WO2024121558A1 - Enzyme having pepulsol synthase activity - Google Patents
Enzyme having pepulsol synthase activity Download PDFInfo
- Publication number
- WO2024121558A1 WO2024121558A1 PCT/GB2023/053149 GB2023053149W WO2024121558A1 WO 2024121558 A1 WO2024121558 A1 WO 2024121558A1 GB 2023053149 W GB2023053149 W GB 2023053149W WO 2024121558 A1 WO2024121558 A1 WO 2024121558A1
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- sequence
- polypeptide
- peplusol
- nucleic acid
- seq
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Ceased
Links
Classifications
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/63—Introduction of foreign genetic material using vectors; Vectors; Use of hosts therefor; Regulation of expression
- C12N15/79—Vectors or expression systems specially adapted for eukaryotic hosts
- C12N15/82—Vectors or expression systems specially adapted for eukaryotic hosts for plant cells, e.g. plant artificial chromosomes (PACs)
- C12N15/8241—Phenotypically and genetically modified plants via recombinant DNA technology
- C12N15/8242—Phenotypically and genetically modified plants via recombinant DNA technology with non-agronomic quality (output) traits, e.g. for industrial processing; Value added, non-agronomic traits
- C12N15/8243—Phenotypically and genetically modified plants via recombinant DNA technology with non-agronomic quality (output) traits, e.g. for industrial processing; Value added, non-agronomic traits involving biosynthetic or metabolic pathways, i.e. metabolic engineering, e.g. nicotine, caffeine
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/63—Introduction of foreign genetic material using vectors; Vectors; Use of hosts therefor; Regulation of expression
- C12N15/79—Vectors or expression systems specially adapted for eukaryotic hosts
- C12N15/80—Vectors or expression systems specially adapted for eukaryotic hosts for fungi
- C12N15/81—Vectors or expression systems specially adapted for eukaryotic hosts for fungi for yeasts
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N9/00—Enzymes; Proenzymes; Compositions thereof; Processes for preparing, activating, inhibiting, separating or purifying enzymes
- C12N9/10—Transferases (2.)
- C12N9/1085—Transferases (2.) transferring alkyl or aryl groups other than methyl groups (2.5)
Definitions
- the disclosure relates to the isolation and characterisation of a triterpene synthase (e.g., a peplusol synthase), polypeptide isolated from Euphorbia peplus; cells, for example plant cells or microbial cells, transformed with nucleic acid encoding said polypeptide; expression vectors including nucleic acid encoding said polypeptide and methods to produce a triterpene alcohol, for example peplusol.
- a triterpene synthase e.g., a peplusol synthase
- polypeptide isolated from Euphorbia peplus cells, for example plant cells or microbial cells, transformed with nucleic acid encoding said polypeptide
- expression vectors including nucleic acid encoding said polypeptide and methods to produce a triterpene alcohol, for example peplusol.
- Euphorbiaceae is a large family of flowering plants found all over the world, with some synthesising diterpene compounds of considerable biological activity such as ingenol mebutate (Euphorbia peplus), resiniferatoxin (E. resinifera), prostratin (E. cornigera), jatrophanes and lathyranes (Jatropha sp.
- Triterpenes have been isolated from plants belonging to the family of Euphorbiaceae. For example, linear and cyclic triterpenes such as peplusol, cycloartol, lanosterol and others (1). Peplusol is a triterpene alcohol with strong antifungal activities (1) that was previously described as being responsible for the physical properties of Euphorbia peplus latex (2).
- Peplusol like other triterpenoids is thought to be derived from two farnesyl diphosphate (FPP) molecules in a reaction that is similar to the production of the well documented squalene molecule.
- FPP farnesyl diphosphate
- Squalene is used in cosmetics and skin care, was originally sourced from shark liver until recently and is now a target for industrial biotechnology to deliver a more sustainable supply.
- This disclosure relates to a triterpene synthase isolated from E. peplus that produces peplusol when transiently expressed in Nicotiana benthamiana (Figure 1) and Saccharomyces cerevisiae ( Figure 2). Therefore, this enzyme has peplusol synthase activity.
- peplusol is significantly increased when a truncated version of 3-hydroxy-3-methylglutaryl coenzyme A (HMG-CoA) reductase (tHMGR) which catalyzes the conversion of HMG-CoA to mevalonate, is co-expressed in plants with the discovered E. peplus peplusol synthase ( Figure 1).
- HMG-CoA 3-hydroxy-3-methylglutaryl coenzyme A
- tHMGR 3-hydroxy-3-methylglutaryl coenzyme A reductase
- nucleic acid molecule that encodes a polypeptide with peplusol synthase activity
- said nucleic acid molecule comprises or consists of a nucleotide sequence selected from the group consisting of: i) a nucleotide sequence as represented by the sequence in SEQ ID NO: 1 or SEQ ID NO: 2; ii) a nucleotide sequence wherein said sequence is degenerate as a result of the genetic code to the nucleotide sequence defined in (i); iii) a nucleic acid molecule the complementary strand of which hybridizes under stringent hybridization conditions to the sequence in SEQ ID NO: 1 or SEQ ID NO: 2 wherein said nucleic acid molecule encodes a polypeptide with peplusol synthase activity; iv) a nucleotide sequence that encodes a polypeptide comprising an amino acid sequence as represented in SEQ ID NO: 3; v) a
- an isolated nucleic acid molecule that encodes a polypeptide with peplusol synthase activity
- said nucleic acid molecule comprises or consists of a nucleotide sequence selected from the group consisting of: i) a nucleotide sequence as represented by the sequence in SEQ ID NO: 6 or SEQ ID NO: 7; ii) a nucleotide sequence wherein said sequence is degenerate as a result of the genetic code to the nucleotide sequence defined in (i); iii) a nucleic acid molecule the complementary strand of which hybridizes under stringent hybridization conditions to the sequence in SEQ ID NO: 6 or SEQ ID NO: 7 wherein said nucleic acid molecule encodes a polypeptide with peplusol synthase activity; iv) a nucleotide sequence that encodes a polypeptide comprising an amino acid sequence as represented in SEQ ID NO: 8; v) a nucleotide
- Hybridization of a nucleic acid molecule occurs when two complementary nucleic acid molecules undergo an amount of hydrogen bonding to each other.
- the stringency of hybridization can vary according to the environmental conditions surrounding the nucleic acids, the nature of the hybridization method, and the composition and length of the nucleic acid molecules used. Calculations regarding hybridization conditions required for attaining particular degrees of stringency are discussed in Sambrook et al., Molecular Cloning: A Laboratory Manual (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY, 2001); and Tijssen, Laboratory Techniques in Biochemistry and Molecular Biology—Hybridization with Nucleic Acid Probes Part I, Chapter 2 (Elsevier, New York, 1993).
- the T m is the temperature at which 50% of a given strand of a nucleic acid molecule is hybridized to its complementary strand.
- the following is an exemplary set of hybridization conditions and is not limiting: Very High Stringency (allows sequences that share at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to hybridize over the disclosed full-length sequence)
- Hybridization 5x SSC at 65 ⁇ C for 16 hours Wash twice: 2x SSC at room temperature (RT) for 15 minutes each Wash twice: 0.5x SSC at 65 ⁇ C for 20 minutes each High Stringency (allows sequences that share at least 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88% or 89% identity to hybridize over the disclosed full-length sequence)
- Hybridization 5x-6x SSC at 65 ⁇ C-70 ⁇ C for 16-20 hours Wash twice: 2x SSC at RT for 5-20 minutes each Wash twice
- nucleic acid molecule according to the invention in the manufacture of a peplusol synthase.
- an isolated polypeptide selected from the group consisting of: i) a polypeptide comprising or consisting of an amino acid sequence as represented in SEQ ID NO: 3; or ii) a modified polypeptide comprising or consisting of a modified amino acid sequence wherein said polypeptide is modified by addition deletion or substitution of at least one amino acid residue of the sequence presented in SEQ ID NO: 3 and which has peplusol synthase activity.
- an isolated polypeptide selected from the group consisting of: i) a polypeptide comprising or consisting of an amino acid sequence as represented in SEQ ID NO: 8; or ii) a modified polypeptide comprising or consisting of a modified amino acid sequence wherein said polypeptide is modified by addition deletion or substitution of at least one amino acid residue of the sequence presented in SEQ ID NO: 8 and which has peplusol synthase activity.
- a modified polypeptide as herein disclosed may differ in amino acid sequence by one or more substitutions, additions, deletions, truncations that may be present in any combination. Among preferred variants are those that vary from a reference polypeptide by conservative amino acid substitutions.
- substitutions are those that substitute a given amino acid by another amino acid of like characteristics.
- the following non-limiting list of amino acids are considered conservative replacements (similar): a) alanine, serine, and threonine; b) glutamic acid and aspartic acid; c) asparagine and glutamine d) arginine and lysine; e) isoleucine, leucine, methionine and valine and f) phenylalanine, tyrosine and tryptophan.
- Most highly preferred are variants that retain or enhance the same biological function and activity as the reference polypeptide from which it varies.
- the variant polypeptides have at least 50% identity, even more preferably at least 55% identity, still more preferably at least 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95% identity, and at least 99% identity with most or the full-length amino acid sequence illustrated herein.
- the variant polypeptides have at least 74%, 75%, 76%, 77%, 78%, 79%, 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% identity, and at least 99% identity with most or the full-length amino acid sequence illustrated herein.
- a vector comprising a nucleic acid molecule according to the invention.
- said nucleic acid molecule is operably linked to a nucleic acid molecule comprising a promoter sequence.
- said nucleic acid sequence comprising a promoter that confers constitutive, regulated or inducible expression on said peplusol synthase.
- said promoter is a heterologous promoter for expression is a heterologous host cell.
- the nucleic acid molecule in the vector is under the control of, and operably linked to, an appropriate promoter or other regulatory elements for transcription in a host cell such as a microbial, (e.g., bacterial, yeast), or plant cell.
- a host cell such as a microbial, (e.g., bacterial, yeast), or plant cell.
- the vector may be a bi- functional expression vector which functions in multiple hosts.
- promoter is meant a nucleotide sequence upstream from the transcriptional initiation site and which contains all the regulatory regions required for transcription. Suitable promoters include constitutive, tissue-specific, inducible, developmental or other promoters for expression in plant cells comprised in plants depending on design.
- Such promoters include viral, fungal, bacterial, animal and plant-derived promoters capable of functioning in plant cells.
- Constitutive promoters include, for example CaMV 35S promoter (Odell et al. (1985) Nature 313, 9810-812); rice actin (McElroy et al. (1990) Plant Cell 2: 163-171); ubiquitin (Christian et al. (1989) Plant Mol. Biol. 18 (675-689); pEMU (Last et al. (1991) Theor Appl. Genet. 81: 581-588); MAS (Velten et al. (1984) EMBO J. 3. 2723-2730); ALS promoter (U.S. Application Seriel No.
- promoters include those in U.S. Patent Nos. 5,608,149; 5,608,144; 5,604,121; 5,569,597; 5,466,785; 5,399,680, 5,268,463; and 5,608,142, each of which is incorporated by reference.
- Chemical-regulated promoters can be used to modulate the expression of a gene in a plant through the application of an exogenous chemical regulator.
- the promoter may be a chemical-inducible promoter, where application of the chemical induced gene expression, or a chemical-repressible promoter, where application of the chemical represses gene expression.
- Chemical-inducible promoters are known in the art and include, but are not limited to, the maize In2-2 promoter, which is activated by benzenesulfonamide herbicide safeners, the maize GST promoter, which is activated by hydrophobic electrophilic compounds that are used as pre-emergent herbicides, and the tobacco PR-1a promoter, which is activated by salicylic acid.
- Other chemical-regulated promoters of interest include steroid-responsive promoters (see, for example, the glucocorticoid-inducible promoter in Schena et al. (1991) Proc. Natl. Acad. Sci. USA 88: 10421-10425 and McNellis et al. (1998) Plant J.
- tissue-specific promoters include those described by Yamamoto et al. (1997) Plant J. 12(2): 255-265; Kawamata et al. (1997) Plant Cell Physiol. 38(7): 792- 803; Hansen et al. (1997) Mol. Gen. Genet.
- operably linked means joined as part of the same nucleic acid molecule, suitably positioned and oriented for transcription to be initiated from the promoter.
- DNA operably linked to a promoter is "under transcriptional initiation regulation" of the promoter.
- the promoter is a tissue specific promoter, an inducible promoter or a developmentally regulated promoter.
- nucleic acid constructs which operate as plant vectors. Specific procedures and vectors previously used with wide success in plants are described by Guerineau and Mullineaux (1993) (Plant transformation and expression vectors.
- Suitable vectors may include plant viral-derived vectors.
- selectable genetic markers may be included in the construct, such as those that confer selectable phenotypes such as resistance to herbicides (e.g., kanamycin, hygromycin, phosphinotricin, chlorsulfuron, methotrexate, gentamycin, spectinomycin, imidazolinones and glyphosate).
- herbicides e.g., kanamycin, hygromycin, phosphinotricin, chlorsulfuron, methotrexate, gentamycin, spectinomycin, imidazolinones and glyphosate.
- said 3-hydroxy-3-methylglutaryl coenzyme A reductase has a truncation of the N-terminal membrane-binding region.
- HMGCR 3-hydroxy-3-methylglutaryl coenzyme A reductase
- EC 1.1.1.34 The enzyme 3-hydroxy-3-methylglutaryl coenzyme A reductase (HMGCR; EC 1.1.1.34) is known in the art and catalyses the reduction of HMG-CoA to mevalonate.
- the truncation of the N-terminal membrane-binding region of HMGCR is known to remove feedback control of HMGCR activity (20, 21).
- the upregulation of the truncated version of HMGCR results in the increased flux through the mevalonate pathway for the production of farnesyl diphosphate (FPP), which is common precursor for all triterpenes, including peplusol.
- FPP farnesyl diphosphate
- said second nucleic acid molecule encodes a polypeptide with 3-hydroxy-3-methylglutaryl coenzyme A activity and wherein said nucleic acid molecule comprises or consists of a nucleotide sequence selected from the group consisting of: i) a nucleotide sequence as represented by the sequence in SEQ ID NO: 4 ii) a nucleotide sequence wherein said sequence is degenerate as a result of the genetic code to the nucleotide sequence defined in (i); iii) a nucleic acid molecule the complementary strand of which hybridizes under stringent hybridization conditions to the sequence in SEQ ID NO: 4 wherein said nucleic acid molecule encodes a polypeptide with 3-hydroxy-3-methylglutaryl coenzyme A activity; iv) a nucleotide sequence that encodes a polypeptide comprising an amino acid sequence as represented in SEQ ID NO: 5 v) a nucleotide sequence that encodes a
- said cell is a plant cell.
- said cell is a microbial cell; preferably a bacterial or fungal cell [e.g., Saccharomyces cerevisiae].
- a plant comprising a plant cell according to the invention.
- a process for the modification of farnesyl diphosphate comprising: i) providing a transgenic plant cell according to the invention; ii) cultivating said plant cell to produce a transgenic plant; and optionally i) harvesting said transgenic plant, or part thereof.
- said modification results in the production of peplusol.
- a method for the production of peplusol comprising: i) providing a microbial cell according to the invention in culture ii) cultivating the microbial cell under conditions that modifies farnesyl diphosphate to produce peplusol; and optionally iii) isolating said peplusol from the microbial cell or cell culture.
- said microbial cell expresses the truncated form of HMGCR.
- said transgenic cell is a microbial cell; preferably a bacterial or fungal cell [e.g., Saccharomyces cerevisiae].
- microorganisms are grown or cultured in the manner with which the skilled worker is familiar, depending on the host organism.
- a liquid medium comprising a carbon source, usually in the form of sugars, a nitrogen source, usually in the form of organic nitrogen sources such as yeast extract or salts such as ammonium sulfate, trace elements such as salts of iron, manganese and magnesium and, if appropriate, vitamins, at temperatures of between 0°C and 100°C, preferably between 10°C and 60°C, while gassing in oxygen.
- the pH of the liquid medium can either be kept constant, regulated during the culturing period, or not.
- the cultures can be grown batchwise, semi-batchwise or continuously. Nutrients can be provided at the beginning of the fermentation or fed in semi- continuously or continuously.
- the triterpene produced can be isolated from the organisms as described above by processes known to the skilled worker, for example by extraction, distillation, crystallization, if appropriate precipitation with salt, and/or chromatography. To this end, the organisms can advantageously be disrupted beforehand.
- the pH value is advantageously kept between pH 4 and 12, preferably between pH 6 and 9, especially preferably between pH 7 and 8.
- the culture medium to be used must suitably meet the requirements of the strains in question.
- these media which can be employed in accordance with the invention usually comprise one or more carbon sources, nitrogen sources, inorganic salts, vitamins and/or trace elements.
- Preferred carbon sources are sugars, such as mono-, di- or polysaccharides. Examples of carbon sources are glucose, fructose, mannose, galactose, ribose, sorbose, ribulose, lactose, maltose, sucrose, raffinose, starch or cellulose.
- Sugars can also be added to the media via complex compounds such as molasses or other by-products from sugar refining.
- the addition of mixtures of a variety of carbon sources may also be advantageous.
- Other possible carbon sources are oils and fats such as, for example, soya oil, sunflower oil, peanut oil and/or coconut fat, fatty acids such as, for example, palmitic acid, stearic acid and/or linoleic acid, alcohols and/or polyalcohols such as, for example, glycerol, methanol and/or ethanol, and/or organic acids such as, for example, acetic acid and/or lactic acid.
- Nitrogen sources are usually organic or inorganic nitrogen compounds or materials comprising these compounds.
- nitrogen sources comprise ammonia in liquid or gaseous form or ammonium salts such as ammonium sulfate, ammonium chloride, ammonium phosphate, ammonium carbonate or ammonium nitrate, nitrates, urea, amino acids or complex nitrogen sources such as cornsteep liquor, soya meal, soya protein, yeast extract, meat extract and others.
- the nitrogen sources can be used individually or as a mixture.
- Inorganic salt compounds which may be present in the media comprise the chloride, phosphorus and sulfate salts of calcium, magnesium, sodium, cobalt, molybdenum, potassium, manganese, zinc, copper and iron.
- Inorganic sulfur-containing compounds such as, for example, sulfates, sulfites, dithionites, tetrathionates, thiosulfates, sulfides, or else organic sulfur compounds such as mercaptans and thiols may be used as sources of sulfur for the production of sulfur- containing fine chemicals, in particular of methionine.
- Phosphoric acid, potassium dihydrogenphosphate or dipotassium hydrogenphosphate or the corresponding sodium-containing salts may be used as sources of phosphorus.
- Chelating agents may be added to the medium to keep the metal ions in solution.
- Particularly suitable chelating agents comprise dihydroxyphenols such as catechol or protocatechuate and organic acids such as citric acid.
- the fermentation media used according to the invention for culturing microorganisms usually also comprise other growth factors such as vitamins or growth promoters, which include, for example, biotin, riboflavin, thiamine, folic acid, nicotinic acid, panthothenate and pyridoxine.
- growth factors and salts are frequently derived from complex media components such as yeast extract, molasses, cornsteep liquor and the like. It is moreover possible to add suitable precursors to the culture medium.
- the exact composition of the media compounds heavily depends on the particular experiment and is decided upon individually for each specific case. Information on the optimization of media can be found in the textbook "Applied Microbiol. Physiology, A Practical Approach" (Editors P.M. Rhodes, P.F.
- Growth media can also be obtained from commercial suppliers, for example Standard 1 (Merck) or BHI (brain heart infusion, DIFCO) and the like. All media components are sterilized, either by heat (20 min at 1.5 bar and 121°C) or by filter sterilization. The components may be sterilized either together or, if required, separately. All media components may be present at the start of the cultivation or added continuously or batchwise, as desired.
- the culture temperature is normally between 15°C and 45°C, preferably at from 25°C to 40°C, and may be kept constant or may be altered during the experiment.
- the pH of the medium should be in the range from 5 to 8.5, preferably around 7.0.
- the pH for cultivation can be controlled during cultivation by adding basic compounds such as sodium hydroxide, potassium hydroxide, ammonia and aqueous ammonia or acidic compounds such as phosphoric acid or sulfuric acid.
- Foaming can be controlled by employing antifoams such as, for example, fatty acid polyglycol esters.
- suitable substances having a selective effect for example antibiotics. Aerobic conditions are maintained by introducing oxygen or oxygen-containing gas mixtures such as, for example, ambient air into the culture.
- the temperature of the culture is normally 20°C to 45°C and preferably 25°C to 40°C.
- the culture is continued until formation of the desired product is at a maximum. This aim is normally achieved within 10 to 160 hours.
- the fermentation broth can then be processed further.
- the biomass may, according to requirement, be removed completely or partially from the fermentation broth by separation methods such as, for example, centrifugation, filtration, decanting or a combination of these methods or be left completely in said broth. It is advantageous to process the biomass after its separation.
- the fermentation broth can also be thickened or concentrated without separating the cells, using known methods such as, for example, with the aid of a rotary evaporator, thin-film evaporator, falling-film evaporator, by reverse osmosis or by nanofiltration. Finally, this concentrated fermentation broth can be processed to obtain the products present therein.
- EpTS Peplusol Synthase
- EpTS E. peplus truncated Peplusol Synthase
- AtSS A. thaliana Squalene Synthase
- EpTS peplus truncated Peplusol Synthase
- EpTS peplus truncated Peplusol Synthase with four conserved regions in Eukaryotic and Prokaryotic squalene synthases highlighted in blue, two DXXED motifs as the substrate binding sites highlighted in brown and residues involved in NADPH recognition highlighted in green, according to Liu et al. 2014. (top) and the prediction of transmembrane helices in protein sequences using TMHMM - 2.0 service from the closely related homologues selected from Figure 3B (bottom).
- Figure 3B E. peplus truncated Peplusol Synthase (EpTS) nucleotide and amino acid identity to Eukaryotic and Prokaryotic homologous squalene synthases.
- FIG. 3C E. peplus full length Peplusol Synthase (EpTS2) nucleotide and amino acid identity to Eukaryotic and Prokaryotic homologous squalene synthases. Functionally characterised squalene synthases marked with asterisk.
- Figure 4A E.
- EpTS peplus truncated Peplusol Synthase nucleotide sequence
- SEQ ID NO: 3 Figure 4B EpTS amino acid sequence
- Figure 4C EpTS truncated peplusol synthase nucleotide sequence (SEQ ID NO: 2) codon optimised for expression in Saccharomyces cerevisiae.
- HMG-CoA truncated 3-hydroxy-3-methylglutaryl coenzyme A reductase
- DNA precipitated with 0.7%vol isopropanol was captured by spooling it on a glass rod and immediately placed in a tube containing 9.5ml of G2 buffer (800 mM guanidine hydrochloride; 30 mM Tris•Cl, pH 8.0; 30 mM EDTA, pH 8.0; 5% Tween 20; 0.5% Triton X-100), premixed with 19 ⁇ l of RNAse A (100 mg/ ⁇ l) and 200 ⁇ l of Qiagen protease (Cat.No.19155) Samples were mixed by vortexing and incubated at 50 ⁇ C for 30-60 min, agitating occasionally.
- G2 buffer 800 mM guanidine hydrochloride; 30 mM Tris•Cl, pH 8.0; 30 mM EDTA, pH 8.0; 5% Tween 20; 0.5% Triton X-100
- chromatin was fixed in place with formaldehyde in the nucleus and then extracted. Fixed chromatin was digested with DNAse I, chromatin ends were repaired and ligated to a biotinylated bridge adapter followed by proximity ligation of adapter containing ends. After proximity ligation, crosslinks were reversed and the DNA purified. Purified DNA was treated to remove biotin that was not internal to ligated fragments. Sequencing libraries were generated using NEB Next Ultra enzymes and Illumina-compatible adapters. Biotin-containing fragments were isolated using streptavidin beads before PCR enrichment of each library. The library was sequenced on an Illumina HiSeq X platform to produce approximately 30x sequence coverage.
- HiRise used MQ>50 reads for scaffolding.
- 2.3 Scaffolding the Assembly with HiRise The input de novo assembly and Dovetail OmniC library reads were used as input data for HiRise, a software pipeline designed specifically for using proximity ligation data to scaffold genome assemblies 6 .
- Dovetail OmniC library sequences were aligned to the draft input assembly using bwa (https://github.com/lh3/bwa).
- the separations of Dovetail Omni C read pairs mapped within draft scaffolds were analyzed byHiRise to produce a likelihood model for genomic distance between read pairs, and the model was used to identify and break putative misjoins, to score prospective joins, and make joins above a threshold.
- Final HiRise assembly contained 8 contigs with L90 of 29.9Mbps which correspond to 8 chromosomes of E. peplus 7 that covered 99.8% of input 286.827Mbps HiFi assembled sequence. Remaining 0.334 Mbp of the sequence was scattered over 149 contigs.
- 2.4 Ab initio genome annotation AUGUSTUS 8 was used for ab initio gene prediction, using model training based on coding sequences from Amaranthus hypochondriacus, Beta vulgaris, Spinacia oleracea and Arabidopsis thaliana. RNAseq data coming from E.
- peplus roots, leaves, main stems, pods and latex cDNA libraries 9 were mapped onto the 286.827Mbp of HiRise genome assembly described above using Bowtie 2 10 .
- Hints with locations of potential intron–exon boundaries were generated from the alignment files with the software package BAM2 hints in the MAKER package 11 .
- MAKER with AUGUSTUS intron–exon boundary hints provided from RNA-seq and isoform sequencing was then used to predict genes in the HiRise genome assembly. Genes were characterized for their putative function by performing a BLAST search of the peptide sequences against the UniProt database.
- PFAM domains and InterProScan ID were added to the gene models using the scripts provided in the MAKER package.
- Ab initio gene prediction yielded 22,470 genes covering 36,440,672bp of total coding region and average length of 1,621Kbp.
- BUSCO Benchmarking Universal Single-Copy Orthologs
- analysis of predicted genes showed 92.5% complete single-copy-, 0.4% complete duplicated, 1.6% fragmented and 5.5% missing BUSCOs.
- Euphorbia peplus candidate gene cloning and transient gene expression in Nicotiana benthamiana was synthesised using total RNA from 100ng of E. peplus latex or stems total RNA using Superscript II reverse transcriptase (Invitrogen, Carlsbad, CA) and random hexamer primers (Invitrogen, Carlsbad, CA). The open reading frame for each gene was then amplified and inserted into the pEAQ-HT expression vector via In-Fusion cloning tools (TaKaRa bio Inc. Kusatsu, Japan), according to the manufacturer’s protocol using the primers detailed in Table 1. In each instance, a 5’-AAAA-3’ Kozak sequence was included immediately upstream of the start codon.
- EpTS EpTS
- sequence annotated as squalene synthase in the genome assembly described in section 2 aboveE. peplus was used to generate a synthetic fragment containing an open reading frame of EpTS (SEQ ID NO: 1) with 5’- (CTGTATATTCTGCCCAAATTCGCG) and 3’- (CCTTTAACTCTGGTTTCATTAAATT) tails facilitating In-Fusion cloning into pEAQ-HT vector (plus 5’-AAAA-3’ Kozak sequence).
- the synthetic fragment was produced by Integrated DNA Technologies (Leuven, Belgium) and inserted into the pEAQ-HT expression vector via In-Fusion cloning tools as mentioned above.
- the expression vectors were transformed into Agrobacterium tumefaciens LBA4404 using the freeze-thaw method.
- Nicotiana benthamiana leaves were infiltrated by vacuum infiltration, using a vacuum pump to apply negative pressure at -0.9 Bar for 1min with equal mixtures of A. tumefaciens cultures at a final OD 600 nm of 1.0 in infiltration buffer (10 mM MgCl 2 , 200 ⁇ M acetosynringone and 0.015% Silwet L-77).
- AvGFP a visual marker for the gene expression
- Metabolites were eluted at 0.35 mL/min and 40°C using a linear gradient from: 99:1 solvent A: solvent B to 1:99 Solvent A: solvent B over 21min, followed by isocratic 1:99 Solvent A:Solvent B for 3min and isocratic 99:1 Solvent A:Solvent B for 4 min (Solvent A: 10mM Ammonium Formate in Acetonitrile/H20 60:40 + 0.1% Formic Acid, Solvent B: 10mM Ammonium Formate in Acetonitrile/2-Propanol 10:90 + 0.1% Formic Acid).
- Pseudomolecular [M+H]+ ions were detected using a Thermo Fisher LTQ-Orbitrap (ThermoFisher, Hemel Hempstead, UK) mass spectrometer fitted with an atmospheric pressure chemical ionization source operating in positive ionization mode under the control of Xcalibur 2.1 software. Data were acquired over the m/z range 50 - 1200 in FTMS centroid mode with resolution set to 7500. Peplusol was identified via LC-MS using plant-purified, NMR-verified standard, and quantified against PMA internal standard as described before. Squalene was identified via GC-MS using commercial standard (Sigma, cat. No.
- Extract was re-dissolved in methanol and 2 ⁇ L aliquot was injected on an Acquity UPLC system (Waters, Elstree, UK) fitted with a Accucore C30, 2.1mm x 100mm, partical size 2.6 ⁇ m column (Thermo Fisher cat. No. 27826—102130).
- Metabolites were eluted at 0.35 mL/min and 40°C using a linear gradient from: 99:1 solvent A: solvent B to 1:99 Solvent A: solvent B over 21min, followed by isocratic 1:99 Solvent A:Solvent B for 3min and isocratic 99:1 Solvent A:Solvent B for 4 min (Solvent A: 10mM Ammonium Formate in methanol/water 60:40 + 0.1 % Formic acid, Solvent B: 10mM Ammonium Formate in methanol/isopropanol 10:90 + 0.1 % Formic acid) with the same mass spectrometer settings as above. Following dilution series of E.
- peplus-purifed peplusol were run in parallel to create 7- point standard curves: 0, 0.15625, 0.3125, 0.625, 1.25, 2.5 and 5mg/ml.
- Standard samples were run on LC-MS as above.
- Standard curves with linear regression R 2 ⁇ 0.998 were used to calculate amount of peplusol in the plant extracts as presented in Figure X.
- Squalene was extracted using method modified from Reed et al.2017 13 . Plant material ground in a Retsch II homogenizer as above, was extracted in 500 ⁇ L of saponification solution (ethanol:water:KOH, 9:1:1, v:v:w) for 2 hours in 65 0 C with intermittent agitation.
- Dried extract was derivatised using a mixture of pyridine (60 ⁇ L), N-Methyl-N- (trimethylSilyl)TriFluoroAcetamide (MSTFA, 30 ⁇ L) and TriMethylsilyl Chloride (TMS, 1 ⁇ L) for 1h at 50 0 C.1 ⁇ L of the derivaitsed extract was analysed by GC-MS using Agilent 6890 Gas Chromatograph GC, (Agilent Technologies UK Ltd, Cheadle, UK) linked to a LECO Pegasus IV Time of Flight Mass Spectrometer TOF-MS, (LECO Instruments, Stockport, UK).
- MSTFA N-Methyl-N- (trimethylSilyl)TriFluoroAcetamide
- TMS TriMethylsilyl Chloride
- the GC oven was fitted with a Restek Zebron ZB-5HT Inferno TM 30M, capillary column (30m, 0.25-mmID, 0.1 ⁇ m film thickness). Helium carrier gas was used at 1ml/min constant flow and transfer line temperature was set to 250°C. The oven temperature was set at 80°C for 5 min and then increased to 270°C at a rate of 12°C min -1 follwed by increase to 310°C at a rate of 6°C min -1 . Mass spectral data was acquired over the m/z range of 50 to 500 in positive electron ionization mode at -70 eV at acquisition rate of 20 spectra/second and 5min acquisition delay.
- EpTS gene expression in Saccharomyces cerevisiae The codon-optimised Euphorbia Peplusol Synthase (EpTS), Euphorbia Peplusol Synthase2 (EpTS2) and Arabidopsis thaliana Squalene Synthase (AtSS, Gene Bank Accession P53799.1) were synthesised as gBlock DNA fragments from IDT (Integrated DNA Technologies Inc.) with overhangs allowing direct insertion into a PmeI digested modified pBEVY-L vector, without PCR amplification, via In-Fusion cloning tools (TaKaRa bio Inc. Kusatsu, Japan), according to the manufacturer’s protocol.
- IDT Integrated DNA Technologies Inc.
- EpTS2 and AtSS were assembled into a PmeI digested modified pBEVY-L vector (Addgene# 51225) under the Gal10 promoter, plasmid was propagated in Top10 E. coli cells (Invitrogen, USA) and sequence confirmed by Sanger sequencing (Eurofins, Europe).
- the constructs were transformed by lithium acetate 14 method into CEN.PK2-1C wild type strain background (MAT ⁇ ura3-52; trp1-289; leu2-3,112; his3 ⁇ 1; MAL2-8c; SUC2) purchased from EUROSCARF (accession no- 30000B).
- CEN.PK2-1C wild type strain background MAT ⁇ ura3-52; trp1-289; leu2-3,112; his3 ⁇ 1; MAL2-8c; SUC2
- the ethyl acetate fraction was transferred to glass tubes and evaporated using a laboratory evaporator (EZ-2 series, Genevac Ltd.)
- the final pellet was dissolved in 200 ⁇ L of methanol and used for LC-MS analysis as described before.
- 4ml of culture was spun down at 3000g for 5min and 3ml of the media, supernatant was extracted with equal volume of ethyl acetate with PMA internal standard (1.6 ⁇ g/mL), evaporated, re-suspended in 200 ⁇ L of methanol and used for LC-MS analysis as described above.
- Standard curves with linear regression R2 ⁇ 0.998 were used to calculate amount of peplusol in the yeast extracts as presented in Figure 7.
- Squalene was extracted from the cell pellets after spinning down 2mL of cultures for 2min at 20,000g in table-top microcentrifuge using protocol modified from Moses et al., 2014 15 .
- Cell pellets were treated with 0.5ml of saponification solution (25% EtOH, 20% KOH) for 2h at 65 0 C with intermittent agitation.500 ⁇ L of hexane was added and samples were vortexed at 1500rpm for 10min.350 ⁇ L of hexane upper phase was transferred into glass vials and dried down in laboratory evaporator (EZ-2 series, Genevac Ltd.).
- Dried extract was derivatised using a mixture of pyridine (60 ⁇ L), N-Methyl-N- (trimethylSilyl)TriFluoroAcetamide (MSTFA, 30 ⁇ L) and TriMethylsilyl Chloride (TMS, 1 ⁇ L) for 1h at 50 0 C.1 ⁇ L of the derivatised extract was analysed by GC-MS using Agilent 6890 Gas Chromatograph GC, (Agilent Technologies UK Ltd, Cheadle, UK) linked to a LECO Pegasus IV Time of Flight Mass Spectrometer TOF-MS, (LECO Instruments, Stockport, UK).
- MSTFA N-Methyl-N- (trimethylSilyl)TriFluoroAcetamide
- TMS TriMethylsilyl Chloride
- the GC oven was fitted with a Restek Zebron ZB-5HT Inferno TM 30M, capillary column (30m, 0.25-mmID, 0.1 ⁇ m film thickness). Helium carrier gas was used at 1ml/min constant flow and transfer line temperature was set to 250°C. The oven temperature was set at 80°C for 5 min and then increased to 270°C at a rate of 12°C min -1 follwed by increase to 310°C at a rate of 6°C min -1 . Mass spectral data was acquired over the m/z range of 50 to 500 in positive electron ionization mode at -70 eV at acquisition rate of 20 spectra/second and 5min acquisition delay.
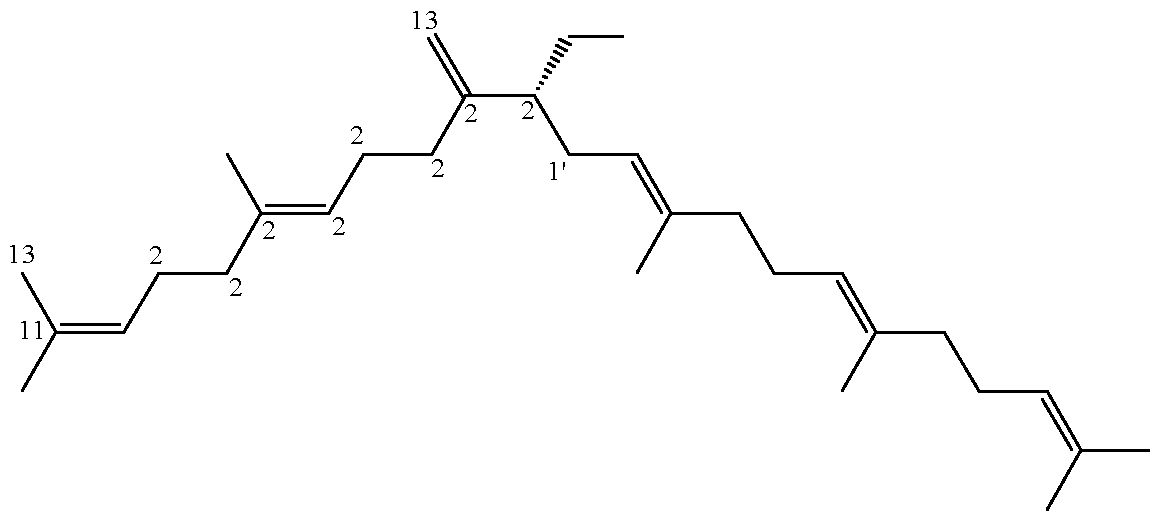
- Peplusol (1) is an unusual linear triterpene alcohol produced by two members of Euphorbia genus: Euphorbia peplus 1 and Euphorbia lateriflora 16 .
- Peplusol content is reaching 5mg/g of Euphorbia peplus latex and is found, albeit in much lower concentrations, in other tissues of the plant, including roots 9
- Previous experiments have shown the moderate antifungal activities of peplusol against agricultural phytopathogenic fungi 17 .
- biosynthetic route to peplusol most likely starts from farnesyl pyrophosphate, a common precursor to all triterpenes.
- Proposed biosynthetic routes involve either: oxidation of squalene, which is suggested by our previous results from gene-silencing in E. peplus 9 , dehydration of a presumed intermediate of squalene synthesis – bisfarensol 18 or alternatively as a direct result of squalene synthase activity.
- Annotation Edit Distance provides a measurement for how well an annotation agrees with overlapping aligned ESTs, mRNA-seq and protein homology data. AED values range from 0 and 1, with 0 denoting perfect agreement of the annotation to aligned evidence, and 1 denoting no evidence support for the annotation. A very low AED scores obtained for most of the 22,470 predicted genes indicated high quality of the E. peplus genome annotation.
- Benchmarking Universal Single-Copy Orthologs (BUSCO) analysis of predicted genes revealed that 94.5% of 255 BUSCO gene searched were found in the annotated HiRise assembly. Analysis of the gene annotations for 158 scaffolds revealed the presence of a single gene annotated as squalene synthase on scaffold 5.
- CDS full length coding sequences
- Transient co-expression of tHMGR has significantly (50-fold) increased production of peplusol reaching 0.16 ⁇ g/mg infiltrated leaf dry weight when compared with infiltrations without tHMGR.
- Squalene was detectable only in samples co-infiltrated with tHMGR and its level was reduced significantly (6.8-fold) when EpTS was co-expressed when compared with empty vector (EV) control.
- EV empty vector
- Reduction in squalene content suggests that EpTS competes with endogenous N. benthamiana squalene synthase for FPP - the common substrate for peplusol and squalene synthesis.
- a second E. peplus genome assembly became publicly available in February 2023.
- Genomic DNA sequence from our own assembly is nearly identical, however, the cDNA annotations differ at the 3' end, which results in the EpTS2 cDNA (Sequence ID No. 6) being 1233bp, which is 60bp longer than the EpTS annotation, and encodes a protein that is 20 amino acids longer (Sequence ID No.8) than the EpTS protein. Differences in the genomic sequence annotations are most likely due to the use of different bioinformatics tools for annotation of the Johnson et al.2023 genome assembly.
- EpTS2 protein sequence contains a full transmembrane domain, understood to be responsible for anchoring squalene synthases in the endoplasmic reticulum.
- EpTS2 is shown to produce peplusol when heterologously expressed in N. benthamiana ( Figure 6) and its peplusol producing activity is very similar to that of EpTS. Peplusol was not detectable either for samples transformed with Arabidopsis thaliana Squalene Synthase 24 (AtSS) or with the Empty Vector.
- Transient co-expression of tHMGR has significantly (15- and 35-fold, respectively) increased production of peplusol for EpTS and EpTS2 which has reached 70 ⁇ g/mg infiltrated leaf dry weight when compared with infiltrations without tHMGR (Figure 6). Squalene was detectable only in samples co- infiltrated with tHMGR and its level was reduced significantly (7- and 12-fold, respectively) when EpTS and EpTS2 were co-expressed when compared with empty vector (EV) control ( Figure 6). Reduction in squalene content was not observed with samples transiently overexpressing AtSS, which suggests that both EpTS and EpTS2 compete with endogenous N.
- Those two constructs and empty vector (EV) control were transformed into haploid wild type CEN.PK-2C background (Y01) as described in Materials and Methods section.
- Three positive clones selected by colony-PCR for each background/construct combination were grown for 72h in liquid cultures as described in Materials and Methods. Extractions were performed on the whole cultures as well as on cell pellets with media separated and peplusol and squalene accumulation was monitored and quantified via GC- and LC-MS techniques as described in Materials and Methods section.
- Peplusol was clearly detectable only in wild type strain Y01 transformed with EpTS and and EpTS2 but not in wild type strain transformed with AtSS or Empty Vector as shown on Figure 2 and Figure 7.
- Peplusol yields were reaching 0.55mg/L when extracted from the whole culture expressing EpTS ( Figure 1) and 0.61 mg/L for strains expressing EpTS2 ( Figure 7).
- Comparison of amount of peplusol released to the media with that retained in the cells indicated that most of the peplusol is retained in the cells ( Figure 2).
- Squalene measurements indicated that AtSS is a functional squalene synthase as its overexpression increases squalene content over 50-fold when compared to empty vector transformed controls ( Figure 2 and Figure 7).
- Example 4 EpTS protein sequence analysis.
- Squalene synthase is a divalent metal-ion-dependent enzyme that catalyzes the two-step reductive ‘head-to-head’ condensation of two molecules of farnesyl pyrophosphate to form squalene using presqualene diphosphate (PSPP) as an intermediate.
- PSPP presqualene diphosphate
- the catalytic mechanism of human squalene synthase has been well described and relies on Mg2 + - mediated condensation of two FPP molecules to form an intermediate with a cyclopropane ring which is followed by NADPH-dependent opening of the cyclopropane ring 23 .
- Functionally characterised plant squalene synthase proteins typically contain two transmembrane domains in the C-terminal region involved in anchoring the protein in the plasma membrane 24 Alignment of cDNA – predicted amino acid sequences for EpTS, its homologues from the Euphorbiaceae family and A. thaliana squalene synthase revealed the EpTS protein has a truncation in the C-terminus. Prediction of transmembrane protein topology with hidden Markov model 25 revealed that the truncated region encodes one of the two predicted transmembrane domains missing in EpTS (Figure 3A).
- EpTS protein sequence alignment also revealed that the two DXXED motifs involved in FPP substrate binding 23 as well as residues involved in NADPH recognition 23 were all conserved in EpTS amino acid sequence (Figure 3A).
- EpTS2 nucleotide sequence is not truncated at the C-terminus and the corresponding protein contains two transmembrane domains which is typical of squalene synthases.
- EpTS2 nucleotide sequence shares between 33 and 79% and EpTS2 amino acid sequence shares between 27 and 73% homology with known Prokaryotic and Eukaryotic squalene synthases respectively (Figure 3C).
- REFERENCES 1 Giner, J. L., Berkowitz, J. D. & Andersson, T. J Nat Prod 63, 267-269, (2000). 2 Loureiro, J., Rodriguez, E., Dolezel, J. & Santos, C. Ann Bot-London 100, 875-888, (2007).
Landscapes
- Health & Medical Sciences (AREA)
- Life Sciences & Earth Sciences (AREA)
- Genetics & Genomics (AREA)
- Engineering & Computer Science (AREA)
- Chemical & Material Sciences (AREA)
- Bioinformatics & Cheminformatics (AREA)
- Organic Chemistry (AREA)
- Zoology (AREA)
- Biotechnology (AREA)
- Wood Science & Technology (AREA)
- Biomedical Technology (AREA)
- General Engineering & Computer Science (AREA)
- Molecular Biology (AREA)
- Biochemistry (AREA)
- Microbiology (AREA)
- General Health & Medical Sciences (AREA)
- Biophysics (AREA)
- Physics & Mathematics (AREA)
- Plant Pathology (AREA)
- Mycology (AREA)
- Medicinal Chemistry (AREA)
- Nutrition Science (AREA)
- Cell Biology (AREA)
- Enzymes And Modification Thereof (AREA)
Abstract
Description
Claims
Priority Applications (1)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| EP23825081.5A EP4630565A1 (en) | 2022-12-07 | 2023-12-06 | Enzyme having peplusol synthase activity |
Applications Claiming Priority (4)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| GBGB2218370.1A GB202218370D0 (en) | 2022-12-07 | 2022-12-07 | Enzyme |
| GB2218370.1 | 2022-12-07 | ||
| GBGB2311804.5A GB202311804D0 (en) | 2023-08-01 | 2023-08-01 | Enzyme |
| GB2311804.5 | 2023-08-01 |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| WO2024121558A1 true WO2024121558A1 (en) | 2024-06-13 |
Family
ID=89222967
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/GB2023/053149 Ceased WO2024121558A1 (en) | 2022-12-07 | 2023-12-06 | Enzyme having pepulsol synthase activity |
Country Status (2)
| Country | Link |
|---|---|
| EP (1) | EP4630565A1 (en) |
| WO (1) | WO2024121558A1 (en) |
Citations (10)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US5268463A (en) | 1986-11-11 | 1993-12-07 | Jefferson Richard A | Plant promoter α-glucuronidase gene construct |
| US5399680A (en) | 1991-05-22 | 1995-03-21 | The Salk Institute For Biological Studies | Rice chitinase promoter |
| US5466785A (en) | 1990-04-12 | 1995-11-14 | Ciba-Geigy Corporation | Tissue-preferential promoters |
| US5569597A (en) | 1985-05-13 | 1996-10-29 | Ciba Geigy Corp. | Methods of inserting viral DNA into plant material |
| US5604121A (en) | 1991-08-27 | 1997-02-18 | Agricultural Genetics Company Limited | Proteins with insecticidal properties against homopteran insects and their use in plant protection |
| US5608149A (en) | 1990-06-18 | 1997-03-04 | Monsanto Company | Enhanced starch biosynthesis in tomatoes |
| US5608144A (en) | 1994-08-12 | 1997-03-04 | Dna Plant Technology Corp. | Plant group 2 promoters and uses thereof |
| US5608142A (en) | 1986-12-03 | 1997-03-04 | Agracetus, Inc. | Insecticidal cotton plants |
| US5789156A (en) | 1993-06-14 | 1998-08-04 | Basf Ag | Tetracycline-regulated transcriptional inhibitors |
| US5814618A (en) | 1993-06-14 | 1998-09-29 | Basf Aktiengesellschaft | Methods for regulating gene expression |
-
2023
- 2023-12-06 EP EP23825081.5A patent/EP4630565A1/en active Pending
- 2023-12-06 WO PCT/GB2023/053149 patent/WO2024121558A1/en not_active Ceased
Patent Citations (10)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US5569597A (en) | 1985-05-13 | 1996-10-29 | Ciba Geigy Corp. | Methods of inserting viral DNA into plant material |
| US5268463A (en) | 1986-11-11 | 1993-12-07 | Jefferson Richard A | Plant promoter α-glucuronidase gene construct |
| US5608142A (en) | 1986-12-03 | 1997-03-04 | Agracetus, Inc. | Insecticidal cotton plants |
| US5466785A (en) | 1990-04-12 | 1995-11-14 | Ciba-Geigy Corporation | Tissue-preferential promoters |
| US5608149A (en) | 1990-06-18 | 1997-03-04 | Monsanto Company | Enhanced starch biosynthesis in tomatoes |
| US5399680A (en) | 1991-05-22 | 1995-03-21 | The Salk Institute For Biological Studies | Rice chitinase promoter |
| US5604121A (en) | 1991-08-27 | 1997-02-18 | Agricultural Genetics Company Limited | Proteins with insecticidal properties against homopteran insects and their use in plant protection |
| US5789156A (en) | 1993-06-14 | 1998-08-04 | Basf Ag | Tetracycline-regulated transcriptional inhibitors |
| US5814618A (en) | 1993-06-14 | 1998-09-29 | Basf Aktiengesellschaft | Methods for regulating gene expression |
| US5608144A (en) | 1994-08-12 | 1997-03-04 | Dna Plant Technology Corp. | Plant group 2 promoters and uses thereof |
Non-Patent Citations (51)
| Title |
|---|
| "Applied Microbiol. Physiology, A Practical Approach", 1997, IRL PRESS, pages: 53 - 73 |
| "Manual of Methods for General Bacteriology", 1981, AMERICAN SOCIETY FOR BACTERIOLOGY |
| CANEVASCNI ET AL., PLANT PHYSIOL., vol. 112, no. 2, 1996, pages 1331 - 1341 |
| CANTAREL, B. L. ET AL., GENOME RES, vol. 18, 2008, pages 188 - 196 |
| CHENG, H.CONCEPCION, G. T.FENG, X.ZHANG, H.LI, H., NAT METHODS, vol. 18, 2021, pages 170 - 175 |
| CHRISTIAN ET AL., PLANT MOL. BIOL., vol. 18, 1989, pages 675 - 689 |
| CZECHOWSKI, T. ET AL., PROC NATL ACAD SCI U S A, vol. 119, 2022, pages 2203890119 |
| DING, C. ET AL., INT J CLIN EXP MED, vol. 8, 2015, pages 12818 - 12825 |
| FAURE, S.CONNOLLY, J. D.FAKUNLE, C. C.PIVA, C., TETRAHEDRON, vol. 56, 2000, pages 9647 - 9653 |
| GATZ ET AL., MOL. GEN. GENET., vol. 227, 1991, pages 229 - 237 |
| GIETZ, R. D.SCHIESTL, R. H., NAT PROTOC, vol. 2, 2007, pages 35 - 37 |
| GINER J-L ET AL: "Nonpolar components of the latex of Euphorbia peplus", JOURNAL OF NATURAL PRODUCTS, AMERICAN CHEMICAL SOCIETY, US, vol. 63, no. 2, 20 January 2000 (2000-01-20), pages 267 - 269, XP002374793, ISSN: 0163-3864, DOI: 10.1021/NP990081G * |
| GINER, J. L.BERKOWITZ, J. D.ANDERSSON, T., J NAT PROD, vol. 63, 2000, pages 267 - 269 |
| GUAN, D. ET AL., BIOINFORMATICS, vol. 36, 2020, pages 2896 - 2898 |
| GUERINEAUMULLINEAUX: "Plant Molecular Biology Labfax", 1993, BIOS SCIENTIFIC PUBLISHERS, article "Plant transformation and expression vectors.", pages: 121 - 148 |
| GUEVARA-GARCIA ET AL., PLANT J., vol. 4, no. 3, 1993, pages 495 - 50 |
| HANSEN ET AL., MOL. GEN. GENET., vol. 254, no. 3, 1997, pages 337 - 343 |
| HIDENOBU UCHIDA ET AL: "Cloning and characterization of a squalene synthase gene from a petroleum plant, Euphorbia tirucalli L", PLANTA ; AN INTERNATIONAL JOURNAL OF PLANT BIOLOGY, SPRINGER, BERLIN, DE, vol. 229, no. 6, 13 March 2009 (2009-03-13), pages 1243 - 1252, XP019715528, ISSN: 1432-2048 * |
| HUA JUAN ET AL: "Chemical profile and defensive function of the latex ofEuphorbia peplus", PHYTOCHEMISTRY, vol. 136, April 2017 (2017-04-01), pages 56 - 64, XP029934517, ISSN: 0031-9422, DOI: 10.1016/J.PHYTOCHEM.2016.12.021 * |
| HUA, J. ET AL., PHYTOCHEMISTRY, vol. 136, 2017, pages 56 - 64 |
| JOHNSON, A. R. ET AL., GENOME BIOL EVOL, vol. 15, 2023 |
| KAWAMATA ET AL., PLANT CELL PHYSIOL., vol. 38, no. 7, 1997, pages 792 - 803 |
| KING, A. J. ET AL., CHEMBIOCHEM, vol. 17, 2016, pages 1593 - 1597 |
| KING, A. J.BROWN, G. D.GILDAY, A. D.LARSON, T. R.GRAHAM, 1. A., PLANT CELL, vol. 26, 2014, pages 3286 - 3298 |
| KROGH, A.LARSSON, B.VON HEIJNE, G.SONNHAMMER, E. L., J MOL BIOL, vol. 305, 2001, pages 567 - 580 |
| LAETSCH, D. R.BLAXTER, M. L., F1000RESEARCH, vol. 6, 2017 |
| LAM, RESULTS PROBL. CELL DIFFER., vol. 20, 1994, pages 181 - 196 |
| LANGMEAD, BSALZBERG, S. L., NAT METHODS, vol. 9, 2012, pages 357 - 359 |
| LAST ET AL., THEOR APPL. GENET., vol. 81, 1991, pages 581 - 588 |
| LIU, C. 1. ET AL., ACTA CRYSTALLOGR D BIOL CRYSTALLOGR, vol. 70, 2014, pages 231 - 241 |
| LOUREIRO, J.RODRIGUEZ, E.DOLEZEL, J.SANTOS, C., ANN BOT-LONDON, vol. 100, 2007, pages 875 - 888 |
| MCELROY ET AL., PLANT CELL, vol. 2, 1990, pages 163 - 171 |
| MCNELLIS ET AL., PLANT J., vol. 14, no. 2, 1998, pages 247 - 257 |
| MOSES, T. ET AL., PROC NATL ACAD SCI U S A, vol. 111, 2014, pages 1634 - 1639 |
| MUTSUOKA ET AL., PROC. NATL. ACAD. SCI. USA, vol. 90, no. 20, 1993, pages 9586 - 9590 |
| NAKASHIMA, T. ET AL., PROC NATL ACAD SCI U S A, vol. 92, 1995, pages 2328 - 2332 |
| NES, W. D.LE, P.VANTAMELEN, E. E.LEOPOLD, E., J. EXP MYCOL, vol. 14, 1990, pages 74 - 77 |
| ODELL ET AL., NATURE, vol. 313, 1985, pages 9810 - 812 |
| OROZCO ET AL., PLANT MOL. BIOL., vol. 23, no. 6, 1993, pages 1129 - 1138 |
| POLAKOWSKI, T.STAHL, U.LANG, C., APPL MICROBIOL BIOT, vol. 49, 1998, pages 66 - 71 |
| PUTNAM, N. H. ET AL., GENOME RES, vol. 26, 2016, pages 342 - 350 |
| REED, J. ET AL., METAB ENG, vol. 42, 2017, pages 185 - 193 |
| RUSSELL ET AL., TRANSGENIC RES., vol. 6, no. 2, 1997, pages 157 - 168 |
| SAINSBURY, F.SAXENA, P.GEISLER, K.OSBOURN, A.LOMONOSSOFF, G. P., METHODS ENZYMOL, vol. 517, 2012, pages 185 - 202 |
| SCHENA ET AL., PROC. NATL. ACAD. SCI. USA, vol. 88, 1991, pages 10421 - 10425 |
| STANKE, M.DIEKHANS, M.BAERTSCH, R.HAUSSLER, D., BIOINFORMATICS, vol. 24, 2008, pages 637 - 644 |
| STERMER, B. A.BIANCHINI, G. M.KORTH, K. L., J LIPID RES, vol. 35, 1994, pages 1133 - 1140 |
| TOMASZ CZECHOWSKI ET AL: "Gene discovery and virus-induced gene silencing revealbranched pathways to major classes of bioactive diterpenoidsinEuphorbia peplus", PROCEEDINGS OF THE NATIONAL ACADEMY OF SCIENCES, vol. 119, no. 21, 1 January 2022 (2022-01-01), XP093119310, ISSN: 0027-8424, Retrieved from the Internet <URL:https://doi.org/10.1073/pnas.2203890119> * |
| VELTEN ET AL., EMBO J., vol. 3, 1984, pages 2723 - 2730 |
| YAMAMOTO ET AL., PLANT CELL PHYSIOL., vol. 35, no. 5, 1994, pages 773 - 778 |
| YAMAMOTO, PLANT J., vol. 12, no. 2, 1997, pages 255 - 265 |
Also Published As
| Publication number | Publication date |
|---|---|
| EP4630565A1 (en) | 2025-10-15 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| AU2019253858B2 (en) | Novel cytochrome p450 fusion protein | |
| Textor et al. | MAM3 catalyzes the formation of all aliphatic glucosinolate chain lengths in Arabidopsis | |
| Baud et al. | Role of WRINKLED1 in the transcriptional regulation of glycolytic and fatty acid biosynthetic genes in Arabidopsis | |
| Han et al. | Effects of overexpression of AaWRKY1 on artemisinin biosynthesis in transgenic Artemisia annua plants | |
| Yoshimoto et al. | Garlic γ-glutamyl transpeptidases that catalyze deglutamylation of biosynthetic intermediate of alliin | |
| EP2257630B1 (en) | Biosynthetic engineering of glucosinolates | |
| US10385354B2 (en) | Papaver somniferum cytochrome P450 | |
| AU2018205190A1 (en) | Genes involved in noscapine production | |
| JP2019533436A (en) | Ciliary process-specific promoter for manipulation of cannabinoids and other compounds in the glandular trichome | |
| Ma et al. | Overexpression of Artemisia annua cinnamyl alcohol dehydrogenase increases lignin and coumarin and reduces artemisinin and other sesquiterpenes | |
| EP2914726B1 (en) | Improved acyltransferase polynucleotides, polypeptides, and methods of use | |
| CN1617880A (en) | Plant cyclopropane fatty acid synthase genes, proteins and uses thereof | |
| Sintupachee et al. | Molecular cloning, bacterial expression and functional characterisation of cytochrome P450 monooxygenase, CYP97C27, and NADPH-cytochrome P450 reductase, CPR I, from Croton stellatopilosus Ohba | |
| US20190218529A1 (en) | Diterpenoid synthesis | |
| Renard et al. | Identification and characterization of thioredoxin h isoforms differentially expressed in germinating seeds of the model legume Medicago truncatula | |
| Liu et al. | Boosting C16 fatty acid biosynthesis of Escherichia coli, yeast and tobacco by tung tree (Vernicia fordii Hemsl.) beta-hydroxyacyl-acyl carrier protein dehydratase gene | |
| CN108795898A (en) | A kind of application of the gene of promotion vegetable seeds flax acid accumulation | |
| EP4630565A1 (en) | Enzyme having peplusol synthase activity | |
| Li et al. | TcWRKY75 participates in pyrethrin biosynthesis by positively regulating the expression of TcCHS, TcAOC, and TcGLIP in Tanacetum cinerariifolium | |
| WO2012085808A1 (en) | Increased avenasterol production | |
| US20050150002A1 (en) | Novel carotenoid hydroxylases for use in engineering carotenoid metabolism in plants | |
| US20080155714A1 (en) | Plant Cyclopropane Fatty Acid Synthase Genes and Uses Thereof | |
| Keyl | The evolution of hydrophobic barriers among land plants | |
| AU2016208335B2 (en) | Plant cytochrome p450 | |
| AU2015200581B2 (en) | Plant cytochrome p450 |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| 121 | Ep: the epo has been informed by wipo that ep was designated in this application |
Ref document number: 23825081 Country of ref document: EP Kind code of ref document: A1 |
|
| WWE | Wipo information: entry into national phase |
Ref document number: 2023825081 Country of ref document: EP |
|
| NENP | Non-entry into the national phase |
Ref country code: DE |
|
| ENP | Entry into the national phase |
Ref document number: 2023825081 Country of ref document: EP Effective date: 20250707 |
|
| WWP | Wipo information: published in national office |
Ref document number: 2023825081 Country of ref document: EP |