WO2024223728A1 - Influenza virus vaccines - Google Patents
Influenza virus vaccines Download PDFInfo
- Publication number
- WO2024223728A1 WO2024223728A1 PCT/EP2024/061356 EP2024061356W WO2024223728A1 WO 2024223728 A1 WO2024223728 A1 WO 2024223728A1 EP 2024061356 W EP2024061356 W EP 2024061356W WO 2024223728 A1 WO2024223728 A1 WO 2024223728A1
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- influenza
- virus
- suitably
- strain
- subtype
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Pending
Links
Classifications
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K39/00—Medicinal preparations containing antigens or antibodies
- A61K39/12—Viral antigens
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P31/00—Antiinfectives, i.e. antibiotics, antiseptics, chemotherapeutics
- A61P31/12—Antivirals
- A61P31/14—Antivirals for RNA viruses
- A61P31/16—Antivirals for RNA viruses for influenza or rhinoviruses
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K39/00—Medicinal preparations containing antigens or antibodies
- A61K2039/51—Medicinal preparations containing antigens or antibodies comprising whole cells, viruses or DNA/RNA
- A61K2039/53—DNA (RNA) vaccination
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K39/00—Medicinal preparations containing antigens or antibodies
- A61K39/12—Viral antigens
- A61K39/145—Orthomyxoviridae, e.g. influenza virus
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N2760/00—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA ssRNA viruses negative-sense
- C12N2760/00011—Details
- C12N2760/16011—Orthomyxoviridae
- C12N2760/16111—Influenzavirus A, i.e. influenza A virus
- C12N2760/16134—Use of virus or viral component as vaccine, e.g. live-attenuated or inactivated virus, VLP, viral protein
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N2760/00—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA ssRNA viruses negative-sense
- C12N2760/00011—Details
- C12N2760/16011—Orthomyxoviridae
- C12N2760/16211—Influenzavirus B, i.e. influenza B virus
- C12N2760/16234—Use of virus or viral component as vaccine, e.g. live-attenuated or inactivated virus, VLP, viral protein
Definitions
- the present invention is inter alia directed to immunogenic compositions, as well as vaccines and kits or kits of parts comprising such, for use in the treatment or prophylaxis of an infection with an Influenza virus comprising nucleic acids, suitably mRNAs, encoding hemagglutinin (HA) antigens derived from strains of Influenza A and B virus.
- the present invention is also directed to a method of eliciting an immune response against an Influenza virus and to a method of treating or preventing a disorder caused by an Influenza virus.
- Influenza viruses are RNA viruses belonging to the family Orthomyxoviridae (NCBI Taxonomy ID: 11308), being sub-divided into e.g. AlphaInfluenzavirus (the genus that includes Influenza A viruses) and BetaInfluenzavirus (the genus that includes Influenza B viruses), that circulate in all parts of the world. Influenza viruses cause acute respiratory illness often during local outbreaks or seasonal epidemics and occasionally during pandemics. Typical Influenza epidemics cause increases in incidence of pneumonia and lower respiratory disease by increased rates of hospitalization or mortality. The elderly or those with underlying chronic diseases are most likely to experience such complications, but young infants also may suffer severe disease. Influenza viruses (mainly Influenza A and B viruses) have a significant impact on global public health, causing millions of cases of severe illness each year, thousands of deaths, and considerable economic losses.
- Influenza viruses such as Influenza A and B viruses
- the best-characterized of these viral proteins are hemagglutinin (HA) and neuraminidase (NA), two large glycoproteins found on the outside of the viral particles.
- HA hemagglutinin
- NA neuraminidase
- NA is an enzyme involved in the release of progeny virus from infected cells.
- HA is a lectin that mediates binding of the virus to target cells and entry of the viral genome into the target cell.
- HA H1-H18 subtypes
- NA N1- N11 subtypes of Influenza A viruses that potentially form 144 HA and NA combinations.
- the Influenza B viruses almost exclusively infect humans.
- the Influenza B viruses are categorized into two distinct lineages: BA/ictoria/2/1987-like (B/Victoria lineage) and B/Yamagata/16/1988-like (B/Yamagata lineage) viruses that have been circulating worldwide since 1983.
- Influenza virus B mutates at a rate 2 to 3 times slower than type A; however, it significantly impacts children and young adults annually.
- Vaccination is currently the most widely used method to prevent Influenza outbreaks, particularly in high-risk population.
- Multivalent live attenuated (FLUMIST, AstraZeneca), inactivated (AFLURIA, FLUAD and FLUCELVAX, Seqirus; FLUARIX and FLULAVAL, GlaxoSmithKline; FLUZONE, Sanofi), or recombinant (FLUBLOK, Sanofi) flu vaccines are already available on the market for active immunization against disease caused by Influenza subtype A viruses and Influenza type B viruses contained in the vaccines.
- HA is the major Influenza virus antigen recognized by neutralizing antibodies
- this glycoprotein has been the focus of currently inactivated and recombinant approved flu vaccines.
- flu vaccines are quadrivalent vaccines, based on 4 HA derived from each of the four strains of Influenza virus specified by health authorities for inclusion in the annual seasonal vaccine (typically two Influenza subtype A strains and two Influenza type B strains), meaning designed to protect against those four different flu virus strains.
- the invention provides an immunogenic composition for use in the treatment or prophylaxis of an infection with an Influenza virus, wherein the immunogenic composition comprises:
- a fourth nucleic acid encoding a HA antigen of a second strain of Influenza B virus, wherein an immune response is elicited against HA antigens of said strains of first and second subtypes of Influenza A virus, said first and, optionally, second strains of Influenza B virus and at least one further HA antigen subtype of Influenza A virus, being different from any of the HA antigen subtypes of Influenza A virus encoded by a nucleic acid present in the composition.
- the invention provides a vaccine for use in the treatment or prophylaxis of an infection with an Influenza virus, comprising an immunogenic composition as defined herein, wherein an immune response is elicited as defined herein.
- the invention provides a kit or kit of parts for use in the treatment or prophylaxis of an infection with an Influenza virus, wherein the kit or kit of parts comprises the nucleic acids, suitably the mRNAs, as defined herein, suitably the mRNAs of (a), (b), (c), (d), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ) as defined herein, optionally comprising a liquid vehicle for solubilizing, and, optionally, technical instructions providing information on administration and dosage of the components, wherein an immune response is elicited as defined herein.
- the kit or kit of parts comprises the nucleic acids, suitably the mRNAs, as defined herein, suitably the mRNAs of (a), (b), (c), (d), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ) as defined herein, optionally comprising a liquid vehicle for solubilizing, and, optionally, technical instructions providing information on administration
- the invention provides a method of eliciting an immune response against an Influenza virus, wherein the method comprises applying or administering to a subject in need thereof an immunogenic composition as defined herein, and wherein an immune response is elicited as defined herein.
- the invention provides a method of treating or preventing a disorder caused by an Influenza virus, wherein the method comprises applying or administering to a subject in need thereof an immunogenic composition as defined herein, and wherein an immune response is elicited as defined herein.
- SEQ ID NO: 1 Amino acid sequence of HA from A/Michigan/45/2015 (H1 N1).
- SEQ ID NO: 2 Amino acid sequence of NA from A/Michigan/45/2015 (H1 N1).
- SEQ ID NO: 3 Amino acid sequence of HA from A/Switzerland/8060/2017 (H3N2).
- SEQ ID NO: 4 Amino acid sequence of NA from A/Switzerland/8060/2017 (H3N2).
- SEQ ID NO: 5 Amino acid sequence of HA from B/Colorado/06/2017.
- SEQ ID NO: 6 Amino acid sequence of NA from B/Colorado/06/2017.
- SEQ ID NO: 7 Amino acid sequence of HA from B/Phuket/3073/2013.
- SEQ ID NO: 8 Amino acid sequence of NA from B/Phuket/3073/2013.
- SEQ ID NO: 10 Amino acid sequence of NA from A/Singapore/INFIMH-16-0019/2016
- SEQ ID NO: 11 Amino acid sequence of HA from A/Brisbane/02/2018 (H1 N1).
- SEQ ID NO: 12 Amino acid sequence of NA from A/Brisbane/02/2018 (H1 N1).
- SEQ ID NO Amino acid sequence of HA from A/Kansas/14/2017 (H3N2).
- SEQ ID NO Amino acid sequence of NA from A/Kansas/14/2017 (H3N2).
- SEQ ID NO: 15 Amino acid sequence of HA from A/South Austral ia/34/2019 (H3N2).
- SEQ ID NO: 16 Amino acid sequence of NA from A/South Austral ia/34/2019 (H3N2).
- SEQ ID NO: 17 Amino acid sequence of HA from B/Washington/02/2019.
- SEQ ID NO: 18 Amino acid sequence of NA from B/Washington/02/2019.
- SEQ ID NO: 19 Amino acid sequence of HA from A/Guangdong-Maonan /SWL1536/2019 (H1 N1).
- SEQ ID NO: 20 Amino acid sequence of NA from A/Guangdong- Maonan /SWL1536/2019 (H1 N1).
- SEQ ID NO: 21 Amino acid sequence of HA from A/Hong Kong/2671/2019 (H3N2).
- SEQ ID NO: 22 Amino acid sequence of NA from A/Hong Kong/2671/2019 (H3N2).
- SEQ ID NO: 23 Amino acid sequence of HA from A/Hawaii/70/2019 (H1 N1).
- SEQ ID NO: 24 Amino acid sequence of NA from A/Hawaii/70/2019 (H1 N1).
- SEQ ID NO: 25 Amino acid sequence of HA from A/Hong Kong/45/2019 (H3N2).
- SEQ ID NO: 26 Amino acid sequence of NA from A/Hong Kong/45/2019 (H3N2).
- SEQ ID NO: 27 Amino acid sequence of HA from A/Victoria/2570/2019 (H1 N1).
- SEQ ID NO: 28 Amino acid sequence of NA from A/Victoria/2570/2019 (H1 N1).
- SEQ ID NO: 29 Amino acid sequence of HA from A/Wisconsin/588/2019 (H1 N1).
- SEQ ID NO: 30 Amino acid sequence of NA from A/Wisconsin/588/2019 (H1 N1).
- SEQ ID NO: 31 Amino acid sequence of HA from A/Cambodia/e0826360/2020 (H3N2).
- SEQ ID NO: 32 Amino acid sequence of NA from A/Cambodia/e0826360/2020 (H3N2).
- SEQ ID NO: 33 Amino acid sequence of HA from A/Darwin/9/2021 (H3N2).
- SEQ ID NO: 34 Amino acid sequence of NA from A/Darwin/9/2021 (H3N2).
- SEQ ID NO: 35 Amino acid sequence of HA from B/Austria/1359417/2021.
- SEQ ID NO: 36 Amino acid sequence of NA from B/Austria/1359417/2021.
- SEQ ID NO: 37 Amino acid sequence of HA from A/Darwin/6/2021 (H3N2).
- SEQ ID NO: 38 Amino acid sequence of NA from A/Darwin/6/2021 (H3N2).
- SEQ ID NO: 39 Amino acid sequence of HA from A/Victoria/4897/2022 (H1 N1).
- SEQ ID NO: 40 Amino acid sequence of NA from A/Victoria/4897/2022 (H1 N1).
- SEQ ID NO: 41 Amino acid sequence of HA from A/Wisconsin/67/2022 (H1 N1).
- SEQ ID NO: 42 Amino acid sequence of NA from A/Wisconsin/67/2022 (H1 N1).
- SEQ ID NO: 43 Amino acid sequence of HA from A/Sydney/5/2021 (H1 N1).
- SEQ ID NO: 44 Amino acid sequence of NA from A/Sydney/5/2021 (H1 N1).
- SEQ ID NO: 45 Amino acid sequence of HA from A/Thailand/8/2022 (H3N2).
- SEQ ID NO: 46 Amino acid sequence of NA from A/Thailand/8/2022 (H3N2).
- SEQ ID NO: 47 Amino acid sequence of HA from A/Massachusetts/18/2022 (H3N2).
- SEQ ID NO: 48 Amino acid sequence of NA from A/Massachusetts/18/2022 (H3N2). DESCRIPTION OF THE FIGURES
- FIG. 1 Domain structure of the Influenza A virus (IAV) HA protein. Domains in HA1 include fusion (F1), vestigial esterase (VE), and receptor-binding domain (RBD). Domains in HA2 include the HA2 ectodomain, transmembrane region (TM), and cytoplasmic tail (CT). The HA head includes the receptor-binding and vestigial esterase subdomains. The stalk (also known as “stem”) contains the HA1 fusion domains and the HA2 ectodomain.
- F1 fusion
- VE vestigial esterase
- RGD receptor-binding domain
- Domains in HA2 include the HA2 ectodomain, transmembrane region (TM), and cytoplasmic tail (CT).
- the HA head includes the receptor-binding and vestigial esterase subdomains.
- the stalk also known as “stem” contains the HA1 fusion domains and the HA2 ectodomain.
- FIG. 2A-B IFN-a release in human PBMCs stimulated with the mRNA-LNP.
- hPBMC from four donors were incubated each in triplicates with 10 pg/ml of the respective mRNA-LNP encoding the HA protein of influenza virus A/Hawaii/70/2019 (H1 N1 pdmO9) in 96-well flat-bottom plate for 24h.
- IFNa was measured via ELISA. Values derived from individual donors (dots) are depicted for each group with line representing the geometric mean with 95% confidence interval (Cl). Between groups geometrical mean ratios (GMR) are depicted on graphs and statistical significance is indicated by a *(CI not containing 1), ns: non-significant.
- GMR geometrical mean ratios
- HA-encoding mRNA-LNP vaccines containing pseudouridine or N1- methylpseudouridine induce substantially lower IFNa levels compared to unmodified mRNA-LNP vaccine in mice.
- GM geometric mean
- Cl 95% confidence interval
- FIG. 3A-B IFN-a release in hPBMCs stimulated with the mRNA-LNP vaccines.
- hPBMC from four donors were incubated each in triplicates with 10 pg/ml of the respective mRNA-LNP vaccines encoding NA protein of influenza virus A/Hong Kong/45/2019 (H3N2) in 96-well flat-bottom plate for 24h.
- IFNa was measured via ELISA. Values derived from individual donors (dots) are depicted for each group with line representing the geometric mean with 95% confidence interval (Cl). Between groups geometrical mean ratios (GMR) are depicted on graphs and statistical significance is indicated by a *(CI not containing 1), ns: nonsignificant.
- GMR geometrical mean ratios
- FIG. 4A-B IFNa release in hPBMCs stimulated with the seasonal influenza 4- and 7- component mRNA vaccine formulations.
- hPBMC from four donors were incubated each in triplicates with 10 pg/ml of the seasonal influenza 4- component (4HA; unmodified, ip and N1-mip) and 7-component (4HA+3NA; unmodified, ip and N1-mip) mRNA-LNP vaccines in 96-well flat-bottom plate for 24h.
- IFNa was measured via ELISA in cell-free supernatants. Values derived from individual donors (dots) are depicted for each group with line representing the geometric mean (GM) with 95% Cl. Between groups geometrical mean ratios (GMR) are depicted on graphs and statistical significance is indicated by a *( Cl not containing 1).
- IFNa levels were determined using ELISA in serum samples collected 18 h after the first immunization. Each dot represents an individual animal, lines depict the geometric mean (GM) with 95% confidence interval (Cl). Between groups geometrical mean ratios (GMR) are depicted on graphs and statistical significance is indicated by a *(CI not containing 1).
- FIG. 5A-D HI titers induced by the 4- or 7-component seasonal influenza mRNA vaccines containing unmodified or modified (ip and N1-mip) nucleosides.
- Control animals received either 0.9% NaCI solution or one tenth of the human dose of the licensed QIVs FLUARIX or FLUZONE High-Dose.
- HI titers against influenza A/Wisconsin/588/2019 H1 N1 pdmO9 (A)
- A/Cambodia/e0826360/2020 (H3N2) (B), B/Washington/02/2019 (C) and B/Phuket/3073/2013 (D) were measured in serum of the mice two weeks post second immunization.
- FIG. 6A-C Nl titers induced by the 7-component seasonal influenza mRNA vaccines containing unmodified or modified (ip and N1-mip) nucleosides.
- Nl titers against influenza A/Wisconsin/588/2019 (H1 N1pdmO9) (A), A/Cambodia/e0826360/2020 (H3N2) (B), B/Washington/02/2019 (C) were measured in serum two weeks post second immunization. Each dot represents an individual animal, lines show the geometrical mean titer (GMT) with 95% confidence interval (Cl).
- FIG. 7A-B 4-component and 7-component seasonal influenza mRNA vaccines induced T cell immune responses after i.m. immunization of mice.
- T cell immune responses were analyzed two weeks post second immunization by intracellular cytokine staining in isolated splenocytes restimulated with 15-mer overlapping peptide libraries spanning the full-length HA or NA proteins of influenza A/Wisconsin/588/2019 (H1 N1pdmO9).
- HA-specific IFNy+TNF+ CD4+ T cells (A) and CD8+ T cells (B) were measured.
- Each dot represents an individual animal, lines depict the geometrical mean (GM) with 95% confidence interval (Cl). Between groups geometrical mean ratio (GMR) are depicted on graphs and statistical significance is indicated by a * (Cl not containing 1).
- GM geometrical mean ratio
- T cell immune responses were analyzed two weeks post second immunization by intracellular cytokine staining in isolated splenocytes restimulated with 15-mer overlapping peptide libraries spanning the full-length HA or NA proteins of influenza A/Wisconsin/588/2019 (H1 N1pdmO9).
- NA-specific IFNy+TNF+ CD4+(A) T cells and CD8+ T cells (B) were measured.
- Each dot represents an individual animal, lines depict the geometrical mean (GM) with 95% confidence interval (Cl). Between groups geometrical mean ratio (GMR) are depicted on graphs and statistical significance is indicated by a * (Cl not containing 1).
- FIG. 9A-D Unmodified and modified 7-component seasonal influenza mRNA vaccines induced HI responses in naive ferrets.
- Two groups were immunized twice by IM route on day 0 and day 28 with full human dose of the licensed split-inactivated QIVs, FLUARIX or FLUZONE HD.
- FIG.10A-C Unmodified and modified 7-component seasonal influenza mRNA vaccines induced Nl responses in naive ferrets.
- Two groups were immunized twice by IM route on day 0 and day 28 with full human dose of the licensed split-inactivated QIVs, FLUARIX or FLUZONE HD.
- FIG. 11 A-D HI responses induced upon i.m. immunization of naive ferrets with 4-component and 8-component seasonal influenza mRNA vaccine formulations.
- HI titers against influenza A/Wisconsin/588/2019 H1 N1pdmO9 (A) , A/Darwin/6/2021 (H3N2) (B), B/Austria/1359417/2021 (C) and B/Phuket/3073/2013 (D) were measured in serum of vaccinated animals collected on Day 55.
- Each dot represents an individual animal, lines depict the geometrical mean (GM) with 95% confidence interval (Cl). Between groups geometrical mean ratio (GMR) are depicted on graphs and statistical significance is indicated by a *(GMR > 3, confidence interval (Cl) not containing 1). The dashed line indicates the HI titer of 40 defined as a surrogate correlate of protection.
- FIG. 12A-D MN titers induced upon i.m. immunization of naive ferrets with 4-component and 8-component seasonal influenza mRNA vaccine formulations.
- MN titers against influenza A/Wisconsin/588/2019 H1 N1 pdmO9 (A), A/Darwin/6/2021 (H3N2) (B), B/Austria/1359417/2021 (C) and B/Phuket/3073/2013 (D) were measured in serum of vaccinated animals collected on Day 55.
- Each dot represents an individual animal, lines depict the geometrical mean (GM) with 95% confidence interval (Cl). Between groups geometrical mean ratio (GMR) are depicted on graphs and statistical significance is indicated by a *(GMR > 3, confidence interval (Cl) not containing 1).
- FIG. 13A-D Nl titers induced upon i.m. immunization of naive ferrets with 8-component seasonal influenza mRNA vaccine formulations.
- Nl titers against influenza A/Wisconsin/588/2019 H1 N1pdmO9 (A), A/Darwin/6/2021 (H3N2) (B), B/Austria/1359417/2021 (C) and B/Phuket/3073/2013 (D) were measured in serum of vaccinated animals collected on Day 55.
- Each dot represents an individual animal, lines depict the geometrical mean (GM) with 95% confidence interval (Cl). Between groups geometrical mean ratio (GMR) are depicted on graphs and statistical significance is indicated by a *(GMR > 3, confidence interval (Cl) not containing 1).
- FIG. 14A-D Viral load in the respiratory tissues post influenza A/Victoria/2570/2019 (H1 N1pdmO9) challenge in ferrets.
- Two groups were immunized twice by IM route on day 0 and day 28 with full human dose of the licensed split-inactivated QIVs, FLUARIX or FLUZONE HD.
- the lower limit of detection (LLOD) for the assay is dependent on the tissue sample weight and the LLOD range is indicated by the dashed lines.
- Severity of bronchiolitis (C) and alveolitis (D) averaged per group (inflammatory changes in the pulmonary parenchyma of animals). For each animal and lung section four tissues slides were evaluated and each dot represents the mean of an individual animal, lines depict the mean of the particular group with 95% confidence intervals (Cis). Between groups difference are depicted on graphs and statistical significance is indicated by a *.
- FIG. 15A-B Viral load in the respiratory tissues post influenza A/Victoria/2570/2019 (H1 N1pdmO9) challenge in ferrets.
- ferrets were challenged with 10 A 5 TCID50 of wild type influenza virus A/Victoria/2570/2019 (H1 N1pdmO9) (via the intratracheal and intranasal route) and euthanized 4 days post challenge infection. Titration of lung tissue (A) and nasal turbinates (B) samples collected on day 4 post challenge with A/Victoria/2570/2019 (H1 N1 pdmO9). Individual titers are shown by group with group mean indicated by solid black line. The lower limit of detection (LLOD) for the assay is dependent on the tissue sample weight and the LLOD range is indicated by the dashed lines.
- LLOD lower limit of detection
- FIG. 16A-C Body temperature changes post influenza A/Victoria/2570/2019 (H1 N1pdmO9) challenge in ferrets.
- FIG. 17A-F Virus titration of throat swabs (A-C) and nose swabs (D-F) Day 0 (pre-challenge) to Day 4 post challenge with influenza A/Victoria/2570/2019 (H1 N1pdmO9).
- FIG. 18A-D Ferrets immunized with 4- and 8-component seasonal influenza mRNA vaccines show reduced macroscopic and microscopic pathological changes in the lungs.
- Four weeks after the last immunization the ferrets were challenged with 10 A 5 TCID50 of wild type influenza virus A/Victoria/2570/2019 (H1 N1pdmO9) (via the intratracheal and intranasal route).
- Relative lung weight on day 4 post challenge with influenza A/Victoria/2570/2019 (H1 N1pdmO9) virus in ferrets Relative lung weights (RLW) reflect the extent and severity of lung lesions since inflamed and/or infiltrated lungs are usually heavier than unaffected lungs. In general RLWs ⁇ 1 are recorded in healthy or slightly pneumonic ferrets. Individual percentage is shown by group with group mean ⁇ SD indicated. A dashed line indicates relative lung weight of 1 .
- FIG. 19A-D 4-component or 7-component seasonal influenza mRNA vaccines induced HI responses against several heterologous influenza strains upon i.m. immunization in mice.
- FIG. 20A-B 4-component or 7-component seasonal influenza mRNA vaccines induced broad heterologous and heterosubtypic HA-binding antibodies upon i.m. immunization in mice.
- HA-specific antibodies were measured in serum of vaccinated animals two weeks post second immunization using multiplex HA-binding assays including recombinant HA proteins from four H1 , one H2, one H5, six H3, one H7, one H10 (A) , two Yamagata and eight Victoria strains (B).
- HA-binding assays including recombinant HA proteins from four H1 , one H2, one H5, six H3, one H7, one H10 (A) , two Yamagata and eight Victoria strains (B).
- FIG. 21A-F 7-component seasonal influenza mRNA vaccine induced heterologous and heterosubtypic Nl responses upon i.m. immunization in mice.
- FIG. 22A-E Heterologous HI responses induced upon i.m. immunization of naive ferrets with 4-component and 8-component Flu Seasonal mRNA vaccine formulations.
- HI titers against influenza A/Brisbane/02/2018 A
- A/California/07/2009 B
- A/South Australia/34/2019 C
- B/Darwin/7/2019 D
- A/Hong Kong/45/2019 E
- GM geometrical mean
- C 95% confidence interval
- the dashed line indicates the HI titer of 40 defined as a surrogate correlate of protection.
- FIG. 23A-G Heterologous Nl responses induced upon i.m. immunization of naive ferrets with 8-component Flu Seasonal mRNA vaccine formulations.
- Nl titers against NA of influenza A/California/07/2009 (A), A/Brevig Mission/1/1918 (B), A/Vietnam/1194/2004 (C), A/Cambodia/e0826360/2020 (D), A/Washington/01/2007 (E), B/Brisbane/60/2008 (F) and B/Washington/02/2019 (G) were measured in serum of vaccinated animals collected on Day 55.
- Each dot represents an individual animal, lines depict the geometrical mean (GM) with 95% confidence interval (Cl). Between groups geometrical mean ratio (GMR) are depicted on graphs and statistical significance is indicated by a *(GMR > 3, confidence interval (Cl) not containing 1).
- the dashed line indicates the Nl titer of 40 defined as a surrogate correlate of protection.
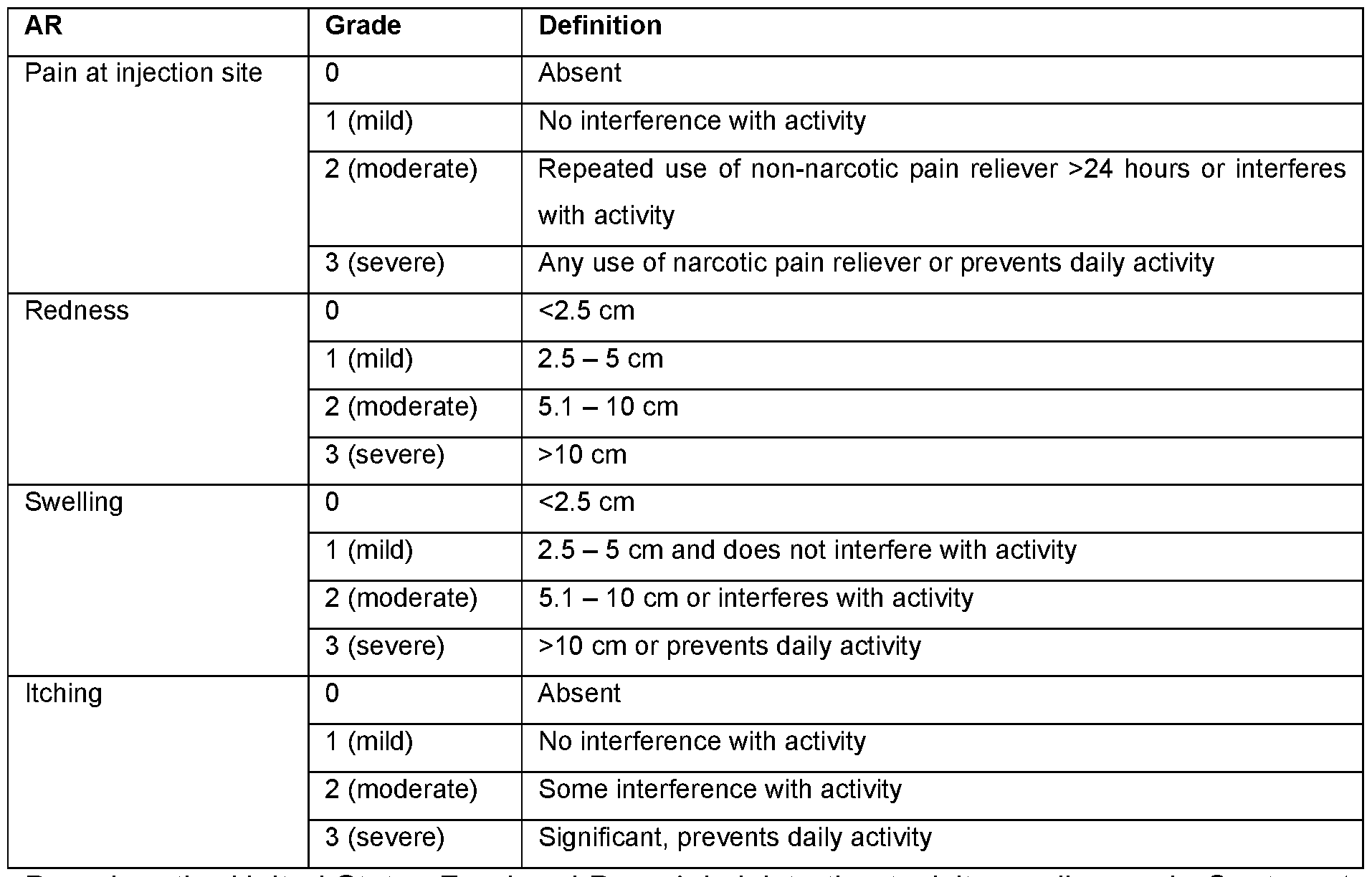
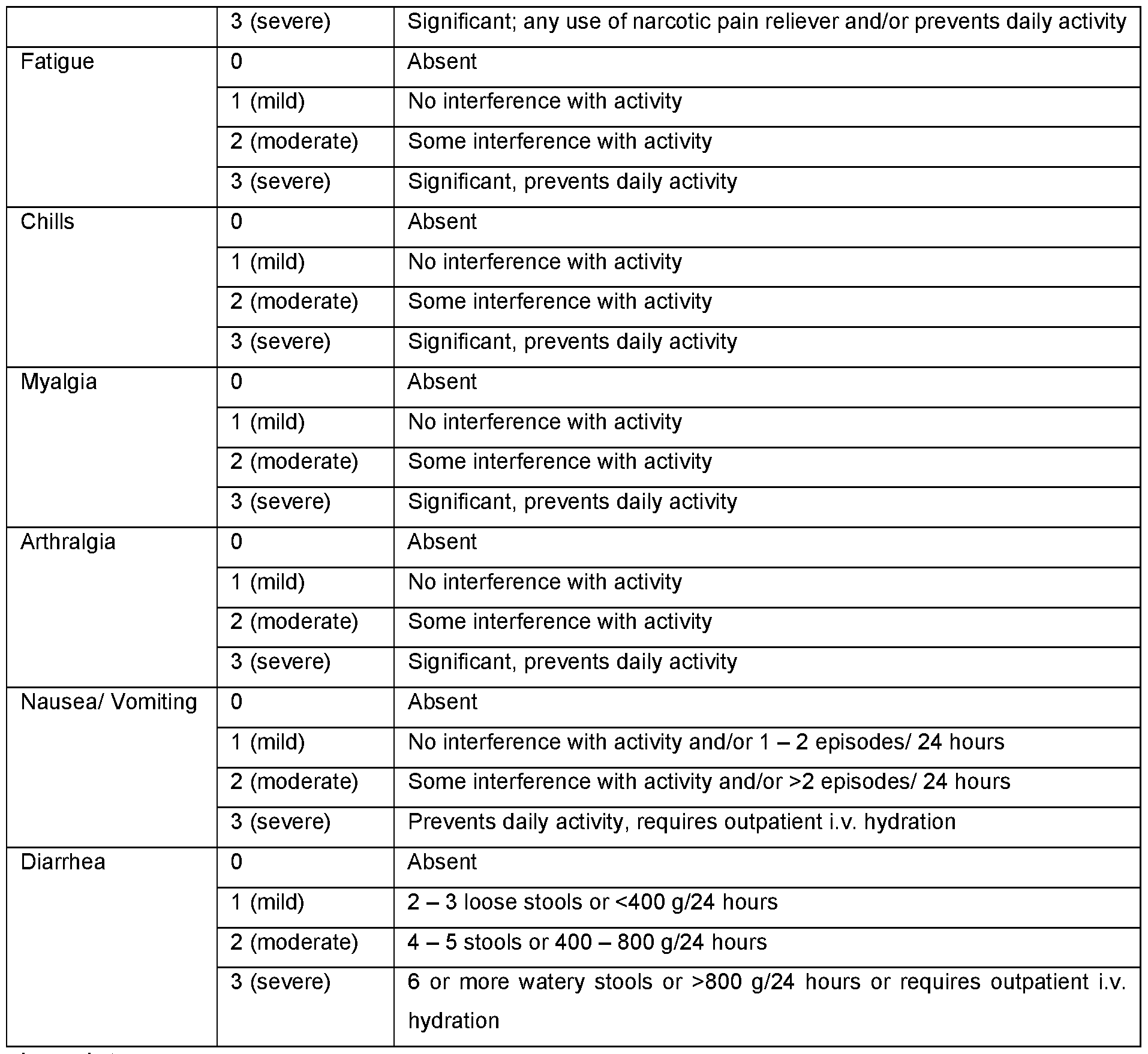
- FIG. 24A-C Reactogenicity assessment of subjects in the CVSQIV Phase I influenza vaccination trial.
- FIG. 24A Solicited Adverse Events in subjects at the indicated mRNA dose levels shown at the bottom of the chart.
- FIG. 24B Solicited Adverse Events in subjects at the indicated mRNA dose levels separated between younger and older adults.
- FIG. 24A-B Grade 0 events at the bottom of the chart above the dose level indication and percentages for increasing grade events arranged vertically.
- FIG. 24C Solicited Adverse Events in subjects at the indicated mRNA dose levels separated for younger and older adults and separated between local and systemic events. Grade 0-1 events at the bottom of the chart above the dose level indication. The percentages for Grade 0-1 versus Grade > 2 are shown.
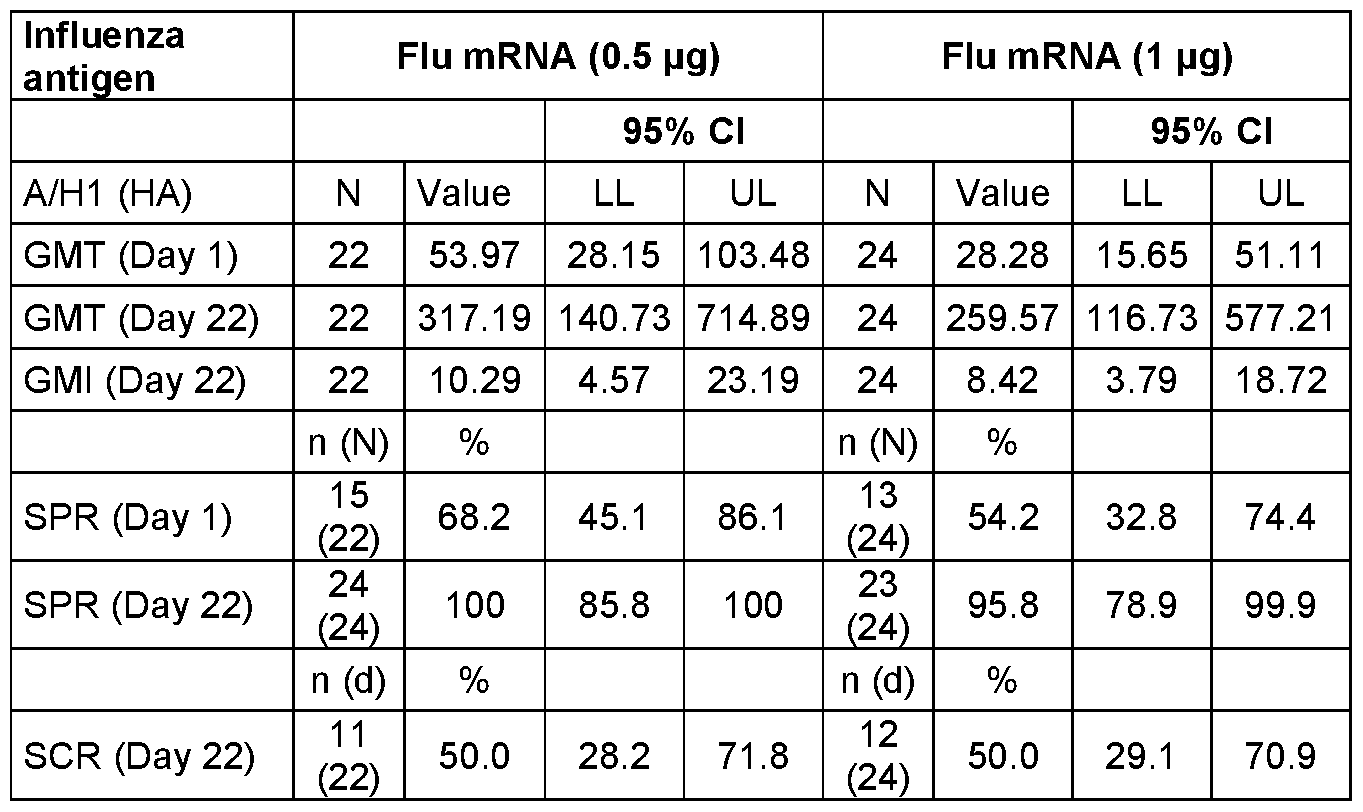
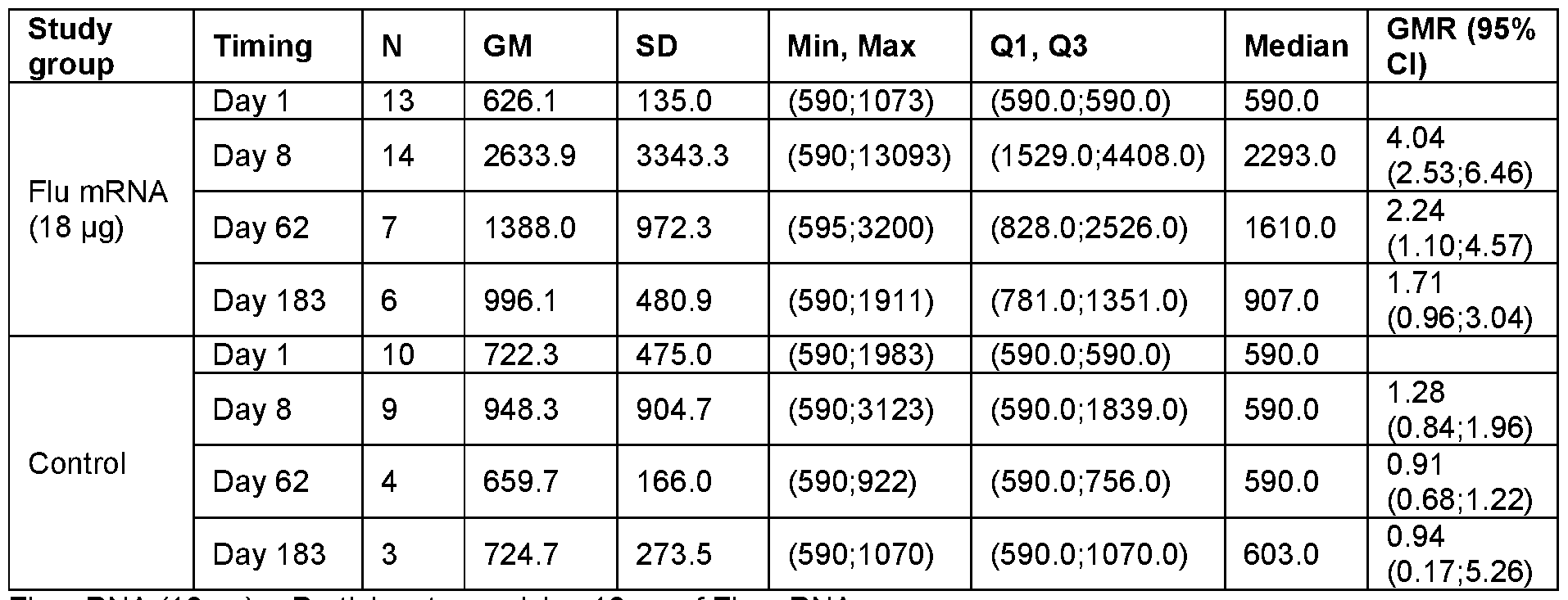
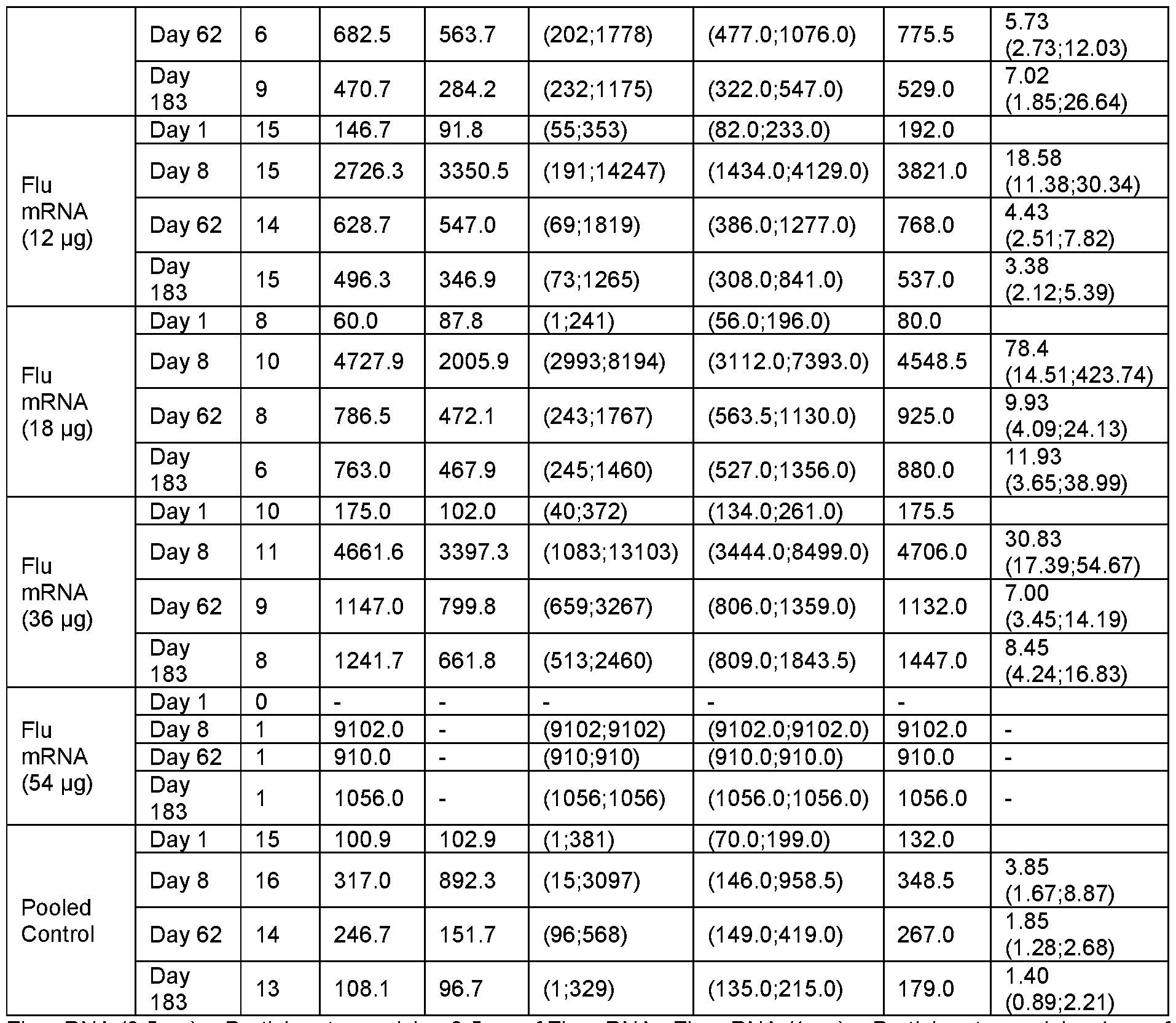
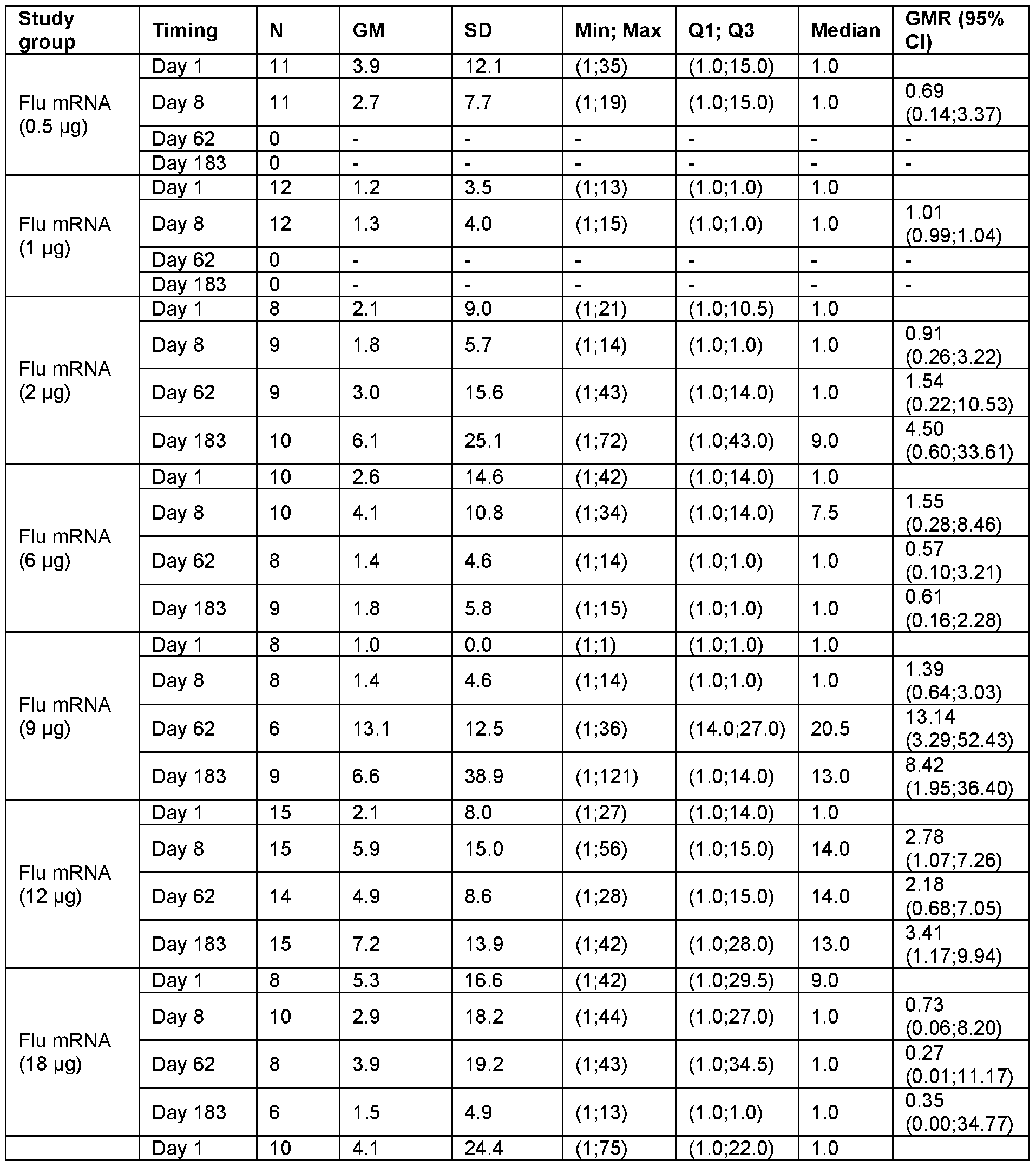
- FIG. 25A-H Graphs show the Geometric Mean Titer (95% Cl) for Hemagglutinin Inhibition Assay (HAI) Assay.
- FIG. 25A, C, E and G show the HAI titers for all subjects at Day 1 , Day 22 and Day 183 at the indicated vaccine mRNA dose levels.
- Data in FIG. 25B, D, F and H are separated between younger adults (YA) and older adults (OA) at the indicated mRNA dose levels.
- Data are shown separately for each of the HA components encoded by the vaccine mRNA: H1 N1 (FIG. 25A- B); H3N2 (FIG. 25C-D); B/Phuket (FIG. 25E-F); and B/Washington (FIG. 25G- H).
- FIG. 26 Seroconversion rates (SCR) from HAI assay.
- the table in upper-left panel shows SCR (is defined as ⁇ 1 :10 pre-vaccination titers, the post-vaccination titers should be >1 :40; if >1 :10 pre-vaccination titers, the post-vaccination titers should be > four-fold increase from baseline).
- Data are shown for each encoded HA, at each dose level and either for all subjects or separated between younger and older adults.
- the graph in the lower left panel shows overall SCR for each encoded HA, at each dose level.
- the graph in the upper right panel shows SCR for each encoded HA, at each dose level, in younger adults.
- the graph in the lower right panel shows SCR for each encoded HA, at each dose level, in older adults.
- FIG. 27 Shows the percentage of study subjects that exhibited a > four-fold increase in anti-HA titer by microneutralization (MN) assay.
- the table in upper-left panel shows the percentage of subjects with > four-fold increase in anti-HA titer by MN assay. Data are shown for each encoded HA, at each dose level, and either for all subjects or separated between younger and older adults.
- the graph in the lower left panel shows overall 4-fold anti-HA increase by MN assay for each encoded HA, at each dose level.
- the graph in the upper right panel shows 4- fold anti-HA increase by MN assay for each encoded HA, at each dose level, in younger adults.
- the graph in the lower right panel shows 4-fold anti-HA increase by MN assay for each encoded HA, at each dose level, in older adults.
- FIG. 28 Shows the percentage of study subjects that exhibited a > four-fold increase in anti-NA titer by enzyme linked lectin assay (ELLA) assay.
- the table in upperleft panel shows the percentage of subjects with > four-fold increase anti-NA titer by ELLA assay. Data are shown for each encoded NA, at each dose level, and either for all subjects or separated between younger and older adults.
- the graph in the lower left panel shows overall 4-fold anti-NA increase by ELLA assay for each encoded NA, at each dose level.
- the graph in the upper right panel shows 4-fold anti-NA increase by ELLA assay for each encoded NA, at each dose level, in younger adults.
- the graph in the lower right panel shows 4- fold anti-NA increase by ELLA assay for each encoded NA, at each dose level, in older adults.
- FIG. 29A-D Shows HI titers induced upon immunization of human healthy adults (18-50 years old) with 1- component, 4-component and 8-component Flu Seasonal mRNA vaccine formulations.
- the control is a Flu D-QIV (FLUARIX, NH 2022- 23).
- FIG. 30A-D Shows Nl titers induced upon immunization of human healthy adults (18-50 years old) with 1- component, 4-component and 8-component Flu Seasonal mRNA vaccine formulations.
- the control is a Flu D-QIV (FLUARIX, NH 2022- 23).
- Nl titers against influenza A/Wisconsin/588/2019 (H1 N1pdmO9) (A), Flu A/Cambodia/e0826360/2020 (H3N2) (B), Flu B/Austria/1359417/2021 (C) and B/Phuket/3073/2013 (D) were measured on Day 29.
- FIG. 31A-D Shows the percentage of human healthy adults (18-50 years old) with solicited events (any; A), local events (B) and systemic events (C) within 7 days of immunization with 1- component, 4-component and 8-component Flu Seasonal mRNA vaccine formulations.
- the control is a Flu D-QIV (FLUARIX, NH 2022- 23).
- (D) shows the overall summary by event including grade 3 events.
- FIG. 32 Shows the percentage of human healthy adults (18-50 years old) with related unsolicited events within 7 days of immunization with 1- component, 4- component and 8-component Flu Seasonal mRNA vaccine formulations.
- the control is a Flu D-QIV (FLUARIX, NH 2022-23).
- sequence listing in electronic format, which is part of the description (WIPO standard ST.26).
- the information contained in the sequence listing is incorporated herein by reference in its entirety.
- sequence listing also provides additional detailed information, e.g. regarding certain structural features, sequence optimizations, GenBank (NCBI) or GISAID (epi) identifiers, or additional detailed information regarding its coding capacity.
- sequences e.g. amino acid sequences or nucleic acid sequences, are explicitly incorporated herein by reference. Accordingly, these sequences constitute an integral part of the underlying description.
- Protective immune responses induced by vaccination against Influenza viruses are primarily directed to the viral HA protein, which is a glycoprotein on the surface of the virus responsible for interaction of the virus with host cell receptors.
- HA proteins on the virus surface are homotrimers of HA protein monomers that are enzymatically cleaved to yield amino-terminal HA1 and carboxy-terminal HA2 polypeptides.
- hemagglutinin proteins are comprised of several domains: a globular head domain, a stalk domain (also referred to as a stem domain), a transmembrane domain, and a cytoplasmic domain (see FIG. 1 , Russell et al., 2021).
- a host cell e.g., a eukaryotic cell such as a human cell
- the hemagglutinin protein recognizes and binds to sialic acid of a receptor on the surface of a host cell facilitating attachment of the virus to the host cell.
- the hemagglutinin protein undergoes a pH-dependent conformational change that allows for the hemagglutinin protein to facilitate fusion of the viral envelope with the endosome membrane of host cell and entry of the viral nucleic acid into the host cell.
- the globular head consists exclusively of the major portion of the HA1 polypeptide, whereas the stem that anchors the HA protein into the viral lipid envelope is comprised of HA2 and part of HA1 .
- the globular head of a HA protein includes two domains: the receptor binding domain (RBD), a domain that includes the sialic acid-binding site, and the vestigial esterase domain, a smaller region just below the RBD.
- RBD receptor binding domain
- Influenza viruses are classified based on the amino acid sequences of the viral hemagglutinin protein and/or the amino acid sequence of the viral neuraminidase (NA). The differences in amino acid sequence between HA proteins of different subtypes are largely found within the sequence of the head domain of the protein.
- the amino acid sequence of the stalk domain is considered to be more conserved between HA subtypes compared to sequences of the head domain. Domains of the HA protein may be predicted using conventional methods known in the art.
- Antibodies against Influenza mainly target variable antigenic sites in the globular head of HA and thus, neutralize only antigenically closely related viruses.
- Current trivalent and quadrivalent influenza vaccines elicit antibody responses to the vaccine strains (e.g.
- homologous immune response or to closely related isolates, but rarely extend to more diverged strains within a subtype/lineage (e.g. heterologous or intrasubtypic immune response) or to strains belonging to different subtypes/lineages (e.g. heterosubtypic immune response).
- the inventors overcame the drawbacks of the prior art by providing an immunogenic composition for use in the treatment or prophylaxis of an infection with an Influenza virus, wherein the immunogenic composition comprises:
- a fourth nucleic acid encoding a HA antigen of a second strain of Influenza B virus, wherein an immune response is elicited against HA antigens of said strains of first and second subtypes of Influenza A virus, said first and, optionally, second strains of Influenza B virus and at least one further HA antigen subtype of Influenza A virus, being different from any of the HA antigen subtypes of Influenza A virus encoded by a nucleic acid present in the composition.
- the immunogenic compositions for use according to the invention induce a broad, rapid, and robust cross-reactive immune response against Influenza virus, such as Influenza A and/or B.
- the immunogenic compositions for use according to the invention induce a broad, rapid, and robust homologous, heterologous and heterosubtypic immune response against Influenza virus, such as Influenza A and/or B.
- the immunogenic compositions for use according to the invention elicit antibody responses to the vaccine strains and to closely related strains, but also to further strains belonging either to the same or to a different subtype/lineage.
- the immunogenic compositions for use according to the invention elicit antibody responses to Influenza virus strains that are antigenically distinct from the vaccine strains.
- the immunogenic compositions for use according to the invention elicit antibody responses to Influenza virus strains being different from any of the strains having an HA encoded by an mRNA present in the composition.
- the immunogenic compositions for use according to the invention elicit antibody responses to Influenza viruses with a different geographical origin and/or year of isolation than any of the strains having an HA and/or NA encoded by an mRNA present in the composition.
- the immunogenic compositions for use according to the invention elicit antibody responses to HA antigen subtype of Influenza A virus, being different from any of the HA antigen subtypes of Influenza A virus encoded by a nucleic acid present in the composition.
- the immunogenic compositions for use according to the invention elicit antibody responses to HA antigen subtype of Influenza A virus, being different from any of the HA antigen of a strain of Influenza B virus, being different from any of the HA antigens of a strain of Influenza B virus encoded by a nucleic acid present in the composition.
- the immunogenic compositions for use according to the invention induce cross-reactive binding and functional anti-HA responses.
- the immunogenic compositions for use according to the invention protect individuals from strains of Influenza virus that are not present in the immunogenic composition.
- the immunogenic compositions for use according to the invention protect individuals from homologous, heterologous and heterosubtypic strains of influenza virus.
- the immunogenic compositions for use according to the invention have at least some of the following advantageous features:
- composition/vaccine for intramuscular administration
- Influenza virus suitably Influenza A virus and/or Influenza B virus;
- Influenza virus Longevity of the induced immune responses against Influenza virus, suitably Influenza A virus and/or Influenza B virus;
- AD antibody dependent enhancement
- nucleic acid-based composition/vaccine Advantageous stability characteristics of the nucleic acid-based composition/vaccine
- the invention relates to an immunogenic composition for use in the treatment or prophylaxis of an infection with an Influenza virus, wherein the immunogenic composition comprises: (a) a first nucleic acid encoding a hemagglutinin (HA) antigen of a strain of a first subtype of Influenza A virus;
- a fourth nucleic acid encoding a HA antigen of a second strain of Influenza B virus, wherein an immune response is elicited against HA antigens of said strains of first and second subtypes of Influenza A virus, said first and, optionally, second strains of Influenza B virus and at least one further HA antigen subtype of Influenza A virus, being different from any of the HA antigen subtypes of Influenza A virus encoded by a nucleic acid present in the composition.
- hemagglutinin hemagglutinin protein
- HA hemagglutinin protein
- said strain of said first subtype of Influenza A virus of (a) and said strain of said second subtype of Influenza A virus of (b) are different.
- Influenza A viruses are divided into subtypes based on the antigenic properties of their hemagglutinin (HA) and neuraminidase (NA) surface proteins.
- HA subtypes e.g. H1 , H2, H3, H4, H5, H6, H7, H8, H9, H10, H11 , H12, H13, H14, H15, H16, H17 and H18 subtypes
- NA subtypes e.g. H1 , H2, H3, H4, H5, H6, H7, H8, H9, H10, H11 , H12, H13, H14, H15, H16, H17 and H18 subtypes
- NA subtypes e.g. H1 , H2, H3, H4, H5, H6, H7, H8, H9, N10 and N11 subtypes
- H1 N1 , H2N2, H3N2, H5N1 , H7N9 or H10N8 e.g. H1 N1 , H
- said first strain of Influenza B virus of (c) and said second strain of Influenza B virus of (d) are identical.
- (c) and (d) are identical.
- said first strain of Influenza B virus of (c) and said second strain of Influenza B virus of (d) are different.
- (c) and (d) are different.
- said first and/or second subtype of Influenza A virus is selected from influenza A viruses characterized by a HA selected from the group consisting of H1 , H2, H3, H4, H5, H6, H7, H8, H9, H10, H11 , H12, H13, H14, H15, H16, H17 and H18, suitably from the group consisting of H1 , H3, H5, H7, H9, and H10, more suitably from the group consisting of H1 and H3.
- a HA selected from the group consisting of H1 , H2, H3, H4, H5, H6, H7, H8, H9, H10, H11 , H12, H13, H14, H15, H16, H17 and H18, suitably from the group consisting of H1 , H3, H5, H7, H9, and H10, more suitably from the group consisting of H1 and H3.
- said first and/or second subtype of Influenza A virus is selected from influenza A viruses characterized by a neuraminidase (NA) selected from the group consisting of N1 , N2, N3, N4, N5, N6, N7, N8, N9, N10, and N11 , suitably selected from the group consisting of N1 , N2, and N8, more suitably selected from the group consisting of N1 and N2.
- NA neuraminidase
- neuroaminidase Asperably, the terms “neuraminidase”, “neuraminidase protein”, and “NA” may be used interchangeably throughout and refer to a neuraminidase protein that may be present on the surface of an Influenza virus.
- said first and/or second subtype of Influenza A virus is selected from the group consisting of H1 N1 , H1 N2, H2N2, H3N1 , H3N2, H3N8, H5N1 , H5N2, H5N3, H5N8, H5N9, H7N1 , H7N2, H7N3, H7N4, H7N7, H7N9, H9N2, H10N7 and H10N8, suitably H1 N1 and H3N2.
- said first subtype of Influenza A virus is a subtype of Influenza A Group 1 , suitably influenza A subtype H1 , H2, H5, H6, H8, H9, H11 , H12, H13, H16, H17 or H18, more suitably H1.
- said first subtype of Influenza A virus is Influenza A H1 N1 subtype.
- said second subtype of Influenza A virus is a subtype of Influenza A Group 2, suitably influenza A subtype H3, H4, H7, H10, H14 and H15, more suitably H3.
- said second subtype of Influenza A virus is Influenza A H3N2 subtype.
- said strain of first and/or second subtype of Influenza A virus is selected from the group consisting of A/Thailand/8/2022 (H3N2)-like virus, A/Massachusetts/18/2022 (H3N2)-like virus, AA/ictoria/4897/2022 (H1 N1)pdmO9-like virus, A/Wisconsin/67/2022 (H1 N1)pdmO9-like virus, A/Sydney/5/2021 (H1 N1)pdmO9-like virus, A/Victoria/2570/2019 (H1 N1)pdmO9-like virus, A/Darwin/9/2021 (H3N2)-like virus, A/Wisconsin/588/2019 (H1 N1)pdmO9-like virus, A/Darwin/6/2021 (H3N2)-like virus, A/Cambodia/e0826360/2020 (H3N2)-like virus, A/Guangdong-Maonan/SWL
- first strain of first subtype of Influenza A virus is a strain of H1 N1 and is selected from the group consisting of A/Victoria/4897/2022 (H1 N1)pdmO9-like virus, A/Wisconsin/67/2022 (H1 N1)pdmO9-like virus, A/Sydney/5/2021 (H1 N1)pdmO9-like virus, A/Beijing/262/95(H1 N1)-like virus, A/New Caledonia/20/99(H1 N1)-like virus, A/Solomon lslands/3/2006 (H1 N1)-like virus, A/Brisbane/59/2007 (H1 N1)-like virus, A/California/7/2009 (H1 N1)-like virus, A/California/7/2009 (H1 N1)pdmO9-like virus, A/Michigan/45/2015 (H1 N1)pdm09-like virus, A/V
- said second strain of Influenza A virus is H3N2 and is selected from the group consisting of A/Thailand/8/2022 (H3N2)-like virus, A/Massachusetts/18/2022 (H3N2)-like virus, A/Sydney/5/97(H3N2)-like virus, A/Moscow/10/99(H3N2)-like virus, A/Panama/2007/99, A/Fujian/411/2002(H3N2)-like virus, A/Wyoming/3/ 2003, A/
- H3N2 Kong/2671/2019 (H3N2)-like virus, A/Hong Kong/45/2019 (H3N2)-like virus, A/Switzerland/9715293/2013 (H3N2)-like virus, A/South Australia/55/2014, A/Norway/466/ 2014, A/Stockholm/6/ 2014, A/Hong Kong/4801/2014 (H3N2)-like virus, A/Singapore/INFIMH- 16-0019/2017 (H3N2)-like virus, A/Switzerland/8060/2017 (H3N2)-like virus, A/Kansas/14/2017 (H3N2)-like virus and A/South Australia/34/2019 (H3N2)-like virus.
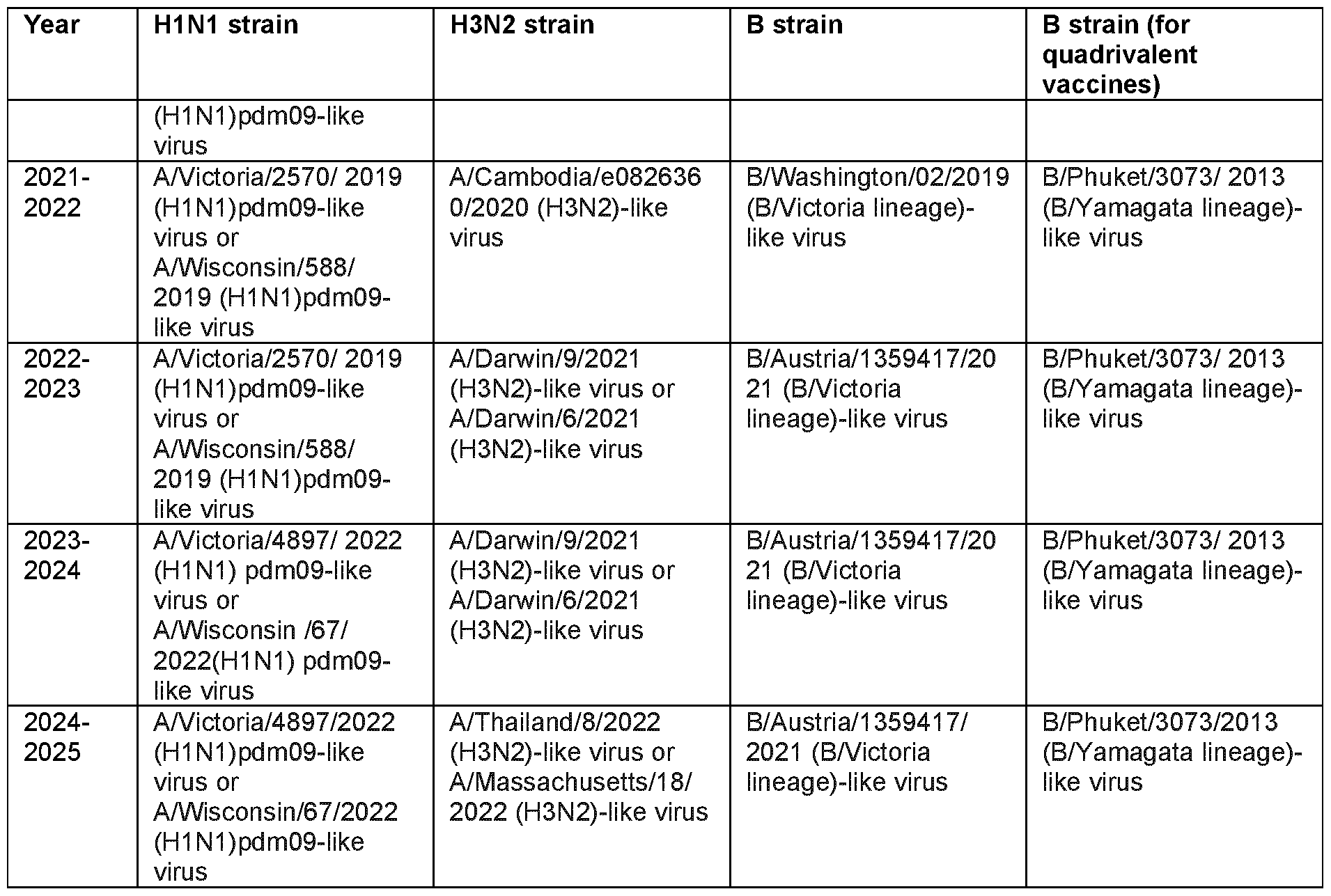
- said strain of said first and/or second subtype of Influenza A virus is selected from an Influenza A virus as listed in Table 1 and/or Table 2.
- said strain of said first and/or second subtype of Influenza A virus is selected from an Influenza A virus which is recommended for Influenza virus vaccine composition by the WHO (https://www.who.int/teams/global-influenza- programme/vaccines/who-recommendations).
- Table 1 Recommended composition of Influenza virus vaccines for use in the 1998- 2025 northern hemisphere influenza season
- Table 2 Recommended composition of Influenza virus vaccines for use in the 1999- 2024 southern hemisphere influenza season
- said first strain of Influenza B virus is selected from the group consisting of B/Victoria lineage and B/Yamagata lineage.
- Influenza B viruses are categorized into two distinct lineages: B/Victoria/2/1987-like (B/Victoria lineage) and B/Yamagata/16/1988-like (B/Yamagata lineage) viruses that have been circulating worldwide since 1983.
- said first and/or second strain of Influenza B virus is selected from the group consisting of B/Beijing/184/93-like virus, B/Harbin/94-like virus, B/Shangdong/7/97-like virus, B/Yamanashi/166/98-like virus, B/Sichuan/379/99-like virus, B/Guangdong/120/2000, B/Johannesburg/5/99, B/Victoria/504/2000, B/Hong Kong/330/2001-like virus, B/Hong Kong/1434/2002, B/Brisbane/32/2002, B/Shanghai/361/2002-like virus, B/Jiangsu/10/2003, B/J ilin/20/2003, B/Malaysia/2506/2004-like virus, B/Malaysia/2506/2004 virus, B/Ohio/1/2005, B/Florida/4/2006-like virus, B/Brisbane/3/2007,
- said first strain of Influenza B virus is selected from an Influenza B virus as listed in Table 1 and/or Table 2.
- said first strain of Influenza B virus is selected from an Influenza B virus which is recommended for Influenza virus vaccine composition by the WHO (https://www.who.int/teams/global-influenza-programme/vaccines/who-recommendations).
- said first strain and said second strain of Influenza B are a strain of B/Victoria lineage.
- said first strain of Influenza B is a strain of B/Victoria lineage.
- said second strain of Influenza B is a strain of B/Yamagata lineage.
- said first strain of Influenza B virus is selected from the group consisting of B/Austria/1359417/2021 (B/Victoria lineage)-like virus, B/Washington/02/2019 (B/Victoria lineage)-like virus, B/Colorado/06/2017-like virus (B/Victoria/2/87 lineage), B/Brisbane/60/2008-like virus and B/Colorado/06/2019 (B/Victoria lineage)-like virus.
- said first subtype of Influenza A virus is of Influenza A H1 N1
- said second subtype of Influenza A virus is of influenza A H3N2
- said first strain of Influenza B virus is of Influenza B/Victoria lineage.
- the immunogenic composition further comprises:
- said first strain of Influenza B virus of (c) and said second strain of Influenza B virus of (d) are identical.
- said first and second strains of Influenza B are a strain of B/Victoria lineage.
- (c) and (d) are identical. In some embodiments, said first strain of Influenza B virus of (c) and said second strain of Influenza B virus of (d) are different.
- said first strain of Influenza B is a strain of B/Victoria lineage.
- said second strain of Influenza B is a strain of B/Yamagata lineage.
- (c) and (d) are different.
- said second strain of Influenza B virus is B/Phuket/3073/2013 (B/Yamagata lineage)-like virus.
- said first subtype of Influenza A virus is of Influenza A H1 N1
- said second subtype of Influenza A virus is of influenza A H3N2
- said first strain of Influenza B virus is of influenza B/Victoria lineage
- said second strain of Influenza B virus is of Influenza B/Yamagata.
- said HA antigen encoded by said first, second, third and/or fourth nucleic acid comprises or consists of an amino acid sequence having at least 90%, 95%, 98% or 99% identity to the amino acid sequence set forth in any one of SEQ ID NO: 1 , 3, 5, 7, 9, 11 , 13, 15, 17, 19, 21 , 23, 25, 27, 29, 31 , 33, 35, 37, 39, 41 , 43, 45 or 47 or fragment or variant thereof.
- said HA antigen encoded by said first, second, third and/or fourth nucleic acid antigen comprises or consists of the amino acid sequence set forth in any one of SEQ ID NO: 1 , 3, 5, 7, 9, 11 , 13, 15, 17, 19, 21 , 23, 25, 27, 29, 31 , 33, 35, 37, 39, 41 , 43, 45 or 47 or fragment or variant thereof.
- said HA antigen encoded by said third and/or fourth nucleic acid comprises or consists of an amino acid sequence having at least 90%, 95%, 98% or 99% identity to the amino acid sequence set forth in any one of SEQ ID NO: 5, 7, 17 or 35, or fragment or variant thereof.
- said HA antigen encoded by said third and/or fourth nucleic acid comprises or consists of an amino acid sequence set forth in any one of SEQ ID NO: 5, 7, 17 or 35, or fragment or variant thereof.
- said HA antigen encoded by said first and/or second nucleic acid comprises or consists of an amino acid sequence having at least 90%, 95%, 98% or 99% identity to the amino acid sequence set forth in any one of SEQ ID NO: 1 , 3, 9, 11 , 13, 15, 19, 21 , 23, 25, 27, 29, 31 , 33, 37, 39, 41 , 43, 45 or 47 or fragment or variant thereof.
- said HA antigen encoded by said first and/or second nucleic acid comprises or consists of an amino acid sequence set forth in any one of SEQ ID NO: 1 , 3, 9, 11 , 13, 15, 19, 21 , 23, 25, 27, 29, 31 , 33, 37, 39, 41 , 43, 45 or 47 or fragment or variant thereof.
- said HA antigen encoded by said first nucleic acid comprises or consists of an amino acid sequence having at least 90%, 95%, 98% or 99% identity to the amino acid sequence set forth in any one of SEQ ID NO: 1 , 11 , 19, 23, 27, 29, 39, 41 or 43, or fragment or variant thereof.
- said HA antigen encoded by said first nucleic acid comprises or consists of an amino acid sequence set forth in any one of SEQ ID NO: 1 , 11 , 19, 23, 27, 29, 39, 41 or 43, or fragment or variant thereof.
- said HA antigen encoded by said second nucleic acid comprises or consists of an amino acid sequence having at least 90%, 95%, 98% or 99% identity to the amino acid sequence set forth in any one of SEQ ID NO: 3, 9, 13, 15, 21 , 25, 31 , 33, 37, 45 or 47 or fragment or variant thereof.
- said HA antigen encoded by said second nucleic acid comprises or consists of an amino acid sequence set forth in any one of SEQ ID NO: 3, 9, 13, 15, 21 , 25, 31 , 33, 37, 45 or 47 or fragment or variant thereof.
- said HA antigen encoded by said first, second, third and/or fourth nucleic acid is a polypeptide comprising a full-length Influenza HA protein.
- said HA antigen encoded by said first, second, third and/or fourth nucleic acid is a polypeptide consisting of a full-length Influenza HA protein.
- said HA antigen encoded by said first, second, third and/or fourth nucleic acid is a fragment of a hemagglutinin protein, such as a truncated hemagglutinin protein.
- the fragment is a headless hemagglutinin, meaning the fragment does not comprise the head domain.
- the fragment comprises a portion of the head domain.
- the fragment is a stalk domain.
- the fragment does not comprise the cytoplasmic domain.
- the fragment does not comprise the transmembrane domain.
- the fragment may be referred to as a soluble or secreted hemagglutinin protein or fragment.
- the composition does not comprise a nucleic acid encoding a HA antigen from an influenza strain which is not recommended by WHO.
- the composition does not comprise a nucleic acid encoding a HA antigen identified or designed by machine learning.
- the elicited immune response is homologous, heterosubtypic, and optionally heterologous or intrasubtypic.
- the elicited immune response is further heterologous or intrasubtypic.
- an immune response is elicited against HA antigens that are antigenically distinct from any of the HA antigen encoded by a nucleic acid present in the compositions.
- Subtypes and lineages of Influenza virus can be further divided into different genetic “’’clades (also called “groups”) and “sub-clades” (also called “sub-groups”) based on the similarity of their HA gene sequences.
- Clades and sub-clades are shown on phylogenetic trees as groups and sub-groups of viruses that usually have similar genetic changes (i.e. nucleotide or amino acid changes) and have a single common ancestor.
- Clades and sub-clades that are genetically different from others are not necessarily antigenically different. Viruses from a specific clade or sub-clade may not have a mutation that impacts host immunity in comparison to other clades or sub-clades.
- antigenic properties is used to describe the immune response triggered by the antigens, e.g. HA and/or NA, on a particular virus.
- said at least one further HA antigen subtype of Influenza A virus is selected from influenza A viruses characterized by a HA selected from the group consisting of H1 , H2, H3, H4, H5, H6, H7, H8, H9, H10, H11 , H12, H13, H14, H15, H16, H17 and H18, suitably from the group consisting of H1 , H2, H3, H5, H7 and H 10, more suitably from the group consisting of H2, H5, H7 and H10.
- said at least one further HA antigen subtype of Influenza A virus is selected from influenza A viruses characterized by a neuraminidase (NA) selected from the group consisting of N1 , N2, N3, N4, N5, N6, N7, N8, N9, N10, and N11 , suitably selected from the group consisting of N1 , N2, N8 and N9.
- NA neuraminidase
- said at least one further HA antigen subtype of Influenza A virus is selected from the group consisting of H1 N1 , H1 N2, H2N2, H3N1 , H3N2, H3N8, H5N1 , H5N2, H5N3, H5N8, H5N9, H7N1 , H7N2, H7N3, H7N4, H7N7, H7N9, H9N2, H10N7 and H10N8, suitably H1 N1 , H2N2, H3N2, H5N1 , H7N9 and H10N8.
- said at least one further HA antigen subtype of Influenza A virus is from Influenza A Group 1 , suitably influenza A subtype H1 , H2, H5, H6, H8, H9, H11 , H12, H13, H16, H17 or H18, more suitably H1 , H2 or H5.
- said at least one further HA antigen subtype of Influenza A virus is of H2N2 or H5N1.
- said at least one further HA antigen subtype of Influenza A virus is from a subtype of Influenza A Group 2, suitably influenza A subtype H3, H4, H7, H10, H14 or H15, more suitably H3, H7 or H10.
- said at least one further HA antigen subtype of Influenza A virus is of H7N9 or H10N8.
- said strain of said at least one further HA antigen subtype of Influenza A virus is selected from the group consisting of H2/Singapore 1957, H5A/ietnam/2004, H7/Shanghai/2013 and H10/Jiangxi Donghu/2013.
- an immune response is elicited against HA antigens of Influenza A subtypes H1 , H3 and at least one, suitably all, HA antigen of Influenza A subtype H2, H5, H7 or H10.
- an immune response is further elicited against at least one further HA antigen of a strain of Influenza B virus, being different from any of the HA antigens of a strain of Influenza B virus encoded by a nucleic acid present in the composition.
- said at least one further HA antigen of a strain of Influenza B virus is derived from a strain selected from the group consisting of B/Victoria lineage and B/Yamagata lineage.
- said at least one further HA antigen of a strain of Influenza B virus is a strain of B/Victoria lineage.
- said at least one further HA antigen of a strain of Influenza B virus is a strain of B/Yamagata lineage.
- said at least one further HA antigen of a strain of Influenza B virus is selected from the group consisting of B/Austria/1359417/2021 (B/Victoria lineage)-like virus, B/Washington/02/2019 (B/Victoria lineage)-like virus, B/Colorado/06/2017-like virus (B/Victoria/2/87 lineage), B/Brisbane/60/2008-like virus, B/Colorado/06/2019 (B/Victoria Iineage)-like virus, B/lllinois/NHRC_FDX51486/2015, B/Oman/4241/2019,
- said at least one further HA antigen of a strain of Influenza B virus is selected from the group consisting of B/Austria/1359417/2021 (B/Victoria lineage)-like virus, B/Washington/02/2019 (B/Victoria lineage)-like virus, B/Colorado/06/2017-like virus (B/Victoria/2/87 lineage), B/Brisbane/60/2008-like virus, B/Colorado/06/2019 (B/Victoria lineage)-like virus, B/lllinois/NHRC_FDX51486/2015, B/Oman/4241/2019, B/lllinois/NHRC_18512/2017, B/California/BRD12452N/2017, B/lndia/Pun-1922338/2019, B/Japan/8858/2019, B/Washington/02/2019 and B/Stockholm/7/2019.
- said at least one further HA antigen of a strain of Influenza B virus is selected from the group consisting of B/Phuket/3073/2013 and B/Quebec/70/2015.
- an immune response is further elicited against at least one HA antigen of a heterologous or intrasubtypic strain of Influenza A virus.
- said heterologous or intrasubtypic strain of Influenza A virus is selected from the group consisting of H1/Michigan/2015, H1/Hawaii/2019, H1/Christchurch/2010, H1/California/2009, H3/Finland/2004, H3/Hong Kong 2019, H3/Perth/2009, H3Bejing/1992, H3/Philippines/1982 and H3/Hong Kong/1968.
- said heterologous or intrasubtypic strain of Influenza A virus is selected from the group consisting of H1/Michigan/2015, H1/Hawaii/2019, H1/Christchurch/2010 and H1/California/2009.
- said heterologous or intrasubtypic strain of Influenza A virus is selected from the group consisting of H3/Finland/2004, H3/Hong Kong 2019, H3/Perth/2009, H3Bejing/1992, H3/Philippines/1982 and H3/Hong Kong/1968.
- an immune response is elicited against at least three, four, five, six, seven, eight, nine, ten, eleven, twelve, thirteen, fourteen, fifteen HA antigens of at least three, four, five, six, seven, eight, nine, ten subtypes of Influenza A virus, including said first and second subtypes of Influenza A virus and/or at least three four, five, six, seven, eight, nine, ten, eleven, twelve, thirteen, fourteen, fifteen HA antigens of Influenza B virus, including against HA antigens of said first strain of Influenza B virus.
- an immune response is elicited against at least three, four, five, six, seven, eight, nine, ten, eleven, twelve, thirteen, fourteen, fifteen HA antigens of at least three, four, five, six, seven, eight, nine, ten subtypes of Influenza A virus, including said first and second subtypes of Influenza A virus and/or at least three four, five, six, seven, eight, nine, ten, eleven, twelve, thirteen, fourteen, fifteen HA antigens of Influenza B virus, including against HA antigens of said first and second strains of Influenza B virus.
- an immune response is elicited against at least one HA antigen of Influenza A from each of subtypes H1, H2, H3, H5, H7 and H10 and at least one HA antigen of Influenza B from B/Victoria lineage.
- an immune response is elicited against at least one HA antigen of Influenza A from each of subtypes H1, H2, H3, H5, H7 and H10, at least one HA antigen of Influenza B from B/Victoria lineage and at least one strain of Influenza B from B/Yamagata lineage.
- the nucleic acids suitably mRNAs, encoding said HA antigens are present in equimolar proportions.
- the ratio of (a):(b):(c) is 1:1:1.
- the ratio of (a):(b):(c):(d) is 1:1:1 :1.
- the nucleic acids suitably mRNAs, encoding said HA antigens are not present in equimolar proportions.
- the dose (e.g. weight dose or molar dose, suitably weight dose) of (c) and/or (d) is different compared to the dose (e.g. weight dose or molar dose, suitably weight dose) of (a) and/or (b).
- the ratio of (a):(b):(c) is comprised between 1:1:1.5 and 1:1:20.
- the ratio of (a):(b):(c) is comprised between 1:1:1.5 and 1:1:5, optionally between 1:1:2 and 1:1:5, optionally between 1:1:3 and 1:1:5, optionally between 1:1:4 and 1:1:5, optionally between 1:1:1.5 and 1:1:4, optionally between 1:1:1.5 and 1:1:3, optionally between 1:1:2 and 1:1:4, optionally between 1:1:2 and 1:1:3.
- the ratio of (a):(b):(c) is greater than 1:1:5 and lower or equal than 1:1:20, optionally greater than 1:1:5 and lower or equal to 1:1:15, optionally greater than 1:1:5 and lower or equal to 1:1:12, optionally greater than 1:1:5 and lower or equal to 1:1:10, optionally greater than 1:1:5 and lower or equal to 1:1:8.
- the ratio of (a):(b):(c) is comprised between 1:1:5.5 and 1:1:20, optionally between 1:1:5.5 and 1:1:15, optionally between 1:1:5.5 and 1:1:12, optionally between 1:1:5.5 and 1:1:10, optionally between 1:1:5.5 and 1:1:8, optionally between 1:1:6 and 1:1:20, optionally between 1:1:6 and 1:1:15, optionally between 1:1:6 and 1:1:12, optionally between 1:1:6 and 1:1:10, optionally between 1:1:6 and 1:1:8, optionally between 1:1:8 and 1:1:20, optionally between 1:1:8 and 1:1:15, optionally between 1:1:8 and 1:1:12, optionally between 1:1:8 and 1:1:10.
- the ratio of (a):(b):(c) is selected from about 1:1:1.5, about 1:1:2, about 1:1:2.2, about 1:1:2.4:2.4, about 1:1:2.6, about 1:1:2.8, about 1:1:3, about 1:1:3.2, about 1:1:3.4, about 1 :1 :3.6, about 1 :1 :3.8, about 1:1:4, about 1 :1 :4.2, about 1 :1 :4.4, about 1 :1 :4.6, about 1:1:4.8, about 1:1:5, about 1:1:5.5, about 1:1:6, about 1:1:6.5, about 1:1:7, about 1:1:7.5, about 1:1:8, about 1:1:8.5, about 1:1:9, about 1:1:9.5, about 1:1:10, about 1:1:10.5, about 1:1:11, about 1:1:11.5, about 1:1:12, about 1:1:12.5, about 1:1:13, about 1:1:13.5, about 1:1:14, about 1:1:14.5, about 1:1:15, about 1:1:15.5, about 1:1:16, about 1:1:16.5, about 1:1:17, about 1:1:17.5, about 1:1:
- the ratio of (a):(b):(c) is 1:1:1.5, 1:1:2, 1:1:2.2, 1:1:2.4, 1:1:2.6, 1:1:2.8, 1:1:3, 1:1:3.2, 1:1:3.4, 1:1:3.6, 1:1:3.8, 1:1:4, 1:1:4.2, 1:1:4.4, 1:1:4.6, 1:1:4.8, 1:1:5, 1:1:5.5, 1:1:6, 1:1:6.5, 1:1:7, 1:1:7.5, 1:1:8, 1:1:8.5, 1:1:9, 1:1:9.5, 1:1:10, 1:1:10.5, 1:1:11, 1:1:11.5, 1:1:12, 1:1:12.5, 1:1:13, 1:1:13.5, 1:1:14, 1:1:14.5, 1:1:15, 1:1:15.5, 1:1:16, 1:1:16.5, 1:1:17, 1:1:17.5, 1:1:18, 1:1:18.5, 1:1:19, 1:1:19.5 or 1:1:20.
- the ratio of (a):(b):(c) is comprised between 1:1:2 and 1:1:4, suitably is about 1:1:2, about 1:1:3 or about 1:1:4, suitably is 1:1:2, 1:1:3 or 1:1:4.
- the ratio of (a):(b):(c) is comprised between 1:1:5.5 and 1:1:20, suitably between 1:1:6 and 1:1:12, suitably between 1:1:6 and 1:1:10, suitably is about 1:1:6, about 1:1:8 or about 1:1:10, suitably is 1:1:6, 1:1:8 or 1:1:10.
- said first subtype of Influenza A virus of (a) is H1
- said second subtype of Influenza A virus of (b) is H3
- the ratio of (a):(b):(c) is comprised between 1:1.5:1 and 1:20:1, optionally between 1:1.5:1 and 1:5:1.
- said first subtype of Influenza A virus of (a) is H1
- said second subtype of Influenza A virus of (b) is H3
- the ratio of (a):(b):(c) is comprised between 1:1.5:1 and 1:5:1, optionally between 1:2:1 and 1:5:1, optionally between 1:3:1 and 1:5:1, optionally between 1:4:1 and 1:5:1, optionally between 1:1.5:1 and 1:4:1, optionally between 1:1.5:1 and 1:3:1, optionally between 1:2:1 and 1:4:1, optionally between 1:2:1 and 1:3:1.
- said first subtype of Influenza A virus of (a) is H1
- said second subtype of Influenza A virus of (b) is H3 and the ratio of (a):(b):(c) is about 1:1.5:1, about 1:2:1, about 1:2.2:1, about 1:2.4:1, about 1:2.6:1, about 1:2.8:1, about 1:3:1, about 1:3.2:1, about 1 :3.4:1 , about 1:3.6:1, about 1 :3.8:1 , about 1:4:1, about 1 :4.2:1 , about 1 :4.4:1 , about 1 :4.6:1 , about 1:4.8:1 or about 1:5:1.
- the ratio of (a):(b):(c) is 1:1.5:1, 1:2:1, 1:2.2:1, 1 :2.4:1, 1 :2.6:1, 1:2.8:1, 1:3:1, 1 :3.2:1, 1 :3.4:1, 1 :3.6:1, 1 :3.8:1, 1:4:1, 1 :4.2:1, 1 :4.4:1, 1:4.6:1, 1:4.8:1 or 1:5:1.
- said first subtype of Influenza A virus of (a) is H1
- said second subtype of Influenza A virus of (b) is H3
- the ratio of (a):(b):(c) is comprised between 1:2:1 and 1:4:1, suitably between 1:2:1 and 1:3:1, suitably is 1:2:1 or 1:3:1.
- said first subtype of Influenza A virus of (a) is H1
- said second subtype of Influenza A virus of (b) is H3
- said ratio of (a):(b):(c) is greater than 1:3:1 and lower or equal to 1 :20: 1.
- said first subtype of Influenza A virus of (a) is H1
- said second subtype of Influenza A virus of (b) is H3
- said ratio of (a):(b):(c) is greater than 1:3:1 and lower or equal to 1:5:1.
- said first subtype of Influenza A virus of (a) is H1
- said second subtype of Influenza A virus of (b) is H3
- the ratio of (a):(b):(c) is comprised between 1:1.5: 1.5 and 1:20:5, optionally between 1:1.5: 1.5 and 1:5:5, optionally between 1:2:2 and 1:5:5, optionally between 1:3:3 and 1:5:5, optionally between 1:4:4 and 1:5:5, optionally between 1:1.5: 1.5 and 1:4:4, optionally between 1:1.5: 1.5 and 1:3:3, optionally between 1:2:2 and 1:4:4, optionally between 1:2:2 and 1:3:3, suitably is 1:2:2, 1:2:3, 1:3:2 or 1:3:3.
- said first subtype of Influenza A virus of (a) is H1
- said second subtype of Influenza A virus of (b) is H3
- the ratio of (a):(b):(c) is comprised between 1 : 1.5:5.5 and 1 :20:20, optionally between 1 : 1.5:5.5 and 1 :5:20, optionally between 1 :2:5.5 and 1 :5:20, optionally between 1 :3:5.5 and 1 :5:20, optionally between 1 :4:5.5 and 1 :5:20, optionally between 1:1.5:5.5 and 1:4:20, optionally between 1:1.5:5.5 and 1:3:20, optionally between 1:2:5.5 and 1:4:20, optionally between 1:2:5.5 and 1:3:20, suitably is 1:2:6, 1:2:8, 1:2:10, 1:3:6, 1:3:8 or 1:3:10.
- the ratio is a weight/weight ratio or a molar ratio.
- the ratio is a weight/weight ratio.
- the ratio of (a):(b):(c):(d) is comprised between 1:1:1.5:1.5 and 1:1:20:20.
- the ratio of (a):(b):(c):(d) is comprised between 1:1:1.5:1.5 and 1:1 :5:5.
- the ratio of (a):(b):(c):(d) is comprised between 1:1:1.5:1.5 and 1:1:5:5, optionally between 1:1:2:2 and 1:1:5:5, optionally between 1:1:3:3 and 1:1:5:5, optionally between 1:1:4:4 and 1 :1 :5:5, optionally between 1:1:1.5: 1.5 and 1 :1 :4:4, optionally between 1:1: 1.5: 1.5 and 1 :1 :3:3, optionally between 1 : 1:2:2 and 1:1 :4:4, optionally between 1:1 :2:2 and 1:1 :3:3.
- the ratio of (a):(b):(c) :(d) is greater than 1 : 1 :5:5 and lower or equal than 1:1:20:20, optionally greater than 1 : 1 :5:5 and lower or equal to 1:1:15:15, optionally greater than 1 : 1 :5:5 and lower or equal to 1:1:12:12, optionally greater than 1 : 1 :5:5 and lower or equal to 1:1:10:10, optionally greater than 1 : 1 :5:5 and lower or equal to 1 : 1 :8:8.
- the ratio of (a):(b):(c) :(d) is comprised between 1:1:5.5:5.5 and 1:1:20:20, optionally between 1:1:5.5:5.5 and 1:1:15:15, optionally between 1:1:5.5:5.5 and 1:1:12:12, optionally between 1:1:5.5:5.5 and 1:1:10:10, optionally between 1:1:5.5:5.5 and 1:1 :8:8, optionally between 1 :1 :6:6 and 1:1:20:20, optionally between 1:1:6:6 and 1:1:15:15, optionally between 1 : 1 :6:6 and 1:1:12:12, optionally between 1 : 1 :6:6 and 1:1:10:10, optionally between 1:1:6:6 and 1:1:8:8, optionally between 1:1:8:8 and 1:1:20:20, optionally between 1:1 :8:8 and 1:1:15:15, optionally between 1 :1:8:8 and 1:1:12:12, optionally between 1:1:8:10:10.
- the ratio of (a):(b):(c):(d) is selected from about 1:1:1.5:1.5, about 1 :1 :2:2, about 1:1:2.2:2.2, about 1:1 :2.4:2.4, about 1:1 :2.6:2.6, about 1:1:2.8:2.8, about 1 : 1 :3:3, about 1 : 1 :3.2:3.2, about 1 : 1 :3.4:3.4, about 1 : 1 :3.6:3.6, about 1 :1 :3.8:3.8, about 1 :1 :4:4, about 1:1:4.2:4.2, about 1:1:4.4:4.4, about 1:1:4.6:4.6, about 1:1:4.8:4.8, about 1 :1 :5:5, about 1:1:5.5:5.5, about 1 :1:6:6, about 1:1:6.5:6.5, about 1 :1:7:7, about 1:1:7.5:7.5, about 1 :1:8:8, about 1:1:8.5:8.5, about 1 :1 :9:9, about 1:1:9.5:9.5, about 1:1:10:10
- the ratio of (a):(b):(c):(d) is 1:1:1.5:1.5, 1:1:2:2, 1:1:2.2:2.2, 1:1:2.4:2.4, 1:1:2.6:2.6, 1:1:2.8:2.8, 1:1:3:3, 1:1:3.2:3.2, 1:1:3.4:3.4, 1:1:3.6:3.6, 1:1:3.8:3.8, 1:1:4:4, 1:1:4.2:4.2, 1:1:4.4:4.4, 1:1:4.6:4.6, 1:1:4.8:4.8, 1:1:5:5, 1:1:5.5:5.5, 1:1:6:6, 1:1:6.5:6.5, 1:1:7:7, 1:1:7.5:7.5, 1:1:8:8, 1:1:8.5:8.5, 1:1:9:9, 1:1:9.5:9.5, 1:1:10:10, 1:1:10.5:10.5, 1:1:11:11, 1:1:11.5:11.5, 1:1:12:12, 1:1:12.5:12.5, 1:1:13:13, 1:1:13.5:13.5, 1:1:14:14, 1:1:14.5:14.5, 1:1:15:15, 1:1:15.5:15.5,
- the ratio of (a):(b):(c):(d) is comprised between 1 : 1:2:2 and 1 :1 :4:4, suitably between 1 :1 :2:2 and 1 :1 :3:3, suitably is 1 :1 :2:2 or 1 :1 :3:3. In some embodiments, the ratio of (a):(b):(c):(d) is comprised between 1:1:2:2 and 1 :1 :4:4, suitably is about 1 :1 :4:4, suitably is 1 :1 :4:4.
- the ratio of (a):(b):(c):(d) is comprised between 1:1:5.5:5.5 and 1:1:20:20, suitably between 1 :1 :6:6 and 1:1:12:12, suitably between 1 :1:6:6 and 1:1:10:10, suitably is 1 :1 :6:6 or 1 :1 :8:8 or 1:1:10:10.
- said first subtype of Influenza A virus of (a) is H1
- said second subtype of Influenza A virus of (b) is H3
- the ratio of (a):(b):(c):(d) is comprised between 1:1.5:1:1 and 1:20:1:1, optionally between 1:1.5:1:1 and 1 :5:1 :1.
- said first subtype of Influenza A virus of (a) is H1
- said second subtype of Influenza A virus of (b) is H3 and the ratio of (a):(b):(c):(d) is comprised between 1:1.5:1:1 and 1:20:1:1, optionally between 1:1.5: 1:1 and 1:5:1 :1, optionally between 1:2:1 :1 and 1:5: 1:1, optionally between 1:3:1 :1 and 1:5:1 :1, optionally between 1:4:1 :1 and 1:5:1 :1, optionally between 1:1.5:1:1 and 1:4:1 :1, optionally between 1:1.5:1:1 and 1:3:1 :1, optionally between 1:2:1 :1 and 1:4:1 :1, optionally between 1:2:1 :1 and 1:3: 1:1.
- said first subtype of Influenza A virus of (a) is H1
- said second subtype of Influenza A virus of (b) is H3 and the ratio of (a):(b):(c):(d) is about 1:1.5:1:1, about 1 :2:1 :1 , about 1:2.2:1:1, about 1:2.4:1:1, about 1 :2.6:1 :1 , about 1:2.8:1:1, about 1 :3:1 :1 , about 1:3.2:1:1, about 1:3.4:1:1, about 1:3.6:1:1, about 1:3.8:1:1, about 1:4:1:1, about 1:4.2:1:1, about 1:4.4:1:1, about 1:4.6:1:1, about 1:4.8:1:1 or about 1 :5:1 :1.
- the ratio of (a):(b):(c):(d) is 1:1.5:1:1, 1:2:1:1, 1:2.2:1:1, 1:2.4:1:1, 1:2.6:1:1, 1:2.8:1:1, 1:3:1:1, 1:3.2:1:1, 1:3.4:1:1, 1:3.6:1:1, 1:3.8:1:1, 1:4:1:1, 1:4.2:1:1, 1:4.4:1:1, 1:4.6:1:1, 1:4.8:1:1 or 1:5:1 :1.
- said first subtype of Influenza A virus of (a) is H1
- said second subtype of Influenza A virus of (b) is H3
- the ratio of (a):(b):(c):(d) is comprised between 1:2:1 :1 and 1:4:1 :1, suitably between 1:2: 1:1 and 1:3: 1:1, suitably is 1:2:1 :1 or 1:3: 1:1.
- said first subtype of Influenza A virus of (a) is H1
- said second subtype of Influenza A virus of (b) is H3
- said ratio of (a):(b):(c):(d) is greater than 1:3:1 :1 and lower or equal to 1 :20: 1:1.
- said first subtype of Influenza A virus of (a) is H1
- said second subtype of Influenza A virus of (b) is H3
- said ratio of (a):(b):(c):(d) is greater than 1:3:1 :1 and lower or equal to 1:5: 1:1.
- said first subtype of Influenza A virus of (a) is H1
- said second subtype of Influenza A virus of (b) is H3 and the ratio of (a):(b):(c):(d) is comprised between 1 :1.5: 1.5: 1.5 and 1 :20:5:5, optionally between 1 :1.5: 1.5: 1.5 and 1 :5:5:5, optionally between 1 :2:2:2 and 1 :5:5:5, optionally between 1 :3:3:3 and 1 :5:5:5, optionally between 1 :4:4:4 and 1 :5:5:5, optionally between 1 :1.5:1.5:1.5 and 1 :4:4:4, optionally between 1 :1.5:1.5:1.5 and 1 :3:3:3, optionally between 1 :2:2:2 and 1 :4:4:4, optionally between 1 :2:2:2 and 1 :3:3:3, suitably is 1 :2:2:2, 1 :2:3:3,
- said first subtype of Influenza A virus of (a) is H1
- said second subtype of Influenza A virus of (b) is H3
- the ratio of (a):(b):(c):(d) is comprised between 1 :1.5:5.5:5.5 and 1 :20:20:20, optionally between 1 :1.5:5.5:5.5 and 1 :5:20:20, optionally between 1 :2:5.5:5.5 and 1 :5:20:20, optionally between 1 :3:5.5:5.5 and 1 :5:20:20, optionally between 1 :4:5.5:5.5 and 1 :5:20:20, optionally between 1 :1.5:5.5:5.5 and 1 :4:20:20, optionally between 1 :1.5:5.5:5.5 and 1 :3:20:20, optionally between 1 :2:5.5:5.5 and 1 :4:20:20, optionally between 1 :2:5.5:5.5 and 1 :3:20:20, suitably is 1 :2
- the ratio is a weight/weight ratio or a molar ratio.
- the ratio is a weight/weight ratio.
- the immunogenic composition further comprises:
- At least one further nucleic acid encoding at least one further antigen, wherein said at least one further antigen is derived from a strain of Influenza virus.
- said strain of Influenza virus from which said at least one further antigen (e) is derived is selected from the group consisting of Influenza A virus and Influenza B virus.
- said strain of Influenza virus from which said at least one further antigen (e) is derived is selected from the group consisting of said first subtype of Influenza A virus, said second subtype of Influenza A virus, said first strain of Influenza B virus and said second strain of Influenza B virus.
- the immunogenic composition is a trivalent composition.
- the immunogenic composition is a quadrivalent composition.
- said at least one further antigen comprises or consists of a peptide or protein selected or derived from an Influenza virus hemagglutinin (HA), neuraminidase (NA), nucleoprotein (NP), matrix protein 1 (M1), matrix protein 2 (M2), non- structural protein 1 (NS1), non-structural protein 2 (NS2), nuclear export protein (NEP), polymerase acidic protein (PA), polymerase basic protein PB1 , PB1-F2, and/or polymerase basic protein 2 (PB2), or an immunogenic fragment or an immunogenic variant thereof.
- said at least one further antigen comprises or consists of a peptide or protein selected or derived from an Influenza virus NA or an immunogenic fragment or an immunogenic variant thereof.
- the immunogenic composition comprises a combination of HA and NA nucleic acids encoding said HA and NA antigens, said at least one further antigen comprising or consisting of a peptide or protein selected or derived from an Influenza virus NA or fragment or variant thereof.
- NA neuraminidase
- NAI Naturally acquired or vaccine-induced NA-inhibiting antibodies have been shown to contribute to influenza disease protection in naturally occurring Influenza or in experimental human challenge studies . NAI antibodies appear to have an independent role in vaccine efficacy/effectiveness as compared to Hemagglutinin inhibition antibodies .
- said NA antigen is a polypeptide comprising a full-length Influenza NA protein.
- said NA antigen is a polypeptide consisting of a full-length Influenza NA protein.
- said NA antigen is a fragment of a neuraminidase protein, such as a truncated neuraminidase protein.
- the composition does not comprise a nucleic acid encoding a NA antigen from an influenza strain which is not recommended by WHO.
- the composition does not comprise a nucleic acid encoding a NA antigen identified or designed by machine learning.
- the nucleic acids suitably mRNAs, encoding said HA and NA antigens are present in equimolar proportions.
- the nucleic acids suitably mRNAs, encoding said HA and NA antigens are not present in equimolar proportions.
- the dose (e.g. weight dose or molar dose, suitably weight dose) of said at least one further nucleic acid, suitably mRNA, encoding said NA antigen is different compared to the dose (e.g. weight dose or molar dose, suitably weight dose) of said nucleic acids, suitably mRNAs, encoding said HA antigens.
- the ratio of HA:NA antigens encoded by nucleic acids, suitably mRNAs is comprised between 4:1 and 1 :4, suitably, 3:1 and 1 :3, suitably 2:1 and 2:1. In some embodiment, the ratio of HA:NA antigens encoded by nucleic acids, suitably mRNAs, is 4:1 or 1 :4.
- the ratio of HA:NA antigens encoded by nucleic acids is 3:1 or 1 :3.
- the ratio of HA:NA antigens encoded by nucleic acids is 2:1 or 1 :2.
- the ratio of HA:NA antigens encoded by nucleic acids is 3:2 or 2:3.
- the ratio of HA:NA antigens encoded by nucleic acids is 4:3 or 3:4.
- the ratio of HA:NA antigens encoded by nucleic acids is about 1 :1.
- the ratio of HA:NA antigens encoded by nucleic acids is 1 :1.
- the ratio is a weight/weight ratio or a molar ratio.
- the ratio is a weight/weight ratio.
- said at least one further antigen comprises or consists of an amino acid sequence having at least 90%, 95%, 98% or 99% identity to the amino acid sequence set forth in any one of SEQ ID NO: 2, 4, 6, 8, 10, 12, 14, 16, 18, 20, 22, 24, 26, 28, 30, 32, 34, 36, 38, 40, 42, 44, 46 or 48 or fragment or variant thereof.
- said at least one further antigen comprises or consists of the amino acid sequence set forth in any one of SEQ ID NO: SEQ ID NO: 2, 4, 6, 8, 10, 12, 14, 16, 18, 20, 22, 24, 26, 28, 30, 32, 34, 36, 38, 40, 42, 44, 46 or 48 or fragment or variant thereof.
- said at least one further antigen comprises or consists of an amino acid sequence having at least 90%, 95%, 98% or 99% identity to the amino acid sequence set forth in any one of SEQ I D NO: 6, 8, 18, 36, or fragment or variant thereof.
- said at least one further antigen comprises or consists of the amino acid sequence set forth in any one of SEQ ID NO: SEQ ID NO: 6, 8, 18, 36, or fragment or variant thereof.
- said at least one further antigen comprises or consists of an amino acid sequence having at least 90%, 95%, 98% or 99% identity to the amino acid sequence set forth in any one of SEQ ID NO: 2, 4, 10, 12, 14, 16, 20, 22, 24, 26, 28, 30, 32, 34, 38, 40, 42, 44, 46 or 48 or fragment or variant thereof.
- said at least one further antigen comprises or consists of the amino acid sequence set forth in any one of SEQ ID NO: SEQ ID NO: 2, 4, 10, 12, 14, 16, 20, 22, 24, 26, 28, 30, 32, 34, 38, 40, 42, 44, 46 or 48 or fragment or variant thereof.
- said at least one further antigen comprises or consists of an amino acid sequence having at least 90%, 95%, 98% or 99% identity to the amino acid sequence set forth in any one of SEQ ID NO: 2, 12, 20, 24, 28, 30, 40, 42 or 44, or fragment or variant thereof.
- said at least one further antigen comprises or consists of the amino acid sequence set forth in any one of SEQ ID NO: 2, 12, 20, 24, 28, 30, 40, 42 or 44, or fragment or variant thereof.
- said at least one further antigen comprises or consists of an amino acid sequence having at least 90%, 95%, 98% or 99% identity to the amino acid sequence set forth in any one of SEQ ID NO: 4, 10, 14, 16, 22, 26, 32, 34, 38, 46 or 48 or fragment or variant thereof.
- said at least one further antigen comprises or consists of the amino acid sequence set forth in any one of SEQ ID NO: SEQ ID NO: 4, 10, 14, 16, 22, 26, 32, 34, 38, 46 or 48 or fragment or variant thereof.
- the immunogenic composition comprises a plurality of (e), such as (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ) as defined herein.
- the composition comprises at least three, four, five, six, seven or eight nucleic acids, suitably mRNAs, encoding at least three, four, five, six, seven or eight antigens, optionally three to eight nucleic acids, suitably mRNAs, encoding three to eight antigens, optionally five to ten nucleic acids, suitably mRNAs, encoding five to ten antigens, optionally seven or eight nucleic acids, suitably mRNAs, encoding seven or eight antigens.
- the composition is a multivalent composition, wherein said antigens of (a), (b), (c), (d) and/or (e) are derived from at least three strains of Influenza virus.
- the composition comprises three nucleic acids, suitably mRNAs, encoding three antigens.
- the composition comprises six nucleic acids, suitably mRNAs, encoding six antigens.
- the immunogenic composition comprises a combination of said first, second and third nucleic acids, suitably mRNAs, encoding said three HA antigens, and three nucleic acids, suitably mRNAs, encoding three NA antigens.
- the composition is a multivalent composition, wherein said antigens of (a), (b), (c), (d) and/or (e) are derived from at least four strains of Influenza virus.
- the composition comprises seven nucleic acids, suitably mRNAs, encoding seven antigens.
- the immunogenic composition comprises a combination of said first, second, third and fourth nucleic acids, suitably mRNAs, encoding said four HA antigens, and three nucleic acids, suitably mRNAs, encoding three NA antigens.
- the immunogenic composition further comprises:
- the ratio of (a):(b):(c):(e 1 ):(e 2 ):(e 3 ) is comprised between 3:3:9: 1 : 1 : 1 and 1 : 1 :3:3:3, suitably between 2:2:6: 1 :1 : 1 and 1 :1 :3:2:2:2, suitably is 2:2:6: 1 : 1 :1 or 1:1:3:2:2:2 or 3:3:6:1:1:1 or 1:1:3:3:3:3.
- the ratio of (a):(b):(c):(e 1 ):(e 2 ):(e 3 ) is comprised between 3:3:24:1:1:1 and 1:1:8:3:3:3, suitably between 2:2:16:1:1:1 and 1:1:8:2:2:2, suitably is 2:2:16:1:1 or 1:1:8:2:2 or 3:3:24:1:1:1 or 1:1:8:3:3:3.
- said first subtype of Influenza A virus of (a) is H1
- said second subtype of Influenza A virus of (b) is H3
- the ratio of (a):(b):(c):(e 1 ):(e 2 ):(e 3 ) is comprised between 3:9:3:1:1:1 and 1 :3:1 :3:3:3, suitably between 2:6:2:1:1 and 1:3:1 :2:2:2, suitably is 2:6:2:1:1:1 or 1:3:1:2:2 or 3:9:3:1:1:1 or 1:3:1:3:3:3.
- said first subtype of Influenza A virus of (a) is H1
- said second subtype of Influenza A virus of (b) is H3
- the ratio of (a):(b):(c):(e 1 ):(e 2 ):(e 3 ) is comprised between 3:15:3:1:1 and 1:5:1:3:3:3, suitably is 2:10:2:1:1 or 3:15:3:1:1:1 or 2:8:2:1:1:1 or 3:12:3:1:1:1.
- said first subtype of Influenza A virus of (a) is H1
- said second subtype of Influenza A virus of (b) is H3
- the ratio of (a):(b):(c):(e 1 ):(e 2 ):(e 3 ) is comprised between 3:9:9:1:1:1 and 1:3:3:3:3, suitably between 2:6:6:1:1 and 1:3:3:2:2:2, suitably is 2:6:6:1:1:1 or 1:3:3:2:2 or 3:9:9:1:1:1 or 1:3:3:3:3:3.
- said first subtype of Influenza A virus of (a) is H1
- said second subtype of Influenza A virus of (b) is H3
- the ratio of (a):(b):(c):(e 1 ):(e 2 ):(e 3 ) is comprised between 3:15:15:1:1:1 and 1:5:5:3:3:3, suitably is 2:10:10:1:1:1 or 3:15:15:1:1:1 or2:8:8:1:1:1 or 3:12:12:1:1:1.
- said first subtype of Influenza A virus of (a) is H1
- said second subtype of Influenza A virus of (b) is H3
- the ratio of (a):(b):(c):(e 1 ):(e 2 ):(e 3 ) is comprised between 3:9:24:1:1:1 and 1:3:8:3:3:3, suitably between 2:6:16:1:1:1 and 1:3:8:2:2:2, suitably is 2:6:16:1:1:1 or 1:3:8:2:2 or 3:9:24:1:1:1 or 1:3:8:3:3:3.
- the ratio of (a):(b):(c):(d):(e 1 ):(e 2 ):(e 3 ) is comprised between 3:3:9:9: 1 : 1 :1 and 1:1:3:3:3:3, suitably between 2:2:6:6:1:1 and 1:1:3:3:2:2:2, suitably is 2:2:6:6:1:1 or 1:1:3:3:2:2 or 3:3:9:9: 1 : 1 : 1 or 1:1:3:3:3:3:3.
- the ratio of (a):(b):(c):(d):(e 1 ):(e 2 ):(e 3 ) is comprised between 3:3:24:24:1:1:1 and 1:1:8:8:3:3:3, suitably between 2:2:16:16:1:1 and 1:1:8:8:2:2:2, suitably is 2:2:16::161:1 or 1:1:8:8:2:2 or 3:3:24:24:1:1:1 or 1:1:8:8:3:3:3.
- said first subtype of Influenza A virus of (a) is H1
- said second subtype of Influenza A virus of (b) is H3
- the ratio of (a):(b):(c):(d):(e 1 ):(e 2 ):(e 3 ) is comprised between 3:9:3:3: 1 :1 : 1 and 1 :3: 1 :1 :3:3:3, suitably between 2:6:2:2:1:1:1 and 1 :3: 1 : 1 :2:2:2, suitably is 2:6:2:2:1:1:1 or 1 :3: 1 :1 :2:2:2 or 3:9:3:3: 1 : 1 : 1 or 1 :3: 1 :1 :3:3:3.
- said first subtype of Influenza A virus of (a) is H1
- said second subtype of Influenza A virus of (b) is H3
- the ratio of (a):(b):(c):(d):(e 1 ):(e 2 ):(e 3 ) is comprised between 3:15:3:3:1:1 and 1 :5:1 : 1 :3:3:3, suitably is 2:10:2:2:1:1:1 or 3:15:3:3:1:1:1 or 2:8:2:2: 1 : 1 : 1 or 3:12:3:3:1:1:1.
- said first subtype of Influenza A virus of (a) is H1
- said second subtype of Influenza A virus of (b) is H3
- the ratio of (a):(b):(c):(d):(e 1 ):(e 2 ):(e 3 ) is comprised between 3:9:9:9: 1 :1 : 1 and 1:3:3:3:3:3, suitably between 2:6:6:6:1:1:1 and 1:3:3:3:2:2:2, suitably is 2:6:6:6:1:1:1 or 1:3:3:3:2:2 or 3:9:9:9: 1 : 1 : 1 or 1:3:3:3:3:3:3.
- said first subtype of Influenza A virus of (a) is H1
- said second subtype of Influenza A virus of (b) is H3
- the ratio of (a):(b):(c):(d):(e 1 ):(e 2 ):(e 3 ) is comprised between 3:15:15:15:1:1:1 and 1:5:5:5:3:3:3, suitably is 2:10:10:10:1:1 or 3:15:15:15:15:1:1:1 or 2:8:8:8:1:1:1 or 3:12:12:12:1:1:1.
- said first subtype of Influenza A virus of (a) is H1
- said second subtype of Influenza A virus of (b) is H3
- the ratio of (a):(b):(c):(d):(e 1 ):(e 2 ):(e 3 ) is comprised between 3:9:24:24:1:1:1 and 1:3:8:8:3:3:3, suitably between 2:6:16:16:1:1:1 and 1:3:8:8:2:2:2, suitably is 2:6:16:16:1:1:1 or 1:3:8:8:2:2 or 3:9:24:24:1:1:1 or 1:3:8:8:3:3:3.
- the composition comprises eight nucleic acids, suitably mRNAs, encoding eight antigens.
- the immunogenic composition comprises a combination of four HA antigens or four nucleic acids, suitably mRNAs, encoding said four HA antigens, and four NA antigens or four nucleic acids, suitably mRNAs, encoding said four NA antigens.
- composition further comprises:
- the ratio of (a):(b):(c):(d):(e 1 ):(e 2 ):(e 3 ):(e 4 ) is comprised between 3:3:9:9: 1 : 1 :1 :1 and 1:1:3:3:3:3:3, suitably between 2:2:6:6: 1 : 1 : 1 : 1 and 1:1:3:3:2:2:2, suitably is 2:2:6:6: 1 : 1 : 1 or 1:1:3:3:2:2:2 or 3:3:9:9: 1 :1 : 1 : 1 or 1:1:3:3:3:3:3:3.
- the ratio of (a):(b):(c):(d):(e 1 ):(e 2 ):(e 3 ):(e 4 ) is comprised between 3:3:24:24:1:1:1:1 and 1:1:8:8:3:3:3, suitably between 2:2:16:16:1:1:1 and 1:1:8:8:2:2:2, suitably is 2:2:16:16:1:1:1:1 or 1:1:8:8:2:2:2 or 3:3:24:24:1:1:1:1 or 1:1:8:8:3:3:3.
- said first subtype of Influenza A virus of (a) is H1
- said second subtype of Influenza A virus of (b) is H3
- the ratio of (a):(b):(c):(d):(e 1 ):(e 2 ):(e 3 ):(e 4 ) is comprised between 3:9:3:3: 1 :1 : 1 : 1 and 1 :3: 1 : 1 :3:3:3, suitably between 2:6:2:2: 1 : 1 : 1 : 1 and 1:3:1 :1:2:2:2, suitably is 2:6:2:2: 1 : 1 : 1 : 1 or 1:3:1:1:2:2:2 or3:9:3:3:1:1:1:1 or 1:3:1 :3:3:3:3.
- said first subtype of Influenza A virus of (a) is H1
- said second subtype of Influenza A virus of (b) is H3
- the ratio of (a):(b):(c):(d):(e 1 ):(e 2 ):(e 3 ):(e 4 ) is comprised between 3:15:3:3:1:1:1 and 1 :5:1 : 1 :3:3:3, suitably is 2:10:2:2:1:1:1:1 or 3:15:3:3:1:1:1:1 or2:8:2:2:1:1:1:1 or 3:12:3:3:1:1:1:1.
- said first subtype of Influenza A virus of (a) is H1
- said second subtype of Influenza A virus of (b) is H3
- the ratio of (a):(b):(c):(d):(e 1 ):(e 2 ):(e 3 ):(e 4 ) is comprised between 3:9:9:9: 1 :1 : 1 : 1 and 1:3:3:3:3:3, suitably between 2:6:6:6: 1 : 1 : 1 : 1 and 1:3:3:3:2:2:2, suitably is 2:6:6:6: 1 : 1 : 1 : 1 or 1:3:3:3:2:2:2 or3:9:9:9:1:1:1:1 or 1:3:3:3:3:3:3:1.
- said first subtype of Influenza A virus of (a) is H1
- said second subtype of Influenza A virus of (b) is H3
- the ratio of (a):(b):(c):(d):(e 1 ):(e 2 ):(e 3 ):(e 4 ) is comprised between 3:15:15:15:1:1:1 and 1:5:5:5:3:3:3, suitably is 2:10:10:10:1:1:1 or 3:15:15:15:15:1:1:1:1 or2:8:8:8:1:1:1:1 or 3:12:12:12:1:1:1:1.
- said first subtype of Influenza A virus of (a) is H1
- said second subtype of Influenza A virus of (b) is H3
- the ratio of (a):(b):(c):(d):(e 1 ):(e 2 ):(e 3 ):(e 4 ) is comprised between 3:9:24:24:1 :1 :1 :1 and 1 :3:8:8:3:3:3:3, suitably between 2:6:16:16:1 :1 :1 :1 and 1 :3:8:8:2:2:2:2, suitably is 2:6:16:16:1 :1 :1 :1 or 1 :3:8:8:2:2:2 or 3:9:24:24:1 :1 :1 :1 or 1 :3:8:8:3:3:3.
- the ratio is a weight/weight ratio or a molar ratio.
- the ratio is a weight/weight ratio.
- an immune response is further elicited against NA antigens of said first and second subtypes of Influenza A virus and said first strain of Influenza B virus, and optionally, at least one further NA antigen of a strain of Influenza A virus and/or Influenza B virus, being different from any of the NA antigen encoded by a nucleic acid present in the composition.
- an immune response is further elicited against NA antigens of said first and second subtypes of Influenza A virus, said first and second strains of Influenza B virus, and optionally, at least one further NA antigen of a strain of Influenza A virus and/or Influenza B virus, being different from any of the NA antigen encoded by a nucleic acid present in the composition.
- an immune response is elicited against NA antigens that are antigenically distinct from any of the NA antigen encoded by a nucleic acid present in the compositions.
- At least one nucleic acid of the immunogenic composition is DNA or RNA, suitably mRNA.
- as least one nucleic acid of the immunogenic composition is a DNA.
- At least one nucleic acid of the immunogenic composition is an artificial nucleic acid, e.g. an artificial DNA or an artificial RNA, suitably mRNA.
- Nucleic acid-based vaccination including DNA or RNA, suitably mRNA, represents a promising technique for novel vaccines against emerging viruses and for the provision of combination vaccines.
- Nucleic acids can be genetically engineered and administered to a human subject.
- Transfected cells directly produce the encoded antigen (e.g. provided by a DNA or an RNA, in particular an mRNA), which results in protective immunological responses.
- Nucleic acids according to the invention e.g. DNAs or RNAs, suitably mRNAs, form the basis for a nucleic acid based immunogenic composition or a nucleic acid based vaccine.
- nucleic acid based immunogenic compositions first aspect
- nucleic acid-based vaccines second aspect
- protein-based vaccines, or live attenuated vaccines are suboptimal for use in developing countries due to their high production costs.
- protein-based vaccines, or live attenuated vaccines require long development times and are not suitable for rapid responses of epidemic virus outbreaks such as e.g. the Influenza virus outbreaks.
- the GISRS recommendation is made six to seven months prior the start of the Influenza season, during which the Influenza viruses may continue to evolve.
- nucleic acid-based immunogenic compositions and vaccines according to the invention allow very fast and cost-effective manufacturing. Therefore, in comparison with known vaccines, compositions/vaccines based on nucleic acids can be produced and manufactured significantly cheaper and faster, which is very advantageous particularly for use in developing countries or in the context of annual epidemics or a global pandemic.
- the nucleic acid-based compositions/vaccines offer the GISRS additional time to monitor circulating viruses and make its recommendation closer to the Influenza season. This extension of the GISRS monitoring timeline should allow the GISRS predictions to be more accurate, resulting in more effective vaccines designated to target circulating viruses closer to Influenza season.
- the different nucleic acid encoding different antigens e.g. of different Influenza strains
- RNA molecules suitably mRNAs
- RNA molecules are considered to be significantly safer than DNA vaccines, as RNAs, suitably mRNAs, are more easily degraded. They are cleared quickly out of the organism and cannot integrate into the genome and influence the cell's gene expression in an uncontrollable manner. It is also less likely for RNA, suitably mRNA, vaccines to cause severe side effects like the generation of autoimmune disease or anti-DNA antibodies.
- Transfection with RNA, suitably mRNA requires only insertion into the cell's cytoplasm, which is easier to achieve than into the nucleus.
- At least one nucleic acid of the immunogenic composition is an RNA.
- RNA is the usual abbreviation for ribonucleic acid. It is a nucleic acid molecule, i.e. a polymer consisting of nucleotide monomers. These nucleotides are usually adenosine-monophosphate (AMP), uridine-monophosphate (UMP), guanosinemonophosphate (GMP) and cytidine-monophosphate (CMP) monomers or analogs thereof, which are connected to each other along a so-called backbone.
- the backbone is typically formed by phosphodiester bonds between the sugar, i.e. ribose, of a first and a phosphate moiety of a second, adjacent monomer.
- the specific order of the monomers i.e. the order of the bases linked to the sugar/phosphate-backbone, is called the RNA sequence.
- the RNA molecule is selected from an antisense RNA, such as an antisense oligonucleotide (ASOs), a small interfering RNA (siRNA), a microRNA (miRNAs), a messenger RNA (mRNA) and an RNA forming part of a single-guide RNA (sgRNA)-mediated CRISPR- Cas system.
- ASOs antisense oligonucleotide
- siRNA small interfering RNA
- miRNAs microRNA
- mRNA messenger RNA
- sgRNA single-guide RNA
- (a) is a first RNA
- (b) is a second RNA
- (c) is a first RNA
- (d) is a fourth RNA.
- (e) is at least one further RNA encoding the at least one further antigen.
- the immunogenic composition comprises a plurality of (e) being RNAs.
- (a), (b), (c), (d), (e), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ) is an RNA.
- RNA is a single-stranded RNA molecule that corresponds to the genetic sequence of a gene and is read by ribosomes in the process of producing a protein.
- mRNA vaccines may utilise non-replicating mRNA or self-replicating RNA (also referred to as self-amplifying mRNA or SAM).
- Non-replicating mRNA-based vaccines typically encode an antigen of interest and contain 5' and 3' untranslated regions (UTRs), a 5’ cap and a poly(A) tail; whereas self-amplifying RNAs also encode viral replication machinery that enables intracellular RNA amplification.
- At least one nucleic acid of the immunogenic composition is a mRNA.
- said first, second, third and/or fourth nucleic acid is a mRNA.
- a dose of each said first, second, third and/or fourth mRNA is 1 to 200 pg, suitably 1 to 60 pg, suitably 2 to 25 pg.
- a dose of each said first, second, third and/or fourth mRNA is 2 to 25 pg, optionally 2 to 18 pg, optionally 2 to 9 pg, optionally 2 to 6 pg, optionally 3 to 25 pg, 3 to 18 pg, 3 to 9 pg, optionally 3 to 6 pg.
- a dose of each said first, second, third and/or fourth mRNA is
- a dose of each said first, second, third and/or fourth mRNA is 3, 6, 9, 12 or 18 pg.
- a dose of each said first, second, third and/or fourth mRNA is 0.5 to 200 pg, optionally 2 to 25 pg, optionally 2 to 30 pg, optionally 2 to 40 pg, optionally 2 to 45 pg, optionally 2 to 50 pg, optionally 2 to 75 pg.
- a dose of each said first, second, third and/or fourth mRNA is 0.5, 1 , 2, 3, 4, 5, 6, 7, 8, 9, 10, 11 , 12, 13, 14, 15, 16, 17, 18, 19, 20, 21 , 22, 23, 24, 25, 30, 35, 36, 37, 38, 39, 40, 45, 46, 47, 48, 49, 50, 55, 60, 65, 70, 71 , 72, 73, 74 or 75 pg, optionally 3, 6, 9, 24, 36, 48 or 72 pg.
- a dose of each said first, second, third and/or fourth mRNA is 3, 6, 9, 24, 36, 48 or 72 pg.
- a dose of said first mRNA is comprised between 0.5 and 15 pg, optionally between 2 and 10 pg.
- a dose of said first mRNA is 0.5, 1 , 2, 3, 4, 5, 6, 7, 8, 9, 10, 11 , 12, 13, 14 or 15 pg.
- a dose of said first mRNA is 3, 6 or 9 pg.
- a dose of said second mRNA is comprised between 0.5 and 15 pg, optionally between 2 and 10 pg.
- a dose of said second mRNA is 0.5, 1 , 2, 3, 4, 5, 6, 7, 8, 9, 10,
- a dose of said second mRNA is 3, 6 or 9 pg.
- a dose of said third mRNA is comprised between 15 and 100 pg, optionally between 20 and 75 pg. In some embodiments, a dose of said third mRNA is 20, 21 , 22, 23, 24, 25, 30, 35, 36, 37, 38, 39, 40, 45, 46, 47, 48, 49, 50, 55, 60, 65, 70, 71 , 72, 73, 74 or 75 pg.
- a dose of said third mRNA is 24, 36, 48 or 72 pg.
- a dose of said fourth mRNA is comprised between 15 and 100 pg, optionally between 20 and 75 pg.
- a dose of said fourth mRNA is 20, 21 , 22, 23, 24, 25, 30, 35, 36, 37, 38, 39, 40, 45, 46, 47, 48, 49, 50, 55, 60, 65, 70, 71 , 72, 73, 74 or 75 pg.
- a dose of said fourth mRNA is 24, 36, 48 or 72 pg.
- said at least one further nucleic acid (e) is an mRNA.
- a dose of each said at least one further mRNA is 1 to 200 pg, suitably 1 to 60 pg, suitably 2 to 25 pg.
- a dose of each said at least one further mRNA is 2 to 25 pg, optionally 2 to 18 pg, optionally 2 to 9 pg, optionally 2 to 6 pg, optionally 3 to 25 pg, 3 to 18 pg, 3 to 9 pg, optionally 3 to 6 pg.
- a dose of each said at least one further mRNA is 1 , 2, 3, 4, 5, 6, 7, 8, 9, 10, 11 , 12, 13, 14, 15, 16, 17, 18, 19, 20, 21 , 22, 23, 24 or 25 pg, optionally 2, 3, 6, 9 or 18 pg.
- a dose of each said at least one further mRNA is 3, 6, 9, 12 or 18 pg.
- a dose of each said at least one further mRNA is 0.5 to 200 pg, optionally 2 to 10 pg, 2 to 25 pg, optionally 2 to 30 pg, optionally 2 to 40 pg, optionally 2 to 45 pg, optionally 2 to 50 pg, optionally 2 to 75 pg.
- a dose of each said at least one further mRNA is 1 , 2, 3, 4, 5, 6, 7, 8, 9, 10, 11 , 12, 13, 14, 15, 16, 17, 18, 19, 20, 21 , 22, 23, 24, 25, 30, 35, 36, 37, 38, 39, 40, 45, 46, 47, 48, 49, 50, 55, 60, 65, 70, 71 , 72, 73, 74or 75 pg, optionally 3, 6, 9, 24, 36, 48 or 72 pg.
- a dose of each at least one further mRNA is 3, 6, 9, 24, 36, 48 or 72 pg.
- a dose of each at least one further mRNA is comprised between 2 to 10 pg, suitably is 3 pg.
- the immunogenic composition comprises a plurality of (e) being mRNAs.
- the composition comprises at least three, four, five, six, seven or eight mRNAs, optionally three to eight mRNAs, optionally five to ten mRNAs, optionally seven or eight mRNAs.
- the third, fourth, fifth, sixth, seventh and/or eighth nucleic acid is an mRNA.
- (a), (b), (c), (d), (e), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ) is an mRNA.
- a dose of each (a), (b), (c), (d), (e), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ) is 1 to 200 pg, suitably 1 to 60 pg, suitably 1 to 25 pg, suitably 2 to 25 pg.
- a dose of each (a), (b), (c), (d), (e), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ) is 1 to 25 pg, optionally 2 to 25 pg, optionally 2 to 18 pg, optionally 2 to 9 pg, optionally 2 to 6 pg, optionally 3 to 25 pg, 3 to 18 pg, 3 to 9 pg, optionally 3 to 6 pg.
- a dose of each (a), (b), (c), (d), (e), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ) is 1 , 2, 3, 4, 5, 6, 7, 8, 9, 10, 11 , 12, 13, 14, 15, 16, 17, 18, 19, 20, 21 , 22, 23, 24 or 25 pg, optionally 1 , 2, 3, 6, 9 or 18 pg.
- a dose of each (a), (b), (c), (d), (e), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ) is 1 , 2, 3, 6, 9, 12 or 18 pg.
- a dose of each (a), (b), (c), (d), (e), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ) is 0.5 to 200 pg, optionally 2 to 10 pg, 2 to 25 pg, optionally 2 to 30 pg, optionally 2 to 40 pg, optionally 2 to 45 pg, optionally 2 to 50 pg, optionally 2 to 75 pg.
- a dose of each (a), (b), (c), (d), (e), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ) is 0.5, 1 , 2, 3, 4, 5, 6, 7, 8, 9, 10, 11 , 12, 13, 14, 15, 16, 17, 18, 19, 20, 21 , 22, 23, 24, 25, 30, 35, 36, 37, 38, 39, 40, 45, 46, 47, 48, 49, 50, 55, 60, 65, 70, 71 , 72, 73, 74 or 75 pg, optionally 3, 6, 9, 24, 36, 48 or 72 pg.
- a dose of each (a), (b), (c), (d), (e), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ) is 3, 6, 9, 24, 36, 48 or 72 pg.
- an immunogenic composition for use in the treatment or prophylaxis of an infection with an Influenza virus, wherein the immunogenic composition comprises:
- a third mRNA encoding a HA antigen of a first strain of Influenza B virus, wherein an immune response is elicited against HA antigens of said strains of first and second subtypes of Influenza A virus, said first strain of Influenza B virus and at least one further HA antigen subtype of Influenza A virus, being different from any of the HA antigen subtypes of Influenza A virus encoded by an mRNA present in the composition.
- an immunogenic composition for use in the treatment or prophylaxis of an infection with an Influenza virus, wherein the immunogenic composition comprises:
- a fourth mRNA encoding a HA antigen of a second strain of Influenza B virus, wherein an immune response is elicited against HA antigens of said strains of first and second subtypes of Influenza A virus, said first and second strains of Influenza B virus and at least one further HA antigen subtype of Influenza A virus, being different from any of the HA antigen subtypes of Influenza A virus encoded by an mRNA present in the composition.
- said first strain of Influenza B virus of (c) and said second strain of Influenza B virus of (d) are identical.
- (c) and (d) are identical.
- said first strain of Influenza B virus of (c) and said second strain of Influenza B virus of (d) are different.
- (c) and (d) are different.
- a dose of each of (c) and (d) is comprised between 15 and 100 pg, optionally between 20 and 75 pg.
- a dose of each of (c) and (d) is 20, 21 , 22, 23, 24, 25, 30, 35, 36, 37, 38, 39, 40, 45, 46, 47, 48, 49, 50, 55, 60, 65, 70, 1 , 72, 73, 74 or 75 pg.
- a dose of each of (c) and (d) is 24, 36, 48 or 72 pg. In some embodiments, a dose of (c) and (d) is 5 to 50 pg, optionally 10 to 40 pg, optionally 12 to 36 pg.
- a dose of (c) and (d) is 10, 11 , 12, 13, 14, 15, 16, 17, 18, 19, 20, 21 , 22, 23, 24, 25, 26, 27, 28, 29, 30, 31 , 32, 33, 34, 35,36, 37, 38, 39 or 40 pg.
- a dose of (c) and (d) is comprised between 20 and 200 pg, optionally between 40 and 150 pg.
- a dose of (c) and (d) is 45, 46, 47, 48, 49, 50, 60, 70, 71 , 72, 73, 74, 75, 80, 90, 95, 96, 97, 98, 99, 100, 110, 120, 130, 140, 141 , 142, 143, 144, 145 or 150 pg-
- a dose of (c) and (d) is 48, 72, 96 or 144 pg.
- a dose of each of (a) and (b) is comprised between 0.5 and 15 pg, optionally between 2 and 10 pg.
- a dose of each of (a) and (b) is 0.5, 1 , 2, 3, 4, 5, 6, 7, 8, 9, 10, 11 , 12, 13, 14 or 15 pg.
- a dose of each of (a) and (b) is 3, 6 or 9 pg.
- a dose of (a) and (b) is 2 to 20 pg, optionally 5 to 15 pg, optionally 6 to 12 pg.
- a dose of (a) and (b) is 2, 3, 4, 5, 6, 7, 8, 9, 10, 11 , 12, 13, 14, 15, 16, 17, 18, 19 or 20 pg.
- a dose of (a) and (b) is 5, 6, 7, 8, 9, 10, 11 , 12, 13, 14 or 15 pg.
- a dose of (a) and (b) is comprised between 2 and 30 pg, optionally between 5 and 20 pg.
- a dose of (a) and (b) is 5, 6, 7, 8, 9, 10, 11 , 12, 13, 14, 15, 16, 17, 18, 19 or 20 pg.
- a dose of (a) and (b) is 6, 12 or 18 pg.
- a dose of (a), (b), (c) and (d) is 5 to 75 pg, optionally 10 to 60 pg, optionally 12 to 48 pg.
- a dose of (a), (b), (c) and (d) is 35 to 75 pg.
- a dose of (a), (b), (c) and (d) is 35, 36, 37, 38, 39, 40, 45, 46, 47, 48, 49, 50, 51 , 52, 53, 54, 55, 70, 71 , 72, 73, 74 or 75 pg.
- a dose of (a), (b), (c) and (d) is 10, 11 , 12, 13, 14, 15, 16, 17, 18, 19, 20, 21 , 22, 24, 24, 25, 26, 27, 28, 29, 30, 45, 46, 47, 48, 49, 50, 55, 60 pg.
- a dose of (a), (b) and (c) is 25 to 150 pg, optionally 25 to 100 pg, optionally 30 to 90 pg.
- a dose of (a), (b) and (c) is 25 to 100 pg.
- a dose of (a), (b) and (c) is 25, 26, 27, 28, 29, 30, 31 , 32, 33, 34, 35, 36, 37, 38, 39, 40, 45, 50, 51 , 52, 53, 54, 55, 56, 57, 58, 59, 60, 65, 70, 75, 80, 85, 86, 87, 88, 89, 90, 91 , 92, 93, 94, 95 or 100 pg.
- a dose of (a), (b) and (c) is 30, 36, 54, 60 or 90 pg.
- a dose of (c) and (d) is 5 to 50 pg, optionally 10 to 40 pg, optionally 12 to 36 pg, and a dose of (a) and (b) is 2 to 20 pg, optionally 5 to 15 pg, optionally 6 to 12 pg.
- a dose of (c) and (d) is 10, 11 , 12, 13, 14, 15, 16, 17, 18, 19, 20, 21 , 22, 23, 24, 25, 26, 27, 28, 29, 30, 31 , 32, 33, 34, 35,36, 37, 38, 39 or 40 pg, and a dose of (a) and (b) is 5, 6, 7, 8, 9, 10, 11 , 12, 13, 14 or 15 pg.
- a dose of (c) is comprised between 15 and 100 pg, optionally between 20 and 75 pg, and a dose of (a) and (b) is comprised between 2 and 30 pg, optionally between 5 and 20 pg.
- a dose of (c) is 24, 36, 48 or 72 pg, and a dose of (a) and (b) is 6, 12 or 18 pg.
- the immunogenic composition further comprises:
- a dose of (e 1 ), (e 2 ) and (e 3 ) is 2 to 50 pg, optionally 2 to 30 pg, optionally 5 to 20, optionally 9 to 18 pg.
- a dose of (e 1 ), (e 2 ) and (e 3 ) is 5, 6, 7, 8, 9, 10, 11 , 12, 13, 14, 15, 16, 17, 18, 19, 20 pg.
- a dose of (e 1 ), (e 2 ) and (e 3 ) is 9 to 36 pg.
- a dose of (e 1 ), (e 2 ) and (e 3 ) is 9, 18, 27 or 36 pg. In some embodiments, a dose of (e 1 ), (e 2 ) and (e 3 ) is 0.5 to 50 pg, optionally 2 to 20 pg, optionally 5 to 15. In some embodiments, a dose of (e 1 ), (e 2 ) and (e 3 ) is 5, 6, 7, 8, 9, 10, 11 , 12, 13, 14 or 15 pg.
- a dose of (e 1 ), (e 2 ) and (e 3 ) is 5 to 15 pg.
- a dose of (e 1 ), (e 2 ) and (e 3 ) is 9 pg.
- a dose of each of (e 1 ), (e 2 ) and (e 3 ) is 0.5 to 20 pg, optionally
- a dose of each of (e 1 ), (e 2 ) and (e 3 ) is 0.5, 1 , 2, 3, 4, 5, 6, 7, 8, 9 or 10 pg.
- a dose of each of (e 1 ), (e 2 ) and (e 3 ) is 1 to 5 pg.
- a dose of each of (e 1 ), (e 2 ) and (e 3 ) is 3 pg.
- the immunogenic composition further comprises:
- a dose of (e 1 ), (e 2 ), (e 3 ) and (e 4 ) is 5 to 50 pg, optionally 10 to 50 pg, optionally 12 to 48 pg.
- a dose of (e 1 ), (e 2 ), (e 3 ) and (e 4 ) is 10, 11 , 12, 13, 14, 15, 20, 21 , 22, 23, 24, 25, 26, 27, 28, 29, 30, 35, 36, 37, 38, 39, 40, 45, 46, 47, 48 pg.
- a dose of (e 1 ), (e 2 ), (e 3 ) and (e 4 ) is comprised between 0.5 to 50 pg, optionally between 2 to 25 pg, optionally between 5 to 20 pg.
- a dose of (e 1 ), (e 2 ), (e 3 ) and (e 4 ) is 5, 6, 7, 8, 9, 10, 11 , 12, 13, 14, 15, 16, 17, 18, 19 or 20 pg.
- a dose of (e 1 ), (e 2 ), (e 3 ) and (e 4 ) is 12 pg.
- a dose of each of (e 1 ), (e 2 ), (e 3 ) and (e 4 ) is 0.5 to 20 pg, optionally 0.5 to 10 pg, optionally 0.5 to 5, optionally 1 to 5 pg, optionally 2 to 5 pg. In some embodiments, a dose of each of (e 1 ), (e 2 ) and (e 3 ) is 0.5, 1 , 2, 3, 4, 5, 6, 7, 8, 9 or 10 pg.
- a dose of each of (e 1 ), (e 2 ), (e 3 ) and (e 4 ) is 1 to 5 pg.
- a dose of each of (e 1 ), (e 2 ), (e 3 ) and (e 4 ) is 3 pg.
- a dose of each (a), (b), (c), (d), (e), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ) is 1 to 200 pg, suitably 1 to 60 pg, suitably 1 to 25 pg, suitably 2 to 25 pg.
- a dose of each (a), (b), (c), (d), (e), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ) is 1 to 25 pg, optionally 2 to 25 pg, optionally 2 to 18 pg, optionally 2 to 9 pg, optionally 2 to 6 pg, optionally 3 to 25 pg, 3 to 18 pg, optionally 3 to 12 pg, optionally 3 to 9 pg, optionally 3 to 6 pg.
- a dose of each (a), (b), (c), (d), (e), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ) is 1 , 2, 3, 4, 5, 6, 7, 8, 9, 10, 11 , 12, 13, 14, 15, 16, 17, 18, 19, 20, 21 , 22, 23, 24 or 25 pg, optionally 1 , 2, 3, 6, 9 or 18 pg.
- a dose of each (a), (b), (c), (d), (e), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ) is 1 , 2, 3, 6, 9, 12 or 18 pg.
- a dose of each (a), (b), (c), (d), (e), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ) is 0.5 to 200 pg, optionally 2 to 10 pg, 2 to 25 pg, optionally 2 to 30 pg, optionally 2 to 40 pg, optionally 2 to 45 pg, optionally 2 to 50 pg, optionally 2 to 75 pg.
- a dose of each (a), (b), (c), (d), (e), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ) is 0.5, 1 , 2, 3, 4, 5, 6, 7, 8, 9, 10, 11 , 12, 13, 14, 15, 16, 17, 18, 19, 20, 21 , 22, 23, 24, 25, 30, 35, 36, 37, 38, 39, 40, 45, 46, 47, 48, 49, 50, 55, 60, 65, 70, 71 , 72, 73, 74 or 75 pg, optionally 3, 6, 9, 24, 36, 48 or 72 pg.
- a dose of each (a), (b), (c), (d), (e), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ) is 3, 6, 9, 24, 36, 48 or 72 pg.
- mRNAs used herein are suitably provided in purified or substantially purified form i.e. substantially free from proteins (e.g., enzymes), other nucleic acids (e.g. DNA and nucleoside phosphate monomers), and the like, generally being at least about 50% pure (by weight), and usually at least 90% pure, such as at least 95% or at least 98% pure.
- mRNAs used herein may be prepared in many ways e.g.
- mRNA may be prepared enzymatically using a DNA template.
- mRNAs used herein may be an artificial nucleic acid.
- the term “artificial nucleic acid” as used herein is intended to refer to a nucleic acid that does not occur naturally. In other words, an artificial nucleic acid may be understood as a non-natural nucleic acid molecule. Such nucleic acid molecules may be non-natural due to its individual sequence (e.g.
- artificial nucleic acid may be designed and/or generated by genetic engineering to correspond to a desired artificial sequence of nucleotides.
- an artificial nucleic acid is a sequence that may not occur naturally, i.e. a sequence that differs from the wild type or reference sequence/the naturally occurring sequence by at least one nucleotide (via e.g. codon modification as further specified below).
- the term “artificial nucleic acid” is not restricted to mean “one single molecule” but is understood to comprise an ensemble of essentially identical nucleic acid molecules. Accordingly, it may relate to a plurality of essentially identical nucleic acid molecules.
- the mRNAs used herein may be a modified and/or stabilized nucleic acid, suitably a modified and/or stabilized artificial mRNA.
- the mRNAs used herein may thus be provided as a “stabilized artificial nucleic acid” or “stabilized coding nucleic acid” that is to say a nucleic acid showing improved resistance to in vivo degradation and/or a nucleic acid showing improved stability in vivo, and/or a nucleic acid showing improved translatability in vivo.
- a nucleic acid showing improved resistance to in vivo degradation and/or a nucleic acid showing improved stability in vivo, and/or a nucleic acid showing improved translatability in vivo.
- specific suitable modifications/adaptations in this context are described which are suitably to “stabilize” the nucleic acid.
- mRNAs used herein may also be codon optimized.
- the mRNAs used herein comprises at least one codon modified coding sequence.
- the coding sequence of the mRNAs used herein is a codon modified coding sequence.
- the amino acid sequence encoded by the codon modified coding sequence is not being modified compared to the amino acid sequence encoded by the corresponding wild type or reference coding sequence.
- the mRNAs of (a), (b), (c), (d), (e), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ) comprises a coding sequence which is a codon modified coding sequence, wherein the amino acid sequence encoded by the codon modified coding sequence is optionally not being modified compared to the amino acid sequence encoded by the corresponding wild type or reference coding sequence.
- mRNAs used herein may be codon optimized for expression in human cells.
- codon optimized is intended modification with respect to codon usage may increase translation efficacy and/or half-life of the nucleic acid.
- the term “codon modified coding sequence” relates to coding sequences that differ in at least one codon (triplets of nucleotides coding for one amino acid) compared to the corresponding wild type or reference coding sequence.
- a codon modified coding sequence in the context of the invention may show improved resistance to in vivo degradation and/or improved stability in vivo, and/or improved translatability in vivo.
- Codon modifications in the broadest sense make use of the degeneracy of the genetic code wherein multiple codons may encode the same amino acid and may be used interchangeably (cf. Table 1 of W02020002525) to optimize/modify the coding sequence for in vivo applications as outlined herein.
- the mRNAs used herein may be modified, wherein the C content of the at least one coding sequence may be increased, suitably maximized, compared to the C content of the corresponding wild type or reference coding sequence (herein referred to as “C maximized coding sequence”).
- the amino acid sequence encoded by the C maximized coding sequence of the mRNA is suitably not modified compared to the amino acid sequence encoded by the respective wild type or reference coding sequence.
- the generation of a C maximized nucleic acid sequences may suitably be carried out using a modification method according to WO20 15/062738. In this context, the disclosure of WO2015/062738 is included herewith by reference.
- the mRNAs used herein may be modified, wherein the codons in the at least one coding sequence may be adapted to human codon usage (herein referred to as “human codon usage adapted coding sequence”). Codons encoding the same amino acid occur at different frequencies in humans. Accordingly, the coding sequence of the mRNAs used herein is suitably modified such that the frequency of the codons encoding the same amino acid corresponds to the naturally occurring frequency of that codon according to the human codon usage.
- the wild type or reference coding sequence is suitably adapted in a way that the codon “GCC” is used with a frequency of 0.40, the codon “GCT” is used with a frequency of 0.28, the codon “GCA” is used with a frequency of 0.22 and the codon “GCG” is used with a frequency of 0.10 etc. (see e.g. Table 1 of W02020002525). Accordingly, such a procedure (as exemplified for Ala) is applied for each amino acid encoded by the coding sequence of the RNA to obtain sequences adapted to human codon usage.
- the mRNAs used herein may be modified, wherein the codon adaptation index (CAI) may be increased or suitably maximised in the at least one coding sequence (herein referred to as “CAI maximized coding sequence”).
- CAI maximized coding sequence all codons of the wild type or reference nucleic acid sequence that are relatively rare in e.g. a human are exchanged for a respective codon that is frequent in the e.g. a human, wherein the frequent codon encodes the same amino acid as the relatively rare codon.
- the most frequent codons are used for each amino acid of the encoded protein (see Table 1 of W02020002525, most frequent human codons are marked with asterisks).
- the mRNAs used herein comprise at least one coding sequence, wherein the codon adaptation index (CAI) of the at least one coding sequence is at least 0.5, at least 0.8, at least 0.9 or at least 0.95.
- the wild type or reference coding sequence may be adapted in a way that the most frequent human codon “GCC” is always used for the amino acid. Accordingly, such a procedure (as exemplified for Ala) may be applied for each amino acid encoded by the coding sequence of the mRNA to obtain CAI maximized coding sequences.
- the mRNAs used herein may be modified, wherein the G/C content of the at least one coding sequence may be modified compared to the G/C content of the corresponding wild type or reference coding sequence (herein referred to as “G/C content modified coding sequence”).
- G/C optimization or “G/C content modification” relate to a nucleic acid that comprises a modified, suitably an increased number of guanosine and/or cytosine nucleotides as compared to the corresponding wild type or reference coding sequence.
- Such an increased number may be generated by substitution of codons containing adenosine or thymidine nucleotides by codons containing guanosine or cytosine nucleotides.
- nucleic acid sequences having an increased G /C content are more stable or show a better expression than sequences having an increased A/ll.
- the amino acid sequence encoded by the G/C content modified coding sequence of the mRNA is suitably not modified as compared to the amino acid sequence encoded by the respective wild type or reference sequence.
- the G/C content of the coding sequence of the nucleic acid is increased by at least 10%, 20%, 30%, suitably by at least 40% compared to the G/C content of the coding sequence of the corresponding wild type or reference nucleic acid sequence.
- the generation of a G/C content optimized mRNA sequence may be carried out using a method according to W02002/098443. In this context, the disclosure of W02002/098443 is included in its full scope in the present invention.
- the mRNAs used herein may be modified by altering the number of A and/or II nucleotides in the nucleic acid sequence with respect to the number of A and/or II nucleotides in the original nucleic acid sequence (e.g. the wild type or reference sequence).
- such an AU alteration is performed to modify the retention time of the individual nucleic acids in a composition, to (i) allow co-purification using a HPLC method, and/or to allow analysis of the obtained nucleic acid composition.
- Such a method is described in detail in published PCT application WO2019092153A1. Claims 1 to 70 of WO2019092153A1 herewith incorporated by reference.
- the modified RNA sequence is selected from C maximized coding sequence, CAI maximized coding sequence, human codon usage adapted coding sequence, G/C content modified (or optimized) sequence, A/U alteration, or any combination thereof.
- the RNA sequence has a G/C content of at least about 45%, 50%, 55%, or 60%.
- the RNA sequence has a G/C content of at least about 50%, 51%, 52%, 53%, 54%, 55%, 56%, 57%, 58%, 59%, 60%, 61%, 62%, 63%, 64%, 65%, 66%, 67%, 68%, 69%, or 70%.
- the mRNA comprising a modified sequence when transfected into mammalian host cells, has a stability of between 12-18 hours, or greater than 18 hours, e.g., 24, 36, 48, 60, 72, or greater than 72 hours and are capable of being expressed by the mammalian host cell (e.g. a muscle cell).
- the mammalian host cell e.g. a muscle cell
- the mRNA comprising a modified RNA sequence is translated into protein, wherein the amount of protein is at least comparable to, or suitably at least 10% more than, or at least 20% more than, or at least 30% more than, or at least 40% more than, or at least 50% more than, or at least 100% more than, or at least 200% or more than the amount of protein obtained by a naturally occurring or wild type or reference coding sequence transfected into mammalian host cells.
- the mRNAs used herein comprise at least one poly(N) sequence, e.g. at least one poly(A) sequence, at least one poly(ll) sequence, at least one poly(C) sequence, or combinations thereof.
- the mRNAs used herein comprise at least one poly(A) sequence.
- a poly A tail e.g., of about 30 adenosine residues or more
- poly(A) sequence “poly(A) tail” or “3’-poly(A) tail” as used herein will be recognized and understood by the person of ordinary skill in the art, and are e.g. intended to be a sequence of adenosine nucleotides, typically located at the 3’-end of a linear RNA (or in a circular RNA), of up to about 1000 adenosine nucleotides.
- the poly(A) sequence is essentially homopolymeric, e.g. a poly(A) sequence of e.g. 100 adenosine nucleotides has essentially the length of 100 nucleotides.
- the poly(A) sequence may be interrupted by at least one nucleotide different from an adenosine nucleotide, e.g. a poly(A) sequence of e.g. 100 adenosine nucleotides may have a length of more than 100 nucleotides (comprising 100 adenosine nucleotides and in addition the at least one nucleotide - or a stretch of nucleotides - different from an adenosine nucleotide).
- a poly(A) sequence of e.g. 100 adenosine nucleotides may have a length of more than 100 nucleotides (comprising 100 adenosine nucleotides and in addition the at least one nucleotide - or a stretch of nucleotides - different from an adenosine nucleotide).
- the poly(A) sequence may comprise about 10 to about 500 adenosine nucleotides, about 10 to about 200 adenosine nucleotides, about 40 to about 200 adenosine nucleotides, or about 40 to about 150 adenosine nucleotides.
- the length of the poly(A) sequence may be at least about or even more than about 10, 50, 64, 75, 100, 200, 300, 400, or 500 adenosine nucleotides.
- the mRNAs used herein comprise at least one poly(A) sequence comprising about 30 to about 200 adenosine nucleotides.
- the poly(A) sequence comprises about 64 adenosine nucleotides (A64).
- the poly(A) sequence comprises about 100 adenosine nucleotides (A100).
- the poly(A) sequence comprises about 150 adenosine nucleotides.
- the mRNAs used herein comprise at least one poly(A) sequence comprising about 100 adenosine nucleotides, wherein the poly(A) sequence is interrupted by non-adenosine nucleotides, suitably by 10 non-adenosine nucleotides (A30- N10-A70).
- the poly(A) sequence as defined herein may be located directly at the 3’ terminus of the mRNA.
- the 3’-terminal nucleotide (that is the last 3’-terminal nucleotide in the polynucleotide chain) is the 3’-terminal A nucleotide of the at least one poly(A) sequence.
- the term “directly located at the 3’ terminus” has to be understood as being located exactly at the 3’ terminus - in other words, the 3’ terminus of the nucleic acid consists of a poly(A) sequence terminating with an A nucleotide.
- the mRNAs used herein comprise a poly(A) sequence of at least 70 adenosine nucleotides, suitably consecutive at least 70 adenosine nucleotides, wherein the 3’- terminal nucleotide is an adenosine nucleotide.
- the poly(A) sequence of the nucleic acid is obtained from a DNA template during RNA in vitro transcription.
- the poly(A) sequence is obtained in vitro by common methods of chemical synthesis without being necessarily transcribed from a DNA template.
- poly(A) sequences are generated by enzymatic polyadenylation of the RNA (after RNA in vitro transcription) using commercially available polyadenylation kits and corresponding protocols known in the art, or alternatively, by using immobilized poly(A)polymerases e.g. using a methods and means as described in WO2016174271.
- the mRNAs used herein comprise at least one poly(C) sequence.
- poly(C) sequence as used herein is intended to be a sequence of cytosine nucleotides of up to about 200 cytosine nucleotides.
- the poly(C) sequence comprises about 10 to about 200 cytosine nucleotides, about 10 to about 100 cytosine nucleotides, about 20 to about 70 cytosine nucleotides, about 20 to about 60 cytosine nucleotides, or about 10 to about 40 cytosine nucleotides.
- the poly(C) sequence comprises about 30 cytosine nucleotides.
- the mRNAs used herein comprise at least one histone stem-loop (hSL) or histone stem loop structure.
- the mRNAs of (a), (b), (c), (d), (e), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ) comprises at least one histone stem-loop.
- histone stem-loop (abbreviated as “hSL” in e.g. the sequence listing) is intended to refer to nucleic acid sequences that form a stem-loop secondary structure predominantly found in histone mRNAs.
- Histone stem-loop sequences/structures may suitably be selected from histone stemloop sequences as disclosed in WO2012019780, the disclosure relating to histone stem-loop sequences/histone stem-loop structures incorporated herewith by reference.
- a histone stemloop sequence that may be used may be derived from formulae (I) or (II) of W02012019780.
- the mRNA comprises at least one histone stem-loop sequence derived from at least one of the specific formulae (la) or (Ila) of the patent application W02012019780.
- the mRNAs used herein does not comprise a hsL as defined herein.
- mRNAs used herein may be modified by the addition of a 5’-cap structure, which suitably stabilizes the RNA and/or enhances expression of the encoded antigen and/or reduces the stimulation of the innate immune system (after administration to a subject).
- 5’-cap structure as used herein will be recognized and understood by the person of ordinary skill in the art, and is e.g. intended to refer to a 5’ modified nucleotide, particularly a guanine nucleotide, positioned at the 5’-end of an RNA, e.g. an mRNA.
- the 5' end of the RNA may be capped with a modified ribonucleotide with the structure m7G (5') ppp (5') N (cap 0 structure) or a derivative thereof, which can be incorporated during RNA synthesis or can be enzymatically engineered after RNA transcription (e.g., by using Vaccinia Virus Capping Enzyme (VCE) consisting of mRNA triphosphatase, guanylyl-transferase and guanine-7-methytransferase, which catalyzes the construction of NT- monomethylated cap 0 structures).
- VCE Vaccinia Virus Capping Enzyme
- Cap 0 structure plays an important role in maintaining the stability and translational efficacy of the RNA molecule.
- the 5' cap of the mRNA molecule may be further modified by a 2'-O-Methyltransferase which results in the generation of a cap 1 structure (m7Gppp [m2'-O] N), which may further increase translation efficacy.
- the 5’-cap structure is connected via a 5’-5’-triphosphate linkage to the RNA.
- capO methylation of the first nucleobase, e.g. m7GpppN
- cap1 additional methylation of the ribose of the adjacent nucleotide of m7GpppN
- cap2 additional methylation of the ribose of the 2nd nucleotide downstream of the m7GpppN
- cap3 additional methylation of the ribose of the 3rd nucleotide downstream of the m7GpppN
- cap4 additional methylation of the ribose of the 4th nucleotide downstream of the m7GpppN
- ARCA anti-reverse cap analogue
- modified ARCA e.g.
- the mRNAs used herein suitably the mRNAs of (a), (b), (c), (d), (e), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ), comprises a 5’ cap, preferably m7G, capO, cap1 , cap2, a modified capO or a modified cap1 structure, suitably a 5’-cap1 structure.
- a 5’-cap (capO or cap1) structure may be formed in chemical RNA synthesis or in RNA in vitro transcription (co-transcriptional capping) using cap analogues.
- cap analogue as used herein will be recognized and understood by the person of ordinary skill in the art, and is e.g. intended to refer to a non-polymerizable dinucleotide or tri-nucleotide that has cap functionality in that it facilitates translation or localization, and/or prevents degradation of a nucleic acid molecule, particularly of an RNA molecule, when incorporated at the 5’-end of the nucleic acid molecule.
- Non-polymerizable means that the cap analogue will be incorporated only at the 5’-terminus because it does not have a 5’ triphosphate and therefore cannot be extended in the 3’-direction by a templatedependent polymerase, particularly, by template-dependent RNA polymerase.
- cap analogues include, but are not limited to, a chemical structure selected from the group consisting of m7GpppG, m7GpppA, m7GpppC; unmethylated cap analogues (e.g. GpppG); dimethylated cap analogue (e.g. m2,7GpppG), trimethylated cap analogue (e.g. m2,2,7GpppG), dimethylated symmetrical cap analogues (e.g. m7Gpppm7G), or anti reverse cap analogues (e.g.
- cap analogues in that context are described in WO2017066793, WO2017066781 , WO2017066791 , WO2017066789, WO2017/053297, WO2017066782, WO2018075827 and WO2017066797 wherein the disclosures referring to cap analogues are incorporated herewith by reference.
- a modified cap1 structure is generated using tri-nucleotide cap analogue as disclosed in WO2017053297, WO2017066793, WO2017066781 ,
- any cap structures derivable from the structure disclosed in claim 1-5 of WO2017053297 may be suitably used to co-transcriptionally generate a modified cap1 structure.
- any cap structures derivable from the structure defined in claim 1 or claim 21 of WO2018075827 may be suitably used to co-transcriptionally generate a modified cap1 structure.
- the mRNAs used herein comprises a cap1 structure.
- the 5’-cap structure may be added co-transcriptionally using trinucleotide cap analogue as defined herein, suitably in an RNA in vitro transcription reaction as defined herein.
- the cap1 structure of the mRNA is formed using co-transcriptional capping using tri-nucleotide cap analogues m7G(5’)ppp(5’)(2’OMeA)pG or m7G(5’)ppp(5’)(2’OMeG)pG.
- a suitable cap1 analogues in that context is m7G(5’)ppp(5’)(2’OMeA)pG.
- the cap1 structure of the mRNA is formed using co- transcriptional capping using tri-nucleotide cap analogue 3’0Me-m7G(5’)ppp(5’)(2’0MeA)pG.
- a capO structure of the mRNAs used herein is formed using co- transcriptional capping using cap analogue 3’0Me-m7G(5’)ppp(5’)G.
- the 5’-cap structure is formed via enzymatic capping using capping enzymes (e.g. vaccinia virus capping enzymes and/or cap-dependent 2’-0 methyltransferases) to generate capO or cap1 or cap2 structures.
- capping enzymes e.g. vaccinia virus capping enzymes and/or cap-dependent 2’-0 methyltransferases
- the 5’-cap structure (capO or cap1) may be added using immobilized capping enzymes and/or cap-dependent 2’-0 methyltransferases using methods and means disclosed in WO2016193226.
- a capping assay as described in published PCT application W02015101416, in particular, as described in claims 27 to 46 of published PCT application W02015101416 can be used.
- Other capping assays that may be used to determine the presence/absence of a capO or a cap1 structure of an RNA are described in PCT/EP2018/08667, or published PCT applications WO2014152673 and WO2014152659.
- the mRNAs used herein comprise an m7G(5’)ppp(5’)(2’OMeA) cap structure.
- the mRNAs comprise a 5’-terminal m7G cap, and an additional methylation of the ribose of the adjacent nucleotide of m7GpppN, in that case, a 2’0 methylated Adenosine.
- about 70%, 75%, 80%, 85%, 90%, 95% of the RNA (species) comprises such a cap1 structure as determined using a capping assay.
- the mRNAs used herein comprise an m7G(5’)ppp(5’)(2’OMeG) cap structure.
- the mRNAs comprise a 5’-terminal m7G cap, and an additional methylation of the ribose of the adjacent nucleotide, in that case, a 2’0 methylated guanosine.
- about 70%, 75%, 80%, 85%, 90%, 95% of the coding RNA (species) comprises such a cap1 structure as determined using a capping assay.
- the first nucleotide of the mRNA sequence may be a 2’0 methylated guanosine or a 2’0 methylated adenosine.
- the mRNAs used herein comprise a ribosome binding site, also referred to as “Kozak sequence”.
- the A/ll (A/T) content in the environment of the ribosome binding site of the mRNAs used herein may be increased compared to the A/ll (A/T) content in the environment of the ribosome binding site of its respective wild type or reference nucleic acid. This modification (an increased A/ll (A/T) content around the ribosome binding site) increases the efficiency of ribosome binding to the mRNA.
- An effective binding of the ribosomes to the ribosome binding site in turn has the effect of an efficient translation the mRNA.
- the mRNAs used herein may comprise at least one heterologous untranslated region (UTR), e.g. a 5’ UTR and/or a 3’ UTR.
- UTR heterologous untranslated region
- UTR untranslated region
- UTR element The term “untranslated region” or “UTR” or “UTR element” will be recognized and understood by the person of ordinary skill in the art, and are e.g. intended to refer to a part of a nucleic acid molecule typically located 5’ or 3’ of a coding sequence.
- An UTR is not translated into protein.
- An UTR may be part of a nucleic acid, e.g. a DNA or an RNA.
- An UTR may comprise elements for controlling gene expression, also called regulatory elements. Such regulatory elements may be, e.g., ribosomal binding sites, miRNA binding sites, promotor elements etc.
- the mRNAs used herein comprise a protein-coding region (“coding sequence” or “cds”), and 5’-UTR and/or 3’-UTR.
- UTRs may harbor regulatory sequence elements that determine nucleic acid, e.g. RNA turnover, stability, and localization.
- UTRs may harbor sequence elements that enhance translation.
- nucleic acid sequences including DNA and RNA
- translation of the nucleic acid into at least one peptide or protein is of paramount importance to therapeutic efficacy.
- Certain combinations of 3’-UTRs and/or 5’-UTRs may enhance the expression of operably linked coding sequences encoding peptides or proteins of the invention.
- Nucleic acid molecules harboring the UTR combinations advantageously enable rapid and transient expression of antigenic peptides or proteins after administration to a subject, suitably after intramuscular administration.
- the mRNA comprising certain combinations of 3’-UTRs and/or 5’- UTRs as provided herein is particularly suitable for administration as a vaccine, in particular, suitable for administration into the muscle, the dermis, or the epidermis of a subject.
- the mRNAs used herein comprise at least one heterologous 5’- UTR and/or at least one heterologous 3’-UTR.
- the heterologous 5’-UTRs or 3’-UTRs may be derived from naturally occurring genes or may be synthetically engineered.
- the mRNA comprises at least one coding sequence as defined herein operably linked to at least one (heterologous) 3’-UTR and/or at least one (heterologous) 5’-UTR.
- the mRNAs used herein comprise at least one heterologous 3’-UTR.
- the mRNAs of (a), (b), (c), (d), (e), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ) comprises a 3’ UTR.
- 3’-untranslated region or “3’-UTR” or “3’-UTR element” will be recognized and understood by the person of ordinary skill in the art, and are e.g. intended to refer to a part of a nucleic acid molecule located 3’ (i.e. downstream) of a coding sequence and which is not translated into protein.
- a 3’-UTR may be part of a nucleic acid, e.g. a DNA or an RNA, located between a coding sequence and an (optional) terminal poly(A) sequence.
- a 3’-UTR may comprise elements for controlling gene expression, also called regulatory elements. Such regulatory elements may be, e.g., ribosomal binding sites, miRNA binding sites etc.
- the mRNAs used herein comprise a 3’-UTR, which may be derivable from a gene that relates to an RNA with enhanced half-life (i.e. that provides a stable RNA).
- a 3’-UTR comprises one or more of a polyadenylation signal, a binding site for proteins that affect a nucleic acid stability of location in a cell, or one or more miRNA or binding sites for miRNAs.
- the mRNAs used herein comprise at least one heterologous 3’-UTR, wherein the at least one heterologous 3’-UTR comprises a nucleic acid sequence is derived or selected from a 3’-UTR of a gene selected from PSMB3, ALB7, alpha-globin (referred to as “muag”), CASP1 , COX6B1 , GNAS, NDUFA1 and RPS9, or from a homolog, a fragment or variant of any one of these genes.
- muag alpha-globin
- the mRNAs of (a), (b), (c), (d), (e), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ) comprises a 3’ UTR comprising or consisting of a nucleic acid sequence derived from a 3’-UTR of a gene selected from PSMB3, ALB7, CASP1 , COX6B1 , GNAS, NDUFA1 and RPS9, or from a homolog, a fragment or a variant of any one of these genes.
- Nucleic acid sequences in that context can be derived from published PCT application W02019077001A1 , in particular, claim 9 of WO2019077001 A1.
- the corresponding 3’-UTR sequences of claim 9 of W02019077001A1 are herewith incorporated by reference.
- the mRNAs used herein may comprise a 3’-UTR as described in WO2016107877, the disclosure of WO2016107877 relating to 3’-UTR sequences herewith incorporated by reference. Suitable 3’-UTRs are SEQ ID NOs: 1-24 and SEQ ID NOs: 49-318 of WO2016107877, or fragments or variants of these sequences.
- the mRNAs used herein comprise a 3’-UTR as described in WO2017036580, the disclosure of WO2017036580 relating to 3’-UTR sequences herewith incorporated by reference.
- Suitable 3’-UTRs are SEQ ID NOs: 152-204 of WO2017036580, or fragments or variants of these sequences.
- the mRNAs used herein comprise a 3’-UTR as described in WO2016022914, the disclosure of WO2016022914 relating to 3’-UTR sequences herewith incorporated by reference.
- Particularly suitable 3’-UTRs are nucleic acid sequences according to SEQ ID NOs: 20-36 of WQ2016022914, or fragments or variants of these sequences.
- the mRNAs used herein comprise at least one heterologous 5’-UTR.
- the mRNAs of (a), (b), (c), (d), (e), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ) comprises a 5’ untranslated region (UTR).
- 5’-untranslated region or “5’-UTR” or “5’-UTR element” will be recognized and understood by the person of ordinary skill in the art, and are e.g. intended to refer to a part of a nucleic acid molecule located 5’ (i.e. “upstream”) of a coding sequence and which is not translated into protein.
- a 5’-UTR may be part of a nucleic acid located 5’ of the coding sequence.
- a 5’-UTR starts with the transcriptional start site and ends before the start codon of the coding sequence.
- a 5’-UTR may comprise elements for controlling gene expression, also called regulatory elements. Such regulatory elements may be, e.g., ribosomal binding sites, miRNA binding sites etc.
- the 5’-UTR may be post-transcriptionally modified, e.g. by enzymatic or post-transcriptional addition of a 5’-cap structure (e.g. for mRNA as defined herein).
- the mRNAs used herein comprise a 5’-UTR, which may be derivable from a gene that relates to an RNA with enhanced half-life (i.e. that provides a stable RNA).
- a 5’-UTR comprises one or more of a binding site for proteins that affect an RNA stability or RNA location in a cell, or one or more miRNA or binding sites for miRNAs.
- the mRNAs used herein suitably the mRNAs of (a), (b), (c), (d), (e), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ), comprise at least one heterologous 5’-UTR, wherein the at least one heterologous 5’-UTR comprises a nucleic acid sequence is derived or selected from a 5’-UTR of gene selected from HSD17B4, RPL32, ASAH1 , ATP5A1 , MP68, NDUFA4, NOSIP, RPL31 , SLC7A3, TLIBB4B, and LIBQLN2, or from a homolog, a fragment or variant of any one of these genes.
- Nucleic acid sequences in that context can be selected from published PCT application W02019077001A1 , in particular, claim 9 of WO2019077001 A1.
- the corresponding 5’-UTR sequences of claim 9 of WO2019077001 A1 are herewith incorporated by reference (e.g., SEQ ID NOs: 1-20 of WO2019077001 A1 , or fragments or variants thereof).
- the mRNAs used herein may comprise a 5’-UTR as described in W02013143700, the disclosure of W02013143700 relating to 5’-UTR sequences herewith incorporated by reference.
- Particularly suitable 5’-UTRs are nucleic acid sequences derived from SEQ ID NOs: 1-1363, SEQ ID NO: 1395, SEQ ID NO: 1421 and SEQ ID NO: 1422 of WQ2013143700, or fragments or variants of these sequences.
- the mRNAs used herein comprise a 5’-UTR as described in WQ2016107877, the disclosure of WQ2016107877 relating to 5’-UTR sequences herewith incorporated by reference.
- Particularly suitable 5’-UTRs are nucleic acid sequences according to SEQ ID NOs: 25-30 and SEQ ID NOs: 319-382 of WQ2016107877, or fragments or variants of these sequences.
- the nucleic acid comprises a 5’-UTR as described in WQ2017036580, the disclosure of WQ2017036580 relating to 5’-UTR sequences herewith incorporated by reference.
- Particularly suitable 5’-UTRs are nucleic acid sequences according to SEQ ID NOs: 1-151 of WQ2017036580, or fragments or variants of these sequences.
- the nucleic acid comprises a 5’-UTR as described in WQ2016022914, the disclosure of WQ2016022914 relating to 5’-UTR sequences herewith incorporated by reference.
- Particularly suitable 5’-UTRs are nucleic acid sequences according to SEQ ID NOs: 3-19 of WQ2016022914, or fragments or variants of these sequences.
- the mRNAs of (a), (b), (c), (d), (e), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ) comprises an heterologous 5’-UTR that comprises or consists of a nucleic acid sequence derived from a 5’-UTR from HSD17B4 and at least one heterologous 3’-UTR comprises or consists of a nucleic acid sequence derived from a 3’-UTR of PSMB3.
- the mRNAs used herein suitably the mRNAs of (a), (b), (c), (d), (e), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ), comprises from 5’ to 3’: i) 5’-cap1 structure; ii) 5’-UTR derived from a 5’-UTR of a HSD17B4 gene; iii) the coding sequence; iv) 3’-UTR derived from a 3’-UTR of a PSMB3 gene; v) optionally, a histone stem-loop sequence; and vi) poly(A) sequence comprising about 100 A nucleotides, wherein the 3’ terminal nucleotide of said RNA is an adenosine.
- RNA suitably mRNA
- of the composition has an RNA integrity ranging from about 40% to about 100%.
- RNA integrity generally describes whether the complete RNA sequence is present in the composition. Low RNA integrity could be due to, amongst others, RNA degradation, RNA cleavage, incorrect or incomplete chemical synthesis of the RNA, incorrect base pairing, integration of modified nucleotides or the modification of already integrated nucleotides, lack of capping or incomplete capping, lack of polyadenylation or incomplete polyadenylation, or incomplete RNA in vitro transcription.
- RNA is a fragile molecule that can easily degrade, which may be caused e.g. by temperature, ribonucleases, pH or other factors (e.g. nucleophilic attacks, hydrolysis etc.), which may reduce the RNA integrity and, consequently, the functionality of the RNA.
- RNA integrity can be determined from a variety of different chromatographic or electrophoretic methods for determining an RNA integrity. Chromatographic and electrophoretic methods are well-known in the art. In case chromatography is used (e.g. RP- HPLC), the analysis of the integrity of the RNA may be based on determining the peak area (or “area under the peak”) of the full length RNA in a corresponding chromatogram. The peak area may be determined by any suitable software which evaluates the signals of the detector system. The process of determining the peak area is also referred to as integration. The peak area representing the full-length RNA is typically set in relation to the peak area of the total RNA in a respective sample. The RNA integrity may be expressed in % RNA integrity.
- RNA integrity may be determined using analytical (RP)HPLC.
- a test sample of the composition comprising lipid based carrier encapsulating RNA may be treated with a detergent (e.g. about 2% Triton X100) to dissociate the lipid based carrier and to release the encapsulated RNA.
- the released RNA may be captured using suitable binding compounds, e.g. Agencourt AMPure XP beads (Beckman Coulter, Brea, CA, USA) essentially according to the manufacturer’s instructions.
- analytical (RP)HPLC may be performed to determine the integrity of RNA.
- the RNA samples may be diluted to a concentration of 0.1 g/l using e.g. water for injection (WFI).
- WFI water for injection
- About 10pl of the diluted RNA sample may be injected into an HPLC column (e.g. a monolithic poly(styrene-divinylbenzene) matrix).
- HPLC column e.g. a monolithic poly(styrene-divinylbenzene) matrix.
- Analytical (RP)HPLC may be performed using standard conditions, for example: Gradient 1 : Buffer A (0.1 M TEAA (pH 7.0)); Buffer B (0.1 M TEAA (pH 7.0) containing 25% acetonitrile).
- RNA integrity in the context of the invention is determined using analytical HPLC, suitably analytical RP-HPLC.
- RNA, suitably mRNA, of the composition has an RNA integrity ranging from about 40% to about 100%. In embodiments, the RNA, suitably mRNA, has an RNA integrity ranging from about 50% to about 100%. In embodiments, the RNA, suitably mRNA, has an RNA integrity ranging from about 60% to about 100%. In embodiments, the RNA, suitably mRNA, has an RNA integrity ranging from about 70% to about 100%. In embodiments, the RNA, suitably mRNA, integrity is for example about 50%, about 60%, about 70%, about 80%, or about 90%. RNA integrity is suitably determined using analytical HPLC, suitably analytical RP-HPLC.
- the RNA, suitably mRNA, of the composition has an RNA integrity of at least about 50%, suitably of at least about 60%, more suitably of at least about 70%, most suitably of at least about 80% or about 90%.
- RNA integrity is suitably determined using analytical HPLC, more suitably analytical RP-HPLC.
- the RNAs suitably mRNAs, used herein does not comprise a replicase element (e.g. a nucleic acid encoding a replicase).
- a replicase element e.g. a nucleic acid encoding a replicase
- the RNAs used herein suitably the mRNAs used herein, suitably the mRNAs of (a), (b), (c), (d), (e), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ), optionally each, are not self-replicating.
- the RNAs used herein, suitably the mRNA used herein, suitably the mRNAs of (a), (b), (c), (d), (e), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ), optionally each, are self-replicating.
- the RNA suitably mRNA, comprises a coding sequence that consists only of G, C, A and II nucleotides and therefore does not comprise modified nucleotides (except of the 5’ terminal cap structure (capO, cap1 , cap2)).
- the mRNAs of (a), (b), (c), (d), (e), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ) does not comprise chemically modified nucleotides.
- the RNAs are modified RNAs, suitably mRNAs, wherein the modification refers to chemical modifications comprising backbone modifications as well as sugar modifications or base modifications.
- a modified RNA may comprise one or more nucleotide analogs or modified nucleotides (nucleotide analogues/modifications, e.g. backbone modifications, sugar modifications or base modifications).
- nucleotide analog or “modified nucleotide” refers to a nucleotide that contains one or more chemical modifications (e.g., substitutions) in or on the nitrogenous base of the nucleoside (e.g. cytosine (C), thymine (T) or uracil (II)), adenine (A) or guanine (G)) and/or one or more chemical modifications in or one the phosphates of the backbone.
- C cytosine
- T thymine
- II uracil
- A adenine
- G guanine
- a nucleotide analog can contain further chemical modifications in or on the sugar moiety of the nucleoside (e.g. ribose, modified ribose, sixmembered sugar analog, or open-chain sugar analog), or the phosphate.
- the preparation of nucleotides and modified nucleotides and nucleosides are well-known in the art, see the following references: US Patent Numbers 4373071 , 4458066, 4500707, 4668777, 4973679, 5047524, 5132418, 5153319, 5262530, 5700642. Many modified nucleosides and modified nucleotides are commercially available.
- a backbone modification as described herein is a modification, in which phosphates of the backbone of the nucleotides of the RNA, suitably the mRNA, are chemically modified.
- a sugar modification as described herein is a chemical modification of the sugar of the nucleotides of the RNA, suitably mRNA.
- a base modification as described herein is a chemical modification of the base moiety of the nucleotides of the RNA, suitably mRNA.
- nucleotide analogues or modifications are suitably selected from nucleotide analogues which are applicable for transcription and/or translation.
- the RNAs suitably the mRNAs, used herein comprise at least one chemical modification.
- the mRNAs of (a), (b), (c), (d), (e), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ) comprises at least one chemical modification.
- Modified nucleobases which can be incorporated into modified nucleosides and nucleotides and be present in the RNA, suitably mRNA, molecules include: m5C (5-methylcytidine), m5U (5-methyluridine), m6A (N6-methyladenosine), s2U (2- thiouridine), Um (2'-O-methyluridine), m1A (1-methyladenosine); m2A (2-methyladenosine); Am (2-1-O-methyladenosine); ms2m6A (2-methylthio-N6-methyladenosine); i6A (N6- isopentenyladenosine); ms2i6A (2-methylthio-N6isopentenyladenosine); io6A (N6-(cis- hydroxyisopentenyl)adenosine); ms2io6A (2-methylthio-N6-(cis- hydroxyis
- the nucleotide analogues/modifications which may be incorporated into a modified RNA, suitably mRNA are selected from 2-amino-6- chloropurineriboside-5’-triphosphate, 2-Aminopurine-riboside-5’-triphosphate; 2- aminoadenosine-5’-triphosphate, 2’-Amino-2’-deoxycytidine-triphosphate, 2-thiocytidine-5’- triphosphate, 2-thiouridine-5’-triphosphate, 2’-Fluorothymidine-5’-triphosphate, 2’-O-Methyl- inosine-5’-triphosphate 4-thiouridine-5’-triphosphate, 5-aminoallylcytidine-5’-triphosphate, 5- aminoallyluridine-5’-triphosphate, 5-bromocytidine-5’-triphosphate, 5-bromouridine-5’- triphosphate, 5-Bromo-2’-deoxycyt
- nucleotides for base modifications selected from the group of base-modified nucleotides consisting of 5- methylcytidine-5’-triphosphate, 7-deazaguanosine-5’-triphosphate, 5-bromocytidine-5’- triphosphate, and pseudouridine-5’-triphosphate, pyridin-4-one ribonucleoside, 5-aza-uridine, 2-thio-5-aza-uridine, 2-thiouridine, 4-thio-pseudouridine, 2-thio-pseudouridine, 5- hydroxyuridine, 3-methyluridine, 5-carboxymethyl-uridine, 1-carboxymethyl-pseudouridine, 5- propynyl-uridine, 1-propynyl-pseudouridine, 5-taurinomethyluridine, 1-taurinomethyl- pseudouridine, 5-taurinomethyl-2-thio-uridine, 1-taurinomethyl-4-thio-uridine
- the chemical modification is selected from pseudouridine, N1- methylpseudouridine, N1-ethylpseudouridine, 2-thiouridine, 4'-thiouridine, 5-methylcytosine, 5- methyluridine, 2-thio-1-methyl-1-deaza-pseudouridine, 2-thio-1-methyl-pseudouridine, 2-thio- 5-aza-uridine , 2-thio-dihydropseudouridine, 2-thio-dihydrouridine, 2-thio-pseudouridine, 4- methoxy-2-thio-pseudouridine, 4-methoxy-pseudouridine, 4-thio-1-methyl-pseudouridine, 4- thio-pseudouridine, 5-aza-uridine, dihydropseudouridine, 5-methoxyuridine and 2'-O-methyl uridine.
- pseudouridine qj
- N1-methylpseudouridine m1 ip
- 5-methylcytosine and 5-methoxyuridine
- pseudouridine ip
- N1- methylpseudouridine m1 ip
- N1 -methylpseudouridine m1 ip
- RNAs suitably mRNAs, used herein have a chemical modification, suitably a chemical modification is in the 5-position of the uracil.
- the RNAs suitably mRNAs, used herein comprise the chemical modification being a uridine modification, preferably wherein 100% of the uridine positions in the mRNA are modified.
- RNAs e.g. pseudouridine (qj), N1- methylpseudouridine (m1 ip), 5-methylcytosine, and/or 5-methoxyuridine into the coding sequence of the RNAs, suitably mRNAs, used herein may be advantageous as unwanted innate immune responses (upon administration of the coding mRNA or the vaccine) may be adjusted or reduced (if required).
- pseudouridine qj
- m1 ip N1- methylpseudouridine
- 5-methylcytosine 5-methoxyuridine
- the coding sequence of the RNAs suitably mRNAs, used herein comprise at least one modified nucleotide selected from pseudouridine (ip) and N1- methylpseudouridine (m1 ip), suitably wherein all uracil nucleotides are replaced by pseudouridine (ip) nucleotides and/or N1-methylpseudouridine (m1 ip) nucleotides, optionally wherein all uracil nucleotides are replaced by pseudouridine ( ⁇ P) nucleotides and/or N1- methylpseudouridine (ml ⁇ P) nucleotides.
- pseudouridine ip
- m1 ip N1- methylpseudouridine
- the RNAs, suitably mRNAs, used herein do not comprise N1- methylpseudouridine (ml ⁇ P) substituted positions. In further embodiments, the RNAs, suitably mRNAs, used herein do not comprise pseudouridine (ip), N1 -methylpseudouridine (m1 ip), 5- methylcytosine, and 5-methoxyuridine substituted position.
- the chemical modification is N1 -methylpseudouridine and/or pseudouridine. In some embodiments, the chemical modification is N1 -methylpseudouridine.
- RNA production is performed under current good manufacturing practice (GMP), implementing various quality control steps on DNA and RNA level, suitably according to W02016180430.
- GMP current good manufacturing practice
- the RNA, suitably mRNA of the invention is a GMP-grade RNA.
- the RNA suitably mRNA
- the RNA may be prepared using any method known in the art, including chemical synthesis such as e.g. solid phase RNA synthesis, as well as in vitro methods, such as RNA in vitro transcription reactions.
- RNA suitably mRNA, used herein is in vitro transcribed RNA.
- RNA in vitro transcription or “in vitro transcription” relate to a process wherein RNA is synthesized in a cell-free system in vitro).
- RNA may be obtained by DNA- dependent in vitro transcription of an appropriate DNA template, which may be a linearized plasmid DNA template or a PCR-amplified DNA template.
- the promoter for controlling RNA in vitro transcription can be any promoter for any DNA-dependent RNA polymerase.
- DNA-dependent RNA polymerases are the T7, T3, SP6, or Syn5 RNA polymerases.
- the DNA template is linearized with a suitable restriction enzyme, before it is subjected to RNA in vitro transcription.
- Reagents used in RNA in vitro transcription typically include: a DNA template (linearized plasmid DNA or PCR product) with a promoter sequence that has a high binding affinity for its respective RNA polymerase such as bacteriophage-encoded RNA polymerases (T7, T3, SP6, or Syn5); ribonucleotide triphosphates (NTPs) for the four bases (adenine, cytosine, guanine and uracil); optionally, a cap analogue as defined herein; optionally, further modified nucleotides as defined herein; a DNA-dependent RNA polymerase capable of binding to the promoter sequence within the DNA template (e.g.
- RNA polymerase T7, T3, SP6, or Syn5 RNA polymerase
- RNase ribonuclease
- a pyrophosphatase to degrade pyrophosphate, which may inhibit RNA in vitro transcription
- MgCI2 which supplies Mg2+ ions as a co-factor for the polymerase
- a buffer TRIS or HEPES
- polyamines such as spermidine at optimal concentrations, e.g. a buffer system comprising TRIS-Citrate as disclosed in W02017109161.
- the nucleotide mixture used in RNA in vitro transcription may additionally comprise modified nucleotides as defined herein.
- suitable modified nucleotides may in particular be selected from pseudouridine (i ), N1-methylpseudouridine (m1 i ), 5-methylcytosine, and 5-methoxyuridine.
- uracil nucleotides in the nucleotide mixture are replaced (either partially or completely) by pseudouridine (i ) and/or N1- methylpseudouridine (m1 ip) to obtain a modified RNA.
- the nucleotide mixture used in RNA in vitro transcription does not comprise modified nucleotides as defined herein.
- the nucleotide mixture used in RNA in vitro transcription only comprises G, C, A and II nucleotides, and, optionally, a cap analog as defined herein.
- the nucleotide mixture i.e. the fraction of each nucleotide in the mixture
- the nucleotide mixture used for RNA in vitro transcription reactions may be optimized for the given RNA sequence, suitably as described in WO2015188933.
- the in vitro transcription has been performed in the presence of a sequence optimized nucleotide mixture and optionally a cap analog.
- the RNA, suitably mRNA, used herein is a purified RNA, suitably mRNA.
- purified RNA or mRNA
- RNA which has a higher purity after certain purification steps (e.g. HPLC, TFF, Oligo d(T) purification, precipitation steps) than the starting material (e.g. in vitro transcribed RNA).
- Typical impurities that are essentially not present in purified RNA comprise peptides or proteins (e.g. enzymes derived from DNA dependent RNA in vitro transcription, e.g.
- RNA polymerases RNases, pyrophosphatase, restriction endonuclease, DNase), spermidine, BSA, abortive RNA sequences, RNA fragments (short double stranded RNA fragments, abortive sequences etc.), free nucleotides (modified nucleotides, conventional NTPs, cap analogue), template DNA fragments, buffer components (HEPES, TRIS, MgCI2) etc.
- Other potential impurities that may be derived from e.g. fermentation procedures comprise bacterial impurities (bioburden, bacterial DNA) or impurities derived from purification procedures (organic solvents etc.).
- purified RNA (or mRNA) has a degree of purity of more than 75%, 80%, 85%, very particularly 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% and most favorably 99% or more.
- the degree of purity may for example be determined by an analytical HPLC, wherein the percentages provided above correspond to the ratio between the area of the peak for the target RNA and the total area of all peaks representing the by-products.
- the degree of purity may for example be determined by an analytical agarose gel electrophoresis or capillary gel electrophoresis.
- the RNA is purified using RP-HPLC, suitably using Reversed- Phase High pressure liquid chromatography (RP-HPLC) with a macroporous styrene/divinylbenzene column (e.g. particle size 30pm, pore size 4000 A) and additionally using a filter cassette with a cellulose based membrane with a molecular weight cutoff of about 100kDa.
- RP-HPLC Reversed- Phase High pressure liquid chromatography
- the RNA may in particular be purified using PUREMESSENGER (CureVac, Tubingen, Germany; RP-HPLC according to W02008077592) and/or tangential flow filtration (as described in WO2016193206) and/or oligo d(T) purification (see WO2016180430).
- the RNA is purified by RP-HPLC and/or TFF to remove double-stranded RNA, non-capped RNA and/or RNA fragments.
- RNA in vitro transcription can lead to an induction of the innate immune response, particularly IFNalpha which is the main factor of inducing fever in vaccinated subjects, which is of course an unwanted side effect.
- Current techniques for immunoblotting of dsRNA via dot Blot, serological specific electron microscopy (SSEM) or ELISA for example) are used for detecting and sizing dsRNA species from a mixture of nucleic acids.
- the RNA suitably mRNA, comprises about 5%, 10%, or 20% less double stranded RNA side products as an RNA, suitably mRNA, that has not been purified with RP-HPLC and/or TFF.
- the RP-HPLC and/or TFF purified RNA comprises about 5%, 10%, or 20% less double stranded RNA side products as an RNA, suitably mRNA, that has been purified with Oligo dT purification, precipitation, filtration and/or A EX.
- RNAs suitably mRNAs
- the RNAs, suitably mRNAs, used herein are complexed, encapsulated, partially encapsulated, or associated with one or more lipids (e.g. cationic lipids and/or neutral lipids), thereby forming lipid-based carriers such as liposomes, lipid nanoparticles (LNPs), lipoplexes, and/or nanoliposomes, suitably lipid nanoparticles.
- lipids e.g. cationic lipids and/or neutral lipids
- the mRNAs of (a), (b), (c), (d), (e), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ) are formulated in a lipid nanoparticle (LNP), either separately or together.
- LNP lipid nanoparticle
- the RNAs, suitably mRNAs, used herein are formulated separately (in any formulation or complexation agent defined herein), suitably wherein the RNAs, suitably mRNAs, used herein are formulated in separate liposomes, lipid nanoparticles (LNP), lipoplexes, and/or nanoliposomes.
- RNAs, suitably mRNAs, used herein are formulated separately (in any formulation or complexation agent defined herein), suitably wherein the RNAs, suitably mRNAs, used herein are formulated in separate liposomes, lipid nanoparticles (LNP), lipoplexes, and/or nanoliposomes.
- LNP lipid nanoparticles
- the RNAs used herein suitably the mRNAs used herein, suitably the mRNAs of (a), (b), (c), (d), (e), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ) are formulated separately.
- the RNAs, suitably mRNAs, used herein are co-formulated (in any formulation or complexation agent defined herein), suitably wherein the RNAs, suitably mRNAs, used herein are formulated in separate liposomes, lipid nanoparticles (LNP), lipoplexes, and/or nanoliposomes.
- LNP lipid nanoparticles
- the RNAs used herein suitably the mRNAs used herein, suitably the mRNAs of (a), (b), (c), (d), (e), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ) are co-formulated, i.e. formulated together.
- lipid nanoparticle refers to a non-virion particle in which nucleic acid molecules, such as RNA, can be encapsulated.
- LNPs are not restricted to any particular morphology, and include any morphology generated when an ionizable (or cationic) lipid and optionally one or more further lipids are combined, e.g. in an aqueous environment and/or in the presence of a nucleic acid, e.g. an RNA.
- a liposome, a lipid complex, a lipoplex and the like are within the scope of a lipid nanoparticle (LNP).
- LNP delivery systems and methods for their preparation are known in the art.
- the particles can include some external RNA, suitably mRNA, (e.g. on the surface of the particles), but desirably at least half of the RNA, suitably mRNA, (and suitably at least 85%, especially at least 95%, such as all of it) is encapsulated.
- LNPs are suitable for intramuscular and/or intradermal administration.
- LNPs typically comprise a cationic lipid and one or more excipients selected from neutral lipids, charged lipids, steroids and polymer conjugated lipids (e.g. PEGylated lipid).
- the RNAs suitably mRNAs, may be encapsulated in the lipid portion of the LNP or an aqueous space enveloped by some or the entire lipid portion of the LNP.
- the RNAs, suitably mRNAs, or a portion thereof may also be associated and complexed with the LNP.
- An LNP may comprise any lipid capable of forming a particle to which the nucleic acids are attached, or in which the one or more nucleic acids are encapsulated.
- the LNP comprising nucleic acids comprises one or more cationic lipids, and one or more stabilizing lipids.
- Stabilizing lipids include neutral lipids and PEGylated lipids.
- the LNP comprises a PEG-modified lipid, a non-cationic lipid, a sterol, and a cationic lipid.
- LNP can, for example, be formed of a mixture of (i) a PEG-modified lipid (ii) a noncationic lipid (iii) a sterol (iv) an ionisable cationic lipid.
- LNP can for example be formed of a mixture of (i) a PEG-modified lipid (ii) a non-cationic lipid (iii) a sterol (iv) a non- ionisable cationic lipid.
- the non-cationic lipid is a neutral lipid.
- the cationic lipid is ionizable.
- LNPs In vivo characteristics and behavior of LNPs can be modified by addition of a hydrophilic polymer coating, e.g. polyethylene glycol (PEG), to the LNP surface to confer steric stabilization.
- a hydrophilic polymer coating e.g. polyethylene glycol (PEG)
- PEG polyethylene glycol
- LNPs can be used for specific targeting by attaching ligands (e.g. antibodies, peptides, and carbohydrates) to its surface or to the terminal end of the attached PEG chains (e.g. via PEGylated lipids or PEGylated cholesterol).
- the RNA is complexed with one or more lipids thereby forming lipid nanoparticles, wherein the LNP (or liposomes, nanoliposomes, lipoplexes) comprises a polymer conjugated lipid, suitably a PEGylated lipid/PEG lipid.
- the LNPs comprise a polymer conjugated lipid.
- polymer conjugated lipid refers to a molecule comprising both a lipid portion and a polymer portion.
- An example of a polymer conjugated lipid is a PEGylated lipid.
- PEGylated lipid or “PEG-modified lipid” refers to a molecule comprising both a lipid portion and a polyethylene glycol portion.
- PEGylated lipids are known in the art and include 1- (monomethoxy-polyethyleneglycol)-2,3-dimyristoylglycerol (PEG-s-DMG) and the like.
- PEGylated lipid and “PEG-modified lipid” are used interchangeably herein.
- a polymer conjugated lipid as defined herein, e.g. a PEG-lipid, may serve as an aggregation reducing lipid.
- the LNP comprises a stabilizing-lipid which is a polyethylene glycol-lipid (PEGylated lipid).
- Suitable polyethylene glycol-lipids include PEG-modified phosphatidylethanolamine, PEG-modified phosphatidic acid, PEG-modified ceramides (e.g. PEG-CerC14 or PEG-CerC20), PEG-modified dialkylamines, PEG-modified diacylglycerols, PEG-modified dialkylglycerols.
- Representative polyethylene glycol-lipids include PEG-c- DOMG, PEG-c-DMA, and PEG-s-DMG.
- the polyethylene glycol-lipid is N- [(methoxy poly(ethylene glycol)2000)carbamyl]-1 ,2-dimyristyloxlpropyl-3-amine (PEG-c- DMA). In some embodiments, the polyethylene glycol-lipid is PEG-2000-DMG. In one embodiment, the polyethylene glycol-lipid is PEG-c-DOMG).
- the LNPs comprise a PEGylated diacylglycerol (PEG-DAG) such as 1-(monomethoxy- polyethyleneglycol)-2,3-dimyristoylglycerol (PEG-DMG), a PEGylated phosphatidylethanoloamine (PEG-PE), a PEG succinate diacylglycerol (PEG-S-DAG) such as
- PEG-DAG PEGylated diacylglycerol
- PEG-DMG 1-(monomethoxy- polyethyleneglycol)-2,3-dimyristoylglycerol
- PEG-PE PEGylated phosphatidylethanoloamine
- PEG-S-DAG PEG succinate diacylglycerol
- 5-DMG a PEGylated ceramide (PEG-cer), or a PEG dialkoxypropylcarbamate such as w- methoxy(polyethoxy)ethyl-N-(2,3di(tetradecanoxy)propyl)carbamate or 2,3- di(tetradecanoxy)propyl-N-(w-methoxy(polyethoxy)ethyl)carbamate.
- PEG-cer PEGylated ceramide
- PEG dialkoxypropylcarbamate such as w- methoxy(polyethoxy)ethyl-N-(2,3di(tetradecanoxy)propyl)carbamate or 2,3- di(tetradecanoxy)propyl-N-(w-methoxy(polyethoxy)ethyl)carbamate.
- the PEG-modified lipid comprises PEG-DMG or PEG-cDMA.
- the PEGylated lipid is suitably derived from formula (IV) of published PCT patent application W02018078053A1. Accordingly, PEGylated lipids derived from formula (IV) of published PCT patent application W02018078053A1 , and the respective disclosure relating thereto, are herewith incorporated by reference.
- the PEG-modified lipid has the formula IV: wherein R 8 and R 9 are each independently a straight or branched, saturated or unsaturated alkyl chain containing from 10 to 30 carbon atoms, wherein the alkyl chain is optionally interrupted by one or more ester bonds; and w has a mean value ranging from 30 to 60.
- the PEG-modified lipid R 8 and R 9 are saturated alkyl chains.
- the RNA is complexed with one or more lipids thereby forming LNPs
- the LNP comprises a polymer conjugated lipid, suitably a PEGylated lipid, wherein the PEG lipid is suitably derived from formula (IVa) of published PCT patent application W02018078053A1.
- PEGylated lipid derived from formula (IVa) of published PCT patent application W02018078053A1 is herewith incorporated by reference.
- the PEG lipid or PEGylated lipid is of formula (IVa): wherein n has a mean value ranging from 30 to 60, such as about 30 ⁇ 2, 32 ⁇ 2, 34 ⁇ 2, 36 ⁇ 2, 38 ⁇ 2, 40 ⁇ 2, 42 ⁇ 2, 44 ⁇ 2, 46 ⁇ 2, 48 ⁇ 2, 50 ⁇ 2, 52 ⁇ 2, 54 ⁇ 2, 56 ⁇ 2, 58 ⁇ 2, or 60 ⁇ 2. In an embodiment n is about 49. In another embodiment n is about 45. In further embodiments, the PEG lipid is of formula (IVa) wherein n is an integer selected such that the average molecular weight of the PEG lipid is about 2000g/mol to about 3000 g/mol or about 2300g/mol to about 2700g/mol, suitably about 2500g/mol.
- the PEG-modified lipid has the formula IVa: wherein n has a mean value ranging from 30 to 60, suitably wherein n has a mean value of about 45, 46, 47, 48, 49, 50, 51 , 52, 53, 54, most suitably wherein n has a mean value of 49 or 45; or wherein n is an integer selected such that the average molecular weight of the PEG lipid is about 2500g/mol.
- the lipid of formula IVa as suitably used herein has the chemical term 2[(polyethylene glycol)-2000]-N,N-ditetradecylacetamide, also referred to as ALC-0159. Further examples of PEG-lipids suitable in that context are provided in LIS20150376115A1 and WO2015199952, each of which is incorporated by reference in its entirety.
- LNPs include less than about 3, 2, or 1 mole percent of PEG or PEG-modified lipid, based on the total moles of lipid in the LNP.
- LNPs comprise from about 0.1 % to about 20% of the PEG- modified lipid on a molar basis, e.g., about 0.5 to about 15%, about 0.5 to about 10%, about 0.5 to about 5%, about 10%, about 5%, about 3.5%, about 3%, about 2,5%, about 2%, about 1.5%, about 1%, about 0.5%, or about 0.3% on a molar basis (based on 100% total moles of lipids in the LNP).
- LNPs comprise from about 1 .0% to about 2.0% of the PEG- modified lipid on a molar basis, e.g., about 1.2 to about 1.9%, about 1.2 to about 1.8%, about 1.3 to about 1 .8%, about 1.4 to about 1.8%, about 1 .5 to about 1 .8%, about 1.6 to about 1.8%, in particular about 1.4%, about 1.5%, about 1.6%, about 1.7%, about 1.8%, about 1.9%, most suitably 1.7% (based on 100% total moles of lipids in the LNP).
- the molar ratio of the cationic lipid to the PEGylated lipid ranges from about 100:1 to about 25:1.
- the LNP comprises a PEG-modified lipid at around 0.5 to 10 molar %, optionally 0.5 to 5 molar % or 0.5 to 3 molar %.
- the LNP comprises one or more additional lipids, which stabilize the formation of particles during their formulation or during the manufacturing process (e.g. neutral lipid and/or one or more steroid or steroid analogue).
- Suitable stabilizing lipids include neutral lipids and anionic lipids.
- neutral lipid refers to any one of a number of lipid species that exist in either an uncharged or neutral zwitterionic form at physiological pH.
- Representative neutral lipids include diacylphosphatidylcholines, diacylphosphatidylethanolamines, ceramides, sphingomyelins, dihydro sphingomyelins, cephalins, and cerebrosides.
- the non-cationic lipid is a neutral lipid, such as 1 ,2-distearoyl- sn-glycero-3-phosphocholine (DSPC), 1 ,2-dipalmitoyl-sn-glycero-3-phosphocholine (DPPC), 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine (POPC), 1 ,2-dioleoyl-sn-glycero-3- phosphoethanolamine (DOPE) or sphingomyelin (SM), preferably the neutral lipid is DSPC.
- DSPC 1 ,2-distearoyl- sn-glycero-3-phosphocholine
- DPPC 1 ,2-dipalmitoyl-sn-glycero-3-phosphocholine
- POPC 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine
- DOPE 1,2-dioleoyl-
- the LNP (or liposome, nanoliposome, lipoplex) comprises one or more neutral lipids, wherein the neutral lipid is selected from the group comprising distearoylphosphatidylcholine (DSPC), dioleoylphosphatidylcholine (DOPC), dipalmitoylphosphatidylcholine (DPPC), dioleoylphosphatidylglycerol (DOPG), dipalmitoylphosphatidylglycerol (DPPG), dioleoyl-phosphatidylethanolamine (DOPE), palmitoyloleoylphosphatidylcholine (POPC), palmitoyloleoyl-phosphatidylethanolamine (POPE) and dioleoyl-phosphatidylethanolamine 4-(N-maleimidomethyl)-cyclohexane- Icarboxylate (DOPE-mal), dipalmitoyl phosphatidyl ethanolamine (DPPE), dipal
- the LNPs comprise a neutral lipid selected from DSPC, DPPC, DMPC, DOPC, POPC, DOPE and SM.
- the molar ratio of the cationic lipid to the neutral lipid ranges from about 2:1 to about 8:1.
- the neutral lipid is 1 ,2-distearoyl-sn-glycero-3-phosphocholine (DSPC).
- DSPC 1 ,2-distearoyl-sn-glycero-3-phosphocholine
- the molar ratio of the cationic lipid to DSPC may be in the range from about 2:1 to about 8:1.
- the steroid is sterol, suitably cholesterol.
- the steroid is cholesterol.
- the molar ratio of the cationic lipid to cholesterol may be in the range from about 2:1 to about 1 :1.
- the cholesterol may be PEGylated.
- the sterol can be about 10mol% to about 60mol% or about 25mol% to about 55mol% or about 25mol% to about 40mol% of the lipid particle. In one embodiment, the sterol is about 10, 15, 20, 25, 30, 35, 40, 45, 50, 55, or about 60mol% of the total lipid present in the lipid particle. In another embodiment, the LNPs include from about 5% to about 50% on a molar basis of the sterol, e.g., about 15% to about 45%, about 20% to about 40%, about 48%, about 40%, about 38.5%, about 35%, about 34.4%, about 31.5% or about 31% on a molar basis (based upon 100% total moles of lipid in the lipid nanoparticle).
- the cationic lipid of an LNP may be ionizable, i.e. it becomes protonated as the pH is lowered below the pK of the ionizable group of the lipid but is progressively more neutral at higher pH values. At pH values below the pK, the lipid is then able to associate with negatively charged nucleic acids.
- the cationic lipid comprises a zwitterionic lipid that assumes a positive charge on pH decrease.
- Such cationic lipids include, but are not limited to, DSDMA, N,N-dioleyl-N,N-dimethylammonium chloride (DODAC), N,N-distearyl-N,N-dimethylammonium bromide (DDAB), 1 ,2- dioleoyltrimethyl ammonium propane chloride (DOTAP) (also known as N-(2,3- dioleoyloxy)propyl)-N,N,N-trimethylammonium chloride and 1 ,2-Dioleyloxy-3- trimethylaminopropane chloride salt), N-(1-(2,3-dioleyloxy)propyl)-N,N,N-trimethylammonium chloride (DOTMA), N,N-dimethyl-2,3-dioleyloxy)propylamine (DODMA), ckk
- DSDMA N,N-dioleyl-N,N-dimethylammonium chloride
- DODAC
- Suitable cationic lipids for use in the compositions and methods of the invention include those described in international patent publications WO2010053572 (and particularly, Cl 2-200 described at paragraph [00225]) and WO2012170930, both of which are incorporated herein by reference, HGT4003, HGT5000, HGTS001 , HGT5001 , HGT5002 (see US20150140070A1).
- the cationic lipid of the liposomes, lipid nanoparticles (LNP), lipoplexes, and/or nanoliposomes may be an amino lipid.
- Representative amino lipids include, but are not limited to, 1 ,2-dilinoleyoxy-3- (dimethylamino)acetoxypropane (DLin-DAC), 1 ,2-dilinoleyoxy-3morpholinopropane (DLin- MA), 1 ,2-dilinoleoyl-3-dimethylaminopropane (DLinDAP), 1 ,2-dilinoleylthio-3- dimethylaminopropane (DLin-S-DMA), 1-linoleoyl-2-linoleyloxy-3dimethylaminopropane (DLin-2-DMAP), 1 ,2-dilinoleyloxy-3-trimethylaminopropane chloride salt (DLin-TMA.CI), 1 ,2- dilinoleoyl-3-trimethylaminopropane chloride salt (DLin-TAP.CI), 1 ,2-dilinoleyloxy-3-(N- methylpiperazino)prop
- the cationic lipid of the liposomes, lipid nanoparticles (LNP), lipoplexes, and/or nanoliposomes may an aminoalcohol lipidoid.
- Aminoalcohol lipidoids may be prepared by the methods described in U.S. Patent No. 8,450,298, herein incorporated by reference in its entirety.
- Suitable (ionizable) lipids can also be the compounds as disclosed in Tables 1 , 2 and 3 and as defined in claims 1-24 of WO2017075531A1 , hereby incorporated by reference.
- suitable lipids can also be the compounds as disclosed in W02015074085A1 (/.e. ATX-001 to ATX-032 or the compounds as specified in claims 1-26), U.S. Appl. Nos. 61/905,724 and 15/614,499 or U.S. Patent Nos. 9,593,077 and 9,567,296 hereby incorporated by reference in their entirety.
- suitable cationic lipids can also be the compounds as disclosed in WO2017117530A1 (/.e. lipids 13, 14, 15, 16, 17, 18, 19, 20, or the compounds as specified in the claims), hereby incorporated by reference in its entirety.
- ionizable or cationic lipids may also be selected from the lipids disclosed in W02018078053A1 (/.e. lipids derived from formula I, II, and III of W02018078053A1 , or lipids as specified in Claims 1 to 12 of W02018078053A1), the disclosure of W02018078053A1 hereby incorporated by reference in its entirety.
- lipids disclosed in Table 7 of W02018078053A1 e.g. lipids derived from formula 1-1 to 1-41
- lipids disclosed in Table 8 of W02018078053A1 e.g.
- formula 11-1 to II-36 may be suitably used in the context of the invention. Accordingly, formula 1-1 to formula 1-41 and formula 11-1 to formula II-36 of W02018078053A1 , and the specific disclosure relating thereto, are herewith incorporated by reference.
- cationic lipids may be derived from formula III of published PCT patent application W02018078053A1. Accordingly, formula III of W02018078053A1 , and the specific disclosure relating thereto, are herewith incorporated by reference.
- the RNA is complexed with one or more lipids thereby forming LNPs (or liposomes, nanoliposomes, lipoplexes), wherein the cationic lipid of the LNP is selected from structures 111-1 to HI-36 of Table 9 of published PCT patent application W02018078053A1. Accordingly, formula 111-1 to HI-36 of W02018078053A1 , and the specific disclosure relating thereto, are herewith incorporated by reference.
- the ionizable cationic lipid has the formula HI: or a pharmaceutically acceptable salt, tautomer or stereoisomer thereof, wherein:
- G 1 and G 2 are each independently unsubstituted C1-C12 alkylene or C1-C12 alkenylene;
- G 3 is CI-C24 alkylene, C1-C24 alkenylene, Cs-Cs cycloalkylene, or Cs-Cs cycloalkenylene;
- R 1 and R 2 are each independently, branched or linear, C6-C24 alkyl or C6-C24 alkenyl;
- R 4 is C1-C12 alkyl
- R 5 is H or C1-C6 alkyl.
- the ionizable cationic lipid has the formula HI: or a pharmaceutically acceptable salt, tautomer or stereoisomer thereof, wherein:
- G 1 and G 2 are each independently unsubstituted C1-C12 alkylene
- G 3 is C1-C24 alkylene; R 1 and R 2 are each independently, branched or linear, C6-C24 alkyl;
- R 3 is OR 5 ;
- R 5 is H.
- the ionizable cationic lipid has the formula III and wherein R 1 , R 2 or both R 1 and R 2 have one of the following structures:
- R 2 has the structure:
- the cationic lipid has the formula:
- the ionizable cationic lipid has the formula:
- the ionizable cationic lipid has the formula 111-3:
- the lipid of formula 111-3 as suitably used herein has the chemical term ((4- hydroxybutyl)azanediyl)bis(hexane-6,1-diyl)bis(2-hexyldecanoate), also referred to as ALC- 0315 i.e. CAS Number 2036272-55-4.
- the cationic lipid as defined herein is present in the LNP in an amount from about 30 mol% to about 80 mol%, suitably about 30 mol% to about 60 mol%, more suitably about 40 mol% to about 55 mol%, more suitably about 47.4 mol%, relative to the total lipid content of the LNP. If more than one cationic lipid is incorporated within the LNP, such percentages apply to the combined cationic lipids.
- the cationic lipid as defined herein is present in the LNP in an amount from about 20 mol% to about 60 mol%.
- the LNP comprises a cationic lipid having the following structure:
- the cationic lipid is present in the LNP in an amount from about 30 mol% to about 70 mol%. In one embodiment, the cationic lipid is present in the LNP in an amount from about 40 mol% to about 60 mol%, such as about 40, 41 , 42, 43, 44, 45, 46, 47, 48, 49, 50, 51 , 52, 53, 54, 55, 56, 57, 58, 59 or 60 mol%, respectively.
- the cationic lipid is present in the LNP in an amount from about 47 mol% to about 48 mol%, such as about 47.0, 47.1 , 47.2, 47.3, 47.4, 47.5, 47.6, 47.7, 47.8, 47.9, 50.0 mol%, respectively, wherein 47.4 mol% are particularly suitable.
- the cationic lipid is present in a ratio of from about 20 mol% to about 70 mol% or 75 mol% or from about 45 mol% to about 65 mol% or about 20, 25, 30, 35, 40, 45, 50, 55, 60, 65, or about 70 mol% of the total lipid present in the LNP.
- the LNPs comprise from about 25% to about 75% on a molar basis of cationic lipid, e.g., from about 20 to about 70%, from about 35 to about 65%, from about 45 to about 65%, about 60%, about 57.5%, about 57.1 %, about 50% or about 40% on a molar basis (based upon 100% total moles of lipid in the lipid nanoparticle).
- the ratio of cationic lipid to nucleic acid, suitably RNA, more suitably mRNA is from about 3 to about 15, such as from about 5 to about 13 or from about 7 to about 11.
- Suitable (cationic or ionizable) lipids are disclosed in W02009086558, W02009127060, W02010048536, W02010054406, W02010088537, W02010129709, WO2011153493, WO 2013063468, US20110256175, US20120128760, US20120027803, US8158601 , WO2016118724, WO2016118725, W02017070613, W02017070620, WO2017099823, WO2012040184, WO2011153120, WO2011149733, WO2011090965, WO2011043913, WO2011022460, WO2012061259, WO2012054365, WO2012044638, WO2010080724, W0201021865, W02008103276, WO2013086373, WO2013086354, US Patent Nos.
- the cationic or ionizable lipid is in embodiments, amino or cationic lipids as defined herein have at least one protonatable or deprotonatable group, such that the lipid is positively charged at a pH at or below physiological pH (e.g. pH 7.4), and neutral at a second pH, suitably at or above physiological pH.
- physiological pH e.g. pH 7.4
- second pH suitably at or above physiological pH.
- Lipids having more than one protonatable or deprotonatable group, or which are zwitterionic, are not excluded and may likewise suitable in the context of the present invention.
- the protonatable lipids have a pKa of the protonatable group in the range of about 4 to about 11 , e.g., a pKa of about 5 to about 7.
- LNPs can comprise two or more (different) cationic lipids as defined herein.
- Cationic lipids may be selected to contribute to different advantageous properties.
- cationic lipids that differ in properties such as amine pKa, chemical stability, half-life in circulation, half-life in tissue, net accumulation in tissue, or toxicity can be used in the LNP (or liposomes, nanoliposomes, lipoplexes).
- the cationic lipids can be chosen so that the properties of the mixed-LNP are more desirable than the properties of a single-LNP of individual lipids.
- the amount of the permanently cationic lipid or lipidoid may be selected taking the amount of the nucleic acid cargo into account. In one embodiment, these amounts are selected such as to result in an N/P ratio of the nanoparticle(s) or of the composition in the range from about 0.1 to about 20, or
- lipid : mRNA weight ratio in the range of 20 to 60, suitably from about 3 to about 15, 5 to about 13, about 4 to about 8 or from about 7 to about 11 ; or
- the N/P ratio is defined as the mole ratio of the nitrogen atoms (“N”) of the basic nitrogen-containing groups of the lipid or lipidoid to the phosphate groups (“P”) of the nucleic acid which is used as cargo.
- the N/P ratio may be calculated on the basis that, for example, 1 pg RNA typically contains about 3 nmol phosphate residues, provided that the RNA exhibits a statistical distribution of bases.
- the “N”-value of the cationic lipid or lipidoid may be calculated on the basis of its molecular weight and the relative content of permanently cationic and - if present - cationisable groups. If more than one cationic lipid is present, the N-value should be calculated on the basis of all cationic lipids comprised in the lipid nanoparticles.
- the composition has a lipid to RNA molar ratio (N/P ratio) of about 2 to about 12, optionally a N/P ratio of 3 to about 8.
- the lipid nanoparticles comprise about 40% cationic lipid LKY750, about 10% zwitterionic lipid DSPC, about 48% cholesterol, and about 2% PEGylated lipid DMG (w/w).
- LNPs comprise: (a) the RNAs, suitably mRNAs, used herein, (b) a cationic lipid, (c) an aggregation reducing agent (such as polyethylene glycol (PEG) lipid or PEG-modified lipid), (d) optionally a non-cationic lipid (such as a neutral lipid), and (e) optionally, a sterol.
- a cationic lipid such as polyethylene glycol (PEG) lipid or PEG-modified lipid
- PEG polyethylene glycol
- a non-cationic lipid such as a neutral lipid
- sterol optionally, a sterol.
- the cationic lipids (as defined above), non-cationic lipids (as defined above), cholesterol (as defined above), and/or PEG-modified lipids (as defined above) may be combined at various relative molar ratios.
- the ratio of cationic lipid to noncationic lipid to cholesterol-based lipid to PEGylated lipid may be between about 30-60:20- 35:20-30:1-15, or at a ratio of about 40:30:25:5, 50:25:20:5, 50:27:20:3, 40:30:20:10, 40:32:20:8, 40:32:25:3 or 40:33:25:2, or at a ratio of about 50:25:20:5, 50:20:25:5, 50:27:20:3 40:30:20: 10,40:30:25:5 or 40:32:20:8, 40:32:25:3 or 40:33:25:2, respectively.
- the LNPs (or liposomes, nanoliposomes, lipoplexes) comprise ALC-0315, the RNAs, suitably mRNAs, used herein, a neutral lipid which is DSPC, a steroid which is cholesterol and a PEGylated lipid which is ALC-0159.
- the LNP comprises a PEG-modified lipid at around 0.5 to 15 molar %, a non-cationic lipid at around 5 to 25 molar %, a sterol at around 25 to 55 molar % and an ionisable cationic lipid at around 20 to 60 molar %.
- the LNP consists essentially of (i) at least one cationic lipid; (ii) a neutral lipid; (iii) a sterol, e.g. , cholesterol; and (iv) a PEG-lipid, e.g. PEG-DMG or PEG-cDMA, in a molar ratio of about 20-60% cationic lipid: 5-25% neutral lipid: 25-55% sterol; 0.5-15% PEG-lipid.
- the RNA suitably mRNA, is complexed with one or more lipids thereby forming lipid nanoparticles, wherein the LNP comprises
- At least one neutral lipid as defined herein suitably 1 ,2-distearoyl-sn-glycero-3- phosphocholine (DSPC);
- At least one polymer conjugated lipid suitably a PEG-lipid as defined herein, e.g.
- PEG-DMG or PEG-cDMA suitably a PEGylated lipid that is or is derived from formula (I a - ALC-0159).
- the mRNA is complexed with one or more lipids thereby forming lipid nanoparticles (LNP), wherein the LNP comprises (i) to (iv) in a molar ratio of about 20- 60% cationic lipid: 5-25% neutral lipid: 25-55% sterol; 0.5-15% polymer conjugated lipid, suitably PEG-lipid.
- LNP lipid nanoparticles
- the lipid nanoparticle (or liposome, nanoliposome, lipoplexe) comprises: a cationic lipid with formula (III-3) and/or PEG lipid with formula (IVa), optionally a neutral lipid, suitably 1 ,2-distearoyl-sn-glycero-3-phosphocholine (DSPC) and optionally a steroid, suitably cholesterol, wherein the molar ratio of the cationic lipid to DSPC is optionally in the range from about 2:1 to 8:1 , wherein the molar ratio of the cationic lipid to cholesterol is optionally in the range from about 2:1 to 1 :1.
- a cationic lipid with formula (III-3) and/or PEG lipid with formula (IVa) optionally a neutral lipid, suitably 1 ,2-distearoyl-sn-glycero-3-phosphocholine (DSPC) and optionally a steroid, suitably cholesterol, wherein the molar ratio of the cationic
- RNA suitably mRNA, lipid nanoparticles (LNPs)
- LNPs lipid nanoparticles
- LNPs are described in the following references: WO2012/006376; WO20 12/030901 ; WO2012/031046; WO2012/031043; WO2012/006378; WO2011/076807; WO20 13/033563; WO2013/006825; WO2014/136086; WO2015/095340; WO2015/095346; W02016/037053, which are also incorporated herein by reference.
- LNPs have a mean diameter of from about 50nm to about 200nm, from about 60nm to about 200nm, from about 70nm to about 200nm, from about 80nm to about 200nm, from about 90nm to about 200nm, from about 90nm to about 190nm, from about 90nm to about 180nm, from about 90nm to about 170nm, from about 90nm to about 160nm, from about 90nm to about 150nm, from about 90nm to about 140nm, from about 90nm to about 130nm, from about 90nm to about 120nm, from about 90nm to about 100nm, from about 70nm to about 90nm, from about 80nm to about 90nm, from about 70nm to about 80nm, or about 30nm, 35nm, 40nm, 45nm, 50nm, 55nm, 60nm, 65nm, 70nm, 75nm, 80nm, 85nm, 90nm
- the LNPs have a polydispersity of 0.4 or less, such as 0.3 or less.
- the PDI is determined by dynamic light scattering.
- the composition has a polydispersity index (PDI) value of less than about 0.4, suitably of less than about 0.3, more suitably of less than about 0.2, most suitably of less than about 0.1.
- PDI polydispersity index
- encapsulated RNA is understood as RNA (suitably mRNA) that is complexed with the lipids forming the LNP and/or that is contained within the interior space of the LNP.
- the proportion of encapsulated RNA can typically be determined using a RiboGreen assay.
- the composition contains less than about 30%, suitably less than 20%, 15%, 10% or 5% non-encapsulated RNA (or free RNA).
- free RNA or “nonencapsulated RNA” is understood as RNA (suitably mRNA) that is not encapsulated in the LNPs as defined herein.
- free RNA may represent a contamination or an impurity.
- an immunogenic composition comprising:
- said immunogenic composition further comprises (d).
- said first subtype of Influenza A virus is a subtype of Influenza A Group 1 , suitably influenza A subtype H1 , H2, H5, H6, H8, H9, H11 , H12, H13, H16, H17 or H18, more suitably H1.
- said first subtype of Influenza A virus is Influenza A H1 N1 subtype.
- said second subtype of Influenza A virus is a subtype of Influenza A Group 2, suitably influenza A subtype H3, H4, H7, H10, H14 and H15, more suitably H3.
- said second strain subtype of Influenza A virus is Influenza A H3N2 subtype.
- said first strain of Influenza B is a strain of B/Victoria lineage.
- said second strain of Influenza B is a strain of B/Yamagata lineage.
- said first, second, third and/or fourth nucleic acid is a messenger ribonucleic acid (mRNA).
- mRNA messenger ribonucleic acid
- the immunogenic composition further comprises:
- At least one further nucleic acid suitably mRNA, encoding at least one further antigen
- said at least one further antigen is derived from a strain of Influenza virus, suitably is selected from the group consisting of Influenza A virus and Influenza B virus, more suitably is selected from the group consisting of said strain of said first subtype of Influenza A virus, said strain of said second subtype of Influenza A virus, said first strain of Influenza B virus and, optionally, said second strain of Influenza B virus.
- said at least one further antigen comprises or consists of a peptide or protein selected or derived from an Influenza virus NA or an immunogenic fragment or an immunogenic variant thereof.
- the composition comprises a plurality of (e).
- the immunogenic composition further comprises:
- a fifth nucleic acid suitably an mRNA, encoding a NA of the first subtype of Influenza A virus
- a sixth nucleic acid suitably an mRNA, encoding a NA of the second subtype of Influenza A virus
- a seventh nucleic acid suitably an mRNA, encoding a NA of the first strain of Influenza B virus; and optionally
- an eighth nucleic acid suitably an mRNA, encoding a NA of the second strain of Influenza B virus.
- the mRNAs of (a), (b), (c), (d), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ) are formulated in a lipid nanoparticle (LNP), each separately or together.
- LNP lipid nanoparticle
- the LNP comprises a PEG-modified lipid, suitably at around 0.5 to 15 molar %, a non-cationic lipid, suitably at around 5 to 25 molar %, a sterol, suitably at around 25 to 55 molar %, and a cationic lipid, suitably an ionizable cationic lipid at around 20 to 60 molar %.
- the mRNAs of (a), (b), (c), (d), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ), optionally each, are not self-replicating.
- the mRNAs of (a), (b), (c), (d), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ) comprises a 5’ untranslated region (UTR), suitably the 5’ UTR comprises or consists of a nucleic acid sequence derived from a 5’-UTR of a gene selected from HSD17B4, RPL32, ASAH1 , ATP5A1 , MP68, NDUFA4, NOSIP, RPL31 , SLC7A3, TUBB4B and UBQLN2, or from a homolog, a fragment or variant of any one of these genes.
- a 5’ UTR comprises or consists of a nucleic acid sequence derived from a 5’-UTR of a gene selected from HSD17B4, RPL32, ASAH1 , ATP5A1 , MP68, NDUFA4, NOSIP, RPL31 , SLC7A3, TUBB4B and UBQLN2, or from
- the mRNAs of (a), (b), (c), (d), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ) comprises a 3’ UTR
- the 3’ UTR comprises or consists of a nucleic acid sequence derived from a 3’-UTR of a gene selected from PSMB3, ALB7, CASP1 , COX6B1 , GNAS, NDUFA1 and RPS9, or from a homolog, a fragment or a variant of any one of these genes.
- the mRNAs of (a), (b), (c), (d), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ) comprises at least one chemical modification, suitably wherein the chemical modification is N1 -methylpseudouridine and/or pseudouridine, suitably N1-methylpseudouridine.
- the invention in a second aspect, relates to a vaccine for use in the treatment or prophylaxis of an infection with an Influenza virus, comprising an immunogenic composition as defined herein, wherein an immune response is elicited as defined herein.
- the vaccine is a nucleic acid-based vaccine, suitably an mRNA-based vaccine.
- the vaccine is a multivalent vaccine.
- the vaccine is a trivalent Influenza virus vaccine (i.e. comprising immunogenic components derived from 3 strains of Influenza virus).
- the trivalent Influenza virus vaccine comprises 3 different nucleic acids, suitably mRNAs, encoding 3 HA antigens.
- the trivalent Influenza virus vaccine comprises 2 different nucleic acids, suitably mRNAs, encoding 2 HA antigens derived from a strain of Influenza A virus, suitably from H1 N1 and/or H3N2, and 1 nucleic acid, suitably mRNA, encoding 1 HA antigen derived from a strain of Influenza B virus, suitably from B/Victoria lineage.
- the trivalent Influenza virus vaccine comprises 6 different nucleic acids, suitably mRNAs, encoding 3 HA and 3 NA antigens.
- the trivalent Influenza virus vaccine comprises 4 different nucleic acids, suitably mRNAs, encoding 2 HA and 2 NA antigens derived from a strain of Influenza A virus, suitably from H1 N1 and/or H3N2, and 2 different nucleic acids, suitably mRNAs, encoding 1 HA and 1 NA antigen derived from a strain of Influenza B virus, suitably from B/Victoria lineage.
- the trivalent Influenza virus vaccine comprises (a), (b) and (c) as defined herein.
- the trivalent Influenza virus vaccine comprises (a), (b), (c), (e 1 ), (e 2 ) and (e 3 ) as defined herein.
- the vaccine is a quadrivalent Influenza virus vaccine (i.e. comprising immunogenic components derived from 4 strains of Influenza virus).
- the quadrivalent Influenza virus vaccine comprises 4 mRNAs encoding 4 HA antigens.
- the quadrivalent Influenza virus vaccine comprises 4 mRNAs encoding 4 HA antigens and 3 mRNAs encoding 3 NA antigens (i.e. seven components quadrivalent Influenza virus vaccine).
- the quadrivalent Influenza virus vaccine comprises 4 mRNAs encoding 4 HA antigens and 4 mRNAs encoding 4 NA antigens (i.e. eight components quadrivalent Influenza virus vaccine).
- the vaccine further comprises at least one antigen or at least one nucleic acid encoding said at least one antigen, such as at least one mRNA encoding an antigen from a further pathogen, suitably the pathogen being a virus, suitably a respiratory virus.
- said antigen is from further virus is selected from the group consisting of Coronavirus (e.g. SARS-CoV-1 , SARS-CoV-2, MERS-CoV), Pneumoviridae virus (e.g. Respiratory syncytial virus, Metapneumovirus) and Paramyxovidirae virus (e.g. Parainfluenza virus, Henipavirus), suitably said antigen from a further virus is a spike protein, or an antigenic fragment thereof, from a SARS-CoV-2 virus or a mRNA encoding a spike protein, or an antigenic fragment thereof, from a SARS-CoV-2 virus.
- Coronavirus e.g. SARS-CoV-1 , SARS-CoV-2, MERS-CoV
- Pneumoviridae virus e.g. Respiratory syncytial virus, Metapneumovirus
- Paramyxovidirae virus e.g. Parainfluenza virus, Henipavirus
- the antigen can be a SARS-CoV- 2 virus spike protein or an antigenic fragment thereof selected from those provided in Table 1 of published PCT application WO2021156267A1 or in Table 1 of published PCT application WO2022137133A1 , each of which is incorporated herein by reference.
- a vaccine comprising an immunogenic composition as defined herein.
- the invention relates to a kit or kit of parts for use in the treatment or prophylaxis of an infection with an Influenza virus, wherein the kit or kit of parts comprises the nucleic acids, suitably the mRNAs, as defined herein, suitably the mRNAs of (a), (b), (c), (d), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ) as defined herein, optionally comprising a liquid vehicle for solubilizing, and, optionally, technical instructions providing information on administration and dosage of the components, wherein an immune response is elicited as defined herein.
- the kit or kit of parts comprises the nucleic acids, suitably the mRNAs, as defined herein, suitably the mRNAs of (a), (b), (c), (d), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ) as defined herein, optionally comprising a liquid vehicle for solubilizing, and, optionally, technical instructions providing information
- kits suitably kits of parts, may be applied e.g. for any of the applications or uses mentioned herein, suitably for the use of the immunogenic composition or the vaccine for the treatment or prophylaxis of an infection or diseases caused by an Influenza virus, suitably Influenza A and/or B virus.
- the immunogenic composition or the vaccine is provided in a separate part of the kit, wherein the immunogenic composition or the vaccine is suitably lyophilised or spray-dried or spray-freeze dried.
- the kit may further contain as a part, a vehicle (e.g. buffer solution) for solubilising the dried or lyophilized nucleic composition or the vaccine.
- a vehicle e.g. buffer solution
- the nucleic acids and/or the mRNAs suitably the mRNAs of (a), (b), (c), (d), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ), as defined herein are formulated separately.
- the nucleic acids and/or the mRNAs suitably the mRNAs of (a), (b), (c), (d), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ), as defined herein are provided as one part of the kit.
- the nucleic acids and/or the mRNAs are each provided as a separate part of the kit.
- the kit or kit of parts comprises at least two, at least three, at least four, at least five, at least six, at least seven, at least eight parts, each containing at least one of the nucleic acids and/or the mRNAs, suitably the mRNAs (a), (b), (c), (d), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ), as defined herein.
- the kit or kit of parts as defined herein comprises a multi-dose container for administration of the composition/the vaccine and/or an administration device (e.g. an injector for intramuscular and/or intradermal injection).
- an administration device e.g. an injector for intramuscular and/or intradermal injection.
- kit or kit of parts comprising the nucleic acids, suitably the mRNAs, as defined herein, suitably the mRNAs of (a), (b), (c), (d), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ) as defined herein, optionally comprising a liquid vehicle for solubilizing, and, optionally, technical instructions providing information on administration and dosage of the components, wherein an immune response is elicited as defined herein.
- the nucleic acids suitably the mRNAs, as defined herein, suitably the mRNAs of (a), (b), (c), (d), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ) as defined herein, optionally comprising a liquid vehicle for solubilizing, and, optionally, technical instructions providing information on administration and dosage of the components, wherein an immune response is elicited as defined herein.
- the nucleic acids suitably mRNAs, as defined herein are coformulated.
- the mRNAs of (a), (b), (c), (d), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ), as defined herein are co-formulated, i.e. formulated together.
- the nucleic acids suitably mRNAs, as defined herein of the kit or kit of parts are formulated separately.
- the mRNAs of (a), (b), (c), (d), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ), as defined herein are formulated separately.
- the antigens or the nucleic acids suitably mRNAs, as defined herein are co-filled.
- the mRNAs of (a), (b), (c), (c 1 ), (c 2 ), (c 3 ), (c 4 ), (c 5 ) and/or (c 6 ), as defined herein are co-filled, i.e. filled together, optionally after being formulated separately.
- the nucleic acids suitably mRNAs, as defined herein are formulated as a bedside mixing formulation.
- the mRNAs suitably the mRNAs of (a), (b), (c), (d), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ), as defined herein are formulated as a bedside mixing formulation.
- a “bedside mixing formulation” must be understood as a formulation wherein some (such as one or more) of the immunogenic components (e.g. mRNA), suitably each, have been formulated (e.g. in LNPs) independently before being mixed to form the bedside mixing formulation.
- the immunogenic components e.g. mRNA
- LNPs LNPs
- the bedside mixing formulation is obtained by a process comprising (1) formulating (e.g. in LNPs) each nucleic acid, suitably mRNAs, independently and (2) mixing each (LN Reformulated nucleic acid, suitably mRNAs.
- the bedside mixing formulation is obtained by a process comprising (1) co-formulating (e.g. in LNPs) said nucleic acid, suitably mRNAs, encoding such derived from a strain of Influenza A virus (2) co-formulating said nucleic acid, suitably mRNAs, encoding such derived from a strain of Influenza B virus, and (3) mixing (LNP-)co-formulated nucleic acids, suitably mRNAs encoding such derived from a strain of Influenza A virus with (LNP-)co-formulated nucleic acids, suitably mRNAs encoding such derived from a strain of Influenza B virus.
- the bedside mixing formulation is obtained by a process comprising (1) co-formulating (e.g. in LNPs) said nucleic acid, suitably mRNAs, encoding such derived from a strain of Influenza A virus (2) formulating each nucleic acid, suitably mRNAs, encoding such derived from a strain of Influenza B virus independently, and (3) mixing (LNP- )co-formulated nucleic acids, suitably mRNAs encoding such derived from a strain of Influenza A virus and each (LNP-)formulated nucleic acids, suitably mRNAs encoding such derived from a strain of Influenza B virus.
- the immunogenic composition may be administered via various suitable routes, including parenteral, such as intramuscular, intradermal, intranasal, or subcutaneous administration.
- parenteral such as intramuscular, intradermal, intranasal, or subcutaneous administration.
- the immunogenic composition, the vaccine or the kit or kit of parts as described herein is administered intramuscularly and/or intradermally.
- intramuscular administration of the immunogenic composition as described herein results in expression of the encoded antigen construct in a subject.
- Administration of the immunogenic composition as described herein results in translation of the mRNA and to a production of the encoded antigen in a subject.
- the immunogenic composition described herein may be provided in liquid or dry (e.g. lyophilised) form.
- the immunogenic composition is provided in liquid form.
- the immunogenic composition is a dry composition.
- dry composition may be a composition that has been lyophilized (e.g. according to WO2016165831 , WO2011069586, WO2022/232585, W02022/101461 , WO2022/076562, WO2012/170889 or W02022/036170), or spray-dried, or freeze-dried (e.g. according to WO2016184575, WO2016184576 or WO2021/216541) to obtain a dry composition, suitably a temperature stable composition, for example in the form of a powder.
- a dry composition suitably a temperature stable composition, for example in the form of a powder.
- the composition according to the invention is a lyophilized, freeze- dried or spray-dried dry composition comprising one or more further excipients selected from cryoprotectants, plasticizers and polymers.
- the lyophilized, freeze-dried or spray- dried dry composition is mixed with a liquid, suitably an aqueous liquid such as sterile water or saline, to form a “reconstituted liquid formulation” prior to administration to a patient.
- lyophilized or spray-dried composition has a water content of less than about 10%.
- lyophilized, freeze-dried or spray-dried composition has a water content of between about 0.5% and 5%.
- the lyophilized, freeze-dried or spray-dried composition is stable for at least 2 months after storage at about 5 °C, suitably for at least 3 months, 4 months, 5 months, 6 months.
- Liquids used for reconstitution will be substantially aqueous, such as water for injection, phosphate buffered saline and the like.
- Buffers may be selected from acetate, citrate, histidine, maleate, phosphate, succinate, tartrate and TRIS.
- the buffer may be a phosphate buffer such as Na/Na2PO4, Na/K2PO4 or K/K2PO4.
- the formulations used in the present invention have a dose volume of between 0.05 ml and 1 ml, such as between 0.1 and 0.6 ml, in particular a dose volume of 0.45 to 0.55 ml, such as 0.5 ml.
- the volumes of the compositions used may depend on the subject, delivery route and location, with smaller doses being given by the intradermal route.
- a typical human dose for administration through routes such as intramuscular is in the region of 200 pl to 750 pl, such as 400 to 600 pl, in particular about 500 pl, such as 500 pl.
- the immunogenic composition as described herein may be provided in various physical containers such as vials or pre-filled syringes.
- the immunogenic composition is provided in the form of a single dose. In other embodiments, the immunogenic composition, the vaccine or the kit or kit of parts is provided in multidose form such containing 2, 5 or 10 doses.
- overage It is common where liquids are to be transferred between containers, such as from a vial to a syringe, to provide ‘an overage’ which ensures that the full volume required can be conveniently transferred.
- the level of overage required will depend on the circumstances, but excessive overage should be avoided to reduce wastage and insufficient overage may cause practical difficulties. Overages may be of the order of 20 to 100 pl per dose, such as 30 pl or 50 pl.
- Stabilisers may be present. Stabilisers may be of particular relevance where multidose containers are provided as doses of the final formulation(s) may be administered to subjects over a period of time.
- Formulations are suitably sterile.
- Approaches for establishing strong and lasting immunity often include repeated immunisation, i.e. boosting an immune response by administration of one or more further doses.
- the immunogenic composition as described herein may therefore be part of a multi-dose administration regime.
- the immunogenic composition as described herein may be provided as a priming dose in a multidose regime, especially a two- or three-dose regime, in particular a two-dose regime.
- the immunogenic composition as described herein may be provided as a boosting dose in a multidose regime, especially a two- or three-dose regime, such as a two-dose regime.
- the time between doses may be two weeks to six months, such as three weeks to three months. Periodic longer-term booster doses may also be provided, such as every 2 to 10 years.
- the immunogenic composition further comprises at least one pharmaceutically acceptable carrier.
- the term “pharmaceutically acceptable carrier” or “pharmaceutically acceptable excipient” as used herein suitably includes the liquid or non-liquid basis of the composition for administration.
- the carrier may be water, e.g. pyrogen-free water; isotonic saline or buffered (aqueous) solutions, e.g. phosphate, citrate etc. buffered solutions.
- Water or suitably a buffer, more suitably an aqueous buffer may be used, containing a sodium salt, suitably at least 50mM of a sodium salt, a calcium salt, suitably at least 0.01 mM of a calcium salt, and optionally a potassium salt, suitably at least 3mM of a potassium salt.
- the sodium, calcium and, optionally, potassium salts may occur in the form of their halogenides, e.g. chlorides, iodides, or bromides, in the form of their hydroxides, carbonates, hydrogen carbonates, or sulfates, etc.
- sodium salts include NaCI, Nal, NaBr, Na2COs, NaHCOs, Na2SC
- examples of the optional potassium salts include KCI, KI, KBr, K2CO3, KHCO3, K2SO4
- examples of calcium salts include CaCh, Cal2, CaBr2, CaCOs, CaSC , Ca(OH)2.
- the immunogenic composition may comprise pharmaceutically acceptable carriers or excipients using one or more pharmaceutically acceptable carriers or excipients to e.g. increase stability, increase cell transfection, permit the sustained or delayed, increase the translation of encoded antigenic peptides or proteins in vivo, and/or alter the release profile of encoded antigenic peptides or proteins protein in vivo.
- excipients can include, without limitation, lipidoids, liposomes, lipid nanoparticles, polymers, lipoplexes, core-shell nanoparticles, peptides, proteins, cells transfected with polynucleotides, hyaluronidase, nanoparticle mimics and combinations thereof.
- one or more compatible solid or liquid fillers or diluents or encapsulating compounds may be used as well, which are suitable for administration to a subject.
- compatible means that the constituents of the composition are capable of being mixed with the at least one nucleic acid of component A and/or component B and, optionally, a plurality of nucleic acids of the composition, in such a manner that no interaction occurs, which would substantially reduce the biological activity or the pharmaceutical effectiveness of the composition under typical use conditions (e.g., intramuscular or intradermal administration).
- Pharmaceutically acceptable carriers or excipients must have sufficiently high purity and sufficiently low toxicity to make them suitable for administration to a subject to be treated.
- Compounds which may be used as pharmaceutically acceptable carriers or excipients may be sugars, such as, for example, lactose, glucose, trehalose, mannose, and sucrose; starches, such as, for example, corn starch or potato starch; dextrose; cellulose and its derivatives, such as, for example, sodium carboxymethylcellulose, ethylcellulose, cellulose acetate; powdered tragacanth; malt; gelatin; tallow; solid glidants, such as, for example, stearic acid, magnesium stearate; calcium sulfate; vegetable oils, such as, for example, groundnut oil, cottonseed oil, sesame oil, olive oil, corn oil and oil from theobroma; polyols, such as, for example, polypropylene glycol, glycerol, sorbitol, mannitol and polyethylene glycol; alginic acid.
- sugars such as, for example, lactose, glucose, tre
- Subjects to which administration of the immunogenic compositions is contemplated include, but are not limited to, humans and/or other primates; mammals, including commercially relevant mammals such as cattle, pigs, horses, sheep, cats, dogs, mice, and/or rats; and/or birds, including commercially relevant birds such as poultry, chickens, ducks, geese, and/or turkeys.
- the immunogenic composition does not exceed a certain proportion of free RNA, suitably mRNA.
- free RNA suitably mRNA or “non-complexed RNA, suitably mRNA” or “non-encapsulated RNA, suitably mRNA” comprise the RNA, suitably mRNA molecules that are not encapsulated in the lipid-based carriers as defined herein.
- free RNA suitably mRNA may represent a contamination or an impurity.
- the immunogenic composition comprises free RNA, suitably mRNA, ranging from about 30% to about 0%.
- the composition comprises about 20% free RNA, suitably mRNA (and about 80% encapsulated RNA, suitably mRNA), about 15% free RNA, suitably mRNA (and about 85% encapsulated RNA, suitably mRNA), about 10% free RNA, suitably mRNA (and about 90% encapsulated RNA, suitably mRNA), or about 5% free RNA, suitably mRNA (and about 95% encapsulated RNA, suitably mRNA).
- the composition comprises less than about 20% free RNA, suitably mRNA, suitably less than about 15% free RNA, suitably mRNA, more suitably less than about 10% free RNA, suitably mRNA, most suitably less than about 5% free RNA, suitably mRNA.
- encapsulated RNA comprises the RNA, suitably mRNA, molecules that are encapsulated in the lipid-based carriers as defined herein.
- the proportion of encapsulated RNA, suitably mRNA, in the context of the invention is typically determined using a RiboGreen assay.
- a single dose of the immunogenic composition is 0.1 to 1000 pg, especially 1 to 500 pg, especially 2 to 500 pg, in particular 10 to 250 pg, suitably 10 to 150 pg of total mRNA.
- a single dose of the immunogenic composition is 10 to 150 pg, optionally 20 to 100 pg, optionally 35 to 100 pg of total mRNA.
- a single dose of the immunogenic composition is 35, 36, 37, 38, 39, 40, 41 , 42, 43, 44, 45, 46, 47, 48, 49, 50, 55, 60, 61 , 62, 63, 64, 65, 66, 67, 68, 69, 70, 75, 80, 85, 90, 95, 96, 97, 98, 99 or 100 pg of total mRNA.
- a single dose of the immunogenic composition is 39, 45, 63, 69 or 99 pg of total mRNA.
- a single dose of the immunogenic composition comprises a mixture of 2, 3, 4, 5, 6, 7, 8, 9 or 10 different mRNA and is 1 to 200 pg, suitably 1 to 60 pg, suitably 1 to 25 pg, suitably 2 to 25 pg, suitably 3 to 18 pg of each mRNA.
- a single dose of the immunogenic composition comprises a mixture of 2, 3, 4, 5, 6, 7, 8, 9 or 10, suitably of 6, different mRNA and is 0.5 to 100 pg, suitably 0.5 to 75 pg, suitably 2 to 75 pg of each mRNA.
- a single dose of the composition is 2 to 500 pg, especially 10 to 250 pg of total mRNA, such as 10 to 75 pg of total mRNA.
- a single dose of the immunogenic composition is 10 to 100 pg of total mRNA.
- a single dose of the composition is 6, 12, 15, 16, 18, 24, 32, 36, 48, 54, 60, 72, 84, 96 or 120 pg of total mRNA.
- a single dose of the composition is 1 to 10 pg of each mRNA for younger adult e.g. 18 to 64 years old.
- a single dose of the composition is 1 , 2, 3, 6 or 9 pg of each mRNA for younger adult e.g. 18 to 64 years old. In some embodiments, a single dose of the composition is 15 to 50 pg of total mRNA for younger adult e.g. 18 to 64 years old.
- a single dose of the composition is 16, 32 or 48 pg of total mRNA for younger adult e.g. 18 to 64 years old.
- a single dose of the composition is 0.5 to 50 pg of each mRNA for younger adult e.g. 18 to 64 years old.
- a single dose of the composition is 0.5, 1 , 2, 3, 4, 5, 6, 7, 8, 9, 10, 11 , 12, 13, 14, 15, 20, 21 , 22, 23, 24, 25, 30, 35, 36, 37, 38, 39, 40, 45, 46, 47, 48, 49 or 50 pg of each mRNA for younger adult e.g. 18 to 64 years old.
- a single dose of the composition is 25 to 75 pg of total mRNA for younger adult e.g. 18 to 64 years old.
- a single dose of the composition is 25, 30, 35, 36, 37, 38, 39, 40, 41 , 42, 43, 44, 45, 46, 47, 48, 49, 50, 55, 60, 65, 66, 67, 68, 69, 70 or 75 pg of total mRNA for younger adult e.g. 18 to 64 years old.
- a single dose of the composition is 2 to 20 pg of each mRNA for older adult e.g. 65 years old and above.
- a single dose of the composition is 2, 3, 6, 9 or 18 pg of each mRNA for older adult e.g. 65 years old and above.
- a single dose of the composition is 30 to 100 pg of total mRNA for older adult e.g. 65 years old and above.
- a single dose of the composition is 32, 48 or 96 pg of total mRNA for older adult e.g. 65 years old and above.
- a single dose of the composition is 0.5 to 100 pg of each mRNA for older adult e.g. 65 years old and above.
- a single dose of the composition is 0.5, 1 , 2, 3, 4, 5, 6, 7, 8, 9, 10, 11 , 12, 13, 14, 15, 20, 21 , 22, 23, 24, 25, 30, 35, 36, 37, 38, 39, 40, 45, 46, 47, 48, 49, 50, 55, 60, 65, 70, 71 , 72, 73, 74, 75, 80, 85, 90, 95 or 100 pg of each mRNA for older adult e.g. 65 years old and above.
- a single dose of the composition is 35 to 150 pg of total mRNA for older adult e.g. 65 years old and above. In some embodiments, a single dose of the composition is 35, 40, 41 , 42, 43, 44, 45, 46, 47, 48, 49, 50, 55, 60, 61 , 62, 63, 64, 65, 66, 67, 68, 69, 70, 75, 80, 85, 90, 95, 96, 97, 98, 99, 100, 110, 120, 130, 140 or 150 pg of total mRNA for older adult e.g. 65 years old and above.
- the nucleic acids and/or the mRNAs suitably the mRNAs of (a), (b), (c), (d), (e 1 ), (e 2 ), (e 3 ) and/or (e 4 ) as described herein are administered at different sites of injection.
- the nucleic acids and/or the mRNAs derived from a strain of Influenza A virus are administered at a site of injection which is different to the site of injection where the nucleic acids and/or the mRNAs derived from a strain of Influenza B virus are administered.
- the nucleic acids and/or the mRNAs derived from a strain of Influenza B virus are administered separately, suitably at different sites of injection.
- the invention in a fourth aspect, relates to a method of eliciting an immune response against an Influenza virus, wherein the method comprises applying or administering to a subject in need thereof an immunogenic composition as defined herein, and wherein an immune response is elicited as defined herein.
- the elicited immune response is an adaptative immune response, suitably a protective adaptative immune response against an Influenza virus, suitably against an Influenza A virus and/or an Influenza B virus.
- the elicited immune response comprises functional antibodies that can effectively neutralize the respective viruses.
- the elicited immune response is a cross-reactive immune response, wherein the functional antibodies that can effectively neutralize the respective viruses further neutralize viruses belonging to same and other Influenza A subtypes and/or Influenza B lineages.
- the cross-reactive immune response is homologous, heterologous and/or heterosubtypic.
- the term “homologous” in the context of an elicited immune response will be recognized and understood by the person of ordinary skill in the art, and is e.g. an immune response which is elicited against the same strain, such as the same Influenza A strain or the same Influenza B strain.
- the immunogenic composition may comprise a HA antigen (or nucleic acid, suitably RNA, suitably mRNA, encoding such) derived from A/Michigan/45/2015 (H1 N1 pdm9) which may elicit an immune response against A/Michigan/45/2015 (H1 N1 pdm9) strain.
- heterologous or “intrasubtypic” in the context of an elicited immune response will be recognized and understood by the person of ordinary skill in the art, and is e.g. an immune response which is elicited against different strains within a subtype (for Influenza A virus) or lineage (for Influenza B virus), such as different Influenza A strains within a subtype such as H1 or H3 subtypes.
- a subtype for Influenza A virus
- lineage for Influenza B virus
- the immunogenic composition may comprise a HA antigen (or nucleic acid, suitably RNA, suitably mRNA, encoding such) derived from A/Michigan/45/2015 (H1 N1 pdm9) which may elicit an immune response against A/New Caledonia/20/1999 (H1 N1) strain.
- a HA antigen or nucleic acid, suitably RNA, suitably mRNA, encoding such
- the term “heterosubtypic” in the context of an elicited immune response will be recognized and understood by the person of ordinary skill in the art, and is e.g. an immune response which is elicited against different strains within one or more different subtypes (for Influenza A virus) or lineages (for Influenza B virus).
- the immunogenic composition may comprise a HA antigen (or nucleic acid, suitably RNA, suitably mRNA, encoding such) derived from A/Michigan/45/2015 (H1 N1pdm9) which may elicit an immune response against HongKong/4801/2014 (H3N2).
- an immune response is elicited against at least three, four, five, six, seven, eight, nine, ten, eleven, twelve, thirteen, fourteen, fifteen HA antigens of at least three, four, five, six, seven, eight, nine, ten subtypes of Influenza A virus, including said first and second subtypes of Influenza A virus and/or at least three four, five, six, seven, eight, nine, ten, eleven, twelve, thirteen, fourteen, fifteen HA antigens of Influenza B virus, including against HA antigens of said first and second strains of Influenza B virus.
- an immune response is elicited against at least one HA antigen of Influenza A from each of subtypes H1 , H2, H3, H5, H7 and H10, at least one HA antigen of Influenza B from B/Victoria lineage and at least one strain of Influenza B from B/Yamagata lineage.
- the elicited immune response comprises broad, functional cellular T-cell responses against the respective viruses.
- the elicited immune response comprises a CD4+ T cell immune response and/or a CD8+ T cell immune response.
- the elicited immune response comprises a well-balanced B cell and T cell response against the respective viruses.
- the elicited immune response comprises antigen-specific immune responses.
- the elicited immune response reduces partially or completely the severity of one or more symptoms and/or time over which one or more symptoms of Influenza virus infection are experienced by the subject.
- the elicited immune response reduces the likelihood of developing an established Influenza virus infection after challenge.
- the elicited immune response slows progression of Influenza, suitably Influenza A and/or B.
- the subject in need is a mammalian subject, suitably a human subject.
- composition, the vaccine or the kit or kit of parts as described herein is administered in an amount effective to induce a T cell response against Influenza A subtypes selected from the group consisting of H1 N1 , H2N2, H3N2, H5N1 , H7N9, H10N8 and any combination thereof.
- composition, the vaccine or the kit or kit of parts as described herein is administered in an amount effective to induce a T cell response against Influenza A subtypes selected from the group consisting of H1 N1 , H2N2, H3N2, H5N1 , H7N9, H10N8 and any combination thereof.
- composition, the vaccine or the kit or kit of parts as described herein is administered in an amount effective to induce a T cell response against Influenza B lineages selected from Influenza B/Yamagata lineage, Influenza B/Victoria lineage and any combination thereof.
- the composition, the vaccine or the kit or kit of parts as described is administered in an amount effective to induce a neutralizing antibody response against Influenza A subtypes selected from the group consisting of H1 N1 , H2N2, H3N2, H5N1 , H7N9, H10N8 and any combination thereof.
- the composition, the vaccine or the kit or kit of parts as described is administered in an amount effective to induce a neutralizing antibody response against Influenza B lineages selected from Influenza B/Yamagata lineage, Influenza B/Victoria lineage and any combination thereof.
- an immune response is elicited against HA antigens that are antigenically distinct from any of the HA antigen encoded by a nucleic acid present in the compositions.
- an immune response is elicited against NA antigens that are antigenically distinct from any of the NA antigen encoded by a nucleic acid present in the compositions.
- an immune response is elicited against HA and NA antigens that are antigenically distinct from any of the HA and NA antigen encoded by a nucleic acid present in the compositions.
- an immune response is elicited against Influenza viruses with a different geographical origin and/or year of isolation than any of the strains having an HA and/or NA encoded by an mRNA present in the composition.
- administration of the immunogenic composition, the vaccine or the kit or kit to a subject elicits neutralizing antibodies and does not elicit disease enhancing antibodies.
- administration of the immunogenic composition, the vaccine or the kit or kit to a subject does not elicit immunopathological effects, like e.g. enhanced disease and/or antibody dependent enhancement (ADE).
- ADE antibody dependent enhancement
- the invention relates to a method of treating or preventing a disorder caused by an Influenza virus, suitably an Influenza A and/or Influenza B, wherein the method comprises applying or administering to a subject in need thereof an immunogenic composition as defined herein, and wherein an immune response is elicited as defined herein.
- Preventing (Inhibiting) or treating a disease, in particular a virus infection relates to inhibiting the full development of a disease or condition, for example, in a subject who is at risk for a disease such as a virus infection.
- Treatment refers to a therapeutic intervention that ameliorates a sign or symptom of a disease or pathological condition after it has begun to develop.
- the term “ameliorating”, with reference to a disease or pathological condition, refers to any observable beneficial effect of the treatment.
- Inhibiting a disease can include preventing or reducing the risk of the disease, such as preventing or reducing the risk of viral infection.
- the beneficial effect can be evidenced, for example, by a delayed onset of clinical symptoms of the disease in a susceptible subject, a reduction in severity of some or all clinical symptoms of the disease, a slower progression of the disease, a reduction in the viral load, an improvement in the overall health or well-being of the subject, or by other parameters that are specific to the particular disease.
- a “prophylactic” treatment is a treatment administered to a subject who does not exhibit signs of a disease or exhibits only early signs for the purpose of decreasing the risk of developing pathology.
- composition, the vaccine or the kit or kit of parts is administered at a therapeutically effective amount.
- the subject in need is a mammalian subject, suitably a human subject.
- composition “comprising” and variants thereof such as “comprises” are to be interpreted as including the stated element (e.g., integer) or elements (e.g., integers) without necessarily excluding any other elements (e.g., integers).
- a composition “comprising” X may consist exclusively of X or may include something additional e.g. X + Y.
- a process comprising a step of mixing two or more components does not require any specific order of mixing.
- components can be mixed in any order. Where there are three components then two components can be combined with each other, and then the combination may be combined with the third component, etc.
- immunogenic fragment or “immunogenic variant” has to be understood as any fragment/variant of the corresponding Influenza antigen that is capable of raising an immune response in a subject. Percentages in the context of numbers should be understood as relative to the total number of the respective items. In other cases, and unless the context dictates otherwise, percentages should be understood as percentages by weight (wt.-%).
- determinants or values do not need to be identical, i.e. 100% the same. Accordingly, “about” means, that a determinant or values may diverge by 1 % to 20%, for example by 1 % to 10%; in particular, by 1%, 2%, 3%, 4%, 5%, 6%, 7%, 8%, 9%, 10%, 11%, 12%, 13%, 14%, 15%, 16%, 17%, 18%, 19%, 20%.
- certain parameters or determinants can slightly vary based on the method how the parameter has been determined. For example, if a certain determinants or value is defined herein to have e.g.
- a length of “about 100 nucleotides” the length may diverge by 1 % to 20%. Accordingly, the skilled person knows that in that specific example, the length may diverge by 1 to 20 nucleotides. Accordingly, a length of “about 100 nucleotides” may encompass sequences ranging from 80 to 120 nucleotides.
- Adaptive immune response The term “adaptive immune response” as used herein will be recognized and understood by the person of ordinary skill in the art, and is e.g. intended to refer to an antigen-specific response of the immune system (the adaptive immune system). Antigen specificity allows for the generation of responses that are tailored to specific pathogens or pathogen-infected cells. The ability to mount these tailored responses is usually maintained in the body by “memory cells” (B-cells).
- B-cells memory cells
- Antigen as used herein will be recognized and understood by the person of ordinary skill in the art, and is e.g. intended to refer to a substance which may be recognized by the immune system, for example by the adaptive immune system, and is capable of triggering an antigen-specific immune response, e.g. by formation of antibodies and/or antigen-specific T cells as part of an adaptive immune response.
- an antigen may be or may comprise a peptide or protein which may be presented by the MHC to T-cells. Also fragments, variants and derivatives of peptides or proteins comprising at least one epitope are understood as antigens.
- Antigenic peptide, polypeptide or protein The term “antigenic peptide or protein” or “immunogenic peptide or protein” will be recognized and understood by the person of ordinary skill in the art, and is e.g. intended to refer to a peptide, protein derived from a (antigenic or immunogenic) protein which stimulates the body’s adaptive immune system to provide an adaptive immune response. Therefore an antigenic/immunogenic peptide or protein comprises at least one epitope (as defined herein) or antigen (as defined herein) of the protein it is derived from.
- cationic means that the respective structure bears a positive charge, either permanently or not permanently, but in response to certain conditions such as pH.
- cationic covers both “permanently cationic” and “cationisable”.
- permanently cationic means, e.g., that the respective compound, or group, or atom, is positively charged at any pH value or hydrogen ion activity of its environment. Typically, the positive charge results from the presence of a quaternary nitrogen atom.
- Cationisable means that a compound, or group or atom, is positively charged at a lower pH and uncharged at a higher pH of its environment. Also in non-aqueous environments where no pH value can be determined, a cationisable compound, group or atom is positively charged at a high hydrogen ion concentration and uncharged at a low concentration or activity of hydrogen ions. It depends on the individual properties of the cationisable or polycationisable compound, in particular the pKa of the respective cationisable group or atom, at which pH or hydrogen ion concentration it is charged or uncharged.
- the fraction of cationisable compounds, groups or atoms bearing a positive charge may be estimated using the so-called Henderson- Hasselbalch equation which is well-known to a person skilled in the art.
- a compound or moiety is cationisable, it is suitable that it is positively charged at a pH value of about 1 to 9, preferably 4 to 9, 5 to 8 or even 6 to 8, for example of a pH value of or below 9, of or below 8, of or below 7, for example at physiological pH values, e.g. about 7.3 to 7.4, i.e. under physiological conditions, particularly under physiological salt conditions of the cell in vivo.
- the cationisable compound or moiety is predominantly neutral at physiological pH values, e.g. about 7.0-7.4, but becomes positively charged at lower pH values.
- the range of pKa for the cationisable compound or moiety is about 5 to about 7.
- Coding sequence/coding region The terms “coding sequence” or “coding region” and the corresponding abbreviation “cds” as used herein will be recognized and understood by the person of ordinary skill in the art, and are e.g. intended to refer to a sequence of several nucleotide triplets, which may be translated into a peptide or protein.
- a coding sequence in the context of the present invention may be an RNA sequence consisting of a number of nucleotides that may be divided by three, which starts with a start codon and which for example terminates with a stop codon.
- nucleic acid derived from (another) nucleic acid
- nucleic acid which is derived from (another) nucleic acid, shares e.g. at least 60%, 70%, 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91 %, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity with the nucleic acid from which it is derived.
- sequence identity is typically calculated for the same types of nucleic acids, i.e.
- RNA sequences for DNA sequences or for RNA sequences.
- a DNA is “derived from” an RNA or if an RNA is “derived from” a DNA
- the RNA sequence in a first step the RNA sequence is converted into the corresponding DNA sequence (in particular by replacing the uracils (II) by thymines (T) throughout the sequence) or, vice versa, the DNA sequence is converted into the corresponding RNA sequence (in particular by replacing the T by II throughout the sequence).
- sequence identity of the DNA sequences or the sequence identity of the RNA sequences is determined.
- nucleic acid “derived from” a nucleic acid also refers to nucleic acid, which is modified in comparison to the nucleic acid from which it is derived, e.g. in order to increase RNA stability even further and/or to prolong and/or increase protein production.
- amino acid sequences e.g. antigenic peptides or proteins
- derived from means that the amino acid sequence, which is derived from (another) amino acid sequence, shares e.g.
- Epitope The term “epitope” (also called “antigen determinant” in the art) as used herein will be recognized and understood by the person of ordinary skill in the art, and is e.g. intended to refer to T cell epitopes and B cell epitopes.
- T cell epitopes or parts of the antigenic peptides or proteins and may comprise fragments preferably having a length of about 6 to about 20 or even more amino acids, e.g. fragments as processed and presented by MHC class I molecules, preferably having a length of about 8 to about 10 amino acids, e.g.
- B cell epitopes are typically fragments located on the outer surface of (native) protein or peptide antigens, preferably having 5 to 15 amino acids, more preferably having 5 to 12 amino acids, even more preferably having 6 to 9 amino acids, which may be recognized by antibodies, i.e. in their native form.
- epitopes of proteins or peptides may furthermore be selected from any of the herein mentioned variants of such proteins or peptides.
- epitopes can be conformational or discontinuous epitopes which are composed of segments of the proteins or peptides as defined herein that are discontinuous in the amino acid sequence of the proteins or peptides as defined herein but are brought together in the three-dimensional structure or continuous or linear epitopes which are composed of a single polypeptide chain.
- Fragment The term “fragment” as used throughout the present specification in the context of a nucleic acid sequence (e.g. RNA or a DNA) or an amino acid sequence may typically be a shorter portion of a full-length sequence of e.g.
- a fragment typically consists of a sequence that is identical to the corresponding stretch within the full-length sequence.
- a particular fragment of a sequence in the context of the present invention consists of a continuous stretch of entities, such as nucleotides or amino acids corresponding to a continuous stretch of entities in the molecule the fragment is derived from, which represents at least 40%, 50%, 60%, 70%, 80%, 90%, 95% of the total (i.e. full-length) molecule from which the fragment is derived (e.g. a virus protein).
- fragment as used throughout the present specification in the context of proteins or peptides may, typically, comprise a sequence of a protein or peptide as defined herein, which is, with regard to its amino acid sequence, N-terminally and/or C-terminally truncated compared to the amino acid sequence of the original protein.
- fragment as used throughout the present specification in the context of RNA sequences may, typically, comprise an RNA sequence that is 5’-terminally and/or 3’-terminally truncated compared to the reference RNA sequence. Such truncation may thus occur either on the amino acid level or correspondingly on the nucleic acid level.
- a sequence identity with respect to such a fragment as defined herein may therefore for example refer to the entire protein or peptide as defined herein or to the entire (coding) nucleic acid molecule of such a protein or peptide. Fragments of proteins or peptides may comprise at least one epitope of those proteins or peptides.
- heterologous refers to a sequence (e.g. RNA, DNA, amino acid) has to be understood as a sequence that is derived from another gene, another allele, or e.g. another species or virus.
- Two sequences are typically understood to be “heterologous” if they are not derivable from the same gene or from the same allele. I.e., although heterologous sequences may be derivable from the same organism or virus, in nature, they do not occur in the same nucleic acid or protein.
- Humoral immune response The terms “humoral immunity” or “humoral immune response” will be recognized and understood by the person of ordinary skill in the art, and are e.g. intended to refer to B-cell mediated antibody production and optionally to accessory processes accompanying antibody production.
- a humoral immune response may be typically characterized, e.g. by Th2 activation and cytokine production, germinal center formation and isotype switching, affinity maturation and memory cell generation.
- Humoral immunity may also refer to the effector functions of antibodies, which include pathogen and toxin neutralization, classical complement activation, and opsonin promotion of phagocytosis and pathogen elimination.
- Identity (of a sequence): The term “identity” as used throughout the present specification in the context of a nucleic acid sequence or an amino acid sequence will be recognized and understood by the person of ordinary skill in the art, and is e.g. intended to refer to the percentage to which two sequences are identical. To determine the percentage to which two sequences are identical, e.g. nucleic acid sequences or amino acid (aa) sequences as defined herein, for example the aa sequences encoded by the nucleic acid sequence as defined herein or the aa sequences themselves, the sequences can be aligned in order to be subsequently compared to one another. Therefore, e.g. a position of a first sequence may be compared with the corresponding position of the second sequence.
- a position in the first sequence is occupied by the same residue as is the case at a position in the second sequence, the two sequences are identical at this position. If this is not the case, the sequences differ at this position. If insertions occur in the second sequence in comparison to the first sequence, gaps can be inserted into the first sequence to allow a further alignment. If deletions occur in the second sequence in comparison to the first sequence, gaps can be inserted into the second sequence to allow a further alignment. The percentage to which two sequences are identical is then a function of the number of identical positions divided by the total number of positions including those positions which are only occupied in one sequence. The percentage to which two sequences are identical can be determined using an algorithm, e.g. an algorithm integrated in the BLAST program.
- Immunogen The terms “immunogen” or “immunogenic” will be recognized and understood by the person of ordinary skill in the art, and are e.g. intended to refer to a compound that is able to stimulate/induce an (adaptive) immune response.
- An immunogen may be a peptide, polypeptide, or protein.
- Immune response The term “immune response” will be recognized and understood by the person of ordinary skill in the art, and is e.g. intended to refer to a specific reaction of the adaptive immune system to a particular antigen (so called specific or adaptive immune response) or an unspecific reaction of the innate immune system (so called unspecific or innate immune response), or a combination thereof.
- innate immune system also known as non-specific or unspecific immune system
- innate immune system will be recognized and understood by the person of ordinary skill in the art, and is e.g. intended to refer to a system typically comprising the cells and mechanisms that defend the host from infection by other organisms in a non-specific manner. This means that the cells of the innate system may recognize and respond to pathogens in a generic way, but unlike the adaptive immune system, it does not confer long-lasting or protective immunity to the host.
- the innate immune system may be activated by ligands of pattern recognition receptor e.g. Toll-like receptors, NOD-like receptors, or RIG-1 like receptors etc.
- Lipidoid compound A lipidoid compound, also simply referred to as lipidoid, is a lipid- like compound, i.e. an amphiphilic compound with lipid-like physical properties. In the context of the present invention, the term lipid is considered to encompass lipidoid compounds.
- nucleic acid, nucleic acid molecule The terms “nucleic acid” or “nucleic acid molecule” as used herein, will be recognized and understood by the person of ordinary skill in the art.
- the term is used synonymously with the term polynucleotide.
- a nucleic acid or a nucleic acid molecule is a polymer comprising or consisting of nucleotide monomers that are covalently linked to each other by phosphodiester-bonds of a sugar/phosphate-backbone.
- the terms “nucleic acid” or “nucleic acid molecule” also encompasses modified nucleic acid (molecules), such as base-modified, sugar-modified or backbone-modified DNA or RNA (molecules) as defined herein.
- Nucleic acid sequence, DNA sequence, RNA sequence The terms “nucleic acid sequence”, “DNA sequence”, “RNA sequence” will be recognized and understood by the person of ordinary skill in the art, and e.g. refer to a particular and individual order of the succession of its nucleotides.
- Permanently cationic The term “permanently cationic” as used herein will be recognized and understood by the person of ordinary skill in the art, and means, e.g., that the respective compound, or group, or atom, is positively charged at any pH value or hydrogen ion activity of its environment. Typically, the positive charge results from the presence of a quaternary nitrogen atom. Where a compound carries a plurality of such positive charges, it may be referred to as permanently polycationic.
- Stabilized RNA refers to an RNA that is modified such, that it is more stable to disintegration or degradation, e.g., by environmental factors or enzymatic digest, such as by exo- or endonuclease degradation, compared to an RNA without such modification.
- a stabilized RNA in the context of the present invention is stabilized in a cell, such as a prokaryotic or eukaryotic cell, preferably in a mammalian cell, such as a human cell.
- the stabilization effect may also be exerted outside of cells, e.g. in a buffer solution etc., e.g., for storage of a composition comprising the stabilized RNA.
- cellular immunity or “cellular immune response” or “cellular T-cell responses” as used herein will be recognized and understood by the person of ordinary skill in the art, and are for example intended to refer to the activation of macrophages, natural killer cells (NK), antigen-specific cytotoxic T-lymphocytes, and the release of various cytokines in response to an antigen.
- cellular immunity is not based on antibodies, but on the activation of cells of the immune system.
- a cellular immune response may be characterized e.g. by activating antigen-specific cytotoxic T-lymphocytes that are able to induce apoptosis in cells, e.g. specific immune cells like dendritic cells or other cells, displaying epitopes of foreign antigens on their surface.
- RNA is the usual abbreviation for ribonucleic acid. It is a nucleic acid molecule, i.e. a polymer consisting of nucleotide monomers. These nucleotides are usually adenosine-monophosphate (AMP), uridine-monophosphate (UMP), guanosinemonophosphate (GMP) and cytidine-monophosphate (CMP) monomers or analogs thereof, which are connected to each other along a so-called backbone. The backbone is typically formed by phosphodiester bonds between the sugar, i.e. ribose, of a first and a phosphate moiety of a second, adjacent monomer.
- AMP adenosine-monophosphate
- UMP uridine-monophosphate
- GMP guanosinemonophosphate
- CMP cytidine-monophosphate
- RNA sequence The specific order of the monomers, i.e. the order of the bases linked to the sugar/phosphate-backbone, is called the RNA sequence.
- RNA can be obtained by transcription of a DNA sequence, e.g. inside a cell or in vitro.
- the RNA may be obtained by RNA in vitro transcription.
- RNA may be obtained by chemical synthesis.
- RNA in vitro transcription or “in vitro transcription” relate to a process wherein RNA is synthesized in a cell-free system in vitro.
- RNA may be obtained by DNA-dependent in vitro transcription of an appropriate DNA template, which is typically a linear DNA template (e.g. linearized plasmid DNA or PCR product).
- the promoter for controlling RNA in vitro transcription can be any promoter for any DNA-dependent RNA polymerase.
- DNA-dependent RNA polymerases are the T7, T3, SP6, or Syn5 RNA polymerases.
- the DNA template is linearized with a suitable restriction enzyme before it is subjected to RNA in vitro transcription.
- Reagents typically used in RNA in vitro transcription include: a DNA template (linearized plasmid DNA or PCR product) with a promoter sequence that has a high binding affinity for its respective RNA polymerase such as bacteriophage-encoded RNA polymerases (T7, T3, SP6, or Syn5); ribonucleotide triphosphates (NTPs) for the four bases (adenine, cytosine, guanine and uracil); optionally, a cap analogue as defined herein; optionally, modified nucleotides as defined herein; a DNA-dependent RNA polymerase capable of binding to the promoter sequence within the DNA template (e.g.
- RNA polymerase T7, T3, SP6, or Syn5 RNA polymerase
- RNase ribonuclease
- MgCh a buffer
- T7, T3, SP6, or Syn5 RNA polymerase optionally, a ribonuclease (RNase) inhibitor to inactivate any potentially contaminating RNase
- RNase ribonuclease
- MgCh a buffer
- TMS or HEPES to maintain a suitable pH value, which can also contain antioxidants (e.g. DTT), and/or polyamines such as spermidine.
- Variant of a sequence:
- the term “variant” as used throughout the present specification in the context of a nucleic acid sequence will be recognized and understood by the person of ordinary skill in the art, and is e.g. intended to refer to a variant of a nucleic acid sequence derived from another nucleic acid sequence.
- a variant of a nucleic acid sequence may exhibit one or more nucleotide deletions, insertions, additions and/or substitutions compared to the nucleic acid sequence from which the variant is derived.
- a variant of a nucleic acid sequence may at least 50%, 60%, 70%, 80%, 90%, or 95% identical to the nucleic acid sequence the variant is derived from.
- the variant is a functional variant in the sense that the variant has retained at least 50%, 60%, 70%, 80%, 90%, or 95% or more of the function of the sequence where it is derived from.
- a “variant” of a nucleic acid sequence may have at least 70%, 75%, 80%, 85%, 90%, 95%, 98% or 99% nucleotide identity over a stretch of at least 10, 20, 30, 50, 75 or 100 nucleotides of such nucleic acid sequence.
- variant as used throughout the present specification in the context of proteins or peptides is e.g. intended to refer to a proteins or peptide variant having an amino acid sequence which differs from the original sequence in one or more mutation(s)/substitution(s), such as one or more substituted, inserted and/or deleted amino acid(s).
- these fragments and/or variants have the same, or a comparable specific antigenic property (immunogenic variants, antigenic variants). Insertions and substitutions are possible, in particular, at those sequence positions which cause no modification to the three-dimensional structure or do not affect the binding region. Modifications to a three-dimensional structure by insertion(s) or deletion(s) can easily be determined e.g.
- a “variant” of a protein or peptide may have at least 70%, 75%, 80%, 85%, 90%, 95%, 98% or 99% amino acid identity over a stretch of at least 10, 20, 30, 50, 75 or 100 amino acids of such protein or peptide.
- a “variant” of a protein or polypeptide may have from 1 to 20, for example from 1 to 10 single amino acid mutations compared to such protein or peptide, for example, 1 , 2, 3, 4, 5, 6, 7, 8, 9, 10, 11 , 12, 13, 15, 16, 17, 18, 19 or 20 single amino acid mutations.
- mutations we mean or include substitution, insertion or deletion.
- a variant of a protein comprises a functional variant of the protein, which means, in the context of the invention, that the variant exerts essentially the same, or at least 40%, 50%, 60%, 70%, 80%, 90% of the immunogenicity as the protein it is derived from.
- Multivalent vaccine/composition the multivalent vaccine or combination of the invention provides more than one valence (e.g. an antigen) derived from more than one virus (e.g. at least one Influenza virus as defined herein and at least one further Influenza virus as defined herein).
- valence e.g. an antigen
- virus e.g. at least one Influenza virus as defined herein and at least one further Influenza virus as defined herein.
- Embodiment 1 An immunogenic composition for use in the treatment or prophylaxis of an infection with an Influenza virus, wherein the immunogenic composition comprises:
- a fourth nucleic acid encoding a HA antigen of a second strain of Influenza B virus, wherein an immune response is elicited against HA antigens of said strains of first and second subtypes of Influenza A virus, said first and, optionally, second strains of Influenza B virus and at least one further HA antigen subtype of Influenza A virus, being different from any of the HA antigen subtypes of Influenza A virus encoded by a nucleic acid present in the composition.
- Embodiment 2 The immunogenic composition for use according to embodiment 1 , wherein the composition further comprises (d) said fourth nucleic acid encoding a HA antigen of a second strain of Influenza B virus and wherein an immune response is further elicited against said HA antigen of said second strain of Influenza B virus.
- Embodiment 3 The immunogenic composition for use according to embodiment 1 or 2, wherein said first and/or second subtype of Influenza A virus is selected from influenza A viruses characterized by a HA selected from the group consisting of H1 , H2, H3, H4, H5, H6, H7, H8, H9, H10, H11 , H12, H13, H14, H15, H16, H17 and H18, suitably from the group consisting of H1 , H3, H5, H7, H9, and H10, more suitably from the group consisting of H1 and H3.
- a HA selected from the group consisting of H1 , H2, H3, H4, H5, H6, H7, H8, H9, H10, H11 , H12, H13, H14, H15, H16, H17 and H18, suitably from the group consisting of H1 , H3, H5, H7, H9, and H10, more suitably from the group consisting of H1 and H3.
- Embodiment 4 The immunogenic composition for use according to any of embodiments 1 to 4.
- said first and/or second subtype of Influenza A virus is selected from influenza A viruses characterized by a neuraminidase (NA) selected from the group consisting of N1 , N2, N3, N4, N5, N6, N7, N8, N9, N10, and N11 , suitably selected from the group consisting of N1 , N2, and N8, more suitably selected from the group consisting of N1 and N2.
- NA neuraminidase
- Embodiment 5 The immunogenic composition for use according to any of embodiments 1 to 4.
- said first and/or second subtype of Influenza A virus is selected from the group consisting of H1 N1 , H1 N2, H2N2, H3N1 , H3N2, H3N8, H5N1 , H5N2, H5N3, H5N8, H5N9, H7N1 , H7N2, H7N3, H7N4, H7N7, H7N9, H9N2, H10N7 and H10N8, suitably H1 N1 and H3N2.
- Embodiment 6 The immunogenic composition for use according to any of embodiments 1 to 4.
- said first subtype of Influenza A virus is a subtype of Influenza A Group 1 , suitably influenza A subtype H1 , H2, H5, H6, H8, H9, H11 , H12, H13, H16, H17 or H18, more suitably H1.
- Embodiment 7 The immunogenic composition for use according to any of embodiments 1 to 4.
- Embodiment 8 The immunogenic composition for use according to any of embodiments 1 to 4.
- said second subtype of Influenza A virus is a subtype of Influenza A Group 2, suitably influenza A subtype H3, H4, H7, H10, H14 and H15, more suitably H3.
- Embodiment 9 The immunogenic composition for use according to any of embodiments 1 to 4.
- Embodiment 10 The immunogenic composition for use according to any of embodiments 1 to 4.
- strain of first and/or second subtype of Influenza A virus is selected from the group consisting of A/Thailand/8/2022 (H3N2)-like virus, A/Massachusetts/18/2022 (H3N2)- like virus, AA/ictoria/4897/2022 (H1 N1)pdmO9-like virus, A/Wisconsin/67/2022 (H1 N1)pdmO9- like virus, A/Sydney/5/2021 (H1 N1)pdmO9-like virus, A/Victoria/2570/2019 (H1 N1)pdmO9-like virus, A/Darwin/9/2021 (H3N2)-like virus, A/Wisconsin/588/2019 (H1 N1)pdmO9-like virus, A/Darwin/6/2021 (H3N2)-like virus, A/Cambodia/e0826360/2020 (H3N2)-like virus, A/Guangdong-Maonan/SWL1536
- Embodiment 11 The immunogenic composition for use according to any of embodiments 1 to 4.
- said first and/or second strain of Influenza B virus is selected from the group consisting of B/Victoria lineage and B/Yamagata lineage.
- Embodiment 12 The immunogenic composition for use according to any of embodiments 1 to 4.
- said first strain of Influenza B is a strain of B/Victoria lineage.
- Embodiment 13 The immunogenic composition for use according to any of embodiments 1 to 4.
- Embodiment 14 The immunogenic composition for use according to any of embodiments 1 to 4.
- said first strain of Influenza B virus is selected from the group consisting of B/Austria/1359417/2021 (B/Victoria lineage)-like virus, B/Washington/02/2019 (B/Victoria lineage)-like virus, B/Colorado/06/2017-like virus (B/Victoria/2/87 lineage), B/Brisbane/60/2008-like virus and B/Colorado/06/2019 (B/Victoria lineage)-like virus.
- Embodiment 15 The immunogenic composition for use according to any of embodiments 1 to 14.
- said second strain of Influenza B is B/Phuket/3073/2013 (B/Yamagata lineage)- like virus.
- Embodiment 16 The immunogenic composition for use according to any of embodiments 1 to 16.
- said first subtype of Influenza A virus is of Influenza A H1 N1
- said second subtype of Influenza A virus is of influenza A H3N2
- said first strain of Influenza B virus is of influenza B/Yamagata lineage
- said second strain of Influenza B virus is of Influenza B/Victoria lineage.
- Embodiment 17 The immunogenic composition for use according to any of embodiments 1 to 4.
- the elicited immune response is homologous, heterosubtypic, and optionally heterologous or intrasubtypic.
- Embodiment 18 The immunogenic composition for use according to any of embodimentl to
- said at least one further HA antigen subtype of Influenza A virus is selected from influenza A viruses characterized by a HA selected from the group consisting of H1 , H2, H3, H4, H5, H6, H7, H8, H9, H10, H11 , H12, H13, H14, H15, H16, H17 and H18, suitably from the group consisting of H1 , H2, H3, H5, H7 and H10, more suitably from the group consisting of H2, H5, H7 and H10.
- Embodiment 19 The immunogenic composition for use according to any of embodiments 1 to 4.
- said at least one further HA antigen subtype of Influenza A virus is selected from influenza A viruses characterized by a neuraminidase (NA) selected from the group consisting of N1 , N2, N3, N4, N5, N6, N7, N8, N9, N10, and N11 , suitably selected from the group consisting of N1 , N2, N8 and N9.
- NA neuraminidase
- Embodiment 20 The immunogenic composition for use according to ant of embodiments 1 to
- said at least one further HA antigen subtype of Influenza A virus is selected from the group consisting of H1 N1 , H1 N2, H2N2, H3N1 , H3N2, H3N8, H5N1 , H5N2, H5N3, H5N8, H5N9, H7N1 , H7N2, H7N3, H7N4, H7N7, H7N9, H9N2, H10N7 and H10N8, suitably H1 N1 , H2N2, H3N2, H5N1 , H7N9 and H10N8.
- Embodiment 21 The immunogenic composition for use according to any of embodiments 1 to
- said at least one further HA antigen subtype of Influenza A virus is from Influenza A Group 1 , suitably influenza A subtype H1 , H2, H5, H6, H8, H9, H11 , H12, H13, H16, H17 or H18, more suitably H1 , H2 or H5.
- Embodiment 22 The immunogenic composition for use according to any of embodiments 1 to 4.
- said at least one further HA antigen subtype of Influenza A virus is from Influenza A Group 2, suitably influenza A subtype H3, H4, H7, H10, H14 or H15, more suitably H3, H7 or H10.
- Embodiment 23 The immunogenic composition for use according to any of embodiments 1 to
- Embodiment 24 The immunogenic composition for use according to any of embodiments 1 to 23, wherein an immune response is further elicited against at least one further HA antigen of a strain of Influenza B virus, being different from any of the HA antigens of a strain of Influenza B virus encoded by a nucleic acid present in the composition.
- Embodiment 25 The immunogenic composition for use according to embodiment 24, wherein said at least one further HA antigen of a strain of Influenza B virus is derived from a strain selected from the group consisting of BA/ictoria lineage and B/Yamagata lineage.
- Embodiment 26 The immunogenic composition for use according to embodiment 24 or 25, wherein said at least one further HA antigen of a strain of Influenza B virus is a strain of BA/ictoria lineage.
- Embodiment 27 The immunogenic composition for use according to any of embodiments 24 to 26, wherein said at least one further HA antigen of a strain of Influenza B virus is a strain of B/Yamagata lineage.
- Embodiment 28 The immunogenic composition for use according to any of embodiments 24 to 27, wherein said at least one further HA antigen of a strain of Influenza B virus is selected from the group consisting of B/Austria/1359417/2021 (B/Victoria lineage)-like virus, B/Washington/02/2019 (BA/ictoria lineage)-like virus, B/Colorado/06/2017-like virus (B/Victoria/2/87 lineage), B/Brisbane/60/2008-like virus, B/Colorado/06/2019 (BA/ictoria lineage)-like virus, B/lllinois/NHRC_FDX51486/2015, B/Oman/4241/2019, B/lllinois/NHRC_18512/2017, B/California/BRD12452N/2017, B/lndia/Pun-1922338/2019, B/Japan/8858/2019, B/Washington/02/2019, B/Stockholm
- Embodiment 29 The immunogenic composition for use according to any of embodiments 1 to
Landscapes
- Health & Medical Sciences (AREA)
- Life Sciences & Earth Sciences (AREA)
- Virology (AREA)
- General Health & Medical Sciences (AREA)
- Medicinal Chemistry (AREA)
- Veterinary Medicine (AREA)
- Public Health (AREA)
- Chemical & Material Sciences (AREA)
- Animal Behavior & Ethology (AREA)
- Pharmacology & Pharmacy (AREA)
- Chemical Kinetics & Catalysis (AREA)
- Nuclear Medicine, Radiotherapy & Molecular Imaging (AREA)
- Organic Chemistry (AREA)
- General Chemical & Material Sciences (AREA)
- Molecular Biology (AREA)
- Pulmonology (AREA)
- Oncology (AREA)
- Communicable Diseases (AREA)
- Immunology (AREA)
- Microbiology (AREA)
- Mycology (AREA)
- Epidemiology (AREA)
- Medicines Containing Antibodies Or Antigens For Use As Internal Diagnostic Agents (AREA)
Abstract
Description
Claims
Priority Applications (2)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| AU2024260120A AU2024260120A1 (en) | 2023-04-27 | 2024-04-25 | Influenza virus vaccines |
| CN202480028133.1A CN121001738A (en) | 2023-04-27 | 2024-04-25 | Influenza vaccine |
Applications Claiming Priority (4)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| GBGB2306220.1A GB202306220D0 (en) | 2023-04-27 | 2023-04-27 | Influenza virus vaccines |
| GB2306220.1 | 2023-04-27 | ||
| GB2314629.3 | 2023-09-25 | ||
| GBGB2314629.3A GB202314629D0 (en) | 2023-09-25 | 2023-09-25 | Influenza virus vaccines |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| WO2024223728A1 true WO2024223728A1 (en) | 2024-10-31 |
Family
ID=90924479
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/EP2024/061356 Pending WO2024223728A1 (en) | 2023-04-27 | 2024-04-25 | Influenza virus vaccines |
Country Status (3)
| Country | Link |
|---|---|
| CN (1) | CN121001738A (en) |
| AU (1) | AU2024260120A1 (en) |
| WO (1) | WO2024223728A1 (en) |
Citations (117)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US4373071A (en) | 1981-04-30 | 1983-02-08 | City Of Hope Research Institute | Solid-phase synthesis of polynucleotides |
| US4458066A (en) | 1980-02-29 | 1984-07-03 | University Patents, Inc. | Process for preparing polynucleotides |
| US4500707A (en) | 1980-02-29 | 1985-02-19 | University Patents, Inc. | Nucleosides useful in the preparation of polynucleotides |
| US4668777A (en) | 1981-03-27 | 1987-05-26 | University Patents, Inc. | Phosphoramidite nucleoside compounds |
| US4973679A (en) | 1981-03-27 | 1990-11-27 | University Patents, Inc. | Process for oligonucleo tide synthesis using phosphormidite intermediates |
| US5047524A (en) | 1988-12-21 | 1991-09-10 | Applied Biosystems, Inc. | Automated system for polynucleotide synthesis and purification |
| US5132418A (en) | 1980-02-29 | 1992-07-21 | University Patents, Inc. | Process for preparing polynucleotides |
| US5153319A (en) | 1986-03-31 | 1992-10-06 | University Patents, Inc. | Process for preparing polynucleotides |
| US5262530A (en) | 1988-12-21 | 1993-11-16 | Applied Biosystems, Inc. | Automated system for polynucleotide synthesis and purification |
| US5700642A (en) | 1995-05-22 | 1997-12-23 | Sri International | Oligonucleotide sizing using immobilized cleavable primers |
| WO2002098443A2 (en) | 2001-06-05 | 2002-12-12 | Curevac Gmbh | Stabilised mrna with an increased g/c content and optimised codon for use in gene therapy |
| WO2008016473A2 (en) | 2006-07-28 | 2008-02-07 | Applera Corporation | Dinucleotide mrna cap analogs |
| WO2008077592A1 (en) | 2006-12-22 | 2008-07-03 | Curevac Gmbh | Method for purifying rna on a preparative scale by means of hplc |
| US7404969B2 (en) | 2005-02-14 | 2008-07-29 | Sirna Therapeutics, Inc. | Lipid nanoparticle based compositions and methods for the delivery of biologically active molecules |
| WO2008103276A2 (en) | 2007-02-16 | 2008-08-28 | Merck & Co., Inc. | Compositions and methods for potentiated activity of biologicaly active molecules |
| WO2008157688A2 (en) | 2007-06-19 | 2008-12-24 | Board Of Supervisors Of Louisiana State University And Agricultural And Mechanical College | Synthesis and use of anti-reverse phosphorothioate analogs of the messenger rna cap |
| WO2009086558A1 (en) | 2008-01-02 | 2009-07-09 | Tekmira Pharmaceuticals Corporation | Improved compositions and methods for the delivery of nucleic acids |
| WO2009127060A1 (en) | 2008-04-15 | 2009-10-22 | Protiva Biotherapeutics, Inc. | Novel lipid formulations for nucleic acid delivery |
| WO2009149253A2 (en) | 2008-06-06 | 2009-12-10 | Uniwersytet Warszawski | Mrna cap analogs |
| US20100036115A1 (en) | 1997-07-23 | 2010-02-11 | Sirna Therapeutics, Inc. | Novel Compositions for the Delivery of Negatively Charged Molecules |
| WO2010021865A1 (en) | 2008-08-18 | 2010-02-25 | Merck Sharp & Dohme Corp. | Novel lipid nanoparticles and novel components for delivery of nucleic acids |
| WO2010048536A2 (en) | 2008-10-23 | 2010-04-29 | Alnylam Pharmaceuticals, Inc. | Processes for preparing lipids |
| WO2010054406A1 (en) | 2008-11-10 | 2010-05-14 | Alnylam Pharmaceuticals, Inc. | Novel lipids and compositions for the delivery of therapeutics |
| WO2010053572A2 (en) | 2008-11-07 | 2010-05-14 | Massachusetts Institute Of Technology | Aminoalcohol lipidoids and uses thereof |
| WO2010080724A1 (en) | 2009-01-12 | 2010-07-15 | Merck Sharp & Dohme Corp. | Novel lipid nanoparticles and novel components for delivery of nucleic acids |
| WO2010088537A2 (en) | 2009-01-29 | 2010-08-05 | Alnylam Pharmaceuticals, Inc. | Improved lipid formulation |
| WO2010129709A1 (en) | 2009-05-05 | 2010-11-11 | Alnylam Pharmaceuticals, Inc. | Lipid compositions |
| US20100324120A1 (en) | 2009-06-10 | 2010-12-23 | Jianxin Chen | Lipid formulation |
| WO2011015347A1 (en) | 2009-08-05 | 2011-02-10 | Biontech Ag | Vaccine composition comprising 5'-cap modified rna |
| US7893302B2 (en) | 2005-02-14 | 2011-02-22 | Sirna Therapeutics, Inc. | Lipid nanoparticle based compositions and methods for the delivery of biologically active molecules |
| WO2011022460A1 (en) | 2009-08-20 | 2011-02-24 | Merck Sharp & Dohme Corp. | Novel cationic lipids with various head groups for oligonucleotide delivery |
| WO2011043913A2 (en) | 2009-10-08 | 2011-04-14 | Merck Sharp & Dohme Corp. | Novel cationic lipids with short lipid chains for oligonucleotide delivery |
| WO2011069586A1 (en) | 2009-12-09 | 2011-06-16 | Curevac Gmbh | Mannose-containing solution for lyophilization, transfection and/or injection of nucleic acids |
| WO2011076807A2 (en) | 2009-12-23 | 2011-06-30 | Novartis Ag | Lipids, lipid compositions, and methods of using them |
| WO2011090965A1 (en) | 2010-01-22 | 2011-07-28 | Merck Sharp & Dohme Corp. | Novel cationic lipids for oligonucleotide delivery |
| US20110256175A1 (en) | 2008-10-09 | 2011-10-20 | The University Of British Columbia | Amino lipids and methods for the delivery of nucleic acids |
| WO2011149733A2 (en) | 2010-05-24 | 2011-12-01 | Merck Sharp & Dohme Corp. | Novel amino alcohol cationic lipids for oligonucleotide delivery |
| WO2011153120A1 (en) | 2010-06-04 | 2011-12-08 | Merck Sharp & Dohme Corp. | Novel low molecular weight cationic lipids for oligonucleotide delivery |
| WO2011153493A2 (en) | 2010-06-03 | 2011-12-08 | Alnylam Pharmaceuticals, Inc. | Biodegradable lipids for the delivery of active agents |
| WO2012006378A1 (en) | 2010-07-06 | 2012-01-12 | Novartis Ag | Liposomes with lipids having an advantageous pka- value for rna delivery |
| WO2012006376A2 (en) | 2010-07-06 | 2012-01-12 | Novartis Ag | Virion-like delivery particles for self-replicating rna molecules |
| WO2012019780A1 (en) | 2010-08-13 | 2012-02-16 | Curevac Gmbh | Nucleic acid comprising or coding for a histone stem-loop and a poly(a) sequence or a polyadenylation signal for increasing the expression of an encoded protein |
| WO2012031046A2 (en) | 2010-08-31 | 2012-03-08 | Novartis Ag | Lipids suitable for liposomal delivery of protein-coding rna |
| WO2012030901A1 (en) | 2010-08-31 | 2012-03-08 | Novartis Ag | Small liposomes for delivery of immunogen-encoding rna |
| WO2012031043A1 (en) | 2010-08-31 | 2012-03-08 | Novartis Ag | Pegylated liposomes for delivery of immunogen-encoding rna |
| WO2012040184A2 (en) | 2010-09-20 | 2012-03-29 | Merck Sharp & Dohme Corp. | Novel low molecular weight cationic lipids for oligonucleotide delivery |
| WO2012044638A1 (en) | 2010-09-30 | 2012-04-05 | Merck Sharp & Dohme Corp. | Low molecular weight cationic lipids for oligonucleotide delivery |
| WO2012054365A2 (en) | 2010-10-21 | 2012-04-26 | Merck Sharp & Dohme Corp. | Novel low molecular weight cationic lipids for oligonucleotide delivery |
| WO2012061259A2 (en) | 2010-11-05 | 2012-05-10 | Merck Sharp & Dohme Corp. | Novel low molecular weight cyclic amine containing cationic lipids for oligonucleotide delivery |
| US20120202871A1 (en) | 2009-07-01 | 2012-08-09 | Protiva Biotherapeutics, Inc. | Cationic lipids and methods for the delivery of therapeutic agents |
| US8283333B2 (en) | 2009-07-01 | 2012-10-09 | Protiva Biotherapeutics, Inc. | Lipid formulations for nucleic acid delivery |
| WO2012170930A1 (en) | 2011-06-08 | 2012-12-13 | Shire Human Genetic Therapies, Inc | Lipid nanoparticle compositions and methods for mrna delivery |
| WO2012170889A1 (en) | 2011-06-08 | 2012-12-13 | Shire Human Genetic Therapies, Inc. | Cleavable lipids |
| WO2013006825A1 (en) | 2011-07-06 | 2013-01-10 | Novartis Ag | Liposomes having useful n:p ratio for delivery of rna molecules |
| WO2013033563A1 (en) | 2011-08-31 | 2013-03-07 | Novartis Ag | Pegylated liposomes for delivery of immunogen-encoding rna |
| US20130064894A1 (en) | 2011-08-31 | 2013-03-14 | Protiva Biotherapeutics, Inc. | Novel cationic lipids and methods of use thereof |
| WO2013059475A1 (en) | 2011-10-18 | 2013-04-25 | Life Technologies Corporation | Alkynyl-derivatized cap analogs, preparation and uses thereof |
| WO2013063468A1 (en) | 2011-10-27 | 2013-05-02 | Massachusetts Institute Of Technology | Amino acid derivates functionalized on the n- terminal capable of forming drug incapsulating microspheres |
| US20130129785A1 (en) | 2010-05-10 | 2013-05-23 | Alnylam Pharmaceuticals, Inc | Methods and compositions for delivery of active agents |
| WO2013086354A1 (en) | 2011-12-07 | 2013-06-13 | Alnylam Pharmaceuticals, Inc. | Biodegradable lipids for the delivery of active agents |
| WO2013086373A1 (en) | 2011-12-07 | 2013-06-13 | Alnylam Pharmaceuticals, Inc. | Lipids for the delivery of active agents |
| US8466122B2 (en) | 2010-09-17 | 2013-06-18 | Protiva Biotherapeutics, Inc. | Trialkyl cationic lipids and methods of use thereof |
| WO2013143700A2 (en) | 2012-03-27 | 2013-10-03 | Curevac Gmbh | Artificial nucleic acid molecules comprising a 5'top utr |
| US20140039032A1 (en) | 2011-12-12 | 2014-02-06 | Kyowa Hakko Kirin Co., Ltd. | Lipid nano particles comprising cationic lipid for drug delivery system |
| WO2014136086A1 (en) | 2013-03-08 | 2014-09-12 | Novartis Ag | Lipids and lipid compositions for the delivery of active agents |
| WO2014152673A1 (en) | 2013-03-14 | 2014-09-25 | Shire Human Genetic Therapies, Inc. | Quantitative assessment for cap efficiency of messenger rna |
| WO2014152659A1 (en) | 2013-03-14 | 2014-09-25 | Shire Human Genetic Therapies, Inc. | Quantitative assessment for cap efficiency of messenger rna |
| WO2015062738A1 (en) | 2013-11-01 | 2015-05-07 | Curevac Gmbh | Modified rna with decreased immunostimulatory properties |
| US20150140070A1 (en) | 2013-10-22 | 2015-05-21 | Shire Human Genetic Therapies, Inc. | Lipid formulations for delivery of messenger rna |
| WO2015074085A1 (en) | 2013-11-18 | 2015-05-21 | Arcturus Therapeutics, Inc. | Ionizable cationic lipid for rna delivery |
| WO2015095346A1 (en) | 2013-12-19 | 2015-06-25 | Novartis Ag | Lipids and lipid compositions for the delivery of active agents |
| WO2015095340A1 (en) | 2013-12-19 | 2015-06-25 | Novartis Ag | Lipids and lipid compositions for the delivery of active agents |
| WO2015101416A1 (en) | 2013-12-30 | 2015-07-09 | Curevac Gmbh | Methods for rna analysis |
| WO2015188933A1 (en) | 2014-06-10 | 2015-12-17 | Curevac Ag | Methods and means for enhancing rna production |
| WO2015199952A1 (en) | 2014-06-25 | 2015-12-30 | Acuitas Therapeutics Inc. | Novel lipids and lipid nanoparticle formulations for delivery of nucleic acids |
| WO2016022914A1 (en) | 2014-08-08 | 2016-02-11 | Moderna Therapeutics, Inc. | Compositions and methods for the treatment of ophthalmic diseases and conditions |
| WO2016037053A1 (en) | 2014-09-05 | 2016-03-10 | Novartis Ag | Lipids and lipid compositions for the delivery of active agents |
| WO2016107877A1 (en) | 2014-12-30 | 2016-07-07 | Curevac Ag | Artificial nucleic acid molecules |
| WO2016118724A1 (en) | 2015-01-21 | 2016-07-28 | Moderna Therapeutics, Inc. | Lipid nanoparticle compositions |
| WO2016118725A1 (en) | 2015-01-23 | 2016-07-28 | Moderna Therapeutics, Inc. | Lipid nanoparticle compositions |
| WO2016165831A1 (en) | 2015-04-17 | 2016-10-20 | Curevac Ag | Lyophilization of rna |
| WO2016174271A1 (en) | 2015-04-30 | 2016-11-03 | Curevac Ag | Immobilized poly(n)polymerase |
| WO2016180430A1 (en) | 2015-05-08 | 2016-11-17 | Curevac Ag | Method for producing rna |
| WO2016184575A1 (en) | 2015-05-20 | 2016-11-24 | Curevac Ag | Dry powder composition comprising long-chain rna |
| WO2016184576A2 (en) | 2015-05-20 | 2016-11-24 | Curevac Ag | Dry powder composition comprising long-chain rna |
| WO2016193226A1 (en) | 2015-05-29 | 2016-12-08 | Curevac Ag | Method for adding cap structures to rna using immobilized enzymes |
| WO2016193206A1 (en) | 2015-05-29 | 2016-12-08 | Curevac Ag | A method for producing and purifying rna, comprising at least one step of tangential flow filtration |
| WO2017036580A1 (en) | 2015-08-28 | 2017-03-09 | Curevac Ag | Artificial nucleic acid molecules |
| WO2017053297A1 (en) | 2015-09-21 | 2017-03-30 | Trilink Biotechnologies, Inc. | Compositions and methods for synthesizing 5'-capped rnas |
| WO2017066797A1 (en) | 2015-10-16 | 2017-04-20 | Modernatx, Inc. | Trinucleotide mrna cap analogs |
| WO2017066782A1 (en) | 2015-10-16 | 2017-04-20 | Modernatx, Inc. | Hydrophobic mrna cap analogs |
| WO2017066793A1 (en) | 2015-10-16 | 2017-04-20 | Modernatx, Inc. | Mrna cap analogs and methods of mrna capping |
| WO2017066781A1 (en) | 2015-10-16 | 2017-04-20 | Modernatx, Inc. | Mrna cap analogs with modified phosphate linkage |
| WO2017066791A1 (en) | 2015-10-16 | 2017-04-20 | Modernatx, Inc. | Sugar substituted mrna cap analogs |
| WO2017066789A1 (en) | 2015-10-16 | 2017-04-20 | Modernatx, Inc. | Mrna cap analogs with modified sugar |
| WO2017070613A1 (en) | 2015-10-22 | 2017-04-27 | Modernatx, Inc. | Human cytomegalovirus vaccine |
| WO2017070620A2 (en) | 2015-10-22 | 2017-04-27 | Modernatx, Inc. | Broad spectrum influenza virus vaccine |
| WO2017075531A1 (en) | 2015-10-28 | 2017-05-04 | Acuitas Therapeutics, Inc. | Novel lipids and lipid nanoparticle formulations for delivery of nucleic acids |
| WO2017099823A1 (en) | 2015-12-10 | 2017-06-15 | Modernatx, Inc. | Compositions and methods for delivery of therapeutic agents |
| WO2017112865A1 (en) | 2015-12-22 | 2017-06-29 | Modernatx, Inc. | Compounds and compositions for intracellular delivery of agents |
| WO2017109161A1 (en) | 2015-12-23 | 2017-06-29 | Curevac Ag | Method of rna in vitro transcription using a buffer containing a dicarboxylic acid or tricarboxylic acid or a salt thereof |
| WO2017117530A1 (en) | 2015-12-30 | 2017-07-06 | Arcturus Therapeutics, Inc. | Ionizable cationic lipid |
| WO2018075827A1 (en) | 2016-10-19 | 2018-04-26 | Arcturus Therapeutics, Inc. | Trinucleotide mrna cap analogs |
| WO2018078053A1 (en) | 2016-10-26 | 2018-05-03 | Curevac Ag | Lipid nanoparticle mrna vaccines |
| JP2019008858A (en) | 2017-06-27 | 2019-01-17 | 花王株式会社 | Polishing liquid composition |
| WO2019077001A1 (en) | 2017-10-19 | 2019-04-25 | Curevac Ag | Novel artificial nucleic acid molecules |
| WO2019092153A1 (en) | 2017-11-08 | 2019-05-16 | Curevac Ag | Rna sequence adaptation |
| WO2020002525A1 (en) | 2018-06-27 | 2020-01-02 | Curevac Ag | Novel lassa virus rna molecules and compositions for vaccination |
| WO2021156267A1 (en) | 2020-02-04 | 2021-08-12 | Curevac Ag | Coronavirus vaccine |
| WO2021216541A1 (en) | 2020-04-20 | 2021-10-28 | Board Of Regents, The University Of Texas System | Biologically active dry powder compositions and method of their manufacture and use |
| WO2022036170A1 (en) | 2020-08-14 | 2022-02-17 | Arcturus Therapeutics, Inc. | Method of lyophilizing lipid nanoparticles |
| WO2022076562A1 (en) | 2020-10-06 | 2022-04-14 | Translate Bio, Inc. | Improved process and formulation of lipid nanoparticles |
| US20220142923A1 (en) * | 2020-11-06 | 2022-05-12 | Sanofi | LIPID NANOPARTICLES FOR DELIVERING mRNA VACCINES |
| WO2022101461A1 (en) | 2020-11-16 | 2022-05-19 | BioNTech SE | Enhanced formulation stabilization and improved lyophilization processes |
| WO2022137133A1 (en) | 2020-12-22 | 2022-06-30 | Curevac Ag | Rna vaccine against sars-cov-2 variants |
| WO2022232585A1 (en) | 2021-04-29 | 2022-11-03 | Modernatx, Inc. | Lyophilization methods for preparing lipid formulated therapeutics |
| WO2022264109A1 (en) * | 2021-06-18 | 2022-12-22 | Sanofi | Multivalent influenza vaccines |
-
2024
- 2024-04-25 WO PCT/EP2024/061356 patent/WO2024223728A1/en active Pending
- 2024-04-25 CN CN202480028133.1A patent/CN121001738A/en active Pending
- 2024-04-25 AU AU2024260120A patent/AU2024260120A1/en active Pending
Patent Citations (129)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US4458066A (en) | 1980-02-29 | 1984-07-03 | University Patents, Inc. | Process for preparing polynucleotides |
| US4500707A (en) | 1980-02-29 | 1985-02-19 | University Patents, Inc. | Nucleosides useful in the preparation of polynucleotides |
| US5132418A (en) | 1980-02-29 | 1992-07-21 | University Patents, Inc. | Process for preparing polynucleotides |
| US4668777A (en) | 1981-03-27 | 1987-05-26 | University Patents, Inc. | Phosphoramidite nucleoside compounds |
| US4973679A (en) | 1981-03-27 | 1990-11-27 | University Patents, Inc. | Process for oligonucleo tide synthesis using phosphormidite intermediates |
| US4373071A (en) | 1981-04-30 | 1983-02-08 | City Of Hope Research Institute | Solid-phase synthesis of polynucleotides |
| US5153319A (en) | 1986-03-31 | 1992-10-06 | University Patents, Inc. | Process for preparing polynucleotides |
| US5262530A (en) | 1988-12-21 | 1993-11-16 | Applied Biosystems, Inc. | Automated system for polynucleotide synthesis and purification |
| US5047524A (en) | 1988-12-21 | 1991-09-10 | Applied Biosystems, Inc. | Automated system for polynucleotide synthesis and purification |
| US5700642A (en) | 1995-05-22 | 1997-12-23 | Sri International | Oligonucleotide sizing using immobilized cleavable primers |
| US20100036115A1 (en) | 1997-07-23 | 2010-02-11 | Sirna Therapeutics, Inc. | Novel Compositions for the Delivery of Negatively Charged Molecules |
| WO2002098443A2 (en) | 2001-06-05 | 2002-12-12 | Curevac Gmbh | Stabilised mrna with an increased g/c content and optimised codon for use in gene therapy |
| US7404969B2 (en) | 2005-02-14 | 2008-07-29 | Sirna Therapeutics, Inc. | Lipid nanoparticle based compositions and methods for the delivery of biologically active molecules |
| US7893302B2 (en) | 2005-02-14 | 2011-02-22 | Sirna Therapeutics, Inc. | Lipid nanoparticle based compositions and methods for the delivery of biologically active molecules |
| WO2008016473A2 (en) | 2006-07-28 | 2008-02-07 | Applera Corporation | Dinucleotide mrna cap analogs |
| WO2008077592A1 (en) | 2006-12-22 | 2008-07-03 | Curevac Gmbh | Method for purifying rna on a preparative scale by means of hplc |
| WO2008103276A2 (en) | 2007-02-16 | 2008-08-28 | Merck & Co., Inc. | Compositions and methods for potentiated activity of biologicaly active molecules |
| WO2008157688A2 (en) | 2007-06-19 | 2008-12-24 | Board Of Supervisors Of Louisiana State University And Agricultural And Mechanical College | Synthesis and use of anti-reverse phosphorothioate analogs of the messenger rna cap |
| WO2009086558A1 (en) | 2008-01-02 | 2009-07-09 | Tekmira Pharmaceuticals Corporation | Improved compositions and methods for the delivery of nucleic acids |
| WO2009127060A1 (en) | 2008-04-15 | 2009-10-22 | Protiva Biotherapeutics, Inc. | Novel lipid formulations for nucleic acid delivery |
| WO2009149253A2 (en) | 2008-06-06 | 2009-12-10 | Uniwersytet Warszawski | Mrna cap analogs |
| WO2010021865A1 (en) | 2008-08-18 | 2010-02-25 | Merck Sharp & Dohme Corp. | Novel lipid nanoparticles and novel components for delivery of nucleic acids |
| US20110256175A1 (en) | 2008-10-09 | 2011-10-20 | The University Of British Columbia | Amino lipids and methods for the delivery of nucleic acids |
| WO2010048536A2 (en) | 2008-10-23 | 2010-04-29 | Alnylam Pharmaceuticals, Inc. | Processes for preparing lipids |
| WO2010053572A2 (en) | 2008-11-07 | 2010-05-14 | Massachusetts Institute Of Technology | Aminoalcohol lipidoids and uses thereof |
| US8450298B2 (en) | 2008-11-07 | 2013-05-28 | Massachusetts Institute Of Technology | Aminoalcohol lipidoids and uses thereof |
| WO2010054406A1 (en) | 2008-11-10 | 2010-05-14 | Alnylam Pharmaceuticals, Inc. | Novel lipids and compositions for the delivery of therapeutics |
| WO2010080724A1 (en) | 2009-01-12 | 2010-07-15 | Merck Sharp & Dohme Corp. | Novel lipid nanoparticles and novel components for delivery of nucleic acids |
| WO2010088537A2 (en) | 2009-01-29 | 2010-08-05 | Alnylam Pharmaceuticals, Inc. | Improved lipid formulation |
| WO2010129709A1 (en) | 2009-05-05 | 2010-11-11 | Alnylam Pharmaceuticals, Inc. | Lipid compositions |
| US20120128760A1 (en) | 2009-05-05 | 2012-05-24 | Alnylam Pharmaceuticals, Inc. | Lipid compositions |
| US20100324120A1 (en) | 2009-06-10 | 2010-12-23 | Jianxin Chen | Lipid formulation |
| US8158601B2 (en) | 2009-06-10 | 2012-04-17 | Alnylam Pharmaceuticals, Inc. | Lipid formulation |
| US20120202871A1 (en) | 2009-07-01 | 2012-08-09 | Protiva Biotherapeutics, Inc. | Cationic lipids and methods for the delivery of therapeutic agents |
| US8569256B2 (en) | 2009-07-01 | 2013-10-29 | Protiva Biotherapeutics, Inc. | Cationic lipids and methods for the delivery of therapeutic agents |
| US8283333B2 (en) | 2009-07-01 | 2012-10-09 | Protiva Biotherapeutics, Inc. | Lipid formulations for nucleic acid delivery |
| WO2011015347A1 (en) | 2009-08-05 | 2011-02-10 | Biontech Ag | Vaccine composition comprising 5'-cap modified rna |
| WO2011022460A1 (en) | 2009-08-20 | 2011-02-24 | Merck Sharp & Dohme Corp. | Novel cationic lipids with various head groups for oligonucleotide delivery |
| WO2011043913A2 (en) | 2009-10-08 | 2011-04-14 | Merck Sharp & Dohme Corp. | Novel cationic lipids with short lipid chains for oligonucleotide delivery |
| WO2011069586A1 (en) | 2009-12-09 | 2011-06-16 | Curevac Gmbh | Mannose-containing solution for lyophilization, transfection and/or injection of nucleic acids |
| WO2011076807A2 (en) | 2009-12-23 | 2011-06-30 | Novartis Ag | Lipids, lipid compositions, and methods of using them |
| WO2011090965A1 (en) | 2010-01-22 | 2011-07-28 | Merck Sharp & Dohme Corp. | Novel cationic lipids for oligonucleotide delivery |
| US20130129785A1 (en) | 2010-05-10 | 2013-05-23 | Alnylam Pharmaceuticals, Inc | Methods and compositions for delivery of active agents |
| WO2011149733A2 (en) | 2010-05-24 | 2011-12-01 | Merck Sharp & Dohme Corp. | Novel amino alcohol cationic lipids for oligonucleotide delivery |
| US20130150625A1 (en) | 2010-05-24 | 2013-06-13 | Brian W. Budzik | Novel Amino Alcohol Cationic Lipids for Oligonucleotide Delivery |
| WO2011153493A2 (en) | 2010-06-03 | 2011-12-08 | Alnylam Pharmaceuticals, Inc. | Biodegradable lipids for the delivery of active agents |
| US20120027803A1 (en) | 2010-06-03 | 2012-02-02 | Alnylam Pharmaceuticals, Inc. | Biodegradable lipids for the delivery of active agents |
| WO2011153120A1 (en) | 2010-06-04 | 2011-12-08 | Merck Sharp & Dohme Corp. | Novel low molecular weight cationic lipids for oligonucleotide delivery |
| WO2012006378A1 (en) | 2010-07-06 | 2012-01-12 | Novartis Ag | Liposomes with lipids having an advantageous pka- value for rna delivery |
| WO2012006376A2 (en) | 2010-07-06 | 2012-01-12 | Novartis Ag | Virion-like delivery particles for self-replicating rna molecules |
| WO2012019780A1 (en) | 2010-08-13 | 2012-02-16 | Curevac Gmbh | Nucleic acid comprising or coding for a histone stem-loop and a poly(a) sequence or a polyadenylation signal for increasing the expression of an encoded protein |
| WO2012031043A1 (en) | 2010-08-31 | 2012-03-08 | Novartis Ag | Pegylated liposomes for delivery of immunogen-encoding rna |
| WO2012030901A1 (en) | 2010-08-31 | 2012-03-08 | Novartis Ag | Small liposomes for delivery of immunogen-encoding rna |
| WO2012031046A2 (en) | 2010-08-31 | 2012-03-08 | Novartis Ag | Lipids suitable for liposomal delivery of protein-coding rna |
| US8466122B2 (en) | 2010-09-17 | 2013-06-18 | Protiva Biotherapeutics, Inc. | Trialkyl cationic lipids and methods of use thereof |
| WO2012040184A2 (en) | 2010-09-20 | 2012-03-29 | Merck Sharp & Dohme Corp. | Novel low molecular weight cationic lipids for oligonucleotide delivery |
| US20130178541A1 (en) | 2010-09-20 | 2013-07-11 | Matthew G. Stanton | Novel low molecular weight cationic lipids for oligonucleotide delivery |
| WO2012044638A1 (en) | 2010-09-30 | 2012-04-05 | Merck Sharp & Dohme Corp. | Low molecular weight cationic lipids for oligonucleotide delivery |
| WO2012054365A2 (en) | 2010-10-21 | 2012-04-26 | Merck Sharp & Dohme Corp. | Novel low molecular weight cationic lipids for oligonucleotide delivery |
| WO2012061259A2 (en) | 2010-11-05 | 2012-05-10 | Merck Sharp & Dohme Corp. | Novel low molecular weight cyclic amine containing cationic lipids for oligonucleotide delivery |
| US20130225836A1 (en) | 2010-11-05 | 2013-08-29 | Merck Sharp & Dohme Corp. | Novel low molecular weight cyclic amine containing cationic lipids for oligonucleotide delivery |
| WO2012170889A1 (en) | 2011-06-08 | 2012-12-13 | Shire Human Genetic Therapies, Inc. | Cleavable lipids |
| WO2012170930A1 (en) | 2011-06-08 | 2012-12-13 | Shire Human Genetic Therapies, Inc | Lipid nanoparticle compositions and methods for mrna delivery |
| WO2013006825A1 (en) | 2011-07-06 | 2013-01-10 | Novartis Ag | Liposomes having useful n:p ratio for delivery of rna molecules |
| US20130064894A1 (en) | 2011-08-31 | 2013-03-14 | Protiva Biotherapeutics, Inc. | Novel cationic lipids and methods of use thereof |
| WO2013033563A1 (en) | 2011-08-31 | 2013-03-07 | Novartis Ag | Pegylated liposomes for delivery of immunogen-encoding rna |
| WO2013059475A1 (en) | 2011-10-18 | 2013-04-25 | Life Technologies Corporation | Alkynyl-derivatized cap analogs, preparation and uses thereof |
| WO2013063468A1 (en) | 2011-10-27 | 2013-05-02 | Massachusetts Institute Of Technology | Amino acid derivates functionalized on the n- terminal capable of forming drug incapsulating microspheres |
| WO2013086354A1 (en) | 2011-12-07 | 2013-06-13 | Alnylam Pharmaceuticals, Inc. | Biodegradable lipids for the delivery of active agents |
| WO2013086373A1 (en) | 2011-12-07 | 2013-06-13 | Alnylam Pharmaceuticals, Inc. | Lipids for the delivery of active agents |
| US20140039032A1 (en) | 2011-12-12 | 2014-02-06 | Kyowa Hakko Kirin Co., Ltd. | Lipid nano particles comprising cationic lipid for drug delivery system |
| WO2013143700A2 (en) | 2012-03-27 | 2013-10-03 | Curevac Gmbh | Artificial nucleic acid molecules comprising a 5'top utr |
| WO2014136086A1 (en) | 2013-03-08 | 2014-09-12 | Novartis Ag | Lipids and lipid compositions for the delivery of active agents |
| WO2014152673A1 (en) | 2013-03-14 | 2014-09-25 | Shire Human Genetic Therapies, Inc. | Quantitative assessment for cap efficiency of messenger rna |
| WO2014152659A1 (en) | 2013-03-14 | 2014-09-25 | Shire Human Genetic Therapies, Inc. | Quantitative assessment for cap efficiency of messenger rna |
| US20150140070A1 (en) | 2013-10-22 | 2015-05-21 | Shire Human Genetic Therapies, Inc. | Lipid formulations for delivery of messenger rna |
| WO2015062738A1 (en) | 2013-11-01 | 2015-05-07 | Curevac Gmbh | Modified rna with decreased immunostimulatory properties |
| WO2015074085A1 (en) | 2013-11-18 | 2015-05-21 | Arcturus Therapeutics, Inc. | Ionizable cationic lipid for rna delivery |
| US9593077B2 (en) | 2013-11-18 | 2017-03-14 | Arcturus Therapeutics, Inc. | Ionizable cationic lipid for RNA delivery |
| US9567296B2 (en) | 2013-11-18 | 2017-02-14 | Arcturus Therapeutics, Inc. | Ionizable cationic lipid for RNA delivery |
| WO2015095346A1 (en) | 2013-12-19 | 2015-06-25 | Novartis Ag | Lipids and lipid compositions for the delivery of active agents |
| WO2015095340A1 (en) | 2013-12-19 | 2015-06-25 | Novartis Ag | Lipids and lipid compositions for the delivery of active agents |
| WO2015101416A1 (en) | 2013-12-30 | 2015-07-09 | Curevac Gmbh | Methods for rna analysis |
| WO2015188933A1 (en) | 2014-06-10 | 2015-12-17 | Curevac Ag | Methods and means for enhancing rna production |
| US20150376115A1 (en) | 2014-06-25 | 2015-12-31 | Acuitas Therapeutics Inc. | Novel lipids and lipid nanoparticle formulations for delivery of nucleic acids |
| WO2015199952A1 (en) | 2014-06-25 | 2015-12-30 | Acuitas Therapeutics Inc. | Novel lipids and lipid nanoparticle formulations for delivery of nucleic acids |
| WO2016022914A1 (en) | 2014-08-08 | 2016-02-11 | Moderna Therapeutics, Inc. | Compositions and methods for the treatment of ophthalmic diseases and conditions |
| WO2016037053A1 (en) | 2014-09-05 | 2016-03-10 | Novartis Ag | Lipids and lipid compositions for the delivery of active agents |
| WO2016107877A1 (en) | 2014-12-30 | 2016-07-07 | Curevac Ag | Artificial nucleic acid molecules |
| WO2016118724A1 (en) | 2015-01-21 | 2016-07-28 | Moderna Therapeutics, Inc. | Lipid nanoparticle compositions |
| WO2016118725A1 (en) | 2015-01-23 | 2016-07-28 | Moderna Therapeutics, Inc. | Lipid nanoparticle compositions |
| WO2016165831A1 (en) | 2015-04-17 | 2016-10-20 | Curevac Ag | Lyophilization of rna |
| WO2016174271A1 (en) | 2015-04-30 | 2016-11-03 | Curevac Ag | Immobilized poly(n)polymerase |
| WO2016180430A1 (en) | 2015-05-08 | 2016-11-17 | Curevac Ag | Method for producing rna |
| WO2016184576A2 (en) | 2015-05-20 | 2016-11-24 | Curevac Ag | Dry powder composition comprising long-chain rna |
| WO2016184575A1 (en) | 2015-05-20 | 2016-11-24 | Curevac Ag | Dry powder composition comprising long-chain rna |
| WO2016193226A1 (en) | 2015-05-29 | 2016-12-08 | Curevac Ag | Method for adding cap structures to rna using immobilized enzymes |
| WO2016193206A1 (en) | 2015-05-29 | 2016-12-08 | Curevac Ag | A method for producing and purifying rna, comprising at least one step of tangential flow filtration |
| WO2017036580A1 (en) | 2015-08-28 | 2017-03-09 | Curevac Ag | Artificial nucleic acid molecules |
| WO2017053297A1 (en) | 2015-09-21 | 2017-03-30 | Trilink Biotechnologies, Inc. | Compositions and methods for synthesizing 5'-capped rnas |
| WO2017066797A1 (en) | 2015-10-16 | 2017-04-20 | Modernatx, Inc. | Trinucleotide mrna cap analogs |
| WO2017066782A1 (en) | 2015-10-16 | 2017-04-20 | Modernatx, Inc. | Hydrophobic mrna cap analogs |
| WO2017066793A1 (en) | 2015-10-16 | 2017-04-20 | Modernatx, Inc. | Mrna cap analogs and methods of mrna capping |
| WO2017066781A1 (en) | 2015-10-16 | 2017-04-20 | Modernatx, Inc. | Mrna cap analogs with modified phosphate linkage |
| WO2017066791A1 (en) | 2015-10-16 | 2017-04-20 | Modernatx, Inc. | Sugar substituted mrna cap analogs |
| WO2017066789A1 (en) | 2015-10-16 | 2017-04-20 | Modernatx, Inc. | Mrna cap analogs with modified sugar |
| WO2017070613A1 (en) | 2015-10-22 | 2017-04-27 | Modernatx, Inc. | Human cytomegalovirus vaccine |
| WO2017070620A2 (en) | 2015-10-22 | 2017-04-27 | Modernatx, Inc. | Broad spectrum influenza virus vaccine |
| US20180311336A1 (en) * | 2015-10-22 | 2018-11-01 | Moderna TX, Inc. | Broad spectrum influenza virus vaccine |
| WO2017075531A1 (en) | 2015-10-28 | 2017-05-04 | Acuitas Therapeutics, Inc. | Novel lipids and lipid nanoparticle formulations for delivery of nucleic acids |
| WO2017099823A1 (en) | 2015-12-10 | 2017-06-15 | Modernatx, Inc. | Compositions and methods for delivery of therapeutic agents |
| WO2017112865A1 (en) | 2015-12-22 | 2017-06-29 | Modernatx, Inc. | Compounds and compositions for intracellular delivery of agents |
| WO2017109161A1 (en) | 2015-12-23 | 2017-06-29 | Curevac Ag | Method of rna in vitro transcription using a buffer containing a dicarboxylic acid or tricarboxylic acid or a salt thereof |
| WO2017117530A1 (en) | 2015-12-30 | 2017-07-06 | Arcturus Therapeutics, Inc. | Ionizable cationic lipid |
| WO2018075827A1 (en) | 2016-10-19 | 2018-04-26 | Arcturus Therapeutics, Inc. | Trinucleotide mrna cap analogs |
| WO2018078053A1 (en) | 2016-10-26 | 2018-05-03 | Curevac Ag | Lipid nanoparticle mrna vaccines |
| JP2019008858A (en) | 2017-06-27 | 2019-01-17 | 花王株式会社 | Polishing liquid composition |
| WO2019077001A1 (en) | 2017-10-19 | 2019-04-25 | Curevac Ag | Novel artificial nucleic acid molecules |
| WO2019092153A1 (en) | 2017-11-08 | 2019-05-16 | Curevac Ag | Rna sequence adaptation |
| WO2020002525A1 (en) | 2018-06-27 | 2020-01-02 | Curevac Ag | Novel lassa virus rna molecules and compositions for vaccination |
| WO2021156267A1 (en) | 2020-02-04 | 2021-08-12 | Curevac Ag | Coronavirus vaccine |
| WO2021216541A1 (en) | 2020-04-20 | 2021-10-28 | Board Of Regents, The University Of Texas System | Biologically active dry powder compositions and method of their manufacture and use |
| WO2022036170A1 (en) | 2020-08-14 | 2022-02-17 | Arcturus Therapeutics, Inc. | Method of lyophilizing lipid nanoparticles |
| WO2022076562A1 (en) | 2020-10-06 | 2022-04-14 | Translate Bio, Inc. | Improved process and formulation of lipid nanoparticles |
| US20220142923A1 (en) * | 2020-11-06 | 2022-05-12 | Sanofi | LIPID NANOPARTICLES FOR DELIVERING mRNA VACCINES |
| WO2022101461A1 (en) | 2020-11-16 | 2022-05-19 | BioNTech SE | Enhanced formulation stabilization and improved lyophilization processes |
| WO2022137133A1 (en) | 2020-12-22 | 2022-06-30 | Curevac Ag | Rna vaccine against sars-cov-2 variants |
| WO2022232585A1 (en) | 2021-04-29 | 2022-11-03 | Modernatx, Inc. | Lyophilization methods for preparing lipid formulated therapeutics |
| WO2022264109A1 (en) * | 2021-06-18 | 2022-12-22 | Sanofi | Multivalent influenza vaccines |
Non-Patent Citations (7)
Also Published As
| Publication number | Publication date |
|---|---|
| CN121001738A (en) | 2025-11-21 |
| AU2024260120A1 (en) | 2025-11-06 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| US12390523B2 (en) | Coronavirus vaccine | |
| US20220211841A1 (en) | Nucleic acid based combination vaccines | |
| US11241493B2 (en) | Coronavirus vaccine | |
| EP4147717A1 (en) | Coronavirus vaccine | |
| US20250064916A1 (en) | Immunogenic compositions and uses thereof | |
| WO2024068545A1 (en) | Influenza virus vaccines | |
| US20240181037A1 (en) | Immunogenic compositions | |
| WO2024223728A1 (en) | Influenza virus vaccines | |
| WO2025132839A1 (en) | Influenza virus vaccines | |
| WO2024223724A1 (en) | Influenza virus vaccines | |
| WO2025011529A2 (en) | Circular rna vaccines for seasonal flu and methods of uses | |
| HK40090415A (en) | Coronavirus vaccine |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| 121 | Ep: the epo has been informed by wipo that ep was designated in this application |
Ref document number: 24722518 Country of ref document: EP Kind code of ref document: A1 |
|
| WWE | Wipo information: entry into national phase |
Ref document number: AU2024260120 Country of ref document: AU |
|
| ENP | Entry into the national phase |
Ref document number: 2024260120 Country of ref document: AU Date of ref document: 20240425 Kind code of ref document: A |
|
| WWE | Wipo information: entry into national phase |
Ref document number: 2024722518 Country of ref document: EP |
|
| NENP | Non-entry into the national phase |
Ref country code: DE |
|
| ENP | Entry into the national phase |
Ref document number: 2024722518 Country of ref document: EP Effective date: 20251127 |
|
| ENP | Entry into the national phase |
Ref document number: 2024722518 Country of ref document: EP Effective date: 20251127 |
|
| ENP | Entry into the national phase |
Ref document number: 2024722518 Country of ref document: EP Effective date: 20251127 |