WO2024138550A1 - Nucleic acid amplification method and use thereof in sequencing - Google Patents
Nucleic acid amplification method and use thereof in sequencing Download PDFInfo
- Publication number
- WO2024138550A1 WO2024138550A1 PCT/CN2022/143516 CN2022143516W WO2024138550A1 WO 2024138550 A1 WO2024138550 A1 WO 2024138550A1 CN 2022143516 W CN2022143516 W CN 2022143516W WO 2024138550 A1 WO2024138550 A1 WO 2024138550A1
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- nucleic acid
- concatemer
- copy
- dna
- stranded dna
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Ceased
Links
Images
Classifications
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12P—FERMENTATION OR ENZYME-USING PROCESSES TO SYNTHESISE A DESIRED CHEMICAL COMPOUND OR COMPOSITION OR TO SEPARATE OPTICAL ISOMERS FROM A RACEMIC MIXTURE
- C12P19/00—Preparation of compounds containing saccharide radicals
- C12P19/26—Preparation of nitrogen-containing carbohydrates
- C12P19/28—N-glycosides
- C12P19/30—Nucleotides
- C12P19/34—Polynucleotides, e.g. nucleic acids, oligoribonucleotides
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q1/00—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions
- C12Q1/68—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions involving nucleic acids
- C12Q1/6844—Nucleic acid amplification reactions
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q1/00—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions
- C12Q1/68—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions involving nucleic acids
- C12Q1/6869—Methods for sequencing
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q1/00—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions
- C12Q1/68—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions involving nucleic acids
- C12Q1/6876—Nucleic acid products used in the analysis of nucleic acids, e.g. primers or probes
-
- C—CHEMISTRY; METALLURGY
- C40—COMBINATORIAL TECHNOLOGY
- C40B—COMBINATORIAL CHEMISTRY; LIBRARIES, e.g. CHEMICAL LIBRARIES
- C40B50/00—Methods of creating libraries, e.g. combinatorial synthesis
- C40B50/06—Biochemical methods, e.g. using enzymes or whole viable microorganisms
Definitions
- the present disclosure relates to the field of molecular biology technology, specifically, the present disclosure relates to the field of nucleic acid amplification, and more specifically, the present disclosure relates to a nucleic acid amplification method, a single-stranded DNA multi-copy concatemer and uses thereof.
- sequencing technology has become one of the most commonly used and important research methods. Since Sanger invented the dideoxy chain termination method, the first generation sequencing technology, in 1977, and the promotion of the Human Genome Project, the first generation sequencing technology has greatly improved the speed and throughput of sequencing from the initial isotope labeling plus plate electrophoresis to fluorescent labeling and capillary electrophoresis and automated imaging systems. However, compared with the growing demand for sequencing, the developed first generation sequencing technology still cannot meet the needs. Therefore, the next generation sequencing technology was invented and developed rapidly in a short period of time. The next generation sequencing technology is also called the second generation sequencing technology or massively parallel sequencing technology.
- the conventional steps of large-scale parallel sequencing technology are to fragment DNA, add adapters with known sequences to form a so-called library, and then load these DNA fragments with adapters onto the sequencing chip.
- Each DNA fragment with adapters in the library occupies a certain position on the chip, so that all the sequences on the chip are sequenced at one time, thereby achieving the purpose of large-scale parallel sequencing.
- an extremely important step is what we call the sequencing template signal amplification step.
- the technologies for amplifying sequencing DNA templates to form so-called base signal acquisition units can be mainly divided into three types: microbead emulsion PCR amplification, solid phase bridge PCR amplification, and DNA nanoball amplification.
- Microbead emulsion PCR amplification is mainly used in Roche's 454 sequencing platform and Thermo Fisher's SOLiD and Ion torrent platforms.
- the specific steps are to mix the DNA library with the adapter sequence with the microbeads with complementary sequences to the adapter, dNTP, primers and DNA polymerase and other reactants, add specific mineral oil and surfactant, and then use an oscillator to oscillate violently to make the reaction system form a stable water-in-oil emulsion.
- each droplet will contain only one magnetic bead and a single-stranded DNA.
- at least 10 to the power of 6 ideal droplets can be formed in 1mL of emulsion.
- the surface of each bead is covered with thousands of copies of the same DNA sequence. Subsequently, the emulsion mixture is broken, and the amplified fragments are still bound to the magnetic beads.
- Solid-phase bridge PCR amplification is mainly used in various Illumina sequencing platforms.
- fragmented DNA is connected to adapters to form a library, and the library with adapters is converted into single strands, which are then diluted to a suitable concentration and added to the sequencing chip.
- the single-stranded DNA fragments are bound to the sequencing chip through complementary adapters. Because the concentration of the library is low enough after dilution, it can be considered that the library fragments are evenly bound to the chip surface, and the positions where each fragment is bound are far enough apart.
- the single-stranded DNA hybridized with the complementary adapter on the chip is amplified to form a double strand, and the unfixed complementary single strand is washed away, leaving the amplified chain, whose free end can hybridize with other nearby adapter primers to form a bridge structure.
- it is denatured into a single strand and interacts with other nearby adapter primers again to form a bridge, becoming the template for the next round of amplification and continuing amplification.
- each single copy of the DNA molecule is amplified nearly a thousand times to become a monoclonal DNA cluster.
- Microbead emulsion PCR amplification The copy number of the DNA sequencing template is high and can be amplified to thousands of copies. However, in order to ensure the copy number, the volume of the microbeads is relatively large, and the diameter is generally in the micrometer range, ranging from 1 micron to tens of microns, which affects the sequencing throughput. The use of small microbeads will increase the number of microbeads contained in a droplet, affecting the accuracy of sequencing. In addition, corresponding instruments are required when preparing droplets, which is costly and complicated to operate.
- DNA nanoball amplification Linear amplification makes the number of DNA template copies in the DNA nanoball not high, generally a few hundred copies.
- the structure of the DNA nanoball is loose, and increasing the number of copies will greatly affect the volume of the DNA nanoball, affecting the improvement of sequencing throughput and sequencing length.
- increasing the number of copies or long-fragment DNA templates requires high continuous synthesis ability of the chain displacement polymerase.
- the structure of the DNA nanoball is loose, and before it is loaded onto the chip, its structure is easily destroyed, resulting in the need for extra caution in its operation, which increases the complexity of the operation.
- the present disclosure aims to solve one of the technical problems in the related art at least to some extent.
- nucleic acid amplification method comprises:
- nucleic acid amplification method comprises:
- the nucleic acid amplification method disclosed in the present invention is easy to operate and does not require complex instruments.
- the obtained single-stranded DNA multiple copies are small in size, large in number of copies, and stable in structure, which effectively solves the problem that the DNA nanoballs obtained by the rolling circle amplification method are difficult to operate, loose in structure, large in size, and low in amplification efficiency. If it is applied to the field of sequencing, it can save the chip usage area, reduce costs, and further improve the accuracy of sequencing due to its large copy number.
- any one primer in the primer pair contains restriction site 1, the other primer does not contain restriction site, or contains restriction site 2 different from restriction site 1;
- the enzyme cleavage site 1 is cleaved by enzyme 1, and the enzyme cleavage site 2 is cleaved by enzyme 2, and enzyme 1 and enzyme 2 are different.
- enzyme 1 is used to cut the restriction site 1 contained in the double-stranded DNA multi-copy concatemer to form a single-stranded gap in the double-stranded DNA loop, and the strand with the gap is digested with a digestive enzyme to obtain a first single-stranded DNA multi-copy concatemer.
- At least one primer in the primer pair carries a modified ribonucleotide or deoxyribonucleotide, and the restriction site is cut by the enzyme to form a gap on any one of the circular DNAs in the double-stranded DNA concatemer.
- the nucleic acid amplification method further comprises, after step (D):
- the amplification and ligation reaction system contains a high temperature resistant DNA ligase.
- the amplification and ligation reaction system contains a high-temperature-intolerant DNA ligase.
- the amplification and ligation reaction system when the amplification and ligation reaction system contains a thermolabile DNA ligase, it further includes a DNA helicase and a single-stranded binding protein.
- the single-stranded binding protein contained in the system can prevent the unwound DNA from self-pairing.
- the enzyme 1 and the enzyme 2 include enzymes capable of cleaving the ribonucleotides and/or uracil deoxyribonucleotides.
- the digestive enzyme comprises an enzyme capable of digesting linear DNA.
- the digestive enzyme comprises a DNA exonuclease.
- the DNA exonuclease includes at least one selected from DNA exonuclease I, DNA exonuclease III, T5 Exonuclease, Exonuclease T, Exonuclease VII, and Lambda Exonuclease.
- Another aspect of the present disclosure provides a method for increasing the copy number of a sequencing template. According to an embodiment of the present disclosure, the method comprises:
- the aforementioned nucleic acid amplification method is used to obtain multiple single-stranded DNA multi-copy concatemers to achieve multi-copy amplification of the sequencing template.
- the above-mentioned nucleic acid amplification method is used to perform multi-copy amplification on the sequencing template, and the obtained single-stranded DNA multi-copy concatemer is loaded onto the sequencing chip as the base signal acquisition unit in sequencing, which has the following advantages: the DNA sequencing template of the DNA multi-copy concatemer has a large number of copies and a small volume, thereby occupying a small area of the sequencing chip or flow cell; the increase in the number of copies of the DNA sequencing template will not affect its volume, and the fragment length of the DNA sequencing template has little effect on the volume of the DNA multi-copy concatemer, which is conducive to improving the sequencing read length.
- the structure of the DNA multi-copy concatemer is stable, convenient for operation and long-term storage, and has low requirements for the DNA polymerase used in the reaction.
- Another aspect of the present disclosure provides a single-stranded DNA multi-copy concatemer.
- the single-stranded DNA multi-copy concatemer is prepared by the aforementioned nucleic acid amplification method.
- the single-stranded DNA circular molecule further comprises a tag sequence, and the tag sequence is located between the two known nucleic acid sequences.
- the single-stranded DNA multi-copy concatemer further comprises a plurality of linear DNA single strands, the 5' end of each linear DNA single strand being an amplification primer capable of hybridizing to a known sequence and a nucleic acid sequence complementary to the nucleic acid fragment to be detected.
- the 5' end of the linear DNA single strand is free, and the 3' end is hybridized on the concatemer.
- the present disclosure provides a nucleic acid composite molecule, which includes the above-mentioned single-stranded DNA multi-copy concatemer and multiple linear DNA single strands, and the 5' end of the linear DNA single strand is an amplification primer that can hybridize to a known sequence and a nucleic acid sequence complementary to the nucleic acid fragment to be detected.
- Another aspect of the present disclosure provides uses of the single-stranded DNA multi-copy concatemer prepared by the aforementioned nucleic acid amplification method, the aforementioned single-stranded DNA multi-copy concatemer, and the aforementioned nucleic acid composite molecule in sequencing.
- the present disclosure provides a method for producing a second strand, the method comprising using the aforementioned single-stranded DNA multi-copy concatemer as a template, hybridizing with a primer that is fully or partially complementary to a known nucleic acid sequence, and performing a rolling circle amplification reaction under the action of a polymerase to generate a second strand.
- the primer is free in the solution.
- the aforementioned single-stranded DNA multi-copy concatemer can be used as a base signal acquisition unit and serially loaded on a sequencing chip.
- the single-stranded DNA multi-copy concatemer is small in size, has a large number of copies, and a stable structure, which effectively solves the problems of difficult operation, loose structure, large size, and low amplification efficiency of DNA nanoballs obtained by rolling circle amplification.
- the aforementioned single-stranded DNA multi-copy concatemer is applied to the sequencing field, saving chip usage area, reducing costs, and improving accuracy.
- Another aspect of the present disclosure provides a single-stranded DNA multi-copy concatemer prepared by the aforementioned nucleic acid amplification method, and uses of the aforementioned single-stranded DNA multi-copy concatemer in template amplification and micro-amplification.
- FIG1 shows a schematic flow chart of a nucleic acid amplification method according to an embodiment of the present disclosure
- FIG2 shows a schematic flow chart of a nucleic acid amplification method according to another embodiment of the present disclosure
- FIG3 shows a technical roadmap of a nucleic acid amplification method according to an embodiment of the present disclosure
- FIG8 is a schematic diagram showing a comparison of the theoretical sizes of DNA multi-copy concatemers and DNA nanoballs obtained from templates of the same length according to an embodiment of the present disclosure.
- base also known as nucleobase, nitrogenous base
- natural bases include adenine (A), guanine (G), cytosine (C), thymine (T), uracil (U); non-natural bases include locked nucleic acids (LNA) and bridged nucleic acids (BNA); base analogs include hypoxanthine, deazaadenine, deazaguanine, deazahypoxanthine, 7-methylguanine, 5,6-dihydrouracil, 5-methylcytosine, 5-hydroxymethylcytosine.
- the base type can be used to represent the nucleotide type in the present disclosure.
- the 5' ends of the primers in the primer pair are all phosphorylated.
- the 5' ends of the primers in the primer pair are all phosphorylated means that the 5' end of each primer in the primer pair is phosphorylated, which facilitates the ligase to connect the 3' end of the chain amplification product to the 5' end of the primer to form a ring.
- the multi-copy concatemer of DNA obtained in step (c) or (D) is used as a template, and a primer complementary to the universal sequence of the first single-stranded DNA multi-copy concatemer linker is used to amplify using the Phi29 enzyme. Relying on the chain displacement ability of the Phi29 enzyme, after one circle of amplification on the single-stranded loop, a second single-stranded DNA multi-copy concatemer complementary to the multi-copy concatemer is displaced.
- the enzyme 1 and enzyme 2 respectively include at least one selected from USER enzyme, RNase H, UDG, RNase HII, and hSMUG1, and enzyme 1 and enzyme 2 are different enzymes.
- the high temperature resistant DNA ligase includes at least one selected from Taq ligase and AMP ligase.
- the digestive enzyme comprises an enzyme capable of digesting single-stranded DNA.
- the method for nucleic acid amplification ( FIG. 4 ) comprises:
- thermostable ligase such as T4 DNA ligase.
- T4 DNA ligase an isothermal amplification system is used, which may include DNA helicase, single-stranded binding protein, etc.
- the primer is free in the solution.
- a primer 1 complementary to the adapter sequence is synthesized, and the 5' end of primer 1 is paired with the adapter sequence using the principle of base complementarity, and a sequencing-by-synthesis reaction is subsequently performed.
- the second strand can be obtained by the following method:
- the adapter as a primer binding site, and the Phi29 enzyme for amplification, relying on the chain displacement ability of the Phi29 enzyme, after amplifying one circle on the single-stranded loop, a second chain complementary to the multi-copy concatemer is displaced.
- Amplification is performed with primer 2, and a ligation reaction is performed after one circle of amplification to generate a two-strand DNA multi-copy concatemer. Then the USER enzyme is used to cut the one chain to form a gap, and then the DNA digestion enzyme is used to remove the entire one-strand DNA multi-copy concatemer. Finally, a two-strand DNA multi-copy concatemer in a single-stranded state is obtained.
- the primer 1 and the primer 2 can have different restriction sites in any order, and in addition to U base and ribonucleic acid base, they can also be 5,6-dihydroxythymine, 5-hydroxyuracil, 5-hydroxymethyluracil, and 5-formyluracil, etc.
- the enzyme 1 and the enzyme 2 include at least one selected from USER enzyme, RNase H, UDG, RNase HII, and hSMUG1, and the enzyme 1 and the enzyme 2 are different enzymes.
- the single-stranded DNA multi-copy concatemer obtained by the above-mentioned nucleic acid amplification method is loaded onto a sequencing chip as a base signal acquisition unit in sequencing, and has the following advantages: the DNA sequencing template of the DNA multi-copy concatemer has a large number of copies and a small volume, thereby occupying a small area of the sequencing chip or the flow tank; the increase in the number of copies of the DNA sequencing template will not affect its volume, and the fragment length of the DNA sequencing template has a small effect on the volume of the DNA multi-copy concatemer, which is conducive to improving the sequencing read length.
- the structure of the DNA multi-copy concatemer is stable, convenient for operation and long-term storage, and has low requirements for the DNA polymerase used in the reaction.
- the DNA multi-copy concatemer does not need to react on the sequencing chip or the flow tank, has low requirements for the chip, and does not need to generate nanowells on the chip and pre-plant oligonucleotides on the chip.
- the above-mentioned nucleic acid amplification method is not only applicable to the field of sequencing, but also to any field of micro-amplification. When there are few experimental materials and the samples are scarce, the disclosed method can be used to efficiently amplify the sample nucleic acid content without changing the nucleic acid abundance.
- Primer2 5’P-GCTCACAGAA/rC//rG//rA//rC/ATGGCTACGATCCGAC-3’ (SEQ ID NO: 2)
- r represents the modification of ribonucleotide and P represents phosphorylation modification.
- Escherichia coli genomic DNA was used as material, the nucleic acid amplification method disclosed in the present invention was adopted, the obtained amplification product was loaded onto a chip for sequencing, and the results were directly quality checked, which was in line with expectations.
- the whole process from obtaining the nucleic acid template to obtaining the amplified product is simple to operate, and the required instruments and consumables are also conveniently available to experimental personnel in the field, which improves the experimental efficiency and reduces the experimental cost.
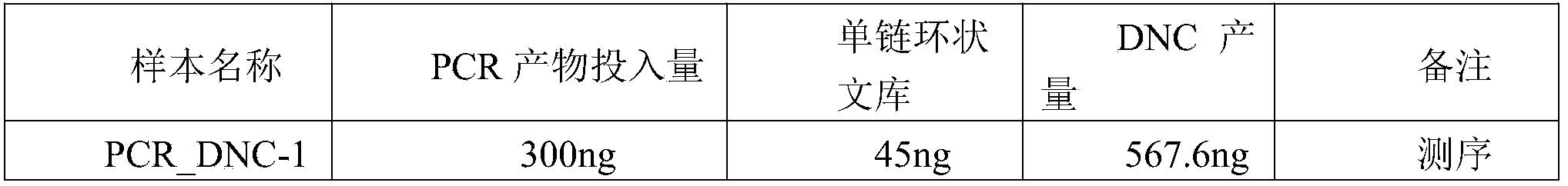
- Table 1 a large amount of DNC can be obtained with the investment of a small amount of DNA library, and it can be seen from Figure 6 that the quality of the obtained DNC meets the standard, so this method can efficiently amplify the nucleic acid template while ensuring the quality of the amplified product.
- Figures 7 and 8 that in sequencing, the method disclosed in the present invention can significantly reduce the chip occupied area. Compared with the prior art, under the chip of the same area, the sequencing read length can be effectively improved, and because the template amplification is not performed on the chip, the chip requirements are relatively low.
- a primer pool for amplifying the tumor hotspot region of the EGFR gene which contains 8 pairs of specific oligonucleotide primers (EGFR_1F-EGFR_8F) and (EGFR_1R-EGFR_8R), with an amplicon size of 100 to 200 bp.
- the sequences of the specific oligonucleotide primers are shown in Table 6 below.
Landscapes
- Chemical & Material Sciences (AREA)
- Life Sciences & Earth Sciences (AREA)
- Organic Chemistry (AREA)
- Health & Medical Sciences (AREA)
- Zoology (AREA)
- Engineering & Computer Science (AREA)
- Wood Science & Technology (AREA)
- Proteomics, Peptides & Aminoacids (AREA)
- Molecular Biology (AREA)
- Biochemistry (AREA)
- Microbiology (AREA)
- Genetics & Genomics (AREA)
- Biotechnology (AREA)
- Bioinformatics & Cheminformatics (AREA)
- General Engineering & Computer Science (AREA)
- General Health & Medical Sciences (AREA)
- Analytical Chemistry (AREA)
- Immunology (AREA)
- Physics & Mathematics (AREA)
- Biophysics (AREA)
- Chemical Kinetics & Catalysis (AREA)
- General Chemical & Material Sciences (AREA)
- Medicinal Chemistry (AREA)
- Measuring Or Testing Involving Enzymes Or Micro-Organisms (AREA)
Abstract
Description
本公开涉及分子生物学技术领域,具体地,本公开涉及核酸扩增领域,更具体地,本公开涉及一种核酸扩增方法、单链DNA多拷贝连环体及其用途。The present disclosure relates to the field of molecular biology technology, specifically, the present disclosure relates to the field of nucleic acid amplification, and more specifically, the present disclosure relates to a nucleic acid amplification method, a single-stranded DNA multi-copy concatemer and uses thereof.
生命科学研究领域,测序技术已经成为最常用最重要的研究手段之一。自从1977桑格发明双脱氧链终止法即第一代测序技术,加上人类基因组计划的推动,一代测序技术从最初的同位素标记加上平板电泳到荧光标记和毛细管电泳和自动化的成像系统,测序的速度和通量大大提升。但是相对于日益增长的测序需求,发展后的一代测序技术还是无法满足。于此,下一代测序技术被发明,并短时间内飞速发展。下一代测序技术又称为二代测序技术或大规模平行测序技术,其主要特点是可以对大量的DNA片段(几百万到几十亿条)同时进行测序,相对于一代的桑格测序,测序速度和通量有了本质上的提升,可以进一步满足科研与应用的测序需求。大规模平行测序技术的常规步骤是将DNA片段化,再加上已知序列的接头,形成所谓的文库,再将这些加上接头的DNA片段加载到测序芯片上,文库中的每个带接头的DNA片段都在芯片上占据一定位置,这样一次性就对芯片上序列都进行测序,从而实现大规模并行测序的目的。在这个过程中,包括一个极其重要的步骤,就是我们所说的测序模板信号放大步骤。这是因为,无论是依靠荧光信号还是离子信号,单个拷贝的DNA序列都不能提供足够的信息强度,所以要在DNA序列加载到芯片上之前,对文库中的每个单拷贝的DNA片段再进行扩增放大,这样的扩增是将单拷贝的DNA序列在固定的空间位置中扩增成单克隆的多拷贝的DNA序列,产生所谓的碱基信号采集单元,来增加信号强度,确保信号可以与背景噪声区分开来;另一方面有助于形成彼此独立的反应中心,可以所有反应中心同时反应,再收集信息,进行信号检测与解读,从而按顺序读取出DNA的片段的序列信息。模板放大的DNA片段的拷贝数目与最终这个多拷贝的模板的占据的空间大小对测序的准确性和测序读长和测序通量都有很大关系。拷贝数越多,则信号越强,信号与背噪声的比就越高,测序的准确性越高。拷贝数越多,越能对抗测序过程中同类拷贝之间的不同步,提高读长。最终这个多拷贝的模板的占据的空间大小越小越能在有限的测序芯片上排列更多的反应中心,越可能增加测序通量。因此,对于大规模并行测序来说,模板放大技术是最重要的核心技术之一。In the field of life science research, sequencing technology has become one of the most commonly used and important research methods. Since Sanger invented the dideoxy chain termination method, the first generation sequencing technology, in 1977, and the promotion of the Human Genome Project, the first generation sequencing technology has greatly improved the speed and throughput of sequencing from the initial isotope labeling plus plate electrophoresis to fluorescent labeling and capillary electrophoresis and automated imaging systems. However, compared with the growing demand for sequencing, the developed first generation sequencing technology still cannot meet the needs. Therefore, the next generation sequencing technology was invented and developed rapidly in a short period of time. The next generation sequencing technology is also called the second generation sequencing technology or massively parallel sequencing technology. Its main feature is that it can sequence a large number of DNA fragments (millions to billions) at the same time. Compared with the first generation of Sanger sequencing, the sequencing speed and throughput have been substantially improved, which can further meet the sequencing needs of scientific research and application. The conventional steps of large-scale parallel sequencing technology are to fragment DNA, add adapters with known sequences to form a so-called library, and then load these DNA fragments with adapters onto the sequencing chip. Each DNA fragment with adapters in the library occupies a certain position on the chip, so that all the sequences on the chip are sequenced at one time, thereby achieving the purpose of large-scale parallel sequencing. In this process, an extremely important step is what we call the sequencing template signal amplification step. This is because, whether relying on fluorescent signals or ion signals, a single copy of the DNA sequence cannot provide sufficient information intensity, so before the DNA sequence is loaded onto the chip, each single copy of the DNA fragment in the library must be amplified. Such amplification is to amplify the single copy of the DNA sequence into a monoclonal multi-copy DNA sequence in a fixed spatial position, generating the so-called base signal acquisition unit to increase the signal intensity and ensure that the signal can be distinguished from the background noise; on the other hand, it helps to form independent reaction centers, so that all reaction centers can react at the same time, collect information, perform signal detection and interpretation, and read the sequence information of the DNA fragments in sequence. The number of copies of the DNA fragment amplified by the template and the size of the space occupied by the multi-copy template have a great impact on the accuracy of sequencing, sequencing read length and sequencing throughput. The more copies there are, the stronger the signal, the higher the signal-to-background noise ratio, and the higher the sequencing accuracy. The more copies there are, the more it can combat the asynchrony between copies of the same type during the sequencing process and improve the read length. The smaller the space occupied by the multi-copy template, the more reaction centers can be arranged on the limited sequencing chip, and the more likely it is to increase the sequencing throughput. Therefore, for large-scale parallel sequencing, template amplification technology is one of the most important core technologies.
从大规模测序技术发明以来,测序DNA模板扩增形成所谓碱基信号采集单元的技术主要可以分为3种,微珠乳液PCR扩增,固相桥式PCR扩增,DNA纳米球扩增。Since the invention of large-scale sequencing technology, the technologies for amplifying sequencing DNA templates to form so-called base signal acquisition units can be mainly divided into three types: microbead emulsion PCR amplification, solid phase bridge PCR amplification, and DNA nanoball amplification.
微珠乳液PCR扩增,主要应用于罗氏的454测序平台和Thermo Fisher的SOLiD和Ion torrent平台等。具体的步骤是将构建带有接头序列的DNA文库与带有于接头互补序列的微珠、dNTP、引物和DNA聚合酶等反应物混合,加入特定的矿物油和表面活性剂,再利用振荡器剧烈振荡,使反应体系形成油包水(water-in-oil)的稳定乳浊液。在理想条件下,每一个液滴将只包含一个磁珠和一条单链DNA,通过控制该步骤的条件,1mL乳液中可以形成至少10的6次方个理想的液滴。经过PCR扩增后,使每个珠子表面覆盖有数千拷贝的相同DNA序列。随后,乳液混合物被打破,扩增的片段依然结合在磁珠上。Microbead emulsion PCR amplification is mainly used in Roche's 454 sequencing platform and Thermo Fisher's SOLiD and Ion torrent platforms. The specific steps are to mix the DNA library with the adapter sequence with the microbeads with complementary sequences to the adapter, dNTP, primers and DNA polymerase and other reactants, add specific mineral oil and surfactant, and then use an oscillator to oscillate violently to make the reaction system form a stable water-in-oil emulsion. Under ideal conditions, each droplet will contain only one magnetic bead and a single-stranded DNA. By controlling the conditions of this step, at least 10 to the power of 6 ideal droplets can be formed in 1mL of emulsion. After PCR amplification, the surface of each bead is covered with thousands of copies of the same DNA sequence. Subsequently, the emulsion mixture is broken, and the amplified fragments are still bound to the magnetic beads.
固相桥式PCR扩增,主要应用于illumina各个测序平台。在固相桥式PCR扩增中,片段化的DNA连接上接头形成文库,将带接头的文库变成单链,再稀释到合适浓度加入到测序芯片上。DNA单链片段通过互补接头结合在测序芯片上,因为文库稀释后浓度足够低,可以认为文库片段均匀的结合在芯片表面,每个片段结合的位置相距足够远。与芯片上互补接头杂交的DNA单链扩增形成双链,洗去未固定的的互补单链,留下扩增后的链,其自由端可以与附近的其他接头引物杂交,形成桥式结构,互补扩增后,经变性成单链,再次与附近的其他接头引物相互作用形成桥,成为下一轮扩增的模板继续扩增。在反复进行扩增后,每个单拷贝的DNA分子得到近千倍扩增,成为单克隆的DNA簇。Solid-phase bridge PCR amplification is mainly used in various Illumina sequencing platforms. In solid-phase bridge PCR amplification, fragmented DNA is connected to adapters to form a library, and the library with adapters is converted into single strands, which are then diluted to a suitable concentration and added to the sequencing chip. The single-stranded DNA fragments are bound to the sequencing chip through complementary adapters. Because the concentration of the library is low enough after dilution, it can be considered that the library fragments are evenly bound to the chip surface, and the positions where each fragment is bound are far enough apart. The single-stranded DNA hybridized with the complementary adapter on the chip is amplified to form a double strand, and the unfixed complementary single strand is washed away, leaving the amplified chain, whose free end can hybridize with other nearby adapter primers to form a bridge structure. After complementary amplification, it is denatured into a single strand and interacts with other nearby adapter primers again to form a bridge, becoming the template for the next round of amplification and continuing amplification. After repeated amplification, each single copy of the DNA molecule is amplified nearly a thousand times to become a monoclonal DNA cluster.
DNA纳米球扩增,主要应用华大智造的各个DNBSEQ测序平台。DNA纳米球扩增是基于滚环扩增,能够在溶液中完成模板扩增。为了实现滚环扩增,DNBSEQ的文库会制备成单链环。这样的单链环文库会被作为模板,在具有链置换能力的聚合酶的反应体系中进行滚环复制,最终得到几百个拷贝的线性DNA复制链,这个复制链会缠绕在一起形成一个纳米球,大大增强了信号强度。DNB纳米球扩增是线性扩增过程,不是指数扩增。DNA nanoball amplification mainly uses various DNBSEQ sequencing platforms manufactured by MGI. DNA nanoball amplification is based on rolling circle amplification, which can complete template amplification in solution. In order to achieve rolling circle amplification, the DNBSEQ library will be prepared into a single-stranded circle. Such a single-stranded circle library will be used as a template for rolling circle replication in the reaction system of a polymerase with chain displacement ability, and finally hundreds of copies of linear DNA replication chains will be obtained. This replication chain will be entangled together to form a nanoball, which greatly enhances the signal intensity. DNB nanoball amplification is a linear amplification process, not an exponential amplification.
上述3种主要的现有技术,的确解决了高通量测序中模板放大技术难题,但同时也存在下述不足:The above three main existing technologies have indeed solved the technical problems of template amplification in high-throughput sequencing, but they also have the following shortcomings:
微珠乳液PCR扩增:得到DNA测序模板的拷贝数高,可以放大到数千拷贝,但是为了保证拷贝数,微珠的体积较大,直径一般都是微米级,从1微米到几十微米不等,影响测序通量。采用小微珠会增加一个液滴中包含2个及以上微珠的情况出现,影响测序准确性。并且在制备液滴时候需要相应的仪器,成本高,操作复杂。Microbead emulsion PCR amplification: The copy number of the DNA sequencing template is high and can be amplified to thousands of copies. However, in order to ensure the copy number, the volume of the microbeads is relatively large, and the diameter is generally in the micrometer range, ranging from 1 micron to tens of microns, which affects the sequencing throughput. The use of small microbeads will increase the number of microbeads contained in a droplet, affecting the accuracy of sequencing. In addition, corresponding instruments are required when preparing droplets, which is costly and complicated to operate.
固相桥式PCR扩增:得到的DNA测序模板(或称簇)的拷贝数较高,有近千拷贝,但是同样如果需要再增加拷贝数,就需要占据更大的芯片面积,导致测序通量和测序长度无法提升。另外需要预先再测序芯片上种植接头,对芯片的要求高。Solid-phase bridge PCR amplification: The copy number of the obtained DNA sequencing template (or cluster) is relatively high, with nearly a thousand copies. However, if the copy number needs to be increased, a larger chip area will be required, resulting in the inability to increase the sequencing throughput and sequencing length. In addition, the sequencing chip needs to be pre-planted with connectors, which places high demands on the chip.
DNA纳米球扩增:线性扩增使得DNA纳米球中的DNA模板拷贝数不高,一般是几百个拷贝。DNA纳米球结构松散,增加拷贝数会很多程度影响DNA纳米球的体积,影响测序通量和测序长度的提升。另外增加拷贝数或者长片段DNA模板对链置换聚合酶的持续合成能力要求高。DNA纳米球结构松散,没有加载到芯片上之前,其结构容易被破环,导致对其操作需要格外小心,增加了操作复杂度。DNA nanoball amplification: Linear amplification makes the number of DNA template copies in the DNA nanoball not high, generally a few hundred copies. The structure of the DNA nanoball is loose, and increasing the number of copies will greatly affect the volume of the DNA nanoball, affecting the improvement of sequencing throughput and sequencing length. In addition, increasing the number of copies or long-fragment DNA templates requires high continuous synthesis ability of the chain displacement polymerase. The structure of the DNA nanoball is loose, and before it is loaded onto the chip, its structure is easily destroyed, resulting in the need for extra caution in its operation, which increases the complexity of the operation.
因此,现有技术在测序过程中模板放大的问题上仍有待改进。Therefore, the existing technology still needs to be improved on the issue of template amplification during sequencing.
发明内容Summary of the invention
本公开旨在至少在一定程度上解决相关技术中的技术问题之一。The present disclosure aims to solve one of the technical problems in the related art at least to some extent.
本公开的一方面提供一种核酸扩增方法,根据本公开的实施方案,所述核酸扩增方法包括:One aspect of the present disclosure provides a nucleic acid amplification method. According to an embodiment of the present disclosure, the nucleic acid amplification method comprises:
(A)将所述核酸与已知序列相连,获得待扩增核酸;(A) connecting the nucleic acid to a known sequence to obtain the nucleic acid to be amplified;
(B)以所述待扩增核酸中含有的已知序列设计引物对;(B) designing a primer pair based on a known sequence contained in the nucleic acid to be amplified;
(C)利用所述待扩增核酸、所述引物对进行扩增连接反应,获得第一双链DNA多拷贝连环体。(C) performing an amplification and ligation reaction using the nucleic acid to be amplified and the primer pair to obtain a first double-stranded DNA multi-copy concatemer.
本公开的一方面提供一种核酸扩增方法,根据本公开的实施方案,所述核酸扩增方法包括:One aspect of the present disclosure provides a nucleic acid amplification method. According to an embodiment of the present disclosure, the nucleic acid amplification method comprises:
(a)利用扩增引物扩增目标核酸,获得扩增产物,其中所述扩增引物的5’末端带有已 知序列;(a) amplifying a target nucleic acid using an amplification primer to obtain an amplification product, wherein the 5' end of the amplification primer has a known sequence;
(d)以所述扩增产物中含有的已知序列设计引物对,其中,所述引物对中的至少一条引物上含有酶切位点;(d) designing a primer pair based on a known sequence contained in the amplified product, wherein at least one primer in the primer pair contains a restriction site;
(c)利用所述扩增产物、所述引物进行扩增连接反应,获得第一双链DNA多拷贝连环体。(c) performing an amplification and ligation reaction using the amplified product and the primers to obtain a first double-stranded DNA multi-copy concatemer.
本发明所公开的核酸扩增方法,其操作方便,不需要复杂的仪器,所得到的单链DNA多拷贝连环,体积小的同时,拷贝数量大,结构稳定,有效的解决了用滚环扩增法获得的DNA纳米球,操作难度大,结构松散,体积较大,扩增效率低的问题。如其应用于测序领域,则更是节约了芯片使用面积,降低成本的同时,因其较大的拷贝数,进一步提高了测序的准确度。The nucleic acid amplification method disclosed in the present invention is easy to operate and does not require complex instruments. The obtained single-stranded DNA multiple copies are small in size, large in number of copies, and stable in structure, which effectively solves the problem that the DNA nanoballs obtained by the rolling circle amplification method are difficult to operate, loose in structure, large in size, and low in amplification efficiency. If it is applied to the field of sequencing, it can save the chip usage area, reduce costs, and further improve the accuracy of sequencing due to its large copy number.
根据本公开的实施方案,当所述引物对中任意一条引物含有酶切位点1时,另一条引物不含酶切位点,或者含有与酶切位点1不同的酶切位点2;According to the embodiments of the present disclosure, when any one primer in the primer pair contains
其中,in,
所述酶切位点1被酶1切割,所述酶切位点2被酶2切割,且酶1与酶2不相同。The
根据本公开的实施方案,步骤(C)或(c)之后进一步包括,According to an embodiment of the present disclosure, step (c) or (c) further comprises,
(D)当所述引物对中任意一条引物含有酶切位点1,且另一条引物不含酶切位点时,利用酶1切割所述双链DNA多拷贝连环体中含有的酶切位点1,在双链DNA环中形成单链缺口,利用消化酶对所述带有缺口的链进行消化,以便获得第一单链DNA多拷贝连环体。(D) When any one of the primers in the primer pair contains a
根据本公开的实施方案,所述引物对中的引物的5’端均含有磷酸化修饰。According to an embodiment of the present disclosure, the 5' ends of the primers in the primer pair all contain phosphorylation modifications.
对引物对中的引物的5’端进行磷酸化修饰,便于连接酶将链状扩增产物的3’端与引物的5’端连接成环。The 5' end of the primer in the primer pair is phosphorylated to facilitate the ligase to connect the 3' end of the chain amplification product to the 5' end of the primer to form a ring.
根据本公开的实施方案,所述引物对中的两条引物至少具有部分重叠互补片段,所述重叠互补片段的Tm值低于所述两条引物的Tm值。由此,能够确保退火时引物与模板优先结合,而不是两引物之间形成二聚体。According to the embodiments of the present disclosure, the two primers in the primer pair have at least partially overlapping complementary fragments, and the Tm value of the overlapping complementary fragments is lower than the Tm value of the two primers. Thus, it can be ensured that the primers preferentially bind to the template during annealing rather than forming a dimer between the two primers.
根据本公开的实施方案,所述引物对中的两条引物之间没有重叠互补片段。According to an embodiment of the present disclosure, there is no overlapping complementary fragment between the two primers in the primer pair.
根据本公开的实施方案,所述酶切位点包括能够被所述酶切割的经修饰的核糖核苷酸或脱氧核糖核苷酸。According to an embodiment of the present disclosure, the enzyme cleavage site includes a modified ribonucleotide or deoxyribonucleotide that can be cleaved by the enzyme.
根据本公开的实施方案,所述引物对中至少一条引物上带有经修饰的核糖核苷酸或脱氧核糖核苷酸,该酶切位点被酶切割,在双链DNA连环体任意一条环状DNA上形成缺口。According to the embodiments of the present disclosure, at least one primer in the primer pair carries a modified ribonucleotide or deoxyribonucleotide, and the restriction site is cut by the enzyme to form a gap on any one of the circular DNAs in the double-stranded DNA concatemer.
根据本公开的实施方案,所述酶切位点包括选自核糖核苷酸、尿嘧啶脱氧核糖核苷酸、5,6二羟基胸腺嘧啶脱氧核糖核苷酸、5-羟基尿嘧啶(5-hydroxyuracil)脱氧核糖核苷酸、5-羟甲基尿嘧啶(5-hydroxymethyluracil)脱氧核糖核苷酸、5-甲酰基尿嘧啶(5-formyluracil)脱氧核糖核苷酸中的至少之一。According to an embodiment of the present disclosure, the enzyme cleavage site includes at least one selected from ribonucleotides, uracil deoxyribonucleotides, 5,6-dihydroxythymine deoxyribonucleotides, 5-hydroxyuracil (5-hydroxyuracil) deoxyribonucleotides, 5-hydroxymethyluracil (5-hydroxymethyluracil) deoxyribonucleotides, and 5-formyluracil (5-formyluracil) deoxyribonucleotides.
根据本公开的实施方案,所述核酸包括选自链状DNA、环状DNA中的至少之一。According to an embodiment of the present disclosure, the nucleic acid includes at least one selected from linear DNA and circular DNA.
根据本公开的实施方案,所述核酸包括链状DNA文库和/或环状DNA文库。According to an embodiment of the present disclosure, the nucleic acid includes a chain DNA library and/or a circular DNA library.
根据本公开的实施方案,所述环状DNA文库为单链环状DNA文库。According to an embodiment of the present disclosure, the circular DNA library is a single-stranded circular DNA library.
根据本公开的实施方案,所述环状DNA文库为双链环状DNA文库。According to an embodiment of the present disclosure, the circular DNA library is a double-stranded circular DNA library.
根据本公开的实施方案,当所述核酸为链状DNA时,步骤(A)进一步包括将与已知序列相连的核酸进行环化反应,获得环化的待扩增核酸。According to an embodiment of the present disclosure, when the nucleic acid is a chain DNA, step (A) further comprises subjecting the nucleic acid connected to the known sequence to a cyclization reaction to obtain a cyclized nucleic acid to be amplified.
本发明步骤(A)中,核酸与已知序列相连后获得的待扩增核酸优选为环化的核酸,这 样便于扩增链的5’端和3’端连接成环,提升连接效率。In step (A) of the present invention, the nucleic acid to be amplified obtained after connecting the nucleic acid to the known sequence is preferably a circularized nucleic acid, which facilitates the connection of the 5' end and the 3' end of the amplified chain into a ring and improves the connection efficiency.
根据本公开的实施方案,所述引物对中的两条引物分别含有酶切位点1和酶切位点2,所述酶切位点2被酶2切割。According to an embodiment of the present disclosure, the two primers in the primer pair contain
根据本公开的实施方案,所述核酸扩增方法在步骤(D)之后进一步包括:According to an embodiment of the present disclosure, the nucleic acid amplification method further comprises, after step (D):
(E)以所述第一单链DNA多拷贝连环体为模板,利用所述引物对中的引物进行扩增连接反应,获得第二双链DNA多拷贝连环体;(E) using the first single-stranded DNA multi-copy concatemer as a template, and using the primers in the primer pair to perform an amplification and ligation reaction to obtain a second double-stranded DNA multi-copy concatemer;
(F)利用酶2切割所述第二双链DNA多拷贝连环体中含有的酶切位点2,在所述第二双链DNA多拷贝连环体中形成缺口单链,利用消化酶对缺口单链进行消化,以便获得第二单链DNA多拷贝连环体。(F) using
根据本公开的实施方案,所述核酸扩增方法在步骤(D)之后进一步包括:According to an embodiment of the present disclosure, the nucleic acid amplification method further comprises, after step (D):
(G)利用引物对中与所述第一单链DNA多拷贝连环体结合的引物和带有链置换能力的酶,对步骤(D)中的所述第一单链DNA多拷贝连环体进行扩增置换连接反应,获得第二单链DNA多拷贝连环体。(G) using the primer in the primer pair that binds to the first single-stranded DNA multi-copy concatemer and an enzyme with strand displacement ability, the first single-stranded DNA multi-copy concatemer in step (D) is subjected to an amplification displacement ligation reaction to obtain a second single-stranded DNA multi-copy concatemer.
根据本公开的实施方案,所述扩增连接反应的体系中含有DNA聚合酶,所述DNA聚合酶不具有5’-3’外切酶活性。所述DNA聚合酶不具有5’到3’外切酶活性,能够保证扩增的效率,防止扩增产物被酶切。According to the embodiments of the present disclosure, the amplification and ligation reaction system contains a DNA polymerase, and the DNA polymerase does not have 5'-3' exonuclease activity. The DNA polymerase does not have 5' to 3' exonuclease activity, which can ensure the efficiency of amplification and prevent the amplification product from being cleaved by enzymes.
根据本公开的实施方案,所述扩增连接反应的体系中含有DNA聚合酶,所述DNA聚合酶包括选自KAPA HiFi HotStart DNA Polymerase、KAPA HiFi HotStart Uracil+Polymerase、BGI Golden High-Fidelity Polymerase、BGI Platinum HiFi Hotstart Polymerase中的至少之一。According to the embodiments of the present disclosure, the amplification and ligation reaction system contains a DNA polymerase, and the DNA polymerase includes at least one selected from KAPA HiFi HotStart DNA Polymerase, KAPA HiFi HotStart Uracil+Polymerase, BGI Golden High-Fidelity Polymerase, and BGI Platinum HiFi Hotstart Polymerase.
根据本公开的实施方案,所述扩增连接反应的体系中含有耐高温DNA连接酶。According to an embodiment of the present disclosure, the amplification and ligation reaction system contains a high temperature resistant DNA ligase.
根据本公开的实施方案,所述耐高温DNA连接酶包括选自Taq ligase、AMP ligase中的至少之一。According to an embodiment of the present disclosure, the high temperature resistant DNA ligase includes at least one selected from Taq ligase and AMP ligase.
根据本公开的实施方案,所述扩增连接反应的体系中含有不耐高温DNA连接酶。According to an embodiment of the present disclosure, the amplification and ligation reaction system contains a high-temperature-intolerant DNA ligase.
根据本公开的实施方案,所述不耐高温DNA连接酶包括选自T4 DNA ligase、E.coli DNA Ligase、T3 DNA Ligase、T7 DNA Ligase中的至少之一。According to the embodiments of the present disclosure, the heat-intolerant DNA ligase includes at least one selected from T4 DNA ligase, E. coli DNA Ligase, T3 DNA Ligase, and T7 DNA Ligase.
根据本公开的实施方案,所述扩增连接反应的体系中含有不耐高温DNA连接酶时,进一步包括DNA解旋酶、单链结合蛋白。体系中含有的单链结合蛋白能够防止解旋的DNA自我配对。According to the embodiment of the present disclosure, when the amplification and ligation reaction system contains a thermolabile DNA ligase, it further includes a DNA helicase and a single-stranded binding protein. The single-stranded binding protein contained in the system can prevent the unwound DNA from self-pairing.
根据本公开的实施方案,所述酶1、酶2包括能够切割所述核糖核苷酸和/或尿嘧啶脱氧核糖核苷酸的酶。According to an embodiment of the present disclosure, the
根据本公开的实施方案,所述酶1、酶2包括选自USER酶、RNase H、UDG、RNase HII、hSMUG1、APE1、Endonuclease VIII、Endonuclease III(Nth)至少之一,且酶1、酶2为不同的酶。According to the embodiments of the present disclosure, the
根据本公开的实施方案,所述消化酶包括能够消化线性DNA的酶。According to an embodiment of the present disclosure, the digestive enzyme comprises an enzyme capable of digesting linear DNA.
根据本公开的实施方案,所述消化酶包括DNA外切酶。According to an embodiment of the present disclosure, the digestive enzyme comprises a DNA exonuclease.
根据本公开的实施方案,所述DNA外切酶包括选自DNA外切酶Ⅰ、DNA外切酶Ⅲ、T5 Exonuclease、Exonuclease T、Exonuclease VII、Lambda Exonuclease中的至少之一。According to the embodiments of the present disclosure, the DNA exonuclease includes at least one selected from DNA exonuclease I, DNA exonuclease III, T5 Exonuclease, Exonuclease T, Exonuclease VII, and Lambda Exonuclease.
本公开另一方面提供一种测序模板拷贝数增加的方法。根据本公开的实施方案,所述方法包括:Another aspect of the present disclosure provides a method for increasing the copy number of a sequencing template. According to an embodiment of the present disclosure, the method comprises:
以具有测序接头序列的单链环状DNA文库和/或双链环状DNA文库作为模板,利用前 面所述的核酸扩增方法,获得多个单链DNA多拷贝连环体,以便实现测序模板的多拷贝扩增。Using a single-stranded circular DNA library and/or a double-stranded circular DNA library with a sequencing adapter sequence as a template, the aforementioned nucleic acid amplification method is used to obtain multiple single-stranded DNA multi-copy concatemers to achieve multi-copy amplification of the sequencing template.
利用上述核酸扩增方法对测序模板进行多拷贝扩增,得到的单链DNA多拷贝连环体,作为测序中的碱基信号采集单元,加载到测序芯片上,有以下优势:DNA多拷贝连环体的DNA测序模板拷贝数多,且其体积小,从而占用测序芯片或流动槽的面积小;DNA测序模板的拷贝数的增加也不会影响其体积,DNA测序模板的片段长度对DNA多拷贝连环体的体积影响小,有利于提升测序读长。并且DNA多拷贝连环体的结构稳定,方便操作以及长期存放,并且对反应用到的DNA聚合酶要求低。The above-mentioned nucleic acid amplification method is used to perform multi-copy amplification on the sequencing template, and the obtained single-stranded DNA multi-copy concatemer is loaded onto the sequencing chip as the base signal acquisition unit in sequencing, which has the following advantages: the DNA sequencing template of the DNA multi-copy concatemer has a large number of copies and a small volume, thereby occupying a small area of the sequencing chip or flow cell; the increase in the number of copies of the DNA sequencing template will not affect its volume, and the fragment length of the DNA sequencing template has little effect on the volume of the DNA multi-copy concatemer, which is conducive to improving the sequencing read length. In addition, the structure of the DNA multi-copy concatemer is stable, convenient for operation and long-term storage, and has low requirements for the DNA polymerase used in the reaction.
本公开另一方面提供一种单链DNA多拷贝连环体。根据本公开的实施方案,所述单链DNA多拷贝连环体由前面所述的核酸扩增方法制备获得。Another aspect of the present disclosure provides a single-stranded DNA multi-copy concatemer. According to an embodiment of the present disclosure, the single-stranded DNA multi-copy concatemer is prepared by the aforementioned nucleic acid amplification method.
根据本公开的实施方案,所述单链DNA多拷贝连环体含有至少两个单链DNA环状分子,其中所述单链DNA环状分子包括两段已知核酸序列和一段待测核酸片段。According to an embodiment of the present disclosure, the single-stranded DNA multi-copy concatemer contains at least two single-stranded DNA circular molecules, wherein the single-stranded DNA circular molecules include two known nucleic acid sequences and a nucleic acid fragment to be detected.
根据本公开的实施方案,所述单链DNA环状分子进一步包括标签序列,所述标签序列位于所述两段已知核酸序列之间。According to an embodiment of the present disclosure, the single-stranded DNA circular molecule further comprises a tag sequence, and the tag sequence is located between the two known nucleic acid sequences.
根据本公开的实施方案,所述单链DNA多拷贝连环体进一步包括多条线性DNA单链,所述线性DNA单链的5’端为能够杂交到已知序列上的扩增引物和一段与所述待测核酸片段互补的核酸序列。According to the embodiments of the present disclosure, the single-stranded DNA multi-copy concatemer further comprises a plurality of linear DNA single strands, the 5' end of each linear DNA single strand being an amplification primer capable of hybridizing to a known sequence and a nucleic acid sequence complementary to the nucleic acid fragment to be detected.
根据本公开的实施方案,所述线性DNA单链的5’端游离,3’端杂交在连环体上。According to an embodiment of the present disclosure, the 5' end of the linear DNA single strand is free, and the 3' end is hybridized on the concatemer.
本公开又一方面提供一种核酸复合分子,所述核酸复合分子包括上述的单链DNA多拷贝连环体和多条线性DNA单链,所述线性DNA单链的5’端为能够杂交到已知序列上的扩增引物和一段与所述待测核酸片段互补的核酸序列。On the other hand, the present disclosure provides a nucleic acid composite molecule, which includes the above-mentioned single-stranded DNA multi-copy concatemer and multiple linear DNA single strands, and the 5' end of the linear DNA single strand is an amplification primer that can hybridize to a known sequence and a nucleic acid sequence complementary to the nucleic acid fragment to be detected.
根据本公开的实施方案,所述线性DNA单链的5’端游离,3’端杂交在连环体上。According to an embodiment of the present disclosure, the 5' end of the linear DNA single strand is free, and the 3' end is hybridized on the concatemer.
本公开另一方面提供前面所述的核酸扩增方法制备获得的单链DNA多拷贝连环体、前面所述的单链DNA多拷贝连环体、前面所述的核酸复合分子在测序中的用途。Another aspect of the present disclosure provides uses of the single-stranded DNA multi-copy concatemer prepared by the aforementioned nucleic acid amplification method, the aforementioned single-stranded DNA multi-copy concatemer, and the aforementioned nucleic acid composite molecule in sequencing.
本公开又一方面提供一种二链生产方法,所述方法包括,以前面所述的单链DNA多拷贝连环体为模板,与已知核酸序列全部或部分互补的引物杂交,在聚合酶的作用下进行滚环扩增反应,生成二链。In another aspect, the present disclosure provides a method for producing a second strand, the method comprising using the aforementioned single-stranded DNA multi-copy concatemer as a template, hybridizing with a primer that is fully or partially complementary to a known nucleic acid sequence, and performing a rolling circle amplification reaction under the action of a polymerase to generate a second strand.
根据本公开的实施方案,所述引物固定在固体支持物上,所述固体支持物可以包括选自:磁珠、多聚物、水凝胶、金属或芯片。According to an embodiment of the present disclosure, the primer is fixed on a solid support, and the solid support may include a material selected from the group consisting of: magnetic beads, polymers, hydrogels, metals, or chips.
根据本公开的实施方案,所述引物游离在溶液中。According to an embodiment of the present disclosure, the primer is free in the solution.
可以将前面所述的单链DNA多拷贝连环体作为碱基信号采集单元,连载在测序芯片上。该单链DNA多拷贝连环体体积小,拷贝数量大,结构稳定,有效的解决了用滚环扩增法获得的DNA纳米球操作难度大、结构松散、体积较大、扩增效率低的问题。前面所述的单链DNA多拷贝连环体应用于测序领域,节约了芯片使用面积,降低成本,提升准确性。The aforementioned single-stranded DNA multi-copy concatemer can be used as a base signal acquisition unit and serially loaded on a sequencing chip. The single-stranded DNA multi-copy concatemer is small in size, has a large number of copies, and a stable structure, which effectively solves the problems of difficult operation, loose structure, large size, and low amplification efficiency of DNA nanoballs obtained by rolling circle amplification. The aforementioned single-stranded DNA multi-copy concatemer is applied to the sequencing field, saving chip usage area, reducing costs, and improving accuracy.
本公开另一方面提供前面所述的核酸扩增方法制备获得的单链DNA多拷贝连环体、前面所述的单链DNA多拷贝连环体在模板放大、微量扩增中的用途。Another aspect of the present disclosure provides a single-stranded DNA multi-copy concatemer prepared by the aforementioned nucleic acid amplification method, and uses of the aforementioned single-stranded DNA multi-copy concatemer in template amplification and micro-amplification.
本公开的附加方面和优点将在下面的描述中部分给出,部分将从下面的描述中变得明显,或通过本公开的实践了解到。Additional aspects and advantages of the present disclosure will be given in part in the following description and in part will be obvious from the following description or learned through practice of the present disclosure.
本公开的上述和/或附加的方面和优点从结合下面附图对实施方案的描述中将变得明显和容易理解,其中:The above and/or additional aspects and advantages of the present disclosure will become apparent and easily understood from the description of the embodiments in conjunction with the following drawings, in which:
图1显示了本公开一个实施方案的核酸扩增方法流程示意图;FIG1 shows a schematic flow chart of a nucleic acid amplification method according to an embodiment of the present disclosure;
图2显示了本公开另一个实施方案的核酸扩增方法流程示意图;FIG2 shows a schematic flow chart of a nucleic acid amplification method according to another embodiment of the present disclosure;
图3显示了本公开一个实施方案的核酸扩增方法技术路线图;FIG3 shows a technical roadmap of a nucleic acid amplification method according to an embodiment of the present disclosure;
图4显示了本公开一个实施方案的形成单链DNA多拷贝连环体方法图;FIG4 shows a diagram of a method for forming multiple copies of single-stranded DNA concatemers according to an embodiment of the present disclosure;
图5显示了本公开一个实施方案的链状核酸模板形成单链DNA多拷贝连环体方法图;FIG5 shows a method for forming multiple copies of single-stranded DNA concatemers from a chained nucleic acid template according to an embodiment of the present disclosure;
图6显示了本公开一个实施方案的DNA多拷贝连环体的电泳图;FIG6 shows an electrophoresis diagram of a DNA multi-copy concatemer according to an embodiment of the present disclosure;
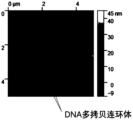
图7显示了本公开一个实施方案的DNA多拷贝连环体测序后芯片的原子力显微图;FIG7 shows an atomic force micrograph of a chip after DNA multi-copy concatemer sequencing according to an embodiment of the present disclosure;
图8显示了本公开一个实施方案的相同长度模板得到DNA多拷贝连环体与DNA纳米球的理论大小的比较示意图。FIG8 is a schematic diagram showing a comparison of the theoretical sizes of DNA multi-copy concatemers and DNA nanoballs obtained from templates of the same length according to an embodiment of the present disclosure.
发明详细描述DETAILED DESCRIPTION OF THE INVENTION
下面详细描述本公开的实施方案。下面描述的实施方案是示例性的,仅用于解释本公开,而不能理解为对本公开的限制。The embodiments of the present disclosure are described in detail below. The embodiments described below are exemplary and are only used to explain the present disclosure, and should not be construed as limiting the present disclosure.
此外,术语“第一”、“第二”仅用于描述目的,而不能理解为指示或暗示相对重要性或者隐含指明所指示的技术特征的数量。由此,限定有“第一”、“第二”的特征可以明示或者隐含地包括至少一个该特征。在本公开的描述中,“多个”的含义是至少两个,例如两个,三个等,除非另有明确具体的限定。In addition, the terms "first" and "second" are used for descriptive purposes only and should not be understood as indicating or implying relative importance or implicitly indicating the number of technical features indicated. Thus, a feature defined as "first" or "second" may explicitly or implicitly include at least one of the features. In the description of the present disclosure, "plurality" means at least two, such as two, three, etc., unless otherwise clearly and specifically defined.
为了更容易理解本发明,以下具体定义了某些技术和科学术语。除显而易见在本文件中的它处另有明确定义,否则本文中使用的所有其它技术和科学术语都具有本发明所属领域的一般技术人员通常理解的含义。In order to make the present invention more easily understood, certain technical and scientific terms are specifically defined below. Unless otherwise clearly defined elsewhere in this document, all other technical and scientific terms used herein have the meanings commonly understood by those skilled in the art to which the present invention belongs.
在本文中,术语“包含”或“包括”为开放式表达,即包括本发明所指明的内容,但并不排除其他方面的内容。In this document, the terms “include” or “comprising” are open expressions, that is, including the contents specified in the present invention but not excluding other contents.
在本文中,术语“任选地”、“任选的”或“任选”通常是指随后所述的事件或状况可以但未必发生,并且该描述包括其中发生该事件或状况的情况,以及其中未发生该事件或状况的情况。As used herein, the terms "optionally", "optional" or "optionally" generally mean that the subsequently described event or circumstance may but need not occur, and that the description includes instances where the event or circumstance occurs and instances where it does not.
术语定义和说明Definitions and explanations of terms
在本公开中,术语“模板”是指待测的核酸分子,表示一定长度的核苷酸的聚合物,核苷酸可以包括核糖核苷酸、脱氧核糖核苷酸、核糖核苷酸或脱氧核糖核苷酸的类似物或衍生物的一种或多种组成;包括单链或双链核酸分子。In the present disclosure, the term "template" refers to the nucleic acid molecule to be tested, which means a polymer of nucleotides of a certain length, and the nucleotides may include one or more components of ribonucleotides, deoxyribonucleotides, analogs or derivatives of ribonucleotides or deoxyribonucleotides; including single-stranded or double-stranded nucleic acid molecules.
在本公开中,术语“测序”又可称为“核酸测序”或“基因测序”,指核酸序列中碱基排列顺序的测定;包括双末端测序、单末端测序和/或配对末端测序等,所称的双末端测序或者配对末端测序可以指同一核酸分子的不完全重叠的任意两段或两个部分的读出;所称的测序包括使核苷酸(包括核苷酸类似物)结合到模板并采集相应的反应信号的过程。In the present disclosure, the term "sequencing" may also be referred to as "nucleic acid sequencing" or "gene sequencing", which refers to the determination of the order of base arrangement in a nucleic acid sequence; it includes double-end sequencing, single-end sequencing and/or paired-end sequencing, etc. The so-called double-end sequencing or paired-end sequencing may refer to the reading of any two segments or two parts that are not completely overlapping of the same nucleic acid molecule; the so-called sequencing includes the process of binding nucleotides (including nucleotide analogs) to templates and collecting corresponding reaction signals.
在本公开中,“核苷酸”指的4种天然核苷酸(dATP、dCTP、dGTP、dTTP)或其衍生物,除非另有明确的限定。In the present disclosure, "nucleotide" refers to the four natural nucleotides (dATP, dCTP, dGTP, dTTP) or their derivatives, unless otherwise specifically defined.
在本公开中,术语“引物”是指:可以与感兴趣的靶序列杂交的寡聚核苷酸或核酸分子;引物是单链寡核苷酸或多核苷酸。In the present disclosure, the term "primer" refers to: an oligonucleotide or a nucleic acid molecule that can hybridize to a target sequence of interest; a primer is a single-stranded oligonucleotide or a polynucleotide.
在本公开中,术语“碱基”,又称核碱基、含氮碱基,包括天然碱基、非天然碱基和碱基类似物。其中,天然碱基包括腺嘌呤(A)、鸟嘌呤(G)、胞嘧啶(C)、胸腺嘧啶(T)、尿嘧啶(U);非天然碱基包括诸如锁定核酸(LNA)和桥接核酸(BNA);碱基类似物包括诸如次黄嘌呤、脱氮腺嘌呤、脱氮鸟嘌呤、脱氮次黄嘌呤、7-甲基鸟嘌呤、5,6-二氢尿嘧啶、5-甲基胞嘧啶、5-羟甲基胞嘧啶。本公开中,由于核苷酸类型通过碱基类型来确定,因此,本公开中可以采用碱基类型来表示核苷酸类型。In the present disclosure, the term "base", also known as nucleobase, nitrogenous base, includes natural bases, non-natural bases and base analogs. Among them, natural bases include adenine (A), guanine (G), cytosine (C), thymine (T), uracil (U); non-natural bases include locked nucleic acids (LNA) and bridged nucleic acids (BNA); base analogs include hypoxanthine, deazaadenine, deazaguanine, deazahypoxanthine, 7-methylguanine, 5,6-dihydrouracil, 5-methylcytosine, 5-hydroxymethylcytosine. In the present disclosure, since the nucleotide type is determined by the base type, the base type can be used to represent the nucleotide type in the present disclosure.
在本公开中,术语“滚环扩增”是新近发展起来的一种恒温核酸扩增方法。以环状DNA为模板,通过一个短的DNA引物(与部分环状模板互补),在酶催化下将dNTPs转变成单链DNA,此单链DNA包含成百上千个重复的模板互补片段。In the present disclosure, the term "rolling circle amplification" is a newly developed isothermal nucleic acid amplification method. Using circular DNA as a template, a short DNA primer (complementary to a portion of the circular template) is used to convert dNTPs into single-stranded DNA under enzyme catalysis. The single-stranded DNA contains hundreds or thousands of repeated template-complementary fragments.
在本公开中,术语“DNA多拷贝连环体”,是利用单链DNA环状文库或双链环状DNA文库进行边扩增边连接的反应,形成几百或上千拷贝的内部连锁在一起的共价链环状DNA。In the present disclosure, the term "DNA multi-copy concatemer" refers to the use of a single-stranded DNA circular library or a double-stranded circular DNA library to perform an amplification and ligation reaction to form hundreds or thousands of copies of internally linked covalently linked circular DNA.
在本公开中,术语“DNA解旋酶”,是一类解开氢键的酶,是由水解ATP供给能量来解开DNA的酶。它们常常依赖于单链的存在,并能识别复制叉的单链结构。一般在DNA或RNA复制过程中起到催化双链DNA或RNA解旋的作用。In the present disclosure, the term "DNA helicase" refers to a class of enzymes that unwind hydrogen bonds, and are enzymes that unwind DNA by hydrolyzing ATP to provide energy. They often rely on the presence of single strands and can recognize the single-stranded structure of replication forks. Generally, they play a role in catalyzing the unwinding of double-stranded DNA or RNA during DNA or RNA replication.
在本公开中,术语“单链结合蛋白”,又称DNA结合蛋白(SSB,single strand DNA-binding protein),是DNA复制所必须的。单链结合蛋白不属于酶,大肠杆菌细胞中的单链结合蛋白由4个相同的亚基组成,相对分子质量为74000,结合单链DNA的跨度约32个核苷酸单位;DNA解旋后,DNA分子只要碱基配对,就有结合成双链的趋向。SSB结合于螺旋酶沿复制叉方向向前推进产生的单链区,防止新形成的单链DNA重新配对形成双链DNA或被核酸酶降解的蛋白质。SSB作用时表现协同效应,保证SSB在下游区段的继续结合。它不像聚合酶那样沿着复制方向向前移动,而是不停的结合,脱离。In the present disclosure, the term "single-strand binding protein", also known as DNA binding protein (SSB), is necessary for DNA replication. Single-strand binding protein is not an enzyme. The single-strand binding protein in Escherichia coli cells is composed of 4 identical subunits, with a relative molecular mass of 74,000, and a span of about 32 nucleotide units for binding to single-stranded DNA; after DNA unwinding, as long as the DNA molecules are base-paired, they tend to bind into double strands. SSB binds to the single-stranded region produced by the helicase moving forward in the direction of the replication fork, preventing the newly formed single-stranded DNA from re-pairing to form double-stranded DNA or proteins degraded by nucleases. SSB exhibits a synergistic effect when acting, ensuring that SSB continues to bind to the downstream segment. It does not move forward in the direction of replication like a polymerase, but instead constantly binds and detaches.
在本公开中,术语“外切酶”即核酸外切酶(exonuclease),是一类能从多核苷酸链的一端开始按序催化水解3、5-磷酸二酯键,降解核苷酸的酶。其水解的最终产物是单个的核苷酸(DNA为dNMP,RNA为NMP)。按作用的特性差异可以将其分为单链的核酸外切酶和双链的核酸外切酶。In the present disclosure, the term "exonuclease" refers to a type of enzyme that can sequentially catalyze the hydrolysis of 3, 5-phosphodiester bonds from one end of a polynucleotide chain to degrade nucleotides. The final product of its hydrolysis is a single nucleotide (dNMP for DNA and NMP for RNA). According to the difference in the characteristics of the action, it can be divided into single-stranded exonucleases and double-stranded exonucleases.
在本公开中,术语“Tm”值,熔解温度(melting temperature,Tm)是指吸光值增加到最大值的一半时的温度称为DNA的解链温度或熔点。Tm值就是DNA的熔解温度,是DNA变性过程中紫外吸收达到最大值一半时的温度。不同序列的DNA,Tm值不同,DNA中C-G含量越高,Tm值就越大。In the present disclosure, the term "Tm" value, melting temperature (Tm) refers to the temperature at which the absorbance value increases to half of the maximum value, which is called the melting temperature or melting point of DNA. The Tm value is the melting temperature of DNA, which is the temperature at which the ultraviolet absorption reaches half of the maximum value during the DNA denaturation process. DNA of different sequences has different Tm values. The higher the C-G content in the DNA, the greater the Tm value.
在本公开中,术语“碱基信号采集单元”指单分子模板经过扩增放大成百上千倍后,形成的单克隆群体,能放大测序的碱基的荧光信号,采集的荧光信号代表原单分子模板的信号。In the present disclosure, the term "base signal collection unit" refers to a monoclonal population formed after a single molecule template is amplified hundreds or thousands of times, which can amplify the fluorescent signal of the sequenced base. The collected fluorescent signal represents the signal of the original single molecule template.
在本公开中,术语“链置换”指在核酸上带上被化学修饰的酶切位点,能在被酶切后将链DNA打开缺口,DNA聚合酶继之延伸缺口3'端并替换下一条DNA链。被替换下来的DNA单链可与引物结合并被DNA聚合酶延伸成双链。该过程不断反复进行,使核酸序列被高效扩增。In the present disclosure, the term "strand displacement" refers to a chemically modified restriction site on a nucleic acid, which can open a gap in the DNA strand after being cleaved by the enzyme, and the DNA polymerase then extends the 3' end of the gap and replaces the next DNA strand. The replaced single-stranded DNA can bind to the primer and be extended into a double strand by the DNA polymerase. This process is repeated continuously, so that the nucleic acid sequence is efficiently amplified.
在本公开中,术语“接头”,指一段已知的短的碱基序列。In the present disclosure, the term "linker" refers to a known short base sequence.
核酸扩增的方法Nucleic Acid Amplification Methods
根据本公开的一些具体实施方案,本公开提出了一种核酸扩增的方法,如图1所示,包括:According to some specific embodiments of the present disclosure, the present disclosure provides a method for nucleic acid amplification, as shown in FIG1 , comprising:
S110、将核酸与已知序列相连,获得待扩增核酸;S110, connecting the nucleic acid to a known sequence to obtain the nucleic acid to be amplified;
S120、以所述待扩增核酸中含有的已知序列设计引物对;S120, designing a primer pair based on a known sequence contained in the nucleic acid to be amplified;
S130、利用所述待扩增核酸、所述引物对进行扩增连接反应,获得双链DNA多拷贝连环体。S130, performing an amplification and ligation reaction using the nucleic acid to be amplified and the primer pair to obtain a double-stranded DNA multi-copy concatemer.
根据本公开的一些具体实施方案,本公开提出了一种核酸扩增的方法,如图2所示,包括:According to some specific embodiments of the present disclosure, the present disclosure provides a method for nucleic acid amplification, as shown in FIG2 , comprising:
S140、利用扩增引物扩增目标核酸,获得扩增产物,其中所述扩增引物的5’末端带有已知序列;S140, amplifying the target nucleic acid using an amplification primer to obtain an amplification product, wherein the 5' end of the amplification primer has a known sequence;
S150、以所述扩增产物中含有的已知序列设计引物对,其中,所述引物对中的至少一条引物上含有酶切位点;S150, designing a primer pair based on the known sequence contained in the amplified product, wherein at least one primer in the primer pair contains a restriction site;
S160、利用所述扩增产物、所述引物进行扩增连接反应,获得第一双链DNA多拷贝连环体。S160, performing an amplification and ligation reaction using the amplified product and the primers to obtain a first double-stranded DNA multi-copy concatemer.
根据本公开的一些更为具体的实施方案,本公开提出了一种核酸扩增的方法,如图3所示,包括:According to some more specific embodiments of the present disclosure, the present disclosure provides a method for nucleic acid amplification, as shown in FIG3 , comprising:
1)提取样本总DNA;1) Extract total DNA from samples;
2)用机械法、超声波法或者酶切法,将DNA打断成大小不一的片段,即待测DNA;2) Use mechanical, ultrasonic or enzyme methods to break the DNA into fragments of different sizes, i.e. the DNA to be tested;
3)在片段的至少一端加上接头序列,进行环化反应,获得环状DNA文库;3) adding an adapter sequence to at least one end of the fragment and performing a circularization reaction to obtain a circular DNA library;
4)根据接头,设计引物对,对两条引物5’端进行磷酸化修饰,并使引物对至少一条引物上带有碱基修饰;4) Designing a primer pair based on the linker, performing phosphorylation modification on the 5' ends of the two primers, and making at least one primer of the primer pair have a base modification;
5)将所得引物与所得环化DNA放入一个体系中进行扩增连接反应,最终获得双链DNA多拷贝连环体;5) placing the obtained primers and the obtained circularized DNA into a system for amplification and ligation reaction, and finally obtaining multiple copies of double-stranded DNA concatemers;
6)将所得DNA多拷贝连环体用可识别引物2中修饰位点的消化酶切割,使其中一条环状DNA呈链状,并用单链DNA消化酶消化,最终得到单链DNA多拷贝连环体。6) The obtained multi-copy concatemer of DNA is cut with a digestive enzyme that can recognize the modified site in
根据本公开的一些实施方案,所述引物对中的引物的5’端均含有磷酸化修饰。“引物对中的引物的5’端均含有磷酸化修饰”是指引物对中的每一条引物的5’端都进行了磷酸化修饰,便于连接酶将链状扩增产物的3’端与引物的5’端连接成环。According to some embodiments of the present disclosure, the 5' ends of the primers in the primer pair are all phosphorylated. "The 5' ends of the primers in the primer pair are all phosphorylated" means that the 5' end of each primer in the primer pair is phosphorylated, which facilitates the ligase to connect the 3' end of the chain amplification product to the 5' end of the primer to form a ring.
根据本公开的一些实施方案,所述引物对中的两条引物之间可以没有重叠互补片段,或者,所述引物对中的两条引物至少具有部分重叠互补片段。关于重叠互补片段的长度并没有特别的限制,根据需要进行调整,同时需要保证重叠互补片段的Tm值低于所述两条引物的Tm值。由此,能够确保退火时引物与模板优先结合,而不是两引物之间形成二聚体。According to some embodiments of the present disclosure, there may be no overlapping complementary fragments between the two primers in the primer pair, or the two primers in the primer pair at least have partially overlapping complementary fragments. There is no particular restriction on the length of the overlapping complementary fragments, which can be adjusted as needed, and it is necessary to ensure that the Tm value of the overlapping complementary fragment is lower than the Tm value of the two primers. Thus, it can be ensured that the primers preferentially bind to the template during annealing, rather than forming a dimer between the two primers.
根据本公开的一些实施方案,所述酶切位点包括能够被所述酶切割的的核糖核苷酸或脱氧核糖核苷酸。所述酶切位点包括选自核糖核苷酸、尿嘧啶脱氧核糖核苷酸、5,6二羟基胸腺嘧啶脱氧核糖核苷酸、5-羟基尿嘧啶脱氧核糖核苷酸、5-羟甲基尿嘧啶脱氧核糖核苷酸、5-甲酰基尿嘧啶脱氧核糖核苷酸中的至少之一。需要说明的是,本发明中所述的“酶切位点”并不限于上述类型,也可以是其他类型的酶切位点,只要能够实现酶的特异性切割即可,这类的酶切位点均包含在本发明的酶切位点种类之内。According to some embodiments of the present disclosure, the restriction site includes a ribonucleotide or deoxyribonucleotide that can be cut by the enzyme. The restriction site includes at least one selected from ribonucleotides, uracil deoxyribonucleotides, 5,6-dihydroxythymine deoxyribonucleotides, 5-hydroxyuracil deoxyribonucleotides, 5-hydroxymethyluracil deoxyribonucleotides, and 5-formyluracil deoxyribonucleotides. It should be noted that the "restriction site" described in the present invention is not limited to the above types, and can also be other types of restriction sites. As long as the specific cutting of the enzyme can be achieved, this type of restriction site is included in the type of restriction site of the present invention.
根据本公开的一些实施方案,所述引物对中至少一条引物上带有酶切位点,该酶切位点能被酶切割,在双链DNA连环体任意一条环状DNA上形成缺口。例如,所述引物对中其中一条引物上带有酶切位点,该酶切位点能被酶切割,在双链DNA连环体的含有酶切位点的链上进行切割形成缺口链;或者,所述引物对中两条引物中含有不同的酶切位点,根 据需要采用相应的酶切割双链DNA连环体的酶切位点,形成缺口链。According to some embodiments of the present disclosure, at least one of the primers in the primer pair carries a restriction site, which can be cut by an enzyme to form a gap on any one of the circular DNAs in the double-stranded DNA concatemer. For example, one of the primers in the primer pair carries a restriction site, which can be cut by an enzyme to cut the strand containing the restriction site in the double-stranded DNA concatemer to form a gap strand; or, the two primers in the primer pair contain different restriction sites, and the restriction sites of the double-stranded DNA concatemer are cut with corresponding enzymes as needed to form a gap strand.
根据本公开的一些实施方案,当所述核酸为链状DNA时,步骤(A)进一步包括将与已知序列相连的核酸进行环化反应,获得环化的待扩增核酸。本发明步骤(A)中,核酸与已知序列相连后获得的待扩增核酸优选为环化的核酸,这样便于扩增链的5’端和3’端连接成环,提升连接效率。According to some embodiments of the present disclosure, when the nucleic acid is a chain DNA, step (A) further comprises cyclizing the nucleic acid connected to the known sequence to obtain a cyclized nucleic acid to be amplified. In step (A) of the present invention, the nucleic acid to be amplified obtained after the nucleic acid is connected to the known sequence is preferably a cyclized nucleic acid, which facilitates the connection of the 5' end and the 3' end of the amplified chain into a ring and improves the connection efficiency.
根据本公开的一些实施方案,当所述核酸为环状DNA时,需要先将环状DNA线性化,获得链状DNA后,在其两端加上已知接头序列,进一步环化反应,获得环化的待扩增核酸。According to some embodiments of the present disclosure, when the nucleic acid is a circular DNA, it is necessary to first linearize the circular DNA to obtain a chain DNA, and then add known linker sequences to both ends thereof for further circularization reaction to obtain a circularized nucleic acid to be amplified.
根据本公开的一些实施方案,可以通过下述方法获得步骤(c)或(D)中所述第一单链DNA多拷贝连环体后的另一条反向单链DNA多拷贝连环体:According to some embodiments of the present disclosure, another reverse single-stranded DNA multi-copy concatemer following the first single-stranded DNA multi-copy concatemer in step (c) or (D) can be obtained by the following method:
方法一method one
制备DNA多拷贝连环体的2条引物具有不同类型的酶切位点,例如引物1带有尿嘧啶脱氧核糖核苷酸,引物2带有核糖核苷酸。在制备一链DNA多拷贝连环体时,在完成扩增连接的放大反应后,使用RNase H将引物2的核糖核苷酸切除,使由引物2扩增的链产生缺口,再利用DNA消化酶将其去除,得到第一单链DNA多拷贝连环体。在完成一链测序以后,洗脱测序过程中生成的互补序列。以引物2进行扩增,扩增一圈后进行连接反应,生成二链的DNA多拷贝连环体。再使用USER酶将一链切开形成缺口,再利用DNA消化酶将整个一链的DNA多拷贝连环体去除。最后得到单链状态的二链DNA多拷贝连环体。The two primers for preparing the multi-copy DNA concatemer have different types of restriction sites, for example,
方法二Method Two
在步骤(C)后,利用酶2切割所述双链DNA多拷贝连环体中含有的酶切位点2,在双链DNA环中形成缺口单链,利用消化酶对所述缺口单链进行消化,以便获得第二单链DNA多拷贝连环体。After step (C),
方法三Method 3
以步骤(c)或(D)获得的DNA多拷贝连环体为模板,用与第一单链DNA多拷贝连环体接头通用序列互补的引物,使用Phi29酶进行扩增,依靠Phi29酶的链置换能力,在单链环上扩增一圈后,置换出与多拷贝连环体互补的第二单链DNA多拷贝连环体。The multi-copy concatemer of DNA obtained in step (c) or (D) is used as a template, and a primer complementary to the universal sequence of the first single-stranded DNA multi-copy concatemer linker is used to amplify using the Phi29 enzyme. Relying on the chain displacement ability of the Phi29 enzyme, after one circle of amplification on the single-stranded loop, a second single-stranded DNA multi-copy concatemer complementary to the multi-copy concatemer is displaced.
根据本公开的一些实施方案,所述酶1,酶2分别包括选自USER酶、RNase H、UDG、RNase HII、hSMUG1中至少一种,且酶1、酶2为不同的酶。According to some embodiments of the present disclosure, the
根据本公开的实施方案,所述扩增连接反应的体系中含有DNA聚合酶,所述DNA聚合酶不具有5’-3’外切酶活性。所述DNA聚合酶不具有5’到3’外切酶活性,能够保证扩增的效率,防止扩增产物被酶切。According to the embodiments of the present disclosure, the amplification and ligation reaction system contains a DNA polymerase, and the DNA polymerase does not have 5'-3' exonuclease activity. The DNA polymerase does not have 5' to 3' exonuclease activity, which can ensure the efficiency of amplification and prevent the amplification product from being cleaved by enzymes.
根据本公开的实施方案,所述扩增连接反应的体系中含有DNA聚合酶,所述DNA聚合酶包括选自KAPA HiFi HotStart DNA Polymerase、KAPA HiFi HotStart Uracil+Polymerase、BGI Golden High-Fidelity Polymerase、BGI Platinum HiFi Hotstart Polymerase中的至少之一。According to the embodiments of the present disclosure, the amplification and ligation reaction system contains a DNA polymerase, and the DNA polymerase includes at least one selected from KAPA HiFi HotStart DNA Polymerase, KAPA HiFi HotStart Uracil+Polymerase, BGI Golden High-Fidelity Polymerase, and BGI Platinum HiFi Hotstart Polymerase.
根据本公开的实施方案,所述扩增连接反应的体系中含有耐高温DNA连接酶。According to an embodiment of the present disclosure, the amplification and ligation reaction system contains a high temperature resistant DNA ligase.
根据本公开的实施方案,所述耐高温DNA连接酶包括选自Taq ligase、AMP ligase中的至少之一。According to an embodiment of the present disclosure, the high temperature resistant DNA ligase includes at least one selected from Taq ligase and AMP ligase.
根据本公开的实施方案,所述扩增连接反应的体系中含有不耐高温DNA连接酶。According to an embodiment of the present disclosure, the amplification and ligation reaction system contains a high-temperature-intolerant DNA ligase.
根据本公开的实施方案,所述不耐高温DNA连接酶选自T4 DNA ligase、E.coli DNA Ligase、T3 DNA Ligase、T7 DNA Ligase中的至少之一。According to the embodiments of the present disclosure, the heat-intolerant DNA ligase is selected from at least one of T4 DNA ligase, E. coli DNA Ligase, T3 DNA Ligase, and T7 DNA Ligase.
根据本公开的实施方案,所述扩增连接反应的体系中含有不耐高温DNA连接酶时,进 一步包括DNA解旋酶、单链结合蛋白。体系中含有的单链结合蛋白能够防止解旋的DNA自我配对。According to the embodiments of the present disclosure, when the amplification and ligation reaction system contains a thermostable DNA ligase, it further includes a DNA helicase and a single-stranded binding protein. The single-stranded binding protein contained in the system can prevent the unwound DNA from self-pairing.
根据本公开的实施方案,所述酶1、酶2包括能够切割所述核糖核苷酸和/或尿嘧啶脱氧核糖核苷酸的酶。According to an embodiment of the present disclosure, the
根据本公开的实施方案,所述酶1、酶2包括选自USER酶、RNase H、UDG、RNase HII、hSMUG1、APE1、Endonuclease VIII、Endonuclease III(Nth)至少之一,且酶1、酶2为不同的酶。According to the embodiments of the present disclosure, the
根据本公开的实施方案,所述消化酶包括能够消化单链DNA的酶。According to an embodiment of the present disclosure, the digestive enzyme comprises an enzyme capable of digesting single-stranded DNA.
根据本公开的实施方案,所述消化酶包括DNA外切酶。According to an embodiment of the present disclosure, the digestive enzyme comprises a DNA exonuclease.
根据本公开的实施方案,所述DNA外切酶包括选自DNA外切酶Ⅰ、DNA外切酶Ⅲ、T5 Exonuclease、Exonuclease T、Exonuclease VII、Lambda Exonuclease中的至少之一。According to the embodiments of the present disclosure, the DNA exonuclease includes at least one selected from DNA exonuclease I, DNA exonuclease III, T5 Exonuclease, Exonuclease T, Exonuclease VII, and Lambda Exonuclease.
根据本公开的一些更为具体的实施方案,所述核酸扩增的方法(图4),包括:According to some more specific embodiments of the present disclosure, the method for nucleic acid amplification ( FIG. 4 ) comprises:
1)使用带有已知接头序列的单链环状DNA文库,或者双链环状DNA文库作为模板。扩增引物根据已知接头进行设计的,2条扩增引物之间可以有重叠互补(互补的片段的Tm值要低于2条引物的Tm值),也可以没有重叠互补的区域。引物的5’端进行磷酸化修饰,并且其中一条引物包括可以后续被对应酶切除的修饰核苷酸,例如尿嘧啶脱氧核糖核苷酸或者核糖核苷酸。1) Use a single-stranded circular DNA library with a known linker sequence or a double-stranded circular DNA library as a template. The amplification primers are designed based on the known linker, and the two amplification primers may have overlapping complementary regions (the Tm value of the complementary fragment is lower than the Tm value of the two primers), or there may be no overlapping complementary regions. The 5' end of the primer is phosphorylated, and one of the primers includes a modified nucleotide that can be subsequently removed by the corresponding enzyme, such as uracil deoxyribonucleotide or ribonucleotide.
2)对单链环状DNA文库或者双链环状DNA文库模板进行扩增和连接反应,并且扩增和连接反应是在一个反应体系中进行。反应体系中除模板包括扩增引物,DNA聚合酶,DNA连接酶,dNTP,Mg 2+离子等,其中DNA聚会酶不具有5’-3’外切酶活性,例如KAPA HiFi HotStart DNA Polymerase或KAPA HiFi HotStart Uracil+Polymerase,DNA连接酶可以是耐高温连接酶,如Taq ligase,AMPligase。也可以是不耐高温的连接酶,如T4 DNA ligase。当使用不耐高温的连接酶时,使用等温扩增体系,可以包括DNA解旋酶,单链结合蛋白等。 2) Amplification and ligation reaction are performed on the single-stranded circular DNA library or double-stranded circular DNA library template, and the amplification and ligation reaction are performed in a reaction system. In addition to the template, the reaction system includes amplification primers, DNA polymerase, DNA ligase, dNTP, Mg 2+ ions, etc., wherein the DNA polymerase does not have 5'-3' exonuclease activity, such as KAPA HiFi HotStart DNA Polymerase or KAPA HiFi HotStart Uracil+Polymerase, and the DNA ligase can be a thermostable ligase, such as Taq ligase, AMPligase. It can also be a thermostable ligase, such as T4 DNA ligase. When a thermostable ligase is used, an isothermal amplification system is used, which may include DNA helicase, single-stranded binding protein, etc.
3)模板在扩增和连接反应的体系中进行反应,经过变性,退火,延伸,连接的过程,其中退火延伸和连接可以在统一的温度下进行,如果采用等温扩增,变性、退火、延伸和连接可以再相同温度下进行,可以利用DNA解旋酶实现DNA变性解链的效果。其中引物于模板进行退火后,DNA聚合酶进行延伸,当延伸到引物的5’端的时候停止,DNA连接酶将延伸链的3’端与引物自身的5’端进行连接,形成共价双链环状DNA。共价双链环状DNA的2条链时相互连锁的,即使进行变性也无法将两条链分离开,2条链会保持纠缠在一起。当再次进行循环扩增连接时,所有的产物都相互连锁在一起,不会分开,最终得到来自同一模板的双链的DNA多拷贝连环体。3) The template reacts in the amplification and ligation reaction system, and goes through the process of denaturation, annealing, extension, and ligation. Annealing, extension, and ligation can be performed at a uniform temperature. If isothermal amplification is used, denaturation, annealing, extension, and ligation can be performed at the same temperature. DNA helicase can be used to achieve the effect of DNA denaturation and unwinding. After the primer is annealed to the template, the DNA polymerase extends and stops when it extends to the 5' end of the primer. The DNA ligase connects the 3' end of the extended chain with the 5' end of the primer itself to form a covalent double-stranded circular DNA. The two chains of the covalent double-stranded circular DNA are interlinked, and even if denaturation is performed, the two chains cannot be separated, and the two chains will remain entangled. When the cyclic amplification and ligation are performed again, all the products are interlinked and will not separate, and finally a double-stranded DNA multi-copy concatemer from the same template is obtained.
4)对得到的双链的多拷贝的DNA连环体的产物进行消化处理,可以得到单链的DNA多拷贝连环体。即使用一系列消化酶切除其中一条引物上带有尿嘧啶脱氧核糖核苷酸或核糖核苷酸等这些修饰,例如使用USER酶切除U碱基,使用RNase H切除核糖核苷酸。让双链DNA链中一条链产生缺口,再利用单链DNA消化酶,例如DNA外切酶Ⅰ、DNA外切酶Ⅲ,对缺口单链进行消化。这些消化反应可以存在于同一反应体系,也可以在不同反应中。对经过消化处理的产物纯化就可以得到单链的DNA多拷贝连环体。4) The obtained double-stranded multi-copy DNA concatemer product is digested to obtain a single-stranded DNA multi-copy concatemer. That is, a series of digestive enzymes are used to remove modifications such as uracil deoxyribonucleotides or ribonucleotides on one of the primers, such as using USER enzyme to remove U bases and using RNase H to remove ribonucleotides. A gap is created in one of the double-stranded DNA chains, and then a single-stranded DNA digestive enzyme, such as DNA exonuclease I and DNA exonuclease III, is used to digest the gapped single chain. These digestion reactions can exist in the same reaction system or in different reactions. Purification of the digested product can obtain a single-stranded DNA multi-copy concatemer.
根据本公开的一些具体实施方案,本公开提出了一种二链生产方法,所述方法包括,以单链DNA多拷贝连环体为模板,与已知核酸序列全部或部分互补的引物杂交,在聚合酶的作用下进行滚环扩增反应,生成二链。According to some specific embodiments of the present disclosure, the present disclosure proposes a method for producing a second chain, which includes using a single-stranded DNA multi-copy concatemer as a template, hybridizing with a primer that is fully or partially complementary to a known nucleic acid sequence, and performing a rolling circle amplification reaction under the action of a polymerase to generate a second chain.
根据本公开的实施方案,所述引物固定在固体支持物上,所述固体支持物可以选自包括:磁珠、多聚物、水凝胶、金属或芯片。According to an embodiment of the present disclosure, the primer is fixed on a solid support, and the solid support may be selected from the group consisting of: magnetic beads, polymers, hydrogels, metals or chips.
根据本公开的实施方案,所述引物游离在溶液中。According to an embodiment of the present disclosure, the primer is free in the solution.
在测序领域中的应用Application in sequencing
根据本公开的实施方案,在所述待扩增模板两端的已知接头中间加上一段随机碱基序列(标签序列),以上述核酸扩增方法获得带有两段已知接头序列、一段标签序列、一段待测序列为测序模板的单链DNA多拷贝连环体(图5),将此含有成百上千拷贝的集合体,作为碱基信号单元,加载到测序芯片上。According to the implementation scheme of the present disclosure, a random base sequence (label sequence) is added in the middle of the known linkers at both ends of the template to be amplified, and the above-mentioned nucleic acid amplification method is used to obtain a single-stranded DNA multi-copy concatemer with two known linker sequences, a label sequence, and a sequence to be tested as a sequencing template (Figure 5). This collection containing hundreds or thousands of copies is loaded onto a sequencing chip as a base signal unit.
根据本公开的实施方案,合成与所述接头序列互补的引物1,利用碱基互补原则,引物1的5’端与接头序列配对,依次往后进行边合成边测序反应。According to the embodiments of the present disclosure, a
根据本公开的具体实施方案,当完成一链测序后,可以以下述方法得到二链:According to a specific embodiment of the present disclosure, after completing the sequencing of one strand, the second strand can be obtained by the following method:
方法一method one
以当前一链的DNA多拷贝连环体为模板,使用接头作为引物结合位点,使用Phi29酶进行扩增,依靠Phi29酶的链置换能力,在单链环上扩增一圈后,置换出与多拷贝连环体互补的二链。Using the current single-strand DNA multi-copy concatemer as a template, the adapter as a primer binding site, and the Phi29 enzyme for amplification, relying on the chain displacement ability of the Phi29 enzyme, after amplifying one circle on the single-stranded loop, a second chain complementary to the multi-copy concatemer is displaced.
方法二Method Two
制备DNA多拷贝连环体的2条引物具有不同类型的酶切位点,例如引物1带有尿嘧啶脱氧核糖核苷酸,引物2带有核糖核苷酸。在制备一链DNA多拷贝连环体时,在完成扩增连接的放大反应后,使用RNase H将引物2的核糖核苷酸切除,使由引物2扩增的链产生缺口,再利用DNA消化酶将其去除,得到单链环的DNA多拷贝连环体。在完成一链测序以后,洗脱测序过程中生成的互补序列。以引物2进行扩增,扩增一圈后进行连接反应,生成二链的DNA多拷贝连环体。再使用USER酶将一链切开形成缺口,再利用DNA消化酶将整个一链的DNA多拷贝连环体去除。最后得到单链状态的二链DNA多拷贝连环体。The two primers for preparing the multi-copy DNA concatemer have different types of restriction sites, for example,
所述引物1和引物2带有不同的酶切位点即可,没有顺序之分,除U碱基,核糖核酸碱基以外,还可以是5,6二羟基胸腺嘧啶,5-hydroxyuracil,5-hydroxymethyluracil,and 5-formyluracil等,所述酶1、酶2包括选自USER酶、RNase H、UDG、RNase HII、hSMUG1中的至少之一,且酶1、酶2为不同的酶。The
上述核酸扩增方法得到的单链DNA多拷贝连环体,作为测序中的碱基信号采集单元,加载到测序芯片上,有以下优势:DNA多拷贝连环体的DNA测序模板拷贝数多,且其体积小,从而占用测序芯片或流动槽的面积小;DNA测序模板的拷贝数的增加也不会影响其体积,DNA测序模板的片段长度对DNA多拷贝连环体的体积影响小,有利于提升测序读长。并且DNA多拷贝连环体的结构稳定,方便操作以及长期存放,并且对反应用到的DNA聚合酶要求低。DNA多拷贝连环体不需要再测序芯片或流动槽上进行反应,对芯片的要求低,不需要在芯片上生成纳米井并在芯片上预先种植寡核苷酸。The single-stranded DNA multi-copy concatemer obtained by the above-mentioned nucleic acid amplification method is loaded onto a sequencing chip as a base signal acquisition unit in sequencing, and has the following advantages: the DNA sequencing template of the DNA multi-copy concatemer has a large number of copies and a small volume, thereby occupying a small area of the sequencing chip or the flow tank; the increase in the number of copies of the DNA sequencing template will not affect its volume, and the fragment length of the DNA sequencing template has a small effect on the volume of the DNA multi-copy concatemer, which is conducive to improving the sequencing read length. In addition, the structure of the DNA multi-copy concatemer is stable, convenient for operation and long-term storage, and has low requirements for the DNA polymerase used in the reaction. The DNA multi-copy concatemer does not need to react on the sequencing chip or the flow tank, has low requirements for the chip, and does not need to generate nanowells on the chip and pre-plant oligonucleotides on the chip.
上述核酸扩增方法,不仅适用于测序领域,还适用于任何微量扩增领域。当实验材料较少,样品较为稀缺时,可用本公开方法,在不改变核酸丰度的同时,高效放大样本核酸含量。The above-mentioned nucleic acid amplification method is not only applicable to the field of sequencing, but also to any field of micro-amplification. When there are few experimental materials and the samples are scarce, the disclosed method can be used to efficiently amplify the sample nucleic acid content without changing the nucleic acid abundance.
实施方案中未注明具体技术或条件的,按照本领域内的文献所描述的技术或条件或者按照产品说明书进行。所用试剂或仪器未注明生产厂商者,均为可以通过市购获得的常规 产品。If no specific technology or conditions are specified in the implementation plan, the technology or conditions described in the literature in this field or the product instructions shall be followed. If the manufacturer of the reagents or instruments used is not specified, they are all conventional products that can be purchased commercially.
实施例1大肠杆菌基因组DNA测序Example 1 Escherichia coli genomic DNA sequencing
实施例中实验材料为大肠杆菌基因组DNAThe experimental material in the embodiment is Escherichia coli genomic DNA
1.实验步骤1. Experimental Procedure
(1)准备MGIEasy酶切DNA文库制备试剂套装(16RXN)(MGI,货号1000006987);(1) Prepare MGIEasy DNA Library Preparation Kit (16RXN) (MGI, Catalog No. 1000006987);
(2)取100ng的大肠杆菌基因组DNA,按照试剂盒说明书,将DNA打断到主带在300到500bp大小,经过磁珠纯化,片段选择,末端修复加A,接头连接、PCR扩增,最后使用试剂盒中的单链环化部分的试剂,对PCR扩增产物进行单链环化,得到最终的适合DNBSEQ测序平台的单链环状DNA文库;(2) Take 100 ng of E. coli genomic DNA, and according to the kit instructions, shear the DNA to a main band of 300 to 500 bp in size, and then undergo magnetic bead purification, fragment selection, end repair and A addition, adapter ligation, and PCR amplification. Finally, use the reagents in the single-stranded circularization part of the kit to circularize the PCR amplification product to obtain the final single-stranded circular DNA library suitable for the DNBSEQ sequencing platform;
(3)取制备好的单链环状DNA文库5ng到200μLPCR管中;(3) Take 5 ng of the prepared single-stranded circular DNA library into a 200 μL PCR tube;
(4)按照表1在冰盒上配制扩增连接反应的反应液;(4) Prepare the amplification and ligation reaction solution on an ice box according to Table 1;
表1Table 1
Primer1:5’P-GCCATGTCGTTCTGTGAGCCAAGG-3’(SEQ ID NO:1)Primer1:5’P-GCCATGTCGTTCTGTGAGCCAAGG-3’(SEQ ID NO:1)
Primer2:5’P-GCTCACAGAA/rC//rG//rA//rC/ATGGCTACGATCCGAC-3’(SEQ ID NO:2)Primer2: 5’P-GCTCACAGAA/rC//rG//rA//rC/ATGGCTACGATCCGAC-3’ (SEQ ID NO: 2)
其中r代表核糖核苷酸的修饰,P代表磷酸化修饰。Where r represents the modification of ribonucleotide and P represents phosphorylation modification.
(5)将配制完成的反应液加入装有单链环状DNA文库的PCR管中,再用无核酸酶水(ThermoFisher,AM9930)补充到最终体积50μL。使用振荡混匀仪进行充分混匀,再瞬时离心,将液体离心到管底;(5) Add the prepared reaction solution to the PCR tube containing the single-stranded circular DNA library, and then add nuclease-free water (ThermoFisher, AM9930) to a final volume of 50 μL. Use an oscillating mixer to mix thoroughly, then centrifuge briefly to centrifuge the liquid to the bottom of the tube;
(6)将离心好的PCR管放置到PCR仪上,按照表2反应程序进行反应;(6) Place the centrifuged PCR tube on a PCR instrument and perform the reaction according to the reaction program in Table 2;
表2Table 2
(7)反应结束后,取出PCR管,瞬时离心后,将PCR管放置在冰盒上;(7) After the reaction is completed, take out the PCR tube, centrifuge it briefly, and place it on an ice box;
(8)按照表3在冰盒上配制扩增消化酶反应液;(8) Prepare the amplification digestion enzyme reaction solution on an ice box according to Table 3;
表3table 3
(9)将配制完成的消化酶反应液加入上一步的PCR管中。使用振荡混匀仪进行充分混匀,再瞬时离心,将液体离心到管底;(9) Add the prepared digestion enzyme reaction solution to the PCR tube from the previous step. Use an oscillating mixer to mix thoroughly, then centrifuge briefly to centrifuge the liquid to the bottom of the tube;
(10)将离心好的PCR管放置到PCR仪上,按照表4反应程序进行反应;(10) Place the centrifuged PCR tube on a PCR instrument and perform the reaction according to the reaction program in Table 4;
表4Table 4
(11)反应结束后,从PCR仪中取出PCR管,瞬时离心将反应液收集至管底;(11) After the reaction is completed, remove the PCR tube from the PCR instrument and centrifuge it briefly to collect the reaction solution at the bottom of the tube;
(12)将反应产物转移到1.5mL离心管中。之后加入150μL的纯化磁珠,用移液器吹打10次到充分混匀后,常温孵育10min;(12) Transfer the reaction product to a 1.5 mL centrifuge tube. Then add 150 μL of purified magnetic beads,
(13)瞬时离心后,将1.5mL离心管放置与磁力架上,静置5min至液体澄清,用移液器吸取并丢弃上清液;(13) After instant centrifugation, place the 1.5 mL centrifuge tube on a magnetic stand and let it stand for 5 min until the liquid becomes clear. Use a pipette to aspirate and discard the supernatant.
(14)向1.5mL离心管中加入200μL 80%乙醇,静置30s,洗涤磁珠;(14) Add 200 μL 80% ethanol to a 1.5 mL centrifuge tube and let stand for 30 seconds to wash the magnetic beads;
(15)重复上一步;(15) Repeat the previous step;
(16)尽量吸干管内液体,有少量残留在管壁时可将1.5mL离心管瞬时离心,在磁力架上分离后,用小量程的移液器将管底液体吸干;(16) Try to drain the liquid in the tube. If there is a small amount of liquid left on the tube wall, centrifuge the 1.5 mL centrifuge tube for a short time. After separation on the magnetic rack, use a small-range pipette to drain the liquid at the bottom of the tube.
(17)保持1.5mL离心管固定于磁力架上,打开1.5mL离心管管盖,室温干燥,直至磁珠表面无反光、无开裂;(17) Keep the 1.5 mL centrifuge tube fixed on the magnetic rack, open the cap of the 1.5 mL centrifuge tube, and dry it at room temperature until the surface of the magnetic beads is not reflective and has no cracks;
(18)将1.5mL离心管从磁力架上取下,加入26μL分子级水进行DNA洗脱,用移液器轻轻吹打10次至完全混匀;(18) Remove the 1.5 mL centrifuge tube from the magnetic stand, add 26 μL of molecular grade water to elute the DNA, and gently pipette 10 times to mix thoroughly;
(19)室温下孵育5min;(19) Incubate at room temperature for 5 min;
(20)瞬时离心,将1.5mL离心管置于磁力架上,静置5min至液体澄清,将25μL上清液转移到新的1.5mL离心管中;(20) Centrifuge briefly, place the 1.5 mL centrifuge tube on a magnetic rack, let stand for 5 min until the liquid becomes clear, and transfer 25 μL of the supernatant to a new 1.5 mL centrifuge tube;
(21)使用 ssDNA Assay Kit荧光定量试剂盒,按照定量试剂盒的操作说明对酶切消化纯化后产物进行定量; (21) Use ssDNA Assay Kit fluorescent quantitative kit, according to the operating instructions of the quantitative kit to quantify the product after enzyme digestion and purification;
(22)取2μL的产物进行电泳检测,使用已经预先加入SYBR Safe DNA凝胶染料(Thermo fisher,S33102)的1.5%的琼脂糖凝胶,再110V电压,跑45分钟,之后再使用bio-rad伯乐Gel Doc XR+凝胶成像系统进行拍照;(22) Take 2 μL of the product for electrophoresis detection, use 1.5% agarose gel that has been pre-added with SYBR Safe DNA gel dye (Thermo Fisher, S33102), run at 110 V for 45 minutes, and then use the Bio-Rad Gel Doc XR+ gel imaging system to take pictures;
(23)准备MGISEQ-2000高通量测序试剂套装(SE100)(MGI,1000012551),按照说明书进行操作;(23) Prepare MGISEQ-2000 high-throughput sequencing reagent set (SE100) (MGI, 1000012551) and operate according to the instructions;
(24)加载过程,只使用试剂盒中的DNB加载缓冲液Ⅱ,吸取对应体积加入制备好的DNA多拷贝连环体产物的PCR管中。之后按照说明书将DNA多拷贝连环体产物加载到测序芯片上后进行测序;(24) During the loading process, only the DNB loading buffer II in the kit was used, and the corresponding volume was added to the PCR tube containing the prepared DNA multi-copy concatemer product. Then, the DNA multi-copy concatemer product was loaded onto the sequencing chip according to the instructions and sequenced;
(25)测序完成后使用原子力显微镜观察测序芯片上加载的DNA多拷贝连环体;(25) After sequencing is completed, atomic force microscopy is used to observe the multi-copy concatemers of DNA loaded on the sequencing chip;
2.实验结果2. Experimental results
(1)单链DNA测序文库经过反应之后得到DNA多拷贝连环体的浓度符合预期的浓度(表5);(1) After the single-stranded DNA sequencing library was reacted, the concentration of the DNA multi-copy concatemers was consistent with the expected concentration (Table 5);
表5单链DNA文库投入量与DNC产量Table 5 Input amount of single-stranded DNA library and DNC yield
(2)电泳图(图6)显示得到的产物的条带过大(~160k左右),没有跑出点样孔,符合预期;(2) The electrophoresis diagram (Figure 6) showed that the band of the obtained product was too large (about ∼160 k) and did not run out of the spotting well, which was in line with expectations;
(3)使用DNBSEQ的测序试剂与芯片进行测序后,得到415M以上的读长序列,Q30质量值能到89%以上,达到预期目标;(3) After sequencing using DNBSEQ sequencing reagents and chips, a read length of more than 415M was obtained, and the Q30 quality value was more than 89%, achieving the expected goal;
(4)使用原子力显微镜对测序后的芯片进行观察,可以观察到明显小于现有测序芯片的200nm直径的修饰范围(图7,图8)。(4) Using an atomic force microscope to observe the chip after sequencing, it can be observed that the modification range is significantly smaller than the 200 nm diameter of the existing sequencing chip (Figure 7, Figure 8).
本实施例,以大肠杆菌基因组DNA为材料,采用本公开核酸扩增的方法,将得到的扩增产物加载到芯片上进行测序,并对结果进行直接质检,符合预期。In this example, Escherichia coli genomic DNA was used as material, the nucleic acid amplification method disclosed in the present invention was adopted, the obtained amplification product was loaded onto a chip for sequencing, and the results were directly quality checked, which was in line with expectations.
本实施例从核酸模板的获取,到扩增产物(单链DNA多拷贝连环体)的获得,全程操作简单,所需仪器耗材,也均为本领域实验人员方便获得的,提高实验效率的同时,降低了实验成本;由表1可知少量DNA文库的投入即可获得大量DNC,图6可知所得DNC质量达标,故而该方法在高效扩增核酸模板的同时,也能保证扩增产物的质量;由图7,图8可知,在测序中,本公开方法可以明显减少芯片占用面积,相比现有技术,在同等面积的芯片下,能有效提升测序读长,且因模板扩增不在芯片上进行,故而对芯片要求相对较低。In this embodiment, the whole process from obtaining the nucleic acid template to obtaining the amplified product (single-stranded DNA multi-copy concatemer) is simple to operate, and the required instruments and consumables are also conveniently available to experimental personnel in the field, which improves the experimental efficiency and reduces the experimental cost. It can be seen from Table 1 that a large amount of DNC can be obtained with the investment of a small amount of DNA library, and it can be seen from Figure 6 that the quality of the obtained DNC meets the standard, so this method can efficiently amplify the nucleic acid template while ensuring the quality of the amplified product. It can be seen from Figures 7 and 8 that in sequencing, the method disclosed in the present invention can significantly reduce the chip occupied area. Compared with the prior art, under the chip of the same area, the sequencing read length can be effectively improved, and because the template amplification is not performed on the chip, the chip requirements are relatively low.
实施例2 EGFR基因肿瘤热点区域测序Example 2 Sequencing of EGFR gene tumor hotspot regions
实施例中实验材料为人基因组DNAThe experimental material in the embodiment is human genomic DNA
1.实验步骤1. Experimental Procedure
(1)设计EGFR基因肿瘤热点区域扩增引物池,该引物池包含8对特异性寡核酸引物(EGFR_1F-EGFR_8F)和(EGFR_1R-EGFR_8R),扩增子大小为100~200bp,特异性寡核酸引物序列如下表6所示。(1) Design a primer pool for amplifying the tumor hotspot region of the EGFR gene, which contains 8 pairs of specific oligonucleotide primers (EGFR_1F-EGFR_8F) and (EGFR_1R-EGFR_8R), with an amplicon size of 100 to 200 bp. The sequences of the specific oligonucleotide primers are shown in Table 6 below.
表6Table 6
注:上述特异性寡核酸引物按照每条特异性寡核酸引物2μM浓度进行混合,得到总浓度为 2μM的特异性寡核酸引物池;Note: The above-mentioned specific oligonucleotide primers were mixed at a concentration of 2 μM for each specific oligonucleotide primer to obtain a specific oligonucleotide primer pool with a total concentration of 2 μM;
(2)取5ng人基因组DNA到PCR管中,按照(表7)所示配置PCR反应体系进行PCR反应,反应程序如下:94℃ 1min;94℃ 30s,58℃ 2min,72℃ 30s,15cycles;72℃ 5min;4℃∞。(2) Take 5 ng of human genomic DNA into a PCR tube and perform PCR reaction according to the PCR reaction system configured as shown in (Table 7). The reaction procedure is as follows: 94℃ for 1 min; 94℃ for 30 s, 58℃ for 2 min, 72℃ for 30 s, 15 cycles; 72℃ for 5 min; 4℃∞.
表7Table 7
(3)反应完后用MGIEasy酶切DNA文库制备试剂套装(16RXN)(MGI,货号1000006987)中的磁珠进行纯化,最后将纯化产物溶于22μL洗脱缓冲液,得到PCR产物。(3) After the reaction, the DNA was purified using the magnetic beads in the MGIEasy DNA Library Preparation Reagent Kit (16RXN) (MGI, Catalog No. 1000006987). Finally, the purified product was dissolved in 22 μL of elution buffer to obtain the PCR product.
(4)取300ng PCR扩增产物,使用MGIEasy酶切DNA文库制备试剂套装(16RXN)(MGI,货号1000006987)中的单链环化部分的试剂进行单链环化,得到适合DNBSEQ测序平台的单链环状DNA文库;(4) Take 300 ng of PCR amplification product and use the single-stranded circularization reagent in the MGIEasy DNA Library Preparation Reagent Set (16RXN) (MGI, Catalog No. 1000006987) to perform single-stranded circularization to obtain a single-stranded circular DNA library suitable for the DNBSEQ sequencing platform;
(5)取制备好的单链环状DNA文库5ng到200μL PCR管中;(5) Take 5 ng of the prepared single-stranded circular DNA library into a 200 μL PCR tube;
(6)按照表8在冰盒上配制扩增连接反应的反应液;(6) Prepare the reaction solution for amplification and ligation reaction on an ice box according to Table 8;
表8Table 8
Primer1:5’P-GCCATGTCGTTCUGUGAGCCAAGG-3’(SEQ ID NO:19)Primer1:5’P-GCCATGTCGTTCUGUGAGCCAAGG-3’(SEQ ID NO:19)
Primer2:5’P-GCTCACAGAA/rC//rG//rA//rC/ATGGCTACGATCCGAC-3’(SEQ ID NO:20)Primer2:5’P-GCTCACAGAA/rC//rG//rA//rC/ATGGCTACGATCCGAC-3’(SEQ ID NO:20)
其中r代表核糖核苷酸的修饰;U代表脱氧核糖尿苷酸;P代表磷酸化修饰。Among them, r represents the modification of ribonucleotide; U represents deoxyribonucleic acid; and P represents phosphorylation modification.
(7)将配制完成的反应液加入装有单链环状DNA文库的PCR管中,再用无核酸酶水(ThermoFisher,AM9930)补充到最终体积50μL。使用振荡混匀仪进行充分混匀,再瞬时离心,将液体离心到管底;(7) Add the prepared reaction solution to the PCR tube containing the single-stranded circular DNA library, and then add nuclease-free water (ThermoFisher, AM9930) to a final volume of 50 μL. Use an oscillating mixer to mix thoroughly, then centrifuge briefly to centrifuge the liquid to the bottom of the tube;
(8)将离心好的PCR管放置到PCR仪上,按照表9反应程序进行反应;(8) Place the centrifuged PCR tube on a PCR instrument and perform the reaction according to the reaction program in Table 9;
表9Table 9
(9)反应结束后,取出PCR管,瞬时离心后,将PCR管放置在冰盒上;(9) After the reaction is completed, take out the PCR tube, centrifuge it briefly, and place it on an ice box;
(10)按照表10在冰盒上配制扩增消化酶反应液;(10) Prepare the amplification digestion enzyme reaction solution on an ice box according to Table 10;
表10Table 10
(11)将配制完成的消化酶反应液加入上一步的PCR管中。使用振荡混匀仪进行充分混匀,再瞬时离心,将液体离心到管底;(11) Add the prepared digestion enzyme reaction solution to the PCR tube from the previous step. Use an oscillating mixer to mix thoroughly, then centrifuge briefly to centrifuge the liquid to the bottom of the tube;
(12)将离心好的PCR管放置到PCR仪上,按照表11反应程序进行反应;(12) Place the centrifuged PCR tube on a PCR instrument and perform the reaction according to the reaction program in Table 11;
表11Table 11
(13)反应结束后,从PCR仪中取出PCR管,瞬时离心将反应液收集至管底;(13) After the reaction is completed, remove the PCR tube from the PCR instrument and centrifuge briefly to collect the reaction solution at the bottom of the tube;
(14)将反应产物转移到1.5mL离心管中。之后加入150μL的纯化磁珠,用移液器吹打10次到充分混匀后,常温孵育10min;(14) Transfer the reaction product to a 1.5 mL centrifuge tube. Then add 150 μL of purified magnetic beads,
(15)瞬时离心后,将1.5mL离心管放置与磁力架上,静置5min至液体澄清,用移液器吸取并丢弃上清液;(15) After instant centrifugation, place the 1.5 mL centrifuge tube on a magnetic stand and let it stand for 5 min until the liquid becomes clear. Use a pipette to aspirate and discard the supernatant.
(16)向1.5mL离心管中加入200μL 80%乙醇,静置30s,洗涤磁珠;(16) Add 200 μL 80% ethanol to a 1.5 mL centrifuge tube and let stand for 30 seconds to wash the magnetic beads;
(17)重复上一步;(17) Repeat the previous step;
(18)尽量吸干管内液体,有少量残留在管壁时可将1.5mL离心管瞬时离心,在磁力架上分离后,用小量程的移液器将管底液体吸干;(18) Try to drain the liquid in the tube. If there is a small amount of liquid left on the tube wall, centrifuge the 1.5 mL centrifuge tube for a short time. After separation on the magnetic rack, use a small-range pipette to drain the liquid at the bottom of the tube.
(19)保持1.5mL离心管固定于磁力架上,打开1.5mL离心管管盖,室温干燥,直至磁珠表面无反光、无开裂;(19) Keep the 1.5 mL centrifuge tube fixed on the magnetic rack, open the cap of the 1.5 mL centrifuge tube, and dry it at room temperature until the surface of the magnetic beads is not reflective and has no cracks;
(20)将1.5mL离心管从磁力架上取下,加入26μL分子级水进行DNA洗脱,用移液器轻轻吹打10次至完全混匀;(20) Remove the 1.5 mL centrifuge tube from the magnetic stand, add 26 μL of molecular grade water to elute the DNA, and gently pipette 10 times to mix thoroughly;
(21)室温下孵育5min;(21) Incubate at room temperature for 5 min;
(22)瞬时离心,将1.5mL离心管置于磁力架上,静置5min至液体澄清,将25μL上清液转移到新的1.5mL离心管中;(22) Centrifuge briefly, place the 1.5 mL centrifuge tube on a magnetic rack, let stand for 5 min until the liquid becomes clear, and transfer 25 μL of the supernatant to a new 1.5 mL centrifuge tube;
(23)使用 ssDNA Assay Kit荧光定量试剂盒,按照定量试剂盒的操作说明对酶切消化纯化后产物进行定量; (23) Use ssDNA Assay Kit fluorescent quantitative kit, according to the operating instructions of the quantitative kit to quantify the product after enzyme digestion and purification;
(24)准备MGISEQ-2000高通量测序试剂套装(PE100)(MGI,1000012554),按照说明书将试剂槽1号孔和2号孔配制完成后结束。(24) Prepare the MGISEQ-2000 high-throughput sequencing reagent set (PE100) (MGI, 1000012554), and complete the preparation of
(25)之后将试剂槽中的5号孔、7号孔的试剂吸出,分子级水清洗2次后,在5号孔加入5mL甲酰胺,在7号孔加入3mL的1μM的合成二链的引物。(25) Then, the reagents in wells 5 and 7 of the reagent reservoir were aspirated, and after washing twice with molecular-grade water, 5 mL of formamide was added to well 5, and 3 mL of 1 μM primer for synthesizing the second chain was added to well 7.
合成二链的引物:5’P-GCTCACAGAACGACATGGCTACGATCCGAC-3’(SEQ ID NO:21)Primer for synthesizing the second strand: 5’P-GCTCACAGAACGACATGGCTACGATCCGAC-3’ (SEQ ID NO: 21)
(26)按照表12配制消化酶反应液,将其加入4号孔中。(26) Prepare the digestive enzyme reaction solution according to Table 12 and add it to well 4.
表12Table 12
(26)按照表13配制二链合成的酶反应液,将其加入15号孔中。(26) Prepare the enzyme reaction solution for double-strand synthesis according to Table 13 and add it to well 15.
表13Table 13
(27)加载过程,只使用试剂盒中的DNB加载缓冲液Ⅱ,吸取对应体积加入制备好的DNA多拷贝连环体产物的PCR管中,将DNA多拷贝连环体产物加载到测序芯片上后进行 测序。(27) During the loading process, only the DNB loading buffer II in the kit was used, and the corresponding volume was added to the PCR tube containing the prepared DNA multi-copy concatemer product. The DNA multi-copy concatemer product was loaded onto the sequencing chip and then sequenced.
(28)完成1链测序后,泵入5号孔的甲酰胺试剂,85℃孵育10min,再泵入清洗试剂进行清洗2次;之后泵入7号孔的合成二链的引物,65℃孵育2min,再泵入清洗试剂进行清洗2次;之后再泵入15号孔的二链合成的酶反应液试剂,65℃孵育10min,再泵入清洗试剂进行清洗2次;最后泵入4号孔的消化酶反应液,37℃孵育30min,最后泵入清洗试剂进行清洗2次,最终完成二链的合成。(28) After completing the sequencing of the first strand, pump the formamide reagent into well 5, incubate at 85°C for 10 min, and then pump the cleaning reagent into well 5 for washing twice; then pump the primer for synthesizing the second strand into well 7, incubate at 65°C for 2 min, and then pump the cleaning reagent into well 7 for washing twice; then pump the enzyme reaction solution reagent for synthesizing the second strand into well 15, incubate at 65°C for 10 min, and then pump the cleaning reagent into well 15 for washing twice; finally, pump the digestion enzyme reaction solution into well 4, incubate at 37°C for 30 min, and finally pump the cleaning reagent into well 15 for washing twice, thereby completing the synthesis of the second strand.
(29)完成2链合成后,按照试剂盒二链测序的后续流程对二链进行测序。(29) After completing the second-strand synthesis, the second strand is sequenced according to the subsequent process of the second-strand sequencing kit.
(30)测序完成后,统计获得的测序数据,数据直接来自于测序仪,包括数据量,反应数据质的Q30值。再对数据进行后续的比对分析,抽取4M的读长序列的数据进行分析,采用BWA软件比对到人参考基因组(hg19),使用samtools对测序数据的数据利用率、比对率、目标区域数据比例等进行统计。(30) After sequencing was completed, the obtained sequencing data were statistically analyzed. The data came directly from the sequencer, including the data volume and the Q30 value reflecting the data quality. The data were then subjected to subsequent alignment analysis. The data of the 4M read length sequence were extracted for analysis and aligned to the human reference genome (hg19) using BWA software. Samtools was used to perform statistics on the data utilization, alignment rate, and target region data ratio of the sequencing data.
2.实验结果2. Experimental results
(1)单链DNA测序文库经过反应之后得到DNA多拷贝连环体的浓度符合预期的浓度(表14);(1) After the single-stranded DNA sequencing library was reacted, the concentration of the DNA multi-copy concatemers was consistent with the expected concentration (Table 14);
表14单链DNA文库投入量与DNC产量Table 14 Input amount of single-stranded DNA library and DNC yield
(2)使用DNBSEQ的测序试剂与芯片进行测序后,得到350M以上的读长序列,其中1链的Q30质量值能到89%以上,二链的Q30质量值能到80%以上,达到预期目标;(2) After sequencing using DNBSEQ sequencing reagents and chips, a read length of more than 350M was obtained, of which the Q30 quality value of the first strand was more than 89%, and the Q30 quality value of the second strand was more than 80%, achieving the expected goal;
(4)测序数据的数据利用率、比对率、目标区域数据比例等进行统计,结果都符合预期,结果见表15。(4) The data utilization rate, alignment rate, and target region data ratio of the sequencing data were statistically analyzed, and the results were in line with expectations. The results are shown in Table 15.
表15Table 15
在本说明书的描述中,参考术语“一个实施方案”、“一些实施方案”、“示例”、“具体示例”、或“一些示例”等的描述意指结合该实施方案或示例描述的具体特征、结构、材料或者特点包含于本公开的至少一个实施方案或示例中。在本说明书中,对上述术语的示意性表述不必须针对的是相同的实施方案或示例。而且,描述的具体特征、结构、材料或者特点可以在任一个或多个实施方案或示例中以合适的方式结合。此外,在不相互矛盾的情况下,本领域的技术人员可以将本说明书中描述的不同实施方案或示例以及不同实施方案或示例的特征进行结合和组合。In the description of this specification, the description with reference to the terms "one embodiment", "some embodiments", "example", "specific example", or "some examples" etc. means that the specific features, structures, materials or characteristics described in conjunction with the embodiment or example are included in at least one embodiment or example of the present disclosure. In this specification, the schematic representations of the above terms do not necessarily refer to the same embodiment or example. Moreover, the specific features, structures, materials or characteristics described may be combined in any one or more embodiments or examples in a suitable manner. In addition, those skilled in the art may combine and combine the different embodiments or examples described in this specification and the features of the different embodiments or examples, without contradiction.
尽管上面已经示出和描述了本公开的实施方案,可以理解的是,上述实施方案是示例性的,不能理解为对本公开的限制,本领域的普通技术人员在本公开的范围内可以对上述实施方案进行变化、修改、替换和变型。Although the embodiments of the present disclosure have been shown and described above, it is to be understood that the above embodiments are exemplary and are not to be construed as limitations of the present disclosure, and that a person skilled in the art may change, modify, substitute and vary the above embodiments within the scope of the present disclosure.
Claims (20)
Priority Applications (2)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| CN202280102856.2A CN120435569A (en) | 2022-12-29 | 2022-12-29 | A nucleic acid amplification method and its use in sequencing |
| PCT/CN2022/143516 WO2024138550A1 (en) | 2022-12-29 | 2022-12-29 | Nucleic acid amplification method and use thereof in sequencing |
Applications Claiming Priority (1)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| PCT/CN2022/143516 WO2024138550A1 (en) | 2022-12-29 | 2022-12-29 | Nucleic acid amplification method and use thereof in sequencing |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| WO2024138550A1 true WO2024138550A1 (en) | 2024-07-04 |
Family
ID=91716051
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/CN2022/143516 Ceased WO2024138550A1 (en) | 2022-12-29 | 2022-12-29 | Nucleic acid amplification method and use thereof in sequencing |
Country Status (2)
| Country | Link |
|---|---|
| CN (1) | CN120435569A (en) |
| WO (1) | WO2024138550A1 (en) |
Citations (5)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN101213311A (en) * | 2005-04-29 | 2008-07-02 | J·克雷格·文特尔研究院 | Amplification and Cloning of Single DNA Molecules Using Rolling Circle Amplification |
| WO2018214036A1 (en) * | 2017-05-23 | 2018-11-29 | 深圳华大基因股份有限公司 | Enrichment method for genomic target region based on rolling circle amplification and application thereof |
| CN112534063A (en) * | 2018-05-22 | 2021-03-19 | 安序源有限公司 | Methods, systems, and compositions for nucleic acid sequencing |
| CN112795620A (en) * | 2019-11-13 | 2021-05-14 | 深圳华大基因股份有限公司 | Double-stranded nucleic acid cyclization method, methylation sequencing library construction method and kit |
| CN113667716A (en) * | 2021-08-27 | 2021-11-19 | 北京医院 | Construction method of sequencing library based on rolling circle amplification and its application |
-
2022
- 2022-12-29 WO PCT/CN2022/143516 patent/WO2024138550A1/en not_active Ceased
- 2022-12-29 CN CN202280102856.2A patent/CN120435569A/en active Pending
Patent Citations (5)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN101213311A (en) * | 2005-04-29 | 2008-07-02 | J·克雷格·文特尔研究院 | Amplification and Cloning of Single DNA Molecules Using Rolling Circle Amplification |
| WO2018214036A1 (en) * | 2017-05-23 | 2018-11-29 | 深圳华大基因股份有限公司 | Enrichment method for genomic target region based on rolling circle amplification and application thereof |
| CN112534063A (en) * | 2018-05-22 | 2021-03-19 | 安序源有限公司 | Methods, systems, and compositions for nucleic acid sequencing |
| CN112795620A (en) * | 2019-11-13 | 2021-05-14 | 深圳华大基因股份有限公司 | Double-stranded nucleic acid cyclization method, methylation sequencing library construction method and kit |
| CN113667716A (en) * | 2021-08-27 | 2021-11-19 | 北京医院 | Construction method of sequencing library based on rolling circle amplification and its application |
Also Published As
| Publication number | Publication date |
|---|---|
| CN120435569A (en) | 2025-08-05 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| US12454719B1 (en) | Linked ligation | |
| CN104838014B (en) | Compositions, methods, systems and kits for target nucleic acid enrichment | |
| US7851158B2 (en) | Enrichment through heteroduplexed molecules | |
| US20240384337A1 (en) | Linked target capture and ligation | |
| KR102843262B1 (en) | Single-channel sequencing method based on autoluminescence | |
| AU2015315103A1 (en) | Methods and compositions for rapid nucleic acid library preparation | |
| CN107922970A (en) | It is enriched with by the target of Single probe primer extend | |
| JP2005509433A (en) | Multiplex PCR | |
| CN104271770A (en) | Targeted DNA enrichment and sequencing | |
| US20260009021A1 (en) | Variant allele enrichment by unidirectional dual probe primer extension | |
| WO2024174850A1 (en) | Sequencing method for nucleic acid molecule | |
| US20240271126A1 (en) | Oligo-modified nucleotide analogues for nucleic acid preparation | |
| WO2024138550A1 (en) | Nucleic acid amplification method and use thereof in sequencing | |
| CN114438185B (en) | A method for double-end amplification sequencing on a chip surface | |
| CN118434882A (en) | A method for generating a labeled nucleic acid molecule population and a kit thereof | |
| CN118451196A (en) | A method for generating a labeled nucleic acid molecule population and a kit thereof | |
| KR20230124636A (en) | Compositions and methods for highly sensitive detection of target sequences in multiplex reactions | |
| WO2021168384A1 (en) | Enhanced detection of target nucleic acids by removal of dna-rna cross contamination | |
| US20260043070A1 (en) | Linked ligation | |
| RU2794177C1 (en) | Method for single-channel sequencing based on self-luminescence | |
| WO2023141604A2 (en) | Methods of molecular tagging for single-cell analysis | |
| CN117940577A (en) | Periodate compositions and methods for chemically cleaving surface-bound polynucleotides | |
| HK40055932B (en) | Single-channel sequencing method based on self-luminescence |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| 121 | Ep: the epo has been informed by wipo that ep was designated in this application |
Ref document number: 22969697 Country of ref document: EP Kind code of ref document: A1 |
|
| WWE | Wipo information: entry into national phase |
Ref document number: 202280102856.2 Country of ref document: CN |
|
| NENP | Non-entry into the national phase |
Ref country code: DE |
|
| WWP | Wipo information: published in national office |
Ref document number: 202280102856.2 Country of ref document: CN |
|
| 122 | Ep: pct application non-entry in european phase |
Ref document number: 22969697 Country of ref document: EP Kind code of ref document: A1 |