WO2014186686A2 - Targeted mutagenesis and genome engineering in plants using rna-guided cas nucleases - Google Patents
Targeted mutagenesis and genome engineering in plants using rna-guided cas nucleases Download PDFInfo
- Publication number
- WO2014186686A2 WO2014186686A2 PCT/US2014/038359 US2014038359W WO2014186686A2 WO 2014186686 A2 WO2014186686 A2 WO 2014186686A2 US 2014038359 W US2014038359 W US 2014038359W WO 2014186686 A2 WO2014186686 A2 WO 2014186686A2
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- nucleic acid
- plant
- acid sequence
- plant cell
- promoter
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Ceased
Links
- KUHNPWUQIFZACX-UHFFFAOYSA-N CC[F]C1CC1 Chemical compound CC[F]C1CC1 KUHNPWUQIFZACX-UHFFFAOYSA-N 0.000 description 1
Classifications
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/63—Introduction of foreign genetic material using vectors; Vectors; Use of hosts therefor; Regulation of expression
- C12N15/79—Vectors or expression systems specially adapted for eukaryotic hosts
- C12N15/82—Vectors or expression systems specially adapted for eukaryotic hosts for plant cells, e.g. plant artificial chromosomes (PACs)
- C12N15/8201—Methods for introducing genetic material into plant cells, e.g. DNA, RNA, stable or transient incorporation, tissue culture methods adapted for transformation
- C12N15/8213—Targeted insertion of genes into the plant genome by homologous recombination
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/10—Processes for the isolation, preparation or purification of DNA or RNA
- C12N15/102—Mutagenizing nucleic acids
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/63—Introduction of foreign genetic material using vectors; Vectors; Use of hosts therefor; Regulation of expression
- C12N15/79—Vectors or expression systems specially adapted for eukaryotic hosts
- C12N15/82—Vectors or expression systems specially adapted for eukaryotic hosts for plant cells, e.g. plant artificial chromosomes (PACs)
- C12N15/8216—Methods for controlling, regulating or enhancing expression of transgenes in plant cells
Definitions
- sequence listing is submitted electronically via EFS-Web as an ASCII formatted sequence listing with a file named 446220SEQLIST.TXT, created on May 13, 2014, and having a size of 62.3 kilobytes, and is filed concurrently with the specification.
- sequence listing contained in this ASCII formatted document is part of the specification and is herein incorporated by reference in its entirety.
- Zinc finger nucleases ZFNs
- TAL effector nucleases TALENs
- the present invention is drawn to methods for targeted mutagenesis and genome engineering in plants using RNA-guided Cas nucleases and assay methods of the rapid testing of the components of the Cas system in plants.
- the invention is based in part on the discovery that the genome of a plant cell can be modified at or in vicinity of a target nucleic acid sequence by introducing into the plant cell a Cas nuclease, particularly a Cas9 nuclease and an engineered, single guide RNA (sgRNA) that is designed for recognition of the target nucleic acid sequence.
- sgRNA single guide RNA
- FIG. 1 Targeted mutagenesis in planta using the Cas9 RNA-guided
- FIG. 2 Measurement of the mutation rate induced by Cas9 RNA-guided endonuclease in N. benthamiana.
- the PDS locus was amplified using non-digested genomic DNA from leaf tissue expressing Cas9 and sgRNA targeting PDS as well as from negative controls (Cas9 plus a sgRNA targeting GFP, and Cas9 on its own).
- the intensity of the uncut band was divided by intensity of all bands in the lane.
- the arrowhead indicates the Mlyl- resistant band; the asterisk indicates the band resulting from star activity of Mlyl.
- Three additional replicates of this experiment are presented in FIG. 8.
- FIG. 3 Cas9 localises to plant nuclei. GFP-Cas9 was expressed in N.
- A Schematic of Arabidopsis U6p::sgRNA construct.
- B Arabidopsis U6 promoter sequence is shown in regular uppercase; the sgRNA transcription start nucleotide is shown in bold lowercase (g); the guide sequence is shown in underlined uppercase; the sgRNA backbone sequence is shown in lowercase italics; the RNA polymerase III terminator sequence is shown in underlined lowercase italics.
- FIG. 5 Replicates of the experiment presented in Figure 1.
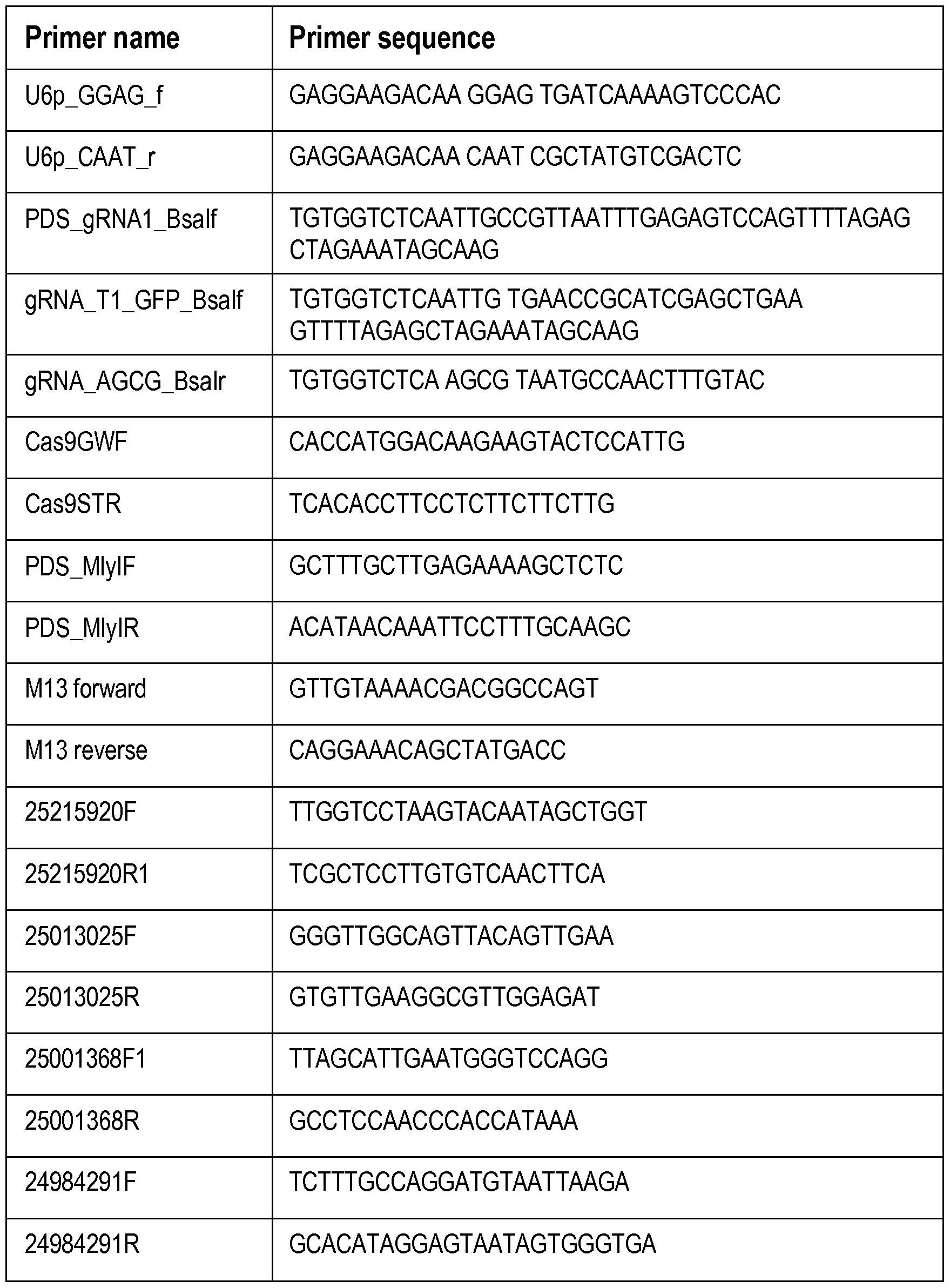
- the PCR primers The PCR primers.
- amplification was done with undigested genomic DNA (a) or Mlyl-digested genomic DNA (b).
- pre-digested genomic DNA co-expression of both Cas9 and sgRNA results in markedly increased levels of the amplicon.
- a total of three independent replicates (1-3) are shown.
- FIG. 6 PCR products amplified on genomic DNA partially digested with Mlyl are resistant to Mlyl digestion only when Cas9 and sgRNA are co-expressed in plant tissue.
- the arrow head indicates the Mlyl- resistant band; the asterisk indicates the band resulting from Mlyl star activity.
- a total of three independent replicates (1-3) are shown.
- FIG. 7 Representative sequence chromatograms of Cas9- induced indels in the Nicotiana benthamiana genome. Regions highlighted in black belong to the target site within the PDS gene.
- FIG. 8 Mutation rate replicates. The PCR was done with the undigested genomic DNA as a template. Resulting amplicons were then digested with Mlyl. The band intensity was quantised with the ImageJ software. The arrowhead indicates the Mlyl% resistant band; the asterisk indicates the band resulting from the star activity of Mlyl.
- FIG. 9 Examples of restriction site loss assays with potential off- targets. Analysis of examples of potential off-targets from groups 1 (a), 2 (b)
- FIG. 10 A large chromosomal deletion can be generated by targeting two adjacent sequences within the PDS locus of Nicotiana benthamiana using transient expression.
- FIG. 11 Large chromosomal deletions can be generated by targeting
- Transgenic plants were produced by agro-transformation of the callus tissue and selection on kanamycin. Agarose gels show PCR bands amplified
- FIG. 12 Sequences of sgRNA constructs of Example 2.
- the AtU6p The AtU6p
- the guide sequence is in italics, the guide sequence is underlined, and the sgRNA
- nucleotide and amino acid sequences listed in the accompanying sequence listing are shown using standard letter abbreviations for nucleotide bases, and three-letter code for amino acids.
- the nucleotide sequences follow the standard convention of beginning at the 5' end of the sequence and proceeding forward (i.e., from left to right in each line) to the 3' end. Only one strand of each nucleotide sequence is shown, but the complementary strand is understood to be included by any reference to the displayed strand.
- amino acid sequences follow the standard convention of beginning at the amino terminus of the sequence and proceeding forward (i.e., from left to right in each line) to the carboxy terminus.
- an "N" in a DNA sequence is intended to mean A, G, C, or T
- an "N” in an RNA sequence is intended to mean A, G, C, or U
- an "N” an amino acid sequence is intended to mean any amino acid, preferably any naturally occurring amino acid in a plant, unless stated otherwise or otherwise apparent from the context of usage.
- SEQ ID NO: 1 sets forth a nucleotide sequence of a human-codon optimized Cas9 (hCas9) coding sequence.
- SEQ ID NO: 2 sets forth the amino acid sequence of the human Cas9 protein encoded by the nucleotide sequence set forth in SEQ ID NO: 1.
- SEQ ID NO: 3 sets forth a nucleotide sequence of the human-codon optimized GFP-hCas9 fusion protein coding seqeuence.
- SEQ ID NO: 4 sets forth the amino acid sequence of the GFP-hCas9 fusion protein encoded by the nucleotide sequence set forth in SEQ ID NO: 3.
- SEQ ID NO: 5 sets forth the nucleotide sequence of the U6p::sgRNA nucleic acid expression construct.
- SEQ ID NO: 6 sets forth an RNA sequence of an sgRNA.
- Embodiments of the invention include, but are not limited to, the following embodiments. 1.
- a method for modifying the genome of a plant, plant part, or plant cell comprising producing at least one plant cell comprising a Cas protein and at least one engineered, single guide RNA (sgRNA) that is capable of recognizing a target nucleic acid sequence, wherein the plant, plant part, plant cell, or descent thereof, comprises in its genome the target nucleic acid sequence.
- sgRNA single guide RNA
- nucleic acid construct for expression of the sgRNA, the nucleic acid construct for expression of the Cas protein, or both are stably incorporated in the genome of the plant cell.
- the target nucleic acid sequence is within a gene or wherein the target nucleic acid sequence affects the level of expression of a gene.
- the gene is selected from the group consisting of a disease resistance gene, a disease susceptibility gene, a gene encoding an enzyme, a gene encoding a structural protein, a gene encoding a receptor, a gene encoding a membrane channel protein, a gene encoding a transcription factor, a gene that is associated with herbicide tolerance, a gene that affects development of male or female reproductive structures, and a gene that affects self-incompatibility.
- the donor nucleic acid molecule comprises a nucleic acid sequence which comprises in a 5' to 3' order: a first homologous region, a donor region, and a second homologous region.
- first homologous region comprises nucleic acid sequence homology to a first genomic region and the second homologous region comprises nucleic acid sequence homology to a second genomic region.
- the donor region comprises at least one transgene.
- the transgene comprises a promoter operably linked to a nucleic acid sequence, and wherein the promoter is capable of driving expression of the operably linked nucleic acid in a plant cell.
- the donor nucleic acid comprises the entire, at least a portion of, or none of, the coding region of a protein encoded by the selectable marker gene.
- nucleic acid construct for expression of the sgR A comprises a promoter operably linked to a nucleic acid sequence encoding the sgRNA and wherein the promoter is capable of driving expression of a nucleic acid sequence in a plant cell.
- nucleic acid construct for expression of the sgR A comprises the nucleic acid sequence of the U6::sgRNA construct set forth in FIG. 4B.
- nucleic acid construct for expression of the Cas protein comprises a promoter operably linked to a nucleic acid sequence encoding the Cas protein and optionally an operably linked eukaryotic nuclear localization signal sequence, wherein the promoter is capable of driving expression of a nucleic acid sequence in a plant cell.
- the promoter is selected from the group consisting of the CaMV 35S promoter, a ubiquitin promoter, an estradiol- inducible promoter, a dexamethasone-inducible promoter, an alcohol-inducible promoter, a heat shock promoter, an inflorescence-specific promoter, the Arabidopsis Apetala2 promoter, and the Arabidopsis Agamous promoter.
- sgRNA comprises a guide nucleic acid sequence that is capable of binding to or recognizing the target nucleic acid or its complement.
- a plant, plant part, seed, or plant cell comprising a modification in its genome that was introduced into the genome according to the method of any one of embodiments 1-56 and 132.
- a method for targeting a fusion protein to a target nucleic acid sequence in the genome of a plant, plant part, or plant cell producing at least one plant cell comprising a fusion protein and an engineered, single guide RNA (sgRNA), wherein the fusion protein comprises a Cas protein domain that is completely or partially defective in nuclease activity and an additional domain, wherein the sgRNA is capable of recognizing the target nucleic acid sequence, whereby the fusion protein is capable of binding to or recognizing the target sequence when in the presence of the sgRNA.
- the Cas protein domain is a Cas9 protein domain or mutant or dervitive of Cas9 protein domain.
- Cas9 protein domain comprises the DIOA amino acid substitution, the H841A amino acid substitution, or both the DIOA and H841A amino acid substitutions.
- nucleic acid construct for expression of the sgRNA, the nucleic acid construct for expression of the fusion protein, or both are stably incorporated in the genome of the plant cell.
- the additional domain is selected from the group consisting of an EAR domain, a SRDX domain, a TOPLESS domain, a cytidine deaminase or other DNA or histone modifying enzyme domain, a MAGI domain, and a RAD54 domain.
- nucleic acid construct for expression of the fusion protein comprises a promoter operably linked to a nucleic acid sequence encoding the fusion protein and optionally an operably linked eukaryotic nuclear localization signal sequence, wherein the promoter is capable of driving expression of a nucleic acid sequence in a plant cell.
- the promoter is a constitutive promoter, an inducible promoter, a developmentally regulated promoter, a tissue-specific promoter, and a seed-specific promoter.
- the promoter is selected from the group consisting of the CaMV 35S promoter, a ubiquitin promoter, an estradiol- inducible promoter, a dexamethasone-inducible promoter, an alcohol-inducible promoter, a heat shock promoter, an inflorescence-specific promoter, the Arabidopsis Apetala2 promoter, and the Arabidopsis Agamous promoter.
- nucleic acid construct for expression of the sgRNA comprises a promoter operably linked to a nucleic acid sequence encoding the sgRNA, wherein the promoter is capable of driving expression of a nucleic acid sequence in a plant cell.
- nucleic acid construct for expression of the sgR A comprises the nucleic acid sequence of the U6: :sgRNA construct set forth in FIG. 4B.
- sgRNA comprises a guide nucleic acid sequence that is capable of binding to or recognizing the target nucleic acid or its complement.
- a plant, plant part, seed, or plant cell comprising the fusion protein and/or sgRNA introduced into the genome according to the method of any one of embodiments 61-93 or comprising at least one modification in its genome that is due to the fusion protein and the sgRNA according to the method of any one of embodiments 61-93.
- a transient assay method for use in testing components of the Cas system in plants comprising the steps of:
- sgRNA engineered, single guide R A
- any one of embodiments 99-104 wherein at least one of the first and the second bacterial strains is a strain from a bacterial species that is selected from the group consisting of Agrobacterium tumefaciens, Agrobacterium rhizogenes, Ensifer adhaerens and other Ensifer spp., Sinorhizobium meliloti, and Mesorhizobium loti.
- step (a) comprises infiltration of the plant tissue with one or more cells of the first bacterial strain and/or the contacting of step (b) comprises infiltration of the plant tissue with one or more cells of the second bacterial strain.
- infiltration comprises use of a syringe to deliver the cells to the plant tissue.
- sgRNA comprises a guide nucleic acid sequence that is capable of binding to or recognizing the target nucleic acid or its complement.
- monitoring the plant tissue for a modification of the genome within the target site comprises determining the nucleotide sequence of all or a part of the target nucleic acid sequence.
- step (c) comprises:
- digesting the genomic DNA with a restriction enzyme that recognizes the restriction enzyme recognition sequence; amplifying the digested DNA by polymerase chain reaction using primers designed to amplify across the target nucleic acid sequence so as to produce PCR products comprising one or more modifications within target nucleic sequence;
- restriction enzyme recognition sequence is a Mlyl restriction enzyme recognition sequence.
- a method for modifying the genome of a plant, plant part, or plant cell comprising producing at least one plant cell comprising a Cas protein and at least two engineered, sgRNAs that are each capable of recognizing a different target nucleic acid sequence, wherein the plant, plant part, plant cell, or descent thereof, comprises in its genome the target nucleic acid sequences.
- ZFNs zinc finger nucleases
- TALENs TAL effector nucleases
- the present invention provides new methods for targeted mutagenesis and genome engineering that do not depend on the use of a ZFNs or a TALENs but instead involve the use of a Cas protein and engineered, single guide RNA (sgRNA) that specifies a targeted nucleic acid sequence .
- sgRNA single guide RNA
- the Cas protein, Cas9 is used for purposes of illustration.
- the present invention is not limited to the use of Cas9.
- Mutants and derivatives of Cas9 as well as other Cas proteins can be used in the methods disclosed herein.
- such other Cas proteins are able to recognize a target nucleic acid sequence when in a plant cell in the presence of an sgRNA that is engineered for recognition of the target sequence.
- the Cas proteins will additional comprise a double-strand endonuclease activity or a single strand endonuclease activity (i.e., a nickase activity). In other embodiments, the Cas protein will not comprise a double-strand endonuclease activity and/or single strand endonuclease activity. In yet other embodiments, a Cas protein domain will be operably linked with at least one additional protein domain to form a fusion protein. Such a fusion protein a can comprise a double-strand endonuclease activity or a single strand endonuclease activity or no endonuclease at all but instead can comprise an additional activity that is due to the additional domain.
- fragments and variants of the disclosed nucleic acid molecules and proteins encoded thereby are also encompassed by the present invention.
- Such fragments and variants of the disclosed nucleic acid molecules and proteins include, for example, nucleic acid molecules encoding mutants and dervivaties of the Cas9 protein and the proteins encoded thereby.
- the nucleic acid and amino acid sequences of Cas9 are disclosed elsewhere herein or otherwise known in the art.
- fragment is intended a portion of the nucleic acid molecule or a portion of the amino acid sequence and hence protein encoded thereby.
- Fragments of a nucleic acid molecule comprising coding sequences may encode protein fragments that retain biological activity of the native protein and hence DNA recognition or binding activity to a target DNA sequence as herein described.
- fragments of a nucleic acid molecule that are useful as hybridization probes generally do not encode proteins that retain biological activity or do not retain promoter activity.
- fragments of a nucleic acid molecule may range from at least about 20 nucleotides, about 50 nucleotides, about 100 nucleotides, and up to the full-length nucleic acid molecule of the invention.
- a variant comprises a nucleic acid molecule having deletions (i.e., truncations) at the 5' and/or 3' end; deletion and/or addition of one or more nucleotides at one or more internal sites in the native polynucleotide; and/or substitution of one or more nucleotides at one or more sites in the native polynucleotide.
- a "native" nucleic acid molecule or polypeptide comprises a naturally occurring nucleiotide sequence or amino acid sequence, respectively.
- nucleic acid molecules conservative variants include those sequences that, because of the degeneracy of the genetic code, encode the amino acid sequence of one of the polypeptides of the invention.
- Variant nucleic acid molecules also include synthetically derived nucleic acid molucules, such as those generated, for example, by using site-directed mutagenesis but which still encode a protein of the invention.
- variants of a particular nucleic acid molecule of the invention will have at least about 70%, 75%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%), 96%o, 97%), 98%>, 99% or more sequence identity to that particular polynucleotide as determined by sequence alignment programs and parameters as described elsewhere herein.
- Variants of a particular nucleic acid molecule of the invention can also be evaluated by comparison of the percent sequence identity between the polypeptide encoded by a variant nucleic acid molecule and the polypeptide encoded by the reference nucleic acid molecule. Percent sequence identity between any two polypeptides can be calculated using sequence alignment programs and parameters described elsewhere herein. Where any given pair of nucleic acid molecule of the invention is evaluated by comparison of the percent sequence identity shared by the two polypeptides that they encode, the percent sequence identity between the two encoded polypeptides is at least about 70%, 75%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or more sequence identity.
- Variant protein is intended to mean a protein derived from the native protein by deletion (so-called truncation) of one or more amino acids at the N-terminal and/or C- terminal end of the native protein; deletion and/or addition of one or more amino acids at one or more internal sites in the native protein; or substitution of one or more amino acids at one or more sites in the native protein.
- Variant proteins encompassed by the present invention are biologically active, that is they continue to possess the desired biological activity of the native protein as described herein. Such variants may result from, for example, genetic polymorphism or from human manipulation.
- Biologically active variants of a protein of the invention will have at least about 70%, 75%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or more sequence identity to the amino acid sequence for the native protein as determined by sequence alignment programs and parameters described elsewhere herein.
- a biologically active variant of a protein of the invention may differ from that protein by as few as 1-15 amino acid residues, as few as 1-10, such as 6-10, as few as 5, as few as 4, 3, 2, or even 1 amino acid residue.
- proteins or polypeptides of the invention may be altered in various ways including amino acid substitutions, deletions, truncations, and insertions. Methods for such manipulations are generally known in the art. For example, amino acid sequence variants and fragments of the proteins can be prepared by mutations in the DNA.
- deletions, insertions, and substitutions of the protein sequences encompassed herein are not expected to produce radical changes in the characteristics of the protein. However, when it is difficult to predict the exact effect of the substitution, deletion, or insertion in advance of doing so, one skilled in the art will appreciate that the effect will be evaluated by routine screening assays as described elsewhere herein or known in the art.
- Variant nucleic acid molecule and proteins also encompass sequences and proteins derived from a mutagenic and recombinogenic procedure such as DNA shuffling.
- Strategies for such DNA shuffling are known in the art. See, for example, Stemmer (1994) Proc. Natl. Acad. Sci. USA 91 : 10747-10751; Stemmer (1994) Nature 370:389-391; Crameri et al. (1997) Nature Biotech. 15:436-438; Moore et al. (1997) J. Mol. Biol. 272:336-347; Zhang et al. (1997) Proc. Natl. Acad. Sci. USA 94:4504-4509; Crameri et al. (1998) Nature 391 :288-291; and U.S. Patent Nos. 5,605,793 and
- oligonucleotide primers can be designed for use in PCR reactions to amplify corresponding nucleic acid molecules from cDNA or genomic DNA extracted from any organism of interest.
- Methods for designing PCR primers and PCR cloning are generally known in the art and are disclosed in Sambrook et al. (1989)
- PCR Protocols A Guide to Methods and Applications (Academic Press, New York); Innis and Gelfand, eds. (1995) PCR Strategies (Academic Press, New York); and Innis and Gelfand, eds. (1999) PCR Methods Manual (Academic Press, New York).
- Known methods of PCR include, but are not limited to, methods using paired primers, nested primers, single specific primers, degenerate primers, gene-specific primers, vector-specific primers, partially-mismatched primers, and the like.
- nucleic acid molecules and proteins of the invention encompass polynucleotide molecules and proteins comprising a nucleotide or an amino acid sequence that is sufficiently identical to the nucleotide sequences or to the amino acid sequence disclosed herein.
- the term "sufficiently identical" is used herein to refer to a first amino acid or nucleotide sequence that contains a sufficient or minimum number of identical or equivalent (e.g., with a similar side chain) amino acid residues or nucleotides to a second amino acid or nucleotide sequence such that the first and second amino acid or nucleotide sequences have a common structural domain and/or common functional activity.
- amino acid or nucleotide sequences that contain a common structural domain having at least about 70% identity, preferably 75% identity, more preferably 85%, 90%, 95%, 96%, 97%, 98% or 99% identity are defined herein as sufficiently identical.
- the sequences are aligned for optimal comparison purposes.
- the two sequences are the same length.
- the percent identity between two sequences can be determined using techniques similar to those described below, with or without allowing gaps. In calculating percent identity, typically exact matches are counted.
- Gapped BLAST can be utilized as described in Altschul et al. (1997) Nucleic Acids Res. 25:3389.
- PSI-Blast can be used to perform an iterated search that detects distant relationships between molecules. See Altschul et al. (1997) supra.
- the default parameters of the respective programs e.g., XBLAST and
- NBLAST NBLAST
- Another preferred, non- limiting example of a mathematical algorithm utilized for the comparison of sequences is the algorithm of Myers and Miller (1988) CABIOS 4: 11-17. Such an algorithm is incorporated into the ALIGN program (version 2.0), which is part of the GCG sequence alignment software package.
- ALIGN program version 2.0
- a PAM120 weight residue table a gap length penalty of 12, and a gap penalty of 4 can be used. Alignment may also be performed manually by inspection.
- sequence identity/similarity values refer to the value obtained using the full-length sequences of the invention and using multiple alignment by mean of the algorithm Clustal W (Nucleic Acid Research, 22(22):4673- 4680, 1994) using the program AlignX included in the software package Vector NTI Suite Version 7 (InforMax, Inc., Bethesda, MD, USA) using the default parameters; or any equivalent program thereof.
- Equivalent program is intended any sequence comparison program that, for any two sequences in question, generates an alignment having identical nucleotide or amino acid residue matches and an identical percent sequence identity when compared to the corresponding alignment generated by
- the methods of the invention involve introducing a polypeptide or
- polynucleotide into a plant is intended to mean presenting to the plant the polynucleotide or polypeptide in such a manner that the sequence gains access to the interior of a cell of the plant.
- the methods of the invention do not depend on a particular method for introducing a sequence into a plant, only that the polynucleotide or polypeptides gains access to the interior of at least one cell of the plant.
- Methods for introducing polynucleotide or polypeptides into plants are known in the art including, but not limited to, stable transformation methods, transient transformation methods, and virus-mediated methods.
- “Stable transformation” is intended to mean that the nucleotide construct introduced into a plant integrates into the genome of the plant and is capable of being inherited by the progeny thereof.
- “Transient transformation” is intended to mean that a polynucleotide is introduced into the plant and does not integrate into the genome of the plant or a polypeptide is introduced into a plant.
- Transformation protocols as well as protocols for introducing polypeptides or polynucleotide sequences into plants may vary depending on the type of plant or plant cell, i.e., monocot or dicot, targeted for transformation. Suitable methods of introducing polypeptides and polynucleotides into plant cells include microinjection (Crossway et al.
- Patent No. 5,736,369 (cereals); Bytebier et al. (1987) Proc. Natl. Acad. Sci. USA
- Plant Cell 4: 1495-1505 electroroporation
- the nucleic acid molecules and constructs of the invention can be provided to a plant using a variety of transient transformation methods.
- transient transformation methods include, but are not limited to, the introduction of a Cas protein and/or one or more sgRNAs directly into the plant or the introduction into the plant of a Cas protein transcript and/or one or more transcripts encoding sgRNAs.
- Such methods include, for example, microinjection or particle bombardment. See, for example, Crossway et al. (1986) Mol Gen. Genet. 202: 179-185; Nomura et al. (1986) Plant Sci. 44:53-58; Hepler et al. (1994) Proc. Natl. Acad. Sci. 91 : 2176-2180 and Hush et al. (1994) The Journal of Cell Science 707:775-784, all of which are herein incorporated by reference.
- the polynucleotide can be transiently transformed into the plant using techniques known in the art. Such techniques include viral vector system and the precipitation of the polynucleotide in a manner that precludes subsequent release of the DNA. Thus, the transcription from the particle-bound DNA can occur, but the frequency with which its released to become integrated into the genome is greatly reduced.
- Such methods include the use particles coated with polyethylimine (PEI; Sigma #P3143).
- the polynucleotide of the invention may be introduced into plants by contacting plants with a virus or viral nucleic acids.
- such methods involve incorporating a nucleotide construct of the invention within a viral DNA or RNA molecule.
- the an nucleic acid molecule of the invention may be initially synthesized as part of a viral polyprotein, which later may be processed by proteolysis in vivo or in vitro to produce the desired recombinant protein.
- promoters of the invention also encompass promoters utilized for transcription by viral RNA polymerases. Methods for introducing polynucleotides into plants and expressing a protein encoded therein, involving viral DNA or RNA molecules, are known in the art. See, for example, U.S. Patent Nos. 5,889,191,
- Methods are known in the art for the targeted insertion of a polynucleotide at a specific location in the plant genome.
- the insertion of the polynucleotide at a desired genomic location is achieved using a site-specific
- the polynucleotide of the invention can be contained in transfer cassette flanked by two non-recombinogenic recombination sites.
- the transfer cassette is introduced into a plant having stably incorporated into its genome a target site which is flanked by two non-recombinogenic recombination sites that correspond to the sites of the transfer cassette.
- An appropriate recombinase is provided and the transfer cassette is integrated at the target site.
- the polynucleotide of interest is thereby integrated at a specific chromosomal position in the plant genome.
- the cells that have been transformed may be grown into plants in accordance with conventional ways. See, for example, McCormick et al. (1986) Plant Cell Reports 5:81-84. These plants may then be grown, and either pollinated with the same transformed strain or different strains, and the resulting progeny having constitutive expression of the desired phenotypic characteristic identified. Two or more generations may be grown to ensure that expression of the desired phenotypic characteristic is stably maintained and inherited and then seeds harvested to ensure expression of the desired phenotypic characteristic has been achieved. In this manner, the present invention provides transformed seed (also referred to as "transgenic seed") having a polynucleotide of the invention, for example, an expression cassette of the invention, stably incorporated into their genome.
- the term plant also includes plant cells, plant protoplasts, plant cell tissue cultures from which plants can be regenerated, plant calli, plant clumps, and plant cells that are intact in plants or parts of plants such as embryos, pollen, ovules, seeds, leaves, flowers, branches, fruit, kernels, ears, cobs, husks, stalks, roots, root tips, anthers, and the like. Grain is intended to mean the mature seed produced by
- Progeny, variants, and mutants of the regenerated plants are also included within the scope of the invention, provided that these parts comprise the introduced
- the present invention may be used for any plant species of interest, including, but not limited to, monocots and dicots.
- plant species of interest include, but are not limited to, corn (Zea mays), Brassica sp. (e.g., B. napus, B.
- rapa, B.juncea particularly those Brassica species useful as sources of seed oil, alfalfa (Medicago sativa), rice (Oryza sativa), rye (Secale cereale), sorghum (Sorghum bicolor, Sorghum vulgare), millet (e.g., pearl millet (Pennisetum glaucum), proso millet (Panicum miliaceum), foxtail millet (Setaria italica), finger millet (Eleusine coracana)), sunflower (Helianthus annuus), safflower (Carthamus tinctorius), wheat (Triticum aestivum), soybean (Glycine max), tobacco (Nicotiana tabacum), potato (Solanum tuberosum), peanuts (Arachis hypogaea), cotton (Gossypium barbadense, Gossypium hirsutum), sweet potato (Ipomoea batatus), cassava (Man
- Vegetables include tomatoes (Lycopersicon esculentum), lettuce (e.g., Lactuca sativa), green beans (Phaseolus vulgaris), lima beans (Phaseolus limensis), peas (Lathyrus spp.), and members of the genus Cucumis such as cucumber (C sativus), cantaloupe (C. cantalupensis), and musk melon (C melo).
- tomatoes Locopersicon esculentum
- lettuce e.g., Lactuca sativa
- green beans Phaseolus vulgaris
- lima beans Phaseolus limensis
- peas Lathyrus spp.
- members of the genus Cucumis such as cucumber (C sativus), cantaloupe (C. cantalupensis), and musk melon (C melo).
- Ornamentals include azalea (Rhododendron spp.), hydrangea (Macrophylla hydrangea), hibiscus (Hibiscus rosasanensis), roses (Rosa spp.), tulips (Tulipa spp.), daffodils (Narcissus spp.), petunias (Petunia hybrida), carnation (Dianthus caryophyllus), poinsettia (Euphorbia pulcherrima), and chrysanthemum.
- Conifers that may be employed in practicing the present invention include, for example, pines such as loblolly pine (Pinus taeda), slash pine (Pinus elliotii), ponderosa pine (Pinus ponderosa), lodgepole pine (Pinus contorta), and Monterey pine (Pinus radiata); Douglas-fir (Pseudotsuga menziesii); Western hemlock (Tsuga canadensis); Sitka spruce (Picea glauca); redwood (Sequoia sempervirens); true firs such as silver fir (Abies amabilis) and balsam fir (Abies balsamea); and cedars such as Western red cedar (Thuja plicata) and Alaska yellow-cedar (Chamaecyparis nootkatensis).
- pines such as loblolly pine (Pinus taeda), slash pine (P
- a “subject plant or plant cell” is one in which genetic alteration, such as transformation, has been effected as to a gene of interest, or is a plant or plant cell which is descended from a plant or cell so altered and which comprises the alteration.
- a “control” or “control plant” or “control plant cell” provides a reference point for measuring changes in phenotype of the subject plant or plant cell.
- a control plant or plant cell may comprise, for example: (a) a wild-type plant or cell, i.e., of the same genotype as the starting material for the genetic alteration which resulted in the subject plant or cell; (b) a plant or plant cell of the same genotype as the starting material but which has been transformed with a null construct (i.e.
- a construct which has no known effect on the trait of interest such as a construct comprising a marker gene
- a construct comprising a marker gene a construct which has no known effect on the trait of interest, such as a construct comprising a marker gene
- a plant or plant cell which is a non-transformed segregant among progeny of a subject plant or plant cell
- a plant or plant cell genetically identical to the subject plant or plant cell but which is not exposed to conditions or stimuli that would induce expression of the gene of interest or (e) the subject plant or plant cell itself, under conditions in which the gene of interest is not expressed.
- benthamiana leaf tissue using standard agroinfiltration protocols 6 and observed a clear nuclear localization (FIG. 3) consistent with the nuclear localization previously observed in human cells 7 .
- the sgRNA was placed under an Arabidopsis U6 promoter (FIG. 4). Both GFP-Cas9 and sgRNA were co-expressed in N. benthamiana leaf tissue using A. tumefaciens as a vector. The tissue was harvested 2 days later and DNA was extracted.
- the assay was robust and reproducible because we detected Mlyl-resistant amplicons in three additional independent experiments using different plants (FIG. 5 and FIG. 6).
- the PCR products from FIG. lb lanes 1 and 4 were cloned into a high copy vector and individual clones sequenced. Sequence analysis of 20 clones derived from the PCR product in lane 1 revealed the presence of indels in 17 of them. The indels can be grouped into nine different types ranging from 1-9 bp deletions to 1 bp insertions (FIG. lc and FIG. 7). All recovered indels abolish the Mlyl restriction site within the target region. Sequences of the 8 clones derived from the control PCR product shown in lane 4 were all wild type.
- the Cas9/sgRNA system may not be as specific as TALEN-induced mutagenesis.
- Cas9 and an engineered sgRNA can direct DNA breaks at defined chromosomal locations in plants.
- the rapid and robust transient assay we developed will enable plant-specific optimization of the Cas9 system.
- the Cas9 system has the potential to simplify the process of plant genome engineering and editing since only a short fragment in the sgRNA needs to be designed to target a new locus. This creates a valuable new tool for plant biologists and breeders, and hastens the prospects of achieving routine targeted genome engineering for basic and applied science. Materials and Methods
- Cas9 was PCR-amplified with primers Cas9GWF and Cas9STR using the clone described in Mali et al. 5 as a template.
- the resulting PCR product was cloned into the pENTR/D-TOPO vector (Life Technologies).
- the entry clone was subsequently recombined into the GW-compatible destination vector pK7WGF2 9 using the LR clonase (Life Technologies) to produce the 35Sp::GFPCas9 construct.
- the Arabidopsis U6 promoter used in this study is the consensus sequence of the 3 U6 promoter variants present in the Arabidopsis genome 10 .
- the AtU6p clone was synthesised with Genescript (FIG. 4).
- the promoter was PCR-amplified with primers U6p_GGAG_f and U6p_CAAT_r and cloned into a level 0 vector 11 via the Bbsl cut- ligation using the Golden Gate (GG) cloning method.
- the sgRNA was PCR-amplified using primers PDS gRNAl Bsalf and gRNA AGCG Bsalr (PDS locus) or primers gRNA T l GFP Bsalf and gRNA AGCG Bsalr (GFP) using the plasmid
- gRNA GFP Tl described in Mali et al. 1 as a template.
- the resulting PCR product was cloned into the pICH86966 vector (kindly provided by S. Marillonnet, based on the modular cloning system described in Weber et al. n under the AtU6p via the Bsal cut- ligation using the GG method.
- Both Cas9 and gRNA GFP Tl plasmids 5 were obtained from Addgene.
- GFP-Cas9 was localised in the leaf tissue of N. benthamiana 2 days post infiltration using the Leica SP5 confocal microscope in accordance with manufacturer's instructions.
- GFP-Cas9 and sgRNA carrying the guide sequence matching a site within the PDS gene of N. benthamiana were transiently co-expressed in the N. benthamiana leaf tissue as follows.
- An Agrobacterium strain carrying a nucleic acid construct of the expression of GFP-Cas9 in plant cells and an Agrobacterium strain carrying a nucleic acid construct of the expression of the sgRNA in plant cells were separately inoculated into L medium containing respective antibiotics for plasmid selection and grown overnight at 28°C. The next day, each of the resulting cultures was centrifuged at 3500 rpm for 10 min, and the resulting pellets of Agrobacteria were separately resuspended in 10 mM MgCl 2 .
- the tissue was harvested at 2 days post infiltration and the genomic DNA extracted using the DNeasy Plant Mini kit (Qiagen). 100 ng of the genomic DNA was then digested overnight with Mlyl restriction enzyme to reduce greatly the background of the non-modified (wild-type) DNA. Upon digesting the reaction mix was desalted using Sepharose CL-6B (Sigma) and then added as a template into a PCR reaction performed with primers PDS MlylF and PDS MlylR with flank the sgRNA target nucleic acid sequence and Phusion DNA polymerase (New England Biolabs). The resulting PCR band was cloned a high copy vector (pCRTM-Blunt II-TOPO vector, Life Technologies, Inc.).
- Plasmids from individual Escherichia coli colonies were sequenced using standard primers Ml 3 forward and Ml 3 reverse. Amplicons with mutated Mlyl sites were recovered only when the Cas9 and sgRNA constructs were co-expressed (FIG. 1 and FIG. 5) but weak bands can be occasionally observed in the negative controls probably due to incomplete digestion. To ensure that the amplicons indeed contain DNA fragments resistant to Mlyl digestion, we performed a partial digestion of the genomic DNA with Mlyl followed by a second Mlyl digestion of the amplicons as shown in FIG. 6.
- chromosomal deletions were generated by targeting two adjacent sequences within the PDS locus of Nicotiana benthamiana using transient expression.
- two sgRNAs were transiently expressed resulted in chromosomal deletion events.
- the sgRNAs were designed for target sequences on each side of the region in the PDS locus that was intended to be deleted.
- FIG. 10A A schematic representation of the PDS locus and targets is shown in FIG. 10A.
- FIG. 10B shows the detection of deletion mutations using the AFLP analysis. The deletions obtained are shown in FIG. IOC.
- the nucleic acid sequence of the sgRNA constructs are shown in FIG. 12.
- chromosomal deletions were generated by targeting two adjacent sequences within the Mlol gene of tomato using transient expression.
- sgRNAs were transiently expressed resulted in chromosomal deletion events.
- the sgRNAs were designed for target sequences on each side of the regions in the Mlol gene that was intended to be deleted as shown (FIG. 11 A).
- FIG. 1 IB shows the detection of deletion mutations by the AFLP analysis
- FIG. 1 1C shows the results from the genotyping of TO transgenic tomato plants. The plants were genotyped for the presence of CRISPR/Cas-induced deletions.
- results demonstrate applicability of the CRISPR/Cas system for purposes of creating large chromosomal deletions in plants.
- Creating a large deletion by targeting with 2 sgRNAs provides a more convenient way to knock-out a gene as such deletion event can be easily detected by the AFLP (PCR band shift) assay.
- Constructs can be quickly tested using the transient expression systems in crops amenable to transient expression (e.g. tomato) before proceeding with generation of stable transgenic lines.
- ability to create large chromosomal deletions provides an opportunity for deleting whole gene clusters as well as deleting certain promoter elements or protein domains.
- NbPDS sgRNAs 1 and 2 NbPDS sgRNAs 1 and 2:
- pICH47732::NOSp::NPTII::OCST pICH47742::2x35S-5'UTR::Cas9::NOST
- pICH47751 ::AtU6p::sgRNA2_S/ oi pICH47761 ::AtU6p::sgRNA3_S/ oi level 1 constructs plus the pELE-4 linker were cut-ligated into the pAGM4723 level 2 vector.
- pICH47732::NOSp::NPTII::OCST pICH47742::2x35S-5'UTR::Cas9::NOST
- pICH47751 ::AtU6p::sgRNA3_S/ oi pICH47761 ::AtU6p::sgRNA4_S/ oi level 1 constructs plus the pELE-4 linker were cut-ligated into the pAGM4723 level 2 vector.
- the resulting level 2 constructs were transformed into the AGL1 strain of A. tumifaciens.
- Tomato transgenic lines were generated with same constructs as the ones used for the transient assay by agrotransformation of the tomato Moneymaker variety using the AGL1 A. tumifaciens strain. Transformants were selected on kanamycin.
- Cas9 and sgRNAs were expressed in the N. benthamiana or tomato Moneymaker leaf tissue.
- the tissue was harvested at 2 days post infiltration and the genomic DNA extracted using the DNeasy Plant Mini kit (Qiagen).
- the genomic DNA extracted from the transiently expressing or stable transgenic tissue was digested with Sacl (New England Biolabs) in the case of N.
Landscapes
- Health & Medical Sciences (AREA)
- Genetics & Genomics (AREA)
- Life Sciences & Earth Sciences (AREA)
- Engineering & Computer Science (AREA)
- Chemical & Material Sciences (AREA)
- Biomedical Technology (AREA)
- Biotechnology (AREA)
- Organic Chemistry (AREA)
- Zoology (AREA)
- Wood Science & Technology (AREA)
- General Engineering & Computer Science (AREA)
- Bioinformatics & Cheminformatics (AREA)
- Plant Pathology (AREA)
- Molecular Biology (AREA)
- Microbiology (AREA)
- Biophysics (AREA)
- Physics & Mathematics (AREA)
- Biochemistry (AREA)
- General Health & Medical Sciences (AREA)
- Cell Biology (AREA)
- Crystallography & Structural Chemistry (AREA)
- Breeding Of Plants And Reproduction By Means Of Culturing (AREA)
- Measuring Or Testing Involving Enzymes Or Micro-Organisms (AREA)
- Peptides Or Proteins (AREA)
- Micro-Organisms Or Cultivation Processes Thereof (AREA)
Abstract
Methods for modifying the genome of plants at a target nucleic acid sequence are provided. Such methods involve producing a plant cell comprising a Cas protein and a single guide RNA (sgRNA) that is capable of recognizing the target nucleic acid sequence in the plant cell. Further provided are methods for targeting fusion proteins to target nucleic acid sequences in the genome of plant. Such methods involve producing a plant cell comprising a fusion protein and an sgRNA designed to recognize the target nucleic acid sequence. The fusion protein is capable of recognizing the target sequence in the presence of the sgRNA. The fusion protein comprises a Cas protein domain that is defective in nuclease activity and an additional domain. Also provided are methods for testing components of the Cas system in plants, modified plants and plant cells, fusion proteins, and nucleic acid molecules encoding such fusion proteins.
Description
TARGETED MUTAGENESIS AND GENOME ENGINEERING IN PLANTS USING
RNA-GUIDED CAS NUCLEASES
REFERENCE TO A SEQUENCE LISTING SUBMITTED AS A TEXT FILE VIA EFS WEB
The official copy of the sequence listing is submitted electronically via EFS-Web as an ASCII formatted sequence listing with a file named 446220SEQLIST.TXT, created on May 13, 2014, and having a size of 62.3 kilobytes, and is filed concurrently with the specification. The sequence listing contained in this ASCII formatted document is part of the specification and is herein incorporated by reference in its entirety.
BACKGROUND OF THE INVENTION
Sustainable intensification of crop production is essential to ensure food demand is matched by supply as the human population continues to increase1. This will require high yielding crop varieties that can be grown sustainably with fewer inputs on less land. Both plant breeding and GM methods make valuable contributions to varietal improvement, but targeted genome engineering promises to be critical to elevating future yields. Most such methods require targeting DNA breaks to defined locations followed by either non-homologous end joining (NHEJ) or homologous recombination . Zinc finger nucleases (ZFNs) and TAL effector nucleases (TALENs) can be engineered to create such breaks, but the recombinant DNA work is complicated by the necessity to create two different DNA binding proteins flanking a sequence of interest, each with a C- terminal Fokl nuclease module. Therefore, new methods for targeted mutagenesis genome engineering that are less complicated and costly than existing methods are desired.
BRIEF SUMMARY OF THE INVENTION
The present invention is drawn to methods for targeted mutagenesis and genome engineering in plants using RNA-guided Cas nucleases and assay methods of the rapid testing of the components of the Cas system in plants. The invention is based in part on the discovery that the genome of a plant cell can be modified at or in vicinity of a target nucleic acid sequence by introducing into the plant cell a Cas nuclease, particularly a Cas9 nuclease and an engineered, single guide RNA (sgRNA) that is designed for recognition of the target nucleic acid sequence. The methods and compositions of the present invention find use in the genome engineering of plants, particularly in the genome engineering of agricultural plants for the development of improved agricultural plant varieties. This discovery and the methods and compositions of the present invention are further described below.
BRIEF DESCRIPTION OF THE DRAWINGS FIG. 1. Targeted mutagenesis in planta using the Cas9 RNA-guided
endonuclease. (A) Scheme illustrating the assay. (B) DNA gel with PCR bands obtained upon amplification using primers flanking the target site within the phytoene desaturase (PDS) gene of Nicotiana benthamiana. In lanes 1-3 the template genomic DNA was pre-digested with Mlyl, while in lane 4 non-digested genomic DNA was used. (C) Alignment of reads with Cas9-induced indels in the PDS gene obtained from lane 1 of FIG. IB. The wild-type sequence is shown at the top. The sgRNA target sequence is shaded in grey. Deleted DNA residues are shown as asterisks. Insertions and substitutions are shown as underlined capitals. The change in length and sequence are shown to the right. PAM, the protospacer-adjacent motif, was selected to follow the consensus sequence NGG. The change in length and sequence are shown to the right. Three additional replicates of this experiment are presented in FIG. 5.
FIG. 2. Measurement of the mutation rate induced by Cas9 RNA-guided endonuclease in N. benthamiana. The PDS locus was amplified using non-digested genomic DNA from leaf tissue expressing Cas9 and sgRNA targeting PDS as well as from negative controls (Cas9 plus a sgRNA targeting GFP, and Cas9 on its own). To measure the mutation rate, the intensity of the uncut band was divided by intensity of all bands in the lane. The arrowhead indicates the Mlyl- resistant band; the asterisk indicates
the band resulting from star activity of Mlyl. Three additional replicates of this experiment are presented in FIG. 8.
FIG. 3. Cas9 localises to plant nuclei. GFP-Cas9 was expressed in N.
benthamiana leaf tissue and visualised 2 days post in:iltration. (a) GFP. (b) Plastids auto:luorescence. (c) Bright field, (d) Overlay of (a), (b) and (c).
FIG. 4. Description of the U6p: :sgRNA construct. (A) Schematic of Arabidopsis U6p::sgRNA construct. (B) Arabidopsis U6 promoter sequence is shown in regular uppercase; the sgRNA transcription start nucleotide is shown in bold lowercase (g); the guide sequence is shown in underlined uppercase; the sgRNA backbone sequence is shown in lowercase italics; the RNA polymerase III terminator sequence is shown in underlined lowercase italics.
FIG. 5. Replicates of the experiment presented in Figure 1. The PCR
amplification was done with undigested genomic DNA (a) or Mlyl-digested genomic DNA (b). In the case of the pre-digested genomic DNA, co-expression of both Cas9 and sgRNA results in markedly increased levels of the amplicon. A total of three independent replicates (1-3) are shown.
FIG. 6. PCR products amplified on genomic DNA partially digested with Mlyl are resistant to Mlyl digestion only when Cas9 and sgRNA are co-expressed in plant tissue. The arrow head indicates the Mlyl- resistant band; the asterisk indicates the band resulting from Mlyl star activity. A total of three independent replicates (1-3) are shown.
FIG. 7. Representative sequence chromatograms of Cas9- induced indels in the Nicotiana benthamiana genome. Regions highlighted in black belong to the target site within the PDS gene.
FIG. 8. Mutation rate replicates. The PCR was done with the undigested genomic DNA as a template. Resulting amplicons were then digested with Mlyl. The band intensity was quantised with the ImageJ software. The arrowhead indicates the Mlyl% resistant band; the asterisk indicates the band resulting from the star activity of Mlyl.
FIG. 9. Examples of restriction site loss assays with potential off- targets. Analysis of examples of potential off-targets from groups 1 (a), 2 (b)
and 3 (c) (Table 1). The PDS positive control is shown in (d).
FIG. 10. A large chromosomal deletion can be generated by targeting two adjacent sequences within the PDS locus of Nicotiana benthamiana using transient expression. A. Schematic representation explaining setup of the
experiment. B. Detection of deletion mutations using the AFLP analysis.
Agarose gel shows PCR bands amplified across targets 1 and 2 using genomic
DNA extracted from respective leaf samples. Cas9, sgR Al and 2 were
expressed in N. benthamiana leaf tissue using the standard agroinfiltration
protocol. In lane 2, Cas9, sgRNAl, and sgRNA2 were expressed from three
separate plasmids, while in lane 4 they were expressed from a single plasmid. C.
Types of deletion mutations identified. Bottom PCR bands from lanes 2 and 4 were cloned into a high copy vector and 15 individual clones were sequenced.
All clones contained deletions that can be grouped in three different types (ml- 3).
FIG. 11. Large chromosomal deletions can be generated by targeting
two adjacent sequences within the Mlol locus of tomato. (A) Schematic
representation explaining the setup of the experiment. (B) Detection of deletion mutations by the AFLP analysis using transient expression in tomato. Cas9 and sgRNAs were expressed in tomato leaf tissue using the standard agro- infiltration protocol. (C) TO tomato transgenic plants genotyped for the presence of CRISPR/Cas-induced deletions. Plants with encircled numbers carry
deletions. Transgenic plants were produced by agro-transformation of the callus tissue and selection on kanamycin. Agarose gels show PCR bands amplified
across respective targets using genomic DNA extracted from tomato leaf
samples. In all cases, Cas9 and both sgRNAs were expressed from a single
plasmid.
FIG. 12. Sequences of sgRNA constructs of Example 2. The AtU6p
sequence is in italics, the guide sequence is underlined, and the sgRNA
backbone is in bold typeface. SEQUENCE LISTING
The nucleotide and amino acid sequences listed in the accompanying sequence listing are shown using standard letter abbreviations for nucleotide bases, and three-letter code for amino acids. The nucleotide sequences follow the standard convention of
beginning at the 5' end of the sequence and proceeding forward (i.e., from left to right in each line) to the 3' end. Only one strand of each nucleotide sequence is shown, but the complementary strand is understood to be included by any reference to the displayed strand. The amino acid sequences follow the standard convention of beginning at the amino terminus of the sequence and proceeding forward (i.e., from left to right in each line) to the carboxy terminus. As used herein and in the accompanying sequence listing, an "N" in a DNA sequence is intended to mean A, G, C, or T, an "N" in an RNA sequence is intended to mean A, G, C, or U, and an "N" an amino acid sequence is intended to mean any amino acid, preferably any naturally occurring amino acid in a plant, unless stated otherwise or otherwise apparent from the context of usage.
SEQ ID NO: 1 sets forth a nucleotide sequence of a human-codon optimized Cas9 (hCas9) coding sequence.
SEQ ID NO: 2 sets forth the amino acid sequence of the human Cas9 protein encoded by the nucleotide sequence set forth in SEQ ID NO: 1.
SEQ ID NO: 3 sets forth a nucleotide sequence of the human-codon optimized GFP-hCas9 fusion protein coding seqeuence.
SEQ ID NO: 4 sets forth the amino acid sequence of the GFP-hCas9 fusion protein encoded by the nucleotide sequence set forth in SEQ ID NO: 3.
SEQ ID NO: 5 sets forth the nucleotide sequence of the U6p::sgRNA nucleic acid expression construct.
SEQ ID NO: 6 sets forth an RNA sequence of an sgRNA.
DETAILED DESCRIPTION OF THE INVENTION
The present invention now will be described more fully hereinafter with reference to the accompanying FIG.s, in which some, but not all embodiments of the inventions are shown. Indeed, the invention may be embodied in many different forms and should not be construed as limited to the embodiments set forth herein; rather, these embodiments are provided so that this disclosure will satisfy applicable legal requirements. Like numbers refer to like elements throughout.
Many modifications and other embodiments of the invention set forth herein will come to mind to one skilled in the art to which these invention pertains having the benefit of the teachings presented in the foregoing descriptions and the associated
drawings. Therefore, it is to be understood that the invention is not to be limited to the specific embodiments disclosed and that modifications and other embodiments are intended to be included within the scope of the appended claims. Although specific terms are employed herein, they are used in a generic and descriptive sense only and not for purposes of limitation.
Embodiments of the invention include, but are not limited to, the following embodiments. 1. A method for modifying the genome of a plant, plant part, or plant cell, the method comprising producing at least one plant cell comprising a Cas protein and at least one engineered, single guide RNA (sgRNA) that is capable of recognizing a target nucleic acid sequence, wherein the plant, plant part, plant cell, or descent thereof, comprises in its genome the target nucleic acid sequence.
2. The method of embodiment 1, wherein the Cas protein is Cas9 or a mutant or derivative of Cas9.
3. The method of embodiment 2, wherein the mutant or derivative is naturally occurring or engineered.
4. The method of any one of embodiments 1-3, wherein the Cas protein is capable of causing a double-strand break in DNA.
5. The method of any one of embodiments 1-3, wherein the Cas protein is capable of causing a single-strand break in DNA.
6. The method of embodiment 2 or 3, wherein the mutant or derivative comprises the DIOA amino acid substitution or the H841A amino acid substitution, and wherein the mutant or derivative is capable of causing a single-strand break in DNA.
7. The method of any one of embodiments 1-6, wherein producing at least one plant cell comprising a Cas protein and an engineered sgRNA, comprises:
(a) introducing into the at least one plant cell or a progenitor cell thereof a nucleic acid construct for expression of the sgRNA in the plant cell, whereby the sgRNA is expressed in the at least one plant cell;
(b) introducing into the at least one a plant cell or a progenitor cell thereof a nucleic acid construct for expression of the Cas protein in the plant cell, whereby the Cas protein is expressed in the at least one plant cell; or
(c) both (a) and (b).
8. The method of any one of embodiments 1-7, wherein the sgR A, the Cas protein, or both are transiently expressed in the plant cell.
9. The method of embodiment 7 or 8, wherein the nucleic acid construct for expression of the sgRNA, the nucleic acid construct for expression of the Cas protein, or both are stably incorporated in the genome of the plant cell.
10. The method of any one of embodiments 1-9, wherein producing at least one plant cell comprising a Cas protein and an engineered sgRNA, comprises:
(i) directly introducing an engineered sgRNA into a plant cell and optionally further comprises virus-mediated delivery of the sgRNA into the plant cell;
(ii) directly introducing the Cas protein into the plant cell; or
(iii) both (i) and (ii).
11. The method of any one of embodiments 1-10, wherein the Cas protein or the nucleic acid construct for expression of the Cas protein is introduced into the plant cell or the progenitor, before, concurrently with, or after introducing the sgRNA or the nucleic acid construct for expression of the sgRNA.
12. The method of any one of embodiments 1-11, wherein at least one modification is introduced into the genome of the plant, plant part, or plant cell, or descendant thereof.
13. The method of embodiment 12, wherein the modification occurs within the target nucleic acid sequence.
14. The method of embodiment 12 or 13, wherein the modification comprises the insertion of at least one base pair in the genome of the plant, plant part, plant cell, or descendant thereof.
15. The method of embodiment 12 or 13, wherein the modification comprises the deletion of at least one base pair in the genome of the plant, plant part, plant cell, or descendant thereof.
16. The method of embodiment 12 or 13, wherein the modification comprises the insertion of at least one base pair in the genome of the plant, plant part, plant cell, or descendant thereof and the deletion of at least one base pair in the genome of the plant, plant part, plant cell, or descendant thereof.
17. The method of any one of embodiments 1-16, wherein the target nucleic acid sequence is within a gene or wherein the target nucleic acid sequence affects the level of expression of a gene.
18. The method of embodiment 17, wherein the gene is selected from the group consisting of a disease resistance gene, a disease susceptibility gene, a gene encoding an enzyme, a gene encoding a structural protein, a gene encoding a receptor, a gene encoding a membrane channel protein, a gene encoding a transcription factor, a gene that is associated with herbicide tolerance, a gene that affects development of male or female reproductive structures, and a gene that affects self-incompatibility.
19. The method of embodiment 17 or 18, wherein modification is a mutation in the gene that results in an alteration or loss of the wild-type function of the gene and/or the protein encoded thereby.
20. The method of embodiment 19, wherein the mutation is a loss-of- function mutation.
21. The method of any one of embodiments 1-20, further comprising introducing into the plant, plant part, plant cell, or descendant thereof a donor nucleic acid .
22. The method of embodiment 21 , wherein the donor nucleic acid molecule or part thereof is incorporated into the genome of the plant, plant part, or plant cell or descendant thereof.
23. The method of embodiment 21 or 22, wherein the donor nucleic acid molecule or part thereof is incorporated into the genome of the plant, plant part, plant cell, or descendant thereof within the target nucleic acid sequence.
24. The method of any one of embodiments 21-23, wherein the donor nucleic acid molecule comprises a nucleic acid sequence which comprises in a 5' to 3' order: a first homologous region, a donor region, and a second homologous region.
25. The method of embodiment 25, wherein the first homologous region comprises nucleic acid sequence homology to a first genomic region and the second homologous region comprises nucleic acid sequence homology to a second genomic region.
26. The method of embodiment 26, wherein the first genomic region and the second genomic region are on opposite sides of the site of incorporation of the donor nucleic acid molecule or part thereof into the genome of the plant, plant part, plant cell, or descendant thereof.
27. The method of any one of embodiments 21-26, wherein the donor region comprises a coding and/or a non-coding nucleic acid sequence.
28. The method of any one of embodiments 21-27, wherein the donor region comprises at least one transgene.
29. The method of embodiment 28, wherein the transgene comprises a promoter operably linked to a nucleic acid sequence, and wherein the promoter is capable of driving expression of the operably linked nucleic acid in a plant cell.
30. The method of embodiment 28 or 29, where the transgene encodes a protein. 31. The method of embodiment 30, wherein the protein is native to the plant, plant part, plant cell.
32. The method of embodiment 30, wherein the protein is a foreign protein that is not native the plant, plant part, or plant cell.
33. The method of any one of embodiments 21-32, wherein the donor nucleic acid comprises or further comprises at least one selectable marker gene.
34. The method of any one of embodiments 22-33, wherein incorporation of the donor nucleic acid into the genome of the plant, plant part, or plant cell or descendant thereof produces a selectable marker gene in the genome of the plant, plant part, or plant cell or descendant thereof.
35. The method of embodiment 34, wherein the donor nucleic acid comprises the entire, at least a portion of, or none of, the coding region of a protein encoded by the selectable marker gene.
36. The method of any one of embodiments 21-35, wherein the donor nucleic acid molecule is single stranded or double stranded.
37. The method of any one of embodiments 21-36, wherein the donor nucleic acid molecule is a linear nucleic acid molecular or a circular nucleic acid molecule.
38. The method of any one of embodiments 7-37, wherein the nucleic acid construct for expression of the sgR A comprises a promoter operably linked to a nucleic acid sequence encoding the sgRNA and wherein the promoter is capable of driving expression of a nucleic acid sequence in a plant cell.
39. The method of embodiment 38, wherein the promoter is an RNA polymerase III promoter.
40. The method of embodiment 38 or 39, wherein the promoter is the Arabidopsis U6 promoter.
41. The method of any one of embodiments 38-40, wherein the promoter comprises the Arabidopsis U6 promoter-derived synthetic nucleic acid sequence set forth in FIG. 4B.
42. The method of any one of embodiments 7-41, wherein nucleic acid construct for expression of the sgR A comprises the nucleic acid sequence of the U6::sgRNA construct set forth in FIG. 4B.
43. The method of any one of embodiments 7-42, wherein the nucleic acid construct for expression of the Cas protein comprises a promoter operably linked to a nucleic acid sequence encoding the Cas protein and optionally an operably linked eukaryotic nuclear localization signal sequence, wherein the promoter is capable of driving expression of a nucleic acid sequence in a plant cell.
44. The method of embodiment 43, wherein the promoter is an RNA polymerase II promoter.
45. The method of embodiments 43 or 44, wherein the promoter is a constitutive promoter, a tissue-specific promoter or an inducible promoter.
46. The method of any one of embodiments 43-45, wherein the promoter is selected from the group consisting of the CaMV 35S promoter, a ubiquitin promoter, an estradiol- inducible promoter, a dexamethasone-inducible promoter, an alcohol-inducible promoter, a heat shock promoter, an inflorescence-specific promoter, the Arabidopsis Apetala2 promoter, and the Arabidopsis Agamous promoter.
47. The method of any one of embodiments 1-46, wherein the target nucleic acid sequence comprises or consists of 19 bases.
48. The method of any one of embodiments 1-47, wherein immediately following the 3' end of the target nucleic acid sequence in the genome of the plant, plant part, or plant cells is a protospacer-adjacent motif (PAM).
49. The method of embodiment 48, wherein the PAM is NGG, wherein N can be A, C, G, or T.
50. The method of embodiment 47, wherein the Cas protein recognizes a PAM that is not NGG, wherein N can be A, C, G, or T.
51. The method of any one of embodiments 1-50, wherein the sgRNA comprises a guide nucleic acid sequence that is capable of binding to or recognizing the target nucleic acid or its complement.
52. The method of embodiment 51 , wherein the guide nucleic acid sequence is identical to the target nucleic acid sequence or its complement, when a U in the guide nucleic acid sequence is equivalent to a T in the target nucleic acid sequence or its complement.
53. The method of embodiment 51 , wherein the guide nucleic acid sequence is not identical to the target nucleic acid sequence or its complement, when a U in the guide nucleic acid sequence is equivalent to a T in the target nucleic acid sequence or its complement.
54. The method of embodiment 51 or 53, wherein there are one or more mismatched bases between the guide nucleic acid sequence and the target nucleic acid sequence or its complement.
55. The method of embodiment 54, wherein the one or more mismatches occur at a position or positions in the target nucleic acid sequence that are located within the first 8, 9, 10, 11, or 12 bases beginning from the 5' end of the target nucleic acid sequence.
56. The method of any of embodiments 1-55, further comprising regenerating the plant cell or descendent thereof into a plant comprising at least one modification in its genome or in the genome of a progeny plant or descendent of the regenerated plant, wherein at least one modification in the genome of a progeny plant or descendent is due to the presence of the sgRNA and the Cas protein in the progeny plant or descendent.
57. A plant, plant part, seed, or plant cell produced by the method of any one of embodiments 1-56 and 132, wherein the genome of the plant, plant part, seed, or plant cell is modified when compared to the genome of a control plant cell, and wherein said control plant cell is plant cell before its genome has been modified by the method.
58. A plant, plant part, seed, or plant cell comprising a modification in its genome that was introduced into the genome according to the method of any one of embodiments 1-56 and 132.
59. Use of the plant, plant part, or seed of embodiment 57 or 58 in agriculture and/or in the production of a human and/or animal food product.
60. Use of the plant, plant part, or seed of embodiment 57 or 58 in a plant breeding method and/or in the production of hybrid seed.
61. A method for targeting a fusion protein to a target nucleic acid sequence in the genome of a plant, plant part, or plant cell, producing at least one plant cell comprising a fusion protein and an engineered, single guide RNA (sgRNA), wherein the fusion protein comprises a Cas protein domain that is completely or partially defective in nuclease activity and an additional domain, wherein the sgRNA is capable of recognizing the target nucleic acid sequence, whereby the fusion protein is capable of binding to or recognizing the target sequence when in the presence of the sgRNA.
62. The method of embodiment 61, wherein the Cas protein domain is a Cas9 protein domain or mutant or dervitive of Cas9 protein domain.
63. The method of embodiment 62, wherein Cas9 protein domain comprises the DIOA amino acid substitution, the H841A amino acid substitution, or both the DIOA and H841A amino acid substitutions.
64. The method of any one of embodiments 61-63, wherein producing the at least one plant cell comprising the fusion protein and the engineered, single guide R A, comprises:
(a) introducing into the at least one plant cell or a progenitor cell thereof a nucleic acid construct for expression of the sgRNA in the plant cell, whereby the sgRNA is expressed in the at least one plant cell;
(b) introducing into the at least one a plant cell or a progenitor cell thereof a nucleic acid construct for expression of the fusion protein in the plant cell, whereby the fusion protein is expressed in the at least one plant cell; or
(c) both (a) and (b).
65. The method of embodiment of embodiment 64, wherein the sgRNA, the fusion protein, or both are transiently expressed in the plant cell.
66. The method of embodiment 64 or 65, wherein the nucleic acid construct for expression of the sgRNA, the nucleic acid construct for expression of the fusion protein, or both are stably incorporated in the genome of the plant cell.
67. The method of any one of embodiments 61-66, wherein producing the at least one plant cell comprising the fusion protein and the sgRNA, comprises:
(i) directly introducing an engineered sgRNA into a plant cell and optionally further comprises virus-mediated delivery of the sgRNA into the plant cell;
(ii) directly introducing the fusion protein into the plant cell; or
(iii) both (i) and (ii).
68. The method of any one of embodiments 61-67, wherein the fusion protein or the nucleic acid construct for expression of the fusion protein is introduced into the plant cell or the progenitor, before, concurrently with, or after introducing the sgRNA or the nucleic acid construct for expression of the sgRNA.
69. The method of any one of embodiments 61-68, wherein the additional domain is selected from the group consisting of a transcriptional activator domain, a transcriptional
repressor or co-repressor domain, a protein domain known to elevate the local mutagenesis rate, and a protein that can elevate the local recombination rate.
70. The method of any one of embodiments 61-69, wherein the additional domain is selected from the group consisting of an EAR domain, a SRDX domain, a TOPLESS domain, a cytidine deaminase or other DNA or histone modifying enzyme domain, a MAGI domain, and a RAD54 domain.
71. The method of any one of embodiments 61-70, wherein the target nucleic acid sequence is within a gene.
72. The method of any one of embodiments 61-71, wherein the target nucleic acid sequence is within a promoter.
73. The method of any one of embodiments 64-72, wherein the nucleic acid construct for expression of the fusion protein comprises a promoter operably linked to a nucleic acid sequence encoding the fusion protein and optionally an operably linked eukaryotic nuclear localization signal sequence, wherein the promoter is capable of driving expression of a nucleic acid sequence in a plant cell.
74. The method of embodiment 71, wherein the promoter is an RNA polymerase II promoter.
75. The method of embodiment 73 or 74, wherein the promoter is a constitutive promoter, an inducible promoter, a developmentally regulated promoter, a tissue-specific promoter, and a seed-specific promoter.
76. The method of any one of embodiments 73-75, wherein the promoter is selected from the group consisting of the CaMV 35S promoter, a ubiquitin promoter, an estradiol- inducible promoter, a dexamethasone-inducible promoter, an alcohol-inducible promoter, a heat shock promoter, an inflorescence-specific promoter, the Arabidopsis Apetala2 promoter, and the Arabidopsis Agamous promoter.
77. The method of any one of embodiments 64-76, wherein the nucleic acid construct for expression of the sgRNA comprises a promoter operably linked to a nucleic acid sequence encoding the sgRNA, wherein the promoter is capable of driving expression of a nucleic acid sequence in a plant cell.
78. The method of embodiment 77, wherein the promoter is an RNA polymerase III promoter.
79. The method of embodiment 77 or 78, wherein the promoter is the Arabidopsis U6 promoter.
80. The method of any one of embodiments 77-79, wherein the promoter comprises the Arabidopsis U6 promoter-derived synthetic nucleic acid sequence set forth in FIG. 4B.
81. The method of any one of embodiments 77-80, wherein nucleic acid construct for expression of the sgR A comprises the nucleic acid sequence of the U6: :sgRNA construct set forth in FIG. 4B.
82. The method of any one of embodiments 61-81, wherein the target nucleic acid sequence comprises or consists of 19 bases.
83. The method of any one of embodiments 61-82, wherein immediately following the 3' end of the target nucleic acid sequence in the genome of the plant, plant part, or plant cells is a protospacer-adjacent motif (PAM).
84. The method of embodiment 83, wherein the PAM is NGG, wherein N can be A, C, G, or T.
85. The method of embodiment 83, wherein the Cas protein recognizes a PAM that is not NGG, wherein N can be A, C, G, or T.
86. The method of any one of embodiments 61-85, wherein the sgRNA comprises a guide nucleic acid sequence that is capable of binding to or recognizing the target nucleic acid or its complement.
87. The method of embodiment 86, wherein the guide nucleic acid sequence is identical to the target nucleic acid sequence or its complement, when a U in the guide nucleic acid sequence is equivalent to a T in the target nucleic acid sequence or its complement.
88. The method of embodiment 86, wherein the guide nucleic acid sequence is not identical to the target nucleic acid sequence or its complement, when a U in the guide nucleic acid sequence is equivalent to a T in the target nucleic acid sequence or its complement.
89. The method of embodiment 86 or 88, wherein there are one or more mismatched bases between the guide nucleic acid sequence and the target nucleic acid sequence or its complement.
90. The method of embodiment 89, wherein the one or more mismatches occur at a position or positions in the target nucleic acid sequence that are located within the first 8, 9, 10, 11, or 12 bases beginning from the 5' end of the target nucleic acid sequence.
91. The method of any one of embodiments 61-90, further comprising introducing at least one additional engineered sgRNA, wherein the sgRNA is capable of recognizing an additional target nucleic acid sequence.
92. The method of embodiment 91, wherein the fusion protein is also capable of binding to or recognizing the additional target sequence when in the presence of the additional sgRNA.
93. The method of any of embodiments 61-92, further comprising regenerating the plant cell or descendent thereof into a plant comprising the fusion protein and/or the sgRNA and/or at least one modification in the genome of the plant cell or descendent that is due the fusion protein and the sgRNA.
94. A plant, plant part, seed, or plant cell produced by the method of any one of embodiments 61-93, wherein the plant, plant part, seed, or plant cell comprises the fusion protein and the engineered sgRNA.
95. A plant, plant part, seed, or plant cell comprising the fusion protein and/or sgRNA introduced into the genome according to the method of any one of embodiments 61-93 or comprising at least one modification in its genome that is due to the fusion protein and the sgRNA according to the method of any one of embodiments 61-93.
96. Use of the plant, plant part, or seed of embodiment 94 or 95 in agriculture and/or in the production of a human and/or animal food product.
97. Use of the plant, plant part, or seed of embodiment 94 or 95 in a plant breeding method and/or in the production of hybrid seed.
98. A fusion protein according to any of embodiments 61-93 or a nucleic acid molecule encoding the fusion protein.
99. A transient assay method for use in testing components of the Cas system in plants, the method comprising the steps of:
(a) contacting a plant tissue with one or more cells of a first bacterial strain comprising a first nucleic acid construct, wherein the first nucleic acid construct comprises an RNA polymerase II promoter that is expressible in a plant cell operably linked to nucleic acid sequence encoding a Cas protein, and wherein a cell of the first bacterial strain is capable of transferring a nucleic acid molecule to a plant cell;
(b) contacting the plant tissue with one or more cells of a second bacterial strain comprising a second nucleic acid construct, wherein the second nucleic acid construct comprises an RNA polymerase III promoter that is expressible in a plant cell
operably linked to nucleic acid sequence encoding an engineered, single guide R A (sgRNA) that is capable of recognizing a target nucleic acid sequence in the genome of plant tissue, and wherein a cell of the second bacterial strain is capable of transferring a nucleic acid molecule to a plant cell; and
(c) monitoring the plant tissue for a modification of the genome within the target site.
100. The method of embodiment 99, wherein the Cas protein is Cas9 or a mutant or derivative of Cas9.
101. The method of embodiment 100, wherein the mutant or derivative is naturally ocurring or engineered.
102. The method of any one of embodiments 99- 101, wherein the Cas protein is capable of causing a double-strand break in DNA.
103. The method of any one of embodiments 99- 101, wherein the Cas protein is capable of causing a single-strand break in DNA.
104. The method of any one of embodiments 100, 101, and 103, wherein the mutant or derivative comprises the D10A amino acid substitution or the H841A amino acid substitution, and wherein the mutant or derivative is capable of causing a single-strand break in DNA.
105. The method of any one of embodiments 99-104, wherein at least one of the first and the second bacterial strains is a strain from a bacterial species that is selected from the group consisting of Agrobacterium tumefaciens, Agrobacterium rhizogenes, Ensifer adhaerens and other Ensifer spp., Sinorhizobium meliloti, and Mesorhizobium loti.
106. The method of any one of embodiments 99-105, wherein at least one of the first and the second bacterial strains is selected from the group consisting of Agrobacterium tumefaciens strain AGL1 and Ensifer adhaerens strain OV14.
107. The method of any one of embodiments 99-106, wherein at least one of the first and the second nucleic acid constructs is contained in a Ti plasmid or an Agrobacterium T-DNA border-containing broad host range binary vector..
108. The method of any one of embodiments 99-107, wherein the contacting of step (a) comprises infiltration of the plant tissue with one or more cells of the first bacterial strain and/or the contacting of step (b) comprises infiltration of the plant tissue with one or more cells of the second bacterial strain.
109. The method of embodiment 108, wherein infiltration comprises use of a syringe to deliver the cells to the plant tissue.
110. The method of any one of embodiments 99-109, wherein steps (a) and (b) are performed at the same time.
111. The method of embodiment 110, wherein the plant tissue is infiltrated with a mixture comprising cells of the first bacterial strain and the second bacterial strain.
112. The method of any one of embodiments 99-111, wherein the plant tissue is from a plant selected from the group consisting of Nicotiana benthamiana, Nicotiana tabacum, Lactuca sativa, Brassica rapa, and any other plant species in which transient expression of genes after delivery of DNA either from bacteria or by other means can be achieved.
113. The method of any one of embodiments 99-112, wherein the plant tissue is leaf tissue.
114. The method of embodiment 112, wherein the leaf tissue is, or is part of, a whole leaf.
115. The method of any one of embodiments 99-114, wherein the plant tissue is attached to a plant or is detached from a plant.
116. The method of any one of embodiments 99-115, wherein the target nucleic acid sequence comprises or consists of 19 bases.
117. The method of any one of embodiments 99-116, wherein immediately following the 3' end of the target nucleic acid sequence in the genome of the plant, plant part, or plant cells is the protospacer-adjacent motif (PAM).
118. The method of embodiment 117, wherein the PAM is NGG, wherein N can be A, C, G, or T.
119. The method of embodiment 117, wherein the Cas protein recognizes a PAM that is not NGG, wherein N can be A, C, G, or T.
120. The method of any one of embodiments 99-119, wherein the sgRNA comprises a guide nucleic acid sequence that is capable of binding to or recognizing the target nucleic acid or its complement.
121. The method of embodiment 120, wherein the guide nucleic acid sequence is identical to the target nucleic acid sequence or its complement, when a U in the guide nucleic acid sequence is equivalent to a T in the target nucleic acid sequence or its complement.
122. The method of embodiment 120, wherein the guide nucleic acid sequence is not identical to the target nucleic acid sequence or its complement, when a U in the guide nucleic acid sequence is equivalent to a T in the target nucleic acid sequence or its complement.
123. The method of embodiment 120 or 123, wherein there are one or more mismatched bases between the guide nucleic acid sequence and the target nucleic acid sequence or its complement.
124. The method of embodiment 123, wherein the one or more mismatches occur at a position or positions in the target nucleic acid sequence that are located within the first 8, 9, 10, 11, or 12 bases beginning from the 5' end of the target nucleic acid sequence.
125. The method of any one of embodiments 99-124, wherein the target nucleic acid sequence is:
(i) native to the genome,
(ii) a foreign nucleic acid sequence that was introduced and stably incorporated into the genome of a plant from which the plant tissue was obtained,
(iii) a foreign nucleic acid sequence that was introduced and stably incorporated into the genome of a progenitor of the plant from which the plant tissue was obtained, or
(iv) co-delivered with the Cas gene and the sgR A.
126. The method of any one of embodiments 99-125, wherein monitoring the plant tissue for a modification of the genome within the target site comprises determining the nucleotide sequence of all or a part of the target nucleic acid sequence.
127. The method of any one of embodiments 99-126, wherein the target nucleic acid sequence comprises a restriction enzyme recognition sequence.
128. The method of embodiment 127, wherein step (c) comprises:
extracting genomic DNA from the plant tissue contacted by the first and second Agrobacterium strains:
digesting the genomic DNA with a restriction enzyme that recognizes the restriction enzyme recognition sequence;
amplifying the digested DNA by polymerase chain reaction using primers designed to amplify across the target nucleic acid sequence so as to produce PCR products comprising one or more modifications within target nucleic sequence;
cloning the PCR products; and
determining the nucleic acid sequence of the cloned PCR products.
129. The method of embodiment 127 or 128, wherein the restriction enzyme recognition sequence is a Mlyl restriction enzyme recognition sequence.
130. The method of embodiment 128 or 129, wherein the DNA is extracted from plant tissue harvested at about 1-48 hours after the later of the contacting steps.
131. The method of embodiment 128 or 129, wherein the DNA is extracted from leaves harvested at about 6, 12, 18, 24, 30, 36, 42, or 48 hours after the later of the contacting steps.
132. The method of any one of embodiments 1-55, wherein the method comprises producing at least one plant cell comprising a Cas protein and at least two engineered, sgRNAs that are each capable of recognizing a different target nucleic acid sequence.
133. A method for modifying the genome of a plant, plant part, or plant cell, the method comprising producing at least one plant cell comprising a Cas protein and at least two engineered, sgRNAs that are each capable of recognizing a different target nucleic acid sequence, wherein the plant, plant part, plant cell, or descent thereof, comprises in its genome the target nucleic acid sequences.
134. The method of embodiment 133, wherein a first sgRNA is capable of recognizing a first target sequence and a second sgRNA is capable of recognizing a second target sequence.
135. The method of embodiment 134, wherein the first target sequence and the second target sequence are on the same chromosome.
136. The method of embodiment 134 or 135, wherein the first target sequence and the second target sequence are within the same gene.
137. The method of any one of embodiments 134-136, wherein the first target sequence and the second target sequence are separated by at least 10, 20, 30, 40, 50, 75, 100, 150, 200, 300, 400, 500, 1000, 1500, 2000, 3000 or more nucleotides.
138. The method of any one of embodiments 134-137, wherein the modification comprises the deletion of at least one base pair in the genome of the plant, plant part, plant cell, or descendant thereof.
139. The method embodiment 138, wherein the deletion occurs between the first target sequence and the second target sequence.
140. The method of any one of embodiments 134-139, wherein at least one of the Cas protein, the first sgR A, and the second sgR A are transiently expressed in the plant cell.
It is an object of the present invention to provide new methods for targeted mutagenesis and genome engineering that are less complicated and costly than existing methods such as, for example, those methods involving zinc finger nucleases (ZFNs) and TAL effector nucleases (TALENs).
The present invention provides new methods for targeted mutagenesis and genome engineering that do not depend on the use of a ZFNs or a TALENs but instead involve the use of a Cas protein and engineered, single guide RNA (sgRNA) that specifies a targeted nucleic acid sequence . In the examples below, the Cas protein, Cas9 is used for purposes of illustration. The present invention is not limited to the use of Cas9. Mutants and derivatives of Cas9 as well as other Cas proteins can be used in the methods disclosed herein. Preferably, such other Cas proteins are able to recognize a target nucleic acid sequence when in a plant cell in the presence of an sgRNA that is engineered for recognition of the target sequence. In some embodiments of the invention, the Cas proteins will additional comprise a double-strand endonuclease activity or a single strand endonuclease activity (i.e., a nickase activity). In other embodiments, the Cas protein will not comprise a double-strand endonuclease activity and/or single strand endonuclease activity. In yet other embodiments, a Cas protein domain will be operably linked with at least one additional protein domain to form a fusion protein. Such a fusion protein a can comprise a double-strand endonuclease activity or a single strand endonuclease activity or no endonuclease at all but instead can comprise an additional activity that is due to the additional domain.
Fragments and variants of the disclosed nucleic acid molecules and proteins encoded thereby are also encompassed by the present invention. Such fragments and variants of the disclosed nucleic acid molecules and proteins include, for example, nucleic acid molecules encoding mutants and dervivaties of the Cas9 protein and the
proteins encoded thereby. The nucleic acid and amino acid sequences of Cas9 are disclosed elsewhere herein or otherwise known in the art. By "fragment" is intended a portion of the nucleic acid molecule or a portion of the amino acid sequence and hence protein encoded thereby. Fragments of a nucleic acid molecule comprising coding sequences may encode protein fragments that retain biological activity of the native protein and hence DNA recognition or binding activity to a target DNA sequence as herein described. Alternatively, fragments of a nucleic acid molecule that are useful as hybridization probes generally do not encode proteins that retain biological activity or do not retain promoter activity. Thus, fragments of a nucleic acid molecule may range from at least about 20 nucleotides, about 50 nucleotides, about 100 nucleotides, and up to the full-length nucleic acid molecule of the invention.
"Variants" is intended to mean substantially similar sequences. For nucleic acid molecules, a variant comprises a nucleic acid molecule having deletions (i.e., truncations) at the 5' and/or 3' end; deletion and/or addition of one or more nucleotides at one or more internal sites in the native polynucleotide; and/or substitution of one or more nucleotides at one or more sites in the native polynucleotide. As used herein, a "native" nucleic acid molecule or polypeptide comprises a naturally occurring nucleiotide sequence or amino acid sequence, respectively. For nucleic acid molecules, conservative variants include those sequences that, because of the degeneracy of the genetic code, encode the amino acid sequence of one of the polypeptides of the invention. Variant nucleic acid molecules also include synthetically derived nucleic acid molucules, such as those generated, for example, by using site-directed mutagenesis but which still encode a protein of the invention. Generally, variants of a particular nucleic acid molecule of the invention will have at least about 70%, 75%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%), 96%o, 97%), 98%>, 99% or more sequence identity to that particular polynucleotide as determined by sequence alignment programs and parameters as described elsewhere herein.
Variants of a particular nucleic acid molecule of the invention (i.e., the reference DNA sequence) can also be evaluated by comparison of the percent sequence identity between the polypeptide encoded by a variant nucleic acid molecule and the polypeptide encoded by the reference nucleic acid molecule. Percent sequence identity between any two polypeptides can be calculated using sequence alignment programs and parameters described elsewhere herein. Where any given pair of nucleic acid molecule of the
invention is evaluated by comparison of the percent sequence identity shared by the two polypeptides that they encode, the percent sequence identity between the two encoded polypeptides is at least about 70%, 75%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or more sequence identity.
"Variant" protein is intended to mean a protein derived from the native protein by deletion (so-called truncation) of one or more amino acids at the N-terminal and/or C- terminal end of the native protein; deletion and/or addition of one or more amino acids at one or more internal sites in the native protein; or substitution of one or more amino acids at one or more sites in the native protein. Variant proteins encompassed by the present invention are biologically active, that is they continue to possess the desired biological activity of the native protein as described herein. Such variants may result from, for example, genetic polymorphism or from human manipulation. Biologically active variants of a protein of the invention will have at least about 70%, 75%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or more sequence identity to the amino acid sequence for the native protein as determined by sequence alignment programs and parameters described elsewhere herein. A biologically active variant of a protein of the invention may differ from that protein by as few as 1-15 amino acid residues, as few as 1-10, such as 6-10, as few as 5, as few as 4, 3, 2, or even 1 amino acid residue.
The proteins or polypeptides of the invention may be altered in various ways including amino acid substitutions, deletions, truncations, and insertions. Methods for such manipulations are generally known in the art. For example, amino acid sequence variants and fragments of the proteins can be prepared by mutations in the DNA.
Techniques to specifically modify DNA sequences in order to obtain a specified codon for a specific amino acid are well known in the art. Methods for mutagenesis and polynucleotide alterations are also well known in the art. See, for example, Kunkel (1985) Proc. Natl. Acad. Sci. USA 82:488-492; Kunkel et al. (1987) Methods in
Enzymol. 154:367-382; U.S. Patent No. 4,873,192; Walker and Gaastra, eds. (1983) Techniques in Molecular Biology (MacMillan Publishing Company, New York) and the references cited therein. Guidance as to appropriate amino acid substitutions that do not affect biological activity of the protein of interest may be found in the model of Dayhoff et al. (1978) Atlas of Protein Sequence and Structure (Natl. Biomed. Res. Found.,
Washington, D.C.), herein incorporated by reference. Conservative substitutions, such as exchanging one amino acid with another having similar properties, may be optimal .
The deletions, insertions, and substitutions of the protein sequences encompassed herein are not expected to produce radical changes in the characteristics of the protein. However, when it is difficult to predict the exact effect of the substitution, deletion, or insertion in advance of doing so, one skilled in the art will appreciate that the effect will be evaluated by routine screening assays as described elsewhere herein or known in the art.
Variant nucleic acid molecule and proteins also encompass sequences and proteins derived from a mutagenic and recombinogenic procedure such as DNA shuffling. Strategies for such DNA shuffling are known in the art. See, for example, Stemmer (1994) Proc. Natl. Acad. Sci. USA 91 : 10747-10751; Stemmer (1994) Nature 370:389-391; Crameri et al. (1997) Nature Biotech. 15:436-438; Moore et al. (1997) J. Mol. Biol. 272:336-347; Zhang et al. (1997) Proc. Natl. Acad. Sci. USA 94:4504-4509; Crameri et al. (1998) Nature 391 :288-291; and U.S. Patent Nos. 5,605,793 and
5,837,458.
In a PCR approaches, oligonucleotide primers can be designed for use in PCR reactions to amplify corresponding nucleic acid molecules from cDNA or genomic DNA extracted from any organism of interest. Methods for designing PCR primers and PCR cloning are generally known in the art and are disclosed in Sambrook et al. (1989)
Molecular Cloning: A Laboratory Manual (2d ed., Cold Spring Harbor Laboratory Press, Plainview, New York). See also Innis et al., eds. (1990) PCR Protocols: A Guide to Methods and Applications (Academic Press, New York); Innis and Gelfand, eds. (1995) PCR Strategies (Academic Press, New York); and Innis and Gelfand, eds. (1999) PCR Methods Manual (Academic Press, New York). Known methods of PCR include, but are not limited to, methods using paired primers, nested primers, single specific primers, degenerate primers, gene-specific primers, vector-specific primers, partially-mismatched primers, and the like.
It is recognized that the nucleic acid molecules and proteins of the invention encompass polynucleotide molecules and proteins comprising a nucleotide or an amino acid sequence that is sufficiently identical to the nucleotide sequences or to the amino acid sequence disclosed herein. The term "sufficiently identical" is used herein to refer to a first amino acid or nucleotide sequence that contains a sufficient or minimum
number of identical or equivalent (e.g., with a similar side chain) amino acid residues or nucleotides to a second amino acid or nucleotide sequence such that the first and second amino acid or nucleotide sequences have a common structural domain and/or common functional activity. For example, amino acid or nucleotide sequences that contain a common structural domain having at least about 70% identity, preferably 75% identity, more preferably 85%, 90%, 95%, 96%, 97%, 98% or 99% identity are defined herein as sufficiently identical.
To determine the percent identity of two amino acid sequences or of two nucleic acids, the sequences are aligned for optimal comparison purposes. The percent identity between the two sequences is a function of the number of identical positions shared by the sequences (i.e., percent identity = number of identical positions/total number of positions (e.g., overlapping positions) x 100). In one embodiment, the two sequences are the same length. The percent identity between two sequences can be determined using techniques similar to those described below, with or without allowing gaps. In calculating percent identity, typically exact matches are counted.
The determination of percent identity between two sequences can be
accomplished using a mathematical algorithm. A preferred, nonlimiting example of a mathematical algorithm utilized for the comparison of two sequences is the algorithm of Karlin and Altschul (1990) Proc. Natl. Acad. Sci. USA 87:2264, modified as in Karlin and Altschul (1993) Proc. Natl. Acad. Sci. USA 90:5873-5877. Such an algorithm is incorporated into the NBLAST and XBLAST programs of Altschul et al. (1990) J. Mol. Biol. 215:403. BLAST nucleotide searches can be performed with the NBLAST program, score = 100, wordlength = 12, to obtain nucleotide sequences homologous to the polynucleotide molecules of the invention. BLAST protein searches can be performed with the XBLAST program, score = 50, wordlength = 3, to obtain amino acid sequences homologous to protein molecules of the invention. To obtain gapped alignments for comparison purposes, Gapped BLAST can be utilized as described in Altschul et al. (1997) Nucleic Acids Res. 25:3389. Alternatively, PSI-Blast can be used to perform an iterated search that detects distant relationships between molecules. See Altschul et al. (1997) supra. When utilizing BLAST, Gapped BLAST, and PSI-Blast programs, the default parameters of the respective programs (e.g., XBLAST and
NBLAST) can be used. See http://www.ncbi.nlm.nih.gov. Another preferred, non- limiting example of a mathematical algorithm utilized for the comparison of sequences is
the algorithm of Myers and Miller (1988) CABIOS 4: 11-17. Such an algorithm is incorporated into the ALIGN program (version 2.0), which is part of the GCG sequence alignment software package. When utilizing the ALIGN program for comparing amino acid sequences, a PAM120 weight residue table, a gap length penalty of 12, and a gap penalty of 4 can be used. Alignment may also be performed manually by inspection. Unless otherwise stated, sequence identity/similarity values provided herein refer to the value obtained using the full-length sequences of the invention and using multiple alignment by mean of the algorithm Clustal W (Nucleic Acid Research, 22(22):4673- 4680, 1994) using the program AlignX included in the software package Vector NTI Suite Version 7 (InforMax, Inc., Bethesda, MD, USA) using the default parameters; or any equivalent program thereof. By "equivalent program" is intended any sequence comparison program that, for any two sequences in question, generates an alignment having identical nucleotide or amino acid residue matches and an identical percent sequence identity when compared to the corresponding alignment generated by
CLUSTALW (Version 1.83) using default parameters (available at the European
Bioinformatics Institute website: http://www.ebi . ac .uk/Too 1 s/c lustalw/inde . html
The methods of the invention involve introducing a polypeptide or
polynucleotide into a plant. "Introducing" is intended to mean presenting to the plant the polynucleotide or polypeptide in such a manner that the sequence gains access to the interior of a cell of the plant. The methods of the invention do not depend on a particular method for introducing a sequence into a plant, only that the polynucleotide or polypeptides gains access to the interior of at least one cell of the plant. Methods for introducing polynucleotide or polypeptides into plants are known in the art including, but not limited to, stable transformation methods, transient transformation methods, and virus-mediated methods.
"Stable transformation" is intended to mean that the nucleotide construct introduced into a plant integrates into the genome of the plant and is capable of being inherited by the progeny thereof. "Transient transformation" is intended to mean that a polynucleotide is introduced into the plant and does not integrate into the genome of the plant or a polypeptide is introduced into a plant.
Transformation protocols as well as protocols for introducing polypeptides or polynucleotide sequences into plants may vary depending on the type of plant or plant cell, i.e., monocot or dicot, targeted for transformation. Suitable methods of introducing
polypeptides and polynucleotides into plant cells include microinjection (Crossway et al.
(1986) Biotechniques 4:320-334), electroporation (Riggs et al. (1986) Proc. Natl. Acad.
Sci. USA 83:5602-5606, Agrobacterium-mediatGd transformation (U.S. Patent No.
5,563,055 and U.S. Patent No. 5,981,840), direct gene transfer (Paszkowski et al. (1984) EMBO J. 3:2717-2722), and ballistic particle acceleration (see, for example, U.S. Patent
Nos. 4,945,050; U.S. Patent No. 5,879,918; U.S. Patent No. 5,886,244; and, 5,932,782;
Tomes et al. (1995) in Plant Cell, Tissue, and Organ Culture: Fundamental Methods, ed.
Gamborg and Phillips (Springer-Verlag, Berlin); McCabe et al. (1988) Biotechnology
6:923-926); and Lecl transformation (WO 00/28058). Also see Weissinger et al. (1988) Ann. Rev. Genet. 22:421-477; Sanford et al. (1987) Particulate Science and Technology
5:27-37 (onion); Christou et al. (1988) Plant Physiol. 87:671-674 (soybean); McCabe et al. (1988) Bio/Technology 6:923-926 (soybean); Finer and McMullen (1991) In Vitro
Cell Dev. Biol. 27P: 175-182 (soybean); Singh et al. (1998) Theor. Appl. Genet. 96:319-
324 (soybean); Datta et al. (1990) Biotechnology 8:736-740 (rice); Klein et al. (1988) Proc. Natl. Acad. Sci. USA 85:4305-4309 (maize); Klein et al. (1988) Biotechnology
6:559-563 (maize); U.S. Patent Nos. 5,240,855; 5,322,783; and, 5,324,646; Klein et al.
(1988) Plant Physiol. 91 :440-444 (maize); Fromm et al. (1990) Biotechnology 8:833-839
(maize); Hooykaas-Van Slogteren et al. (1984) Nature (London) 311 :763-764; U.S.
Patent No. 5,736,369 (cereals); Bytebier et al. (1987) Proc. Natl. Acad. Sci. USA
84:5345-5349 (Liliaceae); De Wet et al. (1985) in The Experimental Manipulation of
Ovule Tissues, ed. Chapman et al. (Longman, New York), pp. 197-209 (pollen);
Kaeppler et al. (1990) Plant Cell Reports 9:415-418 and Kaeppler et al. (1992) Theor.
Appl. Genet. 84:560-566 (whisker-mediated transformation); D'Halluin et al. (1992)
Plant Cell 4: 1495-1505 (electroporation); Li et al. (1993) Plant Cell Reports 12:250-255 and Christou and Ford (1995) Annals of Botany 75:407-413 (rice); Osjoda et al. (1996)
Nature Biotechnology 14:745-750 (maize via Agrobacterium tumefaciens); all of which are herein incorporated by reference.
In specific embodiments, the nucleic acid molecules and constructs of the invention can be provided to a plant using a variety of transient transformation methods. Such transient transformation methods include, but are not limited to, the introduction of a Cas protein and/or one or more sgRNAs directly into the plant or the introduction into the plant of a Cas protein transcript and/or one or more transcripts encoding sgRNAs.
Such methods include, for example, microinjection or particle bombardment. See, for
example, Crossway et al. (1986) Mol Gen. Genet. 202: 179-185; Nomura et al. (1986) Plant Sci. 44:53-58; Hepler et al. (1994) Proc. Natl. Acad. Sci. 91 : 2176-2180 and Hush et al. (1994) The Journal of Cell Science 707:775-784, all of which are herein incorporated by reference. Alternatively, the polynucleotide can be transiently transformed into the plant using techniques known in the art. Such techniques include viral vector system and the precipitation of the polynucleotide in a manner that precludes subsequent release of the DNA. Thus, the transcription from the particle-bound DNA can occur, but the frequency with which its released to become integrated into the genome is greatly reduced. Such methods include the use particles coated with polyethylimine (PEI; Sigma #P3143).
In other embodiments, the polynucleotide of the invention may be introduced into plants by contacting plants with a virus or viral nucleic acids. Generally, such methods involve incorporating a nucleotide construct of the invention within a viral DNA or RNA molecule. It is recognized that the an nucleic acid molecule of the invention may be initially synthesized as part of a viral polyprotein, which later may be processed by proteolysis in vivo or in vitro to produce the desired recombinant protein. Further, it is recognized that promoters of the invention also encompass promoters utilized for transcription by viral RNA polymerases. Methods for introducing polynucleotides into plants and expressing a protein encoded therein, involving viral DNA or RNA molecules, are known in the art. See, for example, U.S. Patent Nos. 5,889,191,
5,889,190, 5,866,785, 5,589,367, 5,316,931, and Porta et al. (1996) Molecular
Biotechnology 5:209-221; herein incorporated by reference.
Methods are known in the art for the targeted insertion of a polynucleotide at a specific location in the plant genome. In one embodiment, the insertion of the polynucleotide at a desired genomic location is achieved using a site-specific
recombination system. See, for example, W099/25821, W099/25854, WO99/25840, W099/25855, and W099/25853, all of which are herein incorporated by reference. Briefly, the polynucleotide of the invention can be contained in transfer cassette flanked by two non-recombinogenic recombination sites. The transfer cassette is introduced into a plant having stably incorporated into its genome a target site which is flanked by two non-recombinogenic recombination sites that correspond to the sites of the transfer cassette. An appropriate recombinase is provided and the transfer cassette is integrated
at the target site. The polynucleotide of interest is thereby integrated at a specific chromosomal position in the plant genome.
The cells that have been transformed may be grown into plants in accordance with conventional ways. See, for example, McCormick et al. (1986) Plant Cell Reports 5:81-84. These plants may then be grown, and either pollinated with the same transformed strain or different strains, and the resulting progeny having constitutive expression of the desired phenotypic characteristic identified. Two or more generations may be grown to ensure that expression of the desired phenotypic characteristic is stably maintained and inherited and then seeds harvested to ensure expression of the desired phenotypic characteristic has been achieved. In this manner, the present invention provides transformed seed (also referred to as "transgenic seed") having a polynucleotide of the invention, for example, an expression cassette of the invention, stably incorporated into their genome.
As used herein, the term plant also includes plant cells, plant protoplasts, plant cell tissue cultures from which plants can be regenerated, plant calli, plant clumps, and plant cells that are intact in plants or parts of plants such as embryos, pollen, ovules, seeds, leaves, flowers, branches, fruit, kernels, ears, cobs, husks, stalks, roots, root tips, anthers, and the like. Grain is intended to mean the mature seed produced by
commercial growers for purposes other than growing or reproducing the species.
Progeny, variants, and mutants of the regenerated plants are also included within the scope of the invention, provided that these parts comprise the introduced
polynucleotides.
The present invention may be used for any plant species of interest, including, but not limited to, monocots and dicots. Examples of plant species of interest include, but are not limited to, corn (Zea mays), Brassica sp. (e.g., B. napus, B. rapa, B.juncea), particularly those Brassica species useful as sources of seed oil, alfalfa (Medicago sativa), rice (Oryza sativa), rye (Secale cereale), sorghum (Sorghum bicolor, Sorghum vulgare), millet (e.g., pearl millet (Pennisetum glaucum), proso millet (Panicum miliaceum), foxtail millet (Setaria italica), finger millet (Eleusine coracana)), sunflower (Helianthus annuus), safflower (Carthamus tinctorius), wheat (Triticum aestivum), soybean (Glycine max), tobacco (Nicotiana tabacum), potato (Solanum tuberosum), peanuts (Arachis hypogaea), cotton (Gossypium barbadense, Gossypium hirsutum), sweet potato (Ipomoea batatus), cassava (Manihot esculenta), coffee (Coffea spp.), coconut (Cocos nucifera), pineapple
(Ananas comosus), citrus trees (Citrus spp.), cocoa (Theobroma cacao), tea (Camellia sinensis), banana (Musa spp.), avocado (Persea americana), fig (Ficus casica), guava (Psidium guajava), mango (Mangifera indica), olive (Olea europaea), papaya (Carica papaya), cashew (Anacardium occidentale), macadamia (Macadamia integrifolid), almond (Prunus amygdalus), sugar beets (Beta vulgaris), sugarcane (Saccharum spp.), oats, barley, vegetables, ornamentals, and conifers.
Vegetables include tomatoes (Lycopersicon esculentum), lettuce (e.g., Lactuca sativa), green beans (Phaseolus vulgaris), lima beans (Phaseolus limensis), peas (Lathyrus spp.), and members of the genus Cucumis such as cucumber (C sativus), cantaloupe (C. cantalupensis), and musk melon (C melo). Ornamentals include azalea (Rhododendron spp.), hydrangea (Macrophylla hydrangea), hibiscus (Hibiscus rosasanensis), roses (Rosa spp.), tulips (Tulipa spp.), daffodils (Narcissus spp.), petunias (Petunia hybrida), carnation (Dianthus caryophyllus), poinsettia (Euphorbia pulcherrima), and chrysanthemum.
Conifers that may be employed in practicing the present invention include, for example, pines such as loblolly pine (Pinus taeda), slash pine (Pinus elliotii), ponderosa pine (Pinus ponderosa), lodgepole pine (Pinus contorta), and Monterey pine (Pinus radiata); Douglas-fir (Pseudotsuga menziesii); Western hemlock (Tsuga canadensis); Sitka spruce (Picea glauca); redwood (Sequoia sempervirens); true firs such as silver fir (Abies amabilis) and balsam fir (Abies balsamea); and cedars such as Western red cedar (Thuja plicata) and Alaska yellow-cedar (Chamaecyparis nootkatensis).
A "subject plant or plant cell" is one in which genetic alteration, such as transformation, has been effected as to a gene of interest, or is a plant or plant cell which is descended from a plant or cell so altered and which comprises the alteration. A "control" or "control plant" or "control plant cell" provides a reference point for measuring changes in phenotype of the subject plant or plant cell.
A control plant or plant cell may comprise, for example: (a) a wild-type plant or cell, i.e., of the same genotype as the starting material for the genetic alteration which resulted in the subject plant or cell; (b) a plant or plant cell of the same genotype as the starting material but which has been transformed with a null construct (i.e. with a construct which has no known effect on the trait of interest, such as a construct comprising a marker gene); (c) a plant or plant cell which is a non-transformed segregant among progeny of a subject plant or plant cell; (d) a plant or plant cell genetically identical to the subject plant or plant cell but which is not exposed to
conditions or stimuli that would induce expression of the gene of interest; or (e) the subject plant or plant cell itself, under conditions in which the gene of interest is not expressed.
The following examples are offered by way of illustration and not by way of limitation.
EXAMPLE 1
Targeted Mutagenesis of Nicotiana benthamiana To test the potential of the Cas9 system to induce gene knock-outs (i.e., loss-of- function mutations) in plants, we took advantage of Agrobacterium turne/adens-mediated transient expression assays (agroinfiltration) to co-express a Cas9 variant with a eukaryotic nuclear localization signal (nls) and an sgRNA in the model plant Nicotiana benthamiana4. First, we constructed a GFP-tagged version of Cas9 using the clone described in Mali et al.5. We expressed GFP-Cas9 in N. benthamiana leaf tissue using standard agroinfiltration protocols6 and observed a clear nuclear localization (FIG. 3) consistent with the nuclear localization previously observed in human cells7. We then generated an sgRNA with the guide sequence matching a 19 bp region within the phytoene desaturase (PDS) gene in the Nicotiana benthamiana (FIG. la). The sgRNA was placed under an Arabidopsis U6 promoter (FIG. 4). Both GFP-Cas9 and sgRNA were co-expressed in N. benthamiana leaf tissue using A. tumefaciens as a vector. The tissue was harvested 2 days later and DNA was extracted. In order to easily detect sgRNA-guided, Cas9-induced mutations at the PDS locus, we used the restriction enzyme site loss method . As the target sequence within the PDS gene overlaps with an Mlyl restriction site, we digested the genomic DNA with Mlyl and then performed a polymerase chain reaction (PCR) with primers flanking the target site. By doing so we greatly reduced unaltered wild type DNA in the sample and enriched for DNA molecules carrying mutations that remove the Mlyl site. As illustrated in FIG. lb, the presence of both Cas9 and the sgRNA resulted in increased levels of the PCR product (lane 1) compared to negative control treatments (lanes 2-3). Non-digested N benthamiana genomic DNA was used as a positive control (lane 4). The assay was robust and reproducible because we detected Mlyl-resistant amplicons in three additional independent experiments using different plants (FIG. 5 and FIG. 6). The PCR products
from FIG. lb lanes 1 and 4 were cloned into a high copy vector and individual clones sequenced. Sequence analysis of 20 clones derived from the PCR product in lane 1 revealed the presence of indels in 17 of them. The indels can be grouped into nine different types ranging from 1-9 bp deletions to 1 bp insertions (FIG. lc and FIG. 7). All recovered indels abolish the Mlyl restriction site within the target region. Sequences of the 8 clones derived from the control PCR product shown in lane 4 were all wild type.
To estimate the efficiency of targeted mutagenesis, we amplified non-digested genomic DNA from Cas9/sgRNA expressing N. benthamiana leaves and negative controls, and subsequently digested the amplicons with Mlyl and subjected them to gel electrophoresis. We then measured the intensity of the uncut band relative to the intensity of all detectable bands in a gel lane as described in Qi et al. (FIG. 2 and FIG. 8). We estimated the mutation rate to be in the range of 1.8 to 2.4% (2.1% average) based on four independent experiments.
Given that the target sequence is 19 bp, the Cas9/sgRNA system may not be as specific as TALEN-induced mutagenesis. We examined 18 off-target sites in the N. benthamiana genome that have 14 to 16 out of 19 bp identity to the targeted PDS sequence and overlap a Mlyl restriction site. None of 18 amplicons showed unambiguous evidence of Cas9/sgRNA dependent Mlyl restriction site loss as observed with the PDS target sequence (Table 1 and FIG. 9). We therefore conclude that any off-target nuclease activity is much lower than activity against the specified target.
These data clearly indicate that Cas9 and an engineered sgRNA can direct DNA breaks at defined chromosomal locations in plants. The rapid and robust transient assay we developed will enable plant-specific optimization of the Cas9 system. Relative to other methods of plant genome engineering, the Cas9 system has the potential to simplify the process of plant genome engineering and editing since only a short fragment in the sgRNA needs to be designed to target a new locus. This creates a valuable new tool for plant biologists and breeders, and hastens the prospects of achieving routine targeted genome engineering for basic and applied science.
Materials and Methods
Construction of Cas9 and sgRNA expressing plasmids
Cas9 was PCR-amplified with primers Cas9GWF and Cas9STR using the clone described in Mali et al.5 as a template. The resulting PCR product was cloned into the pENTR/D-TOPO vector (Life Technologies). The entry clone was subsequently recombined into the GW-compatible destination vector pK7WGF29 using the LR clonase (Life Technologies) to produce the 35Sp::GFPCas9 construct.
The Arabidopsis U6 promoter used in this study is the consensus sequence of the 3 U6 promoter variants present in the Arabidopsis genome10. The AtU6p clone was synthesised with Genescript (FIG. 4). The promoter was PCR-amplified with primers U6p_GGAG_f and U6p_CAAT_r and cloned into a level 0 vector11 via the Bbsl cut- ligation using the Golden Gate (GG) cloning method. The sgRNA was PCR-amplified using primers PDS gRNAl Bsalf and gRNA AGCG Bsalr (PDS locus) or primers gRNA T l GFP Bsalf and gRNA AGCG Bsalr (GFP) using the plasmid
gRNA GFP Tl described in Mali et al.1 as a template. The resulting PCR product was cloned into the pICH86966 vector (kindly provided by S. Marillonnet, based on the modular cloning system described in Weber et al.n under the AtU6p via the Bsal cut- ligation using the GG method. Both Cas9 and gRNA GFP Tl plasmids5 were obtained from Addgene.
Transient gene expression in N. benthamiana and Cas9 intracellular localisation
Transient expression was performed using the AGL1 strain of Agrobacterium
12
tumefaciens as described in Bos et al. . GFP-Cas9 was localised in the leaf tissue of N. benthamiana 2 days post infiltration using the Leica SP5 confocal microscope in accordance with manufacturer's instructions.
Detection of Cas9-induced indels in plant genomic DNA
GFP-Cas9 and sgRNA carrying the guide sequence matching a site within the PDS gene of N. benthamiana were transiently co-expressed in the N. benthamiana leaf tissue as follows. An Agrobacterium strain carrying a nucleic acid construct of the expression of GFP-Cas9 in plant cells and an Agrobacterium strain carrying a nucleic acid construct of the expression of the sgRNA in plant cells were separately inoculated
into L medium containing respective antibiotics for plasmid selection and grown overnight at 28°C. The next day, each of the resulting cultures was centrifuged at 3500 rpm for 10 min, and the resulting pellets of Agrobacteria were separately resuspended in 10 mM MgCl2. The resuspended Agrobacteria were diluted with the 10 mM MgCl2 to OD600=0.5 and then the resuspended Agrobacteria carrying the GFP-Cas9 construct were mixed with the resuspended Agrobacteria carrying the sgR A construct were mixed at a ratio 1 : 1 (v/v). Leaves of 4-week-old N. benthamiana plants were infiltrated with pre-mixed Agrobacteria, and after 48 hours, the leaf tissue was harvested.
The tissue was harvested at 2 days post infiltration and the genomic DNA extracted using the DNeasy Plant Mini kit (Qiagen). 100 ng of the genomic DNA was then digested overnight with Mlyl restriction enzyme to reduce greatly the background of the non-modified (wild-type) DNA. Upon digesting the reaction mix was desalted using Sepharose CL-6B (Sigma) and then added as a template into a PCR reaction performed with primers PDS MlylF and PDS MlylR with flank the sgRNA target nucleic acid sequence and Phusion DNA polymerase (New England Biolabs). The resulting PCR band was cloned a high copy vector (pCRTM-Blunt II-TOPO vector, Life Technologies, Inc.). Plasmids from individual Escherichia coli colonies were sequenced using standard primers Ml 3 forward and Ml 3 reverse. Amplicons with mutated Mlyl sites were recovered only when the Cas9 and sgRNA constructs were co-expressed (FIG. 1 and FIG. 5) but weak bands can be occasionally observed in the negative controls probably due to incomplete digestion. To ensure that the amplicons indeed contain DNA fragments resistant to Mlyl digestion, we performed a partial digestion of the genomic DNA with Mlyl followed by a second Mlyl digestion of the amplicons as shown in FIG. 6.
Measurements of the mutation rate
In order to estimate the rate of Cas9/sgRNA-induced mutations in the PDS locus we used the method described by Qi et al. . The PCR product amplified on the non- digested genomic DNA as a template was digested with Mlyl and run on a gel. The intensity of non-saturated bands was quantified using the ImageJ software
(h tt ://rsb . info .nih. gov/ij) . The mutation rate was calculated by dividing the intensity of the uncut band by the total intensity of all bands in a given lane. Analysis of potential off-targets
In order to select potential off-targets, we searched the N. benthamiana genome database using the NCBI BLASTN online tool against the 19 bp sgRNA target site within the PDS gene (CCGTTAATTTGAGAGTCCA). We then picked 44 potential off- targets with varying numbers of bases (14-16) matching the 19 bp PDS target sequence. None of the matching N. benthamiana sequences had more than 16 out of 19 bp matches to the PDS target sequence. All of the off-targets selected had the Mlyl site present in them. Primers were designed to amplify the potential off-target loci across the off-target site. Out of the 44 loci amplified, 18 gave a PCR band of expected size without nonspecific bands. We then proceeded further with these 18 loci (Table 1). First, we tested if we could amplify the loci using Mlyl-digested genomic DNA extracted from the leaf tissue expressing Cas9+sgRNA(PDS) or Cas9 alone. For 12 out of 18 loci it was possible to amplify a PCR product using the Mlyl-digested genomic DNA and for the 1 out of 12 loci a weak band was observed only in the case of the Cas9+sgRNA(PDS) sample indicating potential off-target activity (FIG. 9c). Nevertheless, in all 18 cases the PCR products could be digested by Mlyl (FIG. 9a-c), while the PCR product from the positive control (PDS) showed clear resistance to Mlyl digestion (FIG. 9d).
Table 1. Summary of the off-target analyses. The Mlyl restriction site is shown in blue.
Table 2. Primers used in the Examples.
EXAMPLE 2
Targeted, Large Chromosomal Deletions in Nicotiana benthamiana and Tomato
Targeted, large chromosomal deletions were generated by targeting two adjacent sequences within the PDS locus of Nicotiana benthamiana using transient expression. To make the large deletions in PDS locus, two sgRNAs were transiently expressed resulted in chromosomal deletion events. The sgRNAs were designed for target sequences on each side of the region in the PDS locus that was intended to be deleted. A schematic representation of the PDS locus and targets is shown in FIG. 10A. FIG. 10B shows the detection of deletion mutations using the AFLP analysis. The deletions obtained are shown in FIG. IOC. The nucleic acid sequence of the sgRNA constructs are shown in FIG. 12.
Targeted, large chromosomal deletions were generated by targeting two adjacent sequences within the Mlol gene of tomato using transient expression. To make the large deletions in Mlol gene, sgRNAs were transiently expressed resulted in chromosomal deletion events. The sgRNAs were designed for target sequences on each side of the regions in the Mlol gene that was intended to be deleted as shown (FIG. 11 A). FIG. 1 IB
shows the detection of deletion mutations by the AFLP analysis and FIG. 1 1C shows the results from the genotyping of TO transgenic tomato plants. The plants were genotyped for the presence of CRISPR/Cas-induced deletions.
The results demonstrate applicability of the CRISPR/Cas system for purposes of creating large chromosomal deletions in plants. Creating a large deletion by targeting with 2 sgRNAs provides a more convenient way to knock-out a gene as such deletion event can be easily detected by the AFLP (PCR band shift) assay. Constructs can be quickly tested using the transient expression systems in crops amenable to transient expression (e.g. tomato) before proceeding with generation of stable transgenic lines. In addition to generating single gene-knockouts, ability to create large chromosomal deletions provides an opportunity for deleting whole gene clusters as well as deleting certain promoter elements or protein domains.
Materials and Methods
Plasmid construction
In the case of the lane 2 (Figure IB), pK7WGF2::Cas9 and
pICH86966::AtU6p::sgRNAl_ D5* [1] were co-expressed with
pICH86966::AtU6p::sgRNA2_ NbPDS in N. benthamiana using the standard agroinfiltration protocol. In the case of the lane 4 (Figure IB), Cas9 and both sgRNAl_A/¾ ZXS' and sgRNA2_A/¾ ZXS' were expressed in N. benthamiana from the single construct.
In order to use the human codon optimised Cas95 in the Golden Gate cloning system11, all Bsal and Bbsl sites had to be removed from its sequence, while preserving the amino acid composition of the protein, in a process called "domestication. The "domesticated" Cas9 was cloned into a level 1 vector pICH47742n.
N. benthamiana PDS locus
NbPDS sgRNAs 1 and 2:
pICH47732 : : AtU6p : :sgKNA2_NbPDS, pICH47742 : :2x35 S- 5'UTR::Cas9::NOST, pICH47751 ::AtU6p::sgR Al_M> D5 level 1 constructs plus the pELE-3 linker [2] were cut-ligated into the pAGM4723 level 2 vector.
Tomato SlMlol locus
SlMlol sgR As 1 and 3:
pICH47732::NOSp::NPTII::OCST, pICH47742::2x35S-5'UTR::Cas9::NOST, pICH47751 ::AtU6p::sgRNAl_S/ oi, pICH47761 ::AtU6p::sgRNA3_S/ oi level 1 constructs plus the pELE-4 linker [2] were cut-ligated into the pAGM4723 level 2 vector.
SlMlol sgRNAs 2 and 3:
pICH47732::NOSp::NPTII::OCST, pICH47742::2x35S-5'UTR::Cas9::NOST, pICH47751 ::AtU6p::sgRNA2_S/ oi, pICH47761 ::AtU6p::sgRNA3_S/ oi level 1 constructs plus the pELE-4 linker were cut-ligated into the pAGM4723 level 2 vector.
SlMlol sgRNAs 3 and 4:
pICH47732::NOSp::NPTII::OCST, pICH47742::2x35S-5'UTR::Cas9::NOST, pICH47751 ::AtU6p::sgRNA3_S/ oi, pICH47761 ::AtU6p::sgRNA4_S/ oi level 1 constructs plus the pELE-4 linker were cut-ligated into the pAGM4723 level 2 vector.
The resulting level 2 constructs were transformed into the AGL1 strain of A. tumifaciens.
Transient gene expression in N. benthamiana and tomato
Transient expression in N. benthamiana and tomato was performed as the AGL1
12
strain of A. tumifaciens as described in Bos et al. .
Generation of stable transgenic tomato lines
Tomato transgenic lines were generated with same constructs as the ones used for the transient assay by agrotransformation of the tomato Moneymaker variety using the AGL1 A. tumifaciens strain. Transformants were selected on kanamycin.
Detection of Cas9-induced deletions in plant genomic DNA
In the case of the transient expression, Cas9 and sgRNAs were expressed in the N. benthamiana or tomato Moneymaker leaf tissue. The tissue was harvested at 2 days post infiltration and the genomic DNA extracted using the DNeasy Plant Mini kit (Qiagen). The genomic DNA extracted from the transiently expressing or stable transgenic tissue was digested with Sacl (New England Biolabs) in the case of N.
benthamiana PDS targets 1 and 2; with Bsrl (New England Biolabs) in the case of
tomato Mlol targets 1 and 3, and 2 and 3; with Ncol (New England Biolabs) in the case of tomato Mlol targets 3 and 4.
50 ng of pre-digested DNA was added in a PCR reaction and amplified with respective primers using Phusion DNA polymerase (New England Biolabs).
PCR products were run on an agarose gel. The DNA from bottom bands in lanes
2 and 4 (Figure 10B) was extracted and cloned into pCR-Blunt II-TOPO vector (Life Technologies). 15 individual clones were sequenced using standard Ml 3 forward and Ml 3 reverse primers. REFERENCES
Each of the publications, patents, and patent applications that listed below or referred to elsewhere herein is incorporated herein in its entirety by reference.
1. Griggs, D. et al. Nature 495, 305-307 (2013).
2. Voytas, D. Annu Rev Plant Biol. DOI: 10.1146/annurev-arplant-042811- 105552
(2013).
3. Mussolino, C, Cathomen, T. Nat Biotechnol . 31, 208-209 (2013).
4. Goodin, M. et al. Mol Plant Microbe Interact. 21, 1015-1026 (2008).
5. Mali, Petal. Science 339, 823-826 (2013).
6. Van der Hoorn, R. Mol Plant Microbe Interact. 13 , 439-446 (2000).
7. Cong, L. et al. Science 339, 819-823 (2013).
8. Qi, Y. et al. Genome Res 23, 547-554 (2013).
9. Karimi, M., Inze, D., Depicker, A. Trends Plant Sci 7, 193-195 (2002).
10. Waibel, F., Filipowicz, W. Nucleic Acids Res 18, 3451-3458 (1990).
11. Weber, E. etal PLoS One 6:el6765 (2011).
12. Bos, J. et al. Plant J 8, 165-176 (2006).
13. Jinek, M. et al., "A Programmable Dual-RNA-Guided DNA Endonuclease in Adaptive Bacterial Immunity", Science, August 17, 2012, pp. 816-821, Vol. 337.
14. Cain, A. et al. , "A CRISPR view of genome sequences", Nature Reviews
Microbiology, 2013, p. 1, Advance Online Publication.
Hwang, W. et al, "Efficient genome editing in zebrafish using a CRISPR-Cas system," Nature Biotechnology, January 29, 2013, pp. 1-3, Nature America, Inc.
Dicarlo, J. et al., "Genome engineering in Saccharomyces cerevisiae using CRISPR-Cas systems," Nucleic Acids Research, 2013, pp. 1-8, Oxford University Press.
Mali, P. et al., "RNA-Guided Human Genome Engineering via Cas9", Sciencexpress, http://www.sciencemag.org content/early/recent , January 3, 2013, pp. 1-5.
U.S. Patent Application Publication No. US 2010/0076057 Al, Sontheimer et al., Publication Date: March 25, 2010.
Lee, H. et al, "RNA-protein analysis using a conditional CRISPR nuclease", PNAS, April 2, 2013, pp. 5416-5421, Vol. 110(14).
Delker, R. et al., "A coming-of-age story: activation-induced cytidine deaminase turns 10", Nature Immunology, November 2009, pp. 1147-1153, Vol. 10(11).
Finney-Manchester, S . et al, "Harnessing mutagenic homologous
recombination for targeted mutagenesis in vivo by TaGTEAM", Nucleic Acids Research, 2013, pp. 1-10, Oxford University Press.
Voytas, D., "Plant Genome Engineering with Sequence-Specific Nucleases," Annu. Rev. Plant Biol, 2013, pp. 30.1-30.24, Vol. 64.
Jander, G., "Ethylmethanesulfonate Satuation Mutagenesis in Arabidopsis to Determine Frequency of Herbicide Resistance", Plant Physiology, January 2003, pp. 139-146, Vol. 131, American Society of Plant Biologists.
Zuo, J., et al, "An estrogen receptor-based transactivator SVE mediates highly inducible gene expression in transgenic plants", The Plant Journal, 2000, pp. 265-273, Vol. 24(2), Blackwell Science Ltd.
Lee, K., et al, "The molecular basis of sulfonylurea herbicide resistance in tobacco", The EMBO Journal, 1988, pp. 1241-1248, Vol. 7(5).
Matsui, K., et al, "Detection of protein-protein interactions in plants using the transrepressive activity of the EAR motif repression domain", The Plant Journal, 2010, pp. 570-578, Vol. 61, Blackwell Publishing Ltd.
Schwechheimer, C, "Understanding gibberellic acid signaling - are we there yet?", Current Opinion in Plant Biology, 2008, pp. 9-15, Vol. 11, Elsevier Ltd. Gawehns, F., "The potential of effector-target genes in breeding for plant innate immunity", Microbial Biotechnology, 2012, pp. 1-7, Society of Applied Microbiology and Blackwell Publishing Ltd.
Aoyama, T., et al., "A glucocorticoid-mediated transcriptional induction system in transgenic plants", The Plant Journal, 1997, pp. 605-612, Vol. 11(3).
Gawehns, F., "The potential of effector-target genes in breeding for plant innate immunity", Microbial Biotechnology, 2012, pp. 223-229, Vol. 6, Society of Applied Microbiology and Blackwell Publishing Ltd.
Hanin, M., et al., "Gene targeting in Arabidopsis" ', The Plant Journal, 2001, pp. 671-677, Vol. 28(6), Blackwell Science Ltd.
Lefeuvre, P. et al., "The Spread of Tomato Yellow Leaf Curl Virus from the Middle East to the World", PLoS Pathogens, October 2010, pp. 1-12, Vol. 6(10).
Durant, S. et al., "Vanillins - a novel family of DNA-PK inhibitors", Nucleic Acids Research, 2003, pp. 5501-5512, Vol. 31(19), Oxford University Press. Qi, L. et al., "Repurposing CRISPR as an RNA-Guided Platform for Sequence- Specific Control of Gene Expression", Cell, February 28, 2013, pp. 1173-1183, Vol. 152, Elsevier Inc.
Wendt, T., et al., "Production of Phytophthora infestans-resistant potato (Solanum tuberosum) utilising Ensifer adhaerens OV14", Transgenic Res., 2012, pp. 567-578, Vol. 21, No. 3.
WO 2011/076933, Mullins et al, Publication Date: June 30, 2011.
EP2361986 Al, Shen, J., et al, Publication Date: Aug 31, 2011.
Chung, S.-M., et al., "Agrobacterium is not alone: gene transfer to plants by viruses and other bacteria" 2006, Trends Plant Sci. 11(1): 1-4, Epub 2005 Nov 15.
Van Ex, F., et al., "Evaluation of seven promoters to achieve germline directed Cre-lox recombination in Arabidopsis thaliana", 2009, Plant Cell Rep., 28(10): 1509-20. doi: 10.1007/s00299-009-0750-y. Epub 2009 Aug 4.
40. Hong, R., et al., "Regulatory elements of the floral homeotic gene AGAMOUS identified by phylogenetic footprinting and shadowing", 2003, Plant Cell, 15(6): 1296-309.
41. Johnson, R., et al. "A rapid assay to quantify the cleavage efficiency of custom- designed nucleases in planta", Plant Mol Biol 2013 Apr 28. Epub.
42. U.S. Pat. App. Pub. No. 2009/0133158, Lahaye et al, Publication Date: May 21, 2009.
43. U.S. Prov. Pat. App. No. 61/791,760; Lahaye, T., filed March 15, 2013.
44. Nekrasov et al. "Targeted mutagenesis in the model plant Nicotiana
benthamiana using Cas9 RNA-guided endonuclease", Nat Biotechnol 2013,
31 :691-693.
The article "a" and "an" are used herein to refer to one or more than one (i.e., to at least one) of the grammatical object of the article. By way of example, "an element" means one or more element. As used herein, the word "comprising," or variations such as "comprises" or "comprising," will be understood to imply the inclusion of a stated element, integer or step, or group of elements, integers or steps, but not the exclusion of any other element, integer or step, or group of elements, integers or steps.
All publications and patent applications mentioned in the specification are indicative of the level of those skilled in the art to which this invention pertains. All publications and patent applications are herein incorporated by reference to the same extent as if each individual publication or patent application was specifically and individually indicated to be incorporated by reference.
Although the foregoing invention has been described in some detail by way of illustration and example for purposes of clarity of understanding, it will be obvious that certain changes and modifications may be practiced within the scope of the appended claims.
Claims
1. A method for modifying the genome of a plant, plant part, or plant cell, the method comprising producing at least one plant cell comprising a Cas protein and at least one engineered, single guide RNA (sgRNA) that is capable of recognizing a target nucleic acid sequence, wherein the plant, plant part, plant cell, or descent thereof, comprises in its genome the target nucleic acid sequence.
2. The method of claim 1 , wherein the Cas protein is Cas9 or a mutant or derivative of Cas9.
3. The method of claim 2, wherein the mutant or derivative is naturally occurring or engineered.
4. The method of any one of claims 1-3, wherein the Cas protein is capable of causing a double-strand break in DNA.
5. The method of any one of claims 1-3, wherein the Cas protein is capable of causing a single-strand break in DNA.
6. The method of claim 2 or 3, wherein the mutant or derivative comprises the DIOA amino acid substitution or the H841A amino acid substitution, and wherein the mutant or derivative is capable of causing a single-strand break in DNA.
7. The method of any one of claims 1-6, wherein producing at least one plant cell comprising a Cas protein and an engineered sgRNA, comprises:
(a) introducing into the at least one plant cell or a progenitor cell thereof a nucleic acid construct for expression of the sgRNA in the plant cell, whereby the sgRNA is expressed in the at least one plant cell;
(b) introducing into the at least one a plant cell or a progenitor cell thereof a nucleic acid construct for expression of the Cas protein in the plant cell, whereby the Cas protein is expressed in the at least one plant cell; or
(c) both (a) and (b).
8. The method of any one of claims 1-7, wherein the sgRNA, the Cas protein, or both are transiently expressed in the plant cell.
9. The method of claim 7 or 8, wherein the nucleic acid construct for expression of the sgRNA, the nucleic acid construct for expression of the Cas protein, or both are stably incorporated in the genome of the plant cell.
10. The method of any one of claims 1-9, wherein producing at least one plant cell comprising a Cas protein and an engineered sgRNA, comprises:
(i) directly introducing an engineered sgRNA into a plant cell and optionally further comprises virus-mediated delivery of the sgRNA into the plant cell;
(ii) directly introducing the Cas protein into the plant cell; or
(iii) both (i) and (ii).
11. The method of any one of claims 1-10, wherein the Cas protein or the nucleic acid construct for expression of the Cas protein is introduced into the plant cell or the progenitor, before, concurrently with, or after introducing the sgRNA or the nucleic acid construct for expression of the sgRNA.
12. The method of any one of claims 1-11, wherein at least one modification is introduced into the genome of the plant, plant part, or plant cell, or descendant thereof.
13. The method of claim 12, wherein the modification occurs within the target nucleic acid sequence.
14. The method of claim 12 or 13, wherein the modification comprises the insertion of at least one base pair in the genome of the plant, plant part, plant cell, or descendant thereof.
15. The method of claim 12 or 13, wherein the modification comprises the deletion of at least one base pair in the genome of the plant, plant part, plant cell, or descendant thereof.
16. The method of claim 12 or 13, wherein the modification comprises the insertion of at least one base pair in the genome of the plant, plant part, plant cell, or descendant thereof and the deletion of at least one base pair in the genome of the plant, plant part, plant cell, or descendant thereof.
17. The method of any one of claims 1-16, wherein the target nucleic acid sequence is within a gene or wherein the target nucleic acid sequence affects the level of expression of a gene.
18. The method of claim 17, wherein the gene is selected from the group consisting of a disease resistance gene, a disease susceptibility gene, a gene encoding an enzyme, a gene encoding a structural protein, a gene encoding a receptor, a gene encoding a membrane channel protein, a gene encoding a transcription factor, a gene that is associated with herbicide tolerance, a gene that affects development of male or female reproductive structures, and a gene that affects self-incompatibility.
19. The method of claim 17 or 18, wherein modification is a mutation in the gene that results in an alteration or loss of the wild-type function of the gene and/or the protein encoded thereby.
20. The method of claim 19, wherein the mutation is a loss-of- function mutation.
21. The method of any one of claims 1-20, further comprising introducing into the plant, plant part, plant cell, or descendant thereof a donor nucleic acid .
22. The method of claim 21 , wherein the donor nucleic acid molecule or part thereof is incorporated into the genome of the plant, plant part, or plant cell or descendant thereof.
23. The method of claim 21 or 22, wherein the donor nucleic acid molecule or part thereof is incorporated into the genome of the plant, plant part, plant cell, or descendant thereof within the target nucleic acid sequence.
24. The method of any one of claims 21-23, wherein the donor nucleic acid molecule comprises a nucleic acid sequence which comprises in a 5' to 3' order: a first homologous region, a donor region, and a second homologous region.
25. The method of claim 25, wherein the first homologous region comprises nucleic acid sequence homology to a first genomic region and the second homologous region comprises nucleic acid sequence homology to a second genomic region.
26. The method of claim 26, wherein the first genomic region and the second genomic region are on opposite sides of the site of incorporation of the donor nucleic acid molecule or part thereof into the genome of the plant, plant part, plant cell, or descendant thereof.
27. The method of any one of claims 21-26, wherein the donor region comprises a coding and/or a non-coding nucleic acid sequence.
28. The method of any one of claims 21-27, wherein the donor region comprises at least one transgene.
29. The method of claim 28, wherein the transgene comprises a promoter operably linked to a nucleic acid sequence, and wherein the promoter is capable of driving expression of the operably linked nucleic acid in a plant cell.
30. The method of claim 28 or 29, where the transgene encodes a protein.
31. The method of claim 30, wherein the protein is native to the plant, plant part, plant cell.
32. The method of claim 30, wherein the protein is a foreign protein that is not native the plant, plant part, or plant cell.
33. The method of any one of claims 21-32, wherein the donor nucleic acid comprises or further comprises at least one selectable marker gene.
34. The method of any one of claims 22-33, wherein incorporation of the donor nucleic acid into the genome of the plant, plant part, or plant cell or descendant thereof produces a selectable marker gene in the genome of the plant, plant part, or plant cell or descendant thereof.
35. The method of claim 34, wherein the donor nucleic acid comprises the entire, at least a portion of, or none of, the coding region of a protein encoded by the selectable marker gene.
36. The method of any one of claims 21-35, wherein the donor nucleic acid molecule is single stranded or double stranded.
37. The method of any one of claims 21-36, wherein the donor nucleic acid molecule is a linear nucleic acid molecular or a circular nucleic acid molecule.
38. The method of any one of claims 7-37, wherein the nucleic acid construct for expression of the sgR A comprises a promoter operably linked to a nucleic acid sequence encoding the sgRNA and wherein the promoter is capable of driving expression of a nucleic acid sequence in a plant cell.
39. The method of claim 38, wherein the promoter is an RNA polymerase III promoter.
40. The method of claim 38 or 39, wherein the promoter is the Arabidopsis U6 promoter.
41. The method of any one of claims 38-40, wherein the promoter comprises the Arabidopsis U6 promoter-derived synthetic nucleic acid sequence set forth in FIG. 4B.
42. The method of any one of claims 7-41, wherein nucleic acid construct for expression of the sgR A comprises the nucleic acid sequence of the U6::sgRNA construct set forth in FIG. 4B.
43. The method of any one of claims 7-42, wherein the nucleic acid construct for expression of the Cas protein comprises a promoter operably linked to a nucleic acid sequence encoding the Cas protein and optionally an operably linked eukaryotic nuclear localization signal sequence, wherein the promoter is capable of driving expression of a nucleic acid sequence in a plant cell.
44. The method of claim 43, wherein the promoter is an RNA polymerase II promoter.
45. The method of claims 43 or 44, wherein the promoter is a constitutive promoter, a tissue-specific promoter or an inducible promoter.
46. The method of any one of claims 43-45, wherein the promoter is selected from the group consisting of the CaMV 35S promoter, a ubiquitin promoter, an estradiol- inducible promoter, a dexamethasone-inducible promoter, an alcohol-inducible promoter, a heat shock promoter, an inflorescence-specific promoter, the Arabidopsis Apetala2 promoter, and the Arabidopsis Agamous promoter.
47. The method of any one of claims 1-46, wherein the target nucleic acid sequence comprises or consists of 19 bases.
48. The method of any one of claims 1-47, wherein immediately following the 3' end of the target nucleic acid sequence in the genome of the plant, plant part, or plant cells is a protospacer-adjacent motif (PAM).
49. The method of claim 48, wherein the PAM is NGG, wherein N can be A, C, G, or T.
50. The method of claim 47, wherein the Cas protein recognizes a PAM that is not NGG, wherein N can be A, C, G, or T.
51. The method of any one of claims 1-50, wherein the sgRNA comprises a guide nucleic acid sequence that is capable of binding to or recognizing the target nucleic acid or its complement.
52. The method of claim 51 , wherein the guide nucleic acid sequence is identical to the target nucleic acid sequence or its complement, when a U in the guide nucleic acid sequence is equivalent to a T in the target nucleic acid sequence or its complement.
53. The method of claim 51 , wherein the guide nucleic acid sequence is not identical to the target nucleic acid sequence or its complement, when a U in the guide nucleic acid sequence is equivalent to a T in the target nucleic acid sequence or its complement.
54. The method of claim 51 or 53, wherein there are one or more mismatched bases between the guide nucleic acid sequence and the target nucleic acid sequence or its complement.
55. The method of claim 54, wherein the one or more mismatches occur at a position or positions in the target nucleic acid sequence that are located within the first 8, 9, 10, 11, or 12 bases beginning from the 5' end of the target nucleic acid sequence.
56. The method of any of claims 1-55, further comprising regenerating the plant cell or descendent thereof into a plant comprising at least one modification in its genome or in the genome of a progeny plant or descendent of the regenerated plant, wherein at least one modification in the genome of a progeny plant or descendent is due to the presence of the sgRNA and the Cas protein in the progeny plant or descendent.
57. A plant, plant part, seed, or plant cell produced by the method of any one of claims 1-56, wherein the genome of the plant, plant part, seed, or plant cell is modified when compared to the genome of a control plant cell, and wherein said control plant cell is plant cell before its genome has been modified by the method.
58. A plant, plant part, seed, or plant cell comprising a modification in its genome that was introduced into the genome according to the method of any one of claims 1-56.
59. Use of the plant, plant part, or seed of claim 57 or 58 in agriculture and/or in the production of a human and/or animal food product.
60. Use of the plant, plant part, or seed of claim 57 or 58 in a plant breeding method and/or in the production of hybrid seed.
61. A method for targeting a fusion protein to a target nucleic acid sequence in the genome of a plant, plant part, or plant cell, producing at least one plant cell comprising a fusion protein and an engineered, single guide R A (sgRNA), wherein the fusion protein comprises a Cas protein domain that is completely or partially defective in nuclease activity and an additional domain, wherein the sgRNA is capable of recognizing the target nucleic acid sequence, whereby the fusion protein is capable of binding to or recognizing the target sequence when in the presence of the sgRNA.
62. The method of claim 61, wherein the Cas protein domain is a Cas9 protein domain or mutant or dervitive of Cas9 protein domain.
63. The method of claim 62, wherein Cas9 protein domain comprises the DIOA amino acid substitution, the H841A amino acid substitution, or both the DIOA and H841A amino acid substitutions.
64. The method of any one of claims 61-63, wherein producing the at least one plant cell comprising the fusion protein and the engineered, single guide RNA, comprises:
(a) introducing into the at least one plant cell or a progenitor cell thereof a nucleic acid construct for expression of the sgRNA in the plant cell, whereby the sgRNA is expressed in the at least one plant cell;
(b) introducing into the at least one a plant cell or a progenitor cell thereof a nucleic acid construct for expression of the fusion protein in the plant cell, whereby the fusion protein is expressed in the at least one plant cell; or
(c) both (a) and (b).
65. The method of claim of claim 64, wherein the sgRNA, the fusion protein, or both are transiently expressed in the plant cell.
66. The method of claim 64 or 65, wherein the nucleic acid construct for expression of the sgRNA, the nucleic acid construct for expression of the fusion protein, or both are stably incorporated in the genome of the plant cell.
67. The method of any one of claims 61-66, wherein producing the at least one plant cell comprising the fusion protein and the sgRNA, comprises:
(i) directly introducing an engineered sgRNA into a plant cell and optionally further comprises virus-mediated delivery of the sgRNA into the plant cell;
(ii) directly introducing the fusion protein into the plant cell; or
(iii) both (i) and (ii).
68. The method of any one of claims 61-67, wherein the fusion protein or the nucleic acid construct for expression of the fusion protein is introduced into the plant cell or the progenitor, before, concurrently with, or after introducing the sgRNA or the nucleic acid construct for expression of the sgRNA.
69. The method of any one of claims 61-68, wherein the additional domain is selected from the group consisting of a transcriptional activator domain, a transcriptional
repressor or co-repressor domain, a protein domain known to elevate the local mutagenesis rate, and a protein that can elevate the local recombination rate.
70. The method of any one of claims 61-69, wherein the additional domain is selected from the group consisting of an EAR domain, a SRDX domain, a TOPLESS domain, a cytidine deaminase or other DNA or histone modifying enzyme domain, a MAGI domain, and a RAD54 domain.
71. The method of any one of claims 61 -70, wherein the target nucleic acid sequence is within a gene.
72. The method of any one of claims 61-71, wherein the target nucleic acid sequence is within a promoter.
73. The method of any one of claims 64-72, wherein the nucleic acid construct for expression of the fusion protein comprises a promoter operably linked to a nucleic acid sequence encoding the fusion protein and optionally an operably linked eukaryotic nuclear localization signal sequence, wherein the promoter is capable of driving expression of a nucleic acid sequence in a plant cell.
74. The method of claim 71, wherein the promoter is an RNA polymerase II promoter.
75. The method of claim 73 or 74, wherein the promoter is a constitutive promoter, an inducible promoter, a developmentally regulated promoter, a tissue-specific promoter, and a seed-specific promoter.
76. The method of any one of claims 73-75, wherein the promoter is selected from the group consisting of the CaMV 35S promoter, a ubiquitin promoter, an estradiol- inducible promoter, a dexamethasone-inducible promoter, an alcohol-inducible promoter, a heat shock promoter, an inflorescence-specific promoter, the Arabidopsis Apetala2 promoter, and the Arabidopsis Agamous promoter.
77. The method of any one of claims 64-76, wherein the nucleic acid construct for expression of the sgRNA comprises a promoter operably linked to a nucleic acid sequence encoding the sgRNA, wherein the promoter is capable of driving expression of a nucleic acid sequence in a plant cell.
78. The method of claim 77, wherein the promoter is an RNA polymerase III promoter.
79. The method of claim 77 or 78, wherein the promoter is the Arabidopsis U6 promoter.
80. The method of any one of claims 77-79, wherein the promoter comprises the Arabidopsis U6 promoter-derived synthetic nucleic acid sequence set forth in FIG. 4B.
81. The method of any one of claims 77-80, wherein nucleic acid construct for expression of the sgRNA comprises the nucleic acid sequence of the U6::sgRNA construct set forth in FIG. 4B.
82. The method of any one of claims 61-81, wherein the target nucleic acid sequence comprises or consists of 19 bases.
83. The method of any one of claims 61-82, wherein immediately following the 3' end of the target nucleic acid sequence in the genome of the plant, plant part, or plant cells is a protospacer-adjacent motif (PAM).
84. The method of claim 83, wherein the PAM is NGG, wherein N can be A, C, G, or T.
85. The method of claim 83, wherein the Cas protein recognizes a PAM that is not NGG, wherein N can be A, C, G, or T.
86. The method of any one of claims 61-85, wherein the sgRNA comprises a guide nucleic acid sequence that is capable of binding to or recognizing the target nucleic acid or its complement.
87. The method of claim 86, wherein the guide nucleic acid sequence is identical to the target nucleic acid sequence or its complement, when a U in the guide nucleic acid sequence is equivalent to a T in the target nucleic acid sequence or its complement.
88. The method of claim 86, wherein the guide nucleic acid sequence is not identical to the target nucleic acid sequence or its complement, when a U in the guide nucleic acid sequence is equivalent to a T in the target nucleic acid sequence or its complement.
89. The method of claim 86 or 88, wherein there are one or more mismatched bases between the guide nucleic acid sequence and the target nucleic acid sequence or its complement.
90. The method of claim 89, wherein the one or more mismatches occur at a position or positions in the target nucleic acid sequence that are located within the first 8, 9, 10, 11, or 12 bases beginning from the 5' end of the target nucleic acid sequence.
91. The method of any one of claims 61-90, further comprising introducing at least one additional engineered sgRNA, wherein the sgRNA is capable of recognizing an additional target nucleic acid sequence.
92. The method of claim 91, wherein the fusion protein is also capable of binding to or recognizing the additional target sequence when in the presence of the additional sgRNA.
93. The method of any of claims 61-92, further comprising regenerating the plant cell or descendent thereof into a plant comprising the fusion protein and/or the
sgRNA and/or at least one modification in the genome of the plant cell or descendent that is due the fusion protein and the sgRNA.
94. A plant, plant part, seed, or plant cell produced by the method of any one of claims 61-93, wherein the plant, plant part, seed, or plant cell comprises the fusion protein and the engineered sgRNA.
95. A plant, plant part, seed, or plant cell comprising the fusion protein and/or sgRNA introduced into the genome according to the method of any one of claims 61-93 or comprising at least one modification in its genome that is due to the fusion protein and the sgRNA according to the method of any one of claims 61-93.
96. Use of the plant, plant part, or seed of claim 94 or 95 in agriculture and/or in the production of a human and/or animal food product.
97. Use of the plant, plant part, or seed of claim 94 or 95 in a plant breeding method and/or in the production of hybrid seed.
98. A fusion protein according to any of claims 61-93 or a nucleic acid molecule encoding the fusion protein.
99. A transient assay method for use in testing components of the Cas system in plants, the method comprising the steps of:
(a) contacting a plant tissue with one or more cells of a first bacterial strain comprising a first nucleic acid construct, wherein the first nucleic acid construct comprises an RNA polymerase II promoter that is expressible in a plant cell operably linked to nucleic acid sequence encoding a Cas protein, and wherein a cell of the first bacterial strain is capable of transferring a nucleic acid molecule to a plant cell;
(b) contacting the plant tissue with one or more cells of a second bacterial strain comprising a second nucleic acid construct, wherein the second nucleic acid construct comprises an RNA polymerase III promoter that is expressible in a plant cell operably linked to nucleic acid sequence encoding an engineered, single guide RNA (sgRNA) that is capable of recognizing a target nucleic acid sequence in the genome of
plant tissue, and wherein a cell of the second bacterial strain is capable of transferring a nucleic acid molecule to a plant cell; and
100. The method of claim 99, wherein the Cas protein is Cas9 or a mutant or derivative of Cas9.
101. The method of claim 100, wherein the mutant or derivative is naturally ocurring or engineered.
102. The method of any one of claims 99-101, wherein the Cas protein is capable of causing a double-strand break in DNA.
103. The method of any one of claims 99- 101, wherein the Cas protein is capable of causing a single-strand break in DNA.
104. The method of any one of claims 100, 101, and 103, wherein the mutant or derivative comprises the D10A amino acid substitution or the H841A amino acid substitution, and wherein the mutant or derivative is capable of causing a single-strand break in DNA.
105. The method of any one of claims 99-104, wherein at least one of the first and the second bacterial strains is a strain from a bacterial species that is selected from the group consisting of Agrobacterium tumefaciens, Agrobacterium rhizogenes, Ensifer adhaerens and other Ensifer spp., Sinorhizobium meliloti, and Mesorhizobium loti.
106. The method of any one of claims 99-105, wherein at least one of the first and the second bacterial strains is selected from the group consisting of Agrobacterium tumefaciens strain AGL1 and Ensifer adhaerens strain OV14.
107. The method of any one of claims 99-106, wherein at least one of the first and the second nucleic acid constructs is contained in a Ti plasmid or an Agrobacterium T-DNA border-containing broad host range binary vector..
108. The method of any one of claims 99-107, wherein the contacting of step
(a) comprises infiltration of the plant tissue with one or more cells of the first bacterial strain and/or the contacting of step (b) comprises infiltration of the plant tissue with one or more cells of the second bacterial strain.
109. The method of claim 108, wherein infiltration comprises use of a syringe to deliver the cells to the plant tissue.
110. The method of any one of claims 99-109, wherein steps (a) and (b) are performed at the same time.
111. The method of claim 110, wherein the plant tissue is infiltrated with a mixture comprising cells of the first bacterial strain and the second bacterial strain.
112. The method of any one of claims 99-111, wherein the plant tissue is from a plant selected from the group consisting of Nicotiana benthamiana, Nicotiana tabacum, Lactuca sativa, Brassica rapa, and any other plant species in which transient expression of genes after delivery of DNA either from bacteria or by other means can be achieved.
113. The method of any one of claims 99-112, wherein the plant tissue is leaf tissue.
114. The method of claim 112, wherein the leaf tissue is, or is part of, a whole leaf.
115. The method of any one of claims 99-114, wherein the plant tissue is attached to a plant or is detached from a plant.
116. The method of any one of claims 99-115, wherein the target nucleic acid sequence comprises or consists of 19 bases.
117. The method of any one of claims 99-116, wherein immediately following the 3' end of the target nucleic acid sequence in the genome of the plant, plant part, or plant cells is the protospacer-adjacent motif (PAM).
118. The method of claim 117, wherein the PAM is NGG, wherein N can be A, C, G, or T.
119. The method of claim 117, wherein the Cas protein recognizes a PAM that is not NGG, wherein N can be A, C, G, or T.
120. The method of any one of claims 99-119, wherein the sgRNA comprises a guide nucleic acid sequence that is capable of binding to or recognizing the target nucleic acid or its complement.
121. The method of claim 120, wherein the guide nucleic acid sequence is identical to the target nucleic acid sequence or its complement, when a U in the guide nucleic acid sequence is equivalent to a T in the target nucleic acid sequence or its complement.
122. The method of claim 120, wherein the guide nucleic acid sequence is not identical to the target nucleic acid sequence or its complement, when a U in the guide nucleic acid sequence is equivalent to a T in the target nucleic acid sequence or its complement.
123. The method of claim 120 or 123, wherein there are one or more mismatched bases between the guide nucleic acid sequence and the target nucleic acid sequence or its complement.
124. The method of claim 123, wherein the one or more mismatches occur at a position or positions in the target nucleic acid sequence that are located within the first 8, 9, 10, 11, or 12 bases beginning from the 5' end of the target nucleic acid sequence.
125. The method of any one of claims 99-124, wherein the target nucleic acid sequence is:
(i) native to the genome,
(ii) a foreign nucleic acid sequence that was introduced and stably incorporated into the genome of a plant from which the plant tissue was obtained,
(iii) a foreign nucleic acid sequence that was introduced and stably incorporated into the genome of a progenitor of the plant from which the plant tissue was obtained, or
(iv) co-delivered with the Cas gene and the sgR A.
126. The method of any one of claims 99-125, wherein monitoring the plant tissue for a modification of the genome within the target site comprises determining the nucleotide sequence of all or a part of the target nucleic acid sequence.
127. The method of any one of claims 99-126, wherein the target nucleic acid sequence comprises a restriction enzyme recognition sequence.
128. The method of claim 127, wherein step (c) comprises:
extracting genomic DNA from the plant tissue contacted by the first and second Agrobacterium strains:
digesting the genomic DNA with a restriction enzyme that recognizes the restriction enzyme recognition sequence;
amplifying the digested DNA by polymerase chain reaction using primers designed to amplify across the target nucleic acid sequence so as to produce PCR products comprising one or more modifications within target nucleic sequence;
cloning the PCR products; and
determining the nucleic acid sequence of the cloned PCR products.
129. The method of claim 127 or 128, wherein the restriction enzyme recognition sequence is a Mlyl restriction enzyme recognition sequence.
130. The method of claim 128 or 129, wherein the DNA is extracted from plant tissue harvested at about 1-48 hours after the later of the contacting steps.
131. The method of claim 128 or 129, wherein the DNA is extracted from leaves harvested at about 6, 12, 18, 24, 30, 36, 42, or 48 hours after the later of the contacting steps.
132. A method for modifying the genome of a plant, plant part, or plant cell, the method comprising producing at least one plant cell comprising a Cas protein and at least two engineered, sgRNAs that are each capable of recognizing a different target nucleic acid sequence, wherein the plant, plant part, plant cell, or descent thereof, comprises in its genome the target nucleic acid sequences.
133. The method of claim 132, wherein a first sgRNA is capable of recognizing a first target sequence and a second sgRNA is capable of recognizing a second target sequence.
134. The method of claim 133, wherein the first target sequence and the second target sequence are on the same chromosome.
135. The method of claim 133 or 134, wherein the first target sequence and the second target sequence are within the same gene.
136. The method of any one of claims 133-135, wherein the first target sequence and the second target sequence are separated by at least 10, 20, 30, 40, 50, 75, 100, 150, 200, 300, 400, 500, 1000, 1500, 2000, 3000 or more nucleotides.
137. The method of any one of claims 133-136, wherein the modification comprises the deletion of at least one base pair in the genome of the plant, plant part, plant cell, or descendant thereof.
138. The method claim 137, wherein the deletion occurs between the first target sequence and the second target sequence.
139. The method of any one of claims 133-138, wherein at least one of the Cas protein, the first sgRNA, and the second sgRNA are transiently expressed in the plant cell.
Applications Claiming Priority (2)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| US201361824748P | 2013-05-17 | 2013-05-17 | |
| US61/824,748 | 2013-05-17 |
Publications (2)
| Publication Number | Publication Date |
|---|---|
| WO2014186686A2 true WO2014186686A2 (en) | 2014-11-20 |
| WO2014186686A3 WO2014186686A3 (en) | 2015-01-08 |
Family
ID=50943610
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/US2014/038359 Ceased WO2014186686A2 (en) | 2013-05-17 | 2014-05-16 | Targeted mutagenesis and genome engineering in plants using rna-guided cas nucleases |
Country Status (1)
| Country | Link |
|---|---|
| WO (1) | WO2014186686A2 (en) |
Cited By (76)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO2015089406A1 (en) * | 2013-12-12 | 2015-06-18 | President And Fellows Of Harvard College | Cas variants for gene editing |
| US9163284B2 (en) | 2013-08-09 | 2015-10-20 | President And Fellows Of Harvard College | Methods for identifying a target site of a Cas9 nuclease |
| WO2015189693A1 (en) * | 2014-06-12 | 2015-12-17 | King Abdullah University Of Science And Technology | Targeted viral-mediated plant genome editing using crispr/cas9 |
| US9228207B2 (en) | 2013-09-06 | 2016-01-05 | President And Fellows Of Harvard College | Switchable gRNAs comprising aptamers |
| US9322006B2 (en) | 2011-07-22 | 2016-04-26 | President And Fellows Of Harvard College | Evaluation and improvement of nuclease cleavage specificity |
| US9322037B2 (en) | 2013-09-06 | 2016-04-26 | President And Fellows Of Harvard College | Cas9-FokI fusion proteins and uses thereof |
| US9359599B2 (en) | 2013-08-22 | 2016-06-07 | President And Fellows Of Harvard College | Engineered transcription activator-like effector (TALE) domains and uses thereof |
| WO2016137774A1 (en) * | 2015-02-25 | 2016-09-01 | Pioneer Hi-Bred International Inc | Composition and methods for regulated expression of a guide rna/cas endonuclease complex |
| US9526784B2 (en) | 2013-09-06 | 2016-12-27 | President And Fellows Of Harvard College | Delivery system for functional nucleases |
| US20170121693A1 (en) * | 2015-10-23 | 2017-05-04 | President And Fellows Of Harvard College | Nucleobase editors and uses thereof |
| CN107043778A (en) * | 2017-03-14 | 2017-08-15 | 广西大学 | Fungal gene pinpoints the nearly covering method in situ of insertion mutation |
| CN107164375A (en) * | 2017-05-25 | 2017-09-15 | 中国科学院天津工业生物技术研究所 | A kind of new guide rna expression box and the application in CRISPR/Cas systems |
| CN107236737A (en) * | 2017-05-19 | 2017-10-10 | 上海交通大学 | The sgRNA sequences of special target arabidopsis ILK2 genes and its application |
| US9834791B2 (en) | 2013-11-07 | 2017-12-05 | Editas Medicine, Inc. | CRISPR-related methods and compositions with governing gRNAS |
| WO2018054911A1 (en) | 2016-09-23 | 2018-03-29 | Bayer Cropscience Nv | Targeted genome optimization in plants |
| WO2018099256A1 (en) * | 2016-12-01 | 2018-06-07 | 中国农业科学院作物科学研究所 | Application of crispr/ncas9 mediated site-directed base substitution in plants |
| US10077453B2 (en) | 2014-07-30 | 2018-09-18 | President And Fellows Of Harvard College | CAS9 proteins including ligand-dependent inteins |
| US10113163B2 (en) | 2016-08-03 | 2018-10-30 | President And Fellows Of Harvard College | Adenosine nucleobase editors and uses thereof |
| US10136649B2 (en) | 2015-05-29 | 2018-11-27 | North Carolina State University | Methods for screening bacteria, archaea, algae, and yeast using CRISPR nucleic acids |
| WO2018220581A1 (en) * | 2017-05-31 | 2018-12-06 | Tropic Biosciences UK Limited | Compositions and methods for increasing shelf-life of banana |
| CN110234770A (en) * | 2017-01-17 | 2019-09-13 | 基础科学研究院 | Identify that base editor is missed the target the method in site by DNA single-strand break |
| EP3115457B1 (en) | 2014-03-05 | 2019-10-02 | National University Corporation Kobe University | Genomic sequence modification method for specifically converting nucleic acid bases of targeted dna sequence, and molecular complex for use in same |
| US10450584B2 (en) | 2014-08-28 | 2019-10-22 | North Carolina State University | Cas9 proteins and guiding features for DNA targeting and genome editing |
| US10450576B2 (en) | 2015-03-27 | 2019-10-22 | E I Du Pont De Nemours And Company | Soybean U6 small nuclear RNA gene promoters and their use in constitutive expression of small RNA genes in plants |
| US10519457B2 (en) | 2013-08-22 | 2019-12-31 | E I Du Pont De Nemours And Company | Soybean U6 polymerase III promoter and methods of use |
| US10584358B2 (en) | 2013-10-30 | 2020-03-10 | North Carolina State University | Compositions and methods related to a type-II CRISPR-Cas system in Lactobacillus buchneri |
| US10676754B2 (en) | 2014-07-11 | 2020-06-09 | E I Du Pont De Nemours And Company | Compositions and methods for producing plants resistant to glyphosate herbicide |
| CN111269998A (en) * | 2020-01-14 | 2020-06-12 | 青海大学 | Molecular marker linked with cabbage type rape limited inflorescence gene Bnsdt2 and application thereof |
| US10711267B2 (en) | 2018-10-01 | 2020-07-14 | North Carolina State University | Recombinant type I CRISPR-Cas system |
| US10745677B2 (en) | 2016-12-23 | 2020-08-18 | President And Fellows Of Harvard College | Editing of CCR5 receptor gene to protect against HIV infection |
| WO2020168102A1 (en) * | 2019-02-15 | 2020-08-20 | Sigma-Aldrich Co. Llc | Crispr/cas fusion proteins and systems |
| US10787654B2 (en) | 2014-01-24 | 2020-09-29 | North Carolina State University | Methods and compositions for sequence guiding Cas9 targeting |
| US10934536B2 (en) | 2018-12-14 | 2021-03-02 | Pioneer Hi-Bred International, Inc. | CRISPR-CAS systems for genome editing |
| CN112852813A (en) * | 2021-02-06 | 2021-05-28 | 南昌大学 | Asparagus U6 gene promoter AspU6p3 and cloning and application thereof |
| US11118189B2 (en) * | 2015-12-02 | 2021-09-14 | Ceres, Inc. | Methods for genetic modification of plants |
| US11155823B2 (en) | 2015-06-15 | 2021-10-26 | North Carolina State University | Methods and compositions for efficient delivery of nucleic acids and RNA-based antimicrobials |
| US11268082B2 (en) | 2017-03-23 | 2022-03-08 | President And Fellows Of Harvard College | Nucleobase editors comprising nucleic acid programmable DNA binding proteins |
| US11286480B2 (en) | 2015-09-28 | 2022-03-29 | North Carolina State University | Methods and compositions for sequence specific antimicrobials |
| US11306324B2 (en) | 2016-10-14 | 2022-04-19 | President And Fellows Of Harvard College | AAV delivery of nucleobase editors |
| US11319532B2 (en) | 2017-08-30 | 2022-05-03 | President And Fellows Of Harvard College | High efficiency base editors comprising Gam |
| US11345924B2 (en) * | 2017-05-11 | 2022-05-31 | Institute Of Genetics And Developmental Biology, Chinese Academy Of Sciences | Creation of herbicide resistant gene and use thereof |
| US11352623B2 (en) | 2020-03-19 | 2022-06-07 | Rewrite Therapeutics, Inc. | Methods and compositions for directed genome editing |
| CN114901302A (en) * | 2019-11-05 | 2022-08-12 | 成对植物服务股份有限公司 | Compositions and methods of RNA-encoded DNA replacement alleles |
| CN114945671A (en) * | 2019-11-01 | 2022-08-26 | 上海蓝十字医学科学研究所 | Method for targeted modification of plant genome sequence |
| US11439712B2 (en) | 2014-04-08 | 2022-09-13 | North Carolina State University | Methods and compositions for RNA-directed repression of transcription using CRISPR-associated genes |
| US11447770B1 (en) | 2019-03-19 | 2022-09-20 | The Broad Institute, Inc. | Methods and compositions for prime editing nucleotide sequences |
| US20220315938A1 (en) * | 2019-08-09 | 2022-10-06 | Regents Of The University Of Minnesota | AUGMENTED sgRNAS AND METHODS FOR THEIR USE TO ENHANCE SOMATIC AND GERMLINE PLANT GENOME ENGINEERING |
| US11542466B2 (en) | 2015-12-22 | 2023-01-03 | North Carolina State University | Methods and compositions for delivery of CRISPR based antimicrobials |
| US11542496B2 (en) | 2017-03-10 | 2023-01-03 | President And Fellows Of Harvard College | Cytosine to guanine base editor |
| US11542509B2 (en) | 2016-08-24 | 2023-01-03 | President And Fellows Of Harvard College | Incorporation of unnatural amino acids into proteins using base editing |
| US11560566B2 (en) | 2017-05-12 | 2023-01-24 | President And Fellows Of Harvard College | Aptazyme-embedded guide RNAs for use with CRISPR-Cas9 in genome editing and transcriptional activation |
| US11560568B2 (en) | 2014-09-12 | 2023-01-24 | E. I. Du Pont De Nemours And Company | Generation of site-specific-integration sites for complex trait loci in corn and soybean, and methods of use |
| US11649443B2 (en) | 2017-09-08 | 2023-05-16 | The Regents Of The University Of California | RNA-guided endonuclease fusion polypeptides and methods of use thereof |
| US11661590B2 (en) | 2016-08-09 | 2023-05-30 | President And Fellows Of Harvard College | Programmable CAS9-recombinase fusion proteins and uses thereof |
| US11732274B2 (en) | 2017-07-28 | 2023-08-22 | President And Fellows Of Harvard College | Methods and compositions for evolving base editors using phage-assisted continuous evolution (PACE) |
| US11795443B2 (en) | 2017-10-16 | 2023-10-24 | The Broad Institute, Inc. | Uses of adenosine base editors |
| US11898179B2 (en) | 2017-03-09 | 2024-02-13 | President And Fellows Of Harvard College | Suppression of pain by gene editing |
| US11912985B2 (en) | 2020-05-08 | 2024-02-27 | The Broad Institute, Inc. | Methods and compositions for simultaneous editing of both strands of a target double-stranded nucleotide sequence |
| US12084676B2 (en) | 2018-02-23 | 2024-09-10 | Pioneer Hi-Bred International, Inc. | Cas9 orthologs |
| US12133884B2 (en) | 2018-05-11 | 2024-11-05 | Beam Therapeutics Inc. | Methods of substituting pathogenic amino acids using programmable base editor systems |
| US12157760B2 (en) | 2018-05-23 | 2024-12-03 | The Broad Institute, Inc. | Base editors and uses thereof |
| US12203123B2 (en) | 2018-10-01 | 2025-01-21 | North Carolina State University | Recombinant type I CRISPR-Cas system and uses thereof for screening for variant cells |
| US12264313B2 (en) | 2018-10-01 | 2025-04-01 | North Carolina State University | Recombinant type I CRISPR-Cas system and uses thereof for genome modification and alteration of expression |
| US12264330B2 (en) | 2018-10-01 | 2025-04-01 | North Carolina State University | Recombinant type I CRISPR-Cas system and uses thereof for killing target cells |
| US12275939B2 (en) | 2017-09-19 | 2025-04-15 | Tropic Biosciences UK Limited | Modifying the specificity of plant non-coding RNA molecules for silencing gene expression |
| US12281338B2 (en) | 2018-10-29 | 2025-04-22 | The Broad Institute, Inc. | Nucleobase editors comprising GeoCas9 and uses thereof |
| US12331330B2 (en) | 2019-10-23 | 2025-06-17 | Pairwise Plants Services, Inc. | Compositions and methods for RNA-templated editing in plants |
| US12338444B2 (en) | 2011-03-23 | 2025-06-24 | Pioneer Hi-Bred International, Inc. | Methods for producing a complex transgenic trait locus |
| US12351837B2 (en) | 2019-01-23 | 2025-07-08 | The Broad Institute, Inc. | Supernegatively charged proteins and uses thereof |
| US12390514B2 (en) | 2017-03-09 | 2025-08-19 | President And Fellows Of Harvard College | Cancer vaccine |
| US12406749B2 (en) | 2017-12-15 | 2025-09-02 | The Broad Institute, Inc. | Systems and methods for predicting repair outcomes in genetic engineering |
| US12435330B2 (en) | 2019-10-10 | 2025-10-07 | The Broad Institute, Inc. | Methods and compositions for prime editing RNA |
| US12454694B2 (en) | 2018-09-07 | 2025-10-28 | Beam Therapeutics Inc. | Compositions and methods for improving base editing |
| US12473543B2 (en) | 2019-04-17 | 2025-11-18 | The Broad Institute, Inc. | Adenine base editors with reduced off-target effects |
| US12522807B2 (en) | 2018-07-09 | 2026-01-13 | The Broad Institute, Inc. | RNA programmable epigenetic RNA modifiers and uses thereof |
| US12529041B2 (en) | 2018-09-07 | 2026-01-20 | Beam Therapeutics Inc. | Compositions and methods for delivering a nucleobase editing system |
Families Citing this family (3)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN105131097B (en) * | 2015-08-31 | 2018-10-09 | 中国林业科学研究院林业研究所 | Chinese catalpa Pistil And Stamen develops relevant CabuAG genes and its protein and application |
| CN107937427A (en) * | 2017-10-20 | 2018-04-20 | 广东石油化工学院 | A kind of homologous repair vector construction method based on CRISPR/Cas9 systems |
| CN110331146B (en) * | 2019-09-05 | 2019-12-10 | 中国科学院天津工业生物技术研究所 | A promoter and expression vector for regulating sgRNA transcription, genome editing system and application thereof |
Family Cites Families (5)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| PT3241902T (en) * | 2012-05-25 | 2018-05-28 | Univ California | METHODS AND COMPOSITIONS FOR MODIFICATION OF TARGETED TARGET DNA BY RNA AND FOR MODULATION DIRECTED BY TRANSCRIPTION RNA |
| CA2877290A1 (en) * | 2012-06-19 | 2013-12-27 | Daniel F. Voytas | Gene targeting in plants using dna viruses |
| WO2014065596A1 (en) * | 2012-10-23 | 2014-05-01 | Toolgen Incorporated | Composition for cleaving a target dna comprising a guide rna specific for the target dna and cas protein-encoding nucleic acid or cas protein, and use thereof |
| KR20150105634A (en) * | 2012-12-12 | 2015-09-17 | 더 브로드 인스티튜트, 인코퍼레이티드 | Engineering and optimization of improved systems, methods and enzyme compositions for sequence manipulation |
| US8697359B1 (en) * | 2012-12-12 | 2014-04-15 | The Broad Institute, Inc. | CRISPR-Cas systems and methods for altering expression of gene products |
-
2014
- 2014-05-16 WO PCT/US2014/038359 patent/WO2014186686A2/en not_active Ceased
Cited By (155)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US12338444B2 (en) | 2011-03-23 | 2025-06-24 | Pioneer Hi-Bred International, Inc. | Methods for producing a complex transgenic trait locus |
| US10323236B2 (en) | 2011-07-22 | 2019-06-18 | President And Fellows Of Harvard College | Evaluation and improvement of nuclease cleavage specificity |
| US12006520B2 (en) | 2011-07-22 | 2024-06-11 | President And Fellows Of Harvard College | Evaluation and improvement of nuclease cleavage specificity |
| US9322006B2 (en) | 2011-07-22 | 2016-04-26 | President And Fellows Of Harvard College | Evaluation and improvement of nuclease cleavage specificity |
| US10508298B2 (en) | 2013-08-09 | 2019-12-17 | President And Fellows Of Harvard College | Methods for identifying a target site of a CAS9 nuclease |
| US9163284B2 (en) | 2013-08-09 | 2015-10-20 | President And Fellows Of Harvard College | Methods for identifying a target site of a Cas9 nuclease |
| US11920181B2 (en) | 2013-08-09 | 2024-03-05 | President And Fellows Of Harvard College | Nuclease profiling system |
| US10954548B2 (en) | 2013-08-09 | 2021-03-23 | President And Fellows Of Harvard College | Nuclease profiling system |
| US12428645B2 (en) | 2013-08-22 | 2025-09-30 | Pioneer Hi-Bred International, Inc. | Methods for producing genetic modifications in a plant genome without incorporating a selectable transgene marker, and compositions thereof |
| US10519457B2 (en) | 2013-08-22 | 2019-12-31 | E I Du Pont De Nemours And Company | Soybean U6 polymerase III promoter and methods of use |
| US11046948B2 (en) | 2013-08-22 | 2021-06-29 | President And Fellows Of Harvard College | Engineered transcription activator-like effector (TALE) domains and uses thereof |
| US11773400B2 (en) | 2013-08-22 | 2023-10-03 | E.I. Du Pont De Nemours And Company | Methods for producing genetic modifications in a plant genome without incorporating a selectable transgene marker, and compositions thereof |
| US9359599B2 (en) | 2013-08-22 | 2016-06-07 | President And Fellows Of Harvard College | Engineered transcription activator-like effector (TALE) domains and uses thereof |
| US12378566B2 (en) | 2013-08-22 | 2025-08-05 | Pioneer Hi-Bred International, Inc. | Plant genome modification using guide RNA/Cas endonuclease systems and methods of use |
| US10227581B2 (en) | 2013-08-22 | 2019-03-12 | President And Fellows Of Harvard College | Engineered transcription activator-like effector (TALE) domains and uses thereof |
| US9999671B2 (en) | 2013-09-06 | 2018-06-19 | President And Fellows Of Harvard College | Delivery of negatively charged proteins using cationic lipids |
| US9388430B2 (en) | 2013-09-06 | 2016-07-12 | President And Fellows Of Harvard College | Cas9-recombinase fusion proteins and uses thereof |
| US12473573B2 (en) | 2013-09-06 | 2025-11-18 | President And Fellows Of Harvard College | Switchable Cas9 nucleases and uses thereof |
| US10682410B2 (en) | 2013-09-06 | 2020-06-16 | President And Fellows Of Harvard College | Delivery system for functional nucleases |
| US9526784B2 (en) | 2013-09-06 | 2016-12-27 | President And Fellows Of Harvard College | Delivery system for functional nucleases |
| US9340800B2 (en) | 2013-09-06 | 2016-05-17 | President And Fellows Of Harvard College | Extended DNA-sensing GRNAS |
| US10597679B2 (en) | 2013-09-06 | 2020-03-24 | President And Fellows Of Harvard College | Switchable Cas9 nucleases and uses thereof |
| US9340799B2 (en) | 2013-09-06 | 2016-05-17 | President And Fellows Of Harvard College | MRNA-sensing switchable gRNAs |
| US9737604B2 (en) | 2013-09-06 | 2017-08-22 | President And Fellows Of Harvard College | Use of cationic lipids to deliver CAS9 |
| US10858639B2 (en) | 2013-09-06 | 2020-12-08 | President And Fellows Of Harvard College | CAS9 variants and uses thereof |
| US11299755B2 (en) | 2013-09-06 | 2022-04-12 | President And Fellows Of Harvard College | Switchable CAS9 nucleases and uses thereof |
| US10912833B2 (en) | 2013-09-06 | 2021-02-09 | President And Fellows Of Harvard College | Delivery of negatively charged proteins using cationic lipids |
| US9322037B2 (en) | 2013-09-06 | 2016-04-26 | President And Fellows Of Harvard College | Cas9-FokI fusion proteins and uses thereof |
| US9228207B2 (en) | 2013-09-06 | 2016-01-05 | President And Fellows Of Harvard College | Switchable gRNAs comprising aptamers |
| US11499169B2 (en) | 2013-10-30 | 2022-11-15 | North Carolina State University | Compositions and methods related to a type-II CRISPR-Cas system in Lactobacillus buchneri |
| US10584358B2 (en) | 2013-10-30 | 2020-03-10 | North Carolina State University | Compositions and methods related to a type-II CRISPR-Cas system in Lactobacillus buchneri |
| US11390887B2 (en) | 2013-11-07 | 2022-07-19 | Editas Medicine, Inc. | CRISPR-related methods and compositions with governing gRNAS |
| US10190137B2 (en) | 2013-11-07 | 2019-01-29 | Editas Medicine, Inc. | CRISPR-related methods and compositions with governing gRNAS |
| US10640788B2 (en) | 2013-11-07 | 2020-05-05 | Editas Medicine, Inc. | CRISPR-related methods and compositions with governing gRNAs |
| US9834791B2 (en) | 2013-11-07 | 2017-12-05 | Editas Medicine, Inc. | CRISPR-related methods and compositions with governing gRNAS |
| US9840699B2 (en) | 2013-12-12 | 2017-12-12 | President And Fellows Of Harvard College | Methods for nucleic acid editing |
| US12215365B2 (en) | 2013-12-12 | 2025-02-04 | President And Fellows Of Harvard College | Cas variants for gene editing |
| US10465176B2 (en) | 2013-12-12 | 2019-11-05 | President And Fellows Of Harvard College | Cas variants for gene editing |
| EP4375373A3 (en) * | 2013-12-12 | 2024-08-21 | President And Fellows Of Harvard College | Cas variants for gene editing |
| AU2014362208B2 (en) * | 2013-12-12 | 2021-02-11 | President And Fellows Of Harvard College | Cas variants for gene editing |
| EP3604511A1 (en) * | 2013-12-12 | 2020-02-05 | President And Fellows Of Harvard College | Cas variants for gene editing |
| US11053481B2 (en) | 2013-12-12 | 2021-07-06 | President And Fellows Of Harvard College | Fusions of Cas9 domains and nucleic acid-editing domains |
| US11124782B2 (en) | 2013-12-12 | 2021-09-21 | President And Fellows Of Harvard College | Cas variants for gene editing |
| WO2015089406A1 (en) * | 2013-12-12 | 2015-06-18 | President And Fellows Of Harvard College | Cas variants for gene editing |
| AU2021200375B2 (en) * | 2013-12-12 | 2023-08-17 | President And Fellows Of Harvard College | Cas variants for gene editing |
| US9068179B1 (en) | 2013-12-12 | 2015-06-30 | President And Fellows Of Harvard College | Methods for correcting presenilin point mutations |
| US10787654B2 (en) | 2014-01-24 | 2020-09-29 | North Carolina State University | Methods and compositions for sequence guiding Cas9 targeting |
| EP3115457B1 (en) | 2014-03-05 | 2019-10-02 | National University Corporation Kobe University | Genomic sequence modification method for specifically converting nucleic acid bases of targeted dna sequence, and molecular complex for use in same |
| US11718846B2 (en) | 2014-03-05 | 2023-08-08 | National University Corporation Kobe University | Genomic sequence modification method for specifically converting nucleic acid bases of targeted DNA sequence, and molecular complex for use in same |
| US11439712B2 (en) | 2014-04-08 | 2022-09-13 | North Carolina State University | Methods and compositions for RNA-directed repression of transcription using CRISPR-associated genes |
| US11584936B2 (en) | 2014-06-12 | 2023-02-21 | King Abdullah University Of Science And Technology | Targeted viral-mediated plant genome editing using CRISPR /Cas9 |
| WO2015189693A1 (en) * | 2014-06-12 | 2015-12-17 | King Abdullah University Of Science And Technology | Targeted viral-mediated plant genome editing using crispr/cas9 |
| US10676754B2 (en) | 2014-07-11 | 2020-06-09 | E I Du Pont De Nemours And Company | Compositions and methods for producing plants resistant to glyphosate herbicide |
| US12398406B2 (en) | 2014-07-30 | 2025-08-26 | President And Fellows Of Harvard College | CAS9 proteins including ligand-dependent inteins |
| US10077453B2 (en) | 2014-07-30 | 2018-09-18 | President And Fellows Of Harvard College | CAS9 proteins including ligand-dependent inteins |
| US10704062B2 (en) | 2014-07-30 | 2020-07-07 | President And Fellows Of Harvard College | CAS9 proteins including ligand-dependent inteins |
| US11578343B2 (en) | 2014-07-30 | 2023-02-14 | President And Fellows Of Harvard College | CAS9 proteins including ligand-dependent inteins |
| US11753651B2 (en) | 2014-08-28 | 2023-09-12 | North Carolina State University | Cas9 proteins and guiding features for DNA targeting and genome editing |
| US10450584B2 (en) | 2014-08-28 | 2019-10-22 | North Carolina State University | Cas9 proteins and guiding features for DNA targeting and genome editing |
| US12173294B2 (en) | 2014-09-12 | 2024-12-24 | Corteva Agriscience Llc | Generation of site specific integration sites for complex trait loci in corn and soybean, and methods of use |
| US11560568B2 (en) | 2014-09-12 | 2023-01-24 | E. I. Du Pont De Nemours And Company | Generation of site-specific-integration sites for complex trait loci in corn and soybean, and methods of use |
| CN107406858A (en) * | 2015-02-25 | 2017-11-28 | 先锋国际良种公司 | Compositions and methods for directing regulated expression of RNA/CAS endonuclease complexes |
| WO2016137774A1 (en) * | 2015-02-25 | 2016-09-01 | Pioneer Hi-Bred International Inc | Composition and methods for regulated expression of a guide rna/cas endonuclease complex |
| US10450576B2 (en) | 2015-03-27 | 2019-10-22 | E I Du Pont De Nemours And Company | Soybean U6 small nuclear RNA gene promoters and their use in constitutive expression of small RNA genes in plants |
| US10136649B2 (en) | 2015-05-29 | 2018-11-27 | North Carolina State University | Methods for screening bacteria, archaea, algae, and yeast using CRISPR nucleic acids |
| US11261451B2 (en) | 2015-05-29 | 2022-03-01 | North Carolina State University | Methods for screening bacteria, archaea, algae, and yeast using CRISPR nucleic acids |
| US11155823B2 (en) | 2015-06-15 | 2021-10-26 | North Carolina State University | Methods and compositions for efficient delivery of nucleic acids and RNA-based antimicrobials |
| US11286480B2 (en) | 2015-09-28 | 2022-03-29 | North Carolina State University | Methods and compositions for sequence specific antimicrobials |
| US12043852B2 (en) | 2015-10-23 | 2024-07-23 | President And Fellows Of Harvard College | Evolved Cas9 proteins for gene editing |
| US20170121693A1 (en) * | 2015-10-23 | 2017-05-04 | President And Fellows Of Harvard College | Nucleobase editors and uses thereof |
| US11214780B2 (en) | 2015-10-23 | 2022-01-04 | President And Fellows Of Harvard College | Nucleobase editors and uses thereof |
| US20180312825A1 (en) * | 2015-10-23 | 2018-11-01 | President And Fellows Of Harvard College | Nucleobase editors and uses thereof |
| US12344869B2 (en) | 2015-10-23 | 2025-07-01 | President And Fellows Of Harvard College | Nucleobase editors and uses thereof |
| US10167457B2 (en) | 2015-10-23 | 2019-01-01 | President And Fellows Of Harvard College | Nucleobase editors and uses thereof |
| EP3365356B1 (en) | 2015-10-23 | 2023-06-28 | President and Fellows of Harvard College | Nucleobase editors and uses thereof |
| US11802287B2 (en) | 2015-12-02 | 2023-10-31 | Ceres, Inc. | Methods for genetic modification of plants |
| US11118189B2 (en) * | 2015-12-02 | 2021-09-14 | Ceres, Inc. | Methods for genetic modification of plants |
| US11542466B2 (en) | 2015-12-22 | 2023-01-03 | North Carolina State University | Methods and compositions for delivery of CRISPR based antimicrobials |
| US11702651B2 (en) | 2016-08-03 | 2023-07-18 | President And Fellows Of Harvard College | Adenosine nucleobase editors and uses thereof |
| US11999947B2 (en) | 2016-08-03 | 2024-06-04 | President And Fellows Of Harvard College | Adenosine nucleobase editors and uses thereof |
| US10947530B2 (en) | 2016-08-03 | 2021-03-16 | President And Fellows Of Harvard College | Adenosine nucleobase editors and uses thereof |
| US10113163B2 (en) | 2016-08-03 | 2018-10-30 | President And Fellows Of Harvard College | Adenosine nucleobase editors and uses thereof |
| US11661590B2 (en) | 2016-08-09 | 2023-05-30 | President And Fellows Of Harvard College | Programmable CAS9-recombinase fusion proteins and uses thereof |
| US11542509B2 (en) | 2016-08-24 | 2023-01-03 | President And Fellows Of Harvard College | Incorporation of unnatural amino acids into proteins using base editing |
| US12084663B2 (en) | 2016-08-24 | 2024-09-10 | President And Fellows Of Harvard College | Incorporation of unnatural amino acids into proteins using base editing |
| WO2018054911A1 (en) | 2016-09-23 | 2018-03-29 | Bayer Cropscience Nv | Targeted genome optimization in plants |
| US11306324B2 (en) | 2016-10-14 | 2022-04-19 | President And Fellows Of Harvard College | AAV delivery of nucleobase editors |
| WO2018099256A1 (en) * | 2016-12-01 | 2018-06-07 | 中国农业科学院作物科学研究所 | Application of crispr/ncas9 mediated site-directed base substitution in plants |
| US10745677B2 (en) | 2016-12-23 | 2020-08-18 | President And Fellows Of Harvard College | Editing of CCR5 receptor gene to protect against HIV infection |
| US11820969B2 (en) | 2016-12-23 | 2023-11-21 | President And Fellows Of Harvard College | Editing of CCR2 receptor gene to protect against HIV infection |
| CN110234770A (en) * | 2017-01-17 | 2019-09-13 | 基础科学研究院 | Identify that base editor is missed the target the method in site by DNA single-strand break |
| US11898179B2 (en) | 2017-03-09 | 2024-02-13 | President And Fellows Of Harvard College | Suppression of pain by gene editing |
| US12390514B2 (en) | 2017-03-09 | 2025-08-19 | President And Fellows Of Harvard College | Cancer vaccine |
| US12516308B2 (en) | 2017-03-09 | 2026-01-06 | President And Fellows Of Harvard College | Suppression of pain by gene editing |
| US11542496B2 (en) | 2017-03-10 | 2023-01-03 | President And Fellows Of Harvard College | Cytosine to guanine base editor |
| US12435331B2 (en) | 2017-03-10 | 2025-10-07 | President And Fellows Of Harvard College | Cytosine to guanine base editor |
| CN107043778A (en) * | 2017-03-14 | 2017-08-15 | 广西大学 | Fungal gene pinpoints the nearly covering method in situ of insertion mutation |
| US11268082B2 (en) | 2017-03-23 | 2022-03-08 | President And Fellows Of Harvard College | Nucleobase editors comprising nucleic acid programmable DNA binding proteins |
| US12195741B2 (en) | 2017-05-11 | 2025-01-14 | Institute Of Genetics And Developmental Biology, Chinese Academy Of Sciences | Creation of herbicide resistant gene and use thereof |
| US11345924B2 (en) * | 2017-05-11 | 2022-05-31 | Institute Of Genetics And Developmental Biology, Chinese Academy Of Sciences | Creation of herbicide resistant gene and use thereof |
| US11560566B2 (en) | 2017-05-12 | 2023-01-24 | President And Fellows Of Harvard College | Aptazyme-embedded guide RNAs for use with CRISPR-Cas9 in genome editing and transcriptional activation |
| CN107236737A (en) * | 2017-05-19 | 2017-10-10 | 上海交通大学 | The sgRNA sequences of special target arabidopsis ILK2 genes and its application |
| CN107164375B (en) * | 2017-05-25 | 2020-12-29 | 中国科学院天津工业生物技术研究所 | A Novel Guide RNA Expression Cassette and Its Application in CRISPR/Cas System |
| CN107164375A (en) * | 2017-05-25 | 2017-09-15 | 中国科学院天津工业生物技术研究所 | A kind of new guide rna expression box and the application in CRISPR/Cas systems |
| WO2018220581A1 (en) * | 2017-05-31 | 2018-12-06 | Tropic Biosciences UK Limited | Compositions and methods for increasing shelf-life of banana |
| EP4461813A3 (en) * | 2017-05-31 | 2025-02-26 | Tropic Biosciences UK Limited | Compositions and methods for increasing shelf-life of banana |
| US11732274B2 (en) | 2017-07-28 | 2023-08-22 | President And Fellows Of Harvard College | Methods and compositions for evolving base editors using phage-assisted continuous evolution (PACE) |
| US12359218B2 (en) | 2017-07-28 | 2025-07-15 | President And Fellows Of Harvard College | Methods and compositions for evolving base editors using phage-assisted continuous evolution (PACE) |
| US11932884B2 (en) | 2017-08-30 | 2024-03-19 | President And Fellows Of Harvard College | High efficiency base editors comprising Gam |
| US11319532B2 (en) | 2017-08-30 | 2022-05-03 | President And Fellows Of Harvard College | High efficiency base editors comprising Gam |
| US11649443B2 (en) | 2017-09-08 | 2023-05-16 | The Regents Of The University Of California | RNA-guided endonuclease fusion polypeptides and methods of use thereof |
| US12415993B2 (en) | 2017-09-08 | 2025-09-16 | The Regents Of The University Of California | RNA-guided endonuclease fusion polypeptides and methods of use thereof |
| US12497601B2 (en) | 2017-09-08 | 2025-12-16 | The Regents Of The University Of California | RNA-guided endonuclease fusion polypeptides and methods of use thereof |
| US11649442B2 (en) | 2017-09-08 | 2023-05-16 | The Regents Of The University Of California | RNA-guided endonuclease fusion polypeptides and methods of use thereof |
| US12331295B2 (en) | 2017-09-19 | 2025-06-17 | Tropic Biosciences UK Limited | Modifying the specificity of plant non-coding RNA molecules for silencing gene expression |
| US12275939B2 (en) | 2017-09-19 | 2025-04-15 | Tropic Biosciences UK Limited | Modifying the specificity of plant non-coding RNA molecules for silencing gene expression |
| US11795443B2 (en) | 2017-10-16 | 2023-10-24 | The Broad Institute, Inc. | Uses of adenosine base editors |
| US12406749B2 (en) | 2017-12-15 | 2025-09-02 | The Broad Institute, Inc. | Systems and methods for predicting repair outcomes in genetic engineering |
| US12084676B2 (en) | 2018-02-23 | 2024-09-10 | Pioneer Hi-Bred International, Inc. | Cas9 orthologs |
| US12133884B2 (en) | 2018-05-11 | 2024-11-05 | Beam Therapeutics Inc. | Methods of substituting pathogenic amino acids using programmable base editor systems |
| US12157760B2 (en) | 2018-05-23 | 2024-12-03 | The Broad Institute, Inc. | Base editors and uses thereof |
| US12522807B2 (en) | 2018-07-09 | 2026-01-13 | The Broad Institute, Inc. | RNA programmable epigenetic RNA modifiers and uses thereof |
| US12454694B2 (en) | 2018-09-07 | 2025-10-28 | Beam Therapeutics Inc. | Compositions and methods for improving base editing |
| US12529041B2 (en) | 2018-09-07 | 2026-01-20 | Beam Therapeutics Inc. | Compositions and methods for delivering a nucleobase editing system |
| US12203123B2 (en) | 2018-10-01 | 2025-01-21 | North Carolina State University | Recombinant type I CRISPR-Cas system and uses thereof for screening for variant cells |
| US11680259B2 (en) | 2018-10-01 | 2023-06-20 | North Carolina State University | Recombinant type I CRISPR-CAS system |
| US10711267B2 (en) | 2018-10-01 | 2020-07-14 | North Carolina State University | Recombinant type I CRISPR-Cas system |
| US12264313B2 (en) | 2018-10-01 | 2025-04-01 | North Carolina State University | Recombinant type I CRISPR-Cas system and uses thereof for genome modification and alteration of expression |
| US12264330B2 (en) | 2018-10-01 | 2025-04-01 | North Carolina State University | Recombinant type I CRISPR-Cas system and uses thereof for killing target cells |
| US12281338B2 (en) | 2018-10-29 | 2025-04-22 | The Broad Institute, Inc. | Nucleobase editors comprising GeoCas9 and uses thereof |
| US12215364B2 (en) | 2018-12-14 | 2025-02-04 | Pioneer Hi-Bred International, Inc. | CRISPR-cas systems for genome editing |
| US10934536B2 (en) | 2018-12-14 | 2021-03-02 | Pioneer Hi-Bred International, Inc. | CRISPR-CAS systems for genome editing |
| US11807878B2 (en) | 2018-12-14 | 2023-11-07 | Pioneer Hi-Bred International, Inc. | CRISPR-Cas systems for genome editing |
| US12365888B2 (en) | 2018-12-14 | 2025-07-22 | Pioneer Hi-Bred International, Inc. | CRISPR-Cas systems for genome editing |
| US12351837B2 (en) | 2019-01-23 | 2025-07-08 | The Broad Institute, Inc. | Supernegatively charged proteins and uses thereof |
| WO2020168102A1 (en) * | 2019-02-15 | 2020-08-20 | Sigma-Aldrich Co. Llc | Crispr/cas fusion proteins and systems |
| US11965184B2 (en) | 2019-02-15 | 2024-04-23 | Sigma-Aldrich Co. Llc | CRISPR/Cas fusion proteins and systems |
| CN113728099A (en) * | 2019-02-15 | 2021-11-30 | 西格马-奥尔德里奇有限责任公司 | CRISPR/CAS fusion proteins and systems |
| US12509680B2 (en) | 2019-03-19 | 2025-12-30 | The Broad Institute, Inc. | Methods and compositions for prime editing nucleotide sequences |
| US11795452B2 (en) | 2019-03-19 | 2023-10-24 | The Broad Institute, Inc. | Methods and compositions for prime editing nucleotide sequences |
| US11447770B1 (en) | 2019-03-19 | 2022-09-20 | The Broad Institute, Inc. | Methods and compositions for prime editing nucleotide sequences |
| US11643652B2 (en) | 2019-03-19 | 2023-05-09 | The Broad Institute, Inc. | Methods and compositions for prime editing nucleotide sequences |
| US12281303B2 (en) | 2019-03-19 | 2025-04-22 | The Broad Institute, Inc. | Methods and compositions for prime editing nucleotide sequences |
| US12473543B2 (en) | 2019-04-17 | 2025-11-18 | The Broad Institute, Inc. | Adenine base editors with reduced off-target effects |
| US20220315938A1 (en) * | 2019-08-09 | 2022-10-06 | Regents Of The University Of Minnesota | AUGMENTED sgRNAS AND METHODS FOR THEIR USE TO ENHANCE SOMATIC AND GERMLINE PLANT GENOME ENGINEERING |
| US12435330B2 (en) | 2019-10-10 | 2025-10-07 | The Broad Institute, Inc. | Methods and compositions for prime editing RNA |
| US12331330B2 (en) | 2019-10-23 | 2025-06-17 | Pairwise Plants Services, Inc. | Compositions and methods for RNA-templated editing in plants |
| CN114945671A (en) * | 2019-11-01 | 2022-08-26 | 上海蓝十字医学科学研究所 | Method for targeted modification of plant genome sequence |
| CN114901302A (en) * | 2019-11-05 | 2022-08-12 | 成对植物服务股份有限公司 | Compositions and methods of RNA-encoded DNA replacement alleles |
| CN111269998A (en) * | 2020-01-14 | 2020-06-12 | 青海大学 | Molecular marker linked with cabbage type rape limited inflorescence gene Bnsdt2 and application thereof |
| US11352623B2 (en) | 2020-03-19 | 2022-06-07 | Rewrite Therapeutics, Inc. | Methods and compositions for directed genome editing |
| US12031126B2 (en) | 2020-05-08 | 2024-07-09 | The Broad Institute, Inc. | Methods and compositions for simultaneous editing of both strands of a target double-stranded nucleotide sequence |
| US11912985B2 (en) | 2020-05-08 | 2024-02-27 | The Broad Institute, Inc. | Methods and compositions for simultaneous editing of both strands of a target double-stranded nucleotide sequence |
| CN112852813B (en) * | 2021-02-06 | 2022-07-29 | 南昌大学 | Asparagus U6 gene promoter AspU6p3 and cloning and application thereof |
| CN112852813A (en) * | 2021-02-06 | 2021-05-28 | 南昌大学 | Asparagus U6 gene promoter AspU6p3 and cloning and application thereof |
Also Published As
| Publication number | Publication date |
|---|---|
| WO2014186686A3 (en) | 2015-01-08 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| US12173294B2 (en) | Generation of site specific integration sites for complex trait loci in corn and soybean, and methods of use | |
| WO2014186686A2 (en) | Targeted mutagenesis and genome engineering in plants using rna-guided cas nucleases | |
| US11459576B2 (en) | Methods for producing a complex transgenic trait locus | |
| US20220177900A1 (en) | Genome modification using guide polynucleotide/cas endonuclease systems and methods of use | |
| US20220364107A1 (en) | Agronomic trait modification using guide rna/cas endonuclease systems and methods of use | |
| AU2020202241B2 (en) | Manipulation of dominant male sterility | |
| US20180002715A1 (en) | Composition and methods for regulated expression of a guide rna/cas endonuclease complex | |
| CA3197672A1 (en) | Engineered cas endonuclease variants for improved genome editing | |
| CA2954626A1 (en) | Compositions and methods for producing plants resistant to glyphosate herbicide | |
| EP3694992A1 (en) | Type i-e crispr-cas systems for eukaryotic genome editing | |
| Lu et al. | Low frequency of zinc-finger nuclease-induced mutagenesis in Populus | |
| CN115151637A (en) | Homologous recombination within the genome | |
| CN115243711B (en) | Two-step gene exchange | |
| CN110959043A (en) | Methods for improving plant agronomic traits using BCS1L gene and guide RNA/CAS endonuclease system |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| 121 | Ep: the epo has been informed by wipo that ep was designated in this application |
Ref document number: 14730704 Country of ref document: EP Kind code of ref document: A2 |
|
| 122 | Ep: pct application non-entry in european phase |
Ref document number: 14730704 Country of ref document: EP Kind code of ref document: A2 |