WO2024064826A1 - Neutralizing antibodies to plasmodium falciparum circumsporozoite protein and their use - Google Patents
Neutralizing antibodies to plasmodium falciparum circumsporozoite protein and their use Download PDFInfo
- Publication number
- WO2024064826A1 WO2024064826A1 PCT/US2023/074791 US2023074791W WO2024064826A1 WO 2024064826 A1 WO2024064826 A1 WO 2024064826A1 US 2023074791 W US2023074791 W US 2023074791W WO 2024064826 A1 WO2024064826 A1 WO 2024064826A1
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- antibody
- seq
- nos
- antigen binding
- binding fragment
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Ceased
Links
Classifications
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K16/00—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies
- C07K16/18—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans
- C07K16/20—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans from protozoa
- C07K16/205—Plasmodium
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P33/00—Antiparasitic agents
- A61P33/02—Antiprotozoals, e.g. for leishmaniasis, trichomoniasis, toxoplasmosis
- A61P33/06—Antimalarials
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K39/00—Medicinal preparations containing antigens or antibodies
- A61K2039/505—Medicinal preparations containing antigens or antibodies comprising antibodies
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/20—Immunoglobulins specific features characterized by taxonomic origin
- C07K2317/21—Immunoglobulins specific features characterized by taxonomic origin from primates, e.g. man
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/30—Immunoglobulins specific features characterized by aspects of specificity or valency
- C07K2317/33—Crossreactivity, e.g. for species or epitope, or lack of said crossreactivity
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/30—Immunoglobulins specific features characterized by aspects of specificity or valency
- C07K2317/34—Identification of a linear epitope shorter than 20 amino acid residues or of a conformational epitope defined by amino acid residues
Definitions
- Plasmodium species that infect humans is transmitted through the bite of an infected female Anopheles mosquito, which introduces Plasmodium sporozoites into the bloodstream of the human host.
- the major protein on the surface of the infecting P. falciparum sporozoites is the circumsporozoite protein (PfCSP) and provides a major target for antibodies and vaccines.
- the sporozoites rapidly reach the liver where they are sequestered by hepatocytes and undergo asexual expansion. One week later, the infected hepatocytes rupture and release mature parasites, the merozoites. These then begin the erythrocytic phase of malaria by attaching to and invading red blood cells, or erythrocytes.
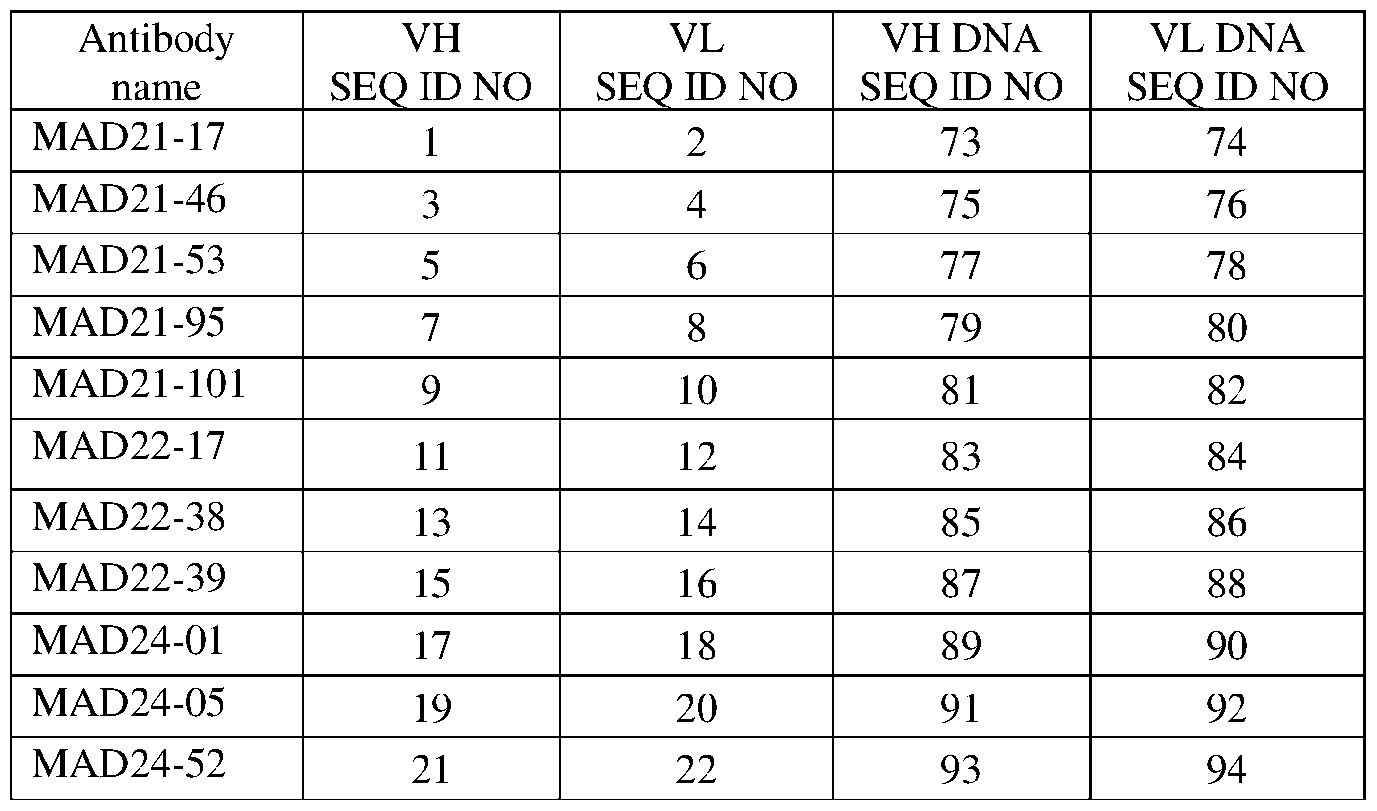
- a monoclonal antibody comprises a heavy chain variable region (V H ) and a light chain variable region (V L ) comprising a heavy chain complementarity determining region (HCDR)1, a HCDR2, and a HCDR3, and a light chain complementarity determining region (LCDR)1, a LCDR2, and a LCDR3 of the VH and VL set forth as SEQ ID NOs: 1 and 2, respectively (MAD21-17), SEQ ID NOs: 3 and 4, respectively (MAD21-46), SEQ ID NOs: 5 and 6, respectively (MAD21-53), SEQ ID NOs: 7 and 8, respectively (MAD21-95), SEQ ID NOs: 9 and 10, respectively (MAD21-101), SEQ ID NOs: 11 and 12, respectively (MAD22-17), SEQ ID NOs: 13 and 14, respectively (MAD22-38), SEQ ID NOs: 15 and 16, respectively (MAD22-39), SEQ ID NOs: 17 and 18, respectively (MAD24-01),
- the monoclonal antibody specifically binds to PfCSP and neutralizes P. falciparum.
- compositions including the antibodies and antigen binding fragments, nucleic acids encoding the antibodies and antigen binding fragments, expression vectors comprising the nucleic acids, and isolated host cells that comprise the nucleic acids.
- the nucleic acid molecule encoding a disclosed antibody or antigen binding fragment can be a cDNA or RNA molecule that encodes the antibody or antigen binding fragment.
- the nucleic acid molecule can be a bicistronic expression construct encoding the VH and VL of the antibody or antigen binding fragment.
- the disclosed antibodies and antigen binding fragments potently neutralize PfCSP expressed on infectious sporozoites in vivo. Accordingly, a method is disclosed for inhibiting (including preventing) P. falciparum infection in a subject.
- the method comprises administering an effective amount (that is, an amount effective to inhibit P. falciparum infection in a subject) of one or more of the disclosed antibodies, antigen binding fragments, nucleic acid molecules, vectors, or compositions, to the subject, such as a subject at risk of or having a P. falciparum infection.
- the antibodies, antigen binding fragments, nucleic acid molecules, vectors, and compositions disclosed herein can be used for a variety of additional purposes, such as for diagnosing P.
- FIG.1 Selected vaccinees show high IgG reactivity to sporozoites after blocking plasma with recombinant PfCSP.
- the data represents polyclonal IgG binding to whole P. falciparum sporozoites after incubation of each plasma sample with recombinant PfCSP.
- FIGs.2A-2B mAbs bind to sporozoites that express Pf CSP. Titration curves of mAb binding to wild-type P.
- FIG.3 Disclosed mAbs recognize an ⁇ 60kDa sporozoite expressed protein. Western blot of P. falciparum sporozoite lysate probed with representative isolated mAbs MAD21-46, MAD22-38 and MAD21-101 as well as known PfCSP mAbs CIS43 and mAb10. The HIV-1 mAb VRC01 was used as a negative control.
- FIGs.4A-4I Disclosed mAbs do not bind canonical PfCSP epitopes.
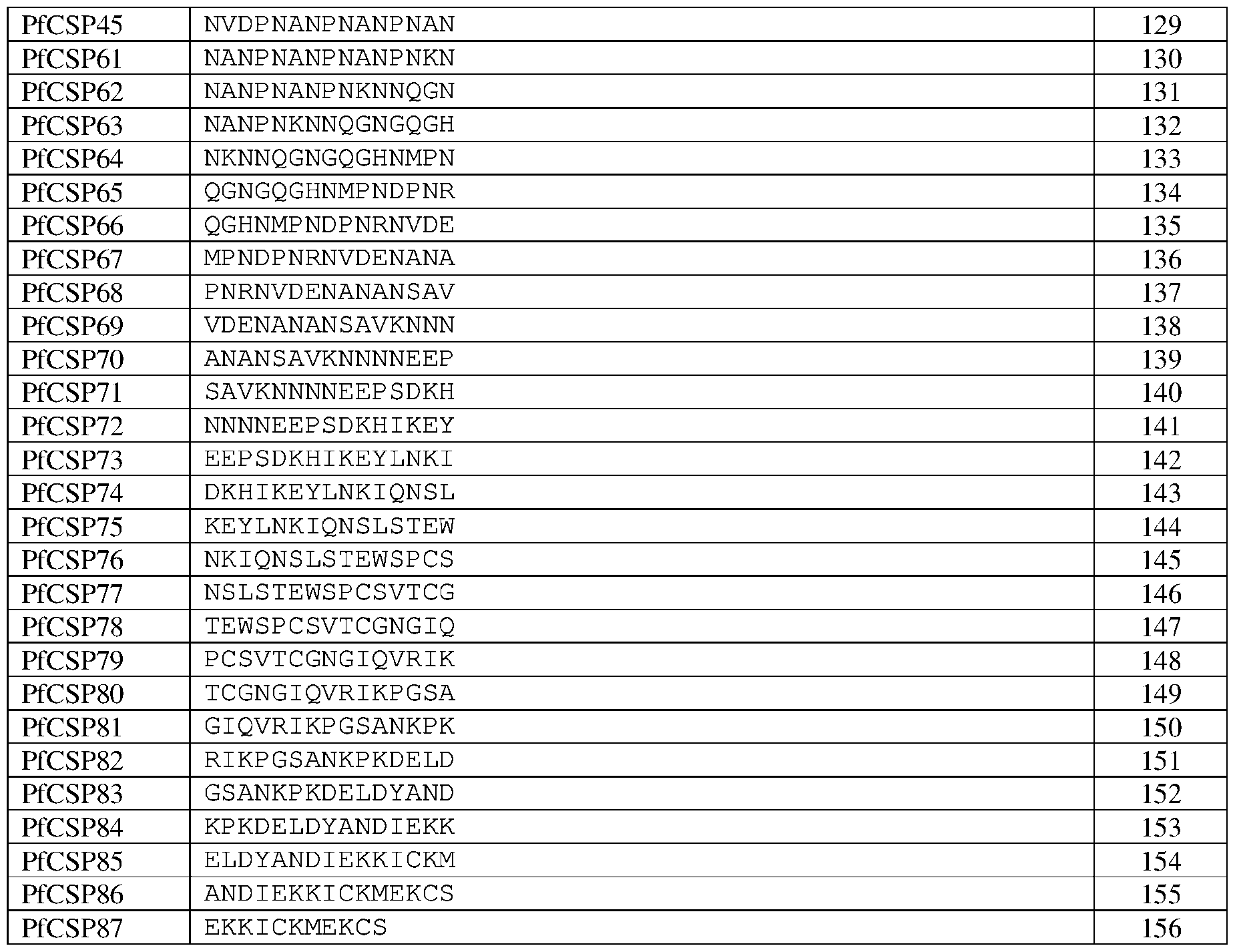
- FIG.5 PfCSP peptide scanning analysis. ELISA analysis of disclosed mAbs binding to a panel of 15-mer overlapping peptides that span the length of PfCSP.
- FIG.6 Disclosed mAbs do not bind recombinant forms of PfCSP.
- FIG.8A-8B The identified mAbs recognize an epitope within the Pf-CSP junctional region and do not show cross-reactivity to the major repeat (NANP) sequences.
- NANP major repeat
- FIG.8A shows the sequences of the PbPfCSP (partial), Pb-PfCSP JRKO (partial), Pb CSP Pf-NANP 12 (partial) and Pb CSP Pf-NANP 4- 5’ (partial) amino acid sequences.
- SEQ ID NOs: 32, 161, 162 and 163 are shown.
- FIG.8B shows results from enzyme linked immunosorbent (ELISA) assays. The findings suggest that the mAb epitope lies within the junction region and that the disclosed mAbs do not cross react with Pf- NANP repeats.
- FIGS.9A-9B The disclosed mAbs recognize a proteolytically processed form of Pf-CSP.
- Binding of the disclosed mAbs to the antigen conjugated beads was measured by flow cytometry in a multiplexed bead-based assay. All mAbs bound to the peptide bearing an N-terminal glutamine which represents the proposed PfCSP N-terminus post-cleavage, and binding was increased towards the peptide bearing an N-terminal pyroglutamic acid. The results evidence that the mAbs target a novel epitope on PfCSP which is dependent upon sporozoite-mediated cleavage and pyroglutamic acid modification of PfCSP. SEQ ID NOs: 164 and 165 are shown.
- SEQ ID NO: 1 is the amino acid sequence of the MAD21-17 VH.
- SEQ ID NO: 2 is the amino acid sequence of the MAD21-17 V L .
- SEQ ID NO: 3 is the amino acid sequence of the MAD21-46 VH.
- SEQ ID NO: 4 is the amino acid sequence of the MAD21-46 VL.
- SEQ ID NO: 5 is the amino acid sequence of the MAD21-53 V H .
- SEQ ID NO: 6 is the amino acid sequence of the MAD21-53 V L .
- SEQ ID NO: 7 is the amino acid sequence of the MAD21-95 V H .
- SEQ ID NO: 8 is the amino acid sequence of the MAD21-95 VL.
- SEQ ID NO: 9 is the amino acid sequence of the MAD21-101 VH.
- SEQ ID NO: 10 is the amino acid sequence of the MAD21-101 VL.
- SEQ ID NO: 11 is the amino acid sequence of the MAD22-17 V H .
- SEQ ID NO: 12 is the amino acid sequence of the MAD22-17 VL.
- SEQ ID NO: 13 is the amino acid sequence of the MAD22-38 VH.
- SEQ ID NO: 14 is the amino acid sequence of the MAD22-38 V L .
- SEQ ID NO: 15 is the amino acid sequence of the MAD22-39 VH.
- SEQ ID NO: 16 is the amino acid sequence of the MAD22-39 V L .
- SEQ ID NO: 17 is the amino acid sequence of the MAD24-01 VH.
- SEQ ID NO: 18 is the amino acid sequence of the MAD24-01 VL.
- SEQ ID NO: 19 is the amino acid sequence of the MAD24-05 V H .
- SEQ ID NO: 20 is the amino acid sequence of the MAD24-05 VL.
- SEQ ID NO: 21 is the amino acid sequence of the MAD24-52 V H .
- SEQ ID NO: 22 is the amino acid sequence of the MAD24-52 V L .
- DIVMTQTPLSSPVTLGQPASISCRSSQSLVHSDGNTYLSWLQQRPGQPPRLLIYKISNRFSGVPDRFSGSGA GTDFTLKISRVEAEDVGVYYCMQATQFPLTFGGGTKVEIK SEQ ID NOs: 23-31 are CDR sequences.
- SEQ ID NO: 32 is a transgenic Plasmodium berghei sporozoite amino acid sequence.
- SEQ ID NOs: 33-67 are CDR sequences.
- SEQ ID NO: 68 is the amino acid sequence of the CIS43 VH.
- SEQ ID NO: 69 is the amino acid sequence of the CIS43 V L .
- SEQ ID NO: 70 is the amino acid sequence of the L9 V H .
- SEQ ID NO: 71 is the amino acid sequence of the L9 VL.
- SEQ ID NO: 72 is an exemplary amino acid sequence for PfCSP (GenBank Acc. No.
- SEQ ID NO: 73 is an exemplary nucleic acid sequence encoding MAD21
- gacatccagatgacccagtctccatcctccctgtctgcatctgtaggagacagagtcaccatcacttgccgggcaaatca cagcattagcagctatttaaattggtatcagcagaaaccagggaaagcccctaagctcctgatctatgctgcatccagtt tgcaaagtggggtcccatcaaggttcagtggcagtggatctgggacagatttcactctcaccatcagcagtctgcaacct gaagattttgcaacttactactgtcaacagagttactataccccatggactttcggccctgggaccaaagtggatatcaa ac SEQ ID NO: 87 is an exemplary nucleic acid sequence encoding MAD22-39 V H .
- gacatccagatgacccagtctccatcctccctgtctgcatctgtaggagacagagtcaccatcacttgccgggcaagtca gagcattatcagctatttaaattggtatcagcagaaaccagggcaagcccctaagctcctgatctatgctgcatccagtt tgcaaagtgggatcccatcaaggttcagtggcagtggatctgggacagatttcactctcaccatcagcagtctgcaacct gaagattttgcaacttactactgtcaacagagttactataccccgtggacgttcggccaagggaccaaggtggaaatcaac SEQ ID NO: 89 is an exemplary nucleic acid sequence encoding MAD24-01 V H .
- Gatattgtgatgacccagactccactctcctcacctgtcacccttggacagccggcctccatctcctgcaggtctagtca aagcctcgtacacagtgatggaaacacctacttgagttggcttcagcagaggccaggccagcctccaagactcctaattt ataagatttctaaccggttctctggggtcccagacagattcagtggggggcagggacagatttcacactgaaaatc agggtggaagctgaggatgtcggggtttattactgcatgcaagctacacaatttcctctcactttcggcggagggac caaggtggagatcaaac SEQ ID NOs: 95-156 are PfCSP peptide sequences
- SEQ ID NO: 157 is the amino acid sequence for the full length heavy chain of MAD21-101.
- SEQ ID NO: 159 is the amino acid sequence of the 317 VH.
- SEQ ID NO: 160 is the amino acid sequence for 317 V L .
- SEQ ID NOs: 161-163 are transgenic Plasmodium berghei sporozoites amino acid sequences.
- SEQ ID Nos: 164-165 are P. falciparum CSP peptide sequences.
- SEQ ID NO: 166 is the amino acid sequence of the 5D5 V H .
- SEQ ID NO: 167 is the amino acid sequence of the 5D5 V L .
- EDLAVYFCQQDYSSPFTFGSGTKLEIK SEQ ID NO: 168 is the amino acid sequence of the mAb10 VH.
- SEQ ID NO: 169 is the amino acid sequence of the mAb10 VL.
- an antigen includes singular or plural antigens and can be considered equivalent to the phrase “at least one antigen.”
- the term “comprises” means “includes.” It is further to be understood that any and all base sizes or amino acid sizes, and all molecular weight or molecular mass values, given for nucleic acids or polypeptides are approximate, and are provided for descriptive purposes, unless otherwise indicated. Although many methods and materials similar or equivalent to those described herein can be used, particular suitable methods and materials are described herein. In case of conflict, the present specification, including explanations of terms, will control.
- 317 Antibody A monoclonal antibody that specifically binds to an epitope on PfCSP and neutralizes malaria infection.
- the CIS317 antibody and methods for its production are described, for example, in Oyen et al. (“Structural basis for antibody recognition of the NANP repeats in Plasmodium falciparum circumsporozoite protein,” Proc Natl Acad Sci USA, 114, E10438-E10445, 2017).
- the amino acid sequences of the heavy and light variable regions of the CIS317 antibody are provided herein as SEQ ID NOs: 159 and 160.
- Administration The introduction of a composition into a subject by a chosen route. Administration can be local or systemic. For example, if the chosen route is intravenous, the composition is administered by introducing the composition into a vein of the subject. Exemplary routes of administration include, but are not limited to, oral, injection (such as subcutaneous, intramuscular, intradermal, intraperitoneal, and intravenous), sublingual, rectal, transdermal (for example, topical), intranasal, vaginal, and inhalation routes.
- Antibody and Antigen Binding Fragment An immunoglobulin, antigen-binding fragment, or derivative thereof, that specifically binds and recognizes an analyte (antigen) such as PfCSP.
- antibody is used herein in the broadest sense and encompasses various antibody structures, including but not limited to monoclonal antibodies, polyclonal antibodies, multispecific antibodies (e.g., bispecific antibodies), and antigen binding fragments, so long as they exhibit the desired antigen-binding activity.
- Non-limiting examples of antibodies include, for example, intact immunoglobulins and variants and fragments thereof that retain binding affinity for the antigen.
- antigen binding fragments include but are not limited to Fv, Fab, Fab', Fab'-SH, F(ab')2; diabodies; linear antibodies; single-chain antibody molecules (e.g. scFv); and multispecific antibodies formed from antibody fragments.
- Antibody fragments include antigen binding fragments either produced by the modification of whole antibodies or those synthesized de novo using recombinant DNA methodologies (see, e.g., Kontermann and Dübel (Eds.), Antibody Engineering, Vols.1-2, 2 nd ed., Springer-Verlag, 2010).
- Antibodies also include genetically engineered forms such as chimeric antibodies (such as humanized murine antibodies) and heteroconjugate antibodies (such as bispecific antibodies).
- An antibody may have one or more binding sites. If there is more than one binding site, the binding sites may be identical to one another or may be different. For instance, a naturally-occurring immunoglobulin has two identical binding sites, a single-chain antibody or Fab fragment has one binding site, while a bispecific or bifunctional antibody has two different binding sites. Typically, a naturally occurring immunoglobulin has heavy (H) chains and light (L) chains interconnected by disulfide bonds. Immunoglobulin genes include the kappa, lambda, alpha, gamma, delta, epsilon and mu constant region genes, as well as the myriad immunoglobulin variable domain genes. There are two types of light chain, lambda ( ⁇ ) and kappa ( ⁇ ).
- Each heavy and light chain contains a constant region (or constant domain) and a variable region (or variable domain).
- the heavy and the light chain variable regions specifically bind the antigen.
- References to “VH” or “VH” refer to the variable region of an antibody heavy chain, including that of an antigen binding fragment, such as Fv, scFv, dsFv or Fab.
- References to “VL” or “VL” refer to the variable domain of an antibody light chain, including that of an Fv, scFv, dsFv or Fab.
- the V H and V L contain a “framework” region interrupted by three hypervariable regions, also called “complementarity-determining regions” or “CDRs” (see, e.g., Kabat et al., Sequences of Proteins of Immunological Interest, 5 th ed., NIH Publication No.91-3242, Public Health Service, National Institutes of Health, U.S. Department of Health and Human Services, 1991).
- CDRs complementarity-determining regions
- amino acid sequence boundaries of a given CDR can be readily determined using any of a number of well-known schemes, including those described by Kabat et al. (Sequences of Proteins of Immunological Interest, 5 th ed., NIH Publication No.91-3242, Public Health Service, National Institutes of Health, U.S. Department of Health and Human Services, 1991; “Kabat” numbering scheme), Al-Lazikani et al., (“Standard conformations for the canonical structures of immunoglobulins,” J. Mol. Bio., 273(4):927-948, 1997; “Chothia” numbering scheme), and Lefranc et al.
- IMGT unique numbering for immunoglobulin and T cell receptor variable domains and Ig superfamily V-like domains Dev. Comp. Immunol., 27(1):55-77, 2003; “IMGT” numbering scheme).
- the CDRs of each chain are typically referred to as CDR1, CDR2, and CDR3 (from the N-terminus to C-terminus), and are also typically identified by the chain in which the particular CDR is located.
- a V H CDR3 is the CDR3 from the V H of the antibody in which it is found
- a V L CDR1 is the CDR1 from the V L of the antibody in which it is found.
- Light chain CDRs are sometimes referred to as LCDR1, LCDR2, and LCDR3.
- Heavy chain CDRs are sometimes referred to as HCDR1, HCDR2, and HCDR3.
- a disclosed antibody includes a heterologous constant domain.
- the antibody includes a constant domain that is different from a native constant domain, such as a constant domain including one or more modifications (such as the “LS” mutations) to increase half-life.
- a “monoclonal antibody” is an antibody obtained from a population of substantially homogeneous antibodies, that is, the individual antibodies comprising the population are identical and/or bind the same epitope, except for possible variant antibodies, for example, containing naturally occurring mutations or arising during production of a monoclonal antibody preparation, such variants generally being present in minor amounts.
- each monoclonal antibody of a monoclonal antibody preparation is directed against a single determinant on an antigen.
- the modifier “monoclonal” indicates the character of the antibody as being obtained from a substantially homogeneous population of antibodies, and is not to be construed as requiring production of the antibody by any particular method.
- the monoclonal antibodies may be made by a variety of techniques, including but not limited to the hybridoma method, recombinant DNA methods, phage-display methods, and methods utilizing transgenic animals containing all or part of the human immunoglobulin loci, such methods and other exemplary methods for making monoclonal antibodies being described herein.
- monoclonal antibodies are isolated from a subject.
- Monoclonal antibodies can have conservative amino acid substitutions which have substantially no effect on antigen binding or other immunoglobulin functions. (See, for example, Greenfield (Ed.), Antibodies: A Laboratory Manual, 2 nd ed.
- a “humanized” antibody or antigen binding fragment includes a human framework region and one or more CDRs from a non-human (such as a mouse, rat, or synthetic) antibody or antigen binding fragment.
- the non-human antibody or antigen binding fragment providing the CDRs is termed a “donor,” and the human antibody or antigen binding fragment providing the framework is termed an “acceptor.”
- all the CDRs are from the donor immunoglobulin in a humanized immunoglobulin. Constant regions need not be present, but if they are, they can be substantially identical to human immunoglobulin constant regions, such as at least about 85-90%, such as about 95% or more identical.
- a “chimeric antibody” is an antibody which includes sequences derived from two different antibodies, which typically are of different species.
- a chimeric antibody includes one or more CDRs and/or framework regions from one human antibody and CDRs and/or framework regions from another human antibody.
- a “fully human antibody” or “human antibody” is an antibody which includes sequences from (or derived from) the human genome, and does not include sequence from another species.
- a human antibody includes CDRs, framework regions, and (if present) an Fc region from (or derived from) the human genome.
- Human antibodies can be identified and isolated using technologies for creating antibodies based on sequences derived from the human genome, for example by phage display or using transgenic animals (see, e.g., Barbas et al. Phage display: A Laboratory Manuel. 1 st Ed. New York: Cold Spring Harbor Laboratory Press, 2004. Print.; Lonberg, Nat. Biotech., 23: 1117-1125, 2005; Lonenberg, Curr. Opin. Immunol., 20:450-459, 2008).
- Antibody or antigen binding fragment that neutralizes P. falciparum An antibody or antigen binding fragment that specifically binds to a P. falciparum antigen (such as PfCSP) in such a way as to inhibit a biological function associated with P.
- PfCSP PfCSP
- an antibody or antigen binding fragment that neutralizes P. falciparum may interfere with the pathogen by binding it in the skin and limiting entry into the blood or entry into the hepatocytes in the liver by interfering with the interaction of the pathogen and one or more cell surface receptors.
- an antibody may interfere with one or more post-attachment interactions of the pathogen with its receptors, for example, by interfering with pathogen internalization by receptor-mediated endocytosis.
- falciparum inhibits sporozoite invasion of hepatocytes, for example, by at least 50% (such as at least 60%, at least 70%, at least 80%, at least 90%, or more) compared to a control antibody or antigen binding fragment.
- an antibody or antigen binding fragment that specifically binds to PfCSP and neutralizes P. falciparum inhibits infection of a human subject by P. falciparum, for example, by at least 50% compared to a control antibody or antigen binding fragment.
- Biological sample A sample obtained from a subject. Biological samples include all clinical samples useful for detection of disease or infection (for example, P.
- Bispecific antibody A recombinant molecule composed of two different antigen binding domains that consequently binds to two different antigenic epitopes.
- Bispecific antibodies include chemically or genetically linked molecules of two antigen-binding domains. The antigen binding domains can be linked using a linker.
- the antigen binding domains can be monoclonal antibodies, antigen-binding fragments (e.g., Fab, scFv), or combinations thereof.
- a bispecific antibody can include one or more constant domains, but does not necessarily include a constant domain.
- Circumsporozoite protein (CSP) The circumsporozoite protein (CSP) is a major malaria parasite surface protein during the sporogonic cycle. PfCSP covers the surface of P. falciparum sporozoites, which are transmitted from the mosquito salivary gland to host hepatocytes.
- An exemplary PfCSP amino acid sequence is provided as SEQ ID NO: 72.
- CIS43 Antibody A monoclonal antibody that specifically binds to an epitope on PfCSP and neutralizes malaria infection.
- the CIS43 antibody and methods for its production are described, for example, in PCT Pub. No. WO 2018/148660.
- the amino acid sequences of the heavy and light variable regions of the CIS43 antibody are provided herein as SEQ ID NOs: 68 and 69.
- Conditions sufficient to form an immune complex Conditions which allow an antibody or antigen binding fragment to bind to its cognate epitope to a detectably greater degree than, and/or to the substantial exclusion of, binding to substantially all other epitopes. Conditions sufficient to form an immune complex are dependent upon the format of the binding reaction and typically are those utilized in immunoassay protocols or those conditions encountered in vivo.
- the conditions employed in the methods are “physiological conditions” which include reference to conditions (e.g., temperature, osmolarity, pH) that are typical inside a living mammal or a mammalian cell. While it is recognized that some organs are subject to extreme conditions, the intra-organismal and intracellular environment normally lies around pH 7 (e.g., from pH 6.0 to pH 8.0, more typically pH 6.5 to 7.5), contains water as the predominant solvent, and exists at a temperature above 0°C and below 50°C.
- physiological conditions e.g., temperature, osmolarity, pH
- Osmolarity is within the range that is supportive of cell viability and proliferation.
- the formation of an immune complex can be detected through conventional methods, for instance immunohistochemistry (IHC), immunoprecipitation (IP), flow cytometry, immunofluorescence microscopy, ELISA, immunoblotting (for example, Western blot), magnetic resonance imaging (MRI), computed tomography (CT) scans, radiography, and affinity chromatography.
- Conjugate A complex of two molecules linked together, for example, linked together by a covalent bond.
- an antibody is linked to an effector molecule; for example, an antibody that specifically binds to CSP from P. falciparum covalently linked to an effector molecule.
- the linkage can be by chemical or recombinant means.
- the linkage is chemical, wherein a reaction between the antibody moiety and the effector molecule has produced a covalent bond formed between the two molecules to form one molecule.
- a peptide linker (short peptide sequence) can optionally be included between the antibody and the effector molecule. Because conjugates can be prepared from two molecules with separate functionalities, such as an antibody and an effector molecule, they are also sometimes referred to as “chimeric molecules.” Conservative variants: “Conservative” amino acid substitutions are those substitutions that do not substantially affect or decrease a function of a protein, such as the ability of the protein to interact with a target protein.
- a PfCSP-specific antibody can include up to 1, 2, 3, 4, 5, 6, 7, 8, 9, or up to 10 conservative substitutions compared to a reference antibody sequence and retain specific binding activity for PfCSP, and/or P. falciparum neutralization activity.
- the term conservative variation also includes the use of a substituted amino acid in place of an unsubstituted parent amino acid. Individual substitutions, deletions or additions which alter, add or delete a single amino acid or a small percentage of amino acids (for instance less than 5%, in some aspects less than 1%) in an encoded sequence are conservative variations where the alterations result in the substitution of an amino acid with a chemically similar amino acid.
- Non-conservative substitutions are those that reduce an activity or function of the PfCSP specific antibody, such as the ability to specifically bind to PfCSP or neutralize P. falciparum.
- Placement in direct physical association includes both in solid and liquid form, which can take place either in vivo or in vitro.
- Contacting includes contact between one molecule and another molecule, for example the amino acid on the surface of one polypeptide, such as an antigen, that contacts another polypeptide, such as an antibody.
- Contacting can also include contacting a cell for example by placing an antibody in direct physical association with a cell.
- Control A reference standard. In some aspects, the control is a negative control, such as sample obtained from a healthy patient not infected with P. falciparum.
- control is a positive control, such as a tissue sample obtained from a patient diagnosed with P. falciparum infection.
- the control is a historical control or standard reference value or range of values (such as a previously tested control sample, such as a group of P. falciparum patients with known prognosis or outcome, or group of samples that represent baseline or normal values).
- a difference between a test sample and a control can be an increase or conversely a decrease.
- the difference can be a qualitative difference or a quantitative difference, for example a statistically significant difference.
- a difference is an increase or decrease, relative to a control, of at least about 5%, such as at least about 10%, at least about 20%, at least about 30%, at least about 40%, at least about 50%, at least about 60%, at least about 70%, at least about 80%, at least about 90%, at least about 100%, at least about 150%, at least about 200%, at least about 250%, at least about 300%, at least about 350%, at least about 400%, or at least about 500%.
- Detectable marker A detectable molecule (also known as a label) that is conjugated directly or indirectly to a second molecule, such as an antibody, to facilitate detection of the second molecule.
- the detectable marker can be capable of detection by ELISA, spectrophotometry, flow cytometry, microscopy or diagnostic imaging techniques (such as CT scans, MRIs, ultrasound, fiberoptic examination, and laparoscopic examination).
- detectable markers include fluorophores, chemiluminescent agents, enzymatic linkages, radioactive isotopes and heavy metals or compounds (for example super paramagnetic iron oxide nanocrystals for detection by MRI).
- Effective amount A quantity of a specific substance sufficient to achieve a desired effect in a subject to whom the substance is administered. For instance, this can be the amount necessary to inhibit a P. falciparum infection, such as the amount necessary to inhibit or prevent P. falciparum sporozoites from invading the liver in the subject or to measurably alter outward symptoms of the P. falciparum infection.
- administration of an effective amount of a disclosed antibody or antigen binding fragment that binds to PfCSP can reduce or inhibit a P.
- falciparum infection for example, as measured by infection of cells, or by number or percentage of subjects infected by the P. falciparum, or by an increase in the survival time of infected subjects, or reduction in symptoms associated with P. falciparum infection
- a desired amount for example by at least 10%, at least 20%, at least 50%, at least 60%, at least 70%, at least 80%, at least 90%, at least 95%, at least 98%, or even at least 100% (elimination or prevention of detectable P. falciparum infection), as compared to a suitable control.
- an effective amount can be determined by varying the dosage and measuring the resulting response, such as, for example, a reduction in pathogen titer. Effective amounts also can be determined through various in vitro, in vivo or in situ immunoassays. An effective amount encompasses a fractional dose that contributes in combination with previous or subsequent administrations to attaining an effective response. For example, an effective amount of an agent can be administered in a single dose, or in several doses, for example daily, during a course of treatment lasting several days or weeks. However, the effective amount can depend on the subject being treated, the severity and type of the condition being treated, and the manner of administration.
- a unit dosage form of the agent can be packaged in an amount, or in multiples of the effective amount, for example, in a vial (e.g., with a pierceable lid) or syringe having sterile components.
- Effector molecule A molecule intended to have or produce a desired effect; for example, a desired effect on a cell to which the effector molecule is targeted. Effector molecules can include, for example, polypeptides and small molecules. In one non-limiting example, the effector molecule is a toxin. Some effector molecules may have or produce more than one desired effect.
- Epitope An antigenic determinant. These are particular chemical groups or peptide sequences on a molecule that are antigenic, i.e.
- an antibody specifically binds a particular antigenic epitope on a polypeptide. In some examples a disclosed antibody specifically binds to an epitope on CSP from P. falciparum.
- Expression Transcription or translation of a nucleic acid sequence.
- an encoding nucleic acid sequence (such as a gene) can be expressed when its DNA is transcribed into RNA or an RNA fragment, which in some examples is processed to become mRNA.
- An encoding nucleic acid sequence (such as a gene) may also be expressed when its mRNA is translated into an amino acid sequence, such as a protein or a protein fragment.
- a heterologous gene is expressed when it is transcribed into an RNA.
- a heterologous gene is expressed when its RNA is translated into an amino acid sequence.
- Regulation of expression can include controls on transcription, translation, RNA transport and processing, degradation of intermediary molecules such as mRNA, or through activation, inactivation, compartmentalization or degradation of specific protein molecules after they are produced.
- Expression Control Sequences Nucleic acid sequences that regulate the expression of a heterologous nucleic acid sequence to which it is operatively linked. Expression control sequences are operatively linked to a nucleic acid sequence when the expression control sequences control and regulate the transcription and, as appropriate, translation of the nucleic acid sequence.
- expression control sequences can include appropriate promoters, enhancers, transcriptional terminators, a start codon (ATG) in front of a protein-encoding gene, splice signals for introns, maintenance of the correct reading frame of that gene to permit proper translation of mRNA, and stop codons.
- control sequences is intended to include, at a minimum, components whose presence can influence expression, and can also include additional components whose presence is advantageous, for example, leader sequences and fusion partner sequences.
- Expression control sequences can include a promoter.
- Expression vector A vector comprising a recombinant polynucleotide comprising expression control sequences operatively linked to a nucleotide sequence to be expressed.
- An expression vector comprises sufficient cis- acting elements for expression; other elements for expression can be supplied by the host cell or in an in vitro expression system.
- expression vectors include cosmids, plasmids (e.g., naked or contained in liposomes) and viruses (e.g., lentiviruses, retroviruses, adenoviruses, and adeno-associated viruses) that incorporate the recombinant polynucleotide.
- a polynucleotide can be inserted into an expression vector that contains a promoter sequence which facilitates the efficient transcription of the inserted genetic sequence of the host.
- Fc region The constant region of an antibody excluding the first heavy chain constant domain. Fc region generally refers to the last two heavy chain constant domains of IgA, IgD, and IgG, and the last three heavy chain constant domains of IgE and IgM. An Fc region may also include part or all of the flexible hinge N-terminal to these domains. For IgA and IgM, an Fc region may or may not include the tailpiece, and may or may not be bound by the J chain.
- the Fc region is typically understood to include immunoglobulin domains C ⁇ 2 and C ⁇ 3 and optionally the lower part of the hinge between C ⁇ 1 and C ⁇ 2. Although the boundaries of the Fc region may vary, the human IgG heavy chain Fc region is usually defined to include residues following C226 or P230 to the Fc carboxyl-terminus, wherein the numbering is according to Kabat.
- the Fc region includes immunoglobulin domains C ⁇ 2 and C ⁇ 3 and optionally the lower part of the hinge between C ⁇ 1 and C ⁇ 2.
- Host cell Cells in which a vector can be propagated and its DNA expressed. The cell may be prokaryotic or eukaryotic. The term also includes any progeny of the subject host cell.
- IgA A polypeptide belonging to the class of antibodies that are substantially encoded by a recognized immunoglobulin alpha gene. In humans, this class or isotype comprises IgA1 and IgA2. IgA antibodies can exist as monomers, polymers (referred to as pIgA) of predominantly dimeric form, and secretory IgA. The constant chain of wild-type IgA contains an 18-amino-acid extension at its C-terminus called the tail piece (tp).
- IgG A polypeptide belonging to the class or isotype of antibodies that are substantially encoded by a recognized immunoglobulin gamma gene. In humans, this class comprises IgG1, IgG2, IgG3, and IgG4.
- Immune complex The binding of antibody or antigen binding fragment (such as a scFv) to a soluble antigen forms an immune complex.
- an immune complex can be detected through conventional methods, for instance immunohistochemistry, immunoprecipitation, flow cytometry, immunofluorescence microscopy, ELISA, immunoblotting (for example, Western blot), magnetic resonance imaging, CT scans, radiography, and affinity chromatography.
- Inhibiting a disease or condition Reducing the full development of a disease or condition in a subject, for example, reducing the full development of a P. falciparum infection in a subject who is at risk of a P. falciparum infection. This includes neutralizing, antagonizing, prohibiting, preventing, restraining, slowing, disrupting, stopping, or reversing progression or severity of the disease or condition.
- inhibiting a disease or condition refers to a prophylactic intervention administered before the disease or condition has begun to develop (for example a treatment initiated in a subject at risk of P. falciparum infection, but not infected by P. falciparum) that reduces subsequent development of the disease or condition and/or ameliorates a sign or symptom of the disease or condition following development.
- the term “ameliorating,” with reference to inhibiting a disease or condition refers to any observable beneficial effect of the prophylactic intervention intended to inhibit the disease or condition.
- the beneficial effect can be evidenced, for example, by a delayed onset of clinical symptoms of the disease or condition in a susceptible subject, a reduction in severity of some or all clinical symptoms of the disease or condition, a slower progression of the disease or condition, an improvement in the overall health or well- being of the subject, a reduction in infection, or by other parameters that are specific to the particular disease or condition.
- the disclosed PfCSP-specific antibodies and antigen binding fragments inhibit the invasion of Plasmodium falciparum sporozoites into human liver cells (hepatocytes).
- the invasion of liver cells is a key event in the infection of a subject with the malaria parasite. Inhibition of the invasion of human liver cells can be measured by one or more of several standard assays (see, for example, Example 1).
- the disclosed PfCSP-specific antibodies and antigen binding fragments can inhibit the invasion of Plasmodium falciparum sporozoites into human liver cells by at least 25%, such as at least 50%, at least 75%, at least 90%, at least 95%, or 100% compared to a suitable control.
- the disclosed PfCSP-specific antibodies and antigen binding fragments inhibit the growth of Plasmodium falciparum in a subject, for example, the antibodies and antigen binding fragments inhibit the multiplication of Plasmodium falciparum in the subject, resulting in a reduction in pathogen load in the subject compared to a relevant control.
- the disclosed PfCSP-specific antibodies and antigen binding fragments can inhibit the growth of Plasmodium falciparum in a subject by at least 25%, such as at least 50%, at least 75%, at least 90%, at least 95%, or 100% compared to a suitable control.
- Isolated A biological component (such as a nucleic acid, peptide, protein or protein complex, for example an antibody) that has been substantially separated, produced apart from, or purified away from other biological components in the cell of the organism in which the component naturally occurs, that is, other chromosomal and extra-chromosomal DNA and RNA, and proteins.
- isolated nucleic acids, peptides and proteins include nucleic acids and proteins purified by standard purification methods.
- nucleic acids, peptides and proteins prepared by recombinant expression in a host cell, as well as, chemically synthesized nucleic acids.
- An isolated nucleic acid, peptide or protein, for example an antibody, can be at least 50%, at least 60%, at least 70%, at least 80%, at least 90%, at least 95%, at least 96%, at least 97%, at least 98%, or at least 99% pure.
- Kabat position A position of a residue in an amino acid sequence that follows the numbering convention delineated by Kabat et al.
- L9 Antibody A monoclonal antibody that specifically binds to an epitope on PfCSP and neutralizes malaria infection. The L9 antibody and methods for its production are described, for example, in PCT Pub. No. WO 2020/227228.
- the amino acid sequences of the heavy and light variable regions of the L9 antibody are provided herein as SEQ ID NOs: 70 and 71.
- Linker A bi-functional molecule that can be used to link two molecules into one contiguous molecule, for example, to link an effector molecule to an antibody.
- Non-limiting examples of peptide linkers include glycine-serine linkers.
- conjugating,” “joining,” “bonding,” or “linking” can refer to making two molecules into one contiguous molecule; for example, linking two polypeptides into one contiguous polypeptide, or covalently attaching an effector molecule or detectable marker radionuclide or other molecule to a polypeptide, such as an scFv.
- the linkage can be either by chemical or recombinant means.
- “Chemical means” refers to a reaction between the antibody moiety and the effector molecule such that there is a covalent bond formed between the two molecules to form one molecule.
- mAb 5D5 Antibody A control monoclonal antibody that specifically binds to an epitope on PfCSP that is used as a control herein. This antibody detects only the un-cleaved form of PfCSP.
- the mAb 5D5 antibody is described, for example, on the internet rcsb.org/structure/6UUD, as available on September 18, 2023.
- the amino acid sequences of the heavy and light variable regions of the mAb 5D5 are provided as SEQ ID NOs: 166 and 167.
- mAb10 Antibody A control monoclonal antibody that specifically binds to an epitope on PfCSP.
- This antibody detects both the cleaved (lower molecular weight) and un-cleaved (higher molecular weight) forms of PfCSP.
- the mAb10 antibody is described, for example, in Kisalu et al., Nat. Med.24, 408-416. 10.1038/nm.4512, 2018 and PCT Publication No. WO 2018/148660.
- the amino acid sequences of the heavy and light variable regions of the mAb10 are provided as SEQ ID NOs: 168 and 169.
- Malaria Malaria is a parasitic infection of humans by the Plasmodium species P. falciparum, P. vivax, P. ovale, P. malariae, and P. knowlesi.
- Infection begins when malaria sporozoites gain access to or are directly injected into the bloodstream of a host by a mosquito. After injection, they migrate to the liver and multiply in hepatocytes for one week. The sporozoites substantially expand in the liver and differentiate to merozoites which are released from the liver into the blood stream, where they infect erythrocytes.
- a schizont is the stage when nuclear division occurs to form individual merozoites which are released to invade other red cells. Malaria clinical symptoms appear during the blood-stage. After several schizogonic cycles, some parasites, instead of becoming schizonts through asexual reproduction, develop into large uninucleate parasites, known as gametocytes. These gametocytes are the sexual blood cell stage forms of the parasite. Sexual development of the malaria parasites involves the female macrogametocyte and the male microgametocyte.
- a mosquito feeds on the blood of an infected host, it can ingest gametocytes within the blood. Fertilization and sexual recombination of the parasite occurs in the mosquito's gut.
- the fertilized parasite which is known as a zygote, then develops into an ookinete.
- the ookinete penetrates the midgut wall of the mosquito and develops into an oocyst, within which many small sporozoites form. When the oocyst ruptures, the sporozoites migrate to the salivary gland of the mosquito via the hemolymph. Once in the saliva of the mosquito, the parasite can be injected into a host, repeating the life cycle.
- Nucleic acid (molecule or sequence): A deoxyribonucleotide or ribonucleotide polymer or combination thereof including without limitation, cDNA, mRNA, genomic DNA, and synthetic (such as chemically synthesized) DNA or RNA.
- the nucleic acid can be double stranded (ds) or single stranded (ss). Where single stranded, the nucleic acid can be the sense strand or the antisense strand.
- Nucleic acids can include natural nucleotides (such as A, T/U, C, and G), and can include analogs of natural nucleotides, such as labeled nucleotides.
- cDNA refers to a DNA that is complementary or identical to an mRNA, in either single stranded or double stranded form.
- Encoding refers to the inherent property of specific sequences of nucleotides in a polynucleotide, such as a gene, a cDNA, or an mRNA, to serve as templates for synthesis of other polymers and macromolecules in biological processes having either a defined sequence of nucleotides (i.e., rRNA, tRNA and mRNA) or a defined sequence of amino acids and the biological properties resulting therefrom.
- a gene encodes a protein if transcription and translation of mRNA produced by that gene produces the protein in a cell or other biological system.
- coding strand the nucleotide sequence of which is identical to the mRNA sequence and is usually provided in sequence listings
- non-coding strand used as the template for transcription
- a “nucleotide sequence encoding an amino acid sequence” includes all nucleotide sequences that are degenerate versions of each other and that encode the same amino acid sequence. Nucleotide sequences that encode proteins and RNA may include introns.
- a first nucleic acid sequence is operably linked with a second nucleic acid sequence when the first nucleic acid sequence is placed in a functional relationship with the second nucleic acid sequence.
- a promoter such as the CMV promoter
- operably linked DNA sequences are contiguous and, where necessary to join two protein-coding regions, in the same reading frame.
- Pharmaceutically acceptable carriers The pharmaceutically acceptable carriers of use are conventional. Remington: The Science and Practice of Pharmacy, 22 nd ed., London, UK: Pharmaceutical Press, 2013, describes compositions and formulations suitable for pharmaceutical delivery of the disclosed agents.
- parenteral formulations usually include injectable fluids that include pharmaceutically and physiologically acceptable fluids such as water, physiological saline, balanced salt solutions, aqueous dextrose, glycerol or the like as a vehicle.
- injectable fluids such as water, physiological saline, balanced salt solutions, aqueous dextrose, glycerol or the like as a vehicle.
- physiologically acceptable fluids such as water, physiological saline, balanced salt solutions, aqueous dextrose, glycerol or the like as a vehicle.
- solid compositions e.g., powder, pill, tablet, or capsule forms
- conventional non-toxic solid carriers can include, for example, pharmaceutical grades of mannitol, lactose, starch, or magnesium stearate.
- compositions to be administered can contain minor amounts of non-toxic auxiliary substances, such as wetting or emulsifying agents, added preservatives (such as non-natural preservatives), and pH buffering agents and the like, for example sodium acetate or sorbitan monolaurate.
- the pharmaceutically acceptable carrier is sterile and suitable for parenteral administration to a subject for example, by injection.
- the active agent and pharmaceutically acceptable carrier are provided in a unit dosage form such as a pill or in a selected quantity in a vial. Unit dosage forms can include one dosage or multiple dosages (for example, in a vial from which metered dosages of the agents can selectively be dispensed).
- Polypeptide A polymer in which the monomers are amino acid residues that are joined together through amide bonds. When the amino acids are alpha-amino acids, either the L-optical isomer or the D- optical isomer can be used, the L-isomers being preferred.
- the terms “polypeptide” or “protein” as used herein are intended to encompass any amino acid sequence and include modified sequences such as glycoproteins.
- a polypeptide includes both naturally occurring proteins, as well as those that are recombinantly or synthetically produced.
- a polypeptide has an amino terminal (N-terminal) end and a carboxy-terminal end. In some aspects, the polypeptide is a disclosed antibody or a fragment thereof.
- a purified peptide preparation is one in which the peptide or protein (such as an antibody) is more enriched than the peptide or protein is in its natural environment within a cell.
- a preparation is purified such that the protein or peptide represents at least 50% of the total peptide or protein content of the preparation.
- a recombinant nucleic acid is one that has a sequence that is not naturally occurring or has a sequence that is made by an artificial combination of two otherwise separated segments of sequence. This artificial combination can be accomplished by chemical synthesis or, more commonly, by the artificial manipulation of isolated segments of nucleic acids, for example, by genetic engineering techniques.
- a recombinant protein is one that has a sequence that is not naturally occurring or has a sequence that is made by an artificial combination of two otherwise separated segments of sequence.
- a recombinant protein is encoded by a heterologous (for example, recombinant) nucleic acid that has been introduced into a host cell, such as a bacterial or eukaryotic cell.
- the nucleic acid can be introduced, for example, on an expression vector having signals capable of expressing the protein encoded by the introduced nucleic acid or the nucleic acid can be integrated into the host cell chromosome.
- Sequence identity The identity between two or more nucleic acid sequences, or two or more amino acid sequences, is expressed in terms of the identity between the sequences.
- Sequence identity can be measured in terms of percentage identity; the higher the percentage, the more identical the sequences.
- Homologs and variants of a VL or a VH of an antibody that specifically binds a target antigen are typically characterized by possession of at least about 75% sequence identity, for example at least about 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% sequence identity counted over the full-length alignment with the amino acid sequence of interest. Any suitable method may be used to align sequences for comparison. Non-limiting examples of programs and alignment algorithms are described in: Smith and Waterman, Adv. Appl. Math.2(4):482-489, 1981; Needleman and Wunsch, J. Mol.
- the NCBI Basic Local Alignment Search Tool (BLAST) (Altschul et al., J. Mol. Biol.215(3):403-410, 1990) is available from several sources, including the National Center for Biological Information and on the Internet, for use in connection with the sequence analysis programs blastp, blastn, blastx, tblastn, and tblastx. Blastn is used to compare nucleic acid sequences, while blastp is used to compare amino acid sequences. Additional information can be found at the NCBI web site. Generally, once two sequences are aligned, the number of matches is determined by counting the number of positions where an identical nucleotide or amino acid residue is present in both sequences.
- BLAST Basic Local Alignment Search Tool
- the percent sequence identity between the two sequences is determined by dividing the number of matches either by the length of the sequence set forth in the identified sequence, or by an articulated length (such as 100 consecutive nucleotides or amino acid residues from a sequence set forth in an identified sequence), followed by multiplying the resulting value by 100.
- bind When referring to an antibody or antigen binding fragment, refers to a binding reaction which determines the presence of a target protein in the presence of a heterogeneous population of proteins and other biologics.
- an antibody binds preferentially to a particular target protein, peptide or polysaccharide (such as an antigen present on the surface of a pathogen, for example PfCSP) and does not bind in a significant amount to other proteins present in the sample or subject.
- Specific binding can be determined by standard methods. See Harlow & Lane, Antibodies, A Laboratory Manual, 2 nd ed., Cold Spring Harbor Publications, New York (2013), for a description of immunoassay formats and conditions that can be used to determine specific immunoreactivity.
- K D refers to the dissociation constant for a given interaction, such as a polypeptide ligand interaction or an antibody antigen interaction.
- K D refers to the concentration of the individual components of the bimolecular interaction divided by the concentration of the complex.
- An antibody that specifically binds to an epitope on PfCSP is an antibody that binds substantially to PfCSP, including cells or tissue expressing PfCSP, substrate to which the PfCSP is attached, or PfCSP in a biological specimen. It is, of course, recognized that a certain degree of non-specific interaction may occur between an antibody and a non-target (such as a cell that does not express PfCSP). Typically, specific binding results in a much stronger association between the antibody and protein or cells bearing the antigen than between the antibody and protein or cells lacking the antigen.
- Specific binding typically results in greater than 2-fold, such as greater than 5-fold, greater than 10-fold, or greater than 100-fold increase in amount of bound antibody (per unit time) to a protein including the epitope or cell or tissue expressing the target epitope as compared to a protein or cell or tissue lacking this epitope.
- Specific binding to a protein under such conditions requires an antibody that is selected for its specificity for a particular protein.
- a variety of immunoassay formats are appropriate for selecting antibodies or other ligands specifically immunoreactive with a particular protein.
- solid-phase ELISA immunoassays are routinely used to select monoclonal antibodies specifically immunoreactive with a protein.
- Subject Living multi-cellular vertebrate organisms, a category that includes human and non- human mammals.
- a subject is a human.
- a subject is selected that is in need of inhibiting a P. falciparum infection.
- the subject is uninfected and at risk of P. falciparum infection.
- Transformed A transformed cell is a cell into which a nucleic acid molecule has been introduced by molecular biology techniques.
- the term transformed and the like encompasses all techniques by which a nucleic acid molecule might be introduced into such a cell, including transduction with viral vectors, transformation with plasmid vectors, and introduction of DNA by electroporation, lipofection, and particle gun acceleration.
- Vector An entity containing a nucleic acid molecule (such as a DNA or RNA molecule) bearing a promoter(s) that is operationally linked to the coding sequence of a protein of interest and can express the coding sequence.
- Non-limiting examples include a naked or packaged (lipid and/or protein) DNA, a naked or packaged RNA, a subcomponent of a virus or bacterium or other microorganism that may be replication- incompetent, or a virus or bacterium or other microorganism that may be replication-competent.
- a vector is sometimes referred to as a construct.
- Recombinant DNA vectors are vectors having recombinant DNA.
- a vector can include nucleic acid sequences that permit it to replicate in a host cell, such as an origin of replication.
- a vector can also include one or more selectable marker genes and other genetic elements.
- Viral vectors are recombinant nucleic acid vectors having at least some nucleic acid sequences derived from one or more viruses.
- a viral vector comprises a nucleic acid molecule encoding a disclosed antibody or antigen binding fragment that specifically binds to PfCSP and neutralizes P. falciparum.
- the viral vector can be an adeno-associated virus (AAV) vector.
- Isolated monoclonal antibodies and antigen binding fragments that specifically bind an epitope on PfCSP are provided.
- the antibodies and antigen binding fragments can be fully human.
- the antibodies and antigen binding fragments can neutralize P. falciparum, for example the disclosed antibodies can inhibit P. falciparum sporozoite infection of hepatocytes in vitro and P. falciparum sporozoite invasion of liver in vivo.
- compositions comprising the antibodies and antigen binding fragments and a pharmaceutically acceptable carrier.
- Nucleic acids encoding the antibodies or antigen binding fragments, expression vectors (such as DNA and RNA vectors for expression and delivery, as well as adeno-associated virus (AAV) viral vectors) comprising these nucleic acids are also provided.
- the antibodies, antigen binding fragments, nucleic acid molecules, host cells, and compositions can be used for research, diagnostic and prophylactic purposes.
- the disclosed antibodies and antigen binding fragments can be used to diagnose a subject with a P. falciparum infection, or can be administered prophylactically to inhibit P. falciparum infection in a subject. 1.
- monoclonal antibodies and antigen binding fragments refers to isolated monoclonal antibodies that include heavy and/or light chain variable domains (or antigen binding fragments thereof) comprising a CDR1, CDR2, and/or CDR3 with reference to the IMMGT numbering scheme (unless the context indicates otherwise).
- CDR numbering schemes such as the Kabat, Chothia or IMGT numbering schemes
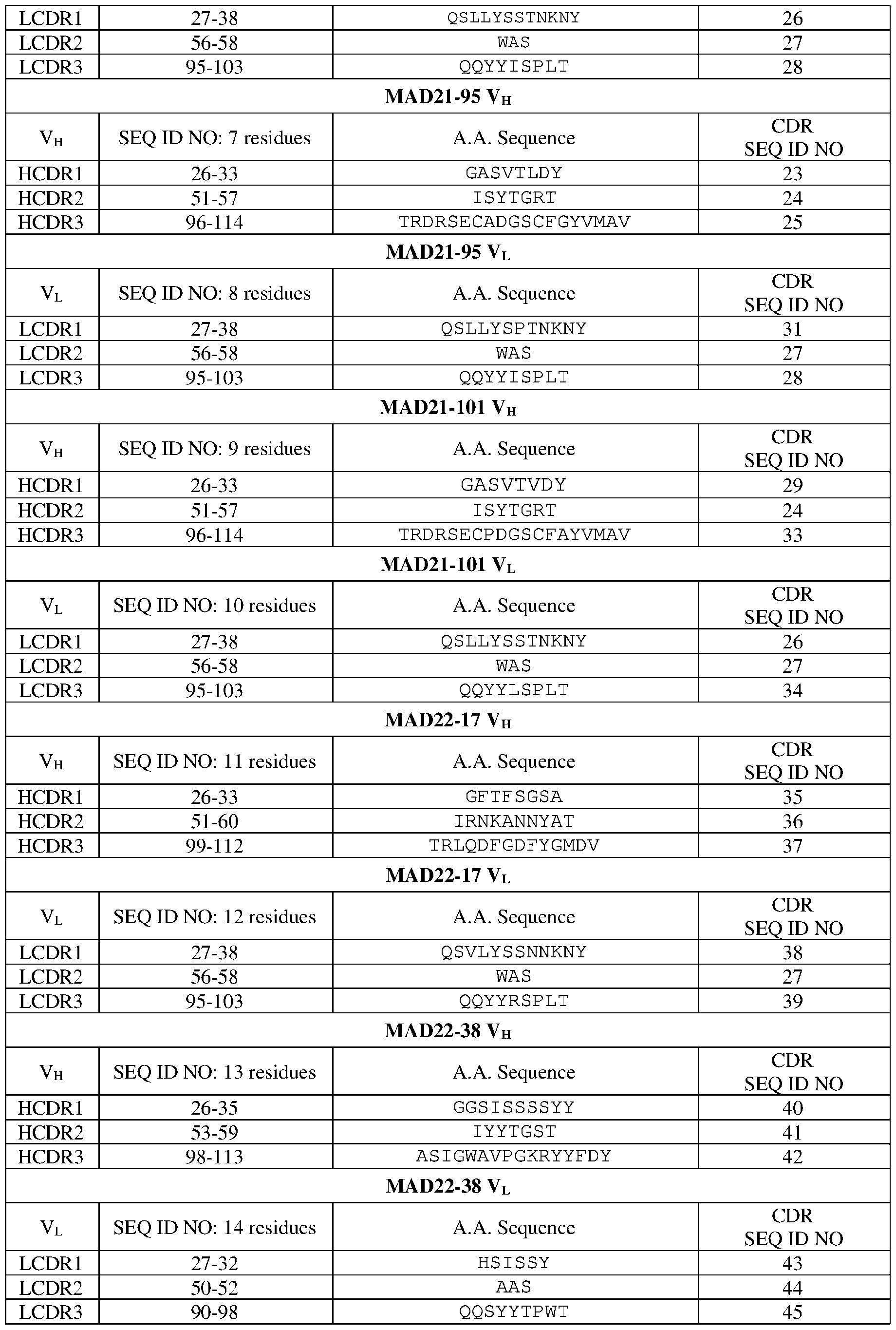
- the amino acid sequence and the CDR positions (according to the IMGT numbering scheme) of the heavy and light chains of exemplary monoclonal antibodies that bind to PfCSP and neutralize P. falciparum are shown in Table 1. Table 1.
- IMGT CDR sequences of PfCSP specific antibodies MAD21-17 VH C DR LCDR1 27-38 QSLLYSSTNKNY 26 LCDR2 56-58 WAS 27 L CDR3 95-103 YYISPLT 28 MAD22-39 VH V H SEQ ID NO: 15 residues AA Se uence CDR a.
- the antibody or antigen binding fragment is based on or derived from the MAD21- 17 antibody, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a VH and a VL comprising the HCDR1, the HCDR2, and the HCDR3, and the LCDR1, the LCDR2, and the LCDR3, respectively (for example, according to IMGT, Kabat, or Chothia), of the MAD21-17 antibody, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a V H comprising an amino acid sequence at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequence set forth as SEQ ID NO: 1, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a VL comprising an amino acid sequence at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequence set forth as SEQ ID NO: 2, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a VH and a VL independently comprising amino acid sequences at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequences set forth as SEQ ID NOs: 1 and 2, respectively, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 23, 24, and 25, respectively, and a V L comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 26, 27, and 28, respectively, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 23, 24, and 25, respectively, a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 26, 27, and 28, respectively, wherein the V H comprises an amino acid sequence at least 90% identical to SEQ ID NO: 1 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 1), the V L comprises an amino acid sequence at least 90% identical to SEQ ID NO: 2 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 2 and the antibody or antigen binding fragment specifically binds to PfCSP and neutralizes P.
- V H comprises an amino acid sequence at least 90% identical to SEQ ID NO: 1 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 1)
- the V L comprises an amino
- the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 23, 24, and 25, respectively, a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 26, 27, and 28, respectively, wherein the framework regions of the V H comprise up to 10 (such as up to 1, up to 2, up to 3, up to 4, up to 5, up to 6, up to 7, up to 8, or up to 9) amino acid substitutions (such as conservative amino acid substitutions) compared to the amino acid sequence set forth as one of SEQ ID NO: 1, and the framework regions of the VL comprise up to 10 (such as up to 1, up to 2, up to 3, up to 4, up to 5, up to 6, up to 7, up to 8, or up to 9) amino acid substitutions (such as conservative amino acid substitutions) compared to the amino acid sequence set forth as one of SEQ ID NO: 2, and the antibody or antigen binding fragment
- the antibody or antigen binding fragment comprises a V H comprising the amino acid sequence set forth as SEQ ID NO: 1, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a VL comprising the amino acid sequence set forth as SEQ ID NO: 2, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a V H and a V L comprising the amino acid sequences set forth as SEQ ID NOs: 1 and 2, respectively, and specifically binds to PfCSP and neutralizes P. falciparum.
- the disclosed antibodies and antigen binding fragments inhibit the invasion of Plasmodium falciparum sporozoites into human liver cells, and/or reduce pathogen load Plasmodium falciparum in a subject, compared to a control.
- the antibody or antigen binding fragment is based on or derived from the MAD21- 46 antibody, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a VH and a VL comprising the HCDR1, the HCDR2, and the HCDR3, and the LCDR1, the LCDR2, and the LCDR3, respectively (for example, according to IMGT, Kabat, or Chothia), of the MAD21-46 antibody, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a V H comprising an amino acid sequence at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequence set forth as SEQ ID NO: 3, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a VL comprising an amino acid sequence at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequence set forth as SEQ ID NO: 4, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a V H and a V L independently comprising amino acid sequences at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequences set forth as SEQ ID NOs: 3 and 4, respectively, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 29, 24, and 25, respectively, and a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 26, 27, and 28, respectively, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 29, 24, and 25, respectively, a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 26, 27, and 28, respectively, wherein the VH comprises an amino acid sequence at least 90% identical to SEQ ID NO: 3 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 3), the V L comprises an amino acid sequence at least 90% identical to SEQ ID NO: 4 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 4 and the antibody or antigen binding fragment specifically binds to PfCSP and neutralizes P.
- the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 29, 24, and 25, respectively, a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 26, 27, and 28, respectively, wherein the framework regions of the V H comprise up to 10 (such as up to 1, up to 2, up to 3, up to 4, up to 5, up to 6, up to 7, up to 8, or up to 9) amino acid substitutions (such as conservative amino acid substitutions) compared to the amino acid sequence set forth as one of SEQ ID NO: 3, and the framework regions of the VL comprise up to 10 (such as up to 1, up to 2, up to 3, up to 4, up to 5, up to 6, up to 7, up to 8, or up to 9) amino acid substitutions (such as conservative amino acid substitutions) compared to the amino acid sequence set forth as one of SEQ ID NO: 4, and the antibody or antigen binding fragment
- the antibody or antigen binding fragment comprises a V H comprising the amino acid sequence set forth as SEQ ID NO: 3, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a VL comprising the amino acid sequence set forth as SEQ ID NO: 4, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a VH and a VL comprising the amino acid sequences set forth as SEQ ID NOs: 3 and 4, respectively, and specifically binds to PfCSP and neutralizes P. falciparum.
- the disclosed antibodies and antigen binding fragments inhibit the invasion of Plasmodium falciparum sporozoites into human liver cells, and/or reduce pathogen load Plasmodium falciparum in a subject, compared to a control.
- the antibody or antigen binding fragment is based on or derived from the MAD21- 53 antibody, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a VH and a VL comprising the HCDR1, the HCDR2, and the HCDR3, and the LCDR1, the LCDR2, and the LCDR3, respectively (for example, according to IMGT, Kabat, or Chothia), of the MAD21-53 antibody, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a V H comprising an amino acid sequence at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequence set forth as SEQ ID NO: 5, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a VL comprising an amino acid sequence at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequence set forth as SEQ ID NO: 6, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a V H and a V L independently comprising amino acid sequences at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequences set forth as SEQ ID NOs: 5 and 6, respectively, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 29, 24, and 30, respectively, and a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 26, 27, and 28, respectively, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a V H comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 29, 24, and 30, respectively, a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 26, 27, and 28, respectively, wherein the VH comprises an amino acid sequence at least 90% identical to SEQ ID NO: 5 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 5), the VL comprises an amino acid sequence at least 90% identical to SEQ ID NO: 6 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 6 and the antibody or antigen binding fragment specifically binds to PfCSP and neutralizes P.
- VH comprises an amino acid sequence at least 90% identical to SEQ ID NO: 5 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 5)
- the VL comprises an
- the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 29, 24, and 30, respectively, a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 26, 27, and 28, respectively, wherein the framework regions of the VH comprise up to 10 (such as up to 1, up to 2, up to 3, up to 4, up to 5, up to 6, up to 7, up to 8, or up to 9) amino acid substitutions (such as conservative amino acid substitutions) compared to the amino acid sequence set forth as one of SEQ ID NO: 5, and the framework regions of the V L comprise up to 10 (such as up to 1, up to 2, up to 3, up to 4, up to 5, up to 6, up to 7, up to 8, or up to 9) amino acid substitutions (such as conservative amino acid substitutions) compared to the amino acid sequence set forth as one of SEQ ID NO: 6, and the antibody or antigen binding fragment
- the antibody or antigen binding fragment comprises a V H comprising the amino acid sequence set forth as SEQ ID NO: 5, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a VL comprising the amino acid sequence set forth as SEQ ID NO: 6, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a VH and a VL comprising the amino acid sequences set forth as SEQ ID NOs: 5 and 6, respectively, and specifically binds to PfCSP and neutralizes P. falciparum.
- the disclosed antibodies and antigen binding fragments inhibit the invasion of Plasmodium falciparum sporozoites into human liver cells, and/or reduce pathogen load Plasmodium falciparum in a subject, compared to a control.
- the antibody or antigen binding fragment is based on or derived from the MAD21- 95 antibody, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a V H and a V L comprising the HCDR1, the HCDR2, and the HCDR3, and the LCDR1, the LCDR2, and the LCDR3, respectively (for example, according to IMGT, Kabat, or Chothia), of the MAD21-95 antibody, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a VH comprising an amino acid sequence at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequence set forth as SEQ ID NO: 7, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a V L comprising an amino acid sequence at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequence set forth as SEQ ID NO: 8, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a VH and a VL independently comprising amino acid sequences at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequences set forth as SEQ ID NOs: 7 and 8, respectively, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 23, 24, and 25, respectively, and a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 31, 27, and 38, respectively, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a V H comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 23, 24, and 25, respectively, a V L comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 31, 27, and 38, respectively, wherein the VH comprises an amino acid sequence at least 90% identical to SEQ ID NO: 7 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 7), the VL comprises an amino acid sequence at least 90% identical to SEQ ID NO: 8 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 8 and the antibody or antigen binding fragment specifically binds to PfCSP and neutralizes P.
- the antibody or antigen binding fragment comprises a V H comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 23, 24, and 25, respectively, a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 31, 27, and 38, respectively, wherein the framework regions of the VH comprise up to 10 (such as up to 1, up to 2, up to 3, up to 4, up to 5, up to 6, up to 7, up to 8, or up to 9) amino acid substitutions (such as conservative amino acid substitutions) compared to the amino acid sequence set forth as one of SEQ ID NO: 7, and the framework regions of the V L comprise up to 10 (such as up to 1, up to 2, up to 3, up to 4, up to 5, up to 6, up to 7, up to 8, or up to 9) amino acid substitutions (such as conservative amino acid substitutions) compared to the amino acid sequence set forth as one of SEQ ID NO: 8, and the antibody or antigen binding
- the antibody or antigen binding fragment comprises a VH comprising the amino acid sequence set forth as SEQ ID NO: 7, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a V L comprising the amino acid sequence set forth as SEQ ID NO: 8, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a VH and a VL comprising the amino acid sequences set forth as SEQ ID NOs: 7 and 8, respectively, and specifically binds to PfCSP and neutralizes P. falciparum.
- the disclosed antibodies and antigen binding fragments inhibit the invasion of Plasmodium falciparum sporozoites into human liver cells, and/or reduce pathogen load Plasmodium falciparum in a subject, compared to a control.
- the antibody or antigen binding fragment is based on or derived from the MAD21- 101 antibody, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a V H and a V L comprising the HCDR1, the HCDR2, and the HCDR3, and the LCDR1, the LCDR2, and the LCDR3, respectively (for example, according to IMGT, Kabat, or Chothia), of the MAD21-101 antibody, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a VH comprising an amino acid sequence at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequence set forth as SEQ ID NO: 9, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a V L comprising an amino acid sequence at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequence set forth as SEQ ID NO: 10, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a VH and a VL independently comprising amino acid sequences at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequences set forth as SEQ ID NOs: 9 and 10, respectively, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a V H comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 29, 24, and 33, respectively, and a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 26, 27, and 34, respectively, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 29, 24, and 33, respectively, a V L comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 26, 27, and 34, respectively, wherein the V H comprises an amino acid sequence at least 90% identical to SEQ ID NO: 9 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 9), the VL comprises an amino acid sequence at least 90% identical to SEQ ID NO: 10 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 10 and the antibody or antigen binding fragment specifically binds to PfCSP and neutralizes P.
- V H comprises an amino acid sequence at least 90% identical to SEQ ID NO: 9 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 9)
- the VL
- the antibody or antigen binding fragment comprises a V H comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 29, 24, and 33, respectively, a V L comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 26, 27, and 34, respectively, wherein the framework regions of the VH comprise up to 10 (such as up to 1, up to 2, up to 3, up to 4, up to 5, up to 6, up to 7, up to 8, or up to 9) amino acid substitutions (such as conservative amino acid substitutions) compared to the amino acid sequence set forth as one of SEQ ID NO: 9, and the framework regions of the VL comprise up to 10 (such as up to 1, up to 2, up to 3, up to 4, up to 5, up to 6, up to 7, up to 8, or up to 9) amino acid substitutions (such as conservative amino acid substitutions) compared to the amino acid sequence set forth as one of SEQ ID NO: 10, and the antibody or antigen
- the antibody or antigen binding fragment comprises a VH comprising the amino acid sequence set forth as SEQ ID NO: 9, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a VL comprising the amino acid sequence set forth as SEQ ID NO: 10, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a V H and a V L comprising the amino acid sequences set forth as SEQ ID NOs: 9 and 10, respectively, and specifically binds to PfCSP and neutralizes P. falciparum.
- the disclosed antibodies and antigen binding fragments inhibit the invasion of Plasmodium falciparum sporozoites into human liver cells, and/or reduce pathogen load Plasmodium falciparum in a subject, compared to a control.
- the antibody or antigen binding fragment is based on or derived from the MAD22- 17 antibody, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a V H and a V L comprising the HCDR1, the HCDR2, and the HCDR3, and the LCDR1, the LCDR2, and the LCDR3, respectively (for example, according to IMGT, Kabat, or Chothia), of the MAD22-17 antibody, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a VH comprising an amino acid sequence at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequence set forth as SEQ ID NO: 11, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a VL comprising an amino acid sequence at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequence set forth as SEQ ID NO: 12, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a VH and a VL independently comprising amino acid sequences at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequences set forth as SEQ ID NOs: 11 and 12, respectively, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a V H comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 35, 36, and 37, respectively, and a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 38, 27, and 39, respectively, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a V H comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 35, 36, and 37, respectively, a V L comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 38, 27, and 39, respectively, wherein the V H comprises an amino acid sequence at least 90% identical to SEQ ID NO: 11 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 11), the VL comprises an amino acid sequence at least 90% identical to SEQ ID NO: 12 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 12 and the antibody or antigen binding fragment specifically binds to PfCSP and neutralizes P.
- V H comprises an amino acid sequence at least 90% identical to SEQ ID NO: 11 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 11)
- the antibody or antigen binding fragment comprises a V H comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 35, 36, and 37, respectively, a V L comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 38, 27, and 39, respectively, wherein the framework regions of the VH comprise up to 10 (such as up to 1, up to 2, up to 3, up to 4, up to 5, up to 6, up to 7, up to 8, or up to 9) amino acid substitutions (such as conservative amino acid substitutions) compared to the amino acid sequence set forth as one of SEQ ID NO: 11, and the framework regions of the VL comprise up to 10 (such as up to 1, up to 2, up to 3, up to 4, up to 5, up to 6, up to 7, up to 8, or up to 9) amino acid substitutions (such as conservative amino acid substitutions) compared to the amino acid sequence set forth as one of SEQ ID NO: 12, and the antibody
- the antibody or antigen binding fragment comprises a VH comprising the amino acid sequence set forth as SEQ ID NO: 11, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a V L comprising the amino acid sequence set forth as SEQ ID NO: 12, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a VH and a VL comprising the amino acid sequences set forth as SEQ ID NOs: 11 and 12, respectively, and specifically binds to PfCSP and neutralizes P. falciparum.
- the disclosed antibodies and antigen binding fragments inhibit the invasion of Plasmodium falciparum sporozoites into human liver cells, and/or reduce pathogen load Plasmodium falciparum in a subject, compared to a control.
- the antibody or antigen binding fragment is based on or derived from the MAD22- 38 antibody, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a V H and a V L comprising the HCDR1, the HCDR2, and the HCDR3, and the LCDR1, the LCDR2, and the LCDR3, respectively (for example, according to IMGT, Kabat, or Chothia), of the MAD22-38 antibody, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a VH comprising an amino acid sequence at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequence set forth as SEQ ID NO: 13, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a V L comprising an amino acid sequence at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequence set forth as SEQ ID NO: 14, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a VH and a VL independently comprising amino acid sequences at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequences set forth as SEQ ID NOs: 13 and 14, respectively, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 40, 41, and 42, respectively, and a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 43, 44, and 45, respectively, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a V H comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 40, 41, and 42, respectively, a V L comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 43, 44, and 45, respectively, wherein the VH comprises an amino acid sequence at least 90% identical to SEQ ID NO: 13 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 13), the VL comprises an amino acid sequence at least 90% identical to SEQ ID NO: 14 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 14 and the antibody or antigen binding fragment specifically binds to PfCSP and neutralizes P.
- VH comprises an amino acid sequence at least 90% identical to SEQ ID NO: 13 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 13)
- the antibody or antigen binding fragment comprises a V H comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 40, 41, and 42, respectively, a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 43, 44, and 45, respectively, wherein the framework regions of the VH comprise up to 10 (such as up to 1, up to 2, up to 3, up to 4, up to 5, up to 6, up to 7, up to 8, or up to 9) amino acid substitutions (such as conservative amino acid substitutions) compared to the amino acid sequence set forth as one of SEQ ID NO: 13, and the framework regions of the V L comprise up to 10 (such as up to 1, up to 2, up to 3, up to 4, up to 5, up to 6, up to 7, up to 8, or up to 9) amino acid substitutions (such as conservative amino acid substitutions) compared to the amino acid sequence set forth as one of SEQ ID NO: 14, and the
- the antibody or antigen binding fragment comprises a VH comprising the amino acid sequence set forth as SEQ ID NO: 13, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a V L comprising the amino acid sequence set forth as SEQ ID NO: 14, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a VH and a VL comprising the amino acid sequences set forth as SEQ ID NOs: 13 and 14, respectively, and specifically binds to PfCSP and neutralizes P. falciparum.
- the disclosed antibodies and antigen binding fragments inhibit the invasion of Plasmodium falciparum sporozoites into human liver cells, and/or reduce pathogen load Plasmodium falciparum in a subject, compared to a control.
- the antibody or antigen binding fragment is based on or derived from the MAD22- 39 antibody, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a V H and a V L comprising the HCDR1, the HCDR2, and the HCDR3, and the LCDR1, the LCDR2, and the LCDR3, respectively (for example, according to IMGT, Kabat, or Chothia), of the MAD22-39 antibody, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a VH comprising an amino acid sequence at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequence set forth as SEQ ID NO: 15, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a V L comprising an amino acid sequence at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequence set forth as SEQ ID NO: 16, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a VH and a VL independently comprising amino acid sequences at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequences set forth as SEQ ID NOs: 15 and 16, respectively, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 46, 47, and 48, respectively, and a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 49, 44, and 45, respectively, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a V H comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 46, 47, and 48, respectively, a V L comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 49, 44, and 45, respectively, wherein the VH comprises an amino acid sequence at least 90% identical to SEQ ID NO: 15 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 15), the VL comprises an amino acid sequence at least 90% identical to SEQ ID NO: 16 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 16 and the antibody or antigen binding fragment specifically binds to PfCSP and neutralizes P.
- VH comprises an amino acid sequence at least 90% identical to SEQ ID NO: 15 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 15)
- the antibody or antigen binding fragment comprises a V H comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 46, 47, and 48, respectively, a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 49, 44, and 45, respectively, wherein the framework regions of the VH comprise up to 10 (such as up to 1, up to 2, up to 3, up to 4, up to 5, up to 6, up to 7, up to 8, or up to 9) amino acid substitutions (such as conservative amino acid substitutions) compared to the amino acid sequence set forth as one of SEQ ID NO: 15, and the framework regions of the V L comprise up to 10 (such as up to 1, up to 2, up to 3, up to 4, up to 5, up to 6, up to 7, up to 8, or up to 9) amino acid substitutions (such as conservative amino acid substitutions) compared to the amino acid sequence set forth as one of SEQ ID NO: 16, and the
- the antibody or antigen binding fragment comprises a VH comprising the amino acid sequence set forth as SEQ ID NO: 15, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a V L comprising the amino acid sequence set forth as SEQ ID NO: 16, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a VH and a VL comprising the amino acid sequences set forth as SEQ ID NOs: 15 and 16, respectively, and specifically binds to PfCSP and neutralizes P. falciparum.
- the disclosed antibodies and antigen binding fragments inhibit the invasion of Plasmodium falciparum sporozoites into human liver cells, and/or reduce pathogen load Plasmodium falciparum in a subject, compared to a control.
- the antibody or antigen binding fragment is based on or derived from the MAD24- 01 antibody, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a V H and a V L comprising the HCDR1, the HCDR2, and the HCDR3, and the LCDR1, the LCDR2, and the LCDR3, respectively (for example, according to IMGT, Kabat, or Chothia), of the MAD24-01 antibody, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a VH comprising an amino acid sequence at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequence set forth as SEQ ID NO: 17, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a V L comprising an amino acid sequence at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequence set forth as SEQ ID NO: 18, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a VH and a VL independently comprising amino acid sequences at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequences set forth as SEQ ID NOs: 17 and 18, respectively, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 50, 51, and 52, respectively, and a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 53, 54, and 55, respectively, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a V H comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 50, 51, and 52, respectively, a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 53, 54, and 55, respectively, wherein the VH comprises an amino acid sequence at least 90% identical to SEQ ID NO: 17 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 17), the VL comprises an amino acid sequence at least 90% identical to SEQ ID NO: 18 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 18 and the antibody or antigen binding fragment specifically binds to PfCSP and neutralizes P.
- VH comprises an amino acid sequence at least 90% identical to SEQ ID NO: 17 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 17)
- the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 50, 51, and 52, respectively, a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 53, 54, and 55, respectively, wherein the framework regions of the VH comprise up to 10 (such as up to 1, up to 2, up to 3, up to 4, up to 5, up to 6, up to 7, up to 8, or up to 9) amino acid substitutions (such as conservative amino acid substitutions) compared to the amino acid sequence set forth as one of SEQ ID NO: 17, and the framework regions of the V L comprise up to 10 (such as up to 1, up to 2, up to 3, up to 4, up to 5, up to 6, up to 7, up to 8, or up to 9) amino acid substitutions (such as conservative amino acid substitutions) compared to the amino acid sequence set forth as one of SEQ ID NO: 18, and the
- the antibody or antigen binding fragment comprises a V H comprising the amino acid sequence set forth as SEQ ID NO: 17, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a VL comprising the amino acid sequence set forth as SEQ ID NO: 18, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a VH and a VL comprising the amino acid sequences set forth as SEQ ID NOs: 17 and 18, respectively, and specifically binds to PfCSP and neutralizes P. falciparum.
- the disclosed antibodies and antigen binding fragments inhibit the invasion of Plasmodium falciparum sporozoites into human liver cells, and/or reduce pathogen load Plasmodium falciparum in a subject, compared to a control.
- the antibody or antigen binding fragment is based on or derived from the MAD24- 05 antibody, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a V H and a V L comprising the HCDR1, the HCDR2, and the HCDR3, and the LCDR1, the LCDR2, and the LCDR3, respectively (for example, according to IMGT, Kabat, or Chothia), of the MAD24-05 antibody, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a VH comprising an amino acid sequence at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequence set forth as SEQ ID NO: 17, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a V L comprising an amino acid sequence at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequence set forth as SEQ ID NO: 18, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a VH and a VL independently comprising amino acid sequences at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequences set forth as SEQ ID NOs: 17 and 18, respectively, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 56, 57, and 58, respectively, and a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 59, 60, and 61, respectively, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a V H comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 56, 57, and 58, respectively, a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 59, 60, and 61, respectively, wherein the VH comprises an amino acid sequence at least 90% identical to SEQ ID NO: 17 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 17), the V L comprises an amino acid sequence at least 90% identical to SEQ ID NO: 18 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 18 and the antibody or antigen binding fragment specifically binds to PfCSP and neutralizes P.
- VH comprises an amino acid sequence at least 90% identical to SEQ ID NO: 17 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO
- the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 56, 57, and 58, respectively, a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 59, 60, and 61, respectively, wherein the framework regions of the VH comprise up to 10 (such as up to 1, up to 2, up to 3, up to 4, up to 5, up to 6, up to 7, up to 8, or up to 9) amino acid substitutions (such as conservative amino acid substitutions) compared to the amino acid sequence set forth as one of SEQ ID NO: 17, and the framework regions of the V L comprise up to 10 (such as up to 1, up to 2, up to 3, up to 4, up to 5, up to 6, up to 7, up to 8, or up to 9) amino acid substitutions (such as conservative amino acid substitutions) compared to the amino acid sequence set forth as one of SEQ ID NO:
- the antibody or antigen binding fragment comprises a V H comprising the amino acid sequence set forth as SEQ ID NO: 17, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a V L comprising the amino acid sequence set forth as SEQ ID NO: 18, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a VH and a VL comprising the amino acid sequences set forth as SEQ ID NOs: 17 and 18, respectively, and specifically binds to PfCSP and neutralizes P. falciparum.
- the disclosed antibodies and antigen binding fragments inhibit the invasion of Plasmodium falciparum sporozoites into human liver cells, and/or reduce pathogen load Plasmodium falciparum in a subject, compared to a control.
- MAD24-52 In some aspects, the antibody or antigen binding fragment is based on or derived from the MAD24- 52 antibody, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a V H and a V L comprising the HCDR1, the HCDR2, and the HCDR3, and the LCDR1, the LCDR2, and the LCDR3, respectively (for example, according to IMGT, Kabat, or Chothia), of the MAD24-52 antibody, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a VH comprising an amino acid sequence at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequence set forth as SEQ ID NO: 19, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a V L comprising an amino acid sequence at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequence set forth as SEQ ID NO: 20, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a V H and a V L independently comprising amino acid sequences at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequences set forth as SEQ ID NOs: 19 and 20, respectively, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 62, 63, and 64, respectively, and a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 65, 66, and 67, respectively, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a V H comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 62, 63, and 64, respectively, a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 65, 66, and 67, respectively, wherein the VH comprises an amino acid sequence at least 90% identical to SEQ ID NO: 19 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 19), the VL comprises an amino acid sequence at least 90% identical to SEQ ID NO: 20 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 20 and the antibody or antigen binding fragment specifically binds to PfCSP and neutralizes P.
- VH comprises an amino acid sequence at least 90% identical to SEQ ID NO: 19 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO
- the antibody or antigen binding fragment comprises a V H comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 62, 63, and 64, respectively, a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 65, 66, and 67, respectively, wherein the framework regions of the VH comprise up to 10 (such as up to 1, up to 2, up to 3, up to 4, up to 5, up to 6, up to 7, up to 8, or up to 9) amino acid substitutions (such as conservative amino acid substitutions) compared to the amino acid sequence set forth as one of SEQ ID NO: 19, and the framework regions of the V L comprise up to 10 (such as up to 1, up to 2, up to 3, up to 4, up to 5, up to 6, up to 7, up to 8, or up to 9) amino acid substitutions (such as conservative amino acid substitutions) compared to the amino acid sequence set forth as one of SEQ ID NO:
- the antibody or antigen binding fragment comprises a VH comprising the amino acid sequence set forth as SEQ ID NO: 19, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a V L comprising the amino acid sequence set forth as SEQ ID NO: 20, and specifically binds to PfCSP and neutralizes P. falciparum.
- the antibody or antigen binding fragment comprises a VH and a VL comprising the amino acid sequences set forth as SEQ ID NOs: 19 and 20, respectively, and specifically binds to PfCSP and neutralizes P. falciparum.
- the disclosed antibodies and antigen binding fragments inhibit the invasion of Plasmodium falciparum sporozoites into human liver cells, and/or reduce pathogen load Plasmodium falciparum in a subject, compared to a control.
- the antibody or antigen binding fragment can be a human antibody or fragment thereof. Chimeric antibodies are also provided.
- the antibody or antigen binding fragment can include any suitable framework region, such as (but not limited to) a human framework region.
- a heterologous framework region such as, but not limited to a mouse or monkey framework region, can be included in the heavy or light chain of the antibodies.
- the antibody can be of any isotype.
- the antibody can be, for example, an IgM or an IgG antibody, such as IgG1, IgG2, IgG3, or IgG4.
- the class of an antibody that specifically binds PfCSP can be switched with another.
- a nucleic acid molecule encoding V L or V H is isolated such that it does not include any nucleic acid sequences encoding the constant region of the light or heavy chain, respectively.
- a nucleic acid molecule encoding VL or VH is then operatively linked to a nucleic acid sequence encoding a CL or CH from a different class of immunoglobulin molecule. This can be achieved, for example, using a vector or nucleic acid molecule that comprises a CL or CH chain.
- an antibody that specifically binds PfCSP, that was originally IgG may be class switched to an IgM. Class switching can be used to convert one IgG subclass to another, such as from IgG 1 to IgG 2, IgG 3, or IgG 4 .
- exemplary full-length heavy and light chain sequences containing the MAD21-101 VH and VL are provided as SEQ ID NOs: 157 and 158, respectively.
- the disclosed antibodies are oligomers of antibodies, such as dimers, trimers, tetramers, pentamers, hexamers, septamers, octomers and so on.
- the antibody or antigen binding fragment can be derivatized or linked to another molecule (such as another peptide or protein).
- the antibody or antigen binding fragment is derivatized such that the binding to P. falciparum is not affected adversely by the derivatization or labeling.
- the antibody or antigen binding fragment can be functionally linked (by chemical coupling, genetic fusion, noncovalent association or otherwise) to one or more other molecular entities, such as another antibody (for example, a bi-specific antibody or a diabody), a detectable marker, an effector molecule, or a protein or peptide that can mediate association of the antibody or antibody portion with another molecule (such as a streptavidin core region or a polyhistidine tag).
- the antibody or antigen binding fragment specifically binds PfCSP with an affinity (e.g., measured by KD) of no more than 1.0 x 10 -8 M, no more than 5.0 x 10 -8 M, no more than 1.0 x 10 -9 M, no more than 5.0 x 10 -9 M, no more than 1.0 x 10 -10 M, no more than 5.0 x 10 -10 M, or no more than 1.0 x 10 -11 M.
- K D can be measured, for example, by a radiolabeled antigen binding assay (RIA) performed with the Fab version of an antibody of interest and its antigen.
- RIA radiolabeled antigen binding assay
- solution binding affinity of Fabs for antigen is measured by equilibrating Fab with a minimal concentration of ( 125 I)-labeled antigen in the presence of a titration series of unlabeled antigen, then capturing bound antigen with an anti-Fab antibody-coated plate (see, e.g., Chen et al., J. Mol. Biol.293(4):865-881, 1999).
- MICROTITER® multi-well plates (Thermo Scientific) are coated overnight with 5 ⁇ g/ml of a capturing anti-Fab antibody (Cappel Labs) in 50 mM sodium carbonate (pH 9.6), and subsequently blocked with 2% (w/v) bovine serum albumin in PBS for two to five hours at room temperature (approximately 23° C.).
- a non-adsorbent plate (NuncTM Catalog #269620) 100 ⁇ M or 26 pM [ 125 I]-antigen are mixed with serial dilutions of a Fab of interest (e.g., consistent with assessment of the anti-VEGF antibody, Fab-12, in Presta et al., Cancer Res.57(20):4593-4599, 1997).
- the Fab of interest is then incubated overnight; however, the incubation may continue for a longer period (e.g., about 65 hours) to ensure that equilibrium is reached. Thereafter, the mixtures are transferred to the capture plate for incubation at room temperature (e.g., for one hour).
- K D can be measured using surface plasmon resonance assays using a BIACORE®- 2000 or a BIACORE®-3000 (BIAcore, Inc., Piscataway, N.J.) at 25° C with immobilized antigen CM5 chips at ⁇ 10 response units (RU). Briefly, carboxymethylated dextran biosensor chips (CM5, BIACORE®, Inc.) are activated with N-ethyl-N′-(3-dimethylaminopropyl)-carbodiimide hydrochloride (EDC) and N- hydroxysuccinimide (NHS) according to the supplier's instructions.
- CM5 carboxymethylated dextran biosensor chips
- EDC N-ethyl-N′-(3-dimethylaminopropyl)-carbodiimide hydrochloride
- NHS N- hydroxysuccinimide
- Antigen is diluted with 10 mM sodium acetate, pH 4.8, to 5 ⁇ g/ml ( ⁇ 0.2 ⁇ M) before injection at a flow rate of 5 l/minute to achieve approximately 10 response units (RU) of coupled protein. Following the injection of antigen, 1 M ethanolamine is injected to block unreacted groups. For kinetics measurements, two-fold serial dilutions of Fab (0.78 nM to 500 nM) are injected in PBS with 0.05% polysorbate 20 (TWEEN-20TM) surfactant (PBST) at 25° C at a flow rate of approximately 25 l/min.
- TWEEN-20TM polysorbate 20
- association rates (kon) and dissociation rates (koff) are calculated using a simple one-to-one Langmuir binding model (BIACORE® Evaluation Software version 3.2) by simultaneously fitting the association and dissociation sensorgrams.
- the equilibrium dissociation constant (KD) is calculated as the ratio koff/kon. See, e.g., Chen et al., J. Mol. Biol.293:865-881 (1999).
- a multi-specific antibody such as a bi-specific antibody, is provided that comprises an antibody or antigen binding fragment as provided herein, and specifically binds to PfCSP.
- Any suitable method can be used to design and produce the multi-specific antibody, such as crosslinking two or more antibodies, antigen binding fragments (such as scFvs) of the same type or of different types.
- Exemplary methods of making multispecific antibodies include those described in PCT Pub. No. WO2013/163427.
- Non-limiting examples of suitable crosslinkers include those that are heterobifunctional, having two distinctly reactive groups separated by an appropriate spacer (such as m-maleimidobenzoyl-N- hydroxysuccinimide ester) or homobifunctional (such as disuccinimidyl suberate).
- the multi-specific antibody may have any suitable format that allows for antigen binding by the antibody or antigen binding fragment as provided herein, such as any one of MAD21-17, MAD21-46, MAD21-53, MAD21-95, MAD21-101, MAD22-17, MAD22-38, MAD22-39, MAD24-01, MAD24-05, or MAD24-52, or an antigen binding fragment thereof.
- Bispecific single chain antibodies can be encoded by a single nucleic acid molecule.
- Non-limiting examples of bispecific single chain antibodies, as well as methods of constructing such antibodies are provided in U.S. Pat. Nos.8,076,459, 8,017,748, 8,007,796, 7,919,089, 7,820,166, 7,635,472, 7,575,923, 7,435,549, 7,332,168, 7,323,440, 7,235,641, 7,229,760, 7,112,324, 6,723,538. Additional examples of bispecific single chain antibodies can be found in PCT application No. WO 99/54440; Mack et al., J. Immunol., 158(8):3965-3970, 1997; Mack et al., Proc.
- bispecific Fab-scFv (“bibody”) molecules are described, for example, in Schoonjans et al. (J. Immunol., 165(12):7050-7057, 2000) and Willems et al. (J. Chromatogr. B Analyt. Technol.
- a scFv molecule can be fused to one of the VL-CL (L) or VH- CH1 chains, e.g., to produce a bibody one scFv is fused to the C-term of a Fab chain.
- Fragments Antigen binding fragments are encompassed by the present disclosure, such as Fab, F(ab')2, and Fv which include a heavy chain and VL and specifically bind PfCSP. These antibody fragments retain the ability to selectively bind with the antigen and are “antigen-binding” fragments.
- Non-limiting examples of such fragments include: (1) Fab, the fragment which contains a monovalent antigen-binding fragment of an antibody molecule, can be produced by digestion of whole antibody with the enzyme papain to yield an intact light chain and a portion of one heavy chain; (2) Fab', the fragment of an antibody molecule can be obtained by treating whole antibody with pepsin, followed by reduction, to yield an intact light chain and a portion of the heavy chain; (3) (Fab')2, the fragment of the antibody that can be obtained by treating whole antibody with the enzyme pepsin without subsequent reduction; F(ab') 2 is a dimer of two Fab' fragments held together by two disulfide bonds; (4) Fv, a genetically engineered fragment containing the VL and VL expressed as two chains; and (5) Single chain antibody (such as scFv), defined as a genetically engineered molecule containing the V H and the V L linked by a suitable polypeptide linker as a genetically fused single chain molecule (see, e.g.,
- VH-domain-linker domain-VL-domain VL-domain-linker domain-VH-domain
- scFV2 A dimer of a single chain antibody
- Antigen binding fragments can be prepared by proteolytic hydrolysis of the antibody or by expression in a host cell (such as an E. coli cell) of DNA encoding the fragment.
- Antigen binding fragments can also be obtained by pepsin or papain digestion of whole antibodies by conventional methods. For example, antigen binding fragments can be produced by enzymatic cleavage of antibodies with pepsin to provide a 5S fragment denoted F(ab') 2 .
- This fragment can be further cleaved using a thiol reducing agent, and optionally a blocking group for the sulfhydryl groups resulting from cleavage of disulfide linkages, to produce 3.5S Fab' monovalent fragments.
- Other methods of cleaving antibodies such as separation of heavy chains to form monovalent light- heavy chain fragments, further cleavage of fragments, or other enzymatic, chemical, or genetic techniques may also be used, so long as the fragments bind to the antigen that is recognized by the intact antibody.
- amino acid sequence variants of the antibodies and antigen binding fragments provided herein are provided. For example, it may be desirable to improve the binding affinity and/or other biological properties of the antibody.
- Amino acid sequence variants of an antibody may be prepared by introducing appropriate modifications into the nucleotide sequence encoding the antibody, or by peptide synthesis. Such modifications include, for example, deletions from, and/or insertions into and/or substitutions of residues within the amino acid sequences of the antibody. Any combination of deletion, insertion, and substitution can be made to arrive at the final construct, provided that the final construct possesses the desired characteristics, e.g., antigen-binding. In some aspects, antibody variants having one or more amino acid substitutions are provided. Sites of interest for substitutional mutagenesis include the CDRs and the framework regions.
- Amino acid substitutions may be introduced into an antibody of interest and the products screened for a desired activity, e.g., retained/improved antigen binding, decreased immunogenicity, or improved ADCC or CDC.
- the variants typically retain amino acid residues necessary for correct folding and stabilizing between the VH and the VL regions, and will retain the charge characteristics of the residues in order to preserve the low pI and low toxicity of the molecules.
- Amino acid substitutions can be made in the V H and the V L regions to increase yield.
- the antibody or antigen binding fragment can include up to 10 (such as up to 1, up to 2, up to 3, up to 4, up to 5, up to 6, up to 7, up to 8, or up to 9) amino acid substitutions (such as conservative amino acid substitutions) in the framework regions of the heavy chain of the antibody, or the light chain of the antibody, or the heavy and light chains of the antibody, compared to known framework regions, or compared to the framework regions of an antibody as provided herein, and maintain the specific binding activity for PfCSP.
- substitutions, insertions, or deletions may occur within one or more CDRs so long as such alterations do not substantially reduce the ability of the antibody to bind antigen.
- each CDR either is unaltered, or contains no more than one, two or three amino acid substitutions.
- the V L and V H segments can be randomly mutated, such as within HCDR3 region or the LCDR3 region, in a process analogous to the in vivo somatic mutation process responsible for affinity maturation of antibodies during a natural immune response.
- in vitro affinity maturation can be accomplished by amplifying VH and VL regions using PCR primers complementary to the HCDR3 or LCDR3, respectively.
- the primers have been “spiked” with a random mixture of the four nucleotide bases at certain positions such that the resultant PCR products encode V H and V L segments into which random mutations have been introduced into the V H and/or V L CDR3 regions. These randomly mutated V H and V L segments can be tested to determine the binding affinity for PfCSP.
- an antibody or antigen binding fragment is altered to increase or decrease the extent to which the antibody or antigen binding fragment is glycosylated. Addition or deletion of glycosylation sites may be conveniently accomplished by altering the amino acid sequence such that one or more glycosylation sites is created or removed. Where the antibody comprises an Fc region, the carbohydrate attached thereto may be altered.
- Native antibodies produced by mammalian cells typically comprise a branched, biantennary oligosaccharide that is generally attached by an N-linkage to Asn297 of the CH2 domain of the Fc region. See, e.g., Wright et al. Trends Biotechnol.15(1):26-32, 1997.
- the oligosaccharide may include various carbohydrates, e.g., mannose, N-acetyl glucosamine (GlcNAc), galactose, and sialic acid, as well as a fucose attached to a GlcNAc in the “stem” of the biantennary oligosaccharide structure.
- modifications of the oligosaccharide in an antibody may be made in order to create antibody variants with certain improved properties.
- antibody variants are provided having a carbohydrate structure that lacks fucose attached (directly or indirectly) to an Fc region.
- the amount of fucose in such antibody may be from 1% to 80%, from 1% to 65%, from 5% to 65% or from 20% to 40%.
- the amount of fucose is determined by calculating the average amount of fucose within the sugar chain at Asn297, relative to the sum of all glycostructures attached to Asn 297 (e.g.
- Asn297 refers to the asparagine residue located at about position 297 in the Fc region; however, Asn297 may also be located about ⁇ 3 amino acids upstream or downstream of position 297, i.e., between positions 294 and 300, due to minor sequence variations in antibodies. Such fucosylation variants may have improved ADCC function. See, e.g., US Patent Publication Nos. US 2003/0157108 (Presta, L.); US 2004/0093621 (Kyowa Hakko Kogyo Co., Ltd).
- Examples of publications related to “defucosylated” or “fucose-deficient” antibody variants include: US 2003/0157108; WO 2000/61739; WO 2001/29246; US 2003/0115614; US 2002/0164328; US 2004/0093621; US 2004/0132140; US 2004/0110704; US 2004/0110282; US 2004/0109865; WO 2003/085119; WO 2003/084570; WO 2005/035586; WO 2005/035778; WO2005/053742; WO 2002/031140; Okazaki et al., J. Mol.
- Antibody variants are further provided with bisected oligosaccharides, e.g., in which a biantennary oligosaccharide attached to the Fc region of the antibody is bisected by GlcNAc. Such antibody variants may have reduced fucosylation and/or improved ADCC function. Examples of such antibody variants are described, e.g., in WO 2003/011878 (Jean-Mairet et al.); U.S. Pat. No.6,602,684 (Umana et al.); and US 2005/0123546 (Umana et al.). Antibody variants with at least one galactose residue in the oligosaccharide attached to the Fc region are also provided.
- the constant region of the antibody (such as any one of MAD21-17, MAD21-46, MAD21-53, MAD21-95, MAD21-101, MAD22-17, MAD22-38, MAD22-39, MAD24-01, MAD24-05, or MAD24-52) comprises one or more amino acid substitutions to optimize in vivo half-life of the antibody.
- the serum half-life of IgG Abs is regulated by the neonatal Fc receptor (FcRn).
- the antibody comprises an amino acid substitution that increases binding to the FcRn.
- substitutions include substitutions at IgG constant regions T250Q and M428L (see, e.g., Hinton et al., J Immunol., 176(1):346-356, 2006); M428L and N434S (the “LS” mutation, see, e.g., Zalevsky, et al., Nature Biotechnol., 28(2):157-159, 2010); N434A (see, e.g., Petkova et al., Int.
- the disclosed antibodies (such as any one of MAD21-17, MAD21-46, MAD21-53, MAD21-95, MAD21-101, MAD22-17, MAD22-38, MAD22-39, MAD24-01, MAD24-05, or MAD24-52) and antigen binding fragments can be linked to or comprise an Fc polypeptide including any of the substitutions listed above, for example, the Fc polypeptide can include the M428L and N434S substitutions.
- the constant region of the antibody comprises one or more amino acid substitutions to optimize ADCC. ADCC is mediated primarily through a set of closely related Fc ⁇ receptors.
- the antibody comprises one or more amino acid substitutions that increase binding to Fc ⁇ RIIIa.
- Non-limiting examples of such substitutions include substitutions at IgG constant regions S239D and I332E (see, e.g., Lazar et al., Proc. Natl., Acad. Sci. U.S.A., 103(11):4005-4010, 2006); and S239D, A330L, and I332E (see, e.g., Lazar et al., Proc. Natl., Acad. Sci. U.S.A., 103(11):4005-4010, 2006). Combinations of the above substitutions are also included, to generate an IgG constant region with increased binding to FcRn and Fc ⁇ RIIIa. The combinations increase antibody half-life and ADCC.
- such combinations include antibodies with the following amino acid substitutions in the Fc region: (1) S239D/I332E and T250Q/M428L; (2) S239D/I332E and M428L/N434S; (3) S239D/I332E and N434A; (4) S239D/I332E and T307A/E380A/N434A; (5) S239D/I332E and M252Y/S254T/T256E; (6) S239D/A330L/I332E and 250Q/M428L; (7) S239D/A330L/I332E and M428L/N434S; (8) S239D/A330L/I332E and N434A; (9) S239D/A330L/I332E and T307A/E380A/N434A; or (10) S239D/A330L/I332E and M252Y/S254
- the antibodies, or an antigen binding fragment thereof is modified such that it is directly cytotoxic to infected cells, or uses natural defenses such as complement, ADCC, or phagocytosis by macrophages.
- an antibody provided herein may be further modified to contain additional nonproteinaceous moieties.
- the moieties suitable for derivatization of the antibody include but are not limited to water soluble polymers.
- Non-limiting examples of water soluble polymers include, but are not limited to, polyethylene glycol (PEG), copolymers of ethylene glycol/propylene glycol, carboxymethylcellulose, dextran, polyvinyl alcohol, polyvinyl pyrrolidone, poly-1,3-dioxolane, poly-1,3,6- trioxane, ethylene/maleic anhydride copolymer, polyaminoacids (either homopolymers or random copolymers), and dextran or poly(n-vinyl pyrrolidone)polyethylene glycol, propropylene glycol homopolymers, prolypropylene oxide/ethylene oxide co-polymers, polyoxyethylated polyols (e.g., glycerol), polyvinyl alcohol, and mixtures thereof.
- PEG polyethylene glycol
- copolymers of ethylene glycol/propylene glycol carboxymethylcellulose
- dextran polyvinyl alcohol
- Polyethylene glycol propionaldehyde may have advantages in manufacturing due to its stability in water.
- the polymer may be of any molecular weight, and may be branched or unbranched.
- the number of polymers attached to the antibody may vary, and if more than one polymer are attached, they can be the same or different molecules. In general, the number and/or type of polymers used for derivatization can be determined based on considerations including, but not limited to, the particular properties or functions of the antibody to be improved, whether the antibody derivative will be used in an application under defined conditions, etc. B.
- the antibodies and antigen binding fragments that specifically bind to PfCSP can be conjugated to an agent, such as an effector molecule or detectable marker. Both covalent and noncovalent attachment means may be used.
- an effector molecule and detectable markers can be used, including (but not limited to) toxins and radioactive agents such as 125 I, 32 P, 14 C, 3 H and 35 S and other labels, target moieties and ligands, etc.
- the choice of a particular effector molecule or detectable marker depends on the particular target molecule or cell, and the desired biological effect.
- the procedure for attaching an effector molecule or detectable marker to an antibody or antigen binding fragment varies according to the chemical structure of the effector.
- Polypeptides typically contain a variety of functional groups, such as carboxyl (-COOH), free amine (-NH 2 ) or sulfhydryl (-SH) groups, which are available for reaction with a suitable functional group on a polypeptide to result in the binding of the effector molecule or detectable marker.
- the antibody or antigen binding fragment is derivatized to expose or attach additional reactive functional groups. The derivatization may involve attachment of any suitable linker molecule.
- the linker is capable of forming covalent bonds to both the antibody or antigen binding fragment and to the effector molecule or detectable marker.
- Suitable linkers include, but are not limited to, straight or branched-chain carbon linkers, heterocyclic carbon linkers, or peptide linkers.
- the linkers may be joined to the constituent amino acids through their side chains (such as through a disulfide linkage to cysteine) or the alpha carbon, or through the amino, and/or carboxyl groups of the terminal amino acids.
- a suitable method for attaching a given agent to an antibody or antigen binding fragment or other polypeptide can be determined.
- the antibody or antigen binding fragment can be conjugated with a detectable marker; for example, a detectable marker capable of detection by ELISA, spectrophotometry, flow cytometry, microscopy or diagnostic imaging techniques (such as CT, computed axial tomography (CAT), MRI, magnetic resonance tomography (MTR), ultrasound, fiberoptic examination, and laparoscopic examination).
- detectable markers include fluorophores, chemiluminescent agents, enzymatic linkages, radioactive isotopes and heavy metals or compounds (for example super paramagnetic iron oxide nanocrystals for detection by MRI).
- useful detectable markers include fluorescent compounds, including fluorescein, fluorescein isothiocyanate, rhodamine, 5-dimethylamine-1-napthalenesulfonyl chloride, phycoerythrin, lanthanide phosphors and the like.
- Bioluminescent markers are also of use, such as luciferase, green fluorescent protein (GFP), and yellow fluorescent protein (YFP).
- An antibody or antigen binding fragment can also be conjugated with enzymes that are useful for detection, such as horseradish peroxidase, ⁇ - galactosidase, luciferase, alkaline phosphatase, glucose oxidase and the like.
- enzymes that are useful for detection
- an antibody or antigen binding fragment is conjugated with a detectable enzyme, it can be detected by adding additional reagents that the enzyme uses to produce a reaction product that can be discerned.
- the agent horseradish peroxidase is present, the addition of hydrogen peroxide and diaminobenzidine leads to a colored reaction product, which is visually detectable.
- An antibody or antigen binding fragment may also be conjugated with biotin, and detected through indirect measurement of avidin or streptavidin binding.
- the avidin itself can be conjugated with an enzyme or a fluorescent label.
- the antibody or antigen binding fragment can be conjugated with a paramagnetic agent, such as gadolinium. Paramagnetic agents such as superparamagnetic iron oxide are also of use as labels.
- Antibodies can also be conjugated with lanthanides (such as europium and dysprosium), and manganese.
- An antibody or antigen binding fragment may also be labeled with a predetermined polypeptide epitope recognized by a secondary reporter (such as leucine zipper pair sequences, binding sites for secondary antibodies, metal binding domains, epitope tags).
- the antibody or antigen binding fragment can also be conjugated with a radiolabeled amino acid, for example, for diagnostic purposes.
- the radiolabel may be used to detect PfCSP expressing cells by radiography, emission spectra, or other diagnostic techniques.
- labels for polypeptides include, but are not limited to, the following radioisotopes: 3 H, 14 C, 35 S, 90 Y, 99m Tc, 111 In, 125 I, 131 I.
- the radiolabels may be detected, for example, using photographic film or scintillation counters, fluorescent markers may be detected using a photodetector to detect emitted illumination.
- Enzymatic labels are typically detected by providing the enzyme with a substrate and detecting the reaction product produced by the action of the enzyme on the substrate, and colorimetric labels are detected by simply visualizing the colored label.
- the average number of effector molecule or detectable marker moieties per antibody or antigen binding fragment in a conjugate can range, for example, from 1 to 20 moieties per antibody or antigen binding fragment. In some aspects, the average number of effector molecules or detectable marker moieties per antibody or antigen binding fragment in a conjugate range from about 1 to about 2, from about 1 to about 3, about 1 to about 8; from about 2 to about 6; from about 3 to about 5; or from about 3 to about 4.
- the loading (for example, effector molecule per antibody ratio) of a conjugate may be controlled in different ways, for example, by: (i) limiting the molar excess of effector molecule-linker intermediate or linker reagent relative to antibody, (ii) limiting the conjugation reaction time or temperature, (iii) partial or limiting reducing conditions for cysteine thiol modification, (iv) engineering by recombinant techniques the amino acid sequence of the antibody such that the number and position of cysteine residues is modified for control of the number or position of linker-effector molecule attachments.
- Nucleic acid molecules for example, cDNA or RNA molecules encoding the amino acid sequences of antibodies, antigen binding fragments, and conjugates that specifically bind to PfCSP (such as any one of MAD21-17, MAD21-46, MAD21-53, MAD21-95, MAD21-101, MAD22-17, MAD22-38, MAD22-39, MAD24-01, MAD24-05, or MAD24-52) are provided. Nucleic acids encoding these molecules can readily be produced using the amino acid sequences provided herein (such as the CDR sequences and VH and VL sequences), sequences available in the art (such as framework or constant region sequences), and the genetic code.
- a nucleic acid molecules can encode the V H , the V L , or both the V H and V L (for example in a bicistronic expression vector) of a disclosed antibody or antigen binding fragment.
- the nucleic acid molecules can be expressed in a host cell (such as a mammalian cell) to produce a disclosed antibody or antigen binding fragment.
- Exemplary nucleic acid sequences encoding the heavy and light chain variable regions of the disclosed antibodies are provided as SEQ ID NOs: 73-94.
- the genetic code can be used to construct a variety of functionally equivalent nucleic acid sequences, such as nucleic acids which differ in sequence but which encode the same antibody sequence, or encode a conjugate or fusion protein including the VL and/or VH nucleic acid sequence.
- Nucleic acid molecules encoding the antibodies, antigen binding fragments, and conjugates that specifically bind to PfCSP can be prepared by any suitable method including, for example, cloning of appropriate sequences or by direct chemical synthesis by standard methods. Chemical synthesis produces a single stranded oligonucleotide. This can be converted into double stranded DNA by hybridization with a complementary sequence or by polymerization with a DNA polymerase using the single strand as a template.
- Exemplary nucleic acids can be prepared by cloning techniques. Examples of appropriate cloning and sequencing techniques can be found, for example, in Green and Sambrook (Molecular Cloning: A Laboratory Manual, 4 th ed., New York: Cold Spring Harbor Laboratory Press, 2012) and Ausubel et al. (Eds.) (Current Protocols in Molecular Biology, New York: John Wiley and Sons, including supplements). Nucleic acids can also be prepared by amplification methods. Amplification methods include the polymerase chain reaction (PCR), the ligase chain reaction (LCR), the transcription-based amplification system (TAS), and the self-sustained sequence replication system (3SR).
- PCR polymerase chain reaction
- LCR ligase chain reaction
- TAS transcription-based amplification system
- 3SR self-sustained sequence replication system
- the nucleic acid molecules can be expressed in a recombinantly engineered cell such as bacteria, plant, yeast, insect and mammalian cells.
- the antibodies, antigen binding fragments, and conjugates can be expressed as individual proteins including the V H and/or V L (linked to an effector molecule or detectable marker as needed), or can be expressed as a fusion protein. Any suitable method of expressing and purifying antibodies and antigen binding fragments may be used; non-limiting examples are provided in Al- Rubeai (Ed.), Antibody Expression and Production, Dordrecht; New York: Springer, 2011).
- An immunoadhesin can also be expressed.
- nucleic acids encoding a VH and VL, and immunoadhesin are provided.
- the nucleic acid sequences can optionally encode a leader sequence.
- the V H - and V L -encoding DNA fragments can be operatively linked to another fragment encoding a flexible linker, e.g., encoding the amino acid sequence (Gly 4 -Ser) 3 , such that the V H and VL sequences can be expressed as a contiguous single-chain protein, with the VL and VH domains joined by the flexible linker (see, e.g., Bird et al., Science, 242(4877):423-426, 1988; Huston et al., Proc. Natl. Acad. Sci.
- cleavage site can be included in a linker, such as a furin cleavage site.
- the single chain antibody may be monovalent, if only a single VH and VL are used, bivalent, if two VH and VL are used, or polyvalent, if more than two VH and VL are used. Bispecific or polyvalent antibodies may be generated that bind specifically to PfCSP and another antigen.
- the encoded VH and VL optionally can include a furin cleavage site between the VH and VL domains.
- One or more DNA sequences encoding the antibodies, antigen binding fragments, or conjugates can be expressed in vitro by DNA transfer into a suitable host cell.
- the cell may be prokaryotic or eukaryotic. Numerous expression systems available for expression of proteins including E.
- coli other bacterial hosts, yeast, and various higher eukaryotic cells such as the COS, CHO, HeLa and myeloma cell lines, can be used to express the disclosed antibodies and antigen binding fragments. Methods of stable transfer, meaning that the foreign DNA is continuously maintained in the host may be used. Hybridomas expressing the antibodies of interest are also encompassed by this disclosure.
- the expression of nucleic acids encoding the antibodies and antigen binding fragments described herein can be achieved by operably linking the DNA or cDNA to a promoter (which is either constitutive or inducible), followed by incorporation into an expression cassette.
- the promoter can be any promoter of interest, including a cytomegalovirus promoter.
- an enhancer such as a cytomegalovirus enhancer
- the cassettes can be suitable for replication and integration in either prokaryotes or eukaryotes.
- Typical expression cassettes contain specific sequences useful for regulation of the expression of the DNA encoding the protein.
- the expression cassettes can include appropriate promoters, enhancers, transcription and translation terminators, initiation sequences, a start codon (i.e., ATG) in front of a protein-encoding gene, splicing signals for introns, sequences for the maintenance of the correct reading frame of that gene to permit proper translation of mRNA, and stop codons.
- the vector can encode a selectable marker, such as a marker encoding drug resistance (for example, ampicillin or tetracycline resistance).
- a selectable marker such as a marker encoding drug resistance (for example, ampicillin or tetracycline resistance).
- drug resistance for example, ampicillin or tetracycline resistance
- expression cassettes which contain, for example, a strong promoter to direct transcription, a ribosome binding site for translational initiation (e.g., internal ribosomal binding sequences), and a transcription/translation terminator.
- a promoter such as the T7, trp, lac, or lambda promoters, a ribosome binding site, and preferably a transcription termination signal.
- control sequences can include a promoter and/or an enhancer derived from, for example, an immunoglobulin gene, HTLV, SV40 or cytomegalovirus, and a polyadenylation sequence, and can further include splice donor and/or acceptor sequences (for example, CMV and/or HTLV splice acceptor and donor sequences).
- the cassettes can be transferred into the chosen host cell by any suitable method such as transformation or electroporation for E. coli and calcium phosphate treatment, electroporation or lipofection for mammalian cells.
- Cells transformed by the cassettes can be selected by resistance to antibiotics conferred by genes contained in the cassettes, such as the amp, gpt, neo and hyg genes.
- Modifications can be made to a nucleic acid encoding a polypeptide described herein without diminishing its biological activity. Some modifications can be made to facilitate the cloning, expression, or incorporation of the targeting molecule into a fusion protein. Such modifications include, for example, termination codons, sequences to create conveniently located restriction sites, and sequences to add a methionine at the amino terminus to provide an initiation site, or additional amino acids (such as poly His) to aid in purification steps.
- the antibodies, antigen binding fragments, and conjugates can be purified according to standard procedures in the art, including ammonium sulfate precipitation, affinity columns, column chromatography, and the like (see, generally, Simpson et al. (Eds.), Basic methods in Protein Purification and Analysis: A Laboratory Manual, New York: Cold Spring Harbor Laboratory Press, 2009).
- the antibodies, antigen binding fragment, and conjugates need not be 100% pure.
- the polypeptides should be substantially free of endotoxin.
- a disclosed antibody such as any one of MAD21-17, MAD21-46, MAD21-53, MAD21-95, MAD21-101, MAD22-17, MAD22-38, MAD22-39, MAD24-01, MAD24-05, or MAD24-52
- the methods can be used pre-exposure or post-exposure.
- P. falciparum infection does not need to be completely eliminated or inhibited for the method to be effective. For example, the method can decrease P.
- falciparum infection by a desired amount, for example by at least 10%, at least 20%, at least 50%, at least 60%, at least 70%, at least 80%, at least 90%, at least 95%, at least 98%, or even at least 100% (elimination or prevention of detectable P. falciparum infection) as compared to P. falciparum infection in the absence of the treatment.
- the subject can also be treated with an effective amount of an additional agent, such as anti-malaria agent.
- administration of an effective amount of a disclosed antibody, antigen binding fragment, conjugate, or nucleic acid molecule inhibits the establishment of P. falciparum infection and/or subsequent P. falciparum disease progression in a subject, which can encompass any statistically significant reduction in P.
- Antibodies and antigen binding fragments thereof are typically administered by intravenous infusion. Doses of the antibody or antigen binding fragment vary, but generally range between about 0.5 mg/kg to about 50 mg/kg, such as a dose of about 1 mg/kg, about 5 mg/kg, about 10 mg/kg, about 20 mg/kg, about 30 mg/kg, about 40 mg/kg, or about 50 mg/kg. In some aspects, the dose of the antibody or antigen binding fragment can be from about 0.5 mg/kg to about 5 mg/kg, such as a dose of about 1 mg/kg, about 2 mg/kg, about 3 mg/kg, about 4 mg/kg or about 5 mg/kg.
- the antibody or antigen binding fragment is administered according to a dosing schedule determined by a medical practitioner. In some examples, the antibody or antigen binding fragment is administered weekly, every two weeks, every three weeks or every four weeks. In some aspects, the method of inhibiting P. falciparum infection in a subject further comprises administration of one or more additional agents to the subject. Additional agents of interest include, but are not limited to, anti-malaria agents. In some aspects, the method of inhibiting P.
- falciparum infection in a subject comprises administration of a first antibody that specifically binds to PfCSP as disclosed herein (such as any one MAD21-17, MAD21-46, MAD21-53, MAD21-95, MAD21-101, MAD22-17, MAD22-38, MAD22-39, MAD24-01, MAD24-05, or MAD24-52) and a second antibody that that specifically binds to PfCSP (such as L9 or CIS43).
- a subject is administered DNA or RNA encoding a disclosed antibody to provide in vivo antibody production, for example using the cellular machinery of the subject. Any suitable method of nucleic acid administration may be used; non-limiting examples are provided in U.S.
- U.S. Patent No.5,880,103 describes several methods of delivery of nucleic acids encoding proteins to an organism.
- One approach to administration of nucleic acids is direct administration with plasmid DNA, such as with a mammalian expression plasmid.
- the nucleotide sequence encoding the disclosed antibody, or antigen binding fragments thereof can be placed under the control of a promoter to increase expression.
- the methods include liposomal delivery of the nucleic acids. Such methods can be applied to the production of an antibody, or antigen binding fragments thereof.
- a disclosed antibody or antigen binding fragment is expressed in a subject using the pVRC8400 vector (described in Barouch et al., J. Virol., 79(14), 8828-8834, 2005).
- a subject such as a human subject at risk of P. falciparum infection
- the AAV viral vector is designed for expression of the nucleic acid molecules encoding a disclosed antibody or antigen binding fragment, and administration of the effective amount of the AAV viral vector to the subject leads to expression of an effective amount of the antibody or antigen binding fragment in the subject.
- Non-limiting examples of AAV viral vectors that can be used to express a disclosed antibody or antigen binding fragment in a subject include those provided in Johnson et al., Nat. Med., 15(8):901-906, 2009 and Gardner et al., Nature, 519(7541):87-91, 2015.
- a nucleic acid encoding a disclosed antibody, or antigen binding fragment thereof is introduced directly into tissue.
- the nucleic acid can be loaded onto gold microspheres by standard methods and introduced into the skin by a device such as Bio-Rad’s HELIOS ⁇ Gene Gun.
- the nucleic acids can be “naked,” consisting of plasmids under control of a strong promoter.
- the DNA is injected into muscle, although it can also be injected directly into other sites.
- Dosages for injection are usually around 0.5 ⁇ g/kg to about 50 mg/kg, and typically are about 0.005 mg/kg to about 5 mg/kg (see, e.g., U.S. Patent No.5,589,466).
- Single or multiple administrations of a composition including a disclosed PfCSP-specific antibody, antigen binding fragment, conjugate, or nucleic acid molecule encoding such molecules can be administered depending on the dosage and frequency as required and tolerated by the patient.
- the dosage can be administered once, but may be applied periodically until either a desired result is achieved or until side effects warrant discontinuation of therapy. Generally, the dose is sufficient to inhibit P.
- the dosage normally lies within a range of circulating concentrations that include the ED 50 , with little or minimal toxicity.
- the dosage can vary within this range depending upon the dosage form employed and the route of administration utilized.
- the effective dose can be determined from cell culture assays and animal studies.
- the PfCSP-specific antibody, antigen binding fragment, conjugate, or nucleic acid molecule encoding such molecules, or a composition including such molecules can be administered to subjects in various ways, including local and systemic administration, such as, e.g., by injection subcutaneously, intravenously, intra-arterially, intraperitoneally, intramuscularly, intradermally, or intrathecally.
- the antibody, antigen binding fragment, conjugate, or nucleic acid molecule encoding such molecules, or a composition including such molecules is administered by a single subcutaneous, intravenous, intra-arterial, intraperitoneal, intramuscular, intradermal or intrathecal injection once a day.
- the antibody, antigen binding fragment, conjugate, or nucleic acid molecule encoding such molecules, or a composition including such molecules can also be administered by direct injection at or near the site of disease.
- a further method of administration is by osmotic pump (e.g., an Alzet pump) or mini-pump (e.g., an Alzet mini-osmotic pump), which allows for controlled, continuous and/or slow-release delivery of the antibody, antigen binding fragment, conjugate, or nucleic acid molecule encoding such molecules, or a composition including such molecules, over a pre-determined period.
- the osmotic pump or mini-pump can be implanted subcutaneously, or near a target site. 2.
- compositions are provided that include one or more of the PfCSP-specific antibody, antigen binding fragment, conjugate, or nucleic acid molecule encoding such molecules, that are disclosed herein in a pharmaceutically acceptable carrier.
- the composition comprises an antibody as provided herein (such as any one of MAD21-17, MAD21-46, MAD21-53, MAD21-95, MAD21-101, MAD22-17, MAD22-38, MAD22-39, MAD24-01, MAD24-05, or MAD24-52).
- the composition comprises an antibody as provided herein (such as any one of MAD21-17, MAD21-46, MAD21-53, MAD21-95, MAD21-101, MAD22-17, MAD22-38, MAD22-39, MAD24-01, MAD24-05, or MAD24-52) and one or more additional PfCSP-specific antibody, such as L9 or CIS43 or 317.
- the compositions are useful, for example, for example, for the inhibition or detection of a P. falciparum infection.
- the compositions can be prepared in unit dosage forms for administration to a subject. The amount and timing of administration are at the discretion of the administering physician to achieve the desired purposes.
- the PfCSP-specific antibody, antigen binding fragment, conjugate, or nucleic acid molecule encoding such molecules can be formulated for systemic or local administration.
- the PfCSP-specific antibody, antigen binding fragment, conjugate, or nucleic acid molecule encoding such molecules is formulated for parenteral administration, such as intravenous administration.
- the antibody, antigen binding fragment, or conjugate thereof, in the composition is at least 70% (such as at least 75%, at least 80%, at least 85%, at least 90%, at least 95%, at least 96%, at least 97%, at least 98% or at least 99%) pure.
- the composition contains less than 10% (such as less than 5%, less than 4%, less than 3%, less than 2%, less than 1%, less than 0.5%, or even less) of macromolecular contaminants, such as other mammalian (e.g., human) proteins.
- the compositions for administration can include a solution of the PfCSP-specific antibody, antigen binding fragment, conjugate, or nucleic acid molecule encoding such molecules, dissolved in a pharmaceutically acceptable carrier, such as an aqueous carrier.
- a pharmaceutically acceptable carrier such as an aqueous carrier.
- aqueous carriers can be used, for example, buffered saline and the like. These solutions are sterile and generally free of undesirable matter.
- These compositions may be sterilized by any suitable technique.
- compositions may contain pharmaceutically acceptable auxiliary substances as required to approximate physiological conditions such as pH adjusting and buffering agents, toxicity adjusting agents and the like, for example, sodium acetate, sodium chloride, potassium chloride, calcium chloride, sodium lactate and the like.
- concentration of antibody in these formulations can vary widely, and will be selected primarily based on fluid volumes, viscosities, body weight and the like in accordance with the particular mode of administration selected and the subject’s needs.
- a typical composition for intravenous administration comprises about 0.01 to about 30 mg/kg of antibody or antigen binding fragment or conjugate per subject per day (or the corresponding dose of a conjugate including the antibody or antigen binding fragment).
- the composition can be a liquid formulation including one or more antibodies, antigen binding fragments (such as an antibody or antigen binding fragment that specifically binds to PfCSP), in a concentration range from about 0.1 mg/ml to about 20 mg/ml, or from about 0.5 mg/ml to about 20 mg/ml, or from about 1 mg/ml to about 20 mg/ml, or from about 0.1 mg/ml to about 10 mg/ml, or from about 0.5 mg/ml to about 10 mg/ml, or from about 1 mg/ml to about 10 mg/ml.
- antigen binding fragments such as an antibody or antigen binding fragment that specifically binds to PfCSP
- Antibodies, or an antigen binding fragment thereof or a conjugate or a nucleic acid encoding such molecules can be provided in lyophilized form and rehydrated with sterile water before administration, although they are also provided in sterile solutions of known concentration.
- the antibody solution, or an antigen binding fragment or a nucleic acid encoding such antibodies or antigen binding fragments can then be added to an infusion bag containing 0.9% sodium chloride, USP, and typically administered at a dosage of from 0.5 to 15 mg/kg of body weight.
- Antibodies, antigen binding fragments, conjugates, or a nucleic acid encoding such molecules can be administered by slow infusion, rather than in an intravenous push or bolus.
- a higher loading dose is administered, with subsequent, maintenance doses being administered at a lower level.
- an initial loading dose of 4 mg/kg may be infused over a period of some 90 minutes, followed by weekly maintenance doses for 4-8 weeks of 2 mg/kg infused over a 30-minute period if the previous dose was well tolerated.
- Controlled-release parenteral formulations can be made as implants, oily injections, or as particulate systems.
- Particulate systems include microspheres, microparticles, microcapsules, nanocapsules, nanospheres, and nanoparticles.
- Microcapsules contain the active protein agent, such as a cytotoxin or a drug, as a central core. In microspheres, the active protein agent is dispersed throughout the particle. Particles, microspheres, and microcapsules smaller than about 1 ⁇ m are generally referred to as nanoparticles, nanospheres, and nanocapsules, respectively.
- Capillaries have a diameter of approximately 5 ⁇ m so that only nanoparticles are administered intravenously. Microparticles are typically around 100 ⁇ m in diameter and are administered subcutaneously or intramuscularly. See, for example, Kreuter, Colloidal Drug Delivery Systems, J. Kreuter (Ed.), New York, NY: Marcel Dekker, Inc., pp.219-342, 1994; and Tice and Tabibi, Treatise on Controlled Drug Delivery: Fundamentals, Optimization, Applications, A. Kydonieus (Ed.), New York, NY: Marcel Dekker, Inc., pp.315-339, 1992. Polymers can be used for ion-controlled release of the antibody compositions disclosed herein.
- Any suitable polymer may be used, such as a degradable or nondegradable polymeric matrix designed for use in controlled drug delivery.
- hydroxyapatite has been used as a microcarrier for controlled release of proteins.
- liposomes are used for controlled release as well as drug targeting of the lipid-capsulated drug. 3.
- Methods of detection and diagnosis Methods are also provided for the detection of the presence of PfCSP in vitro or in vivo.
- the presence of PfCSP is detected in a biological sample from a subject, and can be used to identify a subject with P. falciparum infection.
- the sample can be any sample, including, but not limited to, tissue from biopsies, autopsies and pathology specimens.
- Biological samples also include sections of tissues, for example, frozen sections taken for histological purposes. Biological samples further include body fluids, such as blood, serum, plasma, sputum, spinal fluid or urine.
- the method of detection can include contacting a cell or sample, with an antibody or antigen binding fragment that specifically binds to PfCSP, or conjugate thereof (e.g., a conjugate including a detectable marker) under conditions sufficient to form an immune complex, and detecting the immune complex (e.g., by detecting a detectable marker conjugated to the antibody or antigen binding fragment.
- the antibody or antigen binding fragment is directly labeled with a detectable marker.
- the antibody that binds P is directly labeled with a detectable marker.
- the primary antibody is unlabeled and a secondary antibody or other molecule that can bind the primary antibody is utilized for detection.
- the secondary antibody is chosen that is able to specifically bind the specific species and class of the first antibody.
- the first antibody is a human IgG
- the secondary antibody may be an anti-human-IgG.
- Other molecules that can bind to antibodies include, without limitation, Protein A and Protein G, both of which are available commercially.
- Suitable labels for the antibody, antigen binding fragment or secondary antibody are known and described above, and include various enzymes, prosthetic groups, fluorescent materials, luminescent materials, magnetic agents and radioactive materials.
- the disclosed antibodies or antigen binding fragments thereof are used to test vaccines.
- a vaccine composition including a PfCSP or fragment thereof assumes a conformation including the epitope of a disclosed antibody.
- the method comprises contacting a sample containing the vaccine, such as a PfCSP immunogen, with a disclosed antibody or antigen binding fragment under conditions sufficient for formation of an immune complex, and detecting the immune complex, to detect the vaccine with an PfCSP immunogen including the epitope in the sample.
- the detection of the immune complex in the sample indicates that vaccine component, such as a PfCSP immunogen assumes a conformation capable of binding the antibody or antigen binding fragment.
- Example 1 Highly Protective Anti-Malarial Antibodies This example illustrates the design and assessment of antibodies to PfCSP with protective potency and that bind to a novel PfCSP epitope. Identification of Pf sporozoite specific mAbs. An antigen-agnostic approach was used to isolate functional monoclonal antibodies (mAbs) that target novel epitopes on the Plasmodium falciparum (Pf) sporozoite.
- mAbs monoclonal antibodies
- Plasma IgG reactivity towards whole Pf sporozoites was assessed in parallel for both the blocked and unblocked plasma samples and is represented as the percentage of IgG positive sporozoites. In total, 4 vaccinees showed high IgG reactivity to sporozoites after blocking of plasma (FIG.1). These individuals were therefore selected as donors of interest for the study. Binding of plasma IgG to sporozoites or PfCSP antigens was measured with the IntelliCyt iQue Screener flow cytometer and FACS data were analyzed with FlowJo (Version 10.8.1. Ashland, OR).
- MBCs memory B cells isolated from the four donors of interest and exported MBCs that secreted mAbs which bound to the surface of Pf sporozoites but not to recombinant, mammalian cell expressed PfCSP.
- the respective antibodies were recombinantly expressed as IgG1 mAbs and titrated against wild-type Pf sporozoites. Binding of recombinant mAbs to sporozoites was measured with the IntelliCyt iQue Screener flow cytometer and FACS data were analysed with FlowJo (Version 10.8.1. Ashland, OR).
- the panel consisted of transgenic Pb sporozoites that express the full PfCSP sequence (Pb PfCSP ); transgenic Pb sporozoites that express PfCSP which lacks the ADGNPDP residues in the junctional region (Pb-PfCSP JRKO); chimeric Pb sporozoites with 12 Pf-NANP repeats inserted into the PbCSP open reading frame (Pb CSP Pf-NANP 12 ); and chimeric Pb sporozoites with 4 Pf-NANP repeats and the ADGNPDP residues from the Pf junctional region inserted into the PbCSP open reading frame (Pb CSP Pf-NANP 4-5 ).
- the eleven mAbs were titrated against recombinant peptides that recapitulate sporozoite-mediated, post- translational modifications present on PfCSP. Binding of recombinant mAbs to the antigen conjugated beads was measured by flow cytometry in a multiplexed bead-based assay. All mAbs bound to the peptide bearing an N-terminal glutamine which represents the proposed PfCSP N-terminus post-cleavage (FIG.10).
Landscapes
- Health & Medical Sciences (AREA)
- Chemical & Material Sciences (AREA)
- Tropical Medicine & Parasitology (AREA)
- Organic Chemistry (AREA)
- Medicinal Chemistry (AREA)
- Veterinary Medicine (AREA)
- Life Sciences & Earth Sciences (AREA)
- General Health & Medical Sciences (AREA)
- Molecular Biology (AREA)
- Genetics & Genomics (AREA)
- Biophysics (AREA)
- Proteomics, Peptides & Aminoacids (AREA)
- Biochemistry (AREA)
- Immunology (AREA)
- Chemical Kinetics & Catalysis (AREA)
- General Chemical & Material Sciences (AREA)
- Nuclear Medicine, Radiotherapy & Molecular Imaging (AREA)
- Pharmacology & Pharmacy (AREA)
- Animal Behavior & Ethology (AREA)
- Public Health (AREA)
- Peptides Or Proteins (AREA)
Abstract
Antibodies and antigen binding fragments that specifically bind to P. falciparum circumsporozoite protein are disclosed. Nucleic acids encoding these antibodies, vectors and host cells are also provided. The disclosed antibodies, antigen binding fragments, nucleic acids and vectors can be used, for example, to inhibit a P. falciparum infection.
Description
NEUTRALIZING ANTIBODIES TO PLASMODIUM FALCIPARUM CIRCUMSPOROZOITE PROTEIN AND THEIR USE CROSS REFERENCE TO RELATED APPLICATIONS This claims the benefit of U.S. Provisional Application No.63/409,016, filed September 22, 2022, which is incorporated by reference herein in its entirety. FIELD This relates to monoclonal antibodies and antigen binding fragments that specifically bind to Plasmodium falciparum (P. falciparum or Pf) circumsporozoite protein (PfCSP) and their use, for example, in methods of inhibiting P. falciparum infection in a subject. REFERENCE TO AN ELECTRONIC SEQUENCE LISTING The contents of the electronic sequence listing, entitled “Sequence.xml”, which has a size of 157,652 bytes and a Date of Creation of September 21, 2023, is herein incorporated by reference in its entirety. BACKGROUND Malaria ranks as one of the world’s deadliest infectious diseases, with approximately 300 million cases per year. Malaria in humans is caused by five species of the Plasmodium parasite: P. falciparum, P. vivax, P. ovale, P. knowlesi and P. malariae. P. falciparum causes the most severe form of malaria disease, leading to the death of about ~ 500,000 people annually, most of whom are young children. Each of the Plasmodium species that infect humans is transmitted through the bite of an infected female Anopheles mosquito, which introduces Plasmodium sporozoites into the bloodstream of the human host. The major protein on the surface of the infecting P. falciparum sporozoites is the circumsporozoite protein (PfCSP) and provides a major target for antibodies and vaccines. The sporozoites rapidly reach the liver where they are sequestered by hepatocytes and undergo asexual expansion. One week later, the infected hepatocytes rupture and release mature parasites, the merozoites. These then begin the erythrocytic phase of malaria by attaching to and invading red blood cells, or erythrocytes. The invasion of the erythrocytes by the malarial parasites leads to malarial pathogenesis and clinical infection. While there is no FDA approved vaccine for malaria, the World Health Organization (WHO) recently approved the RTS,S vaccine, which has modest efficacy against malaria. Moreover, malarial parasites are increasingly becoming resistant to antimalarial drugs used to treat the disease. Therefore, preventive interventions to inhibit malaria infection are urgently needed for limiting morbidity, mortality, and ultimately eliminating malaria.
SUMMARY This disclosure provides monoclonal antibodies and antigen binding fragments directed against PfCSP. In an example, data shows that the disclosed antibodies target a novel epitope on PfCSP, and that passive transfer of the disclosed antibodies confers sterile protection in an animal model of P. falciparum infection. In some aspects, a monoclonal antibody is provided that comprises a heavy chain variable region (VH) and a light chain variable region (VL) comprising a heavy chain complementarity determining region (HCDR)1, a HCDR2, and a HCDR3, and a light chain complementarity determining region (LCDR)1, a LCDR2, and a LCDR3 of the VH and VL set forth as SEQ ID NOs: 1 and 2, respectively (MAD21-17), SEQ ID NOs: 3 and 4, respectively (MAD21-46), SEQ ID NOs: 5 and 6, respectively (MAD21-53), SEQ ID NOs: 7 and 8, respectively (MAD21-95), SEQ ID NOs: 9 and 10, respectively (MAD21-101), SEQ ID NOs: 11 and 12, respectively (MAD22-17), SEQ ID NOs: 13 and 14, respectively (MAD22-38), SEQ ID NOs: 15 and 16, respectively (MAD22-39), SEQ ID NOs: 17 and 18, respectively (MAD24-01), SEQ ID NOs: 19 and 20, respectively (MAD24-05), SEQ ID NOs: 21 and 22, respectively (MAD24-52). In several aspects, the monoclonal antibody specifically binds to PfCSP and neutralizes P. falciparum. Also disclosed are compositions including the antibodies and antigen binding fragments, nucleic acids encoding the antibodies and antigen binding fragments, expression vectors comprising the nucleic acids, and isolated host cells that comprise the nucleic acids. In several aspects, the nucleic acid molecule encoding a disclosed antibody or antigen binding fragment can be a cDNA or RNA molecule that encodes the antibody or antigen binding fragment. In additional aspects, the nucleic acid molecule can be a bicistronic expression construct encoding the VH and VL of the antibody or antigen binding fragment. The disclosed antibodies and antigen binding fragments potently neutralize PfCSP expressed on infectious sporozoites in vivo. Accordingly, a method is disclosed for inhibiting (including preventing) P. falciparum infection in a subject. The method comprises administering an effective amount (that is, an amount effective to inhibit P. falciparum infection in a subject) of one or more of the disclosed antibodies, antigen binding fragments, nucleic acid molecules, vectors, or compositions, to the subject, such as a subject at risk of or having a P. falciparum infection. The antibodies, antigen binding fragments, nucleic acid molecules, vectors, and compositions disclosed herein can be used for a variety of additional purposes, such as for diagnosing P. falciparum infection in a subject, or detecting P. falciparum in a sample. The foregoing and other features and advantages of this disclosure will become more apparent from the following detailed description of several aspects, which proceeds with reference to the accompanying figures.
BRIEF DESCRIPTION OF THE FIGURES FIG.1. Selected vaccinees show high IgG reactivity to sporozoites after blocking plasma with recombinant PfCSP. The data represents polyclonal IgG binding to whole P. falciparum sporozoites after incubation of each plasma sample with recombinant PfCSP. FIGs.2A-2B. mAbs bind to sporozoites that express Pf CSP. Titration curves of mAb binding to wild-type P. falciparum sporozoites, wild-type P. berghei sporozoites and transgenic P. berghei sporozoites that express PfCSP. FIG.3. Disclosed mAbs recognize an ~60kDa sporozoite expressed protein. Western blot of P. falciparum sporozoite lysate probed with representative isolated mAbs MAD21-46, MAD22-38 and MAD21-101 as well as known PfCSP mAbs CIS43 and mAb10. The HIV-1 mAb VRC01 was used as a negative control. FIGs.4A-4I. Disclosed mAbs do not bind canonical PfCSP epitopes. Titration curves of mAb binding to (FIGs.4A-4B) recombinant Pf CSP, (FIGs.4C-4E) Pf (NANP)9 repeat peptide, (FIGs.4F-4G) Pf C-terminus and (FIGs.4H-4I) Pf N-terminus. CIS43 was included as a positive control where indicated. FIG.5. PfCSP peptide scanning analysis. ELISA analysis of disclosed mAbs binding to a panel of 15-mer overlapping peptides that span the length of PfCSP. FIG.6. Disclosed mAbs do not bind recombinant forms of PfCSP. ELISA analysis of representative mAb binding to PfCSP produced by mammalian, bacterial, and yeast expression systems. FIGs.7A-7D. A subset of mAbs show protection in an in vivo model of P. falciparum liver invasion. Liver burden (total flux of bioluminescence, photons/sec) was assessed in mice 40h post-infection (n=5 mice per group). The indicated doses of mAbs were administered 2 hours before IV challenge with 2,000 PbPfCSP-GFP/Luc sporozoites. FIG.8A-8B. The identified mAbs recognize an epitope within the Pf-CSP junctional region and do not show cross-reactivity to the major repeat (NANP) sequences. FIG.8A shows the sequences of the PbPfCSP (partial), Pb-PfCSP JRKO (partial), Pb CSP Pf-NANP 12 (partial) and Pb CSP Pf-NANP 4- 5’ (partial) amino acid sequences. SEQ ID NOs: 32, 161, 162 and 163 are shown. FIG.8B shows results from enzyme linked immunosorbent (ELISA) assays. The findings suggest that the mAb epitope lies within the junction region and that the disclosed mAbs do not cross react with Pf- NANP repeats. FIGS.9A-9B. The disclosed mAbs recognize a proteolytically processed form of Pf-CSP. proteins in sporozoite lysate were resolved and subsequent Western blots were developed using representative mAbs (MAD22-39, MAD22-38, MAD24-52, MAD24-01 and MAD21-101), and anti-PfCSP control mAbs mAb10 and 5D5 (Espinosa et al., J Infect Dis 212, 1111-1119.10.1093/infdis/jiv154, 2015). The results confirm that the disclosed mAbs target a proteolytically processed form of PfCSP. FIG.10. The disclosed mAbs preferentially bind Pf-CSP peptides with a truncated N-terminus and pyroglutamic acid modification. Binding of the disclosed mAbs to the antigen conjugated beads was measured by flow cytometry in a multiplexed bead-based assay. All mAbs bound to the peptide bearing an
N-terminal glutamine which represents the proposed PfCSP N-terminus post-cleavage, and binding was increased towards the peptide bearing an N-terminal pyroglutamic acid. The results evidence that the mAbs target a novel epitope on PfCSP which is dependent upon sporozoite-mediated cleavage and pyroglutamic acid modification of PfCSP. SEQ ID NOs: 164 and 165 are shown. SEQUENCES The nucleic and amino acid sequences listed in the accompanying sequence listing are shown using standard letter abbreviations for nucleotide bases, and three letter code for amino acids, as defined in 37 C.F.R.1.822. Only one strand of each nucleic acid sequence is shown, but the complementary strand is understood as included by any reference to the displayed strand. SEQ ID NO: 1 is the amino acid sequence of the MAD21-17 VH. QVQLQESGPGQVKPSETLSLTCTVSGASVTLDYWSWIRQTPERGLEWIGYISYTGRTNYNPSLKSRVTISTD TAKNQVSLRLTSVTAADTAVYYCTRDRSECADGSCFGYVMAVWGHGTTVIVSS SEQ ID NO: 2 is the amino acid sequence of the MAD21-17 VL. DIVMTQSPESLAVSLGERATINCKSNQSLLYSSTNKNYLVWYQQKSGQPPKLLIYWASTRESGVPDRFSGSG SGTDFTLTISSLQAEDVAVYYCQQYYISPLTFGGGTTVEIK SEQ ID NO: 3 is the amino acid sequence of the MAD21-46 VH. QVQLQESGPGQVKPSETLSLTCSVSGASVTVDYWSWIRQTPERGLEWIGYISYTGRTNYNPSLKSRVTISTD TAKNQVSLKLTSVTAADTAVYYCTRDRSECTDGLCFAYVMDVWGHGTTVTVSS SEQ ID NO: 4 is the amino acid sequence of the MAD21-46 VL. DIVMTQSPESLAVSLGERATINCKSNQSLLYSSTNKNYLVWYQQKPGQAPKMLIYWASTRESGVPDRFSGSG SGTDFTLTISSLQAEDVAVYYCQQYYISPLTFGGGTRVEIK SEQ ID NO: 5 is the amino acid sequence of the MAD21-53 VH. QVQLQESGPGQVKPSETLSLTCTVSGASVTVDYWSWIRQTPERGLEWIGYISYTGRTNYNPSLKSRVTISTD TAKNQVSLKLTSVTAADTAVYYCTRDRSECSDGLCFAYVMDVWGHGTTVTVSS SEQ ID NO: 6 is the amino acid sequence of the MAD21-53 VL. DIVMTQSPESLAVSLGERATINCKSNQSLLYSSTNKNYLVWYQQKPGQPPKLLFYWASTRESGVPDRFSGSG SGTDFTLTISSLQAEDVAVYYCQQYYISPLTFGGGTRVEIK
SEQ ID NO: 7 is the amino acid sequence of the MAD21-95 VH. QVQLQESGPGQVKPSETLSLTCTVSGASVTLDYWSWIRQTPERGLEWIGYISYTGRTNYNPSLKSRVTISTD TAKNQVSLRLTSVTAADTAVYYCTRDRSECADGSCFGYVMAVWGHGTTVIVSS SEQ ID NO: 8 is the amino acid sequence of the MAD21-95 VL. DIVMTQSPESLAVSLGERATINCKSNQSLLYSPTNKNYLVWYQQKSGQPPKLLIYWASTRESGVPDRFSGSG SGTDFTLTISSLQAEDVAVYYCQQYYISPLTFGGGTTVEIK SEQ ID NO: 9 is the amino acid sequence of the MAD21-101 VH. QVQLQESGPGQVKPSETLSLTCSVSGASVTVDYWSWIRQTPERGLEWIGYISYTGRTNYNPSLKSRVTISTD TAKNQVSLKLSSVTAADTAVYYCTRDRSECPDGSCFAYVMAVWGHGTTVTVSS SEQ ID NO: 10 is the amino acid sequence of the MAD21-101 VL. DIVMTQSPESLAVSLGERATINCKSNQSLLYSSTNKNYLVWYLQKPGQPPKLLIYWASTRESGVPDRFSGSG SGTDFTLTISSLQAEDVAVYYCQQYYLSPLTFGGGTTVEIK SEQ ID NO: 11 is the amino acid sequence of the MAD22-17 VH. EVQVLESGGGLVQPGGSLKLSCAASGFTFSGSAMHWVRQASGKGLEWVGRIRNKANNYATAYAASVKGRFTV SRDDSENTAYLQMDSLKTEDTAVYYCTRLQDFGDFYGMDVWGPGTTVTVSS SEQ ID NO: 12 is the amino acid sequence of the MAD22-17 VL. DIVMTQSPDSLAVSLGERATINCKSSQSVLYSSNNKNYLAWFQQKLGQPPKLLIYWASTRESGVPDRFSGSG SGTDFTLTISSLQTEDVAVYYCQQYYRSPLTFGGGTKVEIK SEQ ID NO: 13 is the amino acid sequence of the MAD22-38 VH. QLQLQESGPGLVKPAETLSLTCTVSGGSISSSSYYWGWIRQPPGKGLEWIGSIYYTGSTYYNPSLKSRVTIS VDTSKNQFSLKLRSVTAADTAVYYCASIGWAVPGKRYYFDYWGQGTLVTVSS SEQ ID NO: 14 is the amino acid sequence of the MAD22-38 VL. DIQMTQSPSSLSASVGDRVTITCRANHSISSYLNWYQQKPGKAPKLLIYAASSLQSGVPSfRFSGSGSGTDF TLTISSLQPEDFATYYCQQSYYTPWTFGPGTKVDIK SEQ ID NO: 15 is the amino acid sequence of the MAD22-39 VH. QLQLQESGPGLVKPSETLSLTCTVSGGSINNSSYYWGWIRQPPGKGLEWIGSIFYSGTTYYNPSLKSRVTVS VDTSKNQFSLKLNSVTAADTAVYYCVRVGRVDTATRLWWFDPWGQGTLVTVSS
SEQ ID NO: 16 is the amino acid sequence of the MAD22-39 VL. DIQMTQSPSSLSASVGDRVTITCRASQSIISYLNWYQQKPGQAPKLLIYAASSLQSGIPSRFSGSGSGTDFT LTISSLQPEDFATYYCQQSYYTPWTFGQGTKVEIK SEQ ID NO: 17 is the amino acid sequence of the MAD24-01 VH. QVQLVESGGGLVKPGGSLRLSCAASGFTFSDYYMSWIRQAPGKGLEWVSYIGSSNTYTSYAASVKGRFTISR DNAKNSLYLQMNSLRAEDTAVYYCAREIERSGWVRGNDYWGRGTLVTVSS SEQ ID NO: 18 is the amino acid sequence of the MAD24-01 VL. EIVLTQSPATLSLSPGERATLSCRASQNVSTYLAWYQQKPGQAPRLLMYDASKRATGIPARFSGSGSGTEFT LTISSLEPEDFAVYYCQQRRNWPLTFGGGTKVEIK SEQ ID NO: 19 is the amino acid sequence of the MAD24-05 VH. EVQLVESGGGLVQPGGSLRLSCAASGFSFSTYWMTWVRQAPGKGLEWVATVKQDGSEKYYVDSVKGRFTISR DNAKNSLYLQMNSLRAEDTAVYYCARAGYSDYLYYYAMDVWGQGTTVTVSS SEQ ID NO: 20 is the amino acid sequence of the MAD24-05 VL. QSVLTQPPSASGTPGQRVTISCSGSSSNIGSNYVYWYQHLPGTAPKLLIYRNNQRPSGVPDRISGSKSGTSV SLAISGLRSEDEADYYCAVWDDSLSCWVFGGGTKLTVL SEQ ID NO: 21 is the amino acid sequence of the MAD24-52 VH. QVQLVESGGGVVQPGRSLRLSCAASGFTFSSYGMHWVRQAPGKGLEWVAVIWYDGSNKYYADSVKGRFTISR DNSKNTLYLQMNSLRAEDTAVYYCARVGTLWGDRIGGFDYWGQGTLVTVSS SEQ ID NO: 22 is the amino acid sequence of the MAD24-52 VL. DIVMTQTPLSSPVTLGQPASISCRSSQSLVHSDGNTYLSWLQQRPGQPPRLLIYKISNRFSGVPDRFSGSGA GTDFTLKISRVEAEDVGVYYCMQATQFPLTFGGGTKVEIK SEQ ID NOs: 23-31 are CDR sequences. SEQ ID NO: 32 is a transgenic Plasmodium berghei sporozoite amino acid sequence. SEQ ID NOs: 33-67 are CDR sequences. SEQ ID NO: 68 is the amino acid sequence of the CIS43 VH. QVQLVQSGAEVKKPGASVKVSCKASGYTFTSYAIHWVRQAPGQRLEWMGWIKAGNGNTRYSQKFQDRVTITRDTSTTTA YMELSSLRSEDTAVYYCALLTVLTPDDAFDIWGQGTMVTVSS
SEQ ID NO: 69 is the amino acid sequence of the CIS43 VL. DIVMTQSPDSLAVSLGERATINCKSSQSVLYSSNNKNYLAWYQQKPGQPPNLLIYWASTRQSGVPDRFSGSGSGTDFTL TISSLQAEDVAVYYCHQYYSSPLTFGGGTKVEIK SEQ ID NO: 70 is the amino acid sequence of the L9 VH. QVKLVESGGGVVQPGRSLRLSCEASGFIFSTYGMHWVRQAPGKGLEWVAVIWFDGSNIYYADSVKGRFTISRDNSKNTVF MQMDSLRAEDTAVYYCHRNFYDGSGPFDYWGQGTLVTVSS SEQ ID NO: 71 is the amino acid sequence of the L9 VL. DIQMTQSPSTLSASVGDRVTITCRASQFISRWLAWYQQKPGKAPKLLIYKASSLESGVPSRFSGSGSETHFTLTISSLQP DDVATYYCQEYTSYGRTFGQGTKVEIK SEQ ID NO: 72 is an exemplary amino acid sequence for PfCSP (GenBank Acc. No. CAB38998.2, incorporated by reference herein) MMRKLAILSVSSFLFVEALFQEYQCYGSSSNTRVLNELNYDNAGTNLYNELEMNYYGKQENWYSLKKNSRSLGENDDGN NEDNEKLRKPKHKKLKQPADGNPDPNANPNVDPNANPNVDPNANPNVDPNANPNANPNANPNANPNANPNANPNANPNA NPNANPNANPNANPNANPNANPNANPNANPNANPNANPNVDPNANPNANPNANPNANPNANPNANPNANPNANPNANPN ANPNANPNANPNANPNANPNANPNANPNANPNANPNKNNQGNGQGHNMPNDPNRNVDENANANSAVKNNNNEEPSDKHI KEYLNKIQNSLSTEWSPCSVTCGNGIQVRIKPGSANKPKDELDYANDIEKKICKMEKCSSVFNVVNSSIGLIMVLSFLF LN SEQ ID NO: 73 is an exemplary nucleic acid sequence encoding MAD21-17 VH. caggtgcaactgcaggagtcgggcccaggacaggtgaagccttcggagaccctgtccctcacctgcactgtctctggtgc ctccgtcacacttgactactggagctggatccggcagaccccagagaggggactggagtggattggctatatctcttaca ctggaaggaccaactacaacccatccctcaagagtcgagtcaccatctcaacagacacggccaagaaccaggtctccctg aggttgacctctgtgaccgctgcggacacggccgtgtattattgtacgagagatcgatctgaatgtgctgatggttcatg cttcggctacgttatggccgtctggggccacgggaccacggtcatcgtctcctcag SEQ ID NO: 74 is an exemplary nucleic acid sequence encoding MAD21-17 VL. gacatcgtgatgacccagtctccagagtccctggctgtgtctctgggcgagagggccaccatcaactgcaagtccaatca gagtcttttatacagttccacgaataagaactacttagtttggtaccagcagaagtcaggacagcctcctaagttgctca tttactgggcatctacccgggaatccggggtccctgaccgattcagcggcagcgggtctgggacagatttcactctcacc atcagcagcctgcaggctgaggatgtggcagtttattactgtcagcaatattatattagtccgctcacattcggcggagg gaccacggtggagatcaaac SEQ ID NO: 75 is an exemplary nucleic acid sequence encoding MAD21-46 VH. caggtgcagctgcaggagtcgggcccaggacaggtgaagccttcggagaccctgtccctcacctgcagtgtctctggtgc ctccgtcacagttgactactggagctggatccggcagaccccagagaggggactggagtggattggctatatctcttata ctggaagaaccaactacaacccctccctcaagagtcgagtcaccatatcaacagacacggccaagaaccaggtctccctg
aagttgacctctgtgaccgctgcggacacggccgtgtattattgtacgagagatcgatctgaatgtactgatggtttatg cttcgcctacgttatggacgtctggggccacgggaccacggtcaccgtctcctcag SEQ ID NO: 76 is an exemplary nucleic acid sequence encoding MAD21-46 VL. gacatcgtgatgacccagtctccagagtccctggctgtgtctctgggcgagagggccaccatcaactgcaagtccaatca gagtcttttatacagctccaccaataagaattacttagtttggtaccagcagaaaccaggacaggcccctaagatgctca tttactgggcatctacccgggaatccggggtccctgaccgattcagcggcagcgggtctgggacagatttcactctcacc atcagcagcctgcaggctgaagatgtggcagtttattactgtcagcagtattatattagtccgctcaccttcggcggagg gaccagggtggagatcaaac SEQ ID NO: 77 is an exemplary nucleic acid sequence encoding MAD21-53 VH. caggtgcagctgcaggagtcgggcccaggacaggtgaagccttcggagaccctgtccctcacctgcactgtctctggtgc ctccgtcacagttgactactggagctggatccggcagaccccagagaggggactggagtggattggctatatctcttaca ctggaagaaccaactacaacccctccctcaagagtcgagtcaccatatcaacagacacggccaagaaccaggtctccctg aagttgacctctgtgaccgctgcggacacggccgtgtattattgcacgcgagatcgatctgaatgtagtgatggtttatg cttcgcctacgttatggacgtctggggccacgggaccacggtcaccgtctcctcag SEQ ID NO: 78 is an exemplary nucleic acid sequence encoding MAD21-53 VL. gacatcgtgatgacccagtctccagagtccctggctgtgtctctgggcgagagggccaccatcaactgcaagtccaatca gagtcttttatacagctccaccaataagaactacttagtttggtaccagcagaaaccaggacagccccctaagttgctct tttactgggcatctacccgggagtccggggtccctgaccgattcagcggcagcgggtctgggacagatttcactctcacc atcagcagcctgcaggctgaagatgtggcagtttattactgtcagcagtattatattagtccgctcaccttcggcggagg gaccagggtggagatcaaac SEQ ID NO: 79 is an exemplary nucleic acid sequence encoding MAD21-95 VH. caggtgcaactgcaggagtcgggcccaggacaggtgaagccttcggagaccctgtccctcacctgcactgtctctggtgc ctccgtcacacttgactactggagctggatccggcagaccccagagaggggactggagtggattggctatatctcttaca ctggaaggaccaactacaacccatccctcaagagtcgagtcaccatctcaacagacacggccaagaaccaggtctccctg aggttgacctctgtgaccgctgcggacacggccgtgtattattgtacgagagatcgatctgaatgtgctgatggttcatg cttcggctacgttatggccgtctggggccacgggaccacggtcatcgtctcctcag SEQ ID NO: 80 is an exemplary nucleic acid sequence encoding MAD21-95 VL. gacatcgtgatgacccagtctccagagtccctggctgtgtctctgggcgagagggccaccatcaactgcaagtccaatca gagtcttttatacagtcccacgaataagaactacttagtttggtaccagcagaagtcaggacagcctcctaagttgctca tttactgggcatctacccgggaatccggggtccctgaccgattcagcggcagcgggtctgggacagatttcactctcacc atcagcagcctgcaggctgaggatgtggcagtttattactgtcagcaatattatattagtccgctcacattcggcggagg gaccacggtggagatcaaac SEQ ID NO: 81 is an exemplary nucleic acid sequence encoding MAD21-101 VH. caggtgcagctgcaggagtcgggcccaggacaagtgaagccttcggagaccctgtccctcacctgctctgtctctggtgc ctccgtcacagttgactactggagctggatccggcagaccccagagaggggactggagtggattggctatatctcttaca
ctggaagaaccaactacaacccctccctcaagagtcgagtcaccatatcaacagacacggccaagaaccaggtctccctg aagttgagctctgtgaccgctgcggacacggccgtgtattattgtacgagagatcgatctgaatgtcctgatggttcatg cttcgcctacgttatggccgtctggggccacgggaccacggtcaccgtctcctcag SEQ ID NO: 82 is an exemplary nucleic acid sequence encoding MAD21-101 VL. gacatcgtgatgacccagtctccagagtccctggctgtgtctctgggcgagagggccaccatcaactgcaagtccaatca gagtcttttatacagttccaccaataagaactacttagtttggtacctgcagaaaccaggacagcctcctaagttgctca tttactgggcatctacccgggaatccggggtccctgaccgattcagcggcagcgggtctgggacagatttcactctcacc atcagcagcctgcaggctgaggatgtggcagtttattactgtcagcaatattatcttagtccgctcaccttcggcggagg gaccacggtggagatcaaac SEQ ID NO: 83 is an exemplary nucleic acid sequence encoding MAD22-17 VH. gaggtgcaggtgctggagtccgggggaggcttggtccagcctggggggtccctgaaactctcctgtgcagcctccgggtt caccttcagtggctctgctatgcactgggtccgccaggcttccgggaaagggctggagtgggttggccgtattagaaaca aagctaacaattacgcgacagcatatgctgcgtcggtgaaaggcaggttcaccgtctccagagatgactcagagaacacg gcatatctgcaaatggacagcctgaaaaccgaggacacggccgtgtattactgtactaggctccaagacttcggtgactt ctacggtatggacgtctggggcccagggaccacggtcaccgtctcctcag SEQ ID NO: 84 is an exemplary nucleic acid sequence encoding MAD22-17 VL. gacatcgtgatgacccagtctccagactccctggctgtttctctgggcgagagggccaccatcaactgcaagtccagcca gagtgttttatacagctccaacaataagaactacttagcttggttccagcagaaactaggtcagcctcctaagttgctca tttactgggcatctacccgggaatccggggtccctgaccgattcagtggcagcgggtctgggacagatttcactctcacc atcagcagcctgcagactgaggatgtggcagtttattactgtcagcaatattatcgtagtccgctcactttcggcggagg gaccaaggtggagatcaaac SEQ ID NO: 85 is an exemplary nucleic acid sequence encoding MAD22-38 VH. cagctgcagctgcaggagtcgggcccaggactggtgaagcctgcggagaccctgtccctcacctgcactgtctctggtgg ctccatcagcagtagtagttactactggggctggatccgccagcccccagggaaggggctggagtggattgggagtatct attatactgggagcacctactacaacccgtccctcaagagtcgagtcaccatatccgtagacacgtccaagaaccagttc tccctgaagctgcgctctgtgaccgccgcagacacggctgtgtattactgtgcctctataggatgggcagtgcctggtaa aaggtactactttgactactggggccagggaaccctggtcaccgtctcctcgg SEQ ID NO: 86 is an exemplary nucleic acid sequence encoding MAD22-38 VL. gacatccagatgacccagtctccatcctccctgtctgcatctgtaggagacagagtcaccatcacttgccgggcaaatca cagcattagcagctatttaaattggtatcagcagaaaccagggaaagcccctaagctcctgatctatgctgcatccagtt tgcaaagtggggtcccatcaaggttcagtggcagtggatctgggacagatttcactctcaccatcagcagtctgcaacct gaagattttgcaacttactactgtcaacagagttactataccccatggactttcggccctgggaccaaagtggatatcaa ac
SEQ ID NO: 87 is an exemplary nucleic acid sequence encoding MAD22-39 VH. cagctgcagctgcaggagtcgggcccaggactggtgaagccttcggagaccctgtccctcacctgcactgtctctggtgg ctccatcaacaatagtagttactactggggctggatccgccagcccccagggaaggggctggagtggattgggagtatct tttatagtgggaccacctactacaacccgtccctcaagagtcgagtcaccgtatccgtagacacgtccaagaaccagttc tccctgaagctgaactctgtgaccgccgcagacacggctgtgtattactgtgtgagagtgggccgggtcgatacagctac ccgtctgtggtggttcgacccctggggccagggaaccctggtcaccgtctcctcag SEQ ID NO: 88 is an exemplary nucleic acid sequence encoding MAD22-39 VL. gacatccagatgacccagtctccatcctccctgtctgcatctgtaggagacagagtcaccatcacttgccgggcaagtca gagcattatcagctatttaaattggtatcagcagaaaccagggcaagcccctaagctcctgatctatgctgcatccagtt tgcaaagtgggatcccatcaaggttcagtggcagtggatctgggacagatttcactctcaccatcagcagtctgcaacct gaagattttgcaacttactactgtcaacagagttactataccccgtggacgttcggccaagggaccaaggtggaaatcaa ac SEQ ID NO: 89 is an exemplary nucleic acid sequence encoding MAD24-01 VH. caggtgcagttggtggagtctgggggaggcttggtcaagcctggagggtcccttagactctcctgtgcagcctctggatt caccttcagtgactactacatgagctggatccgccaggctccagggaaggggctggagtgggtttcatacattggtagta gtaacacttacacaagctacgcagcctctgtgaagggccgattcaccatctccagagacaacgccaagaactcactgtat ctgcagatgaacagcctgagagccgaggacacggctgtctattactgtgcgagagaaattgaaaggagtggctgggtccg aggcaatgactactggggccggggaaccctggtcaccgtctcctcag SEQ ID NO: 90 is an exemplary nucleic acid sequence encoding MAD24-01 VL. gaaattgtgttgacacagtctccagccaccctgtctttgtctccaggggaaagagccaccctctcctgcagggccagtca gaatgttagtacctacttagcctggtaccaacagaaacctggccaggctcccaggctcctcatgtatgatgcttccaaga gggccactggcatcccagccaggttcagtggcagtgggtctgggacagagttcactctcactatcagcagcctagagcct gaagattttgcagtttattactgtcagcagcgtcgcaactggcctctcactttcggcggagggaccaaggtagagatcaa ac SEQ ID NO: 91 is an exemplary nucleic acid sequence encoding MAD24-05 VH. gaggtgcagctggtggagtctgggggaggcttggtccagcctggggggtccctgagactctcctgtgcagcctctggatt cagttttagtacctattggatgacctgggtccgccaggctccagggaaggggctggagtgggtggccaccgtaaagcaag atggaagtgagaaatactatgtagactctgtgaagggccgattcaccatctccagagacaacgccaagaactcactgtat ctacaaatgaacagcctgagagccgaggacacggctgtttattactgtgcgagagccggatatagtgactacctttacta ctacgctatggacgtctggggccaagggaccacggtcaccgtctcctcag SEQ ID NO: 92 is an exemplary nucleic acid sequence encoding MAD24-05 VL. cagtctgtgctgactcagccaccctcagcgtctgggacccccgggcagagggtcaccatctcttgttctggaagcagctc caacatcggaagtaattatgtatactggtaccagcatctcccaggaacggcccccaaactcctcatctataggaataatc agcggccctcaggggtccctgaccgaatctctggctccaagtctggcacctcagtctccctggccatcagtgggctccgg tccgaggatgaggctgattattattgtgcagtatgggatgacagcctgagttgttgggtgttcggcggagggaccaagct gaccgtcctag
SEQ ID NO: 93 is an exemplary nucleic acid sequence encoding MAD24-52 VH. caggtgcagctggtggagtctgggggaggcgtggtccagcctgggaggtccctgagactctcctgtgcagcgtctggatt caccttcagtagctatggcatgcactgggtccgccaggctccaggcaaggggctggagtgggtggcagttatatggtatg atggaagtaataaatactatgcagactccgtgaagggccgattcaccatctccagagacaattccaagaacacgctgtat ctgcaaatgaacagcctgagagccgaggacacggctgtgtattactgtgcgagagttggaaccctgtggggtgaccgaat tgggggttttgactactggggccagggaaccctggtcaccgtctcctcag SEQ ID NO: 94 is an exemplary nucleic acid sequence encoding MAD24-52 VL. Gatattgtgatgacccagactccactctcctcacctgtcacccttggacagccggcctccatctcctgcaggtctagtca aagcctcgtacacagtgatggaaacacctacttgagttggcttcagcagaggccaggccagcctccaagactcctaattt ataagatttctaaccggttctctggggtcccagacagattcagtggcagtggggcagggacagatttcacactgaaaatc agcagggtggaagctgaggatgtcggggtttattactgcatgcaagctacacaatttcctctcactttcggcggagggac caaggtggagatcaaac SEQ ID NOs: 95-156 are PfCSP peptide sequences. SEQ ID NO: 157 is the amino acid sequence for the full length heavy chain of MAD21-101. QVQLQESGPGQVKPSETLSLTCSVSGASVTVDYWSWIRQTPERGLEWIGYISYTGRTNYNPSLKSRVTISTDTAKNQVS LKLSSVTAADTAVYYCTRDRSECPDGSCFAYVMAVWGHGTTVTVSSASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYF PEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPC PAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTV LHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWESNGQP ENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK SEQ ID NO: 158 is the amino acid sequence for the full length light chain of MAD21-101. DIVMTQSPESLAVSLGERATINCKSNQSLLYSSTNKNYLVWYLQKPGQPPKLLIYWASTRESGVPDRFSGSGSGTDFTL TISSLQAEDVAVYYCQQYYLSPLTFGGGTTVEIKRTVAAPSVFIFPPSDEQLKSGTASVVCLLNNFYPREAKVQWKVDN ALQSGNSQESVTEQDSKDSTYSLSSTLTLSKADYEKHKVYACEVTHQGLSSPVTKSFNRGEC SEQ ID NO: 159 is the amino acid sequence of the 317 VH. QMQLVESGGGVVQPGRSLRLSCTASGFTFSTYAMHWVRQSPGQGLQWVAVISYHSTNKYYEDSVRGRFTISRDNSKNTLY LQMNSLRAEDTAVYYCARDGYSSSFFDFWGQGTLVTVSS SEQ ID NO: 160 is the amino acid sequence for 317 VL. DIQMTQSPSTLSASVGDRVAITCRASQSISRWLAWYQQQPGKAPKLLMSGASVLESGVPSRFSGSGSGTEFTLTISSLQP DDFATYYCQHYNSYFVTFGQGTKVEIK
SEQ ID NOs: 161-163 are transgenic Plasmodium berghei sporozoites amino acid sequences. SEQ ID NOs: 164-165 are P. falciparum CSP peptide sequences. SEQ ID NO: 166 is the amino acid sequence of the 5D5 VH. QVHLQQSGGEVARPGASVKLSCKASGYTFTGYGLSWVKQRTGQGLEWIGEIYPRSGNTYYNEKFKGKATLTADKSSSTAY MELRSLTSEDSAVYFCARSWGNSSFVYWGQGTLVTVSA SEQ ID NO: 167 is the amino acid sequence of the 5D5 VL. SIVMTQTPKFLLVSAGDRVTITCKASQSVTNDVTWYQQKPGQSPKLLIYYASNRYTGVPDRFTGSGYGTDFTFTISTVQA EDLAVYFCQQDYSSPFTFGSGTKLEIK SEQ ID NO: 168 is the amino acid sequence of the mAb10 VH. EVOLVESGGGVVQPGRSLRLSCEASGFTFSTYGMHWVRQAPGKGLEWVAI IWHDGS KKYHADSVRGRFTISRDNSKNTLYLQMNSLRAEDTAVYFCARVGNYGGDWGAGFDYWGQGTLVTVSS SEQ ID NO: 169 is the amino acid sequence of the mAb10 VL. DIQMTQSPSFLSASVGDRVTIACRASQSISSWLAWYQQKPGKAPKLLIYHASSLESGVPSRFSGSASGTEFALTISSLOP DDFATYYCQQYSSYWTFGQGTKVEIK DETAILED DESCRIPTION Malaria is a mosquito-borne parasitic disease causing high morbidity and mortality, primarily in infants and young children in sub-Saharan Africa. Development of a highly effective vaccine or antibodies that can prevent and ultimately eliminate malaria is urgently needed. This disclosure provides monoclonal antibodies and antigen binding fragments directed against PfCSP. Data in the examples show that passive transfer of the disclosed antibodies reduces parasite liver burden in an animal model, and that the disclosed antibodies target a different site than prior antibodies against PfCSP and enhance protection when passively transferred in combination with prior antibodies against PfCSP. Thus, the PfCSP-specific antibodies and antigen binding fragments provided herein are effective for passive prevention of malaria for use in suitable subjects, such as travelers, military personnel, and subjects in elimination campaigns. I. Summary of Terms Unless otherwise noted, technical terms are used according to conventional usage. Definitions of many common terms in molecular biology may be found in Krebs et al. (eds.), Lewin’s genes XII, published by Jones & Bartlett Learning, 2017. As used herein, the singular forms “a,” “an,” and “the,” refer to both the singular as well as plural, unless the context clearly indicates otherwise. For example, the term “an antigen” includes singular or plural antigens and can be considered equivalent to the phrase “at least one antigen.” As used herein, the term “comprises” means “includes.” It is further to be understood that any
and all base sizes or amino acid sizes, and all molecular weight or molecular mass values, given for nucleic acids or polypeptides are approximate, and are provided for descriptive purposes, unless otherwise indicated. Although many methods and materials similar or equivalent to those described herein can be used, particular suitable methods and materials are described herein. In case of conflict, the present specification, including explanations of terms, will control. In addition, the materials, methods, and examples are illustrative only and not intended to be limiting. To facilitate review of the various aspects, the following explanations of terms are provided: 317 Antibody: A monoclonal antibody that specifically binds to an epitope on PfCSP and neutralizes malaria infection. The CIS317 antibody and methods for its production are described, for example, in Oyen et al. (“Structural basis for antibody recognition of the NANP repeats in Plasmodium falciparum circumsporozoite protein,” Proc Natl Acad Sci USA, 114, E10438-E10445, 2017). The amino acid sequences of the heavy and light variable regions of the CIS317 antibody are provided herein as SEQ ID NOs: 159 and 160. About: Unless context indicated otherwise, “about” refers to plus or minus 5% of a reference value. For example, “about” 100 refers to 95 to 105. Administration: The introduction of a composition into a subject by a chosen route. Administration can be local or systemic. For example, if the chosen route is intravenous, the composition is administered by introducing the composition into a vein of the subject. Exemplary routes of administration include, but are not limited to, oral, injection (such as subcutaneous, intramuscular, intradermal, intraperitoneal, and intravenous), sublingual, rectal, transdermal (for example, topical), intranasal, vaginal, and inhalation routes. Antibody and Antigen Binding Fragment: An immunoglobulin, antigen-binding fragment, or derivative thereof, that specifically binds and recognizes an analyte (antigen) such as PfCSP. The term “antibody” is used herein in the broadest sense and encompasses various antibody structures, including but not limited to monoclonal antibodies, polyclonal antibodies, multispecific antibodies (e.g., bispecific antibodies), and antigen binding fragments, so long as they exhibit the desired antigen-binding activity. Non-limiting examples of antibodies include, for example, intact immunoglobulins and variants and fragments thereof that retain binding affinity for the antigen. Examples of antigen binding fragments include but are not limited to Fv, Fab, Fab', Fab'-SH, F(ab')2; diabodies; linear antibodies; single-chain antibody molecules (e.g. scFv); and multispecific antibodies formed from antibody fragments. Antibody fragments include antigen binding fragments either produced by the modification of whole antibodies or those synthesized de novo using recombinant DNA methodologies (see, e.g., Kontermann and Dübel (Eds.), Antibody Engineering, Vols.1-2, 2nd ed., Springer-Verlag, 2010). Antibodies also include genetically engineered forms such as chimeric antibodies (such as humanized murine antibodies) and heteroconjugate antibodies (such as bispecific antibodies). An antibody may have one or more binding sites. If there is more than one binding site, the binding sites may be identical to one another or may be different. For instance, a naturally-occurring
immunoglobulin has two identical binding sites, a single-chain antibody or Fab fragment has one binding site, while a bispecific or bifunctional antibody has two different binding sites. Typically, a naturally occurring immunoglobulin has heavy (H) chains and light (L) chains interconnected by disulfide bonds. Immunoglobulin genes include the kappa, lambda, alpha, gamma, delta, epsilon and mu constant region genes, as well as the myriad immunoglobulin variable domain genes. There are two types of light chain, lambda (λ) and kappa (κ). There are five main heavy chain classes (or isotypes) which determine the functional activity of an antibody molecule: IgM, IgD, IgG, IgA and IgE. Each heavy and light chain contains a constant region (or constant domain) and a variable region (or variable domain). In combination, the heavy and the light chain variable regions specifically bind the antigen. References to “VH” or “VH” refer to the variable region of an antibody heavy chain, including that of an antigen binding fragment, such as Fv, scFv, dsFv or Fab. References to “VL” or “VL” refer to the variable domain of an antibody light chain, including that of an Fv, scFv, dsFv or Fab. The VH and VL contain a “framework” region interrupted by three hypervariable regions, also called “complementarity-determining regions” or “CDRs” (see, e.g., Kabat et al., Sequences of Proteins of Immunological Interest, 5th ed., NIH Publication No.91-3242, Public Health Service, National Institutes of Health, U.S. Department of Health and Human Services, 1991). The sequences of the framework regions of different light or heavy chains are relatively conserved within a species. The framework region of an antibody, that is the combined framework regions of the constituent light and heavy chains, serves to position and align the CDRs in three-dimensional space. The CDRs are primarily responsible for binding to an epitope of an antigen. The amino acid sequence boundaries of a given CDR can be readily determined using any of a number of well-known schemes, including those described by Kabat et al. (Sequences of Proteins of Immunological Interest, 5th ed., NIH Publication No.91-3242, Public Health Service, National Institutes of Health, U.S. Department of Health and Human Services, 1991; “Kabat” numbering scheme), Al-Lazikani et al., (“Standard conformations for the canonical structures of immunoglobulins,” J. Mol. Bio., 273(4):927-948, 1997; “Chothia” numbering scheme), and Lefranc et al. (“IMGT unique numbering for immunoglobulin and T cell receptor variable domains and Ig superfamily V-like domains,” Dev. Comp. Immunol., 27(1):55-77, 2003; “IMGT” numbering scheme). The CDRs of each chain are typically referred to as CDR1, CDR2, and CDR3 (from the N-terminus to C-terminus), and are also typically identified by the chain in which the particular CDR is located. Thus, a VH CDR3 is the CDR3 from the VH of the antibody in which it is found, whereas a VL CDR1 is the CDR1 from the VL of the antibody in which it is found. Light chain CDRs are sometimes referred to as LCDR1, LCDR2, and LCDR3. Heavy chain CDRs are sometimes referred to as HCDR1, HCDR2, and HCDR3. In some aspects, a disclosed antibody includes a heterologous constant domain. For example, the antibody includes a constant domain that is different from a native constant domain, such as a constant domain including one or more modifications (such as the “LS” mutations) to increase half-life.
A “monoclonal antibody” is an antibody obtained from a population of substantially homogeneous antibodies, that is, the individual antibodies comprising the population are identical and/or bind the same epitope, except for possible variant antibodies, for example, containing naturally occurring mutations or arising during production of a monoclonal antibody preparation, such variants generally being present in minor amounts. In contrast to polyclonal antibody preparations, which typically include different antibodies directed against different determinants (epitopes), each monoclonal antibody of a monoclonal antibody preparation is directed against a single determinant on an antigen. Thus, the modifier “monoclonal” indicates the character of the antibody as being obtained from a substantially homogeneous population of antibodies, and is not to be construed as requiring production of the antibody by any particular method. For example, the monoclonal antibodies may be made by a variety of techniques, including but not limited to the hybridoma method, recombinant DNA methods, phage-display methods, and methods utilizing transgenic animals containing all or part of the human immunoglobulin loci, such methods and other exemplary methods for making monoclonal antibodies being described herein. In some examples monoclonal antibodies are isolated from a subject. Monoclonal antibodies can have conservative amino acid substitutions which have substantially no effect on antigen binding or other immunoglobulin functions. (See, for example, Greenfield (Ed.), Antibodies: A Laboratory Manual, 2nd ed. New York: Cold Spring Harbor Laboratory Press, 2014.) A “humanized” antibody or antigen binding fragment includes a human framework region and one or more CDRs from a non-human (such as a mouse, rat, or synthetic) antibody or antigen binding fragment. The non-human antibody or antigen binding fragment providing the CDRs is termed a “donor,” and the human antibody or antigen binding fragment providing the framework is termed an “acceptor.” In one aspect, all the CDRs are from the donor immunoglobulin in a humanized immunoglobulin. Constant regions need not be present, but if they are, they can be substantially identical to human immunoglobulin constant regions, such as at least about 85-90%, such as about 95% or more identical. Hence, all parts of a humanized antibody or antigen binding fragment, except possibly the CDRs, are substantially identical to corresponding parts of natural human antibody sequences. A “chimeric antibody” is an antibody which includes sequences derived from two different antibodies, which typically are of different species. In some examples, a chimeric antibody includes one or more CDRs and/or framework regions from one human antibody and CDRs and/or framework regions from another human antibody. A “fully human antibody” or “human antibody” is an antibody which includes sequences from (or derived from) the human genome, and does not include sequence from another species. In some aspects, a human antibody includes CDRs, framework regions, and (if present) an Fc region from (or derived from) the human genome. Human antibodies can be identified and isolated using technologies for creating antibodies based on sequences derived from the human genome, for example by phage display or using transgenic animals (see, e.g., Barbas et al. Phage display: A Laboratory Manuel. 1st Ed. New York: Cold Spring
Harbor Laboratory Press, 2004. Print.; Lonberg, Nat. Biotech., 23: 1117-1125, 2005; Lonenberg, Curr. Opin. Immunol., 20:450-459, 2008). Antibody or antigen binding fragment that neutralizes P. falciparum: An antibody or antigen binding fragment that specifically binds to a P. falciparum antigen (such as PfCSP) in such a way as to inhibit a biological function associated with P. falciparum that inhibits P. falciparum infection. The antibody can neutralize the activity of P. falciparum at various points during the lifecycle of the pathogen. For example, an antibody or antigen binding fragment that neutralizes P. falciparum may interfere with the pathogen by binding it in the skin and limiting entry into the blood or entry into the hepatocytes in the liver by interfering with the interaction of the pathogen and one or more cell surface receptors. Alternately, an antibody may interfere with one or more post-attachment interactions of the pathogen with its receptors, for example, by interfering with pathogen internalization by receptor-mediated endocytosis. In some aspects, an antibody or antigen binding fragment that specifically binds to PfCSP and neutralizes P. falciparum inhibits sporozoite invasion of hepatocytes, for example, by at least 50% (such as at least 60%, at least 70%, at least 80%, at least 90%, or more) compared to a control antibody or antigen binding fragment. In some aspects, an antibody or antigen binding fragment that specifically binds to PfCSP and neutralizes P. falciparum inhibits infection of a human subject by P. falciparum, for example, by at least 50% compared to a control antibody or antigen binding fragment. Biological sample: A sample obtained from a subject. Biological samples include all clinical samples useful for detection of disease or infection (for example, P. falciparum infection) in subjects, including, but not limited to, cells, tissues, and bodily fluids, such as blood, derivatives and fractions of blood (such as serum), cerebrospinal fluid; as well as biopsied or surgically removed tissue, for example tissues that are unfixed, frozen, or fixed in formalin or paraffin. In a particular example, a biological sample is obtained from a subject having or suspected of having a P. falciparum infection. Bispecific antibody: A recombinant molecule composed of two different antigen binding domains that consequently binds to two different antigenic epitopes. Bispecific antibodies include chemically or genetically linked molecules of two antigen-binding domains. The antigen binding domains can be linked using a linker. The antigen binding domains can be monoclonal antibodies, antigen-binding fragments (e.g., Fab, scFv), or combinations thereof. A bispecific antibody can include one or more constant domains, but does not necessarily include a constant domain. Circumsporozoite protein (CSP): The circumsporozoite protein (CSP) is a major malaria parasite surface protein during the sporogonic cycle. PfCSP covers the surface of P. falciparum sporozoites, which are transmitted from the mosquito salivary gland to host hepatocytes. An exemplary PfCSP amino acid sequence is provided as SEQ ID NO: 72. CIS43 Antibody: A monoclonal antibody that specifically binds to an epitope on PfCSP and neutralizes malaria infection. The CIS43 antibody and methods for its production are described, for example, in PCT Pub. No. WO 2018/148660. The amino acid sequences of the heavy and light variable regions of the CIS43 antibody are provided herein as SEQ ID NOs: 68 and 69.
Conditions sufficient to form an immune complex: Conditions which allow an antibody or antigen binding fragment to bind to its cognate epitope to a detectably greater degree than, and/or to the substantial exclusion of, binding to substantially all other epitopes. Conditions sufficient to form an immune complex are dependent upon the format of the binding reaction and typically are those utilized in immunoassay protocols or those conditions encountered in vivo. See Greenfield (Ed.), Antibodies: A Laboratory Manual, 2nd ed. New York: Cold Spring Harbor Laboratory Press, 2014, for a description of immunoassay formats and conditions. The conditions employed in the methods are “physiological conditions” which include reference to conditions (e.g., temperature, osmolarity, pH) that are typical inside a living mammal or a mammalian cell. While it is recognized that some organs are subject to extreme conditions, the intra-organismal and intracellular environment normally lies around pH 7 (e.g., from pH 6.0 to pH 8.0, more typically pH 6.5 to 7.5), contains water as the predominant solvent, and exists at a temperature above 0°C and below 50°C. Osmolarity is within the range that is supportive of cell viability and proliferation. The formation of an immune complex can be detected through conventional methods, for instance immunohistochemistry (IHC), immunoprecipitation (IP), flow cytometry, immunofluorescence microscopy, ELISA, immunoblotting (for example, Western blot), magnetic resonance imaging (MRI), computed tomography (CT) scans, radiography, and affinity chromatography. Conjugate: A complex of two molecules linked together, for example, linked together by a covalent bond. In one aspect, an antibody is linked to an effector molecule; for example, an antibody that specifically binds to CSP from P. falciparum covalently linked to an effector molecule. The linkage can be by chemical or recombinant means. In one aspect, the linkage is chemical, wherein a reaction between the antibody moiety and the effector molecule has produced a covalent bond formed between the two molecules to form one molecule. A peptide linker (short peptide sequence) can optionally be included between the antibody and the effector molecule. Because conjugates can be prepared from two molecules with separate functionalities, such as an antibody and an effector molecule, they are also sometimes referred to as “chimeric molecules.” Conservative variants: “Conservative” amino acid substitutions are those substitutions that do not substantially affect or decrease a function of a protein, such as the ability of the protein to interact with a target protein. For example, a PfCSP-specific antibody can include up to 1, 2, 3, 4, 5, 6, 7, 8, 9, or up to 10 conservative substitutions compared to a reference antibody sequence and retain specific binding activity for PfCSP, and/or P. falciparum neutralization activity. The term conservative variation also includes the use of a substituted amino acid in place of an unsubstituted parent amino acid. Individual substitutions, deletions or additions which alter, add or delete a single amino acid or a small percentage of amino acids (for instance less than 5%, in some aspects less than 1%) in an encoded sequence are conservative variations where the alterations result in the substitution of an amino acid with a chemically similar amino acid.
The following six groups are examples of amino acids that are considered to be conservative substitutions for one another: 1) Alanine (A), Serine (S), Threonine (T); 2) Aspartic acid (D), Glutamic acid (E); 3) Asparagine (N), Glutamine (Q); 4) Arginine (R), Lysine (K); 5) Isoleucine (I), Leucine (L), Methionine (M), Valine (V); and 6) Phenylalanine (F), Tyrosine (Y), Tryptophan (W). Non-conservative substitutions are those that reduce an activity or function of the PfCSP specific antibody, such as the ability to specifically bind to PfCSP or neutralize P. falciparum. For instance, if an amino acid residue is essential for a function of the protein, even an otherwise conservative substitution may disrupt that activity. Thus, a conservative substitution does not alter the basic function of a protein of interest. Contacting: Placement in direct physical association; includes both in solid and liquid form, which can take place either in vivo or in vitro. Contacting includes contact between one molecule and another molecule, for example the amino acid on the surface of one polypeptide, such as an antigen, that contacts another polypeptide, such as an antibody. Contacting can also include contacting a cell for example by placing an antibody in direct physical association with a cell. Control: A reference standard. In some aspects, the control is a negative control, such as sample obtained from a healthy patient not infected with P. falciparum. In other aspects, the control is a positive control, such as a tissue sample obtained from a patient diagnosed with P. falciparum infection. In still other aspects, the control is a historical control or standard reference value or range of values (such as a previously tested control sample, such as a group of P. falciparum patients with known prognosis or outcome, or group of samples that represent baseline or normal values). A difference between a test sample and a control can be an increase or conversely a decrease. The difference can be a qualitative difference or a quantitative difference, for example a statistically significant difference. In some examples, a difference is an increase or decrease, relative to a control, of at least about 5%, such as at least about 10%, at least about 20%, at least about 30%, at least about 40%, at least about 50%, at least about 60%, at least about 70%, at least about 80%, at least about 90%, at least about 100%, at least about 150%, at least about 200%, at least about 250%, at least about 300%, at least about 350%, at least about 400%, or at least about 500%. Detectable marker: A detectable molecule (also known as a label) that is conjugated directly or indirectly to a second molecule, such as an antibody, to facilitate detection of the second molecule. For example, the detectable marker can be capable of detection by ELISA, spectrophotometry, flow cytometry, microscopy or diagnostic imaging techniques (such as CT scans, MRIs, ultrasound, fiberoptic examination, and laparoscopic examination). Specific, non-limiting examples of detectable markers include fluorophores, chemiluminescent agents, enzymatic linkages, radioactive isotopes and heavy metals or compounds (for
example super paramagnetic iron oxide nanocrystals for detection by MRI). Methods for using detectable markers and guidance in the choice of detectable markers appropriate for various purposes are discussed for example in Green and Sambrook (Molecular Cloning: A Laboratory Manual, 4th ed., New York: Cold Spring Harbor Laboratory Press, 2012) and Ausubel et al. (Eds.) (Current Protocols in Molecular Biology, New York: John Wiley and Sons, including supplements, 2017). Detecting: To identify the existence, presence, or fact of something. Effective amount: A quantity of a specific substance sufficient to achieve a desired effect in a subject to whom the substance is administered. For instance, this can be the amount necessary to inhibit a P. falciparum infection, such as the amount necessary to inhibit or prevent P. falciparum sporozoites from invading the liver in the subject or to measurably alter outward symptoms of the P. falciparum infection. In some aspects, administration of an effective amount of a disclosed antibody or antigen binding fragment that binds to PfCSP can reduce or inhibit a P. falciparum infection (for example, as measured by infection of cells, or by number or percentage of subjects infected by the P. falciparum, or by an increase in the survival time of infected subjects, or reduction in symptoms associated with P. falciparum infection) by a desired amount, for example by at least 10%, at least 20%, at least 50%, at least 60%, at least 70%, at least 80%, at least 90%, at least 95%, at least 98%, or even at least 100% (elimination or prevention of detectable P. falciparum infection), as compared to a suitable control. The effective amount of an antibody or antigen binding fragment that specifically binds PfCSP that is administered to a subject to inhibit P. falciparum infection will vary depending upon a number of factors associated with that subject, for example the overall health and/or weight of the subject. An effective amount can be determined by varying the dosage and measuring the resulting response, such as, for example, a reduction in pathogen titer. Effective amounts also can be determined through various in vitro, in vivo or in situ immunoassays. An effective amount encompasses a fractional dose that contributes in combination with previous or subsequent administrations to attaining an effective response. For example, an effective amount of an agent can be administered in a single dose, or in several doses, for example daily, during a course of treatment lasting several days or weeks. However, the effective amount can depend on the subject being treated, the severity and type of the condition being treated, and the manner of administration. A unit dosage form of the agent can be packaged in an amount, or in multiples of the effective amount, for example, in a vial (e.g., with a pierceable lid) or syringe having sterile components. Effector molecule: A molecule intended to have or produce a desired effect; for example, a desired effect on a cell to which the effector molecule is targeted. Effector molecules can include, for example, polypeptides and small molecules. In one non-limiting example, the effector molecule is a toxin. Some effector molecules may have or produce more than one desired effect. Epitope: An antigenic determinant. These are particular chemical groups or peptide sequences on a molecule that are antigenic, i.e. that elicit a specific immune response. An antibody specifically binds a
particular antigenic epitope on a polypeptide. In some examples a disclosed antibody specifically binds to an epitope on CSP from P. falciparum. Expression: Transcription or translation of a nucleic acid sequence. For example, an encoding nucleic acid sequence (such as a gene) can be expressed when its DNA is transcribed into RNA or an RNA fragment, which in some examples is processed to become mRNA. An encoding nucleic acid sequence (such as a gene) may also be expressed when its mRNA is translated into an amino acid sequence, such as a protein or a protein fragment. In a particular example, a heterologous gene is expressed when it is transcribed into an RNA. In another example, a heterologous gene is expressed when its RNA is translated into an amino acid sequence. Regulation of expression can include controls on transcription, translation, RNA transport and processing, degradation of intermediary molecules such as mRNA, or through activation, inactivation, compartmentalization or degradation of specific protein molecules after they are produced. Expression Control Sequences: Nucleic acid sequences that regulate the expression of a heterologous nucleic acid sequence to which it is operatively linked. Expression control sequences are operatively linked to a nucleic acid sequence when the expression control sequences control and regulate the transcription and, as appropriate, translation of the nucleic acid sequence. Thus, expression control sequences can include appropriate promoters, enhancers, transcriptional terminators, a start codon (ATG) in front of a protein-encoding gene, splice signals for introns, maintenance of the correct reading frame of that gene to permit proper translation of mRNA, and stop codons. The term “control sequences” is intended to include, at a minimum, components whose presence can influence expression, and can also include additional components whose presence is advantageous, for example, leader sequences and fusion partner sequences. Expression control sequences can include a promoter. Expression vector: A vector comprising a recombinant polynucleotide comprising expression control sequences operatively linked to a nucleotide sequence to be expressed. An expression vector comprises sufficient cis- acting elements for expression; other elements for expression can be supplied by the host cell or in an in vitro expression system. Non-limiting examples of expression vectors include cosmids, plasmids (e.g., naked or contained in liposomes) and viruses (e.g., lentiviruses, retroviruses, adenoviruses, and adeno-associated viruses) that incorporate the recombinant polynucleotide. A polynucleotide can be inserted into an expression vector that contains a promoter sequence which facilitates the efficient transcription of the inserted genetic sequence of the host. The expression vector typically contains an origin of replication, a promoter, as well as specific nucleic acid sequences that allow phenotypic selection of the transformed cells. Fc region: The constant region of an antibody excluding the first heavy chain constant domain. Fc region generally refers to the last two heavy chain constant domains of IgA, IgD, and IgG, and the last three heavy chain constant domains of IgE and IgM. An Fc region may also include part or all of the flexible hinge N-terminal to these domains. For IgA and IgM, an Fc region may or may not include the tailpiece, and may or may not be bound by the J chain. For IgG, the Fc region is typically understood to include immunoglobulin domains Cγ2 and Cγ3 and optionally the lower part of the hinge between Cγ1 and Cγ2.
Although the boundaries of the Fc region may vary, the human IgG heavy chain Fc region is usually defined to include residues following C226 or P230 to the Fc carboxyl-terminus, wherein the numbering is according to Kabat. For IgA, the Fc region includes immunoglobulin domains Cα2 and Cα3 and optionally the lower part of the hinge between Cα1 and Cα2. Host cell: Cells in which a vector can be propagated and its DNA expressed. The cell may be prokaryotic or eukaryotic. The term also includes any progeny of the subject host cell. It is understood that all progeny may not be identical to the parental cell since there may be mutations that occur during replication. However, such progeny are included when the term “host cell” is used. IgA: A polypeptide belonging to the class of antibodies that are substantially encoded by a recognized immunoglobulin alpha gene. In humans, this class or isotype comprises IgA1 and IgA2. IgA antibodies can exist as monomers, polymers (referred to as pIgA) of predominantly dimeric form, and secretory IgA. The constant chain of wild-type IgA contains an 18-amino-acid extension at its C-terminus called the tail piece (tp). Polymeric IgA is secreted by plasma cells with a 15-kDa peptide called the J chain linking two monomers of IgA through the conserved cysteine residue in the tail piece. IgG: A polypeptide belonging to the class or isotype of antibodies that are substantially encoded by a recognized immunoglobulin gamma gene. In humans, this class comprises IgG1, IgG2, IgG3, and IgG4. Immune complex: The binding of antibody or antigen binding fragment (such as a scFv) to a soluble antigen forms an immune complex. The formation of an immune complex can be detected through conventional methods, for instance immunohistochemistry, immunoprecipitation, flow cytometry, immunofluorescence microscopy, ELISA, immunoblotting (for example, Western blot), magnetic resonance imaging, CT scans, radiography, and affinity chromatography. Inhibiting a disease or condition: Reducing the full development of a disease or condition in a subject, for example, reducing the full development of a P. falciparum infection in a subject who is at risk of a P. falciparum infection. This includes neutralizing, antagonizing, prohibiting, preventing, restraining, slowing, disrupting, stopping, or reversing progression or severity of the disease or condition. In some aspects, inhibiting a disease or condition refers to a prophylactic intervention administered before the disease or condition has begun to develop (for example a treatment initiated in a subject at risk of P. falciparum infection, but not infected by P. falciparum) that reduces subsequent development of the disease or condition and/or ameliorates a sign or symptom of the disease or condition following development. The term “ameliorating,” with reference to inhibiting a disease or condition refers to any observable beneficial effect of the prophylactic intervention intended to inhibit the disease or condition. The beneficial effect can be evidenced, for example, by a delayed onset of clinical symptoms of the disease or condition in a susceptible subject, a reduction in severity of some or all clinical symptoms of the disease or condition, a slower progression of the disease or condition, an improvement in the overall health or well- being of the subject, a reduction in infection, or by other parameters that are specific to the particular disease or condition.
In some aspects, the disclosed PfCSP-specific antibodies and antigen binding fragments inhibit the invasion of Plasmodium falciparum sporozoites into human liver cells (hepatocytes). The invasion of liver cells is a key event in the infection of a subject with the malaria parasite. Inhibition of the invasion of human liver cells can be measured by one or more of several standard assays (see, for example, Example 1). For example, the disclosed PfCSP-specific antibodies and antigen binding fragments can inhibit the invasion of Plasmodium falciparum sporozoites into human liver cells by at least 25%, such as at least 50%, at least 75%, at least 90%, at least 95%, or 100% compared to a suitable control. In some aspects, the disclosed PfCSP-specific antibodies and antigen binding fragments inhibit the growth of Plasmodium falciparum in a subject, for example, the antibodies and antigen binding fragments inhibit the multiplication of Plasmodium falciparum in the subject, resulting in a reduction in pathogen load in the subject compared to a relevant control. For example, the disclosed PfCSP-specific antibodies and antigen binding fragments can inhibit the growth of Plasmodium falciparum in a subject by at least 25%, such as at least 50%, at least 75%, at least 90%, at least 95%, or 100% compared to a suitable control. Isolated: A biological component (such as a nucleic acid, peptide, protein or protein complex, for example an antibody) that has been substantially separated, produced apart from, or purified away from other biological components in the cell of the organism in which the component naturally occurs, that is, other chromosomal and extra-chromosomal DNA and RNA, and proteins. Thus, isolated nucleic acids, peptides and proteins include nucleic acids and proteins purified by standard purification methods. The term also embraces nucleic acids, peptides and proteins prepared by recombinant expression in a host cell, as well as, chemically synthesized nucleic acids. An isolated nucleic acid, peptide or protein, for example an antibody, can be at least 50%, at least 60%, at least 70%, at least 80%, at least 90%, at least 95%, at least 96%, at least 97%, at least 98%, or at least 99% pure. Kabat position: A position of a residue in an amino acid sequence that follows the numbering convention delineated by Kabat et al. (Sequences of Proteins of Immunological Interest, 5th Edition, Department of Health and Human Services, Public Health Service, National Institutes of Health, Bethesda, NIH Publication No.91-3242, 1991). L9 Antibody: A monoclonal antibody that specifically binds to an epitope on PfCSP and neutralizes malaria infection. The L9 antibody and methods for its production are described, for example, in PCT Pub. No. WO 2020/227228. The amino acid sequences of the heavy and light variable regions of the L9 antibody are provided herein as SEQ ID NOs: 70 and 71. Linker: A bi-functional molecule that can be used to link two molecules into one contiguous molecule, for example, to link an effector molecule to an antibody. Non-limiting examples of peptide linkers include glycine-serine linkers. The terms “conjugating,” “joining,” “bonding,” or “linking” can refer to making two molecules into one contiguous molecule; for example, linking two polypeptides into one contiguous polypeptide, or covalently attaching an effector molecule or detectable marker radionuclide or other molecule to a polypeptide, such as an scFv. The linkage can be either by chemical or recombinant means. “Chemical
means” refers to a reaction between the antibody moiety and the effector molecule such that there is a covalent bond formed between the two molecules to form one molecule. mAb 5D5 Antibody: A control monoclonal antibody that specifically binds to an epitope on PfCSP that is used as a control herein. This antibody detects only the un-cleaved form of PfCSP. The mAb 5D5 antibody is described, for example, on the internet rcsb.org/structure/6UUD, as available on September 18, 2023. The amino acid sequences of the heavy and light variable regions of the mAb 5D5 are provided as SEQ ID NOs: 166 and 167. mAb10 Antibody: A control monoclonal antibody that specifically binds to an epitope on PfCSP. This antibody detects both the cleaved (lower molecular weight) and un-cleaved (higher molecular weight) forms of PfCSP. The mAb10 antibody is described, for example, in Kisalu et al., Nat. Med.24, 408-416. 10.1038/nm.4512, 2018 and PCT Publication No. WO 2018/148660. The amino acid sequences of the heavy and light variable regions of the mAb10 are provided as SEQ ID NOs: 168 and 169. Malaria: Malaria is a parasitic infection of humans by the Plasmodium species P. falciparum, P. vivax, P. ovale, P. malariae, and P. knowlesi. Humans become infected following the bite of an infected mosquito, the host of the malarial parasite. Malaria rarely occurs in humans following a blood transfusion or subsequent to needle-sharing. Clinical manifestations of malarial infection which may occur include blackwater fever, cerebral malaria, respiratory failure, hepatic necrosis, occlusion of myocardial capillaries and death. Infection begins when malaria sporozoites gain access to or are directly injected into the bloodstream of a host by a mosquito. After injection, they migrate to the liver and multiply in hepatocytes for one week. The sporozoites substantially expand in the liver and differentiate to merozoites which are released from the liver into the blood stream, where they infect erythrocytes. When the merozoite matures in the red blood cell, it is known as a trophozoite and, when fully developed, as a schizont. A schizont is the stage when nuclear division occurs to form individual merozoites which are released to invade other red cells. Malaria clinical symptoms appear during the blood-stage. After several schizogonic cycles, some parasites, instead of becoming schizonts through asexual reproduction, develop into large uninucleate parasites, known as gametocytes. These gametocytes are the sexual blood cell stage forms of the parasite. Sexual development of the malaria parasites involves the female macrogametocyte and the male microgametocyte. If a mosquito feeds on the blood of an infected host, it can ingest gametocytes within the blood. Fertilization and sexual recombination of the parasite occurs in the mosquito's gut. The fertilized parasite, which is known as a zygote, then develops into an ookinete. The ookinete penetrates the midgut wall of the mosquito and develops into an oocyst, within which many small sporozoites form. When the oocyst ruptures, the sporozoites migrate to the salivary gland of the mosquito via the hemolymph. Once in the saliva of the mosquito, the parasite can be injected into a host, repeating the life cycle. Nucleic acid (molecule or sequence): A deoxyribonucleotide or ribonucleotide polymer or combination thereof including without limitation, cDNA, mRNA, genomic DNA, and synthetic (such as chemically synthesized) DNA or RNA. The nucleic acid can be double stranded (ds) or single stranded (ss).
Where single stranded, the nucleic acid can be the sense strand or the antisense strand. Nucleic acids can include natural nucleotides (such as A, T/U, C, and G), and can include analogs of natural nucleotides, such as labeled nucleotides. “cDNA” refers to a DNA that is complementary or identical to an mRNA, in either single stranded or double stranded form. “Encoding” refers to the inherent property of specific sequences of nucleotides in a polynucleotide, such as a gene, a cDNA, or an mRNA, to serve as templates for synthesis of other polymers and macromolecules in biological processes having either a defined sequence of nucleotides (i.e., rRNA, tRNA and mRNA) or a defined sequence of amino acids and the biological properties resulting therefrom. Thus, a gene encodes a protein if transcription and translation of mRNA produced by that gene produces the protein in a cell or other biological system. Both the coding strand, the nucleotide sequence of which is identical to the mRNA sequence and is usually provided in sequence listings, and non-coding strand, used as the template for transcription, of a gene or cDNA can be referred to as encoding the protein or other product of that gene or cDNA. Unless otherwise specified, a “nucleotide sequence encoding an amino acid sequence” includes all nucleotide sequences that are degenerate versions of each other and that encode the same amino acid sequence. Nucleotide sequences that encode proteins and RNA may include introns. Operably linked: A first nucleic acid sequence is operably linked with a second nucleic acid sequence when the first nucleic acid sequence is placed in a functional relationship with the second nucleic acid sequence. For instance, a promoter, such as the CMV promoter, is operably linked to a coding sequence if the promoter affects the transcription or expression of the coding sequence. Generally, operably linked DNA sequences are contiguous and, where necessary to join two protein-coding regions, in the same reading frame. Pharmaceutically acceptable carriers: The pharmaceutically acceptable carriers of use are conventional. Remington: The Science and Practice of Pharmacy, 22nd ed., London, UK: Pharmaceutical Press, 2013, describes compositions and formulations suitable for pharmaceutical delivery of the disclosed agents. In general, the nature of the carrier will depend on the particular mode of administration being employed. For instance, parenteral formulations usually include injectable fluids that include pharmaceutically and physiologically acceptable fluids such as water, physiological saline, balanced salt solutions, aqueous dextrose, glycerol or the like as a vehicle. For solid compositions (e.g., powder, pill, tablet, or capsule forms), conventional non-toxic solid carriers can include, for example, pharmaceutical grades of mannitol, lactose, starch, or magnesium stearate. In addition to biologically neutral carriers, pharmaceutical compositions to be administered can contain minor amounts of non-toxic auxiliary substances, such as wetting or emulsifying agents, added preservatives (such as non-natural preservatives), and pH buffering agents and the like, for example sodium acetate or sorbitan monolaurate. In particular examples, the pharmaceutically acceptable carrier is sterile and suitable for parenteral administration to a subject for example, by injection. In some aspects, the active agent and pharmaceutically acceptable carrier
are provided in a unit dosage form such as a pill or in a selected quantity in a vial. Unit dosage forms can include one dosage or multiple dosages (for example, in a vial from which metered dosages of the agents can selectively be dispensed). Polypeptide: A polymer in which the monomers are amino acid residues that are joined together through amide bonds. When the amino acids are alpha-amino acids, either the L-optical isomer or the D- optical isomer can be used, the L-isomers being preferred. The terms “polypeptide” or “protein” as used herein are intended to encompass any amino acid sequence and include modified sequences such as glycoproteins. A polypeptide includes both naturally occurring proteins, as well as those that are recombinantly or synthetically produced. A polypeptide has an amino terminal (N-terminal) end and a carboxy-terminal end. In some aspects, the polypeptide is a disclosed antibody or a fragment thereof. Purified: The term purified does not require absolute purity; rather, it is intended as a relative term. Thus, for example, a purified peptide preparation is one in which the peptide or protein (such as an antibody) is more enriched than the peptide or protein is in its natural environment within a cell. In one aspect, a preparation is purified such that the protein or peptide represents at least 50% of the total peptide or protein content of the preparation. Recombinant: A recombinant nucleic acid is one that has a sequence that is not naturally occurring or has a sequence that is made by an artificial combination of two otherwise separated segments of sequence. This artificial combination can be accomplished by chemical synthesis or, more commonly, by the artificial manipulation of isolated segments of nucleic acids, for example, by genetic engineering techniques. A recombinant protein is one that has a sequence that is not naturally occurring or has a sequence that is made by an artificial combination of two otherwise separated segments of sequence. In several aspects, a recombinant protein is encoded by a heterologous (for example, recombinant) nucleic acid that has been introduced into a host cell, such as a bacterial or eukaryotic cell. The nucleic acid can be introduced, for example, on an expression vector having signals capable of expressing the protein encoded by the introduced nucleic acid or the nucleic acid can be integrated into the host cell chromosome. Sequence identity: The identity between two or more nucleic acid sequences, or two or more amino acid sequences, is expressed in terms of the identity between the sequences. Sequence identity can be measured in terms of percentage identity; the higher the percentage, the more identical the sequences. Homologs and variants of a VL or a VH of an antibody that specifically binds a target antigen are typically characterized by possession of at least about 75% sequence identity, for example at least about 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% sequence identity counted over the full-length alignment with the amino acid sequence of interest. Any suitable method may be used to align sequences for comparison. Non-limiting examples of programs and alignment algorithms are described in: Smith and Waterman, Adv. Appl. Math.2(4):482-489, 1981; Needleman and Wunsch, J. Mol. Biol.48(3):443-453, 1970; Pearson and Lipman, Proc. Natl. Acad. Sci. U.S.A.85(8):2444-2448, 1988; Higgins and Sharp, Gene, 73(1):237-244, 1988; Higgins and Sharp, Bioinformatics, 5(2):151-3, 1989; Corpet, Nucleic Acids Res.16(22):10881-10890, 1988; Huang et al.
Bioinformatics, 8(2):155-165, 1992; and Pearson, Methods Mol. Biol.24:307-331, 1994. Altschul et al., J. Mol. Biol.215(3):403-410, 1990, presents a detailed consideration of sequence alignment methods and homology calculations. The NCBI Basic Local Alignment Search Tool (BLAST) (Altschul et al., J. Mol. Biol.215(3):403-410, 1990) is available from several sources, including the National Center for Biological Information and on the Internet, for use in connection with the sequence analysis programs blastp, blastn, blastx, tblastn, and tblastx. Blastn is used to compare nucleic acid sequences, while blastp is used to compare amino acid sequences. Additional information can be found at the NCBI web site. Generally, once two sequences are aligned, the number of matches is determined by counting the number of positions where an identical nucleotide or amino acid residue is present in both sequences. The percent sequence identity between the two sequences is determined by dividing the number of matches either by the length of the sequence set forth in the identified sequence, or by an articulated length (such as 100 consecutive nucleotides or amino acid residues from a sequence set forth in an identified sequence), followed by multiplying the resulting value by 100. Specifically bind: When referring to an antibody or antigen binding fragment, refers to a binding reaction which determines the presence of a target protein in the presence of a heterogeneous population of proteins and other biologics. Thus, under designated conditions, an antibody binds preferentially to a particular target protein, peptide or polysaccharide (such as an antigen present on the surface of a pathogen, for example PfCSP) and does not bind in a significant amount to other proteins present in the sample or subject. Specific binding can be determined by standard methods. See Harlow & Lane, Antibodies, A Laboratory Manual, 2nd ed., Cold Spring Harbor Publications, New York (2013), for a description of immunoassay formats and conditions that can be used to determine specific immunoreactivity. With reference to an antibody-antigen complex, specific binding of the antigen and antibody has a KD of less than about 10-7 Molar, such as less than about 10-8 Molar, 10-9, or even less than about 10-10 Molar. KD refers to the dissociation constant for a given interaction, such as a polypeptide ligand interaction or an antibody antigen interaction. For example, for the bimolecular interaction of an antibody or antigen binding fragment and an antigen it is the concentration of the individual components of the bimolecular interaction divided by the concentration of the complex. An antibody that specifically binds to an epitope on PfCSP is an antibody that binds substantially to PfCSP, including cells or tissue expressing PfCSP, substrate to which the PfCSP is attached, or PfCSP in a biological specimen. It is, of course, recognized that a certain degree of non-specific interaction may occur between an antibody and a non-target (such as a cell that does not express PfCSP). Typically, specific binding results in a much stronger association between the antibody and protein or cells bearing the antigen than between the antibody and protein or cells lacking the antigen. Specific binding typically results in greater than 2-fold, such as greater than 5-fold, greater than 10-fold, or greater than 100-fold increase in amount of bound antibody (per unit time) to a protein including the epitope or cell or tissue expressing the target epitope as compared to a protein or cell or tissue lacking this epitope. Specific binding to a protein under such conditions requires an antibody that is selected for its specificity for a particular protein. A
variety of immunoassay formats are appropriate for selecting antibodies or other ligands specifically immunoreactive with a particular protein. For example, solid-phase ELISA immunoassays are routinely used to select monoclonal antibodies specifically immunoreactive with a protein. Subject: Living multi-cellular vertebrate organisms, a category that includes human and non- human mammals. In an example, a subject is a human. In an additional example, a subject is selected that is in need of inhibiting a P. falciparum infection. For example, the subject is uninfected and at risk of P. falciparum infection. Transformed: A transformed cell is a cell into which a nucleic acid molecule has been introduced by molecular biology techniques. As used herein, the term transformed and the like (e.g., transformation, transfection, transduction, etc.) encompasses all techniques by which a nucleic acid molecule might be introduced into such a cell, including transduction with viral vectors, transformation with plasmid vectors, and introduction of DNA by electroporation, lipofection, and particle gun acceleration. Vector: An entity containing a nucleic acid molecule (such as a DNA or RNA molecule) bearing a promoter(s) that is operationally linked to the coding sequence of a protein of interest and can express the coding sequence. Non-limiting examples include a naked or packaged (lipid and/or protein) DNA, a naked or packaged RNA, a subcomponent of a virus or bacterium or other microorganism that may be replication- incompetent, or a virus or bacterium or other microorganism that may be replication-competent. A vector is sometimes referred to as a construct. Recombinant DNA vectors are vectors having recombinant DNA. A vector can include nucleic acid sequences that permit it to replicate in a host cell, such as an origin of replication. A vector can also include one or more selectable marker genes and other genetic elements. Viral vectors are recombinant nucleic acid vectors having at least some nucleic acid sequences derived from one or more viruses. In some aspects, a viral vector comprises a nucleic acid molecule encoding a disclosed antibody or antigen binding fragment that specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the viral vector can be an adeno-associated virus (AAV) vector. II. Description of Several Aspects A. Neutralizing Monoclonal Antibodies to PfCSP and Antigen Binding Fragments Thereof Isolated monoclonal antibodies and antigen binding fragments that specifically bind an epitope on PfCSP are provided. The antibodies and antigen binding fragments can be fully human. The antibodies and antigen binding fragments can neutralize P. falciparum, for example the disclosed antibodies can inhibit P. falciparum sporozoite infection of hepatocytes in vitro and P. falciparum sporozoite invasion of liver in vivo. Also disclosed herein are compositions comprising the antibodies and antigen binding fragments and a pharmaceutically acceptable carrier. Nucleic acids encoding the antibodies or antigen binding fragments, expression vectors (such as DNA and RNA vectors for expression and delivery, as well as adeno-associated virus (AAV) viral vectors) comprising these nucleic acids are also provided. The antibodies, antigen binding fragments, nucleic acid molecules, host cells, and compositions can be used for research, diagnostic and prophylactic purposes. For example, the disclosed antibodies and antigen binding fragments can be used
to diagnose a subject with a P. falciparum infection, or can be administered prophylactically to inhibit P. falciparum infection in a subject. 1. Exemplary monoclonal antibodies and antigen binding fragments The discussion of monoclonal antibodies below refers to isolated monoclonal antibodies that include heavy and/or light chain variable domains (or antigen binding fragments thereof) comprising a CDR1, CDR2, and/or CDR3 with reference to the IMMGT numbering scheme (unless the context indicates otherwise). Various CDR numbering schemes (such as the Kabat, Chothia or IMGT numbering schemes) can be used to determine CDR positions. The amino acid sequence and the CDR positions (according to the IMGT numbering scheme) of the heavy and light chains of exemplary monoclonal antibodies that bind to PfCSP and neutralize P. falciparum are shown in Table 1. Table 1. IMGT CDR sequences of PfCSP specific antibodies MAD21-17 VH CDR
LCDR1 27-38 QSLLYSSTNKNY 26 LCDR2 56-58 WAS 27 LCDR3 95-103 YYISPLT 28
MAD22-39 VH VH SEQ ID NO: 15 residues AA Se uence CDR
a. MAD21-17 In some aspects, the antibody or antigen binding fragment is based on or derived from the MAD21- 17 antibody, and specifically binds to PfCSP and neutralizes P. falciparum. For example, the antibody or antigen binding fragment comprises a VH and a VL comprising the HCDR1, the HCDR2, and the HCDR3, and the LCDR1, the LCDR2, and the LCDR3, respectively (for example, according to IMGT, Kabat, or Chothia), of the MAD21-17 antibody, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH comprising an amino acid sequence at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequence set forth as SEQ ID NO: 1, and specifically binds to PfCSP and neutralizes P. falciparum. In more aspects, the antibody or antigen binding fragment comprises a VL comprising an amino acid sequence at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequence set forth as SEQ ID NO: 2, and specifically binds to PfCSP and neutralizes P. falciparum. In additional aspects, the antibody or antigen binding fragment comprises a VH and a VL independently comprising amino acid sequences at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequences set forth as SEQ ID NOs: 1 and 2, respectively, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 23, 24, and 25, respectively, and a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 26, 27, and 28, respectively, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 23, 24, and 25, respectively, a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 26, 27, and 28, respectively, wherein the VH comprises an amino acid sequence at least 90% identical to SEQ ID NO: 1 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 1), the VL comprises an amino acid sequence at least 90% identical to SEQ ID NO: 2 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 2 and the antibody or antigen binding fragment specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 23, 24, and 25, respectively, a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 26, 27, and 28, respectively, wherein the framework regions of the VH comprise up to 10 (such as up to 1, up to 2, up to 3, up to 4, up to 5, up to 6, up to 7, up to 8, or up to 9) amino acid substitutions (such as conservative amino acid substitutions) compared to the amino acid sequence set forth as one of SEQ ID NO: 1, and the framework regions of the VL comprise up to 10 (such as up to 1, up to 2, up to 3, up to 4, up to 5, up to 6, up to 7, up to 8, or up to 9) amino acid substitutions (such as conservative amino acid substitutions) compared to the amino acid sequence set forth as one of SEQ ID NO: 2, and the antibody or antigen binding fragment specifically binds to PfCSP and neutralizes P. falciparum.
In additional aspects, the antibody or antigen binding fragment comprises a VH comprising the amino acid sequence set forth as SEQ ID NO: 1, and specifically binds to PfCSP and neutralizes P. falciparum. In more aspects, the antibody or antigen binding fragment comprises a VL comprising the amino acid sequence set forth as SEQ ID NO: 2, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH and a VL comprising the amino acid sequences set forth as SEQ ID NOs: 1 and 2, respectively, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the disclosed antibodies and antigen binding fragments inhibit the invasion of Plasmodium falciparum sporozoites into human liver cells, and/or reduce pathogen load Plasmodium falciparum in a subject, compared to a control. b. MAD21-46 In some aspects, the antibody or antigen binding fragment is based on or derived from the MAD21- 46 antibody, and specifically binds to PfCSP and neutralizes P. falciparum. For example, the antibody or antigen binding fragment comprises a VH and a VL comprising the HCDR1, the HCDR2, and the HCDR3, and the LCDR1, the LCDR2, and the LCDR3, respectively (for example, according to IMGT, Kabat, or Chothia), of the MAD21-46 antibody, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH comprising an amino acid sequence at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequence set forth as SEQ ID NO: 3, and specifically binds to PfCSP and neutralizes P. falciparum. In more aspects, the antibody or antigen binding fragment comprises a VL comprising an amino acid sequence at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequence set forth as SEQ ID NO: 4, and specifically binds to PfCSP and neutralizes P. falciparum. In additional aspects, the antibody or antigen binding fragment comprises a VH and a VL independently comprising amino acid sequences at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequences set forth as SEQ ID NOs: 3 and 4, respectively, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 29, 24, and 25, respectively, and a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 26, 27, and 28, respectively, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 29, 24, and 25, respectively, a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 26, 27, and 28, respectively, wherein the VH comprises an amino acid sequence at least 90% identical to SEQ ID NO: 3 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 3), the VL comprises an amino acid sequence at least 90% identical to SEQ ID NO:
4 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 4 and the antibody or antigen binding fragment specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 29, 24, and 25, respectively, a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 26, 27, and 28, respectively, wherein the framework regions of the VH comprise up to 10 (such as up to 1, up to 2, up to 3, up to 4, up to 5, up to 6, up to 7, up to 8, or up to 9) amino acid substitutions (such as conservative amino acid substitutions) compared to the amino acid sequence set forth as one of SEQ ID NO: 3, and the framework regions of the VL comprise up to 10 (such as up to 1, up to 2, up to 3, up to 4, up to 5, up to 6, up to 7, up to 8, or up to 9) amino acid substitutions (such as conservative amino acid substitutions) compared to the amino acid sequence set forth as one of SEQ ID NO: 4, and the antibody or antigen binding fragment specifically binds to PfCSP and neutralizes P. falciparum. In additional aspects, the antibody or antigen binding fragment comprises a VH comprising the amino acid sequence set forth as SEQ ID NO: 3, and specifically binds to PfCSP and neutralizes P. falciparum. In more aspects, the antibody or antigen binding fragment comprises a VL comprising the amino acid sequence set forth as SEQ ID NO: 4, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH and a VL comprising the amino acid sequences set forth as SEQ ID NOs: 3 and 4, respectively, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the disclosed antibodies and antigen binding fragments inhibit the invasion of Plasmodium falciparum sporozoites into human liver cells, and/or reduce pathogen load Plasmodium falciparum in a subject, compared to a control. c. MAD21-53 In some aspects, the antibody or antigen binding fragment is based on or derived from the MAD21- 53 antibody, and specifically binds to PfCSP and neutralizes P. falciparum. For example, the antibody or antigen binding fragment comprises a VH and a VL comprising the HCDR1, the HCDR2, and the HCDR3, and the LCDR1, the LCDR2, and the LCDR3, respectively (for example, according to IMGT, Kabat, or Chothia), of the MAD21-53 antibody, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH comprising an amino acid sequence at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequence set forth as SEQ ID NO: 5, and specifically binds to PfCSP and neutralizes P. falciparum. In more aspects, the antibody or antigen binding fragment comprises a VL comprising an amino acid sequence at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequence set forth as SEQ ID NO: 6, and specifically binds to PfCSP and neutralizes P. falciparum. In additional aspects, the antibody or antigen binding fragment comprises a VH and a VL independently comprising amino acid sequences at least 90% (such as at least 95%,
at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequences set forth as SEQ ID NOs: 5 and 6, respectively, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 29, 24, and 30, respectively, and a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 26, 27, and 28, respectively, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 29, 24, and 30, respectively, a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 26, 27, and 28, respectively, wherein the VH comprises an amino acid sequence at least 90% identical to SEQ ID NO: 5 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 5), the VL comprises an amino acid sequence at least 90% identical to SEQ ID NO: 6 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 6 and the antibody or antigen binding fragment specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 29, 24, and 30, respectively, a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 26, 27, and 28, respectively, wherein the framework regions of the VH comprise up to 10 (such as up to 1, up to 2, up to 3, up to 4, up to 5, up to 6, up to 7, up to 8, or up to 9) amino acid substitutions (such as conservative amino acid substitutions) compared to the amino acid sequence set forth as one of SEQ ID NO: 5, and the framework regions of the VL comprise up to 10 (such as up to 1, up to 2, up to 3, up to 4, up to 5, up to 6, up to 7, up to 8, or up to 9) amino acid substitutions (such as conservative amino acid substitutions) compared to the amino acid sequence set forth as one of SEQ ID NO: 6, and the antibody or antigen binding fragment specifically binds to PfCSP and neutralizes P. falciparum. In additional aspects, the antibody or antigen binding fragment comprises a VH comprising the amino acid sequence set forth as SEQ ID NO: 5, and specifically binds to PfCSP and neutralizes P. falciparum. In more aspects, the antibody or antigen binding fragment comprises a VL comprising the amino acid sequence set forth as SEQ ID NO: 6, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH and a VL comprising the amino acid sequences set forth as SEQ ID NOs: 5 and 6, respectively, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the disclosed antibodies and antigen binding fragments inhibit the invasion of Plasmodium falciparum sporozoites into human liver cells, and/or reduce pathogen load Plasmodium falciparum in a subject, compared to a control. d. MAD21-95 In some aspects, the antibody or antigen binding fragment is based on or derived from the MAD21- 95 antibody, and specifically binds to PfCSP and neutralizes P. falciparum. For example, the antibody or
antigen binding fragment comprises a VH and a VL comprising the HCDR1, the HCDR2, and the HCDR3, and the LCDR1, the LCDR2, and the LCDR3, respectively (for example, according to IMGT, Kabat, or Chothia), of the MAD21-95 antibody, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH comprising an amino acid sequence at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequence set forth as SEQ ID NO: 7, and specifically binds to PfCSP and neutralizes P. falciparum. In more aspects, the antibody or antigen binding fragment comprises a VL comprising an amino acid sequence at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequence set forth as SEQ ID NO: 8, and specifically binds to PfCSP and neutralizes P. falciparum. In additional aspects, the antibody or antigen binding fragment comprises a VH and a VL independently comprising amino acid sequences at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequences set forth as SEQ ID NOs: 7 and 8, respectively, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 23, 24, and 25, respectively, and a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 31, 27, and 38, respectively, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 23, 24, and 25, respectively, a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 31, 27, and 38, respectively, wherein the VH comprises an amino acid sequence at least 90% identical to SEQ ID NO: 7 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 7), the VL comprises an amino acid sequence at least 90% identical to SEQ ID NO: 8 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 8 and the antibody or antigen binding fragment specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 23, 24, and 25, respectively, a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 31, 27, and 38, respectively, wherein the framework regions of the VH comprise up to 10 (such as up to 1, up to 2, up to 3, up to 4, up to 5, up to 6, up to 7, up to 8, or up to 9) amino acid substitutions (such as conservative amino acid substitutions) compared to the amino acid sequence set forth as one of SEQ ID NO: 7, and the framework regions of the VL comprise up to 10 (such as up to 1, up to 2, up to 3, up to 4, up to 5, up to 6, up to 7, up to 8, or up to 9) amino acid substitutions (such as conservative amino acid substitutions) compared to the amino acid sequence set forth as one of SEQ ID NO: 8, and the antibody or antigen binding fragment specifically binds to PfCSP and neutralizes P. falciparum. In additional aspects, the antibody or antigen binding fragment comprises a VH comprising the amino acid sequence set forth as SEQ ID NO: 7, and specifically binds to PfCSP and neutralizes P. falciparum. In more aspects, the antibody or antigen binding fragment comprises a VL comprising the
amino acid sequence set forth as SEQ ID NO: 8, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH and a VL comprising the amino acid sequences set forth as SEQ ID NOs: 7 and 8, respectively, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the disclosed antibodies and antigen binding fragments inhibit the invasion of Plasmodium falciparum sporozoites into human liver cells, and/or reduce pathogen load Plasmodium falciparum in a subject, compared to a control. e. MAD21-101 In some aspects, the antibody or antigen binding fragment is based on or derived from the MAD21- 101 antibody, and specifically binds to PfCSP and neutralizes P. falciparum. For example, the antibody or antigen binding fragment comprises a VH and a VL comprising the HCDR1, the HCDR2, and the HCDR3, and the LCDR1, the LCDR2, and the LCDR3, respectively (for example, according to IMGT, Kabat, or Chothia), of the MAD21-101 antibody, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH comprising an amino acid sequence at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequence set forth as SEQ ID NO: 9, and specifically binds to PfCSP and neutralizes P. falciparum. In more aspects, the antibody or antigen binding fragment comprises a VL comprising an amino acid sequence at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequence set forth as SEQ ID NO: 10, and specifically binds to PfCSP and neutralizes P. falciparum. In additional aspects, the antibody or antigen binding fragment comprises a VH and a VL independently comprising amino acid sequences at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequences set forth as SEQ ID NOs: 9 and 10, respectively, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 29, 24, and 33, respectively, and a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 26, 27, and 34, respectively, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 29, 24, and 33, respectively, a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 26, 27, and 34, respectively, wherein the VH comprises an amino acid sequence at least 90% identical to SEQ ID NO: 9 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 9), the VL comprises an amino acid sequence at least 90% identical to SEQ ID NO: 10 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 10 and the antibody or antigen binding fragment specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 29, 24, and 33, respectively, a VL comprising a LCDR1,
a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 26, 27, and 34, respectively, wherein the framework regions of the VH comprise up to 10 (such as up to 1, up to 2, up to 3, up to 4, up to 5, up to 6, up to 7, up to 8, or up to 9) amino acid substitutions (such as conservative amino acid substitutions) compared to the amino acid sequence set forth as one of SEQ ID NO: 9, and the framework regions of the VL comprise up to 10 (such as up to 1, up to 2, up to 3, up to 4, up to 5, up to 6, up to 7, up to 8, or up to 9) amino acid substitutions (such as conservative amino acid substitutions) compared to the amino acid sequence set forth as one of SEQ ID NO: 10, and the antibody or antigen binding fragment specifically binds to PfCSP and neutralizes P. falciparum. In additional aspects, the antibody or antigen binding fragment comprises a VH comprising the amino acid sequence set forth as SEQ ID NO: 9, and specifically binds to PfCSP and neutralizes P. falciparum. In more aspects, the antibody or antigen binding fragment comprises a VL comprising the amino acid sequence set forth as SEQ ID NO: 10, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH and a VL comprising the amino acid sequences set forth as SEQ ID NOs: 9 and 10, respectively, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the disclosed antibodies and antigen binding fragments inhibit the invasion of Plasmodium falciparum sporozoites into human liver cells, and/or reduce pathogen load Plasmodium falciparum in a subject, compared to a control. f. MAD22-17 In some aspects, the antibody or antigen binding fragment is based on or derived from the MAD22- 17 antibody, and specifically binds to PfCSP and neutralizes P. falciparum. For example, the antibody or antigen binding fragment comprises a VH and a VL comprising the HCDR1, the HCDR2, and the HCDR3, and the LCDR1, the LCDR2, and the LCDR3, respectively (for example, according to IMGT, Kabat, or Chothia), of the MAD22-17 antibody, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH comprising an amino acid sequence at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequence set forth as SEQ ID NO: 11, and specifically binds to PfCSP and neutralizes P. falciparum. In more aspects, the antibody or antigen binding fragment comprises a VL comprising an amino acid sequence at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequence set forth as SEQ ID NO: 12, and specifically binds to PfCSP and neutralizes P. falciparum. In additional aspects, the antibody or antigen binding fragment comprises a VH and a VL independently comprising amino acid sequences at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequences set forth as SEQ ID NOs: 11 and 12, respectively, and specifically binds to PfCSP and neutralizes P. falciparum.
In some aspects, the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 35, 36, and 37, respectively, and a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 38, 27, and 39, respectively, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 35, 36, and 37, respectively, a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 38, 27, and 39, respectively, wherein the VH comprises an amino acid sequence at least 90% identical to SEQ ID NO: 11 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 11), the VL comprises an amino acid sequence at least 90% identical to SEQ ID NO: 12 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 12 and the antibody or antigen binding fragment specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 35, 36, and 37, respectively, a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 38, 27, and 39, respectively, wherein the framework regions of the VH comprise up to 10 (such as up to 1, up to 2, up to 3, up to 4, up to 5, up to 6, up to 7, up to 8, or up to 9) amino acid substitutions (such as conservative amino acid substitutions) compared to the amino acid sequence set forth as one of SEQ ID NO: 11, and the framework regions of the VL comprise up to 10 (such as up to 1, up to 2, up to 3, up to 4, up to 5, up to 6, up to 7, up to 8, or up to 9) amino acid substitutions (such as conservative amino acid substitutions) compared to the amino acid sequence set forth as one of SEQ ID NO: 12, and the antibody or antigen binding fragment specifically binds to PfCSP and neutralizes P. falciparum. In additional aspects, the antibody or antigen binding fragment comprises a VH comprising the amino acid sequence set forth as SEQ ID NO: 11, and specifically binds to PfCSP and neutralizes P. falciparum. In more aspects, the antibody or antigen binding fragment comprises a VL comprising the amino acid sequence set forth as SEQ ID NO: 12, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH and a VL comprising the amino acid sequences set forth as SEQ ID NOs: 11 and 12, respectively, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the disclosed antibodies and antigen binding fragments inhibit the invasion of Plasmodium falciparum sporozoites into human liver cells, and/or reduce pathogen load Plasmodium falciparum in a subject, compared to a control. g. MAD22-38 In some aspects, the antibody or antigen binding fragment is based on or derived from the MAD22- 38 antibody, and specifically binds to PfCSP and neutralizes P. falciparum. For example, the antibody or antigen binding fragment comprises a VH and a VL comprising the HCDR1, the HCDR2, and the HCDR3,
and the LCDR1, the LCDR2, and the LCDR3, respectively (for example, according to IMGT, Kabat, or Chothia), of the MAD22-38 antibody, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH comprising an amino acid sequence at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequence set forth as SEQ ID NO: 13, and specifically binds to PfCSP and neutralizes P. falciparum. In more aspects, the antibody or antigen binding fragment comprises a VL comprising an amino acid sequence at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequence set forth as SEQ ID NO: 14, and specifically binds to PfCSP and neutralizes P. falciparum. In additional aspects, the antibody or antigen binding fragment comprises a VH and a VL independently comprising amino acid sequences at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequences set forth as SEQ ID NOs: 13 and 14, respectively, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 40, 41, and 42, respectively, and a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 43, 44, and 45, respectively, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 40, 41, and 42, respectively, a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 43, 44, and 45, respectively, wherein the VH comprises an amino acid sequence at least 90% identical to SEQ ID NO: 13 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 13), the VL comprises an amino acid sequence at least 90% identical to SEQ ID NO: 14 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 14 and the antibody or antigen binding fragment specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 40, 41, and 42, respectively, a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 43, 44, and 45, respectively, wherein the framework regions of the VH comprise up to 10 (such as up to 1, up to 2, up to 3, up to 4, up to 5, up to 6, up to 7, up to 8, or up to 9) amino acid substitutions (such as conservative amino acid substitutions) compared to the amino acid sequence set forth as one of SEQ ID NO: 13, and the framework regions of the VL comprise up to 10 (such as up to 1, up to 2, up to 3, up to 4, up to 5, up to 6, up to 7, up to 8, or up to 9) amino acid substitutions (such as conservative amino acid substitutions) compared to the amino acid sequence set forth as one of SEQ ID NO: 14, and the antibody or antigen binding fragment specifically binds to PfCSP and neutralizes P. falciparum. In additional aspects, the antibody or antigen binding fragment comprises a VH comprising the amino acid sequence set forth as SEQ ID NO: 13, and specifically binds to PfCSP and neutralizes P. falciparum. In more aspects, the antibody or antigen binding fragment comprises a VL comprising the
amino acid sequence set forth as SEQ ID NO: 14, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH and a VL comprising the amino acid sequences set forth as SEQ ID NOs: 13 and 14, respectively, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the disclosed antibodies and antigen binding fragments inhibit the invasion of Plasmodium falciparum sporozoites into human liver cells, and/or reduce pathogen load Plasmodium falciparum in a subject, compared to a control. h. MAD22-39 In some aspects, the antibody or antigen binding fragment is based on or derived from the MAD22- 39 antibody, and specifically binds to PfCSP and neutralizes P. falciparum. For example, the antibody or antigen binding fragment comprises a VH and a VL comprising the HCDR1, the HCDR2, and the HCDR3, and the LCDR1, the LCDR2, and the LCDR3, respectively (for example, according to IMGT, Kabat, or Chothia), of the MAD22-39 antibody, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH comprising an amino acid sequence at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequence set forth as SEQ ID NO: 15, and specifically binds to PfCSP and neutralizes P. falciparum. In more aspects, the antibody or antigen binding fragment comprises a VL comprising an amino acid sequence at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequence set forth as SEQ ID NO: 16, and specifically binds to PfCSP and neutralizes P. falciparum. In additional aspects, the antibody or antigen binding fragment comprises a VH and a VL independently comprising amino acid sequences at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequences set forth as SEQ ID NOs: 15 and 16, respectively, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 46, 47, and 48, respectively, and a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 49, 44, and 45, respectively, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 46, 47, and 48, respectively, a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 49, 44, and 45, respectively, wherein the VH comprises an amino acid sequence at least 90% identical to SEQ ID NO: 15 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 15), the VL comprises an amino acid sequence at least 90% identical to SEQ ID NO: 16 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 16 and the antibody or antigen binding fragment specifically binds to PfCSP and neutralizes P. falciparum.
In some aspects, the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 46, 47, and 48, respectively, a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 49, 44, and 45, respectively, wherein the framework regions of the VH comprise up to 10 (such as up to 1, up to 2, up to 3, up to 4, up to 5, up to 6, up to 7, up to 8, or up to 9) amino acid substitutions (such as conservative amino acid substitutions) compared to the amino acid sequence set forth as one of SEQ ID NO: 15, and the framework regions of the VL comprise up to 10 (such as up to 1, up to 2, up to 3, up to 4, up to 5, up to 6, up to 7, up to 8, or up to 9) amino acid substitutions (such as conservative amino acid substitutions) compared to the amino acid sequence set forth as one of SEQ ID NO: 16, and the antibody or antigen binding fragment specifically binds to PfCSP and neutralizes P. falciparum. In additional aspects, the antibody or antigen binding fragment comprises a VH comprising the amino acid sequence set forth as SEQ ID NO: 15, and specifically binds to PfCSP and neutralizes P. falciparum. In more aspects, the antibody or antigen binding fragment comprises a VL comprising the amino acid sequence set forth as SEQ ID NO: 16, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH and a VL comprising the amino acid sequences set forth as SEQ ID NOs: 15 and 16, respectively, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the disclosed antibodies and antigen binding fragments inhibit the invasion of Plasmodium falciparum sporozoites into human liver cells, and/or reduce pathogen load Plasmodium falciparum in a subject, compared to a control. i. MAD24-01 In some aspects, the antibody or antigen binding fragment is based on or derived from the MAD24- 01 antibody, and specifically binds to PfCSP and neutralizes P. falciparum. For example, the antibody or antigen binding fragment comprises a VH and a VL comprising the HCDR1, the HCDR2, and the HCDR3, and the LCDR1, the LCDR2, and the LCDR3, respectively (for example, according to IMGT, Kabat, or Chothia), of the MAD24-01 antibody, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH comprising an amino acid sequence at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequence set forth as SEQ ID NO: 17, and specifically binds to PfCSP and neutralizes P. falciparum. In more aspects, the antibody or antigen binding fragment comprises a VL comprising an amino acid sequence at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequence set forth as SEQ ID NO: 18, and specifically binds to PfCSP and neutralizes P. falciparum. In additional aspects, the antibody or antigen binding fragment comprises a VH and a VL independently comprising amino acid sequences at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequences set
forth as SEQ ID NOs: 17 and 18, respectively, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 50, 51, and 52, respectively, and a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 53, 54, and 55, respectively, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 50, 51, and 52, respectively, a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 53, 54, and 55, respectively, wherein the VH comprises an amino acid sequence at least 90% identical to SEQ ID NO: 17 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 17), the VL comprises an amino acid sequence at least 90% identical to SEQ ID NO: 18 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 18 and the antibody or antigen binding fragment specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 50, 51, and 52, respectively, a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 53, 54, and 55, respectively, wherein the framework regions of the VH comprise up to 10 (such as up to 1, up to 2, up to 3, up to 4, up to 5, up to 6, up to 7, up to 8, or up to 9) amino acid substitutions (such as conservative amino acid substitutions) compared to the amino acid sequence set forth as one of SEQ ID NO: 17, and the framework regions of the VL comprise up to 10 (such as up to 1, up to 2, up to 3, up to 4, up to 5, up to 6, up to 7, up to 8, or up to 9) amino acid substitutions (such as conservative amino acid substitutions) compared to the amino acid sequence set forth as one of SEQ ID NO: 18, and the antibody or antigen binding fragment specifically binds to PfCSP and neutralizes P. falciparum. In additional aspects, the antibody or antigen binding fragment comprises a VH comprising the amino acid sequence set forth as SEQ ID NO: 17, and specifically binds to PfCSP and neutralizes P. falciparum. In more aspects, the antibody or antigen binding fragment comprises a VL comprising the amino acid sequence set forth as SEQ ID NO: 18, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH and a VL comprising the amino acid sequences set forth as SEQ ID NOs: 17 and 18, respectively, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the disclosed antibodies and antigen binding fragments inhibit the invasion of Plasmodium falciparum sporozoites into human liver cells, and/or reduce pathogen load Plasmodium falciparum in a subject, compared to a control. j. MAD24-05 In some aspects, the antibody or antigen binding fragment is based on or derived from the MAD24- 05 antibody, and specifically binds to PfCSP and neutralizes P. falciparum. For example, the antibody or
antigen binding fragment comprises a VH and a VL comprising the HCDR1, the HCDR2, and the HCDR3, and the LCDR1, the LCDR2, and the LCDR3, respectively (for example, according to IMGT, Kabat, or Chothia), of the MAD24-05 antibody, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH comprising an amino acid sequence at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequence set forth as SEQ ID NO: 17, and specifically binds to PfCSP and neutralizes P. falciparum. In more aspects, the antibody or antigen binding fragment comprises a VL comprising an amino acid sequence at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequence set forth as SEQ ID NO: 18, and specifically binds to PfCSP and neutralizes P. falciparum. In additional aspects, the antibody or antigen binding fragment comprises a VH and a VL independently comprising amino acid sequences at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequences set forth as SEQ ID NOs: 17 and 18, respectively, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 56, 57, and 58, respectively, and a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 59, 60, and 61, respectively, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 56, 57, and 58, respectively, a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 59, 60, and 61, respectively, wherein the VH comprises an amino acid sequence at least 90% identical to SEQ ID NO: 17 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 17), the VL comprises an amino acid sequence at least 90% identical to SEQ ID NO: 18 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 18 and the antibody or antigen binding fragment specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 56, 57, and 58, respectively, a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 59, 60, and 61, respectively, wherein the framework regions of the VH comprise up to 10 (such as up to 1, up to 2, up to 3, up to 4, up to 5, up to 6, up to 7, up to 8, or up to 9) amino acid substitutions (such as conservative amino acid substitutions) compared to the amino acid sequence set forth as one of SEQ ID NO: 17, and the framework regions of the VL comprise up to 10 (such as up to 1, up to 2, up to 3, up to 4, up to 5, up to 6, up to 7, up to 8, or up to 9) amino acid substitutions (such as conservative amino acid substitutions) compared to the amino acid sequence set forth as one of SEQ ID NO: 18, and the antibody or antigen binding fragment specifically binds to PfCSP and neutralizes P. falciparum. In additional aspects, the antibody or antigen binding fragment comprises a VH comprising the amino acid sequence set forth as SEQ ID NO: 17, and specifically binds to PfCSP and neutralizes P.
falciparum. In more aspects, the antibody or antigen binding fragment comprises a VL comprising the amino acid sequence set forth as SEQ ID NO: 18, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH and a VL comprising the amino acid sequences set forth as SEQ ID NOs: 17 and 18, respectively, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the disclosed antibodies and antigen binding fragments inhibit the invasion of Plasmodium falciparum sporozoites into human liver cells, and/or reduce pathogen load Plasmodium falciparum in a subject, compared to a control. k. MAD24-52 In some aspects, the antibody or antigen binding fragment is based on or derived from the MAD24- 52 antibody, and specifically binds to PfCSP and neutralizes P. falciparum. For example, the antibody or antigen binding fragment comprises a VH and a VL comprising the HCDR1, the HCDR2, and the HCDR3, and the LCDR1, the LCDR2, and the LCDR3, respectively (for example, according to IMGT, Kabat, or Chothia), of the MAD24-52 antibody, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH comprising an amino acid sequence at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequence set forth as SEQ ID NO: 19, and specifically binds to PfCSP and neutralizes P. falciparum. In more aspects, the antibody or antigen binding fragment comprises a VL comprising an amino acid sequence at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequence set forth as SEQ ID NO: 20, and specifically binds to PfCSP and neutralizes P. falciparum. In additional aspects, the antibody or antigen binding fragment comprises a VH and a VL independently comprising amino acid sequences at least 90% (such as at least 95%, at least 96%, at least 97%, at least 98%, or at least 99%) identical to the amino acid sequences set forth as SEQ ID NOs: 19 and 20, respectively, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 62, 63, and 64, respectively, and a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 65, 66, and 67, respectively, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 62, 63, and 64, respectively, a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 65, 66, and 67, respectively, wherein the VH comprises an amino acid sequence at least 90% identical to SEQ ID NO: 19 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 19), the VL comprises an amino acid sequence at least 90% identical to SEQ ID NO: 20 (such as 95%, 96%, 97%, 98% or 99% identical to SEQ ID NO: 20 and the antibody or antigen binding fragment specifically binds to PfCSP and neutralizes P. falciparum.
In some aspects, the antibody or antigen binding fragment comprises a VH comprising a HCDR1, a HCDR2, and a HCDR3 as set forth as SEQ ID NOs: 62, 63, and 64, respectively, a VL comprising a LCDR1, a LCDR2, and a LCDR3 as set forth as SEQ ID NOs: 65, 66, and 67, respectively, wherein the framework regions of the VH comprise up to 10 (such as up to 1, up to 2, up to 3, up to 4, up to 5, up to 6, up to 7, up to 8, or up to 9) amino acid substitutions (such as conservative amino acid substitutions) compared to the amino acid sequence set forth as one of SEQ ID NO: 19, and the framework regions of the VL comprise up to 10 (such as up to 1, up to 2, up to 3, up to 4, up to 5, up to 6, up to 7, up to 8, or up to 9) amino acid substitutions (such as conservative amino acid substitutions) compared to the amino acid sequence set forth as one of SEQ ID NO: 20, and the antibody or antigen binding fragment specifically binds to PfCSP and neutralizes P. falciparum. In additional aspects, the antibody or antigen binding fragment comprises a VH comprising the amino acid sequence set forth as SEQ ID NO: 19, and specifically binds to PfCSP and neutralizes P. falciparum. In more aspects, the antibody or antigen binding fragment comprises a VL comprising the amino acid sequence set forth as SEQ ID NO: 20, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the antibody or antigen binding fragment comprises a VH and a VL comprising the amino acid sequences set forth as SEQ ID NOs: 19 and 20, respectively, and specifically binds to PfCSP and neutralizes P. falciparum. In some aspects, the disclosed antibodies and antigen binding fragments inhibit the invasion of Plasmodium falciparum sporozoites into human liver cells, and/or reduce pathogen load Plasmodium falciparum in a subject, compared to a control. 2. Additional Description of Antibodies and Antigen Binding Fragments The antibody or antigen binding fragment can be a human antibody or fragment thereof. Chimeric antibodies are also provided. The antibody or antigen binding fragment can include any suitable framework region, such as (but not limited to) a human framework region. Alternatively, a heterologous framework region, such as, but not limited to a mouse or monkey framework region, can be included in the heavy or light chain of the antibodies. The antibody can be of any isotype. The antibody can be, for example, an IgM or an IgG antibody, such as IgG1, IgG2, IgG3, or IgG4. The class of an antibody that specifically binds PfCSP can be switched with another. In one aspect, a nucleic acid molecule encoding VL or VH is isolated such that it does not include any nucleic acid sequences encoding the constant region of the light or heavy chain, respectively. A nucleic acid molecule encoding VL or VH is then operatively linked to a nucleic acid sequence encoding a CL or CH from a different class of immunoglobulin molecule. This can be achieved, for example, using a vector or nucleic acid molecule that comprises a CL or CH chain. For example, an antibody that specifically binds PfCSP, that was originally IgG may be class switched to an IgM. Class switching can be used to convert one IgG subclass to another, such as from IgG1 to IgG2, IgG3, or IgG4.
In an aspect, exemplary full-length heavy and light chain sequences containing the MAD21-101 VH and VL are provided as SEQ ID NOs: 157 and 158, respectively. In some examples, the disclosed antibodies are oligomers of antibodies, such as dimers, trimers, tetramers, pentamers, hexamers, septamers, octomers and so on. The antibody or antigen binding fragment can be derivatized or linked to another molecule (such as another peptide or protein). In general, the antibody or antigen binding fragment is derivatized such that the binding to P. falciparum is not affected adversely by the derivatization or labeling. For example, the antibody or antigen binding fragment can be functionally linked (by chemical coupling, genetic fusion, noncovalent association or otherwise) to one or more other molecular entities, such as another antibody (for example, a bi-specific antibody or a diabody), a detectable marker, an effector molecule, or a protein or peptide that can mediate association of the antibody or antibody portion with another molecule (such as a streptavidin core region or a polyhistidine tag). a. Binding affinity In several aspects, the antibody or antigen binding fragment specifically binds PfCSP with an affinity (e.g., measured by KD) of no more than 1.0 x 10-8 M, no more than 5.0 x 10-8 M, no more than 1.0 x 10-9 M, no more than 5.0 x 10-9 M, no more than 1.0 x 10-10 M, no more than 5.0 x 10-10 M, or no more than 1.0 x 10-11 M. KD can be measured, for example, by a radiolabeled antigen binding assay (RIA) performed with the Fab version of an antibody of interest and its antigen. In one assay, solution binding affinity of Fabs for antigen is measured by equilibrating Fab with a minimal concentration of (125I)-labeled antigen in the presence of a titration series of unlabeled antigen, then capturing bound antigen with an anti-Fab antibody-coated plate (see, e.g., Chen et al., J. Mol. Biol.293(4):865-881, 1999). To establish conditions for the assay, MICROTITER® multi-well plates (Thermo Scientific) are coated overnight with 5 μg/ml of a capturing anti-Fab antibody (Cappel Labs) in 50 mM sodium carbonate (pH 9.6), and subsequently blocked with 2% (w/v) bovine serum albumin in PBS for two to five hours at room temperature (approximately 23° C.). In a non-adsorbent plate (Nunc™ Catalog #269620), 100 μM or 26 pM [125I]-antigen are mixed with serial dilutions of a Fab of interest (e.g., consistent with assessment of the anti-VEGF antibody, Fab-12, in Presta et al., Cancer Res.57(20):4593-4599, 1997). The Fab of interest is then incubated overnight; however, the incubation may continue for a longer period (e.g., about 65 hours) to ensure that equilibrium is reached. Thereafter, the mixtures are transferred to the capture plate for incubation at room temperature (e.g., for one hour). The solution is then removed and the plate washed eight times with 0.1% polysorbate 20 (TWEEN-20®) in PBS. When the plates have dried, 150 μl/well of scintillant (MicroScint™-20; PerkinEmler) is added, and the plates are counted on a TOPCOUNT™ gamma counter (PerkinEmler) for ten minutes. Concentrations of each Fab that give less than or equal to 20% of maximal binding are chosen for use in competitive binding assays. In another assay, KD can be measured using surface plasmon resonance assays using a BIACORE®- 2000 or a BIACORE®-3000 (BIAcore, Inc., Piscataway, N.J.) at 25° C with immobilized antigen CM5
chips at ~10 response units (RU). Briefly, carboxymethylated dextran biosensor chips (CM5, BIACORE®, Inc.) are activated with N-ethyl-N′-(3-dimethylaminopropyl)-carbodiimide hydrochloride (EDC) and N- hydroxysuccinimide (NHS) according to the supplier's instructions. Antigen is diluted with 10 mM sodium acetate, pH 4.8, to 5 μg/ml (~0.2 μM) before injection at a flow rate of 5 l/minute to achieve approximately 10 response units (RU) of coupled protein. Following the injection of antigen, 1 M ethanolamine is injected to block unreacted groups. For kinetics measurements, two-fold serial dilutions of Fab (0.78 nM to 500 nM) are injected in PBS with 0.05% polysorbate 20 (TWEEN-20™) surfactant (PBST) at 25° C at a flow rate of approximately 25 l/min. Association rates (kon) and dissociation rates (koff) are calculated using a simple one-to-one Langmuir binding model (BIACORE® Evaluation Software version 3.2) by simultaneously fitting the association and dissociation sensorgrams. The equilibrium dissociation constant (KD) is calculated as the ratio koff/kon. See, e.g., Chen et al., J. Mol. Biol.293:865-881 (1999). If the on-rate exceeds 106 M−1 s−1 by the surface plasmon resonance assay above, then the on-rate can be determined by using a fluorescent quenching technique that measures the increase or decrease in fluorescence emission intensity (excitation=295 nm; emission=340 nm, 16 nm band-pass) at 25° C. of a 20 nM anti-antigen antibody (Fab form) in PBS, pH 7.2, in the presence of increasing concentrations of antigen as measured in a spectrometer, such as a stop-flow equipped spectrophometer (Aviv Instruments) or a 8000-series SLM-AMINCO™ spectrophotometer (ThermoSpectronic) with a stirred cuvette. b. Multispecific antibodies In some aspects, a multi-specific antibody, such as a bi-specific antibody, is provided that comprises an antibody or antigen binding fragment as provided herein, and specifically binds to PfCSP. Any suitable method can be used to design and produce the multi-specific antibody, such as crosslinking two or more antibodies, antigen binding fragments (such as scFvs) of the same type or of different types. Exemplary methods of making multispecific antibodies include those described in PCT Pub. No. WO2013/163427. Non-limiting examples of suitable crosslinkers include those that are heterobifunctional, having two distinctly reactive groups separated by an appropriate spacer (such as m-maleimidobenzoyl-N- hydroxysuccinimide ester) or homobifunctional (such as disuccinimidyl suberate). The multi-specific antibody may have any suitable format that allows for antigen binding by the antibody or antigen binding fragment as provided herein, such as any one of MAD21-17, MAD21-46, MAD21-53, MAD21-95, MAD21-101, MAD22-17, MAD22-38, MAD22-39, MAD24-01, MAD24-05, or MAD24-52, or an antigen binding fragment thereof. Bispecific single chain antibodies can be encoded by a single nucleic acid molecule. Non-limiting examples of bispecific single chain antibodies, as well as methods of constructing such antibodies are provided in U.S. Pat. Nos.8,076,459, 8,017,748, 8,007,796, 7,919,089, 7,820,166, 7,635,472, 7,575,923, 7,435,549, 7,332,168, 7,323,440, 7,235,641, 7,229,760, 7,112,324, 6,723,538. Additional examples of bispecific single chain antibodies can be found in PCT application No. WO 99/54440; Mack et al., J. Immunol., 158(8):3965-3970, 1997; Mack et al., Proc. Natl. Acad. Sci. U.S.A., 92(15):7021-7025, 1995; Kufer et al., Cancer Immunol. Immunother., 45(3-4):193-197,
1997; Löffler et al., Blood, 95(6):2098-2103, 2000; and Brühl et al., J. Immunol., 166(4):2420-2426, 2001. Production of bispecific Fab-scFv (“bibody”) molecules are described, for example, in Schoonjans et al. (J. Immunol., 165(12):7050-7057, 2000) and Willems et al. (J. Chromatogr. B Analyt. Technol. Biomed Life Sci.786(1-2):161-176, 2003). For bibodies, a scFv molecule can be fused to one of the VL-CL (L) or VH- CH1 chains, e.g., to produce a bibody one scFv is fused to the C-term of a Fab chain. c. Fragments Antigen binding fragments are encompassed by the present disclosure, such as Fab, F(ab')2, and Fv which include a heavy chain and VL and specifically bind PfCSP. These antibody fragments retain the ability to selectively bind with the antigen and are “antigen-binding” fragments. Non-limiting examples of such fragments include: (1) Fab, the fragment which contains a monovalent antigen-binding fragment of an antibody molecule, can be produced by digestion of whole antibody with the enzyme papain to yield an intact light chain and a portion of one heavy chain; (2) Fab', the fragment of an antibody molecule can be obtained by treating whole antibody with pepsin, followed by reduction, to yield an intact light chain and a portion of the heavy chain; (3) (Fab')2, the fragment of the antibody that can be obtained by treating whole antibody with the enzyme pepsin without subsequent reduction; F(ab')2 is a dimer of two Fab' fragments held together by two disulfide bonds; (4) Fv, a genetically engineered fragment containing the VL and VL expressed as two chains; and (5) Single chain antibody (such as scFv), defined as a genetically engineered molecule containing the VH and the VL linked by a suitable polypeptide linker as a genetically fused single chain molecule (see, e.g., Ahmad et al., Clin. Dev. Immunol., 2012, doi:10.1155/2012/980250; Marbry and Snavely, IDrugs, 13(8):543-549, 2010). The intramolecular orientation of the VH-domain and the VL- domain in a scFv, is not decisive for the provided antibodies (e.g., for the provided multispecific antibodies). Thus, scFvs with both possible arrangements (VH-domain-linker domain-VL-domain; VL-domain-linker domain-VH-domain) may be used. (6) A dimer of a single chain antibody (scFV2), defined as a dimer of a scFV. This has also been termed a “miniantibody.” Any suitable method of producing the above-discussed antinge binding fragments may be used. Non-limiting examples are provided in Harlow and Lane, Antibodies: A Laboratory Manual, 2nd, Cold Spring Harbor Laboratory, New York, 2013. Antigen binding fragments can be prepared by proteolytic hydrolysis of the antibody or by expression in a host cell (such as an E. coli cell) of DNA encoding the fragment. Antigen binding fragments can also be obtained by pepsin or papain digestion of whole antibodies by conventional methods. For example, antigen binding fragments can be produced by enzymatic cleavage of antibodies with pepsin to
provide a 5S fragment denoted F(ab')2. This fragment can be further cleaved using a thiol reducing agent, and optionally a blocking group for the sulfhydryl groups resulting from cleavage of disulfide linkages, to produce 3.5S Fab' monovalent fragments. Other methods of cleaving antibodies, such as separation of heavy chains to form monovalent light- heavy chain fragments, further cleavage of fragments, or other enzymatic, chemical, or genetic techniques may also be used, so long as the fragments bind to the antigen that is recognized by the intact antibody. d. Variants In some aspects, amino acid sequence variants of the antibodies and antigen binding fragments provided herein are provided. For example, it may be desirable to improve the binding affinity and/or other biological properties of the antibody. Amino acid sequence variants of an antibody may be prepared by introducing appropriate modifications into the nucleotide sequence encoding the antibody, or by peptide synthesis. Such modifications include, for example, deletions from, and/or insertions into and/or substitutions of residues within the amino acid sequences of the antibody. Any combination of deletion, insertion, and substitution can be made to arrive at the final construct, provided that the final construct possesses the desired characteristics, e.g., antigen-binding. In some aspects, antibody variants having one or more amino acid substitutions are provided. Sites of interest for substitutional mutagenesis include the CDRs and the framework regions. Amino acid substitutions may be introduced into an antibody of interest and the products screened for a desired activity, e.g., retained/improved antigen binding, decreased immunogenicity, or improved ADCC or CDC. The variants typically retain amino acid residues necessary for correct folding and stabilizing between the VH and the VL regions, and will retain the charge characteristics of the residues in order to preserve the low pI and low toxicity of the molecules. Amino acid substitutions can be made in the VH and the VL regions to increase yield. In some aspects, the antibody or antigen binding fragment can include up to 10 (such as up to 1, up to 2, up to 3, up to 4, up to 5, up to 6, up to 7, up to 8, or up to 9) amino acid substitutions (such as conservative amino acid substitutions) in the framework regions of the heavy chain of the antibody, or the light chain of the antibody, or the heavy and light chains of the antibody, compared to known framework regions, or compared to the framework regions of an antibody as provided herein, and maintain the specific binding activity for PfCSP. In some aspects, substitutions, insertions, or deletions may occur within one or more CDRs so long as such alterations do not substantially reduce the ability of the antibody to bind antigen. For example, conservative alterations (e.g., conservative substitutions as provided herein) that do not substantially reduce binding affinity may be made in CDRs. In some aspects of the variant VH and VL sequences provided above, each CDR either is unaltered, or contains no more than one, two or three amino acid substitutions. To increase binding affinity of the antibody, the VL and VH segments can be randomly mutated, such as within HCDR3 region or the LCDR3 region, in a process analogous to the in vivo somatic mutation
process responsible for affinity maturation of antibodies during a natural immune response. Thus in vitro affinity maturation can be accomplished by amplifying VH and VL regions using PCR primers complementary to the HCDR3 or LCDR3, respectively. In this process, the primers have been “spiked” with a random mixture of the four nucleotide bases at certain positions such that the resultant PCR products encode VH and VL segments into which random mutations have been introduced into the VH and/or VL CDR3 regions. These randomly mutated VH and VL segments can be tested to determine the binding affinity for PfCSP. In some aspects, an antibody or antigen binding fragment is altered to increase or decrease the extent to which the antibody or antigen binding fragment is glycosylated. Addition or deletion of glycosylation sites may be conveniently accomplished by altering the amino acid sequence such that one or more glycosylation sites is created or removed. Where the antibody comprises an Fc region, the carbohydrate attached thereto may be altered. Native antibodies produced by mammalian cells typically comprise a branched, biantennary oligosaccharide that is generally attached by an N-linkage to Asn297 of the CH2 domain of the Fc region. See, e.g., Wright et al. Trends Biotechnol.15(1):26-32, 1997. The oligosaccharide may include various carbohydrates, e.g., mannose, N-acetyl glucosamine (GlcNAc), galactose, and sialic acid, as well as a fucose attached to a GlcNAc in the “stem” of the biantennary oligosaccharide structure. In some aspects, modifications of the oligosaccharide in an antibody may be made in order to create antibody variants with certain improved properties. In one aspect, antibody variants are provided having a carbohydrate structure that lacks fucose attached (directly or indirectly) to an Fc region. For example, the amount of fucose in such antibody may be from 1% to 80%, from 1% to 65%, from 5% to 65% or from 20% to 40%. The amount of fucose is determined by calculating the average amount of fucose within the sugar chain at Asn297, relative to the sum of all glycostructures attached to Asn 297 (e.g. complex, hybrid and high mannose structures) as measured by MALDI-TOF mass spectrometry, as described in WO 2008/077546, for example. Asn297 refers to the asparagine residue located at about position 297 in the Fc region; however, Asn297 may also be located about ±3 amino acids upstream or downstream of position 297, i.e., between positions 294 and 300, due to minor sequence variations in antibodies. Such fucosylation variants may have improved ADCC function. See, e.g., US Patent Publication Nos. US 2003/0157108 (Presta, L.); US 2004/0093621 (Kyowa Hakko Kogyo Co., Ltd). Examples of publications related to “defucosylated” or “fucose-deficient” antibody variants include: US 2003/0157108; WO 2000/61739; WO 2001/29246; US 2003/0115614; US 2002/0164328; US 2004/0093621; US 2004/0132140; US 2004/0110704; US 2004/0110282; US 2004/0109865; WO 2003/085119; WO 2003/084570; WO 2005/035586; WO 2005/035778; WO2005/053742; WO 2002/031140; Okazaki et al., J. Mol. Biol., 336(5):1239-1249, 2004; Yamane- Ohnuki et al., Biotechnol. Bioeng.87(5):614-622, 2004. Examples of cell lines capable of producing defucosylated antibodies include Lec 13 CHO cells deficient in protein fucosylation (Ripka et al., Arch. Biochem. Biophys.249(2):533-545, 1986; US Pat. Appl. No. US 2003/0157108 and WO 2004/056312,
especially at Example 11), and knockout cell lines, such as alpha-1,6-fucosyltransferase gene, FUT8, knockout CHO cells (see, e.g., Yamane-Ohnuki et al., Biotechnol. Bioeng., 87(5): 614-622, 2004; Kanda et al., Biotechnol. Bioeng., 94(4):680-688, 2006; and WO2003/085107). Antibody variants are further provided with bisected oligosaccharides, e.g., in which a biantennary oligosaccharide attached to the Fc region of the antibody is bisected by GlcNAc. Such antibody variants may have reduced fucosylation and/or improved ADCC function. Examples of such antibody variants are described, e.g., in WO 2003/011878 (Jean-Mairet et al.); U.S. Pat. No.6,602,684 (Umana et al.); and US 2005/0123546 (Umana et al.). Antibody variants with at least one galactose residue in the oligosaccharide attached to the Fc region are also provided. Such antibody variants may have improved CDC function. Such antibody variants are described, e.g., in WO 1997/30087; WO 1998/58964; and WO 1999/22764. In several aspects, the constant region of the antibody (such as any one of MAD21-17, MAD21-46, MAD21-53, MAD21-95, MAD21-101, MAD22-17, MAD22-38, MAD22-39, MAD24-01, MAD24-05, or MAD24-52) comprises one or more amino acid substitutions to optimize in vivo half-life of the antibody. The serum half-life of IgG Abs is regulated by the neonatal Fc receptor (FcRn). Thus, in several aspects, the antibody comprises an amino acid substitution that increases binding to the FcRn. Non-limiting examples of such substitutions include substitutions at IgG constant regions T250Q and M428L (see, e.g., Hinton et al., J Immunol., 176(1):346-356, 2006); M428L and N434S (the “LS” mutation, see, e.g., Zalevsky, et al., Nature Biotechnol., 28(2):157-159, 2010); N434A (see, e.g., Petkova et al., Int. Immunol., 18(12):1759-1769, 2006); T307A, E380A, and N434A (see, e.g., Petkova et al., Int. Immunol., 18(12):1759-1769, 2006); and M252Y, S254T, and T256E (see, e.g., Dall’Acqua et al., J. Biol. Chem., 281(33):23514-23524, 2006). The disclosed antibodies (such as any one of MAD21-17, MAD21-46, MAD21-53, MAD21-95, MAD21-101, MAD22-17, MAD22-38, MAD22-39, MAD24-01, MAD24-05, or MAD24-52) and antigen binding fragments can be linked to or comprise an Fc polypeptide including any of the substitutions listed above, for example, the Fc polypeptide can include the M428L and N434S substitutions. In some aspects, the constant region of the antibody comprises one or more amino acid substitutions to optimize ADCC. ADCC is mediated primarily through a set of closely related Fcγ receptors. In some aspects, the antibody comprises one or more amino acid substitutions that increase binding to FcγRIIIa. Non-limiting examples of such substitutions include substitutions at IgG constant regions S239D and I332E (see, e.g., Lazar et al., Proc. Natl., Acad. Sci. U.S.A., 103(11):4005-4010, 2006); and S239D, A330L, and I332E (see, e.g., Lazar et al., Proc. Natl., Acad. Sci. U.S.A., 103(11):4005-4010, 2006). Combinations of the above substitutions are also included, to generate an IgG constant region with increased binding to FcRn and FcγRIIIa. The combinations increase antibody half-life and ADCC. For example, such combinations include antibodies with the following amino acid substitutions in the Fc region: (1) S239D/I332E and T250Q/M428L; (2) S239D/I332E and M428L/N434S; (3) S239D/I332E and N434A; (4) S239D/I332E and T307A/E380A/N434A; (5) S239D/I332E and M252Y/S254T/T256E; (6) S239D/A330L/I332E and 250Q/M428L; (7) S239D/A330L/I332E and M428L/N434S; (8) S239D/A330L/I332E and N434A; (9) S239D/A330L/I332E and T307A/E380A/N434A; or (10)
S239D/A330L/I332E and M252Y/S254T/T256E. In some examples, the antibodies, or an antigen binding fragment thereof is modified such that it is directly cytotoxic to infected cells, or uses natural defenses such as complement, ADCC, or phagocytosis by macrophages. In some aspects, an antibody provided herein may be further modified to contain additional nonproteinaceous moieties. The moieties suitable for derivatization of the antibody include but are not limited to water soluble polymers. Non-limiting examples of water soluble polymers include, but are not limited to, polyethylene glycol (PEG), copolymers of ethylene glycol/propylene glycol, carboxymethylcellulose, dextran, polyvinyl alcohol, polyvinyl pyrrolidone, poly-1,3-dioxolane, poly-1,3,6- trioxane, ethylene/maleic anhydride copolymer, polyaminoacids (either homopolymers or random copolymers), and dextran or poly(n-vinyl pyrrolidone)polyethylene glycol, propropylene glycol homopolymers, prolypropylene oxide/ethylene oxide co-polymers, polyoxyethylated polyols (e.g., glycerol), polyvinyl alcohol, and mixtures thereof. Polyethylene glycol propionaldehyde may have advantages in manufacturing due to its stability in water. The polymer may be of any molecular weight, and may be branched or unbranched. The number of polymers attached to the antibody may vary, and if more than one polymer are attached, they can be the same or different molecules. In general, the number and/or type of polymers used for derivatization can be determined based on considerations including, but not limited to, the particular properties or functions of the antibody to be improved, whether the antibody derivative will be used in an application under defined conditions, etc. B. Conjugates The antibodies and antigen binding fragments that specifically bind to PfCSP (such as any one of MAD21-17, MAD21-46, MAD21-53, MAD21-95, MAD21-101, MAD22-17, MAD22-38, MAD22-39, MAD24-01, MAD24-05, or MAD24-52) can be conjugated to an agent, such as an effector molecule or detectable marker. Both covalent and noncovalent attachment means may be used. Various effector molecules and detectable markers can be used, including (but not limited to) toxins and radioactive agents such as 125I, 32P, 14C, 3H and 35S and other labels, target moieties and ligands, etc. The choice of a particular effector molecule or detectable marker depends on the particular target molecule or cell, and the desired biological effect. The procedure for attaching an effector molecule or detectable marker to an antibody or antigen binding fragment varies according to the chemical structure of the effector. Polypeptides typically contain a variety of functional groups, such as carboxyl (-COOH), free amine (-NH2) or sulfhydryl (-SH) groups, which are available for reaction with a suitable functional group on a polypeptide to result in the binding of the effector molecule or detectable marker. Alternatively, the antibody or antigen binding fragment is derivatized to expose or attach additional reactive functional groups. The derivatization may involve attachment of any suitable linker molecule. The linker is capable of forming covalent bonds to both the antibody or antigen binding fragment and to the effector molecule or detectable marker. Suitable linkers include, but are not limited to, straight or branched-chain carbon linkers, heterocyclic carbon linkers, or
peptide linkers. Where the antibody or antigen binding fragment and the effector molecule or detectable marker are polypeptides, the linkers may be joined to the constituent amino acids through their side chains (such as through a disulfide linkage to cysteine) or the alpha carbon, or through the amino, and/or carboxyl groups of the terminal amino acids. In view of the large number of methods that have been reported for attaching a variety of radiodiagnostic compounds, radiotherapeutic compounds, labels (such as enzymes or fluorescent molecules), toxins, and other agents to antibodies, a suitable method for attaching a given agent to an antibody or antigen binding fragment or other polypeptide can be determined. The antibody or antigen binding fragment can be conjugated with a detectable marker; for example, a detectable marker capable of detection by ELISA, spectrophotometry, flow cytometry, microscopy or diagnostic imaging techniques (such as CT, computed axial tomography (CAT), MRI, magnetic resonance tomography (MTR), ultrasound, fiberoptic examination, and laparoscopic examination). Specific, non- limiting examples of detectable markers include fluorophores, chemiluminescent agents, enzymatic linkages, radioactive isotopes and heavy metals or compounds (for example super paramagnetic iron oxide nanocrystals for detection by MRI). For example, useful detectable markers include fluorescent compounds, including fluorescein, fluorescein isothiocyanate, rhodamine, 5-dimethylamine-1-napthalenesulfonyl chloride, phycoerythrin, lanthanide phosphors and the like. Bioluminescent markers are also of use, such as luciferase, green fluorescent protein (GFP), and yellow fluorescent protein (YFP). An antibody or antigen binding fragment can also be conjugated with enzymes that are useful for detection, such as horseradish peroxidase, β- galactosidase, luciferase, alkaline phosphatase, glucose oxidase and the like. When an antibody or antigen binding fragment is conjugated with a detectable enzyme, it can be detected by adding additional reagents that the enzyme uses to produce a reaction product that can be discerned. For example, when the agent horseradish peroxidase is present, the addition of hydrogen peroxide and diaminobenzidine leads to a colored reaction product, which is visually detectable. An antibody or antigen binding fragment may also be conjugated with biotin, and detected through indirect measurement of avidin or streptavidin binding. It should be noted that the avidin itself can be conjugated with an enzyme or a fluorescent label. The antibody or antigen binding fragment can be conjugated with a paramagnetic agent, such as gadolinium. Paramagnetic agents such as superparamagnetic iron oxide are also of use as labels. Antibodies can also be conjugated with lanthanides (such as europium and dysprosium), and manganese. An antibody or antigen binding fragment may also be labeled with a predetermined polypeptide epitope recognized by a secondary reporter (such as leucine zipper pair sequences, binding sites for secondary antibodies, metal binding domains, epitope tags). The antibody or antigen binding fragment can also be conjugated with a radiolabeled amino acid, for example, for diagnostic purposes. For instance, the radiolabel may be used to detect PfCSP expressing cells by radiography, emission spectra, or other diagnostic techniques. Examples of labels for polypeptides include, but are not limited to, the following radioisotopes: 3H, 14C, 35S, 90Y, 99mTc, 111In, 125I, 131I. The radiolabels may be detected, for example, using photographic film or scintillation counters, fluorescent
markers may be detected using a photodetector to detect emitted illumination. Enzymatic labels are typically detected by providing the enzyme with a substrate and detecting the reaction product produced by the action of the enzyme on the substrate, and colorimetric labels are detected by simply visualizing the colored label. The average number of effector molecule or detectable marker moieties per antibody or antigen binding fragment in a conjugate can range, for example, from 1 to 20 moieties per antibody or antigen binding fragment. In some aspects, the average number of effector molecules or detectable marker moieties per antibody or antigen binding fragment in a conjugate range from about 1 to about 2, from about 1 to about 3, about 1 to about 8; from about 2 to about 6; from about 3 to about 5; or from about 3 to about 4. The loading (for example, effector molecule per antibody ratio) of a conjugate may be controlled in different ways, for example, by: (i) limiting the molar excess of effector molecule-linker intermediate or linker reagent relative to antibody, (ii) limiting the conjugation reaction time or temperature, (iii) partial or limiting reducing conditions for cysteine thiol modification, (iv) engineering by recombinant techniques the amino acid sequence of the antibody such that the number and position of cysteine residues is modified for control of the number or position of linker-effector molecule attachments. C. Polynucleotides and Expression Nucleic acid molecules (for example, cDNA or RNA molecules) encoding the amino acid sequences of antibodies, antigen binding fragments, and conjugates that specifically bind to PfCSP (such as any one of MAD21-17, MAD21-46, MAD21-53, MAD21-95, MAD21-101, MAD22-17, MAD22-38, MAD22-39, MAD24-01, MAD24-05, or MAD24-52) are provided. Nucleic acids encoding these molecules can readily be produced using the amino acid sequences provided herein (such as the CDR sequences and VH and VL sequences), sequences available in the art (such as framework or constant region sequences), and the genetic code. In several aspects, a nucleic acid molecules can encode the VH, the VL, or both the VH and VL (for example in a bicistronic expression vector) of a disclosed antibody or antigen binding fragment. In several aspects, the nucleic acid molecules can be expressed in a host cell (such as a mammalian cell) to produce a disclosed antibody or antigen binding fragment. Exemplary nucleic acid sequences encoding the heavy and light chain variable regions of the disclosed antibodies are provided as SEQ ID NOs: 73-94. The genetic code can be used to construct a variety of functionally equivalent nucleic acid sequences, such as nucleic acids which differ in sequence but which encode the same antibody sequence, or encode a conjugate or fusion protein including the VL and/or VH nucleic acid sequence. Nucleic acid molecules encoding the antibodies, antigen binding fragments, and conjugates that specifically bind to PfCSP can be prepared by any suitable method including, for example, cloning of appropriate sequences or by direct chemical synthesis by standard methods. Chemical synthesis produces a single stranded oligonucleotide. This can be converted into double stranded DNA by hybridization with a
complementary sequence or by polymerization with a DNA polymerase using the single strand as a template. Exemplary nucleic acids can be prepared by cloning techniques. Examples of appropriate cloning and sequencing techniques can be found, for example, in Green and Sambrook (Molecular Cloning: A Laboratory Manual, 4th ed., New York: Cold Spring Harbor Laboratory Press, 2012) and Ausubel et al. (Eds.) (Current Protocols in Molecular Biology, New York: John Wiley and Sons, including supplements). Nucleic acids can also be prepared by amplification methods. Amplification methods include the polymerase chain reaction (PCR), the ligase chain reaction (LCR), the transcription-based amplification system (TAS), and the self-sustained sequence replication system (3SR). The nucleic acid molecules can be expressed in a recombinantly engineered cell such as bacteria, plant, yeast, insect and mammalian cells. The antibodies, antigen binding fragments, and conjugates can be expressed as individual proteins including the VH and/or VL (linked to an effector molecule or detectable marker as needed), or can be expressed as a fusion protein. Any suitable method of expressing and purifying antibodies and antigen binding fragments may be used; non-limiting examples are provided in Al- Rubeai (Ed.), Antibody Expression and Production, Dordrecht; New York: Springer, 2011). An immunoadhesin can also be expressed. Thus, in some examples, nucleic acids encoding a VH and VL, and immunoadhesin are provided. The nucleic acid sequences can optionally encode a leader sequence. To create a scFv the VH- and VL-encoding DNA fragments can be operatively linked to another fragment encoding a flexible linker, e.g., encoding the amino acid sequence (Gly4-Ser)3, such that the VH and VL sequences can be expressed as a contiguous single-chain protein, with the VL and VH domains joined by the flexible linker (see, e.g., Bird et al., Science, 242(4877):423-426, 1988; Huston et al., Proc. Natl. Acad. Sci. U.S.A., 85(16):5879-5883, 1988; McCafferty et al., Nature, 348:552-554, 1990; Kontermann and Dübel (Eds.), Antibody Engineering, Vols.1-2, 2nd ed., Springer-Verlag, 2010; Greenfield (Ed.), Antibodies: A Laboratory Manual, 2nd ed. New York: Cold Spring Harbor Laboratory Press, 2014). Optionally, a cleavage site can be included in a linker, such as a furin cleavage site. The single chain antibody may be monovalent, if only a single VH and VL are used, bivalent, if two VH and VL are used, or polyvalent, if more than two VH and VL are used. Bispecific or polyvalent antibodies may be generated that bind specifically to PfCSP and another antigen. The encoded VH and VL optionally can include a furin cleavage site between the VH and VL domains. One or more DNA sequences encoding the antibodies, antigen binding fragments, or conjugates can be expressed in vitro by DNA transfer into a suitable host cell. The cell may be prokaryotic or eukaryotic. Numerous expression systems available for expression of proteins including E. coli, other bacterial hosts, yeast, and various higher eukaryotic cells such as the COS, CHO, HeLa and myeloma cell lines, can be used to express the disclosed antibodies and antigen binding fragments. Methods of stable transfer, meaning that the foreign DNA is continuously maintained in the host may be used. Hybridomas expressing the antibodies of interest are also encompassed by this disclosure.
The expression of nucleic acids encoding the antibodies and antigen binding fragments described herein can be achieved by operably linking the DNA or cDNA to a promoter (which is either constitutive or inducible), followed by incorporation into an expression cassette. The promoter can be any promoter of interest, including a cytomegalovirus promoter. Optionally, an enhancer, such as a cytomegalovirus enhancer, is included in the construct. The cassettes can be suitable for replication and integration in either prokaryotes or eukaryotes. Typical expression cassettes contain specific sequences useful for regulation of the expression of the DNA encoding the protein. For example, the expression cassettes can include appropriate promoters, enhancers, transcription and translation terminators, initiation sequences, a start codon (i.e., ATG) in front of a protein-encoding gene, splicing signals for introns, sequences for the maintenance of the correct reading frame of that gene to permit proper translation of mRNA, and stop codons. The vector can encode a selectable marker, such as a marker encoding drug resistance (for example, ampicillin or tetracycline resistance). To obtain high level expression of a cloned gene, it is desirable to construct expression cassettes which contain, for example, a strong promoter to direct transcription, a ribosome binding site for translational initiation (e.g., internal ribosomal binding sequences), and a transcription/translation terminator. For E. coli, this can include a promoter such as the T7, trp, lac, or lambda promoters, a ribosome binding site, and preferably a transcription termination signal. For eukaryotic cells, the control sequences can include a promoter and/or an enhancer derived from, for example, an immunoglobulin gene, HTLV, SV40 or cytomegalovirus, and a polyadenylation sequence, and can further include splice donor and/or acceptor sequences (for example, CMV and/or HTLV splice acceptor and donor sequences). The cassettes can be transferred into the chosen host cell by any suitable method such as transformation or electroporation for E. coli and calcium phosphate treatment, electroporation or lipofection for mammalian cells. Cells transformed by the cassettes can be selected by resistance to antibiotics conferred by genes contained in the cassettes, such as the amp, gpt, neo and hyg genes. Modifications can be made to a nucleic acid encoding a polypeptide described herein without diminishing its biological activity. Some modifications can be made to facilitate the cloning, expression, or incorporation of the targeting molecule into a fusion protein. Such modifications include, for example, termination codons, sequences to create conveniently located restriction sites, and sequences to add a methionine at the amino terminus to provide an initiation site, or additional amino acids (such as poly His) to aid in purification steps. Once expressed, the antibodies, antigen binding fragments, and conjugates can be purified according to standard procedures in the art, including ammonium sulfate precipitation, affinity columns, column chromatography, and the like (see, generally, Simpson et al. (Eds.), Basic methods in Protein Purification and Analysis: A Laboratory Manual, New York: Cold Spring Harbor Laboratory Press, 2009). The antibodies, antigen binding fragment, and conjugates need not be 100% pure. Once purified, partially or to homogeneity as desired, if to be used prophylatically, the polypeptides should be substantially free of endotoxin.
Methods for expression of antibodies, antigen binding fragments, and conjugates, and/or refolding to an appropriate active form, from mammalian cells, and bacteria such as E. coli have been described and are applicable to the antibodies disclosed herein. See, e.g., Greenfield (Ed.), Antibodies: A Laboratory Manual, 2nd ed. New York: Cold Spring Harbor Laboratory Press, 2014, Simpson et al. (Eds.), Basic methods in Protein Purification and Analysis: A Laboratory Manual, New York: Cold Spring Harbor Laboratory Press, 2009, and Ward et al., Nature 341(6242):544-546, 1989. D. Methods and Compositions 1. Inhibiting P. falciparum infection Methods are disclosed herein for the inhibition of a P. falciparum infection in a subject. The methods include administering to the subject an effective amount (that is, an amount effective to inhibit P. falciparum infection in the subject) of a disclosed antibody (such as any one of MAD21-17, MAD21-46, MAD21-53, MAD21-95, MAD21-101, MAD22-17, MAD22-38, MAD22-39, MAD24-01, MAD24-05, or MAD24-52), antigen binding fragment, conjugate, or a nucleic acid encoding such an antibody, antigen binding fragment, or conjugate, to a subject at risk of a P. falciparum infection. The methods can be used pre-exposure or post-exposure. P. falciparum infection does not need to be completely eliminated or inhibited for the method to be effective. For example, the method can decrease P. falciparum infection by a desired amount, for example by at least 10%, at least 20%, at least 50%, at least 60%, at least 70%, at least 80%, at least 90%, at least 95%, at least 98%, or even at least 100% (elimination or prevention of detectable P. falciparum infection) as compared to P. falciparum infection in the absence of the treatment. In some aspects, the subject can also be treated with an effective amount of an additional agent, such as anti-malaria agent. In some aspects, administration of an effective amount of a disclosed antibody, antigen binding fragment, conjugate, or nucleic acid molecule, inhibits the establishment of P. falciparum infection and/or subsequent P. falciparum disease progression in a subject, which can encompass any statistically significant reduction in P. falciparum activity (for example, growth or invasion) or symptoms of P. falciparum infection in the subject. Antibodies and antigen binding fragments thereof are typically administered by intravenous infusion. Doses of the antibody or antigen binding fragment vary, but generally range between about 0.5 mg/kg to about 50 mg/kg, such as a dose of about 1 mg/kg, about 5 mg/kg, about 10 mg/kg, about 20 mg/kg, about 30 mg/kg, about 40 mg/kg, or about 50 mg/kg. In some aspects, the dose of the antibody or antigen binding fragment can be from about 0.5 mg/kg to about 5 mg/kg, such as a dose of about 1 mg/kg, about 2 mg/kg, about 3 mg/kg, about 4 mg/kg or about 5 mg/kg. The antibody or antigen binding fragment is administered according to a dosing schedule determined by a medical practitioner. In some examples, the antibody or antigen binding fragment is administered weekly, every two weeks, every three weeks or every four weeks.
In some aspects, the method of inhibiting P. falciparum infection in a subject further comprises administration of one or more additional agents to the subject. Additional agents of interest include, but are not limited to, anti-malaria agents. In some aspects, the method of inhibiting P. falciparum infection in a subject comprises administration of a first antibody that specifically binds to PfCSP as disclosed herein (such as any one MAD21-17, MAD21-46, MAD21-53, MAD21-95, MAD21-101, MAD22-17, MAD22-38, MAD22-39, MAD24-01, MAD24-05, or MAD24-52) and a second antibody that that specifically binds to PfCSP (such as L9 or CIS43). In some aspects, a subject is administered DNA or RNA encoding a disclosed antibody to provide in vivo antibody production, for example using the cellular machinery of the subject. Any suitable method of nucleic acid administration may be used; non-limiting examples are provided in U.S. Patent No.5,643,578, U.S. Patent No.5,593,972 and U.S. Patent No.5,817,637. U.S. Patent No.5,880,103 describes several methods of delivery of nucleic acids encoding proteins to an organism. One approach to administration of nucleic acids is direct administration with plasmid DNA, such as with a mammalian expression plasmid. The nucleotide sequence encoding the disclosed antibody, or antigen binding fragments thereof, can be placed under the control of a promoter to increase expression. The methods include liposomal delivery of the nucleic acids. Such methods can be applied to the production of an antibody, or antigen binding fragments thereof. In some aspects, a disclosed antibody or antigen binding fragment is expressed in a subject using the pVRC8400 vector (described in Barouch et al., J. Virol., 79(14), 8828-8834, 2005). In several aspects, a subject (such as a human subject at risk of P. falciparum infection) can be administered an effective amount of an AAV viral vector that comprises one or more nucleic acid molecules encoding a disclosed antibody or antigen binding fragment. The AAV viral vector is designed for expression of the nucleic acid molecules encoding a disclosed antibody or antigen binding fragment, and administration of the effective amount of the AAV viral vector to the subject leads to expression of an effective amount of the antibody or antigen binding fragment in the subject. Non-limiting examples of AAV viral vectors that can be used to express a disclosed antibody or antigen binding fragment in a subject include those provided in Johnson et al., Nat. Med., 15(8):901-906, 2009 and Gardner et al., Nature, 519(7541):87-91, 2015. In one aspect, a nucleic acid encoding a disclosed antibody, or antigen binding fragment thereof, is introduced directly into tissue. For example, the nucleic acid can be loaded onto gold microspheres by standard methods and introduced into the skin by a device such as Bio-Rad’s HELIOS ^ Gene Gun. The nucleic acids can be “naked,” consisting of plasmids under control of a strong promoter. Typically, the DNA is injected into muscle, although it can also be injected directly into other sites. Dosages for injection are usually around 0.5 µg/kg to about 50 mg/kg, and typically are about 0.005 mg/kg to about 5 mg/kg (see, e.g., U.S. Patent No.5,589,466). Single or multiple administrations of a composition including a disclosed PfCSP-specific antibody, antigen binding fragment, conjugate, or nucleic acid molecule encoding such molecules, can be administered
depending on the dosage and frequency as required and tolerated by the patient. The dosage can be administered once, but may be applied periodically until either a desired result is achieved or until side effects warrant discontinuation of therapy. Generally, the dose is sufficient to inhibit P. falciparum infection without producing unacceptable toxicity to the patient. Data obtained from cell culture assays and animal studies can be used to formulate a range of dosage for use in humans. The dosage normally lies within a range of circulating concentrations that include the ED50, with little or minimal toxicity. The dosage can vary within this range depending upon the dosage form employed and the route of administration utilized. The effective dose can be determined from cell culture assays and animal studies. The PfCSP-specific antibody, antigen binding fragment, conjugate, or nucleic acid molecule encoding such molecules, or a composition including such molecules, can be administered to subjects in various ways, including local and systemic administration, such as, e.g., by injection subcutaneously, intravenously, intra-arterially, intraperitoneally, intramuscularly, intradermally, or intrathecally. In an aspect, the antibody, antigen binding fragment, conjugate, or nucleic acid molecule encoding such molecules, or a composition including such molecules, is administered by a single subcutaneous, intravenous, intra-arterial, intraperitoneal, intramuscular, intradermal or intrathecal injection once a day. The antibody, antigen binding fragment, conjugate, or nucleic acid molecule encoding such molecules, or a composition including such molecules, can also be administered by direct injection at or near the site of disease. A further method of administration is by osmotic pump (e.g., an Alzet pump) or mini-pump (e.g., an Alzet mini-osmotic pump), which allows for controlled, continuous and/or slow-release delivery of the antibody, antigen binding fragment, conjugate, or nucleic acid molecule encoding such molecules, or a composition including such molecules, over a pre-determined period. The osmotic pump or mini-pump can be implanted subcutaneously, or near a target site. 2. Compositions Compositions are provided that include one or more of the PfCSP-specific antibody, antigen binding fragment, conjugate, or nucleic acid molecule encoding such molecules, that are disclosed herein in a pharmaceutically acceptable carrier. In some aspects, the composition comprises an antibody as provided herein (such as any one of MAD21-17, MAD21-46, MAD21-53, MAD21-95, MAD21-101, MAD22-17, MAD22-38, MAD22-39, MAD24-01, MAD24-05, or MAD24-52). In some aspects, the composition comprises an antibody as provided herein (such as any one of MAD21-17, MAD21-46, MAD21-53, MAD21-95, MAD21-101, MAD22-17, MAD22-38, MAD22-39, MAD24-01, MAD24-05, or MAD24-52) and one or more additional PfCSP-specific antibody, such as L9 or CIS43 or 317. The compositions are useful, for example, for example, for the inhibition or detection of a P. falciparum infection. The compositions can be prepared in unit dosage forms for administration to a subject. The amount and timing of administration are at the discretion of the administering physician to achieve the desired purposes. The PfCSP-specific antibody, antigen binding fragment, conjugate, or nucleic acid molecule encoding such
molecules can be formulated for systemic or local administration. In one example, the PfCSP-specific antibody, antigen binding fragment, conjugate, or nucleic acid molecule encoding such molecules, is formulated for parenteral administration, such as intravenous administration. In some aspects, the antibody, antigen binding fragment, or conjugate thereof, in the composition is at least 70% (such as at least 75%, at least 80%, at least 85%, at least 90%, at least 95%, at least 96%, at least 97%, at least 98% or at least 99%) pure. In some aspects, the composition contains less than 10% (such as less than 5%, less than 4%, less than 3%, less than 2%, less than 1%, less than 0.5%, or even less) of macromolecular contaminants, such as other mammalian (e.g., human) proteins. The compositions for administration can include a solution of the PfCSP-specific antibody, antigen binding fragment, conjugate, or nucleic acid molecule encoding such molecules, dissolved in a pharmaceutically acceptable carrier, such as an aqueous carrier. A variety of aqueous carriers can be used, for example, buffered saline and the like. These solutions are sterile and generally free of undesirable matter. These compositions may be sterilized by any suitable technique. The compositions may contain pharmaceutically acceptable auxiliary substances as required to approximate physiological conditions such as pH adjusting and buffering agents, toxicity adjusting agents and the like, for example, sodium acetate, sodium chloride, potassium chloride, calcium chloride, sodium lactate and the like. The concentration of antibody in these formulations can vary widely, and will be selected primarily based on fluid volumes, viscosities, body weight and the like in accordance with the particular mode of administration selected and the subject’s needs. A typical composition for intravenous administration comprises about 0.01 to about 30 mg/kg of antibody or antigen binding fragment or conjugate per subject per day (or the corresponding dose of a conjugate including the antibody or antigen binding fragment). Any suitable method may be used for preparing administrable compositions; non-limiting examples are provided in such publications as Remington: The Science and Practice of Pharmacy, 22nd ed., London, UK: Pharmaceutical Press, 2013. In some aspects, the composition can be a liquid formulation including one or more antibodies, antigen binding fragments (such as an antibody or antigen binding fragment that specifically binds to PfCSP), in a concentration range from about 0.1 mg/ml to about 20 mg/ml, or from about 0.5 mg/ml to about 20 mg/ml, or from about 1 mg/ml to about 20 mg/ml, or from about 0.1 mg/ml to about 10 mg/ml, or from about 0.5 mg/ml to about 10 mg/ml, or from about 1 mg/ml to about 10 mg/ml. Antibodies, or an antigen binding fragment thereof or a conjugate or a nucleic acid encoding such molecules, can be provided in lyophilized form and rehydrated with sterile water before administration, although they are also provided in sterile solutions of known concentration. The antibody solution, or an antigen binding fragment or a nucleic acid encoding such antibodies or antigen binding fragments, can then be added to an infusion bag containing 0.9% sodium chloride, USP, and typically administered at a dosage of from 0.5 to 15 mg/kg of body weight. Considerable experience is available in the art in the administration of antibody drugs, which have been marketed in the U.S. since the approval of Rituximab in 1997. Antibodies, antigen binding fragments, conjugates, or a nucleic acid encoding such molecules, can be
administered by slow infusion, rather than in an intravenous push or bolus. In one example, a higher loading dose is administered, with subsequent, maintenance doses being administered at a lower level. For example, an initial loading dose of 4 mg/kg may be infused over a period of some 90 minutes, followed by weekly maintenance doses for 4-8 weeks of 2 mg/kg infused over a 30-minute period if the previous dose was well tolerated. Controlled-release parenteral formulations can be made as implants, oily injections, or as particulate systems. For a broad overview of protein delivery systems see, Banga, Therapeutic Peptides and Proteins: Formulation, Processing, and Delivery Systems, Lancaster, PA: Technomic Publishing Company, Inc., 1995. Particulate systems include microspheres, microparticles, microcapsules, nanocapsules, nanospheres, and nanoparticles. Microcapsules contain the active protein agent, such as a cytotoxin or a drug, as a central core. In microspheres, the active protein agent is dispersed throughout the particle. Particles, microspheres, and microcapsules smaller than about 1 µm are generally referred to as nanoparticles, nanospheres, and nanocapsules, respectively. Capillaries have a diameter of approximately 5 µm so that only nanoparticles are administered intravenously. Microparticles are typically around 100 µm in diameter and are administered subcutaneously or intramuscularly. See, for example, Kreuter, Colloidal Drug Delivery Systems, J. Kreuter (Ed.), New York, NY: Marcel Dekker, Inc., pp.219-342, 1994; and Tice and Tabibi, Treatise on Controlled Drug Delivery: Fundamentals, Optimization, Applications, A. Kydonieus (Ed.), New York, NY: Marcel Dekker, Inc., pp.315-339, 1992. Polymers can be used for ion-controlled release of the antibody compositions disclosed herein. Any suitable polymer may be used, such as a degradable or nondegradable polymeric matrix designed for use in controlled drug delivery. Alternatively, hydroxyapatite has been used as a microcarrier for controlled release of proteins. In yet another aspect, liposomes are used for controlled release as well as drug targeting of the lipid-capsulated drug. 3. Methods of detection and diagnosis Methods are also provided for the detection of the presence of PfCSP in vitro or in vivo. In one example, the presence of PfCSP is detected in a biological sample from a subject, and can be used to identify a subject with P. falciparum infection. The sample can be any sample, including, but not limited to, tissue from biopsies, autopsies and pathology specimens. Biological samples also include sections of tissues, for example, frozen sections taken for histological purposes. Biological samples further include body fluids, such as blood, serum, plasma, sputum, spinal fluid or urine. The method of detection can include contacting a cell or sample, with an antibody or antigen binding fragment that specifically binds to PfCSP, or conjugate thereof (e.g., a conjugate including a detectable marker) under conditions sufficient to form an immune complex, and detecting the immune complex (e.g., by detecting a detectable marker conjugated to the antibody or antigen binding fragment. In one aspect, the antibody or antigen binding fragment is directly labeled with a detectable marker. In another aspect, the antibody that binds P. falciparum (the primary antibody) is unlabeled and a secondary
antibody or other molecule that can bind the primary antibody is utilized for detection. The secondary antibody is chosen that is able to specifically bind the specific species and class of the first antibody. For example, if the first antibody is a human IgG, then the secondary antibody may be an anti-human-IgG. Other molecules that can bind to antibodies include, without limitation, Protein A and Protein G, both of which are available commercially. Suitable labels for the antibody, antigen binding fragment or secondary antibody are known and described above, and include various enzymes, prosthetic groups, fluorescent materials, luminescent materials, magnetic agents and radioactive materials. In some aspects, the disclosed antibodies or antigen binding fragments thereof are used to test vaccines. For example to test if a vaccine composition including a PfCSP or fragment thereof assumes a conformation including the epitope of a disclosed antibody. Thus provided herein is a method for testing a vaccine, wherein the method comprises contacting a sample containing the vaccine, such as a PfCSP immunogen, with a disclosed antibody or antigen binding fragment under conditions sufficient for formation of an immune complex, and detecting the immune complex, to detect the vaccine with an PfCSP immunogen including the epitope in the sample. In one example, the detection of the immune complex in the sample indicates that vaccine component, such as a PfCSP immunogen assumes a conformation capable of binding the antibody or antigen binding fragment. III. EXAMPLES The following example is provided to illustrate particular features of certain aspects, but the scope of the claims should not be limited to those features exemplified. Example 1 Highly Protective Anti-Malarial Antibodies This example illustrates the design and assessment of antibodies to PfCSP with protective potency and that bind to a novel PfCSP epitope. Identification of Pf sporozoite specific mAbs. An antigen-agnostic approach was used to isolate functional monoclonal antibodies (mAbs) that target novel epitopes on the Plasmodium falciparum (Pf) sporozoite. To down-select donors who were likely to produce Abs that targeted non-canonical epitopes on the surface of Pf sporozoites, we screened plasma samples from a cohort of US vaccinees who had received doses of radiation-attenuated sporozoites (PfSPZ Vaccine, Sanaria). Plasma samples were incubated with excess recombinant PfCSP to block Abs which target canonical PfCSP epitopes. Blocking of these antibodies was confirmed after negligible detection of plasma IgG reactivity to beads conjugated to recombinant PfCSP, (NANP)9 peptide, N-terminus or C-terminus in a multiplex assay. Plasma IgG reactivity towards whole Pf sporozoites was assessed in parallel for both the blocked and unblocked plasma samples and is represented as the percentage of IgG positive sporozoites. In total, 4 vaccinees showed high IgG reactivity to sporozoites after blocking of plasma (FIG.1). These individuals were therefore selected as
donors of interest for the study. Binding of plasma IgG to sporozoites or PfCSP antigens was measured with the IntelliCyt iQue Screener flow cytometer and FACS data were analyzed with FlowJo (Version 10.8.1. Ashland, OR). Using the Beacon optofluidic platform (Berkeley Lights) we assessed the antigen specificity of memory B cells (MBCs) isolated from the four donors of interest and exported MBCs that secreted mAbs which bound to the surface of Pf sporozoites but not to recombinant, mammalian cell expressed PfCSP. The respective antibodies were recombinantly expressed as IgG1 mAbs and titrated against wild-type Pf sporozoites. Binding of recombinant mAbs to sporozoites was measured with the IntelliCyt iQue Screener flow cytometer and FACS data were analysed with FlowJo (Version 10.8.1. Ashland, OR). In total, it was confirmed that 11 of the isolated mAbs bound to the surface of Pf sporozoites. The names of these mAbs, their type, and the corresponding VH and VL sequences are as follows: Antibody VH VL VH DNA VL DNA name SEQ ID NO SEQ ID NO SEQ ID NO SEQ ID NO The i
dentified mAbs bind to sporozoites that express Pf CSP. To further interrogate antigen specificity, the mAbs were titrated against wild-type Plasmodium berghei (Pb) sporozoites and transgenic Pb sporozoites that express PfCSP (PbPfCSP) (FIG.2). All 11 of the mAbs bound to PbPfCSP sporozoites but none of the mAbs showed appreciable binding to wild-type Pb sporozoites, suggesting that PfCSP is a likely target for these mAbs. The identified mAbs recognize an ~60kDa sporozoite expressed protein. To further assess the binding specificity of the identified mAbs, proteins in sporozoite lysate were resolved and subsequent western blots were developed using the representative mAbs MAD21-46, MAD22-38 and MAD21-101, and known PfCSP mAbs CIS43 and mAb10 as probes (FIG.3). Both the western blotting analysis with the anti- CSP antibodies (CIS43 and mAb10) and the representative mAbs (MAD21-46, MAD22-38 and MAD21- 101) detected a band at ~60kDa, suggesting the mAbs likely bind to the same target as CIS43 and mAb10. The identified mAbs do not bind canonical PfCSP epitopes. To further assess the binding specificity of the identified mAbs, the 11 mAbs were titrated against recombinant Pf CSP, Pf (NANP)9
peptide, Pf C-terminus and Pf N-terminus (FIG.4). Binding of recombinant mAbs to the antigen conjugated beads was measured by flow cytometry in a multiplexed bead-based assay as previously described. Overall, none of the mAbs showed strong binding to any of the antigens tested. Taken together with the data in FIGS.2 and 3, these results suggest that the mAbs target a novel, non-canonical PfCSP epitope. Pf CSP peptide scanning analysis. To determine whether an unexplored site within Pf CSP was targeted by the mAbs, two representative mAbs (MAD22-38 and MAD21-101) were assessed using ELISA- based peptide mapping using an array of 69 overlapping peptides (amino acid sequences detailed below) which spanned the length of the entire PfCSP (3D7 strain) (FIG.5). Neither of the mAbs appreciably bound any of the peptides tested. Peptide name Peptide sequence SEQ ID NO
PfCSP45 NVDPNANPNANPNAN 129 PfCSP61 NANPNANPNANPNKN 130 PfCSP62 NANPNANPNKNN GN 131
e a ove ana yses suggest t at t e ent e m s o not target canonca ep topes o previously identified neutralizing Pf mAbs, including CIS43 (junctional tetrapeptide), L9 (NVDP minor repeats) and 317 (NANP major repeats). The identified mAbs do not bind recombinant forms of Pf CSP. To further assess the PfCSP binding of the identified mAbs, the binding of two representative mAbs (MAD22-38 and MAD21-101) to a diverse panel of PfCSP constructs was measured by ELISA (FIG.6). These recombinant forms of PfCSP were produced in HEK293, E. coli (Kastenmuller et al., 2013), P. pastoris (Kastenmuller et al., Infect Immun 81, 789-800.10.1128/IAI.01108-12, 2013) and L. lactis (Singh et al., J Biol Chem 295, 403-414. 10.1074/jbc.RA119.011268, 2020) expression systems. Neither of the representative mAbs bound any of the constructs tested. These results suggest that mAb binding favors the native, sporozoite-expressed form of the unknown epitope. The identified mAbs show protection in an in vivo model of P. falciparum liver invasion. The 11 mAbs were passively transferred to mice and tested for their ability to reduce parasite liver burden after intravenous (IV) challenge with transgenic PbPfCSP sporozoites that express a green fluorescent protein/luciferase fusion protein (Raghunandan et al., Malar J 19, 113.10.1186/s12936-020-03181-0, 2020) (FIGS.7A-7D). Initial screening in the model highlighted that 6 mAbs (MAD22-38, MAD22-39, MAD21- 46, MAD21-53, MAD21-95 and MAD21-101) reduced liver burden when compared to the max burden
condition (FIG.7A). Further challenge experiments (FIGS.7B-D) show that a subset of these mAbs (MAD21-46, MAD21-101 and MAD22-38) may cooperatively improve protection when delivered in combination with known neutralizing PfCSP mAbs CIS43 (Kisalu et al., Nat Med 24, 408-416. 10.1038/nm.4512, 2018), L9 (Wang et al., Immunity 53, 733-744 e738.10.1016/j.immuni.2020.08.014, 2020) or 317 (Oyen et al., Proc Natl Acad Sci U S A 114, E10438-E10445.10.1073/pnas.1715812114, 2017). The identified mAbs recognize an epitope within the Pf-CSP junctional region and do not show cross-reactivity to the major repeat (NANP) sequences. To delineate the domain of PfCSP in which the mAbs’ epitope was located, three representative mAbs (MAD21-46, MAD21-101 and MAD22-38) were assessed by ELISA for binding to a panel of chimeric sporozoites. The panel consisted of transgenic Pb sporozoites that express the full PfCSP sequence (PbPfCSP); transgenic Pb sporozoites that express PfCSP which lacks the ADGNPDP residues in the junctional region (Pb-PfCSP JRKO); chimeric Pb sporozoites with 12 Pf-NANP repeats inserted into the PbCSP open reading frame (Pb CSPPf-NANP 12); and chimeric Pb sporozoites with 4 Pf-NANP repeats and the ADGNPDP residues from the Pf junctional region inserted into the PbCSP open reading frame (Pb CSPPf-NANP 4-5). While the representative mAbs do not bind to wild-type Pb parasites, the ELISA analysis confirmed their binding to the PbPfCSP parasites (FIGS.8A-8B). However, mAb binding was abrogated in parasites which lack the junctional region ADGNPDP residues (Pb-PfCSP JRKO). While the mAbs were not reactive to the Pf-NANP epitope expressed in Pb CSPPf-NANP 12 parasites, binding ability was restored for the Pb CSPPf-NANP 4-5 parasites which express the junctional region ADGNPDP residues in addition to the Pf-NANP epitope. Together, these findings evidence that the mAb epitope lies within the junction region and that the disclosed mAbs do not cross react with Pf-NANP repeats (unlike the monoclonal antibody CIS43). mAbs recognize a proteolytically processed form of Pf-CSP. To interrogate whether post- translational processing of PfCSP was involved in generating the mAb epitope, proteins in sporozoite lysate were resolved and subsequent western blots were developed using representative mAbs (MAD22-39, MAD22-38, MAD24-52, MAD24-01 and MAD21-101), and known anti-PfCSP control mAbs mAb10 and 5D5 (Espinosa et al., J Infect Dis 212, 1111-1119.10.1093/infdis/jiv154, 2015) as probes (FIGS.9A-9B). Western blotting analysis with mAb10 detected both the cleaved (lower molecular weight) and un-cleaved (higher molecular weight) forms of PfCSP while the mAb 5D5 detected only the un-cleaved form of PfCSP. The MW of the band detected by the representative mAbs corresponds to that of the cleaved PfCSP, confirming that the representative mAbs target a proteolytically processed form of PfCSP. mAbs preferentially bind Pf-CSP peptides with a truncated N-terminus and pyroglutamic acid modification. To further interrogate the post-translational modifications involved in generating the epitope, the eleven mAbs were titrated against recombinant peptides that recapitulate sporozoite-mediated, post- translational modifications present on PfCSP. Binding of recombinant mAbs to the antigen conjugated beads was measured by flow cytometry in a multiplexed bead-based assay. All mAbs bound to the peptide bearing an N-terminal glutamine which represents the proposed PfCSP N-terminus post-cleavage (FIG.10). This
binding was increased towards the peptide bearing an N-terminal pyroglutamic acid which represents an additional post-translational modification proposed to occur downstream of PfCSP cleavage (Kolli et al., Proc Natl Acad Sci U S A 119, e2209729119.10.1073/pnas.2209729119, 2022). Taken together, these results evidence that the presently disclosed mAbs target a novel epitope on PfCSP which is dependent upon sporozoite-mediated cleavage and pyroglutamic acid modification of PfCSP. It will be apparent that the precise details described may be varied or modified without departing from the spirit of the described invention. We claim all such modifications and variations that fall within the scope and spirit of the claims below.
Claims
It is claimed: 1. A monoclonal antibody, comprising a) a heavy chain variable region (VH) and a light chain variable region (VL) comprising a heavy chain complementarity determining region (HCDR)1, a HCDR2, and a HCDR3, and a light chain complementarity determining region (LCDR)1, a LCDR2, and a LCDR3 of the VH and VL set forth as SEQ ID NOs: 9 and 10, respectively (MAD21-101); b) a VH and VL comprising a HCDR1, a HCDR2, and a HCDR3, and a LCDR1, a LCDR2, and a LCDR3 of the VH and VL set forth as SEQ ID NOs: 3 and 4, respectively (MAD21-46); c) a VH and VL comprising a HCDR1, a HCDR2, and a HCDR3, and a LCDR1, a LCDR2, and a LCDR3 of the VH and VL set forth as SEQ ID NOs: 13 and 14, respectively (MAD22-38); d) a VH and VL comprising a HCDR1, a HCDR2, and a HCDR3, and a LCDR1, a LCDR2, and a LCDR3 of the VH and VL set forth as SEQ ID NOs: 7 and 8, respectively (MAD21-95); e) a VH and VL comprising a HCDR1, a HCDR2, and a HCDR3, and a LCDR1, a LCDR2, and a LCDR3 of the VH and VL set forth as SEQ ID NOs: 15 and 16, respectively (MAD22-39); f) a VH and VL comprising a HCDR1, a HCDR2, and a HCDR3, and a LCDR1, a LCDR2, and a LCDR3 of the VH and VL set forth as SEQ ID NOs: 11 and 12, respectively (MAD22-17); g) a VH and VL comprising a HCDR1, a HCDR2, and a HCDR3, and a LCDR1, a LCDR2, and a LCDR3 of the VH and VL set forth as SEQ ID NOs: 5 and 6, respectively (MAD21-53); h) a VH and VL comprising a HCDR1, a HCDR2, and a HCDR3, and a LCDR1, a LCDR2, and a LCDR3 of the VH and VL set forth as SEQ ID NOs: 1 and 2, respectively (MAD21-17); i) a VH and VL comprising a HCDR1, a HCDR2, and a HCDR3, and a LCDR1, a LCDR2, and a LCDR3 of the VH and VL set forth as SEQ ID NOs: 17 and 18, respectively (MAD24-01); j) a VH and VL comprising a HCDR1, a HCDR2, and a HCDR3, and a LCDR1, a LCDR2, and a LCDR3 of the VH and VL set forth as SEQ ID NOs: 19 and 20, respectively (MAD24-05); k) a VH and VL comprising a HCDR1, a HCDR2, and a HCDR3, and a LCDR1, a LCDR2, and a LCDR3 of the VH and VL set forth as SEQ ID NOs: 21 and 22, respectively (MAD24-52); and wherein the monoclonal antibody specifically binds to P. falciparum circumsporozoite protein (PfCSP) and neutralizes P. falciparum.
2. The monoclonal antibody of claim 1, wherein the HCDR1, the HCDR2, the HCDR3, the LCDR1, the LCDR2, and the LCDR3 are set forth as: a) SEQ ID NOs: 29, 24, 33, 26, 27, and 34, respectively (MAD21-101); b) SEQ ID NOs: 29, 24, 25, 26, 27, and 28, respectively (MAD21-46); c) SEQ ID NOs: 40, 41, 42, 43, 44, and 45, respectively (MAD22-38); d) SEQ ID NOs: 23, 24, 25, 31, 27, and 28, respectively (MAD21-95); e) SEQ ID NOs: 46, 47, 48, 49, 44, and 45, respectively (MAD22-39); f) SEQ ID NOs: 35, 36, 37, 38, 27, and 39, respectively (MAD22-17);
g) SEQ ID NOs: 29, 24, 30, 26, 27, and 28, respectively (MAD21-53); h) SEQ ID NOs: 23, 24, 25, 26, 27, and 28, respectively (MAD21-17); i) SEQ ID NOs: 50, 51, 52, 53, 54, and 55, respectively (MAD24-01); j) SEQ ID NOs: 56, 57, 58, 59, 60, and 61, respectively (MAD24-05); or k) SEQ ID NOs: 62, 63, 64, 65, 66, and 67, respectively (MAD24-52).
3. The antibody of any one of the prior claims, wherein the VH and the VL comprise the HCDR1, the HCDR2, the HCDR3, the LCDR1, the LCDR2, and the LCDR3 and the amino acid sequences of the framework regions of the VH and the VL are at least 90% identical to those of: a) SEQ ID NOs: 9 and 10, respectively; b) SEQ ID NOs: 3 and 4, respectively; c) SEQ ID NOs: 13 and 14, respectively; d) SEQ ID NOs: 7 and 8, respectively; e)SEQ ID NOs: 15 and 16, respectively ; f) SEQ ID NOs: 11 and 12, respectively; g) SEQ ID NOs: 5 and 6, respectively; h) SEQ ID NOs: 1 and 2, respectively; i) SEQ ID NOs: 17 and 18, respectively; j) SEQ ID NOs: 19 and 20, respectively; or k) SEQ ID NOs: 21 and 22, respectively.
4. The antibody of any one of the prior claims, wherein the VH and the VL comprise amino acid sequences set forth as: a) SEQ ID NOs: 9 and 10, respectively; b) SEQ ID NOs: 3 and 4, respectively; c) SEQ ID NOs: 13 and 14, respectively; d) SEQ ID NOs: 7 and 8, respectively; e)SEQ ID NOs: 15 and 16, respectively ; f) SEQ ID NOs: 11 and 12, respectively; g) SEQ ID NOs: 5 and 6, respectively; h) SEQ ID NOs: 1 and 2, respectively; i) SEQ ID NOs: 17 and 18, respectively; j) SEQ ID NOs: 19 and 20, respectively; or k) SEQ ID NOs: 21 and 22, respectively.
5. The antibody of any one of the prior claims, wherein the antibody comprises a human constant domain.
6. The antibody of any one of the prior claims, wherein the antibody is a human antibody.
7. The antibody of any one of the prior claims, wherein the antibody is an IgG.
8. The antibody of any one of the prior claims, comprising a recombinant constant domain comprising a modification that increases the half-life of the antibody.
9. The antibody of claim 8, wherein the modification increases binding to the neonatal Fc receptor.
10. The isolated monoclonal antibody of claim 9, wherein the recombinant constant domain is an IgG1 constant domain comprising M428L and N434S mutations.
11. An isolated antigen binding fragment of the antibody of any one of the prior claims, wherein the antigen binding fragment comprises the VH and the VL of the antibody, specifically binds to PfCSP, and neutralizes P. falciparum.
12. The antigen binding fragment of claim 11, wherein the antigen binding fragment is a Fv, Fab, F(ab')2, scFV or a scFV2 fragment.
13. The antibody or antigen binding fragment of any one of the prior claims, conjugated to an effector molecule or a detectable marker.
14. The antibody or antigen binding fragment of any one of the prior claims, wherein the antibody or antigen binding fragment inhibits P. falciparum sporozoite entry into the blood from the skin of the subject and/or inhibits P. falciparum sporozoite entry into hepatocytes in the liver of the subject.
15. A bispecific antibody comprising the antibody or antigen binding fragment of any one of the prior claims.
16. An isolated nucleic acid molecule encoding the antibody or antigen binding fragment of any one of the prior claims.
17. The isolated nucleic acid molecule of claim 16, wherein the nucleic acid molecule is RNA.
18. The nucleic acid molecule of claim 16 or claim 17, operably linked to a promoter.
19. A vector comprising the nucleic acid molecule of any of claims 16-18.
20. A host cell comprising the nucleic acid molecule or vector of any one of claims 16-19.
21. A composition for use in inhibiting P. falciparum infection, comprising an effective amount of the antibody, antigen binding fragment, nucleic acid molecule, or vector, of any one of the prior claims, and a pharmaceutically acceptable carrier.
22. The composition of claim 21, further comprising CIS43 antibody, L9 antibody, or 317 antibody.
23. A method of producing an antibody or antigen binding fragment that specifically binds to PfCSP, comprising: expressing one or more nucleic acid molecules encoding the antibody or antigen binding fragment of any one of claims 1-15 in a host cell; and purifying the antibody or antigen binding fragment.
24. A method of detecting the presence of P. falciparum in a biological sample from a human subject, comprising: contacting the biological sample with an effective amount of the antibody or antigen binding fragment of any one of claims 1-15 under conditions sufficient to form an immune complex; and detecting the presence of the immune complex in the biological sample, wherein the presence of the immune complex in the biological sample indicates the presence of the P. falciparum in the sample.
25. The method of claim 24, wherein detecting the detecting the presence of the immune complex in the biological sample indicates that the subject has a P. falciparum infection.
26. A method of inhibiting a P. falciparum infection in a subject, comprising administering an effective amount of the antibody, antigen binding fragment, nucleic acid molecule, vector, or composition of any one of claims 1-19 or 21-22 to the subject, wherein the subject has or is at risk of a P. falciparum infection.
27. The method of claim 26, wherein the subject is at risk of a P. falciparum infection
28. The method of claim 26 or claim 27, wherein the method inhibits P. falciparum sporozoite entry into the blood from the skin of the subject and/or inhibits P. falciparum sporozoite entry into hepatocytes in the liver of the subject
29. Use of the antibody, antigen binding fragment, nucleic acid molecule, vector, or pharmaceutical composition of any one of claims 1-19 or 21-22, to inhibit P. falciparum infection in a subject or to detect the presence of a P. falciparum in a biological sample.
Applications Claiming Priority (2)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| US202263409016P | 2022-09-22 | 2022-09-22 | |
| US63/409,016 | 2022-09-22 |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| WO2024064826A1 true WO2024064826A1 (en) | 2024-03-28 |
Family
ID=88505260
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/US2023/074791 Ceased WO2024064826A1 (en) | 2022-09-22 | 2023-09-21 | Neutralizing antibodies to plasmodium falciparum circumsporozoite protein and their use |
Country Status (1)
| Country | Link |
|---|---|
| WO (1) | WO2024064826A1 (en) |
Citations (45)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US5589466A (en) | 1989-03-21 | 1996-12-31 | Vical Incorporated | Induction of a protective immune response in a mammal by injecting a DNA sequence |
| US5593972A (en) | 1993-01-26 | 1997-01-14 | The Wistar Institute | Genetic immunization |
| US5643578A (en) | 1992-03-23 | 1997-07-01 | University Of Massachusetts Medical Center | Immunization by inoculation of DNA transcription unit |
| WO1997030087A1 (en) | 1996-02-16 | 1997-08-21 | Glaxo Group Limited | Preparation of glycosylated antibodies |
| WO1998058964A1 (en) | 1997-06-24 | 1998-12-30 | Genentech, Inc. | Methods and compositions for galactosylated glycoproteins |
| US5880103A (en) | 1992-08-11 | 1999-03-09 | President And Fellows Of Harvard College | Immunomodulatory peptides |
| WO1999022764A1 (en) | 1997-10-31 | 1999-05-14 | Genentech, Inc. | Methods and compositions comprising glycoprotein glycoforms |
| WO1999054440A1 (en) | 1998-04-21 | 1999-10-28 | Micromet Gesellschaft Für Biomedizinische Forschung Mbh | CD19xCD3 SPECIFIC POLYPEPTIDES AND USES THEREOF |
| WO2000061739A1 (en) | 1999-04-09 | 2000-10-19 | Kyowa Hakko Kogyo Co., Ltd. | Method for controlling the activity of immunologically functional molecule |
| WO2001029246A1 (en) | 1999-10-19 | 2001-04-26 | Kyowa Hakko Kogyo Co., Ltd. | Process for producing polypeptide |
| WO2002031140A1 (en) | 2000-10-06 | 2002-04-18 | Kyowa Hakko Kogyo Co., Ltd. | Cells producing antibody compositions |
| US20020164328A1 (en) | 2000-10-06 | 2002-11-07 | Toyohide Shinkawa | Process for purifying antibody |
| WO2003011878A2 (en) | 2001-08-03 | 2003-02-13 | Glycart Biotechnology Ag | Antibody glycosylation variants having increased antibody-dependent cellular cytotoxicity |
| US20030115614A1 (en) | 2000-10-06 | 2003-06-19 | Yutaka Kanda | Antibody composition-producing cell |
| US6602684B1 (en) | 1998-04-20 | 2003-08-05 | Glycart Biotechnology Ag | Glycosylation engineering of antibodies for improving antibody-dependent cellular cytotoxicity |
| US20030157108A1 (en) | 2001-10-25 | 2003-08-21 | Genentech, Inc. | Glycoprotein compositions |
| WO2003085119A1 (en) | 2002-04-09 | 2003-10-16 | Kyowa Hakko Kogyo Co., Ltd. | METHOD OF ENHANCING ACTIVITY OF ANTIBODY COMPOSITION OF BINDING TO FcϜ RECEPTOR IIIa |
| WO2003084570A1 (en) | 2002-04-09 | 2003-10-16 | Kyowa Hakko Kogyo Co., Ltd. | DRUG CONTAINING ANTIBODY COMPOSITION APPROPRIATE FOR PATIENT SUFFERING FROM FcϜRIIIa POLYMORPHISM |
| WO2003085107A1 (en) | 2002-04-09 | 2003-10-16 | Kyowa Hakko Kogyo Co., Ltd. | Cells with modified genome |
| US6723538B2 (en) | 1999-03-11 | 2004-04-20 | Micromet Ag | Bispecific antibody and chemokine receptor constructs |
| US20040093621A1 (en) | 2001-12-25 | 2004-05-13 | Kyowa Hakko Kogyo Co., Ltd | Antibody composition which specifically binds to CD20 |
| US20040110282A1 (en) | 2002-04-09 | 2004-06-10 | Kyowa Hakko Kogyo Co., Ltd. | Cells in which activity of the protein involved in transportation of GDP-fucose is reduced or lost |
| US20040109865A1 (en) | 2002-04-09 | 2004-06-10 | Kyowa Hakko Kogyo Co., Ltd. | Antibody composition-containing medicament |
| US20040132140A1 (en) | 2002-04-09 | 2004-07-08 | Kyowa Hakko Kogyo Co., Ltd. | Production process for antibody composition |
| WO2004056312A2 (en) | 2002-12-16 | 2004-07-08 | Genentech, Inc. | Immunoglobulin variants and uses thereof |
| WO2005035778A1 (en) | 2003-10-09 | 2005-04-21 | Kyowa Hakko Kogyo Co., Ltd. | PROCESS FOR PRODUCING ANTIBODY COMPOSITION BY USING RNA INHIBITING THE FUNCTION OF α1,6-FUCOSYLTRANSFERASE |
| WO2005035586A1 (en) | 2003-10-08 | 2005-04-21 | Kyowa Hakko Kogyo Co., Ltd. | Fused protein composition |
| US20050123546A1 (en) | 2003-11-05 | 2005-06-09 | Glycart Biotechnology Ag | Antigen binding molecules with increased Fc receptor binding affinity and effector function |
| WO2005053742A1 (en) | 2003-12-04 | 2005-06-16 | Kyowa Hakko Kogyo Co., Ltd. | Medicine containing antibody composition |
| US7229760B2 (en) | 2000-03-24 | 2007-06-12 | Micromet Ag | mRNA amplification |
| US7235641B2 (en) | 2003-12-22 | 2007-06-26 | Micromet Ag | Bispecific antibodies |
| US7323440B2 (en) | 2002-02-13 | 2008-01-29 | Micromet Ag | De-immunized MOG (poly)peptide constructs |
| US7332168B2 (en) | 2000-08-22 | 2008-02-19 | Micromet Ag | Composition for the elimination of autoreactive B-cells |
| WO2008077546A1 (en) | 2006-12-22 | 2008-07-03 | F. Hoffmann-La Roche Ag | Antibodies against insulin-like growth factor i receptor and uses thereof |
| US7435549B1 (en) | 1997-11-17 | 2008-10-14 | Micromet Ag | Method of identifying binding site domains that retain the capacity of binding to an epitope |
| US7635472B2 (en) | 2003-05-31 | 2009-12-22 | Micromet Ag | Pharmaceutical compositions comprising bispecific anti-cd3, anti-cd19 antibody constructs for the treatment of b-cell related disorders |
| US7820166B2 (en) | 2002-10-11 | 2010-10-26 | Micromet Ag | Potent T cell modulating molecules |
| US7919089B2 (en) | 2003-05-31 | 2011-04-05 | Micromet Ag | Pharmaceutical composition comprising a bispecific antibody for EpCAM |
| US8007796B2 (en) | 2005-12-16 | 2011-08-30 | Micromet Ag | Means and methods for the treatment of tumorous diseases |
| US8017748B2 (en) | 2005-04-18 | 2011-09-13 | Micromet Ag | Antibody neutralizers of human granulocyte macrophage colony stimulating factor |
| US8076459B2 (en) | 2003-10-16 | 2011-12-13 | Micromet Ag | Multispecfic deimmunized CD3-binders |
| WO2013163427A1 (en) | 2012-04-25 | 2013-10-31 | The United States Of America, As Represented By The Secretary, Department Of Health & Human Services | Antibodies to treat hiv-1 infection |
| WO2018148660A1 (en) | 2017-02-10 | 2018-08-16 | The United States Of America, As Represented By The Secretary, Department Of Health And Human Services | Neutralizing antibodies to plasmodium falciparum circumsporozoite protein and their use |
| WO2020227228A2 (en) | 2019-05-03 | 2020-11-12 | The United States Of America, As Represented By The Secretary, Department Of Health And Human Services | Neutralizing antibodies to plasmodium falciparum circumsporozoite protein and their use |
| WO2021257665A1 (en) * | 2020-06-19 | 2021-12-23 | The United States Of America, As Represented By The Secretary, Department Of Health And Human Services | Human monoclonal antibody that targets a conserved site on the plasmodium falciparum circumsporozoite protein |
-
2023
- 2023-09-21 WO PCT/US2023/074791 patent/WO2024064826A1/en not_active Ceased
Patent Citations (49)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US5589466A (en) | 1989-03-21 | 1996-12-31 | Vical Incorporated | Induction of a protective immune response in a mammal by injecting a DNA sequence |
| US5643578A (en) | 1992-03-23 | 1997-07-01 | University Of Massachusetts Medical Center | Immunization by inoculation of DNA transcription unit |
| US5880103A (en) | 1992-08-11 | 1999-03-09 | President And Fellows Of Harvard College | Immunomodulatory peptides |
| US5593972A (en) | 1993-01-26 | 1997-01-14 | The Wistar Institute | Genetic immunization |
| US5817637A (en) | 1993-01-26 | 1998-10-06 | The Trustees Of The University Of Pennsylvania | Genetic immunization |
| WO1997030087A1 (en) | 1996-02-16 | 1997-08-21 | Glaxo Group Limited | Preparation of glycosylated antibodies |
| WO1998058964A1 (en) | 1997-06-24 | 1998-12-30 | Genentech, Inc. | Methods and compositions for galactosylated glycoproteins |
| WO1999022764A1 (en) | 1997-10-31 | 1999-05-14 | Genentech, Inc. | Methods and compositions comprising glycoprotein glycoforms |
| US7435549B1 (en) | 1997-11-17 | 2008-10-14 | Micromet Ag | Method of identifying binding site domains that retain the capacity of binding to an epitope |
| US6602684B1 (en) | 1998-04-20 | 2003-08-05 | Glycart Biotechnology Ag | Glycosylation engineering of antibodies for improving antibody-dependent cellular cytotoxicity |
| WO1999054440A1 (en) | 1998-04-21 | 1999-10-28 | Micromet Gesellschaft Für Biomedizinische Forschung Mbh | CD19xCD3 SPECIFIC POLYPEPTIDES AND USES THEREOF |
| US7575923B2 (en) | 1998-04-21 | 2009-08-18 | Micromet Ag | CD19xCD3 specific polypeptides and uses thereof |
| US7112324B1 (en) | 1998-04-21 | 2006-09-26 | Micromet Ag | CD 19×CD3 specific polypeptides and uses thereof |
| US6723538B2 (en) | 1999-03-11 | 2004-04-20 | Micromet Ag | Bispecific antibody and chemokine receptor constructs |
| WO2000061739A1 (en) | 1999-04-09 | 2000-10-19 | Kyowa Hakko Kogyo Co., Ltd. | Method for controlling the activity of immunologically functional molecule |
| WO2001029246A1 (en) | 1999-10-19 | 2001-04-26 | Kyowa Hakko Kogyo Co., Ltd. | Process for producing polypeptide |
| US7229760B2 (en) | 2000-03-24 | 2007-06-12 | Micromet Ag | mRNA amplification |
| US7332168B2 (en) | 2000-08-22 | 2008-02-19 | Micromet Ag | Composition for the elimination of autoreactive B-cells |
| WO2002031140A1 (en) | 2000-10-06 | 2002-04-18 | Kyowa Hakko Kogyo Co., Ltd. | Cells producing antibody compositions |
| US20020164328A1 (en) | 2000-10-06 | 2002-11-07 | Toyohide Shinkawa | Process for purifying antibody |
| US20030115614A1 (en) | 2000-10-06 | 2003-06-19 | Yutaka Kanda | Antibody composition-producing cell |
| WO2003011878A2 (en) | 2001-08-03 | 2003-02-13 | Glycart Biotechnology Ag | Antibody glycosylation variants having increased antibody-dependent cellular cytotoxicity |
| US20030157108A1 (en) | 2001-10-25 | 2003-08-21 | Genentech, Inc. | Glycoprotein compositions |
| US20040093621A1 (en) | 2001-12-25 | 2004-05-13 | Kyowa Hakko Kogyo Co., Ltd | Antibody composition which specifically binds to CD20 |
| US7323440B2 (en) | 2002-02-13 | 2008-01-29 | Micromet Ag | De-immunized MOG (poly)peptide constructs |
| US20040109865A1 (en) | 2002-04-09 | 2004-06-10 | Kyowa Hakko Kogyo Co., Ltd. | Antibody composition-containing medicament |
| US20040110282A1 (en) | 2002-04-09 | 2004-06-10 | Kyowa Hakko Kogyo Co., Ltd. | Cells in which activity of the protein involved in transportation of GDP-fucose is reduced or lost |
| WO2003085119A1 (en) | 2002-04-09 | 2003-10-16 | Kyowa Hakko Kogyo Co., Ltd. | METHOD OF ENHANCING ACTIVITY OF ANTIBODY COMPOSITION OF BINDING TO FcϜ RECEPTOR IIIa |
| WO2003084570A1 (en) | 2002-04-09 | 2003-10-16 | Kyowa Hakko Kogyo Co., Ltd. | DRUG CONTAINING ANTIBODY COMPOSITION APPROPRIATE FOR PATIENT SUFFERING FROM FcϜRIIIa POLYMORPHISM |
| US20040132140A1 (en) | 2002-04-09 | 2004-07-08 | Kyowa Hakko Kogyo Co., Ltd. | Production process for antibody composition |
| US20040110704A1 (en) | 2002-04-09 | 2004-06-10 | Kyowa Hakko Kogyo Co., Ltd. | Cells of which genome is modified |
| WO2003085107A1 (en) | 2002-04-09 | 2003-10-16 | Kyowa Hakko Kogyo Co., Ltd. | Cells with modified genome |
| US7820166B2 (en) | 2002-10-11 | 2010-10-26 | Micromet Ag | Potent T cell modulating molecules |
| WO2004056312A2 (en) | 2002-12-16 | 2004-07-08 | Genentech, Inc. | Immunoglobulin variants and uses thereof |
| US7919089B2 (en) | 2003-05-31 | 2011-04-05 | Micromet Ag | Pharmaceutical composition comprising a bispecific antibody for EpCAM |
| US7635472B2 (en) | 2003-05-31 | 2009-12-22 | Micromet Ag | Pharmaceutical compositions comprising bispecific anti-cd3, anti-cd19 antibody constructs for the treatment of b-cell related disorders |
| WO2005035586A1 (en) | 2003-10-08 | 2005-04-21 | Kyowa Hakko Kogyo Co., Ltd. | Fused protein composition |
| WO2005035778A1 (en) | 2003-10-09 | 2005-04-21 | Kyowa Hakko Kogyo Co., Ltd. | PROCESS FOR PRODUCING ANTIBODY COMPOSITION BY USING RNA INHIBITING THE FUNCTION OF α1,6-FUCOSYLTRANSFERASE |
| US8076459B2 (en) | 2003-10-16 | 2011-12-13 | Micromet Ag | Multispecfic deimmunized CD3-binders |
| US20050123546A1 (en) | 2003-11-05 | 2005-06-09 | Glycart Biotechnology Ag | Antigen binding molecules with increased Fc receptor binding affinity and effector function |
| WO2005053742A1 (en) | 2003-12-04 | 2005-06-16 | Kyowa Hakko Kogyo Co., Ltd. | Medicine containing antibody composition |
| US7235641B2 (en) | 2003-12-22 | 2007-06-26 | Micromet Ag | Bispecific antibodies |
| US8017748B2 (en) | 2005-04-18 | 2011-09-13 | Micromet Ag | Antibody neutralizers of human granulocyte macrophage colony stimulating factor |
| US8007796B2 (en) | 2005-12-16 | 2011-08-30 | Micromet Ag | Means and methods for the treatment of tumorous diseases |
| WO2008077546A1 (en) | 2006-12-22 | 2008-07-03 | F. Hoffmann-La Roche Ag | Antibodies against insulin-like growth factor i receptor and uses thereof |
| WO2013163427A1 (en) | 2012-04-25 | 2013-10-31 | The United States Of America, As Represented By The Secretary, Department Of Health & Human Services | Antibodies to treat hiv-1 infection |
| WO2018148660A1 (en) | 2017-02-10 | 2018-08-16 | The United States Of America, As Represented By The Secretary, Department Of Health And Human Services | Neutralizing antibodies to plasmodium falciparum circumsporozoite protein and their use |
| WO2020227228A2 (en) | 2019-05-03 | 2020-11-12 | The United States Of America, As Represented By The Secretary, Department Of Health And Human Services | Neutralizing antibodies to plasmodium falciparum circumsporozoite protein and their use |
| WO2021257665A1 (en) * | 2020-06-19 | 2021-12-23 | The United States Of America, As Represented By The Secretary, Department Of Health And Human Services | Human monoclonal antibody that targets a conserved site on the plasmodium falciparum circumsporozoite protein |
Non-Patent Citations (57)
| Title |
|---|
| "Antibody Expre sion and Production, Dordrecht", 2011, SPRINGER |
| AHMAD ET AL., CLIN. DEV. IMMUNOL., 2012 |
| AL-LAZIKANI ET AL.: "Standard conformations for the canonical structures of immunoglobulins", J. MOL. BIO., vol. 273, no. 4, 1997, pages 927 - 948, XP004461383, DOI: 10.1006/jmbi.1997.1354 |
| ALTSCHUL ET AL., J. MOL. BIOL., vol. 215, no. 3, 1990, pages 403 - 410 |
| BANGA: "Therapeutic Peptides and Proteins: Formulation, Processing, and Deliver-v Systems", 1995, TECHNOMIC PUBLISHING COMPANY, INC. |
| BAROUCH ET AL., J. VIROL., vol. 79, no. 14, 2005, pages 8828 - 8834 |
| BIRD ET AL., SCIENCE, vol. 242, no. 4877, 1988, pages 423 - 426 |
| BRIIHL ET AL., J. IMMUNOL., vol. 166, no. 4, 2001, pages 2420 - 2426 |
| CHEN ET AL., J. MOL. BIOL., vol. 293, no. 4, 1999, pages 865 - 881 |
| CORPET, NUCLEIC ACIDS RES., vol. 16, no. 22, 1988, pages 10881 - 10890 |
| DALL'ACQUA ET AL., J. BIOL. CHEM., vol. 281, no. 33, 2006, pages 23514 - 23524 |
| ESPINOSA ET AL., J INFECT DIS, vol. 212, 2015, pages 1111 - 1119 |
| GARDNER ET AL., NATURE, vol. 519, no. 7541, 2015, pages 87 - 91 |
| HARLOWLANE: "Remington: The Science and Practice of Pharmacy", 2013, COLD SPRING HARBOR PUBLICATIONS |
| HIGGINSSHARP, BIOINFORMATICS, vol. 5, no. 2, 1989, pages 151 - 3 |
| HIGGINSSHARP, GENE, vol. 73, no. 1, 1988, pages 237 - 244 |
| HINTON ET AL., J IMMUNOL., vol. 176, 2006, pages 346 - 356 |
| HUANG ET AL., BIOINFORMATICS, vol. 8, no. 2, 1992, pages 155 - 165 |
| HUSTON ET AL., PROC. NATL. ACAD. SCI. U.S.A., vol. 85, no. 16, 1988, pages 5879 - 5883 |
| JOHNSON ET AL., NAT. MED., vol. 15, no. 8, 2009, pages 901 - 906 |
| KABAT ET AL.: "NIH Publication No. 91-3242", 1991, PUBLIC HEALTH SERVICE, NATIONAL INSTITUTES OF HEALTH, article "Sequences of Proteins of Immunological Interest" |
| KANDA ET AL., BIOTECHNOL. BIOENG., vol. 94, no. 4, 2006, pages 680 - 688 |
| KASTENMULLER ET AL., INFECT IMMUN, vol. 81, 2013, pages 789 - 800 |
| KISALU ET AL., NAT MED, vol. 24, 2018, pages 408 - 416 |
| KISALU ET AL., NAT. MED., vol. 24, 2018, pages 408 - 416 |
| KOLLI ET AL., PROC NATL ACAD SCI U S A, vol. 119, 2022, pages e2209729119 |
| KREUTER: "Colloidal Drug Delivery Systems", 1994, MARCEL DEKKER, INC., pages: 219 - 342 |
| KUFER ET AL., CANCER IMMUNOL. IMMUNOTHER., vol. 45, no. 3-4, 1997, pages 193 - 197 |
| LAZAR ET AL., PROC. NATL., ACAD. SCI. U.S.A., vol. 103, no. 11, 2006, pages 4005 - 4010 |
| LEFRANC ET AL.: "IMGT unique numbering for immunoglobulin and T cell receptor variable domains and Ig superfamily V-like domains", DEV. COMP. IMMUNOL., vol. 27, no. 1, 2003, pages 55 - 77, XP055585227, DOI: 10.1016/S0145-305X(02)00039-3 |
| LOFFLER ET AL., BLOOD, vol. 95, no. 6, 2000, pages 2098 - 2103 |
| LONBERG, NAT. BIOTECH., vol. 23, 2005, pages 1117 - 1125 |
| LONENBERG, CURR. OPIN. IMMUNOL., vol. 20, 2008, pages 450 - 459 |
| MACK ET AL., J. IMMUNOL., vol. 158, no. 8, 1997, pages 3965 - 3970 |
| MACK ET AL., PROC. NATL. ACAD. SCI. U.S.A., vol. 92, no. 15, 1995, pages 7021 - 7025 |
| MARBRYSNAVELY, IDRUGS, vol. 1-2, no. 8, 2010, pages 543 - 549 |
| MCCAFFERTY ET AL., NATURE, vol. 348, 1990, pages 552 - 554 |
| NEEDLEMANWUNSCH, J. MOL. BIOL., vol. 48, no. 3, 1970, pages 443 - 453 |
| NEVILLE K KISALU ET AL: "A human monoclonal antibody prevents malaria infection by targeting a new site of vulnerability on the parasite", NATURE MEDICINE, vol. 24, no. 4, 19 March 2018 (2018-03-19), New York, pages 408 - 416, XP055717091, ISSN: 1078-8956, DOI: 10.1038/nm.4512 * |
| OKAZAKI ET AL., J. MOL. BIOL., vol. 336, no. 5, 2004, pages 1239 - 1249 |
| OYEN ET AL., PROC NATL ACAD SCI U S A, vol. 114, 2017, pages E10438 - E10445 |
| OYEN ET AL.: "Structural basis for antibody recognition of the NANP repeats in Plasmodium falciparum circumsporozoite protein", PROC NATL ACAD SCI USA, vol. 114, 2017, pages E10438 - E10445, XP055717496, DOI: 10.1073/pnas.1715812114 |
| PEARSON, METHODS MOL. BIOL., vol. 24, 1994, pages 307 - 331 |
| PETKOVA ET AL., INT. IMMUNOL., vol. 18, no. 12, 2006, pages 1759 - 1769 |
| PRESTA ET AL., CANCER RES., vol. 57, no. 20, 1997, pages 4593 - 4599 |
| RAGHUNANDAN ET AL., MALAR J, vol. 19, 2020, pages 113 |
| RIPKA ET AL., ARCH. BIOCHEM. BIOPHYS., vol. 249, no. 2, 1986, pages 533 - 545 |
| SCHOONJANS ET AL., J. IMMUNOL., vol. 165, no. 12, 2000, pages 7050 - 7057 |
| SINGH ET AL., J BIOL CHEM, vol. 295, 2020, pages 403 - 414 |
| SMITHWATERMAN, ADV. APPL. MATH., vol. 2, no. 4, 1981, pages 482 - 489 |
| TICETABIBI: "Treatise on Controlled Drug Delivery: Fundamentals, Optimization, Applications", 1992, MARCEL DEKKER, INC., pages: 315 - 339 |
| WANG ET AL., IMMUNITY, vol. 53, 2020, pages 733 - 744 |
| WARD ET AL., NATURE, vol. 341, no. 6242, 1989, pages 544 - 546 |
| WILLEMS ET AL., J. CHROMATOGR. B ANALYT. TECHNOL. BIOMED LIFE SCI., vol. 786, no. 1-2, 2003, pages 161 - 176 |
| WRIGHT ET AL., TRENDS BIOTECHNOL, vol. 15, no. 1, 1997, pages 26 - 32 |
| YAMANE-OHNUKI ET AL., BIOTECHNOL. BIOENG., vol. 87, no. 5, 2004, pages 614 - 622 |
| ZALEVSKY ET AL., NATURE BIOTECHNOL., vol. 28, no. 2, 2010, pages 157 - 159 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| US11760794B2 (en) | Neutralizing antibodies to plasmodium falciparum circumsporozoite protein and their use | |
| US20250263478A1 (en) | Neutralizing antibodies to plasmodium falciparum circumsporozoite protein and their use | |
| US20250145690A1 (en) | Human monoclonal antibodies that broadly target coronaviruses | |
| US10273288B2 (en) | Neutralizing antibodies to Ebola virus glycoprotein and their use | |
| US20240239873A1 (en) | Neutralizing antibodies to ebola virus glycoprotein and their use | |
| US20230348568A1 (en) | Epstein-barr virus monoclonal antibodies and uses thereof | |
| WO2022132904A1 (en) | Human monoclonal antibodies targeting sars-cov-2 | |
| US20250197483A1 (en) | Neutralizing antibodies to hiv-1 env and their use | |
| EP4638491A1 (en) | Monoclonal antibodies for treating sars-cov-2 infection | |
| WO2021257665A1 (en) | Human monoclonal antibody that targets a conserved site on the plasmodium falciparum circumsporozoite protein | |
| WO2024064826A1 (en) | Neutralizing antibodies to plasmodium falciparum circumsporozoite protein and their use | |
| WO2024243355A1 (en) | Human monoclonal antibodies that target the rh5 complex of blood-stage plasmodium falciparum | |
| US20250011406A1 (en) | Neutralizing antibodies to plasmodium falciparum circumsporozoite protein and their use | |
| WO2025024233A1 (en) | Bispecific antibodies that broadly target coronaviruses | |
| WO2024054822A1 (en) | Engineered sars-cov-2 antibodies with increased neutralization breadth | |
| WO2026030473A1 (en) | West nile virus neutralizing monoclonal antibodies | |
| WO2025137284A2 (en) | Broadly neutralizing antibodies against sars-cov-2 and sars-cov variants | |
| HK40057872A (en) | Neutralizing antibodies to ebola virus glycoprotein and their use |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| 121 | Ep: the epo has been informed by wipo that ep was designated in this application |
Ref document number: 23793183 Country of ref document: EP Kind code of ref document: A1 |
|
| NENP | Non-entry into the national phase |
Ref country code: DE |
|
| 122 | Ep: pct application non-entry in european phase |
Ref document number: 23793183 Country of ref document: EP Kind code of ref document: A1 |