WO2021239666A1 - Therapeutic methods - Google Patents
Therapeutic methods Download PDFInfo
- Publication number
- WO2021239666A1 WO2021239666A1 PCT/EP2021/063764 EP2021063764W WO2021239666A1 WO 2021239666 A1 WO2021239666 A1 WO 2021239666A1 EP 2021063764 W EP2021063764 W EP 2021063764W WO 2021239666 A1 WO2021239666 A1 WO 2021239666A1
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- protein
- seq
- antibody
- gib
- pla2
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Ceased
Links
Classifications
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K39/00—Medicinal preparations containing antigens or antibodies
- A61K39/12—Viral antigens
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K39/00—Medicinal preparations containing antigens or antibodies
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P31/00—Antiinfectives, i.e. antibiotics, antiseptics, chemotherapeutics
- A61P31/12—Antivirals
- A61P31/14—Antivirals for RNA viruses
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K16/00—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies
- C07K16/40—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against enzymes
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K39/00—Medicinal preparations containing antigens or antibodies
- A61K2039/505—Medicinal preparations containing antigens or antibodies comprising antibodies
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K39/00—Medicinal preparations containing antigens or antibodies
- A61K2039/58—Medicinal preparations containing antigens or antibodies raising an immune response against a target which is not the antigen used for immunisation
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N2770/00—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA ssRNA viruses positive-sense
- C12N2770/00011—Details
- C12N2770/20011—Coronaviridae
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N2770/00—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA ssRNA viruses positive-sense
- C12N2770/00011—Details
- C12N2770/20011—Coronaviridae
- C12N2770/20034—Use of virus or viral component as vaccine, e.g. live-attenuated or inactivated virus, VLP, viral protein
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N9/00—Enzymes; Proenzymes; Compositions thereof; Processes for preparing, activating, inhibiting, separating or purifying enzymes
- C12N9/14—Hydrolases (3)
- C12N9/16—Hydrolases (3) acting on ester bonds (3.1)
- C12N9/18—Carboxylic ester hydrolases (3.1.1)
- C12N9/20—Triglyceride splitting, e.g. by means of lipase
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Y—ENZYMES
- C12Y301/00—Hydrolases acting on ester bonds (3.1)
- C12Y301/01—Carboxylic ester hydrolases (3.1.1)
- C12Y301/01004—Phospholipase A2 (3.1.1.4)
Definitions
- the present invention relates to novel compounds, compositions, uses and methods for treating or detecting RNA virus infections in mammals, particularly in human subjects.
- the invention may be in used in a preventive or curative approach, alone or in combination with other treatments.
- sPLA2-GIB is involved in the inactivation of CD4 T cells in HIV infected patients, and that sPLA2-GIB inhibitors are effective for treating disorders associated with an immune deficiency (see WO2015/097140).
- pathogens produce or activate cofactors which bind gC1qR and cause a sensitization of CD4 T cells to inactivation by sPLA2-GIB.
- modulating such cofactors is an effective approach for treating such diseases (WO2019/166412).
- SARS-Cov2 infection can also utilize the sPLA2-GIB pathway.
- the inventors have identified that SARS-Cov2 can act as a sPLA2-GIB cofactor, causing T cell anergy, especially mediated by the spike protein.
- the invention thus provides novel efficient molecules, compositions, methods and uses for detecting and/or treating coronavirus infections as well as other RNA virus infections, particularly SARS-Cov2 infection.
- An object of the invention relates to methods for treating a Group IV RNA vims infection, particularly a coronavirus infection, in a mammal, particularly a human, comprising inhibiting the sPLA2-GIB pathway in said mammal.
- a further object of the invention relates to the use of an inhibitor of the sPLA2-GIB pathway for the manufacture of a medicament for treating a Group IV RNA vims infection, particularly a coronavirus infection.
- Another object of the invention concerns an inhibitor of the sPLA2-GIB pathway for use for treating a Group IV RNA vims infection, particularly a coronavirus infection.
- a further object of the invention relates to any methods for inhibiting a sPLA2-GIB cofactor effect of SARS-Cov2 in a mammal.
- a further object of the invention is a polypeptide comprising a sequence selected from anyone of SEQ ID Nos: 2-208 or an immunogenic-fragment thereof.
- the polypeptide is preferably a polypeptide of less than 50 amino acids, more preferably less than 40 amino acids.
- the polypeptide may consist or consist essentially of anyone of SEQ ID Nos: 2- 208.
- the polypeptides of the invention may be conjugated to any molecule, such as a protein, carrier, surface, device, adjuvant, bead, column, etc.
- a further object of the invention relates to a molecule which binds a Group IV RNA virus, particularly a coronavirus, in a binding domain comprising at least one amino acid residue of any one of SEQ ID Nos: 2-208.
- a further object of the invention relates to an antibody (or a fragment or derivative thereof) which binds a Group IV RNA virus, particularly a coronavirus, wherein said antibody binds an epitope comprising at least one amino acid residue of any one of SEQ ID Nos: 2-208.
- a further object of the invention relates to a vaccine composition
- a vaccine composition comprising an immunogen, wherein the immunogen comprises a sequence selected from anyone of SEQ ID Nos: 2-208 or a fragment thereof.
- a further object of the invention relates to a vaccine composition
- a vaccine composition comprising an immunogen, wherein the immunogen comprises a viral envelope protein with a modified gC1qR binding motif, preferably a deleted and/or mutated gC1qR binding motif.
- the immunogen comprises a viral envelope protein with a modified gC1qR binding motif, preferably a deleted and/or mutated gC1qR binding motif.
- a specific example is a SARS-Cov-2 spike protein having at least one amino acid modification in a domain selected from SEQ ID NO: 156-185, even more preferably in a domain selected from aa 348-362 or 431-445 (by reference to SEQ ID NO: 209).
- the invention also relates to a combination therapy or therapeutic regimen for treating a Group IV RNA virus infection, particularly a coronavirus, comprising (i) an inhibitor of PLA2-GIB pathway, in combination with (ii) at least one further active agent.
- the further active agent may be an antiviral agent, an antibiotic, an anti-inflammatory agent, etc.
- the agents in a combination therapy of the invention may be formulated together or separately, for combined, separate or sequential administration(s).
- the invention also relates to the use of a PLA2-GIB cofactor, or an agonist, fragment, derivative, or mimotope thereof, for the manufacture of a medicament to induce immunosuppression in a subject in need thereof, by increasing the effect of PLA2-GIB on T cells.
- the invention may be used in any mammal, particularly in human subjects. It is suitable to treat Group IV RNA vims infections, particularly coronavirus infections, at any stage of infection, either alone or in combination with other agents/therapies.
- FIG. 1 Position Specific Scoring Matrix (PSSM) defining peptide binding motif to gClqR.
- PSSM Position Specific Scoring Matrix
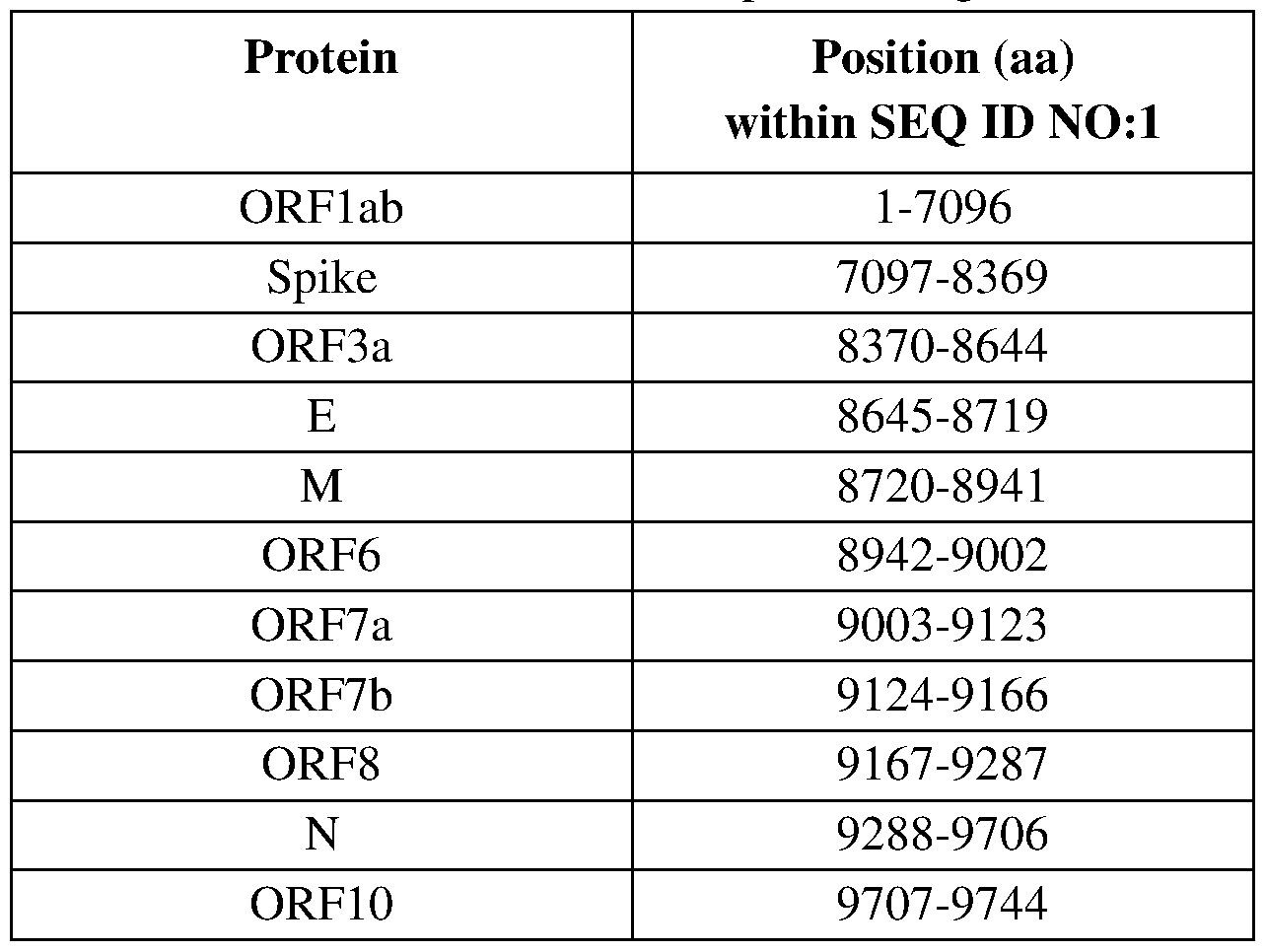
- Figure 2 gClqR:PSSM motifs across the SARS-CoV-2 proteome (SEQ ID NO: 1).
- the graphic reports gC1qR:PSSM scores for every given 15 a.a. window scanning the SARS-CoV-2 proteome (SEQ ID Nol).
- the gC1qR:PSSM score is assigned to the N- terminal amino acid position of every given 15 a.a. window (2A).
- the location of the different open-reading frames within the SARS-CoV-2 proteome are presented below the x-axis.
- FIG. 3 High scoring gClqR:PSSM motifs across the spike protein sequence from SARS-CoV-2 (SEQ ID NO: 209).
- the graphic reports gC1qR:PSSM scores for every given 15 a.a. window scanning the sequence of spike from SARS-CoV-2.
- gC1qR:PSSM scores assigned to 15 a.a. motifs are represented by bars.
- Figure 4 Co-localization of high scoring gClqR:PSSM motifs with solvent accessible domains within the spike protein sequence.
- the graphic reports gC1qR:PSSM scores for every given 15 a.a. window scanning the sequence of spike from SARS-CoV-2.
- gC1qR:PSSM scores assigned to 15 a.a. motifs are represented by bars.
- Solvent accessible scores were calculated according to the method presented in example 3. The location of the different open- reading frames within the SARS-CoV-2 proteome are presented below the x-axis.
- FIG. 5 Co-localization of high scoring gC1qR:PSSM motifs with predicted B cell epitopes (Discotope) within the spike protein sequence.
- the graphic reports gC1qR:PSSM scores for every given 15 a.a. window scanning the sequence of spike from SARS-CoV-2.
- gC1qR:PSSM scores assigned to 15 a.a. motifs are represented by bars.
- Discotope scores were calculated according to the method presented in example 3. The location of the different open- reading frames within the SARS-CoV-2 proteome are presented below the x-axis.
- Figure 6 Co-localization of high scoring gC1qR:PSSM motifs with predicted HLA class II binding motif-rich domains within the spike protein sequence.
- the graphic reports gC1qR:PSSM scores for every given 15 a.a. window scanning the sequence of spike from SARS-CoV-2.
- gC1qR:PSSM scores assigned to 15 a.a. motifs are represented by bars.
- the invention generally relates to novel molecules, compositions, uses and methods for treating or detecting a subject infected by a Group IV RNA virus, particularly SARS- Cov2.
- SARS-Cov-2 exerts PLA2-G1B cofactor effect.
- Applicant identified various peptides in SARS-Cov2 proteins that can bind to gC1qR and cause CD4 T cell anergy.
- Particularly relevant peptides are located in SI protein, such as in the N-terminal domain (NTD) and in the receptor binding domain (RBD).
- NTD N-terminal domain
- RBD receptor binding domain
- These peptides have remarkable immunogenic character (e.g., they overlap with predicted B cell epitopes) and some are present in the RBD and thus represent efficient molecules for inhibiting Group IV RNA virus infections, particularly coronavirus infections, as well as for producing protective antibodies.
- Vaccines against these regions would also be a very potent strategy to protect against SARS-Cov2 infection by inhibiting both PLA2G1B deleterious effect and viral infection.
- PLA2-GIB designates group IB pancreatic phospholipase A2.
- PLA2-GIB has been identified and cloned from various mammalian species. The human PLA2-GIB protein is disclosed, for instance, in Lambeau and Gelb (2008). The sequence is available on Genbank No. NP_000919.
- the amino acid sequence of an exemplary human PLA2-GIB is shown below (SEQ ID NO: 210).
- PLA2-GIB designates preferably human PLA2-GIB.
- the human PLA2-GIB protein may be present under two distinct forms: a pro form (pro- sPLA2-GIB), which is activated by proteolytic cleavage of a pro-peptide, leading to the mature secreted form (sPLA2-GIB).
- the term PLA2-GIB includes any form of the protein, such as the pro-form and/or the mature form.
- the mature secreted form comprises the sequence of amino acid residues 23-148 of SEQ ID NO: 210, or any natural variants thereof.
- Natural variants of a protein include variants resulting e.g., from polymorphism or splicing. Natural variants may also include any protein comprising the sequence of SEQ ID NO: 210, or the sequence of amino acid residues 23-148 of SEQ ID NO: 210, with one or more amino acid substitution(s), addition(s) and/or deletion(s) of one or several (typically 1, 2 or 3) amino acid residues. Variants include naturally-occurring variants having e.g., at least 90% amino acid sequence identity to SEQ ID NO: 210.
- PLA2-GIB has at least one activity selected from induction of formation of membrane microdomains (MMD) in CD4 T cells from healthy subjects, or rendering CD4 T cells of healthy subjects refractory to interleukin signaling, such as refractory to IL-2 signaling or refractory to IL-7 signaling or refractory to IL-4 signaling.
- MMD membrane microdomains
- rendering CD4 T cells of healthy subjects refractory to interleukin-7 signaling comprises a reduction of STAT5A and/or B phosphorylation in said cells by at least about 10%, at least about 20%, at least about 30%, or at least about 40%. In some embodiments rendering CD4 T cells of healthy subjects refractory to interleukin-7 signaling comprises reducing the rate of nuclear translocation of phospho-STAT5A and/or phospho-STAT5B by at least about 20%, at least about 30%, at least about 40%, or at least about 50%.
- sequence identity refers to the quantification (usually percentage) of nucleotide or amino acid residue matches between at least two sequences aligned using a standardized algorithm such as Smith- Waterman alignment (Smith and Waterman (1981) J Mol Biol 147:195-197), CLUSTALW (Thompson et al. (1994) Nucleic Acids Res 22:4673-4680), or BLAST2 (Altschul et al. (1997) Nucleic Acids Res 25:3389-3402).
- BLAST2 may be used in a standardized and reproducible way to insert gaps in one of the sequences in order to optimize alignment and to achieve a more meaningful comparison between them.
- inactivation indicates, in relation to CD4 T cells, that such cells lose at least part of their ability to contribute to the development of an effective immune response. Inactivation may be partial or complete, transient or permanent. Inactivation designates preferably reducing by at least 10%, 20%, 30%, 40%, 50%, 60%, 70%, 80% or more a function of CD4 T cells, particularly pSTAT5 nuclear translocation and/or CD4 T cell’s immunostimulatory activity. Typically, inactive CD4 T cells have no effective pSTAT5 nuclear translocation. In a particular embodiment, an inactive CD4 T cell is an anergic CD4 T cell.
- the term “resistance” (or “insensitivity”) of CD4 T cells to inactivation by sPLA2-GIB indicates, within the context of this invention, that CD4 T cells are essentially not inactivated in vitro when incubated in the presence of 5nM of sPLA2-GIB. Resistance indicates, for instance, that CD4 T cells retain an active nuclear translocation of pSTAT5 when incubated in vitro in the presence of 5nM sPLA2-GIB and interleukin-7. Resistance (or insensitivity) of CD4 T cells to sPLA2-GIB may also indicate that CD4 T cells incubated in vitro with 5nM PLA2-GIB remain immunologically functional, e.g., do not become anergic.
- sPLA2-GIB Pathway designates any component or molecule involved in sPLA2-G IB -mediated T cell anergy. This includes SPLA2-G1B, as well as any cofactor of SPLA2-G1B.
- treatment includes preventive and curative treatments.
- the treatment includes prevention of vims infection, such as reduction of the infection, as well as treatment of an existing infection (e.g., reduction of viral load, reduction of immune deficiency, reduction of CD4 T cell anergy) and related disease(s) or symptom(s) (e.g., pain, fever, cough, respiratory disorders, etc.).
- treatment includes a reduction of mortality caused by or associated with coronavirus infection.
- Group IV RNA virus refers to the Baltimore classification and includes, without limitation, coronaviruses, such as particularly Sars-cov-2, Dengue vims, Chikungunya vims, and Polio vims.
- SARS Cov-2 utilizes the sPLA2-GIB pathway
- SARS-CoV-2 vims (a.k.a 2019-nCoV), is the causative agent of covid-19. There are no approved treatments against SARS-CoV-2 vims infection, and vaccines being developed are not expected to reach patients promptly.
- SARS-CoV-2 vims is a coronavirus, with a fully sequenced single- stranded RNA genome.
- the inventors have surprisingly found that SARS-Cov-2 utilizes the sPLA2-GIB pathway. In particular, the inventors have found that SARS-Cov-2 can act by rendering CD4 T cells sensitive to inactivation by PLA2-GIB.
- Such mechanism involves the binding of a molecule of (or induced by) SARS-Cov-2 to gC1qR at the surface of CD4 T cells, causing sensitization of CD4 T cells to inactivation by physiological concentrations of PLA2-GIB.
- various proteins of SARS-Cov-2 such as the spike protein, contain sequences or regions that can bind to gC1qR and may act as cofactor increasing the sPLA2-GIB inhibitory activity on CD4 T cells.
- Applicant thus identified a novel general approach for treating coronavirus infection, through a modulation of the SPLA2-G1B pathway.
- One aspect of the invention resides in the treatment of a Group IV RNA virus infection, particularly coronavirus infections using an inhibitor of PLA2-G1B.
- PLA2-GIB inhibitors suitable for use in the invention may be any compound that inhibits or neutralizes the expression or activity of PLA2-GIB, such as expression inhibitors, antagonists, or sequestrators.
- Preferred types of inhibitors include PLA2-GIB ligands (covalent or non-covalent), anti-PLA2-GIB antibodies (and fragments and derivatives thereof), nucleic acids encoding anti-PLA2-GIB antibodies (or fragments and derivatives thereof), inhibitory nucleic acids, peptides, or small drugs, soluble receptors, or combination(s) thereof.
- the PLA2-GIB inhibitor can be a PLA2-GIB antigen which, upon administration to the subject, induces the production of anti-PLA2GIB antibodies.
- Inhibiting PLA2-GIB designates typically reducing by at least 10%, 20%, 30%, 40%, 50%, 60%, 70%, 80% or more PLA2-GIB level or activity, as well as completely blocking or suppressing PLA2-GIB level or activity. Depending on the situation, inhibition may be transient, sustained or permanent.
- PLA2-GIB inhibitors are anti-PLA2-GIB antibodies, e.g., antibodies that bind to PLA2-GIB and/or have been generated by immunization of a mammal with a PLA2-GIB antigen.
- Antibodies can be synthetic, monoclonal, or polyclonal and can be made by techniques well known in the art. Such antibodies specifically bind via the antigen-binding sites of the antibody (as opposed to non-specific binding).
- PLA2-GIB polypeptides, fragments, variants, fusion proteins, etc. can be employed as immunogens in producing antibodies immunoreactive therewith. More specifically, the polypeptides, fragments, variants, fusion proteins, etc. contain antigenic determinants or epitopes that elicit the formation of antibodies.
- antibodies is meant to include polyclonal antibodies, monoclonal antibodies, fragments thereof, such as F(ab')2 and Fab fragments, single-chain variable fragments (scFvs), single-domain antibody fragments (VHHs or Nanobodies), bivalent antibody fragments (diabodies), as well as any recombinantly and synthetically produced binding partners, human antibodies or humanized antibodies.
- Antibodies are defined to be specifically binding preferably if they bind to PLA2-GIB with a Ka of greater than or equal to about 10 7 M-l. Affinities of antibodies can be readily determined using conventional techniques, for example those described by Scatchard et al., Ann. N.Y. Acad. Sci., 51:660 (1949).
- Polyclonal antibodies can be readily generated from a variety of sources, for example, horses, cows, donkeys, goats, sheep, dogs, chickens, rabbits, mice, or rats, using procedures that are well known in the art.
- purified PLA2-GIB or a peptide based on the amino acid sequence of PLA2-GIB that is appropriately conjugated is administered to the host animal typically through parenteral injection.
- the immunogenicity of PLA2-GIB can be enhanced through the use of an adjuvant, for example, Freund's complete or incomplete adjuvant. Following booster immunizations, small samples of serum are collected and tested for reactivity to PLA2-GIB polypeptide.
- Examples of various assays useful for such determination include those described in Antibodies: A Laboratory Manual, Harlow and Lane (eds.), Cold Spring Harbor Laboratory Press, 1988; as well as procedures, such as countercurrent Immuno- electrophoresis (CIEP), radioimmunoassay, radio-immunoprecipitation, enzyme-linked immunosorbent assays (ELISA), dot blot assays, and sandwich assays. See U.S. Pat. Nos. 4,376, 110 and 4,486,530.
- Monoclonal antibodies can be readily prepared using well known procedures. See, for example, the procedures described in U.S. Pat. Nos. RE 32,011, 4,902,614, 4,543,439, and 4,411,993; Monoclonal Antibodies, Hybridomas: A New Dimension in Biological Analyses, Plenum Press, Kennett, McKeam, and Bechtol (eds.), 1980.
- the host animals such as mice
- Mouse sera are then assayed by conventional dot blot technique or antibody capture (ABC) to determine which animal is best to fuse.
- ABSC antibody capture
- mice are given an intravenous boost of protein or peptide.
- Mice are later sacrificed and spleen cells fused with commercially available myeloma cells, such as Ag8.653 (ATCC), following established protocols.
- ATCC Ag8.653
- the myeloma cells are washed several times in media and fused to mouse spleen cells at a ratio of about three spleen cells to one myeloma cell.
- the fusing agent can be any suitable agent used in the art, for example, polyethylene glycol (PEG). Fusion is plated out into plates containing media that allows for the selective growth of the fused cells. The fused cells can then be allowed to grow for approximately eight days. Supernatants from resultant hybridomas are collected and added to a plate that is first coated with goat anti-mouse Ig. Following washes, a label, such as a labeled PFA2-GIB polypeptide, is added to each well followed by incubation. Positive wells can be subsequently detected. Positive clones can be grown in bulk culture and supernatants are subsequently purified over a Protein A column (Pharmacia).
- PEG polyethylene glycol
- the monoclonal antibodies of the disclosure can be produced using alternative techniques, such as those described by Alting-Mees et al., "Monoclonal Antibody Expression Fibraries: A Rapid Alternative to Hybridomas", Strategies in Molecular Biology 3:1-9 (1990), which is incorporated herein by reference.
- binding partners can be constructed using recombinant DNA techniques to incorporate the variable regions of a gene that encodes a specific binding antibody. Such a technique is described in Farrick et al., Biotechnology, 7:394 (1989).
- Antigen-binding fragments of such antibodies which can be produced by conventional techniques, are also encompassed by the present invention.
- fragments include, but are not limited to, Fab and F(ab')2 fragments.
- Antibody fragments and derivatives produced by genetic engineering techniques are also provided.
- the monoclonal antibodies include chimeric antibodies, e.g., humanized versions of murine monoclonal antibodies. Such humanized antibodies can be prepared by known techniques, and offer the advantage of reduced immunogenicity when the antibodies are administered to humans.
- a humanized monoclonal antibody comprises the variable region of a murine antibody (or just the antigen binding site thereof) and a constant region derived from a human antibody.
- a humanized antibody fragment can comprise the antigen binding site of a murine monoclonal antibody and a variable region fragment (lacking the antigen-binding site) derived from a human antibody. Procedures for the production of chimeric and further engineered monoclonal antibodies include those described in Riechmann et al.
- Antibodies produced by genetic engineering methods such as chimeric and humanized monoclonal antibodies, comprising both human and non-human portions, which can be made using standard recombinant DNA techniques, can be used.
- Such chimeric and humanized monoclonal antibodies can be produced by genetic engineering using standard DNA techniques known in the art, for example using methods described in Robinson et al. International Publication No. WO 87/02671; Akira, et al. European Patent Application 0184187; Taniguchi, M., European Patent Application 0171496; Morrison et al. European Patent Application 0173494; Neuberger et al. PCT International Publication No. WO 86/01533; Cabilly et al. U.S. Pat. No.
- antibodies In connection with synthetic and semi-synthetic antibodies, such terms are intended to cover but are not limited to antibody fragments, isotype switched antibodies, humanized antibodies (e.g., mouse-human, human-mouse), hybrids, antibodies having plural specificities, and fully synthetic antibody-like molecules.
- Human monoclonal antibodies having human constant and variable regions can be generated by immunizing transgenic animals which contain human immunoglobulin genes. See Jakobovits et al. Ann NY Acad Sci 764:525-535 (1995). Human monoclonal antibodies against PLA2-GIB polypeptides can also be prepared by constructing a combinatorial immunoglobulin library, such as a Fab phage display library or a scFv phage display library, using immunoglobulin light chain and heavy chain cDNAs prepared from mRNA derived from lymphocytes of a subject. See, e.g., McCafferty et al. PCT publication WO 92/01047; Marks et al. (1991) J. Mol.
- a combinatorial library of antibody variable regions can be generated by mutating a known human antibody.
- a variable region of a human antibody known to bind PLA2-GIB can be mutated by, for example, using randomly altered mutagenized oligonucleotides, to generate a library of mutated variable regions which can then be screened to bind to PLA2-GIB.
- Methods of inducing random mutagenesis within the CDR regions of immunoglobin heavy and/or light chains, methods of crossing randomized heavy and light chains to form pairings and screening methods can be found in, for example, Barbas et al. PCT publication WO 96/07754; Barbas et al. (1992) Proc. Nat'l Acad. Sci. USA 89:4457 4461.
- An immunoglobulin library can be expressed by a population of display packages, preferably derived from filamentous phage, to form an antibody display library.
- Examples of methods and reagents particularly amenable for use in generating antibody display library can be found in, for example, Ladner et al. U.S. Pat. No. 5,223,409; Kang et al. PCT publication WO 92/18619; Dower et al. PCT publication WO 91/17271; Winter et al. PCT publication WO 92/20791; Markland et al. PCT publication WO 92/15679; Breitling et al. PCT publication WO 93/01288; McCafferty et al.
- the antibody library is screened to identify and isolate packages that express an antibody that binds a PLA2-GIB polypeptide.
- a display package e.g., filamentous phage
- the primary screening of the library involves panning with an immobilized PLA2-GIB polypeptide and display packages expressing antibodies that bind immobilized PLA2-GIB polypeptide are selected.
- Preferred antibodies for use in the invention are directed to a PLA2-GIB epitope, and/or have been generated by immunization with a polypeptide comprising a PLA2-GIB epitope selected from: the mature PLA2-GIB protein, a fragment of PLA2-GIB comprising at least 8 consecutive amino acid residues of SEQ ID NO: 210 (or the corresponding residues of a natural variant of SEQ ID NO: 210), said fragment preferably comprising at least amino acid 70, amino acid 121, amino acid 50, amino acid 52, amino acid 54, amino acid 71, or a combination thereof (numbering by reference to SEQ ID NO: 210).
- Particular anti-PLA2-GIB antibodies for use in the invention bind mature human PLA2- GIB, even more preferably an epitope comprised in a domain of PLA2-GIB comprising an amino acid residue selected from amino acid 70, amino acid 121, amino acid 50, amino acid 52, amino acid 54, amino acid 71, or a combination thereof (numbering by reference to SEQ ID NO: 210).
- Particular antibodies for use in the invention bind an epitope comprised between amino acid residues 50-71 of mature human PLA2-GIB (by reference to SEQ ID NO: 210) or the corresponding residues of a natural variant of SEQ ID NO: 210.
- Examples of anti-PLA2-GIB antibodies suitable for use in the invention have been disclosed in W02015/097140.
- anti-PLA2-GIB antibodies for use in the present invention bind an epitope comprising at least one amino acid residue selected from W3, R6, K7, K10, C77, Y75, G79 and S80 of human mature PLA2-GIB (numbering by reference to SEQ ID NO: 210), more preferably an epitope comprising at least 2 or at least 3 amino acid residues selected from W3, R6, K7, K10, C77, Y75, G79 and S80 of human mature PLA2-GIB, further more preferably an epitope comprising at least 4, at least 5, at least 6 or at least 7 amino acid residues selected from W3, R6, K7, K10, C77, Y75, G79 and S80 of human mature PLA2-GIB.
- antibodies for use in the invention bind an epitope comprising an amino acid residue comprised between amino acids 1-10 or 75-80 of mature human PLA2-GIB (by reference to SEQ ID NO: 210) or the corresponding residues of a natural variant of SEQ ID NO: 210.
- Such antibodies exhibit potent neutralizing activity and represent valuable therapeutic agents for use in the invention.
- anti-PLA2-GIB antibodies suitable for use in the invention have been disclosed in WO2019/166665, incorporated therein by reference.
- the antibodies or derivatives for use in the invention is monoclonal antibody 14G9 as disclosed in WO2019/166665, or an anti-PLA2-GIB antibody that competitively inhibits binding of monoclonal antibody 14G9 to human PLA2-GIB.
- the antibody may be human or humanized.
- the antibodies or derivatives for use in the invention is monoclonal antibody #2B as disclosed in WO2019/166665, or an anti-PLA2-GIB antibody that competitively inhibits binding of monoclonal antibody #2B to human SPLA2-GIB.
- the antibodies or derivatives for use in the invention is monoclonal antibody #2B1 as disclosed in WO2019/166665, or an anti-PLA2-GIB antibody that competitively inhibits binding of monoclonal antibody #2B1 to human SPLA2-GIB.
- the antibodies or derivatives for use in the invention is monoclonal antibody #2B2 as disclosed in WO2019/166665, or an anti-PLA2-GIB antibody that competitively inhibits binding of monoclonal antibody #2B2 to human SPLA2-GIB.
- competitive inhibits indicates that the antibody can reduce or inhibit or displace the binding of a reference antibody to sPLA2-GIB.
- Competition assays can be performed using standard techniques such as, for instance, competitive ELISA or other binding assays.
- a competitive binding assay involves a purified target antigen, generally bound either to a solid substrate or cells, an unlabeled test antibody and a labeled reference antibody.
- Competitive inhibition is measured by determining the amount of labeled antibody bound in the presence of the test antibody.
- the test antibody is present in excess, such as about 5 to 500 times the amount of reference antibody.
- the test antibody is in 100X excess, and for enzymatic methods, the test antibody in in 10X excess.
- test antibody present in excess inhibits or displaces at least 70% of the binding of the reference antibody to the antigen, it is considered as competitively inhibiting said reference antibody.
- test antibody present in 100X excess inhibits or displaces at least 70%, more preferably at least 80% of the binding of the reference antibody to the antigen in ELISA, it is considered as competitively inhibiting said reference antibody.
- Preferred competing antibodies bind epitopes that share common amino acid residues.
- the inhibitor is a monoclonal antibody comprising: (i) a light chain variable region comprising a CDR-L1, a CDR-L2, a CDR-L3 and a FR- L, wherein the CDR-L1, CDR-L2 and/or CDR-L3 consists, or consists essentially, of the CDR-L1, CDR-L2 and CDR-L3, respectively, of the light chain variable region of SEQ ID NO: 3 or 5 as disclosed in WO2019/166665, and wherein a FR-L is of a human immunoglobulin sequence; and
- a heavy chain variable region comprising a CDR-H1, a CDR-H2, a CDR-H3 and a FR-H, wherein the CDR-H1, CDR-H2 and/or CDR-H3 consists, or consists essentially, of the CDR-H1, CDR-H2 and CDR-H3, respectively, of the heavy chain variable region of SEQ ID NO: 4, 6, 7 or 8 as disclosed in WO2019/166665, and wherein a FR-H is of a human immunoglobulin sequence.
- variable region refers to the amino-terminal domains of the heavy or light chain (“VH” or “VL”), which contain the antigen-binding sites.
- VL or VH generally consists of a framework region (“FR”) interrupted by three hypervariable regions referred to as "complementarity determining regions" or "CDRs".
- FR framework region
- CDRs complementarity determining regions
- CDR-L1 amino acid residues QDVSTA (residues 27-31 of SEQ ID NO: 3),
- CDR-L2 amino acid residues WAS (residues 50-52 of SEQ ID NO: 3),
- CDR-L3 amino acid residues QQDYSTPPT (residues 89-97 of SEQ ID NO: 3).
- CDR-H1 amino acid residues GYTFTNYW (residues 26-33 of SEQ ID NO: 4),
- CDR-H2 amino acid residues IDPSDTRT (residues 51-58 of SEQ ID NO: 4),
- CDR-H3 amino acid residues ARQTLYYEALDY (residues 97-108 of SEQ ID NO: 4).
- the invention uses a monoclonal antibody selected from:
- monoclonal antibody #2B 1 monoclonal antibody #2B 1 ;
- antibody derivative refers to an antibody which retains the antigenic specificity of a reference antibody but wherein one or more amino acid residues are (chemically, or biologically) modified to improve its properties. Examples of such chemical modifications include, e.g. by alkylation, PEGylation, acylation, ester or amide formation, and the like.
- a derivative is an antibody as disclosed herein that is modified to contain one or more additional non-pro teinaceous moieties such as water-soluble polymers.
- water-soluble polymers include, but are not limited to, PEG, copolymers of ethylene glycol/propylene glycol, carboxymethylcellulose, dextran and polyvinyl alcohol.
- Derivatives may also be generated to increase or decrease the extent to which the antibody is glycosylated. Addition or deletion of glycosylation sites to an antibody may be conveniently accomplished by altering the amino acid sequence such that one or more glycosylation sites is created or removed. Where the antibody comprises an Fc region, the carbohydrate attached thereto may be altered. Native antibodies produced by mammalian cells typically comprise a branched, biantennary oligosaccharide that is generally attached by an N-linkage to Asn297 of the CH2 domain of the Fc region (see e.g., Wright et al. TIBTECH, 1997, 15:26-32).
- the oligosaccharide may include various carbohydrates, e.g., mannose, N-acetyl glucosamine (GlcNAc), galactose, and sialic acid, as well as a fucose attached to a GlcNAc in the "stem" of the biantennary oligosaccharide structure.
- modifications of the oligosaccharide in an antibody of the invention may be made in order to create antibody variants with certain improved properties.
- antibody variants having a carbohydrate structure that lacks fucose attached (directly or indirectly) to an Fc region.
- the amount of fucose in such antibody may be from 1% to 80%, from 1% to 65%, from 5% to 65% or from 20% to 40%).
- the amount of fucose is determined by calculating the average amount of fucose within the sugar chain at Asn297, relative to the sum of all glycostructures attached to Asn 297 (e. g. complex, hybrid and high mannose structures) as measured by MALDI-TOF mass spectrometry.
- Asn297 refers to the asparagine residue located at about position 297 in the Fc region (Eu numbering of Fc region residues); however, Asn297 may also be located about ⁇ 3 amino acids upstream or downstream of position 297, i.e., between positions 294 and 300, due to minor sequence variations in antibodies.
- Examples of publications related to "defucosylated” or “fucose-deficient" antibody variants include, but are not limited to, Okazaki et al. J. Mol. Biol. 336: 1239- 1249 (2004) and Yamane-Ohnuki N, Satoh M. mAbs. 2009;1:230-236.
- Examples of cell lines capable of producing defucosylated antibodies include Led 3 CHO cells deficient in protein fucosylation (Ripka et al. Arch. Biochem. Biophys. 249:533-545 (1986)), and knockout cell lines, such as alpha- 1 ,6-fucosyltransferase gene, FUT8, knockout CHO cells (see, e.g., Yamane-Ohnuki et al. Biotech. Bioeng. 87: 614 (2004); Kanda, Y. et al., Biotechnol. Bioeng., 94(4):680-688 (2006)).
- cysteine engineered antibodies e.g., "thioMAbs”
- one or more residues of an antibody are substituted with cysteine residues.
- the substituted residues occur at accessible sites of the antibody.
- reactive thiol groups are thereby positioned at accessible sites of the antibody and may be used to conjugate the antibody to other moieties, such as drug moieties or linker-drug moieties, to create an immunoconjugate, as described further herein.
- any one or more of the following residues may be substituted with cysteine: V205 (Kabat numbering) of the light chain; A118 (EU numbering) of the heavy chain; and S400 (EU numbering) of the heavy chain Fc region.
- Cysteine engineered antibodies may be generated as described, e.g., in U.S. Patent No. 7,521,541.
- the term derivative also includes immunoconjugates comprising an anti-sPLA2-GIB antibody as defined above conjugated to one or more heterologous molecule(s), including but not limited to a cytotoxic agent, a detectable moiety such as a fluorescent moiety, a diagnostic radioisotope or an imaging agent; or to a solid support, such as agarose beads or the like.
- cytotoxic agents include, but are not limited to chemotherapeutic agents or drugs, growth inhibitory agents, toxins (e.g., protein toxins, enzymatically active toxins of bacterial, fungal, plant, or animal origin, or fragments thereof), or radioactive isotopes.
- Conjugates of an antibody and cytotoxic agent may be made using a variety of bifunctional protein coupling agents well known by the skilled person.
- the linker may be a "cleavable linker" facilitating release of a cytotoxic drug in the cell.
- an acid-labile linker, peptidase-sensitive linker, photolabile linker, dimethyl linker or disulfide-containing linker (Chari et al., Cancer Res. 52: 127-131 (1992)) may be used.
- the antibodies for use in the invention are typically “isolated”, e.g., have been separated from at least one component of their natural environment.
- the antibodies may be purified to greater e.g., at least 95%, at least 96%; at least 97%, at least 98% or at least 99% purity as determined by, for example, electrophoretic (e.g., SDS-PAGE, isoelectric focusing (IEF), capillary electrophoresis) or chromatographic (e.g., ion exchange or reverse phase HPLC) techniques.
- electrophoretic e.g., SDS-PAGE, isoelectric focusing (IEF), capillary electrophoresis
- chromatographic e.g., ion exchange or reverse phase HPLC
- Preferred antibodies of the invention are essentially neutralizing antibodies, i.e., they are able to at least partially inhibit an activity of PLA2-GIB .
- sPLA2-GIB catalyzes the hydrolysis of the sn-2 fatty acyl bond of phospholipids to release free fatty acids and lysophospholipids.
- Particular antibodies of the invention inhibit an enzymatic activity of sPLA2-GIB, such as the hydrolysis of the sn-2 fatty acyl bond of phospholipids. Methods for testing such a property are disclosed in detail in the experimental section.
- Particular antibodies for use in the invention inhibit binding of sPLA2-GIB to a substrate thereof.
- Further particular antibodies for use in the invention inhibit sPLA2-GIB -mediated inhibition of IF-7-induced phospho-STAT5 nuclear translocation in CD4 T cells. Methods for testing such a property are disclosed in detail in the experimental section.
- the neutralizing activity of the antibody or derivative can be determined in vitro or in vivo using e.g., binding or biological assays, such as tests as described in the experimental section. Inhibition/neutralization may be complete or partial. In particular, the antibodies may inhibit 10% or more of the tested activity, preferably 20% or more, 30% or more, 40% or more, 50% or more.
- the antibodies are IgG, e.g., gGl, IgG2, IgG3 or IgG4.
- the antibodies or derivatives may be isolated and preserved using conventional methods and media. They may be lyophilized. They may also be frozen.
- the PLA2-GIB inhibitor is or comprises or consists of a nucleic acid molecule encoding an anti-PLA2-GIB antibody, or a light or heavy chain thereof, or a variable domain thereof, or a nucleic acid complementary to said encoding sequence.
- the nucleic acid can be DNA (cDNA or gDNA), RNA, or a mixture thereof. It can be in single stranded form or in duplex form or a mixture of the two. It can comprise modified nucleotides, comprising for example a modified bond, a modified purine or pyrimidine base, or a modified sugar. It can be prepared by any method known to one skilled in the art, including chemical synthesis, recombination, and mutagenesis.
- the nucleic acid according to the invention may be deduced from the sequence of the antibody according to the invention and codon usage may be adapted according to the host cell in which the nucleic acid shall be transcribed. These steps may be carried out according to methods well known to one of skill in the art and some of which are described in the reference manual Sambrook et al. (Sambrook J, Russell D (2001) Molecular cloning: a laboratory manual, Third Edition Cold Spring Harbor).
- the nucleic acid may encode an amino acid sequence comprising the light chain and/or an amino acid sequence comprising the heavy chain of the antibody, or may be complementary to such encoding sequence.
- the present invention further provides a vector comprising a nucleic acid of the invention.
- the vector may comprise several nucleic acids of the invention.
- the vector may comprise a nucleic acid of the invention operably linked to a regulatory region, i.e. a region comprising one or more control sequences.
- the vector may comprise several nucleic acids of the invention operably linked to several regulatory regions.
- control sequences means nucleic acid sequences necessary for expression of a coding region. Control sequences may be endogenous or heterologous. Well-known control sequences and currently used by the person skilled in the art will be preferred. Such control sequences include, but are not limited to, promoter, signal peptide sequence and transcription terminator.
- operably linked means a configuration in which a control sequence is placed at an appropriate position relative to a coding sequence, in such a way that the control sequence directs expression of the coding region.
- the present invention further relates to the use of a nucleic acid or vector according to the invention to transform, transfect or transduce a host cell.
- the present invention also provides a host cell comprising one or several nucleic acids of the invention and/or one or several vectors of the invention.
- host cell also encompasses any progeny of a parent host cell that is not identical to the parent host cell due to mutations that occur during replication.
- Suitable host cells for cloning or expression of antibody-encoding vectors include prokaryotic or eukaryotic cells such as bacteria, yeasts, insect cells, mammalian cells, etc.
- Inhibitory Nucleic acids include prokaryotic or eukaryotic cells such as bacteria, yeasts, insect cells, mammalian cells, etc.
- the PLA2-GIB inhibitor is an inhibitory nucleic acid, i.e., any nucleic acid molecule which inhibits PLA2-GIB gene or protein expression.
- Preferred inhibitory nucleic acids include antisense nucleic acids, short interfering RNAs (siRNAs), small hairpin RNAs (shRNA), microRNAs, aptamers, or ribozymes.
- the inhibitory nucleic acid is a small interfering RNA that prevents translation of PLA2-GIB mRNA.
- the inhibitory nucleic acid is an antisense oligonucleotide that prevents translation of PLA2-GIB mRNA.
- the inhibitory nucleic acid is a small hairpin RNA that prevents translation of PLA2-GIB mRNA.
- siRNA comprise a sense nucleic acid sequence and an anti-sense nucleic acid sequence of the polynucleotide of interest.
- siRNA are constructed such that a single transcript (double stranded RNA) have both the sense and complementary antisense sequences from the target gene.
- the nucleotide sequence of siRNAs may be designed using an siRNA design computer program available from, for example, the Ambion website on the world wide web.
- the length of the antisense oligonucleotide or siRNAs is less than or equal to 10 nucleotides. In some embodiments, the length of the antisense oligonucleotides and siRNAs is as long as the naturally occurring transcript. In some embodiments, the antisense oligonucleotides and siRNAs have 18-30 nucleotides. In some embodiments, the antisense oligonucleotides and siRNAs are less than 25 nucleotides in length.
- Preferred inhibitory nucleic acid molecules comprise a domain having a nucleotide sequence that is perfectly complementary to a region of a PLA2-GIB gene or RNA. Such a domain contains typically from 4 to 20 nucleotides, allowing specific hybridization and optimal inhibition of the gene transcription or RNA translation.
- the sequence of the inhibitory nucleic acids may be derived directly from the sequence of a gene encoding PLA2-GIB. Alternatively, or in addition, inhibitory nucleic acids may hybridize to a regulatory element in a PLA2-GIB gene or RNA, such as a promoter, a splicing site, etc., and prevent effective regulation thereof.
- inhibitory nucleic acid molecules of the present invention include isolated single strand nucleic acid molecules consisting of from 10 to 50 consecutive nucleotides of a sequence encoding SEQ ID NO: 210.
- inhibitory nucleic acid molecules of the invention are antisense nucleic acids consisting of the following nucleotide sequence or the perfectly complementary strand thereof:
- the PLA2-GIB inhibitor is a peptide or small drug that inhibits the activity of PLA2-GIB.
- the peptide or small drug is typically a molecule that selectively binds PLA2-GIB, or a substrate of PLA2-GIB, or a co-factor of PLA2-GIB, or a degradation product or metabolite of PLA2-GIB pathway.
- Peptides preferably contain from 3 to 20 amino acid residues, and their sequence may be identical to a domain of PLA2-GIB (bait peptide) or to a domain of a PLA2-GIB substrate, co-factor, degradation product or metabolite.
- Preferred peptides of the invention contain from 4 to 30 consecutive amino acid residues of SEQ ID NO: 210 (or of a corresponding sequence of a natural variant of SEQ ID NO: 210).
- Most preferred peptides of the invention comprise from 5 to 25 consecutive amino acid residues of SEQ ID NO: 210 (or of a corresponding sequence of a natural variant of SEQ ID NO: 210) and further comprise at least one of the following amino acid residues of SEQ ID NO: 210 (or of a corresponding sequence of a natural variant of SEQ ID NO: 210): amino acid 3, amino acid 6, amino acid 7, amino acid 10, amino acid 70, amino acid 121, amino acid 50, amino acid 52, amino acid 54, amino acid 71, amino acid 75, amino acid 77, amino acid 79, amino acid 80, or a combination thereof.
- Specific examples of peptides of the invention are peptides of less than 25 amino acids comprising anyone of the following sequences:
- peptides for use in the invention include a pentapeptide as disclosed in WO20 17/060405, incorporated therein by reference.
- the compound is a cyclic peptide selected from FLSYK (SEQ ID NO: 231), FLSYR (SEQ ID NO: 232) and (2NapA)LS(2NapA)R (SEQ ID NO: 233).
- the peptides of the invention can comprise peptide, non-peptide and/or modified peptide bonds.
- the peptides comprise at least one peptidomimetic bond selected from intercalation of a methylene (-CH2-) or phosphate (-PO2-) group, secondary amine (-NH-) or oxygen (-0-), alpha-azapeptides, alpha-alkylpeptides, N- alkylpeptides, phosphonamidates, depsipeptides, hydroxymethylenes, hydroxyethylenes, dihydroxyethylenes, hydroxyethylamines, retro-inverso peptides, methyleneoxy, cetomethylene, esters, phosphinates, phosphinics, or phosphonamides.
- the peptides may comprise a protected N-ter and/or C-ter function, for example, by acylation, and/or amidation and/or esterification.
- the peptides of the invention may be produced by techniques known per se in the art such as chemical, biological, and/or genetic synthesis.
- Preferred small drugs are hydrocarbon compounds that selectively bind PLA2-GIB.
- small drugs include indole compounds, such as those disclosed in W02017/037041, incorporated therein by reference.
- the compound is 3-(2-amino-l,2-dioxoethyl)-2-ethyl-l-(phenylmethyl)-lH-indol-4- yl)oxy)acetic acid or a pharmaceutically acceptable salt, hydrate, or prodrug thereof, such as a sodium salt thereof (Varespladib).
- Small drugs and peptides are preferably obtainable by a method comprising: (i) contacting a test compound with PLA2-GIB or a fragment thereof, (ii) selecting a test compound which binds PLA2-GIB or said fragment thereof, and (iii) selecting a compound of (ii) which inhibits an activity of PLA2-GIB.
- a method comprising: (i) contacting a test compound with PLA2-GIB or a fragment thereof, (ii) selecting a test compound which binds PLA2-GIB or said fragment thereof, and (iii) selecting a compound of (ii) which inhibits an activity of PLA2-GIB.
- Small drugs and peptides are also obtainable by a method comprising: (i) contacting a test compound with a PLA2-GIB substrate, co-factor, or degradation product, or a fragment thereof, (ii) selecting a test compound which binds to said PLA2-GIB substrate, co-factor, or degradation product, or a fragment thereof, and (iii) selecting a compound of (ii) which inhibits an activity of PLA2-GIB.
- a method represents a particular object of the invention.
- the PLA2-GIB inhibitor is a PLA2-GIB antigen.
- the subject produces antibodies (or cells) which inhibit PLA2-GIB .
- injection(s) of a PLA2-GIB antigen e.g., an immunogenic PLA2-GIB essentially devoid of biological activity
- a PLA2-GIB antigen e.g., an immunogenic PLA2-GIB essentially devoid of biological activity
- An object of the invention thus resides in a method of treating a solid cancer in a subject having solid cancer, comprising administering to the subject a PLA2-GIB antigen.
- a further object of the invention relates to a PLA2-GIB antigen for use to treat a solid cancer in a subject in need thereof.
- the PLA2-GIB antigen is an inactivated immunogenic molecule that induces an immune response against PLA2-GIB in a subject.
- Inactivation may be obtained e.g., by chemically or physically altering PLA2-GIB or by mutating or truncating the protein, or both; and immunogenicity may be obtained as a result of the inactivation and/or by further conjugating the protein to a suitable carrier or hapten, such as KLH, HSA, polylysine, a viral anatoxin, or the like, and/or by polymerization, or the like.
- the antigen may thus be chemically or physically modified, e.g., to improve its immunogenicity .
- the PLA2-GIB antigen comprises PLA2-GIB or an epitope- containing fragment or mimotope thereof.
- the PLA2-GIB antigen comprises a full length PLA2-GIB protein.
- the PLA2-GIB antigen comprises a protein comprising SEQ ID NO: 210, or a sequence having at least 90% identity to SEQ ID NO: 210.
- the PLA2-GIB antigen comprises a fragment of a PLA2- GIB protein comprising at least 6 consecutive amino acid residues and containing an immunogenic epitope, or a mimotope thereof. In a preferred embodiment, the PLA2-GIB antigen comprises at least from 6 to 20 amino acid residues. Preferred peptides of the invention contain from 4 to 30 consecutive amino acid residues of SEQ ID NO: 210 (or of a corresponding sequence of a natural variant of SEQ ID NO: 210).
- Most preferred peptides of the invention comprise from 5 to 25 consecutive amino acid residues of SEQ ID NO: 210 (or of a corresponding sequence of a natural variant of SEQ ID NO: 210) and further comprise at least one of the following amino acid residues of SEQ ID NO: 210 (or of a corresponding sequence of a natural variant of SEQ ID NO: 210): amino acid 3, 6, 7, 10, 70, 121, 50, 52, 54, 71, 75, 77, 79, 80, or a combination thereof.
- Specific examples of such antigens are the peptides of SEQ ID NO: 222-230 as disclosed above.
- the PLA2-GIB antigen may be in various forms such as in free form, polymerized, chemically or physically modified, and/or coupled (i.e., linked) to a carrier molecule. Coupling to a carrier may increase the immunogenicity and (further) suppress the biological activity of the PLA2-GIB polypeptide.
- the carrier molecule may be any carrier molecule or protein conventionally used in immunology such as for instance KLH (Keyhole limpet hemocyanin), ovalbumin, bovine serum albumin (BSA), a viral or bacterial anatoxin such as toxoid tetanos, toxoid diphteric B cholera toxin, mutants thereof such as diphtheria toxin CRM 197, an outer membrane vesicle protein, a polylysine molecule, or a virus like particle (VLP).
- the carrier is KLH or CRM 197 or a VLP.
- Coupling of PLA2-GIB to a carrier may be performed by covalent chemistry using linking chemical groups or reactions, such as for instance glutaraldehyde, biotin, etc.
- the conjugate or the PLA2-GIB protein or fragment or mimotope is submitted to treatment with formaldehyde in order to complete inactivation of PLA2-GIB.
- the PLA2-GIB antigen comprises a full length PLA2-GIB protein, optionally coupled to a carrier protein.
- the PLA2- GIB antigen comprises a protein comprising SEQ ID NO: 210, or a sequence having at least 90% identity to SEQ ID NO: 210, coupled to a carrier protein.
- the PLA2-GIB antigen comprises an immunogenic peptide or mimotope of PLA2-GIB, optionally coupled to a carrier protein.
- the PLA2-GIB antigen comprises a polypeptide of at least 10 amino acids long comprising at least one of the following amino acid residues of SEQ ID NO: 210 (or of a corresponding sequence of a natural variant of SEQ ID NO: 210): amino acid 70, amino acid 121, amino acid 50, amino acid 52, amino acid 54, amino acid 71, or a combination thereof, optionally coupled to a carrier molecule.
- the immunogenicity of the PLA2-GIB antigen may be tested by various methods, such as by immunization of a non-human animal grafted with human immune cells, followed by verification of the presence of antibodies, or by sandwich ELISA using human or humanized antibodies. The lack of biological activity may be verified by any of the activity tests described in the application.
- the PLA2-GIB antigen has less than 20%, more preferably less than 15%, 10%, 5% or even 1% of the activity of a wild-type PLA2-GIB protein in an in vitro method of (i) induction of formation of membrane microdomains (MMD) in CD4 T cells or (ii) in rendering CD4 T cells refractory to IL-2 signaling or refractory to IL-7 signaling.
- MMD membrane microdomains
- Such molecules and conjugates and vaccines represent potent agents for use to immunize subjects, thereby causing a sustained PLA2-GIB inhibition. Upon repetition, such methods can be used to cause a permanent PLA2-GIB inhibition.
- the invention also relates to an inhibitor of a PLA2-GIB cofactor, for use for treating a Group IV RNA vims infection, particularly a coronavirus infection in a subject in need thereof.
- the invention also relates to the use of an inhibitor of a PLA2-GIB cofactor, for the manufacture of a medicament for treating a Group IV RNA vims infection, particularly a coronavirus infection in a subject in need thereof.
- SARS-Cov-2 can act as (or produce) a cofactor of PLA2-GIB that, in combination with PLA2-GIB, leads to CD4 T cell inactivation.
- cofactor of PLA2-GIB designates any molecule or agent which potentiates or amplifies or mediates an effect of PLA2-GIB, particularly an effect of PLA2-GIB on CD4 T cells.
- Preferred cofactors are molecules which can sensitize CD4 T cells to inactivation by low concentrations of PLA2-GIB.
- the PLA2-GIB cofactor is a ligand of gC1qR.
- the PLA2-GIB cofactor is an agonist of gC1qR, e.g., can induce signaling through gC1qR, more particularly can induce gC1qR- mediated exocytosis.
- the inventors have identified various proteins which can act as cofactor of PLA2-GIB, as listed in Table 3. In particular, such proteins include ORFlab, the Spike protein, ORF3, E protein, ORF7b, ORF8, N protein and M protein.
- Each of said proteins thus represents a relevant target for inhibition according to the invention.
- the inventors also identified various peptides from said proteins which can bind gC1qR, as listed in Table 3. Each of said peptides represents an object of the invention.
- Each of said proteins and peptides can act as PLA2-GIB cofactors in SARS-Cov2 infected patients.
- Each of said peptides represents a strong inhibitor (or be used to construct a strong inhibitor) for use according to the invention
- Each of said proteins or peptides can be used to produce relevant inhibitors, such as antibodies or derivatives thereof, or immunogens for vaccination.
- Each of said proteins and peptides can be used as a reagent to detect the presence of anti- Cov-2 antibodies in a sample, particularly of a protective immunity.
- the PLA2-GIB cofactor is a protein selected from ORFlab, the Spike protein, ORF3, E protein, ORF7b, ORF8, N protein and M protein of SARS-Cov-2, or a gC1qR-binding element of such proteins, such as listed in Table 3. More particularly, the cofactor may be any protein comprising anyone of SEQ ID NOs: 2-209, or any fragment or mimotope thereof.
- fragment in relation to such cofactors, designates preferably a fragment containing a gC1qR-binding element of such a protein, and/or a fragment retaining a capacity of binding gC1qR.
- Preferred fragments contain at least 5 consecutive amino acid residues, typically between 5 and 100, between 5-80, between 5-60, between 5-50, between 5-40, between 5-30, between 5-25, or between 5-20, or even less.
- the PLA2-GIB cofactor is SARS-Cov2 spike (or S) protein, or a fragment or mimotope thereof.
- the amino acid sequence of a Spike protein is depicted as SEQ ID NO: 209. Such sequence is also available as GenBank reference QHD43416.1.
- the PLA2-GIB cofactor is a protein or peptide comprising or consisting of the amino acid sequence of any one of SEQ ID NOs: 156-185 and 209, or a fragment or mimotope thereof.
- the PLA2-GIB cofactor is SARS-Cov2 ORFlab protein, or a fragment or mimotope thereof.
- the PLA2-GIB cofactor is a protein or peptide comprising or consisting of the amino acid sequence of any one of SEQ ID NOs: 2-155, or a fragment or mimotope thereof.
- the PLA2-GIB cofactor is SARS-Cov2 ORF3 protein, or a fragment or mimotope thereof.
- the PLA2-GIB cofactor is a protein or peptide comprising or consisting of the amino acid sequence of any one of SEQ ID NOs: 186-193, or a fragment or mimotope thereof.
- the PLA2-GIB cofactor is SARS-Cov2 protein E, or a fragment or mimotope thereof.
- the PLA2-GIB cofactor is a protein or peptide comprising or consisting of the amino acid sequence of SEQ ID NO: 194, or a fragment or mimotope thereof.
- the PLA2-GIB cofactor is SARS-Cov2 protein M, or a fragment or mimotope thereof.
- the PLA2-GIB cofactor is a protein or peptide comprising or consisting of the amino acid sequence of any one of SEQ ID NOs: 195-202, or a fragment or mimotope thereof.
- the PLA2-GIB cofactor is SARS-Cov2 ORF7b protein, or a fragment or mimotope thereof.
- the PLA2-GIB cofactor is a protein or peptide comprising or consisting of the amino acid sequence of SEQ ID NO: 203, or a fragment or mimotope thereof.
- the PLA2-GIB cofactor is SARS-Cov2 ORF8 protein, or a fragment or mimotope thereof.
- the PLA2-GIB cofactor is a protein or peptide comprising or consisting of the amino acid sequence of SEQ ID NO: 204, or a fragment or mimotope thereof.
- the PLA2-GIB cofactor is SARS-Cov2 protein N, or a fragment or mimotope thereof.
- the PLA2-GIB cofactor is a protein or peptide comprising or consisting of the amino acid sequence of any one of SEQ ID NOs: 205-208, or a fragment or mimotope thereof.
- the invention relates to the treatment of a Group IV RNA virus infection, particularly a coronavirus infection, more particularly SARS-Cov2 infection, in subjects by modulating such a cofactor effect.
- the invention thus provides methods and compositions for treating diseased subjects and/or for restoring/enhancing CD4 T cell activity in subjects using an inhibitor of a PLA2-GIB cofactor.
- the term “inhibitor” of a cofactor designates any molecule or treatment which causes (directly or indirectly) an inhibition of the expression or a function of the cofactor, e.g., cofactor binding to gC1qR or cofactor ability to sensitize CD4 T cells to PLA2-GIB.
- Inhibiting the cofactor designates preferably reducing by at least 10%, 20%, 30%, 40%, 50%, 60%, 70%, 80% or more the expression or a function of the cofactor, as well as completely blocking or suppressing said expression or function. Depending on the situation, the inhibition may be transient, sustained or permanent.
- an inhibitor of the cofactor is a gC1qR inhibitor.
- cofactors bind gC1qR as a target receptor. Blocking or reducing or preventing binding of the cofactor to gC1qR using gC1qR inhibitors can affect the cofactor effect.
- the term “gC1qR inhibitor” designates any molecule or treatment which causes (directly or indirectly) an inhibition of a function of gC1qR, e.g., gC1qR-mediated exocytosis.
- gC1qR designates the receptor for complement Clq at the surface of cells, particularly of CD4 T cells, especially the human form of said receptor.
- gC1qR is also known as Clq binding protein (C1QBP), ASF/SF2-associated protein p32 (SF2P32); Glycoprotein gClqBP; Hyaluronan-binding protein 1 (HABP1); Mitochondrial matrix protein p32; gClq-R protein; p33; ClqBP and GC1QBP.
- C1QBP Clq binding protein
- SF2P32 ASF/SF2-associated protein p32
- Glycoprotein gClqBP Glycoprotein gClqBP
- HABP1 Hyaluronan-binding protein 1
- Mitochondrial matrix protein p32 gClq-R protein
- p33 ClqBP and GC1QBP.
- the amino acid sequence of the receptor was disclosed in the art.
- An exemplary amino acid sequence of human gC1qR is reproduced below (SEQ ID NO: 211):
- gC1qR designates any receptor of SEQ ID NO: 211 (accession number UniProtKB/Swiss-Prot: Q07021.1) above, as well as processed forms and variants thereof. Variants include naturally-occurring variants having e.g., at least 90% amino acid sequence identity to SEQ ID NO: 211.
- gC1qR Upon binding of a cofactor, gC1qR triggers a signaling pathway that results in exocytosis of intracellular vesicles. Without being bound by theory, it is believed that the fusion of these vesicles with the cytoplasmic membrane could change the lipid composition and increase sPLA2-GIB activity on CD4 T cells membrane, resulting in an inhibition of phosphoSTAT5 signaling.
- the fusion of these vesicles with plasma membrane can change the lipid composition and cause sPLA2-GIB activity on CD4 T cells membranes.
- membrane fluidity is increased and cytokines receptors are aggregated in abnormal membrane domain, resulting in a dramatic decrease of cytokine signaling, and anergy of CD4 T cells.
- gC1qR inhibitor thus includes any molecule which binds to gC1qR, or to a partner of gC1qR, and inhibits a function of gC1qR, such as gC1qR- mediated exocytosis.
- the cofactor inhibitor is a molecule which directly inhibits an activity of the cofactor, e.g., which binds the cofactor and/or inhibits binding of the cofactor to its receptor.
- cofactor inhibitors include, for instance, antibodies and variants thereof, synthetic specific ligands, peptides, small drugs, or inhibitory nucleic acids.
- a cofactor inhibitor is an antibody or an antibody variant/fragment having essentially the same antigen specificity, or a nucleic acid encoding such an antibody or variant/fragment.
- the antibody may bind a cofactor, or gC1qR, or a partner of gC1qR, or a gC1qR-binding element thereof, and preferably inhibits a function of the cognate antigen (e.g., gC1qR or the cofactor).
- Antibodies can be synthetic, monoclonal, or polyclonal and can be made by techniques well known per se in the art.
- antibodies is meant to include polyclonal antibodies, monoclonal antibodies, fragments thereof, such as F(ab')2 and Fab fragments, single-chain variable fragments (scFvs), single-domain antibody fragments (VHHs or Nanobodies), bivalent antibody fragments (diabodies), as well as any recombinantly and synthetically produced binding partners, human antibodies or humanized antibodies.
- Antibodies are defined to be specifically binding, preferably if they bind to the cognate antigen with a Ka of greater than or equal to about 10 7 M-l. Affinities of antibodies can be readily determined using conventional techniques, for example those described by Scatchard et al., Ann. N.Y. Acad. Sci., 51:660 (1949).
- Polyclonal antibodies can be readily generated from a variety of sources, for example, horses, cows, donkeys, goats, sheeps, dogs, chickens, rabbits, mice, hamsters, or rats, using procedures that are well known in the art.
- a purified immunogen optionally appropriately conjugated, is administered to the host animal typically through parenteral injection.
- the immunogenicity of immunogen can be enhanced through the use of an adjuvant, for example, Freund's complete or incomplete adjuvant.
- small samples of serum are collected and tested for reactivity to the antigen polypeptide.
- Examples of various assays useful for such determination include those described in Antibodies: A Laboratory Manual, Harlow and Lane (eds.), Cold Spring Harbor Laboratory Press, 1988; as well as procedures, such as countercurrent immuno-electrophoresis (CIEP), radioimmunoassay, radio-immunoprecipitation, enzyme-linked immunosorbent assays (ELISA), dot blot assays, and sandwich assays. See U.S. Pat. Nos. 4,376,110 and 4,486,530.
- Monoclonal antibodies can be readily prepared using well known procedures. See, for example, the procedures described in U.S. Pat. Nos. RE 32,011, 4,902,614, 4,543,439, and 4,411,993; Monoclonal Antibodies, Hybridomas: A New Dimension in Biological Analyses, Plenum Press, Kennett, McKeam, and Bechtol (eds.), 1980. Lor example, the host animals, such as mice, can be injected intraperitoneally at least once and preferably at least twice at about 3 week intervals with isolated and purified immunogen, optionally in the presence of adjuvant. Mouse sera are then assayed by conventional dot blot technique or antibody capture (ABC) to determine which animal is best to fuse.
- ABSC antibody capture
- mice are given an intravenous boost of protein or peptide.
- Mice are later sacrificed and spleen cells fused with commercially available myeloma cells, such as Ag8.653 (ATCC), following established protocols. Briefly, the myeloma cells are washed several times in media and fused to mouse spleen cells at a ratio of about three spleen cells to one myeloma cell.
- the fusing agent can be any suitable agent used in the art, for example, polyethylene glycol (PEG). Fusion is plated out into plates containing media that allows for the selective growth of the fused cells. The fused cells can then be allowed to grow for approximately eight days.
- Monoclonal antibodies may also be produced using alternative techniques, such as those described by Alting-Mees et al. , "Monoclonal Antibody Expression Libraries: A Rapid Alternative to Hybridomas", Strategies in Molecular Biology 3:1-9 (1990), which is incorporated herein by reference.
- binding partners can be constructed using recombinant DNA techniques to incorporate the variable regions of a gene that encodes a specific binding antibody. Such a technique is described in Larrick et ah, Biotechnology, 7:394 (1989).
- Antigen-binding fragments of antibodies which can be produced by conventional techniques, are also encompassed by the present invention.
- fragments include, but are not limited to, Fab and F(ab')2 fragments.
- Antibody fragments and derivatives produced by genetic engineering techniques are also provided.
- the monoclonal antibodies of the invention also include chimeric antibodies, e.g., humanized versions of murine monoclonal antibodies.
- humanized antibodies can be prepared by known techniques, and offer the advantage of reduced immunogenicity when the antibodies are administered to humans.
- a humanized monoclonal antibody comprises the variable region of a murine antibody (or just the antigen binding site thereof) and a constant region derived from a human antibody.
- a humanized antibody fragment can comprise the antigen binding site of a murine monoclonal antibody and a variable region fragment (lacking the antigen-binding site) derived from a human antibody.
- Procedures for the production of chimeric and further engineered monoclonal antibodies include those described in Riechmann et al.
- Such chimeric and humanized monoclonal antibodies can be produced by genetic engineering using standard DNA techniques known in the art, for example using methods described in Robinson et al. International Publication No. WO 87/02671; Akira, et al. European Patent Application 0184187; Taniguchi, M., European Patent Application 0171496; Morrison et al. European Patent Application 0173494; Neuberger et al. PCT International Publication No. WO 86/01533; Cabilly et al. U.S. Pat. No. 4,816,567; Cabilly et al.
- antibodies In connection with synthetic and semi-synthetic antibodies, such terms are intended to cover but are not limited to antibody fragments, isotype switched antibodies, humanized antibodies (e.g., mouse-human, human-mouse), hybrids, antibodies having plural specificities, and fully synthetic antibody-like molecules.
- Human monoclonal antibodies can also be prepared by constructing a combinatorial immunoglobulin library, such as a Fab phage display library or a scFv phage display library, using immunoglobulin light chain and heavy chain cDNAs prepared from mRNA derived from lymphocytes of a subject. See, e.g., McCafferty et al. PCT publication WO 92/01047; Marks et al. (1991) J. Mol. Biol. 222:581 597; and Griffths et al. (1993) EMBO J 12:725 734.
- a combinatorial library of antibody variable regions can be generated by mutating a known human antibody.
- variable region of a human antibody known to bind gC1qR can be mutated by, for example, using randomly altered mutagenized oligonucleotides, to generate a library of mutated variable regions which can then be screened to bind to gC1qR.
- Methods of inducing random mutagenesis within the CDR regions of immunoglobin heavy and/or light chains, methods of crossing randomized heavy and light chains to form pairings and screening methods can be found in, for example, Barbas et al. PCT publication WO 96/07754; Barbas et al. (1992) Proc. Nat'l Acad. Sci. USA 89:44574461.
- Antibodies of the invention may be directed against gC1qR, a gC1qR ligand, or a gC1qR partner, and cause an inhibition of signaling mediated by gC1qR.
- an immunogen may be used comprising gC1qR, a gC1qR ligand, or a gC1qR partner, or a fragment, variant, or fusion molecule thereof.
- Particular antibodies of the invention bind a gC1qR epitope, and/or have been generated by immunization with a polypeptide comprising a gC1qR epitope, selected from the mature gC1qR protein or a fragment of gC1qR comprising at least 8 consecutive amino acid residues thereof.
- Preferred anti-gC1qR antibodies of the invention bind an epitope of a ligand-binding site within gC1qR, thereby interfering with binding of the ligand.
- the antibodies bind an epitope comprised between amino acid residues 76-282 of SEQ ID NO: 211, which contain the gC1qR ligand bind site.
- Clq binding to gC1qR can involve at least three different motifs on gC1qR, namely: amino acid residues 75-96, 190-202 and 144-162 (by reference to SEQ ID NO: 211).
- HCV core protein binding to gC1qR can involve at least two different motifs on gC1qR, namely: amino acid residues 144-148 and 196-202 (by reference to SEQ ID NO: 211).
- HIV gp41 binding to gC1qR can involve at least amino acid residues 174- 180 on gC1qR (by reference to SEQ ID NO: 211).
- an antibody which binds an epitope containing at least one amino acid residue contained in one of said epitopes or close to one of said epitopes.
- antibodies include antibody 60.11, which binds to residues 75-96 of gC1qR; as well as antibody 74.5.2, which binds to an epitope with the residues 204 to 218.
- Preferred gC1qR inhibitors are therefore monoclonal antibodies against gC1qR, more preferably against an epitope of gC1qR located within amino acid residues 76-282 of the protein (by reference to SEQ ID NO: 211), even more preferably an epitope containing an amino acid residue selected from amino acids 75-96, 144-162, 174-180, and 190- 210.
- Preferred antibodies are neutralizing (or antagonist) antibodies, i.e., they prevent or inhibit or reduce binding of a natural ligand to the receptor and/or signaling through the receptor.
- Other particular inhibitors of the invention are antibodies that bind a PLA2-GIB cofactor and/or have been generated by immunization with a PLA2-GIB cofactor or a fragment or variant thereof, and preferably inhibit at least partially an activity of such cofactor, preferably the binding of such a cofactor to gC1qR.
- antibodies of the invention are polyclonal antibodies or monoclonal antibodies, or variants thereof, which bind a protein selected from the proteins listed in Table 3, and inhibit at least partially the binding of said protein to gC1qR.
- Preferred antibodies of the invention are polyclonal antibodies or monoclonal antibodies, or variants thereof, which bind a protein selected from the proteins listed in Table 3, and inhibit at least partially the binding of said protein to gC1qR.
- the inhibitor is an antibody or a variant thereof that binds SARS-Cov-2 spike protein.
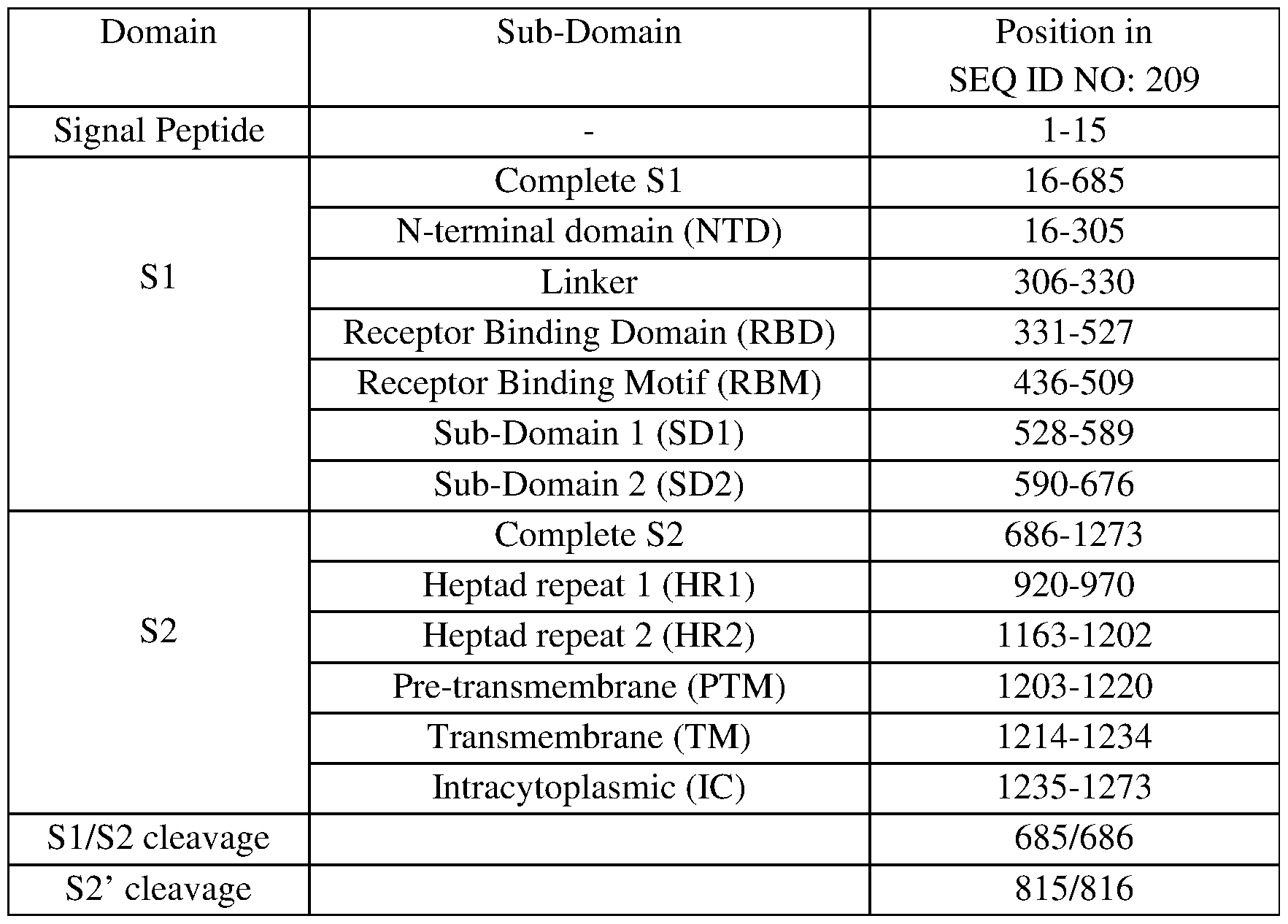
- the antibody binds an epitope comprising an amino acid residue contained in the NTD domain or in the RBD domain of the spike protein (see example 3 for domain position).
- Most preferred antibodies bind an epitope comprising an amino acid residue located in the RBD (amino acid residues 331-527 of SEQ ID NO: 209), or even in the RBM (amino acid residues 436-509 of SEQ ID NO: 209).
- Particular antibodies bind an epitope comprising any amino acid residue of the spike located in anyone of SEQ ID Nos: 156-185.
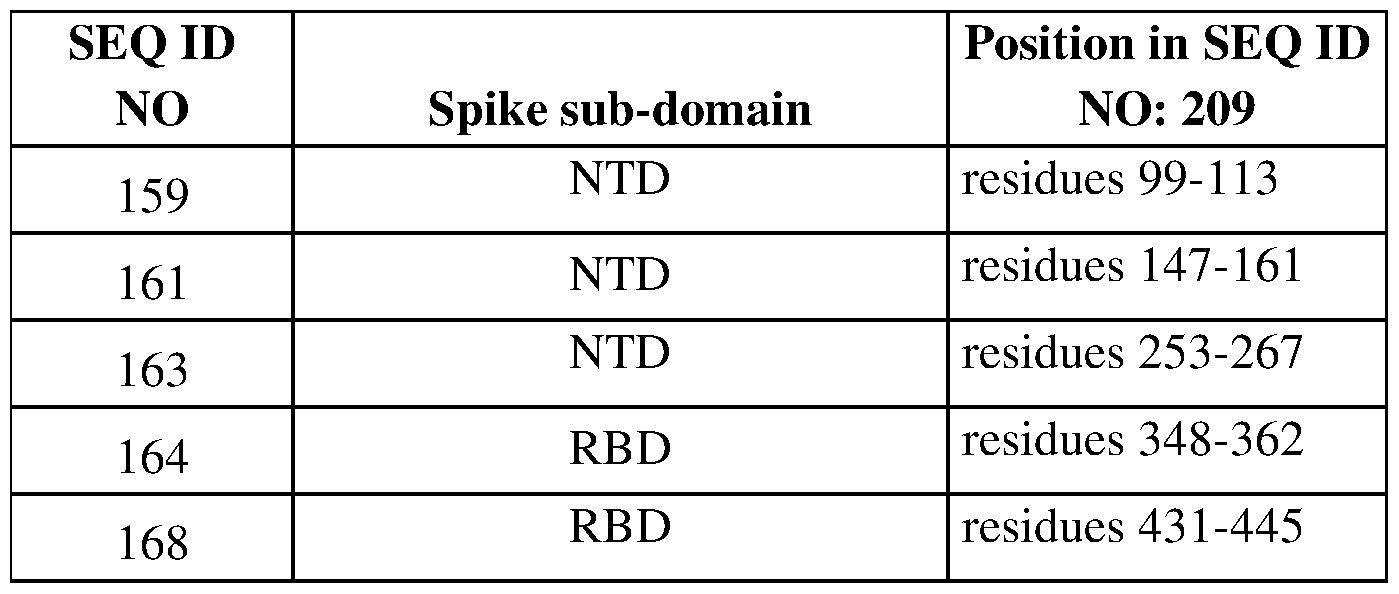
- Particularly preferred antibodies bind an epitope containing at least one amino acid residues of Spike protein located at any of the following positions (by reference to SEQ ID NO: 209): 99-113; 147-161; 253-267; 348-445, 348-362, 431-445.
- Particular antibodies bind an epitope comprising at least one of the following amino acid residues of spike protein (by reference to SEQ ID NO: 209): 431, 432, 433, 434, 435, 436, 437, 438, 438, 440, 441, 442, 443, 444, 445.
- Particular antibodies bind an epitope comprising at least one of the following amino acid residues of spike protein (by reference to SEQ ID NO: 209): 348, 349, 350, 351, 352, 353, 354, 355, 356, 357, 358, 359, 360, 361, 362.
- the inhibitor is an antibody or a variant thereof that binds SARS-Cov-2 ORFlab protein.
- the antibody binds an epitope comprising an amino acid residue contained in anyone of SEQ ID Nos: 2-155.
- the inhibitor is an antibody or a variant thereof that binds SARS-Cov-2 ORF3 protein.
- the antibody binds an epitope comprising an amino acid residue contained in anyone of SEQ ID Nos: 186- 193.
- the inhibitor is an antibody or a variant thereof that binds SARS-Cov-2 protein E. In a further particular embodiment, the antibody binds an epitope comprising an amino acid residue contained in SEQ ID No: 194. In another particular embodiment, the inhibitor is an antibody or a variant thereof that binds SARS-Cov-2 protein M. In a further particular embodiment, the antibody binds an epitope comprising an amino acid residue contained in anyone of SEQ ID Nos: 195-202.
- the inhibitor is an antibody or a variant thereof that binds SARS-Cov-2 protein ORF7b.
- the antibody binds an epitope comprising an amino acid residue contained in SEQ ID No: 203.
- the inhibitor is an antibody or a variant thereof that binds SARS-Cov-2 protein ORF8.
- the antibody binds an epitope comprising an amino acid residue contained in SEQ ID No: 204.
- the inhibitor is an antibody or a variant thereof that binds SARS-Cov-2 protein N.
- the antibody binds an epitope comprising an amino acid residue contained in anyone of SEQ ID Nos: 205-208.
- the cofactor inhibitor is an inhibitory nucleic acid.
- Preferred inhibitory nucleic acids include aptamers which are designed to bind the cofactor, or gC1qR, or a partner of gC1qR, and to inhibit a function thereof.
- the cofactor inhibitor is a peptide that inhibits a function of the cofactor.
- the peptide is typically a molecule that selectively binds a cofactor, a gC1qR, or a partner of gC1qR.
- Peptides preferably contain from 4 to 40 amino acid residues, and their sequence may be identical to a domain of gC1qR or to a domain of a cofactor (bait peptide), or their sequence may contain a variation as compared to the sequence of a domain of gC1qR or to a domain of a cofactor (peptide antagonist).
- Preferred peptides of the invention contain from 4 to 40 consecutive amino acid residues of SEQ ID NO: 211 (gC1qR) or of protein selected from ORFlab, Spike, ORF3, E, M, ORF7b, ORF8 or N. Examples of such peptides are or comprise the amino acid sequence of anyone of SEQ ID Nos: 2-208. Particularly preferred peptides of the invention contain from 4 to 40 consecutive amino acid residues of the spike protein of SEQ ID NO: 209, such as peptides consisting of or comprising anyone of SEQ ID NOs: 156-185.
- Peptides of the invention may be modified to increase their properties.
- the modification may consist of one or more amino acid substitution. Examples of such substitution includes, without limitation, replacement of a charged or reactive amino acid residue by a more neutral residue such as alanine, or conversely.
- the modification may alternatively (or in addition) consist of a chemical modification, such as addition of a chemical group to one (or both) ends of the peptide, or to a lateral chain thereof, or to a peptide bond.
- the peptides of the invention can comprise peptide, non-peptide and/or modified peptide bonds.
- the peptides comprise at least one peptidomimetic bond selected from intercalation of a methylene (-CH2-) or phosphate (- PO2-) group, secondary amine (-NH-) or oxygen (-0-), alpha-azapeptides, alpha- alky lpeptides, N-alkylpeptides, phosphonamidates, depsipeptides, hydroxymethylenes, hydroxyethylenes, dihydroxyethylenes, hydroxyethylamines, retro-inverso peptides, methyleneoxy, cetomethylene, esters, phosphinates, phosphinics, or phosphonamides.
- the peptides may comprise a protected N-ter and/or C-ter function, for example, by acylation, and/or amidation and/or esterification.
- peptides include, for instance the peptide with amino acid residues 144- 162 of SEQ ID NO: 211 (gC1qR) and the peptide with amino acid residues 204-218 of SEQ ID NO: 211 (gC1qR).
- peptides of the invention include peptides comprising a sequence of anyone of SEQ ID NOs: 156-185 with one amino acid substitution, more preferably with at least one amino acid selected from W, I or K replaced with an Alanine.
- the peptides of the invention may be produced by techniques known per se in the art such as chemical, biological, and/or genetic synthesis.
- isolated refers to molecules (e.g., nucleic or amino acid) that are removed from a component of their natural environment, isolated or separated, and are at least 60% free, preferably 75% free, and most preferably 90% free from other components with which they are naturally associated.
- an “isolated” polypeptide (or protein) is for instance a polypeptide separated from a component of its natural environment and, preferably purified to greater than 90% or 95% purity as determined by, for example, electrophoretic (e.g., SDS-PAGE, isoelectric focusing (IEF), capillary electrophoresis) or chromatographic (e.g., ion exchange or reverse phase HPLC) migration.
- electrophoretic e.g., SDS-PAGE, isoelectric focusing (IEF), capillary electrophoresis
- chromatographic e.g., ion exchange or reverse phase HPLC
- inhibitors are small drug inhibitors, such as are hydrocarbon compounds that selectively bind gC1qR or a cofactor.
- Small drugs are preferably obtainable by a method comprising: (i) contacting a test compound with a cell expressing gC1qR, (ii) selecting a test compound which binds gC1qR, and (iii) selecting a compound of (ii) which inhibits an activity of gC1qR.
- a method comprising: (i) contacting a test compound with a cell expressing gC1qR, (ii) selecting a test compound which binds gC1qR, and (iii) selecting a compound of (ii) which inhibits an activity of gC1qR.
- the cofactor inhibitor is a soluble form of gC1qR.
- viruses express the peptide at the surface of the virus to ensure exposure.
- the peptide is inserted in the envelope protein of the virus.
- Viruses that may be used for that purpose include any safe virus for use in humans, such as adenoviruses, AAVs, retroviruses (including lentiviruses), etc. gClqR soluble receptors
- the cofactor inhibitor is a soluble form of gC1qR.
- inhibition of the cofactor in a subject is obtained by using (e.g., vaccinating or immunizing the subject with) an immunogen or an antigen that can induce an immune response in vivo.
- the subject produces antibodies (or immune cells) which inhibit the cofactor (such as its interaction with gC1qR).
- administration(s) of a cofactor immunogen e.g., any immunogenic portion of a cofactor
- These antibodies will inhibit the cofactor effect as immunotherapy or a vaccine prophylaxis.
- An object of the invention thus resides in a method of treating coronavirus infection by vaccinating a subject with an antigen that induces an immune response against a PLA2- GIB cofactor, particularly against SARS-Cov-2 spike antigen.
- a further object of the invention relates to an antigen that induces an immune response against of a PLA2-GIB cofactor, particularly against SARS-Cov-2 spike antigen, for use to vaccinate a subject in need thereof.
- the immunogen of a PLA2-GIB cofactor antigen used for vaccination is an inactivated immunogenic molecule that induces an immune response against the cofactor in a subject.
- Inactivation may be obtained e.g., by chemically or physically altering the cofactor or by mutating or truncating the protein, or both; and immunogenicity may be obtained as a result of the inactivation and/or by further conjugating the protein to a suitable carrier or hapten, such as KLH, HSA, polylysine, a viral anatoxin, or the like, and/or by polymerization, or the like.
- the immunogen may thus be chemically or physically modified, e.g., to improve its immunogenicity.
- the immunogen of a PLA2-GIB cofactor of the invention comprises the entire cofactor or a variant thereof.
- the immunogen comprises a virus envelope protein with a modified gC1qR binding motif.
- Such immunogen thus retains immunogenicity while having no or less PLA2-GIB cofactor effect.
- a further object of the invention thus relates to an immunogen, wherein the immunogen comprises a viral envelope protein with a modified gC1qR binding motif, preferably a deleted and/or mutated gC1qR binding motif.
- the immunogen comprises a viral envelope protein with a modified gC1qR binding motif, preferably a deleted and/or mutated gC1qR binding motif.
- a specific example is a SARS-Cov-2 spike protein having, by reference to SEQ ID NO: 209, at least one amino acid modification in a residue selected from anyone of SEQ ID NOs: 156-185, even more preferably in a residue selected from aa 348-362 or 431-445 (by reference to SEQ ID NO: 209).
- a specific embodiment of the invention is a spike protein comprising the amino acid sequence of SEQ ID NO: 209, or a natural variant thereof, wherein the sequence contains at least one amino acid substitution or deletion in a position selected from: aa99-113, aal47-161, aa253-267, aa348-362, aa431-445.
- the sequence contains at least one deletion of anyone said residues, even more preferably of at least 2, 3, 4, 5, 6, 7, 8, 9, 10, or more consecutive aa residues at said positions.
- a particular protein of the invention is a protein comprising SEQ ID NO: 209 with a deletion, preferably of at most 50 amino acids, spanning all or part of AA431-445.
- Another particular protein of the invention is a protein comprising SEQ ID NO: 209 with a deletion, preferably of at most 50 amino acids, spanning all or part of AA348-362.
- Such antigens retain immunogenicity of the viral protein, such as an ability to induce immune response against the vims when administered in vivo.
- Such antigens have a reduced cofactor effect, such as a cofactor effect reduced by at least 20% as compared to the native protein or virus.
- the immunogen of a PLA2-GIB cofactor comprises a fragment of a cofactor comprising at least 6 consecutive amino acid residues and containing an immunogenic epitope thereof. In a preferred embodiment, the immunogen comprises at least from 6 to 20 amino acid residues.
- Preferred immunogens of the invention comprise or consist of from 4 to 30 consecutive amino acid residues of a protein or peptide selected from anyone of SEQ ID NOs: 2-211.
- the immunogen may be in various forms such as in free form, polymerized, chemically or physically modified, and/or coupled (i.e., linked) to a carrier molecule. Coupling to a carrier may increase the immunogenicity and (further) suppress the biological activity of the immunogen.
- the carrier molecule may be any carrier molecule or protein conventionally used in immunology such as for instance KLH (Keyhole limpet hemocyanin), ovalbumin, bovine serum albumin (BSA), a viral or bacterial anatoxin such as toxoid tetanos, toxoid diphteric B cholera toxin, mutants thereof such as diphtheria toxin CRM 197, an outer membrane vesicle protein, a polylysine molecule, or a vims like particle (VLP).
- the carrier is KLH or CRM 197 or a VLP.
- Coupling of the immunogen to a carrier may be performed by covalent chemistry using linking chemical groups or reactions, such as for instance glutaraldehyde, biotin, etc.
- the conjugate or the immunogen is submitted to treatment with formaldehyde in order to complete inactivation of the cofactor.
- the immunogenicity of the immunogen may be tested by various methods, such as by immunization of a non-human animal grafted with human immune cells, followed by verification of the presence of antibodies, or by sandwich ELISA using human or humanized antibodies. The lack of biological activity may be verified by any of the activity tests described in the application.
- the invention relates to an inactivated and immunogenic PLA2-GIB cofactor.
- the invention relates to a PLA2-GIB cofactor protein or a fragment or variant thereof conjugated to a carrier molecule, preferably to KLH.
- the invention relates to a vaccine comprising an immunogen of PLA2- GIB cofactor, a suitable excipient and, optionally, a suitable adjuvant.
- a further object of the invention relates to a method for inducing the production of antibodies that neutralize the activity of a PLA2-GIB cofactor in a subject in need thereof, the method comprising administering to said subject an effective amount of an immunogen or vaccine as defined above.
- Administration of an immunogen or vaccine of the invention may be by any suitable route, such as by injection, preferably intramuscular, subcutaneous, transdermal, intravenous or intraarterial; by nasal, oral, mucosal or rectal administration.
- the inhibitor of the SPLA2G1B pathway may be used alone, or in combination with one or more other active agents, such as a cytostatic or cytotoxic agent.
- the agent may be an antibiotic, such as any broad-spectrum antibiotic, or an antibiotic with specific spectrum towards the target bacterium.
- antibiotics include, but are not limited to, amoxicillin, clarithromycin, cefuroxime, cephalexin ciprofloxacin, clindamycin, doxycycline, metronidazole, terbinafine, levofloxacin, nitrofurantoin, tetracycline, penicillin and azithromycin.
- the agent may be an anti-fungal agent, such as, without limitation, clotrimazole, butenafine, butoconazole, ciclopirox, clioquinol, clioquinol, clotrimazole, econazole, fluconazole, flucytosine, griseofulvin, haloprogin, itraconazole, ketoconazole, miconazole, naftifine, nystatin, oxiconazole, sulconazole, terbinafine, terconazole, tioconazole, and tolnaftate.
- an anti-fungal agent such as, without limitation, clotrimazole, butenafine, butoconazole, ciclopirox, clioquinol, clioquinol, clotrimazole, econazole, fluconazole, flucytosine, griseofulvin, haloprogin, itraconazole, ketoconazole, miconazole,
- the agent may be an anti-viral agent, such as, without limitation, zidovudine, didanosine, zalcitabine, stavudine, lamivudine, abacavir, tenofovir, nevirapine, delavirdine, efavirenz, saquinavir, ritonavir, indinavir, nelfinavir, saquinavir, amprenavir, and lopinavir.
- the agent may be selected from other drugs such as chloroquine, hydroxychloroquine, Remdesivir, or ABX464, for instance.
- the invention also relates to a composition
- a composition comprising an inhibitor as defined above and, preferably, a pharmaceutically acceptable diluent, excipient or carrier.
- a “pharmaceutical composition” refers to a formulation of a compound of the invention (active ingredient) and a medium generally accepted in the art for the delivery of biologically active compounds to the subject in need thereof.
- a carrier includes all pharmaceutically acceptable carriers, diluents, medium or supports therefore.
- Conventional pharmaceutical practice may be employed to provide suitable formulations or compositions to subjects, for example in unit dosage form.
- the compounds or compositions according to the invention may be formulated in the form of ointment, gel, paste, liquid solutions, suspensions, tablets, gelatin capsules, capsules, suppository, powders, nasal drops, or aerosol, preferably in the form of an injectable solution or suspension.
- the compounds are generally packaged in the form of liquid suspensions, which may be injected via syringes or perfusions, for example.
- the compounds are generally dissolved in saline, physiological, isotonic or buffered solutions, compatible with pharmaceutical use and known to the person skilled in the art.
- the compositions may contain one or more agents or excipients selected from dispersants, solubilizers, stabilizers, preservatives, etc.
- Agents or excipients that can be used in liquid and/or injectable formulations are notably methylcellulose, hydroxymethylcellulose, carboxymethylcellulose, polysorbate 80, mannitol, gelatin, lactose, vegetable oils, acacia, etc.
- the carrier can also be selected for example from methyl-beta-cyclodextrin, a polymer of acrylic acid (such as carbopol), a mixture of polyethylene glycol and polypropylene glycol, monoethanolamine and hydroxymethyl cellulose.
- compositions generally comprise an effective amount of a compound of the invention, e.g., an amount that is effective to modulate directly or indirectly an effect of PLA2-GIB on CD4 T cells.
- Inhibitors are typically used in an amount effective to maintain/restore resistance of CD4 T cells to inactivation by PLA2-GIB.
- compositions according to the invention comprise from about 1 ⁇ g to 1000 mg of a cofactor or modulator, such as from 0.001-0.01, 0.01-0.1, 0.05-100, 0.05-10, 0.05-5, 0.05- 1, 0.1-100, 0.1-1.0, 0.1-5, 1.0-10, 5-10, 10-20, 20-50, and 50-100 mg, for example between 0.05 and 100 mg, preferably between 0.05 and 5 mg, for example 0.05, 0.1, 0.2, 0.3, 0.4, 0.5, 1, 2, 3, 4 or 5 mg.
- a cofactor or modulator such as from 0.001-0.01, 0.01-0.1, 0.05-100, 0.05-10, 0.05-5, 0.05- 1, 0.1-100, 0.1-1.0, 0.1-5, 1.0-10, 5-10, 10-20, 20-50, and 50-100 mg, for example between 0.05 and 100 mg, preferably between 0.05 and 5 mg, for example 0.05, 0.1, 0.2, 0.3, 0.4, 0.5, 1, 2, 3, 4 or 5 mg.
- compositions of the invention can further comprise one or more additional active compounds, for separate, simultaneous or sequential use.
- additional active compounds include, but are not limited to, chemotherapeutic drug, antibiotics, antiparasitic agents, antifungal agents or antiviral agents.
- the inhibitor is used in combination with a modulator of PLA2-GIB.
- the methods or compositions of the invention use a combination of (i) an inhibitor of a PLA2GIB cofactor and (ii) an antibody against PLA2GIB (or a derivative or fragment thereof).
- the inhibitor of a PLA2GIB cofactor in an antibody against the cofactor, or an antibiotic, or an antifungal agent, or an antivirus agent.
- the method or compositions of the invention use a combination of (i) an inhibitor of a PLA2GIB cofactor and (ii) an indole-based inhibitor of PLA2GIB (such as 3-(2-amino-l,2-dioxoethyl)-2-ethyl-l-(phenylmethyl)-lH-indol-4- yl)oxy)acetic acid or a pharmaceutically acceptable salt, hydrate, or prodrug thereof, such as a sodium salt thereof (Varespladib)).
- the inhibitor of a PLA2GIB cofactor in an antibody against the cofactor, or an antibiotic, or an antifungal agent, or an antivirus agent.
- the method or compositions of the invention use a combination of (i) an inhibitor of a PLA2GIB cofactor and (ii) a pentapeptide inhibitor of PLA2GIB (such as a cyclic peptide selected from FLSYK, FLSYR and (2NapA)LS(2NapA)R).
- a pentapeptide inhibitor of PLA2GIB such as a cyclic peptide selected from FLSYK, FLSYR and (2NapA)LS(2NapA)R.
- the inhibitor of a PLA2GIB cofactor in an antibody against the cofactor, or an antibiotic, or an antifungal agent, or an antivirus agent.
- the method or compositions of the invention use a combination of (i) an inhibitor of a PLA2GIB cofactor and (ii) a further active agent such as an antiviral agent, an antibiotic, chloroquine or hydroxychloroquine.
- the invention also relates to a method for preparing a pharmaceutical composition, comprising mixing a cofactor or modulator as previously described and a pharmaceutically acceptable diluent or excipient, and formulating the composition in any suitable form or container (syringe, ampoule, flask, bottle, pouch, etc.).
- the invention also relates to a kit comprising (i) a composition comprising a cofactor or modulator as previously described, (ii) at least one container, and optionally (iii) written instructions for using the kit.
- the compounds and compositions and methods of the invention may be used to treat a variety of diseases caused by a Group IV RNA virus infection, particularly coronavirus infection, more particularly SARS-Cov-2 infection.
- the invention is directed to methods for stimulating an immune response in a subject infected with a Group IV RNA vims, particularly coronavirus.
- the invention is directed to methods for treating an immunodeficiency or an associated disorder in a subject infected with a Group IV RNA virus infection, particularly a coronavirus.
- the inhibitor may be administered by any suitable route.
- administration is by injection, such as systemic or parenteral injection or perfusion, e.g., intramuscular, intravenous, intraarterial, subcutaneous, etc. Administration is typically repeated, or continuous.
- the duration, dosages and frequency of administering compounds or compositions of the invention may be adapted according to the subject and disease.
- a typical regimen comprises a single or repeated administration of an effective amount of an inhibitor over a period of one or several days, up to one year, and including between one week and about six months. It is understood that the dosage of a pharmaceutical compound or composition of the invention administered in vivo will be dependent upon the age, health, sex, and weight of the recipient (subject), kind of concurrent treatment, if any, frequency of treatment, and the nature of the pharmaceutical effect desired.
- the ranges of effectives doses provided herein are not intended to be limiting and represent preferred dose ranges.
- the most preferred dosage will be tailored to the individual subject, as is understood and determinable by one skilled in the relevant arts (see, e.g., Berkowet et al., eds., The Merck Manual, 16 th edition, Merck and Co., Rahway, N.J., 1992; Goodmanetna., eds., Goodman and Oilman’s The pharmacological Basis of Therapeutics, 10 th edition, Pergamon Press, Inc., Elmsford, N.Y., (2001)).
- the invention may be used in any mammal, particularly any human.
- the present invention also provides efficient methods for detecting SARS-Cov-2 infection in a sample. More particularly, the cofactors and antibodies of the inention can be used as reagent for detecting SARS-Cov-2 infection in a sample.
- the invention relates to the use of an antibody of the invention for detecting the presence of SARS-Cov-2 in a sample.
- the invention relates to the use of a polypeptide or molecule of the invention for detecting the presence of anti-SARS-Cov-2 antibodies in a sample.
- the detection is preferably performed in vivo, using conventional techniques such as immunoassays or radioassays.
- detection is through ELISA type assay.
- the sample is contacted with either the polypeptides or antibodies of the invention, and the formation of an immune complex is detected, such as with an antibody.
- the method is typically conducted in any suitable device, such as plate, microplate, tube, etc.
- the reagents are typically coated on a surface of the device.
- the sample may by any biological sample or derivatives thereof (dilutions, concentrate, enriched, aliquotes, etc), such as a biological fluid (e.g., blood, serum, plasma, urine, nasal swab, etc.).
- the invention relates to a method for detecting in vitro the presence of SARS-Cov-2 in a sample, comprising (i) contacting the sample in vitro with an antibody (or fragment or derivative thereof) of the invention, particularly an anti-Spike antibody (or fragment or derivative thereof) of the invention, and (ii) detecting the presence of antibody-antigen type complexes.
- the invention relates to a method for detecting in vitro the presence of anti-SARS-Cov-2 antibodies in a sample, comprising (i) contacting the sample in vitro with a molecule comprising a sequence selected from anyone of SEQ ID Nos: 1-208 or an epitope-containing fragment thereof, particularly a molecule comprising a sequence selected from anyone of SEQ ID Nos: 156-185 or an epitope-containing fragment thereof, and (ii) detecting the presence of antibody-antigen type complexes.
- the invention also relates to a kit comprising (i) an antibody (or fragment or derivative thereof) of the invention, or (ii) a molecule comprising a sequence selected from anyone of SEQ ID Nos: 1-208 or an epitope-containing fragment thereof.
- the kit may further comprise a support and/or reagents and/or instructions for performing an immunoassay.
- Example 1 Definition of the Position Specific Scoring Matrix (PSSM) for the identification of gClqR peptide binding motifs
- PSSM Position Specific Scoring Matrix
- Synthetic peptides designed by the inventors were assayed in vitro for their binding capacity to the receptor for the globular heads of Clq (gC1qR).
- the in vitro assay is based on an enzyme-linked immunosorbent assay (ELISA), as disclosed in Pednekar et al. Mol Immunol. 2016. The sequence of the tested peptides is presented in table 1 below. Table 1:
- Peptide sequences and binding values were used to define an optimized peptide-binding motif to gC1qR using GibbsCluster 2.0 (http://www.cbs.dtu.dk/services/GibbsCluster- 2.01. GibbsCluster 2.0 was run with default parameters except length which was set to 15 amino-acids.
- Cluster 1 obtained from the analysis was selected to generate a Position Specific Scoring Matrix (PSSM), which is disclosed in Figure 1.
- PSSM Position Specific Scoring Matrix
- the matrix displays weight (or "preference") for any given amino acid at each position for a window of 15 amino acids in length.
- Example 2 Identification of gClqR binding motifs (gC1qR:PSSM) within the SARS-CoV-2 proteome.
- PSSM Position Specific Scoring Matrix
- SEQ ID NO:l A SARS-CoV-2 proteome sequence (SEQ ID NO:l) was constructed by co-linear assembly of open-reading frames from the SARS-CoV-2 reference genome (NC_045512.2). Positions of the different open-reading frames (ORFlab, Spike, ORF3a, E, M, ORF6, ORF7a, ORF7b, ORF8, N and ORF10) are presented in table 2. Table 2: Positions of SARS-CoV-2 open-reading frames within SEQ ID NQ:1.
- PSSM search For any selected windows of 15 amino acids, the PSSM returns a single score calculated by adding the relevant values at each position.
- the PSSM score is assigned to the N- terminal amino acid position of the window unless otherwise stated.
- Figure 2A provides the PSSM scores across the SARS-CoV-2 proteome (SEQ ID NO: 1).
- gC1qR-binding motifs were thus identified in the SARS-CoV-2 proteins.
- binding domains were identified in the following proteins: ORFlab, Spike, ORF3, E, ORF7b, ORF8, N and M.
- Each of said proteins thus represents a relevant target for inhibition according to the invention.
- Each of said peptides sequence represents an object of the invention.
- Each of said proteins and peptides can act as PLA2-GIB cofactors in SARS-Cov2 infected patients.
- Each of said peptides represents a strong inhibitor (or be used to construct a strong inhibitor) for use according to the invention
- Each of said proteins or peptides can be used to produce relevant inhibitors, such as antibodies or derivatives thereof.
- Each of said proteins and peptides can be used as a reagent to detect the presence of anti- Cov-2 antibodies in a sample, particularly of a protective immunity.
- Example 3 Analysis of binding peptides from Spike antigen Spike is the principal antigen exposed at the surface of the SARS-CoV-2 virus. It is an ideal target for antibody as it is expressed on the surface of the enveloped virus. Also, the spike antigen interacts with the receptor ACE-2.
- the amino acid sequence of the Spike protein is represented as SEQ ID NO: 209.
- the spike antigen comprises several functional domains, which are disclosed in the following Table: Thirty (30) gC1qR-binding peptides derived from the spike antigen were identified (see example 2). Each of said peptides was further analyzed for selection/optimization.
- Figure 3 provides the PSSM score across the spike antigen (SEQ ID NO: 209). In this figure, PSSM scores are assigned to a 15aa peptide represented by a bar. The figure shows most peptides are located within the following sub-domains of the spike antigen: NTD, RBD, S2 and TM. Each peptide was analyzed for solvent accessibility, co-localization with B cell epitopes and/or with HLA class II binding motif-rich domains. The results are presented below. Said further analysis confirmed the relevance of the claimed therapeutic approach and further led to the selection of most preferred peptides and domains. 3.1. Co-localization of peptides with solvent accessible domains.
- gC1qR binding motifs identified in example 2 were aligned with surface accessible domains within the spike antigen (SEQ ID NO: 209).
- Solvent accessible structure was measured from PDB structure using the “Dictionary of Secondary Structure of Proteins” (DSSP) developed by Kabsch and Sander (Kabsch W, Sander C. Dictionary of protein secondary structure: pattern recognition of hydrogen- bonded and geometrical features. Biopolymers. 1983 Dec;22(12):2577-637.).
- the DSSP program defines secondary structure, geometrical features and solvent exposure of proteins, given atomic coordinates in Protein Data Bank format (PDB) or macromolecular Crystallographic Information File format.
- Solvent exposure is given as the number of water molecules in possible contact with a residue (residue water exposed surface in Angstrom 2 ).
- DSSP software package can be obtained from https://swift.cmbi.umcn.nl/gv/dssp/DSSP 3.html.
- the structure information for the spike antigen from SARS-CoV-2 used for solvent accessibility analysis was obtained from SWISS-MODEL (https://swissmodel.expasy.org/) based on a model number 4 (https://swissmodel.expasv.org/assess/7dVLxC/04) derived the cryo-EM structure of spike (PDB accession number: 6vsb) as established by Wrapp et al (Wrapp D, Wang N, Corbett KS, Goldsmith JA, Hsieh CL, Abiona O, Graham BS, McLellan JS. Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science. 2020 Mar 13;367(6483):1260-1263).
- the 3D model for spike SARS-CoV-2 is covering a sequence corresponding to residues 27 to 1146 from SEQ ID NO:209. For this analysis, only information from chain (a) - monomeric chain in the “up” conformation - was
- Figure 4 presents the co-localization of high scoring gC1qR:PSSM motifs with solvent accessible domains within the spike protein sequence from SARS-CoV-2. It can be shown that most peptides in NTD, RBD and S2 domains are solvent accessible, and thus exposed and suitable for use as medicament or target.
- B cell epitopes were determined in the sequence of the Spike protein and then aligned with the peptide position.
- B cell epitopes are regions of an antigen protein surface that are preferentially recognized by antibodies. B cell epitopes interact with an antibody based on the conformation adopted by the epitope, which is determined by the surface features of the involved epitope residues and the shape or tertiary structure of other segments of the antigen. Based on spatial structure, B-cell epitopes can be categorized as a continuous and discontinuous (nonlinear or conformational) epitopes. In the latter case, amino acid residues are in close contact due to the three-dimensional conformation.
- DiscoTope 2.0 http ://tool s.iedb.org/discotope/
- DiscoTope method incorporating a spatial neighborhood definition and half-sphere exposure as surface measure. J. V. Kringelum, C. Lundegaard, O. Lund, M. Nielsen. 2012. Reliable B cell epitope predictions: impacts of method development and improved benchmarking. PLoS Comput Biol. 8:(12):el002829.
- the method utilizes calculation of surface accessibility (estimated in terms of contact numbers) and an epitope propensity amino acid score.
- the final scores are calculated by combining the propensity scores of residues in spatial proximity and the contact numbers.
- the structure information for the spike antigen from SARS-CoV-2 used for DiscoTope 2.0 analysis was obtained from SWISS-MODEL (https://swissmodel.expasy.org/) based on a model number 4 (https://swissmodel.expasv.org/assess/7dVLxC/04) derived the cryo-EM structure of spike (PDB accession number: 6vsb) as established by Wrapp et al (Wrapp D, Wang N, Corbett KS, Goldsmith JA, Hsieh CL, Abiona O, Graham BS, McLellan JS.
- Figure 5 presents co-localization of high scoring gC1qR-binding peptides with predicted B cell epitopes (Discotope) within the spike protein sequence from SARS-CoV-2.
- peptides in NTD, RBD and S2 domains colocalize with predicted B cell epitopes, thus confirming their relevance as targets for antibody development.
- HLA class II binding motif-rich domains of the SARS-CoV-2 virus spike antigen were determined. Such domains were then aligned with the identified binding peptides.
- HLA Human leukocyte antigen
- HLA -DR Human leukocyte antigen
- -DP Human leukocyte antigen
- -DQ Human leukocyte antigen
- Each locus is comprised of many different allelic variants, which makes the HLA class II molecule highly polymorphic across the human population.
- Peptides presented by HLA class II molecules bind to a binding groove formed by residues of the HLA a- and the b-chains and a 9 amino-acid long core binding motif contained within the peptide antigen sequences.
- HLA class II binding is a prerequisite for the stimulation of CD4+ T-cells.
- One additional challenge is to identify peptide that can bind a broad number of different HLA class II alleles in order to provide the ability of the peptide antigen to be immunogenic in a large proportion of the population.
- the phenotypic frequency of HLA class II molecules is highly variable in the human population.
- HLA class II binding motifs are not evenly distributed along an antigen sequence and tend to form hotspots of overlapping motifs.
- Many different methods to predict the ability of peptide to bind to MHC class II molecules have been developed, including TEPITOPE, TEPITOPEpan, PROPRED, RANKPEP, SVRMHC, NetMHCIIpan, NetMHCII.
- NetMHCII 2.3 has been shown to be among the best methods for predicting binding affinities to a large number of HLA class II molecules (Jensen KK, Andreatta M, Marcatili P, Buus S, Greenbaum JA, Yan Z, Sette A, Peters B, Nielsen M. Improved methods for predicting peptide binding affinity to MHC class II molecules. Immunology. 2018 Jul;154(3):394-406.).
- NetMHCII is based on ensembles of artificial neural networks that are trained on quantitative peptide binding affinity data from the Immune Epitope Database (IEDB).
- IEDB Immune Epitope Database
- NetMHCII 2.3 server predicts binding of peptides to HLA class II molecules providing a range of information including the %-Rank and the core binding motif corresponding to a sequence of 9 amino-acids. Positive binding peptides are considered to have a %-rank below 10.
- NetMHCII 2.3 analysis was performed with a set of 20 HLA-DR alleles, 20 HLA-DQ, 9 HLA-DP: DRal*0101-DR ⁇ 1*0101, DRal*0101-DR ⁇ 1*0301, DRal*0101-DR ⁇ 1*0401, DRal*0101-DR ⁇ 1*0701, DRal*0101-DR ⁇ 1*0801,
- DRal*0101-DR ⁇ 1*0802 DRal*0101-DR ⁇ 1*0901, DRal*0101-DR ⁇ 1*1001, DRal*0101-DR ⁇ 1*1101, DRal*0101-DR ⁇ 1*1201, DRal*0101-DR ⁇ 1*1301, DRal*0101-DR ⁇ 1*1302,, DRal*0101-DR ⁇ 1*150, DRal *0101 -DR ⁇ 1*1602, DRal*0101-DR ⁇ 3*0101, DRal*0101-DR ⁇ 3*0202, DRal*0101-DR ⁇ 3*0301, DRal*0101-DR ⁇ 4*0101, DRal*0101-DR ⁇ 4*0103, DRal*0101-DR ⁇ 5*0101, DQal *0101-DQ ⁇ 1*0501, DQal*0102-DQ ⁇ 1*0501, DQal *0102-DQ ⁇ 1*0502, DQal *0
- Figure 6 presents co-localization results between the peptides and the predicted HLA class II binding motif-rich domains within the spike protein sequence from SARS-CoV- 2.
- Example 4 Detection of anti-SARS-CoV-2 immunity in a sample
- Recombinant proteins and peptides- SARS-Cov-2 SI (Genbank accession #YP_009724390.1, which is identical to QHD43416) recombinant protein was purified from HEK293 cell culture and purchased from antibodies online (SARS-CoV-2 Spike Subunit SI (SARS-CoV-2 SI) (Active) protein, ABIN6952739, a.a. 11-683, purity >90%).
- GCVIAWNSNNLDSKV-COOH were ordered from Covalab (purity >98%).
- Detection of anti-SARS-CoV-2 immunity in a sample- Detection of anti-SARS-Cov-2 SI IgG and IgM antibodies in SARS-Cov-2 patient plasma/serum is performed by ELISA on coated SARS-Cov-2 SI peptides or SI protein (Antibodies online, ABIN6952739).
- Microtiter plates (Nunc Maxisorp flat-bottom microplate, 44-2404-21, Thermofisher Scientific) are coated overnight at 4°C with 100 ⁇ l of a solution at 5 ⁇ g/ml of SI recombinant protein or SARS-Cov-2 peptides in carbonate buffer pH 9.6.
- Casein blocking buffer (37528, Pierce Blocker Casein in PBS, Thermofisher Scientific). After washing, various dilutions in Casein blocking buffer of the anti-Sl specific Ab as positive control for IgM and IgG (Human IgGl or IgM ANTI SARS-COV-2 SPIKE (SI) ANTIBODY (CR3022), IgM: MAB 12423-50, IgG: MAB 12422-100, the native antigen company) at 125ng/ml and plasma/serum from SARS-Cov-2 patients or healthy donors (negative control) at 1:20 and gradually diluted 1:3 are incubated in quadruplicate for 2h at RT in 100 ⁇ l.
- IgM and IgG Human IgGl or IgM ANTI SARS-COV-2 SPIKE (SI) ANTIBODY (CR3022)
- IgM MAB 12423-50
- IgG MAB 12422-100, the native
- Bound antibody are then detected by interaction with biotinylated rabbit anti-human IgG Ab (Anti-Human IgG antibody [EPR12700] (Biotin), ab201842, abeam) for IgG detection or biotinylated F(ab')2-Goat anti-Human IgM (F(ab')2-Goat anti-Human IgM (Heavy chain) Secondary Antibody, Biotin, A24486, Thermofish scientific), both at 1:10,000 dilution, IOOmI/well for 2h, and is revealed by sequential reaction with HRP-Streptavidin at 1:10,000 (Peroxidase labeled Streptavidin ( SA-HRP) UP395888, Interchim, lh, room temp.) followed by the addition of 100 ⁇ l/well TMB substrate (UP664781, Interchim).
- SA-HRP Peroxidase labeled Streptavidin
- GC VIA WN S NNLDS KV in SARS-Cov-2 SI (Genbank accession #YP_009724390.1) are generated by immunization of OF1 mice with the peptide NH2-C- KNNKS WMESEFRVY S -COOH, NH2-C-DS S S GWT AGAAA YYV -COOH or NH2-C- GCVIAWNSNNLDSKV-COOH conjugated to KLH (KLH-SARS-Cov-2_147-161, KLH-SARS-Cov-2_253-267, KLH-SARS-Cov-2_431-445, respectively). Mice are immunized 5 times every 3 weeks.
- First immunization is subcutaneous and intraperitoneal with 50 ⁇ g of peptides and complete freund’s adjuvant. Then mice are intraperitoneally injected three times with 50 ⁇ g of peptides and incomplete freund’s adjuvant.
- the antibody titer in serum is evaluated by EFISA on SARS-Cov-2_147-161, SARS-Cov-2_253-267, SARS-Cov-2_431-445 peptides and SARS-Cov-2 SI protein (ABIN6952739).
- the mouse with the best titer on protein and peptide is selected and injected intravenously with 5 ⁇ g of SARS-Cov-2 peptide (KLH-SARS-Cov-2_147-161,KLH-SARS-Cov-2_253-267 or KLH-SARS-Cov-2_431-445) alone.
- SARS-Cov-2 peptide KLH-SARS-Cov-2_147-161,KLH-SARS-Cov-2_253-267 or KLH-SARS-Cov-2_431-445
- spleen Three days later the spleen is collected and cells are fused with myeloma (Sp2/0-Agl4) cells to generate hybridoma.
- the best 120 positive on BSA-SARS-Cov- 2 peptides are selected and tested another time on BS A-S ARS-Cov-2 peptide and S 1.
- the isotypes of the best clones are determined and they are frozen including 15 clones that are maintained in cell culture to obtain stable clone, 5 clones per peptide. These 15 clones, are seeded in High Yielding Performance Flask (hyperflask) for cell culture to produce 560 ml of supernatant with 10% Low IgG fetal bovine serum (FBS) in DMEM medium. IgG are purified from supernatant by affinity chromatography on protein A in low endotoxin condition ( ⁇ 10 EU / mg, FAF test). Antibodies binding to BSA-SARS-Cov-2 and S 1 is verified and the cDNA sequence coding the variable regions of these antibodies is determined.
- High Yielding Performance Flask hyperflask
- FBS Low IgG fetal bovine serum
- ELISA to screen new anti-Sl antibodies produced- EFISA on peptides or proteins is performed on microtiter plates coated overnight at room temperature (RT) with 50 pi of a solution in PBS (pH7.4) of BSA-SARS-Cov-2_147-161, BSA-SARS-Cov- 2_253-267, BSA-SARS-Cov-2_431-445, SI protein at 2 ⁇ g/ml. After washing, wells are then blocked lh at RT with 150 pi of PBS/2.5% skimmed milk. After washing, various dilutions in PBS/0.05%Tween 20/0.5%BSA of Ab from 100-0.0001 ⁇ g/ml are incubated for 2h at RT in 50 pi.
- Example 6 Inhibition of sPLA2-GIB affects SARS-CoV-2 plasma inhibitor effect
- CD4 T -lymphocytes Purification of Human CD4 T -lymphocytes- Venous blood was obtained from healthy volunteers through the EFS (Etableau Francais du Sang, Centre Necker-Cabanel, Paris). CD4 T-cells were purified from whole blood using RosetteSep Human CD4+ T cell Enrichment Cocktail (Stem Cell, 15062). This cocktail contains mouse and rat monoclonal antibodies purified from mouse ascites fluid or hybridoma culture supernatant, by affinity chromatography using protein A or Protein G sepharose.
- bispecific tetrameric antibody complexes which are directed against cell surface antigens on human hematopoietic cells (CD8, CD16, CD 19, CD36, CD56 CD66b, TCR ⁇ /d) and glycophorin A on red blood cells.
- the rosetteSep antibody cocktail crosslinks unwanted cells in human whole blood to multiple red blood cells, forming immunorosettes . This increases the density of unwanted cells, such that they pellet along with the free red blood cells when centrifuged over a buoyant density medium such as lymphocytes separation medium (Eurobio, CMSMSLOl-01).
- Cells are subsequently resuspended in RPMI 1640 medium (Lonza) supplemented with 5% FBS, 50 mM HEPES pH 7.4, glutamine, penicillin, streptomycin and fungizone (complete medium), counted with a Moxi Z mini automated cell counter (ORFLO, MXZ000). Cells suspension is adjusted at 7xl0 6 cells/ml and equilibrated at least 2 h at 37°C in a 5% CO2 humidified atmosphere.
- the enriched CD4-T cell population is controlled by flow cytometry on a cytoflex (Beckman coulter). The quiescence of recovered CD4 T-cells is controlled by the low level of IL-2Ro ⁇ (CD25).
- CD4 T cells are labeled with anti-Human CD3 eFluor780 (eBioscience, clone UCHT1, 47-0038-42), anti-Human CD25-PE (Biolegend, clone BC96, 302605) and anti-human CD4-PerCP (BD, clone SK3, 345770).
- the enriched CD4-T cell population contains >95% CD3+CD4+ and less than 8% of CD25+.
- PLA2-G1B and SARS-Cov-2 SI protein immunodepletion to study plasma cofactor activity-
- PLA2-G1B and SARS-Cov-2 SI protein role in SARS-Cov-2 patient plasma inhibitory activity plasma are immunodepleted with anti-PLA2GlB antibodies and anti-Sl antibodies.
- 1 ml VP or HD plasma is incubated with 100 ⁇ g of rabbit anti-PLA2GlB pAb (produced at BIOTEM) or control isotype (Rabbit Gamma Globulin, 31887, Invitrogen) to immunodeplete PLA2-G1B, or with 100 ⁇ g of anti-Sl specific Ab (Human IgGl ANTI SARS-COV-2 SPIKE (SI) ANTIBODY (CR3022), MAB 12422- 100, the native antigen company) or the control goat polyclonal antibody (Ultra-LEAFTM Purified Human IgGl Isotype Control Recombinant Antibody, 403502, Biolegends) to immunodeplete SI, in 1.5-ml Eppendorf tubes overnight (Test tube-rotor, 34528, Snijders, Netherland) at 4°C.
- CD4 T- cells are loaded (3.5x10 5 cells/50 ⁇ l in complete medium) on poly-L-Lysine-coated (Sigma, P8920) round coverslips (14mm-diameter, Marienfeld) in 24-well polystyrene plates at 37°C in a thermo-regulated water and mixed with 50pl of a suspension in PBS BSA1% containing peptides, recombinant proteins together with recombinant PLA2-GIB or not or containing SARS-Cov-2 patient plasma (1 or 3%), PLA2-GlB-depleted or Sl- depleted or not plasma (1 or 3%), or healthy donor plasma.
- the cells suspension was either pretreated with 40pl of peptides, recombinant protein dilutions in PBS BSA1% for 15 minutes with subsequent addition of IOmI PLA2-GIB (5nM at the end) for 30 minutes or directly treated with 50m1 of dilution in PBS BSA 1% with peptides or recombinant protein together with PLA2-GIB (5nM at the end) for 45 minutes.
- Cells were activated for 15 minutes with 2 nM recombinant glycosylated human IL-7 (Accrobio System).
- cDNA was obtained from 1 ⁇ g total RNA as follows: total RNA was extracted using QIAshredder (79654, Qiagen) and the RNeasy Kit (74104, Qiagen). Extracted RNA was treated with DNase I (RNase-Free DNase Set, 79254, Qiagen) and reverse transcribed using Superscript III reverse transcriptase (18080-044, Invitrogen) according to the manufacturer’s instructions.
- DNase I RNase-Free DNase Set, 79254, Qiagen
- Superscript III reverse transcriptase 18080-044, Invitrogen
- Example 7 Peptide SEQ ID NOs: 164 and 168 affect SARS-CoV-2
- Purified SARS-Cov-2 particles were purified from African green monkey kidney Vero E6 (ATCC ® CRL-1586) cell culture.
- Vero E6 cells were seeded (60,000/well) in 500- ⁇ l 24-well plates in DMEM 10% FBS. 24h later cells were treated for lh with SARS-Cov-2_348-362 and SARS-Cov-2_431-445 or a control scrambled peptide.
- Percent of [3H] arachidonic acid in CD4 T cells is the (1 minus ratio of [3H] arachidonic acid in the supernatant of CD4 T cells without cells (cpm/ml) on total [3H] arachidonic acid in supernatant and cells (cpm/ml).
- Jurkat E6.1 T cells (ECACC 88042803) or gC1qR KO Jurkat E6.1 T cells were incubated for 17h at 5x10 5 cells/ml with 1 ⁇ Ci/ml of arachidonic acid [5,6,8,9,11,14,15- 3 H(N)] (Perkin Elmer, NET298Z250UC) in RPMI 1640 medium (Lonza) supplemented with 10% FBS, 50 mM HEPES pH 7.4, glutamine, penicillin, streptomycin and fungizone at 2 ml/well in 6-well plates at 37°C in a 5% C02 humidified atmosphere.
- arachidonic acid [5,6,8,9,11,14,15- 3 H(N)]
- Percent of [3H] arachidonic acid in CD4 T cells is the (1 minus ratio of [3H] arachidonic acid in the supernatant of CD4 T cells without cells (cpm/ml) on total [3H] arachidonic acid in supernatant and cells (cpm/ml).
- Results are expressed as PLA2GIB activity (release of [3H] arachidonic acid in the supernatant of cells treated with peptide together with PLA2G1B minus spontaneous release of [3H] arachidonic acid by cells in cpm/ml) or ⁇ PLA2G1B activity with peptides minus activity with Scrambled peptide (release of [3H] arachidonic acid in the supernatant of cells treated with peptide minus release of [3H] arachidonic acid by cells treated with Scrambled in cpm/ml).
- SEQ ID NO:l SARS-CoV-2 proteome (9744 amino acids)
- MFVFF VFFPFV S S QC VNFTTRTQFPP A YTN S FTRG V Y YPDKVFRS S VFHS T QDF FFPFFSNVTWFHAIHVSGTNGTKRFDNPVFPFNDGVYFASTEKSNIIRGWIFGTT FDSKTQSFFIVNNATNVVIKVCEFQFCNDPFFGVYYHKNNKSWMESEFRVYSS ANNCTFEY V S QPFFMDFEGKQGNFKNFREFVFKNIDGYFKIY S KHTPINFVRDF PQGF S AFEPFVDFPIGINITRFQTFF AFHRS YFTPGDS S S GWT AG A A A Y Y V G YFQ PRTFFFKYNENGTITDAVDCAFDPFSETKCTFKSFTVEKGIYQTSNFRVQPTESI VRFPNITNFCPF GE VFN ATRFAS V Y A WNRKRIS NC V AD Y S VF YN S AS FS
Landscapes
- Health & Medical Sciences (AREA)
- Life Sciences & Earth Sciences (AREA)
- Chemical & Material Sciences (AREA)
- General Health & Medical Sciences (AREA)
- Medicinal Chemistry (AREA)
- Immunology (AREA)
- Veterinary Medicine (AREA)
- Pharmacology & Pharmacy (AREA)
- Public Health (AREA)
- Animal Behavior & Ethology (AREA)
- Mycology (AREA)
- Epidemiology (AREA)
- Microbiology (AREA)
- Virology (AREA)
- Organic Chemistry (AREA)
- Molecular Biology (AREA)
- Nuclear Medicine, Radiotherapy & Molecular Imaging (AREA)
- Chemical Kinetics & Catalysis (AREA)
- General Chemical & Material Sciences (AREA)
- Oncology (AREA)
- Communicable Diseases (AREA)
- Biochemistry (AREA)
- Biophysics (AREA)
- Genetics & Genomics (AREA)
- Proteomics, Peptides & Aminoacids (AREA)
- Medicines Containing Antibodies Or Antigens For Use As Internal Diagnostic Agents (AREA)
- Peptides Or Proteins (AREA)
Abstract
The present invention relates to novel therapeutic approaches for treating or preventing diseases in mammals, particularly in human subjects, using a PLA2-GIB cofactor, or a modulator of a PLA2-GIB cofactor.
Description
THERAPEUTIC METHODS
The present invention relates to novel compounds, compositions, uses and methods for treating or detecting RNA virus infections in mammals, particularly in human subjects. The invention may be in used in a preventive or curative approach, alone or in combination with other treatments.
Introduction and background It has been documented by the inventor that sPLA2-GIB is involved in the inactivation of CD4 T cells in HIV infected patients, and that sPLA2-GIB inhibitors are effective for treating disorders associated with an immune deficiency (see WO2015/097140). The inventors have also discovered that pathogens produce or activate cofactors which bind gC1qR and cause a sensitization of CD4 T cells to inactivation by sPLA2-GIB. The inventors have shown that modulating such cofactors is an effective approach for treating such diseases (WO2019/166412).
Continuing their research, applicant surprisingly found that SARS-Cov2 infection can also utilize the sPLA2-GIB pathway. The inventors have identified that SARS-Cov2 can act as a sPLA2-GIB cofactor, causing T cell anergy, especially mediated by the spike protein.
The invention thus provides novel efficient molecules, compositions, methods and uses for detecting and/or treating coronavirus infections as well as other RNA virus infections, particularly SARS-Cov2 infection.
Summary of the invention An object of the invention relates to methods for treating a Group IV RNA vims infection, particularly a coronavirus infection, in a mammal, particularly a human, comprising inhibiting the sPLA2-GIB pathway in said mammal.
A further object of the invention relates to the use of an inhibitor of the sPLA2-GIB pathway for the manufacture of a medicament for treating a Group IV RNA vims infection, particularly a coronavirus infection.
Another object of the invention concerns an inhibitor of the sPLA2-GIB pathway for use for treating a Group IV RNA vims infection, particularly a coronavirus infection.
A further object of the invention relates to any methods for inhibiting a sPLA2-GIB cofactor effect of SARS-Cov2 in a mammal.
A further object of the invention is a polypeptide comprising a sequence selected from anyone of SEQ ID Nos: 2-208 or an immunogenic-fragment thereof. The polypeptide is preferably a polypeptide of less than 50 amino acids, more preferably less than 40 amino acids. The polypeptide may consist or consist essentially of anyone of SEQ ID Nos: 2- 208. The polypeptides of the invention may be conjugated to any molecule, such as a protein, carrier, surface, device, adjuvant, bead, column, etc.
A further object of the invention relates to a molecule which binds a Group IV RNA virus, particularly a coronavirus, in a binding domain comprising at least one amino acid residue of any one of SEQ ID Nos: 2-208.
A further object of the invention relates to an antibody (or a fragment or derivative thereof) which binds a Group IV RNA virus, particularly a coronavirus, wherein said antibody binds an epitope comprising at least one amino acid residue of any one of SEQ ID Nos: 2-208.
A further object of the invention relates to a vaccine composition comprising an immunogen, wherein the immunogen comprises a sequence selected from anyone of SEQ ID Nos: 2-208 or a fragment thereof.
A further object of the invention relates to a vaccine composition comprising an immunogen, wherein the immunogen comprises a viral envelope protein with a modified gC1qR binding motif, preferably a deleted and/or mutated gC1qR binding motif. A specific example is a SARS-Cov-2 spike protein having at least one amino acid modification in a domain selected from SEQ ID NO: 156-185, even more preferably in a domain selected from aa 348-362 or 431-445 (by reference to SEQ ID NO: 209).
The invention also relates to a combination therapy or therapeutic regimen for treating a Group IV RNA virus infection, particularly a coronavirus, comprising (i) an inhibitor of PLA2-GIB pathway, in combination with (ii) at least one further active agent. The further active agent may be an antiviral agent, an antibiotic, an anti-inflammatory agent, etc. The agents in a combination therapy of the invention may be formulated together or separately, for combined, separate or sequential administration(s).
The invention also relates to the use of a PLA2-GIB cofactor, or an agonist, fragment, derivative, or mimotope thereof, for the manufacture of a medicament to induce immunosuppression in a subject in need thereof, by increasing the effect of PLA2-GIB on T cells.
The invention may be used in any mammal, particularly in human subjects. It is suitable to treat Group IV RNA vims infections, particularly coronavirus infections, at any stage of infection, either alone or in combination with other agents/therapies.
Legend to the figures
Figure 1: Position Specific Scoring Matrix (PSSM) defining peptide binding motif to gClqR. The matrix of size P x A is presented, where P is the length of the window and A is the amino-acid one letter code according to the IUB/IUPAC standard.
Figure 2: gClqR:PSSM motifs across the SARS-CoV-2 proteome (SEQ ID NO: 1).
The graphic reports gC1qR:PSSM scores for every given 15 a.a. window scanning the SARS-CoV-2 proteome (SEQ ID Nol). The gC1qR:PSSM score is assigned to the N- terminal amino acid position of every given 15 a.a. window (2A). Only motifs with a gC1qR:PSSM score >=5 are presented in 2B. The location of the different open-reading frames within the SARS-CoV-2 proteome are presented below the x-axis.
Figure 3: High scoring gClqR:PSSM motifs across the spike protein sequence from SARS-CoV-2 (SEQ ID NO: 209). The graphic reports gC1qR:PSSM scores for every given 15 a.a. window scanning the sequence of spike from SARS-CoV-2. gC1qR:PSSM scores assigned to 15 a.a. motifs are represented by bars. Overlapping gC1qR:PSSM motifs are represented by bars of different colors. Only motifs with a gC1qR:PSSM score >=5 are presented. The location of the different open-reading frames within the SARS- CoV-2 proteome are presented below the x-axis.
Figure 4: Co-localization of high scoring gClqR:PSSM motifs with solvent accessible domains within the spike protein sequence. The graphic reports gC1qR:PSSM scores for every given 15 a.a. window scanning the sequence of spike from SARS-CoV-2. gC1qR:PSSM scores assigned to 15 a.a. motifs are represented by bars. Overlapping gC1qR:PSSM motifs are represented by bars of different colors. Only motifs with a gC1qR:PSSM score >=5 are presented. Solvent accessible scores were calculated according to the method presented in example 3. The location of the different open- reading frames within the SARS-CoV-2 proteome are presented below the x-axis. Figure 5: Co-localization of high scoring gC1qR:PSSM motifs with predicted B cell epitopes (Discotope) within the spike protein sequence. The graphic reports gC1qR:PSSM scores for every given 15 a.a. window scanning the sequence of spike from SARS-CoV-2. gC1qR:PSSM scores assigned to 15 a.a. motifs are represented by bars. Overlapping gC1qR:PSSM motifs are represented by bars of different colors. Only motifs with a gC1qR:PSSM score >=5 are presented. Discotope scores were calculated
according to the method presented in example 3. The location of the different open- reading frames within the SARS-CoV-2 proteome are presented below the x-axis.
Figure 6: Co-localization of high scoring gC1qR:PSSM motifs with predicted HLA class II binding motif-rich domains within the spike protein sequence. The graphic reports gC1qR:PSSM scores for every given 15 a.a. window scanning the sequence of spike from SARS-CoV-2. gC1qR:PSSM scores assigned to 15 a.a. motifs are represented by bars. Overlapping gC1qR:PSSM motifs are represented by bars of different colors. Only motifs with a gC1qR:PSSM score >=5 are presented. Number of HLA class II binding motifs were calculated according to the method presented in example 3. The location of the different open-reading frames within the SARS-CoV-2 proteome are presented below the x-axis.
Detailed description of the invention
The invention generally relates to novel molecules, compositions, uses and methods for treating or detecting a subject infected by a Group IV RNA virus, particularly SARS- Cov2.
Applicant have discovered that SARS-Cov-2 exerts PLA2-G1B cofactor effect. Applicant identified various peptides in SARS-Cov2 proteins that can bind to gC1qR and cause CD4 T cell anergy. Particularly relevant peptides are located in SI protein, such as in the N-terminal domain (NTD) and in the receptor binding domain (RBD). These peptides have remarkable immunogenic character (e.g., they overlap with predicted B cell epitopes) and some are present in the RBD and thus represent efficient molecules for inhibiting Group IV RNA virus infections, particularly coronavirus infections, as well as for producing protective antibodies. Vaccines against these regions would also be a very potent strategy to protect against SARS-Cov2 infection by inhibiting both PLA2G1B deleterious effect and viral infection.
Definitions
As used herein, the term “PLA2-GIB” (or “PLA2-G1B” or “PLA2G1B” “SPLA2-G1B”) designates group IB pancreatic phospholipase A2. PLA2-GIB has been identified and cloned from various mammalian species. The human PLA2-GIB protein is disclosed, for instance, in Lambeau and Gelb (2008). The sequence is available on Genbank No. NP_000919.
The amino acid sequence of an exemplary human PLA2-GIB is shown below (SEQ ID NO: 210).
MKLLVLAVLL TVAAADSGIS PRAVWQFRKM IKCVIPGSDP FLEYNNYGCY CGFGGSGTPV DEFDKCCQTH DNCYDQAKKF DSCKFFFDNP YTHTYSYSCS GSAITCSSKN KECEAFICNC DRNAAICFSK APYNKAHKNF DTKKYCQS
Amino acids 1 to 15 of SEQ ID NO: 210 (underlined) are a signal sequence, and amino acids 16 to 22 of SEQ ID NO: 210 (in bold) are a propeptide sequence.
Within the context of the invention, the term “PLA2-GIB” designates preferably human PLA2-GIB.
The human PLA2-GIB protein may be present under two distinct forms: a pro form (pro- sPLA2-GIB), which is activated by proteolytic cleavage of a pro-peptide, leading to the mature secreted form (sPLA2-GIB). The term PLA2-GIB includes any form of the protein, such as the pro-form and/or the mature form. Typically, the mature secreted form comprises the sequence of amino acid residues 23-148 of SEQ ID NO: 210, or any natural variants thereof.
Natural variants of a protein include variants resulting e.g., from polymorphism or splicing. Natural variants may also include any protein comprising the sequence of SEQ ID NO: 210, or the sequence of amino acid residues 23-148 of SEQ ID NO: 210, with one or more amino acid substitution(s), addition(s) and/or deletion(s) of one or several (typically 1, 2 or 3) amino acid residues. Variants include naturally-occurring variants having e.g., at least 90% amino acid sequence identity to SEQ ID NO: 210. Particular variants contain no more than 10 amino acid substitution(s), addition(s), and/or deletion(s) of one or several (typically 1, 2 or 3) amino acid residues as compared to SEQ ID NO: 210. Typical naturally-occurring variants retain a biological activity of PLA2- GIB. In this regard, in some embodiments, PLA2-GIB has at least one activity selected from induction of formation of membrane microdomains (MMD) in CD4 T cells from healthy subjects, or rendering CD4 T cells of healthy subjects refractory to interleukin signaling, such as refractory to IL-2 signaling or refractory to IL-7 signaling or refractory to IL-4 signaling. In some embodiments rendering CD4 T cells of healthy subjects refractory to interleukin-7 signaling comprises a reduction of STAT5A and/or B phosphorylation in said cells by at least about 10%, at least about 20%, at least about 30%, or at least about 40%. In some embodiments rendering CD4 T cells of healthy subjects refractory to interleukin-7 signaling comprises reducing the rate of nuclear translocation of phospho-STAT5A and/or phospho-STAT5B by at least about 20%, at least about 30%, at least about 40%, or at least about 50%.
The term “sequence identity” as applied to nucleic acid or protein sequences, refers to the quantification (usually percentage) of nucleotide or amino acid residue matches between at least two sequences aligned using a standardized algorithm such as Smith- Waterman alignment (Smith and Waterman (1981) J Mol Biol 147:195-197), CLUSTALW (Thompson et al. (1994) Nucleic Acids Res 22:4673-4680), or BLAST2 (Altschul et al. (1997) Nucleic Acids Res 25:3389-3402). BLAST2 may be used in a standardized and reproducible way to insert gaps in one of the sequences in order to optimize alignment and to achieve a more meaningful comparison between them.
The term “inactivation” indicates, in relation to CD4 T cells, that such cells lose at least part of their ability to contribute to the development of an effective immune response. Inactivation may be partial or complete, transient or permanent. Inactivation designates preferably reducing by at least 10%, 20%, 30%, 40%, 50%, 60%, 70%, 80% or more a function of CD4 T cells, particularly pSTAT5 nuclear translocation and/or CD4 T cell’s immunostimulatory activity. Typically, inactive CD4 T cells have no effective pSTAT5 nuclear translocation. In a particular embodiment, an inactive CD4 T cell is an anergic CD4 T cell. The term “resistance” (or “insensitivity”) of CD4 T cells to inactivation by sPLA2-GIB indicates, within the context of this invention, that CD4 T cells are essentially not inactivated in vitro when incubated in the presence of 5nM of sPLA2-GIB. Resistance indicates, for instance, that CD4 T cells retain an active nuclear translocation of pSTAT5 when incubated in vitro in the presence of 5nM sPLA2-GIB and interleukin-7. Resistance (or insensitivity) of CD4 T cells to sPLA2-GIB may also indicate that CD4 T cells incubated in vitro with 5nM PLA2-GIB remain immunologically functional, e.g., do not become anergic. The term “sPLA2-GIB Pathway” designates any component or molecule involved in sPLA2-G IB -mediated T cell anergy. This includes SPLA2-G1B, as well as any cofactor of SPLA2-G1B.
The term “treatment” of a coronavirus infection includes preventive and curative treatments. The treatment includes prevention of vims infection, such as reduction of the infection, as well as treatment of an existing infection (e.g., reduction of viral load, reduction of immune deficiency, reduction of CD4 T cell anergy) and related disease(s) or symptom(s) (e.g., pain, fever, cough, respiratory disorders, etc.). The term treatment includes a reduction of mortality caused by or associated with coronavirus infection.
The term “Group IV RNA virus” refers to the Baltimore classification and includes, without limitation, coronaviruses, such as particularly Sars-cov-2, Dengue vims, Chikungunya vims, and Polio vims.
SARS Cov-2 utilizes the sPLA2-GIB pathway
SARS-CoV-2 vims (a.k.a 2019-nCoV), is the causative agent of covid-19. There are no approved treatments against SARS-CoV-2 vims infection, and vaccines being developed are not expected to reach patients promptly. SARS-CoV-2 vims is a coronavirus, with a fully sequenced single- stranded RNA genome.
The inventors have surprisingly found that SARS-Cov-2 utilizes the sPLA2-GIB pathway. In particular, the inventors have found that SARS-Cov-2 can act by rendering CD4 T cells sensitive to inactivation by PLA2-GIB. Such mechanism involves the binding of a molecule of (or induced by) SARS-Cov-2 to gC1qR at the surface of CD4 T cells, causing sensitization of CD4 T cells to inactivation by physiological concentrations of PLA2-GIB. The inventors surprisingly found that various proteins of SARS-Cov-2, such as the spike protein, contain sequences or regions that can bind to gC1qR and may act as cofactor increasing the sPLA2-GIB inhibitory activity on CD4 T cells.
Applicant thus identified a novel general approach for treating coronavirus infection, through a modulation of the SPLA2-G1B pathway.
Treatment of Group IV RNA virus infections, particularly coronavirus infections, with an inhibitor of SPLA2-G1B
One aspect of the invention resides in the treatment of a Group IV RNA virus infection, particularly coronavirus infections using an inhibitor of PLA2-G1B.
PLA2-GIB inhibitors suitable for use in the invention may be any compound that inhibits or neutralizes the expression or activity of PLA2-GIB, such as expression inhibitors, antagonists, or sequestrators. Preferred types of inhibitors include PLA2-GIB ligands (covalent or non-covalent), anti-PLA2-GIB antibodies (and fragments and derivatives thereof), nucleic acids encoding anti-PLA2-GIB antibodies (or fragments and derivatives thereof), inhibitory nucleic acids, peptides, or small drugs, soluble receptors, or combination(s) thereof. Alternatively, or in addition, the PLA2-GIB inhibitor can be a PLA2-GIB antigen which, upon administration to the subject, induces the production of anti-PLA2GIB antibodies.
Inhibiting PLA2-GIB designates typically reducing by at least 10%, 20%, 30%, 40%, 50%, 60%, 70%, 80% or more PLA2-GIB level or activity, as well as completely blocking or suppressing PLA2-GIB level or activity. Depending on the situation, inhibition may be transient, sustained or permanent.
Antibodies against PLA2-GIB
Specific examples of PLA2-GIB inhibitors are anti-PLA2-GIB antibodies, e.g., antibodies that bind to PLA2-GIB and/or have been generated by immunization of a mammal with a PLA2-GIB antigen.
Antibodies can be synthetic, monoclonal, or polyclonal and can be made by techniques well known in the art. Such antibodies specifically bind via the antigen-binding sites of the antibody (as opposed to non-specific binding). PLA2-GIB polypeptides, fragments,
variants, fusion proteins, etc., can be employed as immunogens in producing antibodies immunoreactive therewith. More specifically, the polypeptides, fragments, variants, fusion proteins, etc. contain antigenic determinants or epitopes that elicit the formation of antibodies.
The term “antibodies” is meant to include polyclonal antibodies, monoclonal antibodies, fragments thereof, such as F(ab')2 and Fab fragments, single-chain variable fragments (scFvs), single-domain antibody fragments (VHHs or Nanobodies), bivalent antibody fragments (diabodies), as well as any recombinantly and synthetically produced binding partners, human antibodies or humanized antibodies.
Antibodies are defined to be specifically binding preferably if they bind to PLA2-GIB with a Ka of greater than or equal to about 107 M-l. Affinities of antibodies can be readily determined using conventional techniques, for example those described by Scatchard et al., Ann. N.Y. Acad. Sci., 51:660 (1949).
Polyclonal antibodies can be readily generated from a variety of sources, for example, horses, cows, donkeys, goats, sheep, dogs, chickens, rabbits, mice, or rats, using procedures that are well known in the art. In general, purified PLA2-GIB or a peptide based on the amino acid sequence of PLA2-GIB that is appropriately conjugated is administered to the host animal typically through parenteral injection. The immunogenicity of PLA2-GIB can be enhanced through the use of an adjuvant, for example, Freund's complete or incomplete adjuvant. Following booster immunizations, small samples of serum are collected and tested for reactivity to PLA2-GIB polypeptide. Examples of various assays useful for such determination include those described in Antibodies: A Laboratory Manual, Harlow and Lane (eds.), Cold Spring Harbor Laboratory Press, 1988; as well as procedures, such as countercurrent Immuno- electrophoresis (CIEP), radioimmunoassay, radio-immunoprecipitation, enzyme-linked immunosorbent assays (ELISA), dot blot assays, and sandwich assays. See U.S. Pat. Nos. 4,376, 110 and 4,486,530.
Monoclonal antibodies can be readily prepared using well known procedures. See, for example, the procedures described in U.S. Pat. Nos. RE 32,011, 4,902,614, 4,543,439, and 4,411,993; Monoclonal Antibodies, Hybridomas: A New Dimension in Biological Analyses, Plenum Press, Kennett, McKeam, and Bechtol (eds.), 1980.
For example, the host animals, such as mice, can be injected intraperitoneally at least once and preferably at least twice at about 3 week intervals with isolated and purified wild-type or mutant PLA2-GIB protein or conjugated PLA2-GIB peptide, optionally in the presence of adjuvant. Mouse sera are then assayed by conventional dot blot technique or antibody capture (ABC) to determine which animal is best to fuse. Approximately two to three weeks later, the mice are given an intravenous boost of protein or peptide. Mice are later sacrificed and spleen cells fused with commercially available myeloma cells,
such as Ag8.653 (ATCC), following established protocols. Briefly, the myeloma cells are washed several times in media and fused to mouse spleen cells at a ratio of about three spleen cells to one myeloma cell. The fusing agent can be any suitable agent used in the art, for example, polyethylene glycol (PEG). Fusion is plated out into plates containing media that allows for the selective growth of the fused cells. The fused cells can then be allowed to grow for approximately eight days. Supernatants from resultant hybridomas are collected and added to a plate that is first coated with goat anti-mouse Ig. Following washes, a label, such as a labeled PFA2-GIB polypeptide, is added to each well followed by incubation. Positive wells can be subsequently detected. Positive clones can be grown in bulk culture and supernatants are subsequently purified over a Protein A column (Pharmacia).
The monoclonal antibodies of the disclosure can be produced using alternative techniques, such as those described by Alting-Mees et al., "Monoclonal Antibody Expression Fibraries: A Rapid Alternative to Hybridomas", Strategies in Molecular Biology 3:1-9 (1990), which is incorporated herein by reference. Similarly, binding partners can be constructed using recombinant DNA techniques to incorporate the variable regions of a gene that encodes a specific binding antibody. Such a technique is described in Farrick et al., Biotechnology, 7:394 (1989).
Antigen-binding fragments of such antibodies, which can be produced by conventional techniques, are also encompassed by the present invention. Examples of such fragments include, but are not limited to, Fab and F(ab')2 fragments. Antibody fragments and derivatives produced by genetic engineering techniques are also provided.
The monoclonal antibodies include chimeric antibodies, e.g., humanized versions of murine monoclonal antibodies. Such humanized antibodies can be prepared by known techniques, and offer the advantage of reduced immunogenicity when the antibodies are administered to humans. In one embodiment, a humanized monoclonal antibody comprises the variable region of a murine antibody (or just the antigen binding site thereof) and a constant region derived from a human antibody. Alternatively, a humanized antibody fragment can comprise the antigen binding site of a murine monoclonal antibody and a variable region fragment (lacking the antigen-binding site) derived from a human antibody. Procedures for the production of chimeric and further engineered monoclonal antibodies include those described in Riechmann et al. (Nature 332:323, 1988), Liu et al. (PNAS 84:3439, 1987), Farrick et al. (Bio/Technology 7:934, 1989), and Winter and Harris (TIPS 14:139, May, 1993). Procedures to generate antibodies transgenically can be found in GB 2,272,440, U.S. Pat. Nos. 5,569,825 and 5,545,806.
Antibodies produced by genetic engineering methods, such as chimeric and humanized monoclonal antibodies, comprising both human and non-human portions, which can be made using standard recombinant DNA techniques, can be used. Such chimeric and humanized monoclonal antibodies can be produced by genetic engineering using standard
DNA techniques known in the art, for example using methods described in Robinson et al. International Publication No. WO 87/02671; Akira, et al. European Patent Application 0184187; Taniguchi, M., European Patent Application 0171496; Morrison et al. European Patent Application 0173494; Neuberger et al. PCT International Publication No. WO 86/01533; Cabilly et al. U.S. Pat. No. 4,816,567; Cabilly et al. European Patent Application 0125023; Better et al., Science 240:1041 1043, 1988; Liu et al., PNAS 84:34393443, 1987; Liu et al., J. Immunol. 139:3521 3526, 1987; Sun et al. PNAS 84:214 218, 1987; Nishimura et al., Cane. Res. 47:999 1005, 1987; Wood et al., Nature 314:446 449, 1985; and Shaw et al., J. Natl. Cancer Inst. 80:1553 1559, 1988); Morrison, S. L., Science 229:1202 1207, 1985; Oi et al., BioTechniques 4:214, 1986; Winter U.S. Pat. No. 5,225,539; Jones et al., Nature 321:552 525, 1986; Verhoeyan et al., Science 239:1534, 1988; and Beidler et al., J. Immunol. 141:4053 4060, 1988.
In connection with synthetic and semi-synthetic antibodies, such terms are intended to cover but are not limited to antibody fragments, isotype switched antibodies, humanized antibodies (e.g., mouse-human, human-mouse), hybrids, antibodies having plural specificities, and fully synthetic antibody-like molecules.
Human monoclonal antibodies having human constant and variable regions can be generated by immunizing transgenic animals which contain human immunoglobulin genes. See Jakobovits et al. Ann NY Acad Sci 764:525-535 (1995). Human monoclonal antibodies against PLA2-GIB polypeptides can also be prepared by constructing a combinatorial immunoglobulin library, such as a Fab phage display library or a scFv phage display library, using immunoglobulin light chain and heavy chain cDNAs prepared from mRNA derived from lymphocytes of a subject. See, e.g., McCafferty et al. PCT publication WO 92/01047; Marks et al. (1991) J. Mol. Biol. 222:581 597; and Griffths et al. (1993) EMBO J 12:725734. In addition, a combinatorial library of antibody variable regions can be generated by mutating a known human antibody. For example, a variable region of a human antibody known to bind PLA2-GIB, can be mutated by, for example, using randomly altered mutagenized oligonucleotides, to generate a library of mutated variable regions which can then be screened to bind to PLA2-GIB. Methods of inducing random mutagenesis within the CDR regions of immunoglobin heavy and/or light chains, methods of crossing randomized heavy and light chains to form pairings and screening methods can be found in, for example, Barbas et al. PCT publication WO 96/07754; Barbas et al. (1992) Proc. Nat'l Acad. Sci. USA 89:4457 4461.
An immunoglobulin library can be expressed by a population of display packages, preferably derived from filamentous phage, to form an antibody display library. Examples of methods and reagents particularly amenable for use in generating antibody display library can be found in, for example, Ladner et al. U.S. Pat. No. 5,223,409; Kang et al. PCT publication WO 92/18619; Dower et al. PCT publication WO 91/17271; Winter et al. PCT publication WO 92/20791; Markland et al. PCT publication WO 92/15679; Breitling et al. PCT publication WO 93/01288; McCafferty et al. PCT publication WO
92/01047; Garrard et al. PCT publication WO 92/09690; Ladner et al. PCT publication WO 90/02809; Fuchs et al. (1991) Bio/Technology 9:1370 1372; Hay et al. (1992) Hum Antibod Hybridomas 3:81 85; Huse et al. (1989) Science 246:1275 1281; Griffths et al. (1993) supra; Hawkins et al. (1992) J Mol Biol 226:889 896; Clackson et al. (1991) Nature 352:624 628; Gram et al. (1992) PNAS 89:3576 3580; Garrad et al. (1991) Bio/Technology 9:1373 1377; Hoogenboom et al. (1991) Nuc Acid Res 19:4133 4137; and Barbas et al. (1991) PNAS 88:7978 7982. Once displayed on the surface of a display package (e.g., filamentous phage), the antibody library is screened to identify and isolate packages that express an antibody that binds a PLA2-GIB polypeptide. In a preferred embodiment, the primary screening of the library involves panning with an immobilized PLA2-GIB polypeptide and display packages expressing antibodies that bind immobilized PLA2-GIB polypeptide are selected.
Preferred antibodies for use in the invention are directed to a PLA2-GIB epitope, and/or have been generated by immunization with a polypeptide comprising a PLA2-GIB epitope selected from: the mature PLA2-GIB protein, a fragment of PLA2-GIB comprising at least 8 consecutive amino acid residues of SEQ ID NO: 210 (or the corresponding residues of a natural variant of SEQ ID NO: 210), said fragment preferably comprising at least amino acid 70, amino acid 121, amino acid 50, amino acid 52, amino acid 54, amino acid 71, or a combination thereof (numbering by reference to SEQ ID NO: 210).
Particular anti-PLA2-GIB antibodies for use in the invention bind mature human PLA2- GIB, even more preferably an epitope comprised in a domain of PLA2-GIB comprising an amino acid residue selected from amino acid 70, amino acid 121, amino acid 50, amino acid 52, amino acid 54, amino acid 71, or a combination thereof (numbering by reference to SEQ ID NO: 210). Particular antibodies for use in the invention bind an epitope comprised between amino acid residues 50-71 of mature human PLA2-GIB (by reference to SEQ ID NO: 210) or the corresponding residues of a natural variant of SEQ ID NO: 210. Examples of anti-PLA2-GIB antibodies suitable for use in the invention have been disclosed in W02015/097140.
Further particular anti-PLA2-GIB antibodies for use in the present invention bind an epitope comprising at least one amino acid residue selected from W3, R6, K7, K10, C77, Y75, G79 and S80 of human mature PLA2-GIB (numbering by reference to SEQ ID NO: 210), more preferably an epitope comprising at least 2 or at least 3 amino acid residues selected from W3, R6, K7, K10, C77, Y75, G79 and S80 of human mature PLA2-GIB, further more preferably an epitope comprising at least 4, at least 5, at least 6 or at least 7 amino acid residues selected from W3, R6, K7, K10, C77, Y75, G79 and S80 of human mature PLA2-GIB. Particular antibodies for use in the invention bind an epitope comprising an amino acid residue comprised between amino acids 1-10 or 75-80 of mature human PLA2-GIB (by reference to SEQ ID NO: 210) or the corresponding residues of a natural variant of SEQ ID NO: 210. Such antibodies exhibit potent
neutralizing activity and represent valuable therapeutic agents for use in the invention. Examples of such anti-PLA2-GIB antibodies suitable for use in the invention have been disclosed in WO2019/166665, incorporated therein by reference. In a particular embodiment, the antibodies or derivatives for use in the invention is monoclonal antibody 14G9 as disclosed in WO2019/166665, or an anti-PLA2-GIB antibody that competitively inhibits binding of monoclonal antibody 14G9 to human PLA2-GIB. The antibody may be human or humanized. In another particular embodiment, the antibodies or derivatives for use in the invention is monoclonal antibody #2B as disclosed in WO2019/166665, or an anti-PLA2-GIB antibody that competitively inhibits binding of monoclonal antibody #2B to human SPLA2-GIB. In another particular embodiment, the antibodies or derivatives for use in the invention is monoclonal antibody #2B1 as disclosed in WO2019/166665, or an anti-PLA2-GIB antibody that competitively inhibits binding of monoclonal antibody #2B1 to human SPLA2-GIB.
In another particular embodiment, the antibodies or derivatives for use in the invention is monoclonal antibody #2B2 as disclosed in WO2019/166665, or an anti-PLA2-GIB antibody that competitively inhibits binding of monoclonal antibody #2B2 to human SPLA2-GIB.
The term “competitively inhibits” indicates that the antibody can reduce or inhibit or displace the binding of a reference antibody to sPLA2-GIB. Competition assays can be performed using standard techniques such as, for instance, competitive ELISA or other binding assays. Typically, a competitive binding assay involves a purified target antigen, generally bound either to a solid substrate or cells, an unlabeled test antibody and a labeled reference antibody. Competitive inhibition is measured by determining the amount of labeled antibody bound in the presence of the test antibody. Usually the test antibody is present in excess, such as about 5 to 500 times the amount of reference antibody. Typically, for ELISA, the test antibody is in 100X excess, and for enzymatic methods, the test antibody in in 10X excess. When a test antibody present in excess inhibits or displaces at least 70% of the binding of the reference antibody to the antigen, it is considered as competitively inhibiting said reference antibody. In a specific embodiment, when a test antibody present in 100X excess inhibits or displaces at least 70%, more preferably at least 80% of the binding of the reference antibody to the antigen in ELISA, it is considered as competitively inhibiting said reference antibody. Preferred competing antibodies bind epitopes that share common amino acid residues.
In a particular embodiment, the inhibitor is a monoclonal antibody comprising:
(i) a light chain variable region comprising a CDR-L1, a CDR-L2, a CDR-L3 and a FR- L, wherein the CDR-L1, CDR-L2 and/or CDR-L3 consists, or consists essentially, of the CDR-L1, CDR-L2 and CDR-L3, respectively, of the light chain variable region of SEQ ID NO: 3 or 5 as disclosed in WO2019/166665, and wherein a FR-L is of a human immunoglobulin sequence; and
(ii) a heavy chain variable region comprising a CDR-H1, a CDR-H2, a CDR-H3 and a FR-H, wherein the CDR-H1, CDR-H2 and/or CDR-H3 consists, or consists essentially, of the CDR-H1, CDR-H2 and CDR-H3, respectively, of the heavy chain variable region of SEQ ID NO: 4, 6, 7 or 8 as disclosed in WO2019/166665, and wherein a FR-H is of a human immunoglobulin sequence.
The “variable region” of an antibody refers to the amino-terminal domains of the heavy or light chain (“VH” or “VL”), which contain the antigen-binding sites. A light or heavy chain variable region (VL or VH) generally consists of a framework region (“FR”) interrupted by three hypervariable regions referred to as "complementarity determining regions" or "CDRs". The extent of the framework region and CDRs have been precisely defined, for example as in Rabat (see "Sequences of Proteins of Immunological Interest," E. Rabat et al., U.S. Department of Health and Human Services, (1983)), and as in Chothia.
With reference to SEQ ID NO: 3 as disclosed in WO2019/166665, the three CDR regions correspond to the following amino acid residues:
. CDR-L1: amino acid residues QDVSTA (residues 27-31 of SEQ ID NO: 3),
. CDR-L2: amino acid residues WAS (residues 50-52 of SEQ ID NO: 3),
. CDR-L3: amino acid residues QQDYSTPPT (residues 89-97 of SEQ ID NO: 3).
With reference to SEQ ID NO: 4 as disclosed in WO2019/166665, the three CDR regions correspond to the following amino acid residues:
. CDR-H1: amino acid residues GYTFTNYW (residues 26-33 of SEQ ID NO: 4),
. CDR-H2: amino acid residues IDPSDTRT (residues 51-58 of SEQ ID NO: 4),
. CDR-H3: amino acid residues ARQTLYYEALDY (residues 97-108 of SEQ ID NO: 4).
In a particular embodiment, the invention uses a monoclonal antibody selected from:
. monoclonal antibody 14G9;
. monoclonal antibody #2;
. monoclonal antibody #2B 1 ;
. monoclonal antibody #2B2; as disclosed in WO2019/166665, and . derivatives thereof.
The term “antibody derivative”, as used herein, refers to an antibody which retains the antigenic specificity of a reference antibody but wherein one or more amino acid residues are (chemically, or biologically) modified to improve its properties.
Examples of such chemical modifications include, e.g. by alkylation, PEGylation, acylation, ester or amide formation, and the like. In particular, a derivative is an antibody as disclosed herein that is modified to contain one or more additional non-pro teinaceous moieties such as water-soluble polymers. Examples of water-soluble polymers include, but are not limited to, PEG, copolymers of ethylene glycol/propylene glycol, carboxymethylcellulose, dextran and polyvinyl alcohol.
Derivatives may also be generated to increase or decrease the extent to which the antibody is glycosylated. Addition or deletion of glycosylation sites to an antibody may be conveniently accomplished by altering the amino acid sequence such that one or more glycosylation sites is created or removed. Where the antibody comprises an Fc region, the carbohydrate attached thereto may be altered. Native antibodies produced by mammalian cells typically comprise a branched, biantennary oligosaccharide that is generally attached by an N-linkage to Asn297 of the CH2 domain of the Fc region (see e.g., Wright et al. TIBTECH, 1997, 15:26-32). The oligosaccharide may include various carbohydrates, e.g., mannose, N-acetyl glucosamine (GlcNAc), galactose, and sialic acid, as well as a fucose attached to a GlcNAc in the "stem" of the biantennary oligosaccharide structure. In some embodiments, modifications of the oligosaccharide in an antibody of the invention may be made in order to create antibody variants with certain improved properties.
In one embodiment, antibody variants are provided having a carbohydrate structure that lacks fucose attached (directly or indirectly) to an Fc region. For example, the amount of fucose in such antibody may be from 1% to 80%, from 1% to 65%, from 5% to 65% or from 20% to 40%). The amount of fucose is determined by calculating the average amount of fucose within the sugar chain at Asn297, relative to the sum of all glycostructures attached to Asn 297 (e. g. complex, hybrid and high mannose structures) as measured by MALDI-TOF mass spectrometry. Asn297 refers to the asparagine residue located at about position 297 in the Fc region (Eu numbering of Fc region residues); however, Asn297 may also be located about ± 3 amino acids upstream or downstream of position 297, i.e., between positions 294 and 300, due to minor sequence variations in antibodies. Examples of publications related to "defucosylated" or "fucose-deficient" antibody variants include, but are not limited to, Okazaki et al. J. Mol. Biol. 336: 1239- 1249 (2004) and Yamane-Ohnuki N, Satoh M. mAbs. 2009;1:230-236. Examples of cell lines capable of producing defucosylated antibodies include Led 3 CHO cells deficient in protein fucosylation (Ripka et al. Arch. Biochem. Biophys. 249:533-545 (1986)), and knockout cell lines, such as alpha- 1 ,6-fucosyltransferase gene, FUT8, knockout CHO cells (see, e.g., Yamane-Ohnuki et al. Biotech. Bioeng. 87: 614 (2004); Kanda, Y. et al., Biotechnol. Bioeng., 94(4):680-688 (2006)).
In certain embodiments, it may be desirable to create cysteine engineered antibodies, e.g., "thioMAbs," in which one or more residues of an antibody are substituted with cysteine residues. In particular embodiments, the substituted residues occur at accessible sites of the antibody. By substituting those residues with cysteine, reactive thiol groups are thereby positioned at accessible sites of the antibody and may be used to conjugate the
antibody to other moieties, such as drug moieties or linker-drug moieties, to create an immunoconjugate, as described further herein. In certain embodiments, any one or more of the following residues may be substituted with cysteine: V205 (Kabat numbering) of the light chain; A118 (EU numbering) of the heavy chain; and S400 (EU numbering) of the heavy chain Fc region. Cysteine engineered antibodies may be generated as described, e.g., in U.S. Patent No. 7,521,541.
The term derivative also includes immunoconjugates comprising an anti-sPLA2-GIB antibody as defined above conjugated to one or more heterologous molecule(s), including but not limited to a cytotoxic agent, a detectable moiety such as a fluorescent moiety, a diagnostic radioisotope or an imaging agent; or to a solid support, such as agarose beads or the like. Examples of cytotoxic agents include, but are not limited to chemotherapeutic agents or drugs, growth inhibitory agents, toxins (e.g., protein toxins, enzymatically active toxins of bacterial, fungal, plant, or animal origin, or fragments thereof), or radioactive isotopes. Conjugates of an antibody and cytotoxic agent may be made using a variety of bifunctional protein coupling agents well known by the skilled person. The linker may be a "cleavable linker" facilitating release of a cytotoxic drug in the cell. For example, an acid-labile linker, peptidase-sensitive linker, photolabile linker, dimethyl linker or disulfide-containing linker (Chari et al., Cancer Res. 52: 127-131 (1992)) may be used.
The antibodies for use in the invention are typically “isolated”, e.g., have been separated from at least one component of their natural environment. In particular, the antibodies may be purified to greater e.g., at least 95%, at least 96%; at least 97%, at least 98% or at least 99% purity as determined by, for example, electrophoretic (e.g., SDS-PAGE, isoelectric focusing (IEF), capillary electrophoresis) or chromatographic (e.g., ion exchange or reverse phase HPLC) techniques. For review of methods for assessment of antibody purity, see, e.g. Flatman et al., J. Chromatogr. B 848:79-87 (2007).
Preferred antibodies of the invention are essentially neutralizing antibodies, i.e., they are able to at least partially inhibit an activity of PLA2-GIB . sPLA2-GIB catalyzes the hydrolysis of the sn-2 fatty acyl bond of phospholipids to release free fatty acids and lysophospholipids. Particular antibodies of the invention inhibit an enzymatic activity of sPLA2-GIB, such as the hydrolysis of the sn-2 fatty acyl bond of phospholipids. Methods for testing such a property are disclosed in detail in the experimental section. Particular antibodies for use in the invention inhibit binding of sPLA2-GIB to a substrate thereof. Further particular antibodies for use in the invention inhibit sPLA2-GIB -mediated inhibition of IF-7-induced phospho-STAT5 nuclear translocation in CD4 T cells. Methods for testing such a property are disclosed in detail in the experimental section.
The neutralizing activity of the antibody or derivative can be determined in vitro or in vivo using e.g., binding or biological assays, such as tests as described in the experimental
section. Inhibition/neutralization may be complete or partial. In particular, the antibodies may inhibit 10% or more of the tested activity, preferably 20% or more, 30% or more, 40% or more, 50% or more.
In preferred embodiments, the antibodies are IgG, e.g., gGl, IgG2, IgG3 or IgG4.
The antibodies or derivatives may be isolated and preserved using conventional methods and media. They may be lyophilized. They may also be frozen.
Nucleic acids, vectors and host cells encoding recombinant antibodies
In another aspect, the PLA2-GIB inhibitor is or comprises or consists of a nucleic acid molecule encoding an anti-PLA2-GIB antibody, or a light or heavy chain thereof, or a variable domain thereof, or a nucleic acid complementary to said encoding sequence. The nucleic acid can be DNA (cDNA or gDNA), RNA, or a mixture thereof. It can be in single stranded form or in duplex form or a mixture of the two. It can comprise modified nucleotides, comprising for example a modified bond, a modified purine or pyrimidine base, or a modified sugar. It can be prepared by any method known to one skilled in the art, including chemical synthesis, recombination, and mutagenesis. The nucleic acid according to the invention may be deduced from the sequence of the antibody according to the invention and codon usage may be adapted according to the host cell in which the nucleic acid shall be transcribed. These steps may be carried out according to methods well known to one of skill in the art and some of which are described in the reference manual Sambrook et al. (Sambrook J, Russell D (2001) Molecular cloning: a laboratory manual, Third Edition Cold Spring Harbor).
The nucleic acid may encode an amino acid sequence comprising the light chain and/or an amino acid sequence comprising the heavy chain of the antibody, or may be complementary to such encoding sequence.
The present invention further provides a vector comprising a nucleic acid of the invention. Optionally, the vector may comprise several nucleic acids of the invention. In particular, the vector may comprise a nucleic acid of the invention operably linked to a regulatory region, i.e. a region comprising one or more control sequences. Optionally, the vector may comprise several nucleic acids of the invention operably linked to several regulatory regions.
The term "control sequences" means nucleic acid sequences necessary for expression of a coding region. Control sequences may be endogenous or heterologous. Well-known control sequences and currently used by the person skilled in the art will be preferred. Such control sequences include, but are not limited to, promoter, signal peptide sequence and transcription terminator.
The term "operably linked" means a configuration in which a control sequence is placed at an appropriate position relative to a coding sequence, in such a way that the control sequence directs expression of the coding region.
The present invention further relates to the use of a nucleic acid or vector according to the invention to transform, transfect or transduce a host cell.
The present invention also provides a host cell comprising one or several nucleic acids of the invention and/or one or several vectors of the invention. The term "host cell" also encompasses any progeny of a parent host cell that is not identical to the parent host cell due to mutations that occur during replication.
Suitable host cells for cloning or expression of antibody-encoding vectors include prokaryotic or eukaryotic cells such as bacteria, yeasts, insect cells, mammalian cells, etc. Inhibitory Nucleic acids
In another embodiment, the PLA2-GIB inhibitor is an inhibitory nucleic acid, i.e., any nucleic acid molecule which inhibits PLA2-GIB gene or protein expression. Preferred inhibitory nucleic acids include antisense nucleic acids, short interfering RNAs (siRNAs), small hairpin RNAs (shRNA), microRNAs, aptamers, or ribozymes. In a particular embodiment, the inhibitory nucleic acid is a small interfering RNA that prevents translation of PLA2-GIB mRNA. In another particular embodiment, the inhibitory nucleic acid is an antisense oligonucleotide that prevents translation of PLA2-GIB mRNA. In another particular embodiment, the inhibitory nucleic acid is a small hairpin RNA that prevents translation of PLA2-GIB mRNA. siRNA comprise a sense nucleic acid sequence and an anti-sense nucleic acid sequence of the polynucleotide of interest. siRNA are constructed such that a single transcript (double stranded RNA) have both the sense and complementary antisense sequences from the target gene. The nucleotide sequence of siRNAs may be designed using an siRNA design computer program available from, for example, the Ambion website on the world wide web.
In some embodiments, the length of the antisense oligonucleotide or siRNAs is less than or equal to 10 nucleotides. In some embodiments, the length of the antisense oligonucleotides and siRNAs is as long as the naturally occurring transcript. In some embodiments, the antisense oligonucleotides and siRNAs have 18-30 nucleotides. In some embodiments, the antisense oligonucleotides and siRNAs are less than 25 nucleotides in length.
Preferred inhibitory nucleic acid molecules comprise a domain having a nucleotide sequence that is perfectly complementary to a region of a PLA2-GIB gene or RNA. Such a domain contains typically from 4 to 20 nucleotides, allowing specific hybridization and optimal inhibition of the gene transcription or RNA translation. The sequence of the inhibitory nucleic acids may be derived directly from the sequence of a gene encoding PLA2-GIB. Alternatively, or in addition, inhibitory nucleic acids may hybridize to a regulatory element in a PLA2-GIB gene or RNA, such as a promoter, a splicing site, etc., and prevent effective regulation thereof.
Specific examples of inhibitory nucleic acid molecules of the present invention include isolated single strand nucleic acid molecules consisting of from 10 to 50 consecutive
nucleotides of a sequence encoding SEQ ID NO: 210. Specific examples of inhibitory nucleic acid molecules of the invention are antisense nucleic acids consisting of the following nucleotide sequence or the perfectly complementary strand thereof:
ATGAAACTCCTTGTGCTAG (SEQ ID NO: 212)
ACAGCGGCATCAGC (SEQ ID NO: 213)
TTCCGCAAAATGATCAA (SEQ ID NO: 214)
CCCGGGGAGTGACCCC (SEQ ID NO: 215)
TACGGCTGCTACTGTGGCTT (SEQ ID NO: 216) GACACATGACAACTGCTACGACC (SEQ ID NO: 217) ACCCACACCTATTCATACTCGT (SEQ ID NO: 218)
ATCACCTGTAGCAGCA (SEQ ID NO: 219)
AGCTCCATATAACAAGGCA (SEQ ID NO: 220)
CAAGAAGTATTGTCAGAG (SEQ ID NO: 221)
Peptides and Small Drugs
In an alternative embodiment, the PLA2-GIB inhibitor is a peptide or small drug that inhibits the activity of PLA2-GIB. The peptide or small drug is typically a molecule that selectively binds PLA2-GIB, or a substrate of PLA2-GIB, or a co-factor of PLA2-GIB, or a degradation product or metabolite of PLA2-GIB pathway.
Peptides preferably contain from 3 to 20 amino acid residues, and their sequence may be identical to a domain of PLA2-GIB (bait peptide) or to a domain of a PLA2-GIB substrate, co-factor, degradation product or metabolite. Preferred peptides of the invention contain from 4 to 30 consecutive amino acid residues of SEQ ID NO: 210 (or of a corresponding sequence of a natural variant of SEQ ID NO: 210). Most preferred peptides of the invention comprise from 5 to 25 consecutive amino acid residues of SEQ ID NO: 210 (or of a corresponding sequence of a natural variant of SEQ ID NO: 210) and further comprise at least one of the following amino acid residues of SEQ ID NO: 210 (or of a corresponding sequence of a natural variant of SEQ ID NO: 210): amino acid 3, amino acid 6, amino acid 7, amino acid 10, amino acid 70, amino acid 121, amino acid 50, amino acid 52, amino acid 54, amino acid 71, amino acid 75, amino acid 77, amino acid 79, amino acid 80, or a combination thereof. Specific examples of peptides of the invention are peptides of less than 25 amino acids comprising anyone of the following sequences:
NNYGCY (SEQ ID NO: 222)
CYCGLG (SEQ ID NO: 223)
YNNY GC YCGLGGS G (SEQ ID NO: 224)
FLE YNNY GC Y CGLGGS GTPV (SEQ ID NO: 225)
QTHDN (SEQ ID NO: 226)
CQTHDNC (SEQ ID NO: 227)
ECEAFICNC (SEQ ID NO: 228)
DRNAAI (SEQ ID NO: 229)
DRNAAICFSKAPYNKAHKNL (SEQ ID NO: 230)
Other peptides for use in the invention include a pentapeptide as disclosed in WO20 17/060405, incorporated therein by reference. In a specific embodiment, the compound is a cyclic peptide selected from FLSYK (SEQ ID NO: 231), FLSYR (SEQ ID NO: 232) and (2NapA)LS(2NapA)R (SEQ ID NO: 233).
The peptides of the invention can comprise peptide, non-peptide and/or modified peptide bonds. In a particular embodiment, the peptides comprise at least one peptidomimetic bond selected from intercalation of a methylene (-CH2-) or phosphate (-PO2-) group, secondary amine (-NH-) or oxygen (-0-), alpha-azapeptides, alpha-alkylpeptides, N- alkylpeptides, phosphonamidates, depsipeptides, hydroxymethylenes, hydroxyethylenes, dihydroxyethylenes, hydroxyethylamines, retro-inverso peptides, methyleneoxy, cetomethylene, esters, phosphinates, phosphinics, or phosphonamides. Also, the peptides may comprise a protected N-ter and/or C-ter function, for example, by acylation, and/or amidation and/or esterification.
The peptides of the invention may be produced by techniques known per se in the art such as chemical, biological, and/or genetic synthesis.
Each of these peptides, in isolated form, represents a particular object of the present invention.
Preferred small drugs are hydrocarbon compounds that selectively bind PLA2-GIB.
Examples of small drugs include indole compounds, such as those disclosed in W02017/037041, incorporated therein by reference. In a particular embodiment, the compound is 3-(2-amino-l,2-dioxoethyl)-2-ethyl-l-(phenylmethyl)-lH-indol-4- yl)oxy)acetic acid or a pharmaceutically acceptable salt, hydrate, or prodrug thereof, such as a sodium salt thereof (Varespladib).
Small drugs and peptides are preferably obtainable by a method comprising: (i) contacting a test compound with PLA2-GIB or a fragment thereof, (ii) selecting a test compound which binds PLA2-GIB or said fragment thereof, and (iii) selecting a compound of (ii) which inhibits an activity of PLA2-GIB. Such a method represents a particular object of the invention.
Small drugs and peptides are also obtainable by a method comprising: (i) contacting a test compound with a PLA2-GIB substrate, co-factor, or degradation product, or a fragment thereof, (ii) selecting a test compound which binds to said PLA2-GIB substrate, co-factor, or degradation product, or a fragment thereof, and (iii) selecting a compound of (ii) which inhibits an activity of PLA2-GIB. Such a method represents a particular object of the invention.
Vaccination
In an alternative (or cumulative) embodiment, the PLA2-GIB inhibitor is a PLA2-GIB antigen. As a result of a vaccination or immunization of the subject with said antigen, the subject produces antibodies (or cells) which inhibit PLA2-GIB . In particular, injection(s) of a PLA2-GIB antigen (e.g., an immunogenic PLA2-GIB essentially devoid of biological activity) can generate antibodies in the treated subject. These antibodies will protect against an excess of PLA2-GIB expression and can be used along as immunotherapy or a vaccine prophylaxis. An object of the invention thus resides in a method of treating a solid cancer in a subject having solid cancer, comprising administering to the subject a PLA2-GIB antigen.
A further object of the invention relates to a PLA2-GIB antigen for use to treat a solid cancer in a subject in need thereof.
In a particular embodiment, the PLA2-GIB antigen is an inactivated immunogenic molecule that induces an immune response against PLA2-GIB in a subject. Inactivation may be obtained e.g., by chemically or physically altering PLA2-GIB or by mutating or truncating the protein, or both; and immunogenicity may be obtained as a result of the inactivation and/or by further conjugating the protein to a suitable carrier or hapten, such as KLH, HSA, polylysine, a viral anatoxin, or the like, and/or by polymerization, or the like. The antigen may thus be chemically or physically modified, e.g., to improve its immunogenicity .
In a preferred embodiment, the PLA2-GIB antigen comprises PLA2-GIB or an epitope- containing fragment or mimotope thereof.
In a particular embodiment, the PLA2-GIB antigen comprises a full length PLA2-GIB protein. In a further particular embodiment, the PLA2-GIB antigen comprises a protein comprising SEQ ID NO: 210, or a sequence having at least 90% identity to SEQ ID NO: 210.
In an alternative embodiment, the PLA2-GIB antigen comprises a fragment of a PLA2- GIB protein comprising at least 6 consecutive amino acid residues and containing an immunogenic epitope, or a mimotope thereof. In a preferred embodiment, the PLA2-GIB antigen comprises at least from 6 to 20 amino acid residues. Preferred peptides of the invention contain from 4 to 30 consecutive amino acid residues of SEQ ID NO: 210 (or of a corresponding sequence of a natural variant of SEQ ID NO: 210). Most preferred
peptides of the invention comprise from 5 to 25 consecutive amino acid residues of SEQ ID NO: 210 (or of a corresponding sequence of a natural variant of SEQ ID NO: 210) and further comprise at least one of the following amino acid residues of SEQ ID NO: 210 (or of a corresponding sequence of a natural variant of SEQ ID NO: 210): amino acid 3, 6, 7, 10, 70, 121, 50, 52, 54, 71, 75, 77, 79, 80, or a combination thereof. Specific examples of such antigens are the peptides of SEQ ID NO: 222-230 as disclosed above.
The PLA2-GIB antigen may be in various forms such as in free form, polymerized, chemically or physically modified, and/or coupled (i.e., linked) to a carrier molecule. Coupling to a carrier may increase the immunogenicity and (further) suppress the biological activity of the PLA2-GIB polypeptide. In this regard, the carrier molecule may be any carrier molecule or protein conventionally used in immunology such as for instance KLH (Keyhole limpet hemocyanin), ovalbumin, bovine serum albumin (BSA), a viral or bacterial anatoxin such as toxoid tetanos, toxoid diphteric B cholera toxin, mutants thereof such as diphtheria toxin CRM 197, an outer membrane vesicle protein, a polylysine molecule, or a virus like particle (VLP). In a preferred embodiment, the carrier is KLH or CRM 197 or a VLP.
Coupling of PLA2-GIB to a carrier may be performed by covalent chemistry using linking chemical groups or reactions, such as for instance glutaraldehyde, biotin, etc. Preferably, the conjugate or the PLA2-GIB protein or fragment or mimotope is submitted to treatment with formaldehyde in order to complete inactivation of PLA2-GIB.
In a particular embodiment, the PLA2-GIB antigen comprises a full length PLA2-GIB protein, optionally coupled to a carrier protein. In a preferred embodiment, the PLA2- GIB antigen comprises a protein comprising SEQ ID NO: 210, or a sequence having at least 90% identity to SEQ ID NO: 210, coupled to a carrier protein.
In another particular embodiment, the PLA2-GIB antigen comprises an immunogenic peptide or mimotope of PLA2-GIB, optionally coupled to a carrier protein. In a more preferred embodiment, the PLA2-GIB antigen comprises a polypeptide of at least 10 amino acids long comprising at least one of the following amino acid residues of SEQ ID NO: 210 (or of a corresponding sequence of a natural variant of SEQ ID NO: 210): amino acid 70, amino acid 121, amino acid 50, amino acid 52, amino acid 54, amino acid 71, or a combination thereof, optionally coupled to a carrier molecule.
The immunogenicity of the PLA2-GIB antigen may be tested by various methods, such as by immunization of a non-human animal grafted with human immune cells, followed by verification of the presence of antibodies, or by sandwich ELISA using human or humanized antibodies. The lack of biological activity may be verified by any of the activity tests described in the application. In a preferred embodiment, the PLA2-GIB antigen has less than 20%, more preferably less than 15%, 10%, 5% or even 1% of the activity of a wild-type PLA2-GIB protein in an in vitro method of (i) induction of formation of membrane microdomains (MMD) in CD4 T cells or (ii) in rendering CD4 T cells refractory to IL-2 signaling or refractory to IL-7 signaling.
Such molecules and conjugates and vaccines represent potent agents for use to immunize subjects, thereby causing a sustained PLA2-GIB inhibition. Upon repetition, such methods can be used to cause a permanent PLA2-GIB inhibition.
Treatment of Group IV RNA virus infection, particularly corona virus infection, with an inhibitor of a sPLA2-G1B cofactor
The inventors found that various proteins of SARS-Cov-2, such as the spike protein, can act as cofactor of PLA2-G1B. Any strategy based on an inhibition of such cofactor effect can be used to treat a Group IV RNA vims infection, particularly coronavirus infections, more particularly SARS-Cov-2 infection.
It is thus an object of the invention to provide methods and compositions for treating a Group IV RNA vims infection, particularly a coronavirus infection in a mammalian subject, comprising administering to the subject an inhibitor of a PLA2-GIB cofactor.
The invention also relates to an inhibitor of a PLA2-GIB cofactor, for use for treating a Group IV RNA vims infection, particularly a coronavirus infection in a subject in need thereof.
The invention also relates to the use of an inhibitor of a PLA2-GIB cofactor, for the manufacture of a medicament for treating a Group IV RNA vims infection, particularly a coronavirus infection in a subject in need thereof.
PLA2-GIB cofactors
The inventors have surprisingly discovered that SARS-Cov-2 can act as (or produce) a cofactor of PLA2-GIB that, in combination with PLA2-GIB, leads to CD4 T cell inactivation.
Within the context of the invention, the term “cofactor” of PLA2-GIB designates any molecule or agent which potentiates or amplifies or mediates an effect of PLA2-GIB, particularly an effect of PLA2-GIB on CD4 T cells. Preferred cofactors are molecules which can sensitize CD4 T cells to inactivation by low concentrations of PLA2-GIB.
In a particular embodiment, the PLA2-GIB cofactor is a ligand of gC1qR. The inventors have indeed demonstrated that ligands of gC1qR at the surface of CD4 T cells act as cofactors of PLA2-GIB, rendering cells more sensitive to inactivation by PLA2-GIB. More particularly, the PLA2-GIB cofactor is an agonist of gC1qR, e.g., can induce signaling through gC1qR, more particularly can induce gC1qR- mediated exocytosis.
In this respect, the inventors have identified various proteins which can act as cofactor of PLA2-GIB, as listed in Table 3. In particular, such proteins include ORFlab, the Spike protein, ORF3, E protein, ORF7b, ORF8, N protein and M protein.
Each of said proteins thus represents a relevant target for inhibition according to the invention.
The inventors also identified various peptides from said proteins which can bind gC1qR, as listed in Table 3. Each of said peptides represents an object of the invention.
Each of said proteins and peptides can act as PLA2-GIB cofactors in SARS-Cov2 infected patients.
Each of said peptides represents a strong inhibitor (or be used to construct a strong inhibitor) for use according to the invention
Each of said proteins or peptides can be used to produce relevant inhibitors, such as antibodies or derivatives thereof, or immunogens for vaccination.
Each of said proteins and peptides can be used as a reagent to detect the presence of anti- Cov-2 antibodies in a sample, particularly of a protective immunity.
In a particular embodiment of the invention, the PLA2-GIB cofactor is a protein selected from ORFlab, the Spike protein, ORF3, E protein, ORF7b, ORF8, N protein and M protein of SARS-Cov-2, or a gC1qR-binding element of such proteins, such as listed in Table 3. More particularly, the cofactor may be any protein comprising anyone of SEQ ID NOs: 2-209, or any fragment or mimotope thereof.
The term “fragment”, in relation to such cofactors, designates preferably a fragment containing a gC1qR-binding element of such a protein, and/or a fragment retaining a capacity of binding gC1qR. Preferred fragments contain at least 5 consecutive amino acid residues, typically between 5 and 100, between 5-80, between 5-60, between 5-50, between 5-40, between 5-30, between 5-25, or between 5-20, or even less.
In a specific embodiment, the PLA2-GIB cofactor is SARS-Cov2 spike (or S) protein, or a fragment or mimotope thereof. The amino acid sequence of a Spike protein is depicted as SEQ ID NO: 209. Such sequence is also available as GenBank reference QHD43416.1. In a particular embodiment, the PLA2-GIB cofactor is a protein or peptide comprising or consisting of the amino acid sequence of any one of SEQ ID NOs: 156-185 and 209, or a fragment or mimotope thereof.
In another specific embodiment, the PLA2-GIB cofactor is SARS-Cov2 ORFlab protein, or a fragment or mimotope thereof. In a particular embodiment, the PLA2-GIB cofactor is a protein or peptide comprising or consisting of the amino acid sequence of any one of SEQ ID NOs: 2-155, or a fragment or mimotope thereof.
In another specific embodiment, the PLA2-GIB cofactor is SARS-Cov2 ORF3 protein, or a fragment or mimotope thereof. In a particular embodiment, the PLA2-GIB cofactor is a protein or peptide comprising or consisting of the amino acid sequence of any one of SEQ ID NOs: 186-193, or a fragment or mimotope thereof.
In another specific embodiment, the PLA2-GIB cofactor is SARS-Cov2 protein E, or a fragment or mimotope thereof. In a particular embodiment, the PLA2-GIB cofactor is a protein or peptide comprising or consisting of the amino acid sequence of SEQ ID NO: 194, or a fragment or mimotope thereof.
In another specific embodiment, the PLA2-GIB cofactor is SARS-Cov2 protein M, or a fragment or mimotope thereof. In a particular embodiment, the PLA2-GIB cofactor is a protein or peptide comprising or consisting of the amino acid sequence of any one of SEQ ID NOs: 195-202, or a fragment or mimotope thereof.
In another specific embodiment, the PLA2-GIB cofactor is SARS-Cov2 ORF7b protein, or a fragment or mimotope thereof. In a particular embodiment, the PLA2-GIB cofactor is a protein or peptide comprising or consisting of the amino acid sequence of SEQ ID NO: 203, or a fragment or mimotope thereof.
In another specific embodiment, the PLA2-GIB cofactor is SARS-Cov2 ORF8 protein, or a fragment or mimotope thereof. In a particular embodiment, the PLA2-GIB cofactor is a protein or peptide comprising or consisting of the amino acid sequence of SEQ ID NO: 204, or a fragment or mimotope thereof.
In another specific embodiment, the PLA2-GIB cofactor is SARS-Cov2 protein N, or a fragment or mimotope thereof. In a particular embodiment, the PLA2-GIB cofactor is a protein or peptide comprising or consisting of the amino acid sequence of any one of SEQ ID NOs: 205-208, or a fragment or mimotope thereof.
Treatments that modulate the cofactor effect
The invention relates to the treatment of a Group IV RNA virus infection, particularly a coronavirus infection, more particularly SARS-Cov2 infection, in subjects by modulating such a cofactor effect. The invention thus provides methods and compositions for treating diseased subjects and/or for restoring/enhancing CD4 T cell activity in subjects using an inhibitor of a PLA2-GIB cofactor.
The term “inhibitor” of a cofactor designates any molecule or treatment which causes (directly or indirectly) an inhibition of the expression or a function of the cofactor, e.g., cofactor binding to gC1qR or cofactor ability to sensitize CD4 T cells to PLA2-GIB. Inhibiting the cofactor designates preferably reducing by at least 10%, 20%, 30%, 40%, 50%, 60%, 70%, 80% or more the expression or a function of the cofactor, as well as completely blocking or suppressing said expression or function. Depending on the situation, the inhibition may be transient, sustained or permanent.
In a particular embodiment, an inhibitor of the cofactor is a gC1qR inhibitor. Indeed, cofactors bind gC1qR as a target receptor. Blocking or reducing or preventing binding of the cofactor to gC1qR using gC1qR inhibitors can affect the cofactor effect. The term “gC1qR inhibitor” designates any molecule or treatment which causes (directly or indirectly) an inhibition of a function of gC1qR, e.g., gC1qR-mediated exocytosis. gC1qR designates the receptor for complement Clq at the surface of cells, particularly of CD4 T cells, especially the human form of said receptor. gC1qR is also known as Clq binding protein (C1QBP), ASF/SF2-associated protein p32 (SF2P32); Glycoprotein gClqBP; Hyaluronan-binding protein 1 (HABP1); Mitochondrial matrix protein p32; gClq-R protein; p33; ClqBP and GC1QBP. The amino acid sequence of the receptor was disclosed in the art. An exemplary amino acid sequence of human gC1qR is reproduced below (SEQ ID NO: 211):
MLPLLRC VPR VLGS S V AGLR A A AP AS PFRQLLQP APRLCTRPFGLLS VR AGS ER RPGLLRPRGPCACGCGCGSLHTDGDKAFVDFLSDEIKEERKIQKHKTLPKMSGG WELELN GTE AKLVRKVAGEKIT VTFNINN S IPPTFDGEEEPS QGQKVEEQEPELT S TPNFV VE VIKNDDGKKAL VLDCH YPEDE V GQEDE AES DIF S IRE V S FQS T GES E WKDTN YTLNTDS LD W ALYDHLMDFL ADRG VDNTFADELVELS T ALEHQE YIT FLEDLKS FVKS Q
The term gC1qR designates any receptor of SEQ ID NO: 211 (accession number UniProtKB/Swiss-Prot: Q07021.1) above, as well as processed forms and variants thereof. Variants include naturally-occurring variants having e.g., at least 90% amino acid sequence identity to SEQ ID NO: 211. Upon binding of a cofactor, gC1qR triggers a signaling pathway that results in exocytosis of intracellular vesicles. Without being bound by theory, it is believed that the fusion of these vesicles with the cytoplasmic membrane could change the lipid composition and increase sPLA2-GIB activity on CD4 T cells membrane, resulting in an inhibition of phosphoSTAT5 signaling. In particular, the fusion of these vesicles with plasma membrane can change the lipid composition and cause sPLA2-GIB activity on CD4 T cells membranes. As a result, membrane fluidity is increased and
cytokines receptors are aggregated in abnormal membrane domain, resulting in a dramatic decrease of cytokine signaling, and anergy of CD4 T cells.
The term gC1qR inhibitor thus includes any molecule which binds to gC1qR, or to a partner of gC1qR, and inhibits a function of gC1qR, such as gC1qR- mediated exocytosis.
In another embodiment, the cofactor inhibitor is a molecule which directly inhibits an activity of the cofactor, e.g., which binds the cofactor and/or inhibits binding of the cofactor to its receptor.
Preferred examples of cofactor inhibitors include, for instance, antibodies and variants thereof, synthetic specific ligands, peptides, small drugs, or inhibitory nucleic acids.
Antibodies
In a first embodiment, a cofactor inhibitor is an antibody or an antibody variant/fragment having essentially the same antigen specificity, or a nucleic acid encoding such an antibody or variant/fragment. The antibody may bind a cofactor, or gC1qR, or a partner of gC1qR, or a gC1qR-binding element thereof, and preferably inhibits a function of the cognate antigen (e.g., gC1qR or the cofactor).
Antibodies can be synthetic, monoclonal, or polyclonal and can be made by techniques well known per se in the art.
The term “antibodies” is meant to include polyclonal antibodies, monoclonal antibodies, fragments thereof, such as F(ab')2 and Fab fragments, single-chain variable fragments (scFvs), single-domain antibody fragments (VHHs or Nanobodies), bivalent antibody fragments (diabodies), as well as any recombinantly and synthetically produced binding partners, human antibodies or humanized antibodies.
Antibodies are defined to be specifically binding, preferably if they bind to the cognate antigen with a Ka of greater than or equal to about 107 M-l. Affinities of antibodies can be readily determined using conventional techniques, for example those described by Scatchard et al., Ann. N.Y. Acad. Sci., 51:660 (1949).
Polyclonal antibodies can be readily generated from a variety of sources, for example, horses, cows, donkeys, goats, sheeps, dogs, chickens, rabbits, mice, hamsters, or rats, using procedures that are well known in the art. In general, a purified immunogen, optionally appropriately conjugated, is administered to the host animal typically through parenteral injection. The immunogenicity of immunogen can be enhanced through the use of an adjuvant, for example, Freund's complete or incomplete adjuvant. Following booster immunizations, small samples of serum are collected and tested for reactivity to
the antigen polypeptide. Examples of various assays useful for such determination include those described in Antibodies: A Laboratory Manual, Harlow and Lane (eds.), Cold Spring Harbor Laboratory Press, 1988; as well as procedures, such as countercurrent immuno-electrophoresis (CIEP), radioimmunoassay, radio-immunoprecipitation, enzyme-linked immunosorbent assays (ELISA), dot blot assays, and sandwich assays. See U.S. Pat. Nos. 4,376,110 and 4,486,530.
Monoclonal antibodies can be readily prepared using well known procedures. See, for example, the procedures described in U.S. Pat. Nos. RE 32,011, 4,902,614, 4,543,439, and 4,411,993; Monoclonal Antibodies, Hybridomas: A New Dimension in Biological Analyses, Plenum Press, Kennett, McKeam, and Bechtol (eds.), 1980. Lor example, the host animals, such as mice, can be injected intraperitoneally at least once and preferably at least twice at about 3 week intervals with isolated and purified immunogen, optionally in the presence of adjuvant. Mouse sera are then assayed by conventional dot blot technique or antibody capture (ABC) to determine which animal is best to fuse. Approximately two to three weeks later, the mice are given an intravenous boost of protein or peptide. Mice are later sacrificed and spleen cells fused with commercially available myeloma cells, such as Ag8.653 (ATCC), following established protocols. Briefly, the myeloma cells are washed several times in media and fused to mouse spleen cells at a ratio of about three spleen cells to one myeloma cell. The fusing agent can be any suitable agent used in the art, for example, polyethylene glycol (PEG). Fusion is plated out into plates containing media that allows for the selective growth of the fused cells. The fused cells can then be allowed to grow for approximately eight days. Supernatants from resultant hybridomas are collected and added to a plate that is first coated with goat anti-mouse Ig. Following washes, a label is added to each well followed by incubation. Positive wells can be subsequently detected. Positive clones can be grown in bulk culture and supernatants are subsequently purified over a Protein A column (Pharmacia). Monoclonal antibodies may also be produced using alternative techniques, such as those described by Alting-Mees et al. , "Monoclonal Antibody Expression Libraries: A Rapid Alternative to Hybridomas", Strategies in Molecular Biology 3:1-9 (1990), which is incorporated herein by reference. Similarly, binding partners can be constructed using recombinant DNA techniques to incorporate the variable regions of a gene that encodes a specific binding antibody. Such a technique is described in Larrick et ah, Biotechnology, 7:394 (1989).
Antigen-binding fragments of antibodies, which can be produced by conventional techniques, are also encompassed by the present invention. Examples of such fragments include, but are not limited to, Fab and F(ab')2 fragments. Antibody fragments and derivatives produced by genetic engineering techniques are also provided.
The monoclonal antibodies of the invention also include chimeric antibodies, e.g., humanized versions of murine monoclonal antibodies. Such humanized antibodies can be prepared by known techniques, and offer the advantage of reduced immunogenicity when
the antibodies are administered to humans. In one embodiment, a humanized monoclonal antibody comprises the variable region of a murine antibody (or just the antigen binding site thereof) and a constant region derived from a human antibody. Alternatively, a humanized antibody fragment can comprise the antigen binding site of a murine monoclonal antibody and a variable region fragment (lacking the antigen-binding site) derived from a human antibody. Procedures for the production of chimeric and further engineered monoclonal antibodies include those described in Riechmann et al. (Nature 332:323, 1988), Liu et al. (PNAS 84:3439, 1987), Larrick et al. (Bio/Technology 7:934, 1989), and Winter and Harris (TIPS 14:139, May, 1993). Procedures to generate antibodies transgenically can be found in GB 2,272,440, U.S. Pat. Nos. 5,569,825 and 5,545,806. Antibodies produced by genetic engineering methods, such as chimeric and humanized monoclonal antibodies, comprising both human and non-human portions, which can be made using standard recombinant DNA techniques, can be used. Such chimeric and humanized monoclonal antibodies can be produced by genetic engineering using standard DNA techniques known in the art, for example using methods described in Robinson et al. International Publication No. WO 87/02671; Akira, et al. European Patent Application 0184187; Taniguchi, M., European Patent Application 0171496; Morrison et al. European Patent Application 0173494; Neuberger et al. PCT International Publication No. WO 86/01533; Cabilly et al. U.S. Pat. No. 4,816,567; Cabilly et al. European Patent Application 0125023; Better et al., Science 240: 1041 1043, 1988; Liu et al., PNAS 84:3439 3443, 1987; Liu et al., J. Immunol. 139:3521 3526, 1987; Sun et al. PNAS 84:214 218, 1987; Nishimura et al., Cane. Res. 47:999 1005, 1987; Wood et al., Nature 314:446 449, 1985; and Shaw et al., J. Natl. Cancer Inst. 80:1553 1559, 1988); Morrison, S. L., Science 229:1202 1207, 1985; Oi et al., BioTechniques 4:214, 1986; Winter U.S. Pat. No. 5,225,539; Jones et al., Nature 321:552525, 1986; Verhoeyan et al., Science 239:1534, 1988; and Beidler et al., J. Immunol. 141:4053 4060, 1988.
In connection with synthetic and semi-synthetic antibodies, such terms are intended to cover but are not limited to antibody fragments, isotype switched antibodies, humanized antibodies (e.g., mouse-human, human-mouse), hybrids, antibodies having plural specificities, and fully synthetic antibody-like molecules.
Human monoclonal antibodies can also be prepared by constructing a combinatorial immunoglobulin library, such as a Fab phage display library or a scFv phage display library, using immunoglobulin light chain and heavy chain cDNAs prepared from mRNA derived from lymphocytes of a subject. See, e.g., McCafferty et al. PCT publication WO 92/01047; Marks et al. (1991) J. Mol. Biol. 222:581 597; and Griffths et al. (1993) EMBO J 12:725 734. In addition, a combinatorial library of antibody variable regions can be generated by mutating a known human antibody. For example, a variable region of a human antibody known to bind gC1qR can be mutated by, for example, using randomly altered mutagenized oligonucleotides, to generate a library of mutated variable regions which can then be screened to bind to gC1qR. Methods of inducing random mutagenesis
within the CDR regions of immunoglobin heavy and/or light chains, methods of crossing randomized heavy and light chains to form pairings and screening methods can be found in, for example, Barbas et al. PCT publication WO 96/07754; Barbas et al. (1992) Proc. Nat'l Acad. Sci. USA 89:44574461. Antibodies of the invention may be directed against gC1qR, a gC1qR ligand, or a gC1qR partner, and cause an inhibition of signaling mediated by gC1qR. For preparing antibodies of the invention, an immunogen may be used comprising gC1qR, a gC1qR ligand, or a gC1qR partner, or a fragment, variant, or fusion molecule thereof. Antibodies to gClqR
Particular antibodies of the invention bind a gC1qR epitope, and/or have been generated by immunization with a polypeptide comprising a gC1qR epitope, selected from the mature gC1qR protein or a fragment of gC1qR comprising at least 8 consecutive amino acid residues thereof. Preferred anti-gC1qR antibodies of the invention bind an epitope of a ligand-binding site within gC1qR, thereby interfering with binding of the ligand. In a particular embodiment, the antibodies bind an epitope comprised between amino acid residues 76-282 of SEQ ID NO: 211, which contain the gC1qR ligand bind site. Clq binding to gC1qR can involve at least three different motifs on gC1qR, namely: amino acid residues 75-96, 190-202 and 144-162 (by reference to SEQ ID NO: 211). HCV core protein binding to gC1qR can involve at least two different motifs on gC1qR, namely: amino acid residues 144-148 and 196-202 (by reference to SEQ ID NO: 211). HIV gp41 binding to gC1qR can involve at least amino acid residues 174- 180 on gC1qR (by reference to SEQ ID NO: 211).
It is thus preferred to use an antibody (or variant thereof) which binds an epitope containing at least one amino acid residue contained in one of said epitopes or close to one of said epitopes. Examples of such antibodies include antibody 60.11, which binds to residues 75-96 of gC1qR; as well as antibody 74.5.2, which binds to an epitope with the residues 204 to 218.
Preferred gC1qR inhibitors are therefore monoclonal antibodies against gC1qR, more preferably against an epitope of gC1qR located within amino acid residues 76-282 of the protein (by reference to SEQ ID NO: 211), even more preferably an epitope containing an amino acid residue selected from amino acids 75-96, 144-162, 174-180, and 190- 210. Preferred antibodies are neutralizing (or antagonist) antibodies, i.e., they prevent or inhibit or reduce binding of a natural ligand to the receptor and/or signaling through the receptor.
Antibodies to a PLA2-GIB cofactor
Other particular inhibitors of the invention are antibodies that bind a PLA2-GIB cofactor and/or have been generated by immunization with a PLA2-GIB cofactor or a fragment or variant thereof, and preferably inhibit at least partially an activity of such cofactor, preferably the binding of such a cofactor to gC1qR.
Particular antibodies of the invention are polyclonal antibodies or monoclonal antibodies, or variants thereof, which bind a protein selected from the proteins listed in Table 3, and inhibit at least partially the binding of said protein to gC1qR. Preferred antibodies of the invention are polyclonal antibodies or monoclonal antibodies, or variants thereof, which bind a protein selected from the proteins listed in Table 3, and inhibit at least partially the binding of said protein to gC1qR.
In a particular embodiment, the inhibitor is an antibody or a variant thereof that binds SARS-Cov-2 spike protein. In a further particular embodiment, the antibody binds an epitope comprising an amino acid residue contained in the NTD domain or in the RBD domain of the spike protein (see example 3 for domain position). Most preferred antibodies bind an epitope comprising an amino acid residue located in the RBD (amino acid residues 331-527 of SEQ ID NO: 209), or even in the RBM (amino acid residues 436-509 of SEQ ID NO: 209). Particular antibodies bind an epitope comprising any amino acid residue of the spike located in anyone of SEQ ID Nos: 156-185.
Particularly preferred antibodies bind an epitope containing at least one amino acid residues of Spike protein located at any of the following positions (by reference to SEQ ID NO: 209): 99-113; 147-161; 253-267; 348-445, 348-362, 431-445. Particular antibodies bind an epitope comprising at least one of the following amino acid residues of spike protein (by reference to SEQ ID NO: 209): 431, 432, 433, 434, 435, 436, 437, 438, 438, 440, 441, 442, 443, 444, 445.
Particular antibodies bind an epitope comprising at least one of the following amino acid residues of spike protein (by reference to SEQ ID NO: 209): 348, 349, 350, 351, 352, 353, 354, 355, 356, 357, 358, 359, 360, 361, 362.
As shown in the experimental section, such residues are highly relevant since they also contain B-cell epitopes and/or T-cell epitopes.
In another particular embodiment, the inhibitor is an antibody or a variant thereof that binds SARS-Cov-2 ORFlab protein. In a further particular embodiment, the antibody
binds an epitope comprising an amino acid residue contained in anyone of SEQ ID Nos: 2-155.
In another particular embodiment, the inhibitor is an antibody or a variant thereof that binds SARS-Cov-2 ORF3 protein. In a further particular embodiment, the antibody binds an epitope comprising an amino acid residue contained in anyone of SEQ ID Nos: 186- 193.
In another particular embodiment, the inhibitor is an antibody or a variant thereof that binds SARS-Cov-2 protein E. In a further particular embodiment, the antibody binds an epitope comprising an amino acid residue contained in SEQ ID No: 194. In another particular embodiment, the inhibitor is an antibody or a variant thereof that binds SARS-Cov-2 protein M. In a further particular embodiment, the antibody binds an epitope comprising an amino acid residue contained in anyone of SEQ ID Nos: 195-202.
In another particular embodiment, the inhibitor is an antibody or a variant thereof that binds SARS-Cov-2 protein ORF7b. In a further particular embodiment, the antibody binds an epitope comprising an amino acid residue contained in SEQ ID No: 203.
In another particular embodiment, the inhibitor is an antibody or a variant thereof that binds SARS-Cov-2 protein ORF8. In a further particular embodiment, the antibody binds an epitope comprising an amino acid residue contained in SEQ ID No: 204.
In another particular embodiment, the inhibitor is an antibody or a variant thereof that binds SARS-Cov-2 protein N. In a further particular embodiment, the antibody binds an epitope comprising an amino acid residue contained in anyone of SEQ ID Nos: 205-208.
Any such antibodies and fragments and derivatives thereof represent particular objects of the invention. Inhibitory Nucleic acids
In an alternative embodiment, the cofactor inhibitor is an inhibitory nucleic acid. Preferred inhibitory nucleic acids include aptamers which are designed to bind the cofactor, or gC1qR, or a partner of gC1qR, and to inhibit a function thereof.
Peptides
In an alternative embodiment, the cofactor inhibitor is a peptide that inhibits a function of the cofactor. The peptide is typically a molecule that selectively binds a cofactor, a gC1qR, or a partner of gC1qR.
Peptides preferably contain from 4 to 40 amino acid residues, and their sequence may be identical to a domain of gC1qR or to a domain of a cofactor (bait peptide), or their sequence may contain a variation as compared to the sequence of a domain of gC1qR or to a domain of a cofactor (peptide antagonist).
Preferred peptides of the invention contain from 4 to 40 consecutive amino acid residues of SEQ ID NO: 211 (gC1qR) or of protein selected from ORFlab, Spike, ORF3, E, M, ORF7b, ORF8 or N. Examples of such peptides are or comprise the amino acid sequence of anyone of SEQ ID Nos: 2-208. Particularly preferred peptides of the invention contain from 4 to 40 consecutive amino acid residues of the spike protein of SEQ ID NO: 209, such as peptides consisting of or comprising anyone of SEQ ID NOs: 156-185.
Peptides of the invention may be modified to increase their properties. The modification may consist of one or more amino acid substitution. Examples of such substitution includes, without limitation, replacement of a charged or reactive amino acid residue by a more neutral residue such as alanine, or conversely. The modification may alternatively (or in addition) consist of a chemical modification, such as addition of a chemical group to one (or both) ends of the peptide, or to a lateral chain thereof, or to a peptide bond. In this regard, the peptides of the invention can comprise peptide, non-peptide and/or modified peptide bonds. In a particular embodiment, the peptides comprise at least one peptidomimetic bond selected from intercalation of a methylene (-CH2-) or phosphate (- PO2-) group, secondary amine (-NH-) or oxygen (-0-), alpha-azapeptides, alpha- alky lpeptides, N-alkylpeptides, phosphonamidates, depsipeptides, hydroxymethylenes, hydroxyethylenes, dihydroxyethylenes, hydroxyethylamines, retro-inverso peptides, methyleneoxy, cetomethylene, esters, phosphinates, phosphinics, or phosphonamides. Also, the peptides may comprise a protected N-ter and/or C-ter function, for example, by acylation, and/or amidation and/or esterification.
Examples of such peptides include, for instance the peptide with amino acid residues 144- 162 of SEQ ID NO: 211 (gC1qR) and the peptide with amino acid residues 204-218 of SEQ ID NO: 211 (gC1qR).
Further specific examples of such peptides of the invention include peptides comprising a sequence of anyone of SEQ ID NOs: 156-185 with one amino acid substitution, more preferably with at least one amino acid selected from W, I or K replaced with an Alanine.
The peptides of the invention may be produced by techniques known per se in the art such as chemical, biological, and/or genetic synthesis.
Each of these peptides, in isolated form, represents a particular object of the present invention. The term "isolated", as used herein, refers to molecules (e.g., nucleic or amino acid) that are removed from a component of their natural environment, isolated or separated, and are at least 60% free, preferably 75% free, and most preferably 90% free
from other components with which they are naturally associated. An "isolated" polypeptide (or protein) is for instance a polypeptide separated from a component of its natural environment and, preferably purified to greater than 90% or 95% purity as determined by, for example, electrophoretic (e.g., SDS-PAGE, isoelectric focusing (IEF), capillary electrophoresis) or chromatographic (e.g., ion exchange or reverse phase HPLC) migration. An "isolated" nucleic acid refers to a nucleic acid molecule separated from a component of its natural environment and/or assembled in a different construct (e.g., a vector, expression cassette, recombinant host, etc.).
Small drugs
Other inhibitors are small drug inhibitors, such as are hydrocarbon compounds that selectively bind gC1qR or a cofactor.
Small drugs are preferably obtainable by a method comprising: (i) contacting a test compound with a cell expressing gC1qR, (ii) selecting a test compound which binds gC1qR, and (iii) selecting a compound of (ii) which inhibits an activity of gC1qR. Such a method represents a particular object of the invention. gClqR soluble receptors
In an alternative embodiment, the cofactor inhibitor is a soluble form of gC1qR. Pseudotyped viruses
Another type of inhibitor is a virus expressing a peptide as defined above. Preferred viruses express the peptide at the surface of the virus to ensure exposure. In a particular embodiment, the peptide is inserted in the envelope protein of the virus. Viruses that may be used for that purpose include any safe virus for use in humans, such as adenoviruses, AAVs, retroviruses (including lentiviruses), etc. gClqR soluble receptors
In an alternative embodiment, the cofactor inhibitor is a soluble form of gC1qR.
Immunogens and vaccination
In an alternative (or cumulative) embodiment, inhibition of the cofactor in a subject is obtained by using (e.g., vaccinating or immunizing the subject with) an immunogen or an antigen that can induce an immune response in vivo. As a result, the subject produces antibodies (or immune cells) which inhibit the cofactor (such as its interaction with gC1qR). In particular, administration(s) of a cofactor immunogen (e.g., any immunogenic portion of a cofactor) can generate antibodies in the treated subject. These antibodies will inhibit the cofactor effect as immunotherapy or a vaccine prophylaxis.
An object of the invention thus resides in a method of treating coronavirus infection by vaccinating a subject with an antigen that induces an immune response against a PLA2- GIB cofactor, particularly against SARS-Cov-2 spike antigen.
A further object of the invention relates to an antigen that induces an immune response against of a PLA2-GIB cofactor, particularly against SARS-Cov-2 spike antigen, for use to vaccinate a subject in need thereof.
In a particular embodiment, the immunogen of a PLA2-GIB cofactor antigen used for vaccination is an inactivated immunogenic molecule that induces an immune response against the cofactor in a subject. Inactivation may be obtained e.g., by chemically or physically altering the cofactor or by mutating or truncating the protein, or both; and immunogenicity may be obtained as a result of the inactivation and/or by further conjugating the protein to a suitable carrier or hapten, such as KLH, HSA, polylysine, a viral anatoxin, or the like, and/or by polymerization, or the like. The immunogen may thus be chemically or physically modified, e.g., to improve its immunogenicity.
In a particular embodiment, the immunogen of a PLA2-GIB cofactor of the invention comprises the entire cofactor or a variant thereof.
In this respect, in a particular embodiment, the immunogen comprises a virus envelope protein with a modified gC1qR binding motif. Such immunogen thus retains immunogenicity while having no or less PLA2-GIB cofactor effect.
A further object of the invention thus relates to an immunogen, wherein the immunogen comprises a viral envelope protein with a modified gC1qR binding motif, preferably a deleted and/or mutated gC1qR binding motif. A specific example is a SARS-Cov-2 spike protein having, by reference to SEQ ID NO: 209, at least one amino acid modification in a residue selected from anyone of SEQ ID NOs: 156-185, even more preferably in a residue selected from aa 348-362 or 431-445 (by reference to SEQ ID NO: 209).
A specific embodiment of the invention is a spike protein comprising the amino acid sequence of SEQ ID NO: 209, or a natural variant thereof, wherein the sequence contains at least one amino acid substitution or deletion in a position selected from: aa99-113, aal47-161, aa253-267, aa348-362, aa431-445. In a more preferred embodiment, the sequence contains at least one deletion of anyone said residues, even more preferably of at least 2, 3, 4, 5, 6, 7, 8, 9, 10, or more consecutive aa residues at said positions.
A particular protein of the invention is a protein comprising SEQ ID NO: 209 with a deletion, preferably of at most 50 amino acids, spanning all or part of AA431-445.
Another particular protein of the invention is a protein comprising SEQ ID NO: 209 with a deletion, preferably of at most 50 amino acids, spanning all or part of AA348-362.
Such antigens retain immunogenicity of the viral protein, such as an ability to induce immune response against the vims when administered in vivo. Such antigens have a reduced cofactor effect, such as a cofactor effect reduced by at least 20% as compared to the native protein or virus.
In an alternative embodiment, the immunogen of a PLA2-GIB cofactor comprises a fragment of a cofactor comprising at least 6 consecutive amino acid residues and containing an immunogenic epitope thereof. In a preferred embodiment, the immunogen comprises at least from 6 to 20 amino acid residues. Preferred immunogens of the invention comprise or consist of from 4 to 30 consecutive amino acid residues of a protein or peptide selected from anyone of SEQ ID NOs: 2-211.
The immunogen may be in various forms such as in free form, polymerized, chemically or physically modified, and/or coupled (i.e., linked) to a carrier molecule. Coupling to a carrier may increase the immunogenicity and (further) suppress the biological activity of the immunogen. In this regard, the carrier molecule may be any carrier molecule or protein conventionally used in immunology such as for instance KLH (Keyhole limpet hemocyanin), ovalbumin, bovine serum albumin (BSA), a viral or bacterial anatoxin such as toxoid tetanos, toxoid diphteric B cholera toxin, mutants thereof such as diphtheria toxin CRM 197, an outer membrane vesicle protein, a polylysine molecule, or a vims like particle (VLP). In a preferred embodiment, the carrier is KLH or CRM 197 or a VLP.
Coupling of the immunogen to a carrier may be performed by covalent chemistry using linking chemical groups or reactions, such as for instance glutaraldehyde, biotin, etc. Preferably, the conjugate or the immunogen is submitted to treatment with formaldehyde in order to complete inactivation of the cofactor.
The immunogenicity of the immunogen may be tested by various methods, such as by immunization of a non-human animal grafted with human immune cells, followed by verification of the presence of antibodies, or by sandwich ELISA using human or humanized antibodies. The lack of biological activity may be verified by any of the activity tests described in the application.
In a particular embodiment, the invention relates to an inactivated and immunogenic PLA2-GIB cofactor.
In a further particular embodiment, the invention relates to a PLA2-GIB cofactor protein or a fragment or variant thereof conjugated to a carrier molecule, preferably to KLH.
In a further aspect, the invention relates to a vaccine comprising an immunogen of PLA2- GIB cofactor, a suitable excipient and, optionally, a suitable adjuvant.
A further object of the invention relates to a method for inducing the production of antibodies that neutralize the activity of a PLA2-GIB cofactor in a subject in need thereof, the method comprising administering to said subject an effective amount of an immunogen or vaccine as defined above.
Administration of an immunogen or vaccine of the invention may be by any suitable route, such as by injection, preferably intramuscular, subcutaneous, transdermal, intravenous or intraarterial; by nasal, oral, mucosal or rectal administration.
Other active agents The inhibitor of the SPLA2G1B pathway may be used alone, or in combination with one or more other active agents, such as a cytostatic or cytotoxic agent.
The agent may be an antibiotic, such as any broad-spectrum antibiotic, or an antibiotic with specific spectrum towards the target bacterium. Examples of antibiotics include, but are not limited to, amoxicillin, clarithromycin, cefuroxime, cephalexin ciprofloxacin, clindamycin, doxycycline, metronidazole, terbinafine, levofloxacin, nitrofurantoin, tetracycline, penicillin and azithromycin.
The agent may be an anti-fungal agent, such as, without limitation, clotrimazole, butenafine, butoconazole, ciclopirox, clioquinol, clioquinol, clotrimazole, econazole, fluconazole, flucytosine, griseofulvin, haloprogin, itraconazole, ketoconazole, miconazole, naftifine, nystatin, oxiconazole, sulconazole, terbinafine, terconazole, tioconazole, and tolnaftate. The agent may be an anti-viral agent, such as, without limitation, zidovudine, didanosine, zalcitabine, stavudine, lamivudine, abacavir, tenofovir, nevirapine, delavirdine, efavirenz, saquinavir, ritonavir, indinavir, nelfinavir, saquinavir, amprenavir, and lopinavir. The agent may be selected from other drugs such as chloroquine, hydroxychloroquine, Remdesivir, or ABX464, for instance.
Compositions & methods of treatment
The invention also relates to a composition comprising an inhibitor as defined above and, preferably, a pharmaceutically acceptable diluent, excipient or carrier.
A “pharmaceutical composition” refers to a formulation of a compound of the invention (active ingredient) and a medium generally accepted in the art for the delivery of biologically active compounds to the subject in need thereof. Such a carrier includes all pharmaceutically acceptable carriers, diluents, medium or supports therefore. Conventional pharmaceutical practice may be employed to provide suitable formulations or compositions to subjects, for example in unit dosage form.
The compounds or compositions according to the invention may be formulated in the form of ointment, gel, paste, liquid solutions, suspensions, tablets, gelatin capsules, capsules, suppository, powders, nasal drops, or aerosol, preferably in the form of an injectable solution or suspension. For injections, the compounds are generally packaged in the form of liquid suspensions, which may be injected via syringes or perfusions, for example. In this respect, the compounds are generally dissolved in saline, physiological, isotonic or buffered solutions, compatible with pharmaceutical use and known to the person skilled in the art. Thus, the compositions may contain one or more agents or excipients selected from dispersants, solubilizers, stabilizers, preservatives, etc. Agents or excipients that can be used in liquid and/or injectable formulations are notably methylcellulose, hydroxymethylcellulose, carboxymethylcellulose, polysorbate 80, mannitol, gelatin, lactose, vegetable oils, acacia, etc. The carrier can also be selected for example from methyl-beta-cyclodextrin, a polymer of acrylic acid (such as carbopol), a mixture of polyethylene glycol and polypropylene glycol, monoethanolamine and hydroxymethyl cellulose.
The compositions generally comprise an effective amount of a compound of the invention, e.g., an amount that is effective to modulate directly or indirectly an effect of PLA2-GIB on CD4 T cells. Inhibitors are typically used in an amount effective to maintain/restore resistance of CD4 T cells to inactivation by PLA2-GIB. Generally, the compositions according to the invention comprise from about 1 μg to 1000 mg of a cofactor or modulator, such as from 0.001-0.01, 0.01-0.1, 0.05-100, 0.05-10, 0.05-5, 0.05- 1, 0.1-100, 0.1-1.0, 0.1-5, 1.0-10, 5-10, 10-20, 20-50, and 50-100 mg, for example between 0.05 and 100 mg, preferably between 0.05 and 5 mg, for example 0.05, 0.1, 0.2, 0.3, 0.4, 0.5, 1, 2, 3, 4 or 5 mg. The dosage may be adjusted by the skilled person depending on the agent and the disease.
The compositions of the invention can further comprise one or more additional active compounds, for separate, simultaneous or sequential use. Examples of additional active compounds include, but are not limited to, chemotherapeutic drug, antibiotics, antiparasitic agents, antifungal agents or antiviral agents.
In another particular embodiment, the inhibitor is used in combination with a modulator of PLA2-GIB.
In a particular embodiment, the methods or compositions of the invention use a combination of (i) an inhibitor of a PLA2GIB cofactor and (ii) an antibody against PLA2GIB (or a derivative or fragment thereof). In a further particular embodiment, the inhibitor of a PLA2GIB cofactor in an antibody against the cofactor, or an antibiotic, or an antifungal agent, or an antivirus agent.
In another particular embodiment, the method or compositions of the invention use a combination of (i) an inhibitor of a PLA2GIB cofactor and (ii) an indole-based inhibitor of PLA2GIB (such as 3-(2-amino-l,2-dioxoethyl)-2-ethyl-l-(phenylmethyl)-lH-indol-4- yl)oxy)acetic acid or a pharmaceutically acceptable salt, hydrate, or prodrug thereof, such as a sodium salt thereof (Varespladib)). In a further particular embodiment, the inhibitor of a PLA2GIB cofactor in an antibody against the cofactor, or an antibiotic, or an antifungal agent, or an antivirus agent.
In another particular embodiment, the method or compositions of the invention use a combination of (i) an inhibitor of a PLA2GIB cofactor and (ii) a pentapeptide inhibitor of PLA2GIB (such as a cyclic peptide selected from FLSYK, FLSYR and (2NapA)LS(2NapA)R). In a further particular embodiment, the inhibitor of a PLA2GIB cofactor in an antibody against the cofactor, or an antibiotic, or an antifungal agent, or an antivirus agent.
In another particular embodiment, the method or compositions of the invention use a combination of (i) an inhibitor of a PLA2GIB cofactor and (ii) a further active agent such as an antiviral agent, an antibiotic, chloroquine or hydroxychloroquine.
The invention also relates to a method for preparing a pharmaceutical composition, comprising mixing a cofactor or modulator as previously described and a pharmaceutically acceptable diluent or excipient, and formulating the composition in any suitable form or container (syringe, ampoule, flask, bottle, pouch, etc.).
The invention also relates to a kit comprising (i) a composition comprising a cofactor or modulator as previously described, (ii) at least one container, and optionally (iii) written instructions for using the kit.
The compounds and compositions and methods of the invention may be used to treat a variety of diseases caused by a Group IV RNA virus infection, particularly coronavirus infection, more particularly SARS-Cov-2 infection.
In a particular embodiment, the invention is directed to methods for stimulating an immune response in a subject infected with a Group IV RNA vims, particularly coronavirus.
In another particular embodiment, the invention is directed to methods for treating an immunodeficiency or an associated disorder in a subject infected with a Group IV RNA virus infection, particularly a coronavirus. For use in the present invention, the inhibitor may be administered by any suitable route. Preferably, administration is by injection, such as systemic or parenteral injection or perfusion, e.g., intramuscular, intravenous, intraarterial, subcutaneous, etc. Administration is typically repeated, or continuous. The duration, dosages and frequency of administering compounds or compositions of the invention may be adapted according to the subject and disease.
A typical regimen comprises a single or repeated administration of an effective amount of an inhibitor over a period of one or several days, up to one year, and including between one week and about six months. It is understood that the dosage of a pharmaceutical compound or composition of the invention administered in vivo will be dependent upon the age, health, sex, and weight of the recipient (subject), kind of concurrent treatment, if any, frequency of treatment, and the nature of the pharmaceutical effect desired. The ranges of effectives doses provided herein are not intended to be limiting and represent preferred dose ranges. However, the most preferred dosage will be tailored to the individual subject, as is understood and determinable by one skilled in the relevant arts (see, e.g., Berkowet et al., eds., The Merck Manual, 16th edition, Merck and Co., Rahway, N.J., 1992; Goodmanetna., eds., Goodman and Oilman’s The pharmacological Basis of Therapeutics, 10th edition, Pergamon Press, Inc., Elmsford, N.Y., (2001)).
The invention may be used in any mammal, particularly any human.
Diagnosis and detection of Coronavirus
The present invention also provides efficient methods for detecting SARS-Cov-2 infection in a sample. More particularly, the cofactors and antibodies of the inention can be used as reagent for detecting SARS-Cov-2 infection in a sample.
In a particular embodiment, the invention relates to the use of an antibody of the invention for detecting the presence of SARS-Cov-2 in a sample. In another particular embodiment, the invention relates to the use of a polypeptide or molecule of the invention for detecting the presence of anti-SARS-Cov-2 antibodies in a sample.
The detection is preferably performed in vivo, using conventional techniques such as immunoassays or radioassays. In a particular embodiment, detection is through ELISA
type assay. In particular, the sample is contacted with either the polypeptides or antibodies of the invention, and the formation of an immune complex is detected, such as with an antibody. The method is typically conducted in any suitable device, such as plate, microplate, tube, etc. The reagents are typically coated on a surface of the device. The sample may by any biological sample or derivatives thereof (dilutions, concentrate, enriched, aliquotes, etc), such as a biological fluid (e.g., blood, serum, plasma, urine, nasal swab, etc.).
In a particular embodiment, the invention relates to a method for detecting in vitro the presence of SARS-Cov-2 in a sample, comprising (i) contacting the sample in vitro with an antibody (or fragment or derivative thereof) of the invention, particularly an anti-Spike antibody (or fragment or derivative thereof) of the invention, and (ii) detecting the presence of antibody-antigen type complexes.
In another particular embodiment, the invention relates to a method for detecting in vitro the presence of anti-SARS-Cov-2 antibodies in a sample, comprising (i) contacting the sample in vitro with a molecule comprising a sequence selected from anyone of SEQ ID Nos: 1-208 or an epitope-containing fragment thereof, particularly a molecule comprising a sequence selected from anyone of SEQ ID Nos: 156-185 or an epitope-containing fragment thereof, and (ii) detecting the presence of antibody-antigen type complexes.
The invention also relates to a kit comprising (i) an antibody (or fragment or derivative thereof) of the invention, or (ii) a molecule comprising a sequence selected from anyone of SEQ ID Nos: 1-208 or an epitope-containing fragment thereof. The kit may further comprise a support and/or reagents and/or instructions for performing an immunoassay.
Further aspects and advantages of the invention will be disclosed in the following experimental section. Examples
Example 1: Definition of the Position Specific Scoring Matrix (PSSM) for the identification of gClqR peptide binding motifs Synthetic peptides designed by the inventors were assayed in vitro for their binding capacity to the receptor for the globular heads of Clq (gC1qR). The in vitro assay is based on an enzyme-linked immunosorbent assay (ELISA), as disclosed in Pednekar et al. Mol Immunol. 2016. The sequence of the tested peptides is presented in table 1 below.
Table 1:
Peptide sequences and binding values were used to define an optimized peptide-binding motif to gC1qR using GibbsCluster 2.0 (http://www.cbs.dtu.dk/services/GibbsCluster- 2.01. GibbsCluster 2.0 was run with default parameters except length which was set to 15 amino-acids. Cluster 1 obtained from the analysis was selected to generate a Position Specific Scoring Matrix (PSSM), which is disclosed in Figure 1. The matrix displays weight (or "preference") for any given amino acid at each position for a window of 15 amino acids in length.
Example 2: Identification of gClqR binding motifs (gC1qR:PSSM) within the SARS-CoV-2 proteome.
The Position Specific Scoring Matrix (PSSM) defined in Example 1 was used to search for the presence of gC1qR:PSSM motif within SARS-CoV-2.
2.1. Construction of SARS-CoV-2 proteome
A SARS-CoV-2 proteome sequence (SEQ ID NO:l) was constructed by co-linear assembly of open-reading frames from the SARS-CoV-2 reference genome (NC_045512.2). Positions of the different open-reading frames (ORFlab, Spike, ORF3a, E, M, ORF6, ORF7a, ORF7b, ORF8, N and ORF10) are presented in table 2.
Table 2: Positions of SARS-CoV-2 open-reading frames within SEQ ID NQ:1.
2.2. PSSM search For any selected windows of 15 amino acids, the PSSM returns a single score calculated by adding the relevant values at each position. The PSSM score is assigned to the N- terminal amino acid position of the window unless otherwise stated. A score >=0 is considered to be representative of a high scoring peptide. More preferably, a score >=5 is considered to be representative of a high scoring peptide.
Figure 2A provides the PSSM scores across the SARS-CoV-2 proteome (SEQ ID NO: 1). Figure 2B and Table 3 provide the 15 amino-acid sequences having a score >=5 across the SARS-CoV-2 proteome. Table 3: High scoring gClqR:PSSM motifs identified within the SARS-CoV-2 proteome (SEQ ID NO: 11.
ORFlab HSLAEYHNESGLKTI 5.43
ORFlab NKC A YW VPR AS ANIG 14.163
ORFlab EGSEGLNDNLLEILQ 7.905
ORFlab ETVKGLDYKAFKQIV 15.849
ORFlab KVTKGKAKKGAWNIG 9.312
ORFlab AKKGAWNIGEQKSIL 7.182
ORFlab AWNIGEQKSILSPLY 6.58
ORFlab TILDGISQYSLRLID 5.971
ORFlab DGISQYSLRLIDAMM 10.664
ORFlab MFTS DLATNNL V VM A 8.335
ORFlab QLTS QWLTNIFGT VY 5.081
ORFlab QWLTNIFGTVYEKLK 5.727
ORFlab KPVLDWLEEKFKEGV 6.886
ORFlab QTFFKLVNKFLALCA 5.853
ORFlab PTKVTFGDDTVIEVQ 6.061
ORFlab IDLDEW S M AT Y YLFD 7.143
ORFlab EPSTQYEY GTEDDY Q 6.22
ORFlab EQEEDWLDDDS QQT V 5.295
ORFlab GG V AG ALNKATNN AM 8.855
ORFlab ATNNAMQVESDDYIA 6.822
ORFlab VYLAVFDKNLYDKLV 6.117
ORFlab VFDKNLYDKLVSSFL 5.651
ORFlab SFLEMKSEKQVEQKI 5.741
ORFlab LGTVS WNLREMLAHA 5.09
ORFlab HFIETIS LAGS YKD W 7.684
ORFlab PNDDTLRVEAFEYYH 5.57
ORFlab NHTKKWKYPQVN GLT 10.282
ORFlab KWKYPQ VN GLTS IKW 11.398
ORFlab KWADNNCYLATALLT 5.738
ORFlab ALIL A Y CNKT V GELG 12.075
ORFlab HANLDSCKRVLNVVC 6.533
ORFlab MYMGTLSYEQFKKGV 6.092
ORFlab HYKHITSKETLYCID 7.022
ORFlab KKDNSYFTEQPIDLV 5.633
ORFlab VPN QP YPN AS FDNFK 7.214
ORFlab YPN AS FDNFKF V CDN 7.033
ORFlab NFKFVCDNIKFADDL 6.828
ORFlab YKPNTWCIRCLWSTK 8.376
ORFlab CIRCLWSTKPVETSN 5.554
ORFlab EDAQGMDNLACEDLK 7.522
ORFlab PNFS KLINIIIWFLL 6.225
ORFlab SYCTGYREGYLNSTN 6.198
ORFlab GYREGYLNSTNVTIA 7.816
ORFlab VTIAT Y CTGS IPC S V 5.773
ORFlab SFKWDLTAFGLVAEW 6.607
ORFlab SYFAVHFISNSWLMW 6.856
ORFlab NS WLMWLIINLV QM A 7.802
ORFlab NGGKGFCKLHNWNCV 6.125
ORFlab AGQKT YERHS LS HF V 7.313
ORFlab AEAELAKNVSLDNVL 5.472
ORFlab AKNVSLDNVLSTFIS 5.55
ORFlab INAQVAKSHNIALIW 5.054
ORFlab NIALIWNVKDFMSLS 6.449
ORFlab NWLKQLIKVTLVFLF 10.659
ORFlab STDTCFANKHADFDT 7.645
ORFlab DTCFANKHADFDTWF 7.231
ORFlab DIS AS I V AGGI V AIV 9.312
ORFlab PLVPFWITIAYIICI 8.032
ORFlab HFYWFFSNYLKRRVV 5.101
ORFlab EAACCHLAKALNDFS 5.697
ORFlab KALNDFSNSGSDVLY 5.374
ORFlab MV QVTCGTTTLN GLW 5.323
ORFlab CGTTTLN GLWLDD VV 5.044
ORFlab NFLV Q AGN V QLRVIG 9.121
ORFlab TIKGSFLNGSCGSVG 8.217
ORFlab VNVLAWLYAAVINGD 5.674
ORFlab RWFLNRFTTTLNDFN 10.363
ORFlab RFTTTLNDFNLVAMK 5.067
ORFlab MRIMT WLDM VDT S LS 6.22
ORFlab M VDT S LS GFKLKDC V 7.366
ORFlab KDC VM Y AS A V VLLIL 5.944
ORFlab VYKVYYGNALDQAIS 6.443
ORFlab YGNALDQAISMWALI 6.237
ORFlab CPIFFITGNTLQCIM 8.46
ORFlab S ID AFKLNIKLLG V G 5.029
ORFlab TMLRKLDNDALNNII 9.314
ORFlab NTCDGTTFTYASALW 8.112
ORFlab DGTTFTYASALWEIQ 7.75
ORFlab Y AS ALWEIQQ V VD AD 6.352
ORFlab NLNRGM VLGS LA AT V 7.131
ORFlab KTNCCRFQEKDEDDN 6.123
ORFlab HFDEGN CDTLKEIL V 8.951
ORFlab CCDDDYFNKKDWYDF 8.935
ORFlab DWYDFVENPDILRVY 10.758
ORFlab QFCDAMRNAGIVGVL 10.771
ORFlab V G VLTLDN QDLN GNW 8.537
ORFlab S FKELL V Y A ADP AMH 5.471
ORFlab TVKPGNFNKDFYDFA 7.19
ORFlab NFNKDFYDFA V S KGF 7.665
ORFlab FFFAQDGNAAISDYD 5.174
ORFlab CYDGGCINANQVIVN 5.264
ORFlab PFNKW GKARLYYDS M 5.71
ORFlab MTNRQFHQKLLKS IA 10.93
ORFlab KFY GGWHNMLKTVY S 13.987
ORFlab PHLMGWDYPKCDRAM 15.621
ORFlab GWDYPKCDRAMPNML 6.593
ORFlab CFNS T Y AS QGL V AS I 6.909
ORFlab SEAKCWTETDLTKGP 6.765
ORFlab MY S VMLTNDNT S R YW 9.425
ORFlab NT S R YWEPEFYE AM Y 8.771
ORFlab DH VIS T S HKLVLS VN 5.671
ORFlab SDNVTDFNAIATCDW 8.163
ORFlab DNVTDFNAIATCDWT 5.252
ORFlab IAT CD WTN AGD YIL A 7.185
ORFlab TVREVLSDRELHLSW 8.024
ORFlab CFDKFKVNSTLEQYV 13.069
ORFlab NYDLSVVNARLRAKH 7.764
ORFlab V VN ARLRAKH Y V YIG 6.931
ORFlab KDKSAQCFKMFYKGV 5.536
ORFlab SCNVNRFNVAITRAK 5.556
ORFlab TGLFKDCSKVITGLH 5.846
ORFlab RHVRAWIGFDVEGCH 8.846
ORFlab DTLKNLS DR V VF VLW 12.76
ORFlab LWAHGFELTSMKYFV 5.023
ORFlab HGFELT S MKYFVKIG 5.863
ORFlab AT CFSTASDTY AC WH 8.789
ORFlab DTYACWHHSIGFDYV 6.188
ORFlab NPKAIKCVPQADVEW 6.57
ORFlab TRLQS LEN V AFN V VN 7.473
ORFlab PEVKILNNLGVDIAA 5.734
ORFlab LNNLGVDIAANTVIW 8.883
ORFlab LGVDIAANTVIWDYK 5.311
ORFlab ANTVIWDYKRDAPAH 8.027
ORFlab AHIS TIG V C S MTDIA 8.491
ORFlab VFFDGRVDGQVDLFR 5.144
ORFlab KCVCSVIDLLLDDFV 7.415
ORFlab DFVEIIKS QDLS V V S 7.414
ORFlab PTGTLLVDSDLNDFV 9.015
ORFlab DLNDFVSDADSTLIG 8.287
ORFlab GDC ATVHT ANKWDLI 9.674
ORFlab ITEHSWNADLYKLMG 8.233
ORFlab SWNADLYKLMGHFAW 7.174
ORFlab MGHFAWWT AFVTN VN 9.577
ORFlab EQIDGYVMHANYIFW 5.142
ORFlab ANYIFWRNTNPIQLS 10.062
ORFlab TNPIQLS SYS LFDMS 5.499
Spike FSNVTWFHAIHVSGT 6.33
Spike PFNDGVYFASTEKSN 8.668
Spike V YFAS TEKS NIIRGW 7.461
Spike NIIRGWIFGTTLDSK 8.95
Spike PFLGVYYHKNNKSWM 11.766
Spike KNNKSWMESEFRVYS 14.037
Spike KNLREFVFKNIDGYF 5.877
Spike DS S S GWT AGAAA YYV 8.336
Spike ASVYAWNRKRISNCV 10.736
Spike AWNRKRISNCVADYS 10.174
Spike FKCY GVSPTKLNDLC 5.503
Spike GKIADYNYKLPDDFT 5.631
Spike GCVIAWNSNNLDSKV 11.056
Spike CVIAWNSNNLDSKVG 5.539
Spike PFERDISTEIY QAGS 6.454
Spike ENSVAYSNNSIAIPT 7.134
Spike TGIAVEQDKNTQEVF 5.393
Spike PPIKDFGGFNFSQIL 9.044
Spike QYTSALLAGTITSGW 7.296
Spike TITSGWTFGAGAALQ 5.587
Spike EN QKLIAN QFN S AIG 5.539
Overall, 207 sequences across the 9744 amino-acids of the SARS-CoV-2 proteome were identified, having a score >=5.
Surprisingly, a substantial number of gC1qR-binding motifs were thus identified in the SARS-CoV-2 proteins. In particular, such binding domains were identified in the following proteins: ORFlab, Spike, ORF3, E, ORF7b, ORF8, N and M.
Each of said proteins thus represents a relevant target for inhibition according to the invention.
Each of said peptides sequence represents an object of the invention.
Each of said proteins and peptides can act as PLA2-GIB cofactors in SARS-Cov2 infected patients.
Each of said peptides represents a strong inhibitor (or be used to construct a strong inhibitor) for use according to the invention
Each of said proteins or peptides can be used to produce relevant inhibitors, such as antibodies or derivatives thereof. Each of said proteins and peptides can be used as a reagent to detect the presence of anti- Cov-2 antibodies in a sample, particularly of a protective immunity.
Example 3: Analysis of binding peptides from Spike antigen Spike is the principal antigen exposed at the surface of the SARS-CoV-2 virus. It is an ideal target for antibody as it is expressed on the surface of the enveloped virus. Also, the spike antigen interacts with the receptor ACE-2.
The amino acid sequence of the Spike protein is represented as SEQ ID NO: 209.
The spike antigen comprises several functional domains, which are disclosed in the following Table:
Thirty (30) gC1qR-binding peptides derived from the spike antigen were identified (see example 2). Each of said peptides was further analyzed for selection/optimization. Figure 3 provides the PSSM score across the spike antigen (SEQ ID NO: 209). In this figure, PSSM scores are assigned to a 15aa peptide represented by a bar. The figure shows most peptides are located within the following sub-domains of the spike antigen: NTD, RBD, S2 and TM. Each peptide was analyzed for solvent accessibility, co-localization with B cell epitopes and/or with HLA class II binding motif-rich domains. The results are presented below. Said further analysis confirmed the relevance of the claimed therapeutic approach and further led to the selection of most preferred peptides and domains. 3.1. Co-localization of peptides with solvent accessible domains.
In this example, gC1qR binding motifs identified in example 2 were aligned with surface accessible domains within the spike antigen (SEQ ID NO: 209). Solvent accessible structure was measured from PDB structure using the “Dictionary of Secondary Structure of Proteins” (DSSP) developed by Kabsch and Sander (Kabsch W, Sander C. Dictionary of protein secondary structure: pattern recognition of hydrogen- bonded and geometrical features. Biopolymers. 1983 Dec;22(12):2577-637.). The DSSP program defines secondary structure, geometrical features and solvent exposure of proteins, given atomic coordinates in Protein Data Bank format (PDB) or macromolecular Crystallographic Information File format. Solvent exposure is given as the number of water molecules in possible contact with a residue (residue water exposed surface in Angstrom2). DSSP software package can be obtained from https://swift.cmbi.umcn.nl/gv/dssp/DSSP 3.html. The structure information for the spike antigen from SARS-CoV-2 used for solvent accessibility analysis was obtained from SWISS-MODEL (https://swissmodel.expasy.org/) based on a model number 4 (https://swissmodel.expasv.org/assess/7dVLxC/04) derived the cryo-EM structure of spike (PDB accession number: 6vsb) as established by Wrapp et al (Wrapp D, Wang N, Corbett KS, Goldsmith JA, Hsieh CL, Abiona O, Graham BS, McLellan JS. Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science. 2020 Mar 13;367(6483):1260-1263). The 3D model for spike SARS-CoV-2 is covering a sequence corresponding to residues 27 to 1146 from SEQ ID NO:209. For this analysis, only information from chain (a) - monomeric chain in the “up” conformation - was selected.
A solvent accessibility value was assigned to every amino-acid position of the sequence and was given in residue water exposed surface in Angstrom2 (A2).
Figure 4 presents the co-localization of high scoring gC1qR:PSSM motifs with solvent accessible domains within the spike protein sequence from SARS-CoV-2. It can be shown that most peptides in NTD, RBD and S2 domains are solvent accessible, and thus exposed and suitable for use as medicament or target.
3.2. Co-localization of peptides with B cell epitopes (Discotope)
In order to further validate the identified peptides, B cell epitopes were determined in the sequence of the Spike protein and then aligned with the peptide position.
B cell epitopes are regions of an antigen protein surface that are preferentially recognized by antibodies. B cell epitopes interact with an antibody based on the conformation adopted by the epitope, which is determined by the surface features of the involved epitope residues and the shape or tertiary structure of other segments of the antigen. Based on spatial structure, B-cell epitopes can be categorized as a continuous and discontinuous (nonlinear or conformational) epitopes. In the latter case, amino acid residues are in close contact due to the three-dimensional conformation. Several computational methods have been developed to predict the presence of B cell epitopes in protein structures such as for example BepiPred, CBtope, Ellipro and ABCpred. BEpro(PEPITO), SEPPA, EPITOPIA, EPCES , EPSVR, Bpredictor, and EPMeta. DiscoTope 1.1, DiscoTope 2.0.
We used DiscoTope 2.0 (http ://tool s.iedb.org/discotope/) in this example (Reliable B cell epitope predictions: Impacts of method development and improved benchmarking Jens Vindahl Kringelum, Claus Lundegaard, Ole Lund, and Morten Nielsen Plos Computational Biology, 2012). DiscoTope method incorporating a spatial neighborhood definition and half-sphere exposure as surface measure. J. V. Kringelum, C. Lundegaard, O. Lund, M. Nielsen. 2012. Reliable B cell epitope predictions: impacts of method development and improved benchmarking. PLoS Comput Biol. 8:(12):el002829. The method utilizes calculation of surface accessibility (estimated in terms of contact numbers) and an epitope propensity amino acid score. The final scores are calculated by combining the propensity scores of residues in spatial proximity and the contact numbers. The structure information for the spike antigen from SARS-CoV-2 used for DiscoTope 2.0 analysis was obtained from SWISS-MODEL (https://swissmodel.expasy.org/) based on a model number 4 (https://swissmodel.expasv.org/assess/7dVLxC/04) derived the cryo-EM structure of spike (PDB accession number: 6vsb) as established by Wrapp et al (Wrapp D, Wang N, Corbett KS, Goldsmith JA, Hsieh CL, Abiona O, Graham BS, McLellan JS. Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science. 2020 Mar 13;367(6483): 1260-1263). The 3D model for spike SARS-CoV-2 is covering a partial sequence corresponding to residues 27 to 1146 from SEQ ID NO: 209. For this analysis, only information from chain (a) - monomeric chain in the “up” conformation - was selected. The Discotope score is assigned to every amino-acid
position of the sequence. To identify B cell epitopes, a default threshold of -3.7 is generally applied.
Figure 5 presents co-localization of high scoring gC1qR-binding peptides with predicted B cell epitopes (Discotope) within the spike protein sequence from SARS-CoV-2.
As can be seen, peptides in NTD, RBD and S2 domains colocalize with predicted B cell epitopes, thus confirming their relevance as targets for antibody development.
3.3. Co-localization of peptides with HLA class II binding motif-rich domains
HLA class II binding motif-rich domains of the SARS-CoV-2 virus spike antigen were determined. Such domains were then aligned with the identified binding peptides.
Human leukocyte antigen (HLA) class II molecules are found on the surface of antigen- presenting cells where they present peptides derived from antigens to CD4+ T cells. HLA molecules are encoded in one of three loci called HLA -DR, -DP and -DQ. Each locus is comprised of many different allelic variants, which makes the HLA class II molecule highly polymorphic across the human population. Peptides presented by HLA class II molecules bind to a binding groove formed by residues of the HLA a- and the b-chains and a 9 amino-acid long core binding motif contained within the peptide antigen sequences. The specificity of the interaction is driven by the ability of lateral chain within the peptide core binding motif to interact with 4 to 5 anchor pockets lining the HLA binding groove. The specificity of the HLA binding groove is influence by the presence of polymorphic residues. Peptide:HLA class II binding is a prerequisite for the stimulation of CD4+ T-cells. There is a strong correlation between peptide: HLA class II binding strength and T cell response but high affinity peptides are rare. One additional challenge is to identify peptide that can bind a broad number of different HLA class II alleles in order to provide the ability of the peptide antigen to be immunogenic in a large proportion of the population. The phenotypic frequency of HLA class II molecules is highly variable in the human population. However, the identification of peptides having a high affinity across a range of highly prevalent HLA class II alleles that would be associated with broad population coverage may be achieved. HLA class II binding motifs are not evenly distributed along an antigen sequence and tend to form hotspots of overlapping motifs. Many different methods to predict the ability of peptide to bind to MHC class II molecules have been developed, including TEPITOPE, TEPITOPEpan, PROPRED, RANKPEP, SVRMHC, NetMHCIIpan, NetMHCII.
Analysis of HLA class II binding motifs across the spike SARS-CoV-2 sequence was performed using NetMHCII 2.3 (http://www.cbs.dtu.dk/services/NetMHCIE). NetMHCII 2.3 has been shown to be among the best methods for predicting binding affinities to a large number of HLA class II molecules (Jensen KK, Andreatta M,
Marcatili P, Buus S, Greenbaum JA, Yan Z, Sette A, Peters B, Nielsen M. Improved methods for predicting peptide binding affinity to MHC class II molecules. Immunology. 2018 Jul;154(3):394-406.). NetMHCII is based on ensembles of artificial neural networks that are trained on quantitative peptide binding affinity data from the Immune Epitope Database (IEDB). NetMHCII 2.3 server predicts binding of peptides to HLA class II molecules providing a range of information including the %-Rank and the core binding motif corresponding to a sequence of 9 amino-acids. Positive binding peptides are considered to have a %-rank below 10. In this example, NetMHCII 2.3 analysis was performed with a set of 20 HLA-DR alleles, 20 HLA-DQ, 9 HLA-DP: DRal*0101-DRβ1*0101, DRal*0101-DRβ1*0301, DRal*0101-DRβ 1*0401, DRal*0101-DRβ1*0701, DRal*0101-DRβ1*0801,
DRal*0101-DRβ 1*0802, DRal*0101-DRβ 1*0901, DRal*0101-DRβ 1*1001, DRal*0101-DRβ1*1101, DRal*0101-DRβ 1*1201, DRal*0101-DRβ 1*1301, DRal*0101-DRβ 1*1302,, DRal*0101-DRβ1*150, DRal *0101 -DRβ 1*1602, DRal*0101-DRβ3*0101, DRal*0101-DRβ3*0202, DRal*0101-DRβ3*0301, DRal*0101-DRβ4*0101, DRal*0101-DRβ4*0103, DRal*0101-DRβ5*0101, DQal *0101-DQβ 1*0501, DQal*0102-DQβ 1*0501, DQal *0102-DQβ 1*0502, DQal *0102-DQβ 1*0602, DQal*0103-DQβ 1*0603, DQal*0104-DQβ 1*0503, DQal *0201-DQβ 1*0202, DQal*0201-DQβ 1*0301, DQal*0201-DQβ 1*0303, DQal *0201-DQβ 1*0402, DQal*0301-DQβ 1*0301, DQal*0301-DQβ 1*0302, DQal *0303-DQβ 1*0402, DQal *0401 -DQβ 1 *0402, DQal *0501-DQβ 1*0201, DQal*0501-DQβ1*0301, DQal*0501-DQβ 1*0302, DQal*0501-DQβ 1*0303, DQal *0501-DQβ 1*0402, DQal *0601 -DQβ 1*0402, DPal*0103-DPβ1*0301, DPal*0103-DPβ 1*0401,, DPal*0103-DPβ1*0402, DPal*0103-DPβ 1*0601, DPal*0201-DPβ 1*0101, DPa 1 *0201 -DPβ 1*0501, DPal*0201-DPβ 1*1401,
DPa 1*0301 -ϋRb 1 *0402 and DPal*0103-1Pβ1*0201. Each 9 amino-acid long, HLA class II core binding motif identified by NetMHCII 2.3 was assigned a %-rank. The number of HLA class II core binding motifs having a %-rank <=10 across the 49 HLA class II alleles was then calculated for each 15 amino-acid long window scanning the spike SARS-CoV-2 sequence (SEQ ID NO:209).
Figure 6 presents co-localization results between the peptides and the predicted HLA class II binding motif-rich domains within the spike protein sequence from SARS-CoV- 2.
The results show a particular interest in peptides located in NTD and RBD sub-domains of the spike antigens. Particularly, the peptides listed in the following table exhibit remarkable immunogenic characteristics:
Example 4: Detection of anti-SARS-CoV-2 immunity in a sample
MATERIALS AND METHODS
Recombinant proteins and peptides- SARS-Cov-2 SI (Genbank accession #YP_009724390.1, which is identical to QHD43416) recombinant protein was purified from HEK293 cell culture and purchased from antibodies online (SARS-CoV-2 Spike Subunit SI (SARS-CoV-2 SI) (Active) protein, ABIN6952739, a.a. 11-683, purity >90%). SARS-Cov-2 peptides SARS-Cov-2_99-l 13 NH2-NIIRGWIFGTTLDSK- COOH, S ARS-Cov-2_ 147-161 NH2-KNNKSWMESEFRVYS-COOH, SARS-Cov- 2_253-267 NH2-DSS S GWT AGAAA YYV-COOH, SARS-Cov-2_348-362 NH2- ASVYAWNRKRISNCV-COOH, SARS-Cov-2_431-445 NH2-
GCVIAWNSNNLDSKV-COOH were ordered from Covalab (purity >98%).
Detection of anti-SARS-CoV-2 immunity in a sample- Detection of anti-SARS-Cov-2 SI IgG and IgM antibodies in SARS-Cov-2 patient plasma/serum is performed by ELISA on coated SARS-Cov-2 SI peptides or SI protein (Antibodies online, ABIN6952739). Microtiter plates (Nunc Maxisorp flat-bottom microplate, 44-2404-21, Thermofisher Scientific) are coated overnight at 4°C with 100 μl of a solution at 5 μg/ml of SI recombinant protein or SARS-Cov-2 peptides in carbonate buffer pH 9.6. Nonspecific binding sites are blocked with 300 μl of Casein blocking buffer (37528, Pierce Blocker Casein in PBS, Thermofisher Scientific). After washing, various dilutions in Casein blocking buffer of the anti-Sl specific Ab as positive control for IgM and IgG (Human IgGl or IgM ANTI SARS-COV-2 SPIKE (SI) ANTIBODY (CR3022), IgM: MAB 12423-50, IgG: MAB 12422-100, the native antigen company) at 125ng/ml and plasma/serum from SARS-Cov-2 patients or healthy donors (negative control) at 1:20 and gradually diluted 1:3 are incubated in quadruplicate for 2h at RT in 100 μl. Bound antibody are then detected by interaction with biotinylated rabbit anti-human IgG Ab (Anti-Human IgG antibody [EPR12700] (Biotin), ab201842, abeam) for IgG detection or biotinylated F(ab')2-Goat anti-Human IgM (F(ab')2-Goat anti-Human IgM (Heavy chain) Secondary Antibody, Biotin, A24486, Thermofish scientific), both at 1:10,000 dilution,
IOOmI/well for 2h, and is revealed by sequential reaction with HRP-Streptavidin at 1:10,000 (Peroxidase labeled Streptavidin ( SA-HRP) UP395888, Interchim, lh, room temp.) followed by the addition of 100 μl/well TMB substrate (UP664781, Interchim).
Example 5 : Production and selection of antibodies
Development of anti- SI antibodies- Antibodies specific for the SARS-Cov-2_147-161 sequence (KNNKSWMESEFRVYS), SARS-Cov-2_253-267 sequence (DSSSGWTAGAAAYYV) and SARS-Cov-2_431-445 sequence
(GC VIA WN S NNLDS KV) in SARS-Cov-2 SI (Genbank accession #YP_009724390.1) are generated by immunization of OF1 mice with the peptide NH2-C- KNNKS WMESEFRVY S -COOH, NH2-C-DS S S GWT AGAAA YYV -COOH or NH2-C- GCVIAWNSNNLDSKV-COOH conjugated to KLH (KLH-SARS-Cov-2_147-161, KLH-SARS-Cov-2_253-267, KLH-SARS-Cov-2_431-445, respectively). Mice are immunized 5 times every 3 weeks. First immunization is subcutaneous and intraperitoneal with 50 μg of peptides and complete freund’s adjuvant. Then mice are intraperitoneally injected three times with 50 μg of peptides and incomplete freund’s adjuvant. The antibody titer in serum is evaluated by EFISA on SARS-Cov-2_147-161, SARS-Cov-2_253-267, SARS-Cov-2_431-445 peptides and SARS-Cov-2 SI protein (ABIN6952739). The mouse with the best titer on protein and peptide is selected and injected intravenously with 5 μg of SARS-Cov-2 peptide (KLH-SARS-Cov-2_147-161,KLH-SARS-Cov-2_253-267 or KLH-SARS-Cov-2_431-445) alone. Three days later the spleen is collected and cells are fused with myeloma (Sp2/0-Agl4) cells to generate hybridoma. Among supernatants from fusion, the best 120 positive on BSA-SARS-Cov- 2 peptides are selected and tested another time on BS A-S ARS-Cov-2 peptide and S 1. The isotypes of the best clones are determined and they are frozen including 15 clones that are maintained in cell culture to obtain stable clone, 5 clones per peptide. These 15 clones, are seeded in High Yielding Performance Flask (hyperflask) for cell culture to produce 560 ml of supernatant with 10% Low IgG fetal bovine serum (FBS) in DMEM medium. IgG are purified from supernatant by affinity chromatography on protein A in low endotoxin condition (<10 EU / mg, FAF test). Antibodies binding to BSA-SARS-Cov-2 and S 1 is verified and the cDNA sequence coding the variable regions of these antibodies is determined.
ELISA to screen new anti-Sl antibodies produced- EFISA on peptides or proteins is performed on microtiter plates coated overnight at room temperature (RT) with 50 pi of a solution in PBS (pH7.4) of BSA-SARS-Cov-2_147-161, BSA-SARS-Cov- 2_253-267, BSA-SARS-Cov-2_431-445, SI protein at 2 μg/ml. After washing, wells are then blocked lh at RT with 150 pi of PBS/2.5% skimmed milk. After washing, various dilutions in PBS/0.05%Tween 20/0.5%BSA of Ab from 100-0.0001 μg/ml are incubated for 2h at RT in 50 pi. Wells are washed and antibody binding is revealed after lh of incubation at RT in 50 pi of PBS/0.05%Tween 20/0.5%BSA with
HRP-conjugated affiniPure F(ab)'2 fragment Goat anti-Mouse lgG (1:5000, Jackson 115-036-008) followed by TMB addition (10 min at RT, SureBlue™ TMB Microwell Peroxidase Substrate Kit, KPL 52-00-01, VWR).
Example 6 : Inhibition of sPLA2-GIB affects SARS-CoV-2 plasma inhibitor effect
Purification of Human CD4 T -lymphocytes- Venous blood was obtained from healthy volunteers through the EFS (Etablissement Francais du Sang, Centre Necker-Cabanel, Paris). CD4 T-cells were purified from whole blood using RosetteSep Human CD4+ T cell Enrichment Cocktail (Stem Cell, 15062). This cocktail contains mouse and rat monoclonal antibodies purified from mouse ascites fluid or hybridoma culture supernatant, by affinity chromatography using protein A or Protein G sepharose. These antibodies are bound in bispecific tetrameric antibody complexes which are directed against cell surface antigens on human hematopoietic cells (CD8, CD16, CD 19, CD36, CD56 CD66b, TCRγ/d) and glycophorin A on red blood cells. The rosetteSep antibody cocktail crosslinks unwanted cells in human whole blood to multiple red blood cells, forming immunorosettes . This increases the density of unwanted cells, such that they pellet along with the free red blood cells when centrifuged over a buoyant density medium such as lymphocytes separation medium (Eurobio, CMSMSLOl-01).
Whole blood is incubated with RosetteSep Human CD4+ T cell Enrichment Cocktail at 50 μl/ml for 20 minutes at room temperature under gentle shaking (100 rpm), diluted with an equal volume of PBS+2% foetal bovine serum (FBS) and mixed gently. The diluted samples are centrifuged 20 minutes at 1200 X g on top of lymphocytes separation medium. The enriched cells are then collected from the density medium at plasma interface and washed twice with PBS+2% FBS. Cells are subsequently resuspended in RPMI 1640 medium (Lonza) supplemented with 5% FBS, 50 mM HEPES pH 7.4, glutamine, penicillin, streptomycin and fungizone (complete medium), counted with a Moxi Z mini automated cell counter (ORFLO, MXZ000). Cells suspension is adjusted at 7xl06 cells/ml and equilibrated at least 2 h at 37°C in a 5% CO2 humidified atmosphere.
The enriched CD4-T cell population is controlled by flow cytometry on a cytoflex (Beckman coulter). The quiescence of recovered CD4 T-cells is controlled by the low level of IL-2Roα (CD25). CD4 T cells are labeled with anti-Human CD3 eFluor780 (eBioscience, clone UCHT1, 47-0038-42), anti-Human CD25-PE (Biolegend, clone BC96, 302605) and anti-human CD4-PerCP (BD, clone SK3, 345770). The enriched CD4-T cell population contains >95% CD3+CD4+ and less than 8% of CD25+.
PLA2-G1B and SARS-Cov-2 SI protein immunodepletion to study plasma cofactor activity- To study the PLA2-G1B and SARS-Cov-2 SI protein role in SARS-Cov-2 patient plasma inhibitory activity, plasma are immunodepleted with anti-PLA2GlB
antibodies and anti-Sl antibodies. Briefly, 1 ml VP or HD plasma is incubated with 100 μg of rabbit anti-PLA2GlB pAb (produced at BIOTEM) or control isotype (Rabbit Gamma Globulin, 31887, Invitrogen) to immunodeplete PLA2-G1B, or with 100 μg of anti-Sl specific Ab (Human IgGl ANTI SARS-COV-2 SPIKE (SI) ANTIBODY (CR3022), MAB 12422- 100, the native antigen company) or the control goat polyclonal antibody (Ultra-LEAF™ Purified Human IgGl Isotype Control Recombinant Antibody, 403502, Biolegends) to immunodeplete SI, in 1.5-ml Eppendorf tubes overnight (Test tube-rotor, 34528, Snijders, Netherland) at 4°C. Then, 200 pi Protein G Sepharose 4 Fast Flow beads (17-0618-01, GE healthcare), that had been washed three times in PBS/1%BSA, is added to each sample and the samples incubated for 3 h at 4°C. Beads are removed by centrifugation of the samples at 400 x g for 2 min at 4°C, collection of the supernatant then centrifugation at 16,100 x g for 15 min at 4°C.
SARS-Cov-2 patient plasma, SI protein or SARS-Cov-2 peptides bioassay on CD4 T cells and labelling of specific proteins for optical microscopy- Equilibrated purified CD4 T- cells are loaded (3.5x105cells/50μl in complete medium) on poly-L-Lysine-coated (Sigma, P8920) round coverslips (14mm-diameter, Marienfeld) in 24-well polystyrene plates at 37°C in a thermo-regulated water and mixed with 50pl of a suspension in PBS BSA1% containing peptides, recombinant proteins together with recombinant PLA2-GIB or not or containing SARS-Cov-2 patient plasma (1 or 3%), PLA2-GlB-depleted or Sl- depleted or not plasma (1 or 3%), or healthy donor plasma. The cells suspension was either pretreated with 40pl of peptides, recombinant protein dilutions in PBS BSA1% for 15 minutes with subsequent addition of IOmI PLA2-GIB (5nM at the end) for 30 minutes or directly treated with 50m1 of dilution in PBS BSA 1% with peptides or recombinant protein together with PLA2-GIB (5nM at the end) for 45 minutes. Cells were activated for 15 minutes with 2 nM recombinant glycosylated human IL-7 (Accrobio System). Cells supernatant was removed and cells were fixed by addition of 500m1 of a 4% paraformaldehyde solution in PBS (Fisher, PFA 32% Electron Microscopy Science, 15714) for 15 minutes at 37 °C and then permeabilized for 20 min in 500m1 of ice-cold 90% methanol/water solution.
Cells were then rehydrated for 15 min in PBS plus 5% fetal bovine serum (FBS) and then labeled. Thus, slides were washed twice after methanol treatment in PBS and rehydrated for 15 min in PBS supplemented with 5% FBS at room temperature. Slides were labelled with primary antibodies (1/120) in 60 μl of PBS 5% FBS for lh, washed in PBS buffer 15 times, 5 times in PBS/FBS buffer and then stained with secondary antibodies (1/300) for lh. Slides were washed 5 times in PBS 5% FBS buffer, rinsed 15 times in PBS and then mounted in fresh Prolong Gold Antifade (ThermoFisher Scientific, P36930) mounting medium for confocal microscopy. The primary antibodies used consisted of rabbit anti-pSTAT5 (pY694, 9359, Cell Signalling), mouse anti-CD4 (BD Pharmingen, 555344) and secondary antibodies were Donkey anti-mouse IgG- AF488 (Invitrogen, A21202) and Donkey anti-rabbit IgG-AF555 (Invitrogen, A31572).
Confocal Microscopy- Images were acquired above the diffraction limit on an inverted laser scanning confocal microscope (LSM700, Zeiss), with an oil-immersion plan- apochromatic 63x/1.4 NA objective lens (Zeiss) for PFA-fixed cells. Images were acquired and analyzed with the ZEN software (Zeiss).
SARS-Cov-2 patient plasma, SI protein or SARS-Cov-2 peptides effect on gene expression in IL-7 -stimulated CD4 T cells- Equilibrated purified CD4 T-cells were seeded at 106cells/ml in 2 ml of 2.5% FBS RPMI medium in 6-well polystyrene plates (10110151, Thermofischer scientific) at 37°C. After lh30 of resting, cells were treated overnight (16 h) with PBS (control buffer), SI protein or peptides, PLA2-G1B protein, PLA2-G1B with SI protein or peptides, SARS-Cov2 patient or healthy donor plasma (1, 3, 5 or 10%), or PLA2-G IB-depleted or S 1-depleted SARS-Cov2 patient. Then cells were stimulated for 6h with IL-7 (2 nM recombinant glycosylated human IL-7, Accrobio System). Gene expression was then analyzed by quantitative real-time PCR. cDNA was obtained from 1 μg total RNA as follows: total RNA was extracted using QIAshredder (79654, Qiagen) and the RNeasy Kit (74104, Qiagen). Extracted RNA was treated with DNase I (RNase-Free DNase Set, 79254, Qiagen) and reverse transcribed using Superscript III reverse transcriptase (18080-044, Invitrogen) according to the manufacturer’s instructions. Specific TaqMan probes from Applied Biosystems were used to quantify IL-7 regulated gene expression and b2m (used as endogenous reference, Hs00984230_ml) mRNA levels by qRT-PCR using TaqMan Universal Master Mix II with UNG (4440038, Thermofisher), according to the manufacturer’s instructions. Triplicate Ct values for IL-7 regulated gene were analyzed using the comparative Ct (ΔΔCt) method (AppliedBiosystems). The relative amount of mRNA (2-ΔΔCt) was obtained by normalizing against the b2m endogenous references.
Example 7: Peptide SEQ ID NOs: 164 and 168 affect SARS-CoV-2
Purified SARS-Cov-2 particles were purified from African green monkey kidney Vero E6 (ATCC® CRL-1586) cell culture.
To test SARS-Cov-2_348-362 and SARS-Cov-2_431-445 peptides effect on SARS- Cov-2 infection viral infection of Vero E6 cells was followed by qRT-PCR. Vero E6 cells were seeded (60,000/well) in 500- μl 24-well plates in DMEM 10% FBS. 24h later cells were treated for lh with SARS-Cov-2_348-362 and SARS-Cov-2_431-445 or a control scrambled peptide. Then cells were infected with SARS-Cov-2 at different multiplicity of infection (MOI) 0.1 , 0.5 or 1 and viral RNA quantity at 8 h, 24 h and 48 h post-infection in cell supernatant and in cells was quantified by qRT-PCR, as described in Zhou et al. Nature 2020. A standard curve from RNA purified directly from viral particles was used to express results in equivalent pfu/ml.
Effect of SARS-Cov-2 SI protein or peptides on PLA2-G1B activity on CD4 T cells
membrane:
PLA2-GIB enzymatic assay on [3H] arachidonic acid labelled CD4 T cells or Jurkat E6.1 T cells - Purified CD4 T-cells were incubated for 16h at 2xl06 cells/ml with lμCi/ml of arachidonic acid [5, 6, 8, 9,11,14,15-3H(N)] (Perkin Elmer, NET298Z250UC) in RPMI 1640 medium (Lonza) supplemented with 10% FBS, 50 mM HEPES pH 7.4, glutamine, penicillin, streptomycin and fungizone at 2 ml/well in 6-well plates at 37 °C in a 5% C02 humidified atmosphere. Cells were washed twice with RPMI with 10% FBS by centrifugation at 580xg for 10 minutes at room temperature and then frozen in 90% FBS 10% DMSO at 107 cells/ml/vial at -80°C. Percent of [3H] arachidonic acid in CD4 T cells is the (1 minus ratio of [3H] arachidonic acid in the supernatant of CD4 T cells without cells (cpm/ml) on total [3H] arachidonic acid in supernatant and cells (cpm/ml).
Jurkat E6.1 T cells (ECACC 88042803) or gC1qR KO Jurkat E6.1 T cells were incubated for 17h at 5x105 cells/ml with 1 μCi/ml of arachidonic acid [5,6,8,9,11,14,15- 3H(N)] (Perkin Elmer, NET298Z250UC) in RPMI 1640 medium (Lonza) supplemented with 10% FBS, 50 mM HEPES pH 7.4, glutamine, penicillin, streptomycin and fungizone at 2 ml/well in 6-well plates at 37°C in a 5% C02 humidified atmosphere. Cells were washed twice with RPMI with 10% FBS by centrifugation at 300xg for 10 minutes at room temperature and then frozen in 90% FBS 10% DMSO at 107 cells/ml/vial at -80°C. Percent of [3H] arachidonic acid in CD4 T cells is the (1 minus ratio of [3H] arachidonic acid in the supernatant of CD4 T cells without cells (cpm/ml) on total [3H] arachidonic acid in supernatant and cells (cpm/ml).
To test PLA2-GIB activity on [3H] arachidonic acid labelled CD4 T lymphocytes, cells were unfrozen in 10% FBS RPMI preheated at 37°C by centrifugation at 580xg for 10 minutes at room temperature, washed twice in 2.5% FBS RPMI, and equilibrated at 2xl05 CD4 T cells/400 μl/well in 24-well polystyrene plates for lh30 at 37°C in a 5% C02 humidified atmosphere. Then IOOmI of recombinant SI protein (Antibodies online, ABIN6952739) or vehicle dilution in 2.5% FBS RPMI was added to each well for 2h. Cells and supernatant were collected in eppendorf tubes and centrifuged at 580xg for 10 minutes at room temperature. The [3H] arachidonic acid released in cell supernatant was quantified in 300m1 with 16 ml of Ultima gold (Perkin Elmer, 6013329) in low diffusion vials (Perkin Elmer, 6000477) on a counter (tri-Carb 2800 TR liquid scintillation analyzer, Perkin Elmer).
To test PLA2-GIB activity on [3H] arachidonic acid labelled Jurkat E6.1 T lymphocytes, cells were unfrozen in 10% FBS RPMI preheated at 37°C by centrifugation at 300xg for 10 minutes at room temperature, washed twice in 2.5% FBS RPMI, and equilibrated at 5x104 or 105 Jurkat E6.1 T
in 24-well polystyrene plates for 1h30 at 37°C in a 5% CO2 humidified atmosphere. For peptide treatments, cells were pretreated for 21h with 50m1 per well of peptide solutions at 110μM or 55μM in 2.5%
FBS RPMI. Then 50 μl per well of PLA2GIB at 630nM or 2mM 2.5% FBS RPMI or medium alone were added for 2h. Cells and supernatant were collected in eppendorf tubes and centrifuged at 580xg for 10 minutes at room temperature. The [3H] arachidonic acid released in cell supernatant was quantified in 300μl with 16 ml of Ultima gold (Perkin Elmer, 6013329) in low diffusion vials (Perkin Elmer, 6000477) on a counter (tri-Carb 2800 TR liquid scintillation analyzer, Perkin Elmer).
Results are expressed as PLA2GIB activity (release of [3H] arachidonic acid in the supernatant of cells treated with peptide together with PLA2G1B minus spontaneous release of [3H] arachidonic acid by cells in cpm/ml) or ΔPLA2G1B activity with peptides minus activity with Scrambled peptide (release of [3H] arachidonic acid in the supernatant of cells treated with peptide minus release of [3H] arachidonic acid by cells treated with Scrambled in cpm/ml).
List of Sequences
SEQ ID NO:l : SARS-CoV-2 proteome (9744 amino acids)
MESLVPGFNEKTHVQLSLPVLQVRDVLVRGFGDSVEEVLSEARQHLKDGTCGL VEVEKGVLPQLEQPYVFIKRSDARTAPHGHVMVELVAELEGIQYGRSGETLGV LVPH V GEIP V A YRKVLLRKN GNKG AGGHS Y G ADLKS FDLGDELGTDP YEDF QE NWNTKHS S GVTRELMRELN GGA YTRYVDNNFCGPDGYPLECIKDLLARAGKA S CTLS EQLDFIDTKRG V Y CCREHEHEIA W YTERS EKS YELQTPFEIKLAKKFDTF NGECPNFVFPLNSIIKTIQPRVEKKKLDGFMGRIRSVYPVASPNECNQMCLSTLM KCDHCGETSWQTGDFVKATCEFCGTENLTKEGATTCGYLPQNAVVKIYCPAC HNSEVGPEHSLAEYHNESGLKTILRKGGRTIAFGGCVFSYVGCHNKCAYWVPR ASANIGCNHTGVVGEGSEGLNDNLLEILQKEKVNINIVGDFKLNEEIAIILASFSA STSAFVETVKGLDYKAFKQIVESCGNFKVTKGKAKKGAWNIGEQKSILSPLYAF ASE A ARV VRS IF S RTLET AQN S VR VLQKA AITILDGIS Q Y S LRLID AMMFT S DLA TNNLVVM A YITGGV V QLT S QWLTNIFGT V YEKLKPVLDWLEEKFKEGVEFLRD GWEIVKFISTCACEIVGGQIVTCAKEIKESVQTFFKLVNKFLALCADSIIIGGAKL KALNLGETFVTHSKGLYRKCVKSREETGLLMPLKAPKEIIFLEGETLPTEVLTEE VVLKTGDLQPLEQPTSEAVEAPLVGTPVCINGLMLLEIKDTEKYCALAPNMMV TNNTFTLKGGAPTKVTFGDDTVIEVQGYKSVNITFELDERIDKVLNEKCSAYTV ELGTE VNEF AC V V AD A VIKTLQP V S ELLTPLGIDLDE W S M AT Y YLFDES GEFKL AS HM Y CS F YPPDEDEEEGDCEEEEFEPS T Q YE Y GTEDD Y QGKPLEF GAT S A ALQ PEEEQEEDWLDDDS QQTV GQQDGSEDN QTTTIQTIVE V QPQLEMELTPVV QTIE VN S FS G YLKLTDN V YIKN ADI VEE AKKVKPT V V VN A AN V YLKHGGG V AG ALN KATNN AMQ VES DD YIATN GPLKV GGS C VLS GHNLAKHCLH V V GPN VNKGEDI QLLKS A YENFN QHE VLLAPLLS AGIFG ADPIHS LR V C VDT VRTN V YL A VFD KNL YDKLVSSFLEMKSEKQVEQKIAEIPKEEVKPFITESKPSVEQRKQDDKKIKACVE E VTTTLEETKFLTENLLLYIDIN GNLHPDS ATLVSDIDITFLKKD APYIV GD VV QE GVLTAVVIPTKKAGGTTEMLAKALRKVPTDNYITTYPGQGLNGYTVEEAKTVL KKC KS AFYILPS IIS NEKQEILGT VS WNLREMLAH AEETRKLMP VC VETKAIV S T IQRKYKGIKIQEG V VD Y G ARFYFYT S KTT V AS LINTLNDLNETL VTMPLG Y VTH GLNLEE A AR YMRS LK VP AT V S VS S PD A VT A YN G YLT S S S KTPEEHFIETIS LAGS YKD W S YS GQS T QLGIEFLKRGDKS V Y YT S NPTTFHLDGE VITFDNLKTLLS LRE VRTIKVFTTVDNINLHTQVVDMSMTYGQQFGPTYLDGADVTKIKPHNSHEGKT FYVLPNDDTLRVEAFEYYHTTDPSFLGRYMSALNHTKKWKYPQVNGLTSIKW ADNNCYLATALLTLQQIELKFNPPALQDAYYRARAGEAANFCALILAYCNKTV GELGDVRETMSYLFQHANLDSCKRVLNVVCKTCGQQQTTLKGVEAVMYMGT LS YEQFKKG V QIPCTCGKQ ATKYL V QQES PF VMMS APP AQ YELKHGTFTC AS E YTGNYQCGHYKHITSKETLYCIDGALLTKSSEYKGPITDVFYKENSYTTTIKPVT YKLDG V V CTEIDPKLDN Y YKKDN S YFTEQPIDL VPN QP YPN AS FDNFKF VCDNI KF ADDLN QLT G YKKP AS RELKVTFFPDLN GD V V AID YKH YTPS FKKG AKLLHK PIV WH VNN ATNKAT YKPNTWCIRCLW S TKP VET S NS FD VLKS ED AQGMDNL A CEDLKPVSEE VVENPTIQKD VLECNVKTTE VV GDIILKPANNSLKITEEV GHTDL
M A A Y VDN S S LTIKKPNELS RVLGLKTLATHGL A A VN S VPWDTIAN Y AKPFLNK VVSTTTNIVTRCLNRVCTNYMPYFFTLLLQLCTFTRSTNSRIKASMPTTIAKNTV KS VGKFCLE AS FN YLKS PNF S KLINIIIWFLLLS VCLGS LIY S T A ALG VLMS NLGM PS Y CTG YREG YLN S TN VTIAT Y CTGS IPC S VCLS GLDS LDT YPS LETIQITIS S FKW DLTAFGLVAEWFLAYILFTRFFYVLGLAAIMQLFFSYFAVHFISNSWLMWLIINL V QM APIS AM VRM YIFF AS FY Y VWKS Y VH V VDGCN S S T CMMC YKRNR ATR VE CTTIVN G VRRS F Y V Y AN GGKGFC KLHN WNC VN CDTFCAGS TFIS DE V ARDLSL QFKRPINPTDQS S YIVDS VT VKN GS IHL YFDKAGQKT YERHS LS HF VNLDNLRA NNTKGSLPINVIVFDGKSKCEESSAKSASVYYSQLMCQPILLLDQALVSDVGDS AE V A VKMFD A Y VNTFS S TFN VPMEKLKTL V AT AE AELAKN V S LDN VLS TFIS A ARQGFVDSDVETKDVVECLKLSHQSDIEVTGDSCNNYMLTYNKVENMTPRDL G ACIDC S ARHIN AQ V AKS HNIALIWN VKDFMS LS EQLRKQIRS A AKKNNLPFKL TCATTRQVVNVVTTKIALKGGKIVNNWLKQLIKVTLVFLFVAAIFYLITPVHVM SKHTDFSSEIIGYKAIDGGVTRDIASTDTCFANKHADFDTWFSQRGGSYTNDKA CPLIA A VITRE V GF V VPGLPGTILRTTN GDFLHFLPRVF S A V GNIC YTPS KLIE YT DFAT S AC VLA AECTIFKD AS GKP VP Y C YDTN VLEGS V A YES LRPDTR Y VLMDG S IIQFPNT YLEGS VRV VTTFDS E Y CRHGT CERS E AG VC VS T S GRW VLNND Y YRS LPGVFCGVDAVNLLTNMFTPLIQPIGALDISASIVAGGIVAIVVTCLAYYFMRFR RAFGE Y S H V V AFNTLLFLMS FT VLCLTP V Y S FLPG V Y S VIYL YLTF YLTND VS FL AHIQWMVMFTPLVPFWITIAYIICISTKHFYWFFSNYLKRRVVFNGVSFSTFEEA ALCTFLLNKEM YLKLRS D VLLPLT Q YNRYL ALYNKYK YFS G AMDTT S YRE A A CCHL AKALNDFS N S GS D VL Y QPPQTS ITS A VLQS GFRKM AFPS GKVEGCM V Q V TCGTTTLNGLWLDDVVYCPRHVICTSEDMLNPNYEDLLIRKSNHNFLVQAGNV QLRVIGHS MQNC VLKLKVDT ANPKTPKYKF VRIQPGQTF S VLAC YN GS PS G V Y QCAMRPNFTIKGSFLNGSCGSVGFNIDYDCVSFCYMHHMELPTGVHAGTDLEG NFY GPFVDRQT AQAAGTDTTITVNVLAWLY AA VIN GDRWFLNRFTTTLNDFNL V AMKYN YEPLTQDH VDILGPLS AQTGIA VLDMC AS LKELLQN GMN GRTILGS A LLEDEFTPFDVVRQCSGVTFQSAVKRTIKGTHHWLLLTILTSLLVLVQSTQWSL FFFLYENAFLPFAMGIIAMSAFAMMFVKHKHAFLCLFLLPSLATVAYFNMVYM PASWVMRIMTWLDMVDTSLSGFKLKDCVMYASAVVLLILMTARTVYDDGAR RVWTLMN VLTL V YK V Y Y GN ALDQ AIS MW ALIIS VT S NY S G V VTT VMFLARGI VFMCVEY CPIFFITGNTLQCIMLVY CFLGYFCTCYFGLFCLLNRYFRLTLGVYD YLV S TQEFR YMN S QGLLPPKN S ID AFKLNIKLLG V GGKPCIKV AT V QS KMS D VK CTS V VLLS VLQQLR VES S S KLW AQC V QLHNDILL AKDTTE AFEKM V S LLS VLLS MQGAVDINKLCEEMLDNRATLQAIASEFSSLPSYAAFATAQEAYEQAVANGDS E V VLKKLKKS LN V AKS EFDRD A AMQRKLEKM ADQ AMT QM YKQ ARS EDKR A KVTSAMQTMLFTMLRKLDNDALNNIINNARDGCVPLNIIPLTTAAKLMVVIPDY NT YKNTCDGTTFT Y AS ALWEIQQ V VD ADS KIV QLS EIS MDN S PNL A WPLIVT AL RAN S A VKLQNNELS P V ALRQMS C A AGTT QT ACTDDN ALA Y YNTTKGGRFVL A LLS DLQDLKW ARFPKS DGT GTIYTELEPPCRFVTDTPKGPKVKYLYFIKGLNNL NRGM VLGS LA AT VRLQ AGN ATE VP AN S T VLS FC AF A VD A AKA YKD YL AS GGQ PITNCVKMLCTHTGTGQAITVTPEANMDQESFGGASCCLYCRCHIDHPNPKGFC
DLKGKY V QIPTTC ANDP V GFTLKNT VCT V C GM WKG Y GCS CDQLREPMLQS AD
AQSFLNRVCGVSAARLTPCGTGTSTDVVYRAFDIYNDKVAGFAKFLKTNCCRF
QEKDEDDNLIDS YF V VKRHTF S NY QHEETI YNLLKDCP A V AKHDFFKFRID GD
MVPHISRQRLTKYTMADLVYALRHFDEGNCDTLKEILVTYNCCDDDYFNKKD
W YDFVENPDILRV Y ANLGER VRQ ALLKT V QFCD AMRN AGI V G VLTLDN QDLN
GNWYDFGDFIQTTPGSGVPVVDSYYSLLMPILTLTRALTAESHVDTDLTKPYIK
WDLLKYDFTEERLKLFDRYFKYWDQTYHPNCVNCLDDRCILHCANFNVLFST
VFPPT S F GPLVRKIF VDG VPFV V S T G YHFRELG V VHN QD VNLHS S RLS FKELLV
YAADPAMHAASGNLLLDKRTTCFSVAALTNNVAFQTVKPGNFNKDFYDFAVS
KGFFKEGSSVELKHFFFAQDGNAAISDYDYYRYNLPTMCDIRQLLFVVEVVDK
YFDCYDGGCINANQVIVNNLDKSAGFPFNKWGKARLYYDSMSYEDQDALFAY
TKRN VIPTITQMNLKY AIS AKNRART V AG VS IC S TMTNRQFHQKLLKS IA ATRG
ATVVIGTSKFYGGWHNMLKTVYSDVENPHLMGWDYPKCDRAMPNMLRIMAS
LVLARKHTT CC S LS HRFYRLANEC AQ VLS EM VMCGGS L Y VKPGGT S S GD ATT
AYANSVFNICQAVTANVNALLSTDGNKIADKYVRNLQHRLYECLYRNRDVDT
DF VNEFY A YLRKHF S MMILS DD A V V CFN S T Y AS QGLV AS IKNFKS VL Y Y QNN V
FMSEAKCWTETDLTKGPHEFCSQHTMLVKQGDDYVYLPYPDPSRILGAGCFVD
DIVKTDGTLMIERFVSLAIDAYPLTKHPNQEYADVFHLYLQYIRKLHDELTGHM
LDM Y S VMLTNDNT S RYWEPEF YE AM YTPHT VLQ A V G AC VLCNS QT S LRC G AC
IRRPFLCCKCC YDH VIS T S HKLVLS VNP Y V CN APGCD VTD VT QL YLGGMS Y YC
KSHKPPISFPLCANGQVFGLYKNTCVGSDNVTDFNAIATCDWTNAGDYILANT
CTERLKLFAAETLKATEETFKLSYGIATVREVLSDRELHLSWEVGKPRPPLNRN
YVFTG YRVTKN S KV QIGEYTFEKGD Y GD A VVYRGTTTYKLNV GD YFVLTSHT
VMPLS APTLVPQEH Y VRITGL YPTLNIS DEFS S N V ANY QKV GMQKY S TLQGPPG
TGKS HFAIGL ALY YPS ARIV YT AC S H A A VD ALCEKALKYLPIDKCS RIIP ARARV
ECFDKFKVN S TLEQ Y VFCT VN ALPETT ADIV VFDEIS M ATN YDLS V VN ARLRAK
HYVYIGDPAQLPAPRTLLTKGTLEPEYFNSVCRLMKTIGPDMFLGTCRRCPAEI
VDTVSALVYDNKLKAHKDKSAQCFKMFYKGVITHDVSSAINRPQIGVVREFLT
RNP A WRKA VFIS P YN S QN A V AS KILGLPTQT VDS S QGS E YD Y VIFTQTTET AHS
CN VNRFN V AITR AKV GILCIMS DRDL YDKLQFT S LEIPRRN V ATLQ AEN VT GLF
KDCSKVITGLHPTQAPTHLSVDTKFKTEGLCVDIPGIPKDMTYRRLISMMGFKM
NYQVNGYPNMFITREEAIRHVRAWIGFDVEGCHATREAVGTNLPLQLGFSTGV
NLV A VPTG Y VDTPNNTDF S RV S AKPPPGDQFKHLIPLM YKGLPWN V VRIKI V Q
MLSDTLKNLSDRVVFVLWAHGFELTSMKYFVKIGPERTCCLCDRRATCFSTAS
DT Y AC WHHS IGFD Y V YNPFMID V QQW GFTGNLQS NHDL YCQ VHGN AH V AS C
DAIMTRCLAVHECFVKRVDWTIEYPIIGDELKINAACRKVQHMVVKAALLADK
FPVLHDIGNPKAIKCVPQADVEWKFYDAQPCSDKAYKIEELFYSYATHSDKFTD
G V CLFWN CN VDRYP AN S IV CRFDTRVLS NLNLPGCDGGS LY VNKH AFHTP AFD
KS AFVNLKQLPFF Y Y S DS PCES HGKQ V V S DID Y VPLKS ATCITRCNLGG A VCRH
HANEYRLYLDAYNMMISAGFSLWVYKQFDTYNLWNTFTRLQSLENVAFNVV
NKGHFDGQQGEVPVSIINNTVYTKVDGVDVELFENKTTLPVNVAFELWAKRNI
KPVPEVKILNNLGVDIAANTVIWDYKRDAPAHISTIGVCSMTDIAKKPTETICAP
LTVFFDGRVDGQVDLFRNARNGVLITEGSVKGLQPSVGPKQASLNGVTLIGEA VKTQFNYYKKVDGVVQQLPETYFTQSRNLQEFKPRSQMEIDFLELAMDEFIER YKLEGYAFEHIVYGDFSHSQLGGLHLLIGLAKRFKESPFELEDFIPMDSTVKNYF ITD AQT GS S KC V C S VIDLLLDDFVEIIKS QDLS V V S KV VKVTID YTEIS FMLW C K DGHVETFYPKLQSSQAWQPGVAMPNLYKMQRMLLEKCDLQNYGDSATLPKGI MMN V AKYTQLC Q YLNTLTLA VP YNMRVIHF GAGS DKG V APGT A VLRQWLPT GTLLVDSDLNDFVSDADSTLIGDCATVHTANKWDLIISDMYDPKTKNVTKEND SKEGFFTYICGFIQQKLALGGSVAIKITEHSWNADLYKLMGHFAWWTAFVTNV NASSSEAFLIGCNYLGKPREQIDGYVMHANYIFWRNTNPIQLSSYSLFDMSKFPL KLRGT A VMS LKEGQINDMILS LLS KGRLIIRENNR V VIS S D VL VNNMFVFL VLLP LV S S QC VNLTTRT QLPP A YTN S FTRG V Y YPDKVFRS S VLHS T QDLFLPFFS NVT WFHAIHVSGTNGTKRFDNPVLPFNDGVYFASTEKSNIIRGWIFGTTLDSKTQSLL IVNN ATN V VIK V CEF QFCNDPFLG V Y YHKNNKS WMES EFR V Y S S ANN CTFE Y V SQPFLMDLEGKQGNFKNLREFVFKNIDGYFKIYSKHTPINLVRDLPQGFSALEPL VDLPIGINITRFQTLL ALHRS YLTPGDS S S GWT AG A A AY YV G YLQPRTFLLKYN EN GTITD A VDC ALDPLS ETKCTLKS FT VEKGI Y QT S NFRV QPTES IVRFPNITNLC PF GE VFN ATRF AS V Y A WNRKRIS NC V AD Y S VL YN S AS FS TFKC Y G VS PTKLND LCFTNVYADSFVIRGDEVRQIAPGQTGKIADYNYKLPDDFTGCVIAWNSNNLDS KVGGNYNYLYRLFRKSNLKPFERDISTEIY QAGSTPCNGVEGFNCYFPLQS Y GF QPTN G V GY QP YRV V VLS FELLH AP AT VCGPKKS TNL VKNKC VNFNFN GLTGTG VLTESNKKFLPFQQFGRDIADTTDAVRDPQTLEILDITPCSFGGVSVITPGTNTSN QVAVLYQDVNCTEVPVAIHADQLTPTWRVYSTGSNVFQTRAGCLIGAEHVNNS YECDIPIG AGIC AS Y QTQTN S PRR ARS V AS QS IIA YTMS LG AEN S V AY S NN S IAIP TNFTIS VTTEILP V S MTKT S VDCTM YICGDS TEC S NLLLQ Y GS FCT QLNRALTGI A VEQDKNTQEVFAQ VKQIYKTPPIKDFGGFNFS QILPDPS KPS KRSFIEDLLFNK VTLADAGFIKQYGDCLGDIAARDLICAQKFNGLTVLPPLLTDEMIAQYTSALLA GTITS GWTFGAG AALQIPFAMQM A YRFN GIGVTQNVLYEN QKLIAN QFNS AIG KIQDS LS STAS ALGKLQD V VN QN AQ ALNTL VKQLS S NFG AIS S VLNDILS RLDK VEAEVQIDRLITGRLQSLQTYVTQQLIRAAEIRASANLAATKMSECVLGQSKRV DFCGKGYHLMSFPQSAPHGVVFLHVTYVPAQEKNFTTAPAICHDGKAHFPREG VF V S N GTH WF VTQRNF YEPQIITTDNTFV S GN CD V VIGI VNNT V YDPLQPELDS F KEELDKYFKNHT S PD VDLGDIS GIN AS V VNIQKEIDRLNE V AKNLNES LIDLQEL GKYEQYIKWPWYIWLGFIAGLIAIVMVTIMLCCMTSCCSCLKGCCSCGSCCKFD EDDS EP VLKG VKLH YTMDLFMRIFTIGT VTLKQGEIKD ATPS DF VR AT ATIPIQ A S LPFGWLIV G V ALL A VF QS AS KIITLKKRW QL ALS KG VHFVCNLLLLF VT V Y S H LLLVAAGLEAPFLYLYALVYFLQSINFVRIIMRLWLCWKCRSKNPLLYDANYFL CWHTNCYDYCIPYNSVTSSIVITSGDGTTSPISEHDYQIGGYTEKWESGVKDCV VLHSYFTSDYYQLYSTQLSTDTGVEHVTFFIYNKIVDEPEEHVQIHTIDGSSGVV NP VMEPIYDEPTTTT S VPLM Y S F V S EETGTLIVN S VLLFLAFV VFLL VTL AILT AL RLC A Y CCNIVN V S L VKPS FY V Y S R VKNLN S S RVPDLLVM ADS N GTIT VEELKKL LEQWNLVIGFLFLTWICLLQFAYANRNRFLYIIKLIFLWLLWPVTLACFVLAAV YRINWITGGIAIAMACLVGLMWLSYFIASFRLFARTRSMWSFNPETNILLNVPLH
GTILTRPLLES ELVIG A VILRGHLRIAGHHLGRCDIKDLPKEIT V AT S RTLS Y YKL GASQRVAGDSGFAAYSRYRIGNYKLNTDHSSSSDNIALLVQMFHLVDFQVTIAE ILLIIMRTFKV S IWNLD YIINLIIKNLS KSLTENKY S QLDEEQPMEIDMKIILFLALI TL AT CELYH Y QEC VRGTT VLLKEPCS S GT YEGN S PFHPLADNKFALT CFS T QFA FACPDG VKH V Y QLR ARS V S PKLFIRQEE V QELY S PIFLIV A AIVFITLCFTLKRKT EMIELSLIDFYLCFLAFLLFLVLIMLIIFWFSLELQDHNETCHAMKFLVFLGIITTV AAFHQECSLQSCTQHQPYVVDDPCPIHFYSKWYIRVGARKSAPLIELCVDEAGS KSPIQ YIDIGN YT V S CLPFTIN C QEPKLGS LV VRC S FYEDFLE YHD VR V VLDFIMS DN GPQN QRN APRITF GGPSDSTGSN QN GERS GARS KQRRPQGLPNNT AS WFT A LTQHGKEDLKFPRGQGVPINTNSSPDDQIGYYRRATRRIRGGDGKMKDLSPRW YFYYLGTGPEAGLPYGANKDGIIWVATEGALNTPKDHIGTRNPANNAAIVLQLP QGTTLPKGF Y AEGS RGGS Q AS S RS S S RS RN S S RN S TPGS S RGTS P ARM AGN GGD AALALLLLDRLN QLES KMS GKGQQQQGQT VTKKS AAEAS KKPRQKRTATKA Y N VTQ AF GRRGPEQTQGNF GDQELIRQGTD YKHWPQIAQFAPS AS AFF GMS RIG MEVTPSGTWLTYTGAIKLDDKDPNFKDQVILLNKHIDAYKTFPPTEPKKDKKK KADETQ ALPQRQKKQQT VTLLP A ADLDDF S KQLQQS MSS ADS TQ AMG YIN VF AFPFTI Y S FFFCRMN S RN YI AQ VD V VNFNFT SEQ ID NO: 209: SARS-CoV-2 spike (1273 amino acids)
MFVFF VFFPFV S S QC VNFTTRTQFPP A YTN S FTRG V Y YPDKVFRS S VFHS T QDF FFPFFSNVTWFHAIHVSGTNGTKRFDNPVFPFNDGVYFASTEKSNIIRGWIFGTT FDSKTQSFFIVNNATNVVIKVCEFQFCNDPFFGVYYHKNNKSWMESEFRVYSS ANNCTFEY V S QPFFMDFEGKQGNFKNFREFVFKNIDGYFKIY S KHTPINFVRDF PQGF S AFEPFVDFPIGINITRFQTFF AFHRS YFTPGDS S S GWT AG A A A Y Y V G YFQ PRTFFFKYNENGTITDAVDCAFDPFSETKCTFKSFTVEKGIYQTSNFRVQPTESI VRFPNITNFCPF GE VFN ATRFAS V Y A WNRKRIS NC V AD Y S VF YN S AS FS TFKC Y GVSPTKFNDFCFTNVYADSFVIRGDEVRQIAPGQTGKIADYNYKFPDDFTGCVI AWN S NNFDS KV GGN YN YLYRFFRKS NFKPFERDIS TEIY Q AGS TPCN G VEGFN C YFPFQS Y GFQPTN G V GY QP YRV V VFS FEFFH AP AT V C GPKKS TNFVKNKC VN FNFNGFTGTGVFTESNKKFFPFQQFGRDIADTTDAVRDPQTFEIFDITPCSFGGV S VITPGTNTS N Q V A VF Y QD VNCTE VP V AIH ADQFTPT WRV Y S TGS N VF QTR AG CFIG AEH VNN S YECDIPIG AGIC AS Y QTQTN S PRR ARS V AS QS IIA YTMS EG AEN S VAYSNNSIAIPTNFTISVTTEIFPVSMTKTSVDCTMYICGDSTECSNFFFQYGSFC TQFNRAFTGIAVEQDKNTQEVFAQVKQIYKTPPIKDFGGFNFSQIFPDPSKPSKR SFIEDFFFNKVTLAD AGFIKQY GDCFGDIAARDFIC AQKFN GLT VLPPLLTDEMI AQYTS ALLAGTITS GWTFG AGAALQIPFAMQMA YRFN GIGVTQN VLYEN QKLI AN QFN S AIGKIQDS LS STAS ALGKLQD V VN QN AQ ALNTL VKQLS S NFG AIS S VL NDILSRLDKVEAEVQIDRLITGRLQSLQTYVTQQLIRAAEIRASANLAATKMSEC VLGQSKRVDFCGKGYHLMSFPQSAPHGVVFLHVTYVPAQEKNFTTAPAICHDG KAHFPREGVFVSNGTHWFVTQRNFYEPQIITTDNTFV S GNCD VVIGIVNNT VYD
PLQPELDS FKEELDKYFKNHT S PD VDLGDIS GINAS V VNIQKEIDRLNE V AKNLN ESLIDLQELGKYEQYIKWPWYIWLGFIAGLIAIVMVTIMLCCMTSCCSCLKGCC SC GS CC KFDEDDS EP VLKG VKLH YT
REFERENCES
Ghebrehiwet B, Jesty J, Vinayagasundaram R, Vinayagasundaram U, Ji Y, Valentino A, Tumma N, Hosszu KH, Peerschke EL Targeting gC1qR domains for therapy against infection and inflammation. Adv Exp Med Biol. 2013;735:97-110.
Pednekar L, Valentino A, Ji Y, Tumma N, Valentino C, Kadoor A, Hosszu KK, Ramadass M, Kew RR, Kishore U, Peerschke El, Ghebrehiwet B . Identification of the gC1qR sites for the HIV-1 viral envelope protein gp41 and the HCV core protein: Implications in viral- specific pathogenesis and therapy. Mol Immunol. 2016 Jun;74: 18-26.
Rossio JL, Esser MT, Suryanarayana K, Schneider DK, Bess JW Jr, Vasquez GM, Wiltrout TA, Chertova E, Grimes MK, Sattentau Q, Arthur LO, Henderson LE, Lifson JD. Inactivation of human immunodeficiency virus type 1 infectivity with preservation of conformational and functional integrity of virion surface proteins. J Virol. 1998 0ct;72(10):7992-8001.
Platt EJ, Wehrly K, Kuhmann SE, Chesebro B, Rabat D. Effects of CCR5 and CD4 cell surface concentrations on infections by macrophagetropic isolates of human immunodeficiency virus type 1. J Virol. 1998 Apr;72(4):2855-64.
Vieillard V, Strominger JL, Debre P. NK cytotoxicity against CD4+ T cells during HIV- 1 infection: A gp41 peptide induces expression of an NKp44 ligand. PNAS 2005 Aug 2; 102(31): 10981-6.
Fausther-Bovendo H, Vieillard V, Sagan S, Bismuth G, Debre P. HIV gp41 engages gC1qR on CD4+ T cells to induce the expression of an NK ligand through the PIP3/H202 pathway. PLoS Pathog. 2010 Jul l;6:el000975.
Kittlesen DJ, Chianese-Bullock KA, Yao ZQ, Braciale TJ, Hahn YS. Interaction between complement receptor gC1qR and hepatitis C virus core protein inhibits T-lymphocyte proliferation. J Clin Invest. 2000 Nov; 106(10): 1239-49.
Large MK, Kittlesen DJ, Hahn YS. Suppression of host immune response by the core protein of hepatitis C virus: possible implications for hepatitis C virus persistence. J Immunol. 1999 Jan 15; 162(2):931-8.
Walls AC, Park YJ, Tortorici MA, Wall A, McGuire AT, Veesler D. Structure, Function, and Antigenicity of the SARSCoV-2 Spike Glycoprotein. Cell. 2020 Apr 16; 181(2):281- 292. e6.
Lan J, Ge J, Yu J, Shan S, Zhou H, Fan S, Zhang Q, Shi X, Wang Q, Zhang L, Wang X.
Structure of the SARS-CoV-2 spike receptor-binding domain bound to the ACE2 receptor. Nature. 2020 Mar 30.
Ou X, Liu Y, Lei X, Li P, Mi D, Ren L, Guo L, Guo R, Chen T, Hu J, Xiang Z, Mu Z, Chen X, Chen J, Hu K, Jin Q, Wang J, Qian Z. Characterization of spike glycoprotein of SARS-CoV-2 on virus entry and its immune cross -reactivity with SARS-CoV. Nat Commun. 2020 Mar 27; 11(1): 1620.
Wrapp D, Wang N, Corbett KS, Goldsmith JA, Hsieh CL, Abiona O, Graham BS, McLellan JS. Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science. 2020 Mar 13;367(6483): 1260-1263.
Claims
1. A compound for use for treating a Group IV RNA virus infection, particularly a coronavirus infection, preferably a SARS-Cov-2 infection, wherein the compound inhibits the sPLA2GIB pathway.
2. The compound for use according to claim 1, wherein the compound is an inhibitor of SPLA2GIB.
3. The compound for use according to claim 2, wherein the compound is selected from: a. an anti-sPLA2GlB antibody or a fragment or derivative thereof which binds PLA2- GIB, preferably a monoclonal antibody, more preferably human or humanized; b. a nucleic acid encoding an antibody of a., c. a soluble receptor, d. an inhibitory nucleic acid, or e. a peptide or small drug.
4. The compound for use according to claim 1, wherein the compound comprises an antigen which can induce an anti-sPLA2GIB immune response in vivo.
5. The compound for use according to claim 1, wherein the compound is an inhibitor of a cofactor of sPLA2-GIB.
6. The compound for use according to claim 5, wherein the PLA2-GIB cofactor is a coronavirus protein selected from proteins ORFlab, Spike, E, M, N, Orf7b, and ORF8, or a molecule comprising a gC1qR-binding element of any one of said proteins.
7. The compound for use according to claim 6, wherein the cofactor is a spike protein of the coronavirus, or a molecule comprising a gC1qR-binding element of a spike virus of the coronavirus.
8. The compound for use according to anyone of claims 5 to 7, wherein the inhibitor inhibits binding of the cofactor to gC1qR, or inhibits expression of the cofactor.
9. The compound for use according to any one of claims 5 to 8, wherein the inhibitor is an antibody or a variant or fragment of an antibody, preferably which binds gC1qR or which binds a protein selected from ORFlab, Spike, E, M, N, Orf7b, and ORF8 and preferably inhibits binding of said protein to gC1qR.
10. The compound for use according to any one of claims 5 to 8, wherein the inhibitor is a peptide or lipopeptide, preferably a peptide which binds gC1qR and inhibits binding to
gC1qR of a protein selected from ORFlab, Spike, E, M, N, Orf7b, and ORF8, more preferably a peptide comprising anyone of SEQ ID Nos: 2-208.
11. The compound for use according to any one of claims 5 to 8, wherein the inhibitor is a nucleic acid or a carbohydrate.
12. The compound for use according to any one of claims 5 to 8, wherein the inhibitor is a recombinant vims expressing a peptide comprising anyone of SEQ ID Nos: 2-208, preferably a virus with a pseudotyped envelope comprising a peptide comprising anyone of SEQ ID Nos: 2-208.
13. The compound for use according to any one of claims 5 to 8, wherein the inhibitor comprises an antigen which can induce an immune response in vivo against the PLA2- GIB cofactor.
14. The compound for use according to any one of the preceding claims, for treating a SARS-Cov-2 infection in a mammal, preferably a human.
15. The compound for use according to anyone of the preceding claims, wherein the compound is administered in combination with another drug or treatment.
16. A peptide of less than about 50 amino acids which comprises a sequence selected from anyone of SEQ ID Nos: 2-208 or an immunogenic fragment thereof.
17. The peptide of claim 16, which is conjugated to an entity, preferably a surface, device, drug, carrier, protein, virus, or adjuvant.
18. A nucleic acid molecule encoding a peptide of claim 16.
19. A compound which inhibits binding of Group IV RNA vims, particularly a coronavirus protein to gC1qR, preferably of a coronavirus Spike protein to gC1qR.
20. The compound of claim 19, which is an antibody or a variant or fragment of an antibody.
21. An antibody which binds a SARS-Cov-2 spike protein, wherein said antibody binds an epitope comprising at least one amino acid residue located in the NTD domain (aa 16- 305, by reference to SEQ ID NO: 209) or in the RBD domain (aa 331-527, by reference to SEQ ID NO: 209) of the spike protein.
22. An antibody which binds a SARS-Cov-2 spike protein, wherein said antibody binds an epitope comprising at least one amino acid residue of the spike protein located in anyone of SEQ ID NOs: 156-185.
23. An antibody which binds a SARS-Cov-2 spike protein, wherein said antibody binds an epitope comprising at least one of the following amino acid residues of the spike protein (by reference to SEQ ID NO: 209): 99-113; 147-161; 253-267; 348-445, 348-362, 431-445.
24. An antibody which binds a SARS-Cov-2 spike protein, wherein said antibody binds an epitope comprising at least one of the following amino acid residue of the spike protein (by reference to SEQ ID NO: 209):: 431, 432, 433, 434, 435, 436, 437, 438, 438, 440, 441, 442, 443, 444, 445.
25. A composition comprising an antibody or a compound of anyone of claims 19-24.
26. A vaccine comprising a peptide of claim 16 or 17 and, optionally, an adjuvant.
27. A vaccine comprising, as immunogen, a virus envelope protein, particularly a Group IV RNA virus, particularly with a modified gC1qR binding motif, said protein retaining immunogenicity while having no or less PLA2-GIB cofactor effect.
28. A vaccine of claim 26, wherein the protein has a deleted and/or mutated gC1qR binding motif.
29. The vaccine of claim 27 or 28, comprising a SARS-Cov-2 spike protein having, by reference to SEQ ID NO: 209, at least one amino acid modification in a residue selected from anyone of SEQ ID NOs: 156-185, even more preferably in a residue selected from aa 348-362 or 431-445 (by reference to SEQ ID NO: 209).
30. A protein comprising the amino acid sequence of SEQ ID NO: 209, or a natural variant thereof, wherein the sequence contains at least one amino acid substitution or deletion in a position selected from: aa99-113, aal47-161, aa253-267, aa348-362, aa431- 445.
31. The protein of claim 30, wherein the sequence contains at least one deletion of anyone said residues, even more preferably of at least 2, 3, 4, 5, 6, 7, 8, 9, 10, or more consecutive aa residues at said positions.
32. A protein comprising SEQ ID NO: 209 with a deletion spanning all or part of AA431- 445.
33. A protein comprising SEQ ID NO: 209 with a deletion spanning all or part of AA348- 362.
34. A vaccine comprising a protein of anyone of claims 30-33.
35. A method of detecting SARS-Cov-2 in a sample, comprising detecting the presence in said sample of antibodies which bind a peptide of claim 16.
36. A method of making inhibitors, comprising (i) providing a SARS-Cov-19 binding agent and (ii) determining if said agent binds a region comprising at least one of the following amino acids (by reference to SEQ ID NO: 209): 99-113; 147-161; 253-267; 348-445, 348-362, 431-445.
37. A combination therapy or therapeutic regimen for treating a disorder caused by a Group IV RNA vims infection, particularly a coronavirus infection, comprising a combination of at least (i) an inhibitor of the sPLA2-GIB pathway, and (ii) another active agent, said agents being for combined, separate or sequential administration.
Applications Claiming Priority (4)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| EP20305547.0 | 2020-05-26 | ||
| EP20305547 | 2020-05-26 | ||
| US202063044428P | 2020-06-26 | 2020-06-26 | |
| US63/044,428 | 2020-06-26 |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| WO2021239666A1 true WO2021239666A1 (en) | 2021-12-02 |
Family
ID=75977765
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/EP2021/063764 Ceased WO2021239666A1 (en) | 2020-05-26 | 2021-05-24 | Therapeutic methods |
Country Status (1)
| Country | Link |
|---|---|
| WO (1) | WO2021239666A1 (en) |
Cited By (1)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO2023104154A1 (en) * | 2021-12-08 | 2023-06-15 | 安达生物药物开发(深圳)有限公司 | Antigenic polypeptide and use thereof |
Citations (33)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US4376110A (en) | 1980-08-04 | 1983-03-08 | Hybritech, Incorporated | Immunometric assays using monoclonal antibodies |
| US4411993A (en) | 1981-04-29 | 1983-10-25 | Steven Gillis | Hybridoma antibody which inhibits interleukin 2 activity |
| EP0125023A1 (en) | 1983-04-08 | 1984-11-14 | Genentech, Inc. | Recombinant immunoglobulin preparations, methods for their preparation, DNA sequences, expression vectors and recombinant host cells therefor |
| US4486530A (en) | 1980-08-04 | 1984-12-04 | Hybritech Incorporated | Immunometric assays using monoclonal antibodies |
| US4543439A (en) | 1982-12-13 | 1985-09-24 | Massachusetts Institute Of Technology | Production and use of monoclonal antibodies to phosphotyrosine-containing proteins |
| USRE32011E (en) | 1981-12-14 | 1985-10-22 | Scripps Clinic And Research Foundation | Ultrapurification of factor VIII using monoclonal antibodies |
| EP0171496A2 (en) | 1984-08-15 | 1986-02-19 | Research Development Corporation of Japan | Process for the production of a chimera monoclonal antibody |
| EP0173494A2 (en) | 1984-08-27 | 1986-03-05 | The Board Of Trustees Of The Leland Stanford Junior University | Chimeric receptors by DNA splicing and expression |
| WO1986001533A1 (en) | 1984-09-03 | 1986-03-13 | Celltech Limited | Production of chimeric antibodies |
| EP0184187A2 (en) | 1984-12-04 | 1986-06-11 | Teijin Limited | Mouse-human chimaeric immunoglobulin heavy chain, and chimaeric DNA encoding it |
| WO1987002671A1 (en) | 1985-11-01 | 1987-05-07 | International Genetic Engineering, Inc. | Modular assembly of antibody genes, antibodies prepared thereby and use |
| US4902614A (en) | 1984-12-03 | 1990-02-20 | Teijin Limited | Monoclonal antibody to human protein C |
| WO1990002809A1 (en) | 1988-09-02 | 1990-03-22 | Protein Engineering Corporation | Generation and selection of recombinant varied binding proteins |
| WO1991017271A1 (en) | 1990-05-01 | 1991-11-14 | Affymax Technologies N.V. | Recombinant library screening methods |
| WO1992001047A1 (en) | 1990-07-10 | 1992-01-23 | Cambridge Antibody Technology Limited | Methods for producing members of specific binding pairs |
| WO1992009690A2 (en) | 1990-12-03 | 1992-06-11 | Genentech, Inc. | Enrichment method for variant proteins with altered binding properties |
| WO1992015679A1 (en) | 1991-03-01 | 1992-09-17 | Protein Engineering Corporation | Improved epitode displaying phage |
| WO1992018619A1 (en) | 1991-04-10 | 1992-10-29 | The Scripps Research Institute | Heterodimeric receptor libraries using phagemids |
| WO1992020791A1 (en) | 1990-07-10 | 1992-11-26 | Cambridge Antibody Technology Limited | Methods for producing members of specific binding pairs |
| WO1993001288A1 (en) | 1991-07-08 | 1993-01-21 | Deutsches Krebsforschungszentrum Stiftung des öffentlichen Rechts | Phagemide for screening antibodies |
| US5223409A (en) | 1988-09-02 | 1993-06-29 | Protein Engineering Corp. | Directed evolution of novel binding proteins |
| US5225539A (en) | 1986-03-27 | 1993-07-06 | Medical Research Council | Recombinant altered antibodies and methods of making altered antibodies |
| GB2272440A (en) | 1990-08-29 | 1994-05-18 | Genpharm Int | Transgenic non-human animals capable of producing heterologous antibodies |
| WO1996007754A1 (en) | 1994-09-02 | 1996-03-14 | The Scripps Research Institute | Methods for producing antibody libraries using universal or randomized immunoglobulin light chains |
| US5545806A (en) | 1990-08-29 | 1996-08-13 | Genpharm International, Inc. | Ransgenic non-human animals for producing heterologous antibodies |
| WO2007044695A2 (en) * | 2005-10-07 | 2007-04-19 | Dana-Farber Cancer Institute | ANTIBODIES AGAINST SARS-CoV AND METHODS OF USE THEREOF |
| US7521541B2 (en) | 2004-09-23 | 2009-04-21 | Genetech Inc. | Cysteine engineered antibodies and conjugates |
| WO2015097140A1 (en) | 2013-12-23 | 2015-07-02 | Institut Pasteur | Therapeutic methods and compositions |
| WO2017037041A1 (en) | 2015-08-31 | 2017-03-09 | Diaccurate | Use of indole compounds to stimulate the immune system |
| WO2017060405A1 (en) | 2015-10-09 | 2017-04-13 | Diaccurate | Use of peptides to stimulate the immune system |
| EP3530282A1 (en) * | 2018-02-27 | 2019-08-28 | Diaccurate | Therapeutic methods |
| WO2019166665A1 (en) | 2018-03-02 | 2019-09-06 | Diaccurate | Therapeutic anti-spla2-gib antibodies and the uses thereof |
| CN111088283A (en) * | 2020-03-20 | 2020-05-01 | 苏州奥特铭医药科技有限公司 | mVSV viral vector, viral vector vaccine thereof and mVSV-mediated novel coronary pneumonia vaccine |
-
2021
- 2021-05-24 WO PCT/EP2021/063764 patent/WO2021239666A1/en not_active Ceased
Patent Citations (36)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US4486530A (en) | 1980-08-04 | 1984-12-04 | Hybritech Incorporated | Immunometric assays using monoclonal antibodies |
| US4376110A (en) | 1980-08-04 | 1983-03-08 | Hybritech, Incorporated | Immunometric assays using monoclonal antibodies |
| US4411993A (en) | 1981-04-29 | 1983-10-25 | Steven Gillis | Hybridoma antibody which inhibits interleukin 2 activity |
| USRE32011E (en) | 1981-12-14 | 1985-10-22 | Scripps Clinic And Research Foundation | Ultrapurification of factor VIII using monoclonal antibodies |
| US4543439A (en) | 1982-12-13 | 1985-09-24 | Massachusetts Institute Of Technology | Production and use of monoclonal antibodies to phosphotyrosine-containing proteins |
| US4816567A (en) | 1983-04-08 | 1989-03-28 | Genentech, Inc. | Recombinant immunoglobin preparations |
| EP0125023A1 (en) | 1983-04-08 | 1984-11-14 | Genentech, Inc. | Recombinant immunoglobulin preparations, methods for their preparation, DNA sequences, expression vectors and recombinant host cells therefor |
| EP0171496A2 (en) | 1984-08-15 | 1986-02-19 | Research Development Corporation of Japan | Process for the production of a chimera monoclonal antibody |
| EP0173494A2 (en) | 1984-08-27 | 1986-03-05 | The Board Of Trustees Of The Leland Stanford Junior University | Chimeric receptors by DNA splicing and expression |
| WO1986001533A1 (en) | 1984-09-03 | 1986-03-13 | Celltech Limited | Production of chimeric antibodies |
| US4902614A (en) | 1984-12-03 | 1990-02-20 | Teijin Limited | Monoclonal antibody to human protein C |
| EP0184187A2 (en) | 1984-12-04 | 1986-06-11 | Teijin Limited | Mouse-human chimaeric immunoglobulin heavy chain, and chimaeric DNA encoding it |
| WO1987002671A1 (en) | 1985-11-01 | 1987-05-07 | International Genetic Engineering, Inc. | Modular assembly of antibody genes, antibodies prepared thereby and use |
| US5225539A (en) | 1986-03-27 | 1993-07-06 | Medical Research Council | Recombinant altered antibodies and methods of making altered antibodies |
| WO1990002809A1 (en) | 1988-09-02 | 1990-03-22 | Protein Engineering Corporation | Generation and selection of recombinant varied binding proteins |
| US5223409A (en) | 1988-09-02 | 1993-06-29 | Protein Engineering Corp. | Directed evolution of novel binding proteins |
| WO1991017271A1 (en) | 1990-05-01 | 1991-11-14 | Affymax Technologies N.V. | Recombinant library screening methods |
| WO1992020791A1 (en) | 1990-07-10 | 1992-11-26 | Cambridge Antibody Technology Limited | Methods for producing members of specific binding pairs |
| WO1992001047A1 (en) | 1990-07-10 | 1992-01-23 | Cambridge Antibody Technology Limited | Methods for producing members of specific binding pairs |
| US5569825A (en) | 1990-08-29 | 1996-10-29 | Genpharm International | Transgenic non-human animals capable of producing heterologous antibodies of various isotypes |
| US5545806A (en) | 1990-08-29 | 1996-08-13 | Genpharm International, Inc. | Ransgenic non-human animals for producing heterologous antibodies |
| GB2272440A (en) | 1990-08-29 | 1994-05-18 | Genpharm Int | Transgenic non-human animals capable of producing heterologous antibodies |
| WO1992009690A2 (en) | 1990-12-03 | 1992-06-11 | Genentech, Inc. | Enrichment method for variant proteins with altered binding properties |
| WO1992015679A1 (en) | 1991-03-01 | 1992-09-17 | Protein Engineering Corporation | Improved epitode displaying phage |
| WO1992018619A1 (en) | 1991-04-10 | 1992-10-29 | The Scripps Research Institute | Heterodimeric receptor libraries using phagemids |
| WO1993001288A1 (en) | 1991-07-08 | 1993-01-21 | Deutsches Krebsforschungszentrum Stiftung des öffentlichen Rechts | Phagemide for screening antibodies |
| WO1996007754A1 (en) | 1994-09-02 | 1996-03-14 | The Scripps Research Institute | Methods for producing antibody libraries using universal or randomized immunoglobulin light chains |
| US7521541B2 (en) | 2004-09-23 | 2009-04-21 | Genetech Inc. | Cysteine engineered antibodies and conjugates |
| WO2007044695A2 (en) * | 2005-10-07 | 2007-04-19 | Dana-Farber Cancer Institute | ANTIBODIES AGAINST SARS-CoV AND METHODS OF USE THEREOF |
| WO2015097140A1 (en) | 2013-12-23 | 2015-07-02 | Institut Pasteur | Therapeutic methods and compositions |
| WO2017037041A1 (en) | 2015-08-31 | 2017-03-09 | Diaccurate | Use of indole compounds to stimulate the immune system |
| WO2017060405A1 (en) | 2015-10-09 | 2017-04-13 | Diaccurate | Use of peptides to stimulate the immune system |
| EP3530282A1 (en) * | 2018-02-27 | 2019-08-28 | Diaccurate | Therapeutic methods |
| WO2019166412A1 (en) | 2018-02-27 | 2019-09-06 | Diaccurate | Modulation of pla2-g1b in therapy |
| WO2019166665A1 (en) | 2018-03-02 | 2019-09-06 | Diaccurate | Therapeutic anti-spla2-gib antibodies and the uses thereof |
| CN111088283A (en) * | 2020-03-20 | 2020-05-01 | 苏州奥特铭医药科技有限公司 | mVSV viral vector, viral vector vaccine thereof and mVSV-mediated novel coronary pneumonia vaccine |
Non-Patent Citations (67)
| Title |
|---|
| "GenBank", Database accession no. QHD43416.1 |
| "Genbank", Database accession no. YP_009724390.1 |
| "Monoclonal Antibodies, Hybridomas: A New Dimension in Biological Analyses", 1980, PLENUM PRESS |
| "Swiss-Prot", Database accession no. Q07021.1 |
| "The Merck Manual", 1992, MERCK AND CO. |
| ALTING-MEES ET AL.: "Monoclonal Antibody Expression Libraries: A Rapid Alternative to Hybridomas", STRATEGIES IN MOLECULAR BIOLOGY, vol. 3, 1990, pages 1 - 9, XP001538956 |
| ALTSCHUL ET AL., NUCLEIC ACIDS RES, vol. 25, 1997, pages 3389 - 3402 |
| BARBAS ET AL., PNAS, vol. 88, 1991, pages 7978 - 7982 |
| BARBAS ET AL., PROC. NAT'L ACAD. SCI. USA, vol. 89, 1992, pages 4457 - 4461 |
| BEIDLER ET AL., J. IMMUNOL., vol. 141, 1988, pages 4053 - 4060 |
| CHARI ET AL., CANCER RES., vol. 52, 1992, pages 127 - 131 |
| CLACKSON ET AL., NATURE, vol. 352, 1991, pages 624 - 628 |
| E. KABAT ET AL.: "Sequences of Proteins of Immunological Interest", 1983, U.S. DEPARTMENT OF HEALTH AND HUMAN SERVICES |
| FAUSTHER-BOVENDO HVIEILLARD VSAGAN SBISMUTH GDEBRE P: "HIV gp41 engages gClqR on CD4+ T cells to induce the expression of an NK ligand through the PIP3/H202 pathway", PLOS PATHOG, vol. 6, 1 July 2010 (2010-07-01), pages el000975 |
| FLATMAN ET AL., J. CHROMATOGR. B, vol. 848, 2007, pages 79 - 87 |
| GARRAD ET AL., BIO/TECHNOLOGY, vol. 9, 1991, pages 1373 - 1377 |
| GHEBREHIWET BJESTY JVINAYAGASUNDARAM RVINAYAGASUNDARAM UJI YVALENTINO ATUMMA NHOSSZU KHPEERSCHKE EI: "Targeting gClqR domains for therapy against infection and inflammation", ADV EXP MED BIOL, vol. 735, 2013, pages 97 - 110 |
| GRAM ET AL., PNAS, vol. 89, 1992, pages 3576 - 3580 |
| GRIFFTHS ET AL., EMBO J, vol. 12, 1993, pages 725 - 734 |
| HAWKINS ET AL., J MOL BIOL, vol. 226, 1992, pages 889 - 896 |
| HAY ET AL., HUM ANTIBOD HYBRIDOMAS, vol. 3, 1992, pages 81 - 85 |
| HOOGENBOOM ET AL., NUC ACID RES, vol. 19, 1991, pages 4133 - 4137 |
| HUSE ET AL., SCIENCE, vol. 246, 1989, pages 1275 - 1281 |
| J. V. KRINGELUMC. LUNDEGAARDO. LUNDM. NIELSEN: "Reliable B cell epitope predictions: impacts of method development and improved benchmarking", PLOS COMPUT BIOL, vol. 8, no. 12, 2012, pages e1002829, XP055113333, DOI: 10.1371/journal.pcbi.1002829 |
| JAKOBOVITS ET AL., ANN NY ACAD SCI, vol. 764, 1995, pages 525 - 535 |
| JENS VINDAHL KRINGELUMCLAUS LUNDEGAARDOLE LUNDMORTEN NIELSEN: "Reliable B cell epitope predictions: Impacts of method development and improved benchmarking", PLOS COMPUTATIONAL BIOLOGY, 2012 |
| JENSEN KKANDREATTA MMARCATILI PBUUS SGREENBAUM JAYAN ZSETTE APETERS BNIELSEN M: "Improved methods for predicting peptide binding affinity to MHC class II molecules", IMMUNOLOGY, vol. 154, no. 3, July 2018 (2018-07-01), pages 394 - 406, XP055715184, DOI: 10.1111/imm.12889 |
| JONES ET AL., NATURE, vol. 321, 1986, pages 552 |
| KABSCH WSANDER C: "Dictionary of protein secondary structure: pattern recognition of hydrogen-bonded and geometrical features", BIOPOLYMERS, vol. 22, no. 12, December 1983 (1983-12-01), pages 2577 - 637, XP055076173, DOI: 10.1002/bip.360221211 |
| KANDA, Y. ET AL., BIOTECHNOL. BIOENG., vol. 94, no. 4, 2006, pages 680 - 688 |
| KITTLESEN DJCHIANESE-BULLOCK KAYAO ZQBRACIALE TJHAHN YS: "Interaction between complement receptor gClqR and hepatitis C virus core protein inhibits T-lymphocyte proliferation", J CLIN INVEST, vol. 106, no. 10, November 2000 (2000-11-01), pages 1239 - 49, XP002685556, DOI: 10.1172/JCI10323 |
| LAN J, GE J, YU J, SHAN S, ZHOU H, FAN S, ZHANG Q, SHI X, WANG Q, ZHANG L, WANG X: "Structure of the SARS-CoV-2 spike receptor-binding domain bound to the ACE2 receptor", NATURE, 30 March 2020 (2020-03-30) |
| LARGE MKKITTLESEN DJHAHN YS: "Suppression of host immune response by the core protein of hepatitis C virus: possible implications for hepatitis C virus persistence", J IMMUNOL., vol. 162, no. 2, 15 January 1999 (1999-01-15), pages 931 - 8 |
| LARRICK ET AL., BIO/TECHNOLOGY, vol. 7, 1989, pages 934 |
| LARRICK ET AL., BIOTECHNOLOGY, vol. 7, 1989, pages 394 |
| LIU ET AL., J. IMMUNOL., vol. 139, 1987, pages 3521 - 3526 |
| MARKS ET AL., J. MOL. BIOL., vol. 222, 1991, pages 581 - 597 |
| MORRISON, S. L., SCIENCE, vol. 229, 1985, pages 1202 - 1207 |
| NISHIMURA ET AL., CANC. RES., vol. 47, 1987, pages 999 - 1005 |
| OI ET AL., BIOTECHNIQUES, vol. 4, 1986, pages 214 |
| OKAZAKI ET AL., J. MOL. BIOL., vol. 336, 2004, pages 1239 - 1249 |
| OU XLIU YLEI XLI PMI DREN LGUO LGUO RCHEN THU J: "Characterization of spike glycoprotein of SARS-CoV-2 on virus entry and its immune cross-reactivity with SARS-CoV", NAT COMMUN, vol. 11, no. 1, 27 March 2020 (2020-03-27), pages 1620 |
| PEDNEKAR ET AL., MOL IMMUNOL, 2016 |
| PEDNEKAR LINA ET AL: "Identification of the gC1qR sites for the HIV-1 viral envelope protein gp41 and the HCV core protein: Implications in viral-specific pathogenesis and therapy", MOLECULAR IMMUNOLOGY, PERGAMON, GB, vol. 74, 23 April 2016 (2016-04-23), pages 18 - 26, XP029568768, ISSN: 0161-5890, DOI: 10.1016/J.MOLIMM.2016.03.016 * |
| PEDNEKAR LVALENTINO AJI YTUMMA NVALENTINO CKADOOR AHOSSZU KKRAMADASS MKEW RRKISHORE U: "Identification of the gClqR sites for the HIV-1 viral envelope protein gp41 and the HCV core protein: Implications in viral-specific pathogenesis and therapy", MOL IMMUNOL, vol. 74, June 2016 (2016-06-01), pages 18 - 26, XP029568768, DOI: 10.1016/j.molimm.2016.03.016 |
| PLATT EJWEHRLY KKUHMANN SECHESEBRO BKABAT D: "Effects of CCR5 and CD4 cell surface concentrations on infections by macrophagetropic isolates of human immunodeficiency virus type 1", J VIROL, vol. 72, no. 4, April 1998 (1998-04-01), pages 2855 - 64 |
| POTHLICHET JULIEN ET AL: "PLA2G1B is involved in CD4 anergy and CD4 lymphopenia in HIV-infected patients", THE JOURNAL OF CLINICAL INVESTIGATION, vol. 130, no. 6, 27 April 2020 (2020-04-27), GB, pages 2872 - 2887, XP055774578, ISSN: 0021-9738, DOI: 10.1172/JCI131842 * |
| RIECHMANN ET AL., NATURE, vol. 332, 1988, pages 323 |
| RIPKA ET AL., ARCH. BIOCHEM. BIOPHYS., vol. 249, 1986, pages 533 - 545 |
| ROSSIO JLESSER MTSURYANARAYANA KSCHNEIDER DKBESS JW JRVASQUEZ GMWILTROUT TACHERTOVA EGRIMES MKSATTENTAU Q: "Inactivation of human immunodeficiency virus type 1 infectivity with preservation of conformational and functional integrity of virion surface proteins", J VIROL, vol. 72, no. 10, October 1998 (1998-10-01), pages 7992 - 8001, XP002120813 |
| SAMBROOK JRUSSELL D: "Goodman and Cilman's The pharmacological Basis of Therapeutics", 2001, COLD SPRING HARBOR |
| SCATCHARD ET AL., ANN. N.Y. ACAD. SCI., vol. 51, 1949, pages 660 |
| SHAW ET AL., J. NATL. CANCER INST., vol. 80, 1988, pages 1553 - 1559 |
| SMITHWATERMAN, J MOL BIOL, vol. 147, 1981, pages 195 - 197 |
| SUN ET AL., PNAS, vol. 84, 1987, pages 3439 - 3443 |
| THOMPSON ET AL., NUCLEIC ACIDS RES, vol. 22, 1994, pages 4673 - 4680 |
| VERHOEYAN ET AL., SCIENCE, vol. 239, 1988, pages 1534 - 1043 |
| VIEILLARD VSTROMINGER JLDEBRE P: "NK cytotoxicity against CD4+ T cells during HIV-1 infection: A gp41 peptide induces expression of an NKp44 ligand", PNAS, vol. 102, no. 31, 2 August 2005 (2005-08-02), pages 10981 - 6, XP002569191, DOI: 10.1073/pnas.0504315102 |
| WALLS ACPARK YJTORTORICI MAWALL AMCGUIRE ATVEESLER D: "Structure, Function, and Antigenicity of the SARSCoV-2 Spike Glycoprotein", CELL, vol. 181, no. 2, 16 April 2020 (2020-04-16), pages 281 - 292 |
| WINTERHARRIS, TIPS, vol. 14, May 1993 (1993-05-01), pages 139 |
| WOOD ET AL., NATURE, vol. 314, 1985, pages 446 - 449 |
| WRAPP DWANG NCORBETT KSGOLDSMITH JAHSIEH CLABIONA OGRAHAM BSMCLELLAN JS: "Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation", SCIENCE, vol. 367, no. 6483, 13 March 2020 (2020-03-13), pages 1260 - 1263 |
| WRIGHT ET AL., TIBTECH, vol. 15, 1997, pages 26 - 32 |
| XIA XUHUA: "Position Weight Matrix, Gibbs Sampler, and the Associated Significance Tests in Motif Characterization and Prediction", SCIENTIFICA, vol. 2012, 1 January 2012 (2012-01-01), US, pages 1 - 15, XP055774650, ISSN: 2090-908X, DOI: 10.6064/2012/917540 * |
| YAMANE-OHNUKI ET AL., BIOTECH. BIOENG., vol. 87, 2004, pages 614 |
| YAMANE-OHNUKI NSATOH M, MABS, vol. 1, 2009, pages 230 - 236 |
| ZHOU ET AL., NATURE, 2020 |
Cited By (1)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO2023104154A1 (en) * | 2021-12-08 | 2023-06-15 | 安达生物药物开发(深圳)有限公司 | Antigenic polypeptide and use thereof |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| JP7171809B2 (en) | Antibodies that potently neutralize hepatitis B virus and uses thereof | |
| JP7375062B2 (en) | Antibodies, uses, and methods | |
| US20230038788A1 (en) | Therapeutic methods and compositions | |
| EP3134111B1 (en) | Middle east respiratory syndrome coronavirus neutralizing antibodies and methods of use thereof | |
| JP5814790B2 (en) | Dengue virus neutralizing antibody and use thereof | |
| KR101407216B1 (en) | Human cytomegalovirus neutralising antibodies and use thereof | |
| US8298545B2 (en) | Anti-autoimmune antibodies for treatment of pemphigus | |
| US11078285B2 (en) | Anti-TEM1 antibodies and uses thereof | |
| SA96170384B1 (en) | Methods of preparation and therapeutic uses (RSV) Human neutralizing monoclonal antibodies with high affinity for the F-protein of respiratory syncytial virus | |
| US20200399392A1 (en) | Inhibitors of pla2-g1b cofactors for treating cancer | |
| WO2021222988A1 (en) | Cell entry-modulating agents and uses therefor | |
| US12503515B2 (en) | Humanized 4-1BB monoclonal antibody and pharmaceutical composition thereof | |
| CN108779168A (en) | HIV bonding agents | |
| WO2021239666A1 (en) | Therapeutic methods | |
| WO2022132887A1 (en) | Human monoclonal antibodies targeting the sars-cov-2 spike protein | |
| EP4328236A1 (en) | Glycosylated rbd and use thereof | |
| US20250304659A1 (en) | HUMAN MONOCLONAL ANTIBODIES TO SEVERE ACUTE RESPIRATORY SYNDROME CORONAVIRUS 1 (SARS-CoV-1) | |
| US12331133B2 (en) | Therapeutic antibodies for treating lung cancer | |
| WO2024026420A1 (en) | Methods and systems for identifying or treating central nervous systems disease | |
| WO2023145874A1 (en) | Novel monoclonal antibody against corona virus | |
| CN116744966A (en) | DNA-encoded antibodies against SARS-COV-2 | |
| HK40073500A (en) | Humanized 4-1bb monoclonal antibody and pharmaceutical composition thereof |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| 121 | Ep: the epo has been informed by wipo that ep was designated in this application |
Ref document number: 21726427 Country of ref document: EP Kind code of ref document: A1 |
|
| NENP | Non-entry into the national phase |
Ref country code: DE |
|
| 122 | Ep: pct application non-entry in european phase |
Ref document number: 21726427 Country of ref document: EP Kind code of ref document: A1 |