RNA molecule which induces an RNAi response; and (iii) the 3’ RNA replication element from the amalgavirus.
- COMSAT amalgavirus satellite
- a COMSAT comprising amalgavirus 5’ and 3’ RNA replication elements flanking a cargo RNA containing a Pepper mild mottle virus (PMMoV) RNAi inducing sequence (e.g., a PMMoV sequence which can form a dsRNA) is provided as SEQ ID NO: 480.
- the elements of the satellite are set forth in Table 21. Table 21. Genetic element Nucleotide position in SEQ ID NO: 480 amal avirus 5’ RNA re lication element 1-128 Example 4.
- An antiviral amalgavirus satellite with antiviral protein cargo is generally provided in 5’ to 3’ orientation as follows: (i) 5’ RNA replication element from the amalgavirus; (ii) an antiviral protein cargo; and (iii) the 3’ RNA replication element from the amalgavirus.
- a COMSAT comprising amalgavirus 5’ and 3’ RNA replication elements flanking an antiviral protein cargo containing the coding sequence of the N gene of tobacco is provided as SEQ ID NO: 479. The elements of the satellite are set forth in Table 22. Table 22.
- Antiviral COMSATs e.g., of Examples 1 to 4
- control RNAs e.g., RNAs lacking the antiviral cargo
- target host plants such as pepper or tomato plants via micro- bombardment (“biolistic” delivery) using a “gene gun” or by bacterially mediated (e.g., by Agrobacterium) transient expression.
- Antiviral COMSATs are prepared either as in vitro transcribed (IVT) products (capped or uncapped), or as Agrobacterium binary vectors.
- IVT in vitro transcribed
- the IVT product is coated onto the surface of gold nanoparticles which are precipitated on inner surface of bullet tubes; these are accelerated at the abaxial surface of the seedlings’ leaves with helium pressure from 100 to 180 psi.
- COMSAT-carrying clones in binary vector are transformed into Agrobacterium GV 2260, and the transformed Agrobacterium is selected by growing at 28°C under appropriate antibiotic selection. The positive transformants are verified by PCR and then grown at 28°C overnight in liquid LB medium with appropriate antibiotics plus rifamycin to maintain the Agrobacterium and prevent contamination. Agrobacterium cells are collected by centrifugation and resuspended in MMA buffer at an OD600 of 0.2, incubated at 28°C on a shaker for 2 hours, and then infiltrated into the abaxial sides of the leaves. Control plants are treated in the same way but with non- COMSAT control IVT transcripts, or empty buffers.
- tissues from systemic leaves are collected and subjected to RNA extraction, followed by cDNA syntheses.
- COMSAT titers are monitored in systemic tissue by qRT-PCR with COMSAT-specific primers.
- the efficacy of an antiviral COMSAT can be tested by challenging the COMSAT-treated plants with the viral pathogen of interest (e.g., cucumber mosaic virus, CMV, or tobacco mosaic virus, TMV), for example, by mechanical infection.
- the viral pathogen of interest e.g., cucumber mosaic virus, CMV, or tobacco mosaic virus, TMV
- Infectious preparations of such acute viral pathogens are prepared in planta (e.g., in Nicotiana benthamiana) or in the form of GFP-fused infectious clones in a binary vector or T7-based vector.
- Suitable viral inoculums can be prepared as infectious sap extracted from an infected plant, as an Agrobacterium-based inoculum, or as IVT products.
- the inoculum is introduced into leaves of the COMSAT-treated plants by rub-inoculation (for infectious sap or IVT product), by agroinfiltration (for GFP-fused infectious clone in binary vector), or by micro-bombardment Agent Ref: P14357WO00 - 134 - (for IVT product or plasmid of infectious clone in binary vector).
- the infectious sap or IVT products are respectively diluted 3 times or to 50ng/microliter in phosphate buffer 0.05M, pH 7.4 and dropped onto leaf adaxial surfaces which is pre-dusted with an abrasive such as carborundum or bentonite.
- the inoculum is gently spread on the abrasive-dusted leaf surface by gloved fingers or cotton buds; after 30 seconds the inoculated leaf is washed with water.
- Micro-bombardment of infectious clones is conducted following previously described methods (see, e.g., delivery techniques described in PCT/US22/78963; see also Bio-Rad Tech Note 2531 “Inoculation of Viral RNA and cDNA to Potato and Tobacco Plants Using the HeliosTM Gene Gun”).
- Infiltration with an Agrobacterium-based viral pathogen inoculum is performed using methods similar to that used for COMSAT infiltration, but the Agrobacterium suspension is diluted at OD600 of 0.1.
- Symptoms and titre of the acute viral pathogens are monitored in the plants, typically over time to confirm progress or decline of viral infection. Effectiveness of antiviral COMSATs is evaluated by comparing relative titres of systemic infected viruses in antiviral COMSAT-treated plants and the control plants. Tissues from inoculated leaves and systemic leaves (distal to the inoculated leaves) are collected, total RNA is extracted, optionally followed by cDNA synthesis. The viral titre can be quantitatively measured by qRT-PCR with virus-specific primers.
- Viral titre or presence can be measured by immunoassay methods; for example, Tobacco mosaic virus (TMV) is routinely detected in plants using a commercially available strip assay (Agdia ImmunoStrip ® for TMV Agdia, Inc., Elkhart, IN, USA).
- TMV Tobacco mosaic virus
- strip assay Agdia ImmunoStrip ® for TMV Agdia, Inc., Elkhart, IN, USA.
- viral titre is qualitatively evaluated by viral disease symptoms, or by proxy measurement (e.g., measuring GFP expressed by GFP-fused viral constructs).
- qRT-PCR measurements are normalized to endogenous reference gene controls (e.g., actin, tubulin, ubiquitin-3, GADPH, or translation elongation factor EF1a, used individually or preferably in multiples, e.g., at least 3 reference genes or at least three deltaCt values).
- endogenous reference gene controls e.g., actin, tubulin, ubiquitin-3, GADPH, or translation elongation factor EF1a, used individually or preferably in multiples, e.g., at least 3 reference genes or at least three deltaCt values.
- the viral titre is compared to that of controls.
- An insect inhibitory amalgavirus satellite with insecticidal protein or RNAi cargo is generally provided in 5’ to 3’ orientation as follows: (i) 5’ RNA replication element from the amalgavirus; (ii) an insecticidal RNA molecule which induces an RNAi response; and (iii) the 3’ RNA replication element from the amalgavirus.
- a COMSAT comprising amalgavirus 5’ and 3’ RNA replication elements flanking a cargo RNA containing an RNAi inducing sequence that targets Colorado potato beetle is provided as SEQ ID NO: 484. The elements of the satellite are set forth in Table 23.
- Table 23 Genetic element Nucleotide position in SEQ ID NO: 484 Agent Ref: P14357WO00 - 135 - ZmHSP101 IRES 617-763 TMV MP 764-1570 provided in 5’ to 3’ orientation as follows: (i) 5’ RNA replication element from the amalgavirus; (ii) an insecticidal protein cargo; and (iii) the 3’ RNA replication element from the amalgavirus. [0340] A COMSAT comprising amalgavirus 5’ and 3’ RNA replication elements flanking an insecticidal cargo protein containing the coding sequence of the Vip3Aa gene is provided as SEQ ID NO: 483. The elements of the satellite are set forth in Table 24. Table 24.
- RNA replication element 1-128 o a e an a e are esta s e n ost pants essent a y as set ort n Example 5.
- the efficacy of an insecticidal or insect inhibitory COMSAT can be tested by challenging the COMSAT and control treated plants with the insect of interest (e.g., Colorado Potato Beetle for plants treated with the Table 23 COMSAT or Fall Armyworm, Corn Earworm, or Black Cutworm for plants treated with the Table 24 COMSAT).
- insect of interest e.g., Colorado Potato Beetle for plants treated with the Table 23 COMSAT or Fall Armyworm, Corn Earworm, or Black Cutworm for plants treated with the Table 24 COMSAT.
- An antifungal amalgavirus satellite with antifungal protein or RNAi cargo is generally provided in 5’ to 3’ orientation as follows: (i) 5’ RNA replication element from the amalgavirus; (ii) an antifungal RNA molecule which induces an RNAi response; and (iii) the 3’ RNA replication element from the amalgavirus.
- a COMSAT comprising amalgavirus 5’ and 3’ RNA replication elements flanking a cargo RNA containing an RNAi inducing sequence that targets a DCL gene of Botrytis cinerea is provided as SEQ ID NO: 486.
- the elements of the satellite are set forth in Table 25.
- Agent Ref P14357WO00 - 136 - Genetic element Nucleotide position in SEQ ID NO: 486 amalgavirus 5’ RNA replication element 1-128 provided in 5’ to 3’ orientation as follows: (i) 5’ RNA replication element from the amalgavirus; (ii) an antifungal protein cargo; and (iii) the 3’ RNA replication element from the amalgavirus.
- a COMSAT comprising amalgavirus 5’ and 3’ RNA replication elements flanking an antifungal cargo protein containing the coding sequence of the CaAMP1 gene is provided as SEQ ID NO: 485.
- the elements of the satellite are set forth in Table 26. Table 26.
- a COMSAT construct is built by Golden Gate assembly of synthesized cargos and other COMSAT components synthesized from a commercial vendor.
- the plasmid is used as a template for a PCR reaction to amplify the expression cassette with a T7 promoter.
- This PCR product is used as the template for an in vitro transcription reaction using a MEGAscriptTM T7 Transcription Kit from Thermo Fisher Scientific.
- RNA or protein cargo is then transformed into leaves of the target crop using gold nanoparticles from Bio-Rad according to the guidelines specified in the Helios gene gun manual or via Agrobacterium mediated transformation.
- Replication and persistence of delivered RNA or expression of protein in local leaves can be determined by RT-PCR from local tissues at 2-weeks post transformation.
- Systemic movement of RNA/protein is determined by qRT-PCR/ELISA in the sample collected from distal untreated leaf samples after 1-, 2-, and 3-months post transformation.
- Agent Ref P14357WO00 - 137 - [0347]
- An DNA molecule encoding an RNA molecule containing a 5’ replicase recognition sequence from amalgavirus; Dicer-like 1 and Dicer-like 2 fragments; ZmHSP101 IRES; TMV movement protein; Flowering Locus T from Arabidopsis thaliana; and a 3’ replicase recognition sequence from amalgavirus is cloned into a pUC19 plasmid backbone and transformed into E. coli DH5a (New England Biolabs; C2987I). The plasmid is extracted, and PCR is used to add a T7 promotor sequence.
- RNA quantity is measured using a Nanodrop spectrophotometer (Thermo Fisher Scientific; 840274200) and the intactness is quality controlled through gel electrophoresis.
- the synthesized and quality-controlled RNA molecule is precipitated onto gold nanoparticles and fired into 4-week-old Solanum lycopersicum ‘Early Girl’ leaves using a gene gun.
- RNA is extracted with the MagMax Plant RNA Isolation kit (Thermo Fisher Scientific; A33899) and quality controlled through Nanodrop spectrophotometer (Thermo Fisher Scientific; 840274200) and RNA ScreenTape Analysis (Agilent; 5067-5576). Quality controlled RNA is used for cDNA synthesis and qRT-PCR is used to amplify the Dicer-like 1 and Dicer-like 2 fragments and reference gene fragment RPL2.
- Amalgavirus COMSAT with cDNA-based cargo expression amplification A DNA molecule encoding an RNA molecule containing a 5' replicase recognition sequence from amalgavirus; Cricket paralysis virus IRES; Avian leukosis virus reverse transcriptase; tRNA primer binding site (tRNA-trp); CaMV 35s promoter; a protein cargo (Superfolder GFP); ZmHSP101 IRES; Tobacco mosaic virus movement protein; and a 3' replicase recognition site from amalgavirus is cloned into a pUC19 plasmid backbone and is transformed into E.
- RNA quantity is measured using a Nanodrop spectrophotometer (Thermo Fisher Scientific; 840274200) and the intactness is quality controlled through gel electrophoresis.
- the synthesized and quality-controlled RNA molecule is precipitated onto gold nanoparticles and fired into the 4-week-old Solanum lycopersicum ‘Early Girl’ leaves using a gene gun.
- Agent Ref P14357WO00 - 138 -
- leaf tissue samples are collected at 2-week and 4-week timepoints: fired leaf # 1; fired leaf # 2; nearest adjacent leaflet; opposite leaf, terminal leaflet; and apical leaf growth.
- Protein (GFP) expression is quantified. Total protein is extracted from collected tissue samples using the Pierce Plant Total Protein Extraction Kit (Thermo Fisher Scientific; A44056) according to manufacturer specifications, and quality controlled by Nanodrop spectrophotometry paired with SDS-PAGE to verify extraction yield and integrity. GFP expression is quantified relative to other protein expressing COMSAT designs as well as non-transfected tissue using the Abcam GFP ELISA Kit (ab171581) according to manufacturer specifications.
- Example 10 Example 10
- RNA Negative-Sense Strand where only the positive-sense strand of a satellite RNA is provided to a plant or plant cell (e.g., by delivery of the RNA itself or by delivery of DNA encoding the satellite RNA’s positive-sense strand), replication of the satellite RNA in a plant or plant cell can be verified by detection of the satellite RNA’s negative-sense strand, for example, by using a negative-sense-strand-specific Taqman® assay.
- a bell pepper endornavirus (BPEV) satellite of 2357 nucleotides was constructed with the following elements in 5’ to 3’ orientation: (1) BPEV 5’ RNA replication element; (2) a first IRES sequence (PSVI IRES, with the addition of spacer nucleotides on the 5’ end); (3) RNA encoding a reporter protein (GFP); (4) a second IRES sequence (ZmHSP IRES); (5) a viral movement protein; (6) a zebrafish sequence (as a non-plant heterologous sequence for detection purposes); (7) an isoleucine tRNA sequence; (8) BPEV 3’ RNA replication element.
- PSVI IRES first IRES sequence
- GFP reporter protein
- ZmHSP IRES ZmHSP IRES
- Similar satellites can be designed and constructed using alternative genetic elements such as those provided elsewhere in this specification, such as other pairs of 5’ and 3’ RNA replication elements, other pairs of HRV 5’ and 3’ replication region sequences, and/or other cargo sequences.
- the positive-sense strand of this BPEV satellite RNA was produced through in vitro transcription (IVT), and biolistically delivered with a gene gun into bell pepper plant leaves. Samples were collected 4 weeks post gene gun firing from a local leaf.
- RNA extraction, library prep, and Illumina® sequencing using a stranded kit (NEBNext® UltraTM II Directional RNA Library Prep Kit for Illumina®, catalogue numbers E7760S or E7760L, New England Biolabs, Ipswich, MA) with added DMSO treatment and plant rRNA depletion.
- Reads were mapped against the BPEV satellite RNA’s sequence and negative-sense reads were identified using the Integrative Genomics Viewer (IGV) browser (Robinson et al., Bioinformatics, 39(1) (2023), btac830). Negative-sense reads mapping to the BPEV satellite were present, confirming that replication of the BPEV satellite had occurred.
- IGV Integrative Genomics Viewer
- BPEV negative-sense reads were also identified in BPEV infected plants that served as a positive control for the analysis.
- the above-described negative-strand detection method can be adapted for use in the detection of negative strand RNA produced by amalgavirus satellite RNAs disclosed herein.
- Agent Ref P14357WO00 - 139 - Example 11.
- RNA Negative-Sense Strand in a Tomato Plant where only the positive-sense strand of a satellite RNA is provided to a plant or plant cell (e.g., by delivery of the RNA itself or by delivery of DNA encoding the satellite RNA’s positive-sense strand), replication of the satellite RNA in a plant or plant cell can be verified by detection of the satellite RNA’s negative-sense strand, for example, by using a negative-sense-strand-specific Taqman® assay.
- Negative sense GFP produced by a satellite RNA containing a GFP protein encoding cargo sequences is detected using Taqman® qPCR.
- the reference sequence that can be used is an endogenous tomato gene, “Tom06” (NCBI Gene ID 101249087, ubiquitin-conjugating enzyme E2-17 kDa, from Solanum lycopersicum).
- Taqman® qPCR primers and probes were ordered from Integrated DNA Technology, Coralville, IA.
- the negative sense GFP probe is labeled with FAM520 dye with ZEN/Iowa Black FQ quencher.
- the Tom06 probe is labeled with SUN5544 with ZEN/Iowa Black FQ quencher. Primers and probes are used at a 2:1 ratio.
- cDNA is synthesized via reverse transcription (30 minutes at 60 degrees C, then 10 minutes at 85 degrees C) using Maxima H Minus First Strand cDNA synthesis Kit (K1652, Thermo Fisher Scientific, Waltham MA) using the RT primers in Table 26. This was followed by treatment (30 minutes at 37 degrees C, then 1 minute at 80 degrees C) with thermolabile exonuclease I (E1050 New England Biolabs, Ipswich MA) to remove excess RT primers. The synthesized cDNA was cleaned with RNAClean XP beads (A63987 Beckman Coulter, Brea CA).
- Taqman® qPCR is carried out using Taqman® Advanced Fast MasterMix (444965, Thermo Fisher Scientific, Waltham MA) and the primer and probes in Table 27.
- Table 27. Description of Reverse Transcriptase Primers RT Primer Name RT Primer Sequence i i i i A AAA AA AG Table 28. Description of qPCR primers and probes qPCR Primer/Probe Name qPCR Primer/Probe Sequence k Q/ Agent Ref: P14357WO00 - 140 - [0356]
- the above-described negative-strand detection method can be adapted for use in the detection of negative strand RNA produced by amalgavirus satellite RNAs disclosed herein.
- An antiviral amalgaviral satellite with an imbedded HRV amplicon designed to promote secondary amplification by Citrus tristeza virus is provided in 5’ to 3’ orientation as follows: (i) 5' replication element from an endornavirus; (ii) an HRV amplicon in either sense or anti-sense orientation comprising a Citrus tristeza virus 5’ replication region (HRV 5’RR) which is operably linked to RNA encoding an antiviral protein cargo or antiviral RNA cargo (e.g., an RNA molecule which induces an RNAi response) which is operably linked to the Citrus tristeza virus 3’ RNA replication region (HRV 3’RR); and (iii) the 3’ RNA replication element from an endornavirus is provided.
- the antiviral protein or RNA cargo can be specifically anti-CTV or have broader antiviral activity.
- the amalgaviral satellite is delivered via application and/or injection into citrus trees (see, e.g., US Patent No.11,844,318, International Patent Application publication WO2023/240208, Al-Rimawi et al. (2019) Antibiotics, 8, 196; doi:10.3390/antibiotics8040196, each incorporated herein by reference in their entireties).
- the amalgaviral satellite is delivered into the vascular system (e.g., xylem) of a citrus tree, for example, by injection directly into the tree’s vascular system.
- Citrus trees containing the amalgavirus or that otherwise are provided with the appropriate rndornaviral RdRP that recognizes the 5’ and 3’ RNA replication element of the amalgaviral satellites (and, in the case of constructs containing an HRV amplicon, also containing or otherwise provided with a CTV RdRP) and that are provided with these antiviral amalgaviral satellites are expected to exhibit resistance to Citrus tristeza virus.
Landscapes
- Health & Medical Sciences (AREA)
- Life Sciences & Earth Sciences (AREA)
- Genetics & Genomics (AREA)
- Engineering & Computer Science (AREA)
- Biomedical Technology (AREA)
- Biotechnology (AREA)
- Bioinformatics & Cheminformatics (AREA)
- Organic Chemistry (AREA)
- Chemical & Material Sciences (AREA)
- Zoology (AREA)
- General Engineering & Computer Science (AREA)
- Wood Science & Technology (AREA)
- Molecular Biology (AREA)
- Microbiology (AREA)
- Plant Pathology (AREA)
- Cell Biology (AREA)
- Biophysics (AREA)
- Biochemistry (AREA)
- General Health & Medical Sciences (AREA)
- Physics & Mathematics (AREA)
- Virology (AREA)
- Micro-Organisms Or Cultivation Processes Thereof (AREA)
- Agricultural Chemicals And Associated Chemicals (AREA)
- Medicines Containing Material From Animals Or Micro-Organisms (AREA)
Abstract
Description
Claims
Applications Claiming Priority (2)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| US202363499722P | 2023-05-03 | 2023-05-03 | |
| US63/499,722 | 2023-05-03 |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| WO2024229398A1 true WO2024229398A1 (en) | 2024-11-07 |
Family
ID=91302521
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/US2024/027769 Pending WO2024229398A1 (en) | 2023-05-03 | 2024-05-03 | Artificial amalgavirus satellite rnas |
Country Status (2)
| Country | Link |
|---|---|
| AR (1) | AR132586A1 (en) |
| WO (1) | WO2024229398A1 (en) |
Citations (147)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US5229114A (en) | 1987-08-20 | 1993-07-20 | The United States Of America As Represented By The Secretary Of Agriculture | Approaches useful for the control of root nodulation of leguminous plants |
| US5304730A (en) | 1991-09-03 | 1994-04-19 | Monsanto Company | Virus resistant plants and method therefore |
| US5322938A (en) | 1987-01-13 | 1994-06-21 | Monsanto Company | DNA sequence for enhancing the efficiency of transcription |
| US5380831A (en) | 1986-04-04 | 1995-01-10 | Mycogen Plant Science, Inc. | Synthetic insecticidal crystal protein gene |
| US5463175A (en) | 1990-06-25 | 1995-10-31 | Monsanto Company | Glyphosate tolerant plants |
| US5512466A (en) | 1990-12-26 | 1996-04-30 | Monsanto Company | Control of fruit ripening and senescence in plants |
| US5516671A (en) | 1993-11-24 | 1996-05-14 | Monsanto Company | Method of controlling plant pathogens |
| US5543576A (en) | 1990-03-23 | 1996-08-06 | Mogen International | Production of enzymes in seeds and their use |
| US5608149A (en) | 1990-06-18 | 1997-03-04 | Monsanto Company | Enhanced starch biosynthesis in tomatoes |
| US5633435A (en) | 1990-08-31 | 1997-05-27 | Monsanto Company | Glyphosate-tolerant 5-enolpyruvylshikimate-3-phosphate synthases |
| US5641876A (en) | 1990-01-05 | 1997-06-24 | Cornell Research Foundation, Inc. | Rice actin gene and promoter |
| US5689052A (en) | 1993-12-22 | 1997-11-18 | Monsanto Company | Synthetic DNA sequences having enhanced expression in monocotyledonous plants and method for preparation thereof |
| US5689041A (en) | 1989-08-10 | 1997-11-18 | Plant Gentic Systems N.V. | Plants modified with barstar for fertility restoration |
| US5716837A (en) | 1995-02-10 | 1998-02-10 | Monsanto Company | Expression of sucrose phosphorylase in plants |
| US5750876A (en) | 1994-07-28 | 1998-05-12 | Monsanto Company | Isoamylase gene, compositions containing it, and methods of using isoamylases |
| US5763241A (en) | 1987-04-29 | 1998-06-09 | Monsanto Company | Insect resistant plants |
| US5763245A (en) | 1991-09-23 | 1998-06-09 | Monsanto Company | Method of controlling insects |
| US5773696A (en) | 1996-03-29 | 1998-06-30 | Monsanto Company | Antifungal polypeptide and methods for controlling plant pathogenic fungi |
| US5850023A (en) | 1992-11-30 | 1998-12-15 | Monsanto Company | Modified plant viral replicase genes |
| US5858742A (en) | 1983-01-17 | 1999-01-12 | Monsanto Company | Chimeric genes for transforming plant cells using viral promoters |
| US5866775A (en) | 1990-09-28 | 1999-02-02 | Monsanto Company | Glyphosate-tolerant 5-enolpyruvyl-3-phosphoshikimate synthases |
| US5869720A (en) | 1993-09-30 | 1999-02-09 | Monsanto Company | Transgenic cotton plants producing heterologous peroxidase |
| US5880275A (en) | 1989-02-24 | 1999-03-09 | Monsanto Company | Synthetic plant genes from BT kurstaki and method for preparation |
| US5942658A (en) | 1993-07-29 | 1999-08-24 | Monsanto Company | Transformed plant with Bacillus thuringiensis toxin gene |
| US5942664A (en) | 1996-11-27 | 1999-08-24 | Ecogen, Inc. | Bacillus thuringiensis Cry1C compositions toxic to lepidopteran insects and methods for making Cry1C mutants |
| US5958745A (en) | 1996-03-13 | 1999-09-28 | Monsanto Company | Methods of optimizing substrate pools and biosynthesis of poly-β-hydroxybutyrate-co-poly-β-hydroxyvalerate in bacteria and plants |
| US5959091A (en) | 1984-12-10 | 1999-09-28 | Monsanto Company | Truncated gene of Bacillus thuringiensis encoding a polypeptide toxin |
| US5981834A (en) | 1988-10-04 | 1999-11-09 | Monsanto Company | Genetically engineering cotton plants for altered fiber |
| US5998700A (en) | 1996-07-02 | 1999-12-07 | The Board Of Trustees Of Southern Illinois University | Plants containing a bacterial Gdha gene and methods of use thereof |
| US6011199A (en) | 1992-12-15 | 2000-01-04 | Commonwealth Scientific | Method for producing fruiting plants with improved fruit flavour |
| US6013864A (en) | 1993-02-03 | 2000-01-11 | Monsanto Company | Plants resistant to infection by luteoviruses |
| US6015940A (en) | 1992-04-07 | 2000-01-18 | Monsanto Company | Virus resistant potato plants |
| US6023013A (en) | 1997-12-18 | 2000-02-08 | Monsanto Company | Insect-resistant transgenic plants |
| US6063597A (en) | 1997-12-18 | 2000-05-16 | Monsanto Company | Polypeptide compositions toxic to coleopteran insects |
| US6063756A (en) | 1996-09-24 | 2000-05-16 | Monsanto Company | Bacillus thuringiensis cryET33 and cryET34 compositions and uses therefor |
| US6072103A (en) | 1997-11-21 | 2000-06-06 | Calgene Llc | Pathogen and stress-responsive promoter for gene expression |
| US6080560A (en) | 1994-07-25 | 2000-06-27 | Monsanto Company | Method for producing antibodies in plant cells |
| US6093695A (en) | 1996-09-26 | 2000-07-25 | Monsanto Company | Bacillus thuringiensis CryET29 compositions toxic to coleopteran insects and ctenocephalides SPP |
| US6107549A (en) | 1998-03-10 | 2000-08-22 | Monsanto Company | Genetically engineered plant resistance to thiazopyr and other pyridine herbicides |
| US6110464A (en) | 1996-11-20 | 2000-08-29 | Monsanto Company | Broad-spectrum δ-endotoxins |
| US6121436A (en) | 1996-12-13 | 2000-09-19 | Monsanto Company | Antifungal polypeptide and methods for controlling plant pathogenic fungi |
| US6140075A (en) | 1994-07-25 | 2000-10-31 | Monsanto Company | Method for producing antibodies and protein toxins in plant cells |
| US6166292A (en) | 1996-04-26 | 2000-12-26 | Ajinomoto Co., Inc. | Raffinose synthetase gene, method of producing raffinose and transgenic plant |
| US6171640B1 (en) | 1997-04-04 | 2001-01-09 | Monsanto Company | High beta-conglycinin products and their use |
| US6228992B1 (en) | 1998-05-18 | 2001-05-08 | Pioneer Hi-Bred International, Inc. | Proteins for control of nematodes in plants |
| US6228623B1 (en) | 1996-03-13 | 2001-05-08 | Monsanto Company | Polyhydroxyalkanoates of narrow molecular weight distribution prepared in transgenic plants |
| US6242241B1 (en) | 1996-11-20 | 2001-06-05 | Monsanto Company | Polynucleotide compositions encoding broad-spectrum δ-endotoxins |
| US6271443B1 (en) | 1996-10-29 | 2001-08-07 | Calgene Llc | Cotton and rice cellulose synthase DNA sequences |
| USRE37543E1 (en) | 1996-08-13 | 2002-02-05 | Monsanto Company | DNA sequence useful for the production of polyhydroxyalkanoates |
| US6372211B1 (en) | 1997-04-21 | 2002-04-16 | Monsanto Technolgy Llc | Methods and compositions for controlling insects |
| US6380462B1 (en) | 1998-08-14 | 2002-04-30 | Calgene Llc | Method for increasing stearate content in soybean oil |
| US6380466B1 (en) | 1997-05-08 | 2002-04-30 | Calgene Llc | Production of improved rapeseed exhibiting yellow-seed coat |
| US6426447B1 (en) | 1990-11-14 | 2002-07-30 | Monsanto Technology Llc | Plant seed oils |
| US6441277B1 (en) | 1997-06-17 | 2002-08-27 | Monsanto Technology Llc | Expression of fructose 1,6 bisphosphate aldolase in transgenic plants |
| US6444876B1 (en) | 1998-06-05 | 2002-09-03 | Calgene Llc | Acyl CoA: cholesterol acyltransferase related nucleic acid sequences |
| US6448476B1 (en) | 1998-11-17 | 2002-09-10 | Monsanto Technology Llc | Plants and plant cells transformation to express an AMPA-N-acetyltransferase |
| US6459018B1 (en) | 1998-06-12 | 2002-10-01 | Monsanto Technology Llc | Polyunsaturated fatty acids in plants |
| US6468523B1 (en) | 1998-11-02 | 2002-10-22 | Monsanto Technology Llc | Polypeptide compositions toxic to diabrotic insects, and methods of use |
| US6483008B1 (en) | 1990-08-15 | 2002-11-19 | Calgene Llc | Methods for producing plants with elevated oleic acid content |
| US6489461B1 (en) | 1999-06-08 | 2002-12-03 | Calgene Llc | Nucleic acid sequences encoding proteins involved in fatty acid beta-oxidation and methods of use |
| US6495739B1 (en) | 1998-07-24 | 2002-12-17 | Calgene Llc | Plant phosphatidic acid phosphatases |
| US6501009B1 (en) | 1999-08-19 | 2002-12-31 | Monsanto Technology Llc | Expression of Cry3B insecticidal protein in plants |
| US6506962B1 (en) | 1999-05-13 | 2003-01-14 | Monsanto Technology Llc | Acquired resistance genes in plants |
| US6518488B1 (en) | 2000-07-21 | 2003-02-11 | Monsanto Technology Llc | Nucleic acid molecules and other molecules associated with the β-oxidation pathway |
| US6531648B1 (en) | 1998-12-17 | 2003-03-11 | Syngenta Participations Ag | Grain processing method and transgenic plants useful therein |
| US6538181B1 (en) | 1990-06-11 | 2003-03-25 | Calgene Llc | Glycogen biosynthetic enzymes in plants |
| US6537750B1 (en) | 1998-08-04 | 2003-03-25 | Cargill Incorporated | Plant fatty acid desaturase promoters |
| US6538178B1 (en) | 1990-06-18 | 2003-03-25 | Monsanto Technology Llc | Increased starch content in plants |
| US6541259B1 (en) | 1999-04-15 | 2003-04-01 | Calgene Llc | Nucleic acid sequences to proteins involved in isoprenoid synthesis |
| US6555655B1 (en) | 1999-05-04 | 2003-04-29 | Monsanto Technology, Llc | Coleopteran-toxic polypeptide compositions and insect-resistant transgenic plants |
| US6573361B1 (en) | 1999-12-06 | 2003-06-03 | Monsanto Technology Llc | Antifungal proteins and methods for their use |
| US6589767B1 (en) | 1997-04-11 | 2003-07-08 | Abbott Laboratories | Methods and compositions for synthesis of long chain polyunsaturated fatty acids |
| US6593293B1 (en) | 1999-09-15 | 2003-07-15 | Monsanto Technology, Llc | Lepidopteran-active Bacillus thuringiensis δ-endotoxin compositions and methods of use |
| US6596538B1 (en) | 1997-06-05 | 2003-07-22 | Calgene Llc | Fatty acyl-CoA: fatty alcohol acyltransferases |
| US6608241B1 (en) | 1985-10-29 | 2003-08-19 | Monsanto Technology Llc | Protection of plants against viral infection |
| US6617496B1 (en) | 1985-10-16 | 2003-09-09 | Monsanto Company | Effecting virus resistance in plants through the use of negative strand RNAs |
| US6620988B1 (en) | 1997-12-18 | 2003-09-16 | Monsanto Technology, Llc | Coleopteran-resistant transgenic plants and methods of their production using modified Bacillus thuringiensis Cry3Bb nucleic acids |
| US6630306B1 (en) | 1996-12-19 | 2003-10-07 | Yale University | Bioreactive allosteric polynucleotides |
| US6639054B1 (en) | 2000-01-06 | 2003-10-28 | Monsanto Technology Llc | Preparation of deallergenized proteins and permuteins |
| US6653530B1 (en) | 1998-02-13 | 2003-11-25 | Calgene Llc | Methods for producing carotenoid compounds, tocopherol compounds, and specialty oils in plant seeds |
| US6657046B1 (en) | 2000-01-06 | 2003-12-02 | Monsanto Technology Llc | Insect inhibitory lipid acyl hydrolases |
| US6660849B1 (en) | 1997-04-11 | 2003-12-09 | Calgene Llc | Plant fatty acid synthases and use in improved methods for production of medium-chain fatty acids |
| US6663906B2 (en) | 1997-06-17 | 2003-12-16 | Monsanto Technology Llc | Expression of fructose 1,6 bisphosphate aldolase in transgenic plants |
| DE10225066A1 (en) * | 2002-06-06 | 2003-12-18 | Basf Plant Science Gmbh | New transgenic double-stranded RNA, useful for expressing proteins or suppressing genes in plants, contains sequences that allow replication through RNA polymerase |
| USRE38446E1 (en) | 1990-07-20 | 2004-02-24 | Calgene, Llc. | Sucrose phosphate synthase (SPS), its process for preparation its cDNA, and utilization of cDNA to modify the expression of SPS in plant cells |
| US6706950B2 (en) | 2000-07-25 | 2004-03-16 | Calgene Llc | Nucleic acid sequences encoding β-ketoacyl-ACP synthase and uses thereof |
| US6713063B1 (en) | 1996-11-20 | 2004-03-30 | Monsanto Technology, Llc | Broad-spectrum δ-endotoxins |
| US6723897B2 (en) | 1998-08-10 | 2004-04-20 | Monsanto Technology, Llc | Methods for controlling gibberellin levels |
| US6723837B1 (en) | 1999-07-12 | 2004-04-20 | Monsanto Technology Llc | Nucleic acid molecule and encoded protein associated with sterol synthesis and metabolism |
| US6770465B1 (en) | 1999-06-09 | 2004-08-03 | Calgene Llc | Engineering B-ketoacyl ACP synthase for novel substrate specificity |
| US6774283B1 (en) | 1985-07-29 | 2004-08-10 | Calgene Llc | Molecular farming |
| US6803501B2 (en) | 2000-03-09 | 2004-10-12 | Monsanto Technology, Llc | Methods for making plants tolerant to glyphosate and compositions thereof using a DNA encoding an EPSPS enzyme from Eleusine indica |
| US6812379B2 (en) | 1998-07-10 | 2004-11-02 | Calgene Llc | Expression of eukaryotic peptides in plant plastids |
| US6822141B2 (en) | 1998-07-02 | 2004-11-23 | Calgene Llc | Diacylglycerol acyl transferase proteins |
| US6828475B1 (en) | 1994-06-23 | 2004-12-07 | Calgene Llc | Nucleic acid sequences encoding a plant cytoplasmic protein involved in fatty acyl-CoA metabolism |
| WO2005049839A2 (en) * | 2003-11-10 | 2005-06-02 | Icon Genetics Ag | Rna virus-derived plant expression system |
| US6946588B2 (en) | 1996-03-13 | 2005-09-20 | Monsanto Technology Llc | Nucleic acid encoding a modified threonine deaminase and methods of use |
| US6949379B2 (en) | 2000-10-20 | 2005-09-27 | Canji, Inc. | Aptamer-mediated regulation of gene expression |
| US7151204B2 (en) | 2001-01-09 | 2006-12-19 | Monsanto Technology Llc | Maize chloroplast aldolase promoter compositions and methods for use thereof |
| US20070011761A1 (en) | 2005-05-19 | 2007-01-11 | Monsanto Technology, L.L.C. | Post-transcriptional regulation of gene expression |
| US20100311168A1 (en) | 2009-04-07 | 2010-12-09 | Dow Agrosciences Llc | Nanoparticle mediated delivery of sequence specific nucleases |
| US8030473B2 (en) | 2005-01-07 | 2011-10-04 | State Of Oregon Acting By And Through The State Board Of Higher Education On Behalf Of Oregon State University | Method to trigger RNA interference |
| US20120023619A1 (en) | 2010-07-07 | 2012-01-26 | Dow Agrosciences Llc | Linear dna molecule delivery using pegylated quantum dots for stable trasformation in plants |
| US20120244569A1 (en) | 2011-03-23 | 2012-09-27 | Dow Agrosciences Llc | Quantum dot carrier peptide conjugates suitable for imaging and delivery applications in plants |
| US8334430B2 (en) | 2005-10-13 | 2012-12-18 | Monsanto Technology Llc | Methods for producing hybrid seed |
| US8395023B2 (en) | 2004-12-21 | 2013-03-12 | Monsanto Technology Llc | Recombinant DNA constructs and methods for controlling gene expression |
| US8404928B2 (en) | 2006-08-31 | 2013-03-26 | Monsanto Technology Llc | Phased small RNAs |
| US8404927B2 (en) | 2004-12-21 | 2013-03-26 | Monsanto Technology Llc | Double-stranded RNA stabilized in planta |
| US8410334B2 (en) | 2007-02-20 | 2013-04-02 | Monsanto Technology Llc | Invertebrate microRNAs |
| US20130102651A1 (en) | 2007-08-28 | 2013-04-25 | California Institute Of Technology | General composition framework for ligand-controlled rna regulatory systems |
| US8455716B2 (en) | 2009-04-20 | 2013-06-04 | Monsanto Technology Llc | Multiple virus resistance in plants |
| US20130145488A1 (en) | 2011-12-06 | 2013-06-06 | Iowa State University Research Foundation, Inc. | Mesoporous silica nanoparticles suitable for co-delivery |
| US20130185823A1 (en) | 2012-01-16 | 2013-07-18 | Academia Sinica | Mesoporous silica nanoparticle-mediated delivery of dna into arabidopsis root |
| US20140096284A1 (en) | 2012-10-01 | 2014-04-03 | Iowa State University Research Foundation, Inc. | Method for the delivery of molecules lyophilized onto microparticles to plant tissues |
| US20140356414A1 (en) | 2013-06-03 | 2014-12-04 | University Of Southern California | Targeted Crosslinked Multilamellar Liposomes |
| US8946511B2 (en) | 2006-10-12 | 2015-02-03 | Monsanto Technology Llc | Plant microRNAs and methods of use thereof |
| US20150040268A1 (en) | 2013-04-25 | 2015-02-05 | Morflora Israel Ltd | Methods and compositions for the delivery of nucleic acids to seeds |
| US20150047074A1 (en) | 2013-08-09 | 2015-02-12 | Massachusetts Institute Of Technology | Nanobionic engineering of organelles and photosynthetic organisms |
| US20150082478A1 (en) | 2013-08-22 | 2015-03-19 | E I Du Pont De Nemours And Company | Plant genome modification using guide rna/cas endonuclease systems and methods of use |
| US9040774B2 (en) | 2008-07-01 | 2015-05-26 | Monsanto Technology Llc | Recombinant DNA constructs encoding ribonuclease cleavage blockers and methods for modulating expression of a target gene |
| US20150208663A1 (en) | 2013-10-15 | 2015-07-30 | Board Of Trustees Of The University Of Arkansas | Method of delivery of bio-active agents to plant cells by using nano-sized materials as carriers |
| US9139838B2 (en) | 2011-07-01 | 2015-09-22 | Monsanto Technology Llc | Methods and compositions for selective regulation of protein expression |
| US9777288B2 (en) | 2013-07-19 | 2017-10-03 | Monsanto Technology Llc | Compositions and methods for controlling leptinotarsa |
| US20170369898A1 (en) | 2014-12-19 | 2017-12-28 | AgBiome, Inc. | Sequences to facilitate incorporation of dna into the genome of an organism |
| US9873888B2 (en) | 2009-10-23 | 2018-01-23 | Monsanto Technology Llc | Transgenic soybean plants and chromosomes |
| US9976152B2 (en) | 2007-06-26 | 2018-05-22 | Monsanto Technology Llc | Temporal regulation of gene expression by microRNAs |
| US10017549B2 (en) | 2008-08-29 | 2018-07-10 | Monsanto Technology Llc | Hemipteran and coleopteran active toxin proteins from Bacillus thuringiensis |
| US20180312854A1 (en) | 2006-05-16 | 2018-11-01 | Monsanto Technology Llc | Use of non-agrobacterium bacterial species for plant transformation |
| US20190032131A1 (en) | 2016-12-12 | 2019-01-31 | Integrated Dna Technologies, Inc. | Genome editing detection |
| US10233217B2 (en) | 2014-10-16 | 2019-03-19 | Monsanto Technology Llc | Chimeric insecticidal proteins toxic or inhibitory to Lepidopteran pests |
| US10378012B2 (en) | 2014-07-29 | 2019-08-13 | Monsanto Technology Llc | Compositions and methods for controlling insect pests |
| US20190352655A1 (en) | 2017-01-28 | 2019-11-21 | Inari Agriculture, Inc. | Novel plant cells, plants, and seeds |
| US10487123B2 (en) | 2014-10-16 | 2019-11-26 | Monsanto Technology Llc | Chimeric insecticidal proteins toxic or inhibitory to lepidopteran pests |
| US10612037B2 (en) | 2016-06-20 | 2020-04-07 | Monsanto Technology Llc | Insecticidal proteins toxic or inhibitory to hemipteran pests |
| US10827755B2 (en) | 2015-11-18 | 2020-11-10 | Monsanto Technology Llc | Insecticidal compositions and methods |
| US11091770B2 (en) | 2014-04-01 | 2021-08-17 | Monsanto Technology Llc | Compositions and methods for controlling insect pests |
| US20210259176A1 (en) | 2004-08-26 | 2021-08-26 | Monsanto Technology Llc | Methods of Seed Breeding Using High Throughput Nondestructive Seed Sampling |
| US11130965B2 (en) | 2016-10-27 | 2021-09-28 | Syngenta Participations Ag | Insecticidal proteins |
| US11136593B2 (en) | 2016-09-09 | 2021-10-05 | Syngenta Participations Ag | Insecticidal proteins |
| US11180774B2 (en) | 2017-01-12 | 2021-11-23 | Syngenta Participations Ag | Insecticidal proteins |
| US11186837B2 (en) | 2004-04-09 | 2021-11-30 | Monsanto Technology Llc | Compositions and methods for controlling Diabrotica |
| WO2021257987A2 (en) * | 2020-06-20 | 2021-12-23 | Halo-Bio Rnai Therapeutics, Inc. | Methods and compositions of rna nanostructures for replication and subgenomic expression by rna-directed rna polymerase |
| US20220221377A1 (en) | 2006-03-02 | 2022-07-14 | Monsanto Technology Llc | Automated Contamination-Free Seed Sampler And Methods Of Sampling, Testing And Bulking Seeds |
| WO2023004435A1 (en) | 2021-07-23 | 2023-01-26 | Flagship Pioneering Innovations Vii, Llc | Compositions for fungal control and related methods |
| WO2023161802A1 (en) | 2022-02-22 | 2023-08-31 | Invaio Sciences International Gmbh | Methods and systems for controlling xylella fastidiosa in olive trees |
| WO2023240208A1 (en) | 2022-06-09 | 2023-12-14 | Invaio Sciences, Inc. | Controlling citrus greening in citrus plants using oxytetracycline |
| US11844318B2 (en) | 2018-07-25 | 2023-12-19 | Invaio Sciences International Gmbh | Injection systems, injection tools and methods for same |
-
2024
- 2024-05-03 AR ARP240101121A patent/AR132586A1/en unknown
- 2024-05-03 WO PCT/US2024/027769 patent/WO2024229398A1/en active Pending
Patent Citations (190)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US5858742A (en) | 1983-01-17 | 1999-01-12 | Monsanto Company | Chimeric genes for transforming plant cells using viral promoters |
| US5959091A (en) | 1984-12-10 | 1999-09-28 | Monsanto Company | Truncated gene of Bacillus thuringiensis encoding a polypeptide toxin |
| US6774283B1 (en) | 1985-07-29 | 2004-08-10 | Calgene Llc | Molecular farming |
| US6617496B1 (en) | 1985-10-16 | 2003-09-09 | Monsanto Company | Effecting virus resistance in plants through the use of negative strand RNAs |
| US6608241B1 (en) | 1985-10-29 | 2003-08-19 | Monsanto Technology Llc | Protection of plants against viral infection |
| US5380831A (en) | 1986-04-04 | 1995-01-10 | Mycogen Plant Science, Inc. | Synthetic insecticidal crystal protein gene |
| US5322938A (en) | 1987-01-13 | 1994-06-21 | Monsanto Company | DNA sequence for enhancing the efficiency of transcription |
| US5763241A (en) | 1987-04-29 | 1998-06-09 | Monsanto Company | Insect resistant plants |
| US6284949B1 (en) | 1987-04-29 | 2001-09-04 | Monsanto Company | Insect-resistant plants comprising a Bacillus thuringiensis gene |
| US5229114A (en) | 1987-08-20 | 1993-07-20 | The United States Of America As Represented By The Secretary Of Agriculture | Approaches useful for the control of root nodulation of leguminous plants |
| US5981834A (en) | 1988-10-04 | 1999-11-09 | Monsanto Company | Genetically engineering cotton plants for altered fiber |
| US7741118B1 (en) | 1989-02-24 | 2010-06-22 | Monsanto Technology Llc | Synthetic plant genes and method for preparation |
| US5880275A (en) | 1989-02-24 | 1999-03-09 | Monsanto Company | Synthetic plant genes from BT kurstaki and method for preparation |
| US5689041A (en) | 1989-08-10 | 1997-11-18 | Plant Gentic Systems N.V. | Plants modified with barstar for fertility restoration |
| US5641876A (en) | 1990-01-05 | 1997-06-24 | Cornell Research Foundation, Inc. | Rice actin gene and promoter |
| US5543576A (en) | 1990-03-23 | 1996-08-06 | Mogen International | Production of enzymes in seeds and their use |
| US6538181B1 (en) | 1990-06-11 | 2003-03-25 | Calgene Llc | Glycogen biosynthetic enzymes in plants |
| US6538178B1 (en) | 1990-06-18 | 2003-03-25 | Monsanto Technology Llc | Increased starch content in plants |
| US5608149A (en) | 1990-06-18 | 1997-03-04 | Monsanto Company | Enhanced starch biosynthesis in tomatoes |
| US6538179B1 (en) | 1990-06-18 | 2003-03-25 | Monsanto Technology Llc | Enhanced starch biosynthesis in seeds |
| US5463175A (en) | 1990-06-25 | 1995-10-31 | Monsanto Company | Glyphosate tolerant plants |
| USRE38446E1 (en) | 1990-07-20 | 2004-02-24 | Calgene, Llc. | Sucrose phosphate synthase (SPS), its process for preparation its cDNA, and utilization of cDNA to modify the expression of SPS in plant cells |
| US6483008B1 (en) | 1990-08-15 | 2002-11-19 | Calgene Llc | Methods for producing plants with elevated oleic acid content |
| US5804425A (en) | 1990-08-31 | 1998-09-08 | Monsanto Company | Glyphosate-tolerant 5-enolpyruvylshikimate-3-phosphate synthases |
| US6248876B1 (en) | 1990-08-31 | 2001-06-19 | Monsanto Company | Glyphosate-tolerant 5-enolpyruvylshikimate-3-phosphate synthases |
| US5633435A (en) | 1990-08-31 | 1997-05-27 | Monsanto Company | Glyphosate-tolerant 5-enolpyruvylshikimate-3-phosphate synthases |
| US5866775A (en) | 1990-09-28 | 1999-02-02 | Monsanto Company | Glyphosate-tolerant 5-enolpyruvyl-3-phosphoshikimate synthases |
| US6225114B1 (en) | 1990-09-28 | 2001-05-01 | Monsanto Company | Modified gene encoding glyphosate-tolerant 5-enolpruvyl-3-phosphoshikimate synthase |
| US6426447B1 (en) | 1990-11-14 | 2002-07-30 | Monsanto Technology Llc | Plant seed oils |
| US5512466A (en) | 1990-12-26 | 1996-04-30 | Monsanto Company | Control of fruit ripening and senescence in plants |
| US5304730A (en) | 1991-09-03 | 1994-04-19 | Monsanto Company | Virus resistant plants and method therefore |
| US5763245A (en) | 1991-09-23 | 1998-06-09 | Monsanto Company | Method of controlling insects |
| US6015940A (en) | 1992-04-07 | 2000-01-18 | Monsanto Company | Virus resistant potato plants |
| US5850023A (en) | 1992-11-30 | 1998-12-15 | Monsanto Company | Modified plant viral replicase genes |
| US6011199A (en) | 1992-12-15 | 2000-01-04 | Commonwealth Scientific | Method for producing fruiting plants with improved fruit flavour |
| US6013864A (en) | 1993-02-03 | 2000-01-11 | Monsanto Company | Plants resistant to infection by luteoviruses |
| US5942658A (en) | 1993-07-29 | 1999-08-24 | Monsanto Company | Transformed plant with Bacillus thuringiensis toxin gene |
| US5869720A (en) | 1993-09-30 | 1999-02-09 | Monsanto Company | Transgenic cotton plants producing heterologous peroxidase |
| US5516671A (en) | 1993-11-24 | 1996-05-14 | Monsanto Company | Method of controlling plant pathogens |
| US5689052A (en) | 1993-12-22 | 1997-11-18 | Monsanto Company | Synthetic DNA sequences having enhanced expression in monocotyledonous plants and method for preparation thereof |
| US6828475B1 (en) | 1994-06-23 | 2004-12-07 | Calgene Llc | Nucleic acid sequences encoding a plant cytoplasmic protein involved in fatty acyl-CoA metabolism |
| US6140075A (en) | 1994-07-25 | 2000-10-31 | Monsanto Company | Method for producing antibodies and protein toxins in plant cells |
| US6080560A (en) | 1994-07-25 | 2000-06-27 | Monsanto Company | Method for producing antibodies in plant cells |
| US5750876A (en) | 1994-07-28 | 1998-05-12 | Monsanto Company | Isoamylase gene, compositions containing it, and methods of using isoamylases |
| US6476295B2 (en) | 1995-02-10 | 2002-11-05 | Monsanto Technology, Llc | Expression of sucrose phosphorylase in plants |
| US5716837A (en) | 1995-02-10 | 1998-02-10 | Monsanto Company | Expression of sucrose phosphorylase in plants |
| US6235971B1 (en) | 1995-02-10 | 2001-05-22 | Monsanto Company | Expression of sucrose phoshorylase in plants |
| US6222098B1 (en) | 1995-02-10 | 2001-04-24 | Monsanto Company | Expression of sucrose phosphorylase in plants |
| US6946588B2 (en) | 1996-03-13 | 2005-09-20 | Monsanto Technology Llc | Nucleic acid encoding a modified threonine deaminase and methods of use |
| US6228623B1 (en) | 1996-03-13 | 2001-05-08 | Monsanto Company | Polyhydroxyalkanoates of narrow molecular weight distribution prepared in transgenic plants |
| US5958745A (en) | 1996-03-13 | 1999-09-28 | Monsanto Company | Methods of optimizing substrate pools and biosynthesis of poly-β-hydroxybutyrate-co-poly-β-hydroxyvalerate in bacteria and plants |
| US5773696A (en) | 1996-03-29 | 1998-06-30 | Monsanto Company | Antifungal polypeptide and methods for controlling plant pathogenic fungi |
| US6215048B1 (en) | 1996-03-29 | 2001-04-10 | Monsanto Company | Nucleic acid sequences encoding an antifungal polypeptide, aly AFP from alyssum and methods for their use |
| US6653280B2 (en) | 1996-03-29 | 2003-11-25 | Monsanto Technology Llc | Antifungal polypeptide AlyAFP from Alyssum and methods for controlling plant pathogenic fungi |
| US6166292A (en) | 1996-04-26 | 2000-12-26 | Ajinomoto Co., Inc. | Raffinose synthetase gene, method of producing raffinose and transgenic plant |
| US5998700A (en) | 1996-07-02 | 1999-12-07 | The Board Of Trustees Of Southern Illinois University | Plants containing a bacterial Gdha gene and methods of use thereof |
| USRE37543E1 (en) | 1996-08-13 | 2002-02-05 | Monsanto Company | DNA sequence useful for the production of polyhydroxyalkanoates |
| US6326351B1 (en) | 1996-09-24 | 2001-12-04 | Monsanto Technology Llc | Bacillus thuringiensis CryET33 and CryET34 compositions and uses therefor |
| US6248536B1 (en) | 1996-09-24 | 2001-06-19 | Monsanto Company | Bacillus thuringiensis CryET33 and CryET34 compositions and uses thereof |
| US6063756A (en) | 1996-09-24 | 2000-05-16 | Monsanto Company | Bacillus thuringiensis cryET33 and cryET34 compositions and uses therefor |
| US6399330B1 (en) | 1996-09-24 | 2002-06-04 | Monsanto Technology Llc | Bacillus thuringiensis cryet33 and cryet34 compositions and uses thereof |
| US6686452B2 (en) | 1996-09-26 | 2004-02-03 | Monsanto Technology Llc | Bacillus thuringiensis CryET29 compositions toxic to coleopteran insects and ctenocephalides SPP |
| US6537756B1 (en) | 1996-09-26 | 2003-03-25 | Monsanto Technology, Llc | Bacillus thuringiensis CryET29 compositions toxic to coleopteran insects and Ctenocephalides SPP |
| US6093695A (en) | 1996-09-26 | 2000-07-25 | Monsanto Company | Bacillus thuringiensis CryET29 compositions toxic to coleopteran insects and ctenocephalides SPP |
| US6271443B1 (en) | 1996-10-29 | 2001-08-07 | Calgene Llc | Cotton and rice cellulose synthase DNA sequences |
| US6576818B1 (en) | 1996-10-29 | 2003-06-10 | Calgene Llc | Plant cellulose synthase and promoter sequences |
| US6713063B1 (en) | 1996-11-20 | 2004-03-30 | Monsanto Technology, Llc | Broad-spectrum δ-endotoxins |
| US6221649B1 (en) | 1996-11-20 | 2001-04-24 | Monsanto Company | Chimeric bacillus thuringiensis-endotoxins and host cells expressing same |
| US6242241B1 (en) | 1996-11-20 | 2001-06-05 | Monsanto Company | Polynucleotide compositions encoding broad-spectrum δ-endotoxins |
| US6538109B2 (en) | 1996-11-20 | 2003-03-25 | Monsanto Technology, Llc | Polynucleotide compositions encoding broad spectrum delta-endotoxins |
| US6521442B2 (en) | 1996-11-20 | 2003-02-18 | Monsanto Technology Llc | Polynucleotide compositions encoding broad spectrum δ-endotoxins |
| US6110464A (en) | 1996-11-20 | 2000-08-29 | Monsanto Company | Broad-spectrum δ-endotoxins |
| US6281016B1 (en) | 1996-11-20 | 2001-08-28 | Monsanto Company | Broad-spectrum insect resistant transgenic plants |
| US6645497B2 (en) | 1996-11-20 | 2003-11-11 | Monsanto Technology, Llc | Polynucleotide compositions encoding broad-spectrum δ endotoxins |
| US6156573A (en) | 1996-11-20 | 2000-12-05 | Monsanto Company | Hybrid Bacillus thuringiensis δ-endotoxins with novel broad-spectrum insecticidal activity |
| US6177615B1 (en) | 1996-11-27 | 2001-01-23 | Monsanto Company | Lepidopteran-toxic polypeptide and polynucleotide compositions and methods for making and using same |
| US6153814A (en) | 1996-11-27 | 2000-11-28 | Monsanto Company | Polypeptide compositions toxic to lepidopteran insects and methods for making same |
| US6809078B2 (en) | 1996-11-27 | 2004-10-26 | Monsanto Technology Llc | Compositions encoding lepidopteran-toxic polypeptides and methods of use |
| US6423828B1 (en) | 1996-11-27 | 2002-07-23 | Monsanto Technology Llc | Nuclei acid and polypeptide compositions encoding lepidopteran-toxic polypeptides |
| US6313378B1 (en) | 1996-11-27 | 2001-11-06 | Monsanto Technology Llc | Lepidopteran-resistent transgenic plants |
| US5942664A (en) | 1996-11-27 | 1999-08-24 | Ecogen, Inc. | Bacillus thuringiensis Cry1C compositions toxic to lepidopteran insects and methods for making Cry1C mutants |
| US6121436A (en) | 1996-12-13 | 2000-09-19 | Monsanto Company | Antifungal polypeptide and methods for controlling plant pathogenic fungi |
| US6316407B1 (en) | 1996-12-13 | 2001-11-13 | Monsanto Company | Antifungal polypeptide from alfalfa and methods for controlling plant pathogenic fungi |
| US6630306B1 (en) | 1996-12-19 | 2003-10-07 | Yale University | Bioreactive allosteric polynucleotides |
| US6171640B1 (en) | 1997-04-04 | 2001-01-09 | Monsanto Company | High beta-conglycinin products and their use |
| US6589767B1 (en) | 1997-04-11 | 2003-07-08 | Abbott Laboratories | Methods and compositions for synthesis of long chain polyunsaturated fatty acids |
| US6660849B1 (en) | 1997-04-11 | 2003-12-09 | Calgene Llc | Plant fatty acid synthases and use in improved methods for production of medium-chain fatty acids |
| US6372211B1 (en) | 1997-04-21 | 2002-04-16 | Monsanto Technolgy Llc | Methods and compositions for controlling insects |
| US6380466B1 (en) | 1997-05-08 | 2002-04-30 | Calgene Llc | Production of improved rapeseed exhibiting yellow-seed coat |
| US6596538B1 (en) | 1997-06-05 | 2003-07-22 | Calgene Llc | Fatty acyl-CoA: fatty alcohol acyltransferases |
| US6441277B1 (en) | 1997-06-17 | 2002-08-27 | Monsanto Technology Llc | Expression of fructose 1,6 bisphosphate aldolase in transgenic plants |
| US6716474B2 (en) | 1997-06-17 | 2004-04-06 | Monsanto Technology Llc | Expression of fructose 1,6 bisphosphate aldolase in transgenic plants |
| US6663906B2 (en) | 1997-06-17 | 2003-12-16 | Monsanto Technology Llc | Expression of fructose 1,6 bisphosphate aldolase in transgenic plants |
| US6072103A (en) | 1997-11-21 | 2000-06-06 | Calgene Llc | Pathogen and stress-responsive promoter for gene expression |
| US6063597A (en) | 1997-12-18 | 2000-05-16 | Monsanto Company | Polypeptide compositions toxic to coleopteran insects |
| US6023013A (en) | 1997-12-18 | 2000-02-08 | Monsanto Company | Insect-resistant transgenic plants |
| US6642030B1 (en) | 1997-12-18 | 2003-11-04 | Monsanto Technology, Llc | Nucleic acid compositions encoding modified Bacillus thuringiensis coleopteran-toxic crystal proteins |
| US6620988B1 (en) | 1997-12-18 | 2003-09-16 | Monsanto Technology, Llc | Coleopteran-resistant transgenic plants and methods of their production using modified Bacillus thuringiensis Cry3Bb nucleic acids |
| US6653530B1 (en) | 1998-02-13 | 2003-11-25 | Calgene Llc | Methods for producing carotenoid compounds, tocopherol compounds, and specialty oils in plant seeds |
| US6107549A (en) | 1998-03-10 | 2000-08-22 | Monsanto Company | Genetically engineered plant resistance to thiazopyr and other pyridine herbicides |
| US6228992B1 (en) | 1998-05-18 | 2001-05-08 | Pioneer Hi-Bred International, Inc. | Proteins for control of nematodes in plants |
| US6444876B1 (en) | 1998-06-05 | 2002-09-03 | Calgene Llc | Acyl CoA: cholesterol acyltransferase related nucleic acid sequences |
| US6459018B1 (en) | 1998-06-12 | 2002-10-01 | Monsanto Technology Llc | Polyunsaturated fatty acids in plants |
| US6822141B2 (en) | 1998-07-02 | 2004-11-23 | Calgene Llc | Diacylglycerol acyl transferase proteins |
| US6812379B2 (en) | 1998-07-10 | 2004-11-02 | Calgene Llc | Expression of eukaryotic peptides in plant plastids |
| US6495739B1 (en) | 1998-07-24 | 2002-12-17 | Calgene Llc | Plant phosphatidic acid phosphatases |
| US6537750B1 (en) | 1998-08-04 | 2003-03-25 | Cargill Incorporated | Plant fatty acid desaturase promoters |
| US6723897B2 (en) | 1998-08-10 | 2004-04-20 | Monsanto Technology, Llc | Methods for controlling gibberellin levels |
| US6380462B1 (en) | 1998-08-14 | 2002-04-30 | Calgene Llc | Method for increasing stearate content in soybean oil |
| US6468523B1 (en) | 1998-11-02 | 2002-10-22 | Monsanto Technology Llc | Polypeptide compositions toxic to diabrotic insects, and methods of use |
| US6448476B1 (en) | 1998-11-17 | 2002-09-10 | Monsanto Technology Llc | Plants and plant cells transformation to express an AMPA-N-acetyltransferase |
| US6531648B1 (en) | 1998-12-17 | 2003-03-11 | Syngenta Participations Ag | Grain processing method and transgenic plants useful therein |
| US6541259B1 (en) | 1999-04-15 | 2003-04-01 | Calgene Llc | Nucleic acid sequences to proteins involved in isoprenoid synthesis |
| US6555655B1 (en) | 1999-05-04 | 2003-04-29 | Monsanto Technology, Llc | Coleopteran-toxic polypeptide compositions and insect-resistant transgenic plants |
| US6506962B1 (en) | 1999-05-13 | 2003-01-14 | Monsanto Technology Llc | Acquired resistance genes in plants |
| US6489461B1 (en) | 1999-06-08 | 2002-12-03 | Calgene Llc | Nucleic acid sequences encoding proteins involved in fatty acid beta-oxidation and methods of use |
| US6770465B1 (en) | 1999-06-09 | 2004-08-03 | Calgene Llc | Engineering B-ketoacyl ACP synthase for novel substrate specificity |
| US6723837B1 (en) | 1999-07-12 | 2004-04-20 | Monsanto Technology Llc | Nucleic acid molecule and encoded protein associated with sterol synthesis and metabolism |
| US6501009B1 (en) | 1999-08-19 | 2002-12-31 | Monsanto Technology Llc | Expression of Cry3B insecticidal protein in plants |
| US6593293B1 (en) | 1999-09-15 | 2003-07-15 | Monsanto Technology, Llc | Lepidopteran-active Bacillus thuringiensis δ-endotoxin compositions and methods of use |
| US6573361B1 (en) | 1999-12-06 | 2003-06-03 | Monsanto Technology Llc | Antifungal proteins and methods for their use |
| US6639054B1 (en) | 2000-01-06 | 2003-10-28 | Monsanto Technology Llc | Preparation of deallergenized proteins and permuteins |
| US6657046B1 (en) | 2000-01-06 | 2003-12-02 | Monsanto Technology Llc | Insect inhibitory lipid acyl hydrolases |
| US6803501B2 (en) | 2000-03-09 | 2004-10-12 | Monsanto Technology, Llc | Methods for making plants tolerant to glyphosate and compositions thereof using a DNA encoding an EPSPS enzyme from Eleusine indica |
| US6518488B1 (en) | 2000-07-21 | 2003-02-11 | Monsanto Technology Llc | Nucleic acid molecules and other molecules associated with the β-oxidation pathway |
| US6706950B2 (en) | 2000-07-25 | 2004-03-16 | Calgene Llc | Nucleic acid sequences encoding β-ketoacyl-ACP synthase and uses thereof |
| US6949379B2 (en) | 2000-10-20 | 2005-09-27 | Canji, Inc. | Aptamer-mediated regulation of gene expression |
| US7151204B2 (en) | 2001-01-09 | 2006-12-19 | Monsanto Technology Llc | Maize chloroplast aldolase promoter compositions and methods for use thereof |
| DE10225066A1 (en) * | 2002-06-06 | 2003-12-18 | Basf Plant Science Gmbh | New transgenic double-stranded RNA, useful for expressing proteins or suppressing genes in plants, contains sequences that allow replication through RNA polymerase |
| WO2005049839A2 (en) * | 2003-11-10 | 2005-06-02 | Icon Genetics Ag | Rna virus-derived plant expression system |
| US11186837B2 (en) | 2004-04-09 | 2021-11-30 | Monsanto Technology Llc | Compositions and methods for controlling Diabrotica |
| US20210259176A1 (en) | 2004-08-26 | 2021-08-26 | Monsanto Technology Llc | Methods of Seed Breeding Using High Throughput Nondestructive Seed Sampling |
| US9708620B2 (en) | 2004-12-21 | 2017-07-18 | Monsanto Technology Llc | Recombinant DNA constructs and methods for controlling gene expression |
| US8395023B2 (en) | 2004-12-21 | 2013-03-12 | Monsanto Technology Llc | Recombinant DNA constructs and methods for controlling gene expression |
| US8404927B2 (en) | 2004-12-21 | 2013-03-26 | Monsanto Technology Llc | Double-stranded RNA stabilized in planta |
| US10793869B2 (en) | 2004-12-21 | 2020-10-06 | Monsanto Technology Llc | Recombinant DNA constructs and methods for controlling gene expression |
| US9018002B2 (en) | 2005-01-07 | 2015-04-28 | State Of Oregon Acting By And Through The State Board Of Higher Education On Behalf Of Oregon State University | Method to trigger RNA interference |
| US8030473B2 (en) | 2005-01-07 | 2011-10-04 | State Of Oregon Acting By And Through The State Board Of Higher Education On Behalf Of Oregon State University | Method to trigger RNA interference |
| US8816061B2 (en) | 2005-01-07 | 2014-08-26 | State Of Oregon Acting By And Through The State Board Of Higher Education On Behalf Of Oregon State University | Method to trigger RNA interference |
| US8476422B2 (en) | 2005-01-07 | 2013-07-02 | State of Oregon acting by and through the State Board of Higher Eduction on behalf of Oregon State University | Method to trigger RNA interference |
| US20070011761A1 (en) | 2005-05-19 | 2007-01-11 | Monsanto Technology, L.L.C. | Post-transcriptional regulation of gene expression |
| US8334430B2 (en) | 2005-10-13 | 2012-12-18 | Monsanto Technology Llc | Methods for producing hybrid seed |
| US10876126B2 (en) | 2005-10-13 | 2020-12-29 | Monsanto Technology Llc | Methods for producing hybrid seed |
| US20220221377A1 (en) | 2006-03-02 | 2022-07-14 | Monsanto Technology Llc | Automated Contamination-Free Seed Sampler And Methods Of Sampling, Testing And Bulking Seeds |
| US20180312854A1 (en) | 2006-05-16 | 2018-11-01 | Monsanto Technology Llc | Use of non-agrobacterium bacterial species for plant transformation |
| US8404928B2 (en) | 2006-08-31 | 2013-03-26 | Monsanto Technology Llc | Phased small RNAs |
| US8946511B2 (en) | 2006-10-12 | 2015-02-03 | Monsanto Technology Llc | Plant microRNAs and methods of use thereof |
| US8410334B2 (en) | 2007-02-20 | 2013-04-02 | Monsanto Technology Llc | Invertebrate microRNAs |
| US9976152B2 (en) | 2007-06-26 | 2018-05-22 | Monsanto Technology Llc | Temporal regulation of gene expression by microRNAs |
| US20130102651A1 (en) | 2007-08-28 | 2013-04-25 | California Institute Of Technology | General composition framework for ligand-controlled rna regulatory systems |
| US9040774B2 (en) | 2008-07-01 | 2015-05-26 | Monsanto Technology Llc | Recombinant DNA constructs encoding ribonuclease cleavage blockers and methods for modulating expression of a target gene |
| US10017549B2 (en) | 2008-08-29 | 2018-07-10 | Monsanto Technology Llc | Hemipteran and coleopteran active toxin proteins from Bacillus thuringiensis |
| US20100311168A1 (en) | 2009-04-07 | 2010-12-09 | Dow Agrosciences Llc | Nanoparticle mediated delivery of sequence specific nucleases |
| US8455716B2 (en) | 2009-04-20 | 2013-06-04 | Monsanto Technology Llc | Multiple virus resistance in plants |
| US9873888B2 (en) | 2009-10-23 | 2018-01-23 | Monsanto Technology Llc | Transgenic soybean plants and chromosomes |
| US20120023619A1 (en) | 2010-07-07 | 2012-01-26 | Dow Agrosciences Llc | Linear dna molecule delivery using pegylated quantum dots for stable trasformation in plants |
| US20120244569A1 (en) | 2011-03-23 | 2012-09-27 | Dow Agrosciences Llc | Quantum dot carrier peptide conjugates suitable for imaging and delivery applications in plants |
| US9139838B2 (en) | 2011-07-01 | 2015-09-22 | Monsanto Technology Llc | Methods and compositions for selective regulation of protein expression |
| US20130145488A1 (en) | 2011-12-06 | 2013-06-06 | Iowa State University Research Foundation, Inc. | Mesoporous silica nanoparticles suitable for co-delivery |
| US20130185823A1 (en) | 2012-01-16 | 2013-07-18 | Academia Sinica | Mesoporous silica nanoparticle-mediated delivery of dna into arabidopsis root |
| US20140096284A1 (en) | 2012-10-01 | 2014-04-03 | Iowa State University Research Foundation, Inc. | Method for the delivery of molecules lyophilized onto microparticles to plant tissues |
| US20150040268A1 (en) | 2013-04-25 | 2015-02-05 | Morflora Israel Ltd | Methods and compositions for the delivery of nucleic acids to seeds |
| US20140356414A1 (en) | 2013-06-03 | 2014-12-04 | University Of Southern California | Targeted Crosslinked Multilamellar Liposomes |
| US9777288B2 (en) | 2013-07-19 | 2017-10-03 | Monsanto Technology Llc | Compositions and methods for controlling leptinotarsa |
| US20150047074A1 (en) | 2013-08-09 | 2015-02-12 | Massachusetts Institute Of Technology | Nanobionic engineering of organelles and photosynthetic organisms |
| US20150082478A1 (en) | 2013-08-22 | 2015-03-19 | E I Du Pont De Nemours And Company | Plant genome modification using guide rna/cas endonuclease systems and methods of use |
| US20150208663A1 (en) | 2013-10-15 | 2015-07-30 | Board Of Trustees Of The University Of Arkansas | Method of delivery of bio-active agents to plant cells by using nano-sized materials as carriers |
| US11091770B2 (en) | 2014-04-01 | 2021-08-17 | Monsanto Technology Llc | Compositions and methods for controlling insect pests |
| US10378012B2 (en) | 2014-07-29 | 2019-08-13 | Monsanto Technology Llc | Compositions and methods for controlling insect pests |
| US10494408B2 (en) | 2014-10-16 | 2019-12-03 | Monsanto Technology Llc | Chimeric insecticidal proteins toxic or inhibitory to lepidopteran pests |
| US10233217B2 (en) | 2014-10-16 | 2019-03-19 | Monsanto Technology Llc | Chimeric insecticidal proteins toxic or inhibitory to Lepidopteran pests |
| US10611806B2 (en) | 2014-10-16 | 2020-04-07 | Monsanto Technology Llc | Chimeric insecticidal proteins toxic or inhibitory to Lepidopteran pests |
| US10669317B2 (en) | 2014-10-16 | 2020-06-02 | Monsanto Technology Llc | Chimeric insecticidal proteins toxic or inhibitory to lepidopteran pests |
| US10494409B2 (en) | 2014-10-16 | 2019-12-03 | Monsanto Technology Llc | Chimeric insecticidal proteins toxic or inhibitory to lepidopteran pests |
| US11267849B2 (en) | 2014-10-16 | 2022-03-08 | Monsanto Technology Llc | Chimeric insecticidal proteins toxic or inhibitory to lepidopteran pests |
| US10487123B2 (en) | 2014-10-16 | 2019-11-26 | Monsanto Technology Llc | Chimeric insecticidal proteins toxic or inhibitory to lepidopteran pests |
| US20170369898A1 (en) | 2014-12-19 | 2017-12-28 | AgBiome, Inc. | Sequences to facilitate incorporation of dna into the genome of an organism |
| US10827755B2 (en) | 2015-11-18 | 2020-11-10 | Monsanto Technology Llc | Insecticidal compositions and methods |
| US11254950B2 (en) | 2016-06-20 | 2022-02-22 | Monsanto Technology Llc | Insecticidal proteins toxic or inhibitory to hemtpteran pests |
| US10612037B2 (en) | 2016-06-20 | 2020-04-07 | Monsanto Technology Llc | Insecticidal proteins toxic or inhibitory to hemipteran pests |
| US11136593B2 (en) | 2016-09-09 | 2021-10-05 | Syngenta Participations Ag | Insecticidal proteins |
| US11130965B2 (en) | 2016-10-27 | 2021-09-28 | Syngenta Participations Ag | Insecticidal proteins |
| US20190032131A1 (en) | 2016-12-12 | 2019-01-31 | Integrated Dna Technologies, Inc. | Genome editing detection |
| US11180774B2 (en) | 2017-01-12 | 2021-11-23 | Syngenta Participations Ag | Insecticidal proteins |
| US20190352655A1 (en) | 2017-01-28 | 2019-11-21 | Inari Agriculture, Inc. | Novel plant cells, plants, and seeds |
| US11844318B2 (en) | 2018-07-25 | 2023-12-19 | Invaio Sciences International Gmbh | Injection systems, injection tools and methods for same |
| WO2021257987A2 (en) * | 2020-06-20 | 2021-12-23 | Halo-Bio Rnai Therapeutics, Inc. | Methods and compositions of rna nanostructures for replication and subgenomic expression by rna-directed rna polymerase |
| WO2023004435A1 (en) | 2021-07-23 | 2023-01-26 | Flagship Pioneering Innovations Vii, Llc | Compositions for fungal control and related methods |
| WO2023161802A1 (en) | 2022-02-22 | 2023-08-31 | Invaio Sciences International Gmbh | Methods and systems for controlling xylella fastidiosa in olive trees |
| WO2023240208A1 (en) | 2022-06-09 | 2023-12-14 | Invaio Sciences, Inc. | Controlling citrus greening in citrus plants using oxytetracycline |
Non-Patent Citations (40)
| Title |
|---|
| ABRAHAMIAN PETER ET AL: "Annual Review of Virology Plant Virus-Derived Vectors: Applications in Agricultural and Medical Biotechnology", ANNU. REV. VIROL, vol. 7, 10 June 2020 (2020-06-10), pages 513 - 535, XP093047962, Retrieved from the Internet <URL:https://www.annualreviews.org/doi/suppl/10.1146/annurev-virology-010720-054958> DOI: 10.1146/annurev-virology-010720- * |
| AL-RIMAWI ET AL., ANTIBIOTICS, vol. 8, 2019, pages 196 |
| ALTSCHUL ET AL., J. MOL. BIOL., vol. 215, 1990, pages 403 - 410 |
| ALTSTEIN, PEPTIDES, vol. 25, 2004, pages 1373 - 1376 |
| AMAN ET AL., GENOME BIOL., vol. 19, 2018, pages 1 - 10 |
| CHOI ET AL., J. CONTROLLED RELEASE, vol. 235, 2016, pages 222 - 235 |
| CONG ET AL., SCIENCE, vol. 339, 2013, pages 819 - 823 |
| DE CESARE ET AL., MBIO, vol. 11, 2020, pages 02123 - 20 |
| DUANIS-ASSAF, PLANT BIOTECHNOL J., vol. 20, no. 1, January 2022 (2022-01-01), pages 226 - 237 |
| EDGAR, NUCLEIC ACIDS RES., vol. 32, no. 5, 2004, pages 1792 - 1797 |
| GEISBERG ET AL., CELL, vol. 156, 2014, pages 812 - 824 |
| GIRALDO ET AL., NATURE MATERIALS, vol. 13, 2014, pages 400 - 409 |
| GRANDI ET AL., FRONT. PLANT SCI.., vol. 14, 2023, pages 2023 |
| HENDEL ET AL., NATURE BIOTECHNOL., 2015, pages 985 - 991 |
| HOFACKER ET AL., MONATSHEFTE FOR CHEMIE CHEM. MONTHLY, vol. 125, 1994, pages 167 - 188 |
| HORIUCHI ET AL., PLANT CELL PHYSIOL, vol. 42, no. 2, 2001, pages 197 - 203 |
| KHAKHAR ARJUN ET AL: "RNA Viral Vectors for Accelerating Plant Synthetic Biology", FRONTIERS IN PLANT SCIENCE, vol. 12, 23 June 2021 (2021-06-23), XP093047125, Retrieved from the Internet <URL:https://www.frontiersin.org/articles/10.3389/fpls.2021.668580/full> DOI: 10.3389/fpls.2021.668580 * |
| KIM, BIOCONJUGATE CHEM., vol. 22, 2011, pages 2558 - 2567 |
| KOCVMILLER, J VIROL., vol. 74, no. 13, July 2000 (2000-07-01), pages 5988 - 96 |
| KRUPOVIC ET AL., BIOL DIRECT., vol. 10, 29 March 2015 (2015-03-29), pages 12 |
| LI, MOL PLANT MICROBE INTERACT., vol. 32, no. 12, December 2019 (2019-12-01), pages 1649 - 1664 |
| MATTEI ET AL., NUCLEIC ACIDS RESEARCH, vol. 42, no. 10, 2014, pages 6146 - 6157 |
| O'BRIANL MLMIS, BMC BIOTECHNOL., vol. 11, 2011, pages 66 - 71 |
| PADMANABHAN CHELLAPPAN ET AL: "Complete Genome Sequence of Southern tomato virus Identified in China Using Next-Generation Sequencing", GENOME ANNOUNCEMENTS, vol. 3, no. 5, 29 October 2015 (2015-10-29), US, XP093189109, ISSN: 2169-8287, Retrieved from the Internet <URL:https://journals.asm.org/doi/pdf/10.1128/genomeA.01226-15> DOI: 10.1128/genomeA.01226-15 * |
| PUTLYAEV ET AL., BIOCHEMISTRY (MOSC, vol. 80, no. 8, pages 1039 - 46 |
| RAMLANZAUNER: "International Workshop on Computing With Biomolecules", 27 August 2008, pages: 75 - 86 |
| RAN ET AL., NATURE PROTOCOLS, vol. 8, 2013, pages 2281 - 2308 |
| RANDOLPH-ANDERSON ET AL.: "Submicron gold particles are superior to larger particles for efficient Biolistic® transformation of organelles and some cell types", BIO-RAD US/EG BULLETIN, 2015 |
| ROBINSON ET AL., BIOINFORMATICS, vol. 39, no. 1, 2023, pages 830 |
| SHEN ET AL., THERANOSTICS, vol. 2, 2012, pages 283 - 294 |
| SMAKOV ET AL., NATURE REVIEWS MICROBIOL, 2017 |
| TANGBREAKER, PROC NATL ACAD SCI USA., vol. 97, no. 11, 23 May 2000 (2000-05-23), pages 5784 - 9 |
| WANG ET AL., J. AM. CHEM. SOC. COMM, vol. 132, 2010, pages 9274 - 9276 |
| WHITHAM, S. ET AL., CELL, vol. 78, 1994, pages 1101 - 1115 |
| WOLTER ET AL., BMC PLANT BIOL., vol. 19, 2019, pages 176 - 183 |
| WONG ET AL., NANO LETT., vol. 16, 2016, pages 1161 - 1172 |
| XING, BMC PLANT BIOL, vol. 14, 2014, pages 327 - 340 |
| YOUNG ET AL.: "Compendium of Herbicide Adjuvants", 2016, PURDUE UNIVERSITY |
| ZHANG ET AL., J. CONTROLLED RELEASE, vol. 123, 2007, pages 1 - 10 |
| ZHAO ET AL., NANOSCALE RES. LETT., vol. 11, 2016, pages 195 - 203 |
Also Published As
| Publication number | Publication date |
|---|---|
| AR132586A1 (en) | 2025-07-16 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| US12102091B2 (en) | Control of plant pests using RNA molecules | |
| US12529067B2 (en) | Methods for modifying plants with a recombinant RNA molecule | |
| BR112020007465A2 (en) | pest control of hemiptera using RNA molecules | |
| US20250230452A1 (en) | Polynucleotides for modifying organisms | |
| US11578338B2 (en) | Hemipteran active insecticidal protein | |
| US20150203866A1 (en) | Methods and compositions for plant pest control | |
| CN118556127A (en) | Polynucleotides for modifying organisms | |
| WO2024229398A1 (en) | Artificial amalgavirus satellite rnas | |
| WO2024229356A2 (en) | Artificial solemoviridae satellite rnas | |
| WO2024229362A1 (en) | ARTIFICIAL MARTELLIVIRALES SATELLITE RNAs | |
| WO2024229351A1 (en) | Artificial secoviridae satellite rnas | |
| WO2024229385A1 (en) | Artificial ghabrivirales satellite rnas | |
| WO2024229359A2 (en) | Artificial tymovirales satellite rnas | |
| WO2024229347A1 (en) | Artificial tombusviridae satellite rnas | |
| WO2024229395A1 (en) | Partitiviral satellite rna amplification systems for plants | |
| WO2024229403A1 (en) | Endornaviral satellite rna amplification systems for plants | |
| US20230189818A1 (en) | Rna-based control of botrytis | |
| CN118922549A (en) | Polynucleotides for modifying organisms |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| 121 | Ep: the epo has been informed by wipo that ep was designated in this application |
Ref document number: 24729552 Country of ref document: EP Kind code of ref document: A1 |
|
| REG | Reference to national code |
Ref country code: BR Ref legal event code: B01A Ref document number: 112025023819 Country of ref document: BR |
|
| WWE | Wipo information: entry into national phase |
Ref document number: 2024729552 Country of ref document: EP |
|
| NENP | Non-entry into the national phase |
Ref country code: DE |
|
| ENP | Entry into the national phase |
Ref document number: 2024729552 Country of ref document: EP Effective date: 20251203 |
|
| ENP | Entry into the national phase |
Ref document number: 2024729552 Country of ref document: EP Effective date: 20251203 |