WO2012138901A1 - Method for enhancing rare-cutting endonuclease efficiency and uses thereof - Google Patents
Method for enhancing rare-cutting endonuclease efficiency and uses thereof Download PDFInfo
- Publication number
- WO2012138901A1 WO2012138901A1 PCT/US2012/032386 US2012032386W WO2012138901A1 WO 2012138901 A1 WO2012138901 A1 WO 2012138901A1 US 2012032386 W US2012032386 W US 2012032386W WO 2012138901 A1 WO2012138901 A1 WO 2012138901A1
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- rare
- cutting endonuclease
- cutting
- endonuclease
- chimeric
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Ceased
Links
Classifications
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N9/00—Enzymes; Proenzymes; Compositions thereof; Processes for preparing, activating, inhibiting, separating or purifying enzymes
- C12N9/14—Hydrolases (3)
- C12N9/16—Hydrolases (3) acting on ester bonds (3.1)
- C12N9/22—Ribonucleases [RNase]; Deoxyribonucleases [DNase]
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2319/00—Fusion polypeptide
Definitions
- the present invention relates to a method for enhancing the efficiency of rare-cutting endonucleases such as meganucleases. More specifically, the present invention concerns a method for the creation of fusion proteins that consist of one or more "enhancer" domains fused to at least one rare-cutting endonuclease-derived scaffold in a single polypeptide chain for simple and efficient vectorization.
- the present invention also relates to engineered chimeric rare-cutting endonucleases, vectors, compositions and kits used to implement the method and use of said chimeric rare-cutting endonucleases according to the invention for various applications ranging from homologous gene targeting to targeted mutagenesis and sequence removal.
- DSBs double-strand breaks
- HR homologous recombination
- SSA Single-strand annealing
- NHEJ non-homologous end joining
- HCT Homologous gene targeting
- Orr- Weaver Szostak et al. 1981
- Orr-Weaver Szostak et al. 1983
- Rothstein 1983 was one of the first methods for rational genome engineering and remains to this day a standard for the generation of engineered cells or knock-out mice (Capecchi 2001).
- An inherently low efficiency has nevertheless prevented it from being used as a routine protocol in most cell types and organisms.
- DSBs induced by biological reagents e.g. meganucleases, ZFNs and TALENs (see below), which cleave DNA by hydrolysis of two phosphodiester bonds
- the DNA can be rejoined in a seamless manner by simple re-ligationof the cohesive ends.
- deleterious insertions or deletions indels of various sizes can occur at the breaks, eventually resulting in gene inactivation (Liang, Han et al. 1998; Lloyd, Plaisier et al. 2005; Doyon, McCammon et al. 2008; Perez, Wang et al. 2008; Santiago, Chan et al. 2008; Kim, Lee et al.
- Targeted mutagenesis based on NEHJ has been used to trigger inactivation of single or even multiple genes in immortalized cell lines (Cost, Freyvert et al. 2010; Liu, Chan et al. 2010).
- this method opens new perspectives for organisms in which the classical HR-based gene knock-out methods have proven inefficient, or at least difficult to establish (Doyon, McCammon et al. 2008; Geurts, Cost et al. 2009; Shukla, Doyon et al. 2009; Yang, Djukanovic et al. 2009; Gao, Smith et al. 2010; Mashimo, Takizawa et al. 2010; Menoret, Iscache et al. 2010).
- Meganucleases also called homing endonucleases (HEs) can be divided into five families based on sequence and structure motifs: LAGLIDADG, GIY-YIG, HNH, His-Cys box and PD-(D/E)XK (Stoddard 2005; Zhao, Bonocora et al. 2007). Structural data are available for at least one member of each family. The most well studied family is that of the LAGLIDADG proteins, with a considerable body of biochemical, genetic and structural work having established that these endonucleases could be used as molecular tools (Stoddard, Scharenberg et al. 2007; Arnould, Delenda et al. 2011).
- Member proteins are composed of domains that adopt a similar ⁇ fold, with the LAGLIDADG motif comprising the terminal region of the first helix and not only contributing to a bipartite catalytic center but also forming the core subunit/subunit interaction (Stoddard 2005). Two such ⁇ / ⁇ domains assemble to form the functional protein, with the ⁇ -strands in each creating a saddle- shaped DNA binding region. The spatial separation of the catalytic center with regions directly interacting with the DNA has allowed for specificity re-engineering (Seligman, Chisholm et al. 2002; Sussman, Chadsey et al. 2004; Arnould, Chames et al.
- LAGLIDADG HEs members from two other families, GIY-YIG and HNH, are of particular interest. Biochemical and structural studies have established that in both families, member proteins can adopt a bipartite fold with distinct functional domains: (1) a catalytic domain responsible mainly for DNA cleavage, and; (2) a DNA-binding domain to provide target specificity (Stoddard 2005; Marcaida, Munoz et al. 2010).
- the related GIY-YIG HEs I- TevI and l-Bmol have been exploited to demonstrate the interchangeability of the DNA-binding region for these enzymes (Liu, Derbyshire et al. 2006).
- IENR1 intron-encoded endonuclease repeat motif found in endonucleases of the GIY-YIG family.
- the catalytic head of l-Basl has sequence similarity to those of the HNH HEs l-Hmul, l-Hmull and l-Twol, all of which function as strand-specific nickases (Landthaler, Begley et al. 2002; Landthaler and Shub 2003; Landthaler, Lau et al.
- HNH motif has also been identified in nonspecific nucleases such the E.coli colicins (e.g. ColE9 and ColE7), EndA from S. pneumoniae, NucA from Anabaena and CAD (Midon, Schafer et al. 2011).
- E.coli colicins e.g. ColE9 and ColE7
- EndA from S. pneumoniae
- NucA from Anabaena
- CAD Mordon, Schafer et al. 2011
- several of these nucleases contain the signature DRGH motif and share structural homology with core elements forming the ⁇ -Me-finger active site motif.
- Zinc-finger nucleases generated by fusing Zinc-finger-based DNA-binding domains to an independent catalytic domain via a flexible linker (Kim, Cha et al. 1996; Smith, Berg et al. 1999; Smith, Bibikova et al. 2000), represent another type of engineered nuclease commonly used to stimulate gene targeting.
- the archetypal ZFNs are based on the catalytic domain of the Type IIS restriction enzyme Fokl and have been successfully used to induce gene correction, gene insertion, and gene deletion.
- Zinc Finger-based DNA binding domains are made of strings of 3 or 4 individual Zinc Fingers, each recognizing a DNA triplet (Pabo, Peisach et al. 2001).
- ZFNs are easy to design, using combinatorial assembly of preexisting Zinc Fingers with known recognition patterns (Choo and Klug 1994; Choo and Klug 1994; Kim, Lee et al. 2009).
- close examination of high resolution structures shows that there are actually crosstalks between units (EIrod-Erickson, Rould et al. 1996), and several methods have been used to assemble ZF proteins by choosing individual Zinc Fingers in a context dependant manner (Greisman and Pabo 1997; Isalan and Choo 2001; Maeder, Thibodeau-Beganny et al. 2008; Ramirez, Foley et al. 2008) to achieve better success rates and reagents of better quality.
- TALE Transcription Activator Like Effectors
- each base pair in the DNA target is contacted by a single repeat, with the specificity resulting from the two variant amino acids of the repeat (the so-called repeat variable dipeptide, RVD).
- RVD repeat variable dipeptide
- TALEN Fokl-based TALE-nuclease
- the functional layout of a Fokl-based TALE-nuclease is essentially that of a ZFN, with the Zinc-finger DNA binding domain being replaced by the TALE domain (Christian, Cermak et al. 2010; Li, Huang et al. 2011).
- DNA cleavage by a TALEN requires two DNA recognition regions flanking an unspecific central region. This central "spacer" DNA region is essential to promote catalysis by the dimerizing Fokl catalytic domain, and extensive effort has been placed into optimizing the distance between the DNA binding sites (Christian, Cermak et al. 2010; Miller, Tan et al. 2011).
- the length of the spacer has been varied from 14 to 30 base pairs, with efficacy in DNA cleavage being interdependent with spacer length as well as TALE scaffold construction (i.e. the nature of the fusion construct used). It is still unknown whether differences in the repeat region (i.e. RVD type and number used) have an impact on the DNA "spacer” requirements or on the efficacy of DNA cleavage by TALENs. Nevertheless, TALE-nucleases have been shown to be active to various extents in cell-based assays in yeast, mammalian cells and plants (Christian, Cermak et al. 2010; Li, Huang et al. 2011; Mahfouz, Li et al. 2011; Miller, Tan et al. 2011).
- the efficiency of rare-cutting endonuclease e.g. their ability to induce a desired event (Homologous gene targeting, targeted mutagenesis, sequence removal or excision) at a locus, depends on several parameters, including the specific activity of the nuclease, probably the accessibility of the target, and the efficacy and outcome of the repair pathway(s) resulting in the desired event (homologous repair for gene targeting, NHEJ pathways for targeted mutagenesis).
- Nucleases (Smith, Bibikova et al. 2000). These ends, which result from hydrolysis of phosphodiester bonds, can be re-ligated in vivo by NHEJ in a seamless way (i.e a scarless re-ligation).
- the restoration of a cleavable target sequence allows for a new cleavage event by the same endonuclease, and thus, a series of futile cycles of cleavage and re-ligation events can take place.
- Indirect evidences have shown that even in the yeast Saccharomyces cerevisiae, such cycles could take place upon continuous cleavage by the HO endonuclease (Lee, Paques et al. 1999).
- DSB resection is important for certain DSB pathways. Extensive DSB resection, resulting in the generation of large single stranded regions (a few hundred nucleotides at least), has been shown in yeast to initiate single strand annealing (Sugawara and Haber 1992) and strand invasion, the ATP-dependant step that initiates many homologous recombination events of DNA duplex invasion by an homologous strand that (White and Haber 1990; Sun, Treco et al. 1991) (for a review of mechanisms, see (Paques and Haber 1999)).
- DSB resection depends on several proteins including BLM/Sgsl and DNA2, EXOI, and the MRN complex (Mrell, Rad50, Nbsl/Xrs2) and is thought to result from different pathways. MRN is involved in a small scale resection process, while two redundant pathways depending on BLM and DNA2 on one hand, and on EXOI on another hand, would be involved in extensive resection (Mimitou and Symington 2008; Nimonkar, Genschel et al. 2011).
- the authors of the present invention have developed a method to significantly enhance the efficiency of rare-cutting endonucleases such as LAGLIDADG-type meganucleases.
- the use of novel "enhancer" domains allows for boosting the overall efficiency of the meganuclease without modifying its specificity determinants (i.e. residues making base-specific contacts to the DNA).
- the invention allows for generating several distinct types of enzymes that can be applied to applications ranging from homologous gene targeting to targeted mutagenesis and sequence removal.
- the present invention relates to a method for enhancing the efficiency of a rare- cutting endonuclease for a DNA target sequence by merging a rare-cutting endonuclease-derived scaffold with at least one enhancer domain to obtain a chimeric rare-cutting endonuclease with an enhanced efficiency for said DNA target sequence.
- the present invention concerns a method for the creation of fusion proteins that consist of engineering a fusion protein between a rare-cutting endonuclease-derived scaffold and at least one enhancer domain wherein said enhancer domain enhances the efficiency of said rare-cutting endonuclease when fused to it.
- the present invention also concerns the creation of functional single polypeptide fusion proteins for simple and efficient vectorization.
- the present invention relates to chimeric rare- cutting endonucleases comprising at least an enhancer domain wherein said enhancer domain enhances the efficiency of said rare-cutting endonuclease when fused to it, thereby obtaining a chimeric rare-cutting endonuclease with enhanced efficiency for a DNA target sequence compared to a corresponding rare-cutting endonuclease lacking said enhancer domain.
- the present invention also relates to engineered rare-cutting endonucleases, vectors, compositions and kits used to implement the method and use of said chimeric rare-cutting endonucleases according to the invention for various applications ranging from homologous gene targeting to targeted mutagenesis and sequence removal.
- Figure 1 Endonuclease-induced gene targeting approaches. Upon cleavage, DNA repair mechanisms may result in one of several outcomes.
- A When a double-strand break is targeted between two direct repeats, HR can result in the deletion of one repeat together with the intervening sequence.
- Gene insertion (B) or correction (C) can be achieved by the introduction of a DNA repair matrix containing sequences homologous to the endogenous sequence surrounding the DNA break. Mutations can be corrected either at or distal to the break, with the frequency of correction decreasing with increasing distance.
- D The misrepair of DNA ends by error-prone NHEJ can result in insertions or deletions of various sizes, leading to gene inactivation.
- Figure 2 Sequences of target DNA recognized by l-Crel.
- C1234 (SEQ ID NO: 3) represents the partially symmetric DNA sequence recognized and cleaved by wild-type l-Crel.
- C1221 (SEQ ID NO: 2) represents a palindromic DNA sequence, derived from C1234 (SEQ ID NO: 3), recognized and cleaved by the l-Crel meganuclease. Nucleotides are numbered outward (-/+) from the center of the target. The nature of the nucleotides at positions -2 to +2 can potentially interfere with the cleavage activity of the protein.
- FIG. 3 Schematic of enhanced rare-cutting endonucleases, such as meganucleases, fusion configurations.
- Rare-cutting endonucleases such as meganucleases, can be enhanced through the addition of a domain to promote existing or alternate activities.
- the order (N- v.s C-terminal) of addition and number of the enhancer domains can vary with the application.
- Enhanced fusion construct are optimized to address or overcome distinct problems.
- A The addition of two enhancer domains to an active meganuclease can enhance DNA binding and/or cleavage activity.
- Such a configuration can be achieved via (i) a single N- or C-terminal fusion to a homodimeric variant; (ii) a single N- or C-terminal fusion to individual monomers of a heterodimer, or; (iii) a double fusion to a monomeric protein.
- B When specificity reengineering precludes maintaining cleavage activity of the meganuclease, the attached enhancer domains can provide alternative functions.
- C) and (D) represent instances of (A) and (B), respectively, when only one enhancer domain is needed or tolerated per fusion protein (e.g. either as an N- or C-terminal fusion or in the context of a single-chain molecule). Fusion junctions (N- vs. C- terminal) and linker designs can vary with the application. Components of the fusion proteins are listed in the legend.
- FIG. 4 Schematic of DNA cleavage, in vivo re-ligation and other repair pathways.
- cleavage by peptidic rare-cutting endonucleases usually result in a DNA double strand break (DSB) with cohesive ends.
- DSB DNA double strand break
- meganucleases from the LAGLIDADG family such as l-Scel and I- Crel, produce DSBs with 3' overhang.
- These cohesive ends can be re-ligated in vivo by NHEJ, resulting in seamless repair, and in the restoration of a cleavable target sequence, which can in turn be processed again by the same endonuclease.
- a series of futile cycles of cleavage and re-ligation events can take place.
- Imprecise NHEJ or homologous recombination can alter or remove the cleavage site, resulting in cycle exit (A).
- Two other ways can also stop the process : (i) Chromosome loss can occur as the consequence of failure to repair the DSB; (ii) a loss of nuclease (degradation, dilution, cell division, etc.).
- B-E Consequences of cleavage of additional phosphodiester bonds. The addition of a single nickase activity (B) or of two nickase activities affecting the same strand (C) would result in a single strand gap, and suppress the cohesive ends, which could in turn affect the spectrum of events.
- the present invention relates to a method for enhancing the efficiency of a rare- cutting endonuclease for a DNA target sequence by merging a rare-cutting endonuclease-derived scaffold with at least one enhancer domain to obtain a chimeric rare-cutting endonuclease with an enhanced efficiency for said DNA target sequence.
- a first aspect of the present invention is a method for enhancing rare-cutting endonuclease efficiency at a genomic locus of interest in a cell comprising the steps of:
- said enhancer domain is fused to the N-terminus part of said rare-cutting endonuclease-derived scaffold. In another preferred embodiment, said enhancer domain is fused to the C-terminus part of said rare-cutting endonuclease-derived scaffold. In another embodiment, two enhancer domains are fused to both N-terminus part of said rare-cutting endonuclease-derived scaffold and C-terminus part of said rare-cutting endonuclease-derived scaffold.
- said enhancer domain(s) are fused to N-terminus and/or C-terminus of a rare-cutting endonuclease-derived monomer scaffold of a homodimeric endonuclease, resulting in a chimeric rare-cutting endonuclease comprising two or four enhancer domains.
- said enhancer domain(s) are fused to N-terminus and/or C-terminus of a rare-cutting endonuclease- derived monomer scaffold of a heterodimeric endonuclease, resulting in a chimeric rare-cutting endonuclease comprising one or two or three or four enhancer domains.
- said enhancer domain(s) are fused to N-terminus and/or C-terminus of a rare-cutting endonuclease- derived scaffold from a single-chain endonuclease, resulting in a chimeric rare-cutting endonuclease comprising one or two enhancer domains.
- said enhancer domain is catalytically active, or not providing functional and/or structural support to said rare-cutting endonuclease-derived scaffold.
- said enhancer domain consists of a protein domain derived from an exonuclease.
- said enhancer domain consists of a protein domain derived from an endonuclease.
- said enhancer domain consists of a protein domain derived from a cleavase.
- said enhancer domain consists of a protein domain derived from a nickase.
- said enhancer domain consists of a protein domain derived from a protein selected from the group consisting of Mmel, Colicin-E7 (CEA7_ECOLX), EndA, Endo I (ENDl_ECOLI), Human Endo G (NUCG_HUMAN), Bovine Endo G (NUCG_BOVIN), .HinPll, l-Basl, I- Bmol, l-Hmul, l-Tevl, l-Tevll, l-Tevlll, l-Twol, R.Mspl, R.Mval, NucA, NucM, Vvn, Vvn_CLS, Staphylococcal nuclease (NUC_STAAU), Staphylococcal nuclease (NUC_STAHY), Micrococcal nuclease (NUC_SHIFL), Endonuclease yncB, Endodeoxyribonuclease I (
- said enhancer domain consists of a peptide derived from CFPl peptide (SEQ ID NO: 112).
- any combinations of two protein domains selected from the group consisting of Mmel, Colicin-E7 (CEA7_ECOLX), EndA, Endo I (ENDl_ECOLI), Human Endo G (NUCG_HUMAN), Bovine Endo G (NUCG_BOVIN), R.HinPll, l-Basl, l-Bmol, l-Hmul, I- Tevl, l-Tevll, l-Tevlll, l-Twol, R.Mspl, R.Mval, NucA, NucM, Vvn, Vvn_CLS, Staphylococcal nuclease (NUC_STAAU), Staphylococcal nuclease (NUC_STAHY), Micrococcal nuclease (NUC_SHIFL), Endonu
- l-Hmul catalytic domain can be fused to the N-terminus part of said rare-cutting endonuclease-derived scaffold and ColE7 can be fused to the C-terminus part of said rare-cutting endonuclease-derived scaffold.
- said enhancer domain consists of a catalytically active derivative of the protein domains listed above and in Table 1, providing functional and/or structural support to said rare-cutting endonuclease- derived scaffold.
- said enhancer domain consists of a catalytically inactive derivative of the protein domains listed above and in Table 1, providing structural support to said rare-cutting endonuclease-derived scaffold.
- said rare-cutting endonuclease-derived scaffold is derived from a meganuclease.
- said meganuclease comprises two identical monomers.
- said meganuclease comprises two non-identical monomers.
- said meganuclease is a single-chain meganuclease.
- said rare-cutting endonuclease-derived scaffold is derived from the group consisting of l-Crel, a functional mutant of l-Crel, a variant of l-Crel or a derivative thereof.
- rare-cutting endonuclease-derived scaffold is a truncated form of wild- type l-Crel (SEQ ID NO: 1).
- rare-cutting endonuclease-derived scaffold comprises first 152, 153, 154 or 155 amino acids residues of wild-type l-Crel (SEQ ID NO: 1).
- rare-cutting endonuclease-derived scaffold comprises residues 2 to 153 of wild-type l-Crel (SEQ ID NO: 1).
- rare-cutting endonuclease-derived scaffold comprises residues 2 to 155 of wild-type I- Crel (SEQ ID NO: 1).
- rare-cutting endonuclease-derived scaffold comprises residues 2 to 153 of wild-type l-Crel (SEQ ID NO: 1) and one or several amino acids substitutions.
- rare-cutting endonuclease-derived scaffold comprises residues 2 to 155 of wild-type l-Crel (SEQ ID NO: 1) and one or several amino acids substitutions.
- rare-cutting endonuclease-derived scaffold of the present invention comprises residues 2 to 153 or residues 2 to 155 of wild-type l-Crel (SEQ ID NO: 1) and one or two or three or four or five or six or seven or eight or nine or ten further amino acid mutations.
- rare-cutting endonuclease-derived scaffold of the present invention comprises residues 2 to 153 of wild-type l-Crel (SEQ ID NO: 1) and K82A mutation.
- said rare-cutting endonuclease-derived scaffold comprises a sequence selected from the group consisting of l-Crel_NFSl (SEQ ID NO: 6); l-Crel_NFS2 (SEQ ID NO: 7); and l-Crel_CFSl (SEQ ID NO: 8).
- said chimeric rare-cutting endonuclease can comprise at least one peptidic linker between said rare-cutting endonuclease-derived scaffold and said at least one enhancer domain.
- said peptidic linker sequence is selected from the group consisting of
- the peptidic linker that can link said enhancer domain to the rare-cutting endonuclease-derived scaffold according to the method of the present invention can be selected from the group consisting of NFSl (SEQ ID NO: 98), NFS2 (SEQ ID NO: 99) and CFSl (SEQ ID NO: 100).
- NFSl SEQ ID NO: 98
- NFS2 SEQ ID NO: 99
- CFSl SEQ ID NO: 100.
- a peptidic linker is not needed to fuse said enhancer domain to said rare-cutting endonuclease-derived scaffold in order to obtain a chimeric rare-cutting endonuclease according to the present invention.
- Table 2 List of peptidic linkers that can be used in chimeric rare-cutting endonuclease.
- Enhancement of efficiency of a chimeric rare-cutting endonuclease according to the present invention, compared to a starting rare-cutting endonuclease, can also be a consequence of a fuctional support by said at least one enhancer domain.
- said functional support can be the consequence of the hydrolysis of additional phosphodiester bonds.
- said functional support can be the hydrolysis of additional phosphodiester bonds by a protein domain derived from a nuclease.
- said functional support can be the hydrolysis of additional phosphodiester bonds by a protein domain derived from an endonuclease.
- said functional support can be the hydrolysis of additional phosphodiester bonds by a protein domain derived from a cleavase. In another more preferred embodiment, said functional support can be the hydrolysis of additional phosphodiester bonds by a protein domain derived from a nickase. In a more preferred embodiment, said functional support can be the hydrolysis of additional phosphodiester bonds by a protein domain derived from an exonuclease.
- enhancement of efficiency of a chimeric rare-cutting endonuclease according to the present invention can be the hydrolysis of additional phosphodiester bonds at the cleavage site to promote exit from such futile cycles of cleavage and re-ligation events via imprecise NHEJ or Homologous Recombination or Single Strand Annealing (SSA).
- SSA Single Strand Annealing
- Said hydrolysis of additional phosphodiester bonds at the cleavage site by said at least one enhancer according to the invention can lead to different types of DSB resection affecting at said DSB cleavage site, one single DNA strand or both DNA strands, affecting either 5' overhangs ends, either 3' overhangs ends, or both ends and depending on the length of said resection.
- adding new nickase or cleavase activities to the existing cleavase activity of a rare-cutting endonuclease enhances the efficiency of the resulting chimeric rare-cutting endonuclease according to the invention, at a genomic locus of interest.
- the addition of a single nickase activity can result in a single strand gap, and suppress the cohesivity of the ends, which can also enhances the efficiency of the resulting chimeric rare-cutting endonuclease at a genomic locus of interest, according to the invention, via stimulation of one or several alternative repair outcomes mentioned above.
- the method according to the invention enhances rare-cutting endonuclease efficiency for a DNA target sequence.
- Enhancement of efficiency of a rare-cutting endonuclease according to the present invention can be a consequence of a structural support by said at least one enhancer domain.
- said structural support enhances the binding of a chimeric rare-cutting endonuclease according to the invention for a DNA target sequence compared to the binding of a starting rare-cutting endonuclease for the same DNA target sequence, as a non-limiting example.
- said structural support by at least one enhancer domain enhances the existing catalytical activity of a chimeric rare-cutting
- said at least one enhancer domain enhances both the binding and the existing catalytical activity of a chimeric rare-cutting endonuclease according to the invention for a DNA target sequence compared to the binding of a starting rare-cutting endonuclease for the same DNA target sequence, as another non-limiting example. All these non- limiting examples lead to a chimeric rare-cutting endonuclease with enhanced efficiency for a DNA target at a genomic locus of interest, according to the present invention.
- the resulting chimeric rare-cutting endonuclease can comprise several enzymatic activities that contribute to the enhancement of its efficiency according to the present invention.
- the addition of one nickase domain to an active rare- cutting endonuclease scaffold derived from a meganuclease results in a chimeric rare-cutting endonuclease with one meganuclease activity and one nickase activity (depicted in figure 4B).
- the addition of two nickase domains to an active rare-cutting endonuclease scaffold derived from a meganuclease results in a chimeric rare-cutting endonuclease with one meganuclease activity and two nickase activities (as depicted in figures 5C and 5D).
- the addition of one cleavase domain to an active rare-cutting endonuclease scaffold derived from a meganuclease results in a chimeric rare-cutting endonuclease with one meganuclease activity and one cleavase activity (figure 4E).
- the addition of two cleavase domains to an active rare-cutting endonuclease scaffold derived from a meganuclease results in a chimeric rare-cutting endonuclease with one meganuclease activity and two cleavase activities.
- enhancement of efficiency of a rare-cutting endonuclease refers to the increase in the detected level of said efficiency, against a target DNA sequence, of a second rare-cutting endonuclease in comparison to the activity of a first rare-cutting endonuclease against the same target DNA sequence.
- Said second rare-cutting endonuclease can be an engineered rare-cutting endonuclease, i.e. a chimeric rare-cutting endonuclease according to the invention or not.
- Said first rare-cutting endonuclease can be a wild-type rare-cutting endonuclease, a starting rare-cutting endonuclease, an engineered rare-cutting endonuclease i.e a chimeric rare-cutting
- Said enhancement of efficiency can be determined, as non-limiting example, by measuring the level of cleavage-induced recombination generated by said rare-cutting endonuclease or said chimeric rare-cutting endonuclease according to the present invention. Said level of cleavage-induced recombination can be measured by a cell-based recombination assay as described in the International PCT Application WO 2004/067736, as non-limiting example.
- enhancement of efficacy in cells can be, but is not necessarily associated with an enhancement of the cleavage activity that could be detected in certain in vitro assays.
- additional phosphodiesterase activities as described in Figure 4 could barely affect the cleavage profile, as detected by in vitro cleavage and separation of the cleavage products on an electrophoresis gel.
- the DSB ends generated in this way could be more prone to induce detectable genomic rearrangements such as targeted mutagenesis (by imprecise NHEJ) or homologous recombination.
- Said enhancement in efficacy of said rare-cutting endonuclease is at least a 5% enhancement compared to the starting scaffold, more preferably at least a 10% enhancement, again more preferably at least a 15% enhancement, again more preferably at least a 20% enhancement, again more preferably at least a 25% enhancement, again more preferably a 50% enhancement, again more preferably a enhancement greater than 50%.
- the enhancement of efficiency of a rare-cutting endonuclease according to the method of the present invention can also be an enhancement in the efficiency for 2NN derivatives of a DNA target sequence by a rare-cutting endonuclease from the LAGLIDADG meganuclease subfamily.
- 2NN derivatives of a DNA target sequence is intended variants of a given DNA target sequence having mutations in the four central nucleotides -2 to +2 as numbered in figure 2.
- the method according to the invention enhances the efficiency for derivatives of the DNA target sequence C1221 (SEQ ID NO: 2) having mutations in the four central nucleotides -2 to +2 as numbered in figure 2, i.e that do not have GTAC sequence at position -2 to +2 as numbered in figure 2.
- the method according to the invention allows the recombination to be initiated at 2NN derivatives that are not normally processed by a starting rare- cutting endonuclease scaffold from the LAGLIDADG meganuclease subfamily.
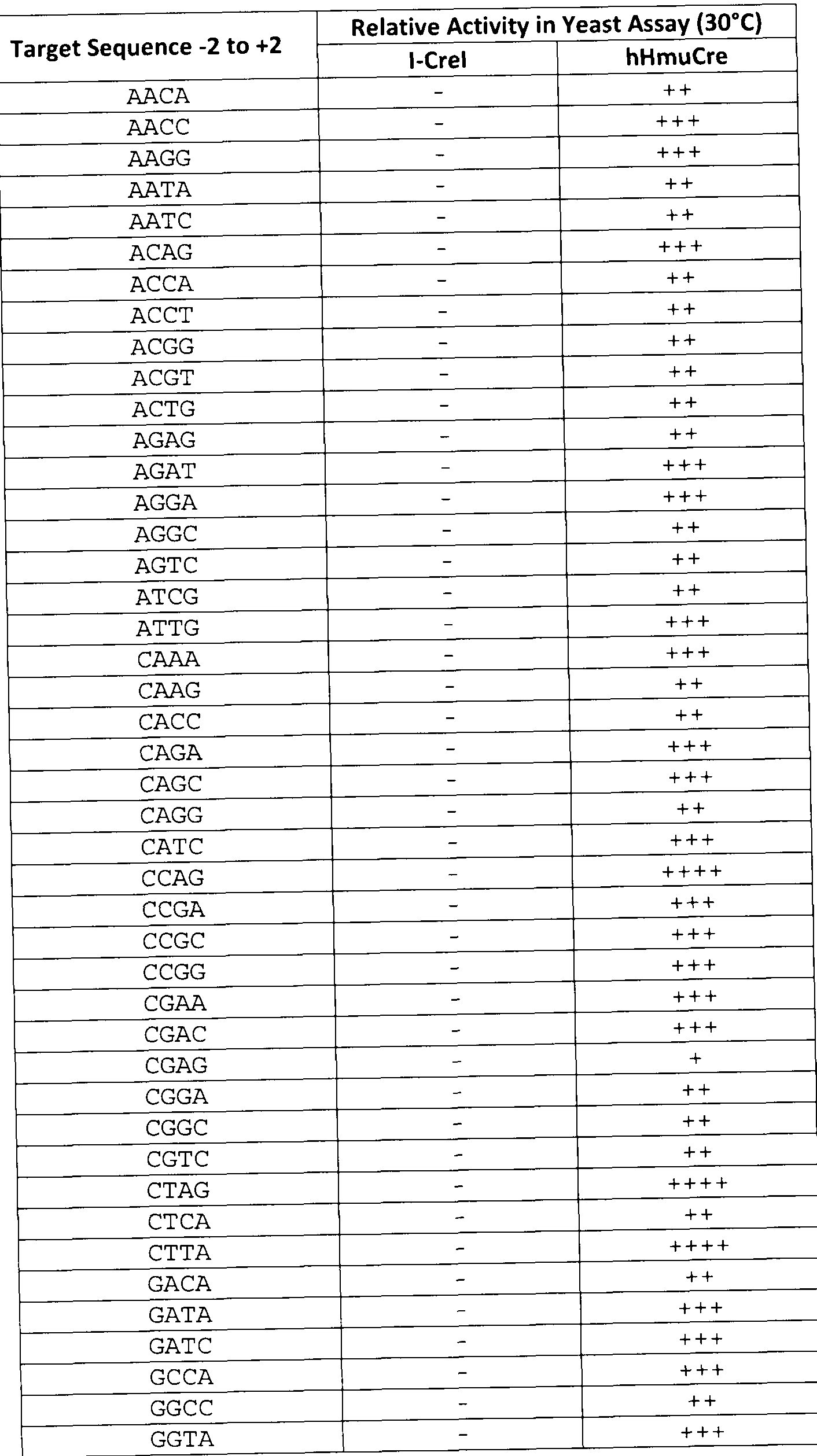
- the method according to the invention allows the processing of 2NN derivatives that have - 2 to +2 target sequences as listed in Table 4 and Table 5, non-originally cleaved by l-Crel.
- this first aspect of the invention is the method according to the invention wherein said chimeric rare-cutting endonuclease is selected from the group consisting of hHmuCre_D0101 (SEQ ID NO: 9), hHmuCre_D0201 (SEQ ID NO: 105), hHmuCre_D0301 (SEQ ID NO: 106), hHmuCre_D0401 (SEQ ID NO: 107), hHmuCre_D0102 (SEQ ID NO: 108), hHmuCre_D0202 (SEQ ID NO: 109), hHmuCre_D0302 (SEQ ID NO: 110) and hHmuCre_D0402 (SEQ ID NO: 111).
- efficiency of rare-cutting endonucleases such as meganucleases can be enhanced through the addition of a domain to promote existing or alternate activities.
- the order (N- v.s C- terminal) of addition and number of the enhancer domains can vary with the application as depicted in Figure 3.
- Enhanced fusion construct are optimized to address or overcome distinct problems.
- Figure 3A The addition of two enhancer domains to an active rare-cutting endonuclease such as a meganuclease can enhance DNA binding and/or cleavage activity.
- Such a configuration can be achieved via (i) a single N- or C-terminal fusion to a homodimeric variant; (ii) a single N- or C-terminal fusion to individual monomers of a heterodimer, or; (iii) a double fusion to a monomeric protein.
- Figure 3B When specificity reengineering precludes maintaining cleavage activity of the rare- cutting endonuclease such as a meganuclease, the attached enhancer domains can provide alternative functions.
- Figure 3C) and (Figure 3D) represent instances of ( Figure 3A) and ( Figure 3B), respectively, when only one enhancer domain is needed or tolerated per fusion protein (e.g. either as an N- or C-terminal fusion or in the context of a single-chain molecule). Fusion junctions (N- vs. C- terminal) and linker designs can vary with the application.
- the present invention can relate to a method for increasing targeted H (and NHEJ) when efficiency is enhanced in a chimeric rare-cutting endonuclease targeting a DNA target sequence according to the invention.
- said efficiency can be enhanced by the addition of a catalytically active cleavase enhancer domain according to the invention, providing functional and/or structural support to the starting rare-cutting endonuclease scaffold according to the invention.
- said efficiency can be enhanced by the addition of two catalytically active cleavase enhancer domains according to the invention, providing functional and/or structural support to the starting rare-cutting endonuclease scaffold according to the invention.
- two cataiyticaily active cleavase enhancer domains can allow an increase in Double-strand break-induced mutagenesis by leading to a loss of genetic information between two nearby DNA double strand breaks and thus preventing any scarless re-ligation of targeted genomic locus of interest by NHEJ.
- the present invention can relate to a method for increasing targeted Homologous Recombination (HR) with less NHEJ (i.e in a more conservative fashion) when at least one single-strand break activity is promoted in a chimeric rare-cutting endonuclease targeting a DNA target sequence according to the invention.
- HR Homologous Recombination
- NHEJ i.e in a more conservative fashion
- the efficiency of said chimeric rare-cutting endonuclease could be enhanced by the addition of a cataiyticaily active nickase enhancer domain according to the invention.
- the efficiency of said chimeric rare-cutting endonuclease could be enhanced by the addition of two cataiyticaily active nickase enhancer domains according to the invention, all nickase enhancer domains being able to cut the same DNA strand. More preferably, the addition of a least one nickase enhancer domain according to the invention can be performed on an inactive rare- cutting endonuclease scaffold which serves as a scaffold for targeted DNA binding.
- the present invention can relate to a method for increasing excision of a single-strand of DNA spanning the starting rare-cutting endonuclease DNA binding region when both one cleavase enhancer domain and one nickase enhancer domain, respectively, are fused to both N-terminus and C-terminus of a starting rare-cutting endonuclease scaffold according to the invention.
- both cleavase enhancer domain and nickase enhancer domain can be fused on an inactive rare-cutting endonuclease scaffold which serves as a scaffold for targeted DNA binding.
- the present invention concerns a method for the creation of fusion proteins that consist of engineering a fusion protein between a rare-cutting endonuclease-derived scaffold and at least one enhancer domain wherein said enhancer domain enhances the efficiency of said chimeric rare-cutting endonuclease when fused to it.
- a method to create a chimeric rare-cutting endonuclease with enhanced efficiency for a DNA target sequence comprising the steps of:
- said enhancer domain is fused to the N-terminus part of said rare-cutting endonuclease-derived scaffold. In another preferred embodiment, said enhancer domain is fused to the C-terminus part of said rare-cutting endonuclease-derived scaffold. In another embodiment, two enhancer domains are fused to both N-terminus part of said rare-cutting endonuclease-derived scaffold and C-terminus part of said rare-cutting endonuclease-derived scaffold.
- said enhancer domain(s) are fused to N-terminus and/or C-terminus of a rare-cutting endonuclease-derived monomer scaffold of a homodimeric endonuclease, resulting in a chimeric rare-cutting endonuclease comprising two or four enhancer domains.
- said enhancer domain(s) are fused to N-terminus and/or C-terminus of a rare-cutting endonuclease- derived monomer scaffold of a heterodimeric endonuclease, resulting in a chimeric rare-cutting endonuclease comprising one or two or three or four enhancer domains.
- said enhancer domain(s) are fused to N-terminus and/or C-terminus of a rare-cutting endonuclease- derived scaffold from a single-chain endonuclease, resulting in a chimeric rare-cutting endonuclease comprising one or two enhancer domains.
- said enhancer domain is catalytically active or not, providing functional and/or structural support to said rare-cutting endonuclease-derived scaffold.
- said enhancer domain consists of a protein domain derived from an exonuclease.
- said enhancer domain consists of a protein domain derived from an endonuclease. In another more preferred embodiment, said enhancer domain consists of a protein domain derived from a cleavase. In another more preferred embodiment, said enhancer domain consists of a protein domain derived from a nickase.
- said enhancer domain consists of a protein domain derived from a protein selected from the group consisting of Mmel, Colicin-E7 (CEA7_ECOLX), EndA, Endo I (ENDl_ECOLI), Human Endo G (NUCG_HUMAN), Bovine Endo G (NUCG_BOVIN), R.HinPll, l-Basl, I- Bmol, l-Hmul, l-Tevl, l-Tevll, l-Tevlll, l-Twol, R.Mspl, R.Mval, NucA, NucM, Vvn, Vvn_CLS, Staphylococcal nuclease (NUC_STAAU), Staphylococcal nuclease (NUC_STAHY), Micrococcal nuclease (NUC_SHIFL), Endonuclease yncB, Endodeoxyribonuclease I (
- said enhancer domain consists of a peptide derived from CFPl peptide (SEQ ID NO: 112).
- any combinations of two protein domains selected from the group consisting of Mmel, Colicin-E7 (CEA7_ECOLX), EndA, Endo I (ENDl_ECOLI), Human Endo G (NUCG_HUMAN), Bovine Endo G (NUCG_BOVIN), R.HinPll, l-Basl, l-Bmol, l-Hmul, I- Tevl, l-Tevll, l-Tevlll, l-Twol, R.Mspl, R.Mval, NucA, NucM, Vvn, Vvn_CLS, Staphylococcal nuclease (NUC_STAAU), Staphylococcal nuclease (NUC_STAHY), Micrococcal nuclease (NUC_SHIFL), Endonu
- l-Hmul catalytic domain can be fused to the N-terminus part of said rare-cutting endonuclease-derived scaffold and ColE7 can be fused to the C-terminus part of said rare-cutting endonuclease-derived scaffold.
- ColE7 can be fused to the C-terminus part of said rare-cutting endonuclease-derived scaffold.
- said enhancer domain consists of a catalytically active derivative of the protein domains listed above and in Table 1, providing functional and/or structural support to said rare- cutting endonuclease-derived scaffold.
- said enhancer domain consists of a catalytically inactive derivative of the protein domains listed above and in Table 1, providing structural support to said rare-cutting endonuclease-derived scaffold.
- said rare-cutting endonuclease-derived scaffold is derived from a meganuclease.
- said meganuclease comprises two identical monomers.
- said meganuclease comprises two non-identical monomers.
- said meganuclease is a single-chain meganuclease.
- said rare-cutting endonuclease-derived scaffold is derived from the group consisting of !-Crel, a functional mutant of l-Crel, a variant of l-Crel or a derivative thereof.
- rare-cutting endonuclease-derived scaffold is a truncated form of wild- type l-Crel (SEQ ID NO: 1).
- rare-cutting endonuclease-derived scaffold comprises first 152, 153, 154 or 155 amino acids residues of wild-type l-Crel (SEQ ID NO: 1).
- rare-cutting endonuclease-derived scaffold comprises residues 2 to 153 of wild-type l-Crel (SEQ ID NO: 1).
- rare-cutting endonuclease-derived scaffold comprises residues 2 to 155 of wild-type I- Crel (SEQ ID NO: 1).
- rare-cutting endonuclease-derived scaffold comprises residues 2 to 153 of wild-type l-Crel (SEQ ID NO: 1) and one or several amino acids substitutions.
- rare-cutting endonuclease-derived scaffold comprises residues 2 to 155 of wild-type l-Crel (SEQ ID NO: 1) and one or several amino acids substitutions.
- rare-cutting endonuclease-derived scaffold of the present invention comprises residues 2 to 153 or residues 2 to 155 of wild-type l-Crel (SEQ ID NO: 1) and one or two or three or four or five or six or seven or eight or nine or ten further amino acid mutations.
- rare-cutting endonuclease-derived scaffold of the present invention comprises residues 2 to 153 of wild-type l-Crel (SEQ ID NO: 1) and K82A mutation.
- said rare-cutting endonuclease-derived scaffold comprises a sequence selected from the group consisting of l-Crel_NFSl (SEQ ID NO: 6); l-Crel_NFS2 (SEQ ID NO: 7); and l-Crel_CFSl (SEQ ID NO: 8).
- said chimeric rare-cutting endonuciease can comprise at least one peptidic linker between said rare-cutting endonuclease-derived scaffold and said at least one enhancer domain.
- said peptidic linker sequence is selected from the group consisting of 2012/032386
- the peptidic linker that can link said enhancer domain to the rare-cutting endonuclease-derived scaffold according to the method of the present invention can be selected from the group consisting of NFS1 (SEQ ID NO: 98), NFS2 (SEQ ID NO: 99) and CFS1 (SEQ ID NO: 100).
- NFS1 SEQ ID NO: 98
- NFS2 SEQ ID NO: 99
- CFS1 SEQ ID NO: 100
- Enhancement of efficiency of a chimeric rare-cutting endonuclease according to the present invention, compared to a starting rare-cutting endonuclease, can also be a consequence of a fuctional support by said at least one enhancer domain.
- said functional support can be the consequence of the hydrolysis of additional phosphodiester bonds.
- said functional support can be the hydrolysis of additional phosphodiester bonds by a protein domain derived from a nuclease.
- said functional support can be the hydrolysis of additional phosphodiester bonds by a protein domain derived from an endonuclease.
- said functional support can be the hydrolysis of additional phosphodiester bonds by a protein domain derived from a cleavase. In another more preferred embodiment, said functional support can be the hydrolysis of additional phosphodiester bonds by a protein domain derived from a nickase. In a more preferred embodiment, said functional support can be the hydrolysis of additional phosphodiester bonds by a protein domain derived from an exonuclease.
- Cleavage by rare-cutting endonucleases usually generates cohesive ends, with 3' overhangs for LAGLIDADG meganucleases ⁇ Chevalier, 2001 #6 ⁇ and 5' overhangs for Zinc Finger Nucleases ⁇ Smith, 2000 #7 ⁇ . These ends, which result from hydrolysis of phosphodiester bonds, can be re-ligated in vivo by NHEJ in a seamless way (i.e a scarless re-ligation).
- the restoration of a cleavable target sequence allows for a new cleavage event by the same endonuclease, and thus, a series of futile cycles of cleavage and re-ligation events can take place (Figure 4A).
- enhancement of efficiency of a chimeric rare-cutting endonuclease according to the present invention can be the hydrolysis of additional
- nickase or cleavase activities to the existing cleavase activity of a rare-cutting endonuclease enhances the efficiency of the resulting chimeric rare-cutting endonuclease according to the invention, at a genomic locus of interest.
- addition of two nickase activities on opposite strands (as depicted on Figure 4D) or of a new cleavase activity generating a second DSB ( Figure 4E) can result in a gap.
- perfect re-ligation is not possible anymore, and one or several alternative repair outcomes such as imprecise NHEJ, Homologous Recombination or SSA for instance, can be stimulated.
- the addition of a single nickase activity can result in a single strand gap, and suppress the cohesivity of the ends, which can also enhances the efficiency of the resulting chimeric rare-cutting endonuclease at a genomic locus of interest, according to the invention, via stimulation of one or several alternative repair outcomes mentioned above.
- the method according to the invention enhances rare-cutting endonuclease efficiency for a DNA target sequence.
- Enhancement of efficiency of a rare-cutting endonuclease according to the present invention can be a consequence of a structural support by said at least one enhancer domain.
- said structural support enhances the binding of a chimeric rare-cutting endonuclease according to the invention for a DNA target sequence compared to the binding of a starting rare-cutting endonuclease for the same DNA target sequence, as a non-limiting example.
- said structural support by at least one enhancer domain enhances the existing catalytical activity of a chimeric rare-cutting endonuclease according to the invention for a DNA target sequence compared to the binding of a starting rare-cutting endonuclease for the same DNA target sequence, as another non-limiting example.
- said at least one enhancer domain enhances both the binding and the existing catalytical activity of a chimeric rare-cutting endonuclease according to the invention for a DNA target sequence compared to the binding of a starting rare-cutting endonuclease for the same DNA target sequence, as another non-limiting example. All these non- limiting examples lead to a chimeric rare-cutting endonuclease with enhanced efficiency for a DNA target at a genomic locus of interest, according to the present invention.
- the resulting chimeric rare-cutting endonuclease can comprise several enzymatic activities that contribute to the enhancement of its efficiency according to the present invention.
- the addition of one nickase domain to an active rare-cutting endonuclease scaffold derived from a meganuclease results in a chimeric rare-cutting endonuclease with one meganuclease activity and one nickase activity (depicted in figure 4B).
- the addition of two nickase domains to an active rare-cutting endonuclease scaffold derived from a meganuclease results in a chimeric rare-cutting endonuclease with one meganuclease activity and two nickase activities (as depicted in figures 5C and 5D).
- the addition of one cleavase domain to an active rare-cutting endonuclease scaffold derived from a meganuclease results in a chimeric rare-cutting endonuclease with one meganuclease activity and one cleavase activity (figure 4E).
- the addition of two cleavase domains to an active rare-cutting endonuclease scaffold derived from a meganuclease results in a chimeric rare-cutting endonuclease with one meganuclease activity and two cleavase activities.
- enhancement of efficiency of a rare-cutting endonuclease refers to the increase in the detected level of said efficiency, against a target DNA sequence, of a second rare-cutting endonuclease in comparison to the activity of a first rare-cutting endonuclease against the same target DNA sequence.
- Said second rare-cutting endonuclease can be an engineered rare-cutting endonuclease, i.e. a chimeric rare-cutting endonuclease according to the invention or not.
- Said first rare-cutting endonuclease can be a wild-type rare-cutting endonuclease, a starting rare-cutting endonuclease, an engineered rare-cutting endonuclease i.e a chimeric rare-cutting endonuclease according to the invention or not, taken as a reference scaffold for enhancement in efficiency. Additional rounds of enhancement can be envisioned on a starting rare-cutting endonuclease scaffold.
- Said enhancement of efficiency can be determined, as non-limiting example, by measuring the level of cleavage-induced recombination generated by said rare-cutting endonuclease or said chimeric rare-cutting endonuclease according to the present invention. Said level of cleavage-induced recombination can be measured by a cell-based recombination assay as described in the International PCT Application WO 2004/067736, as non-limiting example.
- enhancement can be measured by
- 33 of efficacy in cells is not necessarily associated with an enhancement of the cleavage activity that could be detected in certain in vitro assays.
- additional phosphodiesterase activities as described in Figure 4 could barely affect the cleavage profile, as detected by in vitro cleavage and separation of the cleavage products on an electrophoresis gel.
- the DSB ends generated in this way could be more prone to induce detectable genomic rearrangements such as targeted mutagenesis (by imprecise NHEJ) or homologous recombination.
- Said enhancement in efficiency of said rare-cutting endonuclease is at least a 5% enhancement compared to the starting scaffold, more preferably at least a 10% enhancement, again more preferably at least a 15% enhancement, again more preferably at least a 20% enhancement, again more preferably at least a 25% enhancement, again more preferably a 50% enhancement, again more preferably a enhancement greater than 50%.
- the enhancement of efficiency of a rare-cutting endonuclease according to the method of the present invention can also be an enhancement in the efficiency for 2NN derivatives of a DNA target sequence by a rare-cutting endonuclease from the LAGLIDADG meganuclease subfamily.
- 2NN derivatives of a DNA target sequence is intended variants of a given DNA target sequence having mutations in the four central nucleotides -2 to +2 as numbered in figure 2.
- the method according to the invention enhances the efficiency for derivatives of the DNA target sequence C1221 having mutations in the four central nucleotides -2 to +2 as numbered in figure 2, i.e that do not have GTAC sequence at position -2 to +2 as numbered in figure 2.
- the method according to the invention allows recombination to be initiated at 2NN derivatives that are not normally processed by a starting rare-cutting endonuclease scaffold from the LAGLIDADG meganuclease subfamily.
- the method according to the invention allows the processing of 2NN derivatives that have -2 to +2 target sequences as listed in Table 4 and Table 5, non-originally cleaved by l-Crel.
- the present invention also concerns the creation of functional single polypeptide fusion proteins for simple and efficient vectorization.
- the present invention relates to chimeric rare-cutting endonucleases comprising at least an enhancer domain wherein said enhancer domain enhances the efficiency of said rare- cutting endonuclease when fused to it, thereby obtaining a chimeric rare-cutting endonuclease with enhanced efficiency for a DNA target sequence compared to a corresponding rare-cutting endonuclease lacking said enhancer domain.
- a chimeric rare-cutting endonuclease for a DNA target sequence comprising:
- an enhancer domain wherein said enhancer domain enhances the efficiency of said rare-cutting endonuclease for said DNA target sequence when fused to it;
- said enhancer domain is fused to the N-terminus part of said rare-cutting endonuclease-derived scaffold. In another preferred embodiment, said enhancer domain is fused to the C-terminus part of said rare-cutting endonuclease-derived scaffold. In another embodiment, two enhancer domains are fused to both N-terminus part of said rare-cutting endonuclease-derived scaffold and C-terminus part of said rare-cutting endonuclease-derived scaffold.
- said enhancer domain(s) are fused to N-terminus and/or C-terminus of a rare-cutting endonuclease-derived monomer scaffold of a homodimeric endonuclease, resulting in a chimeric rare-cutting endonuclease comprising two or four enhancer domains.
- said enhancer domain(s) are fused to N-terminus and/or C-terminus of a rare-cutting endonuclease- derived monomer scaffold of a heterodimeric endonuclease, resulting in a chimeric rare-cutting endonuclease comprising one or two or three or four enhancer domains.
- said enhancer domain(s) are fused to N-terminus and/or C-terminus of a rare-cutting endonuclease- derived scaffold from a single-chain endonuclease, resulting in a chimeric rare-cutting endonuclease comprising one or two enhancer domains.
- said enhancer domain is catalytically active, providing functional and/or structural support to said rare-cutting endonuclease-derived scaffold.
- said enhancer domain consists of a protein domain derived from an exonuclease.
- said enhancer domain consists of a protein domain derived
- said enhancer domain consists of a protein domain derived from a cleavase. In another more preferred embodiment, said enhancer domain consists of a protein domain derived from a nickase.
- said enhancer domain consists of a protein domain derived from a protein selected from the group consisting of Mmel, Colicin-E7 (CEA7_ECOLX), EndA, Endo I (ENDl_ECOLI), Human Endo G (NUCG_HUMAN), Bovine Endo G (NUCG_BOVIN), R.HinPll, l-Basl, I- Bmol, l-Hmul, l-Tevl, l-Tevll, l-Tevlll, l-Twol, R.Mspl, R.Mval, NucA, NucM, Vvn, Vvn_CLS, Staphylococcal nuclease (NUC_STAAU), Staphylococcal nuclease (NUC_STAHY), Micrococcal nuclease (NUC_SHIFL), Endonuclease yncB, Endodeoxyribonuclease I (
- said enhancer domain consists of a peptide derived from CFPl peptide (SEQ ID NO: 112).
- any combinations of two protein domains selected from the group consisting of Mmel, Colicin-E7 (CEA7_ECOLX), EndA, Endo I (ENDl_ECOLI), Human Endo G (NUCG_HUMAN), Bovine Endo G (NUCG_BOVIN), R.HinPll, l-Basl, l-Bmol, l-Hmul, I- Tevl, l-Tevll, l-Tevlll, l-Twol, R.Mspl, R.Mval, NucA, NucM, Vvn, Vvn_CLS, Staphylococcal nuclease (NUC_STAAU), Staphylococcal nuclease (NUC_STAHY), Micrococcal nuclease (NUC_SHIFL), Endonu
- l-Hmul catalytic domain can be fused to the N-terminus part of said rare-cutting endonuclease-derived scaffold and ColE7 can be fused to the C-terminus part of said rare-cutting endonuclease-derived scaffold.
- said enhancer domain consists of a catalytically active derivative of the protein domains listed above
- said enhancer domain consists of a catalytically inactive derivative of the protein domains listed above and in Table 1, providing structural support to said rare-cutting endonuclease-derived scaffold.
- said rare-cutting endonuclease-derived scaffold is derived from a meganuclease.
- said meganuclease comprises two identical monomers.
- said meganuclease comprises two non-identical monomers.
- said meganuclease is a single-chain meganuclease.
- said rare-cutting endonuclease-derived scaffold is derived from the group consisting of l-Crel, a functional mutant of l-Crel, a variant of l-Crel or a derivative thereof.
- rare-cutting endonuclease-derived scaffold is a truncated form of wild- type l-Crel (SEQ ID NO: 1).
- rare-cutting endonuclease-derived scaffold comprises first 152, 153, 154 or 155 amino acids residues of wild-type l-Crel (SEQ ID NO: 1).
- rare-cutting endonuclease-derived scaffold comprises residues 2 to 153 of wild-type l-Crel (SEQ ID NO: 1).
- rare-cutting endonuclease-derived scaffold comprises residues 2 to 155 of wild-type I- Crel (SEQ ID NO: 1).
- rare-cutting endonuclease-derived scaffold comprises residues 2 to 153 of wild-type l-Crel (SEQ ID NO: 1) and one or several amino acids substitutions.
- rare-cutting endonuclease-derived scaffold comprises residues 2 to 155 of wild-type l-Crel (SEQ ID NO: 1) and one or several amino acids substitutions.
- rare-cutting endonuclease-derived scaffold of the present invention comprises residues 2 to 153 or residues 2 to 155 of wild-type l-Crel (SEQ ID NO: 1) and one or two or three or four or five or six or seven or eight or nine or ten further amino acid mutations.
- rare-cutting endonuclease-derived scaffold of the present invention comprises residues 2 to 153 of wild-type l-Crel (SEQ ID NO: 1) and K82A mutation.
- said rare-cutting endonuclease-derived scaffold comprises a sequence selected from the group consisting of l-Crel_NFSl (SEQ ID NO: 6); l-Crel_NFS2 (SEQ ID NO: 7); and l-Crel_CFSl (SEQ ID NO: 8).
- said chimeric rare-cutting endonuclease can optionally comprise at least one peptidic linker between said rare-cutting endonuclease-derived scaffold and said at least one enhancer domain.
- said peptidic linker sequence is selected from
- the peptidic linker that can link said enhancer domain to the rare-cutting endonuclease-derived scaffold according to the present invention can be selected from the group consisting of NFS1 (SEQ ID NO: 98), NFS2 (SEQ ID NO: 99) and CFS1 (SEQ ID NO: 100).
- NFS1 SEQ ID NO: 98
- NFS2 SEQ ID NO: 99
- CFS1 SEQ ID NO: 100
- Enhancement of efficiency of a chimeric rare-cutting endonuclease according to the present invention, compared to a starting rare-cutting endonuclease, can also be a consequence of a fuctional support by said at least one enhancer domain.
- said functional support can be the consequence of the hydrolysis of additional phosphodiester bonds.
- said functional support can be the hydrolysis of additional phosphodiester bonds by a protein domain derived from a nuclease.
- said functional support can be the hydrolysis of additional phosphodiester bonds by a protein domain derived from an endonuclease.
- said functional support can be the hydrolysis of additional phosphodiester bonds by a protein domain derived from a cleavase. In another more preferred embodiment, said functional support can be the hydrolysis of additional phosphodiester bonds by a protein domain derived from a nickase. In a more preferred embodiment, said functional support can be the hydrolysis of additional phosphodiester bonds by a protein domain derived from an exonuclease.
- Cleavage by rare-cutting endonucleases usually generates cohesive ends, with 3' overhangs for LAGLIDADG meganucleases ⁇ Chevalier, 2001 #6 ⁇ and 5' overhangs for Zinc Finger Nucleases ⁇ Smith, 2000 #7 ⁇ . These ends, which result from hydrolysis of phosphodiester bonds, can be re-ligated in vivo by NHEJ in a seamless way (i.e a scarless re-ligation).
- the restoration of a cleavable target sequence allows for a new cleavage event by the same endonuclease, and thus, a series of futile cycles of cleavage and re-ligation events can take place (Figure 4A).
- enhancement of efficiency of a chimeric rare-cutting endonuclease according to the present invention can be the hydrolysis of additional
- nickase or cleavase activities to the existing cleavase activity of a rare-cutting endonuclease enhances the efficiency of the resulting chimeric rare-cutting endonuclease according to the invention, at a genomic locus of interest.
- addition of two nickase activities on opposite strands (as depicted on Figure 4D) or of a new cleavase activity generating a second DSB ( Figure 4E) can result in a gap.
- perfect re-ligation is not possible anymore, and one or several alternative repair outcomes such as imprecise NHEJ, Homologous Recombination or SSA for instance, can be stimulated.
- the addition of a single nickase activity can result in a single strand gap, and suppress the cohesivity of the ends, which can also enhances the efficiency of the resulting chimeric rare-cutting endonuclease at a genomic locus of interest, according to the invention, via stimulation of one or several alternative repair outcomes mentioned above.
- chimeric rare-cutting endonucleases according to the invention enhances rare-cutting endonuclease efficiency for a DNA target sequence. Enhancement of efficiency of a rare-cutting endonuclease according to the present invention can be a consequence of a structural support by said at least one enhancer domain.
- said structural support enhances the binding of a chimeric rare-cutting endonuclease according to the invention for a DNA target sequence compared to the binding of a starting rare-cutting endonuclease for the same DNA target sequence, as a non-limiting example.
- said structural support by at least one enhancer domain enhances the existing catalytical activity of a chimeric rare-cutting endonuclease according to the invention for a DNA target sequence compared to the binding of a starting rare-cutting endonuclease for the same DNA target sequence, as another non-limiting example.
- said at least one enhancer domain enhances both the binding and the existing catalytical activity of a chimeric rare-cutting endonuclease according to the invention for a DNA target sequence compared to the binding of a starting rare-cutting endonuclease for the same DNA target sequence, as another non- limiting example. All these non-limiting examples lead to a chimeric rare-cutting endonuclease with
- the resulting chimeric rare-cutting endonuclease can comprise several enzymatic activities that contribute to the enhancement of its efficiency according to the present invention.
- the addition of one nickase domain to an active rare-cutting endonuclease scaffold derived from a meganuclease results in a chimeric rare-cutting endonuclease with one meganuclease activity and one nickase activity (depicted in figure 4B).
- the addition of two nickase domains to an active rare-cutting endonuclease scaffold derived from a meganuclease results in a chimeric rare-cutting endonuclease with one meganuclease activity and two nickase activities (as depicted in figures 5C and 5D).
- the addition of one cleavase domain to an active rare-cutting endonuclease scaffold derived from a meganuclease results in a chimeric rare-cutting endonuclease with one meganuclease activity and one cleavase activity (figure 4E).
- the addition of two cleavase domains to an active rare-cutting endonuclease scaffold derived from a meganuclease results in a chimeric rare-cutting endonuclease with one meganuclease activity and two cleavase activities.
- enhancement of efficiency of a rare-cutting endonuclease refers to the increase in the detected level of said efficiency, against a target DNA sequence, of a second rare-cutting endonuclease in comparison to the activity of a first rare-cutting endonuclease against the same target DNA sequence.
- Said second rare-cutting endonuclease can be an engineered rare-cutting endonuclease, i.e. a chimeric rare-cutting endonuclease according to the invention or not.
- Said first rare-cutting endonuclease can be a wild-type rare-cutting endonuclease, a starting rare-cutting endonuclease, an engineered rare-cutting endonuclease i.e a chimeric rare-cutting endonuclease according to the invention or not, taken as a reference scaffold for enhancement in efficiency. Additional rounds of enhancement can be envisioned on a starting rare-cutting endonuclease scaffold.
- Said enhancement of efficiency can be determined, as non-limiting example, by measuring the level of cleavage-induced recombination generated by said rare-cutting endonuclease or said chimeric rare-cutting endonuclease according to the present invention. Said level of cleavage-induced
- enhancement of efficacy in cells can be, but is not necessarily associated with an enhancement of the cleavage activity that could be detected in certain in vitro assays.
- additional phosphodiesterase activities as described in Figure 4 could barely affect the cleavage profile, as detected by in vitro cleavage and separation of the cleavage products on an electrophoresis gel.
- the DSB ends generated in this way could be more prone to induce detectable genomic Rearrangements such as targeted mutagenesis (by imprecise NHEJ) or homologous recombination.
- Said enhancement in efficiency of said rare-cutting endonuclease is at least a 5% enhancement compared to the starting scaffold, more preferably at least a 10% enhancement, again more preferably at least a 15% enhancement, again more preferably at least a 20% enhancement, again more preferably at least a 25% enhancement, again more preferably a 50% enhancement, again more preferably a enhancement greater than 50%.
- the enhancement of efficiency of a chimeric rare-cutting endonuclease according to the present invention can also be an enhancement in the efficiency for 2NN derivatives of a DNA target sequence by a rare-cutting endonuclease from the LAGLIDADG meganuclease subfamily.
- 2NN derivatives of a DNA target sequence is intended variants of a given DNA target sequence having mutations in the four central nucleotides -2 to +2 as numbered in figure 2.
- a chimeric rare-cutting endonuclease according to the invention can enhance the efficiency for derivatives of the DNA target sequence C1221 having mutations in the four central nucleotides -2 to +2 as numbered in figure 2, i.e that do not have GTAC sequence at position -2 to +2 as numbered in figure 2.
- the chimeric rare-cutting endonuclease according to the invention can allow the processing of 2NN derivatives that are not efficiently processed by a starting rare-cutting endonuclease scaffold from the LAGLIDADG meganuclease subfamily.
- the chimeric rare-cutting endonuclease according to the invention allows the induction of recombination by processing 2NN derivatives that have -2 to +2 target sequences as listed in Table 4 and Table 5, non-originally cleaved by l-Crel.
- said chimeric rare-cutting endonuclease is selected from the group consisting of hHmuCre_D0101 (SEQ ID NO: 9), hHmuCre_D0201 (SEQ ID NO: 105), hHmuCre_D0301 (SEQ ID NO: 106), hHmuCre_D0401 (SEQ ID NO: 107), hHmuCre_D0102 (SEQ ID NO: 9), hHmuCre_D0201 (SEQ ID NO: 105), hHmuCre_D0301 (SEQ ID NO: 106), hHmuCre_D0401 (SEQ ID NO: 107), hHmuCre_D0102 (SEQ ID NO: 9), hHmuCre_D0201 (SEQ ID NO: 105), hHmuCre_D0301 (SEQ ID NO: 106), hHmuCre_D0401 (SEQ
- the present invention relates to a method for treatment of a genetic disease caused by a mutation in a specific single double-stranded DNA target sequence in a gene, comprising administering to a subject in need thereof an effective amount of a chimeric rare-cutting endonuclease according to the present invention.
- the present invention relates to a method for inserting a transgene into a specific single double-stranded DNA target sequence of a genomic locus of a cell, tissue or non-human animal wherein at least one chimeric rare-cutting endonuclease of the present invention is introduced in said cell, tissue or non-human animal.
- the chimeric rare-cutting endonuclease of the present invention and a pharmaceutically acceptable excipient are administered in a therapeutically effective amount.
- Such a combination is said to be administered in a "therapeutically effective amount” if the amount administered is physiologically significant.
- An agent is physiologically significant if its presence results in a detectable change in the physiology of the recipient.
- an agent is physiologically significant if its presence results in a decrease in the severity of one or more symptoms of the targeted disease and in a genome correction of the lesion or abnormality.
- Vectors comprising targeting DNA and/or nucleic acid encoding a chimeric rare-cutting endonuclease can be introduced into a cell by a variety of methods (e.g., injection, direct uptake, projectile bombardment, liposomes, electroporation). Chimeric rare-cutting endonucleases can be stably or transiently expressed into cells using expression vectors. Techniques of expression in eukaryotic cells are well known to those in the art. (See Current Protocols in Human Genetics: Chapter 12 "Vectors For Gene Therapy” & Chapter 13 "Delivery Systems for Gene Therapy”).
- the chimeric rare-cutting endonuclease of the present invention is substantially non-immunogenic, i.e., engender little or no adverse immunological response.
- a variety of methods for ameliorating or eliminating deleterious immunological reactions of this sort can be used in accordance with the invention.
- the chimeric rare-cutting endonuclease is substantially free of N-formyl methionine.
- Another way to avoid unwanted immunological reactions is to conjugate chimeric rare-cutting endonuclease to polyethylene glycol (“PEG”) or polypropylene glycol (“PPG”) (preferably of 500 to 20,000 daltons average molecular weight (MW)). Conjugation with PEG or PPG, as described by
- the present invention relates to recombinant polynucleotides encoding the chimeric rare-cutting endonucleases of the present invention, specific vectors (polynucleotidic or not) encoding and/or vectorizing them, compositions and/or kits comprising them, all of them beign used or part of a whole to implement methods of the present invention for enhancing rare-cutting endonuclease cleavage activity at a genomic locus of interest in a cell.
- - Amino acid substitution means the replacement of one amino acid residue with another, for instance the replacement of an Arginine residue with a Glutamine residue in a peptide sequence is an amino acid substitution.
- Efficiency of a rare-cutting endonuclease is the property for said rare-cutting endonuclease of producing a desired event.
- This desired event can be for example Homologous gene targeting, targeted mutagenesis, or sequence removal or excision.
- the efficiency of the desired event depends on several parameters, including the specific activity of the nuclease and the repair pathway resulting in the desired event (efficacy of homologous repair for gene targeting, efficacy and outcome of NHEJ pathways for targeted mutagenesis).
- Efficiency of a rare cutting endonuclease for a locus is intended to mean its abiliy to produce a desired event at this locus.
- Efficiency of a rare cutting endonuclease for a target is intended to mean its abiliy to produce a desired event as a consequence of cleavage of this target.
- Enhancement of efficiency of a rare-cutting endonuclease by creating a chimeric rare- cutting endonuclease with enhanced efficiency according to the present invention is measurable comparatively to the efficiency of the starting rare-cutting endonuclease scaffold in the same conditions.
- Enhancement of efficiency of a chimeric rare-cutting endonuclease can be the consequence of an enhancement of an individual structural parameter or an enhancement of a combination of several individual structural parameters consecutive to the addition of an enhancer domain according to the present invention.
- Such parameters can be as non-limiting examples, binding affinity or capacity of binding, cleavage activity, turn-over of one enzyme (Kcat), Kcat/Km (incorporating the different rate constants of an enzymatic reaction).
- Enhancement of efficiency of a chimeric rare-cutting endonuclease can also be a functional consequence to the addition of an enhancer domain according to the present invention.
- Said additional enhancer domain can provide additional phosphodiester bonds hydrolysis, via cleavase or nickase domains as non-limiting examples, allowing to favour or stimulate one or several alternative repair outcomes.
- enhancement of efficiency can also be an enhancement in th eefficiency for 2NN derivatives of DNA target sequence originally targeted by the parent rare-cutting endonuclease.
- an enhancer domain can provide additional phosphodiester bonds hydrolysis, via cleavase or nickase domains as non-limiting examples, allowing to favour or stimulate one or several alternative repair outcomes.
- enhancement of efficiency can also be an enhancement in th eefficiency for 2NN derivatives of DNA target sequence originally targeted by the parent rare-cutting endonuclease.
- an enhancer domain can provide additional phosphodiester bonds hydrolysis, via cleavase or nick
- enhancement of the efficiency of chimeric rare-cutting endonucleases derived from LAGLIDADG meganuclease subfamily can be the capacity to process 2NN derivatives that are not cleaved in a detectable way by the parent rare-cutting endonuclease.
- enhancement of efficiency of a rare- cutting endonuclease according to the present invention can be measured from an application point of view.
- Chimeric rare-cutting endonucleases according to the present invention being usable for various genome engineering applications, said chimeric rare-cutting endonucleases according to the invention can have an enhanced efficiency for NHEJ, HR, SSA, as non-limiting examples.
- Said chimeric rare-cutting endonucleases according to the invention can also have an enhanced efficiency for a precise genome engineering such as gene targeting, targeted DNA cleavage or targeted gene repression as non-limiting examples.
- Enhanced/increased/improved cleavage activity or binding refers to an increase in the detected level of an endonuclease, a rare-cutting endonuclease or a chimeric rare-cutting endonuclease cleavage activity or binding, see below, against a target DNA sequence or DNA target sequence by a second endonuclease, rare-cutting endonuclease or chimeric rare-cutting endonuclease in comparison to the cleavage activity or binding of a first endonuclease or a wild-type endonuclease (non chimeric) or a first rare-cutting endonuclease or a first chimeric rare-cutting endonuclease against said target DNA sequence.
- the second endonuclease or rare-cutting endonuclease or chimeric rare-cutting endonuclease can be a variant of the first one and can comprise one or more substituted amino acid residues in comparison to the first endonuclease or rare-cutting endonuclease or chimeric rare-cutting endonuclease or additional peptidic sequences or domains. Parameters in this definition are included in the method for enhancing rare-cutting endonuclease efficiency according to the present invention.
- nucleosides are designated as follows: one-letter code is used for designating the base of a nucleoside: a is adenine, t is thymine, c is cytosine, and g is guanine.
- r represents g or a (purine nucleotides)
- k represents g or t
- s represents g or c
- w represents a or t
- m represents a or c
- y represents t or c (pyrimidine nucleotides)
- d represents g, a or t
- v represents g, a or c
- b represents g, t or c
- h represents a, t or c
- n represents g, a, t or c.
- meganuclease is intended a rare-cutting endonuclease subtype having a double- stranded DNA target sequence greater than 12 bp.
- Said meganuclease is either a dimeric enzyme, wherein each domain is on a monomer or a monomeric enzyme comprising the two domains on a single polypeptide.
- “meganuclease domain” is intended the region which interacts with one half of the DNA target of a meganuclease and is able to associate with the other domain of the same meganuclease
- the starting endonuclease, rare-cutting endonuclease, chimeric rare-cutting endonuclease or meganuclease also the starting scaffold with at least a different amino acid or additional peptidic sequences or domains.
- peptide linker by "peptide linker", “peptidic linker” or “peptide spacer” it is intended to mean a peptide sequence which allows the connection of different monomers in a fusion protein and the adoption of the correct conformation for said fusion protein activity and which does not alter the specificity of either of the monomers for their targets.
- Peptide linkers can be of various sizes, from 3 amino acids to 50 amino acids as a non limiting indicative range. Peptide linkers can also be structured or unstructured.
- -variants or mutants of the structural elements of the invention including a rare-cutting endonuclease-derived scaffold, an enhancer domain, or a peptidic linker are contemplated.
- these will be at least 80, 90, 95, 98, or 99% identical to a known element, such as those described herein, or have 1, 2, 3, 4, 5, 6, 7, 8, 9, 10 or more amino acid insertions, deletions or substitutions compared to a corresponding element described herein.
- one cell type related to the chosen cell type or organism is intended a cell type or an organism sharing characteristics with said chosen cell type or said chosen organism; this cell type or organism related to the chosen cell type or organism, can be derived from said chosen cell type or organism or not.
- targeting DNA construct/minimal repair matrix/repair matrix it is intended to mean a DNA construct comprising a first and second portions that are homologous to regions 5' and 3' of the DNA target in situ.
- the DNA construct also comprises a third portion positioned between the first and second portion which comprise some homology with the corresponding DNA sequence in situ or alternatively comprise no homology with the regions 5' and 3' of the DNA target in situ.
- a homologous recombination event is stimulated between the genome containing the targeted gene comprised in the locus of interest and the repair matrix, wherein the
- a functional variant is intended a catalytically active variant of a protein, such variant can have additional properties compared to its parent protein.
- a functional variant of a meganuclease can be able to cleave additional DNA target sequences that are not cleaved by the parent meganuclease.
- a meganuclease variant which is created from a parent meganuclease and hence the peptide sequence of the meganuclease variant is related to (primary sequence level) but derived from (mutations) the peptide sequence of the parent meganuclease.
- mutations encompass deletions or insertions of several amino acid residues; as non-limiting example, a truncated variant of an l-Crel meganuclease is considered as a scaffold derived from l-Crel meganuclease.
- This expression can be applied to an endonuclease, a rare-cutting endonuclease or a chimeric rare-cutting endonuclease.
- l-Crel is intended the wild-type l-Crel having the sequence of pdb accession code lg9y, corresponding to the sequence SEQ ID NO: 1 in the sequence listing.
- Application l-Cre ⁇ variants described can comprise an additional Alanine after the first Methionine of the wild type l-Cre ⁇ sequence (SEQ ID NO: 1). These variants may also comprise two additional Alanine residues and an Aspartic Acid residue after the final Proline of the wild type l-Crei sequence as shown in SEQ ID NO: 4.
- l-Crel site is intended a 22 to 24 bp double-stranded DNA sequence which is cleaved by l-Crel.
- l-Crel sites include the wild-type non-palindromic l-Crel homing site and the derived palindromic sequences such as the sequence 5'- t.nCna.joa.ga.ga.yC.eg.st ⁇ c.Bg ⁇ t. ia +1 c +2 g + 3a + C + 5g +6 t + 7t + 8t +9 t + i 0 g + 1 ia +12 (SEQ ID NO: 2), also called C1221.
- domain or “core domain” is intended the "LAG LIDADG homing endonuclease core domain” which is the characteristic ⁇ fold of the homing endonucleases of the LAGLIDADG family, corresponding to a sequence of about one hundred amino acid residues.
- Said domain comprises four beta-strands ( ⁇ , ⁇ folded in an anti-parallel beta-sheet which interacts with one half of the DNA target.
- This domain is able to associate with another LAGLIDADG homing endonuclease core domain which interacts with the other half of the DNA target to form a functional endonuclease able to cleave said DNA target.
- the dimeric homing the dimeric homing
- LAGLIDADG homing endonuclease core domain corresponds to the residues 6 to 94.
- subdomain is intended the region of a LAGLIDADG homing endonuclease core domain which interacts with a distinct part of a homing endonuclease DNA target half-site.
- beta-hairpin is intended two consecutive beta-strands of the antiparallel beta-sheet of a LAGLIDADG homing endonuclease core domain ( ⁇ 2 0 ⁇ ⁇ 3 ⁇ 4 ) which are connected by a loop or a turn,
- single-chain meganuclease is intended a meganuclease comprising two LAGLIDADG homing endonuclease domains or core domains linked by a peptidic linker or peptidic spacer.
- the single-chain meganuclease is able to cleave a chimeric DNA target sequence comprising one different half of each parent meganuclease target sequence.
- This definition can be enlarged to "single-chain endonuclease” or “single-chain endonuclease derivative” that qualify two endonuclease domains or core domains from non-LAGLIDADG endonucleases linked by a peptidic linker or peptidic spacer.
- cleavage site is intended a double-stranded palindromic, partially palindromic (pseudo-palindromic) or non-palindromic polynucleotide sequence that is recognized and cleaved by a LAGLIDADG homing endonuclease such as l-Crel, or a variant, or a single-chain chimeric meganuclease derived from l-Crel.
- Said DNA target sequence is qualified as "cleavable" by an endonuclease, rare-cutting endonuclease, chimeric rare- cutting endonuclease or meganuclease when recognized within a genomic sequence and known to correspond to the DNA target sequence of a given endonuclease, rare-cutting endonuclease, chimeric rare-cutting endonuclease or meganuclease or a variant of such endonuclease, rare-cutting endonuclease, chimeric rare-cutting endonuclease or meganuclease.
- a distinct DNA location preferably a genomic location but also a portion of genetic material that can exist independently to the main body of genetic material such as plasmids, episomes, virus, transposons or in organelles such as mitochondria or chloroplasts as non-limiting examples, at which at least a double-strand break (cleavage) or a single strand break (nick) is to be induced by the meganuclease, endonuclease, rare-cutting endonuclease or chimeric rare-cutting endonuclease.
- the DNA target is defined by the 5' to 3' sequence of one strand of the double-stranded polynucleotide, as indicate above for C1221 (SEQ ID NO: 2, figure 2). Cleavage of the DNA target occurs at the nucleotides at positions +2 and -2, respectively for the
- Endonuclease refers to any wild-type or variant enzyme capable of catalyzing the hydrolysis (cleavage) of bonds between nucleic acids within a DNA or NA molecule, preferably a DNA molecule.
- Endonucleases can be classified as rare-cutting endonucleases when having typically a polynucleotide recognition site greater than 12base pairs (bp) in length, more preferably of 14-45 bp.
- Rare-cutting endonucleases significantly increase HR by inducing DNA double-strand breaks (DSBs) at a defined locus (Rouet, Smih et al. 1994; Rouet, Smih et al.
- Rare-cutting endonucleases can for example be a homing endonuclease (Paques and Duchateau 2007), a chimeric Zinc-Finger nuclease (ZFN) resulting from the fusion of engineered zinc-finger domains with the catalytic domain of a restriction enzyme such as Fokl (Porteus and Carroll 2005) or a chemical endonuclease (Eisenschmidt, Lanio et al. 2005 ; Arimondo, Thomas et al. 2006; Simon, Cannata et al. 2008).
- a chemical or peptidic cleaver is conjugated either to a polymer of nucleic acids or to another DNA recognizing a specific target sequence, thereby targeting the cleavage activity to a specific sequence.
- Chemical endonucleases also encompass synthetic nucleases like conjugates of orthophenanthroline, a DNA cleaving molecule, and triplex-forming oligonucleotides (TFOs), known to bind specific DNA sequences (Kalish and Glazer 2005). Such chemical endonucleases are comprised in the term "endonuclease" according to the present invention.
- Rare-cutting endonucleases can also be for example TALENs, a new class of chimeric nucleases using a Fokl catalytic domain and a DNA binding domain derived from Transcription Activator Like Effector (TALE), a family of proteins used in the infection process by plant pathogens of the Xanthomonas genus (Boch, Scholze et al. 2009; Moscou and Bogdanove 2009; Christian, Cermak et al. 2010; Li, Huang et al. 2010).
- TALE Transcription Activator Like Effector
- the functional layout of a Fokl-based TALE-nuclease (TALEN) is essentially that of a ZFN, with the Zinc-finger DNA binding domain being replaced by the TALE domain.
- DNA cleavage by a TALEN requires two DNA recognition regions flanking an unspecific central region.
- Rare-cutting endonucleases encompassed in the present invention can also be derived from
- Rare-cutting endonuclease can be a homing endonuclease, also known under the name of meganuclease. Such homing endonucleases are well-known to the art (Stoddard 2005). Homing endonucleases recognize a DNA target sequence and generate a single- or double-strand break. Homing endonucleases are highly specific, recognizing DNA target sites ranging from 12 to 45 base pairs (bp) in length, usually ranging from 14 to 40 bp in length. The homing endonuclease according to the art (Stoddard 2005). Homing endonucleases recognize a DNA target sequence and generate a single- or double-strand break. Homing endonucleases are highly specific, recognizing DNA target sites ranging from 12 to 45 base pairs (bp) in length, usually ranging from 14 to 40 bp in length. The homing endonuclease according to the art (Stodd
- 49 12 032386 to the invention may for example correspond to a LAGLIDADG endonuclease, to a HNH endonuclease, or to a GIY-YIG endonuclease.
- HEs Homing Endonucleases
- proteins families Cholier and Stoddard 2001.
- homing the endonuclease cleaves a cognate allele from which the mobile element is absent, thereby stimulating a homologous recombination event that duplicates the mobile DNA into the recipient locus.
- LAGLIDADG The LAGLIDADG family, named after a conserved peptidic motif involved in the catalytic center, is the most widespread and the best characterized group. Many structures are now available. Whereas most proteins from this family are monomeric and display two LAGLIDADG motifs, a few have only one motif, and thus dimerize to cleave palindromic or pseudo-palindromic target sequences.
- LAGLIDADG peptide is the only conserved region among members of the family, these proteins share a very similar architecture.
- the catalytic core is flanked by two DNA- binding domains with a perfect two-fold symmetry for homodimers such as l-Crel (Chevalier, Monnat et al. 2001), l- sol (Chevalier, Turmel et al. 2003) and l-Ceul (Spiegel, Chevalier et al. 2006) and with a pseudo symmetry for monomers such as l-Scel (Moure, Gimble et al. 2003), l-Dmol (Silva, Dalgaard et al.
- residues 28 to 40 and 44 to 77 of l-Crel were shown to form two partially separable functional subdomains, able to bind distinct parts of a homing endonuclease target half- site (Smith, Grizot et al. 2006); International PCT Applications WO 2007/049095 and WO 2007/057781 (Cellectis)).
- the combination of the two former steps allows a larger combinatorial approach, involving four different subdomains.
- the different subdomains can be modified separately and combined to obtain an entirely redesigned meganuclease variant (heterodimer or single-chain molecule) with chosen specificity.
- couples of novel meganucleases are combined in new molecules
- half-meganucleases cleaving palindromic targets derived from the target one wants to cleave. Then, the combination of such "half-meganucleases” can result in a heterodimeric species cleaving the target of interest.
- Examples of such endonuclease include l-Sce I, l-Chu I, l-Cre I, l-Csm I, Pl-Sce I, Pl-Tli I, Pl-Mtu I, l-Ceu I, l-Sce II, l-Sce III, HO, Pl-Civ I, Pl-Ctr I, Pl-Aae I, Pl-Bsu I, Pl-Dha I, Pl-Dra I, Pl-Mav I, Pl-Mch I, Pl-Mfu I, Pl-Mfl I, Pl-Mga I, Pl-Mgo I, Pl-Min I, Pl-Mka I, Pl-Mle I, Pl-Mma I, Pl-Msh I, Pl-Msm I, Pl-Mth I, Pl-Mtu I, Pl-Mxe I, Pl-Npu I, Pl-Pfu I, Pl-Rma I, Pl-Spb I, Pl-Ss
- a homing endonuclease can be a LAGLIDADG endonuclease such as ⁇ -Scel, ⁇ -Crel, ⁇ -Ceul, I- Msol, and ⁇ -Dmol.
- Said LAGLIDADG endonuclease can be l-Sce I, a member of the family that contains two LAGLIDADG motifs and functions as a monomer, its molecular mass being approximately twice the mass of other family members like l-Crel which contains only one LAGLIDADG motif and functions as homodimers.
- Meganucleases, endonucleases or rare-cutting endonucleases mentioned in the present application encompass both wild-type (naturally-occurring) and variant meganucleases, endonucleases or rare-cutting endonucleases.
- Meganucleases, endonucleases or rare-cutting endonucleases according to the invention can be a "variant" meganuclease, endonuclease or rare- cutting endonuclease, i.e.
- a meganuclease, endonuclease or rare-cutting endonuclease that does not naturally exist in nature and that is obtained by genetic engineering or by random mutagenesis, i.e. an engineered meganuclease, endonuclease or rare-cutting endonuclease.
- This variant meganuclease, endonuclease or rare-cutting endonuclease can for example be obtained by substitution of at least one residue in the amino acid sequence of a wild-type, naturally-occurring, meganuclease, endonuclease or rare-cutting endonuclease with a different amino acid.
- substitution(s) can for example be introduced by site-directed mutagenesis and/or by random mutagenesis.
- such variant meganucleases, endonucleases or rare-cutting endonucleases remain functional, i.e. they retain the capacity of recognizing (binding
- Meganuclease part, endonuclease part or rare-cutting endonuclease part in a chimeric rare- cutting endonuclease according to the present invention can only retain the capacity of recognizing a target sequence, in particular case, the capacity of cleaving a target sequence being provided by an enhancer domain according to the present invention.
- the variant meganuclease, endonuclease or rare-cutting endonuclease according to the invention cleaves a target sequence that is different from the target sequence of the corresponding wild-type meganuclease, endonuclease or rare-cutting endonuclease.
- Methods for obtaining such variant endonucleases with novel specificities are well-known in the art.
- Endonucleases variants may be homodimers (meganuclease comprising two identical monomers) or heterodimers (meganuclease comprising two non-identical monomers). It is understood that the scope of the present invention also encompasses endonuclease variants per se, including heterodimers (WO2006097854), obligate heterodimers (WO2008093249) and single chain meganucleases (WO03078619 and WO2009095793) as non limiting examples, able to cleave one target of interest in a polynucleotide sequence or in a genome and derivatives of these variants according to the present invention, such as chimeric rare-cutting endonucleases derived from these variants.
- the invention also encompasses hybrid variant per se composed of two monomers from different origins (WO03078619).
- Meganucleases, endonucleases or rare-cutting endonucleases with novel specificities can be used in the method according to the present invention for gene targeting and thereby integrating a transgene of interest into a genome at a predetermined location.
- parent meganuclease it is intended to mean a wild type meganuclease or a variant of such a wild type meganuclease with identical properties or alternatively a meganuclease with some altered characteristic in comparison to a wild type version of the same meganuclease.
- This expression can be applied to an endonuclease, a rare-cutting endonuclease or a chimeric rare- cutting endonuclease.
- delivery vector or “ delivery vectors” is intended any delivery vector which can be used in the present invention to put into cell contact ( i.e “contacting") or deliver inside cells or subcellular compartments agents/chemicals and molecules (proteins or nucleic acids) needed in the present invention, such as meganucleases, endonucleases, rare-cutting endonucleases or chimeric rare-cutting endonucleases or plasmidic vectors encoding said meganucleases, endonucleases, rare-cutting endonucleases or chimeric rare-cutting endonucleases, or repair matrix per se or encoded in a plasmidic vector for a gene engineering application for instance.
- liposomal delivery vectors viral delivery vectors, drug delivery vectors, chemical carriers, polymeric carriers, lipoplexes, polyplexes, dendrimers, microbubbles (ultrasound contrast agents), nanoparticles, emulsions or other appropriate transfer vectors.
- delivery vectors allow delivery of molecules, chemicals, macromolecules (genes, proteins), or other vectors such as plasmids, peptides developed by Diatos. In these cases, delivery vectors are molecule carriers.
- delivery vector or “delivery vectors” is also intended delivery methods to perform transfection.
- vector refers to a nucleic acid molecule capable of transporting another nucleic acid to which it has been linked.
- a “vector” in the present invention includes, but is not limited to, a viral vector, a plasmid (i.e a plasmidic vector), a RNA vector or a linear or circular DNA or RNA molecule which may consists of a chromosomal, non chromosomal, semi-synthetic or synthetic nucleic acids.
- Preferred vectors are those capable of autonomous replication (episomal vector) and/or expression of nucleic acids to which they are linked (expression vectors). Large numbers of suitable vectors are known to those of skill in the art and commercially available.
- Viral vectors include retrovirus, adenovirus, parvovirus (e. g. adenoassociated viruses), coronavirus, negative strand RNA viruses such as orthomyxovirus (e. g., influenza virus), rhabdovirus (e. g., rabies and vesicular stomatitis virus), paramyxovirus (e. g. measles and Sendai), positive strand RNA viruses such as picornavirus and alphavirus, and double-stranded DNA viruses including adenovirus, herpesvirus (e. g., Herpes Simplex virus types 1 and 2, Epstein-Barr virus, cytomegalovirus), and poxvirus (e.
- orthomyxovirus e. g., influenza virus
- rhabdovirus e. g., rabies and vesicular stomatitis virus
- paramyxovirus e. g. measles and Sendai
- viruses include Norwalk virus, togavirus, flavivirus, reoviruses, papovavirus, hepadnavirus, and hepatitis virus, for example.
- retroviruses include: avian leukosis-sarcoma, mammalian C-type, B-type viruses, D type viruses, HTLV-BLV group, lentivirus, spumavirus (Coffin, J. M., Retroviridae: The viruses and their replication, In Fundamental Virology, Third Edition, B. N. Fields, et al., Eds., Lippincott-Raven Publishers, Philadelphia, 1996).
- lentiviral vector HIV-Based lentiviral vectors that are very promising for gene delivery because of their relatively large packaging capacity, reduced immunogenicity and their ability to stably transduce with high efficiency a large range of different cell types.
- Lentiviral vectors are usually generated following transient transfection of three (packaging, envelope and transfer) or more plasmids into producer cells.
- lentiviral vectors enter the target cell through the interaction of viral surface glycoproteins with receptors on the cell surface.
- the viral RNA undergoes reverse transcription, which is mediated by the viral reverse transcriptase complex.
- the product of reverse transcription is a double-stranded linear viral DNA, which is the substrate for viral integration in the DNA of infected cells.
- integrated lentiviral vectors or LV
- integrated lentiviral vectors or LV
- non integrative lentiviral vectors or NILV is meant efficient gene delivery vectors that do not integrate the genome of a target cell through the action of the virus integrase.
- One type of preferred vector is an episome, i.e., a nucleic acid capable of extra-chromosomal replication.
- Preferred vectors are those capable of autonomous replication and/or expression of nucleic acids to which they are linked.
- Vectors capable of directing the expression of genes to which they are operatively linked are referred to herein as "expression vectors.
- a vector according to the present invention comprises, but is not limited to, a YAC (yeast artificial chromosome), a BAC (bacterial artificial), a baculovirus vector, a phage, a phagemid, a cosmid, a viral vector, a plasmid, a NA vector or a linear or circular DNA or RNA molecule which may consist of chromosomal, non chromosomal, semi-synthetic or synthetic DNA.
- expression vectors of utility in recombinant DNA techniques are often in the form of "plasmids" which refer generally to circular double stranded DNA loops which, in their vector form are not bound to the chromosome.
- Vectors can comprise selectable markers, for example: neomycin phosphotransferase, histidinol dehydrogenase, dihydrofolate reductase, hygromycin phosphotransferase, herpes simplex virus thymidine kinase, adenosine deaminase, glutamine synthetase, and hypoxanthine-guanine phosphoribosyl transferase for eukaryotic cell culture; TRP1 for 5. cerevisiae; tetracyclin, rifampicin or ampicillin resistance in E. coli.
- selectable markers for example: neomycin phosphotransferase, histidinol dehydrogenase, dihydrofolate reductase, hygromycin phosphotransferase, herpes simplex virus thymidine kinase, adenosine deaminase, glutamine
- said vectors are expression vectors, wherein a sequence encoding a polypeptide of interest is placed under control of appropriate transcriptional and translational control elements to permit production or synthesis of said polypeptide. Therefore, said polynucleotide is comprised in an expression cassette. More particularly, the vector comprises a replication origin, a promoter operatively linked to said encoding polynucleotide, a ribosome binding site, a RNA-splicing site (when genomic DNA is used), a polyadenylation site and a transcription termination site. It also can comprise an enhancer or silencer elements. Selection of the promoter will depend upon the cell in which the polypeptide is expressed. Suitable promoters include tissue specific and/or inducible promoters.
- inducible promoters are: eukaryotic metallothionine promoter which is induced by increased levels of heavy metals, prokaryotic lacZ promoter which is induced in response to isopropyl-p-D-thiogalacto-pyranoside (IPTG) and eukaryotic heat shock promoter which is induced by increased temperature.
- tissue specific promoters are skeletal muscle creatine kinase, prostate-specific antigen (PSA), ct-antitrypsin protease, human surfactant (SP) A and B proteins, ⁇ - casein and acidic whey protein genes.
- Inducible promoters may be induced by pathogens or stress, more preferably by stress like cold, heat, UV light, or high ionic concentrations (reviewed in Potenza C et al. 2004, In vitro Cell Dev Biol 40:1-22). Inducible promoter may be induced by chemicals (reviewed in (Moore, Samalova et al. 2006); (Padidam 2003); (Wang, Zhou et al. 2003); (Zuo and Chua 2000).
- Delivery vectors and vectors can be associated or combined with any cellular permeabilization techniques such as sonoporation or electroporation or derivatives of these techniques.
- cell or cells any prokaryotic or eukaryotic living cells, cell lines derived from these organisms for in vitro cultures, primary cells from animal or plant origin.
- primary cell or “primary cells” are intended cells taken directly from living tissue (i.e. biopsy material) and established for growth in vitro, that have undergone very few population doublings and are therefore more representative of the main functional components and characteristics of tissues from which they are derived from, in comparison to continuous tumorigenic or artificially immortalized cell lines. These cells thus represent a more valuable model to the in vivo state they refer to.
- fungus refers to a fungal, plant or animal cell or a cell line derived from the organisms listed below and established for in vitro culture. More preferably, the fungus is of the genus Aspergillus, Penicillium, Acremonium, Trichoderma, Chrysoporium, Mortierella, Kluyveromyces or Pichia; More preferably, the fungus is of the species Aspergillus niger, Aspergillus nidulans, Aspergillus oryzae, Aspergillus terreus, Penicillium chrysogenum, Penicillium citrinum, Acremonium Chrysogenum, Trichoderma reesei, Mortierella alpine, Chrysosporium lucknowense, Kluyveromyces lactis, Pichia pastoris or Pichia ciferrii.
- the plant is of the genus Arabidospis, Nicotiana, Solanum, lactuca, Brassica, Oryza, Asparagus, Pisum, Medicago, Zea, Hordeum, Secale, Triticum, Capsicum, Cucumis, Cucurbita, Citrullis, Citrus, Sorghum; More preferably, the plant is of the species Arabidospis thaliana, Nicotiana tabaccum, Solanum lycopersicum, Solanum tuberosum, Solanum melongena, Solanum esculentum, Lactuca saliva, Brassica napus, Brassica oleracea, Brassica rapa, Oryza glaberrima, Oryza sativa, Asparagus officinalis, Pisum sativum, Medicago sativa, zea mays, Hordeum vulgare, Secale cereal, Triticum aestivum, Triticum durum, Capsicum sativus, Cu
- the animal cell is of the genus Homo, Rattus, Mus, Sus, Bos, Danio, Canis, Felis, Equus, Salmo, Oncorhynchus, Gallus, Meleagris, Drosophila, Caenorhabditis; more preferably, the animal cell is of the species Homo sapiens, Rattus norvegicus, Mus musculus, Sus scrofa, Bos
- the cell can be a plant cell, a mammalian cell, a fish cell, an insect cell or cell lines derived from these organisms for in vitro cultures or primary cells taken directly from living tissue and established for in vitro culture.
- cell lines can be selected from the group consisting of CHO-K1 cells; HEK293 cells; Caco2 cells; U2-OS cells; NIH 3T3 cells; NSO cells; SP2 cells; CHO-S cells; DG44 cells; K-562 cells, U-937 cells; M C5 cells; IMR90 cells; Jurkat cells; HepG2 cells; HeLa cells; HT-1080 cells; HCT-116 cells; Hu-h7 cells; Huvec cells; Molt 4 cells.
- All these cell lines can be modified by the method of the present invention to provide cell line models to produce, express, quantify, detect, study a gene or a protein of interest; these models can also be used to screen biologically active molecules of interest in research and production and various fields such as chemical, biofuels, therapeutics and agronomy as non-limiting examples.
- Adoptive immunotherapy using genetically engineered T cells is a promising approach for the treatment of malignancies and infectious diseases. Most current approaches rely on gene transfer by random integration of an appropriate T Cell Receptor (TCR) or Chimeric Antigen Receptor (CAR).
- TCR T Cell Receptor
- CAR Chimeric Antigen Receptor
- Targeted approach using rare-cutting endonucleases such as meganucleases is an efficient and safe alternative method to transfer genes into T cells and generate genetically engineered T cells.
- homologous is intended a sequence with enough identity to another one to lead to homologous recombination between sequences, more particularly having at least 95 % identity, preferably 97 % identity and more preferably 99 %.
- identity refers to sequence identity between two nucleic acid molecules or polypeptides. Identity can be determined by comparing a position in each sequence which may be aligned for purposes of comparison. When a position in the compared sequence is occupied by the same base, then the molecules are identical at that position. A degree of similarity or identity between nucleic acid or amino acid sequences is a function of the number of identical or matching nucleotides at positions shared by the nucleic acid sequences.
- Various alignment algorithms and/or programs may be used to calculate the identity between two sequences, including FASTA, or BLAST which are available as a part of the GCG sequence analysis package (University of Wisconsin, Madison, Wis.), and can be used with, e.g., default setting.
- mutant is intended the substitution, deletion, insertion of one or more nucleotides/amino acids in a polynucleotide (cDNA, gene) or a polypeptide sequence.
- Said mutation can affect the coding sequence of a gene or its regulatory sequence. It may also affect the structure of the genomic sequence or the structure/stability of the encoded mRNA.
- double-strand break-induced mutagenesis (DSB-induced mutagenesis) and “targeted mutagenesis” refer to a mutagenesis event consecutive to an NHEJ event following an endonuclease-induced DSB, leading to insertion/deletion at the cleavage site of an endonuclease
- gene is meant the basic unit of heredity, consisting of a segment of DNA arranged in a linear manner along a chromosome, which codes for a specific protein or segment of protein.
- a gene typically includes a promoter, a 5' untranslated region, one or more coding sequences (exons), optionally introns, a 3' untranslated region.
- the gene may further comprise a terminator, enhancers and/or silencers.
- transgene refers to a sequence encoding a polypeptide.
- the polypeptide encoded by the transgene is either not expressed, or expressed but not biologically active, in the cell, tissue or individual in which the transgene is inserted.
- the transgene encodes a therapeutic polypeptide useful for the treatment of an individual.
- gene of interest refers to any nucleotide sequence encoding a known or putative gene product.
- locus is the specific physical location of a DNA sequence (e.g. of a gene) on a chromosome.
- locus usually refers to the specific physical location of an endonuclease's target sequence on a chromosome.
- locus which comprises a target sequence that is recognized and cleaved by an endonuclease according to the invention, is referred to as “locus according to the invention”.
- locus according to the invention is referred to as “locus according to the invention”.
- the expression "genomic locus of interest” is used to qualify a nucleic acid sequence in a genome that can be a putative target for a double-strand break according to the invention.
- genomic locus of interest is intended a native nucleic acid sequence in a genome, i.e. a sequence or allelic variations of this sequence that is naturally present at this genomic locus. It is understood that the considered genomic locus of interest of the present invention can not only qualify a nucleic acid sequence that exists in the main body of genetic material (i.e. in a chromosome) of a cell but also a portion of genetic material that can exist independently to said main body of genetic material such as plasmids, episomes, virus, transposons or in organelles such as mitochondria or chloroplasts as non-limiting examples.
- loss of genetic information is understood the elimination or addition of at least one given DNA fragment (at least one nucleotide) or sequence, bordering the recognition sites of the endonucleases, rare-cutting endonucleases or chimeric rare-cutting endonucleases of the present invention and leading to a change of the original sequence around said endonuclease-, rare-cutting endonuclease- or chimeric rare-cutting endonuclease-cutting sites, within the genomic locus of interest.
- This loss of genetic information can be, as a non-limiting example, the elimination
- this loss of genetic information can also be an excision of a single-strand of DNA spanning the starting rare-cutting endonuclease scaffold DNA binding region of a chimeric rare-cutting endonuclease according to the present invention when both one cleavase enhancer domain and one nickase enhancer domain, respectively, are part of said chimeric rare-cutting endonuclease.
- the starting rare- cutting endonuclease scaffold DNA binding region can be catalytically active or not and the "loss of genetic information" corresponds to the elimination of an intervening DNA fragment between both cleavase and nickase cleavage sites of said chimeric rare-cutting endonuclease.
- two nearby DNA double strand breaks within the genomic locus of interest, is meant two endonucleases cutting sites distant at between 12 bp and 1000 bp.
- fusion protein is intended the result of a well-known process in the art consisting in the joining or merging of two or more genes which originally encode for separate proteins or part of them, the translation of said "fusion gene” resulting in a single polypeptide, said "fusion protein” with functional properties derived from each of the original proteins.
- Imprecise NHEJ is intended the re-ligation of nucleic acid ends generated by a DSB, with insertions or deletions of nucleotides. Imprecise NHEJ is an outcome and not a repair pathway and can result from different NHEJ pathways (Ku dependent or Ku independent as non-limiting examples).
- chimeric rare-cutting endonuclease any fusion protein comprising a rare- cutting endonuclease. Said rare-cutting endonuclease might be at the N- terminus part of said chimeric rare-cutting endonuclease; at the opposite, said rare-cutting endonuclease might be at the C- terminus part of said chimeric rare-cutting endonuclease.
- a “chimeric rare-cutting endonuclease” according to the present invention which comprises two catalytic domains can be described as "bi- functional" or as "bi-functional meganuclease".
- chimeric rare-cutting endonuclease which comprises more than two catalytic domains can be described as “multi- functional” or as “multi-functional meganuclease".
- chimeric rare-cutting endonucleases according to the present invention can be a fusion protein between a rare-cutting endonuclease and one catalytic domain; chimeric rare-cutting endonucleases according to the present invention can also be a fusion protein between a rare-cutting endonuclease and two catalytic domains.
- the rare-cutting endonuclease part of chimeric rare-
- 59 cutting endonucleases according to the present invention can be a meganuclease comprising two identical monomers, two non-identical monomers, or a single chain meganuclease.
- the rare-cutting endonuclease part of chimeric rare-cutting endonucleases according to the present invention can also be the DNA-binding domain of an inactive rare-cutting endonuclease.
- chimeric rare-cutting endonucleases according to the present invention can be derived from a TALE-nuclease (TALEN), i. e.
- a fusion between a DNA-binding domain derived from a Transcription Activator Like Effector (TALE) and one or two catalytic domains are encompassed mutants, variants or derivatives of meganucleases (including monomers and single chain meganucleases), endonucleases, rare-cutting endonucleases or chimeric rare-cutting endonucleases.
- TALE Transcription Activator Like Effector
- hancer domain(s) or “enhancer(s)” are meant protein domains that provide functional and/or structural support to a protein scaffold, a rare-cutting endonuclease-derived scaffold as a non-limiting example, therefore allowing an enhancement in cleavage efficiency of the resulting fusion protein, i.e a chimeric rare-cutting endonuclease, relative to the cleavage efficiency of the wild-type rare-cutting endonuclease or starting rare-endonuclease scaffold.
- a particular domain is an enhancer domain when it provides at least a 5% enhancement in efficiency of the starting scaffold, more preferably 10 %, again more preferably 15 %, again more preferably 20 %, again more preferably 25 %, again more preferably 50%, again more preferably greater than 50%.
- enhancer domains are given in Table 1.
- Enhancer domain(s) according to the present invention can be fused to N-terminus and/or C-terminus of a rare-cutting endonuclease-derived monomer scaffold of a homodimeric endonuclease, resulting in a chimeric rare-cutting endonuclease comprising two or four enhancer domains.
- Enhancer domain(s) according to the present invention can also be fused to N-terminus and/or C-terminus of a rare-cutting endonuclease-derived monomer scaffold of a heterodimeric endonuclease, resulting in a chimeric rare-cutting endonuclease comprising one or two or three or four enhancer domains.
- Enhancer domain(s) according to the present invention can also be fused to N-terminus and/or C-terminus of a rare-cutting endonuclease-derived scaffold of a single-chain endonuclease, resulting in a chimeric rare-cutting endonuclease comprising one or two enhancer domains.
- catalytic domain is intended the protein domain or module of an enzyme containing the active site of said enzyme; by active site is intended the part of said enzyme at which catalysis of the substrate occurs.
- Enzymes, but also their catalytic domains, are classified and named according to the reaction they catalyze.
- the Enzyme Commission number (EC number) is a numerical classification scheme for enzymes, based on the chemical reactions they catalyze
- any catalytic domain can be fused to a rare-cutting endonuclease scaffold or starting rare-endonuclease scaffold or starting scaffold to generate a chimeric rare-cutting endonuclease.
- said catalytic domain can be an enhancer domain according to the present invention. If catalytically active, said enhancer domain can provide functional and/or structural support to rare-cutting endonuclease scaffold when fused to it. If catalytically inactive, said enhancer domain provides structural support to rare-cutting endonuclease scaffold when fused to it.
- nuclease catalytic domain is intended the protein domain comprising the active site of an endonuclease or an exonuclease enzyme.
- Such nuclease catalytic domain can be, for instance, a “cleavase domain” or a “nickase domain”.
- cleavase domain is intended a protein domain whose catalytic activity generates a Double Strand Break (DSB) in a DNA target.
- nickase domain is intended a protein domain whose catalytic activity generates a single strand break in a DNA target sequence.
- Non-limiting examples of such catalytic domains are given in Table 1 with a GenBank or NCBI or UniProtKB/Swiss-Prot number as a reference.
- Protein-fusion scaffolds were designed based on a truncated form of wild-type l-Crel (SEQ ID NO: 1), l-Crel_X (SEQ ID NO: 5) and three different linker polypeptides [NFSl (SEQ ID NO: 98), NFS2 (SEQ ID NO: 99) and CFS1 (SEQ ID NO: 100)] fused to either the N- or C-terminus of the protein. Structure models were generated in all cases, with the goal of designing a "baseline" fusion linker that would traverse the l-Crel parent scaffold surface with little to no effect on its DNA binding or cleavage activities.
- the polypeptide spanning residues 2 to 153 of I- Crel was used, with a K82A mutation to allow for linker placement.
- the C-terminal fusion scaffold contains residues 2 to 155 of wild-type l-Crel.
- the "free" end of the linker i.e. onto which a polypeptide can be linked
- l-Crel_CFSl SEQ ID NO: 8
- yeast assay previously described in International PCT Applications WO 2004/067736 and in (Epinat, Arnould et al. 2003; Chames, Epinat et al. 2005; Arnould, Chames et al. 2006; Smith, Grizot et al. 2006) and found to have activity similar to that of wild-type l-Crel.
- Wild-type l-Hmul functions as a monomeric nickase to generate a strand-specific nick in its target DNA [REF].
- variable length constructs were designed from the N-terminal region of l-Hmul that encompass the entire catalytic domain. Fragments were fused to the N-terminus of either l-Crel_NFSl or l-Crel_NFS2 to generate 8 constructs (SEQ ID NO: 9 and SEQ ID NO: 105-111).
- each "enhanced" meganuclease was assessed using yeast assay described in International PCT Applications WO 2004/067736.
- yeast assay described in International PCT Applications WO 2004/067736.
- all constructs were screened against libraries of the C1221 target DNA ( Figure 2) having regions mutated in the following positions (numbering as in Figure 2): 12NN, lONNNnn, 7NNNnn, 5NNN, 2NN.
- These targets contain nucleotide substitutions at the +/- positions indicated.
- fusion proteins strongly enhanced cleavage of targets relative to the
- Table 3 Activity in Yeast assay for l-Hmul/l-Crel fusions. The activity of the hHmuCre fusion protein relative to that of wild-type l-Crel is shown for various target libraries (see Example 3 text for details). Activity enhancement is based on a direct comparison of the change in activity for targets cleavable by the hHmuCre hybrid. The total number of targets in a given library is indicated next to the library name. The percent of targets enhanced to a given threshold are listed in each column. Assays were performed at two temperatures, 30°C and 37°C. Numbers in parentheses indicate actual target number enhanced versus total targets cleaved at the given temperature.
- Table 4 Additional 2NN targets cleaved by hHmuCre at 37°C. From the 136 possible 2NN targets, hHmuCre is able to cleave a total of 125 (see Table 3) at 37°C. Of these, 13 DNA sequences are not cleaved by wild-type l-Crel at this temperature. Listed are the 2NN sequences and relative activities
- Table 5 Additional 2NN targets cleaved by hHmuCre at 30"C. From the 136 possible 2NN targets, hHmuCre is able to cleave a total of 110 (see Table 3) at 30°C. Of these, 47 DNA sequences are not cleaved by wild-type l-Crel at this temperature. Listed are the 2NN sequences and relative activities for hHmuCre vs wild-type l-Crel.DNA sequence is based on the C1221target with the central four base pairs replaced as indicated. Relative activity is scaled as: -, no activity detectable; +, ⁇ 25% activity; ++, 25% to ⁇ 50% activity; +++, 50% to ⁇ 75% activity; ++++, 75% to 100% activity.
- constructs were generated by adding variable length regions of the CFP1 peptide (SEQ ID NO: 112) to the C-terminus of the protein.
- l-Crel is a homodimer
- each "enhanced" meganuclease was assessed using yeast assay described in International PCT Applications WO 2004/067736 (see Example 1).
- yeast assay described in International PCT Applications WO 2004/067736 (see Example 1).
- all constructs were screened against libraries of the C1221 target DNA ( Figure 2) having regions mutated in the following positions (numbering as in Figure 2): 12NN, lONNNnn, 7NNNnn, 5NNN, 2NN.
- These targets contain nucleotide substitutions at the +/- positions indicated. Fusion proteins enhance cleavage of targets to various degrees relative to the activity of wild-type l-Crel.
- Hybrid restriction enzymes zinc finger fusions to Fok I cleavage
- TAL nucleases hybrid proteins composed of TAL effectors
- TAL nucleases hybrid proteins composed of TAL effectors
- nbtl410 [pii] 10.1038/nbtl410.” Nature Biotechnology 26(7): 808-16.
- thermostable archaeal intron- encoded endonuclease l-Dmol J Mol Biol 286(4): 1123-36.
Landscapes
- Life Sciences & Earth Sciences (AREA)
- Health & Medical Sciences (AREA)
- Chemical & Material Sciences (AREA)
- Wood Science & Technology (AREA)
- Molecular Biology (AREA)
- Engineering & Computer Science (AREA)
- Bioinformatics & Cheminformatics (AREA)
- Organic Chemistry (AREA)
- Zoology (AREA)
- Genetics & Genomics (AREA)
- Medicinal Chemistry (AREA)
- Microbiology (AREA)
- Biotechnology (AREA)
- Biochemistry (AREA)
- General Engineering & Computer Science (AREA)
- General Health & Medical Sciences (AREA)
- Biomedical Technology (AREA)
- Enzymes And Modification Thereof (AREA)
- Micro-Organisms Or Cultivation Processes Thereof (AREA)
Abstract
The present invention relates to a method for enhancing the efficiency of rare-cutting endonucieases such as meganucleases. More specifically, the present invention concerns a method for the creation of fusion proteins that consist of one or more "enhancer" domains fused to at least one rare-cutting endonuclease-derived scaffold in a single polypeptide chain for simple and efficient vectorization. The present invention also relates to engineered chimeric rare-cutting endonucieases, vectors, compositions and kits used to implement the method and use of said chimeric rare-cutting endonucieases according to the invention for various applications ranging from homologous gene targeting to targeted mutagenesis and sequence removal.
Description
METHOD FOR ENHANCING RARE-CUTTING ENDONUCLEASE EFFICIENCY AND USES THEREOF
The present application claims priority to U.S. Serial number 61/472,053 filed April 5, 2011, the entire contents of which are incorporated herein by reference.
Field of the invention
The present invention relates to a method for enhancing the efficiency of rare-cutting endonucleases such as meganucleases. More specifically, the present invention concerns a method for the creation of fusion proteins that consist of one or more "enhancer" domains fused to at least one rare-cutting endonuclease-derived scaffold in a single polypeptide chain for simple and efficient vectorization. The present invention also relates to engineered chimeric rare-cutting endonucleases, vectors, compositions and kits used to implement the method and use of said chimeric rare-cutting endonucleases according to the invention for various applications ranging from homologous gene targeting to targeted mutagenesis and sequence removal.
Background of the invention
Mammalian genomes constantly suffer from various types of damage, of which double-strand breaks (DSBs) are considered the most dangerous (Haber 2000). Repair of DSBs can occur through diverse mechanisms that can depend on cellular context. Repair via homologous recombination (HR) is able to restore the original sequence at the break. Because of its strict dependence on extensive sequence homology, this mechanism is suggested to be active mainly during the S and G2 phases of the cell cycle where the sister chromatids are in close proximity (Sonoda, Hochegger et al. 2006). Single-strand annealing (SSA) is another homology-dependent process that can repair DSBs between direct repeats and thereby promotes deletions (Paques and Haber 1999). Finally, non-homologous end joining (NHEJ) of DIMA is a major pathway for the repair of DSBs that can function throughout the cell cycle and does not depend on homologous recombination (Moore and Haber 1996; Haber 2008). NHEJ seems to comprise at least two different components: (i) a pathway that consists mostly in the direct re-joining of DSB ends, and which depends on the XRCC4, Lig4 and Ku proteins, and; (ii) an alternative NHEJ pathway, which does not depend on XRCC4, Lig4 and Ku, and is especially error- prone, resulting mostly in deletions, with the junctions occurring between micro-homologies (Frank, Sekiguchi et al. 1998; Gao, Sun et al. 1998; Guirouilh-Barbat, Huck et al. 2004; Guirouilh-Barbat, Rass et al. 2007; Haber 2008; McVey and Lee 2008).
Homologous gene targeting (HGT), first described over 25 years ago (Hinnen, Hicks et al. 1978; Orr- Weaver, Szostak et al. 1981; Orr-Weaver, Szostak et al. 1983; Rothstein 1983), was one of the first methods for rational genome engineering and remains to this day a standard for the generation of engineered cells or knock-out mice (Capecchi 2001). An inherently low efficiency has nevertheless prevented it from being used as a routine protocol in most cell types and organisms. To address these issues, an extensive assortment of rational approaches has been proposed with the intent of achieving greater than 1% targeted modifications. Many groups have focused on enhancing the efficacy of HGT, with two major disciplines having become apparent: (i) so-called "matrix optimization" methods, essentially consisting of modifying the targeting vector structure to achieve maximal efficacy, and; (ii) methods involving additional effectors to stimulate HR, generally sequence-specific endonucleases. The field of matrix optimization has covered a wide range of techniques, with varying degrees of success (Russell and Hirata 1998; Inoue, Dong et al. 2001; Hirata, Chamberlain et al. 2002; Taubes 2002; Gruenert, Bruscia et al. 2003; Sangiuolo, Scaldaferri et al. 2008; Bedayat, Abdolmohamadi et al. 2010). Stimulation of HR via nucleases, on the other hand, has repeatedly proven efficient (Paques and Duchateau 2007; Carroll 2008).
For DSBs induced by biological reagents, e.g. meganucleases, ZFNs and TALENs (see below), which cleave DNA by hydrolysis of two phosphodiester bonds, the DNA can be rejoined in a seamless manner by simple re-ligationof the cohesive ends. Alternatively, deleterious insertions or deletions (indels) of various sizes can occur at the breaks, eventually resulting in gene inactivation (Liang, Han et al. 1998; Lloyd, Plaisier et al. 2005; Doyon, McCammon et al. 2008; Perez, Wang et al. 2008; Santiago, Chan et al. 2008; Kim, Lee et al. 2009; Yang, Djukanovic et al. 2009). The nature of this process, which does not rely on site-specific or homologous recombination, gives rise to a third targeted approach based on endonuclease-induced mutagenesis. This approach, as well as the related applications, may be simpler than those based on homologous recombination in that (a) one does not need to introduce a repair matrix, and; (b) efficacy will be less cell-type dependant (in contrast to HR, NHEJ is probably active throughout the cell cycle (Delacote and Lopez 2008). Targeted mutagenesis based on NEHJ has been used to trigger inactivation of single or even multiple genes in immortalized cell lines (Cost, Freyvert et al. 2010; Liu, Chan et al. 2010). In addition, this method opens new perspectives for organisms in which the classical HR-based gene knock-out methods have proven inefficient, or at least difficult to establish (Doyon, McCammon et al. 2008; Geurts, Cost et al. 2009; Shukla, Doyon et al. 2009; Yang, Djukanovic et al. 2009; Gao, Smith et al. 2010; Mashimo, Takizawa et al. 2010; Menoret, Iscache et al. 2010).
Over the last 15 years, the use of meganucleases to successfully induce gene targeting has been well documented, starting from straightforward experiments involving wild-type l-Scel to more refined work involving completely re-engineered enzymes (Stoddard, Scharenberg et al. 2007; Galetto, Duchateau et al. 2009; Marcaida, Munoz et al. 2010; Arnould, Delenda et al. 2011). Meganucleases, also called homing endonucleases (HEs), can be divided into five families based on sequence and structure motifs: LAGLIDADG, GIY-YIG, HNH, His-Cys box and PD-(D/E)XK (Stoddard 2005; Zhao, Bonocora et al. 2007). Structural data are available for at least one member of each family. The most well studied family is that of the LAGLIDADG proteins, with a considerable body of biochemical, genetic and structural work having established that these endonucleases could be used as molecular tools (Stoddard, Scharenberg et al. 2007; Arnould, Delenda et al. 2011). Member proteins are composed of domains that adopt a similar αββαββα fold, with the LAGLIDADG motif comprising the terminal region of the first helix and not only contributing to a bipartite catalytic center but also forming the core subunit/subunit interaction (Stoddard 2005). Two such α/β domains assemble to form the functional protein, with the β-strands in each creating a saddle- shaped DNA binding region. The spatial separation of the catalytic center with regions directly interacting with the DNA has allowed for specificity re-engineering (Seligman, Chisholm et al. 2002; Sussman, Chadsey et al. 2004; Arnould, Chames et al. 2006; Doyon, Pattanayak et al. 2006; Rosen, Morrison et al. 2006; Smith, Grizot et al. 2006; Arnould, Perez et al. 2007). In addition, whereas all known LAGLIDADG proteins analyzed to date act as "cleavases" to cut both strands of the target DNA, recent progress has been made in generating "mega-nickases" that cleave only one strand (Niu, Tenney et al. 2008; McConnell Smith, Takeuchi et al. 2009). Such enzymes can in principle provide similar levels of targeted induced HR with a minimization in the frequency of NHEJ. Although numerous engineering efforts have focused on LAGLIDADG HEs, members from two other families, GIY-YIG and HNH, are of particular interest. Biochemical and structural studies have established that in both families, member proteins can adopt a bipartite fold with distinct functional domains: (1) a catalytic domain responsible mainly for DNA cleavage, and; (2) a DNA-binding domain to provide target specificity (Stoddard 2005; Marcaida, Munoz et al. 2010). The related GIY-YIG HEs I- TevI and l-Bmol have been exploited to demonstrate the interchangeability of the DNA-binding region for these enzymes (Liu, Derbyshire et al. 2006). Analysis of the l-Basl HE revealed that although the N-terminal catalytic domain belongs to the HNH family, the C-terminal DNA-binding region resembles the intron-encoded endonuclease repeat motif (IENR1) found in endonucleases of the GIY-YIG family (Landthaler and Snub 2003). The catalytic head of l-Basl has sequence similarity to
those of the HNH HEs l-Hmul, l-Hmull and l-Twol, all of which function as strand-specific nickases (Landthaler, Begley et al. 2002; Landthaler and Shub 2003; Landthaler, Lau et al. 2004; Shen, Landthaler et al. 2004; Landthaler, Shen et al. 2006). Whereas the above families of proteins contain sequence-specific nucleases, the HNH motif has also been identified in nonspecific nucleases such the E.coli colicins (e.g. ColE9 and ColE7), EndA from S. pneumoniae, NucA from Anabaena and CAD (Midon, Schafer et al. 2011). As well as having the HNH motif, several of these nucleases contain the signature DRGH motif and share structural homology with core elements forming the ββα-Me-finger active site motif. Mutational studies of residues in the HNH/DRGH motifs have confirmed their role in nucleic acid cleavage activity (Ku, Liu et al. 2002; Doudeva, Huang et al. 2006; Eastberg, Eklund et al. 2007; Huang and Yuan 2007). Furthermore, the DNA binding affinity and sequence preference for ColE7 could be effectively altered (Wang, Wright et al. 2009). Such detailed studies illustrate the potential in re-engineering nonspecific nucleases for targeted purposes.
Zinc-finger nucleases (ZFNs), generated by fusing Zinc-finger-based DNA-binding domains to an independent catalytic domain via a flexible linker (Kim, Cha et al. 1996; Smith, Berg et al. 1999; Smith, Bibikova et al. 2000), represent another type of engineered nuclease commonly used to stimulate gene targeting. The archetypal ZFNs are based on the catalytic domain of the Type IIS restriction enzyme Fokl and have been successfully used to induce gene correction, gene insertion, and gene deletion. Zinc Finger-based DNA binding domains are made of strings of 3 or 4 individual Zinc Fingers, each recognizing a DNA triplet (Pabo, Peisach et al. 2001). In theory, one of the major advantages of ZFNs is that they are easy to design, using combinatorial assembly of preexisting Zinc Fingers with known recognition patterns (Choo and Klug 1994; Choo and Klug 1994; Kim, Lee et al. 2009). However, close examination of high resolution structures shows that there are actually crosstalks between units (EIrod-Erickson, Rould et al. 1996), and several methods have been used to assemble ZF proteins by choosing individual Zinc Fingers in a context dependant manner (Greisman and Pabo 1997; Isalan and Choo 2001; Maeder, Thibodeau-Beganny et al. 2008; Ramirez, Foley et al. 2008) to achieve better success rates and reagents of better quality.
Recently, a new class of chimeric nuclease using a Fokl catalytic domain has been described (Christian, Cermak et al. 2010; Li, Huang et al. 2011). The DNA binding domain of these nucleases is derived from Transcription Activator Like Effectors (TALE), a family of proteins used in the infection process by plant pathogens of the Xanthomonas genus. In these DNA binding domains, sequence
specificity is driven by a series of 33-35 amino acids repeats, differing essentially by two positions (Boch, Scholze et al. 2009; Moscou and Bogdanove 2009). Each base pair in the DNA target is contacted by a single repeat, with the specificity resulting from the two variant amino acids of the repeat (the so-called repeat variable dipeptide, RVD). The apparent modularity of these DNA binding domains has been confirmed to a certain extent by modular assembly of designed TALE-derived protein with new specificities (Boch, Scholze et al. 2009; Moscou and Bogdanove 2009). However, one cannot yet rule out a certain level of context dependence of individual repeat/base recognition patterns, as was observed for Zinc Finger proteins (see above). Furthermore, it has been shown that natural TAL effectors can dimerize (Gurlebeck, Szurek et al. 2005) and how this would affect a "dimerization-based" TALE-derived nuclease is currently unknown.
The functional layout of a Fokl-based TALE-nuclease (TALEN) is essentially that of a ZFN, with the Zinc-finger DNA binding domain being replaced by the TALE domain (Christian, Cermak et al. 2010; Li, Huang et al. 2011). As such, DNA cleavage by a TALEN requires two DNA recognition regions flanking an unspecific central region. This central "spacer" DNA region is essential to promote catalysis by the dimerizing Fokl catalytic domain, and extensive effort has been placed into optimizing the distance between the DNA binding sites (Christian, Cermak et al. 2010; Miller, Tan et al. 2011). The length of the spacer has been varied from 14 to 30 base pairs, with efficacy in DNA cleavage being interdependent with spacer length as well as TALE scaffold construction (i.e. the nature of the fusion construct used). It is still unknown whether differences in the repeat region (i.e. RVD type and number used) have an impact on the DNA "spacer" requirements or on the efficacy of DNA cleavage by TALENs. Nevertheless, TALE-nucleases have been shown to be active to various extents in cell-based assays in yeast, mammalian cells and plants (Christian, Cermak et al. 2010; Li, Huang et al. 2011; Mahfouz, Li et al. 2011; Miller, Tan et al. 2011).
In genome engineering experiments, the efficiency of rare-cutting endonuclease, e.g. their ability to induce a desired event (Homologous gene targeting, targeted mutagenesis, sequence removal or excision) at a locus, depends on several parameters, including the specific activity of the nuclease, probably the accessibility of the target, and the efficacy and outcome of the repair pathway(s) resulting in the desired event (homologous repair for gene targeting, NHEJ pathways for targeted mutagenesis).
Cleavage by peptidic rare cutting endonucieases usually generates cohesive ends, with 3' overhangs for LAGLIDADG meganucleases(Chevalier and Stoddard 2001) and 5' overhangs for Zinc Finger
U 2012/032386
Nucleases (Smith, Bibikova et al. 2000). These ends, which result from hydrolysis of phosphodiester bonds, can be re-ligated in vivo by NHEJ in a seamless way (i.e a scarless re-ligation). The restoration of a cleavable target sequence allows for a new cleavage event by the same endonuclease, and thus, a series of futile cycles of cleavage and re-ligation events can take place. Indirect evidences have shown that even in the yeast Saccharomyces cerevisiae, such cycles could take place upon continuous cleavage by the HO endonuclease (Lee, Paques et al. 1999). In mammalian cells, several experiment have shown that perfect re-ligation of compatible cohesive ends resulting from two independent but close l-Scel-induced DSBs is an efficient process (Guirouilh-Barbat, Huck et al. 2004; Guirouilh-Barbat, ass et al. 2007; Bennardo, Cheng et al. 2008; Bennardo, Gunn et al. 2009). Absence of the Ku DNA repair protein does not significantly affect the overall frequency of NHEJ events rejoining the ends from the two DSBs; however it very strongly enhances the contribution of imprecise NHEJ to the repair process in CHO immortalized cells and mouse ES cells (Guirouilh-Barbat, Huck et al. 2004; Guirouilh-Barbat, Rass et al. 2007; Bennardo, Cheng et al. 2008). Furthermore, the absence of Ku stimulates l-Scel-induced events such as imprecise NHEJ (Bennardo, Cheng et al. 2008), single-strand annealing (Bennardo, Cheng et al. 2008) and gene conversion (Pierce, Hu et al. 2001; Bennardo, Cheng et al. 2008) in mouse ES cells . Similar observations shave been made with cells deficient for the XRCC4 repair protein (Pierce, Hu et al. 2001; Guirouilh-Barbat, Rass et al. 2007; Bennardo, Gunn et al. 2009) (although XRCC4 deficiency affects the overal level of NHEJ in CHO cells (Guirouilh-Barbat, Rass et al. 2007)) or for DNA-PK (Pierce, Hu et al. 2001). In contrast, knock-down of CtIP has been shown to suppresses "alt-NHEJ" (a Ku- and XRCC4-independent form of NHEJ more prone to result in imprecise NHEJ), single-strand annealing and gene conversion, while not affecting the overall level of rejoining of two compatible ends generated by l-Scel (Bennardo, Cheng et al. 2008). Thus, competition between different DSB repair pathways can affect the spectrum or repair events resulting from a nuclease-induced DSB.
In addition, DSB resection is important for certain DSB pathways. Extensive DSB resection, resulting in the generation of large single stranded regions (a few hundred nucleotides at least), has been shown in yeast to initiate single strand annealing (Sugawara and Haber 1992) and strand invasion, the ATP-dependant step that initiates many homologous recombination events of DNA duplex invasion by an homologous strand that (White and Haber 1990; Sun, Treco et al. 1991) (for a review of mechanisms, see (Paques and Haber 1999)). In eukaryotic cells DSB resection depends on several proteins including BLM/Sgsl and DNA2, EXOI, and the MRN complex (Mrell, Rad50, Nbsl/Xrs2) and is thought to result from different pathways. MRN is involved in a small scale resection process, while two redundant pathways depending on BLM and DNA2 on one hand, and on EXOI on another
hand, would be involved in extensive resection (Mimitou and Symington 2008; Nimonkar, Genschel et al. 2011). In addition, processing ends involving a damaged nucleotide (resulting from chemical cleavage or from a bulk adduct), requires the CtlP/Sae2 protein together with MN (Sartori, Lukas et al. 2007; Buis, Wu et al. 2008; Hartsuiker, Mizuno et al. 2009). Over-expression of the Trex2 exonuclease was shown to strongly stimulate imperfect NHEJ associated with loss of only a few base pairs (Bennardo, Gunn et al. 2009), while it inhibited various kinds of DNA repair events between distant sequences (such as Single-strand annealing, NHEJ between ends from different breaks, or NHEJ repair of a single DSB involving remote micro-homologies). In the same study, it was suggested that Trex2 did resect the 3' overhangs let by l-Scel in a non processive way. Thus, the type of stimulated pathway could in turn depend on the type of resection (length of resection, single strand vs. double strand, resection of 5' strand vs. 3' strand).
The authors of the present invention have developed a method to significantly enhance the efficiency of rare-cutting endonucleases such as LAGLIDADG-type meganucleases. The use of novel "enhancer" domains allows for boosting the overall efficiency of the meganuclease without modifying its specificity determinants (i.e. residues making base-specific contacts to the DNA). Furthermore, the invention allows for generating several distinct types of enzymes that can be applied to applications ranging from homologous gene targeting to targeted mutagenesis and sequence removal.
T/US2012/032386
Brief summary of the invention
In a general aspect, the present invention relates to a method for enhancing the efficiency of a rare- cutting endonuclease for a DNA target sequence by merging a rare-cutting endonuclease-derived scaffold with at least one enhancer domain to obtain a chimeric rare-cutting endonuclease with an enhanced efficiency for said DNA target sequence. In another aspect, the present invention concerns a method for the creation of fusion proteins that consist of engineering a fusion protein between a rare-cutting endonuclease-derived scaffold and at least one enhancer domain wherein said enhancer domain enhances the efficiency of said rare-cutting endonuclease when fused to it. The present invention also concerns the creation of functional single polypeptide fusion proteins for simple and efficient vectorization. In another aspect, the present invention relates to chimeric rare- cutting endonucleases comprising at least an enhancer domain wherein said enhancer domain enhances the efficiency of said rare-cutting endonuclease when fused to it, thereby obtaining a chimeric rare-cutting endonuclease with enhanced efficiency for a DNA target sequence compared to a corresponding rare-cutting endonuclease lacking said enhancer domain. The present invention also relates to engineered rare-cutting endonucleases, vectors, compositions and kits used to implement the method and use of said chimeric rare-cutting endonucleases according to the invention for various applications ranging from homologous gene targeting to targeted mutagenesis and sequence removal.
Brief description of the figures
In addition to the preceding features, the invention further comprises other features which will emerge from the description which follows, as well as to the appended drawings. A more complete appreciation of the invention and many of the attendant advantages thereof will be readily obtained as the same becomes better understood by reference to the following Figures in conjunction with the detailed description below. Figure 1: Endonuclease-induced gene targeting approaches. Upon cleavage, DNA repair mechanisms may result in one of several outcomes. (A) When a double-strand break is targeted between two direct repeats, HR can result in the deletion of one repeat together with the intervening sequence. Gene insertion (B) or correction (C) can be achieved by the introduction of a DNA repair matrix containing sequences homologous to the endogenous sequence surrounding the DNA break. Mutations can be corrected either at or distal to the break, with the frequency of correction decreasing with increasing distance. (D) The misrepair of DNA ends by error-prone NHEJ can result in insertions or deletions of various sizes, leading to gene inactivation.
Figure 2: Sequences of target DNA recognized by l-Crel. C1234 (SEQ ID NO: 3) represents the partially symmetric DNA sequence recognized and cleaved by wild-type l-Crel. C1221 (SEQ ID NO: 2) represents a palindromic DNA sequence, derived from C1234 (SEQ ID NO: 3), recognized and cleaved by the l-Crel meganuclease. Nucleotides are numbered outward (-/+) from the center of the target. The nature of the nucleotides at positions -2 to +2 can potentially interfere with the cleavage activity of the protein.
Figure 3: Schematic of enhanced rare-cutting endonucleases, such as meganucleases, fusion configurations. Rare-cutting endonucleases, such as meganucleases, can be enhanced through the addition of a domain to promote existing or alternate activities. As each end of the meganuclease scaffold is amenable to fusion, the order (N- v.s C-terminal) of addition and number of the enhancer domains can vary with the application. Enhanced fusion construct are optimized to address or overcome distinct problems. (A) The addition of two enhancer domains to an active meganuclease can enhance DNA binding and/or cleavage activity. Such a configuration can be achieved via (i) a single N- or C-terminal fusion to a homodimeric variant; (ii) a single N- or C-terminal fusion to
individual monomers of a heterodimer, or; (iii) a double fusion to a monomeric protein. (B) When specificity reengineering precludes maintaining cleavage activity of the meganuclease, the attached enhancer domains can provide alternative functions. (C) and (D) represent instances of (A) and (B), respectively, when only one enhancer domain is needed or tolerated per fusion protein (e.g. either as an N- or C-terminal fusion or in the context of a single-chain molecule). Fusion junctions (N- vs. C- terminal) and linker designs can vary with the application. Components of the fusion proteins are listed in the legend.
Figure 4 : Schematic of DNA cleavage, in vivo re-ligation and other repair pathways. In cells, cleavage by peptidic rare-cutting endonucleases usually result in a DNA double strand break (DSB) with cohesive ends. For example, meganucleases from the LAGLIDADG family, such as l-Scel and I- Crel, produce DSBs with 3' overhang. These cohesive ends can be re-ligated in vivo by NHEJ, resulting in seamless repair, and in the restoration of a cleavable target sequence, which can in turn be processed again by the same endonuclease. Thus, a series of futile cycles of cleavage and re-ligation events can take place. Imprecise NHEJ or homologous recombination can alter or remove the cleavage site, resulting in cycle exit (A). Two other ways can also stop the process : (i) Chromosome loss can occur as the consequence of failure to repair the DSB; (ii) a loss of nuclease (degradation, dilution, cell division, etc.). B-E: Consequences of cleavage of additional phosphodiester bonds. The addition of a single nickase activity (B) or of two nickase activities affecting the same strand (C) would result in a single strand gap, and suppress the cohesive ends, which could in turn affect the spectrum of events. Addition of two nickase activities affecting opposite strands (D) or of a new cleavase activity generating a second DSB (E) would result in a double strand gap. As a consequence, perfect re-ligation is not possible anymore, and one or several alternative repair outcomes could be stimulated. The current figure makes no assumption regarding the relative frequencies of these alternative outcomes (imprecise NHEJ, homologous recombination, others...). Solid triangles represent hydrolysis of phosphodiester bonds.
Detailed description of the invention
Unless specifically defined herein below, all technical and scientific terms used herein have the same meaning as commonly understood by a skilled artisan in the fields of gene therapy, biochemistry, genetics, and molecular biology.
2386
All methods and materials similar or equivalent to those described herein can be used in the practice or testing of the present invention, with suitable methods and materials being described herein. All publications, patent applications, patents, and other references mentioned herein are incorporated by reference in their entirety. In case of conflict, the present specification, including definitions, will prevail. Further, the materials, methods, and examples are illustrative only and are not intended to be limiting, unless otherwise specified.
The practice of the present invention will employ, unless otherwise indicated, conventional techniques of cell biology, cell culture, molecular biology, transgenic biology, microbiology, recombinant DNA, and immunology, which are within the skill of the art. Such techniques are explained fully in the literature. See, for example, Current Protocols in Molecular Biology (Frederick M. AUSUBEL, 2000, Wiley and son Inc, Library of Congress, USA); Molecular Cloning: A Laboratory Manual, Third Edition, (Sambrook et al, 2001, Cold Spring Harbor, New York: Cold Spring Harbor Laboratory Press); Oligonucleotide Synthesis (M. J. Gait ed., 1984); Mullis et al. U.S. Pat. No. 4,683,195; Nucleic Acid Hybridization (B. D. Harries & S. J. Higgins eds. 1984); Transcription And Translation (B. D. Hames & S. J. Higgins eds. 1984); Culture Of Animal Cells (R. I. Freshney, Alan R. Liss, Inc., 1987); Immobilized Cells And Enzymes (IRL Press, 1986); B. Perbal, A Practical Guide To Molecular Cloning (1984); the series, Methods In ENZYMOLOGY (J. Abelson and M. Simon, eds. -in- chief, Academic Press, Inc., New York), specifically, Vols.154 and 155 (Wu et al. eds.) and Vol. 185, "Gene Expression Technology" (D. Goeddel, ed.); Gene Transfer Vectors For Mammalian Cells (J. H. Miller and M. P. Calos eds., 1987, Cold Spring Harbor Laboratory); Immunochemical Methods In Cell And Molecular Biology (Mayer and Walker, eds., Academic Press, London, 1987); Handbook Of Experimental Immunology, Volumes l-IV (D. M. Weir and C. C. Blackwell, eds., 1986); and Manipulating the Mouse Embryo, (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y., 1986).
In a general aspect, the present invention relates to a method for enhancing the efficiency of a rare- cutting endonuclease for a DNA target sequence by merging a rare-cutting endonuclease-derived scaffold with at least one enhancer domain to obtain a chimeric rare-cutting endonuclease with an enhanced efficiency for said DNA target sequence.
According to a first aspect of the present invention is a method for enhancing rare-cutting endonuclease efficiency at a genomic locus of interest in a cell comprising the steps of:
(i) Identifying at said genomic locus of interest one DNA target sequence cleavable by one rare-cutting endonuclease;
(ii) Engineering said rare-cutting endonuclease in order to create a chimeric rare-cutting endonuclease wherein said chimeric rare-cutting endonuclease is a fusion protein comprising at least a rare-cutting endonuclease-derived scaffold and at least one enhancer domain and wherein said chimeric rare-cutting endonuclease has an enhanced efficiency for said DNA target sequence relative to the efficiency of the starting rare-cutting endonuclease for said DNA target sequence;
(iii) Contacting said DNA target sequence with said chimeric rare-cutting endonuclease for cleavage within the genomic locus of interest; thereby obtaining a cell in which rare-cutting endonuclease efficiency at a genomic locus of interest in a cell is enhanced.
In an embodiment, said enhancer domain is fused to the N-terminus part of said rare-cutting endonuclease-derived scaffold. In another preferred embodiment, said enhancer domain is fused to the C-terminus part of said rare-cutting endonuclease-derived scaffold. In another embodiment, two enhancer domains are fused to both N-terminus part of said rare-cutting endonuclease-derived scaffold and C-terminus part of said rare-cutting endonuclease-derived scaffold. In another embodiment, said enhancer domain(s) are fused to N-terminus and/or C-terminus of a rare-cutting endonuclease-derived monomer scaffold of a homodimeric endonuclease, resulting in a chimeric rare-cutting endonuclease comprising two or four enhancer domains. In another embodiment, said enhancer domain(s) are fused to N-terminus and/or C-terminus of a rare-cutting endonuclease- derived monomer scaffold of a heterodimeric endonuclease, resulting in a chimeric rare-cutting endonuclease comprising one or two or three or four enhancer domains. In another embodiment, said enhancer domain(s) are fused to N-terminus and/or C-terminus of a rare-cutting endonuclease- derived scaffold from a single-chain endonuclease, resulting in a chimeric rare-cutting endonuclease comprising one or two enhancer domains.
In another preferred embodiment, said enhancer domain is catalytically active, or not providing functional and/or structural support to said rare-cutting endonuclease-derived scaffold. In another
preferred embodiment, said enhancer domain consists of a protein domain derived from an exonuclease. In a more preferred embodiment, said enhancer domain consists of a protein domain derived from an endonuclease. In another more preferred embodiment, said enhancer domain consists of a protein domain derived from a cleavase. In another more preferred embodiment, said enhancer domain consists of a protein domain derived from a nickase.
In a more preferred embodiment, said enhancer domain consists of a protein domain derived from a protein selected from the group consisting of Mmel, Colicin-E7 (CEA7_ECOLX), EndA, Endo I (ENDl_ECOLI), Human Endo G (NUCG_HUMAN), Bovine Endo G (NUCG_BOVIN), .HinPll, l-Basl, I- Bmol, l-Hmul, l-Tevl, l-Tevll, l-Tevlll, l-Twol, R.Mspl, R.Mval, NucA, NucM, Vvn, Vvn_CLS, Staphylococcal nuclease (NUC_STAAU), Staphylococcal nuclease (NUC_STAHY), Micrococcal nuclease (NUC_SHIFL), Endonuclease yncB, Endodeoxyribonuclease I (ENRN_BPT7), Metnase, Nb.BsrDI, BsrDI A, Nt.BspD6l (R.BspD6l large subunit), ss.BspD6l (R.BspD6l small subunit), R.PIel, Mlyl, Alwl, Mval269l, Bsrl, Bsml, Nb.BtsCI, Nt.BtsCI, Rl.Btsl, R2.Btsl, BbvCI subunit 1, BbvCI subunit 2, BpulOI alpha subunit, BpulOI beta subunit, Bmrl, Bfil, hExol (EX01_HUMAN), Yeast Exol (EX01_YEAST), E.coli Exol, Human TREX2, Mouse TREX1, Human TREX1, Bovine TREX1, Rat TREX1, Human DNA2, Yeast DNA2 (DNA2_YEAST) and VP16, as listed in Table 1 (SEQ ID NO: 10 to SEQ ID NO: 66),, a functional mutant, a variant or a derivative of these protein domains thereof. In another more preferred embodiment, said enhancer domain consists of a peptide derived from CFPl peptide (SEQ ID NO: 112). In another more preferred embodiment, any combinations of two protein domains selected from the group consisting of Mmel, Colicin-E7 (CEA7_ECOLX), EndA, Endo I (ENDl_ECOLI), Human Endo G (NUCG_HUMAN), Bovine Endo G (NUCG_BOVIN), R.HinPll, l-Basl, l-Bmol, l-Hmul, I- Tevl, l-Tevll, l-Tevlll, l-Twol, R.Mspl, R.Mval, NucA, NucM, Vvn, Vvn_CLS, Staphylococcal nuclease (NUC_STAAU), Staphylococcal nuclease (NUC_STAHY), Micrococcal nuclease (NUC_SHIFL), Endonuclease yncB, Endodeoxyribonuclease I (ENRN_BPT7), Metnase, Nb.BsrDI, BsrDI A, Nt.BspD6l (R.BspD6l large subunit), ss.BspD6l (R.BspD6l small subunit), R.PIel, Mlyl, Alwl, Mval269l, Bsrl, Bsml, Nb.BtsCI, Nt.BtsCI, Rl.Btsl, R2.Btsl, BbvCI subunit 1, BbvCI subunit 2, BpulOI alpha subunit, BpulOI beta subunit, Bmrl, Bfil, hExol (EX01_HUMAN), Yeast Exol (EX01_YEAST), E.coli Exol, Human TREX2, Mouse TREX1, Human TREX1, Bovine TREX1, Rat TREX1, Human DNA2, Yeast DNA2 (DNA2_YEAST) and VP16, as listed in Table 1 (SEQ ID NO: 10 to SEQ ID NO: 66), a functional mutant, a variant or a derivative of these protein domains thereof, can be fused to both N-terminus part of said rare- cutting endonuclease-derived scaffold and C-terminus part of said rare-cutting endonuclease- derived scaffold, respectively. For example, l-Hmul catalytic domain can be fused to the N-terminus part of said rare-cutting endonuclease-derived scaffold and ColE7 can be fused to the C-terminus
part of said rare-cutting endonuclease-derived scaffold. In another more preferred embodiment, said enhancer domain consists of a catalytically active derivative of the protein domains listed above and in Table 1, providing functional and/or structural support to said rare-cutting endonuclease- derived scaffold. In another preferred embodiment, said enhancer domain consists of a catalytically inactive derivative of the protein domains listed above and in Table 1, providing structural support to said rare-cutting endonuclease-derived scaffold.
In a preferred embodiment, said rare-cutting endonuclease-derived scaffold is derived from a meganuclease. In another preferred embodiment, said meganuclease comprises two identical monomers. In another preferred embodiment, said meganuclease comprises two non-identical monomers. In another preferred embodiment, said meganuclease is a single-chain meganuclease. In a more preferred embodiment, said rare-cutting endonuclease-derived scaffold is derived from the group consisting of l-Crel, a functional mutant of l-Crel, a variant of l-Crel or a derivative thereof. In a more preferred embodiment, rare-cutting endonuclease-derived scaffold is a truncated form of wild- type l-Crel (SEQ ID NO: 1). In another more preferred embodiment, rare-cutting endonuclease- derived scaffold comprises first 152, 153, 154 or 155 amino acids residues of wild-type l-Crel (SEQ ID NO: 1). In another more preferred embodiment, rare-cutting endonuclease-derived scaffold comprises residues 2 to 153 of wild-type l-Crel (SEQ ID NO: 1). In another more preferred embodiment, rare-cutting endonuclease-derived scaffold comprises residues 2 to 155 of wild-type I- Crel (SEQ ID NO: 1). In another more preferred embodiment, rare-cutting endonuclease-derived scaffold comprises residues 2 to 153 of wild-type l-Crel (SEQ ID NO: 1) and one or several amino acids substitutions. In another more preferred embodiment, rare-cutting endonuclease-derived scaffold comprises residues 2 to 155 of wild-type l-Crel (SEQ ID NO: 1) and one or several amino acids substitutions. As a non-limiting example, rare-cutting endonuclease-derived scaffold of the present invention comprises residues 2 to 153 or residues 2 to 155 of wild-type l-Crel (SEQ ID NO: 1) and one or two or three or four or five or six or seven or eight or nine or ten further amino acid mutations. As a non-limiting example, rare-cutting endonuclease-derived scaffold of the present invention comprises residues 2 to 153 of wild-type l-Crel (SEQ ID NO: 1) and K82A mutation. In another more preferred embodiment, said rare-cutting endonuclease-derived scaffold comprises a sequence selected from the group consisting of l-Crel_NFSl (SEQ ID NO: 6); l-Crel_NFS2 (SEQ ID NO: 7); and l-Crel_CFSl (SEQ ID NO: 8).
GENBANK/ NAME SEQ FASTA SEQUENCE
SWISS- ID
PROT ID NO
ACC85607.1 Mmel 10 >gi I 186469979 I gb| ACC85607.11 Mmel [Methylophilus methylotrophus]
NLALSWNEIRRKAIEFSKRWEDASDENSQAKPFLIDFFEVFGITNKRVATFEHAVKKFAKAHKEQSRGFVD
LFWPGILLIEMKSRGKDLDKAYDQALDYFSGIAERDLPRYVLVCDFQRFRLTDLITKESVEFLLKDLYQN
VRSFGFIAGYQTQVIKPQDPINIKAAERMGKLHDTLKLVGYEGHALELYLVRLLFCLFAEDTTIFEKSLF
QEYIETKTLEDGSD1_LAHHINTLFYVLNTPEQKRLKNLDEHIJ^FPYINGKLFEEPLPPAQFDKAMREALL
DLCSLDWSRISPAIFGSLFQSIMDAKKRRNLGAHYTSEANILKLIKPLFLDELWVEFEKVKNN NKLLAF
HKKLRGLTFFDPACGCGNFLVITYRELRLLEIEVLRGLHRGGQQVLDIEHLIQINVDQFFGIEIEEFPAQ
IAQVALWLTDHQMNMKISDEFGNYFARIPLKSTPHILNANALQIDWNDVLEAKKCCFILGNPPFVGKSKQ
TPGQKADLLSVFGNLKSASDLDLVAA YPKAAHYIQTNANIRCAFVSTNSITQGEQVSLLWPLLLSLGIK
INFAHRTFSWTNEASGVAAVHCVI IGFGLKDSDEKI IYEYESINGEPLAIKAKNINPYLRDGVDVIACKR
QQPISKLPSMRYGNKPTDDGNFLFTDEEK QFITNEPSSEKYFRRFVGGDEFINNTSRWCLWLDGADISE
IRAMPLVLARIKKVQEFRLKSSAKPTRQSASTPMKFFYISQPDTDYLLIPETSSENRQFIPIGFVDR VI
SSNATYHIPSAEPLIFGLLSSTMHNCWMR VGGRLESRYRYSASLVYNTFP IQPNEKQSKAIEEAAFAI
LKARSNYPNESLAGLYDPKTMPSELLKAHQKLDKAVDSVYGFKGPNTEIARIAFLFETYQKMTSLLPPEK
EIKKSKGK
Q47112.2 Colicin-E7 11 >gi|l2644448|sp|Q47112.2|CEA7_ECOLX RecName : Full=Colicin-E7
(CEA7_EC MSGGDGRGHNSGAHNTGGNINGGPTGLGGNGGASDGSG SSEN PWGGGSGSGVHWGGGSGHGNGGGNSN
OLX) SGGGSNSSVAAPMAFGFPALAAPGAGTLGISVSGEALSAAIADIFAALKGPFKFSA GIALYGILPSEIA
KDDPlnVIMSKIVTSLPAETVTl QVSTLPLDQATVSVTKRVTDVVKDTRQHIAVVAGVPMSVPVV AKPTR
TPGVFHASFPGVPSLTVSTVKGLPVSTTLPRGITEDKGRTAVPAGFTFGGGSHEAVIRFPKESGQKPVYV
SVTDVLTPAQVKQRQDEEKRLQQEWNDAHPVEVAERNYEQARAELNQA KDVARNQERQAKAVQVYNSRK
SELDAA KTLADAKAEIKQFERFAREPMAAGHRMWQMAGLKAQRAQTDV KKAAFDAAAKEKSDADVA
SSALERRKQKENKEKDAKAKLDKESKRNKPGKATGKGKPV KWLN AGKDLGSPVPDRIANKLRDKEFK
SFDDFRKKFWEEVSKDPELSKQFSRInSINDRMKVGKAPKTRTQDVSGKRTSFELHHEKPISQNGGVYDMDN
ISWTPKRHIDIHRGK
21/1
21/2
IADYIIAKLQHRDTSNIEQLLINKNLKMVEFLSKNTKNDNNFTYSE ESIYNGTYRITNLPSLGRFKFRK
KIAEKSLSGKVKEFNNIVQRYSVGLASSDLPFGVIRKESRNDFINDVCKLYNINDMKI IKELKEDADLIV CMLKGFKPRGDDNRPDRGALPLVA LAGENAQIFTFIYGPLIKGAINLIDQDINKLAKRNGLWKSFVSLS DFIVLDCPI IGESYNEFRLI INKNNKESILRKTSKQQNILVDPTPNHYQENDVDTVIYSIFKYIVPNCFS GMCNPPGGDWSGLSI IRNGHEFRWLSLPRVSENGKRPDHVIQILDLFEKPLLLSIESKEKPNDLEPKIGV QLIKYIEYLFDFTPSVQRKIAGGN EFGNKSLVPNDFILLSAGAFIDYDNLTENDYEKIFEVTGCDLLIA IKNQNNPQKWVIKFKPKNTIAEKLVNYIKLNFKSNIFDTGFFHIEG
ADI24225.1 Nb.BtsCI 46 >gi I 297185870 |gb| ADI24225.11 BtsCI bottom- strand nicking enzyme variant
[synthetic construct]
MKRILYLLTEERPKINIIHQI INLEYKATLHFGAKIVPVMNEENKFTFIYHVKGIEVEGFDAVLIKIVSG HSSFVDYLVFDSNDLKPEKNTITLFDLDQYELDLSYYFGKGWIVRIPSPSDLPKYWEETKTDDHESRNT NAYQRSSKFVFCELYYGKEVKKYMLYDISDGRTLSGTDTHNFGMR LVTNNVNLVGVPN YLPFTDIKEF INEKNRIADNGPSHNVPIRLKLDKEKNVIYISAKLDKGNGKNKNKISNDPNIGAVAI ISATLRNLNWKGD IEIINHNLLPSSISSRSNGNKLLYIMKKLGVRFNNII NWNNIKNNINYFFYNITSEKIVSIYYHLYVED KLSNARVIFDNHAGCGKSYFRTLNNKI IPVGKEIPLPALVIFDSDQNIVKVIAAAKAENVYNGVEQLSTF DKFIESYINKYYPGAAVECSVITWGKSSNPYVSFYLDKDGSAVFL
ADI24224.1 N BtsCI 47 >g I 297185868 |gb| ADI24224.11 BtsCI top-strand nicking enzyme variant
[synthetic construct]
MKRILYLLTEERPKINIIHQIINLEYKATLHFGAKIVPVMNEENKFTFIYHVKGIEVEGFDAVLIKIVSG HSSFVDYLVFDSNDLKPEKNTITLFDLDQYELDLSYYFGKGWIVRIPSPSDLPKYWFETKTDDHESRNT NAYQRSSKFVFCELYYGKEVKKYMLYDISDGRTLSGTDTHNFGMRMLVTNNVNLVGVPNMYLPFTDIKEF INEKNRIADNGPSHNVPIRLKLDKEKNVIYISAKLDKGNGKNKNKISNDPNIGAVAIISATLRNLNWKGD IEIINHNLLPSSISSRSNGNKLLYIMKKLGVRFNNINVNWNNIKNNINYFFYNITSEKIVSIYYHLYVED KLSNARVIFDNHAGCGKSYFRTLNNKI IPVGKEIPLPDLVIFDSDQNIVKVIEAEKAENVYNGVEQLSTF DKFIESYINKYYPGAAVECSVITWGKSSNPYVSFYLDKDGSAVFL
>gi I 85720924 IgbIABC75874.11 Rl.BtsI [Geobacillus thermoglucosidasius] MKITEGIVHVAMRHFLKSNGWKLIAGQYPGGSDDELTALNIVDPWARDNSPDPRRHSLGKIVPDLIAYK NDDLLVIEAKPKYSQDDRDKLLYLLSERKHDFYAALEKFATERNHPELLPVSKLNIIPGLAFSASENKFK KDPGFVYIRVSGIFEAFMEGYDWG
ABC75874.1 Rl.BtsI 48 >gi I 85720924 IgbIABC75874.11 Rl.BtsI [Geobacillus thermoglucosidasius]
21/3
21/4
21/5
RSAELSEDDLLSQYSLSFTKKTKKNSSEGNKSLSFSEVFVPDLVNGPTNKKSVSTPPRTRNKFATFLQRK
NEESGAWVPGTRSRFFCSSDSTDCVSNKVSIQPLDETAVTDKENNLHESEYGDQEGKRLVDTDVARNSS DDIPNNHIPGDHIPDKATVFTDEESYSFESSKFTRTISPPTLGTLRSCFSWSGGLGDFSRTPSPSPSTAL QQFRRKSDSPTSLPE MSDVSQLKSEESSDDESHPLREEACSSQSQESGEFSLQSSNASKLSQCSSKDS DSEESDCNIKLLDSQSDQTSKLRLSHFSKKDTPLRNKVPGLYKSSSADSLSTTKIKPLGPARASGLSKKP ASIQKRKHHNAENKPGLQIKLNEL KNFGFKKDSEKLPPCKKPLSPVRDNIQLTPEAEEDIFNKPECGRV QRAIFQ
P39875.2 Yeast Exol 57 1706421 | | P39875.2 | EX01_YEAST RecName : Full=Exodeoxyribonuclease
(EX01_YE 1; AltName: Full=Exodeoxyribonuclease I; Short=EXO I; Short=Exonuclease AST) I; AltName: F ll=Protein DHS1
MGIQGLLPQLKPIQNPVSLRRYEGEVLAIDGYAWLHRAACSCAYELAMGKPTDKYLQFFIKRFSLLKTFK VEPYLVFDGDAIPVKKSTESK-RRDKRKENKAIAERLWACGEKKNAMDYFQKCVDITPEMAKCI ICYCKLN GIRYIVAPFEADSQMVYLEQKNIVQGIISEDSDLLVFGCRRLITKLNDYGECLEICRDNFIKLPKKFPLG SLTNEEIITMVCLSGCDYTNGIPKVGLITA KLVRRFNTIERIILSIQREGKLMIPDTYINEYEAAVLAF QFQRVFCPIRKKIVSLNEIPLYLKDTESKRKRLYACIGFVIHRETQKKQIVHFDDDIDHHLHLKIAQGDL NPYDFHQPLANREHKLQILASKSNIEFGKTNTTNSEAKVKPIESFFQKMTKLDHNPKVANNIHSLRQAEDK LT1XLAIKRRKLSNANVVQETLKDTRSKFFNKPSMTVVENFKEKGDSIQDFKEDTNSQSLEEPVSESQLSTQ IPSSFITTNLEDDDNLSEEVSEWSDIEEDRKNSEGKTIGNEIYNTDDDGDGDTSEDYSETAESRVPTSS TTSFPGSSQRSISGCTKVLQKFRYSSSFSGVNANRQPLFPRHVNQKSRGMVYVNQNRDDDCDDNDGKNQI TQRPSLRKSLIGARSQRIVIDMKSVDERKSFNSSPILHEESKKRDIETTKSSQARPAVRSISLLSQFVYK GK
BAJ43803.1 E.coli Exol 58 >gi 315136644 dbj BAJ43803.1 exonuclease I [Escherichia coli DH1]
MMNDGKQQSTFLFHDYETFGTHPALDRPAQFAAIRTDSEF VIGEPEVFYCKPADDYLPQPGAVLITGIT PQEARAKGENEAAFAARIHSLFTVPKTCILGYNNVRFDDEVTRNIFYRNFYDPYAWSWQHDNSRWDLLDV MRACYALRPEGINWPENDDGLPSFRLEHLTKANGIEHSNAHDAMADVYATIAMAKLVKTRQPRLFDYLFT HRNKHKLMALIDVPQMKPLVHVSGMFGAWRGNTS VAPLA HPENR AVIMVDLAGDISPLLELDSDTLR ERLYTAKTDLGDNAAVPVKLVHINKCPVLAQANTLRPEDADRLGINRQHCLDNLKILRENPQVREKWAI FAEAEPFTPSDNVDAQLYNGFFSDADRAAMKIVLETEPRNLPALDITFVDKRIEKLLFNYRARNFPGTLD YAEQQRWLEHRRQVFTPEFLQGYADELQMLVQQYADDKEKVALLKALWQYAEEIV
Q9BQ50.1 Human 59 >gi 47606206 s Q9BQ50.1 TREX2_HUMAN RecName: Full=Three prime repair
21/6
21/7
MGSQALPHGHMQTLIFLDLEATGLPYSQPKITELCLLAVHRHALENSSMSEGQPPPVPKPPRWDKLSLC
IAPGKPCSSGASEITGLTTAGLEAHGRQRFNDNLATLLQVFLQRQPQPCCLVAHNGDRYDFPLLQAELAS LSVISPLDGTFCVDSIAALKTLEQASSPSEHGPRKSYSLGSIYTRLYGQAPTDSHTAEGDVLALLSICQW KPQALLQWVDKHARPFSTIKPMYGMAATTGTASPRLCAATTSSPLATANLSPSNGRSRGKRPTSPPPENV PEAPSREGLLAPLGLLTFLTLAIAVLYGIFLASPGQ
AAH63664.1 Human 64 >gi I39793966 |gb|AAH63664.1 I DNA2 protein [Homo sapiens]
DNA2 FAIPASRMEQLNELELLMEKSFWEEAELPAELFQKKWASFPRTVLSTGMDNRYLVLAWTVQNKEGNCE
KRLVITASQSLENKELCILRNDWCSVPVEPGD11HLEGDCTSDT 11DKDFGYLILYPDMLISGTSIASS IRCMRRAVLSETFRSSDPATRQ LIGTVLHEVFQKAINNSFAPEKLQELAFQTIQEIRHLKEMYRLNLSQ DEIKQEVEDYLPSFCK AGDFMHKNTSTDFPQMQLSLPSDNSKDNSTCNIEWKP DIEESIWSPRFGLK GKIDVTVGVKIHRGYKTKYKIMPLELKTGKESNSIEHRSQWLYTLLSQERRADPEAGLLLYLKTGQMYP VPANHLDKRELLKLR QMAFSLFHRISKSATRQKTQLASLPQIIEEEKTCKYCSQIGNCALYSRAVEQQM DCSSVPIVMLPKIEEETQHLKQTHLEYFSLWCLMLTLESQSKDNKK HQNIWLMPASEMEKSGSCIGNLI RMEHVKIVCDGQYLHNFQCKHGAIPVTNLMAGDRVIVSGEERSLFALSRGYVKEIN TTVTCLLDRNLSV LPESTLFRLDQEEKNCDIDTPLGNLSKLMENTFVSKKLRDLI IDFREPQFISYLSSVLPHDAKDTVACIL KGLNKPQRQAMKKVLLSKDYTLIVGMPGTGKTTTICTLVPAPEQVEKGGVS VTEAKLIVFLTSIFVKAG CSPSDIGI IAPYRQQLKI INDLLARSIGMVEVNTVDKYQGRDKSIVLVSFVRSNKDGTVGELLKDWRRLN VAITRAKHKLILLGCVPSLNCYPPLEKLLNHLNSEKLI IDLPSREHESLCHILGDFQRE
P38859.1 Yeast 65 >giI 731738 I spIP38859.1 |DNA2_YEAST RecName : Full=DNA replication ATP- DNA2 dependent helicase DNA2
(DNA2 YE MPGTPQKNKRSASISVSPAKKTEEKEIIQNDSKAILSKQTKRKKKYAFAPINNLNGKNTKVSNASVLKSI AST) AVSQVRNTSRTKDINKAVSKSVKQLPNSQVKPKREMSNLSRHHDFTQDEDGPMEEVIWKYSPLQRDMSDK
TTSAAEYSDDYEDVQNPSSTPIVPNRLKTVLSFTNIQVPNADVNQLIQENGNEQVRPKPAEISTRESLRN IDDILDDIEGDLTIKPTITKFSDLPSSPIKAPNVEKKAEVNAEEVDKMDSTGDSNDGDDSLIDILTQKYV EKRKSESQITIQGNTNQKSGAQESCGKNDNTKSRGEIEDHENVDNQAKTGNAFYENEEDSNCQRIKKNEK IEYNSSDEFSDDSLIELLNETQTQVEPNTIEQDLDKVEKMVSDDLRIATDSTLSAYALRAKSGAPRDGW RLVIVSLRSVELPKIGTQKILECIDGKGEQSSVVVRHPWVYLEFEVGDVIHI IEGKNIENKRLLSDDKNP KTQLANDNLLVLNPDVLFSATSVGSSVGCLRRSILQMQFQDPRGEPSLVMTLGNIVHELLQDSIKYKLSH NKISMEI I IQKLDSLLETYSFSII ICNEEIQYVKELVMKEHAENILYFVNKFVSKSNYGCYTSISGTRRT QPISISNVIDIEENIWSPIYGLKGFLDATVEANVENNKKHIVPLEVKTGKSRSVSYEVQGLIYTLLLNDR YEIPIEFFLLYFTRDKNMTKFPSVLHSIKHILMSRNRMSMNFKHQLQEVFGQAQSRFELPPLLRDSSCDS
21/8
In a preferred embodiment, said chimeric rare-cutting endonuclease can comprise at least one peptidic linker between said rare-cutting endonuclease-derived scaffold and said at least one enhancer domain. In a more preferred embodiment, said peptidic linker sequence is selected from the group consisting of
NFSl, NFS2, CFSl, M2, BQY, QGPSG, LGPDG KA, la8h_l, ldnpA_l, ld8cA_2, lckqA_3, lsbp_l, lev7A_l, lalo_3, lamf l, ladjA_3, lfcdC_l, lal3_2, lg3p_l, lacc_3, lahjB_l, lacc_l, laf7_l, lheiA_l, lbia_2, ligtB_l, lnfkA_l, lau7A_l, lbpoB_l, lb0pA_2, lc05A_2, lgcb_l, lbt3A_l, lb3oB_2, 16vpA_6, ldhx_l, lb8aA_land lqu6A_l, as listed in Table 2 (SEQ ID NO: 67 to SEQ ID NO: 104). In a more preferred embodiment, the peptidic linker that can link said enhancer domain to the rare-cutting endonuclease-derived scaffold according to the method of the present invention can be selected from the group consisting of NFSl (SEQ ID NO: 98), NFS2 (SEQ ID NO: 99) and CFSl (SEQ ID NO: 100). In the scope of the present invention is also encompassed the case where a peptidic linker is not needed to fuse said enhancer domain to said rare-cutting endonuclease-derived scaffold in order to obtain a chimeric rare-cutting endonuclease according to the present invention.
21/10
lbia_2 268 - 276 9 16,089 LDNFINRPV 84 l igtB l 1 1 1 - 1 19 9 19,737 VSSAKTTAP 85 lnfkA l 239 - 248 10 13,228 DSKAPNASNL 86 lau7A 1 103 - 1 12 10 20,486 KRRTTISIAA 87 lbpoB l 138 - 148 1 1 21,645 PVKMFDRHSSL 88 lb0pA_2 625 - 635 1 1 26,462 APAETKAEPMT 89 lc05A 2 135 - 148 14 23,819 YTRLPERSELPAEI 90 lgcbj 57 - 70 14 27,39 VSTDSTPVTNQ SS 91 lbt3A_l 38 - 51 14 28,818 YKLPAVTTM VRPA 92 lb3oB_2 222 - 236 15 20,054 IARTDLKKNRDYPLA 93
16vpA 6 312 - 332 21 23,713 TEEPGAPLTTPPTLHGNQARA 94 l dhx l 81 - 101 21 42,703 ARFTLAVGDNRVLDMASTYFD 95 lb8aA_l 95 - 120 26 31 ,305 IVVLNRAETPLPLDPTG VKAELDTR 96 lqu6A 1 79 - 106 28 51 ,301 ILNKE AVSPLLLTTTNSSEGLSMGNY 97
NFS1 - 20 - GSDITKSKISEKMKGQGPSG 98
NFS2 - 23 - GSDIT SKISEKMKG LGPDGRKA 99
CFS1 - 10 - SLTKSKISGS 100
RM2 - 32 - AAGGSALTAGALSLTAGALSLTAGALSGGGGS 101
BQY - 27 - AAGASSVSASGH1APLSLPSSPPSVGS 102
QGPSG - 5 - QGPSG 103
LGPDGR A - 8 - LGPDGRKA 104
Table 2: List of peptidic linkers that can be used in chimeric rare-cutting endonuclease.
Enhancement of efficiency of a chimeric rare-cutting endonuclease according to the present invention, compared to a starting rare-cutting endonuclease, can also be a consequence of a fuctional support by said at least one enhancer domain. In a preferred embodiment, said functional support can be the consequence of the hydrolysis of additional phosphodiester bonds. In a more preferred embodiment, said functional support can be the hydrolysis of additional phosphodiester bonds by a protein domain derived from a nuclease. In a more preferred embodiment, said functional support can be the hydrolysis of additional phosphodiester bonds by a protein domain derived from an endonuclease. In a more preferred embodiment, said functional support can be the hydrolysis of additional phosphodiester bonds by a protein domain derived from a cleavase. In another more preferred embodiment, said functional support can be the hydrolysis of additional phosphodiester bonds by a protein domain derived from a nickase. In a more preferred embodiment, said functional support can be the hydrolysis of additional phosphodiester bonds by a protein domain derived from an exonuclease.
22
32386
Cleavage by rare-cutting endonucleases usually generates cohesive ends, with 3' overhangs for LAGLIDADG meganucleases (Chevalier and Stoddard 2001) and 5' overhangs for Zinc Finger Nucleases (Smith, Bibikova et al. 2000) These ends, which result from hydrolysis of phosphodiester bonds, can be re-ligated in vivo by N HEJ in a seamless way (i.e a scarless re-ligation). The restoration of a cleavable target sequence allows for a new cleavage event by the same endonuclease, and thus, a series of futile cycles of cleavage and re-ligation events can take place (Figure 4A). In a preferred embodiment, enhancement of efficiency of a chimeric rare-cutting endonuclease according to the present invention, compared to a starting rare-cutting endonuclease, can be the hydrolysis of additional phosphodiester bonds at the cleavage site to promote exit from such futile cycles of cleavage and re-ligation events via imprecise NHEJ or Homologous Recombination or Single Strand Annealing (SSA). Said hydrolysis of additional phosphodiester bonds at the cleavage site by said at least one enhancer according to the invention can lead to different types of DSB resection affecting at said DSB cleavage site, one single DNA strand or both DNA strands, affecting either 5' overhangs ends, either 3' overhangs ends, or both ends and depending on the length of said resection. Thus, adding new nickase or cleavase activities to the existing cleavase activity of a rare-cutting endonuclease enhances the efficiency of the resulting chimeric rare-cutting endonuclease according to the invention, at a genomic locus of interest. As a non-limiting example, addition of two nickase activities on opposite strands (as depicted on Figure 4D) or of a new cleavase activity generating a second DSB (Figure 4E) can result in a gap. As a consequence, perfect re-ligation is not possible anymore, and one or several alternative repair outcomes such as imprecise NHEJ, Homologous Recombination or SSA for instance, can be stimulated. As another non-limiting example, the addition of a single nickase activity can result in a single strand gap, and suppress the cohesivity of the ends, which can also enhances the efficiency of the resulting chimeric rare-cutting endonuclease at a genomic locus of interest, according to the invention, via stimulation of one or several alternative repair outcomes mentioned above.
In this first aspect of the invention, the method according to the invention enhances rare-cutting endonuclease efficiency for a DNA target sequence. Enhancement of efficiency of a rare-cutting endonuclease according to the present invention can be a consequence of a structural support by said at least one enhancer domain. In a preferred embodiment, said structural support enhances the binding of a chimeric rare-cutting endonuclease according to the invention for a DNA target sequence compared to the binding of a starting rare-cutting endonuclease for the same DNA target sequence, as a non-limiting example. In another preferred embodiment, said structural support by at least one enhancer domain enhances the existing catalytical activity of a chimeric rare-cutting
23
endonuclease according to the invention for a DNA target sequence compared to the binding of a starting rare-cutting endonuclease for the same DNA target sequence, as another non-limiting example. In another preferred embodiment, said at least one enhancer domain enhances both the binding and the existing catalytical activity of a chimeric rare-cutting endonuclease according to the invention for a DNA target sequence compared to the binding of a starting rare-cutting endonuclease for the same DNA target sequence, as another non-limiting example. All these non- limiting examples lead to a chimeric rare-cutting endonuclease with enhanced efficiency for a DNA target at a genomic locus of interest, according to the present invention. Figure 4
Depending on number (one to four) of enhancer domains to the starting rare-cutting endonuclease-derived scaffold according to the invention and on their catalytic activities (functional support) or not (structural support), the resulting chimeric rare-cutting endonuclease can comprise several enzymatic activities that contribute to the enhancement of its efficiency according to the present invention. As a non-limiting example, the addition of one nickase domain to an active rare- cutting endonuclease scaffold derived from a meganuclease results in a chimeric rare-cutting endonuclease with one meganuclease activity and one nickase activity (depicted in figure 4B). As another non-limiting example, the addition of two nickase domains to an active rare-cutting endonuclease scaffold derived from a meganuclease results in a chimeric rare-cutting endonuclease with one meganuclease activity and two nickase activities (as depicted in figures 5C and 5D). As a non-limiting example, the addition of one cleavase domain to an active rare-cutting endonuclease scaffold derived from a meganuclease results in a chimeric rare-cutting endonuclease with one meganuclease activity and one cleavase activity (figure 4E). As another non-limiting example, the addition of two cleavase domains to an active rare-cutting endonuclease scaffold derived from a meganuclease results in a chimeric rare-cutting endonuclease with one meganuclease activity and two cleavase activities.
In this aspect of the present invention, enhancement of efficiency of a rare-cutting endonuclease refers to the increase in the detected level of said efficiency, against a target DNA sequence, of a second rare-cutting endonuclease in comparison to the activity of a first rare-cutting endonuclease against the same target DNA sequence. Said second rare-cutting endonuclease can be an engineered rare-cutting endonuclease, i.e. a chimeric rare-cutting endonuclease according to the invention or not. Said first rare-cutting endonuclease can be a wild-type rare-cutting endonuclease, a starting rare-cutting endonuclease, an engineered rare-cutting endonuclease i.e a chimeric rare-cutting
24
12 032386 endonuclease according to the invention or not, taken as a reference scaffold for enhancement in efficiency. Additional rounds of enhancement can be envisioned on a starting rare-cutting endonuclease scaffold. Said enhancement of efficiency can be determined, as non-limiting example, by measuring the level of cleavage-induced recombination generated by said rare-cutting endonuclease or said chimeric rare-cutting endonuclease according to the present invention. Said level of cleavage-induced recombination can be measured by a cell-based recombination assay as described in the International PCT Application WO 2004/067736, as non-limiting example. Importantly, enhancement of efficacy in cells (enhanced generation of targeted mutagenesis or targeted recombination) can be, but is not necessarily associated with an enhancement of the cleavage activity that could be detected in certain in vitro assays. For example, additional phosphodiesterase activities as described in Figure 4 could barely affect the cleavage profile, as detected by in vitro cleavage and separation of the cleavage products on an electrophoresis gel. However, as explained above, and in the legend of Figure 4, the DSB ends generated in this way could be more prone to induce detectable genomic rearrangements such as targeted mutagenesis (by imprecise NHEJ) or homologous recombination.
Said enhancement in efficacy of said rare-cutting endonuclease is at least a 5% enhancement compared to the starting scaffold, more preferably at least a 10% enhancement, again more preferably at least a 15% enhancement, again more preferably at least a 20% enhancement, again more preferably at least a 25% enhancement, again more preferably a 50% enhancement, again more preferably a enhancement greater than 50%.
In another preferred embodiment, the enhancement of efficiency of a rare-cutting endonuclease according to the method of the present invention can also be an enhancement in the efficiency for 2NN derivatives of a DNA target sequence by a rare-cutting endonuclease from the LAGLIDADG meganuclease subfamily. By 2NN derivatives of a DNA target sequence is intended variants of a given DNA target sequence having mutations in the four central nucleotides -2 to +2 as numbered in figure 2. For instance, the method according to the invention enhances the efficiency for derivatives of the DNA target sequence C1221 (SEQ ID NO: 2) having mutations in the four central nucleotides -2 to +2 as numbered in figure 2, i.e that do not have GTAC sequence at position -2 to +2 as numbered in figure 2. In another preferred embodiment, the method according to the invention allows the recombination to be initiated at 2NN derivatives that are not normally processed by a starting rare- cutting endonuclease scaffold from the LAGLIDADG meganuclease subfamily. As a non limiting
25
example, the method according to the invention allows the processing of 2NN derivatives that have - 2 to +2 target sequences as listed in Table 4 and Table 5, non-originally cleaved by l-Crel.
In this first aspect of the invention is the method according to the invention wherein said chimeric rare-cutting endonuclease is selected from the group consisting of hHmuCre_D0101 (SEQ ID NO: 9), hHmuCre_D0201 (SEQ ID NO: 105), hHmuCre_D0301 (SEQ ID NO: 106), hHmuCre_D0401 (SEQ ID NO: 107), hHmuCre_D0102 (SEQ ID NO: 108), hHmuCre_D0202 (SEQ ID NO: 109), hHmuCre_D0302 (SEQ ID NO: 110) and hHmuCre_D0402 (SEQ ID NO: 111).
According to the present invention, efficiency of rare-cutting endonucleases such as meganucleases can be enhanced through the addition of a domain to promote existing or alternate activities. As each end of the meganuclease scaffold, for instance, is amenable to fusion, the order (N- v.s C- terminal) of addition and number of the enhancer domains can vary with the application as depicted in Figure 3. Enhanced fusion construct are optimized to address or overcome distinct problems. (Figure 3A) The addition of two enhancer domains to an active rare-cutting endonuclease such as a meganuclease can enhance DNA binding and/or cleavage activity. Such a configuration can be achieved via (i) a single N- or C-terminal fusion to a homodimeric variant; (ii) a single N- or C-terminal fusion to individual monomers of a heterodimer, or; (iii) a double fusion to a monomeric protein. (Figure 3B) When specificity reengineering precludes maintaining cleavage activity of the rare- cutting endonuclease such as a meganuclease, the attached enhancer domains can provide alternative functions. (Figure 3C) and (Figure 3D) represent instances of (Figure 3A) and (Figure 3B), respectively, when only one enhancer domain is needed or tolerated per fusion protein (e.g. either as an N- or C-terminal fusion or in the context of a single-chain molecule). Fusion junctions (N- vs. C- terminal) and linker designs can vary with the application.
In a preferred embodiment, the present invention can relate to a method for increasing targeted H (and NHEJ) when efficiency is enhanced in a chimeric rare-cutting endonuclease targeting a DNA target sequence according to the invention. In another more preferred embodiment, said efficiency can be enhanced by the addition of a catalytically active cleavase enhancer domain according to the invention, providing functional and/or structural support to the starting rare-cutting endonuclease scaffold according to the invention. In another more preferred embodiment, said efficiency can be enhanced by the addition of two catalytically active cleavase enhancer domains according to the invention, providing functional and/or structural support to the starting rare-cutting endonuclease scaffold according to the invention. In another more preferred embodiment, the addition of at least
26
two cataiyticaily active cleavase enhancer domains according to the invention can allow an increase in Double-strand break-induced mutagenesis by leading to a loss of genetic information between two nearby DNA double strand breaks and thus preventing any scarless re-ligation of targeted genomic locus of interest by NHEJ.
In another preferred embodiment, the present invention can relate to a method for increasing targeted Homologous Recombination (HR) with less NHEJ (i.e in a more conservative fashion) when at least one single-strand break activity is promoted in a chimeric rare-cutting endonuclease targeting a DNA target sequence according to the invention. In a more preferred embodiment, the efficiency of said chimeric rare-cutting endonuclease could be enhanced by the addition of a cataiyticaily active nickase enhancer domain according to the invention. In another more preferred embodiment, the efficiency of said chimeric rare-cutting endonuclease could be enhanced by the addition of two cataiyticaily active nickase enhancer domains according to the invention, all nickase enhancer domains being able to cut the same DNA strand. More preferably, the addition of a least one nickase enhancer domain according to the invention can be performed on an inactive rare- cutting endonuclease scaffold which serves as a scaffold for targeted DNA binding.
In another preferred embodiment, the present invention can relate to a method for increasing excision of a single-strand of DNA spanning the starting rare-cutting endonuclease DNA binding region when both one cleavase enhancer domain and one nickase enhancer domain, respectively, are fused to both N-terminus and C-terminus of a starting rare-cutting endonuclease scaffold according to the invention.
In a more preferred embodiment, both cleavase enhancer domain and nickase enhancer domain can be fused on an inactive rare-cutting endonuclease scaffold which serves as a scaffold for targeted DNA binding.
27
In another aspect, the present invention concerns a method for the creation of fusion proteins that consist of engineering a fusion protein between a rare-cutting endonuclease-derived scaffold and at least one enhancer domain wherein said enhancer domain enhances the efficiency of said chimeric rare-cutting endonuclease when fused to it.
According to this aspect of the invention is a method to create a chimeric rare-cutting endonuclease with enhanced efficiency for a DNA target sequence comprising the steps of:
(i) Engineering a rare-cutting endonuclease-derived scaffold;
(ii) Determining or engineering an enhancer domain wherein said enhancer domain enhances the efficiency of said rare-cutting endonuclease when fused to it;
(iii) Optionally determining or engineering a peptidic linker to fuse said rare-cutting endonuclease-derived scaffold to said enhancer domain; thereby obtaining a chimeric rare-cutting endonuclease with enhanced efficiency for a DNA target sequence.
In an embodiment, said enhancer domain is fused to the N-terminus part of said rare-cutting endonuclease-derived scaffold. In another preferred embodiment, said enhancer domain is fused to the C-terminus part of said rare-cutting endonuclease-derived scaffold. In another embodiment, two enhancer domains are fused to both N-terminus part of said rare-cutting endonuclease-derived scaffold and C-terminus part of said rare-cutting endonuclease-derived scaffold. In another embodiment, said enhancer domain(s) are fused to N-terminus and/or C-terminus of a rare-cutting endonuclease-derived monomer scaffold of a homodimeric endonuclease, resulting in a chimeric rare-cutting endonuclease comprising two or four enhancer domains. In another embodiment, said enhancer domain(s) are fused to N-terminus and/or C-terminus of a rare-cutting endonuclease- derived monomer scaffold of a heterodimeric endonuclease, resulting in a chimeric rare-cutting endonuclease comprising one or two or three or four enhancer domains. In another embodiment, said enhancer domain(s) are fused to N-terminus and/or C-terminus of a rare-cutting endonuclease- derived scaffold from a single-chain endonuclease, resulting in a chimeric rare-cutting endonuclease comprising one or two enhancer domains.
In a preferred embodiment, said enhancer domain is catalytically active or not, providing functional and/or structural support to said rare-cutting endonuclease-derived scaffold. In another preferred embodiment, said enhancer domain consists of a protein domain derived from an exonuclease. In a
28
more preferred embodiment, said enhancer domain consists of a protein domain derived from an endonuclease. In another more preferred embodiment, said enhancer domain consists of a protein domain derived from a cleavase. In another more preferred embodiment, said enhancer domain consists of a protein domain derived from a nickase.
In a more preferred embodiment, said enhancer domain consists of a protein domain derived from a protein selected from the group consisting of Mmel, Colicin-E7 (CEA7_ECOLX), EndA, Endo I (ENDl_ECOLI), Human Endo G (NUCG_HUMAN), Bovine Endo G (NUCG_BOVIN), R.HinPll, l-Basl, I- Bmol, l-Hmul, l-Tevl, l-Tevll, l-Tevlll, l-Twol, R.Mspl, R.Mval, NucA, NucM, Vvn, Vvn_CLS, Staphylococcal nuclease (NUC_STAAU), Staphylococcal nuclease (NUC_STAHY), Micrococcal nuclease (NUC_SHIFL), Endonuclease yncB, Endodeoxyribonuclease I (ENRN_BPT7), Metnase, Nb.BsrDI, BsrDI A, Nt.BspD6l (R.BspD6l large subunit), ss.BspD6l (R.BspD6l small subunit), R.PIel, Mlyl, Alwl, Mval269l, Bsrl, Bsml, Nb.BtsCI, Nt.BtsCI, Rl.Btsl, R2.Btsl, BbvCI subunit 1, BbvCI subunit 2, BpulOI alpha subunit, BpulOI beta subunit, Bmrl, Bfil, hExol (EX01_HUMAN), Yeast Exol (EX01_YEAST), E.coli Exol, Human TREX2, Mouse TREX1, Human TREX1, Bovine TREX1, Rat TREX1, Human DNA2, Yeast DNA2 (DNA2_YEAST) and VP16, as listed in Table 1 (SEQ ID NO: 10 to SEQ ID NO: 66), a functional mutant, a variant or a derivative of these protein domains thereof. In another more preferred embodiment, said enhancer domain consists of a peptide derived from CFPl peptide (SEQ ID NO: 112). In another more preferred embodiment, any combinations of two protein domains selected from the group consisting of Mmel, Colicin-E7 (CEA7_ECOLX), EndA, Endo I (ENDl_ECOLI), Human Endo G (NUCG_HUMAN), Bovine Endo G (NUCG_BOVIN), R.HinPll, l-Basl, l-Bmol, l-Hmul, I- Tevl, l-Tevll, l-Tevlll, l-Twol, R.Mspl, R.Mval, NucA, NucM, Vvn, Vvn_CLS, Staphylococcal nuclease (NUC_STAAU), Staphylococcal nuclease (NUC_STAHY), Micrococcal nuclease (NUC_SHIFL), Endonuclease yncB, Endodeoxyribonuclease I (ENRN_BPT7), Metnase, Nb.BsrDI, BsrDI A, Nt.BspD6l (R.BspD6l large subunit), ss.BspD6l (R.BspD6l small subunit), R.PIel, Mlyl, Alwl, Mval269l, Bsrl, Bsml, Nb.BtsCI, Nt.BtsCI, Rl.Btsl, R2.Btsl, BbvCI subunit 1, BbvCI subunit 2, BpulOI alpha subunit, BpulOI beta subunit, Bmrl, Bfil, hExol (EX01_HUMAN), Yeast Exol (EX01_YEAST), E.coli Exol, Human TREX2, Mouse TREX1, Human TREX1, Bovine TREX1, Rat TREX1, Human DNA2, Yeast DNA2 (DNA2_YEAST) and VP16, as listed in Table 1 (SEQ ID NO: 10 to SEQ ID NO: 66 and SEQ ID NO: 1), a functional mutant, a variant or a derivative of these protein domains thereof, can be fused to both N-terminus part of said rare-cutting endonuclease-derived scaffold and C-terminus part of said rare-cutting endonuclease-derived scaffold, respectively. For example, l-Hmul catalytic domain can be fused to the N-terminus part of said rare-cutting endonuclease-derived scaffold and ColE7 can be fused to the C-terminus part of said rare-cutting endonuclease-derived scaffold. In another more preferred
29
P T/US2012/032386 embodiment, said enhancer domain consists of a catalytically active derivative of the protein domains listed above and in Table 1, providing functional and/or structural support to said rare- cutting endonuclease-derived scaffold. In another preferred embodiment, said enhancer domain consists of a catalytically inactive derivative of the protein domains listed above and in Table 1, providing structural support to said rare-cutting endonuclease-derived scaffold.
In a preferred embodiment, said rare-cutting endonuclease-derived scaffold is derived from a meganuclease. In another preferred embodiment, said meganuclease comprises two identical monomers. In another preferred embodiment, said meganuclease comprises two non-identical monomers. In another preferred embodiment, said meganuclease is a single-chain meganuclease. In a more preferred embodiment, said rare-cutting endonuclease-derived scaffold is derived from the group consisting of !-Crel, a functional mutant of l-Crel, a variant of l-Crel or a derivative thereof. In a more preferred embodiment, rare-cutting endonuclease-derived scaffold is a truncated form of wild- type l-Crel (SEQ ID NO: 1). In another more preferred embodiment, rare-cutting endonuclease- derived scaffold comprises first 152, 153, 154 or 155 amino acids residues of wild-type l-Crel (SEQ ID NO: 1). In another more preferred embodiment, rare-cutting endonuclease-derived scaffold comprises residues 2 to 153 of wild-type l-Crel (SEQ ID NO: 1). In another more preferred embodiment, rare-cutting endonuclease-derived scaffold comprises residues 2 to 155 of wild-type I- Crel (SEQ ID NO: 1). In another more preferred embodiment, rare-cutting endonuclease-derived scaffold comprises residues 2 to 153 of wild-type l-Crel (SEQ ID NO: 1) and one or several amino acids substitutions. In another more preferred embodiment, rare-cutting endonuclease-derived scaffold comprises residues 2 to 155 of wild-type l-Crel (SEQ ID NO: 1) and one or several amino acids substitutions. As a non-limiting example, rare-cutting endonuclease-derived scaffold of the present invention comprises residues 2 to 153 or residues 2 to 155 of wild-type l-Crel (SEQ ID NO: 1) and one or two or three or four or five or six or seven or eight or nine or ten further amino acid mutations. As a non-limiting example, rare-cutting endonuclease-derived scaffold of the present invention comprises residues 2 to 153 of wild-type l-Crel (SEQ ID NO: 1) and K82A mutation. In another more preferred embodiment, said rare-cutting endonuclease-derived scaffold comprises a sequence selected from the group consisting of l-Crel_NFSl (SEQ ID NO: 6); l-Crel_NFS2 (SEQ ID NO: 7); and l-Crel_CFSl (SEQ ID NO: 8).
In a preferred embodiment, said chimeric rare-cutting endonuciease can comprise at least one peptidic linker between said rare-cutting endonuclease-derived scaffold and said at least one enhancer domain. In a more preferred embodiment, said peptidic linker sequence is selected from the group consisting of
2012/032386
NFS1, NFS2, CFS1, RM2, BQY, QGPSG, LGPDGRKA, la8h_l, ldnpA_l, ld8cA_2, lckqA_3, lsbp_l, lev7A_l, lalo_3, lamf_l, ladjA_3, lfcdC_l, lal3_2, lg3p_l, lacc_3, lahjB_l, lacc_l, laf7_l, lheiA_l, lbia_2, ligtB_l, lnfkA_l, lau7A_l, lbpoB_l, lb0pA_2, lc05A_2, lgcb_l, lbt3A_l, lb3oB_2, 16vpA_6, ldhx_l, lb8aA_land lqu6A_l, as listed in Table 2 (SEQ ID NO: 67 to SEQ ID NO: 104). In a more preferred embodiment, the peptidic linker that can link said enhancer domain to the rare-cutting endonuclease-derived scaffold according to the method of the present invention can be selected from the group consisting of NFS1 (SEQ ID NO: 98), NFS2 (SEQ ID NO: 99) and CFS1 (SEQ ID NO: 100). In the scope of the present invention is also encompassed the case where a peptidic linker is not needed to fuse said enhancer domain to said rare-cutting endonuclease-derived scaffold in order to obtain a chimeric rare-cutting endonuclease according to the present invention.
Enhancement of efficiency of a chimeric rare-cutting endonuclease according to the present invention, compared to a starting rare-cutting endonuclease, can also be a consequence of a fuctional support by said at least one enhancer domain. In a preferred embodiment, said functional support can be the consequence of the hydrolysis of additional phosphodiester bonds. In a more preferred embodiment, said functional support can be the hydrolysis of additional phosphodiester bonds by a protein domain derived from a nuclease. In a more preferred embodiment, said functional support can be the hydrolysis of additional phosphodiester bonds by a protein domain derived from an endonuclease. In a more preferred embodiment, said functional support can be the hydrolysis of additional phosphodiester bonds by a protein domain derived from a cleavase. In another more preferred embodiment, said functional support can be the hydrolysis of additional phosphodiester bonds by a protein domain derived from a nickase. In a more preferred embodiment, said functional support can be the hydrolysis of additional phosphodiester bonds by a protein domain derived from an exonuclease.
Cleavage by rare-cutting endonucleases usually generates cohesive ends, with 3' overhangs for LAGLIDADG meganucleases {Chevalier, 2001 #6} and 5' overhangs for Zinc Finger Nucleases {Smith, 2000 #7}. These ends, which result from hydrolysis of phosphodiester bonds, can be re-ligated in vivo by NHEJ in a seamless way (i.e a scarless re-ligation). The restoration of a cleavable target sequence allows for a new cleavage event by the same endonuclease, and thus, a series of futile cycles of cleavage and re-ligation events can take place (Figure 4A). In a preferred embodiment, enhancement of efficiency of a chimeric rare-cutting endonuclease according to the present invention, compared to a starting rare-cutting endonuclease, can be the hydrolysis of additional
31
phosphodiester bonds at the cleavage site to promote exit from such futile cycles of cleavage and re- ligation events via imprecise NHEJ or Homologous Recombination or Single Strand Annealing (SSA). Said hydrolysis of additional phosphodiester bonds at the cleavage site by said at least one enhancer according to the invention can lead to different types of DSB resection affecting at said DSB cleavage site, one single DNA strand or both DNA strands, affecting either 5' overhangs ends, either 3' overhangs ends, or both ends and depending on the length of said resection. Thus, adding new nickase or cleavase activities to the existing cleavase activity of a rare-cutting endonuclease enhances the efficiency of the resulting chimeric rare-cutting endonuclease according to the invention, at a genomic locus of interest. As a non-limiting example, addition of two nickase activities on opposite strands (as depicted on Figure 4D) or of a new cleavase activity generating a second DSB (Figure 4E) can result in a gap. As a consequence, perfect re-ligation is not possible anymore, and one or several alternative repair outcomes such as imprecise NHEJ, Homologous Recombination or SSA for instance, can be stimulated. As another non-limiting example, the addition of a single nickase activity can result in a single strand gap, and suppress the cohesivity of the ends, which can also enhances the efficiency of the resulting chimeric rare-cutting endonuclease at a genomic locus of interest, according to the invention, via stimulation of one or several alternative repair outcomes mentioned above.
In this second aspect of the invention, the method according to the invention enhances rare-cutting endonuclease efficiency for a DNA target sequence. Enhancement of efficiency of a rare-cutting endonuclease according to the present invention can be a consequence of a structural support by said at least one enhancer domain. In a preferred embodiment, said structural support enhances the binding of a chimeric rare-cutting endonuclease according to the invention for a DNA target sequence compared to the binding of a starting rare-cutting endonuclease for the same DNA target sequence, as a non-limiting example. In another preferred embodiment, said structural support by at least one enhancer domain enhances the existing catalytical activity of a chimeric rare-cutting endonuclease according to the invention for a DNA target sequence compared to the binding of a starting rare-cutting endonuclease for the same DNA target sequence, as another non-limiting example. In another preferred embodiment, said at least one enhancer domain enhances both the binding and the existing catalytical activity of a chimeric rare-cutting endonuclease according to the invention for a DNA target sequence compared to the binding of a starting rare-cutting endonuclease for the same DNA target sequence, as another non-limiting example. All these non- limiting examples lead to a chimeric rare-cutting endonuclease with enhanced efficiency for a DNA target at a genomic locus of interest, according to the present invention.
32
Depending on number (one to four) of enhancer domains to the starting rare-cutting endonuclease- derived scaffold according to the invention and on their catalytic activities (functional support) or not (structural support), the resulting chimeric rare-cutting endonuclease can comprise several enzymatic activities that contribute to the enhancement of its efficiency according to the present invention. As a non-limiting example, the addition of one nickase domain to an active rare-cutting endonuclease scaffold derived from a meganuclease results in a chimeric rare-cutting endonuclease with one meganuclease activity and one nickase activity (depicted in figure 4B). As another non- limiting example, the addition of two nickase domains to an active rare-cutting endonuclease scaffold derived from a meganuclease results in a chimeric rare-cutting endonuclease with one meganuclease activity and two nickase activities (as depicted in figures 5C and 5D). As a non-limiting example, the addition of one cleavase domain to an active rare-cutting endonuclease scaffold derived from a meganuclease results in a chimeric rare-cutting endonuclease with one meganuclease activity and one cleavase activity (figure 4E). As another non-limiting example, the addition of two cleavase domains to an active rare-cutting endonuclease scaffold derived from a meganuclease results in a chimeric rare-cutting endonuclease with one meganuclease activity and two cleavase activities.
In this aspect of the present invention, enhancement of efficiency of a rare-cutting endonuclease refers to the increase in the detected level of said efficiency, against a target DNA sequence, of a second rare-cutting endonuclease in comparison to the activity of a first rare-cutting endonuclease against the same target DNA sequence. Said second rare-cutting endonuclease can be an engineered rare-cutting endonuclease, i.e. a chimeric rare-cutting endonuclease according to the invention or not. Said first rare-cutting endonuclease can be a wild-type rare-cutting endonuclease, a starting rare-cutting endonuclease, an engineered rare-cutting endonuclease i.e a chimeric rare-cutting endonuclease according to the invention or not, taken as a reference scaffold for enhancement in efficiency. Additional rounds of enhancement can be envisioned on a starting rare-cutting endonuclease scaffold. Said enhancement of efficiency can be determined, as non-limiting example, by measuring the level of cleavage-induced recombination generated by said rare-cutting endonuclease or said chimeric rare-cutting endonuclease according to the present invention. Said level of cleavage-induced recombination can be measured by a cell-based recombination assay as described in the International PCT Application WO 2004/067736, as non-limiting example. Importantly, enhancement
33
of efficacy in cells (enhanced generation of targeted mutagenesis or targeted recombination) is not necessarily associated with an enhancement of the cleavage activity that could be detected in certain in vitro assays. For example, additional phosphodiesterase activities as described in Figure 4 could barely affect the cleavage profile, as detected by in vitro cleavage and separation of the cleavage products on an electrophoresis gel. However, as explained above, and in the legend of Figure 4, the DSB ends generated in this way could be more prone to induce detectable genomic rearrangements such as targeted mutagenesis (by imprecise NHEJ) or homologous recombination.
Said enhancement in efficiency of said rare-cutting endonuclease is at least a 5% enhancement compared to the starting scaffold, more preferably at least a 10% enhancement, again more preferably at least a 15% enhancement, again more preferably at least a 20% enhancement, again more preferably at least a 25% enhancement, again more preferably a 50% enhancement, again more preferably a enhancement greater than 50%. In another preferred embodiment, the enhancement of efficiency of a rare-cutting endonuclease according to the method of the present invention can also be an enhancement in the efficiency for 2NN derivatives of a DNA target sequence by a rare-cutting endonuclease from the LAGLIDADG meganuclease subfamily. By 2NN derivatives of a DNA target sequence is intended variants of a given DNA target sequence having mutations in the four central nucleotides -2 to +2 as numbered in figure 2. For instance, the method according to the invention enhances the efficiency for derivatives of the DNA target sequence C1221 having mutations in the four central nucleotides -2 to +2 as numbered in figure 2, i.e that do not have GTAC sequence at position -2 to +2 as numbered in figure 2. In another preferred embodiment, the method according to the invention allows recombination to be initiated at 2NN derivatives that are not normally processed by a starting rare-cutting endonuclease scaffold from the LAGLIDADG meganuclease subfamily. As a non limiting example, the method according to the invention allows the processing of 2NN derivatives that have -2 to +2 target sequences as listed in Table 4 and Table 5, non-originally cleaved by l-Crel.
The present invention also concerns the creation of functional single polypeptide fusion proteins for simple and efficient vectorization.
34
2012/032386
In another aspect, the present invention relates to chimeric rare-cutting endonucleases comprising at least an enhancer domain wherein said enhancer domain enhances the efficiency of said rare- cutting endonuclease when fused to it, thereby obtaining a chimeric rare-cutting endonuclease with enhanced efficiency for a DNA target sequence compared to a corresponding rare-cutting endonuclease lacking said enhancer domain.
According to this aspect of the invention is a chimeric rare-cutting endonuclease for a DNA target sequence comprising:
(i) a rare-cutting endonuclease-derived scaffold;
(ii) an enhancer domain wherein said enhancer domain enhances the efficiency of said rare-cutting endonuclease for said DNA target sequence when fused to it;
(iii) Optionally a peptidic linker.
In an embodiment, said enhancer domain is fused to the N-terminus part of said rare-cutting endonuclease-derived scaffold. In another preferred embodiment, said enhancer domain is fused to the C-terminus part of said rare-cutting endonuclease-derived scaffold. In another embodiment, two enhancer domains are fused to both N-terminus part of said rare-cutting endonuclease-derived scaffold and C-terminus part of said rare-cutting endonuclease-derived scaffold. In another embodiment, said enhancer domain(s) are fused to N-terminus and/or C-terminus of a rare-cutting endonuclease-derived monomer scaffold of a homodimeric endonuclease, resulting in a chimeric rare-cutting endonuclease comprising two or four enhancer domains. In another embodiment, said enhancer domain(s) are fused to N-terminus and/or C-terminus of a rare-cutting endonuclease- derived monomer scaffold of a heterodimeric endonuclease, resulting in a chimeric rare-cutting endonuclease comprising one or two or three or four enhancer domains. In another embodiment, said enhancer domain(s) are fused to N-terminus and/or C-terminus of a rare-cutting endonuclease- derived scaffold from a single-chain endonuclease, resulting in a chimeric rare-cutting endonuclease comprising one or two enhancer domains.
In a preferred embodiment, said enhancer domain is catalytically active, providing functional and/or structural support to said rare-cutting endonuclease-derived scaffold. In a more preferred embodiment, said enhancer domain consists of a protein domain derived from an exonuclease. In another more preferred embodiment, said enhancer domain consists of a protein domain derived
35
from an endonuclease. In a more preferred embodiment, said enhancer domain consists of a protein domain derived from a cleavase. In another more preferred embodiment, said enhancer domain consists of a protein domain derived from a nickase. In a more preferred embodiment, said enhancer domain consists of a protein domain derived from a protein selected from the group consisting of Mmel, Colicin-E7 (CEA7_ECOLX), EndA, Endo I (ENDl_ECOLI), Human Endo G (NUCG_HUMAN), Bovine Endo G (NUCG_BOVIN), R.HinPll, l-Basl, I- Bmol, l-Hmul, l-Tevl, l-Tevll, l-Tevlll, l-Twol, R.Mspl, R.Mval, NucA, NucM, Vvn, Vvn_CLS, Staphylococcal nuclease (NUC_STAAU), Staphylococcal nuclease (NUC_STAHY), Micrococcal nuclease (NUC_SHIFL), Endonuclease yncB, Endodeoxyribonuclease I (ENRN_BPT7), Metnase, Nb.BsrDI, BsrDI A, Nt.BspD6l (R.BspD6l large subunit), ss.BspD6l (R.BspD6l small subunit), R.PIel, Mlyl, Alwl, Mval269l, Bsrl, Bsml, Nb.BtsCI, Nt.BtsCI, Rl.Btsl, R2.Btsl, BbvCI subunit 1, BbvCI subunit 2, BpulOI alpha subunit, BpulOI beta subunit, Bmrl, Bfil, hExol (EX01_HUMAN), Yeast Exol (EX01_YEAST), E.coli Exol, Human TREX2, Mouse TREX1, Human TREX1, Bovine TREX1, Rat TREX1, Human DNA2, Yeast DNA2 (DNA2_YEAST) and VP16, as listed in Table 1 (SEQ ID NO: 10 to SEQ ID NO: 66), a functional mutant, a variant or a derivative of these protein domains thereof. In another more preferred embodiment, said enhancer domain consists of a peptide derived from CFPl peptide (SEQ ID NO: 112). In another more preferred embodiment, any combinations of two protein domains selected from the group consisting of Mmel, Colicin-E7 (CEA7_ECOLX), EndA, Endo I (ENDl_ECOLI), Human Endo G (NUCG_HUMAN), Bovine Endo G (NUCG_BOVIN), R.HinPll, l-Basl, l-Bmol, l-Hmul, I- Tevl, l-Tevll, l-Tevlll, l-Twol, R.Mspl, R.Mval, NucA, NucM, Vvn, Vvn_CLS, Staphylococcal nuclease (NUC_STAAU), Staphylococcal nuclease (NUC_STAHY), Micrococcal nuclease (NUC_SHIFL), Endonuclease yncB, Endodeoxyribonuclease I (ENRN_BPT7), Metnase, Nb.BsrDI, BsrDI A, Nt.BspD6l (R.BspD6l large subunit), ss.BspD6l (R.BspD6l small subunit), R.PIel, Mlyl, Alwl, Mval269l, Bsrl, Bsml, Nb.BtsCI, Nt.BtsCI, Rl.Btsl, R2.Btsl, BbvCI subunit 1, BbvCI subunit 2, BpulOI alpha subunit, BpulOI beta subunit, Bmrl, Bfil, hExol (EX01_HUMAN), Yeast Exol (EX01_YEAST), E.coli Exol, Human TREX2, Mouse TREX1, Human TREX1, Bovine TREX1, Rat TREX1, Human DNA2, Yeast DNA2 (DNA2_YEAST) and VP16, as listed in Table 1 (SEQ ID NO: 10 to SEQ ID NO: 66), a functional mutant, a variant or a derivative of these protein domains thereof, can be fused to both N-terminus part of said rare- cutting endonuclease-derived scaffold and C-terminus part of said rare-cutting endonuclease- derived scaffold, respectively. For example, l-Hmul catalytic domain can be fused to the N-terminus part of said rare-cutting endonuclease-derived scaffold and ColE7 can be fused to the C-terminus part of said rare-cutting endonuclease-derived scaffold. In another more preferred embodiment, said enhancer domain consists of a catalytically active derivative of the protein domains listed above
36
and in Table 1, providing functional and/or structural support to said rare-cutting endonuclease- derived scaffold. In another preferred embodiment, said enhancer domain consists of a catalytically inactive derivative of the protein domains listed above and in Table 1, providing structural support to said rare-cutting endonuclease-derived scaffold.
In a preferred embodiment, said rare-cutting endonuclease-derived scaffold is derived from a meganuclease. In another preferred embodiment, said meganuclease comprises two identical monomers. In another preferred embodiment, said meganuclease comprises two non-identical monomers. In another preferred embodiment, said meganuclease is a single-chain meganuclease. In a more preferred embodiment, said rare-cutting endonuclease-derived scaffold is derived from the group consisting of l-Crel, a functional mutant of l-Crel, a variant of l-Crel or a derivative thereof. In a more preferred embodiment, rare-cutting endonuclease-derived scaffold is a truncated form of wild- type l-Crel (SEQ ID NO: 1). In another more preferred embodiment, rare-cutting endonuclease- derived scaffold comprises first 152, 153, 154 or 155 amino acids residues of wild-type l-Crel (SEQ ID NO: 1). In another more preferred embodiment, rare-cutting endonuclease-derived scaffold comprises residues 2 to 153 of wild-type l-Crel (SEQ ID NO: 1). In another more preferred embodiment, rare-cutting endonuclease-derived scaffold comprises residues 2 to 155 of wild-type I- Crel (SEQ ID NO: 1). In another more preferred embodiment, rare-cutting endonuclease-derived scaffold comprises residues 2 to 153 of wild-type l-Crel (SEQ ID NO: 1) and one or several amino acids substitutions. In another more preferred embodiment, rare-cutting endonuclease-derived scaffold comprises residues 2 to 155 of wild-type l-Crel (SEQ ID NO: 1) and one or several amino acids substitutions. As a non-limiting example, rare-cutting endonuclease-derived scaffold of the present invention comprises residues 2 to 153 or residues 2 to 155 of wild-type l-Crel (SEQ ID NO: 1) and one or two or three or four or five or six or seven or eight or nine or ten further amino acid mutations. As a non-limiting example, rare-cutting endonuclease-derived scaffold of the present invention comprises residues 2 to 153 of wild-type l-Crel (SEQ ID NO: 1) and K82A mutation. In another more preferred embodiment, said rare-cutting endonuclease-derived scaffold comprises a sequence selected from the group consisting of l-Crel_NFSl (SEQ ID NO: 6); l-Crel_NFS2 (SEQ ID NO: 7); and l-Crel_CFSl (SEQ ID NO: 8)..
In a preferred embodiment, said chimeric rare-cutting endonuclease can optionally comprise at least one peptidic linker between said rare-cutting endonuclease-derived scaffold and said at least one enhancer domain. In a more preferred embodiment, said peptidic linker sequence is selected from
37
the group consisting of NFS1, NFS2, CFS1, RM2, BQY, QGPSG, LGPDG KA, la8h_l, ldnpA_l, ld8cA_2, lckqA_3, lsbp_l, lev7A_l, lalo_3, lamf_l, ladjA_3, lfcdC_l, lal3_2, lg3p_l, lacc_3, lahjB_l, lacc_l, laf7_l, lheiA_l, lbia_2, ligtB_l, lnfkA_l, lau7A_l, lbpoB_l, lb0pA_2, lc05A_2, lgcb_l, lbt3A_l, lb3oB_2, 16vpA_6, ldhx_l, lb8aA_land lqu6A_l, as listed in Table 2 (SEQ ID NO: 67 to SEQ ID NO: 104). In a more preferred embodiment, the peptidic linker that can link said enhancer domain to the rare-cutting endonuclease-derived scaffold according to the present invention can be selected from the group consisting of NFS1 (SEQ ID NO: 98), NFS2 (SEQ ID NO: 99) and CFS1 (SEQ ID NO: 100). In the scope of the present invention is also encompassed the case where a peptidic linker is not needed to fuse said enhancer domain to said rare-cutting endonuclease-derived scaffold in order to obtain a chimeric rare-cutting endonuclease according to the present invention.
Enhancement of efficiency of a chimeric rare-cutting endonuclease according to the present invention, compared to a starting rare-cutting endonuclease, can also be a consequence of a fuctional support by said at least one enhancer domain. In a preferred embodiment, said functional support can be the consequence of the hydrolysis of additional phosphodiester bonds. In a more preferred embodiment, said functional support can be the hydrolysis of additional phosphodiester bonds by a protein domain derived from a nuclease. In a more preferred embodiment, said functional support can be the hydrolysis of additional phosphodiester bonds by a protein domain derived from an endonuclease. In a more preferred embodiment, said functional support can be the hydrolysis of additional phosphodiester bonds by a protein domain derived from a cleavase. In another more preferred embodiment, said functional support can be the hydrolysis of additional phosphodiester bonds by a protein domain derived from a nickase. In a more preferred embodiment, said functional support can be the hydrolysis of additional phosphodiester bonds by a protein domain derived from an exonuclease.
Cleavage by rare-cutting endonucleases usually generates cohesive ends, with 3' overhangs for LAGLIDADG meganucleases {Chevalier, 2001 #6} and 5' overhangs for Zinc Finger Nucleases {Smith, 2000 #7}. These ends, which result from hydrolysis of phosphodiester bonds, can be re-ligated in vivo by NHEJ in a seamless way (i.e a scarless re-ligation). The restoration of a cleavable target sequence allows for a new cleavage event by the same endonuclease, and thus, a series of futile cycles of cleavage and re-ligation events can take place (Figure 4A). In a preferred embodiment, enhancement of efficiency of a chimeric rare-cutting endonuclease according to the present invention, compared to a starting rare-cutting endonuclease, can be the hydrolysis of additional
38
phosphodiester bonds at the cleavage site to promote exit from such futile cycles of cleavage and re- ligation events via imprecise NHEJ or Homologous Recombination or Single Strand Annealing (SSA). Said hydrolysis of additional phosphodiester bonds at the cleavage site by said at least one enhancer according to the invention can lead to different types of DSB resection affecting at said DSB cleavage site, one single DNA strand or both DNA strands, affecting either 5' overhangs ends, either 3' overhangs ends, or both ends and depending on the length of said resection. Thus, adding new nickase or cleavase activities to the existing cleavase activity of a rare-cutting endonuclease enhances the efficiency of the resulting chimeric rare-cutting endonuclease according to the invention, at a genomic locus of interest. As a non-limiting example, addition of two nickase activities on opposite strands (as depicted on Figure 4D) or of a new cleavase activity generating a second DSB (Figure 4E) can result in a gap. As a consequence, perfect re-ligation is not possible anymore, and one or several alternative repair outcomes such as imprecise NHEJ, Homologous Recombination or SSA for instance, can be stimulated. As another non-limiting example, the addition of a single nickase activity can result in a single strand gap, and suppress the cohesivity of the ends, which can also enhances the efficiency of the resulting chimeric rare-cutting endonuclease at a genomic locus of interest, according to the invention, via stimulation of one or several alternative repair outcomes mentioned above.
In this third aspect of the invention, chimeric rare-cutting endonucleases according to the invention enhances rare-cutting endonuclease efficiency for a DNA target sequence. Enhancement of efficiency of a rare-cutting endonuclease according to the present invention can be a consequence of a structural support by said at least one enhancer domain. In a preferred embodiment, said structural support enhances the binding of a chimeric rare-cutting endonuclease according to the invention for a DNA target sequence compared to the binding of a starting rare-cutting endonuclease for the same DNA target sequence, as a non-limiting example. In another preferred embodiment, said structural support by at least one enhancer domain enhances the existing catalytical activity of a chimeric rare-cutting endonuclease according to the invention for a DNA target sequence compared to the binding of a starting rare-cutting endonuclease for the same DNA target sequence, as another non-limiting example. In another preferred embodiment, said at least one enhancer domain enhances both the binding and the existing catalytical activity of a chimeric rare-cutting endonuclease according to the invention for a DNA target sequence compared to the binding of a starting rare-cutting endonuclease for the same DNA target sequence, as another non- limiting example. All these non-limiting examples lead to a chimeric rare-cutting endonuclease with
39
enhanced efficiency for a DNA target at a genomic locus of interest, according to the present invention.
Depending on number (one to four) of enhancer domains to the starting rare-cutting endonuclease- derived scaffold according to the invention and on their catalytic activities (functional support) or not (structural support), the resulting chimeric rare-cutting endonuclease can comprise several enzymatic activities that contribute to the enhancement of its efficiency according to the present invention. As a non-limiting example, the addition of one nickase domain to an active rare-cutting endonuclease scaffold derived from a meganuclease results in a chimeric rare-cutting endonuclease with one meganuclease activity and one nickase activity (depicted in figure 4B). As another non- limiting example, the addition of two nickase domains to an active rare-cutting endonuclease scaffold derived from a meganuclease results in a chimeric rare-cutting endonuclease with one meganuclease activity and two nickase activities (as depicted in figures 5C and 5D). As a non-limiting example, the addition of one cleavase domain to an active rare-cutting endonuclease scaffold derived from a meganuclease results in a chimeric rare-cutting endonuclease with one meganuclease activity and one cleavase activity (figure 4E). As another non-limiting example, the addition of two cleavase domains to an active rare-cutting endonuclease scaffold derived from a meganuclease results in a chimeric rare-cutting endonuclease with one meganuclease activity and two cleavase activities.
In this aspect of the present invention, enhancement of efficiency of a rare-cutting endonuclease refers to the increase in the detected level of said efficiency, against a target DNA sequence, of a second rare-cutting endonuclease in comparison to the activity of a first rare-cutting endonuclease against the same target DNA sequence. Said second rare-cutting endonuclease can be an engineered rare-cutting endonuclease, i.e. a chimeric rare-cutting endonuclease according to the invention or not. Said first rare-cutting endonuclease can be a wild-type rare-cutting endonuclease, a starting rare-cutting endonuclease, an engineered rare-cutting endonuclease i.e a chimeric rare-cutting endonuclease according to the invention or not, taken as a reference scaffold for enhancement in efficiency. Additional rounds of enhancement can be envisioned on a starting rare-cutting endonuclease scaffold.
Said enhancement of efficiency can be determined, as non-limiting example, by measuring the level of cleavage-induced recombination generated by said rare-cutting endonuclease or said chimeric rare-cutting endonuclease according to the present invention. Said level of cleavage-induced
40
recombination can be measured by a cell-based recombination assay as described in the International PCT Application WO 2004/067736, as non-limiting example. Importantly, enhancement of efficacy in cells (enhanced generation of targeted mutagenesis or targeted recombination) can be, but is not necessarily associated with an enhancement of the cleavage activity that could be detected in certain in vitro assays. For example, additional phosphodiesterase activities as described in Figure 4 could barely affect the cleavage profile, as detected by in vitro cleavage and separation of the cleavage products on an electrophoresis gel. However, as explained above, and in the legend of Figure 4, the DSB ends generated in this way could be more prone to induce detectable genomic Rearrangements such as targeted mutagenesis (by imprecise NHEJ) or homologous recombination.
Said enhancement in efficiency of said rare-cutting endonuclease is at least a 5% enhancement compared to the starting scaffold, more preferably at least a 10% enhancement, again more preferably at least a 15% enhancement, again more preferably at least a 20% enhancement, again more preferably at least a 25% enhancement, again more preferably a 50% enhancement, again more preferably a enhancement greater than 50%.
In another preferred embodiment, the enhancement of efficiency of a chimeric rare-cutting endonuclease according to the present invention can also be an enhancement in the efficiency for 2NN derivatives of a DNA target sequence by a rare-cutting endonuclease from the LAGLIDADG meganuclease subfamily. By 2NN derivatives of a DNA target sequence is intended variants of a given DNA target sequence having mutations in the four central nucleotides -2 to +2 as numbered in figure 2. For instance, a chimeric rare-cutting endonuclease according to the invention can enhance the efficiency for derivatives of the DNA target sequence C1221 having mutations in the four central nucleotides -2 to +2 as numbered in figure 2, i.e that do not have GTAC sequence at position -2 to +2 as numbered in figure 2. In another preferred embodiment, the chimeric rare-cutting endonuclease according to the invention can allow the processing of 2NN derivatives that are not efficiently processed by a starting rare-cutting endonuclease scaffold from the LAGLIDADG meganuclease subfamily. As a non limiting example, the chimeric rare-cutting endonuclease according to the invention allows the induction of recombination by processing 2NN derivatives that have -2 to +2 target sequences as listed in Table 4 and Table 5, non-originally cleaved by l-Crel.
In this third aspect of the invention said chimeric rare-cutting endonuclease is selected from the group consisting of hHmuCre_D0101 (SEQ ID NO: 9), hHmuCre_D0201 (SEQ ID NO: 105), hHmuCre_D0301 (SEQ ID NO: 106), hHmuCre_D0401 (SEQ ID NO: 107), hHmuCre_D0102 (SEQ ID
41
NO: 108), hHmuCre_D0202 (SEQ ID NO: 109), hHmuCre_D0302 (SEQ ID NO: 110) and hHmuCre_D0402 (SEQ ID NO: 111).
In another aspect, the present invention relates to a method for treatment of a genetic disease caused by a mutation in a specific single double-stranded DNA target sequence in a gene, comprising administering to a subject in need thereof an effective amount of a chimeric rare-cutting endonuclease according to the present invention.
In another preferred embodiment, the present invention relates to a method for inserting a transgene into a specific single double-stranded DNA target sequence of a genomic locus of a cell, tissue or non-human animal wherein at least one chimeric rare-cutting endonuclease of the present invention is introduced in said cell, tissue or non-human animal.
For purposes of therapy, the chimeric rare-cutting endonuclease of the present invention and a pharmaceutically acceptable excipient are administered in a therapeutically effective amount. Such a combination is said to be administered in a "therapeutically effective amount" if the amount administered is physiologically significant. An agent is physiologically significant if its presence results in a detectable change in the physiology of the recipient. In the present context, an agent is physiologically significant if its presence results in a decrease in the severity of one or more symptoms of the targeted disease and in a genome correction of the lesion or abnormality. Vectors comprising targeting DNA and/or nucleic acid encoding a chimeric rare-cutting endonuclease can be introduced into a cell by a variety of methods (e.g., injection, direct uptake, projectile bombardment, liposomes, electroporation). Chimeric rare-cutting endonucleases can be stably or transiently expressed into cells using expression vectors. Techniques of expression in eukaryotic cells are well known to those in the art. (See Current Protocols in Human Genetics: Chapter 12 "Vectors For Gene Therapy" & Chapter 13 "Delivery Systems for Gene Therapy").
In one further aspect of the present invention, the chimeric rare-cutting endonuclease of the present invention is substantially non-immunogenic, i.e., engender little or no adverse immunological response. A variety of methods for ameliorating or eliminating deleterious immunological reactions of this sort can be used in accordance with the invention. In a preferred embodiment, the chimeric rare-cutting endonuclease is substantially free of N-formyl methionine. Another way to avoid unwanted immunological reactions is to conjugate chimeric rare-cutting endonuclease to polyethylene glycol ("PEG") or polypropylene glycol ("PPG") (preferably of 500 to 20,000 daltons average molecular weight (MW)). Conjugation with PEG or PPG, as described by
42
Davis et al. (US 4,179,337) for example, can provide non-immunogenic, physiologically active, water soluble chimeric rare-cutting endonuclease conjugates with anti-viral activity. Similar methods also using a polyethylene-polypropylene glycol copolymer are described in Saifer ef al. (US 5,006,333). The present invention also relates to vectors, compositions and kits used to implement the method. More specifically, the present invention relates to recombinant polynucleotides encoding the chimeric rare-cutting endonucleases of the present invention, specific vectors (polynucleotidic or not) encoding and/or vectorizing them, compositions and/or kits comprising them, all of them beign used or part of a whole to implement methods of the present invention for enhancing rare-cutting endonuclease cleavage activity at a genomic locus of interest in a cell.
43
Definitions
- Amino acid residues in a polypeptide sequence are designated herein according to the one- letter code, in which, for example, Q means Gin or Glutamine residue, means Arg or Arginine residue and D means Asp or Aspartic acid residue.
- Amino acid substitution means the replacement of one amino acid residue with another, for instance the replacement of an Arginine residue with a Glutamine residue in a peptide sequence is an amino acid substitution.
- Efficiency of a rare-cutting endonuclease according to the present invention is the property for said rare-cutting endonuclease of producing a desired event.. This desired event can be for example Homologous gene targeting, targeted mutagenesis, or sequence removal or excision. The efficiency of the desired event depends on several parameters, including the specific activity of the nuclease and the repair pathway resulting in the desired event (efficacy of homologous repair for gene targeting, efficacy and outcome of NHEJ pathways for targeted mutagenesis). Efficiency of a rare cutting endonuclease for a locus is intended to mean its abiliy to produce a desired event at this locus. Efficiency of a rare cutting endonuclease for a target is intended to mean its abiliy to produce a desired event as a consequence of cleavage of this target.
-Enhancement of efficiency of a rare-cutting endonuclease by creating a chimeric rare- cutting endonuclease with enhanced efficiency according to the present invention is measurable comparatively to the efficiency of the starting rare-cutting endonuclease scaffold in the same conditions. Enhancement of efficiency of a chimeric rare-cutting endonuclease can be the consequence of an enhancement of an individual structural parameter or an enhancement of a combination of several individual structural parameters consecutive to the addition of an enhancer domain according to the present invention. Such parameters can be as non-limiting examples, binding affinity or capacity of binding, cleavage activity, turn-over of one enzyme (Kcat), Kcat/Km (incorporating the different rate constants of an enzymatic reaction). Enhancement of efficiency of a chimeric rare-cutting endonuclease can also be a functional consequence to the addition of an enhancer domain according to the present invention. Said additional enhancer domain can provide additional phosphodiester bonds hydrolysis, via cleavase or nickase domains as non-limiting examples, allowing to favour or stimulate one or several alternative repair outcomes. For chimeric rare-cutting endonucleases derived from LAGLIDADG meganuclease subfamily, enhancement of efficiency can also be an enhancement in th eefficiency for 2NN derivatives of DNA target sequence originally targeted by the parent rare-cutting endonuclease. As non-limiting example, an
44
enhancement of the efficiency of chimeric rare-cutting endonucleases derived from LAGLIDADG meganuclease subfamily can be the capacity to process 2NN derivatives that are not cleaved in a detectable way by the parent rare-cutting endonuclease. Also, enhancement of efficiency of a rare- cutting endonuclease according to the present invention can be measured from an application point of view. Chimeric rare-cutting endonucleases according to the present invention being usable for various genome engineering applications, said chimeric rare-cutting endonucleases according to the invention can have an enhanced efficiency for NHEJ, HR, SSA, as non-limiting examples. Said chimeric rare-cutting endonucleases according to the invention can also have an enhanced efficiency for a precise genome engineering such as gene targeting, targeted DNA cleavage or targeted gene repression as non-limiting examples.
- Enhanced/increased/improved cleavage activity or binding, refers to an increase in the detected level of an endonuclease, a rare-cutting endonuclease or a chimeric rare-cutting endonuclease cleavage activity or binding, see below, against a target DNA sequence or DNA target sequence by a second endonuclease, rare-cutting endonuclease or chimeric rare-cutting endonuclease in comparison to the cleavage activity or binding of a first endonuclease or a wild-type endonuclease (non chimeric) or a first rare-cutting endonuclease or a first chimeric rare-cutting endonuclease against said target DNA sequence. The second endonuclease or rare-cutting endonuclease or chimeric rare-cutting endonuclease can be a variant of the first one and can comprise one or more substituted amino acid residues in comparison to the first endonuclease or rare-cutting endonuclease or chimeric rare-cutting endonuclease or additional peptidic sequences or domains. Parameters in this definition are included in the method for enhancing rare-cutting endonuclease efficiency according to the present invention.
- Nucleotides are designated as follows: one-letter code is used for designating the base of a nucleoside: a is adenine, t is thymine, c is cytosine, and g is guanine. For the degenerated nucleotides, r represents g or a (purine nucleotides), k represents g or t, s represents g or c, w represents a or t, m represents a or c, y represents t or c (pyrimidine nucleotides), d represents g, a or t, v represents g, a or c, b represents g, t or c, h represents a, t or c, and n represents g, a, t or c.
- by "meganuclease", is intended a rare-cutting endonuclease subtype having a double- stranded DNA target sequence greater than 12 bp. Said meganuclease is either a dimeric enzyme, wherein each domain is on a monomer or a monomeric enzyme comprising the two domains on a single polypeptide.
- by "meganuclease domain" is intended the region which interacts with one half of the DNA target of a meganuclease and is able to associate with the other domain of the same meganuclease
45
which interacts with the other half of the DNA target to form a functional meganuclease able to cleave said DNA target.
- by "endonuclease variant", "rare-cutting endonuclease variant", chimeric rare-cutting endonuclease variant" or "meganuclease variant" or "variant" it is intended an endonuclease, rare- cutting endonuclease, chimeric rare-cutting endonuclease or meganuclease obtained by replacement of at least one residue in the amino acid sequence of the parent endonuclease, rare- cutting endonuclease, chimeric rare-cutting endonuclease or meganuclease, i.e. the starting endonuclease, rare-cutting endonuclease, chimeric rare-cutting endonuclease or meganuclease (also the starting scaffold) with at least a different amino acid or additional peptidic sequences or domains.
- by "peptide linker", "peptidic linker" or "peptide spacer" it is intended to mean a peptide sequence which allows the connection of different monomers in a fusion protein and the adoption of the correct conformation for said fusion protein activity and which does not alter the specificity of either of the monomers for their targets. Peptide linkers can be of various sizes, from 3 amino acids to 50 amino acids as a non limiting indicative range. Peptide linkers can also be structured or unstructured.
-variants or mutants of the structural elements of the invention including a rare-cutting endonuclease-derived scaffold, an enhancer domain, or a peptidic linker are contemplated.
Generally, these will be at least 80, 90, 95, 98, or 99% identical to a known element, such as those described herein, or have 1, 2, 3, 4, 5, 6, 7, 8, 9, 10 or more amino acid insertions, deletions or substitutions compared to a corresponding element described herein.
- by "related to", particularly in the expression "one cell type related to the chosen cell type or organism", is intended a cell type or an organism sharing characteristics with said chosen cell type or said chosen organism; this cell type or organism related to the chosen cell type or organism, can be derived from said chosen cell type or organism or not.
- by "targeting DNA construct/minimal repair matrix/repair matrix" it is intended to mean a DNA construct comprising a first and second portions that are homologous to regions 5' and 3' of the DNA target in situ. The DNA construct also comprises a third portion positioned between the first and second portion which comprise some homology with the corresponding DNA sequence in situ or alternatively comprise no homology with the regions 5' and 3' of the DNA target in situ. Following cleavage of the DNA target, a homologous recombination event is stimulated between the genome containing the targeted gene comprised in the locus of interest and the repair matrix, wherein the
46
12 032386 genomic sequence containing the DNA target is replaced by the third portion of the repair matrix and a variable part of the first and second portions of the repair matrix.
- by "functional variant" is intended a catalytically active variant of a protein, such variant can have additional properties compared to its parent protein. As a non-limiting example, a functional variant of a meganuclease can be able to cleave additional DNA target sequences that are not cleaved by the parent meganuclease.
- by "derived from" or "derivative(s)" it is intended to mean for instance a meganuclease variant which is created from a parent meganuclease and hence the peptide sequence of the meganuclease variant is related to (primary sequence level) but derived from (mutations) the peptide sequence of the parent meganuclease. In this definition, mutations encompass deletions or insertions of several amino acid residues; as non-limiting example, a truncated variant of an l-Crel meganuclease is considered as a scaffold derived from l-Crel meganuclease. This expression can be applied to an endonuclease, a rare-cutting endonuclease or a chimeric rare-cutting endonuclease.
- by "l-Crel" is intended the wild-type l-Crel having the sequence of pdb accession code lg9y, corresponding to the sequence SEQ ID NO: 1 in the sequence listing. In the present Patent
Application l-Cre\ variants described can comprise an additional Alanine after the first Methionine of the wild type l-Cre\ sequence (SEQ ID NO: 1). These variants may also comprise two additional Alanine residues and an Aspartic Acid residue after the final Proline of the wild type l-Crei sequence as shown in SEQ ID NO: 4. These additional residues do not affect the properties of the enzyme and to avoid confusion these additional residues do not affect the numeration of the residues in l-Cre\ or a variant referred in the present Patent Application, as these references exclusively refer to residues of the wild type l-Cre\ enzyme (SEQ ID NO: 1) as present in the variant, so for instance residue 2 of /- Crel is in fact residue 3 of a variant which comprises an additional Alanine after the first Methionine.
- by "l-Crel site" is intended a 22 to 24 bp double-stranded DNA sequence which is cleaved by l-Crel. l-Crel sites include the wild-type non-palindromic l-Crel homing site and the derived palindromic sequences such as the sequence 5'- t.nCna.joa.ga.ga.yC.eg.st^c.Bg^t. ia+1c+2g+3a+ C+5g+6t+7t+8t+9t+i0g+ 1ia+12 (SEQ ID NO: 2), also called C1221.
- by "domain" or "core domain" is intended the "LAG LIDADG homing endonuclease core domain" which is the characteristic αββαββα fold of the homing endonucleases of the LAGLIDADG family, corresponding to a sequence of about one hundred amino acid residues. Said domain comprises four beta-strands (β^β,ιβ^ folded in an anti-parallel beta-sheet which interacts with one half of the DNA target. This domain is able to associate with another LAGLIDADG homing endonuclease core domain which interacts with the other half of the DNA target to form a functional endonuclease able to cleave said DNA target. For example, in the case of the dimeric homing
47
endonuclease l-Crel (163 amino acids), the LAGLIDADG homing endonuclease core domain corresponds to the residues 6 to 94.
- by "subdomain" is intended the region of a LAGLIDADG homing endonuclease core domain which interacts with a distinct part of a homing endonuclease DNA target half-site.
- by "beta-hairpin" is intended two consecutive beta-strands of the antiparallel beta-sheet of a LAGLIDADG homing endonuclease core domain (βιβ2 0Γ β3 β4 ) which are connected by a loop or a turn,
- by "single-chain meganuclease", "single-chain chimeric meganuclease", "single-chain meganuclease derivative", "single-chain chimeric meganuclease derivative" or "single-chain derivative" is intended a meganuclease comprising two LAGLIDADG homing endonuclease domains or core domains linked by a peptidic linker or peptidic spacer. The single-chain meganuclease is able to cleave a chimeric DNA target sequence comprising one different half of each parent meganuclease target sequence. This definition can be enlarged to "single-chain endonuclease" or "single-chain endonuclease derivative" that qualify two endonuclease domains or core domains from non-LAGLIDADG endonucleases linked by a peptidic linker or peptidic spacer.
- by "DNA target", "DNA target sequence", "target DNA sequence", "target sequence" , "target-site", "target" , "site", "site of interest", "recognition site", "polynucleotide recognition site", "recognition sequence", "homing recognition site", "homing site", "cleavage site" is intended a double-stranded palindromic, partially palindromic (pseudo-palindromic) or non-palindromic polynucleotide sequence that is recognized and cleaved by a LAGLIDADG homing endonuclease such as l-Crel, or a variant, or a single-chain chimeric meganuclease derived from l-Crel. Said DNA target sequence is qualified as "cleavable" by an endonuclease, rare-cutting endonuclease, chimeric rare- cutting endonuclease or meganuclease when recognized within a genomic sequence and known to correspond to the DNA target sequence of a given endonuclease, rare-cutting endonuclease, chimeric rare-cutting endonuclease or meganuclease or a variant of such endonuclease, rare-cutting endonuclease, chimeric rare-cutting endonuclease or meganuclease. These terms refer to a distinct DNA location, preferably a genomic location but also a portion of genetic material that can exist independently to the main body of genetic material such as plasmids, episomes, virus, transposons or in organelles such as mitochondria or chloroplasts as non-limiting examples, at which at least a double-strand break (cleavage) or a single strand break (nick) is to be induced by the meganuclease, endonuclease, rare-cutting endonuclease or chimeric rare-cutting endonuclease. For the LAGLIDADG subfamily of rare-cutting endonucleases, the DNA target is defined by the 5' to 3' sequence of one strand of the double-stranded polynucleotide, as indicate above for C1221 (SEQ ID NO: 2, figure 2). Cleavage of the DNA target occurs at the nucleotides at positions +2 and -2, respectively for the
48
sense and the antisense strand. Unless otherwise indicated, the position at which cleavage of the DNA target by an l-Crel-derived variant occurs, corresponds to the cleavage site on the sense strand of the DNA target. - by "DNA target half-site", "half cleavage site" or half-site" is intended the portion of the DNA target which is bound by each LAGLIDADG homing endonuclease core domain.
- The term "endonuclease" refers to any wild-type or variant enzyme capable of catalyzing the hydrolysis (cleavage) of bonds between nucleic acids within a DNA or NA molecule, preferably a DNA molecule. Endonucleases can be classified as rare-cutting endonucleases when having typically a polynucleotide recognition site greater than 12base pairs (bp) in length, more preferably of 14-45 bp. Rare-cutting endonucleases significantly increase HR by inducing DNA double-strand breaks (DSBs) at a defined locus (Rouet, Smih et al. 1994; Rouet, Smih et al. 1994; Choulika, Perrin et al. 1995; Pingoud and Silva 2007). Rare-cutting endonucleases can for example be a homing endonuclease (Paques and Duchateau 2007), a chimeric Zinc-Finger nuclease (ZFN) resulting from the fusion of engineered zinc-finger domains with the catalytic domain of a restriction enzyme such as Fokl (Porteus and Carroll 2005) or a chemical endonuclease (Eisenschmidt, Lanio et al. 2005 ; Arimondo, Thomas et al. 2006; Simon, Cannata et al. 2008). In chemical endonucleases, a chemical or peptidic cleaver is conjugated either to a polymer of nucleic acids or to another DNA recognizing a specific target sequence, thereby targeting the cleavage activity to a specific sequence. Chemical endonucleases also encompass synthetic nucleases like conjugates of orthophenanthroline, a DNA cleaving molecule, and triplex-forming oligonucleotides (TFOs), known to bind specific DNA sequences (Kalish and Glazer 2005). Such chemical endonucleases are comprised in the term "endonuclease" according to the present invention.
Rare-cutting endonucleases can also be for example TALENs, a new class of chimeric nucleases using a Fokl catalytic domain and a DNA binding domain derived from Transcription Activator Like Effector (TALE), a family of proteins used in the infection process by plant pathogens of the Xanthomonas genus (Boch, Scholze et al. 2009; Moscou and Bogdanove 2009; Christian, Cermak et al. 2010; Li, Huang et al. 2010). The functional layout of a Fokl-based TALE-nuclease (TALEN) is essentially that of a ZFN, with the Zinc-finger DNA binding domain being replaced by the TALE domain. As such, DNA cleavage by a TALEN requires two DNA recognition regions flanking an unspecific central region. Rare-cutting endonucleases encompassed in the present invention can also be derived from TALENs.
Rare-cutting endonuclease can be a homing endonuclease, also known under the name of meganuclease. Such homing endonucleases are well-known to the art (Stoddard 2005). Homing endonucleases recognize a DNA target sequence and generate a single- or double-strand break. Homing endonucleases are highly specific, recognizing DNA target sites ranging from 12 to 45 base pairs (bp) in length, usually ranging from 14 to 40 bp in length. The homing endonuclease according
49
12 032386 to the invention may for example correspond to a LAGLIDADG endonuclease, to a HNH endonuclease, or to a GIY-YIG endonuclease.
In the wild, meganucleases are essentially represented by homing endonucleases. Homing Endonucleases (HEs) are a widespread family of natural meganucleases including hundreds of proteins families (Chevalier and Stoddard 2001). These proteins are encoded by mobile genetic elements which propagate by a process called "homing": the endonuclease cleaves a cognate allele from which the mobile element is absent, thereby stimulating a homologous recombination event that duplicates the mobile DNA into the recipient locus. Given their exceptional cleavage properties in terms of efficacy and specificity, they could represent ideal scaffolds to derive novel, highly specific endonucleases.
HEs belong to five major families. The LAGLIDADG family, named after a conserved peptidic motif involved in the catalytic center, is the most widespread and the best characterized group. Many structures are now available. Whereas most proteins from this family are monomeric and display two LAGLIDADG motifs, a few have only one motif, and thus dimerize to cleave palindromic or pseudo-palindromic target sequences.
Although the LAGLIDADG peptide is the only conserved region among members of the family, these proteins share a very similar architecture. The catalytic core is flanked by two DNA- binding domains with a perfect two-fold symmetry for homodimers such as l-Crel (Chevalier, Monnat et al. 2001), l- sol (Chevalier, Turmel et al. 2003) and l-Ceul (Spiegel, Chevalier et al. 2006) and with a pseudo symmetry for monomers such as l-Scel (Moure, Gimble et al. 2003), l-Dmol (Silva, Dalgaard et al. 1999) or \-Ani\ (Bolduc, Spiegel et al. 2003). Both monomers and both domains (for monomeric proteins) contribute to the catalytic core, organized around divalent cations. Just above the catalytic core, the two LAGLIDADG peptides also play an essential role in the dimerization interface. DNA binding depends on two typical saddle-shaped αββαββα folds, sitting on the DNA major groove. Other domains can be found, for example in inteins such as PI-P/ul (lchiyanagi, Ishino et al. 2000) and Pl-Scel (Moure, Gimble et al. 2002), whose protein splicing domain is also involved in DNA binding.
The making of functional chimeric meganucleases, by fusing the N-terminal l-Dmol domain with an l-Crel monomer (Chevalier, Kortemme et al. 2002; Epinat, Arnould et al. 2003); International PCT Application WO 03/078619 (Cellectis) and WO 2004/031346 (Fred Hutchinson Cancer Research Center, Stoddard et al)) have demonstrated the plasticity of LAGLIDADG proteins.
Different groups have also used a semi-rational approach to locally alter the specificity of the l-Crel (Seligman, Stephens et al. 1997; Sussman, Chadsey et al. 2004); International PCT Applications WO 2006/097784, WO 2006/097853, WO 2007/060495 and WO 2007/049156 (Cellectis); (Arnould,
50
12 032386
Chames et al. 2006; Rosen, Morrison et al. 2006; Smith, Grizot et al. 2006), l-5cel (Doyon, Pattanayak et al. 2006), Pl-Scel (Gimble, Moure et al. 2003) and l-Msol (Ashworth, Havranek et al. 2006).
In addition, hundreds of l-Crel derivatives with locally altered specificity were engineered by combining the semi-rational approach and High Throughput Screening:
- Residues Q44, R68 and R70 or Q44, R68, D75 and 177 of l-Crel were mutagenized and a collection of variants with altered specificity at positions ± 3 to 5 of the DNA target (5NNN DNA target) were identified by screening (International PCT Applications WO 2006/097784 and WO 2006/097853 (Cellectis); (Arnould, Chames et al. 2006; Smith, Grizot et al. 2006).
- Residues K28, N30 and Q38 or N30, Y33 and Q38 or K28, Y33, Q38 and S40 of l-Crel were mutagenized and a collection of variants with altered specificity at positions ± 8 to 10 of the DNA target (10NNN DNA target) were identified by screening (Arnould, Chames et al. 2006; Smith, Grizot et al. 2006); International PCT Applications WO 2007/060495 and WO 2007/049156 (Cellectis)).
Two different variants were combined and assembled in a functional heterodimeric endonuclease able to cleave a chimeric target resulting from the fusion of two different halves of each variant DNA target sequence ((Arnould, Chames et al. 2006; Smith, Grizot et al. 2006); International PCT Applications WO 2006/097854 and WO 2007/034262).
Furthermore, residues 28 to 40 and 44 to 77 of l-Crel were shown to form two partially separable functional subdomains, able to bind distinct parts of a homing endonuclease target half- site (Smith, Grizot et al. 2006); International PCT Applications WO 2007/049095 and WO 2007/057781 (Cellectis)).
The combination of mutations from the two subdomains of l-Crel within the same monomer allowed the design of novel chimeric molecules (homodimers) able to cleave a palindromic combined DNA target sequence comprising the nucleotides at positions ± 3 to 5 and ± 8 to 10 which are bound by each subdomain ((Smith, Grizot et al. 2006); International PCT Applications WO 2007/049095 and WO 2007/057781 (Cellectis)).
The method for producing meganuclease variants and the assays based on cleavage-induced recombination in mammal or yeast cells, which are used for screening variants with altered specificity are described in the International PCT Application WO 2004/067736; (Epinat, Arnould et al. 2003; Chames, Epinat et al. 2005; Arnould, Chames et al. 2006). These assays result in a functional LacZ reporter gene which can be monitored by standard methods.
The combination of the two former steps allows a larger combinatorial approach, involving four different subdomains. The different subdomains can be modified separately and combined to obtain an entirely redesigned meganuclease variant (heterodimer or single-chain molecule) with chosen specificity. In a first step, couples of novel meganucleases are combined in new molecules
51
("half-meganucleases") cleaving palindromic targets derived from the target one wants to cleave. Then, the combination of such "half-meganucleases" can result in a heterodimeric species cleaving the target of interest. The assembly of four sets of mutations into heterodimeric endonucleases cleaving a model target sequence or a sequence from different genes has been described in the following Cellectis International patent applications: XPC gene (WO2007/093918), RAG gene (WO2008/010093), HPRT gene (WO2008/059382), beta-2 microglobulin gene (WO2008/102274), Rosa26 gene (WO2008/152523), Human hemoglobin beta gene (WO2009/13622) and Human interleukin-2 receptor gamma chain gene (WO2009019614).
These variants can be used to cleave genuine chromosomal sequences and have paved the way for novel perspectives in several fields, including gene therapy.
Examples of such endonuclease include l-Sce I, l-Chu I, l-Cre I, l-Csm I, Pl-Sce I, Pl-Tli I, Pl-Mtu I, l-Ceu I, l-Sce II, l-Sce III, HO, Pl-Civ I, Pl-Ctr I, Pl-Aae I, Pl-Bsu I, Pl-Dha I, Pl-Dra I, Pl-Mav I, Pl-Mch I, Pl-Mfu I, Pl-Mfl I, Pl-Mga I, Pl-Mgo I, Pl-Min I, Pl-Mka I, Pl-Mle I, Pl-Mma I, Pl-Msh I, Pl-Msm I, Pl-Mth I, Pl-Mtu I, Pl-Mxe I, Pl-Npu I, Pl-Pfu I, Pl-Rma I, Pl-Spb I, Pl-Ssp I, Pl-Fac I, Pl-Mja I, Pl-Pho I, Pi-Tag I, Pl-Thy I, Pl-Tko I, Pl-Tsp I, l-Msol.
A homing endonuclease can be a LAGLIDADG endonuclease such as \-Scel, \-Crel, \-Ceul, I- Msol, and \-Dmol.
Said LAGLIDADG endonuclease can be l-Sce I, a member of the family that contains two LAGLIDADG motifs and functions as a monomer, its molecular mass being approximately twice the mass of other family members like l-Crel which contains only one LAGLIDADG motif and functions as homodimers.
Meganucleases, endonucleases or rare-cutting endonucleases mentioned in the present application encompass both wild-type (naturally-occurring) and variant meganucleases, endonucleases or rare-cutting endonucleases. Meganucleases, endonucleases or rare-cutting endonucleases according to the invention can be a "variant" meganuclease, endonuclease or rare- cutting endonuclease, i.e. a meganuclease, endonuclease or rare-cutting endonuclease that does not naturally exist in nature and that is obtained by genetic engineering or by random mutagenesis, i.e. an engineered meganuclease, endonuclease or rare-cutting endonuclease. This variant meganuclease, endonuclease or rare-cutting endonuclease can for example be obtained by substitution of at least one residue in the amino acid sequence of a wild-type, naturally-occurring, meganuclease, endonuclease or rare-cutting endonuclease with a different amino acid. Said substitution(s) can for example be introduced by site-directed mutagenesis and/or by random mutagenesis. In the frame of the present invention, such variant meganucleases, endonucleases or rare-cutting endonucleases remain functional, i.e. they retain the capacity of recognizing (binding
52
function) and optionally specifically cleaving a target sequence to initiate a gene engineering process. Meganuclease part, endonuclease part or rare-cutting endonuclease part in a chimeric rare- cutting endonuclease according to the present invention can only retain the capacity of recognizing a target sequence, in particular case, the capacity of cleaving a target sequence being provided by an enhancer domain according to the present invention.
The variant meganuclease, endonuclease or rare-cutting endonuclease according to the invention cleaves a target sequence that is different from the target sequence of the corresponding wild-type meganuclease, endonuclease or rare-cutting endonuclease. Methods for obtaining such variant endonucleases with novel specificities are well-known in the art.
Endonucleases variants may be homodimers (meganuclease comprising two identical monomers) or heterodimers (meganuclease comprising two non-identical monomers). It is understood that the scope of the present invention also encompasses endonuclease variants per se, including heterodimers (WO2006097854), obligate heterodimers (WO2008093249) and single chain meganucleases (WO03078619 and WO2009095793) as non limiting examples, able to cleave one target of interest in a polynucleotide sequence or in a genome and derivatives of these variants according to the present invention, such as chimeric rare-cutting endonucleases derived from these variants. The invention also encompasses hybrid variant per se composed of two monomers from different origins (WO03078619).
Meganucleases, endonucleases or rare-cutting endonucleases with novel specificities can be used in the method according to the present invention for gene targeting and thereby integrating a transgene of interest into a genome at a predetermined location.
- by "parent meganuclease" it is intended to mean a wild type meganuclease or a variant of such a wild type meganuclease with identical properties or alternatively a meganuclease with some altered characteristic in comparison to a wild type version of the same meganuclease. This expression can be applied to an endonuclease, a rare-cutting endonuclease or a chimeric rare- cutting endonuclease.
- By " delivery vector" or " delivery vectors" is intended any delivery vector which can be used in the present invention to put into cell contact ( i.e "contacting") or deliver inside cells or subcellular compartments agents/chemicals and molecules (proteins or nucleic acids) needed in the present invention, such as meganucleases, endonucleases, rare-cutting endonucleases or chimeric rare-cutting endonucleases or plasmidic vectors encoding said meganucleases, endonucleases, rare-cutting endonucleases or chimeric rare-cutting endonucleases, or repair matrix per se or encoded in a plasmidic vector for a gene engineering application for instance. It includes, but is not limited to liposomal delivery vectors, viral delivery vectors, drug
delivery vectors, chemical carriers, polymeric carriers, lipoplexes, polyplexes, dendrimers, microbubbles (ultrasound contrast agents), nanoparticles, emulsions or other appropriate transfer vectors. These delivery vectors allow delivery of molecules, chemicals, macromolecules (genes, proteins), or other vectors such as plasmids, peptides developed by Diatos. In these cases, delivery vectors are molecule carriers. By "delivery vector" or "delivery vectors" is also intended delivery methods to perform transfection.
- The terms "vector" or "vectors" refer to a nucleic acid molecule capable of transporting another nucleic acid to which it has been linked. A "vector" in the present invention includes, but is not limited to, a viral vector, a plasmid (i.e a plasmidic vector), a RNA vector or a linear or circular DNA or RNA molecule which may consists of a chromosomal, non chromosomal, semi-synthetic or synthetic nucleic acids. Preferred vectors are those capable of autonomous replication (episomal vector) and/or expression of nucleic acids to which they are linked (expression vectors). Large numbers of suitable vectors are known to those of skill in the art and commercially available.
Viral vectors include retrovirus, adenovirus, parvovirus (e. g. adenoassociated viruses), coronavirus, negative strand RNA viruses such as orthomyxovirus (e. g., influenza virus), rhabdovirus (e. g., rabies and vesicular stomatitis virus), paramyxovirus (e. g. measles and Sendai), positive strand RNA viruses such as picornavirus and alphavirus, and double-stranded DNA viruses including adenovirus, herpesvirus (e. g., Herpes Simplex virus types 1 and 2, Epstein-Barr virus, cytomegalovirus), and poxvirus (e. g., vaccinia, fowlpox and canarypox). Other viruses include Norwalk virus, togavirus, flavivirus, reoviruses, papovavirus, hepadnavirus, and hepatitis virus, for example. Examples of retroviruses include: avian leukosis-sarcoma, mammalian C-type, B-type viruses, D type viruses, HTLV-BLV group, lentivirus, spumavirus (Coffin, J. M., Retroviridae: The viruses and their replication, In Fundamental Virology, Third Edition, B. N. Fields, et al., Eds., Lippincott-Raven Publishers, Philadelphia, 1996).
- By "lentiviral vector" is meant HIV-Based lentiviral vectors that are very promising for gene delivery because of their relatively large packaging capacity, reduced immunogenicity and their ability to stably transduce with high efficiency a large range of different cell types. Lentiviral vectors are usually generated following transient transfection of three (packaging, envelope and transfer) or more plasmids into producer cells. Like HIV, lentiviral vectors enter the target cell through the interaction of viral surface glycoproteins with receptors on the cell surface. On entry, the viral RNA undergoes reverse transcription, which is mediated by the viral reverse transcriptase complex. The product of reverse transcription is a double-stranded linear viral DNA, which is the substrate for viral integration in the DNA of infected cells.
54
2012/032386
By "integrative lentiviral vectors (or LV)", is meant such vectors as non limiting example, that are able to integrate the genome of a target cell.
At the opposite by "non integrative lentiviral vectors (or NILV)" is meant efficient gene delivery vectors that do not integrate the genome of a target cell through the action of the virus integrase.
One type of preferred vector is an episome, i.e., a nucleic acid capable of extra-chromosomal replication. Preferred vectors are those capable of autonomous replication and/or expression of nucleic acids to which they are linked. Vectors capable of directing the expression of genes to which they are operatively linked are referred to herein as "expression vectors. A vector according to the present invention comprises, but is not limited to, a YAC (yeast artificial chromosome), a BAC (bacterial artificial), a baculovirus vector, a phage, a phagemid, a cosmid, a viral vector, a plasmid, a NA vector or a linear or circular DNA or RNA molecule which may consist of chromosomal, non chromosomal, semi-synthetic or synthetic DNA. In general, expression vectors of utility in recombinant DNA techniques are often in the form of "plasmids" which refer generally to circular double stranded DNA loops which, in their vector form are not bound to the chromosome. Large numbers of suitable vectors are known to those of skill in the art. Vectors can comprise selectable markers, for example: neomycin phosphotransferase, histidinol dehydrogenase, dihydrofolate reductase, hygromycin phosphotransferase, herpes simplex virus thymidine kinase, adenosine deaminase, glutamine synthetase, and hypoxanthine-guanine phosphoribosyl transferase for eukaryotic cell culture; TRP1 for 5. cerevisiae; tetracyclin, rifampicin or ampicillin resistance in E. coli. Preferably said vectors are expression vectors, wherein a sequence encoding a polypeptide of interest is placed under control of appropriate transcriptional and translational control elements to permit production or synthesis of said polypeptide. Therefore, said polynucleotide is comprised in an expression cassette. More particularly, the vector comprises a replication origin, a promoter operatively linked to said encoding polynucleotide, a ribosome binding site, a RNA-splicing site (when genomic DNA is used), a polyadenylation site and a transcription termination site. It also can comprise an enhancer or silencer elements. Selection of the promoter will depend upon the cell in which the polypeptide is expressed. Suitable promoters include tissue specific and/or inducible promoters. Examples of inducible promoters are: eukaryotic metallothionine promoter which is induced by increased levels of heavy metals, prokaryotic lacZ promoter which is induced in response to isopropyl-p-D-thiogalacto-pyranoside (IPTG) and eukaryotic heat shock promoter which is induced by increased temperature. Examples of tissue specific promoters are skeletal muscle creatine kinase, prostate-specific antigen (PSA), ct-antitrypsin protease, human surfactant (SP) A and B proteins, β- casein and acidic whey protein genes.
55
Inducible promoters may be induced by pathogens or stress, more preferably by stress like cold, heat, UV light, or high ionic concentrations (reviewed in Potenza C et al. 2004, In vitro Cell Dev Biol 40:1-22). Inducible promoter may be induced by chemicals (reviewed in (Moore, Samalova et al. 2006); (Padidam 2003); (Wang, Zhou et al. 2003); (Zuo and Chua 2000).
Delivery vectors and vectors can be associated or combined with any cellular permeabilization techniques such as sonoporation or electroporation or derivatives of these techniques.
By cell or cells is intended any prokaryotic or eukaryotic living cells, cell lines derived from these organisms for in vitro cultures, primary cells from animal or plant origin.
- By "primary cell" or "primary cells" are intended cells taken directly from living tissue (i.e. biopsy material) and established for growth in vitro, that have undergone very few population doublings and are therefore more representative of the main functional components and characteristics of tissues from which they are derived from, in comparison to continuous tumorigenic or artificially immortalized cell lines. These cells thus represent a more valuable model to the in vivo state they refer to.
In the frame of the present invention, "eukaryotic cells" refer to a fungal, plant or animal cell or a cell line derived from the organisms listed below and established for in vitro culture. More preferably, the fungus is of the genus Aspergillus, Penicillium, Acremonium, Trichoderma, Chrysoporium, Mortierella, Kluyveromyces or Pichia; More preferably, the fungus is of the species Aspergillus niger, Aspergillus nidulans, Aspergillus oryzae, Aspergillus terreus, Penicillium chrysogenum, Penicillium citrinum, Acremonium Chrysogenum, Trichoderma reesei, Mortierella alpine, Chrysosporium lucknowense, Kluyveromyces lactis, Pichia pastoris or Pichia ciferrii.
More preferably the plant is of the genus Arabidospis, Nicotiana, Solanum, lactuca, Brassica, Oryza, Asparagus, Pisum, Medicago, Zea, Hordeum, Secale, Triticum, Capsicum, Cucumis, Cucurbita, Citrullis, Citrus, Sorghum; More preferably, the plant is of the species Arabidospis thaliana, Nicotiana tabaccum, Solanum lycopersicum, Solanum tuberosum, Solanum melongena, Solanum esculentum, Lactuca saliva, Brassica napus, Brassica oleracea, Brassica rapa, Oryza glaberrima, Oryza sativa, Asparagus officinalis, Pisum sativum, Medicago sativa, zea mays, Hordeum vulgare, Secale cereal, Triticum aestivum, Triticum durum, Capsicum sativus, Cucurbita pepo, Citrullus lanatus, Cucumis melo, Citrus aurantifolia, Citrus maxima, Citrus medica, Citrus reticulata.
More preferably the animal cell is of the genus Homo, Rattus, Mus, Sus, Bos, Danio, Canis, Felis, Equus, Salmo, Oncorhynchus, Gallus, Meleagris, Drosophila, Caenorhabditis; more preferably, the animal cell is of the species Homo sapiens, Rattus norvegicus, Mus musculus, Sus scrofa, Bos
56
taurus, Danio rerio, Canis lupus, Felis catus, Equus caballus, Salmo salar, Oncorhynchus mykiss, Gallus gallus, Meleagris gallopavo, Drosophila melanogaster, Caenorhabditis elegans.
In the present invention, the cell can be a plant cell, a mammalian cell, a fish cell, an insect cell or cell lines derived from these organisms for in vitro cultures or primary cells taken directly from living tissue and established for in vitro culture. As non limiting examples cell lines can be selected from the group consisting of CHO-K1 cells; HEK293 cells; Caco2 cells; U2-OS cells; NIH 3T3 cells; NSO cells; SP2 cells; CHO-S cells; DG44 cells; K-562 cells, U-937 cells; M C5 cells; IMR90 cells; Jurkat cells; HepG2 cells; HeLa cells; HT-1080 cells; HCT-116 cells; Hu-h7 cells; Huvec cells; Molt 4 cells.
All these cell lines can be modified by the method of the present invention to provide cell line models to produce, express, quantify, detect, study a gene or a protein of interest; these models can also be used to screen biologically active molecules of interest in research and production and various fields such as chemical, biofuels, therapeutics and agronomy as non-limiting examples. Adoptive immunotherapy using genetically engineered T cells is a promising approach for the treatment of malignancies and infectious diseases. Most current approaches rely on gene transfer by random integration of an appropriate T Cell Receptor (TCR) or Chimeric Antigen Receptor (CAR). Targeted approach using rare-cutting endonucleases such as meganucleases is an efficient and safe alternative method to transfer genes into T cells and generate genetically engineered T cells.
- by "homologous" is intended a sequence with enough identity to another one to lead to homologous recombination between sequences, more particularly having at least 95 % identity, preferably 97 % identity and more preferably 99 %.
- "identity" refers to sequence identity between two nucleic acid molecules or polypeptides. Identity can be determined by comparing a position in each sequence which may be aligned for purposes of comparison. When a position in the compared sequence is occupied by the same base, then the molecules are identical at that position. A degree of similarity or identity between nucleic acid or amino acid sequences is a function of the number of identical or matching nucleotides at positions shared by the nucleic acid sequences. Various alignment algorithms and/or programs may be used to calculate the identity between two sequences, including FASTA, or BLAST which are available as a part of the GCG sequence analysis package (University of Wisconsin, Madison, Wis.), and can be used with, e.g., default setting.
- by "mutation" is intended the substitution, deletion, insertion of one or more nucleotides/amino acids in a polynucleotide (cDNA, gene) or a polypeptide sequence. Said mutation can affect the coding sequence of a gene or its regulatory sequence. It may also affect the structure of the genomic sequence or the structure/stability of the encoded mRNA.
57
- In the frame of the present invention, the expressions "double-strand break-induced mutagenesis" (DSB-induced mutagenesis) and "targeted mutagenesis" refer to a mutagenesis event consecutive to an NHEJ event following an endonuclease-induced DSB, leading to insertion/deletion at the cleavage site of an endonuclease
- By "gene" is meant the basic unit of heredity, consisting of a segment of DNA arranged in a linear manner along a chromosome, which codes for a specific protein or segment of protein. A gene typically includes a promoter, a 5' untranslated region, one or more coding sequences (exons), optionally introns, a 3' untranslated region. The gene may further comprise a terminator, enhancers and/or silencers.
- As used herein, the term "transgene" refers to a sequence encoding a polypeptide.
Preferably, the polypeptide encoded by the transgene is either not expressed, or expressed but not biologically active, in the cell, tissue or individual in which the transgene is inserted. Most preferably, the transgene encodes a therapeutic polypeptide useful for the treatment of an individual.
- The term "gene of interest" or "GOI" refers to any nucleotide sequence encoding a known or putative gene product.
- As used herein, the term "locus" is the specific physical location of a DNA sequence (e.g. of a gene) on a chromosome. The term "locus" usually refers to the specific physical location of an endonuclease's target sequence on a chromosome. Such a locus, which comprises a target sequence that is recognized and cleaved by an endonuclease according to the invention, is referred to as "locus according to the invention". Also, the expression "genomic locus of interest" is used to qualify a nucleic acid sequence in a genome that can be a putative target for a double-strand break according to the invention. By "endogenous genomic locus of interest" is intended a native nucleic acid sequence in a genome, i.e. a sequence or allelic variations of this sequence that is naturally present at this genomic locus. It is understood that the considered genomic locus of interest of the present invention can not only qualify a nucleic acid sequence that exists in the main body of genetic material (i.e. in a chromosome) of a cell but also a portion of genetic material that can exist independently to said main body of genetic material such as plasmids, episomes, virus, transposons or in organelles such as mitochondria or chloroplasts as non-limiting examples.
- By the expression "loss of genetic information" is understood the elimination or addition of at least one given DNA fragment (at least one nucleotide) or sequence, bordering the recognition sites of the endonucleases, rare-cutting endonucleases or chimeric rare-cutting endonucleases of the present invention and leading to a change of the original sequence around said endonuclease-, rare-cutting endonuclease- or chimeric rare-cutting endonuclease-cutting sites, within the genomic locus of interest. This loss of genetic information can be, as a non-limiting example, the elimination
58
of an intervening sequence between two endonuclease-, rare-cutting endonuclease- or chimeric rare-cutting endonuclease-cutting sites. As another non-limiting example, this loss of genetic information can also be an excision of a single-strand of DNA spanning the starting rare-cutting endonuclease scaffold DNA binding region of a chimeric rare-cutting endonuclease according to the present invention when both one cleavase enhancer domain and one nickase enhancer domain, respectively, are part of said chimeric rare-cutting endonuclease. In this case, the starting rare- cutting endonuclease scaffold DNA binding region can be catalytically active or not and the "loss of genetic information" corresponds to the elimination of an intervening DNA fragment between both cleavase and nickase cleavage sites of said chimeric rare-cutting endonuclease.
- By the expression "two nearby DNA double strand breaks" within the genomic locus of interest, is meant two endonucleases cutting sites distant at between 12 bp and 1000 bp.
- By "scarless re-ligation" is intended the perfect re-ligation event, without loss of genetic information (no insertion/deletion events) of the DNA broken ends through NHEJ process after the creation of a double-strand break event.
- By "fusion protein" is intended the result of a well-known process in the art consisting in the joining or merging of two or more genes which originally encode for separate proteins or part of them, the translation of said "fusion gene" resulting in a single polypeptide, said "fusion protein" with functional properties derived from each of the original proteins.
- By "Imprecise NHEJ" is intended the re-ligation of nucleic acid ends generated by a DSB, with insertions or deletions of nucleotides. Imprecise NHEJ is an outcome and not a repair pathway and can result from different NHEJ pathways (Ku dependent or Ku independent as non-limiting examples).
- By "chimeric rare-cutting endonuclease" is meant any fusion protein comprising a rare- cutting endonuclease. Said rare-cutting endonuclease might be at the N- terminus part of said chimeric rare-cutting endonuclease; at the opposite, said rare-cutting endonuclease might be at the C- terminus part of said chimeric rare-cutting endonuclease. A "chimeric rare-cutting endonuclease" according to the present invention which comprises two catalytic domains can be described as "bi- functional" or as "bi-functional meganuclease". A "chimeric rare-cutting endonuclease" according to the present invention which comprises more than two catalytic domains can be described as "multi- functional" or as "multi-functional meganuclease". As non-limiting examples, chimeric rare-cutting endonucleases according to the present invention can be a fusion protein between a rare-cutting endonuclease and one catalytic domain; chimeric rare-cutting endonucleases according to the present invention can also be a fusion protein between a rare-cutting endonuclease and two catalytic domains. As mentioned previously, the rare-cutting endonuclease part of chimeric rare-
59
cutting endonucleases according to the present invention can be a meganuclease comprising two identical monomers, two non-identical monomers, or a single chain meganuclease. The rare-cutting endonuclease part of chimeric rare-cutting endonucleases according to the present invention can also be the DNA-binding domain of an inactive rare-cutting endonuclease. In other non-limiting examples, chimeric rare-cutting endonucleases according to the present invention can be derived from a TALE-nuclease (TALEN), i. e. a fusion between a DNA-binding domain derived from a Transcription Activator Like Effector (TALE) and one or two catalytic domains. In this definition are encompassed mutants, variants or derivatives of meganucleases (including monomers and single chain meganucleases), endonucleases, rare-cutting endonucleases or chimeric rare-cutting endonucleases.
- By "enhancer domain(s)" or "enhancer(s)" are meant protein domains that provide functional and/or structural support to a protein scaffold, a rare-cutting endonuclease-derived scaffold as a non-limiting example, therefore allowing an enhancement in cleavage efficiency of the resulting fusion protein, i.e a chimeric rare-cutting endonuclease, relative to the cleavage efficiency of the wild-type rare-cutting endonuclease or starting rare-endonuclease scaffold. A particular domain is an enhancer domain when it provides at least a 5% enhancement in efficiency of the starting scaffold, more preferably 10 %, again more preferably 15 %, again more preferably 20 %, again more preferably 25 %, again more preferably 50%, again more preferably greater than 50%. Non-limiting examples of such enhancer domains are given in Table 1. Enhancer domain(s) according to the present invention can be fused to N-terminus and/or C-terminus of a rare-cutting endonuclease-derived monomer scaffold of a homodimeric endonuclease, resulting in a chimeric rare-cutting endonuclease comprising two or four enhancer domains. Enhancer domain(s) according to the present invention can also be fused to N-terminus and/or C-terminus of a rare-cutting endonuclease-derived monomer scaffold of a heterodimeric endonuclease, resulting in a chimeric rare-cutting endonuclease comprising one or two or three or four enhancer domains. Enhancer domain(s) according to the present invention can also be fused to N-terminus and/or C-terminus of a rare-cutting endonuclease-derived scaffold of a single-chain endonuclease, resulting in a chimeric rare-cutting endonuclease comprising one or two enhancer domains. - By "catalytic domain" is intended the protein domain or module of an enzyme containing the active site of said enzyme; by active site is intended the part of said enzyme at which catalysis of the substrate occurs. Enzymes, but also their catalytic domains, are classified and named according to the reaction they catalyze. The Enzyme Commission number (EC number) is a numerical classification scheme for enzymes, based on the chemical reactions they catalyze
60
(http://wwwxhem.qmul.ac.uk/iubmb/enzyme/). In the scope of the present invention, any catalytic domain can be fused to a rare-cutting endonuclease scaffold or starting rare-endonuclease scaffold or starting scaffold to generate a chimeric rare-cutting endonuclease. In a preferred embodiment of the present invention, said catalytic domain can be an enhancer domain according to the present invention. If catalytically active, said enhancer domain can provide functional and/or structural support to rare-cutting endonuclease scaffold when fused to it. If catalytically inactive, said enhancer domain provides structural support to rare-cutting endonuclease scaffold when fused to it.
- By "nuclease catalytic domain" is intended the protein domain comprising the active site of an endonuclease or an exonuclease enzyme. Such nuclease catalytic domain can be, for instance, a "cleavase domain" or a "nickase domain". By "cleavase domain" is intended a protein domain whose catalytic activity generates a Double Strand Break (DSB) in a DNA target. By "nickase domain" is intended a protein domain whose catalytic activity generates a single strand break in a DNA target sequence. Non-limiting examples of such catalytic domains are given in Table 1 with a GenBank or NCBI or UniProtKB/Swiss-Prot number as a reference.
The above written description of the invention provides a manner and process of making and using it such that any person skilled in this art is enabled to make and use the same, this enablement being provided in particular for the subject matter of the appended claims, which make up a part of the original description.
As used above, the phrases "selected from the group consisting of," "chosen from," and the like include mixtures of the specified materials.
Where a numerical limit or range is stated herein, the endpoints are included. Also, all values and subranges within a numerical limit or range are specifically included as if explicitly written out.
The above description is presented to enable a person skilled in the art to make and use the invention, and is provided in the context of a particular application and its requirements. Various modifications to the preferred embodiments will be readily apparent to those skilled in the art, and the generic principles defined herein may be applied to other embodiments and applications without departing from the spirit and scope of the invention. Thus, this invention is not intended to be limited to the embodiments shown, but is to be accorded the widest scope consistent with the principles and features disclosed herein.
Having generally described this invention, a further understanding can be obtained by reference to certain specific examples, which are provided herein for purposes of illustration only, and are not intended to be limiting unless otherwise specified.
61
2386
Examples Example 1
Protein-fusion scaffolds were designed based on a truncated form of wild-type l-Crel (SEQ ID NO: 1), l-Crel_X (SEQ ID NO: 5) and three different linker polypeptides [NFSl (SEQ ID NO: 98), NFS2 (SEQ ID NO: 99) and CFS1 (SEQ ID NO: 100)] fused to either the N- or C-terminus of the protein. Structure models were generated in all cases, with the goal of designing a "baseline" fusion linker that would traverse the l-Crel parent scaffold surface with little to no effect on its DNA binding or cleavage activities. For the two N-terminal fusion scaffolds, the polypeptide spanning residues 2 to 153 of I- Crel was used, with a K82A mutation to allow for linker placement. The C-terminal fusion scaffold contains residues 2 to 155 of wild-type l-Crel. For both fusion scaffold types, the "free" end of the linker (i.e. onto which a polypeptide can be linked) is designed to be proximal to the DNA, as determined from models built using the l-Crel/DNA complex structures as a starting point (PDB id: lg9z). The two l-Crel N-terminal fusion scaffolds (l-Crel_NFSl = SEQ ID NO: 6 and l-Crel_NFS2 = SEQ ID NO: 7) and the single C-terminal fusion scaffold (l-Crel_CFSl = SEQ ID NO: 8) were tested in our yeast assay (previously described in International PCT Applications WO 2004/067736 and in (Epinat, Arnould et al. 2003; Chames, Epinat et al. 2005; Arnould, Chames et al. 2006; Smith, Grizot et al. 2006) and found to have activity similar to that of wild-type l-Crel.
Wild-type l-Hmul functions as a monomeric nickase to generate a strand-specific nick in its target DNA [REF]. Guided by biochemical and structural data, variable length constructs were designed from the N-terminal region of l-Hmul that encompass the entire catalytic domain. Fragments were fused to the N-terminus of either l-Crel_NFSl or l-Crel_NFS2 to generate 8 constructs (SEQ ID NO: 9 and SEQ ID NO: 105-111). As l-Crel is a homodimer, all fusion constructs contain two HNH domains (Figure 3A, where "enhancer domain" = the l-Hmul catalytic domain).
The activity of each "enhanced" meganuclease (eMega) was assessed using yeast assay described in International PCT Applications WO 2004/067736. To monitor the effect on activity resulting from the addition of the new enhancer domains, all constructs were screened against libraries of the C1221 target DNA (Figure 2) having regions mutated in the following positions (numbering as in Figure 2): 12NN, lONNNnn, 7NNNnn, 5NNN, 2NN. These targets contain nucleotide substitutions at the +/- positions indicated. In all cases, fusion proteins strongly enhanced cleavage of targets relative to the
62
activity of wild-type l-Crel (Table 3). In addition, the hHmuCre fusion hybrids were able to cleave "2NN" targets not cleaved by wild-type l-Crel at either 37°C (Table 4) or 30°C (Table 5).
Table 3: Activity in Yeast assay for l-Hmul/l-Crel fusions. The activity of the hHmuCre fusion protein relative to that of wild-type l-Crel is shown for various target libraries (see Example 3 text for details). Activity enhancement is based on a direct comparison of the change in activity for targets cleavable by the hHmuCre hybrid. The total number of targets in a given library is indicated next to the library name. The percent of targets enhanced to a given threshold are listed in each column. Assays were performed at two temperatures, 30°C and 37°C. Numbers in parentheses indicate actual target number enhanced versus total targets cleaved at the given temperature.
Table 4: Additional 2NN targets cleaved by hHmuCre at 37°C. From the 136 possible 2NN targets, hHmuCre is able to cleave a total of 125 (see Table 3) at 37°C. Of these, 13 DNA sequences are not cleaved by wild-type l-Crel at this temperature. Listed are the 2NN sequences and relative activities
63
for hHmuCre vs wild-type l-Crel.DNA sequence is based on the C1221target with the central four base pairs replaced as indicated. Relative activity is scaled as: -, no activity detectable; +, <25% activity; ++, 25% to <50% activity; +++, 50% to <75% activity; ++++, 75% to 100% activity.
64
TACA - + +
TATA - + + +
TCCA - + + +
Table 5: Additional 2NN targets cleaved by hHmuCre at 30"C. From the 136 possible 2NN targets, hHmuCre is able to cleave a total of 110 (see Table 3) at 30°C. Of these, 47 DNA sequences are not cleaved by wild-type l-Crel at this temperature. Listed are the 2NN sequences and relative activities for hHmuCre vs wild-type l-Crel.DNA sequence is based on the C1221target with the central four base pairs replaced as indicated. Relative activity is scaled as: -, no activity detectable; +, <25% activity; ++, 25% to <50% activity; +++, 50% to <75% activity; ++++, 75% to 100% activity.
Example 2
Using the truncated l-Cre_X scaffold (SEQ ID NO: 5) as a starting point, constructs were generated by adding variable length regions of the CFP1 peptide (SEQ ID NO: 112) to the C-terminus of the protein. As l-Crel is a homodimer, all fusion constructs contain two CFP1 peptides (Figure 3A, where "enhancer domain" = the CFP1 peptide domain).
The activity of each "enhanced" meganuclease (eMega) was assessed using yeast assay described in International PCT Applications WO 2004/067736 (see Example 1). To monitor the effect on activity resulting from the addition of the new enhancer domains, all constructs were screened against libraries of the C1221 target DNA (Figure 2) having regions mutated in the following positions (numbering as in Figure 2): 12NN, lONNNnn, 7NNNnn, 5NNN, 2NN. These targets contain nucleotide substitutions at the +/- positions indicated. Fusion proteins enhance cleavage of targets to various degrees relative to the activity of wild-type l-Crel.
65
List of cited references
Arimondo, P. B., C. J. Thomas, et al. (2006). "Exploring the cellular activity of camptothecin-triple- helix-forming oligonucleotide conjugates." Mol Cell Biol 26(1): 324-33.
Arnould, S., P. Chames, et al. (2006). Engineering of large numbers of highly specific homing
endonucleases that induce recombination on novel DNA targets. Journal of Molecular
Biology. 355: 443-58.
Arnould, S., P. Chames, et al. (2006). "Engineering of large numbers of highly specific homing
endonucleases that induce recombination on novel DNA targets." J Mol Biol 355(3): 443-58.
Arnould, S., C. Delenda, et al. (2011). "The l-Crel meganuclease and its engineered derivatives: applications from cell modification to gene therapy
gzq083 [pii] 10.1093/protein/gzq083." Protein engineering, design & selection : PEPS 24(1-2): 27-31. Arnould, S., C. Perez, et al. (2007). "Engineered l-Crel derivatives cleaving sequences from the human XPC gene can induce highly efficient gene correction in mammalian cells." Journal of
Molecular Biology 371(1): 49-65.
Ashworth, J., J. J. Havranek, et al. (2006). "Computational redesign of endonuclease DNA binding and cleavage specificity." Nature 441(7093): 656-9.
Bedayat, B., A. Abdolmohamadi, et al. (2010). "Sequence-specific correction of genomic
hypoxanthine-guanine phosphoribosyl transferase mutations in lymphoblasts by small fragment homologous replacement
10.1089/oli.2009.0205." Oligonucleotides 20(1): 7-16.
Bennardo, N., A. Cheng, et al. (2008). "Alternative-NHEJ is a mechanistically distinct pathway of mammalian chromosome break repair." PLoS Genet 4(6): elOOOHO.
Bennardo, N., A. Gunn, et al. (2009). "Limiting the persistence of a chromosome break diminishes its mutagenic potential." PLoS Genet 5(10): el000683.
Boch, J., H. Scholze, et al. (2009). "Breaking the code of DNA binding specificity of TAL-type III
effectors." Science 326(5959): 1509-12.
Boch, J., H. Scholze, et al. (2009). "Breaking the code of DNA binding specificity of TAL-type III
effectors
1178811 [pii] 10.1126/science.1178811." Science 326(5959): 1509-12.
Bolduc, J. M., P. C. Spiegel, et al. (2003). "Structural and biochemical analyses of DNA and NA
binding by a bifunctional homing endonuclease and group I intron splicing factor." Genes De 17(23): 2875-88.
Buis, J., Y. Wu, et al. (2008). "Mrell nuclease activity has essential roles in DNA repair and genomic stability distinct from ATM activation." Cell 135(1): 85-96.
Capecchi, M. R. (2001). "Generating mice with targeted mutations
10.1038/nml001-1086 nmlOOl-1086 [pii]." Nature Medicine 7(10): 1086-90.
Carroll, D. (2008). "Progress and prospects: zinc-finger nucleases as gene therapy agents gt2008145 [pii] 10.1038/gt.2008.145." Gene therapy 15(22): 1463-8.
Chames, P., J. C. Epinat, et al. (2005). "In vivo selection of engineered homing endonucleases using double-strand break induced homologous recombination." Nucleic Acids Res 33(20): el78. Chevalier, B., M. Turmel, et al. (2003). "Flexible DNA target site recognition by divergent homing endonuclease isoschizomers l-Crel and l-Msol." J Mol Biol 329(2): 253-69.
Chevalier, B. S., T. Kortemme, et al. (2002). "Design, activity, and structure of a highly specific
artificial endonuclease." Mol Cell 10(4): 895-905.
Chevalier, B. S., R. J. Monnat, Jr., et al. (2001). "The homing endonuclease l-Crel uses three metals, one of which is shared between the two active sites." Nat Struct Biol 8(4): 312-6.
Chevalier, B. S. and B. L. Stoddard (2001). "Homing endonucleases: structural and functional insight into the catalysts of intron/intein mobility." Nucleic Acids Res 29(18): 3757-74.
66
Choo, Y. and A. Klug ( 1994). "Selection of DNA binding sites for zinc fingers using rationally randomized DNA reveals coded interactions." Proceedings of the National Academy of
Sciences of the United States of America 91(23): 11168-72.
Choo, Y. and A. Klug ( 1994). "Toward a code for the interactions of zinc fingers with DNA: selection of randomized fingers displayed on phage." Proceedings of the National Academy of
Sciences of the United States of America 91(23): 11163-7.
Choulika, A., A. Perrin, et al. ( 1995). "Induction of homologous recombination in mammalian
chromosomes by using the l-Scel system of Saccharomyces cerevisiae." Mol Cell Biol 15(4):
1968-73.
Christian, M., T. Cermak, et al. (2010). "Targeting DNA double-strand breaks with TAL effector
nucleases." Genetics 186(2): 757-61.
Christian, M., T. Cermak, et al. (2010). "Targeting DNA double-strand breaks with TAL effector
nucleases
genetics.110.120717 [pii] 10.1534/genetics.ll0.120717." Genetics 186(2): 757-61.
Cost, G. J., Y. Freyvert, et al. (2010). " BAK and BAX deletion using zinc-finger nucleases yields
apoptosis-resistant CHO cells
10.1002/bit.22541." Biotechnology and Bioengineering 105(2): 330-40.
Delacote, F. and B. S. Lopez (2008). "Importance of the cell cycle phase for the choice of the
appropriate DSB repair pathway, for genome stability maintenance: the trans-S double- strand break repair model
5149 fpiil ." Cell Cycle 7(1) : 33-8.
Doudeva, L. G., H. Huang, et al. (2006). "Crystal structural analysis and metal-dependent stability and activity studies of the ColE7 endonuclease domain in complex with DNA/Zn2+ or inhibitor/Ni2+
15/2/269 [pii] 10.1110/ps.051903406." Protein science : a publication of the Protein Society 15(2):
269-80.
Doyon, J. B., V. Pattanayak, et al. (2006). "Directed evolution and substrate specificity profile of homing endonuclease l-Scel." Journal of the American Chemical Society 128(7): 2477-84. Doyon, Y., J. M . McCammon, et al. (2008). " Heritable targeted gene disruption in zebrafish using designed zinc-finger nucleases
nbtl409 [pii] 10.1038/nbtl409." Nature Biotechnology 26(6) : 702-8.
Eastberg, J. H., J . Eklund, et al. (2007). "M utability of an HNH nuclease imidazole general base and exchange of a deprotonation mechanism ." Biochemistry 46(24): 7215-25.
Eisenschmidt, K., T. Lanio, et al. (2005 ). "Developing a programmed restriction endonuclease for highly specific DNA cleavage." Nucleic Acids Res 33(22): 7039-47.
Elrod-Erickson, M ., M . A. Rould, et al. (1996). "Zif268 protein-DNA complex refined at 1.6 A: a model system for understanding zinc finger-DNA interactions." Structure 4(10): 1171-80.
Epinat, J. C, S. Arnould, et al. (2003). "A novel engineered meganuclease induces homologous
recombination in yeast and mammalian cells." Nucleic Acids Res 31( 11): 2952-62.
Frank, K. M ., J. M. Sekiguchi, et al. ( 1998). "Late embryonic lethality and impaired V(D)J
recombination in mice lacking DNA ligase IV
10.1038/24172." Nature 396(6707): 173-7.
Galetto, R., P. Duchateau, et al. (2009). "Targeted approaches for gene therapy and the emergence of engineered meganucleases
10.1517/14712590903213669." Expert opinion on biological therapy 9( 10): 1289-303.
Gao, H ., J. Smith, et al. (2010). " Heritable targeted mutagenesis in maize using a designed
endonuclease
TPJ4041 [pii] 10.1111/i. l365-313X.2009.04041.x." The Plant journal : for cell and molecular biology
61(1): 176-87.
Gao, Y., Y. Sun, et al. ( 1998). "A critical role for DNA end-joining proteins in both lymphogenesis and neurogenesis
67
S0092-8674(00)81714-6 [pii]." Cell 95(7): 891-902.
Geurts, A. M .( G. J. Cost, et al. (2009). " Knockout rats via embryo microinjection of zinc-finger
nucleases
325/5939/433 [pii] 10.1126/science.1172447." Science 325(5939): 433.
Gimble, F. S., C. M. Moure, et al. (2003). "Assessing the plasticity of DNA target site recognition of the Pl-Scel homing endonuclease using a bacterial two-hybrid selection system." J Mol Biol 334(5): 993-1008.
Greisman, H. A. and C. O. Pabo (1997). "A general strategy for selecting high-affinity zinc finger proteins for diverse DNA target sites." Science 275(5300): 657-61.
Gruenert, D. C, E. Bruscia, et al. (2003). "Sequence-specific modification of genomic DNA by small DNA fragments
10.1172/JCI19773 112/5/637 [pii]." The Journal of clinical investigation 112(5): 637-41.
Guirouilh-Barbat, J., S. Huck, et al. (2004). "Impact of the KU80 pathway on NHEJ-induced genome rearrangements in mammalian cells." Mol Cell 14(5): 611-23.
Guirouilh-Barbat, J ., S. Huck, et al. (2004). " Impact of the KU80 pathway on N HEJ-induced genome rearrangements in mammalian cells
10.1016/j.molcel.2004.05.008 S1097276504002916 [pii]." Molecular Cell 14(5): 611-23.
Guirouilh-Barbat, J ., E. Rass, et al. (2007). "Defects in XRCC4 and KU80 differentially affect the joining of distal nonhomologous ends." Proc Natl Acad Sci U S A 104(52): 20902-7.
Guirouilh-Barbat, J., E. Rass, et al. (2007). "Defects in XRCC4 and KU80 differentially affect the joining of distal nonhomologous ends
0708541104 [pii] 10.1073/pnas.0708541104." Proceedings of the National Academy of Sciences of the United States of America 104(52) : 20902-7.
Gurlebeck, D., B. Szurek, et al. (2005). "Dimerization of the bacterial effector protein AvrBs3 in the plant cell cytoplasm prior to nuclear import
TPJ2370 [pii] 10.1111/j.l365-313X.2005.02370.x." The Plant journal : for cell and molecular biology
42(2): 175-87.
Haber, J. (2000). "Partners and pathwaysrepairing a double-strand break." Trends Genet.16(6): 259- 264.
Haber, J. E. (2008). "Alternative endings
0711334105 [pii] 10.1073/pnas.0711334105." Proceedings of the National Academy of Sciences of the United States of America 105(2): 405-6.
Hartsuiker, E., K. Mizuno, et al. (2009). "CtplCtIP and Rad32Mrell nuclease activity are required for Recl2Spoll removal, but Recl2Spoll removal is dispensable for other M RN-dependent meiotic functions." Mol Cell Biol 29(7): 1671-81.
Hinnen, A., J. B. Hicks, et al. ( 1978). "Transformation of yeast." Proceedings of the National Academy of Sciences of the United States of America 75(4): 1929-33.
Hirata, R., J. Chamberlain, et al. (2002). "Targeted transgene insertion into human chromosomes by adeno-associated virus vectors
10.1038/nbt0702-735 nbt0702-735 [pii]." Nature Biotechnology 20(7): 735-8.
Huang, H. and H. S. Yuan (2007). "The conserved asparagine in the H NH motif serves an important structural role in metal finger endonucleases." Journal of Molecular Biology 368(3): 812-21. Ichiyanagi, K., Y. Ishino, et al. (2000). "Crystal structure of an archaeal intein-encoded homing
endonuclease Pl-Pful." J Mol Biol 300(4): 889-901.
Inoue, N., R. Dong, et al. (2001). "Introduction of single base substitutions at homologous
chromosomal sequences by adeno-associated virus vectors
10.1006/mthe.2001.0283 S1525-0016(01)90283-7 [pii]." Molecular therapy : the journal of the
American Society of Gene Therapy 3(4): 526-30.
Isalan, M . and Y. Choo (2001). "Rapid, high-throughput engineering of sequence-specific zinc finger DNA-binding proteins
S0076-6879(01)40444-7 [pii] ." Methods in Enzymology 340: 593-609.
68
Kalish, J. M. and P. M. Glazer (2005). "Targeted genome modification via triple helix formation." Ann
N Y Acad Sci 1058: 151-61.
Kim, H. J., H. J. Lee, et al. (2009). "Targeted genome editing in human cells with zinc finger nucleases constructed via modular assembly
gr.089417.108 [pii] 10.1101/gr.089417.108." Genome Research 19(7): 1279-88.
Kim, Y. G., J. Cha, et al. (1996). "Hybrid restriction enzymes: zinc finger fusions to Fok I cleavage
domain." Proceedings of the National Academy of Sciences of the United States of America
93(3): 1156-60.
Ku, W. Y., Y. W. Liu, et al. (2002). "The zinc ion in the HN H motif of the endonuclease domain of colicin E7 is not required for DNA binding but is essential for DNA hydrolysis." Nucleic Acids
Research 30(7): 1670-8.
Landthaler, M U. Begley, et al. (2002). "Two self-splicing group I introns in the ribonucleotide
reductase large subunit gene of Staphylococcus aureus phage Twort." Nucleic Acids
Research 30(9): 1935-43.
Landthaler, M ., N. C. Lau, et al. (2004). "Group I intron homing in Bacillus phages SPOl and SP82: a gene conversion event initiated by a nicking homing endonuclease." Journal of Bacteriology 186(13): 4307-14.
Landthaler, M ., B. W. Shen, et al. (2006). "I-Basl and l-Hmul : two phage intron-encoded
endonucleases with homologous DNA recognition sequences but distinct DNA specificities." Journal of Molecular Biology 358(4): 1137-51.
Landthaler, M. and D. A. Shub (2003). "The nicking homing endonuclease l-Basl is encoded by a
group I intron in the DNA polymerase gene of the Bacillus thuringiensis phage Bastille."
Nucleic Acids Research 31(12): 3071-7.
Lee, S. E., F. Paques, et al. (1999). "Role of yeast SIR genes and mating type in directing DNA double- strand breaks to homologous and non-homologous repair paths." Curr Biol 9(14): 767-70.
Li, T., S. Huang, et al. (2010). "TAL nucleases (TALNs): hybrid proteins composed of TAL effectors and
Fokl DNA-cleavage domain." Nucleic Acids Res 39(1): 359-72.
Li, T., S. Huang, et al. (2011). "TAL nucleases (TALNs): hybrid proteins composed of TAL effectors and
Fokl DNA-cleavage domain
gkq704 [pii] 10.1093/nar/gkq704." Nucleic Acids Research 39(1): 359-72.
Liang, F., M . Han, et al. (1998). "Homology-directed repair is a major double-strand break repair pathway in mammalian cells." Proceedings of the National Academy of Sciences of the
United States of America 95(9): 5172-7.
Liu, P. Q., E. M. Chan, et al. (2010). "Generation of a triple-gene knockout mammalian cell line using engineered zinc-finger nucleases
10.1002/bit.22654." Biotechnology and Bioengineering 106(1): 97-105.
Liu, Q., V. Derbyshire, et al. (2006). " Distance determination by G IY-YIG intron endonucleases:
discrimination between repression and cleavage functions." Nucleic Acids Research 34(6):
1755-64.
Lloyd, A., C. L. Plaisier, et al. (2005). "Targeted mutagenesis using zinc-finger nucleases in Arabidopsis 0409339102 [pii] 10.1073/pnas.0409339102." Proceedings of the National Academy of Sciences of the United States of America 102(6): 2232-7.
Maeder, M. L., S. Thibodeau-Beganny, et al. (2008). "Rapid "open-source" engineering of customized zinc-finger nucleases for highly efficient gene modification
S1097-2765(08)00461-9 [pii] 10.1016/j.molcel.2008.06.016." Molecular Cell 31(2): 294-301.
Mahfouz, M . M ., L. Li, et al. (2011). "De novo-engineered transcription activator-like effector (TALE) hybrid nuclease with novel DNA binding specificity creates double-strand breaks
1019533108 [pii] 10.1073/pnas.1019533108." Proceedings of the National Academy of Sciences of the United States of America 108(6): 2623-8.
Marcaida, M . J., I. G . Munoz, et al. (2010). " Homing endonucleases: from basics to therapeutic
applications
69
10.1007/s00018-009-0188-v." Cellular and molecular life sciences : CMLS 67(5): 727-48.
Mashimo, T., A. Takizawa, et al. (2010). "Generation of knockout rats with X-linked severe combined immunodeficiency (X-SCID) using zinc-finger nucleases
10.1371/journal.pone.0008870." PloS one 5(1): e8870.
McConnell Smith, A., . Takeuchi, et al. (2009). "Generation of a nicking enzyme that stimulates site- specific gene conversion from the l-Anil LAGLIDADG homing endonuclease
0810588106 [pii] 10.1073/pnas.0810588106." Proceedings of the National Academy of Sciences of the United States of America 106(13): 5099-104.
McVey, M. and S. E. Lee (2008). "MMEJ repair of double-strand breaks (director's cut): deleted
sequences and alternative endings
S0168-9525(08)00229-l [pii] 10.1016/i.tig.2008.08.007." Trends in genetics : TIG 24(11): 529-38.
Menoret, S., A. L. Iscache, et al. (2010). "Characterization of immunoglobulin heavy chain knockout rats
10.1002/eii.201040939." European Journal of Immunology 40(10): 2932-41.
Midon, M., P. Schafer, et al. (2011). "Mutational and biochemical analysis of the DNA-entry nuclease EndA from Streptococcus pneumoniae
gkq802 [pii] 10.1093/nar/gkq802." Nucleic Acids Research 39(2): 623-34.
Miller, J. C, S. Tan, et al. (2011). "A TALE nuclease architecture for efficient genome editing nbt.1755 [pii] 10.1038/nbt.l755." Nature Biotechnology 29(2): 143-8.
Mimitou, E. P. and L. S. Symington (2008). "Sae2, Exol and Sgsl collaborate in DNA double-strand break processing." Nature 455(7214): 770-4.
Moore, I., M. Samalova, et al. (2006). "Transactivated and chemically inducible gene expression in plants." Plant J 45(4): 651-83.
Moore, J. K. and J. E. Haber (1996). "Cell cycle and genetic requirements of two pathways of
nonhomologous end-joining repair of double-strand breaks in Saccharomyces cerevisiae."
Mol Cell Biol 16(5): 2164-73.
Moscou, M. J. and A. J. Bogdanove (2009). "A simple cipher governs DNA recognition by TAL
effectors." Science 326(5959): 1501.
Moscou, M. J. and A. J. Bogdanove (2009). "A simple cipher governs DNA recognition by TAL
effectors
1178817 [pii] 10.1126/science.1178817." Science 326(5959): 1501.
Moure, C. M., F. S. Gimble, et al. (2002). "Crystal structure of the intein homing endonuclease Pl-Scel bound to its recognition sequence." Nat Struct Biol 9(10): 764-70.
Moure, C. M., F. S. Gimble, et al. (2003). "The crystal structure of the gene targeting homing
endonuclease l-Scel reveals the origins of its target site specificity." J Mol Biol 334(4): 685-
95.
Nimonkar, A. V., J. Genschel, et al. (2011). "BLM-DNA2-RPA-MRN and EXOl-BLM-RPA-MRN
constitute two DNA end resection machineries for human DNA break repair." Genes Dev 25(4): 350-62.
Niu, Y., K. Tenney, et al. (2008). "Engineering variants of the l-Scel homing endonuclease with strand- specific and site-specific DNA-nicking activity
S0022-2836(08)00840-l [pii] 10.1016/i.jmb.2008.07.010." Journal of Molecular Biology 382(1): 188- 202.
Orr-Weaver, T. L., J. W. Szostak, et al. (1981). "Yeast transformation: a model system for the study of recombination." Proceedings of the National Academy of Sciences of the United States of
America 78(10): 6354-8.
Orr-Weaver, T. L., J. W. Szostak, et al. (1983). "Genetic applications of yeast transformation with linear and gapped plasmids." Methods in Enzymology 101: 228-45.
Pabo, C. O., E. Peisach, et al. (2001). "Design and selection of novel Cys2His2 zinc finger proteins 70/1/313 [pii] 10.1146/annurev.biochem.70.1.313." Annual Review of Biochemistry 70: 313-40.
70
Padidam, M. (2003). "Chemically regulated gene expression in plants." Curr Opin Plant Biol 6(2): 169- 77.
Paques, F. and P. Duchateau (2007). "Meganucleases and DNA double-strand break-induced
recombination: perspectives for gene therapy." Current Gene Therapy 7(1): 49-66.
Paques, F. and J. E. Haber (1999). "Multiple pathways of recombination induced by double-strand breaks in Saccharomyces cerevisiae." Microbiol Mol Biol Rev 63(2): 349-404.
Paques, F. and J. E. Haber (1999). "Multiple pathways of recombination induced by double-strand breaks in Saccharomyces cerevisiae." Microbiology and molecular biology reviews : MMBR
63(2): 349-404.
Perez, E. E., J. Wang, et al. (2008). "Establishment of HIV-1 resistance in CD4+ T cells by genome editing using zinc-finger nucleases
nbtl410 [pii] 10.1038/nbtl410." Nature Biotechnology 26(7): 808-16.
Pierce, A. J., P. Hu, et al. (2001). "Ku DNA end-binding protein modulates homologous repair of double-strand breaks in mammalian cells." Genes Dev 15(24): 3237-42.
Pingoud, A. and G. H. Silva (2007). "Precision genome surgery." Nat Biotechnol 25(7): 743-4.
Porteus, M. H. and D. Carroll (2005). "Gene targeting using zinc finger nucleases." Nat Biotechnol
23(8): 967-73.
Ramirez, C. L, J. E. Foley, et al. (2008). "Unexpected failure rates for modular assembly of
engineered zinc fingers
nmeth0508-374 [pii] 10.1038/nmeth0508-374." Nature Methods 5(5): 374-5.
Rosen, L. E., H. A. Morrison, et al. (2006). "Homing endonuclease l-Crel derivatives with novel DNA target specificities." Nucleic Acids Research 34(17): 4791-800.
Rothstein, R. J. (1983). "One-step gene disruption in yeast." Methods in Enzymology 101: 202-11. Rouet, P., F. Smih, et al. (1994). "Expression of a site-specific endonuclease stimulates homologous recombination in mammalian cells." Proc Natl Acad Sci U S A 91(13): 6064-8.
Rouet, P., F. Smih, et al. (1994). "Introduction of double-strand breaks into the genome of mouse cells by expression of a rare-cutting endonuclease." Mol Cell Biol 14(12): 8096-106.
Russell, D. W. and R. K. Hirata (1998). "Human gene targeting by viral vectors
10.1038/ng0498-325." Nature Genetics 18(4): 325-30.
Sangiuolo, F., M. L. Scaldaferri, et al. (2008). "Cftr gene targeting in mouse embryonic stem cells mediated by Small Fragment Homologous Replacement (SFHR)
2904 [pii]." Frontiers in bioscience : a journal and virtual library 13: 2989-99.
Santiago, Y., E. Chan, et al. (2008). "Targeted gene knockout in mammalian cells by using engineered zinc-finger nucleases
0800940105 [pii] 10.1073/pnas.0800940105." Proceedings of the National Academy of Sciences of the United States of America 105(15): 5809-14.
Sartori, A. A., C. Lukas, et al. (2007). "Human CtIP promotes DNA end resection." Nature 450(7169):
509-14.
Seligman, L. M., K. M. Chisholm, et al. (2002). "Mutations altering the cleavage specificity of a
homing endonuclease." Nucleic Acids Research 30(17): 3870-9.
Seligman, L. M., K. M. Stephens, et al. (1997). "Genetic analysis of the Chlamydomonas reinhardtii I-
Crel mobile intron homing system in Escherichia coli." Genetics 147(4): 1653-64.
Shen, B. W., M. Landthaler, et al. (2004). "DNA binding and cleavage by the HNH homing
endonuclease l-Hmul." Journal of Molecular Biology 342(1): 43-56.
Shukla, V. K., Y. Doyon, et al. (2009). "Precise genome modification in the crop species Zea mays using zinc-finger nucleases
nature07992 [pii] 10.1038/nature07992." Nature 459(7245): 437-41.
Silva, G. H., J. Z. Dalgaard, et al. (1999). "Crystal structure of the thermostable archaeal intron- encoded endonuclease l-Dmol." J Mol Biol 286(4): 1123-36.
Simon, P., F. Cannata, et al. (2008). "Sequence-specific DNA cleavage mediated by bipyridine
polyamide conjugates." Nucleic Acids Res 36(11): 3531-8.
71
Smith, J., J. M. Berg, et al. (1999). "A detailed study of the substrate specificity of a chimeric restriction enzyme
gkcl39 [pii]." Nucleic Acids Research 27(2): 674-81.
Smith, J., M. Bibikova, et al. (2000). "Requirements for double-strand cleavage by chimeric
restriction enzymes with zinc finger DNA-recognition domains." Nucleic Acids Res 28(17):
3361-9.
Smith, J., M. Bibikova, et al. (2000). "Requirements for double-strand cleavage by chimeric
restriction enzymes with zinc finger DNA-recognition domains." Nucleic Acids Research 28(17): 3361-9.
Smith, J., S. Grizot, et al. (2006). "A combinatorial approach to create artificial homing endonucleases cleaving chosen sequences." Nucleic Acids Research 34(22): el49.
Sonoda, E., H. Hochegger, et al. (2006). "Differential usage of non-homologous end-joining and
homologous recombination in double strand break repair." DNA Repair (Amst) 5(9-10): 1021-9.
Spiegel, P. C, B. Chevalier, et al. (2006). "The structure of l-Ceul homing endonuclease: Evolving asymmetric DNA recognition from a symmetric protein scaffold." Structure 14(5): 869-80. Stoddard, B. L. (2005). "Homing endonuclease structure and function." Quarterly Reviews of
Biophysics 38(1): 49-95.
Stoddard, B. L, A. M. Scharenberg, et al. (2007). Advances in Engineering Homing Endonucleases for Gene Targeting: Ten Years After Structures. Progress in Gene Therapy: Autologous and
Cancer Stem Cell Gene Therapy. R. Bertolotti and K. Ozawa, World Scientific Publishing Co. Pte. Ltd. 3: 135-68.
Sugawara, N. and J. E. Haber (1992). "Characterization of double-strand break-induced
recombination: homology requirements and single-stranded DNA formation." Mol Cell Biol 12(2): 563-75.
Sun, H., D. Treco, et al. (1991). "Extensive 3'-overhanging, single-stranded DNA associated with the meiosis-specific double-strand breaks at the ARG4 recombination initiation site." Cell 64(6): 1155-61.
Sussman, D., M. Chadsey, et al. (2004). "Isolation and characterization of new homing endonuclease specificities at individual target site positions." Journal of Molecular Biology 342(1): 31-41.
Taubes, G. (2002). "Gene therapy. The strange case of chimeraplasty
10.1126/science.298.5601.2116 298/5601/2116 [pii]." Science 298(5601): 2116-20.
Wang, R., X. Zhou, et al. (2003). "Chemically regulated expression systems and their applications in transgenic plants." Transgenic Res 12(5): 529-40.
Wang, Y. T., J. D. Wright, et al. (2009). "Redesign of high-affinity nonspecific nucleases with altered sequence preference
10.1021/ja907160r." Journal of the American Chemical Society 131(47): 17345-53.
White, C. I. and J. E. Haber (1990). "Intermediates of recombination during mating type switching in Saccharomyces cerevisiae." Embo J 9(3): 663-73.
Yang, M., V. Djukanovic, et al. (2009). "Targeted mutagenesis in the progeny of maize transgenic plants
10.1007/S11103-009-9499-5." Plant Molecular Biology 70(6): 669-79.
Zhao, L, R. P. Bonocora, et al. (2007). "The restriction fold turns to the dark side: a bacterial homing endonuclease with a PD-(D/E)-XK motif." The EMBO Journal 26(9): 2432-42.
Zuo, J. and N. H. Chua (2000). "Chemical-inducible systems for regulated expression of plant genes." Curr Opin Biotechnol 11(2): 146-51.
72
Claims
A method for enhancing rare-cutting endonuclease efficiency at a genomic locus of interest in a cell comprising the steps of:
(i) identifying at said genomic locus of interest one DNA target sequence cleavable by one rare-cutting endonuclease;
(ii) engineering said rare-cutting endonuclease in order to create a chimeric rare-cutting endonuclease wherein said chimeric rare-cutting endonuclease is a fusion protein comprising at least a rare-cutting endonuclease-derived scaffold and at least one enhancer domain and wherein said chimeric rare-cutting endonuclease has an enhanced efficiency for said DNA target sequence relative to the efficiency of the starting rare-cutting endonuclease for said DNA target sequence;
(iii) contacting said DNA target sequence with said chimeric rare-cutting endonuclease for cleavage within the genomic locus of interest; thereby obtaining a cell in which rare-cutting endonuclease efficiency at a genomic locus of interest in a cell is enhanced.
The method according to claim 1 wherein said enhancer domain is fused to the N-terminus part of said rare-cutting endonuclease-derived scaffold.
The method according to claim 1 wherein said enhancer domain is fused to the C-terminus part of said rare-cutting endonuclease-derived scaffold.
The method according to claim 1 wherein two enhancer domains are fused to both N- terminus part of said rare-cutting endonuclease-derived scaffold and C-terminus part of said rare-cutting endonuclease-derived scaffold.
The method according to claim 1 wherein said rare-cutting endonuclease-derived scaffold is derived from the group consisting of l-Crel, a functional mutant of l-Crel, a variant of l-Crel or a derivative thereof.
73
6) The method according to claim 5 wherein said rare-cutting endonuclease-derived scaffold comprises a sequence selected from the group consisting of l-Crel_NFSl (SEQ ID NO: 6); I- Crel_NFS2 (SEQ ID NO: 7) and l-Crel_CFSl (SEQ ID NO: 8).
7) The method according to claim 1 wherein said enhancer domain consists of a protein domain derived from a protein selected from the group consisting of proteins listed in Table 1, a functional mutant, a variant or a derivative thereof.
8) The method according to claim 7 wherein said enhancer domain is l-Hmul, a functional mutant, a variant or a derivative thereof.
9) The method according to claim 1 wherein said chimeric rare-cutting endonuclease further comprises at least one peptidic linker between said rare-cutting endonuclease-derived scaffold and said at least one enhancer domain.
10) The method according to claim 9 wherein said peptidic linker sequence is selected from the group consisting of peptides listed in Table 2.
11) The method according to claim 1 wherein said chimeric rare-cutting endonuclease processes 2NN derivatives of said DNA target sequence that are not cleaved by wild-type rare-cutting endonuclease.
12) The method according to claim 1 wherein said chimeric rare-cutting endonuclease is selected from the group consisting of hHmuCre_D0101 (SEQ ID NO: 9), hHmuCre_D0201 (SEQ ID NO: 105), hHmuCre_D0301 (SEQ ID NO: 106), hHmuCre_D0401 (SEQ ID NO: 107), hHmuCre_D0102 (SEQ ID NO: 108), hHmuCre_D0202 (SEQ ID NO: 109), hHmuCre_D0302 (SEQ ID NO: 110) and hHmuCre_D0402 (SEQ ID NO: 111).
13) A method to create a chimeric rare-cutting endonuclease with enhanced efficiency for a DNA target sequence comprising the steps of:
(i) Engineering a rare-cutting endonuclease-derived scaffold from a rare-cutting endonuclease capable to cleave said DNA target sequence;
(ii) determining or engineering an enhancer domain wherein said enhancer domain enhances the efficiency of said rare-cutting endonuclease when fused to it;
(iii) optionally determining or engineering a peptidic linker to fuse said rare-cutting endonuclease-derived scaffold to said enhancer domain;
74
thereby obtaining a chimeric rare-cutting endonuclease with enhanced efficiency for a DNA target sequence.
14) The method according to claim 13 wherein said enhancer domain is fused to the N-terminus part of said rare-cutting endonuclease-derived scaffold.
15) The method according to claim 13 wherein said enhancer domain is fused to the C-terminus part of said rare-cutting endonuclease-derived scaffold.
16) The method according to claim 13 wherein two enhancer domains are fused to both N- terminus part of said rare-cutting endonuclease-derived scaffold and C-terminus part of said rare-cutting endonuclease-derived scaffold.
17) The method according to claim 13 wherein said rare-cutting endonuclease-derived scaffold is derived from l-Crel, a functional mutant of l-Crel, a variant of l-Crel or a derivative thereof.
18) The method according to claim 17 wherein said rare-cutting endonuclease-derived scaffold comprises a sequence selected from the group consisting of l-Crel_NFSl (SEQ ID NO: 6); I- Crel_NFS2 (SEQ ID NO: 7) and l-Crel_CFSl (SEQ ID NO: 8).
19) The method according to claim 1 wherein said enhancer domain consists of a protein domain derived from a protein selected from the group consisting of proteins listed in Table 1, a functional mutant, a variant or a derivative thereof.
20) The method according to claim 19 wherein said enhancer domain is selected from the group consisting of l-Hmul, a functional mutant, a variant or a derivative thereof.
21) The method according to claim 20 wherein said chimeric rare-cutting endonuclease can comprise a peptidic linker selected from the group consisting of peptides listed in Table 2.
22) The method according to claim 13 wherein said chimeric rare-cutting endonuclease processes 2NN derivatives of said DNA target sequence that are not cleaved by wild-type rare-cutting endonuclease.
23) A chimeric rare-cutting endonuclease for a DNA target sequence comprising:
75
(i) a rare-cutting endonuclease-derived scaffold;
(ii) an enhancer domain wherein said enhancer domain enhances the efficiency of said rare-cutting endonuclease for said DNA target sequence when fused to it;
(iii) optionally a peptidic linker.
24) The chimeric rare-cutting endonuclease according to claim 23 wherein said enhancer domain is fused to the N-terminus part of said rare-cutting endonuclease-derived scaffold.
25) The chimeric rare-cutting endonuclease according to claim 23 wherein said enhancer domain is fused to the C-terminus part of said rare-cutting endonuclease-derived scaffold.
26) The chimeric rare-cutting endonuclease according to claim 23 wherein two enhancer domains are fused to both N-terminus part of said rare-cutting endonuclease-derived scaffold and C-terminus part of said rare-cutting endonuclease-derived scaffold.
27) The chimeric rare-cutting endonuclease according to claim 23 wherein said rare-cutting endonuclease-derived scaffold is derived from l-Crel, a functional mutant of l-Crel, a variant of l-Crel or a derivative thereof.
28) The chimeric rare-cutting endonuclease according to claim 27 wherein said rare-cutting endonuclease-derived scaffold comprises a sequence selected from the group consisting of I- Crel_NFSl (SEQ ID NO: 6); l-Crel_NFS2 (SEQ ID NO: 7) and l-Crel_CFSl (SEQ ID NO: 8).
29) The chimeric rare-cutting endonuclease according to claim 23 wherein said enhancer domain consists of a protein domain derived from a protein selected from the group consisting of SEQ ID NOs: 10-66.
30) The chimeric rare-cutting endonuclease according to claim 23 wherein said enhancer domain is l-Hmul.
31) The chimeric rare-cutting endonuclease according to claim 23 wherein said peptidic linker sequence can be selected from the group consisting of SEQ ID NOs: 67-104
32) The chimeric rare-cutting endonuclease according to claim 23 wherein said chimeric rare- cutting endonuclease processes 2NN derivatives of said DNA target sequence that are not cleaved by wild-type rare-cutting endonuclease.
76
33) The chimeric rare-cutting endonuciease according to claim 23 wherein said chimeric rare- cutting endonuciease is selected from the group consisting of SEQ ID NO: 9 and SEQ ID NOs: 105-109.
34) A recombinant polynucleotide encoding a chimeric rare-cutting endonuciease according to claim 23.
35) A vector comprising a recombinant polynucleotide according to claim 34.
36) A composition comprising a chimeric rare-cutting endonuciease according to claim 23 and a carrier.
37) A pharmaceutical composition comprising a chimeric rare-cutting endonuciease according to claim 23 and a pharmaceutically active carrier. 38) A host cell which comprises a recombinant polynucleotide of claim 34.
39) A non-human transgenic animal which comprises a recombinant polynucleotide of claim 34.
40) A non-human transgenic animal which comprises a vector of claim 35.
41) A transgenic plant which comprises a recombinant polynucleotide of claim 34. 42) A transgenic plant which comprises a vector of claim 35.
43) A method of treatment of a genetic disease caused by a mutation in a specific double- stranded DNA target sequence in a gene comprising administering to a subject in need thereof an effective amount of a chimeric rare-cutting endonuciease according to claim 23.
44) A method for inserting a transgene into a specific double-stranded DNA target sequence of a genomic locus of a cell, tissue or non-human animal wherein at least one chimeric rare- cutting endonuciease according to claim 23 is introduced in said cell, tissue or non-human animal.
45) A kit comprising a chimeric rare-cutting endonuciease according to claim 23 and instructio for use in enhancing efficiency for a DNA target sequence.
77
46) A method for increasing targeted Homologous Recombination by using chimeric rare-cutting endonucleases according to claim 23.
A method for increasing targeted Homologous Recombination with less Non Homologous End-joining by using chimeric rare-cutting endonucleases according to claim 23 wherein at least one enhancer domain is a catalytically active nickase domain.
48) A method for increasing excision of a single-strand of DNA spanning the starting rare-cutting endonuclease binding region by using chimeric rare-cutting endonucleases according to claim 23 wherein:
(i) one enhancer domain is a catalytically active nickase domain;
(ii) one enhancer domain is a catalytically active cleavase domain;
the rare-cutting endonuclease-derived scaffold is an inactive rare-cutting endonuclease scaffold which serves as a scaffold for targeted DNA binding.
49) A chimeric rare-cutting endonuclease for a DNA target sequence comprising:
(i) a rare-cutting endonuclease-derived scaffold;
(ii) an enhancer domain fused to a rare-cutting endonuclease; and optionally
(iii) a peptidic linker.
50) A method for enhancing processing of a nucleic acid sequence compared to the processing of a parent rare-cutting endonuclease comprising:
(i) identifying one DNA target sequence recognized by one rare-cutting endonuclease; (ii) producing a chimeric rare-cutting endonuclease by fusing at least a rare-cutting endonuclease-derived scaffold with at least one enhancer domain;
(iii) selecting a chimeric rare-cutting endonuclease with enhanced efficiency to process a nucleic acid sequence compared to the parent rare-cutting endonuclease;
(iv) contacting a nucleic acid containing said DNA target sequence with said chimeric rare- cutting endonuclease for a time an under conditions sufficient for processing of said nucleic acid.
78
Applications Claiming Priority (2)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| US201161472053P | 2011-04-05 | 2011-04-05 | |
| US61/472,053 | 2011-04-05 |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| WO2012138901A1 true WO2012138901A1 (en) | 2012-10-11 |
Family
ID=45952663
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/US2012/032386 Ceased WO2012138901A1 (en) | 2011-04-05 | 2012-04-05 | Method for enhancing rare-cutting endonuclease efficiency and uses thereof |
Country Status (1)
| Country | Link |
|---|---|
| WO (1) | WO2012138901A1 (en) |
Cited By (4)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO2014118719A1 (en) * | 2013-02-01 | 2014-08-07 | Cellectis | Tevl chimeric endonuclease and their preferential cleavage sites |
| JP2019503198A (en) * | 2016-02-02 | 2019-02-07 | サンガモ セラピューティクス, インコーポレイテッド | Composition for linking a DNA binding domain and a cleavage domain |
| EP4130256A4 (en) * | 2020-06-28 | 2024-04-10 | Jiangsu Conquers Medical Technology Co., Ltd. | Enzyme reaction solution for constructing sequencing library and use thereof |
| WO2025101946A1 (en) * | 2023-11-08 | 2025-05-15 | Precision Biosciences, Inc. | Polypeptide linkers for use in engineered meganucleases |
Citations (27)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US4179337A (en) | 1973-07-20 | 1979-12-18 | Davis Frank F | Non-immunogenic polypeptides |
| US4683195A (en) | 1986-01-30 | 1987-07-28 | Cetus Corporation | Process for amplifying, detecting, and/or-cloning nucleic acid sequences |
| US5006333A (en) | 1987-08-03 | 1991-04-09 | Ddi Pharmaceuticals, Inc. | Conjugates of superoxide dismutase coupled to high molecular weight polyalkylene glycols |
| WO2003078619A1 (en) | 2002-03-15 | 2003-09-25 | Cellectis | Hybrid and single chain meganucleases and use thereof |
| WO2004031346A2 (en) | 2002-09-06 | 2004-04-15 | Fred Hutchinson Cancer Research Center | Methods and compositions concerning designed highly-specific nucleic acid binding proteins |
| WO2004067736A2 (en) | 2003-01-28 | 2004-08-12 | Cellectis | Custom-made meganuclease and use thereof |
| WO2006097854A1 (en) | 2005-03-15 | 2006-09-21 | Cellectis | Heterodimeric meganucleases and use thereof |
| WO2006097784A1 (en) | 2005-03-15 | 2006-09-21 | Cellectis | I-crei meganuclease variants with modified specificity, method of preparation and uses thereof |
| WO2007034262A1 (en) | 2005-09-19 | 2007-03-29 | Cellectis | Heterodimeric meganucleases and use thereof |
| WO2007047859A2 (en) * | 2005-10-18 | 2007-04-26 | Precision Biosciences | Rationally-designed meganucleases with altered sequence specificity and dna-binding affinity |
| WO2007049156A2 (en) | 2005-10-25 | 2007-05-03 | Cellectis | I-CreI HOMING ENDONUCLEASE VARIANTS HAVING NOVEL CLEAVAGE SPECIFICITY AND USE THEREOF |
| WO2007049095A1 (en) | 2005-10-25 | 2007-05-03 | Cellectis | Laglidadg homing endonuclease variants having mutations in two functional subdomains and use thereof |
| WO2007093918A2 (en) | 2006-02-13 | 2007-08-23 | Cellectis | Meganuclease variants cleaving a dna target sequence from a xeroderma pigmentosum gene and uses thereof |
| WO2008010093A2 (en) | 2006-07-18 | 2008-01-24 | Cellectis | Meganuclease variants cleaving a dna target sequence from a rag gene and uses thereof |
| WO2008059382A2 (en) | 2006-11-14 | 2008-05-22 | Cellectis | Meganuclease variants cleaving a dna target sequence from the hprt gene and uses thereof |
| WO2008093249A2 (en) | 2007-02-01 | 2008-08-07 | Cellectis | Obligate heterodimer meganucleases and uses thereof |
| WO2008102274A2 (en) | 2007-02-20 | 2008-08-28 | Cellectis | Meganuclease variants cleaving a dna target sequence from the beta-2-microglobulin gene and uses thereof |
| WO2008130629A2 (en) * | 2007-04-19 | 2008-10-30 | Codon Devices, Inc. | Engineered nucleases and their uses for nucleic acid assembly |
| WO2008152523A1 (en) | 2007-06-06 | 2008-12-18 | Cellectis | Meganuclease variants cleaving a dna target sequence from the mouse rosa26 locus and uses thereof |
| WO2009013622A2 (en) | 2007-07-23 | 2009-01-29 | Cellectis | Meganuclease variants cleaving a dna target sequence from the human hemoglobin beta gene and uses thereof |
| WO2009019614A2 (en) | 2007-08-03 | 2009-02-12 | Cellectis | Meganuclease variants cleaving a dna target sequence from the human interleukin-2 receptor gamma chain gene and uses thereof |
| WO2009059195A2 (en) * | 2007-10-31 | 2009-05-07 | Precision Biosciences | Rationally-designed single-chain meganucleases with non-palindromic recognition sequences |
| WO2009074842A1 (en) * | 2007-12-13 | 2009-06-18 | Cellectis | Improved chimeric meganuclease enzymes and uses thereof |
| WO2009095793A1 (en) | 2008-01-31 | 2009-08-06 | Cellectis | New i-crei derived single-chain meganuclease and uses thereof |
| WO2009154686A1 (en) * | 2008-05-28 | 2009-12-23 | Sangamo Biosciences, Inc. | Compositions for linking dna-binding domains and cleavage domains |
| WO2011064751A1 (en) * | 2009-11-27 | 2011-06-03 | Basf Plant Science Company Gmbh | Chimeric endonucleases and uses thereof |
| WO2012058458A2 (en) * | 2010-10-27 | 2012-05-03 | Cellectis Sa | Method for increasing the efficiency of double-strand break-induced mutagenesis |
-
2012
- 2012-04-05 WO PCT/US2012/032386 patent/WO2012138901A1/en not_active Ceased
Patent Citations (31)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US4179337A (en) | 1973-07-20 | 1979-12-18 | Davis Frank F | Non-immunogenic polypeptides |
| US4683195A (en) | 1986-01-30 | 1987-07-28 | Cetus Corporation | Process for amplifying, detecting, and/or-cloning nucleic acid sequences |
| US4683195B1 (en) | 1986-01-30 | 1990-11-27 | Cetus Corp | |
| US5006333A (en) | 1987-08-03 | 1991-04-09 | Ddi Pharmaceuticals, Inc. | Conjugates of superoxide dismutase coupled to high molecular weight polyalkylene glycols |
| WO2003078619A1 (en) | 2002-03-15 | 2003-09-25 | Cellectis | Hybrid and single chain meganucleases and use thereof |
| WO2004031346A2 (en) | 2002-09-06 | 2004-04-15 | Fred Hutchinson Cancer Research Center | Methods and compositions concerning designed highly-specific nucleic acid binding proteins |
| WO2004067736A2 (en) | 2003-01-28 | 2004-08-12 | Cellectis | Custom-made meganuclease and use thereof |
| WO2006097854A1 (en) | 2005-03-15 | 2006-09-21 | Cellectis | Heterodimeric meganucleases and use thereof |
| WO2006097784A1 (en) | 2005-03-15 | 2006-09-21 | Cellectis | I-crei meganuclease variants with modified specificity, method of preparation and uses thereof |
| WO2006097853A1 (en) | 2005-03-15 | 2006-09-21 | Cellectis | I-crei meganuclease variants with modified specificity, method of preparation and uses thereof |
| WO2007034262A1 (en) | 2005-09-19 | 2007-03-29 | Cellectis | Heterodimeric meganucleases and use thereof |
| WO2007047859A2 (en) * | 2005-10-18 | 2007-04-26 | Precision Biosciences | Rationally-designed meganucleases with altered sequence specificity and dna-binding affinity |
| WO2007049156A2 (en) | 2005-10-25 | 2007-05-03 | Cellectis | I-CreI HOMING ENDONUCLEASE VARIANTS HAVING NOVEL CLEAVAGE SPECIFICITY AND USE THEREOF |
| WO2007049095A1 (en) | 2005-10-25 | 2007-05-03 | Cellectis | Laglidadg homing endonuclease variants having mutations in two functional subdomains and use thereof |
| WO2007057781A2 (en) | 2005-10-25 | 2007-05-24 | Cellectis | Laglidadg homing endonuclease variants having mutations in two functional subdomains and use thereof. |
| WO2007060495A1 (en) | 2005-10-25 | 2007-05-31 | Cellectis | I-crei homing endonuclease variants having novel cleavage specificity and use thereof |
| WO2007093918A2 (en) | 2006-02-13 | 2007-08-23 | Cellectis | Meganuclease variants cleaving a dna target sequence from a xeroderma pigmentosum gene and uses thereof |
| WO2008010093A2 (en) | 2006-07-18 | 2008-01-24 | Cellectis | Meganuclease variants cleaving a dna target sequence from a rag gene and uses thereof |
| WO2008059382A2 (en) | 2006-11-14 | 2008-05-22 | Cellectis | Meganuclease variants cleaving a dna target sequence from the hprt gene and uses thereof |
| WO2008093249A2 (en) | 2007-02-01 | 2008-08-07 | Cellectis | Obligate heterodimer meganucleases and uses thereof |
| WO2008102274A2 (en) | 2007-02-20 | 2008-08-28 | Cellectis | Meganuclease variants cleaving a dna target sequence from the beta-2-microglobulin gene and uses thereof |
| WO2008130629A2 (en) * | 2007-04-19 | 2008-10-30 | Codon Devices, Inc. | Engineered nucleases and their uses for nucleic acid assembly |
| WO2008152523A1 (en) | 2007-06-06 | 2008-12-18 | Cellectis | Meganuclease variants cleaving a dna target sequence from the mouse rosa26 locus and uses thereof |
| WO2009013622A2 (en) | 2007-07-23 | 2009-01-29 | Cellectis | Meganuclease variants cleaving a dna target sequence from the human hemoglobin beta gene and uses thereof |
| WO2009019614A2 (en) | 2007-08-03 | 2009-02-12 | Cellectis | Meganuclease variants cleaving a dna target sequence from the human interleukin-2 receptor gamma chain gene and uses thereof |
| WO2009059195A2 (en) * | 2007-10-31 | 2009-05-07 | Precision Biosciences | Rationally-designed single-chain meganucleases with non-palindromic recognition sequences |
| WO2009074842A1 (en) * | 2007-12-13 | 2009-06-18 | Cellectis | Improved chimeric meganuclease enzymes and uses thereof |
| WO2009095793A1 (en) | 2008-01-31 | 2009-08-06 | Cellectis | New i-crei derived single-chain meganuclease and uses thereof |
| WO2009154686A1 (en) * | 2008-05-28 | 2009-12-23 | Sangamo Biosciences, Inc. | Compositions for linking dna-binding domains and cleavage domains |
| WO2011064751A1 (en) * | 2009-11-27 | 2011-06-03 | Basf Plant Science Company Gmbh | Chimeric endonucleases and uses thereof |
| WO2012058458A2 (en) * | 2010-10-27 | 2012-05-03 | Cellectis Sa | Method for increasing the efficiency of double-strand break-induced mutagenesis |
Non-Patent Citations (149)
| Title |
|---|
| "Gene Transfer Vectors For Mammalian Cells", vol. 185, 1987, COLD SPRING HARBOR LABORATORY, article "Gene Expression Technology" |
| "Immobilized Cells And Enzymes", 1986, IRL PRESS |
| "Manipulating the Mouse Embryo", 1986, COLD SPRING HARBOR LABORATORY PRESS |
| ARIMONDO, P. B.; C. J. THOMAS ET AL.: "Exploring the cellular activity of camptothecin-triple- helix-forming oligonucleotide conjugates.", MOL CELL BIOL, vol. 26, no. 1, 2006, pages 324 - 33, XP055108516 |
| ARNOULD, S.; C. DELENDA ET AL.: "The I-Crel meganuclease and its engineered derivatives: applications from cell modification to gene therapy gzq083 [pii] 10.1093/protein/gzq083.", PROTEIN ENGINEERING, DESIGN & SELECTION : PEDS, vol. 24, no. 1-2, 2011, pages 27 - 31 |
| ARNOULD, S.; C. PEREZ ET AL.: "Engineered I-Crel derivatives cleaving sequences from the human XPC gene can induce highly efficient gene correction in mammalian cells.", JOURNAL OF MOLECULAR BIOLOGY, vol. 371, no. 1, 2007, pages 49 - 65, XP022145891, DOI: doi:10.1016/j.jmb.2007.04.079 |
| ARNOULD, S.; P. CHAMES ET AL.: "Engineering of large numbers of highly specific homing endonucleases that induce recombination on novel DNA targets", JOURNAL OF MOLECULAR BIOLOGY, vol. 355, 2006, pages 443 - 58, XP024950505, DOI: doi:10.1016/j.jmb.2005.10.065 |
| ARNOULD, S.; P. CHAMES ET AL.: "Engineering of large numbers of highly specific homing endonucleases that induce recombination on novel DNA targets.", J MOL BIOL, vol. 355, no. 3, 2006, pages 443 - 58, XP024950505, DOI: doi:10.1016/j.jmb.2005.10.065 |
| ASHWORTH, J.; J. J. HAVRANEK ET AL.: "Computational redesign of endonuclease DNA binding and cleavage specificity.", NATURE, vol. 441, no. 7093, 2006, pages 656 - 9, XP002500320, DOI: doi:10.1038/nature04818 |
| B. D. HARRIES & S. J. HIGGINS: "Nucleic Acid Hybridization", 1984 |
| B. D. HARRIES & S. J. HIGGINS: "Transcription And Translation", 1984 |
| B. PERBAL: "A Practical Guide To Molecular Cloning", 1984 |
| BEDAYAT, B.; A. ABDOLMOHAMADI ET AL.: "Sequence-specific correction of genomic hypoxanthine-guanine phosphoribosyl transferase mutations in lymphoblasts by small fragment homologous replacement10.1089/oli.2009.0205.", OLIGONUCLEOTIDES, vol. 20, no. 1, 2010, pages 7 - 16 |
| BENNARDO, N.; A. CHENG ET AL.: "Alternative-NHEJ is a mechanistically distinct pathway of mammalian chromosome break repair.", PLOS GENET, vol. 4, no. 6, 2008, pages E1000110, XP002680904, DOI: doi:10.1371/JOURNAL.PGEN.1000110 |
| BENNARDO, N.; A. GUNN ET AL.: "Limiting the persistence of a chromosome break diminishes its mutagenic potential.", PLOS GENET, vol. 5, no. 10, 2009, pages E1000683, XP002683881, DOI: doi:10.1371/JOURNAL.PGEN.1000683 |
| BOCH, J.; H. SCHOLZE ET AL.: "Breaking the code of DNA binding specificity of TAL-type III effectors 1178811 [pii] 10.1126/science.ll78811.", SCIENCE, vol. 326, no. 5959, 2009, pages 1509 - 12 |
| BOCH, J.; H. SCHOLZE ET AL.: "Breaking the code of DNA binding specificity of TAL-type III effectors.", SCIENCE, vol. 326, no. 5959, 2009, pages 1509 - 12 |
| BOLDUC, J. M.; P. C. SPIEGEL ET AL.: "Structural and biochemical analyses of DNA and RNA binding by a bifunctional homing endonuclease and group I intron splicing factor.", GENES DEV, vol. 17, no. 23, 2003, pages 2875 - 88 |
| BUIS, J.; Y. WU ET AL.: "Mrell nuclease activity has essential roles in DNA repair and genomic stability distinct from ATM activation.", CELL, vol. 135, no. 1, 2008, pages 85 - 96 |
| CAPECCHI, M. R.: "Generating mice with targeted mutations 10.1038/nm1001-1086 nm1001-1086 [pii].", NATURE MEDICINE, vol. 7, no. 10, 2001, pages 1086 - 90 |
| CARROLL, D.: "Progress and prospects: zinc-finger nucleases as gene therapy agents gt2008145 [pii] 10.1038/gt.2008.145.", GENE THERAPY, vol. 15, no. 22, 2008, pages 1463 - 8 |
| CHAMES, P.; J. C. EPINAT ET AL.: "In vivo selection of engineered homing endonucleases using double-strand break induced homologous recombination.", NUCLEIC ACIDS RES, vol. 33, no. 20, 2005, pages E178, XP002378874 |
| CHEVALIER, B. S.; B. L. STODDARD: "Homing endonucleases: structural and functional insight into the catalysts of intron/intein mobility.", NUCLEIC ACIDS RES, vol. 29, no. 18, 2001, pages 3757 - 74, XP002383228, DOI: doi:10.1093/nar/29.18.3757 |
| CHEVALIER, B. S.; R. J. MONNAT, JR. ET AL.: "The homing endonuclease I-Crel uses three metals, one of which is shared between the two active sites.", NAT STRUCT BIOL, vol. 8, no. 4, 2001, pages 312 - 6, XP008015664, DOI: doi:10.1038/86181 |
| CHEVALIER, B. S.; T. KORTEMME ET AL.: "Design, activity, and structure of a highly specific artificial endonuclease.", MOL CELL, vol. 10, no. 4, 2002, pages 895 - 905, XP002248750, DOI: doi:10.1016/S1097-2765(02)00690-1 |
| CHEVALIER, B.; M. TURMEL ET AL.: "Flexible DNA target site recognition by divergent homing endonuclease isoschizomers I-Crel and I-Msol.", J MOL BIOL, vol. 329, no. 2, 2003, pages 253 - 69 |
| CHOO, Y.; A. KLUG: "Selection of DNA binding sites for zinc fingers using rationally randomized DNA reveals coded interactions.", PROCEEDINGS OF THE NATIONAL ACADEMY OF SCIENCES OF THE UNITED STATES OF AMERICA, vol. 91, no. 23, 1994, pages 11168 - 72, XP002075339, DOI: doi:10.1073/pnas.91.23.11168 |
| CHOO, Y.; A. KLUG: "Toward a code for the interactions of zinc fingers with DNA: selection of randomized fingers displayed on phage.", PROCEEDINGS OF THE NATIONAL ACADEMY OF SCIENCES OF THE UNITED STATES OF AMERICA, vol. 91, no. 23, 1994, pages 11163 - 7, XP002075340, DOI: doi:10.1073/pnas.91.23.11163 |
| CHOULIKA, A.; A. PERRIN ET AL.: "Induction of homologous recombination in mammalian chromosomes by using the I-Scel system of Saccharomyces cerevisiae.", MOL CELL BIOL, vol. 15, no. 4, 1995, pages 1968 - 73, XP000572150 |
| CHRISTIAN, M.; T. CERMAK ET AL.: "Targeting DNA double-strand breaks with TAL effector nucleases genetics.110.120717 [pii] 10.1534/genetics.110.120717.", GENETICS, vol. 186, no. 2, 2010, pages 757 - 61 |
| CHRISTIAN, M.; T. CERMAK ET AL.: "Targeting DNA double-strand breaks with TAL effector nucleases.", GENETICS, vol. 186, no. 2, 2010, pages 757 - 61, XP002632806, DOI: doi:10.1534/GENETICS.110.120717 |
| COFFIN, J. M. ET AL.: "Fundamental Virology, Third Edition,", 1996, LIPPINCOTT-RAVEN PUBLISHERS,, article "Retroviridae: The viruses and their replication" |
| COST, G. J.; Y. FREYVERT ET AL.: "BAK and BAX deletion using zinc-finger nucleases yields apoptosis-resistant CHO cells 10.1002/bit.22541.", BIOTECHNOLOGY AND BIOENGINEERING, vol. 105, no. 2, 2010, pages 330 - 40, XP055004494, DOI: doi:10.1002/bit.22541 |
| D. M. WEIR AND C. C. BLACKWELL,: "Handbook Of Experimental Immunology", vol. I-IV, 1986 |
| DELACOTE, F.; B. S. LOPEZ: "Importance of the cell cycle phase for the choice of the appropriate DSB repair pathway, for genome stability maintenance: the trans-S double-strand break repair model 5149 [pii].", CELL CYCLE, vol. 7, no. 1, 2008, pages 33 - 8 |
| DOUDEVA, L. G.; H. HUANG ET AL.: "Crystal structural analysis and metal-dependent stability and activity studies of the CoIE7 endonuclease domain in complex with DNA/Zn2+ or inhibitor/Ni2+ 15/2/269 [pii] 10.1110/ps.051903406.", PROTEIN SCIENCE : A PUBLICATION OF THE PROTEIN SOCIETY, vol. 15, no. 2, 2006, pages 269 - 80 |
| DOYON, J. B.; V. PATTANAYAK ET AL.: "Directed evolution and substrate specificity profile of homing endonuclease I-Scel.", JOURNAL OF THE AMERICAN CHEMICAL SOCIETY, vol. 128, no. 7, 2006, pages 2477 - 84, XP002611223, DOI: doi:10.1021/JA0575191 |
| DOYON, Y.; J. M. MCCAMMON ET AL.: "Heritable targeted gene disruption in zebrafish using designed zinc-finger nucleases nbtl409 [pii] 10.1038/nbt1409.", NATURE BIOTECHNOLOGY, vol. 26, no. 6, 2008, pages 702 - 8, XP002512110, DOI: doi:10.1038/NBT1409 |
| EASTBERG, J. H.; J. EKLUND ET AL.: "Mutability of an HNH nuclease imidazole general base and exchange of a deprotonation mechanism.", BIOCHEMISTRY, vol. 46, no. 24, 2007, pages 7215 - 25 |
| EISENSCHMIDT, K.; T. LANIO: "Developing a programmed restriction endonuclease for highly specific DNA cleavage.", NUCLEIC ACIDS RES, vol. 33, no. 22, 2005, pages 7039 - 47, XP002383230, DOI: doi:10.1093/nar/gki1009 |
| ELROD-ERICKSON, M.; M. A. ROULD ET AL.: "Zif268 protein-DNA complex refined at 1.6 A: a model system for understanding zinc finger-DNA interactions.", STRUCTURE, vol. 4, no. 10, 1996, pages 1171 - 80 |
| EPINAT, J. C.; S. ARNOULD ET AL.: "A novel engineered meganuclease induces homologous recombination in yeast and mammalian cells.", NUCLEIC ACIDS RES, vol. 31, no. 11, 2003, pages 2952 - 62 |
| FRANK, K. M.; J. M. SEKIGUCHI ET AL.: "Late embryonic lethality and impaired V(D)J recombination in mice lacking DNA ligase IV 10.1038/24172.", NATURE, vol. 396, no. 6707, 1998, pages 173 - 7 |
| FREDERICK M. AUSUBEL: "Current Protocols in Molecular Biology", 2000, WILEY AND SON INC |
| GALETTO, R.; P. DUCHATEAU ET AL.: "Targeted approaches for gene therapy and the emergence of engineered meganucleases 10.1517/14712590903213669.", EXPERT OPINION ON BIOLOGICAL THERAPY, vol. 9, no. 10, 2009, pages 1289 - 303, XP009137203, DOI: doi:10.1517/14712590903213669 |
| GAO, H.; J. SMITH ET AL.: "Heritable targeted mutagenesis in maize using a designed endonuclease TPJ4041 [pii] 10.1111/j.1365-313X.2009.04041.x.", THE PLANT FOR CELL AND MOLECULAR BIOLOGY, vol. 61, no. 1, 2010, pages 176 - 87 |
| GAO, Y.; Y. SUN ET AL.: "A critical role for DNA end-joining proteins in both lymphogenesis and neurogenesis S0092-8674(00)81714-6 [pii].", CELL, vol. 95, no. 7, 1998, pages 891 - 902 |
| GEURTS, A. M.; G. J. COST ET AL.: "Knockout rats via embryo microinjection of zinc-finger nucleases 325/5939/433 [pii] 10.1126/science.1172447.", SCIENCE, vol. 325, no. 5939, 2009, pages 433 |
| GIMBLE, F. S.; C. M. MOURE ET AL.: "Assessing the plasticity of DNA target site recognition of the PI-Scel homing endonuclease using a bacterial two-hybrid selection system.", J MOL BIOL, vol. 334, no. 5, 2003, pages 993 - 1008, XP004474968, DOI: doi:10.1016/j.jmb.2003.10.013 |
| GREISMAN, H. A.; C. O. PABO: "A general strategy for selecting high-affinity zinc finger proteins for diverse DNA target sites.", SCIENCE, vol. 275, no. 5300, 1997, pages 657 - 61, XP002167950, DOI: doi:10.1126/science.275.5300.657 |
| GRUENERT, D. C.; E. BRUSCIA ET AL.: "Sequence-specific modification of genomic DNA by small DNA fragments 10.1172/JC119773 112/5/637 [pii].", THE JOURNAL OF CLINICAL INVESTIGATION, vol. 112, no. 5, 2003, pages 637 - 41, XP002987522, DOI: doi:10.1172/JCI200319773 |
| GUIROUILH-BARBAT, J.; E. RASS ET AL.: "Defects in XRCC4 and KU80 differentially affect the joining of distal nonhomologous ends 0708541104 [pii] 10.1073/pnas.0708541104.", PROCEEDINGS OF THE NATIONAL ACADEMY OF SCIENCES OF THE UNITED STATES OF AMERICA, vol. 104, no. 52, 2007, pages 20902 - 7 |
| GUIROUILH-BARBAT, J.; E. RASS ET AL.: "Defects in XRCC4 and KU80 differentially affect the joining of distal nonhomologous ends.", PROC NATL ACAD SCI U S A, vol. 104, no. 52, 2007, pages 20902 - 7 |
| GUIROUILH-BARBAT, J.; S. HUCK ET AL.: "Impact of the KU80 pathway on NHEJ-induced genome rearrangements in mammalian cells 10.1016/j.molcel.2004.05.008 S1097276504002916 [pii].", MOLECULAR CELL, vol. 14, no. 5, 2004, pages 611 - 23 |
| GUIROUILH-BARBAT, J.; S. HUCK ET AL.: "Impact of the KU80 pathway on NHEJ-induced genome rearrangements in mammalian cells.", MOL CELL, vol. 14, no. 5, 2004, pages 611 - 23 |
| GURLEBECK, D.; B. SZUREK ET AL.: "Dimerization of the bacterial effector protein AvrBs3 in the plant cell cytoplasm prior to nuclear import TPJ2370 [pii] 10.1111/j.1365-313X.2005.02370.x.", THE PLANT JOURNAL: FOR CELL AND MOLECULAR BIOLOGY, vol. 42, no. 2, 2005, pages 175 - 87, XP002555457, DOI: doi:10.1111/J.1365-313X.2005.02370.X |
| HABER, J. E.: "Alternative endings 0711334105 [pii] 10.1073/pnas.0711334105.", PROCEEDINGS OF THE NATIONAL ACADEMY OF SCIENCES OF THE UNITED STATES OF AMERICA, vol. 105, no. 2, 2008, pages 405 - 6 |
| HABER, J.: "Partners and pathwaysrepairing a double-strand break.", TRENDS GENET., vol. 16, no. 6, 2000, pages 259 - 264, XP004200111, DOI: doi:10.1016/S0168-9525(00)02022-9 |
| HARTSUIKER, E.; K. MIZUNO ET AL.: "CtplCtlP and Rad32Mrell nuclease activity are required for Recl2Spoll removal, but Recl2Spoll removal is dispensable for other MRN-dependent meiotic functions.", MOL CELL BIOL, vol. 29, no. 7, 2009, pages 1671 - 81 |
| HINNEN, A.; J. B. HICKS ET AL.: "Transformation of yeast.", PROCEEDINGS OF THE NATIONAL ACADEMY OF SCIENCES OF THE UNITED STATES OF AMERICA, vol. 75, no. 4, 1978, pages 1929 - 33, XP002336016, DOI: doi:10.1073/pnas.75.4.1929 |
| HIRATA, R.; J. CHAMBERLAIN ET AL.: "Targeted transgene insertion into human chromosomes by adeno-associated virus vectors 10.1038/nbt0702-735 nbt0702-735 [pii].", NATURE BIOTECHNOLOGY, vol. 20, no. 7, 2002, pages 735 - 8 |
| HUANG, H.; H. S. YUAN: "The conserved asparagine in the HNH motif serves an important structural role in metal finger endonucleases.", JOURNAL OF MOLECULAR BIOLOGY, vol. 368, no. 3, 2007, pages 812 - 21, XP022020040, DOI: doi:10.1016/j.jmb.2007.02.044 |
| ICHIYANAGI, K.; Y. ISHINO ET AL.: "Crystal structure of an archaeal intein-encoded homing endonuclease PI-Pful.", J MOL BIOL, vol. 300, no. 4, 2000, pages 889 - 901 |
| INOUE, N.; R. DONG ET AL.: "Introduction of single base substitutions at homologous chromosomal sequences by adeno-associated virus vectors 10.1006/mthe.2001.0283 S1525-0016(01)90283-7 [pii].", MOLECULAR THERAPY : THE JOURNAL OF THE AMERICAN SOCIETY OF GENE THERAPY, vol. 3, no. 4, 2001, pages 526 - 30 |
| ISATAN, M.; Y. CHOO: "Rapid, high-throughput engineering of sequence-specific zinc finger DNA-binding proteins 50076-6879(01)40444-7 [pii].", METHODS IN ENZYMOLOGY, vol. 340, 2001, pages 593 - 609, XP009163869, DOI: doi:10.1016/S0076-6879(01)40444-7 |
| J. ABELSON AND M. SIMON: "Methods In ENZYMOLOGY", vol. 154, 155, ACADEMIC PRESS, INC. |
| KALISH, J. M.; P. M. GLAZER: "Targeted genome modification via triple helix formation.", ANN N Y ACAD SCI, vol. 1058, 2005, pages 151 - 61, XP055001413, DOI: doi:10.1196/annals.1359.023 |
| KIM, H. J.; H. J. LEE ET AL.: "Targeted genome editing in human cells with zinc finger nucleases constructed via modular assembly gr.089417.108 [pii] 10.1101/gr.089417.108.", GENOME RESEARCH, vol. 19, no. 7, 2009, pages 1279 - 88, XP055143646, DOI: doi:10.1101/gr.089417.108 |
| KIM, Y. G.; J. CHA ET AL.: "Hybrid restriction enzymes: zinc finger fusions to Fok I cleavage domain.", PROCEEDINGS OF THE NATIONAL ACADEMY OF SCIENCES OF THE UNITED STATES OF AMERICA, vol. 93, no. 3, 1996, pages 1156 - 60, XP002116423, DOI: doi:10.1073/pnas.93.3.1156 |
| KU, W. Y.; Y. W. LIU ET AL.: "The zinc ion in the HNH motif of the endonuclease domain of colicin E7 is not required for DNA binding but is essential for DNA hydrolysis.", NUCLEIC ACIDS RESEARCH, vol. 30, no. 7, 2002, pages 1670 - 8 |
| LANDTHALER, M.; B. W. SHEN ET AL.: "I-Basl and I-Hmul: two phage intron-encoded endonucleases with homologous DNA recognition sequences but distinct DNA specificities.", JOURNAL OF MOLECULAR BIOLOGY, vol. 358, no. 4, 2006, pages 1137 - 51 |
| LANDTHALER, M.; D. A. SHUB: "The nicking homing endonuclease I-Basl is encoded by a group I intron in the DNA polymerase gene of the Bacillus thuringiensis phage Bastille.", NUCLEIC ACIDS RESEARCH, vol. 31, no. 12, 2003, pages 3071 - 7 |
| LANDTHALER, M.; N. C. LAU ET AL.: "Group I intron homing in Bacillus phages SP01 and SP82: a gene conversion event initiated by a nicking homing endonuclease.", JOURNAL OF BACTERIOLOGY, vol. 186, no. 13, 2004, pages 4307 - 14 |
| LANDTHALER, M.; U. BEGLEY ET AL.: "Two self-splicing group I introns in the ribonucleotide reductase large subunit gene of Staphylococcus aureus phage Twort.", NUCLEIC ACIDS RESEARCH, vol. 30, no. 9, 2002, pages 1935 - 43 |
| LEE, S. E.; F. PAQUES ET AL.: "Role of yeast SIR genes and mating type in directing DNA double-strand breaks to homologous and non-homologous repair paths.", CURR BIOL, vol. 9, no. 14, 1999, pages 767 - 70 |
| LI, T.; S. HUANG ET AL.: "TAL nucleases (TALNs): hybrid proteins composed of TAL effectors and Fokl DNA-cleavage domain gkq704 [pii] 10.1093/nar/gkq704.", NUCLEIC ACIDS RESEARCH, vol. 39, no. 1, 2011, pages 359 - 72 |
| LI, T.; S. HUANG ET AL.: "TAL nucleases (TALNs): hybrid proteins composed of TAL effectors and Fokl DNA-cleavage domain.", NUCLEIC ACIDS RES, vol. 39, no. 1, 2010, pages 359 - 72 |
| LIANG, F.; M. HAN ET AL.: "Homology-directed repair is a major double-strand break repair pathway in mammalian cells.", PROCEEDINGS OF THE NATIONAL ACADEMY OF SCIENCES OF THE UNITED STATES OF AMERICA, vol. 95, no. 9, 1998, pages 5172 - 7, XP055024260, DOI: doi:10.1073/pnas.95.9.5172 |
| LIU, P. Q.; E. M. CHAN ET AL.: "Generation of a triple-gene knockout mammalian cell line using engineered zinc-finger nucleases 10.1002/bit.22654.", BIOTECHNOLOGY AND BIOENGINEERING, vol. 106, no. 1, 2010, pages 97 - 105, XP002689346, DOI: doi:10.1002/bit.22654 |
| LIU, Q.; V. DERBYSHIRE ET AL.: "Distance determination by GIY-YIG intron endonucleases: discrimination between repression and cleavage functions.", NUCLEIC ACIDS RESEARCH, vol. 34, no. 6, 2006, pages 1755 - 64, XP002691037, DOI: doi:10.1093/nar/gkl079 |
| LLOYD, A.; C. L. PLAISIER ET AL.: "Targeted mutagenesis using zinc-finger nucleases in Arabidopsis 0409339102 [pii] 10.1073/pnas.0409339102.", PROCEEDINGS OF THE NATIONAL ACADEMY OF SCIENCES OF THE UNITED STATES OF AMERICA, vol. 102, no. 6, 2005, pages 2232 - 7 |
| M. J. GAIT: "Oligonucleotide Synthesis", 1984 |
| MAEDER, M. L.; S. THIBODEAU-BEGANNY ET AL.: "Rapid ''open-source'' engineering of customized zinc-finger nucleases for highly efficient gene modification 51097-2765(08)00461-9 [pii] 10.1016/j.molcel.2008.06.016.", MOLECULAR CELL, vol. 31, no. 2, 2008, pages 294 - 301 |
| MAHFOUZ, M. M.; L. LI ET AL.: "De novo-engineered transcription activator-like effector (TALE) hybrid nuclease with novel DNA binding specificity creates double-strand breaks 1019533108 [pii] 10.1073/pnas.1019533108.", PROCEEDINGS OF THE NATIONAL ACADEMY OF SCIENCES OF THE UNITED STATES OF AMERICA, vol. 108, no. 6, 2011, pages 2623 - 8 |
| MARCAIDA, M. J.; G. MUNOZ ET AL.: "Homing endonucleases: from basics to therapeutic applications 10.1007/s00018-009-0188-y.", CELLULAR AND MOLECULAR LIFE SCIENCES: CMLS, vol. 67, no. 5, 2010, pages 727 - 48, XP019797908 |
| MASHIMO, T.; A. TAKIZAWA ET AL.: "Generation of knockout rats with X-linked severe combined immunodeficiency (X-SCID) using zinc-finger nucleases 10.1371/journal.pone.0008870.", PLOS ONE, vol. 5, no. 1, 2010, pages E8870 |
| MAYER AND WALKER,: "Immunochemical Methods In Cell And Molecular Biology", 1987, ACADEMIC PRESS |
| MCCONNELL SMITH, A.; R. TAKEUCHI ET AL.: "Generation of a nicking enzyme that stimulates site-specific gene conversion from the I-Anil LAGLIDADG homing endonuclease 0810588106 [pii] 10.1073/pnas.0810588106.", PROCEEDINGS OF THE NATIONAL ACADEMY OF SCIENCES OF THE UNITED STATES OF AMERICA, vol. 106, no. 13, 2009, pages 5099 - 104 |
| MCVEY, M.; S. E. LEE: "MMEJ repair of double-strand breaks (director's cut): deleted sequences and alternative endings 50168-9525(08)00229-1 [pii] 10.1016/j.tig.2008.08.007.", TRENDS IN GENETICS : TIG, vol. 24, no. 11, 2008, pages 529 - 38 |
| MENORET, S.; A. L. ISCACHE ET AL.: "Characterization of immunoglobulin heavy chain knockout rats 10.1002/eji.201040939.", EUROPEAN JOURNAL OF IMMUNOLOGY, vol. 40, no. 10, 2010, pages 2932 - 41, XP055129843, DOI: doi:10.1002/eji.201040939 |
| MIDON, M.; P. SCHAFER ET AL.: "Mutational and biochemical analysis of the DNA-entry nuclease EndA from Streptococcus pneumoniae gkq802 [pii] 10.1093/nar/gkq802.", NUCLEIC ACIDS RESEARCH, vol. 39, no. 2, 2011, pages 623 - 34 |
| MILLER, J. C.; S. TAN ET AL.: "A TALE nuclease architecture for efficient genome editing nbt.1755 [pii] 10.1038/nbt.1755.", NATURE BIOTECHNOLOGY, vol. 29, no. 2, 2011, pages 143 - 8 |
| MIMITOU, E. P.; L. S. SYMINGTON: "Sae2, Exol and Sgsl collaborate in DNA double-strand break processing.", NATURE, vol. 455, no. 7214, 2008, pages 770 - 4 |
| MOORE, I.; M. SAMALOVA ET AL.: "Transactivated and chemically inducible gene expression in plants.", PLANT, vol. 45, no. 4, 2006, pages 651 - 83, XP002716440, DOI: doi:10.1111/J.1365-313X.2006.02660.X |
| MOORE, J. K.; J. E. HABER: "Cell cycle and genetic requirements of two pathways of nonhomologous end-joining repair of double-strand breaks in Saccharomyces cerevisiae.", MOL CELL BIOL, vol. 16, no. 5, 1996, pages 2164 - 73, XP055092963 |
| MOSCOU, M. J.; A. J. BOGDANOVE: "A simple cipher governs DNA recognition by TAL effectors 1178817 [pii] 10.1126/science.1178817.", SCIENCE, vol. 326, no. 5959, 2009, pages 1501 |
| MOSCOU, M. J.; A. J. BOGDANOVE: "A simple cipher governs DNA recognition by TAL effectors.", SCIENCE, vol. 326, no. 5959, 2009, pages 1501, XP002599998 |
| MOURE, C. M.; F. S. GIMBLE ET AL.: "Crystal structure of the intein homing endonuclease PI-Scel bound to its recognition sequence.", NAT STRUCT BIOL, vol. 9, no. 10, 2002, pages 764 - 70 |
| MOURE, C. M.; F. S. GIMBLE ET AL.: "The crystal structure of the gene targeting homing endonuclease I-Scel reveals the origins of its target site specificity.", J MOL BIOL, vol. 334, no. 4, 2003, pages 685 - 95, XP004473365, DOI: doi:10.1016/j.jmb.2003.09.068 |
| NIMONKAR, A. V.; J. GENSCHEL ET AL.: "BLM-DNA2-RPA-MRN and EX01-BLM-RPA-MRN constitute two DNA end resection machineries for human DNA break repair.", GENES DEV, vol. 25, no. 4, 2011, pages 350 - 62 |
| NIU, Y.; K. TENNEY ET AL.: "Engineering variants of the I-Scel homing endonuclease with strand-specific and site-specific DNA-nicking activity 50022-2836(08)00840-1 [pii] 10.1016/j.jmb.2008.07.010.", JOURNAL OF MOLECULAR BIOLOGY, vol. 382, no. 1, 2008, pages 188 - 202 |
| ORR-WEAVER, T. L.; J. W. SZOSTAK ET AL.: "Genetic applications of yeast transformation with linear and gapped plasmids.", METHODS IN ENZYMOLOGY, vol. 101, 1983, pages 228 - 45 |
| ORR-WEAVER, T. L.; J. W. SZOSTAK ET AL.: "Yeast transformation: a model system for the study of recombination.", PROCEEDINGS OF THE NATIONAL ACADEMY OF SCIENCES OF THE UNITED STATES OF AMERICA, vol. 78, no. 10, 1981, pages 6354 - 8 |
| PABO, C. 0.; E. PEISACH ET AL.: "Design and selection of novel Cys2His2 zinc finger proteins 70/1/313 [pii] 10.1146/annurev.biochem.70.1.313.", ANNUAL REVIEW OF BIOCHEMISTRY, vol. 70, 2001, pages 313 - 40 |
| PADIDAM, M.: "Chemically regulated gene expression in plants.", CURR OPIN PLANT BIOL, vol. 6, no. 2, 2003, pages 169 - 77, XP002559217, DOI: doi:10.1016/S1369-5266(03)00005-0 |
| PÂQUES FRÉDÉRIC ET AL: "Meganucleases and DNA double-strand break-induced recombination: perspectives for gene therapy", CURRENT GENE THERAPY, BENTHAM SCIENCE PUBLISHERS LTD, NL, vol. 7, no. 1, 1 February 2007 (2007-02-01), pages 49 - 66, XP002493370, ISSN: 1566-5232, DOI: 10.2174/156652307779940216 * |
| PAQUES, F.; J. E. HABER: "Multiple pathways of recombination induced by double-strand breaks in Saccharomyces cerevisiae.", MICROBIOL MOL BIOL REV, vol. 63, no. 2, 1999, pages 349 - 404 |
| PAQUES, F.; J. E. HABER: "Multiple pathways of recombination induced by double-strand breaks in Saccharomyces cerevisiae.", MICROBIOLOGY AND MOLECULAR BIOLOGY REVIEWS : MMBR, vol. 63, no. 2, 1999, pages 349 - 404 |
| PAQUES, F.; P. DUCHATEAU: "Meganucleases and DNA double-strand break-induced recombination: perspectives for gene therapy.", CURRENT GENE THERAPY, vol. 7, no. 1, 2007, pages 49 - 66, XP002493370, DOI: doi:10.2174/156652307779940216 |
| PEREZ, E. E.; J. WANG ET AL.: "Establishment of HIV-1 resistance in CD4+ T cells by genome editing using zinc-finger nucleases nbt1410 [pii] 10.1038/nbt1410.", NATURE BIOTECHNOLOGY, vol. 26, no. 7, 2008, pages 808 - 16, XP055024363, DOI: doi:10.1038/nbt1410 |
| PIERCE, A. J.; P. HU ET AL.: "Ku DNA end-binding protein modulates homologous repair of double-strand breaks in mammalian cells.", GENES DEV, vol. 15, no. 24, 2001, pages 3237 - 42 |
| PINGOUD, A.; G. H. SILVA: "Precision genome surgery.", NAT BIOTECHNOL, vol. 25, no. 7, 2007, pages 743 - 4, XP002657044, DOI: doi:10.1038/nbt0707-743 |
| PORTEUS, M. H.; D. CARROLL: "Gene targeting using zinc finger nucleases.", NAT BIOTECHNOL, vol. 23, no. 8, 2005, pages 967 - 73, XP002467422, DOI: doi:10.1038/nbt1125 |
| POTENZA C ET AL., CELL DEV BIOL, vol. 40, 2004, pages 1 - 22 |
| R. I. FRESHNEY,: "Culture Of Animal Cells", 1987, ALAN R. LISS, INC. |
| RAMIREZ, C. L.; J. E. FOLEY ET AL.: "Unexpected failure rates for modular assembly of engineered zinc fingers nmeth0508-374 [pii] 10.1038/nmeth0508-374.", NATURE METHODS, vol. 5, no. 5, 2008, pages 374 - 5 |
| ROSEN, L. E.; H. A. MORRISON ET AL.: "Homing endonuclease I-Crel derivatives with novel DNA target specificities.", NUCLEIC ACIDS RESEARCH, vol. 34, no. 17, 2006, pages 4791 - 800 |
| ROTHSTEIN, R. J.: "One-step gene disruption in yeast.", METHODS IN ENZYMOLOGY, vol. 101, 1983, pages 202 - 11, XP009084053, DOI: doi:10.1016/0076-6879(83)01015-0 |
| ROUET, P.; F. SMIH ET AL.: "Expression of a site-specific endonuclease stimulates homologous recombination in mammalian cells.", PROC NATL ACAD SCI U S A, vol. 91, no. 13, 1994, pages 6064 - 8, XP002005205, DOI: doi:10.1073/pnas.91.13.6064 |
| ROUET, P.; F. SMIH ET AL.: "Introduction of double-strand breaks into the genome of mouse cells by expression of a rare-cutting endonuclease.", MOL CELL BIOL, vol. 14, no. 12, 1994, pages 8096 - 106, XP002005200 |
| RUSSELL, D. W.; R. K. HIRATA: "Human gene targeting by viral vectors 10.1038/ng0498-325.", NATURE GENETICS, vol. 18, no. 4, 1998, pages 325 - 30, XP001105541, DOI: doi:10.1038/ng0498-325 |
| SAMBROOK ET AL.: "Molecular Cloning: A Laboratory Manual", 2001, COLD SPRING HARBOR LABORATORY PRESS |
| SANGIUOLO, F.; M. L. SCALDAFERRI ET AL.: "Cftr gene targeting in mouse embryonic stem cells mediated by Small Fragment Homologous Replacement (SFHR) 2904 [pii", FRONTIERS IN BIOSCIENCE : A IOURNAL AND VIRTUAL LIBRARY, vol. 13, 2008, pages 2989 - 99 |
| SANTIAGO, Y.; E. CHAN ET AL.: "Targeted gene knockout in mammalian cells by using engineered zinc-finger nucleases 0800940105 [pii] 10.1073/pnas.0800940105.", PROCEEDINGS OF THE NATIONAL ACADEMY OF SCIENCES OF THE UNITED STATES OF AMERICA, vol. 105, no. 15, 2008, pages 5809 - 14 |
| SARTORI, A. A.; C. LUKAS ET AL.: "Human CtlP promotes DNA end resection.", NATURE, vol. 450, no. 7169, 2007, pages 509 - 14 |
| SELIGMAN, L. M.; K. M. CHISHOLM ET AL.: "Mutations altering the cleavage specificity of a homing endonuclease.", NUCLEIC ACIDS RESEARCH, vol. 30, no. 17, 2002, pages 3870 - 9, XP002282592, DOI: doi:10.1093/nar/gkf495 |
| SELIGMAN, L. M.; K. M. STEPHENS ET AL.: "Genetic analysis of the Chlamydomonas reinhardtii I-Crel mobile intron homing system in Escherichia coli.", GENETICS, vol. 147, no. 4, 1997, pages 1653 - 64, XP055039708 |
| SHEN, B. W.; M. LANDTHALER ET AL.: "DNA binding and cleavage by the HNH homing endonuclease I-Hmul.", JOURNAL OF MOLECULAR BIOLOGY, vol. 342, no. 1, 2004, pages 43 - 56, XP004844890, DOI: doi:10.1016/j.jmb.2004.07.032 |
| SHUKLA, V. K.; Y. DOYON ET AL.: "Precise genome modification in the crop species Zea mays using zinc-finger nucleases nature07992 [pii] 10.1038/nature07992.", NATURE, vol. 459, no. 7245, 2009, pages 437 - 41, XP002626698, DOI: doi:10.1038/NATURE07992 |
| SILVA, G. H.; J. Z. DALGAARD ET AL.: "Crystal structure of the thermostable archaeal intron-encoded endonuclease I-Dmol.", J MOL BIOL, vol. 286, no. 4, 1999, pages 1123 - 36, XP004462690, DOI: doi:10.1006/jmbi.1998.2519 |
| SIMON, P.; F. CANNATA ET AL.: "Sequence-specific DNA cleavage mediated by bipyridine polyamide conjugates.", NUCLEIC ACIDS RES, vol. 36, no. 11, 2008, pages 3531 - 8 |
| SMITH, J.; J. M. BERG ET AL.: "A detailed study of the substrate specificity of a chimeric restriction enzyme gkcl39 [pii].", NUCLEIC ACIDS RESEARCH, vol. 27, no. 2, 1999, pages 674 - 81 |
| SMITH, J.; M. BIBIKOVA ET AL.: "Requirements for double-strand cleavage by chimeric restriction enzymes with zinc finger DNA-recognition domains.", NUCLEIC ACIDS RES, vol. 28, no. 17, 2000, pages 3361 - 9, XP002974228, DOI: doi:10.1093/nar/28.17.3361 |
| SMITH, J.; M. BIBIKOVA ET AL.: "Requirements for double-strand cleavage by chimeric restriction enzymes with zinc finger DNA-recognition domains.", NUCLEIC ACIDS RESEARCH, vol. 28, no. 17, 2000, pages 3361 - 9, XP002974228, DOI: doi:10.1093/nar/28.17.3361 |
| SMITH, J.; S. GRIZOT ET AL.: "A combinatorial approach to create artificial homing endonucleases cleaving chosen sequences.", NUCLEIC ACIDS RESEARCH, vol. 34, no. 22, 2006, pages E149, XP002559417, DOI: doi:10.1093/nar/gkl720 |
| SONODA, E.; H. HOCHEGGER ET AL.: "Differential usage of non-homologous end-joining and homologous recombination in double strand break repair.", DNA REPAIR (AMST), vol. 5, no. 9-10, 2006, pages 1021 - 9, XP024977137, DOI: doi:10.1016/j.dnarep.2006.05.022 |
| SPIEGEL, P. C.; B. CHEVALIER ET AL.: "The structure of I-Ceul homing endonuclease: Evolving asymmetric DNA recognition from a symmetric protein scaffold.", STRUCTURE, vol. 14, no. 5, 2006, pages 869 - 80 |
| STODDARD, B. L.: "Homing endonuclease structure and function.", QUARTERLY REVIEWS OF BIOPHYSICS, vol. 38, no. 1, 2005, pages 49 - 95, XP008065807, DOI: doi:10.1017/S0033583505004063 |
| STODDARD, B. L.; A. M. SCHARENBERG ET AL.: "Progress in Gene Therapy: Autologous and Cancer Stem Cell Gene Therapy.", vol. 3, 2007, WORLD SCIENTIFIC PUBLISHING CO. PTE. LTD., article "Advances in Engineering Homing Endonucleases for Gene Targeting: Ten Years After Structures", pages: 135 - 68 |
| SUGAWARA, N.; J. E. HABER: "Characterization of double-strand break-induced recombination: homology requirements and single-stranded DNA formation.", MOL CELL BIOL, vol. 12, no. 2, 1992, pages 563 - 75 |
| SUN, H.; D. TRECO ET AL.: "Extensive 3'-overhanging, single-stranded DNA associated with the meiosis-specific double-strand breaks at the ARG4 recombination initiation site.", CELL, vol. 64, no. 6, 1991, pages 1155 - 61, XP024245310, DOI: doi:10.1016/0092-8674(91)90270-9 |
| SUSSMAN, D.; M. CHADSEY ET AL.: "Isolation and characterization of new homing endonuclease specificities at individual target site positions.", JOURNAL OF MOLECULAR BIOLOGY, vol. 342, no. 1, 2004, pages 31 - 41, XP004844889, DOI: doi:10.1016/j.jmb.2004.07.031 |
| TAUBES, G.: "Gene therapy. The strange case of chimeraplasty 10.1126/science.298.5601.2116 298/5601/2116 [pii].", SCIENCE, vol. 298, no. 5601, 2002, pages 2116 - 20 |
| WANG, R.; X. ZHOU ET AL.: "Chemically regulated expression systems and their applications in transgenic plants.", TRANSGENIC RES, vol. 12, no. 5, 2003, pages 529 - 40, XP002395660, DOI: doi:10.1023/A:1025852307127 |
| WANG, Y. T.; J. D. WRIGHT ET AL.: "Redesign of high-affinity nonspecific nucleases with altered sequence preference 10.1021/ja907160r.", JOURNAL OF THE AMERICAN CHEMICAL SOCIETY, vol. 131, no. 47, 2009, pages 17345 - 53 |
| WHITE, C. I.; J. E. HABER: "Intermediates of recombination during mating type switching in Saccharomyces cerevisiae.", EMBO J, vol. 9, no. 3, 1990, pages 663 - 73 |
| YANG, M.; V. DJUKANOVIC ET AL.: "Targeted mutagenesis in the progeny of maize transgenic plants 10.1007/S11103-009-9499-5.", PLANT MOLECULAR BIOLOGY, vol. 70, no. 6, 2009, pages 669 - 79, XP019686576 |
| ZHAO, L.; R. P. BONOCORA ET AL.: "The restriction fold turns to the dark side: a bacterial homing endonuclease with a PD-(D/E)-XK motif.", THE EMBO JOURNAL, vol. 26, no. 9, 2007, pages 2432 - 42 |
| ZUO, J.; N. H. CHUA: "Chemical-inducible systems for regulated expression of plant genes.", CURR OPIN BIOTECHNOL, vol. 11, no. 2, 2000, pages 146 - 51, XP001039646, DOI: doi:10.1016/S0958-1669(00)00073-2 |
Cited By (11)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO2014118719A1 (en) * | 2013-02-01 | 2014-08-07 | Cellectis | Tevl chimeric endonuclease and their preferential cleavage sites |
| US10738289B2 (en) | 2013-02-01 | 2020-08-11 | Cellectis | Tevi chimeric endonuclease and their preferential cleavage sites |
| JP2019503198A (en) * | 2016-02-02 | 2019-02-07 | サンガモ セラピューティクス, インコーポレイテッド | Composition for linking a DNA binding domain and a cleavage domain |
| EP3411056A4 (en) * | 2016-02-02 | 2019-10-02 | Sangamo Therapeutics, Inc. | COMPOSITIONS FOR BINDING DNA BINDING DOMAINS AND CLEAVAGE DOMAINS |
| US10724020B2 (en) | 2016-02-02 | 2020-07-28 | Sangamo Therapeutics, Inc. | Compositions for linking DNA-binding domains and cleavage domains |
| EP3769775A3 (en) * | 2016-02-02 | 2021-03-17 | Sangamo Therapeutics, Inc. | Compositions for linking dna-binding domains and cleavage domains |
| JP7012650B2 (en) | 2016-02-02 | 2022-01-28 | サンガモ セラピューティクス, インコーポレイテッド | Composition for linking DNA binding domain and cleavage domain |
| JP2022051772A (en) * | 2016-02-02 | 2022-04-01 | サンガモ セラピューティクス, インコーポレイテッド | Composition for linking DNA binding domain and cleavage domain |
| US11920169B2 (en) | 2016-02-02 | 2024-03-05 | Sangamo Therapeutics, Inc. | Compositions for linking DNA-binding domains and cleavage domains |
| EP4130256A4 (en) * | 2020-06-28 | 2024-04-10 | Jiangsu Conquers Medical Technology Co., Ltd. | Enzyme reaction solution for constructing sequencing library and use thereof |
| WO2025101946A1 (en) * | 2023-11-08 | 2025-05-15 | Precision Biosciences, Inc. | Polypeptide linkers for use in engineered meganucleases |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| JP6839691B2 (en) | Methods for Producing Compact TALE-nucleases and Their Use | |
| US9540623B2 (en) | Method for increasing the efficiency of double-strand-break induced mutagenesis | |
| EP2633040B1 (en) | Method for increasing the efficiency of double-strand break-induced mutagenesis | |
| US11220683B2 (en) | Method to overcome DNA chemical modifications sensitivity of engineered TALE DNA binding domains | |
| EP2951295B1 (en) | Tevi chimeric endonucleases and their preferential cleavage sites | |
| WO2012138901A1 (en) | Method for enhancing rare-cutting endonuclease efficiency and uses thereof | |
| HK1192709B (en) | Method for the generation of compact tale-nucleases and uses thereof | |
| HK1192709A (en) | Method for the generation of compact tale-nucleases and uses thereof |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| 121 | Ep: the epo has been informed by wipo that ep was designated in this application |
Ref document number: 12713842 Country of ref document: EP Kind code of ref document: A1 |
|
| NENP | Non-entry into the national phase |
Ref country code: DE |
|
| 122 | Ep: pct application non-entry in european phase |
Ref document number: 12713842 Country of ref document: EP Kind code of ref document: A1 |