WO2012019300A1 - Endometrial cancer biomarkers and methods of identifying and using same - Google Patents
Endometrial cancer biomarkers and methods of identifying and using same Download PDFInfo
- Publication number
- WO2012019300A1 WO2012019300A1 PCT/CA2011/050485 CA2011050485W WO2012019300A1 WO 2012019300 A1 WO2012019300 A1 WO 2012019300A1 CA 2011050485 W CA2011050485 W CA 2011050485W WO 2012019300 A1 WO2012019300 A1 WO 2012019300A1
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- endometrial cancer
- protein
- endometrial
- markers
- marker
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Ceased
Links
Classifications
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K49/00—Preparations for testing in vivo
- A61K49/0002—General or multifunctional contrast agents, e.g. chelated agents
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q1/00—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions
- C12Q1/68—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions involving nucleic acids
- C12Q1/6876—Nucleic acid products used in the analysis of nucleic acids, e.g. primers or probes
- C12Q1/6883—Nucleic acid products used in the analysis of nucleic acids, e.g. primers or probes for diseases caused by alterations of genetic material
- C12Q1/6886—Nucleic acid products used in the analysis of nucleic acids, e.g. primers or probes for diseases caused by alterations of genetic material for cancer
-
- G01N33/5755—
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q2600/00—Oligonucleotides characterized by their use
- C12Q2600/158—Expression markers
-
- G—PHYSICS
- G01—MEASURING; TESTING
- G01N—INVESTIGATING OR ANALYSING MATERIALS BY DETERMINING THEIR CHEMICAL OR PHYSICAL PROPERTIES
- G01N2800/00—Detection or diagnosis of diseases
- G01N2800/52—Predicting or monitoring the response to treatment, e.g. for selection of therapy based on assay results in personalised medicine; Prognosis
-
- G—PHYSICS
- G01—MEASURING; TESTING
- G01N—INVESTIGATING OR ANALYSING MATERIALS BY DETERMINING THEIR CHEMICAL OR PHYSICAL PROPERTIES
- G01N2800/00—Detection or diagnosis of diseases
- G01N2800/56—Staging of a disease; Further complications associated with the disease
Definitions
- the invention relates to endometrial cancer markers, methods for assessing the status of endometrial cell tissue, and methods for the detection, diagnosis, prediction, monitoring, and therapy of endometrial cancer.
- One aspect of the invention relates to biomarkers of endometrial carcinoma and methods for detection, diagnosis, prediction, monitoring, and therapy for endometrial carcinoma and related conditions.
- Endometrial carcinoma is the fourth most prevalent cancer in North America women (10, 1 a), and the most common invasive carcinoma of the female genital tract. Definitive diagnosis is mostly based on histological examination of tissues obtained after a biopsy, an invasive procedure typically performed as a result of investigative diagnosis following abnormal uterine bleeding at presentation.
- EmCa is a heterogeneous disease, classified according to histological cell type, with the most common being endometrioid, serous, and clear cell carcinoma (2a).
- the two main types of EmCa are Type I and Type I I.
- Type I is of endometrioid histology, while the more clinically aggressive Type I I is primarily serous in morphology.
- Clear cell carcinoma is also grouped under Type I I EmCa, and can sometimes be present admixed with endometrioid and serous tumor components (1 1 ).
- EmCa subtypes Given its role in identifying EmCa subtypes, histology is pivotal. Inter-observer studies have shown moderate to excellent reproducibility from sample to sample within a given cell type, when considering all grades and cell types. Reproducibility would undoubtedly improve if only high- grade subtypes were considered. Morphological overlap exists between the subtypes, and, in the case of clear cell and serous, there is morphological and clinicopathological overlap. Increasingly, immunohistochemical markers are used to refine cell type in difficult cases. In addition, biopsy samples are small and may be poorly representative (e.g., diagnostic features that are focal may not be sampled).
- FIGO grading In terms of FIGO grading, there is also the potential issue of inter-observer variability.
- the histological cut points have been defined arbitrarily, and are not the result of an outcome-driven process.
- ovarian cancer a need for subtype-specific therapies has recently been recognized.
- subtype-specific chemotherapeutic agents are also required for the subtypes of endometrial cancers.
- the application of the FIGO grading system requires histological assessment of the percentage of solid growth within the tumor and the degree of cytologic atypia.
- quantitative proteomics may provide distinguishing differences in protein expression levels, as well as surveillance and therapeutic targets specific to given histological types.
- biomarkers are biometric measurements that convey information about the biological condition of the subject being tested.
- a biomarker should be a molecule expressed at a higher concentration by an abnormal tissue, such as a nodule of cancer cells.
- Proteins are good biomarker candidates because protein expression is closely linked to the metabolic state of the cell. This is particularly pertinent to cancer diagnosis: tumor cells that are undergoing rapid uncontrolled proliferation are bound to have biochemical changes that would manifest as changes in expression levels of specific proteins. To identify biomarkers, a quantitative or, at a minimum, semi-quantitative approach is required, as it is expected that all healthy and tumor cells will express a common core of proteins, and that most changes will be in terms of relative expression levels rather than absolute presence or absence of a given protein. Biomarker candidates include proteins for which patterns of differential expression are specific for a disease condition, and could be either causative or symptomatic in nature.
- a proteomics approach to biomarker discovery consists of a quantitative comparison of the proteins expressed between tissues at two or more different states, including different subtypes of the disease. Proteins that are differentially expressed in distinct disease subtypes can be useful as specific diagnostic and prognostic markers, since different subtypes have different etiologies.
- One of the challenges in the quest for biomarkers is the selection of molecules that show both selectivity and specificity for the studied illness, particulaly if the final objective is to assay these biomarkers in a body fluid, such as blood (see, e.g. , references 4a-7a for discussions of biomarker discovery challenges).
- IMS isotope dilution mass spectrometry
- LC chromatography
- ESI electrospray ionization
- MS/MS tandem mass spectrometry
- the peptides identified are mapped to proteins in a so-called “bottom- up” strategy.
- Quantification is performed via the reporter ions, m/z 1 14-1 17, generated from the iTRAQ tags under MS/MS.
- the tagging serves to distinguish proteins originating from the individual samples in the mixture; a comparison among the abundances of the reporter ions results in relative quantification of the given peptide and the protein from which the peptide originates.
- Due to the complexity of tissue samples, a prefractionation of the samples is typically performed using strong cation exchange (SCX) prior to the reverse phase (RP) separation that is coupled online with MS/MS.
- SCX strong cation exchange
- RP reverse phase
- a possible solution is iterative analysis of the same sample, using an exclusion list of identified peptides to instruct the mass spectrometer to ignore those ions that have already been selected in the previous run(s), when choosing ions for MS/MS analysis in subsequent runs.
- Attempts at iterative analysis have been reported for matrix-assisted laser desorption / ionization (MALDI) MS/MS (8, 9) and ESI -MS/MS (7).
- MALDI matrix-assisted laser desorption / ionization
- MS/MS (8, 9)
- ESI -MS/MS ESI -MS/MS (7).
- the last study (7) reported an extensive comparison of four different approaches to creating the exclusion list, ranging from a simple list of identified ions to a complex list of all potential ions from identified proteins plus all selected but unidentified ions.
- a key issue in any iterative approach is the confidence of the peptide identification.
- EmCa endometrial carcinoma
- the inventors identified a number of differentially expressed proteins for use as EmCa biomarkers. As described in greater detail herein, the inventors have further verified and confirmed those results, and identified a new set of biomarkers, using an iterative analytical strategy in which a list of the peptide ions identified in the first run is employed as an exclusion list for the subsequent acquisition. The inventors' strategy allows progressively lower- abundance peptide ions to be selected and identified.
- EmCa multidimensional liquid chromatography-mass spectrometry (LC-MS / MS) to analyze biological samples labelled with isobaric mass tags (iTRAQ)
- LC-MS / MS isobaric mass tags
- the inventors identified proteins that were differentially expressed in EmCa (e.g. , Type I EmCa and grade I I I endometrioid EmCa, clear cell EmCa, and serous EmCa), as compared with normal proliferative samples (control).
- LC-MS / MS isobaric mass tags
- PK pyruvate kinase
- LDHA lactate dehydrogenase A
- AAT Alpha-1 -antitrypsin
- CAG macrophage capping protein
- Other proteins that were downregulated include fibrinogen, haptoglobulin, apolipoprotein A-l and apoliprotein A-l I, and serpin H1 , all of which are involved in extracellular matrix formation, motility, and/or inflammatory reaction.
- Pathway analysis showed that most differentially expressed proteins map to a single network that has transcription factors and kinases linked to tumorigenesis as central protein hubs.
- FIGO grade I II endometrioid In an analysis of different EmCa subtypes, six FIGO grade I II endometrioid, five clear cell, and eight serous carcinomas were compared using iTRAQ labeling and LC-MS/MS analysis. Using their iterative approach, the inventors identified eighteen proteins that showed differential expression in different histological types of high-grade endometrial cancer (Table 8). Fourteen proteins showed a similar trend for all samples, and may be considered suitable candidates for use as endometrial cancer biomarkers (Table 9). Heterogeneous nuclear ribonucleoprotein A2/B1 (hnRNP A2/B1 ) was found to be specifically up-regulated in FIGO grade I I endometrioid carcinomas.
- hnRNP A2/B1 Heterogeneous nuclear ribonucleoprotein A2/B1
- L-lactate dehydrogenase A was observed to be constantly up-regulated in endometrioid samples; this protein is also up-regulated in clear cell carcinomas.
- Hepatoma derived growth factor (HDGF) was up-regulated in the serous endometrial carcinomas (Table 8).
- the inventors have also identified a series of proteins which exhibit varying expression patterns across different subtypes of high-grade endometrial cancers, including grade I I I endometrioid, clear cell, and serous EmCa.
- the invention provides a method for detecting one or more endometrial cancer markers in a subject by: (a) obtaining a sample from a subject; (b) detecting in the sample an amount of at least one of the endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9; and (c) comparing the detected amount with an amount detected for a standard.
- the invention provides a method for diagnosing endometrial cancer in a subject, by comparing: (a) levels of at least one of the endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9; and (b) normal levels of expression of the corresponding endometrial cancer marker or polynucleotide in a control sample, wherein a significant difference in levels of the endometrial cancer marker, relative to the corresponding normal levels, is indicative of the endometrial cancer in the subject.
- the endometrial cancer marker is a protein
- the invention provides a method for diagnosing endometrial cancer in a subject, by: (a) contacting a biological sample obtained from the subject with at least one binding agent that specifically binds to at least one of the endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9; and (b) detecting in the sample amounts of the endometrial cancer marker or polynucleotide or part thereof that binds to the at least one binding agent, relative to a predetermined standard or cut-off value, and thereby determining the presence or absence of the endometrial disease in the subject.
- the binding agent is an antibody.
- the invention provides a method of screening a subject for endometrial cancer by: (a) obtaining a biological sample from a subject; (b) detecting in the sample an amount of at least one of the endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9; and (c) comparing the detected amount of the endometrial cancer marker or polynucleotide with a predetermined standard, wherein detection of a level of the endometrial cancer marker or polynucleotide different from that of the standard is indicative of endometrial cancer.
- the level of the endometrial cancer marker or polynucleotide is significantly higher than the standard and is indicative of endometrial cancer. In certain other embodiments, the level of the endometrial cancer marker or polynucleotide is significantly lower than the standard and is indicative of endometrial cancer.
- the sample is obtained from a tissue, extract, cell culture, cell lysate, lavage fluid, or physiological fluid of the subject. In still other embodiments, the sample is obtained from endometrial tumor tissue.
- the invention provides a method for determining the presence or absence of an endometrial cancer marker associated with an endometrial disease in a subject by: (a) detecting at least one of the endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9; and (b) relating the detected amount of the marker or polynucleotide to the presence of the endometrial disease.
- the polynucleotide is mRNA.
- the polynucleotide is detected by: (a) contacting the sample with at least one oligonucleotide that hybridizes to the polynucleotide; and (b) detecting in the sample levels of at least one nucleic acid that hybridizes to the polynucleotide, relative to a predetermined standard or cut-off value, and thereby determining the presence or absence of the endometrial disease in the subject.
- the polynucleotide is detected with an amplification reaction.
- the amplification reaction includes a polymerase chain reaction employing
- oligonucleotide primers that hybridize to the polynucleotide or to a precursor or complement thereof. Additionally, in certain embodiments, the polynucleotide is detected with at least one oligonucleotide probe that hybridizes to the polynucleotide or to a complement thereof.
- the polynucleotide is detected by: (a) isolating RNA from the sample; (b) combining the RNA with at least one reagent, to convert the RNA to cDNA; (c) treating the cDNA with at least one amplification reaction reagent and at least one primer that hybridizes to the cDNA, to produce at least one amplification product; (d) analyzing the at least one amplification product to detect an amount of RNA encoding the at least one endometrial cancer marker; and (e) comparing the amount of RNA to an amount detected against a panel of expected values for normal tissue derived using similar primers.
- the invention provides a method for diagnosing and monitoring endometrial cancer in a subject by: (a) isolating at least one nucleic acid in a sample from the subject; and (b) detecting at least one of the endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9, wherein the presence of higher or lower levels of the marker or polynucleotide in the sample, compared to a standard or control, is indicative of the disease or prognosis.
- the invention provides a method for monitoring the progression of endometrial cancer in a subject by: (a) detecting in a sample from the subject, at a first time point, at least one of the endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9; (b) repeating step (a) at a subsequent point in time; and (c) comparing levels detected in steps (a) and (b), and thereby monitoring the progression of endometrial cancer.
- the invention provides a method for determining in a subject whether endometrial cancer has metastasized or is likely to metastasize in the future, by comparing:
- the invention provides a method for assessing the aggressiveness or indolence of endometrial cancer, by comparing: (a) levels of at least one of the endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9; and (b) normal levels of the endometrial cancer marker or polynucleotide in a control sample, wherein a significant difference between the levels in the subject sample and normal levels is indicative that the cancer is aggressive or indolent.
- the invention provides a diagnostic composition including an agent that: binds to at least one of the endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9, or hybridizes to at least one polynucleotide encoding at least one of the endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9.
- the invention provides a method for assessing the potential efficacy of a test agent for inhibiting endometrial cancer in a subject, by comparing: (a) levels of at least one of the endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9; and (b) levels of at least one of the endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9 in a second sample obtained from the subject, wherein the second sample has not been exposed to the test agent, and wherein a significant difference in the levels of the endometrial cancer marker or polynucleotide in the first sample, relative to the second sample, is an indication that the test agent inhibits endometrial cancer in the subject.
- the invention provides a method for assessing the efficacy of a therapy for inhibiting endometrial cancer in a subject, by comparing: (a) levels of at least one of the endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9 in a first sample obtained from the subject; and (b) levels of at least one of the endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9 in a second sample obtained from the subject following therapy, wherein a significant difference in the levels of expression of the endometrial cancer marker or polynucleotide in the second sample, relative to the first sample, is an indication that the therapy inhibits endometrial cancer in the subject.
- the invention provides a method for selecting an agent for inhibiting endometrial cancer in a subject by: (a) obtaining a sample including precancer or cancer cells from the subject; (b) separately exposing aliquots of the sample to a plurality of test agents;
- the invention provides a method for inhibiting endometrial cancer in a subject by: (a) obtaining a sample including precancer or cancer cells from the subject; (b) separately maintaining aliquots of the sample in the presence of a plurality of test agents;
- the invention provides a method for assessing the endometrial cancer cell carcinogenic potential of a test compound by: (a) maintaining separate aliquots of endometrial cancer cells in the presence and absence of the test compound; and (b) comparing levels of at least one of the endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9, in the aliquots, wherein a significant difference in levels of the endometrial cancer marker or polynucleotide in the aliquot maintained in the presence of the test compound, relative to levels in the aliquot maintained in the absence of the test compound, is indicative that the test compound possesses endometrial cancer cell carcinogenic potential.
- the invention provides an in vivo method for imaging an endometrial disease by: (a) injecting a subject with one or more agents that bind to an endometrial cancer marker listed in Table 5a, Table 5b, Table 8, and/or Table 9, the agent carrying a label for imaging the endometrial cancer marker; (b) allowing the agent to incubate in vivo and bind to an endometrial cancer marker; and (c) detecting the presence of the label localized to diseased endometrial tissue.
- the agent is an antibody that specifically reacts with an endometrial cancer marker.
- the endometrial cancer marker is listed in Table 5a, Table 5b, Table 8, and/or Table 9.
- the invention provides a set of endometrial cancer markers, including at least 2 of the markers listed in Table 5a, Table 5b, Table 8, and/or Table 9 or polynucleotides encoding same.
- the at least 2 markers are protein endometrial cancer markers (e.g. , L-lactate dehydrogenase A (LDHA), nucleophosmin, heterogenous nuclear ribonucleoprotein A1 , cystatin B, transferrin, nucleolin, villin 2 (ezrin), or galectin-1 ).
- kits comprising the endometrial cancer markers of the invention.
- the invention provides a kit for determining the presence of endometrial cancer in a subject, including a known amount of at least one binding agent that binds to at least one of the protein endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9, or a part thereof, wherein the binding agent includes a detectable substance or binds directly or indirectly to a detectable substance.
- the invention provides a kit for determining the presence of endometrial cancer in a subject, including a known amount of an oligonucleotide that hybridizes to a polynucleotide encoding an endometrial cancer marker listed in Table 5a, Table 5b, Table 8, and/or Table 9, wherein the oligonucleotide is directly or indirectly labelled with a detectable substance.
- the endometrial cancer marker is an EmCa marker.
- the endometrial cancer marker may be a Type I EmCa marker.
- Exemplary EmCa that may be detected by the markers of the present invention include, without limitation, endometrioid EmCa, clear cell EmCa, and serous EmCa.
- the endometrioid EmCa marker is a heterogeneous nuclear ribonucleoprotein (hnRNPA).
- the serous EmCa marker is a hepatoma-derived growth factor.
- the invention provides a use of a marker, or differential expression of a marker, to screen for, diagnose, or monitor the progress, treatment, recurrence, or prognosis of endometrial cancer.
- exemplary markers include, without limitation, L-lactate dehydrogenase A (LDHA), phosphoglycerate kinase, pyruvate kinase isozymes M 1/M2, nucleolin, glucose-6-phosphate isomerase, alpha-enolase, villin 2, isocitrate dehydrogenase 2 (mitochondrial), peroxiredoxin-1 , transketolase, serpin H1 , transferrin, galectin-1 , and lumican, and polynucleotides encoding same.
- LDHA L-lactate dehydrogenase A
- phosphoglycerate kinase phosphoglycerate kinase
- the invention provides an iterative computerized method for identifying at least one protein endometrial cancer marker, by: (a) conducting a first mass spectrometry analysis of peptide ions in normal and diseased tissues, to identify a first set of potential markers based on differential expression; (b) creating an exclusion list consisting of the peptide ions identified in the first set of potential markers, wherein the exclusion list is restricted by a mass window and a time window; (c) conducting a second mass spectrometry analysis of peptide ions in the normal and diseased tissues, to identify a second set of potential markers based on differential expression, wherein the second analysis disregards peptide ions in the exclusion list; (d) updating the exclusion list to consist of the peptide ions identified in the first and second sets of potential markers; and (e) conducting a third mass spectrometry analysis of peptide ions in the normal and diseased tissues, to identify a third set of potential markers based on differential expression, wherein the third
- the method includes the step of calculating a false discovery rate for each identified potential marker. Also provided is a system for identifying at least one protein endometrial cancer marker.
- the system includes at least one computing device including software that, when executed, performs the iterative computerized method of the invention.
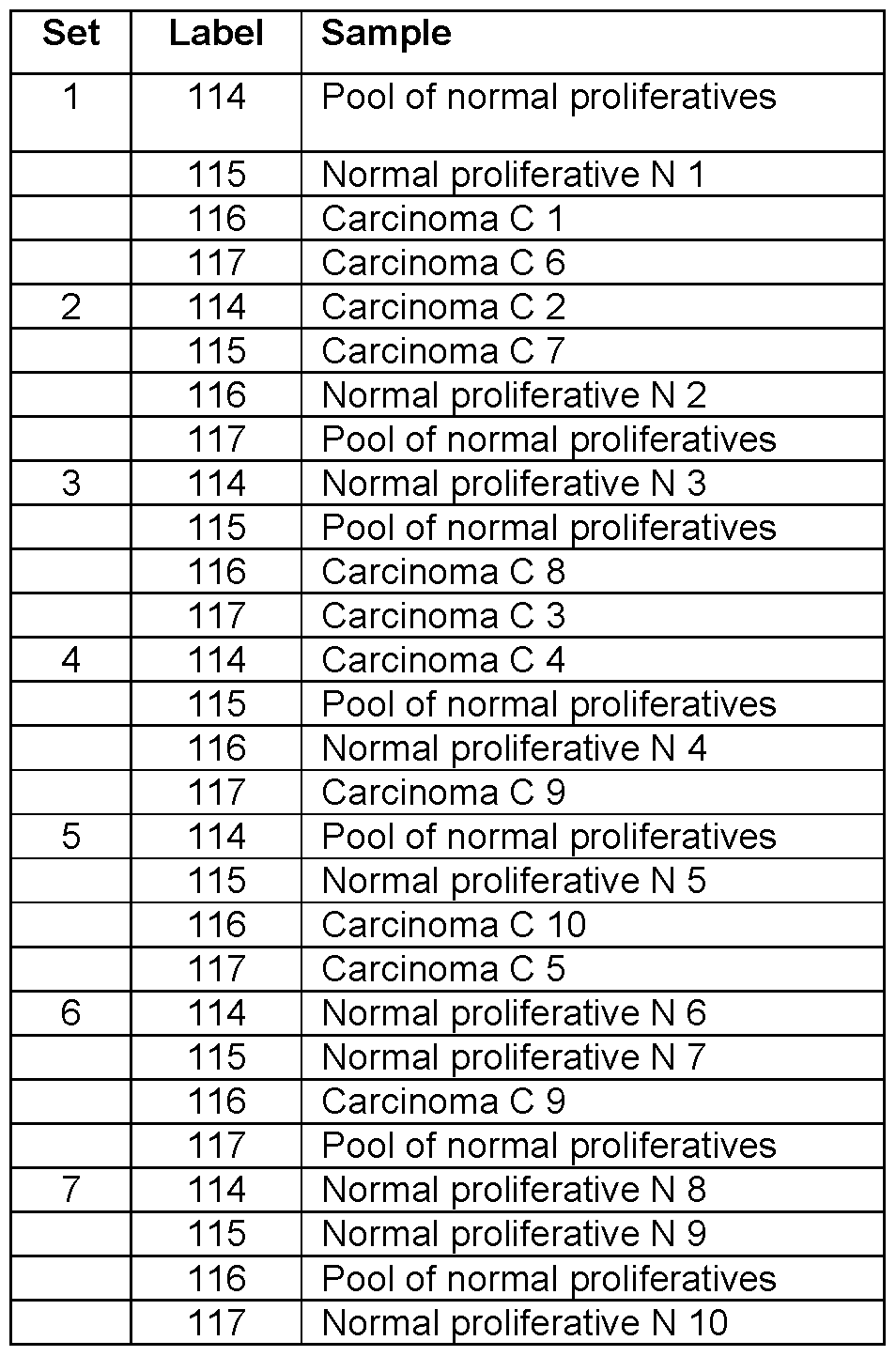
- Table 1 is a description of each set of samples that were used in the analysis of Type I EmCa samples to find EmCa biomarkers.
- Table 2a is the liquid chromatography gradient used for strong cation exchange (SCX) HPLC fractionation in the analysis of Type I EmCa samples.
- Table 2b is the solvent gradient used for reverse-phase (RP) HPLC separation in the analysis of Type I EmCa samples.
- Table 3 is a comparison of protein functions or categories between the proteins identified in the analysis of Type I EmCa samples and those identified in an earlier study.
- Table 4 is a comparison of identified protein numbers across the iterative analysis of iTRAQ sets in the analysis of Type I EmCa samples. Numbers correspond to the unique proteins identified in the forward database. Identifications pointing to reversed sequences have been removed. Values in the "additional" columns do not include the proteins identified in two or more iterations. Percentages are calculated as the ratios of newly-identified proteins over proteins already identified in all previous iterations in %.
- Table 5a is a comparison of the protein mean iTRAQ ratios for the analysis of Type I EmCa samples that meet the criteria described in the Examples. For each set, the protein mean ratios over the three analyses have been calculated, taking into account the iTRAQ p-values from ProteinPilot® quantification. Columns also display the number of samples in which the protein has been identified (#, out of 10).
- Table 5b is a comparison of the protein mean iTRAQ ratios for the analysis of Type I EmCa samples in which the proteins exhibited changes in expression between 40% and 50% or expression ratios between 1.4 and 1.5 or between 0.67 and 0.71.
- Table 6 shows the liquid chromatography gradients used for strong cation exchange (SCX) and HPLC fractionation in the analysis of different EmCa subtypes.
- Table 7 provides the number of proteins identified by the iterative method in the eight sample sets studied in the analysis of different EmCa subtypes.
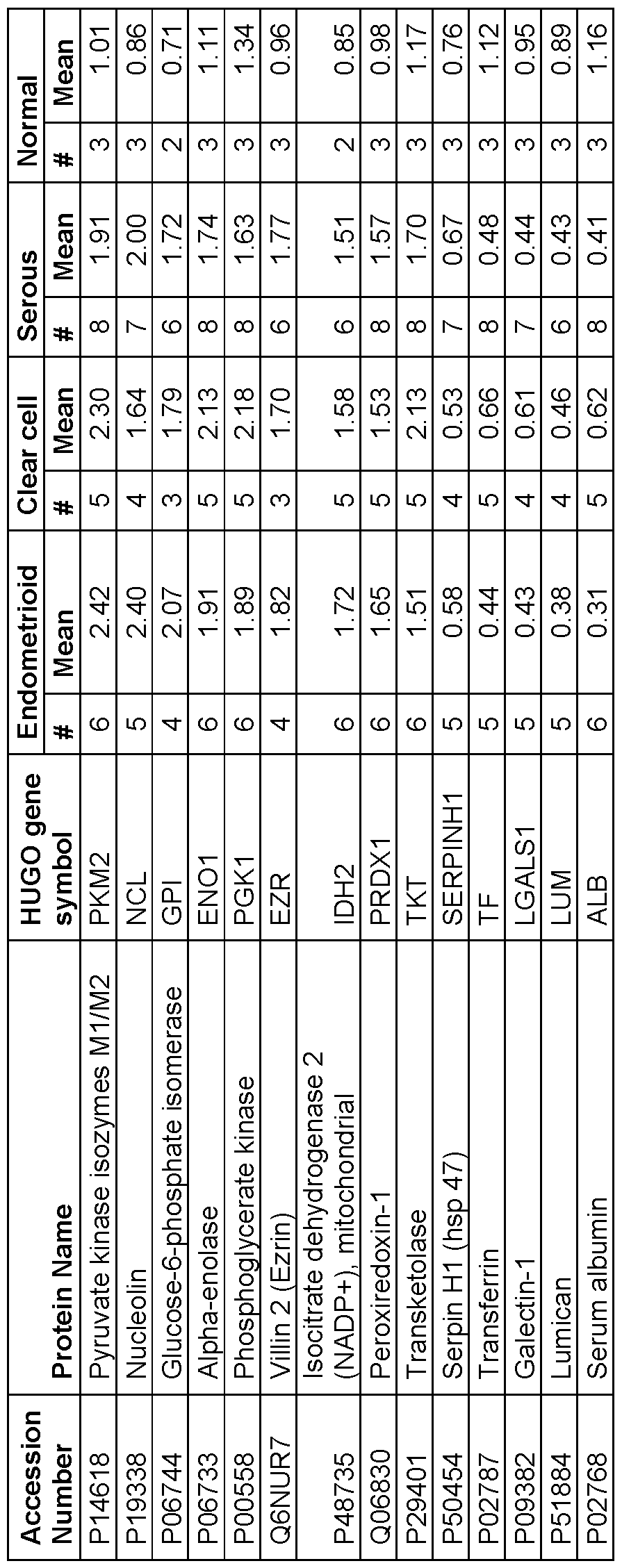
- Table 8 lists the proteins from the EmCa subtype analysis which exhibited divergent expression between EmCa samples as compared to normal proliferative tissues. "#” is the number of samples in the subtype in which the given expression was confidently determined. "Mean” is the average expression ratio.
- Table 9 provides a list of proteins showing similar expression trends regardless of EmCa subtype.
- Table 10 shows how iTRAQ labels were assigned randomly to the pooled reference sample and tissue homogenates in the analysis of different EmCa subtypes. These labeled samples were mixed in sets of four, with each set containing one label of each type, resulting in a total of eight sets.
- Table 1 1 is a comparison of iTRAQ ratios for proteins identified in three different studies of EmCa samples.
- FIG. 1 depicts interaction network #1 for EmCa biomarker candidates. Overexpressed proteins are shown in red; underexpressed proteins are shown in green. Uncolored proteins were added by Ingenuity Pathway Analysis to connect the submitted biomarker candidates.
- AHSG alpha- 2-HS-glycoprotein
- ALDH1 A2 aldehyde dehydrogenase 1 family, member A2
- APOA1 apolipoprotein A-l
- APOA2 apolipoprotein A-l l
- APOH apolipoprotein H (beta-2-glycoprotein I)
- CAP1 CAP, adenylate cyclase-associated protein 1
- CAPG capping protein (actin filament), gelsolin-like
- CAPZB capping protein (actin filament) muscle Z-line, beta
- CD163 CD163 molecule
- CLEC1 1A C-type lectin domain family 11 , member A
- COL1 A2 collagen
- eukaryotic translation initiation factor 4E eukaryotic translation initiation factor 4E; GPLD1 : glycosylphosphatidylinositol specific phospholipase D1 ; HNF1 A: HNF1 homeobox A; HNRNPA1 : heterogeneous nuclear ribonucleoprotein A1 ; HP: haptoglobin; IL1 B: interleukin 1 , beta; IL2: interleukin 2; IL13: interleukin 13; ILF2: interleukin enhancer binding factor 2, 45kDa; ILF3: interleukin enhancer binding factor 3, 90kDa; KITLG: KIT ligand; LCAT: lecithin-cholesterol acyltransferase; LIPC: lipase, hepatic; MYC: v-myc
- NCL nucleolin
- NOS2 nitric oxide synthase 2, inducible
- PCBP1 poly(rC) binding protein 1
- PKM2 pyruvate kinase M1/M2
- PLTP phospholipid transfer protein
- SERPINH1 serpin peptidase inhibitor, clade H (heat shock protein 47); TP53: tumor protein p53.
- FIG. 2 sets forth interaction network #2 for EmCa biomarker candidates. Overexpressed proteins are shown in red; underexpressed proteins are shown in green. Uncolored proteins were added by Ingenuity Pathway Analysis to connect the submitted biomarker candidates.
- A2M alpha-2- macroglobulin
- ACTA2 actin, alpha 2, smooth muscle, aorta
- ALB albumin
- ANXA1 annexin A1
- BAD BCL2-associated agonist of cell death
- CASP7 caspase 7, apoptosis-related cysteine peptidase
- COL1 A1 collagen, type I, alpha 1
- DAPK1 death-associated protein kinase 1
- DUSP1 dual specificity phosphatase 1
- ERK p42/p44 Map Kinase
- EZR Ezrin (villin 2);
- FGG fibrinogen gamma chain
- FLNC filamin C
- PDGFB platelet-derived growth factor beta polypeptide
- PEA15 phosphoprotein enriched in astrocytes 15
- PIAS4 protein inhibitor of activated STAT, 4
- PLD1 phospholipase D1 , phosphatidylcholine-specific
- Raf Raf kinase
- SCARB1 scavenger receptor class B, member 1
- TCHP trichoplein, keratin filament binding
- TF transferrin
- TFRC transferrin receptor (p90).
- FIG. 3 presents interaction network #3 for EmCa biomarker candidates. Overexpressed proteins are shown in red; underexpressed proteins are shown in green. Uncolored proteins were added by Ingenuity Pathway Analysis to connect the submitted biomarker candidates.
- ADAMTS1 ADAM metallopeptidase with thrombospondin type 1 motif
- BIRC5 ADAM metallopeptidase with thrombospondin type 1 motif
- FIG. 4 depicts interaction network for EmCa subtype biomarker candidates. This network was constructed using the proteins from Table 8. Overexpressed proteins are shown in red; underexpressed proteins are shown in green. See Table 8 for more details. Uncolored proteins were added by Ingenuity Pathway Analysis to connect the submitted biomarker candidates.
- ALDH2 ALDH2
- aldehyde dehydrogenase 2 family (mitochondrial); APOA1 : apolipoprotein A-l; BRD2: bromodomain containing 2; C3: complement component 3; C3AR1 : complement component 3a receptor 1 ; CASP1 : caspase 1 , apoptosis-related cysteine peptidase (interleukin 1 , beta, convertase); Caspase 3/7: Caspase group; CDKN2A: cyclin-dependent kinase inhibitor 2A (melanoma, p16, inhibits CDK4); CR2: complement component (3d/Epstein Barr virus) receptor 2; EIF4E: eukaryotic translation initiation factor 4E; FCER2: Fc fragment of IgE, low affinity I I, receptor for (CD23); HDGF: hepatoma- derived growth factor (high-mobility group protein 1-like); HNRNPA1 : heterogeneous nuclear rib
- nucleophosmin nucleolar phosphoprotein B23, numatrin
- ORM 1 orosomucoid 1
- PCBP1 includes EG:5093
- PLA2G2A phospholipase A2, group I IA (platelets, synovial fluid)
- PLTP phospholipid transfer protein
- PPA1 pyrophosphatase (inorganic) 1
- PRDX5 PRDX5
- peroxiredoxin 5 RARA: retinoic acid receptor, alpha; SERPINA1 : serpin peptidase inhibitor, clade A (alpha-1 antitrypsin), member 1 ; SMPD2: sphingomyelin phosphodiesterase 2, neutral membrane ; SOD2: superoxide dismutase 2, mitochondrial; SPHK1 : sphingosine kinase 1 ; TMSB4X: thymosin beta 4, X-linked; TNF: tumor necrosis factor (TNF superfamily, member 2); TP53: tumor protein p53; VCAM 1 : vascular cell adhesion molecule 1 ; VVDR1 : WD repeat domain 1 ; YBX2: Y box binding protein 2.
- FIG. 5 presents interaction network #5 for EmCa biomarker candidates. This network was constructed using the proteins from Table 9. Overexpressed proteins are shown in red;
- ALB albumin
- cytokine signaling 3 SUM01 : SMT3 suppressor of mif two 3 homolog 1 (S. cerevisiae);
- TERC telomerase RNA component;
- TF transferrin;
- TFRC transferrin receptor (p90, CD71 );
- TKT transketolase;
- TNF tumor necrosis factor (TNF superfamily, member 2).
- the inventors previously performed a 40-sample iTRAQ study on EmCa that resulted in the identification of a number of differentially expressed proteins (12).
- tryptic peptides from tumor and normal endometrial tissues were compared after iTRAQ-labeling and two-dimensional LC with online nanoESI and MS/MS.
- the inventors implement a more thorough analysis of their samples, to confirm and expand their earlier results.
- the objectives of the present work were three-fold: first, to identify a larger number of proteins from the inventors' samples than previously achieved (12); second, to discover more proteins as cancer biomarkers than before; and third, to provide further verification of the inventors' previous results on an independent batch of cancer samples.
- the inventors compared normal proliferative tissue samples and Type I EmCa tissue samples employing the same proteomic strategy as described in the earlier study (12).
- the present comparison followed a "drill-down" approach in the proteomic analysis in which each sample was injected three times, and the acquisition software was programmed to exclude the peptides ions identified during previous iterations.
- Exclusion was restricted by means of a mass window and a time window, the latter to reduce the chance that peptide ions having m/z values similar to ones previously identified, but eluting at different times, would be excluded.
- the method described herein is considerably less involved than previous studies, as it did not incorporate the unidentified and expected ions with other charge states.
- the inventors also used the analysis of three different subtypes of EmCa to identify cancer biomarkers that can be used for classifying EmCa into subtypes and for general identification of EmCa.
- the inventors employed a new feature of the database search engine ProteinPilot to monitor the quality of the identifications.
- This new feature the plug-in Proteomics System Performance Evaluation Pipeline (PSPEP), compares every identification to the chance of random matching in a reverse database and calculates the individual false discovery rate (FDR) for each identified protein entry (13). This FDR value was used as validation of the ProteinPilot identification.
- PSPEP Proteomics System Performance Evaluation Pipeline
- the inventors describe herein methods for detecting: the presence of an endometrial disease, including EmCa and similar conditions, in a sample; the absence of a disease in a sample; the stage or grade of a disease; and other characteristics of endometrial diseases that are relevant to prevention, diagnosis, characterization, monitoring, and therapy of endometrial diseases in a patient, including the metastatic potential and indolence or aggressiveness of endometrial cancer in a patient.
- Methods are also provided for: assessing the efficacy of one or more test agents for inhibiting a endometrial disease; assessing the efficacy of a therapy for endometrial diseases, such as EmCa; monitoring the progression of endometrial diseases, such as EmCa; selecting an agent or therapy for inhibiting endometrial diseases, such as EmCa; treating a patient afflicted with EmCa; inhibiting an endometrial disease, such as EmCa; and assessing the disease (e.g. , carcinogenic) potential of a test compound.
- the invention provides marker sets that detect endometrial diseases and uses therefor.
- a marker set may include a plurality of polypeptides, or polynucleotides encoding such polypeptides, including or consisting of at least one marker listed in Table 5a, Table 5b, Table 8, and/or Table 9, and, optionally, 2, 3, 4, 5, 6, and 7 or up to all of the markers listed therein.
- the markers consist of at least 2, 3, 4, or 5 polypeptides listed in Table 5a, Table 5b, Table 8, and/or Table 9 or polynucleotides encoding same.
- the protein marker sets include or consist of protein clusters, or proteins in pathways, including markers listed in Table 5a, Table 5b, Table 8, and/or Table 9 or polynucleotides encoding same.
- the invention provides markers in Table 5a, Table 5b, Table 8, and/or Table 9 that are up-regulated or down-regulated or differentially expressed in cancer samples as compared to the non-cancer samples.
- a "protein endometrial cancer marker” includes the up- regulated and down- regulated protein markers identified in Table 5a, Table 5b, Table 8, and/or Table 9 (up-regulated or down- regulated in cancer samples versus non-cancer samples), as well as native- sequence polypeptides, chimeric polypeptides, and all isoforms, homologs, fragments, and precursors thereof. Also included are modified forms of the proteins and derivatives thereof.
- the invention includes, within its scope, polynucleotides encoding the protein EmCa markers.
- polynucleotides may be referred to herein, variously, as “polynucleotides encoding protein endometrial cancer markers" or
- Protein endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9 (in cancer sample versus non-cancer sample) and polynucleotides encoding the endometrial cancer markers may be used to determine the status of an endometrial cell or tissue and to detect an endometrial disease, such as endometrial cancer.
- the markers can be used for diagnosis, monitoring (including, without limitation, monitoring progression of disease state or success of therapeutic treatment), prognosis, treatment, or classification of an endometrial disease (including, without limitation, cancer, EmCa, and related conditions).
- the markers can also be used to evaluate disease states before surgery or after relapse.
- the invention also contemplates methods for assessing the status of an endometrial tissue, and methods for the diagnosis and therapy of an endometrial disease.
- endometrial cancer can be assessed or characterized, for example, by detecting the presence in a sample of: (a) an endometrial cancer marker or fragment thereof; (b) a metabolite which is produced directly or indirectly by an endometrial cancer marker; (c) a transcribed nucleic acid or fragment thereof having at least a portion with which a polynucleotide endometrial cancer marker is substantially identical; and/or (d) a transcribed nucleic acid or fragment thereof, wherein the nucleic acid hybridizes with a polynucleotide marker encoding a protein endometrial cancer marker.
- the levels of endometrial cancer markers may be determined by methods as described herein and generally known in the art.
- the expression levels of protein endometrial cancer markers may be determined by isolating and determining the level of nucleic acid transcribed from each polynucleotide endometrial cancer marker.
- the levels of protein endometrial cancer markers translated from mRNA transcribed from an endometrial cancer polynucleotide marker may be determined.
- the invention provides a method for characterizing or classifying an endometrial sample by detecting a difference in the expression of a first plurality of endometrial cancer markers relative to a control, the first plurality of such markers including at least 2, 3, 4, or 5 of the endometrial cancer markers corresponding to the markers listed in Table 5a, Table 5b, Table 8, and/or Table 9, and optionally 2, 3, 4, 5, 6, and 7, or up to all of the markers listed therein or those listed in Table 5a, Table 5b, Table 8, and/or Table 9 or those that are up-regulated in cancer versus control tissue as indicated in Table 5a, Table 5b, Table 8, and/or Table 9.
- a method for characterizing endometrial tissue by detecting endometrial cancer markers associated with the endometrial tissue stage or phase, or endometrial disease in a subject by: (a) obtaining a sample from a subject; (b) detecting or identifying in the sample an amount of endometrial cancer markers; and (c) comparing the detected amount with an amount detected for a standard.
- a method for detecting protein endometrial cancer markers or polynucleotide endometrial cancer markers associated with endometrial cancer in a patient by: (a) obtaining a sample from a patient; (b) detecting in the sample amount of protein endometrial cancer markers or polynucleotide endometrial cancer markers; and (c) comparing the detected amount with an amount detected for a standard.
- the term "detect” or “detecting” includes assaying, imaging or otherwise establishing the presence or absence of the target endometrial cancer markers or polynucleotides encoding the markers, subunits thereof, or combinations of reagent bound targets, and the like, or assaying for, imaging, ascertaining, establishing, or otherwise determining one or more factual characteristics of an endometrial tissue phase or endometrial disease including cancer, metastasis, stage, or similar conditions.
- the term encompasses diagnostic, prognostic, and monitoring applications for the endometrial cancer markers and polynucleotides encoding the markers.
- the invention also provides a method of assessing whether a patient is afflicted with or has a pre-disposition for endometrial disease, in particular endometrial cancer, by comparing: (a) levels of endometrial cancer markers associated with the endometrial disease in a sample from the patient; and (b) normal levels of endometrial cancer markers associated with the endometrial disease in samples of the same type obtained from control patients not afflicted with the disease, wherein altered levels of the endometrial cancer markers relative to the corresponding normal levels of such markers is an indication that the patient is afflicted with endometrial disease.
- the invention also provides a method of assessing whether a patient is afflicted with or has a pre-disposition for endometrial disease, in particular endometrial cancer, the method by comparing: (a) levels of protein endometrial cancer markers and/or the polynucleotide endometrial cancer markers associated with the endometrial disease in a sample from the patient; and (b) normal levels of endometrial cancer markers and/or the polynucleotide endometrial cancer markers associated with the endometrial disease in samples of the same type obtained from control patients not afflicted with the disease, wherein altered levels of the endometrial cancer markers and/or the polynucleotide endometrial cancer markers relative to the corresponding normal levels of such markers is an indication that the patient is afflicted with endometrial disease.
- a further embodiment of the invention provides a method for assessing whether a patient is afflicted with or has a pre-disposition for endometrial cancer, where higher levels of protein endometrial cancer markers (e.g. , a heterogeneous nuclear ribonucleoprotein (e.g. , A2/B
- hnRNPA2/B1 L-lactate dehydrogenase A
- LDHA L-lactate dehydrogenase A
- nucleophosmin nucleophosmin
- cystatin B hepatoma derived growth factor and/or calcyphosin
- Another embodiment of the invention provides a method for assessing whether a patient is afflicted with or has a pre-disposition for endometrial cancer, where lower levels of protein endometrial cancer markers (e.g. , fibrinogen (e.g. , alpha, beta, and/or gamma chain), apolipoprotein A1 , haptoglobin (HP), and/or serpin H1 (or heat-shock protein 47 kD (hsp47)) in a sample, relative to the corresponding normal levels, indicate that the patient is afflicted with endometrial cancer.
- protein endometrial cancer markers e.g. , fibrinogen (e.g. , alpha, beta, and/or gamma chain), apolipoprotein A1 , haptoglobin (HP), and/or serpin H1 (or heat-shock protein 47 kD (hsp47)
- fibrinogen e.g. , alpha, beta, and/or
- the invention provides a method for screening a subject for endometrial disease by: (a) obtaining a biological sample from a subject; (b) detecting the amount of endometrial cancer markers associated with the disease in the sample; and (c) comparing the amount of endometrial cancer markers detected to a predetermined standard, where detection of a level of endometrial cancer markers that differs significantly from the standard indicates endometrial disease.
- a significant difference between the levels of endometrial cancer marker levels in a patient and normal levels is an indication that the patient is afflicted with or has a predisposition to endometrial disease.
- the level of endometrial cancer markers is significantly higher compared to the standard and indicative of endometrial disease.
- the level of endometrial cancer markers is significantly lower compared to the standard and indicative of endometrial disease.
- the endometrial cancer markers detected are protein endometrial cancer markers or the polynucleotide endometrial cancer markers.
- the amount of protein endometrial cancer marker(s) e.g. , a heterogeneous nuclear ribonucleoprotein (e.g. , A2/B (hnRNPA2/B1 )), L-lactate dehydrogenase A (LDHA), anterior gradient protein2 homolog, nucleophosmin, cystatin B, hepatoma derived growth factor and/or calcyphosin
- protein endometrial cancer marker(s) e.g. , a heterogeneous nuclear ribonucleoprotein (e.g. , A2/B (hnRNPA2/B1 )), L-lactate dehydrogenase A (LDHA), anterior gradient protein2 homolog, nucleophosmin, cystatin B, hepatoma derived growth factor and/or calcyphosin
- the amount of endometrial cancer marker(s) e.g. , fibrinogen (e.g.
- apolipoprotein A1 alpha, beta, and/or gamma chain
- apolipoprotein A1 haptoglobin (HP)
- serpin H 1 heat-shock protein 47 kD (hsp47)
- the methods are non-invasive for detecting endometrial disease which in turn allow for diagnosis of a variety of conditions or diseases associated with the endometrium.
- the invention provides a non-invasive non-surgical method for detection, diagnosis or prediction of endometrial disease, including, without limitation, endometrial cancer and EmCa, in a subject, by: (a) obtaining a sample of body fluids, including, without limitation, blood, plasma, serum, urine or saliva, and/or a tissue sample from the subject; (b) subjecting the sample to a procedure to detect endometrial cancer markers in the body fluids and/or the tissue sample;
- endometrial disease is detected, diagnosed, or predicted by determination of increased levels of protein endometrial cancer markers, including, without limitation, one or more of the protein endometrial cancer markers indicated to be up-regulated in Table 5a, Table 5b, Table 8, and/or Table 9, when compared to such levels obtained from the control.
- endometrial disease is detected, diagnosed, or predicted by determination of decreased levels of protein endometrial cancer markers, including, without limitation, one or more of the protein endometrial caner marked indicated to be down-regulated in Table 5a, Table 5b, Table 8, and/or Table 9, when compared to such levels obtained from the control.
- protein endometrial cancer markers including, without limitation, one or more of the protein endometrial caner marked indicated to be down-regulated in Table 5a, Table 5b, Table 8, and/or Table 9, when compared to such levels obtained from the control.
- Another aspect of the invention provides for a method for assessing the aggressiveness or indolence of an endometrial disease in particular cancer (e.g. , staging), the method by comparing: (a) levels of endometrial cancer markers with the endometrial disease in a patient sample; and (b) normal levels of the endometrial cancer markers in a control sample.
- a method for assessing the aggressiveness or indolence of an endometrial disease in particular cancer e.g. , staging
- the method by comparing: (a) levels of endometrial cancer markers with the endometrial disease in a patient sample; and (b) normal levels of the endometrial cancer markers in a control sample.
- a significant difference between the levels in the sample and the normal levels is an indication that the endometrial disease, in particular cancer, is aggressive or indolent.
- the levels of endometrial cancer markers are higher than normal levels.
- the levels of endometrial cancer markers are lower than normal levels.
- the endometrial cancer markers are protein endometrial cancer markers or polynucleotide endometrial cancer markers.
- aggressiveness or indolence of an endometrial disease assessed by determination of increased levels of protein endometrial cancer markers including, without limitation, one or more of the protein endometrial cancer markers indicated to be up-regulated in Table 5a, Table 5b, Table 8, and/or Table 9, when compared to such levels obtained from the control.
- aggressiveness or indolence of an endometrial disease assessed by determination of decreased levels of protein endometrial cancer markers including, without limitation, one or more of the protein endometrial caner marked indicated to be down- regulated in Table 5a, Table 5b, Table 8, and/or Table 9, when compared to such levels obtained from the control.
- the invention provides a method for diagnosing and/or monitoring EmCa by comparing: (a) levels of LDHA or a polynucleotide encoding LDHA in a sample from the patient; and (b) normal levels of LDHA or a polynucleotide encoding the same in samples of the same type obtained from control patients not afflicted with endometrial cancer or having a different stage of endometrial cancer, wherein altered levels of LHDA or polynucleotides encoding same compared with the corresponding normal levels is an indication that the patient is afflicted with EmCa.
- the method compares patient-sample levels and normal levels of phosphoglycerate kinase, apolipoprotein A1 , fibrinogen (e.g. , alpha, beta, and/or gamma chain), serpin H1 (or heat- shock protein 47 kD (hsp47), a heterogeneous nuclear ribonucleoprotein (e.g. , A2/B (hnRNPA2/B1 )), or haptoglobin (HP)), or a polynucleotide encoding same.
- Exemplary EmCa which may be diagnosed and/or monitored by these methods include, without limitation, endometrioid EmCa, serous EmCa, or clear cell EmCa.
- the invention provides a method for determining whether a cancer has metastasized or is likely to metastasize in the future, by comparing: (b) levels of endometrial cancer markers in a patient sample; and (b) normal levels (or non-metastatic levels) of the endometrial cancer markers in a control sample.
- a significant difference between the levels in the patient sample and the normal levels is an indication that the cancer has metastasized or is likely to metastasize in the future.
- the invention provides a method for monitoring the progression of endometrial disease, in particular endometrial cancer in a patient, by: (a) detecting endometrial cancer markers or polynucleotides encoding the markers associated with the disease in a sample from the patient at a first time point; (b) repeating step (a) at a subsequent point in time; and (c) comparing the levels detected in (a) and (b), and therefrom monitoring the progression of the endometrial disease.
- the invention contemplates a method for determining the effect of an environmental factor on the endometrial tissue, or endometrial disease, by comparing endometrial cancer endometrial cancer markers in the presence and absence of the environmental factor.
- One embodiment of such method uses protein endometrial cancer marker or polynucleotide endometrial cancer markers.
- the invention also provides a method for assessing the efficacy of a test agent for inhibiting endometrial disease, and a method of selecting an agent for inhibiting endometrial disease.
- the invention provides for a method for assessing the carcinogenic potential of a test compound.
- the invention further contemplates a method of assessing the potential of a test compound to contribute to an endometrial disease by: (a) maintaining separate aliquots of endometrial diseased cells in the presence and absence of the test compound; and (b) comparing the levels of endometrial cancer markers associated with the disease in each of the aliquots.
- the endometrial cancer markers are protein endometrial cancer markers or polynucleotide endometrial cancer markers.
- the invention further relates to a method of assessing the efficacy of a therapy for inhibiting endometrial disease in a patient by comparing: (a) levels of endometrial cancer markers associated with disease in a first sample from the patient obtained from the patient prior to providing at least a portion of the therapy to the patient; and (b) levels of endometrial cancer markers associated with disease in a second sample obtained from the patient following therapy.
- a significant difference between the levels of endometrial cancer markers in the second sample relative to the first sample is an indication that the therapy is efficacious for inhibiting endometrial disease.
- the endometrial cancer markers are protein endometrial cancer markers or polynucleotide endometrial cancer markers.
- the method is used to assess the efficacy of a therapy for inhibiting endometrial disease, including, without limitation, endometrial cancer or EmCa, where lower levels of endometrial cancer markers in the second sample relative to the first sample, is an indication that the therapy is efficacious for inhibiting the disease.
- the endometrial cancer markers are protein endometrial cancer markers or polynucleotide endometrial cancer markers.
- the "therapy” may be any therapy for treating endometrial disease, in particular endometrial cancer, including but not limited to therapeutics, radiation, immunotherapy, gene therapy, and surgical removal of tissue. Therefore, the method can be used to evaluate a patient before, during, and after therapy.
- Certain methods of the invention employ binding agents that specifically recognize endometrial cancer markers.
- the invention provides methods for determining the presence or absence of endometrial disease, in particular endometrial cancer, in a patient, by: (a) contacting a biological sample obtained from a patient with one or more binding agent that specifically binds to one or more endometrial cancer markers associated with the disease; and (b) detecting in the sample an amount of marker that binds to the binding agent, relative to a predetermined standard or cut-off value, and therefrom determine the presence or absence of endometrial disease in the patient from such results.
- the invention relates to a method for diagnosing and monitoring an endometrial disease, in particular endometrial cancer, in a subject by quantifying one or more endometrial cancer markers associated with the disease in a biological sample from the subject, by: (a) reacting the biological sample with one or more binding agent specific for the endometrial cancer markers that are directly or indirectly labelled with a detectable substance; and (b) detecting the detectable substance.
- antibodies are used as binding agents to recognize the protein endometrial cancer markers.
- polynucleotides are used as binding agents to recognize the polynucleotide endometrial cancer markers.
- the invention provides a method of using an antibody to detect expression of one or more protein endometrial cancer marker in a sample, by: (a) combining antibodies specific for one or more protein endometrial cancer marker with a sample under conditions which allow the formation of antibody-protein marker complexes; and (b) detecting complex formation, wherein complex formation indicates expression of the protein endometrial cancer marker in the sample wherein expression may be compared with standards and is diagnostic of an endometrial disease, in particular endometrial cancer.
- Embodiments of the invention may also involve: (a) reacting a biological sample from a subject with antibodies specific for one or more endometrial cancer markers which are directly or indirectly labelled with an enzyme; (b) adding a substrate for the enzyme wherein the substrate is selected so that the substrate, or a reaction product of the enzyme and substrate forms fluorescent complexes; (c) quantifying one or more endometrial cancer markers in the sample by measuring fluorescence of the fluorescent complexes; and (d) comparing the quantified levels to levels obtained for other samples from the subject patient, or control subjects.
- the quantified levels are compared to levels quantified for control subjects, such as normal or benign tumours, without an endometrial disease, particularly endometrial cancer, wherein an increase in endometrial cancer marker levels compared with the control subjects is indicative of endometrial disease.
- the quantified levels are compared to levels quantified for control subjects, such as normal or benign tumours, without an endometrial disease, particularly endometrial cancer, wherein a decrease in endometrial marker levels compared with the control subjects is indicative of endometrial disease.
- the invention provides a method by: (a) incubating a biological sample with first antibodies specific for one or more endometrial cancer markers which are directly or indirectly labelled with a detectable substance, and second antibodies specific for one or more endometrial cancer markers which are immobilized; (b) detecting the detectable substance thereby quantifying endometrial cancer markers in the biological sample; and (c) comparing the quantified endometrial cancer markers with levels for a pre-determined standard.
- the standard may correspond to levels quantified for samples from control subjects without endometrial cancer (normal or benign), with a different disease stage, or from other samples of the subject.
- increased levels of endometrial cancer markers as compared to the standard may be indicative of endometrial cancer.
- lower levels of endometrial cancer markers as compared to a standard may be indicative of endometrial cancer.
- Protein endometrial cancer marker levels can be determined by constructing an antibody microarray in which binding sites include immobilized, preferably monoclonal, antibodies specific to a substantial fraction of marker-derived protein endometrial cancer markers of interest.
- Other methods of the invention employ one or more polynucleotides capable of hybridizing to one or more polynucleotide endometrial cancer markers.
- methods can be used to monitor an endometrial disease, in particular endometrial cancer, by detecting polynucleotide endometrial cancer markers associated with the disease.
- the invention relates to a method for diagnosing and monitoring an endometrial disease, including, without limitation, endometrial cancer, EmCa or related condition, in a sample from a subject, by: (a) isolating nucleic acids, preferably mRNA, from the sample; and (b) detecting polynucleotide endometrial cancer markers associated with endometrial diseases, wherein the presence of different levels of polynucleotide endometrial cancer markers in the sample compared to a standard or control may be indicative of endometrial disease, disease stage, and/or a negative or positive prognosis, such as longer progression-free and overall survival.
- polynucleotide endometrial cancer marker positive tumours are a negative diagnostic indicator.
- Positive tumors can be indicative of endometrial cancer, advanced stage disease, lower progression-free survival, and/or overall survival.
- polynucleotide endometrial cancer marker negative tumours are a negative diagnostic indicator. Negative tumors can be indicative of endometrial cancer, advanced stage disease, lower progression-free survival, and/or overall survival.
- the invention provides methods for determining the presence or absence of an endometrial disease in a subject, by: (a) detecting in the sample levels of nucleic acids that hybridize to one or more polynucleotide endometrial cancer markers associated with the disease; (b) comparing the levels with a pre-determined standard or cut-off value; and (c) determining the presence or absence of endometrial disease in the subject.
- the invention provides methods for determining the presence or absence of endometrial disease, such as EmCa or another endometrial cancer, in a subject, by: (a) contacting a sample obtained from the subject with oligonucleotides that hybridize to
- polynucleotides encoding endometrial cancer markers (b) detecting in the sample a level of nucleic acids that hybridize to the polynucleotides relative to a predetermined cut-off value; and (c) determining the presence or absence of endometrial cancer in the subject.
- the amount of polynucleotides that are mRNA are detected via polymerase chain reaction using, for example, oligonucleotide primers that hybridize to one or more polynucleotide endometrial cancer markers or complements of such polynucleotides.
- the amount of mRNA is detected using a hybridization technique, employing oligonucleotide probes that hybridize to one or more polynucleotide endometrial cancer markers or complements of such polynucleotides.
- the method may be carried out by combining isolated mRNA with reagents to convert to cDNA according to standard methods well known in the art, treating the converted cDNA with amplification reaction reagents (such as cDNA PCR reaction reagents) in a container along with an appropriate mixture of nucleic acid primers; reacting the contents of the container to produce amplification products; and analyzing the amplification products to detect the presence of one or more of the polynucleotide endometrial cancer markers in the sample.
- the analyzing step may be accomplished using Northern Blot analysis to detect the presence of polynucleotide endometrial cancer markers in the sample.
- the analysis step may be further accomplished by quantitatively detecting the presence of polynucleotide endometrial cancer markers in the amplification product, and comparing the quantity of marker detected against a panel of expected values for the known presence or absence of such markers in normal and malignant tissue derived using similar primers.
- the invention provides a method wherein mRNA is detected by: (a) isolating mRNA from a sample and combining the mRNA with reagents to convert it to cDNA; (b) treating the converted cDNA with amplification reaction reagents and nucleic acid primers that hybridize to one or more of the polynucleotide endometrial cancer markers endometrial cancer marker to produce amplification products; (c) analyzing the amplification products for determining the amount of mRNA present encoding the protein endometrial cancer marker; and (d) comparing the determined amount of mRNA to an amount detected against a panel of expected values for normal and diseased tissue (e.g. , malignant tissue) derived using similar methods.
- a panel of expected values for normal and diseased tissue e.g. , malignant tissue
- RT-PCR can be used to amplify the mRNA for protein endometrial cancer markers for detection and analysis.
- Other embodiments of the invention use quantitative RT-PCR to quantitatively determine amount of mRNA for protein endometrial cancer markers.
- Further embodiments of the invention use real time RT-PCR for quantification and analysis.
- the methods described herein utilize the polynucleotide endometrial cancer markers placed on a microarray so that the expression status of each of the markers may be assessed.
- the invention provides a microarray including a defined set of genes encoding protein endometrial cancer markers (i.e. , at least 2, 3, 4, or 5 genes listed in Table 5a, Table 5b, Table 8, and/or Table 9) whose expression is significantly altered by endometrial disease.
- the invention further relates to the use of the microarray as a prognostic tool to predict or diagnose endometrial disease.
- the endometrial microarray discriminates between endometrial diseases resulting from different etiologies.
- the invention provides for oligonucleotide arrays including marker sets described herein.
- the microarrays provided by the present invention may include probes to markers able to distinguish endometrial disease.
- the invention provides oligonucleotide arrays including probes to a subset or subsets of at least 5 to 10 polynucleotide endometrial cancer markers, up to a full set of markers which distinguish endometrial disease.
- the invention also contemplates a method by administering to cells or tissues imaging agents that carry labels for imaging and bind to endometrial cancer markers and optionally other markers of an endometrial disease, and then imaging the cells or tissues.
- the invention provides an in vivo method by administering to a subject an agent that has been constructed to target one or more endometrial cancer markers.
- the invention contemplates an in vivo method by
- administering to a mammal one or more agent that carries a label for imaging and binds to one or more endometrial cancer markers, and then imaging the mammal.
- the invention provides an in vivo method for imaging endometrial cancer, by: (a) injecting a patient with an agent that binds to one or more endometrial cancer markers, the agent carrying a label for imaging the endometrial cancer; (b) allowing the agent to incubate in vivo and bind to one or more endometrial cancer markers associated with the endometrial cancer; and (c) detecting the presence of the label localized to the cancer.
- the agent is an antibody that recognizes a protein endometrial cancer marker.
- the agent is a chemical entity that recognizes an endometrial cancer marker.
- An agent carries a label to image an endometrial marker and optionally other markers.
- labels useful for imaging are radiolabels, fluorescent labels (e.g. , fluorescein and rhodamine), nuclear magnetic resonance active labels, positron emitting isotopes detectable by a positron emission tomography (“PET”) scanner, chemiluminescers such as luciferin, and enzymatic markers such as peroxidase or phosphatase.
- PET positron emission tomography
- chemiluminescers such as luciferin
- enzymatic markers such as peroxidase or phosphatase.
- Short-range radiation emitters such as isotopes detectable by short-range detector probes can also be employed.
- the invention also contemplates the localization or imaging methods described herein using multiple markers for an endometrial disease, including, without limitation, endometrial cancer, EmCa or related conditions.
- kits for carrying out the methods of the invention are for assessing whether a patient is afflicted with an endometrial disease, including, without limitation, endometrial cancer, EmCa or related conditions, and the kit includes reagents for assessing one or more endometrial cancer markers.
- the invention further provides kits including marker sets described herein.
- the kit contains a microarray ready for hybridization to target endometrial cancer polynucleotide markers and the software needed for the data analysis.
- the invention also provides a diagnostic composition including protein endometrial cancer markers or polynucleotide encoding the markers.
- a composition is also provided including a probe that specifically hybridizes to polynucleotide endometrial cancer markers, or a fragment thereof, or an antibody specific for protein endometrial cancer markers or a fragment thereof.
- a composition is provided including polynucleotide endometrial cancer markers specific primer pairs capable of amplifying such polynucleotides using polymerase chain reaction methodologies.
- the probes, primers or antibodies can be labelled with a detectable substance.
- the invention relates to therapeutic applications for endometrial diseases, in particular endometrial cancer, employing protein endometrial cancer markers, and polynucleotide encoding the markers, and/or binding agents for such markers.
- the invention relates to compositions including markers or parts thereof associated with an endometrial disease, or antibodies specific for protein endometrial cancer markers associated with an endometrial disease, and a pharmaceutically acceptable carrier, excipient, or diluent.
- Another of the invention provides a method for treating or preventing an endometrial disease, in particular endometrial cancer, in a patient, by administering to a patient in need thereof, markers or parts thereof associated with endometrial disease, antibodies specific for protein endometrial cancer markers, or a composition of the invention.
- the invention provides a method of treating a patient afflicted with or at risk of developing an endometrial disease (e.g. , endometrial cancer) by inhibiting expression of polynucleotide endometrial cancer markers.
- An embodiment of the invention includes antisense oligonucleotides complementary to polynucleotide endometrial cancer markers delivered to diseased cells for regulation of gene expression.
- the invention provides antibodies specific for protein endometrial cancer markers associated with a disease, such as EmCa, that can be used therapeutically to destroy or inhibit the disease, such as growth of marker expressing cancer cells), or to block marker activity associated with a disease.
- a disease such as EmCa
- the endometrial cancer markers may be used in various immunotherapeutic methods to promote immune-mediated destruction or growth inhibition of tumors expressing the endometrial cancer markers.
- the invention also contemplates a method of using endometrial cancer markers or parts thereof, or antibodies specific for the protein endometrial cancer markers in the preparation or manufacture of a medicament for the prevention or treatment of an endometrial disease, including, without limitation, endometrial cancer, EmCa or related conditions.
- Another aspect of the invention relates to the use of protein endometrial cancer markers, peptides derived therefrom, or chemically produced (synthetic) peptides, or any combination of these molecules, for use in the preparation of vaccines to prevent an endometrial disease and/or to treat an endometrial disease.
- the invention contemplates vaccines for stimulating or enhancing in a subject to whom the vaccine is administered, production of antibodies directed against one or more endometrial cancer markers.
- the invention also provides a method for stimulating or enhancing in a subject production of antibodies directed against one or more endometrial cancer markers by administering to the subject a vaccine of the invention in a dose effective for stimulating or enhancing production of the antibodies.
- the invention further provides a method for treating, preventing, or delaying recurrence of an endometrial disease, including, without limitation, endometrial cancer, EmCa or related conditions by administering to the subject a vaccine of the invention in a dose effective for treating, preventing, or delaying recurrence of an endometrial disease including, without limitation, endometrial cancer, EmCa or related conditions.
- the invention contemplates the methods, compositions, and kits described herein using additional markers associated with an endometrial disease, including, without limitation, endometrial cancer, EmCa or related conditions.
- the methods described herein may be modified by including reagents to detect the additional markers, or polynucleotides for the markers.
- the invention contemplates the methods described herein using multiple markers for endometrial cancer, such as EmCa. Therefore, the invention contemplates a method for analyzing a biological sample for the presence of endometrial cancer markers and other markers that are specific indicators of cancer, in particular endometrial cancer.
- the methods described herein may be modified by including reagents to detect the additional markers, or nucleic acids for the additional markers.
- the methods, compositions, and kits use a panel of the protein endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9 or polynucleotides encoding same.
- biological samples may be obtained from tissues, extracts, cell cultures, cell lysates, lavage fluid, or physiological fluids.
- the sample is an endometrial tumour tissue.
- iTRAQ isobaric tags for relative and absolute quantification
- LC liquid chromatography
- MS / MS tandem mass spectrometry
- PCM potential cancer marker
- EmCa endometrial carcinoma
- LCM laser capture microdissection
- PBS phosphate- buffered saline
- SCX strong cation exchange
- ID internal diameter
- RP reverse phase
- IDA information-dependent acquisition
- TBS tris-buffered saline
- AUC area under the curve
- RSD relative standard deviation
- TMA tissue microarray
- AAT alpha-antitrypsin (serpin A1 )
- AFP alpha fetoprotein
- APO A-1 apolipoprotein A-l
- ATP5A1 ATP synthase subunit alpha, mitochondrial
- ATP5A1 ATP synthase subunit alpha, mitochondrial
- ATP5B ATP synthase subunit beta, mitochondrial
- EmCa endometrial carcinoma
- FDR false discovery rate
- FBPA A fructose-bisphosphate aldolase A
- FIGO Federation Internationale de Gynecologie et d'Obstetrique or International Federation of Gynecology and Obstetrics
- HCC hepatocellular carcinoma
- hnRNPA2B1 heterogeneous nuclear ribonucleoproteins A2/B1 (splice variants of A1 );
- HP haptoglobin
- HRPC hormone-refractory prostate cancer
- HSPC hormone- sensitive prostate cancer
- IPA Ingenuity Pathways Analysis
- LDHA L-lactate dehydrogenase A
- OvCa Ovarian carcinoma
- PGK1 phosphoglycerate kinase 1
- PK pyruvate kinase isoen
- Endometrial disease refers to any disorder, disease, condition, syndrome, or combination of manifestations or symptoms recognized or diagnosed as a disorder of the endometrium, including, without limitation, inflammation, cancer precursors, and carcinoma.
- Endometrial disease includes malignant endometrial cancer, including, without limitation, endometrial carcinomas (EmCa) such as Type I EmCa, endometrial serous carcinoma, and endometrial clear cell carcinoma.
- EmCa endometrial carcinomas
- Type I EmCa endometrial serous carcinoma
- endometrial clear cell carcinoma endometrial clear cell carcinoma
- sample means a material known or suspected of expressing or containing one or more endometrial carcinoma polynucleotide or polypeptide markers.
- a test sample can be used directly as obtained from the source or following a pretreatment to modify the character of the sample.
- the sample can be derived from any biological source, such as tissues, extracts, or cell cultures, including cells (e.g. , tumor cells), cell lysates, and physiological fluids, such as, for example, whole blood, plasma, serum, saliva, ocular lens fluid, cerebral spinal fluid, sweat, urine, milk, ascites fluid, synovial fluid, peritoneal fluid, lavage fluid, and the like.
- the sample can be obtained from animals, preferably mammals, most preferably humans.
- the sample can be treated prior to use, such as preparing plasma from blood, diluting viscous fluids, and the like.
- Methods of treatment can involve freezing, fixation, embedding in paraffin or OCT, sonication, centrifugation, filtration, distillation, extraction, concentration, inactivation of interfering components, the addition of reagents, and the like.
- the sample is a mammalian tissue sample.
- the tissue is endometrial tissue.
- the sample is an endometrial tumour tissue.
- the sample is a human physiological fluid.
- the sample is human serum.
- RNA RNA transcribed from cDNA
- mRNA messenger RNA
- RNA transcribed from cDNA i.e. , cRNA; see, for example. , Linsley & Schelter, Patent Application Ser. No. 09 / 411 ,074, or Patent Nos. 5,545, 522; 5,891 ,636; or 5,716,785).
- RNA may be isolated from eukaryotic cells by procedures involving lysis of the cells and denaturation of the proteins contained in the cells.
- RNA can be separated from DNA by organic extraction, for example, with hot phenol or phenol / chloroform / isoamyl alcohol.
- RNAs such as transfer RNA (tRNA) and ribosomal RNA (rRNA).
- Most mRNAs contain a poly(A) tail at their 3' end allowing them to be enriched by affinity chromatography, for example, using oligo(dT) or poly(U) coupled to a solid support, such as cellulose or SephadexTM (see Ausubel et al. , eds. , 1994, Current Protocols in Molecular Biology, Vol. 2, Current Protocols Publishing, New York).
- Bound poly(A)+mRNA is eluted from the affinity column using 2 mM EDTA / 0.1 % SDS.
- a sample of RNA can include a plurality of different RNA molecules each with a different nucleotide sequence.
- the RNA molecules contain mRNA molecules.
- the mRNA molecules in the RNA sample include at least 100 different nucleotide sequences.
- Target polynucleotides can be detectably labelled at one or more nucleotides using methods known in the art.
- the label is preferably uniformly incorporated along the length of the RNA, and more preferably, is carried out at a high degree of efficiency.
- the detectable label can be a luminescent label, fluorescent label, bio-luminescent label, chemiluminescent label, radiolabel, and colorimetric label.
- the label is a fluorescent label, such as a fluorescein, a phosphor, a rhodamine, or a polymethine dye derivative.
- fluorescent labels include, for example, fluorescent phosphoramidites, such as FluorePrime (Amersham Pharmacia, Piscataway, N.J. ), Fluoredite (Millipore, Bedford, Mass. ), FAM (ABI, Foster City, Calif. ), and Cy3 or Cy5 (Amersham Pharmacia, Piscataway, N.J. ).
- fluorescent phosphoramidites such as FluorePrime (Amersham Pharmacia, Piscataway, N.J. ), Fluoredite (Millipore, Bedford, Mass. ), FAM (ABI, Foster City, Calif. ), and Cy3 or Cy5 (Amersham Pharmacia, Piscataway, N.J. ).
- Target polynucleotides from a patient sample can be labelled differentially from polynucleotides of a standard.
- the standard can include target polynucleotides from normal individuals (i.e. , those not afflicted with or pre-disposed to endometrial disease), in particular pooled from samples from normal individuals.
- the target polynucleotides can be derived from the same individual, but taken at different time points, and thus indicate the efficacy of a treatment by a change in expression of the markers, or lack thereof, during and after the course of treatment.
- subject refers to a warm-blooded animal such as a mammal. In particular, the terms refer to a human. A subject, individual or patient may be afflicted with or suspected of having or being pre-disposed to endometrial disease or a condition as described herein.
- the terms also includes domestic animals bred for food or as pets, including horses, cows, sheep, poultry, fish, pigs, cats, dogs, and zoo animals.
- Methods herein for administering an agent or composition to subjects / individuals / patients contemplate treatment as well as prophylactic use.
- Typical subjects for treatment include persons susceptible to, suffering from or that have suffered a condition or disease described herein.
- suitable subjects for treatment in accordance with the invention are persons that are susceptible to, suffering from or that have suffered endometrial cancer.
- endometrial cancer marker refers to a marker associated with normal or diseased endometrial tissue and includes one or more of polypeptides that are differentially expressed in cancer samples as compared to normal samples and polynucleotides encoding these polypeptides.
- the expression includes native-sequence polypeptides and nucleotides, isoforms, chimeric polypeptides, complexes, and all homologs, fragments, precursors, modified forms, and derivatives of the markers, as applicable.
- An endometrial cancer marker may be associated with an endometrial disease, including endometrial cancer (e.g. , EmCa).
- endometrial cancer marker includes a marker associated with protein endometrial cancer marker listed in Table 5a, Table 5b, Table 8, and/or Table 9, polynucleotides encoding the proteins listed in Table 5a, Table 5b, Table 8, and/or Table 9, and precursors thereof.
- a "native-sequence polypeptide” includes a polypeptide having the same amino acid sequence of a polypeptide derived from nature. Such native-sequence polypeptides can be isolated from nature or can be produced by recombinant or synthetic means. The term specifically encompasses naturally occurring truncated or secreted forms of a polypeptide, polypeptide variants including naturally occurring variant forms (e.g. , alternatively spliced forms or splice variants), and naturally occurring allelic variants.
- polypeptide variant refers to a polypeptide having at least about 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, 97%, 98%, or 99% amino acid sequence identity, particularly at least about 70-80%, more particularly at least about 85%, still more particularly at least about 90%, most particularly at least about 95% amino acid sequence identity with a native- sequence polypeptide.
- Particular polypeptide variants have at least 70-80%, 85%, 90%, 95% amino acid sequence identity to the sequences identified in Table 5a, Table 5b, Table 8, and/or Table 9.
- variants include, for instance, polypeptides wherein one or more amino acid residues are added to, or deleted from, the N- or C-terminus of the full-length or mature sequences of the polypeptide, including variants from other species, but exclude a native-sequence polypeptide.
- variants retain the immunogenic activity of the corresponding native-sequence polypeptide.
- Percent identity of two amino acid sequences, or of two nucleic acid sequences is defined as the percentage of amino acid residues or nucleotides in a candidate sequence that are identical with the amino acid residues in a polypeptide or nucleic acid sequence, after aligning the sequences and introducing gaps, if necessary, to achieve the maximum percent sequence identity, and not considering any conservative substitutions as part of the sequence identity. Alignment for purposes of determining percent amino acid or nucleic acid sequence identity can be achieved in various conventional ways, for instance, using publicly available computer software including the GIG program package (Devereux J. et al.
- BLAST X program is publicly available from NCBI and other sources (BLAST Manual, Altschul, S. et al. , NCBI NLM NIH Bethesda, Md. 20894; Altschul, S. et al. , J. Mol. Biol. 215: 403-410, 1990). Skilled artisans can determine appropriate parameters for measuring alignment, including any algorithms needed to achieve maximal alignment over the full length of the sequences being compared. Methods to determine identity and similarity are codified in publicly-available computer programs.
- allelic variant may also be created by introducing substitutions, additions, or deletions into a polynucleotide encoding a native polypeptide sequence such that one or more amino acid substitutions, additions, or deletions are introduced into the encoded protein. Mutations may be introduced by standard methods, such as site-directed mutagenesis and PCR-mediated mutagenesis. In an embodiment, conservative substitutions are made at one or more predicted non-essential amino acid residues. A "conservative amino acid substitution" is one in which an amino acid residue is replaced with an amino acid residue with a similar side chain.
- Amino acids with similar side chains include amino acids with basic side chains (e.g., Lys, Arg, His), acidic side chains (e.g. , Asp, Glu), uncharged polar side chains (e.g. , Gly, Asp, Glu, Ser, Thr, Tyr, and Cys), non- polar side chains (e.g. , Ala, Val, Leu, Iso, Pro, Trp), beta-branched side chains (e.g. , Thr, Val, Iso), and aromatic side chains (e.g. , Tyr, Phe, Trp, His). Mutations can also be introduced randomly along part or all of the native sequence, for example, by saturation mutagenesis. Following mutagenesis the variant polypeptide can be recombinantly expressed and the activity of the polypeptide may be determined.
- basic side chains e.g., Lys, Arg, His
- acidic side chains e.g. , Asp, Glu
- Polypeptide variants include polypeptides and amino acid sequences sufficiently identical to or derived from the amino acid sequence of a native polypeptide which includes fewer amino acids than the full length polypeptides.
- a portion of a polypeptide can be a polypeptide which is, for example, 10, 15, 20, 25, 30, 35, 40, 45, 50, 60, 70, 80, 90, 100 or more amino acids in length.
- Portions in which regions of a polypeptide are deleted can be prepared by recombinant techniques and can be evaluated for one or more functional activities such as the ability to form antibodies specific for a polypeptide.
- allelic variant may contain conservative amino acid substitutions from the native polypeptide sequence or it may contain a substitution of an amino acid from a corresponding position in a polypeptide homolog, for example, a murine polypeptide.
- An endometrial cancer marker may be part of a chimeric or fusion protein.
- a "chimeric protein” or “fusion protein” includes all or part (preferably biologically active) of an endometrial cancer marker operably linked to a heterologous polypeptide (i.e. , a polypeptide other than an endometrial diseases marker).
- a heterologous polypeptide i.e. , a polypeptide other than an endometrial diseases marker.
- the term "operably linked” is intended to indicate that an endometrial cancer marker and the heterologous polypeptide are fused in-frame to each other.
- the heterologous polypeptide can be fused to the N-terminus or C-terminus of an endometrial cancer marker.
- a useful fusion protein is a GST fusion protein in which an endometrial cancer marker is fused to the C-terminus of GST sequences. Another example of a fusion protein is an
- immunoglobulin fusion protein in which all or part of an endometrial cancer marker is fused to sequences derived from a member of the immunoglobulin protein family.
- Chimeric and fusion proteins can be produced by standard recombinant DNA techniques.
- a modified form of a polypeptide referenced herein includes modified forms of the polypeptides and derivatives of the polypeptides, including post-translationally modified forms such as glycosylated, phosphorylated, acetylated, methylated or lapidated forms of the polypeptides.
- an N-terminal methionine may be cleaved from a polypeptide, and a new N-terminal residue may or may not be acetylated.
- the first residue, methionine can be cleaved and the second first residue, alanine can be N-acetylated.
- An endometrial cancer marker may be prepared by recombinant or synthetic methods, or isolated from a variety of sources, or by any combination of these and similar techniques.
- polynucleotides encoding the protein markers include polynucleotides that encode the protein endometrial cancer markers described herein, polynucelotides that encode a native-sequence polypeptide, a polypeptide variant, including a portion of a polypeptide, a chimeric polypeptide, or an isoform, precursor, complex, modified form, or derivative of any of the polypeptide endometrial cancer markers, and any precursors thereof.
- a polynucleotide endometrial cancer marker includes or consists of one or more polynucleotides encoding the polypeptides listed in Table 5a, Table 5b, Table 8, and/or Table 9.
- Polynucleotide endometrial cancer markers include complementary nucleic acid sequences, and nucleic acids that are substantially identical to these sequences (e.g. , having at least about 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, 97%, 98%, or 99% sequence identity). Polynucleotide endometrial cancer markers also include sequences that differ from a native sequence due to degeneracy in the genetic code. As one example, DNA sequence polymorphisms within the nucleotide sequence of an endometrial cancer marker may result in silent mutations that do not affect the amino acid or nucleotide sequence.
- Variations in one or more nucleotides may exist among individuals within a population due to natural allelic variation.
- DNA sequence polymorphisms may also occur which lead to changes in the amino acid sequence of a polypeptide or the nucleotide sequence of a polynucleotide.
- Polynucleotide endometrial cancer markers also include nucleic acids that hybridize under stringent conditions, preferably high stringency conditions to endometrial cancer polynucleotide markers.
- Appropriate stringency conditions which promote DNA hybridization are known to those skilled in the art, or can be found in Current Protocols in Molecular Biology, John Wiley & Sons, N.Y. (1989), 6.3.1 -6.3.6.
- 6.0 x sodium chloride / sodium citrate (SSC) at about 45°C, followed by a wash of 2.0 x SSC at 50°C may be employed.
- the stringency may be selected based on the conditions used in the wash step.
- the salt concentration in the wash step can be selected from a high stringency of about 0.2 x SSC at 50°C.
- the temperature in the wash step can be at high stringency conditions, at about 65°C.
- Polynucleotide endometrial cancer markers further include truncated nucleic acids or nucleic acid fragments and variant forms of the nucleic acids that arise by alternative splicing of an mRNA corresponding to a DNA.
- the polynucleotide endometrial cancer markers are intended to include DNA and RNA, including, without limitation, mRNA, and can be either double stranded or single stranded.
- a polynucleotide may, but need not, include additional coding or non-coding sequences, or it may, but need not, be linked to other molecules and/or carrier or support materials.
- the polynucleotides for use in the methods of the invention may be of any length suitable for a particular method. In certain applications the term also includes antisense polynucleotides (e.g., RNA or DNA strand with complementary sequences in the reverse orientation to the polynucleotide endometrial cancer markers).
- Statistically different levels may represent levels that are higher or lower than the standard error of the detection assay.
- the levels may be 1.5, 2, 3, 4, 5, 6 or more times higher or lower than the control or standard.
- the levels may be significantly higher compared to the control or standard.
- the levels may be significantly lower compared to the control or standard.
- “Microarray” and “array” refer to nucleic acid or nucleotide arrays or protein or peptide arrays that can be used to detect biomolecules associated with endometrial cancer cell or tissue phase and endometrial disease, for instance to measure gene expression.
- arrays are made in research and manufacturing facilities worldwide, some of which are available commercially.
- spotted arrays and in situ synthesized arrays are two kinds of nucleic acid arrays that differ in the manner in which the nucleic acid materials are placed onto the array substrate.
- a widely used in situ synthesized oligonucleotide array is GeneChipTM made by Affymetrix, Inc.
- Oligonucleotide probes that are 20- or 25-base long can be synthesized in silico on the array substrate. These arrays can achieve high densities (e.g., more than 40,000 genes per cm2).
- spotted arrays have lower densities, but the probes, typically partial cDNA molecules, are much longer than 20- or 25-mers.
- Examples of spotted cDNA arrays include LifeArray made by Incyte Genomics and DermArray made by IntegriDerm (or Invitrogen). Pre-synthesized and amplified cDNA sequences are attached to the substrate of spotted arrays. Protein and peptide arrays also are known (see for example, Zhu et al. , Science 293:2101 (2001 )).
- Binding agent refers to a substance such as a polypeptide or antibody that specifically binds to one or more endometrial cancer markers.
- a substance “specifically binds" to one or more protein endometrial cancer markers if it reacts at a detectable level with one or more endometrial cancer markers, and does not react detectably with peptides containing an unrelated or different sequence. Binding properties may be assessed using an ELISA, which may be readily performed by those skilled in the art (see, for example, Newton et al. , Develop. Dynamics 197: 1 -13, 1993).
- a binding agent may be a ribosome, with or without a peptide component, an aptamer, an RNA molecule, or a polypeptide.
- a binding agent may be a polypeptide that includes one or more endometrial cancer marker sequence, a peptide variant thereof, or a non-peptide mimetic of such a sequence.
- an annexin A1 sequence may be a peptide portion of an annexin A1 that is capable of modulating a function mediated by annexin A1.
- An aptamer includes a DNA or RNA molecule that binds to nucleic acids and proteins.
- An aptamer that binds to a protein (or binding domain) of an endometrial cancer marker or a polynucleotide endometrial cancer marker can be produced using conventional techniques, without undue experimentation.
- aptamers Klug et al. , Mol. Biol. Reports 20:97-107 (1994); Wallis et al. , Chem. Biol. 2:543-552 (1995); Ellington, Curr. Biol. 4:427-429 (1994); Lato et al. , Chem. Biol. 2:291 -303 (1995); Conrad et al. , Mol. Div. 1 :69-78 (1995); and Uphoff et al. , Curr. Opin. Struct. Biol. 6:281 -287 (1996).
- Antibodies for use in the present invention include but are not limited to monoclonal or polyclonal antibodies, immunologically active fragments (e.g. , a Fab or (Fab)2 fragments), antibody heavy chains, humanized antibodies, antibody light chains, genetically engineered single chain Fv molecules (Ladner et al. , US Patent No. 4,946,778), chimeric antibodies, for example, antibodies which contain the binding specificity of murine antibodies, but in which the remaining portions are of human origin, or derivatives, such as enzyme conjugates or labelled derivatives.
- immunologically active fragments e.g. , a Fab or (Fab)2 fragments
- antibody heavy chains e.g., humanized antibodies, antibody light chains, genetically engineered single chain Fv molecules (Ladner et al. , US Patent No. 4,946,778)
- chimeric antibodies for example, antibodies which contain the binding specificity of murine antibodies, but in which the remaining portions are of human origin, or derivatives, such as enzyme conjugates
- Antibodies including monoclonal and polyclonal antibodies, fragments and chimeras, may be prepared using methods known to those skilled in the art. Isolated native or recombinant endometrial cancer markers may be utilized to prepare antibodies. (See, for example, Kohler et al. (1975), Nature 256:495-497; Kozbor et al. (1985), J. Immunol Methods 81 :31 -42; Cote et al. (1983), Proc Natl Acad Sci 80:2026-2030; and Cole et al. (1984), Mol Cell Biol 62: 109-120 for the preparation of monoclonal antibodies; Huse et al.
- antibodies are reactive against a protein endometrial cancer marker if they bind with a Ka of greater than or equal to 10-7 M.
- the invention provides a set of markers correlated with endometrial disease.
- the invention provides a set of markers identified as useful for detection, diagnosis, prevention and therapy of endometrial disease including or consisting of one or more of the markers listed in Table 5a, Table 5b, Table 8, and/or Table 9.
- the invention provides the endometrial cancer markers in Table 5a, Table 5b, Table 8, and/or Table 9 for detection, diagnosis and prognosis of an endometrial disease.
- the invention also provides a method of using endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9 to distinguish endometrial disease.
- the markers include or consist of 1 , 2, 3, 4 or more other markers listed in Table 5a, Table 5b, Table 8, and/or Table 9 or polynucleotides encoding same.
- a marker is provided which is selected from the group consisting of the polypeptides set forth in Table 5a, Table 5b, Table 8, and/or Table 9 which polypeptides are up-regulated biomarkers in endometrial.
- a marker is provided which is selected from the group consisting of at least one marker of Table 5a, Table 5b, Table 8, and/or Table 9.
- a marker is provided which is selected from the group consisting of at least one marker of Table 5a, Table 5b, Table 8, and/or Table 9 and at least 2, 3, 4, 5, 6, 7, 8, 9, or 10 polypeptides set forth in Table 5a, Table 5b, Table 8, and/or Table 9.
- the invention provides marker sets that distinguish endometrial disease and uses thereof.
- the invention provides a method for classifying an endometrial disease including detecting a difference in the expression of a first plurality of endometrial cancer markers relative to a control, the first plurality of endometrial cancer markers including or consisting of at least 2, 3, 4, or 5 of the markers listed in Table 5a, Table 5b, Table 8, and/or Table 9 or polynucleotides encoding same.
- a control includes markers derived from a sample from a patient with no endometrial disease.
- markers provided herein may be used alone or with other markers of endometrial disease, or with markers for other phenotypes or conditions. Additionally, all of the sequences provided herein are representative only; there may be other sequences for particular protein or coding sequences or related sequences. The invention is not intended to be limited to the sequences herein provided.
- Detection Methods A variety of methods can be employed for the diagnostic and prognostic evaluation of endometrial cancer status involving one or more endometrial cancer markers and the polynucleotides encoding the markers, and the identification of subjects with a predisposition to endometrial diseases or that are receptive to in vitro fertilization and embryo transfer procedures. Such methods may, for example, involve endometrial cancer polynucleotide markers, and fragments thereof, and binding agents (e.g. , antibodies) against one or more endometrial cancer markers, including peptide fragments.
- binding agents e.g. , antibodies
- polynucleotides and antibodies may be used, for example, for: (1 ) the detection of the presence of polynucleotide endometrial cancer marker mutations, or the detection of either over- or under-expression of protein endometrial cancer marker mRNA relative to a non-disorder state or different endometrial cancer cell or tissue phase, or the qualitative or quantitative detection of alternatively spliced forms of polynucleotide endometrial cancer marker transcripts which may correlate with certain conditions or susceptibility toward such conditions; and (2) the detection of either an over- or an under-abundance of one or more endometrial cancer markers relative to a non-disorder state or a different endometrial cell or tissue phase or the presence of a modified (e.g.
- the invention also contemplates a method for detecting an endometrial disease, in particular an endometrial cancer, including producing a profile of levels of one or more endometrial cancer marker associated with an endometrial disease and/or polynucleotides encoding the markers, and other markers associated with endometrial disease in cells from a patient, and comparing the profile with a reference to identify a profile for the test cells indicative of disease.
- the endometrial cancer markers are one or more of those listed in Table 5a, Table 5b, Table 8, and/or Table 9 or a precursor thereof.
- the methods described herein may be used to evaluate the probability of the presence of malignant cells, for example, in a group of cells freshly removed from a host. Such methods can be used to detect tumors, quantify their growth, and help in the diagnosis and prognosis of endometrial disease. The methods can be used to detect the presence of cancer metastasis, as well as confirm the absence or removal of all tumor tissue following surgery, cancer chemotherapy, and/or radiation therapy. They can further be used to monitor cancer chemotherapy and tumor reappearance.
- the methods described herein can be adapted for diagnosing and monitoring endometrial tissue status or an endometrial disease by detecting one or more endometrial cancer markers or polynucleotides encoding the markers in biological samples from a subject.
- These applications require that the amount of markers or polynucleotides quantified in a sample from a subject being tested be compared to a predetermined standard or cut-off value.
- the standard may correspond to levels quantified for another sample or an earlier sample from the subject, or levels quantified for a control sample.
- Levels for control samples from healthy subjects, different endometrial tissue phases, or subjects with an endometrial disease may be established by prospective and/or retrospective statistical studies.
- Healthy subjects who have no clinically evident disease or abnormalities may be selected for statistical studies. Diagnosis may be made by a finding of statistically different levels of detected endometrial cancer markers associated with disease or polynucleotides encoding same, compared to a control sample or previous levels quantified for the same subject.
- the methods described herein may also use multiple markers for an endometrial disease, in particular endometrial cancer, EmCa, or similar conditions. Therefore, the invention contemplates a method for analyzing a biological sample for the presence of one or more endometrial cancer markers and polynucleotides encoding the markers, and other markers that are specific indicators of an endometrial disease.
- the methods described herein may be modified by including reagents to detect the additional markers, or polynucleotides for the markers.
- nucleic Acid Methods As noted herein an endometrial disease or phase may be detected based on the level of endometrial cancer polynucleotide markers in a sample.
- PCR polymerase chain reaction
- Probes may be used in hybridization techniques to detect endometrial cancer polynucleotide markers.
- the technique generally involves contacting and incubating nucleic acids (e.g. , recombinant DNA molecules, cloned genes) obtained from a sample from a patient or other cellular source with a probe under conditions favorable for the specific annealing of the probes to complementary sequences in the nucleic acids. After incubation, the non-annealed nucleic acids are removed, and the presence of nucleic acids that have hybridized to the probe if any are detected.
- nucleic acids e.g. , recombinant DNA molecules, cloned genes
- Nucleotide probes for use in the detection of nucleic acid sequences in samples may be constructed using conventional methods known in the art. Suitable probes may be based on nucleic acid sequences encoding at least 5 sequential amino acids from regions of a polynucleotide endometrial cancer marker, preferably they include 10-200, more particularly 10-30, 10-40, 20-50, 40- 80, 50-150, 80-120 nucleotides in length.
- the probes may include DNA or DNA mimics (e.g. , derivatives and analogues) corresponding to a portion of an organism's genome, or complementary RNA or RNA mimics.
- Mimics are polymers including subunits capable of specific, Watson-Crick-like hybridization with DNA, or of specific hybridization with RNA.
- the nucleic acids can be modified at the base moiety, at the sugar moiety, or at the phosphate backbone.
- DNA can be obtained using standard methods such as polymerase chain reaction (PCR) amplification of genomic DNA or cloned sequences.
- PCR polymerase chain reaction
- a nucleotide probe may be labelled with a detectable substance such as a radioactive label that provides for an adequate signal and has sufficient half-life such as 32P, 3H, 14C or the like.
- detectable substances include antigens that are recognized by a specific labelled antibody, fluorescent compounds, enzymes, antibodies specific for a labelled antigen, and luminescent compounds.
- An appropriate label may be selected having regard to the rate of hybridization and binding of the probe to the nucleotide to be detected and the amount of nucleotide available for hybridization.
- Labelled probes may be hybridized to nucleic acids on solid supports such as nitrocellulose filters or nylon membranes as generally described in Sambrook et al.
- the nucleic acid probes may be used to detect endometrial cancer polynucleotide markers, preferably in human cells.
- the nucleotide probes may also be useful in the diagnosis of an endometrial disease involving one or more endometrial cancer polynucleotide markers, in monitoring the progression of such disorder, or monitoring a therapeutic treatment.
- the detection of polynucleotide encoding the markers may involve the amplification of specific gene sequences using an amplification method such as polymerase chain reaction (PCR), followed by the analysis of the amplified molecules using techniques known to those skilled in the art. Suitable primers can be routinely designed by one of skill in the art.
- PCR polymerase chain reaction
- At least two oligonucleotide primers may be employed in a PCR based assay to amplify a portion of a polynucleotide encoding one or more endometrial disease marker derived from a sample, wherein at least one of the oligonucleotide primers is specific for (i.e. , hybridizes to) a polynucleotide encoding the endometrial disease marker.
- the amplified cDNA is then separated and detected using techniques well known in the art, such as gel electrophoresis.
- primers and probes employed in the methods of the invention generally have at least about 60%, preferably at least about 75%, and more preferably at least about 90% identity to a portion of a polynucleotide encoding an endometrial disease marker; that is, they are at least 10 nucleotides, and preferably at least 20 nucleotides in length. In an embodiment the primers and probes are at least about 10-40 nucleotides in length.
- Hybridization and amplification techniques described herein may be used to assay qualitative and quantitative aspects of polynucleotide endometrial cancer marker expression.
- RNA may be isolated from a cell type or tissue known to express an endometrial cancer polynucleotide marker and tested utilizing the hybridization (e.g. , standard Northern analyses) or PCR techniques referred to herein.
- the primers and probes may be used in the above-described methods in situ (i.e. , directly on tissue sections (fixed and/or frozen) of patient tissue obtained from biopsies or resections).
- RNA is extracted from a sample tissue using standard techniques (e.g. , guanidine isothiocyanate extraction as described by Chomcynski and Sacchi, Anal. Biochem.
- cDNA 162 156-159, 1987
- the cDNA is used as a template for a polymerase chain reaction.
- the cDNA is hybridized to a set of primers, at least one of which is specifically designed against an endometrial disease marker sequence.
- a DNA polymerase is employed to extend from the primer, to synthesize a copy of the template.
- the DNA strands are denatured, and the procedure is repeated many times until sufficient DNA is generated to allow visualization by ethidium bromide staining and agarose gel electrophoresis.
- Real time RT-PCR and qRT-PCR may also be used.
- Amplification may be performed on samples obtained from a subject with a suspected endometrial disease and an individual who is not afflicted with an endometrial disease.
- the reaction may be performed on several dilutions of cDNA spanning at least two orders of magnitude.
- a statistically significant difference in expression in several dilutions of the subject sample as compared to the same dilutions of the non-disease sample may be considered positive for the presence of an endometrial disease.
- the invention provides methods for determining the presence or absence of an endometrial disease in a subject including (a) contacting a sample obtained from the subject with oligonucleotides that hybridize polynucleotide encoding the endometrial cancer markers; and (b) detecting in the sample a level of nucleic acids that hybridize to the polynucleotides relative to a predetermined cut-off value, and therefrom determining the presence or absence of an endometrial disease in the subject.
- the endometrial disease is cancer and endometrial cancer markers are one or more of those listed in Table 5a, Table 5b, Table 8, and/or Table 9 or a precursor thereof.
- the invention provides a method wherein mRNA for a protein endometrial cancer marker is detected by: (a) isolating RNA from a sample and combining the RNA with reagents to convert it to cDNA; (b) treating the converted cDNA with amplification reaction reagents and nucleic acid primers that hybridize to one or more mRNA encoding protein endometrial disease markers, to produce amplification products; (d) analyzing the amplification products to detect amounts of mRNA encoding protein endometrial disease markers; and (e) comparing the amount of such RNA to an amount detected against a panel of expected values for normal and malignant tissue derived using similar nucleic acid primers.
- Endometrial cancer marker-positive samples or alternatively higher levels in patients compared to a control may be indicative of late stage disease, and/or that the patient is not responsive to chemotherapy.
- negative samples or lower levels compared to a control e.g. , non-cancerous tissue or negative samples
- endometrial cancer marker-positive samples or alternatively higher levels in patients compared to a control may be indicative of late stage disease, and/or that the patient is not responsive to chemotherapy.
- negative samples or lower levels compared to a control e.g. , non-cancerous tissue or negative samples
- the invention provides methods for determining the presence or absence of endometrial cancer in a subject by: (a) contacting a sample obtained from the subject with oligonucleotides that hybridize to one or more polynucleotide endometrial cancer markers; and (b) detecting in the sample levels of nucleic acids that hybridize to the polynucleotides relative to a predetermined cut-off value, and therefrom determining the presence or absence of endometrial cancer in the subject.
- the endometrial cancer polynucleotide markers encode one or more polypeptides listed in Table 5a, Table 5b, Table 8, and/or Table 9.
- the invention provides a method wherein mRNA for L-lactate dehydrogenase A (LDHA), phosphoglycerate kinase, apolipoprotein A1 , fibrinogen (e.g. , alpha, beta, and/or gamma chain), serpin H 1 (or heat-shock protein 47 kD (hsp47), a heterogeneous nuclear ribonucleoprotein (e.g.
- LDHA L-lactate dehydrogenase A
- apolipoprotein A1 e.g. , alpha, beta, and/or gamma chain
- fibrinogen e.g. , alpha, beta, and/or gamma chain
- serpin H 1 or heat-shock protein 47 kD (hsp47)
- a heterogeneous nuclear ribonucleoprotein e.g.
- haptoglobin is detected by: (a) isolating mRNA from a sample and combining the mRNA with reagents to convert it to cDNA; (b) treating the converted cDNA with amplification reaction reagents and nucleic acid primers that hybridize to a polynucleotide encoding (LDHA), phosphoglycerate kinase, apolipoprotein A1 , fibrinogen (e.g.
- amplification products (c) analyzing the amplification products to detect an amount of the mRNA encoding (LDHA), phosphoglycerate kinase, apolipoprotein A1 , fibrinogen (e.g.
- Protein-positive samples or higher levels, especially significantly higher levels, of polynucleotides encoding a heterogeneous nuclear ribonucleoprotein e.g. , A2/B (hnRNPA2/B1 )
- a heterogeneous nuclear ribonucleoprotein e.g. , A2/B (hnRNPA2/B1 )
- L- lactate dehydrogenase A LDHA
- anterior gradient protein2 homolog e.g. , nucleophosmin
- cystatin B e.g., cystatin B
- hepatoma derived growth factor hepatoma derived growth factor
- calcyphosin e.g., calcyphosin in patients, as compared to a control (e.g. , normal or benign)
- Negative samples or lower levels of polynucleotides encoding fibrinogen e.g.
- apolipoprotein A1 alpha, beta, and/or gamma chain
- apolipoprotein A1 haptoglobin (HP)
- serpin H1 heat-shock protein 47 kD (hsp47) in patients, as compared to a control (e.g. , normal or benign)
- a control e.g. , normal or benign
- hsp47 heat-shock protein 47 kD
- Oligonucleotides or longer fragments derived from an endometrial cancer polynucleotide marker may be used as targets in a microarray.
- the microarray can be used to simultaneously monitor the expression levels of large numbers of genes and to identify genetic variants, mutations, and polymorphisms.
- the information from the microarray may be used to determine gene function, to understand the genetic basis of a disorder, to diagnose a disorder, and to develop and monitor the activities of therapeutic agents.
- the invention also includes an array including one or more endometrial cancer polynucleotide markers (in particular the markers listed in Table 5a, Table 5b, Table 8, and/or Table 9).
- the array can be used to assay expression of endometrial cancer polynucleotide markers in the array.
- the invention allows the quantification of expression of one or more endometrial cancer polynucleotide markers.
- Microarrays typically contain at separate sites nanomolar quantities of individual genes, cDNAs, or ESTs on a substrate (e.g. , nitrocellulose or silicon plate), or photolithographically prepared glass substrate.
- the arrays are hybridized to cDNA probes using conventional techniques with gene- specific primer mixes.
- the target polynucleotides to be analyzed are isolated, amplified and labelled, typically with fluorescent labels, radiolabels or phosphorous label probes. After hybridization is completed, the array is inserted into the scanner, where patterns of hybridization are detected. Data are collected as light emitted from the labels incorporated into the target, which becomes bound to the probe array. Probes that completely match the target generally produce stronger signals than those that have mismatches.
- the sequence and position of each probe on the array are known, and thus by complementarity, the identity of the target nucleic acid applied to the probe array can be determined.
- Microarrays are prepared by selecting polynucleotide probes and immobilizing them to a solid support or surface.
- the probes may include DNA sequences, RNA sequences, copolymer sequences of DNA and RNA, DNA and/or RNA analogues, or combinations thereof.
- the probe sequences may be full or partial fragments of genomic DNA, or they may be synthetic oligonucleotide sequences synthesized either enzymatically in vivo, enzymatically in vitro (e.g. , by PCR), or non- enzymatically in vitro.
- the probe or probes used in the methods of the invention can be immobilized to a solid support or surface which may be either porous or non-porous.
- the probes can be attached to a nitrocellulose or nylon membrane or filter covalently at either the 3' or the 5' end of the polynucleotide probe.
- the solid support may be a glass or plastic surface.
- hybridization levels are measured to microarrays of probes consisting of a solid support on the surface of which are immobilized a population of polynucleotides, such as a population of DNA or DNA mimics, or, alternatively, a population of RNA or RNA mimics.
- a solid support may be a nonporous or, optionally, a porous material such as a gel.
- a microarray including a support or surface with an ordered array of hybridization sites or "probes" each representing one of the markers described herein.
- the microarrays can be addressable arrays, and in particular positionally addressable arrays.
- Each probe of the array is typically located at a known, predetermined position on the solid support such that the identity of each probe can be determined from its position in the array.
- each probe is covalently attached to the solid support at a single site.
- Microarrays used in the present invention are preferably (a) reproducible, allowing multiple copies of a given array to be produced and easily compared with each other; (b) made from materials that are stable under hybridization conditions; (c) small (e.g. , between 1 cm2 and 25 cm2, between 12 cm2 and 13 cm2, or 3 cm2); and (d) include a unique set of binding sites that will specifically hybridize to the product of a single gene in a cell (e.g. , to a specific mRNA, miRNA, or to a specific cDNA derived therefrom).
- larger arrays may be used particularly in screening arrays, and other related or similar sequences will cross hybridize to a given binding site.
- the microarray is an array in which each position represents one of the markers described herein (e.g. , the markers listed in Table 5a, Table 5b, Table 8, and/or Table 9).
- Each position of the array can include a DNA or DNA analogue based on genomic DNA to which a particular RNA or cDNA transcribed from a genetic marker can specifically hybridize.
- a DNA or DNA analogue can be a synthetic oligomer or a gene fragment.
- probes representing each of the endometrial cancer markers and endometrial cancer polynucleotide markers are present on the array.
- the array includes at least 5 of the endometrial cancer polynucleotide markers.
- Probes for the microarray can be synthesized using N-phosphonate or phosphoramidite chemistries (Froehler et al., 1986, Nucleic Acid Res. 14:5399-5407; McBride et al. , 1983, Tetrahedron Lett. 24:246-248). Synthetic sequences are typically between about 10 and about 500 bases, 20-100 bases, or 40-70 bases in length. Synthetic nucleic acid probes can include non-natural bases, such as, without limitation, inosine. Nucleic acid analogues such as peptide nucleic acid may be used as binding sites for hybridization (see, e.g. , Egholm et al. , 1993, Nature 363:566-568; US Patent No. 5,539,083).
- Probes can be selected using an algorithm that takes into account binding energies, base composition, sequence complexity, cross-hybridization binding energies, and secondary structure (see Friend et al. , International Patent Publication WO 01 / 05935, published Jan. 25, 2001 ).
- Positive control probes e.g. , probes known to be complementary and hybridize to sequences in the target polynucleotides
- negative control probes e.g. , probes known to not be complementary and hybridize to sequences in the target polynucleotides
- Positive controls can be synthesized along the perimeter of the array or synthesized in diagonal stripes across the array.
- a reverse complement for each probe can be next to the position of the probe to serve as a negative control.
- the probes can be attached to a solid support or surface, which may be made from glass, plastic (e.g. , polypropylene, nylon), polyacrylamide, nitrocellulose, gel, or other porous or nonporous material.
- the probes can be printed on surfaces such as glass plates (see Schena et al. , 1995, Science 270:467-470). This method may be particularly useful for preparing microarrays of cDNA. (See, also, DeRisi et al. , 1996, Nature Genetics 14:457-460; Shalon et al. , 1996, Genome Res. 6:639- 645; and Schena et al. , 1995, Proc. Natl. Acad. Sci. U.S.A. 93: 10539-1 1286).
- Bioelectronics 1 1 :687-690 oligonucleotides (e.g. , 60-mers) of known sequence are synthesized directly on a surface such as a derivatized glass slide.
- the array produced may be redundant, with several oligonucleotide molecules per RNA.
- Microarrays can be made by other methods including masking (Maskos and Southern, 1992, Nuc. Acids. Res. 20: 1679-1684).
- microarrays of the present invention are produced by synthesizing polynucleotide probes on a support wherein the nucleotide probes are attached to the support covalently at either the 3' or the 5' end of the polynucleotide.
- the invention provides microarrays including a disclosed marker set.
- the invention provides a microarray for distinguishing endometrial disease samples including a positionally-addressable array of polynucleotide probes bound to a support, the polynucleotide probes including a plurality of polynucleotide probes of different nucleotide sequences, each of the different nucleotide sequences including a sequence complementary and hybridizable to a plurality of genes, the plurality consisting of at least 2, 3, 4, 5, or 6 of the genes corresponding to the markers listed in Table 5a, Table 5b, Table 8, and/or Table 9 and optionally at least 2-18, 5-16, 10-15, 13-21 , 2-21 , 2- 32, 22-32, or 13-32 of the genes corresponding to the markers listed in Table 5a, Table 5b, Table 8, and/or Table 9.
- An aspect of the invention provides microarrays including at least 4, 5, or 6 of the polynucleotides en
- the invention provides gene marker sets that distinguish endometrial disease and uses therefor.
- the invention provides a method for classifying an endometrial disease including detecting a difference in the expression of a first plurality of genes relative to a control, the first plurality of genes consisting of at least 3, 4, 5, or 6 of the genes encoding the markers listed in Table 5a, Table 5b, Table 8, and/or Table 9.
- the plurality of genes consists of at least 4 or 5 of the genes encoding the markers listed in Table 5a, Table 5b, Table 8, and/or Table 9 and optionally at least 2-18, 5-16, 10-15, 13-21 , 2-21 , 2-32, 22-32, or 13-32 of the genes
- control includes nucleic acids derived from a pool of samples from individual control patients.
- An aspect of the invention provides microarrays including at least 4, 5, or 6 of the polynucleotides encoding the markers listed in Table 5a, Table 5b, Table 8, and/or Table 9. [00239]
- the invention provides a method for classifying an endometrial disease by calculating the similarity between the expression in a sample of at least 3, 4, 5, or 6 of the endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9 and the expression of the same markers in a control pool.
- the method includes the steps of: (a) labelling nucleic acids derived from a sample, with a first fluorophore to obtain a first pool of fluorophore-labelled nucleic acids; (b) labelling with a second fluorophore a first pool of nucleic acids derived from two or more endometrial disease samples, and a second pool of nucleic acids derived from two or more control samples; (c) contacting the first fluorophore-labelled nucleic acid and the first pool of second fluorophore-labelled nucleic acid with a first microarray under conditions such that hybridization can occur, and contacting the first fluorophore-labelled nucleic acid and the second pool of second fluorophore-labelled nucleic acid with a second microarray under conditions such that hybridization can occur, detecting at each of a plurality of discrete loci on the first microarray a first fluorescent emission signal from the first fluorophore-labelled nucleic acid and a second fluorescent emission signal from
- the array can be used to monitor the time course of expression of one or more endometrial cancer polynucleotide markers in the array. This can occur in various biological contexts such as tumor progression.
- the array is also useful for ascertaining differential expression patterns of endometrial cancer polynucleotide markers, and optionally other markers, in normal and abnormal cells. This may provide a battery of nucleic acids that could serve as molecular targets for diagnosis or therapeutic intervention.
- Binding agents may be used for a variety of diagnostic and assay applications. There are a variety of assay formats known to the skilled artisan for using a binding agent to detect a target molecule in a sample. (For example, see Harlow and Lane, Antibodies: A Laboratory Manual, Cold Spring Harbor Laboratory, 1988). In general, the presence or absence of an endometrial disease (e.g. , cancer) in a subject may be determined by (a) contacting a sample from the subject with a binding agent; (b) detecting in the sample a level of polypeptide that binds to the binding agent; and (c) comparing the level of polypeptide with a predetermined standard or cut-off value.
- an endometrial disease e.g. , cancer
- the binding agent is an antibody.
- Antibodies specifically reactive with one or more endometrial disease marker, or derivatives, such as enzyme conjugates or labelled derivatives, may be used to detect one or more endometrial disease marker in various samples (e.g. , biological materials). They may be used as diagnostic or prognostic reagents and they may be used to detect abnormalities in the level of expression of one or more endometrial disease marker, or abnormalities in the structure, and/or temporal, tissue, cellular, or subcellular location of one or more endometrial disease marker. Antibodies may also be used to screen potentially therapeutic compounds in vitro to determine their effects on disorders (e.g. , cancer) involving one or more endometrial cancer markers, and other conditions. In vitro immunoassays may also be used to assess or monitor the efficacy of particular therapies.
- the invention provides a method for monitoring or diagnosing an endometrial disease (e.g. , cancer) in a subject by quantifying one or more endometrial cancer markers in a biological sample from the subject including reacting the sample with antibodies specific for one or more endometrial cancer markers, which are directly or indirectly labelled with detectable substances and detecting the detectable substances.
- endometrial cancer markers are quantified or measured.
- the invention provides a method for detecting or screening for an endometrial disease (e.g. , cancer) by: (a) obtaining a sample suspected of containing one or more protein endometrial cancer markers associated with an endometrial disease; (b) contacting the sample with antibodies that specifically bind to the protein endometrial cancer markers under conditions effective to bind the antibodies and form complexes; (c) measuring the amount of protein endometrial cancer markers present in the sample by quantifying the amount of the complexes; and (d) comparing the amount of protein endometrial cancer markers present in the samples with the amount of protein endometrial cancer markers in a control, wherein a change or significant difference in the amount of protein endometrial cancer markers in the sample compared with the amount in the control is indicative of an endometrial disease.
- an endometrial disease e.g. , cancer
- the invention contemplates a method for monitoring the progression of an endometrial disease (e.g. , cancer) in an individual, by: (a) contacting antibodies which bind to one or more protein endometrial cancer markers with a sample from the individual so as to form complexes including the antibodies and one or more endometrial cancer markers in the sample; (b) determining or detecting the presence or amount of complex formation in the sample; (c) repeating steps (a) and (b) at a point later in time; and (d) comparing the result of step (b) with the result of step (c), wherein a difference in the amount of complex formation is indicative of disease, disease stage, and/or progression of the disease in the individual.
- an endometrial disease e.g. , cancer
- the amount of complexes may also be compared to a value representative of the amount of the complexes from an individual not at risk of, or afflicted with, an endometrial disease at different stages.
- a significant difference in complex formation may be indicative of advanced disease (e.g. , advanced endometrial cancer, or an unfavourable prognosis).
- the endometrial cancer markers include one or more of those up-regulated in cancer samples as compared to normal samples, as shown in Table 5a, Table 5b, Table 8, and/or Table 9.
- examples include, without limitation, a heterogeneous nuclear ribonucleoprotein (e.g. , A2/B (hnRNPA2/B1 )), L- lactate dehydrogenase A (LDHA), anterior gradient protein2 homolog, nucleophosmin, cystatin B, hepatoma derived growth factor, and calcyphosin, and fragments thereof.
- higher levels of the up-regulated markers are indicative of the prognosis of an endometrial cancer patient's outcome.
- the endometrial cancer markers include one or more of those down- regulated in cancer samples as compared to normal samples, as shown in Table 5a, Table 5b, Table 8, and/or Table 9. Examples include, without limitation, fibrinogen (e.g. , alpha, beta, and/or gamma chain), apolipoprotein A1 , haptoglobin (HP), and serpin H1 (or heat-shock protein 47 kD (hsp47) and fragments thereof.
- fibrinogen e.g. , alpha, beta, and/or gamma chain
- apolipoprotein A1 e.g. , haptoglobin (HP)
- serpin H1 or heat-shock protein 47 kD (hsp47) and fragments thereof.
- lower levels of the down-regulated markers particularly significantly lower levels, compared to a control (normal or benign), are indicative of the prognosis of an endometrial cancer patient's outcome.
- Antibodies may be used in any known immunoassays that rely on the binding interaction between antigenic determinants of one or more endometrial disease marker and the antibodies.
- Immunoassay procedures for in vitro detection of antigens in fluid samples are also well known in the art. (See, for example, Paterson et al. , Int. J. Can. 37:659 (1986) and Burchell et al. , Int. J. Can. 34:763 (1984) for a general description of immunoassay procedures.
- Qualitative and/or quantitative determinations of one or more endometrial disease marker in a sample may be accomplished by competitive or non-competitive immunoassay procedures in either a direct or indirect format.
- Detection of one or more endometrial disease marker using antibodies can be done utilizing immunoassays which are run in either the forward, reverse or simultaneous modes.
- immunoassays are radioimmunoassays (RIA), enzyme immunoassays (e.g. , ELISA),
- an immunoassay for detecting one or more endometrial cancer markers in a biological sample includes contacting binding agents that specifically bind to endometrial cancer markers in the sample under conditions that allow the formation of first complexes including a binding agent and endometrial cancer markers and determining the presence or amount of the complexes as a measure of the amount of endometrial cancer markers contained in the sample.
- the binding agents are labelled differently or are capable of binding to different labels.
- Antibodies may be used to detect and quantify one or more endometrial cancer markers in a sample in order to diagnose and treat pathological states.
- the antibodies may be used in immunohistochemical analyses, for example, at the cellular and sub-subcellular level, to detect one or more endometrial cancer markers, to localize them to particular endometrial cells and tissues (e.g. , tumor cells and tissues), and to specific subcellular locations, and to quantify the level of expression.
- Immunohistochemical methods for the detection of antigens in tissue samples are well known in the art. For example, immunohistochemical methods are described in Taylor, Arch. Pathol. Lab. Med. 102: 1 12 (1978). Briefly, in the context of the invention, a tissue sample obtained from a subject suspected of having an endometrial-related problem is contacted with antibodies, preferably monoclonal antibodies recognizing one or more endometrial cancer markers. The site at which the antibodies are bound is determined by selective staining of the sample by standard
- tissue sample may be normal endometrial tissue, or a cancer tissue or a benign tissue.
- An antibody microarray in which binding sites include immobilized, preferably monoclonal, antibodies specific to a substantial fraction of marker-derived endometrial cancer markers of interest can be utilized in the present invention.
- Antibody arrays can be prepared using methods known in the art (see, for example, Zhu et al. , Science 293:2101 (2001 ) and reference 20).
- Antibodies specific for one or more endometrial marker may be labelled with a detectable substance and localised in biological samples based upon the presence of the detectable substance.
- detectable substances include, but are not limited to, the following: radioisotopes (e.g., 3H, 14C, 35s, 1251, 1311), fluorescent labels (e.g. , FITC, rhodamine, lanthanide phosphors), luminescent labels such as luminol; enzymatic labels (e.g.
- biotinyl groups which can be detected by marked avidin (e.g. , streptavidin containing a fluorescent marker or enzymatic activity that can be detected by optical or colorimetric methods), predetermined polypeptide epitopes recognized by a secondary reporter (e.g. , leucine zipper pair sequences, binding sites for secondary antibodies, metal binding domains, epitope tags).
- labels are attached via spacer arms of various lengths to reduce potential steric hindrance.
- Antibodies may also be coupled to electron dense substances, such as ferritin or colloidal gold, which are readily visualised by electron microscopy.
- One of the ways an antibody can be detectably labelled is to link it directly to an enzyme.
- the enzyme when later exposed to its substrate will produce a product that can be detected.
- detectable substances that are enzymes are horseradish peroxidase, beta- galactosidase, luciferase, alkaline phosphatase, acetylcholinesterase, malate dehydrogenase, ribonuclease, urease, catalase, glucose-6-phosphate, staphylococcal nuclease, delta-5-steriod isomerase, yeast alcohol dehydrogenase, alpha-glycerophosphate, those phosphate isomerase, asparaginase, glucose oxidase, and acetylcholine esterase.
- a fluorescence-emitting metal atom such as Eu (europium) and other lanthanides can be used. These can be attached to the desired molecule by means of metal-chelating groups such as DTPA or EDTA.
- a bioluminescent compound may also be used as a detectable substance.
- Bioluminescence is a type of chemiluminescence found in biological systems where a catalytic protein increases the efficiency of the chemiluminescent reaction. The presence of a bioluminescent molecule is determined by detecting the presence of luminescence. Examples of bioluminescent detectable substances are luciferin, luciferase and aequorin.
- Indirect methods may also be employed in which the primary antigen-antibody reaction is amplified by the introduction of a second antibody, having specificity for the antibody reactive against one or more endometrial cancer markers.
- a second antibody having specificity for the antibody reactive against one or more endometrial cancer markers.
- the antibody having specificity against one or more endometrial disease marker is a rabbit IgG antibody
- the second antibody may be goat anti-rabbit gamma-globulin labelled with a detectable substance as described herein.
- Cytochemical techniques known in the art for localizing antigens using light and electron microscopy may be used to detect one or more endometrial cancer markers.
- antibodies may be labelled with detectable substances and one or more endometrial cancer markers may be localised in tissues and cells based upon the presence of the detectable substances.
- binding agents e.g. , antibodies specific for one or more endometrial cancer markers
- one or more endometrial cancer markers may be immobilized on a carrier or support.
- suitable carriers or supports are agarose, cellulose, nitrocellulose, dextran, Sephadex, Sepharose, liposomes, carboxymethyl cellulose, polyacrylamides, polystyrene, gabbros, filter paper, magnetite, ion-exchange resin, plastic film, plastic tube, glass, polyamine-methyl vinyl-ether-maleic acid copolymer, amino acid copolymer, ethylene-maleic acid copolymer, nylon, silk, etc.
- the support material may have any possible configuration including spherical (e.g. , bead), cylindrical (e.g. , inside surface of a test tube or well, or the external surface of a rod), or flat (e.g. , sheet, test strip).
- the carrier may be in the shape of, for example, a tube, test plate, well, beads, disc, sphere, etc.
- the immobilized antibody may be prepared by reacting the material with a suitable insoluble carrier using known chemical or physical methods, for example, cyanogen bromide coupling.
- An antibody may be indirectly immobilized using a second antibody specific for the antibody.
- mouse antibody specific for an endometrial disease marker may be immobilized using sheep anti-mouse IgG Fc fragment specific antibody coated on the carrier or support.
- one or more endometrial disease marker may be localized by radioautography.
- the results of radioautography may be quantified by determining the density of particles in the radioautographs by various optical methods, or by counting the grains.
- Time-resolved fluorometry may be used to detect a signal.
- the method described by Christopoulos TK and Diamandis EP in Anal Chem 1992:64:342-346 may be used with a conventional time-resolved fluorometer.
- a method wherein one or more endometrial disease marker antibodies are directly or indirectly labelled with enzymes, substrates for the enzymes are added wherein the substrates are selected so that the substrates, or a reaction product of an enzyme and substrate, form fluorescent complexes with a lanthanide metal (e.g. , europium, terbium, samarium, and dysprosium, preferably europium and terbium).
- a lanthanide metal e.g. , europium, terbium, samarium, and dysprosium, preferably europium and terbium.
- a lanthanide metal is added and one or more endometrial cancer markers are quantified in the sample by measuring fluorescence of the fluorescent complexes.
- Enzymes are selected based on the ability of a substrate of the enzyme, or a reaction product of the enzyme and substrate, to complex with lanthanide metals such as europium and terbium.
- Suitable enzymes and substrates that provide fluorescent complexes are described in US Patent No. 5,31 12,922 to Diamandis.
- suitable enzymes include alkaline phosphatase and ⁇ -galactosidase.
- the enzyme is alkaline phosphatase.
- One or more endometrial disease marker antibodies may also be indirectly labelled with an enzyme.
- the antibodies may be conjugated to one partner of a ligand binding pair, and the enzyme may be coupled to the other partner of the ligand binding pair.
- Representative examples include avidin-biotin, and riboflavin-riboflavin binding protein.
- the antibodies are biotinylated, and the enzyme is coupled to streptavidin.
- an antibody specific for endometrial disease marker antibody is labelled with an enzyme.
- the invention provides means for determining one or more endometrial cancer markers in a sample by measuring one or more endometrial cancer markers by immunoassay. It will be evident to a skilled artisan that a variety of immunoassay methods can be used to measure one or more endometrial cancer markers. In general, an immunoassay method may be competitive or non-competitive. Competitive methods typically employ an immobilized or immobilizable antibody to one or more endometrial disease marker and a labelled form of one or more endometrial disease marker. Sample endometrial disease cancer markers and labelled endometrial cancer markers compete for binding to antibodies to endometrial cancer markers.
- the amount of the label in either bound or unbound fraction is measured and may be correlated with the amount of endometrial cancer markers in the test sample in any conventional manner (e.g. , by comparison to a standard curve).
- a non-competitive method is used for the determination of one or more endometrial cancer markers, with the most common method being the "sandwich” method.
- two antibodies to endometrial cancer markers are employed.
- One of the antibodies to endometrial cancer markers is directly or indirectly labelled (sometimes referred to as the “detection antibody”) and the other is immobilized or immobilizable (sometimes referred to as the "capture antibody").
- the capture and detection antibodies can be contacted simultaneously or sequentially with the test sample.
- Sequential methods can be accomplished by incubating the capture antibody with the sample, and adding the detection antibody at a predetermined time thereafter (sometimes referred to as the "forward” method); or the detection antibody can be incubated with the sample first and then the capture antibody added (sometimes referred to as the "reverse” method).
- the capture antibody is separated from the liquid test mixture, and the label is measured in at least a portion of the separated capture antibody phase or the remainder of the liquid test mixture. Generally, it is measured in the capture antibody phase since it includes endometrial disease markers bound by ("sandwiched" between) the capture and detection antibodies.
- the label may be measured without separating the capture antibodies and liquid test mixture.
- one or both of the capture and detection antibodies are polyclonal antibodies or one or both of the capture and detection antibodies are monoclonal antibodies (i.e. , polyclonal / polyclonal, monoclonal / monoclonal, or monoclonal / polyclonal).
- the label used in the detection antibody can be selected from any of those known conventionally in the art.
- the label may be an enzyme or a chemiluminescent moiety, but it can also be a radioactive isotope, a fluorophor, a detectable ligand (e.g.
- the antibody is labelled with an enzyme which is detected by adding a substrate that is selected so that a reaction product of the enzyme and substrate forms fluorescent complexes.
- the capture antibody may be selected so that it provides a means for being separated from the remainder of the test mixture. Accordingly, the capture antibody can be introduced to the assay in an already immobilized or insoluble form, or can be in an immobilizable form, that is, a form which enables immobilization to be accomplished subsequent to introduction of the capture antibody to the assay.
- An immobilized capture antibody may include an antibody covalently or non-covalently attached to a solid phase such as a magnetic particle, a latex particle, a microtiter plate well, a bead, a cuvette, or other reaction vessel.
- a solid phase such as a magnetic particle, a latex particle, a microtiter plate well, a bead, a cuvette, or other reaction vessel.
- An example of an immobilizable capture antibody is antibody which has been chemically modified with a ligand moiety, e.g. , a hapten, biotin, or the like, and which can be subsequently immobilized by contact with an immobilized form of a binding partner for the ligand, e.g. , an antibody, avidin, or the like.
- the capture antibody may be immobilized using a species specific antibody for the capture antibody that is bound to the solid phase.
- Computer Systems can be implemented by use of computer systems and methods described below and known in the art.
- the invention provides computer readable media including one or more endometrial cancer markers, and/or polynucleotides encoding one or more endometrial cancer markers, and optionally other markers (e.g. , markers of endometrial cancer).
- Computer readable media refers to any medium that can be read and accessed directly by a computer, including but not limited to magnetic storage media, such as floppy discs, hard disc storage medium, and magnetic tape; optical storage media such as CD- ROM; electrical storage media such as RAM and ROM; and hybrids of these categories such as magnetic / optical storage media.
- the invention contemplates computer readable medium having recorded thereon markers identified for patients and controls.
- Recorded refers to a process for storing information on computer readable medium.
- the skilled artisan can readily adopt any of the presently known methods for recording information on computer readable medium to generate manufactures including information on one or more endometrial cancer markers, and optionally other markers.
- a variety of data processor programs and formats can be used to store information on one or more endometrial cancer markers, and/or polynucleotides encoding one or more endometrial cancer markers, and other markers on computer readable medium.
- the information can be represented in a word processing text file, formatted in commercially-available software such as WordPerfect and Microsoft Word, or represented in the form of an ASCI I file, stored in a database application, such as SQL, DB2, Sybase, Oracle, or the like.
- Any number of dataprocessor structuring formats e.g. , text file or database
- marker information in computer readable form
- one skilled in the art can use the information in computer readable form to compare marker information obtained during or following therapy with the information stored within the data storage means.
- the invention provides a medium for holding instructions for performing a method for determining or whether a patient has an endometrial disease (e.g. , endometrial cancer) or a predisposition to an endometrial disease (e.g. , cancer), including determining the presence or absence of one or more endometrial cancer markers, and/or polynucleotides encoding one or more endometrial cancer markers, and optionally other markers, and based on the presence or absence of the one or more endometrial cancer markers, and/or polynucleotides encoding one or more endometrial cancer markers, and optionally other markers, determining endometrial receptivity, endometrial disease (e.g. , cancer) or a pre-disposition to an endometrial disease (e.g. , cancer), and optionally recommending a procedure or treatment.
- an endometrial disease e.g. , endometrial cancer
- the invention also provides in an electronic system and/or in a network, a method for determining whether a subject has an endometrial disease (e.g. , cancer) or a pre-disposition to an endometrial disease (e.g. , cancer), including determining the presence or absence of one or more endometrial cancer markers and/or polynucleotides encoding one or more endometrial cancer markers, and optionally other markers (e.g.
- an endometrial disease e.g. , cancer
- a pre-disposition to an endometrial disease e.g. , cancer
- cancer markers based on the presence or absence of the one or more endometrial cancer markers and/or polynucleotides encoding one or more endometrial cancer markers, and optionally other markers, determining whether the subject has an endometrial disease (e.g., cancer) or a pre-disposition to an endometrial disease (e.g. , cancer), and optionally recommending a procedure or treatment.
- endometrial disease e.g., cancer
- a pre-disposition to an endometrial disease e.g. , cancer
- the invention further provides in a network, a method for determining whether a subject is receptive to in vitro fertilization, has an endometrial disease (e.g. , cancer) or a pre-disposition to an endometrial disease (e.g.
- cancer including: (a) receiving phenotypic information on the subject and information on one or more endometrial cancer markers, and/or polynucleotides encoding one or more endometrial cancer markers, and optionally other markers associated with samples from the subject; (b) acquiring information from the network corresponding to the one or more endometrial cancer markers, and/or polynucleotides encoding one or more endometrial cancer markers, and optionally other markers; and (c) based on the phenotypic information and information on the one or more endometrial cancer markers, and/or polynucleotides encoding one or more endometrial cancer markers, and optionally other markers, determining whether the subject is receptive to in vitro fertilization, has an endometrial disease (e.g. , cancer) or a pre-disposition to an endometrial disease (e.g. , cancer); and (d) optionally recommending a procedure or treatment.
- endometrial disease e.
- the invention still further provides a system for identifying selected records that identify a diseased endometrial cell or tissue (e.g. , cancer cell or tissue) or an endometrial cell or tissue phase.
- a system of the invention generally includes a digital computer; a database server coupled to the computer; a database coupled to the database server having data stored therein, the data including records of data including one or more endometrial cancer markers, and/or polynucleotides encoding one or more endometrial cancer markers, and optionally other endometrial cancer markers, and a code mechanism for applying queries based upon a desired selection criteria to the data file in the database to produce reports of records which match the desired selection criteria.
- a method for detecting endometrial cancer tissue or cells using a computer having a processor, memory, display, and input / output devices the method by: (a) creating records of one or more endometrial cancer markers, and/or polynucleotides encoding one or more endometrial cancer markers, and optionally other markers of cancer identified in a sample suspected of containing endometrial cancer cells or tissue; (b) providing a database including records of data including one or more endometrial cancer markers, and/or polynucleotides encoding one or more endometrial cancer markers, and optionally other markers of cancer; and (c) using a code mechanism for applying queries based upon a desired selection criteria to the data file in the database to produce reports of records of step (a) which provide a match of the desired selection criteria of the database of step (b) the presence of a match being a positive indication that the markers of step (a) have been isolated from cells or tissue that are endometrial cancer
- the invention contemplates a business method for determining whether a subject is receptive to in vitro fertilization, has an endometrial disease (e.g. , cancer) or a pre-disposition to endometrial cancer including: (a) receiving phenotypic information on the subject and information on one or more endometrial cancer markers, and/or polynucleotides encoding the markers, and optionally other markers, associated with samples from the subject; (b) acquiring information from a network corresponding to one or more endometrial cancer markers, and/or polynucleotides encoding the markers, and optionally other markers; and (c) based on the phenotypic information, information on one or more endometrial cancer markers, and/or polynucleotides encoding the markers, and optionally other markers, and acquired information, determining whether the subject is receptive to in vitro fertilization, has an endometrial disease (e.g. , cancer) or a pre-disposition
- the computer systems, components, and methods described herein are used to monitor disease or determine the stage of disease.
- Binding agents particularly antibodies, specific for one or more endometrial disease markers may also be used in imaging methodologies in the management of an endometrial disease.
- the invention provides a method for imaging tumors associated with one or more endometrial cancer markers.
- the invention also contemplates imaging methods described herein using multiple markers for an endometrial disease.
- each agent is labelled so that it can be distinguished during the imaging.
- the method is an in vivo method and a subject or patient is administered one or more agents that carry an imaging label and that are capable of targeting or binding to one or more endometrial cancer markers.
- the agent is allowed to incubate in vivo and bind to the endometrial cancer markers associated with endometrial cells or tissues of a particular phase or associated with diseased cells or tissues, (e.g. , endometrial tumor).
- the presence of the label is localized to the endometrial cells or tissues, and the localized label is detected using imaging devices known to those skilled in the art.
- the agent may be an antibody or chemical entity that recognizes the endometrial cancer markers.
- the agent is a polyclonal antibody or monoclonal antibody, or fragments thereof, or constructs thereof including but not limited to, single chain antibodies, bifunctional antibodies, molecular recognition units, and peptides or entities that mimic peptides.
- the antibodies specific for the endometrial cancer markers used in the methods of the invention may be obtained from scientific or commercial sources, or isolated native endometrial cancer markers or recombinant endometrial cancer markers may be utilized to prepare antibodies etc. as described herein.
- An agent may be a peptide that mimics the epitope for an antibody specific for an endometrial disease marker and binds to the marker.
- the peptide may be produced on a commercial synthesizer using conventional solid phase chemistry.
- a peptide may be prepared that includes tyrosine, lysine, or phenylalanine to which N2S2 chelate is complexed (see US Patent No. 4,897,255).
- An anti-endocrine marker peptide conjugate is then combined with a radiolabel (e.g. , sodium 99mTc pertechnetate or sodium 188Re perrhenate) and it may be used to locate an endometrial disease marker producing cell or tissue (e.g.
- the agent carries a label to image the endometrial cancer markers.
- the agent may be labelled for use in radionuclide imaging.
- the agent may be directly or indirectly labelled with a radioisotope.
- radioisotopes that may be used in the invention are the following: 277Ac, 21 1 At, 128Ba, 131 Ba, 7Be, 204Bi, 205Bi, 206Bi, 76Br, 77Br, 82Br, 109Cd, 47Ca, 1 1 C, 14C, 36CI, 48Cr, 51 Cr, 62Cu, 64Cu, 67Cu, 165Dy, 155Eu, 18F, 153Gd, 66Ga, 67Ga, 68Ga, 72Ga, 198Au, 3H, 166HO, 11 1 ln, 1 13mln, 1 15mln, 1231, 1251, 131 1, 189lr, 191 m
- the radioisotope is 131 1, 1251, 1231, 1 11 1, 99mTc, 90Y, 186Re, 188Re, 32P, 153Sm, 67Ga, 201 TI 77Br, or 18F, and is imaged with a photoscanning device.
- Procedures for biological agents with the radioactive isotopes are generally known in the art.
- US Patent No. 4,302,438 describes tritium procedures. Procedures for iodinating, tritium labelling, and 35S especially adapted for murine monoclonal antibodies are described by Goding, J.vV. (supra, pp. 124-126) and the references cited therein. Other procedures for iodinating biological agents, such as antibodies, binding portions thereof, probes, or ligands, are described in the scientific literature (see Hunter and Greenwood, Nature 144:945 (1962); David et al. , Biochemistry 13: 1014- 1021 (1974); and US Patent Nos. 3,867,517 and 4,376, 110).
- An agent may also be labelled with a paramagnetic isotope for purposes of an in vivo method of the invention.
- elements that are useful in magnetic resonance imaging include gadolinium, terbium, tin, iron, or isotopes thereof. See, for example, Schaefer et al. , (1989) JACC 14, 472-480; Shreve et al. , ( 1986) Magn. Reson. Med. 3, 336-340; Wolf, G L , (1984) Physiol. Chem. Phys. Med. N MR 16, 93-95; Wesbey et al. ( 1984), Physiol. Chem. Phys. Med. N MR 16, 145- 155; Runge et al. (1984), Invest. Radiol. 19, 408-415 for discussions on in vivo nuclear magnetic resonance imaging.
- the agent may be administered to the patient, it is localized to the cell or tissue (e.g. , tumor) having an endometrial disease marker with which the agent binds, and is detected or "imaged" in vivo using known techniques such as radionuclear scanning using e.g. , a gamma camera or emission tomography.
- known techniques such as radionuclear scanning using e.g. , a gamma camera or emission tomography.
- a positron emission transaxial tomography scanner such as designated Pet VI located at Brookhaven National Laboratory, can also be used where the radiolabel emits positrons (e.g. , 1 1 C, 18F, 150, and 13N).
- Whole body imaging techniques using radioisotope labelled agents can be used for locating diseased cells and tissues (e.g. , primary tumors and tumors which have metastasized).
- Antibodies specific for endometrial cancer markers, or fragments thereof having the same epitope specificity are bound to a suitable radioisotope, or a combination thereof, and administered parenterally.
- administration preferably is intravenous.
- the bio-distribution of the label can be monitored by scintigraphy, and accumulations of the label are related to the presence of endometrial cancer cells.
- Whole body imaging techniques are described in US Patent Nos.
- agents useful for diagnosis and therapeutic use that can be coupled to antibodies and antibody fragments include metallothionein and fragments (see US Patent No. 4,732,864). These agents are useful in diagnosis staging and visualization of cancer, in particular endometrial cancer, so that surgical and/or radiation treatment protocols can be used more efficiently.
- An imaging agent may carry a bioluminescent or chemiluminescent label.
- Such labels include polypeptides known to be fluorescent, bioluminescent, or chemiluminescent, or, that act as enzymes on a specific substrate (reagent), or can generate a fluorescent, bioluminescent or chemiluminescent molecule.
- bioluminescent or chemiluminescent labels include luciferases, aequorin, obelin, mnemiopsin, berovin, a phenanthridinium ester, and variations thereof and combinations thereof.
- a substrate for the bioluminescent or chemiluminescent polypeptide may also be utilized in a method of the invention.
- the chemiluminescent polypeptide can be luciferase and the reagent luciferin.
- a substrate for a bioluminescent or chemiluminescent label can be administered before, at the same time (e.g. , in the same formulation), or after administration of the agent.
- An imaging agent may include a paramagnetic compound, such as a polypeptide chelated to a metal (e.g. , a metalloporphyrin).
- the paramagnetic compound may also include a monocrystalline nanoparticle, e.g. , a nanoparticle including a lanthanide (e.g. , Gd) or iron oxide; or, a metal ion such as a lanthanide.
- lanthanide refers to elements of atomic numbers 58 to 70, a transition metal of atomic numbers 21 to 29, 42 or 44, a Gd(l l l), a Mn(l l), or an element including a Fe element.
- Paramagnetic compounds can also include a neodymium iron oxide (NdFe03) or a dysprosium iron oxide (DyFe03).
- elements that are useful in magnetic resonance imaging include gadolinium, terbium, tin, iron, or isotopes thereof.
- Schaefer et al. (1989) JACC 14, 472-480; Shreve et al. (1986), Magn. Reson. Med. 3, 336-340; Wolf, G. L. (1984), Physiol. Chem. Phys. Med. NMR 16, 93-95; Wesbey et al. (1984), Physiol. Chem. Phys. Med. NMR 16, 145-155; Runge et al. (1984), Invest. Radiol. 19, 408-415 for discussions on in vivo nuclear magnetic resonance imaging. )
- An image can be generated in a method of the invention by computer assisted tomography (CAT), magnetic resonance spectroscopy (MRS) image, magnetic resonance imaging (MRI), positron emission tomography (PET), single-photon emission computed tomography (SPECT), or bioluminescence imaging (BLI) or equivalent.
- CAT computer assisted tomography
- MRS magnetic resonance spectroscopy
- MRI magnetic resonance imaging
- PET positron emission tomography
- SPECT single-photon emission computed tomography
- BBI bioluminescence imaging
- Computer assisted tomography (CAT) and computerized axial tomography (CAT) systems and devices well known in the art can be utilized in the practice of the invention.
- CAT Computer assisted tomography
- CAT computerized axial tomography
- the invention may also utilize animal imaging modalities, such as MicroCATTM (ImTek, Inc. ).
- Magnetic resonance imaging (MRI) systems and devices well known in the art can be utilized in the practice of the invention.
- MRI methods and devices see, for example, US Patent Nos. 6, 151 ,377; 6, 144,202; 6, 128,522; 6, 127,825; 6, 121 ,775; 6, 1 19,032;
- MRI and supporting devices are commercially available, for example, from Bruker Medical GMBH; Caprius; Esaote Biomedica; Fonar; GE Medical Systems (GEMS); Hitachi Medical Systems America; Intermagnetics General Corporation; Lunar Corp.; MagneVu; Marconi Medicals; Philips Medical Systems; Shimadzu; Siemens; Toshiba America Medical Systems; including imaging systems, by, e.g. , Silicon Graphics.
- the invention may also utilize animal imaging modalities such as micro-MRIs.
- PET Positron emission tomography imaging
- a method of the invention may use the system designated Pet VI located at Brookhaven National Laboratory.
- Pet VI located at Brookhaven National Laboratory.
- Animal imaging modalities such as micro-PETs (Concorde Microsystems, Inc. ) can also be used in the invention.
- Single-photon emission computed tomography (SPECT) systems and devices well known in the art can be utilized in the practice of the invention. See, for example, US Patent Nos. 6, 115,446; 6,072, 177; 5,608,221 ; 5,600, 145; 5,210,421 ; 5, 103,098.
- the methods of the invention may also utilize animal imaging modalities, such as micro-SPECTs.
- Bioluminescence imaging includes bioluminescence, fluorescence, and
- chemiluminescence and other photon detection systems and devices that are capable of detecting bioluminescence, fluorescence, or chemiluminescence.
- Sensitive photon detection systems can be used to detect bioluminescent and fluorescent proteins externally; see, for example, Contag (2000), Neoplasia 2:41 -52; and Zhang (1994), Clin. Exp. Metastasis, 12:87-92.
- the methods of the invention can be practiced using any such photon detection device, or variation or equivalent thereof, or in conjunction with any known photon detection methodology, including visual imaging.
- an intensified charge-coupled device (ICCD) camera coupled to an image processor may be used in the invention.
- ICCD intensified charge-coupled device
- Photon detection devices are also commercially available from Xenogen, Hamamatsue.
- test agents and compounds include but are not limited to peptides such as soluble peptides including Ig-tailed fusion peptides, members of random peptide libraries and combinatorial chemistry-derived molecular libraries made of D- and/or L-configuration amino acids, phosphopeptides (including members of random or partially degenerate, directed phosphopeptide libraries), antibodies (e.g.
- agents or compounds may be endogenous physiological compounds or natural or synthetic compounds.
- the invention provides a method for assessing the potential efficacy of a test agent or therapy for inhibiting an endometrial disease (e.g. , cancer) in a patient, by comparing: (a) levels of one or more protein endometrial cancer markers, and/or polynucleotides encoding endometrial cancer markers, and optionally other markers in a first sample obtained from a patient and exposed to the test agent or therapy; and (b) levels of one or more protein endometrial cancer markers, and/or polynucleotides encoding endometrial cancer markers, and optionally other markers, in a second sample obtained from the patient, wherein the sample is not exposed to the test agent or therapy, wherein a significant difference in the levels of expression of one or more protein endometrial cancer markers, and/or polynucleotides encoding one or more endometrial cancer markers, and optionally the other markers, in the first sample, relative to the second sample, is an indication that the test agent or therapy is potentially eff
- the first and second samples may be portions of a single sample obtained from a patient or portions of pooled samples obtained from a patient.
- the invention provides a method of selecting an agent for inhibiting an endometrial disease (e.g. , cancer) in a patient by: (a) obtaining a sample from the patient;
- an endometrial disease e.g. , cancer
- the invention provides a method of selecting an agent for inhibiting or enhancing endometrial cell or tissue phase in a patient by: (a) obtaining a sample of endometrial cell or tissue in a selected phase; (b) separately maintaining aliquots of the sample in the presence of a plurality of test agents; (c) comparing one or more endometrial cancer markers, and/or
- polynucleotides encoding endometrial cancer markers, and optionally other markers, in each of the aliquots are selected by selecting one of the test agents which alters the levels of one or more endometrial cancer markers, and/or polynucleotides encoding endometrial cancer markers, and optionally other markers in the aliquot containing that test agent, relative to other test agents.
- Still another aspect of the invention provides a method of conducting a drug discovery business by: (a) providing one or more methods or assay systems for identifying agents that inhibit an endometrial disease (e.g. , cancer) or affect an endometrial cell or tissues phase in a patient; (b) conducting therapeutic profiling of agents identified in step (a), or further analogs thereof, for efficacy and toxicity in animals; and (c) formulating a pharmaceutical preparation including one or more agents identified in step (b) as having an acceptable therapeutic profile.
- an endometrial disease e.g. , cancer
- the subject method can also include a step of establishing a distribution system for distributing the pharmaceutical preparation for sale, and may optionally include establishing a sales group for marketing the pharmaceutical preparation.
- the invention also contemplates a method of assessing the potential of a test compound to contribute to an endometrial disease (e.g. , cancer) by: (a) maintaining separate aliquots of cells or tissues from a patient with an endometrial disease (e.g. , cancer) in the presence and absence of the test compound; and (b) comparing one or more endometrial cancer markers and/or polynucleotides encoding the endometrial cancer markers, and optionally other markers in each of the aliquots.
- an endometrial disease e.g. , cancer
- the endometrial cancer markers are protein endometrial cancer markers.
- Kits The invention also contemplates kits for carrying out the methods of the invention. Kits may typically include two or more components required for performing a diagnostic assay.
- Components include but are not limited to compounds, reagents, containers, and/or equipment.
- kits including one or more specific endometrial disease marker polynucleotide, endometrial cancer markers, or antibody described herein, which may be conveniently used, e.g. , in clinical settings to screen and diagnose patients and to screen and identify those individuals exhibiting a predisposition to developing an endometrial disease.
- a container with a kit includes a binding agent as described herein.
- the kit may contain antibodies or antibody fragments which bind specifically to epitopes of one or more endometrial cancer markers and optionally other markers, antibodies against the antibodies labelled with an enzyme; and a substrate for the enzyme.
- the kit may also contain microtiter plate wells, standards, assay diluent, wash buffer, adhesive plate covers, and/or instructions for carrying out a method of the invention using the kit.
- the kit includes antibodies or fragments of antibodies which bind specifically to an epitope of one or more polypeptide listed in Table 5a, Table 5b, Table 8, and/or Table 9 that is up-regulated in cancer samples as compared to normal samples, and means for detecting binding of the antibodies to their epitope associated with tumor cells, either as concentrates (including lyophilized compositions), which may be further diluted prior to use or at the concentration of use, where the vials may include one or more dosages.
- single dosages may be provided in sterilized containers, having the desired amount and concentration of agents. Containers that provide a formulation for direct use, usually do not require other reagents, as for example, where the kit contains a radiolabeled antibody preparation for in vivo imaging.
- a kit may be designed to detect the level of protein EmCa markers or polynucleotides encoding protein endometrial cancer markers in a sample.
- the kit includes one or more polynucleotides encoding one or more polypeptides listed in Table 5a, Table 5b, Table 8, and/or Table 9 that are up-regulated in EmCa samples as compared to normal samples.
- Such kits generally include at least one oligonucleotide probe or primer, as described herein, that hybridizes to a polynucleotide encoding one or more endometrial cancer markers.
- Such an oligonucleotide may be used, for example, within a PCR or hybridization procedure.
- Additional components that may be present within the kit include a second oligonucleotide and/or a diagnostic reagent or container to facilitate detection of a polynucleotide encoding one or more protein endometrial cancer markers.
- the invention provides a kit containing a microarray described herein ready for hybridization to target polynucleotide endometrial cancer markers, plus software for the data analysis of the results.
- the software to be included with the kit includes data analysis methods, in particular mathematical routines for marker discovery, including the calculation of correlation coefficients between clinical categories and marker expression.
- the software may also include mathematical routines for calculating the correlation between sample marker expression and control marker expression, using array-generated fluorescence data, to determine the clinical classification of the sample.
- kits suitable for applying the screening methods of the invention to evaluate compounds may be packaged into convenient kits described herein providing the necessary materials packaged into suitable containers.
- the invention contemplates a kit for assessing the presence of endometrial disease cells, wherein the kit includes antibodies specific for one or more protein endometrial cancer markers, or primers or probes for polynucleotides encoding the endometrial cancer markers and optionally probes, primers or antibodies specific for other markers associated with an endometrial disease (e.g. , cancer).
- the invention relates to a kit for assessing the suitability of each of a plurality of test compounds for inhibiting an endometrial disease (e.g. , cancer) in a patient.
- the kit includes reagents for assessing one or more or protein endometrial cancer markers, polynucleotides encoding the protein endometrial cancer markers, and optionally a plurality of test agents or compounds.
- the invention provides a kit for assessing the potential of a test compound to contribute to an endometrial disease (e.g. , cancer).
- the kit includes endometrial diseased cells (e.g. , cancer cells) and reagents for assessing one or more endometrial cancer markers, polynucleotide encoding the endometrial cancer markers and optionally other markers associated with an endometrial disease.
- One or more endometrial cancer markers may be targets for immunotherapy.
- Immunotherapeutic methods include the use of antibody therapy, in vivo vaccines, and ex vivo immunotherapy approaches.
- the invention provides one or more endometrial disease marker antibodies that may be used systemically to treat an endometrial disease associated with the marker.
- the endometrial disease is endometrial cancer and one or more endometrial disease marker antibodies may be used systemically to treat endometrial cancer.
- antibodies are used that target the tumor cells but not the surrounding non-tumor cells and tissue.
- the endometrial cancer is EmCa.
- the endometrial cancer is Type I EmCa.
- the invention provides a method of treating a patient susceptible to, or having a disease (e.g. , cancer) that expresses one or more endometrial disease marker, in particular, a marker up-regulated in endometrial cancer (for example, an up-regulated marker in Table 5a, Table 5b, Table 8, and/or Table 9), including administering to the patient an effective amount of an antibody that binds specifically to one or more endometrial disease marker.
- a disease e.g. , cancer
- a marker up-regulated in endometrial cancer for example, an up-regulated marker in Table 5a, Table 5b, Table 8, and/or Table 9
- the invention provides a method of inhibiting the growth of tumor cells expressing one or more endometrial cancer markers, including administering to a patient an antibody which binds specifically to one or more endometrial cancer markers in an amount effective to inhibit growth of the tumor cells.
- One or more endometrial disease marker antibodies may also be used in a method for selectively inhibiting the growth of, or killing a cell expressing one or more endometrial disease marker (e.g. , tumor cell expressing one or more endometrial cancer marker) including reacting one or more endometrial disease marker antibody immunoconjugate or immunotoxin with the cell in an amount sufficient to inhibit the growth of, or kill the cell.
- endometrial disease marker e.g. , tumor cell expressing one or more endometrial cancer marker
- unconjugated antibodies to endometrial cancer markers may be introduced into a patient such that the antibodies bind to endometrial cancer marker expressing cancer cells and mediate growth inhibition of such cells (including the destruction thereof), and the tumor, by mechanisms which may include complement-mediated cytolysis, antibody-dependent cellular cytotoxicity, altering the physiologic function of one or more endometrial cancer markers, and/or the inhibition of ligand binding or signal transduction pathways.
- one or more endometrial cancer marker antibodies conjugated to therapeutic agents e.g.
- immunoconjugates may also be used therapeutically to deliver the agent directly to one or more endometrial cancer marker expressing tumor cells and thereby destroy the tumor.
- agents include abrin, ricin A, Pseudomonas exotoxin, or diphtheria toxin; proteins such as tumor necrosis factor, alpha-interferon, beta-interferon, nerve growth factor, platelet derived growth factor, tissue plasminogen activator; and biological response modifiers such as lymphokines, interleukin-1 , interleukin-2, interleukin-6, granulocyte macrophage colony stimulating factor, granulocyte colony stimulating factor, or other growth factors.
- Cancer immunotherapy using one or more endometrial cancer marker antibodies may utilize the various approaches that have been successfully employed for cancers, including but not limited to colon cancer (Arlen et al. , 1998, Crit Rev Immunol 18: 133-138), multiple myeloma (Ozaki et al. , 1997, Blood 90: 3179-3186; Tsunenati et al. , 1997, Blood 90: 2437-2444), gastric cancer (Kasprzyk et al. , 1992, Cancer Res 52: 2771 -2776), B-cell lymphoma (Funakoshi et al. , 1996, J.
- endometrial cancer marker antibodies capable of inhibiting the growth of precancer or cancer cells expressing endometrial cancer markers are administered in a therapeutically effective amount to cancer patients whose lesions or tumors express or overexpress one or more endometrial cancer markers.
- the invention may provide a specific, effective and long-needed treatment for endometrial cancer.
- the antibody therapy methods of the invention may be combined with other therapies including chemotherapy and radiation.
- Patients may be evaluated for the presence and level of expression or overexpression of one or more endometrial cancer markers in diseased cells and tissues (e.g. , tumors), in particular using immunohistochemical assessments of tissue, quantitative imaging as described herein, or other techniques capable of reliably indicating the presence and degree of expression of one or more endometrial disease markers. Immunohistochemical analysis of tumor biopsies or surgical specimens may be employed for this purpose.
- Endometrial disease marker antibodies useful in treating disease include those that are capable of initiating a potent immune response against the disease (e.g., tumor) and those that are capable of direct cytotoxicity.
- endometrial disease marker antibodies may elicit cell lysis by either complement-mediated or antibody-dependent cell cytotoxicity (ADCC) mechanisms, both of which require an intact Fc portion of the immunoglobulin molecule for interaction with effector cell Fc receptor sites or complement proteins.
- ADCC antibody-dependent cell cytotoxicity
- Endometrial disease marker antibodies that exert a direct biological effect on tumor growth may also be useful in the practice of the invention. Such antibodies may not require the complete immunoglobulin to exert the effect.
- Potential mechanisms by which such directly cytotoxic antibodies may act include inhibition of cell growth, modulation of cellular differentiation, modulation of tumor angiogenesis factor profiles, and the induction of apoptosis.
- the mechanism by which a particular antibody exerts an anti-tumor effect may be evaluated using any number of in vitro assays designed to determine ADCC, antibody-dependent macrophage-mediated cytotoxicity (ADMMC), complement-mediated cell lysis, and others known in the art.
- ADMMC antibody-dependent macrophage-mediated cytotoxicity
- the anti-tumor activity of a particular endometrial cancer marker antibody, or combination of endometrial cancer marker antibodies may be evaluated in vivo using a suitable animal model.
- Xenogenic cancer models where human cancer explants or passaged xenograft tissues are introduced into immune compromised animals, such as nude or SCID mice, may be employed.
- the methods of the invention contemplate the administration of single endometrial disease marker antibodies as well as combinations, or "cocktails", of different individual antibodies such as those recognizing different epitopes of other markers.
- Such cocktails may have certain advantages in as much as they contain antibodies that bind to different epitopes of endometrial cancer markers and/or exploit different effector mechanisms or combine directly cytotoxic antibodies with antibodies that rely on immune effector functionality. Such antibodies in combination may exhibit synergistic therapeutic effects.
- the administration of one or more endometrial disease marker specific antibodies may be combined with other therapeutic agents, including but not limited to chemotherapeutic agents, androgen-blockers, and immune modulators (e.g. , IL2, GM-CSF).
- the endometrial disease marker specific antibodies may be administered in their "naked” or unconjugated form, or may have therapeutic agents conjugated to them.
- the endometrial disease marker specific antibodies used in the methods of the invention may be formulated into pharmaceutical compositions including a carrier suitable for the desired delivery method.
- Suitable carriers include any material which when combined with the antibodies retains the function of the antibody and is non-reactive with the subject's immune systems. Examples include any of a number of standard pharmaceutical carriers such as sterile phosphate buffered saline solutions, bacteriostatic water, and the like (see, generally, Remington's Pharmaceutical Sciences 16th Edition, A. Osal. , ed. , 1980).
- One or more endometrial disease marker specific antibody formulations may be administered via any route capable of delivering the antibodies to a disease (e.g. , tumor) site.
- Routes of administration include, but are not limited to, intravenous, intraperitoneal, intramuscular, intratumor, intradermal, and the like.
- the route of administration is by intravenous injection.
- Antibody preparations may be lyophilized and stored as a sterile powder, preferably under vacuum, and then reconstituted in bacteriostatic water containing, for example, benzyl alcohol preservative, or in sterile water prior to injection.
- Treatment will generally involve the repeated administration of the antibody preparation via an acceptable route of administration such as intravenous injection (IV), at an effective dose.
- IV intravenous injection
- Dosages will depend upon various factors generally appreciated by those of skill in the art, including the type of disease and the severity, grade, or stage of the disease, the binding affinity and half life of the antibodies used, the degree of endometrial disease marker expression in the patient, the extent of circulating endometrial disease markers, the desired steady-state antibody concentration level, frequency of treatment, and the influence of any chemotherapeutic agents used in combination with the treatment method of the invention.
- Daily doses may range from about 0.1 to 100 mg / kg.
- Doses in the range of 10-500 mg antibodies per week may be effective and well tolerated, although even higher weekly doses may be appropriate and/or well tolerated.
- a determining factor in defining the appropriate dose is the amount of a particular antibody necessary to be therapeutically effective in a particular context. Repeated administrations may be required to achieve disease inhibition or regression. Direct administration of one or more endometrial disease marker antibodies is also possible and may have advantages in certain situations.
- Patients may be evaluated for serum cancer markers in order to assist in the determination of the most effective dosing regimen and related factors.
- the endometrial cancer assay methods described herein, or similar assays may be used for quantifying circulating endometrial disease marker levels in patients prior to treatment. Such assays may also be used for monitoring throughout therapy, and may be useful to gauge therapeutic success in combination with evaluating other parameters such as serum levels of endometrial cancer markers.
- the invention further provides vaccines formulated to contain one or more endometrial disease marker or fragment thereof.
- the invention provides a method of vaccinating an individual against one or more endometrial disease marker listed in Table 5a, Table 5b, Table 8, and/or Table 9, including the step of inoculating the individual with the marker or fragment thereof that lacks activity, wherein the inoculation elicits an immune response in the individual thereby vaccinating the individual against the marker.
- viral gene delivery systems may be used to deliver one or more polynucleotide endometrial cancer markers or complementary anti-sense versions thereof.
- Various viral gene delivery systems which can be used in the practice of this aspect of the invention include, but are not limited to, vaccinia, fowlpox, canarypox, adenovirus, influenza, poliovirus, adeno- associated virus, lentivirus, and Sindbus virus (Restifo, 1996, Curr. Opin. Immunol. 8: 658-663).
- Non- viral delivery systems may also be employed by using naked DNA encoding one or more endometrial cancer marker or fragment thereof introduced into the patient (e.g. , intramuscularly) to induce an antitumor response.
- Various ex vivo strategies may also be employed.
- One approach involves the use of cells to present one or more endometrial disease marker to a patient's immune system.
- autologous dendritic cells which express MHC class I and I I, may be pulsed with one or more endometrial disease marker or peptides thereof that are capable of binding to MHC molecules, to thereby stimulate the patients' immune systems (see, for example, Tjoa et al. , 1996, Prostate 28: 65- 69; Murphy et al. , 1996, Prostate 29: 371-380).
- Anti-idiotypic endometrial disease marker specific antibodies can also be used in therapy as a vaccine for inducing an immune response to cells expressing one or more endometrial disease marker.
- the generation of anti-idiotypic antibodies is well known in the art and can readily be adapted to generate anti-idiotypic endometrial cancer marker specific antibodies that mimic an epitope on one or more endometrial cancer markers (see, for example, Wagner et al. , 1997, Hybridoma 16: 33-40; Foon et al., 1995, J. Clin Invest 96: 334-342; and Herlyn et al. , 1996, Cancer Immunol Immunother 43: 65-76).
- Such an antibody can be used in anti-idiotypic therapy as presently practiced with other anti-idiotypic antibodies directed against antigens associated with disease (e.g. , tumor antigens).
- Genetic immunization methods may be utilized to generate prophylactic or therapeutic humoral and cellular immune responses directed against cells expressing one or more endometrial cancer marker.
- One or more DNA molecules encoding endometrial cancer markers, constructs including DNA encoding one or more endometrial cancer markers / immunogens and appropriate regulatory sequences may be injected directly into muscle or skin of an individual, such that the cells of the muscle or skin take-up the construct and express the encoded endometrial cancer markers / immunogens.
- the endometrial cancer markers / immunogens may be expressed as cell surface proteins or be secreted. Expression of one or more endometrial cancer markers results in the generation of prophylactic or therapeutic humoral and cellular immunity against the disease (e.g. , cancer).
- Various prophylactic and therapeutic genetic immunization techniques known in the art may be used.
- the invention further provides methods for inhibiting cellular activity (e.g. , cell proliferation, activation, or propagation) of a cell expressing one or more endometrial disease marker.
- This method includes reacting immunoconjugates of the invention (e.g. , a heterogeneous or homogenous mixture) with the cell so that endometrial cancer markers form complexes with the immunoconjugates.
- immunoconjugates of the invention e.g. , a heterogeneous or homogenous mixture
- a subject with a neoplastic or preneoplastic condition can be treated when the inhibition of cellular activity results in cell death.
- the invention provides methods for selectively inhibiting a cell expressing one or more endometrial disease marker by reacting any one or a combination of the immunoconjugates of the invention with the cell in an amount sufficient to inhibit the cell.
- Amounts include those that are sufficient to kill the cell or sufficient to inhibit cell growth or proliferation.
- Vectors derived from retroviruses, adenovirus, herpes or vaccinia viruses, or from various bacterial plasmids may be used to deliver polynucleotides encoding endometrial cancer markers or to a targeted organ, tissue, or cell population. Methods well known to those skilled in the art may be used to construct recombinant vectors that will express antisense polynucleotides for endometrial cancer markers. (See, for example, the techniques described in Sambrook et al. (supra) and Ausubel et al. (supra). )
- Methods for introducing vectors into cells or tissues include those methods discussed herein and which are suitable for in vivo, in vitro and ex vivo therapy.
- vectors may be introduced into stem cells obtained from a patient and clonally propagated for autologous transplant into the same patient (See US Patent Nos. 5,399,493 and 5,437,994). Delivery by transfection and by liposome is well known in the art.
- Genes encoding endometrial cancer markers can be turned off by transfecting a cell or tissue with vectors that express high levels of a desired endometrial disease marker-encoding fragment. Such constructs can inundate cells with untranslatable sense or antisense sequences. Even in the absence of integration into the DNA, such vectors may continue to transcribe RNA molecules until all copies are disabled by endogenous nucleases.
- Modifications of gene expression can be obtained by designing antisense molecules, DNA, RNA or PNA, to the regulatory regions of a gene encoding an endometrial disease marker, i.e. , the promoters, enhancers, and introns.
- oligonucleotides are derived from the transcription initiation site, (e.g. , between -10 and +10 regions of the leader sequence).
- the antisense molecules may also be designed so that they block translation of mRNA by preventing the transcript from binding to ribosomes. Inhibition may also be achieved using "triple helix" base-pairing methodology.
- Triple helix pairing compromises the ability of the double helix to open sufficiently for the binding of polymerases, transcription factors, or regulatory molecules.
- Therapeutic advances using triplex DNA were reviewed by Gee J E et al. , in: Huber and Carr (1994), Molecular and Immunologic Approaches, Futura Publishing Co, Mt Kisco, N.Y.
- Ribozymes are enzymatic RNA molecules that catalyze the specific cleavage of RNA. Ribozymes act by sequence-specific hybridization of the ribozyme molecule to complementary target RNA, followed by endonucleolytic cleavage.
- the invention therefore contemplates engineered hammerhead motif ribozyme molecules that can specifically and efficiently catalyze endonucleolytic cleavage of sequences encoding an endometrial disease marker.
- Specific ribozyme cleavage sites within any potential RNA target may initially be identified by scanning the target molecule for ribozyme cleavage sites which include the following sequences, GUA, GUU, and GUC. Once the sites are identified, short RNA sequences of between 15 and 20 ribonucleotides corresponding to the region of the target gene containing the cleavage site may be evaluated for secondary structural features which may render the oligonucleotide inoperable. The suitability of candidate targets may also be determined by testing accessibility to hybridization with complementary oligonucleotides using ribonuclease protection assays.
- One or more endometrial cancer markers and polynucleotides encoding the markers, and fragments thereof, may be used in the treatment of an endometrial disease (e.g. , cancer) in a subject.
- the endometrial cancer markers and polynucleotides encoding the markers are endometrial cancer markers that are down- regulated in endometrial cancer (e.g. , apolipoprotein A1 , serpin H, or one or more of the down- regulated markers listed in Table 5a, Table 5b, Table 8, and/or Table 9).
- the markers or polynucleotides may be formulated into compositions for administration to subjects suffering from an endometrial disease.
- the invention also relates to a composition including one or more endometrial disease cancer markers or polynucleotides encoding the markers, or a fragment thereof, and a pharmaceutically acceptable carrier, excipient or diluent.
- a method for treating or preventing an endometrial disease in a subject is also provided including administering to a patient in need thereof, one or more endometrial cancer markers or polynucleotides encoding the markers, or a composition of the invention.
- the invention further provides a method of inhibiting an endometrial disease (e.g. , cancer) in a patient by: (a) obtaining a sample including diseased cells from the patient; (b) separately maintaining aliquots of the sample in the presence of a plurality of test agents; (c) comparing levels of one or more endometrial cancer markers, and/or polynucleotides encoding one or more endometrial cancer markers in each aliquot; and (d) administering to the patient at least one of the test agents which alters the levels of the endometrial cancer markers, markers, and/or polynucleotides encoding one or more cancer markers in the aliquot containing that test agent, relative to the other test agents.
- an endometrial disease e.g. , cancer
- An active therapeutic substance described herein may be administered in a convenient manner such as by injection (subcutaneous, intravenous, etc. ), oral administration, inhalation, transdermal application, or rectal administration. Depending on the route of administration, the active substance may be coated in a material to protect the substance from the action of enzymes, acids and other natural conditions that may inactivate the substance.
- Solutions of an active compound as a free base or pharmaceutically acceptable salt can be prepared in an appropriate solvent with a suitable surfactant. Dispersions may be prepared in glycerol, liquid polyethylene glycols, and mixtures thereof, or in oils.
- compositions described herein can be prepared by per se known methods for the preparation of pharmaceutically acceptable compositions which can be administered to subjects, such that an effective quantity of the active substance is combined in a mixture with a pharmaceutically acceptable vehicle.
- Suitable vehicles are described, for example, in Remington's Pharmaceutical Sciences (Remington's Pharmaceutical Sciences, Mack Publishing Company, Easton, Pa. , USA, 1985).
- the compositions include, albeit not exclusively, the active substances in association with one or more pharmaceutically acceptable vehicles or diluents, and contained in buffered solutions with a suitable pH and iso-osmotic with the physiological fluids.
- compositions are indicated as therapeutic agents either alone or in conjunction with other therapeutic agents or other forms of treatment.
- the compositions of the invention may be administered concurrently, separately, or sequentially with other therapeutic agents or therapies.
- the therapeutic activity of compositions, agents, and compounds may be identified using a method of the invention and may be evaluated in vivo using a suitable animal model.
- the inventors' study is a significant advancement in this direction as it places major emphasis on determining the clinical impact of a proteomics based biomarker in diagnosing tumor in patients showing symptoms of endometrial disease.
- Examples 1 -7 the inventors demonstrate the identification of a panel of proteins with consistently differential expression, including apolipoprotein A1 , calcyphosin, cathepsin B, dihydropyrimidinase-like 2, fibrinogen (e.g. , alpha, beta, and gamma chains), haptoglobin, macrophage capping protein (CAP-G), and hsp47, that serve as endometrial cancer biomarkers.
- Examples 8-13 the inventors demonstrate the identification of a panel of proteins with consistently differential expression among different subtypes of EmCa as compared to normal tissues, including a heterogeneous nuclear ribonucleoprotein (e.g.
- A2/B hnRNPA2/B1
- nucleophosmin LDHA
- HDGF hepatoma-derived growth factor
- mitochondrial peroxiredoxin-5 mitochondrial peroxiredoxin-5
- cystatin B cystatin B.
- the inventors demonstrate the identification of a panel of proteins with consistently differential expression across all subtypes of EmCa as compared to normal tissues, including alpha-enolase, glucose-6-phosphate isomerase, phosphoglycerate kinase, villin 2 (ezrin), mitochondrial isocitrate dehyrdogenase 2 (NADP+), transketolase, serpin H1 (hsp47), and nucleolin.
- Endometrial tissues were retrieved from an in-house, dedicated research endometrial tissue bank, as previously described (12). The collection and use of these materials were approved by the Research Ethics Boards of York University, Mount Sinai Hospital, University Health Network, and North York General Hospital. Briefly, samples from hysterectomy specimens were flash frozen in liquid nitrogen within 20 min of devitalizing. A portion of the tissue sample from the mirror face of the histological section was taken for proteomic analysis.
- This material was washed three times in ⁇ 1 mL of PBS with a mixture of protease inhibitors, as described previously (1 mM 4-(2-aminoethyl) benzenesulfonyl fluoride, 10 ⁇ ⁇ leupeptin, 1 ⁇ g/mL aprotinin, and 1 ⁇ pepstatin) (14)). It was then homogenized in 0.5 mL of phosphate-buffered saline (PBS) with protease inhibitors using a hand-held homogenizer, and then flash-frozen in liquid nitrogen and stored at -80°C until use.
- PBS phosphate-buffered saline
- iTRAQ For each sample, 100 ⁇ g of proteins were digested and labelled individually with the appropriate iTRAQ tag (vide infra). Trypsin digestion and labeling were performed according to the manufacturer's protocol, except that the trypsin used was porcine sequencing grade (Promega, Madison, VVI). A total of seven iTRAQ sets were examined (Table 1 ). Each iTRAQ set included the reference sample, one individual normal proliferative sample, and two EmCa samples (one new and the other a previously examined sample), with the exception of the last two sets which included the remaining normal proliferative samples and an EmCa sample. The assignments of the tags to the sample types were randomized to minimize the effects of any m/z- related biases.
- Each iTRAQ set was then separated by SCX fractionation using an HP1050 HPLC instrument (Agilent, Palo Alto, CA) with a 2.1-mm internal diameter x 100-mm-length PolyLC Polysulfoethyl A column packed with 5- ⁇ beads with 300-A pores (The Nest Group, Southborough, MA). A 2.1-mm-internal diameter x 10-mm-length guard column of the same material was fitted immediately upstream of the analytical column. Separation was performed as described previously (14).
- the iTRAQ set was diluted with the loading buffer (Buffer A: 15 mM KH2P04 in 25% acetonitrile, pH 3.0) to a total volume of 1.8 ml_, and the pH was adjusted to 3.0 with phosphoric acid.
- the solution was then filtered using a 0.45- ⁇ syringe filter (Millipore, Cambridge, ON, Canada) before loading onto the column. Separation was performed using a linear binary gradient over 1 h, plus 30 min of column re-equilibration (see Table 2a).
- Buffer A was identical in composition to the loading buffer; Buffer B was Buffer A containing 350 mM potassium chloride; Buffer C was Buffer A containing 1 M potassium chloride.
- the separation column used was a 75 ⁇ m-internal-diameter x 150-mm-length capillary column (Integrafit capillary from New Objective, VVoburn, MA) packed in-house with 3.5- ⁇ C18 beads with 100-A pores from Kromasil (Akzo Nobel/EKA Chemicals Inc. , NY).
- the flow rate used for separation on the RP column was 200 nL/min.
- Solvent A was 5% acetonitrile in 0.1 % formic acid;
- Solvent B was 95% acetonitrile in 0.1 % formic acid.
- the solvent gradient is detailed in Table 2b.
- a new column was used for each iTRAQ set.
- MS/MS spectra were processed by the software ProteinPilot version 2.0.1 , using the Paragon algorithm (15), and against a concatenated Swissprot/Panther database of 66082 distinct human protein entries (132164 entries after the reversed sequences were added). Protein identification was performed using a confidence threshold of 95% (ProteinPilot Unused score > 1.31 ) with M MTS selected as cysteine modification, and with the search option 'emphasis on biological modifications' checked.
- Tolerance windows were set at ⁇ 120 ppm for m/z and ⁇ 720 s for elution time.
- the dynamic exclusion list was maintained for a 30-s time window after the first MS/MS scan at any particular m/z ratio.
- the inventors used the report for the analysis of all 25 fractions to generate a single exclusion list comprising m/z values of peptides and their elution times, before starting the second iteration.
- the report for the second iteration of the same fractions was, in turn, used to generate a second list that was added to the first exclusion list before the third iteration was performed.
- the inventors altered the strategy from using all fractions to generate exclusion lists, to generating exclusion lists from groups of five fractions. This allowed for a more efficient process, with a second group of five fractions being acquired during the computer analysis of the first group.
- This approach not only maximized the use of available mass spectrometer and computer time, but also limited the time between the three RP separations of the same fraction by the chromatographic column. This latter point reduced the risk of a shift in the elution time due to column aging, or worse: replacement following a column failure.
- a final data analysis was performed on the complete series of the 25 data files to generate a data report encompassing all 25 fractions of each iTRAQ set.
- This analysis of all 25 fractions as a single group enhanced the number of positive identifications and the identification confidence, by regrouping different peptides of the same protein scattered across many fractions.
- Iterative iTRAQ results were compared using an Excel-based program from Applied Biosystems (noncommercial product, courtesy of Dr. Sean Seymour). The protein reports were entered into this Excel program; specialized macros then compared the identified proteins to a master list, determining which proteins were on this list, and which were not. By using the results from the first iteration as the master list, the inventors were able to determine how many proteins the second and third iterations added to the global list of identified proteins
- differentially expressed proteins were sorted according to the following criteria: (1 ) they must have been identified in at least half of the EmCa samples; (2) differential expression was signified by a t-test or by iTRAQ ratios that were > 1.5 or ⁇ 0.67 in at least half of the samples in which the protein was identified. Ideally, this trend among the samples was uniform across all samples for each protein; to validate a trend, no more than one outlier was accepted per protein. Proteins that had been found differentially expressed in the inventors' previous study (12) were also specifically selected for comparison.
- the Ingenuity Pathways Analysis software version 7.5 (Ingenuity Systems, Redwood City, CA), was used to interrogate a database of literature on the proteins of interest. Maps of protein interaction network were generated using this software.
- Endometrial tissue samples were retrieved from the University Health Network (UHN) tissue bank and from an in-house, dedicated, research endometrial tissue bank, as described elsewhere (9a). Procurement of patient samples and tissue-banking procedures were all approved by the Research Ethics Boards of York University and the UHN. Briefly, samples were taken from hysterectomy specimens (procured with patient's consent) that had been flash-frozen in liquid nitrogen within 20 min of devitalization. Samples were first sectioned for histological diagnoses.
- UHN University Health Network
- PBS phosphate buffer saline
- a pooled reference sample was created by mixing 100 ⁇ g of protein lysates, retrieved from 10 additional normal proliferative samples analyzed earlier (9a). Tissue homogenates were clarified by centrifugation, and the protein concentrations were determined using a Bradford assay (Bio-Rad, Hercules, CA).
- Each iTRAQ set included one pooled reference sample and any three samples of the different types, with the exception of the last two sets that were each composed of one reference sample and only two individual tissue samples.
- 100 ⁇ g of protein lysate were digested using sequencing grade trypsin (Promega, Madison, VVI) and labeled individually with an iTRAQ tag, as described earlier (9a). These labeled samples were mixed in sets of four, with each set containing one label of each type, resulting in a total of eight sets (Table 15).
- the separation column was a 75 ⁇ m-internal diameter x 150-mm-length capillary column (Integrafrit capillary from New Objective, VVoburn, MA) packed in-house with 3.5- ⁇ C18 beads with 100-A pores from Kromasil (Akzo Nobel/EKA Chemicals Inc. , NY).
- the flow rate used for separation on the RP column was 200 nl_ / min. Gradient elution was performed with solvent A (5% methanol in 0.1 % formic acid) and solvent B (95% methanol in 0.1 % formic acid); details of the gradient are given in Table 9b.
- Samples were analyzed on a Q-STAR Pulsar mass spectrometer (Applied Biosystems / MDS SCIEX, Foster City, CA) in information-dependent acquisition mode, with the scan cycles set up to perform a 1-s MS scan followed by five MS/MS scans of the five most abundant peaks for 2 s each, with a dynamic exclusion period of 30 s.
- Performance of the analytical system including sensitivity, resolution, and mass accuracies, was evaluated and re-optimized at minimum once every 48 h (and more frequently, when necessary), using 80 fmol of a bovine serum albumin tryptic digest.
- MS/MS spectra were processed by ProteinPilot version 2.0.1 , using the Paragon algorithm (12a), and the search was performed against a concatenated Swiss-Prot / Panther database of 66,082 distinct human protein entries (132, 164 entries after addition of the reverse database), with M MTS selected as the cysteine modification, and the search option 'emphasis on biological modifications' checked.
- the protein identification was performed simultaneously on five consecutive fraction acquisitions, using ProteinPilot unused score > 1.31 as the limit (confidence threshold of 95%), for creating the exclusion list.
- ProteinPilot software was used to generate a report of peptides identified after every analysis of five fractions. This report provided the m/z values of the identified ions and elution times of the peptides to be imported into the exclusion list of the inventors' acquisition method.
- Perl® language an in-house tool was developed to sort peptides and remove redundant peptide entries in the summary report, as discussed above.
- the tolerance window was set at ⁇ 120 ppm for m/z and ⁇ 720 s (6 min) for elution time.
- the dynamic exclusion list was maintained for a 30-s time window after the first MS/MS scan at any particular m/z ratio.
- the report generated from the second iteration of these fractions was used to generate a new list of m/z values to be added to the first exclusion list before initiation of the third iteration.
- This approach allowed for a more efficient process, with a second set of five fractions being acquired during software analysis of the first set. Iterative iTRAQ results were compared using custom macros in Excel files from Applied Biosystems (noncommercial product). Using results from the first iteration as a master list, these macros permitted the inventors to determine the additional number of proteins identified in the second and third iterations.
- iTRAQ ratios were analyzed to determine differentially expressed proteins in endometrial cancer and non-cancerous tissue samples. iTRAQ ratios for proteins identified by accession numbers in Matlab (version 7.7.0.471 ) were screened to sort significantly different proteins (p ⁇ 0.05). These ratios were normalized using the ratio for cytoplasmic beta-actin as the control. The mean expression ratio of a given protein in each sample was averaged across different runs.
- the mean expression ratios were used to identify differentially expressed proteins using the following criteria: (1 ) the protein must have been identified in at least three of the samples for each EmCa subtype; (2) at least three of the expression ratios should be ⁇ 0.67 or > 1.5 in one subtype; and (3) only one outlier is accepted (e.g. , one ratio ⁇ 0.67 along an otherwise up-regulated trend).
- the inventors Following the analysis of the first iTRAQ set, the inventors improved the precision of fraction injection and widened the exclusion windows for both time and m/z, from ⁇ 5 to ⁇ 7 min, and from 100 mDa to 120 ppm, to mitigate some of these challenges.
- the inventors chose not to exclude ions based on differences of charge or modification, as this would have increased the exclusion list beyond a practical size; the inventors also reasoned that some redundancy based on differences of charge or modification may increase the confidence of identification. Thus, the inventors' iterative analyses struck a balance between the depth of analysis and tractability.
- the second analysis added 78% more peptides to those identified after the first analysis, while the third analysis added 36% more peptides.
- about 12% of the peptides were common to any two successive analyses; however, only 3 to 6% of the peptides were identified in all three iterations.
- These additional peptides were expected to improve the coverage of already identified proteins.
- this identification via multiple peptides meant that the number of identified proteins increased at a slower pace, as shown in Table 4.
- the second iteration added 34% new proteins on average, while the third iteration added only 14% to the combined list from iterations 1 and 2; thus, there was little incentive to perform further iterations.
- 40% of the proteins identified during the first run were also identified in the next two iterations (Table 4).
- EmCa biomarker candidates Differentially expressed proteins that passed the criteria detailed in Example 7 are considered as EmCa biomarker candidates and are listed in Table 5a. Of the 40 biomarker candidates, 38 were identified with a minimum of two peptides. The two exceptions are included because they were identified with > 99% in confidence and manual inspection showed excellent MS/MS spectral quality. Since the EmCa and normal proliferative tissues were sampled from different individuals - and, therefore, were not identical (i.e., the analyses were not replicates) - the typical concept of standard deviation in quantitative analysis may be misleading as a measure of analytical quality.
- the inventors have opted to express the distributions of combined analytical and individual variability in the following manner: in the analyses, 28% of individual iTRAQ values deviate within ⁇ 10% from the means, 54% within ⁇ 20%, and 88% within ⁇ 50%. These distributions support the inventors' hypothesis that a 50% change in iTRAQ ratios is indicative of differential expression.
- EmCa biomarkers Two of the proteins listed in Table 5a have previously been identified by the inventors as potential EmCa biomarkers (12, 13): alpha-1-antitrypsin (AAT) and macrophage capping protein (CAPG).
- AAT alpha-1-antitrypsin
- CAG macrophage capping protein
- Manual checking of the data for previously identified biomarker candidates proteins yielded pyruvate kinase (PK) and creatinase kinase B (CKB) with expression ratio changes just under 50% (PK, 1.43; and CKB, 0.69) in this study, but meeting the other two criteria (vide supra). Indeed, there is a group of 16 proteins in this category that satisfy almost all of the criteria; these are listed in Table 5b, and may also be considered as EmCa biomarker candidates.
- PK pyruvate kinase
- CKB creatinase kinase B
- AAT and PK as EmCa biomarkers have already been verified on an independent cohort of 148 patients by means of immunohistochemistry on a tissue microarray format (20). Observation of differential expressions for these four proteins (AAT, CAPG, CKB, and PK) further confirms that these proteins are, indeed, excellent biomarker candidates.
- a number of proteins in Table 5b also complement some of the differentially expressed proteins in Table 5a. For example, type I keratin 18 (KRT18, Table 5b) usually forms a dimer with type I I keratin 8 (KRT8, Table 5a); overexpression of both proteins accords with this expectation. Pathway analyses for the differentially expressed proteins in Tables 5a and 5b generated the interaction networks shown in FIGs. 1 -3.
- these proteins appear to group around known key proteins involved in tumorigenesis (e.g. , p53 and interleukins in FIG. 1 , ERK and NF kappa B in FIG. 2, and tumor necrosis factor and Erbb2 in FIG. 3).
- CTSB cathepsin B
- EmCa cathepsin B
- CTSB and other cathepsins have been found upregulated in human cervical and mouse pancreatic cancers (21 , 22). It has been shown that knocking out CTSB in a mouse model of pancreatic islet cell carcinogenesis can result in reduced tumor and blood vessel formation (22).
- Phosphorylation of eukaryotic translation initiation factor 2 subunit 1 (EIF2S1 ) is an important pivotal event for both cell death and NFKB pathway activation.
- EIF2S1 When phosphorylated, this protein will inhibit the translation of a number of non-essential genes, while promoting the expression of GADD153, which will eventually result in cell cycle arrest (23).
- EIF2S1 is also the target of various stress-specific kinases, and its phosphorylation will lead to the activation of NFKB (21 ).
- Astrocytic phosphoprotein PEA-15 is known for inhibiting many apoptosis signaling cascades (25). Its overexpression has been demonstrated to increase the susceptibility of skin cells to tumor-inducing chemicals (22).
- S100 calcium-binding protein A6 (S100A6, calcyclin) is known to be upregulated in pancreatic cancer (26) and has also been suggested to be a prognostic marker of aggressive forms of brain medulloblastoma (27). S100A6 inhibition has been shown to reduce cancer cell proliferation (26).
- calcyphosin (CAPS), which may have a role in ion transport regulation
- calumenin (CALU), which is involved in protein folding and sorting in the endoplasmic reticulum, have been found to be involved in ovarian carcinoma in humans and esophageal carcinoma in rats, respectively (28, 29).
- KRT18 and KRT8 which are known to associate, provide an example in which the differential expressions of two proteins are mutually consistent.
- the KRT18/8 dimer has been shown to be involved in cell signaling in rat epithelial cells, as it binds to 14-3-3 proteins and also interacts with Raf kinase (30, 31 ).
- Other proteins are less directly connected to tumorigenesis, because they are involved in cell motility and cytoskeletal formation - although, like keratin above, they may also be involved in the regulation of cell signaling cascades.
- F-actin capping protein subunit beta (CAPZB) and macrophage capping protein (CAPG), both of which control cell motility by binding to actin filaments (32), were upregulated in the inventors' study.
- Villin 2 (EZR, Table 5b), which has similar actin-binding functions, is upregulated as well.
- EZR Villin 2
- FLNA filamins alpha
- FLNC gamma
- EmCa Proteins acting on the extracellular matrix were also found differentially expressed in EmCa. Downregulation of alpha-1-antitrypsin (AAT or serpin A1 ) was found to be associated with EmCa (12, 13), malignant lymphoma, and liver, lung, stomach, bladder, and gall-bladder cancers (33, 34). Serpin H1 , also known as heat-shock protein 47 kD (hsp47), is a collagen-binding chaperone that is generally co-expressed with collagen precursors (35). A number of proteins are linked to inflammation and the activation of T-cells.
- AAT or serpin A1 alpha-1-antitrypsin
- ILF3 interleukin enhancer- binding factor 3
- IL2 interleukin 2
- ILF3 has been found upregulated in breast cancer (37).
- Another such upregulated protein that is linked to inflammation is annexin A1 (ANXA1 , Table 5b).
- ANXA1 annexin A1
- Table 5b The gene encoding ANXA1 has been found upregulated in intrahepatic cholangiocarcinoma (38).
- annexin A1 is upregulated in mast cells, which are attracted by tumors and infiltrate cancer tissues where they may either impede or promote tumor growth, depending upon the local conditions (39).
- Annexin A1 participates in the local inflammatory regulation, and, therefore, in the immune response to the tumor.
- annexin A1 may result from invasion of the tumor by mast cells or similar partners of the immune system.
- Fibrinogen all 3 chains
- haptoglobin haptoglobin
- apoliproteins A-l and A-l l were found downregulated in EmCa samples.
- Fibrinogen is a blood protein, the synthesis of which is positively regulated by an area of inflammation-related cytokines, including interleukin 1 , interleukin 6, and tumor necrosis factor alpha (TNF) (40).
- TNF tumor necrosis factor alpha
- fibrinogen chains are synthesized by cancer cells in cervical and breast carcinomas (43).
- Haptoglobin is another blood protein, a peptidase involved in free plasma hemoglobin degradation. Its expression and blood concentration are promoted by inflammatory-related cytokines (40), notably by interleukine-1 beta via the protein Myd88 in mouse (44). This protein has been shown to be downregulated in hepatocarcinoma (45) and in AFP-negative hepatoblastoma (46); however, it was upregulated in the more aggressive AFP-positive hepatoblastoma (46) and in Her-2/neu breast cancer (47). The inventors' finding that HP is downregulated in Type I EmCa, which is typically less invasive than Type I I EmCa, is in line with these previous studies.
- apolipoprotein A-l was found to be overexpressed in high-grade dysplastic cervical cells (48), but underexpressed in the serum of ovarian cancer patients (49). Alteration of the level of APOA1 may be a part of the host inflammatory response to the tumor, but the basis for this change is unclear.
- the other lipoprotein, apolipoprotein A-l I (APOA2), is known to be upregulated in alpha fetoprotein (AFP)-positive hepatoblastoma, following the same pattern as HP (46). Upregulation of annexin A1 , together with the downregulations of apolipoproteins, fibrinogens, and haptglobin, could be indicative of suppression of the inflammatory process in tissues surrounding the tumor.
- HNRNPA1 ribonucleoprotein A1 .
- HNRNPA1 also binds DNA telomeres and promotes telomere elongation. Preventing HNRNPA1 binding to telomeres leads to the apoptosis of tumor cells (51 ). As telomere degradation in fast- proliferative cells eventually triggers p53-dependent cell-cycle arrest, it has been postulated that HNRNPA1 action results in cancer cells bypassing cell cycle arrest mechanisms (51 ).
- PK pyruvate kinase
- LDHA lactate dehydrogenase A
- PEP phosphoenol pyruvate
- PK expresses as a number of isoforms, with M 1 and M2 being splice variants of a single gene (52).
- M1 isoform is expressed in skeletal muscle and brain tissue, while the M2 isoform is found in fetal tissue and proliferating cells, especially cancer cells.
- the presence of the M2 isoform of PK is critical during the process of tumorigenesis.
- Protein Identification A total of 1518 non-redundant proteins were identified across the eight iTRAQ sets with a Local FDR value ⁇ 5%. Of these, 1068 were identified by multiple peptides with > 95% confidence scores, at an average of 8.3 peptides per protein. Of the remaining 450 proteins identified by a single peptide, 41 1 peptides showed > 99% confidence scores. Of all proteins identified, 1 194 (79%) were reported with iTRAQ ratios; however, only 732 (48%) had iTRAQ ratios where the p-value was ⁇ 0.05, and these, therefore, were eligible for further analyses.
- the second analysis identified an additional 44% unique peptides on top of the peptides identified in the first analysis; the third iteration added a further 27% unique peptides to the two combined analyses. Eight percent of the peptides were observed in all three analyses despite the use of an exclusion list. These repeated identifications were due to shifts in retention times, peak tailing, multiple charge states, and modifications (e.g., deamidation and methionine oxidation) (10a). Table 7 shows the number of unique proteins identified in the eight sample sets by the iterative analysis. The second and third analyses of the same set identified, on average, 45% and 15% additional proteins, respectively. One notable outlier is set 8, for which the first analysis was sub- optimal.
- the second and third analyses compensated for the first and provided higher numbers of protein identifications, thereby demonstrating one of the advantages of an iterative approach: viz., as each sample is analyzed several times, this approach is self-compensating and consequently results in a more rounded performance.
- Table 8 shows the average expression ratios for the differentially upregulated and downregulated proteins in the three subtypes of high-grade EmCa, along with the number of samples in which each given expression was confidently determined.
- the maximum numbers are: endometrioid, 6; clear cell, 5; serous, 8; and (individual) normal proliferative, 3.
- the 11 reported upregulated and 7 downregulated proteins should permit discrimination among the endometrioid, clear cell, and serous EmCa subtypes, particularly when used as a panel.
- LDHA L-lactate dehydrogenase A
- PGK1 phosphoglycerate kinase 1
- PK pyruvate kinase
- alpha-1 -antitrypsin and apolipoprotein A1 from Table 8 were also found downregulated in one or more previous studies (8a-10a). It should be noted that the last five are blood proteins: their lower concentrations in cancer tissues more likely result from tumors being differently vascularized from healthy tissues, rather than the proteins being underexpressed in the tumors themselves.
- HNRNPA1 and HNRNPA2B1 heterogeneous nuclear ribonucleoproteins A1 and A2/B1 (HNRNPA1 and HNRNPA2B1 , respectively) observed in the endometrioid samples correlate well with elevated levels found in studies on other forms of cancer, including breast, small-cell lung, and ovarian carcinomas (14a).
- This trend is consistent with the known function of these proteins, since protein HNRNPA1 and its splice variants A2/B1 bind telomeres to protect them, and thereby promote cancer cell survival and proliferation by ensuring telomere stabilization.
- nucleophosmin Another protein listed in Table 8 that promotes cell survival is nucleophosmin (NPM 1 ).
- NPM 1 nucleophosmin
- This nucleolar protein is involved in many functions related to embryonic development and genome stability (15a). Its suppression favors tumorigenesis, but its overexpression promotes cell proliferation and cell survival.
- NPM 1 overexpression not only results in re-localization of the p53 protein into the nucleus, but also inhibits p53 accumulation in the mitochondria (16a). Since p53-dependent apoptosis pathways rely on p53 transactivating mitochondrial genes and promoting other changes in mitochondrial activities, this described interaction of NPM1 with p53 is, in effect, anti-apoptotic.
- NPM1 can be posttranslationally modified in a variety of ways, with the acetylated form being the one involved in anti-apoptotic activities (15a).
- Nucleoside diphosphate kinase B (NME2) expression is progesterone-dependent, and reportedly has anti-tumorigenic properties in ovarian cancer: it inhibits the growth of ovarian cancer cell lines and also suppresses cell motility (19a). Upregulation of this protein in EmCa suggests that it is also involved in EmCa, but apparently not in a tumor suppressor capacity.
- Peroxiredoxin-5 (PRDX5) is expressed in response to oxidative stress, and has a protective role during inflammation. Silencing PRDX5 by RNA interference leads to an increase of caspase activity in mitochondria and cell apoptosis (20a). This could be a consequence of the increase of oxidative stress, rather than a direct interaction between PRDX5 and caspase proteins.
- Cystatin B is a cystein proteinase inhibitor, and may actually have a role in cancer: it may act against cancer proliferation and metastasis by inhibiting extracellular matrix degradation by proteinases like cathepsin B (21 a). Its expression in mouse uterus is increased by estradiol (22a). Since estrogen receptors are known to be overexpressed in EmCa (23a-25a), the overexpression of CSTB may be linked to this event.
- Inorganic pyrophosphatase (PPA1 ) is only loosely related to cancer: it has been shown to bind to p53 (26a), but the physiological meaning of this interaction is totally unknown. Given the importance of p53 in EmCa (see below), PPA1 may be a valuable candidate for further study.
- PK has again been observed to be upregulated in all EmCa subtypes (see Table 9). This is typical of the Warburg effect, characterized by the overexpression of pyruvate kinase M2 and the shift of the glycolytic pathway toward lactate production under aerobic conditions, a feature which has been found in many cancer cells (27a). In this light, the upregulation of other proteins related to glucose metabolism is consistent with the Warburg effect. Notably, glucose-6 phosphate isomerase, alpha-enolase, and PGK1 were upregulated in all subtypes (Table 9).
- LDHA is also upregulated in the endometrioid and clear cell subtypes (Table 8). LDHA has previously been shown to be upregulated in human and murine models of lung carcinoma (28a). Since the function of this enzyme is to convert pyruvate into lactate, its upregulation is to be expected following the upregulation of PK. It was initially puzzling that LDHA was not found to be upregulated in the serous subtype; upon a closer inspection of the data, it became apparent that the iTRAQ ratio was, in fact, above 1.5 in four serous carcinoma samples, although close to 1 for the remaining four samples.
- inactivation of the tumor suppressor PTEN is the most often reported alteration. This inactivation is common in estrogen-dependent endometrioid EmCa, generally in conjunction with defects of DNA mismatch repair genes, and is believed to be an early event in carcinogenesis. Less frequent, but still specific to the endometroid subtype, are mutations in the Kras and/or beta-catenin oncogenes. By contrast, an early event in estrogen-independent serous carcinomas is apparently a mutation and overexpression of the protein p53, a central regulator of cell cycle and differentiation.
- EmCa Another important alteration in serous EmCa is the mutation and amplification of HER-2/neu. None of the protein products of these genes was detected in the current study; these are all expected to have low abundances and are located in the nucleus or membranes, compartments that are poorly sampled in current proteomic technologies.
- FIG. 4 was constructed by merging five interacting networks, the top functions of which include cell cycle and death; cancer; molecular transport; cellular development, growth, and development; and signal transduction.
- FIG. 5 comprises a single network centering on cell death and cellular growth and proliferation.
- Adachi O.; Kawai, T.; Takeda, K.; Matsumoto, M.; Tsutsui, H.; Sakagami, M.; Nakanishi, K.; Akira, S. Targeted disruption of the MyD88 gene results in loss of IL-1- and IL-18-mediated function. Immunity 1998, 9(1), 143-150.
- Buffer A 15 mM KH 2 P0 4 in 25% acetonitrile, pH 3.0
- Buffer B Buffer A containing 350 mM KCI
- Buffer C Buffer A containing 1 M KCI
- Solvent A 5% acetonitrile in 0.1 % formic acid
- Solvent B 95% acetonitrile in 0.1 % formic acid. Table 3. Distribution of protein functions for biomarkers identified in the Type I EmCa
- Buffer A 15 mM KH 2 P0 4 in 25% acetonitrile, pH 3.0
- Buffer B Buffer A containing 350 mM KCI
- Buffer C Buffer A containing 1 M KCI
- Solvent A 5% acetonitrile in 0.1 % formic acid
- Solvent B 95% acetonitrile in 0.1 % formic acid Table 7. Number of proteins identified by the iterative method in the EmCa subtype analysis.
Landscapes
- Health & Medical Sciences (AREA)
- Chemical & Material Sciences (AREA)
- Life Sciences & Earth Sciences (AREA)
- Organic Chemistry (AREA)
- Proteomics, Peptides & Aminoacids (AREA)
- Wood Science & Technology (AREA)
- Analytical Chemistry (AREA)
- Zoology (AREA)
- Immunology (AREA)
- General Health & Medical Sciences (AREA)
- Pathology (AREA)
- Genetics & Genomics (AREA)
- Engineering & Computer Science (AREA)
- Epidemiology (AREA)
- Hospice & Palliative Care (AREA)
- Veterinary Medicine (AREA)
- Physics & Mathematics (AREA)
- Animal Behavior & Ethology (AREA)
- Biophysics (AREA)
- Public Health (AREA)
- Microbiology (AREA)
- Biotechnology (AREA)
- Oncology (AREA)
- Molecular Biology (AREA)
- Biochemistry (AREA)
- Bioinformatics & Cheminformatics (AREA)
- General Engineering & Computer Science (AREA)
- Measuring Or Testing Involving Enzymes Or Micro-Organisms (AREA)
Abstract
The invention provides markers for endometrial cancer, polynucleotides encoding same, and precursors thereof. The invention also provides methods for detecting, diagnosing, screening for, monitoring, assessing, and treating endometrial cancers and related disease conditions in a subject. The invention further provides a method of selecting for or assessing efficacy of agents against endometrial cancer, and a method for assessing endometrial cancer cell carcinogenic potential of a compound. The invention further provides localization and imaging methods for endometrial cancers. Also provided are diagnostic compositions and kits for carrying out methods of the invention. In addition, the invention provides therapeutic applications for endometrial cancers which employ protein endometrial cancer markers and polynucleotides encoding same, and binding agents for the markers. Moreover, the invention provides an iterative computerized method and a system for identifying at least one protein endometrial cancer marker.
Description
ENDOMETRIAL CANCER BIOMARKERS AND METHODS
OF IDENTIFYING AND USING SAME
CROSS REFERENCE TO PRIOR APPLICATIONS
[0001] The present application claims priority under the Paris Convention to US Application Number 61/344,508, filed August 10, 2010, the entire contents of which are incorporated herein by refernce.
FIELD OF THE INVENTION
[0002] The invention relates to endometrial cancer markers, methods for assessing the status of endometrial cell tissue, and methods for the detection, diagnosis, prediction, monitoring, and therapy of endometrial cancer. One aspect of the invention relates to biomarkers of endometrial carcinoma and methods for detection, diagnosis, prediction, monitoring, and therapy for endometrial carcinoma and related conditions.
BACKGROUND OF THE INVENTION
[0003] Endometrial carcinoma (EmCa) is the fourth most prevalent cancer in North America women (10, 1 a), and the most common invasive carcinoma of the female genital tract. Definitive diagnosis is mostly based on histological examination of tissues obtained after a biopsy, an invasive procedure typically performed as a result of investigative diagnosis following abnormal uterine bleeding at presentation.
[0004] EmCa is a heterogeneous disease, classified according to histological cell type, with the most common being endometrioid, serous, and clear cell carcinoma (2a). The two main types of EmCa are Type I and Type I I. Type I is of endometrioid histology, while the more clinically aggressive Type I I is primarily serous in morphology. Clear cell carcinoma is also grouped under Type I I EmCa, and can sometimes be present admixed with endometrioid and serous tumor components (1 1 ). All serous and clear cell carcinomas are regarded as high grade, with the endometrioid subtype being further stratified according to the FIGO (Federation Internationale de Gynecologie et d'Obstetrique or International Federation of Gynecology and Obstetrics) grading system - a three-tier system which may be binarised by combining grades I and I I into a low-grade subset, with grade I I I constituting high-grade endometrioid carcinoma. Within the rubric of high-grade endometrial cancers, then, are grade I I I endometrioid, serous, and clear cell carcinomas. The different EmCa cell types reflect distinctive epidemiological risk factors and profiles; they exhibit different biology, behavior, and metastasis profiles, and are associated with different precursor lesions.
[0005] Both cell type and grade are used to guide the extent of surgery and the use of adjuvant therapy. In a retrospective review of 187 high-grade endometrial carcinomas, Soslow et al.
highlighted that serous carcinomas are more likely to be treated with chemotherapy, while grade I II
endometrioid carcinomas are more likely to be subject to adjuvant radiation (3a). Since treatment protocols are cell-type-specific, accurate, reproducible, and biologically relevant histological classification is essential. Some studies have shown similar outcomes among subtypes of high-grade endometrial cancer; however, others have suggested that, even within this category, subtype is critical in determining patient outcome.
[0006] Given its role in identifying EmCa subtypes, histology is pivotal. Inter-observer studies have shown moderate to excellent reproducibility from sample to sample within a given cell type, when considering all grades and cell types. Reproducibility would undoubtedly improve if only high- grade subtypes were considered. Morphological overlap exists between the subtypes, and, in the case of clear cell and serous, there is morphological and clinicopathological overlap. Increasingly, immunohistochemical markers are used to refine cell type in difficult cases. In addition, biopsy samples are small and may be poorly representative (e.g., diagnostic features that are focal may not be sampled).
[0007] In terms of FIGO grading, there is also the potential issue of inter-observer variability. The histological cut points have been defined arbitrarily, and are not the result of an outcome-driven process. In ovarian cancer, a need for subtype-specific therapies has recently been recognized. Certainly, subtype-specific chemotherapeutic agents are also required for the subtypes of endometrial cancers. The application of the FIGO grading system requires histological assessment of the percentage of solid growth within the tumor and the degree of cytologic atypia. Thus, quantitative proteomics may provide distinguishing differences in protein expression levels, as well as surveillance and therapeutic targets specific to given histological types.
[0008] In the past decade, numerous studies have analyzed cancer cells and tissues using a proteomics approach (i.e. , by analysis of the global protein content of the cells or tissues). One frequent objective of these studies has been the identification of markers specific to the tumor tissue, based on comparison with normal tissues (4a). Conceptually, these biomarkers can provide molecular discrimination, aid in the diagnosis of the different types and subtypes of EmCa, and lead to prognosis of patient outcome. In addition, blood-borne biomarkers can aid in diagnosis of disease, perhaps at an earlier stage. An excellent example of this is the prostate specific antigen, PSA, for the diagnosis and recurrence monitoring of prostate cancer (http://www.cancer.gov/cancertopics/types/ prostate). It is expected that attempts at early detection and treatment of cancer would benefit from the discovery of biomarker proteins, specifically those over- or under-expressed in a malignant tissue.
[0009] Numerous proteomics studies undertaken in recent years have attempted to discover biomarkers, including EmCa biomarkers, through comparison of the protein contents of diseased versus non-diseased samples. The various issues, pitfalls, and successes of these proteomics-based biomarker discovery studies have been discussed and reviewed (1 -5). As described by LaBaer (1 ), "biomarkers are biometric measurements that convey information about the biological condition of the
subject being tested". Ideally, a biomarker should be a molecule expressed at a higher concentration by an abnormal tissue, such as a nodule of cancer cells.
[0010] Proteins are good biomarker candidates because protein expression is closely linked to the metabolic state of the cell. This is particularly pertinent to cancer diagnosis: tumor cells that are undergoing rapid uncontrolled proliferation are bound to have biochemical changes that would manifest as changes in expression levels of specific proteins. To identify biomarkers, a quantitative or, at a minimum, semi-quantitative approach is required, as it is expected that all healthy and tumor cells will express a common core of proteins, and that most changes will be in terms of relative expression levels rather than absolute presence or absence of a given protein. Biomarker candidates include proteins for which patterns of differential expression are specific for a disease condition, and could be either causative or symptomatic in nature.
[0011] A proteomics approach to biomarker discovery consists of a quantitative comparison of the proteins expressed between tissues at two or more different states, including different subtypes of the disease. Proteins that are differentially expressed in distinct disease subtypes can be useful as specific diagnostic and prognostic markers, since different subtypes have different etiologies. One of the challenges in the quest for biomarkers is the selection of molecules that show both selectivity and specificity for the studied illness, particulaly if the final objective is to assay these biomarkers in a body fluid, such as blood (see, e.g. , references 4a-7a for discussions of biomarker discovery challenges).
[0012] Currently, quantitative comparisons are conveniently achieved via isotope dilution mass spectrometry (IDMS) by first chemically labeling enzymatically digested samples before combining the control and test samples for single handling. One of the more common labeling reagents is iTRAQ (6). The combined sample set is then separated - typically, by multidimensional liquid
chromatography (LC) - with the last stage coupled to online electrospray ionization (ESI) tandem mass spectrometry (MS/MS). The peptides identified are mapped to proteins in a so-called "bottom- up" strategy. Quantification is performed via the reporter ions, m/z 1 14-1 17, generated from the iTRAQ tags under MS/MS. The tagging serves to distinguish proteins originating from the individual samples in the mixture; a comparison among the abundances of the reporter ions results in relative quantification of the given peptide and the protein from which the peptide originates. Due to the complexity of tissue samples, a prefractionation of the samples is typically performed using strong cation exchange (SCX) prior to the reverse phase (RP) separation that is coupled online with MS/MS.
[0013] One limitation of the foregoing bottom-up approach is the inability of the mass spectrometer to analyze all available peptide ions. Despite two-dimensional chromatographic separation, very often a large number of peptides coelute because of the complexity of tissue samples. The mass spectrometer is typically unable to examine every single peptide that coelutes. The MS software, in particular, tends to favour analysis of the most abundant peptides at the expense of coeluting peptides of lower abundance. As many signaling and regulatory proteins are expressed
in lower concentrations than structural proteins and metabolic enzymes, this shotgun, bottom-up approach tends to miss out on acquisition of the most valuable information (7).
[0014] A possible solution is iterative analysis of the same sample, using an exclusion list of identified peptides to instruct the mass spectrometer to ignore those ions that have already been selected in the previous run(s), when choosing ions for MS/MS analysis in subsequent runs. Attempts at iterative analysis have been reported for matrix-assisted laser desorption / ionization (MALDI) MS/MS (8, 9) and ESI -MS/MS (7). The last study (7) reported an extensive comparison of four different approaches to creating the exclusion list, ranging from a simple list of identified ions to a complex list of all potential ions from identified proteins plus all selected but unidentified ions.
[0015] A key issue in any iterative approach is the confidence of the peptide identification.
Lower-abundance precursors selected in a second pass will tend to have MS/MS spectra of lower intensities than those selected in the first pass. Hence, the signal-to-noise ratios are expected to decrease as more low-abundance ions are selected for MS/MS; consequently, the peptide identifications would be more prone to false positive results. By the same token, the precisions of the iTRAQ ratios would also be lower for those peptides. For at least these reasons, there is a heightened need to assess more thoroughly the confidence levels of these identifications.
SUM MARY OF THE INVENTION
[0016] In a previous iTRAQ-labeling study involving 40 endometrial carcinoma (EmCa) and normal proliferative endometrial tissues, the inventors identified a number of differentially expressed proteins for use as EmCa biomarkers. As described in greater detail herein, the inventors have further verified and confirmed those results, and identified a new set of biomarkers, using an iterative analytical strategy in which a list of the peptide ions identified in the first run is employed as an exclusion list for the subsequent acquisition. The inventors' strategy allows progressively lower- abundance peptide ions to be selected and identified.
[0017] Using multidimensional liquid chromatography-mass spectrometry (LC-MS / MS) to analyze biological samples labelled with isobaric mass tags (iTRAQ), the inventors identified proteins that were differentially expressed in EmCa (e.g. , Type I EmCa and grade I I I endometrioid EmCa, clear cell EmCa, and serous EmCa), as compared with normal proliferative samples (control). In an analysis of Type I EmCa, ten samples were compared against ten normal proliferative samples. One thousand five hundred and twenty-nine (1529) proteins were identified in EmCa samples, including structural proteins, signalling components, enzymes, receptors, transcription factors, and chaperones. Using cutoff values of 1.5 fold for overexpression, and 0.67 fold for underexpression, the inventors were able to identify 26 underexpressed proteins, and 14 proteins that were overexpressed in EmCa, compared to the normal tissues. These cutoff values were used to select proteins for further statistical analyses in previous studies (DeSouza et al. (2007), Mol Cell Proteomics, 6: 1170-1 182; Ralhan et al. (2008), Mol Cell Proteomics, 7: 1162-73), and were found to perform satisfactorily.
[0018] The inventors' iterative analysis identified, on average, 34% and 14% new proteins in the second and third analyses, respectively. Two glycolytic proteins, pyruvate kinase (PK) and lactate dehydrogenase A (LDHA), were up-regulated, in accordance with the Warburg effect in tumor cells (Tables 9 and 8). Alpha-1 -antitrypsin (AAT) and macrophage capping protein (CAPG), which were previously found to be differentially expressed in EmCa, displayed the same respective trends (Table 5a). Other proteins that were downregulated include fibrinogen, haptoglobulin, apolipoprotein A-l and apoliprotein A-l I, and serpin H1 , all of which are involved in extracellular matrix formation, motility, and/or inflammatory reaction. Pathway analysis showed that most differentially expressed proteins map to a single network that has transcription factors and kinases linked to tumorigenesis as central protein hubs.
[0019] In an analysis of different EmCa subtypes, six FIGO grade I II endometrioid, five clear cell, and eight serous carcinomas were compared using iTRAQ labeling and LC-MS/MS analysis. Using their iterative approach, the inventors identified eighteen proteins that showed differential expression in different histological types of high-grade endometrial cancer (Table 8). Fourteen proteins showed a similar trend for all samples, and may be considered suitable candidates for use as endometrial cancer biomarkers (Table 9). Heterogeneous nuclear ribonucleoprotein A2/B1 (hnRNP A2/B1 ) was found to be specifically up-regulated in FIGO grade I I I endometrioid carcinomas. L-lactate dehydrogenase A (LDHA) was observed to be constantly up-regulated in endometrioid samples; this protein is also up-regulated in clear cell carcinomas. Hepatoma derived growth factor (HDGF) was up-regulated in the serous endometrial carcinomas (Table 8). Thus, the inventors have also identified a series of proteins which exhibit varying expression patterns across different subtypes of high-grade endometrial cancers, including grade I I I endometrioid, clear cell, and serous EmCa.
[0020] Accordingly, in one aspect, the invention provides a method for detecting one or more endometrial cancer markers in a subject by: (a) obtaining a sample from a subject; (b) detecting in the sample an amount of at least one of the endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9; and (c) comparing the detected amount with an amount detected for a standard.
[0021] In another aspect, the invention provides a method for diagnosing endometrial cancer in a subject, by comparing: (a) levels of at least one of the endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9; and (b) normal levels of expression of the corresponding endometrial cancer marker or polynucleotide in a control sample, wherein a significant difference in levels of the endometrial cancer marker, relative to the corresponding normal levels, is indicative of the endometrial cancer in the subject.
[0022] In certain embodiments of the invention, the endometrial cancer marker is a protein.
[0023] In yet another aspect, the invention provides a method for diagnosing endometrial cancer in a subject, by: (a) contacting a biological sample obtained from the subject with at least one binding
agent that specifically binds to at least one of the endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9; and (b) detecting in the sample amounts of the endometrial cancer marker or polynucleotide or part thereof that binds to the at least one binding agent, relative to a predetermined standard or cut-off value, and thereby determining the presence or absence of the endometrial disease in the subject. In one embodiment, the binding agent is an antibody.
[0024] In another aspect, the invention provides a method of screening a subject for endometrial cancer by: (a) obtaining a biological sample from a subject; (b) detecting in the sample an amount of at least one of the endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9; and (c) comparing the detected amount of the endometrial cancer marker or polynucleotide with a predetermined standard, wherein detection of a level of the endometrial cancer marker or polynucleotide different from that of the standard is indicative of endometrial cancer.
[0025] In certain embodiments of the invention, the level of the endometrial cancer marker or polynucleotide is significantly higher than the standard and is indicative of endometrial cancer. In certain other embodiments, the level of the endometrial cancer marker or polynucleotide is significantly lower than the standard and is indicative of endometrial cancer. In further embodiments, the sample is obtained from a tissue, extract, cell culture, cell lysate, lavage fluid, or physiological fluid of the subject. In still other embodiments, the sample is obtained from endometrial tumor tissue.
[0026] In a further aspect, the invention provides a method for determining the presence or absence of an endometrial cancer marker associated with an endometrial disease in a subject by: (a) detecting at least one of the endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9; and (b) relating the detected amount of the marker or polynucleotide to the presence of the endometrial disease. In one embodiment, the polynucleotide is mRNA. In another embodiment, the polynucleotide is detected by: (a) contacting the sample with at least one oligonucleotide that hybridizes to the polynucleotide; and (b) detecting in the sample levels of at least one nucleic acid that hybridizes to the polynucleotide, relative to a predetermined standard or cut-off value, and thereby determining the presence or absence of the endometrial disease in the subject.
[0027] In one embodiment, the polynucleotide is detected with an amplification reaction. In other embodiments, the amplification reaction includes a polymerase chain reaction employing
oligonucleotide primers that hybridize to the polynucleotide or to a precursor or complement thereof. Additionally, in certain embodiments, the polynucleotide is detected with at least one oligonucleotide probe that hybridizes to the polynucleotide or to a complement thereof.
[0028] In a further embodiment, the polynucleotide is detected by: (a) isolating RNA from the sample; (b) combining the RNA with at least one reagent, to convert the RNA to cDNA; (c) treating the cDNA with at least one amplification reaction reagent and at least one primer that hybridizes to the cDNA, to produce at least one amplification product; (d) analyzing the at least one amplification product to detect an amount of RNA encoding the at least one endometrial cancer
marker; and (e) comparing the amount of RNA to an amount detected against a panel of expected values for normal tissue derived using similar primers.
[0029] In another aspect, the invention provides a method for diagnosing and monitoring endometrial cancer in a subject by: (a) isolating at least one nucleic acid in a sample from the subject; and (b) detecting at least one of the endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9, wherein the presence of higher or lower levels of the marker or polynucleotide in the sample, compared to a standard or control, is indicative of the disease or prognosis.
[0030] In yet another aspect, the invention provides a method for monitoring the progression of endometrial cancer in a subject by: (a) detecting in a sample from the subject, at a first time point, at least one of the endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9; (b) repeating step (a) at a subsequent point in time; and (c) comparing levels detected in steps (a) and (b), and thereby monitoring the progression of endometrial cancer.
[0031] In a further aspect, the invention provides a method for determining in a subject whether endometrial cancer has metastasized or is likely to metastasize in the future, by comparing:
(a) levels of at least one of the endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9; and (b) normal levels or non-metastatic levels of the endometrial cancer marker or polynucleotide in a control sample, wherein a significant difference between the levels in the subject sample and the normal levels or non-metastatic levels is indicative that the endometrial cancer has metastasized.
[0032] In yet another aspect, the invention provides a method for assessing the aggressiveness or indolence of endometrial cancer, by comparing: (a) levels of at least one of the endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9; and (b) normal levels of the endometrial cancer marker or polynucleotide in a control sample, wherein a significant difference between the levels in the subject sample and normal levels is indicative that the cancer is aggressive or indolent.
[0033] In still another aspect, the invention provides a diagnostic composition including an agent that: binds to at least one of the endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9, or hybridizes to at least one polynucleotide encoding at least one of the endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9.
[0034] In a further aspect, the invention provides a method for assessing the potential efficacy of a test agent for inhibiting endometrial cancer in a subject, by comparing: (a) levels of at least one of the endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9; and (b) levels of at least one of the endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9 in a second sample obtained from the subject, wherein the second sample has not been exposed to the test agent, and wherein a significant difference in the levels of the endometrial cancer marker or
polynucleotide in the first sample, relative to the second sample, is an indication that the test agent inhibits endometrial cancer in the subject.
[0035] In yet another aspect, the invention provides a method for assessing the efficacy of a therapy for inhibiting endometrial cancer in a subject, by comparing: (a) levels of at least one of the endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9 in a first sample obtained from the subject; and (b) levels of at least one of the endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9 in a second sample obtained from the subject following therapy, wherein a significant difference in the levels of expression of the endometrial cancer marker or polynucleotide in the second sample, relative to the first sample, is an indication that the therapy inhibits endometrial cancer in the subject.
[0036] In a further aspect, the invention provides a method for selecting an agent for inhibiting endometrial cancer in a subject by: (a) obtaining a sample including precancer or cancer cells from the subject; (b) separately exposing aliquots of the sample to a plurality of test agents;
(c) comparing levels at least one of the endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9 in the aliquots; and (d) selecting one of the test agents which alters the levels of the endometrial cancer marker or polynucleotide in the aliquot containing that test agent, relative to other test agents.
[0037] In another aspect, the invention provides a method for inhibiting endometrial cancer in a subject by: (a) obtaining a sample including precancer or cancer cells from the subject; (b) separately maintaining aliquots of the sample in the presence of a plurality of test agents;
(c) comparing levels of at least one of the endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9 in the aliquots; and (d) administering to the subject at least one of the test agents which alters the levels of the endometrial cancer marker or polynucleotide in the aliquot containing that test agent, relative to other test agents.
[0038] In yet another aspect, the invention provides a method for assessing the endometrial cancer cell carcinogenic potential of a test compound by: (a) maintaining separate aliquots of endometrial cancer cells in the presence and absence of the test compound; and (b) comparing levels of at least one of the endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9, in the aliquots, wherein a significant difference in levels of the endometrial cancer marker or polynucleotide in the aliquot maintained in the presence of the test compound, relative to levels in the aliquot maintained in the absence of the test compound, is indicative that the test compound possesses endometrial cancer cell carcinogenic potential.
[0039] In a further aspect, the invention provides an in vivo method for imaging an endometrial disease by: (a) injecting a subject with one or more agents that bind to an endometrial cancer marker listed in Table 5a, Table 5b, Table 8, and/or Table 9, the agent carrying a label for imaging the endometrial cancer marker; (b) allowing the agent to incubate in vivo and bind to an endometrial
cancer marker; and (c) detecting the presence of the label localized to diseased endometrial tissue. In one embodiment, the agent is an antibody that specifically reacts with an endometrial cancer marker.
[0040] In certain embodiments of the invention, the endometrial cancer marker is listed in Table 5a, Table 5b, Table 8, and/or Table 9.
[0041] In one aspect, the invention provides a set of endometrial cancer markers, including at least 2 of the markers listed in Table 5a, Table 5b, Table 8, and/or Table 9 or polynucleotides encoding same. In one embodiment, the at least 2 markers are protein endometrial cancer markers (e.g. , L-lactate dehydrogenase A (LDHA), nucleophosmin, heterogenous nuclear ribonucleoprotein A1 , cystatin B, transferrin, nucleolin, villin 2 (ezrin), or galectin-1 ).
[0042] In a further aspect, the invention provides kits comprising the endometrial cancer markers of the invention. By way of example, the invention provides a kit for determining the presence of endometrial cancer in a subject, including a known amount of at least one binding agent that binds to at least one of the protein endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9, or a part thereof, wherein the binding agent includes a detectable substance or binds directly or indirectly to a detectable substance.
[0043] In yet another aspect, the invention provides a kit for determining the presence of endometrial cancer in a subject, including a known amount of an oligonucleotide that hybridizes to a polynucleotide encoding an endometrial cancer marker listed in Table 5a, Table 5b, Table 8, and/or Table 9, wherein the oligonucleotide is directly or indirectly labelled with a detectable substance.
[0044] In certain embodiments of the invention, the endometrial cancer marker is an EmCa marker. By way of example, the endometrial cancer marker may be a Type I EmCa marker.
Exemplary EmCa that may be detected by the markers of the present invention include, without limitation, endometrioid EmCa, clear cell EmCa, and serous EmCa. In one embodiment, the endometrioid EmCa marker is a heterogeneous nuclear ribonucleoprotein (hnRNPA). In another embodiment, the serous EmCa marker is a hepatoma-derived growth factor.
[0045] In a further aspect, the invention provides a use of a marker, or differential expression of a marker, to screen for, diagnose, or monitor the progress, treatment, recurrence, or prognosis of endometrial cancer. Exemplary markers include, without limitation, L-lactate dehydrogenase A (LDHA), phosphoglycerate kinase, pyruvate kinase isozymes M 1/M2, nucleolin, glucose-6-phosphate isomerase, alpha-enolase, villin 2, isocitrate dehydrogenase 2 (mitochondrial), peroxiredoxin-1 , transketolase, serpin H1 , transferrin, galectin-1 , and lumican, and polynucleotides encoding same.
[0046] In another aspect, the invention provides an iterative computerized method for identifying at least one protein endometrial cancer marker, by: (a) conducting a first mass spectrometry analysis of peptide ions in normal and diseased tissues, to identify a first set of potential markers based on
differential expression; (b) creating an exclusion list consisting of the peptide ions identified in the first set of potential markers, wherein the exclusion list is restricted by a mass window and a time window; (c) conducting a second mass spectrometry analysis of peptide ions in the normal and diseased tissues, to identify a second set of potential markers based on differential expression, wherein the second analysis disregards peptide ions in the exclusion list; (d) updating the exclusion list to consist of the peptide ions identified in the first and second sets of potential markers; and (e) conducting a third mass spectrometry analysis of peptide ions in the normal and diseased tissues, to identify a third set of potential markers based on differential expression, wherein the third analysis disregards peptide ions in the updated exclusion list. In one embodiment, the method includes the step of calculating a false discovery rate for each identified potential marker. Also provided is a system for identifying at least one protein endometrial cancer marker. The system includes at least one computing device including software that, when executed, performs the iterative computerized method of the invention.
[0047] Other objects, features, and advantages of the invention will become apparent from the following detailed description. It should be understood, however, that the detailed description, and the specific examples which indicate preferred embodiments of the invention, are given by way of illustration only, since various changes and modifications within the spirit and scope of the invention will become apparent to those skilled in the art from the detailed description which follows.
BRIEF DESCRIPTION OF THE TABLES AND DRAWINGS
[0048] TABLES
[0049] Table 1 is a description of each set of samples that were used in the analysis of Type I EmCa samples to find EmCa biomarkers.
[0050] Table 2a is the liquid chromatography gradient used for strong cation exchange (SCX) HPLC fractionation in the analysis of Type I EmCa samples.
[0051] Table 2b is the solvent gradient used for reverse-phase (RP) HPLC separation in the analysis of Type I EmCa samples.
[0052] Table 3 is a comparison of protein functions or categories between the proteins identified in the analysis of Type I EmCa samples and those identified in an earlier study.
[0053] Table 4 is a comparison of identified protein numbers across the iterative analysis of iTRAQ sets in the analysis of Type I EmCa samples. Numbers correspond to the unique proteins identified in the forward database. Identifications pointing to reversed sequences have been removed. Values in the "additional" columns do not include the proteins identified in two or more iterations. Percentages are calculated as the ratios of newly-identified proteins over proteins already identified in all previous iterations in %.
[0054] Table 5a is a comparison of the protein mean iTRAQ ratios for the analysis of Type I EmCa samples that meet the criteria described in the Examples. For each set, the protein mean ratios over the three analyses have been calculated, taking into account the iTRAQ p-values from ProteinPilot® quantification. Columns also display the number of samples in which the protein has been identified (#, out of 10).
[0055] Table 5b is a comparison of the protein mean iTRAQ ratios for the analysis of Type I EmCa samples in which the proteins exhibited changes in expression between 40% and 50% or expression ratios between 1.4 and 1.5 or between 0.67 and 0.71.
[0056] Table 6 shows the liquid chromatography gradients used for strong cation exchange (SCX) and HPLC fractionation in the analysis of different EmCa subtypes.
[0057] Table 7 provides the number of proteins identified by the iterative method in the eight sample sets studied in the analysis of different EmCa subtypes.
[0058] Table 8 lists the proteins from the EmCa subtype analysis which exhibited divergent expression between EmCa samples as compared to normal proliferative tissues. "#" is the number of samples in the subtype in which the given expression was confidently determined. "Mean" is the average expression ratio.
[0059] Table 9 provides a list of proteins showing similar expression trends regardless of EmCa subtype.
[0060] Table 10 shows how iTRAQ labels were assigned randomly to the pooled reference sample and tissue homogenates in the analysis of different EmCa subtypes. These labeled samples were mixed in sets of four, with each set containing one label of each type, resulting in a total of eight sets.
[0061] Table 1 1 is a comparison of iTRAQ ratios for proteins identified in three different studies of EmCa samples.
[0062] FIGURES
[0063] FIG. 1 depicts interaction network #1 for EmCa biomarker candidates. Overexpressed proteins are shown in red; underexpressed proteins are shown in green. Uncolored proteins were added by Ingenuity Pathway Analysis to connect the submitted biomarker candidates. AHSG: alpha- 2-HS-glycoprotein; ALDH1 A2: aldehyde dehydrogenase 1 family, member A2; APOA1 : apolipoprotein A-l; APOA2: apolipoprotein A-l l; APOH: apolipoprotein H (beta-2-glycoprotein I); CAP1 : CAP, adenylate cyclase-associated protein 1 ; CAPG: capping protein (actin filament), gelsolin-like; CAPZB: capping protein (actin filament) muscle Z-line, beta; CD163: CD163 molecule; CLEC1 1A: C-type lectin domain family 11 , member A; COL1 A2: collagen, type I, alpha 2; CTSB: cathepsin B; EIF4E:
eukaryotic translation initiation factor 4E; GPLD1 : glycosylphosphatidylinositol specific phospholipase
D1 ; HNF1 A: HNF1 homeobox A; HNRNPA1 : heterogeneous nuclear ribonucleoprotein A1 ; HP: haptoglobin; IL1 B: interleukin 1 , beta; IL2: interleukin 2; IL13: interleukin 13; ILF2: interleukin enhancer binding factor 2, 45kDa; ILF3: interleukin enhancer binding factor 3, 90kDa; KITLG: KIT ligand; LCAT: lecithin-cholesterol acyltransferase; LIPC: lipase, hepatic; MYC: v-myc
myelocytomatosis viral oncogene homolog; NCL: nucleolin; NOS2: nitric oxide synthase 2, inducible; PCBP1 : poly(rC) binding protein 1 ; PKM2: pyruvate kinase M1/M2; PLTP: phospholipid transfer protein; S100A6: S100 calcium binding protein A6 (calcyclin); SERPINA1 : alpha-1 antitrypsin;
SERPINH1 : serpin peptidase inhibitor, clade H (heat shock protein 47); TP53: tumor protein p53.
[0064] FIG. 2 sets forth interaction network #2 for EmCa biomarker candidates. Overexpressed proteins are shown in red; underexpressed proteins are shown in green. Uncolored proteins were added by Ingenuity Pathway Analysis to connect the submitted biomarker candidates. A2M: alpha-2- macroglobulin; ACTA2: actin, alpha 2, smooth muscle, aorta; ALB: albumin; ANXA1 : annexin A1 ; BAD: BCL2-associated agonist of cell death; CASP7: caspase 7, apoptosis-related cysteine peptidase; COL1 A1 : collagen, type I, alpha 1 ; DAPK1 : death-associated protein kinase 1 ; DUSP1 : dual specificity phosphatase 1 ; ERK: p42/p44 Map Kinase; EZR: Ezrin (villin 2); FGG: fibrinogen gamma chain; FLNC: filamin C, gamma; FSH: follicle stimulating hormone; HSPB1 : heat shock 27kDa protein 1 ; IgG: Immunoglobulin G; Jnk: Jun kinase; KRT18: keratin 18; KRT8: keratin 8; LDHA: lactate dehydrogenase A; LDLR: low density lipoprotein receptor; Lh: luteinizing hormone; LRP1 : low density lipoprotein-related protein 1 (alpha-2-macroglobulin receptor); MPO: myeloperoxidase; MT3:
metallothionein 3; PDGFB: platelet-derived growth factor beta polypeptide; PEA15: phosphoprotein enriched in astrocytes 15; PIAS4: protein inhibitor of activated STAT, 4; PLD1 : phospholipase D1 , phosphatidylcholine-specific; Raf: Raf kinase; SCARB1 : scavenger receptor class B, member 1 ; TCHP: trichoplein, keratin filament binding; TF: transferrin; TFRC: transferrin receptor (p90).
[0065] FIG. 3 presents interaction network #3 for EmCa biomarker candidates. Overexpressed proteins are shown in red; underexpressed proteins are shown in green. Uncolored proteins were added by Ingenuity Pathway Analysis to connect the submitted biomarker candidates.
[0066] ADAMTS1 : ADAM metallopeptidase with thrombospondin type 1 motif; BIRC5:
baculoviral IAP repeat-containing 5; CD44: CD44 antigen; CETP: cholesteryl ester transfer protein, plasma; COL1 A1 : collagen, type I, alpha 1 ; CP: ceruloplasmin (ferroxidase); DBI: diazepam binding inhibitor (GABA receptor modulator, acyl-Coenzyme A binding protein); EIF2S1 : eukaryotic translation initiation factor 2, subunit 1 alpha, 35kDa; ERBB2: v-erb-b2 erythroblastic leukemia viral oncogene homolog 2; EZR: ezrin (villin 2); FAS: TNF receptor superfamily, member 6; FBLN1 : fibulin 1 ; FGA: fibrinogen alpha chain; FGB: fibrinogen beta chain; FLNA: filamin A, alpha; GC: group-specific component (vitamin D binding protein); hCG: chorionic gonadotropin; ITGAV: integrin, alpha V; ITGB1 : integrin, beta 1 ; LDL: low density lipoprotein; MARCKS: myristoylated alanine-rich protein kinase C substrate; PLAU: plasminogen activator, urokinase; PPP1 R15A: protein phosphatase 1 , regulatory (inhibitor) subunit 15A; SCARB1 : scavenger receptor class B, member 1 ; SFRP4: secreted
frizzled-related protein 4; SHBG: sex hormone-binding globulin; SMAD3: SMAD family member 3; SMAD7: SMAD family member 7; SREBF1 : sterol regulatory element binding transcription factor 1 ; TGFA: transforming growth factor, alpha; TGFBR1 : transforming growth factor, beta receptor 1 ; TNF: tumor necrosis factor; VCAM1 : vascular cell adhesion molecule 1 ; VEGFA: vascular endothelial growth factor A.
[0067] FIG. 4 depicts interaction network for EmCa subtype biomarker candidates. This network was constructed using the proteins from Table 8. Overexpressed proteins are shown in red; underexpressed proteins are shown in green. See Table 8 for more details. Uncolored proteins were added by Ingenuity Pathway Analysis to connect the submitted biomarker candidates. ALDH2:
aldehyde dehydrogenase 2 family (mitochondrial); APOA1 : apolipoprotein A-l; BRD2: bromodomain containing 2; C3: complement component 3; C3AR1 : complement component 3a receptor 1 ; CASP1 : caspase 1 , apoptosis-related cysteine peptidase (interleukin 1 , beta, convertase); Caspase 3/7: Caspase group; CDKN2A: cyclin-dependent kinase inhibitor 2A (melanoma, p16, inhibits CDK4); CR2: complement component (3d/Epstein Barr virus) receptor 2; EIF4E: eukaryotic translation initiation factor 4E; FCER2: Fc fragment of IgE, low affinity I I, receptor for (CD23); HDGF: hepatoma- derived growth factor (high-mobility group protein 1-like); HNRNPA1 : heterogeneous nuclear ribonucleoprotein A1 ; HNRNPA2B1 : heterogeneous nuclear ribonucleoprotein A2/B1 ; HNRNPC: heterogeneous nuclear ribonucleoprotein C (C1/C2); HSPE1 : heat shock 10kDa protein 1 (chaperonin 10); IFNG: interferon, gamma; IgG: immunoglobulin gamma; IGHE: immunoglobulin heavy constant epsilon; IL4: interleukin 4; IL6: interleukin 6 (interferon, beta 2); ITGAX: integrin, alpha X (complement component 3 receptor 4 subunit); ITGB1 BP1 : integrin beta 1 binding protein 1 ; LDHA: lactate dehydrogenase A; MYC: v-myc myelocytomatosis viral oncogene homolog (avian); NFkB (complex): ; NME2: non-metastatic cells 2, protein (NM23B) expressed in; NPM 1 (includes EG:4869):
nucleophosmin (nucleolar phosphoprotein B23, numatrin); ORM 1 : orosomucoid 1 ; PCBP1 (includes EG:5093): poly(rC) binding protein 1 ; PLA2G2A: phospholipase A2, group I IA (platelets, synovial fluid); PLTP: phospholipid transfer protein; PPA1 : pyrophosphatase (inorganic) 1 ; PRDX5:
peroxiredoxin 5; RARA: retinoic acid receptor, alpha; SERPINA1 : serpin peptidase inhibitor, clade A (alpha-1 antitrypsin), member 1 ; SMPD2: sphingomyelin phosphodiesterase 2, neutral membrane ; SOD2: superoxide dismutase 2, mitochondrial; SPHK1 : sphingosine kinase 1 ; TMSB4X: thymosin beta 4, X-linked; TNF: tumor necrosis factor (TNF superfamily, member 2); TP53: tumor protein p53; VCAM 1 : vascular cell adhesion molecule 1 ; VVDR1 : WD repeat domain 1 ; YBX2: Y box binding protein 2.
[0068] FIG. 5 presents interaction network #5 for EmCa biomarker candidates. This network was constructed using the proteins from Table 9. Overexpressed proteins are shown in red;
underexpressed proteins are shown in green. Uncolored proteins were added by Ingenuity Pathway Analysis to connect the submitted biomarker candidates. ALB: albumin; CDKN1 A: cyclin-dependent kinase inhibitor 1 A (p21 , Cip1 ); CR1 : complement component (3b/4b) receptor 1 (Knops blood group); EN01 : enolase 1 , (alpha); EZR: ezrin; GH1 : growth hormone 1 ; GPI: glucose phosphate isomerase;
Histone h4; IFNB1 : interferon, beta 1 , fibroblast; IFNGR1 : interferon gamma receptor 1 ; IFNGR2: interferon gamma receptor 2 (interferon gamma transducer 1 ); IgG: Immunoglobulin gamma; ITGAM: integrin, alpha M (complement component 3 receptor 3 subunit); LAMP2: lysosomal-associated membrane protein 2; LEP: leptin; LGALS1 : lectin, galactoside-binding, soluble, 1 ; LU M: lumican; MAPK9: mitogen-activated protein kinase 9; Mek: mitogen-associated extracellular signal-regulated kinase; MYC: v-myc myelocytomatosis viral oncogene homolog (avian); NCL: nucleolin; NOS2: nitric oxide synthase 2, inducible; PGK1 : phosphoglycerate kinase 1 ; PIAS4: protein inhibitor of activated STAT, 4; PKM2: pyruvate kinase, muscle; PRDX1 : peroxiredoxin 1 ; RPS6: ribosomal protein S6; SERPINH 1 : serpin peptidase inhibitor, clade H (heat shock protein 47), member 1 ; SOCS3:
suppressor of cytokine signaling 3; SUM01 : SMT3 suppressor of mif two 3 homolog 1 (S. cerevisiae); TERC: telomerase RNA component; TF: transferrin; TFRC: transferrin receptor (p90, CD71 ); TKT: transketolase; TNF: tumor necrosis factor (TNF superfamily, member 2).
DETAILED DESCRIPTION OF THE INVENTION
[0069] The inventors previously performed a 40-sample iTRAQ study on EmCa that resulted in the identification of a number of differentially expressed proteins (12). In that study, tryptic peptides from tumor and normal endometrial tissues were compared after iTRAQ-labeling and two-dimensional LC with online nanoESI and MS/MS. In the present study, the inventors implement a more thorough analysis of their samples, to confirm and expand their earlier results.
[0070] The objectives of the present work were three-fold: first, to identify a larger number of proteins from the inventors' samples than previously achieved (12); second, to discover more proteins as cancer biomarkers than before; and third, to provide further verification of the inventors' previous results on an independent batch of cancer samples. As described herein, the inventors compared normal proliferative tissue samples and Type I EmCa tissue samples employing the same proteomic strategy as described in the earlier study (12). In contrast to the earlier study, however, the present comparison followed a "drill-down" approach in the proteomic analysis in which each sample was injected three times, and the acquisition software was programmed to exclude the peptides ions identified during previous iterations. Exclusion was restricted by means of a mass window and a time window, the latter to reduce the chance that peptide ions having m/z values similar to ones previously identified, but eluting at different times, would be excluded. The method described herein is considerably less involved than previous studies, as it did not incorporate the unidentified and expected ions with other charge states. The inventors also used the analysis of three different subtypes of EmCa to identify cancer biomarkers that can be used for classifying EmCa into subtypes and for general identification of EmCa.
[0071] As part of their efforts to improve the confidence of identifications, the inventors employed a new feature of the database search engine ProteinPilot to monitor the quality of the identifications. This new feature, the plug-in Proteomics System Performance Evaluation Pipeline (PSPEP), compares every identification to the chance of random matching in a reverse database and calculates
the individual false discovery rate (FDR) for each identified protein entry (13). This FDR value was used as validation of the ProteinPilot identification.
[0072] Accordingly, the inventors describe herein methods for detecting: the presence of an endometrial disease, including EmCa and similar conditions, in a sample; the absence of a disease in a sample; the stage or grade of a disease; and other characteristics of endometrial diseases that are relevant to prevention, diagnosis, characterization, monitoring, and therapy of endometrial diseases in a patient, including the metastatic potential and indolence or aggressiveness of endometrial cancer in a patient. Methods are also provided for: assessing the efficacy of one or more test agents for inhibiting a endometrial disease; assessing the efficacy of a therapy for endometrial diseases, such as EmCa; monitoring the progression of endometrial diseases, such as EmCa; selecting an agent or therapy for inhibiting endometrial diseases, such as EmCa; treating a patient afflicted with EmCa; inhibiting an endometrial disease, such as EmCa; and assessing the disease (e.g. , carcinogenic) potential of a test compound.
[0073] In one aspect, the invention provides marker sets that detect endometrial diseases and uses therefor. A marker set may include a plurality of polypeptides, or polynucleotides encoding such polypeptides, including or consisting of at least one marker listed in Table 5a, Table 5b, Table 8, and/or Table 9, and, optionally, 2, 3, 4, 5, 6, and 7 or up to all of the markers listed therein. In specific aspects, the markers consist of at least 2, 3, 4, or 5 polypeptides listed in Table 5a, Table 5b, Table 8, and/or Table 9 or polynucleotides encoding same. In a further aspect of the invention, the protein marker sets include or consist of protein clusters, or proteins in pathways, including markers listed in Table 5a, Table 5b, Table 8, and/or Table 9 or polynucleotides encoding same. In one aspect, the invention provides markers in Table 5a, Table 5b, Table 8, and/or Table 9 that are up-regulated or down-regulated or differentially expressed in cancer samples as compared to the non-cancer samples.
[0074] As defined and referred to herein, a "protein endometrial cancer marker" includes the up- regulated and down- regulated protein markers identified in Table 5a, Table 5b, Table 8, and/or Table 9 (up-regulated or down- regulated in cancer samples versus non-cancer samples), as well as native- sequence polypeptides, chimeric polypeptides, and all isoforms, homologs, fragments, and precursors thereof. Also included are modified forms of the proteins and derivatives thereof.
[0075] It is also contemplated that the invention includes, within its scope, polynucleotides encoding the protein EmCa markers. By way of example, such polynucleotides may be referred to herein, variously, as "polynucleotides encoding protein endometrial cancer markers" or
"polynucleotide endometrial cancer markers". The protein endometrial cancer markers and the polynucleotides encoding same are sometimes referred to herein, collectively, as "markers" or "endometrial cancer markers", as discussed in greater detail below.
[0076] Protein endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9 (in cancer sample versus non-cancer sample) and polynucleotides encoding the endometrial cancer markers may be used to determine the status of an endometrial cell or tissue and to detect an endometrial disease, such as endometrial cancer. Thus, the markers can be used for diagnosis, monitoring (including, without limitation, monitoring progression of disease state or success of therapeutic treatment), prognosis, treatment, or classification of an endometrial disease (including, without limitation, cancer, EmCa, and related conditions). The markers can also be used to evaluate disease states before surgery or after relapse. The invention also contemplates methods for assessing the status of an endometrial tissue, and methods for the diagnosis and therapy of an endometrial disease.
[0077] In accordance with another aspect of the invention, endometrial cancer can be assessed or characterized, for example, by detecting the presence in a sample of: (a) an endometrial cancer marker or fragment thereof; (b) a metabolite which is produced directly or indirectly by an endometrial cancer marker; (c) a transcribed nucleic acid or fragment thereof having at least a portion with which a polynucleotide endometrial cancer marker is substantially identical; and/or (d) a transcribed nucleic acid or fragment thereof, wherein the nucleic acid hybridizes with a polynucleotide marker encoding a protein endometrial cancer marker.
[0078] The levels of endometrial cancer markers may be determined by methods as described herein and generally known in the art. The expression levels of protein endometrial cancer markers may be determined by isolating and determining the level of nucleic acid transcribed from each polynucleotide endometrial cancer marker. Alternatively or additionally, the levels of protein endometrial cancer markers translated from mRNA transcribed from an endometrial cancer polynucleotide marker may be determined. In an aspect, the invention provides a method for characterizing or classifying an endometrial sample by detecting a difference in the expression of a first plurality of endometrial cancer markers relative to a control, the first plurality of such markers including at least 2, 3, 4, or 5 of the endometrial cancer markers corresponding to the markers listed in Table 5a, Table 5b, Table 8, and/or Table 9, and optionally 2, 3, 4, 5, 6, and 7, or up to all of the markers listed therein or those listed in Table 5a, Table 5b, Table 8, and/or Table 9 or those that are up-regulated in cancer versus control tissue as indicated in Table 5a, Table 5b, Table 8, and/or Table 9.
[0079] In an aspect of the invention, a method is provided for characterizing endometrial tissue by detecting endometrial cancer markers associated with the endometrial tissue stage or phase, or endometrial disease in a subject by: (a) obtaining a sample from a subject; (b) detecting or identifying in the sample an amount of endometrial cancer markers; and (c) comparing the detected amount with an amount detected for a standard.
[0080] In an embodiment of the invention, a method is provided for detecting protein endometrial cancer markers or polynucleotide endometrial cancer markers associated with endometrial cancer in
a patient by: (a) obtaining a sample from a patient; (b) detecting in the sample amount of protein endometrial cancer markers or polynucleotide endometrial cancer markers; and (c) comparing the detected amount with an amount detected for a standard.
[0081] The term "detect" or "detecting" includes assaying, imaging or otherwise establishing the presence or absence of the target endometrial cancer markers or polynucleotides encoding the markers, subunits thereof, or combinations of reagent bound targets, and the like, or assaying for, imaging, ascertaining, establishing, or otherwise determining one or more factual characteristics of an endometrial tissue phase or endometrial disease including cancer, metastasis, stage, or similar conditions. The term encompasses diagnostic, prognostic, and monitoring applications for the endometrial cancer markers and polynucleotides encoding the markers.
[0082] The invention also provides a method of assessing whether a patient is afflicted with or has a pre-disposition for endometrial disease, in particular endometrial cancer, by comparing: (a) levels of endometrial cancer markers associated with the endometrial disease in a sample from the patient; and (b) normal levels of endometrial cancer markers associated with the endometrial disease in samples of the same type obtained from control patients not afflicted with the disease, wherein altered levels of the endometrial cancer markers relative to the corresponding normal levels of such markers is an indication that the patient is afflicted with endometrial disease.
[0083] In one embodiment, the invention also provides a method of assessing whether a patient is afflicted with or has a pre-disposition for endometrial disease, in particular endometrial cancer, the method by comparing: (a) levels of protein endometrial cancer markers and/or the polynucleotide endometrial cancer markers associated with the endometrial disease in a sample from the patient; and (b) normal levels of endometrial cancer markers and/or the polynucleotide endometrial cancer markers associated with the endometrial disease in samples of the same type obtained from control patients not afflicted with the disease, wherein altered levels of the endometrial cancer markers and/or the polynucleotide endometrial cancer markers relative to the corresponding normal levels of such markers is an indication that the patient is afflicted with endometrial disease.
[0084] A further embodiment of the invention provides a method for assessing whether a patient is afflicted with or has a pre-disposition for endometrial cancer, where higher levels of protein endometrial cancer markers (e.g. , a heterogeneous nuclear ribonucleoprotein (e.g. , A2/B
(hnRNPA2/B1 )), L-lactate dehydrogenase A (LDHA), anterior gradient protein2 homolog, nucleophosmin, cystatin B, hepatoma derived growth factor and/or calcyphosin) in a sample, relative to the corresponding normal levels, indicate that the patient is afflicted with endometrial cancer.
[0085] Another embodiment of the invention provides a method for assessing whether a patient is afflicted with or has a pre-disposition for endometrial cancer, where lower levels of protein endometrial cancer markers (e.g. , fibrinogen (e.g. , alpha, beta, and/or gamma chain), apolipoprotein
A1 , haptoglobin (HP), and/or serpin H1 (or heat-shock protein 47 kD (hsp47)) in a sample, relative to the corresponding normal levels, indicate that the patient is afflicted with endometrial cancer.
[0086] In a further aspect, the invention provides a method for screening a subject for endometrial disease by: (a) obtaining a biological sample from a subject; (b) detecting the amount of endometrial cancer markers associated with the disease in the sample; and (c) comparing the amount of endometrial cancer markers detected to a predetermined standard, where detection of a level of endometrial cancer markers that differs significantly from the standard indicates endometrial disease.
[0087] In an embodiment, a significant difference between the levels of endometrial cancer marker levels in a patient and normal levels is an indication that the patient is afflicted with or has a predisposition to endometrial disease. In a further embodiment, the level of endometrial cancer markers is significantly higher compared to the standard and indicative of endometrial disease. In another embodiment, the level of endometrial cancer markers is significantly lower compared to the standard and indicative of endometrial disease. In another embodiment, the endometrial cancer markers detected are protein endometrial cancer markers or the polynucleotide endometrial cancer markers.
[0088] In a particular embodiment of the invention, the amount of protein endometrial cancer marker(s) (e.g. , a heterogeneous nuclear ribonucleoprotein (e.g. , A2/B (hnRNPA2/B1 )), L-lactate dehydrogenase A (LDHA), anterior gradient protein2 homolog, nucleophosmin, cystatin B, hepatoma derived growth factor and/or calcyphosin) detected is greater than that of a standard and is indicative of endometrial disease, including endometrial cancer. In another embodiment, the amount of endometrial cancer marker(s) (e.g. , fibrinogen (e.g. , alpha, beta, and/or gamma chain), apolipoprotein A1 , haptoglobin (HP), and/or serpin H 1 (or heat-shock protein 47 kD (hsp47)) detected is lower than that of a standard and is indicative of endometrial disease, including endometrial cancer.
[0089] In aspects of the methods of the invention, the methods are non-invasive for detecting endometrial disease which in turn allow for diagnosis of a variety of conditions or diseases associated with the endometrium.
[0090] In particular, the invention provides a non-invasive non-surgical method for detection, diagnosis or prediction of endometrial disease, including, without limitation, endometrial cancer and EmCa, in a subject, by: (a) obtaining a sample of body fluids, including, without limitation, blood, plasma, serum, urine or saliva, and/or a tissue sample from the subject; (b) subjecting the sample to a procedure to detect endometrial cancer markers in the body fluids and/or the tissue sample;
(c) detecting, diagnosing, and predicting endometrial disease by comparing the levels of endometrial cancer markers to the levels of markers obtained from a control subject with no endometrial disease.
[0091] In an embodiment, endometrial disease is detected, diagnosed, or predicted by determination of increased levels of protein endometrial cancer markers, including, without limitation,
one or more of the protein endometrial cancer markers indicated to be up-regulated in Table 5a, Table 5b, Table 8, and/or Table 9, when compared to such levels obtained from the control.
[0092] In another embodiment, endometrial disease is detected, diagnosed, or predicted by determination of decreased levels of protein endometrial cancer markers, including, without limitation, one or more of the protein endometrial caner marked indicated to be down-regulated in Table 5a, Table 5b, Table 8, and/or Table 9, when compared to such levels obtained from the control.
[0093] Another aspect of the invention provides for a method for assessing the aggressiveness or indolence of an endometrial disease in particular cancer (e.g. , staging), the method by comparing: (a) levels of endometrial cancer markers with the endometrial disease in a patient sample; and (b) normal levels of the endometrial cancer markers in a control sample.
[0094] In an embodiment, a significant difference between the levels in the sample and the normal levels is an indication that the endometrial disease, in particular cancer, is aggressive or indolent. In a particular embodiment, the levels of endometrial cancer markers are higher than normal levels. In another particular embodiment, the levels of endometrial cancer markers are lower than normal levels. In another embodiment, the endometrial cancer markers are protein endometrial cancer markers or polynucleotide endometrial cancer markers.
[0095] In an embodiment, aggressiveness or indolence of an endometrial disease assessed by determination of increased levels of protein endometrial cancer markers, including, without limitation, one or more of the protein endometrial cancer markers indicated to be up-regulated in Table 5a, Table 5b, Table 8, and/or Table 9, when compared to such levels obtained from the control.
[0096] In another embodiment, aggressiveness or indolence of an endometrial disease assessed by determination of decreased levels of protein endometrial cancer markers, including, without limitation, one or more of the protein endometrial caner marked indicated to be down- regulated in Table 5a, Table 5b, Table 8, and/or Table 9, when compared to such levels obtained from the control.
[0097] In an embodiment, the invention provides a method for diagnosing and/or monitoring EmCa by comparing: (a) levels of LDHA or a polynucleotide encoding LDHA in a sample from the patient; and (b) normal levels of LDHA or a polynucleotide encoding the same in samples of the same type obtained from control patients not afflicted with endometrial cancer or having a different stage of endometrial cancer, wherein altered levels of LHDA or polynucleotides encoding same compared with the corresponding normal levels is an indication that the patient is afflicted with EmCa. In other embodiments, the method compares patient-sample levels and normal levels of phosphoglycerate kinase, apolipoprotein A1 , fibrinogen (e.g. , alpha, beta, and/or gamma chain), serpin H1 (or heat- shock protein 47 kD (hsp47), a heterogeneous nuclear ribonucleoprotein (e.g. , A2/B (hnRNPA2/B1 )), or haptoglobin (HP)), or a polynucleotide encoding same. Exemplary EmCa which may be diagnosed
and/or monitored by these methods include, without limitation, endometrioid EmCa, serous EmCa, or clear cell EmCa.
[0098] In an aspect, the invention provides a method for determining whether a cancer has metastasized or is likely to metastasize in the future, by comparing: (b) levels of endometrial cancer markers in a patient sample; and (b) normal levels (or non-metastatic levels) of the endometrial cancer markers in a control sample. In one embodiment, a significant difference between the levels in the patient sample and the normal levels is an indication that the cancer has metastasized or is likely to metastasize in the future.
[0099] In another aspect, the invention provides a method for monitoring the progression of endometrial disease, in particular endometrial cancer in a patient, by: (a) detecting endometrial cancer markers or polynucleotides encoding the markers associated with the disease in a sample from the patient at a first time point; (b) repeating step (a) at a subsequent point in time; and (c) comparing the levels detected in (a) and (b), and therefrom monitoring the progression of the endometrial disease.
[00100] The invention contemplates a method for determining the effect of an environmental factor on the endometrial tissue, or endometrial disease, by comparing endometrial cancer endometrial cancer markers in the presence and absence of the environmental factor. One embodiment of such method uses protein endometrial cancer marker or polynucleotide endometrial cancer markers.
[00101] The invention also provides a method for assessing the efficacy of a test agent for inhibiting endometrial disease, and a method of selecting an agent for inhibiting endometrial disease. In another aspect, the invention provides for a method for assessing the carcinogenic potential of a test compound.
[00102] The invention further contemplates a method of assessing the potential of a test compound to contribute to an endometrial disease by: (a) maintaining separate aliquots of endometrial diseased cells in the presence and absence of the test compound; and (b) comparing the levels of endometrial cancer markers associated with the disease in each of the aliquots.
[00103] A significant difference between the levels of endometrial cancer markers or
polynucleotides encoding the markers in an aliquot maintained in the presence of (or exposed to) the test compound relative to the aliquot maintained in the absence of the test compound, indicates that the test compound contributes to endometrial disease. In one embodiment, the endometrial cancer markers are protein endometrial cancer markers or polynucleotide endometrial cancer markers.
[00104] The invention further relates to a method of assessing the efficacy of a therapy for inhibiting endometrial disease in a patient by comparing: (a) levels of endometrial cancer markers associated with disease in a first sample from the patient obtained from the patient prior to providing
at least a portion of the therapy to the patient; and (b) levels of endometrial cancer markers associated with disease in a second sample obtained from the patient following therapy.
[00105] In an embodiment, a significant difference between the levels of endometrial cancer markers in the second sample relative to the first sample is an indication that the therapy is efficacious for inhibiting endometrial disease. In another embodiment, the endometrial cancer markers are protein endometrial cancer markers or polynucleotide endometrial cancer markers.
[00106] In a particular embodiment, the method is used to assess the efficacy of a therapy for inhibiting endometrial disease, including, without limitation, endometrial cancer or EmCa, where lower levels of endometrial cancer markers in the second sample relative to the first sample, is an indication that the therapy is efficacious for inhibiting the disease. In another embodiment, the endometrial cancer markers are protein endometrial cancer markers or polynucleotide endometrial cancer markers.
[00107] The "therapy" may be any therapy for treating endometrial disease, in particular endometrial cancer, including but not limited to therapeutics, radiation, immunotherapy, gene therapy, and surgical removal of tissue. Therefore, the method can be used to evaluate a patient before, during, and after therapy.
[00108] Certain methods of the invention employ binding agents that specifically recognize endometrial cancer markers.
[00109] In another aspect, the invention provides methods for determining the presence or absence of endometrial disease, in particular endometrial cancer, in a patient, by: (a) contacting a biological sample obtained from a patient with one or more binding agent that specifically binds to one or more endometrial cancer markers associated with the disease; and (b) detecting in the sample an amount of marker that binds to the binding agent, relative to a predetermined standard or cut-off value, and therefrom determine the presence or absence of endometrial disease in the patient from such results.
[00110] In a further aspect, the invention relates to a method for diagnosing and monitoring an endometrial disease, in particular endometrial cancer, in a subject by quantifying one or more endometrial cancer markers associated with the disease in a biological sample from the subject, by: (a) reacting the biological sample with one or more binding agent specific for the endometrial cancer markers that are directly or indirectly labelled with a detectable substance; and (b) detecting the detectable substance. In one embodiment, antibodies are used as binding agents to recognize the protein endometrial cancer markers. In a further embodiment, polynucleotides are used as binding agents to recognize the polynucleotide endometrial cancer markers.
[00111] In another aspect the invention provides a method of using an antibody to detect expression of one or more protein endometrial cancer marker in a sample, by: (a) combining
antibodies specific for one or more protein endometrial cancer marker with a sample under conditions which allow the formation of antibody-protein marker complexes; and (b) detecting complex formation, wherein complex formation indicates expression of the protein endometrial cancer marker in the sample wherein expression may be compared with standards and is diagnostic of an endometrial disease, in particular endometrial cancer.
[00112] Embodiments of the invention may also involve: (a) reacting a biological sample from a subject with antibodies specific for one or more endometrial cancer markers which are directly or indirectly labelled with an enzyme; (b) adding a substrate for the enzyme wherein the substrate is selected so that the substrate, or a reaction product of the enzyme and substrate forms fluorescent complexes; (c) quantifying one or more endometrial cancer markers in the sample by measuring fluorescence of the fluorescent complexes; and (d) comparing the quantified levels to levels obtained for other samples from the subject patient, or control subjects.
[00113] In another embodiment the quantified levels are compared to levels quantified for control subjects, such as normal or benign tumours, without an endometrial disease, particularly endometrial cancer, wherein an increase in endometrial cancer marker levels compared with the control subjects is indicative of endometrial disease.
[00114] In a further embodiment the quantified levels are compared to levels quantified for control subjects, such as normal or benign tumours, without an endometrial disease, particularly endometrial cancer, wherein a decrease in endometrial marker levels compared with the control subjects is indicative of endometrial disease.
[00115] In another aspect, the invention provides a method by: (a) incubating a biological sample with first antibodies specific for one or more endometrial cancer markers which are directly or indirectly labelled with a detectable substance, and second antibodies specific for one or more endometrial cancer markers which are immobilized; (b) detecting the detectable substance thereby quantifying endometrial cancer markers in the biological sample; and (c) comparing the quantified endometrial cancer markers with levels for a pre-determined standard.
[00116] The standard may correspond to levels quantified for samples from control subjects without endometrial cancer (normal or benign), with a different disease stage, or from other samples of the subject. In an embodiment, increased levels of endometrial cancer markers as compared to the standard may be indicative of endometrial cancer. In another embodiment, lower levels of endometrial cancer markers as compared to a standard may be indicative of endometrial cancer.
[00117] Protein endometrial cancer marker levels can be determined by constructing an antibody microarray in which binding sites include immobilized, preferably monoclonal, antibodies specific to a substantial fraction of marker-derived protein endometrial cancer markers of interest.
[00118] Other methods of the invention employ one or more polynucleotides capable of hybridizing to one or more polynucleotide endometrial cancer markers. Thus, methods can be used to monitor an endometrial disease, in particular endometrial cancer, by detecting polynucleotide endometrial cancer markers associated with the disease.
[00119] Thus, the invention relates to a method for diagnosing and monitoring an endometrial disease, including, without limitation, endometrial cancer, EmCa or related condition, in a sample from a subject, by: (a) isolating nucleic acids, preferably mRNA, from the sample; and (b) detecting polynucleotide endometrial cancer markers associated with endometrial diseases, wherein the presence of different levels of polynucleotide endometrial cancer markers in the sample compared to a standard or control may be indicative of endometrial disease, disease stage, and/or a negative or positive prognosis, such as longer progression-free and overall survival.
[00120] In embodiments of the invention, polynucleotide endometrial cancer marker positive tumours (e.g., higher levels of the polynucleotides compared to a control normal or benign sample) are a negative diagnostic indicator. Positive tumors can be indicative of endometrial cancer, advanced stage disease, lower progression-free survival, and/or overall survival.
[00121] In embodiments of the invention, polynucleotide endometrial cancer marker negative tumours (e.g., lower levels of the polynucleotides compared to a control normal or benign sample) are a negative diagnostic indicator. Negative tumors can be indicative of endometrial cancer, advanced stage disease, lower progression-free survival, and/or overall survival.
[00122] The invention provides methods for determining the presence or absence of an endometrial disease in a subject, by: (a) detecting in the sample levels of nucleic acids that hybridize to one or more polynucleotide endometrial cancer markers associated with the disease; (b) comparing the levels with a pre-determined standard or cut-off value; and (c) determining the presence or absence of endometrial disease in the subject.
[00123] In an embodiment, the invention provides methods for determining the presence or absence of endometrial disease, such as EmCa or another endometrial cancer, in a subject, by: (a) contacting a sample obtained from the subject with oligonucleotides that hybridize to
polynucleotides encoding endometrial cancer markers; (b) detecting in the sample a level of nucleic acids that hybridize to the polynucleotides relative to a predetermined cut-off value; and (c) determining the presence or absence of endometrial cancer in the subject.
[00124] Within certain embodiments, the amount of polynucleotides that are mRNA are detected via polymerase chain reaction using, for example, oligonucleotide primers that hybridize to one or more polynucleotide endometrial cancer markers or complements of such polynucleotides. Within other embodiments, the amount of mRNA is detected using a hybridization technique, employing
oligonucleotide probes that hybridize to one or more polynucleotide endometrial cancer markers or complements of such polynucleotides.
[00125] When using mRNA detection, the method may be carried out by combining isolated mRNA with reagents to convert to cDNA according to standard methods well known in the art, treating the converted cDNA with amplification reaction reagents (such as cDNA PCR reaction reagents) in a container along with an appropriate mixture of nucleic acid primers; reacting the contents of the container to produce amplification products; and analyzing the amplification products to detect the presence of one or more of the polynucleotide endometrial cancer markers in the sample. For mRNA, the analyzing step may be accomplished using Northern Blot analysis to detect the presence of polynucleotide endometrial cancer markers in the sample. The analysis step may be further accomplished by quantitatively detecting the presence of polynucleotide endometrial cancer markers in the amplification product, and comparing the quantity of marker detected against a panel of expected values for the known presence or absence of such markers in normal and malignant tissue derived using similar primers.
[00126] Therefore, the invention provides a method wherein mRNA is detected by: (a) isolating mRNA from a sample and combining the mRNA with reagents to convert it to cDNA; (b) treating the converted cDNA with amplification reaction reagents and nucleic acid primers that hybridize to one or more of the polynucleotide endometrial cancer markers endometrial cancer marker to produce amplification products; (c) analyzing the amplification products for determining the amount of mRNA present encoding the protein endometrial cancer marker; and (d) comparing the determined amount of mRNA to an amount detected against a panel of expected values for normal and diseased tissue (e.g. , malignant tissue) derived using similar methods.
[00127] In particular embodiments of the invention, RT-PCR can be used to amplify the mRNA for protein endometrial cancer markers for detection and analysis. Other embodiments of the invention use quantitative RT-PCR to quantitatively determine amount of mRNA for protein endometrial cancer markers. Further embodiments of the invention use real time RT-PCR for quantification and analysis.
[00128] In particular embodiments of the invention, the methods described herein utilize the polynucleotide endometrial cancer markers placed on a microarray so that the expression status of each of the markers may be assessed.
[00129] In a particular aspect, the invention provides a microarray including a defined set of genes encoding protein endometrial cancer markers (i.e. , at least 2, 3, 4, or 5 genes listed in Table 5a, Table 5b, Table 8, and/or Table 9) whose expression is significantly altered by endometrial disease. The invention further relates to the use of the microarray as a prognostic tool to predict or diagnose endometrial disease. In an embodiment, the endometrial microarray discriminates between endometrial diseases resulting from different etiologies.
[00130] In an embodiment, the invention provides for oligonucleotide arrays including marker sets described herein. The microarrays provided by the present invention may include probes to markers able to distinguish endometrial disease. In particular, the invention provides oligonucleotide arrays including probes to a subset or subsets of at least 5 to 10 polynucleotide endometrial cancer markers, up to a full set of markers which distinguish endometrial disease.
[00131] The invention also contemplates a method by administering to cells or tissues imaging agents that carry labels for imaging and bind to endometrial cancer markers and optionally other markers of an endometrial disease, and then imaging the cells or tissues.
[00132] In an aspect the invention provides an in vivo method by administering to a subject an agent that has been constructed to target one or more endometrial cancer markers.
[00133] In a particular embodiment, the invention contemplates an in vivo method by
administering to a mammal one or more agent that carries a label for imaging and binds to one or more endometrial cancer markers, and then imaging the mammal.
[00134] In another aspect, the invention provides an in vivo method for imaging endometrial cancer, by: (a) injecting a patient with an agent that binds to one or more endometrial cancer markers, the agent carrying a label for imaging the endometrial cancer; (b) allowing the agent to incubate in vivo and bind to one or more endometrial cancer markers associated with the endometrial cancer; and (c) detecting the presence of the label localized to the cancer. In one embodiment, the agent is an antibody that recognizes a protein endometrial cancer marker. In another embodiment, the agent is a chemical entity that recognizes an endometrial cancer marker.
[00135] An agent carries a label to image an endometrial marker and optionally other markers. Examples of labels useful for imaging are radiolabels, fluorescent labels (e.g. , fluorescein and rhodamine), nuclear magnetic resonance active labels, positron emitting isotopes detectable by a positron emission tomography ("PET") scanner, chemiluminescers such as luciferin, and enzymatic markers such as peroxidase or phosphatase. Short-range radiation emitters, such as isotopes detectable by short-range detector probes can also be employed.
[00136] The invention also contemplates the localization or imaging methods described herein using multiple markers for an endometrial disease, including, without limitation, endometrial cancer, EmCa or related conditions.
[00137] The invention also relates to kits for carrying out the methods of the invention. In an embodiment, a kit is for assessing whether a patient is afflicted with an endometrial disease, including, without limitation, endometrial cancer, EmCa or related conditions, and the kit includes reagents for assessing one or more endometrial cancer markers.
[00138] The invention further provides kits including marker sets described herein. In an aspect of the invention, the kit contains a microarray ready for hybridization to target endometrial cancer polynucleotide markers and the software needed for the data analysis.
[00139] The invention also provides a diagnostic composition including protein endometrial cancer markers or polynucleotide encoding the markers. A composition is also provided including a probe that specifically hybridizes to polynucleotide endometrial cancer markers, or a fragment thereof, or an antibody specific for protein endometrial cancer markers or a fragment thereof. In another aspect, a composition is provided including polynucleotide endometrial cancer markers specific primer pairs capable of amplifying such polynucleotides using polymerase chain reaction methodologies. The probes, primers or antibodies can be labelled with a detectable substance.
[00140] Still further the invention relates to therapeutic applications for endometrial diseases, in particular endometrial cancer, employing protein endometrial cancer markers, and polynucleotide encoding the markers, and/or binding agents for such markers.
[00141] In an aspect, the invention relates to compositions including markers or parts thereof associated with an endometrial disease, or antibodies specific for protein endometrial cancer markers associated with an endometrial disease, and a pharmaceutically acceptable carrier, excipient, or diluent. Another of the invention provides a method for treating or preventing an endometrial disease, in particular endometrial cancer, in a patient, by administering to a patient in need thereof, markers or parts thereof associated with endometrial disease, antibodies specific for protein endometrial cancer markers, or a composition of the invention. In an aspect the invention provides a method of treating a patient afflicted with or at risk of developing an endometrial disease (e.g. , endometrial cancer) by inhibiting expression of polynucleotide endometrial cancer markers.
[00142] An embodiment of the invention includes antisense oligonucleotides complementary to polynucleotide endometrial cancer markers delivered to diseased cells for regulation of gene expression.
[00143] In an aspect, the invention provides antibodies specific for protein endometrial cancer markers associated with a disease, such as EmCa, that can be used therapeutically to destroy or inhibit the disease, such as growth of marker expressing cancer cells), or to block marker activity associated with a disease. In an aspect, the endometrial cancer markers may be used in various immunotherapeutic methods to promote immune-mediated destruction or growth inhibition of tumors expressing the endometrial cancer markers.
[00144] The invention also contemplates a method of using endometrial cancer markers or parts thereof, or antibodies specific for the protein endometrial cancer markers in the preparation or manufacture of a medicament for the prevention or treatment of an endometrial disease, including, without limitation, endometrial cancer, EmCa or related conditions.
[00145] Another aspect of the invention relates to the use of protein endometrial cancer markers, peptides derived therefrom, or chemically produced (synthetic) peptides, or any combination of these molecules, for use in the preparation of vaccines to prevent an endometrial disease and/or to treat an endometrial disease.
[00146] The invention contemplates vaccines for stimulating or enhancing in a subject to whom the vaccine is administered, production of antibodies directed against one or more endometrial cancer markers.
[00147] The invention also provides a method for stimulating or enhancing in a subject production of antibodies directed against one or more endometrial cancer markers by administering to the subject a vaccine of the invention in a dose effective for stimulating or enhancing production of the antibodies.
[00148] The invention further provides a method for treating, preventing, or delaying recurrence of an endometrial disease, including, without limitation, endometrial cancer, EmCa or related conditions by administering to the subject a vaccine of the invention in a dose effective for treating, preventing, or delaying recurrence of an endometrial disease including, without limitation, endometrial cancer, EmCa or related conditions.
[00149] The invention contemplates the methods, compositions, and kits described herein using additional markers associated with an endometrial disease, including, without limitation, endometrial cancer, EmCa or related conditions. The methods described herein may be modified by including reagents to detect the additional markers, or polynucleotides for the markers.
[00150] In particular, the invention contemplates the methods described herein using multiple markers for endometrial cancer, such as EmCa. Therefore, the invention contemplates a method for analyzing a biological sample for the presence of endometrial cancer markers and other markers that are specific indicators of cancer, in particular endometrial cancer. The methods described herein may be modified by including reagents to detect the additional markers, or nucleic acids for the additional markers.
[00151] In embodiments of the invention, the methods, compositions, and kits use a panel of the protein endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9 or polynucleotides encoding same.
[00152] In embodiments of the invention, biological samples may be obtained from tissues, extracts, cell cultures, cell lysates, lavage fluid, or physiological fluids. In further embodiments of the invention, the sample is an endometrial tumour tissue.
[00153] Abbreviation Index. For convenience, certain abbreviations used in the description, tables, figures, and appended claims are defined here: iTRAQ, isobaric tags for relative and absolute quantification; LC, liquid chromatography; MS / MS, tandem mass spectrometry; PCM, potential
cancer marker; EmCa, endometrial carcinoma; LCM, laser capture microdissection; PBS, phosphate- buffered saline; SCX, strong cation exchange; ID, internal diameter; RP, reverse phase; IDA, information-dependent acquisition; TBS, tris-buffered saline; AUC, area under the curve; RSD, relative standard deviation; TMA, tissue microarray; AAT, alpha-antitrypsin (serpin A1 ); AFP, alpha fetoprotein; APO A-1 , apolipoprotein A-l; ATP5A1 , ATP synthase subunit alpha, mitochondrial;
ATP5B, ATP synthase subunit beta, mitochondrial; EmCa, endometrial carcinoma; FDR, false discovery rate; FBPA A, fructose-bisphosphate aldolase A; FIGO, Federation Internationale de Gynecologie et d'Obstetrique or International Federation of Gynecology and Obstetrics; HCC, hepatocellular carcinoma; hnRNPA2B1 , heterogeneous nuclear ribonucleoproteins A2/B1 (splice variants of A1 ); HP, haptoglobin; HRPC, hormone-refractory prostate cancer; HSPC, hormone- sensitive prostate cancer; IPA, Ingenuity Pathways Analysis; LAP3, leucine aminopeptidase 3; LDHA, L-lactate dehydrogenase A; OvCa, Ovarian carcinoma; PGK1 , phosphoglycerate kinase 1 ; PK, pyruvate kinase isoenzymes M1/M2; PSPEP, Proteomics System Performance Evaluation Pipeline (ProteinPilot® software plug-in); S100-A1 1 , Calgizzarin.
[00154] Glossary. For convenience, certain terms employed in the specification, examples, and appended claims are collected here.
[00155] The recitation of numerical ranges by endpoints herein includes all numbers and fractions subsumed within that range (e.g. , 1 to 5 includes 1 , 1.5, 2, 2.75, 3, 3.90, 4, and 5). It is also to be understood that all numbers and fractions thereof are presumed to be modified by the term "about." Furthermore, it is to be understood that "a", "an, " and "the" include plural referents unless the content clearly dictates otherwise. Thus, for example, reference to a composition or method that includes "an endometrial cancer marker" will encompass two or more endometrial cancer markers. The term "about" means plus or minus 0.1 to 50%, 5-50%, or 10-40%, preferably 10-20%, more preferably 10% or 15%, of the number to which reference is being made.
[00156] "Endometrial disease" refers to any disorder, disease, condition, syndrome, or combination of manifestations or symptoms recognized or diagnosed as a disorder of the endometrium, including, without limitation, inflammation, cancer precursors, and carcinoma.
[00157] "Endometrial disease" includes malignant endometrial cancer, including, without limitation, endometrial carcinomas (EmCa) such as Type I EmCa, endometrial serous carcinoma, and endometrial clear cell carcinoma.
[00158] The terms "sample", "biological sample", and the like mean a material known or suspected of expressing or containing one or more endometrial carcinoma polynucleotide or polypeptide markers. A test sample can be used directly as obtained from the source or following a pretreatment to modify the character of the sample. The sample can be derived from any biological source, such as tissues, extracts, or cell cultures, including cells (e.g. , tumor cells), cell lysates, and physiological fluids, such as, for example, whole blood, plasma, serum, saliva, ocular lens fluid,
cerebral spinal fluid, sweat, urine, milk, ascites fluid, synovial fluid, peritoneal fluid, lavage fluid, and the like. The sample can be obtained from animals, preferably mammals, most preferably humans. The sample can be treated prior to use, such as preparing plasma from blood, diluting viscous fluids, and the like. Methods of treatment can involve freezing, fixation, embedding in paraffin or OCT, sonication, centrifugation, filtration, distillation, extraction, concentration, inactivation of interfering components, the addition of reagents, and the like.
[00159] In embodiments of the invention, the sample is a mammalian tissue sample. In particular embodiments, the tissue is endometrial tissue. In further embodiments, the sample is an endometrial tumour tissue.
[00160] In another embodiment the sample is a human physiological fluid. In a particular embodiment, the sample is human serum.
[00161] The samples that may be analyzed in accordance with the invention include
polynucleotides from clinically relevant sources, preferably expressed RNA or a nucleic acid derived therefrom (cDNA or amplified RNA derived from cDNA that incorporates an RNA polymerase promoter). The target polynucleotides can include RNA, including, without limitation, total cellular RNA, miRNA, poly(A)+ messenger RNA (mRNA) or fraction thereof, cytoplasmic mRNA, or RNA transcribed from cDNA (i.e. , cRNA; see, for example. , Linsley & Schelter, Patent Application Ser. No. 09 / 411 ,074, or Patent Nos. 5,545, 522; 5,891 ,636; or 5,716,785). Methods for preparing total and poly(A)+ RNA are well known in the art, and are described generally, for example, in Sambrook et al. , (1989, Molecular Cloning - A Laboratory Manual (2nd Ed.), Vols. 1 -3, Cold Spring Harbor Laboratory, Cold Spring Harbor, N.Y. ) and Ausubel et al. , eds. (1994, Current Protocols in Molecular Biology, Vol. 2, Current Protocols Publishing, New York). RNA may be isolated from eukaryotic cells by procedures involving lysis of the cells and denaturation of the proteins contained in the cells.
Additional steps may be utilized to remove DNA. Cell lysis may be achieved with a non-ionic detergent, followed by microcentrifugation to remove the nuclei and hence the bulk of the cellular DNA. (See Chirgwin et al. , 1979, Biochemistry 18:5294-5299). Poly(A)+RNA can be selected using oligo-dT cellulose (see Sambrook et al. , 1989, Molecular Cloning - A Laboratory Manual (2nd Ed. ), Vols. 1 -3, Cold Spring Harbor Laboratory, Cold Spring Harbor, N.Y. ). In the alternative, RNA can be separated from DNA by organic extraction, for example, with hot phenol or phenol / chloroform / isoamyl alcohol.
[00162] It may be desirable to enrich mRNA with respect to other cellular RNAs, such as transfer RNA (tRNA) and ribosomal RNA (rRNA). Most mRNAs contain a poly(A) tail at their 3' end allowing them to be enriched by affinity chromatography, for example, using oligo(dT) or poly(U) coupled to a solid support, such as cellulose or Sephadex™ (see Ausubel et al. , eds. , 1994, Current Protocols in Molecular Biology, Vol. 2, Current Protocols Publishing, New York). Bound poly(A)+mRNA is eluted from the affinity column using 2 mM EDTA / 0.1 % SDS.
[00163] A sample of RNA can include a plurality of different RNA molecules each with a different nucleotide sequence. In an aspect of the invention, the RNA molecules contain mRNA molecules. In a further aspect of the invention, the mRNA molecules in the RNA sample include at least 100 different nucleotide sequences.
[00164] Target polynucleotides can be detectably labelled at one or more nucleotides using methods known in the art. The label is preferably uniformly incorporated along the length of the RNA, and more preferably, is carried out at a high degree of efficiency. The detectable label can be a luminescent label, fluorescent label, bio-luminescent label, chemiluminescent label, radiolabel, and colorimetric label. In a particular embodiment, the label is a fluorescent label, such as a fluorescein, a phosphor, a rhodamine, or a polymethine dye derivative. Commercially available fluorescent labels include, for example, fluorescent phosphoramidites, such as FluorePrime (Amersham Pharmacia, Piscataway, N.J. ), Fluoredite (Millipore, Bedford, Mass. ), FAM (ABI, Foster City, Calif. ), and Cy3 or Cy5 (Amersham Pharmacia, Piscataway, N.J. ).
[00165] Target polynucleotides from a patient sample can be labelled differentially from polynucleotides of a standard. The standard can include target polynucleotides from normal individuals (i.e. , those not afflicted with or pre-disposed to endometrial disease), in particular pooled from samples from normal individuals. The target polynucleotides can be derived from the same individual, but taken at different time points, and thus indicate the efficacy of a treatment by a change in expression of the markers, or lack thereof, during and after the course of treatment.
[00166] The terms "subject", "individual", and "patient" refer to a warm-blooded animal such as a mammal. In particular, the terms refer to a human. A subject, individual or patient may be afflicted with or suspected of having or being pre-disposed to endometrial disease or a condition as described herein. The terms also includes domestic animals bred for food or as pets, including horses, cows, sheep, poultry, fish, pigs, cats, dogs, and zoo animals.
[00167] Methods herein for administering an agent or composition to subjects / individuals / patients contemplate treatment as well as prophylactic use. Typical subjects for treatment include persons susceptible to, suffering from or that have suffered a condition or disease described herein. In particular, suitable subjects for treatment in accordance with the invention are persons that are susceptible to, suffering from or that have suffered endometrial cancer.
[00168] The expression "endometrial cancer marker" refers to a marker associated with normal or diseased endometrial tissue and includes one or more of polypeptides that are differentially expressed in cancer samples as compared to normal samples and polynucleotides encoding these polypeptides. The expression includes native-sequence polypeptides and nucleotides, isoforms, chimeric polypeptides, complexes, and all homologs, fragments, precursors, modified forms, and derivatives of the markers, as applicable. An endometrial cancer marker may be associated with an endometrial disease, including endometrial cancer (e.g. , EmCa). The expression "endometrial cancer marker"
includes a marker associated with protein endometrial cancer marker listed in Table 5a, Table 5b, Table 8, and/or Table 9, polynucleotides encoding the proteins listed in Table 5a, Table 5b, Table 8, and/or Table 9, and precursors thereof.
[00169] A "native-sequence polypeptide" includes a polypeptide having the same amino acid sequence of a polypeptide derived from nature. Such native-sequence polypeptides can be isolated from nature or can be produced by recombinant or synthetic means. The term specifically encompasses naturally occurring truncated or secreted forms of a polypeptide, polypeptide variants including naturally occurring variant forms (e.g. , alternatively spliced forms or splice variants), and naturally occurring allelic variants.
[00170] The expression "polypeptide variant" refers to a polypeptide having at least about 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, 97%, 98%, or 99% amino acid sequence identity, particularly at least about 70-80%, more particularly at least about 85%, still more particularly at least about 90%, most particularly at least about 95% amino acid sequence identity with a native- sequence polypeptide. Particular polypeptide variants have at least 70-80%, 85%, 90%, 95% amino acid sequence identity to the sequences identified in Table 5a, Table 5b, Table 8, and/or Table 9. Such variants include, for instance, polypeptides wherein one or more amino acid residues are added to, or deleted from, the N- or C-terminus of the full-length or mature sequences of the polypeptide, including variants from other species, but exclude a native-sequence polypeptide. In aspects of the invention variants retain the immunogenic activity of the corresponding native-sequence polypeptide.
[00171] Percent identity of two amino acid sequences, or of two nucleic acid sequences is defined as the percentage of amino acid residues or nucleotides in a candidate sequence that are identical with the amino acid residues in a polypeptide or nucleic acid sequence, after aligning the sequences and introducing gaps, if necessary, to achieve the maximum percent sequence identity, and not considering any conservative substitutions as part of the sequence identity. Alignment for purposes of determining percent amino acid or nucleic acid sequence identity can be achieved in various conventional ways, for instance, using publicly available computer software including the GIG program package (Devereux J. et al. , Nucleic Acids Research 12(1 ): 387, 1984); BLASTP, BLASTN, and FASTA (Altschul, S. F. et al. , J. Molec. Biol. 215: 403-410, 1990). The BLAST X program is publicly available from NCBI and other sources (BLAST Manual, Altschul, S. et al. , NCBI NLM NIH Bethesda, Md. 20894; Altschul, S. et al. , J. Mol. Biol. 215: 403-410, 1990). Skilled artisans can determine appropriate parameters for measuring alignment, including any algorithms needed to achieve maximal alignment over the full length of the sequences being compared. Methods to determine identity and similarity are codified in publicly-available computer programs.
[00172] An allelic variant may also be created by introducing substitutions, additions, or deletions into a polynucleotide encoding a native polypeptide sequence such that one or more amino acid substitutions, additions, or deletions are introduced into the encoded protein. Mutations may be introduced by standard methods, such as site-directed mutagenesis and PCR-mediated mutagenesis.
In an embodiment, conservative substitutions are made at one or more predicted non-essential amino acid residues. A "conservative amino acid substitution" is one in which an amino acid residue is replaced with an amino acid residue with a similar side chain. Amino acids with similar side chains are known in the art and include amino acids with basic side chains (e.g., Lys, Arg, His), acidic side chains (e.g. , Asp, Glu), uncharged polar side chains (e.g. , Gly, Asp, Glu, Ser, Thr, Tyr, and Cys), non- polar side chains (e.g. , Ala, Val, Leu, Iso, Pro, Trp), beta-branched side chains (e.g. , Thr, Val, Iso), and aromatic side chains (e.g. , Tyr, Phe, Trp, His). Mutations can also be introduced randomly along part or all of the native sequence, for example, by saturation mutagenesis. Following mutagenesis the variant polypeptide can be recombinantly expressed and the activity of the polypeptide may be determined.
[00173] Polypeptide variants include polypeptides and amino acid sequences sufficiently identical to or derived from the amino acid sequence of a native polypeptide which includes fewer amino acids than the full length polypeptides. A portion of a polypeptide can be a polypeptide which is, for example, 10, 15, 20, 25, 30, 35, 40, 45, 50, 60, 70, 80, 90, 100 or more amino acids in length.
Portions in which regions of a polypeptide are deleted can be prepared by recombinant techniques and can be evaluated for one or more functional activities such as the ability to form antibodies specific for a polypeptide.
[00174] A naturally occurring allelic variant may contain conservative amino acid substitutions from the native polypeptide sequence or it may contain a substitution of an amino acid from a corresponding position in a polypeptide homolog, for example, a murine polypeptide.
[00175] An endometrial cancer marker may be part of a chimeric or fusion protein. A "chimeric protein" or "fusion protein" includes all or part (preferably biologically active) of an endometrial cancer marker operably linked to a heterologous polypeptide (i.e. , a polypeptide other than an endometrial diseases marker). Within the fusion protein, the term "operably linked" is intended to indicate that an endometrial cancer marker and the heterologous polypeptide are fused in-frame to each other. The heterologous polypeptide can be fused to the N-terminus or C-terminus of an endometrial cancer marker. A useful fusion protein is a GST fusion protein in which an endometrial cancer marker is fused to the C-terminus of GST sequences. Another example of a fusion protein is an
immunoglobulin fusion protein in which all or part of an endometrial cancer marker is fused to sequences derived from a member of the immunoglobulin protein family. Chimeric and fusion proteins can be produced by standard recombinant DNA techniques.
[00176] A modified form of a polypeptide referenced herein includes modified forms of the polypeptides and derivatives of the polypeptides, including post-translationally modified forms such as glycosylated, phosphorylated, acetylated, methylated or lapidated forms of the polypeptides. For example, an N-terminal methionine may be cleaved from a polypeptide, and a new N-terminal residue may or may not be acetylated. In particular, for chaperonin 10 the first residue, methionine, can be cleaved and the second first residue, alanine can be N-acetylated.
[00177] An endometrial cancer marker may be prepared by recombinant or synthetic methods, or isolated from a variety of sources, or by any combination of these and similar techniques.
[00178] As used herein, "polynucleotides encoding the protein markers" include polynucleotides that encode the protein endometrial cancer markers described herein, polynucelotides that encode a native-sequence polypeptide, a polypeptide variant, including a portion of a polypeptide, a chimeric polypeptide, or an isoform, precursor, complex, modified form, or derivative of any of the polypeptide endometrial cancer markers, and any precursors thereof. By way of example, a polynucleotide endometrial cancer marker includes or consists of one or more polynucleotides encoding the polypeptides listed in Table 5a, Table 5b, Table 8, and/or Table 9.
[00179] Polynucleotide endometrial cancer markers include complementary nucleic acid sequences, and nucleic acids that are substantially identical to these sequences (e.g. , having at least about 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, 97%, 98%, or 99% sequence identity). Polynucleotide endometrial cancer markers also include sequences that differ from a native sequence due to degeneracy in the genetic code. As one example, DNA sequence polymorphisms within the nucleotide sequence of an endometrial cancer marker may result in silent mutations that do not affect the amino acid or nucleotide sequence. Variations in one or more nucleotides may exist among individuals within a population due to natural allelic variation. DNA sequence polymorphisms may also occur which lead to changes in the amino acid sequence of a polypeptide or the nucleotide sequence of a polynucleotide.
[00180] Polynucleotide endometrial cancer markers also include nucleic acids that hybridize under stringent conditions, preferably high stringency conditions to endometrial cancer polynucleotide markers. Appropriate stringency conditions which promote DNA hybridization are known to those skilled in the art, or can be found in Current Protocols in Molecular Biology, John Wiley & Sons, N.Y. (1989), 6.3.1 -6.3.6. For example, 6.0 x sodium chloride / sodium citrate (SSC) at about 45°C, followed by a wash of 2.0 x SSC at 50°C may be employed. The stringency may be selected based on the conditions used in the wash step. By way of example, the salt concentration in the wash step can be selected from a high stringency of about 0.2 x SSC at 50°C. In addition, the temperature in the wash step can be at high stringency conditions, at about 65°C.
[00181] Polynucleotide endometrial cancer markers further include truncated nucleic acids or nucleic acid fragments and variant forms of the nucleic acids that arise by alternative splicing of an mRNA corresponding to a DNA.
[00182] The polynucleotide endometrial cancer markers are intended to include DNA and RNA, including, without limitation, mRNA, and can be either double stranded or single stranded. A polynucleotide may, but need not, include additional coding or non-coding sequences, or it may, but need not, be linked to other molecules and/or carrier or support materials. The polynucleotides for use in the methods of the invention may be of any length suitable for a particular method. In certain
applications the term also includes antisense polynucleotides (e.g., RNA or DNA strand with complementary sequences in the reverse orientation to the polynucleotide endometrial cancer markers).
[00183] "Statistically different levels", "significantly altered levels", or "significant difference" in levels of markers in a patient sample compared to a control or standard (e.g. , normal levels or levels in other samples from a patient) may represent levels that are higher or lower than the standard error of the detection assay. In particular embodiments, the levels may be 1.5, 2, 3, 4, 5, 6 or more times higher or lower than the control or standard. In other embodiments of the invention, the levels may be significantly higher compared to the control or standard. In another embodiment, the levels may be significantly lower compared to the control or standard.
[00184] "Microarray" and "array" refer to nucleic acid or nucleotide arrays or protein or peptide arrays that can be used to detect biomolecules associated with endometrial cancer cell or tissue phase and endometrial disease, for instance to measure gene expression. A variety of arrays are made in research and manufacturing facilities worldwide, some of which are available commercially. By way of example, spotted arrays and in situ synthesized arrays are two kinds of nucleic acid arrays that differ in the manner in which the nucleic acid materials are placed onto the array substrate. A widely used in situ synthesized oligonucleotide array is GeneChip™ made by Affymetrix, Inc.
Oligonucleotide probes that are 20- or 25-base long can be synthesized in silico on the array substrate. These arrays can achieve high densities (e.g., more than 40,000 genes per cm2).
Generally spotted arrays have lower densities, but the probes, typically partial cDNA molecules, are much longer than 20- or 25-mers. Examples of spotted cDNA arrays include LifeArray made by Incyte Genomics and DermArray made by IntegriDerm (or Invitrogen). Pre-synthesized and amplified cDNA sequences are attached to the substrate of spotted arrays. Protein and peptide arrays also are known (see for example, Zhu et al. , Science 293:2101 (2001 )).
[00185] "Binding agent" refers to a substance such as a polypeptide or antibody that specifically binds to one or more endometrial cancer markers. A substance "specifically binds" to one or more protein endometrial cancer markers if it reacts at a detectable level with one or more endometrial cancer markers, and does not react detectably with peptides containing an unrelated or different sequence. Binding properties may be assessed using an ELISA, which may be readily performed by those skilled in the art (see, for example, Newton et al. , Develop. Dynamics 197: 1 -13, 1993).
[00186] A binding agent may be a ribosome, with or without a peptide component, an aptamer, an RNA molecule, or a polypeptide. A binding agent may be a polypeptide that includes one or more endometrial cancer marker sequence, a peptide variant thereof, or a non-peptide mimetic of such a sequence. By way of example, an annexin A1 sequence may be a peptide portion of an annexin A1 that is capable of modulating a function mediated by annexin A1.
[00187] An aptamer includes a DNA or RNA molecule that binds to nucleic acids and proteins. An aptamer that binds to a protein (or binding domain) of an endometrial cancer marker or a polynucleotide endometrial cancer marker can be produced using conventional techniques, without undue experimentation. For example, see the following publications describing in vitro selection of aptamers: Klug et al. , Mol. Biol. Reports 20:97-107 (1994); Wallis et al. , Chem. Biol. 2:543-552 (1995); Ellington, Curr. Biol. 4:427-429 (1994); Lato et al. , Chem. Biol. 2:291 -303 (1995); Conrad et al. , Mol. Div. 1 :69-78 (1995); and Uphoff et al. , Curr. Opin. Struct. Biol. 6:281 -287 (1996).
[00188] Antibodies for use in the present invention include but are not limited to monoclonal or polyclonal antibodies, immunologically active fragments (e.g. , a Fab or (Fab)2 fragments), antibody heavy chains, humanized antibodies, antibody light chains, genetically engineered single chain Fv molecules (Ladner et al. , US Patent No. 4,946,778), chimeric antibodies, for example, antibodies which contain the binding specificity of murine antibodies, but in which the remaining portions are of human origin, or derivatives, such as enzyme conjugates or labelled derivatives.
[00189] Antibodies, including monoclonal and polyclonal antibodies, fragments and chimeras, may be prepared using methods known to those skilled in the art. Isolated native or recombinant endometrial cancer markers may be utilized to prepare antibodies. (See, for example, Kohler et al. (1975), Nature 256:495-497; Kozbor et al. (1985), J. Immunol Methods 81 :31 -42; Cote et al. (1983), Proc Natl Acad Sci 80:2026-2030; and Cole et al. (1984), Mol Cell Biol 62: 109-120 for the preparation of monoclonal antibodies; Huse et al. (1989), Science 246: 1275-1281 for the preparation of monoclonal Fab fragments; and, Pound (1998), Immunochemical Protocols, Humana Press, Totowa, NJ for the preparation of phagemid or B-lymphocyte immunoglobulin libraries to identify antibodies). Antibodies specific for an endometrial cancer marker may also be obtained from scientific or commercial sources.
[00190] In an embodiment of the invention, antibodies are reactive against a protein endometrial cancer marker if they bind with a Ka of greater than or equal to 10-7 M.
[00191] Markers. The invention provides a set of markers correlated with endometrial disease. In an aspect, the invention provides a set of markers identified as useful for detection, diagnosis, prevention and therapy of endometrial disease including or consisting of one or more of the markers listed in Table 5a, Table 5b, Table 8, and/or Table 9. In another aspect, the invention provides the endometrial cancer markers in Table 5a, Table 5b, Table 8, and/or Table 9 for detection, diagnosis and prognosis of an endometrial disease. The invention also provides a method of using endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9 to distinguish endometrial disease.
[00192] In an embodiment, the markers include or consist of 1 , 2, 3, 4 or more other markers listed in Table 5a, Table 5b, Table 8, and/or Table 9 or polynucleotides encoding same.
[00193] In embodiments of the invention, a marker is provided which is selected from the group consisting of the polypeptides set forth in Table 5a, Table 5b, Table 8, and/or Table 9 which polypeptides are up-regulated biomarkers in endometrial. In embodiments of the invention, a marker is provided which is selected from the group consisting of at least one marker of Table 5a, Table 5b, Table 8, and/or Table 9.
[00194] In embodiments of the invention, a marker is provided which is selected from the group consisting of at least one marker of Table 5a, Table 5b, Table 8, and/or Table 9 and at least 2, 3, 4, 5, 6, 7, 8, 9, or 10 polypeptides set forth in Table 5a, Table 5b, Table 8, and/or Table 9.
[00195] The invention provides marker sets that distinguish endometrial disease and uses thereof. In an aspect, the invention provides a method for classifying an endometrial disease including detecting a difference in the expression of a first plurality of endometrial cancer markers relative to a control, the first plurality of endometrial cancer markers including or consisting of at least 2, 3, 4, or 5 of the markers listed in Table 5a, Table 5b, Table 8, and/or Table 9 or polynucleotides encoding same. In specific aspects, a control includes markers derived from a sample from a patient with no endometrial disease.
[00196] Any of the markers provided herein may be used alone or with other markers of endometrial disease, or with markers for other phenotypes or conditions. Additionally, all of the sequences provided herein are representative only; there may be other sequences for particular protein or coding sequences or related sequences. The invention is not intended to be limited to the sequences herein provided.
[00197] Detection Methods. A variety of methods can be employed for the diagnostic and prognostic evaluation of endometrial cancer status involving one or more endometrial cancer markers and the polynucleotides encoding the markers, and the identification of subjects with a predisposition to endometrial diseases or that are receptive to in vitro fertilization and embryo transfer procedures. Such methods may, for example, involve endometrial cancer polynucleotide markers, and fragments thereof, and binding agents (e.g. , antibodies) against one or more endometrial cancer markers, including peptide fragments. In particular, the polynucleotides and antibodies may be used, for example, for: (1 ) the detection of the presence of polynucleotide endometrial cancer marker mutations, or the detection of either over- or under-expression of protein endometrial cancer marker mRNA relative to a non-disorder state or different endometrial cancer cell or tissue phase, or the qualitative or quantitative detection of alternatively spliced forms of polynucleotide endometrial cancer marker transcripts which may correlate with certain conditions or susceptibility toward such conditions; and (2) the detection of either an over- or an under-abundance of one or more endometrial cancer markers relative to a non-disorder state or a different endometrial cell or tissue phase or the presence of a modified (e.g. , less than full length) endometrial cancer which correlates with a disorder state or a progression toward a disorder state.
[00198] Recent advances in MS techniques enable proteins from different samples to be compared via labelled tags differing in isotopic composition. Samples are combined and processed in a single batch, allowing performance of comparative quantification (15). Effective labeling strategies include isotope-coded affinity tag (ICAT) ( 16), or the more recent variation that uses isobaric tagging reagent, iTRAQ (17), followed by multidimensional LC-MS/MS analysis. These approaches have been applied successfully to identify new tumor markers for endometrial cancer, pre-malignant lesions, and squamous cell carcinoma of head and neck (8, 9).
[00199] The invention also contemplates a method for detecting an endometrial disease, in particular an endometrial cancer, including producing a profile of levels of one or more endometrial cancer marker associated with an endometrial disease and/or polynucleotides encoding the markers, and other markers associated with endometrial disease in cells from a patient, and comparing the profile with a reference to identify a profile for the test cells indicative of disease. In an aspect, the endometrial cancer markers are one or more of those listed in Table 5a, Table 5b, Table 8, and/or Table 9 or a precursor thereof.
[00200] The methods described herein may be used to evaluate the probability of the presence of malignant cells, for example, in a group of cells freshly removed from a host. Such methods can be used to detect tumors, quantify their growth, and help in the diagnosis and prognosis of endometrial disease. The methods can be used to detect the presence of cancer metastasis, as well as confirm the absence or removal of all tumor tissue following surgery, cancer chemotherapy, and/or radiation therapy. They can further be used to monitor cancer chemotherapy and tumor reappearance.
[00201] The methods described herein can be adapted for diagnosing and monitoring endometrial tissue status or an endometrial disease by detecting one or more endometrial cancer markers or polynucleotides encoding the markers in biological samples from a subject. These applications require that the amount of markers or polynucleotides quantified in a sample from a subject being tested be compared to a predetermined standard or cut-off value. The standard may correspond to levels quantified for another sample or an earlier sample from the subject, or levels quantified for a control sample. Levels for control samples from healthy subjects, different endometrial tissue phases, or subjects with an endometrial disease may be established by prospective and/or retrospective statistical studies. Healthy subjects who have no clinically evident disease or abnormalities may be selected for statistical studies. Diagnosis may be made by a finding of statistically different levels of detected endometrial cancer markers associated with disease or polynucleotides encoding same, compared to a control sample or previous levels quantified for the same subject.
[00202] The methods described herein may also use multiple markers for an endometrial disease, in particular endometrial cancer, EmCa, or similar conditions. Therefore, the invention contemplates a method for analyzing a biological sample for the presence of one or more endometrial cancer markers and polynucleotides encoding the markers, and other markers that are specific indicators of an
endometrial disease. The methods described herein may be modified by including reagents to detect the additional markers, or polynucleotides for the markers.
[00203] Nucleic Acid Methods / Assays. As noted herein an endometrial disease or phase may be detected based on the level of endometrial cancer polynucleotide markers in a sample.
Techniques for detecting polynucleotides such as polymerase chain reaction (PCR) and hybridization assays are well known in the art.
[00204] Probes may be used in hybridization techniques to detect endometrial cancer polynucleotide markers. The technique generally involves contacting and incubating nucleic acids (e.g. , recombinant DNA molecules, cloned genes) obtained from a sample from a patient or other cellular source with a probe under conditions favorable for the specific annealing of the probes to complementary sequences in the nucleic acids. After incubation, the non-annealed nucleic acids are removed, and the presence of nucleic acids that have hybridized to the probe if any are detected.
[00205] Nucleotide probes for use in the detection of nucleic acid sequences in samples may be constructed using conventional methods known in the art. Suitable probes may be based on nucleic acid sequences encoding at least 5 sequential amino acids from regions of a polynucleotide endometrial cancer marker, preferably they include 10-200, more particularly 10-30, 10-40, 20-50, 40- 80, 50-150, 80-120 nucleotides in length.
[00206] The probes may include DNA or DNA mimics (e.g. , derivatives and analogues) corresponding to a portion of an organism's genome, or complementary RNA or RNA mimics. Mimics are polymers including subunits capable of specific, Watson-Crick-like hybridization with DNA, or of specific hybridization with RNA. The nucleic acids can be modified at the base moiety, at the sugar moiety, or at the phosphate backbone.
[00207] DNA can be obtained using standard methods such as polymerase chain reaction (PCR) amplification of genomic DNA or cloned sequences. (See, for example, in Innis et al. , eds., 1990, PCR Protocols: A Guide to Methods and Applications, Academic Press Inc. , San Diego, CA).
Computer programs known in the art can be used to design primers with the required specificity and optimal amplification properties, such as Oligo version 5.0 (National Biosciences). Controlled robotic systems may be useful for isolating and amplifying nucleic acids.
[00208] A nucleotide probe may be labelled with a detectable substance such as a radioactive label that provides for an adequate signal and has sufficient half-life such as 32P, 3H, 14C or the like. Other detectable substances that may be used include antigens that are recognized by a specific labelled antibody, fluorescent compounds, enzymes, antibodies specific for a labelled antigen, and luminescent compounds. An appropriate label may be selected having regard to the rate of hybridization and binding of the probe to the nucleotide to be detected and the amount of nucleotide available for hybridization. Labelled probes may be hybridized to nucleic acids on solid supports such
as nitrocellulose filters or nylon membranes as generally described in Sambrook et al. , 1989, Molecular Cloning, A Laboratory Manual (2nd ed. ). The nucleic acid probes may be used to detect endometrial cancer polynucleotide markers, preferably in human cells. The nucleotide probes may also be useful in the diagnosis of an endometrial disease involving one or more endometrial cancer polynucleotide markers, in monitoring the progression of such disorder, or monitoring a therapeutic treatment.
[00209] The detection of polynucleotide encoding the markers may involve the amplification of specific gene sequences using an amplification method such as polymerase chain reaction (PCR), followed by the analysis of the amplified molecules using techniques known to those skilled in the art. Suitable primers can be routinely designed by one of skill in the art.
[00210] By way of example, at least two oligonucleotide primers may be employed in a PCR based assay to amplify a portion of a polynucleotide encoding one or more endometrial disease marker derived from a sample, wherein at least one of the oligonucleotide primers is specific for (i.e. , hybridizes to) a polynucleotide encoding the endometrial disease marker. The amplified cDNA is then separated and detected using techniques well known in the art, such as gel electrophoresis.
[00211] In order to maximize hybridization under assay conditions, primers and probes employed in the methods of the invention generally have at least about 60%, preferably at least about 75%, and more preferably at least about 90% identity to a portion of a polynucleotide encoding an endometrial disease marker; that is, they are at least 10 nucleotides, and preferably at least 20 nucleotides in length. In an embodiment the primers and probes are at least about 10-40 nucleotides in length.
[00212] Hybridization and amplification techniques described herein may be used to assay qualitative and quantitative aspects of polynucleotide endometrial cancer marker expression. For example, RNA may be isolated from a cell type or tissue known to express an endometrial cancer polynucleotide marker and tested utilizing the hybridization (e.g. , standard Northern analyses) or PCR techniques referred to herein.
[00213] The primers and probes may be used in the above-described methods in situ (i.e. , directly on tissue sections (fixed and/or frozen) of patient tissue obtained from biopsies or resections).
[00214] In an aspect of the invention, a method is provided employing reverse transcriptase- polymerase chain reaction (RT-PCR), in which PCR is applied in combination with reverse transcription. Generally, RNA is extracted from a sample tissue using standard techniques (e.g. , guanidine isothiocyanate extraction as described by Chomcynski and Sacchi, Anal. Biochem.
162: 156-159, 1987) and is reverse transcribed to produce cDNA. The cDNA is used as a template for a polymerase chain reaction. The cDNA is hybridized to a set of primers, at least one of which is specifically designed against an endometrial disease marker sequence. Once the primer and template have annealed a DNA polymerase is employed to extend from the primer, to synthesize a
copy of the template. The DNA strands are denatured, and the procedure is repeated many times until sufficient DNA is generated to allow visualization by ethidium bromide staining and agarose gel electrophoresis. Real time RT-PCR and qRT-PCR may also be used.
[00215] Amplification may be performed on samples obtained from a subject with a suspected endometrial disease and an individual who is not afflicted with an endometrial disease. The reaction may be performed on several dilutions of cDNA spanning at least two orders of magnitude. A statistically significant difference in expression in several dilutions of the subject sample as compared to the same dilutions of the non-disease sample may be considered positive for the presence of an endometrial disease.
[00216] In an embodiment, the invention provides methods for determining the presence or absence of an endometrial disease in a subject including (a) contacting a sample obtained from the subject with oligonucleotides that hybridize polynucleotide encoding the endometrial cancer markers; and (b) detecting in the sample a level of nucleic acids that hybridize to the polynucleotides relative to a predetermined cut-off value, and therefrom determining the presence or absence of an endometrial disease in the subject. In an aspect, the endometrial disease is cancer and endometrial cancer markers are one or more of those listed in Table 5a, Table 5b, Table 8, and/or Table 9 or a precursor thereof.
[00217] The invention provides a method wherein mRNA for a protein endometrial cancer marker is detected by: (a) isolating RNA from a sample and combining the RNA with reagents to convert it to cDNA; (b) treating the converted cDNA with amplification reaction reagents and nucleic acid primers that hybridize to one or more mRNA encoding protein endometrial disease markers, to produce amplification products; (d) analyzing the amplification products to detect amounts of mRNA encoding protein endometrial disease markers; and (e) comparing the amount of such RNA to an amount detected against a panel of expected values for normal and malignant tissue derived using similar nucleic acid primers.
[00218] Endometrial cancer marker-positive samples or alternatively higher levels in patients compared to a control (e.g. , non-cancerous tissue) may be indicative of late stage disease, and/or that the patient is not responsive to chemotherapy. Alternatively, negative samples or lower levels compared to a control (e.g. , non-cancerous tissue or negative samples) may be indicative of progressive disease and shorter overall survival.
[00219] In another embodiment, the invention provides methods for determining the presence or absence of endometrial cancer in a subject by: (a) contacting a sample obtained from the subject with oligonucleotides that hybridize to one or more polynucleotide endometrial cancer markers; and (b) detecting in the sample levels of nucleic acids that hybridize to the polynucleotides relative to a predetermined cut-off value, and therefrom determining the presence or absence of endometrial
cancer in the subject. In an embodiment, the endometrial cancer polynucleotide markers encode one or more polypeptides listed in Table 5a, Table 5b, Table 8, and/or Table 9.
[00220] In particular, the invention provides a method wherein mRNA for L-lactate dehydrogenase A (LDHA), phosphoglycerate kinase, apolipoprotein A1 , fibrinogen (e.g. , alpha, beta, and/or gamma chain), serpin H 1 (or heat-shock protein 47 kD (hsp47), a heterogeneous nuclear ribonucleoprotein (e.g. , A2/B (hnRNPA2/B1 )), or haptoglobin (HP) is detected by: (a) isolating mRNA from a sample and combining the mRNA with reagents to convert it to cDNA; (b) treating the converted cDNA with amplification reaction reagents and nucleic acid primers that hybridize to a polynucleotide encoding (LDHA), phosphoglycerate kinase, apolipoprotein A1 , fibrinogen (e.g. , alpha, beta, and/or gamma chain), serpin H 1 (or heat-shock protein 47 kD (hsp47), a heterogeneous nuclear ribonucleoprotein (e.g. , A2/B (hnRNPA2/B1 )), or haptoglobin (HP), to produce amplification products; (c) analyzing the amplification products to detect an amount of the mRNA encoding (LDHA), phosphoglycerate kinase, apolipoprotein A1 , fibrinogen (e.g. , alpha, beta, and/or gamma chain), serpin H1 (or heat-shock protein 47 kD (hsp47), a heterogeneous nuclear ribonucleoprotein (e.g. , A2/B (hnRNPA2/B1 )), or haptoglobin (HP); and (d) comparing the amount of mRNA to an amount detected against a panel of expected values for normal and malignant tissue derived using similar nucleic acid primers.
[00221] Protein-positive samples or higher levels, especially significantly higher levels, of polynucleotides encoding a heterogeneous nuclear ribonucleoprotein (e.g. , A2/B (hnRNPA2/B1 )), L- lactate dehydrogenase A (LDHA), anterior gradient protein2 homolog, nucleophosmin, cystatin B, hepatoma derived growth factor, and/or calcyphosin in patients, as compared to a control (e.g. , normal or benign), are indicative of endometrial carcinoma. Negative samples or lower levels of polynucleotides encoding fibrinogen (e.g. , alpha, beta, and/or gamma chain), apolipoprotein A1 , haptoglobin (HP), and/or serpin H1 (or heat-shock protein 47 kD (hsp47) in patients, as compared to a control (e.g. , normal or benign), may also be indicative of progressive disease and poor overall survival.
[00222] Oligonucleotides or longer fragments derived from an endometrial cancer polynucleotide marker may be used as targets in a microarray. The microarray can be used to simultaneously monitor the expression levels of large numbers of genes and to identify genetic variants, mutations, and polymorphisms. The information from the microarray may be used to determine gene function, to understand the genetic basis of a disorder, to diagnose a disorder, and to develop and monitor the activities of therapeutic agents.
[00223] The preparation, use, and analysis of microarrays are well known to a person skilled in the art. (See, for example, Brennan, T. M. et al. (1995), US Patent No. 5,474,796; Schena, et al. (1996), Proc. Natl. Acad. Sci. 93: 10614-10619; Baldeschweiler et al. (1995), PCT Application WO 95 / 251 1 16; Shalon, D. et al. (1995), PCT application WO 95 / 35505; Heller, R.A. et al. (1997), Proc. Natl. Acad. Sci. , 94:2150-2155; and Heller, M.J. et al. (1997), US Patent No. 5,605,662).
[00224] Thus, the invention also includes an array including one or more endometrial cancer polynucleotide markers (in particular the markers listed in Table 5a, Table 5b, Table 8, and/or Table 9). The array can be used to assay expression of endometrial cancer polynucleotide markers in the array. The invention allows the quantification of expression of one or more endometrial cancer polynucleotide markers.
[00225] Microarrays typically contain at separate sites nanomolar quantities of individual genes, cDNAs, or ESTs on a substrate (e.g. , nitrocellulose or silicon plate), or photolithographically prepared glass substrate. The arrays are hybridized to cDNA probes using conventional techniques with gene- specific primer mixes. The target polynucleotides to be analyzed are isolated, amplified and labelled, typically with fluorescent labels, radiolabels or phosphorous label probes. After hybridization is completed, the array is inserted into the scanner, where patterns of hybridization are detected. Data are collected as light emitted from the labels incorporated into the target, which becomes bound to the probe array. Probes that completely match the target generally produce stronger signals than those that have mismatches. The sequence and position of each probe on the array are known, and thus by complementarity, the identity of the target nucleic acid applied to the probe array can be determined.
[00226] Microarrays are prepared by selecting polynucleotide probes and immobilizing them to a solid support or surface. The probes may include DNA sequences, RNA sequences, copolymer sequences of DNA and RNA, DNA and/or RNA analogues, or combinations thereof. The probe sequences may be full or partial fragments of genomic DNA, or they may be synthetic oligonucleotide sequences synthesized either enzymatically in vivo, enzymatically in vitro (e.g. , by PCR), or non- enzymatically in vitro.
[00227] The probe or probes used in the methods of the invention can be immobilized to a solid support or surface which may be either porous or non-porous. For example, the probes can be attached to a nitrocellulose or nylon membrane or filter covalently at either the 3' or the 5' end of the polynucleotide probe. The solid support may be a glass or plastic surface. In an aspect of the invention, hybridization levels are measured to microarrays of probes consisting of a solid support on the surface of which are immobilized a population of polynucleotides, such as a population of DNA or DNA mimics, or, alternatively, a population of RNA or RNA mimics. A solid support may be a nonporous or, optionally, a porous material such as a gel.
[00228] In accordance with embodiments of the invention, a microarray is provided including a support or surface with an ordered array of hybridization sites or "probes" each representing one of the markers described herein. The microarrays can be addressable arrays, and in particular positionally addressable arrays. Each probe of the array is typically located at a known, predetermined position on the solid support such that the identity of each probe can be determined from its position in the array. In preferred embodiments, each probe is covalently attached to the solid support at a single site.
[00229] Microarrays used in the present invention are preferably (a) reproducible, allowing multiple copies of a given array to be produced and easily compared with each other; (b) made from materials that are stable under hybridization conditions; (c) small (e.g. , between 1 cm2 and 25 cm2, between 12 cm2 and 13 cm2, or 3 cm2); and (d) include a unique set of binding sites that will specifically hybridize to the product of a single gene in a cell (e.g. , to a specific mRNA, miRNA, or to a specific cDNA derived therefrom). However, it will be appreciated that larger arrays may be used particularly in screening arrays, and other related or similar sequences will cross hybridize to a given binding site.
[00230] In accordance with an aspect of the invention, the microarray is an array in which each position represents one of the markers described herein (e.g. , the markers listed in Table 5a, Table 5b, Table 8, and/or Table 9). Each position of the array can include a DNA or DNA analogue based on genomic DNA to which a particular RNA or cDNA transcribed from a genetic marker can specifically hybridize. A DNA or DNA analogue can be a synthetic oligomer or a gene fragment. In an embodiment, probes representing each of the endometrial cancer markers and endometrial cancer polynucleotide markers are present on the array. In a preferred embodiment, the array includes at least 5 of the endometrial cancer polynucleotide markers.
[00231] Probes for the microarray can be synthesized using N-phosphonate or phosphoramidite chemistries (Froehler et al., 1986, Nucleic Acid Res. 14:5399-5407; McBride et al. , 1983, Tetrahedron Lett. 24:246-248). Synthetic sequences are typically between about 10 and about 500 bases, 20-100 bases, or 40-70 bases in length. Synthetic nucleic acid probes can include non-natural bases, such as, without limitation, inosine. Nucleic acid analogues such as peptide nucleic acid may be used as binding sites for hybridization (see, e.g. , Egholm et al. , 1993, Nature 363:566-568; US Patent No. 5,539,083).
[00232] Probes can be selected using an algorithm that takes into account binding energies, base composition, sequence complexity, cross-hybridization binding energies, and secondary structure (see Friend et al. , International Patent Publication WO 01 / 05935, published Jan. 25, 2001 ).
[00233] Positive control probes (e.g. , probes known to be complementary and hybridize to sequences in the target polynucleotides) and negative control probes (e.g. , probes known to not be complementary and hybridize to sequences in the target polynucleotides) are typically included on the array. Positive controls can be synthesized along the perimeter of the array or synthesized in diagonal stripes across the array. A reverse complement for each probe can be next to the position of the probe to serve as a negative control.
[00234] The probes can be attached to a solid support or surface, which may be made from glass, plastic (e.g. , polypropylene, nylon), polyacrylamide, nitrocellulose, gel, or other porous or nonporous material. The probes can be printed on surfaces such as glass plates (see Schena et al. , 1995, Science 270:467-470). This method may be particularly useful for preparing microarrays of cDNA.
(See, also, DeRisi et al. , 1996, Nature Genetics 14:457-460; Shalon et al. , 1996, Genome Res. 6:639- 645; and Schena et al. , 1995, Proc. Natl. Acad. Sci. U.S.A. 93: 10539-1 1286).
[00235] High-density oligonucleotide arrays containing thousands of oligonucleotides
complementary to defined sequences, at defined locations on a surface can be produced using photolithographic techniques for synthesis in situ (see Fodor et al. , 1991 , Science 251 :767-773; Pease et al. , 1994, Proc. Natl. Acad. Sci. U.S.A. 91 :5022-5026; Lockhart et al. , 1996, Nature
Biotechnology 14: 1675; US Patent Nos. 5,578,832; 5,556,752; and 5,510,270) or other methods for rapid synthesis and deposition of defined oligonucleotides (Blanchard et al. , Biosensors &
Bioelectronics 1 1 :687-690). Using these methods oligonucleotides (e.g. , 60-mers) of known sequence are synthesized directly on a surface such as a derivatized glass slide. The array produced may be redundant, with several oligonucleotide molecules per RNA.
[00236] Microarrays can be made by other methods including masking (Maskos and Southern, 1992, Nuc. Acids. Res. 20: 1679-1684). In an embodiment, microarrays of the present invention are produced by synthesizing polynucleotide probes on a support wherein the nucleotide probes are attached to the support covalently at either the 3' or the 5' end of the polynucleotide.
[00237] The invention provides microarrays including a disclosed marker set. In one embodiment, the invention provides a microarray for distinguishing endometrial disease samples including a positionally-addressable array of polynucleotide probes bound to a support, the polynucleotide probes including a plurality of polynucleotide probes of different nucleotide sequences, each of the different nucleotide sequences including a sequence complementary and hybridizable to a plurality of genes, the plurality consisting of at least 2, 3, 4, 5, or 6 of the genes corresponding to the markers listed in Table 5a, Table 5b, Table 8, and/or Table 9 and optionally at least 2-18, 5-16, 10-15, 13-21 , 2-21 , 2- 32, 22-32, or 13-32 of the genes corresponding to the markers listed in Table 5a, Table 5b, Table 8, and/or Table 9. An aspect of the invention provides microarrays including at least 4, 5, or 6 of the polynucleotides encoding the markers listed in Table 5a, Table 5b, Table 8, and/or Table 9.
[00238] The invention provides gene marker sets that distinguish endometrial disease and uses therefor. In an aspect, the invention provides a method for classifying an endometrial disease including detecting a difference in the expression of a first plurality of genes relative to a control, the first plurality of genes consisting of at least 3, 4, 5, or 6 of the genes encoding the markers listed in Table 5a, Table 5b, Table 8, and/or Table 9. In specific aspects, the plurality of genes consists of at least 4 or 5 of the genes encoding the markers listed in Table 5a, Table 5b, Table 8, and/or Table 9 and optionally at least 2-18, 5-16, 10-15, 13-21 , 2-21 , 2-32, 22-32, or 13-32 of the genes
corresponding to the markers listed in Table 2. In another specific aspect, the control includes nucleic acids derived from a pool of samples from individual control patients. An aspect of the invention provides microarrays including at least 4, 5, or 6 of the polynucleotides encoding the markers listed in Table 5a, Table 5b, Table 8, and/or Table 9.
[00239] The invention provides a method for classifying an endometrial disease by calculating the similarity between the expression in a sample of at least 3, 4, 5, or 6 of the endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9 and the expression of the same markers in a control pool. The method includes the steps of: (a) labelling nucleic acids derived from a sample, with a first fluorophore to obtain a first pool of fluorophore-labelled nucleic acids; (b) labelling with a second fluorophore a first pool of nucleic acids derived from two or more endometrial disease samples, and a second pool of nucleic acids derived from two or more control samples; (c) contacting the first fluorophore-labelled nucleic acid and the first pool of second fluorophore-labelled nucleic acid with a first microarray under conditions such that hybridization can occur, and contacting the first fluorophore-labelled nucleic acid and the second pool of second fluorophore-labelled nucleic acid with a second microarray under conditions such that hybridization can occur, detecting at each of a plurality of discrete loci on the first microarray a first fluorescent emission signal from the first fluorophore-labelled nucleic acid and a second fluorescent emission signal from the first pool of second fluorophore-labelled genetic matter that is bound to the first microarray and detecting at each of the marker loci on the second microarray the first fluorescent emission signal from the first fluorophore-labelled nucleic acid and a third fluorescent emission signal from the second pool of second fluorophore-labelled nucleic acid; (d) determining the similarity of the sample to patient and control pools by comparing the first fluorescence emission signals and the second fluorescence emission signals, and the first emission signals and the third fluorescence emission signals; and (e) classifying the sample as endometrial disease where the first fluorescence emission signals are more similar to the second fluorescence emission signals than to the third fluorescent emission signals, and classifying the sample as non-endometrial disease where the first fluorescence emission signals are more similar to the third fluorescence emission signals than to the second fluorescent emission signals, wherein the first microarray and the second microarray are similar to each other, exact replicas of each other, or are identical, and wherein the similarity is defined by a statistical method such that the cell sample and control are similar where the p value of the similarity is less than 0.01.
[00240] In aspects of the invention, the array can be used to monitor the time course of expression of one or more endometrial cancer polynucleotide markers in the array. This can occur in various biological contexts such as tumor progression.
[00241] The array is also useful for ascertaining differential expression patterns of endometrial cancer polynucleotide markers, and optionally other markers, in normal and abnormal cells. This may provide a battery of nucleic acids that could serve as molecular targets for diagnosis or therapeutic intervention.
[00242] Protein Methods. Binding agents may be used for a variety of diagnostic and assay applications. There are a variety of assay formats known to the skilled artisan for using a binding agent to detect a target molecule in a sample. (For example, see Harlow and Lane, Antibodies: A Laboratory Manual, Cold Spring Harbor Laboratory, 1988). In general, the presence or absence of an
endometrial disease (e.g. , cancer) in a subject may be determined by (a) contacting a sample from the subject with a binding agent; (b) detecting in the sample a level of polypeptide that binds to the binding agent; and (c) comparing the level of polypeptide with a predetermined standard or cut-off value.
[00243] In particular embodiments of the invention, the binding agent is an antibody. Antibodies specifically reactive with one or more endometrial disease marker, or derivatives, such as enzyme conjugates or labelled derivatives, may be used to detect one or more endometrial disease marker in various samples (e.g. , biological materials). They may be used as diagnostic or prognostic reagents and they may be used to detect abnormalities in the level of expression of one or more endometrial disease marker, or abnormalities in the structure, and/or temporal, tissue, cellular, or subcellular location of one or more endometrial disease marker. Antibodies may also be used to screen potentially therapeutic compounds in vitro to determine their effects on disorders (e.g. , cancer) involving one or more endometrial cancer markers, and other conditions. In vitro immunoassays may also be used to assess or monitor the efficacy of particular therapies.
[00244] In an aspect, the invention provides a method for monitoring or diagnosing an endometrial disease (e.g. , cancer) in a subject by quantifying one or more endometrial cancer markers in a biological sample from the subject including reacting the sample with antibodies specific for one or more endometrial cancer markers, which are directly or indirectly labelled with detectable substances and detecting the detectable substances. In a particular embodiment of the invention, endometrial cancer markers are quantified or measured.
[00245] In a further aspect, the invention provides a method for detecting or screening for an endometrial disease (e.g. , cancer) by: (a) obtaining a sample suspected of containing one or more protein endometrial cancer markers associated with an endometrial disease; (b) contacting the sample with antibodies that specifically bind to the protein endometrial cancer markers under conditions effective to bind the antibodies and form complexes; (c) measuring the amount of protein endometrial cancer markers present in the sample by quantifying the amount of the complexes; and (d) comparing the amount of protein endometrial cancer markers present in the samples with the amount of protein endometrial cancer markers in a control, wherein a change or significant difference in the amount of protein endometrial cancer markers in the sample compared with the amount in the control is indicative of an endometrial disease.
[00246] In an embodiment, the invention contemplates a method for monitoring the progression of an endometrial disease (e.g. , cancer) in an individual, by: (a) contacting antibodies which bind to one or more protein endometrial cancer markers with a sample from the individual so as to form complexes including the antibodies and one or more endometrial cancer markers in the sample; (b) determining or detecting the presence or amount of complex formation in the sample; (c) repeating steps (a) and (b) at a point later in time; and (d) comparing the result of step (b) with the result of step
(c), wherein a difference in the amount of complex formation is indicative of disease, disease stage, and/or progression of the disease in the individual.
[00247] The amount of complexes may also be compared to a value representative of the amount of the complexes from an individual not at risk of, or afflicted with, an endometrial disease at different stages. A significant difference in complex formation may be indicative of advanced disease (e.g. , advanced endometrial cancer, or an unfavourable prognosis).
[00248] In aspects of the invention for diagnosis and monitoring endometrial cancer, the endometrial cancer markers include one or more of those up-regulated in cancer samples as compared to normal samples, as shown in Table 5a, Table 5b, Table 8, and/or Table 9. Examples include, without limitation, a heterogeneous nuclear ribonucleoprotein (e.g. , A2/B (hnRNPA2/B1 )), L- lactate dehydrogenase A (LDHA), anterior gradient protein2 homolog, nucleophosmin, cystatin B, hepatoma derived growth factor, and calcyphosin, and fragments thereof. In certain embodiments, higher levels of the up-regulated markers, particularly significantly higher levels, compared to a control (normal or benign), are indicative of the prognosis of an endometrial cancer patient's outcome.
[00249] In other aspects, the endometrial cancer markers include one or more of those down- regulated in cancer samples as compared to normal samples, as shown in Table 5a, Table 5b, Table 8, and/or Table 9. Examples include, without limitation, fibrinogen (e.g. , alpha, beta, and/or gamma chain), apolipoprotein A1 , haptoglobin (HP), and serpin H1 (or heat-shock protein 47 kD (hsp47) and fragments thereof. In certain embodiments, lower levels of the down-regulated markers, particularly significantly lower levels, compared to a control (normal or benign), are indicative of the prognosis of an endometrial cancer patient's outcome.
[00250] Antibodies may be used in any known immunoassays that rely on the binding interaction between antigenic determinants of one or more endometrial disease marker and the antibodies. Immunoassay procedures for in vitro detection of antigens in fluid samples are also well known in the art. (See, for example, Paterson et al. , Int. J. Can. 37:659 (1986) and Burchell et al. , Int. J. Can. 34:763 (1984) for a general description of immunoassay procedures. ) Qualitative and/or quantitative determinations of one or more endometrial disease marker in a sample may be accomplished by competitive or non-competitive immunoassay procedures in either a direct or indirect format.
Detection of one or more endometrial disease marker using antibodies can be done utilizing immunoassays which are run in either the forward, reverse or simultaneous modes. Examples of immunoassays are radioimmunoassays (RIA), enzyme immunoassays (e.g. , ELISA),
immunofluorescence, immunoprecipitation, latex agglutination, hemagglutination, histochemical tests, and sandwich (immunometric) assays. These terms are well understood by those skilled in the art. A person skilled in the art will know, or can readily discern, other immunoassay formats without undue experimentation.
[00251] According to an embodiment of the invention, an immunoassay for detecting one or more endometrial cancer markers in a biological sample includes contacting binding agents that specifically bind to endometrial cancer markers in the sample under conditions that allow the formation of first complexes including a binding agent and endometrial cancer markers and determining the presence or amount of the complexes as a measure of the amount of endometrial cancer markers contained in the sample. In a particular embodiment, the binding agents are labelled differently or are capable of binding to different labels.
[00252] Antibodies may be used to detect and quantify one or more endometrial cancer markers in a sample in order to diagnose and treat pathological states. In particular, the antibodies may be used in immunohistochemical analyses, for example, at the cellular and sub-subcellular level, to detect one or more endometrial cancer markers, to localize them to particular endometrial cells and tissues (e.g. , tumor cells and tissues), and to specific subcellular locations, and to quantify the level of expression.
[00253] Immunohistochemical methods for the detection of antigens in tissue samples are well known in the art. For example, immunohistochemical methods are described in Taylor, Arch. Pathol. Lab. Med. 102: 1 12 (1978). Briefly, in the context of the invention, a tissue sample obtained from a subject suspected of having an endometrial-related problem is contacted with antibodies, preferably monoclonal antibodies recognizing one or more endometrial cancer markers. The site at which the antibodies are bound is determined by selective staining of the sample by standard
immunohistochemical procedures. The same procedure may be repeated on the same sample using other antibodies that recognize one or more endometrial cancer markers. Alternatively, a sample may be contacted with antibodies against one or more endometrial cancer markers simultaneously, provided that the antibodies are labelled differently or are able to bind to a different label. The tissue sample may be normal endometrial tissue, or a cancer tissue or a benign tissue.
[00254] An antibody microarray in which binding sites include immobilized, preferably monoclonal, antibodies specific to a substantial fraction of marker-derived endometrial cancer markers of interest can be utilized in the present invention. Antibody arrays can be prepared using methods known in the art (see, for example, Zhu et al. , Science 293:2101 (2001 ) and reference 20).
[00255] Antibodies specific for one or more endometrial marker may be labelled with a detectable substance and localised in biological samples based upon the presence of the detectable substance. Examples of detectable substances include, but are not limited to, the following: radioisotopes (e.g., 3H, 14C, 35s, 1251, 1311), fluorescent labels (e.g. , FITC, rhodamine, lanthanide phosphors), luminescent labels such as luminol; enzymatic labels (e.g. , horseradish peroxidase, beta- galactosidase, luciferase, alkaline phosphatase, acetylcholinesterase), biotinyl groups (which can be detected by marked avidin (e.g. , streptavidin containing a fluorescent marker or enzymatic activity that can be detected by optical or colorimetric methods), predetermined polypeptide epitopes recognized by a secondary reporter (e.g. , leucine zipper pair sequences, binding sites for secondary antibodies,
metal binding domains, epitope tags). In some embodiments, labels are attached via spacer arms of various lengths to reduce potential steric hindrance. Antibodies may also be coupled to electron dense substances, such as ferritin or colloidal gold, which are readily visualised by electron microscopy.
[00256] One of the ways an antibody can be detectably labelled is to link it directly to an enzyme. The enzyme when later exposed to its substrate will produce a product that can be detected.
Examples of detectable substances that are enzymes are horseradish peroxidase, beta- galactosidase, luciferase, alkaline phosphatase, acetylcholinesterase, malate dehydrogenase, ribonuclease, urease, catalase, glucose-6-phosphate, staphylococcal nuclease, delta-5-steriod isomerase, yeast alcohol dehydrogenase, alpha-glycerophosphate, those phosphate isomerase, asparaginase, glucose oxidase, and acetylcholine esterase.
[00257] For increased sensitivity in an immunoassay system a fluorescence-emitting metal atom such as Eu (europium) and other lanthanides can be used. These can be attached to the desired molecule by means of metal-chelating groups such as DTPA or EDTA.
[00258] A bioluminescent compound may also be used as a detectable substance.
Bioluminescence is a type of chemiluminescence found in biological systems where a catalytic protein increases the efficiency of the chemiluminescent reaction. The presence of a bioluminescent molecule is determined by detecting the presence of luminescence. Examples of bioluminescent detectable substances are luciferin, luciferase and aequorin.
[00259] Indirect methods may also be employed in which the primary antigen-antibody reaction is amplified by the introduction of a second antibody, having specificity for the antibody reactive against one or more endometrial cancer markers. By way of example, if the antibody having specificity against one or more endometrial disease marker is a rabbit IgG antibody, the second antibody may be goat anti-rabbit gamma-globulin labelled with a detectable substance as described herein.
[00260] Methods for conjugating or labelling the antibodies discussed above may be readily accomplished by one of ordinary skill in the art. See, for example, Inman, Methods In Enzymology, Vol. 34, Affinity Techniques, Enzyme Purification: Part B, Jakoby and Wichek (eds. ), Academic Press, New York, p. 30, 1974; and Wilchek and Bayer, "The Avidin-Biotin Complex in Bioanalytical Applications", Anal. Biochem. 171 : 1 -32, 1988, for methods for conjugating or labelling the antibodies with enzyme or ligand binding partner.
[00261] Cytochemical techniques known in the art for localizing antigens using light and electron microscopy may be used to detect one or more endometrial cancer markers. Generally, antibodies may be labelled with detectable substances and one or more endometrial cancer markers may be localised in tissues and cells based upon the presence of the detectable substances.
[00262] In the context of the methods of the invention, the sample, binding agents (e.g. , antibodies specific for one or more endometrial cancer markers), or one or more endometrial cancer markers may be immobilized on a carrier or support. Examples of suitable carriers or supports are agarose, cellulose, nitrocellulose, dextran, Sephadex, Sepharose, liposomes, carboxymethyl cellulose, polyacrylamides, polystyrene, gabbros, filter paper, magnetite, ion-exchange resin, plastic film, plastic tube, glass, polyamine-methyl vinyl-ether-maleic acid copolymer, amino acid copolymer, ethylene-maleic acid copolymer, nylon, silk, etc. The support material may have any possible configuration including spherical (e.g. , bead), cylindrical (e.g. , inside surface of a test tube or well, or the external surface of a rod), or flat (e.g. , sheet, test strip). Thus, the carrier may be in the shape of, for example, a tube, test plate, well, beads, disc, sphere, etc. The immobilized antibody may be prepared by reacting the material with a suitable insoluble carrier using known chemical or physical methods, for example, cyanogen bromide coupling. An antibody may be indirectly immobilized using a second antibody specific for the antibody. For example, mouse antibody specific for an endometrial disease marker may be immobilized using sheep anti-mouse IgG Fc fragment specific antibody coated on the carrier or support.
[00263] Where a radioactive label is used as a detectable substance, one or more endometrial disease marker may be localized by radioautography. The results of radioautography may be quantified by determining the density of particles in the radioautographs by various optical methods, or by counting the grains.
[00264] Time-resolved fluorometry may be used to detect a signal. For example, the method described by Christopoulos TK and Diamandis EP in Anal Chem 1992:64:342-346 may be used with a conventional time-resolved fluorometer.
[00265] In accordance with an embodiment of the invention, a method is provided wherein one or more endometrial disease marker antibodies are directly or indirectly labelled with enzymes, substrates for the enzymes are added wherein the substrates are selected so that the substrates, or a reaction product of an enzyme and substrate, form fluorescent complexes with a lanthanide metal (e.g. , europium, terbium, samarium, and dysprosium, preferably europium and terbium). A lanthanide metal is added and one or more endometrial cancer markers are quantified in the sample by measuring fluorescence of the fluorescent complexes. Enzymes are selected based on the ability of a substrate of the enzyme, or a reaction product of the enzyme and substrate, to complex with lanthanide metals such as europium and terbium. Suitable enzymes and substrates that provide fluorescent complexes are described in US Patent No. 5,31 12,922 to Diamandis. Examples of suitable enzymes include alkaline phosphatase and β-galactosidase. Preferably, the enzyme is alkaline phosphatase.
[00266] Examples of enzymes and substrates for enzymes that provide such fluorescent complexes are described in US Patent No. 5,312,922 to Diamandis. By way of example, when the
antibody is directly or indirectly labelled with alkaline phosphatase the substrate employed in the method may be 4-methylumbelliferyl phosphate, 5-fluorosalicyl phosphate, or diflunisal phosphate. The fluorescence intensity of the complexes is typically measured using a time-resolved fluorometer (e.g. , a CyberFluor 615 Immunoanalyzer (Nordion International, Kanata, Ontario)).
[00267] One or more endometrial disease marker antibodies may also be indirectly labelled with an enzyme. For example, the antibodies may be conjugated to one partner of a ligand binding pair, and the enzyme may be coupled to the other partner of the ligand binding pair. Representative examples include avidin-biotin, and riboflavin-riboflavin binding protein. In an embodiment, the antibodies are biotinylated, and the enzyme is coupled to streptavidin. In another embodiment, an antibody specific for endometrial disease marker antibody is labelled with an enzyme.
[00268] In accordance with an embodiment, the invention provides means for determining one or more endometrial cancer markers in a sample by measuring one or more endometrial cancer markers by immunoassay. It will be evident to a skilled artisan that a variety of immunoassay methods can be used to measure one or more endometrial cancer markers. In general, an immunoassay method may be competitive or non-competitive. Competitive methods typically employ an immobilized or immobilizable antibody to one or more endometrial disease marker and a labelled form of one or more endometrial disease marker. Sample endometrial disease cancer markers and labelled endometrial cancer markers compete for binding to antibodies to endometrial cancer markers. After separation of the resulting labelled endometrial cancer markers that have become bound to antibodies (bound fraction) from that which has remained unbound (unbound fraction), the amount of the label in either bound or unbound fraction is measured and may be correlated with the amount of endometrial cancer markers in the test sample in any conventional manner (e.g. , by comparison to a standard curve).
[00269] In an aspect, a non-competitive method is used for the determination of one or more endometrial cancer markers, with the most common method being the "sandwich" method. In this assay, two antibodies to endometrial cancer markers are employed. One of the antibodies to endometrial cancer markers is directly or indirectly labelled (sometimes referred to as the "detection antibody") and the other is immobilized or immobilizable (sometimes referred to as the "capture antibody"). The capture and detection antibodies can be contacted simultaneously or sequentially with the test sample. Sequential methods can be accomplished by incubating the capture antibody with the sample, and adding the detection antibody at a predetermined time thereafter (sometimes referred to as the "forward" method); or the detection antibody can be incubated with the sample first and then the capture antibody added (sometimes referred to as the "reverse" method). After the necessary incubation(s) have occurred, to complete the assay, the capture antibody is separated from the liquid test mixture, and the label is measured in at least a portion of the separated capture antibody phase or the remainder of the liquid test mixture. Generally, it is measured in the capture antibody phase since it includes endometrial disease markers bound by ("sandwiched" between) the
capture and detection antibodies. In an embodiment, the label may be measured without separating the capture antibodies and liquid test mixture.
[00270] In a typical two-site immunometric assay for endometrial cancer markers, one or both of the capture and detection antibodies are polyclonal antibodies or one or both of the capture and detection antibodies are monoclonal antibodies (i.e. , polyclonal / polyclonal, monoclonal / monoclonal, or monoclonal / polyclonal). The label used in the detection antibody can be selected from any of those known conventionally in the art. The label may be an enzyme or a chemiluminescent moiety, but it can also be a radioactive isotope, a fluorophor, a detectable ligand (e.g. , detectable by a secondary binding by a labelled binding partner for the ligand), and the like. In a particular aspect, the antibody is labelled with an enzyme which is detected by adding a substrate that is selected so that a reaction product of the enzyme and substrate forms fluorescent complexes. The capture antibody may be selected so that it provides a means for being separated from the remainder of the test mixture. Accordingly, the capture antibody can be introduced to the assay in an already immobilized or insoluble form, or can be in an immobilizable form, that is, a form which enables immobilization to be accomplished subsequent to introduction of the capture antibody to the assay. An immobilized capture antibody may include an antibody covalently or non-covalently attached to a solid phase such as a magnetic particle, a latex particle, a microtiter plate well, a bead, a cuvette, or other reaction vessel. An example of an immobilizable capture antibody is antibody which has been chemically modified with a ligand moiety, e.g. , a hapten, biotin, or the like, and which can be subsequently immobilized by contact with an immobilized form of a binding partner for the ligand, e.g. , an antibody, avidin, or the like. In an embodiment, the capture antibody may be immobilized using a species specific antibody for the capture antibody that is bound to the solid phase.
[00271] The above-described immunoassay methods and formats are intended to be exemplary and are not limiting.
[00272] Computer Systems. Analytic methods contemplated herein can be implemented by use of computer systems and methods described below and known in the art. Thus, the invention provides computer readable media including one or more endometrial cancer markers, and/or polynucleotides encoding one or more endometrial cancer markers, and optionally other markers (e.g. , markers of endometrial cancer). "Computer readable media" refers to any medium that can be read and accessed directly by a computer, including but not limited to magnetic storage media, such as floppy discs, hard disc storage medium, and magnetic tape; optical storage media such as CD- ROM; electrical storage media such as RAM and ROM; and hybrids of these categories such as magnetic / optical storage media. Thus, the invention contemplates computer readable medium having recorded thereon markers identified for patients and controls.
[00273] "Recorded" refers to a process for storing information on computer readable medium. The skilled artisan can readily adopt any of the presently known methods for recording information on
computer readable medium to generate manufactures including information on one or more endometrial cancer markers, and optionally other markers.
[00274] A variety of data processor programs and formats can be used to store information on one or more endometrial cancer markers, and/or polynucleotides encoding one or more endometrial cancer markers, and other markers on computer readable medium. For example, the information can be represented in a word processing text file, formatted in commercially-available software such as WordPerfect and Microsoft Word, or represented in the form of an ASCI I file, stored in a database application, such as SQL, DB2, Sybase, Oracle, or the like. Any number of dataprocessor structuring formats (e.g. , text file or database) may be adapted in order to obtain computer readable medium having recorded thereon the marker information.
[00275] By providing the marker information in computer readable form, one can routinely access the information for a variety of purposes. For example, one skilled in the art can use the information in computer readable form to compare marker information obtained during or following therapy with the information stored within the data storage means.
[00276] The invention provides a medium for holding instructions for performing a method for determining or whether a patient has an endometrial disease (e.g. , endometrial cancer) or a predisposition to an endometrial disease (e.g. , cancer), including determining the presence or absence of one or more endometrial cancer markers, and/or polynucleotides encoding one or more endometrial cancer markers, and optionally other markers, and based on the presence or absence of the one or more endometrial cancer markers, and/or polynucleotides encoding one or more endometrial cancer markers, and optionally other markers, determining endometrial receptivity, endometrial disease (e.g. , cancer) or a pre-disposition to an endometrial disease (e.g. , cancer), and optionally recommending a procedure or treatment.
[00277] The invention also provides in an electronic system and/or in a network, a method for determining whether a subject has an endometrial disease (e.g. , cancer) or a pre-disposition to an endometrial disease (e.g. , cancer), including determining the presence or absence of one or more endometrial cancer markers and/or polynucleotides encoding one or more endometrial cancer markers, and optionally other markers (e.g. , cancer markers), and based on the presence or absence of the one or more endometrial cancer markers and/or polynucleotides encoding one or more endometrial cancer markers, and optionally other markers, determining whether the subject has an endometrial disease (e.g., cancer) or a pre-disposition to an endometrial disease (e.g. , cancer), and optionally recommending a procedure or treatment.
[00278] The invention further provides in a network, a method for determining whether a subject is receptive to in vitro fertilization, has an endometrial disease (e.g. , cancer) or a pre-disposition to an endometrial disease (e.g. , cancer) including: (a) receiving phenotypic information on the subject and information on one or more endometrial cancer markers, and/or polynucleotides encoding one or
more endometrial cancer markers, and optionally other markers associated with samples from the subject; (b) acquiring information from the network corresponding to the one or more endometrial cancer markers, and/or polynucleotides encoding one or more endometrial cancer markers, and optionally other markers; and (c) based on the phenotypic information and information on the one or more endometrial cancer markers, and/or polynucleotides encoding one or more endometrial cancer markers, and optionally other markers, determining whether the subject is receptive to in vitro fertilization, has an endometrial disease (e.g. , cancer) or a pre-disposition to an endometrial disease (e.g. , cancer); and (d) optionally recommending a procedure or treatment.
[00279] The invention still further provides a system for identifying selected records that identify a diseased endometrial cell or tissue (e.g. , cancer cell or tissue) or an endometrial cell or tissue phase. A system of the invention generally includes a digital computer; a database server coupled to the computer; a database coupled to the database server having data stored therein, the data including records of data including one or more endometrial cancer markers, and/or polynucleotides encoding one or more endometrial cancer markers, and optionally other endometrial cancer markers, and a code mechanism for applying queries based upon a desired selection criteria to the data file in the database to produce reports of records which match the desired selection criteria.
[00280] In an aspect of the invention a method is provided for detecting endometrial cancer tissue or cells using a computer having a processor, memory, display, and input / output devices, the method by: (a) creating records of one or more endometrial cancer markers, and/or polynucleotides encoding one or more endometrial cancer markers, and optionally other markers of cancer identified in a sample suspected of containing endometrial cancer cells or tissue; (b) providing a database including records of data including one or more endometrial cancer markers, and/or polynucleotides encoding one or more endometrial cancer markers, and optionally other markers of cancer; and (c) using a code mechanism for applying queries based upon a desired selection criteria to the data file in the database to produce reports of records of step (a) which provide a match of the desired selection criteria of the database of step (b) the presence of a match being a positive indication that the markers of step (a) have been isolated from cells or tissue that are endometrial cancer cells or endometrial cancer tissue.
[00281] The invention contemplates a business method for determining whether a subject is receptive to in vitro fertilization, has an endometrial disease (e.g. , cancer) or a pre-disposition to endometrial cancer including: (a) receiving phenotypic information on the subject and information on one or more endometrial cancer markers, and/or polynucleotides encoding the markers, and optionally other markers, associated with samples from the subject; (b) acquiring information from a network corresponding to one or more endometrial cancer markers, and/or polynucleotides encoding the markers, and optionally other markers; and (c) based on the phenotypic information, information on one or more endometrial cancer markers, and/or polynucleotides encoding the markers, and optionally other markers, and acquired information, determining whether the subject is receptive to in
vitro fertilization, has an endometrial disease (e.g. , cancer) or a pre-disposition to an endometrial disease (e.g. , cancer); and (d) optionally recommending a procedure or treatment. In one embodiment, the endometrial cancer markers are protein endometrial cancer markers.
[00282] In an aspect of the invention, the computer systems, components, and methods described herein are used to monitor disease or determine the stage of disease.
[00283] Imaging Methods. Binding agents, particularly antibodies, specific for one or more endometrial disease markers may also be used in imaging methodologies in the management of an endometrial disease.
[00284] In an aspect, the invention provides a method for imaging tumors associated with one or more endometrial cancer markers.
[00285] The invention also contemplates imaging methods described herein using multiple markers for an endometrial disease. Preferably, each agent is labelled so that it can be distinguished during the imaging.
[00286] In an embodiment the method is an in vivo method and a subject or patient is administered one or more agents that carry an imaging label and that are capable of targeting or binding to one or more endometrial cancer markers. The agent is allowed to incubate in vivo and bind to the endometrial cancer markers associated with endometrial cells or tissues of a particular phase or associated with diseased cells or tissues, (e.g. , endometrial tumor). The presence of the label is localized to the endometrial cells or tissues, and the localized label is detected using imaging devices known to those skilled in the art.
[00287] The agent may be an antibody or chemical entity that recognizes the endometrial cancer markers. In an aspect of the invention the agent is a polyclonal antibody or monoclonal antibody, or fragments thereof, or constructs thereof including but not limited to, single chain antibodies, bifunctional antibodies, molecular recognition units, and peptides or entities that mimic peptides. The antibodies specific for the endometrial cancer markers used in the methods of the invention may be obtained from scientific or commercial sources, or isolated native endometrial cancer markers or recombinant endometrial cancer markers may be utilized to prepare antibodies etc. as described herein.
[00288] An agent may be a peptide that mimics the epitope for an antibody specific for an endometrial disease marker and binds to the marker. The peptide may be produced on a commercial synthesizer using conventional solid phase chemistry. By way of example, a peptide may be prepared that includes tyrosine, lysine, or phenylalanine to which N2S2 chelate is complexed (see US Patent No. 4,897,255). An anti-endocrine marker peptide conjugate is then combined with a radiolabel (e.g. , sodium 99mTc pertechnetate or sodium 188Re perrhenate) and it may be used to locate an endometrial disease marker producing cell or tissue (e.g. , tumor).
[00289] The agent carries a label to image the endometrial cancer markers. The agent may be labelled for use in radionuclide imaging. In particular, the agent may be directly or indirectly labelled with a radioisotope. Examples of radioisotopes that may be used in the invention are the following: 277Ac, 21 1 At, 128Ba, 131 Ba, 7Be, 204Bi, 205Bi, 206Bi, 76Br, 77Br, 82Br, 109Cd, 47Ca, 1 1 C, 14C, 36CI, 48Cr, 51 Cr, 62Cu, 64Cu, 67Cu, 165Dy, 155Eu, 18F, 153Gd, 66Ga, 67Ga, 68Ga, 72Ga, 198Au, 3H, 166HO, 11 1 ln, 1 13mln, 1 15mln, 1231, 1251, 131 1, 189lr, 191 mlr, 192lr, 194lr, 52Fe, 55Fe, 59Fe, 177Lu, 150, 191 m-1910s, 109Pd, 32P, 33P, 42K, 226Ra, 186Re, 188Re, 82mRb, 153Sm, 46Sc, 47Sc, 72Se, 75Se, 105Ag, 22Na, 24Na, 89Sr, 35s, 38s, 177Ta, 96Tc, 99mTc, 201 TI, 202TI, 1 13Sn, 117mSn, 121 Sn, 166Yb, 169Yb, 175Yb, 88Y, 90Y, 62Zn and 65Zn. Preferably the radioisotope is 131 1, 1251, 1231, 1 11 1, 99mTc, 90Y, 186Re, 188Re, 32P, 153Sm, 67Ga, 201 TI 77Br, or 18F, and is imaged with a photoscanning device.
[00290] Procedures for biological agents with the radioactive isotopes are generally known in the art. US Patent No. 4,302,438 describes tritium procedures. Procedures for iodinating, tritium labelling, and 35S especially adapted for murine monoclonal antibodies are described by Goding, J.vV. (supra, pp. 124-126) and the references cited therein. Other procedures for iodinating biological agents, such as antibodies, binding portions thereof, probes, or ligands, are described in the scientific literature (see Hunter and Greenwood, Nature 144:945 (1962); David et al. , Biochemistry 13: 1014- 1021 (1974); and US Patent Nos. 3,867,517 and 4,376, 110). Iodinating procedures for agents are described by Greenwood, F. et al. , Biochem. J. 89: 1 14-123 ( 1963); Marchalonis, J. , Biochem. J. 113:299-305 (1969); and Morrison, M. et al. , Immunochemistry, 289-297 (1971 ). 99m Tc-labeling procedures are described by Rhodes, B. et al. in Burchiel, S. et al. (eds. ), Tumor Imaging: The Radioimmunochemical Detection of Cancer, New York: Masson 1 1 1 -123 (1982) and the references cited therein. Labelling of antibodies or fragments with technetium-99m are also described for example in US Patent Nos. 5,317,091 ; 4,478,815; 4,478,818; 4,472,371 ; Re 32,417; and 4,311 ,688. Procedures suitable for 1 1 1 In- biological agents are described by Hnatowich, D.J. et al., J. Immun. Methods, 65: 147-157 (1983), Hnatowich, D. et al. , J. Applied Radiation, 35:554-557 (1984), and Buckley, R.G. et al. , F.E.B.S. 166:202-204 (1984).
[00291] An agent may also be labelled with a paramagnetic isotope for purposes of an in vivo method of the invention. Examples of elements that are useful in magnetic resonance imaging include gadolinium, terbium, tin, iron, or isotopes thereof. See, for example, Schaefer et al. , (1989) JACC 14, 472-480; Shreve et al. , ( 1986) Magn. Reson. Med. 3, 336-340; Wolf, G L , (1984) Physiol. Chem. Phys. Med. N MR 16, 93-95; Wesbey et al. ( 1984), Physiol. Chem. Phys. Med. N MR 16, 145- 155; Runge et al. (1984), Invest. Radiol. 19, 408-415 for discussions on in vivo nuclear magnetic resonance imaging.
[00292] In the case of a radiolabeled agent, the agent may be administered to the patient, it is localized to the cell or tissue (e.g. , tumor) having an endometrial disease marker with which the agent binds, and is detected or "imaged" in vivo using known techniques such as radionuclear scanning
using e.g. , a gamma camera or emission tomography. (See, for example, A. R. Bradwell et al. , "Developments in Antibody Imaging", Monoclonal Antibodies for Cancer Detection and Therapy; R.VV. Baldwin et al. (eds. ), pp. 65-85 (Academic Press 1985). ) A positron emission transaxial tomography scanner, such as designated Pet VI located at Brookhaven National Laboratory, can also be used where the radiolabel emits positrons (e.g. , 1 1 C, 18F, 150, and 13N).
[00293] Whole body imaging techniques using radioisotope labelled agents can be used for locating diseased cells and tissues (e.g. , primary tumors and tumors which have metastasized). Antibodies specific for endometrial cancer markers, or fragments thereof having the same epitope specificity, are bound to a suitable radioisotope, or a combination thereof, and administered parenterally. For endometrial cancer, administration preferably is intravenous. The bio-distribution of the label can be monitored by scintigraphy, and accumulations of the label are related to the presence of endometrial cancer cells. Whole body imaging techniques are described in US Patent Nos.
4,036,945 and 4,31 1 ,688. Other examples of agents useful for diagnosis and therapeutic use that can be coupled to antibodies and antibody fragments include metallothionein and fragments (see US Patent No. 4,732,864). These agents are useful in diagnosis staging and visualization of cancer, in particular endometrial cancer, so that surgical and/or radiation treatment protocols can be used more efficiently.
[00294] An imaging agent may carry a bioluminescent or chemiluminescent label. Such labels include polypeptides known to be fluorescent, bioluminescent, or chemiluminescent, or, that act as enzymes on a specific substrate (reagent), or can generate a fluorescent, bioluminescent or chemiluminescent molecule. Examples of bioluminescent or chemiluminescent labels include luciferases, aequorin, obelin, mnemiopsin, berovin, a phenanthridinium ester, and variations thereof and combinations thereof. A substrate for the bioluminescent or chemiluminescent polypeptide may also be utilized in a method of the invention. For example, the chemiluminescent polypeptide can be luciferase and the reagent luciferin. A substrate for a bioluminescent or chemiluminescent label can be administered before, at the same time (e.g. , in the same formulation), or after administration of the agent.
[00295] An imaging agent may include a paramagnetic compound, such as a polypeptide chelated to a metal (e.g. , a metalloporphyrin). The paramagnetic compound may also include a monocrystalline nanoparticle, e.g. , a nanoparticle including a lanthanide (e.g. , Gd) or iron oxide; or, a metal ion such as a lanthanide. As used herein, "lanthanide" refers to elements of atomic numbers 58 to 70, a transition metal of atomic numbers 21 to 29, 42 or 44, a Gd(l l l), a Mn(l l), or an element including a Fe element. Paramagnetic compounds can also include a neodymium iron oxide (NdFe03) or a dysprosium iron oxide (DyFe03). Examples of elements that are useful in magnetic resonance imaging include gadolinium, terbium, tin, iron, or isotopes thereof. (See, for example, Schaefer et al. , (1989) JACC 14, 472-480; Shreve et al. (1986), Magn. Reson. Med. 3, 336-340; Wolf, G. L. (1984), Physiol. Chem. Phys. Med. NMR 16, 93-95; Wesbey et al. (1984), Physiol. Chem.
Phys. Med. NMR 16, 145-155; Runge et al. (1984), Invest. Radiol. 19, 408-415 for discussions on in vivo nuclear magnetic resonance imaging. )
[00296] An image can be generated in a method of the invention by computer assisted tomography (CAT), magnetic resonance spectroscopy (MRS) image, magnetic resonance imaging (MRI), positron emission tomography (PET), single-photon emission computed tomography (SPECT), or bioluminescence imaging (BLI) or equivalent.
[00297] Computer assisted tomography (CAT) and computerized axial tomography (CAT) systems and devices well known in the art can be utilized in the practice of the invention. (See, for example, US Patent Nos. 6, 151 ,377; 5,946,371 ; 5,446,799; 5,406,479; 5,208,581 ; and 5, 109, 97. ) The invention may also utilize animal imaging modalities, such as MicroCAT™ (ImTek, Inc. ).
[00298] Magnetic resonance imaging (MRI) systems and devices well known in the art can be utilized in the practice of the invention. For a description of MRI methods and devices, see, for example, US Patent Nos. 6, 151 ,377; 6, 144,202; 6, 128,522; 6, 127,825; 6, 121 ,775; 6, 1 19,032;
6, 1 15,446; 6, 1 1 1 ,410; ,602,891 ; 5,555,251 ; 5,455,512; 5,450,010; 5,378,987; 5,214,382; 5,031 ,624; 5,207,222; 4,985,678; 4,906,931 ; and 4,558,279. MRI and supporting devices are commercially available, for example, from Bruker Medical GMBH; Caprius; Esaote Biomedica; Fonar; GE Medical Systems (GEMS); Hitachi Medical Systems America; Intermagnetics General Corporation; Lunar Corp.; MagneVu; Marconi Medicals; Philips Medical Systems; Shimadzu; Siemens; Toshiba America Medical Systems; including imaging systems, by, e.g. , Silicon Graphics. The invention may also utilize animal imaging modalities such as micro-MRIs.
[00299] Positron emission tomography imaging (PET) systems and devices well known in the art can be utilized in the practice of the invention. For example, a method of the invention may use the system designated Pet VI located at Brookhaven National Laboratory. For descriptions of PET systems and devices, see, for example, US Patent Nos. 6, 151 ,377; 6,072, 177; 5,900,636; 5,608,221 ; 5,532,489; 5,272,343; and 5, 103,098. Animal imaging modalities such as micro-PETs (Concorde Microsystems, Inc. ) can also be used in the invention.
[00300] Single-photon emission computed tomography (SPECT) systems and devices well known in the art can be utilized in the practice of the invention. See, for example, US Patent Nos. 6, 115,446; 6,072, 177; 5,608,221 ; 5,600, 145; 5,210,421 ; 5, 103,098. The methods of the invention may also utilize animal imaging modalities, such as micro-SPECTs.
[00301] Bioluminescence imaging includes bioluminescence, fluorescence, and
chemiluminescence and other photon detection systems and devices that are capable of detecting bioluminescence, fluorescence, or chemiluminescence. Sensitive photon detection systems can be used to detect bioluminescent and fluorescent proteins externally; see, for example, Contag (2000), Neoplasia 2:41 -52; and Zhang (1994), Clin. Exp. Metastasis, 12:87-92. The methods of the invention
can be practiced using any such photon detection device, or variation or equivalent thereof, or in conjunction with any known photon detection methodology, including visual imaging. By way of example, an intensified charge-coupled device (ICCD) camera coupled to an image processor may be used in the invention. (See, e.g. , US Patent No. 5,650, 135. ) Photon detection devices are also commercially available from Xenogen, Hamamatsue.
[00302] Screening Methods. The invention also contemplates methods for evaluating test agents or compounds for their ability to inhibit an endometrial disease (e.g. , cancer) or to contribute to an endometrial disease (e.g. , cancer). Test agents and compounds include but are not limited to peptides such as soluble peptides including Ig-tailed fusion peptides, members of random peptide libraries and combinatorial chemistry-derived molecular libraries made of D- and/or L-configuration amino acids, phosphopeptides (including members of random or partially degenerate, directed phosphopeptide libraries), antibodies (e.g. , polyclonal, monoclonal, humanized, anti-idiotypic, chimeric, single chain antibodies, fragments (e.g. , Fab, F(ab)2, and Fab expression library fragments, and epitope-binding fragments thereof), and small organic or inorganic molecules. The agents or compounds may be endogenous physiological compounds or natural or synthetic compounds.
[00303] The invention provides a method for assessing the potential efficacy of a test agent or therapy for inhibiting an endometrial disease (e.g. , cancer) in a patient, by comparing: (a) levels of one or more protein endometrial cancer markers, and/or polynucleotides encoding endometrial cancer markers, and optionally other markers in a first sample obtained from a patient and exposed to the test agent or therapy; and (b) levels of one or more protein endometrial cancer markers, and/or polynucleotides encoding endometrial cancer markers, and optionally other markers, in a second sample obtained from the patient, wherein the sample is not exposed to the test agent or therapy, wherein a significant difference in the levels of expression of one or more protein endometrial cancer markers, and/or polynucleotides encoding one or more endometrial cancer markers, and optionally the other markers, in the first sample, relative to the second sample, is an indication that the test agent or therapy is potentially efficacious for inhibiting an endometrial disease (e.g. , cancer) in the patient.
[00304] The first and second samples may be portions of a single sample obtained from a patient or portions of pooled samples obtained from a patient.
[00305] In an aspect, the invention provides a method of selecting an agent for inhibiting an endometrial disease (e.g. , cancer) in a patient by: (a) obtaining a sample from the patient;
(b) separately maintaining aliquots of the sample in the presence of a plurality of test agents;
(c) comparing one or more endometrial cancer markers, and/or polynucleotides encoding endometrial cancer markers, and optionally other markers, in each of the aliquots; and (d) selecting one of the test agents which alters the levels of one or more endometrial cancer markers, and/or polynucleotides encoding endometrial cancer markers, and optionally other markers in the aliquot containing that test agent, relative to other test agents.
[00306] In a further aspect, the invention provides a method of selecting an agent for inhibiting or enhancing endometrial cell or tissue phase in a patient by: (a) obtaining a sample of endometrial cell or tissue in a selected phase; (b) separately maintaining aliquots of the sample in the presence of a plurality of test agents; (c) comparing one or more endometrial cancer markers, and/or
polynucleotides encoding endometrial cancer markers, and optionally other markers, in each of the aliquots; and (d) selecting one of the test agents which alters the levels of one or more endometrial cancer markers, and/or polynucleotides encoding endometrial cancer markers, and optionally other markers in the aliquot containing that test agent, relative to other test agents.
[00307] Still another aspect of the invention provides a method of conducting a drug discovery business by: (a) providing one or more methods or assay systems for identifying agents that inhibit an endometrial disease (e.g. , cancer) or affect an endometrial cell or tissues phase in a patient; (b) conducting therapeutic profiling of agents identified in step (a), or further analogs thereof, for efficacy and toxicity in animals; and (c) formulating a pharmaceutical preparation including one or more agents identified in step (b) as having an acceptable therapeutic profile.
[00308] In certain embodiments, the subject method can also include a step of establishing a distribution system for distributing the pharmaceutical preparation for sale, and may optionally include establishing a sales group for marketing the pharmaceutical preparation.
[00309] The invention also contemplates a method of assessing the potential of a test compound to contribute to an endometrial disease (e.g. , cancer) by: (a) maintaining separate aliquots of cells or tissues from a patient with an endometrial disease (e.g. , cancer) in the presence and absence of the test compound; and (b) comparing one or more endometrial cancer markers and/or polynucleotides encoding the endometrial cancer markers, and optionally other markers in each of the aliquots.
[00310] A significant difference between the levels of the markers in the aliquot maintained in the presence of (or exposed to) the test compound relative to the aliquot maintained in the absence of the test compound, indicates that the test compound possesses the potential to contribute to an endometrial disease (e.g. , cancer). In one embodiment, the endometrial cancer markers are protein endometrial cancer markers.
[00311] Kits. The invention also contemplates kits for carrying out the methods of the invention. Kits may typically include two or more components required for performing a diagnostic assay.
Components include but are not limited to compounds, reagents, containers, and/or equipment.
[00312] The methods described herein may be performed by utilizing pre-packaged diagnostic kits including one or more specific endometrial disease marker polynucleotide, endometrial cancer markers, or antibody described herein, which may be conveniently used, e.g. , in clinical settings to screen and diagnose patients and to screen and identify those individuals exhibiting a predisposition to developing an endometrial disease.
[00313] In an embodiment, a container with a kit includes a binding agent as described herein. By way of example, the kit may contain antibodies or antibody fragments which bind specifically to epitopes of one or more endometrial cancer markers and optionally other markers, antibodies against the antibodies labelled with an enzyme; and a substrate for the enzyme. The kit may also contain microtiter plate wells, standards, assay diluent, wash buffer, adhesive plate covers, and/or instructions for carrying out a method of the invention using the kit.
[00314] In an aspect of the invention, the kit includes antibodies or fragments of antibodies which bind specifically to an epitope of one or more polypeptide listed in Table 5a, Table 5b, Table 8, and/or Table 9 that is up-regulated in cancer samples as compared to normal samples, and means for detecting binding of the antibodies to their epitope associated with tumor cells, either as concentrates (including lyophilized compositions), which may be further diluted prior to use or at the concentration of use, where the vials may include one or more dosages. Where the kits are intended for in vivo use, single dosages may be provided in sterilized containers, having the desired amount and concentration of agents. Containers that provide a formulation for direct use, usually do not require other reagents, as for example, where the kit contains a radiolabeled antibody preparation for in vivo imaging.
[00315] A kit may be designed to detect the level of protein EmCa markers or polynucleotides encoding protein endometrial cancer markers in a sample. In an embodiment, the kit includes one or more polynucleotides encoding one or more polypeptides listed in Table 5a, Table 5b, Table 8, and/or Table 9 that are up-regulated in EmCa samples as compared to normal samples. Such kits generally include at least one oligonucleotide probe or primer, as described herein, that hybridizes to a polynucleotide encoding one or more endometrial cancer markers. Such an oligonucleotide may be used, for example, within a PCR or hybridization procedure. Additional components that may be present within the kit include a second oligonucleotide and/or a diagnostic reagent or container to facilitate detection of a polynucleotide encoding one or more protein endometrial cancer markers.
[00316] The invention provides a kit containing a microarray described herein ready for hybridization to target polynucleotide endometrial cancer markers, plus software for the data analysis of the results. The software to be included with the kit includes data analysis methods, in particular mathematical routines for marker discovery, including the calculation of correlation coefficients between clinical categories and marker expression. The software may also include mathematical routines for calculating the correlation between sample marker expression and control marker expression, using array-generated fluorescence data, to determine the clinical classification of the sample.
[00317] The reagents suitable for applying the screening methods of the invention to evaluate compounds may be packaged into convenient kits described herein providing the necessary materials packaged into suitable containers.
[00318] The invention contemplates a kit for assessing the presence of endometrial disease cells, wherein the kit includes antibodies specific for one or more protein endometrial cancer markers, or primers or probes for polynucleotides encoding the endometrial cancer markers and optionally probes, primers or antibodies specific for other markers associated with an endometrial disease (e.g. , cancer).
[00319] The invention relates to a kit for assessing the suitability of each of a plurality of test compounds for inhibiting an endometrial disease (e.g. , cancer) in a patient. The kit includes reagents for assessing one or more or protein endometrial cancer markers, polynucleotides encoding the protein endometrial cancer markers, and optionally a plurality of test agents or compounds.
[00320] Additionally the invention provides a kit for assessing the potential of a test compound to contribute to an endometrial disease (e.g. , cancer). The kit includes endometrial diseased cells (e.g. , cancer cells) and reagents for assessing one or more endometrial cancer markers, polynucleotide encoding the endometrial cancer markers and optionally other markers associated with an endometrial disease.
[00321] Therapeutic Applications. One or more endometrial cancer markers may be targets for immunotherapy. Immunotherapeutic methods include the use of antibody therapy, in vivo vaccines, and ex vivo immunotherapy approaches.
[00322] In one aspect, the invention provides one or more endometrial disease marker antibodies that may be used systemically to treat an endometrial disease associated with the marker. In particular, the endometrial disease is endometrial cancer and one or more endometrial disease marker antibodies may be used systemically to treat endometrial cancer. Preferably antibodies are used that target the tumor cells but not the surrounding non-tumor cells and tissue. In a particular embodiment, the endometrial cancer is EmCa. In another embodiment, the endometrial cancer is Type I EmCa.
[00323] Thus, the invention provides a method of treating a patient susceptible to, or having a disease (e.g. , cancer) that expresses one or more endometrial disease marker, in particular, a marker up-regulated in endometrial cancer (for example, an up-regulated marker in Table 5a, Table 5b, Table 8, and/or Table 9), including administering to the patient an effective amount of an antibody that binds specifically to one or more endometrial disease marker.
[00324] In another aspect, the invention provides a method of inhibiting the growth of tumor cells expressing one or more endometrial cancer markers, including administering to a patient an antibody which binds specifically to one or more endometrial cancer markers in an amount effective to inhibit growth of the tumor cells.
[00325] One or more endometrial disease marker antibodies may also be used in a method for selectively inhibiting the growth of, or killing a cell expressing one or more endometrial disease marker
(e.g. , tumor cell expressing one or more endometrial cancer marker) including reacting one or more endometrial disease marker antibody immunoconjugate or immunotoxin with the cell in an amount sufficient to inhibit the growth of, or kill the cell.
[00326] By way of example, unconjugated antibodies to endometrial cancer markers may be introduced into a patient such that the antibodies bind to endometrial cancer marker expressing cancer cells and mediate growth inhibition of such cells (including the destruction thereof), and the tumor, by mechanisms which may include complement-mediated cytolysis, antibody-dependent cellular cytotoxicity, altering the physiologic function of one or more endometrial cancer markers, and/or the inhibition of ligand binding or signal transduction pathways. In addition to unconjugated antibodies to endometrial cancer markers, one or more endometrial cancer marker antibodies conjugated to therapeutic agents (e.g. , immunoconjugates) may also be used therapeutically to deliver the agent directly to one or more endometrial cancer marker expressing tumor cells and thereby destroy the tumor. Examples of such agents include abrin, ricin A, Pseudomonas exotoxin, or diphtheria toxin; proteins such as tumor necrosis factor, alpha-interferon, beta-interferon, nerve growth factor, platelet derived growth factor, tissue plasminogen activator; and biological response modifiers such as lymphokines, interleukin-1 , interleukin-2, interleukin-6, granulocyte macrophage colony stimulating factor, granulocyte colony stimulating factor, or other growth factors.
[00327] Cancer immunotherapy using one or more endometrial cancer marker antibodies may utilize the various approaches that have been successfully employed for cancers, including but not limited to colon cancer (Arlen et al. , 1998, Crit Rev Immunol 18: 133-138), multiple myeloma (Ozaki et al. , 1997, Blood 90: 3179-3186; Tsunenati et al. , 1997, Blood 90: 2437-2444), gastric cancer (Kasprzyk et al. , 1992, Cancer Res 52: 2771 -2776), B-cell lymphoma (Funakoshi et al. , 1996, J. Immunther Emphasis Tumor Immunol 19: 93-101 ), leukemia (Zhong et al. , 1996, Leuk Res 20: 581 - 589), colorectal cancer (Moun et al. , 1994, Cancer Res 54: 6160-6166); Velders et al. , 1995, Cancer Res 55: 4398-4403), and breast cancer (Shepard et al. , 1991 , J. Clin Immunol 1 1 : 1 17-127).
[00328] In the practice of a method of the invention, endometrial cancer marker antibodies capable of inhibiting the growth of precancer or cancer cells expressing endometrial cancer markers are administered in a therapeutically effective amount to cancer patients whose lesions or tumors express or overexpress one or more endometrial cancer markers. The invention may provide a specific, effective and long-needed treatment for endometrial cancer. The antibody therapy methods of the invention may be combined with other therapies including chemotherapy and radiation.
[00329] Patients may be evaluated for the presence and level of expression or overexpression of one or more endometrial cancer markers in diseased cells and tissues (e.g. , tumors), in particular using immunohistochemical assessments of tissue, quantitative imaging as described herein, or other techniques capable of reliably indicating the presence and degree of expression of one or more endometrial disease markers. Immunohistochemical analysis of tumor biopsies or surgical specimens may be employed for this purpose.
[00330] Endometrial disease marker antibodies useful in treating disease (e.g. , cancer) include those that are capable of initiating a potent immune response against the disease (e.g., tumor) and those that are capable of direct cytotoxicity. In this regard, endometrial disease marker antibodies may elicit cell lysis by either complement-mediated or antibody-dependent cell cytotoxicity (ADCC) mechanisms, both of which require an intact Fc portion of the immunoglobulin molecule for interaction with effector cell Fc receptor sites or complement proteins.
[00331] Endometrial disease marker antibodies that exert a direct biological effect on tumor growth may also be useful in the practice of the invention. Such antibodies may not require the complete immunoglobulin to exert the effect. Potential mechanisms by which such directly cytotoxic antibodies may act include inhibition of cell growth, modulation of cellular differentiation, modulation of tumor angiogenesis factor profiles, and the induction of apoptosis. The mechanism by which a particular antibody exerts an anti-tumor effect may be evaluated using any number of in vitro assays designed to determine ADCC, antibody-dependent macrophage-mediated cytotoxicity (ADMMC), complement-mediated cell lysis, and others known in the art.
[00332] The anti-tumor activity of a particular endometrial cancer marker antibody, or combination of endometrial cancer marker antibodies, may be evaluated in vivo using a suitable animal model. Xenogenic cancer models, where human cancer explants or passaged xenograft tissues are introduced into immune compromised animals, such as nude or SCID mice, may be employed.
[00333] The methods of the invention contemplate the administration of single endometrial disease marker antibodies as well as combinations, or "cocktails", of different individual antibodies such as those recognizing different epitopes of other markers. Such cocktails may have certain advantages in as much as they contain antibodies that bind to different epitopes of endometrial cancer markers and/or exploit different effector mechanisms or combine directly cytotoxic antibodies with antibodies that rely on immune effector functionality. Such antibodies in combination may exhibit synergistic therapeutic effects. In addition, the administration of one or more endometrial disease marker specific antibodies may be combined with other therapeutic agents, including but not limited to chemotherapeutic agents, androgen-blockers, and immune modulators (e.g. , IL2, GM-CSF). The endometrial disease marker specific antibodies may be administered in their "naked" or unconjugated form, or may have therapeutic agents conjugated to them.
[00334] The endometrial disease marker specific antibodies used in the methods of the invention may be formulated into pharmaceutical compositions including a carrier suitable for the desired delivery method. Suitable carriers include any material which when combined with the antibodies retains the function of the antibody and is non-reactive with the subject's immune systems. Examples include any of a number of standard pharmaceutical carriers such as sterile phosphate buffered saline solutions, bacteriostatic water, and the like (see, generally, Remington's Pharmaceutical Sciences 16th Edition, A. Osal. , ed. , 1980).
[00335] One or more endometrial disease marker specific antibody formulations may be administered via any route capable of delivering the antibodies to a disease (e.g. , tumor) site. Routes of administration include, but are not limited to, intravenous, intraperitoneal, intramuscular, intratumor, intradermal, and the like. Preferably, the route of administration is by intravenous injection. Antibody preparations may be lyophilized and stored as a sterile powder, preferably under vacuum, and then reconstituted in bacteriostatic water containing, for example, benzyl alcohol preservative, or in sterile water prior to injection.
[00336] Treatment will generally involve the repeated administration of the antibody preparation via an acceptable route of administration such as intravenous injection (IV), at an effective dose. Dosages will depend upon various factors generally appreciated by those of skill in the art, including the type of disease and the severity, grade, or stage of the disease, the binding affinity and half life of the antibodies used, the degree of endometrial disease marker expression in the patient, the extent of circulating endometrial disease markers, the desired steady-state antibody concentration level, frequency of treatment, and the influence of any chemotherapeutic agents used in combination with the treatment method of the invention. Daily doses may range from about 0.1 to 100 mg / kg. Doses in the range of 10-500 mg antibodies per week may be effective and well tolerated, although even higher weekly doses may be appropriate and/or well tolerated. A determining factor in defining the appropriate dose is the amount of a particular antibody necessary to be therapeutically effective in a particular context. Repeated administrations may be required to achieve disease inhibition or regression. Direct administration of one or more endometrial disease marker antibodies is also possible and may have advantages in certain situations.
[00337] Patients may be evaluated for serum cancer markers in order to assist in the determination of the most effective dosing regimen and related factors. The endometrial cancer assay methods described herein, or similar assays, may be used for quantifying circulating endometrial disease marker levels in patients prior to treatment. Such assays may also be used for monitoring throughout therapy, and may be useful to gauge therapeutic success in combination with evaluating other parameters such as serum levels of endometrial cancer markers.
[00338] The invention further provides vaccines formulated to contain one or more endometrial disease marker or fragment thereof.
[00339] In an embodiment, the invention provides a method of vaccinating an individual against one or more endometrial disease marker listed in Table 5a, Table 5b, Table 8, and/or Table 9, including the step of inoculating the individual with the marker or fragment thereof that lacks activity, wherein the inoculation elicits an immune response in the individual thereby vaccinating the individual against the marker.
[00340] The use in anti-cancer therapy of a tumor antigen in a vaccine for generating humoral and cell-mediated immunity is well known and, for example, has been employed in prostate cancer using
human PSMA and rodent PAP immunogens (Hodge et al. , 1995, Int. J. Cancer 63: 231 -237; and Fong et al. , 1997, J. Immunol. 159: 3113-31 17). These and similar methods can be practiced by employing one or more endometrial cancer markers, or fragment thereof, or polynucleotide encoding the markers and recombinant vectors capable of expressing and appropriately presenting endometrial disease marker immunogens.
[00341] By way of example, viral gene delivery systems may be used to deliver one or more polynucleotide endometrial cancer markers or complementary anti-sense versions thereof. Various viral gene delivery systems which can be used in the practice of this aspect of the invention include, but are not limited to, vaccinia, fowlpox, canarypox, adenovirus, influenza, poliovirus, adeno- associated virus, lentivirus, and sindbus virus (Restifo, 1996, Curr. Opin. Immunol. 8: 658-663). Non- viral delivery systems may also be employed by using naked DNA encoding one or more endometrial cancer marker or fragment thereof introduced into the patient (e.g. , intramuscularly) to induce an antitumor response.
[00342] Various ex vivo strategies may also be employed. One approach involves the use of cells to present one or more endometrial disease marker to a patient's immune system. For example, autologous dendritic cells which express MHC class I and I I, may be pulsed with one or more endometrial disease marker or peptides thereof that are capable of binding to MHC molecules, to thereby stimulate the patients' immune systems (see, for example, Tjoa et al. , 1996, Prostate 28: 65- 69; Murphy et al. , 1996, Prostate 29: 371-380).
[00343] Anti-idiotypic endometrial disease marker specific antibodies can also be used in therapy as a vaccine for inducing an immune response to cells expressing one or more endometrial disease marker. The generation of anti-idiotypic antibodies is well known in the art and can readily be adapted to generate anti-idiotypic endometrial cancer marker specific antibodies that mimic an epitope on one or more endometrial cancer markers (see, for example, Wagner et al. , 1997, Hybridoma 16: 33-40; Foon et al., 1995, J. Clin Invest 96: 334-342; and Herlyn et al. , 1996, Cancer Immunol Immunother 43: 65-76). Such an antibody can be used in anti-idiotypic therapy as presently practiced with other anti-idiotypic antibodies directed against antigens associated with disease (e.g. , tumor antigens).
[00344] Genetic immunization methods may be utilized to generate prophylactic or therapeutic humoral and cellular immune responses directed against cells expressing one or more endometrial cancer marker. One or more DNA molecules encoding endometrial cancer markers, constructs including DNA encoding one or more endometrial cancer markers / immunogens and appropriate regulatory sequences may be injected directly into muscle or skin of an individual, such that the cells of the muscle or skin take-up the construct and express the encoded endometrial cancer markers / immunogens. The endometrial cancer markers / immunogens may be expressed as cell surface proteins or be secreted. Expression of one or more endometrial cancer markers results in the generation of prophylactic or therapeutic humoral and cellular immunity against the disease (e.g. ,
cancer). Various prophylactic and therapeutic genetic immunization techniques known in the art may be used.
[00345] The invention further provides methods for inhibiting cellular activity (e.g. , cell proliferation, activation, or propagation) of a cell expressing one or more endometrial disease marker. This method includes reacting immunoconjugates of the invention (e.g. , a heterogeneous or homogenous mixture) with the cell so that endometrial cancer markers form complexes with the immunoconjugates. A subject with a neoplastic or preneoplastic condition can be treated when the inhibition of cellular activity results in cell death.
[00346] In another aspect, the invention provides methods for selectively inhibiting a cell expressing one or more endometrial disease marker by reacting any one or a combination of the immunoconjugates of the invention with the cell in an amount sufficient to inhibit the cell. Amounts include those that are sufficient to kill the cell or sufficient to inhibit cell growth or proliferation.
[00347] Vectors derived from retroviruses, adenovirus, herpes or vaccinia viruses, or from various bacterial plasmids, may be used to deliver polynucleotides encoding endometrial cancer markers or to a targeted organ, tissue, or cell population. Methods well known to those skilled in the art may be used to construct recombinant vectors that will express antisense polynucleotides for endometrial cancer markers. (See, for example, the techniques described in Sambrook et al. (supra) and Ausubel et al. (supra). )
[00348] Methods for introducing vectors into cells or tissues include those methods discussed herein and which are suitable for in vivo, in vitro and ex vivo therapy. For ex vivo therapy, vectors may be introduced into stem cells obtained from a patient and clonally propagated for autologous transplant into the same patient (See US Patent Nos. 5,399,493 and 5,437,994). Delivery by transfection and by liposome is well known in the art.
[00349] Genes encoding endometrial cancer markers can be turned off by transfecting a cell or tissue with vectors that express high levels of a desired endometrial disease marker-encoding fragment. Such constructs can inundate cells with untranslatable sense or antisense sequences. Even in the absence of integration into the DNA, such vectors may continue to transcribe RNA molecules until all copies are disabled by endogenous nucleases.
[00350] Modifications of gene expression can be obtained by designing antisense molecules, DNA, RNA or PNA, to the regulatory regions of a gene encoding an endometrial disease marker, i.e. , the promoters, enhancers, and introns. Preferably, oligonucleotides are derived from the transcription initiation site, (e.g. , between -10 and +10 regions of the leader sequence). The antisense molecules may also be designed so that they block translation of mRNA by preventing the transcript from binding to ribosomes. Inhibition may also be achieved using "triple helix" base-pairing methodology. Triple helix pairing compromises the ability of the double helix to open sufficiently for the binding of
polymerases, transcription factors, or regulatory molecules. Therapeutic advances using triplex DNA were reviewed by Gee J E et al. , in: Huber and Carr (1994), Molecular and Immunologic Approaches, Futura Publishing Co, Mt Kisco, N.Y.
[00351] Ribozymes are enzymatic RNA molecules that catalyze the specific cleavage of RNA. Ribozymes act by sequence-specific hybridization of the ribozyme molecule to complementary target RNA, followed by endonucleolytic cleavage. The invention therefore contemplates engineered hammerhead motif ribozyme molecules that can specifically and efficiently catalyze endonucleolytic cleavage of sequences encoding an endometrial disease marker.
[00352] Specific ribozyme cleavage sites within any potential RNA target may initially be identified by scanning the target molecule for ribozyme cleavage sites which include the following sequences, GUA, GUU, and GUC. Once the sites are identified, short RNA sequences of between 15 and 20 ribonucleotides corresponding to the region of the target gene containing the cleavage site may be evaluated for secondary structural features which may render the oligonucleotide inoperable. The suitability of candidate targets may also be determined by testing accessibility to hybridization with complementary oligonucleotides using ribonuclease protection assays.
[00353] One or more endometrial cancer markers and polynucleotides encoding the markers, and fragments thereof, may be used in the treatment of an endometrial disease (e.g. , cancer) in a subject. In an aspect, the endometrial cancer markers and polynucleotides encoding the markers are endometrial cancer markers that are down- regulated in endometrial cancer (e.g. , apolipoprotein A1 , serpin H, or one or more of the down- regulated markers listed in Table 5a, Table 5b, Table 8, and/or Table 9). The markers or polynucleotides may be formulated into compositions for administration to subjects suffering from an endometrial disease. Therefore, the invention also relates to a composition including one or more endometrial disease cancer markers or polynucleotides encoding the markers, or a fragment thereof, and a pharmaceutically acceptable carrier, excipient or diluent. A method for treating or preventing an endometrial disease in a subject is also provided including administering to a patient in need thereof, one or more endometrial cancer markers or polynucleotides encoding the markers, or a composition of the invention.
[00354] The invention further provides a method of inhibiting an endometrial disease (e.g. , cancer) in a patient by: (a) obtaining a sample including diseased cells from the patient; (b) separately maintaining aliquots of the sample in the presence of a plurality of test agents; (c) comparing levels of one or more endometrial cancer markers, and/or polynucleotides encoding one or more endometrial cancer markers in each aliquot; and (d) administering to the patient at least one of the test agents which alters the levels of the endometrial cancer markers, markers, and/or polynucleotides encoding one or more cancer markers in the aliquot containing that test agent, relative to the other test agents.
[00355] An active therapeutic substance described herein may be administered in a convenient manner such as by injection (subcutaneous, intravenous, etc. ), oral administration, inhalation,
transdermal application, or rectal administration. Depending on the route of administration, the active substance may be coated in a material to protect the substance from the action of enzymes, acids and other natural conditions that may inactivate the substance. Solutions of an active compound as a free base or pharmaceutically acceptable salt can be prepared in an appropriate solvent with a suitable surfactant. Dispersions may be prepared in glycerol, liquid polyethylene glycols, and mixtures thereof, or in oils.
[00356] The compositions described herein can be prepared by per se known methods for the preparation of pharmaceutically acceptable compositions which can be administered to subjects, such that an effective quantity of the active substance is combined in a mixture with a pharmaceutically acceptable vehicle. Suitable vehicles are described, for example, in Remington's Pharmaceutical Sciences (Remington's Pharmaceutical Sciences, Mack Publishing Company, Easton, Pa. , USA, 1985). On this basis, the compositions include, albeit not exclusively, the active substances in association with one or more pharmaceutically acceptable vehicles or diluents, and contained in buffered solutions with a suitable pH and iso-osmotic with the physiological fluids.
[00357] The compositions are indicated as therapeutic agents either alone or in conjunction with other therapeutic agents or other forms of treatment. The compositions of the invention may be administered concurrently, separately, or sequentially with other therapeutic agents or therapies. The therapeutic activity of compositions, agents, and compounds may be identified using a method of the invention and may be evaluated in vivo using a suitable animal model.
[00358] The inventors' study is a significant advancement in this direction as it places major emphasis on determining the clinical impact of a proteomics based biomarker in diagnosing tumor in patients showing symptoms of endometrial disease.
[00359] The invention is described in the following non-limiting Examples, which are set forth to illustrate and to aid in an understanding of the invention, and should not be construed to limit in any way the scope of the invention as defined in the claims which follow thereafter.
[00360] EXAMPLES
[00361] In Examples 1 -7, the inventors demonstrate the identification of a panel of proteins with consistently differential expression, including apolipoprotein A1 , calcyphosin, cathepsin B, dihydropyrimidinase-like 2, fibrinogen (e.g. , alpha, beta, and gamma chains), haptoglobin, macrophage capping protein (CAP-G), and hsp47, that serve as endometrial cancer biomarkers. In Examples 8-13, the inventors demonstrate the identification of a panel of proteins with consistently differential expression among different subtypes of EmCa as compared to normal tissues, including a heterogeneous nuclear ribonucleoprotein (e.g. , A2/B (hnRNPA2/B1 )), nucleophosmin, LDHA, hepatoma-derived growth factor (HDGF), mitochondrial peroxiredoxin-5, and cystatin B. The inventors also demonstrate the identification of a panel of proteins with consistently differential
expression across all subtypes of EmCa as compared to normal tissues, including alpha-enolase, glucose-6-phosphate isomerase, phosphoglycerate kinase, villin 2 (ezrin), mitochondrial isocitrate dehyrdogenase 2 (NADP+), transketolase, serpin H1 (hsp47), and nucleolin.
[00362] EXAMPLE 1 - SAMPLES AND REAGENTS
[00363] Unless otherwise stated, all reagents are from Sigma-Aldrich (St-Louis, MO).
Endometrial tissues were retrieved from an in-house, dedicated research endometrial tissue bank, as previously described (12). The collection and use of these materials were approved by the Research Ethics Boards of York University, Mount Sinai Hospital, University Health Network, and North York General Hospital. Briefly, samples from hysterectomy specimens were flash frozen in liquid nitrogen within 20 min of devitalizing. A portion of the tissue sample from the mirror face of the histological section was taken for proteomic analysis. This material was washed three times in ~1 mL of PBS with a mixture of protease inhibitors, as described previously (1 mM 4-(2-aminoethyl) benzenesulfonyl fluoride, 10 μ Μ leupeptin, 1 μg/mL aprotinin, and 1 μΜ pepstatin) (14)). It was then homogenized in 0.5 mL of phosphate-buffered saline (PBS) with protease inhibitors using a hand-held homogenizer, and then flash-frozen in liquid nitrogen and stored at -80°C until use.
[00364] A total of 20 samples were used in the present study: ten normal proliferative samples, and ten Type I EmCa samples. Five of the latter were also examined in the inventors' previous study (12); their selection was based on a desire to extend verification of earlier results. Retrieved samples were first thawed and clarified by centrifugation, and the protein concentrations were determined by a Bradford-type assay using Bio-Rad's (Mississauga, ON) protein quantification reagent. Aliquots of 100 μg of proteins from each of the ten normal proliferative samples were pooled; these served as the reference sample for use in all the iTRAQ sets. The protein content of this pooled sample was once again determined as above. For each sample, 100 μg of proteins were digested and labelled individually with the appropriate iTRAQ tag (vide infra). Trypsin digestion and labeling were performed according to the manufacturer's protocol, except that the trypsin used was porcine sequencing grade (Promega, Madison, VVI). A total of seven iTRAQ sets were examined (Table 1 ). Each iTRAQ set included the reference sample, one individual normal proliferative sample, and two EmCa samples (one new and the other a previously examined sample), with the exception of the last two sets which included the remaining normal proliferative samples and an EmCa sample. The assignments of the tags to the sample types were randomized to minimize the effects of any m/z- related biases.
[00365] EXAMPLE 2 - STRONG CATION EXCHANGE (SCX) SEPARATION
[00366] Each iTRAQ set was then separated by SCX fractionation using an HP1050 HPLC instrument (Agilent, Palo Alto, CA) with a 2.1-mm internal diameter x 100-mm-length PolyLC Polysulfoethyl A column packed with 5-μηι beads with 300-A pores (The Nest Group, Southborough, MA). A 2.1-mm-internal diameter x 10-mm-length guard column of the same material was fitted
immediately upstream of the analytical column. Separation was performed as described previously (14). Briefly, the iTRAQ set was diluted with the loading buffer (Buffer A: 15 mM KH2P04 in 25% acetonitrile, pH 3.0) to a total volume of 1.8 ml_, and the pH was adjusted to 3.0 with phosphoric acid. The solution was then filtered using a 0.45-μηι syringe filter (Millipore, Cambridge, ON, Canada) before loading onto the column. Separation was performed using a linear binary gradient over 1 h, plus 30 min of column re-equilibration (see Table 2a). Buffer A was identical in composition to the loading buffer; Buffer B was Buffer A containing 350 mM potassium chloride; Buffer C was Buffer A containing 1 M potassium chloride. Fractions were collected every 2 min using an SF-2120 Super Fraction Collector (Advantec MFS, Dublin, CA) after an initial wait of 2 min to accommodate the void volume. This resulted in a total of 30 SCX fractions per iTRAQ set. These fractions were dried by speed vacuuming (Thermo Savant SC110 A, Holbrook, NY) and re-dissolved in a minimal volume of 5% acetonitrile in 0.1 % formic acid: typically, 8 μΙ_ for each fraction No. 6-9, 12 μί for No. 10-13, 16 μΙ_ for No. 14-16, 20 μΙ_ for No. 17-19, 25 μΙ_ for No. 20-21 , and 30 μΙ_ for the last fractions, No. 22-30. Larger volumes in the later fractions were necessitated by the need to dissolve completely the larger salt pellets resulting from their correspondingly higher salt contents. Fractions No. 1 -5 were not analyzed, as past experience showed that these early fractions contain mostly iTRAQ by-products (data not shown).
[00367] EXAMPLE 3 - REVERSE PHASE (RP) LC-MS / MS
[00368] The SCX fractions No. 6-30 of each iTRAQ set were analyzed by online nano LC-MS/MS using the LC Packings Ultimate instrument (Amsterdam, The Netherlands) fitted with a 10-μί sample loop. The autosampler was used in the microliter pick-up mode. For each sample, 1 μί solution was loaded onto a 5-mm reverse phase (RP) C18 precolumn (LC Packings) at 25 μί/ηιίη and washed for 4 min before switching the precolumn in line with the separation column. The separation column used was a 75^m-internal-diameter x 150-mm-length capillary column (Integrafit capillary from New Objective, VVoburn, MA) packed in-house with 3.5-μηι C18 beads with 100-A pores from Kromasil (Akzo Nobel/EKA Chemicals Inc. , NY). The flow rate used for separation on the RP column was 200 nL/min. Solvent A was 5% acetonitrile in 0.1 % formic acid; Solvent B was 95% acetonitrile in 0.1 % formic acid. The solvent gradient is detailed in Table 2b. A new column was used for each iTRAQ set.
[00369] Online MS/MS was accomplished on a QSTAR Pulsar hybrid quadrupole / time-of-flight (QqTOF) tandem mass spectrometer (Applied Biosystems/MDS SCIEX, Foster City, CA) in information-dependent acquisition mode with the scan cycles set up to perform a 1 -s MS scan followed by five MS/MS scans of the five most abundant peaks for 2 s each and with a dynamic exclusion period of 30 s. The performance of the LC-MS/MS system was evaluated at a minimum of every three days (more frequently when deemed necessary) by means of 80 fmol of bovine serum albumin tryptic digest standard. Mass calibration on the TOF was verified at the same time and adjusted when necessary.
[00370] EXAMPLE 4 - PROTEIN IDENTIFICATION
[00371] The MS/MS spectra were processed by the software ProteinPilot version 2.0.1 , using the Paragon algorithm (15), and against a concatenated Swissprot/Panther database of 66082 distinct human protein entries (132164 entries after the reversed sequences were added). Protein identification was performed using a confidence threshold of 95% (ProteinPilot Unused score > 1.31 ) with M MTS selected as cysteine modification, and with the search option 'emphasis on biological modifications' checked.
[00372] EXAMPLE 5 - FALSE DISCOVERY RATE ANALYSIS
[00373] A special plug-in, PSPEP version 0.997 (unsupported product, courtesy of Applied Biosystems) (13), was installed on the inventors' ProteinPilot software. When activated in conjunction with a ProteinPilot protein identification search, this plug-in generated a reverse decoy database and concatenated it to the normal, forward database. The samples were then searched against the combined entries of these two databases. Following the usual ProteinPilot search, a report was automatically generated as an Excel file, calculating the FDR at the peptide and protein levels. Two FDR values were calculated: the global (or average) value, and the local (or instantaneous) value.
[00374] EXAMPLE 6 - ITERATIVE RUNS
[00375] Using the peptide summary report generated by ProteinPilot for a specific set of samples, the m/z values and RP elution times of the identified ions (peptides) were imported into the exclusion list of the inventors' acquisition method. A tool in Perl language, developed in-house, helped in sorting the peptides and removed most of the redundant entries in the peptide summary report.
Tolerance windows were set at ± 120 ppm for m/z and ± 720 s for elution time. The dynamic exclusion list was maintained for a 30-s time window after the first MS/MS scan at any particular m/z ratio. Initially, the inventors used the report for the analysis of all 25 fractions to generate a single exclusion list comprising m/z values of peptides and their elution times, before starting the second iteration. The report for the second iteration of the same fractions was, in turn, used to generate a second list that was added to the first exclusion list before the third iteration was performed. After the second iTRAQ set was analyzed, the inventors altered the strategy from using all fractions to generate exclusion lists, to generating exclusion lists from groups of five fractions. This allowed for a more efficient process, with a second group of five fractions being acquired during the computer analysis of the first group. This approach not only maximized the use of available mass spectrometer and computer time, but also limited the time between the three RP separations of the same fraction by the chromatographic column. This latter point reduced the risk of a shift in the elution time due to column aging, or worse: replacement following a column failure.
[00376] A final data analysis was performed on the complete series of the 25 data files to generate a data report encompassing all 25 fractions of each iTRAQ set. This analysis of all 25
fractions as a single group enhanced the number of positive identifications and the identification confidence, by regrouping different peptides of the same protein scattered across many fractions. Iterative iTRAQ results were compared using an Excel-based program from Applied Biosystems (noncommercial product, courtesy of Dr. Sean Seymour). The protein reports were entered into this Excel program; specialized macros then compared the identified proteins to a master list, determining which proteins were on this list, and which were not. By using the results from the first iteration as the master list, the inventors were able to determine how many proteins the second and third iterations added to the global list of identified proteins
[00377] EXAMPLE 7 - COMPARISON OF NORMAL PROLIFERATIVE vs. EmCa iTRAQ RATIOS
[00378] Statistical analyses of differential expressions of the iTRAQ ratios in cancerous and noncancerous samples were performed as follows. First, all proteins in all runs were matched by accession numbers with the help of a script written in Matlab (version 7.7.0.471 ). Second, all expression ratios were then screened to determine those that were significantly different from one (P=0.05). Third, expression ratios of proteins in each sample were averaged across different runs and the mean expression ratios of proteins were evaluated. Fourth, differential expression of the iTRAQ ratios for proteins in normal and EmCa samples was verified by t-test (P=0.05). Using the mean expression ratios and the result of the t-test, differentially expressed proteins were sorted according to the following criteria: (1 ) they must have been identified in at least half of the EmCa samples; (2) differential expression was signified by a t-test or by iTRAQ ratios that were > 1.5 or < 0.67 in at least half of the samples in which the protein was identified. Ideally, this trend among the samples was uniform across all samples for each protein; to validate a trend, no more than one outlier was accepted per protein. Proteins that had been found differentially expressed in the inventors' previous study (12) were also specifically selected for comparison. The Ingenuity Pathways Analysis software, version 7.5 (Ingenuity Systems, Redwood City, CA), was used to interrogate a database of literature on the proteins of interest. Maps of protein interaction network were generated using this software.
[00379] EXAMPLE 8 - SAMPLES AND REAGENTS
[00380] Endometrial tissue samples were retrieved from the University Health Network (UHN) tissue bank and from an in-house, dedicated, research endometrial tissue bank, as described elsewhere (9a). Procurement of patient samples and tissue-banking procedures were all approved by the Research Ethics Boards of York University and the UHN. Briefly, samples were taken from hysterectomy specimens (procured with patient's consent) that had been flash-frozen in liquid nitrogen within 20 min of devitalization. Samples were first sectioned for histological diagnoses. For proteomics analyses, tissues from the mirror faces of the histological sections were each washed three times in ~1 ml of phosphate buffer saline (PBS, 1X, pH = 7.2) containing a mixture of protease inhibitors, as described previously (1 mM 4-(2-aminoethyl) benzenesulfonyl fluoride, 10 μ Μ leupeptin,
1 μ9ΛηΙ aprotinin, and 1 μ Μ pepstatin) (8a). Washed tissue samples were homogenized in 0.5 ml of PBS containing protease inhibitors using a hand-held homogenizer. These homogenates were flash- frozen in liquid nitrogen and stored at -80°C until further use. The present study was carried out using 22 tissue samples including endometrioid carcinomas (n = 6), clear cell carcinomas (n = 5), serous carcinomas (n = 8a), and normal proliferative samples (n = 3). A pooled reference sample was created by mixing 100 μg of protein lysates, retrieved from 10 additional normal proliferative samples analyzed earlier (9a). Tissue homogenates were clarified by centrifugation, and the protein concentrations were determined using a Bradford assay (Bio-Rad, Hercules, CA).
[00381] To minimize potential tag-related bias in the analysis, iTRAQ labels were assigned randomly to the pooled reference sample and tissue homogenates (n = 22) (Table 15). Each iTRAQ set included one pooled reference sample and any three samples of the different types, with the exception of the last two sets that were each composed of one reference sample and only two individual tissue samples. For each sample, 100 μg of protein lysate were digested using sequencing grade trypsin (Promega, Madison, VVI) and labeled individually with an iTRAQ tag, as described earlier (9a). These labeled samples were mixed in sets of four, with each set containing one label of each type, resulting in a total of eight sets (Table 15).
[00382] EXAMPLE 9 - STRONG CATION EXCHANGE SEPARATION CONDITIONS
[00383] Each set of labeled samples was separated by SCX fractionation using an HP1050 HPLC instrument (Agilent, Palo Alto, CA) with a 2.1-mm internal diameter x 100-mm-length PolyLC
Polysulfoethyl A column packed with 5-μηι beads with 300-A pores (The Nest Group, Southborough, MA). A 2.1-mm-internal diameter x 10-mm-length guard column of the same material was fitted immediately upstream of the analytical column. Separation was performed as described previously (8a). Briefly, each pooled sample set was diluted with the loading buffer (Buffer A: 15 mM KH2P04 in 25% acetonitrile, pH 3.0) to a total volume of 1.8 ml, and the pH was adjusted to 3.0 with phosphoric acid. Samples were then filtered using a 0.45-μηι syringe filter (Millipore, Cambridge, ON, Canada) before loading onto the column. Separation was performed using a linear binary gradient over 1 h, plus 30 min of column re-equilibration (Table 9a). Buffer A was identical in composition to the loading buffer; Buffer B was Buffer A containing 350 mM potassium chloride. Fractions were collected every
2 min using an SF-2120 Super Fraction Collector (Advantec MFS, Dublin, CA) after an initial wait of 2 min to accommodate the void volume. This resulted in a total of 30 SCX fractions per sample set. These fractions were dried by using speed-vac (Thermo Savant SC1 10 A, Holbrook, NY) and re- dissolved in a minimal volume of 5% methanol in 0.1 % formic acid, typically 8-30 μΙ_ for each fraction.
[00384] EXAMPLE 10 - LC-MS/MS RUN CONDITIONS
[00385] The SCX fractions 6 to 25 of each iTRAQ-labeled set were analyzed by on-line nanoflow / microcapillary-LC-MS/MS using the LC Packings Ultimate instrument (Amsterdam, The Netherlands) fitted with a 10-μί sample loop. The autosampler was used in the microliter pick-up mode. For each
sample, one microliter was loaded onto a 5-mm RP C18 precolumn (LC Packings) at 25 μΙ_/ηιίη and washed for 4 min before switching the precolumn in line with the separation column. The separation column was a 75^m-internal diameter x 150-mm-length capillary column (Integrafrit capillary from New Objective, VVoburn, MA) packed in-house with 3.5-μηι C18 beads with 100-A pores from Kromasil (Akzo Nobel/EKA Chemicals Inc. , NY). The flow rate used for separation on the RP column was 200 nl_ / min. Gradient elution was performed with solvent A (5% methanol in 0.1 % formic acid) and solvent B (95% methanol in 0.1 % formic acid); details of the gradient are given in Table 9b. Samples were analyzed on a Q-STAR Pulsar mass spectrometer (Applied Biosystems / MDS SCIEX, Foster City, CA) in information-dependent acquisition mode, with the scan cycles set up to perform a 1-s MS scan followed by five MS/MS scans of the five most abundant peaks for 2 s each, with a dynamic exclusion period of 30 s. Performance of the analytical system, including sensitivity, resolution, and mass accuracies, was evaluated and re-optimized at minimum once every 48 h (and more frequently, when necessary), using 80 fmol of a bovine serum albumin tryptic digest.
[00386] EXAMPLE 1 1 - PROTEIN IDENTIFICATION
[00387] The MS/MS spectra were processed by ProteinPilot version 2.0.1 , using the Paragon algorithm (12a), and the search was performed against a concatenated Swiss-Prot / Panther database of 66,082 distinct human protein entries (132, 164 entries after addition of the reverse database), with M MTS selected as the cysteine modification, and the search option 'emphasis on biological modifications' checked. The protein identification was performed simultaneously on five consecutive fraction acquisitions, using ProteinPilot unused score > 1.31 as the limit (confidence threshold of 95%), for creating the exclusion list. After all fractions of a given iTRAQ set had been acquired, a final search was initiated on all 20 fraction acquisitions in order to increase the number of positive identifications and the identification confidence by regrouping different peptides of the same protein scattered across different fractions. For this search, the Proteomics System Performance Evaluation Pipeline (PSPEP) version 0.997, a special feature of ProteinPilot, was employed (13a). When activated in conjunction with a ProteinPilot protein identification search, this plug-in generates a reverse decoy database and concatenates it to the normal database. The samples were then searched against the entries of these two databases. Following the usual ProteinPilot search, an Excel file report was automatically generated, calculating the false discovery rate (FDR) at the spectrum identification, peptide grouping, and protein grouping levels. The local (or instantaneous) FDR value of 5% was used as a threshold for protein identification. This value is more accurate than the global FDR value that is usually calculated, and, therefore, improves the confidence level of protein identifications (13a).
[00388] EXAMPLE 12 - ITERATIVE RUNS
[00389] ProteinPilot software was used to generate a report of peptides identified after every analysis of five fractions. This report provided the m/z values of the identified ions and elution times of the peptides to be imported into the exclusion list of the inventors' acquisition method. Using the
Perl® language, an in-house tool was developed to sort peptides and remove redundant peptide entries in the summary report, as discussed above. The tolerance window was set at ± 120 ppm for m/z and ± 720 s (6 min) for elution time. The dynamic exclusion list was maintained for a 30-s time window after the first MS/MS scan at any particular m/z ratio. The report generated from the second iteration of these fractions was used to generate a new list of m/z values to be added to the first exclusion list before initiation of the third iteration. This approach allowed for a more efficient process, with a second set of five fractions being acquired during software analysis of the first set. Iterative iTRAQ results were compared using custom macros in Excel files from Applied Biosystems (noncommercial product). Using results from the first iteration as a master list, these macros permitted the inventors to determine the additional number of proteins identified in the second and third iterations.
[00390] EXAMPLE 13 - COMPARISON OF NORMAL PROLIFERATIVE vs. CARCINOMA
[00391] Statistical analysis of iTRAQ ratios was carried out to determine differentially expressed proteins in endometrial cancer and non-cancerous tissue samples. iTRAQ ratios for proteins identified by accession numbers in Matlab (version 7.7.0.471 ) were screened to sort significantly different proteins (p < 0.05). These ratios were normalized using the ratio for cytoplasmic beta-actin as the control. The mean expression ratio of a given protein in each sample was averaged across different runs. The mean expression ratios were used to identify differentially expressed proteins using the following criteria: (1 ) the protein must have been identified in at least three of the samples for each EmCa subtype; (2) at least three of the expression ratios should be < 0.67 or > 1.5 in one subtype; and (3) only one outlier is accepted (e.g. , one ratio < 0.67 along an otherwise up-regulated trend).
[00392] Ingenuity Pathways Analysis, version 7.5 (Ingenuity Systems, Redwood City, CA), was used to interrogate a proprietary database built from published literature materials on the proteins of interest. Protein name abbreviations used herein are the HUGO identifiers and are the same abbreviations used in that database.
[00393] DISCUSSION OF RESULTS
[00394] Results from Examples 1 to 7
[00395] Provided below is a summary of the results obtained by the inventors in connection with the experiments of Examples 1 -7. Without being bound by theory, the results obtained in the experiments of Examples 1-7 are also discussed below.
[00396] Identified Proteins. A total of 1529 proteins or protein groups were identified below the ProteinPilot® 5% error threshold from the seven iTRAQ sets. The first iterative analyses of all of the fractions identified a total of 1137 non-redundant proteins, with the second and third analyses contributing the remaining 392. Of the 1529 proteins, 623 were identified by a single peptide; 423 of these were > 99% in confidence. The remaining 906 proteins were identified, on average, with 6.5
peptides per protein, considering only peptides with > 95% confidence scores. Of the 1529 proteins, 1260 (82%) had iTRAQ ratios reported for EmCa samples. The distribution in cellular functions (Table 3) and the total number of proteins identified were comparable to those in a previous study (12) (1529 vs. 1388 proteins, respectively), although the inventors worked with only half as much starting material (100 μg/sample instead of the 200 μg/sample used previously). These attributes strongly suggest that the current methodology works at least as well as, if not better than, the previous protocol (12).
[00397] Iterative Analysis. As will be outlined later, iterative analysis in combination with the use of an exclusion list led to confirmation of a panel of four biomarker candidates identified previously (12, 13) and identification of additional biomarker candidates. The exclusion list improved the number of protein identifications, although a core set of peptides was detected in all runs of any given sample due to shifts in retention time, peak tailing, multiple charges, and modifications (e.g., de-amidation and methionine oxidation). Following the analysis of the first iTRAQ set, the inventors improved the precision of fraction injection and widened the exclusion windows for both time and m/z, from ± 5 to ± 7 min, and from 100 mDa to 120 ppm, to mitigate some of these challenges. The inventors chose not to exclude ions based on differences of charge or modification, as this would have increased the exclusion list beyond a practical size; the inventors also reasoned that some redundancy based on differences of charge or modification may increase the confidence of identification. Thus, the inventors' iterative analyses struck a balance between the depth of analysis and tractability.
[00398] On average, the second analysis added 78% more peptides to those identified after the first analysis, while the third analysis added 36% more peptides. Within a given set, about 12% of the peptides were common to any two successive analyses; however, only 3 to 6% of the peptides were identified in all three iterations. These additional peptides were expected to improve the coverage of already identified proteins. However, this identification via multiple peptides meant that the number of identified proteins increased at a slower pace, as shown in Table 4. Indeed, the second iteration added 34% new proteins on average, while the third iteration added only 14% to the combined list from iterations 1 and 2; thus, there was little incentive to perform further iterations. 40% of the proteins identified during the first run were also identified in the next two iterations (Table 4).
[00399] Endometrial Cancer Biomarker Candidates. Differentially expressed proteins that passed the criteria detailed in Example 7 are considered as EmCa biomarker candidates and are listed in Table 5a. Of the 40 biomarker candidates, 38 were identified with a minimum of two peptides. The two exceptions are included because they were identified with > 99% in confidence and manual inspection showed excellent MS/MS spectral quality. Since the EmCa and normal proliferative tissues were sampled from different individuals - and, therefore, were not identical (i.e., the analyses were not replicates) - the typical concept of standard deviation in quantitative analysis may be misleading as a measure of analytical quality. The inventors have opted to express the distributions of combined analytical and individual variability in the following manner: in the analyses, 28% of individual iTRAQ
values deviate within ± 10% from the means, 54% within ± 20%, and 88% within ± 50%. These distributions support the inventors' hypothesis that a 50% change in iTRAQ ratios is indicative of differential expression.
[00400] Two of the proteins listed in Table 5a have previously been identified by the inventors as potential EmCa biomarkers (12, 13): alpha-1-antitrypsin (AAT) and macrophage capping protein (CAPG). Manual checking of the data for previously identified biomarker candidates proteins yielded pyruvate kinase (PK) and creatinase kinase B (CKB) with expression ratio changes just under 50% (PK, 1.43; and CKB, 0.69) in this study, but meeting the other two criteria (vide supra). Indeed, there is a group of 16 proteins in this category that satisfy almost all of the criteria; these are listed in Table 5b, and may also be considered as EmCa biomarker candidates. The potentials of AAT and PK as EmCa biomarkers have already been verified on an independent cohort of 148 patients by means of immunohistochemistry on a tissue microarray format (20). Observation of differential expressions for these four proteins (AAT, CAPG, CKB, and PK) further confirms that these proteins are, indeed, excellent biomarker candidates. A number of proteins in Table 5b also complement some of the differentially expressed proteins in Table 5a. For example, type I keratin 18 (KRT18, Table 5b) usually forms a dimer with type I I keratin 8 (KRT8, Table 5a); overexpression of both proteins accords with this expectation. Pathway analyses for the differentially expressed proteins in Tables 5a and 5b generated the interaction networks shown in FIGs. 1 -3. Significantly, these proteins appear to group around known key proteins involved in tumorigenesis (e.g. , p53 and interleukins in FIG. 1 , ERK and NF kappa B in FIG. 2, and tumor necrosis factor and Erbb2 in FIG. 3).
[00401] Many of the differentially expressed proteins identified by the inventors have been associated with other forms of cancers. Notably, cathepsin B (CTSB), which is overexpressed in EmCa, is known to favor tumorigenesis. CTSB and other cathepsins have been found upregulated in human cervical and mouse pancreatic cancers (21 , 22). It has been shown that knocking out CTSB in a mouse model of pancreatic islet cell carcinogenesis can result in reduced tumor and blood vessel formation (22). Phosphorylation of eukaryotic translation initiation factor 2 subunit 1 (EIF2S1 ) is an important pivotal event for both cell death and NFKB pathway activation. When phosphorylated, this protein will inhibit the translation of a number of non-essential genes, while promoting the expression of GADD153, which will eventually result in cell cycle arrest (23). EIF2S1 is also the target of various stress-specific kinases, and its phosphorylation will lead to the activation of NFKB (21 ). Astrocytic phosphoprotein PEA-15 is known for inhibiting many apoptosis signaling cascades (25). Its overexpression has been demonstrated to increase the susceptibility of skin cells to tumor-inducing chemicals (22).
[00402] Three calcium-binding proteins are listed in Table 5a. S100 calcium-binding protein A6 (S100A6, calcyclin) is known to be upregulated in pancreatic cancer (26) and has also been suggested to be a prognostic marker of aggressive forms of brain medulloblastoma (27). S100A6 inhibition has been shown to reduce cancer cell proliferation (26). Likewise, calcyphosin (CAPS),
which may have a role in ion transport regulation, and calumenin (CALU), which is involved in protein folding and sorting in the endoplasmic reticulum, have been found to be involved in ovarian carcinoma in humans and esophageal carcinoma in rats, respectively (28, 29).
[00403] As mentioned earlier, KRT18 and KRT8, which are known to associate, provide an example in which the differential expressions of two proteins are mutually consistent. The KRT18/8 dimer has been shown to be involved in cell signaling in rat epithelial cells, as it binds to 14-3-3 proteins and also interacts with Raf kinase (30, 31 ). Other proteins are less directly connected to tumorigenesis, because they are involved in cell motility and cytoskeletal formation - although, like keratin above, they may also be involved in the regulation of cell signaling cascades. F-actin capping protein subunit beta (CAPZB) and macrophage capping protein (CAPG), both of which control cell motility by binding to actin filaments (32), were upregulated in the inventors' study. Villin 2 (EZR, Table 5b), which has similar actin-binding functions, is upregulated as well. However, not all actin- binding proteins showed the same trend in this study. Two other actin-binding proteins, the filamins alpha (FLNA) and gamma (FLNC), were found to be down regulated. The function of one of the filamins is to crosslink actin filaments; changes in filamin expression may reflect changes in the cancer cell's cytoskeletal integrity.
[00404] Proteins acting on the extracellular matrix were also found differentially expressed in EmCa. Downregulation of alpha-1-antitrypsin (AAT or serpin A1 ) was found to be associated with EmCa (12, 13), malignant lymphoma, and liver, lung, stomach, bladder, and gall-bladder cancers (33, 34). Serpin H1 , also known as heat-shock protein 47 kD (hsp47), is a collagen-binding chaperone that is generally co-expressed with collagen precursors (35). A number of proteins are linked to inflammation and the activation of T-cells. One notable upregulated protein is interleukin enhancer- binding factor 3 (ILF3 or NF90), which improves interleukin 2 (IL2) production in T-cells by stabilizing IL2 Mrna (36). ILF3 has been found upregulated in breast cancer (37). Another such upregulated protein that is linked to inflammation is annexin A1 (ANXA1 , Table 5b). The gene encoding ANXA1 has been found upregulated in intrahepatic cholangiocarcinoma (38). By contrast, it was found to be downregulated in a variety of other cancers, including head and neck, prostate, breast, and larynx (39). Interestingly, the last study also reported that annexin A1 is upregulated in mast cells, which are attracted by tumors and infiltrate cancer tissues where they may either impede or promote tumor growth, depending upon the local conditions (39). Annexin A1 participates in the local inflammatory regulation, and, therefore, in the immune response to the tumor. In the case of laryngeal cancer, annexin A1 may result from invasion of the tumor by mast cells or similar partners of the immune system.
[00405] Fibrinogen (all 3 chains), haptoglobin, and apoliproteins A-l and A-l l were found downregulated in EmCa samples. Fibrinogen is a blood protein, the synthesis of which is positively regulated by an area of inflammation-related cytokines, including interleukin 1 , interleukin 6, and tumor necrosis factor alpha (TNF) (40). TNF also stimulates the adhesion of neutrophils to fibrinogen-
covered cell surface (41 ), most likely by increasing the expression of interleukin 8 (42). By contrast, fibrinogen chains are synthesized by cancer cells in cervical and breast carcinomas (43). Haptoglobin (HP) is another blood protein, a peptidase involved in free plasma hemoglobin degradation. Its expression and blood concentration are promoted by inflammatory-related cytokines (40), notably by interleukine-1 beta via the protein Myd88 in mouse (44). This protein has been shown to be downregulated in hepatocarcinoma (45) and in AFP-negative hepatoblastoma (46); however, it was upregulated in the more aggressive AFP-positive hepatoblastoma (46) and in Her-2/neu breast cancer (47). The inventors' finding that HP is downregulated in Type I EmCa, which is typically less invasive than Type I I EmCa, is in line with these previous studies. Similarly, apolipoprotein A-l (APOA1 ) was found to be overexpressed in high-grade dysplastic cervical cells (48), but underexpressed in the serum of ovarian cancer patients (49). Alteration of the level of APOA1 may be a part of the host inflammatory response to the tumor, but the basis for this change is unclear. The other lipoprotein, apolipoprotein A-l I (APOA2), is known to be upregulated in alpha fetoprotein (AFP)-positive hepatoblastoma, following the same pattern as HP (46). Upregulation of annexin A1 , together with the downregulations of apolipoproteins, fibrinogens, and haptglobin, could be indicative of suppression of the inflammatory process in tissues surrounding the tumor.
[00406] One of the more interesting proteins in Table 5b is heterogeneous nuclear
ribonucleoprotein A1 (HNRNPA1 ), which was found to be upregulated. HNRNPA1 is a protein that is phosphorylated in response to T-cell activation, and, in turn, controls TNF expression (50).
HNRNPA1 also binds DNA telomeres and promotes telomere elongation. Preventing HNRNPA1 binding to telomeres leads to the apoptosis of tumor cells (51 ). As telomere degradation in fast- proliferative cells eventually triggers p53-dependent cell-cycle arrest, it has been postulated that HNRNPA1 action results in cancer cells bypassing cell cycle arrest mechanisms (51 ).
[00407] Two other proteins of interest in Table 5b are the glycolytic enzymes, pyruvate kinase (PK) and lactate dehydrogenase A (LDHA). PK catalyzes the conversion of phosphoenol pyruvate (PEP) to pyruvate and generates ATP from ADP in the process. PK expresses as a number of isoforms, with M 1 and M2 being splice variants of a single gene (52). The M1 isoform is expressed in skeletal muscle and brain tissue, while the M2 isoform is found in fetal tissue and proliferating cells, especially cancer cells. The presence of the M2 isoform of PK is critical during the process of tumorigenesis. The shift in metabolism from oxidative phosphorylation to aerobic glycolysis, along with lactate production, is a common feature of tumor cells, known as the Warburg effect (52). In addition to the inventors' studies that identified overexpression of PKM2 in EmCa (12, 19, 53), other studies also revealed increased expression of PKM2 in lung, pancreatic, breast, oral, renal, and ovarian cancers and in various gastrointestinal malignancies (54-60). Thus, the overexpression of LDHA is generally consistent with that of PKM2. LDHA itself has been found to be upregulated by cDNA analyses in both human pulmonary adenocarcinoma and a mouse model of lung cancer (61 ) and in hepatoblastoma (46). Concomitant upregulations of LDHA and PK, such as those observed
here in EmCa, have previously been observed in liver cancer (38). This trend may reflect increased activity of the glycolysis pathway toward aerobic glycolysis in cancer cells.
[00408] Results from Examples 8 to 13
[00409] Provided below is a summary of the results obtained by the inventors in connection with the experiments of Examples 8-13. Without being bound by theory, the results obtained in the experiments of Examples 8-13 are also discussed below.
[00410] Protein Identification. A total of 1518 non-redundant proteins were identified across the eight iTRAQ sets with a Local FDR value < 5%. Of these, 1068 were identified by multiple peptides with > 95% confidence scores, at an average of 8.3 peptides per protein. Of the remaining 450 proteins identified by a single peptide, 41 1 peptides showed > 99% confidence scores. Of all proteins identified, 1 194 (79%) were reported with iTRAQ ratios; however, only 732 (48%) had iTRAQ ratios where the p-value was < 0.05, and these, therefore, were eligible for further analyses. Notably, in Examples 1-7 above, where a similar iTRAQ / iterative-analysis approach was used to compare Type I EmCa with normal proliferative endometrial tissues, the inventors obtained about as many proteins (1529) from slightly fewer tissue samples (20 samples); there was a similar proportion of proteins having iTRAQ ratios (81 %), but with a lower number of ratios determined with high confidence (32%, p < 0.05). The lower number of identified proteins in the current study was probably a consequence of using the more stringent Local FDR, as calculated by the PSPEP plug-in, instead of the less stringent confidence threshold provided by ProteinPilot.
[00411] On average, the second analysis identified an additional 44% unique peptides on top of the peptides identified in the first analysis; the third iteration added a further 27% unique peptides to the two combined analyses. Eight percent of the peptides were observed in all three analyses despite the use of an exclusion list. These repeated identifications were due to shifts in retention times, peak tailing, multiple charge states, and modifications (e.g., deamidation and methionine oxidation) (10a). Table 7 shows the number of unique proteins identified in the eight sample sets by the iterative analysis. The second and third analyses of the same set identified, on average, 45% and 15% additional proteins, respectively. One notable outlier is set 8, for which the first analysis was sub- optimal. As expected, the second and third analyses compensated for the first and provided higher numbers of protein identifications, thereby demonstrating one of the advantages of an iterative approach: viz., as each sample is analyzed several times, this approach is self-compensating and consequently results in a more rounded performance.
[00412] Differentially expressed proteins. All proteins reported in Tables 8 and 9 were identified with five or more peptides. As noted earlier, proteins which showed at least 50% differential expression in a minimum of three samples in at least one of the subtypes, and which were found in at least three samples of each of the other subtypes, were deemed to be of interest for further examination. For proteins listed in Table 8, 31 % of individual iTRAQ values deviated within ± 10%
from the mean values, 59% within ± 20%, and 94% within ± 50%. For proteins in Table 9, these distributions were 22%, 54%, and 88%, respectively. It should be noted that these variations include both analytical and individual differences. The distributions support the inventors' position that a 50% change in iTRAQ ratios is indicative of differential expression.
[00413] Two types of differentially expressed proteins were identified. Table 8 shows the average expression ratios for the differentially upregulated and downregulated proteins in the three subtypes of high-grade EmCa, along with the number of samples in which each given expression was confidently determined. The maximum numbers are: endometrioid, 6; clear cell, 5; serous, 8; and (individual) normal proliferative, 3. The 11 reported upregulated and 7 downregulated proteins should permit discrimination among the endometrioid, clear cell, and serous EmCa subtypes, particularly when used as a panel.
[00414] A second list of differentially expressed proteins consisting of those displaying consistent trends across all EmCa samples, regardless of histological type, is shown in Table 9. These 14 proteins are potential biomarkers that permit the discrimination of EmCa from normal endometrium. L-lactate dehydrogenase A (LDHA), phosphoglycerate kinase 1 (PGK1 ), and pyruvate kinase (PK) have already been found to be upregulated in one or more previous iTRAQ studies on EmCa tissues (8a-10a). The results here provide additional verification. Similarly, alpha-1 -antitrypsin and apolipoprotein A1 from Table 8, and serum albumin and transferrin from Table 9, were also found downregulated in one or more previous studies (8a-10a). It should be noted that the last five are blood proteins: their lower concentrations in cancer tissues more likely result from tumors being differently vascularized from healthy tissues, rather than the proteins being underexpressed in the tumors themselves.
[00415] The vast majority of the proteins that may eventually permit subtyping of high-grade EmCa (Table 8) have already been linked to cancer. For example, elevated levels of heterogeneous nuclear ribonucleoproteins A1 and A2/B1 (HNRNPA1 and HNRNPA2B1 , respectively) observed in the endometrioid samples correlate well with elevated levels found in studies on other forms of cancer, including breast, small-cell lung, and ovarian carcinomas (14a). This trend is consistent with the known function of these proteins, since protein HNRNPA1 and its splice variants A2/B1 bind telomeres to protect them, and thereby promote cancer cell survival and proliferation by ensuring telomere stabilization. Knocking down the expression of HNRNPA1 and its variants by means of siRNA in cancer cells leads to increased cell death (14a).
[00416] Another protein listed in Table 8 that promotes cell survival is nucleophosmin (NPM 1 ). This nucleolar protein is involved in many functions related to embryonic development and genome stability (15a). Its suppression favors tumorigenesis, but its overexpression promotes cell proliferation and cell survival. Notably, NPM 1 overexpression not only results in re-localization of the p53 protein into the nucleus, but also inhibits p53 accumulation in the mitochondria (16a). Since p53-dependent apoptosis pathways rely on p53 transactivating mitochondrial genes and promoting other changes in
mitochondrial activities, this described interaction of NPM1 with p53 is, in effect, anti-apoptotic. It is, therefore, logical to find NPM 1 overexpressed in many human cancers (16a). NPM1 can be posttranslationally modified in a variety of ways, with the acetylated form being the one involved in anti-apoptotic activities (15a).
[00417] Other upregulated proteins in Table 8 have also been linked to cancer. The 10 kDa heat shock protein (chaperonin 10, HSPE1 ) has been found upregulated in colon cancer (17a); it has also consistently been found to be upregulated in endometrioid carcinoma (8a, 9a). Hepatoma-derived growth factor (HDGF) was found to stimulate proliferation of cultured fibroblasts. Its physiological functions are mostly unstudied. However, it has been shown that HDGF is important for cell differentiation and, moreover, is overexpressed in colorectal cancer (18a). Nucleoside diphosphate kinase B (NME2) expression is progesterone-dependent, and reportedly has anti-tumorigenic properties in ovarian cancer: it inhibits the growth of ovarian cancer cell lines and also suppresses cell motility (19a). Upregulation of this protein in EmCa suggests that it is also involved in EmCa, but apparently not in a tumor suppressor capacity. Peroxiredoxin-5 (PRDX5) is expressed in response to oxidative stress, and has a protective role during inflammation. Silencing PRDX5 by RNA interference leads to an increase of caspase activity in mitochondria and cell apoptosis (20a). This could be a consequence of the increase of oxidative stress, rather than a direct interaction between PRDX5 and caspase proteins. Cystatin B (CSTB) is a cystein proteinase inhibitor, and may actually have a role in cancer: it may act against cancer proliferation and metastasis by inhibiting extracellular matrix degradation by proteinases like cathepsin B (21 a). Its expression in mouse uterus is increased by estradiol (22a). Since estrogen receptors are known to be overexpressed in EmCa (23a-25a), the overexpression of CSTB may be linked to this event. Inorganic pyrophosphatase (PPA1 ) is only loosely related to cancer: it has been shown to bind to p53 (26a), but the physiological meaning of this interaction is totally unknown. Given the importance of p53 in EmCa (see below), PPA1 may be a valuable candidate for further study.
[00418] Other proteins have been linked to a distinct feature in the metabolism of cancer cells rather than cell proliferation or apoptosis. Herein, PK has again been observed to be upregulated in all EmCa subtypes (see Table 9). This is typical of the Warburg effect, characterized by the overexpression of pyruvate kinase M2 and the shift of the glycolytic pathway toward lactate production under aerobic conditions, a feature which has been found in many cancer cells (27a). In this light, the upregulation of other proteins related to glucose metabolism is consistent with the Warburg effect. Notably, glucose-6 phosphate isomerase, alpha-enolase, and PGK1 were upregulated in all subtypes (Table 9). LDHA is also upregulated in the endometrioid and clear cell subtypes (Table 8). LDHA has previously been shown to be upregulated in human and murine models of lung carcinoma (28a). Since the function of this enzyme is to convert pyruvate into lactate, its upregulation is to be expected following the upregulation of PK. It was initially puzzling that LDHA was not found to be upregulated in the serous subtype; upon a closer inspection of the data, it became apparent that the iTRAQ ratio
was, in fact, above 1.5 in four serous carcinoma samples, although close to 1 for the remaining four samples.
[00419] In terms of established specific genetic alterations that are known to be associated with the EmCa subtypes (23a-25a), inactivation of the tumor suppressor PTEN is the most often reported alteration. This inactivation is common in estrogen-dependent endometrioid EmCa, generally in conjunction with defects of DNA mismatch repair genes, and is believed to be an early event in carcinogenesis. Less frequent, but still specific to the endometroid subtype, are mutations in the Kras and/or beta-catenin oncogenes. By contrast, an early event in estrogen-independent serous carcinomas is apparently a mutation and overexpression of the protein p53, a central regulator of cell cycle and differentiation. Another important alteration in serous EmCa is the mutation and amplification of HER-2/neu. None of the protein products of these genes was detected in the current study; these are all expected to have low abundances and are located in the nucleus or membranes, compartments that are poorly sampled in current proteomic technologies.
[00420] Pathway analyses by means of IPA showed that the differentially expressed proteins in Tables 8 and 9 (see FIGs. 4 and 5, respectively) interact with key oncogenes and/or their protein products. The potential biomarkers entered are shown in red (ove rex pressed) and green
(underexpressed), while the interacting proteins discovered by IPA are colorless. Many of these discovered proteins are situated as hubs (interacting with and linking numerous proteins) and are known to play key roles in tumorigenesis - e.g. , p53 (TP53), v-myc myelocytomatosis viral oncogene homolog (MYC), NFKB, interleukins, and tumor necrosis factor (TNF). FIG. 4 was constructed by merging five interacting networks, the top functions of which include cell cycle and death; cancer; molecular transport; cellular development, growth, and development; and signal transduction. FIG. 5 comprises a single network centering on cell death and cellular growth and proliferation. Adding PTEN, Kras, beta-catenin, p53, HER-2/neu, and other oncogenes and proteins (23a-25a) to the differentially expressed proteins in Tables 8 and 9 in IPA produced bigger networks, which confirmed that the interactions are complex and that the differentially expressed proteins identified herein are acted upon or acting on these genes and proteins, either directly or indirectly.
[00421] While the foregoing invention has been described in some detail for purposes of clarity and understanding, it will be appreciated by one skilled in the art, from a reading of the disclosure, that various changes in form and detail can be made without departing from the true scope of the invention in the appended claims. The invention is not to be limited in scope by the specific embodiments described herein, since such embodiments are intended as but single illustrations of one aspect of the invention, and any functionally equivalent embodiments are within the scope thereof. Indeed, various modifications of the invention, in addition to those shown and described herein, will become apparent to those skilled in the art from the foregoing description and
accompanying drawings. Such modifications are intended to fall within the scope of the appended claims.
[00422] All publications, patents, and patent applications referred to herein are incorporated by reference in their entirety to the same extent as if each individual publication, patent, or patent application was specifically and individually indicated to be incorporated by reference in its entirety. Nothing herein is to be construed as an admission that the invention is not entitled to antedate the cited references by virtue of prior invention.
[00423] Set out below are full citations for the references cited herein.
(1 ) LaBaer, J. So, You Want to Look for Biomarkers (Introduction to the Special Biomarkers Issue). J. Proteome Res. 2005, 4, 1053-1059.
(2) Ebert, M. P. A. ; Korc, M. ; Malfertheiner, P. ; Rocken, C. Advances, Challenges, and Limitations in Serum-Proteome-Based Cancer Diagnosis. J. Proteome Res. 2006, 5, 19-25.
(3) Gloerich, J. ; Wevers, R. A. ; Smeitink, J. A. M. ; van Engelen, B. G. ; van den Heuvel, L. P.
Proteomics Approaches to Study Genetic and Metabolic Disorders. J. Proteome Res. 2007, 6, 506- 512.
(4) Lescuyer, P. ; Hochstrasser, D.; Rabilloud, T. How Shall We Use the Proteomics Toolbox for Biomarker Discovery? J. Proteome Res. 2007, 6, 3371 -3376.
(5) Good, D. M. ; Thongboonkerd, V. ; Novak J. ; Bascands, J.-L ; Schanstra, J. P. ; Coon J. J.;
Dominiczak, A. ; Mischak, H. Body Fluid Proteomics for Biomarker Discovery: Lessons from the Past Hold the Key to Success in the Future. J. Proteome Res. 2007, 6, 4549-4555.
(6) Ross, P. L ; Huang, Y. N. ; Marchese, J. N. ; Williamson, B.; Parker, K. ; Hattan, S. ; Khainovski, N. ; Pillai, S. ; Dey, S.; Daniels, S. ; Purkayastha, S. ; Juhasz, P. ; Martin, S. ; Bartlet-Jones, M. ; He, F. ; Jacobson, A. ; Pappin, D. J. Multiplexed Protein Quantitation in Saccharomyces cerevisiae Using Amine-reactive Isobaric Tagging Reagents. Mol. Cell. Proteomics 2004, 3, 1 154-1 169.
(7) Wang, N. ; Li, L. Exploring the Precursor Ion Exclusion Feature of Liquid Chromatography- Electrospray Ionization Quadrupole Time-of-Flight Mass Spectrometry for Improving Protein
Identification in Shotgun Proteome Analysis. Anal. Chem. 2008, 80, 4696-4710.
(8) Scherl, A. ; Francois, P. ; Converset, V. ; Bento, M. ; Burgess, J. A. ; Sanchez, J.-C ; Hochstrasser, D. F.; Schrenzel, J. ; Corthals, G. L. Nonredundant mass spectrometry: A strategy to integrate mass spectrometry acquisition and analysis. Proteomics 2004, 4, 917-927.
(9) Chen, H.-S. ; Rejtar, T. ; Andreev, V. ; Moskovets, E. ; Karger, B. L. Enhanced Characterization of Complex Proteomic Samples Using LC-MALDI MS/MS: Exclusion of Redundant Peptides from MS/MS Analysis in Replicate Runs. Anal. Chem. 2005, 77, 7816-7825.
(10) Canadian Cancer Society, National Cancer Institute Of Canada, Statistics Canada,
Provincial/Territorial Cancer Registries, Public Health Agency Of Canada: Canadian Cancer Statistics 2009. Available on the Canadian Cancer Society website, URL:
http://www.cancer.ca/Ontario/About%20cancer/Cancer%20statistics/Canadian%20Cancer%20Statisti cs.aspx?sc_lang=en&r=1
(11 ) Di Cristofano, A. ; Ellenson, L.H. Endometrial Carcinoma. Annu. Rev. Pathol. Mech. Dis. 2007, 2, 57-85.
(12) DeSouza, L. V. ; Grigull, J. ; Ghanny, S.; Dube, V. ; Romaschin, A. D. ; Colgan, T. J. ; Siu, K. W. M. Endometrial Carcinoma Biomarker discovery and verification using differentially tagged clinical
samples with multidimensional liquid chromatography and tandem mass spectrometry. Mol. Cell. Proteomics 2007, 6, 1170-1182.
(13) DeSouza, L. V.; Diehl, G.; Rodrigues, M. J.; Guo, J.; Romaschin, A. D.; Colgan, T. J.; Siu, K. W. M. Search for Cancer Markers from Endometrial Tissues Using Differentially Labeled Tags iTRAQ and cICAT with Multidimensional Liquid Chromatography and Tandem Mass Spectrometry. J.
Proteome Res.2005, 4, 377-386.
(14) Shilov, I. V.; Seymour, S. L; Patel, A. A.; Loboda, A.; Tang, W. H.; Keating, S. P.; Hunter, C. L; Nuwaysir, L. M.; Schaeffer, D. A. The Paragon Algorithm, a Next Generation Search Engine That Uses Sequence Temperature Values and Feature Probabilities to Identify Peptides from Tandem Mass Spectra. Mol. Cell. Proteomics 2007 , 6, 1638-1655.
(15) Ralhan, R.; DeSouza, L. V.; Matta, A.; Tripathi, S. C; Ghanny, S.; Datta Gupta, S.; Bahadur,
5. ; Siu, K. W. M. Discovery and Verification of Head-and-Neck Cancer Biomarkers by Differential Protein Expression Analysis using iTRAQ-labeling and Multidimensional Liquid Chromatography and Tandem Mass Spectrometry. Mol. Cell. Proteomics 2008, 7, 1162-1173.
(16) Ralhan, R.; DeSouza, L. V.; Matta, A.; Tripathi, S. C; Ghanny, S.; Gupta, S. D.; Thakar, A.; Chauhan, S. S.; Siu, K. W. M. iTRAQ-Multidimensional Liquid Chromatography and Tandem Mass Spectrometry based Identification of Potential Biomarkers of Oral Epithelial Dysplasia and Novel Networks between Inflammation and Premalignancy. J. Proteome Res.2009, 8, 300-309.
(17) Siu, K. W. M.; DeSouza, L. V.; Scorilas, A.; Romaschin, A. D.; Honey, R. J.; Stewart, R.; Pace, K.; Youssef, Y.; Chow, T. F. F.; Yousef, G. M. Differential Protein Expressions in Renal Cell Carcinoma: New Biomarker Discovery by Mass Spectrometry. J. Proteome Res.2009, 8, 3797-3807.
(18) Mukherjee, J.; DeSouza, L. V.; Micaleff, J.; Karim, Z.; Croul, S.; Siu, K. W. M.; Guha, A. Loss of Collapsin Response Mediator Protein 1 (CRMP1) as Detected by iTRAQ Analysis Promotes Invasion of Human Gliomas Expressing Mutant EGFRvlll. Cancer Res.2009, 69, 8555-8562.
(19) DeSouza, L. V.; Taylor, A. M.; Li, W.; Minkoff, M. S.; Romaschin, A. D.; Colgan, T. J.; Siu, K. W. M. Multiple Reaction Monitoring of mTRAQ-Labeled Peptides Enables Absolute Quantification of Endogenous Levels of a Potential Cancer Marker in Cancerous and Normal Endometrial Tissues. J. Proteome Res.2008, 7, 3525-3534.
(20) Dube, V.; Grigull, J.; DeSouza, L. V.; Ghanny, S.; Colgan, T. J.; Romaschin, A. D.; Siu, K. W. Verification of endometrial tissue biomarkers previously discovered using mass spectrometry-based proteomics by means of immunohistochemistry in a tissue microarray format. J Proteome Res.2007,
6, 2648-2655.
(21) Vazquez-Ortiz, G.; Pina-Sanchez, P.; Vazquez, K.; Duenas, A.; Taja, L; Mendoza, P.; Garcia, J. A.; Salcedo, M. Overexpression of cathepsin F, matrix metalloproteinases 11 and 12 in cervical cancer. BMC Cancer 2005, 5(1), 68-78.
(22) Gocheva, V.; Zeng, W.; Ke, D.; Klimstra, D.; Reinheckel, T.; Peters, C; Hanahan, D.; Joyce, J. A. Distinct roles for cysteine cathepsin genes in multistage tumorigenesis. Genes Dev.2006, 20(5), 543- 556.
(23) Oyadomari, S.; Mori, M. Roles of CHOP/GADD153 in endoplasmic reticulum stress. Cell Death Differ.2004, 11, 381-389.
(24) Jiang, H. Y.; Wek, S. A.; McGrath, B. C; Scheuner, D.; Kaufman, R. J.; Cavener, D. R.; Wek, R. C. Phosphorylation of the alpha subunit of eukaryotic initiation factor 2 is required for activation of NF- kappaB in response to diverse cellular stresses. Mol. Cell. Biol.2003, 23, 5651-5663.
(25) Formisano, P.; Perruolo, G.; Libertini, S.; Santopietro, S.; Troncone, G.; Raciti, G. A.; Oriente, F.; Portella, G.; Miele, C; Beguinot, F. Raised expression of the antiapoptotic protein ped/pea-15 increases susceptibility to chemically induced skin tumor development. Oncogene 2005, 24(47), 7012-7021.
(26) Ohuchida, K.; Mizumoto, K.; Ishikawa, N.; Fujii, K.; Konomi, H.; Nagai, E.; Yamaguchi, K.;
Tsuneyoshi, M.; Tanaka, M. The role of S100A6 in pancreatic cancer development and its clinical implication as a diagnostic marker and therapeutic target. Clin. Cancer Res.2005, 11(21), 7785-7793.
(27) Pelc, K.; Vincent, S.; Ruchoux, M. M.; Kiss, R.; Pochet, R.; Sariban, E.; Decaestecker, C;
Heizmann, C. W. Calbindin-d(28k): a marker of recurrence for medulloblastomas. Cancer 2002, 95(2), 410-419.
(28) Hibbs, K.; Skubitz, K. M.; Pambuccian, S. E.; Casey, R. C; Burleson, K. M.; Oegema, T. R.; Thiele, J. J.; Grindle, S. M.; Bliss, R. L; Skubitz, A. P. Differential gene expression in ovarian carcinoma: identification of potential biomarkers. Am. J. Pathol.2004, 165(2), 397-414.
(29) Nishida, K.; Mine, S.; Utsunomiya, T.; Inoue, H.; Okamoto, M.;. Udagawa, H.; Hanai, T.; Mori, M. Global Analysis of Altered Gene Expressions during the Process of Esophageal Squamous Cell Carcinogenesis in the Rat: A Study Combined with a Laser Microdissection and a cDNA Microarray. Cancer Res.2005; 65(2), 401-409.
(30) Ku, N. O.; Liao, J.; Omary, M. B. Phosphorylation of human keratin 18 serine 33 regulates binding to 14-3-3 proteins. EMBO J.1998, 17(7), 1892-1906.
(31) Ku, N. O.; Fu, H.; Omary, M. B. Raf-1 activation disrupts its binding to keratins during cell stress. J. Cell Biol.2004, 166(4), 479-485.
(32) Nakagawa, H.; Liyanarachchi, S.; Davuluri, R. V.; Auer, H.; Martin, E. W.; De La Chapelle, A.; Frankel, W. L. Role of cancer-associated stromal fibroblasts in metastatic colon cancer to the liver and their expression profiles. Oncogene 2004, 23(44), 7366-7377.
(33) Sun, Z.; Yang, P. Role of imbalance between neutrophil elastase and alpha 1 -antitrypsin in cancer development and progression. Lancet Oncol.2004, 5, 182-190.
(34) Ryu, J. W.; Kim, H. J.; Lee, Y. S.; Myong, N. H.; Hwang, C. H.; Lee, G. S.; Yom, H. C. The proteomics approach to find biomarkers in gastric cancer. J. Korean Med. Sci.2003, 18, 505-509.
(35) Hirata, H.; Yamamura, I.; Yasuda, K.; Kobayashi, A.; Tada, N.; Suzuki, M.; Hirayoshi, K.;
Hosokawa, N.; Nagata, K. Separate Cis-acting DNA Elements Control Cell Type- and Tissue-specific Expression of Collagen Binding Molecular Chaperone HSP47. J. Biol. Chem.1999, 274, 35703- 35710.
(36) Shi, L; Godfrey, W. R.; Lin, J.; Zhao, G.; Kao, P. N. NF90 regulates inducible IL-2 gene expression in T cells. J. Exp. Med.2007, 204(5), 971-977.
(37) Tripathi, A.; King, C; de la Morenas, A.; Perry, V. K.; Burke, B.; Antoine, G. A.; Hirsch, E. F.; Kavanah, M.; Mendez, J.; Stone, M.; Gerry, N. P.; Lenburg, M. E.; Rosenberg, C. L. Gene expression abnormalities in histologically normal breast epithelium of breast cancer patients. Int. J. Cancer 2008, 122(7), 1557-1566.
(38) Wang, A.-G.; Yoon, S. Y.; Oh, J.-H.; Jeon, Y.-J.; Kim, M.; Kim, J.-M.; Byun, S.-S.; Yang, J. O.; Kim, J. H.; Kim, D.-G.; Yeom, Y.-l.; Yoo, H.-S.; Kim, Y. S.; Kim, N.-S. Identification of intrahepatic cholangiocarcinoma related genes by comparison with normal liver tissues using expressed sequence tags. Biochem. Biophys. Res. Comm.2006, 345, 1022-1032.
(39) Silistino-Souza, R.; Rodrigues-Lisoni, F. C; Cury, P. M.; Maniglia, J. V.; Raposo, L. S.; Tajara, E. H.; Christian H. C; Oliani, S. M. Annexin 1: differential expression in tumor and mast cells in human larynx cancer. Int. J. Cancer 2007, 120, 2582-2589.
(40) Mansouri, R. M.; Bauge, E.; Staels, B.; Gervois, P. Systemic and Distal Repercussions of Liver- Specific Peroxisome Proliferator-Activated Receptor-{alpha} Control of the Acute-Phase Response. Endocrinology 2008, 149(6), 3215-3223.
(41) Loike, J. D.; Sodeik B, Cao, L; Leucona, S.; Weitz, J. I.; Detmers, P. A.; Wright, S. D.;
Silverstein, S. C. CD11c/CD18 on neutrophils recognizes a domain at the N terminus of the A alpha chain of fibrinogen. Proc. Natl. Acad. Sci. USA 1991, 88(3), 1044-1048.
(42) Oppenheim, J. J.; Zachariae, C. O.; Mukaida, N.; Matsushima, K. Properties of the novel proinflammatory supergene "intercrine" cytokine family. Annu. Rev. Immunol.1991, 9, 617-648.
(43) Rybarczyk B. J.; Simpson-Haidaris P. J. Fibrinogen Assembly, Secretion, and Deposition into Extracellular Matrix by MCF-7 Human Breast Carcinoma Cells. Cancer Res.2000, 60, 2033-2039.
(44) Adachi, O.; Kawai, T.; Takeda, K.; Matsumoto, M.; Tsutsui, H.; Sakagami, M.; Nakanishi, K.; Akira, S. Targeted disruption of the MyD88 gene results in loss of IL-1- and IL-18-mediated function. Immunity 1998, 9(1), 143-150.
(45) Lee, l-N.; Chen, C.-H.; Sheu, J.-C; Lee, H.-S.; Huang, G.-T.; Yu, C.-Y.; Lu, F.-J.; Chow, L.-P. Identification of Human Hepatocellular Carcinoma-Related Biomarkers by Two-Dimensional Difference Gel Electrophoresis and Mass Spectrometry. J. Proteome Res.2005, 4, 2062-069.
(46) Yamada, S.-l.; Ohira, M.; Horie, H.; Ando, K.; Takayasu, H.; Suzuki, Y.; Sugano, S.; Hirata, T.; Goto, T.; Matsunaga, T.; Hiyama, E.; Hayashi, Y.; Ando, H.; Suita, S.; Kaneko, M.; Sasaki, F.;
Hashizume, K.; Ohnuma, N.; Nakagawara, A. Expression profiling and differential screening between hepatoblastomas and the corresponding normal livers: identification of high expression of the PLK1 oncogene as a poor-prognostic indicator of hepatoblastomas. Oncogene 2004, 23, 5901-5911.
(47) Zhang, D.; Tai, L. K.; Wong, L. L; Chiu, L.-L; Sethi, S. K.; Koay, E. S. C. Proteomic Study Reveals That Proteins Involved in Metabolic and Detoxification Pathways Are Highly Expressed in HER-2/neu-positive Breast Cancer. Mol. Cell. Proteomics 2005, 4, 1686-1696.
(48) Gu, Y.; Wu, S.-L.; Meyer, J. L; Hancock, W. S.; Burg, L. J.; Linder, J.; Hanlon, D. W.; Karger, B. L. Proteomic Analysis of High-Grade Dysplastic Cervical Cells Obtained from ThinPrep Slides Using Laser Capture Microdissection and Mass Spectrometry. J. Proteome Res.2007, 6, 4256-4268.
(49) Zhen Zhang, Z.; Bast, Jr., R. C; Yu, Y.; Li, J.; Sokoll, L. J.; Rai, A. J.; Rosenzweig, J. M.;
Cameron, B.; Wang, Y. Y.; Meng, X.-Y.; Berchuck, A.; van Haaften-Day, C; Hacker, N. F.; de Bruijn, H. W. A.; van der Zee, A. G. J.; Jacobs, I. J.; Eric, T. Fung, E. T.; Chan, D. W. Three Biomarkers Identified from Serum Proteomic Analysis for the Detection of Early Stage Ovarian Cancer. Cancer Res.2004, 64, 5882-5890.
(50) Buxade, M.; Parra, J. L; Rousseau, S.; Shpiro, N.; Marquez, R.; Morrice, N.; Bain, J.; Espel, E.; Proud, C. G. The Mnks Are Novel Components in the Control of TNFalpha Biosynthesis and Phosphorylate and Regulate hnRNP A1. Immunity 2005, 23(2), 177-189.
(51) Patry, C; Bouchard, L; Labrecque, P.; Gendron, D.; Lemieux, B.; Toutant, J.; Lapointe, E.; Wellinger, R.; Chabot, B. Small interfering RNA-mediated reduction in heterogeneous nuclear ribonucleoparticule A1/A2 proteins induces apoptosis in human cancer cells but not in normal mortal cell lines. Cancer Res.2003, 63(22), 7679-7688.
(52) Mazurek, S.; Boschek, C. B.; Hugo, F.; Eigenbrodt, E. Pyruvate kinase type M2 and its role in tumor growth and spreading. Semin. Cancer Biol.2005, 15, 300-308.
(53) DeSouza, L. V.; Romaschin, A. D.; Colgan, T. J.; Siu, K. W. M. Absolute Quantification of Potential Cancer Markers in Clinical Tissue Homogenates Using Multiple Reaction Monitoring on a Hybrid Triple Quadrupole/Linear Ion Trap Tandem Mass Spectrometer. Anal. Chem.2009, 81, 3462- 70.
(54) Koukourakis, M. I.; Giatromanolaki, A.; Bougioukas, G.; Sivridis, E: Lung cancer: a comparative study of metabolism related protein expression in cancer cells and tumor associated stroma. Cancer Biol. Ther.2007, 6, 1476-1479.
(55) Cerwenka, H. Tumor M2-pyruvate kinase and pancreatic cancer. Pancreas 2008, 37, 221-222.
(56) Isidoro, A. ; Casado, E. ; Redondo, A. ; Acebo, P.; Espinosa, E. ; Alonso, A. M. ; Cejas, P. ;
Hardisson, D. ; Fresno Vara, J. A. ; Belda-lniesta, C ; Gonzalez-Baron, M. ; Cuezva, J. M: Breast carcinomas fulfill the Warburg hypothesis and provide metabolic markers of cancer prognosis.
Carcinogenesis 2005, 26, 2095-2104.
(57) Wong, T. S. ; Liu, X. B. ; Chung-Wai Ho, A. ; Po-Wing Yuen, A. ; Wai-Man Ng, R. ; Ignace Wei, W. Identification of pyruvate kinase type M2 as potential oncoprotein in squamous cell carcinoma of tongue through microRNA profiling. Int. J. Cancer 2008, 123, 251 -257.
(58) Weinberger, R. ; Appel, B. ; Stein, A. ; Metz, Y. ; Neheman, A. ; Barak, M. The pyruvate kinase isoenzyme M2 (Tu M2-PK) as a tumour marker for renal cell carcinoma. Eur. J. Cancer Care (Engl)
2007, 16, 333-337.
(59) Ahmed, A S. ; Dew, T. ; Lawton, F. G. ; Papadopoulos, A. J. ; Devaja, O. ; Raju, K. S. ; Sherwood, R. A. Tumour M2-PK as a predictor of surgical outcome in ovarian cancer, a prospective cohort study. Eur. J. Gynaecol. Oncol. 2007, 28, 103-108.
(60) Hardt. P. D. ; Ewald, N. Tumor M2 pyruvate kinase: a tumor marker and its clinical application in gastrointestinal malignancy. Expert Rev. Mol. Diagn. 2008, 8, 579-5W85.
(61 ) Stearman, R. S. ; Dwyer-Nield, L ; Zerbe, L ; Blaine, S. A.; Chan, Z. ; Bunn, Jr. , P. A. ; Johnson, G. L ; Hirsch, F. R. ; Merrick, D. T. ; Franklin, W. A. ; Baron, A. E. ; Reith, R. L ; Nemenoff, R. A. ; Malkinson, A. M. ; Geraci, M. W. Analysis of Orthologous Gene Expression between Human Pulmonary
Adenocarcinoma and a Carcinogen-Induced Murine Model. Am. J. Path. 2005, 767, 1763-1775.
(1 a). Canadian Cancer Society, National Cancer Institute Of Canada, Statistics Canada,
Provincial/Territorial Cancer Registries, Public Health Agency Of Canada. Canadian Cancer Statistics
2008. Available on the Canadian Cancer Society website, URL:
http://www.cancer.ca/Ontario/About%20cancer/Cancer%20statistics/Canadian%20Cancer%20Statisti cs.aspx?sc_lang=en&r=1
(2a). Di Cristofano, A. , and Ellenson, L. H. (2007) Endometrial Carcinoma. Annu. Rev. Pathol.
Mech. Dis. 2, 57-85.
(3a). Soslow, R. A. , Bissonnette, J. P. , Wilton, A., Ferguson, S. E. , Alektiar, K. M. , Duska, L. R. , and Oliva E. (2007) Clinicopathologic analysis of 187 high-grade endometrial carcinomas of different histologic subtypes: similar outcomes belie distinctive biologic differences. Am. J. Surg. Pathol. 31 (7), 979-987.
(4a). LaBaer, J. (2005) So, You Want to Look for Biomarkers (Introduction to the Special Biomarkers Issue). J. Proteome Res. 4, 1053-1059.
(5a). Ebert, M. P. A. , Korc, M. , Malfertheiner, P. , and Rocken, C. (2006) Advances, Challenges, and Limitations in Serum-Proteome-Based Cancer Diagnosis. J. Proteome Res. 5, 19-25.
(6a). Lescuyer, P. , Hochstrasser, D., and Rabilloud, T. (2007) How Shall We Use the Proteomics Toolbox for Biomarker Discovery? J. Proteome Res. 6, 3371-3376.
(7a). Good, D. M. , Thongboonkerd, V. , Novak J. , Bascands, J.-L , Schanstra, J. P. , Coon J. J. , Dominiczak, A. , and Mischak, H. (2007) Body Fluid Proteomics for Biomarker Discovery: Lessons from the Past Hold the Key to Success in the Future. J. Proteome Res. 6, 4549-4555.
(8a). DeSouza, L. V. , Diehl, G. , Rodrigues, M. J. , Guo, J. , Romaschin, A. D. , Colgan, T. J. , and Siu, K. W. M. (2005) Search for Cancer Markers from Endometrial Tissues Using Differentially Labeled Tags iTRAQ and cICAT with Multidimensional Liquid Chromatography and Tandem Mass
Spectrometry. J. Proteome Res. 4, 377-386.
(9a). DeSouza, L. V., Grigull, J. , Ghanny, S. , Dube, V. , Romaschin, A. D., Colgan, T. J., and Siu, K. W. M. (2007) Endometrial Carcinoma Biomarker discovery and verification using differentially tagged
clinical samples with multidimensional liquid chromatography and tandem mass spectrometry. Mol. Cell. Proteomics 6, 1170-1182.
(10a). Voisin SN, Krakovska O, Matta A, DeSouza LV, Romaschin AD, Colgan TJ, Siu KWM. (2011) Identification of Novel Molecular Targets for Endometrial Cancer using a Drill-Down LC-MS/MS approach with iTRAQ. PLoS ONE.6(1):e16352.
(11a). Ross, P. L, Huang, Y. N., Marchese, J. N., Williamson, B., Parker, K., Hattan, S., Khainovski, N., Pillai, S., Dey, S., Daniels, S., Purkayastha, S., Juhasz, P., Martin, S., Bartlet-Jones, M., He, F., Jacobson, A., Pappin, D. J. (2004) Multiplexed Protein Quantitation in Saccharomyces cerevisiae Using Amine-reactive Isobaric Tagging Reagents. Mol. Cell. Proteomics 3, 1154-1169.
(12a). Shilov, I. V., Seymour, S. L, Patel, A. A., Loboda, A., Tang, W. H., Keating, S. P., Hunter, C. L, Nuwaysir, L. M., and Schaeffer, D. A. (2007) The Paragon Algorithm, a Next Generation Search Engine That Uses Sequence Temperature Values and Feature Probabilities to Identify Peptides from Tandem Mass Spectra. Mol. Cell. Proteomics 6, 1638-1655.
(13a). Tang, W. H., Shilov, I. V., and Seymour, S. L. (2008) Nonlinear Fitting Method for Determining Local False Discovery Rates from Decoy Database Searches. J. Proteome Res.7 (9), 3661-3667.
(14a). Patry, C, Bouchard, L, Labrecque, P., Gendron, D., Lemieux, B., Toutant, J., Lapointe, E., Wellinger, R., and Chabot, B. (2003) Small Interfering RNA-Mediated Reduction in Heterogeneous Nuclear Ribonucleoparticule A1/A2 Proteins Induces Apoptosis in Human Cancer Cells but not in Normal Mortal Cell Lines. Cancer Res.63, 7679-7688.
(15a). Shandilya, J., Swaminathan, V., Gadad, S. S., Choudhari, R., Kodaganur, G. S., Kundu, T. K. (2009) Acetylated NPM1 localizes in the nucleoplasm and regulates transcriptional activation of genes implicated in oral cancer manifestation. Mol. Cell. Biol., 29 (18), 5115-27.
(16a). Dhar, S. K., St Clair, D. K. (2009) Nucleophosmin Blocks Mitochondrial Localization of p53 and Apoptosis. J. Biol. Chem., 284 (24), 16409-18.
(17a). Gavert, N., Sheffer, M., Raveh, S., Spaderna, S., Shtutman, M., Brabletz, T., Barany, F., Paty, P., Notterman, D., Domany, E., and Ben-Ze'ev, A. (2007) Expression of L1-CAM and ADAM10 in Human Colon Cancer Cells Induces Metastasis. Cancer Res.67 (16), 7703-7712.
(18a). Lepourcelet, M., Tou, L, Cai, L, Sawada, J., Lazar, A. J., Glickman, J. N, Williamson, J. A., Everett, A. D., Redston, M., Fox, E. A., Nakatani, Y., Shivdasani, R. A. (2005) Insights into developmental mechanisms and cancers in the mammalian intestine derived from serial analysis of gene expression and study of the hepatoma-derived growth factor (HDGF). Development, 132 (2), 415-27.
(19a). Syed, V., Mukherjee, K., Lyons-Weiler, J., Lau, K. M., Mashima, T., Tsuruo, T., Ho, S. M. (2005) Identification of ATF-3, caveolin-1, DLC-1, and NM23-H2 as putative antitumorigenic, progesterone-regulated genes for ovarian cancer cells by gene profiling. Oncogene, 24 (10), 1774-87.
(20a). De Simoni, S., Goemaere, J., Knoops, B. (2008) Silencing of peroxiredoxin 3 and peroxiredoxin 5 reveals the role of mitochondrial peroxiredoxins in the protection of human neuroblastoma SH-SY5Y cells toward MPP+. Neurosci. Lett.433 (3), 219-24.
(21a). Schenkels, L. C, Veerman, E. C, Nieuw Amerongen, A. V. (1995) Biochemical composition of human saliva in relation to other mucosal fluids. Crit. Rev. Oral Biol. Med.6 (2), 161-75.
(22a). Watanabe, H., Suzuki, A., Kobayashi, M., Lubahn, D. B., Handa, H., Iguchi, T. (2003) Similarities and differences in uterine gene expression patterns caused by treatment with physiological and non-physiological estrogens. J. Mol. Endocrinol.31 (3), 487-97.
(23a). Doll, A., Abal, M., Rigau, M., Monge, M., Gonzalez, M., Demajo, S., Colas, E., Llaurado, M., Alazzouzi, H., Planaguma, J., Lohmann, M.A., Garcia, J., Castellvi, S., Ramon y Cajal, J., Gil-Moreno,
A., Xercavins, J., Alameda, F., Reventos, J. (2008) Novel molecular profiles of endometrial cancer— new light through old windows. J. Steroid Biochem. Mol. Biol.108, 221-229.
(24a). Engelsen, I. B., Akslen, L. A., and Salvesen, H. B. (2009) Biologic markers in endometrial cancer treatment. APMIS 117, 693-707.
(25a). Bansal, N., Yendluri, V., and Wenham, R. M. (2009) The Molecular Biology of Endometrial Cancers and the Implications for Pathogenesis, Classification, and Targeted Therapies. Cancer Control 16 (1), 8-13.
(26a). Stelzl, U., Worm, U., Lalowski, M., Haenig, C, Brembeck, F. H., Goehler, H., Stroedicke, M., Zenkner, M., Schoenherr, A., Koeppen, S., Timm, J., Mintzlaff, S., Abraham, C, Bock, N., Kietzmann, S., Goedde, A., Toksoz, E., Droege, A., Krobitsch, S., Korn, B., Birchmeier, W., Lehrach, H., Wanker, E. E. (2005) A human protein-protein interaction network: a resource for annotating the proteome. Ce//122 (6), 957-68.
(27a). Christofk, H. R., Heiden, M. G. V., Harris, M. H., Ramanathan, A., Gerszten, R. E., Wei, R., Fleming, M. D., Schreiber, S. L, and Cantley, L. C. (2008) The M2 splice isoform of pyruvate kinase is important for cancer metabolism and tumor growth. Nature 452, 230-234.
(28a). Stearman, R. S., Dwyer-Nield, L, Zerbe, L, Blaine, S. A., Chanm Z., Bunn, P. A., Johnson, G. L, Hirsch, F. R., Merrick, D. T., Franklin, W. A., Baron, A. E., Keith, R. L, Nemenoff, R. A., Malkinson, A. M., Geraci, M. W. (2005) Analysis of Orthologous Gene Expression between Human Pulmonary Adenocarcinoma and a Carcinogen-Induced Murine Model. Am. J. Pathol., 167 (6), 1763-75.
(29a). Dube, V., Grigull, J., DeSouza, L. V., Ghanny, S., Colgan, T. J., Romaschin, A. D., Siu, K. W. M. (2007) Verification of Endometrial Tissue biomarkers previously discovered using mass spectrometry-based proteomics by means of immunohistochemistry in a tissue microarray format. J. Proteome Res.6, 2648-55.
Table 1. iTRAQ labeling of endometrial tissue samples.
Table 2. LC gradients for SCX and RP separation in the Type I EmCa analysis-
Buffer A: 15 mM KH2P04 in 25% acetonitrile, pH 3.0
Buffer B: Buffer A containing 350 mM KCI
Buffer C: Buffer A containing 1 M KCI
Solvent A: 5% acetonitrile in 0.1 % formic acid
Solvent B: 95% acetonitrile in 0.1 % formic acid.
Table 3. Distribution of protein functions for biomarkers identified in the Type I EmCa
analysis.
Table 4. Comparison of identified protein numbers across the iterative analysis of iTRAQ sets for the Type I EmCa analysis.
Total
proteins Additional proteins Addition Common
Common to Common to
All runs Iter 1 Iter 2 Iter 3 all iterations Iter 2 / 1 Iter 3 / 1+2 all iterations
Set 1 888 787 40 61 122 5.1 % 7.4% 15.5%
Set 2 635 434 122 79 162 28.1 % 14.2% 37.3%
Set 3 520 408 14 98 136 3.4% 23.2% 33.3%
Set 4 207 148 51 8 48 34.5% 4.0% 32.4%
Set 5 263 161 49 53 91 30.4% 25.2% 56.5%
Set 6 558 227 265 66 141 116.7% 13.4% 62.1 %
Set 7 554 412 81 61 190 19.7% 12.4% 46.1 %
Average: 34.0% 14.3% 40.5%
Table 5a. EmCa biomarker candidates.
Columns also display the number of samples in which iTRAQ ratios have been obtained (# samples, out of 10).
P02774 Vitamin D-binding protein 7 0.58 8 1.02
Table 5b. Additional EmCa biomarker candidates.
Table 6. LC gradients for SCX and Ci» separation in the EmCa subtype analysis.
Buffer A: 15 mM KH2P04 in 25% acetonitrile, pH 3.0
Buffer B: Buffer A containing 350 mM KCI
Buffer C: Buffer A containing 1 M KCI
Solvent A: 5% acetonitrile in 0.1 % formic acid
Solvent B: 95% acetonitrile in 0.1 % formic acid
Table 7. Number of proteins identified by the iterative method in the EmCa subtype analysis.
Total Additional
proteins proteins Common Addition Common
All runs Iter 1 Iter 2 Iter 3 to all Iter 2 / 1 Iter 3 / 1+2 to all
Set 1 606 376 186 44 214 49% 8% 35%
Set 2 647 404 116 127 277 29% 24% 43%
Set 3 694 529 64 101 273 12% 17% 39%
Set 4 629 433 169 27 242 39% 4% 38%
Set 5 334 261 0 73 54 0% 28% 16%
Set 6 569 358 128 83 247 36% 17% 43%
Set 7 746 517 173 56 310 33% 8% 42%
Set 8 529 180 285 64 159 158% 14% 30%
Average: 45% 15% 36% Median 36% 15% 39%
Table 8. Proteins expressing a divergent trend between carcinoma samples.
Table 9. Proteins showing a similar trend in all carcinoma samples.
** Pool : Normal pooled samples; E1 -E6 : Endometroid ; C1-C5 ; Clear cell carcinoma ; S1-S8: Serous cell carcinoma ; N 1-N3: Normal proliferative endometrium samples.
Table 11. Comparison of iTRAQ mean ratios for proteins identified in three different studies.
* 'N' means that this value has not been statistically confirmed, generally due to lack of sufficient quantification events.
Claims
1. A method for detecting one or more endometrial cancer markers in a subject, comprising:
(a) obtaining a sample from a subject;
(b) detecting in the sample an amount of at least one of the protein endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9 or a polynucleotide encoding same; and
(c) comparing the detected amount with an amount detected for a standard.
2. A method for diagnosing endometrial cancer in a subject, comprising comparing:
(a) levels of at least one of the protein endometrial cancer markers listed in Table 5, Table 5b, Table 8, and/or Table 9, or a polynucleotide encoding same, detected in a sample from the subject; and
(b) normal levels of expression of the corresponding protein marker or polynucleotide in a control sample,
wherein a significant difference in levels of the protein marker or polynucleotide, relative to the corresponding normal levels, is indicative of the endometrial cancer in the subject.
3. The method of claim 1 or 2, wherein the polynucleotide is mRNA.
4. A method for diagnosing endometrial cancer in a subject, comprising:
(a) contacting a biological sample obtained from the subject with at least one binding agent that specifically binds to at least one of the protein endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9 or a polynucleotide encoding same; and
(b) detecting in the sample amounts of the protein marker or polynucleotide that bind to the at least one binding agent, relative to a predetermined standard or cut-off value, and thereby determining the presence or absence of the endometrial disease in the subject.
5. The method of claim 4, wherein the binding agent is an antibody.
6. A method of screening a subject for endometrial cancer, comprising:
(a) obtaining a biological sample from a subject;
(b) detecting in the sample an amount of at least one of the protein endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9 or a polynucleotide encoding same; and
(c) comparing the detected amount of the protein marker or polynucleotide with a predetermined standard, wherein detection of a level of the protein marker or polynucleotide different from that of the standard is indicative of endometrial cancer.
7. The method of claim 6, wherein the level of the protein marker or polynucleotide is significantly higher than the standard and is indicative of endometrial cancer.
8. The method of claim 7, wherein the level of the protein marker or polynucleotide is significantly lower than the standard and is indicative of endometrial cancer.
9. The method of any one of claims 1-8, wherein the sample is obtained from a tissue, extract, cell culture, cell lysate, lavage fluid, or physiological fluid of the subject.
10. The method of claim 9, wherein the sample is obtained from endometrial tumor tissue.
11. A method for determining the presence or absence in a subject of an endometrial cancer marker associated with an endometrial disease, comprising:
(a) detecting in a sample from the subject at least one of the protein endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9 or a polynucleotide encoding same; and
(b) relating the detected amount of the protein marker or polynucleotide to the presence of the endometrial disease.
12. The method of claim 1 1 , wherein the polynucleotide is mRNA.
13. The method of claim 1 1 , wherein the polynucleotide is detected by:
(a) contacting the sample with at least one oligonucleotide that hybridizes to the polynucleotide; and
(b) detecting in the sample levels of at least one nucleic acid that hybridizes to the polynucleotide, relative to a predetermined standard or cut-off value, and thereby determining the presence or absence of the endometrial disease in the subject.
14. The method of claim 12 or 13, wherein the polynucleotide is detected with an amplification reaction.
15. The method of claim 14, wherein the amplification reaction comprises a polymerase chain reaction employing oligonucleotide primers that hybridize to the polynucleotide or to a complement thereof.
16. The method of claim 15 wherein the polynucleotide is detected with at least one oligonucleotide probe that hybridizes to the polynucleotide or to a complement thereof.
17. The method of claim 16, wherein the polynucleotide is detected by:
(a) isolating RNA from the sample;
(b) combining the RNA with at least one reagent, to convert the RNA to cDNA;
(c) treating the cDNA with at least one amplification reaction reagent and at least one primer that hybridizes to the cDNA, to produce at least one amplification product; (d) analyzing the at least one amplification product to detect an amount of RNA encoding the at least one protein endometrial cancer marker; and
(e) comparing the amount of RNA to an amount detected against a panel of expected values for normal tissue derived using similar primers.
18. A method for diagnosing or monitoring prognosis of endometrial cancer in a subject, comprising:
(a) obtaining a tissue sample from the subject; and
(b) detecting in the sample the level of at least one of the protein endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9 or a polynucleotide encoding same, wherein the presence of higher or lower levels of the protein marker or polynucleotide in the sample, compared to a standard or control, is indicative of the disease or prognosis.
19. A method for monitoring the progression of endometrial cancer in a subject, comprising:
(a) detecting in a sample from the subject, at a first time point, at least one of the protein endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9 or a polynucleotide encoding same;
(b) repeating step (a) at a subsequent point in time; and
(c) comparing levels detected in steps (a) and (b), and thereby monitoring the progression of endometrial cancer.
20. A method for determining in a subject whether endometrial cancer has metastasized or is likely to metastasize in the future, comprising comparing:
(a) levels of at least one of the protein endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9, or a polynucleotide encoding same, in a sample from the subject; and
(b) normal levels or non-metastatic levels of the protein marker or polynucleotide in a control sample,
wherein a significant difference between the levels in the subject sample and the normal or non-metastatic levels is indicative that the endometrial cancer has metastasized.
21. A method for assessing the aggressiveness or indolence of endometrial cancer, comprising comparing:
(a) levels of at least one of the protein endometrial cancer markers listed in Table 5a, 5b, Table 8, and/or Table 9, or a polynucleotide encoding same, in a sample from a subject; and
(b) normal levels of the protein marker or polynucleotide in a control sample, wherein a significant difference between the levels in the subject sample and normal levels is indicative that the cancer is aggressive or indolent.
22. A diagnostic composition comprising an agent that:
(i) binds to at least one of the protein endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9; or
(ii) hybridizes to at least one polynucleotide encoding at least one of the protein endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9.
23. A method for assessing the efficacy of a test agent for inhibiting endometrial cancer in a subject, comprising comparing:
(a) levels of at least one of the protein endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9, or a polynucleotide encoding same, in a first sample obtained from the subject and exposed to the test agent; and
(b) levels of at least one of the protein endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9, or a polynucleotide encoding same, in a second sample obtained from the subject, wherein the second sample has not been exposed to the test agent,
wherein a significant difference in the levels of the protein marker or polynucleotide in the first sample, relative to the second sample, is an indication that the test agent inhibits endometrial cancer in the subject.
24. A method for assessing the efficacy of a therapy for inhibiting endometrial cancer in a subject, comprising comparing:
(a) levels of at least one of the protein endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9, or a polynucleotide encoding same, in a first sample obtained from the subject; and
(b) levels of at least one of the protein endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9, or a polynucleotide encoding same, in a second sample obtained from the subject following therapy,
wherein a significant difference in the levels of expression of the protein marker or polynucleotide in the second sample, relative to the first sample, is an indication that the therapy inhibits endometrial cancer in the subject.
25. A method for selecting an agent for inhibiting endometrial cancer in a subject, comprising:
(a) obtaining a sample comprising precancer or cancer cells from the subject;
(b) separately exposing aliquots of the sample to a plurality of test agents;
(c) comparing levels of at least one of the protein endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9, or a polynucleotide encoding same, in the aliquots; and
(d) selecting one of the test agents which alters the levels of the protein marker or polynucleotide in the aliquot containing that test agent, relative to other test agents.
26. A method for inhibiting endometrial cancer in a subject, comprising:
(a) obtaining a sample comprising precancer or cancer cells from the subject;
(b) separately maintaining aliquots of the sample in the presence of a plurality of test agents;
(c) comparing levels of at least one of the protein endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9, or a polynucleotide encoding same, in the aliquots; and
(d) administering to the subject at least one of the test agents which alters the levels of the protein marker or polynucleotide in the aliquot containing that test agent, relative to other test agents.
27. A method for assessing the endometrial cancer cell carcinogenic potential of a test compound, comprising:
(a) maintaining separate aliquots of endometrial cancer cells in the presence and absence of the test compound; and
(b) comparing levels of at least one of the protein endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9, or a polynucleotide encoding same, in the aliquots, wherein a significant difference in levels of the protein marker or polynucleotide in the aliquot maintained in the presence of the test compound, relative to levels in the aliquot maintained in the absence of the test compound, is indicative that the test compound possesses endometrial cancer cell carcinogenic potential.
28. An in vivo method for imaging an endometrial disease, comprising:
(a) injecting a subject with one or more agents that bind to a protein endometrial cancer marker listed in Table 5a, Table 5b, Table 8, and/or Table 9 or a polynucleotide encoding same, wherein the agent carries a label for imaging the endometrial cancer marker;
(b) allowing the agent to incubate in vivo and bind to the protein marker or
polynucleotide; and
(c) detecting the presence of the label localized to diseased endometrial tissue.
29. The method of claim 28, wherein the agent is an antibody that specifically reacts with an endometrial cancer marker.
30. The method of any one of claims 1-29, wherein the at least one endometrial cancer marker is a protein listed in Table 5a, Table 5b, Table 8, and/or Table 9.
31. A set of endometrial cancer markers, comprising at least two of the protein markers listed in Table 5a, Table 5b, Table 8, and/or Table 9 or polynucleotides encoding same.
32. The set of markers of claim 31, wherein the at least 2 protein markers are selected from the group consisting of L-lactate dehydrogenase A (LDHA), phosphoglycerate kinase, nucleolin, glucose- 6-phosphate isomerase, alpha-enolase, villin 2, isocitrate dehydrogenase 2 mitochondrial, peroxiredoxin-1 , transketolase, serpin H1 , transferrin, galectin-1 , lumican, 40S ribosomal protein S25, diazepam binding inhibitor, adenosylhomocysteinase, alpha-2-HS-glycoprotein, alpha-2- macroglobulin, anterior gradient protein 2 homolog, apolipoprotein A-l, apolipoprotein A-l l, astrocytic phosphoprotein PEA-15, beta-2-glycoprotein 1 , calcyphosin, calumenin, cathepsin B, ceruloplasmin, chaperonin containing TCP1 subunit 7 (Eta), collagen type I alpha 1 and 2, dihydropyrimidinase-like 2, eukaryotin translation initiation factor 2 subunitl , fibrinogen alpha, beta and gamma, fibulin-1 , filamin- C, haptoglobin, keratin type II cytoskeletal 8, macrophage capping protein, myristoylated alanine-rich C-kinase substrate, retinal dehydrogenase 2, S100 A6 calcium-binding protein, secreted frizzled- related protein 4, and vitamin D binding protein.
33. A kit for carrying out the method of any one of claims 1 -32.
34. A kit for determining the presence of endometrial cancer in a subject, comprising a known amount of at least one binding agent that binds to at least one of the protein endometrial cancer markers listed in Table 5a, Table 5b, Table 8, and/or Table 9, wherein the binding agent comprises a detectable substance or binds directly or indirectly to a detectable substance.
35. A kit for determining the presence of endometrial cancer in a subject, comprising a known amount of an oligonucleotide that hybridizes to a polynucleotide encoding a protein endometrial cancer marker listed in Table 5a, Table 5b, Table 8, and/or Table 9, wherein the oligonucleotide is directly or indirectly labelled with a detectable substance.
36. Use of a marker to screen for, diagnose, or monitor the progress, treatment, recurrence, or prognosis of endometrial cancer, wherein the marker is L-lactate dehydrogenase A (LDHA), phosphoglycerate kinase, nucleolin, glucose-6-phosphate isomerase, alpha-enolase, villin 2, isocitrate dehydrogenase 2 mitochondrial, peroxiredoxin-1 , transketolase, Serpin H1 , transferrin, galectin-1 , lumican, 40S ribosomal protein S25, diazepam binding inhibitor, adenosylhomocysteinase, alpha-2- HS-glycoprotein, alpha-2-macroglobulin, anterior gradient protein 2 homolog, apolipoprotein A-l, apolipoprotein A-l l, astrocytic phosphoprotein PEA-15, beta-2-glycoprotein 1 , calcyphosin, calumenin, cathepsin B, ceruloplasmin, chaperonin containing TCP1 subunit 7 (Eta), collagen type I alpha 1 and 2, dihydropyrimidinase-like 2, eukaryotin translation initiation factor 2 subunitl , fibrinogen alpha, beta and gamma, fibulin-1 , filamin-C, haptoglobin, keratin type I I cytoskeletal 8, macrophage capping protein, myristoylated alanine-rich C-kinase substrate, retinal dehydrogenase 2, S100 A6 calcium- binding protein, secreted frizzled-related protein 4, or vitamin D binding protein, or a polynucleotide encoding same.
37. Use of differential expression of a marker to screen for, diagnose, or monitor the progress, treatment, recurrence, or prognosis of endometrial cancer, wherein the marker is L-lactate dehydrogenase A (LDHA), phosphoglycerate kinase, nucleolin, glucose-6-phosphate isomerase, alpha-enolase, villin 2, isocitrate dehydrogenase 2 mitochondrial, peroxiredoxin-1 , transketolase, Serpin H1 , transferrin, galectin-1 , lumican, 40S ribosomal protein S25, diazepam binding inhibitor, adenosylhomocysteinase, alpha-2-HS-glycoprotein, alpha-2-macroglobulin, anterior gradient protein 2 homolog, apolipoprotein A-l, apolipoprotein A-l l, astrocytic phosphoprotein PEA-15, beta-2- glycoprotein 1 , calcyphosin, calumenin, cathepsin B, ceruloplasmin, chaperonin containing TCP1 subunit 7 (Eta), collagen type I alpha 1 and 2, dihydropyrimidinase-like 2, eukaryotin translation initiation factor 2 subunitl , fibrinogen alpha, beta and gamma, fibulin-1 , filamin-C, haptoglobin, keratin type I I cytoskeletal 8, macrophage capping protein, myristoylated alanine-rich C-kinase substrate, retinal dehydrogenase 2, S100 A6 calcium-binding protein, secreted frizzled-related protein 4, or vitamin D binding protein, or a polynucleotide encoding same.
38. The method of any one of claims 1-21 and 23-30, the composition of claim 22, the set of markers of any one of claims 31 -32, or the kit of any one of claims 33-35, wherein the endometrial cancer marker is an endometrial carcinoma (EmCa) marker.
39. The method of any one of claims 1-21 and 23-30, the composition of claim 22, the set of markers of any one of claims 31 -32, or the kit of any one of claims 33-35, wherein the endometrial cancer marker is a Type I EmCa marker, an endometrioid EmCa marker, a clear cell EmCa marker, or a serous EmCa marker.
40. The method, composition, set of markers, or kit of claim 39, wherein the endometrioid EmCa marker is a heterogeneous nuclear ribonucleoprotein (hnRNPA).
41. The method, composition, set of markers, or kit of claim 39, wherein the serous EmCa marker is a mitochondrial ATP synthase.
42. An iterative computerized method for identifying at least one protein endometrial cancer marker, comprising:
(a) conducting a first mass spectrometry analysis of peptide ions in normal and diseased tissues, to identify a first set of potential markers based on differential expression;
(b) creating an exclusion list consisting of the peptide ions identified in the first set of potential markers, wherein the exclusion list is restricted by a mass window and a time window;
(c) conducting a second mass spectrometry analysis of peptide ions in the normal and diseased tissues, to identify a second set of potential markers based on differential expression, wherein the second analysis disregards peptide ions in the exclusion list; (d) updating the exclusion list to consist of the peptide ions identified in the first and second sets of potential markers; and
(e) conducting a third mass spectrometry analysis of peptide ions in the normal and diseased tissues, to identify a third set of potential markers based on differential expression, wherein the third analysis disregards peptide ions in the updated exclusion list.
43. The method of claim 42, further comprising the step of calculating a false discovery rate for each identified potential marker.
44. A system for identifying at least one protein endometrial cancer marker, the system comprising at least one computing device including software that, when executed, performs a method comprising:
(a) conducting a first mass spectrometry analysis of peptide ions in normal and diseased tissues, to identify a first set of potential markers based on differential expression;
(b) creating an exclusion list consisting of the peptide ions identified in the first set of potential markers, wherein the exclusion list is restricted by a mass window and a time window;
(c) conducting a second mass spectrometry analysis of peptide ions in the normal and diseased tissues, to identify a second set of potential markers based on differential expression, wherein the second analysis disregards peptide ions in the exclusion list;
(d) updating the exclusion list to consist of the peptide ions identified in the first and second sets of potential markers; and
(e) conducting a third mass spectrometry analysis of peptide ions in the normal and diseased tissues, to identify a third set of potential markers based on differential expression, wherein the third analysis disregards peptide ions in the updated exclusion list.
Applications Claiming Priority (2)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| US34450810P | 2010-08-10 | 2010-08-10 | |
| US61/344,508 | 2010-08-10 |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| WO2012019300A1 true WO2012019300A1 (en) | 2012-02-16 |
Family
ID=45567241
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/CA2011/050485 Ceased WO2012019300A1 (en) | 2010-08-10 | 2011-08-10 | Endometrial cancer biomarkers and methods of identifying and using same |
Country Status (1)
| Country | Link |
|---|---|
| WO (1) | WO2012019300A1 (en) |
Cited By (23)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO2014107002A1 (en) * | 2013-01-04 | 2014-07-10 | 서울대학교 산학협력단 | Lung cancer diagnostic marker |
| EP2787350A1 (en) * | 2013-04-05 | 2014-10-08 | Atlas Antibodies AB | ASRGL1 in endometrial cancer |
| US9506925B2 (en) | 2014-07-02 | 2016-11-29 | Dragon Victory Development Ltd. | Specific biomarker set for non-invasive diagnosis of liver cancer |
| EP3093343A4 (en) * | 2014-01-10 | 2017-12-27 | Juntendo Educational Foundation | Method for assessing lymph node metastatic potential of endometrial cancer |
| US9885718B2 (en) | 2014-07-02 | 2018-02-06 | Dragon Victory Development Ltd. | Specific biomarker set for non-invasive diagnosis of liver cancer |
| WO2020047636A1 (en) * | 2018-09-04 | 2020-03-12 | Centro Nacional De Pesquisa Em Energia E Materiais | Method for metastatic prognosis in oral cancer, biosensor, prognosis kit, biomarkers and their use |
| CN111500732A (en) * | 2020-05-25 | 2020-08-07 | 福建医科大学 | Application and kit of microorganisms as diagnostic markers for endometrial cancer |
| US10822307B2 (en) | 2014-08-28 | 2020-11-03 | Eisai R&D Management Co., Ltd. | High-purity quinoline derivative and method for manufacturing same |
| WO2020223657A1 (en) * | 2019-05-02 | 2020-11-05 | Predictive Technology Group, Inc. | Somatic cancer driver mutations in endometriosis lesions contribute to secondary cancer risk |
| CN113049824A (en) * | 2021-04-20 | 2021-06-29 | 首都医科大学附属北京妇产医院 | Application of apolipoprotein ApoA1 in detection of drug resistance of cervical cancer to platinum chemotherapy |
| US11090386B2 (en) | 2015-02-25 | 2021-08-17 | Eisai R&D Management Co., Ltd. | Method for suppressing bitterness of quinoline derivative |
| CN113493839A (en) * | 2021-09-06 | 2021-10-12 | 北京泱深生物信息技术有限公司 | Application of gene marker combination in diagnosis of endometrial cancer |
| CN113493840A (en) * | 2021-09-07 | 2021-10-12 | 北京泱深生物信息技术有限公司 | Marker for endometrial cancer diagnosis and derivative product and application thereof |
| CN113917149A (en) * | 2021-09-30 | 2022-01-11 | 江苏扬新生物医药有限公司 | Application of gelsolin detection substance in preparation of uterine cancer evaluation detection reagent |
| US11369623B2 (en) | 2015-06-16 | 2022-06-28 | Prism Pharma Co., Ltd. | Anticancer combination of a CBP/catenin inhibitor and an immune checkpoint inhibitor |
| CN115184611A (en) * | 2022-08-01 | 2022-10-14 | 中国医学科学院北京协和医院 | Endometrial cancer stratification related marker and application thereof |
| US11547705B2 (en) | 2015-03-04 | 2023-01-10 | Merck Sharp & Dohme Llc | Combination of a PD-1 antagonist and a VEGF-R/FGFR/RET tyrosine kinase inhibitor for treating cancer |
| US11598776B2 (en) | 2011-06-03 | 2023-03-07 | Eisai R&D Management Co., Ltd. | Biomarkers for predicting and assessing responsiveness of thyroid and kidney cancer subjects to lenvatinib compounds |
| US12220398B2 (en) | 2015-08-20 | 2025-02-11 | Eisai R&D Management Co., Ltd. | Tumor therapeutic agent |
| US12226409B2 (en) | 2017-05-16 | 2025-02-18 | Eisai R&D Management Co., Ltd. | Treatment of hepatocellular carcinoma |
| CN119936393A (en) * | 2024-12-31 | 2025-05-06 | 哈尔滨脉图精准技术有限公司 | Metabolic marker composition for early endometrial cancer diagnosis and its application |
| US12303505B2 (en) | 2017-02-08 | 2025-05-20 | Eisai R&D Management Co., Ltd. | Tumor-treating pharmaceutical composition |
| US12508313B2 (en) | 2009-08-19 | 2025-12-30 | Eisai R&D Management Co., Ltd. | Quinoline derivative-containing pharmaceutical composition |
Citations (3)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CA2550900A1 (en) * | 2003-12-23 | 2005-07-07 | Mount Sinai Hospital | Methods for detecting markers associated with endometrial disease or phase |
| CA2703795A1 (en) * | 2006-10-27 | 2008-05-02 | Mount Sinai Hospital | Endometrial phase or endometrial cancer biomarkers |
| WO2009126969A2 (en) * | 2008-04-11 | 2009-10-15 | Expression Pathology Inc. | Biomarkers for endometrial disease |
-
2011
- 2011-08-10 WO PCT/CA2011/050485 patent/WO2012019300A1/en not_active Ceased
Patent Citations (3)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CA2550900A1 (en) * | 2003-12-23 | 2005-07-07 | Mount Sinai Hospital | Methods for detecting markers associated with endometrial disease or phase |
| CA2703795A1 (en) * | 2006-10-27 | 2008-05-02 | Mount Sinai Hospital | Endometrial phase or endometrial cancer biomarkers |
| WO2009126969A2 (en) * | 2008-04-11 | 2009-10-15 | Expression Pathology Inc. | Biomarkers for endometrial disease |
Non-Patent Citations (4)
| Title |
|---|
| GRISENDI ET AL.: "Nucleophosmin and cancer", NATURE REVIEWS, vol. 6, July 2006 (2006-07-01), pages 493 - 505, XP002559721, DOI: doi:10.1038/nrc1885 * |
| LI ET AL.: "Identification of candidate biomarker proteins released by human endometrial and cervical cencer cells using two- dimensional liquid chromatography/tandem mass spectrometry", J. OF PROTEOME RESEARCH, vol. 6, no. 7, 2007, pages 2615 - 2622, XP002600810, DOI: doi:10.1021/PR0700798 * |
| MONGE ET AL.: "Proteomic approach to ETV5 during endometrial carcinoma invasion reveals a link to oxidative stress", CARCINOGENESIS, vol. 30, no. 8, 14 May 2009 (2009-05-14), pages 1288 - 1297 * |
| VOISIN ET AL.: "Identification of novel molecular targets for endometrial cancer using a drill-down LC-MS/MS approach with iTRAQ, art. e16352", PLOS ONE, vol. 6, no. 1, 31 January 2011 (2011-01-31), pages 1 - 11 * |
Cited By (29)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US12508313B2 (en) | 2009-08-19 | 2025-12-30 | Eisai R&D Management Co., Ltd. | Quinoline derivative-containing pharmaceutical composition |
| US11598776B2 (en) | 2011-06-03 | 2023-03-07 | Eisai R&D Management Co., Ltd. | Biomarkers for predicting and assessing responsiveness of thyroid and kidney cancer subjects to lenvatinib compounds |
| WO2014107002A1 (en) * | 2013-01-04 | 2014-07-10 | 서울대학교 산학협력단 | Lung cancer diagnostic marker |
| KR101456683B1 (en) | 2013-01-04 | 2014-11-06 | 서울대학교산학협력단 | Novel Biomarker for the diagnosis of lung cancer |
| EP2787350A1 (en) * | 2013-04-05 | 2014-10-08 | Atlas Antibodies AB | ASRGL1 in endometrial cancer |
| WO2014161980A1 (en) * | 2013-04-05 | 2014-10-09 | Atlas Antibodies Ab | Asrgl1 in endometrial cancer |
| EP3093343A4 (en) * | 2014-01-10 | 2017-12-27 | Juntendo Educational Foundation | Method for assessing lymph node metastatic potential of endometrial cancer |
| US10620209B2 (en) | 2014-07-02 | 2020-04-14 | Dragon Victory Development Ltd. | Specific biomarker set for non-invasive diagnosis of liver cancer |
| US9885718B2 (en) | 2014-07-02 | 2018-02-06 | Dragon Victory Development Ltd. | Specific biomarker set for non-invasive diagnosis of liver cancer |
| US9506925B2 (en) | 2014-07-02 | 2016-11-29 | Dragon Victory Development Ltd. | Specific biomarker set for non-invasive diagnosis of liver cancer |
| US10822307B2 (en) | 2014-08-28 | 2020-11-03 | Eisai R&D Management Co., Ltd. | High-purity quinoline derivative and method for manufacturing same |
| US11186547B2 (en) | 2014-08-28 | 2021-11-30 | Eisai R&D Management Co., Ltd. | High-purity quinoline derivative and method for manufacturing same |
| US11090386B2 (en) | 2015-02-25 | 2021-08-17 | Eisai R&D Management Co., Ltd. | Method for suppressing bitterness of quinoline derivative |
| US11547705B2 (en) | 2015-03-04 | 2023-01-10 | Merck Sharp & Dohme Llc | Combination of a PD-1 antagonist and a VEGF-R/FGFR/RET tyrosine kinase inhibitor for treating cancer |
| US12083112B2 (en) | 2015-03-04 | 2024-09-10 | Eisai R&D Management Co., Ltd. | Combination of a PD-1 antagonist and a VEGFR/FGFR/RET tyrosine kinase inhibitor for treating cancer |
| US11369623B2 (en) | 2015-06-16 | 2022-06-28 | Prism Pharma Co., Ltd. | Anticancer combination of a CBP/catenin inhibitor and an immune checkpoint inhibitor |
| US12220398B2 (en) | 2015-08-20 | 2025-02-11 | Eisai R&D Management Co., Ltd. | Tumor therapeutic agent |
| US12303505B2 (en) | 2017-02-08 | 2025-05-20 | Eisai R&D Management Co., Ltd. | Tumor-treating pharmaceutical composition |
| US12226409B2 (en) | 2017-05-16 | 2025-02-18 | Eisai R&D Management Co., Ltd. | Treatment of hepatocellular carcinoma |
| WO2020047636A1 (en) * | 2018-09-04 | 2020-03-12 | Centro Nacional De Pesquisa Em Energia E Materiais | Method for metastatic prognosis in oral cancer, biosensor, prognosis kit, biomarkers and their use |
| WO2020223657A1 (en) * | 2019-05-02 | 2020-11-05 | Predictive Technology Group, Inc. | Somatic cancer driver mutations in endometriosis lesions contribute to secondary cancer risk |
| CN111500732A (en) * | 2020-05-25 | 2020-08-07 | 福建医科大学 | Application and kit of microorganisms as diagnostic markers for endometrial cancer |
| CN113049824A (en) * | 2021-04-20 | 2021-06-29 | 首都医科大学附属北京妇产医院 | Application of apolipoprotein ApoA1 in detection of drug resistance of cervical cancer to platinum chemotherapy |
| CN113493839A (en) * | 2021-09-06 | 2021-10-12 | 北京泱深生物信息技术有限公司 | Application of gene marker combination in diagnosis of endometrial cancer |
| CN113493840A (en) * | 2021-09-07 | 2021-10-12 | 北京泱深生物信息技术有限公司 | Marker for endometrial cancer diagnosis and derivative product and application thereof |
| CN113917149A (en) * | 2021-09-30 | 2022-01-11 | 江苏扬新生物医药有限公司 | Application of gelsolin detection substance in preparation of uterine cancer evaluation detection reagent |
| CN113917149B (en) * | 2021-09-30 | 2024-05-24 | 江苏扬新生物医药有限公司 | Application of gelsolin detection substance in the preparation of uterine cancer assessment and detection reagent |
| CN115184611A (en) * | 2022-08-01 | 2022-10-14 | 中国医学科学院北京协和医院 | Endometrial cancer stratification related marker and application thereof |
| CN119936393A (en) * | 2024-12-31 | 2025-05-06 | 哈尔滨脉图精准技术有限公司 | Metabolic marker composition for early endometrial cancer diagnosis and its application |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| WO2012019300A1 (en) | Endometrial cancer biomarkers and methods of identifying and using same | |
| US9863005B2 (en) | Endometrial phase or endometrial cancer biomarkers | |
| EP2252729B1 (en) | Biomarkers for head-and-neck cancers and precancers | |
| EP2251695B1 (en) | Markers associated with endometrial disease | |
| EP2567235B1 (en) | Method for the diagnosis of epithelial cancers by the detection of epicd polypeptide | |
| JP2005120099A (en) | A novel method for diagnosing, monitoring, staging, imaging and treating various cancers | |
| WO2011137519A1 (en) | Methods and compositions for the diagnosis and treatment of thyroid cancer | |
| US20060134120A1 (en) | Multiple marker assay for detection of ovarian cancer | |
| EP2195658A2 (en) | A method of assessing colorectal cancer status in an individual | |
| AU2011250588A1 (en) | Method for the diagnosis of epithelial cancers by the detection of EpICD polypeptide | |
| JP2006500036A (en) | Method for detecting endocrine cancer | |
| US20060141471A1 (en) | Assay for detection of renal cell carcinoma | |
| KR102084658B1 (en) | Metastasis-specific markers for diagnosing prognosis and determining treatment strategies of patient of clear cell renal cell carcinoma | |
| JP2002525031A (en) | Novel methods of diagnosing, monitoring, staging, imaging and treating colorectal cancer | |
| US20160011197A1 (en) | Methods and Compositions for the Diagnosis of a Thyroid Condition | |
| JP2010014689A (en) | Identification of melanoma marker using proteomics analysis | |
| WO2016201555A1 (en) | Methods and compositions for the diagnosis of a thyroid condition | |
| JP2002527757A (en) | Methods for diagnosing, monitoring, staging, imaging and treating prostate cancer | |
| EP4650779A1 (en) | Biomarker for diagnosis or prognosis of squamous cell lung carcinoma using exosomes and use thereof | |
| JP2026502556A (en) | Exosome-based biomarkers for the diagnosis or prognosis of squamous cell lung cancer and their uses | |
| KR20230126529A (en) | Extracellular vesicles-derived miRNA gene biomarkders for diagnosis of pancreatic cancer and use thereof | |
| HK1134690B (en) | Endometrial biomarkers | |
| Class et al. | Patent application title: Methods and Compositions for the Diagnosis of a Thyroid Condition | |
| HK1151073B (en) | Biomarkers for head-and-neck cancers and precancers |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| 121 | Ep: the epo has been informed by wipo that ep was designated in this application |
Ref document number: 11815965 Country of ref document: EP Kind code of ref document: A1 |
|
| NENP | Non-entry into the national phase |
Ref country code: DE |
|
| 122 | Ep: pct application non-entry in european phase |
Ref document number: 11815965 Country of ref document: EP Kind code of ref document: A1 |