WO2024228121A1 - Long interspersed nuclear element r2 for target-specific gene delivery - Google Patents
Long interspersed nuclear element r2 for target-specific gene delivery Download PDFInfo
- Publication number
- WO2024228121A1 WO2024228121A1 PCT/IB2024/054205 IB2024054205W WO2024228121A1 WO 2024228121 A1 WO2024228121 A1 WO 2024228121A1 IB 2024054205 W IB2024054205 W IB 2024054205W WO 2024228121 A1 WO2024228121 A1 WO 2024228121A1
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- polynucleotide
- sequence
- seq
- mrna
- length
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Pending
Links
Classifications
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/63—Introduction of foreign genetic material using vectors; Vectors; Use of hosts therefor; Regulation of expression
- C12N15/79—Vectors or expression systems specially adapted for eukaryotic hosts
- C12N15/85—Vectors or expression systems specially adapted for eukaryotic hosts for animal cells
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/63—Introduction of foreign genetic material using vectors; Vectors; Use of hosts therefor; Regulation of expression
- C12N15/79—Vectors or expression systems specially adapted for eukaryotic hosts
- C12N15/85—Vectors or expression systems specially adapted for eukaryotic hosts for animal cells
- C12N15/86—Viral vectors
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N2800/00—Nucleic acids vectors
- C12N2800/90—Vectors containing a transposable element
Definitions
- the clustered regularly interspaced short palindromic repeat (CRISPR)/Cas9 mediated genome editing technology has seen wild use since its invention due to its simplicity and efficiency, leading to a new era of genetic engineering and gene therapy.
- the CRISPR/Cas9 system is composed of the Cas9 endonuclease and a single-guide RNA (sgRNA), which can guide Cas9 to the complementary sequence and induce double-strand break (DSB) at the target site.
- sgRNA single-guide RNA
- the resulting DSB is repaired independently by two cellular DNA repair pathways, namely non-homologous end-joining (NHEJ) pathway and homology directed repair (HDR) pathway.
- NHEJ non-homologous end-joining
- HDR homology directed repair
- Genome editing using the CRISPR-Cas9 system in human embryos has resulted in many unexpected mutations at or near its target site as well as in the neighboring areas. Furthermore, this system depends on HDR, which is regulated in host cells, indicating that the integration process of the exogenous gene itself cannot be regulated autonomously.
- Transposons also referred to as TE or mobile genetic elements
- TE mobile genetic elements
- the two major groups of transposons are DNA transposons and retrotransposons.
- DNA transposons, or class II elements move via a “cut and paste” mechanism, by directly removing themselves from the original position and pasting at another position.
- Retrotransposons, or class I elements move in a “copy and paste” manner, using RNA as an intermediate. Based on whether they are flanked by long terminal repeat (LTR) or not, retrotransposons are further classified as LTR type or non-LTR type.
- LTR retrotransposons are similar to retroviruses in structure and life cycle, and therefore their retrotransposition mechanisms are well-explained. However, the retrotransposition mechanisms of non-LTR type elements has not been well-clarified.
- Non-LTR retrotransposons are sub-classified into two major groups, namely long interspersed nuclear elements (LINEs) and short interspersed nuclear elements (SINEs).
- LINEs are autonomous elements that encode proteins to mediate their own mobility
- SINEs are nonautonomous elements that do not encode protein and consequently require LINEs for their propagation.
- LINEs can be categorized into two groups, an early branched group with a single open reading frame (ORF) and a recently branched group, with two ORFs (ORF1 and ORF2).
- the endonuclease (EN) domain which cuts the target site DNA
- RT reverse transcriptase
- the EN and RT domains are encoded in the single ORF of an early-branched element or in ORF2 of a recently branched element, respectively.
- Non-LTR retrotransposons are mobilized by a mechanism which is very different from that used by retroviruses and LTR retrotransposons.
- the resultant proteins open reading frame proteins; ORFps
- ORFps open reading frame proteins
- RNP ribonucleoprotein
- RNP ribonucleoprotein
- TPRT target-primed reverse transcription
- the TPRT process is initiated by an encoded EN domain that nicks one strand of DNA at a target site and creates a 3 ’-hydroxyl end, which is used as a primer for reverse transcription of the LINE mRNA onto the DNA target via RT.
- the present disclosure demonstrates that a newly discovered long interspersed nuclear element (LINE) R2 of medaka fish Oryzias latipes, R2O1, can integrate into the 28S rDNA gene of a human cell in a target sequence-specific manner. With modification and/or addition of various non-coding sequences in the retrotransposon, its retrotransposition efficiency in human cells can be greatly improved. Further, additional exogenous genes can be included in the R2O1 retrotransposon, enabling the exogenous genes to be inserted into the target genome, achieving target sequence-specific gene delivery. Accordingly, the R2O1 retrotransposon systems of the present disclosure provide new and improved gene delivery approaches.
- LINE nuclear element
- a polynucleotide that is a messenger RNA (mRNA) or a DNA encoding the mRNA, wherein the mRNA comprises, from 5’ to 3’: a protein-coding sequence or a reverse-complement thereof; a 3’ untranslated region (UTR) having at least 85% sequence identity to SEQ ID NO: 16; and a downstream flanking sequence consisting of a fragment of SEQ ID NO:20 or an RNA sequence having least 85% sequence identity to SEQ ID NO:20, wherein the fragment includes at least 4 consecutive nucleotides of SEQ ID NO:20 starting at a position from 1 to 5.
- the mRNA does not include a poly(A) signal.
- the 3 ’UTR is directly connected to the downstream flanking sequence.
- the downstream flanking sequence has a length of 4 to 180 nucleotides (nt). In some embodiments, the downstream flanking sequence has a length of 10 to 150 nt. In some embodiments, the downstream flanking sequence has a length of 15 to 150 nt. In some embodiments, the downstream flanking sequence has a length of 40 to 90 nt. In some embodiments, the downstream flanking sequence has a length of 50 to 90 nt. In some embodiments, the downstream flanking sequence has a length of 71 to 90 nt.
- the fragment includes at least the four 5’ nucleotides (UAGC) of SEQ ID NO:20.
- the 3 ’UTR has at least 90% sequence identity to SEQ ID NO: 16. In some embodiments, the 3’UTR has at least 95% sequence identity to SEQ ID NO: 16. In some embodiments, the 3’UTR consists of SEQ ID NO: 16.
- the mRNA further comprises, 5’ to the protein-coding sequence, an upstream flanking sequence consisting of a fragment of SEQ ID NO: 18 or an RNA sequence having at least 85% sequence identity to SEQ ID NO: 18.
- the fragment of the upstream flanking sequence ends at a position from 176 to 180 of SEQ ID NO: 18.
- the upstream flanking sequence has a length of 4 to 180 nt.
- the upstream flanking sequence has a length of 10 to 150 nt.
- the upstream flanking sequence has a length of 32 to 150 nt.
- the upstream flanking sequence has a length of 20 to 100 nt.
- the upstream flanking sequence has a length of 32 to 80 nt.
- the mRNA further comprises, 5’ to the protein-coding sequence, a 5’ untranslated region (UTR) comprising nucleotides 233 to 265 of SEQ ID NO: 14 or a nucleic acid sequence having at least 85% sequence identity to nucleotides 233 to 265 of SEQ ID NO: 14.
- the mRNA does not include nucleotides 1 to 87 of SEQ ID NO: 14.
- the mRNA does not include nucleotides 1 to 175 of SEQ ID NO: 14.
- the mRNA does not include nucleotides 1 to 198 of SEQ ID NO: 14.
- the 5 ’UTR comprises nucleotides 216 to 265 of SEQ ID NO: 14 or a nucleic acid sequence having at least 85% sequence identity to nucleotides 216 to 265 of SEQ ID NO: 14.
- the upstream flanking sequence is located 5’ to the 5 ’UTR.
- the protein-coding sequence encodes a Long Interspersed Nuclear Element (LINE) R2 of medaka fish Oryzias latipes (R2O1) open reading frame (ORF), which can be expressed by the mRNA.
- the ORF comprises the amino acid sequence of SEQ ID NO: 15 or an amino acid sequence having at last 85% sequence identity to SEQ ID NO: 15.
- the protein-coding sequence encodes an exogenous therapeutic protein for delivery to a target cell.
- the mRNA further comprises a second protein-encoding sequence or the reverse-complement thereof, wherein the second protein-encoding sequence encodes an exogenous therapeutic protein for delivery to a target cell.
- a method for integrating an exogenous polynucleotide to the genome of a mammalian cell comprising introducing to the cell the polynucleotide of the present disclosure.
- the polynucleotide is a mRNA, a plasmid or an adenoviral vector.
- the mammalian cell is a human cell.
- the human cell is an induced pluripotent stem cell (iPSC).
- iPSC induced pluripotent stem cell
- the polynucleotide is a plasmid or an adenoviral vector and the downstream flanking sequence has a length 4 to 90 nt.
- the human cell is not a stem cell.
- the downstream flanking sequence has a length of 70 to 90 nt.
- the polynucleotide or the reverse-complement thereof encodes a protein that is not an R2O1 ORF, and the method further comprises introducing to the mammalian cell a second polynucleotide encoding the R2O1 ORF.
- FIG. 1 The structure and retrotransposition mechanism of R2O1.
- R2O1 cloned from the medaka fish, Oryzias latipes. It comprises a 5' untranslated region (UTR; 265 bp), a single open reading frame (ORF; 3831 bp) encoding EN and RT domains, and a 3' UTR (108 bp).
- R2O1 is inserted into the 28S rDNA target sequence.
- the rDNA unit is indicated by a black closed rectangle.
- the double-stranded sequences of the precise insertion sites are indicated (upper: 5’- ACTCTCTTAAGGTAGCCAAATGCC-3’, SEQ ID NO:4; lower: 5’-
- GGCATTTGGCTACCTTAAGAGAGT-3 SEQ ID NO:5
- Bottom and top strand cleavage sites generated by EN are indicated by vertical arrows.
- TPRT reverse transcription
- the EN domain cleaves the bottom strand of genomic DNA.
- R2O1 mRNA anneals with the target DNA (28S rDNA) at the 3' junction to synthesize cDNA (5’-TCTTAAGGTAGCCAAATGCC, SEQ ID NO:6; 5’- GGCATTTGCTA-3’, SEQ ID NO:7; 5’-CCTTAAGA-3’, SEQ ID NO:8; 5’- CUCAAGUAGC-3, SEQ ID NO: 9).
- FIG. 2 Locations of primer sets used for the detection of R2O1 retrotransposition. Nested PCR was used to detect the junction site between inserted R2O1 and 28SrDNA. Horizontal arrows indicate the PCR primers used to detect retrotransposition events. Primer sets “g,” “h,” “i,” and “j” were used for 3' junction analysis, while the primer sets “f”, “al”, “a2”, “a3”, “a4” and “a5” were used in 5' junction analysis. The rDNA unit is indicated by a black closed rectangle. The double-strand sequences of the precise insertion sites are indicated (upper: SEQ ID NO:4; lower: SEQ ID NO:5).
- FIG. 3A-D 3' junction PCR of inserted R201-r4.
- A Schematic representation of R2O1 used in the human cell retrotransposition assay.
- R2O1 contains a 4-bp target sequence downstream of the 3' UTR.
- CAG chicken beta-actin promoter and cytomegalovirus immediate-early enhancer-promoter.
- pA poly A signal.
- B Horizontal arrows indicate PCR primers used to detect retrotransposition events.
- the predicted PCR band sizes were: 327 bp under primer set h & i; and 480 bp under primer set g & j, indicated by solid arrowheads, respectively.
- the PCR products indicated by a triangle mark were TA cloned for further analyses.
- R201-ENmut is a construct in which the EN portion of the 1149th amino acid (IPD — IP A) has been mutated (indicated by the dashed line).
- N negative-control PCR conducted with genomic DNA from non-infected cells. Non-specific sequences indicated by open arrowheads. Numbers above the lanes indicate independent experiments.
- Horizontal arrows indicate PCR primers used to detect 5' junctions;
- “f” indicates the forward primer set upstream of the target site at 28S rDNA;
- “al”, “a2”, “a3”, “a4”, and “a5” are the five reverse primer sets at the R2O1 sequences.
- FIG. 5. 5' junction analysis around the insertion site of R201-r4.
- the top of the figure includes a diagram of the 28 S target site, indicated by black boxes, with R201-r4 sequences in the middle, indicated by red. Numbers on the left/right (-/+) of each region indicate the position of the accurate cleavage site of the target DNA regions.
- the accurate reverse transcriptionending site of the sequence (in bases) is indicated by the number on the top of the diagram. Extra sequences derived from the vector sequences at the junction are indicated by white triangles.
- FIG. 6A-B 3' junction analysis of inserted r32R201-r4.
- A Schematic representation of r32R201-r4 used in the human cell retrotransposition assay.
- R2O1 contains 32 bp 28S flanking sequences upstream of the 5' UTR; pA, poly A signal.
- B 3' junction region PCR analysis to detect retrotransposition in 293T cells after infection of cells at different MOIs. The predicted PCR band size of 480 bp is indicated by the arrowhead. Numbers above the lanes indicate independent experiments; N, negative-control. PCR conducted with genomic DNA from non-infected cells.
- FIG. 7. 5' junction analysis around the insertion site of r32R201-r4. 5' junction region around the insertion site of r32R201-r4.
- the top of the figure includes a diagram of 28 S target site indicated by black boxes, with the R201-r4 sequences in the middle, indicated by gray. Numbers on the left/right (-/+) of each region indicate the position of the accurate cleavage site of the target DNA regions. The accurate reverse transcriptionending site of the sequence (in nucleotide bases) is indicated by the number on the top of the diagram. Extra sequences derived from vector sequences at the junction are indicated by white triangles.
- FIG. 8A-C 3' junction analysis of inserted r32R201-r4.
- A Schematic representation of plasmids used in the human cell retrotransposition assay. All plasmids contained a 4-bp target sequence downstream of the 3' UTR.; CAG, chicken beta-actin promoter and cytomegalovirus immediate-early enhancer promoter. The RTmut construct was mutated at the essential amino acid site for RT; pA, poly A signal.
- B The 3' junction region PCR analysis to detect retrotransposition in 293T cells after infecting cells at different MOIs. The predicted PCR band of 480 bp is indicated by the arrowhead. Numbers above the lanes indicate independent experiments; N, negative-control; PCR conducted with genomic DNA from non-infected cells.
- C The 3' junction sequence of the inserted R2O1 plasmid (5’-
- GCGAGTGGTGACTCGCCTCAAGTAGCCAAATGCCTCGTCATCT-3’ SEQ ID NO: 10; 5’- GGCATGATTACAAATCTT-3’, SEQ ID NO: 11; 5’-CCTCGTCATCT-3’, SEQ ID NO: 12).
- the nucleic acid sequences of the PCR product were analyzed through direct sequencing. Shown on top is a diagram of the correct insertion of R2O1 3' end. R2O1 sequences are shown on the left and 28 S target DNA sequences are shown on the right. The initiation sites of reverse transcription are indicated by nucleotide numbers.
- FIG. 9A-B mRNA-mediated R2O1 retrotransposition in 293T cells.
- A Schematic representation of mRNA transcription in vitro.
- pGEM-rl05-R2Ol-rl00 contains a T7 promoter, a WT R2O1, and fused with a 100 bp downstream 28S rDNA sequence of R2O1 insertion site at the 3 '-end.
- pA poly A signal.
- pGEM-R201 RTmut containing T7 promoter and R2O1 RTmut (i) Template DNA for in vitro transcription was amplified via PCR.
- the template at the right (RTmut-r4, RTmut-rlO and RTmut-r71) indicates the RT with 28S target sequences of the same length as in WT templates (4 bp/10 bp/50 bp) attached to the end.
- Six types of 5' capped mRNA were synthesized in vitro.
- FIG. 10A-B In vitro synthesis of mRNA used for iPS cell transfection.
- A Schematic representation of mRNA transcription in vitro pGEM-rl05-R2Ol-rl00 containing T7 promoter; WT R2O1 fused with 100 bp 28 S rDNA target site at the 3 end.
- the template DNA for in vitro transcription was amplified via PCR.
- the template at the left indicates WTs with 28S target sequences of different lengths (4 bp/10 bp/50 bp) attached to the end.
- the templates at the right show RTs with 28S target sequences of the same length as in WT templates (4 bp/10 bp/50 bp) attached to the end.
- the six types of 5' capped mRNA were synthesized in vitro.
- B 3' junction region PCR analysis aimed at detecting retrotransposition in iPS cells 72 hours after transfection.
- the predicted PCR band of size 541 bp (contained 61 bp vector sequences between R2O1 ORF and R2O1 3 UTR) is indicated by the arrowhead. Numbers above the lanes indicate independent experiments; N, negative-control PCR conducted with genomic DNA from non-infected cells.
- FIG. 11 illustrates R2O1 constructs with various non-coding sequences added or deleted.
- the retrotransposition efficiency testing results show that neither the 5 ’UTR nor the poly(A) signal was required for retrotransposition.
- FIG. 12 shows design and testing of R2O1 constructs with 4 consecutive nucleotides of the 28S downstream sequence added to the 3’.
- the 4 consecutive nucleotides are close ( ⁇ 6 bases) to the target insertion site, it promoted more efficient retrotransposition than without the short nucleotides.
- FIG. 13A shows construction of R2O1 retrotransposons with different lengths of the 5’UTR deleted and their impact on retrotransposition efficiency. A deletion between 175 and 198 nucleotides of the 5’ portion of the 5’UTR exhibited the highest retrotransposition efficiency.
- FIG. 13B shows construction of R2O1 retrotransposons with different lengths of the 3’UTR deleted and their impact on retrotransposition efficiency.
- the intact 3’UTR resulted in the highest retrotransposition efficiency.
- FIG. 14A-B show construction and testing of R2O1 constructs with exogenous genes inserted downstream of the ORF. Both tested exogenous genes, EGFP and Gal4, were successfully inserted to the target 28 S rDNA and expressed.
- FIG. 15A-C show that the R2O1 ORF can be expressed separately to enable retrotransposition of a R2O1 construct that embeds an exogenous gene.
- LINEs are randomly inserted throughout the host genome, such as LI and L2. Some elements are inserted into specific sites of repetitive genomic sequences, such as ribosomal DNA (rDNA), telomeric repeats, and microsatellites. Among the site-specific LINEs, the R2 elements specifically insert into the 28 S rDNA.
- rDNA ribosomal DNA
- telomeric repeats telomeric repeats
- microsatellites atellites
- R2 elements are widely distributed among vertebrate species, such as fish and birds but not in mammals.
- an approximately 100-base pair (bp) region around the R2 insertion site in 28S rDNA is strongly conserved across a wide variety of eukaryotes, suggesting the high sequence specificity of this element.
- R2Bm isolated from Bombyx mori, exhibited efficient site-specific retrotransposition activity both in vitro and in vivo, thereby facilitating a better understanding of the general mechanism underlying LINE retrotransposition.
- R2O1 isolated from the medaka fish, Oryzias latipes, includes a 5' untranslated region (UTR; 265 nt), a single open reading frame (ORF; 1276 amino acids) encoding the endonuclease (EN) and reverse transcriptase (RT) domains, and a 3' UTR (111 nt) (FIG. 1A).
- the 28S rDNA target sequence and the exact cleavage sites of R2O1 on both strands are shown in FIG. IB.
- R2O1 mRNA anneals with the target site and initiates the reverse transcription mechanism.
- an adenovirus-mediated R2O1 retrotransposition system was efficient at integrating to the 28 S rDNA gene in human 293T cells.
- mRNA-mediated R2O1 retrotransposition systems could readily integrate to 28S gene in 293T as well as in induced pluripotent stem cells (iPSC).
- the R2O1 (FIG. 1A, Table 2, SEQ ID NO: 13) includes a 5' untranslated region (UTR; 265 nt; SEQ ID NO: 14), a single open reading frame (ORF; 1276 amino acids; SQE ID NO: 15) encoding the endonuclease (EN) and reverse transcriptase (RT) domains, and a 3' UTR (108 nt; SEQ ID NO: 16).
- the 3’UTR is required for efficient retrotransposition (FIG. 13B). Retaining of some portions of the 5 ’UTR, by contrast, actually reduced the retrotransposition efficiency.
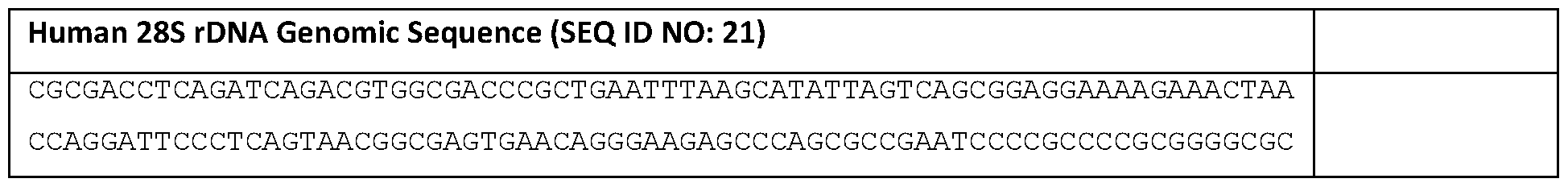
- the R2O1 retrotransposon is capable of target sequence-specific integration. More specifically, in human cells, the R2O1 can insert itself to between 3746 and 3747 of the 28 S gene (and between 3744 and 3745 on the opposite strand) (FIG. IB, Table 3, SEQ ID NO 21).
- the four-nucleotide rDNA sequences immediately downstream the insertion site are ATCG (nucleotides 3747-3750 of SEQ ID NO:21; or nucleotides 1-4 of SEQ ID NO: 19). As shown in the examples, inclusion of these 4 nucleotides (referred to as r4) at the 3’ end of the R2O1 can greatly increase its retrotransposition efficiency. Further testing (e.g., FIG. 12) shows that actually any 4-consecutive nucleotides within the first 8 nucleotides of this “downstream sequence” (SEQ ID NO: 19 or 20) can have similar results.
- the R2O1 retrotransposon integrate itself to a target sequence in the human genome, but it can also further include additional coding sequences so as to achieve delivery of other genes.
- an EGFP and a Gal4 encoding sequence were inserted to the R2O1 retrotransposon, downstream of the R2O1 ORF. Both exogenous genes were integrated to the target genome and expressed (FIG. 14A-B). Note that both genes were inserted backwards into the retrotransposon such that they are not functionally expressed with the R2O1 mRNA, and are only expressed after genomic integration.
- the R2O1 ORF does not need to be included in the retrotransposon itself in order to enable retrotransposition. Therefore, if the gene to be delivered is very large, the R2O1 retrotransposon can just include that gene, while the R2O1 ORF can be delivered separately (FIG. 15A). This way, the technology can allow better control, and delivery of very large exogenous genes to a target cell.
- the instant data demonstrate that the R2O1 transposition system can be suitably used to deliver an exogenous coding sequence to a target cell. Since the integration is target sequence-specific, such a gene delivery technology is highly safe.
- a polynucleotide that is a messenger RNA (mRNA) or a DNA encoding the mRNA.
- the mRNA includes, from 5’ to 3’ (A) a protein-coding sequence or a reversecomplement thereof; (B) a 3’ untranslated region (UTR); and (C) a downstream flanking sequence.
- the mRNA further includes (D) an upstream flanking sequence that is 5’ to (A).
- the mRNA further includes (E) a 5’ UTR that is 5’ to (A).
- the mRNA includes, from 5’ to 3’, (D), (E), (A), (B), and (C). In some embodiments, the mRNA includes, from 5’ to 3’, (D), (A), (B), and (C). In some embodiments, the mRNA includes, from 5’ to 3’, (E), (A), (B), and (C). In some embodiments, the mRNA includes, from 5’ to 3’, (A), (B), and (C). In some embodiments, the mRNA does not include a poly(A) signal. In some embodiments, the mRNA includes a 5’ Cap. In some embodiments, the mRNA does not include any additional sequence or element not specifically recited herein.
- the 3’ untranslated region (UTR) of the R2O1 retrotransposon plays an important role in the retrotransposition.
- the 3 ’UTR has the nucleic acid sequence of SEQ ID NO: 16.
- the 3’UTR has at least 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, 97%, 98%, 99% or 100% sequence identity to SEQ ID Downstream Flanking Sequence
- the downstream flanking sequence refers to a fragment of the 28S (e.g., mammalian 28S such as human 28S) sequence that is downstream of the R2O1 integration site.
- An example 180-nt sequence is provided in SEQ ID NO: 19 (DNA) or SEQ ID NO:20 (RNA).
- the fragment has a length that is at least 4 nt. In some embodiments, the fragment has a length that is at least 5, 10, 15, 20, 25, 30, 35, 40, 45, 50, 55, 60, 65, 70, 71, 75, 80, 85, or 90 nt. In some embodiments, the fragment has a length that is not longer than 10, 15, 20, 25, 30, 35, 40, 45, 50, 55, 60, 65, 70, 71, 75, 80, 85, 90 or 100 nt. In some embodiments, the length is 4-180, 10-150, 15-150, 40-90, 50-90, 70-90 or 71-90 nt, without limitation.
- the fragment is a fragment of SEQ ID NO:20. In some embodiments, the fragment is a fragment of an RNA sequence having least 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, 97%, 98%, 99% or 100% sequence identity to SEQ ID NO:20.
- the fragment starts at position, according to SEQ ID NO:20, that is within positions 1-20, 1-15, 1-10, or 1-5. In some embodiments, the fragment includes at least 4 consecutive nucleotides of SEQ ID NO:20 starting at a position from 1 to 5. In some embodiments, the fragment includes at least the first 4 nucleotides of SEQ ID NO:20 (UAGC).
- the 3’UTR is directly connected to the downstream flanking sequence.
- the polynucleotide (e.g., mRNA) further includes an upstream flanking sequence that is 5’ to the protein-coding sequence.
- the upstream flanking sequence in one embodiment, refers to a fragment of the 28 S (e.g., mammalian 28 S such as human 28S) sequence that is upstream of the R2O1 integration site.
- An example 180-nt sequence is provided in SEQ ID NO: 17 (DNA) or SEQ ID NO: 18 (RNA).
- the fragment has a length that is at least 4 nt. In some embodiments, the fragment has a length that is at least 5, 10, 15, 20, 25, 30, 32, 35, 40, 45, 50, 55, 60, 65, 70, 75, 80, 85, or 90 nt. In some embodiments, the fragment has a length that is not longer than 10, 15, 20, 25, 30, 35, 40, 45, 50, 55, 60, 65, 70, 71, 75, 80, 85, 90 or 100 nt. In some embodiments, the length is 4-180, 10-150, 15-150, 32-150, 20-100, 32-80, 40-90, 50-90, 70-90 or 71-90 nt, without limitation.
- the fragment is a fragment of SEQ ID NO: 18. In some embodiments, the fragment is a fragment of an RNA sequence having least 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, 97%, 98%, 99% or 100% sequence identity to SEQ ID NO: 18.
- the fragment ends at position, according to SEQ ID NO: 18, that is within positions 161-180, 166-180, 170-180, or 176-180. In some embodiments, the fragment includes at least 4 consecutive nucleotides of SEQ ID NO: 18 ending at a position from 176 to 180.
- the polynucleotide (e.g., mRNA) further includes a 5’ untranslated region (UTR) 5’ to the protein-coding sequence.
- UTR untranslated region
- the upstream flanking sequence is 5’ to the 5 ’UTR.
- the wild-type R2O1 5’UTR sequence is provided in SEQ ID NO: 14. As shown in the data, the 5’UTR sequence is not essential to the retrotransposition activity of the R2O1 retrotransposon. Moreover, the 5’ portion of the 5’UTR appeared to inhibit the retrotransposition activity. In some embodiments, therefore, when the 5’UTR is included, it preferably includes nucleotides 233 to 265 of SEQ ID NO: 14 or a nucleic acid sequence having at least 85% sequence identity to nucleotides 233 to 265 of SEQ ID NO: 14. In some embodiments, the 5’UTR includes nucleotides 216 to 265 of SEQ ID NO: 14 or a nucleic acid sequence having at least 85% sequence identity to nucleotides 216 to 265 of SEQ ID NO: 14.
- the polynucleotide does not include nucleotides 1 to 87 of SEQ ID NO: 14. In some embodiments, the polynucleotide (e.g., mRNA) does not include nucleotides 1 to 175 of SEQ ID NO: 14. In some embodiments, the polynucleotide (e.g., mRNA) does not include nucleotides 1 to 198 of SEQ ID NO: 14. In some embodiments, the polynucleotide (e.g., mRNA) does not include nucleotides 88 to 175 of SEQ ID NO: 14. Protein-Coding Sequence
- the wild-type R2O1 includes an R2O1 ORF that has the required EN and RT activities. It has been shown, however, that for the R2O1 retrotransposon to work, the ORF can be expressed from a separate polynucleotide, which enables the “empty” R2O1 retrotransposon to include a large exogenous gene for delivery to the target cell. Accordingly, the R2O1 retrotransposon of the instant disclosure can include a coding sequence that encodes the R2O1, the exogenous gene for delivery, or the combination thereof.
- the protein-coding sequence encodes a Long Interspersed Nuclear Element (LINE) R2 of medaka fish Oryzias latipes (R2O1) open reading frame (ORF), which can be directly expressed by the mRNA (e.g., in the correct orientation for translation).
- the ORF comprises the amino acid sequence of SEQ ID NO: 15 or an amino acid sequence having at last 85% sequence identity to SEQ ID NO: 15.
- the polynucleotide (e.g., mRNA) further includes a second protein-coding sequence which encodes an exogenous protein for delivery to a target cell.
- the second protein-coding sequence is 3’ to the R2O1 ORF-coding sequence.
- the second protein-coding sequence is included in the mRNA in a reverse-complement orientation such that it cannot be directly translated by the mRNA. Rather it is translated after integration to the target genome.
- the polynucleotide (e.g., mRNA) does not encode the R2O1 ORF, and instead encodes an exogenous protein to be expressed in a target cell.
- this coding sequence is included in the correct orientation for translation. In some embodiments, it is in a reverse-complement orientation.
- the protein is a therapeutic protein that can be helpful in treating a disease or condition in a patient.
- examples include:
- G-CSF Filgrastim
- TNF Tumor necrosis factor
- Enzyme replacement therapy enzymes used to treat lysosomal storage disorders, such as Gaucher's disease
- the R2O1 retrotransposition systems disclosed here are suitable for delivering exogenous genes to a target cell, such as mammalian cells.
- a target cell such as mammalian cells.
- target sequence (28S rDNA)-specific integration it can be highly efficient and safe. This new technology, therefore, has promising therapeutic uses as well.
- the present disclosure provides a method for integrating an exogenous polynucleotide to the genome of a mammalian cell.
- the method entails introducing to the cell a R2O1 retrotransposon of the instant disclosure.
- the retrotransposon is delivered to the target cell as an mRNA molecule directly.
- mRNAs may be synthesized according to any of a variety of known methods.
- the mRNAs may be synthesized via in vitro transcription (IVT).
- IVT in vitro transcription
- a linear or circular DNA template containing a promoter, a pool of ribonucleotide triphosphates, a buffer system that may include DTT and magnesium ions, and an appropriate RNA polymerase (e.g., T3, T7 or SP6 RNA polymerase), DNase I, pyrophosphatase, and/or RNase inhibitor.
- RNA polymerase e.g., T3, T7 or SP6 RNA polymerase
- the mRNA may be synthesized as unmodified or modified mRNA.
- mRNAs are modified to enhance stability.
- Modifications of mRNA can include, for example, modifications of the nucleotides of the RNA.
- a modified mRNA can thus include, for example, backbone modifications, sugar modifications or base modifications.
- antibody encoding mRNAs may be synthesized from naturally occurring nucleotides and/or nucleotide analogues (modified nucleotides) including, but not limited to, purines (adenine (A), guanine (G)) or pyrimidines (thymine (T), cytosine (C), uracil (U)), and as modified nucleotides analogues or derivatives of purines and pyrimidines, such as e.g.
- nucleotide analogues modified nucleotides
- purines adenine (A), guanine (G)
- pyrimidines thymine (T), cytosine (C), uracil (U)
- modified nucleotides analogues or derivatives of purines and pyrimidines, such as e.g.
- the mRNAs may contain RNA backbone modifications.
- a backbone modification is a modification in which the phosphates of the backbone of the nucleotides contained in the RNA are modified chemically.
- Exemplary backbone modifications typically include, but are not limited to, modifications from the group consisting of methylphosphonates, methylphosphoramidates, phosphoramidates, phosphorothioates (e.g. cytidine 5 ’-O-(l -thiophosphate)), boranophosphates, positively charged guanidinium groups etc., which means by replacing the phosphodiester linkage by other anionic, cationic or neutral groups.
- the mRNAs may contain sugar modifications.
- a typical sugar modification is a chemical modification of the sugar of the nucleotides it contains including, but not limited to, sugar modifications chosen from the group consisting of 2’- deoxy-2’-fluoro-oligoribonucleotide (2’-fluoro-2’-deoxycytidine 5 ’-triphosphate, 2’-fluoro- 2’-deoxyuridine 5 ’-triphosphate), 2’-deoxy-2’-deamine-oligoribonucleotide (2’-amino-2’- deoxycytidine 5 ’-triphosphate, 2’ -amino-2’ -deoxyuridine 5 ’-triphosphate), 2’-O- alkyloligoribonucleotide, 2’-deoxy-2’-C-alkyloligoribonucleotide (2’-O-methylcytidine 5’- triphosphate, 2’-methyluridine 5 ’-triphosphate),
- the mRNAs may contain modifications of the bases of the nucleotides (base modifications).
- a modified nucleotide which contains a base modification is also called a base-modified nucleotide.
- base-modified nucleotides include, but are not limited to, 2-amino-6-chloropurine riboside 5 ’-triphosphate, 2- aminoadenosine 5 ’-triphosphate, 2-thiocytidine 5 ’-triphosphate, 2-thiouridine 5’- triphosphate, 4-thiouridine 5 ’-triphosphate, 5-aminoallylcytidine 5 ’-triphosphate, 5- aminoallyluridine 5 ’-triphosphate, 5-bromocytidine 5 ’-triphosphate, 5 -bromouridine 5’- triphosphate, 5-iodocytidine 5 ’-triphosphate, 5-iodouridine 5 ’-triphosphate, 5-methylcytidine 5 ’
- the mRNAs include a 5’ cap structure.
- a 5’ cap is typically added as follows: first, an RNA terminal phosphatase removes one of the terminal phosphate groups from the 5’ nucleotide, leaving two terminal phosphates; guanosine triphosphate (GTP) is then added to the terminal phosphates via a guanylyl transferase, producing a 5’5’5 triphosphate linkage; and the 7-nitrogen of guanine is then methylated by a methyltransferase.
- GTP guanosine triphosphate
- cap structures include, but are not limited to, m7G(5’)ppp (5’(A,G(5’)ppp(5)A and G(5)ppp(5’)G.
- the retrotransposon of the present disclosure is delivered by a vector such as a plasmid vector and a viral vector (e.g., adenoviral vector).
- a vector such as a plasmid vector and a viral vector (e.g., adenoviral vector).
- the target mammalian cell is a human cell.
- the human cell is an induced pluripotent stem cell (iPSC).
- iPSC induced pluripotent stem cell
- the downstream flanking sequence of the retrotransposon has a length of at least 40 nt, such as 40 to 90 nt, 45 to 85 nt, or 50 to 80 nt.
- the human cell is not a stem cell.
- the downstream flanking sequence of the retrotransposon has a length of at least 70 nt, such as 70 to 95 nt, 70 to 90 nt, or 71 to 85 nt.
- the R2O1 ORF and the exogenous gene are included in the same retrotransposon polynucleotide. In some embodiments, only the exogenous gene is included in the retrotransposon polynucleotide, while the R2O1 ORF is delivered separately on a different polynucleotide.
- the delivery is in vitro. In some embodiments, the delivery is in vivo or ex vivo. In some embodiments, the retrotransposon polynucleotide is delivered by parenteral means, or locally.
- parenteral refers to modes of administration which include intravenous, intramuscular, intraperitoneal, intrastemal, subcutaneous and intraarticular injection and infusion.
- Delivery can be systemic or local.
- Pulmonary administration can also be employed, e.g., by use of an inhaler or nebulizer, and formulation with an aerosolizing agent.
- Example 1 Target-specific retrotransposition of R2O1 in human cells via viral vector delivery
- This example developed a viral vector delivery system for R2O1 and tested targeted retrotransposition of the R2O1 elements.
- R2O1 Nine full-length sequences of R2 (R2O1) were isolated from the HdrR strain of Japanese medaka fish (Oryzias latipes). Clone F (Accession number: LC349444), which contained minimal amino acid changes in comparison with that of the consensus sequence, was selected for further analyses. Next, R2O1 mRNA was injected into zebrafish embryos, and it displayed the ability to retrotranspose into a specific target sequence in the 28s rDNA locus. Thus, a transgene system with R2O1 was established in zebrafish embryos, and it functioned successfully.
- a recombinant virus (autographa californica nuclear polyhedrosis virus (AcNPV)) was used to deliver R2O1 into human cells, resulting in detection of retrotransposition activity.
- AcNPV autographa californica nuclear polyhedrosis virus
- a retrotransposition assay system was established using an adenovirus-based expression system, to determine whether R2O1 is able to retain its retrotransposon activity in human cells.
- This recombinant adenovirus pAxCAwtit-vectors contained the chicken beta-actin promoter and cytomegalovirus immediate-early enhancer (CAG) promoter.
- the human cell line, 293 T (RCB2202), was purchased from the RIKEN cell bank in Japan. Cells were grown in Dulbecco’ s minimal essential medium (DMEM) supplemented with 10% fetal bovine serum (Gibco) and 1% penicillin/streptomycin (Invitrogen) in 5% CO2 at 37 °C. Cells were grown as adherent cell cultures and passaged every 72 h at a 1 :5 dilution (volume of cells: final volume of medium) until confluency was reached, in order to maintain log-phase growth. Recombinant adenovirus construction
- Adenovirus expression vectors were constructed with pAxCAwtit (TaKaRa) containing the chicken beta-actin promoter and CAG promoter. Furthermore, pENTR201r4, pr32ENTR201r4, or pENTR201-ENmutant were mixed with LR Clonase® (Gateway® Vector Conversion System, Invitrogen) to generate recombinant pAxCAwtit according to the manufacturer’s protocol.
- R2O1 WT wild type was constructed based on R2O1 (Accession number: LC349444). Briefly, full-length R2O1 from medaka fish genomic DNA was amplified using PCR with primers R2O1 5' end and R2O1 3' end. A 4.2 kb amplified fragment, which had less mutations, clone F, was named pGEM-R201-F. A full-length R2O1 fragment, digested with EcoRI and Spel from pGEM-R201-F, was inserted between the EcoRI and Spel sites and a SV40 poly(A) signal was inserted between Not!
- pBluescript II-R2O1 RTmut was constructed by introducing a point mutation using inverse PCR from pR201r4, with primers R2O1-RT(D — V) s and R2O1- RT(D — V), and self-ligated to obtain pR201-RTmut, which was named pBluescript II-R2O1 RTmut.
- pGEM-R201 and pBluescript II-R201-RTmut were amplified using PCR with the /'i RI-R2O15U and R2O13Ur4-EcoRI primers.
- the PCR reaction used consisted of an initial denaturation at 98 °C for 1 min, followed by 25 cycles at 98 °C for 10 s, 55 °C for 30 s, and 72 °C for 30 s, and a final extension at 72 °C for 5 min.
- PCR products were purified using a GenElute PCR Clean-Up Kit (Sigma-Aldrich) and digested using EcoRI in M buffer (TaKaRa) at 37 °C for 1 h.
- the empty vector pCAGGS (RIKKEN, catalog No. RDB0893) was digested using EcoRI in M buffer (TaKaRa) at 37 °C for 1 h.
- electrophoresis in a 1% TAE 40 mM Tris, 20 mM CHiCOONa, ImM EDTA
- target bands were cut out from the gel and purified by a QIAquick Gel Extraction Kit (QIAGEN).
- Plasmid DNA was isolated using a GenElute Plasmid DNA Miniprep Kit (Sigma-Aldrich). Plasmids were sequenced using a BigDye Terminator Cycle Sequencing Kit (Applied Biosystems) on a ABB 130 xl and a 3500/3500 xl Genetic Analyzer (Applied Biosystems). Sequence analysis was performed using a Vector NTI Advance 10 system (Invitrogen).
- the upper aqueous phase was transferred to a new tube, treated with 150 pL isopropanol and gently inverted 50 times. Following centrifugation at 12,000 x g for 1 min at room temperature, the DNA was washed with 70% ethanol and air dried for 5 min. Finally, the pellet was dissolved in Tris- EDTA (TE) buffer and stored.
- TE Tris- EDTA
- Retrotransposition events were analyzed using both 3' and 5' junction PCR of the insertion. PCR reactions were performed with two types of polymerases and two sets of primers. The first polymerase was TaKaRa Ex Taq polymerase in a total volume of 20 pL, which was used according to the manufacturer’s protocol. The 3' junction of the R2O1 insertions was detected using nested PCR to identify retrotransposition events with R2O1 primer sets, h and Hs28S set i (FIG. 2). The second polymerase, KOD One ® PCR Master Mix (TOYOBO), was used for nested PCR in total volumes of 10 pL using the manufacturer’s protocol.
- TOYOBO KOD One ® PCR Master Mix
- the 3' junction of the R2O1 insertions here was detected using R2O1 primer sets, g and Hs28S set j (FIG. 2).
- the 5' junction of the R2O1 insertions was detected via nested PCR at five R2O1 positions.
- the forward primers were specific to an upstream sequence of the 28 S rDNA target site (Hs28S set f).
- the reverse primer sets were specific to five R2O1 positions al, a2, a3, a4, and a5 (FIG. 2).
- the nested-PCR products were used as directed.
- the Taq nested-PCR products were TA cloned into the pGEM-T Easy vector system (Promega) whereas the KOD nested-PCR products were TA cloned into the pTA2 vector from TArget Clone -Plus system (TOYOBO).
- TOYOBO TArget Clone -Plus system
- all PCR primers and unincorporated nucleotides of both PCR products were removed via EXO-SAP, and then the cleaned samples were sequenced using a BigDye Terminator Cycle Sequencing Kit (Applied Biosystems) on an ABB 130 xl and 3500/3500 xl Genetic Analyzer (Applied Biosystems). All sequence analyses were performed using a Geneious Prime system (Digital Biology). Results
- the wild type adenovirus construct, R201-r4, contained a CAG promoter for mammalian cells, a full-length R2O1 transcription unit (5' UTR/ORF/3' UTR) of R2O1, a 4 bp 28S rDNA target flanking sequence at its 3' end, and a polyadenylation signal sequence (PA; FIG. 3A).
- a 4 bp 28S rDNA flanking sequence (r4) added at the end of R2O1 3' UTR was contemplated to increase the annealing efficiency between the template RNA and target DNA.
- an R2O1 mutant construct, R201-ENMut was used as a negative control.
- R2O1- ENMut carried a mutation (IPA to IPD) at the essential amino acid sites for EN, completely abolishing any retrotransposition activities (FIG. 3A).
- Primer sets “h,” “i” and “g,” “j” were used to detect the 3' junction of retrotransposition (FIG. 3B).
- primer sets “g” and “j” were used, the expected PCR band (sized 480 bp), which represents the exact retrotransposition event of R2O1, was detected at MOI 10 and 50 without any non-specific bands. Therefore, primer sets “g” and “j” were selected for further experiments. In addition, the beta-Actin gene was used as an internal control to check whether the genomes of human cells had been equally extracted.
- PCR products denoted by a solid triangle in the images were TA cloned.
- Out of 142 colony PCR products, 38 were sequenced.
- sequences from 14 clones contained both R2O1 and human 28 S rDNA.
- the remaining clones originated from the genomic sequences of adenovirus or 28 S rDNA.
- FIG. 5 The 5' junction regions around the insertion site of R2O1 for the above-mentioned 14 clones is summarized (FIG. 5). Most importantly, 2 clones showed that the full-length of R2O1 was inserted accurately into the target site, whereas the 5' side vector sequence was also reverse transcribed together with R2O1 (FIG. 5; insertion type a & b). Six clones were inserted into the 21 bp ⁇ 191 bp region downstream of the original target site (FIG. 5; insertion type d, e, i, k & 1), whereas 6 clones were inserted into the 5 bp ⁇ 22 bp region upstream of the target, within 28S rDNA (FIG.
- R201 was retrotransposed successfully with higher full-length integration of R201 in 293T cells infected with adenovirus carrying r32R20l-r4
- the truncated regions ranged in length from 71 bp to 3739 bp from the 5'-end of R2O1.
- the remaining clones originated from the genomic sequences of adenovirus or 28S rDNA.
- a comparison between the 5' junction sequence analyses of R201-r4 and r32R201-r4 indicated that two out of 14 clones represented full- length R2O1 insertion in R201-r4 as opposed to five out of 15 clones in r32R201-r4.
- the results showed that flanking sequences at the 5' side of R2O1 may increase integration at the 5' junction.
- retrotransposition of R2O1 in human cells via plasmid transfection was also investigated.
- retrotransposition activity could not be detected with a plasmid carrying the CMV promoter (R201-pCEP4).
- this example changed the promoter and constructed a R2O1 plasmid with a CAG promoter (pCAGGS-R201; FIG. 8A) and detected successful integration of R2O1 after 3 days of plasmid transfection, via 3' junction analysis (FIG. 8B-C).
- expected PCR bands were not detected in samples of RTmut, suggesting that R2O1 retrotransposed via plasmid delivery (FIG. 8B).
- an adenovirus-based in vivo retrotransposition assay indicated that R2O1 isolated from medaka fish, exhibited retrotransposition in human 293T cells. This assay further showed that both the 3 '-end and 5 '-end of R2O1 were inserted into the exact target site of 28S rDNA, showing that a full-length R2O1 unit had been reverse transcribed from 3 '-end of 3' UTR to the 5 '-end of 5' UTR.
- R2 elements are distributed in a wide variety of species ranging from arthropods to vertebrates (birds and fish), where these elements are found inserted into exactly the same target sequence in 28 S rDNA.
- the R2 element had become defunct from the genome of mammals, including that of Homo sapiens.
- this study showed that the R2O1 originating in medaka fish had also retained the ability to insert into the same 28S rDNA target sequence in human cells.
- the R2 element retrotransposed into human 28S rDNA at the 3' side as well as the 5' side of the target.
- a full-length insertion > 4.2 kb
- R2O1 sequence was detected, suggesting that longer genes of interest, which is at least approximately 4 kb, could be knocked-in by this system.
- the analysis of 5' junction sequences of retrotransposed R2Ols also showed 5' truncations at various sites within R2O1 and different insertion sites around the 28 S target site.
- the molecular mechanism of LINE retrotransposition remains unclear.
- the annealing efficiency between template RNA and target DNA may increase R2O1 retrotransposition at the 3' side.
- the 3' junction analysis detected inaccurate cleavage of the bottom strand target in r32R201r4.
- the cleavage of the bottom strand target in R201-r4 appeared to be accurate. This observation indicates that the 32 bp 28S rDNA flanking sequences at the 5' end may affect the recognition or cleavage of the bottom strand target by R2O1.
- This example established an R2O1 retrotransposition system through mRNA- mediated delivery in human cells.
- iPS cell Cellartis Human induced pluripotent stem (iPS) cells (Y00275) were purchased from the TAKARA in Japan, were grown in the Cellartis DEF-CSTM 100 Culture System, which is a complete system for efficient expansion and scale-up manufacturing of human iPS cells in a feeder-free and defined environment, at 37 °C in 5% CO2.
- iPS Cellartis Human induced pluripotent stem
- Template DNA for in vitro transcription was amplified by PCR.
- R201-r4, R201-rlO and R2Ol-r50 were PCR amplified from prl05R2Olrl00, R201-RTmut-rlO and R2O1- RTmut-r50 were PCR amplified from pGEM-R201 RTmut, respectively.
- the forward primers used to transcribe these six types of mRNA, R201-r4, R201-rlO, R2Ol-r50, R2O1- RTmut-r4, R201-RTmut-rlO and R2Ol-RTmut-r50 were T7 5UTR.
- R201-r4 and R201-RTmut-r4 were 28S 4a, for R201-rlO and R201-RTmut-rlO were 28S 10a, for R2Ol-r50 was 28S 50a (Table II-l).
- R2Ol-RTmut-r50 was PCR amplified 3 times using reverse primer 28S 20a, 28S 40a and 28S 50a. PCR products were purified using a PCR clean up kit (sigma) and used for 5’ capped mRNA synthesis in vitro transcription using mMESSAGE mMACHINETM T7 Transcription Kit (Ambion).
- RNA based retrotransposition assay After mRNA synthesis, the template DNA was removed by DNase treatment and a >150 base poly(A) tail was added to RNA transcripts using The Poly(A) Tailing Kit (Ambion). Synthesized mRNA was precipitated with Lithium Chloride. mRNA based retrotransposition assay
- R2O1 mRNA Six types were used in this experiment.
- the linear template DNA was purified from PCR products of the plasmids which contain full-length of WT R2O1 or RTmut R2O1.
- the Capped mRNA was transcribed from the linear DNA template in vitro and transfected into 293 cells to verify the retrotransposition activity.
- Three types are WT R2O1 mRNA which had r4, rlO, and r71 28S flanking sequences at the end (FIG. 9A, Left).
- RTmut R2O1 mRNA which lost its reverse transcription ability
- r4, rlO, or r50 28 S flanking sequences at the end
- FIG. 9A Right
- the expected size of the band was observed at sample R201-r71, but not at R201-r4 or R201-rlO (FIG. 9B, Left panel).
- samples transfected with RTmut mRNA observed no band (FIG. 9B, Right panel), suggesting that R2O1 moves by TPRT and not by recombination events.
- PCR products were TA cloned and sequenced.
- mRNA-based retrotransposition system was tested in iPS cells. Also, 6 types R2O1 mRNA were used in this experiment. Capped mRNA was transcribed from full- length R2O1 construct or RTmut construct in vitro and transfected into iPS cells to verify the retrotransposition activity (FIG. 10A). Three types of mRNA are WT R2O1 mRNA with r4, rlO, and r50 28S flanking sequences at the end (FIG. 10A, Left). 3 types of RTmut R2O1 mRNA, which have r4, rlO, and r50 28S flanking sequences at the end were used as a negative control (FIG. 10A, Right).
- this example transfected those 3 types of WT R2O1 mRNA into iPS cells and extracted their genome after 6 h, 12 h, 24 h post-transfection respectively. After nested-PCR of 3 ’junction, the band of the expected size was only detected at samples that transfected with R2Ol-r50 mRNA after 24h, which suggests that R201 retrotransposition occurred at least after 24h post mRNA transfection.
- R2O1 showed the retrotransposition activity in human 293T cells and iPS cells via an mRNA-based in vivo retrotransposition assay, using mRNA-mediated delivery.
- the summary of sequences of the PCR products showed capped R2O1 mRNA with long 28S rDNA flanking sequences at the 3'-end inserted into the exact target site of 28S rDNA in 293 T cells and iPS cells .
- mRNA- mediated R2O1 retrotransposition may provide a new genome knock-in method in iPS cells, which is important for facilitating the development of gene therapy.
- Example 1 R2O1 constructs with 4 bp 28 S rDNA flanking sequence worked efficiently. This example further demonstrated that the retrotransposition activity was significantly improved by using capped R2O1 mRNA, which has 50 bp 28S rDNA flanking sequences at the 3-end rather than 4 bp or 10 bp. It is contemplated that instead of poly(A) tail, longer 28S rDNA flanking sequences at the 3 ’end of R2O1 mRNA had the added benefit of protecting the mRNA from degradation, which is important for its retrotransposition activity in the human cell.
- the retrotransposition assays were conducted in zebrafish embryos.
- the following mRNA constructs were prepared (see illustrations in FIG. 11), (A) the wild-type (WT) R2O1 retrotransposon with intact 5’UTR and 3’UTR, (B) the R2O1 retrotransposon with a poly(A) signal added to the 3’ end, (C) the R2O1 retrotransposon with four 5’ nucleotides of the 28 S (after insertion the site) (r4) added to the 3’ end, (D) the R2O1 retrotransposon with r4, following by a poly(A) signal, added to the 3’ end, (E) the R2O1 retrotransposon with 10 5’ nucleotides of the 28 S (after insertion the site) (rlO) added to the 3’ end, followed by a poly(A) signal, (F) the R2O1 retrotransposon with 100 5’ nucleotides of the 28 S (after insertion the
- FIG. 15 A The proposed procedure is illustrated in FIG. 15 A, where an R2O1 construct (an exogenous gene and the R2O1 3’UTR) and an R2O1 ORF-expressing construct were separately introduced to a target cell.
- the exogenous gene tested was EGFP, regulated by a SV40 promoter (FIG. 15B).
- FIG. 15C As shown in FIG. 15C, with the wild-type R2O1 ORF (not the inactive mutant) separately expressed in the target cell, the R2O1-EGFP construct was successfully integrated to the target genome and was expressed.
Landscapes
- Health & Medical Sciences (AREA)
- Genetics & Genomics (AREA)
- Life Sciences & Earth Sciences (AREA)
- Engineering & Computer Science (AREA)
- Zoology (AREA)
- Bioinformatics & Cheminformatics (AREA)
- Biomedical Technology (AREA)
- Organic Chemistry (AREA)
- Biotechnology (AREA)
- General Engineering & Computer Science (AREA)
- Chemical & Material Sciences (AREA)
- Wood Science & Technology (AREA)
- Microbiology (AREA)
- Physics & Mathematics (AREA)
- Plant Pathology (AREA)
- Molecular Biology (AREA)
- Biochemistry (AREA)
- General Health & Medical Sciences (AREA)
- Biophysics (AREA)
- Virology (AREA)
- Micro-Organisms Or Cultivation Processes Thereof (AREA)
Abstract
The present disclosure demonstrates that a newly discovered long interspersed nuclear element (LINE) R2 of medaka fish Oryzias latipes, R2Ol, can integrate into the 28S rDNA gene of a human cell in a target sequence-specific manner. With modification and/or addition of various non-coding sequences in the retrotransposon, its retrotransposition efficiency in human cells can be greatly improved. Further, additional exogenous genes can be included in the R2Ol retrotransposon, enabling the exogenous genes to be inserted into the target genome, achieving target sequence-specific gene delivery. Accordingly, the R2Ol retrotransposon systems of the present disclosure provide new and improved gene delivery approaches.
Description
LONG INTERSPERSED NUCLEAR ELEMENT R2 FOR TARGET-SPECIFIC
GENE DELIVERY
CROSS REFERENCE TO RELATED APPLICATIONS
[0001] This application claims the benefit under 35 U.S.C. § 119(e) of the United States Provisional Application Serial Nos. 63/499,626, 63/499,630, and 63/499,631, all filed May 2, 2023, the content of each of which is hereby incorporated by reference in its entirety.
REFERENCE TO AN ELECTRONIC SEQUENCE LISTING
[0002] The content of the electronic sequence listing (350643. xml; Size: 31,173 bytes; and Date of Creation: April 30, 2024) is herein incorporated by reference in its entirety.
BACKGROUND
[0003] The clustered regularly interspaced short palindromic repeat (CRISPR)/Cas9 mediated genome editing technology has seen wild use since its invention due to its simplicity and efficiency, leading to a new era of genetic engineering and gene therapy. The CRISPR/Cas9 system is composed of the Cas9 endonuclease and a single-guide RNA (sgRNA), which can guide Cas9 to the complementary sequence and induce double-strand break (DSB) at the target site. The resulting DSB is repaired independently by two cellular DNA repair pathways, namely non-homologous end-joining (NHEJ) pathway and homology directed repair (HDR) pathway. Genome editing using the CRISPR-Cas9 system in human embryos has resulted in many unexpected mutations at or near its target site as well as in the neighboring areas. Furthermore, this system depends on HDR, which is regulated in host cells, indicating that the integration process of the exogenous gene itself cannot be regulated autonomously.
[0004] The safe use of gene therapy in medical applications requires strict sequencespecificity without generating any off-target effects. Targeted gene knock-in must center on transgene integration at harmless genomic sites and avoid unpredictable phenotypes caused by unexpected integration.
[0005] Transposons, also referred to as TE or mobile genetic elements, are able to move from one chromosomal location to another. The two major groups of transposons are DNA transposons and retrotransposons. DNA transposons, or class II elements, move via a “cut and paste” mechanism, by directly removing themselves from the original position and
pasting at another position. Retrotransposons, or class I elements, move in a “copy and paste” manner, using RNA as an intermediate. Based on whether they are flanked by long terminal repeat (LTR) or not, retrotransposons are further classified as LTR type or non-LTR type. LTR retrotransposons are similar to retroviruses in structure and life cycle, and therefore their retrotransposition mechanisms are well-explained. However, the retrotransposition mechanisms of non-LTR type elements has not been well-clarified.
[0006] Non-LTR retrotransposons are sub-classified into two major groups, namely long interspersed nuclear elements (LINEs) and short interspersed nuclear elements (SINEs). LINEs are autonomous elements that encode proteins to mediate their own mobility, whereas SINEs are nonautonomous elements that do not encode protein and consequently require LINEs for their propagation.
[0007] Based on structural feature, LINEs can be categorized into two groups, an early branched group with a single open reading frame (ORF) and a recently branched group, with two ORFs (ORF1 and ORF2). The endonuclease (EN) domain, which cuts the target site DNA, and reverse transcriptase (RT) domain, which is responsible for reverse transcription of the RNA template, are the two domains essential for retrotransposition of a LINE. The EN and RT domains are encoded in the single ORF of an early-branched element or in ORF2 of a recently branched element, respectively.
[0008] Non-LTR retrotransposons are mobilized by a mechanism which is very different from that used by retroviruses and LTR retrotransposons. After transcription and translation, the resultant proteins (open reading frame proteins; ORFps) combine with their own mRNA in the cytoplasm to form a ribonucleoprotein (RNP) complex and move back to the nucleus, whereupon they are inserted into the target site via a mechanism termed target-primed reverse transcription (TPRT). The TPRT process is initiated by an encoded EN domain that nicks one strand of DNA at a target site and creates a 3 ’-hydroxyl end, which is used as a primer for reverse transcription of the LINE mRNA onto the DNA target via RT.
SUMMARY
[0009] The present disclosure demonstrates that a newly discovered long interspersed nuclear element (LINE) R2 of medaka fish Oryzias latipes, R2O1, can integrate into the 28S rDNA gene of a human cell in a target sequence-specific manner. With modification and/or addition of various non-coding sequences in the retrotransposon, its retrotransposition
efficiency in human cells can be greatly improved. Further, additional exogenous genes can be included in the R2O1 retrotransposon, enabling the exogenous genes to be inserted into the target genome, achieving target sequence-specific gene delivery. Accordingly, the R2O1 retrotransposon systems of the present disclosure provide new and improved gene delivery approaches.
[0010] According to one embodiment of the present disclosure, provided is a polynucleotide that is a messenger RNA (mRNA) or a DNA encoding the mRNA, wherein the mRNA comprises, from 5’ to 3’: a protein-coding sequence or a reverse-complement thereof; a 3’ untranslated region (UTR) having at least 85% sequence identity to SEQ ID NO: 16; and a downstream flanking sequence consisting of a fragment of SEQ ID NO:20 or an RNA sequence having least 85% sequence identity to SEQ ID NO:20, wherein the fragment includes at least 4 consecutive nucleotides of SEQ ID NO:20 starting at a position from 1 to 5. In some embodiments, the mRNA does not include a poly(A) signal.
[0011] In some embodiments, the 3 ’UTR is directly connected to the downstream flanking sequence. In some embodiments, the downstream flanking sequence has a length of 4 to 180 nucleotides (nt). In some embodiments, the downstream flanking sequence has a length of 10 to 150 nt. In some embodiments, the downstream flanking sequence has a length of 15 to 150 nt. In some embodiments, the downstream flanking sequence has a length of 40 to 90 nt. In some embodiments, the downstream flanking sequence has a length of 50 to 90 nt. In some embodiments, the downstream flanking sequence has a length of 71 to 90 nt.
[0012] In some embodiments, the fragment includes at least the four 5’ nucleotides (UAGC) of SEQ ID NO:20.
[0013] In some embodiments, the 3 ’UTR has at least 90% sequence identity to SEQ ID NO: 16. In some embodiments, the 3’UTR has at least 95% sequence identity to SEQ ID NO: 16. In some embodiments, the 3’UTR consists of SEQ ID NO: 16.
[0014] In some embodiments, the mRNA further comprises, 5’ to the protein-coding sequence, an upstream flanking sequence consisting of a fragment of SEQ ID NO: 18 or an RNA sequence having at least 85% sequence identity to SEQ ID NO: 18. In some embodiments, the fragment of the upstream flanking sequence ends at a position from 176 to 180 of SEQ ID NO: 18. In some embodiments, the upstream flanking sequence has a length of 4 to 180 nt. In some embodiments, the upstream flanking sequence has a length of 10 to
150 nt. In some embodiments, the upstream flanking sequence has a length of 32 to 150 nt. In some embodiments, the upstream flanking sequence has a length of 20 to 100 nt. In some embodiments, the upstream flanking sequence has a length of 32 to 80 nt.
[0015] In some embodiments, the mRNA further comprises, 5’ to the protein-coding sequence, a 5’ untranslated region (UTR) comprising nucleotides 233 to 265 of SEQ ID NO: 14 or a nucleic acid sequence having at least 85% sequence identity to nucleotides 233 to 265 of SEQ ID NO: 14. In some embodiments, the mRNA does not include nucleotides 1 to 87 of SEQ ID NO: 14. In some embodiments, the mRNA does not include nucleotides 1 to 175 of SEQ ID NO: 14. In some embodiments, the mRNA does not include nucleotides 1 to 198 of SEQ ID NO: 14.
[0016] In some embodiments, the 5 ’UTR comprises nucleotides 216 to 265 of SEQ ID NO: 14 or a nucleic acid sequence having at least 85% sequence identity to nucleotides 216 to 265 of SEQ ID NO: 14.
[0017] In some embodiments, the upstream flanking sequence is located 5’ to the 5 ’UTR.
[0018] In some embodiments, the protein-coding sequence encodes a Long Interspersed Nuclear Element (LINE) R2 of medaka fish Oryzias latipes (R2O1) open reading frame (ORF), which can be expressed by the mRNA. In some embodiments, the ORF comprises the amino acid sequence of SEQ ID NO: 15 or an amino acid sequence having at last 85% sequence identity to SEQ ID NO: 15.
[0019] In some embodiments, the protein-coding sequence encodes an exogenous therapeutic protein for delivery to a target cell.
[0020] In some embodiments, the mRNA further comprises a second protein-encoding sequence or the reverse-complement thereof, wherein the second protein-encoding sequence encodes an exogenous therapeutic protein for delivery to a target cell.
[0021] Also provided, in one embodiment, is a method for integrating an exogenous polynucleotide to the genome of a mammalian cell, comprising introducing to the cell the polynucleotide of the present disclosure. In some embodiments, the polynucleotide is a mRNA, a plasmid or an adenoviral vector. In some embodiments, the mammalian cell is a human cell. In some embodiments, the human cell is an induced pluripotent stem cell (iPSC).
[0022] In some embodiments, wherein the polynucleotide is mRNA and the downstream flanking sequence has a length of 40 to 90 nt. In some embodiments, the polynucleotide is a plasmid or an adenoviral vector and the downstream flanking sequence has a length 4 to 90 nt. In some embodiments, the human cell is not a stem cell. In some embodiments, the downstream flanking sequence has a length of 70 to 90 nt. In some embodiments, the polynucleotide or the reverse-complement thereof encodes a protein that is not an R2O1 ORF, and the method further comprises introducing to the mammalian cell a second polynucleotide encoding the R2O1 ORF.
BRIEF DESCRIPTION OF THE DRAWINGS
[0023] FIG. 1. The structure and retrotransposition mechanism of R2O1. (A) A schematic representation of R2O1. R2O1, cloned from the medaka fish, Oryzias latipes. It comprises a 5' untranslated region (UTR; 265 bp), a single open reading frame (ORF; 3831 bp) encoding EN and RT domains, and a 3' UTR (108 bp). R2O1 is inserted into the 28S rDNA target sequence. The rDNA unit is indicated by a black closed rectangle. The double-stranded sequences of the precise insertion sites are indicated (upper: 5’- ACTCTCTTAAGGTAGCCAAATGCC-3’, SEQ ID NO:4; lower: 5’-
GGCATTTGGCTACCTTAAGAGAGT-3’, SEQ ID NO:5). Bottom and top strand cleavage sites generated by EN are indicated by vertical arrows. (B) A schematic representation of the R2O1 target-primed reverse transcription (TPRT) mechanism. The EN domain cleaves the bottom strand of genomic DNA. R2O1 mRNA anneals with the target DNA (28S rDNA) at the 3' junction to synthesize cDNA (5’-TCTTAAGGTAGCCAAATGCC, SEQ ID NO:6; 5’- GGCATTTGCTA-3’, SEQ ID NO:7; 5’-CCTTAAGA-3’, SEQ ID NO:8; 5’- CUCAAGUAGC-3, SEQ ID NO: 9).
[0024] FIG. 2. Locations of primer sets used for the detection of R2O1 retrotransposition. Nested PCR was used to detect the junction site between inserted R2O1 and 28SrDNA. Horizontal arrows indicate the PCR primers used to detect retrotransposition events. Primer sets “g,” “h,” “i,” and “j” were used for 3' junction analysis, while the primer sets “f”, “al”, “a2”, “a3”, “a4” and “a5” were used in 5' junction analysis. The rDNA unit is indicated by a black closed rectangle. The double-strand sequences of the precise insertion sites are indicated (upper: SEQ ID NO:4; lower: SEQ ID NO:5). Bottom and top strand cleavage sites generated by endonuclease (EN) domain are indicated by vertical arrows.
[0025] FIG. 3A-D. 3' junction PCR of inserted R201-r4. (A) Schematic representation of R2O1 used in the human cell retrotransposition assay. R2O1 contains a 4-bp target sequence downstream of the 3' UTR. CAG, chicken beta-actin promoter and cytomegalovirus immediate-early enhancer-promoter. R2O1- ENMut mutated at the amino acid sites essential for EN, completely abolishing the retrotransposition activities. pA, poly A signal. (B) Horizontal arrows indicate PCR primers used to detect retrotransposition events. (C) PCR analysis of the 3' junction region to detect retrotransposition in 293T cells after infection of cells at different MOIs. The predicted PCR band sizes were: 327 bp under primer set h & i; and 480 bp under primer set g & j, indicated by solid arrowheads, respectively. The PCR products indicated by a triangle mark were TA cloned for further analyses. (D) Endonuclease (EN) mutant abolishes R2O1 retrotransposition. R201-ENmut is a construct in which the EN portion of the 1149th amino acid (IPD — IP A) has been mutated (indicated by the dashed line). N, negative-control PCR conducted with genomic DNA from non-infected cells. Non-specific sequences indicated by open arrowheads. Numbers above the lanes indicate independent experiments.
[0026] FIG 4. 5' junction PCR of inserted R201-r4. Horizontal arrows indicate PCR primers used to detect 5' junctions; “f” indicates the forward primer set upstream of the target site at 28S rDNA; “al”, “a2”, “a3”, “a4”, and “a5” are the five reverse primer sets at the R2O1 sequences.
[0027] FIG. 5. 5' junction analysis around the insertion site of R201-r4. The 5' junction region around the insertion site of R201-r4. The top of the figure includes a diagram of the 28 S target site, indicated by black boxes, with R201-r4 sequences in the middle, indicated by red. Numbers on the left/right (-/+) of each region indicate the position of the accurate cleavage site of the target DNA regions. The accurate reverse transcriptionending site of the sequence (in bases) is indicated by the number on the top of the diagram. Extra sequences derived from the vector sequences at the junction are indicated by white triangles.
[0028] FIG. 6A-B. 3' junction analysis of inserted r32R201-r4. (A) Schematic representation of r32R201-r4 used in the human cell retrotransposition assay. R2O1 contains 32 bp 28S flanking sequences upstream of the 5' UTR; pA, poly A signal. (B) 3' junction region PCR analysis to detect retrotransposition in 293T cells after
infection of cells at different MOIs. The predicted PCR band size of 480 bp is indicated by the arrowhead. Numbers above the lanes indicate independent experiments; N, negative-control. PCR conducted with genomic DNA from non-infected cells.
[0029] FIG. 7. 5' junction analysis around the insertion site of r32R201-r4. 5' junction region around the insertion site of r32R201-r4. The top of the figure includes a diagram of 28 S target site indicated by black boxes, with the R201-r4 sequences in the middle, indicated by gray. Numbers on the left/right (-/+) of each region indicate the position of the accurate cleavage site of the target DNA regions. The accurate reverse transcriptionending site of the sequence (in nucleotide bases) is indicated by the number on the top of the diagram. Extra sequences derived from vector sequences at the junction are indicated by white triangles.
[0030] FIG. 8A-C. 3' junction analysis of inserted r32R201-r4. (A) Schematic representation of plasmids used in the human cell retrotransposition assay. All plasmids contained a 4-bp target sequence downstream of the 3' UTR.; CAG, chicken beta-actin promoter and cytomegalovirus immediate-early enhancer promoter. The RTmut construct was mutated at the essential amino acid site for RT; pA, poly A signal. (B) The 3' junction region PCR analysis to detect retrotransposition in 293T cells after infecting cells at different MOIs. The predicted PCR band of 480 bp is indicated by the arrowhead. Numbers above the lanes indicate independent experiments; N, negative-control; PCR conducted with genomic DNA from non-infected cells. (C) The 3' junction sequence of the inserted R2O1 plasmid (5’-
GCGAGTGGTGACTCGCCTCAAGTAGCCAAATGCCTCGTCATCT-3’, SEQ ID NO: 10; 5’- GGCATGATTACAAATCTT-3’, SEQ ID NO: 11; 5’-CCTCGTCATCT-3’, SEQ ID NO: 12). The nucleic acid sequences of the PCR product were analyzed through direct sequencing. Shown on top is a diagram of the correct insertion of R2O1 3' end. R2O1 sequences are shown on the left and 28 S target DNA sequences are shown on the right. The initiation sites of reverse transcription are indicated by nucleotide numbers. The cleavage site at 28S, which was nicked by the EN domain, was measured in bases from 5' to 3'; minus indicates downstream of the target.
[0031] FIG. 9A-B mRNA-mediated R2O1 retrotransposition in 293T cells. (A) Schematic representation of mRNA transcription in vitro. pGEM-rl05-R2Ol-rl00 contains a T7 promoter, a WT R2O1, and fused with a 100 bp downstream 28S rDNA sequence of R2O1 insertion site at the 3 '-end. There were 61 bp vector sequences between R2O1 ORF and R2O1 3' UTR. pA; poly A signal. pGEM-R201 RTmut containing T7 promoter and R2O1 RTmut. (i) Template DNA for in vitro transcription was amplified via PCR. The template at the left (R201-r4, R201-rlO and R201-r71) indicates the WT with 28S downstream sequences of different lengths (4 bp/10 bp/71 bp) of the insertion site attached to the end. The template at the right (RTmut-r4, RTmut-rlO and RTmut-r71) indicates the RT with 28S target sequences of the same length as in WT templates (4 bp/10 bp/50 bp) attached to the end. (ii) Six types of 5' capped mRNA were synthesized in vitro. (B) 3' junction region PCR analysis aimed at detecting retrotransposition in 293T cells after a 72 h transfection. The predicted PCR band of size 541 bp (contained 61 bp vector sequences between R2O1 ORF and R2O1 3' UTR) is indicated by the arrowhead; N, negative-control PCR conducted with genomic DNA from non-infected cells.
[0032] FIG. 10A-B In vitro synthesis of mRNA used for iPS cell transfection. (A) Schematic representation of mRNA transcription in vitro pGEM-rl05-R2Ol-rl00 containing T7 promoter; WT R2O1 fused with 100 bp 28 S rDNA target site at the 3 end. There are 61 bp vector sequences between R2O1 ORF and R2O1 3 'UTR; pA, poly A signal; pGEM-R201 RTmut containing T7 promoter and R2O1 RTmut. The template DNA for in vitro transcription was amplified via PCR. The template at the left (R201-r4, R201-rlO and R2O1- r50) indicates WTs with 28S target sequences of different lengths (4 bp/10 bp/50 bp) attached to the end. The templates at the right (RTmut-r4, RTmut-rlO and RTmut-r50) show RTs with 28S target sequences of the same length as in WT templates (4 bp/10 bp/50 bp) attached to the end. (ii) The six types of 5' capped mRNA were synthesized in vitro. (B) 3' junction region PCR analysis aimed at detecting retrotransposition in iPS cells 72 hours after transfection. The predicted PCR band of size 541 bp (contained 61 bp vector sequences between R2O1 ORF and R2O1 3 UTR) is indicated by the arrowhead. Numbers above the lanes indicate independent experiments; N, negative-control PCR conducted with genomic DNA from non-infected cells.
[0033] FIG. 11 illustrates R2O1 constructs with various non-coding sequences added or deleted. The retrotransposition efficiency testing results show that neither the 5 ’UTR nor the
poly(A) signal was required for retrotransposition. The addition of a short 28 S downstream sequences, however, improved retrotransposition.
[0034] FIG. 12 shows design and testing of R2O1 constructs with 4 consecutive nucleotides of the 28S downstream sequence added to the 3’. When the 4 consecutive nucleotides are close (<6 bases) to the target insertion site, it promoted more efficient retrotransposition than without the short nucleotides.
[0035] FIG. 13A shows construction of R2O1 retrotransposons with different lengths of the 5’UTR deleted and their impact on retrotransposition efficiency. A deletion between 175 and 198 nucleotides of the 5’ portion of the 5’UTR exhibited the highest retrotransposition efficiency.
[0036] FIG. 13B shows construction of R2O1 retrotransposons with different lengths of the 3’UTR deleted and their impact on retrotransposition efficiency. The intact 3’UTR resulted in the highest retrotransposition efficiency.
[0037] FIG. 14A-B show construction and testing of R2O1 constructs with exogenous genes inserted downstream of the ORF. Both tested exogenous genes, EGFP and Gal4, were successfully inserted to the target 28 S rDNA and expressed.
[0038] FIG. 15A-C show that the R2O1 ORF can be expressed separately to enable retrotransposition of a R2O1 construct that embeds an exogenous gene.
[0039] It will be recognized that some or all of the figures are schematic representations for purpose of illustration.
DETAILED DESCRIPTION
[0040] The following description sets forth exemplary embodiments of the present technology. It should be recognized, however, that such description is not intended as a limitation on the scope of the present disclosure but is instead provided as a description of exemplary embodiments.
Long Interspersed Nuclear Element (LINE) R2 for Target-Specific Gene Delivery
[0041] Most LINEs are randomly inserted throughout the host genome, such as LI and L2. Some elements are inserted into specific sites of repetitive genomic sequences, such as
ribosomal DNA (rDNA), telomeric repeats, and microsatellites. Among the site-specific LINEs, the R2 elements specifically insert into the 28 S rDNA.
[0042] The instant inventors found that R2 elements are widely distributed among vertebrate species, such as fish and birds but not in mammals. In addition, an approximately 100-base pair (bp) region around the R2 insertion site in 28S rDNA is strongly conserved across a wide variety of eukaryotes, suggesting the high sequence specificity of this element.
[0043] R2Bm, isolated from Bombyx mori, exhibited efficient site-specific retrotransposition activity both in vitro and in vivo, thereby facilitating a better understanding of the general mechanism underlying LINE retrotransposition. R2O1, isolated from the medaka fish, Oryzias latipes, includes a 5' untranslated region (UTR; 265 nt), a single open reading frame (ORF; 1276 amino acids) encoding the endonuclease (EN) and reverse transcriptase (RT) domains, and a 3' UTR (111 nt) (FIG. 1A). The 28S rDNA target sequence and the exact cleavage sites of R2O1 on both strands are shown in FIG. IB. R2O1 mRNA anneals with the target site and initiates the reverse transcription mechanism.
[0044] Sequence analysis shows that R2 elements are not present in the genomes of any mammals, including humans and mice. Considering that the R2 elements in the genomes of human cells have been lost during evolution, the inventors compared the 450 bp 28S rDNA sequence around the putative R2 insertion site in humans with those in medaka and zebrafish (Table 1) Table 1 shows R2 insertion site sequence alignment of three vertebrates, medaka fish (CP020779.1, SEQ ID NO:3), zebrafish (BX537263.17, SEQ ID NO:2), and humans (Ml 1167.1, SEQ ID NO: 1). An approximately 300 base pair (bp) region around the R2 insertion site in 28S rDNA is strongly conserved. A large deletion region was observed at approximately 200-bp downstream of the R2 insertion site in medaka and zebrafish. The insertion site (red) of R2 is between -1 (G) and +1(T).
Table 1. Comparison between the 28S rDNA sequences around the insertion site of R2 element of medaka fish (SEQ ID NO:3), zebrafish (SEQ ID NO:2), and humans (SEQ ID NO:1)
[0045] Despite large deletion regions approximately 200 bp downstream of the R2 insertion site in medaka and zebrafish, the 300 bp sequences around the R2 insertion sites of these three species were found to be completely identical, except for five different nucleotides (5 SNPs). This indicated that the 28S rDNA target sequence is retained in human cells, which had lost the R2 element in the distant past, suggesting that theoretically R2 can integrate into the human 28S rDNA. There has been no report on, however, whether R2O1 can be used to conduct transgenic integration in mammalian cells, including human cells.
[0046] As demonstrated in the appended experimental examples, an adenovirus-mediated R2O1 retrotransposition system was efficient at integrating to the 28 S rDNA gene in human 293T cells. Moreover, mRNA-mediated R2O1 retrotransposition systems could readily integrate to 28S gene in 293T as well as in induced pluripotent stem cells (iPSC).
[0047] Also interestingly, the instant inventors designed and tested many R2O1 constructs with changes to the non-coding regions, and discovered that the retrotransposition efficiency can be greatly improved with appropriate modification of these regions.
[0048] As noted, the R2O1 (FIG. 1A, Table 2, SEQ ID NO: 13) includes a 5' untranslated region (UTR; 265 nt; SEQ ID NO: 14), a single open reading frame (ORF; 1276 amino acids; SQE ID NO: 15) encoding the endonuclease (EN) and reverse transcriptase (RT) domains, and a 3' UTR (108 nt; SEQ ID NO: 16). As shown in Example 3, the 3’UTR is required for efficient retrotransposition (FIG. 13B). Retaining of some portions of the 5 ’UTR, by contrast, actually reduced the retrotransposition efficiency.
[0049] As shown in FIG. 13A, compared to R2O1 having the intact 5’UTR (R2O1 + 4bp), an R2O1 with the first 175 nucleotides deleted (R2O1 Al-175 + 4 bp) actually had greatly increased retrotransposition efficiency. The R2O1 retained high retrotransposition efficiency
even when the first 215 (or even 232) nucleotides were removed. These data, therefore, demonstrate that the majority of the 5’ portion of the 5’UTR is detrimental to the retrotransposition activity of R2O1.
[0050] As shown in Examples 1 and 2, the R2O1 retrotransposon is capable of target sequence-specific integration. More specifically, in human cells, the R2O1 can insert itself to between 3746 and 3747 of the 28 S gene (and between 3744 and 3745 on the opposite strand) (FIG. IB, Table 3, SEQ ID NO 21).
[0051] The four-nucleotide rDNA sequences immediately downstream the insertion site are ATCG (nucleotides 3747-3750 of SEQ ID NO:21; or nucleotides 1-4 of SEQ ID NO: 19). As shown in the examples, inclusion of these 4 nucleotides (referred to as r4) at the 3’ end of the R2O1 can greatly increase its retrotransposition efficiency. Further testing (e.g., FIG. 12) shows that actually any 4-consecutive nucleotides within the first 8 nucleotides of this “downstream sequence” (SEQ ID NO: 19 or 20) can have similar results.
[0052] Also important, in certain situations, depending on the type of polynucleotide or vector used for delivery, inclusion of different lengths of this downstream sequence (SEQ ID NO: 19 or 20) can have different benefits. For instance, while r4 was sufficient for an adenovirus-mediated R2O1 retrotransposition in human cells, when mRNA was directly used, the required downstream sequence was much longer. As shown in FIG. 9B, for 293T cells, the R2O1 mRNA required 71 (r71) nucleotides of SEQ ID NO:20 (> rlO and r4); for iPSC (FIG. 10B), the R2O1 mRNA required 50 (r50) nucleotides of SEQ ID NO:20 (> rlO and r4).
[0053] Likewise, it was discovered that inclusion of a portion of the 28S “upstream sequence” (SEQ ID NO: 17 or 18) of the integration site at the 5’ end of the R2O1 can also be beneficial. As shown in Example 1 and FIG. 6A-B, when a 32-nucleotide fragment of the upstream sequence (149-180, ending at the integration site; referred to as r32) was added to the 5’ end of the R2O1, the retrotransposition efficiency was also increased.
[0054] Not only can the R2O1 retrotransposon integrate itself to a target sequence in the human genome, but it can also further include additional coding sequences so as to achieve delivery of
other genes. In Example 4, an EGFP and a Gal4 encoding sequence were inserted to the R2O1 retrotransposon, downstream of the R2O1 ORF. Both exogenous genes were integrated to the target genome and expressed (FIG. 14A-B). Note that both genes were inserted backwards into the retrotransposon such that they are not functionally expressed with the R2O1 mRNA, and are only expressed after genomic integration.
[0055] Finally, as shown in Example 5, the R2O1 ORF does not need to be included in the retrotransposon itself in order to enable retrotransposition. Therefore, if the gene to be delivered is very large, the R2O1 retrotransposon can just include that gene, while the R2O1 ORF can be delivered separately (FIG. 15A). This way, the technology can allow better control, and delivery of very large exogenous genes to a target cell.
[0056] The instant data, therefore, demonstrate that the R2O1 transposition system can be suitably used to deliver an exogenous coding sequence to a target cell. Since the integration is target sequence-specific, such a gene delivery technology is highly safe.
[0057] According to one embodiment of the present disclosure, therefore, provided is a polynucleotide that is a messenger RNA (mRNA) or a DNA encoding the mRNA. In one embodiment, the mRNA includes, from 5’ to 3’ (A) a protein-coding sequence or a reversecomplement thereof; (B) a 3’ untranslated region (UTR); and (C) a downstream flanking sequence. In some embodiments, the mRNA further includes (D) an upstream flanking sequence that is 5’ to (A). In some embodiments, the mRNA further includes (E) a 5’ UTR that is 5’ to (A). In some embodiments, the mRNA includes, from 5’ to 3’, (D), (E), (A), (B), and (C). In some embodiments, the mRNA includes, from 5’ to 3’, (D), (A), (B), and (C). In some embodiments, the mRNA includes, from 5’ to 3’, (E), (A), (B), and (C). In some embodiments, the mRNA includes, from 5’ to 3’, (A), (B), and (C). In some embodiments, the mRNA does not include a poly(A) signal. In some embodiments, the mRNA includes a 5’ Cap. In some embodiments, the mRNA does not include any additional sequence or element not specifically recited herein.
3’ Untranslated Region (UTR)
[0058] As provided, the 3’ untranslated region (UTR) of the R2O1 retrotransposon plays an important role in the retrotransposition. In some embodiments, the 3 ’UTR has the nucleic acid sequence of SEQ ID NO: 16. In some embodiments, the 3’UTR has at least 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, 97%, 98%, 99% or 100% sequence identity to SEQ ID
Downstream Flanking Sequence
[0059] The downstream flanking sequence, in one embodiment, refers to a fragment of the 28S (e.g., mammalian 28S such as human 28S) sequence that is downstream of the R2O1 integration site. An example 180-nt sequence is provided in SEQ ID NO: 19 (DNA) or SEQ ID NO:20 (RNA).
[0060] In some embodiments, the fragment has a length that is at least 4 nt. In some embodiments, the fragment has a length that is at least 5, 10, 15, 20, 25, 30, 35, 40, 45, 50, 55, 60, 65, 70, 71, 75, 80, 85, or 90 nt. In some embodiments, the fragment has a length that is not longer than 10, 15, 20, 25, 30, 35, 40, 45, 50, 55, 60, 65, 70, 71, 75, 80, 85, 90 or 100 nt. In some embodiments, the length is 4-180, 10-150, 15-150, 40-90, 50-90, 70-90 or 71-90 nt, without limitation.
[0061] In some embodiments, the fragment is a fragment of SEQ ID NO:20. In some embodiments, the fragment is a fragment of an RNA sequence having least 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, 97%, 98%, 99% or 100% sequence identity to SEQ ID NO:20.
[0062] In some embodiments, the fragment starts at position, according to SEQ ID NO:20, that is within positions 1-20, 1-15, 1-10, or 1-5. In some embodiments, the fragment includes at least 4 consecutive nucleotides of SEQ ID NO:20 starting at a position from 1 to 5. In some embodiments, the fragment includes at least the first 4 nucleotides of SEQ ID NO:20 (UAGC).
[0063] In some embodiments, the 3’UTR is directly connected to the downstream flanking sequence.
Upstream Flanking Sequence
[0064] In some embodiments, the polynucleotide (e.g., mRNA) further includes an upstream flanking sequence that is 5’ to the protein-coding sequence. The upstream flanking sequence, in one embodiment, refers to a fragment of the 28 S (e.g., mammalian 28 S such as human 28S) sequence that is upstream of the R2O1 integration site. An example 180-nt sequence is provided in SEQ ID NO: 17 (DNA) or SEQ ID NO: 18 (RNA).
[0065] In some embodiments, the fragment has a length that is at least 4 nt. In some embodiments, the fragment has a length that is at least 5, 10, 15, 20, 25, 30, 32, 35, 40, 45, 50, 55, 60, 65, 70, 75, 80, 85, or 90 nt. In some embodiments, the fragment has a length that
is not longer than 10, 15, 20, 25, 30, 35, 40, 45, 50, 55, 60, 65, 70, 71, 75, 80, 85, 90 or 100 nt. In some embodiments, the length is 4-180, 10-150, 15-150, 32-150, 20-100, 32-80, 40-90, 50-90, 70-90 or 71-90 nt, without limitation.
[0066] In some embodiments, the fragment is a fragment of SEQ ID NO: 18. In some embodiments, the fragment is a fragment of an RNA sequence having least 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, 97%, 98%, 99% or 100% sequence identity to SEQ ID NO: 18.
[0067] In some embodiments, the fragment ends at position, according to SEQ ID NO: 18, that is within positions 161-180, 166-180, 170-180, or 176-180. In some embodiments, the fragment includes at least 4 consecutive nucleotides of SEQ ID NO: 18 ending at a position from 176 to 180.
5’ Untranslated Region (UTR)
[0068] In some embodiments, the polynucleotide (e.g., mRNA) further includes a 5’ untranslated region (UTR) 5’ to the protein-coding sequence. When an upstream flanking sequence is also included, preferably the upstream flanking sequence is 5’ to the 5 ’UTR.
[0069] The wild-type R2O1 5’UTR sequence is provided in SEQ ID NO: 14. As shown in the data, the 5’UTR sequence is not essential to the retrotransposition activity of the R2O1 retrotransposon. Moreover, the 5’ portion of the 5’UTR appeared to inhibit the retrotransposition activity. In some embodiments, therefore, when the 5’UTR is included, it preferably includes nucleotides 233 to 265 of SEQ ID NO: 14 or a nucleic acid sequence having at least 85% sequence identity to nucleotides 233 to 265 of SEQ ID NO: 14. In some embodiments, the 5’UTR includes nucleotides 216 to 265 of SEQ ID NO: 14 or a nucleic acid sequence having at least 85% sequence identity to nucleotides 216 to 265 of SEQ ID NO: 14.
[0070] In some embodiments, the polynucleotide (e.g., mRNA) does not include nucleotides 1 to 87 of SEQ ID NO: 14. In some embodiments, the polynucleotide (e.g., mRNA) does not include nucleotides 1 to 175 of SEQ ID NO: 14. In some embodiments, the polynucleotide (e.g., mRNA) does not include nucleotides 1 to 198 of SEQ ID NO: 14. In some embodiments, the polynucleotide (e.g., mRNA) does not include nucleotides 88 to 175 of SEQ ID NO: 14.
Protein-Coding Sequence
[0071] The wild-type R2O1 includes an R2O1 ORF that has the required EN and RT activities. It has been shown, however, that for the R2O1 retrotransposon to work, the ORF can be expressed from a separate polynucleotide, which enables the “empty” R2O1 retrotransposon to include a large exogenous gene for delivery to the target cell. Accordingly, the R2O1 retrotransposon of the instant disclosure can include a coding sequence that encodes the R2O1, the exogenous gene for delivery, or the combination thereof.
[0072] In one embodiment, the protein-coding sequence encodes a Long Interspersed Nuclear Element (LINE) R2 of medaka fish Oryzias latipes (R2O1) open reading frame (ORF), which can be directly expressed by the mRNA (e.g., in the correct orientation for translation). In some embodiments, the ORF comprises the amino acid sequence of SEQ ID NO: 15 or an amino acid sequence having at last 85% sequence identity to SEQ ID NO: 15.
[0073] In some embodiments, the polynucleotide (e.g., mRNA) further includes a second protein-coding sequence which encodes an exogenous protein for delivery to a target cell. In some embodiments, the second protein-coding sequence is 3’ to the R2O1 ORF-coding sequence. In some embodiments, the second protein-coding sequence is included in the mRNA in a reverse-complement orientation such that it cannot be directly translated by the mRNA. Rather it is translated after integration to the target genome.
[0074] In some embodiments, the polynucleotide (e.g., mRNA) does not encode the R2O1 ORF, and instead encodes an exogenous protein to be expressed in a target cell. In some embodiments, this coding sequence is included in the correct orientation for translation. In some embodiments, it is in a reverse-complement orientation.
[0075] In some embodiments, the protein is a therapeutic protein that can be helpful in treating a disease or condition in a patient. Examples include:
- Insulin - used to treat diabetes
- Phenylalanine hydroxylase - used to treat PKU (Phenylketonuria) disease
- Erythropoietin - used to treat anemia
- Filgrastim (G-CSF) - used to treat neutropenia
- Interferons - used to treat cancer, multiple sclerosis, and viral infections
- Tumor necrosis factor (TNF) inhibitors - used to treat autoimmune diseases, such as rheumatoid arthritis and Crohn's disease
- Monoclonal antibodies - used to treat cancer, autoimmune diseases, and some viral infections
- Follitropin alfa - used to treat infertility
- Coagulation factor - used to treat bleeding disorders, such as Hemophilia
- Enzyme replacement therapy enzymes - used to treat lysosomal storage disorders, such as Gaucher's disease
- Growth hormone - used to treat growth hormone deficiency.
Gene Delivery and Treatments
[0076] As demonstrated, the R2O1 retrotransposition systems disclosed here are suitable for delivering exogenous genes to a target cell, such as mammalian cells. As the delivery results in target sequence (28S rDNA)-specific integration, it can be highly efficient and safe. This new technology, therefore, has promising therapeutic uses as well.
[0077] Accordingly, in embodiment, the present disclosure provides a method for integrating an exogenous polynucleotide to the genome of a mammalian cell. In one embodiment, the method entails introducing to the cell a R2O1 retrotransposon of the instant disclosure. In some embodiments, the retrotransposon is delivered to the target cell as an mRNA molecule directly.
[0078] mRNAs may be synthesized according to any of a variety of known methods. For example, the mRNAs may be synthesized via in vitro transcription (IVT). Briefly, IVT is typically performed with a linear or circular DNA template containing a promoter, a pool of ribonucleotide triphosphates, a buffer system that may include DTT and magnesium ions, and an appropriate RNA polymerase (e.g., T3, T7 or SP6 RNA polymerase), DNase I, pyrophosphatase, and/or RNase inhibitor. The exact conditions will vary according to the specific application.
[0079] The mRNA may be synthesized as unmodified or modified mRNA. Typically, mRNAs are modified to enhance stability. Modifications of mRNA can include, for example, modifications of the nucleotides of the RNA. A modified mRNA can thus include, for example, backbone modifications, sugar modifications or base modifications. In some embodiments, antibody encoding mRNAs (e.g., heavy chain and light chain encoding mRNAs) may be synthesized from naturally occurring nucleotides and/or nucleotide
analogues (modified nucleotides) including, but not limited to, purines (adenine (A), guanine (G)) or pyrimidines (thymine (T), cytosine (C), uracil (U)), and as modified nucleotides analogues or derivatives of purines and pyrimidines, such as e.g. 1-methyl-adenine, 2-methyl- adenine, 2-methylthio-N-6-isopentenyl-adenine, N6-methyl-adenine, N6-isopentenyl- adenine, 2-thio-cytosine, 3-methyl-cytosine, 4-acetyl-cytosine, 5-methyl-cytosine, 2,6- diaminopurine, 1-methyl-guanine, 2-methyl-guanine, 2,2-dimethyl-guanine, 7-methyl- guanine, inosine, 1-methyl-inosine, pseudouracil (5-uracil), dihydro-uracil, 2-thio-uracil, 4- thio-uracil, 5-carboxymethylaminomethyl-2-thio-uracil, 5-(carboxyhydroxymethyl)-uracil, 5- fluoro-uracil, 5-bromo-uracil, 5-carboxymethylaminomethyl-uracil, 5-methyl-2-thio-uracil, 5- methyl-uracil, N-uracil-5-oxyacetic acid methyl ester, 5-methylaminomethyl-uracil, 5- methoxyaminomethyl-2 -thio-uracil, 5 ’ -methoxy carbonylmethyl-uracil, 5-methoxy -uracil, uracil-5-oxyacetic acid methyl ester, uracil-5-oxyacetic acid (v), 1-methyl-pseudouracil, queosine, 13-D-mannosyl-queosine, wybutoxosine, and phosphoramidates, phosphorothioates, peptide nucleotides, methylphosphonates, 7-deazaguanosine, 5- methylcytosine and inosine. The preparation of such analogues is known to a person skilled in the art e.g. from the U.S. Pat. Nos. 4,373,071, 4,401,796, 4,415,732, 4,458,066, 4,500,707, 4,668,777, 4,973,679, 5,047,524, 5,132,418, 5,153,319, 5,262,530 and 5,700,642, the disclosure of which is included here in its full scope by reference.
[0080] In some embodiments, the mRNAs may contain RNA backbone modifications. Typically, a backbone modification is a modification in which the phosphates of the backbone of the nucleotides contained in the RNA are modified chemically. Exemplary backbone modifications typically include, but are not limited to, modifications from the group consisting of methylphosphonates, methylphosphoramidates, phosphoramidates, phosphorothioates (e.g. cytidine 5 ’-O-(l -thiophosphate)), boranophosphates, positively charged guanidinium groups etc., which means by replacing the phosphodiester linkage by other anionic, cationic or neutral groups.
[0081] In some embodiments, the mRNAs may contain sugar modifications. A typical sugar modification is a chemical modification of the sugar of the nucleotides it contains including, but not limited to, sugar modifications chosen from the group consisting of 2’- deoxy-2’-fluoro-oligoribonucleotide (2’-fluoro-2’-deoxycytidine 5 ’-triphosphate, 2’-fluoro- 2’-deoxyuridine 5 ’-triphosphate), 2’-deoxy-2’-deamine-oligoribonucleotide (2’-amino-2’- deoxycytidine 5 ’-triphosphate, 2’ -amino-2’ -deoxyuridine 5 ’-triphosphate), 2’-O-
alkyloligoribonucleotide, 2’-deoxy-2’-C-alkyloligoribonucleotide (2’-O-methylcytidine 5’- triphosphate, 2’-methyluridine 5 ’-triphosphate), 2’-C-alkyloligoribonucleotide, and isomers thereof (2’-aracytidine 5 ’-triphosphate, 2’-arauridine 5 ’-triphosphate), or azidotriphosphates (2’ -azido-2’ -deoxy cytidine 5 ’-triphosphate, 2’-azido-2’-deoxyuridine 5 ’-triphosphate).
[0082] In some embodiments, the mRNAs may contain modifications of the bases of the nucleotides (base modifications). A modified nucleotide which contains a base modification is also called a base-modified nucleotide. Examples of such base-modified nucleotides include, but are not limited to, 2-amino-6-chloropurine riboside 5 ’-triphosphate, 2- aminoadenosine 5 ’-triphosphate, 2-thiocytidine 5 ’-triphosphate, 2-thiouridine 5’- triphosphate, 4-thiouridine 5 ’-triphosphate, 5-aminoallylcytidine 5 ’-triphosphate, 5- aminoallyluridine 5 ’-triphosphate, 5-bromocytidine 5 ’-triphosphate, 5 -bromouridine 5’- triphosphate, 5-iodocytidine 5 ’-triphosphate, 5-iodouridine 5 ’-triphosphate, 5-methylcytidine 5 ’-triphosphate, 5-methyluridine 5 ’-triphosphate, 6-azacytidine 5 ’-triphosphate, 6-azauridine 5 ’-triphosphate, 6-chloropurine riboside 5 ’-triphosphate, 7-deazaadenosine 5 ’-triphosphate, 7-deazaguanosine 5 ’-triphosphate, 8-azaadenosine 5 ’-triphosphate, 8-azidoadenosine 5’- triphosphate, benzimidazole riboside 5 ’-triphosphate, N1 -methyladenosine 5 ’-triphosphate, N1 -methylguanosine 5 ’-triphosphate, N6-methyladenosine 5 ’-triphosphate, 06- methylguanosine 5 ’-triphosphate, pseudouridine 5 ’-triphosphate, puromycin 5 ’-triphosphate or xanthosine 5 ’-triphosphate.
[0083] In some embodiments, the mRNAs include a 5’ cap structure. A 5’ cap is typically added as follows: first, an RNA terminal phosphatase removes one of the terminal phosphate groups from the 5’ nucleotide, leaving two terminal phosphates; guanosine triphosphate (GTP) is then added to the terminal phosphates via a guanylyl transferase, producing a 5’5’5 triphosphate linkage; and the 7-nitrogen of guanine is then methylated by a methyltransferase. Examples of cap structures include, but are not limited to, m7G(5’)ppp (5’(A,G(5’)ppp(5)A and G(5)ppp(5’)G.
[0084] In some embodiments, the retrotransposon of the present disclosure is delivered by a vector such as a plasmid vector and a viral vector (e.g., adenoviral vector).
[0085] In some embodiments, the target mammalian cell is a human cell. In some embodiments, the human cell is an induced pluripotent stem cell (iPSC). When an mRNA is directly introduced to the target stem cell, in a preferred embodiment, the downstream
flanking sequence of the retrotransposon has a length of at least 40 nt, such as 40 to 90 nt, 45 to 85 nt, or 50 to 80 nt.
[0086] In some embodiments, the human cell is not a stem cell. When an mRNA is directly introduced to the target non-stem cell, in a preferred embodiment, the downstream flanking sequence of the retrotransposon has a length of at least 70 nt, such as 70 to 95 nt, 70 to 90 nt, or 71 to 85 nt.
[0087] In some embodiments, the R2O1 ORF and the exogenous gene are included in the same retrotransposon polynucleotide. In some embodiments, only the exogenous gene is included in the retrotransposon polynucleotide, while the R2O1 ORF is delivered separately on a different polynucleotide.
[0088] In some embodiments, the delivery is in vitro. In some embodiments, the delivery is in vivo or ex vivo. In some embodiments, the retrotransposon polynucleotide is delivered by parenteral means, or locally.
[0089] The term “parenteral” as used herein refers to modes of administration which include intravenous, intramuscular, intraperitoneal, intrastemal, subcutaneous and intraarticular injection and infusion.
[0090] Delivery can be systemic or local. In addition, it may be desirable to introduce the retrotransposon polynucleotide of the disclosure into the central nervous system by any suitable route, including intraventricular and intrathecal injection; intraventricular injection may be facilitated by an intraventricular catheter, for example, attached to a reservoir, such as an Ommaya reservoir. Pulmonary administration can also be employed, e.g., by use of an inhaler or nebulizer, and formulation with an aerosolizing agent.
EXAMPLES
[0091] The following examples are included to demonstrate specific embodiments of the disclosure. It should be appreciated by those of skill in the art that the techniques disclosed in the examples which follow represent techniques to function well in the practice of the disclosure, and thus can be considered to constitute specific modes for its practice. However, those of skill in the art should, in light of the present disclosure, appreciate that many changes
can be made in the specific embodiments which are disclosed and still obtain a like or similar result without departing from the spirit and scope of the disclosure.
Example 1: Target-specific retrotransposition of R2O1 in human cells via viral vector delivery
[0092] This example developed a viral vector delivery system for R2O1 and tested targeted retrotransposition of the R2O1 elements.
[0093] Nine full-length sequences of R2 (R2O1) were isolated from the HdrR strain of Japanese medaka fish (Oryzias latipes). Clone F (Accession number: LC349444), which contained minimal amino acid changes in comparison with that of the consensus sequence, was selected for further analyses. Next, R2O1 mRNA was injected into zebrafish embryos, and it displayed the ability to retrotranspose into a specific target sequence in the 28s rDNA locus. Thus, a transgene system with R2O1 was established in zebrafish embryos, and it functioned successfully. In a pilot study, a recombinant virus (autographa californica nuclear polyhedrosis virus (AcNPV)) was used to deliver R2O1 into human cells, resulting in detection of retrotransposition activity. However, the AcNPV-based system had relatively low reproducibility and efficiency.
[0094] In the current study, a retrotransposition assay system was established using an adenovirus-based expression system, to determine whether R2O1 is able to retain its retrotransposon activity in human cells. This recombinant adenovirus pAxCAwtit-vectors contained the chicken beta-actin promoter and cytomegalovirus immediate-early enhancer (CAG) promoter.
Methods and Materials
Cell culture
[0095] The human cell line, 293 T (RCB2202), was purchased from the RIKEN cell bank in Japan. Cells were grown in Dulbecco’ s minimal essential medium (DMEM) supplemented with 10% fetal bovine serum (Gibco) and 1% penicillin/streptomycin (Invitrogen) in 5% CO2 at 37 °C. Cells were grown as adherent cell cultures and passaged every 72 h at a 1 :5 dilution (volume of cells: final volume of medium) until confluency was reached, in order to maintain log-phase growth.
Recombinant adenovirus construction
[0096] Adenovirus expression vectors were constructed with pAxCAwtit (TaKaRa) containing the chicken beta-actin promoter and CAG promoter. Furthermore, pENTR201r4, pr32ENTR201r4, or pENTR201-ENmutant were mixed with LR Clonase® (Gateway® Vector Conversion System, Invitrogen) to generate recombinant pAxCAwtit according to the manufacturer’s protocol. These recombinant pAxCAwtit vectors were digested with NspV, and the linearized adenovirus DNA was transfected into human 293 T cells using FuGENE® HD Transfection Reagent (Promega) according to the manufacturer’s protocol to generate the recombinant adenovirus. Viral titers were assessed using 50% Tissue Culture Infectious Dose (TCID50) assays before infecting the cells. For plasmid construction, PCR was conducted with the iProof™DNA polymerase (Bio-Rad) in total volumes of 20 pL. pBluescript-R2Ol/pBlue script II-R2OI RTmut
[0097] R2O1 WT (wild type) was constructed based on R2O1 (Accession number: LC349444). Briefly, full-length R2O1 from medaka fish genomic DNA was amplified using PCR with primers R2O1 5' end and R2O1 3' end. A 4.2 kb amplified fragment, which had less mutations, clone F, was named pGEM-R201-F. A full-length R2O1 fragment, digested with EcoRI and Spel from pGEM-R201-F, was inserted between the EcoRI and Spel sites and a SV40 poly(A) signal was inserted between Not! and Sad sites in a pBluescript II KS(+) vector to form pBluescript-R2Ol. An RTmutant was constructed by introducing a point mutation using inverse PCR from pR201r4, with primers R2O1-RT(D — V) s and R2O1- RT(D — V), and self-ligated to obtain pR201-RTmut, which was named pBluescript II-R2O1 RTmut. pCAGGS R2Ol/pCAGGS R20l-RTmut
[0098] To create pCAGGSR2Ol and pCAGGS R201-RTmut portions from a full-length R2O1 fragment, pGEM-R201 and pBluescript II-R201-RTmut, were amplified using PCR with the /'i RI-R2O15U and R2O13Ur4-EcoRI primers. The PCR reaction used consisted of an initial denaturation at 98 °C for 1 min, followed by 25 cycles at 98 °C for 10 s, 55 °C for 30 s, and 72 °C for 30 s, and a final extension at 72 °C for 5 min. The PCR products were purified using a GenElute PCR Clean-Up Kit (Sigma-Aldrich) and digested using EcoRI in M buffer (TaKaRa) at 37 °C for 1 h. In parallel, the empty vector pCAGGS (RIKKEN, catalog No. RDB0893) was digested using EcoRI in M buffer (TaKaRa) at 37 °C for 1 h. After electrophoresis in a 1% TAE (40 mM Tris, 20 mM CHiCOONa, ImM EDTA) agarose
gel, target bands were cut out from the gel and purified by a QIAquick Gel Extraction Kit (QIAGEN).
[0099] All purified DNA fragments and the linearized vector were incubated in Solution I (TaKaRa) at an insertion to vector ratio of 10: 1, for 1 h for ligation reaction at 16 °C. Then each ligated product was transformed into 100 pL E. coli competent DH5a cells and incubated on ice for 20 min, and subsequently heat shocked at 42 °C for 45 s and placed on ice for 2 min. After adding 200 pL LB medium and incubating in a water bath at 37 °C for 1 h, 150 pL of each transformate was spread on LB agar plates containing ampicillin and incubated at 37 °C for 16 h. Ampicillin-resistant transformants were picked and cultured overnight in LB medium with 50 mg/mL ampicillin. Plasmid DNA was isolated using a GenElute Plasmid DNA Miniprep Kit (Sigma-Aldrich). Plasmids were sequenced using a BigDye Terminator Cycle Sequencing Kit (Applied Biosystems) on a ABB 130 xl and a 3500/3500 xl Genetic Analyzer (Applied Biosystems). Sequence analysis was performed using a Vector NTI Advance 10 system (Invitrogen).
Cis retrotransposition assay
Adenovirus-based R2OI retrotransposition assay
[0100] Cells at 60% confluency were seeded into 24-well cluster plates one day before infection. Next, cells were infected with two types of adenoviruses, R201-r4, r32R201-r4 and R201-ENmut, using different MOI ratios, 1, 10 and 50 (MOI=1, 10, 50), respectively. Genomic DNA was extracted 72 h post-infection using Gentra Puregene® Kits (QIAGEN).
Plasmid-based R2OI retrotransposition assay
[0101] Transfection of plasmids into cells were performed with Lipofectamine® 3000 Reagent (Invitrogen). Here, 1 pg of pCAGGS R201-r4 or pCAGGS R201-RTmut, Lipofectamine® 3000 Reagent (2 pL each) and P3000 (DNA: P3000 = 1 pg: 2 pL) in OptiMEM medium (Invitrogen) were mixed and incubated for 15 min. Then this mixture was dispensed into each well of the microplate. The transfected cells were harvested after 72 h.
Retrotransposition detection
DNA extraction
[0102] Cells were detached from wells and transferred to a 1.5 mL microcentrifuge tube and centrifuged for 5 min at 3000 x g and washed twice with 1 x phosphate buffered saline (137 mM NaCl, 8.1 mM Na2HPO4, 8.1 mM KC1, 1.47 mM KH2PO4). Using Gentra
Puregene® Kits (QIAGEN), cells were resuspended in 150 pL lysis buffer and mixed by pipetting up and down or vortexing vigorously for 20 s at high speed. Next the preparation was treated with 50 pL protein precipitation solution, and vortexed vigorously for 20 s at high speed. After centrifugation at 12,000 x g for 1 min at room temperature, the upper aqueous phase was transferred to a new tube, treated with 150 pL isopropanol and gently inverted 50 times. Following centrifugation at 12,000 x g for 1 min at room temperature, the DNA was washed with 70% ethanol and air dried for 5 min. Finally, the pellet was dissolved in Tris- EDTA (TE) buffer and stored.
Nested PCR
[0103] Retrotransposition events were analyzed using both 3' and 5' junction PCR of the insertion. PCR reactions were performed with two types of polymerases and two sets of primers. The first polymerase was TaKaRa Ex Taq polymerase in a total volume of 20 pL, which was used according to the manufacturer’s protocol. The 3' junction of the R2O1 insertions was detected using nested PCR to identify retrotransposition events with R2O1 primer sets, h and Hs28S set i (FIG. 2). The second polymerase, KOD One ® PCR Master Mix (TOYOBO), was used for nested PCR in total volumes of 10 pL using the manufacturer’s protocol. The 3' junction of the R2O1 insertions here was detected using R2O1 primer sets, g and Hs28S set j (FIG. 2). The 5' junction of the R2O1 insertions was detected via nested PCR at five R2O1 positions. The forward primers were specific to an upstream sequence of the 28 S rDNA target site (Hs28S set f). The reverse primer sets were specific to five R2O1 positions al, a2, a3, a4, and a5 (FIG. 2).
Sequencing
[0104] For direct sequencing, the nested-PCR products were used as directed. For sequencing of clones, the Taq nested-PCR products were TA cloned into the pGEM-T Easy vector system (Promega) whereas the KOD nested-PCR products were TA cloned into the pTA2 vector from TArget Clone -Plus system (TOYOBO). After colony PCR, all PCR primers and unincorporated nucleotides of both PCR products were removed via EXO-SAP, and then the cleaned samples were sequenced using a BigDye Terminator Cycle Sequencing Kit (Applied Biosystems) on an ABB 130 xl and 3500/3500 xl Genetic Analyzer (Applied Biosystems). All sequence analyses were performed using a Geneious Prime system (Digital Biology).
Results
Adenovirus R20l-r4 retrotransposition assay in human cells
[0105] The 3' junction analyses confirmed the accurate retrotransposition of R201-r4 in human 293 T cells.
[0106] The wild type adenovirus construct, R201-r4, contained a CAG promoter for mammalian cells, a full-length R2O1 transcription unit (5' UTR/ORF/3' UTR) of R2O1, a 4 bp 28S rDNA target flanking sequence at its 3' end, and a polyadenylation signal sequence (PA; FIG. 3A). A 4 bp 28S rDNA flanking sequence (r4) added at the end of R2O1 3' UTR was contemplated to increase the annealing efficiency between the template RNA and target DNA. As a negative control, an R2O1 mutant construct, R201-ENMut, was used. R2O1- ENMut carried a mutation (IPA to IPD) at the essential amino acid sites for EN, completely abolishing any retrotransposition activities (FIG. 3A). Primer sets “h,” “i” and “g,” “j” were used to detect the 3' junction of retrotransposition (FIG. 3B).
[0107] Nested PCR using primer sets “h” and “i”, did not detect a PCR band in the lower multiplicity of infection (MOI) titer of 1, (FIG. 3C, MOI1; 1, 2 and 3 represent three independent experiments). However, at higher MOIs of 10 and 50, the expected PCR band (sized 327 bp), which represented the exact retrotransposition of R2O1 into the 28S rDNA target site, was detected (FIG. 3C, MOI10 and MOI50). Besides the band corresponding to the expected PCR product, a non-specific band was also observed (FIG. 3C; top panel). However, when primer sets “g” and “j” were used, the expected PCR band (sized 480 bp), which represents the exact retrotransposition event of R2O1, was detected at MOI 10 and 50 without any non-specific bands. Therefore, primer sets “g” and “j” were selected for further experiments. In addition, the beta-Actin gene was used as an internal control to check whether the genomes of human cells had been equally extracted.
[0108] In order to certify the accuracy of R2O1 retrotransposition in human cells, TA cloning of the PCR products of sample No.3 of MOHO (MOHO-3) and sample No. 3 of MOI50 (MOI50-3) was conducted using primer sets “h”, “i” and “g”, “j” (FIG. 3C; represented by solid triangle under the images). This example obtained 12 TA-clones of samples using primer sets “h” and “i”, of which nine clones were sequenced, and identified that 6 clones showed the exact 3' junction sequence, representing successful retrotransposition of R2O1. All eight TA-clones of PCR products produced using primer sets “g” and “j,” showed the successful retrotransposition of R2O1. All 14 clones included the
exact 3' junction sequence between R2O1 (the 3 '-terminal region of 3' UTR) and 28S rDNA. The remaining clones showed non-specific sequences from the adenovirus genome. By contrast, in R201-ENMut, which completely abolished target site cleavage, no PCR band was detected (FIG. 3D), indicating an absence of R2O1 retrotransposition activity. The band observed in MOI 50-2 samples using primer sets “h” and “i” corresponded to a non-specific sequence of 28 S rDNA, as confirmed by cloning and sequencing. These results demonstrated that wild-type R2O1 retrotransposed into the human 28 S rDNA, whereas the EN mutant did not. This indicated that the EN domain of R2O1, which is necessary for retrotransposing into the human 28S rDNA, is able to function in human cells as well. Furthermore, this indicated that R2O1 moves via target primed reverse transcription (TPRT), and not via recombination events.
1-2. 5' junction analyses showed that a full-length unit ofR2Ol-r4 inserts into the 28S rDNA [0109] The 5' junctions between the retrotransposed R2O1 and the 28S rDNA target were analyzed using five ORF reverse primers sets al, a2, a3, a4, a5 and a forward primer set, f (FIG. 4). Genomic DNA from 293T cells, which were not infected by the virus construct was used as a negative control. Using the same genomic DNA samples for 3' junction analyses (MOI10-1, MOI10-2, MOI10-3, MOI50-1, MOI50-2, and MOI50-3; FIG. 3C), multiple PCR bands were detected. The PCR products denoted by a solid triangle in the images were TA cloned. Out of 142 colony PCR products, 38 were sequenced. Among these, sequences from 14 clones contained both R2O1 and human 28 S rDNA. The remaining clones originated from the genomic sequences of adenovirus or 28 S rDNA.
[0110] The 5' junction regions around the insertion site of R2O1 for the above-mentioned 14 clones is summarized (FIG. 5). Most importantly, 2 clones showed that the full-length of R2O1 was inserted accurately into the target site, whereas the 5' side vector sequence was also reverse transcribed together with R2O1 (FIG. 5; insertion type a & b). Six clones were inserted into the 21 bp ~ 191 bp region downstream of the original target site (FIG. 5; insertion type d, e, i, k & 1), whereas 6 clones were inserted into the 5 bp ~ 22 bp region upstream of the target, within 28S rDNA (FIG. 5; insertion type c, f, g, h & j). Among the 14 clones, 12 represented 5' truncated R2Ols, each of which terminated its reverse transcription at various sites within the ORF (FIG. 5; insertion type c-1). The truncated regions ranged in length from 217 bp to 4023 bp from the 5'-end of R2O1. This type of 5' truncation is often observed among LINEs.
R201 was retrotransposed successfully with higher full-length integration of R201 in 293T cells infected with adenovirus carrying r32R20l-r4
[oni] To determine whether a longer upstream 28S rDNA sequence would increase the 5' integration of R2O1, 293T cells were infected with an adenovirus construct (r32R201-r4) containing a full-length unit of R201-r4 and 32 bp 28S rDNA 5'-flanking sequence (r32) added at the 5' end of R2O1 5' UTR, (FIG. 6A). The 3' junction analysis results showed that PCR bands of the expected sizes were detected at MOI 10 and 50 (FIG. 6B). Also, there were 2 bands, the sizes of which were longer than the expected size. Direct sequencing indicated that three of the five PCR products (FIG. 6B; MOI 10-2, MOI50-1, and MOI50-3) were exactly transposed into the target site, whereas two products (FIG. 6B; MOI10-3 and MOI50-2) contained some non-specific sequences or vector sequences with the R2O1 insertion. All 5 samples were successfully retrotransposed into the human 28 S target site.
[0112] The 5' junctions between the retrotransposed r32R201-r4 and the 28S rDNA target were also analyzed using the same primer sets used for the R201-r4 samples (FIG. 4) and multiple PCR bands were detected. The PCR products of some were TA cloned. The 5' junction region around the insertion site of r32R201-r4 for all 15 clones after cloning PCR and sequencing is summarized in FIG. 7. In each clone, the black box shows the 28S rDNA sequence and the gray box shows the R2O1 sequence, which was reverse transcribed.
[0113] Here, five clones showed that the full-length of R2O1 was exactly inserted into the target site (FIG. 7; insertion type a). Four clones were inserted into the 43 bp ~ 222 bp region downstream of the original target site (FIG. 7; insertion types d, f & h), respectively, whereas six clones were inserted into the 5 bp ~ 100 bp region upstream of the target, within 28 S rDNA (FIG. 7; insertion types a b, c, e, g & i). Among the 15 clones, 10 represented 5' truncated R2O1 (FIG. 7; insertion type b-i). The truncated regions ranged in length from 71 bp to 3739 bp from the 5'-end of R2O1. The remaining clones originated from the genomic sequences of adenovirus or 28S rDNA. A comparison between the 5' junction sequence analyses of R201-r4 and r32R201-r4 indicated that two out of 14 clones represented full- length R2O1 insertion in R201-r4 as opposed to five out of 15 clones in r32R201-r4. The results showed that flanking sequences at the 5' side of R2O1 may increase integration at the 5' junction.
Plasmid-based in vivo R201 retrotransposition assay in human cells
[0114] In addition to adenovirus-mediated infection described above, retrotransposition of R2O1 in human cells via plasmid transfection was also investigated. Previously, retrotransposition activity could not be detected with a plasmid carrying the CMV promoter (R201-pCEP4). To improve this method, this example changed the promoter and constructed a R2O1 plasmid with a CAG promoter (pCAGGS-R201; FIG. 8A) and detected successful integration of R2O1 after 3 days of plasmid transfection, via 3' junction analysis (FIG. 8B-C). By contrast, expected PCR bands were not detected in samples of RTmut, suggesting that R2O1 retrotransposed via plasmid delivery (FIG. 8B).
[0115] These results suggested that R2O1 retrotransposition using the adenovirus delivery system performed better than that with the plasmid transfection system, indicating the potential of adenovirus as a useful vector for the LINE transfer system in human cells as well.
The retrotransposition ability of adenovirus R2OI in human cells
[0116] In this example, an adenovirus-based in vivo retrotransposition assay indicated that R2O1 isolated from medaka fish, exhibited retrotransposition in human 293T cells. This assay further showed that both the 3 '-end and 5 '-end of R2O1 were inserted into the exact target site of 28S rDNA, showing that a full-length R2O1 unit had been reverse transcribed from 3 '-end of 3' UTR to the 5 '-end of 5' UTR. R2 elements are distributed in a wide variety of species ranging from arthropods to vertebrates (birds and fish), where these elements are found inserted into exactly the same target sequence in 28 S rDNA. Notably, the R2 element had become defunct from the genome of mammals, including that of Homo sapiens. However, this study showed that the R2O1 originating in medaka fish had also retained the ability to insert into the same 28S rDNA target sequence in human cells.
[0117] The example showed that the retrotransposition of R2O1 does not require HDR or donor DNA, but only the EN and RT domains. This indicates that the insertion of R2O1 into the target was moved by the TPRT mechanism, and not by recombination events, suggesting that target genes of interest could be inserted via TPRT using an mRNA template. Compared to HDR, which is a passive process in cells, reverse transcription of mRNA into cDNA via TPRT using the RT domain shows relatively higher processivity and fidelity, which is advantageous for synthesizing long cDNAs at the target site rather than recombination at the target site as donor DNA.
Variable 5' junctions ofR2Ol insertions
[0118] In this example, the R2 element retrotransposed into human 28S rDNA at the 3' side as well as the 5' side of the target. Notably, it is the first time that a full-length insertion (> 4.2 kb) of R2O1 sequence was detected, suggesting that longer genes of interest, which is at least approximately 4 kb, could be knocked-in by this system. In addition, the analysis of 5' junction sequences of retrotransposed R2Ols also showed 5' truncations at various sites within R2O1 and different insertion sites around the 28 S target site. However, the molecular mechanism of LINE retrotransposition remains unclear.
28S rDNA flanking sequences increase the accuracy ofR2Ol insertion integration
[0119] The annealing efficiency between template RNA and target DNA may increase R2O1 retrotransposition at the 3' side. In this example, R2O1 viruses with 4 bp 28S rDNA flanking sequences at the 3 '-end in contracts form of R201-r4 and r32R201r4, retrotransposed R2O1 into the target. The 3' junction analysis detected inaccurate cleavage of the bottom strand target in r32R201r4. However, the cleavage of the bottom strand target in R201-r4 appeared to be accurate. This observation indicates that the 32 bp 28S rDNA flanking sequences at the 5' end may affect the recognition or cleavage of the bottom strand target by R2O1.
Example 2. Target-specific retrotransposition of R2O1 in human cells with mRNA transfection
[0120] This example established an R2O1 retrotransposition system through mRNA- mediated delivery in human cells.
Materials and Methods
Cell culture
[0121] iPS cell: Cellartis Human induced pluripotent stem (iPS) cells (Y00275) were purchased from the TAKARA in Japan, were grown in the Cellartis DEF-CS™ 100 Culture System, which is a complete system for efficient expansion and scale-up manufacturing of human iPS cells in a feeder-free and defined environment, at 37 °C in 5% CO2.
[0122] Both cells are under adherent cultures and passaging is conducted in every 72 h at confluency condition with diluting at a 1 : 5 dilution (volume of cells: final volume of medium) to maintain the log phase growth.
In vitro transcription
[0123] Template DNA for in vitro transcription was amplified by PCR. R201-r4, R201-rlO and R2Ol-r50 were PCR amplified from prl05R2Olrl00, R201-RTmut-rlO and R2O1- RTmut-r50 were PCR amplified from pGEM-R201 RTmut, respectively. The forward primers used to transcribe these six types of mRNA, R201-r4, R201-rlO, R2Ol-r50, R2O1- RTmut-r4, R201-RTmut-rlO and R2Ol-RTmut-r50 were T7 5UTR. The reverse primer for mRNA R201-r4 and R201-RTmut-r4 were 28S 4a, for R201-rlO and R201-RTmut-rlO were 28S 10a, for R2Ol-r50 was 28S 50a (Table II-l). R2Ol-RTmut-r50 was PCR amplified 3 times using reverse primer 28S 20a, 28S 40a and 28S 50a. PCR products were purified using a PCR clean up kit (sigma) and used for 5’ capped mRNA synthesis in vitro transcription using mMESSAGE mMACHINE™ T7 Transcription Kit (Ambion). After mRNA synthesis, the template DNA was removed by DNase treatment and a >150 base poly(A) tail was added to RNA transcripts using The Poly(A) Tailing Kit (Ambion). Synthesized mRNA was precipitated with Lithium Chloride. mRNA based retrotransposition assay
[0124] Cells were seeded at 60% confluence in 24-well cluster plates one day before transfection. For transfection, OptiMEM medium (Invitrogen), was pre-warmed to room temperature.
[0125] Transfection of mRNA into cells was performed with Lipofectamine® MessengerMAX™ (Invitrogen). mRNA Lipofectamine® MessengerMAX™ Reagent (1.5 pL of each) were mixed with 25 pl OptiMEM medium and incubated for 10 min at temperature. 0.5 pg of mRNA R201-r4, R201-rlO, R2Ol-r50, R201-RTmut-r4, R201-RTmut-rlO and R2Ol-RTmut-r50, were mixed with OptiMEM medium in total volume 25 pL, respectably. Then, mixed 25 pL of lipofectamine reagent mixture with 25 pL mRNA mixture and incubated for 5 min at temperature. Mixtures were applied to each well. Helper virus infected the plasmid transfected cells at the same time and continue to incubate for 72 h.
Results mRNA mediated R2OI retrotransposition assay in 293T cells
[0126] Six types of R2O1 mRNA were used in this experiment. The linear template DNA was purified from PCR products of the plasmids which contain full-length of WT R2O1 or RTmut R2O1. The Capped mRNA was transcribed from the linear DNA template in vitro and transfected into 293 cells to verify the retrotransposition activity. Three types are WT R2O1
mRNA which had r4, rlO, and r71 28S flanking sequences at the end (FIG. 9A, Left). As a negative control, the other 3 of them are RTmut R2O1 mRNA, which lost its reverse transcription ability, also have r4, rlO, or r50 28 S flanking sequences at the end (FIG. 9A, Right). After performing nested PCR of the cell genome and amplifying the 3’ junction, the expected size of the band was observed at sample R201-r71, but not at R201-r4 or R201-rlO (FIG. 9B, Left panel). In contrast, samples transfected with RTmut mRNA observed no band (FIG. 9B, Right panel), suggesting that R2O1 moves by TPRT and not by recombination events. PCR products were TA cloned and sequenced. All 8 clones had the same sequences of insertion of R2O1, revealing that the medaka R2O1 was inserted into the exact target sequence of the human 28S rDNA. The above results confirmed that the medaka fish R2O1 retrotransposed into the 28 S rDNA target sequence in 293 T cell host through mRNA- mediated delivery. mRNA mediated R2OI retrotransposition assay in iPS cells.
[0127] Next, the mRNA-based retrotransposition system was tested in iPS cells. Also, 6 types R2O1 mRNA were used in this experiment. Capped mRNA was transcribed from full- length R2O1 construct or RTmut construct in vitro and transfected into iPS cells to verify the retrotransposition activity (FIG. 10A). Three types of mRNA are WT R2O1 mRNA with r4, rlO, and r50 28S flanking sequences at the end (FIG. 10A, Left). 3 types of RTmut R2O1 mRNA, which have r4, rlO, and r50 28S flanking sequences at the end were used as a negative control (FIG. 10A, Right). Every type of mRNA has been experimented with 3 replicates (numbers 1-3 in figures). After performing nested PCR of the cell genome and amplifying the 3’ junction, the expected size of the band was observed only in samples of R2Ol-r50 (FIG. 10B, top panel). In contrast, samples transfected with RTmut mRNA observed no band (FIG. 10B, down panel). PCR products were TA cloned and sequenced. All 8 clones from the PCR products of R2Ol-r50 had the exact sequences of insertion of R2O1 and 28S rDNA sequences. The above results confirmed that R2O1 retrotransposed into the 28S rDNA target sequence in iPS cells host though mRNA-mediated delivery.
[0128] Next, to test when R2O1 retrotransposition occurs after transfection, this example transfected those 3 types of WT R2O1 mRNA into iPS cells and extracted their genome after 6 h, 12 h, 24 h post-transfection respectively. After nested-PCR of 3 ’junction, the band of the expected size was only detected at samples that transfected with R2Ol-r50 mRNA after 24h,
which suggests that R201 retrotransposition occurred at least after 24h post mRNA transfection.
[0129] In this example, R2O1 showed the retrotransposition activity in human 293T cells and iPS cells via an mRNA-based in vivo retrotransposition assay, using mRNA-mediated delivery. The summary of sequences of the PCR products showed capped R2O1 mRNA with long 28S rDNA flanking sequences at the 3'-end inserted into the exact target site of 28S rDNA in 293 T cells and iPS cells .
[0130] It has been reported that the HDR pathway in iPS cells is low, especially for longer genes, which limited the knock-in efficiency of the CRISPR/Cas system. While some recent studies showed the improvement of the knock-in efficiency in ES cells, the DNA-based system remains potential risks, such as tumorigenicity and genome instability. Thus, mRNA- mediated R2O1 retrotransposition may provide a new genome knock-in method in iPS cells, which is important for facilitating the development of gene therapy.
Longer 28S rDNA flanking sequences at capped R2OI mRNA 3 ’-end increase its retrotransposition activity.
[0131] In Example 1, R2O1 constructs with 4 bp 28 S rDNA flanking sequence worked efficiently. This example further demonstrated that the retrotransposition activity was significantly improved by using capped R2O1 mRNA, which has 50 bp 28S rDNA flanking sequences at the 3-end rather than 4 bp or 10 bp. It is contemplated that instead of poly(A) tail, longer 28S rDNA flanking sequences at the 3 ’end of R2O1 mRNA had the added benefit of protecting the mRNA from degradation, which is important for its retrotransposition activity in the human cell.
Example 3. Roles of Structural Elements of the R2O1
[0132] This example examined the impacts of the 5’ and 3’ sequences of the R2O1 retrotransposon, which can be useful for optimizing its applications.
[0133] The retrotransposition assays were conducted in zebrafish embryos. The following mRNA constructs were prepared (see illustrations in FIG. 11), (A) the wild-type (WT) R2O1 retrotransposon with intact 5’UTR and 3’UTR, (B) the R2O1 retrotransposon with a poly(A) signal added to the 3’ end, (C) the R2O1 retrotransposon with four 5’ nucleotides of the 28 S (after insertion the site) (r4) added to the 3’ end, (D) the R2O1 retrotransposon with r4,
following by a poly(A) signal, added to the 3’ end, (E) the R2O1 retrotransposon with 10 5’ nucleotides of the 28 S (after insertion the site) (rlO) added to the 3’ end, followed by a poly(A) signal, (F) the R2O1 retrotransposon with 100 5’ nucleotides of the 28 S (after insertion the site) (rlOO) added to the 3’ end, followed by a poly(A) signal, (F) the R2O1 retrotransposon with r4, but with the 5’UTR deleted, (G) the R2O1 retrotransposon with poly(A), but with the 5’UTR deleted,, and (H) the R2O1 retrotransposon with r4, but with the 5’UTR deleted and without the 5’ mRNA Cap.
[0134] The detected retrotransposition efficiencies are shown on the right portion of FIG. 11. None of the Cap, 5’UTR or the poly(A) appeared to be necessary for the retrotransposition. The 3’ addition of the 28S sequences (r4, rlO, and rlOO), by contrast, is important.
[0135] It was then tested whether the selection of the 28S nucleotides is important. For the r4, 4 consecutive nucleotides from the 28 S sequence, but starting at different locations (1 through 8) downstream of the insertion site, were tested (FIG. 12). As shown in the figure, when the r4 is 5 or more nucleotides away from the target insertion site, retrotransposition did not happen efficiently.
[0136] Further detailed examination of the 5’UTR sequence was conducted, with constructs missing certain portion of the 5’UTR. As shown in FIG. 13A, deletion of the first 175 nucleotides appeared to improve retrotransposition efficiency while deletions of 215 or more 5’ nucleotides appeared to reduce the retrotransposition efficiency. By contrast, even a deletion of 36 nucleotides in the 3’UTR appeared to reduce the retrotransposition efficiency (FIG. 13B)
Example 4. R2O1 for Gene Delivery
[0137] This example tested the capability of R2O1 to carry an exogenous coding sequence to be integrated to the 28S genes in zebrafish.
[0138] In the constructs illustrated in FIG. 14A-B, an exogenous EGFP or Gal4 gene was inserted (in opposite orientation) downstream of the R2O1 ORF within the R2O1 retrotransposon. As shown in the Western blots and microscopic images, the EGFP or Gal4 gene was expressed in the zebrafish. Such results demonstrated the capability of R2O1 as a suitable carrier to knock-in exogenous genes into a target genome.
Example 5. R2O1 ORF Expressed Separately
[0139] This example tested whether an R2O1 ORF can be delivered and expressed separately to enable retrotransposition of an R2O1 construct.
[0140] The proposed procedure is illustrated in FIG. 15 A, where an R2O1 construct (an exogenous gene and the R2O1 3’UTR) and an R2O1 ORF-expressing construct were separately introduced to a target cell. The exogenous gene tested was EGFP, regulated by a SV40 promoter (FIG. 15B). As shown in FIG. 15C, with the wild-type R2O1 ORF (not the inactive mutant) separately expressed in the target cell, the R2O1-EGFP construct was successfully integrated to the target genome and was expressed.
* * *
[0141] Unless otherwise defined, all technical and scientific terms used herein have the same meaning as commonly understood by one of ordinary skill in the art to which this invention belongs.
[0142] The inventions illustratively described herein may suitably be practiced in the absence of any element or elements, limitation or limitations, not specifically disclosed herein. Thus, for example, the terms “comprising”, “including,” “containing”, etc. shall be read expansively and without limitation. Additionally, the terms and expressions employed herein have been used as terms of description and not of limitation, and there is no intention in the use of such terms and expressions of excluding any equivalents of the features shown and described or portions thereof, but it is recognized that various modifications are possible within the scope of the invention claimed.
[0143] Thus, it should be understood that although the present invention has been specifically disclosed by preferred embodiments and optional features, modification, improvement and variation of the inventions embodied therein herein disclosed may be resorted to by those skilled in the art, and that such modifications, improvements and variations are considered to be within the scope of this invention. The materials, methods, and examples provided here are representative of preferred embodiments, are exemplary, and are not intended as limitations on the scope of the invention.
[0144] The invention has been described broadly and generically herein. Each of the narrower species and subgeneric groupings falling within the generic disclosure also form
part of the invention. This includes the generic description of the invention with a proviso or negative limitation removing any subject matter from the genus, regardless of whether or not the excised material is specifically recited herein.
[0145] In addition, where features or aspects of the invention are described in terms of Markush groups, those skilled in the art will recognize that the invention is also thereby described in terms of any individual member or subgroup of members of the Markush group.
[0146] All publications, patent applications, patents, and other references mentioned herein are expressly incorporated by reference in their entirety, to the same extent as if each were incorporated by reference individually. In case of conflict, the present specification, including definitions, will control.
[0147] It is to be understood that while the disclosure has been described in conjunction with the above embodiments, that the foregoing description and examples are intended to illustrate and not limit the scope of the disclosure. Other aspects, advantages and modifications within the scope of the disclosure will be apparent to those skilled in the art to which the disclosure pertains.
Claims
1. A polynucleotide that is a messenger RNA (mRNA) or a DNA encoding the mRNA, wherein the mRNA comprises, from 5’ to 3’: a protein-coding sequence or a reverse-complement thereof; a 3’ untranslated region (UTR) having at least 85% sequence identity to SEQ ID NO: 16; and a downstream flanking sequence consisting of a fragment of SEQ ID NO:20 or an RNA sequence having least 85% sequence identity to SEQ ID NO:20, wherein the fragment includes at least 4 consecutive nucleotides of SEQ ID NO:20 starting at a position from 1 to 5.
2. The polynucleotide of claim 1, wherein the mRNA does not include a poly(A) signal.
3. The polynucleotide of claim 1 or 2, wherein the 3 ’UTR is directly connected to the downstream flanking sequence.
4. The polynucleotide of any preceding claim, wherein the downstream flanking sequence has a length of 4 to 180 nucleotides (nt).
5. The polynucleotide of claim 4, wherein the downstream flanking sequence has a length of 10 to 150 nt.
6. The polynucleotide of claim 4, wherein the downstream flanking sequence has a length of 15 to 150 nt.
7. The polynucleotide of claim 4, wherein the downstream flanking sequence has a length of 40 to 90 nt.
8. The polynucleotide of claim 4, wherein the downstream flanking sequence has a length of 50 to 90 nt.
9. The polynucleotide of claim 4, wherein the downstream flanking sequence has a length of 71 to 90 nt.
10. The polynucleotide of any preceding claim, wherein the fragment includes at least the four 5’ nucleotides (UAGC) of SEQ ID NO:20.
11. The polynucleotide of any preceding claim, wherein the 3 ’UTR has at least 90% sequence identity to SEQ ID NO: 16.
12. The polynucleotide of any preceding claim, wherein the 3 ’UTR has at least 95% sequence identity to SEQ ID NO: 16.
13. The polynucleotide of any preceding claim, wherein the 3 ’UTR consists of SEQ ID NO: 16.
14. The polynucleotide of any preceding claim, wherein the mRNA further comprises, 5’ to the protein-coding sequence, an upstream flanking sequence consisting of a fragment of SEQ ID NO: 18 or an RNA sequence having at least 85% sequence identity to SEQ ID NO: 18.
15. The polynucleotide of claim 14, wherein the fragment of the upstream flanking sequence ends at a position from 176 to 180 of SEQ ID NO: 18.
16. The polynucleotide of claim 14 or 15, wherein the upstream flanking sequence has a length of 4 to 180 nt.
17. The polynucleotide of claim 14 or 15, wherein the upstream flanking sequence has a length of 10 to 150 nt.
18. The polynucleotide of claim 14 or 15, wherein the upstream flanking sequence has a length of 32 to 150 nt.
19. The polynucleotide of claim 14 or 15, wherein the upstream flanking sequence has a length of 20 to 100 nt.
20. The polynucleotide of claim 14 or 15, wherein the upstream flanking sequence has a length of 32 to 80 nt.
21. The polynucleotide of any preceding claim, wherein the mRNA further comprises, 5’ to the protein-coding sequence, a 5’ untranslated region (UTR) comprising nucleotides 233 to 265 of SEQ ID NO: 14 or a nucleic acid sequence having at least 85% sequence identity to nucleotides 233 to 265 of SEQ ID NO: 14.
22. The polynucleotide of claim 21, wherein the mRNA does not include nucleotides 1 to 87 of SEQ ID NO: 14.
23. The polynucleotide of claim 21, wherein the mRNA does not include nucleotides 1 to 175 of SEQ ID NO: 14.
24. The polynucleotide of claim 21, wherein the mRNA does not include nucleotides 1 to 198 of SEQ ID NO: 14.
25. The polynucleotide of any one of claims 21 to 24, wherein the 5 ’UTR comprises nucleotides 216 to 265 of SEQ ID NO: 14 or a nucleic acid sequence having at least 85% sequence identity to nucleotides 216 to 265 of SEQ ID NO: 14.
26. The polynucleotide of any one of claims 14 to 25, wherein the upstream flanking sequence is located 5’ to the 5 ’UTR.
27. The polynucleotide of any preceding claim, wherein the protein-coding sequence encodes a Long Interspersed Nuclear Element (LINE) R2 of medaka fish Oryzias latipes (R2O1) open reading frame (ORF), which can be expressed by the mRNA.
28. The polynucleotide of claim 27, wherein the ORF comprises the amino acid sequence of SEQ ID NO: 15 or an amino acid sequence having at last 85% sequence identity to SEQ ID NO: 15.
29. The polynucleotide of any one of claims 1-26, wherein the protein-coding sequence encodes an exogenous therapeutic protein for delivery to a target cell.
30. The polynucleotide of claim 27 or 28, wherein the mRNA further comprises a second protein-encoding sequence or the reverse-complement thereof, wherein the second proteinencoding sequence encodes an exogenous therapeutic protein for delivery to a target cell.
31. A method for integrating an exogenous polynucleotide to the genome of a mammalian cell, comprising introducing to the cell the polynucleotide of any one of claims 1 to 30.
32. The method of claim 31, wherein the polynucleotide is a mRNA, a plasmid or an adenoviral vector.
33. The method of claim 31, wherein the mammalian cell is a human cell.
34. The method of claim 33, wherein the human cell is an induced pluripotent stem cell (iPSC).
35. The method of claim 34, wherein the polynucleotide is mRNA and the downstream flanking sequence has a length of 40 to 90 nt.
35. The method of claim 34, wherein the polynucleotide is a plasmid or an adenoviral vector and the downstream flanking sequence has a length 4 to 90 nt.
36. The method of claim 33, wherein the human cell is not a stem cell.
37. The method of claim 34, wherein the downstream flanking sequence has a length of 70 to 90 nt.
38. The method of any one of claims 31 to 37, wherein the polynucleotide or the reversecomplement thereof encodes a protein that is not an R2O1 ORF, and the method further comprises introducing to the mammalian cell a second polynucleotide encoding the R2O1 ORF.
Applications Claiming Priority (6)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| US202363499626P | 2023-05-02 | 2023-05-02 | |
| US202363499630P | 2023-05-02 | 2023-05-02 | |
| US202363499631P | 2023-05-02 | 2023-05-02 | |
| US63/499,626 | 2023-05-02 | ||
| US63/499,630 | 2023-05-02 | ||
| US63/499,631 | 2023-05-02 |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| WO2024228121A1 true WO2024228121A1 (en) | 2024-11-07 |
Family
ID=91070249
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/IB2024/054205 Pending WO2024228121A1 (en) | 2023-05-02 | 2024-05-01 | Long interspersed nuclear element r2 for target-specific gene delivery |
Country Status (1)
| Country | Link |
|---|---|
| WO (1) | WO2024228121A1 (en) |
Citations (13)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US4373071A (en) | 1981-04-30 | 1983-02-08 | City Of Hope Research Institute | Solid-phase synthesis of polynucleotides |
| US4401796A (en) | 1981-04-30 | 1983-08-30 | City Of Hope Research Institute | Solid-phase synthesis of polynucleotides |
| US4415732A (en) | 1981-03-27 | 1983-11-15 | University Patents, Inc. | Phosphoramidite compounds and processes |
| US4458066A (en) | 1980-02-29 | 1984-07-03 | University Patents, Inc. | Process for preparing polynucleotides |
| US4500707A (en) | 1980-02-29 | 1985-02-19 | University Patents, Inc. | Nucleosides useful in the preparation of polynucleotides |
| US4668777A (en) | 1981-03-27 | 1987-05-26 | University Patents, Inc. | Phosphoramidite nucleoside compounds |
| US4973679A (en) | 1981-03-27 | 1990-11-27 | University Patents, Inc. | Process for oligonucleo tide synthesis using phosphormidite intermediates |
| US5047524A (en) | 1988-12-21 | 1991-09-10 | Applied Biosystems, Inc. | Automated system for polynucleotide synthesis and purification |
| US5132418A (en) | 1980-02-29 | 1992-07-21 | University Patents, Inc. | Process for preparing polynucleotides |
| US5153319A (en) | 1986-03-31 | 1992-10-06 | University Patents, Inc. | Process for preparing polynucleotides |
| US5262530A (en) | 1988-12-21 | 1993-11-16 | Applied Biosystems, Inc. | Automated system for polynucleotide synthesis and purification |
| US5700642A (en) | 1995-05-22 | 1997-12-23 | Sri International | Oligonucleotide sizing using immobilized cleavable primers |
| WO2022155055A1 (en) * | 2021-01-14 | 2022-07-21 | The Regents Of The University Of California | Site-specific gene modifications |
-
2024
- 2024-05-01 WO PCT/IB2024/054205 patent/WO2024228121A1/en active Pending
Patent Citations (13)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US4458066A (en) | 1980-02-29 | 1984-07-03 | University Patents, Inc. | Process for preparing polynucleotides |
| US5132418A (en) | 1980-02-29 | 1992-07-21 | University Patents, Inc. | Process for preparing polynucleotides |
| US4500707A (en) | 1980-02-29 | 1985-02-19 | University Patents, Inc. | Nucleosides useful in the preparation of polynucleotides |
| US4973679A (en) | 1981-03-27 | 1990-11-27 | University Patents, Inc. | Process for oligonucleo tide synthesis using phosphormidite intermediates |
| US4415732A (en) | 1981-03-27 | 1983-11-15 | University Patents, Inc. | Phosphoramidite compounds and processes |
| US4668777A (en) | 1981-03-27 | 1987-05-26 | University Patents, Inc. | Phosphoramidite nucleoside compounds |
| US4373071A (en) | 1981-04-30 | 1983-02-08 | City Of Hope Research Institute | Solid-phase synthesis of polynucleotides |
| US4401796A (en) | 1981-04-30 | 1983-08-30 | City Of Hope Research Institute | Solid-phase synthesis of polynucleotides |
| US5153319A (en) | 1986-03-31 | 1992-10-06 | University Patents, Inc. | Process for preparing polynucleotides |
| US5047524A (en) | 1988-12-21 | 1991-09-10 | Applied Biosystems, Inc. | Automated system for polynucleotide synthesis and purification |
| US5262530A (en) | 1988-12-21 | 1993-11-16 | Applied Biosystems, Inc. | Automated system for polynucleotide synthesis and purification |
| US5700642A (en) | 1995-05-22 | 1997-12-23 | Sri International | Oligonucleotide sizing using immobilized cleavable primers |
| WO2022155055A1 (en) * | 2021-01-14 | 2022-07-21 | The Regents Of The University Of California | Site-specific gene modifications |
Non-Patent Citations (1)
| Title |
|---|
| KUROKI-KAMI AZUSA ET AL: "Targeted gene knockin in zebrafish using the 28S rDNA-specific non-LTR-retrotransposon R2Ol", MOBILE DNA, vol. 10, no. 1, 22 May 2019 (2019-05-22), XP093018836, Retrieved from the Internet <URL:http://link.springer.com/article/10.1186/s13100-019-0167-2/fulltext.html> DOI: 10.1186/s13100-019-0167-2 * |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| US10927383B2 (en) | Cas9 mRNAs | |
| ES2958832T3 (en) | Plasmid containing a sequence coding for an mRNA with a cleaved poly(A) tail | |
| JP2025066771A (en) | RNA targeting of mutations by suppressor tRNAs and deaminases | |
| CA3149897A1 (en) | Methods and compositions for genomic integration | |
| ES2950736T3 (en) | Viral and non-viral nanoplasmid vectors with improved production | |
| US11884915B2 (en) | Guide RNAs with chemical modification for prime editing | |
| US20230332184A1 (en) | Template guide rna molecules | |
| WO2020198403A2 (en) | Compositions comprising modified circular polyribonucleotides and uses thereof | |
| US20190374567A1 (en) | Reverse Transcriptase Dependent Conversion of RNA Templates Into DNA | |
| JP2004524031A (en) | Bacterial plasmid lacking synthetic gene and CpG | |
| AU2022272235A1 (en) | Methods and compositions for genomic integration | |
| WO2022256578A2 (en) | Circular guide rnas for crispr/cas editing systems | |
| EP4172329A2 (en) | Compositions for genome editing and methods of use thereof | |
| WO2024228121A1 (en) | Long interspersed nuclear element r2 for target-specific gene delivery | |
| CN110551762B (en) | CRISPR/ShaCas9 gene editing system and application thereof | |
| CN110551763B (en) | CRISPR/SlutCas9 gene editing system and application thereof | |
| KR20220018410A (en) | Self-transcribing RNA/DNA system that provides Genome editing in the cytoplasm | |
| KR20230028860A (en) | Self-transcribing RNA/DNA system that provides cDdRp mRNAs in the cytoplasm | |
| KR20220055399A (en) | Self-transcribing RNA/DNA system that provides mRNAs in the cytoplasm | |
| WO2025217194A1 (en) | Repeat dosing of retrotransposons | |
| WO2025085519A1 (en) | R2 retrotransposons for gene writing | |
| Ma et al. | Modulating the cleavage and polyadenylation site: from research tools to therapeutic opportunities | |
| KR20230127069A (en) | Self-transcribing RNA/DNA system that provides mRNAs in the cytoplasm | |
| WO2024173699A2 (en) | Compositions for the treatment of spinal muscular atrophy | |
| WO2024091907A1 (en) | Compositions and methods for modifying the hpv16 genome |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| 121 | Ep: the epo has been informed by wipo that ep was designated in this application |
Ref document number: 24725264 Country of ref document: EP Kind code of ref document: A1 |
|
| NENP | Non-entry into the national phase |
Ref country code: DE |