WO2024163678A2 - Fusion proteins and systems for targeted activation of frataxin (fxn) and related methods - Google Patents
Fusion proteins and systems for targeted activation of frataxin (fxn) and related methods Download PDFInfo
- Publication number
- WO2024163678A2 WO2024163678A2 PCT/US2024/013874 US2024013874W WO2024163678A2 WO 2024163678 A2 WO2024163678 A2 WO 2024163678A2 US 2024013874 W US2024013874 W US 2024013874W WO 2024163678 A2 WO2024163678 A2 WO 2024163678A2
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- seq
- ezfp
- set forth
- sequence
- sequence set
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Ceased
Links
Classifications
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K14/00—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof
- C07K14/435—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans
- C07K14/46—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans from vertebrates
- C07K14/47—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans from vertebrates from mammals
- C07K14/4701—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans from vertebrates from mammals not used
- C07K14/4702—Regulators; Modulating activity
- C07K14/4705—Regulators; Modulating activity stimulating, promoting or activating activity
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P43/00—Drugs for specific purposes, not provided for in groups A61P1/00-A61P41/00
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/11—DNA or RNA fragments; Modified forms thereof; Non-coding nucleic acids having a biological activity
- C12N15/113—Non-coding nucleic acids modulating the expression of genes, e.g. antisense oligonucleotides; Antisense DNA or RNA; Triplex- forming oligonucleotides; Catalytic nucleic acids, e.g. ribozymes; Nucleic acids used in co-suppression or gene silencing
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/11—DNA or RNA fragments; Modified forms thereof; Non-coding nucleic acids having a biological activity
- C12N15/62—DNA sequences coding for fusion proteins
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N9/00—Enzymes; Proenzymes; Compositions thereof; Processes for preparing, activating, inhibiting, separating or purifying enzymes
- C12N9/14—Hydrolases (3)
- C12N9/16—Hydrolases (3) acting on ester bonds (3.1)
- C12N9/22—Ribonucleases [RNase]; Deoxyribonucleases [DNase]
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2319/00—Fusion polypeptide
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2319/00—Fusion polypeptide
- C07K2319/80—Fusion polypeptide containing a DNA binding domain, e.g. Lacl or Tet-repressor
- C07K2319/81—Fusion polypeptide containing a DNA binding domain, e.g. Lacl or Tet-repressor containing a Zn-finger domain for DNA binding
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N2310/00—Structure or type of the nucleic acid
- C12N2310/10—Type of nucleic acid
- C12N2310/20—Type of nucleic acid involving clustered regularly interspaced short palindromic repeats [CRISPR]
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N2320/00—Applications; Uses
- C12N2320/10—Applications; Uses in screening processes
- C12N2320/11—Applications; Uses in screening processes for the determination of target sites, i.e. of active nucleic acids
Definitions
- the present disclosure relates in some aspects to compositions, such as DNA-targeting systems, fusion proteins, guide RNAs (gRNAs), engineered zinc finger proteins (eZFPs) and fusion proteins comprising eZFPs, and pluralities and combinations thereof, that bind to or target a frataxin (FXN) locus.
- compositions such as DNA-targeting systems, fusion proteins, guide RNAs (gRNAs), engineered zinc finger proteins (eZFPs) and fusion proteins comprising eZFPs, and pluralities and combinations thereof, that bind to or target a frataxin (FXN) locus.
- FXN frataxin
- the present disclosure also relates to polynucleotides, vectors, cells and pluralities and combinations thereof, that encode or comprise the DNA-targeting systems, fusion proteins, gRNAs, engineered zinc finger proteins(eZFPs) and fusion proteins comprising eZFPs, or pluralities or combinations thereof, and methods and uses related to the provided compositions, for example, in modulating the expression of FXN, and/or in the treatment or therapy of diseases or disorders that involve the activity, function or expression of FXN, such as Friedreich’s Ataxia (FA).
- FXN Friedreich’s Ataxia
- FA frataxin
- FXN frataxin
- FA is an autosomal recessive neurodegenerative and cardiac disease, and is caused by a trinucleotide repeat expansion mutation in the FXN gene.
- FA can result in ataxia, areflexia, loss of vibratory sense and proprioception, dysarthria, cardiomyopathy, and/or associated arrhythmias, among other symptoms.
- Existing treatment of such genetic disorders are directed towards symptoms and providing support. Treatments that address the fundamental etiology and disease mechanism are needed. Provided are embodiments that meet such needs.
- an engineered zinc finger protein that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the target site is within a target region spanning the genomic coordinates chr9:69, 034, 900-69, 035, 900 from human genome assembly GRCh38 (hg38) (SEQ ID NO:430), or within a target region spanning the genomic coordinates chr9:69, 027, 282-69, 028, 497 from hg38 (SEQ ID NO:431).
- eZFP engineered zinc finger protein
- the target site is within a target region spanning the genomic coordinates chr9:69, 034, 900-69, 035, 900 from hg38 (SEQ ID NO:430). In some of any of the provided embodiments, the target site is within a target region spanning the genomic coordinates chr9:69, 035, 300-69-035, 800 from hg38. In some of any of the provided embodiments, the target site is within a target region spanning the genomic coordinates chr9:69, 035, 350-69, 035, 450 from hg38.
- the target site is within a target region spanning the genomic coordinates chr9:69, 035, 400-69, 035, 450 from hg38. In some of any of the provided embodiments, the target site is within a target region spanning the genomic coordinates chr9:69, 035, 530-69, 035, 580 from hg38. In some of any of the provided embodiments, the target site is within a target region spanning the genomic coordinates chr9:69, 035, 675-69, 035, 725 from hg38.
- the target site is within a target region spanning the genomic coordinates chr9:69, 027, 282-69, 028, 497 from hg38 (SEQ ID NO:431). In some of any of the provided embodiments, the target site is within a target region spanning the genomic coordinates chr9:69, 027, 615-69, 028, 101 from hg38. In some of any of the provided embodiments, the target site is within a target region spanning the genomic coordinates chr9:69, 027, 775-69, 027, 875 from hg38.
- the target site is within a target region spanning the genomic coordinates chr9:69, 027, 795-69, 027, 845 from hg38.
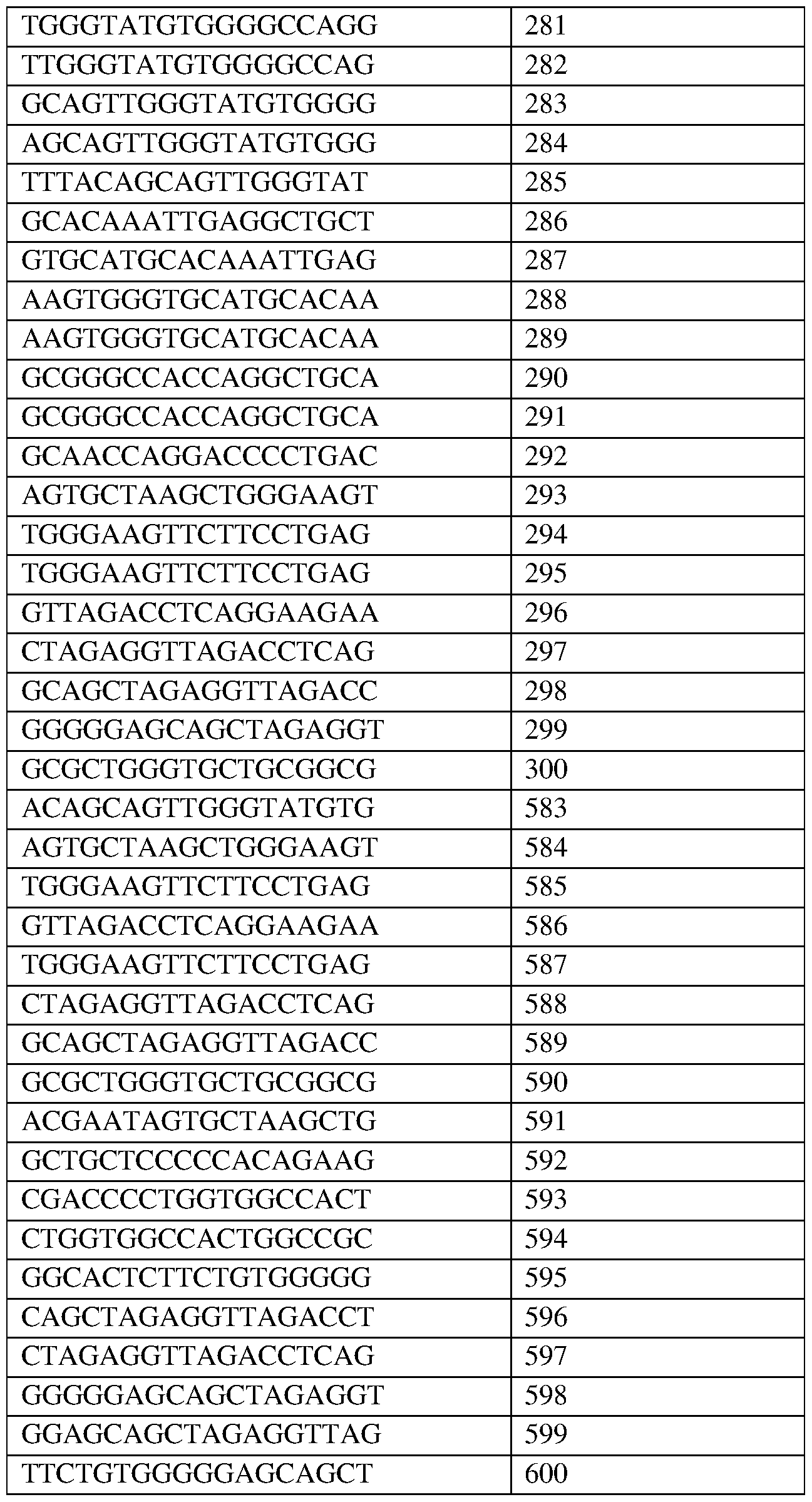
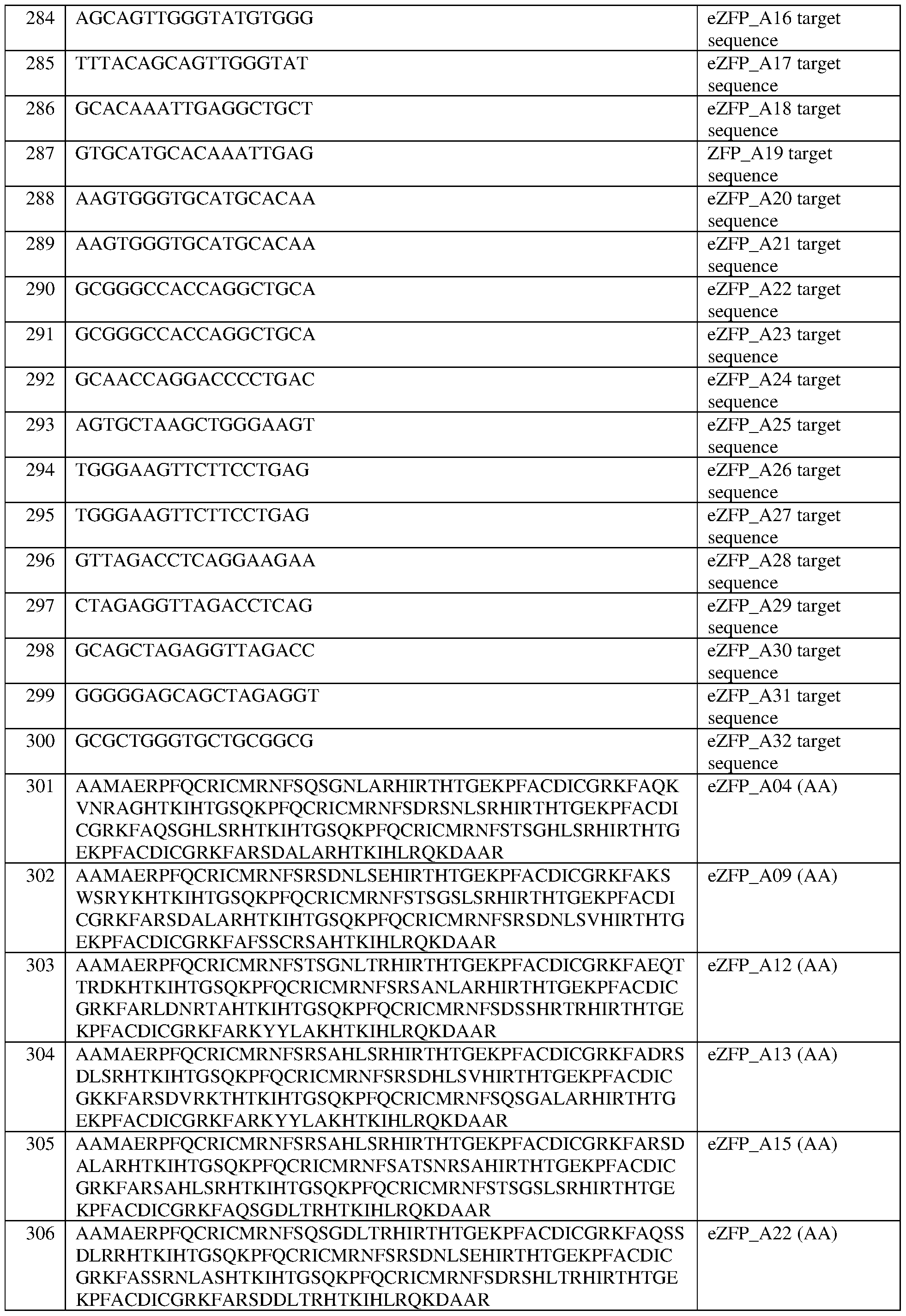
- the target site comprises the nucleotide sequence set forth in any one of SEQ ID NOS:269-300 and 583-600, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
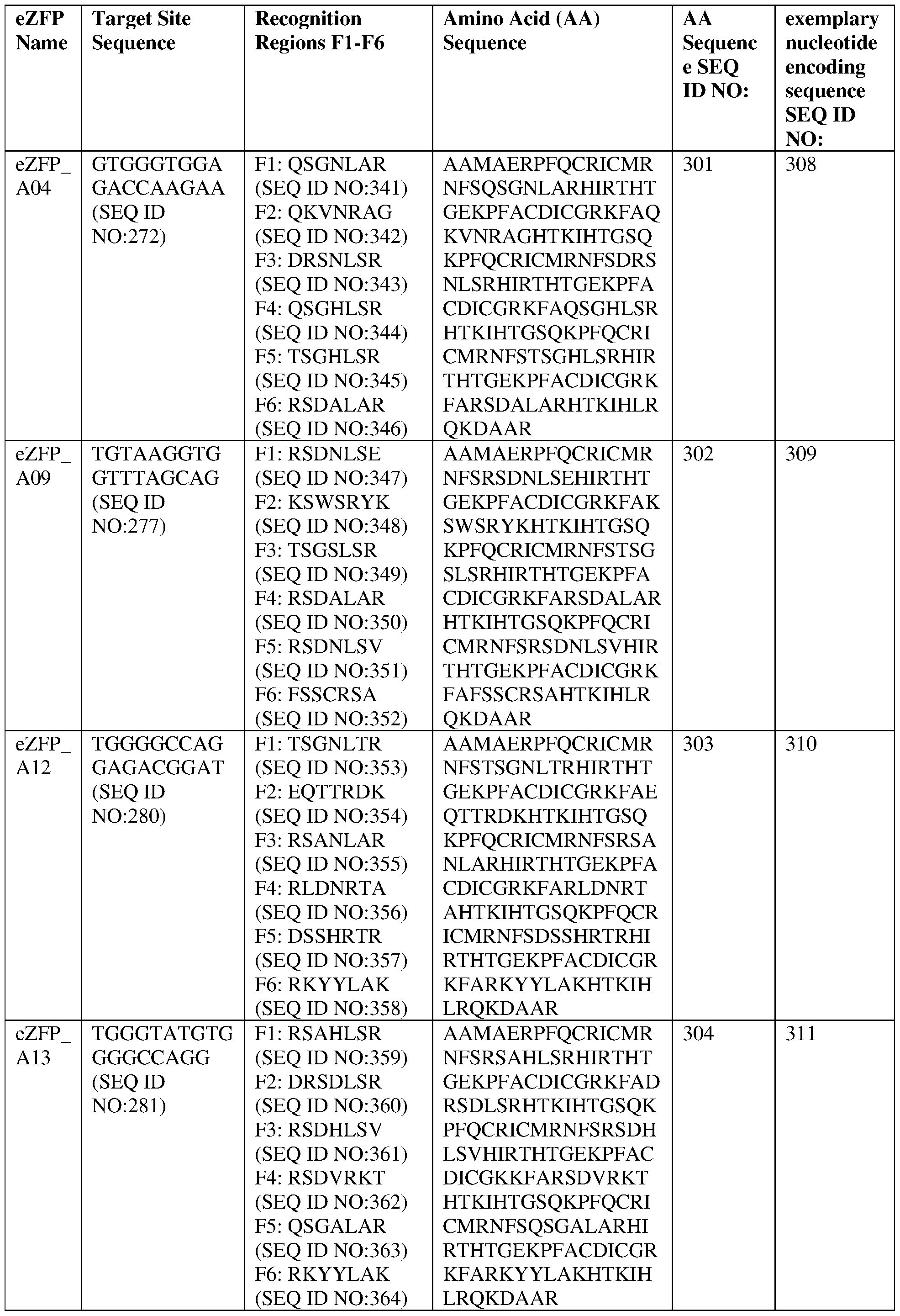
- the target site comprises the nucleotide sequence set forth in SEQ ID NO:272, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N- terminus to C-terminus, selected from F1-F6 as follows: Fl: QSGNLAR (SEQ ID NO:341); F2: QKVNRAG (SEQ ID NO:342); F3: DRSNLSR (SEQ ID NO:343); F4: QSGHLSR (SEQ ID NO:344); F5: TSGHLSR (SEQ ID NO:345); and F6: RSDALAR (SEQ ID NO:346).
- an engineered zinc finger protein that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: QSGNLAR (SEQ ID NO:341); F2: QKVNRAG (SEQ ID NO:342); F3: DRSNLSR (SEQ ID NO:343); F4: QSGHLSR (SEQ ID NO:344); F5: TSGHLSR (SEQ ID NO:345); and F6: RSDALAR (SEQ ID NO:346).
- the eZFP comprises the sequence set forth in SEQ ID NO:301, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:301.
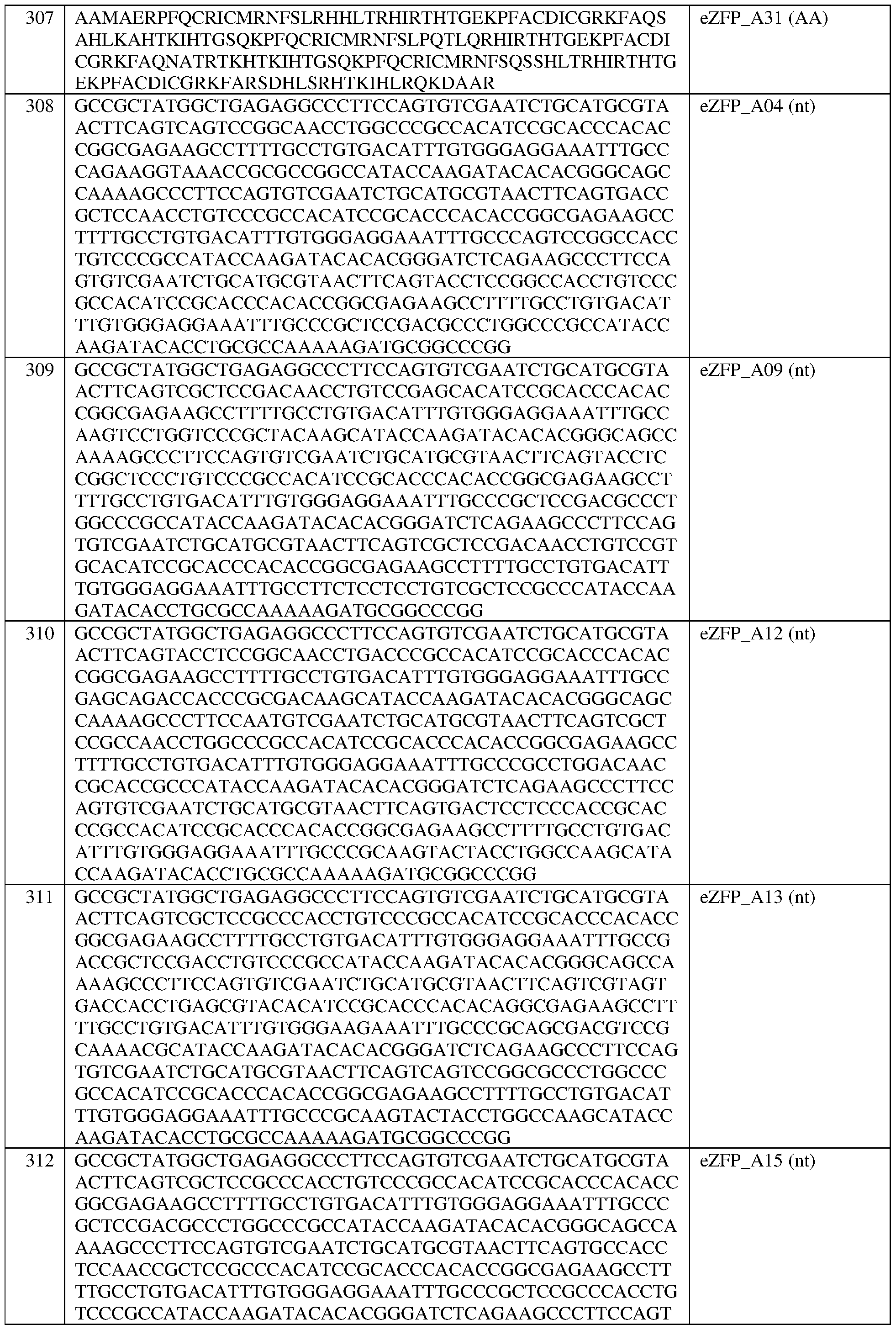
- the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:308 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:308.
- the target site comprises the nucleotide sequence set forth in SEQ ID NO:277, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N- terminus to C-terminus, selected from F1-F6 as follows: Fl: RSDNLSE (SEQ ID NO:347); F2: KSWSRYK (SEQ ID NO:348); F3: TSGSLSR (SEQ ID NO:349); F4: RSDALAR (SEQ ID NO:350); F5: RSDNLSV (SEQ ID NO:351); and F6: FSSCRSA (SEQ ID NO:352).
- an engineered zinc finger protein that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RSDNLSE (SEQ ID NO:347); F2: KSWSRYK (SEQ ID NO:348); F3: TSGSLSR (SEQ ID NO:349); F4: RSDALAR (SEQ ID NO:350); F5: RSDNLSV (SEQ ID NO:351); and F6: FSSCRSA (SEQ ID NO:352).
- the eZFP comprises the sequence set forth in SEQ ID NO:302, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:302.
- the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:309 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:309.
- the target site comprises the nucleotide sequence set forth in SEQ ID NO:280, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N- terminus to C-terminus, selected from F1-F6 as follows: Fl: TSGNLTR (SEQ ID NO:353); F2: EQTTRDK (SEQ ID NO:354); F3: RSANLAR (SEQ ID NO:355); F4: RLDNRTA (SEQ ID NO:356); F5: DSSHRTR (SEQ ID NO:357); and F6: RKYYLAK (SEQ ID NO:358).
- an engineered zinc finger protein that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: TSGNLTR (SEQ ID NO:353); F2: EQTTRDK (SEQ ID NO:354); F3: RSANLAR (SEQ ID NO:355); F4: RLDNRTA (SEQ ID NO:356); F5: DSSHRTR (SEQ ID NO:357); and F6: RKYYLAK (SEQ ID NO:358).
- the eZFP comprises the sequence set forth in SEQ ID NO:303, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:303.
- the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:310 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:310.
- the target site comprises the nucleotide sequence set forth in SEQ ID NO:281, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
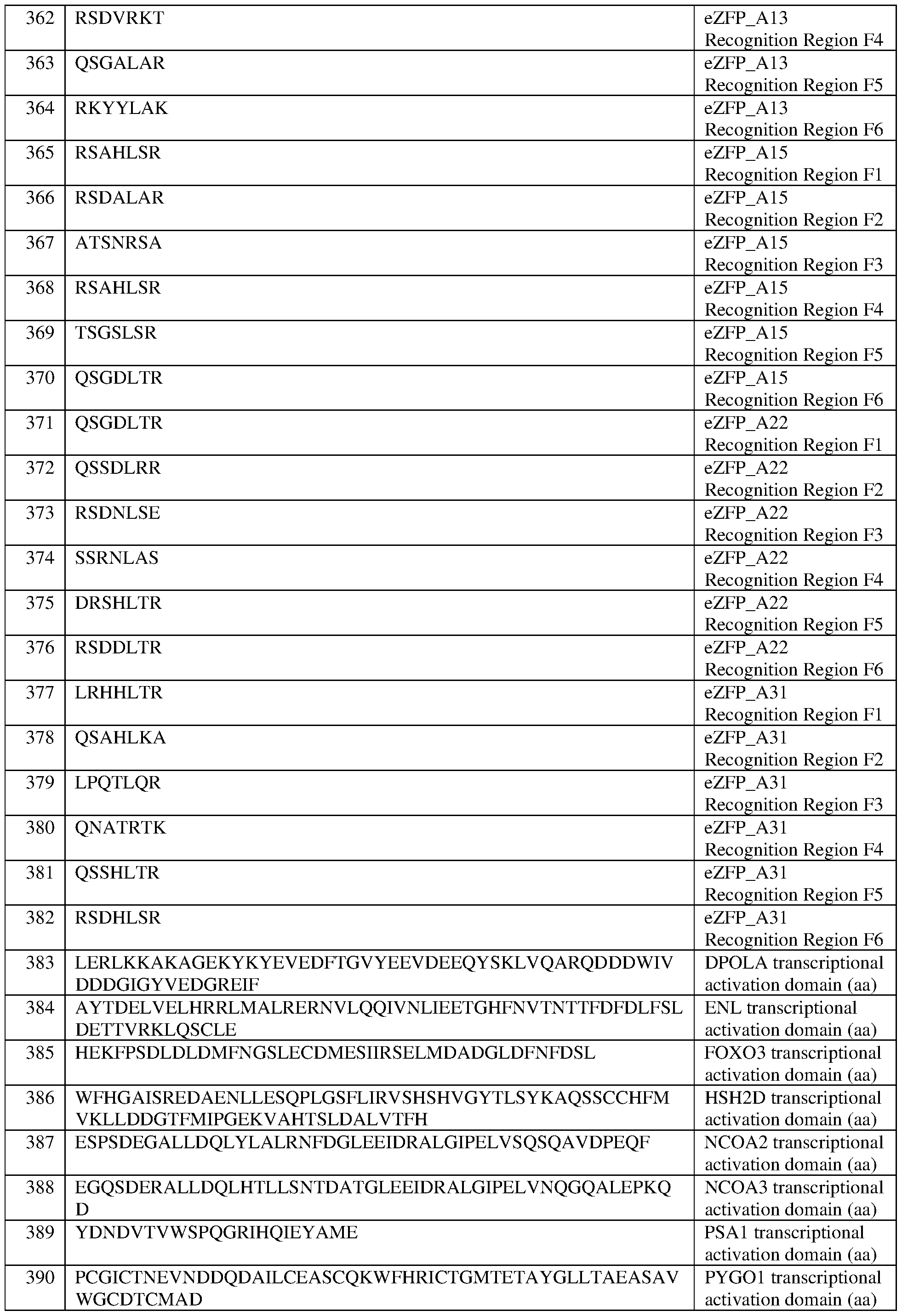
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N- terminus to C-terminus, selected from F1-F6 as follows: Fl: RSAHLSR (SEQ ID NO:359); F2: DRSDLSR (SEQ ID NO:360); F3: RSDHLSV (SEQ ID NO:361); F4: RSDVRKT (SEQ ID NO:362); F5: QSGALAR (SEQ ID NO:363); and F6: RKYYLAK (SEQ ID NO:364).
- an engineered zinc finger protein that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RSAHLSR (SEQ ID NO:359); F2: DRSDLSR (SEQ ID NO:360); F3: RSDHLSV (SEQ ID NO:361); F4: RSDVRKT (SEQ ID NO:362); F5: QSGALAR (SEQ ID NO:363); and F6: RKYYLAK (SEQ ID NO:364).
- the eZFP comprises the sequence set forth in SEQ ID NO:304, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:304.
- the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:311 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:311.
- the target site comprises the nucleotide sequence set forth in SEQ ID NO:283, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N- terminus to C-terminus, selected from F1-F6 as follows: Fl: RSAHLSR (SEQ ID NO:365); F2: RSDALAR (SEQ ID NO:366); F3: ATSNRSA (SEQ ID NO:367); F4: RSAHLSR (SEQ ID NO:368); F5: TSGSLSR (SEQ ID NO:369); and F6: QSGDLTR (SEQ ID NO:370).
- an engineered zinc finger protein that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RSAHLSR (SEQ ID NO:365); F2: RSDALAR (SEQ ID NO:366); F3: ATSNRSA (SEQ ID NO:367); F4: RSAHLSR (SEQ ID NO:368); F5: TSGSLSR (SEQ ID NO:369); and F6: QSGDLTR (SEQ ID NO:370).
- the eZFP comprises the sequence set forth in SEQ ID NO:305, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:305.
- the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:312 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NOG 12.
- the target site comprises the nucleotide sequence set forth in SEQ ID NO:290, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N- terminus to C-terminus, selected from F1-F6 as follows: Fl: QSGDLTR (SEQ ID NO:371); F2: QSSDLRR (SEQ ID NO:372); F3: RSDNLSE (SEQ ID NO:373); F4: SSRNLAS (SEQ ID NO:374); F5: DRSHLTR (SEQ ID NO:375); and F6: RSDDLTR (SEQ ID NO:376).
- an engineered zinc finger protein that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: QSGDLTR (SEQ ID NO:371); F2: QSSDLRR (SEQ ID NO:372); F3: RSDNLSE (SEQ ID NO:373); F4: SSRNLAS (SEQ ID NO:374); F5: DRSHLTR (SEQ ID NO:375); and F6: RSDDLTR (SEQ ID NO:376).
- the eZFP comprises the sequence set forth in SEQ ID NO:306, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:306.
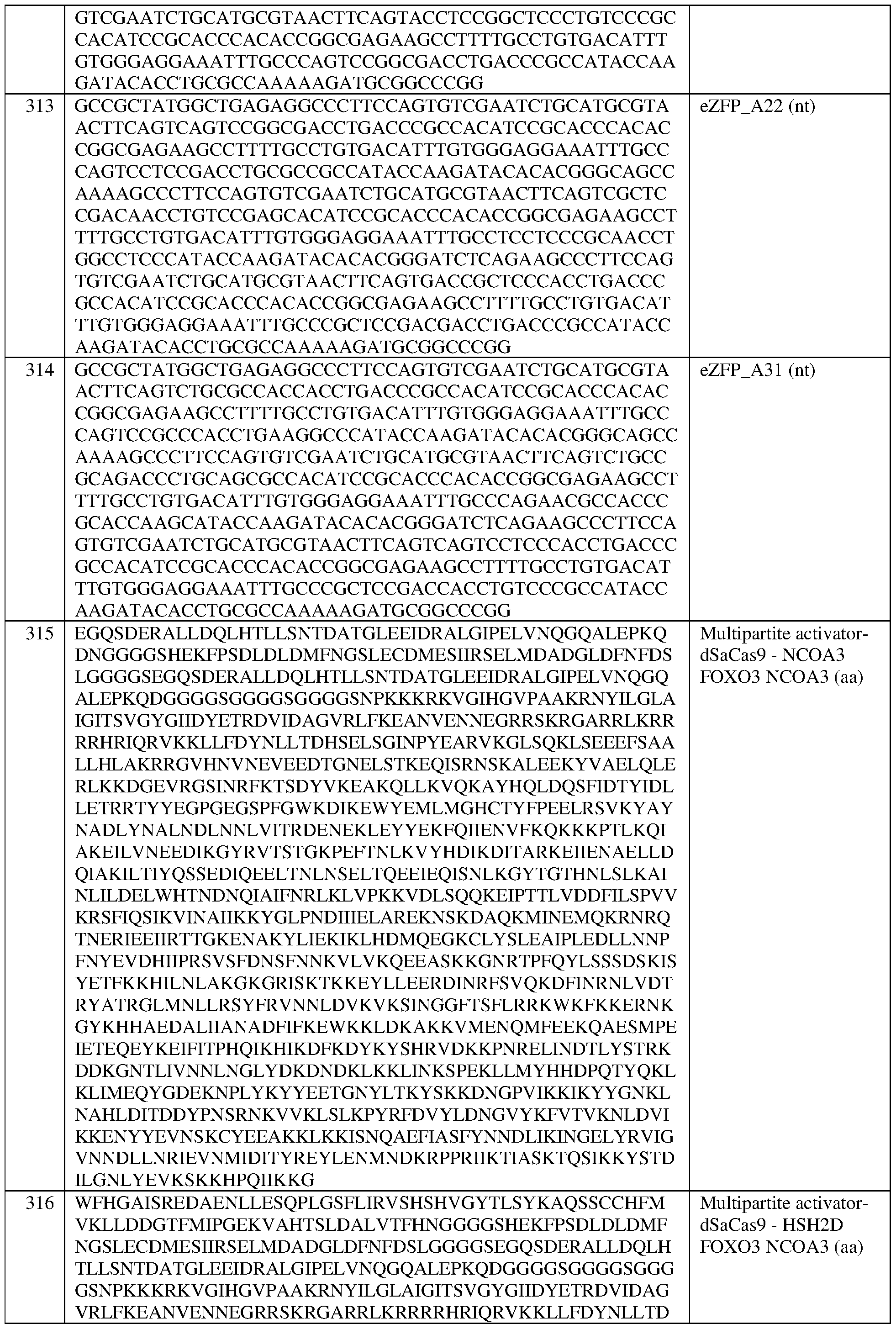
- the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:313 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NOG 13.
- the target site comprises the nucleotide sequence set forth in SEQ ID NO:299, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N- terminus to C-terminus, selected from F1-F6 as follows: Fl: LRHHLTR (SEQ ID NO:377); F2: QSAHLKA (SEQ ID NO:378); F3: LPQTLQR (SEQ ID NO:379); F4: QNATRTK (SEQ ID NO:380); F5: QSSHLTR (SEQ ID NO:381); and F6: RSDHLSR (SEQ ID NO:382).
- an engineered zinc finger protein that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: LRHHLTR (SEQ ID NO:377); F2: QSAHLKA (SEQ ID NO:378); F3: LPQTLQR (SEQ ID NO:379); F4: QNATRTK (SEQ ID NO:380); F5: QSSHLTR (SEQ ID NO:381); and F6: RSDHLSR (SEQ ID NO:382).
- the eZFP comprises the sequence set forth in SEQ ID NO:307, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:307.
- the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:314 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:314.
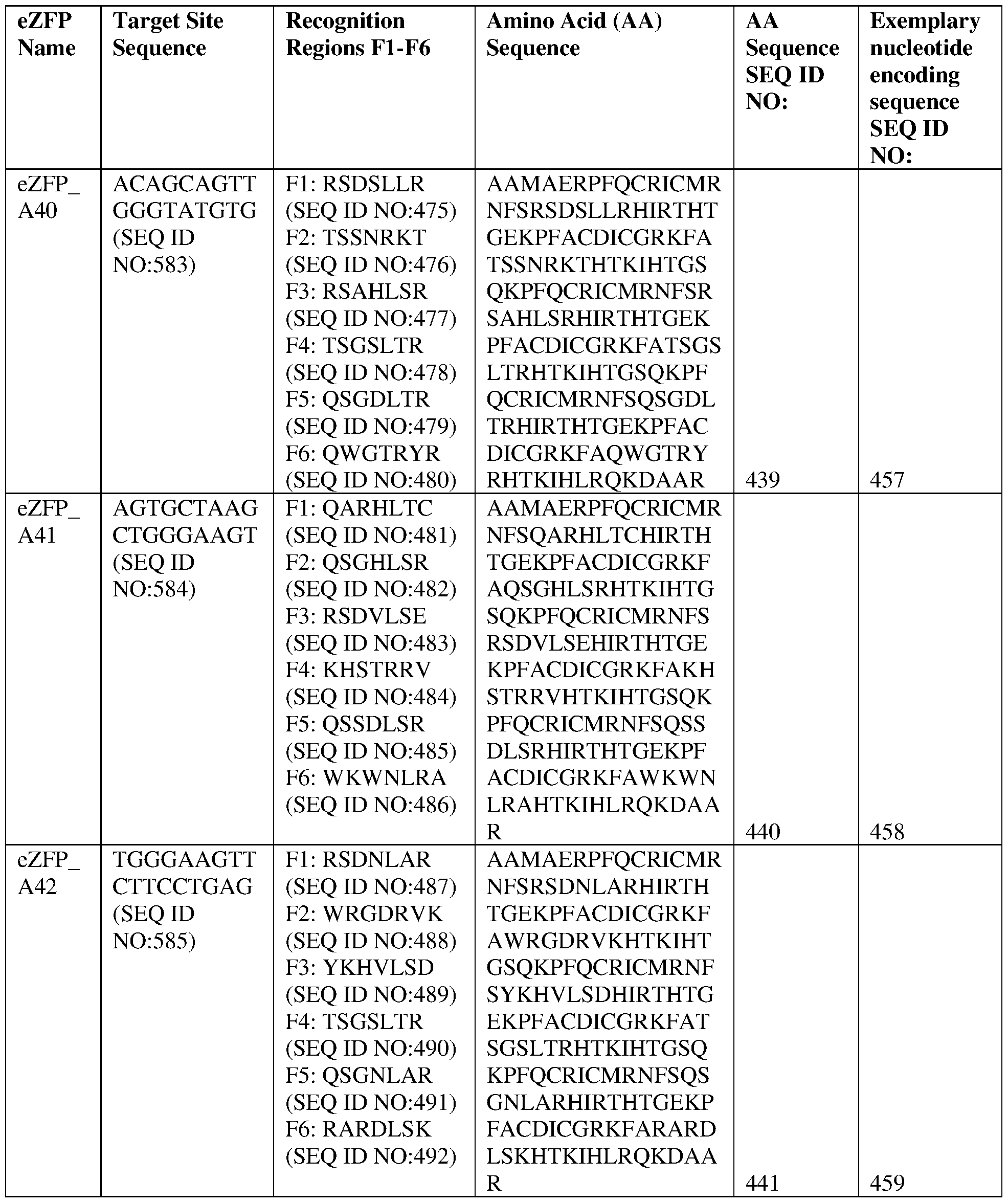
- the target site comprises the nucleotide sequence set forth in SEQ ID NO:583, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N- terminus to C-terminus, selected from F1-F6 as follows: Fl: RSDSLLR (SEQ ID NO:475); F2: TSSNRKT (SEQ ID NO:476); F3: RSAHLSR (SEQ ID NO:477); F4: TSGSLTR (SEQ ID NO:478); F5: QSGDLTR (SEQ ID NO:479); and F6: QWGTRYR (SEQ ID NO:480).
- an engineered zinc finger protein that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RSDSLLR (SEQ ID NO:475); F2: TSSNRKT (SEQ ID NO:476); F3: RSAHLSR (SEQ ID NO:477); F4: TSGSLTR (SEQ ID NO:478); F5: QSGDLTR (SEQ ID NO:479); and F6: QWGTRYR (SEQ ID NO:480).
- the eZFP comprises the sequence set forth in SEQ ID NO:439, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:439.
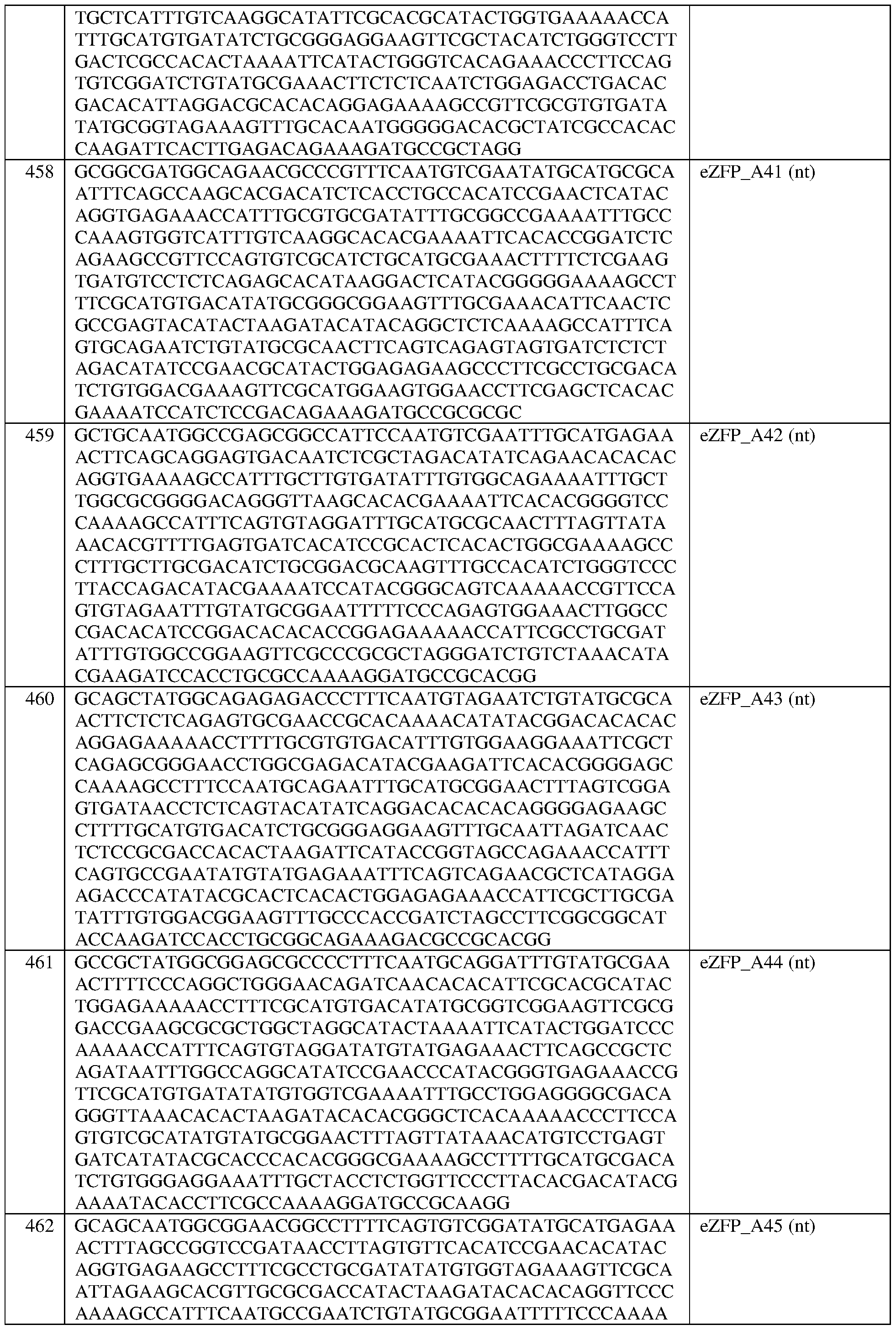
- the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:457 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:457.
- the target site comprises the nucleotide sequence set forth in SEQ ID NO:584, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N- terminus to C-terminus, selected from F1-F6 as follows: Fl: QARHLTC (SEQ ID NO:481); F2: QSGHLSR (SEQ ID NO:482); F3: RSDVLSE (SEQ ID NO:483); F4: KHSTRRV (SEQ ID NO:484); F5: QSSDLSR (SEQ ID NO:485); and F6: WKWNLRA (SEQ ID NO:486).
- an engineered zinc finger protein that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: QARHLTC (SEQ ID NO:481); F2: QSGHLSR (SEQ ID NO:482); F3: RSDVLSE (SEQ ID NO:483); F4: KHSTRRV (SEQ ID NO:484); F5: QSSDLSR (SEQ ID NO:485); and F6: WKWNLRA (SEQ ID NO:486).
- the eZFP comprises the sequence set forth in SEQ ID NO:440, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:440.
- the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:458 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 458.
- the target site comprises the nucleotide sequence set forth in SEQ ID NO:585, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N- terminus to C-terminus, selected from F1-F6 as follows: Fl: RSDNLAR (SEQ ID NO:487); F2: WRGDRVK (SEQ ID NO:488); F3: YKHVLSD (SEQ ID NO:489); F4: TSGSLTR (SEQ ID NO:490); F5: QSGNLAR (SEQ ID NO:491); and F6: RARDLSK (SEQ ID NO:492).
- an engineered zinc finger protein that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RSDNLAR (SEQ ID NO:487); F2: WRGDRVK (SEQ ID NO:488); F3: YKHVLSD (SEQ ID NO:489); F4: TSGSLTR (SEQ ID NO:490); F5: QSGNLAR (SEQ ID NO:491); and F6: RARDLSK (SEQ ID NO:492).
- the eZFP comprises the sequence set forth in SEQ ID NO:441, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:441.
- the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:459 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:459.
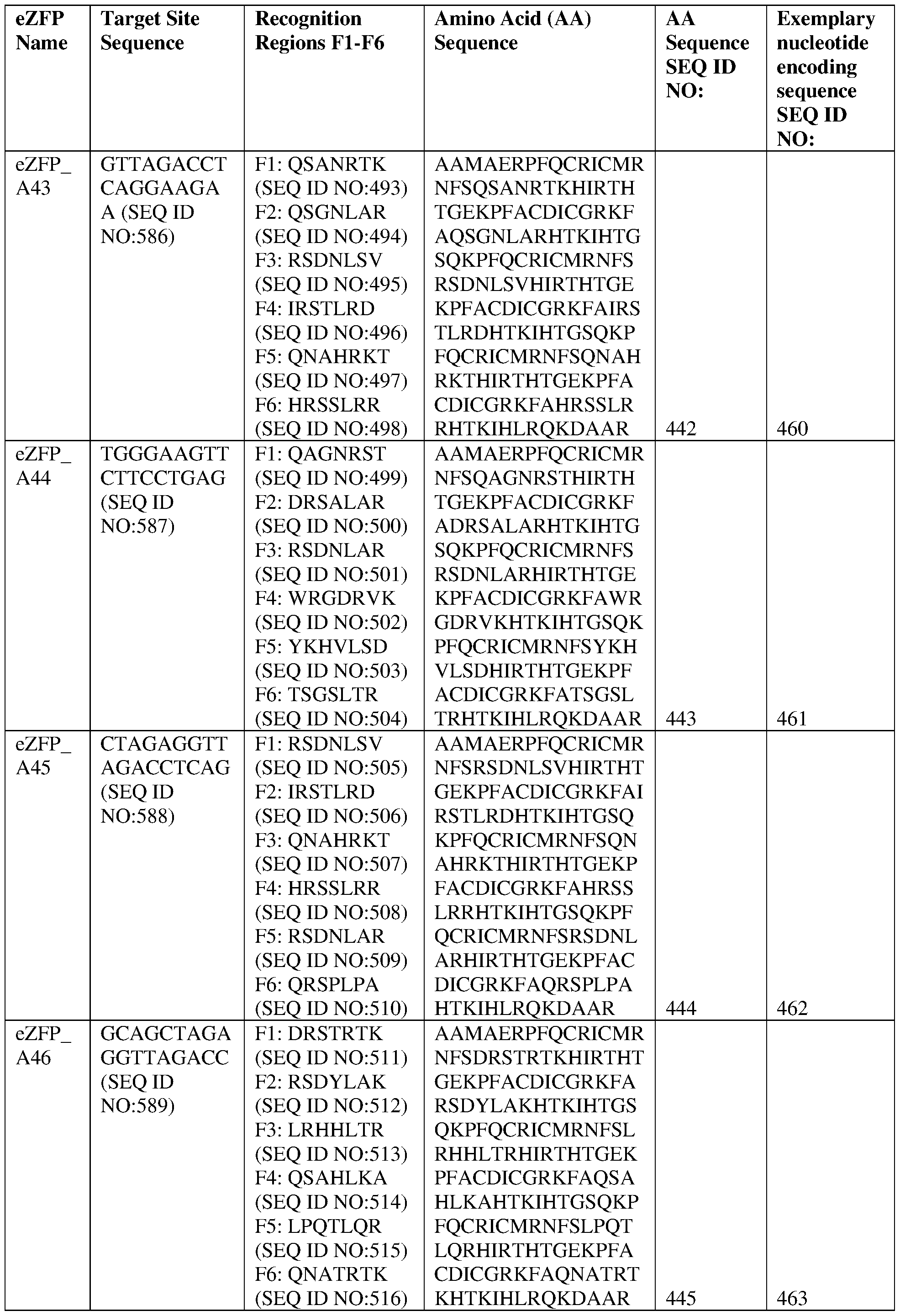
- the target site comprises the nucleotide sequence set forth in SEQ ID NO:586, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N- terminus to C-terminus, selected from F1-F6 as follows: Fl: QSANRTK (SEQ ID NO:493); F2: QSGNEAR (SEQ ID NO:494); F3: RSDNLSV (SEQ ID NO:495); F4: IRSTLRD (SEQ ID NO:496); F5: QNAHRKT (SEQ ID NO:497); and F6: HRSSLRR (SEQ ID NO:498).
- an engineered zinc finger protein that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: QSANRTK (SEQ ID NO:493); F2: QSGNLAR (SEQ ID NO:494); F3: RSDNLSV (SEQ ID NO:495); F4: IRSTLRD (SEQ ID NO:496); F5: QNAHRKT (SEQ ID NO:497); and F6: HRSSLRR (SEQ ID NO:498).
- the eZFP comprises the sequence set forth in SEQ ID NO:442, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:442.
- the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:460 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:460.
- the target site comprises the nucleotide sequence set forth in SEQ ID NO:587, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N- terminus to C-terminus, selected from F1-F6 as follows: Fl: QAGNRST (SEQ ID NO:499); F2: DRSALAR (SEQ ID NG:500); F3: RSDNLAR (SEQ ID NO:501); F4: WRGDRVK (SEQ ID NO:502); F5: YKHVLSD (SEQ ID NO:503); and F6: TSGSLTR (SEQ ID NO:504).
- an engineered zinc finger protein that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: QAGNRST (SEQ ID NO:499); F2: DRSALAR (SEQ ID NG:500); F3: RSDNLAR (SEQ ID NO:501); F4: WRGDRVK (SEQ ID NO:502); F5: YKHVLSD (SEQ ID NO:503); and F6: TSGSLTR (SEQ ID NO:504).
- the eZFP comprises the sequence set forth in SEQ ID NO:443, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:443.
- the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:461 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 461.
- the target site comprises the nucleotide sequence set forth in SEQ ID NO:588, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N- terminus to C-terminus, selected from F1-F6 as follows: Fl: RSDNLSV (SEQ ID NO:505); F2: IRSTLRD (SEQ ID NO:506); F3: QNAHRKT (SEQ ID NO:507); F4: HRSSLRR (SEQ ID NO:508); F5: RSDNLAR (SEQ ID NO:509); and F6: QRSPLPA (SEQ ID NO:510).
- an engineered zinc finger protein that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RSDNLSV (SEQ ID NO:505); F2: IRSTLRD (SEQ ID NO:506); F3: QNAHRKT (SEQ ID NO:507); F4: HRSSLRR (SEQ ID NO:508); F5: RSDNLAR (SEQ ID NO:509); and F6: QRSPLPA (SEQ ID NO:510).
- the eZFP comprises the sequence set forth in SEQ ID NO:444, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:444.
- the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:462 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:462.
- the target site comprises the nucleotide sequence set forth in SEQ ID NO:589, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N- terminus to C-terminus, selected from F1-F6 as follows: Fl: DRSTRTK (SEQ ID NO:511); F2: RSDYLAK (SEQ ID NO:512); F3: LRHHLTR (SEQ ID NO:513); F4: QSAHLKA (SEQ ID NO:514); F5: LPQTLQR (SEQ ID NO:515); and F6: QNATRTK (SEQ ID NO:516).
- an engineered zinc finger protein that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: DRSTRTK (SEQ ID NO:511); F2: RSDYLAK (SEQ ID NO:512); F3: LRHHLTR (SEQ ID NO:513); F4: QSAHLKA (SEQ ID NO:514); F5: LPQTLQR (SEQ ID NO:515); and F6: QNATRTK (SEQ ID NO:516).
- the eZFP comprises the sequence set forth in SEQ ID NO:445, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:445.
- the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:463 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 463.
- the target site comprises the nucleotide sequence set forth in SEQ ID NO:590, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
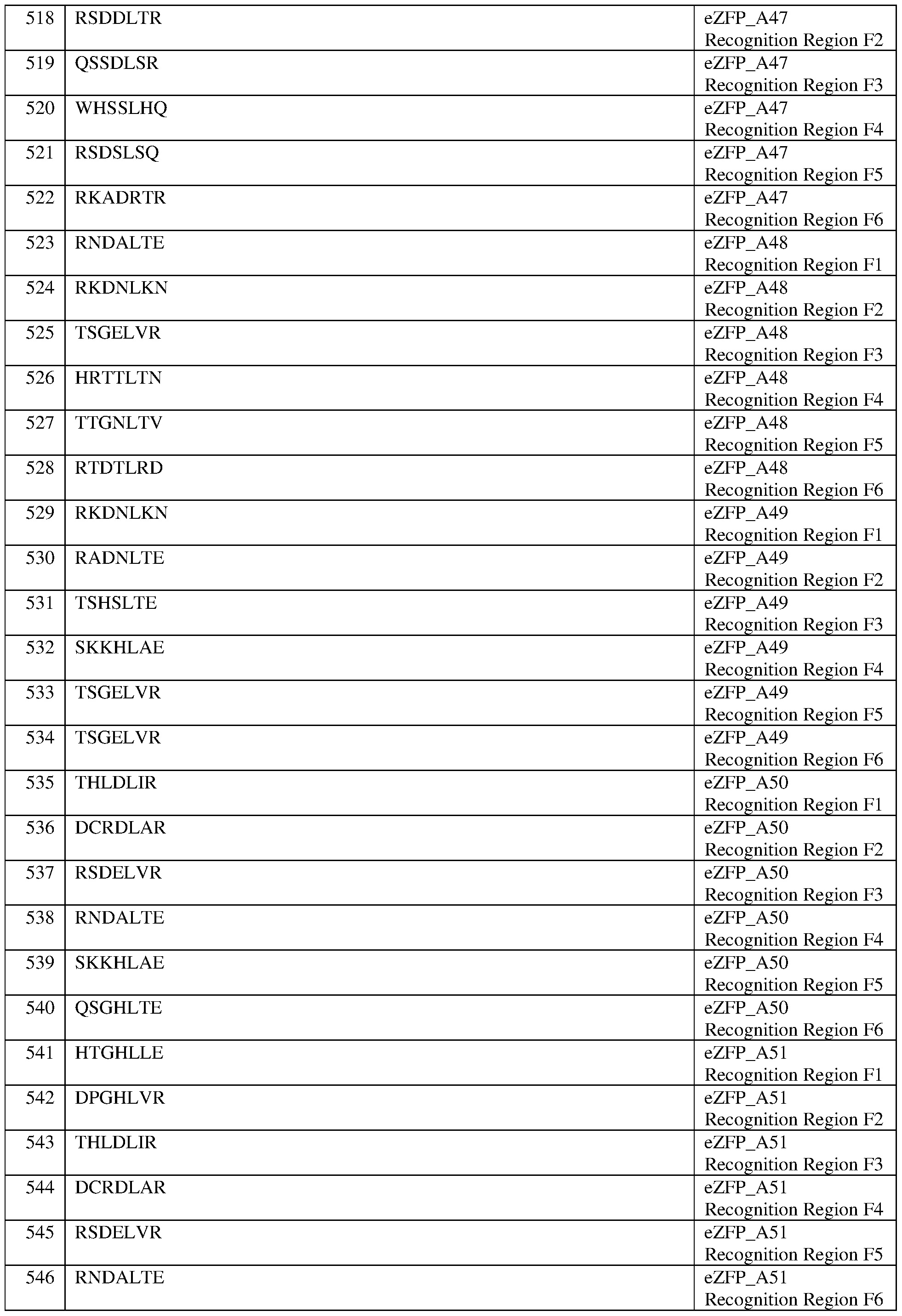
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N- terminus to C-terminus, selected from F1-F6 as follows: Fl: RSADLTR (SEQ ID NO:517); F2: RSDDLTR (SEQ ID NO:518); F3: QSSDLSR (SEQ ID NO:519); F4: WHSSLHQ (SEQ ID NO:520); F5: RSDSLSQ (SEQ ID NO:521); and F6: RKADRTR (SEQ ID NO:522).
- an engineered zinc finger protein that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RSADLTR (SEQ ID NO:517); F2: RSDDLTR (SEQ ID NO:518); F3: QSSDLSR (SEQ ID NO:519); F4: WHSSLHQ (SEQ ID NO:520); F5: RSDSLSQ (SEQ ID NO:521); and F6: RKADRTR (SEQ ID NO:522).
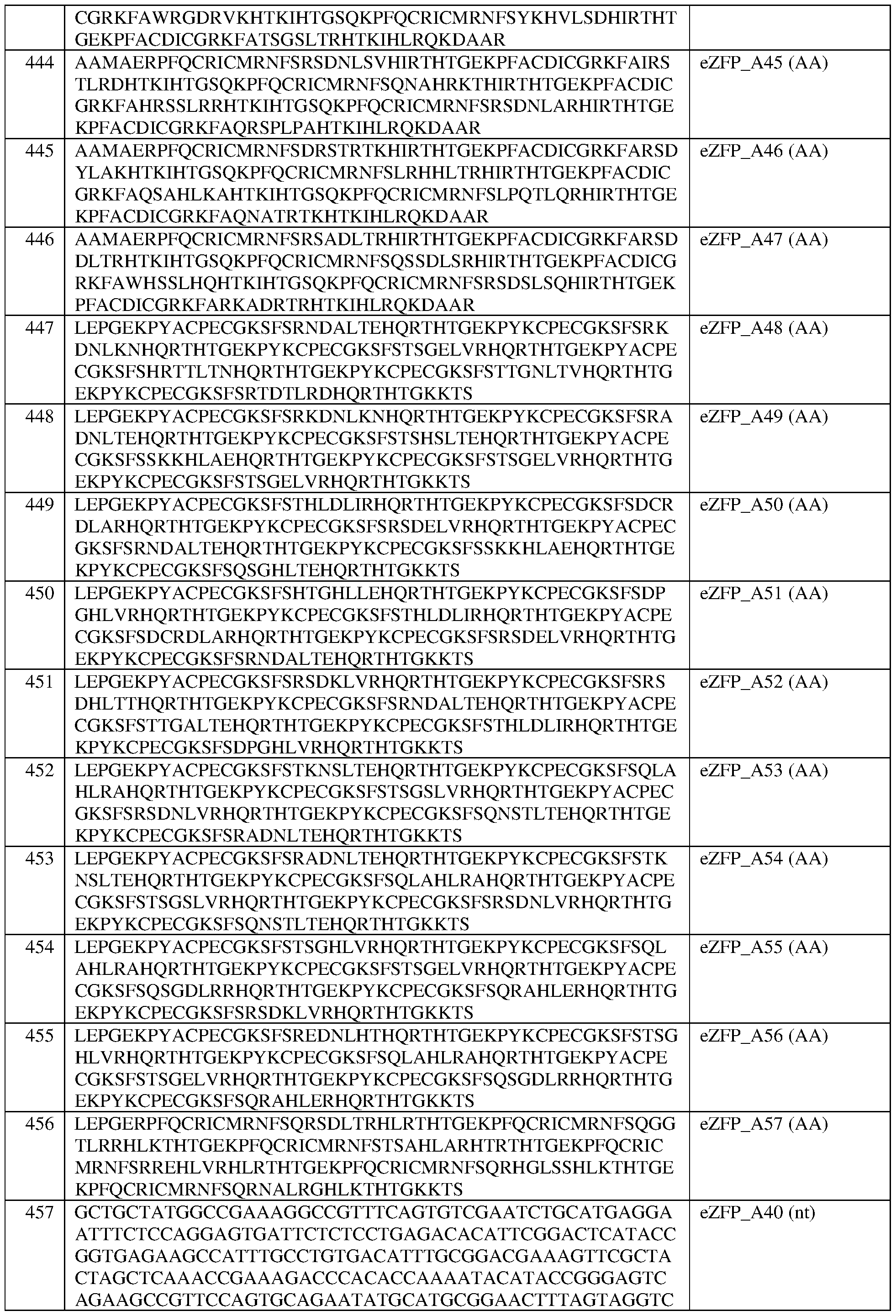
- the eZFP comprises the sequence set forth in SEQ ID NO:446, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:446.
- the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:464 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:464.
- the target site comprises the nucleotide sequence set forth in SEQ ID NO:591, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N- terminus to C-terminus, selected from F1-F6 as follows: Fl: RNDALTE (SEQ ID NO:523); F2: RKDNLKN (SEQ ID NO:524); F3: TSGELVR (SEQ ID NO:525); F4: HRTTLTN (SEQ ID NO:526); F5: TTGNLTV (SEQ ID NO:527); and F6: RTDTLRD (SEQ ID NO:528).
- an engineered zinc finger protein that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RNDALTE (SEQ ID NO:523); F2: RKDNLKN (SEQ ID NO:524); F3: TSGELVR (SEQ ID NO:525); F4: HRTTLTN (SEQ ID NO:526); F5: TTGNLTV (SEQ ID NO:527); and F6: RTDTLRD (SEQ ID NO:528).
- the eZFP comprises the sequence set forth in SEQ ID NO:447, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:447.
- the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:465 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 465.
- the target site comprises the nucleotide sequence set forth in SEQ ID NO:592, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N- terminus to C-terminus, selected from F1-F6 as follows: Fl: RKDNLKN (SEQ ID NO:529); F2: RADNLTE (SEQ ID NO:530); F3: TSHSLTE (SEQ ID NO:531); F4: SKKHLAE (SEQ ID NO:532); F5: TSGELVR (SEQ ID NO:533); and F6: TSGELVR (SEQ ID NO:534).
- an engineered zinc finger protein that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RKDNLKN (SEQ ID NO:529); F2: RADNLTE (SEQ ID NO:530); F3: TSHSLTE (SEQ ID NO:531); F4: SKKHLAE (SEQ ID NO:532); F5: TSGELVR (SEQ ID NO:533); and F6: TSGELVR (SEQ ID NO:534).
- the eZFP comprises the sequence set forth in SEQ ID NO:448, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:448.
- the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:466 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:466.
- the target site comprises the nucleotide sequence set forth in SEQ ID NO:593, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N- terminus to C-terminus, selected from F1-F6 as follows: Fl: THLDLIR (SEQ ID NO:535); F2: DCRDLAR (SEQ ID NO:536); F3: RSDELVR (SEQ ID NO:537); F4: RNDALTE (SEQ ID NO:538); F5: SKKHLAE (SEQ ID NO:539); and F6: QSGHLTE (SEQ ID NO:540).
- an engineered zinc finger protein that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: THLDLIR (SEQ ID NO:535); F2: DCRDLAR (SEQ ID NO:536); F3: RSDELVR (SEQ ID NO:537); F4: RNDALTE (SEQ ID NO:538); F5: SKKHLAE (SEQ ID NO:539); and F6: QSGHLTE (SEQ ID NO:540).
- the eZFP comprises the sequence set forth in SEQ ID NO:449, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:449.
- the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:467 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 467.
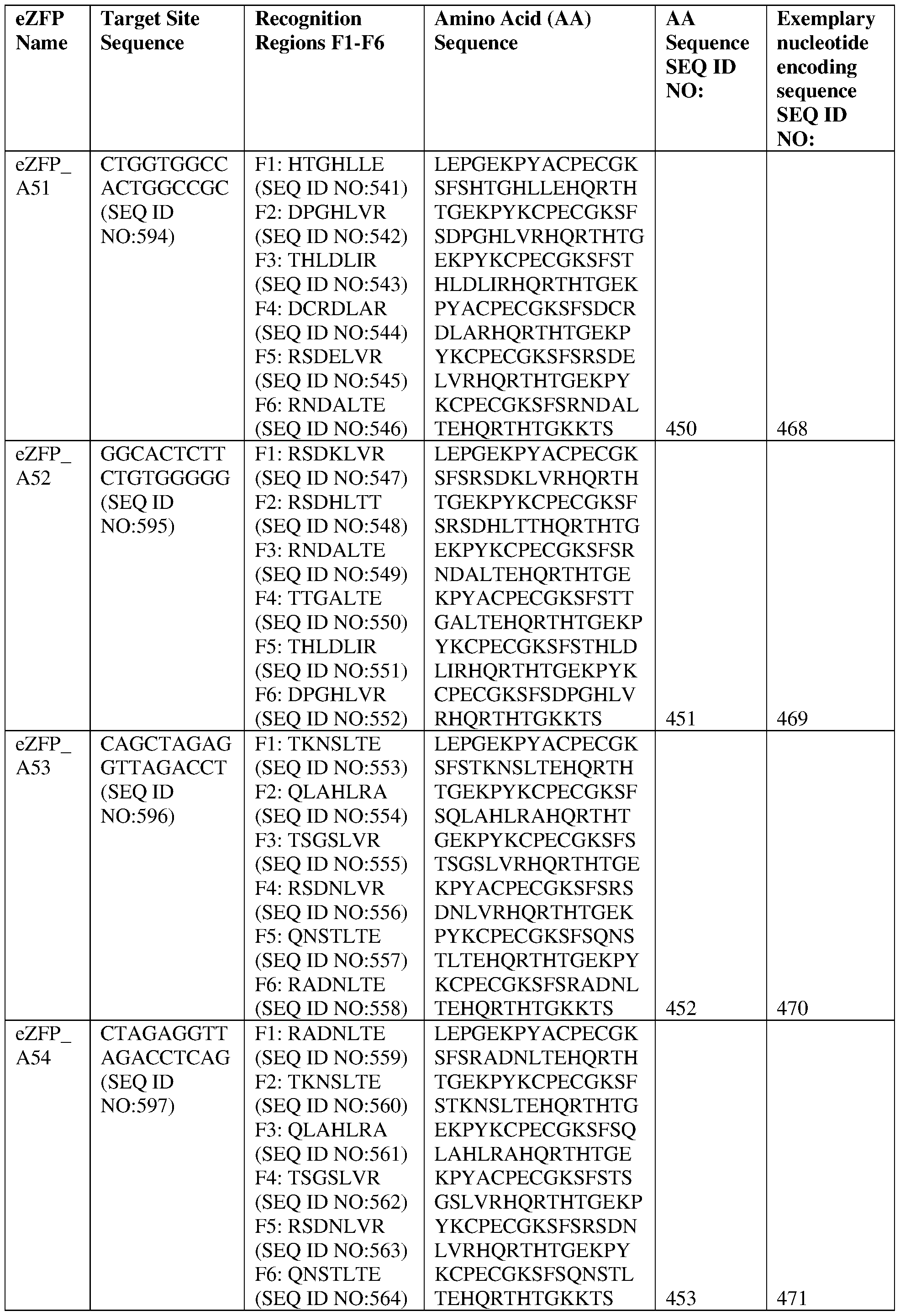
- the target site comprises the nucleotide sequence set forth in SEQ ID NO:594, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N- terminus to C-terminus, selected from F1-F6 as follows: Fl: HTGHLLE (SEQ ID NO:541); F2: DPGHLVR (SEQ ID NO:542); F3: THLDLIR (SEQ ID NO:543); F4: DCRDLAR (SEQ ID NO:544); F5: RSDELVR (SEQ ID NO:545); and F6: RNDALTE (SEQ ID NO:546).
- an engineered zinc finger protein that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: HTGHLLE (SEQ ID NO:541); F2: DPGHLVR (SEQ ID NO:542); F3: THLDLIR (SEQ ID NO:543); F4: DCRDLAR (SEQ ID NO:544); F5: RSDELVR (SEQ ID NO:545); and F6: RNDALTE (SEQ ID NO:546).
- the eZFP comprises the sequence set forth in SEQ ID NO:450, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:450.
- the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:468 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:468.
- the target site comprises the nucleotide sequence set forth in SEQ ID NO:595, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N- terminus to C-terminus, selected from F1-F6 as follows Fl: RSDKLVR (SEQ ID NO:547); F2: RSDHLTT (SEQ ID NO:548); F3: RNDALTE (SEQ ID NO:549); F4: TTGALTE (SEQ ID NO:550); F5: THLDLIR (SEQ ID NO:551); and F6: DPGHLVR (SEQ ID NO:552).
- an engineered zinc finger protein that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RSDKLVR (SEQ ID NO:547); F2: RSDHLTT (SEQ ID NO:548); F3: RNDALTE (SEQ ID NO:549); F4: TTGALTE (SEQ ID NO:550); F5: THLDLIR (SEQ ID NO:551); and F6: DPGHLVR (SEQ ID NO:552).
- the eZFP comprises the sequence set forth in SEQ ID NO:451, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:451.
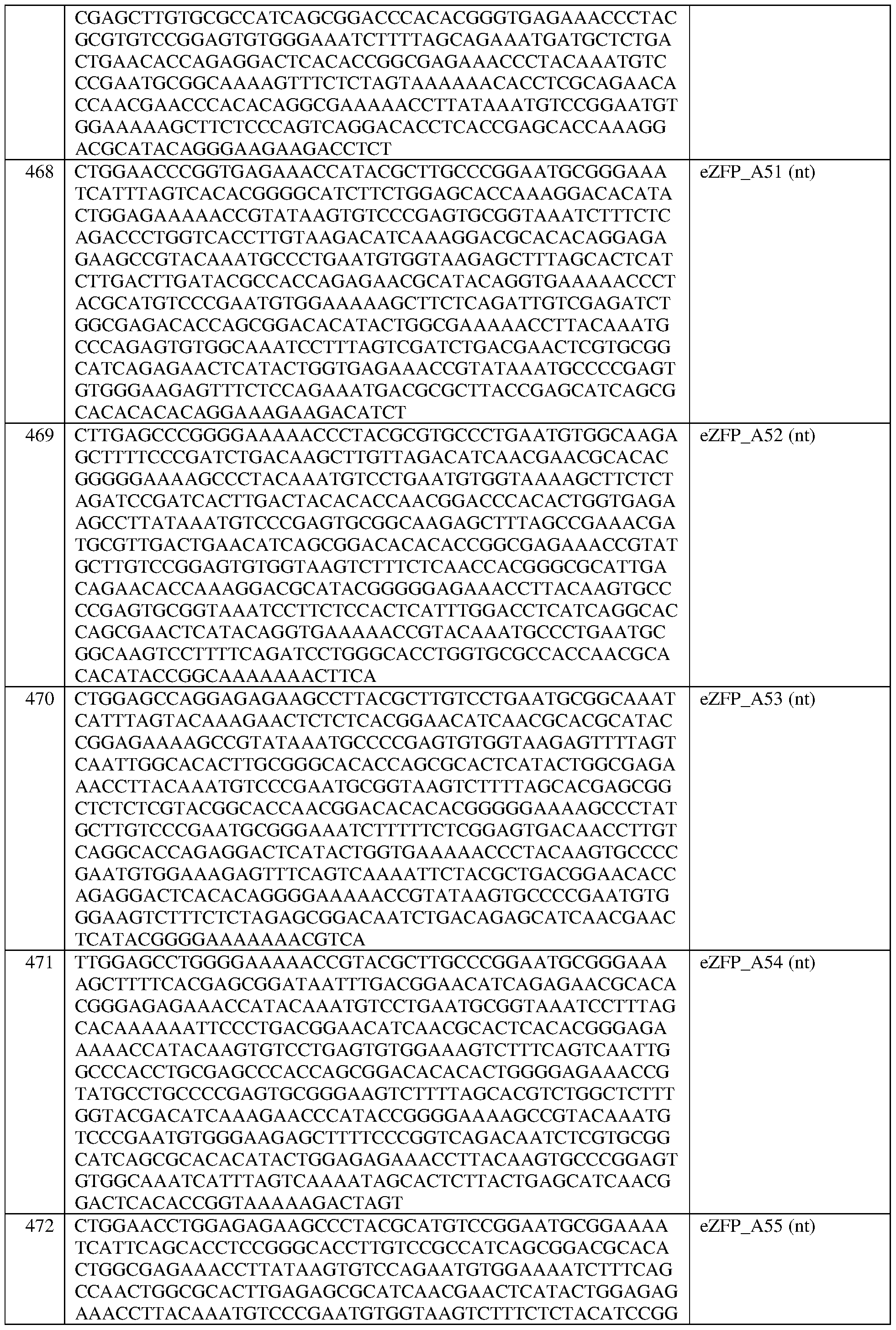
- the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:469 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 469.
- the target site comprises the nucleotide sequence set forth in SEQ ID NO:596, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N- terminus to C-terminus, selected from F1-F6 as follows Fl: TKNSLTE (SEQ ID NO:553); F2: QLAHLRA (SEQ ID NO:554); F3: TSGSLVR (SEQ ID NO:555); F4: RSDNLVR (SEQ ID NO:556); F5: QNSTLTE (SEQ ID NO:557); and F6: RADNLTE (SEQ ID NO:558).
- an engineered zinc finger protein that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: TKNSLTE (SEQ ID NO:553); F2: QLAHLRA (SEQ ID NO:554); F3: TSGSLVR (SEQ ID NO:555); F4: RSDNLVR (SEQ ID NO:556); F5: QNSTLTE (SEQ ID NO:557); and F6: RADNLTE (SEQ ID NO:558).
- the eZFP comprises the sequence set forth in SEQ ID NO:452, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:452.
- the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:470 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:470.
- the target site comprises the nucleotide sequence set forth in SEQ ID NO:597, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N- terminus to C-terminus, selected from F1-F6 as follows Fl: RADNETE (SEQ ID NO:559); F2: TKNSETE (SEQ ID NO:560); F3: QLAHLRA (SEQ ID NO:561); F4: TSGSLVR (SEQ ID NO:562); F5: RSDNLVR (SEQ ID NO:563); and F6: QNSTLTE (SEQ ID NO:564).
- Fl RADNETE
- F2 TKNSETE
- F3 QLAHLRA
- F4 TSGSLVR
- F5 RSDNLVR
- F6 QNSTLTE
- an engineered zinc finger protein that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RADNLTE (SEQ ID NO:559); F2: TKNSLTE (SEQ ID NO:560); F3: QLAHLRA (SEQ ID NO:561); F4: TSGSLVR (SEQ ID NO:562); F5: RSDNLVR (SEQ ID NO:563); and F6: QNSTLTE (SEQ ID NO:564).
- the eZFP comprises the sequence set forth in SEQ ID NO:453, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:453.
- the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:471 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 471.
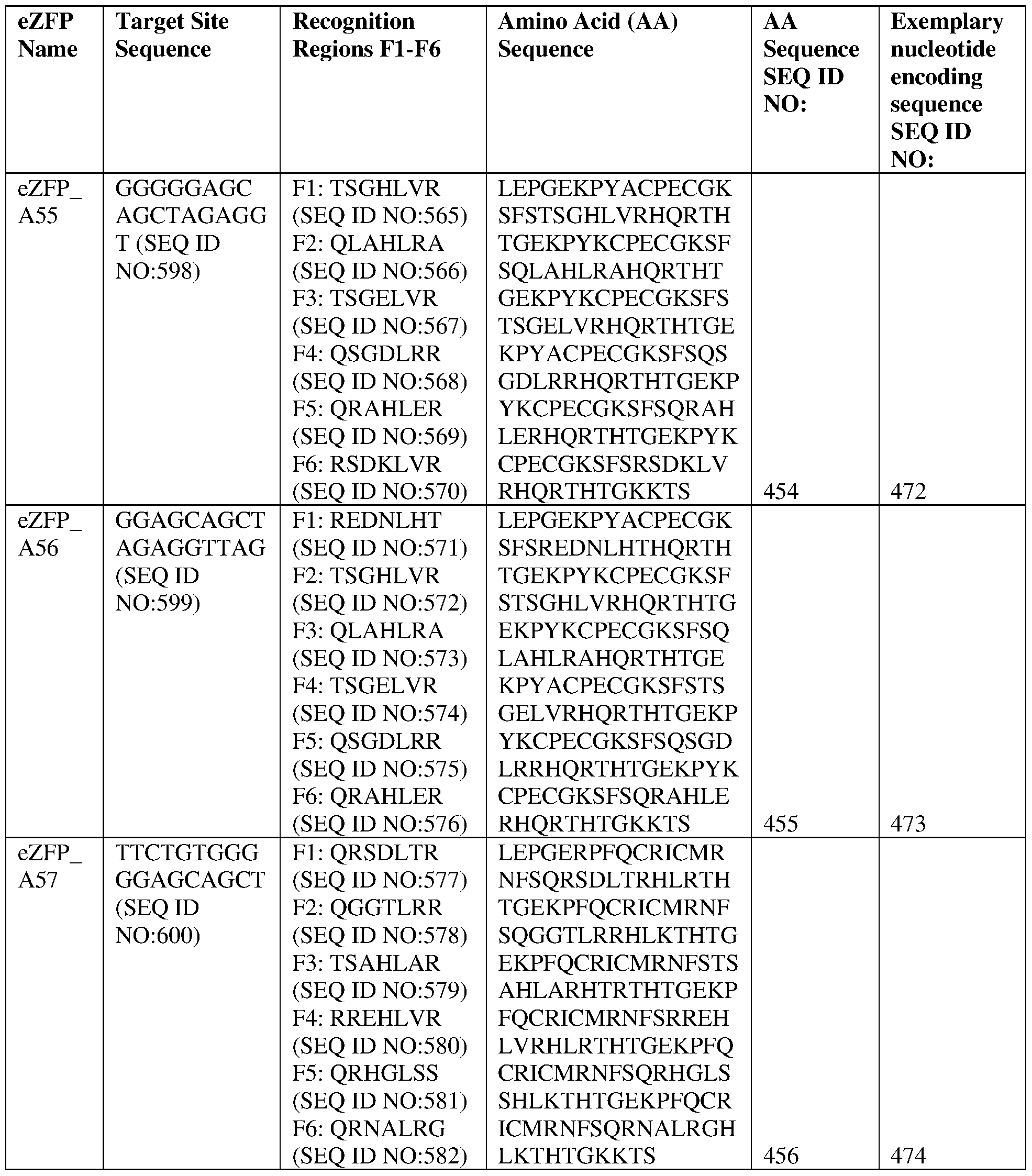
- the target site comprises the nucleotide sequence set forth in SEQ ID NO:598, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N- terminus to C-terminus, selected from F1-F6 as follows Fl: TSGHLVR (SEQ ID NO:565); F2: QLAHLRA (SEQ ID NO:566); F3: TSGELVR (SEQ ID NO:567); F4: QSGDLRR (SEQ ID NO:568); F5: QRAHLER (SEQ ID NO:569); and F6: RSDKLVR (SEQ ID NO:570).
- an engineered zinc finger protein that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: TSGHLVR (SEQ ID NO:565); F2: QLAHLRA (SEQ ID NO:566); F3: TSGELVR (SEQ ID NO:567); F4: QSGDLRR (SEQ ID NO:568); F5: QRAHLER (SEQ ID NO:569); and F6: RSDKLVR (SEQ ID NO:570).
- the eZFP comprises the sequence set forth in SEQ ID NO:454, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:454.
- the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:472 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:472.
- the target site comprises the nucleotide sequence set forth in SEQ ID NO:599, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N- terminus to C-terminus, selected from F1-F6 as follows Fl: REDNEHT (SEQ ID NO:571); F2: TSGHEVR (SEQ ID NO:572); F3: QLAHLRA (SEQ ID NO:573); F4: TSGELVR (SEQ ID NO:574); F5: QSGDLRR (SEQ ID NO:575); and F6: QRAHLER (SEQ ID NO:576).
- Fl REDNEHT
- F2 TSGHEVR
- F3 QLAHLRA
- F4 TSGELVR
- F5 QSGDLRR
- F6 QRAHLER
- an engineered zinc finger protein that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: REDNLHT (SEQ ID NO:571); F2: TSGHLVR (SEQ ID NO:572); F3: QLAHLRA (SEQ ID NO:573); F4: TSGELVR (SEQ ID NO:574); F5: QSGDLRR (SEQ ID NO:575); and F6: QRAHLER (SEQ ID NO:576).
- the eZFP comprises the sequence set forth in SEQ ID NO:455, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:455.
- the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:473 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 473.
- the target site comprises the nucleotide sequence set forth in SEQ ID NO:600, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
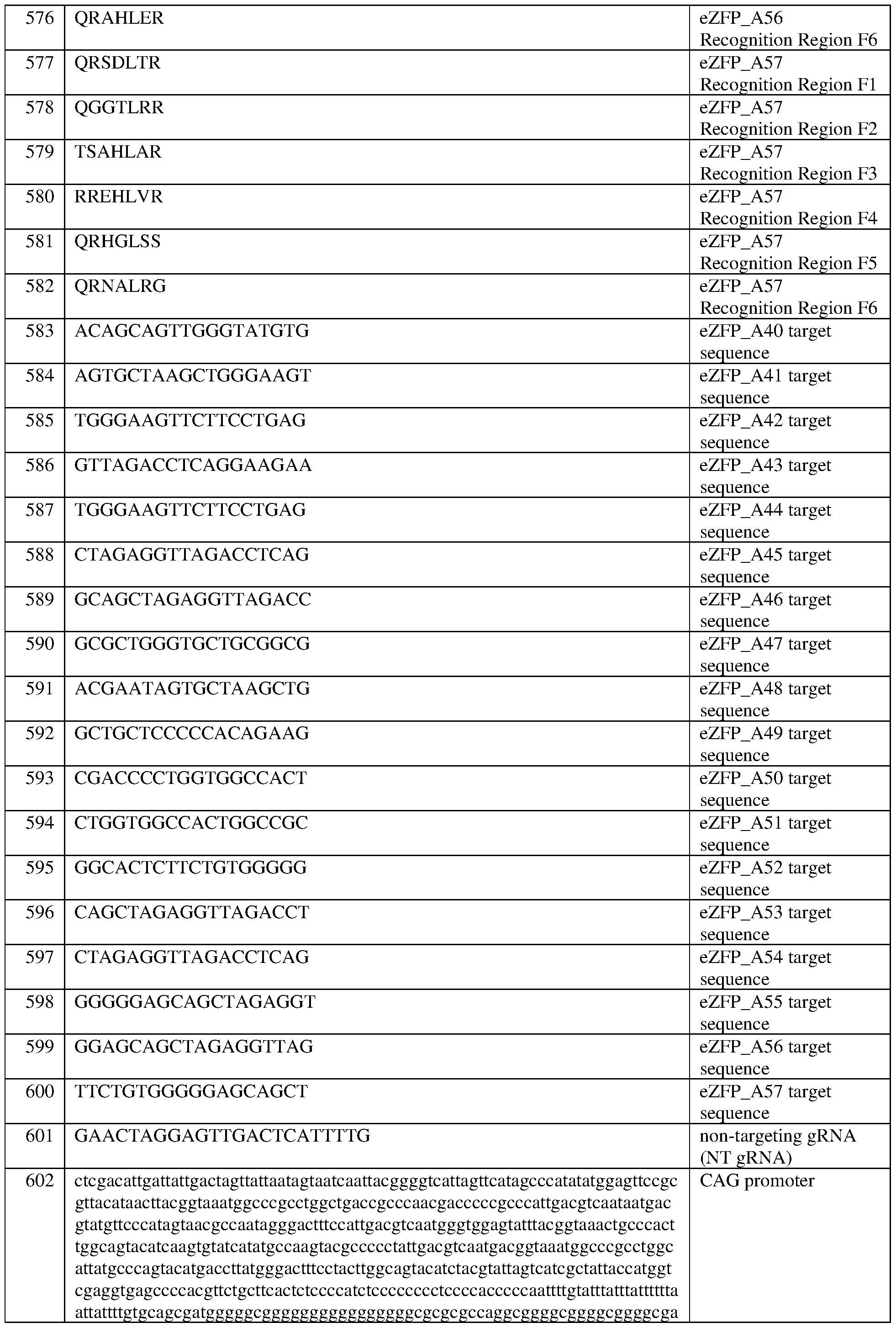
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N- terminus to C-terminus, selected from F1-F6 as follows Fl: QRSDLTR (SEQ ID NO:577); F2: QGGTLRR (SEQ ID NO:578); F3: TSAHLAR (SEQ ID NO:579); F4: RREHLVR (SEQ ID NO:580); F5: QRHGLSS (SEQ ID NO:581); and F6: QRNALRG (SEQ ID NO:582).
- an engineered zinc finger protein that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus
- the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: QRSDLTR (SEQ ID NO:577); F2: QGGTLRR (SEQ ID NO:578); F3: TSAHLAR (SEQ ID NO:579); F4: RREHLVR (SEQ ID NO:580); F5: QRHGLSS (SEQ ID NO:581); and F6: QRNALRG (SEQ ID NO:582).
- the eZFP comprises the sequence set forth in SEQ ID NO:456, or a portion thereof, or an amino acid sequence that has at least 90%, 91%,
- the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:456. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:474 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:474.
- a fusion protein comprising any of the engineered zinc finger proteins (eZFPs) provided herein.
- a fusion protein comprising:
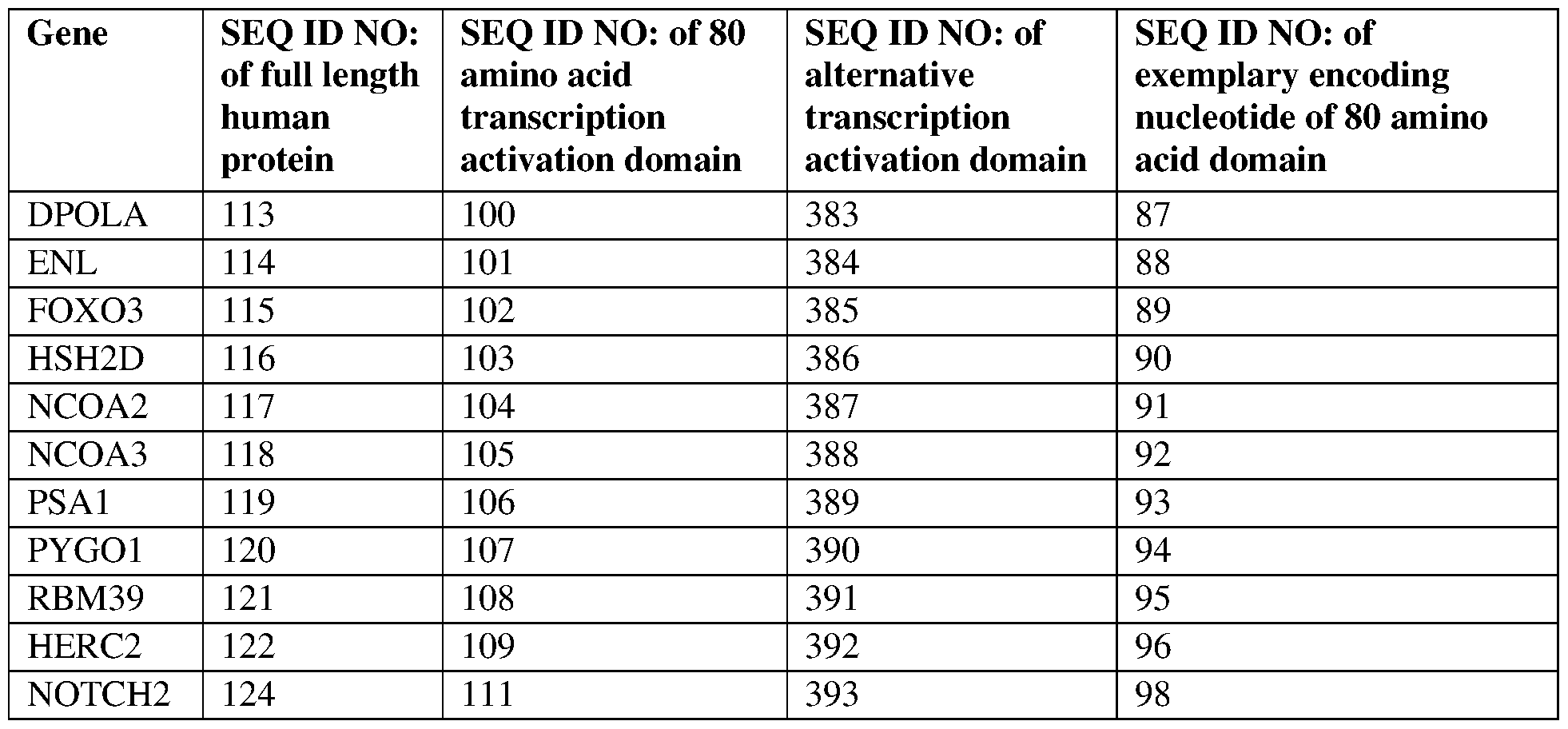
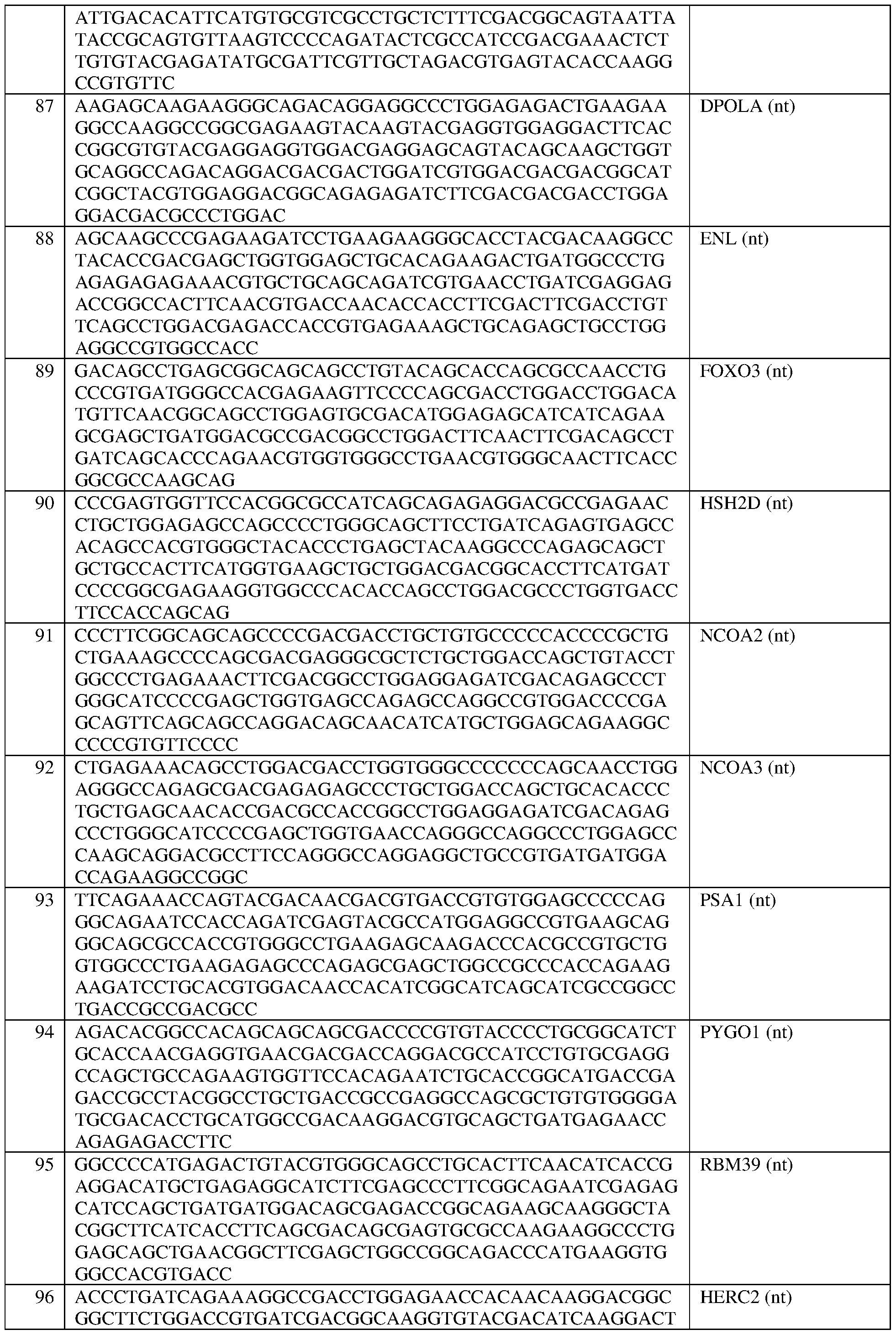
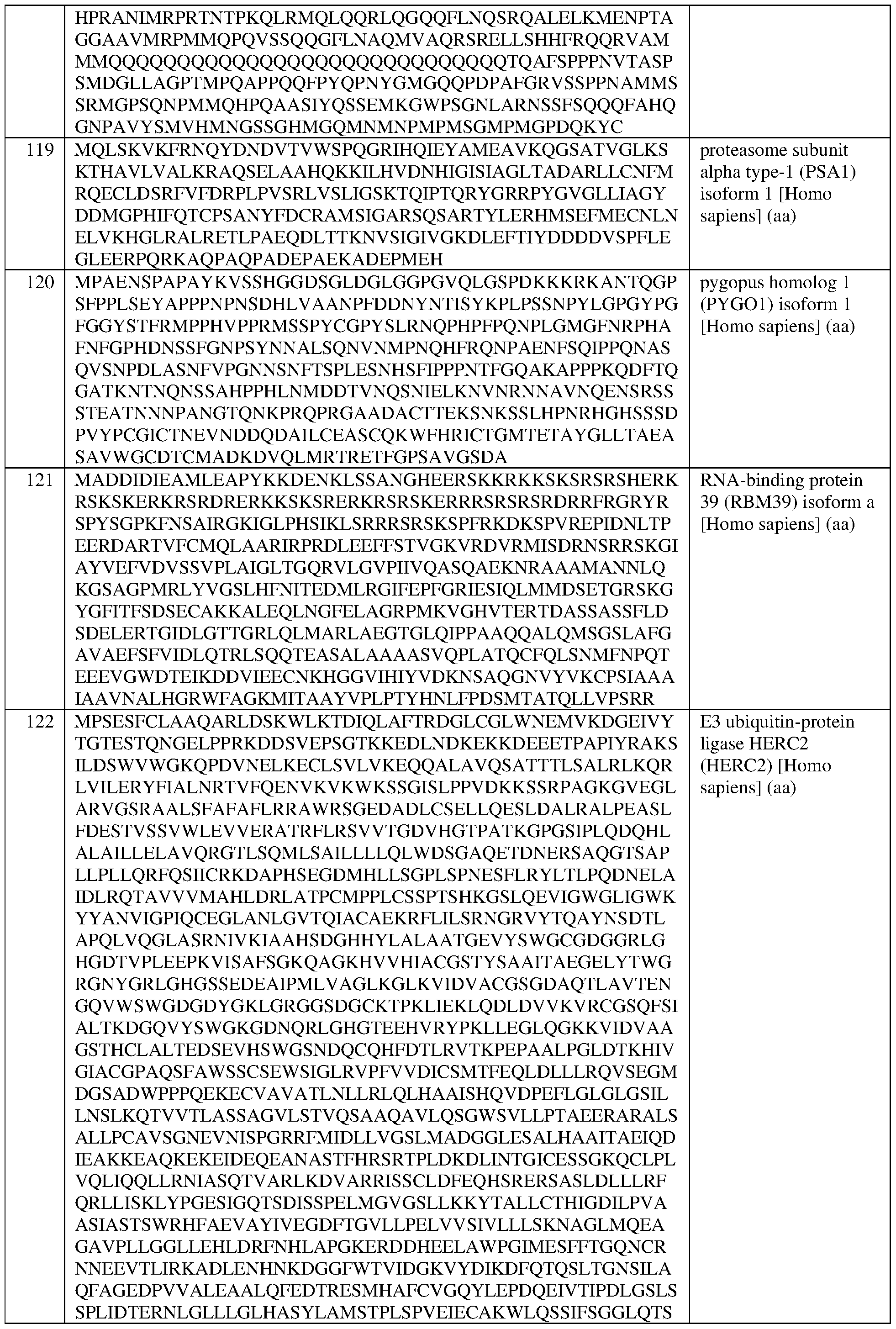
- the at least one epigenetic effector domain comprises: a VP64 domain, a p65 activation domain, a p300 domain, an Rta domain, a CBP domain, a VPR domain, a VPH domain, an HSF1 domain, a TET protein domain, optionally wherein the TET protein is TET1, a SunTag domain, a domain from DPOLA, ENL, FOXO3, HSH2D, NCOA2, NCOA3, PSA1, PYGO1, RBM39, HERC2, or NOTCH2, or a domain, portion, variant, or truncation of any of the foregoing.
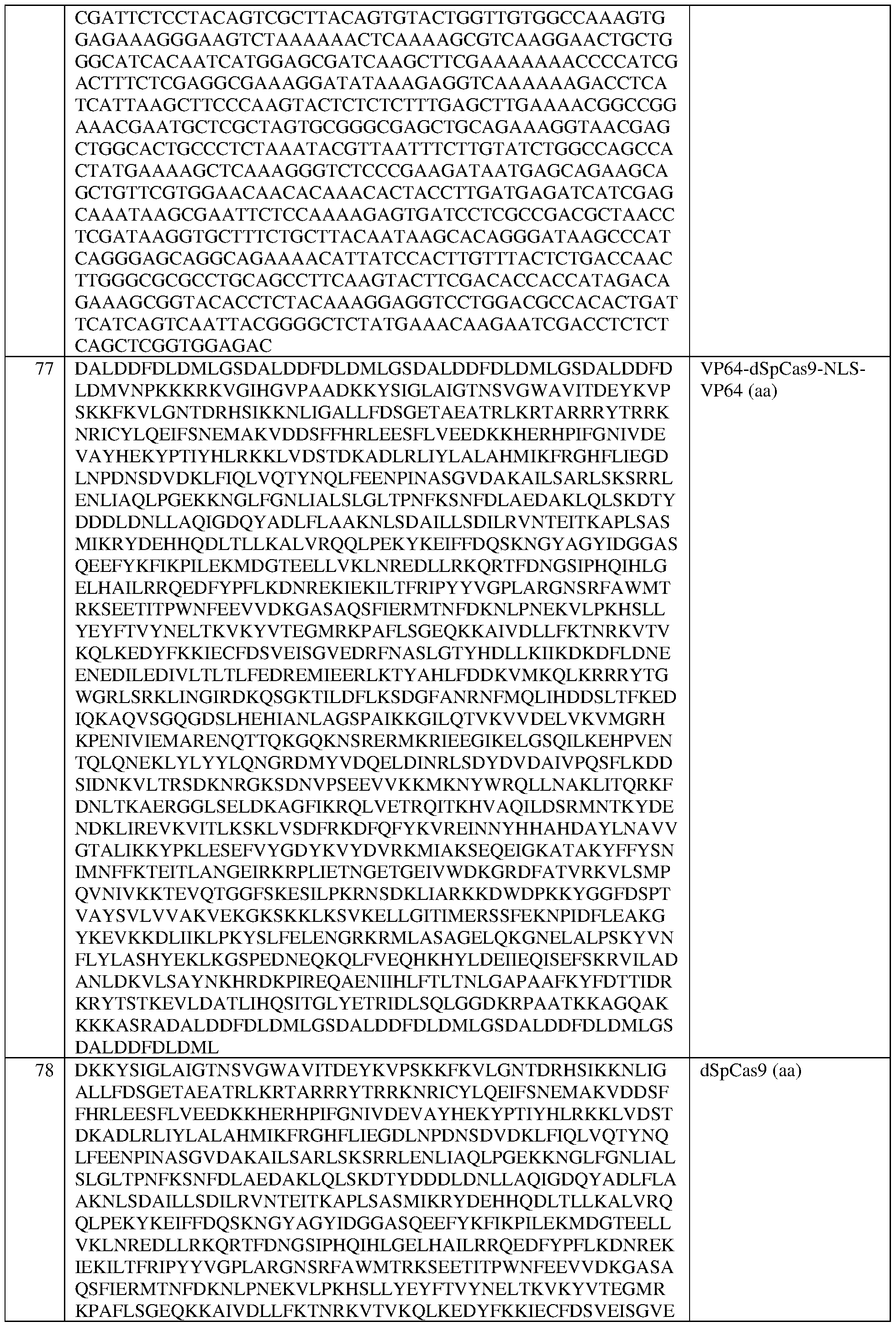
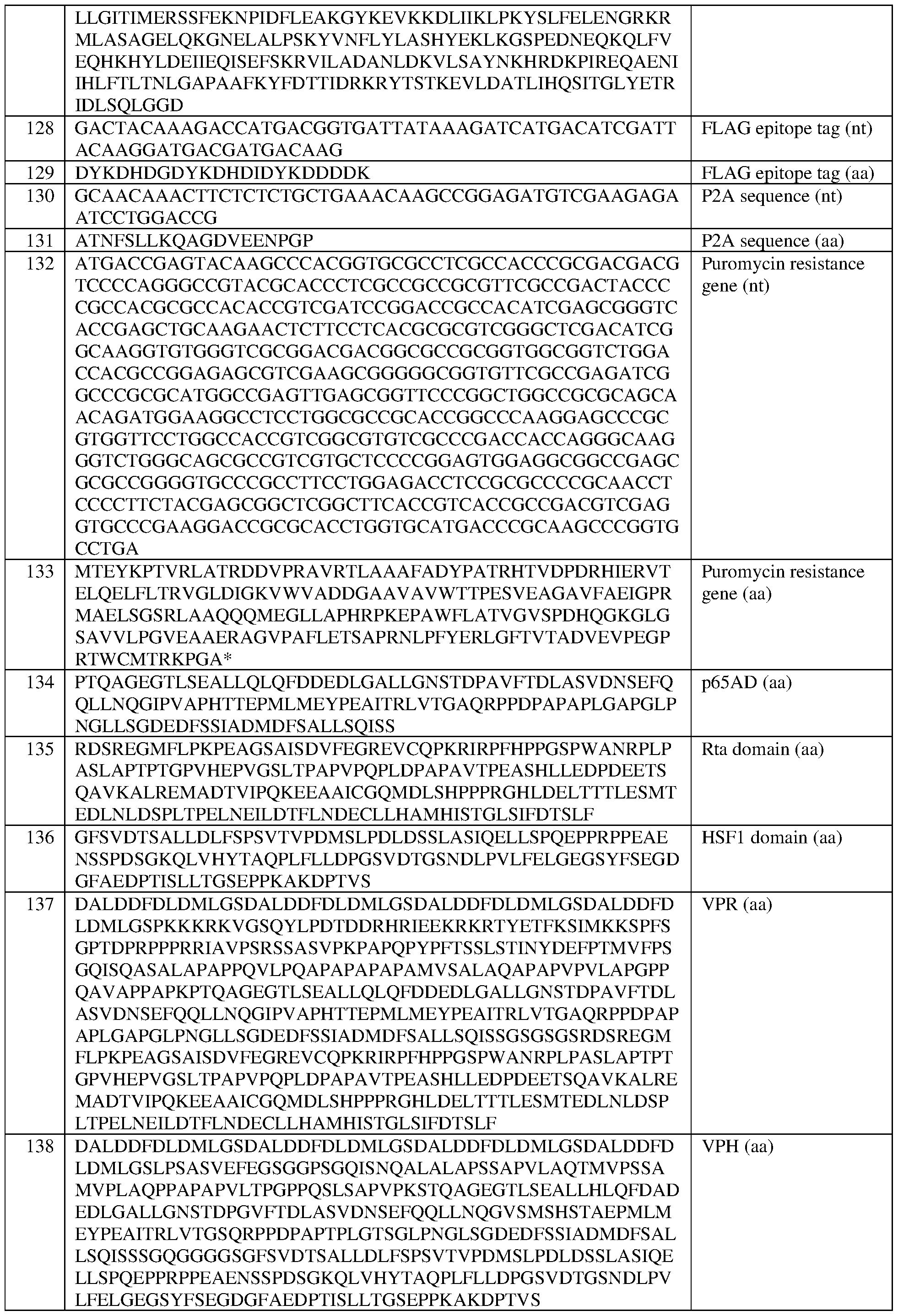
- the at least one epigenetic effector domain comprises the sequence set forth in any of SEQ ID NOS:81, 83, 100-109, 111-122, 124, 125, 134-140, 152, and 383-396, or a domain, portion, variant, or truncation thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
- the at least one effector domain comprises at least one VP16 domain, or a VP16 tetramer (“VP64”) or a variant thereof. In some of any of the provided embodiments, the at least one effector domain comprises VP64. In some of any of the provided embodiments, the at least one effector domain comprises a VP64 domain comprising the sequence set forth in SEQ ID NO:83, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing. In some of any of the provided embodiments, the at least one effector domain comprises a VP64 domain comprising the sequence set forth in SEQ ID NO: 83.
- the at least one epigenetic effector domain comprises: a domain from DPOLA, ENL, FOXO3, HSH2D, NCOA2, NCOA3, PSA1, PYGO1, RBM39, HERC2, or NOTCH2, or a domain, portion, variant, or truncation of any of the foregoing.
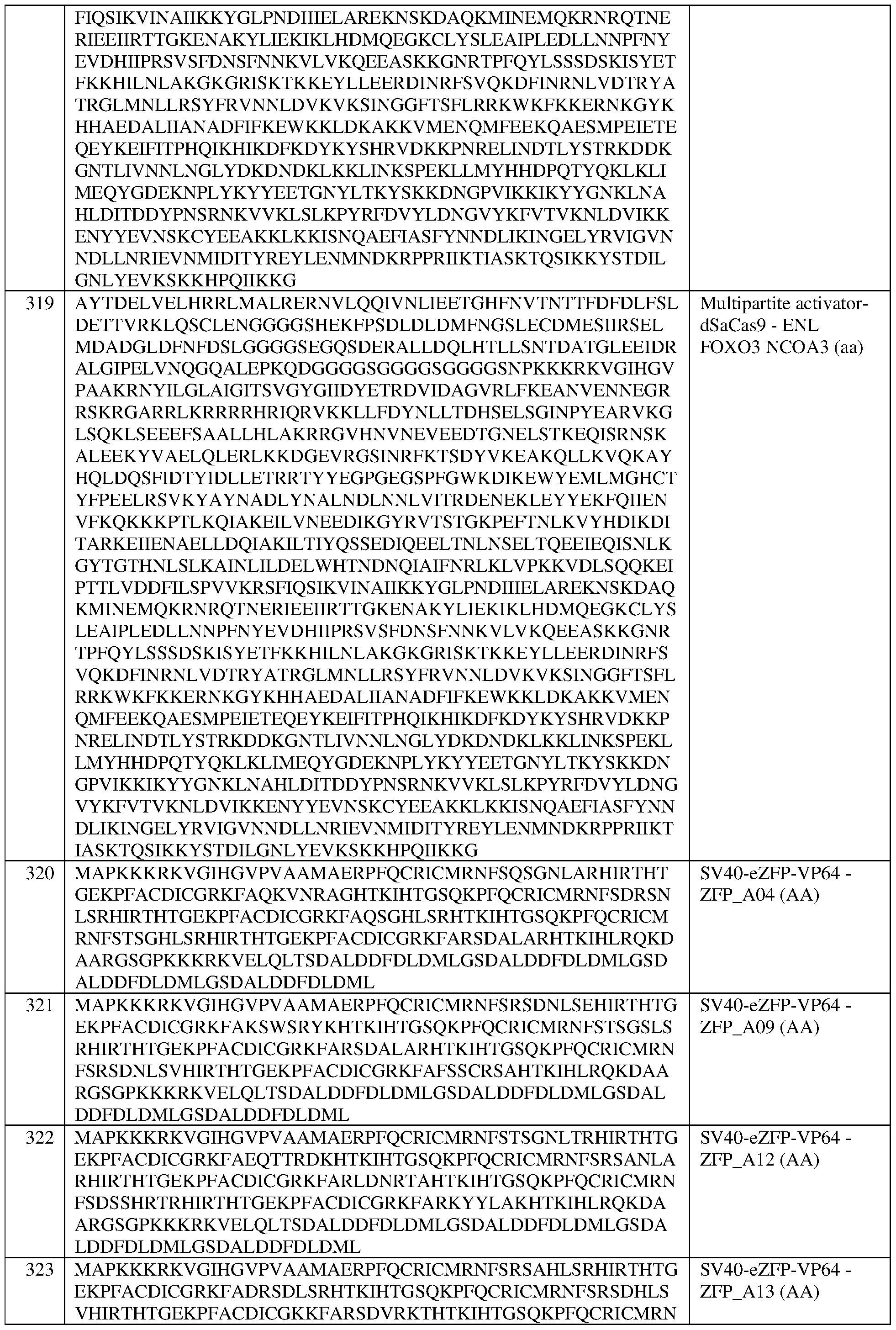
- the at least one epigenetic effector domain comprises the sequence set forth in any of SEQ ID NOS:383-393, or a domain, portion, variant, or truncation thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
- the at least one effector domain comprises a domain from NCOA2, NCOA3, FOXO3, PYGO1, or a portion or variant of any of the foregoing.
- each effector domain of the at least one effector domain is independently selected from an NCOA2 domain, an NCOA3 domain, a FOXO3 domain, and a PYGO1 domain.
- the at least one effector domain comprises a domain from NCOA2 comprising the sequence set forth in SEQ ID NO: 104 or SEQ ID NO:387, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the at least one effector domain comprises a domain from NCOA2 set forth in or SEQ ID NO:387.
- the at least one effector domain comprises a domain from NCOA3 comprising the sequence set forth in SEQ ID NO: 105 or SEQ ID NO:388, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the at least one effector domain comprises a domain from NCOA3 set forth in or SEQ ID NO:388.
- the at least one effector domain comprises a domain from FOXO3 comprising the sequence set forth in SEQ ID NO: 102 or SEQ ID NO:385, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the at least one effector domain comprises a domain from FOXO3 set forth in or SEQ ID NO:385.
- the at least one effector domain comprises a domain from PYGO1 comprising the sequence set forth in SEQ ID NO: 107 or SEQ ID NO:390, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the at least one effector domain comprises a domain from PYGO1 set forth in or SEQ ID NO:390.
- the at least one effector domain is a multipartite effector composed of at least two effector domains.
- the multipartite effector is composed of two effector domains. In some of any of the provided embodiments, the multipartite effector is composed of three effector domains. In some of any of the provided embodiments, the multipartite effector is set forth in any one of SEQ ID NOS:397-418, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
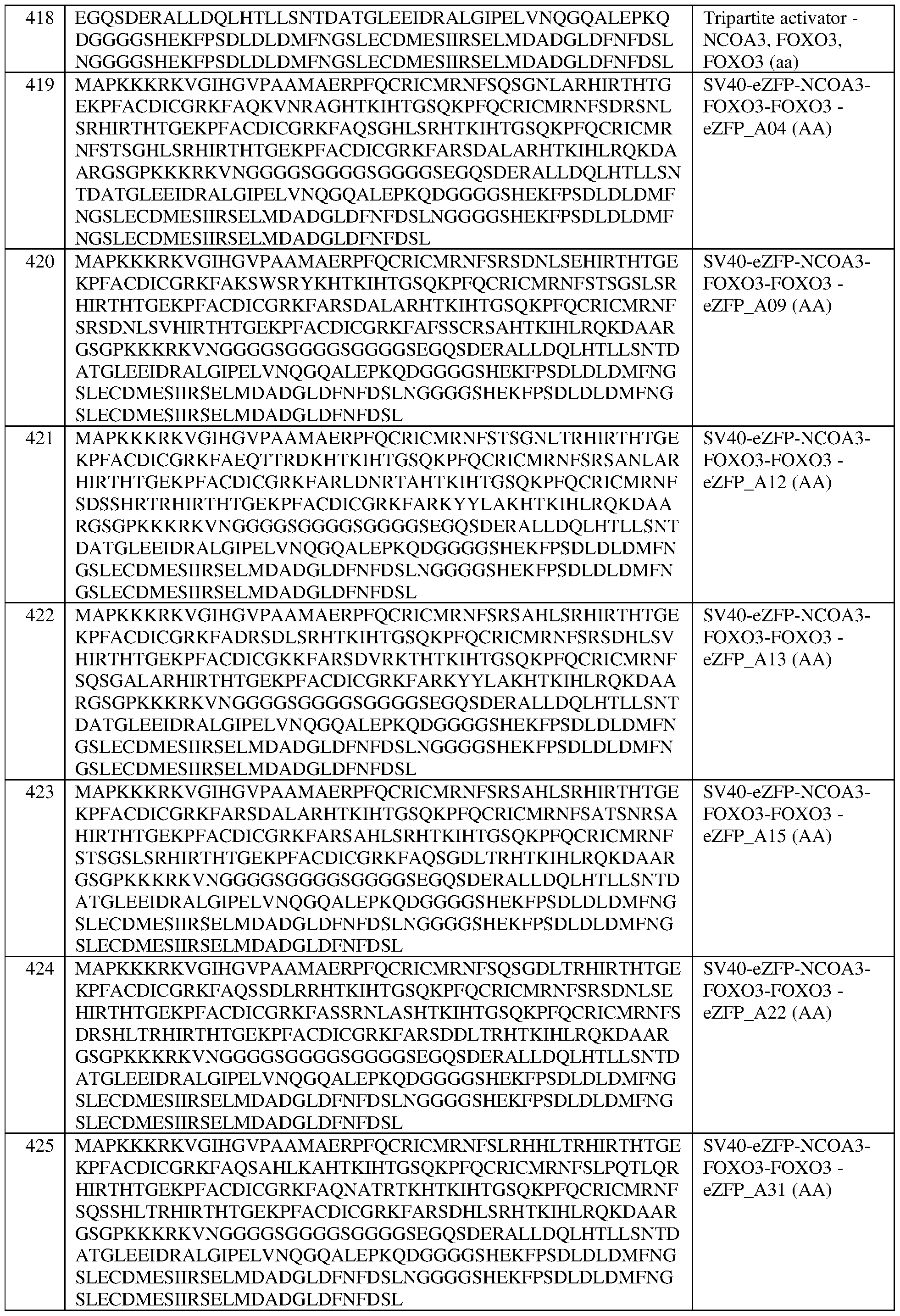
- the multipartite effector is set forth in any one of SEQ ID NOS:411- 418, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
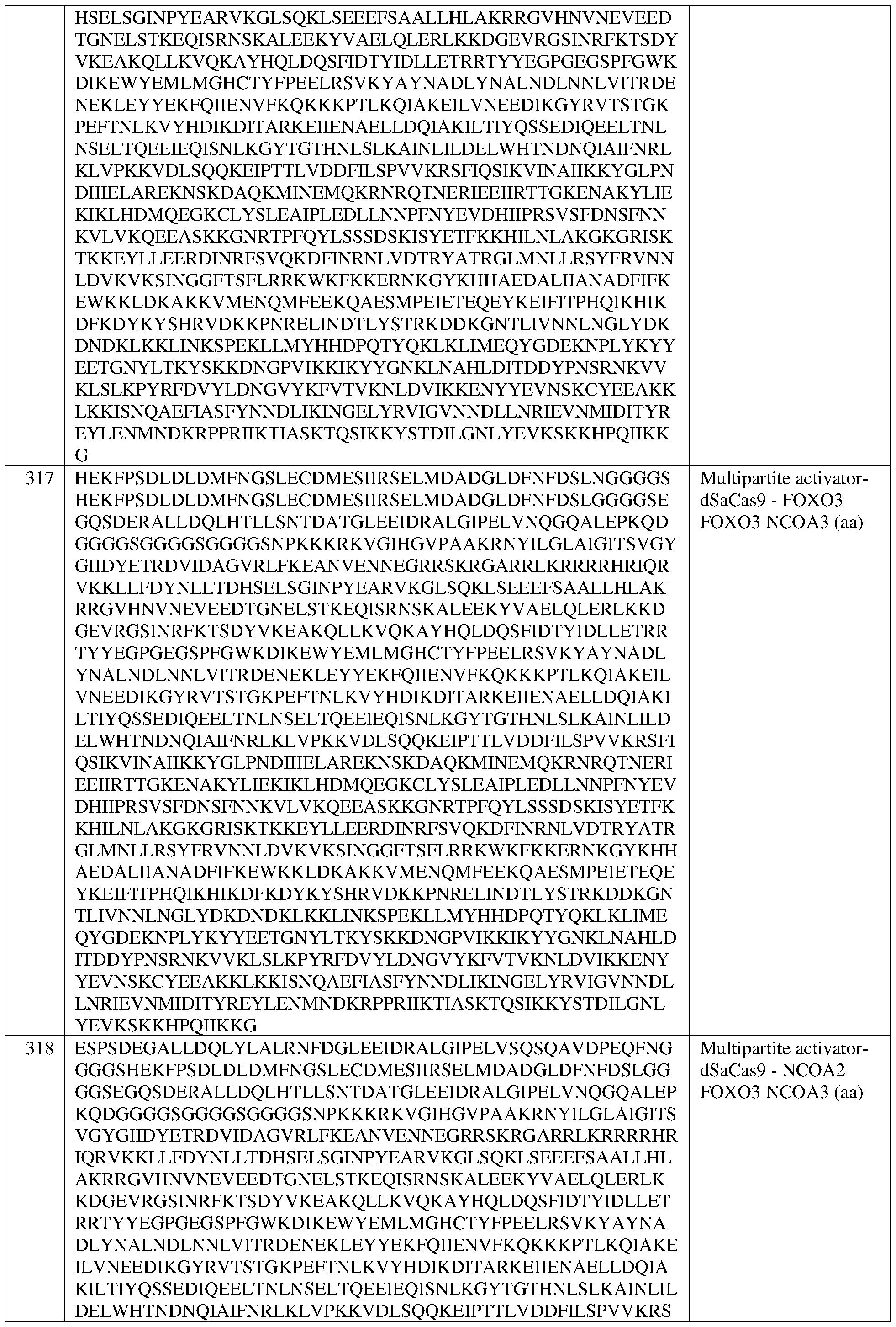
- the multipartite effector comprises, in the N-terminal to C-terminal direction, domains from FOXO3, FOXO3, and NCOA3.
- the multipartite effector comprises the sequence set forth in SEQ ID NO:415, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the multipartite effector comprises, in the N-terminal to C-terminal direction, domains from NCOA3; FOXO3, and FOXO3.
- the multipartite effector comprises the sequence set forth in SEQ ID NO:418, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the multipartite effector comprises, in the N-terminal to C- terminal direction, domains from NC0A3, FOXO3, and NC0A3.
- the multipartite effector comprises the sequence set forth in SEQ ID NO:413, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the multipartite effector comprises, in the N-terminal to C-terminal direction, domains from NCOA2, FOXO3, and NCOA3.
- the multipartite effector comprises the sequence set forth in SEQ ID NO:416, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the multipartite effector comprises, in the N-terminal to C-terminal direction, domains from PYGO1, FOXO3, and NCOA3. In some of any of the provided embodiments, the multipartite effector comprises the sequence set forth in SEQ ID NO:411, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the at least one epigenetic effector domain is fused to the N-terminus of the eZFP. In some of any of the provided embodiments, the at least one epigenetic effector domain is fused to the C-terminus of the eZFP. In some of any of the provided embodiments, the at least one epigenetic effector domain is fused to both the N-terminus and the C- terminus, of the eZFP. In some of any of the provided embodiments, the fusion protein further comprises one or more nuclear localization signals (NLS). In some of any of the provided embodiments, the fusion protein further comprises one or more linkers.
- NLS nuclear localization signals
- the one or more linkers are in between any two of the components of the fusion protein, including the eZFP, any of the at least one effector domains, and the one or more NFS. In some of any of the provided embodiments, the one or more linkers connect the eZFP and the at least one epigenetic effector domain. In some of any of the provided embodiments, the fusion protein further comprises one or more NFS, the eZFP, and the at least one epigenetic effector domain, in order from N-terminus to C-terminus.
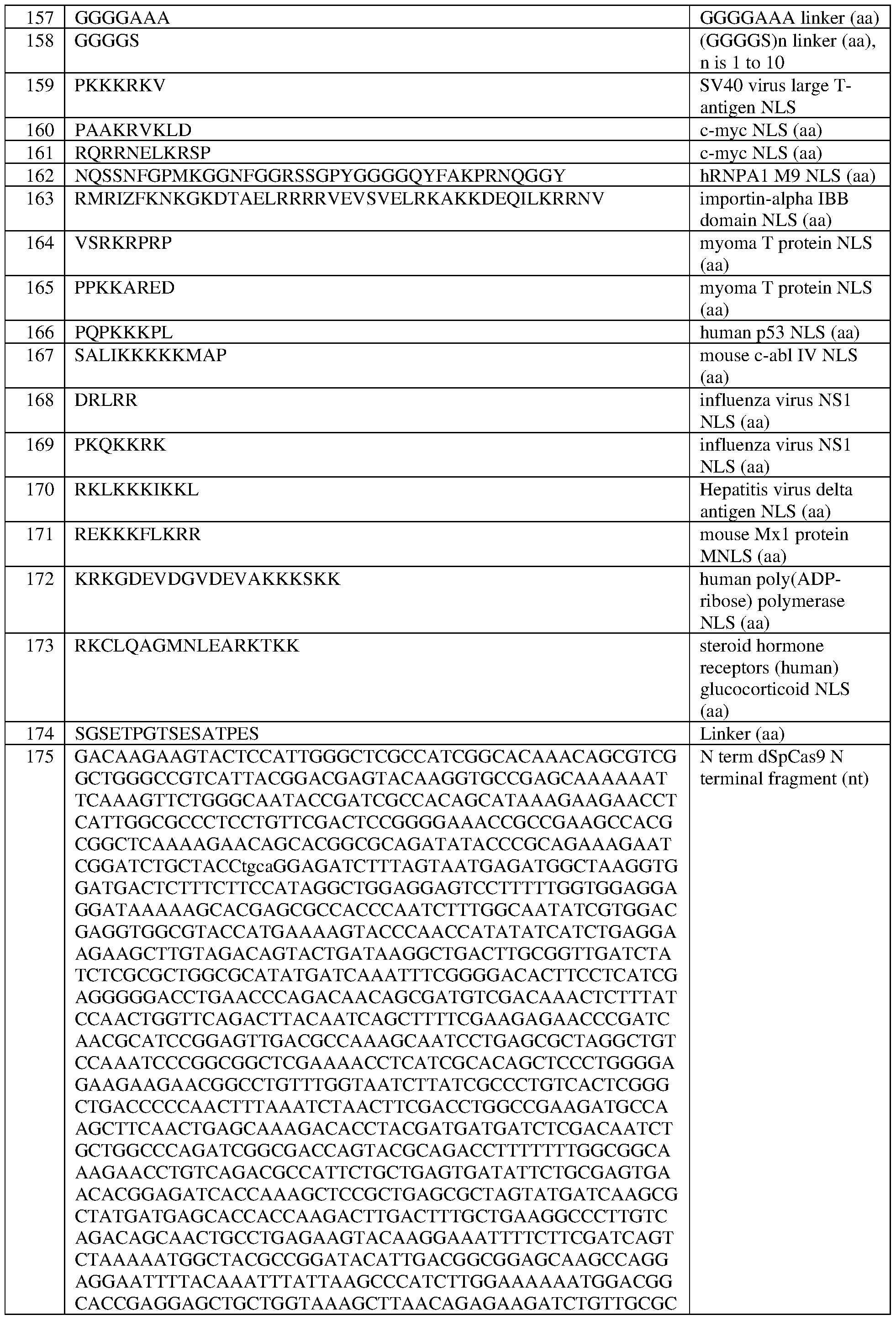
- the one or more NFS comprises a SV40 NFS sequence set forth in SEQ ID NO: 159 or a c-myc NFS sequence set forth in SEQ ID NO: 160.
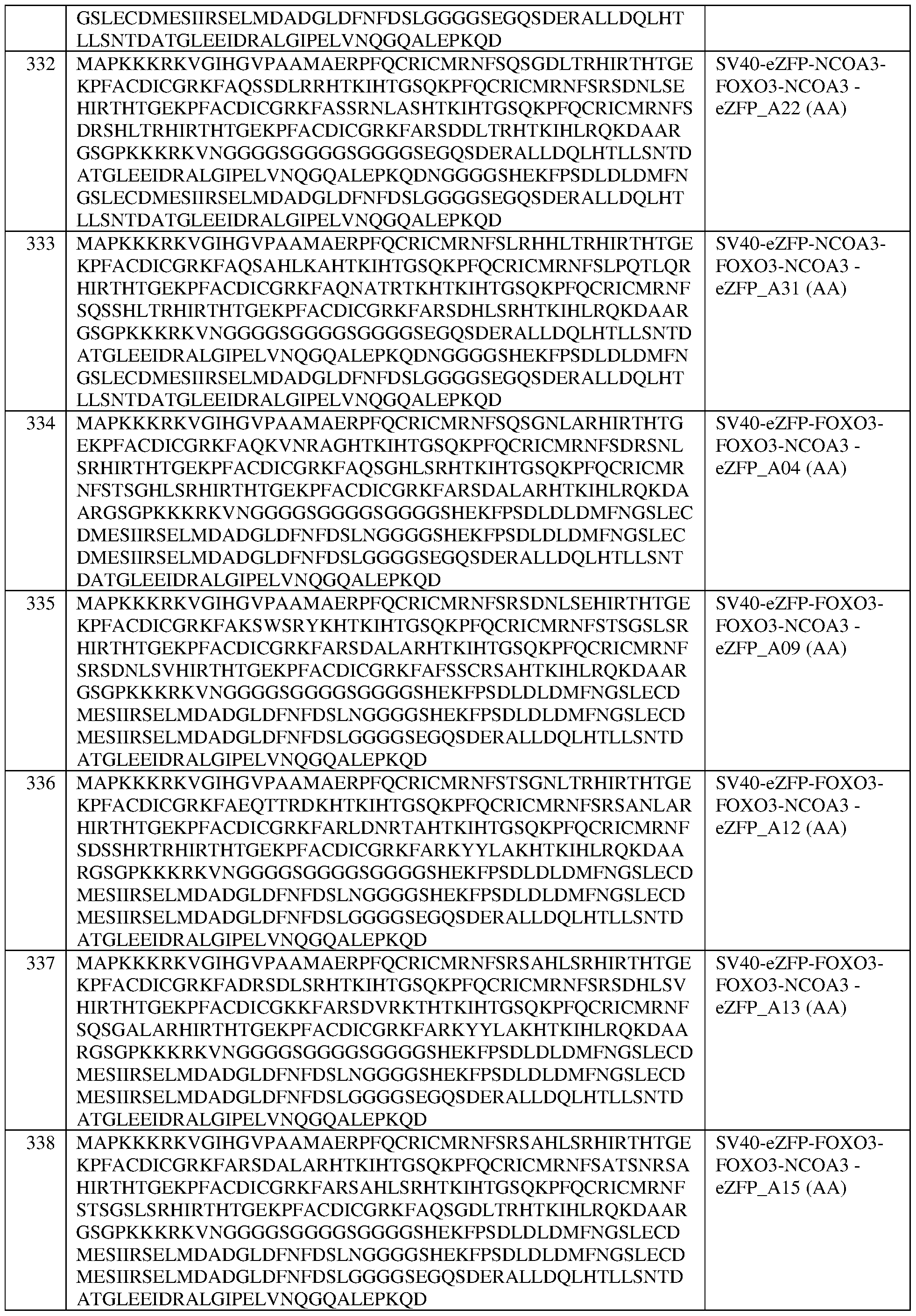
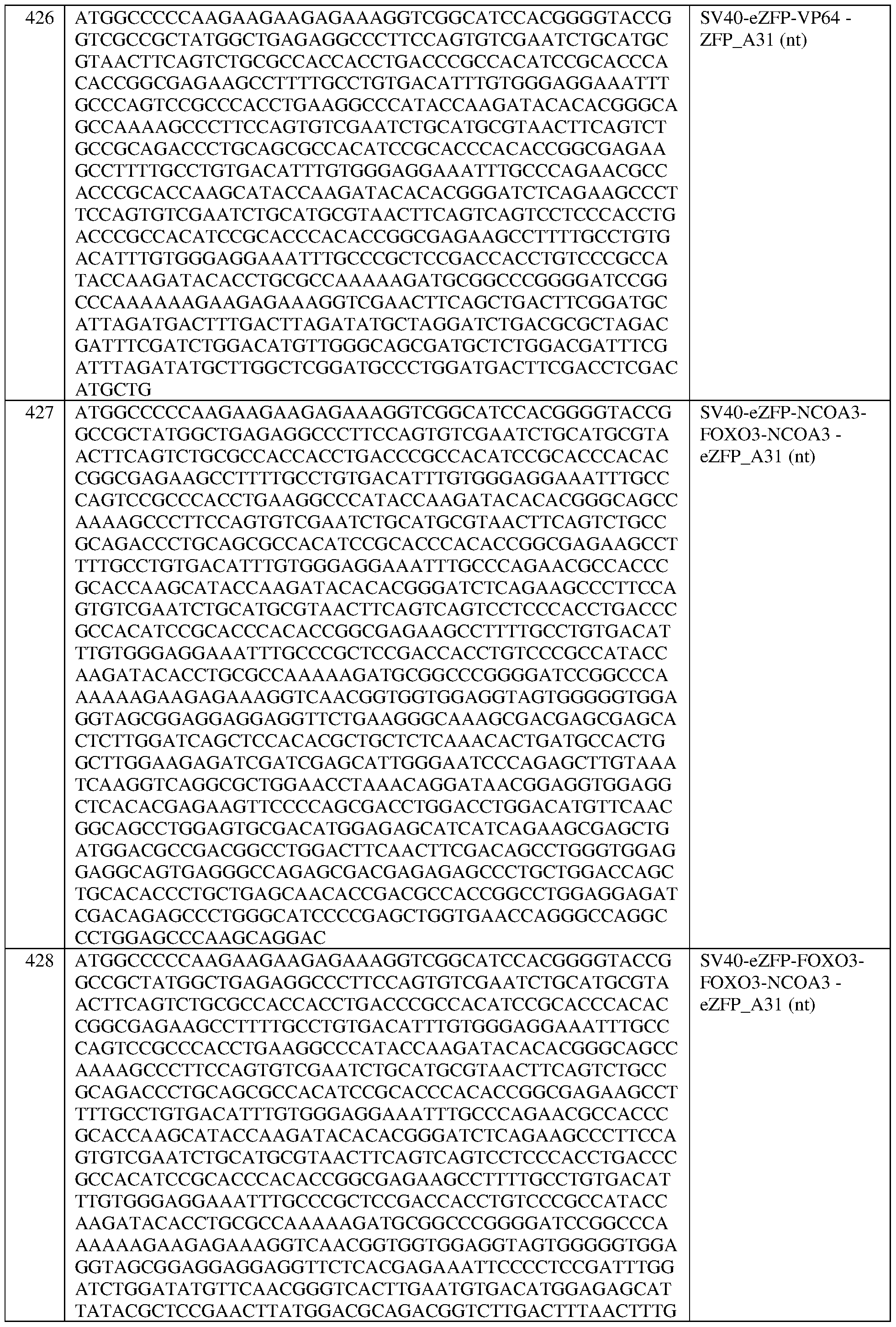
- the fusion protein comprises the sequence set forth in any of SEQ ID NOS:320-340 and 419-425, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
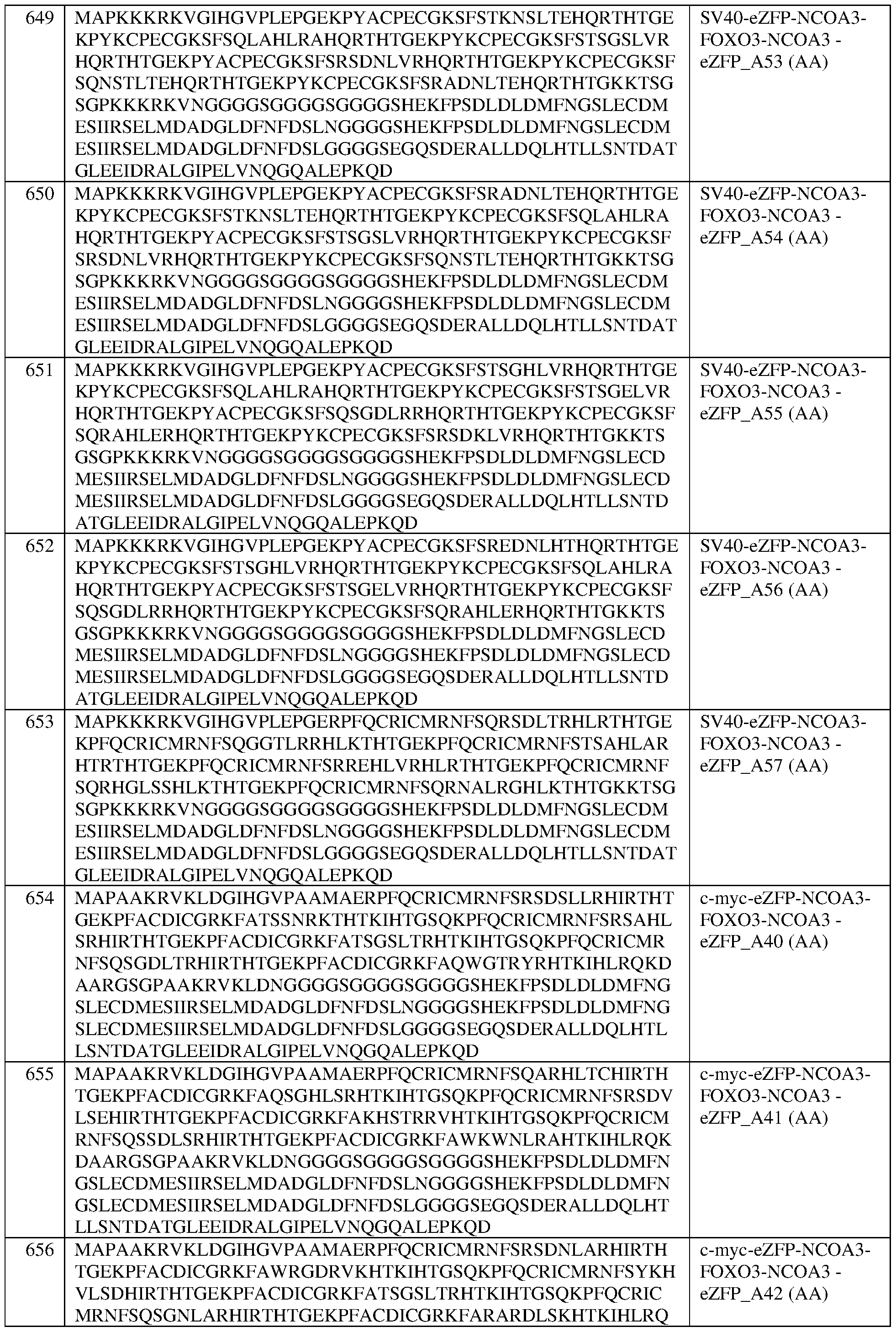
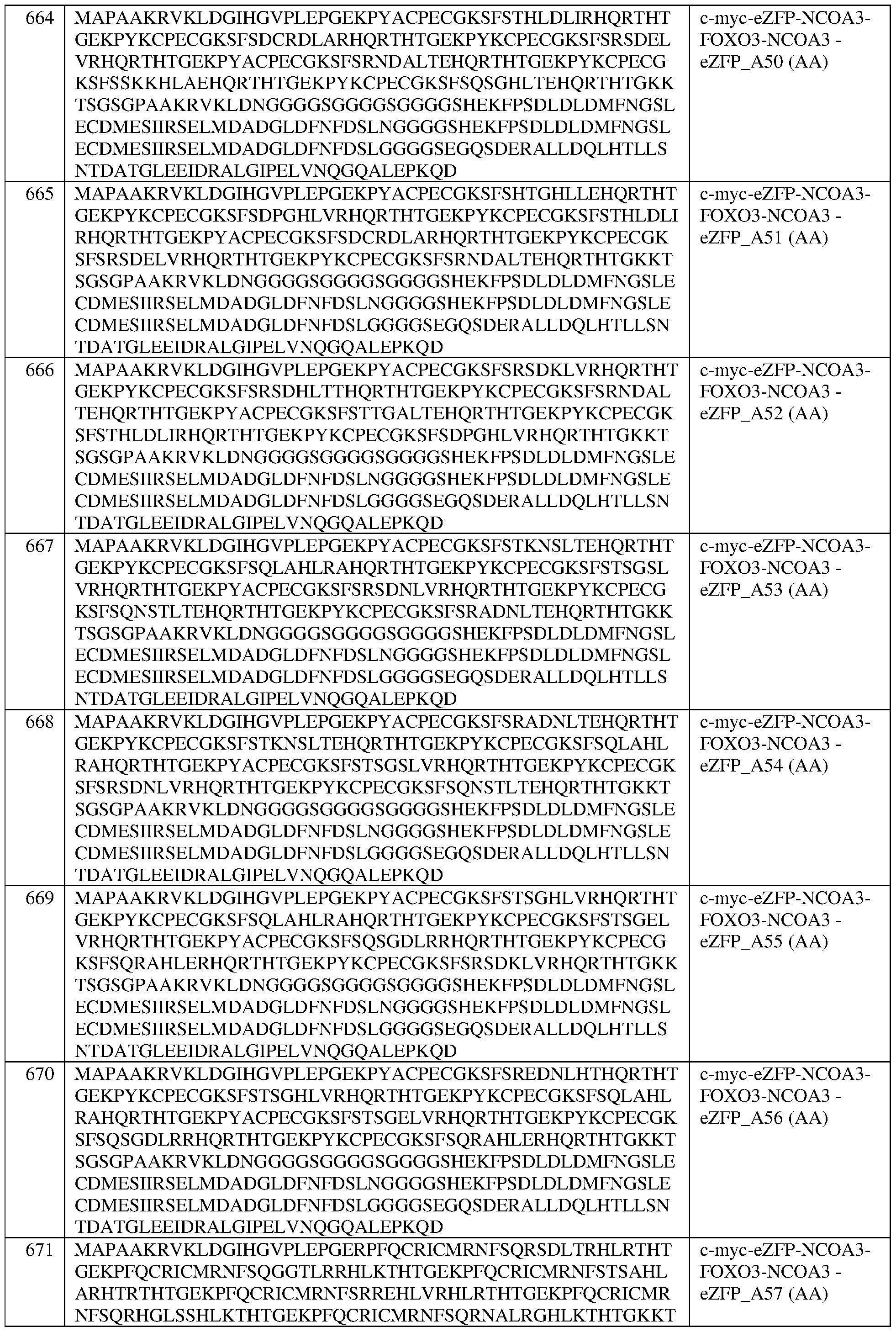
- the fusion protein comprises the sequence set forth in any of SEQ ID NOS:320-340, 419- 425, and 608-671.
- the fusion protein of any of claims 189-232 wherein the fusion protein comprises the sequence set forth in any of SEQ ID NOS:320-340, 419-425, and 608-671. In some of any of the provided embodiments, the fusion protein comprises the sequence set forth in any of SEQ ID NOS:320- 340, 419-425, and 636-653. In some of any of the provided embodiments, the fusion protein comprises the sequence set forth in any of SEQ ID NOS:320-340 and 419-425. In some of any of the provided embodiments, the fusion protein comprises the sequence set forth in any of SEQ ID NOS:636-653.
- the fusion protein comprises the sequence set forth in any of SEQ ID NOS:608-635 and 654-671. In some of any of the provided embodiments, the fusion protein comprises the sequence set forth in any of SEQ ID NOS:608-635. In some of any of the provided embodiments, the fusion protein comprises the sequence set forth in any of SEQ ID NOS:654-671. In some of any of the provided embodiments, the fusion protein comprises the sequence set forth in SEQ ID NO:326, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the fusion protein comprises the sequence set forth in SEQ ID NO: 333, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the fusion protein comprises the sequence set forth in SEQ ID NO:340, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the fusion protein comprises the sequence set forth in SEQ ID NO:425, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the fusion protein comprises the sequence set forth in SEQ ID NO:662, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the fusion protein comprises the sequence set forth in SEQ ID NO:660, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the fusion protein comprises the sequence set forth in SEQ ID NO:658, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- provided herein is a polynucleotide encoding any eZFP, or any fusion protein provided herein, or a portion or component of any of the foregoing. In some aspects, provided herein is a plurality of polynucleotides encoding any eZFP, or any fusion protein provided herein, or a portion or component of any of the foregoing.
- a vector comprising any eZFP, any fusion protein, any polynucleotide, or any plurality of polynucleotides provided herein, or a portion or component of any of the foregoing.
- the vector is a viral vector.
- the vector is an adeno-associated virus (AAV) vector.
- AAV vector is selected from the group consisting of: AAV1, AAV2, AAV3, AAV4, AAV5, AAV6, AAV7, AAV8, AAV9, AAV10, AAV11, AAV12, AAV-DJ, and AAVrh74.
- the AAV vector is AAV6. In some of any of the provided embodiments, the AAV vector is AAV9. In some of any of the provided embodiments, the AAV vector is AAV-DJ. In some of any of the provided embodiments, the AAV vector is AAVrh74. In some of any of the provided embodiments, the vector is a lenti viral vector. In some of any of the provided embodiments, the vector is a non-viral vector. In some of any of the provided embodiments, the non-viral vector is selected from the group consisting of: a lipid nanoparticle, a liposome, an exosome, and a cell penetrating peptide.
- the vector exhibits tropism for a nervous system cell, optionally a neuron, a heart cell, optionally a cardiomyocyte, a skeletal muscle cell, a fibroblast, an induced pluripotent stem cell, and/or a cell derived from any of the foregoing, or for a combination of any of the foregoing cells.
- the vector exhibits tropism for induced pluripotent stem cells.
- the vector exhibits tropism for neurons and cardiomyocytes.
- the vector comprises one vector, or two or more vectors.
- an AAV vector comprising one or both of a) a first nucleic acid comprising an elongation factor alpha short (EFS) promoter operably linked to a sequence encoding a fusion protein comprising (i) a deactivated Cas (dCas) protein and (ii) at least one effector domain that increases transcription of a frataxin (FXN) locus; and b) a second nucleic acid comprising a U6 promoter operably linked to a sequence encoding a guide RNA (gRNA) comprising a gRNA spacer sequence that is capable of hybridizing to a target site in a regulatory DNA element of a FXN locus and/or is complementary to the target site.
- EFS elongation factor alpha short
- dCas deactivated Cas
- FXN frataxin
- the AAV vector comprises both the first nucleic acid and the second nucleic acid.
- the first and second nucleic acid are comprised in a single polynucleotide.
- the EFS promoter comprises the sequence set forth in SEQ ID NO:436, or a sequence having at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to thereto.
- the EFS promoter comprises the sequence set forth in SEQ ID NO:436.
- the U6 promoter is a mini-U6 promoter.
- the mini-U6 promoter comprises the sequence set forth in SEQ ID NO:433, or a sequence having at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to thereto. In some of any of the provided embodiments, the mini-U6 promoter comprises the sequence set forth in SEQ ID NO:433.
- an AAV vector comprising a nucleic acid comprising a promoter selected from an elongation factor alpha short (EFS), CAG, or human elongation factor- 1 alpha (EFla) promoter operably linked to a sequence encoding a fusion protein comprising (i) an eZFP that is capable of hybridizing to a target site in a regulatory DNA element of a frataxin (FXN) locus and/or is complementary to the target site and (ii) at least one effector domain that increases transcription of the frataxin (FXN) locus.
- EFS elongation factor alpha short
- EFla human elongation factor- 1 alpha
- the EFS promoter comprises the sequence set forth in SEQ ID NO:436, or a sequence having at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to thereto.
- the EFS promoter comprises the sequence set forth in SEQ ID NO:436.
- the CAG promoter comprises the sequence set forth in SEQ ID NO:602, or a sequence having at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to thereto.
- the CAG promoter comprises the sequence set forth in SEQ ID NO:602.
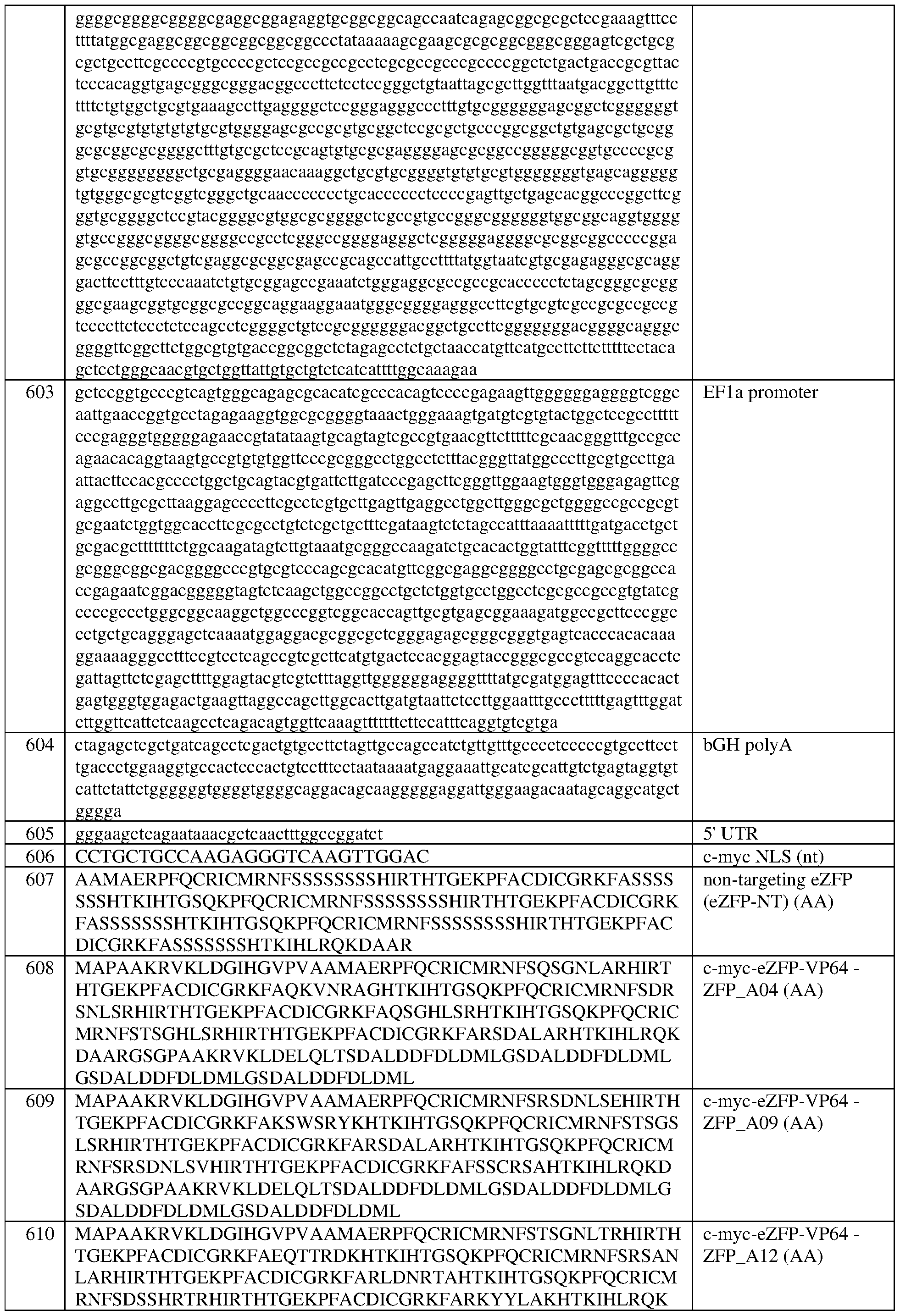
- the EFla promoter comprises the sequence set forth in SEQ ID NO:603, or a sequence having at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to thereto.
- the EFla promoter comprises the sequence set forth in SEQ ID NO:603.
- the nucleic acid further comprises a 5’ untranslated region (UTR) set forth in SEQ ID NO:605.
- the AAV vector further comprises inverted terminal repeats (ITRs).
- ITRs are a first and second ITR, comprising the sequences set forth in SEQ ID NO:434 and SEQ ID NO:435, respectively.
- the single polynucleotide comprises, in the 5’ to 3’ direction, the EFS promoter, the sequence encoding the fusion protein, the U6 promoter, and the sequence encoding the gRNA.
- the single polynucleotide further comprises a first IRT 5’ of the EFS promoter and a second ITR 3’ of the sequence encoding the gRNA.
- the first nucleic acid or the nucleic acid further comprises a polyA sequence selected from a SpA site or a bGH site downstream of the sequence encoding the fusion protein.
- the first nucleic acid or the nucleic acid comprising a sequence encoding a fusion protein further comprises a polyA sequence selected from a SpA site downstream of the sequence encoding the fusion protein.
- the first nucleic acid comprising a sequence encoding a fusion protein further comprises a polyA sequence selected from a bGH site downstream of the sequence encoding the fusion protein.
- the nucleic acid comprising a sequence encoding a fusion protein further comprises a polyA sequence selected from a SpA site or bGH site downstream of the sequence encoding the fusion protein.
- the SpA site comprises the sequence set forth in SEQ ID NO:437.
- the bGH site comprises the sequence set forth in SEQ ID NO: 604.
- the first nucleic acid further comprises a woodchuck hepatitis virus post-transcriptional regulatory element (WPRE) in proximal to the SpA site, optionally wherein the WPRE is located between the sequence encoding the fusion protein and the SpA site.
- WPRE woodchuck hepatitis virus post-transcriptional regulatory element
- the gRNA is capable of complexing with the dCas protein.
- the gRNA comprises a gRNA spacer sequence that is capable of hybridizing to the target site or is complementary to the target site.
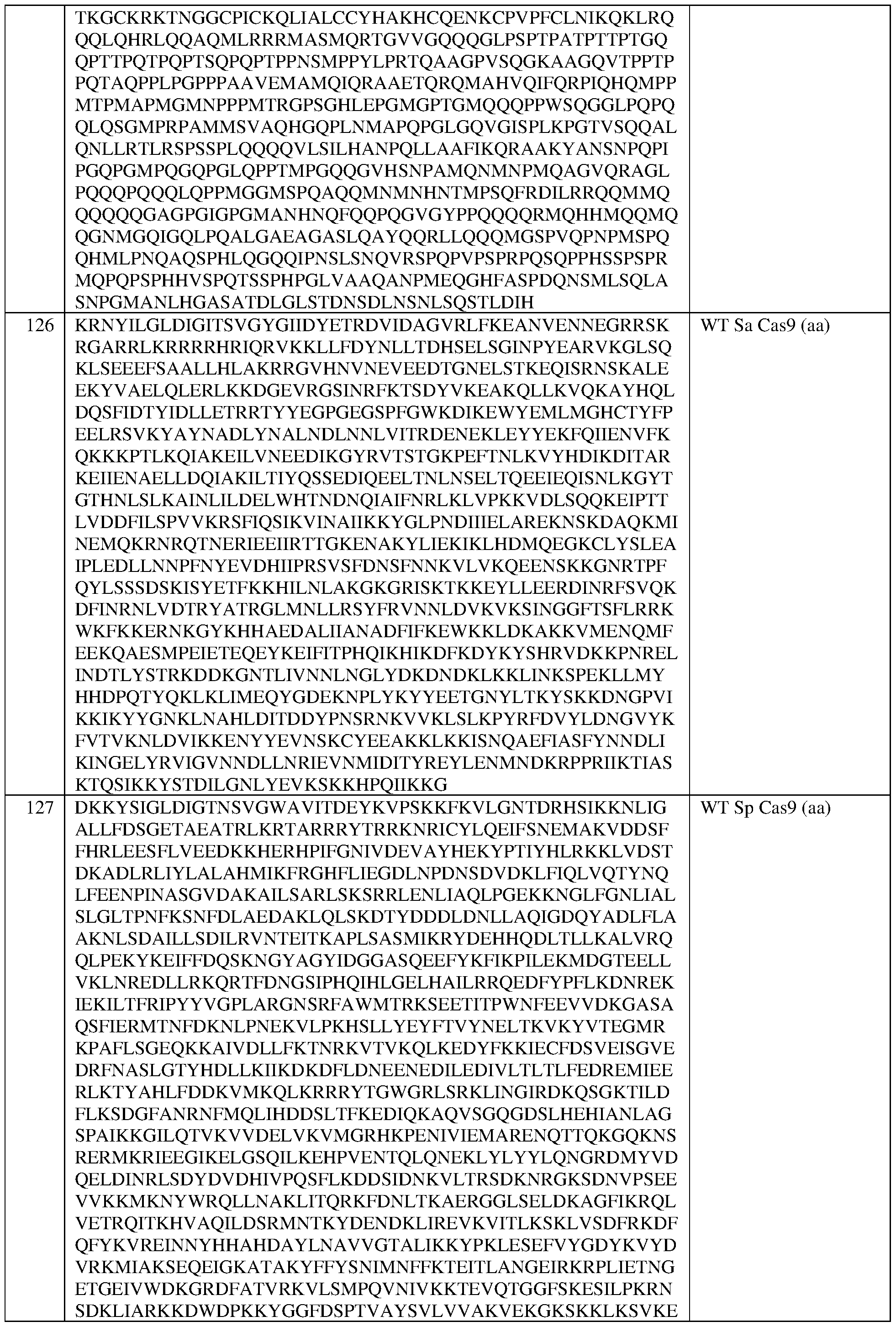
- the dCas protein is a Staphylococcus aureus dCas9 (dSaCas9) protein or a Streptococcus pyogenes dCas9 (dSpCas9) protein.
- the dCas protein is a Staphylococcus aureus dCas9 protein (dSaCas9) that comprises at least one amino acid mutation selected from D10A and N580A, with reference to numbering of positions of SEQ ID NO:73, and/or the dCas protein comprises the sequence set forth in SEQ ID NO:72, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- dSaCas9 protein Staphylococcus aureus dCas9 protein
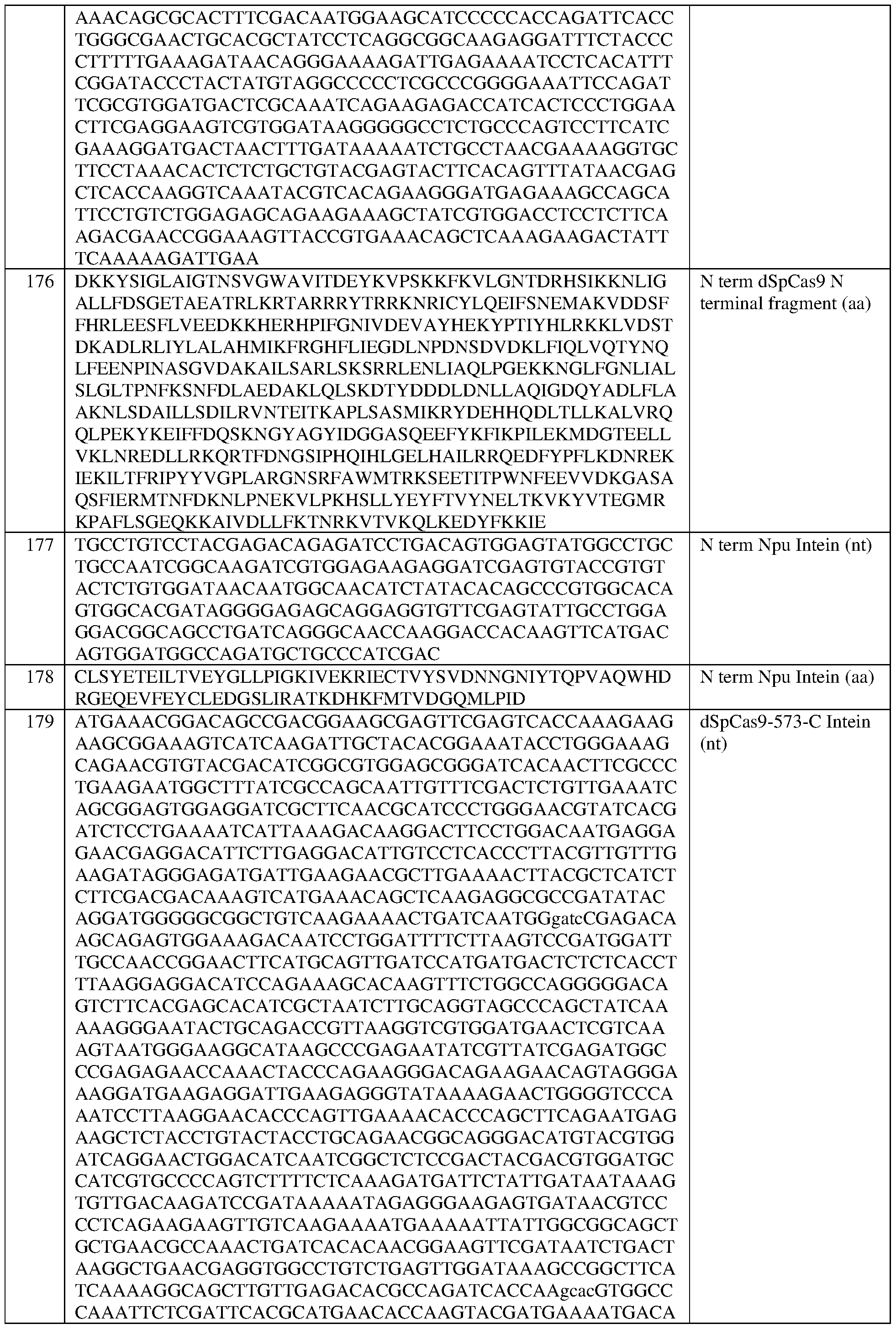
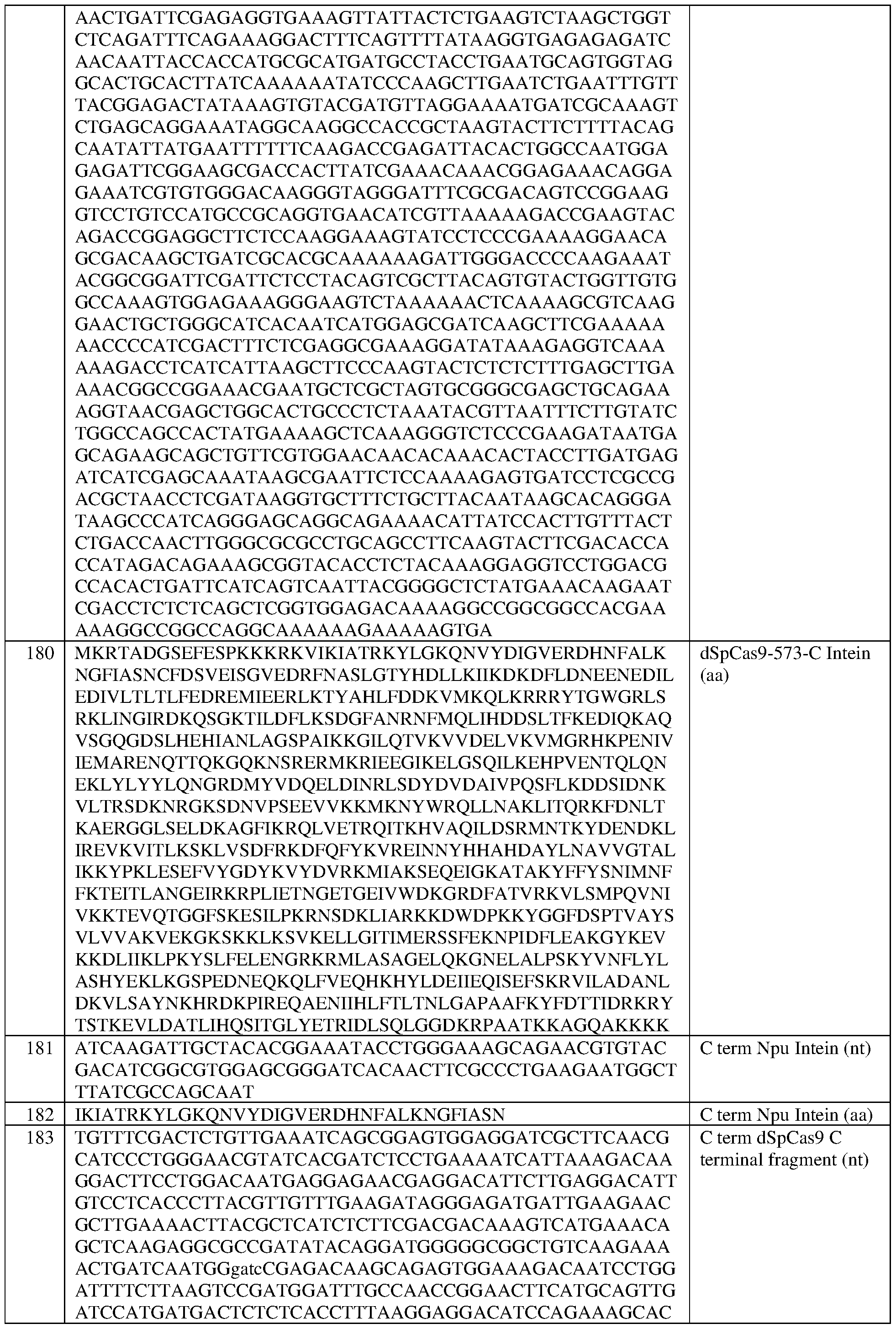
- the dCas is a Streptococcus pyogenes dCas9 (dSpCas9) protein that comprises at least one amino acid mutation selected from D10A and H840A, with reference to numbering of positions of SEQ ID NO:79, and/or the dCas protein comprises the sequence set forth in SEQ ID NO:78, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- dSpCas9 Streptococcus pyogenes dCas9
- the regulatory DNA element is an enhancer.
- the target site is located within a target region spanning the genomic coordinates chr9:69, 027, 282-69, 028, 497 from hg38 (SEQ ID NO:431), optionally wherein the target site is located within a target region spanning the genomic coordinates chr9:69, 027, 615-69, 028, 101 from hg38, optionally wherein the target site is located within a target region spanning the genomic coordinates chr9:69, 027, 825-69, 027, 875.
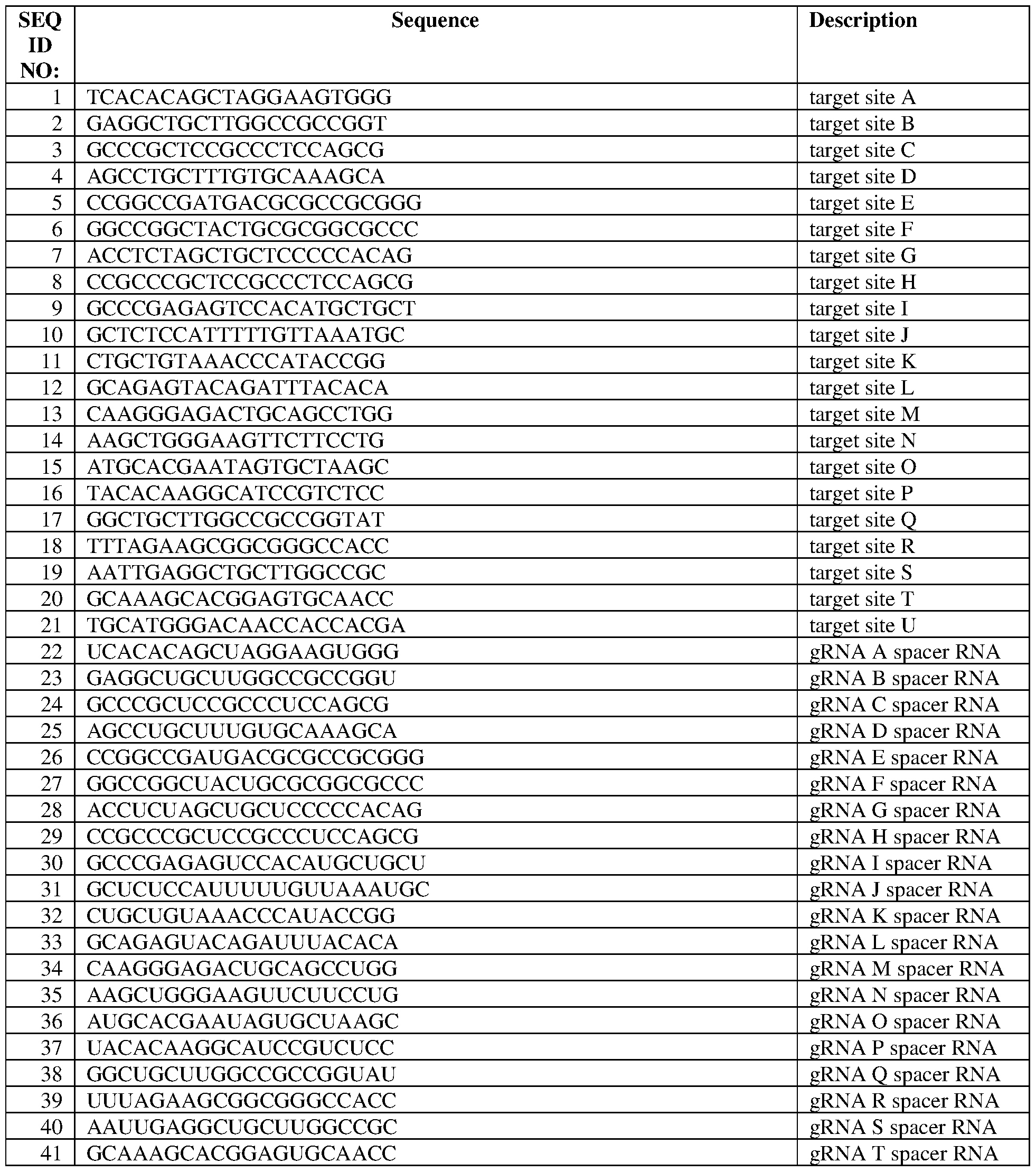
- the target site comprises the sequence set forth in SEQ ID NO:21, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing.
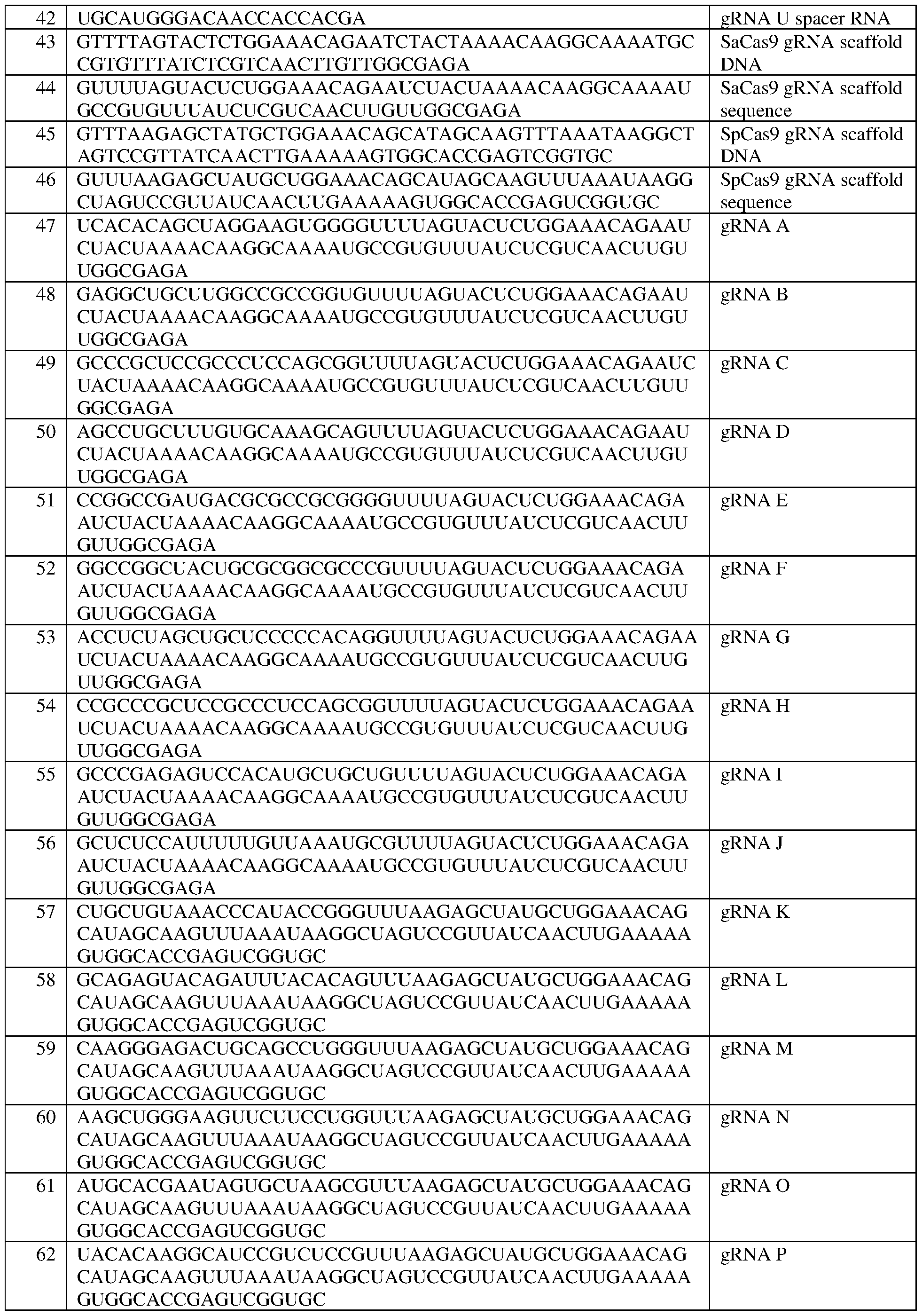
- the gRNA comprises a gRNA spacer sequence comprising the sequence set forth in SEQ ID NO:42, or a contiguous portion thereof of at least 14 nt.
- the gRNA further comprises the sequence set forth in SEQ ID NO:44, optinally wherein the gRNA comprises the sequence set forth in SEQ ID NO:67, optionally wherein the gRNA is the gRNA sequence set forth in SEQ ID NO:67.
- the target site comprises the sequence set forth in any one of SEQ ID NOS: 272 and 277, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing.
- the eZFP comprises the sequence set forth in any one of SEQ ID NOS: 301 and 302.
- the regulatory DNA element is a promoter.
- the target site is within a target region spanning the genomic coordinates chr9:69, 034, 900-69, 035, 900 from hg38 (SEQ ID NO:430), optionally wherein the target site is within a target region spanning the genomic coordinates chr9:69, 035, 300-69-035, 800 from hg38; chr9:69, 035, 350-69, 035, 450 from hg38; or chr9:69, 035, 675-69, 035, 725.
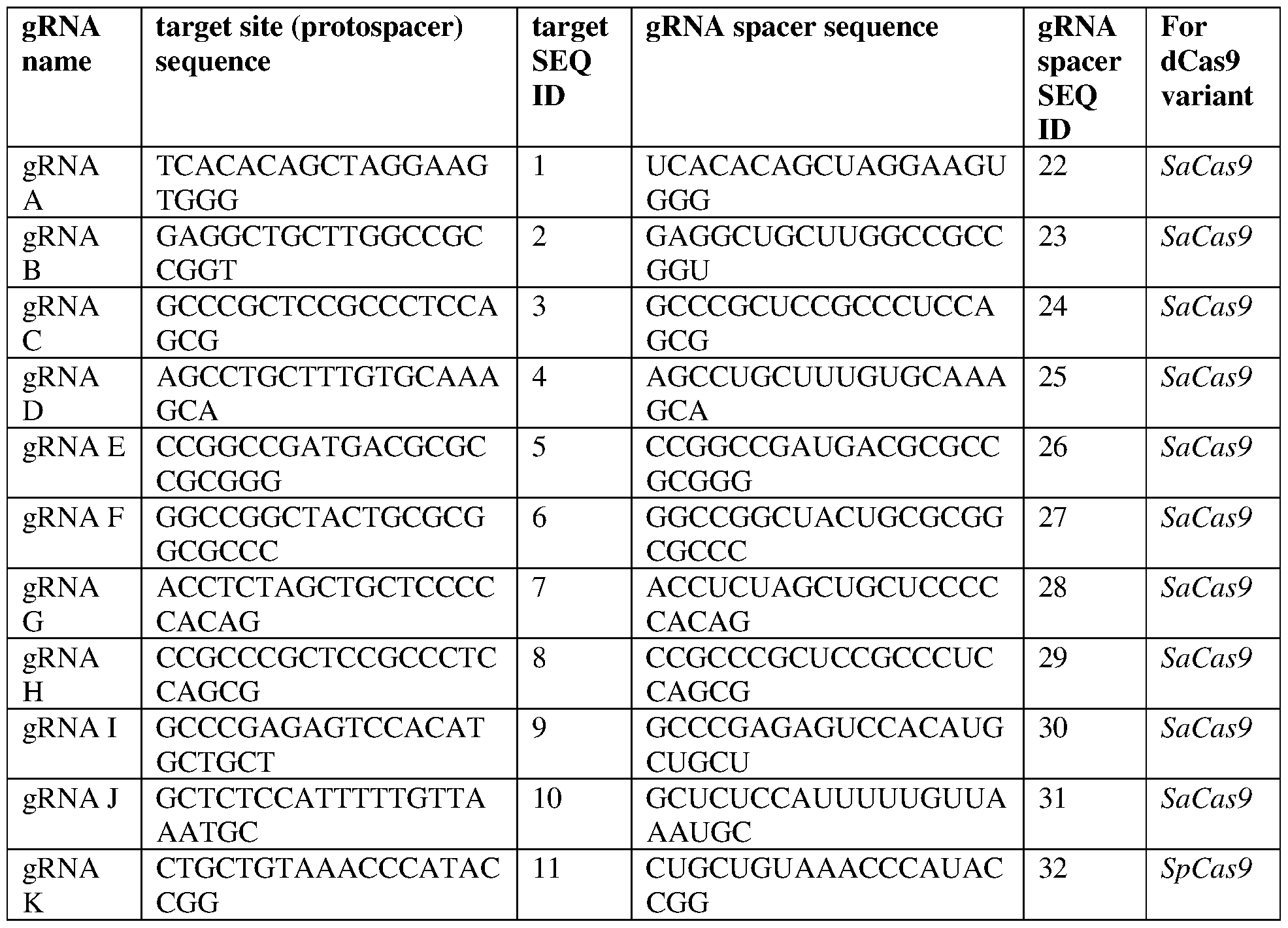
- the target site comprises a sequence selected from any of SEQ ID NOS: 1-10, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing.
- the gRNA comprises a gRNA spacer sequence comprising a sequence selected from any of SEQ ID NOS:22-31, or a contiguous portion thereof of at least 14 nt.
- the gRNA comprises a gRNA spacer sequence comprising SEQ ID NO:22, or a contiguous portion thereof of at least 14 nt.
- the gRNA comprises a gRNA spacer sequence comprising SEQ ID NO:28, or a contiguous portion thereof of at least 14 nt. In some of any of the provided embodiments, the gRNA further comprises the sequence set forth in SEQ ID NO:44, optionally wherein the gRNA comprises a sequence selected from any of SEQ ID NOS:47-56, optionally wherein the gRNA is the gRNA sequence set forth in any of SEQ ID NOS:47- 56, optionally wherein the gRNA is set forth in SEQ ID NO:47 or 53.
- the target site comprises a sequence selected from any of SEQ ID NOS: 11-20, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing.
- the gRNA comprises a gRNA spacer sequence comprising a sequence selected from any of SEQ ID NOS:32-41, or a contiguous portion thereof of at least 14 nt.
- the gRNA further comprises the sequence set forth in SEQ ID NO:46, and/or wherein the gRNA comprises a sequence selected from any of SEQ ID NOS:57-66, optionally wherein the gRNA is the gRNA set forth in any of SEQ ID NOS:57-66.
- the gRNA spacer sequence is between 14 nt and 24 nt, or between 16 nt and 22 nt in length, optionally wherein the gRNA spacer sequence is 18 nt, 19 nt, 20 nt, 21 nt or 22 nt in length.
- the target site comprises the sequence set forth in any one of SEQ ID NOS: 280-283, 290, 299, and 583-600, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing. In some of any of the provided embodiments, the target site comprises the sequence set forth in any one of SEQ ID NOS: 299, 587, 589, and 591, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing.
- the target site comprises the sequence set forth in SEQ ID NO: 299, a contiguous portion thereof of at least 14 nt, or a complementary sequence of the sequence set forth in SEQ ID NO: 299. In some of any of the provided embodiments, the target site comprises the sequence set forth in SEQ ID NO: 587, a contiguous portion thereof of at least 14 nt, or a complementary sequence of the sequence set forth in SEQ ID NO: 587. In some of any of the provided embodiments, the target site comprises the sequence set forth in SEQ ID NO: 589, a contiguous portion thereof of at least 14 nt, or a complementary sequence of the sequence set forth in SEQ ID NO: 589.
- the target site comprises the sequence set forth in SEQ ID NO: 591, a contiguous portion thereof of at least 14 nt, or a complementary sequence of the sequence set forth in SEQ ID NO: 591.
- the eZFP comprises the sequence set forth in any one of SEQ ID NOS: 303-307 and 439-456.
- the eZFP comprises the sequence set forth in any one of SEQ ID NOS: 307, 441, 443, and 445.
- the eZFP comprises the sequence set forth in SEQ ID NO: 307.
- the eZFP comprises the sequence set forth in SEQ ID NO: 441. In some of any of the provided embodiments, the eZFP comprises the sequence set forth in SEQ ID NO: 443. In some of any of the provided embodiments, the eZFP comprises the sequence set forth in SEQ ID NO: 445.
- the at least one effector domain induces transcription activation.
- the at least one epigenetic effector domain comprises: a VP64 domain, a p65 activation domain, a p300 domain, an Rta domain, a CBP domain, a VPR domain, a VPH domain, an HSF1 domain, a TET protein domain, optionally wherein the TET protein is TET1, a SunTag domain, a domain from DPOLA, ENL, FOXO3, HSH2D, NCOA2, NCOA3, PSA1, PYGO1, RBM39, HERC2, or NOTCH2, or a domain, portion, variant, or truncation of any of the foregoing.
- the at least one epigenetic effector domain comprises the sequence set forth in any of SEQ ID NOS:81, 83, 100-109, 111-122, 124, 125, 134-140, 152, and 383-396, or a domain, portion, variant, or truncation thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
- the at least one effector domain is fused to the N-terminus, the C-terminus, or both the N-terminus and the C-terminus, of the dCas protein or eZFP.
- the AAV vector further comprises one or more linkers connecting the dCas protein or eZFP to the at least one effector domain, and/or further comprising one or more nuclear localization signals (NFS).
- the at least one effector domain comprises at least one VP16 domain, or a VP16 tetramer (“VP64”) or a variant thereof. In some of any of the provided embodiments, the at least one effector domain comprises VP64.
- the at least one effector domain comprises the sequence set forth in SEQ ID NO:81 or 83, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing. In some of any of the provided embodiments, the at least one effector domain comprises the sequence set forth in SEQ ID NO: 81 or 83.
- the at least one epigenetic effector domain comprises: a domain from DPOLA, ENL, FOXO3, HSH2D, NCOA2, NCOA3, PSA1, PYGO1, RBM39, HERC2, or NOTCH2, or a domain, portion, variant, or truncation of any of the foregoing.
- the at least one epigenetic effector domain comprises the sequence set forth in any of SEQ ID NOS:383-393, or a domain, portion, variant, or truncation thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
- the at least one effector domain comprises a domain from NCOA2, NCOA3, FOXO3, PYGO1, or a portion or variant of any of the foregoing.
- each effector domain of the at least one effector domain is independently selected from an NCOA2 domain, an NCOA3 domain, a FOXO3 domain, and a PYGO1 domain.
- the at least one effector domain comprises a domain from NCOA2 comprising the sequence set forth in SEQ ID NO: 104 or SEQ ID NO:387, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the at least one effector domain comprises a domain from NCOA2 set forth in or SEQ ID NO:387.
- the at least one effector domain comprises a domain from NCOA3 comprising the sequence set forth in SEQ ID NO: 105 or SEQ ID NO:388, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the at least one effector domain comprises a domain from NCOA3 set forth in or SEQ ID NO:388.
- the at least one effector domain comprises a domain from FOXO3 comprising the sequence set forth in SEQ ID NO: 102 or SEQ ID NO:385, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the at least one effector domain comprises a domain from F0X03 set forth in or SEQ ID NO:385.
- the at least one effector domain comprises a domain from PYG01 comprising the sequence set forth in SEQ ID NO: 107 or SEQ ID NO:390, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the at least one effector domain comprises a domain from PYGO1 set forth in or SEQ ID NO:390.
- the at least one effector domain is a multipartite effector composed of at least two effector domains.
- the multipartite effector is composed of two effector domains.
- the multipartite effector is composed of three effector domains.
- the multipartite effector is set forth in any of SEQ ID NOS:397-418, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
- the multipartite effector is set forth in any of SEQ ID NOS:411-418, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
- the multipartite effector comprises, in the N-terminal to C-terminal direction, domains from FOXO3, FOXO3, and NCOA3.
- the multipartite effector comprises the sequence set forth in SEQ ID NO:415, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the multipartite effector comprises, in the N-terminal to C-terminal direction, domains from NCOA3; FOXO3, and FOXO3.
- the multipartite effector comprises the sequence set forth in SEQ ID NO:418, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the multipartite effector comprises, in the N-terminal to C-terminal direction, domains from NCOA3, FOXO3, and NCOA3.
- the multipartite effector comprises the sequence set forth in SEQ ID NO:413, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the multipartite effector comprises, in the N-terminal to C-terminal direction, domains from NCOA2, FOXO3, and NCOA3.
- the multipartite effector comprises the sequence set forth in SEQ ID NO:416, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the multipartite effector comprises, in the N-terminal to C-terminal direction, domains from PYGO1, FOXO3, and NCOA3.
- the multipartite effector comprises the sequence set forth in SEQ ID NO:411, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the at least one epigenetic effector domain is fused to the N-terminus of the dCas protein or eZFP. In some of any of the provided embodiments, the at least one epigenetic effector domain is fused to the C-terminus of the dCas protein or eZFP.
- the at least one epigenetic effector domain is fused to both the N- terminus and the C-terminus, of the dCas protein or eZFP.
- the one or more linkers are in between any two of the components of the fusion protein, including the eZFP, any of the at least one effector domains, and the one or more NLS.
- the one or more linkers connect the dCas protein and the at least one epigenetic effector domain.
- the fusion protein comprises the sequence set forth in SEQ ID NO:71 or 77, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto, or the sequence set forth in SEQ ID NO:71 or 77, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the fusion protein comprises the sequence set forth in any one of SEQ ID NOS:266-268 and 315-319, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the AAV vector is selected from the group consisting of: AAV1, AAV2, AAV3, AAV4, AAV5, AAV6, AAV7, AAV8, AAV9, AAV10, AAV11, AAV12, AAV-DJ, and AAVrh74.
- the AAV vector is AAV6.
- the AAV vector is AAV9.
- the AAV vector is AAV-DJ.
- the AAV vector is AAVrh74.
- the vector exhibits tropism for a nervous system cell, optionally a neuron, a heart cell, optionally a cardiomyocyte, a skeletal muscle cell, a fibroblast, an induced pluripotent stem cell, and/or a cell derived from any of the foregoing, or for a combination of any of the foregoing cells.
- the vector exhibits tropism for induced pluripotent stem cells.
- the vector exhibits tropism for neurons and cardiomyocytes.
- a cell comprising any eZFP, any fusion protein, any polynucleotide, any plurality of polynucleotides, any vector, or any AAV vector provided herein, or a portion or component of any of the foregoing, or a combination of any of the foregoing.
- the cell is a nervous system cell, optionally a neuron, a heart cell, optionally a cardiomyocyte, a skeletal muscle cell, a fibroblast, an induced pluripotent stem cell, and/or a cell derived from any of the foregoing.
- the cell is from a subject that has or is suspected of having Friedreich’s ataxia (FA).
- a pharmaceutical composition comprising any eZFP, any fusion protein, any polynucleotide, any plurality of polynucleotides, any vector, any AAV vector provided herein, or a portion or component of any of the foregoing, or a combination of any of the foregoing.
- the pharmaceutical composition is for use in treating a disease, condition, or disorder in a subject.
- the disease, condition, or disorder is Friedreich’s ataxia and/or a GAA trinucleotide repeat expansion in the FXN locus.
- the expression of FXN is increased in cells of the subject.
- a method for increasing the expression of FXN in a cell comprising introducing into the cell: any eZFP, any fusion protein, any polynucleotide, any plurality of polynucleotides, any vector, any AAV vector, or any pharmaceutical composition provided herein, or a portion or component of any of the foregoing, or a combination of any of the foregoing.
- the cell is from and/or in a subject that has or is suspected of having Friedreich’s ataxia.
- the cell exhibits reduced expression of FXN in comparison to a reference cell from an individual not having Friedreich’s ataxia and/or a GAA trinucleotide expansion in the FXN locus.
- a method for increasing the expression of FXN in a cell in a subject comprising administering to the subject: any eZFP, any fusion protein, any polynucleotide, any plurality of polynucleotides, any vector, any AAV vector, or any pharmaceutical composition provided herein, or a portion or component of any of the foregoing, or a combination of any of the foregoing.
- provided herein is a method of treating a subject in need thereof, the method comprising administering to the subject: any eZFP, any fusion protein, any polynucleotide, any plurality of polynucleotides, any vector, any AAV vector, or any pharmaceutical composition provided herein, or a portion or component of any of the foregoing, or a combination of any of the foregoing.
- the subject has or is suspected of having Friedreich’s ataxia, and/or a GAA trinucleotide expansion in the FXN locus.
- the introducing or administering is carried out in vivo or ex vivo.
- the cell and/or subject exhibits reduced expression of FXN prior to performing the method.
- the reduced expression of FXN is reduced in comparison to a reference individual not having Friedreich’s ataxia and/or a GAA trinucleotide repeat expansion in the FXN locus, and/or a reference cell therefrom.
- the GAA trinucleotide repeat expansion is in a first intron of a FXN gene, and comprises at least 50, at least 60, at least 70, at least 80, at least 90, at least 100, at least 150, at least 200, at least 300, at least 400, at least 500, at least 600, at least 700, at least 800, at least 900, at least 1000, or more repeated GAA trinucleotides.
- the expression of FXN is increased in the cell and/or subject.
- the expression of FXN is increased in the cell or cells of the subject by at least about 1.2-fold, 1.25-fold, 1.3-fold, 1.4-fold, 1.5-fold, 1.6-fold, 1.7-fold, 1.75-fold, 1.8-fold, 1.9- fold, 2-fold, 2.5-fold, 3-fold, 4-fold, or 5-fold; and/or the expression is increased by less than about 10- fold, 9-fold, 8-fold, 7-fold or 6-fold.
- the expression of FXN is increased in the cell or cells of the subject to a level that is at least at or about 30%, 40%, 50%, 60%, 70%, 75%, 80%, 85%, 90%, 95%, 100%, 105%, 110%, 120%, 125%, 150%, 175%, 200%, 225%, 250%, 300%, 400%, or 500%, of the expression level of FXN in a reference cell from an individual not having Friedreich’ s ataxia and/or a cell not having a GAA trinucleotide repeat expansion in the FXN gene.
- the expression of FXN is increased in the cell or cells of the subject to a level that is less than at or about 200%, 300%, 400%, 500%, 600%, 700%, 800%, 900%, or 1000% of the expression level of FXN in a reference cell from an individual not having Friedreich’s ataxia and/or a cell not having a GAA trinucleotide repeat expansion in the FXN gene.
- the expression is measured by the amount of mRNA encoding the FXN protein, and/or the amount of FXN protein.
- the subject is a human.
- provided herein is a cell comprising an epigenetic modification produced by any method provided herein.
- FIGS. 1A-1C show CRISPR-Cas mediated transcriptional activation of human frataxin in WT-iPSCs using dSaCas9-2xVP64 or dSpCas9-2xVP64 and gRNAs targeted to the frataxin promoter.
- FIG. 1A shows target locations of gRNAs from Table El, targeted to human frataxin gene promoter (Homo sapiens (human) genome assembly GRCh38 (hg38) chr9:69034622-69036670).
- FIGS. IB and 1C show expression of frataxin mRNA following transduction of WT-iPSCs with indicated gRNA and dSaCas9-2xVP64 (FIG. IB) or dSpCas9-2xVP64 (FIG. 1C), as assessed by RT-qPCR.
- FIG. 2 shows CRISPR-Cas mediated transcriptional activation of human frataxin in iPSCs generated from Friedreich’s ataxia patients (FA-iPSCs) using dSaCas9-2xVP64 and promoter-targeting gRNA A or gRNA G, as assessed by RT-qPCR.
- Cell lines harbored expanded GAA trinucleotide repeats left panel: 604/734 repeats; right panel: 867/867 repeats).
- FIG. 3 shows frataxin protein expression levels in WT-iPSCs and FA-iPSCs as assessed by flow cytometry with a mouse monoclonal anti-frataxin AlexaFluor488-conjugated antibody (abeam abl56033).
- FA-iPSCs exhibit reduced frataxin expression in comparison to WT-iPSC control cells.
- FIG. 4 shows scatterplot of results from sequencing analysis for screen of frataxin-saturating gRNA library.
- WT-iPSCs expressing dSaCas9-2xVP64 were transduced with pooled library of gRNAs and subsequently sorted by flow cytometry into populations representing top 10% and bottom 10% of cells based on frataxin protein expression.
- Populations were sequenced to identify gRNAs enriched in the frataxin-high population based on DESeq2.
- Each dot in the scatterplot represents a single gRNA.
- the y- axis represents log fold change in abundance of gRNA in frataxin-high versus frataxin-low population.
- the x-axis represents mean of normalized counts.
- the promoter-targeting gRNA A and enhancertargeting gRNA U were significantly enriched in the frataxin-high population.
- FIG. 5 shows locations of promoter-targeting gRNA A and enhancer-targeting gRNA U with respect to the frataxin gene and indicators of chromatin state H3K4Mel, H3K4Me3, H3K27Ac, and DNase I hypersensitivity based on data from the Encyclopedia of DNA Elements (ENCODE).
- gRNA U targets a region (hg38 chr9:69, 027, 282-69, 028, 497) exhibiting hallmark chromatin signatures of an enhancer element, including H3K4Mel and DNase I hypersensitivity.
- FIG. 6 shows CRISPR-Cas mediated transcriptional activation of frataxin in iPSCs, using dSaCas9-2xVP64 and promoter-targeting gRNA A or enhancer-targeting gRNA U compared to control gRNA or empty gRNA vector control, as assessed by RT-qPCR.
- iPSCs harbored normal GAA trinucleotide repeat lengths.
- FIGS. 7A-7F show combinatorial modulation of frataxin mRNA expression with promotertargeting (gRNA A and gRNA G) and enhancer-targeting (gRNA U) gRNAs, as assessed by RT-qPCR.
- WT-iPSCs or FA-iPSCs with short or long trinucleotide repeats were used to create stable cell lines expressing promoter-targeting gRNA A or gRNA G.
- Cell lines were transduced with different combinations of dSaCas9-2xVP64 and gRNA, as indicated. Results are shown for cell lines WT-gRNA A-iPSC (FIG. 7A), WT-gRNA G-iPSC (FIG.
- FIG. 8 shows the rescue of frataxin mRNA expression with promoter- and enhancertargeting gRNAs in FA-iPSCs, with frataxin expression levels compared to WT-iPSCs, as assessed by RT-qPCR.
- FA-iPSCs stably expressing promoter-targeting gRNA A or gRNA G were transduced with different combinations of dSaCas9-2xVP64 and gRNA, as indicated.
- FIGS. 9A-9C show CRISPR-Cas mediated activation of frataxin protein expression using promoter- and enhancer-targeting gRNAs, as assessed by ELISA.
- FIG. 9A shows frataxin protein expression in WT-iPSCs stably expressing gRNA G (WT-gRNA G-iPSCs), which were (a) left untreated, (b) transduced with dSaCas9-2xVP64 and a control gRNA, or (c) transduced with dSaCas9- 2xVP64 and the frataxin enhancer-targeting gRNA U.
- FIGS. 9A shows frataxin protein expression in WT-iPSCs stably expressing gRNA G (WT-gRNA G-iPSCs), which were (a) left untreated, (b) transduced with dSaCas9-2xVP64 and a control gRNA, or (c) transduced with dSaCa
- FIG. 9B and 9C show rescue of frataxin protein expression in FA-iPSCs harboring short (FIG. 9B) or long (FIG. 9C) expanded trinucleotide GAA repeats in comparison to WT-iPSCs.
- FA-iPSCs stably expressing promoter-targeting gRNAs were transduced with different combinations of dSaCas9-2xVP64 and gRNA, as indicated.
- FIGS. 10A and 10B show scatterplots of results from sequencing analysis for screen of dSaCas9 transcription activator and repressor fusion proteins.
- WT-iPSCs expressing frataxin promotertargeting gRNA A were transduced with pooled libraries of fusion proteins comprising fragments of nuclear localized proteins, fused to the N-terminus (FIG. 10A) or C-terminus (FIG. 10B) of dSaCas9.
- Transduced cells were subsequently sorted by flow cytometry into populations representing top 10% and bottom 10% of cells based on frataxin protein expression.
- Populations were sequenced to identify effectors enriched in the frataxin-high or frataxin-low populations based on DESeq2.
- Each dot in the scatterplots represents a single effector. Effectors in the top and bottom 10% of cells based on frataxin protein expression are indicated by black dots, and other effectors are indicated by gray dots.
- the y-axis represents log fold change in frataxin-high versus frataxin-low populations, x-axis represents mean of normalized counts. Enriched effectors are highlighted in red, as activators (positive log fold change) and repressors (negative log fold change).
- N-terminal screen identified 9 activators and 211 repressors, C- terminal screen identified 5 activators and 208 repressors.
- FIGS. 11A and 11B show transcriptional activation of frataxin in WT-iPSCs facilitated by dSaCas9 transcription activator N-terminal (FIG. 11A) and C-terminal (FIG. 11B) fusion proteins identified in the screen.
- WT-iPSCs stably expressing frataxin promoter-targeting gRNA A were transduced with dSaCas9 fusion proteins comprising indicated effectors, including positive control (2xVP64) and negative control peptides. Expression was assessed by RT-qPCR in comparison to negative control.
- FIGS. 12A and 12B show transcriptional activation of frataxin in FA-iPSCs harboring long trinucleotide repeats facilitated by dSaCas9 transcription activator N-terminal (FIG. 12A) and C-terminal (FIG. 12B) fusion proteins identified in the screen.
- FA-iPSCs stably expressing frataxin promotertargeting gRNA A were transduced with dSaCas9 fusion proteins comprising indicated effectors, including positive control (2xVP64) and negative control peptides. Expression was assessed by RT- qPCR in comparison to negative control.
- FIGS. 13A and 13B show frataxin mRNA expression (FIG. 13A) as assessed by qRT-PCR and frataxin protein expression (FIG. 13B) as assessed by ELISA, in cardiomyocytes derived from FA- iPSCs or WT-iPSCs for the indicated conditions.
- FA-iPSC-derived cardiomyocytes were transduced with dSaCas9-2xVP64 and indicated FXN-targeting gRNA(s).
- Negative control cells were transduced with dSaCas9-2xVP64 and a non-targeting gRNA (NT gRNA), or with a puromycin resistance cassette alone (puro control (FA)). Expression is assessed relative to WT control cells (puro control (WT)).
- FIG. 14 shows frataxin mRNA expression as assessed by qRT-PCR in neurons derived from FA-iPSCs or WT-iPSCs for the indicated conditions.
- FA-iPSC-derived neurons were transduced with dSaCas9-2xVP64 and indicated FXN-targeting gRNA(s).
- Negative control cells were transduced with dSaCas9-2xVP64 and a non-targeting gRNA (NT gRNA), or with a puromycin resistance cassette alone (puro control (FA)). Expression is assessed relative to WT control cells (puro control (WT)).
- FIG. 15 shows frataxin mRNA expression as assessed by qRT-PCR in non-human primate fibroblasts from Crab-eating monkey (Macaco fascicularis) and Rhesus monkey (Macaco mulatto) for the indicated conditions.
- Cells were transduced with dSaCas9-2xVP64 and indicated FXN-targeting gRNA(s).
- Negative control cells were transduced with dSaCas9-2xVP64 and a non-targeting gRNA (NT gRNA), or with a puromycin resistance cassette alone (puro control).
- FIGS. 16A and 16B show frataxin protein expression as assessed by EEISA in the liver of a humanized FA mouse model or healthy control, treated with AAV9 vectors encoding dSaCas9-2xVP64 and indicated FXN-targeting gRNA(s).
- FIG. 16A indicates FXN protein pg per pg of loaded protein into the assay [pg/pg protein]
- FIG. 16B indicates FXN protein levels normalized to the average FXN protein levels from healthy control mice.
- FIG. 17A and 17B show frataxin protein expression as assessed by ELISA in the heart of a humanized FA mouse model or healthy control, treated with AAV9 vectors encoding dSaCas9-2xVP64 and indicated FXN-targeting gRNA(s).
- FIG. 17A indicates FXN protein pg per pg of loaded protein into the assay [pg/pg protein]
- FIG. 17B indicates FXN protein levels normalized to the average FXN protein levels from healthy control mice.
- FIGS. 18A and 18B show succinate dehydrogenase (SDH) enzymatic activity in the heart of a humanized FA mouse model or healthy control, treated with AAV9 vectors encoding dSaCas9-2xVP64 and indicated FXN-targeting gRNA(s).
- SDH succinate dehydrogenase
- FIG. 18A indicates mU of SDH activity per mg loaded protein into the assay [mu/mg]
- FIG. 18B indicates SDH activity normalized to the SDH activity from healthy control mice.
- FIGS. 19A and 19B show AAV vector genome quantification as assessed by ddPCR, in the liver (FIG. 19A) and the heart (FIG. 19B) of the humanized FA mouse model or healthy control, treated with AAV9 vectors encoding dSaCas9-2xVP64 and indicated FXN-targeting gRNA(s).
- AAV9 vectors encoding dSaCas9-2xVP64 and a non-targeting gRNA.
- FIG. 20A shows frataxin protein expression in FA-iPSC-derived cardiomyocytes following AAV-DJ delivery of dSaCas9-2xVP64 and the indicated FXN-targeting gRNAs or a non-targeting gRNA (NT).
- FIG. 20B shows frataxin mRNA expression in FA-iPSC-derived neurons following AAV-DJ delivery of dSaCas9-2xVP64 and the indicated non-targeting or FXN-targeting gRNA at various values for multiplicity of infection (MOI).
- MOI multiplicity of infection
- FIGS. 21A and 21B show the multiplicity of infection (MOI) and vector copy number (VCN) in FA-iPSC-derived cardiomyocytes (FIG. 21A) or FA-iPSC-derived neurons (FIG. 21B) following AAV-DJ delivery of dSaCas9-2xVP64 and the indicated non-targeting or FXN-targeting gRNA.
- MOI multiplicity of infection
- VCN vector copy number
- FIG. 22 shows a schematic illustrating an exemplary dSaCas9-tripartite effector fusion protein, with domains from FOXO3 and NCOA3.
- the first domain (labeled “effector”) can comprise different domains, as described in the Examples.
- FIG. 23 shows frataxin protein expression in FA-iPSC-derived cardiomyocytes following AAV-DJ delivery of dSaCas9 fusion proteins with indicated FXN-targeting gRNA G or non-targeting gRNA (NT). Boxes indicating “tripartite effectors” indicate conditions with dSaCas9 fusion proteins with tripartite effectors comprising the indicated domain (e.g. FOXO3, NCOA2, NCOA3, or PYGO1), followed by a domain from FOXO3 and NCOA3, in the N- to C-terminal direction, e.g. as illustrated in FIG. 22.
- indicated domain e.g. FOXO3, NCOA2, NCOA3, or PYGO1
- FIGS. 24A-24C shows results from FA-iPSC-derived cardiomyocytes following AAV-DJ delivery of the indicated dSaCas9 fusion proteins and gRNA G. Shown are MOI versus % of WT FXN protein expression (FIG. 24A), VCN versus % of WT FXN protein expression (FIG. 24B), or a summary table of the results (FIG. 24C).
- FIG. 25 shows VCN versus % of WT FXN protein expression levels in FA-iPSC-derived cardiomyocytes following AAV-DJ delivery of dSaCas9 fusion proteins with the indicated effectors for transcriptional activation.
- Individual domain names e.g. NCOA3
- FOXO3 and NCOA3 e.g. as illustrated in FIG. 22.
- FIGS. 26A and 26B show FXN protein expression levels (in comparison to WT control) in FA-iPSC-derived cardiomyocytes (FIG. 26A) or FA-iPSC-derived neurons (FIG. 26B) following AAV- DJ delivery of dSaCas9 fusion proteins with the indicated effectors for transcriptional activation paired with FXN-targeting gRNA G or non-targeting gRNA (NT).
- Individual domain names e.g., FOXO3, NCOA2, NCOA3
- FIG. 28 shows FXN mRNA expression in FA-iPSC-derived cardiomyocytes following AAV-DJ delivery of dSaCas9-2xVP64 with or without a FLAG epitope tag and the indicated FXN- targeting or non-targeting gRNA.
- FIG. 29 shows FXN mRNA expression at 48 hours after delivery in FA-iPSCs following delivery via electroporation of mRNA encoding eZFP-VP64 fusion proteins comprising the indicated eZFPs.
- FIGS. 30A and 30B show FXN mRNA expression in FA-iPSCs following delivery via electroporation of mRNA encoding eZFP-VP64 fusion proteins comprising the indicated eZFPs, 48 hours (FIG. 30A) and 72 hours (FIG. 30B) after delivery.
- FIG. 30C shows FXN protein expression (measured using ELISA) in FA-iPSCs following delivery via electroporation of mRNA encoding eZFP-VP64 fusion proteins comprising the indicated eZFPs at 72 hours after delivery.
- FIG. 31 shows a map of genomic regions comprising a FXN promoter (top) and FXN enhancer (bottom) targeted by the indicated eZFPs and gRNAs.
- FIG. 32 shows FXN mRNA expression (measured using RT-qPCR) 7 days after delivery in FA-iPSC-derived cardiomyocytes following AAV-DJ delivery of eZFP-VP64 fusion proteins comprising the indicated eZFPs.
- FIG. 33 shows FXN mRNA expression (left), or VP64 mRNA expression (right) as a measure of ZFP-VP64 fusion protein expression, in FA-iPSC-derived cardiomyocytes following AAV- DJ delivery of eZFP-VP64 fusion proteins comprising the indicated eZFPs. Expression levels were measured using RT-qPCR 7 days after delivery.
- FIG. 34 shows FXN mRNA expression (left) or VP64 or Cas9 expression (right) as a measure of expression of the ZFP-VP64 fusion protein or dSaCas9-VP64 fusion protein, in FA-iPSC- derived cardiomyocytes following AAV-DJ delivery of either a) an eZFP-VP64 fusion protein comprising eZFP_A31 or b) dSaCas9-2xVP64 and gRNA G. Expression levels were measured using RT- qPCR 7 days after delivery.
- FIG. 35 shows FXN mRNA expression in FA-iPSC-derived neurons following AAV-DJ delivery of ZFP-VP64 fusion proteins comprising the indicated eZFPs. Expression levels were measured using RT-qPCR 7 days after delivery.
- FIG. 36 shows VP64 mRNA expression levels as a measure of expression of the indicated ZFP-VP64 fusion proteins, in FA-iPSC-derived neurons following AAV-DJ delivery of ZFP-VP64 fusion proteins. Expression levels were measured using RT-qPCR 7 days after delivery.
- FIG. 37 shows FXN mRNA expression levels (as compared to WT cells) in FA-iPSC- derived cardiomyocytes following AAV-DJ delivery of a) the fusion proteins comprising eZFP_A31 and VP64 or the indicated tripartite effectors, or b) dSaCas9 fusion proteins comprising 2xVP64 or the indicated tripartite effectors with gRNA G. Expression levels were measured using RT-qPCR 7 days after delivery.
- FIG. 38 shows FXN mRNA expression levels in FA-iPSC-derived neurons following AAV- DJ delivery of a) the fusion proteins comprising eZFP_A31 and VP64 or the indicated tripartite effectors, or b) dSaCas9 fusion proteins comprising 2xVP64 or the indicated tripartite effectors with gRNA G. Expression levels were measured using RT-qPCR 7 days after delivery.
- FIG. 39 shows FXN mRNA expression levels in FA-iPSC-derived neurons (as compared to WT control cells) following AAV-DJ delivery of fusion proteins comprising eZFP_A31 and indicated tripartite effectors fused to the C-terminus or N-terminus of the eZFP (left). Schematics of each tested fusion protein are also shown (right). Expression levels were measured using RT-qPCR 7 days after delivery.
- FIG. 40A and FIG. 40B show FXN expression levels in the liver of a humanized FA mouse model treated with AAV9 vectors encoding eZFP_A31 or dCas9 fusion proteins containing transcriptional activator VP64 or tripartite effector NFN.
- the vectors also encoded a FXN-targeting gRNA G.
- FA mouse model (FA model) or healthy control mice (WT) were treated with vehicle only or with AAV9 vectors encoding dCas9-VP64 or dCas9-NFN and a non-targeting (NT) gRNA.
- FIG. 40A shows FXN protein expression levels
- FIG. 40B shows FXN mRNA expression levels relative to the healthy control mice.
- FIG. 41A and FIG. 41B show FXN expression levels in the heart of a humanized FA mouse model treated with AAV9 vectors encoding eZFP_A31 or dCas9 fusion proteins containing transcriptional activator VP64 or tripartite effector NFN.
- the vectors also encoded a FXN-targeting gRNA G.
- a FA mouse model (FA model) or healthy control mice (WT) were treated with vehicle only or with AAV9 vectors encoding dCas9-VP64 or dCas9-NFN and a non-targeting (NT) gRNA.
- FIG. 41A shows FXN protein expression levels
- FIG. 41B shows FXN mRNA expression levels relative to healthy control mice.
- FIG. 42A and FIG. 42B show FXN expression levels in the cerebellum of a humanized FA mouse model treated with AAV9 vectors encoding a dCas9 fusion protein containing transcriptional activator VP64 and a FXN-targeting gRNA G.
- a FA mouse model (FA model) or healthy control mice (WT) were treated with vehicle only or with AAV9 vectors encoding dCas9-VP64 or dCas9-NFN and a non-targeting (NT) gRNA.
- FIG. 42A shows FXN protein expression levels
- FIG. 42B shows FXN mRNA expression levels relative to healthy control mice.
- FIG. 43A and FIG. 43B show FXN expression levels following treatment with AAV9 vectors encoding an exemplary fusion protein containing eZFP_A31 and the tripartite effector NFN and the biodistribution of the AAV9 vectors in different tissues.
- a FA mouse model FA model
- WT healthy control mice
- FXN expression levels were measured in the heart (FIG. 43A; left panel), liver (FIG. 43A; middle panel), and cerebellum (FIG. 43B; left panel).
- Biodistribution of the AAV9 vectors was also assessed relative to expression of heart TBP mRNA in heart and liver (FIG. 43A; right panel) or cerebellum (FIG. 43B; right panel).
- FIG. 44A and FIG. 44B show schematics of exemplary AAVDJ constructs used to deliver a fusion protein containing the exemplary eZFP eZFP_A31 and the tripartite effector NFN.
- the original construct (FIG. 44A; top) included, from N-terminus to C-terminus, an elongation factor la short (EFS) promoter, an SV40 NLS (SEQ ID NO: 159), the exemplary eZFP_A31, another SV40 NLS (SEQ ID NO: 159), the tripartite effector NFN, and the poly(A) sequence from SpA.
- EFS elongation factor la short
- the SV40 NLS was substituted for a c-myc NLS (SEQ ID NO: 160).
- Two optimized constructs using either the SV40 or c-myc NLS (FIG. 44B; top and bottom, respectively) substituted the EFS promoter for either a CAG or EFla promoter and the SpA poly (A) sequence for a bGH poly(A) sequence and inserted a 5’ untranslated region (UTR) following the promoter.
- FIG. 45 shows FXN expression levels in cardiomyocytes following delivery of AAVDJ vectors exemplified in FIG. 44A and FIG. 44B that used EFS, CAG, or EFla promoters to express eZFP_A31 fusion proteins containing either a transcriptional activator VP64 or tripartite effector NFN at different dosages.
- FIG. 46 shows FXN expression levels in HEK293 cells three days following transfection using a screen of different eZFP fusion proteins containing different eZFPs, including a subset of eZFPs set forth in Table 2B, and the tripartite effector VP64.
- cells were (a) left untreated, (b) transfected with GFP, (c) transfected with an empty vector, (d) transfected with exemplary eZFP_A31 fusion proteins eZFP_A31-NFN or eZFP_A31-VP64, or (e) transfected with a dCas9-NFN fusion protein paired with FXN-targeting gRNA G (gG) or a non-targeting gRNA (gNT).
- Asterisks (*) indicate screened eZFPs chosen for further characterization.
- FIG. 47 shows FXN expression levels in HEK293 cells three days following transfection using a screen of different eZFP fusion proteins containing different eZFPs, including a subset of eZFPs set forth in Table 2B, and the tripartite effector NFN.
- cells were (a) left untreated, (b) transfected with GFP, (c) transfected with lipid only, (d) transfected with exemplary eZFP_A31 fusion proteins eZFP_A31-NFN or eZFP_A31-VP64, (e) transfected with a dCas9-NFN fusion protein paired with a non-targeting gRNA (gNT), or (f) treated with PBS.
- Asterisks (*) indicate screened eZFPs chosen for further characterization.
- FIG. 48 shows FXN expression levels in HEK293 cells three days following transfection using a screen of different eZFP fusion proteins containing different eZFPs, including a subset of eZFPs set forth in Table 2B, and the tripartite effector NFN.
- cells were (a) transfected with GFP, (b) transfected with a dCas9-NFN fusion protein paired with a non-targeting gRNA (gNT), or (c) transfected with exemplary eZFP_A31-NFN or eZFP_A48-NFN fusion proteins.
- Asterisks (*) indicate screened eZFPs chosen for further characterization.
- FIG. 49 shows FXN expression levels in HEK293 cells three days following transfection to validate top-performing eZFPs screened in FIG. 46 when fused to the tripartite effector NFN.
- cells were transfected with (a) GFP, (b) vector only, (c) a dCas9-NFN fusion proteins paired with a FXN-targeting gRNA G (gG) or non-targeting gRNA (gNT), or (d) exemplary eZFP_A31 fusion proteins containing either VP64 or NFN.
- FIG. 50 shows FXN expression levels in FA patient fibroblasts (relative to fibroblasts transfected with GFP) to validate eZFP-NFN fusion proteins identified in previous screens.
- Transfection with GFP, exemplary eZFP_A31-VP64 fusion protein, or exemplary eZFP_31-NFN fusion protein were used as controls.
- FIG. 51 shows FXN expression levels in FA patient fibroblasts (relative to fibroblasts transfected with GFP) to validate eZFP-NFN fusion proteins identified in previous screens. Controls included leaving cells untreated or transfecting cells with either GFP or exemplary eZFP_A31-NFN fusion proteins.
- FIG. 52 depicts the level of FXN expression of different screened eZFPs based on target site. All target sites were within the FXN promoter and position is depicted relative to the transcriptional start site (TSS). The target sites of exemplary eZFP_A48 and eZFP_A31 are indicated.
- FIG. 53 shows FXN expression levels in FA-cardiomyocyes following treatment with AAVDJ vectors using either CAG or EFS promoters to express exemplary eZFP-NFN fusion proteins at different doses (3E3, 3E4, or 3E5).
- AAVDJ vectors encoding (a) a fusion protein containing a non-targeting eZFP (eZFP- NT) and the tripartite effector NFN (eZFP-NT-NFN), (b) dCas9-NFN fusion protein paired with a nontargeting gRNA (gNT), or (c) GFP.
- GC genetically correct FA-cardiomyocytes were transduced with AAVDJ vectors encoding GFP or FA-cardiomyocytes were transduced with AAVDJ vectors encoding (a) dCas9-NFN fusion protein paired with FXN-targeting gRNA G (gG), or (b) eZFP_A31 fusion proteins containing either the tripartite effector NFN or the transcriptional activator VP64.
- eZFPs engineered zinc finger proteins
- eZFP fusion proteins bind to or target a frataxin (FXN) locus.
- the DNA-targeting systems include fusion proteins.
- the DNA-targeting systems include guide RNAs (gRNAs).
- the DNA-targeting systems include fusion proteins and gRNAs.
- compositions such as engineered zinc finger proteins (eZFPs), eZFP fusion proteins, DNA- targeting systems, including fusion proteins, gRNAs, and pluralities and combinations thereof, that bind to or target a FXN locus.
- fusion proteins such as eZFP fusion proteins and dCas fusion proteins that bind to or target FXN.
- gRNAs that bind to or target FXN.
- the provided eZFPs, eZFP fusion proteins, DNA-targeting systems, including fusion proteins, and/or gRNAs bind to, target, and/or modulate the expression of FXN.
- methods and uses related to any of the provided compositions and combinations for example, in modulating the expression of FXN, and/or in the treatment or therapy of diseases or disorders associated with the activity, function or expression, for example dysregulation or reduced activity, function or expression of FXN, such as FA.
- the provided embodiments are based on an observation described herein that the level of a human FXN locus expression in cells from FA patients, including in induced pluripotent stem cells (iPSCs) generated from FA patient cells, can be increased or restored, for example using eZFP fusion proteins, or using an exemplary DNA-targeting system comprising a deactivated Cas9 (dCas9)-transcriptional activator fusion protein and a gRNA targeting a promoter region or an enhancer region of a human FXN locus.
- iPSCs induced pluripotent stem cells
- results described herein also show that combinations of two or more DNA-targeting systems targeting different target sites of a human FXN locus can result in a synergistic increase in expression of a human FXN locus.
- the provided embodiments are also based on an observation that certain fusion proteins comprising a DNA-targeting domain and an effector domain is identified based on screening a library of effector domains for their effect in increased expression of a human FXN locus.
- the embodiments described herein demonstrate consistent and effective increase or restoration of FXN expression, in cells from patients with FA, at both the mRNA and protein level, supporting the utility of the approaches in treating FA or other diseases or disorders that are associated with reduced activity, mutation and/or dysregulation of expression of FXN.
- FA frataxin
- FXN frataxin
- FA is an autosomal recessive neurodegenerative and cardiac disease, is caused by a trinucleotide repeat expansion mutation in the FXN locus. FA can result in ataxia, areflexia, loss of vibratory sense and proprioception, dysarthria, cardiomyopathy and/or associated arrhythmias, among other symptoms.
- Existing treatment of FA is directed towards symptoms relief and providing support. Treatments that address the fundamental etiology and disease mechanism are needed.
- eZFPs engineered zinc finger proteins
- eZFP fusion proteins DNA-targeting systems
- gRNAs guide RNAs
- polynucleotides polynucleotides, vectors, cells, kits, and pluralities and combinations thereof, and methods and uses thereof, that meet such needs.
- the provided embodiments offer an advantage of targeting regulatory DNA elements of a frataxin locus within a particular genomic region, such as an enhancer region, for modulating transcription. In some aspects, the provided embodiments offer an advantage of facilitating controlled, additive and/or synergistic activation of FXN by targeting two or more sites within regulatory DNA elements of FXN. In some aspects, the provided embodiments offer an advantage of increasing FXN expression to a level that is therapeutically relevant for subjects having a disease or disorder that involve the activity, function or expression of FXN, such as FA. In some aspects, the provided embodiments also offer an advantage of providing various effector domains which are capable of inducing transcription activation, for example, at a particular target locus such as FXN.
- the provided embodiments offer the ability to fine tune and tightly regulate the level of expression and/or activity of frataxin in a cell or a subject.
- the control of the expression and/or activity of frataxin at a particular level or within an optimal window is critical for the survival and normal function of the subject, as the reduction of expression can result in diseases or disorders such as Friedreich’s Ataxia (FA) and in some cases, substantial overexpression can result in toxicity, organ dysfunction, and reduction of life span. Accordingly, the level of expression and/or activity of frataxin must be fine-tuned to be within an optimal window.
- FA Friedreich’s Ataxia
- Frataxin is a protein (exemplary amino acid sequences of human Frataxin Isoform 1: NCBI NM_000144.4, NP_000135 (210 aa); exemplary amino acid sequences of human Frataxin Isoform 2: NM_181425, NP_852090 (196 aa); and exemplary amino acid sequences of human Frataxin Isoform 3: NM_001161706, NP_001155178, (171 aa), Uniprot Q16595; ENTREZ 2395; Ensembl ENSG00000165060; OMIM: 606829) found in cells throughout the body, with the highest levels in tissues with a high metabolic rate including heart, neurons, spinal cord, liver, pancreas (Langerhans cells), and muscles used for voluntary movement (skeletal muscles).
- exemplary amino acid sequences of human Frataxin Isoform 1 NCBI NM_000144.4, NP_000135 (210 aa)
- frataxin is mainly found in mitochondria.
- frataxin promotes the biosynthesis of heme as well as the assembly and repair of iron-sulfur clusters by delivering Fe2 + to proteins involved in these pathways.
- frataxin also plays a primary role in the protection against oxidative stress through its ability to catalyze the oxidation of Fe2 + to Fe3 + and to store large amounts of the metal in the form of a ferrihydrite mineral.
- frataxin is processed in two steps by mitochondrial processing peptidase (MPP). MPP first cleaves the precursor to intermediate form and subsequently converts the intermediate to a mature protein.
- MPP mitochondrial processing peptidase
- FXN frataxin gene
- GAA guanine-adenine-adenine trinucleotide repeat expansions in the first intron of FXN.
- FA is a degenerative neuromuscular disorder, and can result in ataxia, areflexia, loss of vibratory sense and proprioception, dysarthria, cardiomyopathy and/or associated arrhythmias, among other symptoms, and death.
- the class of mutation most commonly associated with FA is unstable hyper-expansion of a GAA trinucleotide repeat located in the first intron of the frataxin gene. In normal subjects, there are approximately 6-34 repeats, whereas expansions associated with FA are typically 150 or more repeats, and may vary from 44 to 1700 repeats, with most abnormal alleles ranging from 600 to 900 repeats. Increased trinucleotide repeat expansion lengths are associated with decreased FXN levels, earlier onset of disease, and increased disease severity. Patients with fewer repeats (150-200) have milder symptoms than those with longer sections with more triplet repeats (350 to 650). In some severely affected patients there are up to 1700 repeats.
- frataxin expansion mutations are located in an intron, the amino acid sequence of the frataxin protein is not altered.
- approximately 1-3% of FA patients are compound heterozygotes with an expansion on one allele and a conventional mutation (e.g. a missense, nonsense, or deletion mutation) on the other.
- a missense mutation e.g. a missense, nonsense, or deletion mutation
- Some patients with a missense mutation can have less severe symptoms because the mutated protein in still functional. FA symptoms are not observed in heterozygous carriers.
- the term "trinucleotide repeat expansion" means a series of three bases (for example, GAA) repeated at least twice.
- the trinucleotide repeat expansion may be located in intron 1 of a FXN locus, gene or nucleic acid.
- a pathogenic trinucleotide repeat expansion includes at least 66 or 70 repeats of GAA in a FXN nucleic acid and is associated with disease.
- a pathogenic trinucleotide repeat expansion includes at least 67, 68, 69, 70, 75, 80, 85, 90, 95, 100, 150, 200, 250, 300, 400, 500, 800, 1000 or more repeats.
- the repeats are consecutive. In certain examples, the repeats are interrupted by one or more nucleobases. In certain examples, a wild- type trinucleotide repeat expansion includes 12 or fewer repeats of GAA in a FXN nucleic acid. In other examples, a wild-type trinucleotide repeat expansion includes 11, 10, 9, 8, 7, 6, 5, 4, 3, 2, or 1 repeat.
- FXN has been associated with diseases and disorders such as, but not limited to, Alzheimer's Disease, Amyotrophic Lateral Sclerosis, Apraxias, Ataxia, Ataxia Telangiectasia, Hereditary Ataxias, Bloom Syndrome, Brain Neoplasms, Malignant tumor of colon, Dilated Cardiomyopathy, Hypertrophic Cardiomyopathy, Cerebellar Ataxia, Cystic Fibrosis, Diabetes, Diabetes Mellitus, Non-Insulin- Dependent Diabetes Mellitus, Dysarthria, Dystonia, Fragile X Syndrome, Friedreich's Ataxia, Heart Diseases, Cardiomegaly, Hemochromatosis, Herpes Simplex Infections, Huntington Disease, Liver neoplasms, Machado-Joseph Disease, Metabolic Diseases, Myocardial Infarction, Myotonic Dystrophy, nervous system disorder, Neuroblastoma, Neuromuscular Diseases, Pallor, Parkinson Disease, Peripher
- the FXN gene is also associated with a number of other disorders, including hereditary ataxia, X-linked sideroblastic anemia with ataxia, tabes dorsalis, spinocerebellar degeneration, tertiary neurosyphilis, a transferrinemia, spinocerebellar ataxia, scoliosis, hemochromatosis, fragile X syndrome, mitochondrial disorders and cardiomyopathy. Modulating expression of the FXN gene using any of the methods described herein may be used to treat, prevent and/or mitigate the symptoms of the diseases and disorders described herein.
- Frataxin is a mitochondrial iron chaperone suggested to be important for iron-sulfur processing. This protein is found throughout the human body, but is enriched in the heart, spinal cord, liver, pancreas and muscles. Expansions of a trinucleotide GA A repeat region in the FXN gene, and subsequent frataxin protein deficiency, causes Friedreich's Ataxia. The GAA repeat region is located in the middle of an Alu element in the first intron of the FXN gene. In most people, the number of GAA repeats in the FXN gene is fewer than 12. Individuals with 12-33 uninterrupted GAA repeats are said to be asymptomatic.
- the GAA segment is abnormally repeated from 66 to over 1,000 times.
- the number of repeats in the gene correlates with the age of onset and severity of the disease.
- Individuals with fewer than 300 GAA repeats tend to have later symptom onset (after age 25) than those with larger GAA trinucleotide repeats (e.g. 600 to 900 repeats).
- the abnormal repeat expansion can result in an RNA processing defect, which leads to dysregulation of translation and reduced amount of the FXN protein in cells.
- FA is usually diagnosed in the first or second decade and affects 1 in 50,000 people in the United States.
- Friedreich's Ataxia is a progressive movement disorder characterized by loss of strength and sensation, muscle stiffness and impaired speech. Individuals with Friedreich's Ataxia may also have cardiomyopathy, diabetes, vision or hearing loss, and/or scoliosis.
- ZFPs zinc finger proteins
- eZFPs engineered zinc finger proteins
- the eZFPs are capable of binding to, or bind to, a target site in a FXN locus, such as a regulatory element of a FXN locus.
- the eZFP can facilitate specific targeting of effector domains for transcriptional activation to the FXN locus, for example for gene-specific transcriptional activation of FXN.
- fusion proteins comprising the eZFP and one or more other elements, such as the effector domains for transcriptional activation.
- the eZFP facilitates increased FXN expression, for example in connection with compositions and methods for treating a disease or disorder associated with FXN expression, such as Friedreich’s ataxia (FA).
- FA Friedreich’s ataxia
- a zinc finger protein (ZFP), a zinc finger DNA binding protein, or zinc finger DNA binding domain, is a protein, or a domain within a larger protein, that binds DNA in a sequence-specific manner through one or more zinc fingers, which are regions of amino acid sequence within the binding domain, having a structure that is stabilized through coordination of a zinc ion.
- ZFPs are artificial, or engineered, ZFPs (eZFPs), comprising ZFP domains targeting specific DNA sequences, typically 9-18 nucleotides long, generated by assembly of individual zinc fingers.
- ZFPs include those in which a single finger domain is approximately 30 amino acids in length and contains an alpha helix containing two invariant histidine residues coordinated through zinc with two cysteines of a single beta turn, and having two, three, four, five, or six fingers.
- sequence-specificity of a ZFP may be altered by making amino acid substitutions at the four helix positions (-1, 2, 3, and 6) on a zinc finger recognition helix, also called a zinc finger recognition region.
- a ZFP or ZFP-containing molecule such as a fusion protein, can be non-naturally occurring, e.g., is engineered to bind to a target site of choice.
- zinc fingers can be custom-designed (i.e. designed by the user), and/or obtained from a commercial source.
- Various methods for designing zinc finger proteins are available. For example, methods for designing zinc finger proteins to bind to a target DNA sequence of interest are described, for example in Liu, Q. et al., PNAS, 94(l l):5525-30 (1997); Wright, D.A. et al., Nat. Protoc., l(3):1637-52 (2006); Gersbach, C.A. et al., Acc. Chem. Res., 47(8):2309-18 (2014); Bhakta M.S. et al., Methods Mol.
- the target site for an eZFP provided herein is in a FXN promoter.
- the target site is within a target region spanning the genomic coordinates chr9:69, 034, 900-69, 035, 900 from hg38 (SEQ ID NO:430).
- the target site is within a target region spanning the genomic coordinates chr9:69, 035, 300-69-035, 800 from hg38.
- the target site is within a target region spanning the genomic coordinates chr9:69,035,350- 69,035,450 from hg38.
- the target site is within a target region spanning the genomic coordinates chr9:69, 035, 400-69, 035, 450 from hg38. In some embodiments, the target site is within a target region spanning the genomic coordinates chr9:69, 035, 530-69, 035, 580 from hg38. In some embodiments, the target site is within a target region spanning the genomic coordinates chr9:69,035,675- 69,035,725 from hg38.
- the target site for an eZFP provided herein is in a FXN enhancer.
- the target site is within a target region spanning the genomic coordinates chr9:69, 027, 282-69, 028, 497 from hg38 (SEQ ID NO:431).
- the target site is within a target region spanning the genomic coordinates chr9:69, 027, 615-69, 028, 101 from hg38.
- the target site is within a target region spanning the genomic coordinates chr9:69,027,775- 69,027,875 from hg38.
- the target site is within a target region spanning the genomic coordinates chr9:69, 027, 795-69, 027, 845 from hg38.
- the target site for an eZFP provided herein comprises the nucleotide sequence set forth in any one of SEQ ID NOS:269-300 and 583-600, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the target site for an eZFP provided herein comprises the nucleotide sequence set forth in any one of SEQ ID NOS:269- 300 and 583-600.
- the target site is comprised in double-stranded DNA, such as genomic DNA.
- the target site is double-stranded DNA, such as genomic DNA.
- the eZFP is capable of binding to the target site.
- the eZFP binds to the target site.
- the binding is target-specific.
- an eZFP binds to the target site, and not to other sites comprising different sequences.
- an individual eZFP disclosed herein binds to the target site set forth in SEQ ID NO:299, and does not bind to a different target site, such as the target site set forth in SEQ ID NO:269.
- the target site for an eZFP provided herein comprises a sequence set forth in Table 1.
- the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:272, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the sequence set forth in SEQ ID NO: 272.
- the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:277, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the sequence set forth in SEQ ID NO: 277.
- the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:280, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the sequence set forth in SEQ ID NO: 280.
- the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:281, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the sequence set forth in SEQ ID NO: 281.
- the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:283, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the sequence set forth in SEQ ID NO: 283.
- the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:290, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the sequence set forth in SEQ ID NO: 290.
- the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:299, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the sequence set forth in SEQ ID NO: 299.
- the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:583, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the sequence set forth in SEQ ID NO:583.
- the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:584, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the sequence set forth in SEQ ID NO:584.
- the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:585, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the sequence set forth in SEQ ID NO:585.
- the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:586, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the sequence set forth in SEQ ID NO:586.
- the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:587, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the sequence set forth in SEQ ID NO:587.
- the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:588, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the sequence set forth in SEQ ID NO:588.
- the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:589, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the sequence set forth in SEQ ID NO:589.
- the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:590, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the sequence set forth in SEQ ID NO:590.
- the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:591, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the sequence set forth in SEQ ID NO:591.
- the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:592, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the sequence set forth in SEQ ID NO:592.
- the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:593, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the sequence set forth in SEQ ID NO:593.
- the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:594, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the sequence set forth in SEQ ID NO:594.
- the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:595, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the sequence set forth in SEQ ID NO:595. [0150] In some embodiments, the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:596, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the sequence set forth in SEQ ID NO:596.
- the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:597, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the sequence set forth in SEQ ID NO:597.
- the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:598, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the sequence set forth in SEQ ID NO:598.
- the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:599, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the sequence set forth in SEQ ID NO:599.
- the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:600, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the sequence set forth in SEQ ID NO:600.
- the eZFP comprises multiple zinc fingers.
- each zinc finger comprises a recognition region.
- the recognition regions together facilitate sequence-specific binding of the eZFP, for example to a specific target site.
- the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding recognition region Fl through F6, which facilitate sequencespecific binding to a specific target site.
- the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, each comprising a corresponding recognition region F1-F6, as shown in Table 2A and Table 2B.
- the recognition regions F1-F6 facilitate specific binding to the indicated target site sequence in Table 2A and Table 2B.
- the eZFP comprises an amino acid sequence comprising the recognition regions, as shown in Table 2A and Table 2B.
- the eZFP can be encoded by a DNA sequence as shown in Table 2A and Table 2B.
- an eZFP such as eZFP_A04 as described herein.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:272, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:272.
- the target site is double-stranded DNA.
- the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: QSGNLAR (SEQ ID NO:341); F2: QKVNRAG (SEQ ID NO:342); F3: DRSNLSR (SEQ ID NO:343); F4: QSGHLSR (SEQ ID NO:344); F5: TSGHLSR (SEQ ID NO:345); F6: RSDALAR (SEQ ID NO:346).
- the eZFP comprises the amino acid sequence set forth in SEQ ID NO:301, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the eZFP comprises the amino acid sequence set forth in SEQ ID NO:301.
- the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:308, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:308.
- an eZFP such as eZFP_A09 as described herein.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:277, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:277.
- the target site is double-stranded DNA.
- the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: RSDNLSE (SEQ ID NO:347); F2: KSWSRYK (SEQ ID NO:348); F3: TSGSLSR (SEQ ID NO:349); F4: RSDALAR (SEQ ID NO:350); F5: RSDNLSV (SEQ ID NO:351); F6: FSSCRSA (SEQ ID NO:352).
- the eZFP comprises the amino acid sequence set forth in SEQ ID NO:302, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the eZFP comprises the amino acid sequence set forth in SEQ ID NO:302.
- the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:309, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:309.
- an eZFP such as eZFP_A12 as described herein.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:280, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:280.
- the target site is double-stranded DNA.
- the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: TSGNLTR (SEQ ID NO:353); F2: EQTTRDK (SEQ ID NO:354); F3: RSANLAR (SEQ ID NO:355); F4: RLDNRTA (SEQ ID NO:356); F5: DSSHRTR (SEQ ID NO:357); and F6: RKYYLAK (SEQ ID NO:358).
- the eZFP comprises the amino acid sequence set forth in SEQ ID NO:303, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:303. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:310, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:310.
- an eZFP such as eZFP_A13 as described herein.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:281, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:281.
- the target site is double-stranded DNA.
- the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: RSAHESR (SEQ ID NO:359); F2: DRSDESR (SEQ ID NO:360); F3: RSDHESV (SEQ ID NO:361); F4: RSDVRKT (SEQ ID NO:362); F5: QSGAEAR (SEQ ID NO:363); and F6: RKYYLAK (SEQ ID NO:364).
- the eZFP comprises the amino acid sequence set forth in SEQ ID NO:304, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the eZFP comprises the amino acid sequence set forth in SEQ ID NO:304.
- the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:311, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:311.
- an eZFP such as eZFP_A15 as described herein.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:283, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:283.
- the target site is double-stranded DNA.
- the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: RSAHESR (SEQ ID NO:365); F2: RSDAEAR (SEQ ID NO:366); F3: ATSNRSA (SEQ ID NO:367); F4: RSAHESR (SEQ ID NO:368); F5: TSGSLSR (SEQ ID NO:369); and F6: QSGDLTR (SEQ ID NO:370).
- the eZFP comprises the amino acid sequence set forth in SEQ ID NO:305, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the eZFP comprises the amino acid sequence set forth in SEQ ID NO:305.
- the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:312, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:312.
- an eZFP such as eZFP_A22 as described herein.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:290, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:290.
- the target site is double-stranded DNA.
- the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: QSGDETR (SEQ ID NO:371); F2: QSSDERR (SEQ ID NO:372); F3: RSDNESE (SEQ ID NO:373); F4: SSRNEAS (SEQ ID NO:374); F5: DRSHETR (SEQ ID NO:375); and F6: RSDDLTR (SEQ ID NO:376).
- the eZFP comprises the amino acid sequence set forth in SEQ ID NO:306, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the eZFP comprises the amino acid sequence set forth in SEQ ID NO:306.
- the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:313, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:313.
- an eZFP such as eZFP_A31 as described herein.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:299, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:299.
- the target site is double-stranded DNA.
- the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: ERHHETR (SEQ ID NO:377); F2: QSAHEKA (SEQ ID NO:378); F3: EPQTEQR (SEQ ID NO:379); F4: QNATRTK (SEQ ID NO:380); F5: QSSHLTR (SEQ ID NO:381); and F6: RSDHLSR (SEQ ID NO:382).
- the eZFP comprises the amino acid sequence set forth in SEQ ID NO:307, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the eZFP comprises the amino acid sequence set forth in SEQ ID NO:307.
- the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:314, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NOG 14.
- an eZFP such as eZFP_A40 as described herein.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:583, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:583.
- the target site is double-stranded DNA.
- the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: RSDSLLR (SEQ ID NO:475); F2: TSSNRKT (SEQ ID NO:476); F3: RSAHLSR (SEQ ID NO:477); F4: TSGSLTR (SEQ ID NO:478); F5: QSGDLTR (SEQ ID NO:479); and F6: QWGTRYR (SEQ ID NO:480).
- the eZFP comprises the amino acid sequence set forth in SEQ ID NO:439, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the eZFP comprises the amino acid sequence set forth in SEQ ID NO:439.
- the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:457, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:457.
- an eZFP such as eZFP_A41 as described herein.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:584, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:584.
- the target site is double-stranded DNA.
- the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: QARHLTC (SEQ ID NO:481); F2: QSGHLSR (SEQ ID NO:482); F3: RSDVLSE (SEQ ID NO:483); F4: KHSTRRV (SEQ ID NO:484); F5: QSSDLSR (SEQ ID NO:485); and F6: WKWNLRA (SEQ ID NO:486).
- the eZFP comprises the amino acid sequence set forth in SEQ ID NO:440, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:440. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:458, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:458.
- an eZFP such as eZFP_A42 as described herein.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:585, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:585.
- the target site is double-stranded DNA.
- the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: RSDNEAR (SEQ ID NO:487); F2: WRGDRVK (SEQ ID NO:488); F3: YKHVESD (SEQ ID NO:489); F4: TSGSETR (SEQ ID NO:490); F5: QSGNLAR (SEQ ID NO:491); and F6: RARDLSK (SEQ ID NO:492).
- the eZFP comprises the amino acid sequence set forth in SEQ ID NO: 441, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the eZFP comprises the amino acid sequence set forth in SEQ ID NO:441.
- the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:459, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:459.
- an eZFP such as eZFP_A43 as described herein.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:586, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:586.
- the target site is double-stranded DNA.
- the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: QSANRTK (SEQ ID NO:493); F2: QSGNEAR (SEQ ID NO:494); F3: RSDNESV (SEQ ID NO:495); F4: IRSTLRD (SEQ ID NO:496); F5: QNAHRKT (SEQ ID NO:497); and F6: HRSSLRR (SEQ ID NO:498).
- the eZFP comprises the amino acid sequence set forth in SEQ ID NO:442, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:442. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:460, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:460.
- an eZFP such as eZFP_A44 as described herein.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:587, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:587.
- the target site is double-stranded DNA.
- the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: QAGNRST (SEQ ID NO:499); F2: DRSALAR (SEQ ID NO:500); F3: RSDNLAR (SEQ ID NO:501); F4: WRGDRVK (SEQ ID NO:502); F5: YKHVLSD (SEQ ID NO:503); and F6: TSGSLTR (SEQ ID NO:504).
- the eZFP comprises the amino acid sequence set forth in SEQ ID NO: 443, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:443. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:461, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:461.
- an eZFP such as eZFP_A45 as described herein.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:588, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:588.
- the target site is double-stranded DNA.
- the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: RSDNLSV (SEQ ID NO:505); F2: IRSTLRD (SEQ ID NO:506); F3: QNAHRKT (SEQ ID NO:507); F4: HRSSLRR (SEQ ID NO:508); F5: RSDNLAR (SEQ ID NO:509); and F6: QRSPLPA (SEQ ID NO:51Q).
- the eZFP comprises the amino acid sequence set forth in SEQ ID NO:444, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:444. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:462, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:462.
- an eZFP such as eZFP_A46 as described herein.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:589, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:589.
- the target site is double-stranded DNA.
- the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: DRSTRTK (SEQ ID NO:511); F2: RSDYLAK (SEQ ID NO:512); F3: LRHHLTR (SEQ ID NO:513); F4: QSAHLKA (SEQ ID NO:514); F5: LPQTLQR (SEQ ID NO:515); and F6: QNATRTK (SEQ ID NO:516).
- the eZFP comprises the amino acid sequence set forth in SEQ ID NO: 445, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the eZFP comprises the amino acid sequence set forth in SEQ ID NO:445.
- the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:463, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:463.
- an eZFP such as eZFP_A47 as described herein.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:590, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:590.
- the target site is double-stranded DNA.
- the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: RSADLTR (SEQ ID NO:517); F2: RSDDLTR (SEQ ID NO:518); F3: QSSDLSR (SEQ ID NO:519); F4: WHSSLHQ (SEQ ID NO:520); F5: RSDSLSQ (SEQ ID NO:521); and F6: RKADRTR (SEQ ID NO:522).
- the eZFP comprises the amino acid sequence set forth in SEQ ID NO:446, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the eZFP comprises the amino acid sequence set forth in SEQ ID NO:446.
- the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:464, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:464.
- an eZFP such as eZFP_A48 as described herein.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:591, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:591.
- the target site is double-stranded DNA.
- the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: RNDALTE (SEQ ID NO:523); F2: RKDNLKN (SEQ ID NO:524); F3: TSGELVR (SEQ ID NO:525); F4: HRTTLTN (SEQ ID NO:526); F5: TTGNLTV (SEQ ID NO:527); and F6: RTDTLRD (SEQ ID NO:528).
- the eZFP comprises the amino acid sequence set forth in SEQ ID NO: 447, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the eZFP comprises the amino acid sequence set forth in SEQ ID NO:447.
- the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:465, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:465.
- an eZFP such as eZFP_A49 as described herein.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:592, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:592.
- the target site is double-stranded DNA.
- the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: RKDNEKN (SEQ ID NO:529); F2: RADNETE (SEQ ID NO:530); F3: TSHSETE (SEQ ID NO:531); F4: SKKHLAE (SEQ ID NO:532); F5: TSGELVR (SEQ ID NO:533); and F6: TSGELVR (SEQ ID NO:534).
- the eZFP comprises the amino acid sequence set forth in SEQ ID NO:448, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the eZFP comprises the amino acid sequence set forth in SEQ ID NO:448.
- the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:466, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:466.
- an eZFP such as eZFP_A50 as described herein.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:593, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:593.
- the target site is double-stranded DNA.
- the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: THEDEIR (SEQ ID NO:535); F2: DCRDEAR (SEQ ID NO:536); F3: RSDELVR (SEQ ID NO:537); F4: RNDALTE (SEQ ID NO:538); F5: SKKHLAE (SEQ ID NO:539); and F6: QSGHLTE (SEQ ID NO:540).
- the eZFP comprises the amino acid sequence set forth in SEQ ID NO: 449, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the eZFP comprises the amino acid sequence set forth in SEQ ID NO:449.
- the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:467, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:467.
- an eZFP such as eZFP_A51 as described herein.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:594, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:594.
- the target site is double-stranded DNA.
- the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: HTGHEEE (SEQ ID NO:541); F2: DPGHEVR (SEQ ID NO:542); F3: THEDEIR (SEQ ID NO:543); F4: DCRDEAR (SEQ ID NO:544); F5: RSDEEVR (SEQ ID NO:545); and F6: RNDALTE (SEQ ID NO:546).
- the eZFP comprises the amino acid sequence set forth in SEQ ID NO:450, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the eZFP comprises the amino acid sequence set forth in SEQ ID NO:450.
- the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:468, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:468.
- an eZFP such as eZFP_A52 as described herein.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:595, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:595.
- the target site is double-stranded DNA.
- the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: RSDKEVR (SEQ ID NO:547); F2: RSDHETT (SEQ ID NO:548); F3: RNDAETE (SEQ ID NO:549); F4: TTGAETE (SEQ ID NO:550); F5: THLDLIR (SEQ ID NO:551); and F6: DPGHLVR (SEQ ID NO:552).
- the eZFP comprises the amino acid sequence set forth in SEQ ID NO: 451, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the eZFP comprises the amino acid sequence set forth in SEQ ID NO:45E
- the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:469, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:469.
- an eZFP such as eZFP_A53 as described herein.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:596, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:596.
- the target site is double-stranded DNA.
- the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: TKNSLTE (SEQ ID NO:553); F2: QLAHLRA (SEQ ID NO:554); F3: TSGSLVR (SEQ ID NO:555); F4: RSDNLVR (SEQ ID NO:556); F5: QNSTLTE (SEQ ID NO:557); and F6: RADNLTE (SEQ ID NO:558).
- the eZFP comprises the amino acid sequence set forth in SEQ ID NO:452, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:452. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:470, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:470.
- an eZFP such as eZFP_A54 as described herein.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:597, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:597.
- the target site is double-stranded DNA.
- the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: RADNLTE (SEQ ID NO:559); F2: TKNSLTE (SEQ ID NO:560); F3: QLAHLRA (SEQ ID NO:561); F4: TSGSLVR (SEQ ID NO:562); F5: RSDNLVR (SEQ ID NO:563); and F6: QNSTLTE (SEQ ID NO:564).
- the eZFP comprises the amino acid sequence set forth in SEQ ID NO: 453, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:453. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:471, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:471.
- an eZFP such eZFP_A55 as described herein.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:598, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:598.
- the target site is double-stranded DNA.
- the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: TSGHLVR (SEQ ID NO:565); F2: QLAHLRA (SEQ ID NO:566); F3: TSGELVR (SEQ ID NO:567); F4: QSGDLRR (SEQ ID NO:568); F5: QRAHLER (SEQ ID NO:569); and F6: RSDKLVR (SEQ ID NO:570).
- the eZFP comprises the amino acid sequence set forth in SEQ ID NO:454, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:454. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:472, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:472.
- an eZFP such as eZFP_A56 as described herein.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:599, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:599.
- the target site is double-stranded DNA.
- the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: REDNLHT (SEQ ID NO:571); F2: TSGHLVR (SEQ ID NO:572); F3: QLAHLRA (SEQ ID NO:573); F4: TSGELVR (SEQ ID NO:574); F5: QSGDLRR (SEQ ID NO:575); and F6: QRAHLER (SEQ ID NO:576).
- the eZFP comprises the amino acid sequence set forth in SEQ ID NO:455, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the eZFP comprises the amino acid sequence set forth in SEQ ID NO:455.
- the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:473, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:473.
- an eZFP such eZFP_A57 as described herein.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NG:600, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NG:600.
- the target site is double-stranded DNA.
- the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: QRSDLTR (SEQ ID NO:577); F2: QGGTLRR (SEQ ID NO:578); F3: TSAHLAR (SEQ ID NO:579); F4: RREHLVR (SEQ ID NO:580); F5: QRHGLSS (SEQ ID NO:581); and F6: QRNALRG (SEQ ID NO:582).
- the eZFP comprises the amino acid sequence set forth in SEQ ID NO:456, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:456. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:474, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:474.
- DNA-targeting systems comprising a DNA-targeting domain that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus.
- FXN frataxin
- Exemplary components and features of the DNA-targeting systems are provided herein.
- the DNA-targeting system comprises one or more of any of the components described herein, such as one or more DNA-targeting domains, one or more fusion proteins, such as one or more fusion proteins comprising one or more DNA- targeting domains and one or more effector domains, one or more gRNAs, or any component, portion or fragment thereof, or any combination thereof.
- a DNA-targeting system herein comprises an eZFP and/or eZFP fusion protein, such as any described above in Section I.
- the DNA-targeting system comprises one or more of the eZFP fusion proteins.
- the DNA-targeting system comprises at least two eZFP fusion proteins.
- the two eZFP fusion proteins of the DNA-targeting system are any of the eZFP fusion proteins provided herein, such as any of the eZFP fusion proteins shown in Table 2A and Table 2B.
- compositions such as DNA-targeting systems that bind to or target a frataxin (FXN) locus.
- the provided DNA-targeting systems include fusion proteins, such as eZFP fusion proteins or dCas fusion proteins.
- the DNA-targeting system comprises one or more guide RNA (gRNA).
- gRNA guide RNA
- polynucleotides, vectors that encode any of the DNA-targeting systems, fusion proteins and/or components of kits are cells, kits, systems and pluralities and combinations thereof, that comprise any of the DNA-targeting systems, fusion proteins or gRNAs described herein.
- DNA-targeting systems comprising a DNA-targeting domain that binds to a target site in a FXN locus, such as a regulatory DNA element of a frataxin (FXN) locus.
- a target site in a FXN locus such as a regulatory DNA element of a frataxin (FXN) locus.
- binding of the DNA-targeting domain to the target site does not introduce a genetic disruption or a DNA break at or near the target site.
- the provided DNA-targeting systems comprise a fusion protein comprising a DNA-targeting domain and an effector domain, and binds to a target site in a regulatory DNA element of a FXN locus.
- the DNA-targeting system comprises an eZFP, and/or an eZFP fusion protein, which binds to the target site.
- the DNA-targeting system comprises a guide RNA (gRNA), which targets the DNA-targeting domain to the target site (e.g., as in the case of CRISPR/Cas-based DNA-targeting systems).
- gRNA guide RNA
- the provided DNA-targeting systems can lead to an increase of or a restoration of, the activity, function or expression of FXN.
- compositions for example, in modulating the expression of FXN, and/or in the treatment or therapy of diseases or disorders associated with dysregulation or reduced activity, function or expression of FXN, such as FA.
- the DNA-targeting systems are targeted to one or more target sites located within a FXN locus, such as a regulatory DNA element of a FXN locus, such as a promoter or an enhancer. In some embodiments, the DNA-targeting systems are targeted to at least 2, 3, 4, 5, 6, 7, 8, 9 or 10 target sites within a FXN locus. In some embodiments, the DNA-targeting systems are targeted to one or more target sites located within a promoter of a FXN locus, and one or more target sites located within an enhancer of a FXN locus.
- the DNA-targeting system comprises a DNA-targeting domain comprising a Clustered Regularly Interspaced Short Palindromic Repeats associated (Cas)-guide RNA (gRNA) combination comprising (a) a Cas protein or a variant thereof and (b) at least one gRNA; a zinc finger protein (ZFP), such as an engineered zinc finger protein (eZFP); a transcription activator-like effector (TALE); a meganuclease; a homing endonuclease; or an I-Scel enzyme or a variant thereof.
- the DNA-targeting domain comprises a catalytically inactive variant of any of the foregoing.
- the DNA-targeting system comprises a DNA-targeting domain comprising a Cas-gRNA combination comprising (a) a Cas protein or a variant thereof, and (b) at least one gRNA.
- the at least one gRNA comprises at least 2, 3, 4, 5, 6, 7, 8, 9 or 10 gRNAs.
- the gRNAs are targeted to one or more target sites located within a regulatory DNA element of a FXN locus, such as a promoter or an enhancer.
- the gRNAs are targeted to one or more target sites located within a promoter of a FXN locus, and one or more target sites located within an enhancer of a FXN locus.
- the provided embodiments involve modulating transcription of an endogenous FXN locus in a cell. In some aspects, the provided embodiments involve increasing transcription of an endogenous FXN locus in a cell.
- the cell such as the cell to be treated with the provided embodiments, has a GA A trinucleotide repeat expansion in the FXN locus.
- the cell such as the cell to be treated with the provided embodiments, is from or in a subject with Friedreich’s ataxia. In some embodiments, the cell, such as the cell to be treated with the provided embodiments, exhibits reduced expression of FXN compared to a cell from a subject without Friedreich’ s ataxia.
- the expression of FXN is increased at least about 1.2-fold, 1.25-fold, 1.3-fold, 1.4-fold, 1.5-fold, 1.6-fold, 1.7-fold, 1.75-fold, 1.8-fold, 1.9-fold, 2-fold, 2.5-fold, 3-fold, 4-fold, or 5-fold, compared to a cell that has not been introduced or contacted.
- the expression is increased by less than about 10-fold, 9-fold, 8-fold, 7-fold or 6-fold.
- the subject is a human.
- the cell is a heart cell, a skeletal muscle cell, a nervous system cell, or an induced pluripotent stem cell.
- the introducing, contacting or administering is carried out in vivo or ex vivo.
- the DNA-targeting system comprises a DNA-targeting domain and one or more guide RNAs (gRNAs). In some aspects, the DNA-targeting system comprises a fusion protein and one or more gRNAs. In some aspects, the DNA-targeting system comprises a DNA-targeting domain and a gRNA. In some aspects, the DNA-targeting system comprises a fusion protein. In some aspects, the DNA-targeting system comprises a fusion protein and a gRNA. In some aspects, the DNA-targeting system comprises a DNA-targeting domain.
- gRNAs guide RNAs
- binding of the DNA-targeting domain to the target site does not introduce a genetic disruption or a DNA break at or near the target site.
- DNA-targeting systems capable of specifically targeting a target site in a FXN gene or DNA regulatory element thereof, and increasing transcription of the FXN gene.
- the DNA-targeting systems include a DNA-targeting domain that binds to a target site in the FXN gene or regulatory DNA element thereof.
- the DNA- targeting systems additionally include at least one effector domain that is able to epigenetically modify one or more DNA bases of the FXN gene or regulatory element thereof, in which the epigenetic modification results in an increase in transcription of the FXN gene (e.g. activates transcription or increases transcription of FXN compared to the absence of the DNA-targeting system).
- the terms DNA-targeting system and epigenetic-modifying DNA targeting system may be used herein interchangeably.
- the DNA-targeting system includes a fusion protein comprising (a) a DNA-targeting domain capable of being targeted to the target site; and (b) at least one effector domain capable of increasing transcription of the FXN gene.
- the at least one effector domain is a transcription activation domain.
- the DNA-targeting domain comprises or is derived from a CRISPR associated (Cas) protein, zinc finger protein (ZFP), transcription activator-like effectors (TALE), meganuclease, homing endonuclease, I-Scel enzyme, or variants thereof.
- the DNA-targeting domain comprises a catalytically inactive (e.g. nuclease-inactive or nuclease-inactivated) variant of any of the foregoing.
- the DNA-targeting domain comprises a deactivated Cas9 (dCas9) protein or variant thereof that is a catalytically inactivated so that it is inactive for nuclease activity and is not able to cleave the DNA.
- dCas9 deactivated Cas9
- the DNA-targeting domain comprises or is derived from a Cas protein or variant thereof, such as a nuclease-inactive Cas or dCas (e.g. dCas9, and the DNA-targeting system comprises one or more guide RNAs (gRNAs).
- the gRNA comprises a spacer sequence that is capable of targeting and/or hybridizing to the target site.
- the gRNA is capable of complexing with the Cas protein or variant thereof.
- the gRNA directs or recruits the Cas protein or variant thereof to the target site.
- the effector domain comprises a transcription activation domain, and/or is capable of increasing transcription of the gene.
- the effector domain induces, catalyzes or leads to transcription activation, transcription co-activation, or transcription elongation.
- the effector domain is selected from VP64, p65, Rta, p300, CBP, VPR, VPH, Rta, p300, HSF1, a TET protein (e.g. TET1), SunTag, a partially or fully functional fragment or domain thereof, or a combination of any of the foregoing.
- the effector domain may include a domain, portion, or variant of a protein selected from: DPOLA, ENL, FOXO3, HSH2D, NCOA2, NCOA3, PSA1, PYGO1, RBM39, HERC2, and NOTCH2.
- the effector domain is VP64.
- the fusion protein of the DNA-targeting system comprises dCas9-VP64.
- the fusion protein of the DNA-targeting system is an eZFP fusion protein, such as any eZFP fusion protein disclosed herein, such as in Section II.C.
- the DNA-targeting system comprises a DNA-targeting domain.
- the DNA-targeting domain comprises a DNA-binding protein or DNA-binding nucleic acid.
- the DNA-targeting domain specifically binds to or hybridizes to a particular site or position in the genome, e.g., a target, target site, or target position.
- the DNA-targeting domain is coupled to, fused to or complexed with an effector domain, such as any effector domain described herein, for example, in Section II.B.
- the DNA-targeting system comprises various components, such as an RNA-guided nuclease, variant thereof (such as dCas), or fusion protein comprising the RNA-guided nuclease or variant thereof, or a fusion protein comprising a DNA-targeting domain and an effector domain.
- the DNA-targeting system comprises a DNA-targeting molecule that comprises a DNA-binding protein such as one or more zinc finger protein (ZFP) or transcription activator-like effectors (TALEs), fused to an effector domain.
- the DNA-binding protein of the DNA-targeting molecule comprises an eZFP (e.g., the DNA-targeting molecule is an eZFP fusion protein).
- the DNA-targeting system specifically targets at least one target site in a regulatory DNA element of a frataxin (FXN) locus.
- the DNA-targeting system comprises a ZFP, a TAEE, or a CRISPR/Cas9 combination, that specifically binds to, recognizes, or hybridizes to the target site(s).
- the CRISPR/Cas9 system includes an engineered crRNA/tracr RNA (i.e. “single guide RNA”).
- the DNA-targeting system comprises nucleases or variants thereof based on the Argonaute system (e.g., from T. thermophilus, known as ‘TtAgo’ (Swarts et al., (2014) Nature 507(7491): 258-261).
- the DNA-targeting domain comprises a Clustered Regularly Interspaced Short Palindromic Repeats associated (Cas)-guide RNA (gRNA) combination that includes (a) a Cas protein or a variant thereof and (b) at least one gRNA; a zinc finger protein (ZFP) (such as an eZFP); a transcription activator-like effector (TALE); a meganuclease; a homing endonuclease; or a I- Scel enzyme or a variant thereof.
- the DNA-targeting domain comprises a catalytically inactive variant of any of the foregoing.
- the DNA-targeting domain comprises a Cas-gRNA combination that includes (a) a Cas protein or a variant thereof and (b) at least one gRNA.
- the variant Cas protein lacks nuclease activity or is a deactivated Cas (dCas) protein.
- DNA-targeting systems comprising a DNA-targeting domain, that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus and comprises a Cas-guide RNA (gRNA) combination comprising: (a) a variant Cas protein that lacks nuclease activity or that is a deactivated Cas (dCas) protein; and (b) at least one gRNA comprising at least one gRNA spacer sequence that is capable of hybridizing to the target site or is complementary to the target site.
- gRNA Cas-guide RNA
- DNA-targeting systems comprising a DNA-targeting domain that comprises a Cas-guide RNA (gRNA) combination comprising: (a) a Staphylococcus aureus deactivated Cas9 protein (dSaCas9) protein set forth in SEQ ID NO:72 fused to at least one effector domain that induces transcription activation; and (b) at least one gRNA comprising the gRNA spacer sequence set forth in SEQ ID NO:42.
- gRNA Cas-guide RNA
- DNA-targeting systems comprising a DNA-targeting domain that comprises a Cas-guide RNA (gRNA) combination comprising: (a) a Staphylococcus aureus deactivated Cas9 protein (dSaCas9) protein set forth in SEQ ID NO:72 fused to at least one effector domain that induces transcription activation; and (b) at least one gRNA comprising the gRNA spacer sequence set forth in SEQ ID NO:22.
- gRNA Cas-guide RNA
- DNA-targeting systems comprising a DNA-targeting domain that comprises a Cas-guide RNA (gRNA) combination comprising: (a) a Staphylococcus aureus deactivated Cas9 protein (dSaCas9) protein set forth in SEQ ID NO:72 fused to at least one effector domain that induces transcription activation; and (b) at least one gRNA comprising the gRNA spacer sequence set forth in SEQ ID NO:28.
- gRNA Cas-guide RNA
- the provided DNA-targeting systems or fusion proteins comprise a DNA-targeting domain.
- the DNA-targeting domain provides sequence specificity and targets the DNA targeting system or fusion protein to a particular location of the genome, such as a target site specified by a component of the DNA-targeting domain.
- an exemplary DNA- targeting domain comprises a Clustered Regularly Interspaced Short Palindromic Repeats associated (Cas)-guide RNA (gRNA) combination that includes (a) a Cas protein or a variant thereof and (b) at least one gRNA; a zinc finger protein (ZFP) such as an eZFP; a transcription activator-like effector (TALE); a meganuclease; a homing endonuclease; or a I-Scel enzymes or a variant of any of the foregoing.
- the DNA-targeting domain comprises a catalytically inactive variant of any of the foregoing.
- the DNA-targeting domain comprises a Cas-gRNA combination that includes (a) a Cas protein or a variant thereof and (b) at least one gRNA.
- the variant Cas protein lacks nuclease activity or is a deactivated Cas (dCas) protein.
- the gRNA component (such as any described herein) provides the sequence specificity to target the DNA-targeting system, DNA-targeting domain or fusion protein to a target site specified by the gRNA.
- the DNA- targeting domain comprises an eZFP, such as any described herein.
- the DNA-targeting domain comprises a zinc finger protein (ZFP); a transcription activator-like effector (TALE); a meganuclease; a homing endonuclease; or an I-Scel enzyme or a variant thereof.
- ZFP zinc finger protein
- TALE transcription activator-like effector
- the DNA-targeting domain comprises a catalytically inactive variant of any of the foregoing.
- types of DNA- targeting domains include domains from proteins that can recognize nucleic acid sequences (e.g., target site) in a sequence-specific manner.
- a “zinc finger DNA binding protein” (or binding domain) is a protein, or a domain within a larger protein, that binds DNA in a sequence-specific manner through one or more zinc fingers, which are regions of amino acid sequence within the binding domain whose structure is stabilized through coordination of a zinc ion.
- the term zinc finger DNA binding protein is often abbreviated as zinc finger protein or ZFP.
- ZFPs are artificial, or engineered, ZFPs, comprising ZFP domains targeting specific DNA sequences, typically 9-18 nucleotides long, generated by assembly of individual fingers.
- ZFPs include those in which a single finger domain is approximately 30 amino acids in length and contains an alpha helix containing two invariant histidine residues coordinated through zinc with two cysteines of a single beta turn, and having two, three, four, five, or six fingers.
- sequence-specificity of a ZFP may be altered by making amino acid substitutions at the four helix positions (-1, 2, 3, and 6) on a zinc finger recognition helix.
- the ZFP or ZFP-containing molecule is non-naturally occurring, e.g., is engineered to bind to a target site of choice.
- the DNA-targeting system is or comprises a zinc-finger DNA binding domain fused to an effector domain.
- zinc fingers are custom-designed (i.e. designed by the user), or obtained from a commercial source.
- Various methods for designing zinc finger proteins are available. For example, methods for designing zinc finger proteins to bind to a target DNA sequence of interest are described, for example in Liu, Q. et al., PNAS, 94(l l):5525-30 (1997); Wright, D.A. et al., Nat. Protoc., l(3):1637-52 (2006); Gersbach, C.A. et al., Acc. Chem.
- the DNA-targeting domain is a domain from Transcription activator-like effectors (TALEs).
- TALEs are proteins found in Xanthomonas bacteria. TALEs comprise a plurality of repeated amino acid sequences, each repeat having binding specificity for one base in a target sequence. Each repeat comprises a pair of variable residues in position 12 and 13 (repeat variable diresidue; RVD) that determine the nucleotide specificity of the repeat.
- RVDs associated with recognition of the different nucleotides are HD for recognizing C, NG for recognizing T, NI for recognizing A, NN for recognizing G or A, NS for recognizing A, C, G or T, HG for recognizing T, IG for recognizing T, NK for recognizing G, HA for recognizing C, ND for recognizing C, HI for recognizing C, HN for recognizing G, NA for recognizing G, SN for recognizing G or A and YG for recognizing T, TL for recognizing A, VT for recognizing A or G and SW for recognizing A.
- RVDs can be mutated towards other amino acid residues in order to modulate their specificity towards nucleotides A, T, C and G and in particular to enhance this specificity.
- Binding domains with similar modular base-per-base nucleic acid binding properties can also be derived from different bacterial species. These alternative modular proteins may exhibit more sequence variability than TALE repeats.
- a “TALE DNA binding domain” or “TALE” is a polypeptide comprising one or more TALE repeat domains/units.
- the repeat domains each comprising a repeat variable diresidue (RVD), are involved in binding of the TALE to its cognate target DNA sequence.
- a single “repeat unit” (also referred to as a “repeat”) is typically 33-35 amino acids in length and exhibits at least some sequence homology with other TALE repeat sequences within a TALE protein.
- TALE proteins may be designed to bind to a target site using canonical or non-canonical RVDs within the repeat units. See, e.g., U.S. Pat. Nos. 8,586,526 and 9,458,205.
- a TALE is a fusion protein comprising a nucleic acid binding domain derived from a TALE and an effector domain.
- one or more sites in the FXN locus can be targeted by engineered TALEs.
- Zinc finger and TALE DNA-binding domains can be engineered to bind to a predetermined nucleotide sequence, for example via engineering (altering one or more amino acids) of the recognition helix region of a zinc finger protein, by engineering of the amino acids in a TALE repeat involved in DNA binding (the repeat variable diresidue or RVD region), or by systematic ordering of modular DNA- binding domains, such as TALE repeats or ZFP domains.
- engineered zinc finger proteins or TALE proteins are proteins that are non-naturally occurring.
- methods for engineering zinc finger proteins and TALEs are design and selection.
- a designed protein is a protein not occurring in nature whose design/composition results principally from rational criteria. Rational criteria for design include application of substitution rules and computerized algorithms for processing information in a database storing information of existing ZFP or TALE designs (canonical and non- canonical RVDs) and binding data. See, for example, U.S. Pat. Nos.
- the DNA-targeting system comprises at least one effector domain, such as any epigenetic effector domain provided herein, for example as described in Section II.B.
- the DNA-targeting domain or a component thereof is fused to the at least one effector domain.
- a DNA-targeting system comprising a fusion protein comprising: (a) a DNA-targeting domain targeting, or capable of being targeted to, a target site at a FXN locus or a regulatory element thereof, such as any described herein, and (b) at least one effector domain.
- the effector domain leads to an increase in transcription of FXN, or is capable of increasing transcription of FXN.
- the effector domain comprises a transcription activation domain.
- the effector domain comprises a multipartite activator.
- the DNA-targeting domain comprises a Cas-gRNA combination comprising (a) a Cas protein or a variant thereof and (b) at least one gRNA, and the component thereof fused to the at least one effector domain is the Cas protein or a variant thereof.
- the effector domain activates, induces, catalyzes, or leads to demethylation and/or increased transcription of FXN when ectopically recruited to FXN or a DNA regulatory element thereof.
- Exemplary fusion of DNA-targeting domain and at least one effector domain include fusing dCas9 with transcriptional activators such as VP64 (a polypeptide composed of four tandem copies of VP 16, a 16 amino acid transactivation domain of the Herpes simplex virus) can result in robust induction of gene expression.
- the effector domain activates, induces, catalyzes, or leads to demethylation and/or increased transcription of FXN when ectopically recruited to FXN or a DNA regulatory element thereof.
- the effector domain induces, catalyzes or leads to transcription activation, transcription co-activation, transcription elongation, transcription de -repression, histone modification, nucleosome remodeling, chromatin remodeling, reversal of heterochromatin formation, DNA demethylation, or DNA base oxidation.
- the effector domain induces, catalyzes or leads to transcription de-repression, DNA demethylation or DNA base oxidation.
- the effector domain induces transcription de -repression. In some embodiments, the effector domain induces transcription activation. In some embodiments, the effector domain has one of the aforementioned activities itself (i.e. acts directly). In some embodiments, the effector domain recruits and/or interacts with a polypeptide domain that has one of the aforementioned activities (i.e. acts indirectly).
- the effector domain induces, catalyzes or leads to transcription activation, transcription co-activation, transcription elongation, transcription de -repression, transcription factor release, polymerization, histone modification, histone acetylation, histone deacetylation, nucleosome remodeling, chromatin remodeling, reversal of heterochromatin formation, nuclease, signal transduction, proteolysis, ubiquitination, deubiquitination, phosphorylation, dephosphorylation, splicing, nucleic acid association, DNA methylation, DNA demethylation, histone methylation, histone demethylation, or DNA base oxidation.
- the effector domain induces, catalyzes or leads to transcription activation, transcription co-activation, or transcription elongation. In some embodiments, the effector domain induces transcription activation. In some embodiments, the effector domain activates transcription from one or more regulatory elements (e.g., promoters and/or enhancers) from the target locus, e.g., FXN. In some embodiments, the effector domain induces transcription activation. In some embodiments, the effector domain has one of the aforementioned activities itself (i.e. acts or catalyzes directly). In some embodiments, the effector domain recruits and/or interacts with another cellular component (e.g., transcription factor) that has one of the aforementioned activities (i.e. acts or catalyzes indirectly).
- a regulatory elements e.g., promoters and/or enhancers
- the effector domain induces transcription activation.
- the effector domain has one of the aforementioned activities itself (i.e. acts or cata
- gene expression of endogenous mammalian genes can be achieved by targeting a fusion protein comprising a DNA-targeting domain, such as a dCas9, and an effector domain, such as a transcription activation domain, to mammalian genes or regulatory DNA elements thereof (e.g. a promoter or enhancer), e.g. via one or more gRNAs.
- a DNA-targeting domain such as a dCas9
- an effector domain such as a transcription activation domain
- Transcription activation domains as well as activation of target genes by Cas fusion proteins (with a variety of Cas molecules) and the transcription activation domains, are described, for example, in WO 2014/197748, WO 2016/130600, WO 2017/180915, WO 2021/226555, WO 2021/226077, WO 2013/176772, WO 2014/152432, WO 2014/093661, Adli, M. Nat. Commun. 9, 1911 (2018), Perez-Pinera et al. Nat. Methods 10, 973-976 (2013), Mali et al. Nat. Biotechnol. 31, 833-838 (2013), and Maeder et al. Nat. Methods 10, 977-979 (2013).
- the effector domain comprises a transcriptional activator domain described in WO 2021/226077.
- activation or increase in gene expression of FXN is achieved by targeting a fusion protein comprising a DNA-targeting domain, such as a dCas9, and an effector domain, such as a transcription activation domain, to a FXN locus or regulatory DNA elements thereof (e.g. a promoter or enhancer) via one or more gRNAs.
- the one or more target sites of the one or more gRNA is at a FXN locus or regulatory DNA elements thereof (e.g., a promoter or enhancer), for example, as described herein, for example, in Section II.
- Any of a variety of effector domains for transcriptional activation are known and can be used in accord with the provided embodiments as described herein, for example, in Section II.B.
- the effector domain may have transcription activation activity, i.e., a transactivation domain.
- gene expression of endogenous mammalian genes may be achieved by targeting a fusion protein of iCas9 and a transactivation domain to mammalian promoters via combinations of gRNAs.
- the transactivation domain may include a VP16 protein, multiple VP16 proteins, such as a VP48 domain or VP64 domain, or p65 domain of NF kappa B transcription activator activity.
- the fusion protein may be iCas9-VP64.
- the effector domain may comprise a VP64 domain.
- dCas9-VP64 can be targeted to a target site by one or more gRNAs to activate a gene.
- VP64 is a polypeptide composed of four tandem copies of VP16, a 16 amino acid transactivation domain of the Herpes simplex virus.
- VP64 domains, including in dCas fusion proteins, have been described, for example, in WO 2014/197748, WO 2013/176772, WO 2014/152432, and WO 2014/093661.
- the effector domain comprises at least one VP16 domain, or a VP16 tetramer (“VP64”) or a variant thereof. In some embodiments, the effector domain comprises at least one VP16 domain, or a VP16 tetramer (“VP64”) or a variant thereof. In some embodiments, the effector domain comprises the sequence set forth in SEQ ID NO:81 or 83, a domain thereof, a portion thereof, or a variant thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing. An exemplary VP64 domain is set forth in SEQ ID NO:81.
- An exemplary nucleotide sequence encoding the exemplary VP64 domain set forth in SEQ ID NO: 81 is set forth in SEQ ID NO: 80.
- the effector domain comprises the sequence set forth in SEQ ID NO: 81, a domain thereof, a portion thereof, or a variant thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
- An exemplary VP64 domain is set forth in SEQ ID NO:83.
- An exemplary nucleotide sequence encoding the exemplary VP64 domain set forth in SEQ ID NO: 83 is set forth in SEQ ID NO: 82.
- the effector domain comprises the sequence set forth in SEQ ID NO:83, a domain thereof, a portion thereof, or a variant thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
- the provided compositions can target one or more target sites in a FXN locus.
- the target site comprises a specific sequence of nucleotides, such as DNA nucleotides.
- the target site is a DNA regulatory element of the FXN locus, such as a promoter or enhancer.
- compositions, methods and uses such as eZFPs, eZFP fusion proteins, DNA-targeting systems, DNA-targeting domains, components of the DNA-targeting domains, such as at least one gRNA, fusion proteins, and pluralities and combinations thereof, polynucleotides, vectors, cells, and pluralities and combinations thereof, that encode or comprise the DNA-targeting systems, fusion proteins, gRNAs, or pluralities or combinations thereof, that can target one or more particular genomic locations related to the FXN locus, such as a regulatory DNA element of the FXN locus.
- the target site is in a cell, such as any suitable cell.
- the cell is in or from any suitable organism, such as a human, mouse, dog, horse, rabbit, cattle, pig, hamster, gerbil, mouse, ferret, rat, cat, non-human primate, monkey, etc.
- the cell is in or from a human.
- the cell is any suitable cell, such as an immune cell (e.g. a T cell, B cell, or antigen-presenting cell), a liver cell (e.g. a hepatocyte), a cell of a nervous system (e.g. a neuron or glial cell), a heart cell (e.g. a cardiomyocyte) or a stem cell (e.g. an embryonic stem cell or induced pluripotent stem cell).
- an immune cell e.g. a T cell, B cell, or antigen-presenting cell
- a liver cell e.g. a hepatocyte
- a cell of a nervous system e.
- the target site is located in a regulatory DNA element of a frataxin (FXN) locus.
- the target site is located within a promoter, upstream regulatory element (e.g., enhancer), exon, intron, 5’ untranslated region (UTR), 3’ UTR, or downstream regulatory element.
- the target site is located within a FXN locus.
- the target site is located within a regulatory DNA element (e.g. a cis-, trans-, distal, proximal, upstream, or downstream regulatory DNA element) of a FXN locus.
- the target site is located within a promoter, enhancer, exon, intron, untranslated region (UTR), 5’ UTR or 3’ UTR.
- the target site is located within a sequence and/or sequences of unknown or known function that are suspected of being able to control expression of FXN.
- one or more target sites such as one or more target sites located within a regulatory DNA element (e.g. a cis-, trans-, distal, proximal, upstream, or downstream regulatory DNA element) of a FXN locus.
- the target site is located within a promoter, enhancer, exon, intron, untranslated region (UTR), 5’ UTR or 3’ UTR are targeted.
- an exemplary frataxin (FXN) transcript is set forth in RefSeq NM_000144) (transcript variant 1); Gencode Transcript: ENST00000484259.3; Gencode Gene: ENSG00000165060.15.
- Genomic coordinates for an exemplary transcript (including UTRs) for FXN include hg38 chr9:69, 035, 752-69, 079, 076 (Size: 43,325 Total Exon Count: 5 Strand: +).
- Genomic coordinates for the coding region for this transcript variant include hg38 chr9:69, 035, 783-69, 072, 762 (Size: 36,980 Coding Exon Count: 5).
- an exemplary frataxin (FXN) transcript is set forth in RefSeq NM_181425) (transcript variant 2); Gencode Transcript: ENST00000396366.6; Gencode Gene: ENSG00000165060.15.
- Genomic coordinates for an exemplary transcript (including UTRs) for FXN include hg38 chr9:69, 035, 762-69, 073, 022 (Size: 37,261 Total Exon Count: 5 Strand: +).
- Genomic coordinates for the coding region for this transcript variant include hg38 chr9:69, 035, 783-69, 072, 712 (Size: 36,930 Coding Exon Count: 5).
- the regulatory DNA element is located in a genomic region comprising the FXN locus.
- the target site is located within the genomic coordinates hg38 chr9:68,940, 179-69,205,519. In some embodiments, the target site is selected from the sequence set forth in any one of SEQ ID NOS:1-21, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing.
- the regulatory DNA element is an enhancer.
- the target site is located within the genomic coordinates human genome assembly GRCh38 (hg38) chr9:69, 027, 282-69, 028, 497. In some embodiments, the target site is located within the genomic coordinates hg38 chr9:69, 027, 615-69, 028, 101. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO: 21, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing.
- the target site comprises a sequence set forth in any one of SEQ ID NOS:229-243, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises a sequence set forth in any one of SEQ ID NOS:256-265, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing.
- the target site is located within the genomic coordinates hg38 chr9:69, 044, 201-69, 045, 347. In some embodiments, the target site is located within the genomic coordinates hg38 chr9:69, 030, 752-69, 031,507. In some embodiments, the target site is located within the genomic coordinates hg38 chr9:68, 999, 262-69, 000, 023. In some embodiments, the target site is located within the genomic coordinates hg38 chr9:69, 085, 468-69, 086, 426.
- the target site is located within the genomic coordinates hg38 chr9:69, 096, 701-69, 097, 567. In some embodiments, the target site is located within the genomic coordinates hg38 chr9:69, 120, 690-69, 123 ,549. In some embodiments, the target site is located within the genomic coordinates hg38 chr9:69, 130, 392-69, 132, 484.
- the DNA-targeting domain comprises a Cas-gRNA combination comprising (a) a Cas protein or a variant thereof and (b) at least one gRNA; and the gRNA comprises at least one gRNA spacer sequence comprising the sequence set forth in SEQ ID NO:42, or a contiguous portion thereof of at least 14 nt.
- the gRNA further comprises the sequence set forth in SEQ ID NO:44.
- the gRNA comprises the sequence set forth in SEQ ID NO:67.
- the regulatory DNA element is a promoter
- the target site is located within the genomic coordinates hg38 chr9:69, 034, 622-69, 036, 670. In some embodiments, the target site is located within the genomic coordinates hg38 chr9:69, 035, 300-69, 035, 900. In some embodiments, the target site is located within the genomic coordinates hg38 chr9:69, 034, 900-69, 035, 900. In some embodiments, the target site comprises a sequence selected from any one of SEQ ID NOS: 1-10, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing.
- the target site comprises a sequence selected from any one of SEQ ID NOS: 11-20, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises a sequence selected from any one of SEQ ID NOS:244-255, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing.
- the target site is at, near, or within a FXN locus.
- the target site is a sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to all or a portion of the target site sequence described herein.
- the target site is a sequence having at least 80% sequence identity to all or a portion of the target site sequence described herein.
- the target site is a sequence having at least 85% sequence identity to all or a portion of the target site sequence described herein.
- the target site is a sequence having at least 90% sequence identity to all or a portion of the target site sequence described herein. In some aspects, the target site is a sequence having at least 91% sequence identity to all or a portion of the target site sequence described herein. In some aspects, the target site is a sequence having at least 92% sequence identity to all or a portion of the target site sequence described herein. In some aspects, the target site is a sequence having at least 93% sequence identity to all or a portion of the target site sequence described herein. In some aspects, the target site is a sequence having at least 94% sequence identity to all or a portion of the target site sequence described herein.
- the target site is a sequence having at least 95% sequence identity to all or a portion of the target site sequence described herein. In some aspects, the target site is a sequence having at least 96% sequence identity to all or a portion of the target site sequence described herein. In some aspects, the target site is a sequence having at least 97% sequence identity to all or a portion of the target site sequence described herein. In some aspects, the target site is a sequence having at least 98% sequence identity to all or a portion of the target site sequence described herein. In some aspects, the target site is a sequence having at least 99% sequence identity to all or a portion of the target site sequence described herein.
- the target site is a sequence having at least 99.5% sequence identity to all or a portion of the target site sequence described herein. In some aspects, the target site is a sequence having at least 99.9% sequence identity to all or a portion of the target site sequence described herein. In some aspects, the target site is a sequence having 100% sequence identity to all or a portion of the target site sequence described herein.
- the target site is selected from the sequence set forth in any one of SEQ ID NOS:1-21, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing.
- the target site comprises SEQ ID NO:1, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:2, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:3, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:4, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof.
- the target site comprises SEQ ID NO:5, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:6, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:7, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:8, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:9, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof.
- the target site comprises SEQ ID NO: 10, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:11, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO: 12, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO: 13, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO: 14, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof.
- the target site comprises SEQ ID NO: 15, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO: 16, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO: 17, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO: 18, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO: 19, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof.
- the target site comprises SEQ ID NO:20, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:21, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof.
- the target site comprises SEQ ID NO:229, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:230, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:231, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:232, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof.
- the target site comprises SEQ ID NO:233, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:234, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:235, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:236, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof.
- the target site comprises SEQ ID NO:237, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:238, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:239, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:240, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof.
- the target site comprises SEQ ID NO:241, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:242, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:243, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:244, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof.
- the target site comprises SEQ ID NO:245, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:246, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:247, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:248, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof.
- the target site comprises SEQ ID NO:249, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:250, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:251, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:252, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof.
- the target site comprises SEQ ID NO:253, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:254, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:255, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:256, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof.
- the target site comprises SEQ ID NO:257, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:258, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:259, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:260, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof.
- the target site comprises SEQ ID NO:261, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:262, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:263, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:264, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:265, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof.
- the target site comprises SEQ ID NO:1. In some embodiments, the target site comprises SEQ ID NO:2. In some embodiments, the target site comprises SEQ ID NO:3. In some embodiments, the target site comprises SEQ ID NO:4. In some embodiments, the target site comprises SEQ ID NO:5. In some embodiments, the target site comprises SEQ ID NO:6. In some embodiments, the target site comprises SEQ ID NO:7. In some embodiments, the target site comprises SEQ ID NO:8. In some embodiments, the target site comprises SEQ ID NO:9. In some embodiments, the target site comprises SEQ ID NO: 10. In some embodiments, the target site comprises SEQ ID NO: 11. In some embodiments, the target site comprises SEQ ID NO: 12.
- the target site comprises SEQ ID NO: 13. In some embodiments, the target site comprises SEQ ID NO: 14. In some embodiments, the target site comprises SEQ ID NO: 15. In some embodiments, the target site comprises SEQ ID NO: 16. In some embodiments, the target site comprises SEQ ID NO: 17. In some embodiments, the target site comprises SEQ ID NO: 18. In some embodiments, the target site comprises SEQ ID NO: 19. In some embodiments, the target site comprises SEQ ID NO:20. In some embodiments, the target site comprises SEQ ID NO:21.
- the target site comprises SEQ ID NO: 229. In some embodiments, the target site comprises SEQ ID NO:230. In some embodiments, the target site comprises SEQ ID NO:231. In some embodiments, the target site comprises SEQ ID NO:232. In some embodiments, the target site comprises SEQ ID NO:233. In some embodiments, the target site comprises SEQ ID NO:234. In some embodiments, the target site comprises SEQ ID NO: 235. In some embodiments, the target site comprises SEQ ID NO:236. In some embodiments, the target site comprises SEQ ID NO:237. In some embodiments, the target site comprises SEQ ID NO: 238. In some embodiments, the target site comprises SEQ ID NO:239.
- the target site comprises SEQ ID NO:240. In some embodiments, the target site comprises SEQ ID NO: 241. In some embodiments, the target site comprises SEQ ID NO:242. In some embodiments, the target site comprises SEQ ID NO:243. In some embodiments, the target site comprises SEQ ID NO:244. In some embodiments, the target site comprises SEQ ID NO:245. In some embodiments, the target site comprises SEQ ID NO:246. In some embodiments, the target site comprises SEQ ID NO: 247. In some embodiments, the target site comprises SEQ ID NO:248. In some embodiments, the target site comprises SEQ ID NO:249. In some embodiments, the target site comprises SEQ ID NO:250.
- the target site comprises SEQ ID NO:251. In some embodiments, the target site comprises SEQ ID NO:252. In some embodiments, the target site comprises SEQ ID NO: 253. In some embodiments, the target site comprises SEQ ID NO:254. In some embodiments, the target site comprises SEQ ID NO:255. In some embodiments, the target site comprises SEQ ID NO:256. In some embodiments, the target site comprises SEQ ID NO:257. In some embodiments, the target site comprises SEQ ID NO:258. In some embodiments, the target site comprises SEQ ID NO: 259. In some embodiments, the target site comprises SEQ ID NO:260. In some embodiments, the target site comprises SEQ ID NO:261.
- the target site comprises SEQ ID NO:262. In some embodiments, the target site comprises SEQ ID NO:263. In some embodiments, the target site comprises SEQ ID NO:264. In some embodiments, the target site comprises SEQ ID NO: 265.
- the target site comprises a complementary sequence of SEQ ID NO:1. In some embodiments, the target site comprises a complementary sequence of SEQ ID NO:2. In some embodiments, the target site comprises a complementary sequence of SEQ ID NO:3. In some embodiments, the target site comprises a complementary sequence of SEQ ID NO:4. In some embodiments, the target site comprises a complementary sequence of SEQ ID NO:5. In some embodiments, the target site comprises a complementary sequence of SEQ ID NO:6. In some embodiments, the target site comprises a complementary sequence of SEQ ID NO:7. In some embodiments, the target site comprises a complementary sequence of SEQ ID NO:8. In some embodiments, the target site comprises a complementary sequence of SEQ ID NO:9.
- the target site comprises a complementary sequence of SEQ ID NO: 10. In some embodiments, the target site comprises a complementary sequence of SEQ ID NO: 11. In some embodiments, the target site comprises a complementary sequence of SEQ ID NO: 12. In some embodiments, the target site comprises a complementary sequence of SEQ ID NO: 13. In some embodiments, the target site comprises a complementary sequence of SEQ ID NO: 14. In some embodiments, the target site comprises a complementary sequence of SEQ ID NO: 15. In some embodiments, the target site comprises a complementary sequence of SEQ ID NO: 16. In some embodiments, the target site comprises a complementary sequence of SEQ ID NO: 17. In some embodiments, the target site comprises a complementary sequence of SEQ ID NO: 18. In some embodiments, the target site comprises a complementary sequence of SEQ ID NO: 19. In some embodiments, the target site comprises a complementary sequence of SEQ ID NO:20. In some embodiments, the target site comprises a complementary sequence of SEQ ID NO:21.
- the target site comprises the sequence set forth in SEQ ID NO:1, SEQ ID N0:7, or SEQ ID NO:21, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO:1, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO:7, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing.
- the target site comprises the sequence set forth in SEQ ID NO:21, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO:1. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO:7. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO:21. In some embodiments, the target site comprises a complementary sequence of the sequence set forth in SEQ ID NO:1. In some embodiments, the target site comprises a complementary sequence of the sequence set forth in SEQ ID NO: 7. In some embodiments, the target site comprises a complementary sequence of the sequence set forth in SEQ ID NO:21.
- the target site is a target site for an eZFP, such as any eZFP provided herein, for example in Section I, and/or a composition comprising the eZFP, such as an eZFP fusion protein.
- the target site is selected from the sequence set forth in any one of SEQ ID NOS:269-300 and 583-600, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing.
- the target site is in a FXN promoter. In some embodiments, the target site is within a target region spanning the genomic coordinates chr9:69, 034, 900-69, 035, 900 from hg38 (SEQ ID NO:430). In some embodiments, the target site is within a target region spanning the genomic coordinates chr9:69, 035, 300-69-035, 800 from hg38. In some embodiments, the target site is within a target region spanning the genomic coordinates chr9:69, 035, 350-69, 035, 450 from hg38.
- the target site is within a target region spanning the genomic coordinates chr9:69,035,400- 69,035,450 from hg38. In some embodiments, the target site is within a target region spanning the genomic coordinates chr9:69, 035, 530-69, 035, 580 from hg38. In some embodiments, the target site is within a target region spanning the genomic coordinates chr9:69, 035, 675-69, 035, 725 from hg38.
- the target site is in a FXN enhancer. In some embodiments, the target site is within a target region spanning the genomic coordinates chr9:69, 027, 282-69, 028, 497 from hg38 (SEQ ID NO:431). In some embodiments, the target site is within a target region spanning the genomic coordinates chr9:69, 027, 615-69, 028, 101 from hg38. In some embodiments, the target site is within a target region spanning the genomic coordinates chr9:69, 027, 775-69, 027, 875 from hg38. In some embodiments, the target site is within a target region spanning the genomic coordinates chr9:69,027,795- 69,027,845 from hg38.
- the target site comprises the nucleotide sequence set forth in any one of SEQ ID NOS:269-300 and 583-600, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the target site comprises the nucleotide sequence set forth in any one of SEQ ID NOS:269-300 and 583-600.
- the target site is comprised in double-stranded DNA, such as genomic DNA.
- the target site is double-stranded DNA, such as genomic DNA.
- the target site comprises a sequence set forth in Table 1.
- the target site comprises the nucleotide sequence set forth in SEQ ID NO:272, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO: 272. In some embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:277, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO: 277.
- the target site comprises the nucleotide sequence set forth in SEQ ID NO:280, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO: 280. In some embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:281, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO: 281.
- the target site comprises the nucleotide sequence set forth in SEQ ID NO:283, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO: 283. In some embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:290, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO: 290.
- the target site comprises the nucleotide sequence set forth in SEQ ID NO:299, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO: 299. In some embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:583, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO:583.
- the target site comprises the nucleotide sequence set forth in SEQ ID NO:584, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO:584. In some embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:585, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO:585.
- the target site comprises the nucleotide sequence set forth in SEQ ID NO:586, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO:586. In some embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:587, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO:587.
- the target site comprises the nucleotide sequence set forth in SEQ ID NO:588, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO:588. In some embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:589, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO:589.
- the target site comprises the nucleotide sequence set forth in SEQ ID NO:590, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO:590. In some embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:591, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO:591.
- the target site comprises the nucleotide sequence set forth in SEQ ID NO:592, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO:592. In some embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:593, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO:593.
- the target site comprises the nucleotide sequence set forth in SEQ ID NO:594, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO:594. In some embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:595, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO:595.
- the target site comprises the nucleotide sequence set forth in SEQ ID NO:596, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO:596. In some embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:597, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO:597.
- the target site comprises the nucleotide sequence set forth in SEQ ID NO:598, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO:598. In some embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:599, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO:599.
- the target site comprises the nucleotide sequence set forth in SEQ ID NO:600, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO:600.
- the DNA-targeting systems provided herein such as CRISPR/Cas-based and ZFP-based DNA-targeting systems, comprise epigenetic effector domains for targeted transcriptional activation.
- fusion proteins of the DNA-targeting systems such as eZFP fusion proteins or dCas fusion proteins, comprise the epigenetic effector domains.
- multipartite effectors comprising two or more effector domains.
- the DNA-targeting systems such as CRISPR/Cas-based and ZFP-based DNA-targeting systems, comprise the multipartite effectors.
- fusion proteins of the DNA-targeting systems such as eZFP fusion proteins or dCas fusion proteins, comprise the multipartite effectors.
- epigenetic effector domains such as transcriptional activation domains (i.e. effectors for transcriptional activation) and multipartite effectors for transcriptional activation (i.e. multipartite activators).
- the effector domains are provided in fusion proteins (such as eZFP fusion proteins or dCas fusion proteins), and DNA-targeting systems, such as any of the DNA-targeting systems provided herein.
- the transcriptional activation domains facilitate increased expression of FXN, for example when targeted to a FXN locus by an eZFP, eZFP fusion protein, or DNA-targeting system, such as a dCas-based DNA-targeting system.
- fusion proteins, effector proteins and/or DNA-targeting systems that contain two or more of the transcriptional activation domains.
- multipartite effectors for transcriptional activation e.g., multipartite activators, comprising two or more effector domains such as transcriptional activation domains, such as any provided herein.
- the transcriptional activation domains and multipartite activators increase, or are capable of increasing, transcription of an endogenous locus, such as FXN, when recruited to a target site at the endogenous locus, for example increasing transcription of FXN when recruited to a target site for FXN provided herein.
- the transcriptional activation domains and multipartite activators are provided as part of a fusion protein or DNA-targeting system, such as any described herein.
- the transcriptional activation domains and multipartite activators are targeted to FXN to activate, induce, catalyze, or lead to increased transcription of the FXN gene.
- the transcriptional activation domains and multipartite activators are targeted to the target site via a DNA-targeting domain, such as a CRISPR/Cas-based, ZFN- based, or TAEE-based DNA-targeting domain, including any of the DNA-targeting domains described herein, for example, in Section I and II.
- the transcriptional activation domains and/or multipartite effectors are targeted to the target site via an eZFP, such as as eZFP comprised in an eZFP fusion protein that also comprises the transcriptional activation domain and/or multipartite effector.
- a transcriptional activation domain increases transcription of an endogenous locus, such as a FXN locus, when recruited to a target site at the endogenous locus.
- the transcriptional activation domain is a domain that induces, catalyzes, or leads to increased transcription of a gene, such as FXN, when ectopically recruited to the gene or a DNA regulatory element thereof.
- the transcriptional activation domain activates, induces, catalyzes, or leads to transcription activation, transcription co-activation, transcription elongation, or transcription de-repression.
- the transcriptional activation domain induces transcriptional activation.
- the transcriptional activation domain has one of the aforementioned activities itself (i.e., acts directly).
- the effector domain recruits and/or interacts with a polypeptide domain that has one of the aforementioned activities (i.e., acts indirectly).
- activation of gene expression of endogenous genes can be achieved by targeting (e.g., via a CRISPR-based, ZFN-based, or TALE-based DNA-targeting domain) of transcriptional activation domains to a target site for the genes, such as regulatory DNA elements thereof (e.g., a promoter or enhancer).
- a transcriptional activation domain provided herein comprises a domain from a human protein.
- a transcriptional activation domain from a protein comprises any portion of the protein that is capable of acting as a transcriptional activation domain as described herein.
- a transcription activation domain is or comprises a portion, fragment, domain or variant of a human protein, such as a portion, fragment, domain or variant of a human protein selected from among DPOLA, ENL, FOXO3, HSH2D, NCOA2, NCOA3, PSA1, PYGO1, RBM39, HERC2, and NOTCH2, that exhibits transcriptional activation, is capable of inducing or activating transcription from a gene), is a functional transcriptional activation domain, and/or has a function of transcription activation.
- a human protein such as a portion, fragment, domain or variant of a human protein selected from among DPOLA, ENL, FOXO3, HSH2D, NCOA2, NCOA3, PSA1, PYGO1, RBM39, HERC2, and NOTCH2, that exhibits transcriptional activation, is capable of inducing or activating transcription from a gene), is a functional transcriptional activation domain, and/or has a function of transcription activation.
- a transcription activation domain is or comprises a functional portion, a functional fragment, a functional domain or a functional variant of a human protein, such as a portion, fragment, domain or variant of a human protein selected from among DPOLA, ENL, FOXO3, HSH2D, NCOA2, NCOA3, PSA1, PYGO1, RBM39, HERC2, and NOTCH2, that exhibits transcriptional activation, is capable of inducing or activating transcription from a gene), is a functional transcriptional activation domain, and/or has a function of transcription activation.
- a transcription activation domain is or comprises a partially or fully functional portion, a partially or fully functional fragment, a partially or fully functional domain or a partially or fully functional variant of a human protein, such as a portion, fragment, domain or variant of a human protein selected from among DPOLA, ENL, FOXO3, HSH2D, NCOA2, NCOA3, PSA1, PYGO1, RBM39, HERC2, and N0TCH2, that exhibits increases the transcription from a gene by at least 5%, 10%, 20%, 30%, 40%' or 50% 60%, 70%, 80%, 85%, 90%-, or 100% or more, such as 2-fold, 5-fold, 10-fold, 20- fold, 30-fold, 40-fold, 50-fold. 60-fold, 70-fold, 80-fold, 90-fold. 100-fold, 200-fold, 300-fold, 400-food, 500-fold, 1000-fold or more, compared to the absence of the transcriptional activation domain.
- the transcriptional activation domain is 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids in length, or within a range defined by any of the foregoing. In some embodiments, the transcriptional activation domain is at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids in length.
- the transcriptional activation domain is 10, 15, 20, 25, 30, 35, 40, 45, 50, 55, 60, 65, 70, 75, or 80 amino acids in length, or within a range defined by any of the foregoing. In some embodiments, the transcriptional activation domain is at least 10, 15, 20, 25, 30, 35, 40, 45, 50, 55, 60, 65, 70, 75, or 80 amino acids in length, or within a range defined by any of the foregoing. In some embodiments, the transcriptional activation domain is 22, 37, 42, 47, 49, 57, 61, 62, 70, 72, 76, or 80 amino acids in length, or within a range defined by any of the foregoing.
- the transcriptional activation domain is at least 22, 37, 42, 47, 49, 57, 61, 62, 70, 72, 76, or 80 amino acids in length. In some embodiments, the transcriptional activation domain is between 10 and 80, 20 and 70, 30 and 80, 30 and 70, 30 and 60, 40 and 80, 40 and 70, 40 and 60, 40 and 50, 50 and 80, 50 and 70, 50 and 60 amino acids in length.
- the transcriptional activation domain comprises a transcriptional activation domain described in WO 2021/226077.
- a transcriptional activation domain comprises a domain from DPOLA, ENL, FOXO3, HSH2D, NCOA2, NCOA3, PSA1, PYGO1, RBM39, HERC2, or NOTCH2.
- a domain from a gene is referred to as a gene domain.
- a domain from DPOLA may be referred to as a DPOLA domain herein.
- the domain from DPOLA, ENL, FOXO3, HSH2D, NCOA2, NCOA3, PSA1, PYGO1, RBM39, HERC2, or NOTCH2 is or comprises the respective transcriptional activation domains described herein or a partially or fully functional fragment thereof, a domain thereof, or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 25, 30, 35, 40, 45, 50, 55, 60, 65, 70, 75, or 80 amino acids, such as at least 20 amino acids, or a variant thereof.
- the domain from DPOLA, ENL, FOXO3, HSH2D, NCOA2, NCOA3, PSA1, PYGO1, RBM39, HERC2, or NOTCH2 is or comprises the sequence of the respective transcriptional activation domains described herein or a partially or fully functional fragment thereof, a domain thereof, or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 25, 30, 35, 40, 45, 50, 55, 60, 65, 70, 75, or 80 amino acids, such as at least 20 amino acids, or a variant thereof.
- the transcriptional activation domain comprises or is selected from a transcriptional activation domain shown in Table 3, or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of transcriptional activation domain shown in Table 3, or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof
- the transcriptional activation domain comprises or is selected from a transcriptional activation domain shown in Table 3, or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof.
- the transcriptional activation domain comprises or is selected from a transcriptional activation domain shown in Table 3, or a domain or a portion thereof, such as a contiguous portion thereof of 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids in length, or within a range defined by any of the foregoing.
- the transcriptional activation domain comprises or is selected from a transcriptional activation domain shown in Table 3, or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids in length.
- the transcriptional activation domain comprises or is selected from a transcriptional activation domain shown in Table 3, or a domain or a portion thereof, such as a contiguous portion thereof of 10, 15, 20, 25, 30, 35, 40, 45, 50, 55, 60, 65, 70, 75, or 80 amino acids in length, or within a range defined by any of the foregoing.
- the transcriptional activation domain comprises or is selected from a transcriptional activation domain shown in Table 3, or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 25, 30, 35, 40, 45, 50, 55, 60, 65, 70, 75, or 80 amino acids in length, or within a range defined by any of the foregoing.
- the transcriptional activation domain comprises or is selected from a transcriptional activation domain shown in Table 3, or a domain or a portion thereof, such as a contiguous portion thereof of 22, 37, 42, 47, 49, 57, 61, 62, 70, 72, 76, or 80 amino acids in length, or within a range defined by any of the foregoing.
- the transcriptional activation domain comprises or is selected from a transcriptional activation domain shown in Table 3, or a domain or a portion thereof, such as a contiguous portion thereof of at least 22, 37, 42, 47, 49, 57, 61, 62, 70, 72, 76, or 80 amino acids in length.
- the transcriptional activation domain comprises or is selected from a transcriptional activation domain shown in Table 3, or a domain or a portion thereof, such as a contiguous portion thereof of between 10 and 80, 20 and 70, 30 and 80, 30 and 70, 30 and 60, 40 and 80, 40 and 70, 40 and 60, 40 and 50, 50 and 80, 50 and 70, 50 and 60 amino acids in length.
- the transcriptional activation domain is a transcriptional activation domain set forth in Table 3. Table 3 shows a list of human genes and exemplary transcriptional activation domains from each gene.
- any of the provided multipartite effector proteins, fusion proteins, and/or DNA targeting systems comprises a combination of transcriptional activation domains, such as a combination of two or more, such as three or more, such as three or more, of any of transcriptional activation domains shown in Table 3.
- any of the provided multipartite effector proteins, fusion proteins, and/or DNA targeting systems comprises a combination of two or more, such as three or more, of any one of the SEQ ID NOS: set forth in Table 3, or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any one of the SEQ ID NOS: set forth in Table 3, or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49,
- any of the provided multipartite effector proteins, fusion proteins, and/or DNA targeting systems comprises a combination of two or more, such as three or more, of any one of the SEQ ID NOS: set forth in Table 3, or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof.
- any of the provided multipartite effector proteins, fusion proteins, and/or DNA targeting systems, such as a multipartite activator comprises two or more, such as three or more, of any one of the SEQ ID NOS: set forth in Table 3.
- the transcriptional activation domain comprises any one of SEQ ID NOS: 113-122 and 124, or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any one of SEQ ID NOS: 113-122 and 124, or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least
- the transcriptional activation domain comprises any one of SEQ ID NOS:100-109, 111, and 383-393, or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any one of SEQ ID NOS:100-109, 111, and 383-393, or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76
- a transcriptional activation domain comprises a DPOLA domain, i.e. a domain from DPOLA.
- DPOLA refers to the DNA polymerase alpha catalytic subunit protein encoded by the human POLA1 gene.
- DPOLA plays an essential role in the initiation of DNA synthesis.
- An exemplary human DPOLA sequence is set forth in SEQ ID NO: 113.
- An exemplary DPOLA domain sequence is set forth in SEQ ID NO: 100 and SEQ ID NO:383.
- the transcriptional activation domain comprises a sequence set forth in any of SEQ ID NOS: 113, 100, and 383 or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to a sequence set forth in any of SEQ ID NOS: 113, 100, and 383 or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as
- the transcriptional activation domain is or comprises an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO: 100.
- the transcriptional activation domain comprises a contiguous portion of SEQ ID NO: 113 that is at least 80 amino acids in length.
- the transcriptional activation domain comprises SEQ ID NO: 100.
- the transcriptional activation domain is set forth in SEQ ID NO: 100.
- An exemplary nucleotide sequence encoding the transcriptional activation domain set forth in SEQ ID NO: 100 is set forth in SEQ ID NO:87.
- the transcriptional activation domain is or comprises an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO: 383.
- the transcriptional activation domain comprises a contiguous portion of SEQ ID NO: 113 that is at least 61 amino acids in length.
- the transcriptional activation domain comprises SEQ ID NO: 383.
- the transcriptional activation domain is set forth in SEQ ID NO:383.
- a transcriptional activation domain comprises a ENL domain, i.e. a domain from ENL.
- ENL refers to the ENL protein encoded by the human MLLT1 gene.
- ENL functions as a chromatin reader component of the super elongation complex (SEC), a complex which increases the catalytic rate of RNA polymerase II transcription.
- SEC super elongation complex
- An exemplary human ENL sequence is set forth in SEQ ID NO: 114.
- An exemplary ENL domain sequence is set forth in SEQ ID NO: 101 and SEQ ID NO:384.
- the transcriptional activation domain comprises a sequence set forth in any of SEQ ID NOS:114, 101, and 384 or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to a sequence set forth in any of SEQ ID NOS: 114, 101, and 384 or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids
- the transcriptional activation domain is or comprises an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO: 101.
- An exemplary nucleotide sequence encoding the transcriptional activation domain set forth in SEQ ID NO: 101 is set forth in SEQ ID NO:88.
- the transcriptional activation domain comprises a contiguous portion of SEQ ID NO: 114 that is at least 80 amino acids in length.
- the transcriptional activation domain comprises SEQ ID NO: 101.
- the transcriptional activation domain is set forth in SEQ ID NO: 101.
- the transcriptional activation domain is or comprises an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO:384.
- the transcriptional activation domain comprises a contiguous portion of SEQ ID NO: 114 that is at least 62 amino acids in length.
- the transcriptional activation domain comprises SEQ ID NO:384.
- the transcriptional activation domain is set forth in SEQ ID NO:384.
- a transcriptional activation domain comprises a FOXO3 domain, i.e. a domain from FOXO3.
- FOXO3 refers to the Forkhead box protein 03 encoded by the human FOXO3 gene.
- FOXO3 functions as a transcriptional activator that recognizes and binds to specific DNA sequences.
- An exemplary human FOXO3 sequence is set forth in SEQ ID NO: 115.
- An exemplary FOXO3 domain sequence is set forth in SEQ ID NO: 102 and SEQ ID NO:385.
- the transcriptional activation domain comprises a sequence set forth in any of SEQ ID NOS: 115, 102, and 385 or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to a sequence set forth in any of SEQ ID NOS: 115, 102, and 385 or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or
- the transcriptional activation domain is or comprises an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO: 102.
- the transcriptional activation domain comprises a contiguous portion of SEQ ID NO: 115 that is at least 80 amino acids in length.
- the transcriptional activation domain comprises SEQ ID NO: 102.
- the transcriptional activation domain is set forth in SEQ ID NO: 102.
- An exemplary nucleotide sequence encoding the transcriptional activation domain set forth in SEQ ID NO: 102 is set forth in SEQ ID NO: 88.
- the transcriptional activation domain is or comprises an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO:385.
- the transcriptional activation domain comprises a contiguous portion of SEQ ID NO: 115 that is at least 42 amino acids in length.
- the transcriptional activation domain comprises SEQ ID NO:385.
- the transcriptional activation domain is set forth in SEQ ID NO:385.
- a transcriptional activation domain comprises a HSH2D domain, i.e. a domain from HSH2D.
- HSH2D refers to the Hematopoietic SH2 domain-containing protein encoded by the human HSH2D gene.
- HSH2D functions as an adapter protein involved in tyrosine kinase and CD28 signaling.
- An exemplary human HSH2D sequence is set forth in SEQ ID NO: 116.
- An exemplary HSH2D domain sequence is set forth in SEQ ID NO: 103 and SEQ ID NO:386.
- the transcriptional activation domain comprises a sequence set forth in any of SEQ ID NOS: 116, 103, and 386 or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to a sequence set forth in any of SEQ ID NOS: 116, 103, and 386 or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or
- the transcriptional activation domain is or comprises an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO: 103.
- the transcriptional activation domain comprises a contiguous portion of SEQ ID NO: 116 that is at least 80 amino acids in length.
- the transcriptional activation domain comprises SEQ ID NO: 103.
- the transcriptional activation domain is set forth in SEQ ID NO: 103.
- An exemplary nucleotide sequence encoding the transcriptional activation domain set forth in SEQ ID NO: 103 is set forth in SEQ ID NO: 90.
- the transcriptional activation domain is or comprises an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO:386.
- the transcriptional activation domain comprises a contiguous portion of SEQ ID NO: 116 that is at least 76 amino acids in length.
- the transcriptional activation domain comprises SEQ ID NO:386.
- the transcriptional activation domain is set forth in SEQ ID NO:386.
- a transcriptional activation domain comprises a NCOA2 domain, i.e. a domain from NCOA2.
- NCOA2 refers to the Nuclear receptor coactivator 2 protein encoded by the human NCOA2 gene.
- NCOA2 functions as a transcriptional coactivator for steroid receptors and nuclear receptors.
- An exemplary human NCOA2 sequence is set forth in SEQ ID NO: 117.
- An exemplary NCOA2 domain sequence is set forth in SEQ ID NO: 104 and SEQ ID NO:387.
- the transcriptional activation domain comprises a sequence set forth in any of SEQ ID NOS: 117, 104, and 387 or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to a sequence set forth in any of SEQ ID NOS: 117, 104, and 387 or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or
- the transcriptional activation domain is or comprises an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO: 104.
- the transcriptional activation domain comprises a contiguous portion of SEQ ID NO: 117 that is at least 80 amino acids in length.
- the transcriptional activation domain comprises SEQ ID NO: 104.
- the transcriptional activation domain is set forth in SEQ ID NO: 104.
- An exemplary nucleotide sequence encoding the transcriptional activation domain set forth in SEQ ID NO: 104 is set forth in SEQ ID NO: 91.
- the transcriptional activation domain is or comprises an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO:387.
- the transcriptional activation domain comprises a contiguous portion of SEQ ID NO: 117 that is at least 47 amino acids in length.
- the transcriptional activation domain comprises SEQ ID NO:387.
- the transcriptional activation domain is set forth in SEQ ID NO:387.
- a transcriptional activation domain comprises a NCOA3 domain, i.e. a domain from NCOA3.
- NCOA3 refers to the Nuclear receptor coactivator 3 protein encoded by the human NCOA3 gene.
- NCOA3 functions as a transcriptional coactivator for steroid receptors and nuclear receptors.
- An exemplary human NCOA3 sequence is set forth in SEQ ID NO: 118.
- An exemplary NCOA3 domain sequence is set forth in SEQ ID NO: 105 and SEQ ID NO:388.
- the transcriptional activation domain comprises a sequence set forth in any of SEQ ID NOS: 118, 105, and 388 or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to a sequence set forth in any of SEQ ID NOS: 118, 105, and 388 or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or
- the transcriptional activation domain is or comprises an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO: 105.
- the transcriptional activation domain comprises a contiguous portion of SEQ ID NO: 118 that is at least 80 amino acids in length.
- the transcriptional activation domain comprises SEQ ID NO: 105.
- the transcriptional activation domain is set forth in SEQ ID NO: 105.
- An exemplary nucleotide sequence encoding the transcriptional activation domain set forth in SEQ ID NO: 105 is set forth in SEQ ID NO: 92.
- the transcriptional activation domain is or comprises an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO:388.
- the transcriptional activation domain comprises a contiguous portion of SEQ ID NO: 118 that is at least 49 amino acids in length.
- the transcriptional activation domain comprises SEQ ID NO: 388.
- the transcriptional activation domain is set forth in SEQ ID NO:388.
- a transcriptional activation domain comprises a PSA1 domain, i.e. a domain from PSA1.
- PSA1 refers to the Proteasome subunit alpha type-1 protein encoded by the human PSMA1 gene.
- PSA1 functions as a component of the 20S core proteasome complex, which facilitates proteolytic degradation of intracellular proteins.
- An exemplary human PSA1 sequence is set forth in SEQ ID NO: 119.
- An exemplary PSA1 domain sequence is set forth in SEQ ID NO: 106 and SEQ ID NO:389.
- the transcriptional activation domain comprises a sequence set forth in any of SEQ ID NOS: 119, 106, and 389 or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to a sequence set forth in any of SEQ ID NOS: 119, 106, and 389 or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or
- the transcriptional activation domain is or comprises an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO: 106.
- the transcriptional activation domain comprises a contiguous portion of SEQ ID NO: 119 that is at least 80 amino acids in length.
- the transcriptional activation domain comprises SEQ ID NO: 106.
- the transcriptional activation domain is set forth in SEQ ID NO: 106.
- An exemplary nucleotide sequence encoding the transcriptional activation domain set forth in SEQ ID NO: 106 is set forth in SEQ ID NO:93.
- the transcriptional activation domain is or comprises an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO:389.
- the transcriptional activation domain comprises a contiguous portion of SEQ ID NO: 119 that is at least 22 amino acids in length.
- the transcriptional activation domain comprises SEQ ID NO:389.
- the transcriptional activation domain is set forth in SEQ ID NO:389.
- a transcriptional activation domain comprises a PYGO1 domain, i.e. a domain from PYGO1.
- PYGO1 refers to the Pygopus homolog 1 protein encoded by the human PYGO1 gene. PYGO1 is involved in Wnt pathway signal transduction.
- An exemplary human PYGO1 sequence is set forth in SEQ ID NO: 120.
- An exemplary PYGO1 domain sequence is set forth in SEQ ID NO: 107 and SEQ ID NO:390.
- the transcriptional activation domain comprises a sequence set forth in any of SEQ ID NOS:120, 107, and 390 or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to a sequence set forth in any of SEQ ID NOS: 120, 107, and 390 or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as
- the transcriptional activation domain is or comprises an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO: 107.
- the transcriptional activation domain comprises a contiguous portion of SEQ ID NO: 120 that is at least 80 amino acids in length.
- the transcriptional activation domain comprises SEQ ID NO: 107.
- the transcriptional activation domain is set forth in SEQ ID NO: 107.
- An exemplary nucleotide sequence encoding the transcriptional activation domain set forth in SEQ ID NO: 107 is set forth in SEQ ID NO:94.
- the transcriptional activation domain is or comprises an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO:390.
- the transcriptional activation domain comprises a contiguous portion of SEQ ID NO: 120 that is at least 57 amino acids in length.
- the transcriptional activation domain comprises SEQ ID NO:390.
- the transcriptional activation domain is set forth in SEQ ID NO:390.
- a transcriptional activation domain comprises a RBM39 domain, i.e. a domain from RBM39.
- RBM39 refers to the RNA-binding protein 39 protein encoded by the human RBM39 gene.
- RBM39 functions as a RNA-binding protein that acts as a pre-mRNA splicing factor.
- An exemplary human RBM39 sequence is set forth in SEQ ID NO: 121.
- An exemplary RBM39 domain sequence is set forth in SEQ ID NO:108 and SEQ ID NO:391.
- the transcriptional activation domain comprises a sequence set forth in any of SEQ ID NOS:121, 108, and 391 or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to a sequence set forth in any of SEQ ID NOS: 121, 108, and 391 or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80
- the transcriptional activation domain is or comprises an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO: 108.
- the transcriptional activation domain comprises a contiguous portion of SEQ ID NO: 121 that is at least 80 amino acids in length.
- the transcriptional activation domain comprises SEQ ID NO: 108.
- the transcriptional activation domain is set forth in SEQ ID NO: 108.
- An exemplary nucleotide sequence encoding the transcriptional activation domain set forth in SEQ ID NO: 108 is set forth in SEQ ID NO:95.
- the transcriptional activation domain is or comprises an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO:391.
- the transcriptional activation domain comprises a contiguous portion of SEQ ID NO: 121 that is at least 70 amino acids in length.
- the transcriptional activation domain comprises SEQ ID NO:391.
- the transcriptional activation domain is set forth in SEQ ID NO:391.
- a transcriptional activation domain comprises a HERC2 domain, i.e. a domain from HERC2.
- HERC2 refers to the E3 ubiquitin-protein ligase HERC2 protein encoded by the human HERC2 gene.
- HERC2 functions as a regulator of ubiquitin-dependent retention of repair proteins on damaged chromosomes.
- An exemplary human HERC2 sequence is set forth in SEQ ID NO: 122.
- An exemplary HERC2 domain sequence is set forth in SEQ ID NO: 109 and SEQ ID NO:392.
- the transcriptional activation domain comprises a sequence set forth in any of SEQ ID NOS: 122, 109, and 392 or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to a sequence set forth in any of SEQ ID NOS: 122, 109, and 392 or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or
- the transcriptional activation domain is or comprises an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO:109.
- the transcriptional activation domain comprises a contiguous portion of SEQ ID NO: 122 that is at least 80 amino acids in length.
- the transcriptional activation domain comprises SEQ ID NO: 109.
- the transcriptional activation domain is set forth in SEQ ID NO: 109.
- An exemplary nucleotide sequence encoding the transcriptional activation domain set forth in SEQ ID NO: 109 is set forth in SEQ ID NO:96.
- the transcriptional activation domain is or comprises an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO:392.
- the transcriptional activation domain comprises a contiguous portion of SEQ ID NO: 122 that is at least 72 amino acids in length.
- the transcriptional activation domain comprises SEQ ID NO:392.
- the transcriptional activation domain is set forth in SEQ ID NO:392.
- a transcriptional activation domain comprises a NOTCH2 domain, i.e. a domain from NOTCH2.
- NOTCH2 refers to the Neurogenic locus notch homolog protein 2 protein encoded by the human NOTCH2 gene.
- NOTCH2 functions as a receptor for membranebound ligands such as Delta- 1 to regulate cell-fate determination.
- An exemplary human NOTCH2 sequence is set forth in SEQ ID NO: 124.
- An exemplary NOTCH2 domain sequence is set forth in SEQ ID NO:111 and SEQ ID NO:393.
- the transcriptional activation domain comprises a sequence set forth in any of SEQ ID NOS:124, 111, and 393 or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to a sequence set forth in any of SEQ ID NOS:124, 111, and 393 or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as
- the transcriptional activation domain is or comprises an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO: 111.
- the transcriptional activation domain comprises a contiguous portion of SEQ ID NO: 124 that is at least 80 amino acids in length.
- the transcriptional activation domain comprises SEQ ID NO: 111.
- the transcriptional activation domain is set forth in SEQ ID NO: 111.
- An exemplary nucleotide sequence encoding the transcriptional activation domain set forth in SEQ ID NO:111 is set forth in SEQ ID NO: 98.
- the transcriptional activation domain is or comprises an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO:393.
- the transcriptional activation domain comprises a contiguous portion of SEQ ID NO: 124 that is at least 37 amino acids in length.
- the transcriptional activation domain comprises SEQ ID NO:393.
- the transcriptional activation domain is set forth in SEQ ID NO:393.
- transcriptional activation domains A variety of other effector domains for transcriptional activation (e.g., transcriptional activation domains) are known and can be used in accord with or in conjunction with the provided embodiments.
- Other transcriptional activation domains for targeted activation are described, for example, in WO 2014/197748, WO 2016/130600, WO 2017/180915, WO 2021/226555, WO 2021/226077, WO 2013/176772, WO 2014/152432, WO 2014/093661, WO 2021/247570, Adli, M. Nat. Commun. 9, 1911 (2018), Perez-Pinera, P. et al. Nat. Methods 10, 973-976 (2013), Mali, P. et al. Nat. Biotechnol.
- a transcriptional activation domain comprises a domain of a protein selected from among VP64, p65, Rta, p300, CBP, VPR, VPH, HSF1, a TET protein (e.g. TET1), a partially or fully functional fragment or domain thereof, or a combination of any of the foregoing.
- a protein selected from among VP64, p65, Rta, p300, CBP, VPR, VPH, HSF1, a TET protein (e.g. TET1), a partially or fully functional fragment or domain thereof, or a combination of any of the foregoing.
- the transcriptional activation domain comprises a VP64 domain.
- dCas9-VP64 can be targeted to a target site by one or more gRNAs to activate a gene.
- VP64 is a polypeptide composed of four tandem copies of VP 16, a 16 amino acid transactivation domain of the Herpes simplex virus.
- VP64 domains, including in dCas fusion proteins, have been described, for example, in WO 2014/197748, WO 2013/176772, WO 2014/152432, and WO 2014/093661.
- the transcriptional activation domain comprises at least one VP16 domain, or a VP16 tetramer (“VP64”) or a variant thereof.
- an exemplary VP64 domain is set forth in SEQ ID NO:81.
- An exemplary nucleotide sequence encoding the exemplary VP64 domain set forth in SEQ ID NO: 81 is set forth in SEQ ID NO: 80.
- the transcriptional activation domain comprises SEQ ID NO:81, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO:81, or a portion thereof.
- the transcriptional activation domain is set forth in SEQ ID NO:81.
- the transcriptional activation domain comprises a p65 activation domain (p65AD).
- p65AD is the principal transactivation domain of the 65kDa polypeptide of the nuclear form of the NF-KB transcription factor.
- An exemplary sequence of human transcription factor p65 is available at the Uniprot database under accession number Q04206.
- p65 domains, including in dCas fusion proteins, have been described, for example in WO 2017/180915 and Chavez, A. et al. Nat. Methods 12, 326-328 (2015).
- An exemplary p65 activation domain is set forth in SEQ ID NO: 134.
- the transcriptional activation domain comprises SEQ ID NO: 134, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO: 134, or a portion thereof. In some embodiments, the transcriptional activation domain is set forth in SEQ ID NO: 134.
- the transcriptional activation domain comprises an R transactivator (Rta) domain.
- Rta is an immediate-early protein of Epstein-Barr virus (EBV),and is a transcriptional activator that induces lytic gene expression and triggers virus reactivation.
- EBV Epstein-Barr virus
- the Rta domain including in dCas fusion proteins, has been described, for example in WO 2017/180915 and Chavez, A. et al. Nat. Methods 12, 326-328 (2015).
- An exemplary Rta domain is set forth in SEQ ID NO: 135.
- the transcriptional activation domain comprises SEQ ID NO: 135, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO: 135, or a portion thereof. In some embodiments, the transcriptional activation domain is set forth in SEQ ID NO: 135.
- the transcriptional activation domain comprises a CREB-binding protein (CBP) domain or a p300 domain.
- CBP refers to the CREB-binding protein encoded by the human CREBBP gene.
- CBP is a coactivator that interacts with cAMP-response element binding protein (CREB).
- p300 refers to the Histone acetyltransferase p300 protein encoded by the human EP300 gene, and is a coactivator closely related to CBP.
- CBP and p300 each interact with a variety of transcriptional activators to affect gene transcription (Gerritsen, M.E. et al. PNAS 94(7):2927-2932 (1997)).
- the transcriptional activation domain comprises a p300 domain.
- p300 domains (such as the catalytic core of p300) including in dCas fusion proteins for gene activation, has been described, for example, in WO 2016/130600, WO 2017/180915, and Hilton, I.B. et al., Nat. Biotechnol. 33(5):510-517 (2015).
- An exemplary human CBP sequence is set forth in SEQ ID NO:394.
- An exemplary human p300 sequence is set forth in SEQ ID NO: 125.
- An exemplary p300 domain is set forth in SEQ ID NO: 112.
- the transcriptional activation domain comprises any one of SEQ ID NOS:394, 125, and 112, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any one of SEQ ID NOS: 394, 125, and 112, or a portion thereof.
- the transcriptional activation domain comprises SEQ ID NO: 112, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO: 112, or a portion thereof.
- the transcriptional activation domain is set forth in SEQ ID NO: 112.
- the transcriptional activation domain comprises a HSF1 domain.
- HSF1 refers to the Heat shock factor protein 1 protein encoded by the human HSF1 gene. HSF1, including in dCas fusion proteins for gene activation, has been described, for example, in WO 2021/226555, WO 2015/089427, and Konermann et al. Nature 517(7536):583-8 (2015).
- An exemplary human HSF1 sequence is set forth in SEQ ID NO:395.
- An exemplary HSF1 domain sequence is set forth in SEQ ID NO: 136.
- the transcriptional activation domain comprises SEQ ID NO:136 or SEQ ID NO:395, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO: 136 or SEQ ID NO:395, or a portion thereof.
- the transcriptional activation domain comprises SEQ ID NO:136, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO: 136, or a portion thereof.
- the transcriptional activation domain is set forth in SEQ ID NO: 136.
- the transcriptional activation domain comprises the tripartite activator VP64-p65-Rta (also known as VPR).
- VPR comprises three transcription activation domains (VP64, p65, and Rta) fused by short amino acid linkers, and can effectively upregulate target gene expression.
- VPR including in dCas fusion proteins for gene activation, has been described, for example, in WO 2021/226555 and Chavez, A. et al. Nat. Methods 12, 326-328 (2015).
- An exemplary VPR polypeptide is set forth in SEQ ID NO: 137.
- the transcriptional activation domain comprises SEQ ID NO:137, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO: 137, or a portion thereof. In some embodiments, the transcriptional activation domain is set forth in SEQ ID NO: 137.
- the transcriptional activation domain comprises VPH.
- VPH is a tripartite activator polypeptide comprising VP64, mouse p65, and HSF1.
- VPH including in dCas fusion proteins for gene activation, has been described, for example, in WO 2021/226555.
- An exemplary VPH polypeptide is set forth in SEQ ID NO: 138.
- the transcriptional activation domain comprises SEQ ID NO: 138, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO: 138, or a portion thereof.
- the transcriptional activation domain is set forth in SEQ ID NO: 138.
- the transcriptional activation domain has demethylase activity.
- the transcriptional activation domain can include an enzyme that removes methyl (CH3-) groups from nucleic acids, proteins (in particular histones), and other molecules.
- the transcriptional activation domain can convert the methyl group to hydroxymethylcytosine in a mechanism for demethylating DNA.
- the effector domain can catalyze this reaction.
- the transcriptional activation domain that catalyzes this reaction may comprise a domain from a TET protein, for example TET1 (Ten-eleven translocation methylcytosine dioxygenase 1).
- TET1 refers to the Methylcytosine dioxygenase TET1 protein encoded by the human TET1 gene.
- TET1 catalyzes the conversion of the modified genomic base 5 -methylcytosine (5mC) into 5-hydroxymethylcytosine (5hmC) and plays a key role in active DNA demethylation.
- TET1 including in dCas fusion proteins for gene activation, has been described, for example, in WO 2021/226555.
- An exemplary human TET1 sequence is set forth in SEQ ID NO:396.
- An exemplary TET1 catalytic domain is set forth in SEQ ID NO: 139.
- the transcriptional activation domain comprises SEQ ID NO:396 or SEQ ID NO:139, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO:396 or SEQ ID NO: 139, or a portion thereof.
- the transcriptional activation domain comprises SEQ ID NO: 139, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO: 139, or a portion thereof.
- the transcriptional activation domain is set forth in SEQ ID NO: 139.
- the effector domain may comprise a LSD1 domain.
- LSD1 also known as Lysine-specific histone demethylase 1A
- LSD1 is a histone demethylase that can demethylate lysine residues of histone H3, thereby acting as a coactivator or a corepressor, depending on the context.
- LSD1 including in dCas fusion proteins, has been described, for example, in WO 2013/176772, WO 2014/152432, and Kearns, N. A. et al. Nat. Methods. 12(5):401-403 (2015).
- An exemplary LSD1 polypeptide is set forth in SEQ ID NO: 140.
- the effector domain comprises the sequence set forth in SEQ ID NO: 140, a domain thereof, a portion thereof, or a variant thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
- the effector domain may comprise a SunTag domain.
- SunTag is a repeating peptide array, which can recruit multiple copies of an antibody-fusion protein that binds the repeating peptide.
- the antibody-fusion protein may comprise an additional effector domain, such as a transcription activation domain (e.g. VP64), to induce increased transcription of the target gene.
- a transcription activation domain e.g. VP64
- SunTag including in dCas fusion proteins for gene activation, has been described, for example, in WO 2016/011070 and Tanenbaum, M. et al. Cell. 159(3):635-646 (2014).
- An exemplary SunTag effector domain includes a repeating GCN4 peptide having the amino acid sequence LLPKNYHLENEVARLKKLVGER (SEQ ID NO: 152) separated by linkers having the amino acid sequence GGSGG (SEQ ID NO: 153).
- the effector domain comprises the sequence set forth in SEQ ID NO: 152, a domain thereof, a portion thereof, or a variant thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
- the SunTag effector domain recruits an antibody-fusion protein that comprises any of the epigenetic effector domains described herein, such as VP64, and binds the GCN4 peptide. In some embodiments, the SunTag domain recruits one or more epigenetic effector domains for transcriptional activation to the FXN locus, thereby increasing FXN expression.
- the effector domain may have demethylase activity.
- the effector domain may include an enzyme that remove methyl (CH3-) groups from nucleic acids, proteins (in particular histones), and other molecules.
- the effector domain may covert the methyl group to hydroxymethylcytosine in a mechanism for demethylating DNA.
- multipartite effectors for transcriptional activation of FXN for example, multipartite transcriptional activation domains or multipartite activators.
- the epigenetic effector domain is a multipartite effector, such as a multipartite activator.
- the multipartite activator is a fusion protein or a sequence of amino acids comprising two or more transcriptional activation domains, such as any of the effector domains such as transcriptional activation domains provided herein.
- the multipartite activator comprises two or more transcriptional activation domains, each transcriptional activation domain comprising a domain of a protein selected from among NCOA3, ENL, FOXO3, PYGO1, HSH2D, NCOA2, NOTCH2, DPOLA, PSA1, RBM39, and HERC2.
- the multipartite activator is provided as part of a fusion protein or DNA-targeting system, such as any described herein, including an eZFP fusion protein, a ZFP- based DNA-targeting system, and a CRISPR/Cas-based DNA-targeting system.
- the multipartite activator increases transcription of an endogenous locus, such as FXN, when recruited to a target site at the endogenous locus.
- the multipartite activator increases transcription of a FXN gene when recruited (e.g. targeted to) a target site for the FXN gene, such as a regulatory DNA element (e.g. a promoter or enhancer), and/or any target site described herein.
- a multipartite activator may itself be referred to as a transcriptional activation domain herein.
- the multipartite activator induces, catalyzes, or leads to increased transcription of FXN when ectopically recruited to the FXN locus or a DNA regulatory element thereof. In some embodiments, the multipartite activator activates, induces, catalyzes, or leads to: transcription activation, transcription co-activation, transcription elongation, or transcription derepression. In some embodiments, the multipartite activator induces transcriptional activation. In some embodiments, the multipartite activator has one of the aforementioned activities itself (i.e. acts directly). In some embodiments, the multipartite activator recruits and/or interacts with a polypeptide domain that has one of the aforementioned activities (i.e. acts indirectly).
- a multipartite activator provided herein comprises two or more transcriptional activation domains.
- the multipartite activator has an effect that is different from any one of the individual transcriptional activation domains comprised by the multipartite activator alone. The different effect may be quantitatively or qualitatively different.
- the multipartite activator may induce greater, more reliable, or more durable transcriptional activation of a target gene, in comparison to a transcriptional activation domain alone.
- the effect may be context-specific.
- a multipartite activator may induce transcriptional activation in a specific context in which the transcriptional activation domain alone does not induce transcriptional activation to the same degree, at a detectable level, or at all, such as when targeted to a specific gene or target site of the gene.
- a multipartite activator does not necessarily lead to greater activation of a target gene than a transcriptional activation domain alone in every context, but may allow for activation of a target gene in different contexts and to a different degree than the transcriptional activation domain.
- a multipartite activator may have a more durable effect on transcription than a transcriptional activation domain alone.
- a multipartite activator may lead to increased transcription of a target gene in a cell for a longer amount of time, or for a greater number of cell divisions or cell passages.
- the multipartite effector e.g., multipartite activator
- the multipartite effector e.g., multipartite activator
- the multipartite effector, e.g., multipartite activator comprises 4, 5, 6, 7, 8, 9, 10, or more transcriptional activation domains.
- any two or more of the transcriptional activation domains are the same.
- any two or more of the transcriptional activation domains are different.
- the multipartite activator comprises two or more transcriptional activation domains, each transcriptional activation domain comprising a domain of a protein selected from among DPOLA, ENL, FOXO3, HSH2D, NCOA2, NCOA3, PSA1, PYGO1, RBM39, HERC2, or NOTCH2.
- the multipartite activator comprises two or more transcriptional activation domains, wherein one or more of the transcriptional activation domains comprises a domain of a protein selected from among DPOLA, ENL, FOXO3, HSH2D, NCOA2, NCOA3, PSA1, PYGO1, RBM39, HERC2, or N0TCH2.
- the transcriptional activation domain from DPOLA, ENL, FOXO3, HSH2D, NC0A2, NC0A3, PSA1, PYGO1, RBM39, HERC2, or NOTCH2 is or comprises any of the respective transcriptional activation domains described herein, for example, in Section II.B.l, or a partially or fully functional fragment thereof, a domain thereof, or a portion thereof, such as a contiguous portion thereof of at least 30 amino acids, or a variant thereof.
- the transcriptional activation domain from DPOLA, ENL, FOXO3, HSH2D, NCOA2, NCOA3, PSA1, PYGO1, RBM39, HERC2, or NOTCH2 is or comprises any of sequences of the respective transcriptional activation domains described herein, for example, in Section II.B.l, or a partially or fully functional fragment thereof, a domain thereof, or a portion thereof, such as a contiguous portion thereof of at least 30 amino acids, or a variant thereof.
- the multipartite activator further comprises one or more of any of the transcriptional activation domains provided herein, such as VP64, p65, Rta, p300, CBP, VPR, VPH, HSF1, a TET protein (e.g., TET1), a partially or fully functional fragment or domain thereof, or a combination of any of the foregoing.
- the transcriptional activation domains provided herein, such as VP64, p65, Rta, p300, CBP, VPR, VPH, HSF1, a TET protein (e.g., TET1), a partially or fully functional fragment or domain thereof, or a combination of any of the foregoing.
- the multipartite activator is a bipartite activator comprising a first transcriptional activation domain and a second transcriptional activation domain.
- each of the first transcriptional activation domain and the second transcriptional activation domain independently comprises a domain of a protein selected from among DPOLA, ENL, FOXO3, HSH2D, NCOA2, NCOA3, PSA1, PYGO1, RBM39, HERC2, and NOTCH2.
- the first and second transcriptional activation domains are from DPOLA and DPOLA; DPOLA and ENL; DPOLA and FOXO3; DPOLA and HERC2; DPOLA and HSH2D; DPOLA and NCOA2; DPOLA and NCOA3; DPOLA and NOTCH2; DPOLA and PSA1; DPOLA and PYGO1; DPOLA and RBM39; ENL and DPOLA; ENL and ENL; ENL and FOXO3; ENL and HERC2; ENL and HSH2D; ENL and NCOA2; ENL and NCOA3; ENL and NOTCH2; ENL and PSA1; ENL and PYGO1; ENL and RBM39; FOXO3 and DPOLA; FOXO3 and ENL; FOXO3 and FOXO3; FOXO3 and HERC2; FOXO3 and HSH2D; FOXO3 and NCOA3 and HSH2D; FOXO3 and NCOA3 and H
- the multipartite activator is a tripartite activator comprising a first transcriptional activation domain, a second transcriptional activation domain, and a third transcriptional activation domain.
- the first transcriptional activation domain, the second transcriptional activation domain, and the third transcriptional activation domain each independently comprises a domain of a protein selected from among DPOLA, ENL, FOXO3, HSH2D, NCOA2, NCOA3, PSA1, PYGO1, RBM39, HERC2, and NOTCH2.
- the first and second transcriptional domains are the first and second transcriptional domains from any of the bipartite activators described above, and the third transcriptional domain independently comprises a domain of a protein selected from among DPOLA, ENL, FOXO3, HSH2D, NCOA2, NCOA3, PSA1, PYGO1, RBM39, HERC2, and NOTCH2.
- the multipartite activator is a tripartite activator comprising a first transcriptional activation domain, a second transcriptional activation domain, and a third transcriptional activation domain, each independently comprising a domain of a protein selected from among DPOLA, ENL, FOXO3, HSH2D, NCOA2, NCOA3, PSA1, PYGO1, RBM39, HERC2, and NOTCH2.
- the first, second, and third transcriptional activation domains are from NCOA3, NCOA3, and NCOA3; NCOA3, NCOA3, and ENL; NCOA3, NCOA3, and FOXO3; NCOA3, NCOA3, and PYGO1; NCOA3, NCOA3, and HSH2D; NCOA3, NCOA3, and NCOA2; NCOA3, NCOA3, and NOTCH2; NCOA3, ENL, and NCOA3; NCOA3, ENL, and ENL; NCOA3, ENL, and FOXO3; NCOA3, ENL, and PYGO1; NCOA3, ENL, and HSH2D; NCOA3, ENL, and NCOA2; NCOA3, ENL, and NOTCH2; NCOA3, FOXO3, and NCOA3; NCOA3, FOXO3, and ENL; NCOA3, FOXO3, and FOXO3; NCOA3, FOXO3, and PYGO1; NCOA3, FOXO3, and HSH2D; NCOA3, FOXO3, and NCOA2;
- HSH2D, HSH2D, and NCOA2 HSH2D, HSH2D, and NOTCH2; HSH2D, NCOA2, and NCOA3;
- the first, second, and third transcriptional activation domains, respectively are from PYGO1, FOXO3, and NCOA3, respectively. In some embodiments, the first, second, and third transcriptional activation domains, respectively, are from NOTCH2, FOXO3, and NCOA3, respectively. In some embodiments, the first, second, and third transcriptional activation domains, respectively, are from NCOA3, FOXO3, and NCOA3, respectively. In some embodiments, the first, second, and third transcriptional activation domains, respectively, are from HSH2D, FOXO3, and NCOA3, respectively. In some embodiments, the first, second, and third transcriptional activation domains, respectively, are from FOXO3, FOXO3, and NCOA3, respectively.
- the first, second, and third transcriptional activation domains, respectively are from NCOA2, FOXO3, and NCOA3, respectively. In some embodiments, the first, second, and third transcriptional activation domains, respectively, are from ENL, FOXO3, and NCOA3, respectively.
- the first, second, and third transcriptional activation domains, respectively are from FOXO3, FOXO3, and NCOA3, respectively. In some embodiments, the first, second, and third transcriptional activation domains, respectively, are from NCOA3, FOXO3, and FOXO3, respectively. In some embodiments, the first, second, and third transcriptional activation domains, respectively, are from NCOA3, FOXO3, and NCOA3, respectively.
- any of the provided multipartite effector proteins, fusion proteins, and/or DNA targeting systems, such as a multipartite activator comprises a combination of transcriptional activation domains, such as a combination of two or more of any of transcriptional activation domains shown in Table 3.
- any of the provided multipartite effector proteins, fusion proteins, and/or DNA targeting systems comprises a combination of two or more of any one of the SEQ ID NOS: set forth in Table 3, or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any one of the SEQ ID NOS: set forth in Table 3, or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57,
- any of the provided multipartite effector proteins, fusion proteins, and/or DNA targeting systems comprises a combination of two or more of any one of the SEQ ID NOS: set forth in Table 3, or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof.
- any of the provided multipartite effector proteins, fusion proteins, and/or DNA targeting systems, such as a multipartite activator comprises two or more of any one of the SEQ ID NOS: set forth in Table 3.
- any of the provided multipartite effector proteins, fusion proteins, and/or DNA targeting systems comprises a combination of transcriptional activation domains, such as a combination of three or more of any of transcriptional activation domains shown in Table 3.
- any of the provided multipartite effector proteins, fusion proteins, and/or DNA targeting systems comprises a combination of three or more of any one of the SEQ ID NOS: set forth in Table 3, or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any one of the SEQ ID NOS: set forth in Table 3, or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57,
- any of the provided multipartite effector proteins, fusion proteins, and/or DNA targeting systems comprises a combination of three or more of any one of the SEQ ID NOS: set forth in Table 3, or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof.
- any of the provided multipartite effector proteins, fusion proteins, and/or DNA targeting systems, such as a multipartite activator comprises three or more of any one of the SEQ ID NOS: set forth in Table 3.
- the multipartite activator comprises the any one of the SEQ ID NOS: set forth in Table 4, or a domain, portion, or variant thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any one of the SEQ ID NOS: set forth in Table 4.
- the multipartite activator is or comprises any one of the SEQ ID NOS: set forth in Table 4.
- the multipartite activator comprises a combination of transcriptional activation domains, such as any of the combinations of transcriptional activation domains shown in Table 4.
- the multipartite activator comprises any one of SEQ ID NOS:397- 418, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any one of SEQ ID NOS:397-418.
- the multipartite activator is set forth in any one of SEQ ID NOS:397-418, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity any one of SEQ ID NOS:397-418, or a partially or fully functional fragment thereof, a domain thereof, or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 25, 30, 35, 40, 45, 50, 55, 60, 65, 70, 75, or 80 amino acids, or a variant thereof.
- the multipartite activator is set forth in any one of SEQ ID NOS:397-418.
- the multipartite activator comprises domains from NCOA3, FOXO3, and NCOA3, respectively.
- the multipartite activator comprises SEQ ID NO:413, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO:413.
- the multipartite activator is set forth in SEQ ID NO:413.
- the multipartite activator comprises domains from FOXO3, FOXO3, and NCOA3, respectively.
- the multipartite activator comprises SEQ ID NO:415, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO:415.
- the multipartite activator is set forth in SEQ ID NO:415.
- the multipartite activator comprises domains from NCOA3, FOXO3, and FOXO3, respectively.
- the multipartite activator comprises SEQ ID NO:418, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO:418.
- the multipartite activator is set forth in SEQ ID NO:418.
- the DNA-targeting system comprises an eZFP fusion protein, such as any provided herein.
- eZFP fusion proteins such as any provided herein.
- fusion proteins such as fusion proteins targeting, or capable of being targeted to, a FXN locus.
- the fusion proteins comprise an eZFP or a dCas protein.
- the fusion protein comprises an eZFP (i.e. the fusion protein is an eZFP fusion protein), such as any of the eZFPs described herein, for example in Section I.
- the fusion protein further comprises an epigenetic effector domain, such as any of the effector domains for transcriptional activation described herein, for example in Section II.B.
- the fusion protein comprises at least one epigenetic effector domain that increases transcription of the FXN locus.
- the fusion protein comprises more than one effector domain.
- the fusion protein comprises one or more additional elements, such as a nuclear localization signal (NLS) or linker, such as any of the NLSs or linkers described herein.
- the elements of the fusion protein may be arranged in any suitable order within the fusion protein, such as an order from N-terminus to C-terminus.
- the fusion proteins comprising eZFPs provided herein may facilitate increased FXN expression, for example in connection with compositions and methods for treating a disease or disorder associated with FXN expression, such as Friedreich’s ataxia (FA).
- FA Friedreich’s ataxia
- the fusion protein comprising the eZFP binds to, or is capable of binding to, (i.e. targets), any of the target sites provided herein. In some embodiments, the fusion protein binds to the target site. In some embodiments, the fusion protein comprising the eZFP binds to the target site that the eZFP binds to in the absence of the other elements of the fusion protein. Thus, in some embodiments, the eZFP of the fusion protein facilitates target-specific binding of the fusion protein. In some aspects, the fusion protein targets to the target site targeted by any of the eZFPs described herein, such as in Section I. In some embodiments, the fusion protein targets a target site in Table 1.
- the fusion protein targets a target site in Table 2A and Table 2B. In some embodiments, the fusion protein targets a target site comprising the nucleotide sequence set forth in any one of SEQ ID NOS:272, 277, 280, 281, 283, 290, or 299, or a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the fusion protein targets a target site comprising the nucleotide sequence set forth in any one of SEQ ID NOS:583-600, or a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the fusion protein targets a target site comprising the nucleotide sequence set forth in any one of SEQ ID NOS:272, 277, 280, 281, 283, 290, or 299. In some embodiments, the fusion protein targets a target site comprising the nucleotide sequence set forth in any one of SEQ ID NOS:583-600. In some embodiments, the fusion protein targets a target site comprising the nucleotide sequence set forth in SEQ ID NO: 299. In some embodiments, the fusion protein targets a target site comprising the nucleotide sequence set forth in SEQ ID NO: 587.
- the fusion protein targets a target site comprising the nucleotide sequence set forth in SEQ ID NO: 589. In some embodiments, the fusion protein targets a target site comprising the nucleotide sequence set forth in SEQ ID NO: 591. In some embodiments, the target site is double-stranded DNA.
- the fusion protein comprises any of the eZFPs set forth in Table 2A and Table 2B.
- the fusion protein comprises an eZFP comprising the recognition regions Fl- F6 set forth for any of the eZFPs set forth in Table 2A and Table 2B (comprising SEQ ID NOS:341-346, respectively; SEQ ID NOS:347-352, respectively; SEQ ID NOS:353-358, respectively; SEQ ID NOS:359-364, respectively; SEQ ID NOS:365-370, respectively; SEQ ID NOS:371-376, respectively; SEQ ID NOS:377-382, respectively; SEQ ID NOS:475-480, respectively; SEQ ID NOS:481-486, respectively; SEQ ID NOS:487-492, respectively; SEQ ID NOS:493-498, respectively; SEQ ID NOS:499-504, respectively; SEQ ID NOS:505-510, respectively; SEQ ID NOS:511-516
- the fusion protein comprises the amino acid sequence set forth in any one of SEQ ID NOS:301-307 and 439-456, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the fusion protein comprises the amino acid sequence set forth in any one of SEQ ID NOS:301-307 and 439-456.
- the eZFP of the fusion protein is encoded by the nucleotide sequence set forth in any one of SEQ ID NOS:308-314 and 457-474, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP of the fusion protein is encoded by the nucleotide sequence set forth in any one of SEQ ID NOS:308-314 and 457-474.
- a fusion protein comprising an eZFP, such as any of the eZPFs described herein, for example in Table 2A or Table 2B.
- the fusion protein targets a target site comprising the nucleotide sequence set forth in any one of SEQ ID NOS: 272, 277, 280, 281, 283, 290, or 299, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the fusion protein targets a target site comprising the nucleotide sequence set forth in any one of SEQ ID NOS: 583-600, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the fusion protein targets a target site comprising the nucleotide sequence set forth in any one of SEQ ID NOS:272, 277, 280, 281, 283, 290, or 299. In some embodiments, the fusion protein targets a target site comprising the nucleotide sequence set forth in any one of SEQ ID NOS:583-600. In some embodiments, the target site is double-stranded DNA.
- the eZFP of the fusion protein comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequences of the recognition regions F1-F6 comprise: SEQ ID NOS:341-346, respectively; SEQ ID NOS:347-352, respectively; SEQ ID NOS:353-358, respectively; SEQ ID NOS:359-364, respectively; SEQ ID NOS:365-370, respectively; SEQ ID NOS:371-376, respectively; or SEQ ID NOS:377-382, respectively.
- the eZFP of the fusion protein comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequences of the recognition regions F1-F6 comprise: SEQ ID NOS:475-480, respectively; SEQ ID NOS:481-486, respectively; SEQ ID NOS:487-492, respectively; SEQ ID NOS:493-498, respectively; SEQ ID NOS:499-504, respectively; SEQ ID NOS:505-510, respectively; SEQ ID NOS:511-516, respectively; SEQ ID NOS:517-522, respectively; SEQ ID NOS:523-528, respectively; SEQ ID NOS:529-534, respectively; SEQ ID NOS: 535-540, respectively; SEQ ID NOS:541-546, respectively; SEQ ID NOS:547-552, respectively; SEQ ID NOS:553-558, respectively; SEQ ID NOS:
- the eZFP of the fusion protein comprises the amino acid sequence set forth in any one of SEQ ID NOS:301-307, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the eZFP of the fusion protein comprises the amino acid sequence set forth in any one of SEQ ID NOS:439-456, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the eZFP of the fusion protein comprises the amino acid sequence set forth in any one of SEQ ID NOS:301-307. In some embodiments, the eZFP of the fusion protein comprises the amino acid sequence set forth in any one of SEQ ID NOS:439-456. In some embodiments, the eZFP of the fusion protein is encoded by the nucleotide sequence set forth in any one of SEQ ID NOS:308-314, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the eZFP of the fusion protein is encoded by the nucleotide sequence set forth in any one of SEQ ID NOS:457-474, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the eZFP of the fusion protein is encoded by the nucleotide sequence set forth in any one of SEQ ID NOS:308-314.
- the eZFP of the fusion protein is encoded by the nucleotide sequence set forth in any one of SEQ ID NOS:457-474.
- a fusion protein comprising an eZFP, such as eZFP_A31 as described herein.
- the fusion protein targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:299, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the fusion protein targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:299.
- the target site is double-stranded DNA.
- the eZFP of the fusion protein comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: LRHHLTR (SEQ ID NO:377); F2: QSAHLKA (SEQ ID NO:378); F3: LPQTLQR (SEQ ID NO:379); F4: QNATRTK (SEQ ID NO:380); F5: QSSHLTR (SEQ ID NO:381); F6: RSDHLSR (SEQ ID NO:382).
- the eZFP of the fusion protein comprises the amino acid sequence set forth in SEQ ID NO:307, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP of the fusion protein comprises the amino acid sequence set forth in SEQ ID NO:307.
- the eZFP of the fusion protein is encoded by the nucleotide sequence set forth in SEQ ID NO:314, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP of the fusion protein is encoded by the nucleotide sequence set forth in SEQ ID NO:314.
- a fusion protein comprising an eZFP, such as eZFP_A44 as described herein.
- the fusion protein targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:587, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the fusion protein targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:587.
- the target site is double-stranded DNA.
- the eZFP of the fusion protein comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: QAGNRST (SEQ ID NO:499); F2: DRSALAR (SEQ ID NG:500); F3: RSDNLAR (SEQ ID NO:501); F4: WRGDRVK (SEQ ID NO:502); F5: YKHVLSD (SEQ ID NO:503); and F6: TSGSLTR (SEQ ID NO:504).
- the eZFP of the fusion protein comprises the amino acid sequence set forth in SEQ ID NO: 443, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the eZFP of the fusion protein comprises the amino acid sequence set forth in SEQ ID NO:443.
- the eZFP of the fusion protein is encoded by the nucleotide sequence set forth in SEQ ID NO:461, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the eZFP of the fusion protein is encoded by the nucleotide sequence set forth in SEQ ID NO:461.
- a fusion protein comprising an eZFP, such as eZFP_A46 as described herein.
- the fusion protein targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:589, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the fusion protein targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:589.
- the target site is double-stranded DNA.
- the eZFP of the fusion protein comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: DRSTRTK (SEQ ID NO:511); F2: RSDYLAK (SEQ ID NO:512); F3: LRHHLTR (SEQ ID NO:513); F4: QSAHLKA (SEQ ID NO:514); F5: LPQTLQR (SEQ ID NO:515); and F6: QNATRTK (SEQ ID NO:516).
- the eZFP of the fusion protein comprises the amino acid sequence set forth in SEQ ID NO: 445, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP of the fusion protein comprises the amino acid sequence set forth in SEQ ID NO:445.
- the eZFP of the fusion protein is encoded by the nucleotide sequence set forth in SEQ ID NO:463, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP of the fusion protein is encoded by the nucleotide sequence set forth in SEQ ID NO:463.
- a fusion protein comprising an eZFP, such as eZFP_A48 as described herein.
- the fusion protein targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:591, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
- the fusion protein targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:591.
- the target site is double-stranded DNA.
- the eZFP of the fusion protein comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: RNDALTE (SEQ ID NO:523); F2: RKDNLKN (SEQ ID NO:524); F3: TSGELVR (SEQ ID NO:525); F4: HRTTLTN (SEQ ID NO:526); F5: TTGNLTV (SEQ ID NO:527); and F6: RTDTLRD (SEQ ID NO:528).
- the eZFP of the fusion protein comprises the amino acid sequence set forth in SEQ ID NO: 447, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP of the fusion protein comprises the amino acid sequence set forth in SEQ ID NO:447.
- the eZFP of the fusion protein is encoded by the nucleotide sequence set forth in SEQ ID NO:465, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the eZFP of the fusion protein is encoded by the nucleotide sequence set forth in SEQ ID NO:465.
- the fusion protein further comprises one or more nuclear localization signal (NLS), such as any suitable NLS, for example any NLS described herein.
- an NLS may promote nuclear localization of the fusion protein.
- the NLS comprises the amino acid sequence set forth in any one of SEQ ID NOS:85 and 159-173, or an amino acid sequence having at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the NLS comprises the amino acid sequence set forth in any one of SEQ ID NOS:85 and 159-173.
- the NLS is an SV40 NLS.
- the SV40 NLS comprises the amino acid sequence set forth in SEQ ID NO: 159.
- the NLS is a c-myc NLS.
- the c-myc NLS comprises the amino acid sequence set forth in SEQ ID NO: 160.
- a fusion protein described herein comprises one or more nuclear localization sequences (NLSs), such as about or more than about 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, or more NLSs.
- NLSs nuclear localization sequences
- each may be selected independently of the others, such that a single NLS may be present in more than one copy and/or in combination with one or more other NLSs present in one or more copies.
- NLSs include an NLS sequence derived from: the NLS of the SV40 virus large T-antigen, having the sequence PKKKRKV (SEQ ID NO: 159); the NLS from nucleoplasmin (e.g.
- the nucleoplasmin bipartite NLS having the sequence KRPAATKKAGQAKKKK (SEQ ID NO:85); the c-myc NLS having the sequence PAAKRVKLD (SEQ ID NO: 160) or RQRRNELKRSP (SEQ ID NO: 161); the hRNPAl M9 NLS having the sequence NQSSNFGPMKGGNFGGRSSGPYGGGGQYFAKPRNQGGY (SEQ ID NO: 162); the sequence RMRIZFKNKGKDTAELRRRRVEVSVELRKAKKDEQILKRRNV (SEQ ID NO: 163) of the IBB domain from importin-alpha; the sequences VSRKRPRP (SEQ ID NO: 164) and PPKKARED (SEQ ID NO: 165) of the myoma T protein; the sequence PQPKKKPL (SEQ ID NO: 166) of human p53; the sequence SALIKKKKKMAP (SEQ ID NO: 167) of mouse c-abl IV; the sequences D
- the one or more NLSs are of sufficient strength to drive accumulation of the fusion protein in a detectable amount in the nucleus of a eukaryotic cell.
- strength of nuclear localization activity may derive from the number of NLSs in the fusion protein, the particular NLS(s) used, or a combination of these factors.
- Detection of accumulation in the nucleus may be performed by any suitable technique.
- a detectable marker may be fused to the fusion protein, such that location within a cell may be visualized, such as in combination with a means for detecting the location of the nucleus (e.g. a stain specific for the nucleus such as DAPI).
- Cell nuclei may also be isolated from cells, the contents of which may then be analyzed by any suitable process for detecting protein, such as immunohistochemistry, Western blot, or enzyme activity assay. Accumulation in the nucleus may also be determined indirectly, such as by an assay for the effect of the fusion protein (e.g. an assay for altered gene expression activity in a cell transformed with the DNA-targeting system comprising the fusion protein), as compared to a control condition (e.g. an untransformed cell).
- an assay for the effect of the fusion protein e.g. an assay for altered gene expression activity in a cell transformed with the DNA-targeting system comprising the fusion protein
- a control condition e.g. an untransformed cell
- the fusion protein further comprises one or more linker, such as any suitable linker, for example any linker described herein.
- the one or more linkers may connect the eZFP and the at least one epigenetic effector domain.
- each of the one or more linkers are in between any two of the components of the fusion protein, including the eZFP, any of the at least one effector domains, and the one or more NLS.
- a linker may be of any length.
- a linker may be designed to promote or restrict the mobility of components in the fusion protein.
- a linker in the fusion protein has the amino acid sequence set forth in any one of SEQ ID NOS:153-158, 174, 186, 188, and 219, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- a linker in the fusion protein has the amino acid sequence set forth in any one of SEQ ID NOS:153-158, 174, 186, 188, and 219.
- a linker may comprise any amino acid sequence of about 2 to about 100, about 5 to about 80, about 10 to about 60, or about 20 to about 50 amino acids.
- a linker may comprise an amino acid sequence of at least about 2, 3, 4, 5, 10, 15, 20, 25, 30, 35, 40, 45, 50, 55, 60, 65, 70, 75, 80 or 85 amino acids.
- a linker may comprise an amino acid sequence of less than about 100, 90, 80, 70, 60, 50, or 40 amino acids.
- a linker may include sequential or tandem repeats of an amino acid sequence that is 2 to 20 amino acids in length.
- Linkers may be rich in amino acids glycine (G), serine (S), and/or alanine (A).
- Linkers may include, for example, a GS linker.
- An exemplary GS linker is represented by the sequence GGGGS (SEQ ID NO: 158).
- a linker may comprise repeats of a sequence, for example as represented by the formula (GGGGS)n, wherein n is an integer that represents the number of times the GGGGS sequence is repeated (e.g. between 1 and 10 times). The number of times a linker sequence is repeated, for example n in a GS linker, can be adjusted to optimize the linker length and achieve appropriate separation of the functional domains.
- linkers may include, for example, GGGGG (SEQ ID NO: 154), GGAGG (SEQ ID NO: 155), Gly/Ser rich linkers such as GGGGSSS (SEQ ID NO: 156), or Gly/Ala rich linkers such as GGGGAAA (SEQ ID NO: 157), or GSGSG (SEQ ID NO:219).
- the linker is an XTEN linker.
- an XTEN linker is a recombinant polypeptide (e.g., an unstructured recombinant peptide) lacking hydrophobic amino acid residues.
- Exemplary XTEN linkers are described in, for example, Schellenberger et al., Nature Biotechnology 27, 1186-1190 (2009) or WO 2021/247570.
- an exemplary linker comprises a linker described in WO 2021/247570.
- the linker is or comprises the sequence set forth in SEQ ID NO: 186 or SEQ ID NO: 174, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
- the linker comprises the sequence set forth in SEQ ID NO:186, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
- the linker comprises the sequence set forth in SEQ ID NO: 186, or a contiguous portion of SEQ ID NO: 186 of at least 5, 10, 15, 20, 25, 30, 35, 40, 45, 50, 55, 60, 65, 70 or 75 amino acids. In some aspects, the linker consists of the sequence set forth in SEQ ID NO: 186, or a contiguous portion of SEQ ID NO: 186 of at least 5, 10, 15, 20, 25, 30, 35, 40, 45, 50, 55, 60, 65, 70 or 75 amino acids. In some embodiments, the linker comprises the sequence set forth in SEQ ID NO: 186. In some embodiments, the linker consists of the sequence set forth in SEQ ID NO: 186.
- the linker is encoded by a nucleotide sequence set forth in SEQ ID NO: 185.
- the linker comprises the sequence set forth in SEQ ID NO:174, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
- the linker comprises the sequence set forth in SEQ ID NO: 174, or a contiguous portion of SEQ ID NO: 174 of at least 5, 10, or 15 amino acids.
- the linker consists of the sequence set forth in SEQ ID NO: 174, or a contiguous portion of SEQ ID NO: 174 of at least 5, 10, or 15 amino acids. In some embodiments, the linker comprises the sequence set forth in SEQ ID NO: 174. In some embodiments, the linker consists of the sequence set forth in SEQ ID NO: 174. Appropriate linkers may be selected or designed based rational criteria known in the art, for example as described in Chen et al. Adv. Drug Deliv. Rev. 65(10): 1357-1369 (2013).
- a linker comprises the sequence set forth in SEQ ID NO:188, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
- the linker is a self-cleaving peptide, e.g., P2A.
- An exemplary P2A peptide sequence is set forth in SEQ ID NO: 131.
- a P2A peptide sequence is encoded by the nucleotide sequence set forth in SEQ ID NO: 130.
- the fusion protein comprises at least one epigenetic effector domain that increases transcription of the FXN locus.
- the at least one epigenetic effector domain is an effector domain for transcriptional activation and/or a multipartite activator, such as any of the effector domains described herein, for example in Section III.
- the eZFP fusion protein comprises the one or more NLS, the eZFP, and the at least one epigenetic effector domain, in order from N-terminus to C-terminus, optionally wherein the NLS, eZFP, and at least one epigenetic effector domain are separated by one or more linker and/or additional NLS.
- the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:320-340, 419-425, and 608-671, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
- the fusion protein comprises the sequence set forth in any one of SEQ ID NOS:320-340, 419-425, and 608-671.
- the at least one epigenetic effector domain of the eZFP fusion protein comprises a VP64 domain.
- the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:320-326 and 608-614, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
- the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:320-326 and 608-641.
- the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:320-326, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing. In some embodiments, the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:320-326.
- the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:608-641, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
- the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:608-641.
- the eZFP fusion protein comprises the sequence set forth in SEQ ID NO:326.
- the eZFP fusion protein comprises the sequence set forth in SEQ ID NO:340.
- the eZFP fusion protein is encoded by the nucleotide sequence set forth in SEQ ID NO:426, or a nucleotide sequence having at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the eZFP fusion protein is encoded by the nucleotide sequence set forth in SEQ ID NO:426.
- the eZFP fusion protein comprises the sequence set forth in SEQ ID NO:634.
- the at least one epigenetic effector domain of the eZFP fusion protein comprises a multipartite activator comprising, in the N-terminal to C-terminal direction, domains from FOXO3, FOXO3, and NCOA3, respectively, optionally separated by one or more linkers.
- the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:334- 340 and 629-635, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
- the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:334-340 and 629-635. In some embodiments, the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:334-340, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing. In some embodiments, the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:334-340.
- the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS: 629- 635, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
- the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS: 629-635.
- the eZFP fusion protein comprises the sequence set forth in SEQ ID NO:340.
- the eZFP fusion protein is encoded by the nucleotide sequence set forth in SEQ ID NO:428, or a nucleotide sequence having at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the eZFP fusion protein is encoded by the nucleotide sequence set forth in SEQ ID NO:428.
- the eZFP fusion protein comprises the sequence set forth in SEQ ID NO:635.
- the at least one epigenetic effector domain of the eZFP fusion protein comprises a multipartite activator comprising, in the N-terminal to C-terminal direction, domains from NCOA3, FOXO3, and FOXO3, respectively, optionally separated by one or more linkers.
- the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:419- 425 and 615-621, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
- the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:419-425 and 615-621. In some embodiments, the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:419-425, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing. In some embodiments, the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:419-425.
- the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:615- 621, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
- the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:615-621.
- the eZFP fusion protein comprises the sequence set forth in SEQ ID NO: 425.
- the eZFP fusion protein comprises the sequence set forth in SEQ ID NO: 621.
- the at least one epigenetic effector domain of the eZFP fusion protein comprises a multipartite activator comprising, in the N-terminal to C-terminal direction, domains from NCOA3, FOXO3, and NCOA3, respectively, optionally separated by one or more linker and/or NFS.
- the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:327-333, 615-621, and 636-671, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
- the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:327-333, 615-621, and 636-671. In some embodiments, the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:327-333, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing. In some embodiments, the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:327-333.
- the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:615-621, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing. In some embodiments, the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:615-621.
- the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:636-653, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing. In some embodiments, the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS: 636-653.
- the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:654-671, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
- the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:654- 671.
- the eZFP fusion protein comprises the sequence set forth in SEQ ID NO: 333.
- the eZFP fusion protein is encoded by the nucleotide sequence set forth in SEQ ID NO:427, or a nucleotide sequence having at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the eZFP fusion protein is encoded by the nucleotide sequence set forth in SEQ ID NO:427.
- the eZFP fusion protein comprises the sequence set forth in SEQ ID NO:621.
- the eZFP fusion protein comprises the sequence set forth in SEQ ID NO:640.
- the eZFP fusion protein comprises the sequence set forth in SEQ ID NO:658. In some embodiments, the eZFP fusion protein comprises the sequence set forth in SEQ ID NO:642. In some embodiments, the eZFP fusion protein comprises the sequence set forth in SEQ ID NO:660. In some embodiments, the eZFP fusion protein comprises the sequence set forth in SEQ ID NO:644. In some embodiments, the eZFP fusion protein comprises the sequence set forth in SEQ ID NO:662.
- CRISPR/Cas-based DNA-targeting systems are CRISPR/Cas-based DNA-targeting systems.
- the DNA-targeting systems comprise a DNA-targeting domain that binds to a target site in a regulatory DNA element of a FXN locus and comprises a Cas-guide RNA (gRNA) combination.
- the Cas-gRNA combination includes a variant Cas protein that lacks nuclease activity or that is a deactivated Cas (dCas) protein.
- the Cas- gRNA combination includes at least one gRNA comprising a gRNA spacer sequence that is capable of hybridizing to the target site or is complementary to the target site.
- the CRISPR/Cas-based DNA targeting systems comprise an epigenetic effector domain and/or multipartite effector for transcriptional activation, such as any of the effector domains or multipartite effectors described herein, such as in Section II.B.
- the CRISPR/Cas-based DNA-tareting system comprises a fusion protein, such as a dCas fusion protein, comprising the effector domains and/or multipartite effectors, such as any described in Section II.B.
- the DNA-targeting domain comprises a CRISPR-associated (Cas) protein or variant thereof, or comprises a protein that is derived from a Cas protein or variant thereof.
- the Cas protein is nuclease-inactive (i.e. is a dCas protein).
- DNA-targeting systems based on CRISPR/Cas systems, i.e. CRISPR/Cas-based DNA-targeting systems, that are able to bind to a target site in a FXN gene or regulatory DNA element thereof.
- the CRISPR/Cas DNA-targeting domain is nuclease inactive, such as includes a dCas (e.g. dCas9) so that the system binds to the target site in a target gene without mediating nucleic acid cleavage at the target site.
- the CRISPR/Cas-based DNA- targeting systems may be used to modulate expression of FXN in a cell.
- the CRISPR/Cas-based DNA-targeting system can include any known Cas enzyme, such as a nucleaseinactive or dCas.
- the CRISPR/Cas-based DNA-targeting system includes a fusion protein of a nuclease-inactive Cas protein or a variant thereof and an effector domain that increases transcription of a gene (e.g. a transcription activation domain), and at least one gRNA.
- the CRISPR system (also known as CRISPR/Cas system, or CRISPR-Cas system) refers to a conserved microbial nuclease system, found in the genomes of bacteria and archaea, that provides a form of acquired immunity against invading phages and plasmids.
- CRISPR Clustered Regularly Interspaced Short Palindromic Repeats
- spacers are short sequences of foreign DNA that are incorporated into the genome between CRISPR repeats, serving as a 'memory' of past exposures.
- Spacers encode the DNA-targeting portion of RNA molecules that confer specificity for nucleic acid cleavage by the CRISPR system.
- CRISPR loci contain or are adjacent to one or more CRISPR-associated (Cas) genes, which can act as RNA-guided nucleases for mediating the cleavage, as well as non-protein coding DNA elements that encode RNA molecules capable of programming the specificity of the CRISPR-mediated nucleic acid cleavage.
- Cas CRISPR-associated
- CRISPR/Cas systems such as those with Cas9, have been engineered to allow efficient programming of Cas/RNA RNPs to target desired sequences in cells of interest, both for gene-editing and modulation of gene expression.
- the tracrRNA and crRNA have been engineered to form a single chimeric guide RNA molecule, commonly referred to as a guide RNA (gRNA), for example as described in WO 2013/176772, WO 2014/093661, WO 2014/093655, Jinek et al. Science 337(6096):816-21 (2012), or Cong et al. Science 339(6121): 819-23 (2013), and as described herein, for example, in Section II.B.
- gRNA guide RNA
- CRISPR/Cas systems may be multi-protein systems or single effector protein systems. Multi-protein, or Class 1, CRISPR systems include Type I, Type III, and Type IV systems. In some aspects, Class 2 systems include a single effector molecule and include Type II, Type V, and Type VI. In some embodiments, the DNA targeting system comprises components of CRISPR/Cas systems, such as a Type I, Type II, Type III, Type IV, Type V, or Type VI CRISPR system.
- the Cas protein is from a Class 1 CRISPR system (i.e. multiple Cas protein system), such as a Type I, Type III, or Type IV CRISPR system.
- the Cas protein is from a Class 2 CRISPR system (i.e. single Cas protein system), such as a Type II, Type V, or Type VI CRISPR system.
- the Cas protein is derived from a Cas9 protein or variant thereof, for example as described in WO 2013/176772, WO 2014/152432, WO 2014/093661, WO 2014/093655, Jinek, M. et al. Science 337(6096):816-21 (2012), Mali, P. et al. Science 339(6121):823-6 (2013), Cong, L. et al. Science 339(6121):819-23 (2013), Perez-Pinera, P. et al. Nat. Methods 10, 973-976 (2013), or Mali, P. et al. Nat. Biotechnol. 31, 833-838 (2013).
- Type I CRISPR/Cas systems employ a large multisubunit ribonucleoprotein (RNP) complex called Cascade that recognizes double-stranded DNA (dsDNA) targets. After target recognition and verification, Cascade recruits the signature protein Cas3, a fused helicase-nuclease, to degrade DNA.
- RNP ribonucleoprotein
- the Cas protein is from a Type II CRISPR system.
- Exemplary Cas proteins of a Type II CRISPR system include Cas9.
- the Cas protein is from a Cas9 protein or variant thereof, for example as described in WO 2013/176772, WO 2014/152432, WO 2014/093661, WO 2014/093655, Jinek. et al. Science 337(6096):816-21 (2012), Mali et al. Science 339(6121):823-6 (2013), Cong et al. Science 339(6121):819-23 (2013), Perez-Pinera et al. Nat. Methods 10, 973-976 (2013), or Mali et al. Nat.
- RNA molecules and the Cas9 protein form a ribonucleoprotein (RNP) complex to direct Cas9 nuclease activity.
- the CRISPR RNA (crRNA) contains a spacer sequence that is complementary to a target nucleic acid sequence (target site), and that encodes the sequence specificity of the complex.
- the trans-activating crRNA (tracrRNA) base-pairs to a portion of the crRNA and forms a structure that complexes with the Cas9 protein, forming a Cas/RNA RNP complex.
- Cas9 mediates cleavage of target DNA if a correct protospacer-adjacent motif (PAM) is also present at the 3' end of the protospacer.
- PAM protospacer-adjacent motif
- the sequence must be immediately followed by the protospacer- adjacent motif (PAM), a short sequence recognized by the Cas9 nuclease that is required for DNA cleavage.
- the 5. pyogenes CRISPR system may have the PAM sequence for this Cas9 (SpCas9) as 5'-NRG-3', where R is either A or G, and characterized the specificity of this system in human cells.
- SpCas9 the PAM sequence for this Cas9
- a unique capability of the CRISPR/Cas9 system is the straightforward ability to simultaneously target multiple distinct genomic loci by coexpressing a single Cas9 protein with two or more sgRNAs.
- the Streptococcus pyogenes Type II system typically prefers to use an “NGG” sequence, where “N” can be any nucleotide, but also accepts other PAM sequences, such as “NAG” in engineered systems (Hsu et al., Nature Biotechnology (2013) doi:10.1038/nbt.2647).
- NmCas9 derived from Neisseria meningitidis
- NmCas9 normally has a native PAM of NNNNGATT (SEQ ID NO: 143), but has activity across a variety of PAMs, including a highly degenerate NNNNGNNN (SEQ ID NO:222) PAM (Esvelt et al.
- the Cas9 derived from Campylobacter jejuni typically uses 5'-NNNNACAC-3' (SEQ ID NO:226) or 5'-NNNNRYAC-3' (SEQ ID NO: 144) PAM sequences, where “N” can be any nucleotide, “R” can be either guanine (G) or adenine (A), and “Y” can be either cytosine (C) or thymine (T).
- the PAM sequences for spacer targeting depends on the type, ortholog, variant or species of the Cas protein.
- the Cas9 protein comprises a sequence from a Cas9 molecule of S. aureus.
- the Cas9 protein comprises a sequence set forth in SEQ ID NO:73 or SEQ ID NO:126, or a variant thereof, such as an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO:73 or SEQ ID NO: 126.
- the Cas9 protein comprises a sequence from a Cas9 molecule of S. pyogenes.
- the Cas9 protein comprises a sequence set forth in SEQ ID NO:79 or SEQ ID NO: 127, or a variant thereof, such as an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO:79 or SEQ ID NO: 127.
- the RNP complex is multimeric with a helicoid structure similar to Cascade.
- the Type III RNP complex recognizes complementary RNA sequences instead of dsDNA. RNA recognition stimulates a nonspecific DNA cleavage activity of the exemplary Type III CaslO nuclease that is part of the RNP complex, such that DNA cleavage is achieved cotranscriptionally.
- the Cas protein is from a Type V CRISPR system.
- Exemplary Cas proteins of a Type V CRISPR system include Casl2a (also known as Cpfl), Casl2b (also known as C2cl), Casl2e (also known as CasX), Casl2k (also known as C2c5), Casl4a, and Casl4b.
- the Cas protein is from a Casl2 protein (i.e. Cpfl) or variant thereof, for example as described in WO 2017/189308, WO2019/232069 and Zetsche et al. Cell. 163(3):759-71 (2015).
- Exemplary Type V systems include those based on a Casl2 effector, and the C-terminus with only one RuvC endonuclease domain is the defining characteristic of the Type V systems.
- the RuvC nuclease domain cleaves dsDNA adjacent to protospacer adjacent motif (PAM) sequences and singlestranded DNA (ssDNA) nonspecifically.
- PAM protospacer adjacent motif
- ssDNA singlestranded DNA
- the Type V systems can be further divided into subtypes, each characterized by different signature proteins, PAM sequences, and properties.
- Non-limiting exemplary Cas proteins derived from Type V CRISPR systems include Casl2a (Cpfl), UnlCasl2fl, Casl2j (CasPhi, such as CasPhi-2), Casl2k, and CasMini.
- Type V-A includes, for example, Casl2a, which uses “TTTV” (SEQ ID NO: 147) PAM sequence, where “V” is adenine (A), cytosine (C), or guanine (G).
- Type V-F is includes, for example, Casl2f, which can use “TTTR” (SEQ ID NO:228), where “R” is G or A, or “TTTN” (SEQ ID NO:225), where “N” is any nucleotide.
- Type V-K is includes, for example, Casl2k, which uses “GGTT” (SEQ ID NO:227) PAM sequence.
- the Casl2a protein comprises a sequence from a Casl2a molecule of Acidaminococcus sp, such as an AsCasl2a set forth in SEQ ID NO: 191 or SEQ ID NO: 192, or a variant thereof, such as an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO: 191 or SEQ ID NO: 192.
- Non-limiting examples of Cas proteins or Cas orthologs, such as Cas9 orthologs, from other bacterial strains include but are not limited to, Cas proteins identified in Acaryochloris marina MB IC 11017; Acetohalobium arabaticum DSM 5501; Acidaminococcus sp.; Acidithiobacillus caldus; Acidithiobacillus ferrooxidans ATCC 23270; Alicyclobacillus acidocaldarius LAA1; Alicyclobacillus acidocaldarius subsp.
- PCC 8005 Bacillus pseudomycoides DSM 12442; Bacillus selenitireducens MLS10; Burkholderiales bacterium 1_1_47; Caldicrudosiruptor becscii DSM 6725; Campylobacter jejuni; Candidatus Desulforudis audaxviator MP104C; Caldicellulosiruptor hydrothermalis 108; Clostridium phage c-st; Clostridium botulinum A3 str. Loch Maree; Clostridium botulinum Ba4 str. 657; Clostridium difficile QCD-63q42; Crocosphaera watsonii WH 8501; Cyanothece sp.
- PCC 6506 Pelotomaculum_thermopropionicum SI; Petrotoga mobilis SJ95; Polaromonas naphthalenivorans CJ2; Polaromonas sp. JS666; Pseudoalteromonas haloplanktis TAC125; Streptomyces pristinaespiralis ATCC 25486; Streptomyces pristinaespiralis ATCC 25486; Streptococcus thermophilus; Streptomyces viridochromogenes DSM 40736; Streptosporangium roseum DSM 43021; Synechococcus sp. PCC 7335; and Thermosipho africanus TCF52B (Chylinski et al., RNA Biol., 2013; 10(5): 726-737).
- the DNA-targeting systems or fusion proteins comprise a Cas protein, such as a Cas protein set forth in any one of SEQ ID NOS:73, 79, 126, 127, 193, 194, 197-200, and 205- 208, or a variant thereof, such as an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any one of SEQ ID NOS:73, 79, 126, 127, 193, 194, 197-200, and 205-208.
- a Cas protein such as a Cas protein set forth in any one of SEQ ID NOS:73, 79, 126, 127, 193, 194, 197-200, and 205-208.
- the Cas protein of any of the DNA-targeting systems or fusion proteins provided herein comprise a sequence set forth in any one of SEQ ID NOS:73, 79, 126, 127, 193, 194, 197-200, and 205-208, or a variant thereof, such as an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any one of SEQ ID NOS:73, 79, 126, 127, 193, 194, 197-200, and 205-208.
- the Cas protein lacks an initial methionine residue.
- the Cas protein comprises an initial methionine residue.
- the DNA- targeting domain e.g., Cas
- the DNA- targeting domain is a deactivated Cas (dCas), or a nuclease-inactive Cas (iCas).
- the component of the DNA-targeting domain such as a protein component, comprises a Cas9 variant such as a deactivated Cas9 or inactivated Cas9.
- the component of the DNA-targeting domain, such as a protein component comprises a Cas 12a variant such as a deactivated Casl2a (Cpfl) or inactivated Casl2a (Cpfl).
- the Cas9 protein may be mutated so that the nuclease activity is deactivated or inactivated (also referred to as dCas9 or iCas9).
- the Cas protein is a variant that lacks nuclease activity (i.e. is a dCas or iCas protein).
- the Cas protein is mutated so that nuclease activity is reduced or eliminated.
- Such Cas proteins are referred to as deactivated Cas or dead Cas (dCas) or nuclease-inactive Cas (iCas) proteins, as referred to interchangeably herein.
- the variant Cas protein is a variant Cas9 protein that lacks nuclease activity or that is a deactivated Cas9 (dCas9, or iCas9) protein.
- the variant Cas protein is a variant Cpfl protein that lacks nuclease activity or that is a deactivated Casl2a (dCasl2a, or iCasl2a) protein.
- Cas proteins are engineered to be catalytically inactivated or nuclease inactive to allow targeting of Cas/gRNA RNPs without inducing cleavage at the target site. Mutations in Cas proteins can reduce or abolish nuclease activity of the Cas protein, rendering the Cas protein catalytically inactive. Cas proteins with reduced or abolished nuclease activity are referred to as deactivated Cas (dCas), or nuclease-inactive Cas (iCas) proteins, as referred to interchangeably herein.
- dCas deactivated Cas
- iCas nuclease-inactive Cas
- the dCas or iCas can still bind to target site in the DNA in a site- and/or sequence-specific manner, as long as it retains the ability to interact with the guide RNA (gRNA) which directs the Cas- gRNA combination to the target site.
- gRNA guide RNA
- the dCas or iCas exhibits reduced or no endodeoxyribonuclease activity.
- an exemplary dCas or iCas for example dCas9 or iCas9, exhibits less than about 20%, less than about 15%, less than about 10%, less than about 5%, less than about 1%, or less than about 0.1%, of the endodeoxyribonuclease activity of a wild-type Cas protein, e.g., a wild-type Cas9 protein.
- the dCas or iCas exhibits substantially no detectable endodeoxyribonuclease activity.
- an exemplary dCas or iCas for example dCas9 or iCas9, comprises one or more amino acid mutations, substitutions, deletions or insertions at a position corresponding to a position selected from D10, G12, G17, E762, H840, N854, N863, H982, H983, A984, D986, and/or a A987, with reference to a wild-type Streptococcus pyogenes Cas9 (SpCas9), for example, with reference to numbering of positions of a SpCas9 sequence set forth in SEQ ID NO:79.
- SpCas9 wild-type Streptococcus pyogenes Cas9
- the dCas9 or iCas9 comprises one or more amino acid mutations, substitutions, deletions or insertions corresponding to D10A, G12A, G17A, E762A, H840A, N854A, N863A, H982A, H983A, A984A, and/or D986A, with reference to a wild-type Streptococcus pyogenes Cas9 (SpCas9), for example, with reference to numbering of positions of a SpCas9 sequence set forth in SEQ ID NO:79.
- SpCas9 wild-type Streptococcus pyogenes Cas9
- dCas protein lacks an initial methionine residue. In some aspects, the dCas protein comprises an initial methionine residue.
- the dCas9 protein can comprise a sequence from a Cas9 molecule, or variant thereof. In some embodiments, the dCas9 protein can comprise a sequence derived from a Cas9 molecule of .S’, pyogenes, S. thermophilus, S. aureus, N. meningitidis, F. novicida, S. canis, S. auricularis, or variant thereof. In some embodiments, the dCas9 protein comprises a sequence from a Cas9 molecule of .S', aureus. In some embodiments, the dCas9 protein comprises a sequence from a Cas9 molecule of .S'. pyogenes. In some embodiments, the dCas9 protein comprises a sequence from a Cas9 molecule of C. jejuni.
- Exemplary deactivated Cas9 (dCas9) derived from .S'. pyogenes contains silencing mutations of the RuvC and HNH nuclease domains (D10A and H840A), for example as described in WO 2013/176772, WO 2014/093661, Jinek et al. Science 337(6096):816-21 (2012), and Qi et al. Cell 152(5): 1173-83 (2013).
- Exemplary dCas variants derived from the Casl2 system i.e. Cpfl
- WO 2017/189308 and Zetsche et al. Cell 163(3):759-71 (2015).
- Cas orthologs conserved domains that mediate nucleic acid cleavage, such as RuvC and HNH endonuclease domains, are readily identifiable in Cas orthologs, and can be mutated to produce inactive variants, for example as described in Zetsche et al. Cell 163(3):759-71 (2015).
- Other exemplary Cas orthologs or variants include engineered variants based on a Casl2f (also known as Casl4), including those described in Xu et al., Mol. Cell 81(20):4333-4345 (2021).
- the DNA-targeting domain comprises a Cas-gRNA combination that includes (a) a Cas protein or a variant thereof and (b) at least one gRNA.
- the variant Cas protein lacks nuclease activity or is a deactivated Cas (dCas) protein.
- the gRNA is capable of complexing with the Cas protein or variant thereof.
- the gRNA comprises a gRNA spacer sequence that is capable of hybridizing to the target site or is complementary to the target site (e.g., in a FXN locus).
- the Cas protein or a variant thereof is a Cas9 protein or a variant thereof.
- the variant Cas protein is a variant Cas9 protein that lacks nuclease activity or that is a deactivated Cas9 (dCas9) protein.
- the Cas9 protein or a variant thereof is a Staphylococcus aureus Cas9 (SaCas9) protein or a variant thereof.
- the variant Cas9 is a Staphylococcus aureus dCas9 protein (dSaCas9) that comprises at least one amino acid mutation selected from D10A and N580A, with reference to numbering of positions of SEQ ID NO:73.
- the variant Cas9 protein comprises the sequence set forth in SEQ ID NO:72, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the variant Cas9 protein comprises the sequence set forth in SEQ ID NO:72, which lacks an initial methionine residue.
- the variant Cas9 protein comprises the sequence set forth in SEQ ID NO: 189, which includes an initial methionine residue.
- An exemplary nucleotide sequence encoding the variant Cas9 protein is set forth in SEQ ID NO:70.
- the Cas9 protein or variant thereof is a Streptococcus pyogenes Cas9 (SpCas9) protein or a variant thereof.
- the variant Cas9 is a Streptococcus pyogenes dCas9 (dSpCas9) protein that comprises at least one amino acid mutation selected from D10A and H840A, with reference to numbering of positions of SEQ ID NO:79.
- the variant Cas9 protein comprises the sequence set forth in SEQ ID NO:78, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the variant Cas9 protein comprises the sequence set forth in SEQ ID NO:78, which lacks an initial methionine residue. In some embodiments, the variant Cas9 protein comprises the sequence set forth in SEQ ID NO: 190, which includes an initial methionine residue.
- An exemplary nucleotide sequence encoding the variant Cas9 protein is set forth in SEQ ID NO:76.
- the Cas9 protein or variant thereof is a Campylobacter jejuni Cas9 (CjCas9) protein or a variant thereof.
- the variant Cas9 comprises at least one amino acid mutation compared to the sequence set forth in SEQ ID NO:205 or 206.
- the variant Cas9 protein comprises the sequence set forth in SEQ ID NO: 203, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the Cas protein or a variant thereof is a Casl2a protein or a variant thereof.
- the variant Cas protein is a variant Cas 12a protein that lacks nuclease activity or that is a deactivated Casl2a (dCasl2a) protein.
- the Casl2a protein or variant thereof is a Acidaminococcus sp.
- the variant Casl2a is a Acidaminococcus sp.
- the variant Casl2a protein comprises the sequence set forth in SEQ ID NO: 192, which lacks an initial methionine residue. In some embodiments, the variant Casl2a protein comprises the sequence set forth in SEQ ID NO:191, which includes an initial methionine residue.
- the Cas protein or a variant thereof is a CasPhi-2 protein or a variant thereof.
- the variant Cas protein is a variant CasPhi-2 protein that lacks nuclease activity or that is a deactivated CasPhi-2 (dCasPhi-2) protein.
- the variant CasPhi-2 comprises at least one amino acid mutation compared to the sequence set forth in SEQ ID NO: 197 or 198.
- the variant CasPhi-2 protein comprises the sequence set forth in SEQ ID NO: 221, which lacks an initial methionine residue. In some embodiments, the variant CasPhi-2 protein comprises the sequence set forth in SEQ ID NO: 195, which includes an initial methionine residue. In some embodiments, the variant CasPhi-2 protein comprises the sequence set forth in SEQ ID NO:220, which includes an initial methionine residue.
- the Cas protein or a variant thereof is a UnlCasl2fl protein or a variant thereof.
- the variant Cas protein is a variant UnlCasl2fl protein that lacks nuclease activity or that is a deactivated UnlCasl2fl (dUnlCasl2fl) protein.
- the variant UnlCasl2fl comprises at least one amino acid mutation compared to the sequence set forth in SEQ ID NO: 199 or 200.
- the variant UnlCasl2fl protein comprises the sequence set forth in SEQ ID NO:201, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the variant UnlCasl2fl protein comprises the sequence set forth in SEQ ID NO:202, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the variant UnlCasl2fl protein comprises the sequence set forth in SEQ ID NO:202, which lacks an initial methionine residue. In some embodiments, the variant UnlCasl2fl protein comprises the sequence set forth in SEQ ID NO:201, which includes an initial methionine residue.
- the Cas protein or a variant thereof is a Cas 12k protein or a variant thereof.
- the Casl2k protein comprises the sequence set forth in SEQ ID NO:207, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the Casl2k protein comprises the sequence set forth in SEQ ID NO:208, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the Casl2k protein comprises the sequence set forth in SEQ ID NO:208, which lacks an initial methionine residue. In some embodiments, the Casl2k protein comprises the sequence set forth in SEQ ID NO:207, which includes an initial methionine residue.
- the Cas protein or a variant thereof is a CasMini protein or a variant thereof, such as an engineered Cas protein or variant based on a Casl2f (also known as Casl4), including those described in Xu et al., Mol. Cell 81(20):4333-4345 (2021) or set forth in SEQ ID NO:223.
- the variant Cas protein is a variant CasMini protein that lacks nuclease activity or that is a deactivated CasMini (dCasMini) protein.
- the variant CasMini comprises at least one amino acid mutation compared to the sequence set forth in SEQ ID NO:223.
- the variant CasMini protein comprises the sequence set forth in SEQ ID NO:223, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the CasMini protein comprises the sequence set forth in SEQ ID NO: 223.
- the variant CasMini protein comprises the sequence set forth in SEQ ID NO: 209 or 210, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the CasMini protein comprises the sequence set forth in SEQ ID NO:209, which lacks an initial methionine residue. In some embodiments, the CasMini protein comprises the sequence set forth in SEQ ID NO:210, which includes an initial methionine residue.
- DNA-targeting systems in some cases comprising a fusion protein, such as dCas-fusion proteins include fusion of the Cas with an effector domain, such as a transcription activation domain.
- an effector domain such as a transcription activation domain.
- a DNA-targeting system comprising a fusion protein comprising a DNA-targeting domain comprising a nuclease-inactive Cas protein or variant thereof, and an effector domain for increasing or inducing transcriptional activation (i.e. a transcriptional activator) when targeted to a target site in a FXN gene or regulatory element thereof.
- the DNA-targeting system also includes one or more gRNA, provided in combination or as a complex with the dCas protein or variant thereof, for targeting of the DNA-targeting system to the target site.
- the fusion protein is guided to a specific target site sequence of the target gene by the guide RNA, wherein the effector domain mediates targeted epigenetic modification to increase or promote transcription of the target gene.
- gRNAs Guide RNAs
- the gRNA is capable of complexing with the Cas protein or variant thereof.
- the gRNA comprises a gRNA spacer sequence (also known as a spacer sequence or a guide sequence) that is capable of hybridizing to the target site or is complementary to the target site, such as any target site described herein, for example, any target site in a genome.
- the gRNA comprises a scaffold sequence that complexes with or binds to the Cas protein.
- a gRNA specific to a target locus of interest e.g. a regulatory DNA element of a FXN locus
- a target locus of interest e.g. a regulatory DNA element of a FXN locus
- an RNA-guided protein e.g. a Cas protein
- a fusion protein comprising such RNA-guided protein (e.g., a Cas polypeptide)
- the Cas protein e.g. dCas9 is provided in combination or as a complex with one or more guide RNA (gRNA).
- gRNA guide RNA
- the gRNA is a nucleic acid that promotes the specific targeting or homing of the gRNA/Cas RNP complex to the target site, such as any described above.
- a target site of a gRNA may be referred to as a protospacer.
- gRNAs such as gRNAs that target or bind to a target site in a FXN gene or DNA regulatory element thereof, such as any described above in Section ILA.
- the gRNA is capable of complexing with the Cas protein or variant thereof.
- the gRNA comprises a gRNA spacer sequence (i.e. a spacer sequence or a guide sequence) that is capable of hybridizing to the target site, or that is complementary to the target site, such as any target site described in Section II. A or further below.
- the gRNA comprises a scaffold sequence that complexes with or binds to the Cas protein.
- a “gRNA molecule” is a nucleic acid that promotes the specific targeting or homing of a gRNA molecule/Cas9 molecule complex to a target nucleic acid, such as a locus on the genomic DNA of a cell.
- gRNA molecules can be unimolecular (having a single RNA molecule), sometimes referred to herein as “chimeric” gRNAs, or modular (comprising more than one, and typically two, separate RNA molecules).
- a spacer sequence of the guide RNA is any polynucleotide sequences comprising at least a sequence portion that has sufficient complementarity with a target polynucleotide sequence, such as the at the FXN locus in humans, to hybridize with the target sequence at the target site and direct sequence-specific binding of the CRISPR complex to the target sequence.
- target sequence is to a sequence to which a spacer sequence is designed to have complementarity, where hybridization between the target sequence and a spacer sequence of the guide RNA promotes the formation of a CRISPR complex.
- a spacer sequence is selected to reduce the degree of secondary structure within the spacer sequence. Secondary structure may be determined by any suitable polynucleotide folding algorithm.
- a guide RNA specific to a target locus of interest (e.g. at the FXN locus in humans) is used with RNA-guided nucleases or variants thereof, e.g., nuclease-inactive Cas variants, to target the provided DNA-targeting system to the target site or target position.
- RNA-guided nucleases or variants thereof e.g., nuclease-inactive Cas variants
- Methods for designing gRNAs and exemplary spacer sequences are known.
- Exemplary gRNA structures that can be associated with particular RNA-guided nucleases or variants thereof, e.g., nuclease-inactive Cas variants, with particular domains and scaffold regions are also known.
- gRNA molecules comprise a scaffold sequence, e.g., sequences that can be complexed with the Cas protein.
- the scaffold sequence is specific for the Cas protein.
- the gRNA is a chimeric gRNA.
- gRNAs can be unimolecular (i.e. composed of a single RNA molecule), or modular (comprising more than one, and typically two, separate RNA molecules).
- Modular gRNAs can be engineered to be unimolecular, wherein sequences from the separate modular RNA molecules are comprised in a single gRNA molecule, sometimes referred to as a chimeric gRNA, synthetic gRNA, or single gRNA.
- a guide RNA can comprise at least a spacer sequence that hybridizes to a target nucleic acid sequence of interest, and a CRISPR repeat sequence.
- the gRNA also comprises a second RNA called the tracrRNA sequence.
- the CRISPR repeat sequence and tracrRNA sequence hybridize to each other to form a duplex.
- the crRNA forms a duplex.
- the duplex can bind a site-directed polypeptide, such that the guide RNA and site-direct polypeptide form a complex.
- the gRNA can provide target specificity to the complex by virtue of its association with the site-directed polypeptide. The gRNA thus can direct the activity of the site-directed polypeptide.
- the chimeric gRNA is a fusion of two non-coding RNA sequences: a crRNA sequence and a tracrRNA sequence, for example as described in WO 2013/176772, or Jinek, M. et al. Science 337(6096):816-21 (2012).
- the chimeric gRNA mimics the naturally occurring crRNA:tracrRNA duplex involved in the Type II CRISPR/Cas system, wherein the naturally occurring crRNA:tracrRNA duplex acts as a guide for the Cas protein, e.g., Cas9 protein.
- Exemplary types of CRISPR/Cas systems and associated gRNA structures include those described in, for example, Moon et al. Exp.
- the spacer sequence of a gRNA is a polynucleotide sequence comprising at least a portion that has sufficient complementarity with the target site to hybridize with the target site and direct sequence-specific binding of a CRISPR complex to the sequence of the target site. Full complementarity is not necessarily required, provided there is sufficient complementarity to cause hybridization and promote formation of a CRISPR complex.
- the gRNA comprises a spacer sequence that is complementary, e.g., at least 80%, 85%, 90%, 95%, 98%, 99%, or 100% (e.g., fully complementary), to the target site.
- the strand of the target nucleic acid comprising the target site sequence may be referred to as the “complementary strand” of the target nucleic acid.
- the spacer sequence is a user-defined sequence. Guidance on the selection of spacer sequences can be found, e.g., in Fu et al., Nat Biotechnol 2014 32:279-284 and Sternberg et al., Nature 2014 507:62-67.
- the gRNA spacer sequence is between about 14 nt and about 26 nt, between about 14 nt and about 24 nt, or between 16 nt and 22 nt in length. In some embodiments, the gRNA spacer sequence is 14 nt, 15 nt, 16 nt, 17 nt, 18 nt, 19 nt, 20 nt, 21 nt or 22 nt, 23 nt, 24 nt, 25 nt, or 26 nt in length. In some embodiments, the gRNA spacer sequence is 18 nt, 19 nt, 20 nt, 21 nt or 22 nt in length.
- the gRNA spacer sequence is 18 nt in length. In some embodiments, the gRNA spacer sequence is 19 nt in length. In some embodiments, the gRNA spacer sequence is 20 nt in length. In some embodiments, the gRNA spacer sequence is 21 nt in length. In some embodiments, the gRNA spacer sequence is 22 nt in length.
- Methods for designing gRNAs and exemplary targeting domains can include those described in, e.g., International PCT Pub. Nos. WO 2014/197748, WO 2016/130600, WO 2017/180915, WO 2021/226555, WO 2013/176772, WO 2014/152432, WO 2014/093661, WO 2014/093655, WO 2015/089427, WO 2016/049258, WO 2016/123578, WO 2021/076744, WO 2014/191128, WO 2015/161276, WO 2017/193107, and WO 2017/093969.
- a target site of a gRNA may be referred to as a protospacer.
- the spacer is designed to target a protospacer with a specific protospacer-adjacent motif (PAM), i.e. a sequence immediately adjacent to the protospacer that contributes to and/or is required for Cas binding specificity.
- PAM protospacer-adjacent motif
- Different CRISPR/Cas systems have different PAM requirements for targeting.
- 5. pyogenes Cas9 uses the PAM 5’-NGG-3’ (SEQ ID NO:142), where N is any nucleotide. 5.
- aureus Cas9 uses the PAM 5’- NNGRRT-3’ (SEQ ID NO: 143), where N is any nucleotide, and R is G or A.
- N. meningitidis Cas9 uses the PAM 5'-NNNNGATT -3’ (SEQ ID NO: 144), where N is any nucleotide.
- C. jejuni Cas9 uses the PAM 5'-NNNNRYAC-3' (SEQ ID NO: 145) or 5'-NNNNACAC- 3 ’(SEQ ID NO:226), where N is any nucleotide, R is G or A, and Y is C or T. S.
- thermophilus uses the PAM 5’-NNAGAAW-3’ (SEQ ID NO: 146), where N is any nucleotide and W is A or T.
- F. Novicida Cas9 uses the PAM 5’-NGG-3’ (SEQ ID NO: 142), where N is any nucleotide.
- T. denticola Cas9 uses the PAM 5’-NAAAAC-3’ (SEQ ID NO: 146), where N is any nucleotide.
- Casl2a also known as Cpfl
- Casl2a from various species, uses the PAM 5’-TTTV-3’ (SEQ ID NO:148), where V is A, C, or G.
- Phage-derived CasPhi (such as CasPhi-2, also known as Casl2j), uses the PAM 5’-TBN-3’ (SEQ ID NO:224), where N is any nucleotide, and B is G, T, or C.
- Archaeal UnlCasl2fl (also known as Casl4al), uses the PAM 5’- TTTN -3’ (SEQ ID NO:225), where N is any nucleotide.
- a Casl2f protein (also known as Casl4) uses the PAM 5’- TTTR -3’ (SEQ ID NO:228), where R is G or A.
- a Casl2k protein uses the PAM 5’- GGTT -3’ (SEQ ID NO:227).
- Cas proteins may use or be engineered to use different PAMs from those listed above.
- variant SpCas9 proteins may use a PAM selected from: 5’-NGG-3’ (SEQ ID NO:142), 5’-NGAN-3’ (SEQ ID NO:149), 5’-NGNG-3’(SEQ ID NO:15Q), 5’-NGAG-3’(SEQ ID NO:151), or 5’-NGCG-3’(SEQ ID NO:152), where N is any nucleotide.
- Methods for designing or identifying gRNA spacer sequences and/or protospacer sequences in a particular region are known.
- gRNA spacer sequences and/or protospacer sequences can be determined based on the type of Cas protein used and the associated PAM sequence.
- the PAM of a gRNA for complexing with .S', pyogenes Cas9 or variant thereof is set forth in SEQ ID NO: 141.
- the PAM of a gRNA for complexing with .S', aureus Cas9 or variant thereof is set forth in SEQ ID NO: 142.
- the PAM of a gRNA for complexing with a Type V CRISPR/Cas system, such as with Casl2a (also known as Cpfl) or variant thereof is set forth in SEQ ID NO: 147.
- a spacer sequence may be selected to reduce the degree of secondary structure within the spacer sequence.
- Secondary structure may be determined by any suitable polynucleotide folding algorithm.
- the gRNA (including the spacer sequence) will comprise the base uracil (U), whereas DNA encoding the gRNA molecule will comprise the base thymine (T). While not wishing to be bound by theory, in some embodiments, it is believed that the complementarity of the spacer sequence (i.e. guide sequence) with the target sequence contributes to specificity of the interaction of the gRNA molecule/Cas molecule complex with a target nucleic acid. It is understood that in a spacer sequence (i.e. guide sequence) and target sequence pair, the uracil bases in the spacer sequence (i.e. guide sequence) will pair with the adenine bases in the target sequence.
- a gRNA spacer sequence herein may be defined by the DNA sequence encoding the gRNA spacer, and/or the RNA sequence of the spacer.
- the gRNA comprises modified nucleotides, e.g. for increased stability.
- one, more than one, or all of the nucleotides of a gRNA can have a modification, e.g., to render the gRNA less susceptible to degradation and/or improve bio-compatibility.
- the backbone of the gRNA can be modified with a phosphorothioate, or other modification(s).
- a nucleotide of the gRNA can comprise a 2’ modification, e.g., a 2- acetylation, e.g., a 2’ methylation, or other modification(s)
- the gRNA is a concatenation of two non-coding RNA sequences: a crRNA sequence and a tracrRNA sequence.
- the gRNA may target a desired DNA sequence by exchanging the sequence encoding a 20 bp protospacer which confers targeting specificity through complementary base pairing with the desired DNA target.
- gRNA mimics the naturally occurring crRNA:tracrRNA duplex involved in the Type II CRISPR/Cas system (e.g., Cas9).
- This duplex which may include, for example, a 42-nucleotide crRNA and a 75-nucleotide tracrRNA, acts as a guide for the Cas9 protein to cleave the target nucleic acid.
- target region refers to the region of the target gene to which the CRISPR/Cas9-based system targets.
- the CRISPR/Cas9-based system may include two or more gRNAs, wherein the two or more gRNAs target different DNA sequences.
- the target DNA sequences may be overlapping or nonoverlapping.
- the target DNA sequences may be located within or near the same gene or different genes.
- the target sequence or protospacer is followed by a PAM sequence at the 3' end of the protospacer.
- Different Type II systems have differing PAM requirements.
- the Streptococcus pyogenes Type II system uses an “NGG” sequence, where “N” can be any nucleotide.
- the gRNA comprises scaffold sequences.
- the scaffold sequence in some cases including a crRNA sequence and/or a tracrRNA sequence
- different CRISPR/Cas systems have different gRNA scaffold sequences for associating with Cas protein.
- an exemplary scaffold sequence for .S', aureus Cas9 comprises a sequence set forth in SEQ ID NO:44, or a sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to SEQ ID NO:44.
- an exemplary scaffold sequence for .S'. aureus Cas9 comprises a sequence set forth in SEQ ID NO:44.
- an exemplary scaffold sequence for .S', pyogenes Cas9 comprises a sequence set forth in SEQ ID NO:46, or a sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to SEQ ID NO:46.
- an exemplary scaffold sequence for .S', pyogenes Cas9 comprises a sequence set forth in SEQ ID NO:46.
- Casl2a comprises a sequence set forth in SEQ ID NO:211, or a sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to SEQ ID NO:211.
- an exemplary scaffold sequence for CasPhi-2 comprises a sequence set forth in SEQ ID NO:212, or a sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to SEQ ID NO:212.
- an exemplary scaffold sequence for UnlCasl2fl comprises a sequence set forth in SEQ ID NO:213, 214 or 215, or a sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to SEQ ID NO:213, 214 or 215.
- an exemplary scaffold sequence for UnlCasl2fl comprises a sequence set forth in SEQ ID NO:213, or a sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to SEQ ID NO:213.
- an exemplary scaffold sequence for UnlCasl2fl comprises a sequence set forth in SEQ ID NO:214, or a sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to SEQ ID NO:214.
- an exemplary scaffold sequence for UnlCasl2fl comprises a sequence set forth in SEQ ID NO:215, or a sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to SEQ ID NO:215.
- an exemplary scaffold sequence for C comprises a sequence set forth in SEQ ID NO:215, or a sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to SEQ ID NO:215.
- an exemplary scaffold sequence for C comprises a sequence set forth in SEQ ID NO:215, or a sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%,
- jejuni Cas9 comprises a sequence set forth in SEQ ID NO:216, or a sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to SEQ ID NO:216.
- an exemplary scaffold sequence for Cas 12k comprises a sequence set forth in SEQ ID NO:217, or a sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to SEQ ID NO:217.
- an exemplary scaffold sequence for CasMini comprises a sequence set forth in SEQ ID NO:218, or a sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to SEQ ID NO:218.
- a guide RNA that binds a target site in an enhancer region of a frataxin (FXN) locus, wherein the target site is located within the genomic coordinates human genome assembly GRCh38 (hg38) chr9:69, 027, 282-69, 028, 497.
- a guide RNA that binds a target site in an enhancer region of a frataxin (FXN) locus, wherein the target site is located within the genomic coordinates hg38 chr9:69,027,615- 69,028,101.
- the target site comprises the sequence set forth in SEQ ID NO:21, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing.
- DNA encoding the gRNA comprises the sequence set forth in SEQ ID NO:21, a contiguous portion thereof of at least 14 nucleotides, or a complementary sequence of any of the foregoing.
- the gRNA comprises at least one gRNA spacer comprising the sequence set forth in SEQ ID NO:42, or a contiguous portion thereof of at least 14 nt.
- the gRNA further comprises the sequence set forth in SEQ ID NO:44.
- the gRNA comprises the sequence set forth in SEQ ID NO:67.
- gRNA guide RNA
- FXN frataxin locus
- the target site comprises a sequence selected from any one of SEQ ID NOS: 1-10, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing.
- the gRNA comprises at least one gRNA spacer sequence comprising a sequence selected from any one of SEQ ID NOS:22-31, or a contiguous portion thereof of at least 14 nt. In some of any of the provided embodiments, the gRNA further comprises the sequence set forth in SEQ ID NO:44. In some of any of the provided embodiments, the gRNA comprises a sequence selected from any one of SEQ ID NOS:47-56.
- gRNA guide RNA
- FXN frataxin locus
- the target site comprises a sequence selected from any one of SEQ ID NOS: 11-20, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing.
- the gRNA comprises at least one gRNA spacer sequence comprising a sequence selected from any one of SEQ ID NOS:32-41, or a contiguous portion thereof of at least 14 nt. In some of any of the provided embodiments, the gRNA further comprises the sequence set forth in SEQ ID NO:46. In some of any of the provided embodiments, the gRNA comprises a sequence selected from any one of SEQ ID NOS:57-66.
- the DNA-targeting domain comprises a Cas-gRNA combination comprising (a) a Cas protein or a variant thereof and (b) at least one gRNA; and the gRNA comprises at least one gRNA spacer sequence comprising a sequence selected from any one of SEQ ID NOS:22-31, or a contiguous portion thereof of at least 14 nt.
- the gRNA further comprises the sequence set forth in SEQ ID NO:44.
- the gRNA comprises a sequence selected from any one of SEQ ID NOS:47-56.
- the DNA-targeting domain comprises a Cas-gRNA combination comprising (a) a Cas protein or a variant thereof and (b) at least one gRNA; and the gRNA comprises at least one gRNA spacer sequence comprising a sequence selected from any one of SEQ ID NOS:32-41, or a contiguous portion thereof of at least 14 nt.
- the gRNA further comprises the sequence set forth in SEQ ID NO:46.
- the gRNA comprises a sequence selected from any one of SEQ ID NOS:57-66.
- the gRNA comprises a sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to all or a portion of a gRNA sequence or a gRNA spacer sequence described herein.
- the gRNA targets a target site in a FXN locus or a DNA regulatory element thereof that comprises the sequence selected from any one of SEQ ID NO:1-10 and 21, a contiguous portion thereof of at least 14 nucleotides (e.g., 14, 15, 16, 17, 18, 19, 20, 21, or 22 nucleotides), a complementary sequence of any of the foregoing, or a sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to any of the foregoing.
- a target site in a FXN locus or a DNA regulatory element thereof that comprises the sequence selected from any one of SEQ ID NO:1-10 and 21, a contiguous portion thereof of at least 14 nucleotides (e.g., 14, 15, 16, 17, 18, 19, 20, 21, or 22 nucleotides), a complementary sequence of any of the foregoing, or a sequence having at or at least
- the gRNA comprises a spacer sequence comprising the sequence selected from any one of SEQ ID NO:22-31 and 42, a contiguous portion thereof of at least 14 nt (e.g., 14, 15, 16, 17, 18, 19, 20, 21, or 22 nucleotides), or a sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to any of the foregoing.
- the gRNA further comprises a scaffold sequence.
- the scaffold sequence comprises the sequence set forth in SEQ ID NO:44, or a sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to SEQ ID NO:44.
- the gRNA, including a spacer sequence and a scaffold sequence comprises the sequence selected from any one of SEQ ID NO:47-56 and 67, or a sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to all or a portion thereof.
- the gRNA is set forth in the sequence selected from any one of SEQ ID NO:47-56 and 67.
- a provided DNA-targeting system comprises any of the aforementioned gRNAs complexed with a Cas protein, such as a Cas9 protein.
- the Cas9 is a dCas9.
- the dCas9 is a dSaCas9, such as a dSaCas9 set forth in SEQ ID NO:72, or a variant and/or fusion thereof.
- the gRNA targets a target site in a FXN locus or a DNA regulatory element thereof that comprises the sequence selected from any one of SEQ ID NO:229-255, a contiguous portion thereof of at least 14 nucleotides (e.g., 14, 15, 16, 17, 18, 19, 20, 21, or 22 nucleotides), a complementary sequence of any of the foregoing, or a sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to any of the foregoing.
- the gRNA further comprises a scaffold sequence.
- the scaffold sequence comprises the sequence set forth in SEQ ID NO:211, or a sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to SEQ ID NO:211.
- the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:229, and a scaffold sequence of SEQ ID NO:211.
- the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:230, and a scaffold sequence of SEQ ID NO:211.
- the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:231, and a scaffold sequence of SEQ ID NO:211. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:232, and a scaffold sequence of SEQ ID NO:211. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:233, and a scaffold sequence of SEQ ID NO:211. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:234, and a scaffold sequence of SEQ ID NO:211.
- the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:235, and a scaffold sequence of SEQ ID NO:211. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:236, and a scaffold sequence of SEQ ID NO:211. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:237, and a scaffold sequence of SEQ ID NO:211. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:238, and a scaffold sequence of SEQ ID NO:211.
- the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:239, and a scaffold sequence of SEQ ID NO:211. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:240, and a scaffold sequence of SEQ ID NO:211. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:241, and a scaffold sequence of SEQ ID NO:211. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:242, and a scaffold sequence of SEQ ID NO:211.
- the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:243, and a scaffold sequence of SEQ ID NO:211. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:244, and a scaffold sequence of SEQ ID NO:211. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:245, and a scaffold sequence of SEQ ID NO:211. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:246, and a scaffold sequence of SEQ ID NO:211.
- the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:247, and a scaffold sequence of SEQ ID NO:211. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:248, and a scaffold sequence of SEQ ID NO:211. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:249, and a scaffold sequence of SEQ ID NO:211. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:250, and a scaffold sequence of SEQ ID NO:211.
- the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:251, and a scaffold sequence of SEQ ID NO:211. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:252, and a scaffold sequence of SEQ ID NO:211. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:253, and a scaffold sequence of SEQ ID NO:211. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:254, and a scaffold sequence of SEQ ID NO:211.
- the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:255, and a scaffold sequence of SEQ ID NO:211.
- a provided DNA-targeting system comprises any of the aforementioned gRNAs complexed with a Cas protein, such as a Cast 2a (also known as Cpfl) protein.
- the Casl2a is a dCasl2a.
- the dCasl2a is a dSaCasl2a, such as a dSaCasl2a set forth in SEQ ID NO:192, or a variant and/or fusion thereof.
- the gRNA targets a target site in a FXN locus that comprises SEQ ID NO:1, a contiguous portion thereof of at least 14 nucleotides, a complementary sequence of any of the foregoing, or a sequence having at least 90% sequence identity to any of the foregoing.
- the gRNA comprises a spacer sequence comprising SEQ ID NO:22, a contiguous portion thereof of at least 14 nt, or a sequence having at least 90% sequence identity to any of the foregoing.
- the gRNA further comprises a scaffold sequence comprising SEQ ID NO:44, or a sequence having at least 90% sequence identity to SEQ ID NO:44.
- the gRNA including a spacer sequence and a scaffold sequence, comprises SEQ ID NO:47, or a sequence having at least 90% sequence identity to all or a portion thereof.
- the gRNA is set forth in SEQ ID NO:47.
- a provided DNA-targeting system comprises any of the above gRNAs complexed with a Cas protein, such as a Cas9 protein.
- the Cas9 is a dCas9.
- the dCas9 is a dSaCas9, such as a dSaCas9 set forth in SEQ ID NO:72, or a variant and/or fusion thereof.
- the gRNA targets a target site in a FXN locus that comprises SEQ ID NO:2, a contiguous portion thereof of at least 14 nucleotides, a complementary sequence of any of the foregoing, or a sequence having at least 90% sequence identity to any of the foregoing.
- the gRNA comprises a spacer sequence comprising SEQ ID NO:23, a contiguous portion thereof of at least 14 nt, or a sequence having at least 90% sequence identity to any of the foregoing.
- the gRNA further comprises a scaffold sequence comprising SEQ ID NO:44, or a sequence having at least 90% sequence identity to SEQ ID NO:44.
- the gRNA including a spacer sequence and a scaffold sequence, comprises SEQ ID NO:48, or a sequence having at least 90% sequence identity to all or a portion thereof.
- the gRNA is set forth in SEQ ID NO:48.
- a provided DNA-targeting system comprises any of the above gRNAs complexed with a Cas protein, such as a Cas9 protein.
- the Cas9 is a dCas9.
- the dCas9 is a dSaCas9, such as a dSaCas9 set forth in SEQ ID NO:72, or a variant and/or fusion thereof.
- the gRNA targets a target site in a FXN locus that comprises SEQ ID NO:3, a contiguous portion thereof of at least 14 nucleotides, a complementary sequence of any of the foregoing, or a sequence having at least 90% sequence identity to any of the foregoing.
- the gRNA comprises a spacer sequence comprising SEQ ID NO:24, a contiguous portion thereof of at least 14 nt, or a sequence having at least 90% sequence identity to any of the foregoing.
- the gRNA further comprises a scaffold sequence comprising SEQ ID NO:44, or a sequence having at least 90% sequence identity to SEQ ID NO:44.
- the gRNA including a spacer sequence and a scaffold sequence, comprises SEQ ID NO:49, or a sequence having at least 90% sequence identity to all or a portion thereof.
- the gRNA is set forth in SEQ ID NO:49.
- a provided DNA-targeting system comprises any of the above gRNAs complexed with a Cas protein, such as a Cas9 protein.
- the Cas9 is a dCas9.
- the dCas9 is a dSaCas9, such as a dSaCas9 set forth in SEQ ID NO:72, or a variant and/or fusion thereof.
- the gRNA targets a target site in a FXN locus that comprises SEQ ID NO:4, a contiguous portion thereof of at least 14 nucleotides, a complementary sequence of any of the foregoing, or a sequence having at least 90% sequence identity to any of the foregoing.
- the gRNA comprises a spacer sequence comprising SEQ ID NO:25, a contiguous portion thereof of at least 14 nt, or a sequence having at least 90% sequence identity to any of the foregoing.
- the gRNA further comprises a scaffold sequence comprising SEQ ID NO:44, or a sequence having at least 90% sequence identity to SEQ ID NO:44.
- the gRNA including a spacer sequence and a scaffold sequence, comprises SEQ ID NO:50, or a sequence having at least 90% sequence identity to all or a portion thereof.
- the gRNA is set forth in SEQ ID NO:50.
- a provided DNA-targeting system comprises any of the above gRNAs complexed with a Cas protein, such as a Cas9 protein.
- the Cas9 is a dCas9.
- the dCas9 is a dSaCas9, such as a dSaCas9 set forth in SEQ ID NO:72, or a variant and/or fusion thereof.
- the gRNA targets a target site in a FXN locus that comprises SEQ ID NO:5, a contiguous portion thereof of at least 14 nucleotides, a complementary sequence of any of the foregoing, or a sequence having at least 90% sequence identity to any of the foregoing.
- the gRNA comprises a spacer sequence comprising SEQ ID NO:26, a contiguous portion thereof of at least 14 nt, or a sequence having at least 90% sequence identity to any of the foregoing.
- the gRNA further comprises a scaffold sequence comprising SEQ ID NO:44, or a sequence having at least 90% sequence identity to SEQ ID NO:44.
- the gRNA including a spacer sequence and a scaffold sequence, comprises SEQ ID NO:51, or a sequence having at least 90% sequence identity to all or a portion thereof.
- the gRNA is set forth in SEQ ID NO:51.
- a provided DNA-targeting system comprises any of the above gRNAs complexed with a Cas protein, such as a Cas9 protein.
- the Cas9 is a dCas9.
- the dCas9 is a dSaCas9, such as a dSaCas9 set forth in SEQ ID NO:72, or a variant and/or fusion thereof.
- the gRNA targets a target site in a FXN locus that comprises SEQ ID NO:6, a contiguous portion thereof of at least 14 nucleotides, a complementary sequence of any of the foregoing, or a sequence having at least 90% sequence identity to any of the foregoing.
- the gRNA comprises a spacer sequence comprising SEQ ID NO:27, a contiguous portion thereof of at least 14 nt, or a sequence having at least 90% sequence identity to any of the foregoing.
- the gRNA further comprises a scaffold sequence comprising SEQ ID NO:44, or a sequence having at least 90% sequence identity to SEQ ID NO:44.
- the gRNA including a spacer sequence and a scaffold sequence, comprises SEQ ID NO:52, or a sequence having at least 90% sequence identity to all or a portion thereof.
- the gRNA is set forth in SEQ ID NO:52.
- a provided DNA-targeting system comprises any of the above gRNAs complexed with a Cas protein, such as a Cas9 protein.
- the Cas9 is a dCas9.
- the dCas9 is a dSaCas9, such as a dSaCas9 set forth in SEQ ID NO:72, or a variant and/or fusion thereof.
- the gRNA targets a target site in a FXN locus that comprises SEQ ID NO:7, a contiguous portion thereof of at least 14 nucleotides, a complementary sequence of any of the foregoing, or a sequence having at least 90% sequence identity to any of the foregoing.
- the gRNA comprises a spacer sequence comprising SEQ ID NO:28, a contiguous portion thereof of at least 14 nt, or a sequence having at least 90% sequence identity to any of the foregoing.
- the gRNA further comprises a scaffold sequence comprising SEQ ID NO:44, or a sequence having at least 90% sequence identity to SEQ ID NO:44.
- the gRNA including a spacer sequence and a scaffold sequence, comprises SEQ ID NO:53, or a sequence having at least 90% sequence identity to all or a portion thereof.
- the gRNA is set forth in SEQ ID NO:53.
- a provided DNA-targeting system comprises any of the above gRNAs complexed with a Cas protein, such as a Cas9 protein.
- the Cas9 is a dCas9.
- the dCas9 is a dSaCas9, such as a dSaCas9 set forth in SEQ ID NO:72, or a variant and/or fusion thereof.
- the gRNA targets a target site in a FXN locus that comprises SEQ ID NO:8, a contiguous portion thereof of at least 14 nucleotides, a complementary sequence of any of the foregoing, or a sequence having at least 90% sequence identity to any of the foregoing.
- the gRNA comprises a spacer sequence comprising SEQ ID NO:29, a contiguous portion thereof of at least 14 nt, or a sequence having at least 90% sequence identity to any of the foregoing.
- the gRNA further comprises a scaffold sequence comprising SEQ ID NO:44, or a sequence having at least 90% sequence identity to SEQ ID NO:44.
- the gRNA including a spacer sequence and a scaffold sequence, comprises SEQ ID NO:54, or a sequence having at least 90% sequence identity to all or a portion thereof.
- the gRNA is set forth in SEQ ID NO:54.
- a provided DNA-targeting system comprises any of the above gRNAs complexed with a Cas protein, such as a Cas9 protein.
- the Cas9 is a dCas9.
- the dCas9 is a dSaCas9, such as a dSaCas9 set forth in SEQ ID NO:72, or a variant and/or fusion thereof.
- the gRNA targets a target site in a FXN locus that comprises SEQ ID NO:9, a contiguous portion thereof of at least 14 nucleotides, a complementary sequence of any of the foregoing, or a sequence having at least 90% sequence identity to any of the foregoing.
- the gRNA comprises a spacer sequence comprising SEQ ID NO:30, a contiguous portion thereof of at least 14 nt, or a sequence having at least 90% sequence identity to any of the foregoing.
- the gRNA further comprises a scaffold sequence comprising SEQ ID NO:44, or a sequence having at least 90% sequence identity to SEQ ID NO:44.
- the gRNA including a spacer sequence and a scaffold sequence, comprises SEQ ID NO:55, or a sequence having at least 90% sequence identity to all or a portion thereof.
- the gRNA is set forth in SEQ ID NO:55.
- a provided DNA-targeting system comprises any of the above gRNAs complexed with a Cas protein, such as a Cas9 protein.
- the Cas9 is a dCas9.
- the dCas9 is a dSaCas9, such as a dSaCas9 set forth in SEQ ID NO:72, or a variant and/or fusion thereof.
- the gRNA targets a target site in a FXN locus that comprises SEQ ID NO: 10, a contiguous portion thereof of at least 14 nucleotides, a complementary sequence of any of the foregoing, or a sequence having at least 90% sequence identity to any of the foregoing.
- the gRNA comprises a spacer sequence comprising SEQ ID NO:31, a contiguous portion thereof of at least 14 nt, or a sequence having at least 90% sequence identity to any of the foregoing.
- the gRNA further comprises a scaffold sequence comprising SEQ ID NO:44, or a sequence having at least 90% sequence identity to SEQ ID NO:44.
- the gRNA including a spacer sequence and a scaffold sequence, comprises SEQ ID NO:56, or a sequence having at least 90% sequence identity to all or a portion thereof.
- the gRNA is set forth in SEQ ID NO:56.
- a provided DNA-targeting system comprises any of the above gRNAs complexed with a Cas protein, such as a Cas9 protein.
- the Cas9 is a dCas9.
- the dCas9 is a dSaCas9, such as a dSaCas9 set forth in SEQ ID NO:72, or a variant and/or fusion thereof.
- the gRNA targets a target site in a FXN locus that comprises SEQ ID NO: 21, a contiguous portion thereof of at least 14 nucleotides, a complementary sequence of any of the foregoing, or a sequence having at least 90% sequence identity to any of the foregoing.
- the gRNA comprises a spacer sequence comprising SEQ ID NO:42, a contiguous portion thereof of at least 14 nt, or a sequence having at least 90% sequence identity to any of the foregoing.
- the gRNA further comprises a scaffold sequence comprising SEQ ID NO:44, or a sequence having at least 90% sequence identity to SEQ ID NO:44.
- the gRNA including a spacer sequence and a scaffold sequence, comprises SEQ ID NO:67, or a sequence having at least 90% sequence identity to all or a portion thereof.
- the gRNA is set forth in SEQ ID NO:67.
- a provided DNA-targeting system comprises any of the above gRNAs complexed with a Cas protein, such as a Cas9 protein.
- the Cas9 is a dCas9.
- the dCas9 is a dSaCas9, such as a dSaCas9 set forth in SEQ ID NO:72, or a variant and/or fusion thereof.
- the gRNA targets a target site in a FXN locus or a DNA regulatory element thereof that comprises the sequence selected from any one of SEQ ID NO: 11-20, a contiguous portion thereof of at least 14 nucleotides (e.g., 14, 15, 16, 17, 18, 19, 20, 21, or 22 nucleotides), a complementary sequence of any of the foregoing, or a sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to any of the foregoing.
- SEQ ID NO: 11-20 e.g., 14, 15, 16, 17, 18, 19, 20, 21, or 22 nucleotides
- the gRNA comprises a spacer sequence comprising the sequence selected from any one of SEQ ID NO:32-41, a contiguous portion thereof of at least 14 nt (e.g., 14, 15, 16, 17, 18, 19, 20, 21, or 22 nucleotides), or a sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to any of the foregoing.
- the gRNA further comprises a scaffold sequence.
- the scaffold sequence comprises the sequence set forth in SEQ ID NO:46, or a sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to SEQ ID NO:46.
- the gRNA, including a spacer sequence and a scaffold sequence comprises the sequence selected from any one of SEQ ID NO:57-66, or a sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to all or a portion thereof.
- the gRNA is set forth in the sequence selected from any one of SEQ ID NO:57-66.
- a provided DNA-targeting system comprises any of the aforementioned gRNAs complexed with a Cas protein, such as a Cas9 protein.
- the Cas9 is a dCas9.
- the dCas9 is a dSpCas9, such as a dSpCas9 set forth in SEQ ID NO:78, or a variant and/or fusion thereof.
- the gRNA targets a target site in a FXN locus or a DNA regulatory element thereof that comprises the sequence selected from any one of SEQ ID NO:256-265, a contiguous portion thereof of at least 14 nucleotides (e.g., 14, 15, 16, 17, 18, 19, 20, 21, or 22 nucleotides), a complementary sequence of any of the foregoing, or a sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to any of the foregoing.
- the gRNA further comprises a scaffold sequence.
- the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:258, and a scaffold sequence of SEQ ID NO:46. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:259, and a scaffold sequence of SEQ ID NO:46. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:260, and a scaffold sequence of SEQ ID NO:46. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:261, and a scaffold sequence of SEQ ID NO:46.
- the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:262, and a scaffold sequence of SEQ ID NO:46. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:263, and a scaffold sequence of SEQ ID NO:46. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:264, and a scaffold sequence of SEQ ID NO:46. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:265, and a scaffold sequence of SEQ ID NO:46.
- a provided DNA-targeting system comprises any of the aforementioned gRNAs complexed with a Cas protein, such as a Cas9 protein.
- the Cas9 is a dCas9.
- the dCas9 is a dSpCas9, such as a dSpCas9 set forth in SEQ ID NO: 78, or a variant and/or fusion thereof.
- the gRNA targets a target site in a FXN locus that comprises SEQ ID NO: 11, a contiguous portion thereof of at least 14 nucleotides, a complementary sequence of any of the foregoing, or a sequence having at least 90% sequence identity to any of the foregoing.
- the gRNA comprises a spacer sequence comprising SEQ ID NO:32, a contiguous portion thereof of at least 14 nt, or a sequence having at least 90% sequence identity to any of the foregoing.
- the gRNA further comprises a scaffold sequence comprising the sequence set forth in SEQ ID NO:46, or a sequence having at least 90% sequence identity to SEQ ID NO:46.
- the gRNA including a spacer sequence and a scaffold sequence, comprises SEQ ID NO:57, or a sequence having at least 90% sequence identity to all or a portion thereof.
- the gRNA is set forth in SEQ ID NO:57.
- a provided DNA-targeting system comprises any of the above gRNAs complexed with a Cas protein, such as a Cas9 protein.
- the Cas9 is a dCas9.
- the dCas9 is a dSpCas9, such as a dSpCas9 set forth in SEQ ID NO:78, or a variant and/or fusion thereof.
- the gRNA targets a target site in a FXN locus that comprises SEQ ID NO: 12, a contiguous portion thereof of at least 14 nucleotides, a complementary sequence of any of the foregoing, or a sequence having at least 90% sequence identity to any of the foregoing.
- the gRNA comprises a spacer sequence comprising SEQ ID NO:33, a contiguous portion thereof of at least 14 nt, or a sequence having at least 90% sequence identity to any of the foregoing.
- the gRNA further comprises a scaffold sequence comprising the sequence set forth in SEQ ID NO:46, or a sequence having at least 90% sequence identity to SEQ ID NO:46.
- the gRNA including a spacer sequence and a scaffold sequence, comprises SEQ ID NO:58, or a sequence having at least 90% sequence identity to all or a portion thereof.
- the gRNA is set forth in SEQ ID NO:58.
- a provided DNA-targeting system comprises any of the above gRNAs complexed with a Cas protein, such as a Cas9 protein.
- the Cas9 is a dCas9.
- the dCas9 is a dSpCas9, such as a dSpCas9 set forth in SEQ ID NO:78, or a variant and/or fusion thereof.
- the gRNA targets a target site in a FXN locus that comprises SEQ ID NO: 13, a contiguous portion thereof of at least 14 nucleotides, a complementary sequence of any of the foregoing, or a sequence having at least 90% sequence identity to any of the foregoing.
- the gRNA comprises a spacer sequence comprising SEQ ID NO:34, a contiguous portion thereof of at least 14 nt, or a sequence having at least 90% sequence identity to any of the foregoing.
- the gRNA further comprises a scaffold sequence comprising the sequence set forth in SEQ ID NO:46, or a sequence having at least 90% sequence identity to SEQ ID NO:46.
- the gRNA including a spacer sequence and a scaffold sequence, comprises SEQ ID NO:59, or a sequence having at least 90% sequence identity to all or a portion thereof.
- the gRNA is set forth in SEQ ID NO:59.
- a provided DNA-targeting system comprises any of the above gRNAs complexed with a Cas protein, such as a Cas9 protein.
- the Cas9 is a dCas9.
- the dCas9 is a dSpCas9, such as a dSpCas9 set forth in SEQ ID NO:78, or a variant and/or fusion thereof.
- the gRNA targets a target site in a FXN locus that comprises SEQ ID NO: 14, a contiguous portion thereof of at least 14 nucleotides, a complementary sequence of any of the foregoing, or a sequence having at least 90% sequence identity to any of the foregoing.
- the gRNA comprises a spacer sequence comprising SEQ ID NO:35, a contiguous portion thereof of at least 14 nt, or a sequence having at least 90% sequence identity to any of the foregoing.
- the gRNA further comprises a scaffold sequence comprising the sequence set forth in SEQ ID NO:46, or a sequence having at least 90% sequence identity to SEQ ID NO:46.
- the gRNA including a spacer sequence and a scaffold sequence, comprises SEQ ID NO:60, or a sequence having at least 90% sequence identity to all or a portion thereof.
- the gRNA is set forth in SEQ ID NO:60.
- a provided DNA-targeting system comprises any of the above gRNAs complexed with a Cas protein, such as a Cas9 protein.
- the Cas9 is a dCas9.
- the dCas9 is a dSpCas9, such as a dSpCas9 set forth in SEQ ID NO:78, or a variant and/or fusion thereof.
- the gRNA targets a target site in a FXN locus that comprises SEQ ID NO: 15, a contiguous portion thereof of at least 14 nucleotides, a complementary sequence of any of the foregoing, or a sequence having at least 90% sequence identity to any of the foregoing.
- the gRNA comprises a spacer sequence comprising SEQ ID NO:36, a contiguous portion thereof of at least 14 nt, or a sequence having at least 90% sequence identity to any of the foregoing.
- the gRNA further comprises a scaffold sequence comprising the sequence set forth in SEQ ID NO:46, or a sequence having at least 90% sequence identity to SEQ ID NO:46.
- the gRNA including a spacer sequence and a scaffold sequence, comprises SEQ ID NO:61, or a sequence having at least 90% sequence identity to all or a portion thereof.
- the gRNA is set forth in SEQ ID NO:61.
- a provided DNA-targeting system comprises any of the above gRNAs complexed with a Cas protein, such as a Cas9 protein.
- the Cas9 is a dCas9.
- the dCas9 is a dSpCas9, such as a dSpCas9 set forth in SEQ ID NO:78, or a variant and/or fusion thereof.
- the gRNA targets a target site in a FXN locus that comprises SEQ ID NO: 16, a contiguous portion thereof of at least 14 nucleotides, a complementary sequence of any of the foregoing, or a sequence having at least 90% sequence identity to any of the foregoing.
- the gRNA comprises a spacer sequence comprising SEQ ID NO:37, a contiguous portion thereof of at least 14 nt, or a sequence having at least 90% sequence identity to any of the foregoing.
- the gRNA further comprises a scaffold sequence comprising the sequence set forth in SEQ ID NO:46, or a sequence having at least 90% sequence identity to SEQ ID NO:46.
- the gRNA including a spacer sequence and a scaffold sequence, comprises SEQ ID NO:62, or a sequence having at least 90% sequence identity to all or a portion thereof.
- the gRNA is set forth in SEQ ID NO:62.
- a provided DNA-targeting system comprises any of the above gRNAs complexed with a Cas protein, such as a Cas9 protein.
- the Cas9 is a dCas9.
- the dCas9 is a dSpCas9, such as a dSpCas9 set forth in SEQ ID NO:78, or a variant and/or fusion thereof.
- the gRNA targets a target site in a FXN locus that comprises SEQ ID NO: 17, a contiguous portion thereof of at least 14 nucleotides, a complementary sequence of any of the foregoing, or a sequence having at least 90% sequence identity to any of the foregoing.
- the gRNA comprises a spacer sequence comprising SEQ ID NO:38, a contiguous portion thereof of at least 14 nt, or a sequence having at least 90% sequence identity to any of the foregoing.
- the gRNA further comprises a scaffold sequence comprising the sequence set forth in SEQ ID NO:46, or a sequence having at least 90% sequence identity to SEQ ID NO:46.
- the gRNA including a spacer sequence and a scaffold sequence, comprises SEQ ID NO:63, or a sequence having at least 90% sequence identity to all or a portion thereof.
- the gRNA is set forth in SEQ ID NO:63.
- a provided DNA-targeting system comprises any of the above gRNAs complexed with a Cas protein, such as a Cas9 protein.
- the Cas9 is a dCas9.
- the dCas9 is a dSpCas9, such as a dSpCas9 set forth in SEQ ID NO:78, or a variant and/or fusion thereof.
- the gRNA targets a target site in a FXN locus that comprises SEQ ID NO: 18, a contiguous portion thereof of at least 14 nucleotides, a complementary sequence of any of the foregoing, or a sequence having at least 90% sequence identity to any of the foregoing.
- the gRNA comprises a spacer sequence comprising SEQ ID NO:39, a contiguous portion thereof of at least 14 nt, or a sequence having at least 90% sequence identity to any of the foregoing.
- the gRNA further comprises a scaffold sequence comprising the sequence set forth in SEQ ID NO:46, or a sequence having at least 90% sequence identity to SEQ ID NO:46.
- the gRNA including a spacer sequence and a scaffold sequence, comprises SEQ ID NO:64, or a sequence having at least 90% sequence identity to all or a portion thereof.
- the gRNA is set forth in SEQ ID NO:64.
- a provided DNA-targeting system comprises any of the above gRNAs complexed with a Cas protein, such as a Cas9 protein.
- the Cas9 is a dCas9.
- the dCas9 is a dSpCas9, such as a dSpCas9 set forth in SEQ ID NO:78, or a variant and/or fusion thereof.
- the gRNA targets a target site in a FXN locus that comprises SEQ ID NO: 1
- the gRNA comprises a spacer sequence comprising SEQ ID NO:40, a contiguous portion thereof of at least 14 nt, or a sequence having at least 90% sequence identity to any of the foregoing.
- the gRNA further comprises a scaffold sequence comprising the sequence set forth in SEQ ID NO:46, or a sequence having at least 90% sequence identity to SEQ ID NO:46.
- the gRNA including a spacer sequence and a scaffold sequence, comprises SEQ ID NO:65, or a sequence having at least 90% sequence identity to all or a portion thereof.
- the gRNA is set forth in SEQ ID NO:65.
- a provided DNA-targeting system comprises any of the above gRNAs complexed with a Cas protein, such as a Cas9 protein.
- the Cas9 is a dCas9.
- the dCas9 is a dSpCas9, such as a dSpCas9 set forth in SEQ ID NO:78, or a variant and/or fusion thereof.
- the gRNA targets a target site in a FXN locus that comprises SEQ ID NO:20, a contiguous portion thereof of at least 14 nucleotides, a complementary sequence of any of the foregoing, or a sequence having at least 90% sequence identity to any of the foregoing.
- the gRNA comprises a spacer sequence comprising SEQ ID NO:41, a contiguous portion thereof of at least 14 nt, or a sequence having at least 90% sequence identity to any of the foregoing.
- the gRNA further comprises a scaffold sequence comprising the sequence set forth in SEQ ID NO:46, or a sequence having at least 90% sequence identity to SEQ ID NO:46.
- the gRNA including a spacer sequence and a scaffold sequence, comprises SEQ ID NO:66, or a sequence having at least 90% sequence identity to all or a portion thereof.
- the gRNA is set forth in SEQ ID NO:66.
- a provided DNA-targeting system comprises any of the above gRNAs complexed with a Cas protein, such as a Cas9 protein.
- the Cas9 is a dCas9.
- the dCas9 is a dSpCas9, such as a dSpCas9 set forth in SEQ ID NO:78, or a variant and/or fusion thereof.
- gRNAs guide RNAs
- the provided combination of gRNAs include two or more gRNAs, each of which target particular regions of a frataxin (FXN) locus.
- the two or more gRNAs each comprise any of the gRNAs described herein.
- a combination comprising a first gRNA comprising any of the gRNAs described herein, and one or more second gRNAs that binds to a second target site in a regulatory DNA element of a frataxin (FXN) locus.
- the second gRNA comprises any of the gRNAs described herein.
- a combination comprising: a first gRNA that binds a first target site in an enhancer region of a frataxin (FXN) locus, wherein the first target site is located within the genomic coordinates human genome assembly GRCh38 (hg38) chr9:69, 027, 282-69, 028, 497; and a second gRNA that binds a second target site in a promoter region of a FXN locus, wherein the second target site is located within the genomic coordinates hg38 chr9:68, 940, 179-69, 205 ,519.
- FXN frataxin
- the first gRNA comprises a gRNA spacer sequence set forth in SEQ ID NO:42 or a contiguous portion thereof of at least 14 nt.
- the second gRNA comprises a gRNA spacer sequence set forth in any one of SEQ ID NO:22-31 or a contiguous portion thereof of at least 14 nt.
- the second gRNA comprises a gRNA spacer sequence set forth in SEQ ID NO:22 or 28 or a contiguous portion thereof of at least 14 nt.
- the first gRNA comprises a gRNA spacer sequence set forth in SEQ ID NO:42 or a contiguous portion thereof of at least 14 nt
- the second gRNA comprises a gRNA spacer sequence set forth in any one of SEQ ID NO:22-31 or a contiguous portion thereof of at least 14 nt.
- the first gRNA comprises a gRNA spacer sequence set forth in SEQ ID NO:42 or a contiguous portion thereof of at least 14 nt
- the second gRNA comprises a gRNA spacer sequence set forth in SEQ ID NO:22 or a contiguous portion thereof of at least 14 nt.
- the first gRNA comprises a gRNA spacer sequence set forth in SEQ ID NO:42 or a contiguous portion thereof of at least 14 nt
- the second gRNA comprises a gRNA spacer sequence set forth in SEQ ID NO:28 or a contiguous portion thereof of at least 14 nt.
- a combination comprising: a first gRNA that binds a first target site in a promoter region of a FXN locus, wherein the second target site is located within the genomic coordinates hg38 chr9:68,940, 179-69,205,519; and a second gRNA that binds a second target site in a promoter region of a frataxin (FXN) locus, wherein the first target site is located within the genomic coordinates hg38 chr9:68,940, 179-69,205,519.
- FXN frataxin
- the first gRNA comprises a gRNA spacer sequence set forth in any one of SEQ ID NOs:22-31 or a contiguous portion thereof of at least 14 nt. In some embodiments, the first gRNA comprises a gRNA spacer sequence set forth in SEQ ID NO:22 or 28 or a contiguous portion thereof of at least 14 nt. In some embodiments, the second gRNA comprises a gRNA spacer sequence set forth in any one of SEQ ID NOs:22-31 or a contiguous portion thereof of at least 14 nt. In some embodiments, the second gRNA comprises a gRNA spacer sequence set forth in SEQ ID NO:22 or 28 or a contiguous portion thereof of at least 14 nt.
- the combination comprises: the first gRNA comprises a gRNA spacer sequence set forth in any one of SEQ ID NOs:22-31 or a contiguous portion thereof of at least 14 nt; and the second gRNA comprises a gRNA spacer sequence set forth in any one of SEQ ID NOs:22-31 or a contiguous portion thereof of at least 14 nt.
- the combination comprises: the first gRNA comprises a gRNA spacer sequence set forth in SEQ ID NO:22 or a contiguous portion thereof of at least 14 nt; and the second gRNA comprises a gRNA spacer sequence set forth in SEQ ID NO:28 or a contiguous portion thereof of at least 14 nt.
- the first gRNA comprises a gRNA spacer sequence set forth in any one of SEQ ID NOs:32-41 or a contiguous portion thereof of at least 14 nt.
- the second gRNA comprises a gRNA spacer sequence set forth in any one of SEQ ID NOs:32-41 or a contiguous portion thereof of at least 14 nt.
- the combination comprises: the first gRNA comprises a gRNA spacer sequence set forth in any one of SEQ ID NOs:32-41 or a contiguous portion thereof of at least 14 nt; and the second gRNA comprises a gRNA spacer sequence set forth in any one of SEQ ID NOs:32-41 or a contiguous portion thereof of at least 14 nt.
- fusion proteins that include (1) a DNA-targeting domain or a component thereof and (2) at least one effector domain.
- the DNA-targeting domain or component thereof e.g., a protein or polypeptide component of the DNA-targeting domain, such as the eZFP of the eZFP fusion protein, or the Cas component of the Cas-gRNA combination
- the at least one effector domain can be any described herein, for example, in Section II.B.
- the fusion protein is targeted to a target site, for example, one or more target sites at a FXN locus, such as any target site described herein, such as in Section I or Section II, by the DNA- targeting domain.
- a fusion protein comprising an eZFP can be referred to herein as an eZFP fusion protein.
- a fusion protein comprising a dCas protein can be referred to herein as a dCas fusion protein.
- fusion proteins that include (1) a DNA-targeting domain or a component thereof and (2) at least one effector domain, wherein: the DNA-targeting domain or a component thereof binds to a target site in a regulatory DNA element of a frataxin (FXN) locus; and the effector domain induces, catalyzes or leads to transcription activation, transcription co-activation, transcription elongation, transcription de-repression, transcription factor release, polymerization, histone modification, histone acetylation, histone deacetylation, nucleosome remodeling, chromatin remodeling, reversal of heterochromatin formation, nuclease, signal transduction, proteolysis, ubiquitination, deubiquitination, phosphorylation, dephosphorylation, splicing, nucleic acid association, DNA methylation, DNA demethylation, histone methylation, histone demethylation, or DNA base oxidation.
- the fusion protein comprises any of the effector domain
- binding of the DNA-targeting domain or a component thereof to the target site does not introduce a genetic disruption or a DNA break at or near the target site.
- the DNA-targeting domain comprises a Clustered Regularly Interspaced Short Palindromic Repeats associated (Cas)-guide RNA (gRNA) combination comprising (a) a Cas protein or a variant thereof and (b) at least one gRNA; a zinc finger protein (ZFP), such as an eZFP; a transcription activator-like effector (TAEE); a meganuclease; a homing endonuclease; or a I-Scel enzymes or a variant thereof, such as a catalytically inactive variant thereof.
- Cas Clustered Regularly Interspaced Short Palindromic Repeats associated
- gRNA Clustered Regularly Interspaced Short Palindromic Repeats associated
- ZFP zinc finger protein
- TEE transcription activator-like effector
- the DNA-targeting domain comprises an eZFP.
- the DNA-targeting domain comprises a Cas-gRNA combination comprising a Cas protein or a variant thereof and at least one gRNA, and the component of the DNA-targeting domain is a Cas protein or a variant thereof.
- the variant Cas protein lacks nuclease activity or is a deactivated Cas (dCas) protein.
- the gRNA is capable of complexing with the Cas protein or variant thereof.
- the Cas protein or a variant thereof is a Cas9 protein or a variant thereof.
- the variant Cas protein is a variant Cas9 protein that lacks nuclease activity or that is a deactivated Cas9 (dCas9) or a nuclease-inactive Cas9 (iCas9) protein.
- the dCas9 or iCas9 component of the fusion protein includes any described herein.
- the Cas9 protein or a variant thereof is a Staphylococcus aureus Cas9 (SaCas9) protein or a variant thereof.
- the variant Cas9 is a Staphylococcus aureus dCas9 protein (dSaCas9) that comprises at least one amino acid mutation selected from D10A and N580A, with reference to numbering of positions of SEQ ID NO:73.
- the variant Cas9 protein comprises the sequence set forth in SEQ ID NO:72, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the Cas9 protein or variant thereof is a Streptococcus pyogenes Cas9 (SpCas9) protein or a variant thereof.
- the variant Cas9 is a Streptococcus pyogenes dCas9 (dSpCas9) protein that comprises at least one amino acid mutation selected from D10A and H840A, with reference to numbering of positions of SEQ ID NO:79.
- the variant Cas9 protein comprises the sequence set forth in SEQ ID NO:78, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the DNA-targeting domain of the fusion protein is an engineered zinc finger protein (eZFP); a transcription activator-like effector (TALE); a meganuclease; a homing endonuclease; or a I-Scel enzymes or a variant thereof, such as a catalytically inactive variant thereof.
- the DNA-targeting domain of the fusion protein is an eZFP, such as any eZFP described herein, for example in Section I.
- the DNA-targeting domain of the fusion protein is targeted to one or more target sites at a FXN locus, such as one or more target sites described herein, for example, in Sections I and II.
- the DNA-targeting domain of the fusion protein is an engineered zinc finger protein (eZFP); a transcription activator-like effector (TALE); a meganuclease; a homing endonuclease; or a I-Scel enzymes or a variant thereof that is capable of binding to a target site at a FXN locus described herein, in a sequence-specific manner.
- eZFP engineered zinc finger protein
- TALE transcription activator-like effector
- meganuclease a homing endonuclease
- I-Scel enzymes or a variant thereof that is capable of binding to a target site at a FXN locus described herein, in a sequence-specific manner.
- the regulatory DNA element is an enhancer.
- the target site is located within the genomic coordinates human genome assembly GRCh38 (hg38) chr9:69, 027, 282-69, 028, 497.
- the target site is located within the genomic coordinates hg38 chr9:69, 027, 615-69, 028, 101.
- the target site comprises the sequence set forth in SEQ ID NO:21, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing.
- the regulatory DNA element is a promoter.
- the target site is located within the genomic coordinates hg38 chr9:68, 940, 179-69, 205 ,519.
- the target site comprises a sequence selected from any one of SEQ ID NOS: 1-10, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing.
- the target site comprises a sequence selected from any one of SEQ ID NOS: 11-20, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing.
- the target site is any target site provided herein, such as any target site provided in Section I or II.
- the target site is a target site for an eZFP or eZFP fusion protein, such as any target site provided in Section I or II.
- the effector domain induces, catalyzes or leads to transcription activation, transcription co-activation, transcription elongation. In some embodiments, the effector domain induces transcription activation. In some embodiments, the effector domain comprises at least one VP16 domain, or a VP16 tetramer (“VP64”) or a variant thereof. In some embodiments, the effector domain comprises the sequence set forth in SEQ ID NO:81 or 83, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
- the effector domain is selected from a p65 activation domain, a p300 domain, DPOLA, ENL, FOXO3, HSH2D, NCOA2, NCOA3, PSA1, PYGO1, RBM39, HERC2, or NOTCH2, or a domain thereof, a portion thereof or a variant thereof.
- the effector domain comprises a sequence selected from any one of SEQ ID NOS: 113-125, or a domain thereof, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
- the effector domain comprises a sequence selected from any one of SEQ ID NOS: 100-112, or a domain thereof, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
- the variant thereof comprises a truncation thereof.
- the effector domain comprises any one of the effector domains described herein.
- the at least one effector domain is fused to the N-terminus, the C- terminus, or both the N-terminus and the C-terminus, of the DNA-targeting domain or a component thereof (such as a protein or polypeptide component thereof, for example, a Cas component of a Cas- gRNA combination).
- the at least one effector domain is fused to the N-terminus, the C- terminus, or both the N-terminus and the C-terminus, of the DNA-targeting domain or a component thereof.
- the DNA-targeting system also includes one or more linkers connecting the DNA-targeting domain or a component thereof to the at least one effector domain.
- the DNA-targeting system further comprises one or more nuclear localization signals (NLS).
- the fusion protein comprises the sequence set forth in any one of SEQ ID NOs:85 and 159-173, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the NLS comprises the sequence set forth in any one of SEQ ID NOs:85 and 159-173, or a portion thereof.
- the NLS comprises the sequence set forth in SEQ ID NO: 85 or a portion thereof.
- An exemplary nucleotide sequence encoding the NLS set forth in SEQ ID NO: 85 is set forth in SEQ ID NO:84.
- the fusion protein comprises the sequence set forth in SEQ ID NO:77, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the fusion protein further comprises one or more linkers connecting the DNA-targeting domain or a component thereof to the at least one effector domain, and/or further comprises one or more nuclear localization signals (NLS).
- NLS nuclear localization signals
- the fusion protein comprises the sequence set forth in SEQ ID NO:71, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the fusion protein comprises the sequence set forth in SEQ ID NO:77, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the fusion protein includes at least one linker.
- a linker may be included anywhere in the polypeptide sequence of the fusion protein, for example, between the effector domain and the DNA-targeting domain or a component thereof.
- a linker may be of any length and designed to promote or restrict the mobility of components in the fusion protein.
- a linker may comprise any amino acid sequence of about 2 to about 100, about 5 to about 80, about 10 to about 60, or about 20 to about 50 amino acids.
- a linker may comprise an amino acid sequence of at least about 2, 3, 4, 5, 10, 15, 20, 25, 30, 35, 40, 45, 50, 55, 60, 65, 70, 75, 80 or 85 amino acids.
- a linker may comprise an amino acid sequence of less than about 100, 90, 80, 70, 60, 50, or 40 amino acids.
- a linker may include sequential or tandem repeats of an amino acid sequence that is 2 to 20 amino acids in length. Linkers may be rich in amino acids glycine (G), serine (S), and/or alanine (A).
- Linkers may include, for example, a GS linker such as (Gly-Gly-Gly-Gly-Ser)n.
- An exemplary GS linker is represented by the sequence GGGGS (SEQ ID NO: 158).
- a linker may comprise repeats of a sequence, for example as represented by the formula (GGGGS)n, wherein n is an integer that represents the number of times the GGGGS sequence is repeated (e.g. between 1 and 10 times). The number of times a linker sequence is repeated, for example n in a GS linker, can be adjusted to optimize the linker length and achieve appropriate separation of the functional domains.
- linkers may include, for example, Gly-Gly-Gly-Gly-Gly-Gly (SEQ ID NO: 154), Gly-Gly-Ala-Gly-Gly (SEQ ID NO: 155), Gly/Ser rich linkers such as Gly-Gly-Gly-Gly-Ser-Ser-Ser (SEQ ID NO: 156), or Gly/Ala rich linkers such as Gly-Gly-Gly-Gly-Ala-Ala-Ala (SEQ ID NO: 157), or Gly-Ser-Gly-Ser-Gly (SEQ ID NO:219).
- the linker is an XTEN linker.
- an XTEN linker is a recombinant polypeptide (e.g., an unstructured recombinant peptide) lacking hydrophobic amino acid residues.
- exemplary XTEN linkers are described in, for example, Schellenberger et al., Nature Biotechnology 27, 1186-1190 (2009) or WO 2021/247570.
- an exemplary linker comprises a linker described in WO 2021/247570.
- the linker is or comprises the sequence set forth in SEQ ID NO: 186 or SEQ ID NO: 174, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
- the linker comprises the sequence set forth in SEQ ID NO:186, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
- the linker comprises the sequence set forth in SEQ ID NO: 186, or a contiguous portion of SEQ ID NO: 186 of at least 5, 10, 15, 20, 25, 30, 35, 40, 45, 50, 55, 60, 65, 70 or 75 amino acids. In some aspects, the linker consists of the sequence set forth in SEQ ID NO: 186, or a contiguous portion of SEQ ID NO: 186 of at least 5, 10, 15, 20, 25, 30, 35, 40, 45, 50, 55, 60, 65, 70 or 75 amino acids. In some embodiments, the linker comprises the sequence set forth in SEQ ID NO: 186. In some embodiments, the linker consists of the sequence set forth in SEQ ID NO: 186.
- the linker comprises the sequence set forth in SEQ ID NO: 174, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
- the linker comprises the sequence set forth in SEQ ID NO: 174, or a contiguous portion of SEQ ID NO: 174 of at least 5, 10, or 15 amino acids.
- the linker consists of the sequence set forth in SEQ ID NO: 174, or a contiguous portion of SEQ ID NO: 174 of at least 5, 10, or 15 amino acids.
- the linker comprises the sequence set forth in SEQ ID NO: 174.
- the linker consists of the sequence set forth in SEQ ID NO: 174.
- Appropriate linkers may be selected or designed based rational criteria known in the art, for example as described in Chen et al. Adv. Drug Deliv. Rev. 65(10): 1357-1369 (2013).
- a linker comprises the sequence set forth in SEQ ID NO: 188, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
- a linker comprising the sequence set forth in SEQ ID NO: 188 is encoded by the nucleotide sequence set forth in SEQ ID NO: 187.
- a fusion protein described herein comprises one or more nuclear localization sequences (NLSs), such as about or more than about 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, or more NLSs.
- NLSs nuclear localization sequences
- each may be selected independently of the others, such that a single NLS may be present in more than one copy and/or in combination with one or more other NLSs present in one or more copies.
- NLSs include an NLS sequence derived from: the NLS of the SV40 virus large T-antigen, having the sequence PKKKRKV (SEQ ID NO: 159); the NLS from nucleoplasmin (e.g.
- the nucleoplasmin bipartite NLS having the sequence KRPAATKKAGQAKKKK (SEQ ID NO:85); the c-myc NLS having the sequence PAAKRVKLD (SEQ ID NO: 160) or RQRRNELKRSP (SEQ ID NO: 161); the hRNPAl M9 NLS having the sequence NQSSNFGPMKGGNFGGRSSGPYGGGGQYFAKPRNQGGY (SEQ ID NO: 162); the sequence RMRIZFKNKGKDTAELRRRRVEVSVELRKAKKDEQILKRRNV (SEQ ID NO: 163) of the IBB domain from importin-alpha; the sequences VSRKRPRP (SEQ ID NO: 164) and PPKKARED (SEQ ID NO: 165) of the myoma T protein; the sequence PQPKKKPL (SEQ ID NO: 166) of human p53; the sequence SALIKKKKKMAP (SEQ ID NO: 167) of mouse c-abl IV; the sequences D
- the one or more NLSs are of sufficient strength to drive accumulation of the fusion protein in a detectable amount in the nucleus of a eukaryotic cell.
- strength of nuclear localization activity may derive from the number of NLSs in the fusion protein, the particular NLS(s) used, or a combination of these factors.
- Detection of accumulation in the nucleus may be performed by any suitable technique.
- a detectable marker may be fused to the fusion protein, such that location within a cell may be visualized, such as in combination with a means for detecting the location of the nucleus (e.g. a stain specific for the nucleus such as DAPI).
- Cell nuclei may also be isolated from cells, the contents of which may then be analyzed by any suitable process for detecting protein, such as immunohistochemistry, Western blot, or enzyme activity assay. Accumulation in the nucleus may also be determined indirectly, such as by an assay for the effect of the fusion protein (e.g. an assay for altered gene expression activity in a cell transformed with the DNA-targeting system comprising the fusion protein), as compared to a control condition (e.g. an untransformed cell).
- an assay for the effect of the fusion protein e.g. an assay for altered gene expression activity in a cell transformed with the DNA-targeting system comprising the fusion protein
- a control condition e.g. an untransformed cell
- the NLS comprises the sequence set forth in any one of SEQ ID NO:85 and 160-173, or a portion thereof.
- DNA-targeting systems or fusion proteins that comprise a Cas protein or a variant thereof and at least one effector domain, wherein the effector domain increases transcription of the FXN locus.
- the DNA-targeting system or fusion protein comprises one or more tags, linkers and/or NLS sequences.
- exemplary tags, linkers and/or NLS sequences can be any described herein.
- sequences provided herein, including amino acid sequences for the DNA-targeting systems or fusion proteins provided herein contain sequences of one or more tags, linkers and/or NLS sequences.
- it is understood that the exemplary tags, linkers and/or NLS sequences are not required or are not the sole or exclusive tags, linkers and/or NLS sequences that can be employed in the DNA-targeting systems or fusion proteins.
- the DNA-targeting system or fusion protein comprises the sequence set forth in SEQ ID NO:71, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the DNA-targeting system or fusion protein comprises the sequence set forth in SEQ ID NO:71.
- the DNA-targeting system or fusion protein comprises the sequence set forth in SEQ ID NO:77, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the DNA-targeting system or fusion protein comprises the sequence set forth in SEQ ID NO: 77.
- an exemplary nucleotide sequence encoding the DNA-targeting system or fusion protein set forth in SEQ ID NO:77 is set forth in SEQ ID NO:75.
- the DNA-targeting system or fusion protein comprises, from N- to C-terminal order: a dSaCas9 set forth in SEQ ID NO:72, a linker and/or NLS, and a FOXO3 domain set forth in SEQ ID NO: 102.
- the DNA-targeting system or fusion protein comprises, from N- to C-terminal order: a dSaCas9 set forth in SEQ ID NO:72, a linker and/or NLS, and a HSH2D domain set forth in SEQ ID NO: 103.
- the DNA-targeting system or fusion protein comprises, from N- to C-terminal order: a dSaCas9 set forth in SEQ ID NO:72, a linker and/or NLS, and a NCOA2 domain set forth in SEQ ID NO: 104.
- the DNA- targeting system or fusion protein comprises, from N- to C-terminal order: a dSaCas9 set forth in SEQ ID NO:72, a linker and/or NLS, and a NCOA3 domain set forth in SEQ ID NO: 105.
- the DNA-targeting system or fusion protein comprises, from N- to C-terminal order: a dSaCas9 set forth in SEQ ID NO:72, a linker and/or NLS, and a PSA1 domain set forth in SEQ ID NO: 106. In some embodiments, the DNA-targeting system or fusion protein comprises, from N- to C-terminal order: a dSaCas9 set forth in SEQ ID NO:72, a linker and/or NLS, and a PYG01 domain set forth in SEQ ID NO: 107.
- the DNA-targeting system or fusion protein comprises, from N- to C- terminal order: a dSaCas9 set forth in SEQ ID NO:72, a linker and/or NLS, and a RBM39 domain set forth in SEQ ID NO: 108. In some embodiments, the DNA-targeting system or fusion protein comprises, from N- to C-terminal order: a dSaCas9 set forth in SEQ ID NO:72, a linker and/or NLS, and a HERC2 domain set forth in SEQ ID NO: 109.
- the DNA-targeting system or fusion protein comprises, from N- to C-terminal order: a dSaCas9 set forth in SEQ ID NO:72, a linker and/or NLS, and a DMD domain set forth in SEQ ID NO: 110.
- An exemplary nucleotide encoding the DMD domain set forth in SEQ ID NO: 110 is set forth in SEQ ID NO:97.
- the DNA-targeting system or fusion protein comprises, from N- to C-terminal order: a dSaCas9 set forth in SEQ ID NO:72, a linker and/or NLS, and a NOTCH2 domain set forth in SEQ ID NO: 111.
- the DNA- targeting system or fusion protein comprises, from N- to C-terminal order: a dSaCas9 set forth in SEQ ID NO:72, a linker and/or NLS, and a p300 core domain set forth in SEQ ID NO: 112.
- the DNA-targeting system or fusion protein comprises, from N- to C- terminal order: a DPOLA domain set forth in SEQ ID NO: 100, a linker and/or NLS, and a dSaCas9 set forth in SEQ ID NO:72.
- the DNA-targeting system or fusion protein comprises, from N- to C-terminal order: a dSaCas9 set forth in SEQ ID NO:72, a linker and/or NLS, and a ENL domain set forth in SEQ ID NO: 101, a linker and/or NLS, and a dSaCas9 set forth in SEQ ID NO:72.
- the DNA-targeting system or fusion protein comprises, from N- to C-terminal order: a dSaCas9 set forth in SEQ ID NO:72, a linker and/or NLS, and a FOXO3 domain set forth in SEQ ID NO: 102, a linker and/or NLS, and a dSaCas9 set forth in SEQ ID NO:72.
- the DNA-targeting system or fusion protein comprises, from N- to C-terminal order: a dSaCas9 set forth in SEQ ID NO:72, a linker and/or NLS, and a HSH2D domain set forth in SEQ ID NO: 103, a linker and/or NLS, and a dSaCas9 set forth in SEQ ID NO:72.
- the DNA-targeting system or fusion protein comprises, from N- to C-terminal order: a dSaCas9 set forth in SEQ ID NO:72, a linker and/or NLS, and a NCOA2 domain set forth in SEQ ID NO: 104, a linker and/or NLS, and a dSaCas9 set forth in SEQ ID NO:72.
- the DNA-targeting system or fusion protein comprises, from N- to C-terminal order: a dSaCas9 set forth in SEQ ID NO:72, a linker and/or NLS, and a NCOA3 domain set forth in SEQ ID NO: 105, a linker and/or NLS, and a dSaCas9 set forth in SEQ ID NO:72.
- the DNA-targeting system or fusion protein comprises, from N- to C-terminal order: a dSaCas9 set forth in SEQ ID NO:72, a linker and/or NLS, and a PSA1 domain set forth in SEQ ID NO: 106, a linker and/or NLS, and a dSaCas9 set forth in SEQ ID NO:72.
- the DNA-targeting system or fusion protein comprises, from N- to C-terminal order: a dSaCas9 set forth in SEQ ID NO:72, a linker and/or NLS, and a PYGO1 domain set forth in SEQ ID NO: 107, a linker and/or NLS, and a dSaCas9 set forth in SEQ ID NO:72.
- the DNA-targeting system or fusion protein comprises, from N- to C-terminal order: a dSaCas9 set forth in SEQ ID NO:72, a linker and/or NLS, and a RBM39 domain set forth in SEQ ID NO: 108, a linker and/or NLS, and a dSaCas9 set forth in SEQ ID NO:72.
- the DNA-targeting system or fusion protein comprises, from N- to C-terminal order: a dSaCas9 set forth in SEQ ID NO:72, a linker and/or NLS, and a HERC2 domain set forth in SEQ ID NO: 109, a linker and/or NLS, and a dSaCas9 set forth in SEQ ID NO:72.
- the DNA-targeting system or fusion protein comprises, from N- to C-terminal order: a dSaCas9 set forth in SEQ ID NO:72, a linker and/or NLS, and a DMD domain set forth in SEQ ID NO: 110, a linker and/or NLS, and a dSaCas9 set forth in SEQ ID NO:72.
- the DNA-targeting system or fusion protein comprises, from N- to C-terminal order: a dSaCas9 set forth in SEQ ID NO:72, a linker and/or NLS, and a NOTCH2 domain set forth in SEQ ID NO: 111 , a linker and/or NLS, and a dSaCas9 set forth in SEQ ID NO:72.
- the DNA-targeting system or fusion protein comprises, from N- to C-terminal order: a dSaCas9 set forth in SEQ ID NO:72, a linker and/or NLS, and a p300 core domain set forth in SEQ ID NO: 112, a linker and/or NLS, and a dSaCas9 set forth in SEQ ID NO:72.
- the DNA-targeting system or fusion protein comprises a dSaCas9 set forth in SEQ ID NO:72, and any of the epigenetic effector domains and/or multipartite effectors described herein, such as in Section II.B.
- the DNA-targeting system or fusion protein comprises the sequence set forth in any one of SEQ ID NOS:266-268 and 315-319, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
- the DNA-targeting system or fusion protein comprises the sequence set forth in any one of SEQ ID NOS:266-268 and 315-319.
- exemplary linkers or NLS sequences can be any described herein.
- the fusion protein is a split protein, i.e. comprises two or more separate polypeptide domains that interact or self-assemble to form a functional fusion protein.
- the split fusion protein comprises a dCas9 and an effector domain.
- the fusion protein comprises a split dCas9-effector domain fusion protein.
- the split fusion protein is assembled from separate polypeptide domains comprising trans-splicing inteins.
- Inteins are internal protein elements that self-excise from their host protein and catalyze ligation of flanking sequences with a peptide bond.
- the split fusion protein is assembled from a first polypeptide comprising an N-terminal intein and a second polypeptide comprising a C-terminal intein.
- the N terminal intein is the N terminal Npu Intein set forth in SEQ ID NO: 178.
- the C terminal intein is the C terminal Npu intein set forth in SEQ ID NO: 182.
- fusion proteins comprising a first polypeptide of a split variant Cas protein comprising an N-terminal fragment of a Cas protein and an N-terminal Intein, and at least one effector domain, wherein the effector domain induces transcription activation, transcription co-activation, transcription elongation, transcription de-repression, transcription factor release, polymerization, histone modification, histone acetylation, histone deacetylation, nucleosome remodeling, chromatin remodeling, reversal of heterochromatin formation, nuclease, signal transduction, proteolysis, ubiquitination, deubiquitination, phosphorylation, dephosphorylation, splicing, nucleic acid association, DNA methylation, DNA demethylation, histone methylation, histone demethylation, or DNA base oxidation.
- fusion proteins comprising a first polypeptide of a split variant Cas protein comprising an N-terminal fragment of a Cas protein (an exemplary amino acid sequence set forth in SEQ ID NO: 176, and an exemplary nucleotide sequence encoding the N-terminal fragment of the variant Cas protein set forth in SEQ ID NO: 175) and an N-terminal Intein (an exemplary amino acid sequence set forth in SEQ ID NO: 178 and an exemplary nucleotide sequence encoding the N-terminal Intein set forth in SEQ ID NO: 177), and at least one effector domain, wherein the effector domain increases transcription of the FXN locus.
- fusion proteins comprising a second polypeptide of a split variant Cas protein comprising a C-terminal fragment of a Cas protein and a C-terminal Intein and at least one effector domain, wherein the effector domain induces transcription activation, transcription co-activation, transcription elongation, transcription de-repression, transcription factor release, polymerization, histone modification, histone acetylation, histone deacetylation, nucleosome remodeling, chromatin remodeling, reversal of heterochromatin formation, nuclease, signal transduction, proteolysis, ubiquitination, deubiquitination, phosphorylation, dephosphorylation, splicing, nucleic acid association, DNA methylation, DNA demethylation, histone methylation, histone demethylation, or DNA base oxidation.
- fusion proteins comprising a second polypeptide of a split variant Cas protein comprising a C-terminal fragment of a Cas protein and a C-terminal Intein and at least one effector domain, wherein the effector domain increases transcription of the FXN locus.
- the second polypeptide of the split variant Cas protein, and a first polypeptide of the split variant Cas protein comprising an N-terminal fragment of the variant Cas protein and an N-terminal Intein are present in proximity or present in the same cell, the N-terminal Intein and C-terminal Intein self-excise and ligate the N-terminal fragment and the C-terminal fragment of the variant Cas9 to form a full-length variant Cas9 protein.
- the split fusion protein comprises a split dCas9-effector domain fusion protein assembled from two polypeptides.
- the first polypeptide comprises an effector domain catalytic domain and an N-terminal fragment of dSpCas9, followed by an N terminal Npu Intein (effector domain-dSpCas9-573N), and the second polypeptide comprises a C terminal Npu Intein, followed by a C-terminal fragment of dSpCas9 (dSpCas9-573C.
- the N- and C- terminal fragments of the fusion protein are split at position 573Glu of the dSpCas9 molecule, with reference to SEQ ID NO:79.
- the N-terminal Npu Intein (SEQ ID NO: 178) and C- terminal Npu Intein (set forth in SEQ ID NO: 182) may self-excise and ligate the two fragments, thereby forming the full-length dSpCas9-effector domain fusion protein when expressed in a cell.
- the polypeptides of a split protein may interact non-covalently to form a complex that recapitulates the activity of the non-split protein.
- two domains of a Cas enzyme expressed as separate polypeptides may be recruited by a gRNA to form a ternary complex that recapitulates the activity of the full-length Cas enzyme in complex with the gRNA, for example as described in Wright et al. PNAS 112(10):2984-2989 (2015).
- assembly of the split protein is inducible (e.g. light inducible, chemically inducible, small-molecule inducible).
- the two polypeptides of a split fusion protein may be delivered and/or expressed from separate vectors, such as any of the vectors described herein.
- the two polypeptides of a split fusion protein may be delivered to a cell and/or expressed from two separate AAV vectors, i.e. using a split AAV-based approach, for example as described in WO 2017/197238.
- combinations such as combinations of two or more DNA-targeting systems or components thereof.
- combinations of two or more DNA-targeting systems that independently target different target sites at a frataxin (FXN) locus are provided herein.
- the two or more DNA-targeting systems each comprise any of the DNA-targeting systems described herein.
- the DNA-targeting domain is a first DNA-targeting domain
- the DNA-targeting system further comprises one or more second DNA-targeting domains.
- the first DNA-targeting domain binds a first target site in an enhancer of a FXN locus
- the second DNA-targeting domain binds a second target site in a promoter of a FXN locus.
- the provided combination of DNA-targeting systems include two or more DNA-targeting systems, each of which target particular regions of a frataxin (FXN) locus.
- FXN frataxin
- a combination comprising a first DNA-targeting system comprising any of the DNA-targeting systems described herein, and one or more second DNA-targeting systems that binds to a second target site in a regulatory DNA element of a frataxin (FXN) locus.
- the second DNA-targeting system comprises any of the DNA-targeting systems described herein.
- combinations such as combinations of two or more DNA-targeting domains or fusion proteins or components thereof.
- combinations of two or more DNA-targeting domains or fusion proteins that independently target different target sites at a frataxin (FXN) locus are provided herein.
- the two or more DNA-targeting domains or fusion proteins each comprise any of the DNA-targeting domains or fusion proteins described herein.
- the DNA-targeting domain is a first DNA-targeting domain
- the DNA-targeting domain or fusion protein further comprises one or more second DNA-targeting domains.
- the first DNA-targeting domain binds a first target site in an enhancer of a FXN locus
- the second DNA-targeting domain binds a second target site in a promoter of a FXN locus.
- the provided combination of DNA-targeting domains or fusion proteins include two or more DNA-targeting domains or fusion proteins, each of which target particular regions of a frataxin (FXN) locus.
- FXN frataxin
- Also provided herein is a combination, comprising a first DNA-targeting domain or fusion protein comprising any of the DNA-targeting domains or fusion proteins described herein, and one or more second DNA-targeting domains or fusion proteins that binds to a second target site in a regulatory DNA element of a frataxin (FXN) locus.
- the second DNA-targeting domain or fusion protein comprises any of the DNA-targeting domains or fusion proteins described herein.
- DNA-targeting systems that binds to one or more target sites in a regulatory DNA element of a frataxin (FXN) locus
- the DNA-targeting system comprising: a first DNA- targeting domain that binds a first target site in an enhancer of a FXN locus, and a second DNA-targeting domain that binds a second target site in a promoter of a FXN locus.
- exemplary combination of DNA-targeting systems include: (a) a fusion protein comprising a Cas protein or a variant thereof and (b) a combination of gRNAs, such as a first gRNA that is capable of hybridizing to the target site or is complementary to the first target site and a second gRNA that is capable of hybridizing to the target site or is complementary to the second target site.
- gRNAs such as a first gRNA that is capable of hybridizing to the target site or is complementary to the first target site and a second gRNA that is capable of hybridizing to the target site or is complementary to the second target site.
- combinations of DNA-targeting systems comprising one type of Cas protein or variant thereof, such as a dCas9 protein or variant thereof, and two or more different gRNAs, such as a combination of gRNAs, such as any combination of gRNAs described herein.
- DNA-targeting systems comprising one type of Cas protein or variant thereof, such as a dCas9 protein or variant thereof, two or more different types of effector domains, and two or more different gRNAs, such as a combination of gRNAs, such as any combination of gRNAs described herein.
- combinations of DNA-targeting systems comprising one type of Cas protein or variant thereof, such as a dCas9 protein or variant thereof, two or more different types of effector domains, and two or more different gRNAs, such as a combination of gRNAs, such as any combination of gRNAs described herein.
- DNA-targeting systems comprising two or more different type of Cas protein or variant thereof, such as a dCas9 protein or variant thereof, and two or more different gRNAs, such as a combination of gRNAs, such as any combination of gRNAs described herein.
- combinations of DNA-targeting systems comprising two or more different types of DNA-targeting domains and one type of effector domain.
- combinations of DNA-targeting systems comprising two or more different types of DNA-targeting domains and two or more different types of effector domain.
- the first target site is located within the genomic coordinates human genome assembly GRCh38 (hg38) chr9:69, 027, 282-69, 028, 497
- the second target site is located within the genomic coordinates hg38 chr9:68, 940, 179-69, 205, 519.
- the first target site is located within the genomic coordinates hg38 chr9:69, 027, 615-69, 028, 101
- the second target site is located within the genomic coordinates hg38 chr9:68, 940, 179-69, 205, 519.
- the first DNA-targeting domain comprises a first Cas-gRNA combination comprising (a) a first Cas protein or a variant thereof and (b) a first gRNA that is capable of hybridizing to the target site or is complementary to the first target site; and the second DNA-targeting domain comprises a second Cas-gRNA combination comprising (a) a second Cas protein or a variant thereof and (b) a second gRNA that is capable of hybridizing to the target site or is complementary to the second target site.
- the first DNA-targeting domain comprises a first Cas-gRNA combination comprising (a) a first Cas protein or a variant thereof and (b) a first gRNA comprising at least one gRNA spacer sequence set forth in SEQ ID NO:42 or a contiguous portion thereof of at least 14 nt.
- the second DNA-targeting domain comprises a second Cas-gRNA combination comprising (a) a second Cas protein or a variant thereof and (b) a second gRNA comprising at least one gRNA spacer sequence set forth in SEQ ID NO:22 or 28 or a contiguous portion thereof of at least 14 nt.
- the first Cas-gRNA combination comprises (a) a first Cas protein or a variant thereof and (b) a first gRNA comprising at least one gRNA spacer sequence set forth in SEQ ID NO:42 or a contiguous portion thereof of at least 14 nt; and the second Cas-gRNA combination comprises (a) a second Cas protein or a variant thereof and (b) a second gRNA comprising at least one gRNA spacer sequence set forth in SEQ ID NO:22 or a contiguous portion thereof of at least 14 nt.
- the first Cas-gRNA combination comprises (a) a first Cas protein or a variant thereof and (b) a first gRNA comprising at least one gRNA spacer sequence set forth in SEQ ID NO:42 or a contiguous portion thereof of at least 14 nt; and the second Cas-gRNA combination comprises (a) a second Cas protein or a variant thereof and (b) a second gRNA comprising at least one gRNA spacer sequence set forth in SEQ ID NO:28 or a contiguous portion thereof of at least 14 nt.
- the first DNA-targeting domain binds a first target site in a promoter of a FXN locus; and the second DNA-targeting domain binds a second target site in a promoter of a FXN locus.
- DNA-targeting systems that binds to one or more target sites in a regulatory DNA element of a frataxin (FXN) locus
- the DNA-targeting system comprising: a first DNA- targeting domain that binds a first target site in a promoter of a FXN locus; and a second DNA-targeting domain that binds a second target site in a promoter of a FXN locus.
Landscapes
- Health & Medical Sciences (AREA)
- Life Sciences & Earth Sciences (AREA)
- Genetics & Genomics (AREA)
- Engineering & Computer Science (AREA)
- Chemical & Material Sciences (AREA)
- Organic Chemistry (AREA)
- Biomedical Technology (AREA)
- Molecular Biology (AREA)
- Bioinformatics & Cheminformatics (AREA)
- Zoology (AREA)
- Wood Science & Technology (AREA)
- General Health & Medical Sciences (AREA)
- Biotechnology (AREA)
- General Engineering & Computer Science (AREA)
- Biochemistry (AREA)
- Medicinal Chemistry (AREA)
- Biophysics (AREA)
- Microbiology (AREA)
- Plant Pathology (AREA)
- Physics & Mathematics (AREA)
- Pharmacology & Pharmacy (AREA)
- Toxicology (AREA)
- Nuclear Medicine, Radiotherapy & Molecular Imaging (AREA)
- Chemical Kinetics & Catalysis (AREA)
- Animal Behavior & Ethology (AREA)
- Public Health (AREA)
- Veterinary Medicine (AREA)
- General Chemical & Material Sciences (AREA)
- Gastroenterology & Hepatology (AREA)
- Proteomics, Peptides & Aminoacids (AREA)
- Medicines Containing Material From Animals Or Micro-Organisms (AREA)
- Medicines That Contain Protein Lipid Enzymes And Other Medicines (AREA)
- Pharmaceuticals Containing Other Organic And Inorganic Compounds (AREA)
- Peptides Or Proteins (AREA)
Abstract
Provided in some aspects are compositions, such as engineered zinc finger proteins (eZFPs) and fusion proteins comprising the eZFPs, DNA-targeting systems, fusion proteins, guide RNAs (gRNAs), and pluralities and combinations thereof, that bind to or target a frataxin (FXN) locus. In particular, the present disclosure relates to the modulation of expression of the FXN gene. In some aspects, the present disclosure also relates to polynucleotides, vectors, cells and pluralities and combinations thereof, that encode or comprise the eZFPs and fusion proteins comprising the eZFPs, DNA-targeting systems, fusion proteins, gRNAs or pluralities or combinations thereof, and methods and uses related to the provided compositions, for example, in modulating the expression of FXN, and/or in the treatment or therapy of diseases or disorders that involve the activity, function or expression of FXN, such as Friedreich's Ataxia (FA).
Description
FUSION PROTEINS AND SYSTEMS FOR TARGETED ACTIVATION OF FRATAXIN (FXN) AND RELATED METHODS
Cross-Reference to Related Applications
[00011 This application claims priority from U.S. provisional application No. 63/442,756 filed February 1, 2023 and U.S. provisional application No. 63/621,993 filed January 17, 2024, the contents of which are incorporated by reference in their entireties.
Incorporation by Reference of Sequence Listing
[0002] The present application is being filed along with a Sequence Listing in electronic format. The Sequence Listing is provided as a file entitled 224742002640SEQLIST.xml, created January 31, 2024, which is 1,019,999 bytes in size. The information in the electronic format of the Sequence Listing is herein incorporated by reference in its entirety.
Field
[0003] The present disclosure relates in some aspects to compositions, such as DNA-targeting systems, fusion proteins, guide RNAs (gRNAs), engineered zinc finger proteins (eZFPs) and fusion proteins comprising eZFPs, and pluralities and combinations thereof, that bind to or target a frataxin (FXN) locus. In particular, the present disclosure relates to the modulation of expression of the FXN gene. In some aspects, the present disclosure also relates to polynucleotides, vectors, cells and pluralities and combinations thereof, that encode or comprise the DNA-targeting systems, fusion proteins, gRNAs, engineered zinc finger proteins(eZFPs) and fusion proteins comprising eZFPs, or pluralities or combinations thereof, and methods and uses related to the provided compositions, for example, in modulating the expression of FXN, and/or in the treatment or therapy of diseases or disorders that involve the activity, function or expression of FXN, such as Friedreich’s Ataxia (FA).
Background
[0004] Genetic development disorders, including FA, are associated with reduced activity, mutation, and/or dysregulation of expression of the frataxin (FXN) gene. FA is an autosomal recessive neurodegenerative and cardiac disease, and is caused by a trinucleotide repeat expansion mutation in the FXN gene. FA can result in ataxia, areflexia, loss of vibratory sense and proprioception, dysarthria, cardiomyopathy, and/or associated arrhythmias, among other symptoms. Existing treatment of such genetic disorders are directed towards symptoms and providing support. Treatments that address the fundamental etiology and disease mechanism are needed. Provided are embodiments that meet such needs.
Summary
[0005] In some aspects, provided herein is an engineered zinc finger protein (eZFP) that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the target site is within a target region spanning the genomic coordinates chr9:69, 034, 900-69, 035, 900 from human genome assembly GRCh38 (hg38) (SEQ ID NO:430), or within a target region spanning the genomic coordinates chr9:69, 027, 282-69, 028, 497 from hg38 (SEQ ID NO:431). In some of any of the provided embodiments, the target site is within a target region spanning the genomic coordinates chr9:69, 034, 900-69, 035, 900 from hg38 (SEQ ID NO:430). In some of any of the provided embodiments, the target site is within a target region spanning the genomic coordinates chr9:69, 035, 300-69-035, 800 from hg38. In some of any of the provided embodiments, the target site is within a target region spanning the genomic coordinates chr9:69, 035, 350-69, 035, 450 from hg38. In some of any of the provided embodiments, the target site is within a target region spanning the genomic coordinates chr9:69, 035, 400-69, 035, 450 from hg38. In some of any of the provided embodiments, the target site is within a target region spanning the genomic coordinates chr9:69, 035, 530-69, 035, 580 from hg38. In some of any of the provided embodiments, the target site is within a target region spanning the genomic coordinates chr9:69, 035, 675-69, 035, 725 from hg38. In some of any of the provided embodiments, the target site is within a target region spanning the genomic coordinates chr9:69, 027, 282-69, 028, 497 from hg38 (SEQ ID NO:431). In some of any of the provided embodiments, the target site is within a target region spanning the genomic coordinates chr9:69, 027, 615-69, 028, 101 from hg38. In some of any of the provided embodiments, the target site is within a target region spanning the genomic coordinates chr9:69, 027, 775-69, 027, 875 from hg38. In some of any of the provided embodiments, the target site is within a target region spanning the genomic coordinates chr9:69, 027, 795-69, 027, 845 from hg38. In some of any of the provided embodiments, the target site comprises the nucleotide sequence set forth in any one of SEQ ID NOS:269-300 and 583-600, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
[0006] In some of any of the provided embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:272, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some of any of the provided embodiments, the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N- terminus to C-terminus, selected from F1-F6 as follows: Fl: QSGNLAR (SEQ ID NO:341); F2: QKVNRAG (SEQ ID NO:342); F3: DRSNLSR (SEQ ID NO:343); F4: QSGHLSR (SEQ ID NO:344); F5: TSGHLSR (SEQ ID NO:345); and F6: RSDALAR (SEQ ID NO:346). In some aspects, provided herein is an engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: QSGNLAR (SEQ ID NO:341); F2: QKVNRAG (SEQ ID NO:342); F3: DRSNLSR (SEQ ID NO:343); F4: QSGHLSR (SEQ ID NO:344); F5: TSGHLSR (SEQ ID NO:345); and F6: RSDALAR (SEQ ID NO:346). In some of any of the provided embodiments, the eZFP comprises the sequence set forth in SEQ ID NO:301, or a portion thereof, or an amino acid sequence that has at least 90%, 91%,
92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:301. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:308 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:308.
[0007] In some of any of the provided embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:277, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some of any of the provided embodiments, the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N- terminus to C-terminus, selected from F1-F6 as follows: Fl: RSDNLSE (SEQ ID NO:347); F2: KSWSRYK (SEQ ID NO:348); F3: TSGSLSR (SEQ ID NO:349); F4: RSDALAR (SEQ ID NO:350); F5: RSDNLSV (SEQ ID NO:351); and F6: FSSCRSA (SEQ ID NO:352). In some aspects, provided herein is an engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RSDNLSE (SEQ ID NO:347); F2: KSWSRYK (SEQ ID NO:348); F3: TSGSLSR (SEQ ID NO:349); F4: RSDALAR (SEQ ID NO:350); F5: RSDNLSV (SEQ ID NO:351); and F6: FSSCRSA (SEQ ID NO:352). In some of any of the provided embodiments, the eZFP comprises the sequence set forth in SEQ ID NO:302, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:302. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:309 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:309.
[0008] In some of any of the provided embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:280, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some of any of the provided embodiments, the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N- terminus to C-terminus, selected from F1-F6 as follows: Fl: TSGNLTR (SEQ ID NO:353); F2: EQTTRDK (SEQ ID NO:354); F3: RSANLAR (SEQ ID NO:355); F4: RLDNRTA (SEQ ID NO:356); F5: DSSHRTR (SEQ ID NO:357); and F6: RKYYLAK (SEQ ID NO:358). In some aspects, provided herein is an engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: TSGNLTR (SEQ ID NO:353); F2: EQTTRDK (SEQ ID NO:354); F3: RSANLAR (SEQ ID
NO:355); F4: RLDNRTA (SEQ ID NO:356); F5: DSSHRTR (SEQ ID NO:357); and F6: RKYYLAK (SEQ ID NO:358). In some of any of the provided embodiments, the eZFP comprises the sequence set forth in SEQ ID NO:303, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:303. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:310 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:310.
[0009] In some of any of the provided embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:281, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some of any of the provided embodiments, the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N- terminus to C-terminus, selected from F1-F6 as follows: Fl: RSAHLSR (SEQ ID NO:359); F2: DRSDLSR (SEQ ID NO:360); F3: RSDHLSV (SEQ ID NO:361); F4: RSDVRKT (SEQ ID NO:362); F5: QSGALAR (SEQ ID NO:363); and F6: RKYYLAK (SEQ ID NO:364). In some aspects, provided herein is an engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RSAHLSR (SEQ ID NO:359); F2: DRSDLSR (SEQ ID NO:360); F3: RSDHLSV (SEQ ID NO:361); F4: RSDVRKT (SEQ ID NO:362); F5: QSGALAR (SEQ ID NO:363); and F6: RKYYLAK (SEQ ID NO:364). In some of any of the provided embodiments, the eZFP comprises the sequence set forth in SEQ ID NO:304, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:304. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:311 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:311.
[0010] In some of any of the provided embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:283, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some of any of the provided embodiments, the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N- terminus to C-terminus, selected from F1-F6 as follows: Fl: RSAHLSR (SEQ ID NO:365); F2: RSDALAR (SEQ ID NO:366); F3: ATSNRSA (SEQ ID NO:367); F4: RSAHLSR (SEQ ID NO:368); F5: TSGSLSR (SEQ ID NO:369); and F6: QSGDLTR (SEQ ID NO:370). In some aspects, provided herein is an engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element
of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RSAHLSR (SEQ ID NO:365); F2: RSDALAR (SEQ ID NO:366); F3: ATSNRSA (SEQ ID NO:367); F4: RSAHLSR (SEQ ID NO:368); F5: TSGSLSR (SEQ ID NO:369); and F6: QSGDLTR (SEQ ID NO:370). In some of any of the provided embodiments, the eZFP comprises the sequence set forth in SEQ ID NO:305, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:305. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:312 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NOG 12.
[0011] In some of any of the provided embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:290, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some of any of the provided embodiments, the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N- terminus to C-terminus, selected from F1-F6 as follows: Fl: QSGDLTR (SEQ ID NO:371); F2: QSSDLRR (SEQ ID NO:372); F3: RSDNLSE (SEQ ID NO:373); F4: SSRNLAS (SEQ ID NO:374); F5: DRSHLTR (SEQ ID NO:375); and F6: RSDDLTR (SEQ ID NO:376). In some aspects, provided herein is an engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: QSGDLTR (SEQ ID NO:371); F2: QSSDLRR (SEQ ID NO:372); F3: RSDNLSE (SEQ ID NO:373); F4: SSRNLAS (SEQ ID NO:374); F5: DRSHLTR (SEQ ID NO:375); and F6: RSDDLTR (SEQ ID NO:376). In some of any of the provided embodiments, the eZFP comprises the sequence set forth in SEQ ID NO:306, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:306. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:313 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NOG 13.
[0012] In some of any of the provided embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:299, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some of any of the provided embodiments, the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N- terminus to C-terminus, selected from F1-F6 as follows: Fl: LRHHLTR (SEQ ID NO:377); F2:
QSAHLKA (SEQ ID NO:378); F3: LPQTLQR (SEQ ID NO:379); F4: QNATRTK (SEQ ID NO:380); F5: QSSHLTR (SEQ ID NO:381); and F6: RSDHLSR (SEQ ID NO:382). In some aspects, provided herein is an engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: LRHHLTR (SEQ ID NO:377); F2: QSAHLKA (SEQ ID NO:378); F3: LPQTLQR (SEQ ID NO:379); F4: QNATRTK (SEQ ID NO:380); F5: QSSHLTR (SEQ ID NO:381); and F6: RSDHLSR (SEQ ID NO:382). In some of any of the provided embodiments, the eZFP comprises the sequence set forth in SEQ ID NO:307, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:307. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:314 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:314.
[0013] In some of any of the provided embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:583, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some of any of the provided embodiments, the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N- terminus to C-terminus, selected from F1-F6 as follows: Fl: RSDSLLR (SEQ ID NO:475); F2: TSSNRKT (SEQ ID NO:476); F3: RSAHLSR (SEQ ID NO:477); F4: TSGSLTR (SEQ ID NO:478); F5: QSGDLTR (SEQ ID NO:479); and F6: QWGTRYR (SEQ ID NO:480). In some aspects, provided herein is an engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RSDSLLR (SEQ ID NO:475); F2: TSSNRKT (SEQ ID NO:476); F3: RSAHLSR (SEQ ID NO:477); F4: TSGSLTR (SEQ ID NO:478); F5: QSGDLTR (SEQ ID NO:479); and F6: QWGTRYR (SEQ ID NO:480). In some of any of the provided embodiments, the eZFP comprises the sequence set forth in SEQ ID NO:439, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:439. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:457 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:457.
[0014] In some of any of the provided embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:584, a contiguous portion thereof of at least 12 nt, or a complementary
sequence of any of the foregoing. In some of any of the provided embodiments, the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N- terminus to C-terminus, selected from F1-F6 as follows: Fl: QARHLTC (SEQ ID NO:481); F2: QSGHLSR (SEQ ID NO:482); F3: RSDVLSE (SEQ ID NO:483); F4: KHSTRRV (SEQ ID NO:484); F5: QSSDLSR (SEQ ID NO:485); and F6: WKWNLRA (SEQ ID NO:486). In some aspects, provided herein is an engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: QARHLTC (SEQ ID NO:481); F2: QSGHLSR (SEQ ID NO:482); F3: RSDVLSE (SEQ ID NO:483); F4: KHSTRRV (SEQ ID NO:484); F5: QSSDLSR (SEQ ID NO:485); and F6: WKWNLRA (SEQ ID NO:486). In some of any of the provided embodiments, the eZFP comprises the sequence set forth in SEQ ID NO:440, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:440. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:458 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 458.
[0015] In some of any of the provided embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:585, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some of any of the provided embodiments, the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N- terminus to C-terminus, selected from F1-F6 as follows: Fl: RSDNLAR (SEQ ID NO:487); F2: WRGDRVK (SEQ ID NO:488); F3: YKHVLSD (SEQ ID NO:489); F4: TSGSLTR (SEQ ID NO:490); F5: QSGNLAR (SEQ ID NO:491); and F6: RARDLSK (SEQ ID NO:492). In some aspects, provided herein is an engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RSDNLAR (SEQ ID NO:487); F2: WRGDRVK (SEQ ID NO:488); F3: YKHVLSD (SEQ ID NO:489); F4: TSGSLTR (SEQ ID NO:490); F5: QSGNLAR (SEQ ID NO:491); and F6: RARDLSK (SEQ ID NO:492). In some of any of the provided embodiments, the eZFP comprises the sequence set forth in SEQ ID NO:441, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:441. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:459 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided
embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:459.
[0016] In some of any of the provided embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:586, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some of any of the provided embodiments, the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N- terminus to C-terminus, selected from F1-F6 as follows: Fl: QSANRTK (SEQ ID NO:493); F2: QSGNEAR (SEQ ID NO:494); F3: RSDNLSV (SEQ ID NO:495); F4: IRSTLRD (SEQ ID NO:496); F5: QNAHRKT (SEQ ID NO:497); and F6: HRSSLRR (SEQ ID NO:498). In some aspects, provided herein is an engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: QSANRTK (SEQ ID NO:493); F2: QSGNLAR (SEQ ID NO:494); F3: RSDNLSV (SEQ ID NO:495); F4: IRSTLRD (SEQ ID NO:496); F5: QNAHRKT (SEQ ID NO:497); and F6: HRSSLRR (SEQ ID NO:498). In some of any of the provided embodiments, the eZFP comprises the sequence set forth in SEQ ID NO:442, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:442. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:460 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:460.
[0017] In some of any of the provided embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:587, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some of any of the provided embodiments, the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N- terminus to C-terminus, selected from F1-F6 as follows: Fl: QAGNRST (SEQ ID NO:499); F2: DRSALAR (SEQ ID NG:500); F3: RSDNLAR (SEQ ID NO:501); F4: WRGDRVK (SEQ ID NO:502); F5: YKHVLSD (SEQ ID NO:503); and F6: TSGSLTR (SEQ ID NO:504). In some aspects, provided herein is an engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: QAGNRST (SEQ ID NO:499); F2: DRSALAR (SEQ ID NG:500); F3: RSDNLAR (SEQ ID NO:501); F4: WRGDRVK (SEQ ID NO:502); F5: YKHVLSD (SEQ ID NO:503); and F6: TSGSLTR (SEQ ID NO:504). In some of any of the provided embodiments, the eZFP comprises the sequence set forth in SEQ ID NO:443, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:443. In
some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:461 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 461.
[0018] In some of any of the provided embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:588, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some of any of the provided embodiments, the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N- terminus to C-terminus, selected from F1-F6 as follows: Fl: RSDNLSV (SEQ ID NO:505); F2: IRSTLRD (SEQ ID NO:506); F3: QNAHRKT (SEQ ID NO:507); F4: HRSSLRR (SEQ ID NO:508); F5: RSDNLAR (SEQ ID NO:509); and F6: QRSPLPA (SEQ ID NO:510). In some aspects, provided herein is an engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RSDNLSV (SEQ ID NO:505); F2: IRSTLRD (SEQ ID NO:506); F3: QNAHRKT (SEQ ID NO:507); F4: HRSSLRR (SEQ ID NO:508); F5: RSDNLAR (SEQ ID NO:509); and F6: QRSPLPA (SEQ ID NO:510). In some of any of the provided embodiments, the eZFP comprises the sequence set forth in SEQ ID NO:444, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:444. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:462 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:462.
[0019] In some of any of the provided embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:589, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some of any of the provided embodiments, the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N- terminus to C-terminus, selected from F1-F6 as follows: Fl: DRSTRTK (SEQ ID NO:511); F2: RSDYLAK (SEQ ID NO:512); F3: LRHHLTR (SEQ ID NO:513); F4: QSAHLKA (SEQ ID NO:514); F5: LPQTLQR (SEQ ID NO:515); and F6: QNATRTK (SEQ ID NO:516). In some aspects, provided herein is an engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: DRSTRTK (SEQ ID NO:511); F2: RSDYLAK (SEQ ID NO:512); F3: LRHHLTR (SEQ ID NO:513); F4: QSAHLKA (SEQ ID NO:514); F5: LPQTLQR (SEQ ID NO:515); and F6: QNATRTK (SEQ ID NO:516). In some of any of the provided embodiments, the eZFP comprises the sequence set
forth in SEQ ID NO:445, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:445. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:463 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 463.
[0020] In some of any of the provided embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:590, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some of any of the provided embodiments, the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N- terminus to C-terminus, selected from F1-F6 as follows: Fl: RSADLTR (SEQ ID NO:517); F2: RSDDLTR (SEQ ID NO:518); F3: QSSDLSR (SEQ ID NO:519); F4: WHSSLHQ (SEQ ID NO:520); F5: RSDSLSQ (SEQ ID NO:521); and F6: RKADRTR (SEQ ID NO:522). In some aspects, provided herein is an engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RSADLTR (SEQ ID NO:517); F2: RSDDLTR (SEQ ID NO:518); F3: QSSDLSR (SEQ ID NO:519); F4: WHSSLHQ (SEQ ID NO:520); F5: RSDSLSQ (SEQ ID NO:521); and F6: RKADRTR (SEQ ID NO:522). In some of any of the provided embodiments, the eZFP comprises the sequence set forth in SEQ ID NO:446, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:446. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:464 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:464.
[0021 [ In some of any of the provided embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:591, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some of any of the provided embodiments, the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N- terminus to C-terminus, selected from F1-F6 as follows: Fl: RNDALTE (SEQ ID NO:523); F2: RKDNLKN (SEQ ID NO:524); F3: TSGELVR (SEQ ID NO:525); F4: HRTTLTN (SEQ ID NO:526); F5: TTGNLTV (SEQ ID NO:527); and F6: RTDTLRD (SEQ ID NO:528). In some aspects, provided herein is an engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as
follows: Fl: RNDALTE (SEQ ID NO:523); F2: RKDNLKN (SEQ ID NO:524); F3: TSGELVR (SEQ ID NO:525); F4: HRTTLTN (SEQ ID NO:526); F5: TTGNLTV (SEQ ID NO:527); and F6: RTDTLRD (SEQ ID NO:528). In some of any of the provided embodiments, the eZFP comprises the sequence set forth in SEQ ID NO:447, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:447. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:465 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 465.
[0022| In some of any of the provided embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:592, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some of any of the provided embodiments, the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N- terminus to C-terminus, selected from F1-F6 as follows: Fl: RKDNLKN (SEQ ID NO:529); F2: RADNLTE (SEQ ID NO:530); F3: TSHSLTE (SEQ ID NO:531); F4: SKKHLAE (SEQ ID NO:532); F5: TSGELVR (SEQ ID NO:533); and F6: TSGELVR (SEQ ID NO:534). In some aspects, provided herein is an engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RKDNLKN (SEQ ID NO:529); F2: RADNLTE (SEQ ID NO:530); F3: TSHSLTE (SEQ ID NO:531); F4: SKKHLAE (SEQ ID NO:532); F5: TSGELVR (SEQ ID NO:533); and F6: TSGELVR (SEQ ID NO:534). In some of any of the provided embodiments, the eZFP comprises the sequence set forth in SEQ ID NO:448, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:448. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:466 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:466.
[0023] In some of any of the provided embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:593, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some of any of the provided embodiments, the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N- terminus to C-terminus, selected from F1-F6 as follows: Fl: THLDLIR (SEQ ID NO:535); F2: DCRDLAR (SEQ ID NO:536); F3: RSDELVR (SEQ ID NO:537); F4: RNDALTE (SEQ ID NO:538); F5: SKKHLAE (SEQ ID NO:539); and F6: QSGHLTE (SEQ ID NO:540). In some aspects, provided
herein is an engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: THLDLIR (SEQ ID NO:535); F2: DCRDLAR (SEQ ID NO:536); F3: RSDELVR (SEQ ID NO:537); F4: RNDALTE (SEQ ID NO:538); F5: SKKHLAE (SEQ ID NO:539); and F6: QSGHLTE (SEQ ID NO:540). In some of any of the provided embodiments, the eZFP comprises the sequence set forth in SEQ ID NO:449, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:449. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:467 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 467.
[0024] In some of any of the provided embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:594, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some of any of the provided embodiments, the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N- terminus to C-terminus, selected from F1-F6 as follows: Fl: HTGHLLE (SEQ ID NO:541); F2: DPGHLVR (SEQ ID NO:542); F3: THLDLIR (SEQ ID NO:543); F4: DCRDLAR (SEQ ID NO:544); F5: RSDELVR (SEQ ID NO:545); and F6: RNDALTE (SEQ ID NO:546). In some aspects, provided herein is an engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: HTGHLLE (SEQ ID NO:541); F2: DPGHLVR (SEQ ID NO:542); F3: THLDLIR (SEQ ID NO:543); F4: DCRDLAR (SEQ ID NO:544); F5: RSDELVR (SEQ ID NO:545); and F6: RNDALTE (SEQ ID NO:546). In some of any of the provided embodiments, the eZFP comprises the sequence set forth in SEQ ID NO:450, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:450. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:468 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:468.
[0025] In some of any of the provided embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:595, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some of any of the provided embodiments, the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-
terminus to C-terminus, selected from F1-F6 as follows Fl: RSDKLVR (SEQ ID NO:547); F2: RSDHLTT (SEQ ID NO:548); F3: RNDALTE (SEQ ID NO:549); F4: TTGALTE (SEQ ID NO:550); F5: THLDLIR (SEQ ID NO:551); and F6: DPGHLVR (SEQ ID NO:552). In some aspects, provided herein is an engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RSDKLVR (SEQ ID NO:547); F2: RSDHLTT (SEQ ID NO:548); F3: RNDALTE (SEQ ID NO:549); F4: TTGALTE (SEQ ID NO:550); F5: THLDLIR (SEQ ID NO:551); and F6: DPGHLVR (SEQ ID NO:552). In some of any of the provided embodiments, the eZFP comprises the sequence set forth in SEQ ID NO:451, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:451. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:469 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 469.
[0026] In some of any of the provided embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:596, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some of any of the provided embodiments, the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N- terminus to C-terminus, selected from F1-F6 as follows Fl: TKNSLTE (SEQ ID NO:553); F2: QLAHLRA (SEQ ID NO:554); F3: TSGSLVR (SEQ ID NO:555); F4: RSDNLVR (SEQ ID NO:556); F5: QNSTLTE (SEQ ID NO:557); and F6: RADNLTE (SEQ ID NO:558). In some aspects, provided herein is an engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: TKNSLTE (SEQ ID NO:553); F2: QLAHLRA (SEQ ID NO:554); F3: TSGSLVR (SEQ ID NO:555); F4: RSDNLVR (SEQ ID NO:556); F5: QNSTLTE (SEQ ID NO:557); and F6: RADNLTE (SEQ ID NO:558). In some of any of the provided embodiments, the eZFP comprises the sequence set forth in SEQ ID NO:452, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:452. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:470 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:470.
[0027] In some of any of the provided embodiments, the target site comprises the nucleotide
sequence set forth in SEQ ID NO:597, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some of any of the provided embodiments, the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N- terminus to C-terminus, selected from F1-F6 as follows Fl: RADNETE (SEQ ID NO:559); F2: TKNSETE (SEQ ID NO:560); F3: QLAHLRA (SEQ ID NO:561); F4: TSGSLVR (SEQ ID NO:562); F5: RSDNLVR (SEQ ID NO:563); and F6: QNSTLTE (SEQ ID NO:564). In some aspects, provided herein is an engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RADNLTE (SEQ ID NO:559); F2: TKNSLTE (SEQ ID NO:560); F3: QLAHLRA (SEQ ID NO:561); F4: TSGSLVR (SEQ ID NO:562); F5: RSDNLVR (SEQ ID NO:563); and F6: QNSTLTE (SEQ ID NO:564). In some of any of the provided embodiments, the eZFP comprises the sequence set forth in SEQ ID NO:453, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:453. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:471 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 471.
[0028] In some of any of the provided embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:598, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some of any of the provided embodiments, the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N- terminus to C-terminus, selected from F1-F6 as follows Fl: TSGHLVR (SEQ ID NO:565); F2: QLAHLRA (SEQ ID NO:566); F3: TSGELVR (SEQ ID NO:567); F4: QSGDLRR (SEQ ID NO:568); F5: QRAHLER (SEQ ID NO:569); and F6: RSDKLVR (SEQ ID NO:570). In some aspects, provided herein is an engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: TSGHLVR (SEQ ID NO:565); F2: QLAHLRA (SEQ ID NO:566); F3: TSGELVR (SEQ ID NO:567); F4: QSGDLRR (SEQ ID NO:568); F5: QRAHLER (SEQ ID NO:569); and F6: RSDKLVR (SEQ ID NO:570). In some of any of the provided embodiments, the eZFP comprises the sequence set forth in SEQ ID NO:454, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:454. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:472 or a portion thereof, or an amino acid sequence that has at least 90%, 91%,
92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:472.
[0029] In some of any of the provided embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:599, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some of any of the provided embodiments, the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N- terminus to C-terminus, selected from F1-F6 as follows Fl: REDNEHT (SEQ ID NO:571); F2: TSGHEVR (SEQ ID NO:572); F3: QLAHLRA (SEQ ID NO:573); F4: TSGELVR (SEQ ID NO:574); F5: QSGDLRR (SEQ ID NO:575); and F6: QRAHLER (SEQ ID NO:576). In some aspects, provided herein is an engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: REDNLHT (SEQ ID NO:571); F2: TSGHLVR (SEQ ID NO:572); F3: QLAHLRA (SEQ ID NO:573); F4: TSGELVR (SEQ ID NO:574); F5: QSGDLRR (SEQ ID NO:575); and F6: QRAHLER (SEQ ID NO:576). In some of any of the provided embodiments, the eZFP comprises the sequence set forth in SEQ ID NO:455, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:455. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:473 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 473.
[0030] In some of any of the provided embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:600, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some of any of the provided embodiments, the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N- terminus to C-terminus, selected from F1-F6 as follows Fl: QRSDLTR (SEQ ID NO:577); F2: QGGTLRR (SEQ ID NO:578); F3: TSAHLAR (SEQ ID NO:579); F4: RREHLVR (SEQ ID NO:580); F5: QRHGLSS (SEQ ID NO:581); and F6: QRNALRG (SEQ ID NO:582). In some aspects, provided herein is an engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: QRSDLTR (SEQ ID NO:577); F2: QGGTLRR (SEQ ID NO:578); F3: TSAHLAR (SEQ ID NO:579); F4: RREHLVR (SEQ ID NO:580); F5: QRHGLSS (SEQ ID NO:581); and F6: QRNALRG (SEQ ID NO:582). In some of any of the provided embodiments, the eZFP comprises the sequence set forth in SEQ ID NO:456, or a portion thereof, or an amino acid sequence that has at least 90%, 91%,
92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided
embodiments, the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:456. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:474 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:474.
[0031] In some aspects, provided herein is a fusion protein comprising any of the engineered zinc finger proteins (eZFPs) provided herein. In some aspects, provided herein is a fusion protein comprising:
(a) any eZFP provided herein that binds to a target site in a regulatory DNA element of a FXN locus; and
(b) at least one epigenetic effector domain that increases transcription of the FXN locus. In some of any of the provided embodiments, the at least one epigenetic effector domain comprises: a VP64 domain, a p65 activation domain, a p300 domain, an Rta domain, a CBP domain, a VPR domain, a VPH domain, an HSF1 domain, a TET protein domain, optionally wherein the TET protein is TET1, a SunTag domain, a domain from DPOLA, ENL, FOXO3, HSH2D, NCOA2, NCOA3, PSA1, PYGO1, RBM39, HERC2, or NOTCH2, or a domain, portion, variant, or truncation of any of the foregoing. In some of any of the provided embodiments, the at least one epigenetic effector domain comprises the sequence set forth in any of SEQ ID NOS:81, 83, 100-109, 111-122, 124, 125, 134-140, 152, and 383-396, or a domain, portion, variant, or truncation thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
[0032] In some of any of the provided embodiments, the at least one effector domain comprises at least one VP16 domain, or a VP16 tetramer (“VP64”) or a variant thereof. In some of any of the provided embodiments, the at least one effector domain comprises VP64. In some of any of the provided embodiments, the at least one effector domain comprises a VP64 domain comprising the sequence set forth in SEQ ID NO:83, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing. In some of any of the provided embodiments, the at least one effector domain comprises a VP64 domain comprising the sequence set forth in SEQ ID NO: 83.
[0033] In some of any of the provided embodiments, the at least one epigenetic effector domain comprises: a domain from DPOLA, ENL, FOXO3, HSH2D, NCOA2, NCOA3, PSA1, PYGO1, RBM39, HERC2, or NOTCH2, or a domain, portion, variant, or truncation of any of the foregoing. In some of any of the provided embodiments, the at least one epigenetic effector domain comprises the sequence set forth in any of SEQ ID NOS:383-393, or a domain, portion, variant, or truncation thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing. In some of any of the provided embodiments, the at least one effector domain comprises a domain from NCOA2, NCOA3, FOXO3, PYGO1, or a portion or variant of any of the foregoing. In some of any of the provided embodiments, each effector domain of the at least one effector domain is independently selected from an NCOA2 domain, an NCOA3 domain, a FOXO3 domain, and a PYGO1 domain. In some of any of the provided embodiments, the at least one effector
domain comprises a domain from NCOA2 comprising the sequence set forth in SEQ ID NO: 104 or SEQ ID NO:387, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the at least one effector domain comprises a domain from NCOA2 set forth in or SEQ ID NO:387. In some of any of the provided embodiments, the at least one effector domain comprises a domain from NCOA3 comprising the sequence set forth in SEQ ID NO: 105 or SEQ ID NO:388, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the at least one effector domain comprises a domain from NCOA3 set forth in or SEQ ID NO:388. In some of any of the provided embodiments, the at least one effector domain comprises a domain from FOXO3 comprising the sequence set forth in SEQ ID NO: 102 or SEQ ID NO:385, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the at least one effector domain comprises a domain from FOXO3 set forth in or SEQ ID NO:385. In some of any of the provided embodiments, the at least one effector domain comprises a domain from PYGO1 comprising the sequence set forth in SEQ ID NO: 107 or SEQ ID NO:390, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the at least one effector domain comprises a domain from PYGO1 set forth in or SEQ ID NO:390. In some of any of the provided embodiments, the at least one effector domain is a multipartite effector composed of at least two effector domains.
[0034] In some of any of the provided embodiments, the multipartite effector is composed of two effector domains. In some of any of the provided embodiments, the multipartite effector is composed of three effector domains. In some of any of the provided embodiments, the multipartite effector is set forth in any one of SEQ ID NOS:397-418, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing. In some of any of the provided embodiments, the multipartite effector is set forth in any one of SEQ ID NOS:411- 418, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing. In some of any of the provided embodiments, the multipartite effector comprises, in the N-terminal to C-terminal direction, domains from FOXO3, FOXO3, and NCOA3. In some of any of the provided embodiments, the multipartite effector comprises the sequence set forth in SEQ ID NO:415, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the multipartite effector comprises, in the N-terminal to C-terminal direction, domains from NCOA3; FOXO3, and FOXO3. In some of any of the provided embodiments, the multipartite effector comprises the sequence set forth in SEQ ID NO:418, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the multipartite effector comprises, in the N-terminal to C-
terminal direction, domains from NC0A3, FOXO3, and NC0A3. In some of any of the provided embodiments, the multipartite effector comprises the sequence set forth in SEQ ID NO:413, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the multipartite effector comprises, in the N-terminal to C-terminal direction, domains from NCOA2, FOXO3, and NCOA3. In some of any of the provided embodiments, the multipartite effector comprises the sequence set forth in SEQ ID NO:416, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the multipartite effector comprises, in the N-terminal to C-terminal direction, domains from PYGO1, FOXO3, and NCOA3. In some of any of the provided embodiments, the multipartite effector comprises the sequence set forth in SEQ ID NO:411, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
[0035] In some of any of the provided embodiments, the at least one epigenetic effector domain is fused to the N-terminus of the eZFP. In some of any of the provided embodiments, the at least one epigenetic effector domain is fused to the C-terminus of the eZFP. In some of any of the provided embodiments, the at least one epigenetic effector domain is fused to both the N-terminus and the C- terminus, of the eZFP. In some of any of the provided embodiments, the fusion protein further comprises one or more nuclear localization signals (NLS). In some of any of the provided embodiments, the fusion protein further comprises one or more linkers. In some of any of the provided embodiments, the one or more linkers are in between any two of the components of the fusion protein, including the eZFP, any of the at least one effector domains, and the one or more NFS. In some of any of the provided embodiments, the one or more linkers connect the eZFP and the at least one epigenetic effector domain. In some of any of the provided embodiments, the fusion protein further comprises one or more NFS, the eZFP, and the at least one epigenetic effector domain, in order from N-terminus to C-terminus. In some of any of the provided embodiments, the one or more NFS comprises a SV40 NFS sequence set forth in SEQ ID NO: 159 or a c-myc NFS sequence set forth in SEQ ID NO: 160. In some of any of the provided embodiments, the fusion protein comprises the sequence set forth in any of SEQ ID NOS:320-340 and 419-425, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing. In some of any of the provided embodiments, the fusion protein comprises the sequence set forth in any of SEQ ID NOS:320-340, 419- 425, and 608-671. The fusion protein of any of claims 189-232, wherein the fusion protein comprises the sequence set forth in any of SEQ ID NOS:320-340, 419-425, and 608-671. In some of any of the provided embodiments, the fusion protein comprises the sequence set forth in any of SEQ ID NOS:320- 340, 419-425, and 636-653. In some of any of the provided embodiments, the fusion protein comprises the sequence set forth in any of SEQ ID NOS:320-340 and 419-425. In some of any of the provided embodiments, the fusion protein comprises the sequence set forth in any of SEQ ID NOS:636-653. In some of any of the provided embodiments, the fusion protein comprises the sequence set forth in any of
SEQ ID NOS:608-635 and 654-671. In some of any of the provided embodiments, the fusion protein comprises the sequence set forth in any of SEQ ID NOS:608-635. In some of any of the provided embodiments, the fusion protein comprises the sequence set forth in any of SEQ ID NOS:654-671. In some of any of the provided embodiments, the fusion protein comprises the sequence set forth in SEQ ID NO:326, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the fusion protein comprises the sequence set forth in SEQ ID NO: 333, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the fusion protein comprises the sequence set forth in SEQ ID NO:340, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the fusion protein comprises the sequence set forth in SEQ ID NO:425, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the fusion protein comprises the sequence set forth in SEQ ID NO:662, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the fusion protein comprises the sequence set forth in SEQ ID NO:660, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the fusion protein comprises the sequence set forth in SEQ ID NO:658, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
[0036] In some aspects, provided herein is a polynucleotide encoding any eZFP, or any fusion protein provided herein, or a portion or component of any of the foregoing. In some aspects, provided herein is a plurality of polynucleotides encoding any eZFP, or any fusion protein provided herein, or a portion or component of any of the foregoing.
[0037] In some aspects, provided herein is a vector comprising any eZFP, any fusion protein, any polynucleotide, or any plurality of polynucleotides provided herein, or a portion or component of any of the foregoing. In some of any of the provided embodiments, the vector is a viral vector. In some of any of the provided embodiments, the vector is an adeno-associated virus (AAV) vector. In some of any of the provided embodiments, the AAV vector is selected from the group consisting of: AAV1, AAV2, AAV3, AAV4, AAV5, AAV6, AAV7, AAV8, AAV9, AAV10, AAV11, AAV12, AAV-DJ, and AAVrh74. In some of any of the provided embodiments, the AAV vector is AAV6. In some of any of the provided embodiments, the AAV vector is AAV9. In some of any of the provided embodiments, the AAV vector is AAV-DJ. In some of any of the provided embodiments, the AAV vector is AAVrh74. In some of any of the provided embodiments, the vector is a lenti viral vector. In some of any of the provided embodiments, the vector is a non-viral vector. In some of any of the provided embodiments, the non-viral vector is selected from the group consisting of: a lipid nanoparticle, a liposome, an exosome, and a cell
penetrating peptide. In some of any of the provided embodiments, the vector exhibits tropism for a nervous system cell, optionally a neuron, a heart cell, optionally a cardiomyocyte, a skeletal muscle cell, a fibroblast, an induced pluripotent stem cell, and/or a cell derived from any of the foregoing, or for a combination of any of the foregoing cells. In some of any of the provided embodiments, the vector exhibits tropism for induced pluripotent stem cells. In some of any of the provided embodiments, the vector exhibits tropism for neurons and cardiomyocytes. In some of any of the provided embodiments, the vector comprises one vector, or two or more vectors.
[0038] In some aspects, provided herein is an AAV vector comprising one or both of a) a first nucleic acid comprising an elongation factor alpha short (EFS) promoter operably linked to a sequence encoding a fusion protein comprising (i) a deactivated Cas (dCas) protein and (ii) at least one effector domain that increases transcription of a frataxin (FXN) locus; and b) a second nucleic acid comprising a U6 promoter operably linked to a sequence encoding a guide RNA (gRNA) comprising a gRNA spacer sequence that is capable of hybridizing to a target site in a regulatory DNA element of a FXN locus and/or is complementary to the target site. In some of any of the provided embodiments, the AAV vector comprises both the first nucleic acid and the second nucleic acid. In some of any of the provided embodiments, the first and second nucleic acid are comprised in a single polynucleotide. In some of any of the provided embodiments, the EFS promoter comprises the sequence set forth in SEQ ID NO:436, or a sequence having at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to thereto. In some of any of the provided embodiments, the EFS promoter comprises the sequence set forth in SEQ ID NO:436. In some of any of the provided embodiments, the U6 promoter is a mini-U6 promoter. In some of any of the provided embodiments, the mini-U6 promoter comprises the sequence set forth in SEQ ID NO:433, or a sequence having at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to thereto. In some of any of the provided embodiments, the mini-U6 promoter comprises the sequence set forth in SEQ ID NO:433.
[0039] In some aspects, provided herein is an AAV vector comprising a nucleic acid comprising a promoter selected from an elongation factor alpha short (EFS), CAG, or human elongation factor- 1 alpha (EFla) promoter operably linked to a sequence encoding a fusion protein comprising (i) an eZFP that is capable of hybridizing to a target site in a regulatory DNA element of a frataxin (FXN) locus and/or is complementary to the target site and (ii) at least one effector domain that increases transcription of the frataxin (FXN) locus. In some of any of the provided embodiments, the EFS promoter comprises the sequence set forth in SEQ ID NO:436, or a sequence having at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to thereto. In some of any of the provided embodiments, the EFS promoter comprises the sequence set forth in SEQ ID NO:436. In some of any of the provided embodiments, the CAG promoter comprises the sequence set forth in SEQ ID NO:602, or a sequence having at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to thereto. In some of any of the provided embodiments, the CAG promoter comprises the sequence set forth in SEQ ID NO:602. In some of any of the provided embodiments, the EFla promoter comprises the
sequence set forth in SEQ ID NO:603, or a sequence having at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to thereto. In some of any of the provided embodiments, the EFla promoter comprises the sequence set forth in SEQ ID NO:603. In some of any of the provided embodiments, the nucleic acid further comprises a 5’ untranslated region (UTR) set forth in SEQ ID NO:605.
[0040] In some of any of the provided embodiments, the AAV vector further comprises inverted terminal repeats (ITRs). In some of any of the provided embodiments, the ITRs are a first and second ITR, comprising the sequences set forth in SEQ ID NO:434 and SEQ ID NO:435, respectively. In some of any of the provided embodiments, the single polynucleotide comprises, in the 5’ to 3’ direction, the EFS promoter, the sequence encoding the fusion protein, the U6 promoter, and the sequence encoding the gRNA. In some of any of the provided embodiments, the single polynucleotide further comprises a first IRT 5’ of the EFS promoter and a second ITR 3’ of the sequence encoding the gRNA. In some of any of the provided embodiments, the first nucleic acid or the nucleic acid further comprises a polyA sequence selected from a SpA site or a bGH site downstream of the sequence encoding the fusion protein. In some of any of the provided embodiments, the first nucleic acid or the nucleic acid comprising a sequence encoding a fusion protein further comprises a polyA sequence selected from a SpA site downstream of the sequence encoding the fusion protein. In some of any of the provided embodiments, the first nucleic acid comprising a sequence encoding a fusion protein further comprises a polyA sequence selected from a bGH site downstream of the sequence encoding the fusion protein. In some of any of the provided embodiments, the nucleic acid comprising a sequence encoding a fusion protein further comprises a polyA sequence selected from a SpA site or bGH site downstream of the sequence encoding the fusion protein. In some of any of the provided embodiments, the SpA site comprises the sequence set forth in SEQ ID NO:437. In some of any of the provided embodiments, the bGH site comprises the sequence set forth in SEQ ID NO: 604. In some of any of the provided embodiments, the first nucleic acid further comprises a woodchuck hepatitis virus post-transcriptional regulatory element (WPRE) in proximal to the SpA site, optionally wherein the WPRE is located between the sequence encoding the fusion protein and the SpA site. In some of any of the provided embodiments, the gRNA is capable of complexing with the dCas protein. In some of any of the provided embodiments, the gRNA comprises a gRNA spacer sequence that is capable of hybridizing to the target site or is complementary to the target site. In some of any of the provided embodiments, the dCas protein is a Staphylococcus aureus dCas9 (dSaCas9) protein or a Streptococcus pyogenes dCas9 (dSpCas9) protein. In some of any of the provided embodiments, the dCas protein is a Staphylococcus aureus dCas9 protein (dSaCas9) that comprises at least one amino acid mutation selected from D10A and N580A, with reference to numbering of positions of SEQ ID NO:73, and/or the dCas protein comprises the sequence set forth in SEQ ID NO:72, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the dCas is a Streptococcus pyogenes dCas9 (dSpCas9) protein that comprises at least one amino acid mutation selected from D10A and H840A, with reference
to numbering of positions of SEQ ID NO:79, and/or the dCas protein comprises the sequence set forth in SEQ ID NO:78, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
[0041] In some of any of the provided embodiments, the regulatory DNA element is an enhancer. In some of any of the provided embodiments, the target site is located within a target region spanning the genomic coordinates chr9:69, 027, 282-69, 028, 497 from hg38 (SEQ ID NO:431), optionally wherein the target site is located within a target region spanning the genomic coordinates chr9:69, 027, 615-69, 028, 101 from hg38, optionally wherein the target site is located within a target region spanning the genomic coordinates chr9:69, 027, 825-69, 027, 875. In some of any of the provided embodiments, the target site comprises the sequence set forth in SEQ ID NO:21, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing. In some of any of the provided embodiments, the gRNA comprises a gRNA spacer sequence comprising the sequence set forth in SEQ ID NO:42, or a contiguous portion thereof of at least 14 nt. In some of any of the provided embodiments, the gRNA further comprises the sequence set forth in SEQ ID NO:44, optinally wherein the gRNA comprises the sequence set forth in SEQ ID NO:67, optionally wherein the gRNA is the gRNA sequence set forth in SEQ ID NO:67.
[0042] In some of any of the provided embodiments, the target site comprises the sequence set forth in any one of SEQ ID NOS: 272 and 277, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing. In some of any of the provided embodiments, the eZFP comprises the sequence set forth in any one of SEQ ID NOS: 301 and 302.
[0043] In some of any of the provided embodiments, the regulatory DNA element is a promoter. In some of any of the provided embodiments, the target site is within a target region spanning the genomic coordinates chr9:69, 034, 900-69, 035, 900 from hg38 (SEQ ID NO:430), optionally wherein the target site is within a target region spanning the genomic coordinates chr9:69, 035, 300-69-035, 800 from hg38; chr9:69, 035, 350-69, 035, 450 from hg38; or chr9:69, 035, 675-69, 035, 725. In some of any of the provided embodiments, the target site comprises a sequence selected from any of SEQ ID NOS: 1-10, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing. In some of any of the provided embodiments, the gRNA comprises a gRNA spacer sequence comprising a sequence selected from any of SEQ ID NOS:22-31, or a contiguous portion thereof of at least 14 nt. In some of any of the provided embodiments, the gRNA comprises a gRNA spacer sequence comprising SEQ ID NO:22, or a contiguous portion thereof of at least 14 nt. In some of any of the provided embodiments, the gRNA comprises a gRNA spacer sequence comprising SEQ ID NO:28, or a contiguous portion thereof of at least 14 nt. In some of any of the provided embodiments, the gRNA further comprises the sequence set forth in SEQ ID NO:44, optionally wherein the gRNA comprises a sequence selected from any of SEQ ID NOS:47-56, optionally wherein the gRNA is the gRNA sequence set forth in any of SEQ ID NOS:47- 56, optionally wherein the gRNA is set forth in SEQ ID NO:47 or 53. In some of any of the provided embodiments, the target site comprises a sequence selected from any of SEQ ID NOS: 11-20, a
contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing. In some of any of the provided embodiments, the gRNA comprises a gRNA spacer sequence comprising a sequence selected from any of SEQ ID NOS:32-41, or a contiguous portion thereof of at least 14 nt. In some of any of the provided embodiments, the gRNA further comprises the sequence set forth in SEQ ID NO:46, and/or wherein the gRNA comprises a sequence selected from any of SEQ ID NOS:57-66, optionally wherein the gRNA is the gRNA set forth in any of SEQ ID NOS:57-66. In some of any of the provided embodiments, the gRNA spacer sequence is between 14 nt and 24 nt, or between 16 nt and 22 nt in length, optionally wherein the gRNA spacer sequence is 18 nt, 19 nt, 20 nt, 21 nt or 22 nt in length.
[0044] In some of any of the provided embodiments, the target site comprises the sequence set forth in any one of SEQ ID NOS: 280-283, 290, 299, and 583-600, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing. In some of any of the provided embodiments, the target site comprises the sequence set forth in any one of SEQ ID NOS: 299, 587, 589, and 591, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing. In some of any of the provided embodiments, the target site comprises the sequence set forth in SEQ ID NO: 299, a contiguous portion thereof of at least 14 nt, or a complementary sequence of the sequence set forth in SEQ ID NO: 299. In some of any of the provided embodiments, the target site comprises the sequence set forth in SEQ ID NO: 587, a contiguous portion thereof of at least 14 nt, or a complementary sequence of the sequence set forth in SEQ ID NO: 587. In some of any of the provided embodiments, the target site comprises the sequence set forth in SEQ ID NO: 589, a contiguous portion thereof of at least 14 nt, or a complementary sequence of the sequence set forth in SEQ ID NO: 589. In some of any of the provided embodiments, the target site comprises the sequence set forth in SEQ ID NO: 591, a contiguous portion thereof of at least 14 nt, or a complementary sequence of the sequence set forth in SEQ ID NO: 591. In some of any of the provided embodiments, the eZFP comprises the sequence set forth in any one of SEQ ID NOS: 303-307 and 439-456. In some of any of the provided embodiments, the eZFP comprises the sequence set forth in any one of SEQ ID NOS: 307, 441, 443, and 445. In some of any of the provided embodiments, the eZFP comprises the sequence set forth in SEQ ID NO: 307. In some of any of the provided embodiments, the eZFP comprises the sequence set forth in SEQ ID NO: 441. In some of any of the provided embodiments, the eZFP comprises the sequence set forth in SEQ ID NO: 443. In some of any of the provided embodiments, the eZFP comprises the sequence set forth in SEQ ID NO: 445.
[0045] In some of any of the provided embodiments, the at least one effector domain induces transcription activation. In some of any of the provided embodiments, the at least one epigenetic effector domain comprises: a VP64 domain, a p65 activation domain, a p300 domain, an Rta domain, a CBP domain, a VPR domain, a VPH domain, an HSF1 domain, a TET protein domain, optionally wherein the TET protein is TET1, a SunTag domain, a domain from DPOLA, ENL, FOXO3, HSH2D, NCOA2, NCOA3, PSA1, PYGO1, RBM39, HERC2, or NOTCH2, or a domain, portion, variant, or truncation of any of the foregoing. In some of any of the provided embodiments, the at least one epigenetic effector
domain comprises the sequence set forth in any of SEQ ID NOS:81, 83, 100-109, 111-122, 124, 125, 134-140, 152, and 383-396, or a domain, portion, variant, or truncation thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing. In some of any of the provided embodiments, the at least one effector domain is fused to the N-terminus, the C-terminus, or both the N-terminus and the C-terminus, of the dCas protein or eZFP. In some of any of the provided embodiments, the AAV vector further comprises one or more linkers connecting the dCas protein or eZFP to the at least one effector domain, and/or further comprising one or more nuclear localization signals (NFS). In some of any of the provided embodiments, the at least one effector domain comprises at least one VP16 domain, or a VP16 tetramer (“VP64”) or a variant thereof. In some of any of the provided embodiments, the at least one effector domain comprises VP64. In some of any of the provided embodiments, the at least one effector domain comprises the sequence set forth in SEQ ID NO:81 or 83, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing. In some of any of the provided embodiments, the at least one effector domain comprises the sequence set forth in SEQ ID NO: 81 or 83. In some of any of the provided embodiments, the at least one epigenetic effector domain comprises: a domain from DPOLA, ENL, FOXO3, HSH2D, NCOA2, NCOA3, PSA1, PYGO1, RBM39, HERC2, or NOTCH2, or a domain, portion, variant, or truncation of any of the foregoing. In some of any of the provided embodiments, the at least one epigenetic effector domain comprises the sequence set forth in any of SEQ ID NOS:383-393, or a domain, portion, variant, or truncation thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing. In some of any of the provided embodiments, the at least one effector domain comprises a domain from NCOA2, NCOA3, FOXO3, PYGO1, or a portion or variant of any of the foregoing. In some of any of the provided embodiments, each effector domain of the at least one effector domain is independently selected from an NCOA2 domain, an NCOA3 domain, a FOXO3 domain, and a PYGO1 domain. In some of any of the provided embodiments, the at least one effector domain comprises a domain from NCOA2 comprising the sequence set forth in SEQ ID NO: 104 or SEQ ID NO:387, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the at least one effector domain comprises a domain from NCOA2 set forth in or SEQ ID NO:387. In some of any of the provided embodiments, the at least one effector domain comprises a domain from NCOA3 comprising the sequence set forth in SEQ ID NO: 105 or SEQ ID NO:388, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the at least one effector domain comprises a domain from NCOA3 set forth in or SEQ ID NO:388. In some of any of the provided embodiments, the at least one effector domain comprises a domain from FOXO3 comprising the sequence set forth in SEQ ID NO: 102 or SEQ ID NO:385, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity
thereto. In some of any of the provided embodiments, the at least one effector domain comprises a domain from F0X03 set forth in or SEQ ID NO:385. In some of any of the provided embodiments, the at least one effector domain comprises a domain from PYG01 comprising the sequence set forth in SEQ ID NO: 107 or SEQ ID NO:390, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the at least one effector domain comprises a domain from PYGO1 set forth in or SEQ ID NO:390.
[0046] In some of any of the provided embodiments, the at least one effector domain is a multipartite effector composed of at least two effector domains. In some of any of the provided embodiments, the multipartite effector is composed of two effector domains. In some of any of the provided embodiments, the multipartite effector is composed of three effector domains. In some of any of the provided embodiments, the multipartite effector is set forth in any of SEQ ID NOS:397-418, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing. In some of any of the provided embodiments, the multipartite effector is set forth in any of SEQ ID NOS:411-418, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing. In some of any of the provided embodiments, the multipartite effector comprises, in the N-terminal to C-terminal direction, domains from FOXO3, FOXO3, and NCOA3. In some of any of the provided embodiments, the multipartite effector comprises the sequence set forth in SEQ ID NO:415, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the multipartite effector comprises, in the N-terminal to C-terminal direction, domains from NCOA3; FOXO3, and FOXO3. In some of any of the provided embodiments, the multipartite effector comprises the sequence set forth in SEQ ID NO:418, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the multipartite effector comprises, in the N-terminal to C-terminal direction, domains from NCOA3, FOXO3, and NCOA3. In some of any of the provided embodiments, the multipartite effector comprises the sequence set forth in SEQ ID NO:413, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the multipartite effector comprises, in the N-terminal to C-terminal direction, domains from NCOA2, FOXO3, and NCOA3. In some of any of the provided embodiments, the multipartite effector comprises the sequence set forth in SEQ ID NO:416, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the multipartite effector comprises, in the N-terminal to C-terminal direction, domains from PYGO1, FOXO3, and NCOA3. In some of any of the provided embodiments, the multipartite effector comprises the sequence set forth in SEQ ID NO:411, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
[0047] In some of any of the provided embodiments, the at least one epigenetic effector domain is fused to the N-terminus of the dCas protein or eZFP. In some of any of the provided embodiments, the at least one epigenetic effector domain is fused to the C-terminus of the dCas protein or eZFP. In some of any of the provided embodiments, the at least one epigenetic effector domain is fused to both the N- terminus and the C-terminus, of the dCas protein or eZFP. In some of any of the provided embodiments, the one or more linkers are in between any two of the components of the fusion protein, including the eZFP, any of the at least one effector domains, and the one or more NLS. In some of any of the provided embodiments, the one or more linkers connect the dCas protein and the at least one epigenetic effector domain.
[0048] In some of any of the provided embodiments, the fusion protein comprises the sequence set forth in SEQ ID NO:71 or 77, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto, or the sequence set forth in SEQ ID NO:71 or 77, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some of any of the provided embodiments, the fusion protein comprises the sequence set forth in any one of SEQ ID NOS:266-268 and 315-319, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
[0049] In some of any of the provided embodiments, the AAV vector is selected from the group consisting of: AAV1, AAV2, AAV3, AAV4, AAV5, AAV6, AAV7, AAV8, AAV9, AAV10, AAV11, AAV12, AAV-DJ, and AAVrh74. In some of any of the provided embodiments, the AAV vector is AAV6. In some of any of the provided embodiments, the AAV vector is AAV9. In some of any of the provided embodiments, the AAV vector is AAV-DJ. In some of any of the provided embodiments, the AAV vector is AAVrh74. In some of any of the provided embodiments, the vector exhibits tropism for a nervous system cell, optionally a neuron, a heart cell, optionally a cardiomyocyte, a skeletal muscle cell, a fibroblast, an induced pluripotent stem cell, and/or a cell derived from any of the foregoing, or for a combination of any of the foregoing cells. In some of any of the provided embodiments, the vector exhibits tropism for induced pluripotent stem cells. In some of any of the provided embodiments, the vector exhibits tropism for neurons and cardiomyocytes.
[0050] In some aspects, provided herein is a cell comprising any eZFP, any fusion protein, any polynucleotide, any plurality of polynucleotides, any vector, or any AAV vector provided herein, or a portion or component of any of the foregoing, or a combination of any of the foregoing.In some of any of the provided embodiments, the cell is a nervous system cell, optionally a neuron, a heart cell, optionally a cardiomyocyte, a skeletal muscle cell, a fibroblast, an induced pluripotent stem cell, and/or a cell derived from any of the foregoing. In some of any of the provided embodiments, the cell is from a subject that has or is suspected of having Friedreich’s ataxia (FA).
[0051] In some aspects, provided herein is a pharmaceutical composition comprising any eZFP, any fusion protein, any polynucleotide, any plurality of polynucleotides, any vector, any AAV vector
provided herein, or a portion or component of any of the foregoing, or a combination of any of the foregoing. In some of any of the provided embodiments, the pharmaceutical composition is for use in treating a disease, condition, or disorder in a subject. In some of any of the provided embodiments, the disease, condition, or disorder is Friedreich’s ataxia and/or a GAA trinucleotide repeat expansion in the FXN locus. In some of any of the provided embodiments, following administration of the pharmaceutical composition, the expression of FXN is increased in cells of the subject.
[0052] In some aspects, provided herein is a method for increasing the expression of FXN in a cell, the method comprising introducing into the cell: any eZFP, any fusion protein, any polynucleotide, any plurality of polynucleotides, any vector, any AAV vector, or any pharmaceutical composition provided herein, or a portion or component of any of the foregoing, or a combination of any of the foregoing. In some of any of the provided embodiments, the cell is from and/or in a subject that has or is suspected of having Friedreich’s ataxia. In some of any of the provided embodiments, the cell exhibits reduced expression of FXN in comparison to a reference cell from an individual not having Friedreich’s ataxia and/or a GAA trinucleotide expansion in the FXN locus. In some aspects, provided herein is a method for increasing the expression of FXN in a cell in a subject, the method comprising administering to the subject: any eZFP, any fusion protein, any polynucleotide, any plurality of polynucleotides, any vector, any AAV vector, or any pharmaceutical composition provided herein, or a portion or component of any of the foregoing, or a combination of any of the foregoing. In some aspects, provided herein is a method of treating a subject in need thereof, the method comprising administering to the subject: any eZFP, any fusion protein, any polynucleotide, any plurality of polynucleotides, any vector, any AAV vector, or any pharmaceutical composition provided herein, or a portion or component of any of the foregoing, or a combination of any of the foregoing.
[0053] In some of any of the provided embodiments, the subject has or is suspected of having Friedreich’s ataxia, and/or a GAA trinucleotide expansion in the FXN locus. In some of any of the provided embodiments, the introducing or administering is carried out in vivo or ex vivo. In some of any of the provided embodiments, the cell and/or subject exhibits reduced expression of FXN prior to performing the method. In some of any of the provided embodiments, the reduced expression of FXN is reduced in comparison to a reference individual not having Friedreich’s ataxia and/or a GAA trinucleotide repeat expansion in the FXN locus, and/or a reference cell therefrom. In some of any of the provided embodiments, the GAA trinucleotide repeat expansion is in a first intron of a FXN gene, and comprises at least 50, at least 60, at least 70, at least 80, at least 90, at least 100, at least 150, at least 200, at least 300, at least 400, at least 500, at least 600, at least 700, at least 800, at least 900, at least 1000, or more repeated GAA trinucleotides. In some of any of the provided embodiments, following the introducing or administering, the expression of FXN is increased in the cell and/or subject. In some of any of the provided embodiments, the expression of FXN is increased in the cell or cells of the subject by at least about 1.2-fold, 1.25-fold, 1.3-fold, 1.4-fold, 1.5-fold, 1.6-fold, 1.7-fold, 1.75-fold, 1.8-fold, 1.9- fold, 2-fold, 2.5-fold, 3-fold, 4-fold, or 5-fold; and/or the expression is increased by less than about 10-
fold, 9-fold, 8-fold, 7-fold or 6-fold. In some of any of the provided embodiments, the expression of FXN is increased in the cell or cells of the subject to a level that is at least at or about 30%, 40%, 50%, 60%, 70%, 75%, 80%, 85%, 90%, 95%, 100%, 105%, 110%, 120%, 125%, 150%, 175%, 200%, 225%, 250%, 300%, 400%, or 500%, of the expression level of FXN in a reference cell from an individual not having Friedreich’ s ataxia and/or a cell not having a GAA trinucleotide repeat expansion in the FXN gene. In some of any of the provided embodiments, the expression of FXN is increased in the cell or cells of the subject to a level that is less than at or about 200%, 300%, 400%, 500%, 600%, 700%, 800%, 900%, or 1000% of the expression level of FXN in a reference cell from an individual not having Friedreich’s ataxia and/or a cell not having a GAA trinucleotide repeat expansion in the FXN gene. In some of any of the provided embodiments, the expression is measured by the amount of mRNA encoding the FXN protein, and/or the amount of FXN protein. In some of any of the provided embodiments, the subject is a human. In some aspects, provided herein is a cell comprising an epigenetic modification produced by any method provided herein.
Brief Description of the Drawings
[0054] FIGS. 1A-1C show CRISPR-Cas mediated transcriptional activation of human frataxin in WT-iPSCs using dSaCas9-2xVP64 or dSpCas9-2xVP64 and gRNAs targeted to the frataxin promoter. FIG. 1A shows target locations of gRNAs from Table El, targeted to human frataxin gene promoter (Homo sapiens (human) genome assembly GRCh38 (hg38) chr9:69034622-69036670). FIGS. IB and 1C show expression of frataxin mRNA following transduction of WT-iPSCs with indicated gRNA and dSaCas9-2xVP64 (FIG. IB) or dSpCas9-2xVP64 (FIG. 1C), as assessed by RT-qPCR.
[0055] FIG. 2 shows CRISPR-Cas mediated transcriptional activation of human frataxin in iPSCs generated from Friedreich’s ataxia patients (FA-iPSCs) using dSaCas9-2xVP64 and promoter-targeting gRNA A or gRNA G, as assessed by RT-qPCR. Cell lines harbored expanded GAA trinucleotide repeats (left panel: 604/734 repeats; right panel: 867/867 repeats).
[0056] FIG. 3 shows frataxin protein expression levels in WT-iPSCs and FA-iPSCs as assessed by flow cytometry with a mouse monoclonal anti-frataxin AlexaFluor488-conjugated antibody (abeam abl56033). FA-iPSCs exhibit reduced frataxin expression in comparison to WT-iPSC control cells.
[0057] FIG. 4 shows scatterplot of results from sequencing analysis for screen of frataxin-saturating gRNA library. WT-iPSCs expressing dSaCas9-2xVP64 were transduced with pooled library of gRNAs and subsequently sorted by flow cytometry into populations representing top 10% and bottom 10% of cells based on frataxin protein expression. Populations were sequenced to identify gRNAs enriched in the frataxin-high population based on DESeq2. Each dot in the scatterplot represents a single gRNA. The y- axis represents log fold change in abundance of gRNA in frataxin-high versus frataxin-low population. The x-axis represents mean of normalized counts. The promoter-targeting gRNA A and enhancertargeting gRNA U (indicated in the figures) were significantly enriched in the frataxin-high population.
[0058] FIG. 5 shows locations of promoter-targeting gRNA A and enhancer-targeting gRNA U with
respect to the frataxin gene and indicators of chromatin state H3K4Mel, H3K4Me3, H3K27Ac, and DNase I hypersensitivity based on data from the Encyclopedia of DNA Elements (ENCODE). gRNA U targets a region (hg38 chr9:69, 027, 282-69, 028, 497) exhibiting hallmark chromatin signatures of an enhancer element, including H3K4Mel and DNase I hypersensitivity.
[0059] FIG. 6 shows CRISPR-Cas mediated transcriptional activation of frataxin in iPSCs, using dSaCas9-2xVP64 and promoter-targeting gRNA A or enhancer-targeting gRNA U compared to control gRNA or empty gRNA vector control, as assessed by RT-qPCR. iPSCs harbored normal GAA trinucleotide repeat lengths.
[0060] FIGS. 7A-7F show combinatorial modulation of frataxin mRNA expression with promotertargeting (gRNA A and gRNA G) and enhancer-targeting (gRNA U) gRNAs, as assessed by RT-qPCR. WT-iPSCs or FA-iPSCs with short or long trinucleotide repeats were used to create stable cell lines expressing promoter-targeting gRNA A or gRNA G. Cell lines were transduced with different combinations of dSaCas9-2xVP64 and gRNA, as indicated. Results are shown for cell lines WT-gRNA A-iPSC (FIG. 7A), WT-gRNA G-iPSC (FIG. 7B), FA(short)-gRNA A-iPSC (FIG. 7C), FA(short)- gRNA G-iPSC (FIG. 7D), FA(long)-gRNA A-iPSC (FIG. 7E), and FA(long)-gRNA G-iPSC (FIG. 7F).
[0061] FIG. 8 shows the rescue of frataxin mRNA expression with promoter- and enhancertargeting gRNAs in FA-iPSCs, with frataxin expression levels compared to WT-iPSCs, as assessed by RT-qPCR. FA-iPSCs stably expressing promoter-targeting gRNA A or gRNA G were transduced with different combinations of dSaCas9-2xVP64 and gRNA, as indicated.
[0062] FIGS. 9A-9C show CRISPR-Cas mediated activation of frataxin protein expression using promoter- and enhancer-targeting gRNAs, as assessed by ELISA. FIG. 9A shows frataxin protein expression in WT-iPSCs stably expressing gRNA G (WT-gRNA G-iPSCs), which were (a) left untreated, (b) transduced with dSaCas9-2xVP64 and a control gRNA, or (c) transduced with dSaCas9- 2xVP64 and the frataxin enhancer-targeting gRNA U. FIGS. 9B and 9C show rescue of frataxin protein expression in FA-iPSCs harboring short (FIG. 9B) or long (FIG. 9C) expanded trinucleotide GAA repeats in comparison to WT-iPSCs. FA-iPSCs stably expressing promoter-targeting gRNAs were transduced with different combinations of dSaCas9-2xVP64 and gRNA, as indicated.
[0063] FIGS. 10A and 10B show scatterplots of results from sequencing analysis for screen of dSaCas9 transcription activator and repressor fusion proteins. WT-iPSCs expressing frataxin promotertargeting gRNA A were transduced with pooled libraries of fusion proteins comprising fragments of nuclear localized proteins, fused to the N-terminus (FIG. 10A) or C-terminus (FIG. 10B) of dSaCas9. Transduced cells were subsequently sorted by flow cytometry into populations representing top 10% and bottom 10% of cells based on frataxin protein expression. Populations were sequenced to identify effectors enriched in the frataxin-high or frataxin-low populations based on DESeq2. Each dot in the scatterplots represents a single effector. Effectors in the top and bottom 10% of cells based on frataxin protein expression are indicated by black dots, and other effectors are indicated by gray dots. The y-axis represents log fold change in frataxin-high versus frataxin-low populations, x-axis represents mean of
normalized counts. Enriched effectors are highlighted in red, as activators (positive log fold change) and repressors (negative log fold change). N-terminal screen identified 9 activators and 211 repressors, C- terminal screen identified 5 activators and 208 repressors.
[0064] FIGS. 11A and 11B show transcriptional activation of frataxin in WT-iPSCs facilitated by dSaCas9 transcription activator N-terminal (FIG. 11A) and C-terminal (FIG. 11B) fusion proteins identified in the screen. WT-iPSCs stably expressing frataxin promoter-targeting gRNA A were transduced with dSaCas9 fusion proteins comprising indicated effectors, including positive control (2xVP64) and negative control peptides. Expression was assessed by RT-qPCR in comparison to negative control.
[0065] FIGS. 12A and 12B show transcriptional activation of frataxin in FA-iPSCs harboring long trinucleotide repeats facilitated by dSaCas9 transcription activator N-terminal (FIG. 12A) and C-terminal (FIG. 12B) fusion proteins identified in the screen. FA-iPSCs stably expressing frataxin promotertargeting gRNA A were transduced with dSaCas9 fusion proteins comprising indicated effectors, including positive control (2xVP64) and negative control peptides. Expression was assessed by RT- qPCR in comparison to negative control.
[0066] FIGS. 13A and 13B show frataxin mRNA expression (FIG. 13A) as assessed by qRT-PCR and frataxin protein expression (FIG. 13B) as assessed by ELISA, in cardiomyocytes derived from FA- iPSCs or WT-iPSCs for the indicated conditions. FA-iPSC-derived cardiomyocytes were transduced with dSaCas9-2xVP64 and indicated FXN-targeting gRNA(s). Negative control cells were transduced with dSaCas9-2xVP64 and a non-targeting gRNA (NT gRNA), or with a puromycin resistance cassette alone (puro control (FA)). Expression is assessed relative to WT control cells (puro control (WT)).
[0067] FIG. 14 shows frataxin mRNA expression as assessed by qRT-PCR in neurons derived from FA-iPSCs or WT-iPSCs for the indicated conditions. FA-iPSC-derived neurons were transduced with dSaCas9-2xVP64 and indicated FXN-targeting gRNA(s). Negative control cells were transduced with dSaCas9-2xVP64 and a non-targeting gRNA (NT gRNA), or with a puromycin resistance cassette alone (puro control (FA)). Expression is assessed relative to WT control cells (puro control (WT)).
[0068] FIG. 15 shows frataxin mRNA expression as assessed by qRT-PCR in non-human primate fibroblasts from Crab-eating monkey (Macaco fascicularis) and Rhesus monkey (Macaco mulatto) for the indicated conditions. Cells were transduced with dSaCas9-2xVP64 and indicated FXN-targeting gRNA(s). Negative control cells were transduced with dSaCas9-2xVP64 and a non-targeting gRNA (NT gRNA), or with a puromycin resistance cassette alone (puro control).
[0069] FIGS. 16A and 16B show frataxin protein expression as assessed by EEISA in the liver of a humanized FA mouse model or healthy control, treated with AAV9 vectors encoding dSaCas9-2xVP64 and indicated FXN-targeting gRNA(s). As controls, FA mouse model or healthy control mice were treated with AAV9 vectors encoding dSaCas9-2xVP64 and a non-targeting gRNA. FIG. 16A indicates FXN protein pg per pg of loaded protein into the assay [pg/pg protein] , and FIG. 16B indicates FXN protein levels normalized to the average FXN protein levels from healthy control mice.
[0070] FIGS. 17A and 17B show frataxin protein expression as assessed by ELISA in the heart of a humanized FA mouse model or healthy control, treated with AAV9 vectors encoding dSaCas9-2xVP64 and indicated FXN-targeting gRNA(s). As controls, FA mouse model or healthy control mice were treated with AAV9 vectors encoding dSaCas9-2xVP64 and a non-targeting gRNA. FIG. 17A indicates FXN protein pg per pg of loaded protein into the assay [pg/pg protein] , and FIG. 17B indicates FXN protein levels normalized to the average FXN protein levels from healthy control mice.
[0071] FIGS. 18A and 18B show succinate dehydrogenase (SDH) enzymatic activity in the heart of a humanized FA mouse model or healthy control, treated with AAV9 vectors encoding dSaCas9-2xVP64 and indicated FXN-targeting gRNA(s). As controls, FA mouse model or healthy control mice were treated with AAV9 vectors encoding dSaCas9-2xVP64 and a non-targeting gRNA. FIG. 18A indicates mU of SDH activity per mg loaded protein into the assay [mu/mg], and FIG. 18B indicates SDH activity normalized to the SDH activity from healthy control mice.
[0072] FIGS. 19A and 19B show AAV vector genome quantification as assessed by ddPCR, in the liver (FIG. 19A) and the heart (FIG. 19B) of the humanized FA mouse model or healthy control, treated with AAV9 vectors encoding dSaCas9-2xVP64 and indicated FXN-targeting gRNA(s). As controls, FA mouse model or healthy control mice were treated with AAV9 vectors encoding dSaCas9-2xVP64 and a non-targeting gRNA.
[0073] FIG. 20A shows frataxin protein expression in FA-iPSC-derived cardiomyocytes following AAV-DJ delivery of dSaCas9-2xVP64 and the indicated FXN-targeting gRNAs or a non-targeting gRNA (NT).
[0074] FIG. 20B shows frataxin mRNA expression in FA-iPSC-derived neurons following AAV-DJ delivery of dSaCas9-2xVP64 and the indicated non-targeting or FXN-targeting gRNA at various values for multiplicity of infection (MOI).
[0075] FIGS. 21A and 21B show the multiplicity of infection (MOI) and vector copy number (VCN) in FA-iPSC-derived cardiomyocytes (FIG. 21A) or FA-iPSC-derived neurons (FIG. 21B) following AAV-DJ delivery of dSaCas9-2xVP64 and the indicated non-targeting or FXN-targeting gRNA.
[0076] FIG. 22 shows a schematic illustrating an exemplary dSaCas9-tripartite effector fusion protein, with domains from FOXO3 and NCOA3. The first domain (labeled “effector”) can comprise different domains, as described in the Examples.
[0077] FIG. 23 shows frataxin protein expression in FA-iPSC-derived cardiomyocytes following AAV-DJ delivery of dSaCas9 fusion proteins with indicated FXN-targeting gRNA G or non-targeting gRNA (NT). Boxes indicating “tripartite effectors” indicate conditions with dSaCas9 fusion proteins with tripartite effectors comprising the indicated domain (e.g. FOXO3, NCOA2, NCOA3, or PYGO1), followed by a domain from FOXO3 and NCOA3, in the N- to C-terminal direction, e.g. as illustrated in FIG. 22.
[0078] FIGS. 24A-24C shows results from FA-iPSC-derived cardiomyocytes following AAV-DJ
delivery of the indicated dSaCas9 fusion proteins and gRNA G. Shown are MOI versus % of WT FXN protein expression (FIG. 24A), VCN versus % of WT FXN protein expression (FIG. 24B), or a summary table of the results (FIG. 24C).
[0079] FIG. 25 shows VCN versus % of WT FXN protein expression levels in FA-iPSC-derived cardiomyocytes following AAV-DJ delivery of dSaCas9 fusion proteins with the indicated effectors for transcriptional activation. Individual domain names (e.g. NCOA3) stand for tripartite effectors comprising the domain, followed by FOXO3 and NCOA3, e.g. as illustrated in FIG. 22.
[0080] FIGS. 26A and 26B show FXN protein expression levels (in comparison to WT control) in FA-iPSC-derived cardiomyocytes (FIG. 26A) or FA-iPSC-derived neurons (FIG. 26B) following AAV- DJ delivery of dSaCas9 fusion proteins with the indicated effectors for transcriptional activation paired with FXN-targeting gRNA G or non-targeting gRNA (NT). Individual domain names (e.g., FOXO3, NCOA2, NCOA3) stand for tripartite effectors comprising the domain, followed by FOXO3 and NCOA3, e.g. as illustrated in FIG. 22.
[0081] FIG. 27 shows FXN mRNA expression in FA-iPSC-derived cardiomyocytes, or similar cells genetically corrected for the FXN trinucleotide repeat expansion, following AAV-DJ delivery of dSaCas9-2xVP64 and indicated FXN-targeting gRNA or non-targeting gRNA (NT), using a a U6 or mini-U6 promoter for gRNA expression.
[0082] FIG. 28 shows FXN mRNA expression in FA-iPSC-derived cardiomyocytes following AAV-DJ delivery of dSaCas9-2xVP64 with or without a FLAG epitope tag and the indicated FXN- targeting or non-targeting gRNA.
[0083] FIG. 29 shows FXN mRNA expression at 48 hours after delivery in FA-iPSCs following delivery via electroporation of mRNA encoding eZFP-VP64 fusion proteins comprising the indicated eZFPs.
[0084] FIGS. 30A and 30B show FXN mRNA expression in FA-iPSCs following delivery via electroporation of mRNA encoding eZFP-VP64 fusion proteins comprising the indicated eZFPs, 48 hours (FIG. 30A) and 72 hours (FIG. 30B) after delivery.
[0085] FIG. 30C shows FXN protein expression (measured using ELISA) in FA-iPSCs following delivery via electroporation of mRNA encoding eZFP-VP64 fusion proteins comprising the indicated eZFPs at 72 hours after delivery.
[0086] FIG. 31 shows a map of genomic regions comprising a FXN promoter (top) and FXN enhancer (bottom) targeted by the indicated eZFPs and gRNAs.
[0087] FIG. 32 shows FXN mRNA expression (measured using RT-qPCR) 7 days after delivery in FA-iPSC-derived cardiomyocytes following AAV-DJ delivery of eZFP-VP64 fusion proteins comprising the indicated eZFPs.
[0088] FIG. 33 shows FXN mRNA expression (left), or VP64 mRNA expression (right) as a measure of ZFP-VP64 fusion protein expression, in FA-iPSC-derived cardiomyocytes following AAV- DJ delivery of eZFP-VP64 fusion proteins comprising the indicated eZFPs. Expression levels were
measured using RT-qPCR 7 days after delivery.
[0089| FIG. 34 shows FXN mRNA expression (left) or VP64 or Cas9 expression (right) as a measure of expression of the ZFP-VP64 fusion protein or dSaCas9-VP64 fusion protein, in FA-iPSC- derived cardiomyocytes following AAV-DJ delivery of either a) an eZFP-VP64 fusion protein comprising eZFP_A31 or b) dSaCas9-2xVP64 and gRNA G. Expression levels were measured using RT- qPCR 7 days after delivery.
[0090] FIG. 35 shows FXN mRNA expression in FA-iPSC-derived neurons following AAV-DJ delivery of ZFP-VP64 fusion proteins comprising the indicated eZFPs. Expression levels were measured using RT-qPCR 7 days after delivery.
[0091] FIG. 36 shows VP64 mRNA expression levels as a measure of expression of the indicated ZFP-VP64 fusion proteins, in FA-iPSC-derived neurons following AAV-DJ delivery of ZFP-VP64 fusion proteins. Expression levels were measured using RT-qPCR 7 days after delivery.
[0092] FIG. 37 shows FXN mRNA expression levels (as compared to WT cells) in FA-iPSC- derived cardiomyocytes following AAV-DJ delivery of a) the fusion proteins comprising eZFP_A31 and VP64 or the indicated tripartite effectors, or b) dSaCas9 fusion proteins comprising 2xVP64 or the indicated tripartite effectors with gRNA G. Expression levels were measured using RT-qPCR 7 days after delivery.
[0093] FIG. 38 shows FXN mRNA expression levels in FA-iPSC-derived neurons following AAV- DJ delivery of a) the fusion proteins comprising eZFP_A31 and VP64 or the indicated tripartite effectors, or b) dSaCas9 fusion proteins comprising 2xVP64 or the indicated tripartite effectors with gRNA G. Expression levels were measured using RT-qPCR 7 days after delivery.
[0094] FIG. 39 shows FXN mRNA expression levels in FA-iPSC-derived neurons (as compared to WT control cells) following AAV-DJ delivery of fusion proteins comprising eZFP_A31 and indicated tripartite effectors fused to the C-terminus or N-terminus of the eZFP (left). Schematics of each tested fusion protein are also shown (right). Expression levels were measured using RT-qPCR 7 days after delivery.
[0095] FIG. 40A and FIG. 40B show FXN expression levels in the liver of a humanized FA mouse model treated with AAV9 vectors encoding eZFP_A31 or dCas9 fusion proteins containing transcriptional activator VP64 or tripartite effector NFN. For vectors containing dCas9 fusion proteins, the vectors also encoded a FXN-targeting gRNA G. As controls, FA mouse model (FA model) or healthy control mice (WT) were treated with vehicle only or with AAV9 vectors encoding dCas9-VP64 or dCas9-NFN and a non-targeting (NT) gRNA. FIG. 40A shows FXN protein expression levels and FIG. 40B shows FXN mRNA expression levels relative to the healthy control mice.
[0096] FIG. 41A and FIG. 41B show FXN expression levels in the heart of a humanized FA mouse model treated with AAV9 vectors encoding eZFP_A31 or dCas9 fusion proteins containing transcriptional activator VP64 or tripartite effector NFN. For vectors containing dCas9 fusion proteins, the vectors also encoded a FXN-targeting gRNA G. As controls, a FA mouse model (FA model) or
healthy control mice (WT) were treated with vehicle only or with AAV9 vectors encoding dCas9-VP64 or dCas9-NFN and a non-targeting (NT) gRNA. FIG. 41A shows FXN protein expression levels and FIG. 41B shows FXN mRNA expression levels relative to healthy control mice.
[0097] FIG. 42A and FIG. 42B show FXN expression levels in the cerebellum of a humanized FA mouse model treated with AAV9 vectors encoding a dCas9 fusion protein containing transcriptional activator VP64 and a FXN-targeting gRNA G. As controls, a FA mouse model (FA model) or healthy control mice (WT) were treated with vehicle only or with AAV9 vectors encoding dCas9-VP64 or dCas9-NFN and a non-targeting (NT) gRNA. FIG. 42A shows FXN protein expression levels and FIG. 42B shows FXN mRNA expression levels relative to healthy control mice.
[0098] FIG. 43A and FIG. 43B show FXN expression levels following treatment with AAV9 vectors encoding an exemplary fusion protein containing eZFP_A31 and the tripartite effector NFN and the biodistribution of the AAV9 vectors in different tissues. As controls, a FA mouse model (FA model) or healthy control mice (WT) were treated with vehicle only. FXN expression levels were measured in the heart (FIG. 43A; left panel), liver (FIG. 43A; middle panel), and cerebellum (FIG. 43B; left panel). Biodistribution of the AAV9 vectors was also assessed relative to expression of heart TBP mRNA in heart and liver (FIG. 43A; right panel) or cerebellum (FIG. 43B; right panel).
[0099] FIG. 44A and FIG. 44B show schematics of exemplary AAVDJ constructs used to deliver a fusion protein containing the exemplary eZFP eZFP_A31 and the tripartite effector NFN. The original construct (FIG. 44A; top) included, from N-terminus to C-terminus, an elongation factor la short (EFS) promoter, an SV40 NLS (SEQ ID NO: 159), the exemplary eZFP_A31, another SV40 NLS (SEQ ID NO: 159), the tripartite effector NFN, and the poly(A) sequence from SpA. In a slightly modified construct (FIG. 44A; bottom), the SV40 NLS was substituted for a c-myc NLS (SEQ ID NO: 160). Two optimized constructs using either the SV40 or c-myc NLS (FIG. 44B; top and bottom, respectively) substituted the EFS promoter for either a CAG or EFla promoter and the SpA poly (A) sequence for a bGH poly(A) sequence and inserted a 5’ untranslated region (UTR) following the promoter.
[0100] FIG. 45 shows FXN expression levels in cardiomyocytes following delivery of AAVDJ vectors exemplified in FIG. 44A and FIG. 44B that used EFS, CAG, or EFla promoters to express eZFP_A31 fusion proteins containing either a transcriptional activator VP64 or tripartite effector NFN at different dosages.
[0101] FIG. 46 shows FXN expression levels in HEK293 cells three days following transfection using a screen of different eZFP fusion proteins containing different eZFPs, including a subset of eZFPs set forth in Table 2B, and the tripartite effector VP64. As controls, cells were (a) left untreated, (b) transfected with GFP, (c) transfected with an empty vector, (d) transfected with exemplary eZFP_A31 fusion proteins eZFP_A31-NFN or eZFP_A31-VP64, or (e) transfected with a dCas9-NFN fusion protein paired with FXN-targeting gRNA G (gG) or a non-targeting gRNA (gNT). Asterisks (*) indicate screened eZFPs chosen for further characterization.
[0102] FIG. 47 shows FXN expression levels in HEK293 cells three days following transfection
using a screen of different eZFP fusion proteins containing different eZFPs, including a subset of eZFPs set forth in Table 2B, and the tripartite effector NFN. As controls, cells were (a) left untreated, (b) transfected with GFP, (c) transfected with lipid only, (d) transfected with exemplary eZFP_A31 fusion proteins eZFP_A31-NFN or eZFP_A31-VP64, (e) transfected with a dCas9-NFN fusion protein paired with a non-targeting gRNA (gNT), or (f) treated with PBS. Asterisks (*) indicate screened eZFPs chosen for further characterization.
[0103] FIG. 48 shows FXN expression levels in HEK293 cells three days following transfection using a screen of different eZFP fusion proteins containing different eZFPs, including a subset of eZFPs set forth in Table 2B, and the tripartite effector NFN. As controls, cells were (a) transfected with GFP, (b) transfected with a dCas9-NFN fusion protein paired with a non-targeting gRNA (gNT), or (c) transfected with exemplary eZFP_A31-NFN or eZFP_A48-NFN fusion proteins. Asterisks (*) indicate screened eZFPs chosen for further characterization.
[0104] FIG. 49 shows FXN expression levels in HEK293 cells three days following transfection to validate top-performing eZFPs screened in FIG. 46 when fused to the tripartite effector NFN. As controls, cells were transfected with (a) GFP, (b) vector only, (c) a dCas9-NFN fusion proteins paired with a FXN-targeting gRNA G (gG) or non-targeting gRNA (gNT), or (d) exemplary eZFP_A31 fusion proteins containing either VP64 or NFN.
[0105] FIG. 50 shows FXN expression levels in FA patient fibroblasts (relative to fibroblasts transfected with GFP) to validate eZFP-NFN fusion proteins identified in previous screens. Transfection with GFP, exemplary eZFP_A31-VP64 fusion protein, or exemplary eZFP_31-NFN fusion protein were used as controls.
[0106] FIG. 51 shows FXN expression levels in FA patient fibroblasts (relative to fibroblasts transfected with GFP) to validate eZFP-NFN fusion proteins identified in previous screens. Controls included leaving cells untreated or transfecting cells with either GFP or exemplary eZFP_A31-NFN fusion proteins.
[0107] FIG. 52 depicts the level of FXN expression of different screened eZFPs based on target site. All target sites were within the FXN promoter and position is depicted relative to the transcriptional start site (TSS). The target sites of exemplary eZFP_A48 and eZFP_A31 are indicated.
[0108] FIG. 53 shows FXN expression levels in FA-cardiomyocyes following treatment with AAVDJ vectors using either CAG or EFS promoters to express exemplary eZFP-NFN fusion proteins at different doses (3E3, 3E4, or 3E5). As negative controls, FA-cardiomyocytes were either not transduced or transduced with AAVDJ vectors encoding (a) a fusion protein containing a non-targeting eZFP (eZFP- NT) and the tripartite effector NFN (eZFP-NT-NFN), (b) dCas9-NFN fusion protein paired with a nontargeting gRNA (gNT), or (c) GFP. As positive controls, genetically correct (GC) FA-cardiomyocytes were transduced with AAVDJ vectors encoding GFP or FA-cardiomyocytes were transduced with AAVDJ vectors encoding (a) dCas9-NFN fusion protein paired with FXN-targeting gRNA G (gG), or (b) eZFP_A31 fusion proteins containing either the tripartite effector NFN or the transcriptional activator
VP64.
Detailed Description
[0109] Provided herein are engineered zinc finger proteins (eZFPs), eZFP fusion proteins, and DNA-targeting systems that bind to or target a frataxin (FXN) locus. In some aspects, the DNA-targeting systems include fusion proteins. In some aspects, the DNA-targeting systems include guide RNAs (gRNAs). In some aspects, the DNA-targeting systems include fusion proteins and gRNAs. Provided herein are compositions, such as engineered zinc finger proteins (eZFPs), eZFP fusion proteins, DNA- targeting systems, including fusion proteins, gRNAs, and pluralities and combinations thereof, that bind to or target a FXN locus. Also provided are fusion proteins, such as eZFP fusion proteins and dCas fusion proteins that bind to or target FXN. Also provided are gRNAs that bind to or target FXN. In some aspects, the provided eZFPs, eZFP fusion proteins, DNA-targeting systems, including fusion proteins, and/or gRNAs, bind to, target, and/or modulate the expression of FXN. Also provided are polynucleotides, vectors, cells, and pluralities and combinations thereof, that encode or comprise the eZFPs, eZFP fusion proteins, DNA-targeting systems, fusion proteins, gRNAs or components thereof.
[0110] Also provided are methods and uses related to any of the provided compositions and combinations, for example, in modulating the expression of FXN, and/or in the treatment of diseases or disorders associated with reduced activity, mutation and/or dysregulation of expression of FXN, such as Friedreich’s ataxia (FA). In some aspects, also provided are methods and uses related to any of the provided compositions and combinations, for example, in modulating the expression of FXN, and/or in the treatment or therapy of diseases or disorders associated with the activity, function or expression, for example dysregulation or reduced activity, function or expression of FXN, such as FA.
[0111 ] In some aspects, the provided embodiments are based on an observation described herein that the level of a human FXN locus expression in cells from FA patients, including in induced pluripotent stem cells (iPSCs) generated from FA patient cells, can be increased or restored, for example using eZFP fusion proteins, or using an exemplary DNA-targeting system comprising a deactivated Cas9 (dCas9)-transcriptional activator fusion protein and a gRNA targeting a promoter region or an enhancer region of a human FXN locus. In addition, the results described herein also show that combinations of two or more DNA-targeting systems targeting different target sites of a human FXN locus can result in a synergistic increase in expression of a human FXN locus. The provided embodiments are also based on an observation that certain fusion proteins comprising a DNA-targeting domain and an effector domain is identified based on screening a library of effector domains for their effect in increased expression of a human FXN locus. The embodiments described herein demonstrate consistent and effective increase or restoration of FXN expression, in cells from patients with FA, at both the mRNA and protein level, supporting the utility of the approaches in treating FA or other diseases or disorders that are associated with reduced activity, mutation and/or dysregulation of expression of FXN.
[0112] Certain genetic development disorders, including FA, are associated with reduced activity,
mutation and/or dysregulation of expression of a frataxin (FXN) gene. FA is an autosomal recessive neurodegenerative and cardiac disease, is caused by a trinucleotide repeat expansion mutation in the FXN locus. FA can result in ataxia, areflexia, loss of vibratory sense and proprioception, dysarthria, cardiomyopathy and/or associated arrhythmias, among other symptoms. Existing treatment of FA is directed towards symptoms relief and providing support. Treatments that address the fundamental etiology and disease mechanism are needed. Provided are embodiments, including engineered zinc finger proteins (eZFPs), eZFP fusion proteins, DNA-targeting systems, fusion proteins, guide RNAs (gRNAs), polynucleotides, vectors, cells, kits, and pluralities and combinations thereof, and methods and uses thereof, that meet such needs.
[0113] In some aspects, the provided embodiments offer an advantage of targeting regulatory DNA elements of a frataxin locus within a particular genomic region, such as an enhancer region, for modulating transcription. In some aspects, the provided embodiments offer an advantage of facilitating controlled, additive and/or synergistic activation of FXN by targeting two or more sites within regulatory DNA elements of FXN. In some aspects, the provided embodiments offer an advantage of increasing FXN expression to a level that is therapeutically relevant for subjects having a disease or disorder that involve the activity, function or expression of FXN, such as FA. In some aspects, the provided embodiments also offer an advantage of providing various effector domains which are capable of inducing transcription activation, for example, at a particular target locus such as FXN.
[0114] In certain aspects, the provided embodiments offer the ability to fine tune and tightly regulate the level of expression and/or activity of frataxin in a cell or a subject. As described further below, the control of the expression and/or activity of frataxin at a particular level or within an optimal window is critical for the survival and normal function of the subject, as the reduction of expression can result in diseases or disorders such as Friedreich’s Ataxia (FA) and in some cases, substantial overexpression can result in toxicity, organ dysfunction, and reduction of life span. Accordingly, the level of expression and/or activity of frataxin must be fine-tuned to be within an optimal window. Even for the treatment of diseases or disorders associated with reduced expression and/or activity of frataxin, such as in FA, there is a need to tightly regulate the expression and/or activity of frataxin and avoid overexpression. The provided embodiments permit such fine tuning of expression of FXN without the need for introducing additional copies of FXN into the cell, which could result in toxic overexpression of FXN.
[0115] Frataxin is a protein (exemplary amino acid sequences of human Frataxin Isoform 1: NCBI NM_000144.4, NP_000135 (210 aa); exemplary amino acid sequences of human Frataxin Isoform 2: NM_181425, NP_852090 (196 aa); and exemplary amino acid sequences of human Frataxin Isoform 3: NM_001161706, NP_001155178, (171 aa), Uniprot Q16595; ENTREZ 2395; Ensembl ENSG00000165060; OMIM: 606829) found in cells throughout the body, with the highest levels in tissues with a high metabolic rate including heart, neurons, spinal cord, liver, pancreas (Langerhans cells), and muscles used for voluntary movement (skeletal muscles). Within cells, frataxin is mainly found in mitochondria. In some aspects, frataxin promotes the biosynthesis of heme as well as the
assembly and repair of iron-sulfur clusters by delivering Fe2+ to proteins involved in these pathways. In some aspects, frataxin also plays a primary role in the protection against oxidative stress through its ability to catalyze the oxidation of Fe2+ to Fe3+ and to store large amounts of the metal in the form of a ferrihydrite mineral. In some aspects, frataxin is processed in two steps by mitochondrial processing peptidase (MPP). MPP first cleaves the precursor to intermediate form and subsequently converts the intermediate to a mature protein. Thus, in cells, 3 forms exist. For isoform 1, these forms are frataxin (56- 210); frataxin (78-210) and frataxin (81-210), which is the main form of mature frataxin.
[0116] Friedreich’ s ataxia (FA) is a monogenic autosomal recessive neurodegenerative and cardiac disease, associated with trinucleotide repeat expansion mutations in the first intron of the frataxin gene (FXN), which is located on the long arm of chromosome 9. The vast majority of FA patients are homozygous for guanine-adenine-adenine (GAA) trinucleotide repeat expansions in the first intron of FXN. FA is a degenerative neuromuscular disorder, and can result in ataxia, areflexia, loss of vibratory sense and proprioception, dysarthria, cardiomyopathy and/or associated arrhythmias, among other symptoms, and death. The mutation leads to reduced expression of the frataxin mRNA and protein. Frataxin is essential for proper functioning of mitochondria. As noted above, Frataxin is involved in the removal of iron and when Frataxin is reduced, iron builds up and causes free radical damage. Nerve and muscle cells are particularly sensitive to these deleterious effects. FA occurs in approximately 1 in 50,000 persons in European populations but is much more frequent in the province of Quebec in Canada, because of founder effects. Males and females are affected equally. In the classic form, FA symptoms appear during or before the second decade of life. FA is characterized by ataxia, areflexia, loss of vibratory sense and proprioception and dysarthria. Moreover, FA patients often have systemic involvement, with cardiomyopathy, diabetes mellitus, and scoliosis. Early death can result from cardiomyopathy or associated arrhythmias. Degeneration of the dorsal root ganglion cells, their ascending dorsal spinal columns, and the spinocerebellar tracts results in a progressive sensory ataxia. Many patients are wheelchair bound by their third decade of life. Associated oculomotor problems include optic atrophy, square-wave jerks, and difficulty with fixation. Importantly, cognitive abilities are relatively spared. However, many patients suffer from depression.
[0U7| The class of mutation most commonly associated with FA is unstable hyper-expansion of a GAA trinucleotide repeat located in the first intron of the frataxin gene. In normal subjects, there are approximately 6-34 repeats, whereas expansions associated with FA are typically 150 or more repeats, and may vary from 44 to 1700 repeats, with most abnormal alleles ranging from 600 to 900 repeats. Increased trinucleotide repeat expansion lengths are associated with decreased FXN levels, earlier onset of disease, and increased disease severity. Patients with fewer repeats (150-200) have milder symptoms than those with longer sections with more triplet repeats (350 to 650). In some severely affected patients there are up to 1700 repeats. Since frataxin expansion mutations are located in an intron, the amino acid sequence of the frataxin protein is not altered. However, approximately 1-3% of FA patients are compound heterozygotes with an expansion on one allele and a conventional mutation (e.g. a missense,
nonsense, or deletion mutation) on the other. Some patients with a missense mutation can have less severe symptoms because the mutated protein in still functional. FA symptoms are not observed in heterozygous carriers.
[0118] In some aspects, the term "trinucleotide repeat expansion" means a series of three bases (for example, GAA) repeated at least twice. In certain examples, the trinucleotide repeat expansion may be located in intron 1 of a FXN locus, gene or nucleic acid. In certain examples, a pathogenic trinucleotide repeat expansion includes at least 66 or 70 repeats of GAA in a FXN nucleic acid and is associated with disease. In other examples, a pathogenic trinucleotide repeat expansion includes at least 67, 68, 69, 70, 75, 80, 85, 90, 95, 100, 150, 200, 250, 300, 400, 500, 800, 1000 or more repeats. In certain examples, the repeats are consecutive. In certain examples, the repeats are interrupted by one or more nucleobases. In certain examples, a wild- type trinucleotide repeat expansion includes 12 or fewer repeats of GAA in a FXN nucleic acid. In other examples, a wild-type trinucleotide repeat expansion includes 11, 10, 9, 8, 7, 6, 5, 4, 3, 2, or 1 repeat.
[0119] FXN has been associated with diseases and disorders such as, but not limited to, Alzheimer's Disease, Amyotrophic Lateral Sclerosis, Apraxias, Ataxia, Ataxia Telangiectasia, Hereditary Ataxias, Bloom Syndrome, Brain Neoplasms, Malignant tumor of colon, Dilated Cardiomyopathy, Hypertrophic Cardiomyopathy, Cerebellar Ataxia, Cystic Fibrosis, Diabetes, Diabetes Mellitus, Non-Insulin- Dependent Diabetes Mellitus, Dysarthria, Dystonia, Fragile X Syndrome, Friedreich's Ataxia, Heart Diseases, Cardiomegaly, Hemochromatosis, Herpes Simplex Infections, Huntington Disease, Liver neoplasms, Machado-Joseph Disease, Metabolic Diseases, Myocardial Infarction, Myotonic Dystrophy, nervous system disorder, Neuroblastoma, Neuromuscular Diseases, Pallor, Parkinson Disease, Peripheral Neuropathy, Protein Deficiency, Restless Legs Syndrome, Schizophrenia, unspecified Scoliosis, Hereditary Spastic Paraplegia, Spinocerebellar Ataxia, Left Ventricular Hypertrophy, Sensory neuropathy, Tumor Progression, Neurologic Symptoms, Paroxysmal atrial fibrillation, Hypoalbuminemia, Impaired glucose tolerance, Iron Overload, Adenocarcinoma of colon, Depletion of mitochondrial DNA, Ventricular septal hypertrophy, Malignant neoplasm of prostate, Hereditary hemochromatosis, Dystonia Disorders, Congenital Myotonic Dystrophy, Spastic, Neurodegenerative Disorders, Congenital scoliosis, Colon Carcinoma, Central neuroblastoma, Acquired scoliosis, cardiac symptom, Appendicular Ataxia, Mitochondrial Diseases, Heredodegenerative Nervous System Disorders, Spinocerebellar Ataxia Type 1, Cardiomyopathies, Ceruloplasmin deficiency, Hypertrophic Cardiomyopathy Familial, Degenerative disorder, Head titubation, Non-Neoplastic Disorder, X-Linked Bulbo-Spinal Atrophy, Fragile X Tremor/ Ataxia Syndrome, Friedreich's Ataxia With Retained Reflexes, Ataxia With Vitamin E Deficiency, Spinocerebellar Ataxia Autosomal Recessive 1, Friedreich's Ataxia 1, Hereditary Neurodegenerative Disorder, and Spastic Paraplegia Type 7. The FXN gene is also associated with a number of other disorders, including hereditary ataxia, X-linked sideroblastic anemia with ataxia, tabes dorsalis, spinocerebellar degeneration, tertiary neurosyphilis, a transferrinemia, spinocerebellar ataxia, scoliosis, hemochromatosis, fragile X syndrome, mitochondrial disorders and
cardiomyopathy. Modulating expression of the FXN gene using any of the methods described herein may be used to treat, prevent and/or mitigate the symptoms of the diseases and disorders described herein.
[0120] Frataxin is a mitochondrial iron chaperone suggested to be important for iron-sulfur processing. This protein is found throughout the human body, but is enriched in the heart, spinal cord, liver, pancreas and muscles. Expansions of a trinucleotide GA A repeat region in the FXN gene, and subsequent frataxin protein deficiency, causes Friedreich's Ataxia. The GAA repeat region is located in the middle of an Alu element in the first intron of the FXN gene. In most people, the number of GAA repeats in the FXN gene is fewer than 12. Individuals with 12-33 uninterrupted GAA repeats are said to be asymptomatic. However, as these repeats are unstable and more likely to expand during meiosis, such individuals are at increased risk of having affected children. In people with Friedreich's Ataxia, the GAA segment is abnormally repeated from 66 to over 1,000 times. The number of repeats in the gene correlates with the age of onset and severity of the disease. Individuals with fewer than 300 GAA repeats tend to have later symptom onset (after age 25) than those with larger GAA trinucleotide repeats (e.g. 600 to 900 repeats). The abnormal repeat expansion can result in an RNA processing defect, which leads to dysregulation of translation and reduced amount of the FXN protein in cells.
[0121] FA is usually diagnosed in the first or second decade and affects 1 in 50,000 people in the United States. Friedreich's Ataxia is a progressive movement disorder characterized by loss of strength and sensation, muscle stiffness and impaired speech. Individuals with Friedreich's Ataxia may also have cardiomyopathy, diabetes, vision or hearing loss, and/or scoliosis. Currently there is no treatment for Friedreich's Ataxia, only symptom management.
[0122] Provided herein are embodiments that meet such needs.
[0123] All publications, including patent documents, scientific articles and databases, referred to in this application are incorporated by reference in their entirety for all purposes to the same extent as if each individual publication were individually incorporated by reference. If a definition set forth herein is contrary to or otherwise inconsistent with a definition set forth in the patents, applications, published applications and other publications that are herein incorporated by reference, the definition set forth herein prevails over the definition that is incorporated herein by reference.
[0124] The section headings used herein are for organizational purposes only and are not to be construed as limiting the subject matter described.
I. Engineered Zinc Finger Proteins (eZFPs)
[0125] In some aspects, provided herein are zinc finger proteins (ZFPs), such as engineered zinc finger proteins (eZFPs). In some embodiments, the eZFPs are capable of binding to, or bind to, a target site in a FXN locus, such as a regulatory element of a FXN locus. In some aspects, the eZFP can facilitate specific targeting of effector domains for transcriptional activation to the FXN locus, for example for gene-specific transcriptional activation of FXN. In some embodiments, provided herein are fusion proteins comprising the eZFP and one or more other elements, such as the effector domains for
transcriptional activation. Thus, in some aspects the eZFP facilitates increased FXN expression, for example in connection with compositions and methods for treating a disease or disorder associated with FXN expression, such as Friedreich’s ataxia (FA).
In some embodiments, a zinc finger protein (ZFP), a zinc finger DNA binding protein, or zinc finger DNA binding domain, is a protein, or a domain within a larger protein, that binds DNA in a sequence-specific manner through one or more zinc fingers, which are regions of amino acid sequence within the binding domain, having a structure that is stabilized through coordination of a zinc ion. Among the ZFPs are artificial, or engineered, ZFPs (eZFPs), comprising ZFP domains targeting specific DNA sequences, typically 9-18 nucleotides long, generated by assembly of individual zinc fingers. ZFPs include those in which a single finger domain is approximately 30 amino acids in length and contains an alpha helix containing two invariant histidine residues coordinated through zinc with two cysteines of a single beta turn, and having two, three, four, five, or six fingers. Generally, sequence-specificity of a ZFP may be altered by making amino acid substitutions at the four helix positions (-1, 2, 3, and 6) on a zinc finger recognition helix, also called a zinc finger recognition region. Thus, for example, a ZFP or ZFP-containing molecule, such as a fusion protein, can be non-naturally occurring, e.g., is engineered to bind to a target site of choice.
[0126] In some embodiments, zinc fingers can be custom-designed (i.e. designed by the user), and/or obtained from a commercial source. Various methods for designing zinc finger proteins are available. For example, methods for designing zinc finger proteins to bind to a target DNA sequence of interest are described, for example in Liu, Q. et al., PNAS, 94(l l):5525-30 (1997); Wright, D.A. et al., Nat. Protoc., l(3):1637-52 (2006); Gersbach, C.A. et al., Acc. Chem. Res., 47(8):2309-18 (2014); Bhakta M.S. et al., Methods Mol. Biol., 649:3-30 (2010); and Gaj et al., Trends Biotechnol, 31(7):397-405 (2013). In addition, various web-based tools for designing zinc finger proteins to bind to a DNA target sequence of interest are publicly available. See, for example, the Zinc Finger Tools design web site from Scripps available on the world wide web at scripps.edu/barbas/zfdesign/zfdesignhome.php. Various commercial services for designing zinc finger proteins to bind to a DNA target sequence of interest are also available. See, for example, the commercially available services or kits offered by Creative Biolabs (world wide web at creative-biolabs.com/Design-and-Synthesis-of-Artificial-Zinc-Finger-Proteins.html), the Zinc Finger Consortium Modular Assembly Kit available from Addgene (world wide web at addgene.org/kits/zfc-modular-assembly/), or the CompoZr Custom ZFN Service from Sigma Aldrich (world wide web at sigmaaldrich.com/life-science/zinc-finger-nuclease-technology/custom-zfn.html).
[0127] In some embodiments, the target site for an eZFP provided herein is in a FXN promoter. In some embodiments, the target site is within a target region spanning the genomic coordinates chr9:69, 034, 900-69, 035, 900 from hg38 (SEQ ID NO:430). In some embodiments, the target site is within a target region spanning the genomic coordinates chr9:69, 035, 300-69-035, 800 from hg38. In some embodiments, the target site is within a target region spanning the genomic coordinates chr9:69,035,350- 69,035,450 from hg38. In some embodiments, the target site is within a target region spanning the
genomic coordinates chr9:69, 035, 400-69, 035, 450 from hg38. In some embodiments, the target site is within a target region spanning the genomic coordinates chr9:69, 035, 530-69, 035, 580 from hg38. In some embodiments, the target site is within a target region spanning the genomic coordinates chr9:69,035,675- 69,035,725 from hg38.
[0128] In some embodiments, the target site for an eZFP provided herein is in a FXN enhancer. In some embodiments, the target site is within a target region spanning the genomic coordinates chr9:69, 027, 282-69, 028, 497 from hg38 (SEQ ID NO:431). In some embodiments, the target site is within a target region spanning the genomic coordinates chr9:69, 027, 615-69, 028, 101 from hg38. In some embodiments, the target site is within a target region spanning the genomic coordinates chr9:69,027,775- 69,027,875 from hg38. In some embodiments, the target site is within a target region spanning the genomic coordinates chr9:69, 027, 795-69, 027, 845 from hg38.
[0129] In some embodiments, the target site for an eZFP provided herein comprises the nucleotide sequence set forth in any one of SEQ ID NOS:269-300 and 583-600, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the nucleotide sequence set forth in any one of SEQ ID NOS:269- 300 and 583-600. In some embodiments, the target site is comprised in double-stranded DNA, such as genomic DNA. In some embodiments, the target site is double-stranded DNA, such as genomic DNA. In some embodiments, the eZFP is capable of binding to the target site. In some embodiments, the eZFP binds to the target site. In some embodiments, the binding is target-specific. For example, in some embodiments, an eZFP binds to the target site, and not to other sites comprising different sequences. For example, in some embodiments, an individual eZFP disclosed herein binds to the target site set forth in SEQ ID NO:299, and does not bind to a different target site, such as the target site set forth in SEQ ID NO:269. In some embodiments, the target site for an eZFP provided herein comprises a sequence set forth in Table 1.
[0130] In some embodiments, the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:272, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the sequence set forth in SEQ ID NO: 272.
[0131] In some embodiments, the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:277, a contiguous portion thereof of at least 12 nt, or a complementary
sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the sequence set forth in SEQ ID NO: 277.
[0132] In some embodiments, the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:280, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the sequence set forth in SEQ ID NO: 280.
[0133] In some embodiments, the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:281, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the sequence set forth in SEQ ID NO: 281.
[01341 In some embodiments, the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:283, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the sequence set forth in SEQ ID NO: 283.
[0135] In some embodiments, the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:290, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the sequence set forth in SEQ ID NO: 290.
[0136] In some embodiments, the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:299, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the sequence set forth in SEQ ID NO: 299.
[0137] In some embodiments, the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:583, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the sequence set forth in SEQ ID NO:583.
[0138] In some embodiments, the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:584, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the sequence set forth in SEQ ID NO:584.
[0139] In some embodiments, the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:585, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the sequence set forth in SEQ ID NO:585.
[0140] In some embodiments, the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:586, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein
comprises the sequence set forth in SEQ ID NO:586.
[0141] In some embodiments, the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:587, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the sequence set forth in SEQ ID NO:587.
[0142] In some embodiments, the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:588, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the sequence set forth in SEQ ID NO:588.
[0143] In some embodiments, the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:589, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the sequence set forth in SEQ ID NO:589.
[0144] In some embodiments, the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:590, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the sequence set forth in SEQ ID NO:590.
[0145] In some embodiments, the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:591, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the sequence set forth in SEQ ID NO:591.
[0146] In some embodiments, the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:592, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the sequence set forth in SEQ ID NO:592.
[0147] In some embodiments, the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:593, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the sequence set forth in SEQ ID NO:593.
[0148] In some embodiments, the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:594, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the sequence set forth in SEQ ID NO:594.
[0149] In some embodiments, the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:595, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the sequence set forth in SEQ ID NO:595.
[0150] In some embodiments, the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:596, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the sequence set forth in SEQ ID NO:596.
[ 151] In some embodiments, the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:597, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the sequence set forth in SEQ ID NO:597.
[0152] In some embodiments, the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:598, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the sequence set forth in SEQ ID NO:598.
[0153] In some embodiments, the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:599, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the sequence set forth in SEQ ID NO:599.
[0154] In some embodiments, the target site for an eZFP provided herein comprises the nucleotide sequence set forth in SEQ ID NO:600, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site for an eZFP provided herein comprises the sequence set forth in SEQ ID NO:600.
[0155] In some embodiments, the eZFP comprises multiple zinc fingers. In some embodiments, each zinc finger comprises a recognition region. In some embodiments, the recognition regions together facilitate sequence-specific binding of the eZFP, for example to a specific target site. In some embodiments, the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding recognition region Fl through F6, which facilitate sequencespecific binding to a specific target site.
[0156] In some embodiments, characteristics of eZFPs targeting specific target sites provided herein are shown in Table 2A and Table 2B. In some embodiments, the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, each comprising a corresponding recognition region F1-F6, as shown in Table 2A and Table 2B. In some embodiments, the recognition regions F1-F6 facilitate specific binding to the indicated target site sequence in Table 2A and Table 2B. In some embodiments, the eZFP comprises an amino acid sequence comprising the recognition regions, as shown in Table 2A and Table 2B. In some embodiments, the eZFP can be encoded by a DNA sequence as shown in Table 2A and Table 2B.
[0157] In some embodiments, provided herein is an eZFP, such as eZFP_A04 as described herein. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:272, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:272. In some embodiments, the target site is double-stranded DNA. In some embodiments, the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino
acid sequence of each zinc finger recognition region is as follows: Fl: QSGNLAR (SEQ ID NO:341); F2: QKVNRAG (SEQ ID NO:342); F3: DRSNLSR (SEQ ID NO:343); F4: QSGHLSR (SEQ ID NO:344); F5: TSGHLSR (SEQ ID NO:345); F6: RSDALAR (SEQ ID NO:346). In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:301, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:301. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:308, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:308.
[01583 In some embodiments, provided herein is an eZFP, such as eZFP_A09 as described herein. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:277, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:277. In some embodiments, the target site is double-stranded DNA. In some embodiments, the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: RSDNLSE (SEQ ID NO:347); F2: KSWSRYK (SEQ ID NO:348); F3: TSGSLSR (SEQ ID NO:349); F4: RSDALAR (SEQ ID NO:350); F5: RSDNLSV (SEQ ID NO:351); F6: FSSCRSA (SEQ ID NO:352). In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:302, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:302. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:309, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:309.
[01593 In some embodiments, provided herein is an eZFP, such as eZFP_A12 as described herein. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:280, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:280. In some embodiments, the target site is double-stranded DNA. In some embodiments, the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: TSGNLTR (SEQ ID NO:353); F2: EQTTRDK (SEQ ID NO:354); F3: RSANLAR (SEQ ID NO:355); F4: RLDNRTA (SEQ ID NO:356); F5: DSSHRTR (SEQ ID NO:357); and F6: RKYYLAK (SEQ ID NO:358). In some embodiments, the
eZFP comprises the amino acid sequence set forth in SEQ ID NO:303, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:303. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:310, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:310.
[0160] In some embodiments, provided herein is an eZFP, such as eZFP_A13 as described herein. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:281, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:281. In some embodiments, the target site is double-stranded DNA. In some embodiments, the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: RSAHESR (SEQ ID NO:359); F2: DRSDESR (SEQ ID NO:360); F3: RSDHESV (SEQ ID NO:361); F4: RSDVRKT (SEQ ID NO:362); F5: QSGAEAR (SEQ ID NO:363); and F6: RKYYLAK (SEQ ID NO:364). In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:304, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:304. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:311, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:311.
[01 1] In some embodiments, provided herein is an eZFP, such as eZFP_A15 as described herein. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:283, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:283. In some embodiments, the target site is double-stranded DNA. In some embodiments, the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: RSAHESR (SEQ ID NO:365); F2: RSDAEAR (SEQ ID NO:366); F3: ATSNRSA (SEQ ID NO:367); F4: RSAHESR (SEQ ID NO:368); F5: TSGSLSR (SEQ ID NO:369); and F6: QSGDLTR (SEQ ID NO:370). In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:305, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID
NO:305. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:312, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:312.
[0162] In some embodiments, provided herein is an eZFP, such as eZFP_A22 as described herein. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:290, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:290. In some embodiments, the target site is double-stranded DNA. In some embodiments, the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: QSGDETR (SEQ ID NO:371); F2: QSSDERR (SEQ ID NO:372); F3: RSDNESE (SEQ ID NO:373); F4: SSRNEAS (SEQ ID NO:374); F5: DRSHETR (SEQ ID NO:375); and F6: RSDDLTR (SEQ ID NO:376). In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:306, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:306. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:313, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:313.
[0163] In some embodiments, provided herein is an eZFP, such as eZFP_A31 as described herein. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:299, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:299. In some embodiments, the target site is double-stranded DNA. In some embodiments, the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: ERHHETR (SEQ ID NO:377); F2: QSAHEKA (SEQ ID NO:378); F3: EPQTEQR (SEQ ID NO:379); F4: QNATRTK (SEQ ID NO:380); F5: QSSHLTR (SEQ ID NO:381); and F6: RSDHLSR (SEQ ID NO:382). In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:307, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:307. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:314, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP is encoded by the nucleotide sequence
set forth in SEQ ID NOG 14.
[0164] In some embodiments, provided herein is an eZFP, such as eZFP_A40 as described herein. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:583, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:583. In some embodiments, the target site is double-stranded DNA. In some embodiments, the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: RSDSLLR (SEQ ID NO:475); F2: TSSNRKT (SEQ ID NO:476); F3: RSAHLSR (SEQ ID NO:477); F4: TSGSLTR (SEQ ID NO:478); F5: QSGDLTR (SEQ ID NO:479); and F6: QWGTRYR (SEQ ID NO:480). In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:439, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:439. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:457, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:457.
[0165] In some embodiments, provided herein is an eZFP, such as eZFP_A41 as described herein. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:584, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:584. In some embodiments, the target site is double-stranded DNA. In some embodiments, the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: QARHLTC (SEQ ID NO:481); F2: QSGHLSR (SEQ ID NO:482); F3: RSDVLSE (SEQ ID NO:483); F4: KHSTRRV (SEQ ID NO:484); F5: QSSDLSR (SEQ ID NO:485); and F6: WKWNLRA (SEQ ID NO:486). In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:440, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:440. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:458, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:458.
[0166] In some embodiments, provided herein is an eZFP, such as eZFP_A42 as described herein. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ
ID NO:585, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:585. In some embodiments, the target site is double-stranded DNA. In some embodiments, the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: RSDNEAR (SEQ ID NO:487); F2: WRGDRVK (SEQ ID NO:488); F3: YKHVESD (SEQ ID NO:489); F4: TSGSETR (SEQ ID NO:490); F5: QSGNLAR (SEQ ID NO:491); and F6: RARDLSK (SEQ ID NO:492). In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO: 441, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:441. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:459, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:459.
[0167] In some embodiments, provided herein is an eZFP, such as eZFP_A43 as described herein. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:586, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:586. In some embodiments, the target site is double-stranded DNA. In some embodiments, the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: QSANRTK (SEQ ID NO:493); F2: QSGNEAR (SEQ ID NO:494); F3: RSDNESV (SEQ ID NO:495); F4: IRSTLRD (SEQ ID NO:496); F5: QNAHRKT (SEQ ID NO:497); and F6: HRSSLRR (SEQ ID NO:498). In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:442, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:442. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:460, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:460.
[01 8] In some embodiments, provided herein is an eZFP, such as eZFP_A44 as described herein. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:587, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:587. In some embodiments, the target site is double-stranded DNA. In some
embodiments, the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: QAGNRST (SEQ ID NO:499); F2: DRSALAR (SEQ ID NO:500); F3: RSDNLAR (SEQ ID NO:501); F4: WRGDRVK (SEQ ID NO:502); F5: YKHVLSD (SEQ ID NO:503); and F6: TSGSLTR (SEQ ID NO:504). In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO: 443, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:443. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:461, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:461.
[0169] In some embodiments, provided herein is an eZFP, such as eZFP_A45 as described herein. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:588, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:588. In some embodiments, the target site is double-stranded DNA. In some embodiments, the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: RSDNLSV (SEQ ID NO:505); F2: IRSTLRD (SEQ ID NO:506); F3: QNAHRKT (SEQ ID NO:507); F4: HRSSLRR (SEQ ID NO:508); F5: RSDNLAR (SEQ ID NO:509); and F6: QRSPLPA (SEQ ID NO:51Q). In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:444, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:444. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:462, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:462.
[0170] In some embodiments, provided herein is an eZFP, such as eZFP_A46 as described herein. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:589, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:589. In some embodiments, the target site is double-stranded DNA. In some embodiments, the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: DRSTRTK (SEQ ID NO:511); F2:
RSDYLAK (SEQ ID NO:512); F3: LRHHLTR (SEQ ID NO:513); F4: QSAHLKA (SEQ ID NO:514); F5: LPQTLQR (SEQ ID NO:515); and F6: QNATRTK (SEQ ID NO:516). In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO: 445, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:445. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:463, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:463.
[0171] In some embodiments, provided herein is an eZFP, such as eZFP_A47 as described herein. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:590, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:590. In some embodiments, the target site is double-stranded DNA. In some embodiments, the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: RSADLTR (SEQ ID NO:517); F2: RSDDLTR (SEQ ID NO:518); F3: QSSDLSR (SEQ ID NO:519); F4: WHSSLHQ (SEQ ID NO:520); F5: RSDSLSQ (SEQ ID NO:521); and F6: RKADRTR (SEQ ID NO:522). In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:446, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:446. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:464, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:464.
[0172] In some embodiments, provided herein is an eZFP, such as eZFP_A48 as described herein. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:591, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:591. In some embodiments, the target site is double-stranded DNA. In some embodiments, the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: RNDALTE (SEQ ID NO:523); F2: RKDNLKN (SEQ ID NO:524); F3: TSGELVR (SEQ ID NO:525); F4: HRTTLTN (SEQ ID NO:526); F5: TTGNLTV (SEQ ID NO:527); and F6: RTDTLRD (SEQ ID NO:528). In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO: 447, or a portion
thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:447. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:465, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:465.
[0173] In some embodiments, provided herein is an eZFP, such as eZFP_A49 as described herein. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:592, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:592. In some embodiments, the target site is double-stranded DNA. In some embodiments, the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: RKDNEKN (SEQ ID NO:529); F2: RADNETE (SEQ ID NO:530); F3: TSHSETE (SEQ ID NO:531); F4: SKKHLAE (SEQ ID NO:532); F5: TSGELVR (SEQ ID NO:533); and F6: TSGELVR (SEQ ID NO:534). In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:448, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:448. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:466, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:466.
[0174] In some embodiments, provided herein is an eZFP, such as eZFP_A50 as described herein. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:593, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:593. In some embodiments, the target site is double-stranded DNA. In some embodiments, the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: THEDEIR (SEQ ID NO:535); F2: DCRDEAR (SEQ ID NO:536); F3: RSDELVR (SEQ ID NO:537); F4: RNDALTE (SEQ ID NO:538); F5: SKKHLAE (SEQ ID NO:539); and F6: QSGHLTE (SEQ ID NO:540). In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO: 449, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:449. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID
NO:467, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:467.
[0175] In some embodiments, provided herein is an eZFP, such as eZFP_A51 as described herein. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:594, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:594. In some embodiments, the target site is double-stranded DNA. In some embodiments, the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: HTGHEEE (SEQ ID NO:541); F2: DPGHEVR (SEQ ID NO:542); F3: THEDEIR (SEQ ID NO:543); F4: DCRDEAR (SEQ ID NO:544); F5: RSDEEVR (SEQ ID NO:545); and F6: RNDALTE (SEQ ID NO:546). In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:450, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:450. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:468, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:468.
[0176] In some embodiments, provided herein is an eZFP, such as eZFP_A52 as described herein. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:595, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:595. In some embodiments, the target site is double-stranded DNA. In some embodiments, the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: RSDKEVR (SEQ ID NO:547); F2: RSDHETT (SEQ ID NO:548); F3: RNDAETE (SEQ ID NO:549); F4: TTGAETE (SEQ ID NO:550); F5: THLDLIR (SEQ ID NO:551); and F6: DPGHLVR (SEQ ID NO:552). In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO: 451, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:45E In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:469, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:469.
[0177] In some embodiments, provided herein is an eZFP, such as eZFP_A53 as described herein. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:596, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:596. In some embodiments, the target site is double-stranded DNA. In some embodiments, the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: TKNSLTE (SEQ ID NO:553); F2: QLAHLRA (SEQ ID NO:554); F3: TSGSLVR (SEQ ID NO:555); F4: RSDNLVR (SEQ ID NO:556); F5: QNSTLTE (SEQ ID NO:557); and F6: RADNLTE (SEQ ID NO:558). In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:452, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:452. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:470, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:470.
[0178] In some embodiments, provided herein is an eZFP, such as eZFP_A54 as described herein. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:597, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:597. In some embodiments, the target site is double-stranded DNA. In some embodiments, the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: RADNLTE (SEQ ID NO:559); F2: TKNSLTE (SEQ ID NO:560); F3: QLAHLRA (SEQ ID NO:561); F4: TSGSLVR (SEQ ID NO:562); F5: RSDNLVR (SEQ ID NO:563); and F6: QNSTLTE (SEQ ID NO:564). In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO: 453, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:453. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:471, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:471.
[0179] In some embodiments, provided herein is an eZFP, such eZFP_A55 as described herein. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:598, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the
foregoing. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:598. In some embodiments, the target site is double-stranded DNA. In some embodiments, the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: TSGHLVR (SEQ ID NO:565); F2: QLAHLRA (SEQ ID NO:566); F3: TSGELVR (SEQ ID NO:567); F4: QSGDLRR (SEQ ID NO:568); F5: QRAHLER (SEQ ID NO:569); and F6: RSDKLVR (SEQ ID NO:570). In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:454, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:454. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:472, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:472.
[0180] In some embodiments, provided herein is an eZFP, such as eZFP_A56 as described herein. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:599, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:599. In some embodiments, the target site is double-stranded DNA. In some embodiments, the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C- terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: REDNLHT (SEQ ID NO:571); F2: TSGHLVR (SEQ ID NO:572); F3: QLAHLRA (SEQ ID NO:573); F4: TSGELVR (SEQ ID NO:574); F5: QSGDLRR (SEQ ID NO:575); and F6: QRAHLER (SEQ ID NO:576). In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:455, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:455. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:473, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:473.
[0181] In some embodiments, provided herein is an eZFP, such eZFP_A57 as described herein. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NG:600, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the eZFP targets a target site comprising the nucleotide sequence set forth in SEQ ID NG:600. In some embodiments, the target site is double-stranded DNA. In some embodiments, the eZFP comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C-
terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: QRSDLTR (SEQ ID NO:577); F2: QGGTLRR (SEQ ID NO:578); F3: TSAHLAR (SEQ ID NO:579); F4: RREHLVR (SEQ ID NO:580); F5: QRHGLSS (SEQ ID NO:581); and F6: QRNALRG (SEQ ID NO:582). In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:456, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP comprises the amino acid sequence set forth in SEQ ID NO:456. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:474, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP is encoded by the nucleotide sequence set forth in SEQ ID NO:474.
II. DNA-Targeting Systems for Targeting a Frataxin Locus
[0182] Provided herein are DNA-targeting systems comprising a DNA-targeting domain that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus. Exemplary components and features of the DNA-targeting systems are provided herein. In some aspects, the DNA-targeting system comprises one or more of any of the components described herein, such as one or more DNA-targeting domains, one or more fusion proteins, such as one or more fusion proteins comprising one or more DNA- targeting domains and one or more effector domains, one or more gRNAs, or any component, portion or fragment thereof, or any combination thereof.
[0183] In some embodiments, a DNA-targeting system herein comprises an eZFP and/or eZFP fusion protein, such as any described above in Section I. In some embodiments, the DNA-targeting system comprises one or more of the eZFP fusion proteins. In some embodiments, the DNA-targeting system comprises at least two eZFP fusion proteins. In some embodiments, the two eZFP fusion proteins of the DNA-targeting system are any of the eZFP fusion proteins provided herein, such as any of the eZFP fusion proteins shown in Table 2A and Table 2B.
[0184] Provided herein are compositions such as DNA-targeting systems that bind to or target a frataxin (FXN) locus. In some aspects, the provided DNA-targeting systems include fusion proteins, such as eZFP fusion proteins or dCas fusion proteins. In some embodiments, the DNA-targeting system comprises one or more guide RNA (gRNA). In some aspects, provided are polynucleotides, vectors that encode any of the DNA-targeting systems, fusion proteins and/or components of kits. In some embodiments, provided are cells, kits, systems and pluralities and combinations thereof, that comprise any of the DNA-targeting systems, fusion proteins or gRNAs described herein.
[0185] Provided herein are DNA-targeting systems comprising a DNA-targeting domain that binds to a target site in a FXN locus, such as a regulatory DNA element of a frataxin (FXN) locus. In some of any of the embodiments provided herein, binding of the DNA-targeting domain to the target site does not introduce a genetic disruption or a DNA break at or near the target site. In some aspects, the provided
DNA-targeting systems comprise a fusion protein comprising a DNA-targeting domain and an effector domain, and binds to a target site in a regulatory DNA element of a FXN locus. In some embodiments, the DNA-targeting system comprises an eZFP, and/or an eZFP fusion protein, which binds to the target site. In some aspects, the DNA-targeting system comprises a guide RNA (gRNA), which targets the DNA-targeting domain to the target site (e.g., as in the case of CRISPR/Cas-based DNA-targeting systems). In some aspects, when administered to a subject or delivered or introduced into a cell that exhibits dysregulation or reduced activity, function or expression of FXN, the provided DNA-targeting systems can lead to an increase of or a restoration of, the activity, function or expression of FXN. In some aspects, also provided are methods and uses related to any of the provided compositions, for example, in modulating the expression of FXN, and/or in the treatment or therapy of diseases or disorders associated with dysregulation or reduced activity, function or expression of FXN, such as FA.
[0186] In some embodiments, the DNA-targeting systems are targeted to one or more target sites located within a FXN locus, such as a regulatory DNA element of a FXN locus, such as a promoter or an enhancer. In some embodiments, the DNA-targeting systems are targeted to at least 2, 3, 4, 5, 6, 7, 8, 9 or 10 target sites within a FXN locus. In some embodiments, the DNA-targeting systems are targeted to one or more target sites located within a promoter of a FXN locus, and one or more target sites located within an enhancer of a FXN locus.
[0187] In some embodiments, the DNA-targeting system comprises a DNA-targeting domain comprising a Clustered Regularly Interspaced Short Palindromic Repeats associated (Cas)-guide RNA (gRNA) combination comprising (a) a Cas protein or a variant thereof and (b) at least one gRNA; a zinc finger protein (ZFP), such as an engineered zinc finger protein (eZFP); a transcription activator-like effector (TALE); a meganuclease; a homing endonuclease; or an I-Scel enzyme or a variant thereof. In some aspects, the DNA-targeting domain comprises a catalytically inactive variant of any of the foregoing. In some embodiments, the DNA-targeting system comprises a DNA-targeting domain comprising a Cas-gRNA combination comprising (a) a Cas protein or a variant thereof, and (b) at least one gRNA. In some embodiments, the at least one gRNA comprises at least 2, 3, 4, 5, 6, 7, 8, 9 or 10 gRNAs. In some embodiments, the gRNAs are targeted to one or more target sites located within a regulatory DNA element of a FXN locus, such as a promoter or an enhancer. In some embodiments, the gRNAs are targeted to one or more target sites located within a promoter of a FXN locus, and one or more target sites located within an enhancer of a FXN locus.
[0188] In some aspects, the provided embodiments involve modulating transcription of an endogenous FXN locus in a cell. In some aspects, the provided embodiments involve increasing transcription of an endogenous FXN locus in a cell. In some embodiments, the cell, such as the cell to be treated with the provided embodiments, has a GA A trinucleotide repeat expansion in the FXN locus. In some embodiments, the cell, such as the cell to be treated with the provided embodiments, is from or in a subject with Friedreich’s ataxia. In some embodiments, the cell, such as the cell to be treated with the provided embodiments, exhibits reduced expression of FXN compared to a cell from a subject without
Friedreich’ s ataxia.
[0189] In some aspects, in a cell introduced with or contacted with any of the eZFPs, eZFP fusion proteins, DNA-targeting systems, gRNAs, combinations, fusion proteins, polynucleotides, plurality of polynucleotides, vectors, plurality of vectors or components or portions thereof provided herein, the expression of FXN is increased at least about 1.2-fold, 1.25-fold, 1.3-fold, 1.4-fold, 1.5-fold, 1.6-fold, 1.7-fold, 1.75-fold, 1.8-fold, 1.9-fold, 2-fold, 2.5-fold, 3-fold, 4-fold, or 5-fold, compared to a cell that has not been introduced or contacted. In some embodiments, the expression is increased by less than about 10-fold, 9-fold, 8-fold, 7-fold or 6-fold.
[0190] In some embodiments, the subject is a human. In some embodiments, the cell is a heart cell, a skeletal muscle cell, a nervous system cell, or an induced pluripotent stem cell. In some embodiments, the introducing, contacting or administering is carried out in vivo or ex vivo.
[0191] In some aspects, the DNA-targeting system comprises a DNA-targeting domain and one or more guide RNAs (gRNAs). In some aspects, the DNA-targeting system comprises a fusion protein and one or more gRNAs. In some aspects, the DNA-targeting system comprises a DNA-targeting domain and a gRNA. In some aspects, the DNA-targeting system comprises a fusion protein. In some aspects, the DNA-targeting system comprises a fusion protein and a gRNA. In some aspects, the DNA-targeting system comprises a DNA-targeting domain.
[0192] In some embodiments, binding of the DNA-targeting domain to the target site does not introduce a genetic disruption or a DNA break at or near the target site.
[0193] In some embodiments, provided are DNA-targeting systems capable of specifically targeting a target site in a FXN gene or DNA regulatory element thereof, and increasing transcription of the FXN gene. In some embodiments, the DNA-targeting systems include a DNA-targeting domain that binds to a target site in the FXN gene or regulatory DNA element thereof. In provided embodiments, the DNA- targeting systems additionally include at least one effector domain that is able to epigenetically modify one or more DNA bases of the FXN gene or regulatory element thereof, in which the epigenetic modification results in an increase in transcription of the FXN gene (e.g. activates transcription or increases transcription of FXN compared to the absence of the DNA-targeting system). Hence, the terms DNA-targeting system and epigenetic-modifying DNA targeting system may be used herein interchangeably. In some embodiments, the DNA-targeting system includes a fusion protein comprising (a) a DNA-targeting domain capable of being targeted to the target site; and (b) at least one effector domain capable of increasing transcription of the FXN gene. For instance, the at least one effector domain is a transcription activation domain.
[0194] In some embodiments, the DNA-targeting domain comprises or is derived from a CRISPR associated (Cas) protein, zinc finger protein (ZFP), transcription activator-like effectors (TALE), meganuclease, homing endonuclease, I-Scel enzyme, or variants thereof. In some embodiments, the DNA-targeting domain comprises a catalytically inactive (e.g. nuclease-inactive or nuclease-inactivated) variant of any of the foregoing. In some embodiments, the DNA-targeting domain comprises a
deactivated Cas9 (dCas9) protein or variant thereof that is a catalytically inactivated so that it is inactive for nuclease activity and is not able to cleave the DNA.
[0195] In some embodiments, the DNA-targeting domain comprises or is derived from a Cas protein or variant thereof, such as a nuclease-inactive Cas or dCas (e.g. dCas9, and the DNA-targeting system comprises one or more guide RNAs (gRNAs). In some embodiments, the gRNA comprises a spacer sequence that is capable of targeting and/or hybridizing to the target site. In some embodiments, the gRNA is capable of complexing with the Cas protein or variant thereof. In some aspects, the gRNA directs or recruits the Cas protein or variant thereof to the target site.
[0196] In some embodiments, the effector domain comprises a transcription activation domain, and/or is capable of increasing transcription of the gene. In some embodiments, the effector domain induces, catalyzes or leads to transcription activation, transcription co-activation, or transcription elongation. In some aspects, the effector domain is selected from VP64, p65, Rta, p300, CBP, VPR, VPH, Rta, p300, HSF1, a TET protein (e.g. TET1), SunTag, a partially or fully functional fragment or domain thereof, or a combination of any of the foregoing. In some embodiments, the effector domain may include a domain, portion, or variant of a protein selected from: DPOLA, ENL, FOXO3, HSH2D, NCOA2, NCOA3, PSA1, PYGO1, RBM39, HERC2, and NOTCH2. In some embodiments, the effector domain is VP64. In some embodiments, the fusion protein of the DNA-targeting system comprises dCas9-VP64. In some embodiments, the fusion protein of the DNA-targeting system is an eZFP fusion protein, such as any eZFP fusion protein disclosed herein, such as in Section II.C.
[0197] In some embodiments, the DNA-targeting system comprises a DNA-targeting domain. In some embodiments the DNA-targeting domain comprises a DNA-binding protein or DNA-binding nucleic acid. In some embodiments, the DNA-targeting domain specifically binds to or hybridizes to a particular site or position in the genome, e.g., a target, target site, or target position. In some aspects, the DNA-targeting domain is coupled to, fused to or complexed with an effector domain, such as any effector domain described herein, for example, in Section II.B.
[0198] In some embodiments, the DNA-targeting system comprises various components, such as an RNA-guided nuclease, variant thereof (such as dCas), or fusion protein comprising the RNA-guided nuclease or variant thereof, or a fusion protein comprising a DNA-targeting domain and an effector domain. In some embodiments, the DNA-targeting system comprises a DNA-targeting molecule that comprises a DNA-binding protein such as one or more zinc finger protein (ZFP) or transcription activator-like effectors (TALEs), fused to an effector domain. In some embodiments, the DNA-binding protein of the DNA-targeting molecule comprises an eZFP (e.g., the DNA-targeting molecule is an eZFP fusion protein).
[0199] In some embodiments, the DNA-targeting system specifically targets at least one target site in a regulatory DNA element of a frataxin (FXN) locus. In some embodiments, the DNA-targeting system comprises a ZFP, a TAEE, or a CRISPR/Cas9 combination, that specifically binds to, recognizes, or hybridizes to the target site(s). In some embodiments, the CRISPR/Cas9 system includes an
engineered crRNA/tracr RNA (i.e. “single guide RNA”). In some embodiments, the DNA-targeting system comprises nucleases or variants thereof based on the Argonaute system (e.g., from T. thermophilus, known as ‘TtAgo’ (Swarts et al., (2014) Nature 507(7491): 258-261).
[0200] In some embodiments, the DNA-targeting domain comprises a Clustered Regularly Interspaced Short Palindromic Repeats associated (Cas)-guide RNA (gRNA) combination that includes (a) a Cas protein or a variant thereof and (b) at least one gRNA; a zinc finger protein (ZFP) (such as an eZFP); a transcription activator-like effector (TALE); a meganuclease; a homing endonuclease; or a I- Scel enzyme or a variant thereof. In some embodiments, the DNA-targeting domain comprises a catalytically inactive variant of any of the foregoing. In some embodiments, the DNA-targeting domain comprises a Cas-gRNA combination that includes (a) a Cas protein or a variant thereof and (b) at least one gRNA. In some embodiments, the variant Cas protein lacks nuclease activity or is a deactivated Cas (dCas) protein.
[0201] Also provided herein are DNA-targeting systems comprising a DNA-targeting domain, that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus and comprises a Cas-guide RNA (gRNA) combination comprising: (a) a variant Cas protein that lacks nuclease activity or that is a deactivated Cas (dCas) protein; and (b) at least one gRNA comprising at least one gRNA spacer sequence that is capable of hybridizing to the target site or is complementary to the target site.
[0202] Also provided herein are DNA-targeting systems comprising a DNA-targeting domain that comprises a Cas-guide RNA (gRNA) combination comprising: (a) a Staphylococcus aureus deactivated Cas9 protein (dSaCas9) protein set forth in SEQ ID NO:72 fused to at least one effector domain that induces transcription activation; and (b) at least one gRNA comprising the gRNA spacer sequence set forth in SEQ ID NO:42.
[0203] Also provided herein are DNA-targeting systems comprising a DNA-targeting domain that comprises a Cas-guide RNA (gRNA) combination comprising: (a) a Staphylococcus aureus deactivated Cas9 protein (dSaCas9) protein set forth in SEQ ID NO:72 fused to at least one effector domain that induces transcription activation; and (b) at least one gRNA comprising the gRNA spacer sequence set forth in SEQ ID NO:22.
[0204] Also provided herein are DNA-targeting systems comprising a DNA-targeting domain that comprises a Cas-guide RNA (gRNA) combination comprising: (a) a Staphylococcus aureus deactivated Cas9 protein (dSaCas9) protein set forth in SEQ ID NO:72 fused to at least one effector domain that induces transcription activation; and (b) at least one gRNA comprising the gRNA spacer sequence set forth in SEQ ID NO:28.
[0205] In some embodiments, the provided DNA-targeting systems or fusion proteins comprise a DNA-targeting domain. In some aspects, the DNA-targeting domain provides sequence specificity and targets the DNA targeting system or fusion protein to a particular location of the genome, such as a target site specified by a component of the DNA-targeting domain. In some embodiments, an exemplary DNA- targeting domain comprises a Clustered Regularly Interspaced Short Palindromic Repeats associated
(Cas)-guide RNA (gRNA) combination that includes (a) a Cas protein or a variant thereof and (b) at least one gRNA; a zinc finger protein (ZFP) such as an eZFP; a transcription activator-like effector (TALE); a meganuclease; a homing endonuclease; or a I-Scel enzymes or a variant of any of the foregoing. In some embodiments, the DNA-targeting domain comprises a catalytically inactive variant of any of the foregoing. In some embodiments, the DNA-targeting domain comprises a Cas-gRNA combination that includes (a) a Cas protein or a variant thereof and (b) at least one gRNA. In some embodiments, the variant Cas protein lacks nuclease activity or is a deactivated Cas (dCas) protein. In some aspects, for a DNA-targeting domain that comprises a Cas-gRNA combination, the gRNA component (such as any described herein) provides the sequence specificity to target the DNA-targeting system, DNA-targeting domain or fusion protein to a target site specified by the gRNA. In some embodiments, the DNA- targeting domain comprises an eZFP, such as any described herein.
[0206] In some of any of the provided embodiments, the DNA-targeting domain comprises a zinc finger protein (ZFP); a transcription activator-like effector (TALE); a meganuclease; a homing endonuclease; or an I-Scel enzyme or a variant thereof. In some embodiments, the DNA-targeting domain comprises a catalytically inactive variant of any of the foregoing. In some aspects, types of DNA- targeting domains include domains from proteins that can recognize nucleic acid sequences (e.g., target site) in a sequence-specific manner.
[0207] In some embodiments, a “zinc finger DNA binding protein” (or binding domain) is a protein, or a domain within a larger protein, that binds DNA in a sequence-specific manner through one or more zinc fingers, which are regions of amino acid sequence within the binding domain whose structure is stabilized through coordination of a zinc ion. The term zinc finger DNA binding protein is often abbreviated as zinc finger protein or ZFP. Among the ZFPs are artificial, or engineered, ZFPs, comprising ZFP domains targeting specific DNA sequences, typically 9-18 nucleotides long, generated by assembly of individual fingers. ZFPs include those in which a single finger domain is approximately 30 amino acids in length and contains an alpha helix containing two invariant histidine residues coordinated through zinc with two cysteines of a single beta turn, and having two, three, four, five, or six fingers. Generally, sequence-specificity of a ZFP may be altered by making amino acid substitutions at the four helix positions (-1, 2, 3, and 6) on a zinc finger recognition helix. Thus, for example, the ZFP or ZFP-containing molecule is non-naturally occurring, e.g., is engineered to bind to a target site of choice.
[0208] In some cases, the DNA-targeting system is or comprises a zinc-finger DNA binding domain fused to an effector domain. In some embodiments, zinc fingers are custom-designed (i.e. designed by the user), or obtained from a commercial source. Various methods for designing zinc finger proteins are available. For example, methods for designing zinc finger proteins to bind to a target DNA sequence of interest are described, for example in Liu, Q. et al., PNAS, 94(l l):5525-30 (1997); Wright, D.A. et al., Nat. Protoc., l(3):1637-52 (2006); Gersbach, C.A. et al., Acc. Chem. Res., 47(8):2309-18 (2014); Bhakta M.S. et al., Methods Mol. Biol., 649:3-30 (2010); and Gaj et al., Trends Biotechnol, 31(7):397-405 (2013). In addition, various web-based tools for designing zinc finger proteins to bind to a DNA target
sequence of interest are publicly available. See, for example, the Zinc Finger Tools design web site from Scripps available on the world wide web at scripps.edu/barbas/zfdesign/zfdesignhome.php. Various commercial services for designing zinc finger proteins to bind to a DNA target sequence of interest are also available. See, for example, the commercially available services or kits offered by Creative Biolabs (world wide web at creative-biolabs.com/Design-and-Synthesis-of-Artificial-Zinc-Finger-Proteins.html), the Zinc Finger Consortium Modular Assembly Kit available from Addgene (world wide web at addgene.org/kits/zfc-modular-assembly/), or the CompoZr Custom ZFN Service from Sigma Aldrich (world wide web at sigmaaldrich.com/life-science/zinc-finger-nuclease-technology/custom-zfn.html). For example, platforms for zinc-finger construction are available that provide specifically targeted zinc fingers for thousands of targets. See, e.g., Gaj et al., Trends in Biotechnology, 2013, 31(7), 397-405. Some gene-specific engineered zinc fingers are available commercially. In some cases, commercially available zinc fingers are used or are custom designed.
[0209] In some aspects, the DNA-targeting domain is a domain from Transcription activator-like effectors (TALEs). TALEs are proteins found in Xanthomonas bacteria. TALEs comprise a plurality of repeated amino acid sequences, each repeat having binding specificity for one base in a target sequence. Each repeat comprises a pair of variable residues in position 12 and 13 (repeat variable diresidue; RVD) that determine the nucleotide specificity of the repeat. In some embodiments, RVDs associated with recognition of the different nucleotides are HD for recognizing C, NG for recognizing T, NI for recognizing A, NN for recognizing G or A, NS for recognizing A, C, G or T, HG for recognizing T, IG for recognizing T, NK for recognizing G, HA for recognizing C, ND for recognizing C, HI for recognizing C, HN for recognizing G, NA for recognizing G, SN for recognizing G or A and YG for recognizing T, TL for recognizing A, VT for recognizing A or G and SW for recognizing A. In some embodiments, RVDs can be mutated towards other amino acid residues in order to modulate their specificity towards nucleotides A, T, C and G and in particular to enhance this specificity. Binding domains with similar modular base-per-base nucleic acid binding properties can also be derived from different bacterial species. These alternative modular proteins may exhibit more sequence variability than TALE repeats.
[0210| In some embodiments, a “TALE DNA binding domain” or “TALE” is a polypeptide comprising one or more TALE repeat domains/units. The repeat domains, each comprising a repeat variable diresidue (RVD), are involved in binding of the TALE to its cognate target DNA sequence. A single “repeat unit” (also referred to as a “repeat”) is typically 33-35 amino acids in length and exhibits at least some sequence homology with other TALE repeat sequences within a TALE protein. TALE proteins may be designed to bind to a target site using canonical or non-canonical RVDs within the repeat units. See, e.g., U.S. Pat. Nos. 8,586,526 and 9,458,205.
[0211] In some embodiments, a TALE is a fusion protein comprising a nucleic acid binding domain derived from a TALE and an effector domain. In some embodiments, one or more sites in the FXN locus can be targeted by engineered TALEs.
[0212] Zinc finger and TALE DNA-binding domains can be engineered to bind to a predetermined nucleotide sequence, for example via engineering (altering one or more amino acids) of the recognition helix region of a zinc finger protein, by engineering of the amino acids in a TALE repeat involved in DNA binding (the repeat variable diresidue or RVD region), or by systematic ordering of modular DNA- binding domains, such as TALE repeats or ZFP domains. Therefore, engineered zinc finger proteins or TALE proteins are proteins that are non-naturally occurring. Non-limiting examples of methods for engineering zinc finger proteins and TALEs are design and selection. A designed protein is a protein not occurring in nature whose design/composition results principally from rational criteria. Rational criteria for design include application of substitution rules and computerized algorithms for processing information in a database storing information of existing ZFP or TALE designs (canonical and non- canonical RVDs) and binding data. See, for example, U.S. Pat. Nos. 9,458,205; 8,586,526; 6,140,081; 6,453,242; and 6,534,261; see also WO 98/53058; WO 98/53059; WO 98/53060; WO 02/016536 and WO 03/016496.
[0213] In some embodiments, the DNA-targeting system comprises at least one effector domain, such as any epigenetic effector domain provided herein, for example as described in Section II.B. In some embodiments, the DNA-targeting domain or a component thereof is fused to the at least one effector domain. In some embodiments, provided herein is a DNA-targeting system comprising a fusion protein comprising: (a) a DNA-targeting domain targeting, or capable of being targeted to, a target site at a FXN locus or a regulatory element thereof, such as any described herein, and (b) at least one effector domain. In some aspects, the effector domain leads to an increase in transcription of FXN, or is capable of increasing transcription of FXN. In some aspects, the effector domain comprises a transcription activation domain. In some aspects, the effector domain comprises a multipartite activator.
[0214] In some embodiments, the DNA-targeting domain comprises a Cas-gRNA combination comprising (a) a Cas protein or a variant thereof and (b) at least one gRNA, and the component thereof fused to the at least one effector domain is the Cas protein or a variant thereof.
[0215] In some aspects, the effector domain activates, induces, catalyzes, or leads to demethylation and/or increased transcription of FXN when ectopically recruited to FXN or a DNA regulatory element thereof. Exemplary fusion of DNA-targeting domain and at least one effector domain include fusing dCas9 with transcriptional activators such as VP64 (a polypeptide composed of four tandem copies of VP 16, a 16 amino acid transactivation domain of the Herpes simplex virus) can result in robust induction of gene expression.
[0216] In some aspects, the effector domain activates, induces, catalyzes, or leads to demethylation and/or increased transcription of FXN when ectopically recruited to FXN or a DNA regulatory element thereof. In some embodiments, the effector domain induces, catalyzes or leads to transcription activation, transcription co-activation, transcription elongation, transcription de -repression, histone modification, nucleosome remodeling, chromatin remodeling, reversal of heterochromatin formation, DNA demethylation, or DNA base oxidation. In some embodiments, the effector domain induces, catalyzes or
leads to transcription de-repression, DNA demethylation or DNA base oxidation. In some embodiments, the effector domain induces transcription de -repression. In some embodiments, the effector domain induces transcription activation. In some embodiments, the effector domain has one of the aforementioned activities itself (i.e. acts directly). In some embodiments, the effector domain recruits and/or interacts with a polypeptide domain that has one of the aforementioned activities (i.e. acts indirectly).
[0217] In some embodiments, the effector domain induces, catalyzes or leads to transcription activation, transcription co-activation, transcription elongation, transcription de -repression, transcription factor release, polymerization, histone modification, histone acetylation, histone deacetylation, nucleosome remodeling, chromatin remodeling, reversal of heterochromatin formation, nuclease, signal transduction, proteolysis, ubiquitination, deubiquitination, phosphorylation, dephosphorylation, splicing, nucleic acid association, DNA methylation, DNA demethylation, histone methylation, histone demethylation, or DNA base oxidation. In some embodiments, the effector domain induces, catalyzes or leads to transcription activation, transcription co-activation, or transcription elongation. In some embodiments, the effector domain induces transcription activation. In some embodiments, the effector domain activates transcription from one or more regulatory elements (e.g., promoters and/or enhancers) from the target locus, e.g., FXN. In some embodiments, the effector domain induces transcription activation. In some embodiments, the effector domain has one of the aforementioned activities itself (i.e. acts or catalyzes directly). In some embodiments, the effector domain recruits and/or interacts with another cellular component (e.g., transcription factor) that has one of the aforementioned activities (i.e. acts or catalyzes indirectly).
[0218] In some aspects, gene expression of endogenous mammalian genes, such as human genes, can be achieved by targeting a fusion protein comprising a DNA-targeting domain, such as a dCas9, and an effector domain, such as a transcription activation domain, to mammalian genes or regulatory DNA elements thereof (e.g. a promoter or enhancer), e.g. via one or more gRNAs. Any of a variety of effector domains for transcriptional activation (e.g. transcription activation domains) are known and can be used in accord with the provided embodiments. Transcription activation domains, as well as activation of target genes by Cas fusion proteins (with a variety of Cas molecules) and the transcription activation domains, are described, for example, in WO 2014/197748, WO 2016/130600, WO 2017/180915, WO 2021/226555, WO 2021/226077, WO 2013/176772, WO 2014/152432, WO 2014/093661, Adli, M. Nat. Commun. 9, 1911 (2018), Perez-Pinera et al. Nat. Methods 10, 973-976 (2013), Mali et al. Nat. Biotechnol. 31, 833-838 (2013), and Maeder et al. Nat. Methods 10, 977-979 (2013).
[0219] In some embodiments, the effector domain comprises a transcriptional activator domain described in WO 2021/226077.
[0220] In some aspects, activation or increase in gene expression of FXN is achieved by targeting a fusion protein comprising a DNA-targeting domain, such as a dCas9, and an effector domain, such as a transcription activation domain, to a FXN locus or regulatory DNA elements thereof (e.g. a promoter or
enhancer) via one or more gRNAs. In some aspects, the one or more target sites of the one or more gRNA is at a FXN locus or regulatory DNA elements thereof (e.g., a promoter or enhancer), for example, as described herein, for example, in Section II. A. Any of a variety of effector domains for transcriptional activation (e.g. transcription activation domains) are known and can be used in accord with the provided embodiments as described herein, for example, in Section II.B.
[0221] The effector domain may have transcription activation activity, i.e., a transactivation domain. For example, gene expression of endogenous mammalian genes, such as human genes, may be achieved by targeting a fusion protein of iCas9 and a transactivation domain to mammalian promoters via combinations of gRNAs. The transactivation domain may include a VP16 protein, multiple VP16 proteins, such as a VP48 domain or VP64 domain, or p65 domain of NF kappa B transcription activator activity. For example, the fusion protein may be iCas9-VP64. In some embodiments, the effector domain may comprise a VP64 domain. For example, dCas9-VP64 can be targeted to a target site by one or more gRNAs to activate a gene. VP64 is a polypeptide composed of four tandem copies of VP16, a 16 amino acid transactivation domain of the Herpes simplex virus. VP64 domains, including in dCas fusion proteins, have been described, for example, in WO 2014/197748, WO 2013/176772, WO 2014/152432, and WO 2014/093661.
[0222] In some embodiments, the effector domain comprises at least one VP16 domain, or a VP16 tetramer (“VP64”) or a variant thereof. In some embodiments, the effector domain comprises at least one VP16 domain, or a VP16 tetramer (“VP64”) or a variant thereof. In some embodiments, the effector domain comprises the sequence set forth in SEQ ID NO:81 or 83, a domain thereof, a portion thereof, or a variant thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing. An exemplary VP64 domain is set forth in SEQ ID NO:81. An exemplary nucleotide sequence encoding the exemplary VP64 domain set forth in SEQ ID NO: 81 is set forth in SEQ ID NO: 80. In some embodiments, the effector domain comprises the sequence set forth in SEQ ID NO: 81, a domain thereof, a portion thereof, or a variant thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing. An exemplary VP64 domain is set forth in SEQ ID NO:83. An exemplary nucleotide sequence encoding the exemplary VP64 domain set forth in SEQ ID NO: 83 is set forth in SEQ ID NO: 82. In some embodiments, the effector domain comprises the sequence set forth in SEQ ID NO:83, a domain thereof, a portion thereof, or a variant thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
A. Target Sites for the Frataxin Locus
[0223] In some embodiments, the provided compositions, including the eZFPs, fusion proteins (such as eZFP fusion proteins and dCas fusion proteins), and DNA-targeting systems, can target one or more target sites in a FXN locus. In some embodiments, the target site comprises a specific sequence of
nucleotides, such as DNA nucleotides. In some embodiments, the target site is a DNA regulatory element of the FXN locus, such as a promoter or enhancer. In some aspects, provided are compositions, methods and uses, such as eZFPs, eZFP fusion proteins, DNA-targeting systems, DNA-targeting domains, components of the DNA-targeting domains, such as at least one gRNA, fusion proteins, and pluralities and combinations thereof, polynucleotides, vectors, cells, and pluralities and combinations thereof, that encode or comprise the DNA-targeting systems, fusion proteins, gRNAs, or pluralities or combinations thereof, that can target one or more particular genomic locations related to the FXN locus, such as a regulatory DNA element of the FXN locus.
[0224] In some embodiments, the target site is in a cell, such as any suitable cell. In some embodiments, the cell is in or from any suitable organism, such as a human, mouse, dog, horse, rabbit, cattle, pig, hamster, gerbil, mouse, ferret, rat, cat, non-human primate, monkey, etc. In some embodiments, the cell is in or from a human. In some embodiments, the cell is any suitable cell, such as an immune cell (e.g. a T cell, B cell, or antigen-presenting cell), a liver cell (e.g. a hepatocyte), a cell of a nervous system (e.g. a neuron or glial cell), a heart cell (e.g. a cardiomyocyte) or a stem cell (e.g. an embryonic stem cell or induced pluripotent stem cell).
[0225] In some embodiments, the target site is located in a regulatory DNA element of a frataxin (FXN) locus. In some embodiments, the target site is located within a promoter, upstream regulatory element (e.g., enhancer), exon, intron, 5’ untranslated region (UTR), 3’ UTR, or downstream regulatory element.
[0226] In some embodiments, the target site is located within a FXN locus. In some embodiments the target site is located within a regulatory DNA element (e.g. a cis-, trans-, distal, proximal, upstream, or downstream regulatory DNA element) of a FXN locus. In some embodiments, the target site is located within a promoter, enhancer, exon, intron, untranslated region (UTR), 5’ UTR or 3’ UTR. In some embodiments the target site is located within a sequence and/or sequences of unknown or known function that are suspected of being able to control expression of FXN.
[0227] In some embodiments one or more target sites, such as one or more target sites located within a regulatory DNA element (e.g. a cis-, trans-, distal, proximal, upstream, or downstream regulatory DNA element) of a FXN locus. In some embodiments, the target site is located within a promoter, enhancer, exon, intron, untranslated region (UTR), 5’ UTR or 3’ UTR are targeted.
[0228] In some aspects, an exemplary frataxin (FXN) transcript is set forth in RefSeq NM_000144) (transcript variant 1); Gencode Transcript: ENST00000484259.3; Gencode Gene: ENSG00000165060.15. Genomic coordinates for an exemplary transcript (including UTRs) for FXN include hg38 chr9:69, 035, 752-69, 079, 076 (Size: 43,325 Total Exon Count: 5 Strand: +). Genomic coordinates for the coding region for this transcript variant include hg38 chr9:69, 035, 783-69, 072, 762 (Size: 36,980 Coding Exon Count: 5).
[0229] In some aspects, an exemplary frataxin (FXN) transcript is set forth in RefSeq NM_181425) (transcript variant 2); Gencode Transcript: ENST00000396366.6; Gencode Gene:
ENSG00000165060.15. Genomic coordinates for an exemplary transcript (including UTRs) for FXN include hg38 chr9:69, 035, 762-69, 073, 022 (Size: 37,261 Total Exon Count: 5 Strand: +). Genomic coordinates for the coding region for this transcript variant include hg38 chr9:69, 035, 783-69, 072, 712 (Size: 36,930 Coding Exon Count: 5).
[0230] In some embodiments, the regulatory DNA element is located in a genomic region comprising the FXN locus.
[0231] In some embodiments, the target site is located within the genomic coordinates hg38 chr9:68,940, 179-69,205,519. In some embodiments, the target site is selected from the sequence set forth in any one of SEQ ID NOS:1-21, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing.
[0232] In some embodiments, the regulatory DNA element is an enhancer.
[0233] In some embodiments, the target site is located within the genomic coordinates human genome assembly GRCh38 (hg38) chr9:69, 027, 282-69, 028, 497. In some embodiments, the target site is located within the genomic coordinates hg38 chr9:69, 027, 615-69, 028, 101. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO: 21, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises a sequence set forth in any one of SEQ ID NOS:229-243, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises a sequence set forth in any one of SEQ ID NOS:256-265, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing.
[0234] In some embodiments, the target site is located within the genomic coordinates hg38 chr9:69, 044, 201-69, 045, 347. In some embodiments, the target site is located within the genomic coordinates hg38 chr9:69, 030, 752-69, 031,507. In some embodiments, the target site is located within the genomic coordinates hg38 chr9:68, 999, 262-69, 000, 023. In some embodiments, the target site is located within the genomic coordinates hg38 chr9:69, 085, 468-69, 086, 426. In some embodiments, the target site is located within the genomic coordinates hg38 chr9:69, 096, 701-69, 097, 567. In some embodiments, the target site is located within the genomic coordinates hg38 chr9:69, 120, 690-69, 123 ,549. In some embodiments, the target site is located within the genomic coordinates hg38 chr9:69, 130, 392-69, 132, 484.
[0235] In some embodiments, the DNA-targeting domain comprises a Cas-gRNA combination comprising (a) a Cas protein or a variant thereof and (b) at least one gRNA; and the gRNA comprises at least one gRNA spacer sequence comprising the sequence set forth in SEQ ID NO:42, or a contiguous portion thereof of at least 14 nt. In some embodiments, the gRNA further comprises the sequence set forth in SEQ ID NO:44. In some embodiments, the gRNA comprises the sequence set forth in SEQ ID NO:67.
[0236] In some embodiments, the regulatory DNA element is a promoter.
[0237] In some embodiments, the target site is located within the genomic coordinates hg38 chr9:69, 034, 622-69, 036, 670. In some embodiments, the target site is located within the genomic
coordinates hg38 chr9:69, 035, 300-69, 035, 900. In some embodiments, the target site is located within the genomic coordinates hg38 chr9:69, 034, 900-69, 035, 900. In some embodiments, the target site comprises a sequence selected from any one of SEQ ID NOS: 1-10, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises a sequence selected from any one of SEQ ID NOS: 11-20, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises a sequence selected from any one of SEQ ID NOS:244-255, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing.
[0238] In some embodiments, the target site is at, near, or within a FXN locus. In some embodiments, the target site is a sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to all or a portion of the target site sequence described herein. In some aspects, the target site is a sequence having at least 80% sequence identity to all or a portion of the target site sequence described herein. In some aspects, the target site is a sequence having at least 85% sequence identity to all or a portion of the target site sequence described herein. In some aspects, the target site is a sequence having at least 90% sequence identity to all or a portion of the target site sequence described herein. In some aspects, the target site is a sequence having at least 91% sequence identity to all or a portion of the target site sequence described herein. In some aspects, the target site is a sequence having at least 92% sequence identity to all or a portion of the target site sequence described herein. In some aspects, the target site is a sequence having at least 93% sequence identity to all or a portion of the target site sequence described herein. In some aspects, the target site is a sequence having at least 94% sequence identity to all or a portion of the target site sequence described herein. In some aspects, the target site is a sequence having at least 95% sequence identity to all or a portion of the target site sequence described herein. In some aspects, the target site is a sequence having at least 96% sequence identity to all or a portion of the target site sequence described herein. In some aspects, the target site is a sequence having at least 97% sequence identity to all or a portion of the target site sequence described herein. In some aspects, the target site is a sequence having at least 98% sequence identity to all or a portion of the target site sequence described herein. In some aspects, the target site is a sequence having at least 99% sequence identity to all or a portion of the target site sequence described herein. In some aspects, the target site is a sequence having at least 99.5% sequence identity to all or a portion of the target site sequence described herein. In some aspects, the target site is a sequence having at least 99.9% sequence identity to all or a portion of the target site sequence described herein. In some aspects, the target site is a sequence having 100% sequence identity to all or a portion of the target site sequence described herein.
[0239] In some embodiments, the target site is selected from the sequence set forth in any one of SEQ ID NOS:1-21, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing.
[0240] In some embodiments, the target site comprises SEQ ID NO:1, a contiguous portion thereof
of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:2, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:3, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:4, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:5, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:6, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:7, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:8, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:9, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO: 10, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:11, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO: 12, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO: 13, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO: 14, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO: 15, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO: 16, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO: 17, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO: 18, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO: 19, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:20, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:21, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof.
[0241] In some embodiments, the target site comprises SEQ ID NO:229, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:230, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:231, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:232, a contiguous portion thereof of at least 14 nt, or a complementary sequence
of thereof. In some embodiments, the target site comprises SEQ ID NO:233, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:234, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:235, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:236, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:237, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:238, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:239, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:240, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:241, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:242, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:243, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:244, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:245, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:246, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:247, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:248, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:249, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:250, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:251, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:252, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:253, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:254, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:255, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:256, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:257, a contiguous portion
thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:258, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:259, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:260, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:261, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:262, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:263, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:264, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof. In some embodiments, the target site comprises SEQ ID NO:265, a contiguous portion thereof of at least 14 nt, or a complementary sequence of thereof.
[0242] In some embodiments, the target site comprises SEQ ID NO:1. In some embodiments, the target site comprises SEQ ID NO:2. In some embodiments, the target site comprises SEQ ID NO:3. In some embodiments, the target site comprises SEQ ID NO:4. In some embodiments, the target site comprises SEQ ID NO:5. In some embodiments, the target site comprises SEQ ID NO:6. In some embodiments, the target site comprises SEQ ID NO:7. In some embodiments, the target site comprises SEQ ID NO:8. In some embodiments, the target site comprises SEQ ID NO:9. In some embodiments, the target site comprises SEQ ID NO: 10. In some embodiments, the target site comprises SEQ ID NO: 11. In some embodiments, the target site comprises SEQ ID NO: 12. In some embodiments, the target site comprises SEQ ID NO: 13. In some embodiments, the target site comprises SEQ ID NO: 14. In some embodiments, the target site comprises SEQ ID NO: 15. In some embodiments, the target site comprises SEQ ID NO: 16. In some embodiments, the target site comprises SEQ ID NO: 17. In some embodiments, the target site comprises SEQ ID NO: 18. In some embodiments, the target site comprises SEQ ID NO: 19. In some embodiments, the target site comprises SEQ ID NO:20. In some embodiments, the target site comprises SEQ ID NO:21.
[02431 In some embodiments, the target site comprises SEQ ID NO: 229. In some embodiments, the target site comprises SEQ ID NO:230. In some embodiments, the target site comprises SEQ ID NO:231. In some embodiments, the target site comprises SEQ ID NO:232. In some embodiments, the target site comprises SEQ ID NO:233. In some embodiments, the target site comprises SEQ ID NO:234. In some embodiments, the target site comprises SEQ ID NO: 235. In some embodiments, the target site comprises SEQ ID NO:236. In some embodiments, the target site comprises SEQ ID NO:237. In some embodiments, the target site comprises SEQ ID NO: 238. In some embodiments, the target site comprises SEQ ID NO:239. In some embodiments, the target site comprises SEQ ID NO:240. In some embodiments, the target site comprises SEQ ID NO: 241. In some embodiments, the target site comprises SEQ ID NO:242. In some embodiments, the target site comprises SEQ ID NO:243. In some
embodiments, the target site comprises SEQ ID NO:244. In some embodiments, the target site comprises SEQ ID NO:245. In some embodiments, the target site comprises SEQ ID NO:246. In some embodiments, the target site comprises SEQ ID NO: 247. In some embodiments, the target site comprises SEQ ID NO:248. In some embodiments, the target site comprises SEQ ID NO:249. In some embodiments, the target site comprises SEQ ID NO:250. In some embodiments, the target site comprises SEQ ID NO:251. In some embodiments, the target site comprises SEQ ID NO:252. In some embodiments, the target site comprises SEQ ID NO: 253. In some embodiments, the target site comprises SEQ ID NO:254. In some embodiments, the target site comprises SEQ ID NO:255. In some embodiments, the target site comprises SEQ ID NO:256. In some embodiments, the target site comprises SEQ ID NO:257. In some embodiments, the target site comprises SEQ ID NO:258. In some embodiments, the target site comprises SEQ ID NO: 259. In some embodiments, the target site comprises SEQ ID NO:260. In some embodiments, the target site comprises SEQ ID NO:261. In some embodiments, the target site comprises SEQ ID NO:262. In some embodiments, the target site comprises SEQ ID NO:263. In some embodiments, the target site comprises SEQ ID NO:264. In some embodiments, the target site comprises SEQ ID NO: 265.
[0244] In some embodiments, the target site comprises a complementary sequence of SEQ ID NO:1. In some embodiments, the target site comprises a complementary sequence of SEQ ID NO:2. In some embodiments, the target site comprises a complementary sequence of SEQ ID NO:3. In some embodiments, the target site comprises a complementary sequence of SEQ ID NO:4. In some embodiments, the target site comprises a complementary sequence of SEQ ID NO:5. In some embodiments, the target site comprises a complementary sequence of SEQ ID NO:6. In some embodiments, the target site comprises a complementary sequence of SEQ ID NO:7. In some embodiments, the target site comprises a complementary sequence of SEQ ID NO:8. In some embodiments, the target site comprises a complementary sequence of SEQ ID NO:9. In some embodiments, the target site comprises a complementary sequence of SEQ ID NO: 10. In some embodiments, the target site comprises a complementary sequence of SEQ ID NO: 11. In some embodiments, the target site comprises a complementary sequence of SEQ ID NO: 12. In some embodiments, the target site comprises a complementary sequence of SEQ ID NO: 13. In some embodiments, the target site comprises a complementary sequence of SEQ ID NO: 14. In some embodiments, the target site comprises a complementary sequence of SEQ ID NO: 15. In some embodiments, the target site comprises a complementary sequence of SEQ ID NO: 16. In some embodiments, the target site comprises a complementary sequence of SEQ ID NO: 17. In some embodiments, the target site comprises a complementary sequence of SEQ ID NO: 18. In some embodiments, the target site comprises a complementary sequence of SEQ ID NO: 19. In some embodiments, the target site comprises a complementary sequence of SEQ ID NO:20. In some embodiments, the target site comprises a complementary sequence of SEQ ID NO:21.
[0245] In some embodiments, the target site comprises the sequence set forth in SEQ ID NO:1, SEQ
ID N0:7, or SEQ ID NO:21, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO:1, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO:7, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO:21, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO:1. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO:7. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO:21. In some embodiments, the target site comprises a complementary sequence of the sequence set forth in SEQ ID NO:1. In some embodiments, the target site comprises a complementary sequence of the sequence set forth in SEQ ID NO: 7. In some embodiments, the target site comprises a complementary sequence of the sequence set forth in SEQ ID NO:21.
[0246] In some embodiments, the target site is a target site for an eZFP, such as any eZFP provided herein, for example in Section I, and/or a composition comprising the eZFP, such as an eZFP fusion protein. In some embodiments, the target site is selected from the sequence set forth in any one of SEQ ID NOS:269-300 and 583-600, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing.
[0247] In some embodiments, the target site is in a FXN promoter. In some embodiments, the target site is within a target region spanning the genomic coordinates chr9:69, 034, 900-69, 035, 900 from hg38 (SEQ ID NO:430). In some embodiments, the target site is within a target region spanning the genomic coordinates chr9:69, 035, 300-69-035, 800 from hg38. In some embodiments, the target site is within a target region spanning the genomic coordinates chr9:69, 035, 350-69, 035, 450 from hg38. In some embodiments, the target site is within a target region spanning the genomic coordinates chr9:69,035,400- 69,035,450 from hg38. In some embodiments, the target site is within a target region spanning the genomic coordinates chr9:69, 035, 530-69, 035, 580 from hg38. In some embodiments, the target site is within a target region spanning the genomic coordinates chr9:69, 035, 675-69, 035, 725 from hg38.
[0248] In some embodiments, the target site is in a FXN enhancer. In some embodiments, the target site is within a target region spanning the genomic coordinates chr9:69, 027, 282-69, 028, 497 from hg38 (SEQ ID NO:431). In some embodiments, the target site is within a target region spanning the genomic coordinates chr9:69, 027, 615-69, 028, 101 from hg38. In some embodiments, the target site is within a target region spanning the genomic coordinates chr9:69, 027, 775-69, 027, 875 from hg38. In some embodiments, the target site is within a target region spanning the genomic coordinates chr9:69,027,795- 69,027,845 from hg38.
[0249] In some embodiments, the target site comprises the nucleotide sequence set forth in any one of SEQ ID NOS:269-300 and 583-600, a contiguous portion thereof of at least 12 nt, or a complementary
sequence of any of the foregoing. In some embodiments, the target site comprises the nucleotide sequence set forth in any one of SEQ ID NOS:269-300 and 583-600. In some embodiments, the target site is comprised in double-stranded DNA, such as genomic DNA. In some embodiments, the target site is double-stranded DNA, such as genomic DNA. In some embodiments, the target site comprises a sequence set forth in Table 1.
[0250] In some embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:272, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO: 272. In some embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:277, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO: 277. In some embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:280, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO: 280. In some embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:281, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO: 281. In some embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:283, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO: 283. In some embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:290, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO: 290. In some embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:299, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO: 299. In some embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:583, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO:583. In some embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:584, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO:584. In some embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:585, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO:585. In some embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:586, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In
some embodiments, the target site comprises the sequence set forth in SEQ ID NO:586. In some embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:587, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO:587. In some embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:588, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO:588. In some embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:589, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO:589. In some embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:590, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO:590. In some embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:591, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO:591. In some embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:592, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO:592. In some embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:593, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO:593. In some embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:594, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO:594. In some embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:595, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO:595. In some embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:596, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO:596. In some embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:597, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO:597. In some embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:598, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO:598. In some
embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:599, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO:599. In some embodiments, the target site comprises the nucleotide sequence set forth in SEQ ID NO:600, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO:600.
B. Epigenetic Effector Domains for Targeted Transcriptional Activation
[0251] In some aspects, the DNA-targeting systems provided herein, such as CRISPR/Cas-based and ZFP-based DNA-targeting systems, comprise epigenetic effector domains for targeted transcriptional activation. In some aspects, fusion proteins of the DNA-targeting systems, such as eZFP fusion proteins or dCas fusion proteins, comprise the epigenetic effector domains. In some aspects, provided herein are multipartite effectors comprising two or more effector domains. In some aspects, the DNA-targeting systems, such as CRISPR/Cas-based and ZFP-based DNA-targeting systems, comprise the multipartite effectors. In some aspects, fusion proteins of the DNA-targeting systems, such as eZFP fusion proteins or dCas fusion proteins, comprise the multipartite effectors.
[0252] In some aspects, provided herein are epigenetic effector domains, such as transcriptional activation domains (i.e. effectors for transcriptional activation) and multipartite effectors for transcriptional activation (i.e. multipartite activators). In some aspects, the effector domains are provided in fusion proteins (such as eZFP fusion proteins or dCas fusion proteins), and DNA-targeting systems, such as any of the DNA-targeting systems provided herein. In some aspects, the transcriptional activation domains facilitate increased expression of FXN, for example when targeted to a FXN locus by an eZFP, eZFP fusion protein, or DNA-targeting system, such as a dCas-based DNA-targeting system. Also provided are fusion proteins, effector proteins and/or DNA-targeting systems that contain two or more of the transcriptional activation domains. In some aspects, provided herein are multipartite effectors for transcriptional activation, e.g., multipartite activators, comprising two or more effector domains such as transcriptional activation domains, such as any provided herein. In some aspects, the transcriptional activation domains and multipartite activators increase, or are capable of increasing, transcription of an endogenous locus, such as FXN, when recruited to a target site at the endogenous locus, for example increasing transcription of FXN when recruited to a target site for FXN provided herein. In some aspects, the transcriptional activation domains and multipartite activators are provided as part of a fusion protein or DNA-targeting system, such as any described herein. In some aspects, the transcriptional activation domains and multipartite activators are targeted to FXN to activate, induce, catalyze, or lead to increased transcription of the FXN gene. In some aspects, the transcriptional activation domains and multipartite activators are targeted to the target site via a DNA-targeting domain, such as a CRISPR/Cas-based, ZFN- based, or TAEE-based DNA-targeting domain, including any of the DNA-targeting domains described herein, for example, in Section I and II. In some aspects, the transcriptional activation domains and/or
multipartite effectors are targeted to the target site via an eZFP, such as as eZFP comprised in an eZFP fusion protein that also comprises the transcriptional activation domain and/or multipartite effector.
Z Transcriptional A clival ion Domains
[0253] In some aspects, provided herein are transcriptional activation domains. In some aspects, a transcriptional activation domain increases transcription of an endogenous locus, such as a FXN locus, when recruited to a target site at the endogenous locus. In some embodiments, the transcriptional activation domain is a domain that induces, catalyzes, or leads to increased transcription of a gene, such as FXN, when ectopically recruited to the gene or a DNA regulatory element thereof. In some embodiments, the transcriptional activation domain activates, induces, catalyzes, or leads to transcription activation, transcription co-activation, transcription elongation, or transcription de-repression. In some embodiments, the transcriptional activation domain induces transcriptional activation. In some embodiments, the transcriptional activation domain has one of the aforementioned activities itself (i.e., acts directly). In some embodiments, the effector domain recruits and/or interacts with a polypeptide domain that has one of the aforementioned activities (i.e., acts indirectly).
[0254] In some aspects, activation of gene expression of endogenous genes, such as human genes, can be achieved by targeting (e.g., via a CRISPR-based, ZFN-based, or TALE-based DNA-targeting domain) of transcriptional activation domains to a target site for the genes, such as regulatory DNA elements thereof (e.g., a promoter or enhancer). 0255 In some embodiments, a transcriptional activation domain provided herein comprises a domain from a human protein. In some embodiments, a transcriptional activation domain from a protein comprises any portion of the protein that is capable of acting as a transcriptional activation domain as described herein. In some embodiments, a transcription activation domain is or comprises a portion, fragment, domain or variant of a human protein, such as a portion, fragment, domain or variant of a human protein selected from among DPOLA, ENL, FOXO3, HSH2D, NCOA2, NCOA3, PSA1, PYGO1, RBM39, HERC2, and NOTCH2, that exhibits transcriptional activation, is capable of inducing or activating transcription from a gene), is a functional transcriptional activation domain, and/or has a function of transcription activation. In some embodiments, a transcription activation domain is or comprises a functional portion, a functional fragment, a functional domain or a functional variant of a human protein, such as a portion, fragment, domain or variant of a human protein selected from among DPOLA, ENL, FOXO3, HSH2D, NCOA2, NCOA3, PSA1, PYGO1, RBM39, HERC2, and NOTCH2, that exhibits transcriptional activation, is capable of inducing or activating transcription from a gene), is a functional transcriptional activation domain, and/or has a function of transcription activation. In some embodiments, a transcription activation domain is or comprises a partially or fully functional portion, a partially or fully functional fragment, a partially or fully functional domain or a partially or fully functional variant of a human protein, such as a portion, fragment, domain or variant of a human protein selected from among DPOLA, ENL, FOXO3, HSH2D, NCOA2, NCOA3, PSA1, PYGO1, RBM39,
HERC2, and N0TCH2, that exhibits increases the transcription from a gene by at least 5%, 10%, 20%, 30%, 40%' or 50% 60%, 70%, 80%, 85%, 90%-, or 100% or more, such as 2-fold, 5-fold, 10-fold, 20- fold, 30-fold, 40-fold, 50-fold. 60-fold, 70-fold, 80-fold, 90-fold. 100-fold, 200-fold, 300-fold, 400-food, 500-fold, 1000-fold or more, compared to the absence of the transcriptional activation domain.
[0256] In some embodiments, the transcriptional activation domain is 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids in length, or within a range defined by any of the foregoing. In some embodiments, the transcriptional activation domain is at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids in length. In some embodiments, the transcriptional activation domain is 10, 15, 20, 25, 30, 35, 40, 45, 50, 55, 60, 65, 70, 75, or 80 amino acids in length, or within a range defined by any of the foregoing. In some embodiments, the transcriptional activation domain is at least 10, 15, 20, 25, 30, 35, 40, 45, 50, 55, 60, 65, 70, 75, or 80 amino acids in length, or within a range defined by any of the foregoing. In some embodiments, the transcriptional activation domain is 22, 37, 42, 47, 49, 57, 61, 62, 70, 72, 76, or 80 amino acids in length, or within a range defined by any of the foregoing. In some embodiments, the transcriptional activation domain is at least 22, 37, 42, 47, 49, 57, 61, 62, 70, 72, 76, or 80 amino acids in length. In some embodiments, the transcriptional activation domain is between 10 and 80, 20 and 70, 30 and 80, 30 and 70, 30 and 60, 40 and 80, 40 and 70, 40 and 60, 40 and 50, 50 and 80, 50 and 70, 50 and 60 amino acids in length.
[0257] In some embodiments, the transcriptional activation domain comprises a transcriptional activation domain described in WO 2021/226077.
[0258] In some embodiments, a transcriptional activation domain comprises a domain from DPOLA, ENL, FOXO3, HSH2D, NCOA2, NCOA3, PSA1, PYGO1, RBM39, HERC2, or NOTCH2. In some aspects, a domain from a gene is referred to as a gene domain. For example, a domain from DPOLA may be referred to as a DPOLA domain herein. In any of the provided embodiments, the domain from DPOLA, ENL, FOXO3, HSH2D, NCOA2, NCOA3, PSA1, PYGO1, RBM39, HERC2, or NOTCH2, is or comprises the respective transcriptional activation domains described herein or a partially or fully functional fragment thereof, a domain thereof, or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 25, 30, 35, 40, 45, 50, 55, 60, 65, 70, 75, or 80 amino acids, such as at least 20 amino acids, or a variant thereof. In any of the provided embodiments, the domain from DPOLA, ENL, FOXO3, HSH2D, NCOA2, NCOA3, PSA1, PYGO1, RBM39, HERC2, or NOTCH2, is or comprises the sequence of the respective transcriptional activation domains described herein or a partially or fully functional fragment thereof, a domain thereof, or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 25, 30, 35, 40, 45, 50, 55, 60, 65, 70, 75, or 80 amino acids, such as at least 20 amino acids, or a variant thereof.
[0259] In some embodiments, the transcriptional activation domain comprises or is selected from a transcriptional activation domain shown in Table 3, or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61,
62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of transcriptional activation domain shown in Table 3, or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof. In some embodiments, the transcriptional activation domain comprises or is selected from a transcriptional activation domain shown in Table 3, or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof.
[0260] In some embodiments, the transcriptional activation domain comprises or is selected from a transcriptional activation domain shown in Table 3, or a domain or a portion thereof, such as a contiguous portion thereof of 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids in length, or within a range defined by any of the foregoing. In some embodiments, the transcriptional activation domain comprises or is selected from a transcriptional activation domain shown in Table 3, or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids in length. In some embodiments, the transcriptional activation domain comprises or is selected from a transcriptional activation domain shown in Table 3, or a domain or a portion thereof, such as a contiguous portion thereof of 10, 15, 20, 25, 30, 35, 40, 45, 50, 55, 60, 65, 70, 75, or 80 amino acids in length, or within a range defined by any of the foregoing. In some embodiments, the transcriptional activation domain comprises or is selected from a transcriptional activation domain shown in Table 3, or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 25, 30, 35, 40, 45, 50, 55, 60, 65, 70, 75, or 80 amino acids in length, or within a range defined by any of the foregoing. In some embodiments, the transcriptional activation domain comprises or is selected from a transcriptional activation domain shown in Table 3, or a domain or a portion thereof, such as a contiguous portion thereof of 22, 37, 42, 47, 49, 57, 61, 62, 70, 72, 76, or 80 amino acids in length, or within a range defined by any of the foregoing. In some embodiments, the transcriptional activation domain comprises or is selected from a transcriptional activation domain shown in Table 3, or a domain or a portion thereof, such as a contiguous portion thereof of at least 22, 37, 42, 47, 49, 57, 61, 62, 70, 72, 76, or 80 amino acids in length. In some embodiments, the transcriptional activation domain comprises or is selected from a transcriptional activation domain shown in Table 3, or a domain or a portion thereof, such as a contiguous portion thereof of between 10 and 80, 20 and 70, 30 and 80, 30 and 70, 30 and 60, 40 and 80, 40 and 70, 40 and 60, 40 and 50, 50 and 80, 50 and 70, 50 and 60 amino acids in length. In some embodiments, the transcriptional activation domain is a transcriptional activation domain set forth in Table 3. Table 3 shows a list of human genes and exemplary transcriptional activation domains from each gene.
[0261] In some embodiments, any of the provided multipartite effector proteins, fusion proteins,
and/or DNA targeting systems, such as a multipartite activator, comprises a combination of transcriptional activation domains, such as a combination of two or more, such as three or more, such as three or more, of any of transcriptional activation domains shown in Table 3. In some embodiments, any of the provided multipartite effector proteins, fusion proteins, and/or DNA targeting systems, such as a multipartite activator, comprises a combination of two or more, such as three or more, of any one of the SEQ ID NOS: set forth in Table 3, or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any one of the SEQ ID NOS: set forth in Table 3, or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof. In some embodiments, any of the provided multipartite effector proteins, fusion proteins, and/or DNA targeting systems, such as a multipartite activator, comprises a combination of two or more, such as three or more, of any one of the SEQ ID NOS: set forth in Table 3, or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof. In some embodiments, any of the provided multipartite effector proteins, fusion proteins, and/or DNA targeting systems, such as a multipartite activator, comprises two or more, such as three or more, of any one of the SEQ ID NOS: set forth in Table 3.
[0262] In some embodiments, the transcriptional activation domain comprises any one of SEQ ID NOS: 113-122 and 124, or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino
acids, such as at least 20 amino acids, or a variant thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any one of SEQ ID NOS: 113-122 and 124, or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof.
[0263] In some embodiments, the transcriptional activation domain comprises any one of SEQ ID NOS:100-109, 111, and 383-393, or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any one of SEQ ID NOS:100-109, 111, and 383-393, or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof.
[0264] In some embodiments, a transcriptional activation domain comprises a DPOLA domain, i.e. a domain from DPOLA. In some aspects, DPOLA refers to the DNA polymerase alpha catalytic subunit protein encoded by the human POLA1 gene. DPOLA plays an essential role in the initiation of DNA synthesis. An exemplary human DPOLA sequence is set forth in SEQ ID NO: 113. An exemplary DPOLA domain sequence is set forth in SEQ ID NO: 100 and SEQ ID NO:383. In some embodiments, the transcriptional activation domain comprises a sequence set forth in any of SEQ ID NOS: 113, 100, and 383 or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to a sequence set forth in any of SEQ ID NOS: 113, 100, and 383 or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof. In some embodiments, the transcriptional activation domain is or comprises an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO: 100. In some embodiments, the transcriptional activation domain comprises a contiguous portion of SEQ ID NO: 113 that is at least 80 amino acids in length. In some embodiments, the transcriptional activation domain comprises SEQ ID NO: 100. In some embodiments, the transcriptional activation domain is set forth in SEQ ID NO: 100. An exemplary nucleotide sequence encoding the transcriptional activation domain set forth in SEQ ID NO: 100 is set forth in SEQ ID NO:87. In some embodiments, the transcriptional activation domain is or comprises an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO: 383. In some embodiments, the transcriptional activation domain comprises a contiguous portion of SEQ ID NO: 113 that is at least 61 amino acids in length. In some embodiments, the transcriptional activation domain comprises SEQ ID NO: 383. In some embodiments,
the transcriptional activation domain is set forth in SEQ ID NO:383.
[0265] In some embodiments, a transcriptional activation domain comprises a ENL domain, i.e. a domain from ENL. In some aspects, ENL refers to the ENL protein encoded by the human MLLT1 gene. ENL functions as a chromatin reader component of the super elongation complex (SEC), a complex which increases the catalytic rate of RNA polymerase II transcription. An exemplary human ENL sequence is set forth in SEQ ID NO: 114. An exemplary ENL domain sequence is set forth in SEQ ID NO: 101 and SEQ ID NO:384. In some embodiments, the transcriptional activation domain comprises a sequence set forth in any of SEQ ID NOS:114, 101, and 384 or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to a sequence set forth in any of SEQ ID NOS: 114, 101, and 384 or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof. In some embodiments, the transcriptional activation domain is or comprises an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO: 101. An exemplary nucleotide sequence encoding the transcriptional activation domain set forth in SEQ ID NO: 101 is set forth in SEQ ID NO:88. In some embodiments, the transcriptional activation domain comprises a contiguous portion of SEQ ID NO: 114 that is at least 80 amino acids in length. In some embodiments, the transcriptional activation domain comprises SEQ ID NO: 101. In some embodiments, the transcriptional activation domain is set forth in SEQ ID NO: 101. In some embodiments, the transcriptional activation domain is or comprises an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO:384. In some embodiments, the transcriptional activation domain comprises a contiguous portion of SEQ ID NO: 114 that is at least 62 amino acids in length. In some embodiments, the transcriptional activation domain comprises SEQ ID NO:384. In some embodiments, the transcriptional activation domain is set forth in SEQ ID NO:384.
[02663 In some embodiments, a transcriptional activation domain comprises a FOXO3 domain, i.e. a domain from FOXO3. In some aspects, FOXO3 refers to the Forkhead box protein 03 encoded by the human FOXO3 gene. FOXO3 functions as a transcriptional activator that recognizes and binds to specific DNA sequences. An exemplary human FOXO3 sequence is set forth in SEQ ID NO: 115. An exemplary FOXO3 domain sequence is set forth in SEQ ID NO: 102 and SEQ ID NO:385. In some embodiments, the transcriptional activation domain comprises a sequence set forth in any of SEQ ID NOS: 115, 102, and 385 or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to a sequence set forth in
any of SEQ ID NOS: 115, 102, and 385 or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof. In some embodiments, the transcriptional activation domain is or comprises an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO: 102. In some embodiments, the transcriptional activation domain comprises a contiguous portion of SEQ ID NO: 115 that is at least 80 amino acids in length. In some embodiments, the transcriptional activation domain comprises SEQ ID NO: 102. In some embodiments, the transcriptional activation domain is set forth in SEQ ID NO: 102. An exemplary nucleotide sequence encoding the transcriptional activation domain set forth in SEQ ID NO: 102 is set forth in SEQ ID NO: 88. In some embodiments, the transcriptional activation domain is or comprises an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO:385. In some embodiments, the transcriptional activation domain comprises a contiguous portion of SEQ ID NO: 115 that is at least 42 amino acids in length. In some embodiments, the transcriptional activation domain comprises SEQ ID NO:385. In some embodiments, the transcriptional activation domain is set forth in SEQ ID NO:385.
[0267] In some embodiments, a transcriptional activation domain comprises a HSH2D domain, i.e. a domain from HSH2D. In some aspects, HSH2D refers to the Hematopoietic SH2 domain-containing protein encoded by the human HSH2D gene. HSH2D functions as an adapter protein involved in tyrosine kinase and CD28 signaling. An exemplary human HSH2D sequence is set forth in SEQ ID NO: 116. An exemplary HSH2D domain sequence is set forth in SEQ ID NO: 103 and SEQ ID NO:386. In some embodiments, the transcriptional activation domain comprises a sequence set forth in any of SEQ ID NOS: 116, 103, and 386 or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to a sequence set forth in any of SEQ ID NOS: 116, 103, and 386 or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof. In some embodiments, the transcriptional activation domain is or comprises an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO: 103. In some embodiments, the transcriptional activation domain comprises a contiguous portion of SEQ ID NO: 116 that is at least 80 amino acids in length. In some embodiments, the transcriptional activation domain comprises SEQ ID NO: 103. In some embodiments, the transcriptional activation domain is set forth in SEQ ID NO: 103. An exemplary nucleotide sequence encoding the transcriptional activation domain set forth in SEQ ID NO: 103 is set forth in SEQ ID NO: 90. In some embodiments, the transcriptional activation domain is or comprises an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO:386. In some embodiments, the
transcriptional activation domain comprises a contiguous portion of SEQ ID NO: 116 that is at least 76 amino acids in length. In some embodiments, the transcriptional activation domain comprises SEQ ID NO:386. In some embodiments, the transcriptional activation domain is set forth in SEQ ID NO:386.
[0268] In some embodiments, a transcriptional activation domain comprises a NCOA2 domain, i.e. a domain from NCOA2. In some aspects, NCOA2 refers to the Nuclear receptor coactivator 2 protein encoded by the human NCOA2 gene. NCOA2 functions as a transcriptional coactivator for steroid receptors and nuclear receptors. An exemplary human NCOA2 sequence is set forth in SEQ ID NO: 117. An exemplary NCOA2 domain sequence is set forth in SEQ ID NO: 104 and SEQ ID NO:387. In some embodiments, the transcriptional activation domain comprises a sequence set forth in any of SEQ ID NOS: 117, 104, and 387 or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to a sequence set forth in any of SEQ ID NOS: 117, 104, and 387 or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof. In some embodiments, the transcriptional activation domain is or comprises an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO: 104. In some embodiments, the transcriptional activation domain comprises a contiguous portion of SEQ ID NO: 117 that is at least 80 amino acids in length. In some embodiments, the transcriptional activation domain comprises SEQ ID NO: 104. In some embodiments, the transcriptional activation domain is set forth in SEQ ID NO: 104. An exemplary nucleotide sequence encoding the transcriptional activation domain set forth in SEQ ID NO: 104 is set forth in SEQ ID NO: 91. In some embodiments, the transcriptional activation domain is or comprises an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO:387. In some embodiments, the transcriptional activation domain comprises a contiguous portion of SEQ ID NO: 117 that is at least 47 amino acids in length. In some embodiments, the transcriptional activation domain comprises SEQ ID NO:387. In some embodiments, the transcriptional activation domain is set forth in SEQ ID NO:387.
[0269] In some embodiments, a transcriptional activation domain comprises a NCOA3 domain, i.e. a domain from NCOA3. In some aspects, NCOA3 refers to the Nuclear receptor coactivator 3 protein encoded by the human NCOA3 gene. NCOA3 functions as a transcriptional coactivator for steroid receptors and nuclear receptors. An exemplary human NCOA3 sequence is set forth in SEQ ID NO: 118. An exemplary NCOA3 domain sequence is set forth in SEQ ID NO: 105 and SEQ ID NO:388. In some embodiments, the transcriptional activation domain comprises a sequence set forth in any of SEQ ID NOS: 118, 105, and 388 or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof, or an amino acid sequence that has at least
90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to a sequence set forth in any of SEQ ID NOS: 118, 105, and 388 or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof. In some embodiments, the transcriptional activation domain is or comprises an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO: 105. In some embodiments, the transcriptional activation domain comprises a contiguous portion of SEQ ID NO: 118 that is at least 80 amino acids in length. In some embodiments, the transcriptional activation domain comprises SEQ ID NO: 105. In some embodiments, the transcriptional activation domain is set forth in SEQ ID NO: 105. An exemplary nucleotide sequence encoding the transcriptional activation domain set forth in SEQ ID NO: 105 is set forth in SEQ ID NO: 92. In some embodiments, the transcriptional activation domain is or comprises an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO:388. In some embodiments, the transcriptional activation domain comprises a contiguous portion of SEQ ID NO: 118 that is at least 49 amino acids in length. In some embodiments, the transcriptional activation domain comprises SEQ ID NO: 388. In some embodiments, the transcriptional activation domain is set forth in SEQ ID NO:388.
[0270] In some embodiments, a transcriptional activation domain comprises a PSA1 domain, i.e. a domain from PSA1. In some aspects, PSA1 refers to the Proteasome subunit alpha type-1 protein encoded by the human PSMA1 gene. PSA1 functions as a component of the 20S core proteasome complex, which facilitates proteolytic degradation of intracellular proteins. An exemplary human PSA1 sequence is set forth in SEQ ID NO: 119. An exemplary PSA1 domain sequence is set forth in SEQ ID NO: 106 and SEQ ID NO:389. In some embodiments, the transcriptional activation domain comprises a sequence set forth in any of SEQ ID NOS: 119, 106, and 389 or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to a sequence set forth in any of SEQ ID NOS: 119, 106, and 389 or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof. In some embodiments, the transcriptional activation domain is or comprises an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO: 106. In some embodiments, the transcriptional activation domain comprises a contiguous portion of SEQ ID NO: 119 that is at least 80 amino acids in length. In some embodiments, the transcriptional activation domain comprises SEQ ID NO: 106. In some embodiments, the transcriptional activation domain is set forth in SEQ ID NO: 106. An exemplary nucleotide sequence encoding the transcriptional activation domain set forth in SEQ ID NO: 106 is set forth in SEQ ID NO:93. In some embodiments, the transcriptional activation domain is or comprises an amino acid sequence that has at
least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO:389. In some embodiments, the transcriptional activation domain comprises a contiguous portion of SEQ ID NO: 119 that is at least 22 amino acids in length. In some embodiments, the transcriptional activation domain comprises SEQ ID NO:389. In some embodiments, the transcriptional activation domain is set forth in SEQ ID NO:389.
[0271] In some embodiments, a transcriptional activation domain comprises a PYGO1 domain, i.e. a domain from PYGO1. In some aspects, PYGO1 refers to the Pygopus homolog 1 protein encoded by the human PYGO1 gene. PYGO1 is involved in Wnt pathway signal transduction. An exemplary human PYGO1 sequence is set forth in SEQ ID NO: 120. An exemplary PYGO1 domain sequence is set forth in SEQ ID NO: 107 and SEQ ID NO:390. In some embodiments, the transcriptional activation domain comprises a sequence set forth in any of SEQ ID NOS:120, 107, and 390 or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to a sequence set forth in any of SEQ ID NOS: 120, 107, and 390 or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof. In some embodiments, the transcriptional activation domain is or comprises an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO: 107. In some embodiments, the transcriptional activation domain comprises a contiguous portion of SEQ ID NO: 120 that is at least 80 amino acids in length. In some embodiments, the transcriptional activation domain comprises SEQ ID NO: 107. In some embodiments, the transcriptional activation domain is set forth in SEQ ID NO: 107. An exemplary nucleotide sequence encoding the transcriptional activation domain set forth in SEQ ID NO: 107 is set forth in SEQ ID NO:94. In some embodiments, the transcriptional activation domain is or comprises an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO:390. In some embodiments, the transcriptional activation domain comprises a contiguous portion of SEQ ID NO: 120 that is at least 57 amino acids in length. In some embodiments, the transcriptional activation domain comprises SEQ ID NO:390. In some embodiments, the transcriptional activation domain is set forth in SEQ ID NO:390.
[0272] In some embodiments, a transcriptional activation domain comprises a RBM39 domain, i.e. a domain from RBM39. In some aspects, RBM39 refers to the RNA-binding protein 39 protein encoded by the human RBM39 gene. RBM39 functions as a RNA-binding protein that acts as a pre-mRNA splicing factor. An exemplary human RBM39 sequence is set forth in SEQ ID NO: 121. An exemplary RBM39 domain sequence is set forth in SEQ ID NO:108 and SEQ ID NO:391. In some embodiments, the transcriptional activation domain comprises a sequence set forth in any of SEQ ID NOS:121, 108, and 391 or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25,
30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to a sequence set forth in any of SEQ ID NOS: 121, 108, and 391 or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof. In some embodiments, the transcriptional activation domain is or comprises an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO: 108. In some embodiments, the transcriptional activation domain comprises a contiguous portion of SEQ ID NO: 121 that is at least 80 amino acids in length. In some embodiments, the transcriptional activation domain comprises SEQ ID NO: 108. In some embodiments, the transcriptional activation domain is set forth in SEQ ID NO: 108. An exemplary nucleotide sequence encoding the transcriptional activation domain set forth in SEQ ID NO: 108 is set forth in SEQ ID NO:95. In some embodiments, the transcriptional activation domain is or comprises an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO:391. In some embodiments, the transcriptional activation domain comprises a contiguous portion of SEQ ID NO: 121 that is at least 70 amino acids in length. In some embodiments, the transcriptional activation domain comprises SEQ ID NO:391. In some embodiments, the transcriptional activation domain is set forth in SEQ ID NO:391.
[0273] In some embodiments, a transcriptional activation domain comprises a HERC2 domain, i.e. a domain from HERC2. In some aspects, HERC2 refers to the E3 ubiquitin-protein ligase HERC2 protein encoded by the human HERC2 gene. HERC2 functions as a regulator of ubiquitin-dependent retention of repair proteins on damaged chromosomes. An exemplary human HERC2 sequence is set forth in SEQ ID NO: 122. An exemplary HERC2 domain sequence is set forth in SEQ ID NO: 109 and SEQ ID NO:392. In some embodiments, the transcriptional activation domain comprises a sequence set forth in any of SEQ ID NOS: 122, 109, and 392 or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to a sequence set forth in any of SEQ ID NOS: 122, 109, and 392 or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof. In some embodiments, the transcriptional activation domain is or comprises an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO:109. In some embodiments, the transcriptional activation domain comprises a contiguous portion of SEQ ID NO: 122 that is at least 80 amino acids in length. In some embodiments, the transcriptional activation domain comprises SEQ ID NO: 109. In some embodiments, the transcriptional activation domain is set forth in SEQ ID NO: 109. An exemplary nucleotide sequence encoding the transcriptional activation
domain set forth in SEQ ID NO: 109 is set forth in SEQ ID NO:96. In some embodiments, the transcriptional activation domain is or comprises an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO:392. In some embodiments, the transcriptional activation domain comprises a contiguous portion of SEQ ID NO: 122 that is at least 72 amino acids in length. In some embodiments, the transcriptional activation domain comprises SEQ ID NO:392. In some embodiments, the transcriptional activation domain is set forth in SEQ ID NO:392.
[0274] In some embodiments, a transcriptional activation domain comprises a NOTCH2 domain, i.e. a domain from NOTCH2. In some aspects, NOTCH2 refers to the Neurogenic locus notch homolog protein 2 protein encoded by the human NOTCH2 gene. NOTCH2 functions as a receptor for membranebound ligands such as Delta- 1 to regulate cell-fate determination. An exemplary human NOTCH2 sequence is set forth in SEQ ID NO: 124. An exemplary NOTCH2 domain sequence is set forth in SEQ ID NO:111 and SEQ ID NO:393. In some embodiments, the transcriptional activation domain comprises a sequence set forth in any of SEQ ID NOS:124, 111, and 393 or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to a sequence set forth in any of SEQ ID NOS:124, 111, and 393 or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof. In some embodiments, the transcriptional activation domain is or comprises an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO: 111. In some embodiments, the transcriptional activation domain comprises a contiguous portion of SEQ ID NO: 124 that is at least 80 amino acids in length. In some embodiments, the transcriptional activation domain comprises SEQ ID NO: 111. In some embodiments, the transcriptional activation domain is set forth in SEQ ID NO: 111. An exemplary nucleotide sequence encoding the transcriptional activation domain set forth in SEQ ID NO:111 is set forth in SEQ ID NO: 98. In some embodiments, the transcriptional activation domain is or comprises an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO:393. In some embodiments, the transcriptional activation domain comprises a contiguous portion of SEQ ID NO: 124 that is at least 37 amino acids in length. In some embodiments, the transcriptional activation domain comprises SEQ ID NO:393. In some embodiments, the transcriptional activation domain is set forth in SEQ ID NO:393.
[0275] A variety of other effector domains for transcriptional activation (e.g., transcriptional activation domains) are known and can be used in accord with or in conjunction with the provided embodiments. Other transcriptional activation domains for targeted activation are described, for example, in WO 2014/197748, WO 2016/130600, WO 2017/180915, WO 2021/226555, WO 2021/226077, WO
2013/176772, WO 2014/152432, WO 2014/093661, WO 2021/247570, Adli, M. Nat. Commun. 9, 1911 (2018), Perez-Pinera, P. et al. Nat. Methods 10, 973-976 (2013), Mali, P. et al. Nat. Biotechnol. 31, 833— 838 (2013), Maeder, M. L. et al. Nat. Methods 10, 977-979 (2013), Gilbert, L. A. et al. Cell 154(2):442- 451 (2013), and Nunez, J.K. et al. Cell 184(9):2503-2519 (2021).
[0276] In some embodiments, a transcriptional activation domain comprises a domain of a protein selected from among VP64, p65, Rta, p300, CBP, VPR, VPH, HSF1, a TET protein (e.g. TET1), a partially or fully functional fragment or domain thereof, or a combination of any of the foregoing.
[0277] In some embodiments, the transcriptional activation domain comprises a VP64 domain. For example, dCas9-VP64 can be targeted to a target site by one or more gRNAs to activate a gene. VP64 is a polypeptide composed of four tandem copies of VP 16, a 16 amino acid transactivation domain of the Herpes simplex virus. VP64 domains, including in dCas fusion proteins, have been described, for example, in WO 2014/197748, WO 2013/176772, WO 2014/152432, and WO 2014/093661. In some embodiments, the transcriptional activation domain comprises at least one VP16 domain, or a VP16 tetramer (“VP64”) or a variant thereof. An exemplary VP64 domain is set forth in SEQ ID NO:81. An exemplary nucleotide sequence encoding the exemplary VP64 domain set forth in SEQ ID NO: 81 is set forth in SEQ ID NO: 80. In some embodiments, the transcriptional activation domain comprises SEQ ID NO:81, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO:81, or a portion thereof. In some embodiments, the transcriptional activation domain is set forth in SEQ ID NO:81.
[0278] In some embodiments, the transcriptional activation domain comprises a p65 activation domain (p65AD). p65AD is the principal transactivation domain of the 65kDa polypeptide of the nuclear form of the NF-KB transcription factor. An exemplary sequence of human transcription factor p65 is available at the Uniprot database under accession number Q04206. p65 domains, including in dCas fusion proteins, have been described, for example in WO 2017/180915 and Chavez, A. et al. Nat. Methods 12, 326-328 (2015). An exemplary p65 activation domain is set forth in SEQ ID NO: 134. In some embodiments, the transcriptional activation domain comprises SEQ ID NO: 134, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO: 134, or a portion thereof. In some embodiments, the transcriptional activation domain is set forth in SEQ ID NO: 134.
[0279] In some embodiments, the transcriptional activation domain comprises an R transactivator (Rta) domain. Rta is an immediate-early protein of Epstein-Barr virus (EBV),and is a transcriptional activator that induces lytic gene expression and triggers virus reactivation. The Rta domain, including in dCas fusion proteins, has been described, for example in WO 2017/180915 and Chavez, A. et al. Nat. Methods 12, 326-328 (2015). An exemplary Rta domain is set forth in SEQ ID NO: 135. In some embodiments, the transcriptional activation domain comprises SEQ ID NO: 135, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO: 135, or a portion thereof. In some embodiments, the transcriptional
activation domain is set forth in SEQ ID NO: 135.
[0280] The transcriptional activation domain comprises a CREB-binding protein (CBP) domain or a p300 domain. In some aspects, CBP refers to the CREB-binding protein encoded by the human CREBBP gene. CBP is a coactivator that interacts with cAMP-response element binding protein (CREB). In some aspects, p300 refers to the Histone acetyltransferase p300 protein encoded by the human EP300 gene, and is a coactivator closely related to CBP. CBP and p300 each interact with a variety of transcriptional activators to affect gene transcription (Gerritsen, M.E. et al. PNAS 94(7):2927-2932 (1997)). In some embodiments, the transcriptional activation domain comprises a p300 domain. p300 domains (such as the catalytic core of p300) including in dCas fusion proteins for gene activation, has been described, for example, in WO 2016/130600, WO 2017/180915, and Hilton, I.B. et al., Nat. Biotechnol. 33(5):510-517 (2015). An exemplary human CBP sequence is set forth in SEQ ID NO:394. An exemplary human p300 sequence is set forth in SEQ ID NO: 125. An exemplary p300 domain is set forth in SEQ ID NO: 112. In some embodiments, the transcriptional activation domain comprises any one of SEQ ID NOS:394, 125, and 112, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any one of SEQ ID NOS: 394, 125, and 112, or a portion thereof. In some embodiments, the transcriptional activation domain comprises SEQ ID NO: 112, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO: 112, or a portion thereof. In some embodiments, the transcriptional activation domain is set forth in SEQ ID NO: 112.
[0281] In some embodiments, the transcriptional activation domain comprises a HSF1 domain. In some aspects, HSF1 refers to the Heat shock factor protein 1 protein encoded by the human HSF1 gene. HSF1, including in dCas fusion proteins for gene activation, has been described, for example, in WO 2021/226555, WO 2015/089427, and Konermann et al. Nature 517(7536):583-8 (2015). An exemplary human HSF1 sequence is set forth in SEQ ID NO:395. An exemplary HSF1 domain sequence is set forth in SEQ ID NO: 136. In some embodiments, the transcriptional activation domain comprises SEQ ID NO:136 or SEQ ID NO:395, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO: 136 or SEQ ID NO:395, or a portion thereof. In some embodiments, the transcriptional activation domain comprises SEQ ID NO:136, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO: 136, or a portion thereof. In some embodiments, the transcriptional activation domain is set forth in SEQ ID NO: 136.
[0282] In some embodiments, the transcriptional activation domain comprises the tripartite activator VP64-p65-Rta (also known as VPR). VPR comprises three transcription activation domains (VP64, p65, and Rta) fused by short amino acid linkers, and can effectively upregulate target gene expression. VPR, including in dCas fusion proteins for gene activation, has been described, for example, in WO 2021/226555 and Chavez, A. et al. Nat. Methods 12, 326-328 (2015). An exemplary VPR polypeptide is set forth in SEQ ID NO: 137. In some embodiments, the transcriptional activation domain comprises SEQ
ID NO:137, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO: 137, or a portion thereof. In some embodiments, the transcriptional activation domain is set forth in SEQ ID NO: 137.
[0283] In some embodiments, the transcriptional activation domain comprises VPH. VPH is a tripartite activator polypeptide comprising VP64, mouse p65, and HSF1. VPH, including in dCas fusion proteins for gene activation, has been described, for example, in WO 2021/226555. An exemplary VPH polypeptide is set forth in SEQ ID NO: 138. In some embodiments, the transcriptional activation domain comprises SEQ ID NO: 138, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO: 138, or a portion thereof. In some embodiments, the transcriptional activation domain is set forth in SEQ ID NO: 138.
[0284| In some embodiments, the transcriptional activation domain has demethylase activity. The transcriptional activation domain can include an enzyme that removes methyl (CH3-) groups from nucleic acids, proteins (in particular histones), and other molecules. Alternatively, the transcriptional activation domain can convert the methyl group to hydroxymethylcytosine in a mechanism for demethylating DNA. The effector domain can catalyze this reaction. For example, the transcriptional activation domain that catalyzes this reaction may comprise a domain from a TET protein, for example TET1 (Ten-eleven translocation methylcytosine dioxygenase 1). In some aspects, TET1 refers to the Methylcytosine dioxygenase TET1 protein encoded by the human TET1 gene. TET1 catalyzes the conversion of the modified genomic base 5 -methylcytosine (5mC) into 5-hydroxymethylcytosine (5hmC) and plays a key role in active DNA demethylation. TET1, including in dCas fusion proteins for gene activation, has been described, for example, in WO 2021/226555. An exemplary human TET1 sequence is set forth in SEQ ID NO:396. An exemplary TET1 catalytic domain is set forth in SEQ ID NO: 139. In some embodiments, the transcriptional activation domain comprises SEQ ID NO:396 or SEQ ID NO:139, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO:396 or SEQ ID NO: 139, or a portion thereof. In some embodiments, the transcriptional activation domain comprises SEQ ID NO: 139, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO: 139, or a portion thereof. In some embodiments, the transcriptional activation domain is set forth in SEQ ID NO: 139.
[0285] In some embodiments, the effector domain may comprise a LSD1 domain. LSD1 (also known as Lysine-specific histone demethylase 1A) is a histone demethylase that can demethylate lysine residues of histone H3, thereby acting as a coactivator or a corepressor, depending on the context. LSD1, including in dCas fusion proteins, has been described, for example, in WO 2013/176772, WO 2014/152432, and Kearns, N. A. et al. Nat. Methods. 12(5):401-403 (2015). An exemplary LSD1 polypeptide is set forth in SEQ ID NO: 140. In some embodiments, the effector domain comprises the sequence set forth in SEQ ID NO: 140, a domain thereof, a portion thereof, or a variant thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99%
sequence identity to any of the foregoing.
[0286] In some embodiments, the effector domain may comprise a SunTag domain. SunTag is a repeating peptide array, which can recruit multiple copies of an antibody-fusion protein that binds the repeating peptide. The antibody-fusion protein may comprise an additional effector domain, such as a transcription activation domain (e.g. VP64), to induce increased transcription of the target gene. SunTag, including in dCas fusion proteins for gene activation, has been described, for example, in WO 2016/011070 and Tanenbaum, M. et al. Cell. 159(3):635-646 (2014). An exemplary SunTag effector domain includes a repeating GCN4 peptide having the amino acid sequence LLPKNYHLENEVARLKKLVGER (SEQ ID NO: 152) separated by linkers having the amino acid sequence GGSGG (SEQ ID NO: 153). In some embodiments, the effector domain comprises the sequence set forth in SEQ ID NO: 152, a domain thereof, a portion thereof, or a variant thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing. In some embodiments, the SunTag effector domain recruits an antibody-fusion protein that comprises any of the epigenetic effector domains described herein, such as VP64, and binds the GCN4 peptide. In some embodiments, the SunTag domain recruits one or more epigenetic effector domains for transcriptional activation to the FXN locus, thereby increasing FXN expression.
[0287] The effector domain may have demethylase activity. The effector domain may include an enzyme that remove methyl (CH3-) groups from nucleic acids, proteins (in particular histones), and other molecules. Alternatively, the effector domain may covert the methyl group to hydroxymethylcytosine in a mechanism for demethylating DNA.
2. Multipartite Effectors for Transcriptional A ctivation
[0288] In some aspects, provided herein are multipartite effectors for transcriptional activation of FXN, for example, multipartite transcriptional activation domains or multipartite activators. In some aspects, the epigenetic effector domain is a multipartite effector, such as a multipartite activator. In some aspects, the multipartite activator is a fusion protein or a sequence of amino acids comprising two or more transcriptional activation domains, such as any of the effector domains such as transcriptional activation domains provided herein. In some aspects, the multipartite activator comprises two or more transcriptional activation domains, each transcriptional activation domain comprising a domain of a protein selected from among NCOA3, ENL, FOXO3, PYGO1, HSH2D, NCOA2, NOTCH2, DPOLA, PSA1, RBM39, and HERC2. In some aspects, the multipartite activator is provided as part of a fusion protein or DNA-targeting system, such as any described herein, including an eZFP fusion protein, a ZFP- based DNA-targeting system, and a CRISPR/Cas-based DNA-targeting system.
[0289] In some aspects, the multipartite activator increases transcription of an endogenous locus, such as FXN, when recruited to a target site at the endogenous locus. For example, the multipartite activator increases transcription of a FXN gene when recruited (e.g. targeted to) a target site for the FXN gene, such as a regulatory DNA element (e.g. a promoter or enhancer), and/or any target site described
herein. Thus, in some aspects, a multipartite activator may itself be referred to as a transcriptional activation domain herein. In some embodiments, the multipartite activator induces, catalyzes, or leads to increased transcription of FXN when ectopically recruited to the FXN locus or a DNA regulatory element thereof. In some embodiments, the multipartite activator activates, induces, catalyzes, or leads to: transcription activation, transcription co-activation, transcription elongation, or transcription derepression. In some embodiments, the multipartite activator induces transcriptional activation. In some embodiments, the multipartite activator has one of the aforementioned activities itself (i.e. acts directly). In some embodiments, the multipartite activator recruits and/or interacts with a polypeptide domain that has one of the aforementioned activities (i.e. acts indirectly).
[0290] In some aspects, a multipartite activator provided herein comprises two or more transcriptional activation domains. In some aspects, the multipartite activator has an effect that is different from any one of the individual transcriptional activation domains comprised by the multipartite activator alone. The different effect may be quantitatively or qualitatively different. The multipartite activator may induce greater, more reliable, or more durable transcriptional activation of a target gene, in comparison to a transcriptional activation domain alone. The effect may be context-specific. For example, a multipartite activator may induce transcriptional activation in a specific context in which the transcriptional activation domain alone does not induce transcriptional activation to the same degree, at a detectable level, or at all, such as when targeted to a specific gene or target site of the gene. Thus, a multipartite activator does not necessarily lead to greater activation of a target gene than a transcriptional activation domain alone in every context, but may allow for activation of a target gene in different contexts and to a different degree than the transcriptional activation domain. A multipartite activator may have a more durable effect on transcription than a transcriptional activation domain alone. For example, a multipartite activator may lead to increased transcription of a target gene in a cell for a longer amount of time, or for a greater number of cell divisions or cell passages.
[0291] In some embodiments, the multipartite effector, e.g., multipartite activator, is a bipartite effector, e.g., bipartite activator (i.e. comprising two transcriptional activation domains). In some embodiments, the multipartite effector, e.g., multipartite activator, is a tripartite effector, e.g., tripartite activator (i.e. comprising three transcriptional activation domains). In some embodiments, the multipartite effector, e.g., multipartite activator comprises 4, 5, 6, 7, 8, 9, 10, or more transcriptional activation domains. In some embodiments, any two or more of the transcriptional activation domains are the same. In some embodiments, any two or more of the transcriptional activation domains are different.
[0292] In some embodiments, the multipartite activator comprises two or more transcriptional activation domains, each transcriptional activation domain comprising a domain of a protein selected from among DPOLA, ENL, FOXO3, HSH2D, NCOA2, NCOA3, PSA1, PYGO1, RBM39, HERC2, or NOTCH2. In some embodiments, the multipartite activator comprises two or more transcriptional activation domains, wherein one or more of the transcriptional activation domains comprises a domain of a protein selected from among DPOLA, ENL, FOXO3, HSH2D, NCOA2, NCOA3, PSA1, PYGO1,
RBM39, HERC2, or N0TCH2. In some aspects, the transcriptional activation domain from DPOLA, ENL, FOXO3, HSH2D, NC0A2, NC0A3, PSA1, PYGO1, RBM39, HERC2, or NOTCH2, is or comprises any of the respective transcriptional activation domains described herein, for example, in Section II.B.l, or a partially or fully functional fragment thereof, a domain thereof, or a portion thereof, such as a contiguous portion thereof of at least 30 amino acids, or a variant thereof. In some aspects, the transcriptional activation domain from DPOLA, ENL, FOXO3, HSH2D, NCOA2, NCOA3, PSA1, PYGO1, RBM39, HERC2, or NOTCH2, is or comprises any of sequences of the respective transcriptional activation domains described herein, for example, in Section II.B.l, or a partially or fully functional fragment thereof, a domain thereof, or a portion thereof, such as a contiguous portion thereof of at least 30 amino acids, or a variant thereof.
[02933 In some embodiments, the multipartite activator further comprises one or more of any of the transcriptional activation domains provided herein, such as VP64, p65, Rta, p300, CBP, VPR, VPH, HSF1, a TET protein (e.g., TET1), a partially or fully functional fragment or domain thereof, or a combination of any of the foregoing.
[0294] In some embodiments, the multipartite activator is a bipartite activator comprising a first transcriptional activation domain and a second transcriptional activation domain. In some aspects, each of the first transcriptional activation domain and the second transcriptional activation domain independently comprises a domain of a protein selected from among DPOLA, ENL, FOXO3, HSH2D, NCOA2, NCOA3, PSA1, PYGO1, RBM39, HERC2, and NOTCH2. In some embodiments, the first and second transcriptional activation domains, respectively, are from DPOLA and DPOLA; DPOLA and ENL; DPOLA and FOXO3; DPOLA and HERC2; DPOLA and HSH2D; DPOLA and NCOA2; DPOLA and NCOA3; DPOLA and NOTCH2; DPOLA and PSA1; DPOLA and PYGO1; DPOLA and RBM39; ENL and DPOLA; ENL and ENL; ENL and FOXO3; ENL and HERC2; ENL and HSH2D; ENL and NCOA2; ENL and NCOA3; ENL and NOTCH2; ENL and PSA1; ENL and PYGO1; ENL and RBM39; FOXO3 and DPOLA; FOXO3 and ENL; FOXO3 and FOXO3; FOXO3 and HERC2; FOXO3 and HSH2D; FOXO3 and NCOA2; FOXO3 and NCOA3; FOXO3 and NOTCH2; FOXO3 and PSA1; FOXO3 and PYGO1; FOXO3 and RBM39; HERC2 and DPOLA; HERC2 and ENL; HERC2 and FOXO3; HERC2 and HERC2; HERC2 and HSH2D; HERC2 and NCOA2; HERC2 and NCOA3; HERC2 and NOTCH2; HERC2 and PSA1; HERC2 and PYGO1; HERC2 and RBM39; HSH2D and DPOLA; HSH2D and ENL; HSH2D and FOXO3; HSH2D and HERC2; HSH2D and HSH2D; HSH2D and NCOA2; HSH2D and NCOA3; HSH2D and NOTCH2; HSH2D and PSA1; HSH2D and PYGO1; HSH2D and RBM39; NCOA2 and DPOLA; NCOA2 and ENL; NCOA2 and FOXO3; NCOA2 and HERC2; NCOA2 and HSH2D; NCOA2 and NCOA2; NCOA2 and NCOA3; NCOA2 and NOTCH2; NCOA2 and PSA1; NCOA2 and PYGO1; NCOA2 and RBM39; NCOA3 and DPOLA; NCOA3 and ENL; NCOA3 and FOXO3; NCOA3 and HERC2; NCOA3 and HSH2D; NCOA3 and NCOA2; NCOA3 and NCOA3; NCOA3 and NOTCH2; NCOA3 and PSA1; NCOA3 and PYGO1; NCOA3 and RBM39; NOTCH2 and DPOLA; NOTCH2 and ENL; NOTCH2 and FOXO3; NOTCH2 and HERC2; NOTCH2
and HSH2D; N0TCH2 and NC0A2; N0TCH2 and NC0A3; N0TCH2 and N0TCH2; N0TCH2 and PSA1; N0TCH2 and PYG01; NOTCH2 and RBM39; PSA1 and DPOLA; PSA1 and ENL; PSA1 and FOXO3; PSA1 and HERC2; PSA1 and HSH2D; PSA1 and NCOA2; PSA1 and NCOA3; PSA1 and NOTCH2; PSA1 and PSA1; PSA1 and PYGO1; PSA1 and RBM39; PYGO1 and DPOLA; PYGO1 and ENL; PYGO1 and FOXO3; PYGO1 and HERC2; PYGO1 and HSH2D; PYGO1 and NCOA2; PYGO1 and NCOA3; PYGO1 and NOTCH2; PYGO1 and PSA1; PYGO1 and PYGO1; PYGO1 and RBM39; RBM39 and DPOLA; RBM39 and ENL; RBM39 and FOXO3; RBM39 and HERC2; RBM39 and HSH2D; RBM39 and NCOA2; RBM39 and NCOA3; RBM39 and NOTCH2; RBM39 and PSA1; RBM39 and PYGO1; or RBM39 and RBM39, respectively.
[0295] In some embodiments, the multipartite activator is a tripartite activator comprising a first transcriptional activation domain, a second transcriptional activation domain, and a third transcriptional activation domain. In some aspects, the first transcriptional activation domain, the second transcriptional activation domain, and the third transcriptional activation domain each independently comprises a domain of a protein selected from among DPOLA, ENL, FOXO3, HSH2D, NCOA2, NCOA3, PSA1, PYGO1, RBM39, HERC2, and NOTCH2. In some embodiments, the first and second transcriptional domains are the first and second transcriptional domains from any of the bipartite activators described above, and the third transcriptional domain independently comprises a domain of a protein selected from among DPOLA, ENL, FOXO3, HSH2D, NCOA2, NCOA3, PSA1, PYGO1, RBM39, HERC2, and NOTCH2.
[0296] In some embodiments, the multipartite activator is a tripartite activator comprising a first transcriptional activation domain, a second transcriptional activation domain, and a third transcriptional activation domain, each independently comprising a domain of a protein selected from among DPOLA, ENL, FOXO3, HSH2D, NCOA2, NCOA3, PSA1, PYGO1, RBM39, HERC2, and NOTCH2. In some embodiments, the first, second, and third transcriptional activation domains, respectively, are from NCOA3, NCOA3, and NCOA3; NCOA3, NCOA3, and ENL; NCOA3, NCOA3, and FOXO3; NCOA3, NCOA3, and PYGO1; NCOA3, NCOA3, and HSH2D; NCOA3, NCOA3, and NCOA2; NCOA3, NCOA3, and NOTCH2; NCOA3, ENL, and NCOA3; NCOA3, ENL, and ENL; NCOA3, ENL, and FOXO3; NCOA3, ENL, and PYGO1; NCOA3, ENL, and HSH2D; NCOA3, ENL, and NCOA2; NCOA3, ENL, and NOTCH2; NCOA3, FOXO3, and NCOA3; NCOA3, FOXO3, and ENL; NCOA3, FOXO3, and FOXO3; NCOA3, FOXO3, and PYGO1; NCOA3, FOXO3, and HSH2D; NCOA3, FOXO3, and NCOA2; NCOA3, FOXO3, and NOTCH2; NCOA3, PYGO1, and NCOA3; NCOA3, PYGO1, and ENL; NCOA3, PYGO1, and FOXO3; NCOA3, PYGO1, and PYGO1; NCOA3, PYGO1, and HSH2D; NCOA3, PYGO1, and NCOA2; NCOA3, PYGO1, and NOTCH2; NCOA3, HSH2D, and NCOA3; NCOA3, HSH2D, and ENL; NCOA3, HSH2D, and FOXO3; NCOA3, HSH2D, and PYGO1; NCOA3, HSH2D, and HSH2D; NCOA3, HSH2D, and NCOA2; NCOA3, HSH2D, and NOTCH2; NCOA3, NCOA2, and NCOA3; NCOA3, NCOA2, and ENL; NCOA3, NCOA2, and FOXO3; NCOA3, NCOA2, and PYGO1; NCOA3, NCOA2, and HSH2D; NCOA3, NCOA2, and NCOA2; NCOA3,
NCOA2, and NOTCH2; NCOA3, NOTCH2, and NCOA3; NCOA3, NOTCH2, and ENL; NCOA3, NOTCH2, and FOXO3; NCOA3, NOTCH2, and PYGO1; NCOA3, NOTCH2, and HSH2D; NCOA3, NOTCH2, and NCOA2; NCOA3, NOTCH2, and NOTCH2; ENL, NCOA3, and NCOA3; ENL, NCOA3, and ENL; ENL, NCOA3, and FOXO3; ENL, NCOA3, and PYGO1; ENL, NCOA3, and HSH2D; ENL, NCOA3, and NCOA2; ENL, NCOA3, and NOTCH2; ENL, ENL, and NCOA3; ENL, ENL, and ENL; ENL, ENL, and FOXO3; ENL, ENL, and PYGO1; ENL, ENL, and HSH2D; ENL, ENL, and NCOA2; ENL, ENL, and NOTCH2; ENL, FOXO3, and NCOA3; ENL, FOXO3, and ENL; ENL, FOXO3, and FOXO3; ENL, FOXO3, and PYGO1; ENL, FOXO3, and HSH2D; ENL, FOXO3, and NCOA2; ENL, FOXO3, and NOTCH2; ENL, PYGO1, and NCOA3; ENL, PYGO1, and ENL; ENL, PYGO1, and FOXO3; ENL, PYGO1, and PYGO1; ENL, PYGO1, and HSH2D; ENL, PYGO1, and NCOA2; ENL, PYGO1, and NOTCH2; ENL, HSH2D, and NCOA3; ENL, HSH2D, and ENL; ENL, HSH2D, and FOXO3; ENL, HSH2D, and PYGO1; ENL, HSH2D, and HSH2D; ENL, HSH2D, and NCOA2; ENL, HSH2D, and NOTCH2; ENL, NCOA2, and NCOA3; ENL, NCOA2, and ENL; ENL, NCOA2, and FOXO3; ENL, NCOA2, and PYGO1; ENL, NCOA2, and HSH2D; ENL, NCOA2, and NCOA2; ENL, NCOA2, and NOTCH2; ENL, NOTCH2, and NCOA3; ENL, NOTCH2, and ENL; ENL, NOTCH2, and FOXO3; ENL, NOTCH2, and PYGO1; ENL, NOTCH2, and HSH2D; ENL, NOTCH2, and NCOA2; ENL, NOTCH2, and NOTCH2; FOXO3, NCOA3, and NCOA3; FOXO3, NCOA3, and ENL; FOXO3, NCOA3, and FOXO3; FOXO3, NCOA3, and PYGO1; FOXO3, NCOA3, and HSH2D; FOXO3, NCOA3, and NCOA2; FOXO3, NCOA3, and NOTCH2; FOXO3, ENL, and NCOA3; FOXO3, ENL, and ENL; FOXO3, ENL, and FOXO3; FOXO3, ENL, and PYGO1; FOXO3, ENL, and HSH2D;
FOXO3, ENL, and NCOA2; FOXO3, ENL, and NOTCH2; FOXO3, FOXO3, and NCOA3; FOXO3, FOXO3, and ENL; FOXO3, FOXO3, and FOXO3; FOXO3, FOXO3, and PYGO1; FOXO3, FOXO3, and HSH2D; FOXO3, FOXO3, and NCOA2; FOXO3, FOXO3, and NOTCH2; FOXO3, PYGO1, and NCOA3; FOXO3, PYGO1, and ENL; FOXO3, PYGO1, and FOXO3; FOXO3, PYGO1, and PYGO1; FOXO3, PYGO1, and HSH2D; FOXO3, PYGO1, and NCOA2; FOXO3, PYGO1, and NOTCH2; FOXO3, HSH2D, and NCOA3; FOXO3, HSH2D, and ENL; FOXO3, HSH2D, and FOXO3; FOXO3, HSH2D, and PYGO1; FOXO3, HSH2D, and HSH2D; FOXO3, HSH2D, and NCOA2; FOXO3, HSH2D, and NOTCH2; FOXO3, NCOA2, and NCOA3; FOXO3, NCOA2, and ENL; FOXO3, NCOA2, and FOXO3; FOXO3, NCOA2, and PYGO1; FOXO3, NCOA2, and HSH2D; FOXO3, NCOA2, and NCOA2; FOXO3, NCOA2, and NOTCH2; FOXO3, NOTCH2, and NCOA3; FOXO3, NOTCH2, and ENL; FOXO3, NOTCH2, and FOXO3; FOXO3, NOTCH2, and PYGO1; FOXO3, NOTCH2, and HSH2D; FOXO3, NOTCH2, and NCOA2; FOXO3, NOTCH2, and NOTCH2; PYGO1, NCOA3, and NCOA3; PYGO1, NCOA3, and ENL; PYGO1, NCOA3, and FOXO3; PYGO1, NCOA3, and PYGO1; PYGO1, NCOA3, and HSH2D; PYGO1, NCOA3, and NCOA2; PYGO1, NCOA3, and NOTCH2; PYGO1, ENL, and NCOA3; PYGO1, ENL, and ENL; PYGO1, ENL, and FOXO3; PYGO1, ENL, and PYGO1; PYGO1, ENL, and HSH2D; PYGO1, ENL, and NCOA2; PYGO1, ENL, and NOTCH2;
PYG01, FOXO3, and NCOA3; PYGO1, FOXO3, and ENL; PYGO1, FOXO3, and FOXO3; PYGO1,
F0X03, and PYG01; PYGO1, FOXO3, and HSH2D; PYGO1, FOXO3, and NCOA2; PYGO1, FOXO3, and NOTCH2; PYGO1, PYGO1, and NCOA3; PYGO1, PYGO1, and ENL; PYGO1, PYGO1, and FOXO3; PYGO1, PYGO1, and PYGO1; PYGO1, PYGO1, and HSH2D; PYGO1, PYGO1, and NCOA2; PYGO1, PYGO1, and NOTCH2; PYGO1, HSH2D, and NCOA3; PYGO1, HSH2D, and ENL; PYGO1, HSH2D, and FOXO3; PYGO1, HSH2D, and PYGO1; PYGO1, HSH2D, and HSH2D; PYGO1, HSH2D, and NCOA2; PYGO1, HSH2D, and NOTCH2; PYGO1, NCOA2, and NCOA3; PYGO1, NCOA2, and ENL; PYGO1, NCOA2, and FOXO3; PYGO1, NCOA2, and PYGO1; PYGO1, NCOA2, and HSH2D; PYGO1, NCOA2, and NCOA2; PYGO1, NCOA2, and NOTCH2; PYGO1, NOTCH2, and NCOA3; PYGO1, NOTCH2, and ENL; PYGO1, NOTCH2, and FOXO3; PYGO1, NOTCH2, and PYGO1; PYGO1, NOTCH2, and HSH2D; PYGO1, NOTCH2, and NCOA2; PYGO1, NOTCH2, and NOTCH2; HSH2D, NCOA3, and NCOA3; HSH2D, NCOA3, and ENL; HSH2D, NCOA3, and FOXO3; HSH2D, NCOA3, and PYGO1; HSH2D, NCOA3, and HSH2D; HSH2D, NCOA3, and NCOA2; HSH2D, NCOA3, and NOTCH2; HSH2D, ENL, and NCOA3; HSH2D, ENL, and ENL; HSH2D, ENL, and FOXO3; HSH2D, ENL, and PYGO1; HSH2D, ENL, and HSH2D; HSH2D, ENL, and NCOA2; HSH2D, ENL, and NOTCH2; HSH2D, FOXO3, and NCOA3; HSH2D, FOXO3, and ENL; HSH2D, FOXO3, and FOXO3; HSH2D, FOXO3, and PYGO1; HSH2D, FOXO3, and HSH2D; HSH2D, FOXO3, and NCOA2; HSH2D, FOXO3, and NOTCH2; HSH2D, PYGO1, and NCOA3; HSH2D, PYGO1, and ENL; HSH2D, PYGO1, and FOXO3; HSH2D, PYGO1, and PYGO1; HSH2D, PYGO1, and HSH2D; HSH2D, PYGO1, and NCOA2; HSH2D, PYGO1, and NOTCH2; HSH2D, HSH2D, and NCOA3; HSH2D, HSH2D, and ENL; HSH2D, HSH2D, and FOXO3; HSH2D, HSH2D, and PYGO1; HSH2D, HSH2D, and HSH2D;
HSH2D, HSH2D, and NCOA2; HSH2D, HSH2D, and NOTCH2; HSH2D, NCOA2, and NCOA3;
HSH2D, NCOA2, and ENL; HSH2D, NCOA2, and FOXO3; HSH2D, NCOA2, and PYGO1; HSH2D, NCOA2, and HSH2D; HSH2D, NCOA2, and NCOA2; HSH2D, NCOA2, and NOTCH2; HSH2D, NOTCH2, and NCOA3; HSH2D, NOTCH2, and ENL; HSH2D, NOTCH2, and FOXO3; HSH2D, NOTCH2, and PYGO1; HSH2D, NOTCH2, and HSH2D; HSH2D, NOTCH2, and NCOA2; HSH2D, NOTCH2, and NOTCH2; NCOA2, NCOA3, and NCOA3; NCOA2, NCOA3, and ENL; NCOA2, NCOA3, and FOXO3; NCOA2, NCOA3, and PYGO1; NCOA2, NCOA3, and HSH2D; NCOA2, NCOA3, and NCOA2; NCOA2, NCOA3, and NOTCH2; NCOA2, ENL, and NCOA3; NCOA2, ENL, and ENL; NCOA2, ENL, and FOXO3; NCOA2, ENL, and PYGO1; NCOA2, ENL, and HSH2D;
NCOA2, ENL, and NCOA2; NCOA2, ENL, and NOTCH2; NCOA2, FOXO3, and NCOA3; NCOA2, FOXO3, and ENL; NCOA2, FOXO3, and FOXO3; NCOA2, FOXO3, and PYGO1; NCOA2, FOXO3, and HSH2D; NCOA2, FOXO3, and NCOA2; NCOA2, FOXO3, and NOTCH2; NCOA2, PYGO1, and NCOA3; NCOA2, PYGO1, and ENL; NCOA2, PYGO1, and FOXO3; NCOA2, PYGO1, and PYGO1; NCOA2, PYGO1, and HSH2D; NCOA2, PYGO1, and NCOA2; NCOA2, PYGO1, and NOTCH2; NCOA2, HSH2D, and NCOA3; NCOA2, HSH2D, and ENL; NCOA2, HSH2D, and FOXO3; NCOA2, HSH2D, and PYGO1; NCOA2, HSH2D, and HSH2D; NCOA2, HSH2D, and NCOA2; NCOA2, HSH2D, and NOTCH2; NCOA2, NCOA2, and NCOA3; NCOA2, NCOA2, and ENL; NCOA2,
NC0A2, and F0X03; NC0A2, NC0A2, and PYG01; NCOA2, NCOA2, and HSH2D; NCOA2, NCOA2, and NCOA2; NCOA2, NCOA2, and NOTCH2; NCOA2, NOTCH2, and NCOA3; NCOA2, NOTCH2, and ENL; NCOA2, NOTCH2, and FOXO3; NCOA2, NOTCH2, and PYGO1; NCOA2, NOTCH2, and HSH2D; NCOA2, NOTCH2, and NCOA2; NCOA2, NOTCH2, and NOTCH2; NOTCH2, NCOA3, and NCOA3; NOTCH2, NCOA3, and ENL; NOTCH2, NCOA3, and FOXO3; NOTCH2, NCOA3, and PYGO1; NOTCH2, NCOA3, and HSH2D; NOTCH2, NCOA3, and NCOA2; NOTCH2, NCOA3, and NOTCH2; NOTCH2, ENL, and NCOA3; NOTCH2, ENL, and ENL; NOTCH2, ENL, and FOXO3; NOTCH2, ENL, and PYGO1; NOTCH2, ENL, and HSH2D; NOTCH2, ENL, and NCOA2; NOTCH2, ENL, and NOTCH2; NOTCH2, FOXO3, and NCOA3; NOTCH2, FOXO3, and ENL; NOTCH2, FOXO3, and FOXO3; NOTCH2, FOXO3, and PYGO1; NOTCH2, FOXO3, and HSH2D; NOTCH2, FOXO3, and NCOA2; NOTCH2, FOXO3, and NOTCH2; NOTCH2, PYGO1, and NCOA3; NOTCH2, PYGO1, and ENL; NOTCH2, PYGO1, and FOXO3; NOTCH2, PYGO1, and PYGO1; NOTCH2, PYGO1, and HSH2D; NOTCH2, PYGO1, and NCOA2; NOTCH2, PYGO1, and NOTCH2; NOTCH2, HSH2D, and NCOA3; NOTCH2, HSH2D, and ENL; NOTCH2, HSH2D, and FOXO3; NOTCH2, HSH2D, and PYGO1; NOTCH2, HSH2D, and HSH2D; NOTCH2, HSH2D, and NCOA2; NOTCH2, HSH2D, and NOTCH2; NOTCH2, NCOA2, and NCOA3; NOTCH2, NCOA2, and ENL; NOTCH2, NCOA2, and FOXO3; NOTCH2, NCOA2, and PYGO1; NOTCH2, NCOA2, and HSH2D; NOTCH2, NCOA2, and NCOA2; NOTCH2, NCOA2, and NOTCH2; NOTCH2, NOTCH2, and NCOA3; NOTCH2, NOTCH2, and ENL; NOTCH2, NOTCH2, and FOXO3; NOTCH2, NOTCH2, and PYGO1; NOTCH2, NOTCH2, and HSH2D; NOTCH2, NOTCH2, and NCOA2; or NOTCH2, NOTCH2, and NOTCH2, respectively.
[0297] In some embodiments, the first, second, and third transcriptional activation domains, respectively, are from PYGO1, FOXO3, and NCOA3, respectively. In some embodiments, the first, second, and third transcriptional activation domains, respectively, are from NOTCH2, FOXO3, and NCOA3, respectively. In some embodiments, the first, second, and third transcriptional activation domains, respectively, are from NCOA3, FOXO3, and NCOA3, respectively. In some embodiments, the first, second, and third transcriptional activation domains, respectively, are from HSH2D, FOXO3, and NCOA3, respectively. In some embodiments, the first, second, and third transcriptional activation domains, respectively, are from FOXO3, FOXO3, and NCOA3, respectively. In some embodiments, the first, second, and third transcriptional activation domains, respectively, are from NCOA2, FOXO3, and NCOA3, respectively. In some embodiments, the first, second, and third transcriptional activation domains, respectively, are from ENL, FOXO3, and NCOA3, respectively.
[0298] In some embodiments, the first, second, and third transcriptional activation domains, respectively, are from FOXO3, FOXO3, and NCOA3, respectively. In some embodiments, the first, second, and third transcriptional activation domains, respectively, are from NCOA3, FOXO3, and FOXO3, respectively. In some embodiments, the first, second, and third transcriptional activation domains, respectively, are from NCOA3, FOXO3, and NCOA3, respectively.
[0299] In some embodiments, any of the provided multipartite effector proteins, fusion proteins, and/or DNA targeting systems, such as a multipartite activator, comprises a combination of transcriptional activation domains, such as a combination of two or more of any of transcriptional activation domains shown in Table 3. In some embodiments, any of the provided multipartite effector proteins, fusion proteins, and/or DNA targeting systems, such as a multipartite activator, comprises a combination of two or more of any one of the SEQ ID NOS: set forth in Table 3, or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any one of the SEQ ID NOS: set forth in Table 3, or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof. In some embodiments, any of the provided multipartite effector proteins, fusion proteins, and/or DNA targeting systems, such as a multipartite activator, comprises a combination of two or more of any one of the SEQ ID NOS: set forth in Table 3, or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof. In some embodiments, any of the provided multipartite effector proteins, fusion proteins, and/or DNA targeting systems, such as a multipartite activator, comprises two or more of any one of the SEQ ID NOS: set forth in Table 3.
[0300] In some embodiments, any of the provided multipartite effector proteins, fusion proteins, and/or DNA targeting systems, such as a multipartite activator, comprises a combination of transcriptional activation domains, such as a combination of three or more of any of transcriptional activation domains shown in Table 3. In some embodiments, any of the provided multipartite effector proteins, fusion proteins, and/or DNA targeting systems, such as a multipartite activator, comprises a combination of three or more of any one of the SEQ ID NOS: set forth in Table 3, or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any one of the SEQ ID NOS: set forth in Table 3, or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof. In some embodiments, any of the provided multipartite effector proteins, fusion proteins, and/or DNA targeting systems, such as a multipartite activator, comprises a combination of three or more of any one of the SEQ ID NOS: set forth in Table 3, or a domain or a portion thereof, such as a contiguous portion thereof of at least 10, 15, 20, 22, 25, 30, 35, 37, 40, 42, 45, 47, 49, 50, 55, 57, 60, 61, 62, 65, 70, 72, 75, 76, or 80 amino acids, such as at least 20 amino acids, or a variant thereof. In some
embodiments, any of the provided multipartite effector proteins, fusion proteins, and/or DNA targeting systems, such as a multipartite activator, comprises three or more of any one of the SEQ ID NOS: set forth in Table 3.
[0301] In some embodiments, the multipartite activator comprises the any one of the SEQ ID NOS: set forth in Table 4, or a domain, portion, or variant thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any one of the SEQ ID NOS: set forth in Table 4. In some embodiments, the multipartite activator is or comprises any one of the SEQ ID NOS: set forth in Table 4. In some embodiments, the multipartite activator comprises a combination of transcriptional activation domains, such as any of the combinations of transcriptional activation domains shown in Table 4.
[0302| In some embodiments, the multipartite activator comprises any one of SEQ ID NOS:397- 418, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any one of SEQ ID NOS:397-418. In some embodiments, the multipartite activator is set forth in any one of SEQ ID NOS:397-418, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity any one of SEQ ID NOS:397-418, or a partially or fully functional fragment thereof, a domain thereof, or a portion thereof,
such as a contiguous portion thereof of at least 10, 15, 20, 25, 30, 35, 40, 45, 50, 55, 60, 65, 70, 75, or 80 amino acids, or a variant thereof. In some embodiments, the multipartite activator is set forth in any one of SEQ ID NOS:397-418.
[0303] In some embodiments, the multipartite activator comprises domains from NCOA3, FOXO3, and NCOA3, respectively. In some embodiments, the multipartite activator comprises SEQ ID NO:413, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO:413. In some embodiments, the multipartite activator is set forth in SEQ ID NO:413.
[0304] In some embodiments, the multipartite activator comprises domains from FOXO3, FOXO3, and NCOA3, respectively. In some embodiments, the multipartite activator comprises SEQ ID NO:415, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO:415. In some embodiments, the multipartite activator is set forth in SEQ ID NO:415.
[0305] In some embodiments, the multipartite activator comprises domains from NCOA3, FOXO3, and FOXO3, respectively. In some embodiments, the multipartite activator comprises SEQ ID NO:418, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO:418. In some embodiments, the multipartite activator is set forth in SEQ ID NO:418.
C. eZFP Fusion Proteins
[0306] In some aspects, the DNA-targeting system comprises an eZFP fusion protein, such as any provided herein. In some aspects, provided are DNA-targeting systems comprising eZFP fusion proteins.
[0307] In some aspects, provided herein are fusion proteins, such as fusion proteins targeting, or capable of being targeted to, a FXN locus. In some embodiments, the fusion proteins comprise an eZFP or a dCas protein. In some embodiments, the fusion protein comprises an eZFP (i.e. the fusion protein is an eZFP fusion protein), such as any of the eZFPs described herein, for example in Section I. In some embodiments, the fusion protein further comprises an epigenetic effector domain, such as any of the effector domains for transcriptional activation described herein, for example in Section II.B. In some embodiments, the fusion protein comprises at least one epigenetic effector domain that increases transcription of the FXN locus. In some embodiments, the fusion protein comprises more than one effector domain. In some embodiments, the fusion protein comprises one or more additional elements, such as a nuclear localization signal (NLS) or linker, such as any of the NLSs or linkers described herein. In some aspects, the elements of the fusion protein may be arranged in any suitable order within the fusion protein, such as an order from N-terminus to C-terminus. In some aspects, the fusion proteins comprising eZFPs provided herein may facilitate increased FXN expression, for example in connection with compositions and methods for treating a disease or disorder associated with FXN expression, such as Friedreich’s ataxia (FA).
[0308] In some aspects, the fusion protein comprising the eZFP binds to, or is capable of binding to, (i.e. targets), any of the target sites provided herein. In some embodiments, the fusion protein binds to the target site. In some embodiments, the fusion protein comprising the eZFP binds to the target site that the eZFP binds to in the absence of the other elements of the fusion protein. Thus, in some embodiments, the eZFP of the fusion protein facilitates target-specific binding of the fusion protein. In some aspects, the fusion protein targets to the target site targeted by any of the eZFPs described herein, such as in Section I. In some embodiments, the fusion protein targets a target site in Table 1. In some embodiments, the fusion protein targets a target site in Table 2A and Table 2B. In some embodiments, the fusion protein targets a target site comprising the nucleotide sequence set forth in any one of SEQ ID NOS:272, 277, 280, 281, 283, 290, or 299, or a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the fusion protein targets a target site comprising the nucleotide sequence set forth in any one of SEQ ID NOS:583-600, or a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the fusion protein targets a target site comprising the nucleotide sequence set forth in any one of SEQ ID NOS:272, 277, 280, 281, 283, 290, or 299. In some embodiments, the fusion protein targets a target site comprising the nucleotide sequence set forth in any one of SEQ ID NOS:583-600. In some embodiments, the fusion protein targets a target site comprising the nucleotide sequence set forth in SEQ ID NO: 299. In some embodiments, the fusion protein targets a target site comprising the nucleotide sequence set forth in SEQ ID NO: 587. In some embodiments, the fusion protein targets a target site comprising the nucleotide sequence set forth in SEQ ID NO: 589. In some embodiments, the fusion protein targets a target site comprising the nucleotide sequence set forth in SEQ ID NO: 591. In some embodiments, the target site is double-stranded DNA.
[0309] In some aspects, the fusion protein comprises any of the eZFPs set forth in Table 2A and Table 2B. In some aspects, the fusion protein comprises an eZFP comprising the recognition regions Fl- F6 set forth for any of the eZFPs set forth in Table 2A and Table 2B (comprising SEQ ID NOS:341-346, respectively; SEQ ID NOS:347-352, respectively; SEQ ID NOS:353-358, respectively; SEQ ID NOS:359-364, respectively; SEQ ID NOS:365-370, respectively; SEQ ID NOS:371-376, respectively; SEQ ID NOS:377-382, respectively; SEQ ID NOS:475-480, respectively; SEQ ID NOS:481-486, respectively; SEQ ID NOS:487-492, respectively; SEQ ID NOS:493-498, respectively; SEQ ID NOS:499-504, respectively; SEQ ID NOS:505-510, respectively; SEQ ID NOS:511-516, respectively; SEQ ID NOS:517-522, respectively; SEQ ID NOS:523-528, respectively; SEQ ID NOS:529-534, respectively; SEQ ID NOS: 535-540, respectively; SEQ ID NOS:541-546, respectively; SEQ ID NOS:547-552, respectively; SEQ ID NOS:553-558, respectively; SEQ ID NOS:559-564, respectively; SEQ ID NOS:565-570, respectively; SEQ ID NOS: 571-576, respectively; or SEQ ID NOS: 577-582, respectively). In some embodiments, the fusion protein comprises the amino acid sequence set forth in any one of SEQ ID NOS:301-307 and 439-456, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some
embodiments, the fusion protein comprises the amino acid sequence set forth in any one of SEQ ID NOS:301-307 and 439-456. In some embodiments, the eZFP of the fusion protein is encoded by the nucleotide sequence set forth in any one of SEQ ID NOS:308-314 and 457-474, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP of the fusion protein is encoded by the nucleotide sequence set forth in any one of SEQ ID NOS:308-314 and 457-474.
[0310] In some aspects, provided herein is a fusion protein comprising an eZFP, such as any of the eZPFs described herein, for example in Table 2A or Table 2B. In some embodiments, the fusion protein targets a target site comprising the nucleotide sequence set forth in any one of SEQ ID NOS: 272, 277, 280, 281, 283, 290, or 299, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the fusion protein targets a target site comprising the nucleotide sequence set forth in any one of SEQ ID NOS: 583-600, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the fusion protein targets a target site comprising the nucleotide sequence set forth in any one of SEQ ID NOS:272, 277, 280, 281, 283, 290, or 299. In some embodiments, the fusion protein targets a target site comprising the nucleotide sequence set forth in any one of SEQ ID NOS:583-600. In some embodiments, the target site is double-stranded DNA. In some embodiments, the eZFP of the fusion protein comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequences of the recognition regions F1-F6 comprise: SEQ ID NOS:341-346, respectively; SEQ ID NOS:347-352, respectively; SEQ ID NOS:353-358, respectively; SEQ ID NOS:359-364, respectively; SEQ ID NOS:365-370, respectively; SEQ ID NOS:371-376, respectively; or SEQ ID NOS:377-382, respectively. In some embodiments, the eZFP of the fusion protein comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequences of the recognition regions F1-F6 comprise: SEQ ID NOS:475-480, respectively; SEQ ID NOS:481-486, respectively; SEQ ID NOS:487-492, respectively; SEQ ID NOS:493-498, respectively; SEQ ID NOS:499-504, respectively; SEQ ID NOS:505-510, respectively; SEQ ID NOS:511-516, respectively; SEQ ID NOS:517-522, respectively; SEQ ID NOS:523-528, respectively; SEQ ID NOS:529-534, respectively; SEQ ID NOS: 535-540, respectively; SEQ ID NOS:541-546, respectively; SEQ ID NOS:547-552, respectively; SEQ ID NOS:553-558, respectively; SEQ ID NOS:559-564, respectively; SEQ ID NOS:565-570, respectively; SEQ ID NOS: 571-576, respectively; or SEQ ID NOS: 577-582, respectively. In some embodiments, the eZFP of the fusion protein comprises the amino acid sequence set forth in any one of SEQ ID NOS:301-307, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP of the fusion protein comprises the amino acid sequence set forth in any one of SEQ ID NOS:439-456, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some
embodiments, the eZFP of the fusion protein comprises the amino acid sequence set forth in any one of SEQ ID NOS:301-307. In some embodiments, the eZFP of the fusion protein comprises the amino acid sequence set forth in any one of SEQ ID NOS:439-456. In some embodiments, the eZFP of the fusion protein is encoded by the nucleotide sequence set forth in any one of SEQ ID NOS:308-314, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP of the fusion protein is encoded by the nucleotide sequence set forth in any one of SEQ ID NOS:457-474, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP of the fusion protein is encoded by the nucleotide sequence set forth in any one of SEQ ID NOS:308-314. In some embodiments, the eZFP of the fusion protein is encoded by the nucleotide sequence set forth in any one of SEQ ID NOS:457-474.
[0311] In some aspects, provided herein is a fusion protein comprising an eZFP, such as eZFP_A31 as described herein. In some embodiments, the fusion protein targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:299, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the fusion protein targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:299. In some embodiments, the target site is double-stranded DNA. In some embodiments, the eZFP of the fusion protein comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: LRHHLTR (SEQ ID NO:377); F2: QSAHLKA (SEQ ID NO:378); F3: LPQTLQR (SEQ ID NO:379); F4: QNATRTK (SEQ ID NO:380); F5: QSSHLTR (SEQ ID NO:381); F6: RSDHLSR (SEQ ID NO:382). In some embodiments, the eZFP of the fusion protein comprises the amino acid sequence set forth in SEQ ID NO:307, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP of the fusion protein comprises the amino acid sequence set forth in SEQ ID NO:307. In some embodiments, the eZFP of the fusion protein is encoded by the nucleotide sequence set forth in SEQ ID NO:314, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP of the fusion protein is encoded by the nucleotide sequence set forth in SEQ ID NO:314.
[0312] In some aspects, provided herein is a fusion protein comprising an eZFP, such as eZFP_A44 as described herein. In some embodiments, the fusion protein targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:587, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the fusion protein targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:587. In some embodiments, the target site is double-stranded DNA. In some embodiments, the eZFP of the fusion protein comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc
finger recognition region is as follows: Fl: QAGNRST (SEQ ID NO:499); F2: DRSALAR (SEQ ID NG:500); F3: RSDNLAR (SEQ ID NO:501); F4: WRGDRVK (SEQ ID NO:502); F5: YKHVLSD (SEQ ID NO:503); and F6: TSGSLTR (SEQ ID NO:504). In some embodiments, the eZFP of the fusion protein comprises the amino acid sequence set forth in SEQ ID NO: 443, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP of the fusion protein comprises the amino acid sequence set forth in SEQ ID NO:443. In some embodiments, the eZFP of the fusion protein is encoded by the nucleotide sequence set forth in SEQ ID NO:461, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP of the fusion protein is encoded by the nucleotide sequence set forth in SEQ ID NO:461.
[03133 In some aspects, provided herein is a fusion protein comprising an eZFP, such as eZFP_A46 as described herein. In some embodiments, the fusion protein targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:589, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the fusion protein targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:589. In some embodiments, the target site is double-stranded DNA. In some embodiments, the eZFP of the fusion protein comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: DRSTRTK (SEQ ID NO:511); F2: RSDYLAK (SEQ ID NO:512); F3: LRHHLTR (SEQ ID NO:513); F4: QSAHLKA (SEQ ID NO:514); F5: LPQTLQR (SEQ ID NO:515); and F6: QNATRTK (SEQ ID NO:516). In some embodiments, the eZFP of the fusion protein comprises the amino acid sequence set forth in SEQ ID NO: 445, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP of the fusion protein comprises the amino acid sequence set forth in SEQ ID NO:445. In some embodiments, the eZFP of the fusion protein is encoded by the nucleotide sequence set forth in SEQ ID NO:463, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP of the fusion protein is encoded by the nucleotide sequence set forth in SEQ ID NO:463.
[0314] In some aspects, provided herein is a fusion protein comprising an eZFP, such as eZFP_A48 as described herein. In some embodiments, the fusion protein targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:591, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing. In some embodiments, the fusion protein targets a target site comprising the nucleotide sequence set forth in SEQ ID NO:591. In some embodiments, the target site is double-stranded DNA. In some embodiments, the eZFP of the fusion protein comprises six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, each comprising a corresponding zinc finger recognition region Fl through F6, and the amino acid sequence of each zinc finger recognition region is as follows: Fl: RNDALTE (SEQ ID NO:523); F2: RKDNLKN (SEQ ID
NO:524); F3: TSGELVR (SEQ ID NO:525); F4: HRTTLTN (SEQ ID NO:526); F5: TTGNLTV (SEQ ID NO:527); and F6: RTDTLRD (SEQ ID NO:528). In some embodiments, the eZFP of the fusion protein comprises the amino acid sequence set forth in SEQ ID NO: 447, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP of the fusion protein comprises the amino acid sequence set forth in SEQ ID NO:447. In some embodiments, the eZFP of the fusion protein is encoded by the nucleotide sequence set forth in SEQ ID NO:465, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP of the fusion protein is encoded by the nucleotide sequence set forth in SEQ ID NO:465. In some aspects, the fusion protein further comprises one or more nuclear localization signal (NLS), such as any suitable NLS, for example any NLS described herein. In some aspects, an NLS may promote nuclear localization of the fusion protein.
[0315] In some embodiments, the NLS comprises the amino acid sequence set forth in any one of SEQ ID NOS:85 and 159-173, or an amino acid sequence having at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the NLS comprises the amino acid sequence set forth in any one of SEQ ID NOS:85 and 159-173.
[0316] In some embodiments, the NLS is an SV40 NLS. In some embodiments, the SV40 NLS comprises the amino acid sequence set forth in SEQ ID NO: 159.
[0317] In some embodiments, the NLS is a c-myc NLS. In some embodiments, the c-myc NLS comprises the amino acid sequence set forth in SEQ ID NO: 160.
[0318] In some embodiments, a fusion protein described herein comprises one or more nuclear localization sequences (NLSs), such as about or more than about 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, or more NLSs. When more than one NLS is present, each may be selected independently of the others, such that a single NLS may be present in more than one copy and/or in combination with one or more other NLSs present in one or more copies. Non-limiting examples of NLSs include an NLS sequence derived from: the NLS of the SV40 virus large T-antigen, having the sequence PKKKRKV (SEQ ID NO: 159); the NLS from nucleoplasmin (e.g. the nucleoplasmin bipartite NLS) having the sequence KRPAATKKAGQAKKKK (SEQ ID NO:85); the c-myc NLS having the sequence PAAKRVKLD (SEQ ID NO: 160) or RQRRNELKRSP (SEQ ID NO: 161); the hRNPAl M9 NLS having the sequence NQSSNFGPMKGGNFGGRSSGPYGGGGQYFAKPRNQGGY (SEQ ID NO: 162); the sequence RMRIZFKNKGKDTAELRRRRVEVSVELRKAKKDEQILKRRNV (SEQ ID NO: 163) of the IBB domain from importin-alpha; the sequences VSRKRPRP (SEQ ID NO: 164) and PPKKARED (SEQ ID NO: 165) of the myoma T protein; the sequence PQPKKKPL (SEQ ID NO: 166) of human p53; the sequence SALIKKKKKMAP (SEQ ID NO: 167) of mouse c-abl IV; the sequences DRLRR (SEQ ID NO: 168) and PKQKKRK (SEQ ID NO: 169) of the influenza virus NS1; the sequence RKLKKKIKKL (SEQ ID NO: 170) of the Hepatitis virus delta antigen; the sequence REKKKFLKRR (SEQ ID NO: 171) of the mouse Mxl protein; the sequence KRKGDEVDGVDEVAKKKSKK (SEQ ID NO: 172) of the
human poly(ADP-ribose) polymerase; and the sequence RKCLQAGMNLEARKTKK (SEQ ID NO: 173) of the steroid hormone receptors (human) glucocorticoid. In general, the one or more NLSs are of sufficient strength to drive accumulation of the fusion protein in a detectable amount in the nucleus of a eukaryotic cell. In general, strength of nuclear localization activity may derive from the number of NLSs in the fusion protein, the particular NLS(s) used, or a combination of these factors. Detection of accumulation in the nucleus may be performed by any suitable technique. For example, a detectable marker may be fused to the fusion protein, such that location within a cell may be visualized, such as in combination with a means for detecting the location of the nucleus (e.g. a stain specific for the nucleus such as DAPI). Cell nuclei may also be isolated from cells, the contents of which may then be analyzed by any suitable process for detecting protein, such as immunohistochemistry, Western blot, or enzyme activity assay. Accumulation in the nucleus may also be determined indirectly, such as by an assay for the effect of the fusion protein (e.g. an assay for altered gene expression activity in a cell transformed with the DNA-targeting system comprising the fusion protein), as compared to a control condition (e.g. an untransformed cell).
[0319] In some embodiments, the fusion protein further comprises one or more linker, such as any suitable linker, for example any linker described herein. In some embodiments, the one or more linkers may connect the eZFP and the at least one epigenetic effector domain. In some embodiments, each of the one or more linkers are in between any two of the components of the fusion protein, including the eZFP, any of the at least one effector domains, and the one or more NLS. In some embodiments, a linker may be of any length. In some embodiments, a linker may be designed to promote or restrict the mobility of components in the fusion protein.
[0320] In some embodiments, a linker in the fusion protein has the amino acid sequence set forth in any one of SEQ ID NOS:153-158, 174, 186, 188, and 219, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, a linker in the fusion protein has the amino acid sequence set forth in any one of SEQ ID NOS:153-158, 174, 186, 188, and 219.
[0321] A linker may comprise any amino acid sequence of about 2 to about 100, about 5 to about 80, about 10 to about 60, or about 20 to about 50 amino acids. A linker may comprise an amino acid sequence of at least about 2, 3, 4, 5, 10, 15, 20, 25, 30, 35, 40, 45, 50, 55, 60, 65, 70, 75, 80 or 85 amino acids. A linker may comprise an amino acid sequence of less than about 100, 90, 80, 70, 60, 50, or 40 amino acids. A linker may include sequential or tandem repeats of an amino acid sequence that is 2 to 20 amino acids in length. Linkers may be rich in amino acids glycine (G), serine (S), and/or alanine (A). Linkers may include, for example, a GS linker. An exemplary GS linker is represented by the sequence GGGGS (SEQ ID NO: 158). A linker may comprise repeats of a sequence, for example as represented by the formula (GGGGS)n, wherein n is an integer that represents the number of times the GGGGS sequence is repeated (e.g. between 1 and 10 times). The number of times a linker sequence is repeated, for example n in a GS linker, can be adjusted to optimize the linker length and achieve appropriate
separation of the functional domains. Other examples of linkers may include, for example, GGGGG (SEQ ID NO: 154), GGAGG (SEQ ID NO: 155), Gly/Ser rich linkers such as GGGGSSS (SEQ ID NO: 156), or Gly/Ala rich linkers such as GGGGAAA (SEQ ID NO: 157), or GSGSG (SEQ ID NO:219).
[0322] In some embodiments, the linker is an XTEN linker. In some aspects, an XTEN linker is a recombinant polypeptide (e.g., an unstructured recombinant peptide) lacking hydrophobic amino acid residues. Exemplary XTEN linkers are described in, for example, Schellenberger et al., Nature Biotechnology 27, 1186-1190 (2009) or WO 2021/247570. In some embodiments, an exemplary linker comprises a linker described in WO 2021/247570. In some aspects, the linker is or comprises the sequence set forth in SEQ ID NO: 186 or SEQ ID NO: 174, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing. In some embodiments, the linker comprises the sequence set forth in SEQ ID NO:186, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing. In some aspects, the linker comprises the sequence set forth in SEQ ID NO: 186, or a contiguous portion of SEQ ID NO: 186 of at least 5, 10, 15, 20, 25, 30, 35, 40, 45, 50, 55, 60, 65, 70 or 75 amino acids. In some aspects, the linker consists of the sequence set forth in SEQ ID NO: 186, or a contiguous portion of SEQ ID NO: 186 of at least 5, 10, 15, 20, 25, 30, 35, 40, 45, 50, 55, 60, 65, 70 or 75 amino acids. In some embodiments, the linker comprises the sequence set forth in SEQ ID NO: 186. In some embodiments, the linker consists of the sequence set forth in SEQ ID NO: 186. In some embodiments, the linker is encoded by a nucleotide sequence set forth in SEQ ID NO: 185. In some embodiments, the linker comprises the sequence set forth in SEQ ID NO:174, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing. In some aspects, the linker comprises the sequence set forth in SEQ ID NO: 174, or a contiguous portion of SEQ ID NO: 174 of at least 5, 10, or 15 amino acids. In some aspects, the linker consists of the sequence set forth in SEQ ID NO: 174, or a contiguous portion of SEQ ID NO: 174 of at least 5, 10, or 15 amino acids. In some embodiments, the linker comprises the sequence set forth in SEQ ID NO: 174. In some embodiments, the linker consists of the sequence set forth in SEQ ID NO: 174. Appropriate linkers may be selected or designed based rational criteria known in the art, for example as described in Chen et al. Adv. Drug Deliv. Rev. 65(10): 1357-1369 (2013). In some embodiments, a linker comprises the sequence set forth in SEQ ID NO:188, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
[0323] In some embodiments, the linker is a self-cleaving peptide, e.g., P2A. An exemplary P2A peptide sequence is set forth in SEQ ID NO: 131. In some embodiments, a P2A peptide sequence is encoded by the nucleotide sequence set forth in SEQ ID NO: 130.
[0324] In some embodiments, the fusion protein comprises at least one epigenetic effector domain that increases transcription of the FXN locus. In some embodiments, the at least one epigenetic effector domain is an effector domain for transcriptional activation and/or a multipartite activator, such as any of
the effector domains described herein, for example in Section III.
[0325] In some embodiments, the eZFP fusion protein comprises the one or more NLS, the eZFP, and the at least one epigenetic effector domain, in order from N-terminus to C-terminus, optionally wherein the NLS, eZFP, and at least one epigenetic effector domain are separated by one or more linker and/or additional NLS. In some embodiments, the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:320-340, 419-425, and 608-671, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing. In some embodiments, the fusion protein comprises the sequence set forth in any one of SEQ ID NOS:320-340, 419-425, and 608-671.
[0326] In some embodiments, the at least one epigenetic effector domain of the eZFP fusion protein comprises a VP64 domain. In some embodiments, the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:320-326 and 608-614, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing. In some embodiments, the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:320-326 and 608-641. In some embodiments, the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:320-326, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing. In some embodiments, the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:320-326. In some embodiments, the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:608-641, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing. In some embodiments, the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:608-641. In some embodiments, the eZFP fusion protein comprises the sequence set forth in SEQ ID NO:326. In some embodiments, the eZFP fusion protein comprises the sequence set forth in SEQ ID NO:340. In some embodiments, the eZFP fusion protein is encoded by the nucleotide sequence set forth in SEQ ID NO:426, or a nucleotide sequence having at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP fusion protein is encoded by the nucleotide sequence set forth in SEQ ID NO:426. In some embodiments, the eZFP fusion protein comprises the sequence set forth in SEQ ID NO:634.
[0327] In some embodiments, the at least one epigenetic effector domain of the eZFP fusion protein comprises a multipartite activator comprising, in the N-terminal to C-terminal direction, domains from FOXO3, FOXO3, and NCOA3, respectively, optionally separated by one or more linkers. In some embodiments, the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:334- 340 and 629-635, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing. In some embodiments, the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:334-340 and 629-635. In some embodiments, the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID
NOS:334-340, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing. In some embodiments, the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:334-340. In some embodiments, the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS: 629- 635, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing. In some embodiments, the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS: 629-635. In some embodiments, the eZFP fusion protein comprises the sequence set forth in SEQ ID NO:340. In some embodiments, the eZFP fusion protein is encoded by the nucleotide sequence set forth in SEQ ID NO:428, or a nucleotide sequence having at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP fusion protein is encoded by the nucleotide sequence set forth in SEQ ID NO:428. In some embodiments, the eZFP fusion protein comprises the sequence set forth in SEQ ID NO:635.
[0328] In some embodiments, the at least one epigenetic effector domain of the eZFP fusion protein comprises a multipartite activator comprising, in the N-terminal to C-terminal direction, domains from NCOA3, FOXO3, and FOXO3, respectively, optionally separated by one or more linkers. In some embodiments, the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:419- 425 and 615-621, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing. In some embodiments, the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:419-425 and 615-621. In some embodiments, the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:419-425, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing. In some embodiments, the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:419-425. In some embodiments, the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:615- 621, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing. In some embodiments, the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:615-621. In some embodiments, the eZFP fusion protein comprises the sequence set forth in SEQ ID NO: 425. In some embodiments, the eZFP fusion protein comprises the sequence set forth in SEQ ID NO: 621.
[0329] In some embodiments, the at least one epigenetic effector domain of the eZFP fusion protein comprises a multipartite activator comprising, in the N-terminal to C-terminal direction, domains from NCOA3, FOXO3, and NCOA3, respectively, optionally separated by one or more linker and/or NFS. In some embodiments, the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:327-333, 615-621, and 636-671, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing. In some embodiments, the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID
NOS:327-333, 615-621, and 636-671. In some embodiments, the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:327-333, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing. In some embodiments, the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:327-333. In some embodiments, the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:615-621, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing. In some embodiments, the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:615-621. In some embodiments, the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:636-653, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing. In some embodiments, the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS: 636-653. In some embodiments, the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:654-671, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing. In some embodiments, the eZFP fusion protein comprises the sequence set forth in any one of SEQ ID NOS:654- 671. In some embodiments, the eZFP fusion protein comprises the sequence set forth in SEQ ID NO: 333. In some embodiments, the eZFP fusion protein is encoded by the nucleotide sequence set forth in SEQ ID NO:427, or a nucleotide sequence having at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the eZFP fusion protein is encoded by the nucleotide sequence set forth in SEQ ID NO:427. In some embodiments, the eZFP fusion protein comprises the sequence set forth in SEQ ID NO:621. In some embodiments, the eZFP fusion protein comprises the sequence set forth in SEQ ID NO:640. In some embodiments, the eZFP fusion protein comprises the sequence set forth in SEQ ID NO:658. In some embodiments, the eZFP fusion protein comprises the sequence set forth in SEQ ID NO:642. In some embodiments, the eZFP fusion protein comprises the sequence set forth in SEQ ID NO:660. In some embodiments, the eZFP fusion protein comprises the sequence set forth in SEQ ID NO:644. In some embodiments, the eZFP fusion protein comprises the sequence set forth in SEQ ID NO:662.
[0330] In some aspects, provided herein are CRISPR/Cas-based DNA-targeting systems.
[0331] In some embodiments, the DNA-targeting systems comprise a DNA-targeting domain that binds to a target site in a regulatory DNA element of a FXN locus and comprises a Cas-guide RNA (gRNA) combination. In some embodiments, the Cas-gRNA combination includes a variant Cas protein that lacks nuclease activity or that is a deactivated Cas (dCas) protein. In some embodiments, the Cas- gRNA combination includes at least one gRNA comprising a gRNA spacer sequence that is capable of hybridizing to the target site or is complementary to the target site.
[0332] In some aspects, the CRISPR/Cas-based DNA targeting systems comprise an epigenetic effector domain and/or multipartite effector for transcriptional activation, such as any of the effector
domains or multipartite effectors described herein, such as in Section II.B. In some embodiments, the CRISPR/Cas-based DNA-tareting system comprises a fusion protein, such as a dCas fusion protein, comprising the effector domains and/or multipartite effectors, such as any described in Section II.B.
[0333] In some aspects, the DNA-targeting domain comprises a CRISPR-associated (Cas) protein or variant thereof, or comprises a protein that is derived from a Cas protein or variant thereof. In particular embodiments here, the Cas protein is nuclease-inactive (i.e. is a dCas protein).
[0334] In some aspects, provided herein are DNA-targeting systems based on CRISPR/Cas systems, i.e. CRISPR/Cas-based DNA-targeting systems, that are able to bind to a target site in a FXN gene or regulatory DNA element thereof. In some embodiments, the CRISPR/Cas DNA-targeting domain is nuclease inactive, such as includes a dCas (e.g. dCas9) so that the system binds to the target site in a target gene without mediating nucleic acid cleavage at the target site. The CRISPR/Cas-based DNA- targeting systems may be used to modulate expression of FXN in a cell. In some embodiments, the CRISPR/Cas-based DNA-targeting system can include any known Cas enzyme, such as a nucleaseinactive or dCas. In some embodiments, the CRISPR/Cas-based DNA-targeting system includes a fusion protein of a nuclease-inactive Cas protein or a variant thereof and an effector domain that increases transcription of a gene (e.g. a transcription activation domain), and at least one gRNA.
[0335] The CRISPR system (also known as CRISPR/Cas system, or CRISPR-Cas system) refers to a conserved microbial nuclease system, found in the genomes of bacteria and archaea, that provides a form of acquired immunity against invading phages and plasmids. Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR) refers to loci containing multiple repeating DNA elements that are separated by non-repeating DNA sequences called spacers. Spacers are short sequences of foreign DNA that are incorporated into the genome between CRISPR repeats, serving as a 'memory' of past exposures. Spacers encode the DNA-targeting portion of RNA molecules that confer specificity for nucleic acid cleavage by the CRISPR system. CRISPR loci contain or are adjacent to one or more CRISPR-associated (Cas) genes, which can act as RNA-guided nucleases for mediating the cleavage, as well as non-protein coding DNA elements that encode RNA molecules capable of programming the specificity of the CRISPR-mediated nucleic acid cleavage.
[0336] CRISPR/Cas systems, such as those with Cas9, have been engineered to allow efficient programming of Cas/RNA RNPs to target desired sequences in cells of interest, both for gene-editing and modulation of gene expression. The tracrRNA and crRNA have been engineered to form a single chimeric guide RNA molecule, commonly referred to as a guide RNA (gRNA), for example as described in WO 2013/176772, WO 2014/093661, WO 2014/093655, Jinek et al. Science 337(6096):816-21 (2012), or Cong et al. Science 339(6121): 819-23 (2013), and as described herein, for example, in Section II.B. The spacer sequence of the gRNA can be chosen by a user to target the Cas/gRNA RNP complex to a desired locus, e.g. a desired target site in the target gene, e.g., FXN. CRISPR/Cas systems may be multi-protein systems or single effector protein systems. Multi-protein, or Class 1, CRISPR systems include Type I, Type III, and Type IV systems. In some aspects, Class 2 systems include a single effector
molecule and include Type II, Type V, and Type VI. In some embodiments, the DNA targeting system comprises components of CRISPR/Cas systems, such as a Type I, Type II, Type III, Type IV, Type V, or Type VI CRISPR system. In some embodiments, the Cas protein is from a Class 1 CRISPR system (i.e. multiple Cas protein system), such as a Type I, Type III, or Type IV CRISPR system. In some embodiments, the Cas protein is from a Class 2 CRISPR system (i.e. single Cas protein system), such as a Type II, Type V, or Type VI CRISPR system.
[0337] In some embodiments, the Cas protein is derived from a Cas9 protein or variant thereof, for example as described in WO 2013/176772, WO 2014/152432, WO 2014/093661, WO 2014/093655, Jinek, M. et al. Science 337(6096):816-21 (2012), Mali, P. et al. Science 339(6121):823-6 (2013), Cong, L. et al. Science 339(6121):819-23 (2013), Perez-Pinera, P. et al. Nat. Methods 10, 973-976 (2013), or Mali, P. et al. Nat. Biotechnol. 31, 833-838 (2013). Various CRISPR/Cas systems and associated Cas proteins for use in gene editing and regulation have been described, for example in Moon et al. Exp. Mol. Med. 51, 1-11 (2019), Zhang, F. Q. Rev. Biophys. 52, E6 (2019), and Makarova et al. Methods Mol. Biol. 1311:47-75 (2015).
[0338] Type I CRISPR/Cas systems employ a large multisubunit ribonucleoprotein (RNP) complex called Cascade that recognizes double-stranded DNA (dsDNA) targets. After target recognition and verification, Cascade recruits the signature protein Cas3, a fused helicase-nuclease, to degrade DNA.
[0339] Iln some embodiments, the Cas protein is from a Type II CRISPR system. Exemplary Cas proteins of a Type II CRISPR system include Cas9. In some embodiments, the Cas protein is from a Cas9 protein or variant thereof, for example as described in WO 2013/176772, WO 2014/152432, WO 2014/093661, WO 2014/093655, Jinek. et al. Science 337(6096):816-21 (2012), Mali et al. Science 339(6121):823-6 (2013), Cong et al. Science 339(6121):819-23 (2013), Perez-Pinera et al. Nat. Methods 10, 973-976 (2013), or Mali et al. Nat. Biotechnol. 31, 833-838 (2013). In Type II CRISPR/Cas systems with the Cas protein Cas9, two RNA molecules and the Cas9 protein form a ribonucleoprotein (RNP) complex to direct Cas9 nuclease activity. The CRISPR RNA (crRNA) contains a spacer sequence that is complementary to a target nucleic acid sequence (target site), and that encodes the sequence specificity of the complex. The trans-activating crRNA (tracrRNA) base-pairs to a portion of the crRNA and forms a structure that complexes with the Cas9 protein, forming a Cas/RNA RNP complex. Cas9 mediates cleavage of target DNA if a correct protospacer-adjacent motif (PAM) is also present at the 3' end of the protospacer. For protospacer targeting, the sequence must be immediately followed by the protospacer- adjacent motif (PAM), a short sequence recognized by the Cas9 nuclease that is required for DNA cleavage.
[0340] Different Type II systems have differing PAM requirements. The 5. pyogenes CRISPR system may have the PAM sequence for this Cas9 (SpCas9) as 5'-NRG-3', where R is either A or G, and characterized the specificity of this system in human cells. A unique capability of the CRISPR/Cas9 system is the straightforward ability to simultaneously target multiple distinct genomic loci by coexpressing a single Cas9 protein with two or more sgRNAs. For example, the Streptococcus
pyogenes Type II system typically prefers to use an “NGG” sequence, where “N” can be any nucleotide, but also accepts other PAM sequences, such as “NAG” in engineered systems (Hsu et al., Nature Biotechnology (2013) doi:10.1038/nbt.2647). Similarly, the Cas9 derived from Neisseria meningitidis (NmCas9) normally has a native PAM of NNNNGATT (SEQ ID NO: 143), but has activity across a variety of PAMs, including a highly degenerate NNNNGNNN (SEQ ID NO:222) PAM (Esvelt et al. Nature Methods (2013) doi:10.1038/nmeth.2681). In another example, the Cas9 derived from Campylobacter jejuni typically uses 5'-NNNNACAC-3' (SEQ ID NO:226) or 5'-NNNNRYAC-3' (SEQ ID NO: 144) PAM sequences, where “N” can be any nucleotide, “R” can be either guanine (G) or adenine (A), and “Y” can be either cytosine (C) or thymine (T). In some aspects, the PAM sequences for spacer targeting depends on the type, ortholog, variant or species of the Cas protein.
[03411 In some embodiments, the Cas9 protein comprises a sequence from a Cas9 molecule of S. aureus. In some embodiments, the Cas9 protein comprises a sequence set forth in SEQ ID NO:73 or SEQ ID NO:126, or a variant thereof, such as an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO:73 or SEQ ID NO: 126. In some embodiments, the Cas9 protein comprises a sequence from a Cas9 molecule of S. pyogenes. In some embodiments, the Cas9 protein comprises a sequence set forth in SEQ ID NO:79 or SEQ ID NO: 127, or a variant thereof, such as an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO:79 or SEQ ID NO: 127.
[0342] In Type III systems, the RNP complex is multimeric with a helicoid structure similar to Cascade. In contrast to Type I CRISPR/Cas systems, the Type III RNP complex recognizes complementary RNA sequences instead of dsDNA. RNA recognition stimulates a nonspecific DNA cleavage activity of the exemplary Type III CaslO nuclease that is part of the RNP complex, such that DNA cleavage is achieved cotranscriptionally.
[0343] In some embodiments, the Cas protein is from a Type V CRISPR system. Exemplary Cas proteins of a Type V CRISPR system include Casl2a (also known as Cpfl), Casl2b (also known as C2cl), Casl2e (also known as CasX), Casl2k (also known as C2c5), Casl4a, and Casl4b. In some embodiments, the Cas protein is from a Casl2 protein (i.e. Cpfl) or variant thereof, for example as described in WO 2017/189308, WO2019/232069 and Zetsche et al. Cell. 163(3):759-71 (2015).
[0344] Exemplary Type V systems include those based on a Casl2 effector, and the C-terminus with only one RuvC endonuclease domain is the defining characteristic of the Type V systems. The RuvC nuclease domain cleaves dsDNA adjacent to protospacer adjacent motif (PAM) sequences and singlestranded DNA (ssDNA) nonspecifically. The Type V systems can be further divided into subtypes, each characterized by different signature proteins, PAM sequences, and properties. Non-limiting exemplary Cas proteins derived from Type V CRISPR systems include Casl2a (Cpfl), UnlCasl2fl, Casl2j (CasPhi, such as CasPhi-2), Casl2k, and CasMini. For example, Type V-A includes, for example, Casl2a, which uses “TTTV” (SEQ ID NO: 147) PAM sequence, where “V” is adenine (A), cytosine (C), or guanine (G). Type V-F is includes, for example, Casl2f, which can use “TTTR” (SEQ ID NO:228),
where “R” is G or A, or “TTTN” (SEQ ID NO:225), where “N” is any nucleotide. Type V-K is includes, for example, Casl2k, which uses “GGTT” (SEQ ID NO:227) PAM sequence.
[0345] In some embodiments, the Casl2a protein comprises a sequence from a Casl2a molecule of Acidaminococcus sp, such as an AsCasl2a set forth in SEQ ID NO: 191 or SEQ ID NO: 192, or a variant thereof, such as an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO: 191 or SEQ ID NO: 192.
[0346] Non-limiting examples of Cas proteins or Cas orthologs, such as Cas9 orthologs, from other bacterial strains include but are not limited to, Cas proteins identified in Acaryochloris marina MB IC 11017; Acetohalobium arabaticum DSM 5501; Acidaminococcus sp.; Acidithiobacillus caldus; Acidithiobacillus ferrooxidans ATCC 23270; Alicyclobacillus acidocaldarius LAA1; Alicyclobacillus acidocaldarius subsp. acidocaldarius DSM 446; Allochromatium vinosum DSM 180; Ammonifex degensii KC4; Anabaena variabilis ATCC 29413; Arthrospira maxima CS-328; Arthrospira platensis str. Paraca; Arthrospira sp. PCC 8005; Bacillus pseudomycoides DSM 12442; Bacillus selenitireducens MLS10; Burkholderiales bacterium 1_1_47; Caldicelulosiruptor becscii DSM 6725; Campylobacter jejuni; Candidatus Desulforudis audaxviator MP104C; Caldicellulosiruptor hydrothermalis 108; Clostridium phage c-st; Clostridium botulinum A3 str. Loch Maree; Clostridium botulinum Ba4 str. 657; Clostridium difficile QCD-63q42; Crocosphaera watsonii WH 8501; Cyanothece sp. ATCC 51142; Cyanothece sp. CCY0110; Cyanothece sp. PCC 7424; Cyanothece sp. PCC 7822; Exiguobacterium sibiricum 255-15; Finegoldia magna ATCC 29328; Ktedonobacter racemifer DSM 44963; Lactobacillus delbrueckii subsp. bulgaricus PB2003/044-T3-4; Lactobacillus salivarius ATCC 11741; Listeria innocua; Lyngbya sp. PCC 8106; Marinobacter sp. ELB17; Methanohalobium evestigatum Z-7303; Microcystis phage Ma-LMMOl; Microcystis aeruginosa NIES-843; Microscilla marina ATCC 23134; Microcoleus chthonoplastes PCC 7420; Neisseria meningitidis; Nitrosococcus halophilus Nc4; Nocardiopsis dassonvillei subsp. dassonvillei DSM 43111; Nodularia spumigena CCY9414; Nostoc sp. PCC 7120; Oscillatoria sp. PCC 6506; Pelotomaculum_thermopropionicum SI; Petrotoga mobilis SJ95; Polaromonas naphthalenivorans CJ2; Polaromonas sp. JS666; Pseudoalteromonas haloplanktis TAC125; Streptomyces pristinaespiralis ATCC 25486; Streptomyces pristinaespiralis ATCC 25486; Streptococcus thermophilus; Streptomyces viridochromogenes DSM 40736; Streptosporangium roseum DSM 43021; Synechococcus sp. PCC 7335; and Thermosipho africanus TCF52B (Chylinski et al., RNA Biol., 2013; 10(5): 726-737).
[0347] In some embodiments, the DNA-targeting systems or fusion proteins comprise a Cas protein, such as a Cas protein set forth in any one of SEQ ID NOS:73, 79, 126, 127, 193, 194, 197-200, and 205- 208, or a variant thereof, such as an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any one of SEQ ID NOS:73, 79, 126, 127, 193, 194, 197-200, and 205-208. In some embodiments, the Cas protein of any of the DNA-targeting systems or fusion proteins provided herein comprise a sequence set forth in any one of SEQ ID NOS:73, 79, 126, 127, 193, 194, 197-200, and 205-208, or a variant thereof, such as an amino acid sequence that has at
least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any one of SEQ ID NOS:73, 79, 126, 127, 193, 194, 197-200, and 205-208. In some aspects, the Cas protein lacks an initial methionine residue. In some aspects, the Cas protein comprises an initial methionine residue.
[0348] In some aspects, in the provided DNA-targeting systems and fusion proteins, the DNA- targeting domain, e.g., Cas, is a deactivated Cas (dCas), or a nuclease-inactive Cas (iCas). In some embodiments, the component of the DNA-targeting domain, such as a protein component, comprises a Cas9 variant such as a deactivated Cas9 or inactivated Cas9. In some embodiments, the component of the DNA-targeting domain, such as a protein component, comprises a Cas 12a variant such as a deactivated Casl2a (Cpfl) or inactivated Casl2a (Cpfl). In some aspects, the Cas9 protein may be mutated so that the nuclease activity is deactivated or inactivated (also referred to as dCas9 or iCas9). In some aspects, the Cas protein is a variant that lacks nuclease activity (i.e. is a dCas or iCas protein). In some embodiments, the Cas protein is mutated so that nuclease activity is reduced or eliminated. Such Cas proteins are referred to as deactivated Cas or dead Cas (dCas) or nuclease-inactive Cas (iCas) proteins, as referred to interchangeably herein. In some embodiments, the variant Cas protein is a variant Cas9 protein that lacks nuclease activity or that is a deactivated Cas9 (dCas9, or iCas9) protein. In some embodiments, the variant Cas protein is a variant Cpfl protein that lacks nuclease activity or that is a deactivated Casl2a (dCasl2a, or iCasl2a) protein.
[0349] In some embodiments, Cas proteins are engineered to be catalytically inactivated or nuclease inactive to allow targeting of Cas/gRNA RNPs without inducing cleavage at the target site. Mutations in Cas proteins can reduce or abolish nuclease activity of the Cas protein, rendering the Cas protein catalytically inactive. Cas proteins with reduced or abolished nuclease activity are referred to as deactivated Cas (dCas), or nuclease-inactive Cas (iCas) proteins, as referred to interchangeably herein. In some aspects, the dCas or iCas can still bind to target site in the DNA in a site- and/or sequence-specific manner, as long as it retains the ability to interact with the guide RNA (gRNA) which directs the Cas- gRNA combination to the target site.
[0350] In some aspects, the dCas or iCas exhibits reduced or no endodeoxyribonuclease activity. For example, an exemplary dCas or iCas, for example dCas9 or iCas9, exhibits less than about 20%, less than about 15%, less than about 10%, less than about 5%, less than about 1%, or less than about 0.1%, of the endodeoxyribonuclease activity of a wild-type Cas protein, e.g., a wild-type Cas9 protein. In some embodiments, the dCas or iCas, for example dCas9 or iCas9, exhibits substantially no detectable endodeoxyribonuclease activity. In some embodiments, an exemplary dCas or iCas, for example dCas9 or iCas9, comprises one or more amino acid mutations, substitutions, deletions or insertions at a position corresponding to a position selected from D10, G12, G17, E762, H840, N854, N863, H982, H983, A984, D986, and/or a A987, with reference to a wild-type Streptococcus pyogenes Cas9 (SpCas9), for example, with reference to numbering of positions of a SpCas9 sequence set forth in SEQ ID NO:79. In some aspects, the dCas9 or iCas9 comprises one or more amino acid mutations, substitutions, deletions or insertions corresponding to D10A, G12A, G17A, E762A, H840A, N854A, N863A, H982A, H983A,
A984A, and/or D986A, with reference to a wild-type Streptococcus pyogenes Cas9 (SpCas9), for example, with reference to numbering of positions of a SpCas9 sequence set forth in SEQ ID NO:79. Corresponding positions for mutations can be determined based on sequence alignments and determination of sequence conservation, for example, as described in WO 2013/171772 for Cas9 proteins from various species. In some aspects, the dCas protein lacks an initial methionine residue. In some aspects, the dCas protein comprises an initial methionine residue.
[0351] In some embodiments, the dCas9 protein can comprise a sequence from a Cas9 molecule, or variant thereof. In some embodiments, the dCas9 protein can comprise a sequence derived from a Cas9 molecule of .S’, pyogenes, S. thermophilus, S. aureus, N. meningitidis, F. novicida, S. canis, S. auricularis, or variant thereof. In some embodiments, the dCas9 protein comprises a sequence from a Cas9 molecule of .S', aureus. In some embodiments, the dCas9 protein comprises a sequence from a Cas9 molecule of .S'. pyogenes. In some embodiments, the dCas9 protein comprises a sequence from a Cas9 molecule of C. jejuni.
[0352] Exemplary deactivated Cas9 (dCas9) derived from .S'. pyogenes contains silencing mutations of the RuvC and HNH nuclease domains (D10A and H840A), for example as described in WO 2013/176772, WO 2014/093661, Jinek et al. Science 337(6096):816-21 (2012), and Qi et al. Cell 152(5): 1173-83 (2013). Exemplary dCas variants derived from the Casl2 system (i.e. Cpfl) are described, for example in WO 2017/189308 and Zetsche et al. Cell 163(3):759-71 (2015). Conserved domains that mediate nucleic acid cleavage, such as RuvC and HNH endonuclease domains, are readily identifiable in Cas orthologs, and can be mutated to produce inactive variants, for example as described in Zetsche et al. Cell 163(3):759-71 (2015). Other exemplary Cas orthologs or variants include engineered variants based on a Casl2f (also known as Casl4), including those described in Xu et al., Mol. Cell 81(20):4333-4345 (2021).
[0353] In some embodiments, the DNA-targeting domain comprises a Cas-gRNA combination that includes (a) a Cas protein or a variant thereof and (b) at least one gRNA. In some embodiments, the variant Cas protein lacks nuclease activity or is a deactivated Cas (dCas) protein. In some embodiments, the gRNA is capable of complexing with the Cas protein or variant thereof. In some embodiments, the gRNA comprises a gRNA spacer sequence that is capable of hybridizing to the target site or is complementary to the target site (e.g., in a FXN locus).
[0354] In some embodiments, the Cas protein or a variant thereof is a Cas9 protein or a variant thereof. In some embodiments, the variant Cas protein is a variant Cas9 protein that lacks nuclease activity or that is a deactivated Cas9 (dCas9) protein. In some embodiments, the Cas9 protein or a variant thereof is a Staphylococcus aureus Cas9 (SaCas9) protein or a variant thereof. In some embodiments, the variant Cas9 is a Staphylococcus aureus dCas9 protein (dSaCas9) that comprises at least one amino acid mutation selected from D10A and N580A, with reference to numbering of positions of SEQ ID NO:73. In some embodiments, the variant Cas9 protein comprises the sequence set forth in SEQ ID NO:72, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99%
sequence identity thereto. In some embodiments, the variant Cas9 protein comprises the sequence set forth in SEQ ID NO:72, which lacks an initial methionine residue. In some embodiments, the variant Cas9 protein comprises the sequence set forth in SEQ ID NO: 189, which includes an initial methionine residue. An exemplary nucleotide sequence encoding the variant Cas9 protein is set forth in SEQ ID NO:70.
[0355] In some embodiments, the Cas9 protein or variant thereof is a Streptococcus pyogenes Cas9 (SpCas9) protein or a variant thereof. In some embodiments, the variant Cas9 is a Streptococcus pyogenes dCas9 (dSpCas9) protein that comprises at least one amino acid mutation selected from D10A and H840A, with reference to numbering of positions of SEQ ID NO:79. In some embodiments, the variant Cas9 protein comprises the sequence set forth in SEQ ID NO:78, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the variant Cas9 protein comprises the sequence set forth in SEQ ID NO:78, which lacks an initial methionine residue. In some embodiments, the variant Cas9 protein comprises the sequence set forth in SEQ ID NO: 190, which includes an initial methionine residue. An exemplary nucleotide sequence encoding the variant Cas9 protein is set forth in SEQ ID NO:76.
[0356] In some embodiments, the Cas9 protein or variant thereof is a Campylobacter jejuni Cas9 (CjCas9) protein or a variant thereof. In some embodiments, the variant Cas9 comprises at least one amino acid mutation compared to the sequence set forth in SEQ ID NO:205 or 206. In some embodiments, the variant Cas9 protein comprises the sequence set forth in SEQ ID NO: 203, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the variant Cas9 protein comprises the sequence set forth in SEQ ID NO:204, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the variant Cas9 protein comprises the sequence set forth in SEQ ID NO:204, which lacks an initial methionine residue. In some embodiments, the variant Cas9 protein comprises the sequence set forth in SEQ ID NO:203, which includes an initial methionine residue.
[0357] In some embodiments, the Cas protein or a variant thereof is a Casl2a protein or a variant thereof. In some embodiments, the variant Cas protein is a variant Cas 12a protein that lacks nuclease activity or that is a deactivated Casl2a (dCasl2a) protein. In some embodiments, the Casl2a protein or variant thereof is a Acidaminococcus sp. Cas 12a (AsCasl2a) protein or a variant thereof. In some embodiments, the variant Casl2a is a Acidaminococcus sp. dCasl2a (dAsCasl2a) protein that comprises at least one amino acid mutation compared to the sequence set forth in SEQ ID NO: 193 or 194. In some embodiments, the variant Casl2a protein comprises the sequence set forth in SEQ ID NO:191, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the variant Casl2a protein comprises the sequence set forth in SEQ ID NO: 192, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the variant Casl2a protein
comprises the sequence set forth in SEQ ID NO: 192, which lacks an initial methionine residue. In some embodiments, the variant Casl2a protein comprises the sequence set forth in SEQ ID NO:191, which includes an initial methionine residue.
[0358] In some embodiments, the Cas protein or a variant thereof is a CasPhi-2 protein or a variant thereof. In some embodiments, the variant Cas protein is a variant CasPhi-2 protein that lacks nuclease activity or that is a deactivated CasPhi-2 (dCasPhi-2) protein. In some embodiments, the variant CasPhi-2 comprises at least one amino acid mutation compared to the sequence set forth in SEQ ID NO: 197 or 198. In some embodiments, the variant CasPhi-2 protein comprises the sequence set forth in SEQ ID NO: 195, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the variant CasPhi-2 protein comprises the sequence set forth in SEQ ID NO: 196, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the variant CasPhi- 2 protein comprises the sequence set forth in SEQ ID NO:220, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the variant CasPhi-2 protein comprises the sequence set forth in SEQ ID NO:221, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the variant CasPhi-2 protein comprises the sequence set forth in SEQ ID NO: 196, which lacks an initial methionine residue. In some embodiments, the variant CasPhi-2 protein comprises the sequence set forth in SEQ ID NO: 221, which lacks an initial methionine residue. In some embodiments, the variant CasPhi-2 protein comprises the sequence set forth in SEQ ID NO: 195, which includes an initial methionine residue. In some embodiments, the variant CasPhi-2 protein comprises the sequence set forth in SEQ ID NO:220, which includes an initial methionine residue.
[0359] In some embodiments, the Cas protein or a variant thereof is a UnlCasl2fl protein or a variant thereof. In some embodiments, the variant Cas protein is a variant UnlCasl2fl protein that lacks nuclease activity or that is a deactivated UnlCasl2fl (dUnlCasl2fl) protein. In some embodiments, the variant UnlCasl2fl comprises at least one amino acid mutation compared to the sequence set forth in SEQ ID NO: 199 or 200. In some embodiments, the variant UnlCasl2fl protein comprises the sequence set forth in SEQ ID NO:201, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the variant UnlCasl2fl protein comprises the sequence set forth in SEQ ID NO:202, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the variant UnlCasl2fl protein comprises the sequence set forth in SEQ ID NO:202, which lacks an initial methionine residue. In some embodiments, the variant UnlCasl2fl protein comprises the sequence set forth in SEQ ID NO:201, which includes an initial methionine residue.
[0360] In some embodiments, the Cas protein or a variant thereof is a Cas 12k protein or a variant thereof. In some embodiments, the Casl2k protein comprises the sequence set forth in SEQ ID NO:207,
or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the Casl2k protein comprises the sequence set forth in SEQ ID NO:208, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the Casl2k protein comprises the sequence set forth in SEQ ID NO:208, which lacks an initial methionine residue. In some embodiments, the Casl2k protein comprises the sequence set forth in SEQ ID NO:207, which includes an initial methionine residue.
[0361] In some embodiments, the Cas protein or a variant thereof is a CasMini protein or a variant thereof, such as an engineered Cas protein or variant based on a Casl2f (also known as Casl4), including those described in Xu et al., Mol. Cell 81(20):4333-4345 (2021) or set forth in SEQ ID NO:223. In some embodiments, the variant Cas protein is a variant CasMini protein that lacks nuclease activity or that is a deactivated CasMini (dCasMini) protein. In some embodiments, the variant CasMini comprises at least one amino acid mutation compared to the sequence set forth in SEQ ID NO:223. In some embodiments, the variant CasMini protein comprises the sequence set forth in SEQ ID NO:223, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the CasMini protein comprises the sequence set forth in SEQ ID NO: 223. In some embodiments, the variant CasMini protein comprises the sequence set forth in SEQ ID NO: 209 or 210, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the CasMini protein comprises the sequence set forth in SEQ ID NO:209, which lacks an initial methionine residue. In some embodiments, the CasMini protein comprises the sequence set forth in SEQ ID NO:210, which includes an initial methionine residue.
[0362] DNA-targeting systems, in some cases comprising a fusion protein, such as dCas-fusion proteins include fusion of the Cas with an effector domain, such as a transcription activation domain. Any of a variety of effector domains, for example those that increase transcription from the target locus, e.g., FXN locus, including any described herein, can be used.
[0363] In some aspects, provided is a DNA-targeting system comprising a fusion protein comprising a DNA-targeting domain comprising a nuclease-inactive Cas protein or variant thereof, and an effector domain for increasing or inducing transcriptional activation (i.e. a transcriptional activator) when targeted to a target site in a FXN gene or regulatory element thereof. In some aspects, the DNA-targeting system also includes one or more gRNA, provided in combination or as a complex with the dCas protein or variant thereof, for targeting of the DNA-targeting system to the target site. In some embodiments, the fusion protein is guided to a specific target site sequence of the target gene by the guide RNA, wherein the effector domain mediates targeted epigenetic modification to increase or promote transcription of the target gene.
1. Guide RNAs (gRNAs)
[0364] Provided herein are gRNAs, such as gRNAs that target or can bind to a regulatory DNA element of a FXN locus. In some embodiments, the gRNA is capable of complexing with the Cas protein or variant thereof. In some embodiments, the gRNA comprises a gRNA spacer sequence (also known as a spacer sequence or a guide sequence) that is capable of hybridizing to the target site or is complementary to the target site, such as any target site described herein, for example, any target site in a genome. In some embodiments, the gRNA comprises a scaffold sequence that complexes with or binds to the Cas protein. In some embodiments, a gRNA specific to a target locus of interest (e.g. a regulatory DNA element of a FXN locus) is used to recruit an RNA-guided protein (e.g. a Cas protein) or variant thereof or a fusion protein comprising such RNA-guided protein (e.g., a Cas polypeptide), to the target site.
[0365| In some embodiments, the Cas protein (e.g. dCas9) is provided in combination or as a complex with one or more guide RNA (gRNA). In some aspects, the gRNA is a nucleic acid that promotes the specific targeting or homing of the gRNA/Cas RNP complex to the target site, such as any described above. In some embodiments, a target site of a gRNA may be referred to as a protospacer.
[0366] Provided herein are gRNAs, such as gRNAs that target or bind to a target site in a FXN gene or DNA regulatory element thereof, such as any described above in Section ILA. In some embodiments, the gRNA is capable of complexing with the Cas protein or variant thereof. In some embodiments, the gRNA comprises a gRNA spacer sequence (i.e. a spacer sequence or a guide sequence) that is capable of hybridizing to the target site, or that is complementary to the target site, such as any target site described in Section II. A or further below. In some embodiments, the gRNA comprises a scaffold sequence that complexes with or binds to the Cas protein.
[0367] In some aspects, a “gRNA molecule” is a nucleic acid that promotes the specific targeting or homing of a gRNA molecule/Cas9 molecule complex to a target nucleic acid, such as a locus on the genomic DNA of a cell. gRNA molecules can be unimolecular (having a single RNA molecule), sometimes referred to herein as “chimeric” gRNAs, or modular (comprising more than one, and typically two, separate RNA molecules). In general, a spacer sequence of the guide RNA, is any polynucleotide sequences comprising at least a sequence portion that has sufficient complementarity with a target polynucleotide sequence, such as the at the FXN locus in humans, to hybridize with the target sequence at the target site and direct sequence-specific binding of the CRISPR complex to the target sequence. In some embodiments, in the context of formation of a CRISPR complex, “target sequence” is to a sequence to which a spacer sequence is designed to have complementarity, where hybridization between the target sequence and a spacer sequence of the guide RNA promotes the formation of a CRISPR complex. Full complementarity is not necessarily required, provided there is sufficient complementarity to cause hybridization and promote formation of a CRISPR complex. Generally, a spacer sequence is selected to reduce the degree of secondary structure within the spacer sequence. Secondary structure may be determined by any suitable polynucleotide folding algorithm.
[0368] In some embodiments, a guide RNA (gRNA) specific to a target locus of interest (e.g. at the
FXN locus in humans) is used with RNA-guided nucleases or variants thereof, e.g., nuclease-inactive Cas variants, to target the provided DNA-targeting system to the target site or target position. Methods for designing gRNAs and exemplary spacer sequences are known. Exemplary gRNA structures that can be associated with particular RNA-guided nucleases or variants thereof, e.g., nuclease-inactive Cas variants, with particular domains and scaffold regions, are also known. In some aspects, gRNA molecules comprise a scaffold sequence, e.g., sequences that can be complexed with the Cas protein. In some aspects, the scaffold sequence is specific for the Cas protein.
[0369] In some embodiments, the gRNA is a chimeric gRNA. In general, gRNAs can be unimolecular (i.e. composed of a single RNA molecule), or modular (comprising more than one, and typically two, separate RNA molecules). Modular gRNAs can be engineered to be unimolecular, wherein sequences from the separate modular RNA molecules are comprised in a single gRNA molecule, sometimes referred to as a chimeric gRNA, synthetic gRNA, or single gRNA. A guide RNA can comprise at least a spacer sequence that hybridizes to a target nucleic acid sequence of interest, and a CRISPR repeat sequence. In Type II systems, the gRNA also comprises a second RNA called the tracrRNA sequence. In the Type II guide RNA (gRNA), the CRISPR repeat sequence and tracrRNA sequence hybridize to each other to form a duplex. In the Type V guide RNA (gRNA), the crRNA forms a duplex. In both systems, the duplex can bind a site-directed polypeptide, such that the guide RNA and site-direct polypeptide form a complex. The gRNA can provide target specificity to the complex by virtue of its association with the site-directed polypeptide. The gRNA thus can direct the activity of the site-directed polypeptide.
[0370] In some embodiments, the chimeric gRNA is a fusion of two non-coding RNA sequences: a crRNA sequence and a tracrRNA sequence, for example as described in WO 2013/176772, or Jinek, M. et al. Science 337(6096):816-21 (2012). In some embodiments, the chimeric gRNA mimics the naturally occurring crRNA:tracrRNA duplex involved in the Type II CRISPR/Cas system, wherein the naturally occurring crRNA:tracrRNA duplex acts as a guide for the Cas protein, e.g., Cas9 protein. Exemplary types of CRISPR/Cas systems and associated gRNA structures include those described in, for example, Moon et al. Exp. Mol. Med. 51, 1-11 (2019), Zhang, F. Q. Rev. Biophys. 52, E6 (2019), Makarova et al. Methods Mol. Biol. 1311:47-75 (2015), WO 2013/176772, or Jinek, M. et al. Science 337(6096):816-21 (2012).
[0371] In some aspects, the spacer sequence of a gRNA is a polynucleotide sequence comprising at least a portion that has sufficient complementarity with the target site to hybridize with the target site and direct sequence-specific binding of a CRISPR complex to the sequence of the target site. Full complementarity is not necessarily required, provided there is sufficient complementarity to cause hybridization and promote formation of a CRISPR complex. In some embodiments, the gRNA comprises a spacer sequence that is complementary, e.g., at least 80%, 85%, 90%, 95%, 98%, 99%, or 100% (e.g., fully complementary), to the target site. The strand of the target nucleic acid comprising the target site sequence may be referred to as the “complementary strand” of the target nucleic acid. In some aspects,
the spacer sequence is a user-defined sequence. Guidance on the selection of spacer sequences can be found, e.g., in Fu et al., Nat Biotechnol 2014 32:279-284 and Sternberg et al., Nature 2014 507:62-67.
[0372] In some embodiments, the gRNA spacer sequence is between about 14 nt and about 26 nt, between about 14 nt and about 24 nt, or between 16 nt and 22 nt in length. In some embodiments, the gRNA spacer sequence is 14 nt, 15 nt, 16 nt, 17 nt, 18 nt, 19 nt, 20 nt, 21 nt or 22 nt, 23 nt, 24 nt, 25 nt, or 26 nt in length. In some embodiments, the gRNA spacer sequence is 18 nt, 19 nt, 20 nt, 21 nt or 22 nt in length. In some embodiments, the gRNA spacer sequence is 18 nt in length. In some embodiments, the gRNA spacer sequence is 19 nt in length. In some embodiments, the gRNA spacer sequence is 20 nt in length. In some embodiments, the gRNA spacer sequence is 21 nt in length. In some embodiments, the gRNA spacer sequence is 22 nt in length.
[03733 Methods for designing gRNAs and exemplary targeting domains can include those described in, e.g., International PCT Pub. Nos. WO 2014/197748, WO 2016/130600, WO 2017/180915, WO 2021/226555, WO 2013/176772, WO 2014/152432, WO 2014/093661, WO 2014/093655, WO 2015/089427, WO 2016/049258, WO 2016/123578, WO 2021/076744, WO 2014/191128, WO 2015/161276, WO 2017/193107, and WO 2017/093969.
[0374] A target site of a gRNA may be referred to as a protospacer. In some aspects, the spacer is designed to target a protospacer with a specific protospacer-adjacent motif (PAM), i.e. a sequence immediately adjacent to the protospacer that contributes to and/or is required for Cas binding specificity. Different CRISPR/Cas systems have different PAM requirements for targeting. For example, in some embodiments, 5. pyogenes Cas9 uses the PAM 5’-NGG-3’ (SEQ ID NO:142), where N is any nucleotide. 5. aureus Cas9 uses the PAM 5’- NNGRRT-3’ (SEQ ID NO: 143), where N is any nucleotide, and R is G or A. N. meningitidis Cas9 uses the PAM 5'-NNNNGATT -3’ (SEQ ID NO: 144), where N is any nucleotide. C. jejuni Cas9 uses the PAM 5'-NNNNRYAC-3' (SEQ ID NO: 145) or 5'-NNNNACAC- 3 ’(SEQ ID NO:226), where N is any nucleotide, R is G or A, and Y is C or T. S. thermophilus uses the PAM 5’-NNAGAAW-3’ (SEQ ID NO: 146), where N is any nucleotide and W is A or T. F. Novicida Cas9 uses the PAM 5’-NGG-3’ (SEQ ID NO: 142), where N is any nucleotide. T. denticola Cas9 uses the PAM 5’-NAAAAC-3’ (SEQ ID NO: 146), where N is any nucleotide. Casl2a (also known as Cpfl) from various species, uses the PAM 5’-TTTV-3’ (SEQ ID NO:148), where V is A, C, or G. Phage-derived CasPhi (such as CasPhi-2, also known as Casl2j), uses the PAM 5’-TBN-3’ (SEQ ID NO:224), where N is any nucleotide, and B is G, T, or C. Archaeal UnlCasl2fl (also known as Casl4al), uses the PAM 5’- TTTN -3’ (SEQ ID NO:225), where N is any nucleotide. A Casl2f protein (also known as Casl4) uses the PAM 5’- TTTR -3’ (SEQ ID NO:228), where R is G or A. A Casl2k protein uses the PAM 5’- GGTT -3’ (SEQ ID NO:227). Cas proteins may use or be engineered to use different PAMs from those listed above. For example, variant SpCas9 proteins may use a PAM selected from: 5’-NGG-3’ (SEQ ID NO:142), 5’-NGAN-3’ (SEQ ID NO:149), 5’-NGNG-3’(SEQ ID NO:15Q), 5’-NGAG-3’(SEQ ID NO:151), or 5’-NGCG-3’(SEQ ID NO:152), where N is any nucleotide. Methods for designing or identifying gRNA spacer sequences and/or protospacer sequences in a particular region, are known.
gRNA spacer sequences and/or protospacer sequences can be determined based on the type of Cas protein used and the associated PAM sequence.
[0375] In some embodiments, the PAM of a gRNA for complexing with .S', pyogenes Cas9 or variant thereof is set forth in SEQ ID NO: 141. In some embodiments, the PAM of a gRNA for complexing with .S', aureus Cas9 or variant thereof is set forth in SEQ ID NO: 142. In some embodiments, the PAM of a gRNA for complexing with a Type V CRISPR/Cas system, such as with Casl2a (also known as Cpfl) or variant thereof is set forth in SEQ ID NO: 147.
[0376] A spacer sequence may be selected to reduce the degree of secondary structure within the spacer sequence. Secondary structure may be determined by any suitable polynucleotide folding algorithm.
[0377| In some embodiments, the gRNA (including the spacer sequence) will comprise the base uracil (U), whereas DNA encoding the gRNA molecule will comprise the base thymine (T). While not wishing to be bound by theory, in some embodiments, it is believed that the complementarity of the spacer sequence (i.e. guide sequence) with the target sequence contributes to specificity of the interaction of the gRNA molecule/Cas molecule complex with a target nucleic acid. It is understood that in a spacer sequence (i.e. guide sequence) and target sequence pair, the uracil bases in the spacer sequence (i.e. guide sequence) will pair with the adenine bases in the target sequence. A gRNA spacer sequence herein may be defined by the DNA sequence encoding the gRNA spacer, and/or the RNA sequence of the spacer.
[0378] In some embodiments, the gRNA comprises modified nucleotides, e.g. for increased stability. In some embodiments, one, more than one, or all of the nucleotides of a gRNA can have a modification, e.g., to render the gRNA less susceptible to degradation and/or improve bio-compatibility. By way of non-limiting example, the backbone of the gRNA can be modified with a phosphorothioate, or other modification(s). In some cases, a nucleotide of the gRNA can comprise a 2’ modification, e.g., a 2- acetylation, e.g., a 2’ methylation, or other modification(s)
[0379] In some embodiments the gRNA is a concatenation of two non-coding RNA sequences: a crRNA sequence and a tracrRNA sequence. The gRNA may target a desired DNA sequence by exchanging the sequence encoding a 20 bp protospacer which confers targeting specificity through complementary base pairing with the desired DNA target. gRNA mimics the naturally occurring crRNA:tracrRNA duplex involved in the Type II CRISPR/Cas system (e.g., Cas9). This duplex, which may include, for example, a 42-nucleotide crRNA and a 75-nucleotide tracrRNA, acts as a guide for the Cas9 protein to cleave the target nucleic acid. The “target region”, “target sequence” or “protospacer” as used interchangeably herein refers to the region of the target gene to which the CRISPR/Cas9-based system targets. The CRISPR/Cas9-based system may include two or more gRNAs, wherein the two or more gRNAs target different DNA sequences. The target DNA sequences may be overlapping or nonoverlapping. The target DNA sequences may be located within or near the same gene or different genes. The target sequence or protospacer is followed by a PAM sequence at the 3' end of the protospacer. Different Type II systems have differing PAM requirements. For example, the Streptococcus
pyogenes Type II system uses an “NGG” sequence, where “N” can be any nucleotide.
[0380] In some aspects, the gRNA comprises scaffold sequences. In some aspects, the scaffold sequence (in some cases including a crRNA sequence and/or a tracrRNA sequence) will be different depending on the Cas protein. In some aspects, different CRISPR/Cas systems have different gRNA scaffold sequences for associating with Cas protein. In some embodiments, an exemplary scaffold sequence for .S', aureus Cas9 comprises a sequence set forth in SEQ ID NO:44, or a sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to SEQ ID NO:44. In some embodiments, an exemplary scaffold sequence for .S'. aureus Cas9 comprises a sequence set forth in SEQ ID NO:44. In some embodiments, an exemplary scaffold sequence for .S', pyogenes Cas9 comprises a sequence set forth in SEQ ID NO:46, or a sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to SEQ ID NO:46. In some embodiments, an exemplary scaffold sequence for .S', pyogenes Cas9 comprises a sequence set forth in SEQ ID NO:46. In some embodiments, an exemplary scaffold sequence for Acidaminococcus sp. Casl2a comprises a sequence set forth in SEQ ID NO:211, or a sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to SEQ ID NO:211. In some embodiments, an exemplary scaffold sequence for CasPhi-2 comprises a sequence set forth in SEQ ID NO:212, or a sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to SEQ ID NO:212. In some embodiments, an exemplary scaffold sequence for UnlCasl2fl comprises a sequence set forth in SEQ ID NO:213, 214 or 215, or a sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to SEQ ID NO:213, 214 or 215. In some embodiments, an exemplary scaffold sequence for UnlCasl2fl comprises a sequence set forth in SEQ ID NO:213, or a sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to SEQ ID NO:213. In some embodiments, an exemplary scaffold sequence for UnlCasl2fl comprises a sequence set forth in SEQ ID NO:214, or a sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to SEQ ID NO:214. In some embodiments, an exemplary scaffold sequence for UnlCasl2fl comprises a sequence set forth in SEQ ID NO:215, or a sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to SEQ ID NO:215. In some embodiments, an exemplary scaffold sequence for C. jejuni Cas9 comprises a sequence set forth in SEQ ID NO:216, or a sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to SEQ ID NO:216. In some embodiments, an exemplary scaffold sequence for Cas 12k comprises a sequence set forth in SEQ ID NO:217, or a sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to SEQ ID NO:217. In some embodiments, an exemplary scaffold sequence for CasMini comprises a sequence set forth in SEQ ID NO:218, or a
sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to SEQ ID NO:218.
[0381] In some aspects, the gRNA can target the DNA-targeting system to direct the activities of an associated polypeptide (e.g., fusion protein, DNA-targeting system, effector domain, etc.) to a specific target site within a target nucleic acid (e.g., regulatory DNA element of a FXN locus).
[0382] In some aspects, provided herein is a guide RNA (gRNA) that binds a target site in an enhancer region of a frataxin (FXN) locus, wherein the target site is located within the genomic coordinates human genome assembly GRCh38 (hg38) chr9:69, 027, 282-69, 028, 497. In some aspects, provided herein is a guide RNA (gRNA) that binds a target site in an enhancer region of a frataxin (FXN) locus, wherein the target site is located within the genomic coordinates hg38 chr9:69,027,615- 69,028,101.
[0383] In some embodiments the target site comprises the sequence set forth in SEQ ID NO:21, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing. In some embodiments DNA encoding the gRNA comprises the sequence set forth in SEQ ID NO:21, a contiguous portion thereof of at least 14 nucleotides, or a complementary sequence of any of the foregoing. In some embodiments the gRNA comprises at least one gRNA spacer comprising the sequence set forth in SEQ ID NO:42, or a contiguous portion thereof of at least 14 nt. In some embodiments the gRNA further comprises the sequence set forth in SEQ ID NO:44. In some embodiments the gRNA comprises the sequence set forth in SEQ ID NO:67.
[0384] Also provided herein is a guide RNA (gRNA) that binds a target site in a regulatory DNA element of a frataxin (FXN) locus, such as a FXN promoter, wherein the target site comprises a sequence selected from any one of SEQ ID NOS: 1-10, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing.
[0385] In some of any of the provided embodiments, the gRNA comprises at least one gRNA spacer sequence comprising a sequence selected from any one of SEQ ID NOS:22-31, or a contiguous portion thereof of at least 14 nt. In some of any of the provided embodiments, the gRNA further comprises the sequence set forth in SEQ ID NO:44. In some of any of the provided embodiments, the gRNA comprises a sequence selected from any one of SEQ ID NOS:47-56.
[0386] Also provided herein is a guide RNA (gRNA) that binds a target site in a regulatory DNA element of a frataxin (FXN) locus, such as a FXN promoter, wherein the target site comprises a sequence selected from any one of SEQ ID NOS: 11-20, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing.
[0387] In some of any of the provided embodiments, the gRNA comprises at least one gRNA spacer sequence comprising a sequence selected from any one of SEQ ID NOS:32-41, or a contiguous portion thereof of at least 14 nt. In some of any of the provided embodiments, the gRNA further comprises the sequence set forth in SEQ ID NO:46. In some of any of the provided embodiments, the gRNA comprises a sequence selected from any one of SEQ ID NOS:57-66.
[0388] In some embodiments, the DNA-targeting domain comprises a Cas-gRNA combination comprising (a) a Cas protein or a variant thereof and (b) at least one gRNA; and the gRNA comprises at least one gRNA spacer sequence comprising a sequence selected from any one of SEQ ID NOS:22-31, or a contiguous portion thereof of at least 14 nt. In some embodiments, the gRNA further comprises the sequence set forth in SEQ ID NO:44. In some embodiments, the gRNA comprises a sequence selected from any one of SEQ ID NOS:47-56.
[0389] In some embodiments, the DNA-targeting domain comprises a Cas-gRNA combination comprising (a) a Cas protein or a variant thereof and (b) at least one gRNA; and the gRNA comprises at least one gRNA spacer sequence comprising a sequence selected from any one of SEQ ID NOS:32-41, or a contiguous portion thereof of at least 14 nt. In some embodiments, the gRNA further comprises the sequence set forth in SEQ ID NO:46. In some embodiments, the gRNA comprises a sequence selected from any one of SEQ ID NOS:57-66.
[0390] In some embodiments, the gRNA comprises a sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to all or a portion of a gRNA sequence or a gRNA spacer sequence described herein.
[0391] In some embodiments, the gRNA targets a target site in a FXN locus or a DNA regulatory element thereof that comprises the sequence selected from any one of SEQ ID NO:1-10 and 21, a contiguous portion thereof of at least 14 nucleotides (e.g., 14, 15, 16, 17, 18, 19, 20, 21, or 22 nucleotides), a complementary sequence of any of the foregoing, or a sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to any of the foregoing. In some embodiments, the gRNA comprises a spacer sequence comprising the sequence selected from any one of SEQ ID NO:22-31 and 42, a contiguous portion thereof of at least 14 nt (e.g., 14, 15, 16, 17, 18, 19, 20, 21, or 22 nucleotides), or a sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to any of the foregoing. In some embodiments, the gRNA further comprises a scaffold sequence. In some embodiments, the scaffold sequence comprises the sequence set forth in SEQ ID NO:44, or a sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to SEQ ID NO:44. In some embodiments, the gRNA, including a spacer sequence and a scaffold sequence, comprises the sequence selected from any one of SEQ ID NO:47-56 and 67, or a sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to all or a portion thereof. In some embodiments, the gRNA is set forth in the sequence selected from any one of SEQ ID NO:47-56 and 67. In some embodiments, a provided DNA-targeting system comprises any of the aforementioned gRNAs complexed with a Cas protein, such as a Cas9 protein. In some embodiments, the Cas9 is a dCas9. In some embodiments, the dCas9 is a dSaCas9, such as a dSaCas9 set forth in SEQ ID NO:72, or a variant and/or fusion thereof.
[0392] In some embodiments, the gRNA targets a target site in a FXN locus or a DNA regulatory
element thereof that comprises the sequence selected from any one of SEQ ID NO:229-255, a contiguous portion thereof of at least 14 nucleotides (e.g., 14, 15, 16, 17, 18, 19, 20, 21, or 22 nucleotides), a complementary sequence of any of the foregoing, or a sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to any of the foregoing. In some embodiments, the gRNA further comprises a scaffold sequence. In some embodiments, the scaffold sequence comprises the sequence set forth in SEQ ID NO:211, or a sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to SEQ ID NO:211. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:229, and a scaffold sequence of SEQ ID NO:211. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:230, and a scaffold sequence of SEQ ID NO:211. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:231, and a scaffold sequence of SEQ ID NO:211. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:232, and a scaffold sequence of SEQ ID NO:211. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:233, and a scaffold sequence of SEQ ID NO:211. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:234, and a scaffold sequence of SEQ ID NO:211. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:235, and a scaffold sequence of SEQ ID NO:211. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:236, and a scaffold sequence of SEQ ID NO:211. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:237, and a scaffold sequence of SEQ ID NO:211. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:238, and a scaffold sequence of SEQ ID NO:211. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:239, and a scaffold sequence of SEQ ID NO:211. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:240, and a scaffold sequence of SEQ ID NO:211. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:241, and a scaffold sequence of SEQ ID NO:211. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:242, and a scaffold sequence of SEQ ID NO:211. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:243, and a scaffold sequence of SEQ ID NO:211. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:244, and a scaffold sequence of SEQ ID NO:211. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:245, and a scaffold sequence of SEQ ID NO:211. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:246, and a scaffold sequence of SEQ ID NO:211. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:247, and a scaffold sequence of SEQ ID NO:211. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:248, and a scaffold sequence of SEQ ID NO:211. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:249, and a scaffold sequence of SEQ ID NO:211. In some
embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:250, and a scaffold sequence of SEQ ID NO:211. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:251, and a scaffold sequence of SEQ ID NO:211. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:252, and a scaffold sequence of SEQ ID NO:211. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:253, and a scaffold sequence of SEQ ID NO:211. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:254, and a scaffold sequence of SEQ ID NO:211. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:255, and a scaffold sequence of SEQ ID NO:211. In some embodiments, a provided DNA-targeting system comprises any of the aforementioned gRNAs complexed with a Cas protein, such as a Cast 2a (also known as Cpfl) protein. In some embodiments, the Casl2a is a dCasl2a. In some embodiments, the dCasl2a is a dSaCasl2a, such as a dSaCasl2a set forth in SEQ ID NO:192, or a variant and/or fusion thereof.
[0393] In some aspects, the gRNA targets a target site in a FXN locus that comprises SEQ ID NO:1, a contiguous portion thereof of at least 14 nucleotides, a complementary sequence of any of the foregoing, or a sequence having at least 90% sequence identity to any of the foregoing. In some aspects, the gRNA comprises a spacer sequence comprising SEQ ID NO:22, a contiguous portion thereof of at least 14 nt, or a sequence having at least 90% sequence identity to any of the foregoing. In some aspects, the gRNA further comprises a scaffold sequence comprising SEQ ID NO:44, or a sequence having at least 90% sequence identity to SEQ ID NO:44. In some aspects, the gRNA, including a spacer sequence and a scaffold sequence, comprises SEQ ID NO:47, or a sequence having at least 90% sequence identity to all or a portion thereof. In some aspects, the gRNA is set forth in SEQ ID NO:47. In some aspects, a provided DNA-targeting system comprises any of the above gRNAs complexed with a Cas protein, such as a Cas9 protein. In some aspects, the Cas9 is a dCas9. In some aspects, the dCas9 is a dSaCas9, such as a dSaCas9 set forth in SEQ ID NO:72, or a variant and/or fusion thereof.
[0394] In some aspects, the gRNA targets a target site in a FXN locus that comprises SEQ ID NO:2, a contiguous portion thereof of at least 14 nucleotides, a complementary sequence of any of the foregoing, or a sequence having at least 90% sequence identity to any of the foregoing. In some aspects, the gRNA comprises a spacer sequence comprising SEQ ID NO:23, a contiguous portion thereof of at least 14 nt, or a sequence having at least 90% sequence identity to any of the foregoing. In some aspects, the gRNA further comprises a scaffold sequence comprising SEQ ID NO:44, or a sequence having at least 90% sequence identity to SEQ ID NO:44. In some aspects, the gRNA, including a spacer sequence and a scaffold sequence, comprises SEQ ID NO:48, or a sequence having at least 90% sequence identity to all or a portion thereof. In some aspects, the gRNA is set forth in SEQ ID NO:48. In some aspects, a provided DNA-targeting system comprises any of the above gRNAs complexed with a Cas protein, such as a Cas9 protein. In some aspects, the Cas9 is a dCas9. In some aspects, the dCas9 is a dSaCas9, such as a dSaCas9 set forth in SEQ ID NO:72, or a variant and/or fusion thereof.
[0395] In some aspects, the gRNA targets a target site in a FXN locus that comprises SEQ ID NO:3,
a contiguous portion thereof of at least 14 nucleotides, a complementary sequence of any of the foregoing, or a sequence having at least 90% sequence identity to any of the foregoing. In some aspects, the gRNA comprises a spacer sequence comprising SEQ ID NO:24, a contiguous portion thereof of at least 14 nt, or a sequence having at least 90% sequence identity to any of the foregoing. In some aspects, the gRNA further comprises a scaffold sequence comprising SEQ ID NO:44, or a sequence having at least 90% sequence identity to SEQ ID NO:44. In some aspects, the gRNA, including a spacer sequence and a scaffold sequence, comprises SEQ ID NO:49, or a sequence having at least 90% sequence identity to all or a portion thereof. In some aspects, the gRNA is set forth in SEQ ID NO:49. In some aspects, a provided DNA-targeting system comprises any of the above gRNAs complexed with a Cas protein, such as a Cas9 protein. In some aspects, the Cas9 is a dCas9. In some aspects, the dCas9 is a dSaCas9, such as a dSaCas9 set forth in SEQ ID NO:72, or a variant and/or fusion thereof.
[0396] In some aspects, the gRNA targets a target site in a FXN locus that comprises SEQ ID NO:4, a contiguous portion thereof of at least 14 nucleotides, a complementary sequence of any of the foregoing, or a sequence having at least 90% sequence identity to any of the foregoing. In some aspects, the gRNA comprises a spacer sequence comprising SEQ ID NO:25, a contiguous portion thereof of at least 14 nt, or a sequence having at least 90% sequence identity to any of the foregoing. In some aspects, the gRNA further comprises a scaffold sequence comprising SEQ ID NO:44, or a sequence having at least 90% sequence identity to SEQ ID NO:44. In some aspects, the gRNA, including a spacer sequence and a scaffold sequence, comprises SEQ ID NO:50, or a sequence having at least 90% sequence identity to all or a portion thereof. In some aspects, the gRNA is set forth in SEQ ID NO:50. In some aspects, a provided DNA-targeting system comprises any of the above gRNAs complexed with a Cas protein, such as a Cas9 protein. In some aspects, the Cas9 is a dCas9. In some aspects, the dCas9 is a dSaCas9, such as a dSaCas9 set forth in SEQ ID NO:72, or a variant and/or fusion thereof.
[0397] In some aspects, the gRNA targets a target site in a FXN locus that comprises SEQ ID NO:5, a contiguous portion thereof of at least 14 nucleotides, a complementary sequence of any of the foregoing, or a sequence having at least 90% sequence identity to any of the foregoing. In some aspects, the gRNA comprises a spacer sequence comprising SEQ ID NO:26, a contiguous portion thereof of at least 14 nt, or a sequence having at least 90% sequence identity to any of the foregoing. In some aspects, the gRNA further comprises a scaffold sequence comprising SEQ ID NO:44, or a sequence having at least 90% sequence identity to SEQ ID NO:44. In some aspects, the gRNA, including a spacer sequence and a scaffold sequence, comprises SEQ ID NO:51, or a sequence having at least 90% sequence identity to all or a portion thereof. In some aspects, the gRNA is set forth in SEQ ID NO:51. In some aspects, a provided DNA-targeting system comprises any of the above gRNAs complexed with a Cas protein, such as a Cas9 protein. In some aspects, the Cas9 is a dCas9. In some aspects, the dCas9 is a dSaCas9, such as a dSaCas9 set forth in SEQ ID NO:72, or a variant and/or fusion thereof.
[0398] In some aspects, the gRNA targets a target site in a FXN locus that comprises SEQ ID NO:6, a contiguous portion thereof of at least 14 nucleotides, a complementary sequence of any of the
foregoing, or a sequence having at least 90% sequence identity to any of the foregoing. In some aspects, the gRNA comprises a spacer sequence comprising SEQ ID NO:27, a contiguous portion thereof of at least 14 nt, or a sequence having at least 90% sequence identity to any of the foregoing. In some aspects, the gRNA further comprises a scaffold sequence comprising SEQ ID NO:44, or a sequence having at least 90% sequence identity to SEQ ID NO:44. In some aspects, the gRNA, including a spacer sequence and a scaffold sequence, comprises SEQ ID NO:52, or a sequence having at least 90% sequence identity to all or a portion thereof. In some aspects, the gRNA is set forth in SEQ ID NO:52. In some aspects, a provided DNA-targeting system comprises any of the above gRNAs complexed with a Cas protein, such as a Cas9 protein. In some aspects, the Cas9 is a dCas9. In some aspects, the dCas9 is a dSaCas9, such as a dSaCas9 set forth in SEQ ID NO:72, or a variant and/or fusion thereof.
[0399| In some aspects, the gRNA targets a target site in a FXN locus that comprises SEQ ID NO:7, a contiguous portion thereof of at least 14 nucleotides, a complementary sequence of any of the foregoing, or a sequence having at least 90% sequence identity to any of the foregoing. In some aspects, the gRNA comprises a spacer sequence comprising SEQ ID NO:28, a contiguous portion thereof of at least 14 nt, or a sequence having at least 90% sequence identity to any of the foregoing. In some aspects, the gRNA further comprises a scaffold sequence comprising SEQ ID NO:44, or a sequence having at least 90% sequence identity to SEQ ID NO:44. In some aspects, the gRNA, including a spacer sequence and a scaffold sequence, comprises SEQ ID NO:53, or a sequence having at least 90% sequence identity to all or a portion thereof. In some aspects, the gRNA is set forth in SEQ ID NO:53. In some aspects, a provided DNA-targeting system comprises any of the above gRNAs complexed with a Cas protein, such as a Cas9 protein. In some aspects, the Cas9 is a dCas9. In some aspects, the dCas9 is a dSaCas9, such as a dSaCas9 set forth in SEQ ID NO:72, or a variant and/or fusion thereof.
[0400] In some aspects, the gRNA targets a target site in a FXN locus that comprises SEQ ID NO:8, a contiguous portion thereof of at least 14 nucleotides, a complementary sequence of any of the foregoing, or a sequence having at least 90% sequence identity to any of the foregoing. In some aspects, the gRNA comprises a spacer sequence comprising SEQ ID NO:29, a contiguous portion thereof of at least 14 nt, or a sequence having at least 90% sequence identity to any of the foregoing. In some aspects, the gRNA further comprises a scaffold sequence comprising SEQ ID NO:44, or a sequence having at least 90% sequence identity to SEQ ID NO:44. In some aspects, the gRNA, including a spacer sequence and a scaffold sequence, comprises SEQ ID NO:54, or a sequence having at least 90% sequence identity to all or a portion thereof. In some aspects, the gRNA is set forth in SEQ ID NO:54. In some aspects, a provided DNA-targeting system comprises any of the above gRNAs complexed with a Cas protein, such as a Cas9 protein. In some aspects, the Cas9 is a dCas9. In some aspects, the dCas9 is a dSaCas9, such as a dSaCas9 set forth in SEQ ID NO:72, or a variant and/or fusion thereof.
[0401] In some aspects, the gRNA targets a target site in a FXN locus that comprises SEQ ID NO:9, a contiguous portion thereof of at least 14 nucleotides, a complementary sequence of any of the foregoing, or a sequence having at least 90% sequence identity to any of the foregoing. In some aspects,
the gRNA comprises a spacer sequence comprising SEQ ID NO:30, a contiguous portion thereof of at least 14 nt, or a sequence having at least 90% sequence identity to any of the foregoing. In some aspects, the gRNA further comprises a scaffold sequence comprising SEQ ID NO:44, or a sequence having at least 90% sequence identity to SEQ ID NO:44. In some aspects, the gRNA, including a spacer sequence and a scaffold sequence, comprises SEQ ID NO:55, or a sequence having at least 90% sequence identity to all or a portion thereof. In some aspects, the gRNA is set forth in SEQ ID NO:55. In some aspects, a provided DNA-targeting system comprises any of the above gRNAs complexed with a Cas protein, such as a Cas9 protein. In some aspects, the Cas9 is a dCas9. In some aspects, the dCas9 is a dSaCas9, such as a dSaCas9 set forth in SEQ ID NO:72, or a variant and/or fusion thereof.
[0402] In some aspects, the gRNA targets a target site in a FXN locus that comprises SEQ ID NO: 10, a contiguous portion thereof of at least 14 nucleotides, a complementary sequence of any of the foregoing, or a sequence having at least 90% sequence identity to any of the foregoing. In some aspects, the gRNA comprises a spacer sequence comprising SEQ ID NO:31, a contiguous portion thereof of at least 14 nt, or a sequence having at least 90% sequence identity to any of the foregoing. In some aspects, the gRNA further comprises a scaffold sequence comprising SEQ ID NO:44, or a sequence having at least 90% sequence identity to SEQ ID NO:44. In some aspects, the gRNA, including a spacer sequence and a scaffold sequence, comprises SEQ ID NO:56, or a sequence having at least 90% sequence identity to all or a portion thereof. In some aspects, the gRNA is set forth in SEQ ID NO:56. In some aspects, a provided DNA-targeting system comprises any of the above gRNAs complexed with a Cas protein, such as a Cas9 protein. In some aspects, the Cas9 is a dCas9. In some aspects, the dCas9 is a dSaCas9, such as a dSaCas9 set forth in SEQ ID NO:72, or a variant and/or fusion thereof.
[0403] In some aspects, the gRNA targets a target site in a FXN locus that comprises SEQ ID NO: 21, a contiguous portion thereof of at least 14 nucleotides, a complementary sequence of any of the foregoing, or a sequence having at least 90% sequence identity to any of the foregoing. In some aspects, the gRNA comprises a spacer sequence comprising SEQ ID NO:42, a contiguous portion thereof of at least 14 nt, or a sequence having at least 90% sequence identity to any of the foregoing. In some aspects, the gRNA further comprises a scaffold sequence comprising SEQ ID NO:44, or a sequence having at least 90% sequence identity to SEQ ID NO:44. In some aspects, the gRNA, including a spacer sequence and a scaffold sequence, comprises SEQ ID NO:67, or a sequence having at least 90% sequence identity to all or a portion thereof. In some aspects, the gRNA is set forth in SEQ ID NO:67. In some aspects, a provided DNA-targeting system comprises any of the above gRNAs complexed with a Cas protein, such as a Cas9 protein. In some aspects, the Cas9 is a dCas9. In some aspects, the dCas9 is a dSaCas9, such as a dSaCas9 set forth in SEQ ID NO:72, or a variant and/or fusion thereof.
[0404] In some embodiments, the gRNA targets a target site in a FXN locus or a DNA regulatory element thereof that comprises the sequence selected from any one of SEQ ID NO: 11-20, a contiguous portion thereof of at least 14 nucleotides (e.g., 14, 15, 16, 17, 18, 19, 20, 21, or 22 nucleotides), a complementary sequence of any of the foregoing, or a sequence having at or at least 80%, 85%, 90%,
91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to any of the foregoing. In some embodiments, the gRNA comprises a spacer sequence comprising the sequence selected from any one of SEQ ID NO:32-41, a contiguous portion thereof of at least 14 nt (e.g., 14, 15, 16, 17, 18, 19, 20, 21, or 22 nucleotides), or a sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to any of the foregoing. In some embodiments, the gRNA further comprises a scaffold sequence. In some embodiments, the scaffold sequence comprises the sequence set forth in SEQ ID NO:46, or a sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to SEQ ID NO:46. In some embodiments, the gRNA, including a spacer sequence and a scaffold sequence, comprises the sequence selected from any one of SEQ ID NO:57-66, or a sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to all or a portion thereof. In some embodiments, the gRNA is set forth in the sequence selected from any one of SEQ ID NO:57-66. In some embodiments, a provided DNA-targeting system comprises any of the aforementioned gRNAs complexed with a Cas protein, such as a Cas9 protein. In some embodiments, the Cas9 is a dCas9. In some embodiments, the dCas9 is a dSpCas9, such as a dSpCas9 set forth in SEQ ID NO:78, or a variant and/or fusion thereof.
[0405] In some embodiments, the gRNA targets a target site in a FXN locus or a DNA regulatory element thereof that comprises the sequence selected from any one of SEQ ID NO:256-265, a contiguous portion thereof of at least 14 nucleotides (e.g., 14, 15, 16, 17, 18, 19, 20, 21, or 22 nucleotides), a complementary sequence of any of the foregoing, or a sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to any of the foregoing. In some embodiments, the gRNA further comprises a scaffold sequence. In some embodiments, the scaffold sequence comprises the sequence set forth in SEQ ID NO:46, or a sequence having at or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, 99.9%, or 100% sequence identity to SEQ ID NO:46. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:256, and a scaffold sequence of SEQ ID NO:46. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:257, and a scaffold sequence of SEQ ID NO:46. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:258, and a scaffold sequence of SEQ ID NO:46. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:259, and a scaffold sequence of SEQ ID NO:46. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:260, and a scaffold sequence of SEQ ID NO:46. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:261, and a scaffold sequence of SEQ ID NO:46. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:262, and a scaffold sequence of SEQ ID NO:46. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:263, and a scaffold sequence of SEQ ID NO:46. In some embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:264, and a scaffold sequence of SEQ ID NO:46. In some
embodiments, the gRNA comprises, in 5' to 3' order, a spacer targeting SEQ ID NO:265, and a scaffold sequence of SEQ ID NO:46. In some embodiments, a provided DNA-targeting system comprises any of the aforementioned gRNAs complexed with a Cas protein, such as a Cas9 protein. In some embodiments, the Cas9 is a dCas9. In some embodiments, the dCas9 is a dSpCas9, such as a dSpCas9 set forth in SEQ ID NO: 78, or a variant and/or fusion thereof.
[0406] In some embodiments, the gRNA targets a target site in a FXN locus that comprises SEQ ID NO: 11, a contiguous portion thereof of at least 14 nucleotides, a complementary sequence of any of the foregoing, or a sequence having at least 90% sequence identity to any of the foregoing. In some embodiments, the gRNA comprises a spacer sequence comprising SEQ ID NO:32, a contiguous portion thereof of at least 14 nt, or a sequence having at least 90% sequence identity to any of the foregoing. In some embodiments, the gRNA further comprises a scaffold sequence comprising the sequence set forth in SEQ ID NO:46, or a sequence having at least 90% sequence identity to SEQ ID NO:46. In some embodiments, the gRNA, including a spacer sequence and a scaffold sequence, comprises SEQ ID NO:57, or a sequence having at least 90% sequence identity to all or a portion thereof. In some embodiments, the gRNA is set forth in SEQ ID NO:57. In some embodiments, a provided DNA-targeting system comprises any of the above gRNAs complexed with a Cas protein, such as a Cas9 protein. In some embodiments, the Cas9 is a dCas9. In some embodiments, the dCas9 is a dSpCas9, such as a dSpCas9 set forth in SEQ ID NO:78, or a variant and/or fusion thereof.
[0407] In some embodiments, the gRNA targets a target site in a FXN locus that comprises SEQ ID NO: 12, a contiguous portion thereof of at least 14 nucleotides, a complementary sequence of any of the foregoing, or a sequence having at least 90% sequence identity to any of the foregoing. In some embodiments, the gRNA comprises a spacer sequence comprising SEQ ID NO:33, a contiguous portion thereof of at least 14 nt, or a sequence having at least 90% sequence identity to any of the foregoing. In some embodiments, the gRNA further comprises a scaffold sequence comprising the sequence set forth in SEQ ID NO:46, or a sequence having at least 90% sequence identity to SEQ ID NO:46. In some embodiments, the gRNA, including a spacer sequence and a scaffold sequence, comprises SEQ ID NO:58, or a sequence having at least 90% sequence identity to all or a portion thereof. In some embodiments, the gRNA is set forth in SEQ ID NO:58. In some embodiments, a provided DNA-targeting system comprises any of the above gRNAs complexed with a Cas protein, such as a Cas9 protein. In some embodiments, the Cas9 is a dCas9. In some embodiments, the dCas9 is a dSpCas9, such as a dSpCas9 set forth in SEQ ID NO:78, or a variant and/or fusion thereof.
[0408] In some embodiments, the gRNA targets a target site in a FXN locus that comprises SEQ ID NO: 13, a contiguous portion thereof of at least 14 nucleotides, a complementary sequence of any of the foregoing, or a sequence having at least 90% sequence identity to any of the foregoing. In some embodiments, the gRNA comprises a spacer sequence comprising SEQ ID NO:34, a contiguous portion thereof of at least 14 nt, or a sequence having at least 90% sequence identity to any of the foregoing. In some embodiments, the gRNA further comprises a scaffold sequence comprising the sequence set forth
in SEQ ID NO:46, or a sequence having at least 90% sequence identity to SEQ ID NO:46. In some embodiments, the gRNA, including a spacer sequence and a scaffold sequence, comprises SEQ ID NO:59, or a sequence having at least 90% sequence identity to all or a portion thereof. In some embodiments, the gRNA is set forth in SEQ ID NO:59. In some embodiments, a provided DNA-targeting system comprises any of the above gRNAs complexed with a Cas protein, such as a Cas9 protein. In some embodiments, the Cas9 is a dCas9. In some embodiments, the dCas9 is a dSpCas9, such as a dSpCas9 set forth in SEQ ID NO:78, or a variant and/or fusion thereof.
[0409] In some embodiments, the gRNA targets a target site in a FXN locus that comprises SEQ ID NO: 14, a contiguous portion thereof of at least 14 nucleotides, a complementary sequence of any of the foregoing, or a sequence having at least 90% sequence identity to any of the foregoing. In some embodiments, the gRNA comprises a spacer sequence comprising SEQ ID NO:35, a contiguous portion thereof of at least 14 nt, or a sequence having at least 90% sequence identity to any of the foregoing. In some embodiments, the gRNA further comprises a scaffold sequence comprising the sequence set forth in SEQ ID NO:46, or a sequence having at least 90% sequence identity to SEQ ID NO:46. In some embodiments, the gRNA, including a spacer sequence and a scaffold sequence, comprises SEQ ID NO:60, or a sequence having at least 90% sequence identity to all or a portion thereof. In some embodiments, the gRNA is set forth in SEQ ID NO:60. In some embodiments, a provided DNA-targeting system comprises any of the above gRNAs complexed with a Cas protein, such as a Cas9 protein. In some embodiments, the Cas9 is a dCas9. In some embodiments, the dCas9 is a dSpCas9, such as a dSpCas9 set forth in SEQ ID NO:78, or a variant and/or fusion thereof.
[0410] In some embodiments, the gRNA targets a target site in a FXN locus that comprises SEQ ID NO: 15, a contiguous portion thereof of at least 14 nucleotides, a complementary sequence of any of the foregoing, or a sequence having at least 90% sequence identity to any of the foregoing. In some embodiments, the gRNA comprises a spacer sequence comprising SEQ ID NO:36, a contiguous portion thereof of at least 14 nt, or a sequence having at least 90% sequence identity to any of the foregoing. In some embodiments, the gRNA further comprises a scaffold sequence comprising the sequence set forth in SEQ ID NO:46, or a sequence having at least 90% sequence identity to SEQ ID NO:46. In some embodiments, the gRNA, including a spacer sequence and a scaffold sequence, comprises SEQ ID NO:61, or a sequence having at least 90% sequence identity to all or a portion thereof. In some embodiments, the gRNA is set forth in SEQ ID NO:61. In some embodiments, a provided DNA-targeting system comprises any of the above gRNAs complexed with a Cas protein, such as a Cas9 protein. In some embodiments, the Cas9 is a dCas9. In some embodiments, the dCas9 is a dSpCas9, such as a dSpCas9 set forth in SEQ ID NO:78, or a variant and/or fusion thereof.
[0411] In some embodiments, the gRNA targets a target site in a FXN locus that comprises SEQ ID NO: 16, a contiguous portion thereof of at least 14 nucleotides, a complementary sequence of any of the foregoing, or a sequence having at least 90% sequence identity to any of the foregoing. In some embodiments, the gRNA comprises a spacer sequence comprising SEQ ID NO:37, a contiguous portion
thereof of at least 14 nt, or a sequence having at least 90% sequence identity to any of the foregoing. In some embodiments, the gRNA further comprises a scaffold sequence comprising the sequence set forth in SEQ ID NO:46, or a sequence having at least 90% sequence identity to SEQ ID NO:46. In some embodiments, the gRNA, including a spacer sequence and a scaffold sequence, comprises SEQ ID NO:62, or a sequence having at least 90% sequence identity to all or a portion thereof. In some embodiments, the gRNA is set forth in SEQ ID NO:62. In some embodiments, a provided DNA-targeting system comprises any of the above gRNAs complexed with a Cas protein, such as a Cas9 protein. In some embodiments, the Cas9 is a dCas9. In some embodiments, the dCas9 is a dSpCas9, such as a dSpCas9 set forth in SEQ ID NO:78, or a variant and/or fusion thereof.
[0412] In some embodiments, the gRNA targets a target site in a FXN locus that comprises SEQ ID NO: 17, a contiguous portion thereof of at least 14 nucleotides, a complementary sequence of any of the foregoing, or a sequence having at least 90% sequence identity to any of the foregoing. In some embodiments, the gRNA comprises a spacer sequence comprising SEQ ID NO:38, a contiguous portion thereof of at least 14 nt, or a sequence having at least 90% sequence identity to any of the foregoing. In some embodiments, the gRNA further comprises a scaffold sequence comprising the sequence set forth in SEQ ID NO:46, or a sequence having at least 90% sequence identity to SEQ ID NO:46. In some embodiments, the gRNA, including a spacer sequence and a scaffold sequence, comprises SEQ ID NO:63, or a sequence having at least 90% sequence identity to all or a portion thereof. In some embodiments, the gRNA is set forth in SEQ ID NO:63. In some embodiments, a provided DNA-targeting system comprises any of the above gRNAs complexed with a Cas protein, such as a Cas9 protein. In some embodiments, the Cas9 is a dCas9. In some embodiments, the dCas9 is a dSpCas9, such as a dSpCas9 set forth in SEQ ID NO:78, or a variant and/or fusion thereof.
[0413] In some embodiments, the gRNA targets a target site in a FXN locus that comprises SEQ ID NO: 18, a contiguous portion thereof of at least 14 nucleotides, a complementary sequence of any of the foregoing, or a sequence having at least 90% sequence identity to any of the foregoing. In some embodiments, the gRNA comprises a spacer sequence comprising SEQ ID NO:39, a contiguous portion thereof of at least 14 nt, or a sequence having at least 90% sequence identity to any of the foregoing. In some embodiments, the gRNA further comprises a scaffold sequence comprising the sequence set forth in SEQ ID NO:46, or a sequence having at least 90% sequence identity to SEQ ID NO:46. In some embodiments, the gRNA, including a spacer sequence and a scaffold sequence, comprises SEQ ID NO:64, or a sequence having at least 90% sequence identity to all or a portion thereof. In some embodiments, the gRNA is set forth in SEQ ID NO:64. In some embodiments, a provided DNA-targeting system comprises any of the above gRNAs complexed with a Cas protein, such as a Cas9 protein. In some embodiments, the Cas9 is a dCas9. In some embodiments, the dCas9 is a dSpCas9, such as a dSpCas9 set forth in SEQ ID NO:78, or a variant and/or fusion thereof.
[0414] In some embodiments, the gRNA targets a target site in a FXN locus that comprises SEQ ID
NO: 19, a contiguous portion thereof of at least 14 nucleotides, a complementary sequence of any of the
foregoing, or a sequence having at least 90% sequence identity to any of the foregoing. In some embodiments, the gRNA comprises a spacer sequence comprising SEQ ID NO:40, a contiguous portion thereof of at least 14 nt, or a sequence having at least 90% sequence identity to any of the foregoing. In some embodiments, the gRNA further comprises a scaffold sequence comprising the sequence set forth in SEQ ID NO:46, or a sequence having at least 90% sequence identity to SEQ ID NO:46. In some embodiments, the gRNA, including a spacer sequence and a scaffold sequence, comprises SEQ ID NO:65, or a sequence having at least 90% sequence identity to all or a portion thereof. In some embodiments, the gRNA is set forth in SEQ ID NO:65. In some embodiments, a provided DNA-targeting system comprises any of the above gRNAs complexed with a Cas protein, such as a Cas9 protein. In some embodiments, the Cas9 is a dCas9. In some embodiments, the dCas9 is a dSpCas9, such as a dSpCas9 set forth in SEQ ID NO:78, or a variant and/or fusion thereof.
[0415] In some embodiments, the gRNA targets a target site in a FXN locus that comprises SEQ ID NO:20, a contiguous portion thereof of at least 14 nucleotides, a complementary sequence of any of the foregoing, or a sequence having at least 90% sequence identity to any of the foregoing. In some embodiments, the gRNA comprises a spacer sequence comprising SEQ ID NO:41, a contiguous portion thereof of at least 14 nt, or a sequence having at least 90% sequence identity to any of the foregoing. In some embodiments, the gRNA further comprises a scaffold sequence comprising the sequence set forth in SEQ ID NO:46, or a sequence having at least 90% sequence identity to SEQ ID NO:46. In some embodiments, the gRNA, including a spacer sequence and a scaffold sequence, comprises SEQ ID NO:66, or a sequence having at least 90% sequence identity to all or a portion thereof. In some embodiments, the gRNA is set forth in SEQ ID NO:66. In some embodiments, a provided DNA-targeting system comprises any of the above gRNAs complexed with a Cas protein, such as a Cas9 protein. In some embodiments, the Cas9 is a dCas9. In some embodiments, the dCas9 is a dSpCas9, such as a dSpCas9 set forth in SEQ ID NO:78, or a variant and/or fusion thereof.
2. Combinations of gRNAs
[0416] Also provided herein are combinations, including combinations of two or more guide RNAs (gRNAs). In some aspects, the provided combination of gRNAs include two or more gRNAs, each of which target particular regions of a frataxin (FXN) locus. In some aspects, the two or more gRNAs each comprise any of the gRNAs described herein.
[0417] Also provided herein is a combination, comprising a first gRNA comprising any of the gRNAs described herein, and one or more second gRNAs that binds to a second target site in a regulatory DNA element of a frataxin (FXN) locus. In some embodiments, the second gRNA comprises any of the gRNAs described herein.
[0418] Also provided herein is a combination, comprising: a first gRNA that binds a first target site in an enhancer region of a frataxin (FXN) locus, wherein the first target site is located within the genomic coordinates human genome assembly GRCh38 (hg38) chr9:69, 027, 282-69, 028, 497; and a second gRNA
that binds a second target site in a promoter region of a FXN locus, wherein the second target site is located within the genomic coordinates hg38 chr9:68, 940, 179-69, 205 ,519.
[0419] In some embodiments, the first gRNA comprises a gRNA spacer sequence set forth in SEQ ID NO:42 or a contiguous portion thereof of at least 14 nt. In some embodiments, the second gRNA comprises a gRNA spacer sequence set forth in any one of SEQ ID NO:22-31 or a contiguous portion thereof of at least 14 nt. In some embodiments, the second gRNA comprises a gRNA spacer sequence set forth in SEQ ID NO:22 or 28 or a contiguous portion thereof of at least 14 nt.
[0420] In some of any of the provided embodiments, e.g., of a combination provided herein: the first gRNA comprises a gRNA spacer sequence set forth in SEQ ID NO:42 or a contiguous portion thereof of at least 14 nt; and the second gRNA comprises a gRNA spacer sequence set forth in any one of SEQ ID NO:22-31 or a contiguous portion thereof of at least 14 nt. In some of any of the provided embodiments, e.g., of a combination provided herein: the first gRNA comprises a gRNA spacer sequence set forth in SEQ ID NO:42 or a contiguous portion thereof of at least 14 nt; and the second gRNA comprises a gRNA spacer sequence set forth in SEQ ID NO:22 or a contiguous portion thereof of at least 14 nt. In some of any of the provided embodiments, e.g., of a combination provided herein: the first gRNA comprises a gRNA spacer sequence set forth in SEQ ID NO:42 or a contiguous portion thereof of at least 14 nt; and the second gRNA comprises a gRNA spacer sequence set forth in SEQ ID NO:28 or a contiguous portion thereof of at least 14 nt.
[0421] Also provided herein is a combination, comprising: a first gRNA that binds a first target site in a promoter region of a FXN locus, wherein the second target site is located within the genomic coordinates hg38 chr9:68,940, 179-69,205,519; and a second gRNA that binds a second target site in a promoter region of a frataxin (FXN) locus, wherein the first target site is located within the genomic coordinates hg38 chr9:68,940, 179-69,205,519.
[0422] In some embodiments, the first gRNA comprises a gRNA spacer sequence set forth in any one of SEQ ID NOs:22-31 or a contiguous portion thereof of at least 14 nt. In some embodiments, the first gRNA comprises a gRNA spacer sequence set forth in SEQ ID NO:22 or 28 or a contiguous portion thereof of at least 14 nt. In some embodiments, the second gRNA comprises a gRNA spacer sequence set forth in any one of SEQ ID NOs:22-31 or a contiguous portion thereof of at least 14 nt. In some embodiments, the second gRNA comprises a gRNA spacer sequence set forth in SEQ ID NO:22 or 28 or a contiguous portion thereof of at least 14 nt.
[0423] In some embodiments, the combination comprises: the first gRNA comprises a gRNA spacer sequence set forth in any one of SEQ ID NOs:22-31 or a contiguous portion thereof of at least 14 nt; and the second gRNA comprises a gRNA spacer sequence set forth in any one of SEQ ID NOs:22-31 or a contiguous portion thereof of at least 14 nt.
[0424] In some embodiments, the combination comprises: the first gRNA comprises a gRNA spacer sequence set forth in SEQ ID NO:22 or a contiguous portion thereof of at least 14 nt; and the second gRNA comprises a gRNA spacer sequence set forth in SEQ ID NO:28 or a contiguous portion thereof of
at least 14 nt.
[0425] In some embodiments, the first gRNA comprises a gRNA spacer sequence set forth in any one of SEQ ID NOs:32-41 or a contiguous portion thereof of at least 14 nt. In some embodiments, the second gRNA comprises a gRNA spacer sequence set forth in any one of SEQ ID NOs:32-41 or a contiguous portion thereof of at least 14 nt. In some embodiments, the combination comprises: the first gRNA comprises a gRNA spacer sequence set forth in any one of SEQ ID NOs:32-41 or a contiguous portion thereof of at least 14 nt; and the second gRNA comprises a gRNA spacer sequence set forth in any one of SEQ ID NOs:32-41 or a contiguous portion thereof of at least 14 nt.
3. Fusion Proteins
[0426] Provided are fusion proteins that include (1) a DNA-targeting domain or a component thereof and (2) at least one effector domain. In some aspects, the DNA-targeting domain or component thereof (e.g., a protein or polypeptide component of the DNA-targeting domain, such as the eZFP of the eZFP fusion protein, or the Cas component of the Cas-gRNA combination) can be any described herein. In some aspects, the at least one effector domain can be any described herein, for example, in Section II.B. In some aspects, the fusion protein is targeted to a target site, for example, one or more target sites at a FXN locus, such as any target site described herein, such as in Section I or Section II, by the DNA- targeting domain. In some embodiments, a fusion protein comprising an eZFP can be referred to herein as an eZFP fusion protein. In some aspects, a fusion protein comprising a dCas protein can be referred to herein as a dCas fusion protein.
[0427] Provided are fusion proteins that include (1) a DNA-targeting domain or a component thereof and (2) at least one effector domain, wherein: the DNA-targeting domain or a component thereof binds to a target site in a regulatory DNA element of a frataxin (FXN) locus; and the effector domain induces, catalyzes or leads to transcription activation, transcription co-activation, transcription elongation, transcription de-repression, transcription factor release, polymerization, histone modification, histone acetylation, histone deacetylation, nucleosome remodeling, chromatin remodeling, reversal of heterochromatin formation, nuclease, signal transduction, proteolysis, ubiquitination, deubiquitination, phosphorylation, dephosphorylation, splicing, nucleic acid association, DNA methylation, DNA demethylation, histone methylation, histone demethylation, or DNA base oxidation. In some embodiments, the fusion protein comprises any of the effector domains described herein.
[0428] In some embodiments, binding of the DNA-targeting domain or a component thereof to the target site does not introduce a genetic disruption or a DNA break at or near the target site. In some embodiments, the DNA-targeting domain comprises a Clustered Regularly Interspaced Short Palindromic Repeats associated (Cas)-guide RNA (gRNA) combination comprising (a) a Cas protein or a variant thereof and (b) at least one gRNA; a zinc finger protein (ZFP), such as an eZFP; a transcription activator-like effector (TAEE); a meganuclease; a homing endonuclease; or a I-Scel enzymes or a variant thereof, such as a catalytically inactive variant thereof. In some embodiments, the DNA-targeting domain
comprises an eZFP. In some embodiments, the DNA-targeting domain comprises a Cas-gRNA combination comprising a Cas protein or a variant thereof and at least one gRNA, and the component of the DNA-targeting domain is a Cas protein or a variant thereof. In some embodiments, the variant Cas protein lacks nuclease activity or is a deactivated Cas (dCas) protein.
[0429] In some embodiments, the gRNA is capable of complexing with the Cas protein or variant thereof. In some embodiments, the Cas protein or a variant thereof is a Cas9 protein or a variant thereof. In some embodiments, the variant Cas protein is a variant Cas9 protein that lacks nuclease activity or that is a deactivated Cas9 (dCas9) or a nuclease-inactive Cas9 (iCas9) protein. In some aspects, the dCas9 or iCas9 component of the fusion protein includes any described herein.
[0430] In some embodiments, the Cas9 protein or a variant thereof is a Staphylococcus aureus Cas9 (SaCas9) protein or a variant thereof. In some embodiments, the variant Cas9 is a Staphylococcus aureus dCas9 protein (dSaCas9) that comprises at least one amino acid mutation selected from D10A and N580A, with reference to numbering of positions of SEQ ID NO:73. In some embodiments, the variant Cas9 protein comprises the sequence set forth in SEQ ID NO:72, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
[0431] In some embodiments, the Cas9 protein or variant thereof is a Streptococcus pyogenes Cas9 (SpCas9) protein or a variant thereof. In some embodiments, the variant Cas9 is a Streptococcus pyogenes dCas9 (dSpCas9) protein that comprises at least one amino acid mutation selected from D10A and H840A, with reference to numbering of positions of SEQ ID NO:79. In some embodiments, the variant Cas9 protein comprises the sequence set forth in SEQ ID NO:78, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
[0432] In some embodiments, the DNA-targeting domain of the fusion protein is an engineered zinc finger protein (eZFP); a transcription activator-like effector (TALE); a meganuclease; a homing endonuclease; or a I-Scel enzymes or a variant thereof, such as a catalytically inactive variant thereof. In some embodiments, the DNA-targeting domain of the fusion protein is an eZFP, such as any eZFP described herein, for example in Section I. In some aspects, the DNA-targeting domain of the fusion protein is targeted to one or more target sites at a FXN locus, such as one or more target sites described herein, for example, in Sections I and II. In some aspects, the DNA-targeting domain of the fusion protein is an engineered zinc finger protein (eZFP); a transcription activator-like effector (TALE); a meganuclease; a homing endonuclease; or a I-Scel enzymes or a variant thereof that is capable of binding to a target site at a FXN locus described herein, in a sequence-specific manner.
[0433] In some embodiments, the regulatory DNA element is an enhancer. In some embodiments, the target site is located within the genomic coordinates human genome assembly GRCh38 (hg38) chr9:69, 027, 282-69, 028, 497. In some embodiments, the target site is located within the genomic coordinates hg38 chr9:69, 027, 615-69, 028, 101. In some embodiments, the target site comprises the sequence set forth in SEQ ID NO:21, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing.
[0434] In some embodiments, the regulatory DNA element is a promoter. In some embodiments, the target site is located within the genomic coordinates hg38 chr9:68, 940, 179-69, 205 ,519. In some embodiments, the target site comprises a sequence selected from any one of SEQ ID NOS: 1-10, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing. In some embodiments, the target site comprises a sequence selected from any one of SEQ ID NOS: 11-20, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing.
[0435] In some embodiments, the target site is any target site provided herein, such as any target site provided in Section I or II. In some embodiments, the target site is a target site for an eZFP or eZFP fusion protein, such as any target site provided in Section I or II.
[0436] In some embodiments, the effector domain induces, catalyzes or leads to transcription activation, transcription co-activation, transcription elongation. In some embodiments, the effector domain induces transcription activation. In some embodiments, the effector domain comprises at least one VP16 domain, or a VP16 tetramer (“VP64”) or a variant thereof. In some embodiments, the effector domain comprises the sequence set forth in SEQ ID NO:81 or 83, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
[0437] In some embodiments, the effector domain is selected from a p65 activation domain, a p300 domain, DPOLA, ENL, FOXO3, HSH2D, NCOA2, NCOA3, PSA1, PYGO1, RBM39, HERC2, or NOTCH2, or a domain thereof, a portion thereof or a variant thereof. In some embodiments, the effector domain comprises a sequence selected from any one of SEQ ID NOS: 113-125, or a domain thereof, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing. In some embodiments, the effector domain comprises a sequence selected from any one of SEQ ID NOS: 100-112, or a domain thereof, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing. In some embodiments, the variant thereof comprises a truncation thereof. In some aspects, the effector domain comprises any one of the effector domains described herein.
[0438] In some embodiments, the at least one effector domain is fused to the N-terminus, the C- terminus, or both the N-terminus and the C-terminus, of the DNA-targeting domain or a component thereof (such as a protein or polypeptide component thereof, for example, a Cas component of a Cas- gRNA combination).
[0439] In some embodiments, the at least one effector domain is fused to the N-terminus, the C- terminus, or both the N-terminus and the C-terminus, of the DNA-targeting domain or a component thereof. In some embodiments, the DNA-targeting system also includes one or more linkers connecting the DNA-targeting domain or a component thereof to the at least one effector domain. In some embodiments, the DNA-targeting system further comprises one or more nuclear localization signals (NLS).
[0440] In some embodiments, the fusion protein comprises the sequence set forth in any one of SEQ ID NOs:85 and 159-173, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the NLS comprises the sequence set forth in any one of SEQ ID NOs:85 and 159-173, or a portion thereof. In some embodiments, the NLS comprises the sequence set forth in SEQ ID NO: 85 or a portion thereof. An exemplary nucleotide sequence encoding the NLS set forth in SEQ ID NO: 85 is set forth in SEQ ID NO:84.
[0441] In some embodiments, the fusion protein comprises the sequence set forth in SEQ ID NO:77, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
[0442] In some embodiments, the fusion protein further comprises one or more linkers connecting the DNA-targeting domain or a component thereof to the at least one effector domain, and/or further comprises one or more nuclear localization signals (NLS).
[0443] In some embodiments, the fusion protein comprises the sequence set forth in SEQ ID NO:71, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the fusion protein comprises the sequence set forth in SEQ ID NO:77, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
[0444] In some embodiments, the fusion protein includes at least one linker. A linker may be included anywhere in the polypeptide sequence of the fusion protein, for example, between the effector domain and the DNA-targeting domain or a component thereof. A linker may be of any length and designed to promote or restrict the mobility of components in the fusion protein.
[0445] A linker may comprise any amino acid sequence of about 2 to about 100, about 5 to about 80, about 10 to about 60, or about 20 to about 50 amino acids. A linker may comprise an amino acid sequence of at least about 2, 3, 4, 5, 10, 15, 20, 25, 30, 35, 40, 45, 50, 55, 60, 65, 70, 75, 80 or 85 amino acids. A linker may comprise an amino acid sequence of less than about 100, 90, 80, 70, 60, 50, or 40 amino acids. A linker may include sequential or tandem repeats of an amino acid sequence that is 2 to 20 amino acids in length. Linkers may be rich in amino acids glycine (G), serine (S), and/or alanine (A). Linkers may include, for example, a GS linker such as (Gly-Gly-Gly-Gly-Ser)n. An exemplary GS linker is represented by the sequence GGGGS (SEQ ID NO: 158). A linker may comprise repeats of a sequence, for example as represented by the formula (GGGGS)n, wherein n is an integer that represents the number of times the GGGGS sequence is repeated (e.g. between 1 and 10 times). The number of times a linker sequence is repeated, for example n in a GS linker, can be adjusted to optimize the linker length and achieve appropriate separation of the functional domains. Other examples of linkers may include, for example, Gly-Gly-Gly-Gly-Gly (SEQ ID NO: 154), Gly-Gly-Ala-Gly-Gly (SEQ ID NO: 155), Gly/Ser rich linkers such as Gly-Gly-Gly-Gly-Ser-Ser-Ser (SEQ ID NO: 156), or Gly/Ala rich linkers such as Gly-Gly-Gly-Gly-Ala-Ala-Ala (SEQ ID NO: 157), or Gly-Ser-Gly-Ser-Gly (SEQ ID NO:219).
[0446] In some embodiments, the linker is an XTEN linker. In some aspects, an XTEN linker is a recombinant polypeptide (e.g., an unstructured recombinant peptide) lacking hydrophobic amino acid residues. Exemplary XTEN linkers are described in, for example, Schellenberger et al., Nature Biotechnology 27, 1186-1190 (2009) or WO 2021/247570. In some embodiments, an exemplary linker comprises a linker described in WO 2021/247570. In some aspects, the linker is or comprises the sequence set forth in SEQ ID NO: 186 or SEQ ID NO: 174, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing. In some embodiments, the linker comprises the sequence set forth in SEQ ID NO:186, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing. In some aspects, the linker comprises the sequence set forth in SEQ ID NO: 186, or a contiguous portion of SEQ ID NO: 186 of at least 5, 10, 15, 20, 25, 30, 35, 40, 45, 50, 55, 60, 65, 70 or 75 amino acids. In some aspects, the linker consists of the sequence set forth in SEQ ID NO: 186, or a contiguous portion of SEQ ID NO: 186 of at least 5, 10, 15, 20, 25, 30, 35, 40, 45, 50, 55, 60, 65, 70 or 75 amino acids. In some embodiments, the linker comprises the sequence set forth in SEQ ID NO: 186. In some embodiments, the linker consists of the sequence set forth in SEQ ID NO: 186. In some embodiments, the linker comprises the sequence set forth in SEQ ID NO: 174, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing. In some aspects, the linker comprises the sequence set forth in SEQ ID NO: 174, or a contiguous portion of SEQ ID NO: 174 of at least 5, 10, or 15 amino acids. In some aspects, the linker consists of the sequence set forth in SEQ ID NO: 174, or a contiguous portion of SEQ ID NO: 174 of at least 5, 10, or 15 amino acids. In some embodiments, the linker comprises the sequence set forth in SEQ ID NO: 174. In some embodiments, the linker consists of the sequence set forth in SEQ ID NO: 174. Appropriate linkers may be selected or designed based rational criteria known in the art, for example as described in Chen et al. Adv. Drug Deliv. Rev. 65(10): 1357-1369 (2013). In some embodiments, a linker comprises the sequence set forth in SEQ ID NO: 188, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing. In some embodiments, a linker comprising the sequence set forth in SEQ ID NO: 188 is encoded by the nucleotide sequence set forth in SEQ ID NO: 187.
[0447] In some embodiments, a fusion protein described herein comprises one or more nuclear localization sequences (NLSs), such as about or more than about 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, or more NLSs. When more than one NLS is present, each may be selected independently of the others, such that a single NLS may be present in more than one copy and/or in combination with one or more other NLSs present in one or more copies. Non-limiting examples of NLSs include an NLS sequence derived from: the NLS of the SV40 virus large T-antigen, having the sequence PKKKRKV (SEQ ID NO: 159); the NLS from nucleoplasmin (e.g. the nucleoplasmin bipartite NLS) having the sequence KRPAATKKAGQAKKKK (SEQ ID NO:85); the c-myc NLS having the sequence PAAKRVKLD
(SEQ ID NO: 160) or RQRRNELKRSP (SEQ ID NO: 161); the hRNPAl M9 NLS having the sequence NQSSNFGPMKGGNFGGRSSGPYGGGGQYFAKPRNQGGY (SEQ ID NO: 162); the sequence RMRIZFKNKGKDTAELRRRRVEVSVELRKAKKDEQILKRRNV (SEQ ID NO: 163) of the IBB domain from importin-alpha; the sequences VSRKRPRP (SEQ ID NO: 164) and PPKKARED (SEQ ID NO: 165) of the myoma T protein; the sequence PQPKKKPL (SEQ ID NO: 166) of human p53; the sequence SALIKKKKKMAP (SEQ ID NO: 167) of mouse c-abl IV; the sequences DRLRR (SEQ ID NO: 168) and PKQKKRK (SEQ ID NO: 169) of the influenza virus NS1; the sequence RKLKKKIKKL (SEQ ID NO: 170) of the Hepatitis virus delta antigen; the sequence REKKKFLKRR (SEQ ID NO: 171) of the mouse Mxl protein; the sequence KRKGDEVDGVDEVAKKKSKK (SEQ ID NO: 172) of the human poly(ADP-ribose) polymerase; and the sequence RKCLQAGMNLEARKTKK (SEQ ID NO: 173) of the steroid hormone receptors (human) glucocorticoid. In general, the one or more NLSs are of sufficient strength to drive accumulation of the fusion protein in a detectable amount in the nucleus of a eukaryotic cell. In general, strength of nuclear localization activity may derive from the number of NLSs in the fusion protein, the particular NLS(s) used, or a combination of these factors. Detection of accumulation in the nucleus may be performed by any suitable technique. For example, a detectable marker may be fused to the fusion protein, such that location within a cell may be visualized, such as in combination with a means for detecting the location of the nucleus (e.g. a stain specific for the nucleus such as DAPI). Cell nuclei may also be isolated from cells, the contents of which may then be analyzed by any suitable process for detecting protein, such as immunohistochemistry, Western blot, or enzyme activity assay. Accumulation in the nucleus may also be determined indirectly, such as by an assay for the effect of the fusion protein (e.g. an assay for altered gene expression activity in a cell transformed with the DNA-targeting system comprising the fusion protein), as compared to a control condition (e.g. an untransformed cell).
[0448] In some embodiments, the NLS comprises the sequence set forth in any one of SEQ ID NO:85 and 160-173, or a portion thereof.
[0449] In some aspects, provided are DNA-targeting systems or fusion proteins that comprise a Cas protein or a variant thereof and at least one effector domain, wherein the effector domain increases transcription of the FXN locus.
In some embodiments, the at least one effector domain is fused to the N-terminus, the C-terminus, or both the N-terminus and the C-terminus, of the DNA-targeting domain or a component thereof (such as a protein or polypeptide component thereof, for example, a Cas component of a Cas-gRNA combination). In some embodiments, the DNA-targeting system also includes one or more linkers connecting the DNA- targeting domain or a component thereof to the at least one effector domain, and/or further comprising one or more nuclear localization signals (NLS).
[0450] In some aspects, the DNA-targeting system or fusion protein comprises one or more tags, linkers and/or NLS sequences. In some embodiments, exemplary tags, linkers and/or NLS sequences can be any described herein.
In some cases, sequences provided herein, including amino acid sequences for the DNA-targeting systems or fusion proteins provided herein, contain sequences of one or more tags, linkers and/or NLS sequences. In some aspects, it is understood that the exemplary tags, linkers and/or NLS sequences are not required or are not the sole or exclusive tags, linkers and/or NLS sequences that can be employed in the DNA-targeting systems or fusion proteins. In some aspects, sequences containing tags, linkers and/or NLS sequences are exemplary, and are not limited to the specific tags, linkers and/or NLS sequences contained in the described sequences. In some aspects, alternative tags, linkers and/or NLS sequences can be can be employed in the DNA-targeting systems or fusion proteins, or the DNA-targeting system or fusion protein in some cases does not contain or lacks a tag, linker and/or NLS. In some aspects, alternative tags, linkers and/or NLS sequences include other known tags, linkers and/or NLS sequences that have similar function or serve similar purposes.
[0451] In some embodiments, the DNA-targeting system or fusion protein comprises the sequence set forth in SEQ ID NO:71, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
In some embodiments, the DNA-targeting system or fusion protein comprises the sequence set forth in SEQ ID NO:71.
[0452] In some embodiments, the DNA-targeting system or fusion protein comprises the sequence set forth in SEQ ID NO:77, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the DNA-targeting system or fusion protein comprises the sequence set forth in SEQ ID NO: 77. In some embodiments, an exemplary nucleotide sequence encoding the DNA-targeting system or fusion protein set forth in SEQ ID NO:77 is set forth in SEQ ID NO:75.
In some embodiments, the DNA-targeting system or fusion protein comprises, from N- to C-terminal order: a dSaCas9 set forth in SEQ ID NO:72, a linker and/or NLS, and a DPOLA domain set forth in SEQ ID NO: 100. In some embodiments, the DNA-targeting system or fusion protein comprises, from N- to C-terminal order: a dSaCas9 set forth in SEQ ID NO:72, a linker and/or NLS, and a ENL domain set forth in SEQ ID NO: 101. In some embodiments, the DNA-targeting system or fusion protein comprises, from N- to C-terminal order: a dSaCas9 set forth in SEQ ID NO:72, a linker and/or NLS, and a FOXO3 domain set forth in SEQ ID NO: 102. In some embodiments, the DNA-targeting system or fusion protein comprises, from N- to C-terminal order: a dSaCas9 set forth in SEQ ID NO:72, a linker and/or NLS, and a HSH2D domain set forth in SEQ ID NO: 103. In some embodiments, the DNA-targeting system or fusion protein comprises, from N- to C-terminal order: a dSaCas9 set forth in SEQ ID NO:72, a linker and/or NLS, and a NCOA2 domain set forth in SEQ ID NO: 104. In some embodiments, the DNA- targeting system or fusion protein comprises, from N- to C-terminal order: a dSaCas9 set forth in SEQ ID NO:72, a linker and/or NLS, and a NCOA3 domain set forth in SEQ ID NO: 105. In some embodiments, the DNA-targeting system or fusion protein comprises, from N- to C-terminal order: a dSaCas9 set forth in SEQ ID NO:72, a linker and/or NLS, and a PSA1 domain set forth in SEQ ID NO: 106. In some
embodiments, the DNA-targeting system or fusion protein comprises, from N- to C-terminal order: a dSaCas9 set forth in SEQ ID NO:72, a linker and/or NLS, and a PYG01 domain set forth in SEQ ID NO: 107. In some embodiments, the DNA-targeting system or fusion protein comprises, from N- to C- terminal order: a dSaCas9 set forth in SEQ ID NO:72, a linker and/or NLS, and a RBM39 domain set forth in SEQ ID NO: 108. In some embodiments, the DNA-targeting system or fusion protein comprises, from N- to C-terminal order: a dSaCas9 set forth in SEQ ID NO:72, a linker and/or NLS, and a HERC2 domain set forth in SEQ ID NO: 109. In some embodiments, the DNA-targeting system or fusion protein comprises, from N- to C-terminal order: a dSaCas9 set forth in SEQ ID NO:72, a linker and/or NLS, and a DMD domain set forth in SEQ ID NO: 110. An exemplary nucleotide encoding the DMD domain set forth in SEQ ID NO: 110 is set forth in SEQ ID NO:97. In some embodiments, the DNA-targeting system or fusion protein comprises, from N- to C-terminal order: a dSaCas9 set forth in SEQ ID NO:72, a linker and/or NLS, and a NOTCH2 domain set forth in SEQ ID NO: 111. In some embodiments, the DNA- targeting system or fusion protein comprises, from N- to C-terminal order: a dSaCas9 set forth in SEQ ID NO:72, a linker and/or NLS, and a p300 core domain set forth in SEQ ID NO: 112.
[0453] In some embodiments, the DNA-targeting system or fusion protein comprises, from N- to C- terminal order: a DPOLA domain set forth in SEQ ID NO: 100, a linker and/or NLS, and a dSaCas9 set forth in SEQ ID NO:72. In some embodiments, the DNA-targeting system or fusion protein comprises, from N- to C-terminal order: a dSaCas9 set forth in SEQ ID NO:72, a linker and/or NLS, and a ENL domain set forth in SEQ ID NO: 101, a linker and/or NLS, and a dSaCas9 set forth in SEQ ID NO:72. In some embodiments, the DNA-targeting system or fusion protein comprises, from N- to C-terminal order: a dSaCas9 set forth in SEQ ID NO:72, a linker and/or NLS, and a FOXO3 domain set forth in SEQ ID NO: 102, a linker and/or NLS, and a dSaCas9 set forth in SEQ ID NO:72. In some embodiments, the DNA-targeting system or fusion protein comprises, from N- to C-terminal order: a dSaCas9 set forth in SEQ ID NO:72, a linker and/or NLS, and a HSH2D domain set forth in SEQ ID NO: 103, a linker and/or NLS, and a dSaCas9 set forth in SEQ ID NO:72. In some embodiments, the DNA-targeting system or fusion protein comprises, from N- to C-terminal order: a dSaCas9 set forth in SEQ ID NO:72, a linker and/or NLS, and a NCOA2 domain set forth in SEQ ID NO: 104, a linker and/or NLS, and a dSaCas9 set forth in SEQ ID NO:72. In some embodiments, the DNA-targeting system or fusion protein comprises, from N- to C-terminal order: a dSaCas9 set forth in SEQ ID NO:72, a linker and/or NLS, and a NCOA3 domain set forth in SEQ ID NO: 105, a linker and/or NLS, and a dSaCas9 set forth in SEQ ID NO:72. In some embodiments, the DNA-targeting system or fusion protein comprises, from N- to C-terminal order: a dSaCas9 set forth in SEQ ID NO:72, a linker and/or NLS, and a PSA1 domain set forth in SEQ ID NO: 106, a linker and/or NLS, and a dSaCas9 set forth in SEQ ID NO:72. In some embodiments, the DNA-targeting system or fusion protein comprises, from N- to C-terminal order: a dSaCas9 set forth in SEQ ID NO:72, a linker and/or NLS, and a PYGO1 domain set forth in SEQ ID NO: 107, a linker and/or NLS, and a dSaCas9 set forth in SEQ ID NO:72. In some embodiments, the DNA-targeting system or fusion protein comprises, from N- to C-terminal order: a dSaCas9 set forth in SEQ ID NO:72, a linker
and/or NLS, and a RBM39 domain set forth in SEQ ID NO: 108, a linker and/or NLS, and a dSaCas9 set forth in SEQ ID NO:72. In some embodiments, the DNA-targeting system or fusion protein comprises, from N- to C-terminal order: a dSaCas9 set forth in SEQ ID NO:72, a linker and/or NLS, and a HERC2 domain set forth in SEQ ID NO: 109, a linker and/or NLS, and a dSaCas9 set forth in SEQ ID NO:72. In some embodiments, the DNA-targeting system or fusion protein comprises, from N- to C-terminal order: a dSaCas9 set forth in SEQ ID NO:72, a linker and/or NLS, and a DMD domain set forth in SEQ ID NO: 110, a linker and/or NLS, and a dSaCas9 set forth in SEQ ID NO:72. In some embodiments, the DNA-targeting system or fusion protein comprises, from N- to C-terminal order: a dSaCas9 set forth in SEQ ID NO:72, a linker and/or NLS, and a NOTCH2 domain set forth in SEQ ID NO: 111 , a linker and/or NLS, and a dSaCas9 set forth in SEQ ID NO:72. In some embodiments, the DNA-targeting system or fusion protein comprises, from N- to C-terminal order: a dSaCas9 set forth in SEQ ID NO:72, a linker and/or NLS, and a p300 core domain set forth in SEQ ID NO: 112, a linker and/or NLS, and a dSaCas9 set forth in SEQ ID NO:72.
In some embodiments, the DNA-targeting system or fusion protein, comprises a dSaCas9 set forth in SEQ ID NO:72, and any of the epigenetic effector domains and/or multipartite effectors described herein, such as in Section II.B. In some embodiments, the DNA-targeting system or fusion protein comprises the sequence set forth in any one of SEQ ID NOS:266-268 and 315-319, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the DNA-targeting system or fusion protein comprises the sequence set forth in any one of SEQ ID NOS:266-268 and 315-319.
[0454] In some aspects, exemplary linkers or NLS sequences can be any described herein.
4. Split Fusion Proteins
[0455] In some embodiments, the fusion protein is a split protein, i.e. comprises two or more separate polypeptide domains that interact or self-assemble to form a functional fusion protein. In some aspects, the split fusion protein comprises a dCas9 and an effector domain. In some aspects, the fusion protein comprises a split dCas9-effector domain fusion protein.
[0456] In some embodiments, the split fusion protein is assembled from separate polypeptide domains comprising trans-splicing inteins. Inteins are internal protein elements that self-excise from their host protein and catalyze ligation of flanking sequences with a peptide bond. In some embodiments, the split fusion protein is assembled from a first polypeptide comprising an N-terminal intein and a second polypeptide comprising a C-terminal intein. In some embodiments, the N terminal intein is the N terminal Npu Intein set forth in SEQ ID NO: 178. In some embodiments, the C terminal intein is the C terminal Npu intein set forth in SEQ ID NO: 182.
[0457] Also provided are fusion proteins comprising a first polypeptide of a split variant Cas protein comprising an N-terminal fragment of a Cas protein and an N-terminal Intein, and at least one effector domain, wherein the effector domain induces transcription activation, transcription co-activation,
transcription elongation, transcription de-repression, transcription factor release, polymerization, histone modification, histone acetylation, histone deacetylation, nucleosome remodeling, chromatin remodeling, reversal of heterochromatin formation, nuclease, signal transduction, proteolysis, ubiquitination, deubiquitination, phosphorylation, dephosphorylation, splicing, nucleic acid association, DNA methylation, DNA demethylation, histone methylation, histone demethylation, or DNA base oxidation. Also provided are fusion proteins comprising a first polypeptide of a split variant Cas protein comprising an N-terminal fragment of a Cas protein (an exemplary amino acid sequence set forth in SEQ ID NO: 176, and an exemplary nucleotide sequence encoding the N-terminal fragment of the variant Cas protein set forth in SEQ ID NO: 175) and an N-terminal Intein (an exemplary amino acid sequence set forth in SEQ ID NO: 178 and an exemplary nucleotide sequence encoding the N-terminal Intein set forth in SEQ ID NO: 177), and at least one effector domain, wherein the effector domain increases transcription of the FXN locus. In some aspects, the first polypeptide of the split variant Cas protein, and a second polypeptide (dSpCas9-573-C; amino acid sequence set forth in SEQ ID NO: 180, and an exemplary nucleotide sequence encoding dSpCas9-573-C set forth in SEQ ID NO: 179) of the split variant Cas protein comprising a C-terminal fragment of the variant Cas protein (an exemplary amino acid sequence set forth in SEQ ID NO: 184, and an exemplary nucleotide sequence encoding the C- terminal fragment of the variant Cas protein set forth in SEQ ID NO: 183) and a C-terminal Intein (an exemplary amino acid sequence set forth in SEQ ID NO: 182 and an exemplary nucleotide sequence encoding the C-terminal Intein set forth in SEQ ID NO: 181), are present in proximity or present in the same cell, the N-terminal Intein and C-terminal Intein self-excise and ligate the N-terminal fragment and the C-terminal fragment of the variant Cas9 to form a full-length variant Cas9 protein.
[0458] Also provided are fusion proteins comprising a second polypeptide of a split variant Cas protein comprising a C-terminal fragment of a Cas protein and a C-terminal Intein and at least one effector domain, wherein the effector domain induces transcription activation, transcription co-activation, transcription elongation, transcription de-repression, transcription factor release, polymerization, histone modification, histone acetylation, histone deacetylation, nucleosome remodeling, chromatin remodeling, reversal of heterochromatin formation, nuclease, signal transduction, proteolysis, ubiquitination, deubiquitination, phosphorylation, dephosphorylation, splicing, nucleic acid association, DNA methylation, DNA demethylation, histone methylation, histone demethylation, or DNA base oxidation. Also provided are fusion proteins comprising a second polypeptide of a split variant Cas protein comprising a C-terminal fragment of a Cas protein and a C-terminal Intein and at least one effector domain, wherein the effector domain increases transcription of the FXN locus. In some aspects, the second polypeptide of the split variant Cas protein, and a first polypeptide of the split variant Cas protein comprising an N-terminal fragment of the variant Cas protein and an N-terminal Intein, are present in proximity or present in the same cell, the N-terminal Intein and C-terminal Intein self-excise and ligate the N-terminal fragment and the C-terminal fragment of the variant Cas9 to form a full-length variant Cas9 protein.
[0459] In some embodiments, the split fusion protein comprises a split dCas9-effector domain fusion protein assembled from two polypeptides. In an exemplary embodiment, the first polypeptide comprises an effector domain catalytic domain and an N-terminal fragment of dSpCas9, followed by an N terminal Npu Intein (effector domain-dSpCas9-573N), and the second polypeptide comprises a C terminal Npu Intein, followed by a C-terminal fragment of dSpCas9 (dSpCas9-573C. The N- and C- terminal fragments of the fusion protein are split at position 573Glu of the dSpCas9 molecule, with reference to SEQ ID NO:79. In some aspects, the N-terminal Npu Intein (SEQ ID NO: 178) and C- terminal Npu Intein (set forth in SEQ ID NO: 182) may self-excise and ligate the two fragments, thereby forming the full-length dSpCas9-effector domain fusion protein when expressed in a cell.
[0460] In some embodiments, the polypeptides of a split protein may interact non-covalently to form a complex that recapitulates the activity of the non-split protein. For example, two domains of a Cas enzyme expressed as separate polypeptides may be recruited by a gRNA to form a ternary complex that recapitulates the activity of the full-length Cas enzyme in complex with the gRNA, for example as described in Wright et al. PNAS 112(10):2984-2989 (2015). In some embodiments, assembly of the split protein is inducible (e.g. light inducible, chemically inducible, small-molecule inducible).
[0461] In some aspects, the two polypeptides of a split fusion protein may be delivered and/or expressed from separate vectors, such as any of the vectors described herein. In some embodiments, the two polypeptides of a split fusion protein may be delivered to a cell and/or expressed from two separate AAV vectors, i.e. using a split AAV-based approach, for example as described in WO 2017/197238.
[0462] Approaches for the rationale design of split proteins and their delivery, including Cas proteins and fusions thereof, are described, for example, in WO 2016/114972, WO 2017/197238, Zetsche, et al. Nat. Biotechnol. 33(2): 139-42 (2015), Wright et al. PNAS 112(10):2984-2989 (2015), Truong, et al. Nucleic Acids Res. 43, 6450-6458 (2015), and Fine et al. Sci. Rep. 5, 10777 (2015).
D. Combinations of DNA-targeting systems or fusion proteins
[0463] Also provided are combinations, such as combinations of two or more DNA-targeting systems or components thereof. In some aspects, provided herein are combinations of two or more DNA-targeting systems that independently target different target sites at a frataxin (FXN) locus. In some aspects, the two or more DNA-targeting systems each comprise any of the DNA-targeting systems described herein.
[0464] In some embodiments, the DNA-targeting domain is a first DNA-targeting domain, and the DNA-targeting system further comprises one or more second DNA-targeting domains. In some embodiments, the first DNA-targeting domain binds a first target site in an enhancer of a FXN locus, and the second DNA-targeting domain binds a second target site in a promoter of a FXN locus.
[0465] In some aspects, the provided combination of DNA-targeting systems include two or more DNA-targeting systems, each of which target particular regions of a frataxin (FXN) locus.
[0466] Also provided herein is a combination, comprising a first DNA-targeting system comprising
any of the DNA-targeting systems described herein, and one or more second DNA-targeting systems that binds to a second target site in a regulatory DNA element of a frataxin (FXN) locus. In some embodiments, the second DNA-targeting system comprises any of the DNA-targeting systems described herein.
[0467] Also provided are combinations, such as combinations of two or more DNA-targeting domains or fusion proteins or components thereof. In some aspects, provided herein are combinations of two or more DNA-targeting domains or fusion proteins that independently target different target sites at a frataxin (FXN) locus. In some aspects, the two or more DNA-targeting domains or fusion proteins each comprise any of the DNA-targeting domains or fusion proteins described herein.
[0468] In some embodiments, the DNA-targeting domain is a first DNA-targeting domain, and the DNA-targeting domain or fusion protein further comprises one or more second DNA-targeting domains. In some embodiments, the first DNA-targeting domain binds a first target site in an enhancer of a FXN locus, and the second DNA-targeting domain binds a second target site in a promoter of a FXN locus.
[0469] In some aspects, the provided combination of DNA-targeting domains or fusion proteins include two or more DNA-targeting domains or fusion proteins, each of which target particular regions of a frataxin (FXN) locus.
[0470] Also provided herein is a combination, comprising a first DNA-targeting domain or fusion protein comprising any of the DNA-targeting domains or fusion proteins described herein, and one or more second DNA-targeting domains or fusion proteins that binds to a second target site in a regulatory DNA element of a frataxin (FXN) locus. In some embodiments, the second DNA-targeting domain or fusion protein comprises any of the DNA-targeting domains or fusion proteins described herein.
[0471] Also provided herein are DNA-targeting systems that binds to one or more target sites in a regulatory DNA element of a frataxin (FXN) locus, the DNA-targeting system comprising: a first DNA- targeting domain that binds a first target site in an enhancer of a FXN locus, and a second DNA-targeting domain that binds a second target site in a promoter of a FXN locus.
[0472] In some aspects, exemplary combination of DNA-targeting systems include: (a) a fusion protein comprising a Cas protein or a variant thereof and (b) a combination of gRNAs, such as a first gRNA that is capable of hybridizing to the target site or is complementary to the first target site and a second gRNA that is capable of hybridizing to the target site or is complementary to the second target site. In some aspects, also provided herein are combinations of DNA-targeting systems comprising one type of Cas protein or variant thereof, such as a dCas9 protein or variant thereof, and two or more different gRNAs, such as a combination of gRNAs, such as any combination of gRNAs described herein. In some aspects, also provided herein are combinations of DNA-targeting systems comprising one type of Cas protein or variant thereof, such as a dCas9 protein or variant thereof, two or more different types of effector domains, and two or more different gRNAs, such as a combination of gRNAs, such as any combination of gRNAs described herein. In some aspects, also provided herein are combinations of
DNA-targeting systems comprising two or more different type of Cas protein or variant thereof, such as a
dCas9 protein or variant thereof, and two or more different gRNAs, such as a combination of gRNAs, such as any combination of gRNAs described herein. In some aspects, also provided herein are combinations of DNA-targeting systems comprising two or more different types of DNA-targeting domains and one type of effector domain. In some aspects, also provided herein are combinations of DNA-targeting systems comprising two or more different types of DNA-targeting domains and two or more different types of effector domain.
[0473] In some embodiments, the first target site is located within the genomic coordinates human genome assembly GRCh38 (hg38) chr9:69, 027, 282-69, 028, 497, and the second target site is located within the genomic coordinates hg38 chr9:68, 940, 179-69, 205, 519. In some embodiments, the first target site is located within the genomic coordinates hg38 chr9:69, 027, 615-69, 028, 101, and the second target site is located within the genomic coordinates hg38 chr9:68, 940, 179-69, 205, 519.
[0474] In some embodiments, the first DNA-targeting domain comprises a first Cas-gRNA combination comprising (a) a first Cas protein or a variant thereof and (b) a first gRNA that is capable of hybridizing to the target site or is complementary to the first target site; and the second DNA-targeting domain comprises a second Cas-gRNA combination comprising (a) a second Cas protein or a variant thereof and (b) a second gRNA that is capable of hybridizing to the target site or is complementary to the second target site. In some embodiments, the first DNA-targeting domain comprises a first Cas-gRNA combination comprising (a) a first Cas protein or a variant thereof and (b) a first gRNA comprising at least one gRNA spacer sequence set forth in SEQ ID NO:42 or a contiguous portion thereof of at least 14 nt. In some embodiments, the second DNA-targeting domain comprises a second Cas-gRNA combination comprising (a) a second Cas protein or a variant thereof and (b) a second gRNA comprising at least one gRNA spacer sequence set forth in SEQ ID NO:22 or 28 or a contiguous portion thereof of at least 14 nt.
[0475] In some embodiments, the first Cas-gRNA combination comprises (a) a first Cas protein or a variant thereof and (b) a first gRNA comprising at least one gRNA spacer sequence set forth in SEQ ID NO:42 or a contiguous portion thereof of at least 14 nt; and the second Cas-gRNA combination comprises (a) a second Cas protein or a variant thereof and (b) a second gRNA comprising at least one gRNA spacer sequence set forth in SEQ ID NO:22 or a contiguous portion thereof of at least 14 nt. In some embodiments, the first Cas-gRNA combination comprises (a) a first Cas protein or a variant thereof and (b) a first gRNA comprising at least one gRNA spacer sequence set forth in SEQ ID NO:42 or a contiguous portion thereof of at least 14 nt; and the second Cas-gRNA combination comprises (a) a second Cas protein or a variant thereof and (b) a second gRNA comprising at least one gRNA spacer sequence set forth in SEQ ID NO:28 or a contiguous portion thereof of at least 14 nt.
[0476] In some embodiments, the first DNA-targeting domain binds a first target site in a promoter of a FXN locus; and the second DNA-targeting domain binds a second target site in a promoter of a FXN locus.
[0477] Also provided herein are DNA-targeting systems that binds to one or more target sites in a
regulatory DNA element of a frataxin (FXN) locus, the DNA-targeting system comprising: a first DNA- targeting domain that binds a first target site in a promoter of a FXN locus; and a second DNA-targeting domain that binds a second target site in a promoter of a FXN locus.
[0478] In some embodiments, the first target site and the second target site independently are located within the genomic coordinates hg38 chr9:68, 940, 179-69, 205 ,519. In some embodiments, the first target site and the second target site are different.
[0479] In some embodiments, the first DNA-targeting domain comprises a first Cas-gRNA combination comprising (a) a first Cas protein or a variant thereof and (b) a first gRNA that is capable of hybridizing to the target site or is complementary to the first target site; and the second DNA-targeting domain comprises a second Cas-gRNA combination comprising (a) a second Cas protein or a variant thereof and (b) a second gRNA that is capable of hybridizing to the target site or is complementary to the second target site. In some embodiments, the first DNA-targeting domain comprises a first Cas-gRNA combination comprising (a) a first Cas protein or a variant thereof and (b) a first gRNA comprising at least one gRNA spacer sequence set forth in SEQ ID NO:22 or a contiguous portion thereof of at least 14 nt. In some embodiments, the second DNA-targeting domain comprises a second Cas-gRNA combination comprising (a) a second Cas protein or a variant thereof and (b) a second gRNA comprising at least one gRNA spacer sequence set forth in SEQ ID NO:28 or a contiguous portion thereof of at least 14 nt.
[0480] In some embodiments, the first Cas-gRNA combination comprises (a) a first Cas protein or a variant thereof and (b) a first gRNA comprising at least one gRNA spacer sequence set forth in SEQ ID NO:22 or a contiguous portion thereof of at least 14 nt; and the second Cas-gRNA combination comprises (a) a second Cas protein or a variant thereof and (b) a second gRNA comprising at least one gRNA spacer sequence set forth in SEQ ID NO:28 or a contiguous portion thereof of at least 14 nt.
[0481] In some embodiments, the first Cas protein or a variant thereof and/or the second Cas protein or a variant thereof is a variant Cas9 protein that lacks nuclease activity or that is a deactivated Cas9 (dCas9) protein.
[0482] In some embodiments, the first variant Cas protein and/or the second variant Cas protein is a Staphylococcus aureus dCas9 protein (dSaCas9) that comprises at least one amino acid mutation selected from D10A and N580A, with reference to numbering of positions of SEQ ID NO:73; or comprises the sequence set forth in SEQ ID NO:72, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
[0483] In some embodiments, the first variant Cas protein and/or the second variant Cas protein is a Streptococcus pyogenes dCas9 (dSpCas9) protein that comprises at least one amino acid mutation selected from D10A and H840A, with reference to numbering of positions of SEQ ID NO:79; or comprises the sequence set forth in SEQ ID NO:78, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
[0484] In some embodiments, the first Cas protein and the second Cas protein are the same. In some
embodiments, the first Cas protein and the second Cas protein are identical. In some embodiments, the first Cas protein and the second Cas protein are different. In some embodiments, the first Cas protein and the second Cas protein are from different species.
[0485] In some embodiments, the first Cas protein or a variant thereof and/or the second Cas protein or a variant thereof is fused to at least one effector domain.
[0486] In some embodiments, the effector domain induces, catalyzes or leads to transcription activation, transcription co-activation, transcription elongation, transcription de -repression, transcription factor release, polymerization, histone modification, histone acetylation, histone deacetylation, nucleosome remodeling, chromatin remodeling, reversal of heterochromatin formation, nuclease, signal transduction, proteolysis, ubiquitination, deubiquitination, phosphorylation, dephosphorylation, splicing, nucleic acid association, DNA methylation, DNA demethylation, histone methylation, histone demethylation, or DNA base oxidation. In some embodiments, the effector domain induces transcription activation.
[0487] In some embodiments, all of the components of the combination of DNA-targeting systems, DNA-targeting domains or fusion proteins provided herein are encoded in one polynucleotide. In some embodiments, all of the components of the combination of DNA-targeting systems, DNA-targeting domains or fusion proteins provided herein are encoded in multiple individual polynucleotides, such as a first polynucleotide and a second polynucleotide. In some aspects, first DNA-targeting system, DNA- targeting domain or fusion protein and the second DNA-targeting system, DNA-targeting domain or fusion protein are encoded in one polynucleotide, such as a first polynucleotide. In some embodiments, the first DNA-targeting system, domain or fusion protein and the second DNA-targeting system, domain or fusion protein are encoded in one polynucleotide, such as a first polynucleotide. In some embodiments, the first Cas protein and the second Cas protein are encoded in a first polynucleotide. In some embodiments, the first Cas protein and the second Cas protein are encoded by the same nucleotide sequence. In some embodiments, the first gRNA and the second gRNA are encoded in a first polynucleotide. In some embodiments, the first Cas protein and the second Cas protein are encoded by the same nucleotide sequence, and the Cas protein, the first gRNA, and the second gRNA are encoded in a first polynucleotide. In some embodiments, the first DNA-targeting domain is encoded in a first polynucleotide and the second DNA-targeting domain is encoded in a second polynucleotide. In some embodiments, the first Cas protein is encoded in a first polynucleotide and the second Cas protein is encoded in a second polynucleotide. In some embodiments, the first gRNA is encoded in a first polynucleotide and the second gRNA is encoded in a second polynucleotide. In some embodiments, the first Cas protein and the first gRNA are encoded in a first polynucleotide, and the second Cas protein and the second gRNA are encoded in a second polynucleotide.
III. POLYNUCLEOTIDES, VECTORS AND DELIVERY OF DNA-TARGETING SYSTEMS
[0488] Provided are polynucleotides encoding any of the eZFPs provided herein, any of the eZFP
fusion proteins described herein, any of the DNA-targeting systems described herein, any of the gRNAs described herein, any of the combinations described herein, and/or any of the fusion proteins described herein, or a portion or a component of any of the foregoing. In some aspects, the polynucleotides can encode any of the components of the DNA-targeting systems, and/or any nucleic acid or proteinaceous molecule necessary to carry out aspects of the methods of the disclosure can comprise a vector (e.g., a recombinant expression vector). In some of any embodiments, provided are polynucleotides encoding any of the fusion proteins described herein (such as eZFP fusion proteins or dCas fusion proteins). Also provided herein are polynucleotides encoding any of the gRNAs or combinations of gRNAs described herein.
A. Nucleic Acids
[0489] Provided are polynucleotides encoding any of the eZFPs provided herein, any of the eZFP fusion proteins described herein, any of the DNA-targeting systems comprising eZFPs or eZFP fusion proteins described herein, or a portion or component thereof, or a combination of any of the foregoing.
[0490] Also provided are any of the DNA-targeting systems described herein, including a protein component of the DNA-targeting system (e.g., Cas protein or a variant thereof) and the at least one gRNA, such as one or more RNAs.
[0491] In some embodiments, provided are polynucleotides comprising the gRNAs described herein. In some embodiments, the gRNA is transcribed from a genetic construct (i.e. vector or plasmid) in the target cell. In some embodiments, the gRNA is produced by in vitro transcription and delivered to the target cell. In some embodiments, the gRNA comprises one or more modified nucleotides for increased stability. In some embodiments, the gRNA is delivered to the target cell pre-complexed as a RNP with the fusion protein.
[0492] In some embodiments, a provided polynucleotide encodes a fusion protein as described herein that includes (a) a DNA-targeting domain capable of being targeted to a target site of a target gene as described; and (b) at least one effector domain capable of reducing transcription of the gene. In some embodiments, the fusion protein includes a fusion protein of a Cas protein or variant thereof and at least one effector domain capable of increasing transcription of a gene. In a particular example, the Cas is a deactivated Cas (dCas), such as dCas9. In some embodiments, the dCas9 is a dSpCas9. Examples of such domains and fusion proteins include any as described in Section II.C.
[0493] In some embodiments, a provided polynucleotide encodes an eZFP fusion protein as described herein.
[0494] In some embodiments, the polynucleotide is DNA. In some embodiments, the polynucleotide is RNA. In some embodiments, the polynucleotide is mRNA.
[0495] In some embodiments, the polynucleotide is a gRNA. In some embodiments, the gRNA is provided as RNA and a polynucleotide encoding the dCas fusion protein is mRNA. In some aspects, the mRNA is 5' capped and/or 3' polyadenylated. In some embodiments, a polynucleotide provided herein is
DNA. In some aspects, the DNA is present in a vector.
[0496] In some embodiments, the polynucleotide encodes the fusion protein and one or more gRNAs or a combination of gRNAs.
[0497] In some embodiments, the polynucleotide as provided herein can be codon optimized for efficient translation into protein in the eukaryotic cell or animal of interest. For example, codons can be optimized for expression in humans, mice, rats, hamsters, cows, pigs, cats, dogs, fish, amphibians, plants, yeast, insects, and others.
[0498] In some embodiments, the polynucleotide comprises the sequence set forth in SEQ ID NO:68 or 74, or a sequence having at least about 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity thereto. In some embodiments, the polynucleotide comprises the sequence set forth in SEQ ID NO:68. In some embodiments, the polynucleotide comprises the sequence set forth in SEQ ID NO:74.
[0499] In some embodiments, the polynucleotide comprises the sequence set forth in any of SEQ ID NOS:308-314 and 457-474, or a sequence having at least about 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity thereto. In some embodiments, the polynucleotide comprises the sequence set forth in any of SEQ ID NOS:308-314 and 457-474. In some embodiments, the polynucleotide comprises the sequence set forth in any of SEQ ID NOS:426-429, or a sequence having at least about 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity thereto. In some embodiments, the polynucleotide comprises the sequence set forth in any of SEQ ID NOS:426-429.
[0500] Also provided are polynucleotides encoding a first DNA-targeting system, a first Cas protein and/or a first gRNA of any of the DNA-targeting systems described herein or any of the combinations described herein.
[0501] Provided are polynucleotides encoding a second DNA-targeting system, a second Cas protein and/or a second gRNA of any of the DNA-targeting systems described herein or any of the combinations described herein.
[0502] Provided are polynucleotides that include any of the polynucleotides described herein, and one or more additional polynucleotides encoding an additional portion or an additional component of any of the DNA-targeting systems described herein, any of the gRNAs described herein, any of the combinations described herein, or any of the fusion proteins described herein, or a portion or a component of any of the foregoing.
[0503] Also provided herein are pluralities of polynucleotides, comprising: (a) a polynucleotide encoding a first DNA-targeting system, a first Cas protein and/or a first gRNA of any of the a DNA- targeting system disclosed herein or any of the combinations of gRNAs disclosed herein, and (b) a polynucleotide encoding a second DNA-targeting system, a second Cas protein and/or a second gRNA of any of the DNA-targeting system disclosed herein or any of the combinations of gRNAs disclosed herein.
[0504] Provided are pluralities of polynucleotides, that includes a first polynucleotide comprising
any of the polynucleotides described herein; and a second polynucleotide comprising any of the polynucleotides described herein.
[0505] In some embodiments, the first DNA-targeting domain and the second DNA-targeting domain are encoded in a first polynucleotide. In some embodiments, the first Cas protein and the second Cas protein are encoded in a first polynucleotide. In some embodiments, the first Cas protein and the second Cas protein are encoded by the same nucleotide sequence. In some embodiments, the first gRNA and the second gRNA are encoded in a first polynucleotide. In some embodiments, the first Cas protein and the second Cas protein are encoded by the same nucleotide sequence, and the Cas protein, the first gRNA, and the second gRNA are encoded in a first polynucleotide.
[0506] In some embodiments, the first DNA-targeting domain is encoded in a first polynucleotide and the second DNA-targeting domain is encoded in a second polynucleotide. In some embodiments, the first Cas protein is encoded in a first polynucleotide and the second Cas protein is encoded in a second polynucleotide. In some embodiments, the first gRNA is encoded in a first polynucleotide and the second gRNA is encoded in a second polynucleotide. In some embodiments, the first Cas protein and the first gRNA are encoded in a first polynucleotide, and the second Cas protein and the second gRNA are encoded in a second polynucleotide.
B. Vectors
[0507] Provided are vectors that include any of the polynucleotides described herein, any of the pluralities of polynucleotides described herein, or a first polynucleotide or a second polynucleotide of any of the pluralities of polynucleotides described herein, or a portion or a component of any of the foregoing. In some embodiments, the vector comprises any of the eZFPs, fusion proteins (such as eZFP fusion proteins or dCas fusion proteins), gRNAs, or DNA-targeting systems provided herein, components thereof, or a combination of any of the foregoing. In some embodiments, the vector comprises a polynucleotide encoding any of the eZFPs, fusion proteins (such as eZFP fusion proteins or dCas fusion proteins), gRNAs, or DNA-targeting systems provided herein, components thereof, or a combination of any of the foregoing. In some embodiments, the vector is any suitable AAV vector. In some embodiments, the AAV vector is AAV-DJ, AAV9, AAV6, or AAVrh74.
[0508] Also provided herein is a vector that comprises or contains any of the provided polynucleotides. In some embodiments, the vector comprises a genetic construct, such as a plasmid or an expression vector. The vector can be a self-inactivating vector that either inactivates the viral sequences or the components of the CRISPR machinery or other elements.
[0509] In some embodiments, the expression vector comprising the sequence encoding the fusion protein of a CRISPR/Cas-based DNA-targeting system provided herein further comprises a nucleic acid sequence encoding at least one gRNA. In some embodiments, the expression vector comprises a nucleic acid sequence or combination of nucleic acid sequences encoding two or more gRNAs, such as two gRNAs. In some embodiments, the expression vector comprises a nucleic acid sequence or combination
of nucleic acid sequences encoding three gRNAs. In some cases, the sequence encoding the gRNA is operably linked to at least one transcriptional control sequence or transcriptional regulatory sequence (e.g., cis-regulatory sequence) for expression of the gRNA in the cell. In some aspects, DNA encoding the gRNA can be operably linked to a promoter sequence (i.e. a promoter for the gRNA) that is recognized by RNA polymerase III (Pol III). Examples of suitable Pol III promoters include, but are not limited to, mammalian U6, U3, Hl, and 7SL RNA promoters, or variants thereof. In some aspects, if the expression vector comprises nucleic acid sequences encoding two or more gRNAs, each gRNA is operably linked to an identical Pol III promoter, or different Pol III promoters.
[0510] In some embodiments, provided is a vector containing a polynucleotide that encodes a fusion protein comprising a DNA-targeting domain comprising a dCas and at least one effector domain capable of increasing transcription of a gene, and a polynucleotide or combination of polynucleotides encoding a gRNA, or a plurality of gRNAs, such as two, three, or four or more gRNAs, or such as two, three, or four or more different gRNAs. In some embodiments, the dCas is a dCas9, such as dSaCas9 or dSpCas9. In some embodiments, the polynucleotide encodes a fusion protein that includes a dSaCas9 set forth in SEQ ID NO:72. In some embodiments, the polynucleotide encodes a fusion protein that includes a dSpCas9 set forth in SEQ ID NO:78. In some embodiments, the polynucleotide(s) encodes one or more a gRNAs described herein, for example in or a plurality of gRNAs, each gRNA as described in Section II.C.l.
[0511] In some examples, a polynucleotide and/or a vector described herein can comprise one or more transcription and/or translation control elements. Depending on the host/vector system utilized, any of a number of suitable transcription and translation control elements, including constitutive and inducible promoters, transcription enhancer elements, transcription terminators, etc. can be used in the expression vector.
[0512] Non-limiting examples of suitable eukaryotic promoters (i.e., promoters functional in a eukaryotic cell) include those from cytomegalovirus (CMV) immediate early, herpes simplex virus (HSV) thymidine kinase, early and late SV40, long terminal repeats (LTRs) from retrovirus, human elongation factor- 1 promoter (EFl), human elongation factor- 1 promoter alpha (EFla), a hybrid construct comprising the cytomegalovirus (CMV) enhancer fused to the chicken beta-actin promoter (CAG), murine stem cell virus promoter (MSCV), phosphoglycerate kinase- 1 locus promoter (PGK), mouse metallothionein-I, and human elongation factor alpha short promoter (EFS).
[0513] For expressing small RNAs, including guide RNAs used in connection with the DNA- targeting systems, various promoters such as RNA polymerase III promoters, including for example U6 and Hl, can be advantageous. Additional information and approaches include those described in, e.g., Ma, H. et al., Molecular Therapy — Nucleic Acids 3, el61 (2014) doi:10.1038/mtna.2014.12. In some embodiments, the promoter for the gRNA is a U6 promoter (e.g., as set forth in SEQ ID NO:432). In some embodiments, the promoter for the gRNA is a mini-U6 promoter (e.g., as set forth in SEQ ID NO:433). In some aspects, a mini-U6 promoter is an engineered promoter based on a U6 promoter, which is smaller in size than a U6 promoter. In some aspects, the smaller size of the mini-U6 promoter is
advantageous for expression vectors (such as AAV vectors) that have size constraints for polynucleotides to be delivered.
[0514] The expression vector can also contain a ribosome binding site for translation initiation and a transcription terminator. The expression vector can also comprise appropriate sequences for amplifying expression. The expression vector can also include nucleotide sequences encoding non-native tags (e.g., histidine tag, hemagglutinin tag, green fluorescent protein, etc.) that are fused to the site-directed polypeptide, thus resulting in a fusion protein.
[0515] A promoter can be an inducible promoter (e.g., a heat shock promoter, tetracycline-regulated promoter, steroid-regulated promoter, metal-regulated promoter, estrogen receptor-regulated promoter, etc.). The promoter can be a constitutive promoter (e.g., CMV promoter, UBC promoter). In some cases, the promoter can be a spatially restricted and/or temporally restricted promoter (e.g., a tissue specific promoter, a cell type specific promoter (e.g. nervous system specific promoter), etc.). In some embodiments, the promoter is a human elongation factor alpha short promoter EFS promoter. In some embodiments, the promoter is a CAG promoter. In some embodiments, the promoter is a EFla promoter.
[0516] In some examples, vectors can be capable of directing the expression of nucleic acids to which they are operatively linked. Such vectors are referred to herein as “recombinant expression vectors”, or more simply “expression vectors”, which serve equivalent functions.
[0517] Exemplary expression vectors contemplated include, but are not limited to, viral vectors based on vaccinia virus, poliovirus, adenovirus, adeno-associated virus, SV40, herpes simplex virus, human immunodeficiency virus, retrovirus (e.g., Murine Leukemia Virus, spleen necrosis virus, and vectors derived from retroviruses such as Rous Sarcoma Virus, Harvey Sarcoma Virus, avian leukosis virus, a lentivirus, human immunodeficiency virus, myeloproliferative sarcoma virus, and mammary tumor virus) and other recombinant vectors. Other vectors contemplated for eukaryotic target cells include, but are not limited to, the vectors pXTl, pSG5, pSVK3, pBPV, pMSG, and pSVLSV40 (Pharmacia). Other vectors can be used so long as they are compatible with the host cell.
[0518] In some embodiments, the vector is a viral vector, such as an adeno-associated virus (AAV) vector, a retroviral vector, a lentiviral vector, or a gammaretroviral vector. N some embodiments, the viral vector is an adeno-associated virus (AAV) vector. In some embodiments, the AAV vector is selected from among an AAV1, AAV2, AAV3, AAV4, AAV5, AAV6, AAV7, AAV8, or AAV9 vector. In some embodiments, the AAV vector is an AAV6, AAV9, AAV-DJ, or AAVrh74 vector. In some embodiments, the vector is a lentiviral vector. In some embodiments, the vector is a non- viral vector, for example a lipid nanoparticle, a liposome, an exosome, or a cell penetrating peptide.
[0519] In some embodiments, the vector comprises one vector, or two or more vectors.
[0520] In some aspects, provided herein are pluralities of vectors that comprise any of the vectors described herein, and one or more additional vectors comprising one or more additional polynucleotides encoding an additional portion or an additional component of any of the DNA-targeting systems described herein, any of the gRNAs described herein, any of the combinations described herein, any of
the eZFPs described herein, or any of the fusion proteins described herein (such as dCas or eZFP fusion proteins), or a portion or a component of any of the foregoing.
[0521] Provided are pluralities of vectors, that include: a first vector comprising any of the polynucleotides described herein; and a second vector comprising any of the polynucleotides described herein. Also provided herein are pluralities of vectors, comprising: a first vector comprising a polynucleotide encoding a first DNA-targeting system, a first Cas protein and/or a first gRNA of any of the DNA-targeting system described herein or any of the combinations of gRNAs described herein; and; a second vector comprising a polynucleotide encoding a second DNA-targeting system, a second Cas protein and/or a second gRNA of any of the DNA-targeting system described herein or any of the combinations of gRNAs described herein.
[0522| Also provided herein are pluralities of vectors, comprising: a first vector comprising a polynucleotide encoding a first eZFP or eZFP fusion protein provided herein, and a second vector comprising a polynucleotide encoding a second eZFP or eZFP fusion protein provided herein.
[0523] In some embodiments, polynucleotides can be cloned into a suitable vector, such as an expression vector or vectors. The expression vector can be any suitable recombinant expression vector, and can be used to transform or transfect any suitable cell. Suitable vectors include those designed for propagation and expansion or for expression or both, such as plasmids and viruses.
[0524] In some embodiments, the vector can be a vector of the pUC series (Fermentas Life Sciences), the pBluescript series (Stratagene, LaJolla, Calif.), the pET series (Novagen, Madison, Wis.), the pGEX series (Pharmacia Biotech, Uppsala, Sweden), or the pEX series (Clontech, Palo Alto, Calif.). In some embodiments, animal expression vectors include pEUK-Cl, pMAM and pMAMneo (Clontech). In some embodiments, a viral vector is used, such as a lenti viral or retroviral vector. In some embodiments, the recombinant expression vectors can be prepared using standard recombinant DNA techniques. In some embodiments, vectors can contain regulatory sequences, such as transcription and translation initiation and termination codons, which are specific to the type of host into which the vector is to be introduced, as appropriate and taking into consideration whether the vector is DNA- or RNA- based. In some embodiments, the vector can contain a nonnative promoter operably linked to the nucleotide sequence encoding the recombinant receptor. In some embodiments, the promoter can be a non- viral promoter or a viral promoter, such as a cytomegalovirus (CMV) promoter, an SV40 promoter, an RSV promoter, and a promoter found in the long-terminal repeat of the murine stem cell virus. Other promoters also are contemplated.
[0525] In some embodiments, recombinant nucleic acids are transferred into cells using recombinant infectious virus particles, such as, e.g., vectors derived from simian virus 40 (SV40), adenoviruses, or adeno-associated virus (AAV). In some embodiments, recombinant nucleic acids are transferred into cells (e.g. central nervous system cells, such as neurons) using recombinant lentiviral vectors or retroviral vectors, such as gamma-retroviral vectors (see, e.g., Koste et al. (2014) Gene Therapy 2014 Apr 3. Doi: 10.1038/gt.2014.25; Carlens et al. (2000) Exp Hematol 28(10): 1137-46; Alonso-Camino et al. (2013)
Mol Ther Nucl Acids 2, e93; Park et al., Trends Biotechnol. 2011 November 29(11): 550-557.
[0526] In some embodiments, the retroviral vector has a long terminal repeat sequence (LTR), e.g., a retroviral vector derived from the Moloney murine leukemia virus (MoMLV), myeloproliferative sarcoma virus (MPSV), murine embryonic stem cell virus (MESV), murine stem cell virus (MSCV), spleen focus forming virus (SFFV), or adeno-associated virus (AAV). Most retroviral vectors are derived from murine retroviruses. In some embodiments, the retroviruses include those derived from any avian or mammalian cell source. The retroviruses typically are amphotropic, meaning that they are capable of infecting host cells of several species, including humans. In one embodiment, the gene to be expressed replaces the retroviral gag, pol and/or env sequences. A number of illustrative retroviral systems have been described (e.g., U.S. Pat. Nos. 5,219,740; 6,207,453; 5,219,740; Miller and Rosman (1989) BioTechniques 7:980-990; Miller, A. D. (1990) Human Gene Therapy 1:5-14; Scarpa et al. (1991) Virology 180:849-852; Burns et al. (1993) Proc. Natl. Acad. Sci. USA 90:8033-8037; and Boris-Fawrie and Temin (1993) Cur. Opin. Genet. Develop. 3: 102-109.
[0527] In some embodiments, the vector is a lentiviral vector. In some embodiments, the lenti viral vector is an integrase-deficient lentiviral vector. In some embodiments, the lentiviral vector is a recombinant lentiviral vector. In some embodiments, the lenti virus is selected or engineered for a desired tropism (e.g. for central nervous system tropism, or tropism for a heart cell, such as a cardiomyocyte, a skeletal muscle cell, a nervous system cell, such as a neuron, a fibroblast, or an induced pluripotent stem cell). In some embodiments, the cell for any of the provided compositions, such as DNA-targeting systems, fusion proteins, gRNAs, polynucleotides and/or vectors to be delivered is a heart cell, a skeletal muscle cell, a nervous system cell, or an induced pluripotent stem cell. Methods of lentiviral production, transduction, and engineering are known, for example as described in Kasaraneni, N. et al. Sci. Rep. 8(l):10990 (2018), Ghaleh, H.E.G. et al. Biomed. Pharmacother. 128:110276 (2020), and Milone, M.C. et al. Eeukemia. 32(7): 1529-1541 (2018). Additional methods for lentiviral transduction are described, for example in Wang et al. (2012) J. Immunother. 35(9): 689-701; Cooper et al. (2003) Blood. 101: 1637- 1644; Verhoeyen et al. (2009) Methods Mol Biol. 506: 97-114; and Cavalieri et al. (2003) Blood. 102(2): 497-505.
[0528] In some embodiments, recombinant nucleic acids are transferred into cells (e.g. central nervous system cells, such as neurons, or a heart cell, a skeletal muscle cell, a nervous system cell, or an induced pluripotent stem cell) via electroporation (see, e.g., Chicaybam et al, (2013) PloS ONE 8(3): e60298 and Van Tedeloo et al. (2000) Gene Therapy 7(16): 1431-1437). In some embodiments, recombinant nucleic acids are transferred into cells via transposition (see, e.g., Manuri et al. (2010) Hum Gene Ther 21(4): 427-437; Sharma et al. (2013) Molec Ther Nucl Acids 2, e74; and Huang et al. (2009) Methods Mol Biol 506: 115-126). Other methods of introducing and expressing genetic material into immune cells include calcium phosphate transfection (e.g., as described in Current Protocols in Molecular Biology, John Wiley & Sons, New York. N.Y.), protoplast fusion, cationic liposome-mediated transfection; tungsten particle-facilitated microparticle bombardment (Johnston, Nature, 346: 776-777
(1990)); and strontium phosphate DNA co-precipitation (Brash et al., Mol. Cell Biol., 7: 2031-2034 (1987)).
1. AAV vectors
[0529] In some embodiments, the viral vector is an AAV vector. In some embodiments, the AAV vector is selected from among an AAV1, AAV2, AAV3, AAV4, AAV5, AAV6, AAV7, AAV8, AAV9, AAV10, AAV11, AAV12, AAV-DJ, or AAVrh74 vector. In some embodiments, the AAV vector is an AAV vector engineered for central nervous system (CNS) tropism. In some embodiments, the AAV vector is an AAV5 vector or an AAV9 vector. In some embodiments, the AAV vector is an AAV6 vector. In some aspects, the AAV vector is an AAV9 vector. In some aspects, the AAV vector is an AAV-DJ vector. In some embodiments, the AAV vector is an AAVrh74 vector.
[0530] In some embodiments, the AAV is selected or engineered for a desired tropism (e.g. for central nervous system tropism, or tropism for a heart cell, such as a cardiomyocyte, a skeletal muscle cell, a nervous system cell, such as a neuron, a fibroblast, or an induced pluripotent stem cell (iPSC)). In some embodiments, the AAV is exhibits tropism for a cardiomyocyte. In some embodiments, the AAV is exhibits tropism for a nervous system cell. In some embodiments, the AAV is exhibits tropism for a cell of the central nervous system (CNS). In some embodiments, the AAV is exhibits tropism for a neuron. In some embodiments, the AAV is exhibits tropism for a fibroblast. In some embodiments, the AAV is exhibits tropism for an iPSC. In some embodiments, the AAV exhibits tropism at least for any of the above cell types or a combination thereof.
[0531] In some aspects, nucleic acids or polynucleotides encoding any of the DNA-targeting systems, guide RNAs, eZFPs, fusion proteins (such as eZFP fusion proteins or dCas fusion proteins), or components, portions or combinations thereof can be delivered to cells or subjects using gene delivery vectors, such as viral vectors. In some aspects, provided herein are viral vectors that comprise any of the nucleic acids or polynucleotides described herein, any of the pluralities of nucleic acids or polynucleotides described herein, or a first polynucleotide or a second polynucleotide of any of the pluralities of polynucleotides described herein, or a portion or a component of any of the foregoing.
[0532] Examples of virions that can be employed to deliver any of the nucleic acids or polynucleotides provided herein include but are not limited to retroviral virions, lenti viral virions, adenovirus virions, herpes virus virions, alphavirus virions, and adeno-associated virus (AAV) virions. AAV is a 4.7 kb, single-stranded DNA virus. Recombinant virions based on AAV (rAAV virions) are associated with excellent clinical safety, since wild-type AAV is nonpathogenic and has no etiologic association with any known diseases. In addition, AAV offers the capability for highly efficient delivery and sustained expression of the delivered nucleic acid, composition or component thereof, in numerous tissues, including the nervous system, eye, muscle, lung and brain.
[0533] A “recombinant AAV vector (recombinant adeno-associated viral vector)” in some aspects refers to a polynucleotide vector comprising one or more heterologous sequences (i.e., nucleic acid
sequence not of AAV origin) that are flanked by at least one AAV inverted terminal repeat sequences (ITR). In some aspects, the recombinant nucleic acid is flanked by two inverted terminal repeat sequences (ITRs). In some aspects, ITRs are guanine-cytosine-rich sutructures that are involved in the replication and encapsidation of the AAV genome. Exemplary ITRs are set forth in SEQ ID NOS:434 and 435. In some aspects, the ITRs flank (e.g., are placed 5’ and 3’ to) the heterologous sequences for expression in the host cell. In some aspects, the 5’ and 3’ ITRs comprise the sequences set forth in SEQ ID NOS:434 and 435, respectively. Such recombinant viral vectors can be replicated and packaged into infectious viral particles when present in a host cell that has been infected with a suitable helper virus (or that is expressing suitable helper functions) and that is expressing AAV rep and cap gene products (i.e., AAV Rep and Cap proteins). When a recombinant viral vector is incorporated into a larger polynucleotide (e.g., in a chromosome or in another vector such as a plasmid used for cloning or transfection), then the recombinant viral vector may be referred to as a “pro-vector” which can be “rescued” by replication and encapsidation in the presence of AAV packaging functions and suitable helper functions. A recombinant viral vector can be in any of a number of forms, including, but not limited to, plasmids, linear artificial chromosomes, complexed with lipids, encapsulated within liposomes, and encapsidated in a viral particle, for example, an AAV particle. A recombinant viral vector can be packaged into an AAV virus capsid to generate a “recombinant adeno-associated viral particle (recombinant viral particle)”.
[0534] An “rAAV virus” or “rAAV viral particle” refers to a viral particle composed of at least one AAV capsid protein and an encapsidated rAAV vector genome.
[0535] “AAV helper functions” refer to functions that allow AAV to be replicated and packaged by a host cell for producing viruses. AAV helper functions can be provided in any of a number of forms, including, but not limited to, helper virus or helper virus genes which aid in AAV replication and packaging. Other AAV helper functions are known, such as genotoxic agents.
[0536] A “helper virus” for AAV refers to a virus that allows AAV (which is a defective parvovirus) to be replicated and packaged by a host cell for producing viruses. A helper virus provides “helper functions” which allow for the replication of AAV. A number of such helper viruses have been identified, including adenoviruses, herpesviruses, poxviruses such as vaccinia and baculovirus. The adenoviruses encompass a number of different subgroups, although Adenovirus type 5 of subgroup C (Ad5) is most commonly used. Numerous adenoviruses of human, non-human mammalian and avian origin are known and are available from depositories such as the ATCC. Viruses of the herpes family, which are also available from depositories such as ATCC, include, for example, herpes simplex viruses (HSV), Epstein-Barr viruses (EBV), cytomegaloviruses (CMV) and pseudorabies viruses (PRV). Examples of adenovirus helper functions for the replication of AAV include El A functions, E1B functions, E2A functions, VA functions and E4orf6 functions. Baculoviruses available from depositories include Autographa californica nuclear polyhedrosis virus.
[0537] A preparation of rAAV is said to be “substantially free” of helper virus if the ratio of
infectious AAV particles to infectious helper virus particles is at least about 102:1; at least about 104:1, at least about 106:1; or at least about 108:1 or more. In some aspects, preparations are also free of equivalent amounts of helper virus proteins (i.e., proteins as would be present as a result of such a level of helper virus if the helper virus particle impurities noted above were present in disrupted form). Viral and/or cellular protein contamination can generally be observed as the presence of Coomassie staining bands on SDS gels (e.g., the appearance of bands other than those corresponding to the AAV capsid proteins VP1, VP2 and VP3).
[0538] In some aspects, the recombinant viral particles for delivery of any of the provided nucleic acids, compositions or components thereof comprise a self-complementary AAV (sc A AV) genome. In some aspects, the recombinant AAV genome comprises a first heterologous polynucleotide sequence (e.g., coding strand) and a second heterologous polynucleotide sequence (e.g., the noncoding or antisense strand) wherein the first heterologous polynucleotide sequence can form intrastrand base pairs with the second polynucleotide sequence along most or all of its length. In some aspects, the first heterologous polynucleotide sequence and a second heterologous polynucleotide sequence are linked by a sequence that facilitates intrastrand base-pairing; e.g., a hairpin DNA structure. Hairpin structures are known, for example in siRNA molecules. In some aspects, the first heterologous polynucleotide sequence and a second heterologous polynucleotide sequence are linked by a mutated ITR. In some aspects, the scAAV viral particles comprise a monomeric form of an scAAV genome. In some aspects, the scAAV viral particles comprise the dimeric form of and scAAV genome. In some aspects, AUC as described herein is used to detect the presence of rAAV particles comprising the monomeric form of an scAAV genome. In some aspects, AUC as described herein is used to detect the presence of rAAV particles comprising the dimeric form of an scAAV genome. In some aspects, the packaging of scAAV genomes into capsid is monitored by AUC.
[0539] In some aspects, the rAAV particles comprise an AAV1 capsid, an AAV2 capsid, an AAV3 capsid, an AAV4 capsid, an AAV5 capsid, an AAV6 capsid (e.g., a wild-type AAV6 capsid, or a variant AAV6 capsid such as ShHIO, as described in US 2012/0164106), an AAV7 capsid, an AAV8 capsid, an AAVrh8 capsid, an AAVrh8R, an AAV9 capsid (e.g., a wild-type AAV9 capsid, or a modified AAV9 capsid as described in US 2013/0323226), an AAV10 capsid, an AAVrhlO capsid, an AAV11 capsid, an AAV12 capsid, a tyrosine capsid mutant, a heparin binding capsid mutant, an AAV2R471A capsid, an AAVAAV2/2-7m8 capsid, an AAV-DJ capsid (e.g., an AAV-DJ/8 capsid, an AAV-DJ/9 capsid, or any other AAV-DJ capsid, such as any of the capsids described, for example, in US 2012/0066783 or Mao, Y. et al., BMC Biotechnol. 16:1 (2016)), an AAV2 N587A capsid, an AAV2 E548A capsid, an AAV2 N708A capsid, an AAV V708K capsid, a goat AAV capsid, an AAV1/AAV2 chimeric capsid, a bovine AAV capsid, a mouse AAV capsid, or an AAV capsid described in US Pat. 8,283,151 or WO 2003/042397. In some of the above embodiments described herein, the rAAV particles comprise at least one AAV1 ITR, AAV2 ITR, AAV3 ITR, AAV4 ITR, AAV5 ITR, AAV6 ITR, AAV7 ITR, AAV8 ITR, AAVrh8 ITR, AAV9 ITR, AAV 10 ITR, AAVrhlO ITR, AAV 11 ITR, AAV 12 ITR, AAV-DJ ITR, goat
AAV ITR, bovine AAV ITR, or mouse AAV ITR. In some aspects, the rAAV particles comprise ITRs from one AAV serotype and AAV capsid from another serotype. For example, the rAAV particles may comprise the nucleic acid to be delivered (e.g., encoding any of the DNA-targeting systems, fusion proteins, gRNA, compositions or components thereof) flanked by at least one AAV2 ITR encapsidated into an AAV9 capsid. Such combinations may be referred to as pseudotyped rAAV particles. Exemplary AAV vectors include those described, for example, in WO 2020/113034, US 20220001028, US 20220001028, US 20210317474, and US 20160097061.
[0540] In some aspects, the viral particle is a recombinant AAV particle comprising a nucleic acid to be delivered flanked by one or two ITRs. The nucleic acid is encapsidated in the AAV particle. The AAV particle also comprises capsid proteins. In some aspects, the nucleic acid comprises the protein coding sequence or RNA-expressing sequences to be delivered (e.g., any of the DNA-targeting systems, fusion proteins, gRNA, compositions or components thereof) operatively linked components in the direction of transcription, control sequences including transcription initiation and termination sequences, thereby forming an expression cassette. The expression cassette is flanked on the 5' and 3' end by at least one functional AAV ITR sequences. By “functional AAV ITR sequences” it is meant that the ITR sequences function as intended for the rescue, replication and packaging of the AAV virion. See Davidson et al., PNAS, 2000, 97(7)3428-32; Passini et al., J. Virol., 2003, 77(12):7034-40; and Pechan et al., Gene Ther., 2009, 16:10-16, all of which are incorporated herein in their entirety by reference. For practicing some aspects of the invention, the recombinant vectors comprise at least all of the sequences of AAV essential for encapsidation and the physical structures for infection by the rAAV. AAV ITRs for use in the vectors of the invention need not have a wild-type nucleotide sequence (e.g., as described in Kotin, Hum. Gene Ther., 1994, 5:793-801), and may be altered by the insertion, deletion or substitution of nucleotides or the AAV ITRs may be derived from any of several AAV serotypes. More than 40 serotypes of AAV are currently known, and new serotypes and variants of existing serotypes continue to be identified. See Gao et al., PNAS, 2002, 99(18): 11854-6; Gao et al., PNAS, 2003, 100(10): 6081-6; and Bossis et al., J. Virol., 2003, 77(12):6799-810. Use of any AAV serotype is considered within the scope of the present invention. In some aspects, a rAAV vector is a vector derived from an AAV serotype, including without limitation, AAV1, AAV2, AAV3, AAV4, AAV5, AAV6, AAV7, AAV8, AAV9, AAVrh.8, AAVrh.10, AAV11, AAV12, a tyrosine capsid mutant, a heparin binding capsid mutant, an AAV2R471A capsid, an AAVAAV2/2-7m8 capsid, an AAV-DJ capsid, an AAV2 N587A capsid, an AAV2 E548A capsid, an AAV2 N708A capsid, an AAV V708K capsid, a goat AAV capsid, an AAV1/AAV2 chimeric capsid, a bovine AAV capsid, or a mouse AAV capsid, or the like. In some aspects, the nucleic acid in the AAV comprises an ITR of AAV1, AAV2, AAV3, AAV4, AAV5, AAV6, AAV7, AAV8, AAV9, AAVrh.8, AAVrhlO, AAV11, AAV12 or the like. In further embodiments, the rAAV particle comprises capsid proteins of AAV1, AAV2, AAV3, AAV4, AAV5, AAV6, AAV7, AAV8, AAV9, AAVrh.8, AAVrh.10, AAV11, AAV12 or the like. In further embodiments, the rAAV particle comprises capsid proteins of an AAV serotype from Clades A-F (Gao, et al. J. Virol. 2004,
78(12):6381).
[0541 ] Different AAV serotypes are used to optimize transduction of particular target cells or to target specific cell types within a particular target tissue (e.g., a diseased tissue). A rAAV particle can comprise viral proteins and viral nucleic acids of the same serotype or a mixed serotype. For example, a rAAV particle can comprise AAV9 capsid proteins and at least one AAV2 ITR or it can comprise AAV2 capsid proteins and at least one AAV9 ITR. In yet another example, a rAAV particle can comprise capsid proteins from both AAV9 and AAV2, and further comprise at least one AAV2 ITR. Any combination of AAV serotypes for production of a rAAV particle is provided herein as if each combination had been expressly stated herein.
[0542] In some aspects, the AAV comprises at least one AAV1 ITR and capsid protein from any of AAV-DJ, AAV2, AAV3, AAV4, AAV5, AAV6, AAV7, AAV8, AAV9, AAVrh.8, AAVrhlO, AAV11, and/or AAV12. In some aspects, the AAV comprises at least one AAV2 ITR and capsid protein from any of AAV-DJ, AAV1, AAV3, AAV4, AAV5, AAV6, AAV7, AAV8, AAV9, AAVrh.8, AAVrhlO, AAV11, and/or AAV12. In some aspects, the AAV comprises at least one AAV3 ITR and capsid protein from any of AAV-DJ, AAV1, AAV2, AAV4, AAV5, AAV6, AAV7, AAV8, AAV9, AAVrh.8, AAVrhlO, AAV11, and/or AAV12. In some aspects, the AAV comprises at least one AAV4 ITR and capsid protein from any of AAV-DJ, AAV1, AAV2, AAV3, AAV5, AAV6, AAV7, AAV8, AAV9, AAVrh.8, AAVrhlO, AAV11, and/or AAV12. In some aspects, the AAV comprises at least one AAV5 ITR and capsid protein from any of AAV-DJ, AAV1, AAV2, AAV3, AAV4, AAV6, AAV7, AAV8, AAV9, AAVrh.8, AAVrhlO, AAV11, and/or AAV12. In some aspects, the AAV comprises at least one AAV6 ITR and capsid protein from any of AAV-DJ, AAV1, AAV2, AAV3, AAV4, AAV5, AAV7, AAV8, AAV9, AAVrh.8, AAVrhlO, AAV11, and/or AAV12. In some aspects, the AAV comprises at least one AAV7 ITR and capsid protein from any of AAV-DJ, AAV1, AAV2, AAV3, AAV4, AAV5, AAV6, AAV8, AAV9, AAVrh.8, AAVrhlO, AAV11, and/or AAV12. In some aspects, the AAV comprises at least one AAV8 ITR and capsid protein from any of AAV-DJ, AAV1, AAV2, AAV3, AAV4, AAV5, AAV6, AAV7, AAV9, AAVrh.8, AAVrhlO, AAV11, and/or AAV12. In some aspects, the AAV comprises at least one AAV9 ITR and capsid protein from any of AAV-DJ, AAV1, AAV2, AAV3, AAV4, AAV5, AAV6, AAV7, AAV8, AAVrh.8, AAVrhlO, AAV11, and/or AAV12. In some aspects, the AAV comprises at least one AAVrh8 ITR and capsid protein from any of AAV-DJ, AAV1, AAV2, AAV3, AAV4, AAV5, AAV6, AAV8, AAV9, AAVrhlO, AAV11, and/or AAV12. In some aspects, the AAV comprises at least one AAVrhlO ITR and capsid protein from any of AAV-DJ, AAV1, AAV2, AAV3, AAV4, AAV5, AAV6, AAV7, AAV8, AAV9, AAV11, and/or AAV12. In some aspects, the AAV comprises at least one AAV11 ITR and capsid protein from any of AAV-DJ, AAV1, AAV2, AAV3, AAV4, AAV5, AAV6, AAV7, AAV8, AAVrh8, AAV9, AAVrhlO, and/or AAV12. In some aspects, the AAV comprises at least one AAV12 ITR and capsid protein from any of AAV-DJ, AAV1, AAV2, AAV3, AAV4, AAV5, AAV6, AAV7, AAV8, AAV rh8, AAV9, AAVrhlO, and/or AAV11. In some aspects, the AAV comprises at least one AAV-DJ ITR and capsid protein from any of AAV1,
AAV2, AAV3, AAV4, AAV5, AAV6, AAV7, AAV8, AAV rh8, AAV9, AAVrhlO, and/or AAV11. [0543] In some aspects, the viral particles comprise a recombinant self-complementing genome.
AAV viral particles with self-complementing genomes and methods of use of self-complementing AAV genomes are described in US Patent Nos. 6,596,535; 7,125,717; 7,765,583; 7,785,888; 7,790,154; 7,846,729; 8,093,054; and 8,361,457; and Wang Z., et al., (2003) Gene Ther 10:2105-2111, each of which are incorporated herein by reference in its entirety. A rAAV comprising a self-complementing genome will quickly form a double stranded DNA molecule by virtue of its partially complementing sequences (e.g., complementing coding and non-coding strands). In some aspects, an AAV viral particle comprises an AAV genome, wherein the rAAV genome comprises a first heterologous polynucleotide sequence (e.g., a coding strand) and a second heterologous polynucleotide sequence (e.g., the noncoding or antisense strand) wherein the first heterologous polynucleotide sequence can form intrastrand base pairs with the second polynucleotide sequence along most or all of its length. In some aspects, the first heterologous polynucleotide sequence and a second heterologous polynucleotide sequence are linked by a sequence that facilitates intrastrand base-pairing; e.g., a hairpin DNA structure. Hairpin structures include, for example in siRNA molecules. In some aspects, the first heterologous polynucleotide sequence and a second heterologous polynucleotide sequence are linked by a mutated ITR (e.g., the right ITR). The mutated ITR comprises a deletion of the D region comprising the terminal resolution sequence. As a result, on replicating an AAV viral genome, the rep proteins will not cleave the viral genome at the mutated ITR and as such, a recombinant viral genome comprising the following in 5' to 3' order will be packaged in a viral capsid: an AAV ITR, the first heterologous polynucleotide sequence including regulatory sequences, the mutated AAV ITR, the second heterologous polynucleotide in reverse orientation to the first heterologous polynucleotide and a third AAV ITR.
10544] Methods for production of rAAV vectors, including transfection, stable cell line production, and infectious hybrid virus production systems which include adenovirus-AAV hybrids, herpesvirus- AAV hybrids (Conway, JE et al., (1997) J. Virology 71(ll):8780-8789) and baculovirus-AAV hybrids can be employed. Typically, rAAV production cultures for the production of rAAV virus particles all require; 1) suitable host cells, including, for example, human-derived cell lines such as HeLa, A549, or 293 cells, or insect-derived cell lines such as SF-9, in the case of baculovirus production systems; 2) suitable helper virus function, provided by wild-type or mutant adenovirus (such as temperature sensitive adenovirus), herpes virus, baculovirus, or a plasmid construct providing helper functions; 3) AAV rep and cap genes and gene products; 4) a nucleic acid to be delivered (such as any of the DNA-targeting systems, fusion proteins, compositions or components thereof) flanked by at least one AAV ITR sequences; and 5) suitable media and media components to support rAAV production. In some aspects, the AAV rep and cap gene products may be from any AAV serotype. In general, but not obligatory, the AAV rep gene product is of the same serotype as the ITRs of the rAAV vector genome as long as the rep gene products may function to replicated and package the rAAV genome. Suitable media may be used for the production of rAAV vectors. These media include, without limitation, media produced by
Hyclone Laboratories and JRH including Modified Eagle Medium (MEM), Dulbecco's Modified Eagle Medium (DMEM), custom formulations such as those described in U.S. Patent No. 6,566,118, and Sf- 900 II SFM media as described in U.S. Patent No. 6,723,551. In some aspects, the AAV helper functions are provided by adenovirus or HSV. In some aspects, the AAV helper functions are provided by baculovirus and the host cell is an insect cell (e.g., Spodoptera frugiperda (Sf9) cells).
[0545] Suitable rAAV production culture media of the present invention may be supplemented with serum or serum-derived recombinant proteins at a level of 0.5%-20% (v/v or w/v). Alternatively, rAAV vectors may be produced in serum-free conditions which may also be referred to as media with no animal-derived products. Commercial or custom media designed to support production of rAAV vectors may also be supplemented with one or more cell culture components, including without limitation glucose, vitamins, amino acids, and or growth factors, in order to increase the titer of rAAV in production cultures.
[0546] rAAV production cultures can be grown under a variety of conditions (over a wide temperature range, for varying lengths of time, and the like) suitable to the particular host cell being utilized. rAAV production cultures include attachment-dependent cultures which can be cultured in suitable attachment-dependent vessels such as, for example, roller bottles, hollow fiber filters, microcarriers, and packed-bed or fluidized-bed bioreactors. rAAV vector production cultures may also include suspension-adapted host cells such as HeLa, 293, and SF-9 cells which can be cultured in a variety of ways including, for example, spinner flasks, stirred tank bioreactors, and disposable systems such as the Wave bag system.
[0547] rAAV vector particles of the invention may be harvested from rAAV production cultures by lysis of the host cells of the production culture or by harvest of the spent media from the production culture, provided the cells are cultured under conditions to cause release of rAAV particles into the media from intact cells, as described in U.S. Patent No. 6,566,118). Suitable methods of lysing cells include for example multiple freeze/thaw cycles, sonication, microfluidization, and treatment with chemicals, such as detergents and/or proteases.
[0548] In some aspects, recombinant viral particles for delivery of the nucleic acids, compositions or components thereof are highly purified, suitably buffered, and concentrated. In some aspects, the viral particles are concentrated to at least about 1 x 107 vg/mL to about 9 x 1013 vg/mL or any concentration therebetween.
[0549] In some aspects, adeno-associated virus (AAV)-based vectors are generally used vector system for neurologic gene therapy, with an excellent safety record established in multiple clinical trials (Kaplitt et al., (2007) Lancet 369:2097-2105; Eberling et al., (2008) Neurology 70:1980-1983; Fiandaca et al., (2009) Neuroimage 47 Suppl. 2:T27-35). In some cases, effective treatment of neurologic disorders has been hindered by problems associated with the delivery of AAV vectors to affected cell populations. This delivery issue has been especially problematic for disorders involving the cerebral cortex. Simple injections do not distribute AAV vectors effectively, relying on diffusion, which is effective only within
a 1- to 3-mm radius. An alternative method, convection-enhanced delivery (CED) (Nguyen et al., (2003) J. Neurosurg. 98:584-590), has been used clinically in gene therapy (AAV2-hAADC) for Parkinson's disease (Fiandaca et al., (2008) Exp. Neurol. 209:51-57). The underlying principle of CED involves pumping infusate into brain parenchyma under sufficient pressure to overcome the hydrostatic pressure of interstitial fluid, thereby forcing the infused particles into close contact with the dense perivasculature of the brain. Pulsation of these vessels acts as a pump, distributing the particles over large distances throughout the parenchyma (Hadaczek et al., (2006) Hum. Gene Ther. 17:291-302). To increase the safety and efficacy of CED a reflux-resistant cannula (Krauze et al., (2009) Methods Enzymol. 465:349- 362) can be employed along with monitored delivery with real-time MRI. Monitored delivery allows for the quantification and control of aberrant events, such as cannula reflux and leakage of infusate into ventricles (Eberling et al., (2008) Neurology 70:1980-1983; Fiandaca et al., (2009) Neuroimage 47 Suppl. 2:T27-35; Saito et al., (2011) Journal of Neurosurgery Pediatrics 7:522-526).
[0550] In some aspects, the nucleic acid to be delivered is operably linked to a promoter. In some aspects, the promoter expresses the nucleic acid to be delivered in a cell of the CNS. In some aspects, the promoter expresses the nucleic acid to be delivered in a brain cell. In some aspects, the promoter expresses the nucleic acid to be delivered in a neuron and/or a glial cell. In some aspects, the neuron is a medium spiny neuron of the caudate nucleus, a medium spiny neuron of the putamen, a neuron of the cortex layer IV and/or a neuron of the cortex layer V. In some aspects, the glial cell is an astrocyte. In some aspects, the promoter is a CBA promoter, a minimum CBA promoter, a CMV promoter or a GUSB promoter. In some aspects, the promoter is inducible. In further embodiments, the rAAV vector comprises one or more of an enhancer, a splice donor/splice acceptor pair, a matrix attachment site, or a polyadenylation signal.
[0551] In some aspects, the methods for delivering a recombinant adeno-associated viral (rAAV) particle to the central nervous system of a subject involve administering the rAAV particle to the striatum, wherein the rAAV particle comprises a rAAV vector encoding a nucleic acid to be delivered that is expressed in at least the cerebral cortex and striatum of the subject. In some aspects, methods for delivering a rAAV particle to the central nervous system of a subject involve administering the rAAV particle to the striatum, wherein the rAAV particle comprises an rAAV vector encoding a nucleic acid to be delivered that is expressed in at least the cerebral cortex and striatum of the subject and wherein the rAAV particle comprises an AAV serotype 1 (AAV1) capsid. In some aspects, methods for delivering a rAAV particle to the central nervous system of a subject comprise administering the rAAV particle to the striatum, wherein the rAAV particle comprises an rAAV vector encoding a nucleic acid to be delivered that is expressed in at least the cerebral cortex and striatum of the subject and wherein the rAAV particle comprises an AAV serotype 2 (AAV2) capsid. In some aspects, methods for treating a central nervous system-related disease in a subject involve administering a rAAV particle to the striatum, wherein the rAAV particle comprises a rAAV vector encoding a nucleic acid to be delivered that is expressed in at least the cerebral cortex and striatum of the subject. In some aspects, the subject is a human.
[0552] In some aspects, a rAAV particle is administered to one or more regions of the central nervous system (CNS). In some aspects, the rAAV particle is administered to the striatum. The striatum is known as a region of the brain that receives inputs from the cerebral cortex (the term “cortex” may be used interchangeably herein) and sends outputs to the basal ganglia (the striatum is also referred to as the striate nucleus and the neostriatum). In some aspects, the striatum controls both motor movements and emotional control/motivation and has been implicated in many neurological diseases, such as Huntington’s disease. Several cell types of interest are located in the striatum, including without limitation spiny projection neurons (also known as medium spiny neurons), GABAergic interneurons, and cholinergic interneurons. Medium spiny neurons make up most of the striatal neurons. These neurons are GABAergic and express dopamine receptors. Each hemisphere of the brain contains a striatum.
[0553] In some aspects, important substructures of the striatum include the caudate nucleus and the putamen. In some aspects, the rAAV particle is administered to the caudate nucleus (the term “caudate” may be used interchangeably herein). The caudate nucleus is known as a structure of the dorsal striatum. The caudate nucleus has been implicated in control of functions such as directed movements, spatial working memory, memory, goal-directed actions, emotion, sleep, language, and learning. Each hemisphere of the brain contains a caudate nucleus.
[0554] In some aspects, the rAAV particle is administered to the putamen. Along with the caudate nucleus, the putamen is known as a structure of the dorsal striatum. The putamen comprises part of the lenticular nucleus and connects the cerebral cortex with the substantia nigra and the globus pallidus. Highly integrated with many other structures of the brain, the putamen has been implicated in control of functions such as learning, motor learning, motor performance, motor tasks, and limb movements. Each hemisphere of the brain contains a putamen.
[0555] In some aspects, rAAV particles may be administered to one or more sites of the striatum. In some aspects, the rAAV particle is administered to the putamen and the caudate nucleus of the striatum. In some aspects, the rAAV particle is administered to the putamen and the caudate nucleus of each hemisphere of the striatum. In some aspects, the rAAV particle is administered to at least one site in the caudate nucleus and two sites in the putamen.
[0556] In some aspects, the rAAV particle is administered to one hemisphere of the brain. In some aspects, the rAAV particle is administered to both hemispheres of the brain. For example, in some aspects, the rAAV particle is administered to the putamen and the caudate nucleus of each hemisphere of the striatum. In some aspects, the composition containing rAAV particles is administered to the striatum of each hemisphere. In some aspects, the composition containing rAAV particles is administered to striatum of the left hemisphere or the striatum of the right hemisphere and/or the putamen of the left hemisphere or the putamen of the right hemisphere. In some aspects, the composition containing rAAV particles is administered to any combination of the caudate nucleus of the left hemisphere, the caudate nucleus of the right hemisphere, the putamen of the left hemisphere and the putamen of the right
hemisphere.
[0557] In some aspects, the methods involving administration to CNS an effective amount of recombinant viral particles to the striatum can be employed for delivery, wherein the rAAV particle comprises a rAAV vector encoding a nucleic acid to be delivered that is expressed in at least the cerebral cortex and striatum. In some aspects, the viral titer of the rAAV particles is at least about any of 5 x 1012, 6 x 1012, 7 x 1012, 8 x 1012, 9 x 1012, 10 x 1012, 11 x 1012, 15 x 1012, 20 x 1012, 25 x 1012, 30 x 1012, or 50 x 1012 genome copies/mL. In some aspects, the viral titer of the rAAV particles is about any of 5 x 1012 to 6 x 1012, 6 x 1012to 7 x 1012, 7 x 1012 to 8 x 1012, 8 x 1012 to 9 x 1012, 9 x 1012 to 10 x 1012, 10 x 1012 to 11 x 1012, 11 x 1012 to 15 x 1012, 15 x 1012 to 20 x 1012, 20 x 1012 to 25 x 1012, 25 x 1012 to 30 x 1012, 30 x 1012 to 50 x 1012, or 50 x 1012 to 100 x 1012 genome copies/mL. In some aspects, the viral titer of the rAAV particles is about any of 5 x 1012 to 10 x 1012, 10 x 1012 to 25 x 1012, or 25 x 1012 to 50 x 1012 genome copies/mL. In some aspects, the viral titer of the rAAV particles is at least about any of 5 x 109,
6 x 109, 7 x 109, 8 x 109, 9 x 109, 10 x 109, 11 x 109, 15 x 109, 20 x 109, 25 x 109, 30 x 109, or 50 x 109 transducing units/mL. In some aspects, the viral titer of the rAAV particles is about any of 5 x 109 to 6 x 109, 6 x 109 to 7 x 109, 7 x 109 to 8 x 109, 8 x 109 to 9 x 109, 9 x 109 to 10 x 109, 10 x 109 to 11 x 109, 11 x 109 to 15 x 109, 15 x 109 to 20 x 109, 20 x 109 to 25 x 109, 25 x 109 to 30 x 109, 30 x 109 to 50 x 109 or 50 x 109 to 100 x 109 transducing units/mL. In some aspects, the viral titer of the rAAV particles is about any of 5 x 109 to 10 x 109, 10 x 109 to 15 x 109, 15 x 109 to 25 x 109, or 25 x 109 to 50 x 109 transducing units/mL. In some aspects, the viral titer of the rAAV particles is at least any of about 5 x 1010, 6 x IO10,
7 x IO10, 8 x IO10, 9 x IO10, 10 x IO10, 11 x IO10, 15 x IO10, 20 x IO10, 25 x IO10, 30 x IO10, 40 x IO10, or 50 x IO10 infectious units/mL. In some aspects, the viral titer of the rAAV particles is at least any of about 5 x IO10 to 6 x IO10, 6 x IO10 to 7 x IO10, 7 x IO10 to 8 x IO10, 8 x IO10 to 9 x IO10, 9 x IO10 to 10 x IO10, 10 x IO10 to 11 x IO10, 11 x IO10 to 15 x IO10, 15 x IO10 to 20 x IO10, 20 x IO10 to 25 x IO10, 25 x IO10 to 30 x IO10, 30 x IO10 to 40 x IO10, 40 x IO10 to 50 x IO10, or 50 x IO10 to 100 x IO10 infectious units/mL. In some aspects, the viral titer of the rAAV particles is at least any of about 5 x IO10 to 10 x IO10, 10 x IO10 to 15 x IO10, 15 x IO10 to 25 x IO10, or 25 x IO10 to 50 x IO10 infectious units/mL.
[0558] In some aspects, an effective amount of recombinant viral particles is administered to the striatum, wherein the rAAV particle comprises a rAAV vector encoding a nucleic acid to be delivered that is expressed in at least the cerebral cortex and striatum. In some aspects, the dose of viral particles administered to the individual is at least about any of 1 x 108 to about 1 x 1013 genome copies/kg of body weight. In some aspects, the dose of viral particles administered to the individual is about 1 x 108 to 1 x 1013 genome copies/kg of body weight.
[0559] In some aspects, an effective amount of recombinant viral particles is administered to the striatum, wherein the rAAV particle comprises a rAAV vector encoding a nucleic acid to be delivered that is expressed in at least the cerebral cortex and striatum. In some aspects, the total amount of viral particles administered to the individual is at least about 1 x 109 to about 1 x 1014 genome copies. In some aspects, the total amount of viral particles administered to the individual is about 1 x 109 to about 1 x
1014 genome copies.
2. Exemplary A A V Vectors
[0560] In some embodiments, an AAV vector provided herein can comprise nucleic acids encoding any of the eZFPs, fusion proteins (such as eZFP fusion proteins or dCas fusion proteins), gRNAs, DNA- targeting systems provided herein, or a portion or component thereof, or a combination of any of the foregoing. In some embodiments, the AAV vector comprises a single polynucleotide encoding any CRISPR/Cas-based DNA-targeting system provided herein, such as a DNA-targeting system comprising a) a fusion protein comprising (i) a deactivated Cas (dCas) protein and (ii) at least one effector domain that increases transcription of a frataxin (FXN) locus; and b) a guide RNA (gRNA) comprising a gRNA spacer sequence that is capable of hybridizing to a target site in a regulatory DNA element of a FXN locus and/or is complementary to the target site. IN some embodiments, the AAV vector is an AAV6, AAV9, AAV-DJ, or AAVrh74. In some embodiments, the AAV vector is an AAV6 vector. In some embodiments, the AAV vector is an AAV9 vector. In some embodiments, the AAV vector is an AAV-DJ vector. In some embodiments, the AAV vector is an AAVrh74 vector.
[0561] In some embodiments, the AAV vector comprises a nucleic acid, such as a first nucleic acid, comprising a sequence encoding a fusion protein, such as any fusion protein provided herein. In some embodiments, the fusion protein comprises a dCas protein (such as dSaCas9) and at least one epigenetic effector domain for increasing transcription of a FXN locus. In some embodiments, the fusion protein is a dSaCas9-2xVP64 fusion protein. In some embodiments, the fusion protein is encoded by the sequence set forth in SEQ ID NO:69, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the fusion protein is encoded by the sequence set forth in SEQ ID NO:69. In some embodiments, the fusion protein comprises the amino acid sequence set forth in SEQ ID NO:71, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the fusion protein comprises the amino acid sequence set forth in SEQ ID NO: 71. In some embodiments, the nucleic acid, such as the first nucleic acid, further comprises a promoter operably linked to the sequence encoding the fusion protein, i.e. a promoter for the fusion protein. In some embodiments, the promoter for the fusion protein is any suitable promoter, such as a promoter described herein. In some embodiments, the promoter is a human elongation factor alpha short (EFS) promoter. In some embodiments, the promoter comprises the sequence set forth in SEQ ID NO:436, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the promoter comprises the sequence set forth in SEQ ID NO:436. In some embodiments, the promoter is a CAG promoter. In some embodiments, the promoter comprises the sequence set forth in SEQ ID NO:602, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the promoter comprises the sequence set forth in SEQ ID NO:602. In some embodiments, the promoter is an EFla
promoter. In some embodiments, the promoter comprises the sequence set forth in SEQ ID NO:603, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the promoter comprises the sequence set forth in SEQ ID NO: 603. In some embodiments, the sequence encoding the fusion protein is operably linked to a poly(A) site, such as any suitable poly(A) site. In some embodiments, the poly(A) site is a synthetic (such as engineered) poly(A) (SpA) (See, e.g., Levitt N, Briggs D, Gil A, Proudfoot NJ. “Definition of an efficient synthetic poly(A) site.” Genes Dev. 1989;3:1019-1025). In some embodiments, the SpA is engineered for efficient expression of the operably linked coding sequence. In some embodiments, the SpA is engineered to have a small size (i.e. small number of nucleotides), which facilitates AAV packaging. In some embodiments, the SpA comprises the sequence set forth in SEQ ID NO:437 or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the SpA comprises the sequence set forth in SEQ ID NO:437. In some embodiments, the poly(A) site is bovine growth hormone (bGH) poly(A). In some embodiments, the bGH poly(A) comprises the sequence set forth in SEQ ID NO:604 or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the bGH poly(A) comprises the sequence set forth in SEQ ID NO:604. In some embodiments, the nucleic acid encoding the fusion protein further comprises any suitable additional regulatory element, such as a regulatory element that promotes expression of the fusion protein. In some embodiments, the nucleic acid encoding the fusion protein further comprises a woodchuck hepatitis virus post-transcriptional regulatory element (WPRE) element (see, e.g., Choi JH, et al., “Optimization of AAV expression cassettes to improve packaging capacity and transgene expression in neurons.” Mol Brain. 2014 Mar 11 ;7: 17. doi: 10.1186/1756-6606-7-17.) In some embodiments, the WPRE element is placed downstream of the sequence encoding the fusion protein, proximal to the poly(A) site. In some embodiments, the WPRE element comprises the sequence set forth in SEQ ID NO:438, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the WPRE element comprises the sequence set forth in SEQ ID NO:438.
[0562] In some embodiments, the nucleic acid encoding the fusion protein further comprises a 5’ untranslated region (UTR). In some embodiments, the 5’ UTR is placed downstream of the sequence encoding the promoter, such as the CAG or EFla promoter. In some embodiments, the 5’ UTR comprises the sequence set forth in SEQ ID NO:605, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the 5’ UTR comprises the sequence set forth in SEQ ID NO: 605.
[0563] In some embodiments, the AAV vector comprises a nucleic acid, such as a second nucleic acid, comprising a sequence encoding a gRNA, such as any gRNA provided herein. In some embodiments, the gRNA targets the target site set forth in SEQ ID NO:1, 7, or 21. In some embodiments, the gRNA comprises a gRNA spacer sequence set forth in SEQ ID NO:22, 28, or 42. In some embodiments, the gRNA comprises a scaffold sequence set forth in SEQ ID NO:44. In some
embodiments, the gRNA comprises the sequence set forth in SEQ ID NO:47, 54, or 67. In some embodiments, the gRNA gRNA A, gRNA G, or gRNA U, as described herein, e.g. in the Examples. In some embodiments, the gRNA is operably linked to a promoter for the gRNA. In some embodiments, the promoter for the gRNA is a promoter that is recognized by RNA polymerase III (Pol III), such as a U6 promoter. In some embodiments, the U6 promoter comprises the sequence set forth in SEQ ID NO:432, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the U6 promoter comprises the sequence set forth in SEQ ID NO:432. In some embodiments, the promoter is an engineered U6 promoter, such as a mini-U6 promoter. In some embodiments, the mini-U6 promoter comprises the sequence set forth in SEQ ID NO:433, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the mini-U6 promoter comprises the sequence set forth in SEQ ID NO:433.
[0564] In some embodiments, the AAV vector comprises a nucleic acid comprising a sequence encoding a fusion protein, such as any fusion protein provided herein. In some embodiments, the fusion protein comprises an eZFP protein (such as eZFP_A31) and at least one epigenetic effector domain for increasing transcription of a FXN locus. In some embodiments, the fusion protein is an eZFP_A31- NCOA3-FOXO3-NCOA3 fusion protein. In some embodiments, the fusion protein is encoded by the sequence set forth in SEQ ID NO:427, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the fusion protein is encoded by the sequence set forth in SEQ ID NO: 427. In some embodiments, the fusion protein comprises the amino acid sequence set forth in SEQ ID NO:333, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the fusion protein comprises the amino acid sequence set forth in SEQ ID NO:333. In some embodiments, the nucleic acid further comprises a promoter operably linked to the sequence encoding the fusion protein, i.e. a promoter for the fusion protein. In some embodiments, the promoter for the fusion protein is any suitable promoter, such as a promoter described herein. In some embodiments, the promoter is a human elongation factor alpha short (EFS) promoter. In some embodiments, the promoter comprises the sequence set forth in SEQ ID NO:436, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the promoter comprises the sequence set forth in SEQ ID NO:436. In some embodiments, the promoter is a CAG promoter. In some embodiments, the promoter comprises the sequence set forth in SEQ ID NO:602, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the promoter comprises the sequence set forth in SEQ ID NO:602. In some embodiments, the promoter is a EFla promoter. In some embodiments, the promoter comprises the sequence set forth in SEQ ID NO:603, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the promoter comprises the sequence set forth in SEQ
ID NO: 603. In some embodiments, the sequence encoding the fusion protein is operably linked to a poly(A) site, such as any suitable poly(A) site. In some embodiments, the poly(A) site is a synthetic (such as engineered) poly(A) (SpA) (See, e.g., Levitt N, Briggs D, Gil A, Proudfoot NJ. “Definition of an efficient synthetic poly(A) site.” Genes Dev. 1989;3:1019-1025). In some embodiments, the SpA is engineered for efficient expression of the operably linked coding sequence. In some embodiments, the SpA is engineered to have a small size (i.e. small number of nucleotides), which facilitates AAV packaging. In some embodiments, the SpA comprises the sequence set forth in SEQ ID NO:437 or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the SpA comprises the sequence set forth in SEQ ID NO:437. In some embodiments, the poly(A) site is bovine growth hormone (bGH) poly(A). In some embodiments, the bGH poly(A) comprises the sequence set forth in SEQ ID NO:604 or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the bGH poly(A) comprises the sequence set forth in SEQ ID NO:604. In some embodiments, the nucleic acid encoding the fusion protein further comprises any suitable additional regulatory element, such as a regulatory element that promotes expression of the fusion protein. In some embodiments, the nucleic acid encoding the fusion protein further comprises a woodchuck hepatitis virus post-transcriptional regulatory element (WPRE) element (see, e.g., Choi JH, et al., “Optimization of AAV expression cassettes to improve packaging capacity and transgene expression in neurons.” Mol Brain. 2014 Mar 11 ;7: 17. doi: 10.1186/1756-6606-7-17.) In some embodiments, the WPRE element is placed downstream of the sequence encoding the fusion protein, proximal to the poly(A) site. In some embodiments, the WPRE element comprises the sequence set forth in SEQ ID NO:438, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the WPRE element comprises the sequence set forth in SEQ ID NO:438. In some embodiments, the nucleic acid encoding the fusion protein furth comprises a 5’ untranslated region (UTR). In some embodiments, the 5’ UTR is placed downstream of the promoter, such as a CAG or EFla promoter. In some embodiments, the 5’ UTR comprises the sequence set forth in SEQ ID NO:605, or a nucleotide sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto. In some embodiments, the 5’ UTR comprises the sequence set forth in SEQ ID NO:605.
[0565] In some embodiments, the AAV vector comprises a single polynucleotide, such as a single polynucleotide comprising the first nucleic acid or second nucleic acid described above in this section. In some embodiments, the AAV vector comprises a single polynucleotide comprising the first nucleic acid and the second nucleic acid. In some embodiments, the single polynucleotide comprises, in the 5’ to 3’ direction, the promoter for the fusion protein, the sequence encoding the fusion protein, the SpA site, the promoter for the gRNA, and the sequence encoding the gRNA. In some embodiments, the single polynucleotide further comprises the WPRE element in between the sequence encoding the fusion protein and the SpA site. In some embodiments, the promoter for the fusion protein is an EFS promoter.
In some embodiments, the fusion protein is dSaCas9-2xVP64. In some embodiments, the promoter for the gRNA is a mini-U6 promoter. In some embodiments, the gRNA gRNA A, gRNA G, or gRNA U, as described herein. In some embodiments, the single polynucleotide comprises a 5’ITR and a 3’ITR that flank the first and second nucleic acids. In some embodiments, the 5’ITR and 3’ITR are set forth in SEQ ID NO:434 and 435, respectively. In some embodiments, the AAV vector comprises a single polynucleotide, such as a single polynucleotide comprising a nucleic acid comprising a sequence encoding a fusion protein, such as any fusion protein comprising an eZFP protein provided herein. In some embodiments, the single polynucleotide, in the 5’ to 3’ direction, the promoter for the fusion protein, the sequence encoding the fusion protein, and the poly(A) site. In some embodiments, the single polynucleotide further comprises the 5’ UTR between the promoter and the sequence encoding the fusion protein. In some embodiments, the single polynucleotide further comprises a NLS. In some embodiments, the NLS comprises a SV40 NLS set forth in SEQ ID NO: 159. In some embodiments, the NLS comprises a c-myc NLS set forth in SEQ ID NO: 160. In some embodiments, the NLS is upstream of the sequence encoding the fusion protein. In some embodiments, the NLS is between the sequence encoding the fusion protein. In some embodiments, the poly(A) site is a SpA set forth in SEQ ID NO:337. In some embodiments, the poly(A) site is a bGH set forth in SEQ ID NO:604.
3. Non- viral vectors
[0566] In some embodiments, the vector is a non-viral vector. In some aspects, exemplary non-viral vectors include polymers, lipids, peptides, inorganic materials, and hybrid systems. In some aspects, the non-viral vector is a lipid nanoparticle (LNP), a liposome, an exosome, or a cell penetrating peptide. In some aspects, the non-viral vector is a lipid nanoparticle (LNP). In some aspects, the LNP can be used for delivery to the liver. Exemplary non-viral vectors include those described in WO 2020/051561, US 20210301274, Zu et al., The AAPS Journal volume 23, Article number: 78 (2021), and Sung et al., Biomaterials Research volume 23, Article number: 8 (2019), Nyamay’Antu et al., Cell & Gene Therapy Insights 2019; 5(S 1):51 -57, and Yin et al., Nature Reviews Genetics 15:541-555 (2014).
[0567] In some embodiments, the vector is a non-viral vector selected from: a lipid nanoparticle, a liposome, an exosome, or a cell penetrating peptide.
[0568] In some embodiments, a vector described herein is or comprises a lipid nanoparticle (LNP). In some embodiments, any of the eZFPs, eZFP fusion proteins, DNA-targeting systems, gRNAs, Cas- gRNA combinations, polynucleotides, fusion proteins, or components thereof described herein, are incorporated in lipid nanoparticles (LNPs), such as for delivery. In some embodiments, the lipid nanoparticle is a vector for delivery. In some embodiments, the nanoparticle may comprise at least one lipid. The lipid may be selected from, but is not limited to, DLin-DMA, DLin-K-DMA, 98N12- 5, C12- 200, DLin-MC3-DMA, DLin-KC2-DMA, DODMA, PLGA, PEG, PEG-DMG and PEGylated lipids. In another aspect, the lipid may be a cationic lipid such as, but not limited to, DLin-DMA, DLin-D-DMA, DLin-MC 3 -DMA, DLin-KC2-DMA and DODMA.
[0569] Lipid nanoparticles can be used for the delivery of encapsulated or associated (e.g., complexed) therapeutic agents, including nucleic acids and proteins, such as those encoding and/or comprising CRISPR/Cas systems. See, e.g., US Patent No. 10,723,692, US Patent No. 10,941,395, and WO 2015/035136.
[0570] In some embodiments, the provided methods involve use of a lipid nanoparticle (LNP) comprising mRNA, such as mRNA encoding a protein component of any of the provided DNA-targeting systems, for example any of the fusion proteins provided herein. In some embodiments, the mRNA can be produced using methods known in the art such as in vitro transcription. In some embodiments of the method, the mRNA comprises a 5' cap. In some embodiments, the 5’ cap is an altered nucleotide on the 5’ end of primary transcripts such as messenger RNA. In some aspects, the 5’ caps of the mRNA improves one or more of RNA stability and processing, mRNA metabolism, the processing and maturation of an RNA transcript in the nucleus, transport of mRNA from the nucleus to the cytoplasm, mRNA stability, and efficient translation of mRNA to protein. In some embodiments, a 5’ cap can be a naturally-occurring 5’ cap or one that differs from a naturally-occurring cap of an mRNA. A 5’ cap may be any 5' cap known to a skilled artisan. In certain embodiments, the 5' cap is selected from the group consisting of an Anti-Reverse Cap Analog (ARCA) cap, a 7-methyl-guanosine (7mG) cap, a CleanCap® analog, a vaccinia cap, and analogs thereof. For instance, the 5’ cap may include, without limitation, an anti-reverse cap analogs (ARCA) (US7074596), 7-methyl-guanosine, CleanCap® analogs, such as Cap 1 analogs (Trilink; San Diego, CA), or enzymatically capped using, for example, a vaccinia capping enzyme or the like. In some embodiments, the mRNA may be polyadenylated. The mRNA may contain various 5’ and 3’ untranslated sequence elements to enhance expression of the encoded protein and/or stability of the mRNA itself. Such elements can include, for example, posttranslational regulatory elements such as a woodchuck hepatitis virus post-transcriptional regulatory element (WPRE). In some embodiments, the mRNA comprises at least one nucleoside modification. The mRNA may contain modifications of naturally-occurring nucleosides to nucleoside analogs. Any nucleoside analogs known in the art are envisioned. Such nucleoside analogs can include, for example, those described in US 8,278,036. In certain embodiments of the method, the nucleoside modification is selected from the group consisting of a modification from uridine to pseudouridine and uridine to Nl- methyl pseudouridine. In particular embodiments of the method the nucleoside modification is from uridine to pseudouridine.
[0571] In some embodiments, LNPs useful for in the present methods comprise a cationic lipid selected from DLin-DMA ( 1 ,2-dilinoleyloxy-3 -dimethylaminopropane), DLin-MC3 -DM A (dilinoleylmethyl-4-dimethylaminobutyrate) , DLin-KC2-DMA (2,2-dilinoleyl-4-(2- dimethylaminoethyl)-[l,3]-dioxolane), DODMA (1,2- dioleyloxy-N,N-dimethyl-3- aminopropane), SS- OP (Bis[2-(4-{2-[4-(cis-9 octadecenoyloxy)phenylacetoxy]ethyl}piperidinyl)ethyl] disulfide), and derivatives thereof. DLin-MC3-DMA and derivatives thereof are described, for example, in WO 2010/144740. DODMA and derivatives thereof are described, for example, in US 7,745,651 and Mok et al. (1999), Biochimica et Biophysica Acta, 1419(2): 137-150. DLin-DMA and derivatives thereof are
described, for example, in US 7,799,565. DLin-KC2-DMA and derivatives thereof are described, for example, in US 9,139,554. SS-OP (NOF America Corporation, White Plains, NY) is described, for example, at https://www.nofamerica.com/store/index.php?dispatch=products.view&product_id=962. Additional and non-limiting examples of cationic lipids include methylpyridiyl-dialkyl acid (MPDACA), palmitoyl-oleoyl-nor-arginine (PONA), guanidino-dialkyl acid (GUADACA), l,2-di-0-octadecenyl-3- trimethylammonium propane (DOTMA), 1,2- dioleoyl-3-trimethylammonium-propane (DOTAP), Bis{2- [N-methyl-N-(a-D- tocopherolhemisuccinatepropyl) amino] ethyl} disulfide (SS-33/3AP05), Bis{2-[4-(a- D- tocopherolhemisuccinateethyl)piperidyl] ethyl} disulfide (SS33/4PE15), Bis{2-[4-(cis-9- octadecenoateethyl)-l-piperidinyl] ethyl} disulfide (SS18/4PE16), and Bis{2-[4-(cis,cis-9,12- octadecadienoateethyl)-l-piperidinyl] ethyl} disulfide (SS18/4PE13). In further embodiments, the lipid nanoparticles also comprise one or more non-cationic lipids and a lipid conjugate.
[0572} In some embodiments, the molar concentration of the cationic lipid is from about 20% to about 80%, from about 30% to about 70%, from about 40% to about 60%, from about 45% to about 55%, or about 20%, about 25%, about 30%, about 35%, about 40%, about 45%, about 50%, about 55%, about 60%, about 65%, about 70%, about 75%, or about 80% of the total lipid molar concentration, wherein the total lipid molar concentration is the sum of the cationic lipid, the non-cationic lipid, and the lipid conjugate molar concentrations. In certain embodiments, the lipid nanoparticles comprise a molar ratio of cationic lipid to any of the polynucleotides of from about 1 to about 20, from about 2 to about 16, from about 4 to about 12, from about 6 to about 10, or about 1, about 2, about 3, about 4, about 5, about 6, about 7, about 8, about 9, about 10, about 11, about 12, about 13, about 14, about 15, about 16, about 17, about 18, about 19, or about 20.
[0573] In some embodiments, the lipid nanoparticles can comprise at least one non-cationic lipid. In particular embodiments, the molar concentration of the non-cationic lipids is from about 20% to about 80%, from about 30% to about 70%, from about 40% to about 70%, from about 40% to about 60%, from about 46% to about 50%, or about 20%, about 25%, about 30%, about 35%, about 40%, about 45%, about 48.5%, about 50%, about 55%, about 60%, about 65%, about 70%, about 75%, or about 80% of the total lipid molar concentration. Non-cationic lipids include, in some embodiments, phospholipids and steroids.
[0574] In some embodiments, phospholipids useful for the lipid nanoparticles described herein include, but are not limited to, l,2-Distearoyl-sn-glycero-3-phosphocholine (DSPC), 1,2-Didecanoyl-sn- glycero-3- phosphocholine (DDPC), l,2-Dierucoyl-sn-glycero-3-phosphate(Sodium Salt) (DEPA-NA), l,2-Dierucoyl-sn-glycero-3-phosphocholine (DEPC), l,2-Dierucoyl-sn-glycero-3- phosphoethanolamine (DEPE), l,2-Dierucoyl-sn-glycero-3[Phospho-rac-(l-glycerol)(Sodium Salt) (DEPG-NA), 1,2-Dilinoleoyl- sn-glycero-3-phosphocholine (DLOPC), 1,2-Dilauroyl-sn- glycero-3-phosphate(Sodium Salt) (DLPA- NA), l,2-Dilauroyl-sn-glycero-3-phosphocholine (DLPC), l,2-Dilauroyl-sn-glycero-3- phosphoethanolamine (DLPE), 1,2-Dilauroyl-sn- glycero-3[Phospho-rac-(l-glycerol.)(Sodium Salt)
(DLPG-NA), 1 ,2-Dilauroyl-sn-glycero- 3[Phospho-rac-(l-glycerol)(Ammonium Salt) (DLPG-NH4), 1,2-
Dilauroyl-sn-glycero-3- phosphoserine(Sodium Salt) (DLPS-NA), l,2-Dimyristoyl-sn-glycero-3- phosphate(SodiumSalt) (DMPA-NA), l,2-Dimyristoyl-sn-glycero-3-phosphocholine (DMPC), 1,2- Dimyristoyl- sn-glycero-3-phosphoethanolamine (DMPE), l,2-Dimyristoyl-sn-glycero-3 [Phospho-rac-(l- glycerol)(Sodium Salt) (DMPG-NA), l,2-Dimyristoyl-sn-glycero-3[Phospho-rac-(l- glycerol)( Ammonium Salt) (DMPG-NH4), l,2-Dimyristoyl-sn-glycero-3[Phospho-rac-(l- glycerol)(Sodium/ Ammonium Salt) (DMPG-NH4/NA), l,2-Dimyristoyl-sn-glycero-3- phosphoserine(Sodium Salt) (DMPS-NA), l,2-Dioleoyl-sn-glycero-3-phosphate(Sodium Salt) (DOPA- NA), l,2-Dioleoyl-sn-glycero-3-phosphocholine (DOPC), 1 ,2-Dioleoyl-sn- glycero-3- phosphoethanolamine (DOPE), l,2-Dioleoyl-sn-glycero-3[Phospho-rac-(l- glycerol)(Sodium Salt) (DOPG-NA), l,2-Dioleoyl-sn-glycero-3-phosphoserine(Sodium Salt) (DOPS-NA), 1,2-Dipalmitoyl-sn- glycero-3-phosphate(Sodium Salt) (DPPA-NA), 1,2- Dipalmitoyl-sn-glycero-3-phosphocholine (DPPC), 1 ,2-Dipalmitoyl-sn-glycero-3-phosphoethanolamine (DPPE) , 1 ,2-Dipalmitoyl-sn-glycero- 3 [Phospho- rac-(l-glycerol)(Sodium Salt) (DPPG-NA), 1 ,2-Dipalmitoyl-sn-glycero- 3[Phospho-rac-(l- glycerol)(Ammonium Salt) (DPPG-NH4), l,2-Dipalmitoyl-sn-glycero-3- phosphoserine(Sodium Salt) (DPPS-NA), l,2-Distearoyl-sn-glycero-3-phosphate(Sodium Salt) (DSPA-NA), 1,2-Distearoyl-sn- glycero-3-phosphoethanolamine (DSPE), 1 ,2- Distearoyl-sn-glycero-3 [Phospho-rac-(l-glycerol)(Sodium Salt) (DSPG-NA), 1 ,2-Distearoyl- sn-glycero-3[Phospho-rac-(l-glycerol)(Ammonium Salt) (DSPG- NH4), 1 ,2-Distearoyl-sn- glycero-3-phosphoserine(Sodium Salt) (DSPS-NA), Egg-PC (EPC), Hydrogenated Egg PC (HEPC), Hydrogenated Soy PC (HSPC), l-Myristoyl-sn-glycero-3- phosphocholine (LY S OPCM YRIS TIC ), l-Palmitoyl-sn-glycero-3-phosphocholine (LYSOPCPALMITIC), 1- Stearoyl-sn-glycero-3-phosphocholine (LYSOPC STEARIC), l-Myristoyl-2- palmitoyl-sn- glycero3 -phosphocholine (MPPC), l-Myristoyl-2-stearoyl-sn-glycero-3-phosphocholine (MSPC), l-Palmitoyl-2-myristoyl-sn-glycero-3-phosphocholine (PMPC), l-Palmitoyl-2- oleoyl-sn- glycero-3-phosphocholine (POPC), l-Palmitoyl-2-oleoyl-sn-glycero-3- phosphoethanolamine (POPE), 1- Palmitoyl-2-oleoyl-sn-glycero-3[Phospho-rac-(l- glycerol)] (Sodium Salt) (POPG-NA), l-Palmitoyl-2- stearoyl-sn-glycero-3-phosphocholine (PS PC), l-Stearoyl-2-myristoyl-sn-glycero-3-phosphocholine (SMPC), l-Stearoyl-2-oleoyl- sn-glycero-3-phosphocholine (SOPC), and l-Stearoyl-2-palmitoyl-sn- glycero-3- phosphocholine (SPPC). In particular embodiments, the phospholipid is DSPC. In particular embodiments, the phospholipid is DOPE. In particular embodiments, the phospholipid is DOPC.
[0575] In some embodiments, the non-cationic lipids comprised by the lipid nanoparticles include one or more steroids. Steroids useful for the lipid nanoparticles described herein include, but are not limited to, cholestanes such as cholesterol, cholanes such as cholic acid, pregnanes such as progesterone, androstanes such as testosterone, and estranes such as estradiol. Further steroids include, but are not limited to, cholesterol (ovine), cholesterol sulfate, desmosterol-d6, cholesterol-d7, lathosterol-d7, desmosterol, stigmasterol, lanosterol, dehydrocholesterol, dihydrolanosterol, zymosterol, lathosterol, zymosterol-d5, 14-demethyl-lanosterol, 14-demethyl-lanosterol-d6, 8(9)- dehydrocholesterol, 8(14)- dehydrocholesterol, diosgenin, DHEA sulfate, DHEA, lanosterol- d6, dihydrolanosterol-d7, campesterol-
d6, sitosterol, lanosterol-95, Dihydro FF-MAS-d6, zymostenol-d7, zymostenol, sitostanol, campestanol, campesterol, 7-dehydrodesmosterol, pregnenolone, sitosterol-d7, Dihydro T-MAS, Delta 5-avenasterol, Brassicasterol, Dihydro FF-MAS, 24-methylene cholesterol, cholic acid derivatives, cholesteryl esters, and glycosylated sterols. In particular embodiments, the lipid nanoparticles comprise cholesterol.
[0576] In some embodiments, the lipid nanoparticles comprise a lipid conjugate. Such lipid conjugates include, but are not limited to, ceramide PEG derivatives such as C8 PEG2000 ceramide, C16 PEG2000 ceramide, C8 PEG5000 ceramide, C16 PEG5000 ceramide, C8 PEG750 ceramide, and C16 PEG750 ceramide, phosphoethanolamine PEG derivatives such as 16:0 PEG5000PE, 14:0 PEG5000 PE, 18:0 PEG5000 PE, 18:1 PEG5000 PE, 16:0 PEG3000 PE, 14:0 PEG3000 PE, 18:0 PEG3000 PE, 18:1 PEG3000 PE, 16:0 PEG2000 PE, 14:0 PEG2000 PE, 18:0 PEG2000 PE, 18:1 PEG2000 PE 16:0 PEG1000 PE, 14:0 PEG1000 PE, 18:0 PEG1000 PE, 18:1 PEG 1000 PE, 16:0 PEG750 PE, 14:0 PEG750 PE, 18:0 PEG750 PE, 18:1 PEG750 PE, 16:0 PEG550 PE, 14:0 PEG550 PE, 18:0 PEG550 PE, 18:1 PEG550 PE, 16:0 PEG350 PE, 14:0 PEG350 PE, 18:0 PEG350 PE, and 18:1 PEG350, sterol PEG derivatives such as Chol-PEG600, and glycerol PEG derivatives such as DMG-PEG5000, DSG- PEG5000, DPG- PEG5000, DMG-PEG3000, DSG-PEG3000, DPG-PEG3000, DMG-PEG2000, DSG- PEG2000, DPG-PEG2000, DMG-PEG1000, DSG-PEG1000, DPG-PEG1000, DMG- PEG750, DSG- PEG750, DPG-PEG750, DMG-PEG550, DSG-PEG550, DPG-PEG550, DMG-PEG350, DSG-PEG350, and DPG-PEG350. In some embodiments, the lipid conjugate is a DMG-PEG. In some particular embodiments, the lipid conjugate is DMG- PEG2000. In some particular embodiments, the lipid conjugate is DMG-PEG5000.
[0577] It is within the level of a skilled artisan to select the cationic lipids, non-cationic lipids and/or lipid conjugates which comprise the lipid nanoparticle, as well as the relative molar ratio of such lipids to each other, such as based upon the characteristics of the selected lipid(s), the nature of the delivery to the intended target cells, and the characteristics of the nucleic acids and/or proteins to be delivered. Additional considerations include, for example, the saturation of the alkyl chain, as well as the size, charge, pH, pKa, fusogenicity and toxicity of the selected lipid(s). Thus, the molar ratios of each individual component may be adjusted accordingly.
[0578] The lipid nanoparticles for use in the method can be prepared by various techniques which are known to a skilled artisan. Nucleic acid-lipid particles and methods of preparation are disclosed in, for example, U.S. Patent Publication Nos. 20040142025 and 20070042031.
[0579] In some embodiments, the lipid nanoparticles will have a size within the range of about 25 to about 500 nm. In some embodiments, the lipid nanoparticles have a size from about 50 nm to about 300 nm, or from about 60 nm to about 120 nm. The size of the lipid nanoparticles may be determined by quasi-electric light scattering (QELS) as described in Bloomfield, Ann. Rev. Biophys. Bioeng., 10: 421A150 (1981). A variety of methods are known in the art for producing a population of lipid nanoparticles of particular size ranges, for example, sonication or homogenization. One such method is described in U.S. Pat. No. 4,737,323.
[0580] In some embodiments, the lipid nanoparticles comprise a cell targeting molecule such as, for example, a targeting ligand (e.g., antibodies, scFv proteins, DART molecules, peptides, aptamers, and the like) anchored on the surface of the lipid nanoparticle that selectively binds the lipid nanoparticles to the targeted cell, such as any cell described herein.
[0581] In some embodiments, the vector exhibits tropism for one or more cell types. For example, the vector may exhibit liver cell and/or hepatocyte tropism, neural cell (e.g. neuron or glia) tropism, immune cell tropism, or tropism for any suitable cell type.
[0582] In some aspects, provided herein are pluralities of vectors that comprise any of the vectors described herein, and one or more additional vectors. In some embodiments, the one or more additional vectors comprise one or more additional polynucleotides encoding any additional eZFP, eZFP fusion protein, transcriptional activation domain, multipartite effector such as multipartite activator, DNA- targeting domain, gRNA, fusion protein, DNA-targeting system, or a portion, component, or combination thereof. In some aspects, provided are pluralities of vectors, that include: a first vector comprising any of the polynucleotides described herein; a second vector comprising any of the polynucleotides described herein; and optionally one or more additional vectors comprising any of the polynucleotides described herein.
[0583] In some aspects, vectors provided herein may be referred to as delivery vehicles. In some aspects, any of the eZFPs, eZFP fusion proteins, DNA-targeting systems, components thereof, or polynucleotides disclosed herein can be packaged into or on the surface of delivery vehicles for delivery to cells. Delivery vehicles contemplated include, but are not limited to, nanospheres, liposomes, quantum dots, nanoparticles, polyethylene glycol particles, hydrogels, and micelles. As described in the art, a variety of targeting moieties can be used to enhance the preferential interaction of such vehicles with desired cell types or locations.
[0584] Any suitable method of introducing a nucleic acid into a host cell may be used, including methods that can be used to introduce a nucleic acid (e.g., an expression construct) into a cell. Suitable methods include, include e.g., viral or bacteriophage infection, transfection, conjugation, protoplast fusion, lipofection, electroporation, calcium phosphate precipitation, polyethyleneimine (PEI) -mediated transfection, DEAE-dextran mediated transfection, liposome-mediated transfection, particle gun technology, calcium phosphate precipitation, direct micro injection, nanoparticle-mediated nucleic acid delivery, and the like. In some embodiments, the composition may be delivered by mRNA delivery and ribonucleoprotein (RNP) complex delivery. Direct delivery of the RNP complex, including the DNA- targeting domain complexed with the sgRNA, can eliminate the need for intracellular transcription and translation and can offer a robust platform for host cells with low transcriptional and translational activity. The RNP complexes can be introduced into the host cell by any of the methods known in the art.
[0585] Nucleic acids or RNPs of the disclosure can be incorporated into a host using virus-like particles (VLP). VLPs contain normal viral vector components, such as envelope and capsids, but lack the viral genome. For instance, nucleic acids expressing the Cas and sgRNA can be fused to the viral
vector components such as gag and introduced into producer cells. The resulting virus-like particles containing the sgRNA-expressing vectors can infect the host cell for efficient editing.
[0586] Introduction of the complexes, polypeptides, and nucleic acids of the disclosure can occur by protein transduction domains (PTDs). PTDs, including the human immunodeficiency virus- 1 TAT, herpes simplex virus- 1 VP22, Drsophila Antennapedia Antp, and the poluarginines, are peptide sequences that can cross the cell membrane, enter a host cell, and deliver the complexes, polypeptides, and nucleic acids into the cell.
[0587] Introduction of the complexes, polypeptides, and nucleic acids of the disclosure into cells can occur by viral or bacteriophage infection, transfection, conjugation, protoplast fusion, lipofection, electroporation, nucleofection, calcium phosphate precipitation, polyethyleneimine (PEI)-mediated transfection, DEAE-dextran mediated transfection, liposome-mediated transfection, particle gun technology, calcium phosphate precipitation, direct micro-injection, nanoparticle-mediated nucleic acid delivery, and the like, for example as described in WO 2017/193107, WO 2016/123578, WO 2014/152432, WO 2014/093661, WO 2014/093655, or WO 2021/226555.
[0588] Various methods for the introduction of polynucleotides are well known and may be used with the provided methods and compositions. Exemplary methods include those for transfer of polynucleotides encoding the eZFPs, eZFP fusion proteins, and/or DNA targeting systems provided herein, including via viral, e.g., retroviral or lentiviral, transduction, transposons, and electroporation.
C. Pharmaceutical Compositions and Formulations
[0589] Also provided are compositions, such as pharmaceutical compositions and formulations for administration, that include any of the eZFPs, eZFP fusion proteins, AAV vectors, and/or DNA-targeting systems described herein, any of the gRNAs described herein, any of the combinations described herein, any of the fusion proteins described herein, any of the polynucleotides described herein, any of the pluralities of polynucleotides described herein, any of the vectors described herein, any of the pluralities of vectors described herein, or a portion or a component of any of the foregoing. In some aspects, the pharmaceutical composition comprises one or more pharmaceutically acceptable carriers.
[0590] In some aspects, the pharmaceutical composition contains one or more eZFP, eZFP fusion protein, AAV vectors, and/or DNA-targeting system provided herein or a component thereof. In some aspects, the pharmaceutical composition comprises one or more vectors, e.g., viral vectors that contain polynucleotides that encode one or more components of the eZFPs, eZFP fusion proteins, AAV vectors, and/or DNA-targeting systems provided herein. Such compositions can be used in accord with the provided methods, and/or with the provided articles of manufacture or compositions, such as in the prevention or treatment of diseases, conditions, and disorders, or in detection, diagnostic, and prognostic methods.
[0591] The term “pharmaceutical formulation” refers to a preparation which is in such form as to permit the biological activity of an active ingredient contained therein to be effective, and which contains
no additional components which are unacceptably toxic to a subject to which the formulation would be administered.
[0592] A “pharmaceutically acceptable carrier” refers to an ingredient in a pharmaceutical formulation, other than an active ingredient, which is nontoxic to a subject. A pharmaceutically acceptable carrier includes, but is not limited to, a buffer, excipient, stabilizer, or preservative.
[0593] In some aspects, the choice of carrier is determined in part by the particular cell or agent and/or by the method of administration. Accordingly, there are a variety of suitable formulations. For example, the pharmaceutical composition can contain preservatives. Suitable preservatives may include, for example, methylparaben, propylparaben, sodium benzoate, and benzalkonium chloride. In some aspects, a mixture of two or more preservatives is used. The preservative or mixtures thereof are typically present in an amount of about 0.0001% to about 2% by weight of the total composition. Carriers are described, e.g., by Remington’s Pharmaceutical Sciences 16th edition, Osol, A. Ed. (1980). Pharmaceutically acceptable carriers are generally nontoxic to recipients at the dosages and concentrations employed, and include, but are not limited to: buffers such as phosphate, citrate, and other organic acids; antioxidants including ascorbic acid and methionine; preservatives (such as octadecyldimethylbenzyl ammonium chloride; hexamethonium chloride; benzalkonium chloride; benzethonium chloride; phenol, butyl or benzyl alcohol; alkyl parabens such as methyl or propyl paraben; catechol; resorcinol; cyclohexanol; 3-pentanol; and m-cresol); low molecular weight (less than about 10 residues) polypeptides; proteins, such as serum albumin, gelatin, or immunoglobulins; hydrophilic polymers such as polyvinylpyrrolidone; amino acids such as glycine, glutamine, asparagine, histidine, arginine, or lysine; monosaccharides, disaccharides, and other carbohydrates including glucose, mannose, or dextrins; chelating agents such as EDTA; sugars such as sucrose, mannitol, trehalose or sorbitol; salt-forming counter-ions such as sodium; metal complexes (e.g. Zn-protein complexes); and/or non-ionic surfactants such as polyethylene glycol (PEG).
[0594] The pharmaceutical composition in some embodiments contains components in amounts effective to treat or prevent the disease or condition, such as a therapeutically effective or prophylactically effective amount. Therapeutic or prophylactic efficacy in some embodiments is monitored by periodic assessment of treated subjects. For repeated administrations over several days or longer, depending on the condition, the treatment is repeated until a desired suppression of disease symptoms occurs. However, other dosage regimens may be useful and can be determined. The desired dosage can be delivered by a single bolus administration of the composition, by multiple bolus administrations of the composition, or by continuous infusion administration of the composition.
[0595] The composition can be administered by any suitable means, for example, by bolus infusion, by injection, e.g., intravenous or subcutaneous injections, intraocular injection, periocular injection, subretinal injection, intravitreal injection, trans-septal injection, subscleral injection, intrachoroidal injection, intracameral injection, subconjectval injection, subconjuntival injection, sub-Tenon’s injection, retrobulbar injection, peribulbar injection, or posterior juxtascleral delivery. In some embodiments, they
are administered by parenteral, intrapulmonary, and intranasal, and, if desired for local treatment, intralesional administration. Parenteral infusions include intramuscular, intravenous, intraarterial, intraperitoneal, or subcutaneous administration. In some embodiments, a given dose is administered by a single bolus administration of the composition. In some embodiments, it is administered by multiple bolus administrations of the composition, for example, over a period of no more than 3 days, or by continuous infusion administration of the composition.
[0596] For the prevention or treatment of disease, the appropriate dosage may depend on the type of disease to be treated, the type of agent or agents, the type of cells or recombinant receptors, the severity and course of the disease, whether the agent or cells are administered for preventive or therapeutic purposes, previous therapy, the subject’s clinical history and response to the agent or the cells, and the discretion of the attending physician. The compositions are in some embodiments suitably administered to the subject at one time or over a series of treatments.
[0597] Formulations include those for oral, intravenous, intraperitoneal, subcutaneous, pulmonary, transdermal, intramuscular, intranasal, buccal, sublingual, or suppository administration. In some embodiments, the agent or cell populations are administered parenterally. The term “parenteral,” as used herein, includes intravenous, intramuscular, subcutaneous, rectal, vaginal, and intraperitoneal administration. In some embodiments, the agent or cell populations are administered to a subject using peripheral systemic delivery by intravenous, intraperitoneal, or subcutaneous injection.
[0598] Compositions in some embodiments are provided as sterile liquid preparations, e.g., isotonic aqueous solutions, suspensions, emulsions, dispersions, or viscous compositions, which may in some aspects be buffered to a selected pH. Liquid preparations are normally easier to prepare than gels, other viscous compositions, and solid compositions. Additionally, liquid compositions are somewhat more convenient to administer, especially by injection. Viscous compositions, on the other hand, can be formulated within the appropriate viscosity range to provide longer contact periods with specific tissues. Liquid or viscous compositions can comprise carriers, which can be a solvent or dispersing medium containing, for example, water, saline, phosphate buffered saline, polyol (for example, glycerol, propylene glycol, liquid polyethylene glycol) and suitable mixtures thereof.
[0599] Sterile injectable solutions can be prepared by incorporating the agent or cells in a solvent, such as in admixture with a suitable carrier, diluent, or excipient such as sterile water, physiological saline, glucose, dextrose, or the like.
[0600] The formulations to be used for in vivo administration are generally sterile. Sterility may be readily accomplished, e.g., by filtration through sterile filtration membranes.
IV. METHODS OF MODULATING AND METHODS OF TREATMENT
[0601 [ Provided herein are methods of treatment, e.g., including administering any of the compositions, such as pharmaceutical compositions described herein. In some aspects, also provided are methods of administering any of the compositions described herein to a subject, such as a subject that has
a disease or disorder. The compositions, such as pharmaceutical compositions, described herein are useful in a variety of therapeutic, diagnostic and prophylactic indications. For example, the compositions are useful in treating a variety of diseases and disorders in a subject. Such methods and uses include therapeutic methods and uses, for example, involving administration of the compositions, to a subject having a disease, condition, or disorder, such as Friedreich’s ataxia (FA). In some embodiments, the compositions are administered in an effective amount to effect treatment of the disease or disorder. Uses include uses of the compositions in such methods and treatments, and in the preparation of a medicament in order to carry out such therapeutic methods. In some embodiments, the methods are carried out by administering the compositions, to the subject having or suspected of having the disease or condition. In some embodiments, the methods thereby treat the disease or condition or disorder in the subject. Also provided are therapeutic methods for administering the cells and compositions to subjects, e.g., patients.
[0602] Also provided herein are methods for modulating the expression of frataxin (FXN) in a cell, that involve: introducing any of the eZFPs described herein, any of the eZFP fusion proteins described herein, any of the DNA-targeting systems described herein, any of the gRNAs described herein, any of the combinations described herein, any of the fusion proteins described herein, any of the polynucleotides described herein, any of the pluralities of polynucleotides described herein, any of the vectors described herein including any of the AAV vectors described herein, any of the pluralities of vectors described herein, or a portion or a component of any of the foregoing, into the cell.
[0603] In some embodiments, the cell is from a subject that has or is suspected of having Friedreich’s ataxia (FA).
[0604] Also provided herein are methods for modulating the expression of frataxin (FXN) in a subject, the method comprising: administering any of the eZFPs described herein, any of the eZFP fusion proteins described herein, any of the DNA-targeting systems described herein, any of the gRNAs described herein, any of the combinations of gRNAs described herein, any of the fusion proteins described herein, any of the polynucleotides described herein, any of the pluralities of polynucleotides described herein, any of the vectors described herein including any of the AAV vectors, any of the plurality of vectors described herein, or a portion or a component of any of the foregoing, to the subject.
[0605] In some embodiments, the subject has or is suspected of having Friedreich’s ataxia (FA).
[0606] Also provided herein are methods of treating Friedreich’s ataxia (FA), the method comprising: administering any of the eZFPs described herein, any of the eZFP fusion proteins described herein, any of the DNA-targeting systems described herein, any of the gRNAs described herein, any of the combinations described herein, any of the fusion proteins described herein, any of the polynucleotides described herein, any of the pluralities of polynucleotides described herein, any of the vectors described herein including any of the AAV vectors, any of the pluralities of vectors described herein, or a portion or a component of any of the foregoing, to a subject that has or is suspected of having FA.
[0607] In some embodiments, a cell in the subject exhibits reduced expression of FXN compared to a cell from a normal subject. In some embodiments, a cell in the subject has a GAA trinucleotide repeat
expansion in the FXN gene. In some embodiments, the cell is a heart cell, a skeletal muscle cell, a nervous system cell, or an induced pluripotent stem cell. In some embodiments, the introducing, contacting or administering is carried out in vivo or ex vivo. In some embodiments, following the introducing, contacting or administering, the expression of frataxin (FXN) is increased in the cell or the subject. In some embodiments, the expression of frataxin (FXN) is increased at least about 1.2-fold, 1.25- fold, 1.3-fold, 1.4-fold, 1.5-fold, 1.6-fold, 1.7-fold, 1.75-fold, 1.8-fold, 1.9-fold, 2-fold, 2.5-fold, 3-fold, 4-fold, or 5-fold. In some embodiments, the expression is increased by less than about 10-fold, 9-fold, 8- fold, 7-fold or 6-fold. In some embodiments, the subject is a human.
[0608] Also provided herein is a pharmaceutical composition comprising any of the eZFPs described herein, any of the eZFP fusion proteins described herein, any of the DNA-targeting systems described herein, any of the gRNAs described herein, any of the combinations described herein, any of the fusion proteins described herein, any of the polynucleotides described herein, any of the pluralities of polynucleotides described herein, any of the vectors described herein including any of the AAV vectors described herein, any of the pluralities of vectors described herein, or a portion or a component of any of the foregoing.
[0609] In some of any of the provided embodiments, the pharmaceutical composition is for use in treating Friedreich’s ataxia (FA). In some of any of the provided embodiments, the pharmaceutical composition is for use in the manufacture of a medicament for treating Friedreich’s ataxia (FA).
[0610] In some of any of the provided embodiments, the pharmaceutical composition is to be administered to a subject.
[0611] Also provided herein is the use of any of the pharmaceutical compositions described herein for treating Friedreich’s ataxia (FA).
[0612] Also provided herein is the use of any of the pharmaceutical compositions described herein in the manufacture of a medicament for treating Friedreich’s ataxia (FA).
[0613] In some of any embodiments, the pharmaceutical composition is to be administered to a subject.
[0614] In some embodiments, the subject has or is suspected of having Friedreich’s ataxia (FA). In some embodiments, a cell in the subject exhibits reduced expression of FXN compared to a cell from a normal subject. In some embodiments, a cell in the subject has a GAA trinucleotide repeat expansion in the FXN gene.
[0615] In some embodiments, the cell is a heart cell, a skeletal muscle cell, a nervous system cell, or an induced pluripotent stem cell. In some embodiments, the administration is carried out in vivo or ex vivo.
[0616] In some embodiments, following the administration, the expression of frataxin (FXN) is increased in the cell or the subject. In some embodiments, the expression is increased at least about 1.2- fold, 1.25-fold, 1.3-fold, 1.4-fold, 1.5-fold, 1.6-fold, 1.7-fold, 1.75-fold, 1.8-fold, 1.9-fold, 2-fold, 2.5- fold, 3-fold, 4-fold, or 5-fold. In some embodiments, the expression is increased by less than about 10-
fold, 9-fold, 8-fold, 7-fold or 6-fold. In some embodiments, the subject is a human.
[0617] Also provided herein are cells comprising any of the eZFPs described herein, any of the eZFP fusion proteins described herein, any of the DNA-targeting systems described herein, any of the gRNAs described herein, any of the combinations described herein, any of the fusion proteins described herein, any of the polynucleotides described herein, any of the pluralities of polynucleotides described herein, any of the vectors described herein, any of the pluralities of vectors described herein, or a portion or a component of any of the foregoing.
[0618] In some embodiments, the cell is a heart cell, a skeletal muscle cell, a nervous system cell, or an induced pluripotent stem cell. In some embodiments, the cell is from a subject that has or is suspected of having Friedreich’s ataxia (FA).
[0619| In some aspects, methods of treating of treating a disease or disorder, such as diseases or disorders associated with dysregulation or reduced activity, function or expression of FXN, such as Friedreich’s ataxia (FA), in an individual or a subject, involve administering to the individual or the subject AAV particles. The AAV particles may be administered to a particular tissue of interest, or it may be administered systemically. In some aspects, an effective amount of the AAV particles may be administered parenterally. Parenteral routes of administration may include without limitation intravenous, intraosseous, intra-arterial, intracerebral, intramuscular, intrathecal, subcutaneous, intracerebro ventricular, and others. In some aspects, an effective amount of AAV particles may be administered through one route of administration. In some aspects, an effective amount of AAV particles may be administered through a combination of more than one route of administration. In some aspects, the individual is a mammal. In some aspects, the individual is a human.
[0620] An effective amount of AAV particles comprising an oversized AAV genome is administered, depending on the objectives of treatment. For example, where a low percentage of transduction can achieve the desired therapeutic effect, then the objective of treatment is generally to meet or exceed this level of transduction. In some instances, this level of transduction can be achieved by transduction of only about 1 to 5% of the target cells of the desired tissue type, In some aspects at least about 20% of the cells of the desired tissue type, In some aspects at least about 50%, In some aspects at least about 80%, In some aspects at least about 95%, In some aspects at least about 99% of the cells of the desired tissue type. As a guide, the number of particles administered per injection is generally between about 1 x 106 and about 1 x 1014 particles, between about 1 x 107 and 1 x 1013 particles, between about 1 x 109 and 1 x 1012 particles or about 1 x 109 particles, about 1 x 1010 particles, or about 1 x 10” particles. The rAAV composition may be administered by one or more administrations, either during the same procedure or spaced apart by days, weeks, months, or years. One or more of any of the routes of administration described herein may be used. In some aspects, multiple vectors may be used to treat the human.
[0621 ] Methods to identify cells transduced by AAV viral particles can be employed; for example, immunohistochemistry or the use of a marker such as enhanced green fluorescent protein can be used to
detect transduction of viral particles; for example viral particles comprising a rAAV capsid with one or more substitutions of amino acids.
[0622] In some aspects the AAV viral particles comprising an oversized AAV genome with are administered to more than one location simultaneously or sequentially. In some aspects, multiple injections of rAAV viral particles are no more than one hour, two hours, three hours, four hours, five hours, six hours, nine hours, twelve hours or 24 hours apart.
[0623] In some aspects, provided are compositions, methods and related uses, that can be employed to modulate the expression of FXN, such as in a cell or a subject. In some aspects, the provided compositions, methods and uses can be employed to activate or increase the expression of FXN in the cell or the subject. In some aspects, the subject has or is suspected of having a disease or disorder associated with reduced activity, inactivation, mutation and/or dysregulation of expression of the FXN gene, such as FA. In some aspects, disease or disorders such as Friedreich's Ataxia (FA) is associated with reduced FXN expression levels compared to in a normal individual. In some aspects, by modulating, such as by activating or increasing the expression of FXN, the provided compositions, methods and uses can be employed to treat or ameliorate the disease or disorder associated with reduced activity, mutation and/or dysregulation of FXN.
[0624] In some aspects, by modulating, such as by activating or increasing the expression of FXN, the provided compositions, methods and uses can be employed to restore or recover the expression or activity of FXN in a subject or a cell with a disease or disorder associated with reduced activity, mutation and/or dysregulation of FXN, such that the expression or activity of FXN is increased at least about 1.2- fold, 1.25-fold, 1.3-fold, 1.4-fold, 1.5-fold, 1.6-fold, 1.7-fold, 1.75-fold, 1.8-fold, 1.9-fold, 2-fold, 2.5- fold, 3-fold, 4-fold, or 5-fold, compared to the expression or activity of FXN in the subject or cell with the disease or disorder in the absence of the provided compositions or uses. In some aspects, the expression or activity is increased by less than about 10-fold, 9-fold, 8-fold, 7-fold or 6-fold. In some aspects, by modulating, such as by activating or increasing the expression of FXN, the provided compositions, methods and uses can be employed to restore or recover the expression or activity of FXN in a subject or a cell with a disease or disorder associated with reduced activity, mutation and/or dysregulation of FXN, such that the expression or activity of FXN is increased to at least at or about 30%, 40%, 50%, 60%, 70%, 75%, 80%, 85%, 90%, 95%, 100%, 105%, 110%, 120%, 125%, 150%, 175%, 200%, 225%, 250%, 300%, 400%, or 500%, of the expression or activity of FXN in an individual or a cell without the disease or disorder or in a wild-type cell. Increasing the expression of FXN mRNA and/or protein, can lead to recovery or restoration of expression of the FXN gene and be used for treatment and/or prevention of such diseases or disorders.
[0625] In certain cases, overexpression of FXN can also lead to toxicity in cells and tissues, organ dysregulation and/or reduction of life span (see, e.g., Vannocci et al., Disease Models & Mechanisms (2018) 11, dmm032706; Belbellaa et al., Molecular Therapy - Methods & Clinical Development (2020) 19:120-138; Navarro et al., PLoS ONE 6(7): e21017). In some aspects, the provided embodiments
modulate, such as by activate or increase the expression of FXN, however, the level of expression does not result in substantial overexpression of FXN to a level that would result in toxicity, organ dysregulation, cell death or reduction of life span. For example, the expression or activity of FXN is increased by less than about 10-fold, 9-fold, 8-fold, 7-fold or 6-fold, compared to the expression or activity of FXN in the subject or cell with the disease or disorder in the absence of the provided compositions or uses. In certain aspects, the expression or activity of FXN is increased to at least at or about 30%, 40%, 50%, 60%, 70%, 75%, 80%, 85%, 90%, 95%, 100%, 105%, 110%, 120%, 125%, 150%, 175%, 200%, 225%, 250%, 300%, 400%, or 500%, of the expression or activity of FXN in an individual or a cell without the disease or disorder or in a wild-type cell. In certain aspects, the expression or activity of FXN does not exceed about 500%, 600%, 700%, 800%, 900%, or 1000% of the expression or activity of FXN in an individual or a cell without the disease or disorder or in a wild-type cell. In some aspects, the provided embodiments offer the ability to fine tune and tightly regulate the expression level of FXN, for example, for treating diseases or disorders associated with reduced activity, inactivation, mutation and/or dysregulation of expression of the FXN gene, such as FA.
V. KITS AND ARTICLES OF MANUFACTURE
[0626] Also provided are articles of manufacture, systems, apparatuses, and kits useful in performing the provided embodiments. In some embodiments, the provided articles of manufacture or kits contain one or more components of the one or more components of the DNA-targeting system provided herein. In some embodiments, the articles of manufacture or kits include polypeptides, nucleic acids, vectors and/or polynucleotides useful in performing the provided methods.
[0627] In some embodiments, the articles of manufacture or kits include one or more containers, typically a plurality of containers, packaging material, and a label or package insert on or associated with the container or containers and/or packaging, generally including instructions for use, e.g., instructions for introducing or administering.
[0628] Also provided are articles of manufacture, systems, apparatuses, and kits useful in administering the provided compositions, e.g., pharmaceutical compositions, e.g., for use in therapy or treatment. In some embodiments, the articles of manufacture or kits provided herein contain vectors and/or plurality of vectors, such as any vectors and/or plurality of vectors described herein. In some aspects, the articles of manufacture or kits provided herein can be used for administration of the vectors and/or plurality of vectors, and can include instructions for use.
[0629] The articles of manufacture and/or kits containing cells or cell compositions for therapy, may include a container and a label or package insert on or associated with the container. Suitable containers include, for example, bottles, vials, syringes, IV solution bags, etc. The containers may be formed from a variety of materials such as glass or plastic. The container in some embodiments holds a composition which is by itself or combined with another composition effective for treating, preventing and/or diagnosing the condition. In some embodiments, the container has a sterile access port. Exemplary
containers include an intravenous solution bags, vials, including those with stoppers pierceable by a needle for injection, or bottles or vials for orally administered agents. The label or package insert may indicate that the composition is used for treating a disease or condition. The article of manufacture may further include a package insert indicating that the compositions can be used to treat a particular condition. Alternatively, or additionally, the article of manufacture may further include another or the same container comprising a pharmaceutically-acceptable buffer. It may further include other materials such as other buffers, diluents, filters, needles, and/or syringes.
VI. DEFINITIONS
[06303 Unless defined otherwise, all terms of art, notations and other technical and scientific terms or terminology used herein are intended to have the same meaning as is commonly understood by one of ordinary skill in the art to which the claimed subject matter pertains. In some cases, terms with commonly understood meanings are defined herein for clarity and/or for ready reference, and the inclusion of such definitions herein should not necessarily be construed to represent a substantial difference over what is generally understood in the art.
[06311 As used herein, the singular forms “a,” “an,” and “the” include plural referents unless the context clearly dictates otherwise. For example, “a” or “an” means “at least one” or “one or more.” It is understood that aspects and variations described herein include “consisting” and/or “consisting essentially of’ aspects and variations.
[0632] Throughout this disclosure, various aspects of the claimed subject matter are presented in a range format. It should be understood that the description in range format is merely for convenience and brevity and should not be construed as an inflexible limitation on the scope of the claimed subject matter. Accordingly, the description of a range should be considered to have specifically disclosed all the possible sub-ranges as well as individual numerical values within that range. For example, where a range of values is provided, it is understood that each intervening value, between the upper and lower limit of that range and any other stated or intervening value in that stated range is encompassed within the claimed subject matter. The upper and lower limits of these smaller ranges may independently be included in the smaller ranges, and are also encompassed within the claimed subject matter, subject to any specifically excluded limit in the stated range. Where the stated range includes one or both of the limits, ranges excluding either or both of those included limits are also included in the claimed subject matter. This applies regardless of the breadth of the range.
[0633] The term “about” as used herein refers to the usual error range for the respective value readily known. Reference to “about” a value or parameter herein includes (and describes) embodiments that are directed to that value or parameter per se. For example, description referring to “about X” includes description of “X”. In some embodiments, “about” may refer to ±25%, ±20%, ±15%, ±10%, ±5%, or ±1%.
[0634] In some aspects, corresponding positions of the one or more modifications, such as one or
more substitutions, can be determined in reference to positions of a reference amino acid sequence or a reference nucleotide sequence. As used herein, recitation that nucleotides or amino acid positions “correspond to” nucleotides or amino acid positions in a disclosed sequence, such as set forth in the Sequence listing, refers to nucleotides or amino acid positions identified upon alignment with the disclosed sequence to maximize identity using a standard alignment algorithm, such as the GAP algorithm or other available algorithms. By aligning the sequences, corresponding residues can be identified, for example, using conserved and identical amino acid residues as guides. In general, to identify corresponding positions, the sequences of amino acids are aligned so that the highest order match is obtained (see, e.g. : Computational Molecular Biology, Lesk, A.M., ed., Oxford University Press, New York, 1988; Biocomputing: Informatics and Genome Projects, Smith, D.W., ed., Academic Press, New York, 1993; Computer Analysis of Sequence Data, Part I, Griffin, A.M., and Griffin, H.G., eds., Humana Press, New Jersey, 1994; Sequence Analysis in Molecular Biology, von Heinje, G., Academic Press, 1987; and Sequence Analysis Primer, Gribskov, M. and Devereux, J., eds., M Stockton Press, New York, 1991; Carrillo et al. (1988) SIAM J Applied Math 48: 1073). Alignment for determining corresponding positions can be obtained in various ways, for instance, using publicly available computer software such as BLAST, BLAST-2, ALIGN or Megalign (DNASTAR) software. Appropriate parameters for aligning sequences can be determined, including any algorithms needed to achieve maximal alignment over the full length of the sequences being compared. For example, corresponding residues can be determined by alignment of a reference sequence that is a wild-type Cas protein by available alignment methods. By aligning the sequences, one skilled in the art can identify corresponding residues, for example, using conserved and/or identical amino acid residues as guides.
[0635] The term “vector,” as used herein, refers to a nucleic acid molecule capable of propagating another nucleic acid to which it is linked. The term includes the vector as a self-replicating nucleic acid structure as well as the vector incorporated into the genome of a host cell into which it has been introduced. Certain vectors are capable of directing the expression of nucleic acids to which they are operatively linked. Such vectors are referred to herein as “expression vectors.” Among the vectors are viral vectors, such as adenoviral vectors.
[0636| As used herein, “percent (%) amino acid sequence identity” and “percent identity” when used with respect to an amino acid sequence (reference polypeptide sequence) is defined as the percentage of amino acid residues in a candidate sequence that are identical with the amino acid residues in the reference polypeptide sequence, after aligning the sequences and introducing gaps, if necessary, to achieve the maximum percent sequence identity, and not considering any conservative substitutions as part of the sequence identity. Alignment for purposes of determining percent amino acid sequence identity can be achieved in various known ways, in some embodiments, using publicly available computer software such as BLAST, BLAST-2, ALIGN or Megalign (DNASTAR) software. Appropriate parameters for aligning sequences can be determined, including any algorithms needed to achieve maximal alignment over the full length of the sequences being compared.
[0637] In some embodiments, “operably linked” may include the association of components, such as a DNA sequence, e.g. a heterologous nucleic acid) and a regulatory sequence(s), in such a way as to permit gene expression when the appropriate molecules (e.g. transcriptional activator proteins) are bound to the regulatory sequence. Hence, it means that the components described are in a relationship permitting them to function in their intended manner.
[0638] An amino acid substitution may include replacement of one amino acid in a polypeptide with another amino acid. The substitution may be a conservative amino acid substitution or a non-conservative amino acid substitution. Amino acid substitutions may be introduced into a binding molecule, e.g., antibody, of interest and the products screened for a desired activity, e.g., retained/improved antigen binding, decreased immunogenicity, or improved ADCC or CDC.
[0639] Amino acids generally can be grouped according to the following common side-chain properties:
(1) hydrophobic: Norleucine, Met, Ala, Vai, Leu, He;
(2) neutral hydrophilic: Cys, Ser, Thr, Asn, Gin;
(3) acidic: Asp, Glu;
(4) basic: His, Lys, Arg;
(5) residues that influence chain orientation: Gly, Pro;
(6) aromatic: Trp, Tyr, Phe.
[0640] In some embodiments, conservative substitutions can involve the exchange of a member of one of these classes for another member of the same class. In some embodiments, non-conservative amino acid substitutions can involve exchanging a member of one of these classes for another class.
[0641] As used herein, a composition refers to any mixture of two or more products, substances, or compounds, including cells. It may be a solution, a suspension, liquid, powder, a paste, aqueous, nonaqueous or any combination thereof.
[0642] As used herein, a “subject” is a mammal, such as a human or other animal, and typically is human.
VII. EXEMPLARY EMBODIMENTS
[0643] Among the provided embodiments are:
1. An engineered zinc finger protein (eZFP) that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the target site is within a target region spanning the genomic coordinates chr9:69, 034, 900-69, 035, 900 from human genome assembly GRCh38 (hg38) (SEQ ID NO:430), or within a target region spanning the genomic coordinates chr9:69, 027, 282-69, 028, 497 from hg38 (SEQ ID NO:431).
2. The eZFP of embodiment 1, wherein the target site is within a target region spanning the genomic coordinates chr9:69, 034, 900-69, 035, 900 from hg38 (SEQ ID NO:430).
3. The eZFP of embodiment 1, wherein the target site is within a target region spanning the genomic coordinates chr9:69, 035, 300-69-035, 800 from hg38.
4. The eZFP of embodiment 1, wherein the target site is within a target region spanning the genomic coordinates chr9:69, 035, 350-69, 035, 450 from hg38.
5. The eZFP of embodiment 1, wherein the target site is within a target region spanning the genomic coordinates chr9:69, 035, 400-69, 035, 450 from hg38.
6. The eZFP of embodiment 1, wherein the target site is within a target region spanning the genomic coordinates chr9:69, 035, 530-69, 035, 580 from hg38.
7. The eZFP of embodiment 1, wherein the target site is within a target region spanning the genomic coordinates chr9:69, 035, 675-69, 035, 725 from hg38.
8. The eZFP of embodiment 1, wherein the target site is within a target region spanning the genomic coordinates chr9:69, 027, 282-69, 028, 497 from hg38 (SEQ ID NO:431).
9. The eZFP of embodiment 1, wherein the target site is within a target region spanning the genomic coordinates chr9:69, 027, 615-69, 028, 101 from hg38.
10. The eZFP of embodiment 1, wherein the target site is within a target region spanning the genomic coordinates chr9:69, 027, 775-69, 027, 875 from hg38.
11. The eZFP of embodiment 1, wherein the target site is within a target region spanning the genomic coordinates chr9:69, 027, 795-69, 027, 845 from hg38.
12. The eZFP of embodiment 1, wherein the target site comprises the nucleotide sequence set forth in any one of SEQ ID NOS:269-300 and 583-600, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
13. The eZFP of embodiment 1, wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:272, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
14. The eZFP of embodiment 13, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: QSGNLAR (SEQ ID NO:341); F2: QKVNRAG (SEQ ID NO:342); F3: DRSNLSR (SEQ ID NO:343); F4: QSGHLSR (SEQ ID NO:344); F5: TSGHLSR (SEQ ID NO:345); and F6: RSDALAR (SEQ ID NO:346).
15. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: QSGNLAR (SEQ ID NO:341); F2: QKVNRAG (SEQ ID NO:342); F3: DRSNLSR (SEQ ID NO:343); F4: QSGHLSR (SEQ ID NO:344); F5: TSGHLSR (SEQ ID NO:345); and F6: RSDALAR (SEQ ID NO:346).
16. The eZFP of any of embodiments 13-15, wherein the eZFP comprises the sequence set forth in SEQ ID NO:301, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
17. The eZFP of any of embodiments 13-16, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:301.
18. The eZFP of any of embodiments 13-17, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:308 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
19. The eZFP of any of embodiments 13-18, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:308.
20. The eZFP of embodiment 1, wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:277, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
21. The eZFP of embodiment 20, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RSDNLSE (SEQ ID NO:347); F2: KSWSRYK (SEQ ID NO:348); F3: TSGSLSR (SEQ ID NO:349); F4: RSDALAR (SEQ ID NO:350); F5: RSDNLSV (SEQ ID NO:351); and F6: FSSCRSA (SEQ ID NO:352).
22. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RSDNLSE (SEQ ID NO:347); F2: KSWSRYK (SEQ ID NO:348); F3: TSGSLSR (SEQ ID NO:349); F4: RSDALAR (SEQ ID NO:350); F5: RSDNLSV (SEQ ID NO:351); and F6: FSSCRSA (SEQ ID NO:352).
23. The eZFP of any of embodiments 20-22, wherein the eZFP comprises the sequence set forth in SEQ ID NO:302, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
24. The eZFP of any of embodiments 20-23, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:302.
25. The eZFP of any of embodiments 20-24, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:309 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
26. The eZFP of any of embodiments 20-25, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:309.
27. The eZFP of embodiment 1, wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:280, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
28. The eZFP of embodiment 27, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: TSGNLTR (SEQ ID NO:353); F2: EQTTRDK (SEQ ID NO:354); F3: RSANLAR (SEQ ID NO:355); F4: RLDNRTA (SEQ ID NO:356); F5: DSSHRTR (SEQ ID NO:357); and F6: RKYYLAK (SEQ ID NO:358).
29. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: TSGNLTR (SEQ ID NO:353); F2: EQTTRDK (SEQ ID NO:354); F3: RSANLAR (SEQ ID NO:355); F4: RLDNRTA (SEQ ID NO:356); F5: DSSHRTR (SEQ ID NO:357); and F6: RKYYLAK (SEQ ID NO:358).
30. The eZFP of any of embodiments 27-29, wherein the eZFP comprises the sequence set forth in SEQ ID NO:303, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
31. The eZFP of any of embodiments 27-30, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:303.
32. The eZFP of any of embodiments 27-31, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NOG 10 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
33. The eZFP of any of embodiments 27-32, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NOG 10.
34. The eZFP of embodiment 1, wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:281, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
35. The eZFP of embodiment 34, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RSAHLSR (SEQ ID NO:359); F2: DRSDLSR (SEQ ID NO:360); F3: RSDHLSV (SEQ ID NO:361); F4: RSDVRKT (SEQ ID NO:362); F5: QSGALAR (SEQ ID NO:363); and F6: RKYYLAK (SEQ ID NO:364).
36. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RSAHLSR (SEQ ID NO:359); F2: DRSDLSR (SEQ ID NO:360); F3: RSDHLSV (SEQ ID NO:361); F4: RSDVRKT (SEQ ID NO:362); F5: QSGALAR (SEQ ID NO:363); and F6: RKYYLAK (SEQ ID NO:364).
37. The eZFP of any of embodiments 34-36, wherein the eZFP comprises the sequence set forth in SEQ ID NO:304, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
38. The eZFP of any of embodiments 34-37, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:304.
39. The eZFP of any of embodiments 34-38, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 311 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
40. The eZFP of any of embodiments 34-39, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 311.
41. The eZFP of embodiment 1, wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:283, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
42. The eZFP of embodiment 41, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RSAHLSR (SEQ ID NO:365); F2: RSDALAR (SEQ ID NO:366); F3: ATSNRSA (SEQ ID NO:367); F4: RSAHLSR (SEQ ID NO:368); F5: TSGSLSR (SEQ ID NO:369); and F6: QSGDLTR (SEQ ID NO:370).
43. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RSAHLSR (SEQ ID NO:365); F2: RSDALAR (SEQ ID NO:366); F3: ATSNRSA (SEQ ID NO:367); F4: RSAHLSR (SEQ ID NO:368); F5: TSGSLSR (SEQ ID NO:369); and F6: QSGDLTR (SEQ ID NO:370).
44. The eZFP of any of embodiments 41-43, wherein the eZFP comprises the sequence set forth in SEQ ID NO:305, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
45. The eZFP of any of embodiments 41-44, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:305.
46. The eZFP of any of embodiments 41-45, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NOG 12 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
47. The eZFP of any of embodiments 41-46, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NOG 12.
48. The eZFP of embodiment 1, wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:290, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
49. The eZFP of embodiment 48, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: QSGDLTR (SEQ ID NO:371); F2: QSSDLRR (SEQ ID NO:372); F3: RSDNLSE (SEQ ID NO:373); F4: SSRNLAS (SEQ ID NO:374); F5: DRSHLTR (SEQ ID NO:375); and F6: RSDDLTR (SEQ ID NO:376).
50. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: QSGDLTR (SEQ ID NO:371); F2: QSSDLRR (SEQ ID NO:372); F3: RSDNLSE (SEQ ID NO:373); F4: SSRNLAS (SEQ ID NO:374); F5: DRSHLTR (SEQ ID NO:375); and F6: RSDDLTR (SEQ ID NO:376).
51. The eZFP of any of embodiments 48-50, wherein the eZFP comprises the sequence set forth in SEQ ID NO:306, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
52. The eZFP of any of embodiments 48-51, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:306.
53. The eZFP of any of embodiments 48-52, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NOG 13 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
54. The eZFP of any of embodiments 48-53, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NOG 13.
55. The eZFP of embodiment 1, wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:299, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
56. The eZFP of embodiment 55, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: LRHHLTR (SEQ ID NO:377); F2: QSAHLKA (SEQ ID NO:378); F3: LPQTLQR (SEQ ID NO:379); F4: QNATRTK (SEQ ID NO:380); F5: QSSHLTR (SEQ ID NO:381); and F6: RSDHLSR (SEQ ID NO:382).
57. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: LRHHLTR (SEQ ID NO:377); F2: QSAHLKA (SEQ ID NO:378); F3: LPQTLQR (SEQ ID NO:379); F4: QNATRTK (SEQ ID NO:380); F5: QSSHLTR (SEQ ID NO:381); and F6: RSDHLSR (SEQ ID NO:382).
58. The eZFP of any of embodiments 55-57, wherein the eZFP comprises the sequence set forth in SEQ ID NO:307, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
59. The eZFP of any of embodiments 55-58, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:307.
60. The eZFP of any of embodiments 55-59, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:314 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
61. The eZFP of any of embodiments 55-60, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:314.
62. The eZFP of embodiment 1, wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:583, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
63. The eZFP of embodiment 62, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RSDSLLR (SEQ ID NO:475); F2: TSSNRKT (SEQ ID NO:476); F3: RSAHLSR (SEQ ID NO:477); F4: TSGSLTR (SEQ ID NO:478); F5: QSGDLTR (SEQ ID NO:479); and F6: QWGTRYR (SEQ ID NO:480).
64. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RSDSLLR (SEQ ID NO:475); F2: TSSNRKT (SEQ ID NO:476); F3: RSAHLSR (SEQ ID NO:477); F4: TSGSLTR (SEQ ID NO:478); F5: QSGDLTR (SEQ ID NO:479); and F6: QWGTRYR (SEQ ID NO:480).
65. The eZFP of any of embodiments 62-64, wherein the eZFP comprises the sequence set forth in SEQ ID NO:439, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
66. The eZFP of any of embodiments 62-65, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:439.
67. The eZFP of any of embodiments 62-66, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:457 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
68. The eZFP of any of embodiments 62-67, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:457.
69. The eZFP of embodiment 1, wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:584, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
70. The eZFP of embodiment 69, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: QARHLTC (SEQ ID NO:481); F2: QSGHLSR (SEQ ID NO:482); F3: RSDVLSE (SEQ ID NO:483); F4: KHSTRRV (SEQ ID NO:484); F5: QSSDLSR (SEQ ID NO:485); and F6: WKWNLRA (SEQ ID NO:486).
71. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: QARHLTC (SEQ ID NO:481); F2: QSGHLSR (SEQ ID NO:482); F3: RSDVLSE (SEQ ID NO:483); F4: KHSTRRV (SEQ ID NO:484); F5: QSSDLSR (SEQ ID NO:485); and F6: WKWNLRA (SEQ ID NO:486).
72. The eZFP of any of embodiments 69-71, wherein the eZFP comprises the sequence set forth in SEQ ID NO:440, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
73. The eZFP of any of embodiments 69-72, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:440.
74. The eZFP of any of embodiments 69-73, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 458 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
75. The eZFP of any of embodiments 69-74, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 458.
76. The eZFP of embodiment 1, wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:585, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
77. The eZFP of embodiment 77, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RSDNLAR (SEQ ID NO:487); F2: WRGDRVK (SEQ ID NO:488); F3: YKHVLSD (SEQ ID NO:489); F4: TSGSLTR (SEQ ID NO:490); F5: QSGNLAR (SEQ ID NO:491); and F6: RARDLSK (SEQ ID NO:492).
78. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RSDNLAR (SEQ ID NO:487); F2: WRGDRVK (SEQ ID NO:488); F3: YKHVLSD (SEQ ID NO:489); F4: TSGSLTR (SEQ ID NO:490); F5: QSGNLAR (SEQ ID NO:491); and F6: RARDLSK (SEQ ID NO:492).
79. The eZFP of any of embodiments 76-78, wherein the eZFP comprises the sequence set forth in SEQ ID NO:441, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
80. The eZFP of any of embodiments 76-79, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:441.
81. The eZFP of any of embodiments 76-80, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:459 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
82. The eZFP of any of embodiments 76-81, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:459.
83. The eZFP of embodiment 1, wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:586, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
84. The eZFP of embodiment 83, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: QSANRTK (SEQ ID NO:493); F2: QSGNLAR (SEQ ID NO:494); F3: RSDNLSV (SEQ ID NO:495); F4: IRSTLRD (SEQ ID NO:496); F5: QNAHRKT (SEQ ID NO:497); and F6: HRSSLRR (SEQ ID NO:498).
85. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: QSANRTK (SEQ ID NO:493); F2: QSGNLAR (SEQ ID NO:494); F3: RSDNLSV (SEQ ID NO:495); F4: IRSTLRD (SEQ ID NO:496); F5: QNAHRKT (SEQ ID NO:497); and F6: HRSSLRR (SEQ ID NO:498).
86. The eZFP of any of embodiments 83-85, wherein the eZFP comprises the sequence set forth in SEQ ID NO:442, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
87. The eZFP of any of embodiments 83-86, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:442.
88. The eZFP of any of embodiments 83-87, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 460 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
89. The eZFP of any of embodiments 83-88, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:460.
90. The eZFP of embodiment 1, wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:587, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
91. The eZFP of embodiment 90, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: QAGNRST (SEQ ID NO:499); F2: DRSALAR (SEQ ID NO:500); F3: RSDNLAR (SEQ ID NO:501); F4: WRGDRVK (SEQ ID NO:502); F5: YKHVLSD (SEQ ID NO:503); and F6: TSGSLTR (SEQ ID NO:504).
92. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: QAGNRST (SEQ ID NO:499); F2: DRSALAR (SEQ ID NG:500); F3: RSDNLAR (SEQ ID NO:501); F4: WRGDRVK (SEQ ID NO:502); F5: YKHVLSD (SEQ ID NO:503); and F6: TSGSLTR (SEQ ID NO:504).
93. The eZFP of any of embodiments 90-92, wherein the eZFP comprises the sequence set forth in SEQ ID NO:443, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
94. The eZFP of any of embodiments 90-93, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:443.
95. The eZFP of any of embodiments 90-94, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 461 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
96. The eZFP of any of embodiments 90-95, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 461.
97. The eZFP of embodiment 1, wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:588, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
98. The eZFP of embodiment 97, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RSDNLSV (SEQ ID NO:505); F2: IRSTLRD (SEQ ID NO:506); F3: QNAHRKT (SEQ ID NO:507); F4: HRSSLRR (SEQ ID NO:508); F5: RSDNLAR (SEQ ID NO:509); and F6: QRSPLPA (SEQ ID NQ:510).
99. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RSDNLSV (SEQ ID NO:505); F2: IRSTLRD (SEQ ID NO:506); F3: QNAHRKT (SEQ ID NO:507); F4: HRSSLRR (SEQ ID NO:508); F5: RSDNLAR (SEQ ID NO:509); and F6: QRSPLPA (SEQ ID NQ:510).
100. The eZFP of any of embodiments 97-99, wherein the eZFP comprises the sequence set forth in SEQ ID NO:444, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
101. The eZFP of any of embodiments 97-100, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:444.
102. The eZFP of any of embodiments 97-101, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 462 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
103. The eZFP of any of embodiments 97-102, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:462.
104. The eZFP of embodiment 1, wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:589, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
105. The eZFP of embodiment 104, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: DRSTRTK (SEQ ID NO:511); F2: RSDYLAK (SEQ ID NO:512); F3: LRHHLTR (SEQ ID NO:513); F4: QSAHLKA (SEQ ID NO:514); F5: LPQTLQR (SEQ ID NO:515); and F6: QNATRTK (SEQ ID NO:516).
106. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: DRSTRTK (SEQ ID NO:511); F2: RSDYLAK (SEQ ID NO:512); F3: LRHHLTR (SEQ ID NO:513); F4: QSAHLKA (SEQ ID NO:514); F5: LPQTLQR (SEQ ID NO:515); and F6: QNATRTK (SEQ ID NO:516).
107. The eZFP of any of embodiments 104-106, wherein the eZFP comprises the sequence set forth in SEQ ID NO:445, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
108. The eZFP of any of embodiments 104-107, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:445.
109. The eZFP of any of embodiments 104-108, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 463 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
110. The eZFP of any of embodiments 104-109, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 463.
111. The eZFP of embodiment 1, wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:590, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
112. The eZFP of embodiment 111, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RSADLTR (SEQ ID NO:517); F2: RSDDLTR (SEQ ID NO:518); F3: QSSDLSR (SEQ ID NO:519); F4: WHSSLHQ (SEQ ID NO:520); F5: RSDSLSQ (SEQ ID NO:521); and F6: RKADRTR (SEQ ID NO:522).
113. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RSADLTR (SEQ ID NO:517); F2: RSDDLTR (SEQ ID NO:518); F3: QSSDLSR (SEQ ID NO:519); F4: WHSSLHQ (SEQ ID NO:520); F5: RSDSLSQ (SEQ ID NO:521); and F6: RKADRTR (SEQ ID NO:522).
114. The eZFP of any of embodiments 111-113, wherein the eZFP comprises the sequence set forth in SEQ ID NO:446, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
115. The eZFP of any of embodiments 111-114, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:446.
116. The eZFP of any of embodiments 111-115, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 464 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
117. The eZFP of any of embodiments 111-116, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:464.
118. The eZFP of embodiment 1, wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:591, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
119. The eZFP of embodiment 118, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RNDALTE (SEQ ID NO:523); F2: RKDNLKN (SEQ ID NO:524); F3: TSGELVR (SEQ ID NO:525); F4: HRTTLTN (SEQ ID NO:526); F5: TTGNLTV (SEQ ID NO:527); and F6: RTDTLRD (SEQ ID NO:528).
120. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RNDALTE (SEQ ID NO:523); F2: RKDNLKN (SEQ ID NO:524); F3: TSGELVR (SEQ ID NO:525); F4: HRTTLTN (SEQ ID NO:526); F5: TTGNLTV (SEQ ID NO:527); and F6: RTDTLRD (SEQ ID NO:528).
121. The eZFP of any of embodiments 118-120, wherein the eZFP comprises the sequence set forth in SEQ ID NO:447, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
122. The eZFP of any of embodiments 118-121, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:447.
123. The eZFP of any of embodiments 118-122, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 465 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
124. The eZFP of any of embodiments 118-123, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 465.
125. The eZFP of embodiment 1, wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:592, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
126. The eZFP of embodiment 125, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RKDNLKN (SEQ ID NO:529); F2: RADNLTE (SEQ ID NO:530); F3: TSHSLTE (SEQ ID NO:531); F4: SKKHLAE (SEQ ID NO:532); F5: TSGELVR (SEQ ID NO:533); and F6: TSGELVR (SEQ ID NO:534).
127. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RKDNLKN (SEQ ID NO:529); F2: RADNLTE (SEQ ID NO:530); F3: TSHSLTE (SEQ ID NO:531); F4: SKKHLAE (SEQ ID NO:532); F5: TSGELVR (SEQ ID NO:533); and F6: TSGELVR (SEQ ID NO:534).
128. The eZFP of any of embodiments 125-127, wherein the eZFP comprises the sequence set forth in SEQ ID NO:448, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
129. The eZFP of any of embodiments 125-128, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:448.
130. The eZFP of any of embodiments 125-129, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 466 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
131. The eZFP of any of embodiments 125-130, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:466.
132. The eZFP of embodiment 1, wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:593, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
133. The eZFP of embodiment 132, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: THLDLIR (SEQ ID NO:535); F2: DCRDLAR (SEQ ID NO:536); F3: RSDELVR (SEQ ID NO:537); F4: RNDALTE (SEQ ID NO:538); F5: SKKHLAE (SEQ ID NO:539); and F6: QSGHLTE (SEQ ID NO:540).
134. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: THLDLIR (SEQ ID NO:535); F2: DCRDLAR (SEQ ID NO:536); F3: RSDELVR (SEQ ID NO:537); F4: RNDALTE (SEQ ID NO:538); F5: SKKHLAE (SEQ ID NO:539); and F6: QSGHLTE (SEQ ID NO:540).
135. The eZFP of any of embodiments 132-134, wherein the eZFP comprises the sequence set forth in SEQ ID NO:449, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
136. The eZFP of any of embodiments 132-135, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:449.
137. The eZFP of any of embodiments 132-136, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 467 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
138. The eZFP of any of embodiments 132-137, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 467.
139. The eZFP of embodiment 1, wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:594, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
140. The eZFP of embodiment 139, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: HTGHLLE (SEQ ID NO:541); F2: DPGHLVR (SEQ ID NO:542); F3: THLDLIR (SEQ ID NO:543); F4: DCRDLAR (SEQ ID NO:544); F5: RSDELVR (SEQ ID NO:545); and F6: RNDALTE (SEQ ID NO:546).
141. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: HTGHLLE (SEQ ID NO:541); F2: DPGHLVR (SEQ ID NO:542); F3: THLDLIR (SEQ ID NO:543); F4: DCRDLAR (SEQ ID NO:544); F5: RSDELVR (SEQ ID NO:545); and F6: RNDALTE (SEQ ID NO:546).
142. The eZFP of any of embodiments 139-141, wherein the eZFP comprises the sequence set forth in SEQ ID NO:450, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
143. The eZFP of any of embodiments 139-142, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:450.
144. The eZFP of any of embodiments 139-143, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:468 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
145. The eZFP of any of embodiments 139-144, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:468.
146. The eZFP of embodiment 1, wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:595, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
147. The eZFP of embodiment 146, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RSDKLVR (SEQ ID NO:547); F2: RSDHLTT (SEQ ID NO:548); F3: RNDALTE (SEQ ID NO:549); F4: TTGALTE (SEQ ID NO:550); F5: THLDLIR (SEQ ID NO:551); and F6: DPGHLVR (SEQ ID NO:552).
148. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RSDKLVR (SEQ ID NO:547); F2: RSDHLTT (SEQ ID NO:548); F3: RNDALTE (SEQ ID NO:549); F4: TTGALTE (SEQ ID NO:550); F5: THLDLIR (SEQ ID NO:551); and F6: DPGHLVR (SEQ ID NO:552).
149. The eZFP of any of embodiments 146-148, wherein the eZFP comprises the sequence set forth in SEQ ID NO:451, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
150. The eZFP of any of embodiments 146-149, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:451.
151. The eZFP of any of embodiments 146-150, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 469 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
152. The eZFP of any of embodiments 146-151, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 469.
153. The eZFP of embodiment 1, wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:596, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
154. The eZFP of embodiment 153, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: TKNSLTE (SEQ ID NO:553); F2: QLAHLRA (SEQ ID NO:554); F3: TSGSLVR (SEQ ID NO:555); F4: RSDNLVR (SEQ ID NO:556); F5: QNSTLTE (SEQ ID NO:557); and F6: RADNLTE (SEQ ID NO:558).
155. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: TKNSLTE (SEQ ID NO:553); F2: QLAHLRA (SEQ ID NO:554); F3: TSGSLVR (SEQ ID NO:555); F4: RSDNLVR (SEQ ID NO:556); F5: QNSTLTE (SEQ ID NO:557); and F6: RADNLTE (SEQ ID NO:558).
156. The eZFP of any of embodiments 153-155, wherein the eZFP comprises the sequence set forth in SEQ ID NO:452, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
157. The eZFP of any of embodiments 153-156, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:452.
158. The eZFP of any of embodiments 153-157, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 470 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
159. The eZFP of any of embodiments 153-158, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:470.
160. The eZFP of embodiment 1, wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:597, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
161. The eZFP of embodiment 160, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RADNLTE (SEQ ID NO:559); F2: TKNSLTE (SEQ ID NO:560); F3: QLAHLRA (SEQ ID NO:561); F4: TSGSLVR (SEQ ID NO:562); F5: RSDNLVR (SEQ ID NO:563); and F6: QNSTLTE (SEQ ID NO:564).
162. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RADNLTE (SEQ ID NO:559); F2: TKNSLTE (SEQ ID NO:560); F3: QLAHLRA (SEQ ID NO:561); F4: TSGSLVR (SEQ ID NO:562); F5: RSDNLVR (SEQ ID NO:563); and F6: QNSTLTE (SEQ ID NO:564).
163. The eZFP of any of embodiments 160-162, wherein the eZFP comprises the sequence set forth in SEQ ID NO:453, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
164. The eZFP of any of embodiments 160-163, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:453.
165. The eZFP of any of embodiments 160-164, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 471 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
166. The eZFP of any of embodiments 160-165, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 471.
167. The eZFP of embodiment 1, wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:598, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
168. The eZFP of embodiment 167, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: TSGHLVR (SEQ ID NO:565); F2: QLAHLRA (SEQ ID NO:566); F3: TSGELVR (SEQ ID NO:567); F4: QSGDLRR (SEQ ID NO:568); F5: QRAHLER (SEQ ID NO:569); and F6: RSDKLVR (SEQ ID NO:570).
169. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: TSGHLVR (SEQ ID NO:565); F2: QLAHLRA (SEQ ID NO:566); F3: TSGELVR (SEQ ID NO:567); F4: QSGDLRR (SEQ ID NO:568); F5: QRAHLER (SEQ ID NO:569); and F6: RSDKLVR (SEQ ID NO:570).
170. The eZFP of any of embodiments 167-169, wherein the eZFP comprises the sequence set forth in SEQ ID NO:454, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
171. The eZFP of any of embodiments 167-170, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:454.
172. The eZFP of any of embodiments 167-171, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 472 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
173. The eZFP of any of embodiments 167-172, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:472.
174. The eZFP of embodiment 1, wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:599, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
175. The eZFP of embodiment 174, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: REDNLHT (SEQ ID NO:571); F2: TSGHLVR (SEQ ID NO:572); F3: QLAHLRA (SEQ ID NO:573); F4: TSGELVR (SEQ ID NO:574); F5: QSGDLRR (SEQ ID NO:575); and F6: QRAHLER (SEQ ID NO:576).
176. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: REDNLHT (SEQ ID NO:571); F2: TSGHLVR (SEQ ID NO:572); F3: QLAHLRA (SEQ ID NO:573); F4: TSGELVR (SEQ ID NO:574); F5: QSGDLRR (SEQ ID NO:575); and F6: QRAHLER (SEQ ID NO:576).
177. The eZFP of any of embodiments 174-176, wherein the eZFP comprises the sequence set forth in SEQ ID NO:455, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
178. The eZFP of any of embodiments 174-177, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:455.
179. The eZFP of any of embodiments 174-178, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 473 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
180. The eZFP of any of embodiments 174-179, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 473.
181. The eZFP of embodiment 1, wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:600, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
182. The eZFP of embodiment 181, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: QRSDLTR (SEQ ID NO:577); F2: QGGTLRR (SEQ ID NO:578); F3: TSAHLAR (SEQ ID NO:579); F4: RREHLVR (SEQ ID NO:580); F5: QRHGLSS (SEQ ID NO:581); and F6: QRNALRG (SEQ ID NO:582).
183. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: QRSDLTR (SEQ ID NO:577); F2: QGGTLRR (SEQ ID NO:578); F3: TSAHLAR (SEQ ID NO:579); F4: RREHLVR (SEQ ID NO:580); F5: QRHGLSS (SEQ ID NO:581); and F6: QRNALRG (SEQ ID NO:582).
184. The eZFP of any of embodiments 181-183, wherein the eZFP comprises the sequence set forth in SEQ ID NO:456, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
185. The eZFP of any of embodiments 181-184, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:456.
186. The eZFP of any of embodiments 181-185, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 474 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
187. The eZFP of any of embodiments 181-186, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:474.
188. A fusion protein comprising the engineered zinc finger protein (eZFP) of any of embodiments 1-187.
189. A fusion protein comprising:
(a) the engineered zinc finger protein of any of embodiments 1-187 that binds to a target site in a regulatory DNA element of a FXN locus; and
(b) at least one epigenetic effector domain that increases transcription of the FXN locus.
190. The fusion protein of embodiment 189, wherein the at least one epigenetic effector domain comprises: a VP64 domain, a p65 activation domain, a p300 domain, an Rta domain, a CBP domain, a VPR domain, a VPH domain, an HSF1 domain, a TET protein domain, optionally wherein the TET protein is TET1, a SunTag domain, a domain from DPOLA, ENL, FOXO3, HSH2D, NCOA2, NCOA3, PSA1, PYGO1, RBM39, HERC2, or NOTCH2, or a domain, portion, variant, or truncation of any of the foregoing.
191. The fusion protein of embodiment 189 or 190, wherein the at least one epigenetic effector domain comprises the sequence set forth in any of SEQ ID NOS:81, 83, 100-109, 111-122, 124, 125, 134-140, 152, and 383-396, or a domain, portion, variant, or truncation thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
192. The fusion protein of any of embodiments 189-191, wherein the at least one effector domain comprises at least one VP16 domain, or a VP16 tetramer (“VP64”) or a variant thereof.
193. The fusion protein of any of embodiments 189-192, wherein the at least one effector domain comprises VP64.
194. The fusion protein of any of embodiments 189-193, wherein the at least one effector domain comprises a VP64 domain comprising the sequence set forth in SEQ ID NO: 83, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
195. The fusion protein of any of embodiments 189-194, wherein the at least one effector domain comprises a VP64 domain comprising the sequence set forth in SEQ ID NO: 83.
196. The fusion protein of any of embodiments 189-195, wherein the at least one epigenetic effector domain comprises: a domain from DPOLA, ENL, F0X03, HSH2D, NC0A2, NC0A3, PSA1, PYG01, RBM39, HERC2, or N0TCH2, or a domain, portion, variant, or truncation of any of the foregoing.
197. The fusion protein of any of embodiments 189-196, wherein the at least one epigenetic effector domain comprises the sequence set forth in any of SEQ ID NOS:383-393, or a domain, portion, or variant thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
198. The fusion protein of any of embodiments 189-197, wherein the at least one effector domain comprises a domain from NCOA2, NCOA3, FOXO3, PYGO1, or a portion or variant of any of the foregoing.
199. The fusion protein of any of embodiments 189-198, wherein each effector domain of the at least one effector domain is independently selected from an NCOA2 domain, an NCOA3 domain, a FOXO3 domain, and a PYGO1 domain.
200. The fusion protein of any of embodiments 189-199, wherein the at least one effector domain comprises a domain from NCOA2 comprising the sequence set forth in SEQ ID NO: 104 or SEQ ID NO:387, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
201. The fusion protein of any of embodiments 189-200, wherein the at least one effector domain comprises a domain from NCOA2 set forth in or SEQ ID NO:387.
202. The fusion protein of any of embodiments 189-201, wherein the at least one effector domain comprises a domain from NCOA3 comprising the sequence set forth in SEQ ID NO: 105 or SEQ ID NO:388, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
203. The fusion protein of any of embodiments 189-202, wherein the at least one effector domain comprises a domain from NCOA3 set forth in or SEQ ID NO:388.
204. The fusion protein of any of embodiments 189-203, wherein the at least one effector domain comprises a domain from FOXO3 comprising the sequence set forth in SEQ ID NO: 102 or SEQ ID NO:385, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
205. The fusion protein of any of embodiments 189-204, wherein the at least one effector domain comprises a domain from FOXO3 set forth in or SEQ ID NO:385.
206. The fusion protein of any of embodiments 189-205, wherein the at least one effector domain comprises a domain from PYGO1 comprising the sequence set forth in SEQ ID NO: 107 or SEQ ID NO:390, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
207. The fusion protein of any of embodiments 189-206, wherein the at least one effector domain comprises a domain from PYGO1 set forth in or SEQ ID NO:390.
208. The fusion protein of any of embodiments 189-207, wherein the at least one effector domain is a multipartite effector composed of at least two effector domains.
209. The fusion protein of embodiment 208, wherein the multipartite effector is composed of two effector domains.
210. The fusion protein of embodiment 208, wherein the multipartite effector is composed of three effector domains.
211. The fusion protein of any of embodiments 208-210, wherein the multipartite effector is set forth in any one of SEQ ID NOS:397-418, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
212. The fusion protein of any of embodiments 208-211, wherein the multipartite effector is set forth in any one of SEQ ID NOS:411-418, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
213. The fusion protein of any of embodiments 208-212, wherein the multipartite effector comprises, in the N-terminal to C-terminal direction, domains from FOXO3, FOXO3, and NCOA3, respectively.
214. The fusion protein of embodiment 213, wherein the multipartite effector comprises the sequence set forth in SEQ ID NO:415, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
215. The fusion protein of any of embodiments 208-212, wherein the multipartite effector comprises, in the N-terminal to C-terminal direction, domains from NCOA3, FOXO3, and FOXO3, respectively.
216. The fusion protein of embodiment 215, wherein the multipartite effector comprises the sequence set forth in SEQ ID NO:418, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
217. The fusion protein of any of embodiments 208-212, wherein the multipartite effector comprises, in the N-terminal to C-terminal direction, domains from NCOA3, FOXO3, and NCOA3, respectively.
218. The fusion protein of embodiment 217, wherein the multipartite effector comprises the sequence set forth in SEQ ID NO:413, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
219. The fusion protein of any of embodiments 208-212, wherein the multipartite effector comprises, in the N-terminal to C-terminal direction, domains from NCOA2, FOXO3, and NCOA3, respectively.
220. The fusion protein of embodiment 219, wherein the multipartite effector comprises the sequence set forth in SEQ ID NO:416, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
221. The fusion protein of any of embodiments 208-212, wherein the multipartite effector comprises, in the N-terminal to C-terminal direction, domains from PYGO1, FOXO3, and NCOA3, respectively.
222. The fusion protein of embodiment 221, wherein the multipartite effector comprises the sequence set forth in SEQ ID NO:411, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
223. The fusion protein of any of embodiments 189-222, wherein the at least one epigenetic effector domain is fused to the N-terminus of the eZFP.
224. The fusion protein of any of embodiments 189-222, wherein the at least one epigenetic effector domain is fused to the C-terminus of the eZFP.
225. The fusion protein of any of embodiments 189-224, wherein the at least one epigenetic effector domain is fused to both the N-terminus and the C-terminus of the eZFP.
226. The fusion protein of any of embodiments 189-225, further comprising one or more nuclear localization signals (NFS).
227. The fusion protein of any of embodiments 189-226, further comprising one or more linkers.
228. The fusion protein of embodiment 227, wherein the one or more linkers are in between any two of the components of the fusion protein, including the eZFP, any of the at least one effector domains, and the one or more NFS.
229. The fusion protein of embodiment 227 or 228, wherein the one or more linkers connect the eZFP and the at least one epigenetic effector domain.
230. The fusion protein of any of embodiments 226-229, comprising the one or more NFS, the eZFP, and the at least one epigenetic effector domain, in order from N-terminus to C-terminus.
231. The fusion protein of embodiment 230, wherein the one or more NFS comprises a SV40 NFS sequence set forth in SEQ ID NO: 159 or a c-myc NFS sequence set forth in SEQ ID NO: 160.
232. The fusion protein of any of embodiments 189-231 , wherein the fusion protein comprises the sequence set forth in any of SEQ ID NOS:320-340, 419-425, and 608-671, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
233. The fusion protein of any of embodiments 189-232, wherein the fusion protein comprises the sequence set forth in any of SEQ ID NOS:320-340, 419-425, and 608-671.
234. The fusion protein of any of embodiments 189-233, wherein the fusion protein comprises the sequence set forth in any of SEQ ID NOS:320-340, 419-425, and 636-653.
235. The fusion protein of any of embodiments 189-234, wherein the fusion protein comprises the sequence set forth in any of SEQ ID NOS: 320-340 and 419-425.
236. The fusion protein of any of embodiments 189-234, wherein the fusion protein comprises the sequence set forth in any of SEQ ID NOS:636-653.
237. The fusion protein of any of embodiments 189-233, wherein the fusion protein comprises the sequence set forth in any of SEQ ID NOS:608-635 and 654-671.
238. The fusion protein of any of embodiments 189-233 and 237, wherein the fusion protein comprises the sequence set forth in any of SEQ ID NOS:608-635.
239. The fusion protein of any of embodiments 129-233 and 237, wherein the fusion protein comprises the sequence set forth in any of SEQ ID NOS: 654-671.
240. The fusion protein of any of embodiments 189-235, wherein the fusion protein comprises the sequence set forth in SEQ ID NO:326, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
241. The fusion protein of any of embodiments 189-235, wherein the fusion protein comprises the sequence set forth in SEQ ID NO:333, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
242. The fusion protein of any of embodiments 189-235, wherein the fusion protein comprises the sequence set forth in SEQ ID NO:340, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
243. The fusion protein of any of embodiments 189-235, wherein the fusion protein comprises the sequence set forth in SEQ ID NO:425, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
244. The fusion protein of any of embodiments 189-233, 237, and 239, wherein the fusion protein comprises the sequence set forth in SEQ ID NO:662, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
245. The fusion protein of any of embodiments 189-233, 237, and 239, wherein the fusion protein comprises the sequence set forth in SEQ ID NO:660, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
246. The fusion protein of any of embodiments 189-233, 237, and 239, wherein the fusion protein comprises the sequence set forth in SEQ ID NO: 658, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
247. A polynucleotide encoding the eZFP of any of embodiments 1-187, the fusion protein of any of embodiments 188-246, or a portion or component of any of the foregoing.
248. A plurality of polynucleotides encoding the eZFP of any of embodiments 1-187, the fusion protein of any of embodiments 188-246, or a portion or component of any of the foregoing.
249. A vector comprising the eZFP of any of embodiments 1-187, the fusion protein of any of claims 188-246, the polynucleotide of claim 247, the plurality of polynucleotides of claim 248, or a portion or component of any of the foregoing.
250. The vector of embodiment 249, wherein the vector is a viral vector.
251. The vector of embodiment 249 or 250, wherein the vector is an adeno-associated virus (AAV) vector.
252. The vector of embodiment 251 , wherein the AAV vector is selected from the group consisting of: AAV1, AAV2, AAV3, AAV4, AAV5, AAV6, AAV7, AAV8, AAV9, AAV10, AAV11, AAV 12, AAV-DJ, and AAVrh74.
253. The vector of embodiment 251 or 252, wherein the AAV vector is AAV6.
254. The vector of embodiment 251 or 252, wherein the AAV vector is AAV9.
255. The vector of embodiment 251 or 252, wherein the AAV vector is AAV-DJ.
256. The vector of embodiment 251 or 252, wherein the AAV vector is AAVrh74.
257. The vector of embodiment 249 or 250, wherein the vector is a lentiviral vector.
258. The vector of embodiment 249, wherein the vector is a non- viral vector.
259. The vector of embodiment 258, wherein the non-viral vector is selected from the group consisting of: a lipid nanoparticle, a liposome, an exosome, and a cell penetrating peptide.
260. The vector of any of embodiments 249-259, wherein the vector exhibits tropism for a nervous system cell, optionally a neuron, a heart cell, optionally a cardiomyocyte, a skeletal muscle cell, a fibroblast, an induced pluripotent stem cell, and/or a cell derived from any of the foregoing, or for a combination of any of the foregoing cells.
261. The vector of any of embodiments 249-260, wherein the vector exhibits tropism for induced pluripotent stem cells.
262. The vector of any of embodiments 249-261, wherein the vector exhibits tropism for neurons and cardiomyocytes.
263. The vector of any of embodiments 249-262, wherein the vector comprises one vector, or two or more vectors.
264. An AAV vector comprising one or both of: a) a first nucleic acid comprising an elongation factor alpha short (EFS) promoter operably linked to a sequence encoding a fusion protein comprising (i) a deactivated Cas (dCas) protein and (ii) at least one effector domain that increases transcription of a frataxin (FXN) locus; and b) a second nucleic acid comprising a U6 promoter operably linked to a sequence encoding a guide RNA (gRNA) comprising a gRNA spacer sequence that is capable of hybridizing to a target site in a regulatory DNA element of a FXN locus and/or is complementary to the target site.
265. The AAV vector of embodiment 264, wherein the AAV vector comprises both the first nucleic acid and the second nucleic acid.
266. The AAV vector of embodiment 264 or 265, wherein the first and second nucleic acid are comprised in a single polynucleotide.
267. The AAV vector of any of embodiments 264-266, wherein the EFS promoter comprises the sequence set forth in SEQ ID NO:436, or a sequence having at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to thereto.
268. The AAV vector of any of embodiments 264-267, wherein the EFS promoter comprises the sequence set forth in SEQ ID NO:436.
269. The AAV vector of any of embodiments 264-268, wherein the U6 promoter is a mini-U6 promoter.
270. The AAV vector of any of embodiments 264-269, wherein the mini-U6 promoter comprises the sequence set forth in SEQ ID NO:433, or a sequence having at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to thereto.
271. The AAV vector of any of embodiments 264-270, wherein the mini-U6 promoter comprises the sequence set forth in SEQ ID NO:433.
272. An AAV vector comprising a nucleic acid comprising a promoter selected from an elongation factor alpha short (EFS), CAG, or human elongation factor-1 alpha (EFla) promoter operably linked to a sequence encoding a fusion protein comprising (i) an eZFP that is capable of hybridizing to a target site in a regulatory DNA element of a frataxin (FXN) locus and/or is complementary to the target site and (ii) at least one effector domain that increases transcription of the frataxin (FXN) locus.
273. The AAV vector of any of embodiments 264-272, wherein the EFS promoter comprises the sequence set forth in SEQ ID NO:436, or a sequence having at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to thereto.
274. The AAV vector of any of embodiments 264-273, wherein the EFS promoter comprises the sequence set forth in SEQ ID NO:436.
275. The AAV vector of embodiment 272, wherein the CAG promoter comprises the sequence set forth in SEQ ID NO:602, or a sequence having at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to thereto.
276. The AAV vector of embodiment 272 or 275, wherein the CAG promoter comprises the sequence set forth in SEQ ID NO:602.
277. The AAV vector of embodiment 272, wherein the EFla promoter comprises the sequence set forth in SEQ ID NO:603, or a sequence having at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to thereto.
278. The AAV vector of embodiment 272 or 277, wherein the EFla promoter comprises the sequence set forth in SEQ ID NO: 603.
279. The AAV vector of any of embodiments 272-278, wherein the nucleic acid further comprises a 5’ untranslated region (UTR) set forth in SEQ ID NO: 605.
280. The AAV vector of any of embodiments 264-279, wherein the AAV vector further comprises inverted terminal repeats (ITRs).
281. The AAV vector of embodiment 280, wherein the ITRs are a first and second ITR, comprising the sequences set forth in SEQ ID NO:434 and SEQ ID NO:435, respectively.
282. The AAV vector of any of embodiments 266-271 , 280, and 281 , wherein the single polynucleotide comprises, in the 5’ to 3’ direction, the EFS promoter, the sequence encoding the fusion protein, the U6 promoter, and the sequence encoding the gRNA.
283. The AAV vector of any of embodiments 266-271 and 280-282, wherein the single polynucleotide further comprises a first IRT 5’ of the EFS promoter and a second ITR 3’ of the sequence encoding the gRNA.
284. The AAV vector of any of embodiments 264-283, wherein the first nucleic acid or the nucleic acid further comprises a polyA sequence selected from a SpA site or a bGH site downstream of the sequence encoding the fusion protein.
285. The AAV vector of any of embodiments 264-284, wherein the first nucleic acid or the nucleic acid comprising a sequence encoding a fusion protein further comprises a polyA sequence selected from a SpA site downstream of the sequence encoding the fusion protein.
286. The AAV vector of any of embodiments 264-271, 273, 274, and 282-284, wherein the first nucleic acid comprising a sequence encoding a fusion protein further comprises a polyA sequence selected from a bGH site downstream of the sequence encoding the fusion protein.
287. The AAV vector of any of embodiments 272-281, 284, and 285, wherein the nucleic acid comprising a sequence encoding a fusion protein further comprises a polyA sequence selected from a SpA site or bGH site downstream of the sequence encoding the fusion protein.
288. The AAV vector of any of embodiments 284, 285, and 287, wherein the SpA site comprises the sequence set forth in SEQ ID NO:437.
289. The AAV vector of any of embodiments 284, 286, and 287, wherein the bGH site comprises the sequence set forth in SEQ ID NO:604.
290. The AAV vector of any of embodiments 264-271, 273, 274, 280-285, and 288, wherein the first nucleic acid further comprises a woodchuck hepatitis virus post-transcriptional regulatory element (WPRE) in proximal to the SpA site, optionally wherein the WPRE is located between the sequence encoding the fusion protein and the SpA site.
291. The AAV vector of any of embodiments 264-271, 273, 274, 280-286, and 288-290, wherein the gRNA is capable of complexing with the dCas protein.
292. The AAV vector of any of embodiments 264-271, 273, 274, 280-286, and 288-291, wherein the gRNA comprises a gRNA spacer sequence that is capable of hybridizing to the target site or is complementary to the target site.
293. The AAV vector of any of embodiments 264-271, 273, 274, 280-286, and 288-292, wherein the dCas protein is a Staphylococcus aureus dCas9 (dSaCas9) protein or a Streptococcus pyogenes dCas9 (dSpCas9) protein.
294. The AAV vector of any of embodiments 264-271, 273, 274, 280-286, and 288-293, wherein the dCas protein is a Staphylococcus aureus dCas9 protein (dSaCas9) that comprises at least one amino acid mutation selected from D10A and N580A, with reference to numbering of positions of SEQ ID NO:73, and/or the dCas protein comprises the sequence set forth in SEQ ID NO:72, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
295. The AAV vector of any of embodiments 264-271, 273, 274, 280-286, and 288-294, wherein the dCas is a Streptococcus pyogenes dCas9 (dSpCas9) protein that comprises at least one amino acid mutation selected from D10A and H840A, with reference to numbering of positions of SEQ ID NO: 79, and/or the dCas protein comprises the sequence set forth in SEQ ID NO: 78, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
296. The AAV vector of any of embodiments 264-295, wherein the regulatory DNA element is an enhancer.
297. The AAV vector of any of embodiments 264-296, wherein the target site is located within a target region spanning the genomic coordinates chr9:69, 027, 282-69, 028, 497 from hg38 (SEQ ID NO:431), optionally wherein the target site is located within a target region spanning the genomic coordinates chr9:69, 027, 615-69, 028, 101 from hg38, optionally wherein the target site is located within a target region spanning the genomic coordinates chr9:69, 027, 825-69, 027, 875.
298. The AAV vector of any of embodiments 264-271, 273, 274, 280-286, and 288-297, wherein the target site comprises the sequence set forth in SEQ ID NO:21, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing.
299. The AAV vector of any of embodiments 264-271, 273, 274, 280-286, and 288-298, wherein the gRNA comprises a gRNA spacer sequence comprising the sequence set forth in SEQ ID NO:42, or a contiguous portion thereof of at least 14 nt.
300. The AAV vector of any of embodiments 264-271, 273, 274, 280-286, and 288-299, wherein the gRNA further comprises the sequence set forth in SEQ ID NO:44, optinally wherein the gRNA comprises the sequence set forth in SEQ ID NO:67, optionally wherein the gRNA is the gRNA sequence set forth in SEQ ID NO: 67.
301. The AAV vector of any of embodiments 272-281, 284, 285, 287-289, 296, and 297, wherein the target site comprises the sequence set forth in any one of SEQ ID NOS: 272 and 277, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing.
302. The AAV vector of any of embodiments 272-281, 284, 285, 287-289, 296, and 297, wherein the eZFP comprises the sequence set forth in any one of SEQ ID NOS: 301 and 302.
303. The AAV vector of any of embodiments 264-295, wherein the regulatory DNA element is a promoter.
304. The AAV vector of any of embodiments 264-295 and 303, wherein the target site is within a target region spanning the genomic coordinates chr9:69, 034, 900-69, 035, 900 from hg38 (SEQ ID NO:430), optionally wherein the target site is within a target region spanning the genomic coordinates chr9:69, 035, 300-69-035, 800 from hg38; chr9:69, 035, 350-69, 035, 450 from hg38; or chr9:69,035,675- 69,035,725.
305. The AAV vector of any of embodiments 264-271, 273, 274, 280-286, 288-295, 303, and
304, wherein the target site comprises a sequence selected from any of SEQ ID NOS: 1-10, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing.
306. The AAV vector of any of embodiments 264-271, 273, 274, 280-286, 288-295, and 303-
305, wherein the gRNA comprises a gRNA spacer sequence comprising a sequence selected from any of SEQ ID NOS:22-31, or a contiguous portion thereof of at least 14 nt.
307. The AAV vector of any of embodiments 264-271, 273, 274, 280-286, 288-295, and 303-
306, wherein the gRNA comprises a gRNA spacer sequence comprising SEQ ID NO:22, or a contiguous portion thereof of at least 14 nt.
308. The AAV vector of any of embodiments 264-271, 273, 274, 280-286, 288-295, and 303- 306, wherein the gRNA comprises a gRNA spacer sequence comprising SEQ ID NO:28, or a contiguous portion thereof of at least 14 nt.
309. The AAV vector of any of embodiments 264-271, 273, 274, 280-286, 288-295, 303, and 304, wherein the gRNA further comprises the sequence set forth in SEQ ID NO:44, optionally wherein the gRNA comprises a sequence selected from any of SEQ ID NOS:47-56, optionally wherein the gRNA is the gRNA sequence set forth in any of SEQ ID NOS:47-56, optionally wherein the gRNA is set forth in SEQ ID NO:47 or 53.
310. The AAV vector of any of embodiments 264-271, 273, 274, 280-286, 288-295, 303, and 304, wherein the target site comprises a sequence selected from any of SEQ ID NOS: 11-20, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing.
311. The AAV vector of any of embodiments 264-271, 273, 274, 280-286, 288-295, 303, 304, and 310, wherein the gRNA comprises a gRNA spacer sequence comprising a sequence selected from any of SEQ ID NOS:32-41, or a contiguous portion thereof of at least 14 nt.
312. The AAV vector of any of embodiments 264-271, 273, 274, 280-286, 288-295, 303, 304, 310 and 311, wherein the gRNA further comprises the sequence set forth in SEQ ID NO:46, and/or wherein the gRNA comprises a sequence selected from any of SEQ ID NOS:57-66, optionally wherein the gRNA is the gRNA set forth in any of SEQ ID NOS:57-66.
313. The AAV vector of any of embodiments 264-271, 273, 274, 280-286, 288-300, and 303- 312, wherein the gRNA spacer sequence is between 14 nt and 24 nt, or between 16 nt and 22 nt in length, optionally wherein the gRNA spacer sequence is 18 nt, 19 nt, 20 nt, 21 nt or 22 nt in length.
314. The AAV vector of any of embodiments 272-281, 284, 285, 287-289, 303, and 304, wherein the target site comprises the sequence set forth in any one of SEQ ID NOS: 280-283, 290, 299, and 583-600, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing.
315. The AAV vector of any of embodiments 272-281, 284, 285, 287-289, 303, 304, and 314, wherein the target site comprises the sequence set forth in any one of SEQ ID NOS: 299, 587, 589, and 591, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing.
316. The AAV vector of any of embodiments 272-281, 284, 285, 287-289, 303, 304, 314, and 315, wherein the target site comprises the sequence set forth in SEQ ID NO: 299, a contiguous portion thereof of at least 14 nt, or a complementary sequence of the sequence set forth in SEQ ID NO: 299.
317. The AAV vector of any of embodiments 272-281, 284, 285, 287-289, 303, 304, 314, and 315, wherein the target site comprises the sequence set forth in SEQ ID NO: 587, a contiguous portion thereof of at least 14 nt, or a complementary sequence of the sequence set forth in SEQ ID NO: 587.
318. The AAV vector of any of embodiments 272-281, 284, 285, 287-289, 303, 304, 314, and 315, wherein the target site comprises the sequence set forth in SEQ ID NO: 589, a contiguous portion thereof of at least 14 nt, or a complementary sequence of the sequence set forth in SEQ ID NO: 589.
319. The AAV vector of any of embodiments 272-281, 284, 285, 287-289, 303, 304, 314, and 315, wherein the target site comprises the sequence set forth in SEQ ID NO: 591, a contiguous portion thereof of at least 14 nt, or a complementary sequence of the sequence set forth in SEQ ID NO: 591.
320. The AAV vector of any of embodiments 272-281, 284, 285, 287-289, 303, 304, and 314, wherein the eZFP comprises the sequence set forth in any one of SEQ ID NOS: 303-307 and 439-456.
321. The AAV vector of any of embodiments 272-281, 284, 285, 287-289, 303, 304, 314, 315, and 320, wherein the eZFP comprises the sequence set forth in any one of SEQ ID NOS: 307, 441, 443, and 445.
322. The AAV vector of any of embodiments 272-281, 284, 285, 287-289, 303, 304, 314, 315, 320, and 321, wherein the eZFP comprises the sequence set forth in SEQ ID NO: 307.
323. The AAV vector of any of embodiments 272-281, 284, 285, 287-289, 303, 304, 314, 315, 320, and 321, wherein the eZFP comprises the sequence set forth in SEQ ID NO: 441.
324. The AAV vector of any of embodiments 272-281, 284, 285, 287-289, 303, 304, 314, 315, 320, and 321, wherein the eZFP comprises the sequence set forth in SEQ ID NO: 443.
325. The AAV vector of any of embodiments 272-281, 284, 285, 287-289, 303, 304, 314, 315, 320, and 321, wherein the eZFP comprises the sequence set forth in SEQ ID NO: 445.
326. The AAV vector of any of embodiments 264-325, wherein the at least one effector domain induces transcription activation.
327. The AAV vector of any of embodiments 264-326, wherein the at least one epigenetic effector domain comprises: a VP64 domain, a p65 activation domain, a p300 domain, an Rta domain, a CBP domain, a VPR domain, a VPH domain, an HSF1 domain, a TET protein domain, optionally
wherein the TET protein is TET1, a SunTag domain, a domain from DPOLA, ENL, FOXO3, HSH2D, NCOA2, NCOA3, PSA1, PYGO1, RBM39, HERC2, or NOTCH2, or a domain, portion, variant, or truncation of any of the foregoing.
328. The AAV vector of any of embodiments 264-327, wherein the at least one epigenetic effector domain comprises the sequence set forth in any of SEQ ID NOS:81, 83, 100-109, 111-122, 124, 125, 134-140, 152, and 383-396, or a domain, portion, variant, or truncation thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
329. The AAV vector of any of embodiments 264-328 wherein the at least one effector domain is fused to the N-terminus, the C-terminus, or both the N-terminus and the C-terminus, of the dCas protein or eZFP.
330. The AAV vector of any of embodiments 264-329, further comprising one or more linkers connecting the dCas protein or eZFP to the at least one effector domain, and/or further comprising one or more nuclear localization signals (NLS).
331. The AAV vector of any of embodiments 264-330 wherein the at least one effector domain comprises at least one VP16 domain, or a VP16 tetramer (“VP64”) or a variant thereof.
332. The AAV vector of any of embodiments 264-331, wherein the at least one effector domain comprises VP64.
333. The AAV vector of any of embodiments 264-332, wherein the at least one effector domain comprises a VP64 domain comprising the sequence set forth in SEQ ID NO:81 or 83, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
334. The AAV vector of any of embodiments 264-333, wherein the at least one effector domain comprises the sequence set forth in SEQ ID NO:81 or 83.
335. The AAV vector of any of embodiments 264-334, wherein the at least one epigenetic effector domain comprises: a domain from DPOLA, ENL, FOXO3, HSH2D, NCOA2, NCOA3, PSA1, PYGO1, RBM39, HERC2, or NOTCH2, or a domain, portion, variant, or truncation of any of the foregoing.
336. The AAV vector of any of embodiments 264-335, wherein the at least one epigenetic effector domain comprises the sequence set forth in any of SEQ ID NOS:383-393, or a domain, portion, or variant thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
337. The AAV vector of any of embodiments 264-336, wherein the at least one effector domain comprises a domain from NCOA2, NCOA3, FOXO3, PYGO1, or a portion or variant of any of the foregoing.
338. The AAV vector of any of embodiments 264-337, wherein each effector domain of the at least one effector domain is independently selected from an NCOA2 domain, an NCOA3 domain, a FOXO3 domain, and a PYGO1 domain.
339. The AAV vector of any of embodiments 264-338, wherein the at least one effector domain comprises a domain from NCOA2 comprising the sequence set forth in SEQ ID NO: 104 or SEQ ID NO:387, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
340. The AAV vector of any of embodiments 264-339, wherein the at least one effector domain comprises a domain from NCOA2 set forth in or SEQ ID NO:387.
341. The AAV vector of any of embodiments 264-340, wherein the at least one effector domain comprises a domain from NCOA3 comprising the sequence set forth in SEQ ID NO: 105 or SEQ ID NO:388, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
342. The AAV vector of any of embodiments 264-341, wherein the at least one effector domain comprises a domain from NCOA3 set forth in or SEQ ID NO:388.
343. The AAV vector of any of embodiments 264-342, wherein the at least one effector domain comprises a domain from FOXO3 comprising the sequence set forth in SEQ ID NO: 102 or SEQ ID NO:385, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
344. The AAV vector of any of embodiments 264-343, wherein the at least one effector domain comprises a domain from FOXO3 set forth in or SEQ ID NO:385.
345. The AAV vector of any of embodiments 264-344, wherein the at least one effector domain comprises a domain from PYGO1 comprising the sequence set forth in SEQ ID NO: 107 or SEQ ID NO:390, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
346. The AAV vector of any of embodiments 264-345, wherein the at least one effector domain comprises a domain from PYGO1 set forth in or SEQ ID NO:390.
347. The AAV vector of any of embodiments 264-346, wherein the at least one effector domain is a multipartite effector composed of at least two effector domains.
348. The AAV vector of embodiment 347, wherein the multipartite effector is composed of two effector domains.
349. The AAV vector of embodiment 348, wherein the multipartite effector is composed of three effector domains.
350. The AAV vector of any of embodiments 347-349, wherein the multipartite effector is set forth in any of SEQ ID NOS:397-418, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
351. The AAV vector of any of embodiments 347-350, wherein the multipartite effector is set forth in any of SEQ ID NOS:411-418, a portion thereof, or an amino acid sequence that has at least 90%,
91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
352. The AAV vector of any of embodiments 347-351, wherein the multipartite effector comprises, in the N-terminal to C-terminal direction, domains from FOXO3, FOXO3, and NCOA3.
353. The AAV vector of embodiment 352, wherein the multipartite effector comprises the sequence set forth in SEQ ID NO:415, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
354. The AAV vector of any of embodiments 347-351, wherein the multipartite effector comprises, in the N-terminal to C-terminal direction, domains from NCOA3, FOXO3, and FOXO3.
355. The AAV vector of embodiment 354, wherein the multipartite effector comprises the sequence set forth in SEQ ID NO:418, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
356. The AAV vector of any of embodiments 347-351, wherein the multipartite effector comprises, in the N-terminal to C-terminal direction, domains from NCOA3, FOXO3, and NCOA3.
357. The AAV vector of embodiment 356, wherein the multipartite effector comprises the sequence set forth in SEQ ID NO:413, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
358. The AAV vector of any of embodiments 347-351, wherein the multipartite effector comprises, in the N-terminal to C-terminal direction, domains from NCOA2, FOXO3, and NCOA3.
359. The AAV vector of embodiment 358, wherein the multipartite effector comprises the sequence set forth in SEQ ID NO:416, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
360. The AAV vector of any of embodiments 347-351, wherein the multipartite effector comprises, in the N-terminal to C-terminal direction, domains from PYGO1, FOXO3, and NCOA3.
361. The AAV vector of embodiment 360, wherein the multipartite effector comprises the sequence set forth in SEQ ID NO:411, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
362. The AAV vector of any of embodiments 264-361, wherein the at least one epigenetic effector domain is fused to the N-terminus of the dCas protein or eZFP.
363. The AAV vector of any of embodiments 264-361, wherein the at least one epigenetic effector domain is fused to the C-terminus of the dCas protein or eZFP.
364. The AAV vector of any of embodiments 264-361, wherein the at least one epigenetic effector domain is fused to both the N-terminus and the C-terminus, of the dCas protein or eZFP.
365. The AAV vector of any of embodiments 264-361, wherein the one or more linkers are in between any two of the components of the fusion protein, including the dCas protein or eZFP, any of the at least one effector domains, and the one or more NLS.
366. The AAV vector of any of embodiments 264-365, wherein the one or more linkers connect the dCas protein or eZFP and the at least one epigenetic effector domain.
367. The AAV vector of any of embodiments 264-366, wherein the fusion protein comprises the sequence set forth in SEQ ID NO:71 or 77, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto, or the sequence set forth in SEQ ID NO:71 or 77, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
368. The AAV vector of any of embodiments 264-367, wherein the fusion protein comprises the sequence set forth in any one of SEQ ID NOS:266-268 and 315-319, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
369. The AAV vector of any of embodiments 264-368, wherein the AAV vector is selected from the group consisting of: AAV1, AAV2, AAV3, AAV4, AAV5, AAV6, AAV7, AAV8, AAV9, AAV10, AAV11, AAV12, AAV-DJ, and AAVrh74.
370. The AAV vector of any of embodiments 264-369, wherein the AAV vector is AAV6.
371. The AAV vector of any of embodiments 264-369, wherein the AAV vector is AAV9.
372. The AAV vector of any of embodiments 264-369, wherein the AAV vector is AAV-DJ.
373. The AAV vector of any of embodiments 264-369, wherein the AAV vector is AAVrh74.
374. The AAV vector of any of embodiments 264-373, wherein the vector exhibits tropism for a nervous system cell, optionally a neuron, a heart cell, optionally a cardiomyocyte, a skeletal muscle cell, a fibroblast, an induced pluripotent stem cell, and/or a cell derived from any of the foregoing, or for a combination of any of the foregoing cells.
375. The AAV vector of any of embodiments 264-374, wherein the vector exhibits tropism for induced pluripotent stem cells.
376. The AAV vector of any of embodiments 264-375, wherein the vector exhibits tropism for neurons and cardiomyocytes.
377. A cell comprising the eZFP of any of embodiments 1-187, the fusion protein of any of embodiments 186-246, the polynucleotide of embodiment 247, the plurality of polynucleotides of embodiment 248, the vector of any of embodiments 249-263, the AAV vector of any of embodiments 264-376, or a portion or component of any of the foregoing, or a combination of any of the foregoing.
378. The cell of embodiment 377, wherein the cell is a nervous system cell, optionally a neuron, a heart cell, optionally a cardiomyocyte, a skeletal muscle cell, a fibroblast, an induced pluripotent stem cell, and/or a cell derived from any of the foregoing.
379. The cell of embodiment 377 or 378, wherein the cell is from a subject that has or is suspected of having Friedreich’s ataxia (FA).
380. A pharmaceutical composition comprising the eZFP of any of embodiments 1-187, the fusion protein of any of embodiments 188-246, the polynucleotide of embodiment 247, the plurality of polynucleotides of embodiment 248, the vector of any of embodiments 249-263, the AAV vector of any
of embodiments 264-376, or a portion or component of any of the foregoing, or a combination of any of the foregoing.
381. The pharmaceutical composition of embodiment 380, for use in treating a disease, condition, or disorder in a subject.
382. The pharmaceutical composition of embodiment 381, wherein the disease, condition, or disorder is Friedreich’s ataxia and/or a GAA trinucleotide repeat expansion in the FXN locus.
383. The pharmaceutical composition of embodiment 381 or 382, wherein following administration of the pharmaceutical composition, the expression of FXN is increased in cells of the subject.
384. A method for increasing the expression of FXN in a cell, the method comprising introducing into the cell: the eZFP of any of embodiments 1-187, the fusion protein of any of embodiments 188-246, the polynucleotide of embodiment 247, the plurality of polynucleotides of embodiment 248, the vector of any of embodiments 249-263, the AAV vector of any of embodiments 264-376, the pharmaceutical composition of any of embodiments 380-383, or a portion or component of any of the foregoing, or a combination of any of the foregoing.
385. The method of embodiment 384, wherein the cell is from and/or in a subject that has or is suspected of having Friedreich’s ataxia.
386. The method of embodiment 384 or 385, wherein the cell exhibits reduced expression of FXN in comparison to a reference cell from an individual not having Friedreich’s ataxia and/or a GAA trinucleotide expansion in the FXN locus.
387. A method for increasing the expression of FXN in a cell in a subject, the method comprising administering to the subject: the eZFP of any of embodiments 1-187, the fusion protein of any of embodiments 188-246, the polynucleotide of embodiment 247, the plurality of polynucleotides of embodiment 248, the vector of any of embodiments 249-263, the AAV vector of any of embodiments 264-376, the pharmaceutical composition of any of embodiments 380-383, or a portion or component of any of the foregoing, or a combination of any of the foregoing.
388. A method of treating a subject in need thereof, the method comprising administering to the subject: the eZFP of any of embodiments 1-187, the fusion protein of any of embodiments 188-246, the polynucleotide of embodiment 247, the plurality of polynucleotides of embodiment 248, the vector of any of embodiments 249-263, the AAV vector of any of embodiments 264-376, the pharmaceutical composition of any of embodiments 380-383, or a portion or component of any of the foregoing, or a combination of any of the foregoing.
389. The method of embodiment 387 or 388, wherein the subject has or is suspected of having Friedreich’s ataxia, and/or a GAA trinucleotide expansion in the FXN locus.
390. The method of any of embodiments 384-389, wherein the introducing or administering is carried out in vivo or ex vivo.
391. The method of any of embodiments 384-390, wherein the cell and/or subject exhibits reduced expression of FXN prior to performing the method.
392. The method of embodiment 391, wherein the reduced expression of FXN is reduced in comparison to a reference individual not having Friedreich’ s ataxia and/or a GAA trinucleotide repeat expansion in the FXN locus, and/or a reference cell therefrom.
393. The method of any of embodiments 386 and 389-392, wherein the GAA trinucleotide repeat expansion is in a first intron of a FXN gene, and comprises at least 50, at least 60, at least 70, at least 80, at least 90, at least 100, at least 150, at least 200, at least 300, at least 400, at least 500, at least 600, at least 700, at least 800, at least 900, at least 1000, or more repeated GAA trinucleotides.
394. The method of any of embodiments 384-393, wherein following the introducing or administering, the expression of FXN is increased in the cell and/or subject.
395. The method of embodiment 394, wherein the expression of FXN is increased in the cell or cells of the subject by at least about 1.2-fold, 1.25-fold, 1.3-fold, 1.4-fold, 1.5-fold, 1.6-fold, 1.7-fold, 1.75-fold, 1.8-fold, 1.9-fold, 2-fold, 2.5-fold, 3-fold, 4-fold, or 5-fold; and/or the expression is increased by less than about 10-fold, 9-fold, 8-fold, 7-fold or 6-fold.
396. The method of embodiment 394 or 395, wherein the expression of FXN is increased in the cell or cells of the subject to a level that is at least at or about 30%, 40%, 50%, 60%, 70%, 75%, 80%, 85%, 90%, 95%, 100%, 105%, 110%, 120%, 125%, 150%, 175%, 200%, 225%, 250%, 300%, 400%, or 500%, of the expression level of FXN in a reference cell from an individual not having Friedreich’s ataxia and/or a cell not having a GAA trinucleotide repeat expansion in the FXN gene.
397. The method of any of embodiments 394-396, wherein the expression of FXN is increased in the cell or cells of the subject to a level that is less than at or about 200%, 300%, 400%, 500%, 600%, 700%, 800%, 900%, or 1000% of the expression level of FXN in a reference cell from an individual not having Friedreich’s ataxia and/or a cell not having a GAA trinucleotide repeat expansion in the FXN gene.
398. The method of any of embodiments 394-397, wherein the expression is measured by the amount of mRNA encoding the FXN protein, and/or the amount of FXN protein.
399. The method of any of embodiments 385-399, wherein the subject is a human.
400. A cell comprising an epigenetic modification produced by the method of any of embodiments 384-399.
VIII. EXAMPLES
[0644] The following examples are included for illustrative purposes only and are not intended to limit the scope of the invention.
Example 1: CRISPR-Cas mediated transcriptional activation of frataxin (FXN) in induced pluripotent stem cells (iPSCs)
[0645] Guide RNAs (gRNAs) targeting the human frataxin (FXN) gene promoter were generated and transduced with deactivated Cas9 (dCas9)-transcriptional activator fusion proteins into wild-type (WT) and Friedreich’ s ataxia (FA) patient-derived cells and frataxin mRNA expression was assessed.
A. dSpCas9-2xVP64, dSaCas9-2xVP64 and gRNA Constructs
[0646] Plasmids encoding the heterologous fusion proteins dSpCas9-2xVP64 or dSaCas9-2xVP64 were prepared. Fusion proteins included a modified Cas9 engineered to lack endonuclease activity (deactivated Cas9, dCas9) from .S', aureus (dSaCas9) or .S', pyogenes (dSpCas9), fused with 2 copies of the transcriptional activator VP64 (2xVP64), one copy at each of the N- and C-terminals.
[0647] Plasmids were prepared, each encoding a gRNA targeting one of multiple sequences in the human frataxin gene promoter region (Homo sapiens (human) genome assembly GRCh38 (hg38) chr9:69, 034, 622-69, 036, 670), as shown in FIG. 1A. The gRNAs included a 20-22 nucleotide (nt) spacer sequence and a constant scaffold sequence. Different gRNAs were designed for dSaCas9-2xVP64 and dSpCas9-2xVP64, based on the protospacer-adjacent motif (PAM) sequences 5’-NNGRRT-3’ or 5’- NGG-3’, respectively. gRNAs for dSaCas9 further comprised the scaffold sequence set forth in SEQ ID NO:44 (encoded by SEQ ID NO:43). gRNAs for dSpCas9 further comprised the scaffold sequence set forth in SEQ ID NO:46 (encoded by SEQ ID NO:45). The promoter-targeting gRNAs are indicated in
Table El.
B. Delivery of dCas9-2xVP64 and gRNAs to iPSCs
[0648] Wild-type induced pluripotent stem cells (iPSCs) cultured on Matrigel (Corning) with mTeSR Plus medium (Stemcell Technologies) were transduced using lentivirus with a plasmid encoding one of the frataxin promoter-targeting gRNAs described in Table El above and corresponding dSaCas9- 2xVP64 (e.g., as set forth in SEQ ID NO:71) or dSpCas9-2xVP64 (e.g., as set forth in SEQ ID NO:77). Cells were incubated for 2 days, then cultured in the presence of 1 pg/mL puromycin to enrich for transduced cells for an additional 2 days. Cells were then split using Accutase cell dissociation reagent (ThermoFisher #A1110501) and cultured for an additional 3 days prior to assessing frataxin expression levels.
C. Assessing frataxin expression in iPSCs cultured with dCas9-2xVP64 and gRNA
[0649] Frataxin mRNA expression levels were assessed by RT-qPCR. Approximately 30,000 - 60,000 transduced iPSCs per condition were harvested and assayed by RT-qPCR using a Cells-to-Ci kit (ThermoFisher) to assess relative changes in expression of frataxin mRNA. For each condition, frataxin (FXN) expression was normalized to GAPDH expression. Taqman probes were used for FXN (ThermoFisher, Cat#: Hs00175940_ml) and GAPDH (ThermoFisher, Cat#: Hs02786624_gl). Results were assessed as fold-increase expression of FXN relative to control cells that were not transduced with gRNA.
[0650] Frataxin promoter-targeted gRNA A and gRNA G with dSaCas9-2xVP64 induced a greater
than 2-fold increase in frataxin mRNA expression, compared to controls without gRNA (FIG. IB). Similar results were obtained for gRNA K, gRNA L, and gRNA R with dSpCas9-2xVP64 (FIG. 1C).
[0651] The results indicate that a dCas9-effector fusion protein with gRNA targeted to the frataxin promoter can increase frataxin mRNA expression levels in human iPSCs.
D. Transcriptional activation of frataxin in Friedreich’s ataxia patient-derived iPSCs (FA-iPSCs) harboring large repeat expansions
[0652] Friedreich’ s ataxia (FA) is associated with GAA trinucleotide expansion mutations in the first intron of the frataxin gene. Trinucleotide expansion results in reduced frataxin expression, leading to disease. Increasing trinucleotide expansion lengths are associated with decreased frataxin expression levels, earlier disease onset, and increased disease severity.
[0653] iPSCs generated from Friedreich’s ataxia patients iPSCs (FA-iPSCs) containing different lengths of expanded GAA trinucleotide repeats (FA-iPSC 1 with 604/734 repeats; FA-iPSC 2 with 867/867 repeats) in the frataxin gene were (a) left untreated, (b) transduced with dSaCas9-2xVP64 and a control gRNA, or (c) transduced with dSaCas9-2xVP64 and promoter-targeting gRNA A and gRNA G. Frataxin mRNA expression was assessed by RT-qPCR as described above. FA-iPSCs transduced with the promoter-targeting gRNAs exhibited increased frataxin mRNA expression in comparison to FA- iPSCs that were untreated or transduced with control gRNA (FIG. 2). The results indicated that dCas9- transcriptional activator with gRNA targeted to the frataxin promoter can increase frataxin mRNA expression levels in the context of FA-iPSCs with various lengths of GAA trinucleotide expansion mutations. The results support the utility of frataxin-targeting gRNAs and dCas9-transcriptional activator fusion proteins in restoring frataxin expression levels in cells from FA patients and potential treatment of FA.
Example 2; A saturating screen of gRNAs targeting frataxin for CRISPR-based transcriptional activation
[0654] A saturating library of gRNAs was designed and generated for a broad genomic region including and surrounding the frataxin gene. gRNAs were tested in a pooled format in iPSCs expressing dCas9-transcriptional activator fusion proteins, to identify gRNAs that facilitate increased frataxin expression.
A. Design and generation of frataxin-saturating gRNA library
[0655] A saturating library of SaCas9 gRNAs was designed and generated, targeting a broad genomic region of approximately 250 kb including and surrounding the frataxin gene (genomic region: hg38 chr9:68, 940, 179-69, 205 ,519). gRNAs were designed according to the PAM sequence for SaCas9 (5’-NNGRRT-3’). gRNAs were excluded if spacers contained 5 or more continuous T nucleotides to avoid premature termination of the gRNA transcripts. gRNAs with a high level of predicted off-target activity in human cells were also excluded. In total, 6173 gRNAs, including 300 control gRNAs, were
designed, cloned, and tested for ability to transcriptionally activate the expression of frataxin, together with dSaCas9-2xVP64.
B. Flow cytometry assay for frataxin protein expression levels
[0656] WT-iPSCs and long GAA trinucleotide repeat FA-iPSCs were assessed by flow cytometry for levels of frataxin protein expression. Cells were fixed with 4% paraformaldehyde (PF A) at room temperature, permeabilized with MeOH overnight at -20°C and labeled with a mouse monoclonal antifrataxin fluorophore-conjugated antibody (abeam abl56033). Cells were then assessed by flow cytometry for frataxin protein expression levels. The results showed that FA-iPSCs exhibited reduced frataxin expression in comparison to WT-iPSC control cells (FIG. 3).
C. Screen of frataxin-saturating gRNA library for transcriptional activation of frataxin
[0657] gRNAs with strong activity of increasing frataxin expression were identified from screening the saturating gRNA library. WT-iPSCs were transduced with a lentiviral construct containing dSaCas9- 2xVP64 and cultured in the presence of puromycin to select for transduced cells. The WT-iPSCs expressing dSaCas9-2xVP64 were then transduced using lentivirus with the pooled gRNA library at a multiplicity of infection (MOI) of 0.4 (i.e., 0.4 viral particles per cell) and incubated for 1 day. Cells were then cultured for 7 days in the presence of 1 pg/mL puromycin to enrich for transduced cells. Cells were then fixed with 4% PFA at room temperature, permeabilized with MeOH overnight at -20°C and labeled with a mouse monoclonal anti-frataxin fluorophore-conjugated antibody (abeam abl56033). Cells were sorted by flow cytometry into 2 populations, representing the top 10% and bottom 10% of cells based on frataxin protein expression.
[0658] Sequencing was performed to compare the abundance of specific gRNAs between the frataxin-low and frataxin-high population, and to identify gRNAs enriched in the frataxin-high population. Genomic DNA was isolated from the sorted populations. Targeted PCR was performed to amplify the gRNA spacers and append sequencing adapters. Each sample was barcoded separately. Samples were then sequenced using an Illumina MiSeq System. Three sequencing replicates of the frataxin-high population were compared to three replicates of the frataxin-low population using DEseq2, a method for detecting differentially expressed transcripts.
[0659] gRNAs enriched in the frataxin-high population were identified based on sequencing analysis (FIG. 4). gRNAs that were enriched in the frataxin-high population included the promotertargeting gRNA A from Example 1 above, and a second gRNA (gRNA U; target sequence set forth in SEQ ID NO:21, spacer sequence set forth in SEQ ID NO:41, e.g. as shown in Table El). gRNA U targeted a region approximately 7kb upstream of the frataxin promoter. This region (hg38 chr9:69, 027, 282-69, 028, 497) exhibits hallmark chromatin signatures of an enhancer element, including specific histone modifications (H3K4Mel) and DNase I hypersensitivity (FIG. 5). Although similar enhancer-like signatures were present in the screened region (for example, hg38 chr9:69,044,201-
69,045,347 and hg38 chr9: 69,030,752-69,031,507), gRNAs targeting these regions were not identified as significantly enriched in the frataxin-high population.
[0660] The results showed that a gRNA targeting an upstream enhancer region of the frataxin gene, together with a dCas9-transcriptional activator fusion protein, leads to a strong increase in frataxin expression, supporting the utility of the exemplary DNA-targeting system and gRNAs in restoring frataxin expression levels in FA patient cells and potential treatment of FA.
Example 3: Combinatorial modulation of frataxin expression
[0661 ] An exemplary frataxin enhancer region-targeting gRNA (gRNA U, described in Example 2C above) was tested alone and in combination with promoter-targeting gRNAs in iPSCs, for transcriptional activation of frataxin.
D. Enhancer region-targeting gRNA
[0662] WT-iPSCs were transduced with dSaCas9-2xVP64 and (a) gRNA A (promoter-targeting, see Example 1 above), (b) gRNA U (enhancer-targeting, see Example 2C above), (c) a control gRNA (not frataxin-targeting), or (d) an empty gRNA vector. Cells were incubated for 2 days, then cultured in the presence of 1 pg/mL puromycin to enrich for transduced cells for an additional 2 days. Cells were then split using Accutase cell dissociation reagent (ThermoFisher #A1110501) and cultured for an additional 3 days prior to assessing frataxin expression levels. Relative frataxin mRNA expression levels were assessed by RT-qPCR as described above.
[0663] Cells transduced with gRNA A or gRNA U exhibited a greater than 1.5-fold increase in frataxin mRNA expression in comparison to cells transduced with the control gRNA or empty gRNA vector (FIG. 6).
E. iPSC lines stably expressing frataxin promoter-targeting gRNAs
[0664] WT-iPSCs or FA-iPSCs harboring short or long frataxin trinucleotide repeats were transduced with lentivirus containing expression vectors for frataxin promoter-targeting gRNAs A or G. Cells were then selected for transduced cells with blasticidin and stable cell lines were established expressing the promoter-targeting gRNAs, as indicated in Table E2.
F. Combinatorial modulation of frataxin mRNA expression with promoter- and enhancer-
targeting gRNAs
[0665] The enhancer-targeting gRNA U was assessed for activation of frataxin expression when paired with promoter-targeting gRNAs in the gRNA-expressing iPSCs.
[0666] Cells were transduced using lenti virus according to the following conditions: (a) control lentivirus conferring puromycin resistance only, (b) dSaCas9-2xVP64 and a non-frataxin targeting control gRNA, or (c) dSaCas9-2xVP64 and enhancer-targeting gRNA U. After transduction, cells were incubated for 2 days, then cultured in the presence of 1 pg/mL puromycin to enrich for transduced cells for an additional 2 days. Cells were then split using Accutase cell dissociation reagent (ThermoFisher #A1110501) and cultured for an additional 3 days prior to assessing frataxin expression levels. Approximately 60,000 cells per condition were harvested and assessed for relative frataxin mRNA expression levels by RT-qPCR as described above.
[0667] For all cell lines tested, cells transduced with dSaCas9-2xVP64 and gRNA U exhibited increased expression of frataxin in comparison to both control-transduced and dSaCas9-2xVP64/control gRNA-transduced cells, as shown for cell lines WT-gRNA A-iPSC (FIG. 7A), WT-gRNA G-iPSC (FIG. 7B), FA(short)-gRNA A-iPSC (FIG. 7C), FA(short)-gRNA G-iPSC (FIG. 7D), FA(long)-gRNA A-iPSC (FIG. 7E), and FA(long)-gRNA G-iPSC (FIG. 7F).
[0668] The results indicate that an exemplary enhancer region-targeting gRNA (gRNA U) facilitates further transcriptional activation of frataxin when paired with exemplary promoter-targeting gRNAs (gRNA A or gRNA G), both in the context of WT-iPSCs and FA-iPSCs with short and long trinucleotide repeats. These results support the utility of the exemplary DNA-targeting system and gRNAs in restoring frataxin expression levels in FA patient cells and potential treatment of FA.
Example 4; Rescue of frataxin mRNA and protein expression in FA-iPSCs
[0669] Frataxin mRNA and protein expression were compared between WT-iPSCs and FA-iPSCs. FA-iPSCs were transduced with dCas9-transcriptional activator fusion protein and gRNAs, and assessed for rescue of frataxin expression in comparison to WT-iPSCs.
A. Rescue of frataxin mRNA expression with promoter- and enhancer-targeting gRNAs in FA- iPSCs
[0670] FA-iPSC cell lines containing long trinucleotide repeats and transduced with gRNAs, (FA(long)-gRNA A-iPSC and FA(long)-gRNA G-iPSC described in Table E2 above), exhibited severely reduced frataxin mRNA expression levels in comparison to WT-iPSCs, as assessed by RT-qPCR (approximately 15% WT expression for both lines) (FIG. 8). Transduction of FA(long)-gRNA A-iPSC and FA(long)-gRNA G-iPSC with dSaCas9-2xVP64 and a control gRNA increased frataxin mRNA expression to approximately 35% and approximately 50% WT levels, respectively. Transduction of FA(long)-gRNA G-iPSC with dSaCas9-2xVP64 and enhancer-targeted gRNA U further increased frataxin mRNA expression to levels comparable to WT (FIG. 8).
B. Rescue of frataxin protein expression with promoter- and enhancer-targeting gRNAs in FA- iPSCs
[0671] Modulation of frataxin protein expression using promoter- and enhancer-targeting gRNAs in WT- and FA-iPSCs was assessed. For protein measurement experiments, cells were transduced and incubated for 2 days, then cultured in the presence of 1 pg/mL puromycin to enrich for transduced cells for an additional 2 days. Cells were then split using Accutase cell dissociation reagent (ThermoFisher #A1110501) and cultured for an additional 3 days prior to assessing frataxin protein expression levels. Approximately 0.5 to 1.0 million cells per condition were harvested and assessed for frataxin protein expression levels. Frataxin protein was measured by a Human Frataxin ELISA Kit (abeam #abl76112) and normalized to total protein as assessed by Bicinchoninic acid (BCA) assay.
[0672| WT-gRNA G-iPSCs were (a) left untreated, (b) transduced with dSaCas9-2xVP64 and a control gRNA, or (c) transduced with dSaCas9-2xVP64 and the frataxin enhancer-targeting gRNA U. Transduced cells exhibited a greater than 2-fold increase in Frataxin protein expression (FIG. 9A).
[0673] FA(short)-gRNA A-iPSCs (as described in Table E2 above) expressed Frataxin protein at approximately 30% of WT expression levels (FIG. 9B). Transduction of dSaCas9-2xVP64 into FA(short)-gRNA A-iPSCs significantly increased frataxin protein expression to up to approximately 75% of WT levels. In addition, FA(short)-gRNA G-iPSCs transduced with dSaCas9-2xVP64 and enhancer- targeted gRNA U exhibited increased expression of frataxin protein in comparison to the same cell line transduced with dSaCas9-2xVP64 and a control gRNA (FIG. 9B). These results indicate that gRNA U facilitates further increase of frataxin protein expression when paired with a promoter-targeting gRNA, rescuing the reduced frataxin protein expression in the context of FA-associated trinucleotide repeat expansion.
[0674] FA(long)-gRNA A-iPSC and FA(long)-gRNA G-iPSC cells each expressed Frataxin protein at approximately 15% of WT expression levels (FIG. 9C). Transduction of FA(long)-gRNA A-iPSCs with dSaCas9-2xVP64 and gRNA G increased frataxin protein expression to approximately 30% of WT levels. Transduction of FA(long)-gRNA G-iPSC with dSaCas9-2xVP64 and gRNA U also increased frataxin protein expression to approximately 35% of WT levels. Cells in both of these conditions expressed frataxin at higher levels than cells expressing dSaCas9-2xVP64 and a single gRNA (FIG. 9C).
[0675] These results showed a substantial increase of frataxin mRNA and protein expression in FA- iPSCs, facilitated by combinations of gRNAs targeted to promoter and enhancer regions of the frataxin gene, in some cases, to a level similar to WT levels. Friedreich’s ataxia (FA) patients expressing frataxin protein above 35% of the frataxin protein levels observed in individuals with no frataxin trinucleotide repeat display reduced or no symptoms of FA. The results showing increased frataxin protein expression to at least 35% of WT levels in FA-iPSCs with long trinucleotide repeats supports therapeutic utility of the dCas9-transcriptional activator fusion protein and exemplary gRNAs targeting frataxin.
Example 5; Large-scale screen for dSaCas9 transcriptional activator and repressor fusion
proteins
[0676] A library of plasmids was generated encoding fusion proteins comprising fragments of nuclear localized proteins, fused to the N-terminus or C-terminus of dCas9. The libraries were screened separately in a pooled format to identify activators and repressors of frataxin in combination with an exemplary promoter-targeting gRNA.
A. dSaCas9-effector screen
[0677] A library of plasmids was generated encoding fusion proteins comprising protein fragments of nuclear localized proteins fused to the N-terminus of dSaCas9. A second library was generated with the protein fragments fused to the C-terminus of dSaCas9. The two libraries were each screened separately in a pooled format using iPSCs expressing an exemplary frataxin promoter-targeting gRNA A, as described in Example 3B above (WT-gRNA A-iPSC cells).
[0678] dSaCas9 fusion proteins comprising nuclear localized protein fragments facilitating transcriptional activation (i.e. transcriptional activation domains) or repression (i.e. transcriptional repressors) of frataxin expression were identified as follows. WT-gRNA A-iPSC cells were transduced using lentivirus with the pooled dSaCas9-effector library at a multiplicity of infection (MOI) of 0.4 and incubated for 1 day. Cells were then cultured in the presence of 1 pg/mL puromycin to enrich for transduced cells for 7 days. Cells were then fixed with 4% PFA at room temperature, permeabilized with MeOH overnight at -20°C and labeled with a mouse monoclonal anti-frataxin fluorophore-conjugated antibody (abeam abl56033). Cells were sorted by flow cytometry into 2 populations comprising the top 10% and bottom 10% of cells based on frataxin protein expression.
[0679] Sequencing was performed to identify effectors (i.e., nuclear localized protein fragments) enriched in the frataxin-high population (activators) and frataxin-low population (repressors). Genomic DNA was isolated from the sorted populations. Targeted PCR was performed to amplify the protein fragment sequences and append sequencing adapters. Each sample was barcoded separately. Samples were then sequenced using an Illumina MiSeq System. Three replicates of the frataxin-high and frataxin- low population were compared using DEseq2, a method for detecting differentially expressed transcripts.
[0680] Based on sequencing results, 9 activators and 211 repressors of frataxin expression were identified from the N-terminal protein fragment screen (FIG. 10A). 5 activators and 208 repressors were identified from the C-terminal protein fragment screen (FIG. 10B). 3 of the activator domains were identified in both screens, including fragments from the proteins NCOA2, NCOA3, and PYGO1.
B. Effector Validation
[0681 ] dSaCas9-effector fusion proteins (i.e., comprising nuclear localized protein fragments) identified in the screen described above were verified individually. WT-gRNA A-iPSCs were transduced with dSaCas9-2xVP64 (positive control) or the identified fusion proteins containing transcriptional activators (e.g. the transcriptional activation domains shown in Table 3). from the screen described
above. Resulting frataxin mRNA expression was measured by RT-qPCR as described above, in comparison to control cells transduced with a dSaCas9 fusion protein having no effect on frataxin expression levels (control peptide; amino acid sequence set forth in SEQ ID NO:99 and encoded by the nucleotide sequence set forth in SEQ ID NO: 86) as determined in the effector screen (N-terminal fusions: FIG. 11A; C-terminal fusions: FIG. 11B). dSaCas9 effector fusion proteins comprising fragments of the genes NC0A2 and PY0G1 led to upregulation of frataxin mRNA. The results showed that several of the identified dCas9-effector fusion proteins and an exemplary frataxin promoter-targeting gRNA resulted in an increase in frataxin mRNA expression. dSaCas9-effector fusion proteins were also tested in FA(long)- gRNA A-iPSCs (N-terminal fusions: FIG. 12A; C-terminal fusions: FIG. 12B). dSaCas9 effector fusion proteins comprising fragments of the genes F0X03, HSH2D, and HERC2 led to upregulation of frataxin mRNA.
[0682] The results show that dCas9 effector fusion proteins identified in the screen and an exemplary frataxin promoter-targeting gRNA can facilitate upregulation of frataxin expression in WT- and FA-iPSCs. These results support the utility of the exemplary DNA-targeting system and gRNAs in restoring frataxin expression levels in FA patient cells and potential treatment of FA.
Example 6: Rescue of frataxin expression in differentiated cells derived from FA-iPSCs
[0683] Various differentiated cell types derived from FA-iPSCs, including cardiomyocytes and neurons, were transduced with a dCas9-transcriptional activator fusion protein and gRNAs, and assessed for rescue of frataxin expression in comparison to WT-iPSC-derived cardiomyocytes and neurons, respectively.
A. Rescue of frataxin expression in FA-iPSC-derived cardiomyocytes
[0684] FA-iPSC-derived cardiomyocytes harboring expanded GAA trinucleotide repeats (867/867) were transduced with dSaCas-2xVP64 and frataxin (FXN) promoter and enhancer-targeting gRNAs (gRNA A; gRNA G; gRNA A + gRNA G; or gRNA G + gRNA U) using lentivirus, generally as described in Examples 1-4 above. Control cells were transduced with a non-targeting gRNA (NT gRNA), or with a puromycin resistance cassette alone (puro control). WT-iPSC-derived cardiomyocytes were included as controls to assess rescue of FXN expression (WT line + puro control). Transduced cells were enriched via puromycin selection from day 3 to day 10 post-transduction, and cells were harvested for analysis on day 10.
[0685] FXN mRNA expression levels were assessed by RT-qPCR (normalized to GAPDH mRNA) in comparison to FXN mRNA expression levels in WT control cells.
[0686] As shown in FIG. 13A, FA-iPSC-derived cardiomyocytes exhibited substantially reduced FXN mRNA expression levels in comparison to WT controls. In comparison, transduction of FA-iPSC- derived cardiomyocytes with dSaCas9-2xVP64 and FXN promoter- or enhancer-targeting gRNAs (gRNA G; gRNA A + gRNA G; or gRNA G + gRNA U) increased FXN mRNA expression in
comparison to control FA-iPSC-derived cardiomyocytes (puro control). Furthermore, FXN mRNA expression was increased to levels comparable to or greater than levels in WT control cells.
[0687] FXN protein expression levels were assessed by ELISA (normalized to total protein levels) in comparison to FXN protein expression levels in WT control cells.
[0688] As shown in FIG. 13B, FA-iPSC-derived cardiomyocytes exhibited substantially reduced FXN protein expression levels in comparison to WT controls. However, transduction of FA-iPSC- derived cardiomyocytes with dSaCas9-2xVP64 and FXN promoter- and enhancer-targeting gRNAs (gRNA A; gRNA G; gRNA A + gRNA G; or gRNA G + gRNA U) increased FXN protein expression in comparison to control FA-iPSC-derived cardiomyocytes (puro control). Furthermore, FXN protein expression was increased to levels comparable to or greater than levels in WT control cells.
[0689| These results showed a substantial increase of frataxin mRNA and protein expression in FA- iPSC-derived cardiomyocytes, when transduced with a dCas9-transcriptional activator and individual or combinations of gRNAs targeted to promoter and enhancer regions of the frataxin gene, including to levels comparable to or higher than WT levels.
B. Rescue of frataxin expression in FA-iPSC-derived neurons
[0690] FA-iPSC-derived neurons harboring expanded GAA trinucleotide repeats (867/867) were transduced with dSaCas-2xVP64 and FXN promoter and enhancer-targeting gRNAs (gRNA A; gRNA G; gRNA A + gRNA G; or gRNA G + gRNA U) using lentivirus, generally as described in Examples 1-4 above. Control cells were transduced with a non-targeting gRNA (NT gRNA), or with a puromycin resistance cassette (amino acid sequence set forth in SEQ ID NO: 133; exemplary nucleotide sequence encoding puromycin resistance cassette set forth in SEQ ID NO: 132) alone (puro control). WT-iPSC- derived neurons were included as controls to assess rescue of FXN expression (WT line + puro control). Transduced cells were enriched via puromycin selection from day 3 to day 10 post-transduction, and cells were harvested for analysis on day 10.
[0691] Similar to the assessment in differentiated cardiomyocytes, FXN mRNA expression levels were assessed by RT-qPCR (normalized to GAPDH) in comparison to FXN mRNA expression levels in WT control cells.
[0692] As shown in FIG. 14, FA-iPSC-derived neurons exhibited substantially reduced FXN mRNA expression levels in comparison to WT controls. Transduction of FA-iPSC-derived neurons with dSaCas9-2xVP64 and FXN promoter- and enhancer-targeting gRNAs (gRNA A; gRNA G; gRNA A + gRNA G; or gRNA G + gRNA U) increased FXN mRNA expression in comparison to control FA-iPSC- derived neurons (puro control). Furthermore, FXN mRNA expression was increased to levels comparable to levels in WT control cells.
[0693] These results showed a substantial increase of frataxin mRNA expression in FA-iPSC- derived neurons, when transduced with a dCas9-transcriptional activator and individual or combinations of gRNAs targeted to promoter and enhancer regions of the frataxin gene, including to levels comparable
to or higher than WT levels.
C. Conclusion
[0694] These results together showed that transduction with a dCas9-transcriptional activator and individual or combinations of gRNAs targeted to promoter and enhancer regions leads to a substantial increase of frataxin mRNA expression in FA-iPSC-derived cardiomyocytes and neurons, and frataxin protein expression in FA-iPSC-derived cardiomyocytes, including rescuing mRNA or protein expression levels to levels comparable to or higher than WT levels.
[0695] In some cases, Friedreich’s ataxia (FA) patients expressing frataxin protein above 35% of levels observed in individuals with no FA display reduced or no symptoms of FA. The results showing increased frataxin expression to at least 35% or higher of WT levels in FA-iPSC-derived cardiomyocytes supports the therapeutic utility of the dCas9-transcriptional activator fusion protein and exemplary gRNAs targeting frataxin, including in heart cells, such as cardiomyocytes. These results further support therapeutic utility of the exemplary dCas9-transcriptional activator fusion protein and exemplary gRNAs targeting frataxin, including in various cell types, including heart cells, such as cardiomyocytes, and in neurons.
Example 7; Transcriptional activation of frataxin in non-human primate fibroblasts
[0696] Cells from different organisms, such as non-human primates, were transduced with a dCas9- transcriptional activator fusion protein and FXN promoter-targeting gRNAs, and assessed for transcriptional activation of FXN.
[0697] Non-human primate fibroblast cell lines, including those shown in Table E3, were obtained and cultured for the following experiments.
[0698] Lentiviral vectors for expression in the non-human primate cell lines were designed and cloned, each comprising nucleic acid sequences encoding dSaCas9-2xVP64 and gRNA A, gRNA G, both gRNA A and gRNA G, described in Examples 1-4 above, which also target conserved sequences of the non-human primate FXN locus, or a non-targeting gRNA (NT gRNA). Vectors further encoded a puromycin resistance cassette for enrichment of transduced cells. The vectors were transduced into the non-human primate cell lines, and transduced cells were enriched using 6 pg/mL puromycin from day 2 to day 7 post-transduction.
[0699] Cells were collected 7 days post-transduction and analyzed by RT-qPCR for expression of
FXN mRNA. RT-qPCR Taqman probes for FXN expression (and control GAPDH expression) are shown in Table E3 for the non-human primate cell lines. FXN expression was compared to expression in control cells transduced with a puromycin resistance cassette alone.
[0700] As shown in FIG. 15, transduction of non-human primate cell lines with the dCas9- transcriptional activator fusion protein and FXN promoter-targeting gRNAs led to increased transcription of FXN in comparison to control cells. These results further support the utility of the exemplary dCas9- transcriptional activator fusion protein and exemplary gRNAs targeting frataxin, in increasing transcription of FXN in different cell types from different organisms.
Example 8: Transcriptional activation of frataxin in vivo in a Friedreich’s ataxia (FA) mouse model
[0701] A humanized mouse model for Friedreich’s ataxia (FA) was treated with a dCas9- transcriptional activator fusion protein and frataxin (FXN) targeting gRNAs and assessed for in vivo transcriptional activation of FXN.
[0702] Adeno-associated virus (AAV) vectors were designed and generated to evaluate the in vivo effect of the dCas9-transcriptional activator fusion protein and different exemplary gRNAs described in Examples 1-4 above. Each single-stranded AAV9 (ssAAV9) vector included nucleic acid sequences encoding dSaCas9-2xVP64 and one of the following: 1) promoter-targeting gRNA A (see Example 1 above); 2) promoter-targeting gRNA G (see Example 1 above); 3) enhancer-targeting gRNA U (see Example 2C above); and 4) a non-targeting gRNA (NT gRNA). For the combination test group, ssAAV9 vectors encoding gRNA U and ssAAV9 vectors encoding gRNA G were combined and tested along with the other groups.
[0703] 6-8 week old male and female humanized FA mouse model, Fxnem2 ■l l "lz-' Tg(FXN)YG8Pook/800J (n= 4M/4F, “FA mouse model”) and healthy control wild type humanized mouse model Fxnem2 ■l l "lz-' Tg(FXN)Y47Pook/J (n=4M/4F, “Healthy”) were intravenously injected with one of the ssAAV9 viral vectors at a total of 1 x 1014 viral genomes (vg)/kg (0.5 x 1014 vg/kg each for the vectors containing gRNA G and gRNA U for the combination group). Mice were euthanized 30 days after infusion, and heart and liver tissues were collected, weighted and snap frozen in liquid nitrogen. The frozen heart and liver tissues were homogenized and assayed for human FXN protein by ELISA (Abeam© cat#abl76112). To assess biodistribution of the AAV9 vectors, DNA was extracted from the heart and liver tissue samples and AAV9 vector genome quantification by droplet digital (ddPCR). The heart tissue homogenate was also assessed by Succinate Dehydrogenase (SDH) in vitro enzymatic activity assay (Abeam© cat#ab228560).
[0704] In the liver of the FA mouse model, as shown in FIG. 16A (expressed as FXN protein pg per pg of loaded protein into the assay [pg/pg protein]) and FIG. 16B (normalized to the average FXN protein levels from healthy control mice), treatment with dSaCas9-2xVP64 and FXN promoter-targeting gRNAs or a combination of FXN promoter- and enhancer-targeting gRNAs significantly increased FXN
protein levels compared to treatment with dSaCas9-2xVP64 and non-targeting gRNA control.
[0705] In the heart of the FA mouse model, as shown in FIG. 17A (expressed as FXN protein pg per pg of loaded protein into the assay [pg/pg protein]) and FIG. 17B (normalized to the average FXN protein levels from healthy control mice), treatment with dSaCas9-2xVP64 and FXN promoter-targeting gRNAs or a combination of FXN promoter- and enhancer-targeting gRNAs resulted in an increase in FXN protein levels compared to treatment with dSaCas9-2xVP64 and non-targeting gRNA control.
[0706] SDH enzymatic activity was assessed in the heart homogenates of the FA mouse model, as reduced SDH activity impairs mitochondrial function and has been reported to occur in heart autopsies from FA patients. As shown in FIG. 18A (expressed as mU of SDH activity per mg loaded protein into the assay [mu/mg]) and FIG. 18B (normalized to the SDH activity from healthy control mice), treatment with dSaCas9-2xVP64 and FXN promoter-targeting gRNAs or a combination of FXN promoter- and enhancer-targeting gRNAs significantly increased SDH enzymatic activity compared treatment with dSaCas9-2xVP64 and non-targeting gRNA control, rescuing the reduced SDH enzymatic activity to near the levels observed in the healthy control mice.
[0707] Biodistribution of the administered AAV9 vectors in are shown in FIG. 19A (liver) and FIG. 19B (heart). The biodistribution patterns of the administered vectors were consistent with the known tropism of AAV9, and no difference was observed based on mouse (FA model or healthy) or encoded gRNA.
[0708] The results showed that in vivo treatment with an exemplary dCas9-transcriptional activator fusion protein and exemplary gRNAs targeting different target sites in frataxin led to an increase in expression of FXN protein in various tissues, and also rescued the SDH enzymatic activity levels in the heart, in a mouse model of FA. The results further support the utility of the exemplary dCas9- transcriptional activator fusion protein and exemplary gRNAs targeting frataxin in an in vivo treatment for FA.
Example 9: TALE-mediated transcriptional activation of FXN
[0709] Fusion proteins containing DNA-targeting domains based on transcription activator-like effector (TALE) binding domains that target the frataxin (FXN) locus are designed, generated, and assessed for their effect in activation of FXN in cells.
[0710] TALE-based DNA-targeting domains targeting regulatory elements of FXN, including promoter-targeting and enhancer-targeting TALE DNA-targeting domains, are designed, based on available methods for designing TALE targeting specific target sequences. Exemplary TALE DNA- targeting domains target sequences within the genomic coordinates human genome assembly GRCh38 (hg38) chr9:68,940, 179-69,205,519. Exemplary FXN promoter-targeting TALE DNA-targeting domains target sequences within the genomic coordinates hg38 chr9:69, 034, 622-69, 036, 670; chr9:69,034,900- 69,035,900; or chr9:69, 035, 300-69, 035, 900. Exemplary FXN promoter-targeting TALE DNA-targeting domains also target sequences within a target region spanning the genomic coordinates chr9:69,034,900-
69,035,900 from hg38 (SEQ ID NO:430); chr9:69, 035, 300-69-035, 800 from hg38; chr9:69,035,350- 69,035,450 from hg38; or chr9:69, 035, 400-69, 035, 450 from hg38. Exemplary FXN enhancer-targeting TALE DNA-targeting domains target sequences within the genomic coordinates hg38 chr9:69,027,282- 69,028,497 or chr9:69, 027, 615-69, 028, 101. Exemplary FXN enhancer-targeting TALE DNA-targeting domains also target sequences within a target region spanning the genomic coordinates chr9:69,027,282- 69,028,497 from hg38 (SEQ ID NO:431); chr9:69, 027, 615-69, 028, 101 from hg38; chr9:69,027,775- 69,027,875 from hg38; chr9:69, 027, 795-69, 027, 845 from hg38. The exemplary genomic regions specified above contain multiple sequentially tiled target sites, with TALEs targeting one of the tiled target sites in the region. Fusion proteins are designed, each comprising one of the designed TALE DNA-targeting domains fused to one or more effector domains for transcriptional activation of FXN, such as any of the effector domains described herein, such as 2 copies of the transcriptional activator VP64 (FXN-targeting TALE-2xVP64).
[0711] Viral vectors, including lentiviral vectors, are designed and cloned, each comprising nucleic acid sequences encoding a FXN-targeting TALE fusion protein (e.g. a TALE-2xVP64). Vectors further encode a selectable marker (e.g. puromycin resistance cassette). FA-iPSCs, generally described in Examples 1-4 above, are transduced with each individual FXN-targeting TALE fusion protein or combinations thereof, and enriched for transduced cells e.g. using puromycin selection). Negative control cells are transduced with nucleic acid sequences encoding a non-targeting TALE fusion protein, the FXN-targeting TALE DNA-targeting domains without an effector domain for transcriptional activation, or the selectable marker alone. Cells are harvested and analyzed for FXN mRNA expression by RT-qPCR and FXN protein expression by ELISA, in comparison to negative control cells. In some examples, expression is also compared to WT-iPSCs transduced with the selectable marker alone.
[0712] FA-iPSCs transduced with one of the FXN-targeting TALE fusion proteins are assessed for activation of frataxin expression using RT-qPCR (mRNA) and ELISA (protein).
[0713] Increased FXN expression in FA-iPSCs in comparison to negative control cells, and FXN protein expression in FA-iPSCs of at least 35% WT-iPSC levels, supports the utility of fusion proteins comprising FXN-targeting TAEE DNA-targeting domains and transcriptional activators, for targeted transcriptional activation of FXN and increased FXN protein expression, including in connection with therapies for FA.
Example 10: AAV delivery of dCas9-2xVP64 and gRNAs for increasing FXN expression in FA-iPSC-derived cardiomyocytes and neurons
[071 ] Adeno-associated virus (AAV) vectors comprising nucleotides encoding a dCas9-effector fusion protein for transcriptional activation and a gRNA targeting regulatory elements of a frataxin (FXN) locus were used to transduce differentiated cells derived from FA-iPSCs, which were assessed for increased FXN expression.
[0715] AAV delivery of a dSaCas9-2xVP64 (e.g. as set forth in SEQ ID NO:71) and FXN targeting
gRNAs was assessed for the capacity to increase FXN protein levels in FA-iPSC-derived cardiomyocytes harboring expanded GAA trinucleotide repeats (867/867). FA-iPSC-derived cardiomyocytes were transduced with AAVDJ vectors comprising nucleic acids encoding dSaCas9-2xVP64 and one of the following gRNAs: gRNA A (promoter targeting, SEQ ID NO: 1), gRNA G (promoter targeting, SEQ ID NO: 7), gRNA U (enhancer targeting, SEQ ID NO: 21), both gRNA G and gRNA U, or a non-targeting (NT) gRNA. The AAV vectors comprised an EFS promoter operably linked to the dCas fusion protein, and U6 promoter for driving expression of the gRNA. Cells were harvested on day 7 post-transduction to assess FXN protein levels by ELISA, generally as described in Example 4, and vector copy number (VCN).
[0716] As shown in FIG. 20A, AAVDJ delivery of dSaCas-2xVP64 and gRNA G into FA-derived cardiomyocytes resulted in a robust AAV dose-dependent increase in FXN protein levels in comparison to cells expressing the non-targeting (NT) gRNA. Cells expressing gRNA G at the highest AAV dosage reached approximately 80% of WT FXN protein levels whereas cells expressing the NT gRNA remained at approximately 40%. Cells expressing gRNA U or gRNA U+G also exhibited a dose dependent increase in FXN protein, and cells expressing gRNA A also exhibited increased FXN protein levels in comparison to cells expressing the non-targeting gRNA.
[0717] Next, FA-iPSC-derived neurons were transduced with an AAVDJ vector encoding dSaCas9- 2xVP64 and either gRNA G or the non-targeting (NT) gRNA at a multiplicity of infection (MOI) of IxlO6, 3xl06, IxlO7, or 3xl07. Cells were harvested on day 7 post-transduction to assess FXN mRNA expression levels by RT-qPCR (normalized to GAPDH) and analyzed as fold change in comparison to FXN mRNA expression levels in WT control cells.
[0718] As shown in FIG. 20B, FA-iPSC-derived neurons expressing dSaCas-2xVP64 and gRNA G exhibited a greater than 2-fold increase in FXN mRNA expression at 3xl07 MOI compared to FA-iPSC- derived neurons expressing the non-targeting (NT) gRNA at the lowest MOI. FA-iPSC-derived neurons expressing gRNA G did not exhibit increased FXN mRNA expression at the lower MOIs.
[0719] The transduction efficiency of the AAVDJ vector encoding dSaCas9-2xVP64 and either gRNA U or the non-targeting (NT) gRNA was investigated at various MOIs. As shown in FIG. 21A, the calculated vector copy number (VCN) for FA-iPSC-derived cardiomyocytes expressing gRNA U scaled linearly with increasing MOI. In contrast, the calculated VCN for FA-iPSC-derived neurons expressing gRNA U plateaued at the second lowest MOI tested, and indicated a lower transduction efficiency as compared to FA-derived cardiomyocytes (FIG. 21B).
[0720] Together, these results demonstrated that AAVDJ delivery of dSaCas9-2xVP64 and a FXN targeting gRNA can substantially increase frataxin expression in FA-iPSC-derived cardiomyocytes and neurons. The observed increase in frataxin protein and mRNA levels supports the therapeutic utility of AAV delivery of dCas9-effector fusions and the gRNAs for transcriptional activation of FXN in differentiated cells.
Example 11: AAV delivery of dCas9-multipartite effector fusions and gRNAs for increasing FXN expression in cardiomyocvtes and neurons
[0721] dCas9 fusion proteins comprising multipartite effectors for transcriptional activation of FXN were tested in differentiated cells derived from FA-iPSCs. FA-iPSC-derived cells were transduced with an AAV vector encoding a dCas9-multipartite effector fusion and a FXN targeting gRNA and assessed for increased FXN protein levels.
A. Identification of multipartite transcriptional effectors that facilitate increased FXN protein levels in FA-derived cardiomyocytes
[0722| Multipartite effector domains comprising three transcriptional activation domains (i.e. tripartite effectors) were designed and used to generate a series of dSaCas9-tripartite effector fusion proteins. Each dSaCas9-tripartite effector fusion included in the N- to C-terminus direction, i) the tripartite effector, containing a first transcriptional activation domain that contained one of a PYGO1 domain (e.g. SEQ ID NO:390), a FOXO3 domain (e.g. SEQ ID NO:385), an NCOA2 domain (e.g. SEQ ID NO:387), and an NCOA3 domain (e.g. SEQ ID NO:388), a second transcriptional activation domain comprising a FOXO3 domain (e.g. as set forth in SEQ ID NO: 385), a the third transcriptional activation domain comprising an NCOA3 domain (e.g. as set forth in SEQ ID NO:388), and ii) a dSaCas9 protein, generally as shown in FIG. 22. The multipartite effectors comprised domains from PYGO1, FOXO3, and NCOA3, respectively (i.e. PYGO1-FOXO3-NCOA3; e.g. as set forth in SEQ ID NO:411), NCOA3, FOXO3, and NCOA3, respectively (i.e. NCOA3-FOXO3-NCOA3; e.g. as set forth in SEQ ID NO:413), FOXO3, FOXO3, and NCOA3, respectively (i.e. FOXO3-FOXO3-NCOA3; e.g. as set forth in SEQ ID NO:415), or NCOA2, FOXO3, NCOA3, respectively (i.e. NCOA2-FOXO3-NCOA3; e.g. as set forth in SEQ ID NO:416). The individual effector domains were separated by linker sequences, sch as a GGGGS linker (e.g. SEQ ID NO: 158). Additionally, effectors composed of 1, 2, or 3 tandem copies of NCOA3 (i.e. lxNCOA3, 2xNCOA3, and 3xNCOA3, respectively) also were tested in fusion proteins.
[0723] FA-iPSC-derived cardiomyocytes were transduced with an AAVDJ vector encoding gRNA G (SEQ ID NO:53, targeting SEQ ID NO:7) or a non-targeting (NT) guide RNA and one of the dSaCas9- tripartite effector fusions or dSaCas9 fusions with 1, 2, or 3 tandem copies of NCOA3, at a MOI of 3xl04 or 3xl05. The effect of dCas9-multipartite effectors on FXN protein levels was compared to a reference dSaCas9-2xVP64 fusion protein in the presence of the same gRNA. Wild- type-derived (WT) and FA- iPSC-derived cardiomyocyte cells not transduced with the full DNA-targeting system were also included as positive and negative controls, respectively. Cells were harvested on day 7 post-transduction to assess FXN protein levels by ELISA, generally as described in Example 4, and vector copy number (VCN).
[0724] As shown in FIG. 23, there was a robust increase in FXN protein levels at both MOIs in FA- iPSC-derived cardiomyocytes expressing the exemplary FXN promoter-targeting gRNA (gRNA G) and several of the dSaCas9-tripartite effector fusions, including for NCOA3-FOXO3-NCOA3, NCOA2- FOXO3-NCOA3, and FOXO3-FOXO3-NCOA3 tripartite effectors. These tripartite effector fusions
outperformed dSaCas9-2xVP64 in both MOI conditions, with the NCOA3-FOXO3-NCOA3 tripartite effector fusion exhibiting the strongest increase in FXN protein levels.
[0725] At an MOI of 3xlOA4, FA-iPSC-derived cardiomyocytes expressing guide RNA G and the NCOA3-FOXO3-NCOA3-daSaCas9 tripartite effector fusion increased FXN protein levels to approximately 60% of WT FXN protein levels whereas FA-cardiomyocytes expressing guide RNA G and dSaCas9-2xVP64 increased FXN protein levels to approximately 30% of WT FXN protein levels. A combination of guide RNA G and the NCOA2-FOXO3-NCOA3-daSaCas9 tripartite effector fusion increased FX protein levels to approximately 50% of WT FXN protein levels and a combination of guide RNA G and the FOXO3-FOXO3-NCOA3-daSaCas9 tripartite effector fusion increased FXN protein levels to approximately 40% of WT FXN protein levels.
[07263 At 3xl0A5 MOI, the NCOA3-FOXO3-NCOA3-daSaCas9 tripartite effector fusion further increased FXN protein levels to almost 90% of WT when paired with guide RNA G. The NCOA2- FOXO3-NCOA3-daSaCas9 tripartite effector fusion increased FXN protein levels to approximately 70% of WT, and the FOXO3-FOXO3-NCOA3-daSaCas9 tripartite effector fusion increased FXN protein levels to approximately 65% of WT. In contrast, dSaCas9-2xVP64 increased FXN protein levels to 50% of WT FXN protein levels under similar conditions.
[0727] dSaCas9 effector fusions with activation domains composed solely of 1, 2, or 3 copies of NCOA3 did not substantially increase FXN protein levels in FA-iPSC-derived cardiomyocytes expressing guide RNA G at 3xlOA4 MOI. However, at 3xl0A5 MOI, dSaCas9 effector fusions with activation domains comprising 2 or 3 copies of NCOA3 increased FXN protein levels to 40% and 50% of WT in FA-iPSC-derived cardiomyocytes expressing guide RNA G.
B. The tripartite NCOA3 effector increased FXN levels more effectively than 2xVP64.
[0728] The VCN was calculated for FA-iPSC-derived cardiomyocytes expressing gRNA G and either the NCOA3-FOXO3-NCOA3-daSaCas9 or dSaCas9-2xVP64 effector fusion at various MOIs. As shown in FIG. 24A, NCOA3-FOXO3-NCOA3-daSaCas9 exhibited a higher VCN than dSaCas9- 2xVP64 at both MOIs tested. As shown in FIG. 24B, NCOA3-FOXO3-NCOA3 still outperformed 2xVP64 at similar VCNs. Furthermore, NCOA3-FOXO3-NCOA3-daSaCas9 reached a similar VCN as dSaCas9-2xVP64 at a 10-fold lower MOI (FIG. 24C). The numeric values for the data represented in FIG. 24A and FIG. 24B are shown in FIG. 24C and Table E4.
Table E4. Transduction efficiency and resulting FXN expression in FA-iPSC-derived cardiomyocytes transduced with gRNA G and NCOA3-FOXO3-NCOA3-daSaCas9 or dSaCas9- 2xVP64
[0729] Further, the NCOA3-FOXO3-NCOA3-daSaCas9 tripartite fusion outperformed other tripartite effector fusions and dSaCas9-2xVP64 when adjusted for VCN (FIG. 25).
C. AAV delivery of dSaCas9-tripartite effector fusions increases FXN protein and mRNA levels in FA-derived cells
[0730] Next, AAV delivery of the dSaCas9-tripartite effector fusions was assessed for the capacity to increase FXN protein levels in FA-iPSC-derived cardiomyocytes harboring expanded GAA trinucleotide repeats (867/867). FA-iPSC-derived cardiomyocytes were transduced with AAVDJ vectors encoding gRNA G or a non-targeting gRNA (NT) and one of the following dSaCas9-tripartite effector fusions: FOXO3-FOXO3-NCOA3-dSaCas9, NCOA2-FOXO3-NCOA3-dSaCas9, or NCOA3-FOXO3- NCOA3-dSaCas9. dSaCas9-2xVP64 was included as a control and a GFP control construct was used to transduce healthy (CTR GFP) and FA-derived cardiomyocytes (GFP). Cells were harvested on day 7 post-transduction to assess FXN protein levels by ELISA, generally as described in Example 4.
[0731] As shown in FIG. 26A, all dSaCas9 fusions with all three of the tripartite effectors increased FXN protein levels to a greater extent than dSaCas9-2xVP64 in FA-iPSC-derived cardiomyocytes expressing gRNA G. Expression of gRNA G and NCOA3-FOXO3-NCOA3-dSaCas9 increased FXN protein levels to approximately 80% of WT levels in FA-iPSC-derived cardiomyocytes. In comparison, the dSaCas9-2xVP64 increased FXN protein levels to approximately 50% of WT, while the negative GFP-transduced and non-targeting (NT) gRNA controls all remained at approximately 25% of WT FXN levels.
[0732] AAV delivery of the tripartite effector fusions was assessed for the capacity to increase FXN expression in FA-iPSC-derived neurons. FA-iPSC-derived neurons were transduced with AAVDJ vectors encoding either gRNA G or the non-targeting (NT) gRNA and FOXO3-FOXO3-NCOA3- dSaCas9, NCOA3-FOXO3-NCOA3-dSaCas9 or dSaCas9-2xVP64. Healthy (CTR GFP) and FA-derived (GFP) neurons were transduced with a GFP construct as positive and negative controls. Cells were harvested on day 7 post-transduction to assess FXN mRNA expression levels by RT-qPCR (normalized to GAPDH) and analyzed as fold change in comparison to FXN mRNA expression levels in WT control cells. As shown in FIG. 26B, each of the effector fusion proteins increased FXN mRNA levels in cells expressing gRNA G, in comparison to the NT gRNA control. The FOXO3-FOXO3-NCOA3-dSaCas9 and NCOA3-FOXO3-NCOA3-dSaCas9 tripartite effector fusions increased FXN mRNA levels to a similar or greater extent as dSaCas9-2xVP64 in FA-iPSC-derived neurons expressing gRNA G.
[0733] In summary, these results demonstrated that the tripartite effectors described herein function as potent activators of FXN expression when targeted to a FXN locus, and with activity similar to or exceeding that of 2xVP64. Further, dSaCas9-tripartite effector fusions paired with a FXN targeting gRNA were shown to increase both FXN protein and mRNA levels in FA-iPSC-derived cardiomyocytes
and neurons. The tripartite effectors were derived from human proteins, in comparison to the virus- derived VP64 effectors. The results support the therapeutic utility of the tripartite effectors for targeted activation of FXN expression, including in fusion proteins capable of targeting or being targeted to a FXN locus.
Example 12; Size minimization of dCas9-gRNA construct
[0734] AAV vectors for expression of dSaCas9 and FXN-targeting gRNAs were designed, cloned and tested. Specific elements in a construct for AAV-mediated delivery of a dCas9-effector fusion for transcriptional activation and gRNA were replaced or removed to decrease construct size while maintaining ability to increase FXN expression in FA-iPSC-derived differentiated cells. Constructs were transduced into FA-iPSC-derived cardiomyocytes and assessed for ability to FXN and dCas9 mRNA levels compared to the original construct.
A. mini-U6 promoter for gRNA expression
[0735] Constructs for AAV-mediated delivery of a dCas9-effector fusion and gRNA were generated. The constructs comprised a single polynucleotide containing a first nucleic acid comprising a human elongation factor alpha short (EFS) promoter (SEQ ID NO:436) operably linked to a sequence encoding dSaCas9-2xVP64, and a second nucleic acid sequence comprising a U6 promoter (SEQ ID NO:432) or mini-U6 promoter (SEQ ID NO:433) operably linked to an exemplary FXN-targeting gRNA (gRNA A) or non-targeting gRNA (NT). The first and second nucleic acid sequences were separated by an SpA element.
[0736] Constructs comprising the 249 bp U6 promoter (SEQ ID NO:432) driving expression of gRNA A was replaced with a 112 bp mini-U6 promoter (SEQ ID NO:433) in a dCas9-gRNA vector. Cardiomyocytes derived from either FA iPSCs or genetically corrected (GC) FA iPSCs were transduced with AAVDJ vectors encoding dSaCas9-2xVP64 and either the U6 or mini-U6 promoter driving expression of gRNA A or a non-targeting (NT) gRNA. Cells were harvested on day 7 post-transduction to assess FXN mRNA expression levels by RT-qPCR (normalized to GAPDH). mRNA expression levels were normalized to control FA-cardiomyocytes expressing a GFP construct for analysis.
[0737] As shown in FIG. 27, FA-cardiomyocytes expressing gRNA A driven by the mini-U6 promoter increased FXN mRNA levels to a similar extent as the original U6 promoter. In genetically corrected (GC) FA-cardiomyocytes, there were no major differences in FXN mRNA expression across cell lines regardless of the promoter-gRNA combination expressed.
[0738] These data indicate that replacing the U6 promoter with the mini-U6 promoter decreases the size of the construct for AAV-mediated delivery, while maintaining the ability to activate FXN expression. The data also indicate the utility of AAV vectors comprising the described nucleic acids.
B. dSaCas9-2xVP64 increased FXN mRNA without a FLAG epitope tag
[0739] AAV vectors for delivering gRNA and dCas9-2xVP64 with or without a FLAG epitope tag
(amino acid sequence set forth in SEQ ID NO: 129; an exemplary nucleotide sequence encoding a FLAG tag set forth in SEQ ID NO: 128) were assessed for ability to increase FXN expression levels.
Cardiomyocytes derived from FA iPSCs were transduced with AAVDJ vectors encoding gRNA A or non-targeting (NT) gRNA and dSaCas9-2xVP64 with or without a FEAG tag. As shown in FIG. 28, FA- iPSC-derived cardiomyocytes exhibited increased FXN levels following delivery of the dSaCas9- 2xVP64 fusion protein with gRNA A in comparison to the non-targeting gRNA, both with and without the FEAG tag.
[0740] Together, these data demonstrate the therapeutic utility of constructs for AAV-mediated delivery of dCas9 and gRNA, including with the EFS promoter and mini-U6 promoter, and exclusion of a FEAG epitope tag, in order to minimize size of the construct while maintaining the ability to increase FXN expression. Given the size restrictions of many AAV vectors used in gene therapy, smaller dSaCas9-gRNA constructs can expand the range of vectors that can be employed for therapeutic delivery to patient cells.
Example 13: Fusion proteins containing engineered zinc finger proteins and VP64 transcriptional activation domains for targeted activation of FXN expression
[0741] Engineered zinc finger proteins (eZFPs) were designed to bind enhancer and promoter elements of FXN. Fusion proteins comprising the eZFPs and VP64 (eZFP-VP64 fusion proteins) were transduced into cells derived from FA patients and assessed for the capacity to increase FXN expression.
[0742] eZFPs were designed, targeting FXN promoter and enhancer regions, as described in Section I and II. eZFPs included eZFPs targeting the target sites set forth in SEQ ID NOS:269-300, and included the eZFPs set forth in Table 2A. eZFP-VP64 fusion proteins comprising the eZFPs were designed (e.g. as set forth in SEQ ID NOS:320-326), and mRNA encoding the fusion proteins was generated and electroporated into FA-iPSCs harboring expanded GAA trinucleotide repeats (867/867). FA-iPSCs expressing a eZFP-VP64 fusion protein were assessed by qPCR for FXN mRNA levels after 48 or 72 hours. FXN mRNA expression levels were normalized to FA-iPSCs expressing dSaCas9-VP64 without a guide RNA (dCas9 only). Several of the eZFP-VP64 fusion proteins led to increased FXN mRNA levels in FA-iPSCs after 48 hours (FIG. 29). Seven eZFP fusion proteins were selected for further analysis.
[0743] As shown in FIG. 30A, several eZFP-VP64 constructs (including eZFP_A04, eZFP_A09, eZFP_A12, eZFP_A13, eZFP_A15, eZFP_A22, and eZFP_A31) led to increased FXN mRNA levels in FA-iPSCs after 48 hours. At 72 hours post-electroporation, cells with four of the seven eZFP-VP64 fusion proteins (eZFP_A04, eZFP_A09, eZFP_A12, and eZFP_A31) still exhibited an increase in FXN mRNA levels (FIG. 30B).
[0744] FXN protein levels were also assessed by EEISA after 72 hours. As shown in FIG. 30C, FA-iPSCs expressing several of the eZFP-VP64 fusion proteins exhibited increased FXN protein levels. Notably, VP64 fusions with eZFP_A04, eZFP_A09, eZFP_A22 and eZFP_A31 increased FXN protein levels to a similar or greater extent as dSaCas9-2xVP64 and gRNA G. Further, eZFP_A04 and
eZFP_A09 bound sequences in close proximity to the enhancer target site for gRNA U, while the binding site for eZFP_A31 overlapped with the promoter target site for gRNA G (FIG. 31).
[0745] Next, AAVDJ vectors encoding the eZFP-VP64 fusion proteins were transduced into FA- iPSC-derived cardiomyocytes at multiple MOIs. Cells were harvested on day 7 post-transduction to assess FXN mRNA expression levels by RT-qPCR (normalized to GAPDH). FXN expression was normalized to FXN mRNA expression levels in FA-iPSC-derived cardiomyocytes expressing a GFP control construct. As shown in FIG. 32, eZFP_A31 dramatically increased FXN mRNA expression in FA-iPSC-derived cardiomyocytes. Notably, expression of eZFP_A31-VP64 (SEQ ID NO:326) resulted in a greater than 2-fold increase in FXN mRNA, and increased FXN expression to a greater extent than dSaCas9-2xVP64+gRNA G.
[07463 To assess expression levels of the eZFP-VP64 fusion proteins themselves, VP64 mRNA expression in FA-iPSC-derived cardiomyocytes was also assessed by qPCR. As shown in FIG. 33, eZFP_A04 and eZFP_A09 VP64 fusion proteins did not increase FXN mRNA levels but were expressed to a similar extent as the eZFP_A31 VP64 fusion protein. The eZFP_A31 VP64 fusion protein also increased FXN expression more than dSaCas9-2xVP64+gRNA G at multiple MOIs despite similar expression levels of VP64 and Cas9 in FA-cardiomyocytes (FIG. 34).
[0747] Next, FA-iPSC-derived neurons were transduced at IxlO6 MOI with AAVDJ vectors encoding the eZFP-VP64 fusion proteins. Cells were harvested on day 7 post-transduction to assess FXN mRNA expression levels by RT-qPCR (normalized to GAPDH). FXN mRNA expression was normalized to FXN mRNA expression levels for negative control FA-iPSC-derived neurons not transduced with an eZFP-VP64. As shown in FIG. 35, only the eZFP_A31 VP64 fusion protein increased FXN mRNA expression in FA-neurons. dSaCas9-2xVP64+gRNA G had no effect on FXN mRNA levels in FA-iPSC-derived neurons at the same MOI (MOI=lxl06). VP64 mRNA levels were also assessed and confirmed that the eZFP_A31 VP64 fusion protein was expressed at similar levels in FA-iPSC-derived neurons as fusion proteins with several of the other eZFPs that had no impact on FXN mRNA expression (FIG. 36).
[0748] Together, these results demonstrate robust activation of FXN expression in FA-iPSCs. In addition, eZFP_31-VP64 (SEQ ID NO:326, comprising the eZFP_A31 set forth in SEQ ID NO:307) increased FXN mRNA and protein levels in both FA-iPSCs and more than one cell type differentiated from the FA-iPSCs (neurons and cardiomyocytes), including to a similar or greater extent than a dCas9- based DNA-targeting system for transcriptional activation of FXN. Of note is that the target sequence for eZFP_A31 overlaps with the target sequence of gRNA G, indicating that this regulatory region was identified as particularly responsive to the engineered transcriptional activators. In some aspects, an eZFP-based transcriptional activator (i.e. eZFP fusion protein) is more compact than a dCas9-based activator (i.e. dCas fusion protein), does not require a cognate gRNA for DNA targeting, and is derived from human proteins. These features provide technical advantages for packaging a eZFP transcriptional activator into size constrained AAV vectors for gene therapy. In general, these data support the
therapeutic utility of the eZFPs, including in fusion proteins for targeted activation of FXN expression.
Example 14: Fusion proteins containing eZFPs and tripartite effector domains for targeted activation of FXN expression
[0749] eZFP fusion proteins comprising tripartite effectors for transcriptional activation and an eZFP targeting a FXN promoter or enhancer region were designed and tested for transcriptional activation of FXN. FA-derived cells were transduced with an AAV vector encoding the eZFP-tripartite effector fusion proteins and assessed for FXN expression.
[0750] eZFP fusion proteins comprising an exemplary FXN-targeting eZFP (eZFP_A31; SEQ ID NO:307), and tripartite effectors were designed. The tripartite effectors of the eZFP fusion proteins comprised, from N-terminus to C-terminus: FOXO3, FOXO3, and NCOA3 (i.e. FOXO3-FOXO3- NCOA3; SEQ ID NO:415); NCOA3, FOXO3, and FOXO3 (i.e. NCOA3-FOXO3-FOXO3; SEQ ID NO:418); or NCOA3, FOXO3, and NCOA3 (i.e. NCOA3-FOXO3-NCOA3; SEQ ID NO:413).
[0751] FA-derived cardiomyocytes and neurons were transduced with AAVDJ vectors encoding eZFP_A31 fused to VP64 or the tripartite effectors, or dSaCas9 fused to VP64 (dSaCas9-2xVP64) or the tripartite effectors and gRNA G. Non-targeting (NT) gRNA and a GFP construct were included as negative controls for FA-derived cells. Cells were harvested on day 7 post-transduction to assess FXN mRNA expression levels by RT-qPCR (normalized to GAPDH) compared to FXN mRNA levels in WT cells.
[0752] As shown in FIG. 37, all tested fusion proteins with eZFP_A31 increased FXN mRNA levels in FA-iPSC-derived cardiomyocytes. eZFP_A31 fused with NCOA3-FOXO3-NCOA3 (i.e. eZFP_A31-NCOA3-FOXO3-NCOA3; e.g. SEQ ID NO:333) increased FXN mRNA to approximately 80% of WT levels, eZFP_A31 fused with FOXO3-FOXO3-NCOA3 (i.e. eZFP_A31-FOXO3-FOXO3- NCOA3; e.g. SEQ ID NO:340) increased FXN mRNA to approximately 60% of WT, and eZFP_A31 fused with VP64 (i.e. eZFP_A31-VP64; e.g. SEQ ID NO:326) increased FXN mRNA to approximately 50% of WT. In contrast, the three dSaCas9-effector fusions paired with gRNA G each increased FXN mRNA to approximately 50% of WT in FA-iPSC-derived cardiomyocytes.
[0753] As shown in FIG. 38, all three of the eZFP effector fusions also increased FXN mRNA levels in FA-iPSC-derived neurons. Notably, FA-neurons expressing the eZFP_A31 effector fusion proteins exhibited 100% or more of WT FXN mRNA levels. The dSaCas9-effector fusions paired with gRNA G all increased FXN mRNA levels to approximately 60% of WT in FA-iPSC-derived neurons.
[0754] Various formats of a FXN-targeting eZFP tripartite effector fusion were generated, with different tripartite effectors fused at the C-terminus or the N-terminus of eZFP_A31, including the following effectors: NCOA3-FOXO3-FOXO3 (e.g. as set forth in SEQ ID NO:418), FOXO3-FOXO3- NCOA3 (e.g. as set forth in SEQ ID NO:415), and NCOA3-FOXO3-NCOA3 (e.g. as set forth in SEQ ID NO:413).
[0755] As shown in FIG. 39, all eZFP tripartite effector fusion protein formats with the exemplary
eZFP_A31 resulted in increased FXN mRNA in FA-neurons. C-terminal fusions of the tripartite effectors led to a stronger increase of FXN mRNA than N-terminal fusions. Further, the C-terminal tripartite fusions increased FXN expression more than eZFP_A31-VP64 and fully restored FXN mRNA to WT levels in FA-iPSC-derived neurons.
[0756] In summary, these results demonstrated that eZFP-tripartite effector fusions targeted to FXN are potent transcriptional activators and can increase FXN expression levels to a similar or greater extent than dCas9-based transcriptional activation in FA-iPSC-derived cells. The eZFP tripartite effector fusion proteins with the exemplary eZFP eZFP_A31 also increased FXN mRNA levels to greater or similar levels as eZFP VP64 fusion proteins. Notably, all eZFP effector fusion proteins restored FXN mRNA to 100% or more of WT levels in FA-iPSC-derived neurons. The fusion proteins containing the eZFP and tripartite effectors form a compact and human-derived fusion protein well-suited to AAV delivery, including for increasing FXN expression and/or for treatment of Friedreich ataxia (FA). The results generally support the therapeutic utility of the eZFPs and eZFP fusion proteins provided herein for targeted activation of FXN expression, including for Friedreich’s ataxia.
Example 15; Validation of tripartite transcriptional effector in vivo in a Friedreich’s ataxia (FA) mouse model
[0757] The humanized mouse model for Friedreich’s ataxia (FA) previously described in Example 8 was treated with dCas9 and eZFP_A31 fusion proteins containing either the tripartite effector NCOA3- FOXO3-NCOA3 (NFN; SEQ ID NO:413) or transcriptional activator VP64 and assessed for in vivo transcriptional activation of frataxin (FXN).
[0758] Adeno-associated virus (AAV) vectors were generated to evaluate the in vivo effect of dSaCas9 and eZFP_A31 when fused to VP64 or the tripartite effector NFN, as described in Example 10, 11, 13, and 14. Each single-stranded AAV9 (ssAAV9) vector included nucleic acid sequences encoding one of the fusion proteins described above. For fusion proteins that contained dSaCas9, the vector further encoded either a FXN promoter-targeting gRNA G (SEQ ID NO:53) or a non-targeting gRNA (NT gRNA; SEQ ID NO: 601) as a control.
[0759] For expression in heart or liver, 6-8 week old male and female humanized FA mouse model, Fxnem2 1Lutzy Tg(FXN)YG8Pook/800J (n= 4M/4F, “FA model”) and healthy control wild type humanized mouse model Fxnem2 ■l l "lz-' Tg(FXN)Y47Pook/J (n=4M/4F, “WT”) were intravenously injected with one of the ssAAV9 viral vectors at a total of 1 x 1014 viral genomes (vg)/kg, or a vehicle as a control. For expression in the cerebellum, mouse models were delivered via an intracerebellar route with one of the ssAAV9 viral vectors at a total of 1 x 1010 viral genomes (vg), or a vehicle as a control.
[0760] Mice were euthanized 30 days after infusion, and heart, liver, and cerebellum tissues were collected, weighted, and snap frozen in liquid nitrogen. The frozen heart, liver, and cerebellum tissues were homogenized and assayed for human FXN protein by ELISA (Abeam© cat#abl76112). Human
FXN mRNA expression levels were assessed by RT-qPCR (normalized to mouse TBP mRNA) in comparison to human FXN mRNA expression levels in the WT healthy mouse model. To assess biodistribution of the AAV9 vectors, DNA was extracted from the heart, liver, and cerebellum tissue samples and AAV9 vector genome quantified by droplet digital (ddPCR).
[0761] In the liver of the FA mouse model, as shown in FIG. 40A (expressed as FXN protein pg per pg of loaded protein into the assay [pg/pg protein] normalized to the average FXN protein levels from WT healthy control mice) and FIG. 40B (expressed as the fold-change of FXN mRNA levels normalized to average FXN mRNA levels from WT healthy control mice), treatment with eZFP fusion proteins, including those fused to VP64 or the tripartite effector NFN, increased FXN protein and mRNA levels compared to a vehicle control in either the WT or FA mouse models or treatment with dCas9 fusion proteins with gRNA G or NT gRNA.
[0762] In the heart of the FA mouse model, as shown in FIG. 41A (expressed as FXN protein pg per pg of loaded protein into the assay [pg/pg protein] normalized to the average FXN protein levels from WT healthy control mice) and FIG. 41B (expressed as the fold-change of FXN mRNA levels normalized to average FXN mRNA levels from WT healthy control mice), treatment with both eZFP fusion proteins resulted in an increase in FXN mRNA levels compared to a vehicle control in either the WT or FA mouse models or treatment with dCas9 fusion proteins with gRNA G and NT gRNA. Notably, treatment with the eZFP fusion protein containing the tripartite effector NFN resulted in the highest increase in FXN mRNA levels among the experimental conditions.
[0763] In the cerebellum of the FA mouse model, only dCas9 fusion proteins containing either the transcriptional activator VP64 or the tripartite effector NFN were assessed. As shown in FIG. 42A (expressed as FXN protein pg per pg of loaded protein into the assay [pg/pg protein] normalized to the average FXN protein levels from WT healthy control mice) and FIG. 42B (expressed as the fold-change of FXN mRNA levels normalized to average FXN mRNA levels from WT healthy control mice), treatment with dCas9-VP64 and gRNA G resulted in an increase in FXN protein and mRNA levels as compared to use of a NT gRNA or a vehicle control in either WT or FA mouse models.
[0764] Follow-up experiments were also performed to show FXN modulation relative to the wildtype healthy control in each tissue type using the exemplary eZFP fusion protein containing eZFP_A31 and the tripartite effector NFN (eZFP_A31-NFN) and validate the construct localized in the correct tissue following infusion. In each experiment, FXN expression levels were measured in a WT healthy control mouse following infusion with a vehicle only (WT vehicle) as a positive control and in a FA model mouse following infusion with a vehicle only (FA model vehicle) as a negative control.
[0765] As shown in FIG. 43A, treatment with eZFP_A31-NFN increased FXN protein levels compared to a saline control in both the heart (left panel) and liver (middle panel) of the FA mouse model. Successful localization of each eZFP_A31-NFN construct in either the heart or liver following intravenous infusion was validated by measuring transgene expression levels of the tripartite effector NFN in either the heart or liver, which were normalized to mouse TBP mRNA levels in the heart (right
panel). Heart samples following infusion showed comparable NFN and TBP expression levels, indicating eZFP_A31-NFN was successfully delivered to the heart. In contrast, liver samples following infusion showed an overexpression of NFN as compared to heart TBP, indicating that eZFP_A31-NFN was successfully delivered to the liver.
[0766] As shown in FIG. 43B, treatment with eZFP_A31-NFN increased FXN protein levels compared to a saline control in the cerebellum (left panel). Successful localization of each eZFP_A31- NFN fusion protein construct in the cerebellum following intracerebellar infusion was measured using the same analysis method previously described for FIG. 43A. Cerebellum samples following infusion showed an overexpression of NFN as compared to heart TBP, indicating that eZFP_A31-NFN was successfully delivered to the cerebellum.
[0767| In summary, these results showed that in vivo treatment with eZFP-tripartite effector fusions targeted to FXN are potent transcriptional activators and can increase FXN expression levels to a similar or greater extent than dCas9-based transcriptional activation in various tissues. The results further support the therapeutic utility of eZFP and tripartite transcriptional effector fusion proteins provided herein for targeting frataxin in an in vivo treatment for FA.
Example 16: Optimization of ZFP-tripartite effector construct
[0768] AAV vectors for expression of the exemplary eZFP fusion protein containing eZFP_A31 and tripartite effector NCOA3-FOXO3-NCOA3 (eZFP_A31-NFN) were designed, cloned, and tested. Specific elements in a construct for A AV-mediated delivery of an eZFP fusion protein for transcriptional activation were replaced to decrease immunogenicity and increase transgene expression. Constructs were then transduced into Friedreich’ s ataxia (FA)-iPSC-derived cardiomyocytes and assessed for the ability to increase frataxin (FXN) mRNA levels compared to the original construct.
[0769] The original construct for AAV-mediated delivery of eZFP_A31-NFN was a single polynucleotide containing, in sequential order: (a) human elongation factor alpha short (EFS) promoter nucleotide sequence (SEQ ID NO:436), (b) a nucleic acid sequence encoding an SV40 NLS (amino acid sequence set forth in SEQ ID NO: 159), (c) a nucleic acid sequence encoding eZFP_A31 (amino acid sequence set forth in SEQ ID NO:307), (d) a nucleic acid sequence encoding SV40 NLS (amino acid sequence set forth in SEQ ID NO: 159), (e) a nucleic acid sequence encoding NFN (amino acid sequence set forth in SEQ ID NO:413), and (f) a SpA polyA nucleotide sequence (SEQ ID NO:437).
[0770] To decrease immunogenicity of the NLS, a new construct was developed that replaced each SV40 NLS sequence with a different NLS sequence c-myc (amino acid sequence set forth in SEQ ID NO: 160; an exemplary nucleotide sequence set forth in SEQ ID NO:606), as exemplified in FIG. 44A. To increase transgene expression, two new constructs were developed using either the original construct or the construct designed to decrease immunogenicity as the starting construct by: (1) substituting the EFS promoter for either a CAG promoter (exemplary nucleotide sequence set forth in SEQ ID NO: 602) or EFla promoter (exemplary nucleotide sequence set forth in SEQ ID NO:603), (2) substituting the SpA
polyA sequence for a bGH polyA sequence (exemplary nucleotide sequence set forth in SEQ ID NO:604), and (3) adding a 5’ UTR nucleotide sequence (SEQ ID NO:605) between the promoter and the NLS sequence closest to the N-terminal. The two additional constructs designed to increase transgene expression are schematically represented in FIG. 44B.
[0771] Cardiomyocytes derived from either FA iPSCs were transduced with AAVDJ vectors encoding eZFP_A31-NFN in any of the four constructs previously described at different dosages. AAVDJs vectors encoding the original construct but substituting NFN for the transcriptional activator VP64 were also used as a control. Cells were harvested on day 7 post-transduction to assess FXN mRNA expression levels by RT-qPCR (normalized to human TBP mRNA).
[0772] As shown in FIG. 45, treatment of FA-cardiomyocytes with any vector encoding ZFP31- NFN increased FXN mRNA levels in a dose-dependent manner. However, different AAV elements or production methods led to different increases in FXN mRNA levels. The use of a EFla or a CAG promoter led to higher increases in FXN mRNA levels as compared to the use of an EFS promoter.
[0773] The results indicate that using larger promoters, such as EFla and CAG promoters compared to an EFS promoter, improved FXN modulation. Overall, use of the CAG promoter improved FXN modulation by 30x, as compared to the original construct. These data demonstrate the therapeutic utility of A AV-mediated delivery of eZFP-tripartite transcriptional effector fusion proteins, including the optimization of AAV constructs to increase FXN modulation.
Example 17; Screening of additional ZFPs for targeted activation of FXN expression
[0774] Based on the robust activation of FXN expression using fusion proteins containing eZFP_A31 as set forth in SEQ ID NO:307 (see Example 13), additional engineered zinc finger proteins (eZFPs) were designed to target sites in the FXN promoter region proximal to the target site of eZFP_A31 (SEQ ID NO:299). Fusion proteins containing the additional eZFPs and transcriptional activation domains were initially screened in HEK293 cells, then top hits were screened in Friedreich’s ataxia (FA) patient fibroblasts and FA-iPSC-derived cardiomyocytes.
A. Identification of additional ZFPs that facilitate increased FXN modulation in HEK293
[0775] eZFPs were designed, including eZFPs targeting the target sites set forth in SEQ ID NOS:583-600, and included the eZFPs set forth in Table 2B.
[0776] eZFP-VP64 and eZFP-NFN fusion proteins containing additionally designed eZFPs, including the exemplary eZFPs set forth in Table 2B, were designed, and plasmids encoding the fusion proteins were generated and transfected into HEK293 cells seeded at 20,000 cells per well of a 96-well plate one day prior. As controls, exemplary eZFP fusion protein containing an eZFP_A31 fused with the tripartite effector NCOA3-FOXO3-NCOA3 (eZFP_A31-NFN) or transcriptional activator VP64 (eZFP_A31-VP64), or dCas9-NFN fusion proteins with either gRNA G (gG) or a non-targeting gRNA (gNT) were also designed, generated in plasmids, and tested. Additionally, some of the screens also
measured FXN levels following either no treatment (untreated), transfection with GFP, treatment with PBS, and transfection with either an empty vector or lipid only as additional negative controls. HEK293 cells were harvested three days post-transfection and FXN mRNA levels were assessed by qPCR. FXN mRNA expression levels were normalized expression of TBP and presented as fold-change relative to HEK293 cells transfected with a plasmid encoding GFP.
[0777] As shown in FIGS. 46-48, several of the eZFP-VP64 and eZFP-NFN fusion proteins led to increased FXN mRNA levels in HEK293 cells after 72 hours. Some eZFPs appeared to result in higher FXN mRNA levels as compared to eZFP_A31 , which had previously been the best-performing eZFP tested (see Example 14). Several eZFPs from the eZFP fusion proteins indicated with an asterisk (“*”) were selected for further validation when fused with the tripartite effector NFN or in other cell types.
B. Validation of top eZFPs that facilitate increased FXN modulation
[0778] Exemplary eZFP hits identified in Example 17.A were further validated for their ability to modulate FXN when fused to the tripartite effector NCOA3-FOXO3-NCOA3 (NFN) and/or in FA patient fibroblasts.
[0779] Exemplary eZFPs from the eZFP-VP64 fusion proteins indicated by an asterisk in FIG. 46 were selected to be used in designing new fusion proteins containing one of the exemplary eZFPs and the tripartite effector NFN. Plasmids encoding the eZFP-NFN fusion proteins were generated and transfected into HEK293 cells seeded at 20,000 cells per well of a 96-well plate one day prior. As controls, transfection with dCas9-NFN fusion proteins with either FXN promoter-targeting gRNA G (gG) or a non-targeting gRNA (gNT), transfection with GFP, or transfection with a vector only were also tested. HEK293 cells were harvested three days post-transfection and FXN mRNA levels were assessed by qPCR. FXN mRNA expression levels were normalized to HEK293 cells transfected with a plasmid encoding GFP. As shown in FIG. 49, all tested constructs led to an increase in FXN mRNA levels. Several of the tested eZFPs, including eZFP_A40, eZFP_A41, eZFP_A45, and eZFP_A46, outperformed the previous best-performing exemplary eZFP eZFP_A31 in increasing FXN mRNA levels.
[0780] The eZFP-NFN fusion proteins previously tested in FIG. 49 and the highest-performing eZFP-NFN fusion proteins in FIGS. 47 and 48 (as indicated by an asterisk) were also further validated in FA patient fibroblasts. mRNA encoding fusion proteins containing each of the selected eZFP-NFN fusion proteins were generated and transfected into FA fibroblasts. As controls, transfections of vector only, GFP, or eZFP_A31 fusion proteins containing either the tripartite effector NFN or the transcriptional activator VP64 were used. Cells were collected on Day 2 post-transfection and FXN mRNA levels were assessed by qRT-PCR and presented as fold-change relative to FA fibroblasts transfected with GFP. As shown in FIGS. 50 and 51, several eZFP-NFN fusion proteins resulted in higher FXN mRNA levels. Notably, eZFP_A48 and eZFP_A46 fusion proteins led to the highest FXN mRNA increases, outperforming exemplary constructs containing eZFP_A31 (eZFP_A31-NFN and eZFP_A31-VP64) that were the best-performing eZFP previously tested in Example 14.
[0781] These results demonstrated that eZFP fusion proteins targeted to FXN are potent transcriptional activators in FA patient fibroblasts cells. The exemplary eZFPs eZFP_A48 and eZFP_A46 also increased FXN mRNA levels to greater or similar levels as eZFP_A31. Of note, the most effective eZFPs appeared to cluster near the regulatory region proximal to the target site of eZFP_A31 (FIG. 52). The results generally support the therapeutic utility of the eZFPs and eZFP fusion proteins provided herein for targeted activation of FXN expression, including for Friedreich’s ataxia.
C. Validation of top ZFPs that facilitate increased FXN modulation using optimized AAV constructs
[0782] Optimized AAV vectors (see Example 16) for expression of fusion proteins containing exemplary eZFPs that performed well in Example 17. A and/or 17.B, such as eZFP_A31, eZFP_A48, eZFP_A44, and eZFP_A46 were used to assess FXN modulation in FA-iPSC-derived cardiomyocytes. Each exemplary fusion protein was expressed using a CAG promoter and a c-myc NLS in the optimized AAV construct described in Example 16.
[0783] Cardiomyocytes derived from either FA iPSCs or genetically corrected (GC) FA iPSCs were transduced with AAVDJ vectors encoding the fusion proteins previously described at different dosages. As negative controls, FA iPSCs were either not transduced or transduced with AAVDJ vectors encoding (a) a fusion protein containing a non-targeting eZFP (eZFP-NT; SEQ ID NO: 607) and the tripartite effector NFN (eZFP-NT-NFN) using the optimized AAV construct described in Example 16, (b) a fusion protein containing dCas9 and the tripartite effector NFN paired with a non-targeting gRNA (gNT) using the AAV construct described in Example 10, or (c) GFP. As positive controls, genetically correct (GC) FA iPSCs were transduced with AAVDJ vectors encoding GFP or FA iPSCs were transduced with AAVDJ vectors encoding (a) a fusion protein containing dCas9 and the tripartite effector NFN paired with gRNA G (gG) using the AAV construct described in Example 10 or (b) eZFP_A31 fusion proteins containing either the tripartite effector NFN or the transcriptional activator VP64 using the originally designed AAV construct described in Example 16. Cells were harvested on day 7 post-transduction to assess FXN mRNA expression levels by RT-qPCR (normalized to TBP). mRNA expression levels were normalized to control FA-cardiomyocytes expressing a GFP construct for analysis.
[0784] As shown in FIG. 53, treatment of FA-cardiomyocytes with any vector encoding a fusion protein containing one of the exemplary eZFPs eZFP_A31, eZFP_A48, eZFP_A44, or eZFP_A46 increased FXN mRNA levels in a dose-dependent manner. The exemplary AAV containing eZFP_A48 and the tripartite effector NFN in the optimized AAV construct using a CAG promoter led to the highest increase in FXN mRNA of all constructs tested.
[0785] In summary, these results demonstrated that eZFP-NFN fusion proteins targeted to FXN are potent transcriptional activators in FA-iPSC-derived cells. The eZFP tripartite effector fusion protein with the exemplary eZFP_A48 expressed using an optimized AAV construct that included a CAG promoter also significantly increased FXN mRNA levels as compared to all other constructs tested. The
results generally support the therapeutic utility of the eZFPs and eZFP fusion proteins provided herein for targeted activation of FXN expression, including for Friedreich’s ataxia.
[0786] The present invention is not intended to be limited in scope to the particular disclosed embodiments, which are provided, for example, to illustrate various aspects of the invention. Various modifications to the compositions and methods described will become apparent from the description and teachings herein. Such variations may be practiced without departing from the true scope and spirit of the disclosure and are intended to fall within the scope of the present disclosure.
Claims
1. An engineered zinc finger protein (eZFP) that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the target site is within a target region spanning the genomic coordinates chr9:69, 034, 900-69, 035, 900 from human genome assembly GRCh38 (hg38) (SEQ ID NO:430), or within a target region spanning the genomic coordinates chr9:69, 027, 282-69, 028, 497 from hg38 (SEQ ID NO:431).
2. The eZFP of claim 1 , wherein the target site is within a target region spanning the genomic coordinates chr9:69, 034, 900-69, 035, 900 from hg38 (SEQ ID NO:430).
3. The eZFP of claim 1, wherein the target site is within a target region spanning the genomic coordinates chr9:69, 035, 300-69-035, 800 from hg38.
4. The eZFP of claim 1 , wherein the target site is within a target region spanning the genomic coordinates chr9:69, 035, 350-69, 035, 450 from hg38.
5. The eZFP of claim 1, wherein the target site is within a target region spanning the genomic coordinates chr9:69, 035, 400-69, 035, 450 from hg38.
6. The eZFP of claim 1 , wherein the target site is within a target region spanning the genomic coordinates chr9:69, 035, 530-69, 035, 580 from hg38.
7. The eZFP of claim 1, wherein the target site is within a target region spanning the genomic coordinates chr9:69, 035, 675-69, 035, 725 from hg38.
8. The eZFP of claim 1, wherein the target site is within a target region spanning the genomic coordinates chr9:69, 027, 282-69, 028, 497 from hg38 (SEQ ID NO:431).
9. The eZFP of claim 1, wherein the target site is within a target region spanning the genomic coordinates chr9:69, 027, 615-69, 028, 101 from hg38.
10. The eZFP of claim 1, wherein the target site is within a target region spanning the genomic coordinates chr9:69, 027, 775-69, 027, 875 from hg38.
11. The eZFP of claim 1 , wherein the target site is within a target region spanning the genomic coordinates chr9:69, 027, 795-69, 027, 845 from hg38.
12. The eZFP of claim 1, wherein the target site comprises the nucleotide sequence set forth in any one of SEQ ID NOS:269-300 and 583-600, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
13. The eZFP of claim 1, wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:272, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
14. The eZFP of claim 13, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: QSGNLAR (SEQ ID NO:341); F2: QKVNRAG (SEQ ID NO:342); F3:
DRSNLSR (SEQ ID NO:343); F4: QSGHLSR (SEQ ID NO:344); F5: TSGHLSR (SEQ ID NO:345); and F6: RSDALAR (SEQ ID NO:346).
15. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: QSGNLAR (SEQ ID NO:341); F2: QKVNRAG (SEQ ID NO:342); F3: DRSNLSR (SEQ ID NO:343); F4: QSGHLSR (SEQ ID NO:344); F5: TSGHLSR (SEQ ID NO:345); and F6: RSDALAR (SEQ ID NO:346).
16. The eZFP of any of claims 13-15, wherein the eZFP comprises the sequence set forth in SEQ ID NO:301, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
17. The eZFP of any of claims 13-16, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:301.
18. The eZFP of any of claims 13-17, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:308 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
19. The eZFP of any of claims 13-18, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:308.
20. The eZFP of claim 1 , wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:277, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
21. The eZFP of claim 20, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RSDNLSE (SEQ ID NO:347); F2: KSWSRYK (SEQ ID NO:348); F3: TSGSLSR (SEQ ID NO:349); F4: RSDALAR (SEQ ID NO:350); F5: RSDNLSV (SEQ ID NO:351); and F6: FSSCRSA (SEQ ID NO:352).
22. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RSDNLSE (SEQ ID NO:347); F2: KSWSRYK (SEQ ID NO:348); F3: TSGSLSR (SEQ ID NO:349); F4: RSDALAR (SEQ ID NO:350); F5: RSDNLSV (SEQ ID NO:351); and F6: FSSCRSA (SEQ ID NO:352).
23. The eZFP of any of claims 20-22, wherein the eZFP comprises the sequence set forth in SEQ ID NO:302, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
24. The eZFP of any of claims 20-23, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:302.
25. The eZFP of any of claims 20-24, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:309 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
26. The eZFP of any of claims 20-25, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:309.
27. The eZFP of claim 1, wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:280, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
28. The eZFP of claim 27, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: TSGNETR (SEQ ID NO:353); F2: EQTTRDK (SEQ ID NO:354); F3: RSANEAR (SEQ ID NO:355); F4: REDNRTA (SEQ ID NO:356); F5: DSSHRTR (SEQ ID NO:357); and F6: RKYYLAK (SEQ ID NO:358).
29. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: TSGNETR (SEQ ID NO:353); F2: EQTTRDK (SEQ ID NO:354); F3: RSANLAR (SEQ ID NO:355); F4: RLDNRTA (SEQ ID NO:356); F5: DSSHRTR (SEQ ID NO:357); and F6: RKYYLAK (SEQ ID NO:358).
30. The eZFP of any of claims 27-29, wherein the eZFP comprises the sequence set forth in SEQ ID NO:303, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
31. The eZFP of any of claims 27-30, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:303.
32. The eZFP of any of claims 27-31, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NOG 10 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
33. The eZFP of any of claims 27-32, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NOG 10.
34. The eZFP of claim 1 , wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:281, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
35. The eZFP of claim 34, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RSAHLSR (SEQ ID NO:359); F2: DRSDLSR (SEQ ID NO:360); F3: RSDHLSV (SEQ ID NO:361); F4: RSDVRKT (SEQ ID NO:362); F5: QSGALAR (SEQ ID NO:363); and F6: RKYYLAK (SEQ ID NO:364).
36. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RSAHLSR (SEQ ID NO:359); F2: DRSDLSR (SEQ ID NO:360); F3: RSDHLSV (SEQ ID NO:361); F4: RSDVRKT (SEQ ID NO:362); F5: QSGALAR (SEQ ID NO:363); and F6: RKYYLAK (SEQ ID NO:364).
37. The eZFP of any of claims 34-36, wherein the eZFP comprises the sequence set forth in SEQ ID NO:304, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
38. The eZFP of any of claims 34-37, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:304.
39. The eZFP of any of claims 34-38, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:311 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
40. The eZFP of any of claims 34-39, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:311.
41. The eZFP of claim 1, wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:283, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
42. The eZFP of claim 41, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RSAHLSR (SEQ ID NO:365); F2: RSDALAR (SEQ ID NO:366); F3: ATSNRSA (SEQ ID NO:367); F4: RSAHLSR (SEQ ID NO:368); F5: TSGSLSR (SEQ ID NO:369); and F6: QSGDLTR (SEQ ID NO:370).
43. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RSAHLSR (SEQ ID NO:365); F2: RSDALAR (SEQ ID NO:366); F3: ATSNRSA (SEQ ID NO:367); F4: RSAHLSR (SEQ ID NO:368); F5: TSGSLSR (SEQ ID NO:369); and F6: QSGDLTR (SEQ ID NO:370).
44. The eZFP of any of claims 41-43, wherein the eZFP comprises the sequence set forth in SEQ ID NO:305, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
45. The eZFP of any of claims 41-44, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:305.
46. The eZFP of any of claims 41-45, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:312 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
47. The eZFP of any of claims 41-46, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:312.
48. The eZFP of claim 1, wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:290, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
49. The eZFP of claim 48, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: QSGDETR (SEQ ID NO:371); F2: QSSDERR (SEQ ID NO:372); F3: RSDNESE (SEQ ID NO:373); F4: SSRNEAS (SEQ ID NO:374); F5: DRSHLTR (SEQ ID NO:375); and F6: RSDDLTR (SEQ ID NO:376).
50. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: QSGDETR (SEQ ID NO:371); F2: QSSDERR (SEQ ID NO:372); F3: RSDNLSE (SEQ ID NO:373); F4: SSRNLAS (SEQ ID NO:374); F5: DRSHLTR (SEQ ID NO:375); and F6: RSDDLTR (SEQ ID NO:376).
51. The eZFP of any of claims 48-50, wherein the eZFP comprises the sequence set forth in SEQ ID NO:306, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
52. The eZFP of any of claims 48-51, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:306.
53. The eZFP of any of claims 48-52, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:313 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
54. The eZFP of any of claims 48-53, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:313.
55. The eZFP of claim 1, wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:299, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
56. The eZFP of claim 55, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: LRHHLTR (SEQ ID NO:377); F2: QSAHLKA (SEQ ID NO:378); F3: LPQTLQR (SEQ ID NO:379); F4: QNATRTK (SEQ ID NO:380); F5: QSSHLTR (SEQ ID NO:381); and F6: RSDHLSR (SEQ ID NO:382).
57. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: LRHHLTR (SEQ ID NO:377); F2: QSAHLKA (SEQ ID NO:378); F3: LPQTLQR (SEQ ID NO:379); F4: QNATRTK (SEQ ID NO:380); F5: QSSHLTR (SEQ ID NO:381); and F6: RSDHLSR (SEQ ID NO:382).
58. The eZFP of any of claims 55-57, wherein the eZFP comprises the sequence set forth in SEQ ID NO:307, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
59. The eZFP of any of claims 55-58, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:307.
60. The eZFP of any of claims 55-59, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:314 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
61. The eZFP of any of claims 55-60, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:314.
62. The eZFP of claim 1 , wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:583, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
63. The eZFP of claim 62, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RSDSLLR (SEQ ID NO:475); F2: TSSNRKT (SEQ ID NO:476); F3: RSAHLSR (SEQ ID NO:477); F4: TSGSLTR (SEQ ID NO:478); F5: QSGDLTR (SEQ ID NO:479); and F6: QWGTRYR (SEQ ID NO:480).
64. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RSDSLLR (SEQ ID NO:475); F2: TSSNRKT (SEQ ID NO:476); F3: RSAHLSR (SEQ ID NO:477); F4: TSGSLTR (SEQ ID NO:478); F5: QSGDLTR (SEQ ID NO:479); and F6: QWGTRYR (SEQ ID NO:480).
65. The eZFP of any of claims 62-64, wherein the eZFP comprises the sequence set forth in SEQ ID NO:439, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
66. The eZFP of any of claims 62-65, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:439.
67. The eZFP of any of claims 62-66, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:457 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
68. The eZFP of any of claims 62-67, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:457.
69. The eZFP of claim 1, wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:584, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
70. The eZFP of claim 69, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: QARHETC (SEQ ID NO:481); F2: QSGHESR (SEQ ID NO:482); F3: RSDVESE (SEQ ID NO:483); F4: KHSTRRV (SEQ ID NO:484); F5: QSSDLSR (SEQ ID NO:485); and F6: WKWNLRA (SEQ ID NO:486).
71. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: QARHETC (SEQ ID NO:481); F2: QSGHESR (SEQ ID NO:482); F3: RSDVLSE (SEQ ID NO:483); F4: KHSTRRV (SEQ ID NO:484); F5: QSSDLSR (SEQ ID NO:485); and F6: WKWNLRA (SEQ ID NO:486).
72. The eZFP of any of claims 69-71, wherein the eZFP comprises the sequence set forth in SEQ ID NO:440, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
73. The eZFP of any of claims 69-72, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:440.
74. The eZFP of any of claims 69-73, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:458 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
75. The eZFP of any of claims 69-74, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:458.
76. The eZFP of claim 1, wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:585, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
77. The eZFP of claim 76, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RSDNLAR (SEQ ID NO:487); F2: WRGDRVK (SEQ ID NO:488); F3: YKHVLSD (SEQ ID NO:489); F4: TSGSLTR (SEQ ID NO:490); F5: QSGNLAR (SEQ ID NO:491); and F6: RARDLSK (SEQ ID NO:492).
78. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RSDNLAR (SEQ ID NO:487); F2: WRGDRVK (SEQ ID NO:488); F3: YKHVLSD (SEQ ID NO:489); F4: TSGSLTR (SEQ ID NO:490); F5: QSGNLAR (SEQ ID NO:491); and F6: RARDLSK (SEQ ID NO:492).
79. The eZFP of any of claims 76-78, wherein the eZFP comprises the sequence set forth in SEQ ID NO:441, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
80. The eZFP of any of claims 76-79, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:441.
81. The eZFP of any of claims 76-80, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:459 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
82. The eZFP of any of claims 76-81, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:459.
83. The eZFP of claim 1, wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:586, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
84. The eZFP of claim 83, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: QSANRTK (SEQ ID NO:493); F2: QSGNLAR (SEQ ID NO:494); F3: RSDNLSV (SEQ ID NO:495); F4: IRSTLRD (SEQ ID NO:496); F5: QNAHRKT (SEQ ID NO:497); and F6: HRSSLRR (SEQ ID NO:498).
85. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: QSANRTK (SEQ ID NO:493); F2: QSGNLAR (SEQ ID NO:494); F3: RSDNLSV (SEQ ID NO:495); F4: IRSTLRD (SEQ ID NO:496); F5: QNAHRKT (SEQ ID NO:497); and F6: HRSSLRR (SEQ ID NO:498).
86. The eZFP of any of claims 83-85, wherein the eZFP comprises the sequence set forth in SEQ ID NO:442, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
87. The eZFP of any of claims 83-86, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:442.
88. The eZFP of any of claims 83-87, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:460 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
89. The eZFP of any of claims 83-88, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:460.
90. The eZFP of claim 1 , wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:587, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
91. The eZFP of claim 90, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: QAGNRST (SEQ ID NO:499); F2: DRSAEAR (SEQ ID NO:500); F3: RSDNEAR (SEQ ID NO:501); F4: WRGDRVK (SEQ ID NO:502); F5: YKHVLSD (SEQ ID NO:503); and F6: TSGSLTR (SEQ ID NO:504).
92. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: QAGNRST (SEQ ID NO:499); F2: DRSAEAR (SEQ ID NG:500); F3: RSDNLAR (SEQ ID NO:501); F4: WRGDRVK (SEQ ID NO:502); F5: YKHVLSD (SEQ ID NO:503); and F6: TSGSLTR (SEQ ID NO:504).
93. The eZFP of any of claims 90-92, wherein the eZFP comprises the sequence set forth in SEQ ID NO:443, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
94. The eZFP of any of claims 90-93, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:443.
95. The eZFP of any of claims 90-94, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:461 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
96. The eZFP of any of claims 90-95, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:461.
97. The eZFP of claim 1, wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:588, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
98. The eZFP of claim 97, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RSDNLSV (SEQ ID NO:505); F2: IRSTLRD (SEQ ID NO:506); F3: QNAHRKT (SEQ ID NO:507); F4: HRSSLRR (SEQ ID NO:508); F5: RSDNLAR (SEQ ID NO:509); and F6: QRSPLPA (SEQ ID NO:51Q).
99. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RSDNLSV (SEQ ID NO:505); F2: IRSTLRD (SEQ ID NO:506); F3: QNAHRKT (SEQ ID NO:507); F4: HRSSLRR (SEQ ID NO:508); F5: RSDNLAR (SEQ ID NO:509); and F6: QRSPLPA (SEQ ID NO:51Q).
100. The eZFP of any of claims 97-99, wherein the eZFP comprises the sequence set forth in SEQ ID NO:444, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
101. The eZFP of any of claims 97-100, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:444.
102. The eZFP of any of claims 97-101, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:462 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
103. The eZFP of any of claims 97-102, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:462.
104. The eZFP of claim 1, wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:589, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
105. The eZFP of claim 104, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: DRSTRTK (SEQ ID NO:511); F2: RSDYLAK (SEQ ID NO:512); F3: LRHHLTR (SEQ ID NO:513); F4: QSAHLKA (SEQ ID NO:514); F5: LPQTLQR (SEQ ID NO:515); and F6: QNATRTK (SEQ ID NO:516).
106. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: DRSTRTK (SEQ ID NO:511); F2: RSDYLAK (SEQ ID NO:512); F3: LRHHLTR (SEQ ID NO:513); F4: QSAHLKA (SEQ ID NO:514); F5: LPQTLQR (SEQ ID NO:515); and F6: QNATRTK (SEQ ID NO:516).
107. The eZFP of any of claims 104-106, wherein the eZFP comprises the sequence set forth in SEQ ID NO:445, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
108. The eZFP of any of claims 104-107, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:445.
109. The eZFP of any of claims 104-108, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 463 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
110. The eZFP of any of claims 104-109, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 463.
111. The eZFP of claim 1 , wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:590, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
112. The eZFP of claim 111, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RSADETR (SEQ ID NO:517); F2: RSDDETR (SEQ ID NO:518); F3: QSSDESR (SEQ ID NO:519); F4: WHSSEHQ (SEQ ID NO:520); F5: RSDSLSQ (SEQ ID NO:521); and F6: RKADRTR (SEQ ID NO:522).
113. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RSADETR (SEQ ID NO:517); F2: RSDDETR (SEQ ID NO:518); F3: QSSDLSR (SEQ ID NO:519); F4: WHSSLHQ (SEQ ID NO:520); F5: RSDSLSQ (SEQ ID NO:521); and F6: RKADRTR (SEQ ID NO:522).
114. The eZFP of any of claims 111-113, wherein the eZFP comprises the sequence set forth in SEQ ID NO:446, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
115. The eZFP of any of claims 111-114, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:446.
116. The eZFP of any of claims 111-115, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 464 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
117. The eZFP of any of claims 111-116, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:464.
118. The eZFP of claim 1, wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:591, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
119. The eZFP of claim 118, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RNDALTE (SEQ ID NO:523); F2: RKDNLKN (SEQ ID NO:524); F3: TSGELVR (SEQ ID NO:525); F4: HRTTLTN (SEQ ID NO:526); F5: TTGNLTV (SEQ ID NO:527); and F6: RTDTLRD (SEQ ID NO:528).
120. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RNDALTE (SEQ ID NO:523); F2: RKDNLKN (SEQ ID NO:524); F3: TSGELVR (SEQ ID NO:525); F4: HRTTLTN (SEQ ID NO:526); F5: TTGNLTV (SEQ ID NO:527); and F6: RTDTLRD (SEQ ID NO:528).
121. The eZFP of any of claims 118-120, wherein the eZFP comprises the sequence set forth in SEQ ID NO:447, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
122. The eZFP of any of claims 118-121, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:447.
123. The eZFP of any of claims 118-122, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 465 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
124. The eZFP of any of claims 118-123, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 465.
125. The eZFP of claim 1, wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:592, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
126. The eZFP of claim 125, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RKDNLKN (SEQ ID NO:529); F2: RADNLTE (SEQ ID NO:530); F3: TSHSLTE (SEQ ID NO:531); F4: SKKHLAE (SEQ ID NO:532); F5: TSGELVR (SEQ ID NO:533); and F6: TSGELVR (SEQ ID NO:534).
127. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RKDNLKN (SEQ ID NO:529); F2: RADNLTE (SEQ ID NO:530); F3: TSHSLTE (SEQ ID NO:531); F4: SKKHLAE (SEQ ID NO:532); F5: TSGELVR (SEQ ID NO:533); and F6: TSGELVR (SEQ ID NO:534).
128. The eZFP of any of claims 125-127, wherein the eZFP comprises the sequence set forth in SEQ ID NO:448, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
129. The eZFP of any of claims 125-128, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:448.
130. The eZFP of any of claims 125-129, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 466 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
131. The eZFP of any of claims 125-130, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:466.
132. The eZFP of claim 1, wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:593, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
133. The eZFP of claim 132, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: THEDEIR (SEQ ID NO:535); F2: DCRDEAR (SEQ ID NO:536); F3: RSDEEVR (SEQ ID NO:537); F4: RNDAETE (SEQ ID NO:538); F5: SKKHLAE (SEQ ID NO:539); and F6: QSGHLTE (SEQ ID NO:540).
134. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: THEDEIR (SEQ ID NO:535); F2: DCRDEAR (SEQ ID NO:536); F3: RSDELVR (SEQ ID NO:537); F4: RNDALTE (SEQ ID NO:538); F5: SKKHLAE (SEQ ID NO:539); and F6: QSGHLTE (SEQ ID NO:540).
135. The eZFP of any of claims 132-134, wherein the eZFP comprises the sequence set forth in SEQ ID NO:449, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
136. The eZFP of any of claims 132-135, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:449.
137. The eZFP of any of claims 132-136, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 467 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
138. The eZFP of any of claims 132-137, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 467.
139. The eZFP of claim 1, wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:594, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
140. The eZFP of claim 139, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: HTGHLLE (SEQ ID NO:541); F2: DPGHLVR (SEQ ID NO:542); F3: THLDLIR (SEQ ID NO:543); F4: DCRDLAR (SEQ ID NO:544); F5: RSDELVR (SEQ ID NO:545); and F6: RNDALTE (SEQ ID NO:546).
141. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: HTGHLLE (SEQ ID NO:541); F2: DPGHLVR (SEQ ID NO:542); F3: THLDLIR (SEQ ID NO:543); F4: DCRDLAR (SEQ ID NO:544); F5: RSDELVR (SEQ ID NO:545); and F6: RNDALTE (SEQ ID NO:546).
142. The eZFP of any of claims 139-141, wherein the eZFP comprises the sequence set forth in SEQ ID NO:450, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
143. The eZFP of any of claims 139-142, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:450.
144. The eZFP of any of claims 139-143, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:468 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
145. The eZFP of any of claims 139-144, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:468.
146. The eZFP of claim 1, wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:595, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
147. The eZFP of claim 146, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RSDKLVR (SEQ ID NO:547); F2: RSDHLTT (SEQ ID NO:548); F3: RNDALTE (SEQ ID NO:549); F4: TTGALTE (SEQ ID NO:550); F5: THLDLIR (SEQ ID NO:551); and F6: DPGHLVR (SEQ ID NO:552).
148. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RSDKLVR (SEQ ID NO:547); F2: RSDHLTT (SEQ ID NO:548); F3: RNDALTE (SEQ ID NO:549); F4: TTGALTE (SEQ ID NO:550); F5: THLDLIR (SEQ ID NO:551); and F6: DPGHLVR (SEQ ID NO:552).
149. The eZFP of any of claims 146-148, wherein the eZFP comprises the sequence set forth in SEQ ID NO:451, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
150. The eZFP of any of claims 146-149, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:451.
151. The eZFP of any of claims 146-150, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 469 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
152. The eZFP of any of claims 146-151, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 469.
153. The eZFP of claim 1, wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:596, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
154. The eZFP of claim 153, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: TKNSETE (SEQ ID NO:553); F2: QEAHERA (SEQ ID NO:554); F3: TSGSEVR (SEQ ID NO:555); F4: RSDNLVR (SEQ ID NO:556); F5: QNSTLTE (SEQ ID NO:557); and F6: RADNLTE (SEQ ID NO:558).
155. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: TKNSLTE (SEQ ID NO:553); F2: QLAHLRA (SEQ ID NO:554); F3: TSGSLVR (SEQ ID NO:555); F4: RSDNLVR (SEQ ID NO:556); F5: QNSTLTE (SEQ ID NO:557); and F6: RADNLTE (SEQ ID NO:558).
156. The eZFP of any of claims 153-155, wherein the eZFP comprises the sequence set forth in SEQ ID NO:452, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
157. The eZFP of any of claims 153-156, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:452.
158. The eZFP of any of claims 153-157, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 470 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
159. The eZFP of any of claims 153-158, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:470.
160. The eZFP of claim 1, wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:597, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
161. The eZFP of claim 160, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RADNLTE (SEQ ID NO:559); F2: TKNSLTE (SEQ ID NO:560); F3: QLAHLRA (SEQ ID NO:561); F4: TSGSLVR (SEQ ID NO:562); F5: RSDNLVR (SEQ ID NO:563); and F6: QNSTLTE (SEQ ID NO:564).
162. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: RADNLTE (SEQ ID NO:559); F2: TKNSLTE (SEQ ID NO:560); F3: QLAHLRA (SEQ ID NO:561); F4: TSGSLVR (SEQ ID NO:562); F5: RSDNLVR (SEQ ID NO:563); and F6: QNSTLTE (SEQ ID NO:564).
163. The eZFP of any of claims 160-162, wherein the eZFP comprises the sequence set forth in SEQ ID NO:453, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
164. The eZFP of any of claims 160-163, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:453.
165. The eZFP of any of claims 160-164, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 471 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
166. The eZFP of any of claims 160-165, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 471.
167. The eZFP of claim 1, wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:598, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
168. The eZFP of claim 167, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: TSGHLVR (SEQ ID NO:565); F2: QLAHLRA (SEQ ID NO:566); F3: TSGELVR (SEQ ID NO:567); F4: QSGDLRR (SEQ ID NO:568); F5: QRAHLER (SEQ ID NO:569); and F6: RSDKLVR (SEQ ID NO:570).
169. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: TSGHLVR (SEQ ID NO:565); F2: QLAHLRA (SEQ ID NO:566); F3: TSGELVR (SEQ ID NO:567); F4: QSGDLRR (SEQ ID NO:568); F5: QRAHLER (SEQ ID NO:569); and F6: RSDKLVR (SEQ ID NO:570).
170. The eZFP of any of claims 167-169, wherein the eZFP comprises the sequence set forth in SEQ ID NO:454, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
171. The eZFP of any of claims 167-170, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:454.
172. The eZFP of any of claims 167-171, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 472 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
173. The eZFP of any of claims 167-172, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:472.
174. The eZFP of claim 1, wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:599, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
175. The eZFP of claim 174, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: REDNEHT (SEQ ID NO:571); F2: TSGHEVR (SEQ ID NO:572); F3: QEAHERA (SEQ ID NO:573); F4: TSGELVR (SEQ ID NO:574); F5: QSGDLRR (SEQ ID NO:575); and F6: QRAHLER (SEQ ID NO:576).
176. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: REDNLHT (SEQ ID NO:571); F2: TSGHLVR (SEQ ID NO:572); F3: QLAHLRA (SEQ ID NO:573); F4: TSGELVR (SEQ ID NO:574); F5: QSGDLRR (SEQ ID NO:575); and F6: QRAHLER (SEQ ID NO:576).
177. The eZFP of any of claims 174-176, wherein the eZFP comprises the sequence set forth in SEQ ID NO:455, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
178. The eZFP of any of claims 174-177, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:455.
179. The eZFP of any of claims 174-178, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 473 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
180. The eZFP of any of claims 174-179, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 473.
181. The eZFP of claim 1 , wherein the target site comprises the nucleotide sequence set forth in SEQ ID NO:600, a contiguous portion thereof of at least 12 nt, or a complementary sequence of any of the foregoing.
182. The eZFP of claim 181, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: QRSDLTR (SEQ ID NO:577); F2: QGGTLRR (SEQ ID NO:578); F3: TSAHLAR (SEQ ID NO:579); F4: RREHLVR (SEQ ID NO:580); F5: QRHGLSS (SEQ ID NO:581); and F6: QRNALRG (SEQ ID NO:582).
183. An engineered zinc finger protein (eZFP), that binds to a target site in a regulatory DNA element of a frataxin (FXN) locus, wherein the eZFP comprises a zinc finger recognition region comprising six zinc fingers denoted Fl through F6 in order from N-terminus to C-terminus, selected from F1-F6 as follows: Fl: QRSDLTR (SEQ ID NO:577); F2: QGGTLRR (SEQ ID NO:578); F3: TSAHLAR (SEQ ID NO:579); F4: RREHLVR (SEQ ID NO:580); F5: QRHGLSS (SEQ ID NO:581); and F6: QRNALRG (SEQ ID NO:582).
184. The eZFP of any of claims 181-183, wherein the eZFP comprises the sequence set forth in SEQ ID NO:456, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
185. The eZFP of any of claims 181-184, wherein the engineered zinc finger protein comprises the sequence set forth in SEQ ID NO:456.
186. The eZFP of any of claims 181-185, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO: 474 or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
187. The eZFP of any of claims 181-186, wherein the engineered zinc finger protein is encoded by the sequence set forth in SEQ ID NO:474.
188. A fusion protein comprising the engineered zinc finger protein (eZFP) of any of claims 1-187.
189. A fusion protein comprising:
(a) the engineered zinc finger protein of any of claims 1-187 that binds to a target site in a regulatory DNA element of a FXN locus; and
(b) at least one epigenetic effector domain that increases transcription of the FXN locus.
190. The fusion protein of claim 189, wherein the at least one epigenetic effector domain comprises: a VP64 domain, a p65 activation domain, a p300 domain, an Rta domain, a CBP domain, a VPR domain, a VPH domain, an HSF1 domain, a TET protein domain, optionally wherein the TET protein is TET1, a SunTag domain, a domain from DPOLA, ENL, FOXO3, HSH2D, NCOA2, NCOA3, PSA1, PYGO1, RBM39, HERC2, or NOTCH2, or a domain, portion, variant, or truncation of any of the foregoing.
191. The fusion protein of claim 189 or 190, wherein the at least one epigenetic effector domain comprises the sequence set forth in any of SEQ ID NOS:81, 83, 100-109, 111-122, 124, 125, 134-140, 152, and 383-396, or a domain, portion, variant, or truncation thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
192. The fusion protein of any of claims 189-191, wherein the at least one effector domain comprises at least one VP16 domain, or a VP16 tetramer (“VP64”) or a variant thereof.
193. The fusion protein of any of claims 189-192, wherein the at least one effector domain comprises VP64.
194. The fusion protein of any of claims 189-193, wherein the at least one effector domain comprises a VP64 domain comprising the sequence set forth in SEQ ID NO:83, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
195. The fusion protein of any of claims 189-194, wherein the at least one effector domain comprises a VP64 domain comprising the sequence set forth in SEQ ID NO:83.
196. The fusion protein of any of claims 189-195, wherein the at least one epigenetic effector domain comprises: a domain from DPOLA, ENL, FOXO3, HSH2D, NCOA2, NCOA3, PSA1, PYGO1, RBM39, HERC2, or NOTCH2, or a domain, portion, variant, or truncation of any of the foregoing.
197. The fusion protein of any of claims 189-196, wherein the at least one epigenetic effector domain comprises the sequence set forth in any of SEQ ID NOS:383-393, or a domain, portion, or variant thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
198. The fusion protein of any of claims 189-197, wherein the at least one effector domain comprises a domain from NCOA2, NCOA3, FOXO3, PYGO1, or a portion or variant of any of the foregoing.
199. The fusion protein of any of claims 189-198, wherein each effector domain of the at least one effector domain is independently selected from an NCOA2 domain, an NCOA3 domain, a FOXO3 domain, and a PYGO1 domain.
200. The fusion protein of any of claims 189-199, wherein the at least one effector domain comprises a domain from NCOA2 comprising the sequence set forth in SEQ ID NO: 104 or SEQ ID NO:387, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
201. The fusion protein of any of claims 189-200, wherein the at least one effector domain comprises a domain from NCOA2 set forth in or SEQ ID NO:387.
202. The fusion protein of any of claims 189-201, wherein the at least one effector domain comprises a domain from NCOA3 comprising the sequence set forth in SEQ ID NO: 105 or SEQ ID NO:388, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
203. The fusion protein of any of claims 189-202, wherein the at least one effector domain comprises a domain from NCOA3 set forth in or SEQ ID NO:388.
204. The fusion protein of any of claims 189-203, wherein the at least one effector domain comprises a domain from FOXO3 comprising the sequence set forth in SEQ ID NO: 102 or SEQ ID NO:385, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
205. The fusion protein of any of claims 189-204, wherein the at least one effector domain comprises a domain from FOXO3 set forth in or SEQ ID NO:385.
206. The fusion protein of any of claims 189-205, wherein the at least one effector domain comprises a domain from PYGO1 comprising the sequence set forth in SEQ ID NO: 107 or SEQ ID NO:390, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
207. The fusion protein of any of claims 189-206, wherein the at least one effector domain comprises a domain from PYGO1 set forth in or SEQ ID NO:390.
208. The fusion protein of any of claims 189-207, wherein the at least one effector domain is a multipartite effector composed of at least two effector domains.
209. The fusion protein of claim 208, wherein the multipartite effector is composed of two effector domains.
210. The fusion protein of claim 208, wherein the multipartite effector is composed of three effector domains.
211. The fusion protein of any of claims 208-210, wherein the multipartite effector is set forth in any one of SEQ ID NOS:397-418, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
212. The fusion protein of any of claims 208-211, wherein the multipartite effector is set forth in any one of SEQ ID NOS:411-418, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
213. The fusion protein of any of claims 208-212, wherein the multipartite effector comprises, in the N-terminal to C-terminal direction, domains from FOXO3, FOXO3, and NCOA3, respectively.
214. The fusion protein of claim 213, wherein the multipartite effector comprises the sequence set forth in SEQ ID NO:415, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
215. The fusion protein of any of claims 208-212, wherein the multipartite effector comprises, in the N-terminal to C-terminal direction, domains from NCOA3, FOXO3, and FOXO3, respectively.
216. The fusion protein of claim 215, wherein the multipartite effector comprises the sequence set forth in SEQ ID NO:418, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
217. The fusion protein of any of claims 208-212, wherein the multipartite effector comprises, in the N-terminal to C-terminal direction, domains from NCOA3, FOXO3, and NCOA3, respectively.
218. The fusion protein of claim 217, wherein the multipartite effector comprises the sequence set forth in SEQ ID NO:413, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
219. The fusion protein of any of claims 208-212, wherein the multipartite effector comprises, in the N-terminal to C-terminal direction, domains from NCOA2, FOXO3, and NCOA3, respectively.
220. The fusion protein of claim 219, wherein the multipartite effector comprises the sequence set forth in SEQ ID NO:416, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
221. The fusion protein of any of claims 208-212, wherein the multipartite effector comprises, in the N-terminal to C-terminal direction, domains from PYGO1, FOXO3, and NCOA3, respectively.
222. The fusion protein of claim 221, wherein the multipartite effector comprises the sequence set forth in SEQ ID NO:411, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
223. The fusion protein of any of claims 189-222, wherein the at least one epigenetic effector domain is fused to the N-terminus of the eZFP.
224. The fusion protein of any of claims 189-222, wherein the at least one epigenetic effector domain is fused to the C-terminus of the eZFP.
225. The fusion protein of any of claims 189-224, wherein the at least one epigenetic effector domain is fused to both the N-terminus and the C-terminus of the eZFP.
226. The fusion protein of any of claims 189-225, further comprising one or more nuclear localization signals (NFS).
227. The fusion protein of any of claims 189-226, further comprising one or more linkers.
228. The fusion protein of claim 227, wherein the one or more linkers are in between any two of the components of the fusion protein, including the eZFP, any of the at least one effector domains, and the one or more NFS.
229. The fusion protein of claim 227 or 228, wherein the one or more linkers connect the eZFP and the at least one epigenetic effector domain.
230. The fusion protein of any of claims 226-229, comprising the one or more NFS, the eZFP, and the at least one epigenetic effector domain, in order from N-terminus to C-terminus.
231. The fusion protein of claim 230, wherein the one or more NFS comprises a SV40 NFS sequence set forth in SEQ ID NO: 159 or a c-myc NFS sequence set forth in SEQ ID NO: 160.
232. The fusion protein of any of claims 189-231, wherein the fusion protein comprises the sequence set forth in any of SEQ ID NOS:320-340, 419-425, and 608-671, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
233. The fusion protein of any of claims 189-232, wherein the fusion protein comprises the sequence set forth in any of SEQ ID NOS:320-340, 419-425, and 608-671.
234. The fusion protein of any of claims 189-233, wherein the fusion protein comprises the sequence set forth in any of SEQ ID NOS:320-340, 419-425, and 636-653.
235. The fusion protein of any of claims 189-234, wherein the fusion protein comprises the sequence set forth in any of SEQ ID NOS:320-340 and 419-425.
236. The fusion protein of any of claims 189-234, wherein the fusion protein comprises the sequence set forth in any of SEQ ID NOS:636-653.
237. The fusion protein of any of claims 189-233, wherein the fusion protein comprises the sequence set forth in any of SEQ ID NOS:608-635 and 654-671.
238. The fusion protein of any of claims 189-233 and 237, wherein the fusion protein comprises the sequence set forth in any of SEQ ID NOS:608-635.
239. The fusion protein of any of claims 129-233 and 237, wherein the fusion protein comprises the sequence set forth in any of SEQ ID NOS: 654-671.
240. The fusion protein of any of claims 189-235, wherein the fusion protein comprises the sequence set forth in SEQ ID NO:326, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
241. The fusion protein of any of claims 189-235, wherein the fusion protein comprises the sequence set forth in SEQ ID NO:333, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
242. The fusion protein of any of claims 189-235, wherein the fusion protein comprises the sequence set forth in SEQ ID NO:340, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
243. The fusion protein of any of claims 189-235, wherein the fusion protein comprises the sequence set forth in SEQ ID NO:425, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
244. The fusion protein of any of claims 189-233, 237, and 239, wherein the fusion protein comprises the sequence set forth in SEQ ID NO:662, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
245. The fusion protein of any of claims 189-233, 237, and 239, wherein the fusion protein comprises the sequence set forth in SEQ ID NO:660, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
246. The fusion protein of any of claims 189-233, 237, and 239, wherein the fusion protein comprises the sequence set forth in SEQ ID NO:658, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
247. A polynucleotide encoding the eZFP of any of claims 1-187, the fusion protein of any of claims 188-246, or a portion or component of any of the foregoing.
248. A plurality of polynucleotides encoding the eZFP of any of claims 1-187, the fusion protein of any of claims 188-246, or a portion or component of any of the foregoing.
249. A vector comprising the eZFP of any of claims 1-187, the fusion protein of any of claims 188-246, the polynucleotide of claim 247, the plurality of polynucleotides of claim 248, or a portion or component of any of the foregoing.
250. The vector of claim 249, wherein the vector is a viral vector.
251. The vector of claim 249 or 250, wherein the vector is an adeno-associated virus (AAV) vector.
252. The vector of claim 251 , wherein the AAV vector is selected from the group consisting of: AAV1, AAV2, AAV3, AAV4, AAV5, AAV6, AAV7, AAV8, AAV9, AAV10, AAV11, AAV12, AAV-DJ, and AAVrh74.
253. The vector of claim 251 or 252, wherein the AAV vector is AAV6.
254. The vector of claim 251 or 252, wherein the AAV vector is AAV9.
255. The vector of claim 251 or 252, wherein the AAV vector is AAV-DJ.
256. The vector of claim 251 or 252, wherein the AAV vector is AAVrh74.
257. The vector of claim 249 or 250, wherein the vector is a lentiviral vector.
258. The vector of claim 249, wherein the vector is a non-viral vector.
259. The vector of claim 258, wherein the non-viral vector is selected from the group consisting of: a lipid nanoparticle, a liposome, an exosome, and a cell penetrating peptide.
260. The vector of any of claims 249-259, wherein the vector exhibits tropism for a nervous system cell, optionally a neuron, a heart cell, optionally a cardiomyocyte, a skeletal muscle cell, a fibroblast, an induced pluripotent stem cell, and/or a cell derived from any of the foregoing, or for a combination of any of the foregoing cells.
261. The vector of any of claims 249-260, wherein the vector exhibits tropism for induced pluripotent stem cells.
262. The vector of any of claims 249-261, wherein the vector exhibits tropism for neurons and cardiomyocytes.
263. The vector of any of claims 249-262, wherein the vector comprises one vector, or two or more vectors.
264. An AAV vector comprising one or both of: a) a first nucleic acid comprising an elongation factor alpha short (EFS) promoter operably linked to a sequence encoding a fusion protein comprising (i) a deactivated Cas (dCas) protein and (ii) at least one effector domain that increases transcription of a frataxin (FXN) locus; and b) a second nucleic acid comprising a U6 promoter operably linked to a sequence encoding a guide RNA (gRNA) comprising a gRNA spacer sequence that is capable of hybridizing to a target site in a regulatory DNA element of a FXN locus and/or is complementary to the target site.
265. The AAV vector of claim 264, wherein the AAV vector comprises both the first nucleic acid and the second nucleic acid.
266. The AAV vector of claim 264 or 265, wherein the first and second nucleic acid are comprised in a single polynucleotide.
267. The AAV vector of any of claims 264-266, wherein the EFS promoter comprises the sequence set forth in SEQ ID NO:436, or a sequence having at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to thereto.
268. The AAV vector of any of claims 264-267, wherein the EFS promoter comprises the sequence set forth in SEQ ID NO:436.
269. The AAV vector of any of claims 264-268, wherein the U6 promoter is a mini-U6 promoter.
270. The AAV vector of any of claims 264-269, wherein the mini-U6 promoter comprises the sequence set forth in SEQ ID NO:433, or a sequence having at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to thereto.
271. The AAV vector of any of claims 264-270, wherein the mini-U6 promoter comprises the sequence set forth in SEQ ID NO: 433.
272. An AAV vector comprising a nucleic acid comprising a promoter selected from an elongation factor alpha short (EFS), CAG, or human elongation factor-1 alpha (EFla) promoter operably linked to a sequence encoding a fusion protein comprising (i) an eZFP that is capable of hybridizing to a target site in a regulatory DNA element of a frataxin (FXN) locus and/or is complementary to the target site and (ii) at least one effector domain that increases transcription of the frataxin (FXN) locus.
273. The AAV vector of any of claims 264-272, wherein the EFS promoter comprises the sequence set forth in SEQ ID NO:436, or a sequence having at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to thereto.
274. The AAV vector of any of claims 264-273, wherein the EFS promoter comprises the sequence set forth in SEQ ID NO:436.
275. The AAV vector of claim 272, wherein the CAG promoter comprises the sequence set forth in SEQ ID NO:602, or a sequence having at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to thereto.
276. The AAV vector of claim 272 or 275, wherein the CAG promoter comprises the sequence set forth in SEQ ID NO:602.
277. The AAV vector of claim 272, wherein the EFla promoter comprises the sequence set forth in SEQ ID NO:603, or a sequence having at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to thereto.
278. The AAV vector of claim 272 or 277, wherein the EFla promoter comprises the sequence set forth in SEQ ID NO: 603.
279. The AAV vector of any of claims 272-278, wherein the nucleic acid further comprises a 5’ untranslated region (UTR) set forth in SEQ ID NO: 605.
280. The AAV vector of any of claims 264-279, wherein the AAV vector further comprises inverted terminal repeats (ITRs).
281. The AAV vector of claim 280, wherein the ITRs are a first and second ITR, comprising the sequences set forth in SEQ ID NO:434 and SEQ ID NO:435, respectively.
282. The AAV vector of any of claims 266-271 , 280, and 281 , wherein the single polynucleotide comprises, in the 5’ to 3’ direction, the EFS promoter, the sequence encoding the fusion protein, the U6 promoter, and the sequence encoding the gRNA.
283. The AAV vector of any of claims 266-271 and 280-282, wherein the single polynucleotide further comprises a first IRT 5’ of the EFS promoter and a second ITR 3’ of the sequence encoding the gRNA.
284. The AAV vector of any of claims 264-283, wherein the first nucleic acid or the nucleic acid further comprises a polyA sequence selected from a SpA site or a bGH site downstream of the sequence encoding the fusion protein.
285. The AAV vector of any of claims 264-284, wherein the first nucleic acid or the nucleic acid comprising a sequence encoding a fusion protein further comprises a polyA sequence selected from a SpA site downstream of the sequence encoding the fusion protein.
286. The AAV vector of any of claims 264-271, 273, 274, and 282-284, wherein the first nucleic acid comprising a sequence encoding a fusion protein further comprises a polyA sequence selected from a bGH site downstream of the sequence encoding the fusion protein.
287. The AAV vector of any of claims 272-281, 284, and 285, wherein the nucleic acid comprising a sequence encoding a fusion protein further comprises a polyA sequence selected from a SpA site or bGH site downstream of the sequence encoding the fusion protein.
288. The AAV vector of any of claims 284, 285, and 287, wherein the SpA site comprises the sequence set forth in SEQ ID NO:437.
289. The AAV vector of any of claims 284, 286, and 287, wherein the bGH site comprises the sequence set forth in SEQ ID NO:604.
290. The AAV vector of any of claims 264-271, 273, 274, 280-285, and 288, wherein the first nucleic acid further comprises a woodchuck hepatitis virus post-transcriptional regulatory element (WPRE) in proximal to the SpA site, optionally wherein the WPRE is located between the sequence encoding the fusion protein and the SpA site.
291. The AAV vector of any of claims 264-271, 273, 274, 280-286, and 288-290, wherein the gRNA is capable of complexing with the dCas protein.
292. The AAV vector of any of claims 264-271, 273, 274, 280-286, and 288-291, wherein the gRNA comprises a gRNA spacer sequence that is capable of hybridizing to the target site or is complementary to the target site.
293. The AAV vector of any of claims 264-271, 273, 274, 280-286, and 288-292, wherein the dCas protein is a Staphylococcus aureus dCas9 (dSaCas9) protein or a Streptococcus pyogenes dCas9 (dSpCas9) protein.
294. The AAV vector of any of claims 264-271, 273, 274, 280-286, and 288-293, wherein the dCas protein is a Staphylococcus aureus dCas9 protein (dSaCas9) that comprises at least one amino acid mutation selected from D10A and N580A, with reference to numbering of positions of SEQ ID NO:73,
and/or the dCas protein comprises the sequence set forth in SEQ ID NO:72, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
295. The AAV vector of any of claims 264-271, 273, 274, 280-286, and 288-294, wherein the dCas is a Streptococcus pyogenes dCas9 (dSpCas9) protein that comprises at least one amino acid mutation selected from D10A and H840A, with reference to numbering of positions of SEQ ID NO:79, and/or the dCas protein comprises the sequence set forth in SEQ ID NO:78, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
296. The AAV vector of any of claims 264-295, wherein the regulatory DNA element is an enhancer.
297. The AAV vector of any of claims 264-296, wherein the target site is located within a target region spanning the genomic coordinates chr9:69, 027, 282-69, 028, 497 from hg38 (SEQ ID NO:431), optionally wherein the target site is located within a target region spanning the genomic coordinates chr9:69, 027, 615-69, 028, 101 from hg38, optionally wherein the target site is located within a target region spanning the genomic coordinates chr9:69, 027, 825-69, 027, 875.
298. The AAV vector of any of claims 264-271, 273, 274, 280-286, and 288-297, wherein the target site comprises the sequence set forth in SEQ ID NO: 21, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing.
299. The AAV vector of any of claims 264-271, 273, 274, 280-286, and 288-298, wherein the gRNA comprises a gRNA spacer sequence comprising the sequence set forth in SEQ ID NO:42, or a contiguous portion thereof of at least 14 nt.
300. The AAV vector of any of claims 264-271, 273, 274, 280-286, and 288-299, wherein the gRNA further comprises the sequence set forth in SEQ ID NO:44, optinally wherein the gRNA comprises the sequence set forth in SEQ ID NO:67, optionally wherein the gRNA is the gRNA sequence set forth in SEQ ID NO:67.
301. The AAV vector of any of claims 272-281, 284, 285, 287-289, 296, and 297, wherein the target site comprises the sequence set forth in any one of SEQ ID NOS: 272 and 277, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing.
302. The AAV vector of any of claims 272-281, 284, 285, 287-289, 296, and 297, wherein the eZFP comprises the sequence set forth in any one of SEQ ID NOS: 301 and 302.
303. The AAV vector of any of claims 264-295, wherein the regulatory DNA element is a promoter.
304. The AAV vector of any of claims 264-295 and 303, wherein the target site is within a target region spanning the genomic coordinates chr9:69, 034, 900-69, 035, 900 from hg38 (SEQ ID NO:430), optionally wherein the target site is within a target region spanning the genomic coordinates chr9:69, 035, 300-69-035, 800 from hg38; chr9:69, 035, 350-69, 035, 450 from hg38; or chr9:69,035,675- 69,035,725.
305. The AAV vector of any of claims 264-271, 273, 274, 280-286, 288-295, 303, and 304, wherein the target site comprises a sequence selected from any of SEQ ID NOS: 1-10, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing.
306. The AAV vector of any of claims 264-271, 273, 274, 280-286, 288-295, and 303-305, wherein the gRNA comprises a gRNA spacer sequence comprising a sequence selected from any of SEQ ID NOS:22-31, or a contiguous portion thereof of at least 14 nt.
307. The AAV vector of any of claims 264-271, 273, 274, 280-286, 288-295, and 303-306, wherein the gRNA comprises a gRNA spacer sequence comprising SEQ ID NO:22, or a contiguous portion thereof of at least 14 nt.
308. The AAV vector of any of claims 264-271, 273, 274, 280-286, 288-295, and 303-306, wherein the gRNA comprises a gRNA spacer sequence comprising SEQ ID NO:28, or a contiguous portion thereof of at least 14 nt.
309. The AAV vector of any of claims 264-271, 273, 274, 280-286, 288-295, 303, and 304, wherein the gRNA further comprises the sequence set forth in SEQ ID NO:44, optionally wherein the gRNA comprises a sequence selected from any of SEQ ID NOS:47-56, optionally wherein the gRNA is the gRNA sequence set forth in any of SEQ ID NOS:47-56, optionally wherein the gRNA is set forth in SEQ ID NO:47 or 53.
310. The AAV vector of any of claims 264-271, 273, 274, 280-286, 288-295, 303, and 304, wherein the target site comprises a sequence selected from any of SEQ ID NOS: 11-20, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing.
311. The AAV vector of any of claims 264-271, 273, 274, 280-286, 288-295, 303, 304, and 310, wherein the gRNA comprises a gRNA spacer sequence comprising a sequence selected from any of SEQ ID NOS:32-41, or a contiguous portion thereof of at least 14 nt.
312. The AAV vector of any of claims 264-271, 273, 274, 280-286, 288-295, 303, 304, 310 and 311, wherein the gRNA further comprises the sequence set forth in SEQ ID NO:46, and/or wherein the gRNA comprises a sequence selected from any of SEQ ID NOS:57-66, optionally wherein the gRNA is the gRNA set forth in any of SEQ ID NOS:57-66.
313. The AAV vector of any of claims 264-271, 273, 274, 280-286, 288-300, and 303-312, wherein the gRNA spacer sequence is between 14 nt and 24 nt, or between 16 nt and 22 nt in length, optionally wherein the gRNA spacer sequence is 18 nt, 19 nt, 20 nt, 21 nt or 22 nt in length.
314. The AAV vector of any of claims 272-281, 284, 285, 287-289, 303, and 304, wherein the target site comprises the sequence set forth in any one of SEQ ID NOS: 280-283, 290, 299, and 583-600, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing.
315. The AAV vector of any of claims 272-281, 284, 285, 287-289, 303, 304, and 314, wherein the target site comprises the sequence set forth in any one of SEQ ID NOS: 299, 587, 589, and 591, a contiguous portion thereof of at least 14 nt, or a complementary sequence of any of the foregoing.
316. The AAV vector of any of claims 272-281, 284, 285, 287-289, 303, 304, 314, and 315, wherein the target site comprises the sequence set forth in SEQ ID NO: 299, a contiguous portion thereof of at least 14 nt, or a complementary sequence of the sequence set forth in SEQ ID NO: 299.
317. The AAV vector of any of claims 272-281, 284, 285, 287-289, 303, 304, 314, and 315, wherein the target site comprises the sequence set forth in SEQ ID NO: 587, a contiguous portion thereof of at least 14 nt, or a complementary sequence of the sequence set forth in SEQ ID NO: 587.
318. The AAV vector of any of claims 272-281, 284, 285, 287-289, 303, 304, 314, and 315, wherein the target site comprises the sequence set forth in SEQ ID NO: 589, a contiguous portion thereof of at least 14 nt, or a complementary sequence of the sequence set forth in SEQ ID NO: 589.
319. The AAV vector of any of claims 272-281, 284, 285, 287-289, 303, 304, 314, and 315, wherein the target site comprises the sequence set forth in SEQ ID NO: 591, a contiguous portion thereof of at least 14 nt, or a complementary sequence of the sequence set forth in SEQ ID NO: 591.
320. The AAV vector of any of claims 272-281, 284, 285, 287-289, 303, 304, and 314, wherein the eZFP comprises the sequence set forth in any one of SEQ ID NOS: 303-307 and 439-456.
321. The AAV vector of any of claims 272-281, 284, 285, 287-289, 303, 304, 314, 315, and 320, wherein the eZFP comprises the sequence set forth in any one of SEQ ID NOS: 307, 441, 443, and 445.
322. The AAV vector of any of claims 272-281, 284, 285, 287-289, 303, 304, 314, 315, 320, and 321, wherein the eZFP comprises the sequence set forth in SEQ ID NO: 307.
323. The AAV vector of any of claims 272-281, 284, 285, 287-289, 303, 304, 314, 315, 320, and 321, wherein the eZFP comprises the sequence set forth in SEQ ID NO: 441.
324. The AAV vector of any of claims 272-281, 284, 285, 287-289, 303, 304, 314, 315, 320, and 321, wherein the eZFP comprises the sequence set forth in SEQ ID NO: 443.
325. The AAV vector of any of claims 272-281, 284, 285, 287-289, 303, 304, 314, 315, 320, and 321, wherein the eZFP comprises the sequence set forth in SEQ ID NO: 445.
326. The AAV vector of any of claims 264-325, wherein the at least one effector domain induces transcription activation.
327. The AAV vector of any of claims 264-326, wherein the at least one epigenetic effector domain comprises: a VP64 domain, a p65 activation domain, a p300 domain, an Rta domain, a CBP domain, a VPR domain, a VPH domain, an HSF1 domain, a TET protein domain, optionally wherein the TET protein is TET1, a SunTag domain, a domain from DPOLA, ENL, FOXO3, HSH2D, NCOA2, NCOA3, PSA1, PYGO1, RBM39, HERC2, or NOTCH2, or a domain, portion, variant, or truncation of any of the foregoing.
328. The AAV vector of any of claims 264-327, wherein the at least one epigenetic effector domain comprises the sequence set forth in any of SEQ ID NOS:81, 83, 100-109, 111-122, 124, 125, 134-140, 152, and 383-396, or a domain, portion, variant, or truncation thereof, or an amino acid
sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
329. The AAV vector of any of claims 264-328 wherein the at least one effector domain is fused to the N-terminus, the C-terminus, or both the N-terminus and the C-terminus, of the dCas protein or eZFP.
330. The AAV vector of any of claims 264-329, further comprising one or more linkers connecting the dCas protein or eZFP to the at least one effector domain, and/or further comprising one or more nuclear localization signals (NLS).
331. The AAV vector of any of claims 264-330 wherein the at least one effector domain comprises at least one VP16 domain, or a VP16 tetramer (“VP64”) or a variant thereof.
332. The AAV vector of any of claims 264-331, wherein the at least one effector domain comprises VP64.
333. The AAV vector of any of claims 264-332, wherein the at least one effector domain comprises a VP64 domain comprising the sequence set forth in SEQ ID NO:81 or 83, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
334. The AAV vector of any of claims 264-333, wherein the at least one effector domain comprises the sequence set forth in SEQ ID NO:81 or 83.
335. The AAV vector of any of claims 264-334, wherein the at least one epigenetic effector domain comprises: a domain from DPOLA, ENL, FOXO3, HSH2D, NCOA2, NCOA3, PSA1, PYGO1, RBM39, HERC2, or NOTCH2, or a domain, portion, variant, or truncation of any of the foregoing.
336. The AAV vector of any of claims 264-335, wherein the at least one epigenetic effector domain comprises the sequence set forth in any of SEQ ID NOS:383-393, or a domain, portion, or variant thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
337. The AAV vector of any of claims 264-336, wherein the at least one effector domain comprises a domain from NCOA2, NCOA3, FOXO3, PYGO1, or a portion or variant of any of the foregoing.
338. The AAV vector of any of claims 264-337, wherein each effector domain of the at least one effector domain is independently selected from an NCOA2 domain, an NCOA3 domain, a FOXO3 domain, and a PYGO1 domain.
339. The AAV vector of any of claims 264-338, wherein the at least one effector domain comprises a domain from NCOA2 comprising the sequence set forth in SEQ ID NO: 104 or SEQ ID NO:387, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
340. The AAV vector of any of claims 264-339, wherein the at least one effector domain comprises a domain from NCOA2 set forth in or SEQ ID NO:387.
341. The AAV vector of any of claims 264-340, wherein the at least one effector domain comprises a domain from NCOA3 comprising the sequence set forth in SEQ ID NO: 105 or SEQ ID NO:388, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%,
95%, 96%, 97%, 98%, or 99% sequence identity thereto.
342. The AAV vector of any of claims 264-341, wherein the at least one effector domain comprises a domain from NC0A3 set forth in or SEQ ID NO:388.
343. The AAV vector of any of claims 264-342, wherein the at least one effector domain comprises a domain from F0X03 comprising the sequence set forth in SEQ ID NO: 102 or SEQ ID NO:385, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
344. The AAV vector of any of claims 264-343, wherein the at least one effector domain comprises a domain from FOXO3 set forth in or SEQ ID NO:385.
345. The AAV vector of any of claims 264-344, wherein the at least one effector domain comprises a domain from PYGO1 comprising the sequence set forth in SEQ ID NO: 107 or SEQ ID NO:390, or a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
346. The AAV vector of any of claims 264-345, wherein the at least one effector domain comprises a domain from PYGO1 set forth in or SEQ ID NO:390.
347. The AAV vector of any of claims 264-346, wherein the at least one effector domain is a multipartite effector composed of at least two effector domains.
348. The AAV vector of claim 347, wherein the multipartite effector is composed of two effector domains.
349. The AAV vector of claim 348, wherein the multipartite effector is composed of three effector domains.
350. The AAV vector of any of claims 347-349, wherein the multipartite effector is set forth in any of SEQ ID NOS:397-418, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
351. The AAV vector of any of claims 347-350, wherein the multipartite effector is set forth in any of SEQ ID NOS:411-418, a portion thereof, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any of the foregoing.
352. The AAV vector of any of claims 347-351, wherein the multipartite effector comprises, in the N-terminal to C-terminal direction, domains from FOXO3, FOXO3, and NCOA3.
353. The AAV vector of claim 352, wherein the multipartite effector comprises the sequence set forth in SEQ ID NO:415, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
354. The AAV vector of any of claims 347-351, wherein the multipartite effector comprises, in the N-terminal to C-terminal direction, domains from NCOA3, FOXO3, and FOXO3.
355. The AAV vector of claim 354, wherein the multipartite effector comprises the sequence set forth in SEQ ID NO:418, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
356. The AAV vector of any of claims 347-351, wherein the multipartite effector comprises, in the N-terminal to C-terminal direction, domains from NCOA3, FOXO3, and NCOA3.
357. The AAV vector of claim 356, wherein the multipartite effector comprises the sequence set forth in SEQ ID NO:413, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
358. The AAV vector of any of claims 347-351, wherein the multipartite effector comprises, in the N-terminal to C-terminal direction, domains from NCOA2, FOXO3, and NCOA3.
359. The AAV vector of claim 358, wherein the multipartite effector comprises the sequence set forth in SEQ ID NO:416, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
360. The AAV vector of any of claims 347-351, wherein the multipartite effector comprises, in the N-terminal to C-terminal direction, domains from PYGO1, FOXO3, and NCOA3.
361. The AAV vector of claim 360, wherein the multipartite effector comprises the sequence set forth in SEQ ID NO:411, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
362. The AAV vector of any of claims 264-361, wherein the at least one epigenetic effector domain is fused to the N-terminus of the dCas protein or eZFP.
363. The AAV vector of any of claims 264-361, wherein the at least one epigenetic effector domain is fused to the C-terminus of the dCas protein or eZFP.
364. The AAV vector of any of claims 264-361, wherein the at least one epigenetic effector domain is fused to both the N-terminus and the C-terminus, of the dCas protein or eZFP.
365. The AAV vector of any of claims 264-361, wherein the one or more linkers are in between any two of the components of the fusion protein, including the dCas protein or eZFP, any of the at least one effector domains, and the one or more NLS.
366. The AAV vector of any of claims 264-365, wherein the one or more linkers connect the dCas protein or eZFP and the at least one epigenetic effector domain.
367. The AAV vector of any of claims 264-366, wherein the fusion protein comprises the sequence set forth in SEQ ID NO:71 or 77, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto, or the sequence set forth in SEQ ID NO:71 or 77, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
368. The AAV vector of any of claims 264-367, wherein the fusion protein comprises the sequence set forth in any one of SEQ ID NOS:266-268 and 315-319, or an amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity thereto.
369. The AAV vector of any of claims 264-368, wherein the AAV vector is selected from the group consisting of: AAV1, AAV2, AAV3, AAV4, AAV5, AAV6, AAV7, AAV8, AAV9, AAV10, AAV11, AAV 12, AAV-DJ, and AAVrh74.
370. The AAV vector of any of claims 264-369, wherein the AAV vector is AAV6.
371. The AAV vector of any of claims 264-369, wherein the AAV vector is AAV9.
372. The AAV vector of any of claims 264-369, wherein the AAV vector is AAV-DJ.
373. The AAV vector of any of claims 264-369, wherein the AAV vector is AAVrh74.
374. The AAV vector of any of claims 264-373, wherein the vector exhibits tropism for a nervous system cell, optionally a neuron, a heart cell, optionally a cardiomyocyte, a skeletal muscle cell, a fibroblast, an induced pluripotent stem cell, and/or a cell derived from any of the foregoing, or for a combination of any of the foregoing cells.
375. The AAV vector of any of claims 264-374, wherein the vector exhibits tropism for induced pluripotent stem cells.
376. The AAV vector of any of claims 264-375, wherein the vector exhibits tropism for neurons and cardiomyocytes.
377. A cell comprising the eZFP of any of claims 1-187, the fusion protein of any of claims 186-246, the polynucleotide of claim 247, the plurality of polynucleotides of claim 248, the vector of any of claims 249-263, the AAV vector of any of claims 264-376, or a portion or component of any of the foregoing, or a combination of any of the foregoing.
378. The cell of claim 377, wherein the cell is a nervous system cell, optionally a neuron, a heart cell, optionally a cardiomyocyte, a skeletal muscle cell, a fibroblast, an induced pluripotent stem cell, and/or a cell derived from any of the foregoing.
379. The cell of claim 377 or 378, wherein the cell is from a subject that has or is suspected of having Friedreich’s ataxia (FA).
380. A pharmaceutical composition comprising the eZFP of any of claims 1-187, the fusion protein of any of claims 188-246, the polynucleotide of claim 247, the plurality of polynucleotides of claim 248, the vector of any of claims 249-263, the AAV vector of any of claims 264-376, or a portion or component of any of the foregoing, or a combination of any of the foregoing.
381. The pharmaceutical composition of claim 380, for use in treating a disease, condition, or disorder in a subject.
382. The pharmaceutical composition of claim 381, wherein the disease, condition, or disorder is Friedreich’s ataxia and/or a GAA trinucleotide repeat expansion in the FXN locus.
383. The pharmaceutical composition of claim 381 or 382, wherein following administration of the pharmaceutical composition, the expression of FXN is increased in cells of the subject.
384. A method for increasing the expression of FXN in a cell, the method comprising introducing into the cell: the eZFP of any of claims 1-187, the fusion protein of any of claims 188-246, the polynucleotide of claim 247, the plurality of polynucleotides of claim 248, the vector of any of claims
249-263, the AAV vector of any of claims 264-376, the pharmaceutical composition of any of claims 380-383, or a portion or component of any of the foregoing, or a combination of any of the foregoing.
385. The method of claim 384, wherein the cell is from and/or in a subject that has or is suspected of having Friedreich’s ataxia.
386. The method of claim 384 or 385, wherein the cell exhibits reduced expression of FXN in comparison to a reference cell from an individual not having Friedreich’s ataxia and/or a GA A trinucleotide expansion in the FXN locus.
387. A method for increasing the expression of FXN in a cell in a subject, the method comprising administering to the subject: the eZFP of any of claims 1-187, the fusion protein of any of claims 188-246, the polynucleotide of claim 247, the plurality of polynucleotides of claim 248, the vector of any of claims 249-263, the AAV vector of any of claims 264-376, the pharmaceutical composition of any of claims 380-383, or a portion or component of any of the foregoing, or a combination of any of the foregoing.
388. A method of treating a subject in need thereof, the method comprising administering to the subject: the eZFP of any of claims 1-187, the fusion protein of any of claims 188-246, the polynucleotide of claim 247, the plurality of polynucleotides of claim 248, the vector of any of claims 249-263, the AAV vector of any of claims 264-376, the pharmaceutical composition of any of claims 380-383, or a portion or component of any of the foregoing, or a combination of any of the foregoing.
389. The method of claim 387 or 388, wherein the subject has or is suspected of having Friedreich’s ataxia, and/or a GAA trinucleotide expansion in the FXN locus.
390. The method of any of claims 384-389, wherein the introducing or administering is carried out in vivo or ex vivo.
391. The method of any of claims 384-390, wherein the cell and/or subject exhibits reduced expression of FXN prior to performing the method.
392. The method of claim 391, wherein the reduced expression of FXN is reduced in comparison to a reference individual not having Friedreich’ s ataxia and/or a GAA trinucleotide repeat expansion in the FXN locus, and/or a reference cell therefrom.
393. The method of any of claims 386 and 389-392, wherein the GAA trinucleotide repeat expansion is in a first intron of a FXN gene, and comprises at least 50, at least 60, at least 70, at least 80, at least 90, at least 100, at least 150, at least 200, at least 300, at least 400, at least 500, at least 600, at least 700, at least 800, at least 900, at least 1000, or more repeated GAA trinucleotides.
394. The method of any of claims 384-393, wherein following the introducing or administering, the expression of FXN is increased in the cell and/or subject.
395. The method of claim 394, wherein the expression of FXN is increased in the cell or cells of the subject by at least about 1.2-fold, 1.25-fold, 1.3-fold, 1.4-fold, 1.5-fold, 1.6-fold, 1.7-fold, 1.75- fold, 1.8-fold, 1.9-fold, 2-fold, 2.5-fold, 3-fold, 4-fold, or 5-fold; and/or the expression is increased by less than about 10-fold, 9-fold, 8-fold, 7-fold or 6-fold.
396. The method of claim 394 or 395, wherein the expression of FXN is increased in the cell or cells of the subject to a level that is at least at or about 30%, 40%, 50%, 60%, 70%, 75%, 80%, 85%, 90%, 95%, 100%, 105%, 110%, 120%, 125%, 150%, 175%, 200%, 225%, 250%, 300%, 400%, or 500%, of the expression level of FXN in a reference cell from an individual not having Friedreich’s ataxia and/or a cell not having a GAA trinucleotide repeat expansion in the FXN gene.
397. The method of any of claims 394-396, wherein the expression of FXN is increased in the cell or cells of the subject to a level that is less than at or about 200%, 300%, 400%, 500%, 600%, 700%, 800%, 900%, or 1000% of the expression level of FXN in a reference cell from an individual not having Friedreich’ s ataxia and/or a cell not having a GAA trinucleotide repeat expansion in the FXN gene.
398. The method of any of claims 394-397, wherein the expression is measured by the amount of mRNA encoding the FXN protein, and/or the amount of FXN protein.
399. The method of any of claims 385-398, wherein the subject is a human.
400. A cell comprising an epigenetic modification produced by the method of any of claims 384-399.
Applications Claiming Priority (4)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| US202363442756P | 2023-02-01 | 2023-02-01 | |
| US63/442,756 | 2023-02-01 | ||
| US202463621993P | 2024-01-17 | 2024-01-17 | |
| US63/621,993 | 2024-01-17 |
Publications (2)
| Publication Number | Publication Date |
|---|---|
| WO2024163678A2 true WO2024163678A2 (en) | 2024-08-08 |
| WO2024163678A3 WO2024163678A3 (en) | 2024-09-12 |
Family
ID=90364060
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/US2024/013874 Ceased WO2024163678A2 (en) | 2023-02-01 | 2024-01-31 | Fusion proteins and systems for targeted activation of frataxin (fxn) and related methods |
Country Status (1)
| Country | Link |
|---|---|
| WO (1) | WO2024163678A2 (en) |
Cited By (1)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO2025022013A3 (en) * | 2023-07-26 | 2025-04-10 | Imperial College Innovations Ltd | Zinc finger peptides, encoded nucleic acids, methods and uses |
Citations (61)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US4737323A (en) | 1986-02-13 | 1988-04-12 | Liposome Technology, Inc. | Liposome extrusion method |
| US5219740A (en) | 1987-02-13 | 1993-06-15 | Fred Hutchinson Cancer Research Center | Retroviral gene transfer into diploid fibroblasts for gene therapy |
| WO1998053058A1 (en) | 1997-05-23 | 1998-11-26 | Gendaq Limited | Nucleic acid binding proteins |
| WO1998053059A1 (en) | 1997-05-23 | 1998-11-26 | Medical Research Council | Nucleic acid binding proteins |
| US6140081A (en) | 1998-10-16 | 2000-10-31 | The Scripps Research Institute | Zinc finger binding domains for GNN |
| US6207453B1 (en) | 1996-03-06 | 2001-03-27 | Medigene Ag | Recombinant AAV vector-based transduction system and use of same |
| WO2002016536A1 (en) | 2000-08-23 | 2002-02-28 | Kao Corporation | Bactericidal antifouling detergent for hard surface |
| US6453242B1 (en) | 1999-01-12 | 2002-09-17 | Sangamo Biosciences, Inc. | Selection of sites for targeting by zinc finger proteins and methods of designing zinc finger proteins to bind to preselected sites |
| WO2003016496A2 (en) | 2001-08-20 | 2003-02-27 | The Scripps Research Institute | Zinc finger binding domains for cnn |
| US6534261B1 (en) | 1999-01-12 | 2003-03-18 | Sangamo Biosciences, Inc. | Regulation of endogenous gene expression in cells using zinc finger proteins |
| US6566118B1 (en) | 1997-09-05 | 2003-05-20 | Targeted Genetics Corporation | Methods for generating high titer helper-free preparations of released recombinant AAV vectors |
| WO2003042397A2 (en) | 2001-11-13 | 2003-05-22 | The Trustees Of The University Of Pennsylvania | A method of detecting and/or identifying adeno-associated virus (aav) sequences and isolating novel sequences identified thereby |
| US6596535B1 (en) | 1999-08-09 | 2003-07-22 | Targeted Genetics Corporation | Metabolically activated recombinant viral vectors and methods for the preparation and use |
| US6723551B2 (en) | 2001-11-09 | 2004-04-20 | The United States Of America As Represented By The Department Of Health And Human Services | Production of adeno-associated virus in insect cells |
| US20040142025A1 (en) | 2002-06-28 | 2004-07-22 | Protiva Biotherapeutics Ltd. | Liposomal apparatus and manufacturing methods |
| US7074596B2 (en) | 2002-03-25 | 2006-07-11 | Board Of Supervisors Of Louisiana State University And Agricultural And Mechanical College | Synthesis and use of anti-reverse mRNA cap analogues |
| US20070042031A1 (en) | 2005-07-27 | 2007-02-22 | Protiva Biotherapeutics, Inc. | Systems and methods for manufacturing liposomes |
| US7745651B2 (en) | 2004-06-07 | 2010-06-29 | Protiva Biotherapeutics, Inc. | Cationic lipids and methods of use |
| US7765583B2 (en) | 2005-02-28 | 2010-07-27 | France Telecom | System and method for managing virtual user domains |
| US7790154B2 (en) | 2000-06-01 | 2010-09-07 | The University Of North Carolina At Chapel Hill | Duplexed parvovirus vectors |
| US7799565B2 (en) | 2004-06-07 | 2010-09-21 | Protiva Biotherapeutics, Inc. | Lipid encapsulated interfering RNA |
| WO2010144740A1 (en) | 2009-06-10 | 2010-12-16 | Alnylam Pharmaceuticals, Inc. | Improved lipid formulation |
| US20120066783A1 (en) | 2006-03-30 | 2012-03-15 | The Board Of Trustees Of The Leland Stanford Junior University | Aav capsid library and aav capsid proteins |
| US20120164106A1 (en) | 2010-10-06 | 2012-06-28 | Schaffer David V | Adeno-associated virus virions with variant capsid and methods of use thereof |
| US8278036B2 (en) | 2005-08-23 | 2012-10-02 | The Trustees Of The University Of Pennsylvania | RNA containing modified nucleosides and methods of use thereof |
| US8283151B2 (en) | 2005-04-29 | 2012-10-09 | The United States Of America, As Represented By The Secretary, Department Of Health And Human Services | Isolation, cloning and characterization of new adeno-associated virus (AAV) serotypes |
| US8586526B2 (en) | 2010-05-17 | 2013-11-19 | Sangamo Biosciences, Inc. | DNA-binding proteins and uses thereof |
| WO2013171772A1 (en) | 2012-05-17 | 2013-11-21 | Vass Technologies S.R.L. | Modular-based, concrete floor or roofing building structure |
| WO2013176772A1 (en) | 2012-05-25 | 2013-11-28 | The Regents Of The University Of California | Methods and compositions for rna-directed target dna modification and for rna-directed modulation of transcription |
| US20130323226A1 (en) | 2011-02-17 | 2013-12-05 | The Trustees Of The University Of Pennsylvania | Compositions and Methods for Altering Tissue Specificity and Improving AAV9-Mediated Gene Transfer |
| WO2014093655A2 (en) | 2012-12-12 | 2014-06-19 | The Broad Institute, Inc. | Engineering and optimization of systems, methods and compositions for sequence manipulation with functional domains |
| WO2014093661A2 (en) | 2012-12-12 | 2014-06-19 | The Broad Institute, Inc. | Crispr-cas systems and methods for altering expression of gene products |
| WO2014152432A2 (en) | 2013-03-15 | 2014-09-25 | The General Hospital Corporation | Rna-guided targeting of genetic and epigenomic regulatory proteins to specific genomic loci |
| WO2014191128A1 (en) | 2013-05-29 | 2014-12-04 | Cellectis | Methods for engineering t cells for immunotherapy by using rna-guided cas nuclease system |
| WO2014197748A2 (en) | 2013-06-05 | 2014-12-11 | Duke University | Rna-guided gene editing and gene regulation |
| WO2015035136A2 (en) | 2013-09-06 | 2015-03-12 | President And Fellows Of Harvard College | Delivery system for functional nucleases |
| WO2015089427A1 (en) | 2013-12-12 | 2015-06-18 | The Broad Institute Inc. | Crispr-cas systems and methods for altering expression of gene products, structural information and inducible modular cas enzymes |
| US9139554B2 (en) | 2008-10-09 | 2015-09-22 | Tekmira Pharmaceuticals Corporation | Amino lipids and methods for the delivery of nucleic acids |
| WO2015161276A2 (en) | 2014-04-18 | 2015-10-22 | Editas Medicine, Inc. | Crispr-cas-related methods, compositions and components for cancer immunotherapy |
| WO2016011070A2 (en) | 2014-07-14 | 2016-01-21 | The Regents Of The University Of California | A protein tagging system for in vivo single molecule imaging and control of gene transcription |
| WO2016049258A2 (en) | 2014-09-25 | 2016-03-31 | The Broad Institute Inc. | Functional screening with optimized functional crispr-cas systems |
| US20160097061A1 (en) | 2012-05-04 | 2016-04-07 | Novartis Ag | Viral vectors for the treatment of retinal dystrophy |
| WO2016114972A1 (en) | 2015-01-12 | 2016-07-21 | The Regents Of The University Of California | Heterodimeric cas9 and methods of use thereof |
| WO2016123578A1 (en) | 2015-01-30 | 2016-08-04 | The Regents Of The University Of California | Protein delivery in primary hematopoietic cells |
| WO2016130600A2 (en) | 2015-02-09 | 2016-08-18 | Duke University | Compositions and methods for epigenome editing |
| US9458205B2 (en) | 2011-11-16 | 2016-10-04 | Sangamo Biosciences, Inc. | Modified DNA-binding proteins and uses thereof |
| WO2017093969A1 (en) | 2015-12-04 | 2017-06-08 | Novartis Ag | Compositions and methods for immunooncology |
| WO2017180915A2 (en) | 2016-04-13 | 2017-10-19 | Duke University | Crispr/cas9-based repressors for silencing gene targets in vivo and methods of use |
| WO2017189308A1 (en) | 2016-04-19 | 2017-11-02 | The Broad Institute Inc. | Novel crispr enzymes and systems |
| WO2017193107A2 (en) | 2016-05-06 | 2017-11-09 | Juno Therapeutics, Inc. | Genetically engineered cells and methods of making the same |
| WO2017197238A1 (en) | 2016-05-12 | 2017-11-16 | President And Fellows Of Harvard College | Aav split cas9 genome editing and transcriptional regulation |
| WO2019232069A1 (en) | 2018-05-30 | 2019-12-05 | Emerson Collective Investments, Llc | Cell therapy |
| WO2020051561A1 (en) | 2018-09-07 | 2020-03-12 | Beam Therapeutics Inc. | Compositions and methods for delivering a nucleobase editing system |
| WO2020113034A1 (en) | 2018-11-30 | 2020-06-04 | Avexis, Inc. | Aav viral vectors and uses thereof |
| US10723692B2 (en) | 2014-06-25 | 2020-07-28 | Acuitas Therapeutics, Inc. | Lipids and lipid nanoparticle formulations for delivery of nucleic acids |
| US10941395B2 (en) | 2014-06-10 | 2021-03-09 | Massachusetts Institute Of Technology | Method for gene editing |
| WO2021076744A1 (en) | 2019-10-15 | 2021-04-22 | The Regents Of The University Of California | Gene targets for manipulating t cell behavior |
| US20210317474A1 (en) | 2017-11-08 | 2021-10-14 | Novartis Ag | Means and method for producing and purifying viral vectors |
| WO2021226077A2 (en) | 2020-05-04 | 2021-11-11 | The Board Of Trustees Of The Leland Stanford Junior University | Compositions, systems, and methods for the generation, identification, and characterization of effector domains for activating and silencing gene expression |
| WO2021226555A2 (en) | 2020-05-08 | 2021-11-11 | Duke University | Chromatin remodelers to enhance targeted gene activation |
| WO2021247570A2 (en) | 2020-06-02 | 2021-12-09 | The Regents Of The University Ofcalifornia | Compositions and methods for gene editing |
Family Cites Families (3)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| AU2015234204A1 (en) * | 2014-03-20 | 2016-10-06 | Universite Laval | CRISPR-based methods and products for increasing frataxin levels and uses thereof |
| EP3883954A4 (en) * | 2018-11-21 | 2022-08-10 | Stridebio, Inc. | RECOMBINANT VIRAL VECTORS AND NUCLEIC ACIDS FOR THEIR PRODUCTION |
| US20220348893A1 (en) * | 2019-09-23 | 2022-11-03 | Omega Therapeutics, Inc | Methods and compositions for modulating frataxin expression and treating friedrich's ataxia |
-
2024
- 2024-01-31 WO PCT/US2024/013874 patent/WO2024163678A2/en not_active Ceased
Patent Citations (69)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US4737323A (en) | 1986-02-13 | 1988-04-12 | Liposome Technology, Inc. | Liposome extrusion method |
| US5219740A (en) | 1987-02-13 | 1993-06-15 | Fred Hutchinson Cancer Research Center | Retroviral gene transfer into diploid fibroblasts for gene therapy |
| US6207453B1 (en) | 1996-03-06 | 2001-03-27 | Medigene Ag | Recombinant AAV vector-based transduction system and use of same |
| WO1998053059A1 (en) | 1997-05-23 | 1998-11-26 | Medical Research Council | Nucleic acid binding proteins |
| WO1998053060A1 (en) | 1997-05-23 | 1998-11-26 | Gendaq Limited | Nucleic acid binding proteins |
| WO1998053058A1 (en) | 1997-05-23 | 1998-11-26 | Gendaq Limited | Nucleic acid binding proteins |
| US6566118B1 (en) | 1997-09-05 | 2003-05-20 | Targeted Genetics Corporation | Methods for generating high titer helper-free preparations of released recombinant AAV vectors |
| US6140081A (en) | 1998-10-16 | 2000-10-31 | The Scripps Research Institute | Zinc finger binding domains for GNN |
| US6534261B1 (en) | 1999-01-12 | 2003-03-18 | Sangamo Biosciences, Inc. | Regulation of endogenous gene expression in cells using zinc finger proteins |
| US6453242B1 (en) | 1999-01-12 | 2002-09-17 | Sangamo Biosciences, Inc. | Selection of sites for targeting by zinc finger proteins and methods of designing zinc finger proteins to bind to preselected sites |
| US7846729B2 (en) | 1999-08-09 | 2010-12-07 | Genzyme Corporation | Metabolically activated recombinant viral vectors and methods for their preparation and use |
| US6596535B1 (en) | 1999-08-09 | 2003-07-22 | Targeted Genetics Corporation | Metabolically activated recombinant viral vectors and methods for the preparation and use |
| US8093054B2 (en) | 1999-08-09 | 2012-01-10 | Genzyme Corporation | Metabolically activated recombinant viral vectors and methods for their preparation and use |
| US7785888B2 (en) | 1999-08-09 | 2010-08-31 | Genzyme Corporation | Metabolically activated recombinant viral vectors and methods for their preparation and use |
| US7125717B2 (en) | 1999-08-09 | 2006-10-24 | Targeted Genetics Corporation | Metabolically activated recombinant viral vectors and methods for their preparation and use |
| US8361457B2 (en) | 2000-06-01 | 2013-01-29 | The University Of North Carolina At Chapel Hill | Duplexed parvovirus vectors |
| US7790154B2 (en) | 2000-06-01 | 2010-09-07 | The University Of North Carolina At Chapel Hill | Duplexed parvovirus vectors |
| WO2002016536A1 (en) | 2000-08-23 | 2002-02-28 | Kao Corporation | Bactericidal antifouling detergent for hard surface |
| WO2003016496A2 (en) | 2001-08-20 | 2003-02-27 | The Scripps Research Institute | Zinc finger binding domains for cnn |
| US6723551B2 (en) | 2001-11-09 | 2004-04-20 | The United States Of America As Represented By The Department Of Health And Human Services | Production of adeno-associated virus in insect cells |
| WO2003042397A2 (en) | 2001-11-13 | 2003-05-22 | The Trustees Of The University Of Pennsylvania | A method of detecting and/or identifying adeno-associated virus (aav) sequences and isolating novel sequences identified thereby |
| US7074596B2 (en) | 2002-03-25 | 2006-07-11 | Board Of Supervisors Of Louisiana State University And Agricultural And Mechanical College | Synthesis and use of anti-reverse mRNA cap analogues |
| US20040142025A1 (en) | 2002-06-28 | 2004-07-22 | Protiva Biotherapeutics Ltd. | Liposomal apparatus and manufacturing methods |
| US7745651B2 (en) | 2004-06-07 | 2010-06-29 | Protiva Biotherapeutics, Inc. | Cationic lipids and methods of use |
| US7799565B2 (en) | 2004-06-07 | 2010-09-21 | Protiva Biotherapeutics, Inc. | Lipid encapsulated interfering RNA |
| US7765583B2 (en) | 2005-02-28 | 2010-07-27 | France Telecom | System and method for managing virtual user domains |
| US8283151B2 (en) | 2005-04-29 | 2012-10-09 | The United States Of America, As Represented By The Secretary, Department Of Health And Human Services | Isolation, cloning and characterization of new adeno-associated virus (AAV) serotypes |
| US20070042031A1 (en) | 2005-07-27 | 2007-02-22 | Protiva Biotherapeutics, Inc. | Systems and methods for manufacturing liposomes |
| US8278036B2 (en) | 2005-08-23 | 2012-10-02 | The Trustees Of The University Of Pennsylvania | RNA containing modified nucleosides and methods of use thereof |
| US20120066783A1 (en) | 2006-03-30 | 2012-03-15 | The Board Of Trustees Of The Leland Stanford Junior University | Aav capsid library and aav capsid proteins |
| US9139554B2 (en) | 2008-10-09 | 2015-09-22 | Tekmira Pharmaceuticals Corporation | Amino lipids and methods for the delivery of nucleic acids |
| WO2010144740A1 (en) | 2009-06-10 | 2010-12-16 | Alnylam Pharmaceuticals, Inc. | Improved lipid formulation |
| US8586526B2 (en) | 2010-05-17 | 2013-11-19 | Sangamo Biosciences, Inc. | DNA-binding proteins and uses thereof |
| US20120164106A1 (en) | 2010-10-06 | 2012-06-28 | Schaffer David V | Adeno-associated virus virions with variant capsid and methods of use thereof |
| US20130323226A1 (en) | 2011-02-17 | 2013-12-05 | The Trustees Of The University Of Pennsylvania | Compositions and Methods for Altering Tissue Specificity and Improving AAV9-Mediated Gene Transfer |
| US9458205B2 (en) | 2011-11-16 | 2016-10-04 | Sangamo Biosciences, Inc. | Modified DNA-binding proteins and uses thereof |
| US20160097061A1 (en) | 2012-05-04 | 2016-04-07 | Novartis Ag | Viral vectors for the treatment of retinal dystrophy |
| WO2013171772A1 (en) | 2012-05-17 | 2013-11-21 | Vass Technologies S.R.L. | Modular-based, concrete floor or roofing building structure |
| WO2013176772A1 (en) | 2012-05-25 | 2013-11-28 | The Regents Of The University Of California | Methods and compositions for rna-directed target dna modification and for rna-directed modulation of transcription |
| WO2014093661A2 (en) | 2012-12-12 | 2014-06-19 | The Broad Institute, Inc. | Crispr-cas systems and methods for altering expression of gene products |
| WO2014093655A2 (en) | 2012-12-12 | 2014-06-19 | The Broad Institute, Inc. | Engineering and optimization of systems, methods and compositions for sequence manipulation with functional domains |
| WO2014152432A2 (en) | 2013-03-15 | 2014-09-25 | The General Hospital Corporation | Rna-guided targeting of genetic and epigenomic regulatory proteins to specific genomic loci |
| WO2014191128A1 (en) | 2013-05-29 | 2014-12-04 | Cellectis | Methods for engineering t cells for immunotherapy by using rna-guided cas nuclease system |
| WO2014197748A2 (en) | 2013-06-05 | 2014-12-11 | Duke University | Rna-guided gene editing and gene regulation |
| WO2015035136A2 (en) | 2013-09-06 | 2015-03-12 | President And Fellows Of Harvard College | Delivery system for functional nucleases |
| WO2015089427A1 (en) | 2013-12-12 | 2015-06-18 | The Broad Institute Inc. | Crispr-cas systems and methods for altering expression of gene products, structural information and inducible modular cas enzymes |
| WO2015161276A2 (en) | 2014-04-18 | 2015-10-22 | Editas Medicine, Inc. | Crispr-cas-related methods, compositions and components for cancer immunotherapy |
| US10941395B2 (en) | 2014-06-10 | 2021-03-09 | Massachusetts Institute Of Technology | Method for gene editing |
| US10723692B2 (en) | 2014-06-25 | 2020-07-28 | Acuitas Therapeutics, Inc. | Lipids and lipid nanoparticle formulations for delivery of nucleic acids |
| WO2016011070A2 (en) | 2014-07-14 | 2016-01-21 | The Regents Of The University Of California | A protein tagging system for in vivo single molecule imaging and control of gene transcription |
| WO2016049258A2 (en) | 2014-09-25 | 2016-03-31 | The Broad Institute Inc. | Functional screening with optimized functional crispr-cas systems |
| WO2016114972A1 (en) | 2015-01-12 | 2016-07-21 | The Regents Of The University Of California | Heterodimeric cas9 and methods of use thereof |
| WO2016123578A1 (en) | 2015-01-30 | 2016-08-04 | The Regents Of The University Of California | Protein delivery in primary hematopoietic cells |
| WO2016130600A2 (en) | 2015-02-09 | 2016-08-18 | Duke University | Compositions and methods for epigenome editing |
| WO2017093969A1 (en) | 2015-12-04 | 2017-06-08 | Novartis Ag | Compositions and methods for immunooncology |
| WO2017180915A2 (en) | 2016-04-13 | 2017-10-19 | Duke University | Crispr/cas9-based repressors for silencing gene targets in vivo and methods of use |
| WO2017189308A1 (en) | 2016-04-19 | 2017-11-02 | The Broad Institute Inc. | Novel crispr enzymes and systems |
| WO2017193107A2 (en) | 2016-05-06 | 2017-11-09 | Juno Therapeutics, Inc. | Genetically engineered cells and methods of making the same |
| WO2017197238A1 (en) | 2016-05-12 | 2017-11-16 | President And Fellows Of Harvard College | Aav split cas9 genome editing and transcriptional regulation |
| US20210317474A1 (en) | 2017-11-08 | 2021-10-14 | Novartis Ag | Means and method for producing and purifying viral vectors |
| WO2019232069A1 (en) | 2018-05-30 | 2019-12-05 | Emerson Collective Investments, Llc | Cell therapy |
| WO2020051561A1 (en) | 2018-09-07 | 2020-03-12 | Beam Therapeutics Inc. | Compositions and methods for delivering a nucleobase editing system |
| US20210301274A1 (en) | 2018-09-07 | 2021-09-30 | Beam Therapeutics Inc. | Compositions and Methods for Delivering a Nucleobase Editing System |
| WO2020113034A1 (en) | 2018-11-30 | 2020-06-04 | Avexis, Inc. | Aav viral vectors and uses thereof |
| US20220001028A1 (en) | 2018-11-30 | 2022-01-06 | Novartis Ag | Aav viral vectors and uses thereof |
| WO2021076744A1 (en) | 2019-10-15 | 2021-04-22 | The Regents Of The University Of California | Gene targets for manipulating t cell behavior |
| WO2021226077A2 (en) | 2020-05-04 | 2021-11-11 | The Board Of Trustees Of The Leland Stanford Junior University | Compositions, systems, and methods for the generation, identification, and characterization of effector domains for activating and silencing gene expression |
| WO2021226555A2 (en) | 2020-05-08 | 2021-11-11 | Duke University | Chromatin remodelers to enhance targeted gene activation |
| WO2021247570A2 (en) | 2020-06-02 | 2021-12-09 | The Regents Of The University Ofcalifornia | Compositions and methods for gene editing |
Non-Patent Citations (92)
| Title |
|---|
| "Biocomputing: Informatics and Genome Projects", 1993, ACADEMIC PRESS |
| "Computer Analysis of Sequence Data", 1994, HUMANA PRESS |
| "Remington's Pharmaceutical Sciences", 1980 |
| "Sequence Analysis in Molecular Biology", 1987, ACADEMIC PRESS |
| "Sequence Analysis Primer", 1991, M STOCKTON PRESS |
| "Uniprot", Database accession no. Q04206 |
| ADLI, M., NAT. COMMUN., vol. 9, 2018, pages 1911 |
| ALONSO-CAMINO ET AL., MOL THER NUCL ACIDS, vol. 2, 2013, pages e93 |
| BELBELLAA ET AL., MOLECULAR THERAPY - METHODS & CLINICAL DEVELOPMENT, vol. 19, 2020, pages 120 - 138 |
| BHAKTA M.S. ET AL., METHODS MOL. BIOL., vol. 649, 2010, pages 3 - 30 |
| BLOOMFIELD, ANN. REV. BIOPHYS. BIOENG., vol. 10, 1981, pages 421A150 |
| BORIS-LAWRIETEMIN, CUR. OPIN. GENET. DEVELOP., vol. 3, 1993, pages 102 - 109 |
| BRASH ET AL., MOL. CELL BIOL., vol. 7, 1987, pages 2031 - 2034 |
| BURNS ET AL., PROC. NATL. ACAD. SCI. USA, vol. 90, 1993, pages 8033 - 8037 |
| CARLENS ET AL., EXP HEMATOL, vol. 28, no. 10, 2000, pages 1137 - 46 |
| CARRILLO ET AL., SIAM J APPLIED MATH, vol. 48, 1988, pages 1073 |
| CAVALIERI ET AL., BLOOD., vol. 102, no. 2, 2003, pages 1637 - 1644 |
| CHAVEZ, A. ET AL., NAT. METHODS, vol. 12, 2015, pages 326 - 328 |
| CHEN ET AL., ADV. DRUG DELIV. REV., vol. 65, no. 10, 2013, pages 1357 - 1369 |
| CHICAYBAM ET AL., PLOS ONE, vol. 8, no. 3, 2013, pages e60298 |
| CHOI JH ET AL.: "Optimization of AAV expression cassettes to improve packaging capacity and transgene expression in neurons.", MOL BRAIN., vol. 7, 11 March 2014 (2014-03-11), pages 17, XP021180027, DOI: 10.1186/1756-6606-7-17 |
| CHYLINSKI ET AL., RNA BIOL., vol. 10, no. 5, 2013, pages 726 - 737 |
| CONG, L. ET AL., SCIENCE, vol. 339, no. 6121, 2013, pages 823 - 23 |
| CONWAY, JE ET AL., J. VIROLOGY, vol. 71, no. 11, 1997, pages 8780 - 8789 |
| DAVIDSON ET AL., PNAS, vol. 97, no. 7, 2000, pages 3428 - 32 |
| EBERLING ET AL., NEUROLOGY, vol. 70, 2008, pages 1980 - 1983 |
| ESVELT ET AL., NATURE METHODS, 2013 |
| FIANDACA ET AL., EXP. NEUROL., vol. 209, 2008, pages 51 - 57 |
| FIANDACA ET AL., NEUROIMAGE, vol. 47, 2009 |
| FINE ET AL., SCI. REP., vol. 5, 2015, pages 10777 |
| FU ET AL., NAT BIOTECHNOL, vol. 32, 2014, pages 279 - 284 |
| GAJ ET AL., TRENDS BIOTECHNOL, vol. 31, no. 7, 2013, pages 397 - 405 |
| GAJ ET AL., TRENDS IN BIOTECHNOLOGY, vol. 31, no. 7, 2013, pages 397 - 405 |
| GAO ET AL., J. VIROL., vol. 78, no. 12, 2004, pages 6381 |
| GAO ET AL., PNAS, vol. 100, no. 10, 2003, pages 6081 - 6 |
| GAO ET AL., PNAS, vol. 99, no. 18, 2002, pages 11854 - 6 |
| GERRITSEN, M.E. ET AL., PNAS, vol. 94, no. 11, 1997, pages 2927 - 2932 |
| GERSBACH, C.A. ET AL., ACC. CHEM. RES., vol. 47, no. 8, 2014, pages 2309 - 18 |
| GERSBACH, C.A. ET AL., CHEM. RES., vol. 47, no. 8, 2014, pages 2309 - 18 |
| GHALEH, H.E.G. ET AL., BIOMED. PHARMACOTHER., vol. 128, 2020, pages 110276 |
| GILBERT, L. A. ET AL., CELL, vol. 152, no. 5, 2013, pages 1173 - 451 |
| HADACZEK ET AL., HUM. GENE THER., vol. 17, 2006, pages 291 - 302 |
| HILTON, I.B. ET AL., NAT. BIOTECHNOL., vol. 33, no. 2, 2015, pages 139 - 517 |
| HSU ET AL., NATURE BIOTECHNOLOGY, 2013 |
| HUANG ET AL., METHODS MOL BIOL, vol. 506, 2009, pages 115 - 126 |
| JINEK, M. ET AL., SCIENCE, vol. 337, no. 6096, 2012, pages 816 - 21 |
| JOHNSTON, NATURE, vol. 346, 1990, pages 776 - 777 |
| KAPLITT ET AL., LANCET, vol. 369, 2007, pages 2097 - 2105 |
| KASARANENI, N. ET AL., SCI. REP., vol. 8, no. 1, 2018, pages 10990 |
| KEARNS, N. A. ET AL., NAT. METHODS., vol. 12, no. 5, 2015, pages 401 - 403 |
| KONERMANN ET AL., NATURE, vol. 517, no. 7536, 2015, pages 583 - 8 |
| KOSTE ET AL., GENE THERAPY, 3 April 2014 (2014-04-03) |
| KOTIN, HUM, GENE THER., vol. 5, 1994, pages 793 - 801 |
| KRAUZE ET AL., METHODS ENZYMOL., vol. 465, 2009, pages 349 - 362 |
| MA, H. ET AL., MOLECULAR THERAPY-NUCLEIC ACIDS, vol. 3, 2014, pages 161 |
| MAKAROVA ET AL., METHODS MOL. BIOL., vol. 1311, 2015, pages 47 - 75 |
| MALI, P. ET AL., NAT. BIOTECHNOL., vol. 31, 2013, pages 833 - 838 |
| MANURI ET AL., HUM GENE THER, vol. 21, no. 4, 2010, pages 427 - 437 |
| MILLER, A. D., HUMAN GENE THERAPY, vol. 1, 1990, pages 5 - 14 |
| MILLERROSMAN, BIOTECHNIQUES, vol. 7, 1989, pages 980 - 990 |
| MILONE, M.C. ET AL., LEUKEMIA., vol. 32, no. 7, 2018, pages 1529 - 1541 |
| MOK ET AL., BIOCHIMICA ET BIOPHYSICA ACTA, vol. 1419, no. 2, 1999, pages 137 - 150 |
| MOON ET AL., EXP. MOL. MED., vol. 51, 2019, pages 1 - 11 |
| NAVARRO ET AL., PLOS ONE, vol. 6, no. 7, pages e21017 |
| NGUYEN ET AL., J. NEUROSURG., vol. 98, 2003, pages 584 - 590 |
| NUNEZ ET AL., J.K. ET AL. CELL, vol. 184, no. 9, 2021, pages 2503 - 2519 |
| NYAMAY'ANTU ET AL., CELL & GENE THERAPY INSIGHTS, vol. 5, 2019, pages 51 - 57 |
| PARK ET AL., TRENDS BIOTECHNOL., vol. 11, 29 November 2011 (2011-11-29), pages 550 - 557 |
| PASSINI ET AL., J. VIROL., vol. 77, no. 12, 2003, pages 6799 - 810 |
| PECHAN ET AL., GENE THER., vol. 16, 2009, pages 10 - 16 |
| PEREZ-PINERA, P. ET AL., NAT. METHODS, vol. 10, 2013, pages 977 - 979 |
| SAITO ET AL., JOURNAL OF NEUROSURGERY PEDIATRICS, vol. 7, 2011, pages 522 - 526 |
| SCARPA ET AL., VIROLOGY, vol. 180, 1991, pages 849 - 852 |
| SCHELLENBERGER ET AL., NATURE BIOTECHNOLOGY, vol. 27, 2009, pages 1186 - 1190 |
| SHARMA ET AL., MOLEC THER NUCL ACIDS, vol. 2, 2013, pages e74 |
| STERNBERG ET AL., NATURE, vol. 507, no. 7491, 2014, pages 258 - 261 |
| SUNG ET AL., BIOMATERIALS RESEARCH, vol. 23, no. 8, 2019 |
| TANENBAUM, M. ET AL., CELL., vol. 159, no. 3, 2014, pages 635 - 646 |
| TRUONG. ET AL., NUCLEIC ACIDS RES., vol. 43, 2015, pages 6450 - 6458 |
| VAN TEDELOO ET AL., GENE THERAPY, vol. 7, no. 16, 2000, pages 1431 - 1437 |
| VANNOCCI ET AL., DISEASE MODELS & MECHANISMS, 2018, pages 11 |
| VERHOEYEN ET AL., METHODS MOL BIOL., vol. 506, 2009, pages 97 - 114 |
| WANG ET AL., J. IMMUNOTHER., vol. 35, no. 9, 2012, pages 689 - 701 |
| WANG Z. ET AL., GENE THER, vol. 10, 2003, pages 2105 - 2111 |
| WRIGHT ET AL., PNAS, vol. 112, no. 10, 2015, pages 2984 - 2989 |
| WRIGHT, D.A. ET AL., NAT. PROTOC., vol. 1, no. 3, 2006, pages 1637 - 52 |
| XU ET AL., MOL. CELL, vol. 81, no. 20, 2021, pages 4333 - 4345 |
| Y. ET AL., BMC BIOTECHNOL., vol. 16, no. 1, pages 2016 |
| YIN ET AL., NATURE REVIEWS GENETICS, vol. 15, 2014, pages 541 - 555 |
| ZETSCHE ET AL., CELL, vol. 163, no. 3, 2015, pages 759 - 71 |
| ZHANG, F. Q., REV. BIOPHYS., 2019, pages 52 |
| ZU ET AL., THE AAPS JOURNAL, vol. 23, no. 78, 2021 |
Cited By (1)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO2025022013A3 (en) * | 2023-07-26 | 2025-04-10 | Imperial College Innovations Ltd | Zinc finger peptides, encoded nucleic acids, methods and uses |
Also Published As
| Publication number | Publication date |
|---|---|
| WO2024163678A3 (en) | 2024-09-12 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| US20240254483A1 (en) | Compositions and methods for modulating expression of frataxin (fxn) | |
| US20240252684A1 (en) | Compositions and methods for modulating expression of methyl-cpg binding protein 2 (mecp2) | |
| WO2024015881A2 (en) | Compositions, systems, and methods for targeted transcriptional activation | |
| US12037617B2 (en) | Methods and compositions for modulating a genome | |
| AU2008216018B2 (en) | Mitochondrial nucleic acid delivery systems | |
| WO2023039440A9 (en) | Hbb-modulating compositions and methods | |
| CN113710693B (en) | DNA binding domain transactivator and its use | |
| US20240066080A1 (en) | Protoparvovirus and tetraparvovirus compositions and methods for gene therapy | |
| US20210189426A1 (en) | Crispr interference based htt allelic suppression and treatment of huntington disease | |
| WO2024163678A2 (en) | Fusion proteins and systems for targeted activation of frataxin (fxn) and related methods | |
| WO2023147558A2 (en) | Crispr methods for correcting bag3 gene mutations in vivo | |
| US20240002839A1 (en) | Crispr sam biosensor cell lines and methods of use thereof | |
| WO2024163683A2 (en) | Systems, compositions, and methods for modulating expression of methyl-cpg binding protein 2 (mecp2) and x-inactive specific transcript (xist) | |
| US20250302994A1 (en) | Erythroparvovirus with a modified genome for gene therapy | |
| US20230279398A1 (en) | Treating human t-cell leukemia virus by gene editing | |
| US20250295810A1 (en) | Erythroparvovirus with a modified capsid for gene therapy | |
| US20250135032A1 (en) | Crispr methods for correcting bag3 gene mutations in vivo | |
| WO2025235643A1 (en) | Profiling of gene therapy agents | |
| Moço | Improving adeno-associated virus serotype 6 production and transduction for gene delivery and CAR T-cell therapy | |
| WO2023235726A2 (en) | Crispr interference therapeutics for c9orf72 repeat expansion disease | |
| CN118556123A (en) | HBB modulating compositions and methods |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| NENP | Non-entry into the national phase |
Ref country code: DE |
|
| 122 | Ep: pct application non-entry in european phase |
Ref document number: 24710964 Country of ref document: EP Kind code of ref document: A2 |