WO2024123842A1 - Systems and methods for the treatment of hemoglobinopathies - Google Patents
Systems and methods for the treatment of hemoglobinopathies Download PDFInfo
- Publication number
- WO2024123842A1 WO2024123842A1 PCT/US2023/082613 US2023082613W WO2024123842A1 WO 2024123842 A1 WO2024123842 A1 WO 2024123842A1 US 2023082613 W US2023082613 W US 2023082613W WO 2024123842 A1 WO2024123842 A1 WO 2024123842A1
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- cells
- subject
- rbc
- population
- certain embodiments
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Ceased
Links
Classifications
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P7/00—Drugs for disorders of the blood or the extracellular fluid
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K35/00—Medicinal preparations containing materials or reaction products thereof with undetermined constitution
- A61K35/12—Materials from mammals; Compositions comprising non-specified tissues or cells; Compositions comprising non-embryonic stem cells; Genetically modified cells
- A61K35/14—Blood; Artificial blood
- A61K35/18—Erythrocytes
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K35/00—Medicinal preparations containing materials or reaction products thereof with undetermined constitution
- A61K35/12—Materials from mammals; Compositions comprising non-specified tissues or cells; Compositions comprising non-embryonic stem cells; Genetically modified cells
- A61K35/28—Bone marrow; Haematopoietic stem cells; Mesenchymal stem cells of any origin, e.g. adipose-derived stem cells
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/11—DNA or RNA fragments; Modified forms thereof; Non-coding nucleic acids having a biological activity
- C12N15/113—Non-coding nucleic acids modulating the expression of genes, e.g. antisense oligonucleotides; Antisense DNA or RNA; Triplex- forming oligonucleotides; Catalytic nucleic acids, e.g. ribozymes; Nucleic acids used in co-suppression or gene silencing
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N2310/00—Structure or type of the nucleic acid
- C12N2310/10—Type of nucleic acid
- C12N2310/20—Type of nucleic acid involving clustered regularly interspaced short palindromic repeats [CRISPR]
Definitions
- Hb Hemoglobin
- RBCs red blood cells
- HbF is largely replaced by adult hemoglobin (HbA), a tetrameric protein in which the ⁇ -globin chains of HbF are replaced with beta ( ⁇ )-globin chains, through a process known as globin switching.
- HbA hemoglobin
- ⁇ -globin chains of HbF are replaced with beta ( ⁇ )-globin chains
- the average adult makes less than 1% HbF out of total hemoglobin (Thein 2009).
- the ⁇ -hemoglobin gene is located on chromosome 16, while the ⁇ -hemoglobin gene (HBB), A gamma (A ⁇ )-globin chain (HBG1, also known as gamma globin A), and G gamma (G ⁇ )-globin chain (HBG2, also known as gamma globin G) are located on chromosome 11 within the globin gene cluster (also referred to as the globin locus).
- HBB ⁇ -hemoglobin gene
- a ⁇ A gamma
- G ⁇ G gamma globin chain
- HBB2 gamma globin G
- Mutations in HBB can cause hemoglobin disorders (i.e., ⁇ -hemoglobinopathies) including sickle cell disease (SCD) and beta-thalassemia ( ⁇ -Thal).
- SCD is the most common inherited hematologic disease in the United States, affecting approximately 80,000 people (Brousseau 2010). SCD is most common in people of African ancestry, for whom the prevalence of SCD is 1 in 500. In Africa, the prevalence of SCD is 15 million (Aliyu 2008).
- SCD is also more common in people of Indian, Saudi Arabian and Mediterranean descent. In those of Hispanic-American descent, the prevalence of sickle cell disease is 1 in 1,000 (Lewis 2014).
- SCD is caused by a single homozygous mutation in the HBB gene, c.17A>T (HbS mutation).
- the sickle mutation is a point mutation (GAG>GTG) on HBB that results in substitution of valine for glutamic acid at amino acid position 6 in exon 1.
- the valine at position 6 of the ⁇ -hemoglobin chain is hydrophobic and causes a change in conformation of the ⁇ -globin protein when it is not bound to oxygen.
- HbS proteins This change of conformation causes HbS proteins to polymerize in the absence of oxygen, leading to deformation (i.e., sickling) of RBCs.
- SCD is inherited in an autosomal recessive manner, so that only patients with two HbS alleles have the disease.
- Heterozygous subjects have sickle cell trait, and may suffer from anemia and/or painful crises if they are severely dehydrated or oxygen deprived.
- Sickle shaped RBCs cause multiple symptoms, including anemia, sickle cell crises, vaso- occlusive crises, aplastic crises, and acute chest syndrome.
- Sickle shaped RBCs are less elastic than wild-type RBCs and therefore cannot pass as easily through capillary beds and cause occlusion and ischemia (i.e., vaso-occlusion).
- Vaso-occlusive crisis occurs when sickle cells obstruct blood flow in the capillary bed of an organ leading to pain, ischemia, and necrosis. These episodes typically last 5-7 days.
- the spleen plays a role in clearing dysfunctional RBCs, and is therefore typically enlarged during early childhood and subject to frequent vaso-occlusive crises. By the end of childhood, the spleen in SCD patients is often infarcted, which leads to autosplenectomy. Hemolysis is a constant feature of SCD and causes anemia.
- Sickle cells survive for 10-20 days in circulation, while healthy RBCs survive for 90-120 days.
- SCD subjects are transfused as necessary to maintain adequate hemoglobin levels. Frequent transfusions place subjects at risk for infection with HIV, Hepatitis B, and Hepatitis C. Subjects may also suffer from acute chest crises and infarcts of extremities, end organs, and the central nervous system. [0009] Subjects with SCD have decreased life expectancies. The prognosis for patients with SCD is steadily improving with careful, life-long management of crises and anemia. As of 2001, the average life expectancy of subjects with sickle cell disease was the mid-to-late 50’s.
- Thalassemias e.g., ⁇ -Thal, ⁇ -Thal, and ⁇ / ⁇ -Thal
- ⁇ -Thal is estimated to affect approximately 1 in 100,000 people worldwide. Its prevalence is higher in certain populations, including those of European descent, where its prevalence is approximately 1 in 10,000.
- ⁇ -Thal major the more severe form of the disease, is life-threatening unless treated with lifelong blood transfusions and chelation therapy. In the United States, there are approximately 3,000 subjects with ⁇ -Thal major.
- ⁇ -Thal intermedia does not require blood transfusions, but it may cause growth delay and significant systemic abnormalities, and it frequently requires lifelong chelation therapy.
- HbA makes up the majority of hemoglobin in adult RBCs, approximately 3% of adult hemoglobin is in the form of HbA2, an HbA variant in which the two ⁇ -globin chains are replaced with two delta ( ⁇ )-globin chains.
- ⁇ -Thal is associated with mutations in the ⁇ hemoglobin gene (HBD) that cause a loss of HBD expression. Co-inheritance of the HBD mutation can mask a diagnosis of ⁇ -Thal (i.e., ⁇ / ⁇ -Thal) by decreasing the level of HbA2 to the normal range (Bouva 2006).
- ⁇ / ⁇ -Thal is usually caused by deletion of the HBB and HBD sequences in both alleles. In homozygous ( ⁇ o/ ⁇ o ⁇ o/ ⁇ o) patients, HBG is expressed, leading to production of HbF alone. [0011] Like SCD, ⁇ -Thal is caused by mutations in the HBB gene.
- HBB mutations leading to ⁇ -Thal are: c.-136C>G, c.92+1G>A, c.92+6T>C, c.93-21G>A, c.118C>T, c.316-106C>G, c.25_26delAA, c.27_28insG, c.92+5G>C, c.118C>T, c.135delC, c.315+1G>A, c.- 78A>G, c.52A>T, c.59A>G, c.92+5G>C, c.124_127delTTCT, c.316-197C>T, c.-78A>G, c.52A>T, c.124_127delTTCT, c.316-197C>T, c.-138C>T, c.-79A>G, c.92+5G>C, c.75
- ⁇ -Thal intermedia results from mutations in the 5’ or 3’ untranslated region of HBB, mutations in the promoter region or polyadenylation signal of HBB, or splicing mutations within the HBB gene.
- Patient genotypes are denoted ⁇ o/ ⁇ + or ⁇ +/ ⁇ +.
- ⁇ o represents absent expression of a ⁇ - globin chain;
- ⁇ + represents a dysfunctional but present ⁇ -globin chain.
- Phenotypic expression varies among patients. Since there is some production of ⁇ -globin, ⁇ -Thal intermedia results in less precipitation of ⁇ -globin chains in the erythroid precursors and less severe anemia than ⁇ -Thal major.
- ⁇ -Thal intermedia subjects generally present between the ages of 2-6 years. They do not generally require blood transfusions. However, bone abnormalities occur due to chronic hypertrophy of the erythroid lineage to compensate for chronic anemia. Subjects may have fractures of the long bones due to osteoporosis.
- Extramedullary erythropoiesis is common and leads to enlargement of the spleen, liver, and lymph nodes. It may also cause spinal cord compression and neurologic problems. Subjects also suffer from lower extremity ulcers and are at increased risk for thrombotic events, including stroke, pulmonary embolism, and deep vein thrombosis.
- Treatment of ⁇ -Thal intermedia includes splenectomy, folic acid supplementation, hydroxyurea therapy, and radiotherapy for extramedullary masses. Chelation therapy is used in subjects who develop iron overload. [0016] Life expectancy is often diminished in ⁇ -Thal patients. Subjects with ⁇ -Thal major who do not receive transfusion therapy generally die in their second or third decade.
- HSCs hematopoietic stem cells
- SUMMARY [0018] Provided herein in certain aspects are methods of inducing expression of fetal hemoglobin (HbF) in a population of cells, e.g., CD34+ hematopoietic stem and progenitor cells (HSPCs) and/or red blood cells, in a subject.
- HbF fetal hemoglobin
- the subject may be suffering from a ⁇ - hemoglobinopathy.
- the ⁇ -hemoglobinopathy may be sickle cell disease (SCD) or ⁇ -Thal.
- the method may comprise administering to the subject a population of modified cells comprising a plurality of modified CD34+ or hematopoietic stem cells comprising an indel in an HBG promoter, thereby inducing expression of HbF in the population of cells (e.g., HSPCs or RBCs).
- HbF as a percentage of total hemoglobin (% HbF) in the subject may be about 5%, 10%, 15%, 20%, 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, or 80% HbF.
- the % HbF in the subject may be from about 10% to about 30%, from about 20% to about 40%, from about 30% to about 50%, from about 40% to about 60%, from about 50% to about 70%, from about 60% to about 80%, from about 10% to about 20%, from about 20% to about 30%, from about 30% to about 40%, from about 40% to about 50%, from about 50% to about 60%, from about 60% to about 70%, from about 70% to about 80% HbF, or a range defined by any of the two preceding values.
- a concentration of total hemoglobin in the subject may be about 10 g/dL, 11 g/dL, 12 g/dL, 13 g/dL, 14 g/dL, 15 g/dL, 16 g/dL, 17 g/dL, 18 g/dL, 19 g/dL, or 20 g/dL.
- a concentration of total hemoglobin in the subject may be from about 10.0 to about 20.0 g/dL, from about 13.6 to about 18.0 g/dL (for males), from about 12.0 to about 16.0 g/dL (for females), or a range defined by any of the two preceding values.
- a percentage of F-cells among circulating RBCs in the subject may be about 5%, 10%, 15%, 20%, 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, 100% F-cells.
- a percentage of F-cells among circulating RBCs in the subject may be from about 50% to about 100%, from about 60% to about 100%, from about 70% to about 100%, from about 80% to about 100%, from about 50% to about 60%, from about 60% to about 70%, from about 70% to about 80%, from about 80% to about 90%, from about 90% to about 100%, or a range defined by any of the two preceding values.
- a Hemoglobin F concentration in the subject may be about 1 g/dL, 2 g/dL, 3 g/dL, 4 g/dL, 5 g/dL, 6 g/dL, 7 g/dL, 8 g/dL, 9 g/dL, 10 g/dL, 11 g/dL, 12 g/dL, 13 g/dL, 14 g/dL, 15 g/dL, 16 g/dL, 17 g/dL, 18 g/dL, 19 g/dL, or 20 g/dL.
- a Hemoglobin F concentration in the subject may be from about 1g/dL to about 20 g/dL, or a range 5 164554159.2 Attorney Docket No.: 118945.8026.WO00 defined by any of the two preceding values.
- a mean corpuscular HbF (pg/RBC) in the subject may be about 1.0 pg/RBC, 2.0 pg/RBC, 3.0 pg/RBC, 4.0 pg/RBC, 5.0 pg/RBC, 6.0 pg/RBC, 7.0 pg/RBC, 8.0 pg/RBC, 9.0 pg/RBC, 10.0 pg/RBC, 11.0 pg/RBC, 12.0 pg/RBC, 13.0 pg/RBC, 14.0 pg/RBC 15.0 pg/RBC, 16.0 pg/RBC, 17.0 pg/RBC, 18.0 pg/RBC, 19.0 pg/RBC, 20.0 pg/RBC, 25.0 pg/RBC, or 30.0 pg/RBC.
- a mean corpuscular HbF (pg/RBC) in the subject may be from about 1.0 pg/RBC to about 5.00 pg/RBC, from about 5.0 pg/RBC to about 10.00 pg/RBC, from about 10.0 pg/RBC to about 15.00 pg/RBC, from about 15.0 pg/RBC to about 20.00 pg/RBC, from about 20.0 pg/RBC to about 25.00 pg/RBC, from about 25.0 pg/RBC to about 30.00 pg/RBC, or a range defined by any of the two preceding values.
- a mean corpuscular HbF (pg/RBC) in the subject may be ⁇ 10.0 pg/RBC.
- a mean proportion of HbF as a percentage of total hemoglobin (Hb) in the subject is about 5%, 10%, 15%, 20%, 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%.
- a mean proportion of HbS as a percentage of total hemoglobin (Hb) in the subject is about 5%, 10%, 15%, 20%, 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%.
- a mean proportion of HbA as a percentage of total hemoglobin (Hb) in the subject is about 1%, 2%, 3%, 4%, 5%, 10%, 15%, 20%, 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%.
- a mean proportion of HbA2 as a percentage of total hemoglobin (Hb) in the subject is about 1%, 2%, 3%, 4%, 5%.
- a lactate dehydrogenase (U/L) in the subject is about 110 U/L, 120 U/L, 130 U/L, 140 U/L, 150 U/L, 160 U/L, 170 U/L, 180 U/L, 190 U/L, 200 U/L, 210 U/L, 220 U/L, or 230 U/L.
- a lactate dehydrogenase (U/L) in the subject is from about 110 U/L to about 230 U/L, or a range defined by any of the two preceding values.
- an indirect bilirubin ( ⁇ mol/L) in the subject is about 0.0 ⁇ mol/L, 1 ⁇ mol/L, 2 ⁇ mol/L, 3 ⁇ mol/L, 4 ⁇ mol/L, 5 ⁇ mol/L, 6 ⁇ mol/L, 7 ⁇ mol/L, 8 ⁇ mol/L, 9 ⁇ mol/L, 10 ⁇ mol/L, 11 ⁇ mol/L, 12 ⁇ mol/L, 13 ⁇ mol/L, 14 ⁇ mol/L, 15 ⁇ mol/L, 16 ⁇ mol/L, or 17 ⁇ mol/L.
- an indirect bilirubin ( ⁇ mol/L) in the subject is from about 0.0 ⁇ mol/L to about 16.6 ⁇ mol/L, or a range defined by the two preceding values.
- a haptoglobin (g/L) in the subject is about 0.3 g/L, 0.4 g/L, 0.5 g/L, 0.6 g/L, 0.7 g/L, 0.8 g/L, 0.9 g/L, 1.0 g/L, 1.1 g/L, 1.2 g/L, 1.3 g/L, 1.4 g/L, 1.5 g/L, 1.6 g/L, 1.7 g/L, 1.8 g/L, 1.9 g/L, or 2.0 g/L.
- a haptoglobin (g/L) in the subject is from about 0.3 g/L to about 2.0 g/L or a range defined by the two preceding values.
- a reticulocyte count (%) in the subject is about 0.3%, 0.4%, 0.5%, 0.6%, 0.7%, 0.8%, 0.9%, 1.0%, 1.1%, 1.2%, 1.3%, 1.4%, 1.5%, 1.6%, 1.7%, 1.8%, 1.9%, 2.0%, 2.1%, 2.2%, or 2.3%.
- a reticulocyte count (%) in the subject is from about 0.3 % to about 2.3% or a range defined by the two preceding values.
- normalization of total hemoglobin in the subject occurs by at least 1 month, 1.5 months, 2 months, 3 months, 4 months, 5 months after administering the population of modified cells 6 164554159.2 Attorney Docket No.: 118945.8026.WO00 to the subject.
- the subject may undergo myeloablative conditioning with busulfan prior to administering the population of modified cells.
- administering the population of modified cells may comprise a single infusion of the modified population of cells.
- the population of modified cells may be about ⁇ 1 x 10 6 cells/kg, ⁇ 2 x 10 6 cells/kg, ⁇ 3 x 10 6 cells/kg, ⁇ 4 x 10 6 cells/kg, ⁇ 5 x 10 6 cells/kg, ⁇ 6 x 10 6 cells/kg, ⁇ 7 x 10 6 cells/kg, ⁇ 8 x 10 6 cells/kg, ⁇ 9 x 10 6 cells/kg, ⁇ 10 x 10 6 cells/kg, ⁇ 11 x 10 6 cells/kg, ⁇ 12 x 10 6 cells/kg, ⁇ 13 x 10 6 cells/kg, ⁇ 14 x 10 6 cells/kg, ⁇ 15 x 10 6 cells/kg, ⁇ 16 x 10 6 cells/kg, ⁇ 17 x 10 6 cells/kg, ⁇ 18 x 10 6 cells/kg, ⁇ 19 x 10 6 cells/kg, ⁇ 20 x 10 6 cells/kg, ⁇ 21 x 10 6 cells/kg, ⁇ 22 x 10 6 cells/kg,
- the % HbF may be determined 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30 months after administering the population of modified cells to the subject.
- the concentration of total hemoglobin may be determined 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30 months after administering the population of modified cells to the subject.
- the Hemoglobin F concentration may be determined 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30 months after administering the population of modified cells to the subject.
- the percentage of F-cells may be determined 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30 months after administering the population of modified cells to the subject.
- the mean corpuscular HbF (pg/RBC) may be determined 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30 months after administering the population of modified cells to the subject.
- the mean proportion of HbF as a percentage of total hemoglobin is determined 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30 months after administering the population of modified cells to the subject.
- the mean proportion of HbS as a percentage of total hemoglobin is determined 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30 months after administering the population of modified cells to the subject.
- the mean proportion of HbA as a percentage of total hemoglobin is determined 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30 months after administering the population of modified cells to the subject.
- the mean proportion of HbA2 as a percentage of total hemoglobin is determined 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30 months after administering the population of modified cells to the subject.
- the lactate dehydrogenase U/L is determined 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30 months after administering the population of modified cells to the subject.
- the indirect bilirubin is determined 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30 months after administering the population of modified cells to the subject.
- the haptoglobin is determined 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30 months after administering the population of modified cells to the subject.
- the reticulocyte count (%) is determined 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30 months after administering the population of modified cells to the subject.
- an RNP complex comprising a guide RNA (gRNA) and a Cpf1 protein may be delivered to a population of unmodified cells comprising a plurality of unmodified CD34+ or hematopoietic stem cells from the subject to generate the population of modified cells.
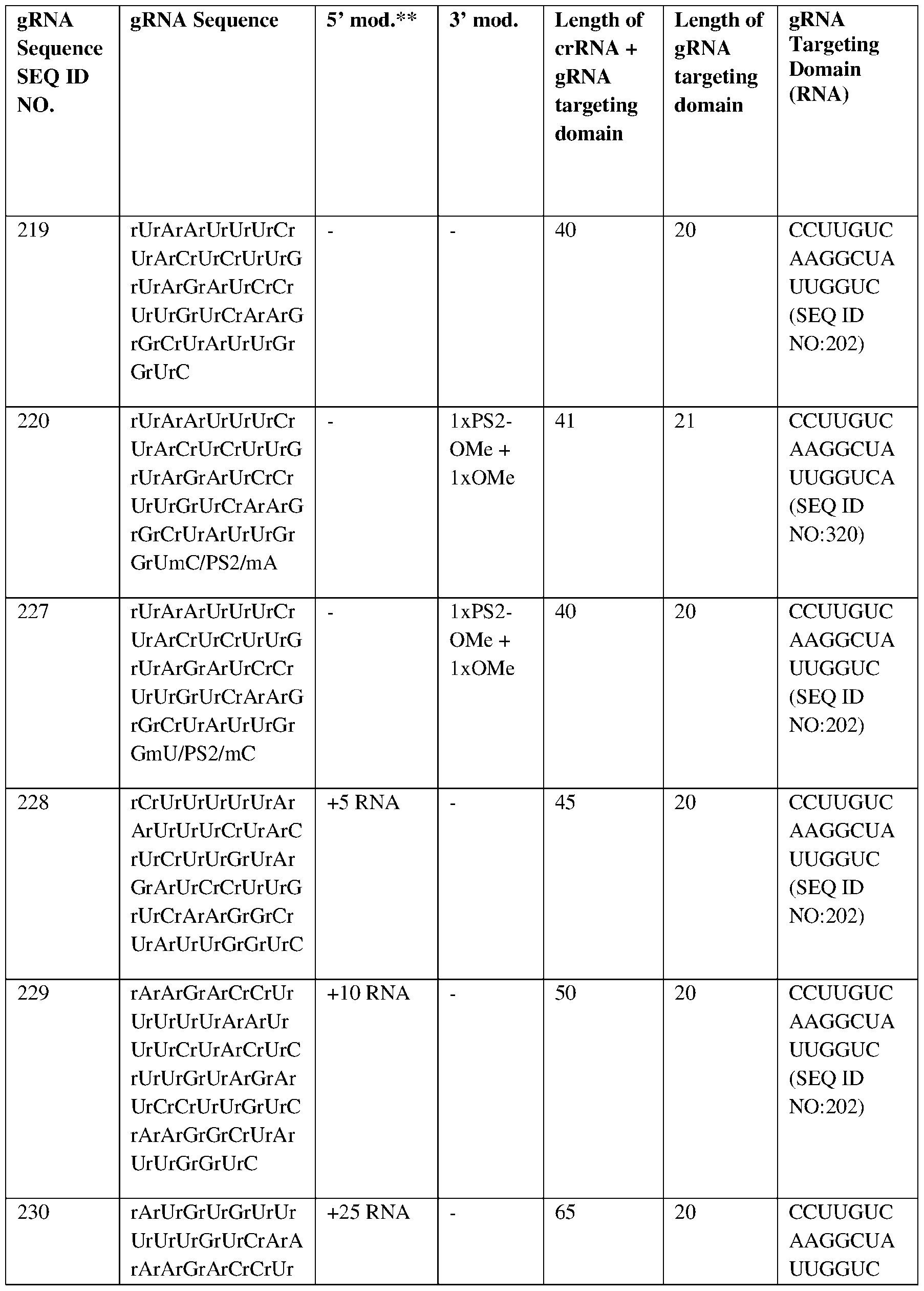
- the gRNA may comprise a targeting domain comprising SEQ ID NO:320.
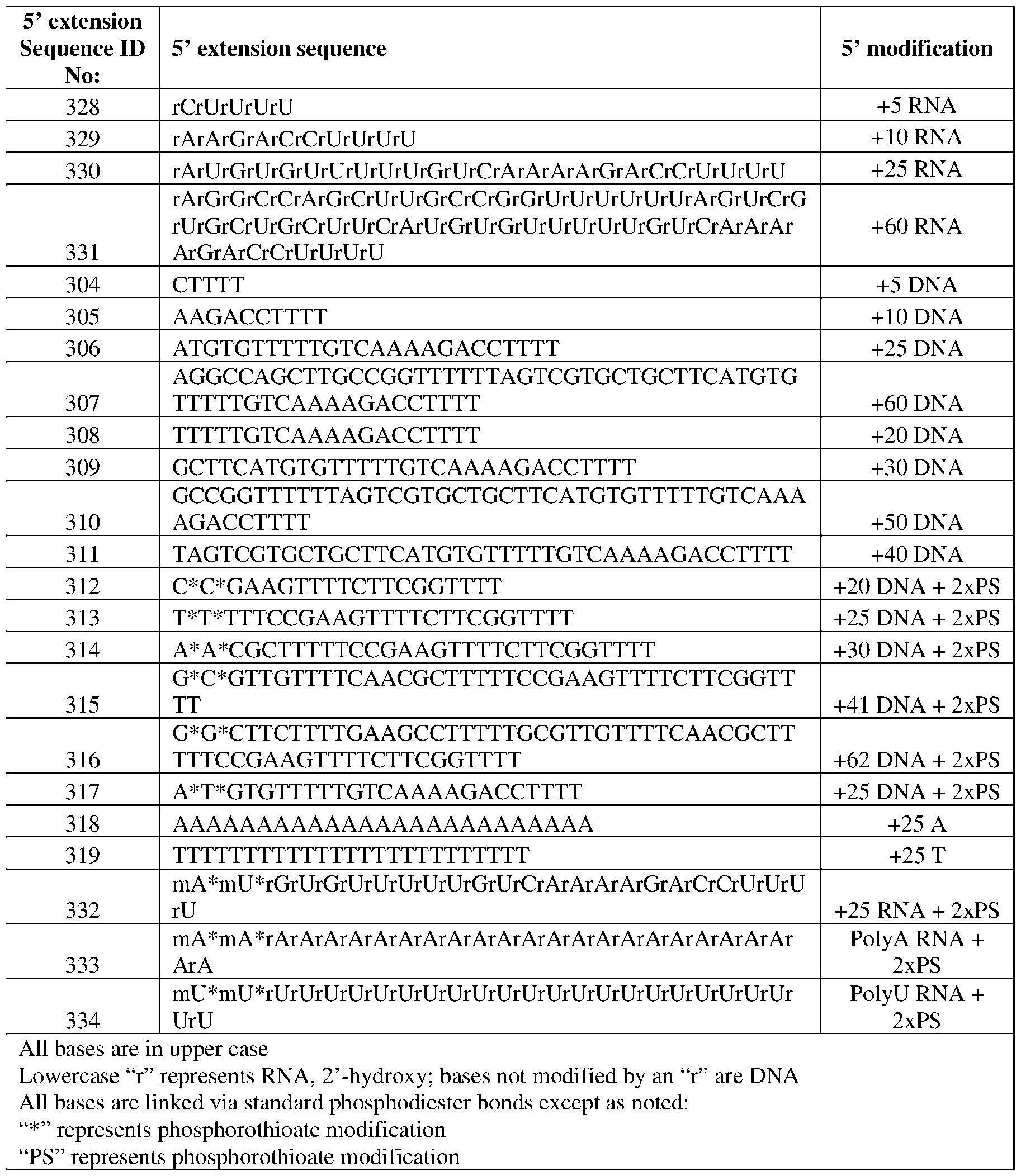
- the gRNA may comprise a 5’ end and a 3’ end, a DNA extension at the 5’ end.
- the gRNA may comprise a 2’-O-methyl, phosphorothioate modification, or both at the 3’ end.
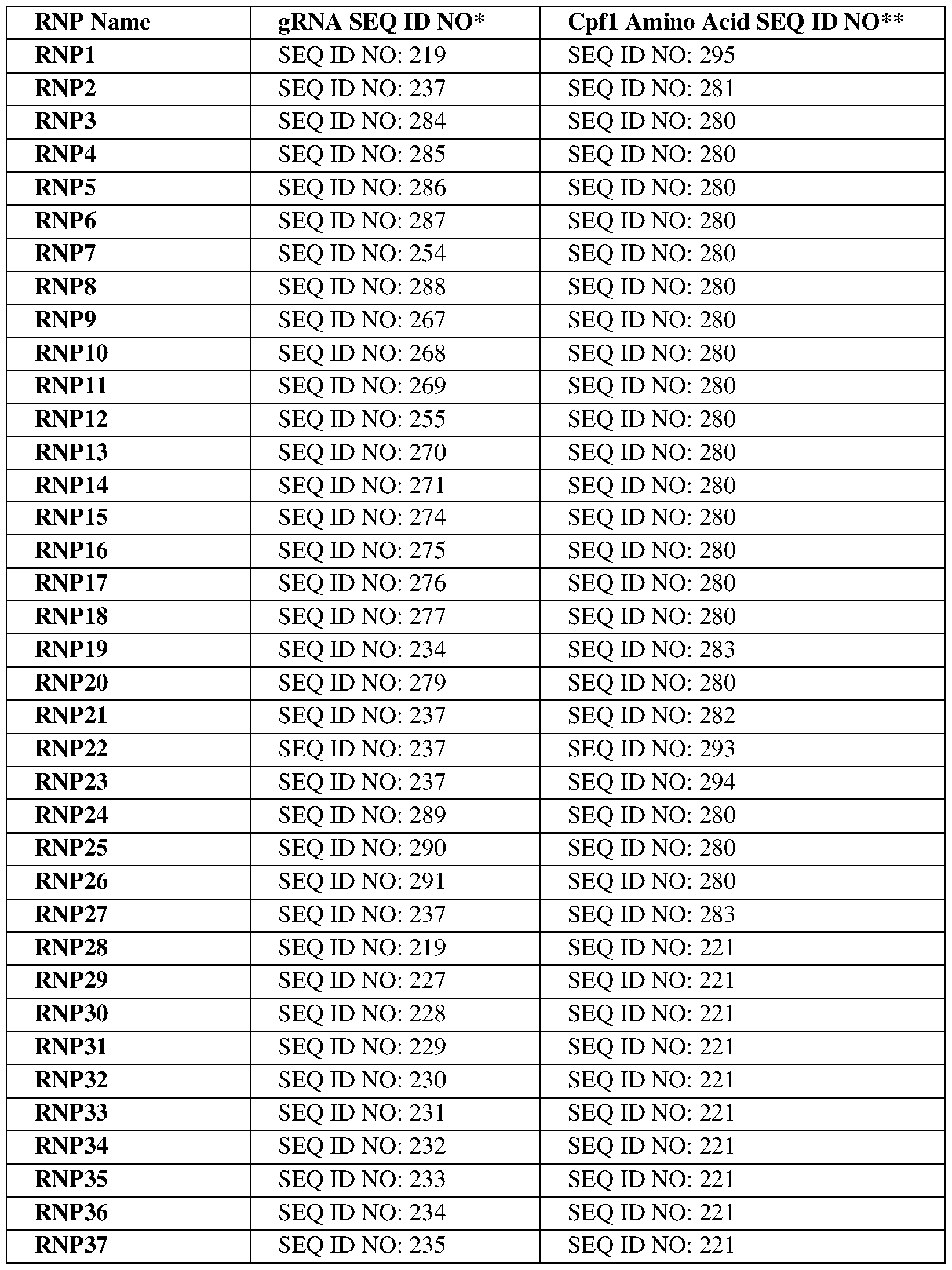
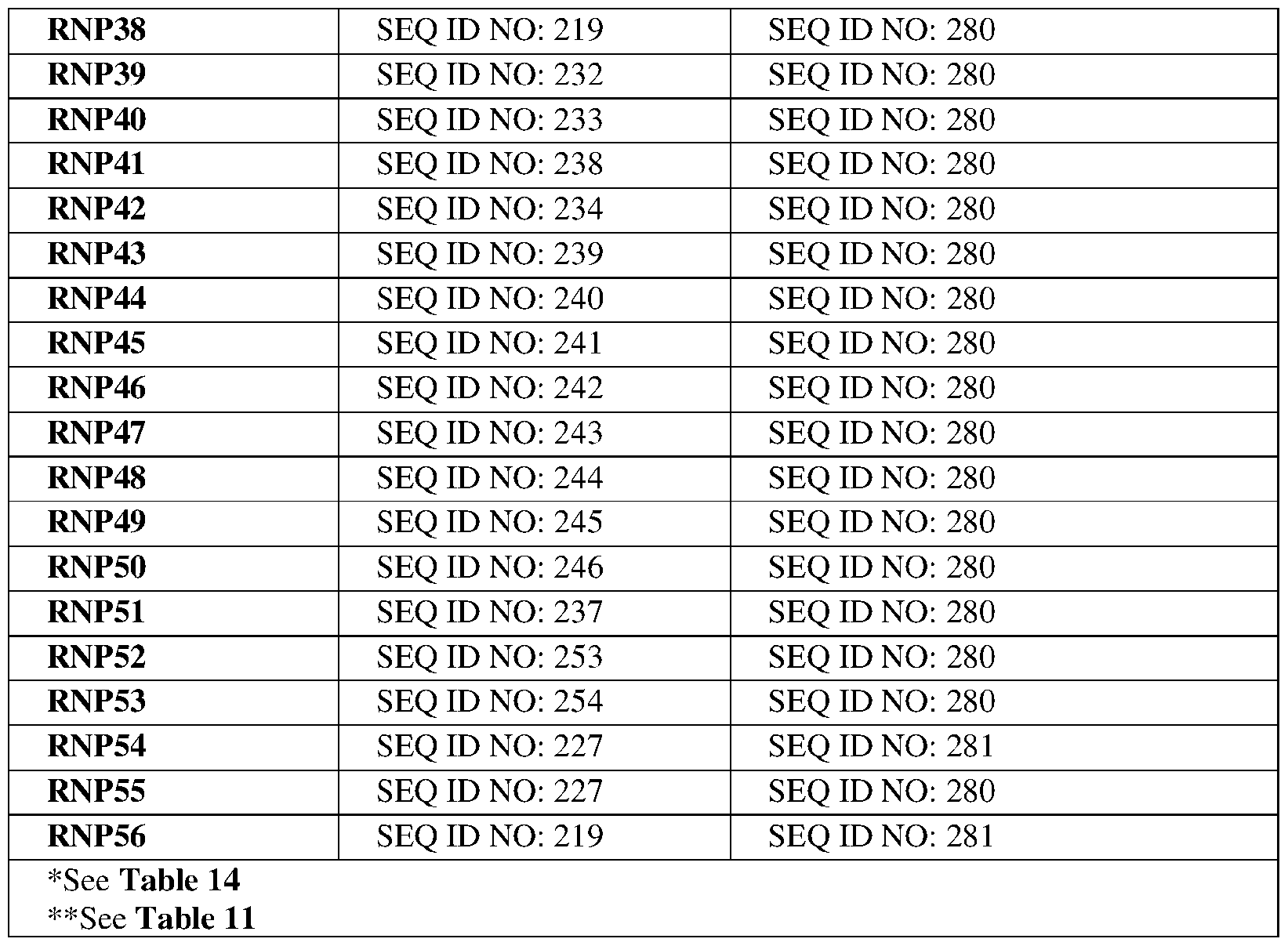
- the DNA extension may comprise a sequence selected from the group consisting of SEQ ID NOs:304-319. In certain embodiments, the DNA extension may comprise a sequence selected from the group consisting of SEQ ID NOs:304-319. In certain embodiments, the gRNA may comprise SEQ ID NO:237. In certain embodiments, the Cpf1 protein may comprise a sequence selected from the group consisting of SEQ ID NO:200, 201, 205-215, 221, 222-226, 280-283, 293- 295. In certain embodiments, the Cpf1 protein may comprise SEQ ID NO:283.
- the Cpf1 protein may be encoded by a sequence comprising a sequence selected from the group consisting of SEQ ID NOs:216-218, 296-303. In certain embodiments, the Cpf1 protein may be encoded by a sequence comprising SEQ ID NO:300.
- the indel in the HBG gene promoter may be in a CCAAT box target region.
- the subject has a single homozygous mutation in the HBB gene, c.17A>T (HbS mutation). In certain embodiments, the subject may be suffering from severe sickle cell disease.
- kits for producing a modified population of peripheral blood nucleated cells in a subject comprising: administering to the subject a population of modified cells comprising a plurality of modified CD34+ or hematopoietic stem cells comprising an indel in an HBG promoter, thereby producing the modified population of peripheral blood nucleated cells, wherein the modified population of peripheral blood nucleated cells as a percentage of the total population of peripheral blood nucleated cells is at least 50%, 55%, 60%, 65%, 8 164554159.2 Attorney Docket No.: 118945.8026.WO00 70%, 75%, 80%, 85%, 90% or 95%.
- the subject may undergo myeloablative conditioning with busulfan prior to administering the population of modified cells.
- methods of generating a population of F-cells in a subject comprising: administering to the subject a population of modified cells comprising a plurality of modified CD34+ or hematopoietic stem cells comprising an indel in an HBG promoter, wherein the percentage of F-cells among circulating red blood cells six months after said administering is at least 90%.
- mean corpuscular HbF per F-cell in the population of F-cells is at least 18 picograms (pg).
- the subject may exhibit no vaso-occlusive events (VOEs) within six months after said administering. In certain embodiments, the subject may exhibit no VOEs within one year after said administering.
- VOEs vaso-occlusive events
- Provided herein in certain aspects are methods of treating a ⁇ -hemoglobinopathy in a subject in need thereof.
- the ⁇ -hemoglobinopathy may be sickle cell disease (SCD) or ⁇ -Thal.
- the method may comprise administering to the subject a population of modified cells comprising a plurality of modified CD34+ or hematopoietic stem cells comprising an indel in an HBG promoter, thereby inducing expression of HbF in the population of cells (e.g., HSPCs or RBCs).
- HbF as a percentage of total hemoglobin (% HbF) in the subject may be about 5%, 10%, 15%, 20%, 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, or 80% HbF.
- the % HbF in the subject may be from about 10% to about 30%, from about 20% to about 40%, from about 30% to about 50%, from about 40% to about 60%, from about 50% to about 70%, from about 60% to about 80%, from about 10% to about 20%, from about 20% to about 30%, from about 30% to about 40%, from about 40% to about 50%, from about 50% to about 60%, from about 60% to about 70%, from about 70% to about 80% HbF, or a range defined by any of the two preceding values.
- a concentration of total hemoglobin in the subject may be about 10 g/dL, 11 g/dL, 12 g/dL, 13 g/dL, 14 g/dL, 15 g/dL, 16 g/dL, 17 g/dL, 18 g/dL, 19 g/dL, or 20 g/dL.
- a concentration of total hemoglobin in the subject may be from about 10.0 to about 20.0 g/dL, from about 13.6 to about 18.0 g/dL (for males), from about 12.0 to about 16.0 g/dL (for females), or a range defined by any of the two preceding values.
- a percentage of F-cells among circulating RBCs in the subject may be about 5%, 10%, 15%, 20%, 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, 100% F-cells.
- a percentage of F-cells among circulating RBCs in the subject may be from about 50% to about 100%, from about 60% to about 100%, from about 70% to about 100%, from about 80% to about 100%, from about 50% to about 60%, from about 60% to about 70%, from about 70% to about 80%, from 9 164554159.2 Attorney Docket No.: 118945.8026.WO00 about 80% to about 90%, from about 90% to about 100%, or a range defined by any of the two preceding values.
- a Hemoglobin F concentration in the subject may be about 1 g/dL, 2 g/dL, 3 g/dL, 4 g/dL, 5 g/dL, 6 g/dL, 7 g/dL, 8 g/dL, 9 g/dL, 10 g/dL, 11 g/dL, 12 g/dL, 13 g/dL, 14 g/dL, 15 g/dL, 16 g/dL, 17 g/dL, 18 g/dL, 19 g/dL, or 20 g/dL.
- a Hemoglobin F concentration in the subject may be from about 1g/dL to about 20 g/dL, or a range defined by any of the two preceding values.
- a mean corpuscular HbF (pg/RBC) in the subject may be about 1.0 pg/RBC, 2.0 pg/RBC, 3.0 pg/RBC, 4.0 pg/RBC, 5.0 pg/RBC, 6.0 pg/RBC, 7.0 pg/RBC, 8.0 pg/RBC, 9.0 pg/RBC, 10.0 pg/RBC, 11.0 pg/RBC, 12.0 pg/RBC, 13.0 pg/RBC, 14.0 pg/RBC 15.0 pg/RBC, 16.0 pg/RBC, 17.0 pg/RBC, 18.0 pg/RBC, 19.0 pg/RBC, 20.0 pg/RBC, 2
- a mean corpuscular HbF (pg/RBC) in the subject may be from about 1.0 pg/RBC to about 5.00 pg/RBC, from about 5.0 pg/RBC to about 10.00 pg/RBC, from about 10.0 pg/RBC to about 15.00 pg/RBC, from about 15.0 pg/RBC to about 20.00 pg/RBC, from about 20.0 pg/RBC to about 25.00 pg/RBC, from about 25.0 pg/RBC to about 30.00 pg/RBC, or a range defined by any of the two preceding values.
- a mean corpuscular HbF (pg/RBC) in the subject may be ⁇ 10.0 pg/RBC.
- a mean proportion of HbF as a percentage of total hemoglobin (Hb) in the subject is about 5%, 10%, 15%, 20%, 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%.
- a mean proportion of HbS as a percentage of total hemoglobin (Hb) in the subject is about 5%, 10%, 15%, 20%, 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%.
- a mean proportion of HbA as a percentage of total hemoglobin (Hb) in the subject is about 1%, 2%, 3%, 4%, 5%, 10%, 15%, 20%, 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%.
- a mean proportion of HbA2 as a percentage of total hemoglobin (Hb) in the subject is about 1%, 2%, 3%, 4%, 5%.
- a lactate dehydrogenase (U/L) in the subject is about 110 U/L, 120 U/L, 130 U/L, 140 U/L, 150 U/L, 160 U/L, 170 U/L, 180 U/L, 190 U/L, 200 U/L, 210 U/L, 220 U/L, or 230 U/L.
- a lactate dehydrogenase (U/L) in the subject is from about 110 U/L to about 230 U/L, or a range defined by any of the two preceding values.
- an indirect bilirubin ( ⁇ mol/L) in the subject is about 0.0 ⁇ mol/L, 1 ⁇ mol/L, 2 ⁇ mol/L, 3 ⁇ mol/L, 4 ⁇ mol/L, 5 ⁇ mol/L, 6 ⁇ mol/L, 7 ⁇ mol/L, 8 ⁇ mol/L, 9 ⁇ mol/L, 10 ⁇ mol/L, 11 ⁇ mol/L, 12 ⁇ mol/L, 13 ⁇ mol/L, 14 ⁇ mol/L, 15 ⁇ mol/L, 16 ⁇ mol/L, or 17 ⁇ mol/L.
- an indirect bilirubin ( ⁇ mol/L) in the subject is from about 0.0 ⁇ mol/L to about 16.6 ⁇ mol/L, or a range defined by the two preceding values.
- a haptoglobin (g/L) in the subject is about 0.3 g/L, 0.4 g/L, 0.5 g/L, 0.6 g/L, 0.7 g/L, 0.8 g/L, 0.9 g/L, 1.0 g/L, 1.1 g/L, 1.2 g/L, 1.3 g/L, 1.4 g/L, 1.5 g/L, 1.6 g/L, 1.7 g/L, 1.8 g/L, 1.9 g/L, or 2.0 g/L.
- a haptoglobin (g/L) in the subject is from about 0.3 g/L to about 2.0 g/L or a range defined by the two preceding values.
- a reticulocyte count (%) in the subject 10 164554159.2 Attorney Docket No.: 118945.8026.WO00 is about 0.3%, 0.4%, 0.5%, 0.6%, 0.7%, 0.8%, 0.9%, 1.0%, 1.1%, 1.2%, 1.3%, 1.4%, 1.5%, 1.6%, 1.7%, 1.8%, 1.9%, 2.0%, 2.1%, 2.2%, or 2.3%.
- a reticulocyte count (%) in the subject is from about 0.3 % to about 2.3% or a range defined by the two preceding values.
- normalization of total hemoglobin in the subject occurs by at least 1 month, 1.5 months, 2 months, 3 months, 4 months, 5 months after administering the population of modified cells to the subject.
- the subject may undergo myeloablative conditioning with busulfan prior to administering the population of modified cells.
- administering the population of modified cells may comprise a single infusion of the modified population of cells.
- the population of modified cells may be about ⁇ 1 x 10 6 cells/kg, ⁇ 2 x 10 6 cells/kg, ⁇ 3 x 10 6 cells/kg, ⁇ 4 x 10 6 cells/kg, ⁇ 5 x 10 6 cells/kg, ⁇ 6 x 10 6 cells/kg, ⁇ 7 x 10 6 cells/kg, ⁇ 8 x 10 6 cells/kg, ⁇ 9 x 10 6 cells/kg, ⁇ 10 x 10 6 cells/kg, ⁇ 11 x 10 6 cells/kg, ⁇ 12 x 10 6 cells/kg, ⁇ 13 x 10 6 cells/kg, ⁇ 14 x 10 6 cells/kg, ⁇ 15 x 10 6 cells/kg, ⁇ 16 x 10 6 cells/kg, ⁇ 17 x 10 6 cells/kg, ⁇ 18 x 10 6 cells/kg, ⁇ 19 x 10 6 cells/kg, ⁇ 20 x 10 6 cells/kg, ⁇ 21 x 10 6 cells/kg, ⁇ 22 x 10 6 cells/kg,
- the % HbF may be determined 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30 months after administering the population of modified cells to the subject.
- the concentration of total hemoglobin may be determined 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30 months after administering the population of modified cells to the subject.
- the Hemoglobin F concentration may be determined 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30 months after administering the population of modified cells to the subject.
- the percentage of F-cells may be determined 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30 months after administering the population of modified cells to the subject.
- the mean corpuscular HbF (pg/RBC) may be determined 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30 months after administering the population of modified cells to the subject.
- the mean proportion of HbF as a percentage of total hemoglobin is determined 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30 months after administering the population of modified cells to the subject.
- the mean proportion of HbS as a percentage of total hemoglobin is determined 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30 months after administering the population of modified cells to the subject.
- the mean proportion of HbA as a 11 164554159.2 Attorney Docket No.: 118945.8026.WO00 percentage of total hemoglobin is determined 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30 months after administering the population of modified cells to the subject.
- the mean proportion of HbA2 as a percentage of total hemoglobin is determined 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30 months after administering the population of modified cells to the subject.
- the lactate dehydrogenase (U/L) is determined 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30 months after administering the population of modified cells to the subject.
- the indirect bilirubin is determined 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30 months after administering the population of modified cells to the subject.
- the haptoglobin is determined 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30 months after administering the population of modified cells to the subject.
- the reticulocyte count (%) is determined 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30 months after administering the population of modified cells to the subject.
- an RNP complex comprising a guide RNA (gRNA) and a Cpf1 protein may be delivered to a population of unmodified cells comprising a plurality of unmodified CD34+ or hematopoietic stem cells from the subject to generate the population of modified cells.
- the gRNA may comprise a targeting domain comprising SEQ ID NO:320.
- the gRNA may comprise a 5’ end and a 3’ end, a DNA extension at the 5’ end.
- the gRNA may comprise a 2’-O-methyl, phosphorothioate modification, or both at the 3’ end.
- the DNA extension may comprise a sequence selected from the group consisting of SEQ ID NOs:304-319. In certain embodiments, the DNA extension may comprise a sequence selected from the group consisting of SEQ ID NOs:304-319. In certain embodiments, the gRNA may comprise SEQ ID NO:237. In certain embodiments, the Cpf1 protein may comprise a sequence selected from the group consisting of SEQ ID NO:200, 201, 205-215, 221, 222-226, 280-283, 293- 295. In certain embodiments, the Cpf1 protein may comprise SEQ ID NO:283.
- the Cpf1 protein may be encoded by a sequence comprising a sequence selected from the group consisting of SEQ ID NOs:216-218, 296-303. In certain embodiments, the Cpf1 protein may be encoded by a sequence comprising SEQ ID NO:300.
- the indel in the HBG gene promoter may be in a CCAAT box target region.
- the subject has a single homozygous mutation in the HBB gene, c.17A>T (HbS mutation). In certain embodiments, the subject may be suffering from severe sickle cell disease. [0022] This listing is intended to be exemplary and illustrative rather than comprehensive and limiting.
- Fig.1 depicts, in schematic form, HBG1 and HBG2 gene(s) in the context of the ⁇ -globin gene cluster on human chromosome 11. Each gene in the ⁇ -globin gene cluster is transcriptionally regulated by a proximal promoter.
- a ⁇ and/or G ⁇ expression is activated by engagement between the proximal promoter with the distal strong erythroid-specific enhancer, the locus control region (LCR).
- LCR locus control region
- Long- range transactivation by the LCR is thought to be mediated by alteration of chromatin configuration/confirmation.
- the LCR is marked by 4 erythroid specific Dnase I hypersensitive sites (HS1-4) and 2 distal enhancer elements (5’ HS and 3’ HS1).
- HS1-4 erythroid specific Dnase I hypersensitive sites
- 5’ HS and 3’ HS1 distal enhancer elements
- Figs.2A-2B depict HBG1 and HBG2 genes, coding sequences (CDS) and small deletions and point mutations in and upstream of the HBG1 and HBG2 proximal promoters that have been identified in patients and associated with elevation of fetal hemoglobin (HbF). Core elements within the proximal promoters (CAAT box, 13 nt sequence) that have been deleted in some patients with hereditary persistence of fetal hemoglobin (HPFH). The ‘target sequence’ region of each locus, which has been screened for gRNA binding target sites, is also identified. [0027] Fig.3 depicts editing in the HBG distal CCAAT box region by RNP27 (Table 12).
- RNP27 comprises a gRNA comprising the sequence set forth in SEQ ID NO:237 complexed with a Cpf1 protein comprising the sequence set forth in SEQ ID NO:283.
- RNP27 targets HBG1 and HBG2 promoters on chromosome 11, which are 4.9 kb apart.
- the distal CCAAT box is highlighted with a box.
- the protospacer adjacent motif (PAM) is bolded in black.
- the target sequence of the RNP27 gRNA is underlined.
- the RNP27 cleavage site and the resulting 5’ overhangs are indicated with dotted arrows.
- the point mutations associated with HPFH are bolded.
- Figs.4A-E depict laboratory parameters for Subjects 1 or 2.
- Fig.4A depicts the total hemoglobin and hemoglobin fractionation data for Subject 1 at baseline and at 1, 1.5, 2, 3, 4, and 5 months after autologous RNP27 edited CD34+ cell infusion.
- HbF The lower portion of the bar (“HbF,” light grey) represents the mean proportion of HbF as a percentage of total hemoglobin (i.e., 5.0% (baseline), 18.7% (1M), 28.1% (1.5 M), 33.5% (2M), 36.7% (3M), 42.6% (4M), 45.4% (5M));
- the middle portion of the bar (“HbS,” dark grey) represents the mean proportion of HbS as a percentage of total Hb (i.e., 89.1% (baseline), 18.2% (1M), 26.3% (1.5 M), 32.2% (2M), 37.5% (3M), 49.9% (4M), 51.8% (5M));
- Hb g/dL
- Labels indicate mean proportion of HbS and HbF as a percentage of total Hb.
- Mean total Hb concentrations are shown directly above bars.
- Hb hemoglobin
- HbF fetal hemoglobin
- HbS sickle hemoglobin
- RBC red blood cell.
- the X axis shows months after autologous RNP27 edited CD34+ cell infusion.
- the dotted lines show the 13.6-18.0 g/dL normal range for men (Central laboratory reference range).
- Fig.4B depicts the total hemoglobin levels over time for Subject 1 at baseline and at 1, 1.5, 2, 3, 4, 5, 6, and 8 months after autologous RNP27 edited CD34+ cell infusion.
- the first portion of the bar (“HbF”) represents the mean proportion of HbF as a percentage of total hemoglobin (i.e., 5.0% (baseline), 18.7% (1M), 28.1% (1.5 M), 33.5% (2M), 36.7% (3M), 42.6% (4M), 45.4% (5M), 44.3% (6M), 44.1% (8M));
- the second portion of the bar (“HbS”) represents the mean proportion of HbS as a percentage of total Hb (i.e., 89.1% (baseline), 18.2% (1M), 26.3% (1.5 M), 32.2% (2M), 37.5% (3M), 49.9% (4M), 51.8% (5M), 52.1% (6M), 54.4% (8M));
- the third portion of the bar (“HbA”) represents HbA (i.e., 6.3% (baseline), 61.7% (1M), 44.7% (1.5 M), 33.5% (2M), 23.4% (3M), 4.2% (4M), 1.9% (5M), 2.0% (6M), 1.9% (8
- Hb g/dL
- Labels indicate mean proportion of HbS and HbF as a percentage of total Hb.
- Mean total Hb concentrations are shown directly above bars.
- Hb hemoglobin
- HbF fetal hemoglobin
- HbS sickle hemoglobin.
- the X axis shows months after autologous RNP27 edited CD34+ cell infusion.
- the region identified between dotted lines indicates the 13.6-18.0 g/dL normal range for men (Central laboratory reference range).
- Fig.4C depicts the percentage of F-cells for Subject 1 at baseline and at 1, 1.5, 2, 3, 4, 5, 6, and 8 months after autologous RNP27 edited CD34+ cell infusion.
- the X axis shows months after autologous RNP27 edited CD34+ cell infusion. Higher pancellularity indicates more red blood cells express HbF for potential clinical benefit.
- Fig.4D depicts the percentage of 14 164554159.2 Attorney Docket No.: 118945.8026.WO00 mean corpuscular HbF (pg) concentration for Subject 1 at baseline and at 1, 1.5, 2, 3, 4, and 5 months after autologous RNP27 edited CD34+ cell infusion.
- Mean Corpuscular HbF: (MCH (pg)*HbF(%))/100% HbF(pg) per RBC. 10 pg/RBC is the threshold for protection from sickling and is shown by a dotted line (Steinberg 2014).
- the X axis shows months after autologous RNP27 edited CD34+ cell infusion.
- Fig.4E depicts the mean corpuscular HbF (pg) per F-cell for Subject 1 at baseline and at 1, 1.5, 2, 3, 4, 5, 6, and 8 months after autologous RNP27 edited CD34+ cell infusion.
- Mean corpuscular HbF (pg) is shown directly above bars.
- Mean Corpuscular HbF: (MCH (pg)*HbF(%))/100% HbF(pg) per RBC.
- 10 pg/RBC is the threshold for protection from sickling and is shown by a dotted line (Steinberg 2014).
- the X axis shows months after autologous RNP27 edited CD34+ cell infusion.
- Fig.4F depicts hemoglobin fractionation (%) data for Subject 1 at various study days before and after autologous RNP27 edited CD34+ cell infusion.
- the lower portion of the bar (“HbF”) represents the mean proportion of HbF as a percentage of total hemoglobin (i.e., 3.5% (Study Day (“SD”) -147), 5.0% (SD -106), 4.6% (SD -85), 1.4% (SD -14), 1.3% (SD 1), 18.7% (SD 31), 28.1% (SD 45), 33.5% (SD 59), 36.7% (SD 80), 42.6% (SD 122), 45.5% (SD 150), 44.3 (SD 191);
- the middle portion of the bar (“Other”) represents other hemoglobin as a percentage of total hemoglobin (i.e., 24% (SD -147), 5.9% (SD -106), 9.6% (SD -85), 54.9% (SD -14), 73.2% (SD 1), 63.1% (SD 31), 45.6% (SD
- Labels indicate mean proportion of HbS and HbF as a percentage of total Hb.
- the percentage of F-Cells, total hemoglobin (g/dL), and MCH-F/F-cell (pg) are shown directly above the bar graph.
- Hb hemoglobin
- HbF fetal hemoglobin
- HbS sickle hemoglobin
- MCH mean corpuscular HbF
- RBC red blood cell.
- the X axis shows Study Days (“SD”) -147, -106, -85, -14, 1, 31, 45, 59, 80, 122, 150, and 191.
- Study Day 1 represents the day on which autologous RNP27 edited CD34+ cell infusion was administered.
- Fig.4G depicts the total hemoglobin levels over time for Subject 2 at baseline and at 1, 1.5, 2, 3, 4, and 5 months after autologous RNP27 edited CD34+ cell infusion.
- the first (lower) portion of the bar (“HbF”) represents the mean proportion of HbF as a percentage of total hemoglobin (i.e., 10.8% (baseline), 8.2% (1M), 17.0% (1.5 M), 25.8% (2M), 38.6% (3M), 46.2% (4M), 51.2% (5M));
- the second portion of the bar (“HbS”) represents the mean proportion of HbS as a percentage of total Hb (i.e., 77.7% (baseline), 5.4% (1M), 13.6% (1.5 M), 22.4% (2M), 33.2% (3M), 42.1% (4M), 46.7% (5M));
- the third portion of the bar (“HbA”) represents HbA (i.e., 10.2% (baseline), 84.8% (1M), 67.7% (1.5 M), 50.5% (2M), 28.8% (3M), 10.6% (4M), 1.9% (5M));
- the 164554159.2 Attorney Docket No.: 118945.8026.WO00 fourth portion of
- Hb g/dL
- Labels indicate mean proportion of HbS and HbF as a percentage of total Hb.
- Mean total Hb concentrations are shown directly above bars.
- Hb hemoglobin
- HbF fetal hemoglobin
- HbS sickle hemoglobin.
- the X axis shows months after autologous RNP27 edited CD34+ cell infusion. The region identified between dotted lines indicates the 12.0-16.0 g/dL normal range for women (Central laboratory reference range).
- Fig.4H depicts the percentage of F-cells for Subject 2 at baseline and at 1, 1.5, 2, 3, and 4 months after autologous RNP27 edited CD34+ cell infusion.
- the X axis shows months after autologous RNP27 edited CD34+ cell infusion.
- FIG.4I depicts the mean corpuscular HbF (pg) per F-cell for Subject 2 at baseline and at 1, 1.5, 2, 3, and 4 months after autologous RNP27 edited CD34+ cell infusion.
- Mean corpuscular HbF (pg) is shown directly above bars.
- Mean Corpuscular HbF: (MCH (pg)*HbF(%))/100% HbF(pg) per RBC.
- 10 pg/RBC is the threshold for protection from sickling and is shown by a dotted line (Steinberg 2014).
- the X axis shows months after autologous RNP27 edited CD34+ cell infusion.
- Fig.4J depicts hemoglobin fractionation (%) data for Subject 2 at various study days before and after autologous RNP27 edited CD34+ cell infusion.
- the lower portion of the bar (“HbF”) represents the mean proportion of HbF as a percentage of total hemoglobin (i.e., 10.5% (Study Day (“SD”) -450), 10.8% (SD -434), 8.4% (SD -9), 2.7% (SD -1), 8.2% (SD 34), 17.0% (SD 44), 25.8% (SD 62), 38.6% (SD 93);
- the middle portion of the bar (“Other”) represents other hemoglobin as a percentage of total hemoglobin (i.e., 15.7% (Study Day (“SD”) -450), 11.5% (SD -434), 53.4% (SD -9), 87.6% (SD -1), 86.4% (SD 34), 69.4% (SD 44), 51.8% (SD 62), 28.2% (SD 93); and the top portion of the bar (“HbS
- Labels indicate mean proportion of HbS and HbF as a percentage of total Hb.
- the percentage of F-Cells, total hemoglobin (g/dL), and MCH-F/F-cell (pg) are shown directly above the bar graph.
- Hb hemoglobin
- HbF fetal hemoglobin
- HbS sickle hemoglobin
- MCH mean corpuscular HbF
- RBC red blood cell.
- the X axis shows Study Days (“SD”) -450, -434, -9, 1, 34, 44, 62, and 93.
- Study Day 1 represents the day on which autologous RNP27 edited CD34+ cell infusion was administered.
- Figs.5A-5E depict the mean hemoglobin levels over time for subjects at baseline and months after autologous RNP27 edited CD34+ cell infusion.
- Figs.5A-5D the mean hemoglobin levels over time is depicted for subjects at baseline and months after autologous RNP27 edited CD34+ cell 16 164554159.2 Attorney Docket No.: 118945.8026.WO00 infusion.
- Hb g/dL
- Labels indicate mean proportion of HbF as a percentage of total Hb.
- Mean total Hb concentrations are shown directly above bars.
- Hb hemoglobin
- HbF fetal hemoglobin
- HbS sickle hemoglobin.
- the X axis shows months after autologous RNP27 edited CD34+ cell infusion, represented as a black vertical dotted line.
- the region identified with grey horizontal band indicates the 13.6-18.0 g/dL normal range for men patients (Central laboratory reference range) (Subjects 1 and 4) or 12.0-16.0 g/dL normal range for female patients (Subjects 2 and 3).
- the dark grey arrow shows the day of last red blood cell (RBC) transfusion.
- Fig.5E the mean hemoglobin levels over time is depicted for subjects at baseline and months after autologous RNP27 edited CD34+ cell infusion. Bars show mean total Hb (g/dL). Labels indicate mean proportion of HbF as a percentage of total Hb. Mean total Hb concentrations are shown directly above bars.
- the region identified with grey horizontal band indicates the 13.6-18.0 g/dL normal range for men patients (Central laboratory reference range) or 12.0-16.0 g/dL normal range for female patients.
- Fig.5A depicts data for Subject 1.
- the first (lowest) portion of the bar (“HbF,” medium grey) represents the mean proportion of HbF as a percentage of total hemoglobin (5.0% (baseline), 18.7% (1M), 28.1% (1.5M), 33.5% (2M), 36.7% (3M), 42.6% (4M), 45.4% (5M), 44.3% (6M), 44.1% (8M), 43.4% (10M));
- the second portion of the bar (“HbS,” black) represents the mean proportion of HbS as a percentage of total Hb;
- the third portion of the bar (“HbA/Transfused blood,” light grey) represents HbA;
- the fourth portion of the bar represents HbA2 (“HbA2,” medium grey);
- the fifth portion of the bar (only found at 4M) represents other hemoglobin (“Other Hb,” dark grey).
- Fig.5B depicts data for Subject 2.
- the first (lowest) portion of the bar (“HbF,” medium grey) represents the mean proportion of HbF as a percentage of total hemoglobin (10.8% (baseline), 8.2% (1M), 17.0% (1.5M), 25.8% (2M), 38.6% (3M), 46.2% (4M), 51.2% (5M), 51.3% (6M));
- the second portion of the bar (“HbS,” black) represents the mean proportion of HbS as a percentage of total Hb
- the third portion of the bar (“HbA/Transfused blood,” light grey) represents HbA
- the fourth (top) portion of the bar represents HbA2 (“HbA2,” medium grey).
- Fig.5C depicts data for Subject 3.
- the first (lowest) portion of the bar (“HbF,” medium grey) represents the proportion of HbF as a percentage of total hemoglobin (2.9% (baseline), 12.1% (1M), 23.7% (1.5M), 31.5% (2M), 42.6% (3M));
- the second portion of the bar (“HbS,” black) represents the mean proportion of HbS as a percentage of total Hb;
- the third portion of the bar (“HbA/Transfused blood,” light grey) represents HbA;
- the fourth (top) portion of the bar represents HbA2 (“HbA2,” medium grey).
- Bars show mean Hb (g/dL).
- Fig.5D depicts data for Subject 4.
- the first (lowest) portion of the bar (“HbF,” medium grey) represents the mean proportion of HbF as a percentage of total hemoglobin (6.1% (baseline), 6.7% (1M), 17.0% (1.5M), 26.6% (2M)); the second portion of the bar (“HbS,” black) represents the mean proportion of HbS as a percentage of total Hb; the third portion of the bar (“HbA/Transfused blood,” light grey) represents HbA; the fourth (top) portion of the bar represents HbA2 (“HbA2,” medium grey).
- Figs.6A-6B depicts the percentage of F-cells and mean corpuscular HbF for Subjects 1-4 at baseline and at various months after autologous RNP27 edited CD34+ cell infusion.
- Fig.6A depicts the percentage of F-cells for Subjects 1-4 at baseline and at various months after autologous RNP27 edited CD34+ cell infusion. Subject 1, medium grey dashed line; Subject 2, dark grey dashed line; Subject 3, solid grey line; and Subject 4, solid black line.
- the X axis shows months after autologous RNP27 edited CD34+ cell infusion, represented as a black vertical dotted line. An increasing percentage of F-cells indicates that more RBCs are protected from sickling for potential clinical benefit. *Data for Subject 2 at 3 months post-RNP27 edited CD34+ cell infusion are not available due to sample quality (hemolyzed sample).
- Fig.6B depicts the mean corpuscular HbF for Subjects 1- 4 at baseline and at various months after autologous RNP27 edited CD34+ cell infusion. Subject 1, medium grey long dashed line; Subject 2, black solid line; Subject 3, solid grey line; and Subject 4, light grey short dashed line.
- the X axis shows months after autologous RNP27 edited CD34+ cell infusion, represented as an grey vertical dotted line. Patients reach sickling-protective levels of HbF in F-cells by 1 month. *Data for Subject 2 at 3 months post-RNP27 edited CD34+ cell infusion are not available due to sample quality (hemolyzed sample).
- Fig.7 depicts the percentage of HBG1 and HBG2 promoter editing for Subjects 1-4 at various months after autologous RNP27 edited CD34+ cell infusion. Subject 1, medium grey long dashed line; Subject 2, black solid line; Subject 3, lower dot; and Subject 4, upper dot.
- the X axis shows months after autologous RNP27 edited CD34+ cell infusion.
- Figs.8A-8C depicts the percentage of HBG1 and HBG2 promoter editing.
- Fig.8A depicts the percentage of HBG1 and HBG2 promoter editing in the drug product and bone marrow (BM) of Subject 1.
- the bone marrow was sampled at 4 months post-RNP27 edited CD34+ cell infusion in the phase 1/2 clinical study.
- Fig.8B depicts the percentage of HBG1 and HBG2 promoter editing in the patient peripheral blood nucleated cells of Subject 1 (dark grey medium dash line), Subject 2 (black solid line), Subject 3 (medium grey solid line), Subject 4 (light grey medium dash line), Subject 5 (dark grey long dashed line), Subject 7 (light grey dotted line), Subject 8 (dark grey dotted line).
- the X axis shows months after autologous RNP27 edited CD34+ cell infusion.
- Fig.8C depicts the percentage of HBG1 and HBG2 promoter editing in the BM of Subject 1 (dark grey medium dashed line), Subject 2 (dark grey solid line), Subject 3 (grey dot), Subject 4 (dot at ⁇ 90%).
- Figs.9A-9C depicts the safety profile for subjects treated with RNP27 edited CD34+ cell infusion.
- Fig.9A depicts the safety profile for Subjects 1-4 treated with RNP27 edited CD34+ cell 18 164554159.2 Attorney Docket No.: 118945.8026.WO00 infusion.
- the safety profile is consistent with that of HSCT and myeloablative conditioning with bulsulfan.
- E number of events; HSCT, hematopoietic stem cell transplantation; TEAE, treatment emergent adverse event; TESAE, treatment emergent serious adverse event.
- Fig.9B depicts the safety profile for Subjects 1-10 treated with RNP27 edited CD34+ cell infusion.
- Fig.9C depicts the vaso-occlusive events (VOE) profile for Subjects 1-10 treated with RNP27 edited CD34+ cell infusion. At the time of the most recent data cutoff, all 10 patients who had been infused with RNP27 edited CD34+ cells and reached the Month 1 visit were completely VOE-free.
- Fig.10 depicts the sequences of Cpf1 proteins set forth in Table 11. Nuclear localization sequences are shown as bolded letters, six-histidine sequences are shown as underlined letters.
- NLS sequences e.g., appending two or more nNLS sequences or combinations of nNLS and sNLS sequences (or other NLS sequences) to either the N-terminal/C-terminal positions, as well as sequences with and without purification sequences, e.g., six-histidine sequences, are within the scope of the instantly disclosed subject matter.
- DETAILED DESCRIPTION Definitions and Abbreviations [0035] Unless otherwise specified, each of the following terms has the meaning associated with it in this section.
- An indel may be the product of the repair of a DNA double strand break, such as a double strand break formed by a genome editing system of the present disclosure.
- An indel is most commonly formed when a break is repaired by an “error prone” repair pathway such as the NHEJ pathway described below.
- “Gene conversion” refers to the alteration of a DNA sequence by incorporation of an endogenous homologous sequence (e.g. a homologous sequence within a gene array).
- Gene correction refers to the alteration of a DNA sequence by incorporation of an exogenous homologous sequence, such as an exogenous single-or double stranded donor template DNA.
- Indels, gene conversion, gene correction, and other genome editing outcomes are typically assessed by sequencing (most commonly by “next-gen” or “sequencing-by-synthesis” methods, though Sanger sequencing may still be used) and are quantified by the relative frequency of numerical changes (e.g., ⁇ 1, ⁇ 2 or more bases) at a site of interest among all sequencing reads.
- DNA samples for sequencing may be prepared by a variety of methods known in the art, and may involve the amplification of sites of interest by polymerase chain reaction (PCR), the capture of DNA ends generated by double strand breaks, as in the GUIDEseq process described in Tsai 2016 (incorporated by reference herein) or by other means well known in the art. Genome editing outcomes may also be assessed by in situ hybridization methods such as the FiberCombTM system commercialized by Genomic Vision (Bagneux, France), and by any other suitable methods known in the art.
- PCR polymerase chain reaction
- Alt-HDR “alternative homology-directed repair,” or “alternative HDR” are used interchangeably to refer to the process of repairing DNA damage using a homologous nucleic acid (e.g., an endogenous homologous sequence, e.g., a sister chromatid, or an exogenous nucleic acid, e.g., a template nucleic acid).
- Alt-HDR is distinct from canonical HDR in that the process utilizes different pathways from canonical HDR, and can be inhibited by the canonical HDR mediators, RAD51 and BRCA2.
- Alt-HDR is also distinguished by the involvement of a single-stranded or nicked homologous nucleic acid template, whereas canonical HDR generally involves a double- stranded homologous template.
- “Canonical HDR,” “canonical homology-directed repair” or “cHDR” refer to the process of repairing DNA damage using a homologous nucleic acid (e.g., an endogenous homologous sequence, 20 164554159.2 Attorney Docket No.: 118945.8026.WO00 e.g., a sister chromatid, or an exogenous nucleic acid, e.g., a template nucleic acid).
- Canonical HDR typically acts when there has been significant resection at the double strand break, forming at least one single stranded portion of DNA.
- cHDR typically involves a series of steps such as recognition of the break, stabilization of the break, resection, stabilization of single stranded DNA, formation of a DNA crossover intermediate, resolution of the crossover intermediate, and ligation. The process requires RAD51 and BRCA2, and the homologous nucleic acid is typically double- stranded.
- HDR as used herein encompasses both canonical HDR and alt-HDR.
- Non-homologous end joining refers to ligation mediated repair and/or non- template mediated repair including canonical NHEJ (cNHEJ) and alternative NHEJ (altNHEJ), which in turn includes microhomology-mediated end joining (MMEJ), single-strand annealing (SSA), and synthesis-dependent microhomology-mediated end joining (SD-MMEJ).
- Replacement or “replaced,” when used with reference to a modification of a molecule (e.g. a nucleic acid or protein), does not require a process limitation but merely indicates that the replacement entity is present.
- Subject means a human, mouse, or non-human primate.
- a human subject can be any age (e.g., an infant, child, young adult, or adult), and may suffer from a disease, or may be in need of alteration of a gene.
- “Treat,” “treating,” and “treatment” mean the treatment of a disease in a subject (e.g., a human subject), including one or more of inhibiting the disease, i.e., arresting or preventing its development or progression; relieving the disease, i.e., causing regression of the disease state; relieving one or more symptoms of the disease; and curing the disease.
- kits refers to any collection of two or more components that together constitute a functional unit that can be employed for a specific purpose.
- one kit according to this disclosure can include a guide RNA complexed or able to complex with an RNA-guided nuclease, and accompanied by (e.g. suspended in, or suspendable in) a pharmaceutically acceptable carrier.
- the kit may include a booster element.
- the kit can be used to introduce the complex into, for example, a cell or a subject, for the purpose of causing a desired genomic alteration in such cell or subject.
- the components of a kit can be packaged 21 164554159.2 Attorney Docket No.: 118945.8026.WO00 together, or they may be separately packaged.
- Kits according to this disclosure also optionally include directions for use (DFU) that describe the use of the kit e.g., according to a method of this disclosure.
- the DFU can be physically packaged with the kit, or it can be made available to a user of the kit, for instance by electronic means.
- polynucleotide refers to a series of nucleotide bases (also called “nucleotides”) in DNA and RNA, and mean any chain of two or more nucleotides.
- the polynucleotides, nucleotide sequences, nucleic acids etc. can be chimeric mixtures or derivatives or modified versions thereof, single-stranded or double-stranded. They can be modified at the base moiety, sugar moiety, or phosphate backbone, for example, to improve stability of the molecule, its hybridization parameters, etc.
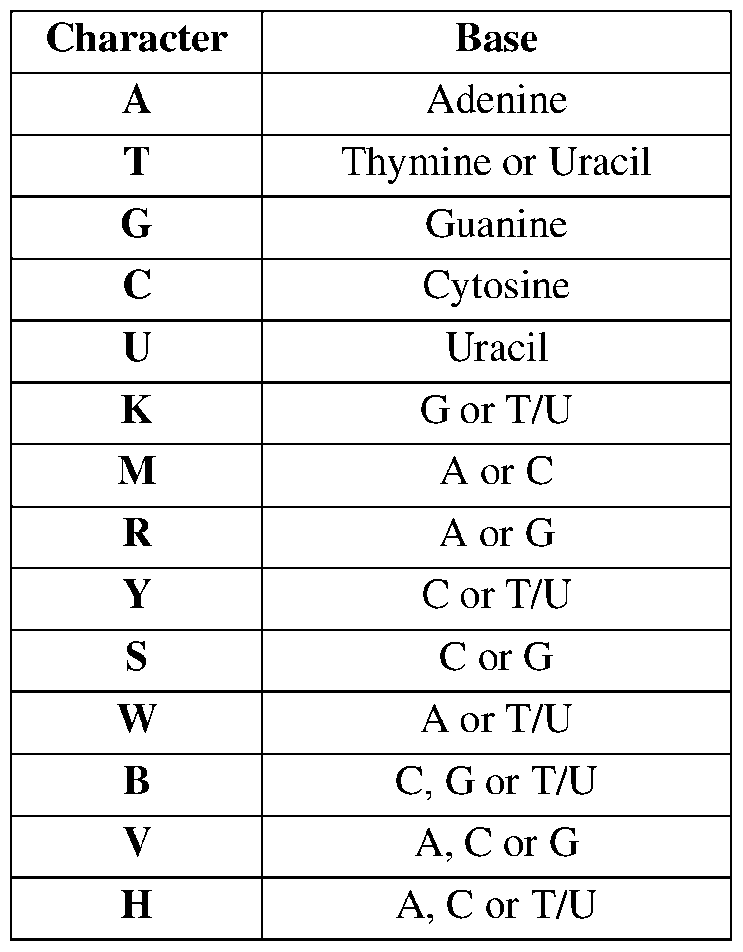
- a nucleotide sequence typically carries genetic information, including, but not limited to, the information used by cellular machinery to make proteins and enzymes. These terms include double- or single-stranded genomic DNA, RNA, any synthetic and genetically manipulated polynucleotide, and both sense and antisense polynucleotides. These terms also include nucleic acids containing modified bases. [0052] Conventional IUPAC notation is used in nucleotide sequences presented herein, as shown in Table 1, below (see also Cornish-Bowden A, Nucleic Acids Res.1985 May 10; 13(9):3021-30, incorporated by reference herein).
- T denotes “Thymine or Uracil” in those instances where a sequence may be encoded by either DNA or RNA, for example in gRNA targeting domains.
- Table 1 IUPAC nucleic acid notation Character Base A Ad i 164554159.2 Attorney Docket No.: 118945.8026.WO00 D A, G or T/U N A, C, G or T/U [0053]
- protein erchangeably to refer to a sequential chain of amino ac s e oge e va pep e o s. e terms include individual proteins, groups or complexes of proteins that associate together, as well as fragments or portions, variants, derivatives and analogs of such proteins.
- CCAAT box target region refers to a sequence that is 5’ of the transcription start site (TSS) of the HBG1 and/or HBG2 gene.
- TSS transcription start site
- the distal CCAAT box of HBG1 and HBG2 is shown in the schematic in Fig.3.
- CCAAT boxes are highly conserved motifs within the promoter region of ⁇ -like and ⁇ -like globin genes. The regions within or near the CCAAT box play important roles in globin gene regulation.
- the ⁇ -globin distal CCAAT box is associated with hereditary persistence of fetal hemoglobin.
- a number of transcription factors have been reported to bind to the duplicated CCAAT box region of the ⁇ -globin promoter, e.g., NF-Y, COUP-TFII (NF- E3), CDP, GATA1/NF-E1 and DRED (Martyn 2017). While not wishing to be bound by theory, it is believed that the binding sites of the transcriptional activator NF-Y overlaps with transcriptional repressors at the ⁇ -globin promoter.
- HPFH mutations present within the distal ⁇ -globin promoter region may alter the competitive binding of those factors and thus contribute to the increased ⁇ -globin expression and elevated levels of HbF.
- Genomic locations provided herein for HBG1 and HBG2 are based on the coordinates provided in NCBI Reference Sequence NC_000011, “Homo sapiens chromosome 11, GRCh38.p12 Primary Assembly,” (Version NC_000011.10).
- the distal CCAAT box of HBG1 and HBG2 is positioned at HBG1 and HBG2 c.- 111 to -115 (Genomic location is Hg38 Chr11:5,249,968 to Chr11:5,249,972 and Hg38 Chr11:5,254,892 to Chr11:5,254,896, respectively).
- the HBG1 c.-111 to -115 region is exemplified in SEQ ID NO:198 (HBG1) at positions 2823-2827
- the HBG2 c.-111 to -115 region is exemplified in SEQ ID NO:199 (HBG2) at positions 2747-2751.
- the “CCAAT box target region” denotes the region that is at or near the distal CCAAT box and includes the nucleotides of the distal CCAAT box and 25 nucleotides upstream (5’) and 25 nucleotides downstream (3’) of the distal CCAAT box (i.e., HBG1/2 c.-86 to -140) (Genomic location is Hg38 Chr11:5249943 to Hg38 Chr11:5249997 and Hg38 Chr11:5254867 to Hg38 Chr11:5254921, respectively).
- HBG1 c.-86 to -140 region is exemplified in SEQ ID NO:198 (HBG1) at positions 2798-2852, and the HBG2 c.-86 to -140 region is exemplified in SEQ ID NO:199 (HBG2) at positions 2723-2776.
- the “CCAAT box target region” denotes the region that is at or near the distal CCAAT box and includes the nucleotides of the distal CCAAT box and 5 nucleotides 23 164554159.2 Attorney Docket No.: 118945.8026.WO00 upstream (5’) and 5 nucleotides downstream (3’) of the distal CCAAT box (i.e., HBG1/2 c.-106 to - 120 (Genomic location is Hg38 Chr11:5249963 to Hg38 Chr11:5249977 (HGB1 and Hg38 Chr11:5254887 to Hg38 Chr11:5254901, respectively)).
- the HBG1 c.-106 to -120 region is exemplified in SEQ ID NO:198 (HBG1) at positions 2818-2832, and the HBG2 c.-106 to -120 region is exemplified in SEQ ID NO:199 (HBG2) at positions 2742-2756.
- CCAAT box target site alteration refers to alterations (e.g., deletions, insertions, mutations) of one or more nucleotides of the CCAAT box target region.
- Examples of exemplary CCAAT box target region alterations include, without limitation, the 1 nt deletion, 4 nt deletion, 11nt deletion, 13 nt deletion, and 18 nt deletion, and -117 G>A alteration.
- CCAAT box and “CAAT box” can be used interchangeably.
- the notations “c.-114 to -102 region,” “c.-102 to -114 region,” “-102:-114,” “13 nt target region” and the like refer to a sequence that is 5’ of the transcription start site (TSS) of the HBG1 and/or HBG2 gene at the genomic location Hg38 Chr11:5,249,959 to Hg38 Chr11:5,249,971 and Hg38 Chr11:5,254,883 to Hg38 Chr11:5,254,895, respectively.
- TSS transcription start site
- the HBG1 c.-102 to -114 region is exemplified in SEQ ID NO:198 (HBG1) at positions 2824-2836 and the HBG2 c.-102 to -114 region is exemplified in SEQ ID NO:199 (HBG2) at positions 2748-2760.
- the term “13 nt deletion” and the like refer to deletions of the 13 nt target region.
- the notations “c.-121 to -104 region,” “c.-104 to -121 region,” “-104:-121,” “18 nt target region,” and the like refer to a sequence that is 5’ of the transcription start site (TSS) of the HBG1 and/or HBG2 gene at the genomic location Hg38 Chr11:5,249,961 to Hg38 Chr11:5,249,978 and Hg38 Chr11:5,254,885 to Hg38 Chr11: 5,254,902, respectively.
- TSS transcription start site
- HBG1 c.-104 to -121 region is exemplified in SEQ ID NO:198 (HBG1) at positions 2817-2834
- HBG2 c.-104 to -121 region is exemplified in SEQ ID NO:199 (HBG2) at positions 2741-2758.
- the term “18 nt deletion” and the like refer to deletions of the 18 nt target region.
- the notations “c.-105 to -115 region,” “c.-115 to -105 region,” “-105:-115,” “11 nt target region,” and the like refer to a sequence that is 5’ of the transcription start site (TSS) of the HBG1 and/or HBG2 gene at the genomic location Hg38 Chr11:5,249,962 to Hg38 Chr11:5,249,972 and Hg38 Chr11:5,254,886 to Hg38 Chr11:5,254,896, respectively.
- TSS transcription start site
- the HBG1 c.-105 to -115 region is exemplified in SEQ ID NO:198 (HBG1) at positions 2823-2833, and the HBG2 c.-105 to -115 region is exemplified in SEQ ID NO:199 (HBG2) at positions 2747-2757.
- the term “11 nt deletion” and the like refer to deletions of the 11 nt target region.
- the notations “c.-115 to -112 region,” “c.-112 to -115 region,” “-112:-115,” “4 nt target region,” and the like refer to a sequence that is 5’ of the transcription start site (TSS) of the HBG1 and/or HBG2 gene at the genomic location Hg38 Chr11:5,249,969 to Hg38 Chr11:5,249,972 and 24 164554159.2 Attorney Docket No.: 118945.8026.WO00 Hg38 Chr11:5,254,893 to Hg38 Chr11:5,254,896, respectively.
- the HBG1 c.-112 to -115 region is exemplified in SEQ ID NO:198 at positions 2823-2826, and the HBG2 c.-112 to -115 region is exemplified in SEQ ID NO:199 (HBG2) at positions 2747-2750.
- the term “4 nt deletion” and the like refer to deletions of the 4 nt target region.
- the notations “c.-116 region,” “HBG-116,” “1 nt target region,” and the like refer to a sequence that is 5’ of the transcription start site (TSS) of the HBG1 and/or HBG2 gene at the genomic location Hg38 Chr11:5,249,973 and Hg38 Chr11:5,254,897, respectively.
- TSS transcription start site
- the HBG1 c.-116 region is exemplified in SEQ ID NO:198 at position 2822, and the HBG2 c.-116 region is exemplified in SEQ ID NO:199 (HBG2) at position 2746.
- the term “1 nt deletion” and the like refer to deletions of the 1 nt target region.
- the notations “c.-117 G>A region,” “HBG-117 G>A,” “-117 G>A target region” and the like refer to a sequence that is 5’ of the transcription start site (TSS) of the HBG1 and/or HBG2 gene at the genomic location Hg38 Chr11:5,249,974 to Hg38 Chr11:5,249,974 and Hg38 Chr11:5,254,898 to Hg38 Chr11:5,254,898, respectively.
- TSS transcription start site
- the HBG1 c.-117 G>A region is exemplified by a substitution from guanine (G) to adenine (A) in SEQ ID NO:198 at position 2821
- the HBG2 c.-117 G>A region is exemplified by a substitution from G to A in SEQ ID NO:199 (HBG2) at position 2745.
- the term “-117 G>A alteration” and the like refer to a substitution from G to A at the -117G>A target region.
- proximal HBG1/2 promoter target sequence denotes the region within 50, 100, 200, 300, 400, or 500 bp of a proximal HBG1/2 promoter sequence including the 13 nt target region.
- the various embodiments of this disclosure generally relate to genome editing systems configured to introduce alterations (e.g., a deletion or insertion, or other mutation) into chromosomal 25 164554159.2 Attorney Docket No.: 118945.8026.WO00 DNA that enhance transcription of the HBG1 and/or HBG2 genes, which encode the A ⁇ and G ⁇ subunits of hemoglobin, respectively.
- the disclosure generally relates to the use of RNP complexes comprising a gRNA complexed to a Cpf1 protein.
- the gRNA may be unmodified or modified.
- the gRNA may comprise a sequence set forth in Table 14.
- a Cpf1 protein may comprise a sequence set forth in SEQ ID NOs:200, 201, 205-215, 221-226, 280-283, 293-295 (Cpf1 polypeptide sequences).
- a Cpf1 protein may be encoded by a sequence comprising SEQ ID NOs:216-218, 296-303 (Cpf1 polynucleotide sequences).
- the RNP complex may comprise an RNP complex set forth in Table 12.
- the RNP complex may include a gRNA comprising the sequence set forth in SEQ ID NO:237 and a Cpf1 protein comprising a sequence set forth in SEQ ID NO:283 (RNP27, Table 12).
- HPFH Hereditary Persistence of Fetal Hemoglobin
- HbF expression can be induced through point mutations in an ⁇ –globin regulatory element that is associated with a naturally occurring HPFH variant, including, for example, HBG1 c.-114 C>T; c.-117 G>A; c.-158 C>T; c.-167 C>T; c.-170 G>A; c.-175 T>G; c.-175 T>C; c.-195 C>G; c.- 196 C>T; c.-197 C>T; c.-198 T>C; c.-201 C>T; c.-202 C>T; c.-211 C>T, c.-251 T>C; or c.-499 T>A; or HBG2 c.-109 G>T; c.-110 A>C; c.-114 C>A; c.-114 C>T; c.-114 C>G; c.-157 C>T; c.-158
- Naturally occurring mutations at the distal CCAAT box motif found within the promoter of the HBG1 and/or HBG2 genes have also been shown to result in continued ⁇ –globin expression and the HPFH condition. It is thought that alteration (mutation or deletion) of the CCAAT box may disrupt the binding of one or more transcriptional repressors, resulting in continued expression of the ⁇ –globin gene and elevated HbF expression (Martyn 2017). For example, a naturally occurring 13 base pair del c.-114 to -102 (“13 nt deletion”) has been shown to be associated with elevated levels of HbF (Martyn 2017).
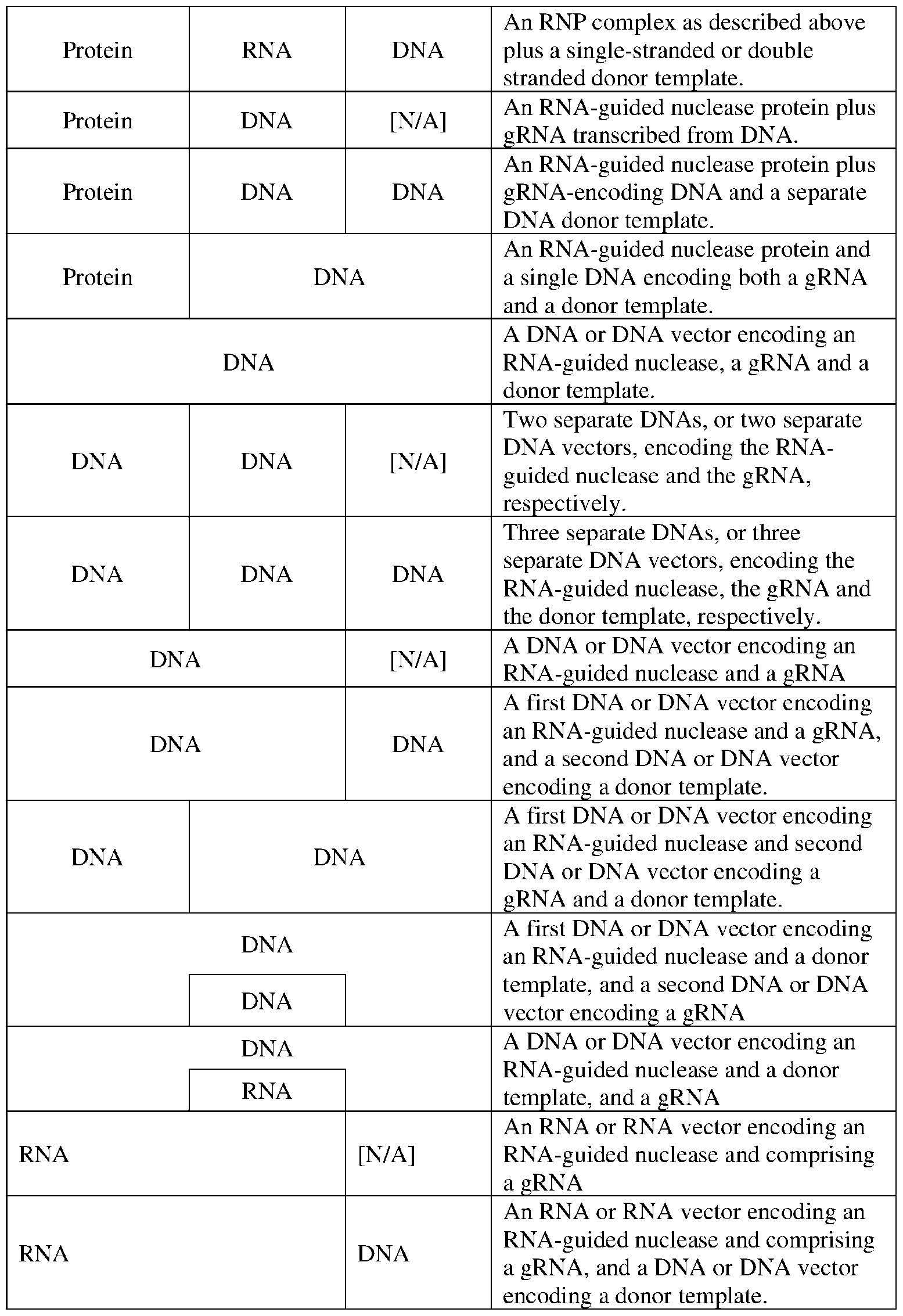
- the genome editing systems of this disclosure can include an RNA-guided nuclease such as Cpf1 protein and one or more gRNAs having a targeting domain that is complementary to a sequence in or near the target region, and optionally one or more of a DNA donor template that encodes a specific mutation (such as a deletion or insertion) in or near the target region, and/or an agent that enhances the efficiency with which such mutations are generated including, without limitation, a random oligonucleotide, a small molecule agonist or antagonist of a gene product involved in DNA repair or a DNA damage response, or a peptide agent.
- an RNA-guided nuclease such as Cpf1 protein and one or more gRNAs having a targeting domain that is complementary to a sequence in or near the target region, and optionally one or more of a DNA donor template that encodes a specific mutation (such as a deletion or insertion) in or near the target region, and/or an agent that enhances the efficiency with which such mutation
- hemoglobinopathies by gene therapy and/or genome editing is complicated by the fact that the cells that are phenotypically affected by the disease, erythrocytes or RBCs, are enucleated, and do not contain genetic material encoding either the aberrant hemoglobin protein (Hb) subunits nor the A ⁇ or G ⁇ subunits targeted in the exemplary genome editing approaches described above.
- Hb hemoglobin protein
- This complication is addressed, in certain embodiments of this disclosure, by the alteration of cells that are competent to differentiate into, or otherwise give rise to, erythrocytes.
- Cells within the erythroid lineage that are altered according to various embodiments of this disclosure include, without limitation, hematopoietic stem and progenitor cells (HSCs), erythroblasts (including basophilic, polychromatic and/or orthochromatic erythroblasts), proerythroblasts, polychromatic erythrocytes or reticulocytes, embryonic stem (ES) cells, and/or induced pluripotent stem (iPSC) cells.
- HSCs hematopoietic stem and progenitor cells
- erythroblasts including basophilic, polychromatic and/or orthochromatic erythroblasts
- proerythroblasts include polychromatic erythrocytes or reticulocytes
- ES embryonic stem
- iPSC induced pluripotent stem
- alterations that result in induction of A ⁇ and/or G ⁇ expression are obtained through the use of a genome editing system comprising an RNA-guided nuclease and at least one gRNA having a targeting domain complementary to a sequence within the CCAAT box target region of HBG1 and/or HBG2 or proximate thereto (e.g., within 10, 20, 30, 40, or 50, 100, 200, 300, 400 or 500 bases of the CCAAT box target region).
- the RNA- guided nuclease and gRNA form a complex that is capable of associating with and altering the CCAAT box target region or a region proximate thereto.
- Suitable gRNAs and gRNA targeting domains directed to the CCAAT box target region of HBG1 and/or HBG2 or proximate thereto include those set forth herein.
- alterations that result in induction of A ⁇ and/or G ⁇ expression are obtained through the use of a genome editing system comprising an RNA-guided nuclease and at least one gRNA having a targeting domain complementary to a sequence within the 13 nt target region of HBG1 and/or HBG2 or proximate thereto (e.g., within 10, 20, 30, 40, or 50, 100, 200, 300, 400 or 500 bases of the 13 nt target region).
- RNA-guided nuclease and gRNA form a complex that is capable of associating with and altering the 13 nt target region or a 27 164554159.2 Attorney Docket No.: 118945.8026.WO00 region proximate thereto.
- suitable gRNAs and gRNA targeting domains directed to the 13 nt target region of HBG1 and/or HBG2 or proximate thereto for use in the embodiments disclosed herein include those set forth herein.
- the genome editing system can be implemented in a variety of ways, as is discussed below in detail.
- a genome editing system of this disclosure can be implemented as a ribonucleoprotein complex or a plurality of complexes in which multiple gRNAs are used.
- This ribonucleoprotein complex can be introduced into a target cell using art-known methods, including electroporation, as described in commonly-assigned International Patent Publication No. WO 2016/182959 by Jennifer Gori ("Gori"), published Nov.17, 2016, which is incorporated by reference in its entirety herein.
- the ribonucleoprotein complexes within these compositions are introduced into target cells by art-known methods, including without limitation electroporation (e.g.
- ribonucleoprotein complexes are formed within the target cells themselves following introduction of nucleic acids encoding the RNA-guided nuclease and/or gRNA. These and other delivery modalities are described in general terms below and in Gori. [0073] Cells that have been altered ex vivo according to this disclosure can be manipulated (e.g.
- an autologous transplant includes the steps of obtaining, from the subject, a plurality of cells, either circulating in peripheral blood, or within the marrow or other tissue (e.g. spleen, skin, etc.), and optionally manipulating those cells to enrich for cells in the erythroid lineage (e.g.
- the cells are, optionally or additionally, expanded, transduced with a transgene, exposed to a cytokine or other peptide or small molecule agent, and/or frozen/thawed prior to transduction with a genome editing system targeting the CCAAT box target region, the 13 nt target region, and/or proximal HBG1/2 promoter target sequence.
- the genome editing system can be implemented or delivered to the cells in any suitable format, including as a 28 164554159.2 Attorney Docket No.: 118945.8026.WO00 ribonucleoprotein complex, as separated protein and nucleic acid components, and/or as nucleic acids encoding the components of the genome editing system.
- CD34+ hematopoietic stem and progenitor cells that have been edited using the genome editing methods disclosed herein may be used for the treatment of a ⁇ - hemoglobinopathy in a subject in need thereof.
- the ⁇ -hemoglobinopathy may be severe sickle cell disease (SCD) or thalassemia, such as ⁇ -thalassemia, ⁇ -thalassemia, or ⁇ / ⁇ - thalassemia.
- an exemplary protocol for treatment of a ⁇ -hemoglobinopathy may include harvesting CD34+ HSPCs from a subject in need thereof, ex vivo editing of the autologous CD34+ HSPCs using the genome editing methods disclosed herein, followed by reinfusion of the edited autologous CD34+ HSPCs into the subject.
- treatment with edited autologous CD34+ HSPCs may result in increased HbF induction.
- a subject may discontinue treatment with hydroxyurea, if applicable, and receive blood transfusions to maintain sufficient hemoglobin (Hb) levels.
- a subject may be administered intravenous plerixafor (e.g., 0.24 mg/kg) to mobilize CD34+ HSPCs from bone marrow into peripheral blood.
- a subject may undergo one or more leukapheresis cycles.
- a subject may undergo approximately one month between leukapheresis cycles, with one cycle defined as two plerixafor-mobilized leukapheresis collections performed on consecutive days.
- a subject may undergo approximately one month between leukapheresis cycles, with one cycle defined as two, three, or more plerixafor-mobilized leukapheresis collections performed on consecutive dates or with one day, two days, three days, four days, or five days between leukapheresis collections.
- the number of leukapheresis cycles performed for a subject may be the number required to achieve a dose of edited autologous CD34+ HSPCs (e.g., ⁇ 2 x 10 6 cells/kg, ⁇ 3 x 10 6 cells/kg, ⁇ 4 x 10 6 cells/kg, ⁇ 5 x 10 6 cells/kg, ⁇ 6 x 10 6 cells/kg, ⁇ 7 x 10 6 cells/kg, ⁇ 8 x 10 6 cells/kg, ⁇ 9 x 10 6 cells/kg, ⁇ 10 x 10 6 cells/kg, ⁇ 11 x 10 6 cells/kg, ⁇ 12 x 10 6 cells/kg, ⁇ 13 x 10 6 cells/kg, ⁇ 14 x 10 6 cells/kg, ⁇ 15 x 10 6 cells/kg, ⁇ 16 x 10 6 cells/kg, ⁇ 17 x 10 6 cells/kg, ⁇ 18 x 10 6 cells/kg, ⁇ 19 x 10 6 cells/kg, ⁇ 20 x 10 6 cells
- the CD34+ HSPCs harvested from the subject may be edited using any of the genome editing methods discussed herein.
- any one or more of the gRNAs and one or more of the RNA-guided nucleases disclosed herein may be used in the genome editing methods.
- the treatment may include an autologous stem cell transplant.
- a subject may undergo myeloablative conditioning with busulfan conditioning (e.g., dose-adjusted based on first-dose pharmacokinetic analysis, with a test dose of 1 mg/kg).
- conditioning may occur for four consecutive days.
- edited autologous CD34+ HSPCs may be administered to the subject.
- the busulfan washout period may be, for example, 1 day, 2 days, 3 days, 4 days, 5 days, 6 days, 7 days, 8 days, 9 days, 10 days, 11 days, 12 days, 13 days, 14 days or longer.
- edited autologous CD34+ HSPCs e.g., about ⁇ 1 x 10 6 cells/kg, ⁇ 2 x 10 6 cells/kg, ⁇ 3 x 10 6 cells/kg, ⁇ 4 x 10 6 cells/kg, ⁇ 5 x 10 6 cells/kg, ⁇ 6 x 10 6 cells/kg, ⁇ 7 x 10 6 cells/kg, ⁇ 8 x 10 6 cells/kg, ⁇ 9 x 10 6 cells/kg, ⁇ 10 x 10 6 cells/kg, ⁇ 11 x 10 6 cells/kg, ⁇ 12 x 10 6 cells/kg, ⁇ 13 x 10 6 cells/kg, ⁇ 14 x 10 6 cells/kg, ⁇ 15 x 10 6 cells/kg, ⁇ 16 x 10 6 cells/kg, ⁇ 17 x 10 6 cells/kg, ⁇ 18 x 10 6 cells/kg, ⁇ 19 x 10 6 cells/kg, ⁇ 20 x 10 6 cells/kg, ⁇
- the edited autologous CD34+ HSPCs may be manufactured and cryopreserved for a particular subject.
- a subject may attain neutrophil engraftment following a sequential myeloablative conditioning regimen and infusion of edited autologous CD34+ cells.
- Neutrophil engraftment may be defined as three consecutive measurements of ANC ⁇ 0.5 x 10 9 /L on different days.
- a genome editing system may include, or may be co- delivered with, one or more factors that improve the viability of the cells during and after editing, including without limitation an aryl hydrocarbon receptor antagonist such as StemRegenin-1 (SR1), UM171, LGC0006, alpha-napthoflavone, and CH-223191, and/or an innate immune response antagonist such as cyclosporin A, dexamethasone, reservatrol, a MyD88 inhibitory peptide, an RNAi agent targeting Myd88, a B18R recombinant protein, a glucocorticoid, OxPAPC, a TLR antagonist, rapamycin, BX795, and a RLR shRNA.
- an aryl hydrocarbon receptor antagonist such as StemRegenin-1 (SR1), UM171, LGC0006, alpha-napthoflavone, and CH-223191
- an innate immune response antagonist such as cyclosporin A, dexamethasone,
- the cells following delivery of the genome editing system, are optionally manipulated e.g. to enrich for HSCs and/or cells in the erythroid lineage and/or for edited cells, to expand them, freeze/thaw, or otherwise prepare the cells for return to the subject.
- the edited cells are then returned 30 164554159.2 Attorney Docket No.: 118945.8026.WO00 to the subject, for instance in the circulatory system by means of intravenous delivery or delivery or into a solid tissue such as bone marrow.
- alteration of the CCAAT box target region, the 13 nt target region, and/or proximal HBG1/2 promoter target sequence using the compositions, methods and genome editing systems of this disclosure results in significant induction, among hemoglobin-expressing cells, of A ⁇ and/or G ⁇ subunits (referred to interchangeably as HbF expression), e.g. at least 5%, 10%, 15%, 20%, 25%, 30%, 35%, 40%, 45%, 50% or greater induction of A ⁇ and/or G ⁇ subunit expression relative to unmodified controls.
- This induction of protein expression is generally the result of alteration of the CCAAT box target region, 13 nt target region, and/or proximal HBG1/2 promoter target sequence (expressed, e.g.
- the functional effects of alterations caused or facilitated by the genome editing systems and methods of the present disclosure can be assessed in any number of suitable ways.
- HBG1 and HBG2 mRNA can be assessed by digital droplet PCR (ddPCR), which is performed on cDNA samples obtained by reverse transcription of mRNA harvested from treated or untreated samples.
- Primers for HBG1, HBG2, HBB, and/or HBA may be used individually or multiplexed using methods known in the art.
- ddPCR analysis of samples may be conducted using the QX200TM ddPCR system commercialized by Bio Rad (Hercules, CA), and associated protocols published by BioRad.
- Fetal hemoglobin protein may be assessed by high pressure liquid chromatography (HPLC), for example, according to the methods discussed on pp.143- 44 in Chang 2017 (incorporated by reference herein), or fast protein liquid chromatography (FPLC), using ion-exchange and/or reverse phase columns to resolve HbF, HbB and HbA and/or A ⁇ and G ⁇ globin chains as is known in the art.
- HPLC high pressure liquid chromatography
- FPLC fast protein liquid chromatography
- the rate at which the CCAAT box target region is altered in the target cells can be modified by the use of optional genome editing system components such as oligonucleotide donor templates.
- Donor template design is described in general terms below under the heading “Donor template design.”
- the embodiments described herein may be used in all classes of vertebrate including, but not limited to, primates, mice, rats, rabbits, pigs, dogs, and cats. 31 164554159.2 Attorney Docket No.: 118945.8026.WO00 [0084]
- This overview has focused on a handful of exemplary embodiments that illustrate the principles of genome editing systems and CRISPR-mediated methods of altering cells. For clarity, however, this disclosure encompasses modifications and variations that have not been expressly addressed above, but will be evident to those of skill in the art. With that in mind, the following disclosure is intended to illustrate the operating principles of genome editing systems more generally.
- Genome editing systems refers to any system having RNA-guided DNA editing activity. Genome editing systems of the present disclosure include at least two components adapted from naturally occurring CRISPR systems: a guide RNA (gRNA) and an RNA-guided nuclease.
- gRNA guide RNA
- Genome editing systems can be implemented (e.g. administered or delivered to a cell or a subject) in a variety of ways, and different implementations may be suitable for distinct applications.
- a genome editing system is implemented, in certain embodiments, as a protein/RNA complex (a ribonucleoprotein, or RNP), which can be included in a pharmaceutical composition that optionally includes a pharmaceutically acceptable carrier and/or an encapsulating agent, such as, without limitation, a lipid or polymer micro- or nano-particle, micelle, or liposome.
- a protein/RNA complex a ribonucleoprotein, or RNP
- RNP ribonucleoprotein
- an encapsulating agent such as, without limitation, a lipid or polymer micro- or nano-particle, micelle, or liposome.
- a genome editing system is implemented as one or more nucleic acids encoding the 32 164554159.2 Attorney Docket No.: 118945.8026.WO00 RNA-guided nuclease and guide RNA components described above (optionally with one or more additional components); in certain embodiments, the genome editing system is implemented as one or more vectors comprising such nucleic acids, for instance a viral vector such as an adeno-associated virus (see section below under the heading “Implementation of genome editing systems: delivery, formulations, and routes of administration”); and in certain embodiments, the genome editing system is implemented as a combination of any of the foregoing.
- the genome editing systems of the present disclosure can be targeted to a single specific nucleotide sequence, or may be targeted to — and capable of editing in parallel — two or more specific nucleotide sequences through the use of two or more guide RNAs.
- the use of multiple gRNAs is referred to as “multiplexing” throughout this disclosure, and can be employed to target multiple, unrelated target sequences of interest, or to form multiple SSBs or DSBs within a single target domain and, in some cases, to generate specific edits within such target domain.
- multiplexing can be employed to target multiple, unrelated target sequences of interest, or to form multiple SSBs or DSBs within a single target domain and, in some cases, to generate specific edits within such target domain.
- WO 2015/138510 by Maeder et al. (“Maeder”), which is incorporated by reference herein, describes a genome editing system for correcting a point mutation (C.2991+1655A to G) in the human CEP290 gene that results in the creation of a cryptic splice site, which in turn reduces or eliminates the function of the gene.
- the genome editing system of Maeder utilizes two guide RNAs targeted to sequences on either side of (i.e. flanking) the point mutation, and forms DSBs that flank the mutation. This, in turn, promotes deletion of the intervening sequence, including the mutation, thereby eliminating the cryptic splice site and restoring normal gene function.
- Cotta-Ramusino WO 2016/073990 by Cotta-Ramusino et al.
- Cotta-Ramusino describes a genome editing system that utilizes two gRNAs in combination with a Cas9 nickase (a Cas9 that makes a single strand nick such as S.
- the dual-nickase system of Cotta-Ramusino is configured to make two nicks on opposite strands of a sequence of interest that are offset by one or more nucleotides, which nicks combine to create a double strand break having an overhang (5’ in the case of Cotta-Ramusino, though 3’ overhangs are also possible).
- the overhang in turn, can facilitate homology directed repair events in some circumstances.
- a gRNA targeted to a nucleotide sequence encoding Cas9 (referred to as a “governing RNA”), which can be included in a genome editing system comprising one or more additional gRNAs to permit transient expression of a Cas9 that might otherwise be constitutively expressed, for example in some virally transduced cells.
- governing RNA nucleotide sequence encoding Cas9
- Genome editing systems can, in some instances, form double strand breaks that are repaired by cellular DNA double-strand break mechanisms such as NHEJ or HDR. These mechanisms are described throughout the literature (see, e.g., Davis & Maizels 2014 (describing Alt-HDR); Frit 2014 (describing Alt-NHEJ); Iyama & Wilson 2013 (describing canonical HDR and NHEJ pathways generally)).
- NHEJ DNA double-strand break mechanism
- DSBs such systems optionally include one or more components that promote or facilitate a particular mode of double-strand break repair or a particular repair outcome.
- Cotta-Ramusino also describes genome editing systems in which a single stranded oligonucleotide “donor template” is added; the donor template is incorporated into a target region of cellular DNA that is cleaved by the genome editing system, and can result in a change in the target sequence.
- editing of HBG1 and HBG2 promoters may occur.
- editing of HBG1 and HBG2 promoters in peripheral blood nucleated cells post RNP27 edited CD34+ cell infusion may occur.
- editing levels in peripheral blood nucleated cells may be >80%, >85%, >90%, >95%, >98%.
- editing of HBG1 and HBG2 promoters in bone marrow post RNP27 edited CD34+ cell infusion may occur.
- editing levels in bone marrow may be >80%, >85%, >90%, >95%, >98%.
- Guide RNA (gRNA) molecules [0093] The terms “guide RNA” and “gRNA” refer to any nucleic acid that promotes the specific association (or “targeting”) of an RNA-guided nuclease such as a Cpf1 protein to a target sequence such as a genomic or episomal sequence in a cell.
- gRNAs can be unimolecular (comprising a single RNA molecule, and referred to alternatively as chimeric), or modular (comprising more than one, and typically two, separate RNA molecules, such as a crRNA and a tracrRNA, which are usually associated with one another, for instance by duplexing).
- gRNAs and their component parts are described throughout the literature (see, e.g., Briner 2014, which is incorporated by reference; Cotta- Ramusino).
- Examples of modular and unimolecular gRNAs that may be used according to the embodiments herein include, without limitation, the sequences set forth in SEQ ID NOs:21-23 and 30-43.
- gRNA proximal and tail domains examples include, without limitation, the sequences set forth in SEQ ID NOs:24-29.
- type II CRISPR systems generally comprise an RNA-guided nuclease protein such as Cas9, a CRISPR RNA (crRNA) that includes a 5’ region that is complementary to a 34 164554159.2 Attorney Docket No.: 118945.8026.WO00 foreign sequence, and a trans-activating crRNA (tracrRNA) that includes a 5’ region that is complementary to, and forms a duplex with, a 3’ region of the crRNA.
- crRNA CRISPR RNA
- tracrRNA trans-activating crRNA
- Guide RNAs include a “targeting domain” that is fully or partially complementary to a target domain within a target sequence, such as a DNA sequence in the genome of a cell where editing is desired.
- Targeting domains are referred to by various names in the literature, including without limitation “guide sequences” (Hsu 2013, incorporated by reference herein), “complementarity regions” (Cotta-Ramusino), “spacers” (Briner 2014) and generically as “crRNAs” (Jiang).
- targeting domains are typically 10-30 nucleotides in length, and in certain embodiments are 16-24 nucleotides in length (for instance, 16, 17, 18, 19, 20, 21, 22, 23 or 24 nucleotides in length), and are at or near the 5’ terminus of in the case of a Cas9 gRNA, and at or near the 3’ terminus in the case of a Cpf1 gRNA.
- gRNAs typically (but not necessarily, as discussed below) include a plurality of domains that may influence the formation or activity of gRNA/Cas9 complexes.
- first and secondary complementarity domains of a gRNA interacts with the recognition (REC) lobe of Cas9 and can mediate the formation of Cas9/gRNA complexes (Nishimasu 2014; Nishimasu 2015; both incorporated by reference herein).
- first and/or second complementarity domains may contain one or more poly-A tracts, which can be recognized by RNA polymerases as a termination signal.
- first and second complementarity domains are, therefore, optionally modified to eliminate these tracts and promote the complete in vitro transcription of gRNAs, for instance through the use of A-G swaps as described in Briner 2014, or A-U swaps. These and other similar modifications to the first and second complementarity domains are within the scope of the present disclosure.
- Cas9 gRNAs typically include two or more additional duplexed regions that are involved in nuclease activity in vivo but not necessarily in vitro. (Nishimasu 2015).
- a first stem-loop one near the 3’ portion of the second complementarity domain is referred to variously as the “proximal domain,” (Cotta-Ramusino) “stem loop 1” (Nishimasu 2014 and 2015) and the “nexus” (Briner 2014).
- One or more additional stem loop 35 164554159.2 Attorney Docket No.: 118945.8026.WO00 structures are generally present near the 3’ end of the gRNA, with the number varying by species: S. pyogenes gRNAs typically include two 3’ stem loops (for a total of four stem loop structures including the repeat:anti-repeat duplex), while S. aureus and other species have only one (for a total of three stem loop structures).
- CRISPR CRISPR from Prevotella and Franciscella 1
- a gRNA for use in a Cpf1 genome editing system generally includes a targeting domain and a complementarity domain (alternately referred to as a “handle”).
- the targeting domain is usually present at or near the 3’ end, rather than the 5’ end as described above in connection with Cas9 gRNAs (the handle is at or near the 5’ end of a Cpf1 gRNA).
- the handle is at or near the 5’ end of a Cpf1 gRNA.
- gRNAs can be defined, in broad terms, by their targeting domain sequences, and skilled artisans will appreciate that a given targeting domain sequence can be incorporated in any suitable gRNA, including a unimolecular or chimeric gRNA, or a gRNA that includes one or more chemical modifications and/or sequential modifications (substitutions, additional nucleotides, truncations, etc.). Thus, for economy of presentation in this disclosure, gRNAs may be described solely in terms of their targeting domain sequences. [0100] More generally, skilled artisans will appreciate that some aspects of the present disclosure relate to systems, methods and compositions that can be implemented using multiple RNA-guided nucleases.
- the term gRNA should be understood to encompass any suitable gRNA that can be used with any RNA-guided nuclease, and not only those gRNAs that are compatible with a particular species of Cas9 or Cpf1.
- the term gRNA can, in certain embodiments, include a gRNA for use with any RNA-guided nuclease occurring in a Class 2 CRISPR system, such as a type II or type V or CRISPR system, or an RNA- guided nuclease derived or adapted therefrom.
- gRNA design may involve the use of a software tool to optimize the choice of potential target sequences corresponding to a user’s target sequence, e.g., to minimize total off-target activity across the genome.
- Targeting domain sequences of gRNAs that were designed to target disruption of the CCAAT box target region include, but are not limited to, SEQ ID NO:202.
- gRNAs comprising the sequence set forth in SEQ ID NO:202 may be complexed with a Cpf1 protein to generate alterations at the CCAAT box target region.
- gRNAs comprising any of the Cpf1 gRNAs set forth in Table 14 may be complexed with a Cpf1 protein forming an RNP (“gRNA-Cpf1-RNP”) to generate alterations at the CCAAT box target region.
- the Cpf1 protein may be His-AsCpf1-nNLS (SEQ ID NO: 200) or His-AsCpf1-sNLS- sNLS (SEQ ID NO:201).
- the Cpf1 protein of the gRNA-Cpf1-RNP may comprise a sequence set forth in SEQ ID NOs:200, 201, 205-215, 221, 222-226 (Cpf1 polypeptide sequences).
- the Cpf1 protein may be encoded by a sequence set forth in SEQ ID NOs:216-218 (Cpf1 polynucleotide sequences).
- gRNA modifications [0103] The activity, stability, or other characteristics of gRNAs can be altered through the incorporation of certain modifications. As one example, transiently expressed or delivered nucleic acids can be prone to degradation by, e.g., cellular nucleases. Accordingly, the gRNAs described herein can contain one or more modified nucleosides or nucleotides which introduce stability toward nucleases. While not wishing to be bound by theory it is also believed that certain modified gRNAs described herein can exhibit a reduced innate immune response when introduced into cells.
- Certain exemplary modifications discussed in this section can be included at any position within a gRNA sequence including, without limitation at or near the 5’ end (e.g., within 1-10, 1-5, or 1-2 nucleotides of the 5’ end) and/or at or near the 3’ end (e.g., within 1-10, 1-5, or 1-2 nucleotides of the 3’ end).
- modifications are positioned within functional motifs, such as the repeat- 37 164554159.2 Attorney Docket No.: 118945.8026.WO00 anti-repeat duplex of a Cas9 gRNA, a stem loop structure of a Cas9 or Cpf1 gRNA, and/or a targeting domain of a gRNA.
- the 5’ end of a gRNA can include a eukaryotic mRNA cap structure or cap analog (e.g., a G(5’)ppp(5’)G cap analog, a m7G(5’)ppp(5’)G cap analog, or a 3’-O-Me- m7G(5’)ppp(5’)G anti reverse cap analog (ARCA)), as shown below:
- the cap or cap can or in vitro of the gRNA.
- the 5’ end of the gRNA can lack a 5’ triphosphate group.
- in vitro transcribed gRNAs can be phosphatase-treated (e.g., using calf intestinal alkaline phosphatase) to remove a 5’ triphosphate group.
- Another common modification involves the addition, at the 3’ end of a gRNA, of a plurality (e.g., 1-10, 10-20, or 25-200) of adenine (A) residues referred to as a polyA tract.
- the polyA tract can be added to a gRNA during chemical synthesis, following in vitro transcription using a polyadenosine polymerase (e.g., E.
- RNAs can be modified at a 3’ terminal U ribose.
- the two terminal hydroxyl groups of the U ribose can be oxidized to aldehyde groups and a concomitant opening of the ribose ring to afford a modified nucleoside as shown below: 164554159.2 Attorney Docket No.: 118945.8026.WO00 wherein “U” can be an unmodified or modified uridine.
- the 3’ terminal U ribose can be modified with a 2’3’ cyclic phosphate as shown below: wherein “U” can be an unmodified or [0111]
- Guide RNAs can contain 3’ nucleotides which can be stabilized against degradation, e.g., by incorporating one or more of the modified nucleotides described herein.
- uridines can be replaced with modified uridines, e.g., 5-(2-amino)propyl uridine, and 5-bromo uridine, or with any of the modified uridines described herein; adenosines and guanosines can be replaced with modified adenosines and guanosines, e.g., with modifications at the 8-position, e.g., 8-bromo guanosine, or with any of the modified adenosines or guanosines described herein.
- sugar-modified ribonucleotides can be incorporated into the gRNA, e.g., wherein the 2’ OH-group is replaced by a group selected from H, -OR, -R (wherein R can be, e.g., alkyl, cycloalkyl, aryl, aralkyl, heteroaryl or sugar), halo, -SH, -SR (wherein R can be, e.g., alkyl, cycloalkyl, aryl, aralkyl, heteroaryl or sugar), amino (wherein amino can be, e.g., NH2; alkylamino, dialkylamino, heterocyclyl, arylamino, diarylamino, heteroarylamino, diheteroarylamino, or amino acid); or cyano (-CN).
- R can be, e.g., alkyl, cycloalkyl, aryl, aralkyl, heteroaryl
- the phosphate backbone can be modified as described herein, e.g., with a phosphorothioate (PhTx) group.
- one or more of the nucleotides of the gRNA can each independently be a modified or unmodified nucleotide including, but not limited to 2’-sugar modified, such as, 2’-O-methyl, 2’-O-methoxyethyl, or 2’-Fluoro modified including, e.g., 2’-F or 2’-O-methyl, adenosine (A), 2’-F or 2’-O-methyl, cytidine (C), 2’-F or 2’-O- methyl, uridine (U), 2’-F or 2’-O-methyl, thymidine (T), 2’-F or 2’-O-methyl, guanosine (G), 2’-O- methoxyethyl-5-methyluridine (Teo
- Guide RNAs can also include “locked” nucleic acids (LNA) in which the 2’ OH-group can be connected, e.g., by a C1-6 alkylene or C1-6 heteroalkylene bridge, to the 4’ carbon of the same ribose sugar.
- LNA locked nucleic acids
- Any suitable moiety can be used to provide such bridges, include without limitation methylene, propylene, ether, or amino bridges; O-amino (wherein amino can be, e.g., NH 2 ; alkylamino, dialkylamino, heterocyclyl, arylamino, diarylamino, heteroarylamino, or diheteroarylamino, ethylenediamine, or polyamino) and aminoalkoxy or O(CH2)n-amino (wherein 39 164554159.2 Attorney Docket No.: 118945.8026.WO00 amino can be, e.g., NH 2 ; alkylamino, dialkylamino, heterocyclyl, arylamino, diarylamino, heteroarylamino, or diheteroarylamino, ethylenediamine, or polyamino).
- a gRNA can include a modified nucleotide which is multicyclic (e.g., tricyclo; and “unlocked” forms, such as glycol nucleic acid (GNA) (e.g., R-GNA or S-GNA, where ribose is replaced by glycol units attached to phosphodiester bonds), or threose nucleic acid (TNA, where ribose is replaced with ⁇ -L-threofuranosyl-(3’ ⁇ 2’)).
- GAA glycol nucleic acid
- TAA threose nucleic acid
- gRNAs include the sugar group ribose, which is a 5-membered ring having an oxygen.
- Exemplary modified gRNAs can include, without limitation, replacement of the oxygen in ribose (e.g., with sulfur (S), selenium (Se), or alkylene, such as, e.g., methylene or ethylene); addition of a double bond (e.g., to replace ribose with cyclopentenyl or cyclohexenyl); ring contraction of ribose (e.g., to form a 4-membered ring of cyclobutane or oxetane); ring expansion of ribose (e.g., to form a 6- or 7-membered ring having an additional carbon or heteroatom, such as for example, anhydrohexitol, altritol, mannitol, cyclohexanyl, cyclohexenyl, and morpholino that also has a phosphoramidate backbone).
- replacement of the oxygen in ribose e.g., with
- a gRNA comprises a 4’-S, 4’-Se or a 4’-C-aminomethyl-2’-O-Me modification.
- deaza nucleotides e.g., 7-deaza-adenosine
- O- and N-alkylated nucleotides e.g., N6-methyl adenosine, can be incorporated into the gRNA.
- gRNAs as used herein may be modified or unmodified gRNAs.
- a gRNA may include one or more modifications.
- the one or more modifications may include a phosphorothioate linkage modification, a phosphorodithioate (PS2) linkage modification, a 2’-O-methyl modification, or combinations thereof.
- the one or more modifications may be at the 5’ end of the gRNA, at the 3’ end of the gRNA, or combinations thereof.
- a gRNA modification may comprise one or more phosphorodithioate (PS2) linkage modifications.
- PS2 phosphorodithioate
- a gRNA used herein includes one or more or a stretch of deoxyribonucleic acid (DNA) bases, also referred to herein as a “DNA extension.”
- a gRNA used herein includes a DNA extension at the 5’ end of the gRNA, the 3’ end of the gRNA, or a combination thereof.
- the DNA extension may be 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 40 164554159.2 Attorney Docket No.: 118945.8026.WO00 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, 99, or 100 DNA bases long.
- the DNA extension may be 1, 2, 3, 4, 5, 10, 15, 20, or 25 DNA bases long.
- the DNA extension may include one or more DNA bases selected from adenine (A), guanine (G), cytosine (C), or thymine (T).
- the DNA extension includes the same DNA bases.
- the DNA extension may include a stretch of adenine (A) bases.
- the DNA extension may include a stretch of thymine (T) bases.
- the DNA extension includes a combination of different DNA bases.
- a DNA extension may comprise a sequence set forth in Table 13.
- a DNA extension may comprise a sequence set forth in SEQ ID NOs:304-319.
- a gRNA used herein includes a DNA extension as well as one or more phosphorothioate linkage modifications, one or more phosphorodithioate (PS2) linkage modifications, one or more 2’-O-methyl modifications, or combinations thereof.
- the one or more modifications may be at the 5’ end of the gRNA, at the 3’ end of the gRNA, or combinations thereof.
- a gRNA including a DNA extension may comprise a sequence set forth in Table 13 that includes a DNA extension.
- a gRNA including a DNA extension may comprise the sequence set forth in SEQ ID NO:237.
- a gRNA including a DNA extension may comprise a sequence selected from the group consisting of SEQ ID NOs:232- 246, 253, 254, 260, 261, 264, 267-270, 272-273, 275-276, 278-279, 284-288, and 292.
- any DNA extension may be used herein, so long as it does not hybridize to the target nucleic acid being targeted by the gRNA and it also exhibits an increase in editing at the target nucleic acid site relative to a gRNA which does not include such a DNA extension.
- a gRNA used herein includes one or more or a stretch of ribonucleic acid (RNA) bases, also referred to herein as an “RNA extension.”
- RNA extension also referred to herein as an “RNA extension.”
- a gRNA used herein includes an RNA extension at the 5’ end of the gRNA, the 3’ end of the gRNA, or a combination thereof.
- the RNA extension may be 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, 99, or 100 RNA bases long.
- the RNA extension may be 1, 2, 3, 4, 5, 10, 15, 20, or 25 RNA bases long.
- the RNA extension may include one or more RNA bases selected from adenine (rA), guanine (rG), cytosine (rC), or uracil (rU), in which the “r” represents RNA, 2’-hydroxy.
- rA adenine
- rG guanine
- rC cytosine
- uracil uracil
- the RNA extension includes the same RNA bases.
- the RNA extension may include a stretch of adenine (rA) bases.
- the RNA extension includes a combination of different RNA bases.
- an RNA extension may comprise a sequence set forth in Table 14.
- an RNA extension may comprise a sequence set forth in 328-334.
- a gRNA used herein includes an RNA extension as well as one or more phosphorothioate linkage modifications, one or more phosphorodithioate (PS2) linkage modifications, one or more 2’-O-methyl modifications, or combinations thereof.
- the one or more modifications may be at the 5’ end of the gRNA, at the 3’ end of the gRNA, or combinations thereof.
- a gRNA including a RNA extension may comprise a sequence set forth in Table 13 that includes an RNA extension.
- gRNAs including an RNA extension at the 5’ end of the gRNA may comprise a sequence selected from the group consisting of SEQ ID NOs:228-231, 289-291.
- gRNAs including an RNA extension at the 3’ end of the gRNA may comprise a sequence selected from the group consisting of SEQ ID NOs:256-261, 265, 267, 284-286.
- gRNAs used herein may also include an RNA extension and a DNA extension. In certain embodiments, the RNA extension and DNA extension may both be at the 5’ end of the gRNA, the 3’ end of the gRNA, or a combination thereof.
- the RNA extension is at the 5’ end of the gRNA and the DNA extension is at the 3’ end of the gRNA. In certain embodiments, the RNA extension is at the 3’ end of the gRNA and the DNA extension is at the 5’ end of the gRNA.
- a gRNA which includes both a phosphorothioate modification at the 3’ end as well as a DNA extension at the 5’ end is complexed with a RNA-guided nuclease, e.g., Cpf1, to form an RNP, which is then employed to edit a hematopoietic stem cell (HSC) or a CD34+ cell ex vivo (i.e., outside the body of a subject from whom such a cell is derived), at the HBG locus.
- HSC hematopoietic stem cell
- CD34+ cell ex vivo i.e., outside the body of a subject from whom such a cell is derived
- RNA-guided nucleases include, but are not limited to, naturally-occurring Class 2 CRISPR nucleases such as Cpf1, and Cas9, as well as other nucleases derived or obtained therefrom. [0125] In functional terms, RNA-guided nucleases are defined as those nucleases that: (a) interact with (e.g.
- RNA-guided nucleases can be defined, in broad terms, by their PAM specificity and cleavage activity, even though variations may exist between individual RNA-guided nucleases that share the same PAM specificity or cleavage activity.
- RNA-guided nuclease should be understood as a generic term, and not limited to any particular type (e.g. Cas9 vs. Cpf1), species (e.g. S. pyogenes vs. S. aureus) or variation (e.g. full-length vs. truncated or split; naturally-occurring PAM specificity vs. engineered PAM specificity, etc.) of RNA-guided nuclease.
- any suitable RNA-guided nuclease having a certain PAM specificity and/or cleavage activity.
- the term RNA-guided nuclease should be understood as a generic term, and not limited to any particular type (e.g. Cas9 vs. Cpf1), species (e.g. S. pyogenes vs. S. aureus) or variation (e.g. full-length vs. truncated or split; naturally-occurring P
- the RNA-guided nuclease may be Cas- ⁇ (Pausch 2020).
- Various RNA-guided nucleases may require different sequential relationships between PAMs and protospacers.
- Cas9s recognize PAM sequences that are 3’ of the protospacer.
- Cpf1 on the other hand, generally recognizes PAM sequences that are 5’ of the protospacer.
- RNA- guided nucleases can also recognize specific PAM sequences. S.
- aureus Cas9 for instance, recognizes a PAM sequence of NNGRRT or NNGRRV, wherein the N residues are immediately 3’ of the region recognized by the gRNA targeting domain.
- S. pyogenes Cas9 recognizes NGG PAM sequences.
- F. novicida Cpf1 recognizes a TTN PAM sequence.
- PAM sequences have been identified for a variety of RNA-guided nucleases, and a strategy for identifying novel PAM sequences has been described by Shmakov 2015.
- engineered RNA-guided nucleases can have PAM specificities that differ from the PAM specificities of reference molecules (for instance, in the case of an engineered RNA-guided nuclease, the reference molecule may be the naturally occurring variant from which the RNA-guided nuclease is derived, or the naturally occurring variant having the greatest amino acid sequence homology to the engineered RNA-guided nuclease).
- PAMs that may be used according to the embodiments herein include, without limitation, the sequences set forth in SEQ ID NOs:191-197.
- RNA-guided nucleases can be characterized by their DNA cleavage activity: naturally-occurring RNA-guided nucleases typically form DSBs in target nucleic acids, but engineered variants have been produced that generate only SSBs (discussed above and in Ran & Hsu 2013, incorporated by reference herein), or that do not cut at all.
- Cas9 [0129] Crystal structures have been determined for S. pyogenes Cas9 (Jinek 2014), and for S. aureus Cas9 in complex with a unimolecular guide RNA and a target DNA (Nishimasu 2014; Anders 2014; and Nishimasu 2015).
- a naturally occurring Cas9 protein comprises two lobes: a recognition (REC) lobe and a nuclease (NUC) lobe; each of which comprise particular structural and/or functional domains.
- the REC lobe comprises an arginine-rich bridge helix (BH) domain, and at least one REC domain (e.g. a REC1 domain and, optionally, a REC2 domain).
- the REC lobe does not share structural similarity with other known proteins, indicating that it is a unique functional domain.
- the NUC lobe comprises a RuvC domain, an HNH domain, and a PAM-interacting (PI) domain.
- the RuvC domain shares structural similarity to retroviral integrase superfamily members and cleaves the non-complementary (i.e. bottom) strand of the target nucleic acid.
- the HNH domain may be formed from two or more split RuvC motifs (such as RuvC I, RuvCII, and RuvCIII in S. pyogenes and S. aureus).
- the HNH domain meanwhile, is structurally similar to HNN endonuclease motifs, and cleaves the complementary (i.e. top) strand of the target nucleic acid.
- the PI domain as its name suggests, contributes to PAM specificity.
- Examples of Cas9 RuvC-like and Cas9 HNH-like domain polypeptide sequences are set forth in SEQ ID NOs:7-15, 44-115 (RuvC-like domains) and SEQ ID NOs:16-20, 116-190 (HNH-like domains).
- Cas9 While certain functions of Cas9 are linked to (but not necessarily fully determined by) the specific domains set forth above, these and other functions may be mediated or influenced by other Cas9 domains, or by multiple domains on either lobe.
- the repeat:antirepeat duplex of the gRNA falls into a groove between the REC and NUC lobes, and nucleotides in the duplex interact with amino acids in the BH, PI, and REC domains.
- Cpf1 [0133] The crystal structure of Acidaminococcus sp. Cpf1 in complex with crRNA and a double- stranded (ds) DNA target including a TTTN PAM sequence has been solved by Yamano 2016 (incorporated by reference herein).
- Cpf1 like Cas9, has two lobes: a REC (recognition) lobe, and a NUC (nuclease) lobe.
- the REC lobe includes REC1 and REC2 domains, which lack similarity to any known protein structures.
- the NUC lobe meanwhile, includes three RuvC domains (RuvC-I, -II and -III) and a BH domain.
- the Cpf1 REC lobe lacks an HNH domain
- 44 164554159.2 Attorney Docket No.: 118945.8026.WO00 includes other domains that also lack similarity to known protein structures: a structurally unique PI domain, three Wedge (WED) domains (WED-I, -II and -III), and a nuclease (Nuc) domain.
- WED Wedge
- Nuc nuclease
- a Cpf1 protein may include one or more modifications.
- the modifications may be, without limitation, one or more mutations in a Cpf1 nucleotide sequence or Cpf1 amino acid sequence, one or more additional sequences such as a His tag or a nuclear localization signal (NLS), or a combination thereof.
- the Cpf1 protein may be derived from a Cpf1 protein selected from the group consisting of Acidaminococcus sp. strain BV3L6 Cpf1 protein (AsCpf1), Lachnospiraceae bacterium ND2006 Cpf1 protein (LbCpf1), and Lachnospiraceae bacterium MA2020 (Lb2Cpf1).
- the Cpf1 protein may comprise a sequence selected from the group consisting of SEQ ID NOs:213-215, having the codon-optimized nucleic acid sequences of SEQ ID NOs:216-218, respectively.
- the Cpf1 protein may comprise a nuclear localization signal (NLS).
- NLS sequences useful in connection with the methods and compositions disclosed herein will comprise an amino acid sequence capable of facilitating protein import into the cell nucleus.
- NLS sequences useful in connection with the methods and compositions disclosed herein are known in the art. Examples of such NLS sequences include the nucleoplasmin NLS having the amino acid sequence: KRPAATKKAGQAKKKK (SEQ ID NO:203) and the simian virus 40 “SV40” NLS having the amino acid sequence PKKKRKV (SEQ ID NO:204).
- the NLS sequence of the Cpf1 protein is positioned at or near the C- terminus of the Cpf1 protein sequence.
- the Cpf1 protein can be selected from the following: His-AsCpf1-nNLS (SEQ ID NO:200); His-AsCpf1-sNLS (SEQ ID NO:205) and His-AsCpf1-sNLS-sNLS (SEQ ID NO:201), where “His” refers to a six-histidine purification sequence, “AsCpf1” refers to the Acidaminococcus sp.
- nNLS refers to the nucleoplasmin NLS
- sNLS refers to the SV40 NLS. Additional permutations of the identity and C-terminal positions of NLS sequences, e.g., appending two or more nNLS sequences or combinations of nNLS and sNLS sequences (or other NLS sequences), as well as sequences with and 45 164554159.2 Attorney Docket No.: 118945.8026.WO00 without purification sequences, e.g., six-histidine sequences, are within the scope of the instantly disclosed subject matter.
- the NLS sequence of the Cpf1 protein may be positioned at or near the N-terminus of the Cpf1 protein sequence.
- the Cpf1 protein may be selected from the following: His-sNLS-AsCpf1 (SEQ ID NO:206), His-sNLS-sNLS-AsCpf1 (SEQ ID NO:207), and sNLS-sNLS-AsCpf1 (SEQ ID NO:208).
- the Cpf1 protein may comprise NLS sequences positioned at or near both the N-terminus and C-terminus of the Cpf1 protein sequence.
- the Cpf1 protein may be selected from the following: His-sNLS-AsCpf1-sNLS (SEQ ID NO:209) and His-sNLS-sNLS-AsCpf1-sNLS-sNLS (SEQ ID NO:210).
- the Cpf1 protein may comprise an alteration (e.g., a deletion or substitution) at one or more cysteine residues of the Cpf1 protein sequence.
- Cpf1 protein may comprise an alteration at a position selected from the group consisting of: C65, C205, C334, C379, C608, C674, C1025, and C1248.
- the Cpf1 protein may comprise a substitution of one or more cysteine residues for a serine or alanine.
- the Cpf1 protein may comprise an alteration selected from the group consisting of: C65S, C205S, C334S, C379S, C608S, C674S, C1025S, and C1248S.
- the Cpf1 protein may comprise an alteration selected from the group consisting of: C65A, C205A, C334A, C379A, C608A, C674A, C1025A, and C1248A.
- the Cpf1 protein may comprise alterations at positions C334 and C674 or C334, C379, and C674.
- the Cpf1 protein may comprise the following alterations: C334S and C674S, or C334S, C379S, and C674S.
- the Cpf1 protein may comprise the following alterations: C334A and C674A, or C334A, C379A, and C674A.
- the Cpf1 protein may comprise both one or more cysteine residue alteration as well as the introduction of one or more NLS sequences, e.g., His-AsCpf1-nNLS Cys-less (SEQ ID NO:211) or His-AsCpf1-nNLS Cys-low (SEQ ID NO:212).
- the Cpf1 protein comprising a deletion or substitution in one or more cysteine residues exhibits reduced aggregation.
- other Cpf1 proteins known in the art may be used with the methods and systems described herein.
- the Cpf1 may be Cpf1 containing the mutation S542R/K548V/N552R (“Cpf1 RVR”). Cpf1 RVR has been shown to cleave target sites with a TATV PAM. In certain embodiments, the Cpf1 may be Cpf1 containing the mutation S542R/K607R (“Cpf1 RR”). Cpf1 RR has been shown to cleave target sites with a TYCV/CCCC PAM. [0143] In some embodiments, a Cpf1 is used herein, wherein the Cpf1 comprises mutations at one or more residues of AsCpf1 (Acidaminococcus sp.
- BV3L6 selected from the group consisting of 11, 12, 13, 14, 15, 16, 17, 34, 36, 39, 40, 43, 46, 47, 50, 54, 57, 58, 111, 126, 127, 128, 129, 130, 131, 132, 133, 134, 135, 136, 157, 158, 159, 160, 161, 162, 163, 164, 165, 166, 167, 168, 169, 170, 171, 172, 173, 174, 175, 176, 177, 178, 532, 533, 534, 535, 536, 537, 538, 539, 540, 541, 542, 543, 544, 545, 546, 547, 548, 549, 550, 551, 552, 553, 554, 555, 556, 565, 566, 567, 568, 569, 570, 571, 572, 573, 574, 575, 592, 593, 594, 595, 596, 597
- a Cpf1 as used herein may include any of the Cpf1 proteins described in International Publication Number WO 2017/184768 A1 by Zhang et al. (“ ⁇ 768 Publication”), which is incorporated by reference herein.
- a Cpf1 protein also referred to herein as a Cpf1 molecule used herein may comprise any of the sequences set forth in SEQ ID NOs:200, 201, 205-215, 221, 222-226, 280- 283, 293-295 (Cpf1 polypeptide sequences).
- a Cpf1 protein may be encoded by a sequence comprising any of SEQ ID NOs:216-218, 296-303 (Cpf1 polynucleotide sequences).
- Table 11 sets forth exemplary Cpf1 amino acid and nucleotide sequences. These sequences are set forth in Fig. 10, which details the positions of six-histidine sequences (underlined letters) and NLS sequences (bolded letters).
- NLS sequences e.g., appending two or more nNLS sequences or combinations of nNLS and sNLS sequences (or other NLS sequences) to either the N-terminal/C-terminal positions, as well as sequences with and without purification sequences, e.g., six-histidine sequences, are within the scope of the instantly disclosed subject matter.
- any of the Cpf1 proteins disclosed herein may be complexed with one or more gRNA comprising the targeting domain set forth in SEQ ID NOs: 202 and/or 327 to alter a CCAAT box target region.
- any of the Cpf1 proteins disclosed herein may be complexed with one or more gRNA comprising a sequence set forth in Table 14.
- the Cpf1 protein may be His-AsCpf1-nNLS (comprising SEQ ID NO:200) or His- AsCpf1-sNLS-sNLS (comprising SEQ ID NO:201).
- a Cpf1 protein used herein may comprise any of the sequences set forth in SEQ ID NOs:200, 201, 205-215, 221, 222-226, 280-283, 293-295 (Cpf1 polypeptide sequences).
- a Cpf1 protein used herein may be encoded by any of the sequences set forth in SEQ ID NOs:216-218, 296-303 (Cpf1 polynucleotide sequences).
- the Cpf1 protein may comprise the sequence set forth in SEQ ID NO:283. [0147]
- the Cpf1 protein may include a Cpf1 described in Kleinstiver 2019.
- the Cpf1 protein may be enAsCas12a.
- the Cpf1 protein may cleave target sites with a TTTV PAM.
- the Cpf1 protein may cleave target sites with a NWYN PAM.
- inactivation of a RuvC domain of a Cas9 will result in a nickase that cleaves the complementary or top strand as shown below (where C denotes the site of cleavage).
- C denotes the site of cleavage.
- inactivation of a Cas9 HNH domain results in a nickase that cleaves the bottom or non-complementary strand.
- Modifications of PAM specificity relative to naturally occurring Cas9 reference molecules has been described by Kleinstiver et al. for both S. pyogenes (Kleinstiver 2015a) and S. aureus 48 164554159.2 Attorney Docket No.: 118945.8026.WO00 (Kleinstiver 2015b).
- RNA-guided nucleases have been split into two or more parts, as described by Zetsche 2015 and Fine 2015 (both incorporated by reference herein). [0153] RNA-guided nucleases can be, in certain embodiments, size-optimized or truncated, for instance via one or more deletions that reduce the size of the nuclease while still retaining gRNA association, target and PAM recognition, and cleavage activities.
- RNA guided nucleases are bound, covalently or non-covalently, to another polypeptide, nucleotide, or other structure, optionally by means of a linker. Exemplary bound nucleases and linkers are described by Guilinger 2014, incorporated by reference herein for all purposes.
- RNA-guided nucleases also optionally include a tag, such as, but not limited to, a nuclear localization signal to facilitate movement of RNA-guided nuclease protein into the nucleus.
- the RNA-guided nuclease can incorporate C- and/or N-terminal nuclear localization signals. Nuclear localization sequences are known in the art and are described in Maeder and elsewhere.
- nucleic acids encoding RNA-guided nucleases have been described previously (see, e.g., Cong 2013; Wang 2013; Mali 2013; Jinek 2012).
- a nucleic acid encoding an RNA-guided nuclease can be a synthetic nucleic acid sequence.
- the synthetic nucleic acid molecule can be chemically modified.
- an mRNA encoding an RNA-guided nuclease will have one or more (e.g., all) of the following properties: it can be capped; polyadenylated; and substituted with 5-methylcytidine and/or pseudouridine.
- Synthetic nucleic acid sequences can also be codon optimized, e.g., at least one non-common codon or less-common codon has been replaced by a common codon.
- the synthetic nucleic acid can direct the synthesis of an optimized messenger mRNA, e.g., optimized for expression in a mammalian expression system, e.g., described herein. Examples of codon optimized Cas9 coding sequences are presented in Cotta-Ramusino.
- a nucleic acid encoding an RNA-guided nuclease may comprise a nuclear localization sequence (NLS). Nuclear localization sequences are known in the art.
- the subject may be suffering from a ⁇ - hemoglobinopathy.
- the ⁇ -hemoglobinopathy may be sickle cell disease (SCD) or ⁇ -Thal.
- the method may comprise administering to the subject a population of modified cells comprising a plurality of modified CD34+ or hematopoietic stem cells comprising an indel in an HBG promoter, thereby inducing expression of HbF in the population of cells (e.g., HSPCs or RBCs).
- HbF as a percentage of total hemoglobin (% HbF) in the subject may be about 5%, 10%, 15%, 20%, 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, or 80% HbF.
- the % HbF in the subject may be from about 10% to about 30%, from about 20% to about 40%, from about 30% to about 50%, from about 40% to about 60%, from about 50% to about 70%, from about 60% to about 80%, from about 10% to about 20%, from about 20% to about 30%, from about 30% to about 40%, from about 40% to about 50%, from about 50% to about 60%, from about 60% to about 70%, from about 70% to about 80% HbF, or a range defined by any of the two preceding values.
- a concentration of total hemoglobin in the subject may be about 10 g/dL, 11 g/dL, 12 g/dL, 13 g/dL, 14 g/dL, 15 g/dL, 16 g/dL, 17 g/dL, 18 g/dL, 19 g/dL, or 20 g/dL.
- a concentration of total hemoglobin in the subject may be from about 10.0 to about 20.0 g/dL, from about 13.6 to about 18.0 g/dL (for males), from about 12.0 to about 16.0 g/dL (for females), or a range defined by any of the two preceding values.
- a percentage of F-cells among circulating RBCs in the subject may be about 5%, 10%, 15%, 20%, 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, 100% F-cells.
- a percentage of F-cells among circulating RBCs in the subject may be from about 50% to about 100%, from about 60% to about 100%, from about 70% to about 100%, from about 80% to about 100%, 50 164554159.2 Attorney Docket No.: 118945.8026.WO00 from about 50% to about 60%, from about 60% to about 70%, from about 70% to about 80%, from about 80% to about 90%, from about 90% to about 100%, or a range defined by any of the two preceding values.
- a Hemoglobin F concentration in the subject may be about 1 g/dL, 2 g/dL, 3 g/dL, 4 g/dL, 5 g/dL, 6 g/dL, 7 g/dL, 8 g/dL, 9 g/dL, 10 g/dL, 11 g/dL, 12 g/dL, 13 g/dL, 14 g/dL, 15 g/dL, 16 g/dL, 17 g/dL, 18 g/dL, 19 g/dL, or 20 g/dL.
- a Hemoglobin F concentration in the subject may be from about 1g/dL to about 20 g/dL, or a range defined by any of the two preceding values.
- a mean corpuscular HbF (pg/RBC) in the subject may be about 1.0 pg/RBC, 2.0 pg/RBC, 3.0 pg/RBC, 4.0 pg/RBC, 5.0 pg/RBC, 6.0 pg/RBC, 7.0 pg/RBC, 8.0 pg/RBC, 9.0 pg/RBC, 10.0 pg/RBC, 11.0 pg/RBC, 12.0 pg/RBC, 13.0 pg/RBC, 14.0 pg/RBC 15.0 pg/RBC, 16.0 pg/RBC, 17.0 pg/RBC, 18.0 pg/RBC, 19.0 pg/RBC, 20.0 pg/RBC, 2
- a mean corpuscular HbF (pg/RBC) in the subject may be from about 1.0 pg/RBC to about 5.00 pg/RBC, from about 5.0 pg/RBC to about 10.00 pg/RBC, from about 10.0 pg/RBC to about 15.00 pg/RBC, from about 15.0 pg/RBC to about 20.00 pg/RBC, from about 20.0 pg/RBC to about 25.00 pg/RBC, from about 25.0 pg/RBC to about 30.00 pg/RBC, or a range defined by any of the two preceding values.
- a mean corpuscular HbF (pg/RBC) in the subject may be ⁇ 10.0 pg/RBC.
- a mean proportion of HbF as a percentage of total hemoglobin (Hb) in the subject is about 5%, 10%, 15%, 20%, 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%.
- a mean proportion of HbS as a percentage of total hemoglobin (Hb) in the subject is about 5%, 10%, 15%, 20%, 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%.
- a mean proportion of HbA as a percentage of total hemoglobin (Hb) in the subject is about 1%, 2%, 3%, 4%, 5%, 10%, 15%, 20%, 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%.
- a mean proportion of HbA2 as a percentage of total hemoglobin (Hb) in the subject is about 1%, 2%, 3%, 4%, 5%.
- a lactate dehydrogenase (U/L) in the subject is about 110 U/L, 120 U/L, 130 U/L, 140 U/L, 150 U/L, 160 U/L, 170 U/L, 180 U/L, 190 U/L, 200 U/L, 210 U/L, 220 U/L, or 230 U/L.
- a lactate dehydrogenase (U/L) in the subject is from about 110 U/L to about 230 U/L, or a range defined by any of the two preceding values.
- an indirect bilirubin ( ⁇ mol/L) in the subject is about 0.0 ⁇ mol/L, 1 ⁇ mol/L, 2 ⁇ mol/L, 3 ⁇ mol/L, 4 ⁇ mol/L, 5 ⁇ mol/L, 6 ⁇ mol/L, 7 ⁇ mol/L, 8 ⁇ mol/L, 9 ⁇ mol/L, 10 ⁇ mol/L, 11 ⁇ mol/L, 12 ⁇ mol/L, 13 ⁇ mol/L, 14 ⁇ mol/L, 15 ⁇ mol/L, 16 ⁇ mol/L, or 17 ⁇ mol/L.
- an indirect bilirubin ( ⁇ mol/L) in the subject is from about 0.0 ⁇ mol/L to about 16.6 ⁇ mol/L, or a range defined by the two preceding values.
- a haptoglobin (g/L) in the subject is about 0.3 g/L, 0.4 g/L, 0.5 g/L, 0.6 g/L, 0.7 g/L, 0.8 g/L, 0.9 g/L, 1.0 g/L, 1.1 g/L, 1.2 g/L, 1.3 g/L, 1.4 g/L, 1.5 g/L, 1.6 g/L, 1.7 g/L, 1.8 g/L, 1.9 g/L, or 2.0 g/L.
- a haptoglobin (g/L) in the subject is from about 0.3 g/L to about 2.0 g/L or a range 51 164554159.2 Attorney Docket No.: 118945.8026.WO00 defined by the two preceding values.
- a reticulocyte count (%) in the subject is about 0.3%, 0.4%, 0.5%, 0.6%, 0.7%, 0.8%, 0.9%, 1.0%, 1.1%, 1.2%, 1.3%, 1.4%, 1.5%, 1.6%, 1.7%, 1.8%, 1.9%, 2.0%, 2.1%, 2.2%, or 2.3%.
- a reticulocyte count (%) in the subject is from about 0.3 % to about 2.3% or a range defined by the two preceding values.
- normalization of total hemoglobin in the subject occurs by at least 1 month, 1.5 months, 2 months, 3 months, 4 months, 5 months after administering the population of modified cells to the subject.
- the subject may undergo myeloablative conditioning with busulfan prior to administering the population of modified cells.
- administering the population of modified cells may comprise a single infusion of the modified population of cells.
- the population of modified cells may be about ⁇ 1 x 10 6 cells/kg, ⁇ 2 x 10 6 cells/kg, ⁇ 3 x 10 6 cells/kg, ⁇ 4 x 10 6 cells/kg, ⁇ 5 x 10 6 cells/kg, ⁇ 6 x 10 6 cells/kg, ⁇ 7 x 10 6 cells/kg, ⁇ 8 x 10 6 cells/kg, ⁇ 9 x 10 6 cells/kg, ⁇ 10 x 10 6 cells/kg, ⁇ 11 x 10 6 cells/kg, ⁇ 12 x 10 6 cells/kg, ⁇ 13 x 10 6 cells/kg, ⁇ 14 x 10 6 cells/kg, ⁇ 15 x 10 6 cells/kg, ⁇ 16 x 10 6 cells/kg, ⁇ 17 x 10 6 cells/kg, ⁇ 18 x 10 6 cells/kg, ⁇ 19 x 10 6 cells/kg, ⁇ 20 x 10 6 cells/kg, ⁇ 21 x 10 6 cells/kg, ⁇ 22 x 10 6 cells/kg,
- the % HbF may be determined 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30 months after administering the population of modified cells to the subject.
- the concentration of total hemoglobin may be determined 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30 months after administering the population of modified cells to the subject.
- the Hemoglobin F concentration may be determined 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30 months after administering the population of modified cells to the subject.
- the percentage of F-cells may be determined 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30 months after administering the population of modified cells to the subject.
- the mean corpuscular HbF (pg/RBC) may be determined 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30 months after administering the population of modified cells to the subject.
- the mean proportion of HbF as a percentage of total hemoglobin is determined 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30 months after administering the population of modified cells to the subject.
- the mean proportion of HbS as a percentage of total hemoglobin is determined 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30 months after administering the 52 164554159.2 Attorney Docket No.: 118945.8026.WO00 population of modified cells to the subject.
- the mean proportion of HbA as a percentage of total hemoglobin is determined 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30 months after administering the population of modified cells to the subject.
- the mean proportion of HbA2 as a percentage of total hemoglobin is determined 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30 months after administering the population of modified cells to the subject.
- the lactate dehydrogenase (U/L) is determined 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30 months after administering the population of modified cells to the subject.
- the indirect bilirubin is determined 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30 months after administering the population of modified cells to the subject.
- the haptoglobin is determined 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30 months after administering the population of modified cells to the subject.
- the reticulocyte count (%) is determined 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30 months after administering the population of modified cells to the subject.
- an RNP complex comprising a guide RNA (gRNA) and a Cpf1 protein may be delivered to a population of unmodified cells comprising a plurality of unmodified CD34+ or hematopoietic stem cells from the subject to generate the population of modified cells.
- the gRNA may comprise a targeting domain comprising SEQ ID NO:320.
- the gRNA may comprise a 5’ end and a 3’ end, a DNA extension at the 5’ end.
- the gRNA may comprise a 2’-O-methyl, phosphorothioate modification, or both at the 3’ end.
- the DNA extension may comprise a sequence selected from the group consisting of SEQ ID NOs:304-319. In certain embodiments, the DNA extension may comprise a sequence selected from the group consisting of SEQ ID NOs:304-319. In certain embodiments, the gRNA may comprise SEQ ID NO:237. In certain embodiments, the Cpf1 protein may comprise a sequence selected from the group consisting of SEQ ID NO:200, 201, 205-215, 221, 222-226, 280-283, 293- 295. In certain embodiments, the Cpf1 protein may comprise SEQ ID NO:283.
- the Cpf1 protein may be encoded by a sequence comprising a sequence selected from the group consisting of SEQ ID NOs:216-218, 296-303. In certain embodiments, the Cpf1 protein may be encoded by a sequence comprising SEQ ID NO:300.
- the indel in the HBG gene promoter may be in a CCAAT box target region.
- the subject has a single homozygous mutation in the HBB gene, c.17A>T (HbS mutation). In certain embodiments, the subject may be suffering from severe sickle cell disease.
- kits for producing a modified population of peripheral blood nucleated cells in a subject comprising: administering to the subject a 53 164554159.2 Attorney Docket No.: 118945.8026.WO00 population of modified cells comprising a plurality of modified CD34+ or hematopoietic stem cells comprising an indel in an HBG promoter, thereby producing the modified population of peripheral blood nucleated cells, wherein the modified population of peripheral blood nucleated cells as a percentage of the total population of peripheral blood nucleated cells is at least 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90% or 95%.
- the subject may undergo myeloablative conditioning with busulfan prior to administering the population of modified cells.
- methods of generating a population of F-cells in a subject comprising: administering to the subject a population of modified cells comprising a plurality of modified CD34+ or hematopoietic stem cells comprising an indel in an HBG promoter, wherein the percentage of F-cells among circulating red blood cells six months after said administering is at least 90%.
- mean corpuscular HbF per F-cell in the population of F-cells is at least 18 pg.
- the subject may exhibit no vaso- occlusive events (VOEs) within six months after said administering. In certain embodiments, the subject may exhibit no VOEs within one year after said administering.
- VOEs vaso- occlusive events
- a VOE is defined as acute episode of pain with no medically determined cause other than a vaso-occlusion.
- the subject may experience no VOEs 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30 months after administering the population of modified cells to the subject.
- severe VOEs include an acute episode of pain with no cause other than a vaso- occlusion, resulting in either: a ⁇ 24-hour hospital or Emergency Room (ER) observation unit visit; ⁇ 2 visits to a day unit or ER over 72 hours, with both visits requiring oral or parenteral opioids or parenteral non-steroidal anti-inflammatory drugs (NSAIDs); or four visits to any medical facility over any time period (with or without medication requirement) for episodes of pain due to priapism.
- the subject may experience no severe VOEs 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30 months after administering the population of modified cells to the subject.
- the subject may experience fewer than 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, or 20 VOEs 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30 months after administering the population of modified cells to the subject.
- the subject may experience fewer than 2, 3, 4, 5, 6, 7, 8, 9, or 10 severe VOEs 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30 months after administering the population of modified cells to the subject.
- the subject may experience fewer than 5, 10, 15, 20, 25, 30, 35, 40, 45, 50, 55, 60, 65 treatment emergent adverse events (TEAEs) 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30 months after administering the population of modified cells to the subject.
- TEAEs treatment emergent adverse events
- the subject 54 164554159.2 Attorney Docket No.: 118945.8026.WO00 may experience fewer than 2, 3, 4, 5, 6, 7, 8, 9, or 10 serious treatment emergent adverse events (TEAEs) 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30 months after administering the population of modified cells to the subject.
- RNA-guided nucleases can be evaluated by standard methods known in the art. See, e.g. Cotta-Ramusino. The stability of RNP complexes may be evaluated by differential scanning fluorimetry, as described below. Differential Scanning Fluorimetry (DSF) [0165] The thermostability of ribonucleoprotein (RNP) complexes comprising gRNAs and RNA- guided nucleases can be measured via DSF. The DSF technique measures the thermostability of a protein, which can increase under favorable conditions such as the addition of a binding RNA molecule, e.g., a gRNA.
- DSF Differential Scanning Fluorimetry
- a DSF assay can be performed according to any suitable protocol, and can be employed in any suitable setting, including without limitation (a) testing different conditions (e.g. different stoichiometric ratios of gRNA: RNA-guided nuclease protein, different buffer solutions, etc.) to identify optimal conditions for RNP formation; and (b) testing modifications (e.g. chemical modifications, alterations of sequence, etc.) of an RNA-guided nuclease and/or a gRNA to identify those modifications that improve RNP formation or stability.
- different conditions e.g. different stoichiometric ratios of gRNA: RNA-guided nuclease protein, different buffer solutions, etc.
- modifications e.g. chemical modifications, alterations of sequence, etc.
- One readout of a DSF assay is a shift in melting temperature of the RNP complex; a relatively high shift suggests that the RNP complex is more stable (and may thus have greater activity or more favorable kinetics of formation, kinetics of degradation, or another functional characteristic) relative to a reference RNP complex characterized by a lower shift.
- a threshold melting temperature shift may be specified, so that the output is one or more RNPs having a melting temperature shift at or above the threshold.
- the threshold can be 5-10°C (e.g.5°, 6°, 7°, 8°, 9°, 10°) or more, and the output may be one or more RNPs characterized by a melting temperature shift greater than or equal to the threshold.
- DSF assay conditions Two non-limiting examples of DSF assay conditions are set forth below: [0168] To determine the best solution to form RNP complexes, a fixed concentration (e.g.2 ⁇ M) of Cas9 in water+10x SYPRO Orange® (Life Technologies cat#S-6650) is dispensed into a 384 well plate. An equimolar amount of gRNA diluted in solutions with varied pH and salt is then added.
- a fixed concentration e.g.2 ⁇ M
- SYPRO Orange® Life Technologies cat#S-6650
- the second assay consists of mixing various concentrations of gRNA with fixed concentration (e.g.2 ⁇ M) Cas9 in optimal buffer from assay 1 above and incubating (e.g. at RT for 10’) in a 384 well plate.
- Genome editing strategies [0170] The genome editing systems described above are used, in various embodiments of the present disclosure, to generate edits in (i.e. to alter) targeted regions of DNA within or obtained from a cell.
- Genome editing strategies that involve the formation of SSBs or DSBs are characterized by repair outcomes including: (a) deletion of all or part of a targeted region; (b) insertion into or replacement of all or part of a targeted region; or (c) interruption of all or part of a targeted region. This grouping is not intended to be limiting, or to be binding to any particular theory or model, and is offered solely for economy of presentation.
- Replacement of a targeted region generally involves the replacement of all or part of the existing sequence within the targeted region with a homologous sequence, for instance through gene correction or gene conversion, two repair outcomes that are mediated by HDR pathways.
- HDR is promoted by the use of a donor template, which can be single-stranded or double stranded, as described in greater detail below.
- Single or double stranded templates can be exogenous, in which case they will promote gene correction, or they can be endogenous (e.g.
- Exogenous templates can have asymmetric overhangs (i.e. the portion of the template that is complementary to the site of the DSB may be offset in a 3’ or 5’ direction, rather than being centered within the donor template), for instance as described by Richardson 2016 (incorporated by reference herein).
- the template is single 56 164554159.2 Attorney Docket No.: 118945.8026.WO00 stranded, it can correspond to either the complementary (top) or non-complementary (bottom) strand of the targeted region.
- Gene conversion and gene correction are facilitated, in some cases, by the formation of one or more nicks in or around the targeted region, as described in Ran & Hsu 2013 and Cotta-Ramusino.
- a dual-nickase strategy is used to form two offset SSBs that, in turn, form a single DSB having an overhang (e.g. a 5’ overhang).
- Interruption and/or deletion of all or part of a targeted sequence can be achieved by a variety of repair outcomes.
- a sequence can be deleted by simultaneously generating two or more DSBs that flank a targeted region, which is then excised when the DSBs are repaired, as is described in Maeder for the LCA10 mutation.
- a sequence can be interrupted by a deletion generated by formation of a double strand break with single-stranded overhangs, followed by exonucleolytic processing of the overhangs prior to repair.
- One specific subset of target sequence interruptions is mediated by the formation of an indel within the targeted sequence, where the repair outcome is typically mediated by NHEJ pathways (including Alt-NHEJ). NHEJ is referred to as an “error prone” repair pathway because of its association with indel mutations.
- a DSB is repaired by NHEJ without alteration of the sequence around it (a so-called “perfect” or “scarless” repair); this generally requires the two ends of the DSB to be perfectly ligated. Indels, meanwhile, are thought to arise from enzymatic processing of free DNA ends before they are ligated that adds and/or removes nucleotides from either or both strands of either or both free ends. [0176] Because the enzymatic processing of free DSB ends may be stochastic in nature, indel mutations tend to be variable, occurring along a distribution, and can be influenced by a variety of factors, including the specific target site, the cell type used, the genome editing strategy used, etc.
- deletions formed by repair of a single DSB are most commonly in the 1-50 bp range, but can reach greater than 100-200 bp. Insertions formed by repair of a single DSB tend to be shorter and often include short duplications of the sequence immediately surrounding the break site. However, it is possible to obtain large insertions, and in these cases, the inserted sequence has often been traced to other regions of the genome or to plasmid DNA present in the cells. [0177] Indel mutations – and genome editing systems configured to produce indels – are useful for interrupting target sequences, for example, when the generation of a specific final sequence is not required and/or where a frameshift mutation would be tolerated.
- indels can be characterized by (a) their relative and absolute frequencies in the genomes of cells contacted with genome editing systems and (b) the distribution of numerical differences relative to the unedited sequence, e.g. ⁇ 1, ⁇ 2, ⁇ 3, etc.
- multiple gRNAs can be screened to identify those gRNAs that most efficiently drive cutting at a target site based on an indel readout under controlled conditions.
- Guides that produce indels at or above a threshold frequency, or that produce a particular distribution of indels, can be selected for further study and development.
- Indel frequency and distribution can also be useful as a readout for evaluating different genome editing system implementations or formulations and delivery methods, for instance by keeping the gRNA constant and varying certain other reaction conditions or delivery methods.
- Multiplex Strategies [0178] Genome editing systems according to this disclosure may also be employed for multiplex gene editing to generate two or more DSBs, either in the same locus or in different loci.
- RNA-guided nucleases and gRNAs disclosed herein may be used in genome editing systems for multiplex gene editing. Strategies for editing that involve the formation of multiple DSBs, or SSBs, are described in, for instance, Cotta-Ramusino.
- multiple gRNAs and an RNA-guided nuclease may be used in genome editing systems to introduce alterations (e.g., deletions, insertions) into the CCAAT box target region of HBG1 and/or HBG2.
- the RNA-guided nuclease may be a Cpf1 protein.
- Donor template design is described in detail in the literature, for instance in Cotta-Ramusino.
- DNA oligomer donor templates oligodeoxynucleotides or ODNs
- ssODNs single stranded
- dsODNs double-stranded
- donor templates generally include regions that are homologous to regions of DNA within or near (e.g.
- homology arms flanking or adjoining
- 58 164554159.2 Attorney Docket No.: 118945.8026.WO00 [0181]
- the homology arms can have any suitable length (including 0 nucleotides if only one homology arm is used), and 3’ and 5’ homology arms can have the same length, or can differ in length.
- homology arm lengths can be influenced by a variety of factors, such as the desire to avoid homologies or microhomologies with certain sequences such as Alu repeats or other very common elements.
- a 5’ homology arm can be shortened to avoid a sequence repeat element.
- a 3’ homology arm can be shortened to avoid a sequence repeat element.
- both the 5’ and the 3’ homology arms can be shortened to avoid including certain sequence repeat elements.
- some homology arm designs can improve the efficiency of editing or increase the frequency of a desired repair outcome.
- a replacement sequence in donor templates have been described elsewhere, including in Cotta-Ramusino.
- a replacement sequence can be any suitable length (including zero nucleotides, where the desired repair outcome is a deletion), and typically includes one, two, three or more sequence modifications relative to the naturally-occurring sequence within a cell in which editing is desired.
- One common sequence modification involves the alteration of the naturally-occurring sequence to repair a mutation that is related to a disease or condition of which treatment is desired.
- Another common sequence modification involves the alteration of one or more sequences that are complementary to, or then, the PAM sequence of the RNA-guided nuclease or the targeting domain of the gRNA(s) being used to generate an SSB or DSB, to reduce or eliminate repeated cleavage of the target site after the replacement sequence has been incorporated into the target site.
- a linear ssODN can be configured to (i) anneal to the nicked strand of the target nucleic acid, (ii) anneal to the intact strand of the target nucleic acid, (iii) anneal to the plus strand of the target nucleic acid, and/or (iv) anneal to the minus strand of the target nucleic acid.
- An ssODN may have any suitable length, e.g., about, at least, or no more than 80-200 nucleotides (e.g., 80, 90, 100, 110, 120, 130, 140, 150, 160, 170, 180, 190, or 200 nucleotides).
- a template nucleic acid can also be a nucleic acid vector, such as a viral genome or circular double stranded DNA, e.g., a plasmid.
- Nucleic acid vectors comprising donor templates can include other coding or non-coding elements.
- a template nucleic acid can be delivered as part of a viral genome (e.g. in an AAV or lentiviral genome) that includes certain genomic backbone elements (e.g. inverted terminal repeats, in the case of an AAV genome) and optionally includes additional sequences coding for a gRNA and/or an RNA-guided nuclease.
- the donor template can be adjacent to, or flanked by, target sites recognized by 59 164554159.2 Attorney Docket No.: 118945.8026.WO00 one or more gRNAs, to facilitate the formation of free DSBs on one or both ends of the donor template that can participate in repair of corresponding SSBs or DSBs formed in cellular DNA using the same gRNAs.
- Exemplary nucleic acid vectors suitable for use as donor templates are described in Cotta-Ramusino, which is incorporated by reference. [0185] Whatever format is used, a template nucleic acid can be designed to avoid undesirable sequences.
- one or both homology arms can be shortened to avoid overlap with certain sequence repeat elements, e.g., Alu repeats, LINE elements, etc.
- silent, non-pathogenic SNPs may be included in the ssODN donor template to allow for identification of a gene editing event.
- a donor template may be a non-specific template that is non- homologous to regions of DNA within or near a target sequence to be cleaved.
- a donor template or template nucleic acid refers to a nucleic acid sequence which can be used in conjunction with an RNA nuclease molecule and one or more gRNA molecules to alter (e.g., delete, disrupt, or modify) a target DNA sequence.
- the template nucleic acid results in an alteration (e.g., deletion) at the CCAAT box target region of HBG1 and/or HBG2.
- the alteration is a non-naturally occurring alteration.
- the template nucleic acid is an ssODN.
- the ssODN is a positive strand or a negative strand.
- the 5’ homology arm comprises a 5’ phosphorothioate (PhTx) modification.
- the 3’ homology arm comprises a 3’ PhTx modification.
- the template nucleic acid comprises a 5’ and 3’ PhTx modification.
- the ssODNs for introducing alterations (e.g., deletions) at the CCAAT box target region may be used in conjunction with an RNA nuclease and one or more gRNAs that target the CCAAT target region.
- Target cells [0191] Genome editing systems according to this disclosure can be used to manipulate or alter a cell, e.g., to edit or alter a target nucleic acid.
- the manipulating can occur, in various embodiments, in vivo or ex vivo.
- a variety of cell types can be manipulated or altered according to the embodiments of this disclosure, and in some cases, such as in vivo applications, a plurality of cell types are altered or manipulated, for example by delivering genome editing systems according to this disclosure to a plurality of cell types. In other cases, however, it may be desirable to limit manipulation or alteration 60 164554159.2 Attorney Docket No.: 118945.8026.WO00 to a particular cell type or types.
- the cell may be an embryonic stem cell, induced pluripotent stem cell (iPSC), hematopoietic stem/progenitor cell (HSPC), or other stem or progenitor cell type that differentiates into a cell type of relevance to a given application or indication.
- iPSC induced pluripotent stem cell
- HSPC hematopoietic stem/progenitor cell
- the cell being altered or manipulated is, variously, a dividing cell or a non- dividing cell, depending on the cell type(s) being targeted and/or the desired editing outcome.
- the cells can be used (e.g. administered to a subject) immediately, or they can be maintained or stored for later use. Those of skill in the art will appreciate that cells can be maintained in culture or stored (e.g. frozen in liquid nitrogen) using any suitable method known in the art.
- the genome editing systems of this disclosure can be implemented in any suitable manner, meaning that the components of such systems, including without limitation the RNA-guided nuclease, gRNA, and optional donor template nucleic acid, can be delivered, formulated, or administered in any suitable form or combination of forms that results in the transduction, expression or introduction of a genome editing system and/or causes a desired repair outcome in a cell, tissue or subject.
- Tables 2 and 3 set forth several, non-limiting examples of genome editing system implementations. Those of skill in the art will appreciate, however, that these listings are not comprehensive, and that other implementations are possible.
- the table lists several exemplary implementations of a genome editing system comprising a single gRNA and an optional donor template.
- genome editing systems according to this disclosure can incorporate multiple gRNAs, multiple RNA-guided nucleases, and other components such as proteins, and a variety of implementations will be evident to the skilled artisan based on the principles illustrated in the table.
- [N/A] indicates that the genome editing system does not include the indicated component.
- Nucleic acids encoding the various elements of a genome editing system according to the present disclosure can be administered to subjects or delivered into cells by art-known methods or as described herein.
- RNA-guided nuclease-encoding and/or gRNA-encoding DNA can be delivered by, e.g., vectors (e.g., viral or non-viral vectors), non-vector based methods (e.g., using naked DNA or DNA complexes), or a combination thereof.
- vectors e.g., viral or non-viral vectors
- non-vector based methods e.g., using naked DNA or DNA complexes
- Nucleic acids encoding genome editing systems or components thereof can be delivered directly to cells as naked DNA or RNA, for instance by means of transfection or electroporation, or can be conjugated to molecules (e.g., N-acetylgalactosamine) promoting uptake by the target cells (e.g., erythrocytes, HSCs).
- molecules e.g., N-acetylgalactosamine
- Nucleic acid vectors such as the vectors summarized in Table 3, can also be used.
- Nucleic acid vectors can comprise one or more sequences encoding genome editing system components, such as an RNA-guided nuclease, a gRNA and/or a donor template.
- a vector can also comprise a sequence encoding a signal peptide (e.g., for nuclear localization, nucleolar localization, or mitochondrial localization), associated with (e.g., inserted into or fused to) a sequence coding for a protein.
- a nucleic acid vectors can include a Cas9 coding sequence that includes one or more nuclear localization sequences (e.g., a nuclear localization sequence from SV40).
- the nucleic acid vector can also include any suitable number of regulatory/control elements, e.g., promoters, enhancers, introns, polyadenylation signals, Kozak consensus sequences, or internal ribosome entry sites (IRES). These elements are well known in the art, and are described in Cotta- Ramusino.
- Nucleic acid vectors according to this disclosure include recombinant viral vectors. Exemplary viral vectors are set forth in Table 3, and additional suitable viral vectors and their use and production are described in Cotta-Ramusino. Other viral vectors known in the art can also be used.
- viral particles can be used to deliver genome editing system components in nucleic acid and/or peptide form.
- non-viral vectors can be used to deliver nucleic acids encoding genome editing systems according to the present disclosure.
- One important category of non-viral nucleic acid vectors are nanoparticles, which can be organic or inorganic. Nanoparticles are well 64 164554159.2 Attorney Docket No.: 118945.8026.WO00 known in the art, and are summarized in Cotta-Ramusino. Any suitable nanoparticle design can be used to deliver genome editing system components or nucleic acids encoding such components.
- organic (e.g. lipid and/or polymer) nanoparticles can be suitable for use as delivery vehicles in certain embodiments of this disclosure.
- lipids for use in nanoparticle formulations, and/or gene transfer are shown in Table 4 and Table 5 lists exemplary polymers for use in gene transfer and/or nanoparticle formulations.
- Table 4 Lipids Used for Gene Transfer Lipid Abbreviation Feature 12-Dioleo l-sn- l cero-3- hos hatid lcholine DOPC Hel er 65 164554159.2 Attorney Docket No.: 118945.8026.WO00 rac-[2(2,3-Dihexadecyloxypropyl- CLIP-6 Cationic oxymethyloxy)ethyl]trimethylammonium bromide Polymer Abbreviation 164554159.2 Attorney Docket No.: 118945.8026.WO00 Poly( ⁇ -aminoester) Poly(4-hydroxy-L-proline ester) PHP selectively target certain cell types.
- targeting modifications can include e.g., cell specific antigens, monoclonal antibodies, single chain antibodies, aptamers, polymers, sugars (e.g., N- acetylgalactosamine (GalNAc)), and cell penetrating peptides.
- Such vectors also optionally use fusogenic and endosome-destabilizing peptides/polymers, undergo acid-triggered conformational changes (e.g., to accelerate endosomal escape of the cargo), and/or incorporate a stimuli-cleavable polymer, e.g., for release in a cellular compartment.
- a stimuli-cleavable polymer e.g., for release in a cellular compartment.
- disulfide-based cationic polymers that are cleaved in the reducing cellular environment can be used.
- one or more nucleic acid molecules are delivered.
- the nucleic acid molecule is delivered at the same time as one or more of the components of the genome editing system.
- the nucleic acid molecule is delivered before or after (e.g., less than about 30 minutes, 1 hour, 2 hours, 3 hours, 6 hours, 9 hours, 12 hours, 1 day, 2 days, 3 days, 1 week, 2 weeks, 67 164554159.2 Attorney Docket No.: 118945.8026.WO00 or 4 weeks) one or more of the components of the genome editing system are delivered.
- the nucleic acid molecule is delivered by a different means than one or more of the components of the genome editing system, e.g., the RNA-guided nuclease component and/or the gRNA component, are delivered.
- the nucleic acid molecule can be delivered by any of the delivery methods described herein.
- the nucleic acid molecule can be delivered by a viral vector, e.g., an integration-deficient lentivirus, and the RNA-guided nuclease molecule component and/or the gRNA component can be delivered by electroporation, e.g., such that the toxicity caused by nucleic acids (e.g., DNAs) can be reduced.
- the nucleic acid molecule encodes a therapeutic protein, e.g., a protein described herein.
- the nucleic acid molecule encodes an RNA molecule, e.g., an RNA molecule described herein.
- RNPs and/or RNA encoding genome editing system components
- RNPs complexes of gRNAs and RNA-guided nucleases
- RNAs encoding RNA- guided nucleases and/or gRNAs can be delivered into cells or administered to subjects by art-known methods, some of which are described in Cotta-Ramusino.
- RNA-guided nuclease-encoding and/or gRNA-encoding RNA can be delivered, e.g., by microinjection, electroporation, transient cell compression or squeezing (see, e.g., Lee 2012).
- Lipid-mediated transfection, peptide-mediated delivery, GalNAc- or other conjugate-mediated delivery, and combinations thereof, can also be used for delivery in vitro and in vivo.
- a protective, interactive, non-condensing (PINC) system may be used for delivery.
- PINC protective, interactive, non-condensing
- In vitro delivery via electroporation comprises mixing the cells with the RNA encoding RNA- guided nucleases and/or gRNAs, with or without donor template nucleic acid molecules, in a cartridge, chamber or cuvette and applying one or more electrical impulses of defined duration and amplitude.
- Systems and protocols for electroporation are known in the art, and any suitable electroporation tool and/or protocol can be used in connection with the various embodiments of this disclosure.
- Genome editing systems, or cells altered or manipulated using such systems can be administered to subjects by any suitable mode or route, whether local or systemic.
- Systemic modes of administration include oral and parenteral routes.
- Parenteral routes include, by way of example, intravenous, intramarrow, intrarterial, intramuscular, intradermal, subcutaneous, intranasal, and intraperitoneal routes.
- Components administered systemically can be modified or formulated to target, e.g., HSCs, hematopoietic stem/progenitor cells, or erythroid progenitors or precursor cells.
- Local modes of administration include, by way of example, intramarrow injection into the 68 164554159.2 Attorney Docket No.: 118945.8026.WO00 trabecular bone or intrafemoral injection into the marrow space, and infusion into the portal vein.
- significantly smaller amounts of the components can exert an effect when administered locally (for example, directly into the bone marrow) compared to when administered systemically (for example, intravenously).
- Local modes of administration can reduce or eliminate the incidence of potentially toxic side effects that may occur when therapeutically effective amounts of a component are administered systemically.
- Administration can be provided as a periodic bolus (for example, intravenously) or as continuous infusion from an internal reservoir or from an external reservoir (for example, from an intravenous bag or implantable pump).
- Components can be administered locally, for example, by continuous release from a sustained release drug delivery device.
- components can be formulated to permit release over a prolonged period of time.
- a release system can include a matrix of a biodegradable material or a material which releases the incorporated components by diffusion.
- the components can be homogeneously or heterogeneously distributed within the release system.
- release systems can be useful, however, the choice of the appropriate system will depend upon rate of release required by a particular application. Both non-degradable and degradable release systems can be used.
- Suitable release systems include polymers and polymeric matrices, non-polymeric matrices, or inorganic and organic excipients and diluents such as, but not limited to, calcium carbonate and sugar (for example, trehalose). Release systems may be natural or synthetic. However, synthetic release systems are preferred because generally they are more reliable, more reproducible and produce more defined release profiles.
- the release system material can be selected so that components having different molecular weights are released by diffusion through or degradation of the material.
- Representative synthetic, biodegradable polymers include, for example: polyamides such as poly(amino acids) and poly(peptides); polyesters such as poly(lactic acid), poly(glycolic acid), poly(lactic-co-glycolic acid), and poly(caprolactone); poly(anhydrides); polyorthoesters; polycarbonates; and chemical derivatives thereof (substitutions, additions of chemical groups, for example, alkyl, alkylene, hydroxylations, oxidations, and other modifications routinely made by those skilled in the art), copolymers and mixtures thereof.
- polyamides such as poly(amino acids) and poly(peptides)
- polyesters such as poly(lactic acid), poly(glycolic acid), poly(lactic-co-glycolic acid), and poly(caprolactone)
- poly(anhydrides) polyorthoesters
- polycarbonates and chemical derivatives thereof (substitutions, additions of chemical groups, for example, alkyl, alkylene, hydroxylation
- Representative synthetic, non-degradable polymers include, for example: polyethers such as poly(ethylene oxide), poly(ethylene glycol), and poly(tetramethylene oxide); vinyl polymers-polyacrylates and polymethacrylates such as methyl, ethyl, other alkyl, hydroxyethyl methacrylate, acrylic and methacrylic acids, and others such as poly(vinyl alcohol), poly(vinyl pyrolidone), and poly(vinyl acetate); poly(urethanes); cellulose and its derivatives such as alkyl, hydroxyalkyl, ethers, esters, nitrocellulose, and various cellulose acetates; polysiloxanes; and any chemical derivatives thereof (substitutions, additions of chemical groups, for 69 164554159.2 Attorney Docket No.: 118945.8026.WO00 example, alkyl, alkylene, hydroxylations, oxidations, and other modifications routinely made by those skilled in the art), copolymers and mixture
- Poly(lactide-co-glycolide) microsphere can also be used.
- the microspheres are composed of a polymer of lactic acid and glycolic acid, which are structured to form hollow spheres.
- the spheres can be approximately 15-30 microns in diameter and can be loaded with components described herein.
- genome editing systems, system components and/or nucleic acids encoding system components are delivered with a block copolymer such as a poloxamer or a poloxamine. Multi-modal or differential delivery of components [0213] Skilled artisans will appreciate, in view of the instant disclosure, that different components of genome editing systems disclosed herein can be delivered together or separately and simultaneously or non-simultaneously.
- Different or differential modes as used herein refer to modes of delivery that confer different pharmacodynamic or pharmacokinetic properties on the subject component molecule, e.g., a RNA- guided nuclease molecule, gRNA, template nucleic acid, or payload.
- the modes of delivery can result in different tissue distribution, different half-life, or different temporal distribution, e.g., in a selected compartment, tissue, or organ.
- Some modes of delivery e.g., delivery by a nucleic acid vector that persists in a cell, or in progeny of a cell, e.g., by autonomous replication or insertion into cellular nucleic acid, result in more persistent expression of and presence of a component.
- examples include viral, e.g., AAV or lentivirus, delivery.
- the components of a genome editing system e.g., a RNA-guided nuclease and a gRNA
- a gRNA can be delivered by such modes.
- the RNA-guided nuclease molecule component can be delivered by a mode which results in less persistence or less exposure to the body or a particular compartment or tissue or organ.
- a first mode of delivery is used to deliver a first component and a second mode of delivery is used to deliver a second component.
- the first mode of delivery confers a first pharmacodynamic or pharmacokinetic property.
- the first pharmacodynamic property can be, e.g., distribution, persistence, or exposure, of the component, or of a nucleic acid that 70 164554159.2 Attorney Docket No.: 118945.8026.WO00 encodes the component, in the body, a compartment, tissue or organ.
- the second mode of delivery confers a second pharmacodynamic or pharmacokinetic property.
- the second pharmacodynamic property can be, e.g., distribution, persistence, or exposure, of the component, or of a nucleic acid that encodes the component, in the body, a compartment, tissue or organ.
- the first pharmacodynamic or pharmacokinetic property e.g., distribution, persistence or exposure
- the first mode of delivery is selected to optimize, e.g., minimize, a pharmacodynamic or pharmacokinetic property, e.g., distribution, persistence or exposure.
- the second mode of delivery is selected to optimize, e.g., maximize, a pharmacodynamic or pharmacokinetic property, e.g., distribution, persistence or exposure.
- the first mode of delivery comprises the use of a relatively persistent element, e.g., a nucleic acid, e.g., a plasmid or viral vector, e.g., an AAV or lentivirus. As such vectors are relatively persistent product transcribed from them would be relatively persistent.
- the second mode of delivery comprises a relatively transient element, e.g., an RNA or protein.
- the first component comprises gRNA
- the delivery mode is relatively persistent, e.g., the gRNA is transcribed from a plasmid or viral vector, e.g., an AAV or lentivirus. Transcription of these genes would be of little physiological consequence because the genes do not encode for a protein product, and the gRNAs are incapable of acting in isolation.
- the second component a RNA-guided nuclease molecule, is delivered in a transient manner, for example as mRNA or as protein, ensuring that the full RNA-guided nuclease molecule/gRNA complex is only present and active for a short period of time.
- the components can be delivered in different molecular form or with different delivery vectors that complement one another to enhance safety and tissue specificity.
- Use of differential delivery modes can enhance performance, safety, and/or efficacy, e.g., the likelihood of an eventual off-target modification can be reduced.
- Delivery of immunogenic components, e.g., Cas9 molecules, by less persistent modes can reduce immunogenicity, as peptides from the bacterially-derived Cas enzyme are displayed on the surface of the cell by MHC molecules.
- a two-part delivery system can alleviate these drawbacks.
- Differential delivery modes can be used to deliver components to different, but overlapping target regions. The formation active complex is minimized outside the overlap of the target regions.
- a first component e.g., a gRNA is delivered by a first delivery mode that results in a first spatial, e.g., tissue, distribution.
- a second component e.g., a RNA-guided nuclease molecule is delivered by a second delivery mode that results in a second spatial, e.g., tissue, distribution.
- the first mode comprises a first element selected from a liposome, nanoparticle, e.g., polymeric nanoparticle, and a nucleic acid, e.g., viral vector.
- the second mode comprises a second element selected from the group.
- the first mode of delivery comprises a first targeting element, e.g., a cell specific receptor or an antibody, and the second mode of delivery does not include that element.
- the second mode of delivery comprises a second targeting element, e.g., a second cell specific receptor or second antibody.
- Example 1 Autologous CD34+ Cells Edited by RNP27 at the HBG1/HBG2 Promoter Show Early, Promising Efficacy and Safety in a Phase 1/2 Study in Patients with Severe Sickle Cell Disease
- SCD Sickle cell disease
- RBCs red blood cells
- VOEs vaso-occlusive events
- RNP27 (Table 12) comprises a gRNA comprising the sequence set forth in SEQ ID NO:237 complexed with a Cpf1 protein comprising the sequence set forth in SEQ ID NO:283 (nucleotide 72 164554159.2 Attorney Docket No.: 118945.8026.WO00 sequence is SEQ ID NO:300).
- RNP27 cleaves the genomic DNA in the distal CCAAT box region of ⁇ -globin gene promoters (HBG1 and HBG2) on chromosome 11.
- the key endpoints of the study are (1) rate of severe vaso-occlusive events (VOEs) requiring medical attention and (2) safety and tolerability of autologous RNP27 edited CD34+ cells.
- VOE is defined as an acute episode of pain with no medically determined cause other than a vaso-occlusion.
- a severe VOE includes an acute episode of pain with no cause other than a vaso-occlusion, resulting in either: a ⁇ 24-hour hospital or Emergency Room (ER) observation unit visit; ⁇ 2 visits to a day unit or ER over 72 hours, with both visits requiring oral or parenteral opioids or parenteral non-steroidal anti- inflammatory drugs (NSAIDs); or four visits to any medical facility over any time period (with or without medication requirement) for episodes of pain due to priapism.
- NSAIDs parenteral non-steroidal anti- inflammatory drugs
- Mutations point mutations and the 13 bp hereditary persistence of fetal hemoglobin [HPFH] deletion
- HPFH fetal hemoglobin
- HbF fetal hemoglobin
- the insertions and/or deletions (indels) generated by RNP27 in this region act by an analogous mechanism to HPFH mutations in de- repressing ⁇ -globin expression in adults.
- autologous RNP27 edited CD34+ cells administered as a one-time, autologous CD34+ cell therapy has the potential to lead to sustained, elevated levels of HbF in SCD patients, ameliorating the disease phenotypes.
- 40 subjects are 18–50 years old with a diagnosis of severe SCD defined as a history of ⁇ 2 severe vaso-occlusive events (VOEs) per year in the 2 year-period prior to provision of informed consent.
- VOEs vaso-occlusive events
- the key endpoints in the study are (1) proportion of patients achieving complete resolution of severe vaso-occlusive events (VOEs) requiring medical attention (Time Frame: up to 2 years post infusion), and (2) safety and tolerability of treatment with RNP27 edited CD34+ cells.
- the secondary outcome measures of the study include: 1. Proportion of subjects with mean HbF > 20% (HbF/Hb) compared with pre-conditioning Baseline (time frame: up to 2 years post RNP27 edited CD34+ cell infusion); 2.
- Diagnosis of severe sickle cell disease as defined by: • Documented severe SCD genotype ( ⁇ S / ⁇ S , ⁇ S / ⁇ 0 , or ⁇ S / ⁇ + ) • History of at least two severe vaso-occlusive crisis events per year requiring medical attention despite hydroxyurea or other supportive care measures in the two year- period prior to provision of informed consent; 2. Karnofsky Performance Status ⁇ 80. [0237] Exclusion criteria include: 1. Available 10/10 HLA-matched related donor; 2. Prior hematopoietic stem cell transplantation (HSCT) or contraindications to autologous HSCT; 3.
- HSCT Prior hematopoietic stem cell transplantation
- HSCs hematopoietic stem cells
- busulfan any contraindications to the use of busulfan and any other medicinal products required during the myeloablative conditioning, including hypersensitivity to the active substances or to any of the excipients; 4. Unable to receive red blood cell (RBC) transfusion for any reason; 5. Unable or unwilling to comply with standard of care changes in background medical treatment in preparation of, during, or following HSCT, including and not limited to discontinuation of hydroxyurea, voxelotor, crizanlizumab, or L-glutamine; 6.
- CD34+ cells are enriched using the CliniMACS Plus system, aliquoted, cryopreserved in Cryostor CS10, and stored in liquid nitrogen vapor phase.
- CD34+ cells are thawed, cultured for 2 days in complete media consisting of X-VIVO 10, supplemented with 1 X Glutamax, 100 ng/mL stem cell factor (SCF), 100 ng/mL thrombopoietin (TPO), and 100 ng/mL FMS-like tyrosine kinase 3 ligand (Flt3L), mixed with RNP (at a gRNA/protein molar ratio of 2.4) to a final concentration of 6 ⁇ M, and electroporated with a Maxcyte GT electroporation device per manufacturer’s instruction.
- SCF stem cell factor
- TPO thrombopoietin
- Flt3L FMS-like tyrosine kinase 3 ligand
- subjects receive a single infusion of autologous RNP27 edited CD34+ cells ( ⁇ 3 ⁇ 10 6 CD34+ cells/kg): For example, Subject 1 received a dose of 10.0 x 10 6 RNP27 edited CD34+ cells/kg, Subject 2 received a dose of 4.0 x 10 6 RNP27 edited CD34+ cells/kg, Subject 3 received a dose of 4.1 x 10 6 RNP27 edited CD34+ cells/kg, and 75 164554159.2 Attorney Docket No.: 118945.8026.WO00 Subject 4 received a dose of 3.7 x 10 6 RNP27 edited CD34+ cells/kg.
- Subjects 1 and 2 experienced 4 and 3 vaso-occlusive events, respectively, annually prior to enrollment in the trial.
- Subject 1 had > 10 outpatient infusions, emergency department (ED) visits, or hospital admissions for pain crises in the first year prior to treatment and > 30 outpatient infusions, ED visits, or hospital admissions for pain crisis in the second year prior to treatment.
- Subject 2 had ⁇ 10 outpatient infusions, ED visits, or hospital admissions for pain crises in the first year prior to treatment and > 2 ED visits or hospital admissions for pain crises in the second year prior to treatment.
- the modified cells had decreased the patients’ sickle cell crises to zero.
- Demographics of Subjects 1-4 are set forth in Table 7.
- the infusion and engraftment results for subjects 1-11 are shown in Table 8.
- neutrophil and platelet engraftment were achieved within a mean of 23.7 and 24.3 days of infusion, respectively (Table 8).
- neutrophil and platelet engraftment were achieved within 23 and 19 days of infusion, respectively (Table 9).
- neutrophil and platelet engraftment were achieved within 29 and 37 days of infusion, respectively (Table 9).
- neutrophil and platelet engraftment were achieved within 23 and 23 days of infusion, respectively (Table 9).
- neutrophil and platelet engraftment were achieved within 24 and 28 days of infusion, respectively (Table 9).
- HbF level and % F-cells were 45.5% and 96.0%, respectively, vs 5.0% and 31.4% at baseline (Fig.4A, Fig.4C).
- HbF level and % F-cells were 44.3% and 96.5% respectively and at the 8 months follow-up visit, HbF level and % F-cells were 44.1% and 95.5% respectively (Fig.4A, Fig.4C).
- the % HbF target is 30 for clinical benefit though lower levels can benefit the patient (Steinberg 2014).
- Fig.4F shows hemoglobin fractionation (%) data (HbF, HbS and other hemoglobin as a percentage of total hemoglobin), the percentage of F-Cells, total hemoglobin (g/dL), and MCH-F/F-cell (pg) for Subject 1 at various study days before and after autologous RNP27 edited CD34+ cell infusion.
- Results from Subject 1 show HbF expressed in > 95% of RBCs with concentration above anti-sickling threshold (10 pg/RBC).
- the data for Subject 1 also indicate a robust increase of HbF, well above the sickling threshold, rapidly normalizing total hemoglobin, well into normal range.
- Fig.4G shows hemoglobin fractionation (%) data (HbF, HbS and other hemoglobin as a percentage of total hemoglobin), the percentage of F-Cells, total hemoglobin (g/dL), and MCH-F/F-cell (pg) for Subject 2 at various study days before and after autologous RNP27 edited CD34+ cell infusion.
- Fetal hemoglobin was expressed in >85% F-cells of RBCs with concentration above anti-sickling threshold (10 pg/RBC) (Fig.4H).
- the mean corpuscular HbF increase was clinically meaningful for SCD patients as the mean corpuscular HbF reached above 10 pg/RBC (Figs.4I, 4J) (10 pg/RBC is the threshold for protection from sickling (Steinberg 2014)).
- Data for up to 6 months post-infusion with autologous RNP27 edited CD34+ cells for Subject 2 is shown in Fig.5B.
- the data for Subject 2 indicate a robust increase of HbF, well above the sickling threshold, rapidly normalizing total hemoglobin, well into normal range.
- Data for up to 18 months post-infusion with autologous RNP27 edited CD34+ cells for 10 subjects (5 male patients, 5 female patients) is shown in Fig.5E.
- patients with SCD show clinically meaningful improvements in HbF levels with total Hb returning to the normal range.
- Subjects show pancellular expression of HbF in RBCs (see Fig.6A for Subjects 1-4). Mean corpuscular HbF is depicted in Fig.6B (Subjects 1-4), and is above the 10 pg/RBC threshold for protection from sickling.
- Results indicated that editing CD34 + cells using RNP27 resulted in ⁇ 80% editing and no detectable off-target editing. High levels of HBG1 and HBG2 promoter editing in peripheral blood nucleated cells post RNP27 edited CD34+ cell infusion were observed, predicting durable clinical benefit (Fig.7). Editing levels in peripheral blood nucleated cells were >80% in both Subjects 1 and 2. Editing levels in Subject 1 bone marrow were >80% (Fig.8A).
- HbF reached > 45.4% at Month 5 after RNP27 edited CD34+ cell infusion.
- total hemoglobin reached 16.4 g/dL, F-cell pancellularity was 96%, and mean corpuscular HbF rose to 13.8 pg/RBC, exceeding the 10 pg/RBC threshold to suppress RBC sickling.
- Example 2 Treatment of ⁇ -hemoglobinopathy using edited hematopoietic stem cells
- the methods and genome editing systems disclosed herein may be used for the treatment of a ⁇ -hemoglobinopathy, such as sickle cell disease or beta-thalassemia, in a patient in need thereof.
- genome editing may be performed on cells derived from the patient in an autologous 81 164554159.2 Attorney Docket No.: 118945.8026.WO00 procedure. Correction of the patient’s cells ex-vivo and reintroduction of the cells into the patient may result in increased HbF expression and treatment of the ⁇ -hemoglobinopathy.
- HSCs may be extracted from the bone marrow of a patient with a ⁇ - hemoglobinopathy using techniques that are well-known to skilled artisans.
- the HSCs may be modified using methods disclosed herein for genome editing.
- RNPs comprised of guide RNAs (gRNA) that target one or more regions in the HBG gene complexed with an RNA-guided nuclease may be used to edit the HSCs.
- the RNA-guided nuclease may be a Cpf1 protein.
- the Cpf1 protein may comprise a sequence set forth in SEQ ID NOs:200, 201, 205-215, 221, 222-226, 280-283, 293-295 (Cpf1 polypeptide sequences).
- the Cpf1 protein may be encoded by a sequence comprising SEQ ID NOs:216-218, 296-303 (Cpf1 polynucleotide sequences).
- the Cpf1 protein may comprise the sequence set forth in SEQ ID NO:283.
- the gRNA may be a modified or unmodified gRNA.
- the gRNA may comprise a sequence set forth in Table 13.
- the gRNA may comprise the sequence set forth in SEQ ID NO:237.
- the RNP complex may comprise an RNP complex set forth in Table 13.
- the RNP complex may include a gRNA comprising the sequence set forth in SEQ ID NO:237 and a Cpf1 protein comprising the sequence set forth in SEQ ID NO:283 (RNP27, Table 13).
- modified HSCs have an increase in the frequency or level of an indel in the human HBG1 gene, HBG2 gene, or both, relative to unmodified HSCs.
- the modified HSCs can differentiate into erythroid cells that express an increased level of HbF.
- a population of the modified HSCs may be selected for reintroduction into the patient via transfusion or other methods known to skilled artisans.
- the population of modified HSCs for reintroduction may be selected based on, for example, increased HbF expression of the erythroid progeny of the modified HSCs or increased indel frequency of the modified HSCs.
- any form of ablation prior to reintroduction of the cells may be used to enhance engraftment of the modified HSCs.
- peripheral blood stem cells can be extracted from a patient with a ⁇ -hemoglobinopathy using techniques that are well-known to skilled artisans (e.g., apheresis or leukapheresis) and stem cells can be removed from the PBSCs.
- the genome editing methods described above can be performed on the stem cells and the modified stem cells can be reintroduced into the patient as described above.
- UrUrUrUrArArUr (SEQ ID UrUrCrUrArCrUrC NO:202) 164554159.2 Attorney Docket No.: 118945.8026.WO00 TCGTGCTGCTTC UUGGUC ATGTGTTTTTGT (SEQ ID 164554159.2 Attorney Docket No.: 118945.8026.WO00 TCAAAAGACCT (SEQ ID TTTrUrArArUrUr NO:202) 164554159.2 Attorney Docket No.: 118945.8026.WO00 CGGTTTTrUrArAr (SEQ ID UrUrUrCrUrArCrU NO:202) 90 164554159.2 Attorney Docket No.: 118945.8026.WO00 251 mUm
Landscapes
- Health & Medical Sciences (AREA)
- Life Sciences & Earth Sciences (AREA)
- Medicinal Chemistry (AREA)
- Immunology (AREA)
- Public Health (AREA)
- General Health & Medical Sciences (AREA)
- Cell Biology (AREA)
- Developmental Biology & Embryology (AREA)
- Pharmacology & Pharmacy (AREA)
- Animal Behavior & Ethology (AREA)
- Veterinary Medicine (AREA)
- Chemical & Material Sciences (AREA)
- Engineering & Computer Science (AREA)
- Hematology (AREA)
- Zoology (AREA)
- Virology (AREA)
- Biotechnology (AREA)
- Biomedical Technology (AREA)
- Epidemiology (AREA)
- Bioinformatics & Cheminformatics (AREA)
- Diabetes (AREA)
- Chemical Kinetics & Catalysis (AREA)
- General Chemical & Material Sciences (AREA)
- Nuclear Medicine, Radiotherapy & Molecular Imaging (AREA)
- Organic Chemistry (AREA)
- Medicines Containing Material From Animals Or Micro-Organisms (AREA)
Abstract
Description
Claims
Priority Applications (1)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| EP23841142.5A EP4630554A1 (en) | 2022-12-05 | 2023-12-05 | Systems and methods for the treatment of hemoglobinopathies |
Applications Claiming Priority (14)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| US202263386167P | 2022-12-05 | 2022-12-05 | |
| US63/386,167 | 2022-12-05 | ||
| US202363481996P | 2023-01-27 | 2023-01-27 | |
| US63/481,996 | 2023-01-27 | ||
| US202363487700P | 2023-03-01 | 2023-03-01 | |
| US63/487,700 | 2023-03-01 | ||
| US202363490729P | 2023-03-16 | 2023-03-16 | |
| US63/490,729 | 2023-03-16 | ||
| US202363498218P | 2023-04-25 | 2023-04-25 | |
| US63/498,218 | 2023-04-25 | ||
| US202363507093P | 2023-06-08 | 2023-06-08 | |
| US63/507,093 | 2023-06-08 | ||
| US202363593938P | 2023-10-27 | 2023-10-27 | |
| US63/593,938 | 2023-10-27 |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| WO2024123842A1 true WO2024123842A1 (en) | 2024-06-13 |
Family
ID=89619050
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/US2023/082613 Ceased WO2024123842A1 (en) | 2022-12-05 | 2023-12-05 | Systems and methods for the treatment of hemoglobinopathies |
Country Status (2)
| Country | Link |
|---|---|
| EP (1) | EP4630554A1 (en) |
| WO (1) | WO2024123842A1 (en) |
Citations (7)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO2015070083A1 (en) | 2013-11-07 | 2015-05-14 | Editas Medicine,Inc. | CRISPR-RELATED METHODS AND COMPOSITIONS WITH GOVERNING gRNAS |
| WO2015138510A1 (en) | 2014-03-10 | 2015-09-17 | Editas Medicine., Inc. | Crispr/cas-related methods and compositions for treating leber's congenital amaurosis 10 (lca10) |
| WO2016073990A2 (en) | 2014-11-07 | 2016-05-12 | Editas Medicine, Inc. | Methods for improving crispr/cas-mediated genome-editing |
| WO2016182959A1 (en) | 2015-05-11 | 2016-11-17 | Editas Medicine, Inc. | Optimized crispr/cas9 systems and methods for gene editing in stem cells |
| WO2017184768A1 (en) | 2016-04-19 | 2017-10-26 | The Broad Institute Inc. | Novel crispr enzymes and systems |
| WO2021119040A1 (en) * | 2019-12-08 | 2021-06-17 | Editas Medicine, Inc. | Modified cells and methods for the treatment of hemoglobinopathies |
| WO2023014727A1 (en) * | 2021-08-02 | 2023-02-09 | Editas Medicine, Inc. | Systems and methods for the treatment of hemoglobinopathies |
-
2023
- 2023-12-05 EP EP23841142.5A patent/EP4630554A1/en active Pending
- 2023-12-05 WO PCT/US2023/082613 patent/WO2024123842A1/en not_active Ceased
Patent Citations (7)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO2015070083A1 (en) | 2013-11-07 | 2015-05-14 | Editas Medicine,Inc. | CRISPR-RELATED METHODS AND COMPOSITIONS WITH GOVERNING gRNAS |
| WO2015138510A1 (en) | 2014-03-10 | 2015-09-17 | Editas Medicine., Inc. | Crispr/cas-related methods and compositions for treating leber's congenital amaurosis 10 (lca10) |
| WO2016073990A2 (en) | 2014-11-07 | 2016-05-12 | Editas Medicine, Inc. | Methods for improving crispr/cas-mediated genome-editing |
| WO2016182959A1 (en) | 2015-05-11 | 2016-11-17 | Editas Medicine, Inc. | Optimized crispr/cas9 systems and methods for gene editing in stem cells |
| WO2017184768A1 (en) | 2016-04-19 | 2017-10-26 | The Broad Institute Inc. | Novel crispr enzymes and systems |
| WO2021119040A1 (en) * | 2019-12-08 | 2021-06-17 | Editas Medicine, Inc. | Modified cells and methods for the treatment of hemoglobinopathies |
| WO2023014727A1 (en) * | 2021-08-02 | 2023-02-09 | Editas Medicine, Inc. | Systems and methods for the treatment of hemoglobinopathies |
Non-Patent Citations (80)
| Title |
|---|
| "NCBI", Database accession no. NC 000011 |
| AHERN ET AL., BR J HAEMATOL, vol. 25, no. 4, 1973, pages 437 - 444 |
| AKINBAMI, HEMOGLOBIN, vol. 40, 2016, pages 64 - 65 |
| ALIYU ET AL., AM J HEMATOL, vol. 83, 2008, pages 63 - 70 |
| ANGASTINIOTISMODELL, ANN N Y ACAD SCI, vol. 850, 1998, pages 251 - 269 |
| ANONYMOUS: "AsCas12a Gene Editing of HBG1/2 Promoters with EDIT-301 Results in Rapid and Sustained Normalization of Hemoglobin and Increased Fetal Hemoglobin in Patients with Severe Sickle Cell Disease and Transfusion-Dependent Beta-Thalassemia | Blood | American Society of Hematology", 28 November 2023 (2023-11-28), XP093148120, Retrieved from the Internet <URL:https://ashpublications.org/blood/article/142/Supplement%201/4996/500993/AsCas12a-Gene-Editing-of-HBG1-2-Promoters-with> [retrieved on 20240404] * |
| ANONYMOUS: "Preclinical Development of EDIT301, an Autologous Cell Therapy Comprising AsCas12a-RNP Modified Mobilized Peripheral Blood-CD34 + Cells for the Potential Treatment of Transfusion Dependent Beta Thalassemia | Blood | American Society of Hematology", 5 November 2021 (2021-11-05), XP093148051, Retrieved from the Internet <URL:https://ashpublications.org/blood/article/138/Supplement%201/1858/480833/Preclinical-Development-of-EDIT301-an-Autologous> [retrieved on 20240404] * |
| BARBOSA ET AL., BRAZ J MED BIO RES, vol. 43, no. 8, 2010, pages 705 - 711 |
| BARNETT CRISTI ET AL: "Editas Medicine Announces Positive Safety and Efficacy Data from the First Two Patients Treated in the RUBY Trial of EDIT-301 for the Treatment of Severe Sickle Cell Disease", 6 December 2022 (2022-12-06), XP093148117, Retrieved from the Internet <URL:https://ir.editasmedicine.com/node/11366/pdf> [retrieved on 20240404] * |
| BAUER ET AL., NAT. MED., vol. 25, no. 5, 2019, pages 776 - 783 |
| BERNAUDIN ET AL., BLOOD, vol. 110, 2007, pages 2749 - 2756 |
| BOTHMER ET AL., CRISPR J, vol. 3, no. 3, 2020, pages 177 - 187 |
| BOUVA, HEMATOLOGICA, vol. 91, no. 1, 2006, pages 129 - 132 |
| BRINER ET AL., MOL CELL, vol. 56, no. 2, 2014, pages 333 - 339 |
| BROUSSEAU, AM J HEMATOL, vol. 85, no. 1, 2010, pages 77 - 78 |
| CALDECOTT, NAT REV GENET, vol. 9, no. 8, 2008, pages 619 - 631 |
| CHANG ET AL., MOL THER METHODS CLIN DEV, vol. 4, 2017, pages 137 - 148 |
| CHASSANIDIS, ANN HEMATOL, vol. 88, no. 6, 2009, pages 549 - 555 |
| CHYLINSKI ET AL., RNA BIOL, vol. 10, no. 5, 2013, pages 726 - 737 |
| CORNISH-BOWDEN A, NUCLEIC ACIDS RES., vol. 13, no. 9, 10 May 1985 (1985-05-10), pages 3021 - 30 |
| COSTA ET AL., CAD SAUDE PUBLICA, vol. 18, no. 5, 2002, pages 1469 - 1471 |
| DAVISMAIZELS 2, PROC NATL ACAD SCI USA, vol. 111, no. 10, 2014, pages E924 - 932 |
| DE DREUZY E ET AL: "S251: Preclinical data supporting the initiation of the edit-301 phase i ruby clinical trial for the potential treatment of sickle cell disease", HEMASPHERE; 26TH CONGRESS OF THE EUROPEAN HEMATOLOGY ASSOCIATION, EHA 2021; 20210609 TO 20210617; VIRTUAL, WOLTERS KLUWER HEALTH, US, vol. 5, no. SUPPL 2, 1 June 2021 (2021-06-01), pages 84, XP009553216, ISSN: 2572-9241, DOI: 10.1097/HS9.0000000000000566 * |
| DE DREUZY EDOUARD ET AL: "EDIT-301: An Experimental Autologous Cell Therapy Comprising Cas12a-RNP Modified mPB-CD34+ Cells for the Potential Treatment of SCD", BLOOD, AMERICAN SOCIETY OF HEMATOLOGY, US, vol. 134, 13 November 2019 (2019-11-13), pages 4636, XP086663530, ISSN: 0006-4971, DOI: 10.1182/BLOOD-2019-130256 * |
| DE DREUZY EDOUARD ET AL: "Robust Pre-Clinical Results and Large-Scale Manufacturing Process for Edit-301: An Autologous Cell Therapy for the Potential Treatment of SCD", BLOOD, vol. 136, no. Supplement 1, 5 November 2020 (2020-11-05), US, pages 45 - 46, XP055790434, ISSN: 0006-4971, Retrieved from the Internet <URL:https://ashpublications.org/blood/article/136/Supplement%201/45/471184/Robust-Pre-Clinical-Results-and-Large-Scale> DOI: 10.1182/blood-2020-140073 * |
| FINE ET AL., SCI REP., vol. 5, 2015, pages 10777 |
| FRANGOUL HAYDAR ET AL: "AsCas12a Gene Editing of HBG1/2 Promoters with Edit-301 Results in Early and Sustained Normalization of Hemoglobin and Increased Fetal Hemoglobin in Patients with Severe Sickle Cell Disease and Transfusion-Dependent Beta-Thalassemia", TRANSPLANTATION AND CELLULAR THERAPY, ELSEVIER, AMSTERDAM, NL, vol. 30, no. 2, 1 February 2024 (2024-02-01), XP087465368, ISSN: 2666-6367, [retrieved on 20240207], DOI: 10.1016/J.JTCT.2023.12.312 * |
| FRIT ET AL., DNA REPAIR (AMST, vol. 17, 2014, pages 81 - 97 |
| GAO ET AL., NAT BIOTECHNOL., vol. 35, no. 8, 2017, pages 789 - 792 |
| GIANNOUKOS ET AL., BMC GENOMICS, vol. 19, no. 1, 2018, pages 212 |
| GIARRATANA ET AL., BLOOD, vol. 118, 2011, pages 5071 - 5079 |
| GIARRATANA ET AL., NAT BIOTECHNOL., vol. 23, no. 1, 2005, pages 69 - 74 |
| GUILINGER ET AL., NAT BIOTECHNOL, vol. 32, 2014, pages 577 - 582 |
| HEATH J: "EDIT-301: AN AUTOLOGOUS CELL THERAPY TO PROMOTE FETAL HEMOGLOBIN EXPRESSION FOR THE POTENTIAL TREATMENT OF SICKLE CELL DISEASE", HEMASPHERE, vol. 4, no. Suppl. 1, 1 June 2020 (2020-06-01), pages 107, XP055790442, Retrieved from the Internet <URL:https://journals.lww.com/hemasphere/Citation/2020/06001/Abstract_Book__25th_Congress_of_the_European.1.aspx> * |
| HEIGWER ET AL., NAT METHODS, vol. 11, no. 2, 2014, pages 122 - 123 |
| IYAMAWILSON, DNA REPAIR (AMST, vol. 12, no. 8, 2013, pages 620 - 636 |
| JIANG ET AL., NAT BIOTECHNOL, vol. 31, no. 3, 2013, pages 233 - 239 |
| JIAO B ET AL., BLOOD ADV, 2023 |
| JINEK ET AL., SCIENCE, vol. 337, no. 6096, 2012, pages 816 - 821 |
| JINEK ET AL., SCIENCE, vol. 343, no. 6176, 2014, pages 1247997 |
| KATO GJ ET AL., NAT REV DIS PRIMERS, vol. 4, 2018, pages 18010 |
| KLEINSTIVER ET AL., NAT BIOTECHNOL, vol. 33, no. 12, 2015, pages 1293 - 1298 |
| KLEINSTIVER ET AL., NAT BIOTECHNOL, vol. 37, no. 3, 2019, pages 276 - 282 |
| KLEINSTIVER ET AL., NATURE, vol. 523, no. 7561, 2015, pages 481 - 485 |
| KLEINSTIVER ET AL., NATURE, vol. 533, no. 7603, 2016, pages 420 - 424 |
| KOSICKI ET AL., NAT BIOTECHNOL, vol. 36, no. 8, 2018, pages 765 - 771 |
| LEE ET AL., NANO LETT, vol. 12, no. 12, 2012, pages 6322 - 6327 |
| LEWIS, MEDICAL-SURGICAL NURSING: ASSESSMENT AND MANAGEMENT OF CLINICAL PROBLEMS, 2014 |
| LI, CELL RES, vol. 18, no. 1, 2008, pages 85 - 98 |
| MAKAROVA ET AL., NAT REV MICROBIOL, vol. 9, no. 6, 2011, pages 467 - 477 |
| MALI ET AL., SCIENCE, vol. 339, no. 6121, 2013, pages 823 - 826 |
| MANTOVANI ET AL., NUCLEIC ACIDS RES, vol. 16, no. 16, 1988, pages 7783 - 7797 |
| MARTEIJN ET AL., NAT REV MOL CELL BIOL, vol. 15, no. 7, 2014, pages 465 - 481 |
| MARTYN ET AL., BIOCHIM BIOPHYS ACTA, vol. 1860, no. 5, 2017, pages 525 - 536 |
| MASALA, METHODS ENZYMOL., vol. 231, 1994, pages 21 - 44 |
| METAIS ET AL., BLOOD ADV., vol. 3, no. 21, 2019, pages 3379 - 92 |
| METAIS JEAN-YVES ET AL: "CRISPR-Cas9 Genome Editing of [gamma]-Globin Promoters in Human Hematopoietic Stem Cells to Induce Erythrocyte Fetal Hemoglobin for Treatment of [beta]-Hemoglobinopathies", BLOOD, AMERICAN SOCIETY OF HEMATOLOGY, US, vol. 134, 13 November 2019 (2019-11-13), pages 2066, XP086672014, ISSN: 0006-4971, DOI: 10.1182/BLOOD-2019-131318 * |
| NISHIMASU ET AL., CELL, vol. 156, no. 5, 2014, pages 935 - 949 |
| NISHIMASU ET AL., CELL, vol. 162, 2015, pages 1113 - 1126 |
| NOTTA ET AL., SCIENCE, vol. 333, no. 6039, 2011, pages 218 - 21 |
| PAUSCH ET AL., SCIENCE, vol. 369, no. 6501, 2020, pages 333 - 337 |
| RICHARDSON ET AL., NAT BIOTECHNOL, vol. 34, no. 5, 2016, pages 483 - 344 |
| RONGRONG LIU ET AL.: "Preliminary Result of the Safety and Efficacy of Autologous HBG1/2 Promoter-Modified CD34+ Hematopoietic Stem and Progenitor Cells (RM-001) in Transfusion-Dependent [Beta]eta-Thalassemia | Blood | American Society of Hematology", 15 November 2022 (2022-11-15), XP093148044, Retrieved from the Internet <URL:https://ashpublications.org/blood/article/140/Supplement%201/4915/490587/Preliminary-Result-of-the-Safety-and-Efficacy-of> [retrieved on 20240404] * |
| SHMAKOV ET AL., MOLECULAR CELL, vol. 60, no. 3, 2015, pages 385 - 397 |
| STEINBERG ET AL., BLOOD., vol. 123, no. 4, 2014, pages 481 - 485 |
| STERNBERG ET AL., NATURE, vol. 507, no. 7490, 2014, pages 569 - 573 |
| STROHKENDL ET AL., MOL. CELL, vol. 71, 2018, pages 816 - 824 |
| SUPERTI-FURGA ET AL., EMBO J, vol. 7, no. 10, 1988, pages 3099 - 3107 |
| THEIN, HUM MOL GENET, vol. 18, no. R2, 2009, pages R216 - 223 |
| THORPE ET AL., BR J HAEMATOL., vol. 87, no. 1, 1994, pages 125 - 132 |
| WABER ET AL., BLOOD, vol. 67, no. 2, 1986, pages 551 - 554 |
| WALTER ET AL., NEJM, vol. 335, 1996, pages 369 - 76 |
| WANG ET AL., CELL, vol. 153, no. 4, 2013, pages 1380 - 1389 |
| WANG L ET AL: "INITIAL SAFETY AND EFFICACY STUDY OF RM-001, AUTOLOGOUS HBG1/2 PROMOTER-MODIFIED CD34+ HEMATOPOIETIC STEM AND PROGENITOR CELLS, IN TRANSFUSION-DEPENDENT BETATHALASSEMIA", vol. 6, no. Supplement 3, 1 June 2022 (2022-06-01), pages 2569 - 2570, XP009553232, ISSN: 2572-9241, Retrieved from the Internet <URL:https://onlinelibrary.wiley.com/doi/epdf/10.1097/01.HS9.0000852292.38263.b8> * |
| WASTNEDGE E ET AL., J GLOB HEALTH, vol. 8, no. 2, 2018, pages 021103 |
| WEBER ET AL., SCI ADV., vol. 6, no. 7, 2020, pages eaay9392 |
| WILLIAMS TN ET AL., ANNU REV GENOMICS HUM GENET, vol. 19, 2018, pages 113 - 147 |
| XIAO ET AL., BIOINFORMATICS, vol. 30, no. 10, 2014, pages 1180 - 1182 |
| XU ET AL., GENES DEV, vol. 24, no. 8, 2010, pages 783 - 798 |
| YAMANO ET AL., CELL, vol. 165, no. 4, 2016, pages 949 - 962 |
Also Published As
| Publication number | Publication date |
|---|---|
| EP4630554A1 (en) | 2025-10-15 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| US11963982B2 (en) | CRISPR/RNA-guided nuclease systems and methods | |
| US20250179488A1 (en) | Systems and methods for the treatment of hemoglobinopathies | |
| US20240417706A1 (en) | Systems and methods for the treatment of hemoglobinopathies | |
| US20240191197A1 (en) | Modified cells and methods for the treatment of hemoglobinopathies | |
| US20220073951A1 (en) | Systems and methods for the treatment of hemoglobinopathies | |
| US20220047637A1 (en) | Systems and methods for the treatment of hemoglobinopathies | |
| US20250250588A1 (en) | Systems and methods for the treatment of hemoglobinopathies | |
| US20200263206A1 (en) | Targeted integration systems and methods for the treatment of hemoglobinopathies | |
| CA3164055A1 (en) | Modified cells and methods for the treatment of hemoglobinopathies | |
| US20240335476A1 (en) | Systems and methods for the treatment of hemoglobinopathies | |
| WO2024123842A1 (en) | Systems and methods for the treatment of hemoglobinopathies | |
| WO2024254566A2 (en) | Systems and methods for the treatment of hemoglobinopathies | |
| US20260035693A1 (en) | Systems and methods for the treatment of hemoglobinopathies | |
| CN117940566A (en) | Systems and methods for treating hemoglobinopathies |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| 121 | Ep: the epo has been informed by wipo that ep was designated in this application |
Ref document number: 23841142 Country of ref document: EP Kind code of ref document: A1 |
|
| WWE | Wipo information: entry into national phase |
Ref document number: 2023841142 Country of ref document: EP |
|
| NENP | Non-entry into the national phase |
Ref country code: DE |
|
| ENP | Entry into the national phase |
Ref document number: 2023841142 Country of ref document: EP Effective date: 20250707 |
|
| WWP | Wipo information: published in national office |
Ref document number: 2023841142 Country of ref document: EP |