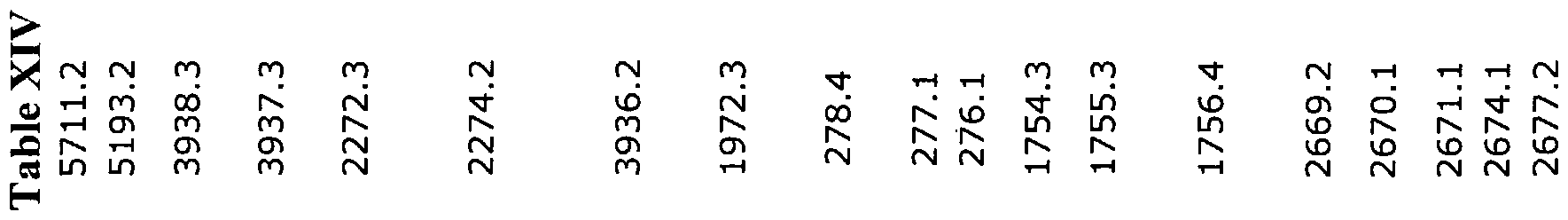GENOME OF LEGIONELLA PNEUMOPHILA PARIS AND LENS STRAIN - DIAGNOSTIC AND EPIDEMIOLOGICAL APPLICATIONS
The object of the invention is the genomic sequence and nucleotidic sequences coding for polypeptides of Legionella pneumophila Paris and Lens strain, such as polypeptides of cellular surface, especially specific between these two strains and/or relative to the Philadelphia strain, or implied in the virulence or in the biosynthesis of polysaccharides having a cellular envelope, as well as vectors including said sequences and cells transformed by these vectors. The invention likewise concerns processes for detection of these nucleic acids or polypeptides and diagnostic kits or kits for typing bacteria of the Legionella genre, especially the Legionella pneumophila species, such as the Paris and Lens strain, between them and/or relative to the Philadelphia strain. The invention especially concerns a specific repeated nucleic sequence of the Legionella pneumophila species and its utilization as an analysis target in processes for detection of the presence of these bacteria. The aim of the invention also is a method for selection of compounds capable of modulating the biosynthesis of these polysaccharides having a cellular envelope utilizing said nucleotidic sequences or said polypeptides. The invention finally comprises des pharmaceutical compositions, especially vaccinal, for the prevention and/or treatment of bacterial infections, in particular by the Legionella pneumophila Paris and/or Lens strain. Legionella is a bacteria of the environment responsible for legionellosis and Pontiac fever. The epidemiological data indicate that probably only certain isolates are capable of causing clinical cases. The L. pneumophila species seems to have a more significant virulence than the other species, by being responsible for 90% of the cases of legionellosis. At the centre of this species, among the fifteen serogroups, the isolates of serogroup 1 are associated with 80% of cases. To date, transmission from person to person has never been observed and measures for preventing legionellosis are thus concentrated on elimination of this pathogen from water circulation or from water-cooling towers in air-conditioning systems. In order to establish a rational policy for prevention, it is necessary to prevent the risk associated with each strain. In this optic, it would be desirable to be able to have processes or diagnostic kits available based on the recognition of protein or specific nucleic acid of this genre legio species pathogen or of a particular strain (or again subspecies) of this species.
In effect, the interest in using these specific sequences in the domain of diagnostics or epidemiology rests on the possibility of analyzing a large number of sequences at the same time and very rapidly for: - classification or typing of bacteria as a function of the presence of a sequence or of a profile of sequences characteristic of a genre, species or strain (sub-species) of bacteria, in particular in association with the gravity or not of pathologies which such bacteria can induce in case of infection in mammals, especially in humans; and - simultaneous comparison of sequence or profile of sequences between different genres, species or strain (sub-species) of bacteria, pathogenic or not, especially enabling identification of a gene, or the corresponding proteic sequence, or a profile of genes whereof the presence and/or expression in a bacteria is specific to a genre, species or strain of bacteria, and/or to its pathogenicity or not, especially by means of tools such as DNA chips or, if required, protein chips, on which these specific sequences are immobilized. These specific sequences can be specific sequences of the Legionella genre, or of a pathogenic bacteria of the Legionella genre and/or of the Legionella pneumophila species, or again of a bacteria of the Legionella pneumophila species Paris and/or Lens strain or again specific to a bacteria of the Legionella pneumophila species Paris and/or Lens strain relative to the Philadelphia strain. This information is widely utilized especially to rapidly identify the presence or not of the pathogenic bacteria, the gravity of the infection which it can cause, the treatment adapted to an infection, and/or the necessity and the means to be put in place to decontaminate the objects, circuits or fluids which are contaminated or could be contaminated This information will likewise be widely used for epidemiological studies relative to this genre of bacteria. This is just one of the aspects of the present invention. In a first stage, the inventors have studied and attempted to comprehend the genomic diversity at the centre of the Legionella genre by complete sequencing of the L. pneumophila serogroup 1 strain, Paris strain and Lens strain found in different French departements (2) and low-level sequencing of covering of two strains not belonging to the L. pneumophila species. The strains selected are L. longbeachae, responsible for cases of legionellosis essentially in Australia, and L. anisa frequently found in water circulation but not found in patients.
The complete sequencing has likewise been carried out within the scope of the invention of an epidemic L.pneumophila serogroup 1 strain known as « Lens strain », responsible for a major epidemic in France with 86 cases and 17 deaths between November 2003 and January 2004. The genes found to be variable between the strains of Legionella, as well as preserved genes having a functional tie to the virulence of L.pneumophila can be utilized to manufacture DNA chips. A large number of isolates of various origins isolated from the environment or originating from clinical cases could be analyzed by using this tool to identify markers enabling the two categories of strains to be discriminated. The comparison of endemic and epidemic isolates will likewise provide bases for understanding the specificities of these strains and in particular the adaptability and stability of the Paris strain. This approach also helps identify new functions necessary to the virulence of Legionella in humans and aid understanding of the different stages of this disease. The object of the invention is to allow development of novel tools for typing the strains of Legionella. These tools could be of the DNA "chip" type or of another type. The novel characteristics of these typing tools will be the following:
* Rapidity and simplicity of use
* High capacity for discrimination between the strains * Possibility for providing information on the genomic content of the strain analyzed and possibly prevent the risk associated with contamination by Legionella. The inventors have, during this study, brought to light genes found to be variable between the strains of Legionella, as well as preserved genes having a functional tie to wall or cellular envelope, or the virulence of L. pneumophila, genes which will be able to be used for carrying out these processes or diagnostic kits, especially for producing biochips with protein or DNA. A large number of isolates of diverse origin isolated from the environment or originating from clinical cases could be analyzed by using these tools for the puφose of identifying markers for discriminating the two categories of strains. This approach will also help identify new functions necessary for the virulence of Legionella in humans and aid in comprehension of the different stages of this disease. The object of the invention is thus to allow development of novel tools for typing of strains of Legionella. These tools could especially be of the protein or
DNA/RNA biochip type. The novel characteristics of these typing tools will be the following:
- rapidity and simplicity of use;
- high capacity for discrimination between strains; and - possibility of supplying information on the genomic content of the strain analyzed and of possibly preventing the risk associated with contamination by Legionella. The inventors have, in the first instance, sequenced the complete genome of L. pneumophila Paris strain in the form of 56 contigs (SEQ ID No. 1 to 56), a sequence made up of a long chromosome of around 3.65 Mb and a long plasmid of around 36 kb. The inventors have likewise identified on these contigs (SEQ ID No. 1 to 56) the nucleic sequences coding for proteins with their respective function (cf Table I with the annotated sequences). The inventors have additionally compared these sequences to the sequences of the genome of L. pneumophila Philadelphia strain available at the website http://genome3.cpmc.columbia.edu ~legion/index.html and revealed their presence or not in the sequences of this Philadelphia strain. The sequences of the genome of L. pneumophila Philadelphia strain available on the website http://genome3.cpmc.columbia.edu/~legion/index.html correspond to the sequences of the 51 contigs identified in the list of sequences under the SEQ ID Nos. 3456 to 3506. This comparison, made from the available genomic sequence of L. pneumophila Philadelphia strain and the proteic sequences obtained by the inventors from the 6 possible reading frames, has revealed that some 88 % of these two genomes are very strongly preserved (95 to 100 % of proteic identity), the remaining 12 % being specific to each strain (cf. Tables I and IV). These results thus demonstrate that there is a wide genomic diversity within the L. pneumophila species. A serine protease autotransporter homolog in which is inserted ten repetitions in tandem of a pattern of 60 amino acids was especially identified among the genes specific to the Paris strain. The autotransporters are secretion systems for the Gram negative bacteria in which the N- terminal and C-terminal parts respectively enable secretion across the internal membrane and the formation of pores in the external membrane. The central part of the protein can then remain exposed at the level of the cellular surface or can be split and salted out in the external medium. The role of certain autotransporters in the virulence of the negative Gram bacteria has already been shown; furthermore, work on the autotransporters of the enterobacteria has helped identify a group of serine protease
present solely in pathogenic bacterias whereof the diversity of functions could be linked to the adaptation to the niche occupied by the pathogen. In addition, the inventors have revealed a very wide inter-species diversity from the sequencing of a pathogenic strain of L. longbeachae, which causes very few cases of legionellosis in France, but which is the major source of legionellosis in Australia, and that of a non-pathogenicic L. anisa strain. We were able to identify to date 703 ORFs of the L. longbeachae strain, whereof 53 % are specific to it; the majority of ORFs preserved between the L. longbeachae and L. pneumophila Paris strains have a percentage of proteic homology greater than 80%. The preliminary results obtained by the inventors on the non-pathogenic L. anisa strain have helped identify 54 % of specific sequences and un percentage of proteic homology greater than 70 % for the sequences preserved between the L. anisa and L. pneumophila Paris strains. Tables I and II (« bestblast » obtained for each of the nucleic and proteic sequences corresponding to the annotated ORFs) and X to XXI hereinbelow comprise for each of the ORFs identified either in the Paris strain (Tables I and XIV), or in the Lens strain (Table XVI), its position on the contigs or chromosomes, and, if required, the existence of a peptide signal, the best result of the blast on prot (Best-Blastp). The ORFs: - specific to the L. pneumophila Paris strain relative to the L. pneumophila Philadelphia strain; - specific to the L. pneumophila Paris strain relative to the L. pneumophila Philadelphia and Lens strains (Table XVII); - specific to the L. pneumophila Lens strain relative to the L. pneumophila Philadelphia and Paris strains (Table XVIII); - specific to the L. pneumophila Philadelphia strain relative to the L. pneumophila Paris and Lens strains (Table XI); were identified in considering as specific the ORFs having a percentage of proteic identity less than 75 %. In the event where ORF is preserved in the two genomes, the percentage of identity between the two proteins is mentioned. hi this Table I, the ORFs present in the partial sequence of the L. longbeachae strain have likewise been noted. Finally, the ORFs specific to the Legionella genre were identified by considering as specific the ORFs having a percentage of proteic identity with sequences of the mprot bank less than 25 %.
05/049642
In Table XIX, the ORFs present at the same time in the Paris and Lens strain were indicated, though absent in the Philadelphia strain. In Table XX, the ORFs present at the same time in the Paris and Philadelphia strain were indicated, though absent in the Lens strain. Finally, in Table XXI, the ORFs present at the same time in the Philadelphia and
Lens strain were indicated, though absent in the Paris strain. In conclusion, the diversity revealed by the content of the present patent application helps define proteic probes, such as antibodies, or DNA probes for developing a typing tool. The utilization of this tool on a large number of strains isolated from patients and strains isolated from the environment will enable this tool to be validated, a tool which will aid in predicting the risk associated with a strain by discriminating in a certain manner the strains isolated from patients of other strains. Among the significant families of proteins of Legionella pneumophila Paris strain the family of surface proteins or that of proteins implied in the biosynthesis of surface polysaccharides can be cited, or again that of proteins implied in the virulence of these bacteria. The process of evolution has allowed the development of a number of unique mechanisms on the Gram+bacteria, by which they can immobilize proteins on their surface. The functions of these different proteins of cellular walls are extremely diverse. However, many proteins linked covalently to the surface of the Gram+ pathogens are estimated to be important for the survival of the pathogen inside the infected host. The study of Legionella pneumophila Paris strain demands novel approaches, in particular genetic, to improve understanding of the different metabolic paths of this organism. Accordingly, it is object of the present invention to divulge the complete sequence of the genome of Legionella pneumophila Paris strain (Collection de the Institut Pasteur CIP 107-629-T), a sequence obtained from a collection of clones (BAC) filed on 19 November 2003 with the Collection Nationale de Cultures de Microorganisms (CNCM) [National Collection of Microorganism Cultures], 25 rue du Docteur Roux, 75724 Paris Cedex 15, France, according to the arrangements of the Budapest Treaty and registered under file number 1-3137, as well as all the genes contained in said genome. It is also another object of the present invention to divulge the complete sequence of the genome of Legionella pneumophila Lens strain, a sequence obtained from a collection of clones (BAC) filed on 23 September 2004 with the Collection
05/049642
Nationale de Cultures de Microorganisms (CNCM) [National Collection of Microorganism Cultures], 25 rue du Docteur Roux, 75724 Paris Cedex 15, France, according to the arrangements of the Budapest Treaty and registered under file number 1-3306, as well as all the genes contained in said genome. In effect, knowledge of the genome of these organisms enables the interactions between the different genes, the different proteins, and the different metabolic paths to be better defined. In effect, and contrary to divulging isolated sequences, the complete genomic sequence of an organism forms a whole, allowing all the information necessary to this organism to grow and function to be obtained immediately. If the present invention provides the nucleotidic sequence of the genome of
Legionella pneumophila Paris strain (Collection de the Institut Pasteur CEP 107-629-T), having been the object of a filing of a collection of clones (BAC) covering this genome with the C.N.C.M. in Paris on 19 November 2003 and registered under file number I- 3138, and likewise provides the nucleotidic sequence of the genome of Legionella pneumophila Lens strain, a sequence obtained from a collection of clones (BAC) filed on 23 September 2004 with CNCM, and likewise provides certain polypeptide sequences coded by these two genomes, the specialist will be able to determine the other ORFs, by utilizing known methods, and appropriate software. In the set of claims hereinbelow, the term « nucleotidic sequence » will especially be able to be replaced by the term « polynucleotide » without modifying the object and the scope of the set of claims such as filed. The present invention thus relates to: - a genomic nucleotidic sequence of Legionella pneumophila Paris strain characterized in that it is selected among the sequences SEQ ID 3507 and 3508, SEQ ID N° 55 and the sequences SEQ ID N° 1 to SEQ ID N° 54, and SEQ ID N° 56; - a genomic nucleotidic sequence of Legionella pneumophila Lens strain characterized in that it is selected among the sequences SEQ ID 6733 and 6734. The present invention likewise relates to an isolated or purified nucleotidic sequence: (A) of Legionella pneumophila Paris strain, characterized in that it is selected among: a) a nucleotidic sequence comprising at least one sequence having 80 % identity with the sequences SEQ ID 3507 and 3508, and SEQ ID N° 1 to SEQ ID N° 56;
05/049642
b) a nucleotidic sequence hybridizing in very stringent conditions with the sequences SEQ ID 3507 and 3508, and SEQ ID N° 1 to SEQ ID N° 56; c) a nucleotidic sequence complementing sequences SEQ ED 3507 and 3508 and SEQ ID N° 1 to SEQ ID N° 56, or complementing a nucleotidic sequence such as defined in a), or b), or a corresponding nucleotidic sequence of RNA; and d) a nucleotidic sequence of at least 15 nucleotides of fragment representative of sequences SEQ ID 3507 and 3508, and SEQ ID N° 1 to SEQ ID N° 56, or of fragment representative of their sequence, or (B) of Legionella pneumophila Lens strain, characterized in that it is selected among: a) a nucleotidic sequence comprising at least one sequence having 80 % of identity with the sequences SEQ ID 6733 and 6734; b) a nucleotidic sequence hybridizing in very stringent conditions with the sequences SEQ ID 6733 and 6734; c) a nucleotidic sequence complementing sequences SEQ ED 6733 and
6734; or complementing a nucleotidic sequence such as defined in a), or b), or a corresponding nucleotidic sequence of RNA; and d) a nucleotidic sequence of at least 15 nucleotides of fragment representative of sequences SEQ ID 6733 and 6734; or of fragment representative of their sequence. More particularly, the object of the present invention likewise is the nucleotidic sequences characterized in that they originate from sequences SEQ ID 3507 and 3508, and SEQ ID N° 1 to SEQ ID N° 56, and in that they code for a polypeptide selected from amongst the polypeptides of sequences SEQ ID 3509 to SEQ ID 6732, and SEQ ID N° 56 to SEQ ID N° 3455, preferably coding for a secreted enzyme likewise present in the Lens strain and Philadelphia strain of sequences SEQ ID 3675, 4267, 4292 and 6477, preferably coding for a polypeptide present on the surface of Legionella pneumophila Paris strain of sequence SEQ ID Nos. 3410, 704, 746, 2267, 2751, 3192, 3218, 3221, 3222, 3317, 3324, 136, 171, 310, 337, 481, 527 652, 664, 893, 972, 1148, 1298, 1361, 1503, 1521, 1576, 1651, 1755, 1847, 1877, 2224, 2406, 2843, 2930, 3037, 3139, 3157, 3165, 3181, preferably coding for a polypeptide present on the specific surface of Legionella pneumophila Paris strain relative to the Philadelphia strain, especially of sequence SEQ ID Nos. 3410, 171, 337, 481, 652, 1148, 1521, 2843, 3037, 3181, or one of its representative fragments of at least 5 amino acids, or coding for a
/049642
polypeptide implied in the biosynthesis of polyssacharides having a cellular envelope of sequence SEQ ID Nos. 1126, 3218, 288, 632, 917, 1503, 1555, 1877, 1928, 1963, 2204, 2212, 2243, 2324, 2378, 2410, 2411, or again the nucleic sequences of the rtxA gene, especially those coding for the polypeptides of sequences SEQ ID Nos. 3410, 3037, 3165 and 3181. The present invention also relates more generally to the nucleotidic sequences derived from the sequences SEQ ID Nos. 3507 and 3508, and SEQ ID N° 1 to SEQ ID N° 56, and coding for a polypeptide of Legionella pneumophila Paris strain such that they can be isolated from these sequences SEQ ID Nos. 3507 and 3508, and SEQ ID N° 1 to SEQ ID N° 56. In addition, the nucleotidic sequences characterized in that they comprise a nucleotidic sequence selected among: a) a nucleotidic sequence coding for a polypeptide selected from amongst the sequences SEQ ID Nos. 3509 to 6732, and SEQ ID N° 56 to SEQ ID N° 3455; b) a nucleotidic sequence comprising at least 80 %, 85 %, 90 %, 95 % or 98 % of identity with a nucleotidic sequence coding for a polypeptide selected from amongst the sequences SEQ ID Nos. 3509 to 6732, and SEQ ID N° 56 to SEQ ID N° 3455; c) a nucleotidic sequence hybridizing in very stringent conditions with a nucleotidic sequence coding for a polypeptide selected from amongst the sequences SEQ ID Nos. 3509 to 6732, and SEQ ID N° 56 to SEQ ID N° 3455; d) a complementary nucleotidic sequence or RNA corresponding to a sequence such as defined in a), b) or c); e) a nucleotidic sequence of a representative fragment having at least 15 nucleotides of a sequence such as defined in a) or d); and f) a modified nucleotidic sequence of a sequence such as defined in a), d) or e), are likewise objects of the invention. Nucleic acid, nucleic sequence or nucleic acid, polynucleotide, oligonucleotide, polynucleotidic sequence, nucleotidic sequence, terms which will be employed indifferently in the present description, are understood to designate precise chaining of nucleotides, modified or not, effectively defining a fragment or a region of a nucleic acid, comprising or not non-natural nucleotides, and able to correspond just as well to a double-strand DNA, a single-strand DNA as transcription products of said DNAs.
Therefore, the nucleic sequences according to the invention likewise encompass the
PNA (Peptid Nucleic Acid), or similar.
05/049642 10
It must be understood that the present invention does not relate to the nucleotidic sequences in their natural chromosomic environment, that is, in the natural state. These are sequences which were isolated and/or purified, that is they were sampled directly or indirectly, for example by copy, their environment having been at least partially modified. It is understood to likewise designate the nucleic acids obtained by chemical synthesis. « Percentage of identity » between two sequences of nucleic acids or amino acids in the sense of the present invention is understood to designate a percentage of nucleotides or residues of identical amino acids between the two sequences to be compared, obtained after the best alignment, this percentage being purely statistical and the differences between the two sequences being distributed randomly and over their entire length. "Best alignment" or " optimal alignment " is understood to designate the alignment for which the percentage of identity determined hereinafter is the highest. The comparisons of sequences between two sequences of nucleic acids or amino acids are traditionally made by comparing these sequences after they were aligned in optimal fashion, said comparison being made by segment or by « window of comparison » to identify and compare the local regions of similarity of sequence. The optimal alignment of the sequences for comparison can be made, apart from manually, by means of the local of de Smith and Waterman (1981, Ad. App. Math. 2:482), by means of the local homology algorithm of Neddleman and Wunsch (1970, J. Mol. Biol. 48:443), by means of the similarity search method of Pearson and Lipman (1988, Proc. Natl. Acad. Sci. USA 85:2444), by means of software utilizing these algorithms (GAP, BESTFIT, BLAST P, BLAST N, FASTA and TFASTA in the Wisconsin Genetics Software Package, Genetics Computer Group, 575 Science Dr., Madison, WI). To obtain optimal alignment, the program BLAST is preferably used, with the BLOSUM 62 matrix. The PAM or PAM250 matrices can likewise be used. The percentage of identity between two sequences of nucleic acids or amino acids is determined by comparing these two sequences aligned optimally in which the sequence of nucleic acids or amino acids to be compared can comprise additions or deletions relative to the reference sequence for optimal alignment between these two sequences. The percentage of identity is calculated by determining the number of identical positions for which the nucleotide or the residue of amino acid is identical between the two sequences, by dividing this number of identical positions by the total
number of positions compared and by multiplying the result obtained by 100 to obtain the percentage of identity between these two sequences. Nucleic sequences having a percentage of identity of at least 80 %, preferably 85 % or 90 %, more preferably 95 % or 98 % or 99 %, after optimal alignment with a reference sequence are understood to designate he nucleic sequences exhibiting, relative to the nucleic reference sequence, certain modifications such as in particular deletion, truncation, elongation, chimeric fusion and/or substitution, especially specific, and whereof the nucleic sequence has at least 80 %, preferably 85 %, 90 %, 95 %, 98 % or 99 %, of identity after optimal alignment with the nucleic reference sequence. These are preferably sequences whereof the complementary sequences are capable of being hybridized specifically with the reference sequences. Preferably, the specific hybridization conditions or stringent conditions will be such that they ensure at least 80 %, preferably 85 %, 90 %, 95 %, 98 % or 99 % of identity after optimal alignment between one of the two sequences and the sequence complementary to the other. Hybridization in very stringent conditions signifies that the temperature and ionic force conditions are selected in such a way that they permit hybridization to be maintained between two fragments of complementary DNA. By way of illustration, very stringent conditions of the hybridization stage for the puφose of defining the polynucleotide fragments described hereinabove, are advantageously the following. DNA-DNA or DNA-RNA hybridization is performed in two stages: (1) prehybridization at 42°C for 3 hours in phosphate buffer (20 mM, pH 7.5) containing 5 x SSC (1 x SSC corresponds to a solution 0.15 M NaCI + 0.015 M sodium citrate), 50 % formamide, 7 % sodium dodecyl sulfate (SDS), 10 x Denhardt's, 5 % dextran sulfate and 1 % DNA salmon sperm; (2) hybridization per se for 20 hours at a temperature depending on the size of the probe (i.e.: 42°C, for a probe of size > 100 nucleotides) followed by 2 washes of 20 minutes at 20°C in 2 x SSC + 2 % SDS, 1 wash of 20 minutes at 20°C in 0.1 x SSC + 0.1 % SDS. The last wash is done in 0.1 x SSC + 0.1 % SDS for 30 minutes at 60°C for a probe of size > 100 nucleotides. The very stringent hybridization conditions described hereinabove for a polynucleotide of defined size can be adapted by the specialist for oligonucleotides of larger or smaller size, according to the teaching of Sambrook et al, (1989, Molecular cloning: a laboratory manual. 2nd Ed. Cold Spring Harbor). In addition, fragment representative of sequences according to the invention is understood to designate any nucleotide fragment having at least 15 nucleotides,
preferably at least 20, 25, 30, 50, 75, 100, 150, 300 and 450 consecutive nucleotides of the sequence from which it originates. Representative fragment is understood in particular to be a nucleic sequence coding for a biologically active fragment of a polypeptide, such as defined hereinbelow. Representative fragment is likewise understood to be the intergenic sequences, and in particular the nucleotidic sequences bearing the regulation signals (promoters, terminators, or enhancers, ...), or again probe or primer sequences aiding in specifically detecting or amplifying the nucleic sequences coding for the polypeptides of sequences SEQ ID Nos. 3509 to 6732, and SEQ ID N° 56 to SEQ ID N° 3455. Of said representative fragments those are preferred having nucleotidic sequences corresponding to open reading frames, known as ORF sequences (ORF for « Open Reading Frame »), included in general between an initiation codon and a stop codon, or between two stop codons, and coding for polypeptides, preferably of at least 100 amino acids, such as for example, without limiting them, the ORF sequences to be described hereinafter. The representative fragments according to the invention can be obtained for example by specific amplification such as PCR or after digestion by appropriate restriction enzymes of nucleotidic sequences according to the invention, this method being described in particular in the work of Sambrook et al.. Said representative fragments can likewise be obtained by chemical synthesis as long as their size is not too significant, according to methods well known to the specialist. The representative genome fragments of Legionella pneumophila Paris strain according to the invention likewise comprise at least one fragment of at least 15 nucleotides or more as cited above for the fragments resulting from enzymatic cutting at the level of a restriction site. Of course, expression proteins such as RNA or proteins are understood according to the present invention. Among the sequences containing inventive sequences, or representative fragments, are likewise understood the sequences which are naturally framed by sequences which present at least 80 %, 85 %, 90 %, 95 % or 98 % of identity with the sequences according to the invention. Modified nucleotidic sequence is understood as any nucleotidic sequence obtained by mutagenesis according to techniques well known to the specialist, and comprising des, preferably a maximum 10 %, 7.5 %, 5 %, 2.5 %, 1 %, 0.5 %, 0.1 % or even less than 0.01 %, of modified nucleotides, relative to normal sequences, for
example mutations in the regulating and/or promoting sequences of the expression of the polypeptide, especially leading to modification of the rate of expression or activity of said polypeptide. Modified nucleotidic sequence is likewise understood as any nucleotidic sequence coding for a polypeptide modified such as defined hereinbelow. The representative fragments according to the invention can likewise be probes or primers, which can be utilized in processes for detection, identification, dosage or amplification of nucleic sequences. In a preferred manner the invention is relative to a nucleotidic sequence coding for a polypeptide according to the invention. In a preferred manner the invention is relative to a nucleotidic sequence according to the invention, characterized in that it codes for a specific polypeptide of a bacteria of the Legionella genre, or one of its fragments of at least 5 amino acids, or its complementary nucleic sequence. In a preferred manner the invention is relative to a nucleotidic sequence according to the invention, characterized in that it codes for a specific polypeptide of a pathogenic bacteria of the Legionella genre and/or of the Legionella pneumophila species, or one of its fragments of at least 5 amino acids, or its complementary nucleic sequence. In a preferred manner the invention is relative to a nucleotidic sequence according to the invention, characterized in that it codes for a specific polypeptide of a bacteria of the Legionella pneumophila species Paris strain, or one of its fragments of at least 5 amino acids, or its complementary nucleic sequence. In a preferred manner the invention is relative to a nucleotidic sequence according to the invention, characterized in that it codes for a specific polypeptide of a bacteria of the Legionella pneumophila species Paris strain relative to the Philadelphia strain, or one of its fragments of at least 5 amino acids, in particular selected from amongst the polypeptides of sequence SEQ ID Nos. 3410, 171, 337, 481, 652, 1148, 1521, 2843, 3037, 3181 or one of its fragments of at least 5 amino acids, or its complementary nucleic sequence. In a preferred manner the invention is relative to a nucleotidic sequence according to the invention, characterized in that it codes for a specific polypeptide of a bacteria of the Legionella pneumophila species Paris strain relative to the Lens and Philadelphia strains, or one of its fragments of at least 5 amino acids, in particular
selected from amongst the polypeptides whereof the sequences are indicated in Table XVII or one of their fragments of at least 5 amino acids, or their complementary nucleic sequence. In a preferred manner the invention is relative to a nucleotidic sequence according to the invention, characterized in that it codes for a surface polypeptide of Legionella pneumophila Paris strain, or one of its fragments of at least 5 amino acids, in particular selected from amongst the sequence polypeptides SEQ ID Nos. 3410, 704, 746, 2267, 2751, 3192, 3218, 3221, 3222, 3317, 3324, 136, 171, 310, 337, 481, 527 652, 664, 893, 972, 1148, 1298, 1361, 1503, 1521, 1576, 1651, 1755, 1847, 1877, 2224, 2406, 2843, 2930, 3037, 3139, 3157, 3165, 3181, or one of its fragments of at least 5 amino acids, or its complementary nucleic sequence. In a preferred manner the invention is relative to a nucleotidic sequence according to the invention, characterized in that it codes for a polypeptide of specific surface of Legionella pneumophila Paris strain relative to the Philadelphia strain, selected from amongst the polypeptides of sequences SEQ ID Nos. 3410, 171, 337, 481, 652, 1148, 1521, 2843, 3037, 3181, or one of its fragments of at least 5 amino acids, or its complementary nucleic sequence. In a preferred manner the invention is relative to a nucleotidic sequence according to the invention, characterized in that it codes for a polypeptide implied in the biosynthesis of polysaccharide having a cellular envelope of Legionella pneumophila
Paris strain, in particular selected from amongst the polypeptides of sequence SEQ TD Nos. 1126, 3218, 288, 632, 917, 1503, 1555, 1877, 1928, 1963, 2204, 2212, 2243, 2324, 2378, 2410, 2411, or one of its representative fragments of at least 5 amino acids, or its complementary nucleic sequence. In a preferred manner the invention is relative to a nucleotidic sequence according to the invention, characterized in that it codes for a polypeptide f Legionella pneumophila Paris strain coded by the rtxA gene of sequence SEQ ID Nos. 3410, 3037, 3165 and 3181, or one of its representative fragments of at least 5 amino acids, or its complementary nucleic sequence. The rtxA gene of Legionella pneumophila was demonstrated as being implied in virulence. This gene codes for a protein of 1208 aa. Four ORFs whereof the references are given hereinbelow (SEQ ID 3410 - 3037 - 3165 - 3181) correspond to this gene in the Paris strain which would code for a protein of at least 4000 aa. Comparison with the Philadelphia strain shows the presence of a homologous gene for the N- and C-terminal
parts. The central part of the protein is constituted by repetitions of a pattern of around 20 aa different between the two strains. In another aspect, in a preferred manner, the invention is relative to a nucleotidic sequence according to the invention characterized in that it codes for a polypeptide of Legionella pneumophila Paris strain, or one of its representative fragments of at least 5 amino acids implied in the biosynthesis of the amino acids, in the biosynthesis of the cofactors, prosthetic and transporter groups, implied in the cellular machinery, implied in the central intermediate metabolism, implied in the energetic metabolism, implied in the metabolism of fatty acids and phospholipids, implied in the metabolism of nucleotides, purines, pyrimidines or nucleosides, implied in the functions of regulation, implied in the process of replication, implied in the process of transcription, implied in the process of translation, implied in the process of transport and bonding of proteins, implied in adaptation to atypical conditions, implied in sensitivity to medications and the like, or implied in the functions relative to transposons. Owing to the genomic sequence presented in the present invention, the specialist will know how to identify the genes coding for proteins regulating transcription of the genes of Legionella pneumophila Paris strain. In addition, Table I provides the list of the open reading phases (ORF for « Open Reading Frame) annotated and identified on the genome of Legionella pneumophila Paris strain (SEQ ID N° 1 to SEQ ID N° 55), with especially their position on said genome, and, in Table II, the putative functions which can be attributed to them by utilizing customary techniques for comparing the genomic (« Bestblast »). All the same, such a list must not be considered as limiting, where one protein can have several roles in the cell. Modifying the structure or the integrity of these genes could help modify the expression of the target genes controlled by the target promoters of these regulators. Thus, the expert will be able to select the regulator(s) pertinent for the required application as well as their target, thus allowing optimization of the expression of genes of interest. The utilization of the tools described above such as the DNA chips, also registers all the genes whereof the regulation is modified by inactivation of certain genes. It is thus possible to select a set of control sequences responding to the same type of regulation. These sequences can then be used to control the expression of genes of interest. In general, the list of sequences SEQ ID, or their corresponding coding nucleic sequence could be determined by the specialist from the most probable putative
5/049642 16
functions determined for each of the sequences SEQ ID in Tables I and XIV hereinbelow for each of the classes of activity classified hereinbelow. It is important to note all the same that a living organism is a whole and must be taken as such. Accordingly, so as to develop and exhibit its properties, any organism has need of interaction between the different metabolic paths. Therefore, the abovementioned classification must not be considered as limiting, with a gene able to be implied in two distinct metabolic paths. The invention likewise relates to polypeptides coded by a nucleotidic sequence according to the invention, preferably by a fragment representative of the sequences SEQ ID Nos. 3509 to 6732, the sequence SEQ ID N° 55 or sequences SEQ ID N° 1 to
SEQ ID N° 54, and SEQ ID N° 56, and corresponding to an ORF sequence, such as described in Table XIV (coding for one of the sequences SEQ ID N° 3509 to SEQ ID
N° 6732), and in Table I (coding for one of the sequences SEQ ID N° 57 to SEQ ID
N° 3455). In particular, polypeptides of Legionella pneumophila Paris strain, characterized in that they are selected from amongst the following polypeptides: - polypeptides of sequences SEQ ED Nos. 3509 to 6732 and of sequences SEQ ID N° 56 to SEQ ID N° 3455; - preferably enzymes secreted by Legionella pneumophila Paris, Lens and Philadelphia strains, especially sequences SEQ ID Nos. 3675, 4267, 4292 and 6477; - preferably polypeptides present on the surface of Legionella pneumophila Paris strain, especially of sequence SEQ ID Nos. 3410, 704, 746, 2267, 2751, 3192, 3218, 3221, 3222, 3317, 3324, 136, 171, 310, 337, 481, 527 652, 664, 893, 972, 1148, 1298, 1361, 1503, 1521, 1576, 1651, 1755, 1847, 1877, 2224, 2406, 2843, 2930, 3037, 3139, 3157, 3165, 3181, and still more preferred polypeptides present on the specific surface of Legionella pneumophila Paris strain relative to the Philadelphia strain, especially those of sequences SEQ ID Nos. 3410, 171, 337, 481, 652, 1148, 1521, 2843, 3037, 3181, or one of its representative fragments of at least 5 amino acids; - polypeptides implied in the biosynthesis of polysaccharides having a cellular envelope, especially of sequence SEQ ID Nos. 1126, 3218, 288, 632, 917, 1503, 1555,
1877, 1928, 1963, 2204, 2212, 2243, 2324, 2378, 2410, 2411; - or again polypeptides of sequence SEQ ID Nos. 3410, 3037, 3165 and 3181, coded by the rtxA gene.
5/049642 17
The invention likewise comprises polypeptides characterized in that they comprise a polypeptide selected from amongst: a) a polypeptide of sequences SEQ ID Nos. 3509 to 6732, and of sequences SEQ ID N° 56 to SEQ ID N° 3455; b) a polypeptide having at least 80 % preferably 85 %, 90 %, 95 % and
98 % of identity with an polypeptide of sequences SEQ ID Nos. 3509 to 6732, and of sequences SEQ ID N° 56 to SEQ ED N° 3455; c) a fragment of at least 5 amino acids, preferably biologically active, of one such as defined in b); d) a biologically active fragment of a polypeptide of sequence SEQ ID
N° 56 to SEQ ED N° 3455; and e) a modified polypeptide of a polypeptide of sequences SEQ ID Nos. 3509 to 6732, and of sequences SEQ ID N° 56 to SEQ ID N° 3455, comprising at most 10 % modified amino acids, preferably 7.5 %, 5 %, 2.5 %, 1 %, 0.5 %, 0, 1 % or again 0.01 %. The nucleotidic sequences coding for the abovedescribed polypeptides are likewise an object of the invention. In the present description, the terms polypeptides, polypeptide sequences, peptides and proteins are interchangeable. It must be understood that the invention does not relate to polypeptides in the natural form, that is, that they are not taken in their natural environment, rather they were isolated or obtained by purification from natural sources, or else obtained by genetic recombination, or by chemical synthesis, and that they can then comprise non- natural amino acids such as will be described hereinbelow. Polypeptide having a certain percentage of identity with another, likewise designated by homologous polypeptide, is understood to designate those polypeptides having, relative to natural polypeptides, certain modifications, in particular deletion, addition or substitution of at least one amino acid, truncation, elongation, a chimeric solution and/or a mutation, or polypeptides having post-translational modifications. Of the homologous polypeptides preference is given to those whereof the sequence of amino acids has at least 80 %, preferably 85 %, 90 %, 95 %, 98 % and 99 % of identity with the sequences of amino acids of the polypeptides according to the invention. In the case of substitution, one or more consecutive or non-consecutive amino acid(s) are replaced by « equivalent » amino acids. The expression « equivalent amino acids» in
5/049642 18
this case endeavors to designate any amino acid likely to be substituted for one of the amino acids of the base structure without however essentially modifying the biological activities of the corresponding peptides and such as they will be defined hereinbelow. These equivalent amino acids can be determined either by being supported on their structural homology with the amino acids for which they are substituted, or on results of comparative assays of biological activity between the different polypeptides capable of being effected. By way of example, mention is made of the possibilities of substitution for being carried out without the resulting extensive modification of the biological activity of the corresponding modified polypeptide. Thus leucine can be replaced by valine or isoleucine, aspartic acid by glutamine acid, glutamine by asparagine, arginine by lysine, etc., the inverse substitutions naturally being envisaged under the same conditions. The homologous polypeptides correspond likewise to the polypeptides coded by the homologous or identical nucleotidic sequences, such as defined previously and thus comprise in the present definition mute polypeptides or corresponding to inter or intra species variations, able to exist in Legionella, and which correspond especially to truncations, substitutions, deletions and/or additions, of at least one residue of amino acids. It is understood that the percentage of identity between two polypeptides is calculated in the same way as between two sequences of nucleic acids. Thus, the percentage of identity between two polypeptides is calculated after optimal alignment of these two sequences, on a maximum homology window. To define said maximum homology window, the same algorithms as for the nucleic acid sequences can be utilized. Biologically active fragment of a polypeptide according to the invention is understood to mean in particular a fragment of polypeptide, such as defined hereinbelow, having at least one of the biological characteristics of the polypeptides according to the invention, especially in that it capable of exerting in general an even partial activity, such as for example: - enzymatic (metabolic) activity or an activity able to be implied in the biosynthesis or biodegradation of organic or inorganic compounds; structural activity (cellular envelope, coping molecule, ribosome); transport activity (energy, ion); or in the secretion of protein;
activity in the process of replication, amplification, preparation, transcription, translation or maturation, especially of DNA, RNA or proteins. Fragment of polypeptides according to the invention is understood to mean a polypeptide comprising a minimum of 5 amino acids, preferably 6, 7, 8, 9, 10, 15, 20, 25, 30, 40, 50, 75, 100, 150 acids and 200 amino acids. The fragments of polypeptides can correspond to isolated or purified fragments naturally present in the strains of Legionella, or to fragments which can be obtained by cleavage of said polypeptide by a proteolitic enzyme such as trypsine or chymotrypsine or collagenase, by a chemical reagent (cyanogen bromide, CNBr) or by placing said polypeptide in a highly acidic environment (for example at pH = 2.5). Polypeptidic fragments can likewise be prepared by chemical synthesis, from hosts transformed by a vector of expression according to the invention which contains a nucleic acid allowing expression of said fragment, and placed under the control of the appropriate elements of regulation and/or expression. « Modified polypeptide» of a polypeptide according to the invention is understood to mean a polypeptide obtained by genetic recombination or by chemical synthesis such as described below, which has at least one modification relative to the normal sequence, preferably at most 10 % of amino acids modified relative to the normal sequence, preferably even at most 7.5 %, 5 %, 2.5 %, 1 %, 0.5 %, 0, 1 % or again 0.01 %. These modifications can be especially made to amino acids necessary for the specificity or efficacy of the activity, or at the origin of the structural conformation, the charge, or the hydrophobicity of the polypeptide according to the invention. Polypeptides of equivalent, augmented or diminished activity, or of equivalent, narrower or wider specificity can thus be created. Among the polypeptides modified, those polypeptides in which up to five amino acids can be modified, truncated at the N end or C terminal, or else deleted, or added, must be mentioned. As is indicated, the object of the modifications of a polypeptide especially are: to permit its usage in biosynthesis or biodegradation processes of organic or inorganic compounds, - to permit its usage in processes of replication, amplification, repair and transcription, translation, or maturation especially of DNA, RNA, or proteins, to permit its improved secretion, to modify its solubility, efficacy or specificity of activity, or again to facilitate its purification.
Chemical synthesis likewise has the advantage of being able to utilize non- natural amino acids or non-peptidic bonds. Therefore, it can be interesting to utilize non-natural amino acids, for example in the D form, or analogs of amino acids, especially sulphurized forms. In another aspect, the invention is preferably relative to a polypeptide according to the invention, characterized in that it is a specific polypeptide of a bacteria of the Legionella genre, or one of its fragments of at least 5 amino acids. In another aspect, the invention is preferably relative to a polypeptide according to the invention, characterized in that it is a specific polypeptide of a pathogenic bacteria of the Legionella geme and or of the Legionella pneumophila species, or one of its representative fragments of at least 5 amino acids. In another aspect, preferably, the invention is relative to a polypeptide according to the present invention, characterized in that it is a specific polypeptide of a bacteria of the species Legionella pneumophila Paris strain, or one of its representative fragments of at least 5 amino acids. In another aspect, preferably, the invention is relative to a polypeptide according to the present invention, characterized in that it is a specific polypeptide of a bacteria of the Legionella pneumophila species Paris strain relative to the Philadelphia strain, or one of its representative fragments of at least 5 amino acids, in particular selected from amongst the polypeptides of sequences SEQ ID Nos. 3410, 171, 337, 481, 652, 1148, 1521, 2843, 3037, 3181, or one of its representative fragments of at least 5 amino acids. In another aspect, preferably, the invention is relative to a polypeptide according to the present invention, characterized in that it is a specific polypeptide of a bacteria of the Legionella pneumophila species Paris strain relative to the Lens and Philadelphia strains, or one of its representative fragments of at least 5 amino acids, in particular selected from amongst the polypeptides whereof the sequence is indicated in Table XVII, or one of their representative fragments of at least 5 amino acids. In another aspect, preferably, the invention is relative to a polypeptide according to the present invention, characterized in that it is a surface polypeptide of Legionella pneumophila Paris strain, or one of its representative fragments of at least 5 amino acids, in particular selected from amongst the polypeptides of sequence SEQ ID Nos. 3410, 704, 746, 2267, 2751, 3192, 3218, 3221, 3222, 3317, 3324, 136, 171, 310, 337, 481, 527 652, 664, 893, 972, 1148, 1298, 1361, 1503, 1521, 1576, 1651, 1755, 1847,
5/049642 21
1877, 2224, 2406, 2843, 2930, 3037, 3139, 3157, 3165, 3181, or one of its representative fragments of at least 5 amino acids. In another aspect, preferably, the invention is relative to a polypeptide according to the present invention, characterized in that it is a polypeptide of specific surface of Legionella pneumophila Paris strain relative to the Philadelphia strain, selected from amongst the polypeptides of sequence SEQ ID Nos. 3410, 171, 337, 481, 652, 1148, 1521, 2843, 3037, 3181, or one of its representative fragments of at least 5 amino acids. In another aspect, preferably, the invention is relative to a polypeptide according to the invention, characterized in that it is a polypeptide Legionella pneumophila Paris strain implied in the biosynthesis of polysaccharide having a cellular envelope of Legionella pneumophila Paris strain, in particular selected from amongst the polypeptides of sequence SEQ ED Nos. 1126, 3218, 288, 632, 917, 1503, 1555, 1877, 1928, 1963, 2204, 2212, 2243, 2324, 2378, 2410, 2411, or one of its representative fragments of at least 5 amino acids. In another aspect, preferably, the invention is relative to a polypeptide according to the invention, characterized in that it is a polypeptide de Legionella pneumophila Paris strain coded by the rtxA gene de sequence SEQ ID Nos. 3410, 3037, 3165 and 3181, or one of its representative fragments of at least 5 amino acids, or its complementary nucleic sequence. In another aspect, preferably, the invention is relative to a polypeptide according to the invention, characterized in that it is a polypeptide of Legionella pneumophila Paris strain, or one of its representative fragments of at least 5 amino acids, implied in the biosynthesis of amino acids, in the biosynthesis of cofactors, prosthetic and transporter groups, implied in the cellular machinery, implied in the central intermediary metabolism, implied in the energetic metabolism, implied in the metabolism of fatty acids and phospholipids, implied in the metabolism of nucleotides, purins, pyrimidins or nucleosides, implied in the functions of regulation, implied in the process of replication, implied in the process of transcription, implied in the process of translation, implied in the process of transport and binding of proteins, implied in the adaptation to atypical conditions, implied in the sensitivity to drugs and the like, or implied in the functions relatives to transposons. The object of the present invention is likewise the nucleotidic sequences and/or polypeptides according to the invention, characterized in that said sequences are registered on a registration support whose form and nature facilitate reading, analysis
and/or exploitation of said sequence(s). These supports can likewise contain other information extracted from the present invention, especially the analogies with already known sequences, and/or information concerning the nucleotidic sequences and/or polypeptides of other microorganisms for the puφose of facilitating comparative analysis and exploitation of the results obtained. Among said registration supports, preference is given in particular to those supports readable by a computer, such as magnetic, optic, electric or hybrid supports, in particular information discs, CD-ROM, servers. Such registration supports are likewise an object of the invention. The registration supports according to the invention, with the information contributed, are very useful for the choice of primers or nucleotidic probes for determining genes in Legionella pneumophila Paris strain or strains close to this organism. Similarly, the utilization of these supports for the study of genetic polymoφhism of a strain close to Legionella pneumophila Paris strain, in particular by determination of the colinearity regions, is very useful as far as these supports provide not only the nucleotidic sequence of the genome of Legionella pneumophila Paris strain, but likewise the genomic organization in said sequence. Thus, the uses of registration supports according to the invention are likewise objects of the invention. The homology analysis between different sequences is completed in effect advantageously by means of software for sequence comparisons, such as BlastP or BlastN software, or other software well known to the specialist. A probe or primer is defined, in the sense of the invention, as being a fragment of single-strand nucleic acids or a denatured double-strand fragment comprising for example 12 bases at several kb, especially 15 at several hundreds of bases, preferably from 15 to 50 or 100 bases, and possess a hybridization specificity in conditions determined to form a hybridization complex with a nucleic acid target. The probes and primers according to the invention can be marked directly or indirectly by a radioactive or non-radioactive compound using methods well known to the specialist, to obtain a detectable and/or quantifiable signal. The non-marked sequences of polynucleotides according to the invention can be utilized directly as probe or primer. The sequences are generally marked to obtain sequences utilizable for numerous applications. The marking of the primers or probes according to the invention is done by radioactive elements or by non-radioactive molecules.
Examples of the radioactive isotopes used are 32P, 33P, 35S, 3H or 125I. The non- radioactive entities are selected among ligands such as biotin, avidin, streptavidin, dioxygenin, haptenes, colorants, luminescent agents such as radioluminescent, chemoluminescent, bioluminescent, fluorescent, phosphorescent agents. The polynucleotides according to the invention can thus be utilized as primer and or probe in processes in particular making use of the PCR technique (amplification in chain by polymerase) (Rolfs et al, 1991, Berlin: Springer- Verlag). This technique requires the choice of pairs of oligonucleotide primers framing the fragment to be amplified. For example, reference can be made to the technique described in the U.S. patent N° 4,683,202. The amplified fragments can be identified, for example after electrophoresis in agarose or polyacrylamide gel, or according to a chromatographic technique such as filtration on gel or ion exchange chromatography, then sequenced. The specificity of the amplification can be controlled by using as primer the inventive nucleotidic sequences of polynucleotides as matrix, plasmids containing these sequences or even the derived amplification products. The amplified nucleotide fragments can be utilized as reagents in hybridization reactions so as to reveal the presence, in a biological sample, of a nucleic acid target of sequence complementary to those of said amplified nucleotide fragments. The aim of the invention is likewise the nucleic acids capable of being obtained by amplification by means of primers according to the invention. Other amplification techniques for the nucleic acid target can be advantageously employed as an alternative to PCR (PCR-like) by means of a couple of primers of nucleotidic sequences according to the invention. PCR-like is understood to mean all the methods making use of direct or indirect reproductions of the sequences of nucleic acids, or else in which the marking systems were amplified; these techniques are well known, and in general are amplification of DNA by a polymerase; when the original sample is a RNA reverse transcription should be previously carried out. There are currently numerous processes enabling this amplification, such as for example the SDA technique (Strand Displacement Amplification) or brine displacement amplification "technique (Walker et al, 1992, Nucleic Acids Res. 20:1691), the TAS technique (Transcription-based Amplification System) described by Kwoh et al. (1989, Proc. Natl. Acad. Sci. USA, 86:1173), the 3SR technique (Self-Sustained Sequence Replication) described by Guatelli et al. (1990, Proc. Natl. Acad. Sci. USA 87:1874), the NASBA technique (Nucleic Acid Sequence Based Amplification) described by Kievitis et al.
(1991, J. Virol. Methods, 35:273), the TMA technique (Transcription Mediated Amplification), the LCR technique (Ligase Chain Reaction) described by Landegren et al. (1988, Science 241 :1077), the RCR technique (Repair Chain Reaction) described by Segev (1992, Kessler C. Springer Verlag, Berlin, New-York, 197-205), the CPR technique (Cycling Probe Reaction) described by Duck et al. (1990, Biotechniques, 9:142), the Q-beta-replicase amplification technique described by Miele et al. (1983, J. Mol. Biol., 171 :281). Certain of these techniques have since been refined. In the event where the polynucleotide target to be detected is a RNAm, prior to use of an amplification reaction by means of the primers according to the invention or prior to use of a detection process by means of the inventive probes, an enzyme of inverse transcriptase type is used advantageously in order to obtain a DNAc from the RNAm contained in the biological sample. The DNAc obtained will then serve as target for the primers or the probes used in the process of amplification or detection according to the invention. The technique of hybridization of probes can be executed in various ways
(Matthews et al, 1988, Anal. Biochem., 169:1-25). The most general method consists of immobilizing the nucleic acid extracted from cells of different tissues or cells in culture on a support (such as nitrocellulose, nylon, polystyrene) and of incubating, in well-defined conditions, the nucleic acid target immobilized with the probe. After hybridization, the probe excess is eliminated and the hybrid molecules formed are detected by the appropriate method (measuring of radioactivity, fluorescence or enzymatic activity associated with the probe). In accordance with another operating mode of the nucleic probes according to the invention, the latter can be utilized as capture probes. In this case, a probe, known as «capture probe», is immobilized on a support and serves to capmre via specific hybridization the nucleic acid target obtained from the biological sample to be tested and the nucleic acid target is then detected due to a second probe, known as (detection probe», marked by an easily detectable element. Of the possibly interesting fragments of nucleic acids, anti-sense oligonucleotides should thus be cited in particular, that is, whereof the structure ensures, via hybridization with the sequence target, inhibition of the expression of the corresponding product. Sense oligonucleotides which, by interaction with proteins implied in regulating the expression of the corresponding product, will cause either inhibition or activation of this expression, should likewise be cited.
In a preferred way the probes or primers according to the invention are immobilized on a support, covalently or non-covalently. In particular, the support can be a DNA chip or a high-density filter, likewise objects of the present invention. The interest in using DNA chips or, if required, protein chips in the domain of diagnostics or epidemiology rests as mentioned previously on the possibility of analyzing a large number of sequences at the same time and very rapidly, for: - classification or typing of bacteria as a function of the presence of a sequence or of a profile of sequences characteristic of the genre, especially of the pathogenicity or not of the genre, especially Legionella, or of the species, especially Legionella pneumophila, or specific to a bacteria of the Legionella genre and/or of the Legionella pneumophila species, or specific to a bacteria of the Legionella pneumophila subspecies Paris strain, or specific to a bacteria of the Legionella pneumophila sub-species Paris strain relative to the Philadelphia strain and/or Lens strain, or even specific to a bacteria of the Legionella pneumophila sub-species Paris and Lens strain relative to the Philadelphia strain, in particular, in association with the gravity or not of the pathologies which such bacteria can induce in case of infection in mammals, especially in humans; or - simultaneous comparison of sequence or of profile of sequences between different genres, species or strain of bacteria, pathogenic or not, allowing especially identification of a gene, or the corresponding proteic sequence, or a profile of genes whereof the presence and/or the expression in a bacteria is specific according to its genre, its species or its sub-species or strain of bacteria, and/or its pathogenicity or not. This information is largely useful especially for rapidly identifying the presence or not of a pathogenic bacteria, the gravity of the infection it can cause, the treatment adapted to an infection, and/or the necessity and the means for implementing contaminated circuits or fluids or able to be contaminated for decontaminating the objects. This information will likewise be largely useful to epidemiological studies relative to this genre of bacteria. DNA chip or high-density filter is understood to mean a support on which DNA sequences are fixed, each of them able to be marked by its geographic location. These chips or filters differ principally in their size, the material of the support, and possibly the number of DNA sequences fixed thereto. The probes or primers according to the first invention can be fixed on solid supports, in particular the DNA chips, by means of different fabrication processes. In
particular, in situ synthesis can be carried out by photochemical addressing or via ink jet. Other techniques consist of carrying out ex situ synthesis and fixing the probes on the support of the DNA chip by mechanical or electronic addressing, or by ink jet. These different processes are well known to the specialist. In effect, numerous techniques or devices for analysis of biological samples were developed in recent years, in particular for parallel analysis of several quantities of nucleic acids, especially following the development of the genomic. Among these techniques or devices, the supports enabling high-rate analysis of nucleic acids, such as biochips, or DNA chips (also called « micro- or macroarrays », or even « DNA chip ») were the object of numerous studies. These biochips can be made in particular from a support, generally solid and functionalized, on which given nucleic acids (nucleic probes) were fixed by covalent bond and localized, and on which nucleic probes are fixed specifically respectively by matching (or specific hybridization) or by recognition of an affinity site of the nucleic acids which are to be detected or identified in the biological sample. Particular examples of the documents describing techniques relative to bioDNA chips are: - the review article by Wang J. (Nucleic Acids Research, 28, 16:3011-3016, 2000), which has an abstract making the point on the main known techniques relative to DNA chips, - the patent document issued under N° US 6,030,782, which describes grafting with a mercaptosilanized surface, of nucleic acids modified by a sulhydryl or disulfide group, and the article by Bamdad (Biophysical Journal, 75:1997-2003, 1998), which describes obtaining surfaces having DNA by incoφoration of composite molecules, DNA-thiols, in auto-assembled monolayers (« self-assembled monolayers or SAMs »); - the international patent application published under N° W0 00/43539 which proposes immobilizing molecules, such as oligonucleotides, by means of polyfunctional polymers (« polymer brushes ») thus enabling the grafting density to be increased. These polymers can be obtained from hydroxyethyl, acrylamide mefhacrylate, or vinyl pyrrolidone; - the international patent application published under N° WO 00/36145 describes a fabrication method of DNA chips, comprising polymerization on a substrate of metallic layer type, a copolymer of pyrrol and functionalized pyrrol, fixing a reticulation agent on the functionalized pyrrol, then fixing a biological probe (such as an
oligonucleotide). The reticulation agent can be bifunctional, and for example have an ester function of the N-hydroxysuccinimide and a maleimide function; - the international patent application published under N° WO 98/20020 which likewise describes the high-density immobilization of nucleic acids on solid supports, this time by placing in contact of a nucleic acid containing a thiol group with support having a group reacting with this thiol, possibly by way of a reticulation agent; - the article by Penchovsky et al. (Nucleic Acids Research, 28, 22, e98, 2000), which describes a method for immobilization of oligonucleotides on animated balls of latex, by means of a reticulation agent which reacts under the action of light; and - the international patent applications published under N° WO 99/16907,
WO 00/40593 and WO 00/44939 filed by the company Surmodics (which produces lames for registering oligonucleotides functionalized with an amine). These applications describe especially the fixing of nucleic acids on surfaces such as glass, by way of a polymer skeleton to which one or more « photochemically active » groups are fixed on one side of the polymer (for grafting on the surface) and « thermochemically active » on the other side (for grafting with the functionalized nucleic acid). A nucleotidic sequence (probe or primer) according to the invention thus enables detection and/or amplification of specific nucleic sequences. In particular, detection of said sequences is made easier when the probe is fixed on a DNA chip, or to a high- density filter. The utilization of DNA chips or of high-density filters in effect helps determine expression of genes in an organism having a genomic sequence close to the genome of Legionella pneumophila Paris strain (Collection of the Pasteur Institute CIP 107-629-T). The genomic sequence of Legionella pneumophila Paris strain, completed by identification of all the genes of this organism, such as presented in the present invention, serves as a base for the construction of these DNA chips or filter. The preparation of these filters or chips consists of synthesizing oligonucleotides, corresponding to the 5' and 3' ends of the genes. These oligonucleotides are selected by utilizing the genomic sequence and its annotations divulged by the present invention. The matching temperature of these oligonucleotides at the corresponding places on the DNA must be approximately the same for each oligonucleotide. This aids in preparing fragments of DNA corresponding to each gene by the utilization of condition of appropriate PCR in a highly automated environment.
5/049642 28
The amplified fragments are then immobilized on filters or supports made of glass, silicon or synthetic polymers and these media are utilized for hybridization. The availability of such filters and/or chips and of the annotated corresponding genomic sequence allows study of the expression of large ensembles, or of the totality of genes in the microorganisms of the Legionella genre or of the Legionella pneumophila species, by preparing the complementary DNA, and by hybridizing it to the DNA or to the oligonucleotides immobilized on the filters or the chips. Likewise, the filters and/or the chips allow study of the variability of the strains or species, by preparing the DNA of these organisms and by hybridizing it to the DNA or to the oligonucleotides immobilized on the filters or the chips. The differences between the genomic sequences of the different strains or species can extensively affect the intensity of the hybridization and, as a consequence, perturb inteφretation of the results. It can thus be necessary to have the precise sequence of the genes of the strain to be studied. The method for detecting the genes described in detail hereinbelow, implying determination of the sequence of random fragments of a genome, and organizing it according to the sequences of the complete genome of the Legionella pneumophila strain divulged in the present invention, can be very useful. The nucleotide, or proteic, sequences according to the invention can be likewise utilized in DNA chips, or, if required, protein chips for performing mutation analysis. This analysis is based on the constitution of chips, especially DNA chips, capable of analyzing each base of a nucleotidic sequence according to the invention. For this puφose the techniques of micro-sequencing on DNA chip especially could be used. The mutations are detected by extension of immobilized primers hybridizing analyzed sequences to the matrix, just in a position adjacent to that of the desired mute nucleotide. A single-strand matrix, RNA or DNA, of the sequences for analysis will be advantageously prepared according to classic methods, from products amplified according to PCR-type techniques. The matrices of single-strand DNA, or RNA thus obtained are then deposited on the DNA chip, in conditions enabling their specific hybridization to the immobilized primers. A thermostable polymerase, for example Tth or the Taq DNA polymerase, specifically extends the end 3' of the primer immobilized with an analog of complementary marked nucleotide of the nucleotide in a variable position of the site; for example a thermal cycle is created in the presence of fluorescent dideoxyribonucleotides. The experimental conditions will be adapted especially to the
chips employed, to the immobilized primers, to the polymerases employed, and to the marking system selected. An advantage of microsequencing, relative to techniques based on hybridization of probes, is that it allows all the variable nucleotides to be identified with optimal discrimination in conditions of homogeneous reactions; when used on DNA chips, it permits optimal resolution and specificity for routine and industrial detection of mutations in multiplex. The utilization of high-density filters and/or chips thus provides new knowledge on the regulation of genes in organisms of industrial importance, and in particular the legionelloses propagated in diverse conditions. It also allows rapid identification of the differences between the genome of the strains utilized in multiples industrial applications. In addition, a DNA chip or a filter can be an interesting extremely tool for determination, detection and/or identification of a microorganism. Therefore, the DNA chips according to the invention which further contain at least one nucleotidic sequence of a microorganism other than Legionella pneumophila Paris strain, immobilized on the support of said chip are likewise preferred. Preferably, the selected microorganism is selected from among the species of bacteria of the Legionella genre (hereinbelow designated at times as bacteria associated with Legionella pneumophila), or the subspecies of Legionella pneumophila, or again the variants of Legionella pneumophila Paris strain. A DNA chip or a filter according to the invention is a very useful element of certain kits or is necessary for the detection and/or identification of microorganisms, in particular the bacteria belonging to the Legionella pneumophila species Paris strain or the associated microorganisms, likewise objects of the invention. Besides, the DNA chips or the filters according to the invention, containing probes or specific primers of Legionella pneumophila Paris strain, compared in particular to the Philadelphia strain, are highly advantageous elements of kits or necessary for the detection and/or quantification of the expression of genes of Legionella pneumophila Paris strain (or of associated microorganisms). In effect, control of the expression of genes is a critical point for optimizing the growth and yield of a strain, either allowing the expression of one or more novel genes, or by modifying the expression of genes already present in the cell. The present invention provides all the sequences naturally active in Legionella pneumophila Paris strain enabling the expression of the genes. It thus allows all the sequences expressed in
Legionella pneumophila Paris strain to be determined. It likewise provides a tool for locating the genes whereof the expression follows a given pattern. To achieve this, the DNA of all or part of the genes of Legionella pneumophila Paris strain can be amplified thanks to primers according to the invention, then fixed to a support, such as for example glass or nylon or a DNA chip, so as to construct a tool allowing the expression profile of these genes to be followed. This tool, constituted by this support containing the coding sequences serves as hybridization matrix to a mixture of marked molecules reflecting the RNA messengers expressed in the cell (in particular the probes marked according to the invention). By repeating this experience at different instants and by combining all these data via suitable processing, the expression profiles of all these genes are obtained. The knowledge of the sequences which follow a given regulation pattern can also be of benefit for direct searching, for example by homology, for other sequences following globally, but slightly different for the same regulation pattern. By way of complement, it is possible to isolate each control sequence present upstream of the segments serving as probes and to follow their activity by means of appropriate means such as a reference gene (luciferase, β-galactosidase, GFP for « Green Fluorescent Protein »). These isolated sequences can then be modified and assembled by metabolic engineering with sequences of interest with a view to their optimal expression. The aim of the invention likewise is the cloning and/or expression vectors, which contain a nucleotidic sequence according to the invention. In particular, those nucleotidic sequences are preferred which code for polypeptides having a cellular envelope or surface, or implied in the cellular machinery, in particular secretion, central intermediary metabolism, in particular production of sugar, energetic metabolism, the synthesis process of Vitamin B12, transcription and translation, synthesis of polypeptides. The vectors according to the invention preferably comprise elements which permit the expression and/or secretion of the nucleotidic sequences in a determined host cell. The vector must comprise a promoter, signals for initiation and termination of translation, thus of the appropriate regulation regions of transcription. It must be able to be kept stable in the host cell and can possibly have particular signals which specify secretion of the translated protein. These different elements are selected and optimized
5/049642 31
by the specialist as a function of the cellular host utilized. To this effect, the nucleotidic sequences according to the invention can be inserted into autonomous replication vectors within the selected host, or they can be integrative vectors of the selected host. Such vectors are prepared by methods currently utilized by the specialist, and the resulting clones can be introduced to an appropriate host using standard methods, such as lipofection, electroporation, thermal choc, or chemical methods. The vectors according to the invention are for example vectors of plasmidic or viral origin. They are useful for transforming host cells so as to clone or express the nucleotidic sequences according to the invention. The invention likewise comprises the host cells transformed by a vector according to the invention. The cellular host can be selected from amongst prokaryotic or eukaryotic systems, for example bacterial cells, but likewise yeast cells or animal cells, in particular the cells of mammals. The cells of insects or plant cells can likewise be used here. The host cells preferred according to the invention are in particular prokaryotic cells, preferably bacteria such as E. coli, or again belonging to the Legionella genre, to the Legionella pneumophila species Paris strain, or microorganisms associated with the Legionella pneumophila species Paris strain. The invention relates likewise to animals, except human, which comprise a cell transformed according to the invention. The transformed cells according to the invention are utilizable in preparation processes for recombinant polypeptides according to the invention. The preparation processes of a polypeptide according to the invention in recombinant form, characterized in that they utilize a vector and/or a cell transformed by a vector according to the invention are themselves included in the present invention. Preferably, a cell transformed by a vector according to the invention is cultivated in conditions permitting expression of said polypeptide and said recombinant peptide is recovered. The host cells according to the invention can likewise be utilizes for preparation of nutritive compositions, themselves an object of the present invention. As was mentioned, the cellular host can be selected from amongst prokaryotic or eukaryotic systems. In particular, it is possible to identify nucleotidic sequences according to the invention, facilitating secretion in such a prokaryotic or eukaryotic system. A vector according to the invention carrying such a sequence can thus be used advantageously for production of recombinant proteins, to be secreted. In fact,
purification of these recombinant proteins of interest will be facilitated by the fact that they are present in the supernatant of the culture cellular rather than inside the host cells. The polypeptides according to the invention can likewise be prepared by chemical synthesis. Such a preparation process is likewise an object of the invention. The specialist is aware of synthesis chemical processes, for example techniques utilizing solid phases (see especially Steward et al, 1984, Solid phase peptides synthesis, Pierce Chem. Company, Rockford, 111, 2nd ed., (1984)) or techniques utilizing partial solid phases, by condensing fragments or by synthesis in classic solution. The polypeptides obtained by chemical synthesis and capable of comprising corresponding non-natural amino acids are likewise included in the invention. The invention is further relative to hybrid polypeptides having at least one polypeptide or one of its fragments according to the invention, and a sequence of a polypeptide for inducing an immune response in humans or animals. Advantageously, the antigenic determinant is such that it is capable of inducing a humoral and/or cellular response. Such a determinant could comprise a polypeptide or one of its fragments according to the invention in glycosylated form utilized with a view to obtaining immunogenic compositions capable of inducing synthesis of antibodies directed against multiple epitopes. Said polypeptides or their glycosylated fragments likewise part of the invention. These hybrid molecules can be made up in part by a carrier molecule of polypeptides or of their fragments according to the invention, associated with a possibly immunogenic part, in particular an epitope of the diphtheric toxin, tetanic toxin, a surface antigen of the hepatitis B virus (patent FR 79 21811), the antigen VP1 of the poliomyelitus virus or any other toxin or viral or bacterial antigen. The synthesis processes of hybrid molecules encompass the methods utilized in genetics to construct hybrid nucleotidic sequences coding for the desired polypeptidic sequences. For example, reference can be made advantageously to the technique for obtaining genes coding for fusion proteins described by Minton in 1984. Said hybrid nucleotidic sequences coding for a hybrid polypeptide, as well as the hybrid polypeptides according to the invention characterized in that these are recombinant polypeptides obtained by the expression of said hybrid nucleotidic sequences, are likewise part of the invention.
5/049642 33
The invention likewise comprises the vectors characterized in that they contain one of said hybrid nucleotidic sequences. The host cells transformed by said vectors, the transgenic animals comprising one of said transformed cells as well as the preparation processes for recombinant polypeptides utilizing said vectors, said transformed cells and or said transgenic animals are likewise naturally part of the invention. The coupling between a polypeptide according to the invention and an immunogenic polypeptide can be made chemically, or biologically. Therefore, according to the invention, it is possible to introduce one or more binding element(s), especially amino acids for facilitating the coupling reactions between the polypeptide according to the invention, and the immunostimulator polypeptide, the covalent coupling of the immunostimulator antigen able to be formed at the N or C-terminal end of the polypeptide according to the invention. The bifunctional reagents allowing this coupling are determined as a function of the end selected for making this coupling, and the coupling techniques are well known to the specialist. The conjugates originating from a coupling of peptides can likewise be prepared by genetic recombination. The (conjugated) hybrid peptide can in effect be produced by recombinant DNA techniques, by insertion in or addition to the DNA sequence coding for the polypeptide according to the invention, of a sequence coding for the antigenic, immunogenic or haptenic peptide(s). These preparation techniques for hybrid peptides by genetic recombination are well known to the specialist (see for example Makrides, 1996, Microbiological Reviews 60:512-538). Preferably, said immune polypeptide is selected in the group of peptides containing anatoxins, especially diphteric toxoid or tetanic toxoid, the proteins derived from Streptococcus (as the binding protein to human seralbumin), the membranous OMPA proteins and the complexes of proteins of external membranes, the vesicles of external membranes or the proteins of thermal shocks. The hybrid polypeptides according to the invention are very useful to obtain monoclonal or polyclonal antibodies, capable of specifically recognizing the polypeptides according to the invention. In effect, a hybrid polypeptide according to the invention allows potentiation of the immune response, against the polypeptide according to the invention coupled to the immunogenic molecule. Such monoclonal or polyclonal antibodies, their fragments, or chimeric antibodies, recognizing the polypeptides according to the invention, are likewise objects of the invention.
The specific monoclonal antibodies can be obtained according to the classic method of hybridome culture described by Kδhler and Milstein (1975, Nature 256, 495). The antibodies according to the invention are for example chimeric antibodies, humanized antibodies, Fab, or F(ab')2 fragments. It can likewise be in the form of immunoconjugate or antibodies marked so as to provide a detectable and/or quantifiable signal. Therefore, the antibodies according to the invention can be employed in a process for detection and/or identification of bacteria belonging to the Legionella pneumophila species Paris and/or Lens strain and/or Philadelphia, or to an associated microorganism in a biological sample, characterized in that it comprises the following stages: a) contact of the biological sample with an antibody according to the invention; b) evidence of the possibly formed antigen-antibody complex. The antibodies according to the present invention are likewise utilizable for detecting an expression of a gene of Legionella pneumophila Paris and/or Lens strain and/or Philadelphia strain, or of associated microorganisms. In effect, the presence of the expression product of a gene recognized by a specific antibody of said product expression can be detected by the presence of an antigen-antibody complex formed after contact of the strain of Legionella pneumophila Paris, Lens or Philadelphia strain, or of the associated microorganism with an antibody according to the invention. The bacterial strain utilized can were « prepared », that is, centrifuged, lysated, placed in an appropriate reagent for the constitution of the medium prone to immunological reaction. In particular, a detection process for expression in the gene is preferred, corresponding to a Western blot, capable of being carried out after electrophoresis on polyacrylamide gel of a lysate of the bacterial strain, in the presence or in the absence of reductory conditions (SDS-PAGE). After migration and separation of the proteins on the polyacrylamide gel, said proteins are transferred to an appropriate membrane (for example made of nylon) and the presence of the protein or of the polypeptide of interest is detected, by contact of said membrane with an antibody according to the invention. Therefore, the present invention likewise comprises the kits or the necessary for implementing a process such as described (detection of the expression of a gene of Legionella pneumophila Paris and/or Lens strain and/or Philadelphia, or of an associated microorganism, or for detection and or identification of bacteria belonging to
5/049642 35
the Legionella pneumophila species Paris and/or Lens and/or Philadelphia strain, or an associated microorganism), comprising the following elements: a) a polyclonal or monoclonal antibody according to the invention; b) possibly, the reagents for the constitution of the medium prone to immunological reaction; c) possibly, the reagents allowing use of the antigen-antibody complexes produced by the immunological reaction. The polypeptides and the antibodies according to the invention can advantageously be immobilized on a support, especially a protein chip. This type of protein chip is an object of the invention, and can likewise contain at least one polypeptide of a microorganism other than Legionella pneumophila Paris and/or Lens strain and/or Philadelphia strain, or an antibody directed against a compound of a microorganism other than Legionella pneumophila Paris and/or Lens and/or Philadelphia strain. The high-density protein or filter chips containing proteins according to the invention can be constructed in the same way as the DNA chips according to the invention. In practice, synthesis of the polypeptides fixed directly on the protein chip, or ex situ synthesis followed by a stage for fixing the synthesized polypeptide on said chip can be carried out. This latter method is preferable, when proteins of significant size are to be fixed on the support, which are advantageously prepared by genetic engineering. All the same, if it is preferred to fix only peptides on the support of said chip, it can be more interesting to proceed with synthesis of said peptides directly in situ. The protein chips according to the invention can be advantageously utilized in kits or necessary for the detection and/or identification of bacteria associated with the Legionella pneumophila species Paris and/or Lens strain and/or Philadelphia strain, or with a microorganism, more generally in kits or necessary for the detection and/or identification of microorganisms. When the polypeptides according to the invention are fixed on DNA chips, the presence of antibodies is searched for in the samples tested, with the fixation of an antibody according to the invention on the support of the protein chip allowing identification of the protein whereof said antibody is specific. Preferably, an antibody according to the invention is fixed on the support of the protein chip, and the presence of the corresponding antigen, specific to Legionella pneumophila Paris and/or Lens strain and/or Philadelphia strain or of an associated microorganism is detected.
A protein chip described hereinabove can be utilized for the detection of products of genes, for establishing an expression profile of said genes, as a complement to a DNA chip according to the invention. The protein chips according to the invention are likewise extremely useful for proteomic experiments, which studies the interactions between the different proteins of a given microorganism. In a simplified manner, peptides representative of the different proteins of an organism are fixed on a support. Next, said support is put in contact with marked proteins, and after an optional rinsing stage, interactions are detected between said marked proteins and the peptides fixed on the protein chip. Therefore, the protein chips comprising a polypeptidic sequence according to the invention or an antibody according to the invention are an object of the invention, as well as kits or necessary containing them. The present invention likewise covers a process for detection and/or identification of bacteria belonging to the Legionella pneumophila species Paris and/or Lens and/or Philadelphia strain, or to an associated microorganism in a biological sample, which uses a nucleotidic sequence according to the invention. It should be understood that the term biological sample relates in the present invention to the samples taken from a living organism (in particular blood, tissue, organs or the like taken a mammal) or a sample containing biological material, that is, DNA. Such a biological sample especially includes all fluids (liquid or air-borne), or any object, such as conduits for fluid, filters for fluids, or any object capable of being implied in the fluid supply in buildings, nutritional compositions containing bacteria or other. The process for detection and/or identification utilizing the nucleotidic sequences according to the invention can be diverse in nature. A process comprising the following stages is preferred: a) possibly, isolation of DNA from the biological sample to be analyzed, or obtaining DNAc from the RNA of the biological sample; b) specific amplification of the DNA of bacteria belonging to the Legionella pneumophila species Paris strain, Lens, Philadelphia or to a microorganism associated by means of at least one primer according to the invention; c) revealing amplification products. This process is based on specific amplification of DNA, in particular by a chain amplification reaction.
Likewise, a process comprising the following stages is preferred: a) contact by a nucleotidic probe according to the invention with a biological sample, the nucleic acid contained in the biological sample having, if required, previously been made accessible to hybridization, in conditions permitting hybridization of the probe to the nucleic acid of bacteria belonging to the Legionella pneumophila species Paris strain, or to an associated microorganism; b) revealing the hybrid possibly formed between the nucleotidic probe and the DNA of the biological sample. Such a process must not be limited to detection of the presence of the DNA contained in the biological sample tested, and it can likewise be used for detecting the RNA contained in said sample. This process in particular includes the Southern and Northern blot. Another preferred process according to the invention comprises the following stages: a) contact by a nucleotidic probe immobilized on a support according to the invention with a biological sample, the nucleic acid of the sample, having, if required, been previously made accessible to hybridization, in conditions allowing hybridization of the probe to the nucleic acid of bacteria belonging to the Legionella pneumophila species Paris strain or to an associated microorganism; b) contact by the hybrid formed between the nucleotidic probe immobilized on a support and the nucleic acid contained in the biological sample, if required after elimination of the DNA from the biological sample not having hybridized with the probe, with a nucleotidic probe marked according to the invention; c) revealing the novel hybrid formed at stage b). This process is advantageously utilized with a DNA chip according to the invention, the desired nucleic acid hybridizing with a probe present on the surface of said chip, and being detected by utilization of a marked probe. This process is advantageously implemented by combining a previous amplification stage of DNA or of complementary DNA obtained possibly by inverse transcription, by means of primers according to the invention. Therefore, the present invention likewise includes the kits or necessary for the detection and/or identification of bacteria belonging to the Legionella pneumophila species Paris and or Lens and/or Philadelphia strain, or to an associated microorganism, characterized in that it comprises the following elements:
a) a nucleotidic probe according to the invention; b) possibly, the reagents necessary for using a hybridization reaction; c) possibly, at least one primer according to the invention as well as the reagents necessary for amplification reaction of the DNA. Similarly, the present invention likewise includes the kits or necessary for the detection and/or identification of bacteria belonging to the Legionella pneumophila species Paris strain or to an encircled microorganism, characterized in that it comprises the following elements: a) a nucleotidic probe, known as capture probe, according to the invention; b) an oligonucleotidic probe, known as revelation probe, according to the invention; c) possibly, at least one primer according to the invention as well as the reagents necessary for amplification reaction of the DNA. Finally, the kits or necessary for the detection and/or identification of bacteria belonging to the Legionella pneumophila species Paris and/or Lens and/or Philadelphia strain, or to an associated microorganism, characterized in that it comprises the following elements: a) at least one primer according to the invention; b) possibly, the reagents necessary for performing an amplification reaction of DNA; c) possibly, a compound enabling the sequence of the amplified fragment, more particularly an oligonucleotidic probe according to the invention, to be verified, are likewise objects of the present invention. Preferably, said primers and/or probes and or polypeptides and/or antibodies according to the present invention utilized in the processes and/or kits or necessary according to the present invention are selected from amongst the primers and/or probes and/or polypeptides and/or antibodies specific to the Legionella pneumophila species Paris and/or Lens and/or Philadelphia strain. In a preferred manner these elements are selected from amongst the nucleotidic sequences coding for a secreted protein, among the polypeptides secreted, or among the antibodies directed against exported polypeptides, such as those implied in the wall or the cellular envelope of Legionella pneumophila Paris and/or Lens and/or Philadelphia strain. The object of the present invention is likewise the strains of Legionella pneumophila Paris or Lens strain, and/or associated microorganisms containing one or
more mutation(s), at most less than 10 % mutation (or again less as cited for the modifications of polypeptides) in a nucleotidic sequence according to the invention, in particular an ORF sequence, or their regulatory elements (in particular promoters). According to the present invention, the strains of Legionella pneumophila Paris or Lens strain having one or more mutation(s) in the nucleotidic sequences coding for polypeptides implied in the machine cellular, in particular secretion, central intermediary metabolism, energetic metabolism, the process of synthesizing of the amino acids, transcription and translation, synthesis of the polypeptides, are preferred. Said mutations can lead to inactivation of the gene, or in particular when they are situated in the regulatory elements of said gene, at overexpression of the latter. The invention relates further to utilizing a compound selection method capable of inhibiting the expression of genes implied in the biosynthesis of polysaccharides having a cellular envelope of bacteria of the Legionella pneumophila species Paris strain, characterized in that it comprises the following stages: a) contact by said compound with a bacteria of said Paris strain, said bacteria being in conditions and in medium appropriate to its culture; b) determination of the capacity of said compound to inhibit the expression of the genes coding for the proteins of SEQ ID Nos. 1126, 3218, 288, 632, 917, 1503, 1555, 1877, 1928, 1963, 2204, 2212, 2243, 2324, 2378, 2410, 2411; c) by means of a process according to the invention in which said antibody is directed specifically against a polypeptide implied in the biosynthesis of the polysaccharides, or by means of a process according to the invention in which the probes or primers are specific to a nucleic sequence coding for a polypeptide implied in the biosynthesis of the polysaccharides; d) selection of organic or inorganic compound capable of modulating, regulating, inducing or inhibiting the expression of genes, and/or of modifying the cellular replication of eukaryotic or prokaryotic cells or capable of inducing, inhibiting or aggravating the pathologies associated with infection by Legionella pneumophila Paris strain or one of its associated microorganisms. The invention likewise comprises a method for selection of compounds capable of binding to a polypeptide or one of its fragments according to the invention, capable of binding to a nucleotidic sequence according to the invention, or capable of recognizing an inventive antibody, and/or capable of modulating, regulating, inducing or inhibiting the expression of genes, and or modifying the growth or cellular
5/049642 40
replication of eukaryotic or prokaryotic cells, or capable of inducing, inhibiting or aggravating in a animal or human organism the pathologies bound to infection by Legionella pneumophila Paris strain or Lens or Philadelphia strains, or one of its associated microorganisms, characterized in that it comprises the following stages: a) contact by said compound with said polypeptide, said nucleotidic sequence, with a cell transformed according to the invention and or administration of said compound to an animal transformed according to the invention; b) determination of the capacity of said compound to be bound with said polypeptide or said nucleotidic sequence, or to modulate, regulate, induce or inhibit the expression of genes, or to modulate the growth or cellular replication, or induce, inhibit or aggravate in said transformed animal the pathologies bound to infection by Legionella pneumophila Paris strain or Lens or Philadelphia strains strain, or one of its associated microorganisms. The cells and/or the animals transformed according to the invention, could advantageously serve as a model and be utilized in processes for studying, identifying and/or selecting compounds capable of being responsible for pathologies induced or aggravated by Legionella pneumophila Paris strain or Lens or Philadelphia strains strain, or capable of preventing and/or treating these pathologies such as for example genital, ocular or systemic diseases, especially of the lymphatic system. In particular, the transformed host cells, especially the bacteria of the family of Legionellae whereof the transformation by a vector according to the invention can for example grow or inhibit its infectious capacity, or modulate the pathologies habitually induced or aggravated by the infection, could be utilized for infecting animals in which the appearance of pathologies will be followed. These animals not transformed, infected for example with transformed Legionellae bacteria, will be able to serve as a study model.
In the same manner, the animals transformed according to the invention will be able to be utilized in selection processes for compounds capable of preventing and/or treating diseases due to Legionella. Said processes utilizing said transformed cells and/or transformed animals are part of the invention. The compounds capable of being selected can be organic compounds such as polypeptides or hydrates of carbon or any other already known organic or inorganic compounds, or novel organic compounds elaborated by molecular modeling techniques and obtained by chemical or biochemical synthesis, these techniques being known to the specialist.
Said selected compounds will be able to be utilized for modulating the growth and/or cellular replication of Legionella pneumophila Paris and/or Lens and/or Philadelphia strain, or any other associated microorganism and thus to control infection by these microorganisms. Said compounds according to the invention will likewise be utilized for modulating the growth and/or cellular replication of all eukaryotic or prokaryotic cells, especially tumoral cells and infectious microorganisms, for which said compounds will prove to be active, the methods determining said modulations being well known to the specialist. Compound capable of modulating the growth of a microorganism is understood to mean any compound allowing to intervene, modify, limit and/or reduce the development, growth, rate of proliferation and/or the viability of said microorganism. This modulation can be realized for example by an agent capable of binding to a protein and thus inhibit or potentialize its biological activity, or capable of binding to a membranous protein of the external surface of a microorganism and blocking penetration of said microorganism in the host cell or benefiting the action of the immune system of the infected organism directed against said microorganism. This modulation can likewise be realized by an agent capable of binding to a nucleotidic sequence of DNA or RNA of a microorganism and for example blocking the expression of a polypeptide whereof the biological or structural activity is necessary to the growth or to the reproduction of said microorganism. For these screening methods, likewise associated microorganism in the present selection method is understood to mean any microorganism whereof the gene expression can be modulated, regulated, induced or inhibited, or whereof the growth or cellular replication can be likewise modulated by a compound of the invention. Likewise, associated microorganism in the present invention is understood to mean any microorganism comprising nucleotidic sequences or polypeptides according to the invention. These microorganisms can in certain cases comprise polypeptides, or nucleotidic sequences identical or homologous to those of the invention will likewise be able to be detected and/or identified by the processes or detection and/or identification kit according to the invention and likewise serve as target for the compounds of the invention. The invention relates to the compounds capable of being selected by a selection method according to the invention.
The invention likewise relates to a pharmaceutical composition comprising a compound selected from amongst the following compounds: a) a nucleotidic sequence according to the invention; b) a polypeptide according to the invention; c) a vector according to the invention; d) an antibody according to the invention; and e) a compound capable of being selected by a selection method according to the invention, possibly in association with a pharmaceutically acceptable vehicle. Efficacious quantity is understood to mean an adequate quantity of said compound or antibodies, or of polypeptide of the invention, allowing to modulate the growth of Legionella pneumophila Paris and/or Lens and/or Philadelphia strain, or of an associated microorganism. The invention also relates to a pharmaceutical composition according to the invention for the prevention or treatment of an infection by a bacteria belonging to the Legionella pneumophila species Paris strain or Lens or Philadelphia strains, or by an associated microorganism. The further aim of the invention is an immunogenic and/or vaccinal composition, characterized in that it comprises one or more polypeptides according to the invention and/or one or more hybrid polypeptides according to the invention. The invention also comprises utilization of a cell transformed according to the invention, for the preparation of a vaccinal composition. The aim of the invention likewise is a vaccinal composition, characterized in that it contains a nucleotidic sequence according to the invention, a vector according to the invention and/or a cell transformed according to the invention. The invention likewise relates to vaccinal compositions according to the invention, for the prevention or treatment of an infection by a bacteria belonging to the Legionella pneumophila species Paris strain or Lens or Philadelphia strains, or by an associated microorganism. In a preferred manner the immunogenic and/or vaccinal compositions according to the invention for preventing and/or treating infection by Legionella pneumophila Paris strain or Lens or Philadelphia strains, or by an associated microorganism will be selected from among the immunogenic and/or vaccinal compositions comprising a polypeptide or one of its fragments corresponding to a protein, or one of its fragments, of the cellular envelope of Legionella pneumophila Paris strain or Lens or Philadelphia
strains. The vaccinal compositions comprising nucleotidic sequences will preferably likewise comprise nucleotidic sequences coding for a polypeptide or one of its fragments corresponding to a protein, or one of its fragments, of the cellular envelope of Legionella pneumophila Paris strain or Lens or Philadelphia strains. Of these preferred immunogenic and/or vaccinal compositions, the most preferred are those comprising a polypeptide or one of its fragments, or a nucleotidic sequence or one of its fragments whereof the sequences are selected from among the nucleotidic sequences or amino acids identified in this functional group and listed previously. The polypeptides of the invention or their fragments entering the immunogenic compositions according to the invention can be selected by techniques known to the specialist such as for example on the capacity of said polypeptides to stimulate the T cells, which translates for example by their proliferation or the secretion of interleukins, and which terminates with the production of antibodies directed against said polypeptides. In mice, in which a ponderal dose of the vaccinal composition comparable to the dose utilized in humans is administered, the reaction antibody is tested by taking serum followed by studying the formation of a complex between the antibodies present in the serum and the antigen of the vaccinal composition, according to customary techniques. According to the present invention, said vaccinal compositions will preferably be in association with a pharmaceutically acceptable vehicle and, if required, with one or more adjuvants of appropriate immunity. These days, diverse types of vaccines are available for protecting humans against infectious diseases: attenuated living microorganisms (M. bovis - BCG for tuberculosis), inactive microorganisms (flu virus), acellular extracts (Bordetella pertussis for pertussis), recombinant proteins (surface antigen of the hepatitis B viras), polyosides (pneumococci). Vaccines prepared from synthesis peptides or genetically modified microorganisms expressing heterologous antigens are in experimentation. Still more recently, recombined plasmidic DNAs carrying coding for protector antigens were proposed as an alternative vaccinal strategy. This type of vaccination is realized with a particular plasmid deriving from a plasmid of E. coli which does not replicate in vivo and which codes solely for the vaccinating protein. Animals were immunized by simply injecting naked plasmidic DNA into the muscle. This technique results in expression of the vaccinal protein in situ and to an immune response of cellular type (CTL) and of
5/049642 44
humoral type (antibodies). This double induction of the immune response is one of the principal advantages of the vaccination technique with naked DNA. The vaccinal compositions comprising nucleotidic sequences or vectors in which said sequences are inserted are described in particular in international application N° WO 90/11092 and likewise in international application N° WO 95/11307. The nucleotidic sequence making up the vaccinal composition according to the invention can be injected into the host after having been coupled to compounds which benefit penetration of this polynucleotide inside the cell or its transport as far as the cellular nucleus. The resulting conjugates can be encapsulated in polymer microparticles, as described in international application N° WO 94/27238 (Medisorb Technologies International). According to another embodiment of the vaccinal composition according to the invention, the nucleotidic sequence, preferably a DNA, is complexed with DEAE- dextran, with nuclear proteins, with lipids or encapsulated in liposomes or introduced in the form of a gel facilitating its transfection in the cells. The polynucleotide or the vector according to the invention can also be in suspension in a buffer solution or be associated with liposomes. Advantageously, such a vaccine will be prepared according to the technique described by Tacson et al. or Huygen et al. in 1996 or again according to the technique described by Davis et al. in the international application N° WO 95/11307. Such a vaccine can likewise be prepared in the form of a composition containing a vector according to the invention, placed under the control of regulation elements allowing its expression in humans or animals. For example, the polypeptidic antigen of interest, the pcDNA3 plasmid or the pcDNAl/neo plasmid could be utilized as an in vivo expression vector, both marketed by Invitrogen (R & D Systems, Abingdon, UK). Such a vaccine will comprise advantageously, apart from the recombinant vector, a saline solution, for example a sodium chloride solution. Pharmaceutically acceptable vehicle is understood to mean a compound or a combination of compounds entering a pharmaceutical or vaccinal composition not causing secondary reactions and which enables for example ease of administration of active compound, an increase in its life expectancy and/or its efficacy in the organism, augmentation of its solubility in solution or again an improvement in its preservation. These pharmaceutically acceptable vehicles are well known and will be adapted by the
, ,_,„05/049642 45
specialist as a function of the nature and mode of administration of the selected active compound. As for vaccinal formulations, these can comprise adjuvants of appropriate immunity which are known to the specialist, such as for example aluminum hydroxide, a representative of the family of muramyl peptides as one of the peptidic derivatives of N-acetyl-muramyl, a bacterial lysate, or even the incomplete Freund adjuvant. Preferably, these compounds will be administered systemically, in particular intravenously, intramuscularly, intradermally or subcutaneously, or orally. In a more preferred way, the vaccinal composition comprising polypeptides according to the invention will be administered in several doses, spread out over time, intradermically or subcutaneously. Their administration methods, posologies and optimal galenic forms can be determined according to the criteria generally taken into account in setting up treatment adapted to a patient such as for example age or body weight of the patient, the seriousness of the general status, tolerance to treatment and the secondary effects. The invention comprises utilizing an inventive composition, for the treatment or prevention of diseases brought on or aggravated by the presence of Legionella pneumophila Paris strain or Lens or Philadelphia strains. The invention comprises the utilization of a composition according to the invention for the treatment or prevention of systemic diseases, induced or aggravated by the presence of Legionella pneumophila Paris strain or Lens or Philadelphia strains. Additionally, an object of the present invention likewise is a genomic DNA bank of a bacteria of the species Legionella pneumophila Paris strain, characterized in that this is the bank deposited with the CNCM on November 19, 2003, under the order number 1-3138. Additionally, an object of the present invention likewise is a genomic DNA bank of a bacteria of the species Legionella pneumophila Lens strain, characterized in that this is the bank deposited with the CNCM on September 23, 2004, under the order number 1-3306. Additionally, an object of the present invention likewise is a vector or a host cell as claimed in Claim 38 or 42, characterized in that this is the vector or the cell deposited with the CNCM on November 19, 2003, under the order number 1-3137. One of the advantages of using the BAC system relative to a cosmides system is that the plasmid utilized is present only in a maximum two copies per transformed cell,
which reduces the potential for recombination between DNA fragments, and more importantly, which eliminates the risk of lethal overexpression of bacterial cloned genes. Nevertheless, the presence of BAC as a single copy signifies that the plasmidic DNA must be extracted from a large volume of culture in order to obtain enough DNA for the sequence. In addition, the stability and fidelity of maintenance of the clones in a BAC bank enable identification of genomic differences among different strains of Legionella, and identification of these genetic differences which can be responsible for the phenotypical variations observed between the different strains. The genomic DNA banks described in the present invention effectively cover the genome of Legionella pneumophila Paris and Lens strains. All the same, although it is possible that certain regions have not been able to be cloned in said bank, by virtue of lethality problems in Escherichia coli, these regions can easily be amplified and identified by the specialist, by utilizing oligonucleotides specific to the sequences of the ends of the different clones which form the contigs. Additionally, an object of the present invention likewise is a method for isolating a polynucleotide of interest present in a bacteria of the Legionella genre and absent from a bacteria of another genre, or present in a pathogenic bacteria of the Legionella genre and absent from a non-pathogenic bacteria of the Legionella genre, or again present in a bacteria of the Legionella pneumophila species and absent from a bacteria of any other species of the Legionella genre, or again present in a bacteria of the Legionella pneumophila species Paris and/or Lens and/or Philadelphia strain and absent from a bacteria of the Legionella pneumophila species of any other strain, characterized in that it utilizes at least the BAC bank deposited on November 19, 2003 (1-3138) with the C.N.C.M and the BAC bank deposited on September 23, 2004 (I- 3306) with the CNCM according to the invention. Said method is preferably characterized in that it comprises the following stages: a) isolating at least one polynucleotide contained in a clone of said DNA bank deposited with the CNCM on November 19 2003, under the order number 1-3138 or contained in a clone of said BAC bank deposited on September 23 2004 under the number 1-3306; b) isolating: at least one genomic polynucleotide or DNAc of a second bacteria of another genre or of the Legionella genre, said second bacteria of the Legionella genre belonging to a different strain of the Paris strain or, alternatively,
at least one polynucleotide contained in a clone of a DNA bank based on a BAC prepared from the genome of a second bacteria of another geme or of the
Legionella genre, said second bacteria of the Legionella geme belonging to a different strain of the Paris and/or Lens and/or Philadelphia strain; c) hybridizing the polynucleotide of stage a) to the polynucleotide of stage b); d) selecting the polynucleotides of stage a) which do not have the hybridization complex form with the polynucleotides of stage b); and e) characterizing the selected polynucleotide. The polynucleotide of stage a) can be prepared by the digestion of at least one recombinant BAC clone with an appropriate restriction enzyme, and optionally, amplification of the resulting polynucleotide insert. Therefore, the method of the invention enables the specialist to effect comparative genomic studies between the different strains or species of the Legionella geme, for example between the pathogenic strains and their non-pathogenic equivalent. In particular, it is possible to study and determine the regions of polymoφhism between said strains.
LEGENDS OF THE FIGURES Figure 1 : Circular genomic map of the line L. pneumophila Paris and specific genes of the L. pneumophila Lens line. From the exterior: circle 1 : genes of Paris line on the chains + and - respectively. Red line, inversion in line Lens. Color code: green, genes of Paris line, black, rRNA operons, red, known virulence genes; the numbers indicate their position: 1 lvh-lvr secretion system type IV (lvrABC, lvhB2B3B4B5, IvrD, lvhB6B8B9B10BHD4, IvrE); 2 dot icm secretion system type IV (icmTSRQOMLKEGCDJBF); 3 mip, 4 IspA, 5 IspDE, 6 htrA, 7 IspFGHIJK, 8 enhABC, 9 dot/icm secretion system type IV (icmVWX and dotABCD), 10 momp; circle 2: specific genes of the Lens line relative to the Paris line; 3: bias G/C (G+C/G-C) of the Paris line; circle 4: G+C content of the Paris line with <32,5% G+C in light yellow, between 32.5% and 44.1 % in yellow and with >44.1 % G+C in dark yellow. The scale (Mb) is indicated on the outside, the origin of the replication in position 0. Figure 2: Phylogenetic tree of a multiple sequential comparison of kinase domains from Legionella pneumophila of Paris line to other prokaryotic and eukaryotic kinases by utilizing the MEGA program. The calculation was made by utilizing the
Poisson correction as a distance process and as a tree construction process the Neighbor-joining distance method. 0.2 indicates the 2 amino acid substitutions for 10 sites. The Nφrot access numbers, the names of genes and the names of organisms are indicated on the pattern. The numbers indicate the priming values. Figure 3: Schema representative the genome core and the sole gene complement of L. pneumophila Paris, Lens and Philadelphia lines. The orthologous genes were defined by the most adequate reciprocated FASTA comparisons. The threshold was defined at a maximum of 80 % of sequence identities and at a length ratio of 0.75 to 1.33. The coding sequences of Philadelphia were determined by the Genmark predictions utilizing the « CAAT-box » program and the sequence obtained on the site http ://www. genome3. cpmccolumbia.edu/-legion/proi ect/ (latest version). Figure 4: A. Comparison of the protein-coding RTX genes of L. pneumophila AA100, Paris and Lens lines. The sequence of the rtxA locus of line AA100 was obtained from the NCBI database (AAD41583). The dotted lines indicate that the correct number of repetitions is uncertain. B. Consensus sequences of the highly preserved repeated patterns of Paris and Lens lines. The amino-acid sequences in black indicate 11 amino-acids of the preserved N-terminal sequence of Paris and Lens lines, the amino-acids sequences in color represent the repeated patterns of each line (same color as for A). The underlined amino-acids indicate the positions which can change among the repetitions. Figure 5: Pattern illustration of the different stages of intracellular growth of L. pneumophila in the macrophages. The different phases are numbered 1) Adhesion and invasion of L. pneumophila in the host cell 2) The phagosome does not fuse with the lysosomes but recruits organelles and converts to a compartment of rugged endoplasmic reticulum type. 3) Intracellular replication, non-flagellated L. pneumophila inside a phagosome. 4) Release of L. pneumophila. Flagellated. In red: Different important stages in the infectious cycle of L. pneumophila. In blue: Hypothesis indicating the stages at which the identified proteins could interfere in this cycle. Figures 6 and 7: Southern Blot showing the specificity of the repeated sequence SEQ ID N° 7074 in L. pneumophila; the legend of Figure 6 given by Table XXV and that of Figure 7 by Table XXVI.
EXAMPLES
Example 1 : Materials and methods
1. Construction of banks
Shotgun bank of small fragments (size 1.5 to 2.5 kb) The chromosomal DNA of the strains studied was prepared by a classic method including proteinase K treatment and phenol extraction (9). Approximately 36 ug of DNA were broken by nebulization (1 minute under pressure of 1 bar) (4). The ends of the DNA fragments were rendered free by having the DNA-polymerase of the bacteriophage T4 act for 15 minutes at 37°C in the presence of 4 tri -phosphate nucleotides. The enzyme was inactivated by incubation of 15 ran at 75°C. Adaptors (invitrogen Cat. N° 408-18) were ligatured to these ends. After ligature, the fragments of chromosomal DNA of a size between 1500 and 2500 base pairs were purified after electrophoresis on agarose gel. The vector utilized for construction of the bank, pcDNA2.1 (Invitrogen), was digested by the BstXl enzyme and purified by geneclean (BIO- 101) after electrophoresis on agarose gel. The chromosomal DNA and the purified vector were ligatured by action of the ligase of the bacteriophage T4. The ligation mixture was introduced by transformation to the strain of Escherichia coli XL2-blue (Stratagene). Environ 4000 colonies are obtained per ul of the ligation mixture. Bank of average fragments (size 5 to 10 kb) The bank was constructed by the technique of 'partial fill in' in the vector pSYX34 (12). The chromosomal DNA of the strain L. pneumophila Paris was prepared by partial digestion by SαuIIIA (Roche). After precipitation of the DNA in sodium acetate and the stage of partial fill-in with the A and G nucleotides by utilizing the
Klenow enzyme, the fragments of chromosomal DNA having a size of between 5000 and 10000 base pairs were purified after electrophoresis on agarose gel and geneclean. The vector is prepared in the same way by partial digestion with the Sail enzyme, precipitation in sodium acetate then reaction of partial fill-in with the C and T nucleotides and purification on agarose gel and geneclean. The fragments of chromosomal DNA and the purified vector were ligatured by action of the ligase of the bacteriophage T4. The ligation mixture was introduced by transformation to the strain of Escherichia coli XLIO-Gold (Stratagene). Around 4000 colonies are obtained per ul of the ligation mixture. The two ends of around 4000 fragments of this bank were sequenced. Bank of large fragments (size 25 to 90 kb) The bank of large fragments was constructed as described previously (4) by utilizing the plndigo BAC vector (Epicentre). Briefly, in order to avoid mechanical
breaking of the DNA molecules the cells were included in agarose blocks in which DNA extraction is performed directly. For the preparation of large-sized fragments we performed partial digestion by HindUT (Roche) and separation by electrophoresis in pulsed fields. Fragments of sizes between 40 and 80 and between 80 and 130 kb were excised from the gel, purified by agarase treatment and ligature with the vector. The ligation mixture was introduced by electroporation to the strain of Escherichia coli DH10B (Gibco BRL). 1300 colonies were stored. The plasmidic DNAs of these 1300 colonies were extracted and the two ends of the cloned fragments were sequenced. 2. Preparation of plasmids and sequencing. The plasmids were prepared by a semi-automatic preparation method developed at the GMP laboratory and based on the alkaline lysis method (2). The chromosomal inserts were sequenced from their two ends by utilizing the T7 and universal primer in following the recommendations of the supplier (Applied-Biosystems). The sequences were determined by utilizing automatic sequencers of type 3700 (Applied-Biosystem). 3. Assembling of sequences. The sequences were assembled by utilizing the software suite developed at the University of Washington, Phred, Phrap and Consed (5, 8). The sequence completion was done by utilizing the software suite CAAT-box (7). The finishing stage corresponds to resequencing of the regions where the sequence is only slightly secure and sequencing of the regions located between the contigs. It was carried out either by sequencing PCR products or by operating on the bank clones. The sequences of oligonucleotides were defined by utilizing the consed and Primo software (8, 10). 4. Annotation of sequences. The identification of the phases coding (CDS) was done by utilizing the software suite CAAT-box (7). This program combines the results of different methods: - (i) identification of open reading phases and their tri as a function of their size; - (ii) analysis of the probability of being coding by utilizing the Genemark software (11); (iii) identification of translation start (initiation codon and fixing sequence of the ribosome); and - (iv) the % of identity of the proteic sequence deduced with the proteic sequences contained in sequence banks by utilizing BLASTP software.
The functions of the proteins coded by the identified coding phases were predicted by analysis of the search results of similarities in the non-redundant NCBI bank (http://www.ncbi.nlm.nih.gov/BLAST/) by utilizing BLASTP software (1). 5. Comparison of the genomes - identification of the CDS specific to the strain of L. pneumophila Paris strain. All the proteic sequences deduced from the predicted coding phases of each genome was compared to all the proteic sequences possibly coded by the other genome by using BLASTP software. A threshold of 75 % of identity on the totality of the length of the protein was retained to identify the specific proteins of an isolate. This very high value was retained since it best allows discrimination of the orthologous genes from the paralogous genes (6). For the proteic sequences for which the sequence preservation is high (> at 70 %) the preservation of the nucleotidic sequences of the genes will also be high and could give a signal in low-stringency hybridization conditions. It will be necessary to consider this eventuality in the analysis of the test result. Example 2: Deposit of biological material The following organisms were deposited on November 19, 2003 with the Collection Nationale de Culmres de Microorganisms (CNCM) [National Collection of Microorganism Culmres], 25 rue du Docteur Roux, 75724 Paris Cedex 15, France, according to the dispositions of the Budapest Treaty: - Clone of a shotgun bank, clone in the pCDNA vector, of the genome of
Legionella pneumophila Paris strain (Pasteur Institute Collection CIP 107-629-T), registered under the file number 1-3137. The insert of this clone is at a size of 14.2 kb and contains a gene coding for an autotransporter called led0019A07; BAC DNA bank (1248 clones) of the genome of Legionella pneumophila Paris strain (Pasteur Institute Collection CIP 107-629-T), registered under file number 1-3138.
Said bank BAC (1-3138) was made in the E.coli DH10B strain (Grant et al, PNAS, 87:4645, 1990). The inserts of this bank were cloned in the pBelo BAC-Kan vector (Mozo et al, Mol. Gen. Genet., 1998, 258:562-70) and have an average size of between 1.5 and 2.5 kb. The total of these inserts corresponds to complete coverage of the genome.
Example 3: Annotations of sequences
1. Genes specific to L. pneumophila Paris strain relative to the L. pneumophila
Philadelphia strain
No significant identity between the nucleotidic sequence of the gene of L. pneumophila Paris strain and the genome of L. pneumophila Philadelphia strain.
Table VII: Example of annotation of sequences in the case of proteic and nucleic sequence of L. pneumophila Paris strain not having % of significant identity with respectively proteic and nucleic sequences of L. pneumophila Philadelphia strain
2. Genes common to the two strains L. pneumophila Paris strain and Philadelphia strain for which the % of identity of deduced nucleic and proteic sequences is less than 75 %
Table VIII: Example of annotation of sequences in the case of proteic and nucleic sequences of genes common to the two strains of L. pneumophila Paris strain and Philadelphia strain for which the % of identity of the deduced nucleic and proteic sequences is less than 75 %
3. Genes common to L. pneumophila Paris strain and Philadelphia strain for which the % of identity of the deduced nucleic and proteic sequences is greater than 75 %
Table IX: Example of annotation of sequences in the case of proteic and nucleic sequences of genes common to the two strains of L. pneumophila Paris strain and Philadelphia strain for which the % of identity of the deduced nucleic and proteic sequences is greater than 75 %
Example 4: Example of alignment of sequences Presented hereinbelow are the alignments of sequences preserved in the Paris and Philadelphia strains. For each of the six examples which follow, we present an alignment of the nucleotidic sequences as well as alignment of the sequences of amino acids. The alignment of the sequences of amino acids is obtained by aligning the translated sequence of the ORF present in the Paris strain with the sequence originating from translation in the six phases of the contigs of the Philadelphia sequence. The sequence homology of these ORFs present in the two strains is very strong, as much in amino acids as in nucleotides.
TBLASTN 2.2.6 [Apr-09-2003]
Query= 1764.3 CONTIG=Contig42 P0SCDS1=13736 POSCDS2=14869 SENS=p, Seq Id : 555 (216 letters)
Database: contigsLpPhiladelphia 51 sequences; 3,410,887 total letters
Searching. done Score E Sequences producing significant alignments: (bits) Value
LpPhiladelphιa_Contig49 433 e-123
>LpPhiladelphia_Contig49 Length = 376826 Score = 433 bits (1114), Expect = e-123 Identities = 215/215 (100%), Positives = 215/215 (100%) Frame = -2
Query: 1 HLALFDETYIKTILILLSICVLKKFILRYDMILNDIVLYDNFFMTFDYKDNFMSKGPYQS 60 HLALFDETYIKTILILLSICVLKKFILRYDMILNDIVLYDNFFMTFDYKDNFMSKGPYQS Sbjct: 297565 HLALFDETYIKTILILLSICVLKKFILRYDMILNDIVLYDNFFMTFDYKDNFMSKGPYQS 297386
Query: 61 FANRLISALKDRGYTASRSPNGICIKTLAEFTGASEQICRRYIRGDALPDYEKVKQLAFH 120 FANRLISALKDRGYTASRSPNGICIKTLAEFTGASEQICRRYIRGDALPDYEKVKQLAFH Sbjct: 297385 FANRLISALKDRGYTASRSPNGICIKTLAEFTGASEQICRRYIRGDALPDYEKVKQLAFH 297206
Query: 121 LQVNPGWLLFGEDENATTKKNEVDEKLLHYILKQSHHLYPISQGSNDDYADFVLGLIKEV 180 LQVNPGWLLFGEDENATTKKNEVDEKLLHYILKQSHHLYPISQGSNDDYADFVLGLIKEV Sbjct: 297205 LQV PG LLFGEDENATTKKNEVDEKLLHYILKQSHHLYPISQGSNDDYADFVLGLIKEV 297026
Query: 181 KAIDTSENNLLKIIDLAIGSISSYEEKRKKHSHAV 215 KAIDTSENNLLKIIDLAIGSISSYEEKRKKHSHAV Sbjct: 297025 KAIDTSENNLLKIIDLAIGSISSYEEKRKKHSHAV 296921
Database: contigsLpPhiladelphia Posted date: Nov 20, 2003 10:38 AM Number of letters in database: 3,410,887 Number of sequences in database: 51
Lambda K H 0.321 0.139 0.402
Gapped
Lambda K H 0.267 0.0410 0.140
Matrix: BLOSUM62
Query= 1764.3 CONTIG=Contig42 P0SCDS1=13736 POSCDS2=14869 SENS=p (1134 letters)
Database: /home/Gmp/rusniok/projets/legionella/pourBrevet- 191103/contigsLpPhiladelphia 51 sequences; 3,410,887 total letters
Searching done Score E Sequences producing significant alignments: (bits) Value
LpPhiladelphia_Contig35 2248 0.0
>LpPhiladelphia_Contig35 „ Length = 48622 Score = 2248 bits (1134), Expect = 0.0 Identities = 1134/1134 (100%) Strand = Plus / Plus
Query: 1 atgatcagaaaaataatttatgttacaggtactcgtgccgattatggactgatgagagaa 60 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 7091 atgatcagaaaaataatttatgttacaggtactcgtgccgattatggactgatgagagaa 7150
Query: 61 gtactaaaaagattacaccagtcagaagacattgacttatcgatttgtgtcactggtatg 120 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 7151 gtactaaaaagattacaccagtcagaagacattgacttatcgatttgtgtcactggtatg 7210
Query: 121 catcttgatgctttgtatggaaatacagttaacgaaattaaagcagatcagttctcaata 180 I I I I I I I I I I I I I I I I I I I I I I I I 1 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 7211 catcttgatgctttgtatggaaatacagttaacgaaattaaagcagatcagttctcaata 7270
Query: 181 tgcggcattattcctgttgatcttgccaatgctcagcatagttctatggcaaaagctatc 240 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 7271 tgcggcattattcctgttgatcttgccaatgctcagcatagttctatggcaaaagctatc 7330
Query: 241 ggccatgaacttttgggattcaccgaggtattcgaaagtgaaactcctgatgtcgtttta 300 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 7331 ggccatgaacttttgggattcaccgaggtattcgaaagtgaaactcctgatgtcgtttta 7390
Query: 301 ttgctgggagatcgaggagaaatgcttgctgcggccatagcagcgatacatttaaatatc 360 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 7391 ttgctgggagatcgaggagaaatgcttgctgcggccatagcagcgatacatttaaatatc 7450
Query: 361 ccggttgtacatctgcacggaggagagcgctctggaaccgttgatgaaatggtaaggcat 420 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 7451 ccggttgtacatctgcacggaggagagcgctctggaaccgttgatgaaatggtaaggcat 7510
Query: 421 gcgatttccaaattatctcattatcattttgtcgcaacagaggcatccaaacaacgattg 480 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 7511 gcgatttccaaattatctcattatcattttgtcgcaacagaggcatccaaacaacgattg 7570
Query: 481 attagaatgggtgagaaagaagaaaccatttttcaggttggtgctccaggcttggatgaa 540 I I I I I I I I 1 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 7571 attagaatgggtgagaaagaagaaaccatttttcaggttggtgctccaggcttggatgaa 7630
Query: 541 atcatgcagtataaaacgtctacacgtgatgtgtttaatcaacgttatggatttgatcct 600 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 7631 atcatgcagtataaaacgtctacacgtgatgtgtttaatcaacgttatggatttgatcct 7690
Query: 601 gacaaaaaaatctgtttattaatctatcacccggttgttcaagaagttgactcgattaaa 660 I I I I I I I I 111 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I 1 I I I I I I I I I I 1 I I I I I I I Sbjct: 7691 gacaaaaaaatctgtttattaatctatcacccggttgttcaagaagttgactcgattaaa 7750
Query: 661 attcaatttcaaagcgtgattcaggcagcactcgctacaaatttacagattatttgcctt 720 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 7751 attcaatttcaaagcgtgattcaggcagcactcgctacaaatttacagattatttgcctt 7810
Query: 721 gagcctaattccgatacgggtggtcatttaattcgagaagtgattcaggaatatattgat 780 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 7811 gagcctaattccgatacgggtggtcatttaattcgagaagtgattcaggaatatattgat 7870
Query: 781 catcctgatgttagaattatcaagcacttacatcgtccggaatttattgattgtcttgca 840 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I 1 I I I I I I I I I I I Sbjct: 7871 catcctgatgttagaattatcaagcacttacatcgtccggaatttattgattgtcttgca 7930
Query: 841 aattctgatgtgatgctgggaaattccagtagtggcatcatagaggcagcctcatttaac 900 1 I I I I I I I I 11 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 7931 aattctgatgtgatgctgggaaattccagtagtggcatcatagaggcagcctcatttaac 7990
Query: 901 ctgaacgtagttaatgttggaagcaggcaaaatttaagagaacgaagcgacaatgtcatt 960 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 7991 ctgaacgtagttaatgttggaagcaggcaaaatttaagagaacgaagcgacaatgtcatt 8050
Query: 961 gatgttgatgttacttatgatgctattttgactggtctaagagaagcgctaaataaaccc 1020 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 8051 gatgttgatgttacttatgatgctattttgactggtctaagagaagcgctaaataaaccc 8110
Query: 1021 aagataaaatactctaactgttatggggatggaaaaacgagtgaaaggtgttatcaattg 1080 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 8111 aagataaaatactctaactgttatggggatggaaaaacgagtgaaaggtgttatcaattg 8170
Query: 1081 ttaaaaactatccctttgcactcacaaatattgaataaatgcaatgcatactaa 1134 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I
Sbj ct : 8171 ttaaaaactatccctttgcactcacaaatattgaataaatgcaatgcatactaa 8224
TBLASTN 2.2.6 [Apr-09-2003]
Query= 1864.3 CONTIG=Contig42 POSCDS1=77740 POSCDS2=79155 SENS=p, Seq Id : 622 (489 letters)
Database: contigsLpPhiladelphia 51 sequences; 3,410,887 total letters
Searching . done Score E
Sequences producing significant alignments: (bits) Value
LpPhiladelρhia_Contig49 1003 0.0
>LpPhiladelphia_Contig49 Length = 376826 Score = 1003 bits (2594), Expect = 0.0 Identities = 488/488 (100%), Positives = 488/488 (100%)
Frame = +2
Query: 1 . KLSLPLIRLWQLSRSKHMFKPQGLYDYICQQ QEEILPSLCDYIKIPNKSPHFDAK EEH 60 ■ KLSLPLIRLWQLSRSKHMFKPQGLYDYICQQWQEEILPSLCDYIKIPNKSPHFDAKWEEH Sbjct: 21029 KLSLPLIRL QLSRSKHMFKPQGLYDYICQQWQEEILPSLCDYIKIPNKSPHFDAK EEH 21208
Query: 61 GYMEQAVNHIANWCKSHAPKGMTLEIVRLKNRTPLLFMEIPGQIDDTVLLYGHLDKQPEM 120 GYMEQAVNHIANWCKSHAPKGMTLEIVRLKNRTPLLFMEIPGQIDDTVLLYGHLDKQPEM Sbjct: 21209 GYMEQAVNHIANWCKSHAPKGMTLEIVRLKNRTPLLFMEIPGQIDDTVLLYGHLDKQPEM 21388
Query: 121 SG SDDLHPWKPVLKNGLLYGRGGADDGYSAYASLTAIRALEQQGLPYPRCILIIEACEE 180 SG SDDLHPWKPVLKNGLLYGRGGADDGYSAYASLTAIRALEQQGLPYPRCILIIEACEE Sbjct: 21389 SGWSDDLHPWKPVLKNGLLYGRGGADDGYSAYASLTAIRALEQQGLPYPRCILIIEACEE 21568
Query: 181 SGSΫDLPFYIELLKERIGKPSLVICLDSGAGNYEQL MTTSLRGNLVGKLTVELINEGVH 240 SGSYDLPFYIELLKERIGKPSLVICLDSGAGNYEQL MTTSLRGNLVGKLTVELINEGVH Sbjct: 21569 SGSYDLPFYIELLKERIGKPSLVICLDSGAGNYEQLWMTTSLRGNLVGKLTVELINEGVH 21748
Query: 241 SGSASGIVADSFRVARQLISRIEDENTGEIKLPQLYCDIPDERIKQAKQCAEILGEQVYS 300 SGSASGIVADSFRVARQLISRIEDENTGEIKLPQLYCDIPDERIKQAKQCAEILGEQVYS Sbjct: 21749 SGSASGIVADSFRVARQLISRIEDENTGEIKLPQLYCDIPDERIKQAKQCAEILGEQVYS 21928
Query: 301 EFP IDSAKPVIQDKQQLILNRTWRPALTVTGADGFPAIADAGNVMRPVTSLKLSMRLPP 360 EFPWIDSAKPVIQDKQQLILNRTWRPALTVTGADGFPAIADAGNVMRPVTSLKLSMRLPP Sbjct: 21929 EFPWIDSAKPVIQDKQQLILNRT RPALTVTGADGFPAIADAGNVMRPVTSLKLSMRLPP 22108
Query: 361 LVDPEAASVAMEKALTQNPPYNAKVDFKIQNGGSKG NAPLLSD LAKAASEASMTYYDK 420 LVDPEAASVAMEKALTQNPPYNAKVDFKIQNGGSKGWNAPLLSD LAKAASEASMTYYDK Sbjct: 22109 LVDPEAASVAMEKALTQNPPYNAKVDFKIQNGGSKGWNAPLLSDWLAKAASEASMTYYDK 22288
Query: 421 PAAYMGEGGTIPFMSMLGEQFPKAQFMITGVLGPHSNAHGPNEFLHLDMVKKLTSCVSYV 480 PAAYMGEGGTIPFMSMLGEQFPKAQFMITGVLGPHSNAHGPNEFLHLDMVKKLTSCVSYV Sbjct: 22289 PAAYMGEGGTIPFMSMLGEQFPKAQFMITGVLGPHSNAHGPNEFLHLDMVKKLTSCVSYV 22468
Query: 481 LYSFSQKK 488 LYSFSQKK Sbjct: 22469 LYSFSQKK 22492
Database: contigsLpPhiladelphia Posted date: Nov 20, 2003 10:38 AM Number of letters in database: 3,410,887 Number of sequences in database: 51
Lambda K H 0 . 318 0. 136 0. 419
Gapped
Lambda K H 0.267 0.0410 0.140
Matrix: BLOSUM62
Query= 1864.3 CONTIG=Contig42 POSCDS1=77740 POSCDS2=79155 SENS=p (1416 letters)
Database: /hbme/Gmp/rusniok/projets/legionella/pourBrevet- 191103/contigsLpPhiladelphia 51 sequences; 3,410,887 total letters
Searching done Score E Sequences producing significant alignments: (bits) Value
LpPhiladelphia_Contig49 2807 0.0
>LpPhiladelphia_Contig49 Length = 376826 Score = 2807 bits (1416), Expect = 0.0 Identities = 1416/1416 (100%) Strand = Plus / Plus
Query: 1 atgttcaaaccccaaggattgtatgattacatatgccaacagtggcaagaagagatattg 60 I I I I I I I I I I I I I I I I I I II I I I I I I I I 1 I I I I I I I I I I I I I I I 1 I I I I I II I I I II I I I Sbjct: 21080 atgttcaaaccccaaggattgtatgattacatatgccaacagtggcaagaagagatattg 21139
Query: 61 ccaagtttatgtgactacataaaaatccctaataaatctcctcactttgatgcaaaatgg 120 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I II I I I I I Sbjct: 21140 ccaagtttatgtgactacataaaaatccctaataaatctcctcactttgatgcaaaatgg 21199
Query: 121 gaagaacatggttatatggagcaggcagttaatcacattgccaattggtgtaagtcgcat 180 I I I I I I I I I I I I I I I I I II I I I I I I I I I I I I I I II I I II I I I II I I I I I I I II II I I I I I Sbjct: 21200 gaagaacatggttatatggagcaggcagttaatcacattgccaattggtgtaagtcgcat 21259
Query: 181 gctcccaaaggaatgactctggaaattgttcgcctgaaaaataggactccattactattt 240 II I I II I I I I II II II I I I II I I II I I I I II I I I I I I I I I I I I I I I I I I II I I I I I I I I I Sbjct: 21260 gctcccaaaggaatgactctggaaattgttcgcctgaaaaataggactccattactattt 21319
Query: 241 atggaaattccaggccaaattgatgacactgtgttgctttatgggcacttggataaacaa 300 I I I I I I I I 11 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 21320 atggaaattccaggccaaattgatgacactgtgttgctttatgggcacttggataaacaa 21379
Query: 301 cctgagatgtcaggctggagtgacgatttacatccatggaaacccgtattgaaaaatgga 360 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I II I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 21380 cctgagatgtcaggctggagtgacgatttacatccatggaaacccgtattgaaaaatgga 21439
Query: 361 ttgttatacggaagaggaggggcagatgatggatattctgcttatgcatcactcacggct 420 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I II I II I I II II I I I I I II I I I I I I I Sbjct: 21440 ttgttatacggaagaggaggggcagatgatggatattctgcttatgcatcactcacggct 21499
Query: 421 attcgcgccttggaacagcaaggtttgccatatcctcgttgtatattaatcatcgaagcg 480 I I II I I I I II I I I II I I I I I I I I I I I I I I I I I I I II I I I I I I I I I I I I I I I I II I I I II I Sbjct: 21500 attcgcgccttggaacagcaaggtttgccatatcctcgttgtatattaatcatcgaagcg 21559
Query: 481 tgtgaggaaagtggcagttacgatttgcctttttatattgagttgctgaaagagcgtatt 540 I I I I I I I I I I I I I II I I I I I II I I I I I 1 I I I 1 I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 21560 tgtgaggaaagtggcagttacgatttgcctttttatattgagttgctgaaagagcgtatt 21619
Query: 541 ggtaaaccatcattggttatttgtcttgattccggagcaggtaattatgagcagttatgg 600 I I I I I I I I I I 1 I I II I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I II I I I I I I I I I I Sbjct: 21620 ggtaaaccatcattggttatttgtcttgattccggagcaggtaattatgagcagttatgg 21679
Query: 601 atgactacgtcattacgcggtaatttggtcggtaagttaactgttgaattaattaatgag 660 I I I I I I I II I I I I II I I I I I I I I I I I I II I II I I I I I I I I I I I I I II I I I I I II II I I I I Sbjct: 21680 atgactacgtcattacgcggtaatttggtcggtaagttaactgttgaattaattaatgag 21739
Query: 661 ggcgttcattctgggagcgccagtggtatagtggcagacagtttcagagtagctcggcaa 720 I I I I I I I M I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I II Sbjct: 21740 ggcgttcattctgggagcgccagtggtatagtggcagacagtttcagagtagctcggcaa 21799
Query: 721 ttgatcagcaggatagaggacgaaaacaccggagagataaaattacctcagttgtattgt 780 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 21800 ttgatcagcaggatagaggacgaaaacaccggagagataaaattacctcagttgtattgt 21859
Query: 781 gatattcctgatgagagaataaaacaagcgaaacaatgtgcggaaattctaggtgaacaa 840 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 21860 gatattcctgatgagagaataaaacaagcgaaacaatgtgcggaaattctaggtgaacaa 21919
Query: 841 gtttatagcgaatttccatggatagattctgccaaacccgttattcaagacaaacagcaa 900 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 21920 gtttatagcgaatttccatggatagattctgccaaacccgttattcaagacaaacagcaa 21979
Query: 901 ttaatattaaacagaacatggcgccctgccttgacggtgactggtgcagatgggtttcca 960 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 21980 ttaatattaaacagaacatggcgccctgccttgacggtgactggtgcagatgggtttcca 22039
Query: 961 gcgatagctgatgcagggaacgtaatgcgccctgttacgtctttgaaattatccatgcgc 1020 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 22040 gcgatagctgatgcagggaacgtaatgcgccctgttacgtctttgaaattatccatgcgc 22099
Query: 1021 cttccaccactggttgatccagaagcagcttctgttgctatggaaaaagccctgacccaa 1080 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 22100 cttccaccactggttgatccagaagcagcttctgttgctatggaaaaagccctgacccaa 22159
Query: 1081 aaccctccctataatgcaaaggttgattttaaaatacaaaatggagggtccaagggatgg 1140 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 22160 aaccctccctataatgcaaaggttgattttaaaatacaaaatggagggtccaagggatgg 22219
Query: 1141 aatgctcctttgctttccgattggttagcgaaagcggcatctgaagcatcaatgacttat 1200 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 22220 aatgctcctttgctttccgattggttagcgaaagcggcatctgaagcatcaatgacttat 22279
Query: 1201 tatgataaacctgctgcttacatgggagaggggggcaccattccatttatgagtatgcta 1260 I I I I I I I I I I I I I I I I I I I I I I I I I II I I I I I I I I I I I I I I I I I I II I I I I I I I I I I I I I Sbjct: 22280 tatgataaacctgctgcttacatgggagaggggggcaccattccatttatgagtatgcta 22339
Query: 1261 ggcgagcaatttcccaaagcacaatttatgataactggtgttttaggcccccattccaat 1320 I I I I I I I I I I I I I I I I I I I I I I I I I II I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 22340 ggcgagcaatttcccaaagcacaatttatgataactggtgttttaggcccccattccaat 22399
Query: 1321 gctcatggtccgaacgagttcttacatttggacatggtaaaaaaactcacctcatgtgtc 1380 I II I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 22400 gctcatggtccgaacgagttcttacatttggacatggtaaaaaaactcacctcatgtgtc 22459
Query: 1381 tcgtacgttctttatagtttttcacagaaaaaataa 1416 I I I II I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 22460 tcgtacgttctttatagtttttcacagaaaaaataa 22495
TBLASTN 2.2.6 [Apr-09-2003J
Query= 1865.3 CONTIG=Contig42 P0SCDS1=76674 POSCDS2=77765 SENS=p, Seq Id : 623 (367 letters)
Database: contigsLpPhiladelphia 51 sequences; 3,410,887 total letters
Searching . done Score E Sequences producing significant alignments: (bits) Value
LpPhiladelphia_Contig49 718 0.0
>LpPhiladelphia_Contig49 Length = 376826 Score = 718 bits (1853), Expect = 0.0 Identities = 366/366 (100%), Positives = 366/366 (100%) Frame = +1
Query: 1 GNIMSPΞIVFTGGGTAGHVTPNIALIKEFRKEG NVEYIGSVSGIEKEMIEPLDIPFHGV 60 GNIMSPSIVFTGGGTAGHVTPNIALIKEFRKEGWNVEYIGSVSGIEKEMIEPLDIPFHGV Sbjct: 20005 GNIMSPSIVFTGGGTAGHVTPNIALIKEFRKEG NVEYIGSVSGIEKEMIEPLDIPFHGV 20184
Query: 61 SSGKLRRYFSLKNLLDPFKIVLGIIQSSLLFYKIKPDVVFSKGGFVAFPVWGA LNRIP 120 SSGKLRRYFSLKNLLDPFKIVLGIIQSSLLFYKIKPDVVFSKGGFVAFPVWGA LNRIP Sbjct: 20185 SSGKLRRYFSLKNLLDPFKIVLGIIQSSLLFYKIKPDWFSKGGFVAFPWVGA LNRIP 20364
Query: 121 VVAHESDMSPGLANRLSFPFVNKICLTFDAGKKYFKRQDKIEVTGTPIRQQLLTGNRMKG 180 VVAHESDMSPGLANRLSFPFVNKICLTFDAGKKYFKRQDKIEVTGTPIRQQLLTGNRMKG Sbjct: 20365 WAHESDMSPGLANRLSFPFVNKICLTFDAGKKYFKRQDKIEVTGTPIRQQLLTGNRMKG 20544
Query: 181 LELCGFNSSKPCLLVVGGSLGAGSINSCIRSALKQLTSEFQVIHLCGKGKLDSSLVGVEG 240 LELCGFNSSKPCLLVVGGSLGAGSINSCIRSALKQLTSEFQVIHLCGKGKLDSSLVGVEG Sbjct: 20545 LELCGFNSSKPCLLWGGSLGAGSINSCIRSALKQLTSEFQVIHLCGKGKLDSSLVGVEG 20724
Query: 241 YCQFEYANEELADLFAASSVVISRAGANSLYEILALGKPHILIPISSQVSRGDQIQNARY 300 YCQFEYANEELADLFAASSWISRAGANSLYEILALGKPHILIPISSQVSRGDQIQNARY Sbjct: 20725 YCQFEYANEELADLFAASSWISRAGANSLYEILALGKPHILIPISSQVSRGDQIQNARY 20904
Query: 301 FQGLGISVVIQDELLKADVLLQAVQDVMRKKDEIDNKIKALKIESATDKIVAIIKEQAHV 360 FQGLGISWIQDELLKADVLLQAVQDVMRKKDEIDNKIKALKIESATDKIVAIIKEQAHV Sbjct: 20905 FQGLGISWIQDELLKADVLLQAVQDVMRKKDEIDNKIKALKIESATDKIVAIIKEQAHV 21084
Query: 361 QTPRIV 366 QTPRIV Sbjct: 21085 QTPRIV 21102
Database: contigsLpPhiladelphia Posted date: Nov 20, 2003 10:38 AM Number of letters in database: 3,410,887 Number of sequences in database: 51
Lambda K H 0.321 0.139 0.399
Gapped
Lambda K H 0.267 0.0410 0.140
Matrix: BLOSUM62
Query= 1865.3 CONTIG=Contig 2 P0SCDS1=76674 POSCDS2=77765 SENS=p (1092 letters)
Database : /home/G p/rusniok/projets/legionella/pourBrevet- 191103/contigsLpPhiladelphia 51 sequences; 3,410,887 total letters
Searching done Score E Sequences producing significant alignments: (bits) Value
LpPhiladelphia_Contig49 2165 0.0
>LpPhiladelphia_Contig49 Length = 376826 Score = 2165 bits (1092), Expect = 0.0 Identities = 1092/1092 (100%) Strand = Plus / Plus
Query: 1 atgagcccaagtattgtttttaccgggggaggaactgccggacatgtaacgcctaatatc 60 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 20014 atgagcccaagtattgtttttaccgggggaggaactgccggacatgtaacgcctaatatc 20073
Query: 61 gctttgattaaggaatttcgaaaagaaggctggaatgtagaatatatcggctctgtttcc 120 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I II I I I I I Sbjct: 20074 gctttgattaaggaatttcgaaaagaaggctggaatgtagaatatatcggctctgtttcc 20133
Query: 121 ggaattgaaaaggagatgattgagccgctggacattccttttcatggggtcagtagcggt 180 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 20134 ggaattgaaaaggagatgattgagccgctggacattccttttcatggggtcagtagcggt 20193
Query: 181 aaattgcgcaggtattttagtttgaagaacttgcttgatcctttcaaaattgttctggga 240 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 20194 aaattgcgcaggtattttagtttgaagaacttgcttgatcctttcaaaattgttctggga 20253
Query: 241 attattcaatcttctttgctattttataaaatcaaacccgatgtggttttttcaaaaggt 300 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 20254 attattcaatcttctttgctattttataaaatcaaacccgatgtggttttttcaaaaggt 20313
Query: 301 ggctttgtagcctttcctgtggttgtaggcgcctggttaaatcgaattcctgttgtcgct 360 I I I I I I I I II I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I II I I I I I Sbjct: 20314 ggctttgtagcctttcctgtggttgtaggcgcctggttaaatcgaattcctgttgtcgct 20373
Query: 361 catgagtctgatatgagcccaggacttgcgaatcgcctatcctttcctttcgtcaataaa 420 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 20374 catgagtctgatatgagcccaggacttgcgaatcgcctatcctttcctttcgtcaataaa 20433
Query: 421 atatgtcttacttttgatgctggcaaaaaatactttaagcgtcaggataaaatagaagtg 480 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 20434 atatgtcttacttttgatgctggcaaaaaatactttaagcgtcaggataaaatagaagtg 20493
Query: 481 acgggtactccaattcgtcaacagctattaactggaaatcgaatgaaaggattggagtta 540 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 20494 acgggtactccaattcgtcaacagctattaactggaaatcgaatgaaaggattggagtta 20553
Query: 541 tgcggatttaattcctccaaaccttgcctgcttgtagtgggaggaagcttaggggctggt 600 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 20554 tgcggatttaattcctccaaaccttgcctgcttgtagtgggaggaagcttaggggctggt 20613
Query: 601 tcaattaacagttgtattcgaagcgcattgaaacaattgacatcagaatttcaagtcatt 660 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 20614 tcaattaacagttgtattcgaagcgcattgaaacaattgacatcagaatttcaagtcatt 20673
Query: 661 catctttgtggcaagggaaaacttgattcttcattggttggtgtggagggatattgccaa 720 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 20674 catctttgtggcaagggaaaacttgattcttcattggttggtgtggagggatattgccaa 20733
Query: 721 tttgaatacgccaatgaagagttggctgatctgttcgctgcttcttctgtggtgatttct 780 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 20734 tttgaatacgccaatgaagagttggctgatctgttcgctgcttcttctgtggtgatttct 20793
Query: 781 cgagcaggagctaattctttgtatgaaatattagcattaggaaaaccacatatcttaatt 840 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 20794 cgagcaggagctaattctttgtatgaaatattagcattaggaaaaccacatatcttaatt 20853
Query: 841 ccaatctcttcacaagtaagcagaggagatcaaattcagaatgcaaggtacttccaggga 900 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 20854 ccaatctcttcacaagtaagcagaggagatcaaattcagaatgcaaggtacttccaggga 20913
Query: 901 ttgggaataagcgttgtgattcaggacgagttattgaaagctgatgttctattacaggca 960 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 20914 ttgggaataagcgttgtgattcaggacgagttattgaaagctgatgttctattacaggca 20973
Query: 961 gtacaggacgtaatgcgaaaaaaagatgaaatagataataaaatcaaagcattaaaaatt 1020 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 20974 gtacaggacgtaatgcgaaaaaaagatgaaatagataataaaatcaaagcattaaaaatt 21033
Query: 1021 gagtctgccactgataagattgtggcaattatcaaggagcaagcacatgttcaaacccca 1080 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 21034 gagtctgccactgataagattgtggcaattatcaaggagcaagcacatgttcaaacccca 21093
Query: 1081 aggattgtatga 1092 I I I I I I I I I I I I Sbjct: 21094 aggattgtatga 21105
TBLASTN 2.2.6 [Apr-09-2003 ]
Query= 2066.5 C0NTIG=Contig46 P0SCDS1=56766 POSCDS2=57173 SENS=p, Seq Id : 732 (150 letters)
Database: contigsLpPhiladelphia 51 sequences; 3,410,887 total letters
Searching. done Score E Sequences producing significant alignments: (bits) Value
LpPhiladelphia_Contig49 323 4e-90
>LpPhiladelphia_Contig49 Length = 376826 Score = 323 bits (828), Expect = 4e-90 Identities = 149/149 (100%), Positives = 149/149 (100%) Frame = +1
Query: 1 IMYLRLLALSALCFVTSPIWSFTCIYTLVKDNCWTDYDVTVDVIEDSTSKTLLTLTAPKG 60 IMYLRLLALSALCFVTSPI SFTCIYTLVKDNC TDYDVTVDVIEDSTSKTLLTLTAPKG Sbjct: 89377 IMYLRLLALSALCFVTSPI SFTCIYTLVKDNCWTDYDVTVDVIEDSTSKTLLTLTAPKG 89556
Query: 61 KSWARGTFNCEAAEGLRYVAQFSPVF QNDVGKTYPALRNWYLPAKVNPGDLA TIPVCF 120 KSWARGTFNCEAAEGLRYVAQFSPVFWQNDVGKTYPALRNWYLPAKVNPGDLAWTIPVCF Sbjct: 89557 KS ARGTFNCEAAEGLRYVAQFSPVFWQNDVGKTYPALRNWYLPAKVNPGDLAWTIPVCF 89736
Query: 121 PADFAQVPFPPNVAGNCKCNFKNIPDPKL 149 PADFAQVPFPPNVAGNCKCNFKNIPDPKL Sbjct: 89737 PADFAQVPFPPNVAGNCKCNFKNIPDPKL 89823
Database: contigsLpPhiladelphia Posted date: Nov 20, 2003 10:38 AM Number of letters in database: 3,410,887 Number of sequences in database: 51
Lambda K H 0.323 0. 138 0. 473
Gapped
Lambda K H 0.267 0.0410 0.140
Matrix: BLOSUM62
Query= 2066.5 CONTIG=Contig46 P0SCDS1=56766 POSCDS2=57173 SENS=p (408 letters)
Database: /home/Gmp/rusniok/projets/legionella/pourBrevet- 191103/contigsLpPhiladelphia 51 sequences; 3,410,887 total letters
Searching done Score E Sequences producing significant alignments: (bits) Value
LpPhiladelphia_Contig49 809 0.0
>LpPhiladelphia_Contig49 Length = 376826 Score = 809 bits (408), Expect = 0.0 Identities = 408/408 (100%)
Strand = Plus / Plus
Query: 1 gtgactagcccaatttggtctttcacatgcatctatactttggttaaagacaattgttgg 60 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I II I I I I I I I I I I I I Sbjct: 89419 gtgactagcccaatttggtctttcacatgcatctatactttggttaaagacaattgttgg 89478
Query: 61 actgattatgatgttactgtcgatgtcattgaagattctacgtcaaaaactttgttgaca 120 I I I I I I I I I I I I I I I I I II I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 89479 actgattatgatgttactgtcgatgtcattgaagattctacgtcaaaaactttgttgaca 89538
Query: 121 cttaccgctcccaaaggaaaatcatgggctagaggtactttcaattgtgaggctgctgaa 180 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 89539 cttaccgctcccaaaggaaaatcatgggctagaggtactttcaattgtgaggctgctgaa 89598
Query: 181 gggttgagatatgtcgctcaattttcgcctgtcttttggcaaaatgatgttggaaaaact 240 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I II I I I I I I I I I I I I I I I I Sbjct: 89599 gggttgagatatgtcgctcaattttcgcctgtcttttggcaaaatgatgttggaaaaact 89658
Query: 241 tacccggcattaagaaattggtatttaccagcaaaagtgaatcctggagatttggcctgg 300 I I I I I I I I I I I I I I I I II I I I I I I I I I I I I I I I I I I I I II I I I I I I I I I I I I I I I I I I I I Sbjct: 89659 tacccggcattaagaaattggtatttaccagcaaaagtgaatcctggagatttggcctgg 89718
Query: 301 actatcccggtttgttttccggcagattttgctcaagttccctttccacctaatgtagca 360 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 89719 actatcccggtttgttttccggcagattttgctcaagttccctttccacctaatgtagca 89778
Query: 361 ggaaactgtaagtgcaacttcaagaacattcctgatcccaagctttaa 408 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 89779 ggaaactgtaagtgcaacttcaagaacattcctgatcccaagctttaa 89826
TBLASTN 2.2.6 [Apr-09-2003]
Query= 3159.2 CONTIG=Contig46 P0SCDS1=34563 POSCDS2=35318 SENS=p, Seq Id : 1433 (265 letters)
Database: contigsLpPhiladelphia 51 sequences; 3,410,887 total letters
Searching . done Score E Sequences producing significant alignments: (bits) Value
LpPhiladelphia_Contig49 537 e-154
>LpPhiladelphia_Contig49 Length = 376826 Score = 537 bits (1383), Expect = e-154 Identities = 264/264 (100%), Positives = 264/264 (100%) Frame = +1
Query: 1 CWESFFLILLFPM KILYQLASPKNFYNYAGRLIP LAVSALTTMAIGMV GLVFAPPD 60 CWESFFLILLFPMWKILYQLAΞPKNFYNYAGRLIPWLAVSALTTMAIGMVWGLVFAPPD Sbjct: 67177 CWESFFLILLFPM KILYQLASPKNFYNYAGRLIPWLAVSALTTMAIGMV GLVFAPPD 67356
Query: 61 YQQGDAYRIIFVHVPSAFLSMALYA MGFLAILLLV RIKMAGLLIHKVAQLGACMAFLA 120 YQQGDAYRIIFVHVPSAFLSMALYA MGFLAILLLVWRIKMAGLLIHKVAQLGACMAFLA , Sbjct: 67357 YQQGDAYRIIFVHVPSAFLSMALYAWMGFLAILLLVWRIKMAGLLIHKVAQLGACMAFLA 67536
Query: 121 LITGSIWGKPMWGAWWVWDARLTSELILLLLYLAILATYQAVKNKEDGDKIIAILALVGL 180 LITGSI GKPMWGAWWV DARLTSELILLLLYLAILATYQAVKNKEDGDKIIAILALVGL Sbjct: 67537 LITGSI GKPM GA V DARLTSELILLLLYLAILATYQAVKNKEDGDKIIAILALVGL 67716
Query: 181 IDLPIIHYSVY WNTLHQGATLSVFAKPKIALSMLYPLLITLLGFFLYSL IILEKARNE 240 IDLPIIHYSVYW NTLHQGATLSVFAKPKIALSMLYPLLITLLGFFLYSL IILEKARNE Sbjct: 67717 IDLPIIHYSVY WNTLHQGATLSVFAKPKIALSMLYPLLITLLGFFLYSL IILEKARNE 67896
Query: 241 VLFRERKQSWVKIQFEEESDESVF 264 VLFRERKQS VKIQFEEESDESVF Sbjct: 67897 VLFRERKQSWVKIQFEEESDESVF 67968
Database: contigsLpPhiladelphia Posted date: Nov 20, 2003 10:38 AM Number of letters in database: 3,410,887 Number of sequences in database: 51
Lambda K H 0.330 0.143 0.468
Gapped
Lambda K H 0.267 0.0410 0.140
Matrix: BLOSUM62
Query= 3159.2 CONTIG=Contig46 P0SCDS1=34563 POSCDS2=35318 SENS=p (756 letters)
Database: /home/Gmp/rusniok/projets/legionella/pourBrevet- 191103/contigsLpPhiladelphia 51 sequences; 3,410,887 total letters
Searching done Score E Sequences producing significant alignments: (bits) Value
LpPhiladelphia_Contig49 1499 0.0
>LpPhiladelphia_Contig49 Length = 376826 Score = 1499 bits (756), Expect = 0.0 Identities = 756/756 (100%) Strand = Plus / Plus
Query: 1 atgtggaagatattgtatcagttggcatcgccaaaaaatttttataactacgcgggacgt 60 II I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 67216 atgtggaagatattgtatcagttggcatcgccaaaaaatttttataactacgcgggacgt 67275
Query: 61 ctcattccctggttggcagtcagtgctttgactaccatggccattggtatggtttgggga 120 II I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 67276 ctcattccctggttggcagtcagtgctttgactaccatggccattggtatggtttgggga 67335
Query: 121 ttggtatttgctccaccagattatcagcaaggggatgcataccgaattatttttgttcat 180 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I
Sbjct: 67336 ttggtatttgctccaccagattatcagcaaggggatgcataccgaattatttttgttcat 67395
Query: 181 gtacccagcgcttttttatcaatggcattgtatgcctggatggggtttctggccatttta 240 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I
Sbjct: 67396 gtacccagcgcttttttatcaatggcattgtatgcctggatggggtttctggccatttta 67455
Query: 241 ttgttggtgtggcgtatcaaaatggcagggcttttgattcataaggtcgcgcaattaggt 300 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I
Sbjct: 67456 ttgttggtgtggcgtatcaaaatggcagggcttttgattcataaggtcgcgcaattaggt 67515
Query: 301 gcctgcatggcatttcttgctttaattacagggagcatttggggtaaacccatgtggggt 360 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I
Sbjct: 67516 gcctgcatggcatttcttgctttaattacagggagcatttggggtaaacccatgtggggt 67575
Query: 361 gcctggtgggtatgggatgcccgcctgacctcagaattaatacttttgttgctctatctg 420 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I
Sbjct: 67576 gcctggtgggtatgggatgcccgcctgacctcagaattaatacttttgttgctctatctg 67635
Query: 421 gcaattctggctacctatcaagcggtaaaaaataaagaagatggagataaaataatagca 480 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I
Sbjct: 67636 gcaattctggctacctatcaagcggtaaaaaataaagaagatggagataaaataatagca 67695
Query: 481 attttagctttggtgggtttaattgatttaccaataattcattattcagtttattggtgg 540 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I
Sbjct: 67696 attttagctttggtgggtttaattgatttaccaataattcattattcagtttattggtgg 67755
Query: 541 aatactttacaccaaggtgcaactttatctgtgtttgccaaacccaaaattgctctcagt 600 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I
Sbjct: 67756 aatactttacaccaaggtgcaactttatctgtgtttgccaaacccaaaattgctctcagt 67815
Query: 601 atgttgtatccattgttaatcactttgctgggttttttcttgtattccttatggatcatt 660 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I
Sbjct: 67816 atgttgtatccattgttaatcactttgctgggttttttcttgtattccttatggatcatt 67875
Query: 661 ttggaaaaagcacgtaatgaagtcttattcagggagagaaagcaatcatgggttaagatt 720 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I
Sbjct: 67876 ttggaaaaagcacgtaatgaagtcttattcagggagagaaagcaatcatgggttaagatt 67935
Query: 721 caatttgaggaagagtctgatgaatcagttttttga 756 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 67936 caatttgaggaagagtctgatgaatcagttttttga 67971
TBLASTN 2.2.6 [Apr-09-2003]
Query= 4774.1 CONTIG=Contig46 POSCDS1=50654 POSCDS2=50950 SENS=m, Seq Id : 2523
(103 letters )
Database: contigsLpPhiladelphia 51 sequences; 3,410,887 total letters
Searching . done Score E Sequences producing significant alignments: (bits) Value
LpPhiladelphia_Contig49 205 4e-55
>LpPhiladelphia_Contig49 Length = 376826 Score = 205 bits (522), Expect = 4e-55 Identities = 102/102 (100%), Positives = 102/102 (100%) Frame = -1
Query: 1 RRSKMPEIHTLDNPYITILTIFVLACFVGYYW KVTPALHTPLMSVTNAISSIIILGAL 60 RRSKMPEIHTLDNPYITILTIFVLACFVGYYWWKVTPALHTPLMSVTNAISSIIILGAL Sbjct: 83615 RRSKMPEIHTLDNPYITILTIFVLACFVGYYW KVTPALHTPLMSVTNAISSIIILGAL 83436
Query: 61 IAAGSELIGCITWLGGIAIFITSINIFGGFWTQRMLRMYKK 102 IAAGSELIGCITWLGGIAIFITSINIFGGFWTQRMLRMYKK Sbjct: 83435 IAAGSELIGCITWLGGIAIFITSINIFGGFWTQRMLRMYKK 83310
Database: contigsLpPhiladelphia Posted date: Nov 20, 2003 10:38 AM Number of letters in database: 3,410,887 Number of sequences in database: 51
Lambda K H 0 . 330 0 . 144 0 . 442
Gapped
Lambda K H 0.267 0.0410 0.140
Matrix: BLOSUM62
Query= 4774.1 CONTIG=Contig46 POSCDS1=50654 POSCDS2=50950 SENS=m (297 letters)
Database: /home/Gmp/rusniok/projets/legionella/pour Brevet- 191103/contigsLpPhiladelphia 51 sequences; 3,410,887 total letters
Searching done Score E Sequences producing significant alignments: (bits) Value
LpPhiladelphia_Contig49 589 e-169
>LpPhiladelphia_Contig49 Length = 376826 Score = 589 bits (297), Expect = e-169 Identities = 297/297 (100%) Strand = Plus / Minus
Query: 1 atgcctgaaattcatacacttgataatccttatattacaatattaaccattttcgtactg 60 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 83603 atgcctgaaattcatacacttgataatccttatattacaatattaaccattttcgtactg 83544
Query: 61 gcctgttttgtaggttattatgtggtttggaaagtaacaccggctttacatacaccccta 120 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 83543 gcctgttttgtaggttattatgtggtttggaaagtaacaccggctttacatacaccccta 83484
Query: 121 atgtcagtaaccaatgccatatccagtattattatacttggtgctttaattgctgcagga 180 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 83483 atgtcagtaaccaatgccatatccagtattattatacttggtgctttaattgctgcagga 83424
Query: 181 agtgaattgatcggatgcataacctggttaggtggcatagccatattcattacttcaatt 240 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 83423 agtgaattgatcggatgcataacctggttaggtggcatagccatattcattacttcaatt 83364
Query: 241 aatatttttggtggctttgtagtaactcaacgcatgcttcgcatgtataaaaaataa 297 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 83363 aatatttttggtggctttgtagtaactcaacgcatgcttcgcatgtataaaaaataa 83307
Exemple 5: Other examples of alignment fo sequences
4546.3 (Seq ID 2365) 3009.2 (Seq ID 1331)
BLASTN 2.2.6 [Apr-09-2003]
Query= Lp Paris Contig48_66441-70516 (4076 letters)
Database: /local/http/htdocs/IPF_Gmp/legionella/blastdb/contig 51 sequences; 3,410,887 total letters
Searching . one Score E Sequences producing significant alignments: (bits) Value
>37 Length = 67519 Score = 1439 bits (726), Expect = 0.0 Identities = 784/802 (97%), Gaps = 1/802 (0%) Strand = Plus / Plus
Query: 3275 gcaaatatggtaaaaatgttgacattaattagtggagtatgagatatttattttgcgagg 3334 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 63920 gcaaatatggtaaaaatgttgacattaatcagtggagtatgagatatttattttgcgagg 63979
Query: 3335 ttggacttgctatgtgtcattccactgggtcaattggcgacacactgattggccctttct 3394 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 63980 ttggacttgctatgtgtcattccactgggtcaattggcgacacactgattggccctttct 64039
Query: 3395 attatcctgaaatcctgacaagagctctctatggcttaatctataagctgcttgtgatta 3454 I I I I I I I I III I I I I I I I II I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 64040 attatcctaaaaccctgacatgaactctctatggcttaatctataagctgcttgtgatta 64099
Query: 3455 atttcatcgcaatataagccattaaaataccgctaagtaactctattttttgccatactt 3514 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 64100 atttcatcgcaatataagccattaaaataccactaagtaactctattttttgccatactt 64159
Query: 3515 tattttgagttaacaggtttgaaaaataacgagtagtcatcgttaatgaactgaaccaaa 3574 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 64160 tattttgggttaataggtttgaaaaataacgagtagtcatcgttaatgaactgaaccaaa 64219
Query: 3575 tcatgctggcagaaattacccctgctaaaaaagccagtttatggtcaggatattgactgc 3634 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 64220 tcatgctggcagaaattacccctgctaaaaaagccagtttatggtcaggatattgactgc 64279
Query: 3635 tgccgctacccacaatcaccagggtgtctataatggcgtgaggattaagcagactaaacc 3694 I I I I I I I I I I I I II I I I I I I I I II I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 64280 tgccgctgcccacaatcaccagggtatctataatggcgtgaggattaagcagactaaacc 64339
Query: 3695 ccagggcaaataaaatgatttgcattcttgtatgaggctgatgtgtttctacaacggttt 3754 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 64340 ccagggcaaataaaatgatttgcattcttgtatgaggctgatgtgtttctacaacggttt 64399
Query: 3755 gtttgtttttggacagcgcgctttttaagttttttattgcataataaattaaaaaggcag 3814 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 64400 gtttgtttttggacaacgcgctttttaagttttttattgcataataaattaaaaaggcag 64459
Query: 3815 agcctagccataccatccagatttgcaagtttggatgagccaaaagcaattgatgtaaac 3874 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 64460 agcctagccataccatccagatttgcaagtttggatgagccaaaagcaattgatgtaatc 64519
Query: 3875 cagcgacactgccgcacaccaaaatgacatcacagaaaaagcaaatgacggcagagagag 3934 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 64520 cagcgacactgccgcacaccaaaatgacatcgcagaaaaagcaaatgacggcagagagag 64579
Query: 3935 cggcatgattttttcgcgcaccttgcctgataagaaagacattttgcggacctaaggcca 3994 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 64580 ccgcatgattttttcgcgcaccttgcctgataagaaaaacattttgcggacctaaggcca 64639
Query: 3995 ttatcaaagataatcccaagagtaatccattaaaataaatcaacataattgcatcaggat 4054 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 64640 ttatcaatgataatcccaagagtaatccattaaaataaatcaacataattgcatcaggac 64699
Query: 4055 agtaaaaaaaaaggcgattata 4076 I I I I I I I I I I I I I I I I I I I I I Sbjct: 64700 agt-aaaaaaaaggcgattata 64720
Score = 957 bits (483), Expect = 0.0 Identities = 495/499 (99%) Strand = Plus / Plus
Query: 1 ctacaaattttgcaaggttattaaatagtggttttcatctggcggcctattgtttttttg 60 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 63429 ctacaaattttgcaaggtaattaaatagtggttttcatctggcggcctattgtttttttg 63488
Query: 61 ggaaagccataagcattctgccaattgatccatgattaaatgttcaacagccatgggatc 120 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 63489 ggaaagccataagcattctgccaattgatccatgattaaatgttcaacagccatgggatc 63548
Query: 121 ctggtatttttctattaacttggtgtaaacagtacggatgccttgtggcctatcggtcgt 180 II I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 63549 ctgatatttttctattaacttggtgtaaacagtacggatgccttgtggcctatcggtcgc 63608
Query: 181 tacttgttcgcgaacggcaagatgaagccccatatgaaggaagggattggtttcgcctaa 240 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 63609 tacttgttcgcgaacggcaagatgaagccccatatgaaggaagggattggtttcgcctaa 63668
Query: 241 ttcaggataataagtatgttcaggaaaagattgaatttgttcaatgactttatggtattc 300 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 63669 ttcaggataataagtatgttcaggaaaagatggaatttgttcaatgactttatggtattc 63728
Query: 301 cggatgatcaagaatcacttgggcaatttctttttccaagggagaaagttcttttttatt 360 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 63729 cggatgatcaagaatcacttgggcaatttctttttccaagggagaaagttcttttttatt 63788
Query: 361 ctggtacttattccagctgataaaaaatagctgtcgagtttcttgtactgtatcgccgta 420 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 63789 ctggtacttattccagctgataaaaaatagctgtcgagtttcttgtactgtatcgccgta 63848
Query: 421 aaacataatggcccgattgatataaaatgatccattttaactgaataaaaaagtaacaac 480 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 63849 aaacataatggcccgattgatataaaatgatccattttaactgaataaaaaagtaacaac 63908
Query: 481 aatgttgatgtgcaaatat 499 I I I I I I I I I I I I I I I I I I I Sbjct: 63909 aatgttgatgtgcaaatat 63927
Database: /local/http/htdocs/IPF_Gmp/legionella/blastdb/contig Posted date: Sep 10, 2003 12:44 PM Number of letters in database: 3,410,887 Number of sequences in database: 51
Lambda K H 1.37 0.711 1.31
Gapped
Lambda K H 1.37 0.711 1.31
Matrix: blastn matrix:!
Alignment of a portion of contig48 (Seq Id 48) of the Paris strain with all the contigs of the
Philadelphia strain.
The positions of this fragment in the contig are indicated in the line starting with « Query= ». In this example, the first position (noted 1 in the alignment) of the fragment is thus position 50760 in contig48 of the Paris strain. This fragment terminates in position 56335 in contig48. 50760 should thus be added to the position indicated by the alignment to have the position in the total sequence of the contig. The positions in the contig of the Philadelphia strain are unchanged.
In this exemple, nous pouvons voir que the regions of the Paris strain between the positions
561(+50760) and 2096(+50760) and between 2622(+50760) and 3981(+50760) are absent from the Philadelphia strain. These regions thus contain the following ORFs, specific to the Paris strain:
322.3 (Seq ID 1466) 321.3 (Seq ID 1462) 3005.2 (Seq ID 1329) 5208.1 (Seq ID 2782)
BLASTN 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lip an (1997), "Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= Lp Paris Contig48_50760-56335 (5576 letters)
Database : /local/http/htdocs/IPF_Gmp/legionella/blastdb/contig 51 sequences; 3,410,887 total letters Searching . done Score E
Sequences producing significant alignments: (bits) Value 37 2892 0.0
Lp 1 1 i 1 1 1 8 2588 5000 37 "■"■ e ■ 8 25ΘΘ 5ΘΘΘ Lp Scoring colors S>θ S>5β S>188 S>15θ S>2ΘΘ
>37 Length = 67519 Score = 2892 bits (1459), Expect = 0.0 Identities = 1561/1595 (97%) Strand = Plus / Plus
Query: 3982 cttatggcaaatatttatccctcaaagcgtttctcaataagccattgttaccatgaacca 4041 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 50163 cttatggcaaatatttatccctcaaaatgtttctcaataagtcattgttaccttgaacca 50222
Query: 4042 gggtaagctttaattttcttaaacaaaatggagtatttagtcctcccttagatggaagct 4101 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 50223 gggtaagctttaattttcttaaacaaaatggagtatatagtcctctcttagatggaagct 50282
Query: 4102 ctttcactgctttgctgagcaaatatttgacgctcctcactgattaattcaatccatttt 4161 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 50283 ctttcactgctttgctgagcagatatttgacgctcctcactgattaattcaatccatttt 50342
Query: 4162 ttaggtgtcatgtcgttttgtatcaataagttgatcgctgcactaggatattgactttgt 4221 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 50343 ttaggtgtcatgtcgttttgtatcaataagttgatcgctgcactaggatattgactttgt 50402
Query: 4222 gtaatgacaaaataaaagtaagcggtcaatttgcttctgcataaagcccaaccacttttt 4281 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 50403 gtaatggcaaaataaaagtaagcggtcaatttgcttctgcataaagcccaaccacttttt 50462
Query: 4282 gcgacacattgttgcaaactagcaaattcaatttgatcttctttgactaattgttgcagc 4341 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I
Sbjct: 50463 gcgacacattgttgcaaactaggaaattcaatttgatcttctttgactaattgttgcagc 50522
Query: 4342 ttttgaatcgtgtcagggatacatgcatcaaaaaacaatgtggtcccttgctgtccccat 4401 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 50523 ttttgaatcgtgttagggatgcatgcatcaaaaaataatgtggtcccttgctgtccccat 50582
Query: 4402 acgtcacttaatataattctttttaaaatgttcatgtcacaaatttgttgctcatgggga 4461 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 50583 acgtcacttaatataattctttttaaaatgttcaagtcacaaatttgttgctcttgcgaa 50642
Query: 4462 ataggtttctcaattctatgtgttaccggttgccccaacatggaatgtgtccattgctga 4521 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 50643 ataggtttctcaattctatgtgttaccggttgccctaacatggaatgtgtccattgttga 50702
Query: 4522 taataagcccgaccgtaaattaatataccacccaaaataaaatcaaaggcagttggggct 4581 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 50703 taataagctcgaccgtaaattaatataccacccaaaataaaatcaaaggcagttggggct 50762
Query: 4582 gcataccaaagagcagtgaagaatttggatttgatgacgtccactttggggtagggatac 4641 I I I I I I I I I I I I I I I I I I I I I I I II I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 50763 gcataccaaagagcagtgaagaaattagatttgatgacgtccactttggggtagggatac 50822
Query: 4642 tgatggttattggttttaatgatatgtttgcgatcgtattgcatttttaaaggttcatat 4701 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 50823 tgatggttattggttttaatgatatgtttgcgatcgtattgcatttttaaaggttcatat 50882
Query: 4702 ttaaccaaggatttagcctggtattgattctggccaatacttatcgtttcttccttcttt 4761 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 50883 ttaaccagggatttagcttggtattgattctgaccaatacttatcgtttcttccttcttc 50942
Query: 4762 ggtattcctccatcgactattgctaaatgaaagttatgatttaaaacatgttcataaccc 4821 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 50943 ggtactcctccatcgactattgctaaatgaaagttatgatttaaaacatgttcataaccc 51002
Query: 4822 agaaaatgacaatcaggatttttagataaggtatttgctaaatgttgtcgcaatgcctcc 4881 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 51003 agaaaatgacaatcaggatttttagataaggtatttgctaaatgttgtcgcaatgcctcc 51062
Query: 4882 ccgactttaatattaggatcataaaaatctggatgtttggctaaattcattacccaaaat 4941 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 51063 ccgactttaatattaggatcataaaaatctggatgtttggctaaattcattacccaaaat 51122
Query: 4942 acattgactttattcttttcaccagcaggaatatctcgtaacatcattccccaacaatac 5001 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 51123 acattgactttattcttttcaccagcaggaatatctcgtaacatcattccccaacaatac 51182
Query: 5002 ccaataacttcgtcgtttttatccttggcaataaaacaaatagtctctttataatgtaac 5061 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 51183 ccaataacttcgtcgtttttatccttggcaataaaacaaatagtctctttataatgtaac 51242
Query: 5062 attgaagataggacgatatcccctataaaaggcaaatcgccaaaatttgctccatgtgct 5121 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I
Sbjct: 51243 attgaagataggacgatatcccctataaaaggcaaatcgccaaaatttgctccatgggct 51302
Query: 5122 aaagaatcgatacgtttaatttcacccattgctttattaagttcagcagtattattggga 5181 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 51303 aaagaatcgatacgtttaatttcccccattgctttattaagttcagcagtattattggga 51362
Query: 5182 tctaaaatctgaacttctttgaagccatagggcttgccttccatcatttttaacgaaaaa 5241 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 51363 tctaaaatctgaacttctttgaagccatagggcttaccttccatcatttttaacgaaaaa 51422
Query: 5242 ttaaaggtacattcgccatctaactttttgtggtgtttaattttaagaggattttgattt 5301 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 51423 ttaaaggtacattcgccatctaactttttgtggtgtttaattttaagaggattttgattt 51482
Query: 5302 ctgcaatttaacagatctgtccataattgatgaatgtaaagtaatctttcatcatttggt 5361 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 51483 ctgcaatttaacagatctgtccataattgatgaatgtaaagtaatttttcatcatttggt 51542
Query: 5362 gaggtatttcgcaaattgattaattgctgggtgaaacgatgcacatattctgtatggaga 5421 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 51543 gaggtatttcgcaaattgattaattgttgggtgaaacgatgcacatattctgtatggaga 51602
Query: 5422 taaatagcatggctactgaaatcaacaacaaactctgccgtcattaattttttgatttcg 5481 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 51603 taaatagcatggctactgaaatcaacaacaaactctgccgtcattaattttttgatttcg 51662
Query: 5482 ggtaagtgatattcaacaggatctgtatatgattttactattgcttctagtgaactccat 5541 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 51663 gataagtgatattcaacaggatctgtatatgattttactattgcctctaatgaactccat 51722
Query: 5542 ttgcgttcaagtacatcgatttttgaatacgacat 5576 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 51723 ttgcgttcaagtacatcgatttttgaatacgacat 51757
Score = 930 bits (469), Expect = 0.0 Identities = 543/569 (95%), Gaps = 9/569 (1%) Strand = Plus / Plus
Query: 1 atggcgaaatcattgtcgcaattagattctgctaatttgctcccctgttttaacatggaa 60 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 46540 atggcgaaaccattgtcgcaattggactctgctaatttgctcccctgttttaacatagag 46599
Query: 61 caagcagaacgcattggaaaacaaatcaataagctcttacagcatgagttttgcgaggaa 120 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 46600 caagcagaacgcattggaaaacaaatcaataaactcttacagcatgaattttgcgaggaa 46659
Query: 121 aacatcaatccaaagaaatttgcctctatcagtcacaatatcctgcccaaaattatgaca 180 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 46660 aacatcaatccaaagaaatttgcctctatcagtcacaatatcctgcccaaaattatgaca 46719
Query: 181 gaaacatttttaggagtaaccccgccagaaaactggcagcaattaagcgacgatattata 240 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I
Sbjct: 46720 gaaacatttttaggagtaacccctccagaaaactggcagcaattaagcgacgatattata 46779
Query: 241 aaaaactgcatcgcaaacaagaatctatgcaaaaaagcagctcgcaaagagctggaagaa 300 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 46780 aaaaactgcatcgcaaacaagaatctatgcaaaaaagcagctcgcaaagagttggaagaa 46839
Query: 301 tgcatcaaaccgagaattcctttgatcctgatacaatttggcccctggcttgctcaaaat 360 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 46840 tgcatcaaaccgagaattcctttgatcctgatacaatttggcccctggcttgcccaaaat 46899
Query: 361 tgtccacaattgaataagtctttgattgaacaatggccaaataaacaggctactctcaaa 420 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I MINIMI Sbjct: 46900 tgtccacaattgaataagtctttgattgaacaatggccaaataaacaggccactctcaaa 46959
Query: 421 aagataattaatgaaaacaaaagtgccgagtaatcgagacaggcttaattacgagagtta 480 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 46960 aagataattaatgaaaacaaaagtgctgagtaatcgaggcaggcttaattacgagagtta 47019
Query: 481 tgcctgataaaaccacatttatttacctaat ttcatcaaatataactcacc 531 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I III Sbjct: 47020 tgccggataaaacaacatttatttacctaattcacctaacttcatcaaatataacttacc 47079
Query: 532 gatatgatctacaaccaagttcttaaaac 560 I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 47080 gatatgatctacaaccaggttcttaaaac 47108
Score = 656 bits (331), Expect = 0.0 Identities = 423/451 (93%), Gaps = 2/451 (0%) Strand = Plus / Plus
Query: 2172 tatatggccgataaaatttgccagggtcaataaatagtattctgatggttaggtaataaa 2231 I I I I II I I I I I I I I I I I I I I I I I I I I II I II I I I I I I I I I I I I I I I III I I I I I I I Sbjct: 47927 tatatgaccgataaaatttgccagggtcagtaaatagtattctgatggctagataataaa 47986
Query: 2232 tgatgaagagttcttttggtgaaatgaataaaagaccccttttttattgagcgactctta 2291 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I II I I I I I I I I I I I I I I I I I I I I I Sbjct: 47987 tgatgaaaagttcttttgatgaaatgaataaaagaacc-ttttttattgagcgactctta 48045
Query: 2292 aaaa-gccatttgctttattctgtgcttttggcaagtgacatgatcgcattcaggttttg 2350 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I III I I I I I I I I I I I I I I I Sbjct: 48046 aaaaagccatttgctttatcctgtgcttttggcaagcagtatggtcgcattcaggctttg 48105
Query: 2351 tttacatgctaggtgtggttttttcccgaggcggttgtgagtttgataccataggtttta 2410 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 48106 tttacatgctaggtgtagttttttcctgaggcggttgtgggcttgataccataggtttta 48165
Query: 2411 ttgaataggcgccgcttgggaaaaaacaattgtcagtaaggattgcccctgtaatcagat 2470 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 48166 ttgaatagacgccacttgggaaaaaacaattgtcggcaaggattgcccctgtaatcagct 48225
Query: 2471 gggtaagccgattggcttcggcacatctcgaatcagtgacaacctctttcgctttttcat 2530 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I II I I I I I I I I I I I I I II I I I II I I I I I I I I
Sbjct: 48226 gggtaagccgattggcttcggcacatctcgaatcagtggcaacctctttcgctttttcat 48285
Query: 2531 ctttatttcgcataacaatcctgtgaagttaatctttgcagaggacaccatgatggtttc 2590 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 48286 ctttatttcgcataacaatcctgtgaagttaatctttgcagaggacactatgatggtttc 48345
Query: 2591 atgtcatacaaacgaagcaaccggataccga 2621 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 48346 atgtcatacaaacgaagcaaccgggtaccga 48376
Score = 147 bits (74), Expect = 9e-35 Identities = 98/106 (92%) Strand = Plus / Plus
Query: 2097 acctttctggagtttcccaactacaagatgatactgcgttataataactccatttattat 2156 I I I I I I I I I I I II I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 44922 accttcctggcgtttcccaactacaagatgatcctacgttataataactccatttattat 44981
Query: 2157 actggggctatcgagtatatggccgataaaatttgccagggtcaat 2202 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 44982 gccggggctatcgggtatatggctgataaaatttgccagggtcaat 45027
Score = 109 bits (55), Expect = 2e-23 Identities = 64/67 (95%) Strand = Plus / Plus
Query: 3653 cgaatccttacaggaaaacgaagcttatggaagtccaacaaggaagaggtagtaaattca 3712 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 48378 cgaatccttacaggaaaacgaagcttatggaaggccaacaaggaacagggagtaaattca 48437
Query: 3713 taacgcc 3719 I I I I I I I Sbjct: 48438 taacgcc 48444
Database : /local/http/htdocs/IPF_Gmp/legionella/blastdb/contig Posted date: Sep 10, 2003 12:44 PM Number of letters in database: 3,410,887 Number of sequences in database: 51
Lambda K H 1.37 0.711 1.31
Gapped
Lambda K H 1.37 0.711 1.31
Matrix: blastn matrix:! -3
Alignment of a portion of the contig45 (Seq Id 45) of the Paris strain with all the contigs of the Philadelphia strain.
The positions of this fragment in the contig are indicated in the line beginning with « Query= ». In this example, the first position (noted 1 in the alignment) of the fragment is thus position 44722 in the contig45 of the Paris strain. This fragment terminates at position 52680 in the contig45. It is thus required to add 44722 a to the indicated by the alignment to have the position in the total sequence of the contig. The positions in the contig of the Philadelphia strain unchanged.
In this example, we can see that the region of the Paris strain between the positions 1333(+44722) and 6899(+44722) is absent from the Philadelphia strain. This region thus contains the following ORFs, specific to the Paris strain:
4927.1 (Seq ID 2623) 413.5 (Seq ID 2069)
415.2 (Seq ID 2087)
417.3 (Seq ID 2102)
BLASTN 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of protein database search programs", Nucleic Acids Res. 25:3389-3402.
Query= Lp Paris Contig45_44722-52680 (7959 letters)
Database: /local/http/htdocs/IPF_Gmp/legionella/blastdb/contig 51 sequences; 3,410,887 total letters
Searching. done Score E
Sequences producing significant alignments: (bits) Value
48 1943 0.0 49 94 2e-l£
Lp 1 1 1 1 1 1 1 1 1 1 1 θ 2500 5000 75ΘΘ 48 49 θ 258Θ 5080 750Θ L i 1 1 1 1 Lp 1 1 1 1 1 1 1 1 1 Scoring colors S>8 S>58 S>lθθ S>158 S>2βθ
>48 Length = 263853
Score = 1943 bits (980), Expect = 0.0 Identities = 1040/1060 (98%) Strand = Plus / Minus
Query: 6900 attctaatggtggccagaggcagaatcgaactgccgacacgaggattttcagtcctctgc 6959 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 55098 attctaatggtggccagaggcagaatcgaactgccgacacgaggattttcagtcctctgc 55039
Query: 6960 tctaccgactgagctatctggccactcaaggcttgctattaaactccgagatcgactttg 7019 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 55038 tctaccgactgagctatctggccactcaaggcttgctattaaactccgagatcgactttg 54979
Query: 7020 agtcaagtgttgattaagttttttttgaaaataattttttttggtaggctatggcaatct 7079 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 54978 agtcaagtgttgattaagttttttttgaaaataattttttttggtaggctatggcaattt 54919
Query: 7080 ctgctcctagaaaagcgttgtttttgataagctcaatattagcttccaggctcttgcctg 7139 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 54918 ctgctcctagaaaagcgttgtttttgataagctcaatattagcttccaggctcttgcccg 54859
Query: 7140 cagttaattcagcaatacgttttaacaggaagggggttaatgattttccgctcatgtgtt 7199 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 54858 cagttaattcagcaatacgttttaacaggaagggggttaatgattttccgctcatgtgtt 54799
Query: 7200 tggcttcttcatgagcctgttttatgtatggactgatttcctcatcagatagttccgctg 7259 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 54798 tggcttcatcatgagcctgttttatgtatggactgatttcctcatcggatagttcggctg 54739
Query: 7260 atactggaatggggtttgcgacgactattccgtttttcatgttcaatttctgttgaattg 7319 I I I I I I I I M I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 54738 atactggaatggggtttgcgacgactattccatttttcatattcaatttctgttgaattg 54679
Query: 7320 acatgagatttgctacttcctcgaccgaatttaagcgttgtgggactggtattccactcg 7379 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 54678 acatgagatttgctacttcctcggccgagtttaagcgttgtggaactggtattccactcg 54619
Query: 7380 atctgctgtaaaaagcagggaattcgtctgtggcataacctatgaccggcaccccaaacg 7439 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 54618 atctgctataaaaagcagggaattcgtctgtggcataacctatgaccggcaccccaaacg 54559
Query: 7440 tttcaagaacttccaatgtttttggtaagtcgagaatcgattttgcgccagaacagacta 7499 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 54558 tttcaagaacttccaatgtttttggtaagtcgagaatcgattttgcgccagaacagacta 54499
Query: 7500 cggtaactggcgtattggatagttctataagatcagctgaaatatcaaaactcattgtca 7559 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 54498 cggtaactggcgtattggatagctctataagatcagctgaaatatcaaaactcatcgtca 54439
Query: 7560 cgtcttgatgaacaccacctatacctccggtgacaaatagggggagcctagccatatggg 7619 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 54438 cgtcttgatgaacaccacctatacctccggtgacaaatagggggagcttagccatatggg 54379
Query: 7620 cacagaacatggttgctgctaccgttgtgctggcagtcactttacgagataatacaaaag 7679 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 54378 cacaaaacatggttgctgctaccgtcgtgctggcagtcactttacgagataatacaaaag 54319
Query: 7680 aaatgtctctgcgagaggcttttattacttctttttgcagcgcgagatgctccataactt 7739 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 54318 aaatgtctctgcgagaggcttttattacttctttttgaagcgcgagatgttccataactt 54259
Query: 7740 cttgagttaaaccaatacggattttcccttggtgcatcgctatagtagctggaatggcgc 7799 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 54258 cttgagttaaaccaacacggattttcccttggtgcatcgctatagtagctggaatggcgc 54199
Query: 7800 cttgtctacgaataatattttcaacttctattgccgtagttaaattatcagggtagggca 7859 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 54198 cttgtctacgaataatattttcaacctctattgccgtagttaaattatcagggtagggca 54139
Query: 7860 ttccatgagagataatggtcgactcaagagcaacaattgggtttttatcattgatagcat 7919 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 54138 ttccatgagagataatggtcgactcaagagcaacaattgggtttttatcattgatagcat 54079
Query: 7920 ccagtacttcttcgttaaattccaacaagtcatgaaacat 7959 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 54078 ccagtacttcttcgttaaattccaacaagtcatgaaacat 54039
>49 Length = 376826 Score = 93.7 bits (47), Expect = 2e-18 Identities = 102/119 (85%), Gaps = 1/119 (0%) Strand = Plus / Plus
Query: 1215 atttgccctgtgtattgtttagtgttggtcgagcggttcactctctgttgaaacccggta 1274 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I II Sbjct: 207683 atttgccctgtctttggtttagtgttgaacgagcggctcactcaaagatgaaacctggcc 207742
Query: 1275 aaa-ccgtaaagctcgaagaagggggcaaatcaatcgttataaggcaaacgatcccgcc 1332 III II I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 207743 aaagccataaagctcgaagtagggggcaaatcaatcgttataaggcaaacgatctcgcc 207801
Database: /local/http/htdocs/IPF_Gmp/legionella/blastdb/contig Posted date: Sep 10, 2003 12:44 PM Number of letters in database: 3,410,887 Number of sequences in database: 51
Lambda K H 1.37 0.711 1.31
Gapped
Lambda K H 1.37 0.711 1.31
Matrix: blastn matrix:! -3
Alignment of a portion of contig39 (Seq Id 39) of the Paris strain with all the contigs of the
Philadelphia strain.
The positions of this fragment in the contig are indicated in the line starting with « Query= ». In this example, the first position (noted 1 in the alignment) of the fragment is thus position 3990 in contig39 of the Paris strain. This fragment terminates at position 8972 in contig39. 3990 should this be added to the position indicated by the alignment to have the position in the total sequence of the contig. The positions in the contig of the Philadelphia strain are unchanged.
In this example, we can see that the region of the Paris strain between the positions 1264(+3990) and 4465(+3990) is absent from the Philadelphia strain. This region thus contains the following
ORFs, specific to the Paris strain:
3396.1 (Seq ID 1588)
3395.2 (Seq ID 1587) 3394.1 (Seq ID 1586)
BLASTN 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of protein database search programs", Nucleic Acids Res. 25:3389-3402.
Query= Lp Paris Contig39_3990-8972 (4983 letters)
Database : /local/httρ/htdocs/IPF_Gmp/legionella/blastdb/contig 51 sequences; 3,410,887 total letters
Searching. done Score E
Sequences producing significant alignments: (bits) Value
38 2123 0. .0 49 638 0. ,0
Lp M I M I I M M M I M I I M M M M M M M I I M M I II M M M 8 588 lθθθ 1500 2Θ00 2588 38ΘΘ 35ΘΘ 4ΘΘΘ 45ΘΘ 38 49 θ 588 lθβθ 1588 2ΘΘΘ 2588 38ΘΘ 3588 4ΘΘΘ 4588 I I I I 1 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Lp Scoring ■■■■■■■■■■■■■■H colors S>θ S>5Θ S>lθθ S>15β S>2ΘΘ
>38 Length = 17003
Score = 2123 bits (1071), Expect = 0.0 Identities = 1215/1263 (96%) Strand = Plus / Minus
Query: 1 ctttggttctacatgagcttgcctgagatgtttgcccttatcgattgcaacaatttttac 60 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 12277 ctttggttctacatgagcttgcctgagatgtttgcccttatcgattgcaacaatttttac 12218
Query: 61 gccagttgtgagcgtttgtttcgtcctgatttaaaggacgtccccatcgtggtgctatcc 120 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 12217 gccagttgtgagcgtttgtttcgtcctgatttaaaggacgtccccatcgtggtgctatcc 12158
Query: 121 aataatgatggctgttgtatcgcacgctcgaatgaagccaaagcattgggcattgccatg 180 I I I I I I I I I I I I I I I I I I I I I I I I I I II I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 12157 aataacgatggctgttgtatcgcacgctcgaatgaagccaaagcattgggcattgccatg 12098
Query: 181 ggcgagccgtacttcaaaattaaacatttgtgcaaacagcatggagtgaaagctttttcc 240 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I II Sbjct: 12097 ggcgagccgtacttcaaaattaaacatttgtgcaaacagcatggagtgaaagctttttcc 12038
Query: 241 tcaaattatacgctgtatggcaacatgagtcatcgtgtgatgtgcactattgaagaagcc 300 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 12037 tcaaattatacgctgtatggcaacatgagtcatcgtgtgatgtgcactattgaagaagcc 11978
Query: 301 tggccccatatagaagtttactcgattgatgaagcgtttcttgatttaaggagtttaccg 360 I I I I I I I I I I I I I I I I I I I I I I I I I I II I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 11977 tggccccatatagaagtttactcgattgatgaagcgtttcttgatttaaggagtttaccg 11918
Query: 361 gttgatagccatgattcgttttgcgagcagttacaaaagaaaatcttgaagcacacagga 420 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 11917 gttgatagctatgattcgttttgcgagcagttacaaaagaaaatcttgaagcacacagga 11858
Query: 421 atacccacttccatcggtattggacctactaaaacactagctaaagccgccaatcattta 480 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 11857 atacccacttccatcggtattggacctactaaaacactagctaaagccgccaatcattta 11798
Query: 481 tgcaaaaaagtttataaaatccctgtgtttaatatcacctcgaatcgtgggcggttattg 540 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 11797 tgcaaaaaagtttataaaattcctgtgtttaatatcacctcgaatcgtgggcggttattg 11738
Query: 541 caacagatttccgttggggacatttggggagtagggcggcaatgggccaataaattaatt 600 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 11737 caacagatttccgttggggacatttggggagtagggcggcaatgggccaataaattaatt 11678
Query: 601 tcgcgaggcattcatacggcttatgatttggcaatgaccaatcctcaccttctgaagaaa 660 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 11677 tcgcgaggcattcatacggcttatgatttggcaatgaccaatcctcaccttctgaagaaa 11618
Query: 661 tgttttaacgtcgtgttgatgcgtaccgccatggagcttcaaggaattgcttgtggcggt 720 I I I I I I I I I I I I II I I I I I I I I II II I I I I I I I I I I I I II I I I I I I I I I I I I I I II Sbjct: 11617 tgttttaacgccgtgttgatgcgaactgctatggagcttcaaggaattgcttgtggcggt 11558
Query: 721 ttagaggcaatagagcctaagcaaagtattatgtcatctaaaagttttggtcagatgcaa 780 I I I I I I I II I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 11557 ttagaggtaatagagcctaagcaaagtattatgtcatccaaaagttttggtcagatgcaa 11498
Query: 781 actcaacttgcttcgattgaggaatcaatcagtagccattgtgcccgtgcggtggagaaa 840 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 11497 actcaaattgcttcgattgaggaatcaatcagtagccattgtgctcgtgcggtggagaaa 11438
Query: 841 atgcgtcgccagcaattggtggcgaagcgcctggttgtatttgtgcatacgaaccgattt 900 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 11437 atgcgtcgccaacaattggtggcgacgcgtctggttgtatttgtgcatacgaaccgattt 11378
Query: 901 cgcgaagatttggcacagcactttcagtccatcgaatttaagctgattaatcctacagat 960 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 11377 cgggaagatttggctcagcactttcagtccatcgaatttaagctgattaatcctacagat 11318
Query: 961 gatttgcgcttaattaccaaaatggccaagcgatgtctgcaacgcatttttaaaccaggg 1020 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 11317 gatttgcgcttaattaccaaaatagccaaaagatgtctgcaacgcatttttaaaccaggg 11258
Query: 1021 tattactataaaaaggcaggagtatgtcttgaggacttaattcccaaaaacccacgacag 1080 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 11257 tattactataaaaaggcaggggtatgtcttgaagacttaattcctaaaaaaccacgacag 11198
Query: 1081 ctggatatgttttatcaaccaagtgacgagcatctaaaccacacggaacaattgatggcg 1140 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 11197 ctggatatgtttcatcaaccaagtgatgagcatctaaaacacaccgaacaattgatgggt 11138
Query: 1141 gtctttgaccaaatcaatcaaaaatacggacgaagtacaatccgcctcgcggcagagggt 1200 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 11137 gtctttgaccaaatcaatcaaaagtatggaagaagcacgattcggttagccgccgaaggc 11078
Query: 1201 tattcaaaaccttgggcgatgcgtgctgaactgaaatcgcctgcctataccacacgatgg 1260 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 11077 tattcaaaaccctgggagatgcgtgctgagctgaaatcacctgcttataccacgcgatgg 11018
Query: 1261 tct 1263 I I I Sbjct: 11017 tct 11015
>49 Length = 376826 Score = 638 bits (322), Expect = 0.0 Identities = 469/518 (90%) Strand = Plus / Plus
Query: 4466 gaacaataatcactgataaaaagatcttgagcaaaagtctcaaaatcaaaatagcagatc 4525 I I I I I I I I' I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 203775 gaacaataatcactaataaaaagatcccgagcaaaagcctcaaaatcaaaatagcagctc 203834
Query: 4526 aaactatccggcatttgatgggcataacagtcatcaaataattgcctggcaaaatcaacc 4585 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I
Sbjct: 203835 aaactatccggtatttgatgggcataacagtcatcaaataactgtctggcaaaatcaacc 203894
Query: 4586 tctgaatcataacaaccttggtaatgatcctccaacatggtttgtgcatcatctaccgag 4645 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 203895 tctgaatcatagcaaccttggtaatgatcctccaacatggtttgtgcatcttctaccgaa 203954
Query: 4646 taatcacagagcagggctaatcctaactctccgtgttcttgaatgaatgaagcatattcc 4705 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 203955 taatcgcagagaagggctaagcctagctctccatgttcttgaataaatgaagcatactcc 204014
Query: 4706 acaatgttgcttattccttcatattcatgaattctcatgctgccgaaaccttcataatcg 4765 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 204015 acaatattacttattgactcgtactcatggattttgatgctgccgaatccttcataatcg 204074
Query: 4766 tggatggcaaattcctcagcattggtttctgggctattatccaacatttcccagatttct 4825 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 204075 tgtatggcatattcttcagcattgggctcagggctgttatccagcatttcccagatttct 204134
Query: 4826 ttcatgatgtcatcttcgctttgagtagcatctatccatacaccatgcagtatggcgttg 4885 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 204135 tccataatgtcatcttcgctttgtgtggcatctatccaaacaccatgcaggatggcattg 204194
Query: 4886 ttgtaagaggctaaacaggcgacgtagattgaaggggtgtccatgggattatctccttgt 4945 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 204195 ttgtatgaggctaaacacgcgacgtagattgaaggggtgtccatgggattatttccttgt 204254
Query: 4946 attaagggagctatcccacacgggagcttgctcccgtg 4983 I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I I Sbjct: 204255 attaagggagcaatcccacacgggagcttactcccgtg 204292
Database: /local/http/htdocs/IPF_Gmp/legionella/blastdb/contig Posted date: Sep 10, 2003 12:44 PM Number of letters in database: 3,410,887 Number of sequences in database: 51
Lambda K H 1.37 0.711 1.31
Gapped
Lambda K H 1.37 0.711 1.31
Matrix: blastn matrix:!
Example 6: RESULTS See hereinbelow Tables V, VI, I and II hereinbelow.
Example 7 : Sequencing and annotation of the genome of L. pneumophila Lens strain Comparison of the sequences of the genomes of L. pneumophila Paris strain, Lens strain and Philadelphia strain
(http://genome3.cpmc.columbia.edu/~legion/index.html). three strains of serogroup 1, shows that around 88% of these genomes are very strongly preserved (95 to 100% of proteic identity), whereas the remaining 12% are specific to each strain. These results suggest that there is a large genomic diversity at the very centre of the L. pneumophila species. The Table XVI hereinbelow comprises for each of the ORFs identified in the
Lens strain its position on the genome, the existence of a peptide signal, the best result of the blast on mprot (Best-Blastp). The ORFs specific to the strain L. pneumophila
Lens relative to the strain L. pneumophila Paris were identified in considering as specific the ORFs having a percentage of proteic homology less than 75%. In the case where the ORF is preserved in the two genomes, the percentage of homology between the two proteins is mentioned. Finally, the ORFs specific to the Legionella geme were identified in considering as specific the ORFs having a percentage of proteic homology with sequences of the bank mprot less than 25 %. In conclusion, these results help define DNA probes for developing a typing tool. The utilization of this tool on a large number of strains isolated from patients and strains isolated from the environment can confirm if this tool can predict the risk associated with a strain by definitely discriminating the strains isolated from patients of the other strains.
Material supplied:
• The complete sequence of the genome of Legionella pneumophila Lens strain made up of the long chromosome of around 3.33 Mb and of a long plasmid of around 60 kb. • A list of specific coding phases of L. pneumophila Lens strain annotated with their nucleotidic sequences.
Materials and methods.
1. Construction of the shotgun bank of small fragments (size 1.5 to 2.5 kb) The chromosomal DNA of the strains studied was prepared by a classic method including proteinase K treatment and phenol extraction (9). Around 40 ug of DNA were broken by nebulization (1 minute under pressure of 1 bar) (4). The ends of the fragments of DNA were rendered free by having the DNA-polymerase of the bacteriophage T4 act for 15 minutes at 37°C in the presence of the 4 tri-phosphate nucleotides. The enzyme was inactivated by incubation of 15 mn at 75°C. Adaptors (invitrogen Cat. N° 408-18) have then been ligatured to these ends. After ligature, the fragments of chromosomal DNA having a size between 1500 and 2500 base pairs were purified after electrophoresis on agarose gel. The vector utilized for construction of the bank, pcDNA2.1 (Invitrogen), was digested by the enzyme BstXl and purified by geneclean (BIO-101) after electrophoresis on agarose gel. The chromosomal DNA and the purified vector were ligatured by action of the ligase of the bacteriophage T4. The ligation mixture was introduced by transformation in the strain of Escherichia coli XL2- blue (Stratagene). About 4000 colonies are obtained per ul of the ligation mixture.
2. Preparation of plasmids and sequencing The plasmids were prepared from bacterial colonies with the «TempliPhi DNA sequencing template amplification» kit marketed by Amersham Bioscience. The chromosomal inserts were sequenced from their two ends by utilizing the universal primer T7 by following the recommendations of the supplier (Applied-Biosystems). The sequences were determined by utilizing automatic sequencers of type 3700 (Applied- Biosystem).
3. Assembling of sequences The sequences were assembled by utilizing the set of software developed at the
University of Washington, Phred, Phrap and Consed (5, 8). The sequence was finished by utilizing the set of CAAT-box software (7). The finishing stage corresponds to resequencing the regions where the sequence is only slightly secure and sequencing of the regions situated between the contigs. It was done either by sequencing PCR products or by operating on the clones of the bank. The oligonucleotidic sequences were defined by utilizing consed and Pri o software (8, 10). 3. Annotation of sequences The identification of the coding phases (CDS) was completed by utilizing the set of CAAT-box software (7). This program combines the results of different methods: (i)
identification of open reading phases and their tri as a function of their size, (ii) analysis of the probability of being coded by utilizing Genemark software (11), (iii) identification of a start in translation (initiation codon and fixing sequence of the ribosome), (iv) similarity of the proteic sequence deduced with the proteic sequences contained in the sequence banks by utilizing BLASTP software. The functions of the proteins coded by the coding phases identified were predicted by analysis of the research results of similarities in the non-redundant bank of the NCBI (http://www.ncbi.nlm.nih.gov/BLAST/) by utilizing BLASTP software (1). 4. Comparison of the genomes - identification of the CDS specific to the strain of L. pneumophila Paris strain The set of proteic sequences deduced from the predicted coding phases each genome was compared to the set of proteic sequences possibly coded by the other genome by utilizing BLASTP software. A threshold of 75% of identity on the totality of the length of the protein was retained for identifying the proteins specific to an isolate. This very high value was kept since it best allows the orthologous genes of the paralogous genes (6) to be discriminated. For the proteic sequences for which the sequence preservation is high (> at 70%) the preservation of the nucleotidic sequences of the genes will also be high and could give a signal in hybridization conditions of low stringency. It will be necessary to take into consideration this eventuality in the analysis of the test result. 5. Examples of annotations 5.1. Genes specific to L. pneumophila Lens strain. There is no significant similarity between the nucleotidic sequence of the gene of L. pneumophila Lens strain and the genome of L. pneumophila Paris strain.
5.2. Genes common to the two strains for which the similarity (identity) of the deduced proteic sequences is less than 75% and value of the similarity at the nucleotide level.
5.4. Genes common to L. pneumophila Lens strain and Paris strain.
1. Altschul, S. F., T. L. Madden, A. A. Schaffer, J. Zhang, Z. Zhang, W. Miller, and D. J. Lipman. 1997. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. [Review] [90 refs]. Nucleic Acids Research. 25:3389-402. 2. Aurell, H., J. Etienne, F. Forey, M. Reyrolle, P. Girardo, P. Farge, B. Decludt, C. Campese, F. Vandenesch and S. Jarraud. 2003. Legionella pneumophila serogroup 1 strain Paris: Endemic distribution throughout France. J Clin Microbiol. 41 : 3320-3322 3. Birnboim, H. C. 1983. A rapid alkaline extraction method for the isolation of plasmid DNA. Methods Enzymol. 100:243-255. 4. Buchrieser, C, C. Rusniok, L. Frangeul, E. Couve, A. Billault, F. Kunst, E. Carniel, and P. Glaser. 1999. The 102 kb locus of Yersinia pestis : sequence analysis and comparison of selected regions among different Yersinia pestis and Yersinia pseudotuberculosis strains. Infect. Immun. 67:4851-4861. 5. Ewing, B., and P. Green. 1998. Base-calling of automated sequencer traces using phred. II. Error probabilities. Genome Res. 8:186-194. 6. Fitch, W. S. 1970. Distingishing homologous from analogous proteins. Syst. Zool 19:99-113.
7. Frangeul L, Glaser P, Rusniok C, Buchrieser C, Duchaud E, Dehoux P and Kunst F. 2004. CAAT-Box, Contigs-Assembly and Annotation tool-box for genome sequencing projects. Bioinformatics. in press.
8. Gordon, D., C. Abajian, and P. Green. 1998. Consed: a graphical tool for sequence finishing. Genome Res. 8:195-202.
10. Li, P., K. C. Kupfer, C. J. Davies, D. Burbee, G. A. Evans, and H. R. Garner. 1997. PRIMO: A primer design program that applies base quality statistics for automated large-scale DNA sequencing. Genomics. 40:476-485.
11. Lukashin, A. V., and M. Borodovsky. 1998. GeneMark.hmm: new solutions for gene finding. Nucleic A cids Res. 15:1107-1115.
12. Xu, S.Y., and Fomenkov, A. 1994. Construction of pSClOl derivatives with Camr and Tetr for selection or LacZ for blue/white screening. Biotechniques 17:57.
Example 8: Proof in the genome of Legionella pneumophila of the exploitation of functions of the host cell and of the high genomic plasticity Legionella pneumophila, the causal agent of legionnaires disease, replicates in the form of an intracellular parasite of the amoeba and persists in the environment in the form of a free living microbe. Analyzed here are the complete genomic sequences of L. pneumophila Paris (3 503 610 bp; 3 077 genes), an endemic strain predominant in France, and L. pneumophila Lens (3 345 687 bp; 2932 genes), an epidemic strain responsible for a major epidemic in France. A striking characteristic of the genome of L. pneumophila is its plasticity. Three different plasmids were identified, and -13 % of each genome is different to the other strain. The Paris strain codes for a unique secretion system of type V, and its secretion system Lvh of type IV is coded by a region of 36 kb which can be either carried on a multicopy plasmid or be integrated into the chromosome. The genetic mobility can be a mechanism which increases the multiplicity of L. pneumophila. A large number of genes codes for proteins or patterns of eukaryotic type provided to modulate the functions of the host cell to the advantage of the pathogen, comprising the repeated sequences of tetratrico peptide, ankyrin, F box, serin- threonin kinase proteins, apyrases and a sphingosine-1 -phosphate lyase. Therefore, the genome reflects the history and the lifestyle of L. pneumophila, a human pathogen of the macrophages which has co-evolved with soft-water amoeba. L. pneumophila is the causal agent of the legionellosis, an atypical pneumonia, which can be fatal if it is not rapidly treated1. This Gram-negative facultative
5/049642 89
intracellular pathogen can at the same time adapt to the aquatic environment and to the intracellular medium of the phagocytary cells of the human host2. When inhaled in contaminated aerosols, L. pneumophila can reach the alveolae of the lungs where they swamped by macrophages. By opposition to the majority of bacteria, which are destroyed, L. pneumophila can multiply within the phagosome of the macrophage and in the end kill off the macrophage, resulting in legionellosis3. L. pneumophila and other legionelles are inhabitants of natural aquatic biotopes and artificial aqueous systems, such as the cooling towers of air conditioners3. Legionelles were detected by culture in soft-water environments at 40 % and by PCR in soft-water environments up to 80 %, where the bacteria are known to survive and replicate by intracellular means in free living protozoons, often within aquatic biological films4. Its capacity for exploiting the base cellular mechanisms of a large spectrum of protozoic eukaryotic hosts likewise allows legionelles to infect human cells5. In fact, it was shown that the capacity of L. pneumophila to multiply by intracellular means in amoeba contributes to the disease, even though little is known of things about the mechanisms governing the host-microbe interactions. Inversely, the biphasic life cycle of L. pneumophila, the changing of replicating parasitic cells into transmissible extracellular forms and the complex regulatory network, which governs these changes, are understood in part6. The Legionella genre comprises 48 species, but more than 90 % of the cases of clinical legionellosis are caused by L. pneumophila even more arresting, up to 84 % are caused by the serogroup 1 of L. pneumophila1. We have determined the complete genomic sequences of two clinical isolates of the serogroup 1, the Paris and Lens strains, to provide knowledge of the genetic characteristics of L. pneumophila, and to identify the properties which were selected in niches specific to the pathogenicity and the life cycle of L. pneumophila. The Paris strain is the only endemic strain known to date, accounting for 12.7 % of the cases of legionellosis in France and for 33 % of those occurring in the Paris region . It is associated with the nosocomial and community diseases occurring in the form of epidemics or sporadic cases. From November 2003 to January 2004 the Lens strain caused an epidemic of 86 cases resulting in 17 deaths in the north of France, suggesting that it is particularly efficacious for causing the disease in humans. The genomic comparatives of an endemic isolate and of an epidemic isolate supplying the bases for comprehending the specificity of strain, and can give indices for the particular adaptability and stability of the Paris strain.
Results
General characteristics The Paris strain and the Lens strain of L. pneumophila each contain a circular chromosome of ~3 503 610 bp and ~3 345 687 bp, respectively, with an average G+C content of 38 % (Table XXII, Figure 1, Gen-Bank/EMBL access numbers CR628336, CR628337). L. pneumophila Paris separates a plasmid of 131 885 bp and the Lens strain contains another plasmid of 59 832 bp (Gen-Bank/EMBL access numbers CR628338, CR628339. In the chromosome of J. pneumophila Paris, 3077 genes were identified and 2 932 in that of J. pneumophila Lens. No function was able to be predicted for 42.1 % (1354) of the genes of L. pneumophila Paris and 44.1 % (1320) of the genes of L. pneumophila Lens, a proportion similar to that found in the majority of the other bacterial genomic sequences. A high proportion of the genes provided (21 % for the Paris strain, 20.4 % for the Lens strain) is unique to the Legionella geme and can thus code the specific functions of Legionella.
Exploitation and modulation of the functions of the host cell A fascinating question is to know how Legionella to decompose the functions of the host to enter, survive, replicate and evade amoeba or alveolar macrophages. Within its genome L. pneumophila codes for an abundance of proteins of eukaryotic type. In effect, 30 have the highest similarity with eukaryotic proteins (Table XXIII) and 32 genes code for proteins with eukaryotic domains implied in protein-protein interactions (Table XXIV). We reveal here proteins provided for diverting eukaryotic regulatory paths or for being secreted in eukaryotic cells, making strong candidates of them for directing the invasion of Legionella, for traveling in the host cell, or modulating or being subtracted from functions of the host cell. The repeated sequences of tetratrico peptide (TPR) are repeated patterns of the 34 degenerated amino-acids present in networks in tandem of 3 to 16 patterns, which form hooks for facilitating the protein-protein interactions. The TPR proteins contribute to control of the cellular cycle, to repression of transcription, to response to stress, to inhibition of the protein kinase, to the transport of mitochondrial and peroxisomal protein and to neurogenesis9. The Sel-1 repeated sequences represent a sub-family of TPR sequences. In L. pneumophila five proteins containing Sel-1 domains were identified. Two of them (EnhC and LidL) were previously implied in interaction with the host cells or in precocious signaling of events which regulate the scheduling
decisions of £. pneumophila in the macrophages10'11. As a consequence, the three newly identified proteins are likewise in al likelihood implied in the host-pathogen interactions (Table XXIV). After internalizing, L. pneumophila manipulates the endosome-lysosome degradation path of the host for surviving and replicating within a vacuole derived from the endoplasmic reticulum (ER). A protein of L. pneumophila, RalF, thought to contribute to recruitment of the ER contains a eukaryotic domain Sec 7. RalF, a substrate of the secretion system of type IN is required by the regulatory protein ARF for associating with the phagosomes of L. pneumophila12. The two strains of L. pneumophila code for three serin/threonin kinase proteins of eukaryotic type (STPK) (Table XXIN). The multisequence comparisons of the domains of the kinase from the Paris strain of L. pneumophila and other prokaryotic and eukaryotic STPK, have revealed that Lpp2626 and Lppl439 of L. pneumophila aggregate in the group of eukaryotic STPK, close to the STPK originating from Entamoeba histolytica (Figure 2). Mycobacterium tuberculosis, which as L. pneumophila blocks the phagosome-lysosome fusion, produces eleven STPK of eukaryotic type13. In particular, the STPK PknG of M. tuberculosis is an inhibitor of the phagosome-lysosome fusion and a promoter of intracellular survival14. The STPK domain of Lpp0267/Lpl0262 is related to the PknG and to the STPK YpkA of Y. pseudotuberculosis (Figure 2), an enzyme which is translocated in the eukaryotic cells where it corrects the defenses of the host by interfering with the transduction paths of the eukaryotic signal15. This suggests that the STPK of L. pneumophila can likewise modulate the transduction mechanisms of the eukaryotic signal and can modify the routing paths of the host cell. Twenty proteins contain ankyrin domains (Table XXIN), sequences repeated in tandem of around 33 amino acids which represent one of the modular interaction protein-protein patterns, the most current being eukaryotic. To date, the only prokaryotic genomes known for coding large families of proteins of the ankyrin domain are Coxiella burnetii and Wolbachia pipientis, which code 13 and 23 elements, respectively16,17. Similar to L. pneumophila, C burnetii is an intracellular pathogen which is extremely well-adapted to life inside the eukaryotic phagolysosome, and W. pipientis is an "endosymbiont" parasite living in the reproductive cells of a large variety of arthropods. Therefore, the ankyrin domains can be implied in a common microbial mechanism for manipulating the physiology of the host cell. A possible function of the proteins containing the repeated ankyrin sequences of L. pneumophila is to modify the
interactions with the host cytoskeleton, given that it is thought that numerous eukaryotic ankyrin proteins are binders between the membranous proteins and the cytoskeleton18 and are important for targeting proteins towards the plasmic membrane or towards the endoplasmic reticulum. The ankyrin domains are likewise compounds of transcription regulators, suggesting that they can influence the expression of the genes of the host cell as proposed for Ehrlichia phagocytophila19. In effect, one of the repeated ankyrin proteins of L. pneumophila (Lpp3991/Lpl0559) likewise contains a eukaryotic SET domain, known for binding the chromatin host and for influencing the expression of the genes of the host cell20. In opposition to the ankyrin proteins of W. pipientis, none of the repeated ankyrin proteins of L. pneumophila contains a peptide signal17. Instead of this, certain of them could be secreted by means of the secretion system of type TV, a way which is independent of the typical targeting signals. The final stages in the intracellular life cycle of L. pneumophila are to kill and escape from its host cell, a mechanism which is still not understood. One class of proteins which can affect the control of the division of the host cell (Lpp2082/Lpl2072, Lpp2486 and Lpp0233/Lpl 10234) separates eukaryotic F box domains, sites of protein- protein interactions. Generally, Lpl2072, Lpp2486 and Lpp0233/Lpll0234) separates eukaryotic are associated with other interaction domains21. Similarly, two of the identified F box proteins are associated with a repeated ankyrin sequence or a double- spooling pattern, respectively (Table XXIV). The proteins of the F box, assembled in SCF ubiquitin-ligase complexes, determine which substrates are going to be targets for ubiquitination and subsequent proteolysis by the proteasome. Given that the targeted substrates comprise promoters and inhibitors of the cellular cycle as well as transduction compounds of the signal , the F box proteins can regulate the division and cellular differentiation. To our knowledge, the only protein of prokaryotic F box described is VirF of Agrobacterium tumefaciens, a protein which, it is thought, interacts with the proteins of the host by means of its F box domain to target it for proteolysis23. Another pattern implied in the eukaryotic ubiquitination is the U box pattern. The protein Lpp2887, present in the Paris strain but not in the Lens strain, contains such a pattern. It is apparently the first recognized in a prokaryotic organism. Additional proteins of eukaryotic type identified in the genome of the two strains of L. pneumophila are sphingosine-1 -phosphate lyase (Lpp2128/Lpl2102) and two secreted apyrases (Lppl033, Lppl880/Lpll000, Lpll869), suggesting that L. pneumophila modulates the cycle of the host cell to its advantage. The broadly
expressed lyase sphingosine-1 -phosphate enzyme catalyses the essentially irreversible splitting of the sphingosine-1 -phosphate signaling molecule, a product of the degradation of sphingomyeline which regulates cellular proliferation and cellular death in the eukaryote. In effect, the overexpression of the sphingosine-1 -phosphate lyase can induce apoptosis in the eukaryote, identifying this enzyme as a double modulator of the metabolism of the sphingosine-1 -phosphate and of the ceramid, as well as a regulator of the decisions of cellular direction24. In addition, the sphingosine-phosphate plays a central role in the development of the Dictyostelium discoideum amoeba, given that interruption to the gene results in aberrant distribution of the actin, an abnormal moφhogenetic phenotype and a viability occurring during the stationary phase25. The two apyrases of L. pneumophila are the only ones identified in the prokaryote. The family of apyrase protein comprises enzymes capable of splitting the tri- and diphosphates nucleotides (NTP and NDP) in a calcium- or magnesium-dependent manner. The apyrase was isolated in the autophagy26 vacuole suggesting that these two proteins influence the destiny of the phagosome of L. pneumophila in diminishing the concentration of NTP and NDP during parasitism of the cell. 246 proteins (7.6 %) in the Paris strain and 231 proteins (1.1 %) in the Lens strain have likewise been identified with double-spooling domains provided (CC), many of which likewise show slight similarities with eukaryotic proteins. The CC domains facilitate the protein-protein interactions either for the multimerisation of protein or the macromolecular recognition. Therefore, double-spooling domains can target proteins at the appropriate locale in the eukaryotic host. Interestingly, all the new substrates (SidA- H, SdeC) of the secretion system of type IV identified by Luo and Isberg27 as well as LidA, LepA and LepB contain double-spooling domains. To affect the eukaryotic cell, L. pneumophila must translocate these proteins of eukaryotic type to the host cytoplasm. As a consequence these proteins are candidate substrates for the secretion system of type IV, as shown for VirF in A. tumefaciensn or 1
RalF in L. pneumophila .
Secretion system At the centre of the pathogenesis of L. pneumophila are the loci dot/icm, which together direct the assembling of secretion apparatus of type IV. Even though the two strains contain the complete loci dot/icm, their sequences have variations. In effect, the comparison of the sequence of dot/icm genes of 18 different strains of L. pneumophila
has identified a wide range of variations in sequence, and has placed the strains in seven phylogenetic different groups29. However, no correlation with the virulence is apparent. A novel factor of putative virulence of L. pneumophila, limited to the Paris strain, is a provided auto-transporter protein. Lpp0779 contains several marks of secretion systems of type V comprising a peptide of N-terminal head for secretion across the internal membrane and a specialized C-terminal domain which forms a pore in the other membrane through which the domain clone passes to the surface of the cell30. Its domain clone is a compound of known repeated sequences of haemagglutinin to be implied in the cell-cell aggregation and extremely similar to those of the auto- transporter of AIDA-I and Ag43 of Escherichia coli, two proteins implied in the virulence. The bacterial surface protein AIDA-I facilitates adherence to the mammal cells31, while Ag43 imparts not only a low level of adhesion to certain mammal cells, but likewise facilitates auto-aggregation which is important for the formation of biological film32. In a similar way, the auto-transporter of L. pneumophila can be implied in adhesion to the host cell and the formation of biological film. In opposition to AIDA-I and Ag43, the auto-transporter of L. pneumophila does not have an RGD pattern implied in the bond with the human integrins, and can thus have another interaction domain. The auto-transporter was acquired in the same way by horizontal gene transfer as suggested from its numerous IS upstream and from GC contents of 41 % which exceed the average of the genome of 38 %. Studies of the distribution of this gene in clinical and environmental strains of L. pneumophila in combination with the study of its function must provide knowledge of its importance. In addition, the two strains of L. pneumophila contain a secretion method by translocation (Tat) with combined arginine (TatAB and TatC) and completes the secretion systems of type I and II. The system of type II coded by the genes IspA, IspD-J is required for the secretion of several enzymes such as lipase A and B (Lpp0533, Lppl l59/Lpl0509, Lpll l64), phosphatase acid Map and SurE (Lppl l20, Lppl245/Lpll l24, lpll245), lysophospholipase A . (Lpp2291/Lpl2264) and phospholipase PlaB (Lppl568/Lpl422), proteins which are all present in the two strains.
Metabolism The metabolic paths utilized by L. pneumophila for multiplying inside the eukaryotic cells are not known. The bacteria seems to prefer the proteic substrates, given that a large number of absoφtion and degradation systems of oligopeptide and
5/049642 95
amino acid are coded in the genome. In particular, apart from the elastase homologue ProA of Pseudomonas aeruginosa (Lpp0532/lpl0508), three secreted paralogue metalloproteases as well as 46 additional peptidases are present. By way of opposition, systems for the absoφtion of sugar are rare, even though the complete ways of Embden-Meyerhof and Entner-Doudoroff are present. In the two strains of L. pneumophila no absoφtion system of type PTS was identified. However, certain of the transport systems of type 55 ABC can be implied in the absoφtion of sugar, given that the bacteria has some systems for the degradation of complex sugars, such as trehalase, polysaccharide deacetylase, glucoamylase of type eukaryotic (Lpρ0489/Lpl0465), β-hexosaminidase and chitinases (Lppl 117/Lpll 121). The two strains code for proteins highly homologous to transporters of glycerol phosphate ABC (Lppl 696, Lppl 695, Lppl694/Lpll695, Lpll694, Lpll693), and for a hexose phosphate transporter (Lpp2623/Lpl2474), which can be important during intracellular growth. We have likewise identified several enzymes probably implied in the utilization of meso- inositol, which can interfere with the signaling of the host cell facilitated by this intracellular messenger. L. pneumophila is provided for coding for an extensive aerobic respiratory chain constituted by NADH deshydrogenase, cytochrome-dependent succinate deshydrogenase, ubiquinol-cytochrome reductase and four terminal oxydases, which guarantee the capacity to adapt to changing oxygen tensions (one cytochrome aai, two quinol-cytochromes of type bd and one quinol cytochrome oxidase of type o). The latter oxidase is absent in the Lens strain. Systems implied in anaerobic respiration are apparently absent in all the strains. The two genomes code for an ATP synthase of type F0Fι typical of γ-proteobacteria, whereas the Paris strain codes for a second ATP synthase similar to the non-characterized systems of archeobacteria and bacteria marines. L. pneumophila likewise codes for at least four sodium/proton anti-carriers (Lppl464, Lpp2448, Lpp0868, Lpp0667/Lpll519, Lpl2304, Lpl0839, Lpl0651), which modulate presumably the proton and sodium gradient across the cytoplasmic membrane. As a consequence, a sodium motor force can be utilized for the cellular activities. In this respect, the presence of a compound of polar flagellar motors of type sodium, MotY, thus two significantly different aggregates of gene MotA-MotB, leads to the prediction that mobility can be activated by the sodium motor forces as well as the proton forces. One particular characteristic of L. pneumophila is differentiation in a mature intracellular form which accumulates inclusions of poly-hydroxybutyrate in the form of
carbon and energy reserve. As a consequence, the Paris strain codes for four paralogue poly-hydroxybutyrate synthases and the Lens strain codes for three paralogue poly- hydroxybutyrate synthases (Lpp2323, Lpp2038, Lpp2214, Lpp0650/Lpll055a-b, Lpl2186, Lpl0634).
Physiological adaptation and regulation of the gene In accord with this intracellular life style the regulatory repertoire is rather small. Analysis of the genome has identified 92 transcription regulators (79 in the Lens strain), which represent only 3.0 % of the genes provided. L. pneumophila codes for six sigma putative factors, the homologous of rpoD, rpoH, rpoS, rpoN, fliA and the sigma factor rpoE of type ECF. The number of systems with two compounds (13 histidine kinases and 14 response regulators) is likewise low. The most abundant class of regulators belongs to the GGDEF/EAL (23) family. Present in numerous bacteria, comprising Vibrio cholerae (41), P. aeruginosa (33), Wolinella succinogenes (26), and E. coli (19), these regulators contain two sub- domains, GGDEF and EAL. Of the 23 regulators identified, 10 separate only one GGDEF domain, 3 in the Paris strain and 2 in the Lens strain contain an EAL domain, and 10 in the Paris strain and 11 in the Lens strain present a combination of the two. The role of these regulators in L. pneumophila is unknown, but in other bacteria these regulators play a role in aggregation, the formation of biological film or mobility by muscular contraction. In L. pneumophila, the cyclic AMP can likewise translate the cellular signals given that the genome codes for five adenylate cyclases of class III (Lppl446, Lppl 131, Lppl704, Lppl277, Lpp0730/Lpll538, Lpll l35, Lpll703, Lpll276, Lpl0710). In P. aeruginosa, CyaB, an adenylate cyclase of class III, is implied in the regulation of the virulence of gene 33. However, L. pneumophila does not contain the orthologue of Vfr, the dependent AMPc regulator of P. aeruginosa, but it codes for five proteins with AMPc bonding patterns (Lpp3069, Lppl482, Lpp2063, LppOόl l, Lρp2777/Lpl2926, Lpll501, Lpl2053, Lρl0592a-b-c, Lpl2648). As for P. aeruginosa, these adenylate cyclases of class III can comprise environmental signals extending from the nutritional content of the surrounding medium to the presence of host cells and can control the expression of the virulence of the gene in consequence.
5/049642 97
Heightened plasticity of the genomes of L. pneumophila The two genomes exhibit an astonishingly high plasticity and a diversity of the genome. Comparison of the chromosomes has identified a preserved skeleton of 2664 genes but 280 and 428 genes (10 and 14 %) are specific to the strain (Figure 3). Given that the two strains analyzed belong to the same species and the same serogroup, this diversity was unexpected. For example the comparison of the genomes of two strains of Salmonella enterica of serotype Typhi identifies only 2 % for each of the genes specific to the strain34. The specific genetic equipment of the Paris strain contains a certain number of regulators (three homologous of CsrA, 13 transcription regulators), of additional repeated ankyrin sequences and proteins of type eukaryotic (Table XXIII and Table XXIV) as well as several restriction modification genes (modification methylases of the DNA, endonucleases), which can explain the low competence (personal observation) and the high genomic stability of the Paris strain. The Lens strain contains fewer specific regulators (4) and four specific proteins with eukaryotic domains (Table XXIII), two of which are repeated sequences of ankyrin proteins, suggesting that the Paris strain is a particularly well-equipped strain. The genomes of L. pneumophila have undergone rearrangements of multiple genomes. The important synteny in the genome between the Paris and Lens strain is interrupted by inversion of 260 kb, insertion of 130 kb in the Paris strain (or deletion in the Lens strain) and by deletions and smaller multiple insertions. The fragment of 130 kb is flanked by an ARnt gene and codes for a putative integrase, suggesting a structure similar to the islets of pathogenicity of the enterobacteria. It contains an ATP synthase, chemiosmotic flow systems (cebABC, cecABC) and the genes cadAl, ctpA, copAl, copA2 coding for ATP-dependent flow pumps, as was proven to be induced in the macrophages35 and separate the pφA-lvrABC gene aggregate, present within a pathogenicity islet of 65 kb in the Philadelphia strain36. With the exception of abovementioned genes, this pathogenicity islet is absent from the Paris strain and from the Lens strain. However, the corresponding chromosomal site in the Paris strain is the insertion site of an integrative plasmid discussed below. Therefore, these two regions can be hot points for genomic rearrangements. Genomic variation is likewise evident from its network of mobile elements comprising ten integrases, 58 insertion sequences (34 complete and 24 truncated) thus as proteins relevant to phages. In addition, the genomes contain a large number of repeated sequences organized in the form of repeated inverse sequences, which recall the ERIC sequences of the enterobacteria.
These LeRIC (Repeated Intergenic Consensus of Legionella) fall into 7 classes present in numerous copies (for example 80, 18, 18, 25, 9, 9 and 6 in the Paris strain). L. pneumophila Paris and Lens contain lvh, a region which codes for a second secretion system of type IV previously characterized in the Philadelphia strain37. One interesting observation is that the lvh region of L. pneumophila Paris is coded on a region of 36 kb which exists either integrated into the chromosome or excised in the form of a multicopy plasmid (unpublished data). This pattern is similar to that described for the unstable element of 30 kb of the Olda strain, which is possibly phage-derived and is implied in the phase variation38. The GC content of the lvh region (43 %) is different to that of the remainder of the chromosome (38 %) and it contains certain genes related to phages, suggesting possible phagic origin. However, the exact excision and integration mode is still not understood. An attractive hypothesis is that the integration and excision of particular regions of the chromosome is a mechanism specific to L. pneumophila for boosting versatility. The second plasmid of the Paris strain (132 kb) comprises known virulence factors, mobile genetic elements and genes of antibiotic resistance. The regulator system with two lφR-lskS compounds present on this plasmid was found on a plasmid of Legionella longbeachae implied in the virulence of these species39. Heightened preservation (93 to 98 % of protein identity) of the six gene sequences on the plasmid of 135 kb along L. longbeachae can indicate a recent horizontal transfer between L. pneumophila and L. longbeachae. L. pneumophila Lens contains a plasmid of 60 kb which codes for several proteins homologous to the transfer region of the F plasmid of E. coli. All three plasmids of the Paris and Lens strain code for a paralogue of CsrA, a repressor of the transmission traits and activator of replication40. Although the role of the plasmids in the virulence of L. pneumophila has yet to be determined, the correlation between strains containing a plasmid and the viralence in a mouse model was described ' . In addition, L. pneumophila strains with plasmids seem to persist longer in the environment than those strains not having plasmids42. The identification in the clinical isolates of the plasmids coding for factors of putative virulence is another indication of the importance of the plasmids for the pathogenicity of Legionella. The genomes of L. pneumophila display a plasticity likewise at the gene level. The loci enh, implied in the entry in the host cells43, are present in the Paris and Lens strains. One of the proteins coded by these loci is RtxA, which contributes to entry,
adherence, cytotoxicity, pore formation43 and intracellular routing in the amoeba44. Unlike the AA10010 strain in the Paris strain, rtxA is fused with aφB and a second protein with about 30 sequences repeated in tandem extremely preserved de 549 bp. A similar structure is coded by the Lens strain, however we have identified two patterns within the repeated region, the two being different to that of the Paris strain. However, the number of repetitions seems to be the same (Figure 4). It is possible that the variations in number and sequence of the repeated sequences contribute to the multiplicity and likewise to the virulence of L. pneumophila. In accordance with the relative plasticity in their genome, the strains of L. pneumophila code for organelles of type IV bacterial pili, which are required for natural competence for the transformation of the DNA45. The organization of the genes coding for type IV bacterial pili is similar to that of P. aeruginosa where they are crucial for bacterial adherence and colonization of the mucosal surfaces and for mobility by muscular contraction. Another mechanism in L. pneumophila contributing to the plasticity of the genome is conjugated transfer facilitated by the secretion of type IV of plasmids46 and chromosomal DNA47.
Conclusion Analysis of the sequences of the genome of the clinical of L. pneumophila Paris and Lens strains and its comparison identifies L. pneumophila as an extremely versatile organism, which demonstrates a plasticity and an extensive genomic diversity. The excision and integration of plasmids or genes can be a mechanism which L. pneumophila exploits for adapting to different environments. Its large cohort of proteins of eukaryotic type is provided for manipulating the host cell to the advantage of the pathogen (Figure 5). Eucaryotic proteins of putative origin have likewise been identified in other intracellular pathogens, comprising Coxiella, Wolinella, Agrobacterium, Mycobacterium and Ehrlichia, but currently L. pneumophila sequences them as prokaryotic with the greatest variety of proteins of eukaryotic type. Presumably, during its co-evolution with free living amoeba, the L. pneumophila pathogen acquires DNA by horizontal transfer from its host or by convergent evolution. These proteins can then likewise contribute to the infection of human macrophages. By being based on the genomic sequences future comparative and functional studies are going to enable survival tactics of intracellular parasites to be defined, and to identify the special attributes of endemic and epidemic L. pneumophila. To combat the menace of L.
5/049642 100
pneumophila resistant to chemical products widely utilized for decontamining public water systems, comprising hospitals, the genomic sequence can stimulate the identification of targets for novel active biocides against L. pneumophila.
Methods Preparation and sequencing of the DNA. Paris and Lens strains of L. pneumophila were cultivated on agar BCYE at 37°C over 3 days and the chromosomal DNA was isolated by utilizing standard protocols. The cloning, sequencing and assembling were completed as described previously48. For the two genomes, two libraries (inserts of 1-2 kb and 2-3 kb) were generated by random mechanical chiseling of the genomic DNA and cloning in pcDNA-2,1 (Invitrogen). A hook was obtained by terminal sequencing of clones from a BAC library constructed as described previously49 by utilizing plndigoBac (Epicentre) as vector. For the Paris strain of L. pneumophila an insert library of average size (8-10 kb) was constructed in the low-number vector of copies pSYX34. The purification Plasmide DNA was produced either with Montage Plasmide Miniprep96 Kit (Millipore) or by utilizing the DNA sequencing matrix amplification kit TempliPhi (Amersham Biosciences). The sequencing reactions were created by utilizing the sequencing reactions kit ABI PRISM BigDye Terminator and an analyzer 3700 or a 3730 XI Genetique Analyzer (Applied Biosystems). 47,200 sequences for the Paris strain of L. pneumophila, and 47,231 sequences for the Lens strain each originating from four libraries were obtained and assembled and finished as described previously 48. Annotation and analysis. The definition and annotation of the coding sequences (CDS) is as was described previously48 by utilizing the Boϊte CAAT 50 software. All the CDS provided were checked visually. The predictions on function were based on preferred BLASTp similarity and on analysis of patterns by utilizing the PFAM, Prosite and SMART databases. We have identified orthologous genes by better BDERNIERE reciprocal correlation and FASTA comparisons. For identification of the double- spooling domains the publicly available software PairCoil and Coilscan were utilized. The pseudogenes have one or more mutations which prevent complete translation. Repetitive sequences of DNA were identified by BLASTN comparisons of the intergenic regions and of the complete genome. MFOLD software was utilized to predict the folding of the single sheet of DNA molecules.
URL. The sequence and annotation of the two genomes of L. pneumophila are at http://genolist.pasteur.fr/LegioList. For annotation and analysis we used PairCoil http.V/paircoil. lcs.mit.edu/cgi-bin/paircoil and Coilscan http://www.biology.wustl.edu/gcg/coilscan.html Bibliography for Example 7:
1. McDade, J.E. et al. Legionnaires' disease: isolation of a bacterium and demonstration of its role in other respiratory disease. N. Engl. J. Med. 297, 1197-1203 (1977).
2. Fields, B.S. The social life of Legionellae. in Legionella (ed. Marre R., Kwaik, Y.A., Bartlett, C, Cianciotto, N.P., Fields, B.S., Frosch, M., Hacker, J. & Luck, P.C.)
135-142 (ASM Press, Washington D.C., 2002).
3. Steinert, M., Hentschel, U. & Hacker, J. Legionella pneumophila: an aquatic microbe goes astray. FEMS Microbiol. Rev. 26, 149-162 (2002).
4. Fields, B.S., Benson, R.F. & Besser, R.E. Legionella and Legionnaires' disease: 25 years of investigation. Clin. Microbiol. Rev. 15, 506-526 (2002).
5. Atlas, R.M. Legionella: from environmental habitats to disease pathology, detection and control. Environ. Microbiol. 1, 283-293 (1999).
6. Molofsky, A.B. & Swanson, M.S. Differentiate to thrive: lessons from the Legionella pneumophila life cycle. Mol. Microbiol. 53, 29-40. (2004). 7. Yu, V.L. et al. Distribution of Legionella species and serogroups isolated by culture in patients with sporadic community-acquired legionellosis: an international collaborative survey. J. Infect. Dis. 186, 127-128 (2002).
8. Aurell, H. et al. Legionella pneumophila Serogroup 1 Strain Paris: Endemic Distribution Throughout France. J. Clin. Microbiol. 41, 3320-3322 (2003). 9. Goebl, M. & Yanagida, M. The TPR snap helix: a novel protein repeat pattern from mitosis to transcription. Trends Biochem. Sci. 16, 173-177 (1991).
10. Cirillo, S.L.G., Lum, J. & Cirillo, J.D. Identification of a novel loci involved in entry by Legionella pneumophila. Microbiology 146, 1345-1359 (2000).
11. Conover, G.M., Derre, L, Vogel, J.P. & Isberg, R.R. The Legionella pneumophila LidA protein: a translocated substrate of the Dot/Icm system associated with maintenance of bacterial integrity. Mol. Microbiol. 48, 305-321 (2003).
12. Nagai, H., Kagan, J.C, Zhu, X., Kahn, R.A. & Roy, CR. A bacterial guanine nucleotide exchange factor activates ARF on Legionella phagosomes. Science 295, 679- 682 (2002).
13. Cole, S.T. et al. Deciphering the biology of Mycobacterium tuberculosis from the complete genomic sequence. Nature 393, 537-544 (1998).
14. Walburger, A. et al. Protein Kinase G from Pathogenic Mycobacteria Promotes Survival Within Macrophages. Science 304,1800-1804 (2004). 15. Barz, C, Abahji, T.N., Trulzsch, K. & Heesemann, J. The Yersinia Ser/Thr protein kinase YpkA/YopO directly interacts with the small GTPases RhoA and Rac-1. FEBS Lett. 482, 139-143 (2000).
16. Seshadri, R. et al. Complete genomic sequence of the Q-fever pathogen Coxiella burnetii. Proc. Natl. Acad. Sci. U S A 100, 5455-5460 (2003). 17. Wu, M. et al. Phylogenomics of the Reproductive Parasite Wolbachia pipientis wMel: A Streamlined Genome Overrun by Mobile Genetic Elements. PLoS Biol 2, E69 (2004).
18. Hryniewicz-Jankowska, A., Czogalla, A., Bok, E. & Sikorsk, A.F. Ankyrins, multifunctional proteins involved in many cellular pathways. Folia Histochem. Cytobiol. 40, 239-249 (2002).
19. Caturegli P, A.K., Walls, J.J., Bakken, J.S., Madigan, J.E., Popov, V.L., Dumler, J.S. ankA: an Ehrlichia phagocytophila group gene encoding a cytoplasmic protein antigen with ankyrin repeats. Infect. Immun. 68, 5277-5283 (2000).
20. Jenuwein, T., Laible, G., Dorn, R. & Reuter, G. SET domain proteins modulate chromatin domains in eu- and heterochromatin. Cell. Mol. Life Sci. 54, 80-93 (1998).
21. Kipreos, E.T. & Pagano, M. The F-box protein family. Genome Biol. 1, Epub 2000 Nov 10 (2000).
22. Craig, K.L. & Tyers, M. The F-box: a new pattern for ubiquitin dependent proteolysis in cell cycle regulation and signal transduction. Prog. Biophys. Mol. Biol. 72, 299-328 (1999).
23. Schrammeijer, B. et al. Interaction of the virulence protein VirF of Agrobacterium tumefaciens with plant homologs of the yeast Skpl protein. Curr. Biol. 11, 258-262 (2001).
24. Reiss, U. et al. Sphingosine-phosphate lyase enhances stress-induced ceramide generation and apoptosis. J. Biol. Chem. 279, 1281-1290 (2004).
25. Li, G., Foote, C, Alexander, S. & Alexander, H. Sphingosine-1 -phosphate lyase has a central role in the development of Dictyostelium discoideum. Development 128, 3473-3483 (2001).
26. Biederbick, A., Rose, S. & Elsasser, H.P. A human intracellular apyrase-like
5/049642 103
protein, LALP70, localizes to lysosomal/autophagic vacuoles. J. Cell. Sci. 112, 2473- 2484 (1999).
27. Luo, Z.Q. & Isberg, R.R. Multiple substrates of the Legionella pneumophila Dot/Icm system identified by interbacterial protein transfer. Proc. Natl. Acad. Sci. U S A 101, 841-846 (2004).
28. Vergunst, A.C et al. VirB/D4-dependent protein translocation from Agrobacterium into plant cells. Science 290, 979-982 (2000).
29. Morozova, I. et al. Comparative sequence analysis of the icm/dot genes in Legionella. Plasmid 51, 127-147 (2004). 30. Desvaux, M., Parham, N.J. & Henderson, LR. The autotransporter secretion system. Res. Microbiol. 155, 53-60 (2004).
31. Benz, I. & Schmidt, M.A. AIDA-I, the adhesin involved in diffuse adherence of the diarrhoeagenic Escherichia coli strain 2787 (O126:H27), is synthesized via a precursor molecule. Mol. Microbiol. 6, 1539-1546 (1992). 32. Klemm, P., Hjerrild, L., Gjermansen, M. & Schembri, M.A. Structure- function analysis of the self-recognizing Antigen 43 autotransporter protein from Escherichia coli. Mol Microbiol. 2 51, 283-296 (2004).
33. Smith, R.S., Wolfgang, M.C. & Lory, S. An adenylate cyclase-controlled signaling network regulates Pseudomonas aeruginosa virulence in a mouse model of acute pneumonia. Infect. Immun. 72, 1677- 1684 (2004).
34. Deng, W. et al. Comparative Genomics of Salmonella enterica Serovar Typhi Strain. J. Bacteriol. 185, 2330-2337 (2003).
35. Rankin, S., Li, Z. & Isberg, R.R. Macrophage-induced genes of Legionella pneumophila: protection from reactive intermediates and solute imbalance during intracellular growth. Infect. Immun. 70, 3637-3648 (2002).
36. Brassinga, A.K. et al. A 65-Kilobase Pathogenicity Island Is Unique to Philadelphia- 1 Strains of Legionella pneumophila. J. Bacteriol. 185, 4630-3637 (2003).
37. Segal, G., Russo, J.J. & Shuman, H.A. Relationships between a new type IN secretion system and the icm/dot viralence system of Legionella pneumophila. Mol. Microbiol. 34, 799-809 (1999).
38. Luneberg, E. et al. Chromosomal insertion and excision of a 30 kb unstable genetic element is responsible for phase variation of lipopolysaccharide and other virulence determinants in Legionella pneumophila. Mol. Microbiol. 39, 1259-1271 (2001).
39. Doyle, R.M. & Heuzenroeder, M.W. A mutation in an ompR-like gene on a Legionella longbeachae serogroup 1 plasmid attenuates virulence. Int. J. Med. Microbiol. 292, 227-239 (2002).
40. Molofsky, A.B. & Swanson, M.S. Legionella pneumophila CsrA is a pivotal repressor of transmission traits and activator of replication. Mol. Microbiol. 50, 445-461
(2003).
41. Bezanson, G., Fernandez, R., Haldane, D., Burbridge, S. & Marrie, T. Virulence of patient and water isolates of Legionella pneumophila in guinea pigs and mouse L929 cells varies with bacterial genotype. Can. J. Microbiol. 40, 426-431 (1994). 42. Brown, A., Vickers, R.M., Elder, E.M., Lema, M. & Garrity, G.M. Plasmid and surface antigen markers of endemic and epidemic Legionella pneumophila strains. J. Clin. Microbiol. 16, 230-235 (1982).
43. Cirillo, S.L., Bermudez, L.E., El-Etr, S.H., Duhamel, G.E. & Cirillo, J.D. Legionella pneumophila entry rtxA gene is involved in virulence. Infect. Immun. 69, 508-517 (2001).
44. Cirillo, S.L., Yan, L., Littman, M., Samrakandi, M.M. & Cirillo, J.D. Role of the Legionella pneumophila rtxA gene in amoebae. Microbiology 148, 1667-1677 (2002).
45. Stone, B.J. & Kwaik, Y.A. Natural competence for DNA transformation by Legionella pneumophila and its association with expression of type IV pili. J. Bacteriol. 181, 1395-1402 (1999).
46. Vogel, J.P., Andrews, H.L., Wong, S.K. & Isberg, R.R. Conjugative transfer by the viralence system of Legionella pneumophila. Science 279, 873-876 (1998).
47. Miyamoto, H., Yoshida, S., Taniguchi, H. & Shuman, H.A. Virulence conversion of Legionella pneumophila by conjugal transfer of chromosomal DNA. J. Bacteriol. 185, 6712-6718 (2003).
48. Glaser, P. et al. Comparative genomics of Listeria species. Science 294, 849-852 (2001).
49. Buchrieser, C. et al. The 102-kilobase pgm locus of Yersinia pestis: sequence analysis and comparison of selected regions among different Yersinia pestis and Yersinia pseudotuberculosis strains. Infect. Immun. 67, 4851-4861 (1999).
50. Frangeul, L. et al. CAAT-Box, Contigs-Assembly and Annotation tool-box for genome sequencing projects. Bioinformatics 20, 790-797 (2004).
Table XXII : General characteristics of the two Legionella pneumophila genomes
5 Table XXIII: Proteins having the greatest similarity with eukaryotic proteins
L. pneumophila Produit provided Lφ.Lens G-C Percentage of protein identity Paris 61% over the entire length (AAR06292.1 lppl647 purC lpll640 38% Nicotania tabacum) exoA 58% over the entire length (EAA20230.1 lpp0702 lpl0684 39% exodeoxyribonuclease III Plasmodium yoelii yoelii) Precursor protein bond to 45% of 50% of the protein (AAL07519 lpp0321 34% RNA Solarium tubeosum) 50% over the entire length (AAB16855.1 lppll57 pyruvate decarboxylase lplll62 39% Arabidopsis thaliana) protein de biosynthesis de 49% over the entire length (AAC64375.1 lppl522 lpll461 38% thiamine NMT-1 Botryotiniafuckelianά) nuoE NADH 49% of 82% of the protein (BAA25988.1 lpp2832 lpl2701 38% dehydrogenase I chain E Homo sapiens) protein with hypersensitive 48% over the entire length (AAN17462.1 plpp0050 36% induced response Hordeum vulgare subsp. Vulgare) 48% over the entire length (XP_306643.1 lpp0634 hypothetical protein lpl0618 39% Anopheles gambiae) 45% over the entire length (NP_189431.2 lpP0965 protease lpl0935 39% Arabidopsis thaliana) 44%o over the entire length 372144.1 lpp2748 phytanoyl-coA dioxygenase lpl2621 36% B9 RRIBE®3SW
lpp0489 glucoamylase 32%, over the entire length (P42042 Arxula lpl0465 39% adeninivorans) lpp0955 cytokinin oxydase 32% over the entire length (NP_484368.1 lpl0925 39% Nostoc sp.) lpp0578 phytanoyl coA dioxygenase 31% over the entire length (EAA70100.1 lpl0554 36% Gibberella zeae) lpp0379 hypothetical protein 31% over the entire length (CAD21525.1 Lpl0354 39% Taenia solium) ectonucleoside triphosphate lpp 1033 diphosphohydrolase 25%ι over the entire length - nucleoside IpllOOO 40% (apyrase) phosphatase signature (Q9MYU4 Sus scrofa) 6-pyruvoyl-tetrahydropterin lpp2923 lpl2777 34% 26% over the entire length (NP 703938.1 synthase Plasmodium falciparum) lpp3071 38% over the entire length (AAF56122.1 zinc metalloproteinase lpl2927 38% Drosophila melanogaster) methyl trans ferase bonded to lpp2134 24% over the entire length (BAC98835.1 SAM lpl2109 35% Bombyx mori) ectonucleoside triphosphate lppl880 diphosphohydrolase 26% over the entire length (CAE70887.1 lpll869 39% (apyrase) Caenorhabditis briggsae) methyltransferase bonded to lpp2747 lpl2620 35% 33% of 56% of the protein (EAA20288.1 SAM Plasmodium yoelii yoelii) lpp2468 Cytochrome P450 20% of 75% of the protein (NP_487786.1 lpl2326 39% Nostoc sp.) protein bound to the nuclear lppl824 34% 19% of 40% of the protein (NP 082559.1 Mus membrane musculus) lpp 1665 uracyl DNA glycosylase lpll659 36% 27% of 80% of the protein (EAA36774.1 Giardia lamblia) condensation chromosome lppl959 Model of preserved chromosome condensation Ipl 1953 41% type 1 regulator
Lpp indicates the coding sequences (CDS) provided from L. pneumophilastrain Paris; Ipl indicates the coding sequences (CDS) provided from L. pneumophilastrain Lens, the lines in gray indicate the proteins which are likewise mentioned in Table 2B in terms of their preserved eukaryotic domains; the access numbers to the proteins are indicated between parentheses
Table XXIV: coding domains of L. pneumophila preferably protein found in eukaryotic proteins
L. pneumophila L. pneumophila Content identified unit putative function Paris Lens G-C enhC (lpp2692) enhC (lpl2564) 21 domains sel-1 39%
UdL(lppll 74)- UdL (lplll80) - 6 omains SP1- 1
EnhC paralog EnhC paralog 38% lppl310- EnhC lpll307 - E hC paralog paralog 4 domains sel-1 41% Invasion and traffic in lpp2174- EnhC lpl!303 - EnhC host cells paralog paralog 3 domains sel-1 40% lpll059 - EnhC paralog 7 domains sel-1 45% ralF (lpp!932) ralF lpll919 domain sec7 34% lpp0267 lpl0262 domain ser/thr kinase protein 38% lpp2626 lpl2481 domain ser/thr kinase protein 32% lppl439 lpll545 domain ser/thr kinase protein 36% lpp2065 lp2055 ankyrin repetition 37% lpp0037 lpl0038 ankyrin repetition 38% plpp0098 - ankyrin repetition 37% lpp2058 Ipl2048 ankyrin repetition 38% lpp0750 lpl0732 ankyrin repetition 35% lpp2061 lpl2051 ankyrin repetition 39% lpp2270 lpl2242 ankyrin repetition 34% lpp0503 lpl0479 ankyrin repetition 38% lpp 1905 - ankyrin repetition 35% lpp 1683 Ipl 1682 ankyrin repetition + domain SET Modulation functions of 33% host cells lpp2248 lpl2219 ankyrin repetition 39% lpp0202 - ankyrin repetition 38% lpp0469 Ipl0445 ankyrin repetition 38% lpp2517 lpl2370 ankyrin repetition 36% lppl 100 - ankyrin repetition 48% lpp0126 IplOlll ankyrin repetition 39% lpp0356 - ankyrin repetition 38% lpp2522 Ipl2375 ankyrin repetition 39% lpp0547 lpl0523 ankyrin repetition 40%
- Ipl 1681 ankyrin repetition 34%
- lpl2344 ankyrin repetition 35%
- lpl2058 ankyrin repetition 40%
Ipp2082 lpl2072 domain F-box + ankyrin repetition 36% lpp2486 - domain F-box + superhelice 34% lpp0233 Ipl0234 domain F-box 39% Control division, evasion lpp2887 - 2 domains U-box 35% host cells lpp2128 Ipp2102
Sphingosine-1- Sphingosine-1- phosphate lyase phosphate lyase 41%
Lpp indicates the coding sequences (CDS) provided from L. pneumophilastrain Paris; Ipl indicates the coding sequences (CDS) provided from L. pneumophilastrain Lens; the numbers indicate the number of domains identified in a protein; ser/thr indicates threonine serin
Example 9: Repeated sequences of DNA and secreted enzymes characteristic of Legionella pneumophyla A) Repeated sequence A repeated sequence was identified in the genome of the Legionella pneumophila Paris strain then sequenced completely. This sequence (SEQ ID 7074) is of 122 bp and is repeated 86 times in the genome of the L. pneumophila Paris strain. The preservation is from 81 to 100 % (0 to 19 mismatch) over the entire length for 53 copies and from 70 to 80 % over a length of at least 100 nucleotides for 33 copies. In the L. pneumophila Lens strain, there are 62 copies whereof the preservation of 29 copies is from 81 % to 95 % over the entire length. We have determined oligonucleotides specific to this sequence for its amplification by PCR. Tests on 15 strains each belonging to one of the serotypes of L. pneumophila (serotypes 1 to 15) have shown that this sequence is present in all serotypes of L. pneumophila. The test of 11 strains belonging to other species of the Legionella geme (L. miedadei, L. dumoffii, L. gormanii, L. longbeachae serogroup 1, L. jordanis, L. anisa, L. erythre, L. rubriluccns, L. quinlivani, L. moravica, L. taurinensis) with these oligonucleotides, thus databank research have shown that this sequence is specific to the Legionella pneumophila species and that it will be able to thus serve as an identifier of the species (see Tables XXV and XXVI and Figures 6 and 7). The high number of copies of this sequence in the genome will enable amelioration of the sensitivity of a PCR test or by hybridization, compared to a present sample in a single copy. SEQ ID 7074:
AGGACTTACGAAAAACCCCAAGATCAAGGCAAAAAATGTTTTTAATGAGG GAGTTTAGATAAACTAAATAACCGAATTAAAAACTTTTTTTAACAAAGAG ATTGGGATTTTTCGTAAGTCCT.
This sequence is an interesting target for diagnostics of L. pneumophila diagnostic by PCR or by other methods equivalent to PCR.
Among the primers used for amplification and detection of this repeated sequence, the following couple of primers can especially be cited:
5/049642 109
SEQ ID N° 7075: GAAAAACCCCAAGATCAAGGC and SEQ ID N° 7076: AGGACTTACGAAAAACCCCAA.
Table XXV: List of the DNA of the 15 reference serogroups of Legionella pneumophila
Number Name labo n°ATCC L pneumophila sgl Rl ATCC 33152 Lpneumoph la sg3 R2 ATCC 33155 L pneumophila sg2 Togus 1 R6 ATCC 33154 L pneumophila sg4 LosAngelesl R7 ATCC 33156 L pneumoph la sg5 subsp fraseri R8 ATCC 33216 L pneumophila sgό Chicago 2 R10 ATCC 33215 Lpneumoph la sg7 Chicago 8 R12 ATCC 33283 L pneumophila sg8 Concord 3 R13 ATCC 33096 L pneumophila sg9 IN-23-G1-E2 R14 ATCC 35289 L pneumophila sglO Leiden 1 R15 ATCC 43283 Lpneumoph la sgll 797-PA-H R18 ATCC 43130 Lpneumoph la sg!2 570-CO-H R19 ATCC 43290 L pneumophila sg!3 82A3105 R20 ATCC 43736 L pneumophila sg!4 1169-MN-H R21 ATCC 43073 L pneumophila sgl 5 Lansing3 R22 ATCC 35251
110
Table XXNI : list of the DΝA of the 11 reference species Legionella non pneumophila
B) Enzymes secreted The enzymes secreted and common to the three strains of L. pneumophila (Paris, Lens and Philadelphia) whereof the sequences are identified hereinbelow can be utilized especially as a target in colorimetric tests (or for their being made available) for detection of the presence or not of Legionella in a biological sample.
- the sequence lpp0489 (SEQ ID 4292) which codes for a precursor of glucoamylase (Glucan 1 ,4-alpha-glucosidase) of eukaryotic cell without homologue in the bacteria;
- the sequence lppl 117 (SEQ ID 6477) which codes for a potential secreted chitinase; and
- the sequences lppl 033 (SEQ ID 4267) and lppl 880 (SEQ ID 3675) which code for a protein similar to an ectonucleoside triphosphate diphosphohydrolase (apyrase) secreted from a eukaryotic cell.
Bibliography (beyond Example 8)
1. Altschul, S. F., T. L. Madden, A. A. Schaffer, J. Zhang, Z. Zhang, W. Miller, and D. J. Lipman. 1997. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. [Review] [90 refs]. Nucleic Acids Research. 25:3389-402.
2. Aurell, H., J. Etienne, F. Forey, M. Reyrolle, P. Girardo, P. Farge, B. Decludt, C. Campese, F. Vandenesch and S. Jarraud. 2003. Legionella pneumophila serogroup 1 strain Paris: Endemic distribution throughout France. J Clin Microbiol. 41 : 3320-3322
3. Birnboim, H. C. 1983. A rapid alkaline extraction method for the isolation of plasmid DNA. Methods Enzymol. 100:243-255.
4. Buchrieser, C, C. Rusniok, L. Frangeul, E. Couve, A. Billault, F. Kunst, E. Camiel, and P. Glaser. 1999. The 102 kb locus of Yersinia pestis : sequence analysis and comparison of selected regions among different Yersinia pestis and Yersinia pseudotuberculosis strains. Infect. Immun. 67:4851-4861.
5. Ewing, B., and P. Green. 1998. Base-calling of automated sequencer traces using phred. II. Error probabilities. Genome Res. 8:186-194. 6. Fitch, W. S. 1970. Distingishing homologous from analogous proteins. Syst. Zool
19:99-113.
7. Frangeul L, Glaser P, Rusniok C, Buchrieser C, Duchaud E, Dehoux P and Kunst F.
2004. CAAT-Box, Contigs-Assembly and Annotation tool-box for genome sequencing projects. Bioinformatics. in press. 8. Gordon, D., C. Abajian, and P. Green. 1998. Consed: a graphical tool for sequence finishing. Genome Res. 8:195-202.
10. Li, P., K. C. Kupfer, C. J. Davies, D. Burbee, G. A. Evans, and H. R. Garner. 1997.
PREVIO: A primer design program that applies base quality statistics for automated large-scale DNA sequencing. Genomics. 40:476-485. 11. Lukashin, A. V., and M. Borodovsky. 1998. GeneMark.hmm: new solutions for gene finding. Nucleic Acids Res. 15:1107- 1115.
12. Xu, S.Y., and Fomenkov, A. 1994. Construction of pSClOl derivatives with Camr and Tetr for selection or LacZ for blue/white screening. Biotechniques 17:57.
Table I: Correspondence of the numbers attributed to the genes of the Paris strain on its contigs with the numbers of SEQ ID identified in the list of sequences and position of nucleic sequences coding these genes on the sequence of these contigs with their putative function, as well as their specificity relative to the Philadelphia strain
Table II « Best-BlastP »: Putative function of the ORFs identified for the Paris strain
Table III : Correspondence of the numbers attributed to the contigs with the numbers of the SEQ ID identified in the list of sequences
Table IV: Correspondence between the contigs of Legionella pneumophila Philadelphia strain and numbering of their sequence in the list of sequences
Contig of the Philadelphia strain Seq Id Contigl SEQ ID N°3456 Contig2 SEQ ID N°3457 Contig3 SEQ ID N°3458 Contig4 SEQ ID N°3459 Contig5 SEQ ED N°3460 Contigό SEQIDN°3461 Contig7 SEQ ID N°3462 Contigδ SEQ ID N°3463 Contig9 SEQ ID N°3464 Contigl 0 SEQIDN°3465 Contigl 1 SEQ ID N°3466 Contigl 2 SEQIDN°3467 Contigl 3 SEQIDN°3468 Contigl 4 SEQ ID N°3469 Contigl 5 SEQIDN°3470 Contigl 6 SEQIDN°3471 Contigl 7 SEQIDN°3472 Contigl 8 SEQIDN°3473 Contigl 9 SEQIDN°3474 Contig20 SEQ ID N°3475 Contig21 SEQIDN°3476 Contig22 SEQ ID N°3477 Contig23 SEQ ID N°3478 Contig24 SEQ ID N°3479 Contig25 SEQ ID N°3480 Contig26 SEQIDN°3481 Contig27 SEQ ID N°3482 Contig28 SEQ ID N°3483 Contig29 SEQ ID N°3484 Contig30 SEQ ID N°3485 Contig31 SEQIDN°3486 Contig32 SEQ ID N°3487 Contig33 SEQ ID N°3488 Contig34 SEQ ID N°3489 Contig35 SEQID °3490 Contig36 SEQIDN03491 Contig37 SEQ ID N°3492 Contig38 SEQ ID N°3493 Contig39 SEQ ID N°3494 Contig40 SEQ ID N°3495 Contig41 SEQ ID N°3496 Contig42 SEQ ID N°3497 Contig43 SEQ ID N°3498 Contig44 SEQ ID N°3499
Contig45 SEQ ID N°3500 Contig46 SEQ ED N°3501 Contig47 SEQ ID N°3502 Contig48 SEQ ED N°3503 Contig49 SEQ ID N°3504 Contig50 SEQ ED N°3505 Contig51 SEQ ID N°3506
Table V: Surface proteins of Legionella pneumophila Paris strain The proteins of surfaces specific to the Paris strain are indicated in bold.
Table VI: Proteins implied in biosynthesis of polysaccharides having a cellular envelope of Legionella pneumophila
Table X: Correspondence of the numbers attributed to the contigs of L. pneumophila Philadelphia with the numbers of the SEQ ID identified in the list of sequences
Contigl SEQ ID N∞7061 Contig2 SEQ ED Nco7062 Contig3 SEQ ID Noo7063 Contig4 SEQ ID Nco7064 Contig5 SEQ ED Noo7065 Contig6 SEQ ED Noo7066 Contig7 SEQ ED Noo7067 Contigδ SEQ ED Noo7068 Contig9 SEQ ED Noo7069 Contigl 0 SEQ LD N∞7070 Contigl 1 SEQ ED Noo7071 Contig 12 SEQ ED Noo7072 Contigl 3 SEQ ED Nco7073
Table XI: List of the seq uences of L. pneumop this strain relative to the Paris and Lens strains and position of these sequences on the contigs
Indication on the specifics of the Philadelphia strain
IPF Lp Philadelphia Contig SEQ ED Position 1 Position2 1007.1 CONTIG9 SEQ ID N°7069 1062411 1062962 10563.1 CONTIG13 SEQ ID N°7073 1463 2446 3775.3 CONTIG13 SEQ ED N°7073 1463 2446 1067.1 CONTIG7 SEQ ID N°7067 163133 163567 1980.3 CONTIG7 SEQ ID N°7067 163628 163918 1102.1 CONTIG7 SEQ ID N°7067 189792 190835 1109.1 CONTIG7 SEQ ED N°7067 195874 198036 4935.1 CONTIG9 SEQ ED N°7069 1604318 1605460 7686.1 CONTIG9 SEQ ID N°7069 1604318 1605460 1771.2 CONTIG8 SEQ ID N°7068 552424 553377 1773.1 CONTIG9 SEQ ID N°7069 961264 962745 1296.1 CONTIG9 SEQ ED N°7069 961264 962745 1297.1 CONTIG9 SEQ ID N°7069 959864 960817 1298.1 CONTIG9 SEQ ID N°7069 959562 959810 1302.1 CONTIG9 SEQ ED N°7069 958145 958699 1303.1 CONTIG9 SEQ ID N°7069 957452 957922 1307.1 CONTIG9 SEQ ED N°7069 956035 956523 1309.1 CONTIG9 SEQ ID N°7069 955209 955589 1310.1 CONTIG9 SEQ ED N°7069 954726 955034 1312.1 CONTIG9 SEQ ID N°7069 953857 954711 1313.1 CONTIG9 SEQ ED N°7069 953085 953864 1315.1 CONTIG9 SEQ ED N°7069 952598 953161
1319.1 CONTIG9 SEQ ID N°7069 950926 951858
1320.1 CONTIG9 SEQ ID N°7069 948772 950907
1321.1 CONTIG9 SEQ ED N°7069 948180 948743
1322.1 CONTIG9 SEQ ID N°7069 947726 948187
1323.1 CONTIG9 SEQ ID N°7069 947134 947706
1324.1 CONTIG9 SEQ ID N°7069 946011 946685
1325.1 CONTIG9 SEQ ID N°7069 945182 945862
1327.1 CONTIG9 SEQ ID N°7069 944125 945009
1328.1 CONTIG9 SEQ ID N°7069 943800 944288
1330.1 CONTIG9 SEQ ID N°7069 943233 943535
1331.1 CONTIG9 SEQ ID N°7069 942938 943243
1332.1 CONTIG9 SEQ ID N°7069 942371 942934
1333.1 CONTIG9 SEQ ID N°7069 941655 942368
1334.1 CONTIG9 SEQ ID N°7069 940367 941653
1337.1 CONTIG9 SEQ ID N°7069 940003 940242
1339.1 CONTIG9 SEQ ID N°7069 937439 940036
1340.1 CONTIG9 SEQ ID N°7069 937111 937446
1341.1 CONTIG9 SEQ ID N°7069 936506 937114
1342.1 CONTIG9 SEQ ID N°7069 935514 936509
1343.1 CONTIG9 SEQ ID N°7069 934830 935513
1344.1 CONTIG9 SEQ ID N°7069 933019 934890
1345.1 CONTIG9 SEQ ID N°7069 932244 933026
1346.1 CONTIG9 SEQ ID N°7069 931832 932254
1347.1 CONTIG9 SEQ ID N°7069 930450 931835
2328.1 CONTIG3 SEQ ID N°7063 749 1696
1348.1 CONTIG9 SEQ ID N°7069 927724 930444
1350.1 CONTIG9 SEQ ED N°7069 927256 927720
1353.1 CONTIG9 SEQ ID N°7069 925299 927188
1355.1 CONTIG9 SEQ ID N°7069 919333 925278
1356.1 CONTIG9 SEQ ID N°7069 918010 919137
1358.1 CONTIG9 SEQ ID N°7069 916581 917417
1359.1 CONTIG9 SEQ ID N°7069 916079 916570
1360.1 CONTIG9 SEQ ID N°7069 914469 916049
1361.1 CONTIG9 SEQ ID N°7069 913897 914271
1365.1 CONTIG9 SEQ ID N°7069 911529 913049
1366.1 CONTIG9 SEQ ID N°7069 910007 911458
1367.1 CONTIG9 SEQ ID N°7069 909405 909701
1368.1 CONTIG9 SEQ ID N°7069 908527 909309
1369.1 CONTIG9 SEQ ID N°7069 908004 908504
1370.1 CONTIG9 SEQ ID N°7069 907151 907993
1371.1 CONTIG9 SEQ ID N°7069 905264 907069
1373.1 CONTIG9 SEQ ID N°7069 903498 905264
1376.1 CONTIG9 SEQ ID N°7069 902217 902987
1378.1 CONTIG9 SEQ ED N°7069 900968 901987
1380.1 CONTIG9 SEQ ID N°7069 899639 900985
1382.1 CONTIG9 SEQ ID N°7069 898691 899314
1383.1 CONTIG9 SEQ ID N°7069 898266 898532
1384.1 CONTIG9 SEQ ID N°7069 897979 898281
1386.1 CONTIG9 SEQ ED N°7069 897062 898066
1435.1 CONTIG9 SEQ ID N°7069 864915 866060
1492.1 CONTIG9 SEQ ID N°7069 821283 821579
1494.1 CONTIG9 SEQ ID N°7069 820792 821109
1500.1 CONTIG9 SEQ ED N°7069 814942 815592
1501.1 CONTIG9 SEQ ED N°7069 814765 815241
1503.1 CONTIG9 SEQ ID N°7069 813201 813485
1518.1 CONTIG9 SEQ ID N°7069 800919 803852
1519.1 CONTIG9 SEQ ID N°7069 799375 800655
1520.1 CONTIG9 SEQ ED N°7069 796924 799023
1538.1 CONTIG9 SEQ ID N°7069 775795 776787
1627.1 CONTIG8 SEQ ID N°7068 454194 455027
1631.1 CONTIG8 SEQ ID N°7068 456917 457408
1635.1 CONTIG8 SEQ ID N°7068 459876 460751
1663.1 CONTIG8 SEQ ED N°7068 477274 477810
1674.1 CONTIG8 SEQ ID N°7068 488306 490213
1676.1 CONTIG8 SEQ ID N°7068 491037 491288
1723.1 CONTIG8 SEQ ED N°7068 522733 523200
1724.1 CONTIG8 SEQ ED N°7068 523306 523560
1725.1 CONTIG8 SEQ ID N°7068 523670 523945
1749.1 CONTIG8 SEQ ID N°7068 537066 537341
1760.1 CONTIG8 SEQ ID N°7068 544546 545574
1767.1 CONTIG8 SEQ ID N°7068 549553 550482
1772.1 CONTIG6 SEQ ID N°7066 779 1804
1785.1 CONTIG9 SEQ ID N°7069 970464 970808
1787.1 CONTIG9 SEQ ID N°7069 970976 971416
1892.1 CONTIG7 SEQ ID N°7067 76315 77967
3771.3 CONTIG13 SEQ ED N°7073 43581 46268
1952.1 CONTIG9 SEQ ID N°7069 1328145 1328393
1981.1 CONTIG9 SEQ ED N°7069 1347920 1348357
2005.1 CONTIG9 SEQ ED N°7069 1364695 1366614
2006.1 CONTIG9 SEQ ID N°7069 1366816 1368231
2026.1 CONTIG9 SEQ ID N°7069 1380350 1380739
2059.1 CONTIG9 SEQ ID N°7069 1407671 1408075
2066.1 CONTIG9 SEQ ID N°7069 1418764 1421481
2083.1 CONTIG9 SEQ ED N°7069 1436828 1439185
2084.1 CONTIG9 SEQ ID N°7069 1439377 1440240
2086.1 CONTIG9 SEQ ED N°7069 1440346 1441680
2125.2 CONTIG9 SEQ ID N°7069 1470612 1471472
2132.1 CONTIG11 SEQ ED N°7071 144883 145770
2133.1 CONTIG 11 SEQ ID N°7071 144142 144882
2134.1 CONTIG11 SEQ ED N°7071 143221 143718
2135.1 CONTIG11 SEQ ID N°7071 143204 144145
2141.1 CONTIG11 SEQ ID N°7071 137310 137879
2202.1 CONTIG11 SEQ ID N°7071 87995 89587
2311.1 CONTIG8 SEQ ED N°7068 553457 554560
2312.1 CONTIG8 SEQ ED N°7068 554678 555862
2314.1 CONTIG8 SEQ ID N°7068 555930 556454
2315.1 CONTIG8 SEQ ED N°7068 556495 556839
2316.1 CONTIG8 SEQ ID N°7068 556651 558360
2317.1 CONTIG8 SEQ ED N°7068 558425 558796
2319.1 CONTIG8 SEQ ED N°7068 559260 560681
2321.1 CONTIG8 SEQ ID N°7068 560851 562347
2323.1 CONTIG8 SEQ ID N°7068 562541 562981
2324.1 CONTIG8 SEQ ID N°7068 563118 563813
2325.1 CONTIG8 SEQ ID N°7068 563899 564930
2326.1 CONTIG8 SEQ ID N°7068 564927 566123
2330.1 CONTIG8 SEQ ID N°7068 567385 567954
2333.1 CONTIG8 SEQ ID N°7068 568120 569727
2337.1 CONTIG8 SEQ ID N°7068 572901 573461
2343.1 CONTIG8 SEQ ID N°7068 575955 578648
2345.1 CONTIG8 SEQ ED N°7068 578635 579891
2346.1 CONTIG8 SEQ ID N°7068 579888 581024
2348.1 CONTIG8 SEQ ED N°7068 581063 584191
2350.1 CONTIG8 SEQ ID N°7068 584584 586371
2357.1 CONTIG8 SEQ ID N°7068 590170 591033
2373.1 CONTIG8 SEQ ID N°7068 601883 602209
2385.1 CONTIG8 SEQ ID N°7068 613763 615529
2465:1 CONTIG12 SEQ ID N°7072 96957 97913
2466.1 CONTIG12 SEQ ID N°7072 97888 98478
2467.1 CONTIG12 SEQ ID N°7072 98499 99803
2468.1 CONTIG12 SEQ ED N°7072 99820 100638
2469.1 CONTIG12 SEQ ID N°7072 100647 101573
2470.1 CONTIG12 SEQ ID N°7072 101679 102866
2471.1 CONTIG12 SEQ ID N°7072 103108 104379
2472.1 CONTIG 12 SEQ ID N°7072 104380 105558
2473.1 CONTIG12 SEQ ID N°7072 105588 106370
2498.1 CONTIG12 SEQ ID N°7072 124135 124545
2500.1 CONTIG12 SEQ ED N°7072 123924 126110
2684.2 CONTIG8 SEQ ID N°7068 740173 740430
2710.1 CONTIG9 SEQ ED N°7069 1778548 1780281
2743.1 CONTIG9 SEQ ID N°7069 1810120 1810353
2769.1 CONTIG9 SEQ ID N°7069 1826652 1828562
2782.3 CONTIG8 SEQ ID N°7068 78469 79875
5631.2 CONTIG8 SEQ ID N°7068 78469 79875
2784.1 CONTIG8 SEQ ID N°7068 73057 73398
2889.1 CONTIG13 SEQ ED N°7073 19409 20020
2890.1 CONTIG 13 SEQ ID N°7073 18305 19162
2892.1 CONTIG13 SEQ ED N°7073 17653 17907
2894.1 CONTIG13 SEQ ID N°7073 15676 17652
2895.1 CONTIG13 SEQ ED N°7073 15195 15683
2896.1 CONTIG13 SEQ ED N°7073 14143 14952
3353.1 CONTIG13 SEQ ED N°7073 14143 14952
2915.1 CONTIG13 SEQ ED N°7073 21609 22910
3909.2 CONTIG 13 SEQ ED N°7073 21609 22910
2916.1 CONTIG13 SEQ ID N°7073 23001 24515
3908.2 CONTIG13 SEQ ED N°7073 23001 24515
2917.1 CONTIG13 SEQ ID N°7073 24645 24908
2918.1 CONTIG13 SEQ ID N°7073 24971 26290
2919.1 CONTIG13 SEQ ED N°7073 26453 26965
2920.1 CONTIG 13 SEQ ID N°7073 27050 27442
2921.1 CONTIG13 SEQ ED N°7073 27535 28215
3018.1 CONTIG13 SEQ ID N°7073 41565 42278
10565.1 CONTIG9 SEQ ID N°7069 1320956 1322047
1947.2 CONTIG9 SEQ ED N°7069 1320956 1322047
3019.1 CONTIG9 SEQ ID N°7069 1320956 1322047
3020.1 CONTIG9 SEQ ID N°7069 1320447 1320905
3021.1 CONTIG9 SEQ ID N°7069 1319365 1320159
3023.1 CONTIG9 SEQ ID N°7069 1317520 1319235
3024.1 CONTIG9 SEQ ID N°7069 1316653 1317507
3025.1 CONTIG9 SEQ ID N°7069 1315445 1316551
3027.1 CONTIG9 SEQ ED N°7069 1314906 1315190
3046.1 CONTIG9 SEQ ID N°7069 1298926 1299282
3047.1 CONTIG9 SEQ ID N°7069 1298615 1299712
3087.1 CONTIG9 SEQ ID N°7069 1016602 1017420
3097.1 CONTIG11 SEQ ID N°7071 183622 184182
3100.1 CONTIG11 SEQ ID N°7071 182447 182740
3101.1 CONTIG11 SEQ ID N°7071 182080 182409
3102.1 CONTIG11 SEQ ID N°7071 180595 181965
3103.1 CONTIG11 SEQ ED °7071 179123 180463
3104.1 CONTIG11 SEQ ID N°7071 177735 179093
3105.1 CONTIG11 SEQ ED °7071 176469 177605
3317.1 CONTIG9 SEQ ED N°7069 1863795 1864346
2685.2 CONTIG13 SEQ ID N°7073 12622 13683
3356.1 CONTIG13 SEQ ID N°7073 12622 13683
3357.1 CONTIG13 SEQ ID N°7073 11421 12638
3362.1 CONTIG13 SEQ ID N°7073 7035 7835
3364.1 CONTIG13 SEQ ID N°7073 5298 6497
3365.1 CONTIG13 SEQ ID N°7073 4728 5141 34.1 CONTIG9 SEQ ID N°7069 40333 43707 35.1 CONTIG9 SEQ ID N°7069 44063 44371 3538.1 CONTIG9 SEQ ID N°7069 1883947 1885137 3539.1 CONTIG9 SEQ ID N°7069 1885249 1885500 3541.1 CONTIG9 SEQ ID N°7069 1887373 1888815 3542.1 CONTIG9 SEQ ID N°7069 1888997 1889311 3544.1 CONTIG9 SEQ ID N°7069 1889663 1889962 3545.1 CONTIG9 SEQ ID N°7069 1890063 1890284 3548.1 CONTIG9 SEQ ID N°7069 1891102 1892148 3550.2 CONTIG9 SEQ ID N°7069 1893944 1895122 3551.1 CONTIG9 SEQ ID N°7069 1895356 1895775 3552.2 CONTIG9 SEQ ID N°7069 1895768 1896094 3683.1 CONTIG8 SEQ ID N°7068 129540 131426 37.1 CONTIG9 SEQ ED N°7069 44561 44827 3740.2 CONTIG9 SEQ ED N°7069 1025799 1027007 2327.1 CONTIG12 SEQ ED N°7072 10 987 3742.1 CONTIG 12 SEQ ED N°7072 10 987 1944.1 CONTIG13 SEQ ID N°7073 999 1205 3793.1 CONTIG9 SEQ ID N°7069 1491228 1491776 3794.1 CONTIG9 SEQ ID N°7069 1490472 1491212 3795.4 CONTIG9 SEQ ID N°7069 1489860 1490459 3797.2 CONTIG9 SEQ ED N°7069 1921869 1923518 3798.1 CONTIG9 SEQ ID N°7069 1923682 1924194
3799.1 CONTIG9 SEQ ID N°7069 1924210 1924524
3800.1 CONTIG9 SEQ ID N°7069 1924550 1926376
3803.1 CONTIG9 SEQ ID N°7069 1926518 1927483
3827.1 CONTIG 13 SEQ ED N°7073 2622 3332
3829.2 CONTIG 13 SEQ ID N°7073 3307 4557
3847.1 CONTIG9 SEQ ID N°7069 1482616 1483233
3848.1 CONTIG9 SEQ ID N°7069 1481860 1482336
3849.3 CONTIG9 SEQ ID N°7069 1480223 1481680
3850.1 CONTIG9 SEQ ID N°7069 1479788 1480216
3890.1 CONTIG 13 SEQ ID N°7073 20841 21641 39.1 CONTIG9 SEQ ID N°7069 44906 45655
8044.1 CONTIG7 SEQ ID N°7067 16618 17850
3986.2 CONTIG5 SEQ ID N°7065 2 2959
4124.2 CONTIG5 SEQ ID N°7065 2 2959
3997.1 CONTIG9 SEQ ID N°7069 1920982 1921893 40.1 CONTIG9 SEQ ID N°7069 45836 46858
4018.1 CONTIG9 SEQ ED N°7069 1913464 1914630
4019.1 CONTIG9 SEQ ED N°7069 1912844 1913476
4022.2 CONTIG9 SEQ ED N°7069 1911062 1911376
4064.1 CONTIG9 SEQ ED N°7069 1486212 1486631
4065.1 CONTIG9 SEQ ED N°7069 1485967 1486812
4066.1 CONTIG9 SEQ ID N°7069 1486860 1487468
4067.4 CONTIG9 SEQ ED N°7069 1487626 1488717 41.1 CONTIG9 SEQ ID N°7069 46950 47837 43.1 CONTIG9 SEQ ED N°7069 48387 50369
4936.1 CONTIG9 SEQ ID N°7069 1605170 1605475 504.1 CONTIG9 SEQ ED N°7069 473249 474757 506.1 CONTIG9 SEQ ED N°7069 475061 476458 507.1 CONTIG9 SEQ ID N°7069 476469 477350 508.1 CONTIG9 SEQ ID N°7069 477545 477874 518.1 CONTIG9 SEQ ID N°7069 485931 487241 519.1 CONTIG9 SEQ ID N°7069 487186 487464 521.1 CONTIG9 SEQ ID N°7069 487597 487989 669.1 CONTIG9 SEQ ID N°7069 604354 605595
6736.1 CONTIG6 SEQ ID N°7066 1824 2537
6874.1 CONTIG8 SEQ ID N°7068 575999 576325 750.1 CONTIG9 SEQ ID N°7069 1271952 1274519 751.1 CONTIG9 SEQ ID N°7069 1271377 1271823 752.1 CONTIG9 SEQ ID N°7069 1270905 1271255 760.1 CONTIG9 SEQ ID N°7069 1266574 1267239 761.1 CONTIG9 SEQ ID N°7069 1266303 1266707 764.1 CONTIG9 SEQ ID N°7069 1263381 1263866 765.1 CONTIG9 SEQ ID N°7069 1261771 1262709 774.1 CONTIG9 SEQ ED N°7069 1258317 1260068 778.1 CONTIG9 SEQ ID N°7069 1256902 1257267
8067.2 CONTIG9 SEQ ED N°7069 1488930 1489379
8073.2 CONTIG9 SEQ ED N°7069 1484592 1484990
10294.1 CONTIG8 SEQ ID N°7068 119556 120962
7817.1 CONTIG8 SEQ ID N°7068 119556 120962
8134.1 CONTIG8 SEQ ID N°7068 119556 120962
874.1 CONTIG9 SEQ ED N°7069 1177144 1178064 875.1 CONTIG9 SEQ ID N°7069 1176375 1176995 876.1 CONTIG9 SEQ ED N°7069 1175483 1176361 1769.1 CONTIG3 SEQ ED N°7063 21 716 9388.1 CONTIG3 SEQ ID N°7063 21 716 4934.1 CONTIG9 SEQ ED N°7069 1819527 1820453 2798.2 CONTIG8 SEQ ED N°7068 71193 77576 3630.3 CONTIG 12 SEQ ID N°7072 11054 16705
Table XII: Position of former cc mtigs of the Paris strain on the genomi c sequence o chromosome of the Paris strain of sequence SEQ ID 3507 and of the plasmid of the Paris strain of sequence SEQ ID 3508 posl pos2 former contig SEQ ID of former a chromosome 1 44600 41 SEQ ID Ncχ41 44600 161000 56 SEQ ED N oo56 162000 223000 54 SEQ ID N oo54 223000 232000 39 SEQ ED Noo39 236000 424000 49 SEQ ED No49 437000 469000 37 SEQ ED N0037 464000 758000 53 SEQ ID N oo53 758000 781000 45 SEQ LD Noc45 789000 879000 43 SEQ ID Noc43 883000 901000 36 SEQ ID Noo36 898000 986000 42 SEQ ID No<42 990000 1160000 46 SEQ JD NOC46 1160000 1214000 39 SEQ ID Noo39 1214000 1352000 54 SEQ ID N oo54 1352000 1670000 52 SEQ ID Noo52 1670000 1736000 40 SEQ ID Noβrø 1736000 2040000 55 SEQ ID N oo55 2044000 2093000 56 SEQ ID Noo56 2093000 2204722 45 SEQ ID No45 2204000 2298000 50 SEQ ID Noo50 2298000 2656000 56 SEQ ID Noo56 2656000 2740000 50 SEQ ID No δO 2740000 2753600 33 SEQ ID N0033 2753600 2954000 47 SEQ ID Noc47 2954000 3178000 51 SEQ ID Noo51 3178000 3289000 44 SEQ ID No44 3289000 3449000 48 SEQ JD NOC48 3449000 3463000 34 SEQ ID Noo34 3463000 3503610 41 SEQ ED Nc41 plasmid 1 131900 55(positionl to 132400) SEQ ED N oo55
Table XIII: Correspondence of the numbers attributed to the chromosome and au plasmid of the Paris and Lens strain with the SEQ ID numbers identified in the list of sequences
SeqID=3507 chromosome of the Paris strain
SeqID=3508 plasmid of the Paris strain
SeqID=6733 chromosome of the Lens strain
SeqID=6734 plasmid of the Lens strain
Table XEV: Correspondence of the numbers attributed to the genes of the Paris strain on its chromosome of sequence SEQ ID 3507 and on its plasmid of sequence SEQ ED 3508 with the numbers of the SEQ ID identified in the list of sequences and position of the nucleic sequences coding these genes on the sequence of the chromosome and of the plasmid with their putative function
Table XV: Nature of the class listed in the "Class" column in Tables XIV and XVI
1. Cellular envelope and cellular processes 1.1 Cellular wall and external membrane
1.2 Proteins of transport/bond and lipoproteins
1.3 Sensors (transduction of signal)
1.4 Bioenergy of membrane
1.5 Mobility and chimiotaxia 1.6 Secretion of protein
1.7 Cellular division
1.8 Structures of cellular surface and pili
2. Intermediary metabolism
2.1 Metabolism of glucides and related molecules
2.1.1 Specific ways
2.1.2 Principal glycolytic ways
2.1.3 TCA cycle 2.2 Metabolism of aminoacids and related molecules
2.3 Metabolism of nucleotides and nucleic acids
2.4 Metabolism of lipids
2.5 Metabolism of coenzymes and prosthetic groups
2.6 Metabolism of phosphate
3. Information paths
3.1 Replication of DNA
3.2 Repair and restriction/modification of DNA
3.3 Recombination of DNA
3.4 Segregation and encapsidation of DNA
3.5 Synthesis of RNA
3.5.1 Initiation
3.5.2 Regulation
3.5.3 Elongation
3.5.4 Termination
3.6 Modification of RNA
3.7 Synthesis of protein
3.7.1 Ribosomal proteins
3.7.2 Synthetases of aminoacyl-tRNA
3.7.3 Initiation
3.7.4 Elongation
3.7.5 Termination
3.8 Modification of protein
3.9 Folding of protein
4. Other functions
4.1 Adaptation to atypical conditions
4.2 Detoxification
4.3 Toxins
4.4 Functions relating to phage
4.5 Transposon, IS, Plasmid
4.6 Various
5. Similar to unknown proteins
5.1 Of Legionella (similar though not the si
5.2 Of other organisms
6. No similarity
Similar to the enzyme IIN of the PTS XXXX-specific system Similar to the transcriptional regulator (family xx)
Similar to the transportor ABC (protein of ATP bond) Similar to the transportor ABC (permease) Similar to the transportor ABC (bond protein) Similar to the response regulator with two compounds Similar to the histidine kinase sensor with two compounds
Protein bound to putative peptidoglycane (LPXTG pattern)
Table XVI: Correspondence of the numbers attributed to the specific genes of the Lens strain relative to the Paris and Philadelphia strains on its chromosome of sequence SEQ ID 6733 and on its plasmid of sequence SEQ ID 6734 with the numbers of SEQ ID identified in the list of sequences and position of the nucleic sequences coding these genes on the sequence of the chromosome and the plasmid with their putative function
Table XVII: List of the specific sequences of the Paris strain relative to the Lens and Philadelphia strains with their Pasteur Institute « ORF » correspondence number and accession number in the gene banks
1056.1 SEQID=3544 EMBL_NAME=lppl800 1067.1 SEQID=3551 EMBL_NAME=lppl877 1069.2 SEQID=3552 EMBL_NAME=lppl878 1076.3 SEQID=6591 EMBL_NAME=plpp0105 1077.1 SEQID=6592 EMBL_NAME=plpp0104 1078.2 SEQID=6593 EMBL_NAME=plpp0103 1080.2 SEQID=3558 EMBL_NAME=lpp3012 1081.2 SEQID=3559 EMBL_NAME=lpp3011 11.2 SEQID=3573 EMBL_NAME=lpp0196 114.2 SEQID=3598 EMBL_NAME=lpp2957 115.2 SEQID=3603 EMBL_NAME=lpp2956 116.1 SEQED=3609 EMBL_NAME=lpp2955 1160.3 SEQID=3610 EMBL_NAME=lpp0125 1171.4 SEQID=6594 EMBL_NAME=plpp0017 1172.2 SEQED=6595 EMBL_NAME=plpp0018 118.1 SEQED=3618 EMBL_NAME=lpp2954 1183.4 SEQID=3621 EMBL_NAME=lpp0356 1213.2 SEQED=3638 EMBL_NAME=lpp0257 1235.3 SEQED=6598 EMBL_NAME=ρlρρ0121 1237.2 SEQED=6600 EMBL_NAME=plpp0119 1299.3 SEQID=6601 EMBL_NAME=plpp0036 13.1 SEQID=3688 EMBL_NAME=lρρ0195 1342.3 SEQID=6602 EMBL_NAME=plpp0034 1344.4 SEQID=6603 EMBL_NAME=plpp0033 1362.3 SEQID=6604 EMBL_NAME=plpp0098 1364.2 SEQID=6605 EMBL_NAME=plpp0099 1372.2 SEQID=3726 EMBL_NAME=lpp2385 1373.1 SEQID=3727 EMBL_NAME=lpp2384 1375.2 SEQID=3728 EMBL_NAME=lpp2383 1376.2 SEQID=3729 EMBL_NAME=lpp2382 1387.2 SEQED=3735 EMBL_NAME=lpp0079 1388.2 SEQID=3736 EMBL_NAME=lpp0080 139.6 SEQID=3737 EMBL_NAME=lppl l00 1392.2 SEQID=3740 EMBL_NAME=lppl097 1394.2 SEQID-3741 EMBL_NAME=lpp2557 1429.4 SEQID=3766 EMBL_NAME=lpp2442 1522.2 SEQID=6606 EMBL_NAME=plpp0127 1523.2 SEQID=6607 EMBL_NAME=plpp0128 1524.3 SEQED=6608 EMBL_NAME=plpp0129 1566.3 SEQID=3846 EMBL_NAME=lpp2490 1570.4 SEQID=3850 EMBL_NAME=lpp0077 1599.5 SEQID=6610 EMBL_NAME=plpp0039 1623.4 SEQID=3881 EMBL_NAME=lpp2394 1624.5 SEQID=3882 EMBL_NAME=lpp2395
163.1 SEQID=3887 EMBL_NAME=lpp2344
1655.2 SEQID=3905 EMBL_NAME=lppl895
1683.2 SEQID=3923 EMBL_NAME=lpp0046 172.1 SEQID=3946 EMBL_NAME=lpp2978
1735.1 SEQID=3955 EMBL_NAME=lppl862 1761.4 SEQID=6611 EMBL_NAME=plpp0074
1779.3 SEQID=3985 EMBL_NAME=lpp2040
1787.4 SEQID=3992 EMBL_NAME=lppl824 18.1 SEQID=4000 EMBL_NAME=lpp0192
1803.2 SEQID=4003 EMBL_NAME=lpp2374 1815.2 SEQID=4010 EMBL_NAME=lpp2986
1848.5 SEQID=6613 EMBL_NAME=plpp0124 1849.4 SEQID=6614 EMBL_NAME=plpp0125 1852.2 SEQID=6615 EMBL_NAME=plpp0126
1891.2 SEQID=4056 EMBL_NAME=lpp3007 19.1 SEQID=4061 EMBL_NAME=lpρ0191
1920.3 SEQID=4074 EMBL_NAME=lpp2405 1923.2 SEQID=4075 EMBL_NAME=lpp2406
1924.4 SEQED=4076 EMBL_NAME=lpp2407 2.1 SEQID=4112 EMBL_NAME=lpp0163 20.1 SEQID=4113 EMBL_NAME=lpp0190 2006.2 SEQED=6616 EMBL_NAME=plpp0032 2027.2 SEQID=4125 EMBL_NAME=lppl910 2030.2 SEQED=4127 EMBL_NAME=lpp0241 2051.2 SEQED=6617 EMBL_NAME=plpp0035 2078.2 SEQED=4152 EMBL_NAME=lpp2456
2112.2 SEQID=4167 EMBL_NAME=lpp3006
2153.3 SEQID=4196 EMBL_NAME=lpρ2427 2172.2 SEQID=6619 EMBL_NAME=plpp0107
2173.2 SEQID=6620 EMBL_NAME=plpp0106
2174.1 SEQID=6621 EMBL_NAME=plpp0102
2175.3 SEQID=6622 EMBL_NAME=plpp0101
2176.3 SEQID=6623 EMBL_NAME=plpp0139
2177.2 SEQE>=6624 EMBL_NAME=plpp0140 220.1 SEQID=4215 EMBL_NAME=lpp2859 221.1 SEQED=4220 EMBL_NAME=lpp2860
2212.4 SEQED=6626 EMBL_NAME=plpp0038 222.1 SEQID=4223 EMBL_NAME=lpp2861 223.1 SEQID=4227 EMBL_NAME=lpp2862
2239.3 SEQID=4233 EMBL_NAME=lppl330 2244.2 SEQID=6627 EMBL_NAME=plpp0052 2245.2 SEQID=6628 EMBL_NAME=plpp0053 2247.7 SEQID=6629 EMBL_NAME=plpp0054 2258.2 SEQID=6630 EMBL_NAME=plpp0130 2260.2 SEQID=6631 EMBL_NAME=plpp0131 2270.2 SEQID=4248 EMBL_NAME=lpp0082
2271.1 SEQID=4249 EMBL_NAME=lpp0081
2351.2 SEQID=4293 EMBL_NAME=lpp3016 2367.2 SEQID=4306 EMBL_NAME=lpp2386 2374.2 SEQID=4312 EMBL_NAME=lpp2987
24.1 SEQED=4325 EMBL_NAME=lρp0189
2414.2 SEQED=4338 EMBL_NAME=lpp2879
2428.2 SEQED=4349 EMBL_NAME=lppl870
2438.2 SEQED=4356 EMBL_NAME=lpp0074
2439.2 SEQID=4357 EMBL_NAME=lpp0073
2441.2 SEQID=4359 EMBL_NAME=lpp0072
2461.2 SEQID=4373 EMBL_NAME=lpp0769
2462.2 SEQID=4374 EMBL_NAME=lpp0770
2464.2 SEQID=4375 EMBL_NAME=lpp0771
2483.2 SEQID=4389 EMBL_NAME=lpp0052
2488.2 SEQID=4392 EMBL_NAME=lpp0336
2489.2 SEQID=4393 EMBL_NAME=lpp0337
2490.2 SEQED=4395 EMBL_NAME=lpp0338 25.1 SEQID=4399 EMBL_NAME=lpp0188 254.2 SEQID=6633 EMBL_NAME=plpp0028 255.1 SEQID=6634 EMBL_NAME=plpp0029 2555.5 SEQID=4432 EMBL_NAME=lpp2147
256.1 SEQID=6635 EMBL_NAME=plpp0030
2573.3 SEQID=6636 EMBL_NAME=plpp0094
258.2 SEQID=6637 EMBL_NAME=plpp0031 26.1 SEQID=4457 EMBL_NAME=lpp0187 2605.3 SEQID=4461 EMBL_NAME=lpp0198 2616.1 SEQID=4464 EMBL_NAME=lpp0201
2622.1 SEQID=4468 EMBL_NAME=lpp0197
2625.2 SEQID=4469 EMBL_NAME=lpp2376 2626.1 SEQID=4470 EMBL_NAME=lpp2375
2659.3 SEQID=6638 EMBL_NAME=plpp0022
2660.1 SEQID=6639 EMBL_NAME=plpp0023
2661.2 SEQID=6640 EMBL_NAME=plpp0024 2662.2 SEQID-6641 EMBL_NAME=plpp0025 2665.1 SEQID=6642 EMBL_NAME=plpp0040
2666.1 SEQID=6643 EMBL_NAME=plpp0041
2667.2 SEQID=6644 EMBL_NAME=plpp0042 27.1 SEQID=4506 EMBL_NAME=lpp0186
2712.1 SEQID=4515 EMBL_NAME=lpp2620
2713.2 SEQID=4516 EMBL_NAME=lpp2621
2726.1 SEQID=4522 EMBL_NAME=lpp2636
2727.2 SEQID=4523 EMBL_NAME=lpp0256
2767.1 SEQID=4547 EMBL_NAME=lppl561
2815.2 SEQED=4575 EMBL_NAME=lpp3070
2830.3 SEQID=4586 EMBL_NAME=lppl928 2873.1 SEQED=6645 EMBL_NAME=plpp0109
2874.1 SEQED=6646 EMBL_NAME=plpp0108
2877.2 SEQID=6647 EMBL_NAME=plpp0100
2878.2 SEQID=6648 EMBL_NAME=plpp0097
2879.4 SEQID=6649 EMBL_NAME=plpp0096
2881.3 SEQED=6650 EMBL_NAME=plpp0095 29.1 SEQID=4611 EMBL_NAME=lpp0185 2919.1 SEQED=4624 EMBL_NAME=lpp0144 2932.3 SEQID=4633 EMBL_NAME=lpp2880
2951.2 SEQID=4643 EMBL_NAME=lpp2139
2972.3 SEQID=4663 EMBL_NAME=lpp2122 2973.2 SEQID=4664 EMBL_NAME=lpp2121 2975.2 SEQID=4665 EMBL_NAME=lpp2120 2976.1 SEQID=4666 EMBL_NAME=lpp2119 2978.1 SEQID=4667 EMBL_NAME=lpp2118 2982.1 SEQID=4668 EMBL_NAME=lpp2117 2983.1 SEQID=4669 EMBL_NAME=lpp2116 2985.1 SEQID=4670 EMBL_NAME=lpp2115 2986.1 SEQID=4671 EMBL_NAME=lpp2114 2987.1 SEQID=4672 EMBL_NAME=lpp2113 2988.1 SEQID=4673 EMBL_NAME=lpp2112
2990.1 SEQID=4674 EMBL_NAME=lpp2111 3.1 SEQID=4683 EMBL_NAME=lpp0162 30.1 SEQID=4684 EMBL_NAME=lpp0184
3005.2 SEQID=4689 EMBL_NAME=lpp2983 3009.2 SEQID=4691 EMBL_NAME=lpp2968 3011.2 SEQID=4692 EMBL_NAME=lpp0587 3016.1 SEQID=4696 EMBL_NAME=lpp0583 3017.1 SEQID=4697 EMBL_NAME=lpp0582
3023.1 SEQID=4700 EMBL_NAME=lpp0576
3030.2 SEQID=6651 EMBL_NAME=plpp0132 3031.2 SEQID=6652 EMBL_NAME=plpp0133 3035.2 SEQLD=6653 EMBL_NAME=plpp0134 3036.2 SEQED=6654 EMBL_NAME=plpp0138
3101.1 SEQED=4743 EMBL_NAME=lpp0303
3109.2 SEQID=4748 EMBL_NAME=lpp0297 3110.1 SEQID=4749 EMBL_NAME=lpp0296
3113.1 SEQID=4750 EMBL_NAME=lpp0295
3114.2 SEQID=4751 EMBL_NAME=lpp0294 3116.2 SEQID=4752 EMBL_NAME=lpp0293 3134.1 SEQID=6655 EMBL_NAME=plpp0072 3135.1 SEQID=6656 EMBL_NAME=plpp0071 3136.1 SEQ1D=6657 EMBL_NAME=plpp0065
3137.1 SEQID=6658 EMBL_NAME=plpp0064
3138.2 SEQED=6659 EMBL_NAME=plpp0063 3231.1 SEQED=4815 EMBL_NAME=lpp2319 3232.1 SEQED=4816 EMBL_NAME=lpp2318
3233.1 SEQID=4817 EMBL_NAME=lpp2317
3234.3 SEQID=4818 EMBL_NAME=lpp2316
3257.2 SEQID=4835 EMBL_NAME=lpp2409
3258.2 SEQID=4836 EMBL_NAME=lpp2408
3299.4 SEQID=4860 EMBL_NAME=lpp2179 3338.1 SEQID=4884 EMBL_NAME=lpp2484 3343.1 SEQID=4887 EMBL_NAME=lpp2486
3348.3 SEQID=6662 EMBL_NAME=plpp0078 3349.3 SEQID=6663 EMBL_NAME=plpp0079 3350.3 SEQID=6664 EMBL_NAME=plpp0080 3351.3 SEQID=6665 EMBL_NAME=plpp0081 3352.1 SEQID=6666 EMBL_NAME=plpp0082
3353.1 SEQED=6667 EMBL_NAME=plpp0083
3354.1 SEQLD=6668 EMBL_NAME=plpρ0084
3355.2 SEQED=6669 EMBL_NAME=plpp0089 3356.2 SEQED=6670 EMBL_NAME=plpp0090 3357.2 SEQID=6671 EMBL_NAME=plpp0091
3394.1 SEQID=4910 EMBL_NAME=lppl041 34.1 SEQID-4916 EMBL_NAME=lpp0182
3406.2 SEQID=4920 EMBL_NAME=lppl050
3411.2 SEQID=4922 EMBL_NAME=lpp2059 3466.1 SEQID=4957 EMBL_NAME=lpp2864 3486.1 SEQID=4969 EMBL_NAME=lppl l lO 3498.1 SEQED=6672 EMBL_NAME=plpp0019 3530.1 SEQED=4998 EMBL_NAME=lppl823 3532.1 SEQID=4999 EMBL_NAME=lppl822 3584.1 SEQID=5034 EMBL_NAME=lppl098 3631.4 SEQID=6675 EMBL_NAME=plpp0058
3632.3 SEQID=6676 EMBL_NAME=plpp0051
3653.3 SEQID=5083 EMBL_NAME=lpp2412 3655.1 SEQID=5084 EMBL_NAME=lpp2413
3658.1 SEQID=5086 EMBL_NAME=lpp2420
3659.2 SEQID=5087 EMBL_NAME=lpp2424
3661.4 SEQID=5089 EMBL_NAME=lpp2425 3664.1 SEQID=5091 EMBL_NAME=lpp3018
3665.1 SEQID=5092 EMBL_NAME=lpp3019
3666.2 SEQID=5093 EMBL_NAME=lpp3020 3729.1 SEQID=5130 EMBL_NAME=lpp0204 3730.1 SEQID=5131 EMBL_NAME=lpp0205 3731.1 SEQED=5132 EMBL_NAME=lpp0206
3732.1 SEQID=5133 EMBL_NAME=lpp0207
3735.2 SEQID=5135 EMBL_NAME=lpp0209
3764.3 SEQID=5153 EMBL_NAME=lppl868 3851.1 SEQID=5203 EMBL_NAME=lppl492 3887.1 SEQED=5217 EMBL_NAME=lpp2439 3888.1 SEQED=5218 EMBL_NAME=lρρ2438
3889.1 SEQID=5219 EMBL_NAME=lpp2437
3890.2 SEQJD=5221 EMBL_NAME=lpp2436 3891.2 SEQID=5222 EMBL_NAME=lpp2435 399.2 SEQID=5282 EMBL_NAME=lppl560 3993.2 SEQID=5284 EMBL_NAME=lppl640 4.1 SEQED=5288 EMBL_NAME=lpp0161
400.1 SEQID=5289 EMBL_NAME=lppl559
401.2 SEQID=5296 EMBL_NAME=lppl558 4037.2 SEQID=5314 EMBL_NAME=lpp0065 4096.1 SEQID=5355 EMBL_NAME=lpp0772 4097.1 SEQID=5356 EMBL_NAME=lpp0773 4098.1 SEQED=5357 EMBL_NAME=lpp0774
4101.1 SEQED=5359 EMBL_NAME=lpp0775
4104.2 SEQED=5360 EMBL_NAME=lpp2390 4105.1 SEQID=5361 EMBL_NAME=lpp2389 4106.1 SEQID=5362 EMBL_NAME=lpp2388
4107.1 SEQID=5363 EMBL_NAME=lpp2387
4108.1 SEQID=5364 EMBL_NAME=lpp2381
4109.2 SEQID=5365 EMBL_NAME=lpp2380
4123.2 SEQID=5374 EMBL_NAME=lppl077 4127.1 SEQID=5376 EMBL_NAME=lppl075 4128.1 SEQID=5377 EMBL_NAME=lppl074
4129.1 SEQED=5378 EMBL_NAME=lppl073 413.5 SEQID=5379 EMBL_NAME=lppl905
4130.3 SEQED=5380 EMBL_NAME=lppl072
4144.3 SEQID=5393 EMBL_NAME=lpp0412
415.2 SEQED=5397 EMBL_NAME=lppl906
4179.2 SEQED=5420 EMBL_NAME=lppl615 4209.2 SEQID=5437 EMBL_NAME=lppl566
4233.4 SEQID=5452 EMBL_NAME=lpp2396 4234.4 SEQID=5453 EMBL_NAME=lpp2397 4238.1 SEQID=5455 EMBL_NAME=lpp2399 4239.4 SEQID=5456 EMBL_NAME=lpp2400 4244.1 SEQID=5460 EMBL_NAME=lppl407
4248.1 SEQED=5463 EMBL_NAME=lpp3010
425.1 SEQID=5465 EMBL_NAME=lpp2430
4251.2 SEQID=5466 EMBL_NAME=lpp3008
426.3 SEQID=5470 EMBL_NAME=lpp2428
4281.2 SEQED=5481 EMBL_NAME=lpp2047
4282.3 SEQED=5482 EMBL_NAME=lpp2046 4284.3 SEQID=5483 EMBL_NAME=lpp2045
4285.1 SEQID=5484 EMBL_NAME=lpp2044
4421.2 SEQID=5579 EMBL_NAME=lpp0211 4422.1 SEQID=5580 EMBL_NAME=lpp0212
4424.1 SEQID=5581 EMBL_NAME=lpp0213
4479.2 SEQED=5614 EMBL_NAME=lppl573 4481.1 SEQID=6677 EMBL_NAME=plpp0050
4482.1 SEQID=6678 EMBL_NAME=plpp0049
4483.2 SEQLD=6679 EMBL_NAME-plpp0048
4490.2 SEQID=5622 EMBL_NAME=lpp0068
4491.3 SEQID=5623 EMBL_NAME=lpp0069 4492.3 SEQID=5624 EMBL_NAME=lpp0070
4493.1 SEQID=5625 EMBL_NAME=lpp0071
4496.2 SEQID=5626 EMBL_NAME=lpp0075
4500.1 SEQID=5631 EMBL_NAME=lpp2085
4516.3 SEQID=5639 EMBL_NAME=lpp0078
4533.2 SEQID=5654 EMBL_NAME=lpp0779
4546.3 SEQID=5661 EMBL_NAME=lpp2967 4549.2 SEQID=6680 EMBL_NAME=plpp0118 4553.2 SEQID=5665 EMBL_NAME=lpp0291
4636.4 SEQID=5715 EMBL_NAME=lpp2168 4660.2 SEQID=5733 EMBL_NAME=lppl956
474.2 SEQED=5787 EMBL_NAME=lpp0777 475.1 SEQID=5794 EMBL_NAME=lpp0778 4770.1 SEQID=5809 EMBL_NAME=lpp0439 4785.1 SEQID=5820 EMBL_NAME=lppl625
2 131
4822.3 SEQID=5848 EMBL_NAME=lpp2393
485.3 SEQID=6681 EMBL_NAME=plρp0026
486.2 SEQID=6682 EMBL_NAME=plpp0027
4860.1 SEQID=5865 EMBL_NAME=lpp0983 489.1 SEQID=5886 EMBL_NAME=lpp0456
4913.2 SEQID=5896 EMBL_NAME=lppl054
4914.3 SEQID=5897 EMBL_NAME=lppl053
4927.1 SEQID=5905 EMBL_NAME=lppl904
4945.2 SEQID=6683 EMBL_NAME=plpp0003 4947.2 SEQID=6684 EMBL_NAME=plpp0002
502.3 SEQID=5956 EMBL_NAME=lppl l22 5037.2 SEQED=5967 EMBL_NAME=lpp3077 505.3 SEQID=5976 EMBL_NAME=lpp2418 507.3 SEQID=5992 EMBL_NAME=lpp2416 5078.2 SEQID=6685 EMBL_NAME=plpp0004 5081.5 SEQID=6687 EMBL_NAME=plpp0005
5088.2 SEQED=6000 EMBL_NAME=lppl576
509.1 SEQED=6001 EMBL_NAME=lpp2415
510.2 SEQID=6008 EMBL_NAME=lpp2414
5113.3 SEQID=6689 EMBL_NAME=plpp0061 5114.3 SEQID=6690 EMBL_NAME=plpp0062 5146.2 SEQED=6022 EMBL_NAME=lpp0292 5151.1 SEQED=6692 EMBL_NAME=plpp0123 5152.1 SEQID=6693 EMBL_NAME=plpp0122
5153.1 SEQID=6694 EMBL_NAME=plpp0093
5154.2 SEQID=6695 EMBL_NAME=plpp0092 52.1 SEQID=6035 EMBL_NAME=lpp0168 5201.2 SEQID=6038 EMBL_NAME=lppl219 5202.2 SEQID=6039 EMBL_NAME=lppl220 5204.2 SEQID=6040 EMBL_NAME=lppl221
5208.1 SEQID=6042 EMBL_NAME=lpp2984
5224.2 SEQED=6045 EMBL_NAME=lppl929
5242.3 SEQID=6054 EMBL_NAME=lpρ2125
5288.1 SEQID=6069 EMBL_NAME=lpp0588
5297.2 SEQID=6073 EMBL_NAME=lpp2492
5307.3 SEQID=6075 EMBL_NAME=lppl682 5321.2 SEQED=6086 EMBL_NAME=lppl052 5322.2 SEQED=6087 EMBL_NAME=lppl051 5387.1 SEQID=6109 EMBL_NAME=lpp0083
5390.1 SEQID=6112 EMBL_NAME=lpp0089
5474.2 SEQID=6145 EMBL_NAME=lpp2315
5517.1 SEQID=6167 EMBL_NAME=lpp2391
5554.3 SEQID=6187 EMBL_NAME=lppl216 5555.3 SEQID=6188 EMBL_NAME=lppl217 5576.3 SEQID=6197 EMBL_NAME=lpp2401
5605.2 SEQID=6211 EMBL_NAME=lpp0076
5621.1 SEQID=6219 EMBL_NAME=lpp2433
5623.2 SEQED=6220 EMBL_NAME=lpp2434
5626.3 SEQID=6221 EMBL_NAME=lpp2404 5634.2 SEQID=6224 EMBL_NAME=lpp0432
5660.1 SEQED=6239 EMBL_NAME=lppl577 567.6 SEQID=6696 EMBL_NAME=plpp0037
5675.2 SEQID=6243 EMBL_NAME=lpp2123 5677.2 SEQID=6244 EMBL_NAME=lpp2124 5688.2 SEQID=6249 EMBL_NAME=lpp2587 577.2 SEQID=6270 EMBL_NAME=lpp0203 5812.1 SEQED=6278 EMBL_NAME=lpp2377 5813.1 SEQID=6279 EMBL_NAME=lpp2378 5814.1 SEQID=6280 EMBL_NAME=lpp2379 5822.1 SEQID=6282 EMBL_NAME=lpp2863
5844.1 SEQED=6697 EMBL_NAME=plpp0047
5846.2 SEQID=6698 EMBL_NAME=plpp0057 5874.2 SEQID=6289 EMBL_NAME=lppl563
5890.1 SEQID=6292 EMBL_NAME=lpp2411 59.1 SEQID=6298 EMBL_NAME=lpp0164
591.4 SEQID=6304 EMBL_NAME=lpp2885
5987.2 SEQID=6315 EMBL_NAME=lpp2403 6015.1 SEQID=6699 EMBL_NAME=plpp0060 6016.1 SEQED=6700 EMBL_NAME=plpp0059 6029.1 SEQID=6321 EMBL_NAME=lpp0543 6052.1 SEQID=6326 EMBL_NAME=lppl218 6071.1 SEQID=6329 EMBL_NAME=lpp0290
6072.1 SEQID=6330 EMBL_NAME=lpp0289
6117.2 SEQID=6335 EMBL_NAME=lpp0202
616.5 SEQID=6340 EMBL_NAME=lpp2884
6165.1 SEQID=6342 EMBL_NAME=lpp2245a
6217.2 SEQID=6352 EMBL_NAME=lppl564
6224.3 SEQID=6701 EMBL_NAME=plpp0021 6227.1 SEQID=6702 EMBL_NAME=plpp0008 6229.1 SEQID=6703 EMBL_NAME=plpp0015 6230.1 SEQID=6704 EMBL_NAME=plpp0016 6231.3 SEQID=6705 EMBL_NAME=plpp0020 6263.1 SEQID=6357 EMBL_NAME=lpp0298 6273.1 SEQED=6359 EMBL_NAME=lpp3045 6314.1 SEQID=6365 EMBL_NAME=lppl908 6429.1 SEQID=6706 EMBL_NAME=plpp0141a
659.2 SEQED=6707 EMBL_NAME=plpp0135
660.1 SEQID=6708 EMBL_NAME=plpp0136
661.3 SEQID=6709 EMBL_NAME=plpp0137 678.3 SEQID=6398 EMBL_NAME=lppl081
680.3 SEQID=6399 EMBL_NAME=lppl080
681.4 SEQID=6400 EMBL_NAME=lppl079 682.3 SEQID=6401 EMBL_NAME=lppl078 7.1 SEQID=6412 EMBL_NAME=lpp0160
713.2 SEQID=6710 EMBL_NAME=plpp0046 715.1 SEQID=6711 EMBL_NAME=plpp0045
716.1 SEQED=6712 EMBL_NAME=plpp0044
717.3 SEQID=6713 EMBL_NAME=plpp0043
737.2 SEQED=6714 EMBL_NAME=plpp0141b
739.5 SEQID=6715 EMBL_NAME=plpp0001
750.2 SEQ1D=6437 EMBL_NAME=lpp2613 752.2 SEQID=6438 EMBL_NAME=lpp2614 761.2 SEQED=6716 EMBL_NAME=plpp0110 762.2 SEQID=6717 EMBL_NAME=plpp0111 764.1 SEQID=6718 EMBL_NAME=plpp0112 765.1 SEQID=6719 EMBL_NAME=plpp0113 766.4 SEQID=6720 EMBL_NAME=plpp0114 783.2 SEQID=6721 EMBL_NAME=plpp0055 785.4 SEQID=6722 EMBL_NAME=plpp0056 789.4 SEQID=6456 EMBL_NAME=lpp2245b 796.2 SEQID=6723 EMBL_NAME=plpp0068 862.1 SEQID=6496 EMBL_NAME=lpp2057 863.1 SEQID=6497 EMBL_NAME=lpp2056 875.5 SEQID=6505 EMBL_NAME=lpp2314 876.1 SEQID=6506 EMBL_NAME=lpp2313 879.3 SEQID=6507 EMBL_NAME=lpp2312 884.3 SEQID=6726 EMBL_NAME=plpp0117 885.3 SEQID=6727 EMBL_NAME=plpp0116 886.4 SEQID=6728 EMBL_NAME=plpp0115 905.2 SEQED=6527 EMBL_NAME=lpp2473 946.3 SEQID=6555 EMBL_NAME=lpp2423 947.3 SEQID=6556 EMBL_NAME=lpp2422 948.3 SEQID=6557 EMBL_NAME=lpp2421 956.2 SEQID=6729 EMBL_NAME=plpp0088 957.2 SEQID=6730 EMBL_NAME=plpp0087 959.2 SEQID=6731 EMBL_NAME=plpp0086 960.2 SEQID=6732 EMBL_NAME=plpp0085 99.1 SEQID=6582 EMBL_NAME=lpp2365
Table XNIII: List of the specific sequences of the Lens strain relative to the Paris and Philadelphia strains with their « ORF » Institut Pasteur correspondence number and accession number in the gene banks
1001.1 SEQID=6735 EMBL ΝAME= =lpll928 1002.1 SEQED=6736 EMBL"NAME= =lpll927 1003.1 SEQID=6737 EMBL"NAME= =lpll926 102.1 SEQID=6738 EMBL"NAME= =lpl0183 1020.1 SEQID=6739 EMBL"NAME= =lpll915 103.1 SEQID=6740 EMBL"NAME= =lpl0182 1040.1 SEQID=6742 EMBL NAME= =lpll900 1041.1 SEQID=6743 EMBL"NAME= =lpll899 1045.1 SEQID=6745 EMBL"NAME= =lpll896 1047.1 SEQID=6746 EMBL"NAME= =lpll895 1048.1 SEQID=6747 EMBL"NAME= =lpll894 1049.2 SEQID=6748 EMBL"NAME= =lpl0064 105.1 SEQID=6749 EMBL"NAME= =lpl0181 1050.1 SEQID=6750 EMBL NAME= =lpl0063 1059.1 SEQID=6751 EMBL"NAME= =lp!0057
106.1 SEQED=6752 EMBL NAME= =lpl0180
110.1 SEQID=6754 EMBL NAME= ;lpl0177
1105.2 SEQID=6755 EMBL NAME= =lpl0019
111.1 SEQED=6756 EMBL "NAME= =lpl0176
112.1 SEQID=6757 EMBL"NAME= =lpl0175
113.1 SEQED=6758 EMBL"NAME= =lp!0174
115.1 SEQED=6760 EMBL"NAME= =lpl0173
116.1 SEQID=6761 EMBL"NAME= =lpl0172
1204.1 SEQED=6763 EMBL NAME= =lpl2879
1206.1 SEQED=6764 EMBL NAME= =lpl2877
1207.1 SEQID=6765 EMBL NAME= =lpl2876
1209.1 SEQID=6766 EMBL~NAME= =lpl2875
1217.1 SEQED=6767 EMBL NAME= =lpl2869
1219.1 SEQED=6768 EMBL NAME= =lpl2868
1221.1 SEQED=6769 EMBL NAME= =lpl2867
1237.2 SEQID=6770 EMBL NAME= =lpl2857
1251.1 SEQID=6773 EMBL "NAME= =lpl2848
1252.1 SEQID=6774 EMBL"NAME= =lpl2847
1255.2 SEQID=6775 EMBL NAME= =lpl2845
1258.1 SEQID=6776 EMBL NAME= ipl2843
126.1 SEQED=6777 EMBL NAME= =lρl0165
127.1 SEQID=6778 EMBL "NAME= =lpl0164
1274.1 SEQED=6779 EMBL "NAME= =lpl2842
1275.1 SEQED=6780 EMBL"NAME= ipl2841
1276.1 SEQED=6781 EMBL"NAME= =lpl2840
1278.1 SEQID=6782 EMBL NAME= =lpl2839
1279.1 SEQED=6783 EMBL NAME= =lpl2838
1280.1 SEQID=6784 EMBL NAME= =lpl2837
1283.1 SEQID=6786 EMBL NAME= =lρl2835
1284.1 SEQID=6787 EMBL NAME= =lpl2834
1285.1 SEQID=6788 EMBL "NAME= ipl2833
1296.1 SEQID=6789 EMBL NAME= =lpl2827
1297.1 SEQID=6790 EMBL NAME= =lpl2826
1321.1 SEQID=6791 EMBL "NAME= =lpl2806
1422.1 SEQID=6792 EMBL "NAME= =lpl2741
1535.1 SEQID=6795 EMBL NAME= =lpl0612
154.1 SEQID=6796 EMBL NAME= =lρl0146
156.1 SEQED=6797 EMBL NAME= =lpl0145
157.1 SEQED=6798 EMBL NAME= =lpl0144
159.1 SEQED=6799 EMBL NAME= =lpl0143
1697.1 SEQED=6800 EMBL NAME= =lpl0718
1718.1 SEQED=6801 EMBL NAME= =lρl0729
1824.1 SEQID=6803 EMBL NAME= lpl0801
1826.1 SEQID=6805 EMBL NAME= =lpl0803
1827.1 SEQED=6806 EMBL NAME= =lpl0804
1834.1 SEQED=6809 EMBL NAME= lpioδi i
1955.1 SEQID=6810 EMBL NAME= lpl0904
2110.1 SEQED=6812 EMBL NAME= lpll015
2137.1 SEQED=6816 EMBL NAME= =lpll037
2141.1 SEQED=6818 EMBL NAME= Ipl 1041
2142.2 SEQID=6819 EMBL NAME=lpll042
2145.1 SEQED=6821 EMBL NAME=lpll043a
2152.2 SEQED=6823 EMBL NAME=lpll048
2155.1 SEQED=6824 EMBL NAME=lpll050
2170.1 SEQED=6827 EMBL NAME=lpll059
2180.1 SEQID=6828 EMBL NAME=lpll067
2181.1 SEQID=6829 EMBL NAME=lpll068
2182.1 SEQED=6830 EMBL NAME=lpll069
2184.1 SEQID=6831 EMBL NAME=lpll070
2186.2 SEQED=6832 EMBL NAME=lpll071
2189.1 SEQLD=6833 EMBL NAME=lpll073
2194.1 SEQED=6835 EMBL NAME=lpll076
2195.1 SEQID=6836 EMBL NAME=lpll077
2198.1 SEQED=6837 EMBL NAME=lpll080
2199.1 SEQID=6838 EMBL NAME=lpll081
2200.1 SEQID=6839 EMBL NAME=lpll082
2201.1 SEQLD=6840 EMBL NAME=lpll083
2202.1 SEQED=6841 EMBL NAME=lpll084
2203.1 SEQED=6842 EMBL NAME=lpll085
2206.1 SEQED=6844 EMBL NAME=lpll087
2207.1 SEQID=6845 EMBL NAME=lpll088
2208.1 SEQED=6846 EMBL NAME=lpll089
2210.1 SEQID=6848 EMBL NAME=lpll091
2211.1 SEQID=6849 EMBL NAME=lpll092
2213.1 SEQID=6850 EMBL NAME=lpll093
2224.1 SEQID=6851 EMBL NAME=lpll l01
2241.1 SEQED=6853 EMBL NAME=lpl0199
2242.1 SEQID=6854 EMBL NAME=lpl0200
2245.1 SEQED=6856 EMBL NAME=lpl0202
2253.1 SEQID=6860 EMBL NAME=lpl0207
2259.1 SEQID=6861 EMBL NAME=lpl0211
2260.1 SEQED=6862 EMBL NAME=lpl0212
2440.1 SEQID=6866 EMBL NAME=lpl2546
2441.1 SEQID=6867 EMBL NAME=lpl2545
2516.1 SEQID=6868 EMBL NAME=lpl2497
2518.1 SEQID=6870 EMBL NAME=lpl2495
2521.1 SEQED=6871 EMBL NAME=lpl2494
2523.1 SEQID=6872 EMBL NAME=lpl2493
2524.1 SEQID=6873 EMBL NAME=lpl2492
2525.1 SEQLD=6874 EMBL NAME=lpl2491
2526.1 SEQID=6875 EMBL NAME=lpl2490
2527.1 SEQID=6876 EMBL NAME=lpl2489
2529.1 SEQED=6877 EMBL NAME=lpl2488
2530.1 SEQED=6878 EMBL NAME=lpl2487
2531.1 SEQED=6879 EMBL NAME=lpl2486
2532.1 SEQID=6880 EMBL NAME=lpl2485
2533.1 SEQED=6881 EMBL NAME=lpl2484
2534.1 SEQED=6882 EMBL NAME=lpl2483
2540.1 SEQED=6884 EMBL NAME=lpl2477
2541.1 SEQED=6885 EMBL NAME=lpl2476
2547.1 SEQED=6886 EMBL NAME=lpl2472
2584.1 SEQID=6888 EMBL NAME=lpl2445
2640.1 SEQED=6890 EMBL NAME=lpl2399
2658.1 SEQED=6891 EMBL NAME=lpl2385
266.1 SEQLD=6892 EMBL NAME=lpl0069
267.1 SEQID=6894 EMBL NAME=lpl0068
269.1 SEQID=6895 EMBL NAME=lpl0067
270.1 SEQID=6897 EMBL NAME=lpl0066
2701.1 SEQLD=6898 EMBL NAME=lpl2354
2708.1 SEQID=6900 EMBL NAME=lpl2350
2717.1 SEQID=6901 EMBL NAME=lpl2344
2719.1 SEQID=6902 EMBL NAME=lpl2343
272.1 SEQID=6903 EMBL NAME=lpl0065
2720.1 SEQID=6904 EMBL NAME=lpl2342
2722.1 SEQID=6905 EMBL NAME=lpl2341
2723.1 SEQID=6906 EMBL NAME=lpl2340
273.2 SEQID=6907 EMBL NAME=lpll893
2738.1 SEQID=6908 EMBL NAME=lpl2330
2749.1 SEQID=6909 EMBL NAME=lpl2323
2775.1 SEQID=6910 EMBL NAME=lpl2309
2782.1 SEQID=6912 EMBL NAME=lpl2305
2795.1 SEQID=6913 EMBL NAME=lpl2295
2796.1 SEQED=6914 EMBL NAME=lpl2294
2798.1 SEQID=6915 EMBL NAME=lpl2293
2801.1 SEQED=6916 EMBL NAME=lpl2292
2802.1 SEQID=6917 EMBL NAME=lpl2291
2805.1 SEQID=6918 EMBL NAME=lpl2289
2806.1 SEQED=6919 EMBL NAME=lpl2288
2808.1 SEQID=6921 EMBL NAME=lpl2286
2810.1 SEQID=6923 EMBL NAME=lpl2284
3002.1 SEQID=6925 EMBL NAME=lpl2148
3042.1 SEQID=6926 EMBL NAME=lpl2114
3054.1 SEQID=6927 EMBL NAME=lpl2107
3056.1 SEQID=6928 EMBL NAME=lpl2106
3057.1 SEQID=6929 EMBL NAME=lpl2105
3064.1 SEQID=6930 EMBL NAME=lpl2100
3065.1 SEQID=6931 EMBL NAME=lpl2099
3066.1 SEQID=6932 EMBL NAME=lpl2098
3140.1 SEQID=6933 EMBL NAME=lpl2049
3155.1 SEQID=6935 EMBL NAME=lpl2038
3156.1 SEQID=6936 EMBL NAME=lpl2037
3158.1 SEQID=6937 EMBL NAME=lpl2036
3160.1 SEQJD=6938 EMBL NAME=lpl2035
3161.1 SEQID=6939 EMBL NAME=lpl2034
3162.1 SEQED=6940 EMBL NAME=lpl2033
3417.1 SEQED=6942 EMBL NAME=lpl0216
3420.1 SEQED=6943 EMBL NAME=lpl0217
3422.1 SEQLD=6944 EMBL NAME=lpl0218
3435.1 SEQID=6945 EMBL NAME=lpl0226
3728.1 SEQID=6948 EMBL NAME=lpl0552
3743.2 SEQID=6949 EMBL NAME=lpl0565
3744.1 SEQED=6950 EMBL NAME=lpl0566
3745.1 SEQID=6951 EMBL NAME=lpl0567
3747.2 SEQLD=6952 EMBL NAME=lpl0568a
3748.2 SEQID=6953 EMBL NAME=lpl0568b
3815.2 SEQID=6957 EMBL NAME=plpl0037
3818.1 SEQID=6959 EMBL NAME=plpl0039
3820.1 SEQID=6960 EMBL NAME=plpl0040
3822.1 SEQID=6961 EMBL NAME=plpl0042
3823.1 SEQID=6962 EMBL NAME=plpl0043
3843.1 SEQED=6972 EMBL NAME=plpl0053
3848.2 SEQED=6973 EMBL NAME=plpl0001
3850.1 SEQED=6974 EMBL NAME=plpl0002
3863.1 SEQID=6978 EMBL NAME=plpl0014
3865.1 SEQID=6979 EMBL NAME=plpl0015
3866.1 SEQID=6980 EMBL NAME=plpl0016
3868.1 SEQID=6981 EMBL NAME=plpl0017
3870.1 SEQID=6982 EMBL NAME=plpl0018
3871.1 SEQID=6983 EMBL NAME=plpl0019
3873.1 SEQID=6984 EMBL NAME=plpl0020
3874.1 SEQED=6985 EMBL NAME=plpl0021
3875.1 SEQID=6986 EMBL NAME=plpl0022
3877.1 SEQID=6987 EMBL NAME=plpl0023
3878.1 SEQID=6988 EMBL NAME=plpl0024
3880.1 SEQED=6989 EMBL NAME=plpl0025
3881.1 SEQID=6990 EMBL NAME=plpl0026
3882.1 SEQID=6991 EMBL NAME=plpl0027
3884.1 SEQED=6992 EMBL NAME=plpl0028
3886.2 SEQLD=6993 EMBL NAME=plpl0029
3888.1 SEQED=6994 EMBL NAME=plpl0030
3890.1 SEQID=6995 EMBL NAME=plpl0031
3891.2 SEQID=6996 EMBL NAME=plpl0032
3893.1 SEQID=6997 EMBL NAME=plpl0033
3894.1 SEQED=6998 EMBL NAME=plpl0034
3949.1 SEQID=7002 EMBL NAME=lpll l58
3976.1 SEQED=7003 EMBL NAME=lpll l38
3987.1 SEQID=7004 EMBL NAME=lpll l32
4007.1 SEQED=7006 EMBL NAME=lpll l l6
4172.1 SEQID=7014 EMBL NAME=lpl0194
4173.1 SEQID=7015 EMBL NAME=lpl0193
4175.1 SEQID=7017 EMBL NAME=lpl0191
4181.1 SEQID=7019 EMBL NAME=lpl0189
4182.1 SEQID=7020 EMBL NAME=lpl0188
4185.1 SEQED=7021 EMBL NAME=lpl0187
4189.2 SEQID=7023 EMBL NAME=lpll424
4196.1 SEQID=7024 EMBL NAME=lpll417
4197.1 SEQED=7025 EMBL NAME=lpll416
4248.1 SEQID=7030 EMBL NAME=lpll942
4250.1 SEQED=7031 EMBL NAME=lpll943
4251.1 SEQED=7032 EMBL NAME=lpll944
4252.1 SEQID=7033 EMBL NAME==lpl1945
4253.1 SEQED=7034 EMBL NAME==lpl1946
4254.1 SEQID=7035 EMBL NAME==lpl1947
560.1 SEQED=7037 EMBL NAME==lpl1681
561.1 SEQED=7038 EMBL NAME==lpl1680
562.1 SEQED=7039 EMBL NAME==lpl1679
564.1 SEQID=7040 EMBL NAME==lpl1678
688.1 SEQED=7042 EMBL NAME==lpl1588
689.1 SEQID=7043 EMBL"NAME==lpl1587
692.1 SEQID=7045 EMBL NAME==lpl1585
699.1 SEQID=7048 EMBL NAME==lpl1581
700.1 SEQED=7049 EMBL"NAME==lpl1580
703.1 SEQLD=7050 EMBL"NAME==lpl1579
709.1 SEQED=7053 EMBL NAME==lpl1575
711.1 SEQID=7054 EMBL"NAME=lpl1574
760.1 SEQID=7055 EMBL"NAME= =ipi1537
761.1 SEQID=7056 EMBL NAME= =lpl1536
898.1 SEQID=7057 EMBL"NAME==lpl1425
995.2 SEQID=7059 EMBL"NAME=lpl1933
997.1 SEQED=7060 EMBL"NAME4pl1931
2144.1 SEQID=6820 EMBL NAME=lpl1043b
Table XIX: List of the sequences present in the Paris and Lens strain though absent in the Philadelphia strain with their Pasteur Instimte « ORF » correspondence number and accession number in the gene banks
1082.3 SEQED=3560 EMBL_NAME=lpp 1844 1090.1 SEQID=3565 EMBL_N AME=lpp 1061 1156.3 SEQID=3606 EMBL_NAME=lppl 106 119.1 SEQID=3625 EMBL_NAME=lpp2953 121.1 SEQED=3635 EMBL_NAME=lpp2952 1225.2 SEQID=6596 EMBL_NAME=plpp0013 1226.2 SEQID=6597 EMBL_NAME=plpp0012 131.2 SEQID=3694 EMBL_NAME=lppl 099 1469.4 SEQID=3786 EMBL_NAME=lppl 843 15.1 SEQID=3803 EMBL_NAME=lppO 194 1560.2 SEQJD=3842 EMBL_NAME=lpp2477 16.1 SEQJD=3869 EMBL_NAME=lpp0193 1737.2 SEQED=3956 EMBL_NAME=lpp 1863 1875.2 SEQID=4045 EMBL_NAME=lpp2529 2.1 SEQED=4112 EMBL_NAME=lppO 163 2026.1 SEQED=4124 EMBL_NAME=lppl 909 2039.1 SEQED=4131 EMBL_NAME=lpp0243 2275.4 SEQID=6632 EMBL_NAME=plpp0014 2357.4 SEQID=4299 EMBL_NAME=lpp2054 2427.4 SEQID=4348 EMBL_NAME=lppl 869 2453.4 SEQID=4367 EMBL_NAME=lpp2450 2649.1 SEQID=4483 EMBL_NAME=lpp0667 321.3 SEQID=4804 EMBL_NAME=lpp2981 3248.1 SEQED=4829 EMBL_NAME=lpp2478 33.1 SEQED=4861 EMBL_NAME=lpp0183 3395.3 SEQED=4911 EMBL_NAME=lppl 042 3396.1 SEQID=4912 EMBL_NAME=lppl 043 3401.2 SEQID=4917 EMBL_NAME=lppl 047 341.6 SEQID=4921 EMBL_NAME=lpp0024 3413.2 SEQED=4923 EMBL_NAME=lpp2060 3414.3 SEQID=4924 EMBL_NAME=lpp2061 3499.1 SEQED=6673 EMBL_NAME=plpp0011 3500.3 SEQID=6674 EMBL_NAME=plpp0010 3563.3 SEQED=5023 EMBL_NAME=lpp0158 3594.1 SEQID=5041 EMBL_NAME=lpp0639 3600.2 SEQID=5048 EMBL_NAME=lpp2449 3601.1 SEQID=5049 EMBL_NAME=lpp2448 3657.1 SEQID=5085 EMBL_NAME=lpp2419 3734.1 SEQID=5134 EMBL_NAME=lpp0208 3744.2 SEQID=5142 EMBL_NAME=lpp 1850 3763.1 SEQID=5152 EMBL_NAME=lpp 1867 3871.1 SEQID=5211 EMBL_NAME=lρρ2070 3872.1 SEQID=5212 EMBL_NAME=lpp2069 3878.1 SEQED=5215 EMBL_NAME=lρρ2066
4039.2 SEQID=5315 EMBL_NAME=lpp0064 4040.1 SEQID=5316 EMBL_NAME=lpp0063 4045.2 SEQID=5320 EMBL_NAME=lpρ0059 417.3 SEQID=5412 EMBL_NAME=lppl 907 4276.2 SEQID=5479 EMBL_NAME=lρρ2048 4532.2 SEQED=5653 EMBL_NAME=lpp2049 4763.2 SEQID=5803 EMBL_NAME=lppl 088 4764.2 SEQID=5804 EMBL_NAME=lppl 087 5056.3 SEQID=5980 EMBL_NAME=lppl 578 5058.2 SEQID=5981 EMBL_NAME=lppl 579 5059.3 SEQID=5982 EMBL_NAME=lppl 580 506.3 SEQID=5983 EMBL_NAME=lpp2417 5080.6 SEQID=6686 EMBL_NAME=plpp0006 5087.2 SEQID=5999 EMBL_NAME=lpp2016 5106.3 SEQID=6688 EMBL_NAME=plpp0007 5147.1 SEQID=6691 EMBL_NAME=plpp0009 5176.1 SEQID=6026 EMBL_NAME=lpp0712 5382.1 SEQID=6107 EMBL_NAME=lpp 1086 5388.1 SEQED=6110 EMBL_NAME=lpp0084 5404.2 SEQED=6117 EMBL_NAME=lpp2443 5504.4 SEQED=6163 EMBL_NAME=lpp2053 553.1 SEQID=6173 EMBL_NAME=lpp2920 5584.2 SEQED=6200 EMBL_N AME=lpp 1450 5609.1 SEQED=6214 EMBL_NAME=lpp2153 58.1 SEQED=6273 EMBL_NAME=lpp0165 6036.1 SEQID=6322 EMBL_NAME=lppl449 650.4 SEQID=6385 EMBL_NAME=lρρ0668 651.3 SEQED=6386 EMBL_NAME=lpp0669 860.2 SEQID=6495 EMBL_NAME=lρρ2058 9.2 SEQID=6521 EMBL_NAME=lpp0159
Table XX: List of the sequences present in the Paris and Philadelphia strain though absent in the Lens strain with their Pasteur Instimte « ORF » correspondence number and accession number in the gene banks
102.1 SEQID=3519 EMBL NAME=lpp2364 103.1 SEQID=3525 EMBL NAME=lpp2363 104.1 SEQID=3533 EMBL NAME=lpp2362 107.1 SEQID=3553 EMBL NAME=lpρ2361 109.1 SEQID=3564 EMBL NAME=lpp2360 1107.2 SEQID=3578 EMBL NAME=lppl601 1109.3 SEQID=3579 EMBL NAME=lppl600 111.2 SEQID=3580 EMBL NAME=lpp2359 1111.3 SEQID=3581 EMBL NAME=lppl599 1211.3 SEQID=3637 EMBL NAME=lpp0258 1236.2 SEQID=6599 EMBL NAME=plpp0120 1334.3 SEQID=3707 EMBL NAME=lpp0094 1335.2 SEQID=3708 EMBL NAME=lρρ0095
1423.2 SEQID=3763 EMBL NAME= =lpp2328
1424.2 SEQED=3764 EMBL NAME= =lpp2329
1425.2 SEQED=3765 EMBL~NAME= =lpp2330
1432.3 SEQID=3767 EMBL NAME= =lpp2441
152.3 SEQED=3818 EMBL"NAME= =lpp2354
1548.2 SEQED=3834 EMBL NAME= =lpp2331
1549.1 SEQID=3835 EMBL NAME= -ipp2332
1550.2 SEQID=3836 EMBL NAME= =lpp2333
158.2 SEQED=3854 EMBL NAME= =lpp2341
159.1 SEQID=3862 EMBL"NAME= =lpp2342
160.1 SEQID=3870 EMBL"NAME= =lpp2343
1628.3 SEQID=3885 EMBL NAME= =lpp2339
1629.1 SEQID=3886 EMBL"NAME= =lpp2338
1631.3 SEQLD=3888 EMBL"NAME= =lpp2337
1635.4 SEQID=3891 EMBL"NAME= =lppl309
1639.4 SEQID=3894 EMBL"NAME-- =lppl308
168.1 SEQID=3920 EMBL ~NAME= =lpp2346
1682.4 SEQID=3922 EMBL"NAME= =lpp0045
169.1 SEQID=3927 EMBL" "NAME= =lpρ2347
1703.1 SEQED=3936 EMBL" NAME= =lppl942
1775.3 SEQID=3983 EMBL" NAME= =lpρl l30
1847.3 SEQID=4029 EMBL NAME= =lppl089
1887.2 SEQID=4053 EMBL NAME-- =lppl940
190.3 SEQID=4062 EMBL" NAME= =lpp0234
1960.2 SEQID=4096 EMBL NAME= =lpp2340
1961.3 SEQID=4097 EMBL NAME-- =lpp2355
2031.2 SEQID=4128 EMBL NAME= =lpp0330
2054.2 SEQED=4138 EMBL ~NAME= =lpp2603
2079.2 SEQID=4153 EMBL ~NAME= =lpp2455
2143.5 SEQID=4186 EMBL "NAME= =lpp2615
2169.2 SEQID=4205 EMBL NAME= =lppl890
227.2 SEQID=4247 EMBL_NAME= =lpp2358
228.1 SEQID=4253 EMBL NAME= =lpp2357
2544.3 SEQID=4425 EMBL" ~NAME= =lpp2192
2591.3 SEQID=4451 EMBL _NAME= =lpp2779
2637.1 SEQED=4474 EMBL _NAMfr =lpp2336
2639.1 SEQID=4475 EMBL _NAME= =lpp2327
2646.1 SEQID=4480 EMBL NAME -lpp0673
2730.1 SEQID=4526 EMBL"NAME =lpp0251
2808.1 SEQID=4572 EMBL_NAME =lpp0321
2849.1 SEQID=4596 EMBL_NAME =lppl912
2938.4 SEQID=4636 EMBL NAME =lpp2883
2991.1 SEQID=4675 EMBL_NAME =lpp2110
2992.2 SEQID=4676 EMBL_NAME =lpp2109
3163.1 SEQID=4778 EMBL NAME =lpp0331
3190.1 SEQID=4795 EMBL NAME =lppl007
3191.1 SEQED=4796 EMBL "NAME =lppl006
3205.3 SEQID=4802 EMBL _NAME =lpp2131
3207.5 SEQED=4803 EMBL _NAME =lpp2132
3250.1 SEQED=4832 EMBL NAME =lpp2474
3508.2 SEQID=4982 EMBL NAME=lpp2780
3588.2 SEQID=5036 EMBL NAME=lppl090
3663.1 SEQID=5090 EMBL NAME=lpp3017
3699.3 SEQID=5114 EMBL NAME=lppl404
3701.1 SEQID=5116 EMBL NAME=lppl403
371.1 SEQID=5123 EMBL NAME=lpp2039
3783.1 SEQJD=5160 EMBL NAME=lppl l44
3822.2 SEQID=5187 EMBL NAME=lpp0851
384.5 SEQID=5197 EMBL NAME=lpp0716
3884.2 SEQID=5216 EMBL NAME=lpp2440
402.2 SEQID=5304 EMBL NAME=lppl557
4030.1 SEQID=5310 EMBL NAME=lpp0096
4083.4 SEQJD=5348 EMBL NAME=lppl603
4084.3 SEQID=5349 EMBL NAME=lppl602
4220.2 SEQID=5443 EMBL NAME=lppl852
423.2 SEQID=5449 EMBL NAME=lpp2432
424.1 SEQID=5457 EMBL NAME=lpp2431
4242.3 SEQID=5458 EMBL NAME=lppl405
4249.1 SEQID=5464 EMBL NAME=lpp3009
4453.2 SEQID=5597 EMBL NAME=lppl447
4517.2 SEQED=5640 EMBL NAME=lpp2497
4528.2 SEQED=5649 EMBL NAME=lpp2052
4530.1 SEQED=5651 EMBL NAME=lpp2051
4559.1 SEQID=5667 EMBL NAME=lpp0287
4591.2 SEQED=5689 EMBL NAME=lpp2508
4819.2 SEQID=5845 EMBL NAME=lpp2498
4965.2 SEQID=5926 EMBL NAME=lpp0829c
4966.2 SEQID=5927 EMBL NAME=lpp0830
5060.2 SEQID=5984 EMBL NAME=lpp0835
5289.2 SEQID=6070 EMBL NAME=lpp0589
5328.1 SEQID=6088 EMBL NAME=lppl565
5337.1 SEQID=6090 EMBL NAME=lpp2896
5340.1 SEQID=6091 EMBL NAME=lppl947
538.1 SEQID=6105 EMBL NAME=lpp0859
5496.2 SEQID=6157 EMBL NAME=lpp0829b
5656.2 SEQID=6238 EMBL NAME=lpp2887
5723.2 SEQID=6261 EMBL NAME=lppl944
5871.4 SEQJD=6288 EMBL NAME=lpp2037
590.4 SEQID=6299 EMBL NAME=lpp2886
592.3 SEQID=6305 EMBL NAME=lpp2348
5920.3 SEQID=6306 EMBL NAME=lpp2311
593.2 SEQID=6308 EMBL NAME=lpρ2349
594.1 SEQED=6309 EMBL NAME=lpp2350
5999.1 SEQED=6317 EMBL NAME=lpp0039
6002.2 SEQID=6318 EMBL NAME=lpp0881
6110.1 SEQID=6334 EMBL NAME=lpρ0829a
615.5 SEQID=6337 EMBL NAME=lppl562
6151.1 SEQED=6338 EMBL NAME=lpp0038
6159.1 SEQID=6339 EMBL NAME=lpp2410
6160.1 SEQID=6341 EMBL NAME=lpp2402
143
6178.1 SEQED=6343 EMBL NAME= lpp0210 6180.1 SEQID=6345 EMBL NAME= lppl035 6186.1 SEQED=6346 EMBL" NAME= lpp0882 6195.2 SEQID=6350 EMBL NAME= lpp0717 6285.1 SEQID=6362 EMBL NAME= lpp2895 6309.1 SEQID=6364 EMBL NAME= lpp 1945 6318.1 SEQID=6366 EMBL" NAME= lppl 851 6320.1 SEQID=6368 EMBL NAME= lppl813 6322.1 SEQLD=6369 EMBL NAME= lppl812 743.4 SEQID=6432 EMBL NAME= lpp2368 744.4 SEQID=6433 EMBL NAME= lpp2369 818.2 SEQID=6469 EMBL NAME= lpp2335 819.2 SEQID=6470 EMBL NAME= lpp2334 864.1 SEQID=6498 EMBL" NAME= lpp2055 901.2 SEQID=6525 EMBL" NAME= lpp2471 938.3 SEQED=6550 EMBL" NAME= lpp 1253 96.6 SEQED=6562 EMBL NAME= lpp2367 97.2 SEQID=6568 EMBL" NAME= lpp2366 979.3 SEQED=6574 EMBL" NAME= lpp2494 980.1 SEQID=6575 EMBL" NAME= lpp2495 981.2 SEQID=6576 EMBL NAME= lpp2496
Table XXI: List of the sequences present in the Philadelphia and Lens strain though absent in the Paris strain with their Pasteur Institute « ORF » correspondence number and accession number in the gene banks
1038.1 SEQID=6741 EMBL NAME= =lpll901 1043.1 SEQID=6744 EMBL"NAME= =lpll897 1073.1 SEQID=6753 EMBL NAME= =lpl0044 1130.1 SEQID=6759 EMBL NAME= =lpl2933 117.1 SEQID=6762 EMBL NAME= =lplO171 124.1 SEQID=6771 EMBL""NAME =lplO167 125.1 SEQID=6772 EMBL NAME =lpl0166 1282.1 SEQID=6785 EMBL NAME =lpl2836 1434.1 SEQID=6793 EMBL"NAME =lpl2732 148.1 SEQID=6794 EMBL"NAME =lpl0150 175.1 SEQID=6802 EMBL NAME= =lpl0132 1825.1 SEQID=6804 EMBL""NAME =lpl0802 1828.1 SEQED=6807 EMBL"NAME =lpl0805 1829.1 SEQED=6808 EMBL"NAME^ =lpl0806 2100.1 SEQED=6811 EMBL NAME-- =lpllOO6 2134.1 SEQID=6813 EMBL NAME: =lpll034 2135.1 SEQID=6814 EMBL NAME: =lpll035 2136.1 SEQID=6815 EMBL"NAME: =lpll036 2139.1 SEQID=6817 EMBL"NAME= =lpll039 2151.1 SEQID=6822 EMBL"NAME= =lpllO47 2157.1 SEQID=6825 EMBL"NAME= -lpllO51 2168.1 SEQED=6826 EMBL"NAME= =lpll058
2192.1 SEQID=6834 EMBL_NAME=lpl 1075
2.205.1 SEQID=6843 EMBL_NAME=lpll086
2209.1 SEQID=6847 EMBL_NAME=lpll090
2238.1 SEQID=6852 EMBL_NAME=lpll l lO
2244.1 SEQID=6855 EMBL_NAME=lpl0201
2246.1 SEQED=6857 EMBL_NAME=lpl0203
2247.1 SEQID=6858 EMBL_NAME=lpl0204
2248.1 SEQID=6859 EMBL_NAME=lpl0205
2261.1 SEQID=6863 EMBL_NAME=lpl0213
2392.1 SEQED=6864 EMBL_NAME=lpl2580
2422.1 SEQID=6865 EMBL_NAME=lpl2558
2517.1 SEQED=6869 EMBL_NAME=lpl2496
2539.1 SEQID=6883 EMBL_NAME=lpl2478
2558.1 SEQID=6887 EMBL_NAME=lρl2465
2587.1 SEQED=6889 EMBL_NAME=lpl2443
2660.1 SEQID=6893 EMBL_NAME=lpl2384
2696.1 SEQED=6896 EMBL_NAME=lpl2358
2706.1 SEQID=6899 EMBL_NAME=lpl2351
2777.1 SEQLD=6911 EMBL_NAME=lpl2308
2807.1 SEQED=6920 EMBL_NAME=lpl2287
2809.1 SEQID=6922 EMBL_NAME=lpl2285
293.1 SEQID=6924 EMBL_NAME=lpl 1879
3151.1 SEQID=6934 EMBL_NAME=lpl2042
322.1 SEQID=6941 EMBL_NAME=lpll857
3536.1 SEQID=6946 EMBL_NAME=lpl0286
3537.1 SEQED=6947 EMBL_NAME=lpl0287
3749.2 SEQID=6954 EMBL_NAME=lpl0569
3788.1 SEQED=6955 EMBL_NAME=lpl0593
3793.1 SEQID=6956 EMBL_NAME=lpl0596
3816.1 SEQID=6958 EMBL_NAME=plpl0038
3827.1 SEQED=6963 EMBL_NAME=plpl0044
3828.1 SEQID=6964 EMBL_NAME=plpl0045
3831.1 SEQID=6965 EMBL_NAME=plpl0046
3835.1 SEQID=6966 EMBL_NAME=plpl0047
3836.1 SEQID=6967 EMBL_NAME=plpl0048
3837.1 SEQID=6968 EMBL_NAME=plpl0049
3839.1 SEQID=6969 EMBL_NAME=plpl0050
3840.1 SEQID=6970 EMBL_NAME=plpl0051
3841.1 SEQID=6971 EMBL_NAME=plpl0052
3859.1 SEQID=6975 EMBL_NAME=plpl0011
3861.1 SEQID=6976 EMBL_NAME=plplOO 12
3862.1 SEQID=6977 EMBL_NAME=plplOO 13
3895.1 SEQID=6999 EMBL_NAME=plpl0035
3896.1 SEQED=7000 EMBL_NAME=plpl0036
3937.1 SEQED=7001 EMBL_NAME=lpll l65
3995.1 SEQLD=7005 EMBL_NAME=lpll l25
4011.1 SEQID=7007 EMBL_NAME=lpll l l3
4085.1 SEQED=7008 EMBL_NAME=lpl0408
4093.1 SEQID=7009 EMBL_NAME=lpl0415
4111.1 SEQED=7010 EMBL_NAME=lpl0432
2005/049642 145
4169.1 SEQID=7011 EMBL NAME=lpl0197 4170.1 SEQID=7012 EMBL NAME=lpl0196 4171.2 SEQID=7013 EMBL NAME=lpl0195 4174.1 SEQID=7016 EMBL NAME=lpl0192 4177.1 SEQID=7018 EMBL NAME=lpl0190 4188.1 SEQID=7022 EMBL NAME=lpl0185 4203.1 SEQID=7026 EMBL NAME=lpll412 4229.2 SEQED=7027 EMBL NAME=lpll393 4236.2 SEQLD=7028 EMBL NAME=lpll965 4237.1 SEQED=7029 EMBL NAME=lpll934 442.1 SEQID=7036 EMBL NAME=lpll768 566.1 SEQID=7041 EMBL NAME=lpll676 690.1 SEQLD=7044 EMBL NAME=lpll586 694.1 SEQID=7046 EMBL NAME=lpll584 697.1 SEQID=7047 EMBL NAME=lpll582 705.1 SEQID=7051 EMBL NAME=lpll578 707.1 SEQED=7052 EMBL NAME=lpll577 899.2 SEQID=7058 EMBL NAME=lpl2032
. . ^ . . „. . Specific to % homology _ . _ Position Position Signal „ . , A . . Presence in L. Legionella SEQ ID No. ORF Cont Sens „ „ Paris/ retained , , 1 2 P ι ι i - nnu longbeachae specific Philadelphia ORFs
SEQ ID No. 3098 5906.1 Contig29 m 2 541
SEQ ID No. 3097 5905.1 Contig29 P 747 1703 97%
SEQ ID No. 3181 6131.1 Contig31 P 1 2313 +
SEQ ID No. 3180 6130.1 Contig32 m 74 1048 +
SEQ ID No. 3179 6128.1 Contig32 m 1045 2223 +
SEQ ID No. 3178 6125.1 Contig32 m 2799 3074 +
SEQ ID No. 3177 6123.1 Contig32 m 3005 3130 + +
SEQ ID No. 3176 6122.1 Contig32 P 3158 3457 + +
SEQ ID No. 3175 6121.1 Contig33 P 1 570 +
SEQ ID No. 1494 3257.2 Contig33 P 933 1391 +
SEQ ID No. 1495 3258.2 Contig33 P 1363 2307 +
SEQ ID No. 659 1924.4 Contig33 m 2329 2898 +
SEQ ID No. 658 1923.2 Contig33 m 2892 3113 +
SEQ ID No. 657 1920.3 Contig33 m 3738 4808 +
SEQ ID No. 2987 5626.2 Contig33 m 4971 5936 +
SEQ ID No. 3122 5987.1 Contig33 m 5866 6501 + +
SEQ ID No. 3089 5898.2 Contig34 m 2 862 + + +
SEQ ID No. 2792 523.2 Contig34 m 1137 2051 + 94%
SEQ ID No. 2795 524.2 Contig34 P 2016 3128 98%
SEQ ID No. 2804 526.2 Contig34 P 3100 3735 97%
SEQ ID No. 2812 528.2 Contig34 m 3861 4526 99%
SEQ ID No. 2816 529.2 Contig34 m 4519 4866 100%
SEQ ID No. 1736 3609.1 Contig34 P 5143 5745 99%
SEQ ID No. 1737 3610.1 Contig34 P 5784 6974 99%
SEQ ID No. 1738 3611.2 Contig34 m 7103 7888 98%
SEQ ID No. 1739 3612.2 Contig34 m 8026 8814 96%
SEQ ID No. 1740 3613.3 Contig34 P 9024 10349 97%
SEQ ID No. 1651 3496.2 Contig36 P 96 590 100%
SEQ ID No. 1650 3494.2 Contig36 P 706 1026 + 100%
SEQ ID No. 789 216.2 Contig36 P 1033 2307 100%
SEQ ID No. 782 215.1 Contig36 P 2289 2777 100%
SEQ ID No. 773 214.1 Contig36 P 2823 3278 100%
SEQ ID No. 765 213.1 Contig36 P 3226 4227 100%
SEQ ID No. 759 212.1 Contig36 P 4308 4799 100%
SEQ ID No. 755 211.1 Contig36 P 4667 5851 100%
SEQ ID No. 746 209.2 Contig36 m 5843 6589 + 100%
SEQ ID No. 2930 5526.2 Contig36 m 6590 7255 + 100%
SEQ ID No. 1037 2549.3 Contig36 P 7391 8842 99%
SEQ ID No. 1530 331.2 Contig36 m 8894 10513 100%
SEQ ID No. 1542 333.3 Contig36 P 10752 14054 100%
SEQ ID No. 3021 5690.2 Contig36 P 14241 14747 + 100%
SEQ ID No. 2651 4972.3 Contig36 P 14883 15563 100%
SEQ ID No. 2649 4969.3 Contig36 P 15689 16312
SEQ ID No. 2648 4968.2 Contig36 P 16308 16703 100%
SEQ ID No. 2597 4883.2 Contig37 P 189 647 100%
SEQ ID No. 2598 4884.1 Contig37 P 629 1198 99%
SEQ ID No. 2599 4885.2 Contig37 P 1299 1742 100%
SEQ ID No. 2968 5592.2 Contig37 P 1743 2447 100%
SEQ ID No. 753 2106.2 Contig37 P 2608 3153 100%
SEQ ID No. 752 2104.1 Contig37 P 3175 3564 100%
SEQ ID No. 130 1106.5 Contig37 P 3626 7762 99%
SEQ ID No. 489 1663.6 Contig37 P 7788 12056 99%
SEQ ID No. 2971 5596.3 Contig37 P 12164 12550 100%
SEQ ID No. 2970 5595.2 Contig37 P 12544 13098 100%
SEQ ID No. 2969 5593.1 Contig37 m 12853 13182 98%
SEQ ID No. 2305 4468.2 Contig37 P 13110 15197 99% +
SEQ ID No. 2304 4466.3 Contig37 P 15176 16408 100% +
SEQ ID No. 2303 4465.3 Contig37 m 15697 16179 100%
SEQ ID No. 2302 4464.1 Contig37 P 16396 16731 100%
SEQ ID No. 2301 4463.2 Contig37 P 16754 17416 99%
SEQ ID No. 2300 4462.3 Contig37 P 17413 18021 99%
SEQ ID No. 3084 5892.1 Contig37 P 18006 18314 100%
SEQ ID No. 3085 5894.3 Contig37 P 18311 19153 100%
SEQ ID No. 3086 5895.3 Contig37 P 19163 19450 100%
SEQ ID No. 187 1199.4 Contig37 P 19451 19795 100%
SEQ ID No. 188 1201.1 Contig37 P 19786 20454 100%
SEQ ID No. 189 1202.2 Contig37 P 20459 20884 - 100%
SEQ ID No. 2073 4133.1 Contig37 P 20866 21078 +
SEQ ID No. 2074 4134.1 Contig37 P 21059 21334 _ 100% ,
SEQ ID No. 2075 4135.1 Contig37 P 21381 21788 - 100% .
SEQ ID No. 2076 4136.1 Contig37 P 21789 22130 _ 99% .
SEQ ID No. 2077 4138.1 Contig37 P 22137 22697 - 100%
SEQ ID No. 2078 4139.1 Contig37 P 22698 23012 _ 100% .
SEQ ID No. 2079 4140.1 Contig37 P 22960 23430 _ 99% ,
SEQ ID No. 2080 4141.1 Contig37 P 23421 23987 - 100%
SEQ ID No. 2081 4142.3 Contig37 P 23941 24357 _ 100% _
SEQ ID No. 2082 4143.3 Contig37 P 24358 24873 _ 100%
SEQ ID No. 2083 4144.3 Contig37 P 24855 25061 +
SEQ ID No. 2084 4145.3 Contig37 P 25049 25495 _ 100% .
SEQ ID No. 2085 4148.3 Contig37 P 25483 26826 _ 99% ,
SEQ ID No. 2086 4149.1 Contig37 P 27019 27390 _ 100%
SEQ ID No. 2088 4151.1 Contig37 P 27411 27812 - 100% _
SEQ ID No. 2089 4153.1 Contig37 P 27826 28449 + .
SEQ ID No. 2090 4154.2 Contig37 P 28468 29460 _ 99%
SEQ ID No. 2941 5540.1 Contig37 P 29461 29862 . 99% +
SEQ ID No. 2940 5539.1 Contig37 m 29484 29870 _ 98% +
SEQ ID No. 2939 5538.2 Contig37 m 29929 30444 _ 99%
SEQ ID No. 3087 5896.1 Contig37 m 30492 30977 +
SEQ ID No. 3088 5897.1 Contig37 P 30844 31131 +
SEQ ID No. 3173 6119.1 Contig39 m 1 531 +
SEQ ID No. 2988 5630.1 Contig39 P 577 1083 _ 91%
SEQ ID No. 3048 5774.2 Contig39 P 1095 2222 _ 79% +
SEQ ID No. 741 2080.3 Contig39 P 2669 3373 - 94%
SEQ ID No. 1955 397.2 Contig39 P 3491 4015 - 100% +
SEQ ID No. 1962 398.2 Contig39 P 3990 5279 _ 96% +
SEQ ID No. 1586 3394.1 Contig39 P 6091 6318 + +
SEQ ID No. 1587 3395.2 Contig39 m 6445 7326 + +
SEQ ID No. 1588 3396.1 Contig39 P 7740 8249 + +
SEQ ID No. 1589 3397.1 Contig39 m 8412 8972 _ 89%
SEQ ID No. 1590 3398.1 Contig39 m 8984 9424 - 87%
SEQ ID No. 1591 3399.1 Contig39 m 9506 9724 » 92% +
SEQ ID No. 3106 5935.1 Contig39 P 9719 9898 - 87% +
SEQ ID No. 1593 3401.2 Cont g39 P 10196 10510 - SEQ ID No. 1594 3402.2 Cont g39 m 10565 11563 - 95% SEQ ID No. 1595 3404.2 Cont g39 m 11457 12302 - 91% SEQ ID No. 1596 3406.2 Conti g39 P 12883 13545 + SEQ ID No. 2832 5322.1 Conti g39 m 13558 13917 - + SEQ ID No. 2831 5321.1 Conti g39 m 13921 14205 - + SEQ ID No. 2615 4914.2 Conti g39 P 14302 14595 + + + SEQ ID No. 2614 4913.2 Conti g39 m 14931 15887 + + SEQ ID No. 3226 677.6 Conti g39 m 16069 18072 + 95% SEQ ID No. 3225 675.6 Conti g39 m 18045 18512 89% + SEQ ID No. 3224 673.3 Conti g39 m 18568 20133 - 97% + SEQ ID No. 1712 3575.2 Conti g39 m 20124 21500 + 98% SEQ ID No. 1711 3574.1 Conti g39 m 21501 22601 - 95% SEQ ID No. 116 1089.2 Conti g39 m 22468 25227 - 96% SEQ ID No. 118 1090.1 Cont g39 m 25228 25578 87% SEQ ID No. 119 1091.2 Conti g39 m 25532 26836 + 95% SEQ ID No. 120 1092.3 Cont g39 m 26788 27693 - 98% SEQ ID No. 2684 5031.2 Conti g39 m 27551 28276 - 97% + SEQ ID No. 2685 5032.1 Conti g39 m 28200 28625 - 98% + SEQ ID No. 2686 5033.1 Conti g39 m 28574 28948 + 99% + SEQ ID No. 2829 5319.1 Conti g39 m 29202 29537 - 97% + SEQ ID No. 2828 5318.2 Conti g39 m 29538 30179 - 96% SEQ ID No. 2072 4132.3 Conti g39 m 30161 30625 - 99% SEQ ID No. 2071 4131.3 Conti g39 m 30592 31416 + 94% SEQ ID No. 3049 5776.1 Conti g39 m 31406 31666 - + SEQ ID No. 2070 4130.2 Conti g39 m 31603 32019 + + SEQ ID No. 2068 4129.1 Cont g39 m 31995 32513 + + SEQ ID No. 2067 4128.1 Conti g39 m 32401 32640 - + SEQ ID No. 2066 4127.1 Conti g39 m 32613 33146 - + + SEQ ID No. 2065 4126.2 Conti g39 m 32977 33888 - + + SEQ ID No. 2064 4123.2 Conti g39 P 34046 34300 75% SEQ ID No. 3230 682.2 Cont g39 P 34304 35275 - + SEQ ID No. 3229 681.3 Conti ιg39 m 35343 36095 - + SEQ ID No. 3228 680.3 Conti g39 m 36188 36706 - + SEQ ID No. 3227 678.3 Conti g39 m 36741 37847 + SEQ ID No. 2639 4955.4 Conti g39 m 37811 39142 - 85%
SEQ ID No. 2638 4954.2 Contig39 m 39111 39632 - 93%
SEQ ID No. 2637 4952.1 Contig39 m 39610 40287 - 81%
SEQ ID No. 2636 4951.2 Contig39 m 40214 40783 - 90%
SEQ ID No. 2858 5382.1 Contig39 P 40834 41157 + +
SEQ ID No. 2515 4764.2 Contig39 P 41153 41398 + +
SEQ ID No. 2514 4763.2 Contig39 m 41667 42947 +
SEQ ID No. 609 1847.3 Contig39 P 43268 45091 - 97%
SEQ ID No. 1718 3588.2 Contig39 P 45192 45809 - 98%
SEQ ID No. 1717 3586.2 Contig39 P 46123 47250 - 88%
SEQ ID No. 207 1229.3 Contig39 P 47244 47789 - 95%
SEQ ID No. 206 1227.1 Contig39 P 48300 49013 - 95%
SEQ ID No. 926 2383.4 Contig39 P 49143 49373 + +
SEQ ID No. 309 1390.5 Contig39 P 49497 49979 - 96%
SEQ ID No. 310 1391.1 Contig39 P 50090 50704 - 97%
SEQ ID No. 311 1392.2 Contig39 m 50780 51103 +
SEQ ID No. 1716 3584.1 Contig39 m 51193 51399 +
SEQ ID No. 260 131.2 Contig39 m 51668 52141 +
SEQ ID No. 3172 6117.1 Contig39 P 52705 55920 +
SEQ ID No. 3047 577.2 Contig39 P 56130 57014 +
SEQ ID No. 1816 3729.1 Contig39 m 57019 57618 +
SEQ ID No. 1817 3730.1 Contig39 m 57909 58367 +
SEQ ID No. 1818 3731.1 Contig39 m 58368 58904 +
SEQ ID No. 1819 3732.1 Contig39 P 59289 60173 +
SEQ ID No. 1820 3734.1 Contig39 P 60441 61568 +
SEQ ID No. 1821 3735.2 Contig39 P 61850 63517 +
SEQ ID No. 3079 5874.1 Contig40 m 2 553 +
SEQ ID No. 3185 615.5 Contig40 P 534 1337 +
SEQ ID No. 1176 2767.1 Contig40 m 2006 3424 + +
SEQ ID No. 1969 399.2 Contig40 m 3481 5943 + +
SEQ ID No. 1978 400.1 Contig40 m 5934 6746 +
SEQ ID No. 1985 401.2 Contig40 P 6795 8066 +
SEQ ID No. 1993 402.2 Contig40 P 8321 9913 + + +
SEQ ID No. 760 2120.2 Contig40 P 10617 11687 - 97% +
SEQ ID No. 107 1075.3 Contig40 P 11840 13171 - 99%
SEQ ID No. 106 1074.3 Contig40 P 13158 15182 - 98%
SEQ ID No. 761 2121.2 Contig40 P 15227 15976 + - 97%
SEQ ID No. 1175 2765.2 Contig40 m 16013 16903 95% +
SEQ ID No. 1174 2763.2 Contig40 m 16914 17111 +
SEQ ID No. 1173 2761.1 Contig40 P 17411 17758 100%
SEQ ID No. 1172 2760.1 Contig40 P 17730 17999
SEQ ID No. 1170 2759.1 Contig40 P 17938 18912 98%
SEQ ID No. 1169 2757.1 Contig40 P 18913 19374 98%
SEQ ID No. 1168 2755.1 Contig40 P 19564 21600 98%
SEQ ID No. 1167 2754.2 Contig40 m 21696 22277 99%
SEQ ID No. 1166 2753.1 Contig40 m 22278 22781 98%
SEQ ID No. 1165 2752.1 Contig40 m 22782 23303 + 100%
SEQ ID No. 1164 2751.1 Contig40 m 23593 24363 99%
SEQ ID No. 3351 859.2 Contig40 P 24703 26091 99%
SEQ ID No. 3350 858.2 Contig40 P 26045 26566 97%
SEQ ID No. 371 1493.6 Contig40 P 26618 31495 99%
SEQ ID No. 509 1697.2 Contig40 P 31501 32064 98%
SEQ ID No. 510 1698.2 Contig40 m 32095 33180 96%
SEQ ID No. 2882 5429.1 Contig40 P 33195 33449 97%
SEQ ID No. 1047 2562.2 Contig40 P 33385 33834 95%
SEQ ID No. 1046 2560.3 Contig40 P 33902 34498 99%
SEQ ID No. 1044 2559.3 Contig40 P 34609 35655 99%
SEQ ID No. 2881 5428.1 Contig40 P 35656 36054 100%
SEQ ID No. 2880 5427.1 Contig40 P 36055 36756 100%
SEQ ID No. 2879 5426.2 Contig40 P 36750 37220 100%
SEQ ID No. 1010 2504.4 Contig40 P 37208 38215 99%
SEQ ID No. 1009 2501.2 Contig40 P 38209 39480 + 99%
SEQ ID No. 1572 3367.1 Contig40 P 39438 40316 98%
SEQ ID No. 3241 699.3 Contig40 m 40345 41484 96%
SEQ ID No. 3243 701.2 Contig40 m 41465 42997 96%
SEQ ID No. 3244 702.2 Contig40 m 42928 43755 96%
SEQ ID No. 1571 3365.2 Contig40 m 43915 44994 95%
SEQ ID No. 1570 3363.2 Contig40 m 44988 45944 99%
SEQ ID No. 1569 3362.1 Contig40 P 46177 46929 97%
SEQ ID No. 1568 3360.1 Contig40 P 46986 47717 + 89%
SEQ ID No. 184 1190.4 Contig40 m 47773 49917 + 96%
SEQ ID No. 1566 3359.2 Contig40 m 50090 52309 99%
SEQ ID No. 3026 5701.2 Contig40 m 52481 53623 99%
SEQ ID No. 2799 5250.2 Conti g40 m 53596 54588 100% SEQ ID No. 1029 2535.2 Cont g40 m 54563 55675 100% SEQ ID No. 1030 2536.2 Contig40 P 55808 57139 95% SEQ ID No. 2349 4523.1 Contig40 P 57063 57629 96% SEQ ID No. 2350 4524.1 Contig40 P 57787 58506 91% SEQ ID No. 2351 4525.2 Contig40 m 58554 60278 98% SEQ ID No. 2730 5094.2 Contig40 P 60305 61030 98% SEQ ID No. 2729 5093.3 Cont g40 P 61021 61755 98% SEQ ID No. 2728 5092.3 Contig40 P 61746 62498 98% SEQ ID No. 2800 5253.1 Contig40 m 62587 63249 99% SEQ ID No. 2472 4705.3 Contig40 P 63281 64168 98% SEQ ID No. 2473 4706.3 Cont g40 P 64231 64728 97% SEQ ID No. 2474 4709.1 Contig40 P 64725 65897 99% SEQ ID No. 2475 4710.1 Contig40 P 66072 66860 98% SEQ ID No. 2476 4711.1 Contig40 P 66832 67404 98% SEQ ID No. 2477 4712.2 Cont g40 P 67379 68674 97% SEQ ID No. 2801 5254.1 Cont g40 P 68650 69366 99% SEQ ID No. 2802 5255.1 Contig40 P 69367 70545 99% SEQ ID No. 2658 4987.2 Contig40 P 70541 71938 99% SEQ ID No. 2584 4865.2 Contig40 m 72015 72449 100% SEQ ID No. 3074 5857.1 Cont g40 m 72492 72932 97% SEQ ID No. 3073 5856.1 Contig40 m 72914 73102 + SEQ ID No. 3080 5875.1 Contig4i m 1 303 + SEQ ID No. 1882 3837.3 Contig4i P 521 2047 92% SEQ ID No. 1881 3835.2 Contig4i P 2208 2951 96% SEQ ID No. 1051 2569.3 Contig4i P 3162 3887 95% SEQ ID No. 1050 2568.1 Contig4i m 3950 4378 98% SEQ ID No. 1049 2567.2 Contig4i m 4563 5330 99% SEQ ID No. 1880 3834.1 Contig4i P 5573 6613 96% SEQ ID No. 1879 3832.1 Contiig4i P 6654 7868 97% SEQ ID No. 1878 3830.2 Contig4i P 7864 8817 96% SEQ ID No. 1549 334.5 Cont g4i m 8929 11910 97% SEQ ID No. 1557 335.1 Contig4i P 12010 12723 98% SEQ ID No. 1567 336.1 Contig4i P 12734 13735 99% SEQ ID No. 1574 337.2 Contig4i m 13745 15148 92% SEQ ID No. 1583 339.4 Contig4i P 15296 15880
SEQ ID No. 1597 341.6 Contig41 P 16002 16445
SEQ ID No. 2856 5380.3 Contig41 P 16574 17140 97%
SEQ ID No. 2263 4395.2 Contig41 m 17210 19366 99%
SEQ ID No. 2264 4396.1 Contig41 m 19330 19821 98%
SEQ ID No. 2265 4398.2 Contig41 m 20000 21013 99%
SEQ ID No. 2996 5644.3 Contig41 m 21132 22817 98%
SEQ ID No. 2619 4919.1 Contig41 m 23043 24431 99%
SEQ ID No. 2618 4918.2 Contig41 m 24412 25359 99%
SEQ ID No. 2995 5642.1 Contig41 m 25302 26252 99%
SEQ ID No. 1911 3895.2 Contig41 m 26209 27321 99%
SEQ ID No. 1910 3892.3 Contig41 m 27384 28613 99%
SEQ ID No. 2784 5216.2 Contig41 m 29032 29880 97%
SEQ ID No. 3375 891.3 Contig41 P 29993 31594 98%
SEQ ID No. 3374 889.1 Contig41 P 31734 32231 99%
SEQ ID No. 3373 888.3 Contig41 m 32336 33607 99%
SEQ ID No. 3444 986.3 Contig41 m 33608 33868 100%
SEQ ID No. 3445 989.3 Contig41 P 34035 35312 98%
SEQ ID No. 3447 991.2 Contig41 P 35832 36698
SEQ ID No. 3400 925.3 Contig41 P 36685 38610 99%
SEQ ID No. 84 1040.3 Contig41 P 38592 39653 96%
SEQ ID No. 85 1041.3 Contig41 m 40023 42467 99%
SEQ ID No. 939 2406.3 Contig41 m 42737 43828 97%
SEQ ID No. 938 2404.2 Contig41 m 43795 44931 98%
SEQ ID No. 937 2402.3 Contig41 m 44912 46285 100%
SEQ ID No. 2687 5037.1 Contig41 P 46971 47156
SEQ ID No. 2688 5038.1 Contig41 P 47157 47501 100%
SEQ ID No. 2689 5039.2 Contig41 P 47465 47713 100%
SEQ ID No. 2690 5040.2 Contig41 P 47714 49393 99%
SEQ ID No. 173 1174.2 Contig41 P 49394 50734 98%
SEQ ID No. 174 1176.5 Contig41 P 51021 52868 98%
SEQ ID No. 1048 2565.4 Contig41 m 52865 53674 97%
SEQ ID No. 1205 2815.2 Contig41 m 53870 55639
SEQ ID No. 1206 2816.2 Contig41 P 56182 58359 99%
SEQ ID No. 1207 2818.1 Contig41 m 58410 59606 98%
SEQ ID No. 1209 2820.1 Contig41 m 59607 60461 98%
SEQ ID No. 1210 2822.1 Contig41 P 60447 62309 97%
SEQ ID No. 763 2127.2 Contig41 P 62316 62687 99% SEQ ID No. 764 2129.1 Contig41 P 62699 63313 98% SEQ ID No. 766 2130.1 Contig41 P 63314 63889 98% SEQ ID No. 767 2132.2 Contig41 P 63867 64478 98% SEQ ID No. 768 2133.1 Contig41 m 64548 64796 98% SEQ ID No. 769 2135.1 Contig41 P 64932 65342 99% SEQ ID No. 770 2136.1 Contig41 P 65284 66171 100%
SEQ ID No. 771 2137.1 Contig41 P 66209 66487 100%
SEQ ID No. 772 2138.2 Contig41 P 66491 67009 100%
SEQ ID No. 774 2140.2 Contig41 P 66999 67553 98%
SEQ ID No. 412 1552.3 Contig41 P 67559 69124 100%
SEQ ID No. 413 1555.2 Contig41 P 69211 70080 99%
SEQ ID No. 414 1556.2 Contig41 P 70081 71469 100%
SEQ ID No. 1211 2824.1 Contig41 P 71460 71903 100%
SEQ ID No. 3389 910.4 Contig41 P 72529 74787 98%
SEQ ID No. 3388 909.2 Contig41 m 74894 75811 98%
SEQ ID No. 758 2119.2 Contig41 P 75964 77661 98%
SEQ ID No. 1212 2826.1 Contig41 P 78109 79644 99% +
SEQ ID No. 1213 2827.3 Contig41 P 79904 82522 99% +
SEQ ID No. 1214 2828.3 Contig41 P 82606 82911
SEQ ID No. 2927 5521.1 Contig42 P 297 830 100%
SEQ ID No. 2926 5520.1 Contig42 P 831 1391 +
SEQ ID No. 1793 3684.2 Contig42 P 1367 2128 100% +
SEQ ID No. 1794 3685.1 Contig42 P 2210 2536 100%
SEQ ID No. 1795 3686.1 Contig42 P 2533 4083 100%
SEQ ID No. 1796 3687.1 Contig42 P 4196 5224 100%
SEQ ID No. 1797 3688.1 Contig42 m 5536 5817 100%
SEQ ID No. 3298 776.3 Contig42 m 6700 8211 100%
SEQ ID No. 3299 778.2 Contig42 m 8247 9074 100%
SEQ ID No. 3300 779.4 Contig42 P 9150 10607 99%
SEQ ID No. 2062 4119.2 Contig42 P 10583 11368 100%
SEQ ID No. 2061 4118.3 Contig42 P 11344 12006 100%
SEQ ID No. 2060 4116.3 Contig42 P 12007 12717 100%
SEQ ID No. 2059 4115.1 Contig42 P 12639 13736 100%
SEQ ID No. 555 1764.3 Contig42 P 13730 14869 100%
SEQ ID No. 556 1765.4 Contig42 P 14850 15467 100%
SEQ ID No. 592 1818.4 Contig42 P 15448 16980 100%
SEQ ID No. 2058 4113.1 Contig42 P 17043 17633 100%
SEQ ID No. 2057 4112.1 Contig42 P 17634 18518 100%
SEQ ID No. 2056 4111.2 Contig42 P 18460 19557 100%
SEQ ID No. 2920 5500.1 Contig42 P 19572 21095 100%
SEQ ID No. 2918 5499.1 Contig42 P 21199 22125 100%
SEQ ID No. 2917 5498.1 Contig42 P 22116 23102 100%
SEQ ID No. 2916 5497.1 Contig42 P 23074 24090 100%
SEQ ID No. 3171 6110.1 Contig42 P 24187 24447 100% +
SEQ ID No. 2915 5496.2 Contig42 P 24434 24973 100% +
SEQ ID No. 2646 4965.2 Contig42 P 24949 25446 100% +
SEQ ID No. 2647 4966.2 Contig42 m 25547 27379 100% +
SEQ ID No. 666 1935.4 Contig42 m 27346 28494 100% +
SEQ ID No. 665 1934.4 Contig42 m 28475 29512 100% +
SEQ ID No. 2530 4782.3 Contig42 m 29466 30560 100%
SEQ ID No. 2932 5528.2 Contig42 m 30570 31292 100% +
SEQ ID No. 2705 5060.2 Contig42 P 31581 33122 100% +
SEQ ID No. 2706 5061.4 Contig42 P 33331 35037 100% +
SEQ ID No. 2091 4156.3 Contig42 P 34991 35854 100%
SEQ ID No. 2092 4157.2 Contig42 P 35845 37275 100%
SEQ ID No. 2093 4158.1 Contig42 P 37269 38429 100%
SEQ ID No. 2094 4159.1 Contig42 m 38443 39345 100%
SEQ ID No. 287 1359.2 Contig42 P 40114 41187 100%
SEQ ID No. 2095 4160.3 Contig42 m 41300 42163 100%
SEQ ID No. 2096 4161.3 Contig42 m 42188 43207 100%
SEQ ID No. 2934 5530.2 Contig42 P 43383 44126 100%
SEQ ID No. 2803 5256.2 Contig42 P 44544 44879 100%
SEQ ID No. 1877 3827.2 Contig42 m 45005 46999 100%
SEQ ID No. 1876 3826.1 Contig42 m 47150 48148 100%
SEQ ID No. 977 2457.2 Contig42 m 48149 48919 100%
SEQ ID No. 976 2456.2 Contig42 P 48965 49921 100%
SEQ ID No. 1875 3824.2 Contig42 P 49884 51209 100%
SEQ ID No. 1874 3822.2 Contig42 P 51526 52095 100%
SEQ ID No. 1873 3821.1 Contig42 P 52422 53600 100%
SEQ ID No. 880 2320.3 Contig42 P 53588 55039 99%
SEQ ID No. 879 2319.3 Contig42 m 55138 56562 100%
SEQ ID No. 941 2409.3 Contig42 P 56686 57420 + 100% -
SEQ ID No. 943 2410.2 Contig42 P 57574 58845 + 100% -
SEQ ID No. 1196 2795.1 Contig42 P 58861 59223 100% -
SEQ ID No. 2866 540.3 Contig42 P 59184 59567 100% -
SEQ ID No. 2862 539.3 Contig42 P 59488 60189 100% -
SEQ ID No. 2855 538.1 Contig42 P 60700 61350 99% +
SEQ ID No. 2850 537.2 Contig42 P 61594 62016 + 100% +
SEQ ID No. 1082 262.3 Contig42 m 62121 63479 100% -
SEQ ID No. 1089 263.1 Contig42 m 63539 65284 100% + -
SEQ ID No. 1093 264.2 Contig42 m 65185 66576 100% + .
SEQ ID No. 3331 829.2 Contig42 P 66690 68411 + 100% -
SEQ ID No. 1195 2794.2 Contig42 m 68614 70335 100% -
SEQ ID No. 1194 2793.1 Contig42 m 70547 71560 100% -
SEQ ID No. 1193 2792.1 Contig42 P 71391 71660 +
SEQ ID No. 1192 2791.1 Contig42 P 71734 74157 100% -
SEQ ID No. 1191 2788.1 Contig42 P 74393 75838 100% -
SEQ ID No. 1190 2785.1 Contig42 P 75829 76677 100% -
SEQ ID No. 623 1865.3 Contig42 P 76665 77765 100% -
SEQ ID No. 622 1864.3 Contig42 P 77689 79155 100% -
SEQ ID No. 1189 2784.3 Contig42 P 79238 79564 + 100% -
SEQ ID No. 2569 4841.2 Contig42 P 79724 80797 100% -
SEQ ID No. 2568 4837.1 Contig42 P 80754 81683 100% -
SEQ ID No. 2567 4836.1 Contig42 P 81668 82159 100% -
SEQ ID No. 2566 4835.4 Contig42 m 82095 82778 100% -
SEQ ID No. 2565 4834.4 Contig42 P 82282 82476 100% +
SEQ ID No. 3001 5650.3 Contig42 P 82949 83656 100% -
SEQ ID No. 3000 5649.1 Contig42 P 83688 85022 100% -
SEQ ID No. 2999 5648.1 Contig42 m 84306 84641 100% -
SEQ ID No. 2998 5647.1 Contig42 P 85391 85759 100% -
SEQ ID No. 2997 5645.2 Contig42 P 85779 88058 100% -
SEQ ID No. 3170 6108.1 Contig42 P 88359 88874
SEQ ID No. 342 1440.2 Contig43 P 2 1249 + 100% -
SEQ ID No. 70 1021.3 Contig43 P 1363 4413 100% -
SEQ ID No. 1519 3295.2 Contig43 P 4645 7437 100% -
SEQ ID No. 531 1733.3 Contig43 P 7541 8806 + 100% -
SEQ ID No. 1518 3291.2 Contig43 P 8871 9356 100% +
SEQ ID No. 1517 3290.1 Contig43 m 9457 10470 - 99%
SEQ ID No. 1516 3288.3 Contig43 P 10583 11272 - 97%
SEQ ID No. 1515 3285.3 Contig43 P 11250 12326 - 95%
SEQ ID No. 2358 4533.2 Contig43 m 12519 18197 + +
SEQ ID No. 2505 475.1 Contig43 P 18407 18700 + +
SEQ ID No. 2498 474.2 Contig43 P 18701 19033 + +
SEQ ID No. 2049 4101.1 Contig43 P 19176 19379 +
SEQ ID No. 2047 4098.1 Contig43 P 19655 20044 - +
SEQ ID No. 2046 4097.1 Contig43 P 20045 20302 - +
SEQ ID No. 2045 4096.1 Contig43 m 20278 20760 +
SEQ ID No. 983 2464.2 Contig43 m 20673 21026 +
SEQ ID No. 982 2462.2 Contig43 P 21047 21406 - +
SEQ ID No. 981 2461.2 Contig43 P 21548 21760 +
SEQ ID No. 2809 5273.1 Contig43 m 21791 22153 - +
SEQ ID No. 980 2460.3 Contig43 m 22069 23802 + 99%
SEQ ID No. 1463 3212.2 Contig43 m 23799 24497 + 98%
SEQ ID No. 74 1028.3 Contig43 m 24521 25792 - 97%
SEQ ID No. 76 1030.3 Contig43 m 25732 26601 - 96%
SEQ ID No. 77 1032.3 Contig43 P 26819 27790 - 99%
SEQ ID No. 78 1034.3 Contig43 P 27791 28777 + 96%
SEQ ID No. 762 2122.2 Contig43 P 28807 30006 + 98%
SEQ ID No. 293 1367.3 Contig43 m 30046 30639 + 97%
SEQ ID No. 295 1369.2 Contig43 m 30471 30911 - 97%
SEQ ID No. 294 1368.2 Contig43 P 30475 30666 96%
SEQ ID No. 296 1370.5 Contig43 m 31018 31629 - 99%
SEQ ID No. 1071 2599.4 Contig43 m 31671 33359 - 99% +
SEQ ID No. 1073 2600.4 Contig43 m 33387 34418 - 99% +
SEQ ID No. 1464 3217.3 Contig43 m 34496 35749 - 96%
SEQ ID No. 1465 3218.1 Contig43 P 35810 36511 -
SEQ ID No. 1467 3221.1 Contig43 P 36457 37890 - 100%
SEQ ID No. 1468 3223.1 Contig43 P 37875 38759 - 98% +
SEQ ID No. 1469 3224.3 Contig43 P 38946 41255 + 100% +
SEQ ID No. 491 1666.3 Contig43 P 41339 43054 - 96% +
SEQ ID No. 1348 304.2 Contig43 m 43061 45910 - 95% +
SEQ ID No. 1324 300.2 Contig43 P 46000 47781 - 98% SEQ ID No. 593 1819.6 Contig43 m 48139 52452 - 95%
SEQ ID No. 974 2450.3 Contig43 m 52511 54343 98%
SEQ ID No. 340 1438.4 Contig43 P 54447 56333 98%
SEQ ID No. 2783 521.2 Contig43 m 56667 58832 98%
SEQ ID No. 2776 520.1 Contig43 m 59041 59676 99%
SEQ ID No. 2771 519.3 Contig43 m 59604 61277 100%
SEQ ID No. 1235 2869.1 Contig43 m 61278 61583 100%
SEQ ID No. 1234 2868.1 Contig43 P 61643 63514 98%
SEQ ID No. 1233 2866.1 Contig43 P 63598 64923 97%
SEQ ID No. 1052 2570.2 Contig43 m 64993 66153 99%
SEQ ID No. 1053 2571.2 Contig43 m 66126 67073 97%
SEQ ID No. 1232 2863.1 Contig43 m 67045 67788 94%
SEQ ID No. 1231 2862.1 Contig43 m 67685 68956 96%
SEQ ID No. 156 1148.2 Contig43 m 69161 71929 98%
SEQ ID No. 541 1746.2 Contig43 P 71971 72792 91%
SEQ ID No. 542 1747.2 Contig43 m 73047 73409 100%
SEQ ID No. 1230 2861.1 Contig43 P 73575 74438 97%
SEQ ID No. 969 2443.2 Contig43 P 74407 75246 95%
SEQ ID No. 968 2442.2 Contig43 m 75266 76591 99%
SEQ ID No. 2502 4746.1 Contig43 m 76572 76862 97%
SEQ ID No. 2503 4748.1 Contig43 m 76881 77648 99%
SEQ ID No. 2504 4749.2 Contig43 m 77687 79966 98%
SEQ ID No. 1442 3172.2 Contig43 m 80019 80918 99%
SEQ ID No. 3066 5824.1 Contig43 m 80785 81201
SEQ ID No. 1443 3173.1 Contig43 P 81385 81654 98%
SEQ ID No. 1444 3174.1 Contig43 P 81648 82529 96%
SEQ ID No. 1445 3175.1 Contig43 P 82516 83301 94%
SEQ ID No. 1446 3177.1 Contig43 P 83277 83762 98%
SEQ ID No. 1447 3178.1 Contig43 P 83973 84629 98%
SEQ ID No. 1448 3179.1 Contig43 P 84611 86452 99%
SEQ ID No. 3169 6107.1 Contig43 m 86295 86489
SEQ ID No. 261 1314.3 Contig43 m 86777 88240 99%
SEQ ID No. 1450 3181.2 Contig43 m 88225 89379 99%
SEQ ID No. 2762 5167.4 Contig43 m 89396 89701 +
SEQ ID No. 3168 6106.1 Contig43 m 89715 89972 +
SEQ ID No. 1275 2938.3 Contig44 P 205 3858 98% +
SEQ ID No. 1274 2934.1 Contig44 m 3925 5202 97% +
SEQ ID No. 1273 2933.2 Contig44 m 5328 6305
SEQ ID No. 1272 2932.2 Contig44 m 6328 8805 +
SEQ ID No. 2833 5324.1 Contig44 m 8763 9419 +
SEQ ID No. 946 2414.2 Contig44 m 9382 10485 +
SEQ ID No. 945 2413.1 Contig44 m 10652 10867 +
SEQ ID No. 944 2412.2 Contig44 m 11014 12684 97%
SEQ ID No. 1074 2602.2 Contig44 m 12685 13578 100%
SEQ ID No. 1075 2604.3 Contig44 m 13579 15414 99%
SEQ ID No. 3281 755.3 Contig44 m 15517 16344 97%
SEQ ID No. 3280 754.2 Contig44 m 16437 17450 98%
SEQ ID No. 3279 753.2 Contig44 P 17613 18584 98%
SEQ ID No. 1270 2929.1 Contig44 P 18836 19348 99%
SEQ ID No. 1269 2928.2 Contig44 m 19362 20231 99%
SEQ ID No. 1268 2926.1 Contig44 P 20406 20564
SEQ ID No. 1267 2925.1 Contig44 P 20611 22620 99%
SEQ ID No. 1266 2924.3 Contig44 m 22704 24785 96%
SEQ ID No. 3065 5823.2 Contig44 P 24801 26072 99%
SEQ ID No. 845 226.4 Contig44 m 26153 27034 96%
SEQ ID No. 1634 3466.1 Contig44 P 27076 27321 +
SEQ ID No. 3064 5822.1 Contig44 P 27230 27472 +
SEQ ID No. 826 223.1 Contig44 m 27631 28530 +
SEQ ID No. 822 222.1 Contig44 P 28556 29101 +
SEQ ID No. 817 221.1 Contig44 P 29082 29993 +
SEQ ID No. 812 220.1 Contig44 P 29972 30685 +
SEQ ID No. 649 1906.2 Contig44 m 31080 32615 99%
SEQ ID No. 648 1905.2 Contig44 m 32600 33892 99%
SEQ ID No. 349 1454.2 Contig44 m 34019 35995 100%
SEQ ID No. 2846 536.3 Contig44 P 36111 38714 98%
SEQ ID No. 2835 533.2 Contig44 P 38923 39735 96%
SEQ ID No. 2830 532.2 Contig44 m 39785 40900 99%
SEQ ID No. 784 2151.2 Contig44 P 41026 42417 96%
SEQ ID No. 1456 3196.3 Contig44 P 42479 43987 99%
SEQ ID No. 884 2324.4 Contig44 P 44152 45291 98% +
SEQ ID No. 3063 5821.1 Contig44 P 45218 45577 +
SEQ ID No. 883 2323.2 Contig44 m 45462 46322 97% +
SEQ ID No. 882 2322.2 Contig44 m 46545 47288 98%
SEQ ID No. 1457 3198.1 Contig44 m 47234 47797 - 100%
SEQ ID No. 1458 3199.1 Contig44 m 48054 49808 - 97%
SEQ ID No. 3218 663.2 Contig44 P 49960 51945 - 98%
SEQ ID No. 3219 665.2 Contig44 m 51996 52349 - 98%
SEQ ID No. 3220 666.2 Contig44 P 52242 52985 - 100%
SEQ ID No. 1459 3201.1 Contig44 P 53078 55018 + - 100%
SEQ ID No. 3196 632.2 Contig44 P 54982 55926 - 98%
SEQ ID No. 3197 633.3 Contig44 P 55984 57357 - 100%
SEQ ID No. 2805 5266.2 Contig44 m 57427 60807 - 89%
SEQ ID No. 3029 5719.2 Contig44 P 60988 61779 - 98%
SEQ ID No. 1300 297.2 Contig44 P 61710 62090 - 99%
SEQ ID No. 1289 296.1 Contig44 P 62252 62626 + - 99%
SEQ ID No. 1281 295.1 Contig44 P 62864 63349 - 100%
SEQ ID No. 1276 294.1 Contig44 P 63372 64055 - 99% +
SEQ ID No. 1271 293.2 Contig44 P 64030 65301 - 98% +
SEQ ID No. 1263 292.1 Contig44 P 65323 65865 - 99% +
SEQ ID No. 1255 291.4 Contig44 P 65866 67161 - 99%
SEQ ID No. 66 1011.5 Contig44 P 67175 69529 - 98%
SEQ ID No. 65 1010.2 Contig44 P 69530 70564 - 97%
SEQ ID No. 64 1009.2 Contig44 P 70565 71083 - 98%
SEQ ID No. 63 1006.3 Contig44 P 71092 71754 + - 96%
SEQ ID No. 1742 3617.2 Contig44 P 71658 72077 - 100%
SEQ ID No. 1743 3618.2 Contig44 P 72070 74055 + - 99%
SEQ ID No. 318 1402.2 Contig44 P 74062 75576 - 100%
SEQ ID No. 317 1400.2 Contig44 P 75570 77072 - 99%
SEQ ID No. 1744 3619.1 Contig44 P 77256 77771 . - - 100%
SEQ ID No. 1745 3621.2 Contig44 P 77720 79261 - 99%
SEQ ID No. 530 1730.3 Contig44 P 79326 81956 - 99%
SEQ ID No. 1927 3920.1 Contig44 P 81957 82331 - 100%
SEQ ID No. 1926 3919.1 Contig44 P 82297 83229 - 98%
SEQ ID No. 1925 3917.1 Contig44 P 83374 83652 +
SEQ ID No. 3113 5957.1 Contig44 P 83695 84627 +
SEQ ID No. 1924 3916.3 Contig44 P 84579 85955 - 97%
SEQ ID No. 1017 2517.2 Contig44 P 86081 87181 + - 95%
SEQ ID No. 1018 2518.4 Contig44 m 87240 87869 - 99%
SEQ ID No. 1020 2520.4 Contig44 m 87791 88135 - 98%
SEQ ID No. 2691 5041.2 Contig44 P 88262 88798 - 100%
SEQ ID No. 2692 5042.1 Contig44 m 88860 89207 - 98%
SEQ ID No. 2693 5043.3 Contig44 P 88939 90213 + - 98%
SEQ ID No. 2708 5064.2 Contig44 P 90483 91244 - 97%
SEQ ID No. 2707 5062.3 Contig44 P 91473 92150 - 94%
SEQ ID No. 3062 5818.1 Contig44 m 91982 92296 +
SEQ ID No. 2142 4232.3 Contig44 m 92543 93277 - 97%
SEQ ID No. 2141 4231.2 Contig44 P 93455 94969 - 97% +
SEQ ID No. 3399 923.3 Contig44 m 95217 97235 - 97% +
SEQ ID No. 3398 922.2 Contig44 m 97211 98074 + - 98%
SEQ ID No. 3397 921.3 Contig44 m 98064 98678 - 98%
SEQ ID No. 3061 5816.1 Contig44 m 98666 99019 - 99%
SEQ ID No. 3338 84.4 Contig44 m 99038 100735 - 96%
SEQ ID No. 3327 82.2 Contig44 m 101521 101775 - 100%
SEQ ID No. 1157 2741.2 Contig44 m 102053 102259 +
SEQ ID No. 1158 2742.2 Contig44 P 102309 102515 +
SEQ ID No. 3305 79.1 Contig44 P 102657 102878 +
SEQ ID No. 3292 77.1 Contig44 P 103230 103448 + +
SEQ ID No. 3031 5720.1 Contig45 P 1 204 + +
SEQ ID No. 3032 5723.1 Contig45 P 218 523 +
SEQ ID No. 3033 5724.2 Contig45 P 651 2195 - 89%
SEQ ID No. 513 1703.1 Contig45 m 2232 3020 - 98%
SEQ ID No. 514 1704.4 Contig45 m 3189 5213 - 97%
SEQ ID No. 636 1887.2 Contig45 m 5447 7078 - 90%
SEQ ID No. 637 1889.2 Contig45 m 7392 8750 - 91 %
SEQ ID No. 1614 3432.2 Contig45 P 8936 10051 - 95%
SEQ ID No. 1613 3431.3 Contig45 P 10009 11346 - 98%
SEQ ID No. 839 2250.2 Contig45 P 11577 12230 + - 95% +
SEQ ID No. 1611 3429.1 Contig45 P 12616 14904 - 95% +
SEQ ID No. 1610 3428.1 Contig45 m 15087 15905 - 96%
SEQ ID No. 1609 3426.1 Contig45 m 16078 16782 - 97%
SEQ ID No. 1608 3423.3 Contig45 m 17254 18540 - 91 %
SEQ ID No. 1607 3421.2 Contig45 P 18675 20015 - 96% +
SEQ ID No. 1605 3419.3 Contig45 m 20115 20882 - 92% +
SEQ ID No. 2788 5224.2 Contig45 P 21155 21526 + +
SEQ ID No. 1216 2830.3 Contig45 P 21481 22104 + +
SEQ ID No. 1217 2831.1 Contig45 P 22331 23041 82%
SEQ ID No. 1218 2833.1 Contig45 P 23134 24129 91%
SEQ ID No. 508 1695.2 Contig45 m 24311 25426 92%
SEQ ID No. 507 1693.2 Contig45 P 25663 25947 91%
SEQ ID No. 1219 2835.1 Contig45 P 26014 26910 91%
SEQ ID No. 1220 2838.1 Contig45 P 26911 28107 92%
SEQ ID No. 1222 2840.1 Contig45 P 28121 29647 91%
SEQ ID No. 1223 2841.4 Contig45 P 29814 31133 95%
SEQ ID No. 288 1360.6 Contig45 m 31446 32963 96%
SEQ ID No. 289 1361.3 Contig45 P 33558 34631 98%
SEQ ID No. 246 1286.3 Contig45 P 34628 36538 98%
SEQ ID No. 245 1284.2 Contig45 P 36772 37059 96%
SEQ ID No. 244 1283.2 Contig45 P 37016 38341 98%
SEQ ID No. 1224 2847.4 Contig45 P 38461 39105 95% +
SEQ ID No. 1225 2848.4 Contig45 m 39163 39426 +
SEQ ID No. 1226 2849.1 Contig45 P 39757 40530 93% +
SEQ ID No. 711 2029.2 Contig45 m 41434 42672 95% +
SEQ ID No. 710 2027.2 Contig45 m 42859 44145
SEQ ID No. 709 2026.1 Contig45 P 44695 44994 80% +
SEQ ID No. 2102 417.3 Contig45 m 45095 45925 + +
SEQ ID No. 2087 415.2 Contig45 P 46532 47668 +
SEQ ID No. 2069 413.5 Contig45 m 48015 50384 +
SEQ ID No. 2623 4927.1 Contig45 m 50814 51575 +
SEQ ID No. 2624 4929.2 Contig45 m 51754 52683 98%
SEQ ID No. 2626 4930.2 Contig45 m 52692 52979 100%
SEQ ID No. 2404 4604.3 Contig45 m 53037 53615 99%
SEQ ID No. 3306 790.3 Contig45 m 53670 56318 95% +
SEQ ID No. 1259 2915.2 Contig45 m 56619 59432 97% +
SEQ ID No. 1915 390.2 Contig45 P 59678 60028 99%
SEQ ID No. 1907 389.3 Contig45 P 59965 61974 98%
SEQ ID No. 1258 2914.1 Contig45 P 61952 62623 97%
SEQ ID No. 482 1655.2 Contig45 P 62532 62795 100%
SEQ ID No. 483 1657.1 Contig45 P 62789 63544 98%
SEQ ID No. 3035 5730.1 Contig45 m 63648 63923 97% +
SEQ ID No. 484 1658.2 Contig45 P 63762 65222 98% +
SEQ ID No. 153 1142.4 Contig45 m 65328 66779 99%
SEQ ID No. 1257 2911.3 Contig45 m 66824 67198 99%
SEQ ID No. 795 2169.2 Contig45 P 67426 67854 + 97%
SEQ ID No. 796 2170.1 Contig45 P 68018 68461 + 97%
SEQ ID No. 3449 993.2 Contig45 m 68458 69726 99%
SEQ ID No. 3448 992.2 Contig45 P 68996 69385 100%
SEQ ID No. 3450 994.2 Contig45 m 69800 72601 + 99%
SEQ ID No. 1256 2910.1 Contig45 P 72535 74043 98%
SEQ ID No. 1254 2909.1 Contig45 P 74272 75552 98%
SEQ ID No. 1253 2908.2 Contig45 P 75581 76588 98%
SEQ ID No. 1252 2906.2 Contig45 P 76746 77390 97%
SEQ ID No. 1251 2905.2 Contig45 P 77708 79522 98%
SEQ ID No. 241 1279.3 Contig45 P 79527 80066 96%
SEQ ID No. 240 1278.2 Contig45 m 80071 81282 99%
SEQ ID No. 754 2108.2 Contig45 P 81438 82382 99%
SEQ ID No. 102 1069.2 Contig45 P 82510 83586 +
SEQ ID No. 101 1067.1 Contig45 P 83801 84055 +
SEQ ID No. 100 1066.2 Contig45 m 84221 85126 94%
SEQ ID No. 1735 3608.1 Contig45 m 85559 85888 100%
SEQ ID No. 1734 3607.2 Contig45 m 85993 87189 97%
SEQ ID No. 2763 5173.1 Contig45 m 87262 89307 96%
SEQ ID No. 1842 3765.1 Contig45 m 89250 90209 98%
SEQ ID No. 958 2429.2 Contig45 m 90197 91141 99%
SEQ ID No. 957 2428.2 Contig45 P 91233 91619 +
SEQ ID No. 956 2427.4 Contig45 m 91643 92011 +
SEQ ID No. 1841 3764.3 Contig45 P 92316 92675 +
SEQ ID No. 1840 3763.1 Contig45 P 92636 93379 +
SEQ ID No. 1839 3761.1 Contig45 P 93268 93459 +
SEQ ID No. 1838 3760.3 Contig45 m 93555 95363 + 99%
SEQ ID No. 2032 4075.3 Contig45 m 95380 95859 98%
SEQ ID No. 2033 4076.1 Contig45 m 96123 96674 95%
SEQ ID No. 534 1737.2 Contig45 P 96822 97934 +
SEQ ID No. 533 1735.1 Contig45 P 98103 98318 + +
SEQ ID No. 532 1734.2 Contig45 P 98319 98483 + +
SEQ ID No. 2034 4078.1 Contig45 m 98517 99893 97%
SEQ ID No. 2036 4080.2 Contig45 P 100247 101554 + 97%
SEQ ID No. 2764 5174.2 Contig45 m 101564 102016 + 99%
SEQ ID No. 496 1675.3 Contig45 m 102037 102357 99% SEQ ID No. 495 1673.5 Contig45 P 102412 104136 97% SEQ ID No. 492 1669.3 Contig45 m 104200 105201 98% SEQ ID No. 493 1670.3 Contig45 P 105391 106725 96% SEQ ID No. 2135 4222.3 Contig45 m 106907 107263 97% SEQ ID No. 2134 4221.3 Contig45 P 107390 107692 97% SEQ ID No. 2133 4220.2 Contig45 m 107870 108175 + SEQ ID No. 1884 384.3 Contig45 m 108189 109274 + SEQ ID No. 502 1685.3 Contig45 m 109460 110584 98% SEQ ID No. 3390 914.3 Contig45 m 110599 112401 98% SEQ ID No. 3391 915.3 Contig45 m 112344 113216 + 97% SEQ ID No. 2765 5176.1 Contig45 m 113642 113920 SEQ ID No. 2766 5177.2 Contig45 P 114007 114765 99% SEQ ID No. 3036 5735.2 Contig45 P 114761 115966 + 98% SEQ ID No. 2972 5598.2 Contig45 P 116104 117363 + 99% SEQ ID No. 3166 6098.1 Contig45 P 117531 117845 SEQ ID No. 2256 4387.3 Contig45 P 117746 118360 93% SEQ ID No. 2257 4388.1 Contig45 m 118391 119677 100% SEQ ID No. 87 1044.3 Contig45 m 119649 120962 99% SEQ ID No. 86 1042.5 Contig45 m 121153 122400 100% SEQ ID No. 2973 5600.2 Contig45 m 122515 122787 100% SEQ ID No. 3082 588.3 Contig45 m 122879 123670 96% SEQ ID No. 3076 587.2 Contig45 m 123642 124427 98% SEQ ID No. 3072 585.2 Contig45 m 124480 124830 99% SEQ ID No. 3068 583.2 Contig45 m 125324 125962 98% SEQ ID No. 3165 6097.1 Contig45 m 126102 134465 + SEQ ID No. 3164 6094.1 Contig46 m 94 366 + SEQ ID No. 2413 4620.1 Contig46 P 136 441 + SEQ ID No. 2411 4618.1 Contig46 P 588 1376 100% SEQ ID No. 2410 4616.2 Contig46 P 1343 2314 + 99% SEQ ID No. 3012 5672.1 Contig46 P . 2296 2640 + + SEQ ID No. 2409 4615.3 Contig46 m 2641 3060 100% + SEQ ID No. 2907 5479.2 Contig46 m 3146 3784 100% SEQ ID No. 1830 3747.2 Contig46 m 3742 4659 + 100% SEQ ID No. 1831 3748.1 Contig46 m 4653 6005 100% SEQ ID No. 1832 3750.2 Contig46 P 6131 7858 + 100%
SEQ ID No. 1833 3752.2 Contig46 P 7797 8543 + 100%
SEQ ID No. 1834 3753.2 Contig46 P 8556 10478 + 100%
SEQ ID No. 1835 3754.2 Contig46 P 10557 11339 + 100%
SEQ ID No. 1836 3756.2 Contig46 P 11314 12657 - 100%
SEQ ID No. 2306 4469.2 Contig46 m 12750 13640 - 100%
SEQ ID No. 3308 793.2 Contig46 m 13696 14010 100% +
SEQ ID No. 3309 794.2 Contig46 m 13863 14069 +
SEQ ID No. 3310 795.2 Contig46 m 14074 14865 - 100%
SEQ ID No. 2308 4470.1 Contig46 m 14843 15916 - 100%
SEQ ID No. 352 1457.2 Contig46 m 15855 16466 - 98%
SEQ ID No. 351 1456.2 Contig46 m 16467 17201 - 100%
SEQ ID No. 350 1455.2 Contig46 m 17189 17725 + 100%
SEQ ID No. 2309 4471.3 Contig46 m 17679 18281 + 100%
SEQ ID No. 3116 5978.2 Contig46 m 18245 18802 - 100%
SEQ ID No. 3095 5903.2 Contig46 m 18786 19751 - 100% +
SEQ ID No. 3024 5696.2 Contig46 P 20113 20913 - 100% +
SEQ ID No. 3023 5695.1 Contig46 P 20906 21688 - 100%
SEQ ID No. 3022 5694.2 Contig46 P 21689 22165 + 100%
SEQ ID No. 2912 5489.2 Contig46 P 22170 22781 + 100%
SEQ ID No. 2219 4341.3 Contig46 P 22769 23065 - 100%
SEQ ID No. 2220 4342.1 Contig46 P 23446 23700 -
SEQ ID No. 2221 4344.1 Contig46 P 23660 24961 - 100%
SEQ ID No. 2222 4345.1 Contig46 P 24955 25716 - 100%
SEQ ID No. 366 1484.2 Contig46 m 25905 26603 - 100%
SEQ ID No. 367 1485.4 Contig46 m 26579 27880 + 100%
SEQ ID No. 2573 4848.3 Contig46 m 27829 28851 + 100%
SEQ ID No. 2572 4847.2 Contig46 P 29223 29828 100%
SEQ ID No. 123 1096.3 Contig46 P 30003 31439 - 100%
SEQ ID No. 122 1094.1 Contig46 m 31483 31767 - 100%
SEQ ID No. 121 1093.3 Contig46 m 31987 32967 - 100%
SEQ ID No. 2376 4564.2 Contig46 P 32890 33672 - 100%
SEQ ID No. 1069 2597.4 Contig46 P 33657 34349 - 100%
SEQ ID No. 1433 3159.2 Contig46 P 34524 35318 + 100%
SEQ ID No. 1432 3158.1 Contig46 P 35182 35460 -
SEQ ID No. 1431 3157.1 Contig46 P 35466 35897 + 100%
SEQ ID No. 1430 3153.3 Contig46 P 35885 37846 + 100%
SEQ ID No. 1429 3152.1 Contig46 P 37789 38376 + 100%
SEQ ID No. 744 2085.2 Contig46 P 38340 38777 + 100%
SEQ ID No. 743 2083.2 Contig46 P 38725 39456 + 100%
SEQ ID No. 742 2082.2 Contig46 P 39407 39649 - 100%
SEQ ID No. 614 1853.2 Contig46 P 39690 40109 + 100%
SEQ ID No. 3454 998.3 Contig46 m 40865 42739 - 100% +
SEQ ID No. 3455 999.2 Contig46 P 42796 43959 - 100% +
SEQ ID No. 58 1000.2 Contig46 P 43955 44752 - 100% +
SEQ ID No. 406 1542.2 Contig46 P 44743 45813 - 100%
SEQ ID No. 404 1539.2 Contig46 m 45979 46878 + 100%
SEQ ID No. 209 1230.2 Contig46 P 47057 48649 - 100%
SEQ ID No. 210 1231.3 Contig46 m 48715 49140 - 100%
SEQ ID No. 2522 4773.3 Contig46 m 49233 50660 + 100%
SEQ ID No. 2523 4774.1 Contig46 m 50654 50962 - 100%
SEQ ID No. 2524 4775.2 Contig46 m 50943 52103 - 100%
SEQ ID No. 2525 4776.3 Contig46 m 52470 53033 - 100%
SEQ ID No. 3163 6093.1 Contig46 m 53278 53583 - 100%
SEQ ID No. 728 2060.3 Contig46 P 53835 54983 - 100%
SEQ ID No. 729 2062.2 Contig46 m 55045 55689 + 100% +
SEQ ID No. 731 2065.2 Contig46 P 56118 56603 - 100% +
SEQ ID No. 730 2064.3 Contig46 m 56324 56575 - 100% +
SEQ ID No. 732 2066.5 Contig46 P 56724 57173 - 100% +
SEQ ID No. 733 2067.5 Contig46 P 57300 58376 - 100%
SEQ ID No. 2026 4068.2 Contig46 P 58873 60171 - 100%
SEQ ID No. 2025 4067.2 Contig46 m 59556 59759 100%
SEQ ID No. 2190 4303.1 Contig46 m 60495 61640 - 99%
SEQ ID No. 2189 4302.1 Contig46 m 61621 62454 - 100%
SEQ ID No. 2188 4301.1 Contig46 m 62455 63360 - 100%
SEQ ID No. 332 1420.2 Contig46 m 63415 64569 - 100%
SEQ ID No. 333 1421.3 Contig46 P 64694 67030 - 100%
SEQ ID No. 2888 5446.3 Contig46 P 67332 68681 - 100%
SEQ ID No. 2889 5447.1 Contig46 P 68866 69651 - 100%
SEQ ID No. 600 1834.4 Contig46 P 69560 70978 - 100%
SEQ ID No. 3117 5979.1 Contig46 m 70998 71240 100%
SEQ ID No. 2890 5448.1 Contig46 P 71292 71837 - 100%
SEQ ID No. 2380 4571.2 Contig46 P 71819 72220 - 100%
SEQ ID No. 1055 2576.2 Contig46 m 72302 73279 100%
SEQ ID No. 1056 2578.2 Contig46 m 73534 74286 100%
SEQ ID No. 2381 4572.2 Contig46 m 74477 75556 100%
SEQ ID No. 2382 4573.4 Contig46 m 75541 77133 100%
SEQ ID No. 3096 5904.2 Contig46 P 77481 78152 100%
SEQ ID No. 2673 5010.2 Contig46 P 78136 79059 100%
SEQ ID No. 2674 5015.2 Contig46 P 79272 79613 100%
SEQ ID No. 2896 5460.1 Contig46 P 79588 80679 99%
SEQ ID No. 1959 3974.2 Contig46 P 80784 81344 100%
SEQ ID No. 1958 3972.2 Contig46 P 81539 82297 100%
SEQ ID No. 1957 3971.2 Contig46 m 82354 82878 100%
SEQ ID No. 1956 3970.1 Contig46 m 82868 83266 100%
SEQ ID No. 1954 3969.1 Contig46 m 83279 83998 100%
SEQ ID No. 1953 3968.1 Contig46 P 84058 84474 100%
SEQ ID No. 1952 3967.1 Contig46 P 84548 85108 100%
SEQ ID No. 1951 3966.1 Contig46 P 85386 86165 100%
SEQ ID No. 1950 3965.1 Contig46 P 86726 87241 100%
SEQ ID No. 1949 3964.1 Contig46 P 87172 88179 100%
SEQ ID No. 1948 3962.2 Contig46 P 88158 88514 100%
SEQ ID No. 2897 5462.2 Contig46 P 88889 90562 99%
SEQ ID No. 3205 643.3 Contig46 P 90559 92034 100%
SEQ ID No. 3206 644.2 Contig46 m 92079 93134 100%
SEQ ID No. 3207 645.4 Contig46 m 93128 93838 100%
SEQ ID No. 3208 647.5 Contig46 P 94033 94683 100%,
SEQ ID No. 2579 4859.2 Contig46 P 94693 95967 99% +
SEQ ID No. 2581 4860.1 Contig46 P 95971 96123 +
SEQ ID No. 2924 5515.2 Contig46 P 96370 97125 99%
SEQ ID No. 1939 3942.2 Contig46 P 97130 98077 99%
SEQ ID No. 3191 623.2 Contig46 P 98038 99210 98%
SEQ ID No. 3190 622.3 Contig46 m 99282 101681 99%
SEQ ID No. 2923 5514.2 Contig46 m 101816 102838 89%
SEQ ID No. 2865 5398.3 Contig46 P 103240 104304 99%
SEQ ID No. 1697 3555.2 Contig46 P 104305 104856 99%
SEQ ID No. 1696 3554.4 Contig46 P 104831 105469 99%
SEQ ID No. 642 1895.6 Contig46 P 105421 106050 96%
SEQ ID No. 643 1896.5 Contig46 P 106051 108150 99%
SEQ ID No. 1695 3551.1 Contig46 P 108590 109123 100%
SEQ ID No. 1694 3550.1 Contig46 P 109101 110219 98%
SEQ ID No. 1693 3548.2 Contig46 P 110195 111667 97%
SEQ ID No. 1692 3547.1 Contig46 P 111746 112207 99%
SEQ ID No. 1691 3545.1 Contig46 P 112319 113317 98%
SEQ ID No. 370 1490.4 Contig46 P 113424 116225 99%
SEQ ID No. 369 1488.2 Contig46 m 114650 114994 98%
SEQ ID No. 830 2233.3 Contig46 P 116207 116686 98%
SEQ ID No. 829 2232.1 Contig46 m 116985 117551 98%
SEQ ID No. 242 1280.3 Contig46 m 117725 119929 98% +
SEQ ID No. 292 1366.3 Contig46 m 120156 123287 97% +
SEQ ID No. 1750 3629.2 Contig46 P 123454 124431 99%
SEQ ID No. 1455 3193.2 Contig46 P 124409 125110 98%
SEQ ID No. 1454 3191.1 Contig46 m 125166 126344 97%
SEQ ID No. 1453 3190.1 Contig46 m 126503 127390 94%
SEQ ID No. 788 2159.2 Contig46 P 127607 128428 99%
SEQ ID No. 455 1615.2 Contig46 P 128397 130232 99% +
SEQ ID No. 790 2160.2 Contig46 P 130192 131184 98% +
SEQ ID No. 2111 418.4 Contig46 m 131572 132249 100%
SEQ ID No. 2116 419.2 Contig46 P 132432 133271 98%
SEQ ID No. 2128 421.2 Contig46 P 133283 134725 99%
SEQ ID No. 2132 422.3 Contig46 P 134726 135649 98%
SEQ ID No. 605 1841.2 Contig46 P 135667 137343 98%
SEQ ID No. 1452 3186.2 Contig46 m 137322 140789 99%
SEQ ID No. 1451 3184.3 Contig46 m 140860 142122 99%
SEQ ID No. 89 1048.3 Contig46 m 142126 143316 99%
SEQ ID No. 88 1047.2 Contig46 m 143301 144233 98%
SEQ ID No. 886 2330.2 Contig46 P 144380 146443 98%
SEQ ID No. 887 2333.3 Contig46 P 146545 148728 98%
SEQ ID No. 2683 5030.2 Contig46 P 148587 149702 99%
SEQ ID No. 1931 3927.2 Contig46 P 149681 151072 98%
SEQ ID No. 469 1636.2 Contig46 P 151141 154593 99%
SEQ ID No. 1930 3925.2 Contig46 m 154679 155944 99%
SEQ ID No. 1929 3924.2 Contig46 P 156244 157134 98%
SEQ ID No. 243 1282.3 Contig46 P 157074 158642 97%
SEQ ID No. 1928 3923.2 Contig46 P 158658 160574 99%
SEQ ID No. 2570 4844.2 Contig46 m 160720 161328 93% +
SEQ ID No. 2571 4845.4 Contig46 m 161597 163333 96% +
SEQ ID No. 2161 4255.1 Contig46 m 163658 164020 100% +
SEQ ID No. 2160 4254.2 Contig46 P 163752 164525 96% +
SEQ ID No. 2159 4253.1 Contig46 m 164822 166306 100%
SEQ ID No. 873 2309.2 Contig46 P 166657 167811 97%
SEQ ID No. 872 2308.2 Contig46 P 168348 168671 +
SEQ ID No. 3162 6089.1 Contig46 m 168652 169203 +
SEQ ID No. 3120 5985.1 Contig47 m 21 398 +
SEQ ID No. 3083 5890.1 Contig47 m 596 778 +
SEQ ID No. 1768 3653.3 Contig47 m 801 1010 +
SEQ ID No. 1769 3655.1 Contig47 m 1280 1993 + +
SEQ ID No. 2733 510.2 Contig47 m 1971 2675 + +
SEQ ID No. 2726 509.1 Contig47 m 2774 3367 +
SEQ ID No. 2713 507.3 Contig47 m 3397 3723 93%
SEQ ID No. 2704 506.3 Contig47 P 4108 4518 + +
SEQ ID No. 2697 505.3 Contig47 m 4559 5503 + +
SEQ ID No. 1770 3657.1 Contig47 m 5811 6452 +
SEQ ID No. 1771 3658.1 Contig47 P 6810 7100 +
SEQ ID No. 3417 948.3 Contig47 P 7066 7596 +
SEQ ID No. 3416 947.3 Contig47 m 7565 7927 +
SEQ ID No. 3415 946.3 Contig47 m 7965 8639 +
SEQ ID No. 1772 3659.2 Contig47 m 8757 9299 +
SEQ ID No. 1774 3661.4 Contig47 m 9263 10255 +
SEQ ID No. 2984 5620.3 Contig47 P 10879 11130 96%
SEQ ID No. 786 2153.3 Contig47 P 11124 11363 + +
SEQ ID No. 2163 426.3 Contig47 P 11669 12370 + +
SEQ ID No. 2156 425.1 Contig47 m 12661 12954 + +
SEQ ID No. 2148 424.1 Contig47 m 12935 13252 96%
SEQ ID No. 2140 423.2 Contig47 m 13262 14566 +
SEQ ID No. 2985 5621.1 Contig47 m 14589 15851 + +
SEQ ID No. 2986 5623.2 Contig47 m 16093 17499 + +
SEQ ID No. 1909 3891.2 Contig47 m 17492 18145 +
SEQ ID No. 1908 3890.2 Contig47 P 18284 19642 +
SEQ ID No. 1906 3889.1 Contig47 m 19632 20417 +
SEQ ID No. 1905 3888.1 Contig47 m 20390 20593 +
SEQ ID No. 1904 3887.1 Contig47 m 20607 21833 +
SEQ ID No. 1903 3884.2 Contig47 P 22460 22942 96% +
SEQ ID No. 338 1432.3 Contig47 P 23126 24049 98%
SEQ ID No. 337 1429.4 Contig47 m 24232 25806 + + +
SEQ ID No. 2868 5404.2 Contig47 m 26133 27047 + +
SEQ ID No. 2867 5402.1 Contig47 P 27185 28633 86% +
SEQ ID No. 1028 2533.3 Contig47 P 28867 29787 84%
SEQ ID No. 1027 2530.3 Contig47 P 29880 31814 95%
SEQ ID No. 1733 3602.2 Contig47 P 31894 33795 87%
SEQ ID No. 1732 3601.1 Contig47 P 33973 35130 +
SEQ ID No. 1731 3600.2 Contig47 P 35321 36292 +
SEQ ID No. 975 2453.4 Contig47 P 36235 39030 +
SEQ ID No. 480 1652.4 Contig47 P 39121 39687 91%
SEQ ID No. 481 1653.3 Contig47 m 39802 40740 + 96%
SEQ ID No. 3148 606.3 Contig47 m 40856 42601 + 95%
SEQ ID No. 3174 612.2 Contig47 P 42746 44959 96%
SEQ ID No. 740 2079.2 Contig47 m 45270 46547 93% +
SEQ ID No. 739 2078.2 Contig47 m 46681 46965 +
SEQ ID No. 1783 3673.2 Contig47 m 47107 48360 94%
SEQ ID No. 3057 5809.1 Contig47 P 48148 48435 79% +
SEQ ID No. 2617 4917.2 Contig47 m 48380 49735 94% +
SEQ ID No. 2616 4916.2 Contig47 P 49917 50837 95%
SEQ ID No. 3004 5655.2 Contig47 m 50958 51446 99%
SEQ ID No. 3003 5654.2 Contig47 P 51771 52181 94%
SEQ ID No. 2628 4935.2 Contig47 m 52309 53451 97%
SEQ ID No. 2627 4934.3 Contig47 m 53749 54948 98%
SEQ ID No. 3056 5808.1 Contig47 m 55003 55581 97%
SEQ ID No. 3427 964.3 Contig47 m 55765 57123 90%
SEQ ID No. 3421 953.2 Contig47 P 57651 59225 89%
SEQ ID No. 3420 952.1 Contig47 m 59202 60149 98%
SEQ ID No. 3419 950.2 Contig47 m 60306 61823 85%
SEQ ID No. 1493 3252.1 Contig47 P 62076 63032 98%
SEQ ID No. 1492 3251.1 Contig47 P 62996 63430 99%
SEQ ID No. 3383 901.2 Contig47 m 63514 64692 87% +
SEQ ID No. 3384 903.2 Contig47 P 64977 66098 94% +
SEQ ID No. 3385 905.2 Contig47 m 66185 67045 +
SEQ ID No. 1491 3250.1 Contig47 m 67309 67623 96% SEQ ID No. 1489 3249.2 Contig47 P 67801 68496 98% SEQ ID No. 418 1562.3 Cont g47 P 68820 69641 97% + SEQ ID No. 3161 6087.1 Contig47 m 69577 69876 + + SEQ ID No. 417 1560.2 Cont g47 m 69761 70600 + SEQ ID No. 1488 3248.1 Contig47 m 70560 71483 + SEQ ID No. 1487 3246.2 Contig47 P 71805 73802 95% SEQ ID No. 619 1861.2 Contig47 m 73905 74735 96% SEQ ID No. 618 1860.2 Contig47 P 75237 76346 92% SEQ ID No. 1546 3336.2 Contig47 P 76579 77034 93% + SEQ ID No. 1547 3337.2 Contig47 m 77247 77543 90% + SEQ ID No. 1548 3338.1 Contig47 P 77766 78158 + + SEQ ID No. 1550 3341.2 Cont g47 P 78359 81394 + + SEQ ID No. 1551 3343.1 Contig47 m 81608 82249 + + SEQ ID No. 1552 3345.2 Contig47 m 82458 84782 98% + SEQ ID No. 1062 2585.2 Cont g47 P 84972 86738 95% + SEQ ID No. 422 1567.2 Contig47 m . 86827 88173 92% + SEQ ID No. 421 1566.3 Contig47 m 88505 89068 SEQ ID No. 490 1664.2 Cont g47 m 89346 90161 87% + SEQ ID No. 2818 5297.2 Cont g47 P 90488 91873 + SEQ ID No. 2345 4519.3 Contig47 P 92237 93889 97% SEQ ID No. 3438 979.3 Cont g47 P 93871 95094 94% SEQ ID No. 3439 980.1 Conti g47 P 95095 95814 98% SEQ ID No. 3440 981.2 Conti g47 P 95787 96434 84% SEQ ID No. 2344 4517.2 Conti g47 P 96391 97248 92% SEQ ID No. 2558 4819.2 Contig47 P 97143 98192 93% SEQ ID No. 2560 4821.2 Contig47 P 98266 99219 94% SEQ ID No. 2575 4850.2 Contig47 P 99385 101181 93% SEQ ID No. 2576 4851.2 Contig47 P 101415 101942 94% SEQ ID No. 897 2347.2 Contig47 P 101957 103012 92% + SEQ ID No. 896 2346.1 Contig47 P 103046 103429 91 % + SEQ ID No. 895 2345.2 Cont g47 m 103536 103991 96% + SEQ ID No. 275 1336.3 Contig47 m 104195 105385 98% SEQ ID No. 2136 4225.1 Cont g47 P 105500 105769 94% SEQ ID No. 881 2321.3 Contig47 m 105652 106263 98% SEQ ID No. 2871 5412.1 Cont g47 P 106268 107311 97%
SEQ ID No.2395 4591.2 Contig47 P 107343 107822 94% SEQ ID No. 2394 4590.1 Contig47 P 107862 108332 100% SEQ ID No. 2392 4589.1 Contig47 m 108456 109088 96% + SEQ ID No. 2391 4588.2 Cont g47 m 109240 109881 99% + SEQ ID No. 2390 4587.2 Cont g47 P 109974 110486 97% + SEQ ID No. 2872 5413.1 Contig47 P 110505 110849 100% + SEQ ID No. 949 2419.3 Cont g47 P 110960 111712 98% SEQ ID No. 948 2418.2 Cont g47 P 111783 112523 97% SEQ ID No. 947 2415.2 Cont g47 m 112603 113577 95% SEQ ID No. 2103 4170.1 Contig47 m 113730 116510 97% SEQ ID No. 2101 4168.2 Contig47 P 116698 117213 98% + SEQ ID No. 2529 4780.2 Contig47 m 117261 117761 95% + SEQ ID No. 2528 4779.1 Cont g47 P 118015 118479 97% SEQ ID No. 2527 4778.1 Contig47 P 118604 119269 88% + SEQ ID No. 2526 4777.2 Contig47 P 119435 120775 96% + SEQ ID No. 2873 5417.1 Contig47 m 120900 121331 99% SEQ ID No. 2233 4357.3 Contig47 m 121371 122663 97% SEQ ID No. 2234 4358.2 Cont g47 m 122641 123207 96% SEQ ID No. 2964 5586.1 Contig47 m 123420 123806 96% SEQ ID No. 998 2485.3 Contig47 P 123935 124582 97% SEQ ID No. 629 1878.3 Contig47 P 124766 125719 98% SEQ ID No. 628 1875.2 Contig47 P 125840 127069 SEQ ID No. 2236 4363.2 Contig47 P 127205 127927 98% SEQ ID No. 2717 5076.2 Cont g47 m 127987 128469 87% SEQ ID No. 2718 5077.3 Contig47 m 128463 129851 99% SEQ ID No. 1767 3652.2 Contig47 m 129748 130578 98% SEQ ID No. 1766 3651.1 Contig47 m 130551 131399 98% SEQ ID No. 1765 3650.1 Contig47 m 131389 132657 97% SEQ ID No. 1011 2506.2 Cont g47 m 132564 133628 98% SEQ ID No. 1012 2508.3 Contig47 m 133619 134782 98% SEQ ID No. 1763 3647.2 Contig47 m 134722 134958 SEQ ID No. 264 1320.3 Cont g47 m 134959 137187 97% SEQ ID No. 1762 3645.3 Contig47 m 137198 137989 94% SEQ ID No. 3055 5807.1 Cont g47 m 137956 138348 99% SEQ ID No. 2471 4704.2 Contig47 m 138309 139100 98% SEQ ID No. 2470 4703.2 Contig47 P 139054 139539
SEQ ID No. 2469 4702.1 Contig47 P 139494 140090 94%
SEQ ID No. 2468 4701.2 Contig47 m 140087 140926 93%
SEQ ID No. 2507 4752.2 Contig47 m 141084 142007 98%
SEQ ID No. 2508 4754.1 Contig47 P 142255 143601 97%
SEQ ID No. 2509 4755.2 Contig47 m 143628 144191 98%
SEQ ID No. 2024 4066.2 Contig47 P 144222 144818 96%
SEQ ID No. 2023 4065.3 Contig47 P 144790 146553 98%
SEQ ID No. 2022 4063.1 Contig47 m 146659 148047 97%
SEQ ID No. 2021 4061.1 Contig47 m 148048 148587 99%
SEQ ID No. 2020 4060.1 Contig47 m 148533 149945 97%
SEQ ID No. 488 1661.3 Contig47 P 149794 150489 98%
SEQ ID No. 487 1660.2 Contig47 P 150510 150761 89% +
SEQ ID No. 485 1659.4 Contig47 P 150829 154725 96% +
SEQ ID No. 835 2242.1 Contig47 m 154849 155424 98% +
SEQ ID No. 312 1394.2 Contig47 m 155492 155962
SEQ ID No. 313 1396.3 Contig47 P 156048 157058 98%
SEQ ID No. 314 1397.2 Contig47 P 157308 157814 99%
SEQ ID No. 1867 3811.1 Contig47 m 157863 158390
SEQ ID No. 1866 3810.1 Contig47 P 158615 159328 97%
SEQ ID No. 587 1807.3 Contig47 m 159379 160806 100%
SEQ ID No. 586 1806.2 Contig47 P 160979 161524 99%
SEQ ID No. 585 1805.3 Contig47 m 161682 162344 98%
SEQ ID No. 2368 4551.2 Contig47 m 162316 162594 100%
SEQ ID No. 2369 4552.4 Contig47 P 162799 163758 89% +
SEQ ID No. 2844 5354.3 Contig47 P 164077 164319 98% +
SEQ ID No. 356 1467.5 Contig47 P 164422 165021 98%
SEQ ID No. 355 1466.2 Contig47 P 165058 165801 98%
SEQ ID No. 2430 4645.2 Contig47 P 165886 167118 99%
SEQ ID No. 2429 4644.3 Contig47 m 167115 168464 95% +
SEQ ID No. 2126 4206.2 Contig47 P 168685 171516 97% +
SEQ ID No. 2125 4205.2 Contig47 P 171437 172408 91 % +
SEQ ID No. 2124 4203.2 Contig47 P 172808 174076 100%
SEQ ID No. 2123 4200.2 Contig47 m 174124 174822 99% +
SEQ ID No. 301 1377.5 Contig47 m 175216 177651 97% +
SEQ ID No. 302 1378.5 Contig47 m 177711 178910 98% +
SEQ ID No. 2843 5349.3 Contig47 P 179148 181841 +
SEQ ID No. 2286 4431.2 Contig47 P 181989 184718 - 91 %
SEQ ID No. 2746 5129.4 Contig47 m 185037 188228 + - 99%
SEQ ID No. 2589 4873.3 Contig47 m 188249 189448 _ 98%
SEQ ID No. 2588 4872.3 Contig47 m 189409 190986 + _ 98%
SEQ ID No. 3160 6086.1 Contig47 m 191153 191521 . 98%
SEQ ID No. 3016 5685.2 Contig47 m 191522 192799 _ 98%
SEQ ID No. 3017 5686.2 Contig47 P 192951 193427 _ 98%
SEQ ID No. 3018 5687.1 Contig47 P 193621 193965 _ 99% +
SEQ ID No. 3019 5688.2 Contig47 P 194152 194715 - 79% +
SEQ ID No. 2958 5569.1 Contig47 m 194808 195188 + _ 98% +
SEQ ID No. 2957 5566.1 Contig47 P 195344 195577 + +
SEQ ID No. 99 1065.3 Contig47 m 195618 196814 _ 96%
SEQ ID No. 98 1063.1 Contig47 m 197173 198501 _ 98% +
SEQ ID No. 2664 4998.2 Contig47 m 198880 200607 - 96% +
SEQ ID No. 2663 4995.1 Contig47 P 200711 201430 . 98% +
SEQ ID No. 1604 3418.1 Contig48 P 1 852 - 96% +
SEQ ID No. 723 2053.2 Contig48 P 913 1896 + . 98% +
SEQ ID No. 2379 457.2 Contig48 P 2184 3791 _ 94%
SEQ ID No. 2362 454.1 Contig48 P 3988 5505 _ 93%
SEQ ID No. 2354 453.3 Contig48 P 5528 6304 _ 98%
SEQ ID No. 1603 3417.1 Contig48 P 6528 7109 _ 100%
SEQ ID No. 1602 3416.1 Contιg48 m 7125 7490 _ 100%
SEQ ID No. 1601 3415.2 Contig48 m 7689 8420 _ 97%
SEQ ID No. 1782 3671.2 Contig48 m 8410 9171 _ 96%
SEQ ID No. 1781 3670.2 Contig48 P 9712 10764 _ 97%
SEQ ID No. 1780 3669.1 Contig48 m 10859 11821 _ 99%
SEQ ID No. 1779 3667.1 Contig48 P 12363 12653 . 100%
SEQ ID No. 1778 3666.2 Contig48 P 12768 14036 +
SEQ ID No. 1777 3665.1 Contig48 P 14015 14953 +
SEQ ID No. 1776 3664.1 Contig48 P 15069 16226 +
SEQ ID No. 1775 3663.1 Contig48 P 16350 17441 +
SEQ ID No. 900 2351.2 Contig48 P 17599 18432 . 94%
SEQ ID No. 899 2350.2 Contig48 m 17731 18108 _ 83%
SEQ ID No. 901 2352.2 Contig48 P 18433 19479 _ 88%
SEQ ID No. 902 2353.3 Contig48 m 19451 20626 - 95%
SEQ ID No. 2153 4247.2 Contig48 P 20746 21615 - 76%
SEQ ID No. 111 1080.2 Contig48 P 21518 22444 +
SEQ ID No. 112 1081.2 Contig48 m 22650 23873 +
SEQ ID No. 2154 4248.1 Contig48 P 24264 24533 +
SEQ ID No. 2155 4249.1 Contig48 P 24436 25785 96%
SEQ ID No. 2157 4250.1 Contig48 m 25684 26193 + +
SEQ ID No. 2158 4251.2 Contig48 m 26261 27592 + +
SEQ ID No. 639 1891.2 Contig48 m 27597 28370 +
SEQ ID No. 756 2112.2 Contig48 P 28628 30067 +
SEQ ID No. 3337 838.3 Contig48 m 30280 31686 99%
SEQ ID No. 3339 840.2 Contig48 P 31793 32653 95%
SEQ ID No. 3340 841.3 Contig48 P 32857 33192 100%
SEQ ID No. 806 2193.4 Contig48 P 33494 34777 100%
SEQ ID No. 807 2195.1 Contig48 P 34920 36389 99%
SEQ ID No. 808 2196.1 Contig48 P 36362 37102 99%
SEQ ID No. 809 2197.1 Contig48 P 37090 37314
SEQ ID No. 810 2198.1 Contig48 m 37352 37684 99%
SEQ ID No. 811 2199.2 Contig48 m 37725 38468 98%
SEQ ID No. 1197 2799.1 Contig48 m 38556 38960 99% +
SEQ ID No. 1198 2800.1 Contig48 P 39290 40063 97% +
SEQ ID No. 689 1986.2 Contig48 P 40082 41437 97%
SEQ ID No. 688 1983.1 Contig48 P 41412 42260 99%
SEQ ID No. 2035 408.3 Contig48 P 42286 43932 + 99%
SEQ ID No. 2042 409.2 Contig48 P 43914 45074 98%
SEQ ID No. 2048 410.1 Contig48 P 45028 45756 99%
SEQ ID No. 2063 412.3 Contig48 P 45684 46946 + 98%
SEQ ID No. 919 2373.2 Contig48 P 47260 48243 + 99%
SEQ ID No. 920 2374.2 Contig48 P 48553 49464 +
SEQ ID No. 590 1815.2 Contig48 P 49732 50319 +
SEQ ID No. 591 1817.2 Contig48 P 50616 51212 98%
SEQ ID No. 2782 5208.1 Contig48 P 51661 52008 +
SEQ ID No. 1329 3005.2 Contig48 P 51996 52523 +
SEQ ID No. 1462 321.3 Contig48 P 53567 54463 +
SEQ ID No. 1466 322.3 Contig48 P 54569 54796 + +
SEQ ID No. 1472 323.2 Contig48 m 54848 56335 97% +
SEQ ID No. 719 2044.2 Contig48 P 57557 59509 98%
SEQ ID No. 523 172.1 Contig48 m 59557 59775
SEQ ID No. 537 174.1 Contig48 P 60382 61275 - 96%
SEQ ID No. 551 176.1 Contig48 m 61299 62291 - 96% +
SEQ ID No. 566 178.1 Contig48 P 62390 62704 - 100% +
SEQ ID No. 574 179.1 Contig48 m 62747 63022 - 95%
SEQ ID No. 581 180.1 Contig48 m 63013 63207 + +
SEQ ID No. 594 182.3 Contig48 m 63200 64855 - 99%
SEQ ID No. 1330 3008.1 Contig48 m 64939 65592 - 98%
SEQ ID No. 1059 2580.2 Contig48 m 65564 66331 - 99%
SEQ ID No. 1057 2579.2 Contig48 m 66441 66896 - 98%
SEQ ID No. 1331 3009.2 Contig48 m 67167 68123 + +
SEQ ID No. 2365 4546.3 Contig48 P 68632 69690 +
SEQ ID No. 916 2369.3 Contig48 m 69875 70516 + - 100%
SEQ ID No. 917 2371.1 Contig48 m 70539 71579 - 97%
SEQ ID No. 918 2372.3 Contig48 m 71633 72022 - 97%
SEQ ID No. 2364 4542.4 Contig48 P 72350 73981 + - 99%
SEQ ID No. 503 1689.3 Contig48 P 74054 75268 + - 100%
SEQ ID No. 505 1691.3 Contig48 P 75250 76881 - 99%
SEQ ID No. 546 1752.2 Contig48 P 76882 77436 + - 100%
SEQ ID No. 545 1750.3 Contig48 P 77589 78464 - 97%
SEQ ID No. 1243 2884.1 Contig48 m 78487 78768 - 94% +
SEQ ID No. 152 114.2 Contig48 P 79316 79507 + +
SEQ ID No. 157 115.2 Contig48 P 79533 80096 +
SEQ ID No. 163 116.1 Contig48 P 80429 80992 +
SEQ ID No. 3159 6083.1 Contig48 P 81259 81510 + +
SEQ ID No. 170 117.1 Contig48 P 81479 81859 + +
SEQ ID No. 176 118.1 Contig48 P 81813 82490 + +
SEQ ID No. 183 119.1 Contig48 P 82812 83327 +
SEQ ID No. 194 121.1 Contig48 m 83648 84016 +
SEQ ID No. 3043 5765.1 Contig48 m 84091 84564 +
SEQ ID No. 199 122.3 Contig48 m 84504 85001 + - 100%
SEQ ID No. 687 1979.1 Contig48 m 85012 85803 - 99%
SEQ ID No. 686 1978.1 Contig48 m 85785 86507 - 97%
SEQ ID No. 685 1976.2 Contig48 m 86408 88291 - 98%
SEQ ID No. 793 2166.2 Contig48 m 88413 90341 - 97%
SEQ ID No. 794 2167.2 Contig48 m 90554 92077 - 99%
SEQ ID No. 1242 2882.2 Contig48 P 92029 92766 + - 99%
SEQ ID No. 3044 5766.1 Contig48 P 92800 93363 98%
SEQ ID No. 278 1341.2 Contig48 P 93650 94402 98%
SEQ ID No. 277 1339.2 Contig48 m 94458 95033 100%
SEQ ID No. 276 1338.3 Contig48 P 95203 97308 98%
SEQ ID No. 3282 757.1 Contig48 P 97275 97922 99%
SEQ ID No. 3283 758.1 Contig48 P 97900 98550 98%
SEQ ID No. 3284 760.2 Contig48 P 98779 100542 99%
SEQ ID No. 1741 3614.3 Contig48 m 100655 101728 99%
SEQ ID No. 2119 4194.3 Contig48 P 101959 103278 99%
SEQ ID No. 3330 825.4 Contig48 m 103305 104558 98%
SEQ ID No. 2885 5438.1 Contig48 171 104563 105147
SEQ ID No. 3329 823.4 Contig48 m 105148 105969 98%
SEQ ID No. 3328 821.2 Contig48 P 106064 106960 98%
SEQ ID No. 3158 6081.1 Contig48 m 107058 107270
SEQ ID No. 1946 396.4 Contig48 m 107186 108082 98%
SEQ ID No. 1934 393.1 Contig48 P 108405 108956 100% +
SEQ ID No. 3379 897.2 Contig48 P 108957 111251 99% +
SEQ ID No. 3380 898.2 Contig48 P 111179 112054 98%
SEQ ID No. 3381 899.3 Contig48 P 112066 112842 98%
SEQ ID No. 1814 3720.2 Contig48 P 112836 113633 97%
SEQ ID No. 2884 5437.2 Contig48 P 113664 113978 94% +
SEQ ID No. 3045 5767.1 Contig48 m 113682 114005 93% +
SEQ ID No. 3046 5769.2 Contig48 m 114096 114503 99%
SEQ ID No. 423 1569.6 Contig48 m 114485 116662 99%
SEQ ID No. 951 2422.3 Contig48 m 116663 117175 98%
SEQ ID No. 952 2423.2 Contig48 m 117160 118278 98%
SEQ ID No. 2922 551.2 Contig48 P 118410 119204 94%
SEQ ID No. 2933 553.1 Contig48 P 119189 119620
SEQ ID No. 2946 555.2 Contig48 m 119679 120698 96%
SEQ ID No. 1360 3058.1 Contig48 P 120708 121190 100%
SEQ ID No. 863 2291.2 Contig48 P 121301 122362 100%
SEQ ID No. 862 2290.1 Contig48 P 122363 123283 98%
SEQ ID No. 861 2289.2 Contig48 P 123222 123761 98%
SEQ ID No. 1359 3057.1 Contig48 P 123677 124783 99%
SEQ ID No. 1358 3055.1 Contig48 P 124770 125705 98%
SEQ ID No. 1357 3054.3 Contig48 P 125624 126583 92%
SEQ ID No. 1356 3053.3 Contig48 m 126062 126499 86%
SEQ ID No. 3157 6079.1 Contig48 m 126066 126479 83%
SEQ ID No. 736 2070.4 Contig48 P 126562 128229 98%
SEQ ID No. 68 1014.2 Contig48 P 128457 129848 98%
SEQ ID No. 67 1013.1 Contig48 m 128699 129295 97% +
SEQ ID No. 734 2068.3 Contig48 P 130078 131586 98% +
SEQ ID No. 2448 4669.2 Contig48 P 131759 132040 97%
SEQ ID No. 2447 4668.1 Contig48 m 132141 133445 98%
SEQ ID No. 2446 4666.2 Contig48 P 133569 134588 98%
SEQ ID No. 365 1482.4 Contig48 P 134694 135689 98%
SEQ ID No. 714 2034.2 Contig48 m 136325 137179 96%
SEQ ID No. 2836 5334.1 Contig48 P 137595 138359 83%
SEQ ID No. 1427 315.3 Contig48 P 137887 138900
SEQ ID No. 1441 317.1 Contig48 m 138934 139956 99% +
SEQ ID No. 1449 318.2 Contig48 m 139970 141388 99% +
SEQ ID No. 589 1812.4 Contig48 m 141531 142739 99%
SEQ ID No. 866 2295.3 Contig48 m 142854 143399 96%
SEQ ID No. 867 2297.3 Contig48 m 143330 143803 99%
SEQ ID No. 2611 4907.2 Contig48 m 143912 144691 98%
SEQ ID No. 2837 5337.1 Contig48 m 144919 145224
SEQ ID No. 2838 5338.2 Contig48 m 145238 146323 100%
SEQ ID No. 2165 4266.3 Contig48 P 146551 147867 99%
SEQ ID No. 3258 723.3 Contig48 P 148217 150046 99%
SEQ ID No. 3257 722.3 Contig48 P 150016 150708 97%
SEQ ID No. 3256 719.4 Contig48 P 150832 153201 99%
SEQ ID No. 3011 5671.2 Contig48 P 153153 153584 100%
SEQ ID No. 3009 5669.2 Contig48 P 153548 155221 99% +
SEQ ID No. 3008 5668.2 Contig48 m 155278 157158 97% +
SEQ ID No. 3005 5656.2 Contig48 P 157395 158135 99%
SEQ ID No. 3092 590.4 Contig48 P 158250 161354 +
SEQ ID No. 2398 4599.2 Contig48 m 161403 162059 +
SEQ ID No. 3156 6074.1 Contig49 P 3 614 +
SEQ ID No. 2278 4421.2 Contig49 P 599 1414 +
SEQ ID No. 2279 4422.1 Contig49 P 1499 1792 + +
SEQ ID No. 2280 4424.1 Contig49 P 1861 2124 + +
SEQ ID No. 2281 4425.1 Contig49 P 2147 2680 94%
SEQ ID No. 2282 4427.1 Contig49 m 3129 3356 + +
SEQ ID No. 2283 4428.1 Contig49 P 3412 3774 + +
SEQ ID No. 2284 4429.2 Contig49 m 3570 3890 91% +
SEQ ID No. 597 1827.3 Contig49 P 4176 5615 +
SEQ ID No. 596 1826.2 Contig49 P 5708 6622 95% +
SEQ ID No. 2621 4920.4 Contig49 m 6970 7851 94%
SEQ ID No. 256 1302.2 Contig49 m 7878 9074 99%
SEQ ID No. 257 1303.3 Contig49 m 9075 9848 97%
SEQ ID No. 1709 3570.2 Contig49 m 9817 10080
SEQ ID No. 931 2392.2 Contig49 m 10206 10697 96% +
SEQ ID No. 930 2391.2 Conιig49 P 10897 1 898 84% +
SEQ ID No. 1710 3572.1 Contig49 m 12027 12509 96%
SEQ ID No. 428 1578.3 Contig49 P 12817 13644 92%
SEQ ID No. 429 1579.4 Contig49 m 13726 14193 92%
SEQ ID No. 2694 ' 5044.2 Contig49 m 14349 15251 98%
SEQ ID No. 2695 5046.2 Contig49 m 15500 15922 98%
SEQ ID No. 2696 5047.3 Contig49 m 16011 16757 98%
SEQ ID No. 1156 2739.3 Contig49 m 16733 17521 99%
SEQ ID No. 1155 2738.2 Contig49 m 17487 18263 99% +
SEQ ID No. 1154 2737.2 Contig49 m 18508 19089 93% +
SEQ ID No. 645 190.3 Contig49 P 19228 19989 93% +
SEQ ID No. 651 191.1 Contig49 m 20119 20982 98%
SEQ ID No. 662 193.3 Contig49 P 21045 22139 97%
SEQ ID No. 675 195.3 Contig49 P 22114 22965 98%
SEQ ID No. 683 197.3 Contig49 P 22941 24404 96%
SEQ ID No. 691 199.1 Contig49 P 24389 25549 96%
SEQ ID No. 700 201.2 Contig49 P 25543 26778 97%
SEQ ID No. 712 2030.2 Contig49 m 26936 28129
SEQ ID No. 718 2041.2 Contig49 P 28480 29592 94%
SEQ ID No. 716 2039.1 Contig49 P 29682 30515
SEQ ID No. 2629 494.2 Contig49 m 30690 31535 95%
SEQ ID No. 2642 496.1 Contig49 P 31594 32652 85% +
SEQ ID No. 2650 497.1 Contig49 m 32699 33202 90% +
SEQ ID No. 2655 498.4 Contig49 P 33556 34560 92%
SEQ ID No. 3042 5761.1 Contig49 P 34545 35045
SEQ ID No. 715 2037.3 Contig49 P 35275 36714 96%
SEQ ID No. 1153 2733.1 Contig49 P 36823 37893 89%
SEQ ID No. 1152 2732.1 Contig49 m 38090 38974 93% +
SEQ ID No. 1151 2730.1 Contig49 P 39326 40288 97% +
SEQ ID No. 182 1188.2 Contig49 m 40523 42844 98%
SEQ ID No. 663 1930.3 Contig49 m 42892 43923 94% +
SEQ ID No. 720 2049.2 Contig49 m 44183 45454 89% +
SEQ ID No. 1149 2728.1 Contig49 P 45792 46145 97%
SEQ ID No. 1148 2727.2 Contig49 P 46358 49045 +
SEQ ID No. 197 1213.2 Contig49 P 49373 50596 +
SEQ ID No. 196 1211.3 Contig49 P 50962 52008 86%
SEQ ID No. 3404 930.3 Contig49 P 52100 53491 99%
SEQ ID No. 3405 932.1 Contig49 P 53437 54480 98%
SEQ ID No. 3406 935.2 Contig49 m 54481 55269 99%
SEQ ID No. 3407 936.3 Contig49 m 55263 55808 97%
SEQ ID No. 1177 2769.2 Contig49 m 55789 57639 96%
SEQ ID No. 1179 2770.1 Contig49 P 57644 58357 96%
SEQ ID No. 1180 2771.2 Contig49 P 58350 59543 97%
SEQ ID No. 3104 5926.1 Contig49 P 59524 59913
SEQ ID No. 1181 2772.1 Contig49 m 59918 60340 98%
SEQ ID No. 3214 657.3 Contig49 P 60518 63580 96%
SEQ ID No. 3213 656.3 Contig49 P 63581 65569 94% 4-
SEQ ID No. 1655 350.3 Contig49 P 65586 66236 98% +
SEQ ID No. 1664 351.1 Contig49 m 66378 66824 91 %
SEQ ID No. 1685 354.1 Contig49 m 66885 68309 97%
SEQ ID No. 1708 357.2 Contig49 m 68395 69129 98%
SEQ ID No. 1719 359.2 Contig49 m 69080 69700 95%
SEQ ID No. 1182 2774.1 Contig49 P 69554 70459 97%
SEQ ID No. 1183 2775.1 Contig49 P 70567 70794
SEQ ID No. 1184 2777.1 Contig49 m 70880 71971 94%
SEQ ID No. 1185 2778.2 Contig49 m 71956 72480 97%
SEQ ID No. 1187 2780.2 Contig49 m 72516 73382 97%
SEQ ID No. 1188 2781.1 Contig49 m 73383 73757 99%
SEQ ID No. 478 1649.3 Contig49 m 73750 76110 94%
SEQ ID No. 479 1650.2 Contig49 m 76053 77585 96%
SEQ ID No. 1063 2587.2 Contig49 P 77654 78526 96%
SEQ ID No. 669 1940.2 Contig49 P 78565 80157 98%
SEQ ID No. 670 1943.3 Contig49 P 80136 81140 + 98%
SEQ ID No. 2375 4561.2 Contig49 P 81131 82534 98%
SEQ ID No. 2374 4560.2 Contig49 P 82681 83823 92%
SEQ ID No. 2373 4559.1 Contig49 m 83994 85562 + 98%
SEQ ID No. 2372 4558.1 Contig49 m 85707 86597 97%
SEQ ID No. 3155 6072.1 Contig49 P 86701 86892 +
SEQ ID No. 3154 6071.1 Contig49 P 86822 87034 +
SEQ ID No. 2371 4554.2 Contig49 m 87282 87542 +
SEQ ID No. 2370 4553.2 Contig49 P 87651 88352 + +
SEQ ID No. 2752 5146.2 Contig49 P 88433 89740 + +
SEQ ID No. 1402 3116.2 Contig49 m 89715 90125 +
SEQ ID No. 1401 3114.2 Contig49 P 90204 91163 +
SEQ ID No. 1400 3113.1 Contig49 P 91157 93160 +
SEQ ID No. 1399 3110.1 Contig49 P 93148 93756 +
SEQ ID No. 1398 3109.2 Contig49 P 93743 94096 +
SEQ ID No. 1397 3107.2 Contig49 m 94191 94412 +
SEQ ID No. 1396 3106.2 Contig49 m 94616 96460 96%
SEQ ID No. 1395 3105.2 Contig49 m 96450 96755
SEQ ID No. 1394 3104.2 Contig49 m 96892 99459 97%
SEQ ID No. 1393 3102.2 Contig49 P 99671 100228 98%
SEQ ID No. 1392 3101.1 Contig49 P 100453 102348
SEQ ID No. 2751 514.5 Contig49 m 102459 107018 95% +
SEQ ID No. 2235 436.4 Contig49 P 104371 104940 96% +
SEQ ID No. 783 2150.1 Contig49 m 107367 107894 97%
SEQ ID No. 201 1222.5 Contig49 m 107944 109923 96%
SEQ ID No. 877 2315.3 Contig49 P 110157 110999 93%
SEQ ID No. 878 2317.2 Contig49 P 111016 112482 97%
SEQ ID No. 1199 2804.1 Contig49 P 112445 113809 98%
SEQ ID No. 858 2279.2 Contig49 P 113883 114656
SEQ ID No. 525 1722.2 Contig49 P 114908 115852 97%
SEQ ID No. 526 1723.5 Contig49 P 115955 116539 98%
SEQ ID No. 3153 6070.1 Contig49 P 116472 116879
SEQ ID No. 1200 2806.1 Contig49 m 117016 117618 99%
SEQ ID No. 3223 670.1 Contig49 m 117945 119357 97%
SEQ ID No. 1385 309.2 Contig49 m 119391 122780 98%
SEQ ID No. 1391 310.1 Contig49 P 122902 123675 96%
SEQ ID No. 1406 312.3 Contig49 P 123904 124869 - 98%
SEQ ID No. 1201 2807.1 Contig49 P 124936 125370 - 95%
SEQ ID No. 962 2434.2 Contig49 m 125457 126668 - 97%
SEQ ID No. 961 2432.2 Contig49 m 126730 127992 - 97%
SEQ ID No. 1202 2808.1 Contig49 P 128188 128580 - 98%
SEQ ID No. 953 2424.3 Contig49 P 128735 129358 - 97%
SEQ ID No. 954 2425.2 Contig49 P 129470 130117 - 97%
SEQ ID No. 955 2426.2 Contig49 P 130209 130778 - 94%
SEQ ID No. 1203 2810.1 Contig49 P 130891 132279 + - 98%
SEQ ID No. 1204 2811.2 Contig49 P 132257 133306 - 98%
SEQ ID No. 1764 365.4 Contig49 P 133354 134355 - 98%
SEQ ID No. 1758 364.2 Contig49 P 134520 136475 - 98%
SEQ ID No. 1730 360.3 Contig49 P 136479 138278 - 97%
SEQ ID No. 713 2031.2 Contig49 P 138614 138955 +
SEQ ID No. 1436 3163.1 Contig49 P 138969 139274 +
SEQ ID No. 1437 3165.1 Contig49 P 139471 139872 - 98%
SEQ ID No. 1438 3166.1 Contig49 m 139958 140611 - 96%
SEQ ID No. 1439 3167.1 Contig49 P 140837 142129 - 100%
SEQ ID No. 1440 3169.1 Contig49 P 142184 142906 - 98%
SEQ ID No. 1000 2488.2 Contig49 m 143043 144032 + +
SEQ ID No. 1001 2489.2 Contig49 m 144070 144399 + +
SEQ ID No. 1003 2490.2 Contig49 P 144461 144931 + +
SEQ ID No. 2483 4719.2 Contig49 m 145036 146628 + - 95%
SEQ ID No. 2482 4718.1 Contig49 P 146787 147245 - 96%
SEQ ID No. 2481 4716.2 Contig49 m 147260 148393 - 96%
SEQ ID No. 1128 2695.2 Contig49 P 148478 149104 - 93%
SEQ ID No. 1129 2696.2 Contig49 P 149173 150738 - 95%
SEQ ID No. 420 1564.3 Contig49 P 150804 152222 - 99%
SEQ ID No. 419 1563.3 Contig49 P 152206 152847 - 95%
SEQ ID No. 1130 2698.1 Contig49 m 152930 153349 - 98%
SEQ ID No. 1133 2700.2 Contig49 m 153313 154554 - 97%
SEQ ID No. 1134 2702.2 Contig49 P 154618 155514 - 97%
SEQ ID No. 1496 326.2 Contig49 P 155911 159276 - 96% +
SEQ ID No. 1490 325.1 Contig49 P 159347 160909 - 94% +
SEQ ID No. 1482 324.3 Contig49 m 160975 161676 +
SEQ ID No. 999 2486.3 Contig49 m 161677 163272 - 97%
SEQ ID No. 1135 2704.4 Contig49 m 163203 164651 - 97%
SEQ ID No. 1136 2705.3 Contig49 m 164641 165852 - 98%
SEQ ID No. 1137 2706.2 Contig49 m 165977 166876 - 94%
SEQ ID No. 179 1183.4 Contig49 P 166928 168976 +
SEQ ID No. 180 1184.2 Contig49 m 168987 170183 + - 95%
SEQ ID No. 181 1186.2 Contig49 m
SEQ ID No. 638 189.3 Contig49 P
SEQ ID No. 630 188.1 Contig49 m 172539 173675 - 96% +
SEQ ID No. 624 187.1 Contig49 P 173879 174583 - 94% +
SEQ ID No. 612 185.1 Contig49 P 174610 175461 + - 98%
SEQ ID No. 603 184.2 Contig49 P 175678 175977 +
SEQ ID No. 682 1968.1 Contig49 P 176650 176925 + - 97%
SEQ ID No. 681 1966.2 Contig49 m 176980 177600 - 98%
SEQ ID No. 1138 2709.1 Contig49 P 177567 178601 - 98%
SEQ ID No. 2166 4267.5 Contig49 m 178670 180760 - 99%
SEQ ID No. 359 1474.4 Contig49 m 180783 181997 - 99%
SEQ ID No. 358 1473.4 Contig49 m 182064 182630 - 98%
SEQ ID No. 2167 4269.3 Contig49 m 182594 183289 - 95%
SEQ ID No. 650 1908.4 Contig49 P 183369 185828 - 99%
SEQ ID No. 2169 4272.3 Contig49 P 185939 186634 - 98%
SEQ ID No. 2204 4323.1 Contig49 m 186671 187360 - 98%
SEQ ID No. 2203 4322.1 Contig49 m 187330 188436 - 96%
SEQ ID No. 2202 4321.2 Contig49 P 188394 191069 - 99%
SEQ ID No. 2201 4320.1 Contig49 P 191146 192534 + - 99%
SEQ ID No. 224 1251.2 Contig49 P 192522 193505 - 97%
SEQ ID No. 223 1250.1 Contig49 P 193501 193998 - 93%
SEQ ID No. 221 1249.2 Contig49 m 194005 194559 +
SEQ ID No. 3184 6134.1 Contigδ m 112 720 +
SEQ ID No. 3183 6133.1 Contigδ m 943 1218 +
SEQ ID No. 3182 6132.1 Contigδ m 1232 1546 +
SEQ ID No. 3152 6066.1 ContigδO P 3 491 +
SEQ ID No. 2960 5576.3 ContigδO P 879 1850 +
SEQ ID No. 2147 4239.4 Contig50 P 1918 2205 + +
SEQ ID No. 2146 4238.1 ContigδO m 2294 2842 + +
SEQ ID No. 2145 4236.3 Contig50 m 2778 4787 +
SEQ ID No. 2144 4234.3 ContigδO m 4788 5267 +
SEQ ID No. 2143 4233.3 ContigδO P 5113 6362 + +
SEQ ID No. 459 1624.4 ContigδO m 53δδ 7022 - + +
SEQ ID No. 458 1623.4 ContigδO m 6937 8328 - +
SEQ ID No. 2561 4822.3 ContigδO m 8329 9436 - +
SEQ ID No. 247 1287.3 Contig50 m 9299 12076 - +
SEQ ID No. 2925 5517.1 ContigδO m 12084 12476 + +
SEQ ID No. 2050 4104.2 ContigδO m 12430 13818 - +
SEQ ID No. 2051 4105.1 ContigδO m 13740 14561 + +
SEQ ID No. 2052 4106.1 ContigδO m 14509 15180 + + +
SEQ ID No. 2053 4107.1 ContigδO m 15176 15646 - + +
SEQ ID No. 914 2367.2 ContigδO m 1δδ3δ 15933 + + +
SEQ ID No. 297 1372.2 ContigδO m 15924 16229 - + +
SEQ ID No. 298 1373.1 ContigδO m 16190 16525 - + +
SEQ ID No. 299 1375.2 ContigδO m 16526 17281 + +
SEQ ID No. 300 1376.2 Contig50 m 17215 17766 - +
SEQ ID No. 2054 4108.1 ContigδO m 17675 18538 + +
SEQ ID No. 2055 4109.2 ContigδO m 18522 19115 + +
SEQ ID No. 3060 5814.1 ContigδO m 19109 195δδ + +
SEQ ID No. 3059 5813.1 ContigδO m 19640 19761 - +
SEQ ID No. 3058 5812.1 ContigδO m 19767 20260 - + +
SEQ ID No. 1086 2625.2 ContigδO m 20086 21020 - + +
SEQ ID No. 1087 2626.1 ContigδO P 21133 21804 - +
SEQ ID No. 583 1803.2 Contig50 P 21826 22089 +
SEQ ID No. 584 1804.2 ContigδO P 222δδ 23602 + 97%
SEQ ID No. 1088 2627.1 ContigδO P 23499 24765 + 98%
SEQ ID No. 1090 2631.4 ContigδO P 24766 27710 - 99%
SEQ ID No. 3112 5953.1 ContigδO P 27632 27916 -
SEQ ID No. 3273 747.3 ContigδO P 27917 30127 - 99%
SEQ ID No. 2761 5164.1 ContigδO P 30361 30561 + +
SEQ ID No. 3272 744.3 ContigδO P 30803 31477 - +
SEQ ID No. 3271 743.3 ContigδO m 31600 31961 - 94%
SEQ ID No. 3111 5952.1 ContigδO P 32298 32483
SEQ ID No. 3425 96.6 ContigδO m 32726 33612 - 99%
SEQ ID No. 3432 97.2 ContigδO m 33710 33985 97% +
SEQ ID No. 3446 99.1 ContigδO m 34081 34362 94% +
SEQ ID No. 2760 5162.1 ContigδO P 34102 34254 - 100% +
SEQ ID No. 69 102.1 Contig50 P 34385 37093 - - 96%
SEQ ID No. 7δ 103.1 ContigδO P 37068 38339 + - 98%
SEQ ID No. 83 104.1 ContigδO P 38294 39469 + - 98%
SEQ ID No. 103 107.1 ContigδO P 39536 42687 - - 98%
SEQ ID No. 117 109.1 ContigδO P 42543 44045 - - 94%
SEQ ID No. 133 111.2 ContigδO P 43985 45656 - - 96%
SEQ ID No. 861 227.2 ContigδO P 45623 46464 - - 93%
SEQ ID No. 859 228.1 ContigδO P 46477 46656 - - 97%
SEQ ID No. 892 234.2 ContigδO P 46705 48930 - - 99%
SEQ ID No. 680 1961.3 ContigδO P 49060 49960 - - 96%
SEQ ID No. 390 152.3 ContigδO P 50084 60461 + - 98%
SEQ ID No. 398 153.2 ContigδO P 60462 51799 + - 99%
SEQ ID No. 405 154.1 ContigδO P 51796 52764 + - 96%
SEQ ID No. 424 157.2 Contig50 P 52765 55980 + - 98%
SEQ ID No. 3107 594.1 ContigδO P 56966 56429 + - 96% +
SEQ ID No. 3105 593.2 ContigδO m 56568 56918 - - 96% +
SEQ ID No. 3102 692.3 ContigδO P 57113 58796 - - 94%
SEQ ID No. 504 169.1 ContigδO m 58808 59020 - +
SEQ ID No. 497 168.1 ContigδO P 59116 59687 - - 97%
SEQ ID No. 486 166.2 ContigδO P 59537 62800 - - 95%
SEQ ID No. 464 163.1 ContigδO P 62978 64276 - +
SEQ ID No. 447 160.1 ContigδO m 64504 64863 +
SEQ ID No. 438 169.1 Contig50 P 65005 66316 - 89%
SEQ ID No. 430 168.2 ContigδO P 65304 66987 - - 98%
SEQ ID No. 679 1960.2 ContigδO P 65972 66484 + - 92%
SEQ ID No. 462 1628.3 ContigδO m 66650 67516 + - 92%
SEQ ID No. 463 1629.1 Contig50 m 67365 68138 + - 95%
SEQ ID No. 46δ 1631.3 ContigδO m 68126 70117 - - 97%
SEQ ID No. 1091 2637.1 Contig50 P 70315 70731 - - 100% +
SEQ ID No. 3326 818.2 ContigδO m 70763 71647 - - 94% +
SEQ ID No. 3326 819.2 ContigδO m 71690 73171 - - 100%
SEQ ID No. 411 1550.2 ContigδO m 73149 73937 - - 97%
SEQ ID No. 410 1549.1 ContigδO m 73898 74191 + - 100%
SEQ ID No. 409 1548.2 ContigδO m 74170 74871 - - 99%
SEQ ID No. 336 1425.2 ContigδO m 74865 75168 - - 97%
SEQ ID No. 335 1424.2 Contig50 m 75133 75661 - - 96%
SEQ ID No. 334 1423.2 ContigδO m 76552 76979 - 97%
SEQ ID No. 1092 2639.1 ContigδO m 77314 77775 + - 95%
SEQ ID No. 1470 3226.2 ContigδO m 78822 79086 - 97%
SEQ ID No. 708 2024.2 ContigδO P 79435 79977 - 98%
SEQ ID No. 363 148.2 ContigδO P 79937 81463 - 96%
SEQ ID No. 376 150.2 ContigδO P 81726 82706 +
SEQ ID No. 2902 5473.1 ContigδO P 82669 83673 +
SEQ ID No. 382 151.2 ContigδO m 83861 84686 - 97%
SEQ ID No. 1471 3228.1 ContigδO m 84690 84905 +
SEQ ID No. 1473 3230.1 ContigδO m 84994 85468 - 90%
SEQ ID No. 1474 3231.1 ContigδO m 85682 85800 +
SEQ ID No. 1475 3232.1 ContigδO m 85971 86342 +
SEQ ID No. 1476 3233.1 ContigδO m 86294 87133 +
SEQ ID No. 1477 3234.3 ContigδO m 87124 87846 +
SEQ ID No. 2903 5474.2 ContigδO P 87816 88361 +
SEQ ID No. 3363 875.6 Contig50 P 88330 89613 + +
SEQ ID No. 3364 876.1 ContigδO P 89606 90100 +
SEQ ID No. 3365 879.2 ContigδO P 90587 91051 +
SEQ ID No. 2982 5615.1 ContigδO P 91012 91698 +
SEQ ID No. 3103 5920.2 ContigδO P 91731 92939 - 97%
SEQ ID No. 3245 704.4 ContigδO m 93538 94440 - 98%
SEQ ID No. 3246 705.1 ContigδO m 94413 95726 - 99%
SEQ ID No. 3247 706.2 ContigδO P 95685 96212 - 97%
SEQ ID No. 3248 707.2 ContigδO m 95702 96757 - 97%
SEQ ID No. 693 1992.2 ContigδO P 96880 97956 - 96%
SEQ ID No. 1637 3472.1 ContigδO m 98100 98402 - 97%
SEQ ID No. 1638 3473.1 ContigδO m 98526 99380 - 92%
SEQ ID No. 1639 3476.1 ContigδO P 99459 100595 - 98%
SEQ ID No. 1640 3477.2 ContigδO P 100596 101333 - 99%
SEQ ID No. 1753 3633.3 ContigδO P 101324 102286 + - 98%
SEQ ID No. 1754 3634.1 ContigδO P 102287 102892 - 100%
SEQ ID No. 1756 3636.1 ContigδO P 102978 103520 + - 100%
SEQ ID No. 1756 3637.1 ContigδO P 103508 104485 + - 98%
SEQ ID No. 1757 3638.2 ContigδO P 104476 105136 - 97%
SEQ ID No. 1759 3640.2 ContigδO P 105120 106100 - 99%
SEQ ID No. 673 1947.3 ContigδO P 106081 106968 - 98%
SEQ ID No. 674 1949.4 ContigδO m 106981 107274 100%
SEQ ID No. 828 2231.3 ContigδO m 107472 108779 99%
SEQ ID No. 827 2230.2 ContigδO m 108772 109404 99%
SEQ ID No. 1760 3641.1 ContigδO m 109367 110134 99%
SEQ ID No. 1761 3642.2 ContigδO m 110241 111392 99%
SEQ ID No. 2724 5087.2 ContigδO P 111723 112805
SEQ ID No. 270 1330.5 ContigδO m 113327 115469 98%
SEQ ID No. 269 1328.2 ContigδO P 115459 116604 100%
SEQ ID No. 268 1325.2 ContigδO P 116708 118477 98%
SEQ ID No. 167 1163.2 ContigδO P 118435 119172 99%
SEQ ID No. 166 1162.1 ContigδO m 119212 119568 98%
SEQ ID No. 166 1161.2 ContigδO P 119787 120848 98%
SEQ ID No. 1897 3867.2 ContigδO m 121146 122486 99%
SEQ ID No. 1896 3863.2 ContigδO P 122748 123362 98%
SEQ ID No. 1976 3999.3 ContigδO P 123480 125441 99%
SEQ ID No. 1975 3998.3 ContigδO m 125026 126495 96%
SEQ ID No. 3357 866.2 ContigδO P 126624 126769 100%
SEQ ID No. 3366 865.2 ContigδO P 126853 127977 98%
SEQ ID No. 460 1625.3 ContigδO P 128135 129334 99%
SEQ ID No. 461 1626.3 ContigδO P 129335 130678 99%
SEQ ID No. 2299 4457.2 ContigδO P 130769 131809 97%
SEQ ID No. 3030 572.2 ContigδO P 131938 133629 98%
SEQ ID No. 3034 573.1 ContigδO m 133783 134067 100%
SEQ ID No. 3038 674.2 ContigδO m 134078 134701 99%
SEQ ID No. 2806 5268.2 ContigδO m 134740 135327 98%
SEQ ID No. 1405 3119.1 ContigδO m 135659 136453 94%
SEQ ID No. 1404 3118.1 ContigδO P 135939 136310 96%
SEQ ID No. 1407 3121.2 ContigδO m 136447 137154 95%
SEQ ID No. 1408 3124.2 ContigδO P 137300 138337 99% +
SEQ ID No. 1409 3127.1 ContigδO m 138666 139430 100% +
SEQ ID No. 2491 473.2 ContigδO P 139666 140541 97%
SEQ ID No. 2484 472.1 ContigδO P 140538 141176 98%
SEQ ID No. 2467 470.3 ContigδO P 141191 141430
SEQ ID No. 2458 469.3 ContigδO P 141568 143721 99%
SEQ ID No. 2454 468.3 ContigδO P 143679 144107 98%
SEQ ID No. 1410 3128.1 Contig50 P 144071 144601 99%
SEQ ID No. 1015 2510.2 ContigδO P 144653 146515 97%
SEQ ID No. 1013 2509.3 ContigδO P 146573 147010 98%
SEQ ID No. 1411 3129.2 ContigδO P 147083 148138 99%
SEQ ID No. 1412 3130.1 ContigδO P 148182 149018 97%
SEQ ID No. 1413 3131.1 ContigδO P 149156 149499 + 100%
SEQ ID No. 3418 949.3 ContigδO P 149500 151377 + 99% +
SEQ ID No. 780 2148.1 ContigδO P 151378 152307 100% +
SEQ ID No. 779 2147.1 ContigδO m 152486 152835 100%
SEQ ID No. 778 2146.2 ContigδO m 152863 154005 98%
SEQ ID No. 1414 3132.1 ContigδO P 154060 164446 95% +
SEQ ID No. 1415 3133.2 ContigδO P 154385 156191 97% +
SEQ ID No. 2807 5269.1 ContigδO P 155067 155423 94% +
SEQ ID No. 2808 5270.2 ContigδO P 155164 155745 95% +
SEQ ID No. 2244 4372.3 ContigδO P 155848 167060 + 98%
SEQ ID No. 2243 4371.1 ContigδO m 157168 168067 99%
SEQ ID No. 2242 4370.1 ContigδO P 168463 159020 98%
SEQ ID No. 2241 4369.1 ContigδO P 159079 159760 88%
SEQ ID No. 2240 4367.1 ContigδO P 159936 160592 82%
SEQ ID No. 2239 4366.1 Contig50 m 160879 161298 91 %
SEQ ID No. 2238 4365.4 ContigδO m 161394 161951 86%
SEQ ID No. 2974 5602.3 ContigδO P 162005 162481 97%
SEQ ID No. 3320 808.4 ContigδO P 162640 165525 78%
SEQ ID No. 3319 807.2 ContigδO P 165692 167020 93%
SEQ ID No. 3318 806.2 ContigδO m 167060 167803 97%
SEQ ID No. 2037 4082.2 ContigδO m 167817 169130 92%
SEQ ID No. 529 1726.4 ContigδO m 169200 170873 93%
SEQ ID No. 2911 5484.4 ContigδO m 170954 172321 94%
SEQ ID No. 2378 4567.2 ContigδO P 172530 174488 91 %
SEQ ID No. 2377 4566.1 ContigδO m 174558 175169 96%
SEQ ID No. 3094 5901.3 ContigδO m 175331 176827 82%
SEQ ID No. 3093 5900.3 ContigδO P 176579 176953 84%
SEQ ID No. 2444 4663.3 ContigδO m 177123 178034 + 97%
SEQ ID No. 2443 4662.1 ContigδO m 178485 178745 + +
SEQ ID No. 2442 4660.2 ContigδO m 179180 179396 + +
SEQ ID No. 2441 4659.2 ContigδO P 179570 179960 93% +
SEQ ID No. 2440 4658.2 ContigδO P 180138 180696 96%
SEQ ID No. 2439 4657.1 ContigδO P 180604 181254 91 %
SEQ ID No. 2438 4656.2 ContigδO P 181681 183482 89% +
SEQ ID No. 3151 6064.1 ContigδO P 183848 184141 92% +
SEQ ID No. 2842 5344.2 ContigδO m 184198 184968 90%
SEQ ID No. 2841 5342.2 ContigδO m 186006 185681 94%
SEQ ID No. 2840 5341.1 ContigδO P 186774 186055 96%
SEQ ID No. 2839 5340.1 ContigδO P 186391 187947 81 %
SEQ ID No. 2657 4984.2 ContigδO m 188128 188745 97%
SEQ ID No. 3150 6062.1 ContigδO P 188912 189430 +
SEQ ID No. 1533 3314.2 Contig51 P 46 753 + +
SEQ ID No. 1534 3316.1 Contig51 m 1154 2896 96% +
SEQ ID No. 726 2057.3 Contig51 P 3624 4694 99%
SEQ ID No. 3437 977.3 Contig51 P 4604 5275 94%
SEQ ID No. 3436 976.3 Contig51 P 6276 5569 94%
SEQ ID No. 3435 974.2 Contigδl P 6479 6735 98%
SEQ ID No. 725 2056.1 Contigδl P 6907 7809 96%
SEQ ID No. 234 1270.2 Contigδl m 7842 8312 99%
SEQ ID No. 235 1271.3 Contigδl P 8263 8610 100%
SEQ ID No. 3119 5983.1 Contigδl P 8510 8743
SEQ ID No. 236 1272.4 Contigδl P 8710 9696 97%
SEQ ID No. 724 2054.2 Contigδl P 9882 10940 97%
SEQ ID No. 2898 6467.1 Contigδl P 11030 11305 95% +
SEQ ID No. 1536 3319.1 Contigδl m 11064 12434 96% +
SEQ ID No. 2645 4962.2 Contigδl m 13063 13476 96% +
SEQ ID No. 2644 4961.1 Contigδl m 13669 14061 96%
SEQ ID No. 2643 4960.2 Contigδl m 14167 15003 96% +
SEQ ID No. 2632 4943.2 Contigδl m 15275 15778 92% +
SEQ ID No. 2631 4942.1 Contig51 m 15834 16097 +
SEQ ID No. 2630 4941.2 Contigδl m 16060 16500 97% +
SEQ ID No. 775 2141.2 Contigδl m 16809 16967 +
SEQ ID No. 3274 748.3 Contigδl P 17140 18522 98%
SEQ ID No. 3275 749.1 Contigδl P 18871 19188 94% +
SEQ ID No. 3276 750.2 Contigδl m 19419 19598 +
SEQ ID No. 3277 751.2 Contigδl P 19507 19821 93% +
SEQ ID No. 3278 752.2 Contig51 m 19664 19837 92% +
SEQ ID No. 3149 6061.1 Contig51 P 19993 20172 94% +
SEQ ID No. 776 2143.5 Contigδl m 20064 21584 96%
SEQ ID No. 871 2302.2 Contigδl m 21648 21992 98%
SEQ ID No. 178 1182.3 Contigδl m 21993 22934 99%
SEQ ID No. 177 1180.2 Contigδl m 22973 23767 98%
SEQ ID No. 175 1179.1 Contigδl m 23986 24621 83%
SEQ ID No. 1140 2712.1 Contigδl m 26013 25828 +
SEQ ID No. 1141 2713.2 Contigδl m 26993 27000 +
SEQ ID No. 1142 2717.1 Contigδl m 27436 29141 97%
SEQ ID No. 1143 2718.1 Contigδl P 29268 30647 98%
SEQ ID No. 992 2477.1 Contigδl P 30907 31392 98%
SEQ ID No. 993 2479.2 Contigδl m 31862 32737 95%
SEQ ID No. 267 1324.4 Contigδl P 33284 34675 91%
SEQ ID No. 737 2073.4 Contigδl m 34700 35431 98%
SEQ ID No. 305 1386.3 Contigδl m 36892 36275 95%
SEQ ID No. 304 1385.2 Contigδl m 36672 37336 90% +
SEQ ID No. 738 2075.1 Contigδl m 37420 37815 100% +
SEQ ID No. 185 1193.2 Contigδl m 37917 38414 98% +
SEQ ID No. 186 1194.3 Contigδl m 38526 39716 100% +
SEQ ID No. 1144 2721.3 Contigδl m 39661 41946 98% +
SEQ ID No. 1145 2724.3 Contigδl P 42127 43302 97%
SEQ ID No. 1146 2725.1 Contigδl P 43206 43565 100%
SEQ ID No. 1147 2726.1 Contigδl P 43667 43867 +
SEQ ID No. 323 141.4 Contigδl P 44028 46769 96% +
SEQ ID No. 331 142.2 Contigδl P 46762 47526 97%
SEQ ID No. 341 144.3 Contigδl P 47728 48822 98%
SEQ ID No. 303 1380.3 Contigδl P 49013 50593 97%
SEQ ID No. 1615 3437.1 Contigδl m 50610 51767 96%
SEQ ID No. 1617 3442.3 Contigδl m 52678 54371 99%
SEQ ID No. 1618 3443.1 Contigδl m 54463 55263 98%
SEQ ID No. 1619 3444.1 Contigδl m 55371 56877 98%
SEQ ID No. 804 2188.2 Contigδl m 56047 66799 100%
SEQ ID No. 3290 767.3 Contigδl m 56793 58106 97%
SEQ ID No. 3291 769.2 Contigδl m 58073 59020 98%
SEQ ID No. 805 2191.4 Contigδl m 59014 69529 100%
SEQ ID No. 1620 3445.3 Contigδl P 59617 60657 99%
SEQ ID No. 1621 3446.1 Contigδl P 60510 61739 98%
SEQ ID No. 1622 3447.1 Contigδl P 61720 62166 97%
SEQ ID No. 2777 5200.2 Contigδl P 62234 64525 99%
SEQ ID No. 1503 3271.3 Contigδl m 64736 66802 99%
SEQ ID No. 1504 3272.1 Contigδl m 66845 67288 100%
SEQ ID No. 1505 3273.1 Contigδl P 67395 67838 100%
SEQ ID No. 1506 3275.1 Contigδl P 67835 69160 97% +
SEQ ID No. 1507 3276.2 Contigδl m 69289 69438 93% +
SEQ ID No. 1508 3277.2 Contigδl P 69443 70258 98%
SEQ ID No. 1509 3278.1 Contigδl m 70313 70750 98%
SEQ ID No. 1511 3280.1 Contigδl m 70844 71740 99%
SEQ ID No. 1512 3281.1 Contigδl P 71741 72040 97%
SEQ ID No. 3201 637.3 Contigδl m 71849 73939 99%
SEQ ID No. 3200 636.2 Contigδl m 74029 74943 100%
SEQ ID No. 3199 635.5 Contigδl m 75191 76447 100%
SEQ ID No. 3198 634.4 Contigδl P 75669 76205 99%
SEQ ID No. 1513 3282.3 Contigδl m 76582 77901 100%
SEQ ID No. 1514 3283.1 Contigδl m 77820 78542 100%
SEQ ID No. 354 1463.2 Contigδl m 78555 79679 99%
SEQ ID No. 353 1461.3 Contigδl m 79633 80634 99%
SEQ ID No. 907 2358.3 Contigδl m 80568 81986 99%
SEQ ID No. 512 1701.5 Contigδl m 81987 83204 99%
SEQ ID No. 2931 5527.2 Contigδl m 83168 83683
SEQ ID No. 511 1699.4 Contigδl m 83619 84545 97%
SEQ ID No. 1798 3694.2 Contigδl m 84535 85659 99%
SEQ ID No. 3231 683.2 Contigδl m 85735 87123 99%
SEQ ID No. 3232 684.5 Contigδl m 87313 88107 99%
SEQ ID No. 1799 3697.4 Contigδl m 88097 91603 98%
SEQ ID No. 1428 3150.1 Contigδl P 91763 92446 99%
SEQ ID No. 1426 3149.1 Contigδl m 92548 93645 99%
SEQ ID No. 1425 3148.1 Contigδl P 93871 94731 98%
SEQ ID No. 1424 3147.1 Contigδl m 94805 95329 100%
SEQ ID No. 3259 727.3 Contigδl m 95296 98532 99%
SEQ ID No. 750 2098.2 Contigδl m 98628 98909 95%
SEQ ID No. 749 2095.2 Contigδl P 99001 100194 95%
SEQ ID No. 1423 3146.2 Contigδl m 100234 101040 98%
SEQ ID No. 1422 3145.1 Contigδl P 100490 100798 98%
SEQ ID No. 3366 880.3 Contigδl m 100986 102164 99%
SEQ ID No. 3367 881.2 Contigδl m 102053 103141 + 98%
SEQ ID No. 3368 882.3 Contigδl P 103303 104787 98%
SEQ ID No. 824 2221.3 Contigδl P 104753 105202 97%
SEQ ID No. 2245 4373.3 Contigδl m 106349 105768 97%
SEQ ID No. 1033 2542.2 Contigδl m 105689 107066 98%
SEQ ID No. 2246 4374.4 Contigδl m 107058 108635 99%
SEQ ID No. 3054 5806.2 Contigδl P 108987 109253 100%
SEQ ID No. 2557 4818.3 Contigδl m 109314 110570 98% +
SEQ ID No. 2556 4817.4 Contigδl m 110877 112283 98% +
SEQ ID No. 2100 4167.4 Contigδl m 112666 116331 + 98%
SEQ ID No. 2099 4166.1 Contigδl m 116296 116864 + 97%
SEQ ID No. 2098 4164.1 Contigδl m 116858 117592 + 99%
SEQ ID No. 2097 4162.2 Contigδl P 117747 120260 97%
SEQ ID No. 3193 628.3 Contigδl P 120526 121188 98%
SEQ ID No. 3192 627.1 Contigδl P 121353 122504 93%
SEQ ID No. 820 2217.3 Contigδl m 122528 124399 99%
SEQ ID No. 384 1511.3 Contigδl m 124396 125073 99%
SEQ ID No. 383 1510.3 Contigδl m 125180 126028 99%
SEQ ID No. 381 1608.4 Contigδl P 126402 127535 98%
SEQ ID No. 1579 3386.1 Contigδl m 127494 128615 99%
SEQ ID No. 1580 3386.1 Contigδl m 128733 129023 100%
SEQ ID No. 641 1894.2 Contig51 m 129024 129398 100%
SEQ ID No. 640 1893.2 Contigδl P 129560 130237 100%
SEQ ID No. 842 2253.2 Contigδl P 130333 130923 98%
SEQ ID No. 841 2252.1 Contig51 P 130939 132033 99%
SEQ ID No. 1581 3388.1 Contig51 P 132396 133510 98%
SEQ ID No. 1582 3389.1 Contigδl P 133644 134612 99%
SEQ ID No. 1584 3390.1 Contigδl m 134639 134893 100%
SEQ ID No. 1586 3391.2 Contigδl m 134835 137117 99%
SEQ ID No. 2768 5180.1 Contig51 m 137087 137341
SEQ ID No. 1690 3544.2 Contigδl m 137403 138500 98%
SEQ ID No. 1689 3643.1 Contigδl m 138482 139159 96%
SEQ ID No. 1688 3542.1 Contig51 P 139288 140169 99%
SEQ ID No. 1687 3541.1 Contig51 P 140157 140939 98%
SEQ ID No. 1370 307.2 Contigδl P 140927 141469 97%
SEQ ID No. 1362 306.1 Contig51 m 141483 142922 - 99%
SEQ ID No. 1355 305.2 Contigδl m 142891 143622 - 99%
SEQ ID No. 527 1724.3 Contigδl P 143835 144766 - 98%
SEQ ID No. 528 1725.3 Contigδl m 144891 145793 - 100%
SEQ ID No. 2183 4294.3 Contigδl m 146041 147027 - 99%
SEQ ID No. 890 2337.3 Contigδl m 146964 147656 - 99%
SEQ ID No. 891 2339.4 Contigδl m 147601 148743 - 98%
SEQ ID No. 2184 4296.3 Contigδl P 148722 150056 - 99%
SEQ ID No. 2185 4296.1 Contigδl P 150005 151357 - 98%
SEQ ID No. 2186 4297.2 Contigδl P 151324 151899 - 97%
SEQ ID No. 2767 5178.1 Contigδl P 151974 152465 + 99%
SEQ ID No. 911 2362.3 Contigδl P 152437 153357 + 99%
SEQ ID No. 912 2364.1 Contigδl P 153306 154490 - 99%
SEQ ID No. 913 2365.2 Contigδl P 154521 156299 + 95%
SEQ ID No. 524 1720.3 Contigδl m 166341 157147 - 99%
SEQ ID No. 1684 3538.1 Contig51 m 157168 158121 - 98%
SEQ ID No. 239 1277.2 Contigδl P 158205 159371 - 99%
SEQ ID No. 238 1275.2 Contigδl P 159645 160532 - 97%
SEQ ID No. 237 1273.4 Contigδl P 160659 161378 - 90%
SEQ ID No. 1686 3540.2 Contigδl P 161362 163212 - 93%
SEQ ID No. 1715 3582.2 Contigδl P 163197 164072 - 89%
SEQ ID No. 1714 3581.1 Contigδl m 164333 165085 - 93%
SEQ ID No. 559 1770.2 Contigδl m 166180 168293 + 85%
SEQ ID No. 560 1772.1 Contigδl m 168284 168760 - 89%
SEQ ID No. 561 1773.1 Contigδl P 168849 169307 - 99%
SEQ ID No. 562 1774.2 Contigδl P 169304 170725 + 87%
SEQ ID No. 1713 3578.2 Contigδl m 170732 172456 - 88%
SEQ ID No. 1887 3846.2 Contigδl P 172718 175288 - 95%
SEQ ID No. 1150 273.1 Contigδl P 175615 176069 - 98%
SEQ ID No. 1139 271.1 Contigδl m 176141 177055 - 98%
SEQ ID No. 1132 270.1 Contigδl m 177065 177934 - 99%
SEQ ID No. 1126 269.1 Contigδl m 178010 179101 - 98%
SEQ ID No. 1120 268.1 Contig51 m 179040 179276 -
SEQ ID No. 1115 267.3 Contigδl m 179257 180225 + 98%
SEQ ID No. 2793 5232.1 Contigδl m 180191 181312 - 93%
SEQ ID No. 1403 3117.2 Contig51 m 181268 181983 -
SEQ ID No. 208 123.2 Contigδl m 181917 183347 + 99%
SEQ ID No. 216 124.1 Contigδl m 183344 183853 100%
SEQ ID No. 222 125.2 Contigδl m 183952 185502 98%
SEQ ID No. 707 2022.1 Contigδl P 185614 185687 98%
SEQ ID No. 706 2021.1 Contigδl m 185661 185980 99%
SEQ ID No. 705 2020.1 Contigδl m 185969 186597 99%
SEQ ID No. 703 2019.2 Contig51 m 186827 187576 + 99%
SEQ ID No. 3240 698.2 Contig51 m 187570 188787 99%
SEQ ID No. 3239 697.1 Contig51 m 188796 189424 + 98%
SEQ ID No. 3238 696.2 Contigδl m 189586 190029 100%
SEQ ID No. 2791 5229.1 Contig51 m 190023 190493 99%
SEQ ID No. 2578 4856.2 Contig51 m 190797 191123 96%
SEQ ID No. 2577 4855.2 Contigδl m 191116 191436 100%
SEQ ID No. 940 2407.3 Contigδl m 191412 193793 98%
SEQ ID No. 2336 4503.1 Contigδl m 194051 195133 99% +
SEQ ID No. 2337 4505.3 Contigδl m 195204 195566 100% +
SEQ ID No. 2790 5227.2 Contigδl m 195579 195845
SEQ ID No. 2789 5226.2 Contig51 m 195800 196339 99%
SEQ ID No. 2622 4923.2 Contig51 m 196361 198343 99%
SEQ ID No. 1669 3516.2 Contig51 P 198784 199077 100%
SEQ ID No. 1668 3515.2 Contigδl P 199397 199897 98%
SEQ ID No. 1667 3514.1 Contigδl P 199903 200154
SEQ ID No. 1666 3612.1 Contigδl m 200543 202108 97%
SEQ ID No. 1665 3611.2 Contig51 P 202374 203537 98%
SEQ ID No. 1663 3609.2 Contigδl P 203513 204547 98%
SEQ ID No. 1064 2590.3 Contigδl P 204548 205471 99%
SEQ ID No. 1065 2591.3 Contig51 m 205599 206756 98%
SEQ ID No. 1662 3508.2 Contigδl m 206726 207028 97%
SEQ ID No. 1661 3507.1 Contigδl m 207278 207652 98%
SEQ ID No. 1660 3506.1 Contigδl P 207724 208368 100% +
SEQ ID No. 1659 3504.1 Contigδl m 208578 209129 99% +
SEQ ID No. 1658 3503.2 Contigδl m 209099 210298 99% +
SEQ ID No. 1657 3502.3 Contigδl m 210312 210839 + 97%
SEQ ID No. 3053 5804.1 Contigδl m 210800 211321 + 100%
SEQ ID No. 388 1517.3 Contig51 P 211384 212667 98%
SEQ ID No. 389 1519.2 Contig51 m 212748 213812 100%
SEQ ID No. 391 1521.2 Contigδl P 213997 214617 97%
SEQ ID No. 2276 4415.3 Contigδl m 214705 215997 100%
SEQ ID No. 2205 4325.3 Contigδl P 216364 217314 97%
SEQ ID No. 2206 4326.1 Contigδl P 217278 218063 96%
SEQ ID No. 2207 4327.1 Contigδl P 218064 219191 97%
SEQ ID No. 2208 4328.1 Contigδl P 219146 220381 + 98%
SEQ ID No. 2499 4740.2 Contigδl m 220517 221473 99%
SEQ ID No. 2497 4739.1 Contigδl m 221437 222102 99%
SEQ ID No. 2496 4737.1 Contigδl m 222089 222667 100%
SEQ ID No. 2495 4736.2 Contigδl m 222642 223916 98%
SEQ ID No. 2494 4734.2 Contigδl P 223928 224914 97%
SEQ ID No. 2887 5440.1 Contigδl m 224949 226127 97% +
SEQ ID No. 3202 64.3 Contigδl m 226279 228249 97% +
SEQ ID No. 3209 65.1 Contigδl m 228321 228539 + +
SEQ ID No. 2951 5556.1 Contigδ2 P 224 484 + +
SEQ ID No. 2950 5556.2 Contig52 P 358 1134 + +
SEQ ID No. 3147 6052.1 Contigδ2 m 1146 1837 +
SEQ ID No. 2778 5201.2 Contig52 m 1785 2489 +
SEQ ID No. 2779 5202.2 Contigδ2 m 2453 2761 +
SEQ ID No. 2780 5204.2 Contig52 m 2762 3238 +
SEQ ID No. 2295 4444.2 Contigδ2 P 3519 4844 98%
SEQ ID No. 2294 4442.1 Contigδ2 m 4989 5946 98%
SEQ ID No. 2293 4441.1 Contig52 P 6110 6532 99%
SEQ ID No. 2292 4440.1 Contigδ2 P 6424 6957 98%
SEQ ID No. 621 1863.2 Contigδ2 P 6950 7645 99%
SEQ ID No. 620 1862.2 Contig52 P 7735 9072 96%
SEQ ID No. 3315 801.3 Contigδ2 P 9080 9829 94%
SEQ ID No. 3316 802.3 Contig52 P 9968 10765 99%
SEQ ID No. 3317 803.3 Contigδ2 P 10775 11470 + 99%
SEQ ID No. 2781 5206.1 Contig52 P 11494 12612 + 98%
SEQ ID No. 1421 3142.2 Contigδ2 P 12546 13499 94%
SEQ ID No. 401 1532.3 Contig52 P 13507 15498 97%
SEQ ID No. 402 1533.1 Contig52 P 15602 16737 97%
SEQ ID No. 220 1246.2 Contigδ2 P 16920 18671 80%
SEQ ID No. 219 1245.1 Contig52 P 18783 19034 87% +
SEQ ID No. 218 1244.2 Contigδ2 P 19306 20388 89% +
SEQ ID No. 329 1417.1 Contig52 P 20421 20891 - 100%
SEQ ID No. 330 1418.3 Contigδ2 m 20994 22823 + - 90%
SEQ ID No. 3146 6049.1 Contig52 m 22855 23631 +
SEQ ID No. 1285 2954.3 Contigδ2 m 23498 24508 - 99%
SEQ ID No. 3131 601.4 Contig52 P 24684 25916 +
SEQ ID No. 3127 600.3 Contigδ2 P 25859 26539 +
SEQ ID No. 3118 598.2 Contig52 P 26619 26939 + - 100%
SEQ ID No. 3115 597.2 Contigδ2 P 26966 27346 + - 98%
SEQ ID No. 3114 596.3 Contig52 m 27403 29127 - 99%
SEQ ID No. 1286 2956.2 Contig52 P 29162 29581 - 99%
SEQ ID No. 1287 2958.2 Contig52 P 29575 30342 - 98%
SEQ ID No. 1288 2969.1 Contigδ2 P 30346 31104 + - 97%
SEQ ID No. 1290 2960.1 Contigδ2 P 31112 32216 - 98%
SEQ ID No. 1291 2961.1 Contigδ2 m 32318 33571 - 98%
SEQ ID No. 1292 2962.1 Contigδ2 P 33682 34458 - 98%
SEQ ID No. 1006 2495.2 Contigδ2 P 34434 34982 - 98%
SEQ ID No. 1005 2493.2 Contig52 P 34913 36578 - 98%
SEQ ID No. 1004 2492.2 Contigδ2 P 35616 35948 + - 95%
SEQ ID No. 3409 938.3 Contig52 m 35989 37614 - 90%
SEQ ID No. 3408 937.3 Contigδ2 m 37875 39305 + - 99%
SEQ ID No. 437 1589.3 Contig52 m 39287 39994 - 99%
SEQ ID No. 439 1590.1 Contigδ2 m 40025 40582 + - 98%
SEQ ID No. 440 1691.4 Contig52 m 40813 42264 - 98%
SEQ ID No. 1293 2963.2 Contigδ2 m 42366 43133 - 98%
SEQ ID No. 1294 2964.1 Contig52 m 43134 43991 - 98%
SEQ ID No. 1295 2965.1 Contigδ2 P 44163 44477 - 100% +
SEQ ID No. 1296 2966.1 Contig52 m 44480 45376 - 97% +
SEQ ID No. 1297 2967.1 Contigδ2 m 45421 45705 - 100% +
SEQ ID No. 1298 2968.3 Contig52 P 45890 48583 - 76%
SEQ ID No. 2892 5455.1 Contigδ2 m 48635 49105 - 98%
SEQ ID No. 787 2156.3 Contig52 m 49102 50409 - 98%
SEQ ID No. 538 1740.3 Contigδ2 P 50644 51432 - 100%
SEQ ID No. 536 1739.2 Contig52 P 51433 62056 - 99%
SEQ ID No. 535 1738.3 Contigδ2 P 52040 53257 - 99%
SEQ ID No. 408 1547.2 Contigδ2 P 63250 54101 - 99%
SEQ ID No. 407 1545.3 Contig52 P 54083 56763 - 99%
SEQ ID No. 1531 3311.1 Contig52 P 56764 57133 98% SEQ ID No. 831 2235.2 Contig52 P 57282 59225 97% SEQ ID No. 1632 3312.1 Contigδ2 P 59352 59669 98% SEQ ID No. 2937 6634.2 Contigδ2 m 59770 61257 99% SEQ ID No. 2121 4197.2 Contigδ2 m 61258 63633 99% SEQ ID No. 2938 5635.1 Contigδ2 P 62267 62506 97% SEQ ID No. 346 1448.2 Contigδ2 P 63680 64834 96% SEQ ID No. 346 1449.3 Contig52 m 64855 67023 + 98% SEQ ID No. 2120 4195.3 Contigδ2 m 67104 68273 + 99% SEQ ID No. 2734 5100.2 Contig52 m 68261 69484 96% SEQ ID No. 1678 3378.2 Contigδ2 m 69468 71318 98% SEQ ID No. 1677 3376.1 Contigδ2 m 71343 72434 96% SEQ ID No. 1773 366.2 Contig52 P 72814 73131 91 % SEQ ID No. 1788 368.2 Contigδ2 m 73211 75703 93% SEQ ID No. 1576 3374.1 Contig52 m 75741 76907 94% SEQ ID No. 3441 982.3 Contig52 m 77475 78893 + 98% SEQ ID No. 3442 983.1 Contig52 m 78868 79611 98% SEQ ID No. 3443 985.2 Contig52 m 79583 80569 99% SEQ ID No. 1576 3371.2 Contig52 m 80548 81924 100% SEQ ID No. 2553 4813.2 Contig52 m 81928 83229 99% SEQ ID No. 1573 3368.2 Contig52 P 82091 82516 97% SEQ ID No. 2564 4814.2 Contigδ2 m 83467 84225 99% SEQ ID No. 2565 4815.2 Contigδ2 m 84536 84952 99% SEQ ID No. 255 130.4 Contig52 m 84956 86611 98% SEQ ID No. 249 129.3 Contigδ2 m 86644 86940 96% SEQ ID No. 227 126.2 Contig52 m 87008 88483 98% SEQ ID No. 2433 465.2 Contig52 P 88842 89729 99% SEQ ID No. 2425 464.2 Contig52 P 89686 90996 95% SEQ ID No. 2420 463.2 Contιgδ2 m 90941 91528 95% SEQ ID No. 2412 462.2 Contig52 P 90969 91781 + 94% SEQ ID No. 2400 460.2 Contigδ2 P 91747 92295 98% SEQ ID No. 2393 459.3 Contig52 m 92270 92908 99% SEQ ID No. 2386 458.3 Contigδ2 m 92893 93930 98% SEQ ID No. 1279 2943.1 Contigδ2 m 93920 94501 + 100% SEQ ID No. 1278 2942.1 Contigδ2 m 94512 96995 99% SEQ ID No. 1277 2941.1 Contig52 m 97049 98665 97%
SEQ ID No. 3108 5944.1 Contig52 P 98412 98600 96%
SEQ ID No. 198 1216.3 Contigδ2 P 98687 99826 - 100%
SEQ ID No. 200 1220.3 Contigδ2 P 99827 101356 - 98%
SEQ ID No. 761 2103.2 Contig52 P 101440 103845 + 99%
SEQ ID No. 470 1638.2 Contigδ2 P 103846 105045 - 99%
SEQ ID No. 471 1639.4 Contig52 m 105168 106336 - 91 % +
SEQ ID No. 468 1635.4 Contig52 m 106495 109404 - 92% +
SEQ ID No. 2909 5481.1 Contigδ2 P 109609 110751 + 97%
SEQ ID No. 2445 4665.2 Contig52 m 110890 113277 - 98%
SEQ ID No. 2178 4288.2 Contigδ2 m 113583 114581 + 97%
SEQ ID No. 2179 4289.2 Contig52 m 114538 115264 + 99%
SEQ ID No. 2181 4291.1 Contigδ2 m 115152 116566 + 98%
SEQ ID No. 3293 770.3 Contig52 m 115526 116104 - 97%
SEQ ID No. 3294 771.4 Contig52 m 115998 116444 - 100%
SEQ ID No. 3295 772.4 Contigδ2 m 116524 117759 - 99% +
SEQ ID No. 3296 774.3 Contig52 P 117790 119229 - 100% +
SEQ ID No. 3297 775.2 Contigδ2 P 119230 119802 + 99%
SEQ ID No. 2182 4293.2 Contig52 P 119803 120579 - 99%
SEQ ID No. 2910 5482.2 Contigδ2 P 120600 121508 - 98%
SEQ ID No. 1920 3909.3 Contig52 P 121603 123297 - 97%
SEQ ID No. 1921 3911.2 Contigδ2 P 123484 125370 - 99%
SEQ ID No. 925 2382.2 Contig52 m 125482 125799 - 100%
SEQ ID No. 924 2381.2 Contigδ2 m 125894 127072 - 96%
SEQ ID No. 922 2377.2 Contig52 m 127035 128015 - 99%
SEQ ID No. 921 2375.3 Contigδ2 P 128218 128793 - 98%
SEQ ID No. 1922 3913.2 Contig52 m 128906 130032 + 100%
SEQ ID No. 1923 3914.3 Contigδ2 m 130020 131921 + 99%
SEQ ID No. 832 2239.3 Contig52 P 132081 133076 -
SEQ ID No. 833 2240.2 Contigδ2 m 133339 133824 - 98%
SEQ ID No. 834 2241.2 Contig52 m 133818 134192 - 99%
SEQ ID No. 2253 4384.2 Contigδ2 m 134261 135727 - 99%
SEQ ID No. 3145 6044.1 Contig52 P 135288 135446 96%
SEQ ID No. 2254 4385.1 Contigδ2 P 135914 137590 - 99%
SEQ ID No. 2255 4386.2 Contig52 P 137614 139124 - 98%
SEQ ID No. 2585 4867.2 Contigδ2 P 139087 140626 - 92%
SEQ ID No. 2586 4868.2 Contig52 P 140406 141347 - 98%
SEQ ID No. 2587 4869.3 Contig52 P 141336 141775 100%
SEQ ID No. 2945 5549.2 Contigδ2 P 141660 142481 99%
SEQ ID No. 2944 5548.1 Contig52 m 142688 143068 100%
SEQ ID No. 2943 5546.1 Contigδ2 P 143354 143830 100%
SEQ ID No. 2590 4874.2 Contig52 P 143907 145362 77%
SEQ ID No. 2591 4875.1 Contig52 m 145418 145843 96%
SEQ ID No. 2592 4877.2 Contig52 m 145946 147057 98%
SEQ ID No. 2419 4629.2 Contig52 P 147294 147728 98%
SEQ ID No. 2418 4628.2 Contig52 P 147706 148005 100%
SEQ ID No. 2417 4627.2 Contig52 P 147984 149039 100%
SEQ ID No. 2416 4626.1 Contig52 P 149012 149989 99%
SEQ ID No. 2415 4625.2 Contig52 P 149995 150963 98%
SEQ ID No. 2645 4803.2 Contig52 P 150976 151721 99%
SEQ ID No. 2546 4804.1 Contig52 P 151758 152064 100%
SEQ ID No. 2547 4805.1 Contig52 P 152038 153312 + 99%
SEQ ID No. 2548 4806.4 Contig52 P 153313 154311 + 97%
SEQ ID No. 3144 6042.1 Contig52 P 154287 164949 97%
SEQ ID No. 2641 4958.3 Contigδ2 P 164943 165861 96%
SEQ ID No. 2640 4957.2 Contig52 P 156962 156318 98%
SEQ ID No. 282 1346.3 Contigδ2 P 156342 157166 98%
SEQ ID No. 281 1345.3 Contig52 P 157275 158378 97%
SEQ ID No. 1670 3517.3 Contig52 P 158419 159726 99%
SEQ ID No. 347 1450.2 Contig52 m 160244 161539 98%
SEQ ID No. 1671 3519.1 Contig52 P 161680 162921 98%
SEQ ID No. 1672 3520.1 Contig52 m 163009 164394 + 97%
SEQ ID No. 1673 3521.2 Contig52 P 164484 165635 97%
SEQ ID No. 1674 3522.1 Contig52 m 165688 166524 99%
SEQ ID No. 1675 3524.1 Contig52 P 166701 167300 98%
SEQ ID No. 1676 3525.1 Contigδ2 P 167624 168361 99%
SEQ ID No. 1677 3527.1 Contig52 P 168334 168732 100%
SEQ ID No. 2139 4228.2 Contigδ2 m 168692 170233 99%
SEQ ID No. 2138 4227.1 Contig52 P 170259 171749 98%
SEQ ID No. 96 1059.2 Contigδ2 m 171843 173126 99%
SEQ ID No. 97 1060.2 Contig52 P 173322 174221 94%
SEQ ID No. 2137 4226.2 Contigδ2 P 174266 176896 98%
SEQ ID No. 932 2395.3 Contig52 P 176847 177977 94%
SEQ ID No. 464 1611.5 Contig52 P 177956 179017 98%
SEQ ID No. 3143 6040.1 Contigδ2 P 178884 179286
SEQ ID No. 3081 5876.2 Contigδ2 P 179252 179977 99%
SEQ ID No. 3141 6039.1 Contigδ2 P 179971 181719 100%
SEQ ID No. 2966 5588.2 Contigδ2 P 181821 182123 100%
SEQ ID No. 2325 4489.3 Contigδ2 P 182170 183339 100%
SEQ ID No. 2324 4488.1 Contigδ2 P 183570 184718 100%
SEQ ID No. 2323 4487.1 Contigδ2 P 184719 185408 97%
SEQ ID No. 578 1795.3 Contig52 P 185529 188249 98%
SEQ ID No. 2542 4800.2 Contigδ2 P 188179 188973 98%
SEQ ID No. 2643 4801.1 Contigδ2 P 188964 189461 93%
SEQ ID No. 2544 4802.2 Contig52 P 189554 190114 94%
SEQ ID No. 2965 5587.3 Contig52 P 190081 190959 98%
SEQ ID No. 2735 5103.4 Contig52 m 191093 191497 100%
SEQ ID No. 2736 5104.3 Contigδ2 m 191571 193376 97%
SEQ ID No. 577 1794.3 Contig52 m 193397 194182 97%
SEQ ID No. 576 1793.3 Contigδ2 m 194158 195009 99%
SEQ ID No. 2894 6457.1 Contigδ2 m 195010 195438 99%
SEQ ID No. 2670 5003.2 Contigδ2 m 195444 196262 97%
SEQ ID No. 2671 5005.3 Contig52 m 196424 197824 99%
SEQ ID No. 2893 6466.1 Contig52 m 197784 198467 99%
SEQ ID No. 1812 3712.2 Contig52 m 198472 199365 98%
SEQ ID No. 1811 3711.1 Contigδ2 m 199322 199801 100%
SEQ ID No. 1810 3710.1 Contigδ2 m 199795 200760 98%
SEQ ID No. 1808 3709.1 Contig52 P 201015 201863 96%
SEQ ID No. 1807 3708.1 Contigδ2 P 201767 202480 99%
SEQ ID No. 1806 3706.2 Contig52 P 202699 203940 99%
SEQ ID No. 1805 3705.1 Contigδ2 P 203934 204731 98%
SEQ ID No. 1804 3703.1 Contig52 P 204682 205314 98%
SEQ ID No. 1803 3702.1 Contig52 P 205289 206050 99%
SEQ ID No. 1802 3701.1 Contig52 P 206268 207071 97%
SEQ ID No. 1800 3699.3 Contigδ2 P 207467 210069 96%
SEQ ID No. 2149 4242.2 Contig52 P 210147 211148 95%
SEQ ID No. 2150 4243.2 Contig52 m 211272 211772 95%
SEQ ID No. 2151 4244.1 Contig52 P 211897 212304
SEQ ID No. 2152 4246.2 Contig52 P 212331 213194 96%
SEQ ID No. 266 1323.2 Contig52 P 213309 213839 98%
SEQ ID No. 265 1322.2 Contig52 m 213961 216397 97%
SEQ ID No. 395 1525.3 Contig52 m 215376 216646 98%
SEQ ID No. 396 1528.4 Contig52 P 216734 218086 98%
SEQ ID No. 400 1531.3 Contig52 P 217993 220296 99%
SEQ ID No. 399 1530.2 Contig52 m 220337 220723 98%
SEQ ID No. 397 1529.2 Contigδ2 P 220737 221964 99%
SEQ ID No. 1540 3327.1 Contig52 P 221944 222768 96%
SEQ ID No. 1541 3328.1 Contigδ2 P 222755 224497 98%
SEQ ID No. 1543 3331.1 Contigδ2 P 224570 226568 96%
SEQ ID No. 433 1584.3 Contigδ2 P 225621 228359 99%
SEQ ID No. 1544 3333.1 Contigδ2 P 228335 228760 99%
SEQ ID No. 1545 3334.2 Contigδ2 P 228944 229807 97%
SEQ ID No. 2678 5021.3 Contig52 m 229852 230601 100%
SEQ ID No. 2676 6019.1 Contigδ2 m 230836 231474 99%
SEQ ID No. 2675 5018.2 Contig52 m 231413 231898 98%
SEQ ID No. 2464 4696.3 Contigδ2 m 231883 233508 99%
SEQ ID No. 2463 4695.1 Contig52 P 233502 233720
SEQ ID No. 2462 4694.1 Contigδ2 P 233593 235266 98%
SEQ ID No. 2461 4692.5 Contigδ2 m 236687 236966 98%
SEQ ID No. 444 1598.6 Contigδ2 P 236937 237929 100%
SEQ ID No. 443 1595.3 Contig52 P 237930 239072 98%
SEQ ID No. 442 1594.2 Contig52 P 239053 239775 97%
SEQ ID No. 989 2472.2 Contigδ2 P 239763 240410 96%
SEQ ID No. 990 2473.1 Contig52 P 240545 240859 100%
SEQ ID No. 991 2474.1 Contigδ2 P 240853 241485 99%
SEQ ID No. 2609 4901.3 Contigδ2 P 241486 243285 99%
SEQ ID No. 2608 4900.3 Contigδ2 P 243257 246211 98%
SEQ ID No. 2247 4376.3 Contig52 P 246162 246848 97%
SEQ ID No. 2248 4377.4 Contigδ2 m 246921 247985 96% +
SEQ ID No. 3140 6038.1 Contig52 P 247180 247419 96% +
SEQ ID No. 3433 971.2 Contigδ2 m 248160 249020 98% +
SEQ ID No. 3434 972.3 Contigδ2 P 249094 250731 96%
SEQ ID No. 2604 4891.2 Contigδ2 m 250785 251615 99%
SEQ ID No. 2602 4889.1 Contig52 m 251793 252218 100%
SEQ ID No. 2601 4888.1 Contigδ2 m 252280 252804 96%
SEQ ID No. 2600 4887.3 Contig52 P 262888 SEQ ID No. 2716 253461 5075.4 Contig52 97% P 253679 256276 SEQ ID No. 3139 6037.1 Contig52 94% P 255546 255887 SEQ ID No. 2715 5072.2 Contigδ2 81% m 255557 SEQ ID No. 2714 256015 5071.2 Contigδ2 82% P 256486 SEQ ID No. 2296 257611 4452.2 Contig52 88% P 257614 SEQ ID No. 2297 259002 4453.2 Contigδ2 94% P 259133 SEQ ID No. 2298 260563 4454.4 Contigδ2 78% m 260752 261759 SEQ ID No. 3138 6036.1 Contigδ2 95% P 262003 SEQ ID No. 2963 262335 + 5584.2 Contigδ2 m + 262416 SEQ ID No. 2662 263513 + 4993.2 Contigδ2 m + 263713 SEQ ID No. 2661 264231 '4992.2 Contigδ2 99% P 264173 SEQ ID No. 2660 264472 4990.3 Contigδ2 98% + P 264835 SEQ ID No. 984 266655 2466.3 Contigδ2 94% + P 266848 SEQ ID No. 2389 269556 4584.3 Contig52 96% m 269691 SEQ ID No. 2953 270662 5559.3 Contigδ2 98% m 270701 + SEQ ID No. 2954 271255 5660.3 Contig52 97% m + 271292 SEQ ID No. 2965 271954 5563.1 Contig52 98% m 272495 SEQ ID No. 2956 272969 5664.1 Contig52 100% m 273293 SEQ ID No. 3387 274750 908.4 Contigδ2 99% m 274806 SEQ ID No. 2237 276458 4364.2 Contigδ2 97% P 274900 SEQ ID No. 3386 275220 907.4 Contigδ2 97% m 276498 SEQ ID No. 2388 279179 4581.2 Contigδ2 99% m 279189 SEQ ID No. 2387 279581 4580.1 Contig52 99% m 279617 SEQ ID No. 2385 280120 4578.1 Contigδ2 90% m 280373 SEQ ID No. 2384 281584 4577.2 Contig52 99% m 281864 SEQ ID No. 2258 282706 4389.2 Contigδ2 98% P 282782 SEQ ID No. 2259 284104 4391.1 Contig52 99% P 284105 SEQ ID No. 2260 284953 4392.1 Contigδ2 97% P 284954 SEQ ID No. 2261 285220 4393.1 Contig52 P 285109 SEQ ID No. 2262 285819 4394.4 Contigδ2 96% P 286823 SEQ ID No. 2699 287865 5052.4 Contigδ2 98% P 287980 SEQ ID No. 2698 288597 5050.2 Contigδ2 96% P 288634 289383 SEQ ID No. 1883 3838.3 Contig52 98% P 289389 SEQ ID No. 1885 291629 3840.1 Contigδ2 99% P 291630 SEQ ID No. 1886 292769 3841.2 Contig52 99% P 292848 295433 85%
SEQ ID No. 3341 843.2 Conti g52 m 295436 296004 99% SEQ ID No. 3342 844.2 Conti gδ2 m 296047 296436 99% SEQ ID No. 3343 845.4 Cont gδ2 m 296556 297314 98% SEQ ID No. 1002 249.4 Conti g52 P 297549 299282 98% SEQ ID No. 3071 5848.1 Cont gδ2 m 298199 298495 96% SEQ ID No. 1008 260.1 Contigδ2 P 299279 300508 99% SEQ ID No. 1014 261.2 Cont gδ2 P 300542 301417 99% SEQ ID No. 1019 262.2 Contigδ2 m 301431 302669 98% SEQ ID No. 1912 3898.2 Contigδ2 m 302744 303340 98% + SEQ ID No. 1913 3899.2 Contigδ2 m 303353 303835 98% + SEQ ID No. 1916 3901.2 Contigδ2 P 303849 304757 99% SEQ ID No. 1917 3902.3 Contigδ2 m 304877 306370 98% SEQ ID No. 1893 3857.3 Contigδ2 m 306327 307457 98% SEQ ID No. 1892 3855.1 Contigδ2 m 307661 308464 98% SEQ ID No. 315 1398.2 Contig52 m 308532 310538 97% SEQ ID No. 316 1399.3 Cont gδ2 m 310525 312153 99% SEQ ID No. 1891 3853.2 Cont gδ2 m 312450 313760 98% SEQ ID No. 1890 3861.1 Contig52 P 314121 314300 SEQ ID No. 1889 3860.1 Cont gδ2 m 314297 314740 97% SEQ ID No. 1888 3849.1 Contig52 m 314722 315453 99% SEQ ID No. 2582 4863.4 Cont gδ2 P 316679 317031 99% SEQ ID No. 2583 4864.2 Contigδ2 P 317013 317423 SEQ ID No. 2891 5463.2 Contig53 m 2 1057 99% SEQ ID No. 1486 3244.2 Cont gδ3 P 1210 1977 99% SEQ ID No. 3187 618.3 Conti'g53 P 2045 2461 100% SEQ ID No. 3188 619.1 Contigδ3 P 2371 3357 97% SEQ ID No. 3189 621.5 Contigδ3 P 3335 4654 99% SEQ ID No. 3233 687.5 Cont gδ3 P 4656 5932 98% SEQ ID No. 3234 688.1 Cont g53 P 5916 6770 99% SEQ ID No. 1485 3243.1 Contigδ3 m 6880 7188 99% + SEQ ID No. 1484 3242.3 Contig53 m 7361 10084 98% + SEQ ID No. 2989 5633.2 Contigδ3 m 10360 11202 98% SEQ ID No. 2990 5634.2 Cont g53 m 11203 11337 SEQ ID No. 2991 5636.1 Contigδ3 m 11490 11912 99% SEQ ID No. 2992 5637.1 Cont g53 P 12003 12689 99% SEQ ID No. 2516 4766.2 Cont gδ3 P 12665 13120 100%
SEQ ID No. 2517 4767.2 Contig53 P 13062 13379 100%
SEQ ID No. 2518 4768.1 Contig53 m 13467 13820 99%
SEQ ID No. 2519 4769.1 Contig53 P 14000 14419 100%
SEQ ID No. 2520 4770.1 Contig53 P 14512 14700
SEQ ID No. 2621 4771.2 Contig53 P 14837 16913 96%
SEQ ID No. 2993 5638.2 Contig53 m 16224 16634 100% +
SEQ ID No. 2864 5377.2 Contigδ3 P 16834 17466 95% +
SEQ ID No. 2863 5376.2 Contig53 m 17524 21840 99% +
SEQ ID No. 1375 3077.1 Contigδ3 m 21958 22719 97%
SEQ ID No. 1376 3078.2 Contig53 m 22682 23938 99%
SEQ ID No. 1377 3080.2 Contigδ3 m 23895 25349 100%
SEQ ID No. 1378 3081.2 Contig53 m 25561 26097 98%
SEQ ID No. 1379 3082.1 Contigδ3 m 26262 26786 98%
SEQ ID No. 1380 3083.1 Contig53 m 26732 27307 97%
SEQ ID No. 840 2251.3 Contigδ3 m 27542 28039 92%
SEQ ID No. 1381 3085.2 Contigδ3 P 28077 30950 98%
SEQ ID No. 1382 3086.1 Contigδ3 P 31106 31714 100%
SEQ ID No. 1383 3087.1 Contigδ3 P 31707 32744 99%
SEQ ID No. 1384 3088.1 Contig53 m 32771 33550 100%
SEQ ID No. 2594 488.2 Contigδ3 m 33534 34451 99%
SEQ ID No. 2603 489.1 Contigδ3 m 34530 34715 +
SEQ ID No. 2613 491.1 Contigδ3 m 34679 36737 97% +
SEQ ID No. 2620 492.4 Contig53 m 35824 36363 98%
SEQ ID No. 2852 5373.1 Contigδ3 m 36256 36521
SEQ ID No. 2625 493.4 Contigδ3 m 36559 37296 96%
SEQ ID No. 1386 3091.1 Contigδ3 m 37453 37764 100% +
SEQ ID No. 3075 5867.1 Contig53 P 37577 37750 94% +
SEQ ID No. 515 1705.2 Contigδ3 P 37874 38746 98% +
SEQ ID No. 516 1706.2 Contig53 m 38847 39317 98%
SEQ ID No. 517 1707.2 Contigδ3 m 39318 39686 100%
SEQ ID No. 1387 3093.1 Contig53 m 39710 40480 98%
SEQ ID No. 1388 3095.1 Contigδ3 m 40462 41001 98%
SEQ ID No. 1389 3096.1 Contig53 m 40977 41246 100%
SEQ ID No. 1390 3097.3 Contigδ3 m 41324 42712 99%
SEQ ID No. 1947 3960.2 Contigδ3 m 42957 43676 93%
SEQ ID No. 1945 3959.2 Contigδ3 P 43842 45401 97%
SEQ ID No.3270 742.3 Conti gδ3 P 45729 47141 + 99% SEQ ID No. 3269 741.2 Cont gδ3 P 47095 47721 - 97% SEQ ID No. 3268 740.2 Conti gδ3 P 47919 48266 - 100% SEQ ID No. 819 2213.2 Conti gδ3 m 48480 48932 + 100% SEQ ID No. 1944 3958.1 Conti gδ3 m 48942 49568 + 97% SEQ ID No. 1943 3957.1 Conti g53 m 49907 50113 + SEQ ID No. 1942 3956.2 Conti g53 P 50175 50906 98% SEQ ID No. 2929 5524.1 Conti gδ3 P 50884 51435 - 99% SEQ ID No. 2928 5623.2 Cont g53 P 51398 52447 + 97% SEQ ID No. 935 2400.2 Conti g53 P 62389 53321 - 99% SEQ ID No. 936 2401.2 Conti g53 P 53322 53972 + 98% SEQ ID No. 361 1476.2 Cont gδ3 P 54335 55246 - 100% + SEQ ID No. 360 1475.2 Conti gδ3 m 55250 55640 - 100% + SEQ ID No. 690 1989.2 Cont gδ3 P 55808 57301 - 98% SEQ ID No. 692 1990.1 Cont gδ3 P 57129 68001 - 97% SEQ ID No. 942 241.2 Conti g53 P 57980 59827 - 98% SEQ ID No. 934 240.1 Cont gδ3 P 59751 60809 - 99% SEQ ID No. 929 239.1 Cont gδ3 P 60784 61458 - 97% SEQ ID No. 923 238.1 Conti gδ3 P 61569 62993 - 98% SEQ ID No. 908 236.1 Cont gδ3 m 62888 63082 98% SEQ ID No. 898 235.2 Conti gδ3 m 63083 64465 - 98% SEQ ID No. 1299 2969.1 Conti g53 m 64558 64863 - 98% SEQ ID No. 1301 2970.2 Cont g53 m 64881 65435 - 100% SEQ ID No. 3392 916.3 Conti g53 m 65548 66546 - 98% SEQ ID No. 3393 917.3 Conti gδ3 m 66654 66896 - 100% SEQ ID No. 3394 918.3 Conti gδ3 P 66802 66963 100% SEQ ID No. 3395 919.1 Conti g53 P 67100 67537 - 92% SEQ ID No. 3396 920.2 Conti gδ3 m 67640 68068 - 97% SEQ ID No. 2437 4656.3 Cont g53 P 68083 69438 + 99% SEQ ID No. 283 1363.3 Conti gδ3 P 69533 70615 - 99% + SEQ ID No. 284 1364.2 Conti g53 P 70542 71525 - 99% + SEQ ID No. 2436 4663.2 Cont gδ3 P 71489 72391 - 98% SEQ ID No. 2435 4662.2 Conti gδ3 P 72396 72793 - 97% SEQ ID No. 2287 4432.2 Cont gδ3 m 72795 73247 - 97% SEQ ID No. 2288 4433.1 Cont gδ3 P 73488 74099 - 98% SEQ ID No. 2289 4434.1 Conti g53 m 74253 75062 - 92%
SEQ ID No. 2290 4435.4 Contig53 P 75262 77324 - 94% +
SEQ ID No. 2339 451.3 Contig53 m 78146 79099 - 97% +
SEQ ID No. 2334 460.1 Contigδ3 P 79261 79500 -
SEQ ID No. 2316 448.1 Contig53 P 79662 79931 - 100%
SEQ ID No. 2307 447.2 Contig53 P 79932 80276 - 100%
SEQ ID No. 2291 444.2 Contigδ3 P 80346 80738 - 96%
SEQ ID No. 2285 443.3 Contig53 P 80802 81404 - 99%
SEQ ID No. 1703 3562.3 Contig53 P 81579 82721 - 99%
SEQ ID No. 1702 3561.2 Contig53 P 82656 85069 - 99%
SEQ ID No. 3362 874.2 Contigδ3 P 85470 86042 + 99%
SEQ ID No. 3361 873.2 Contigδ3 P 86027 86338 + 96%
SEQ ID No. 3360 872.2 Contigδ3 P 86339 86992 - 100%
SEQ ID No. 3359 870.3 Contigδ3 P 86938 88077 + 98%
SEQ ID No. 3358 869.2 Contig53 m 87390 87602 94%
SEQ ID No. 3222 668.4 Contig53 P 88067 91228 + 97%
SEQ ID No. 3221 667.3 Contigδ3 m 89749 90402 95%
SEQ ID No. 1536 3322.2 Contig53 P 91234 92052 - 98%
SEQ ID No. 1537 3323.1 Contigδ3 P 92027 92644 - 100%
SEQ ID No. 1538 3324.1 Contigδ3 P 92655 93068 + 99%
SEQ ID No. 2217 434.3 Contigδ3 P 93248 93892 - 99%
SEQ ID No. 2209 433.1 Contigδ3 P 93919 96957 - 99%
SEQ ID No. 2200 432.1 Contigδ3 m 94806 95345 99% +
SEQ ID No. 2194 431.1 Contigδ3 m 95892 96227 99% +
SEQ ID No. 104 1071.3 Contig53 m 97056 98368 - 98%
SEQ ID No. 105 1072.4 Contigδ3 m 98314 101307 - 99%
SEQ ID No. 1539 3326.2 Contig53 m 101235 102038 - 100%
SEQ ID No. 2794 5238.1 Contigδ3 m 102305 103894 - 98%
SEQ ID No. 3411 940.4 Contigδ3 m 103921 104814 - 98%
SEQ ID No. 3412 941.3 Contigδ3 m 104792 106147 - 100%
SEQ ID No. 3413 943.3 Contigδ3 m 106148 106648 - 98%
SEQ ID No. 3414 944.3 Contig53 m 106644 107087 - 99%
SEQ ID No. 1932 3928.1 Contigδ3 P 107241 109073 - 99%
SEQ ID No. 1933 3929.2 Contig53 P 109621 111270 + 99%
SEQ ID No. 1868 3814.1 Contigδ3 m 111272 112147 + 98%
SEQ ID No. 3431 969.2 Contig53 P 112570 113343 - 99%
SEQ ID No. 3430 967.2 Contigδ3 m 113522 114547 - 99%
SEQ ID No. 3429 966.2 Contig53 m 114682 115431 99%
SEQ ID No. 3428 965.3 Contig53 P 115497 116042 + 100%
SEQ ID No. 1869 3815.2 Contig53 m 116186 116506 - 100%
SEQ ID No. 1870 3816.2 Contig53 P 116694 117437 + 99%
SEQ ID No. 1871 3818.1 Contig53 m 117666 117950 - 100%
SEQ ID No. 1872 3819.2 Contig53 m 118126 118455 - 100%
SEQ ID No. 2452 4676.2 Contig53 m 118452 119846 . 99%
SEQ ID No. 3137 6029.1 Contig53 m 120064 120221 -
SEQ ID No. 2451 4673.1 Contig53 m 120233 120505 - 100%,
SEQ ID No. 2450 4672.1 Contig53 P 120729 121451 - 99%
SEQ ID No. 2449 4671.3 Contig53 P 121408 122550 . 97%
SEQ ID No. 2722 5082.3 Contigδ3 P 122604 124115 - 98%
SEQ ID No. 141 1123.2 Contig53 P 124321 125466 - 99%
SEQ ID No. 140 1122.1 Contigδ3 P 125461 126383 + 99%
SEQ ID No. 781 2149.4 Contig53 P 126515 127813 - 99%
SEQ ID No. 2711 6068.3 Contigδ3 P 128031 128603 - 96%
SEQ ID No. 2712 5069.3 Contig53 m 128803 129348 - 97%
SEQ ID No. 3067 5829.2 Contigδ3 P 129374 130126 + 94%
SEQ ID No. 2348 4522.4 Contig53 P 130084 130770 - 95%
SEQ ID No. 2347 4521.1 Contigδ3 P 130727 131422 - 98%
SEQ ID No. 2346 4520.1 Contig53 P 131377 132636 - 99%
SEQ ID No. 362 1479.3 Contigδ3 P 132452 133864 - 97%
SEQ ID No. 364 1481.3 Contig53 P 133846 134982 - 96%
SEQ ID No. 601 1838.2 Contigδ3 m 135071 136582 + 93%
SEQ ID No. 2431 4647.2 Contig53 P 136661 137836 + 97%
SEQ ID No. 2359 4535.2 Contigδ3 m 137909 139276 + 100%
SEQ ID No. 2360 4536.1 Contig53 m 139327 140496 + 99%
SEQ ID No. 2361 4538.2 Contigδ3 m 140456 142003 - 99%
SEQ ID No. 2363 4640.2 Contig53 m 142103 142414 + 94%
SEQ ID No. 735 207.3 Contig53 m 142476 143771 + 98%
SEQ ID No. 727 206.3 Contig53 P 144091 144813 - 95%
SEQ ID No. 721 205.1 Contig53 P 144761 145621 + 97%
SEQ ID No. 717 204.1 Contig53 P 145639 147006 - 98%
SEQ ID No. 704 202.3 Contig53 P 147129 149489 + 99%
SEQ ID No. 3264 732.3 Contigδ3 P 149594 150103 + 100%
SEQ ID No. 3263 731.2 Contig53 P 150075 151127 - 87%
SEQ ID No. 3262 730.1 Contig53 P 151262 151707 97%
SEQ ID No. 3261 729.1 Contig53 P 151701 152474 99%
SEQ ID No. 3260 728.2 Contig53 P 152475 152882 99%
SEQ ID No. 1341 3024.1 Contigδ3 P 152886 154184 99%
SEQ ID No. 1340 3023.1 Contig53 P 154337 164546
SEQ ID No. 1339 3022.1 Contigδ3 P 154705 154980 100%
SEQ ID No. 233 1268.2 Contigδ3 P 156302 156252 98%
SEQ ID No. 232 1267.3 Contigδ3 P 156234 157100 97%
SEQ ID No. 231 1265.5 Contig53 I71 157090 157923 97%
SEQ ID No. 1338 3018.1 Contig53 m 158359 169219 96% +
SEQ ID No. 1337 3017.1 Contig53 m 159194 159644 + +
SEQ ID No. 1336 3016.1 Contigδ3 m 159545 160066 + +
SEQ ID No. 1335 3015.1 Contigδ3 m 159996 160766 97%
SEQ ID No. 1334 3014.1 Contigδ3 m 160720 161361 97%
SEQ ID No. 1333 3013.2 Contig53 m 161325 161795 96%
SEQ ID No. 1332 3011.2 Contigδ3 m 161789 162874 +
SEQ ID No. 2814 5288.1 Contig53 m 162784 163212 +
SEQ ID No. 2815 5289.2 Contig53 m 163191 163865 94%
SEQ ID No. 2824 5312.2 Contig53 P 164165 164791 96%
SEQ ID No. 2825 6313.1 Contig53 m 164821 165249 87%
SEQ ID No. 2607 4897.2 Contig53 m 165348 166508 96%
SEQ ID No. 2606 4895.1 Contigδ3 P 166781 167182 93%
SEQ ID No. 2605 4894.1 Contig53 P 167095 167523 96%
SEQ ID No. 671 1945.4 Contigδ3 P 167507 169294 99%
SEQ ID No. 672 1946.2 Contig53 P 169298 170029 99%
SEQ ID No. 161 1157.4 Contigδ3 P 170063 172897 98%
SEQ ID No. 1635 3468.2 Contig53 P 172891 174162 99%
SEQ ID No. 416 1559.2 Contigδ3 P 174213 175388 99%
SEQ ID No. 415 1557.3 Contig53 P 175448 176338 99% +
SEQ ID No. 905 2356.2 Contigδ3 P 176752 177408 100% +
SEQ ID No. 571 1786.2 Contigδ3 P 177979 178392 100% +
SEQ ID No. 570 1784.2 Contig53 m 178463 178897 100% +
SEQ ID No. 569 1782.3 Contigδ3 P 179471 180784 99%
SEQ ID No. 2826 5316.1 Contigδ3 P 180979 182469 99%
SEQ ID No. 2743 6123.3 Contig53 P 182641 182940 100%
SEQ ID No. 2744 6124.3 Contigδ3 m 183002 183481
SEQ ID No. 2745 5127.4 Contigδ3 m 183406 183953 98%
SEQ ID No. 1964 3982.2 Contigδ3 m 184332 184934 98%
SEQ ID No! 1963 3980.1 Contig53 P 184907 185500 98%
SEQ ID No! 1961 3979.2 Contigδ3 P 185478 187220 98%
SEQ ID No. 1960 3977.2 Contig53 m 187318 189588 97%
SEQ ID No. 372 1494.2 Contigδ3 m 189743 190537 99%
SEQ ID No. 373 1495.1 Contig53 m 190543 190902 99%
SEQ ID No. 374 1497.2 Contigδ3 m 191202 192272 98%
SEQ ID No. 2827 5317.2 Contigδ3 m 192709 193332 97%
SEQ ID No. 451 1606.2 Contigδ3 m 193404 194234 98%
SEQ ID No. 452 1607.2 Contig53 m 194228 194600
SEQ ID No. 453 1608.2 Contigδ3 P 194570 195826 99%
SEQ ID No. 2396 4597.1 Contig53 m 195867 196286 100%
SEQ ID No. 2397 4698.2 Contigδ3 P 196513 197271 99%
SEQ ID No. 2490 4729.3 Contigδ3 P 197390 198196 100%
SEQ ID No. 2489 4727.1 Contig53 P 198224 198724 99%
SEQ ID No. 2488 4726.1 Contigδ3 P 198889 199224 98%
SEQ ID No. 2487 4725.2 Contig53 m 199470 200573 99%
SEQ ID No. 2486 4724.2 Contigδ3 P 200568 201215 100%
SEQ ID No. 2485 4721.2 Contig53 P 201197 201745 99%
SEQ ID No. 2593 4878.2 Contigδ3 m 202045 203496 99%
SEQ ID No. 1228 2859.2 Contigδ3 P 203633 204940 99%
SEQ ID No. 1227 2868.1 Contig53 m 205210 205464 +
SEQ ID No. 3321 81.3 Contigδ3 m 205742 205948 +
SEQ ID No. 3314 80.3 Contigδ3 P 205998 206204 +
SEQ ID No. 1159 2743.2 Contigδ3 P 206346 206567 +
SEQ ID No. 1160 2746.1 Contig53 P 206934 207152 + +
SEQ ID No. 3235 692.1 Contigδ3 P 209993 210160 97% +
SEQ ID No. 3236 693.1 Contig53 m 210303 210845 96%
SEQ ID No. 3237 694.3 Contigδ3 m 210846 212210 98%
SEQ ID No. 1161 2747.1 Contig53 m 212435 213457 96%
SEQ ID No. 1162 2748.1 Contigδ3 P 213767 214243 97%
SEQ ID No. 1163 2749.4 Contig53 m 214284 215646 99%
SEQ ID No. 552 1760.4 Contigδ3 m 215547 216278 99%
SEQ ID No. 550 1769.4 Contigδ3 P 216391 217173 99%
SEQ ID No. 1720 3691.2 Contig53 P 217448 218038 100%
SEQ ID No. 1721 3592.1 Contig53 P 218007 218450 - 99%
SEQ ID No. 1722 3593.1 Contigδ3 P 218407 219393 - 98%
SEQ ID No. 1723 3594.1 Contig53 m 219607 219902 +
SEQ ID No. 950 2421.2 Contig53 m 220002 221526 - 97%
SEQ ID No. 1724 3596.1 Contig53 m 221578 221850 - 97%
SEQ ID No. 1725 3696.1 Contigδ3 P 221926 222321 - 100%
SEQ ID No. 1726 3597.3 Contigδ3 m 222331 222921 - 98%
SEQ ID No. 1727 3598.3 Contig53 P 223164 223358 +
SEQ ID No. 1728 3599.4 Contigδ3 P 223351 224187 - 99%
SEQ ID No. 3039 5741.1 Contig53 m 224214 224969 - 99%
SEQ ID No. 1106 2657.3 Contigδ3 P 225046 227253 - 98%
SEQ ID No. 1105 2656.1 Contig53 P 227254 227673 - 97%
SEQ ID No. 1104 2654.1 Contigδ3 P 227496 227683 - 90% +
SEQ ID No. 1103 2653.1 Contig53 m 227848 228192 - 93% +
SEQ ID No. 2569 482.2 Contigδ3 m 228193 230004 - 98%
SEQ ID No. 2550 481.1 Contig53 P 230365 230837 - 99%
SEQ ID No. 2541 480.3 Contigδ3 P 230827 232278 - 99%
SEQ ID No. 677 1951.4 Contig53 P 232262 233032 - 100%
SEQ ID No. 676 1950.2 Contigδ3 P 233014 234315 - 100%
SEQ ID No. 1606 342.2 Contigδ3 P 234284 235549 - 100%
SEQ ID No. 1612 343.1 Contigδ3 P 235534 235995 - 100%
SEQ ID No. 1616 344.1 Contig53 P 236025 236402 - 100%
SEQ ID No. 1630 346.2 Contigδ3 P 236327 237358 - 100%
SEQ ID No. 1636 347.1 Contig53 m 237431 238357 - 100%
SEQ ID No. 1649 349.2 Contigδ3 m 238592 239380 - 100%
SEQ ID No. 1102 2652.2 Contigδ3 m 239239 240562 - 100%
SEQ ID No. 1101 2650.1 Contigδ3 P 241173 242501 + - 100%
SEQ ID No. 1021 2521.4 Contig53 m 242561 243616 - 100%
SEQ ID No. 1022 2522.1 Contigδ3 m 243808 244155 + - 100%
SEQ ID No..1023 2623.1 Contig53 P 244324 244743 - 100%
SEQ ID No. 1024 2624.2 Contig53 m 244779 245492 - 97%
SEQ ID No. 1100 2649.1 Contig53 P 245702 246898 +
SEQ ID No. 3210 650.3 Contigδ3 P 246861 248021 +
SEQ ID No. 3211 651.2 Contig53 m 248091 248534 +
SEQ ID No. 3212 652.1 Contigδ3 P 248604 249497 - 96%
SEQ ID No. 1099 2648.2 Contig53 P 249860 250981 + - 99%
SEQ ID No. 1098 2647.2 Contig53 m 261141 251743 100%
SEQ ID No. 1097 2646.1 Contigδ3 P 251907 252410 100%
SEQ ID No. 1096 2645.1 Contig53 m 252611 253491 100%
SEQ ID No. 248 1288.2 Contigδ3 m 253575 255041 100%
SEQ ID No. 656 1915.3 Contig53 m 255160 257109 100%
SEQ ID No. 654 1913.2 Contigδ3 m 257214 257630 100%
SEQ ID No. 1095 2644.1 Contig53 P 257737 258120 + 100%
SEQ ID No. 1094 2643.1 Contigδ3 P 258216 269628 100%
SEQ ID No. 339 1434.3 Contigδ3 m 259654 261867 100%
SEQ ID No. 697 2000.1 Contigδ3 m 261939 262439 100%
SEQ ID No. 1249 290.2 Contig53 m 262400 265921 + 100%
SEQ ID No. 1229 286.2 Contigδ3 m 265922 266464 + 100%
SEQ ID No. 1221 284.2 Contigδ3 m 266431 267564 100%
SEQ ID No. 1215 283.1 Contigδ3 m 267495 268037 100%
SEQ ID No. 1208 282.2 Contigδ3 m 268047 268784 100%
SEQ ID No. 632 1882.2 Contigδ3 m 268976 269923 100%
SEQ ID No. 633 1883.4 Contigδ3 m 270013 271392 100%
SEQ ID No. 2759 5169.2 Contigδ3 P 271638 273131 100%
SEQ ID No. 660 1926.2 Contigδ3 P 273283 273951 98%
SEQ ID No. 661 1928.2 Contigδ3 P 273981 275270 99%
SEQ ID No. 3322 813.2 Contig53 P 275271 276566 100%
SEQ ID No. 3323 815.2 Contigδ3 P 276554 277786 97%
SEQ ID No. 1895 3860.2 Contigδ3 P 277948 278538 100%
SEQ ID No. 1894 3859.2 Contig53 P 278522 279865 99%
SEQ ID No. 1127 2690.2 Contigδ3 P 280065 281204 97% +
SEQ ID No. 1125 2688.2 Contigδ3 P 281205 281639 + +
SEQ ID No. 2758 5156.1 Contigδ3 m 281789 282058 + +
SEQ ID No. 694 1998.3 Contig53 m 282043 283029 98%
SEQ ID No. 3037 5739.3 Contig53 P 283472 288685 +
SEQ ID No. 3410 94.7 Contig53 P 288516 295643 +
SEQ ID No. 1085 2624.1 Contig54 P 3 554 +
SEQ ID No. 67 10.1 Contig54 P 888 1235 +
SEQ ID No. 3382 9.1 Contig54 P 1049 1562 +
SEQ ID No. 3242 7.1 Contig54 P 1711 5829 +
SEQ ID No. 1977 4.1 Contig54 P 5766 6818 +
SEQ ID No. 1322 3.1 Contig54 P 6822 7124 +
SEQ ID No. 695 2.1 Contig54 P 7066 7698 +
SEQ ID No. 3194 63.1 Contig54 m 9066 9281 + +
SEQ ID No. 3167 61.1 Contig54 P 9769 10068 + +
SEQ ID No. 3126 60.1 Contigδ4 m 9838 10008 + +
SEQ ID No. 3091 59.1 Contig54 P 10665 11108 +
SEQ ID No. 3052 58.1 Contigδ4 m 11098 11802 +
SEQ ID No. 3025 57.1 Contig54 P 11919 12803 91 % +
SEQ ID No. 2919 56.1 Contigδ4 P 12574 13131 96% +
SEQ ID No. 2775 62.1 Contig54 P 13132 13347
SEQ ID No. 2666 60.1 Contigδ4 P 13299 13646 82%
SEQ ID No. 2540 48.1 Contig54 P 13647 13937
SEQ ID No. 2466 47.1 Contigδ4 P 13831 16404 94%
SEQ ID No. 2399 46.1 Contig54 P 16389 17111 93%
SEQ ID No. 2333 45.1 Contigδ4 P 17248 17664 80%
SEQ ID No. 2266 44.1 Contigδ4 P 17665 18708 94%
SEQ ID No. 2187 43.1 Contigδ4 P 18684 19421 95%
SEQ ID No. 2122 42.1 Contigδ4 P 19373 20170 95%
SEQ ID No. 1914 39.1 Contigδ4 P 20125 21258 93%
SEQ ID No. 1801 37.1 Contig54 P 21218 22264 91 %
SEQ ID No. 1729 36.1 Contigδ4 P 22191 24158 98%
SEQ ID No. 1654 35.1 Contig54 P 24187 25053 98% +
SEQ ID No. 1592 34.1 Contigδ4 P 25266 26048 + +
SEQ ID No. 1522 33.1 Contig54 m 26172 28868 +
SEQ ID No. 1323 30.1 Contigδ4 P 28991 29266 +
SEQ ID No. 1248 29.1 Contigδ4 P 29232 29522 +
SEQ ID No. 1131 27.1 Contigδ4 P 29610 30263 +
SEQ ID No. 1072 26.1 Contig54 P 30264 31076 +
SEQ ID No. 1007 25.1 Contigδ4 m 31103 31582 +
SEQ ID No. 933 24.1 Contigδ4 m 31557 33023 +
SEQ ID No. 696 20.1 Contigδ4 m 32977 33345 +
SEQ ID No. 644 19.1 Contigδ4 m 33353 34066 +
SEQ ID No. 580 18.1 Contig54 P 34124 34408 +
SEQ ID No. 446 16.1 Contigδ4 P 34418 34645 +
SEQ ID No. 375 15.1 Contigδ4 P 34591 35802 +
SEQ ID No. 254 13.1 Contig54 P 35966 36853 +
SEQ ID No. 126 11.2 Contιgδ4 P 36783 37802 +
SEQ ID No. 1084 2622.1 Contig54 P 38133 40073 +
SEQ ID No. 1076 2605.3 Contig54 P 40025 42568 +
SEQ ID No. 1083 2620.1 Contig54 P 42674 43009 97%
SEQ ID No. 1081 2619.1 Contig54 P 42970 43425 +
SEQ ID No. 1080 2616.1 Contig54 P 43401 44285 +
SEQ ID No. 308 139.5 Contig54 P 44404 48882 +
SEQ ID No. 1079 2611.1 Contigδ4 P 49022 49240 83% +
SEQ ID No. 1078 2609.2 Contig54 P 49931 50938 94% +
SEQ ID No. 1077 2607.2 Contigδ4 P 51047 52288 98%
SEQ ID No. 1648 3489.2 Contig54 m 52297 53340 94%
SEQ ID No. 432 1582.2 Contig54 P 53466 53990 98%
SEQ ID No. 431 1581.4 Contig54 m 54089 55366 87%
SEQ ID No. 160 1156.3 Contig54 m 55687 56997
SEQ ID No. 159 1153.1 Contig54 m 57180 57608 98%
SEQ ID No. 158 1152.2 Contig54 m 57786 58682 92%
SEQ ID No. 1647 3488.1 Contig54 m 58908 61016 86% +
SEQ ID No. 1646 3486.1 Contig54 m 61261 61521 +
SEQ ID No. 1645 3485.1 Contig54 P 61495 62223 87% +
SEQ ID No. 1644 3484.1 Contig54 m 62317 62967 93% +
SEQ ID No. 1643 3483.3 Contig54 m 63171 63917 92%
SEQ ID No. 588 1809.4 Contig54 m 64253 66679 84%
SEQ ID No. 1642 3481.2 Contigδ4 m 66646 66918 100%
SEQ ID No. 1641 3480.4 Contig54 m 66923 68722 98%
SEQ ID No. 3334 833.3 Contigδ4 P 68959 71343 93%
SEQ ID No. 757 2116.2 Contig54 m 71452 71955 96%
SEQ ID No. 2659 499.2 Contig54 m 72017 73159 97%
SEQ ID No. 2667 500.2 Contig54 P 73382 74464 98%
SEQ ID No. 2672 601.2 Contig54 P 74623 75396 96% +
SEQ ID No. 2677 502.3 Contig54 m 75598 76065 +
SEQ ID No. 2682 503.3 Contig54 P 76245 77588 98%
SEQ ID No. 1435 3161.1 Contig54 m 77714 78490 99%
SEQ ID No. 1434 3160.1 Contig54 m 78711 79331 87%
SEQ ID No. 90 1049.2 Contig54 P 79487 80278 96%
SEQ ID No. 91 1050.4 Contig54 P 80420 83893 98%
SEQ ID No. 915 2368.5 Contig54 m 83988 85529 94%
SEQ ID No. 564 1778.5 Contig54 P 85555 86043 100%
SEQ ID No. 563 1775.3 Contigδ4 P 86468 88030 94%
SEQ ID No. 1860 3801.2 Contig54 m 88195 89649 + 98%
SEQ ID No. 1859 3800.4 Contig54 m 89808 90146 100%
SEQ ID No. 3307 791.5 Contigδ4 m 90299 91600 97%
SEQ ID No. 1858 3798.1 Contig54 m 91480 91875 99%
SEQ ID No. 1857 3797.1 Contigδ4 m 92077 93171 96%
SEQ ID No. 1856 3796.3 Contig54 m 93107 93736 99%
SEQ ID No. 1855 3793.3 Contigδ4 P 93867 94790 + 94%
SEQ ID No. 1854 3792.1 Contig54 m 94897 95136 + +
SEQ ID No. 1853 3791.1 Contigδ4 m 95344 96318 94% +
SEQ ID No. 1852 3789.2 Contig54 m 96578 97606 98%
SEQ ID No. 1851 3788.1 Contigδ4 m 97597 98373 99%
SEQ ID No. 1850 3785.2 Contigδ4 m 98361 99386 99%
SEQ ID No. 1849 3784.2 Contig54 m 99175 100311 98%
SEQ ID No. 1848 3783.1 Contigδ4 m 100438 100734 94%
SEQ ID No. 904 2355.2 Contig54 m 100931 101824 97%
SEQ ID No. 1847 3780.2 Contigδ4 m 102158 102685 95%
SEQ ID No. 379 1505.4 Contigδ4 P 102899 105229 96%
SEQ ID No. 380 1507.3 Contigδ4 P 105338 106840 98%
SEQ ID No. 1626 3454.1 Contig54 P 106841 107404 91%
SEQ ID No. 1625 3453.1 Contig54 P 107695 109209 90%
SEQ ID No. 1624 3451.1 Contig54 P 109351 110169 95%
SEQ ID No. 1623 3450.2 Contigδ4 P 110251 111336 98%
SEQ ID No. 2750 5135.2 Contig54 m 111413 112909 93% +
SEQ ID No. 1035 2545.2 Contig54 m 113096 114103 90% +
SEQ ID No. 1036 2546.4 Contigδ4 m 113946 115461 94%
SEQ ID No. 1935 3933.3 Contig54 P 115898 117001 96%
SEQ ID No. 229 1261.2 Contigδ4 P 117232 118920 99%
SEQ ID No. 230 1264.3 Contigδ4 m 118978 120174 + 95%
SEQ ID No. 2322 4485.2 Contigδ4 P 120515 121345 + 98%
SEQ ID No. 2321 4484.1 Contig54 P 121446 122228 97%
SEQ ID No. 3378 895.3 Contigδ4 P 122576 123634 98%
SEQ ID No. 3377 894.3 Contig54 m 123736 124005 98%
SEQ ID No. 823 2220.2 Contigδ4 P 124301 124963 99%
SEQ ID No. 821 2219.2 Contig54 m 124648 125064 98%
SEQ ID No. 2199 4319.2 Contigδ4 m 125030 125764 + 98%
SEQ ID No. 144 1 126.3 Contig54 P 126926 127218 SEQ ID No. 143 1125.2 96% Contig54 P 127188 128384 SEQ ID No. 142 1124.2 97% Contig54 P 128626 129426 SEQ ID No. 2198 4316.4 97% Contig54 m 129491 131527 SEQ ID No. 2857 5381.1 97% Contig54 + P 129885 130109 SEQ ID No. 2338 4508.4 100% Contig54 + P 131646 132248 SEQ ID No. 129 1105.2 98% Contig54 m 132369 134687 SEQ ID No. 2129 4211.2 96% Contig54 m 134827 136197 SEQ ID No. 2130 4212.1 92% Contig54 P 136221 137318 SEQ ID No. 151 1137.2 97% Contig54 P 137371 137796 SEQ ID No. 150 1136.3 98% Contigδ4 P 137907 139430 SEQ ID No. 2131 4214.3 99% Contigδ4 P 139523 1411 15 SEQ ID No. 2886 544.2 96% Contigδ4 P 141103 142440 SEQ ID No. 2883 543.1 96% Contig54 P 142846 143115 SEQ ID No. 986 2468.2 Contigδ4 100% m 143563 144497 SEQ ID No. 986 2469.2 97% Contig54 m 144767 146230 SEQ ID No. 988 2471.3 99% Contigδ4 P 146263 147339 SEQ ID No. 599 1832.4 98% Contig54 P 147324 147938 SEQ ID No. 598 1831.3 98% Contigδ4 P 147869 149143 SEQ ID No. 1816 3721.1 97% Contig54 P 149133 149618 SEQ ID No. 1633 3465.2 100% Contigδ4 P 149702 151381 SEQ ID No. 1632 3463.1 100% Contigδ4 P 161330 152202 SEQ ID No. 1631 3461.4 100% Contigδ4 P 162359 154266 SEQ ID No. 865 2293.5 97% Contig54 P 154352 SEQ ID No. 322 156745 1409.3 97% Contigδ4 P 157120 157902 SEQ ID No. 324 1410.2 99% + Contig54 m 158050 158997 SEQ ID No. 1629 3459.3 98% + Contigδ4 m 159106 160590 SEQ ID No. 1628 3456.4 98% Contigδ4 m 160503 161009 SEQ ID No. 1627 3455.3 Contig54 m 161013 162041 SEQ ID No. 449 1602.2 97% Contigδ4 P 162071 163246 SEQ ID No. 448 1601.3 97% Contig54 P 163332 164519 SEQ ID No. 2813 5282.3 98% Contigδ4 P 164626 164874 SEQ ID No. 3135 6019.1 Contig54 P 164875 165402 SEQ ID No. 1844 3771.2 98% Contigδ4 m 165491 166123 SEQ ID No. 3347 854.3 99% Contig54 m 166111 166785 SEQ ID No. 3050 5780.1 96% Contigδ4 m 166583 166912
SEQ ID No. 3346 853.1 Contigδ4 m 166886 167625 98%
SEQ ID No. 3345 852.3 Contig54 m 167598 168224 98%
SEQ ID No. 1845 3772.2 Contigδ4 m 168194 169256 97%
SEQ ID No. 127 1101.4 Contig54 m 169230 170324 97%
SEQ ID No. 128 1102.2 Contigδ4 m 170325 171641 96%
SEQ ID No. 843 2255.2 Contig54 m 171626 172522 96%
SEQ ID No. 1846 3778.2 Contigδ4 m 172504 172809 98%
SEQ ID No. 2947 5550.3 Contigδ4 P 173129 174709 98%
SEQ ID No. 2948 6562.2 Contigδ4 P 174713 175861 99%
SEQ ID No. 2275 4409.1 Contig54 m 176087 176728 100%
SEQ ID No. 2274 4408.1 Contigδ4 m 176920 177180 98%
SEQ ID No. 2273 4407.1 Contig54 P 176950 177201
SEQ ID No. 2272 4406.1 Contigδ4 P 177700 178284 99%
SEQ ID No. 2271 4405.1 Contigδ4 P 178381 178899 98%
SEQ ID No. 2270 4404.1 Contig54 m 178929 179393 94%
SEQ ID No. 2269 4403.2 Contig54 P 179417 180214 91 %
SEQ ID No. 2268 4402.2 Contig54 P 180142 180759 92%
SEQ ID No. 1070 2598.3 Contig54 m 180883 181581 + 97%
SEQ ID No. 2741 5115.2 Contigδ4 P 181805 182791 99%
SEQ ID No. 2742 5116.3 Contig54 P 182843 183412 97%
SEQ ID No. 2949 5554.2 Contig54 P 183792 184052 + +
SEQ ID No. 3134 6018.1 Contig54 m 184053 184367 + +
SEQ ID No. 2785 5217.1 Contigδδ P 45 509 +
SEQ ID No. 1107 2659.2 Contigδδ P 502 996 +
SEQ ID No. 1108 2660.1 Contigδδ P 983 1276 +
SEQ ID No. 1109 2661.2 Contigδδ P 1221 1838 +
SEQ ID No. 1110 2662.2 Contigδδ P 1823 2554 + + +
SEQ ID No. 2574 485.3 Contigδδ P 2536 4023 + + + +
SEQ ID No. 2580 486.2 Contigδδ P 3956 4243 + +
SEQ ID No. 1032 254.2 Contigδδ P 4093 6849 +
SEQ ID No. 1038 256.1 Contigδδ P 6821 7174 + +
SEQ ID No. 1045 256.1 Contigδδ P 7135 7797 +
SEQ ID No. 1058 258.2 Contigδδ P 7683 8804 +
SEQ ID No. 698 2006.2 Contigδδ P 8818 9483 + +
SEQ ID No. 280 1344.3 Contigδδ P 9401 10501 + +
SEQ ID No. 2786 5219.1 Contigδδ P 10464 11285 +
SEQ ID No. 279 1342.3 Contig55 P 11279 12070 +
SEQ ID No. 722 2051.2 Contigδδ P 12067 12564 +
SEQ ID No. 263 1299.3 Contigδδ P 12561 13958 + +
SEQ ID No. 3010 567.5 Contigδδ P 13877 16723 + +
SEQ ID No. 818 2212.4 Contigδδ P 16754 18805 + +
SEQ ID No. 445 1599.4 Contigδδ P 18806 24001 + +
SEQ ID No. 2787 5220.1 Contigδδ P 23872 24702 +
SEQ ID No. 1111 2665.1 Contigδδ P 24873 25163 + +
SEQ ID No. 1112 2666.1 Contigδδ m 25296 25622 + +
SEQ ID No. 1113 2667.1 Contigδδ m 25607 26068 + +
SEQ ID No. 3256 717.2 Contigδδ P 26412 26885 + +
SEQ ID No. 3254 716.1 Contigδδ P 26900 27130 + +
SEQ ID No. 3253 715.1 Contigδδ P 27284 27616 + +
SEQ ID No. 3252 713.2 Contigδδ P 27686 28033 + +
SEQ ID No. 3069 5844.1 Contigδδ m 28321 28734 +
SEQ ID No. 2320 4483.2 Contigδδ m 28848 29474 +
SEQ ID No. 2319 4482.1 Contigδδ P 29686 30615 +
SEQ ID No. 2318 4481.1 Contigδδ m 30697 31317 +
SEQ ID No. 1752 3632.3 Contigδδ m 31318 34410 +
SEQ ID No. 836 2244.2 Contigδδ P 34509 35120 +
SEQ ID No. 837 2245.2 Contigδδ P 35114 36196 +
SEQ ID No. 838 2247.5 Contigδδ P 36168 37979 +
SEQ ID No. 3301 783.2 Contigδδ m 38018 38377 +
SEQ ID No. 3302 785.4 Contigδδ m 38404 39363 + +
SEQ ID No. 3070 5846.2 Contigδδ m 39236 39436 + +
SEQ ID No. 1751 3631.4 Contigδδ m 39466 39723 + +
SEQ ID No. 3133 6016.1 Contigδδ m 39777 40088 +
SEQ ID No. 3132 6015.1 Contigδδ P 40166 40540 + +
SEQ ID No. 2739 5113.3 Contig55 P 40350 40982 + +
SEQ ID No. 2740 5114.3 Contigδδ P 40999 41910 + +
SEQ ID No. 1420 3138.2 Contigδδ P 41939 42838 + +
SEQ ID No. 1419 3137.1 Contigδδ P 42851 43375 + +
SEQ ID No. 1418 3136.1 Contigδδ P 43426 43786 +
SEQ ID No. 3313 799.3 Contigδδ P 43996 45474 + +
SEQ ID No. 3312 798.2 Contig55 P 45371 47254 + +
SEQ ID No. 3311 796.2 Contigδδ P 47244 48029 + +
SEQ ID No. 745 2087.2 Contigδδ P 48030 48989 +
SEQ ID No. 803 2180.2 Contigδδ P 48966 50470 +
SEQ ID No. 1417 3135.1 Contigδδ P 50443 50679 +
SEQ ID No. 1416 3134.1 Contigδδ P 50672 51244 +
SEQ ID No. 554 1762.2 Contigδδ P 51309 52265 +
SEQ ID No. 553 1761.4 Contigδδ P 52416 55592 +
SEQ ID No. 426 1576.2 Contigδδ P 56896 56345 +
SEQ ID No. 1553 3346.2 Contigδδ P 56297 58270 +
SEQ ID No. 1564 3347.1 Contigδδ P 58332 59138 +
SEQ ID No. 1556 3348.2 Contigδδ P 59139 60737 +
SEQ ID No. 1566 3349.3 Contigδδ P 60738 61637 +
SEQ ID No. 1568 3350.3 Contig55 P 61771 62142 +
SEQ ID No. 1559 3351.3 Contig55 P 62162 63334 +
SEQ ID No. 1560 3352.1 Contig55 m 63363 63833 +
SEQ ID No. 1561 3353.1 Contig55 P 63922 65490 + +
SEQ ID No. 1562 3354.1 Contig55 P 65511 66116 + +
SEQ ID No. 3426 960.2 Contigδδ P 66069 66436 +
SEQ ID No. 3424 959.2 Contigδδ P 66489 66674 +
SEQ ID No. 3423 957.2 Contigδδ m 66768 67700 +
SEQ ID No. 3422 956.2 Contigδδ m 67701 68516 +
SEQ ID No. 1563 3356.2 Contigδδ m 68675 69007 +
SEQ ID No. 1564 3356.2 Contigδδ m 68952 69212 +
SEQ ID No. 1565 3357.2 Contigδδ m 69452 69934 +
SEQ ID No. 2757 5154.2 Contigδδ m 70538 71368 +
SEQ ID No. 2756 5153.1 Contigδδ m 71343 71813 +
SEQ ID No. 1054 2673.3 Contigδδ m 71767 72372 +
SEQ ID No. 1241 2881.2 Contigδδ m 72633 73565 + +
SEQ ID No. 1240 2879.3 Contigδδ m 73627 74961 + +
SEQ ID No. 1239 2878.2 Contigδδ P 75141 75842 +
SEQ ID No. 290 1362.3 Contigδδ P 76079 77905 +
SEQ ID No. 291 1364.2 Contigδδ m 77909 79549 +
SEQ ID No. 1238 2877.2 Contigδδ m 79612 80280 +
SEQ ID No. 800 2175.3 Contigδδ m 80356 80577 +
SEQ ID No. 799 2174.1 Contigδδ m 80578 80886 +
SEQ ID No. 110 1078.2 Contigδδ P 81022 82008 +
SEQ ID No. 109 1077.1 Contigδδ P 82086 82808 +
SEQ ID No. 108 1076.3 Conti g55 P 82869 83252 + SEQ ID No. 798 2173.2 Conti g55 P 83159 83683 + SEQ ID No. 797 2172.2 Conti g55 P 83680 84168 + SEQ ID No. 1237 2874.1 Conti gδδ P 84158 84934 + SEQ ID No. 1236 2873.1 Cont gδδ m 84965 85960 + SEQ ID No. 3285 761.2 Cont gδδ m 85953 86924 + + SEQ ID No. 3286 762.2 Conti gδδ P 86954 87676 + + SEQ ID No. 3287 764.1 Conti gδδ P 87747 88268 + + SEQ ID No. 3288 765.1 Conti gδδ m 88340 89080 + SEQ ID No. 3289 766.3 Cont gδδ m 89231 90691 + + SEQ ID No. 3371 886.3 Cont gδδ m 90736 91527 + + SEQ ID No. 3370 885.2 Conti gδδ P 91494 93173 + SEQ ID No. 3369 884.3 Conti gδδ P 93167 94135 + SEQ ID No. 2367 4549.2 Conti gδδ P 94210 95664 + SEQ ID No. 213 1237.2 Conti gδδ P 95677 96117 + SEQ ID No. 212 1236.2 Conti gδδ m 96236 97093 96% SEQ ID No. 211 1235.3 Conti gδδ m 97094 97873 + SEQ ID No. 2755 5152.1 Conti gδδ m 97828 98214 + SEQ ID No. 2754 6151.1 Conti gδδ m 98262 99167 + SEQ ID No. 610 1848.4 Contigδδ m 99168 100433 + SEQ ID No. 611 1849.3 Contigδδ m 100339 100893 + SEQ ID No. 613 1852.2 Contigδδ m 100904 101302 + SEQ ID No. 392 1522.2 Contigδδ m 101257 102432 + SEQ ID No. 393 1523.2 Cont gδδ m 102483 102815 + SEQ ID No. 394 1524.3 Cont gδδ m 102816 103892 + SEQ ID No. 844 2258.2 Contigδδ m 104112 105362 + SEQ ID No. 846 2260.2 Contigδδ m 105337 106014 + SEQ ID No. 1344 3030.2 Contigδδ m 106015 106557 + SEQ ID No. 1345 3031.2 Contigδδ m 106523 106960 + SEQ ID No. 1346 3035.2 Contigδδ P 107019 107624 + SEQ ID No. 3215 659.2 Contigδδ m 107629 108609 + SEQ ID No. 3216 660.1 Contigδδ m 108579 109169 + SEQ ID No. 3217 661.3 Contigδδ m 109132 110268 + SEQ ID No. 1347 3036.2 Cont gδδ m 110272 110826 + SEQ ID No. 801 2176.3 Cont gδδ m 110897 111364 + SEQ ID No. 802 2177.2 Contigδδ P 111374 112039 +
SEQ ID No. 3265 737.1 Contig55 m 112373 113413 + SEQ ID No. 3266 738.2 Contig55 m 113352 113906 + SEQ ID No. 3267 739.4 Contigδδ m 113977 114864 + SEQ ID No. 2634 4947.2 Contigδδ P 115163 116416 + SEQ ID No. 2633 4945.2 Contigδδ P 116406 117425 + SEQ ID No. 2719 5078.2 Contigδδ P 117433 118752 + SEQ ID No. 2721 5081.3 Contigδδ P 118891 119469 + SEQ ID No. 2720 5080.4 Contig55 P 119493 120956 + SEQ ID No. 2737 5106.3 Contig55 m 120958 122001 + SEQ ID No. 2738 5108.2 Contigδδ + P 122110 122373 + SEQ ID No. 2753 5147.1 + Contιg55 P 122374 123642 + SEQ ID No. 1656 3500.3 Contigδδ + P 123593 124777 + SEQ ID No. 1653 3499.1 + Contigδδ P 124882 125178 + SEQ ID No. 205 1226.2 Contigδδ + P 125159 126205 + SEQ ID No. 204 1225.2 Contigδδ P 126202 127263 + SEQ ID No. 856 2275.3 + Contigδδ P 127253 128290 + SEQ ID No. 857 2277.2 + Contig55 P 128618 128899 + SEQ ID No. 171 1171.3 Contigδδ P 128900 129670 + SEQ ID No. 172 1172.2 Contigδδ P 129671 130108 + SEQ ID No. 1652 3498.1 Contigδδ + P 130156 130584 + SEQ ID No. 3186 616.4 Contigδδ + P 130624 131685 + SEQ ID No. 3100 591.3 Contigδδ P 131637 132476 + SEQ ID No. 2834 5328.1 Contigδδ m 132518 133048 + SEQ ID No. 2127 4209.2 Contigδδ m 133049 134221 + SEQ ID No. 868 2298.2 Contigδδ m 134262 135566 SEQ ID No. 79 1035.3 95% Contigδδ P 136010 137455 SEQ ID No. 2311 4473.4 97% Contigδδ m 137527 138237 SEQ ID No. 2312 4474.1 99% Contigδδ m 138206 138715 SEQ ID No. 2313 4475.1 100% Contigδδ P 138772 139452 SEQ ID No. 2314 4477.2 98% Contigδδ P 139582 142005 SEQ ID No. 2315 4479.2 98% Contigδδ P 142158 142640 SEQ ID No. 2317 4480.4 99% Contigδδ m 142691 143992 SEQ ID No. 2727 5090.2 97% Contigδδ m 143953 145032 SEQ ID No. 2725 5088.2 99% Contig55 m 145313 146738 + SEQ ID No. 3006 5660.1 Contig55 P 145939 146925 + SEQ ID No. 2701 5056.2 Contigδδ P 147059 147427 +
SEQ ID No. 2702 5058.2 Contig55 m 147683 147949 +
SEQ ID No. 2703 5059.3 Contig55 m 147916 148292 +
SEQ ID No. 3136 602.4 Contig55 m 148411 149406 98%
SEQ ID No. 3142 604.3 Contig55 m 149666 150327 96%
SEQ ID No. 2414 4623.2 Contig55 m 150381 150938 94%
SEQ ID No. 2700 5054.2 Contig55 m 150905 151864 95%
SEQ ID No. 1982 4004.2 Contigδδ m 151866 153652 98%
SEQ ID No. 1981 4003.1 Contigδδ m 153639 155102 96%
SEQ ID No. 202 1223.2 Contigδδ P 155439 156341 80%
SEQ ID No. 203 1224.2 Contigδδ m 156401 157246 + 97%
SEQ ID No. 1980 4002.2 Contigδδ m 157251 159182 + 95%
SEQ ID No. 1979 4000.2 Contigδδ m 159161 159628 97%
SEQ ID No. 2192 4307.2 Contigδδ m 159772 161094 95% +
SEQ ID No. 847 2261.3 Contig55 m 161261 162034 93% +
SEQ ID No. 124 1098.3 Contigδδ m 162153 163196 97%
SEQ ID No. 125 1099.2 Contigδδ P 163387 164445 94%
SEQ ID No. 2191 4305.2 Contigδδ m 164470 164892 90%
SEQ ID No. 2962 5582.4 Contigδδ m 165100 167310 94%
SEQ ID No. 2041 4088.2 Contigδδ P 167432 168376 97%
SEQ ID No. 2040 4087.2 Contig55 m 168470 169795 97%
SEQ ID No. 134 1111.3 Contigδδ m 169738 170574 98%
SEQ ID No. 132 1109.3 Contigδδ m 170559 171554 97%
SEQ ID No. 131 1107.2 Contigδδ m 171541 172536 97%
SEQ ID No. 2039 4084.2 Contigδδ P 172650 173840 97%
SEQ ID No. 2038 4083.3 Contigδδ P 173810 174493 98%
SEQ ID No. 427 1577.6 Contigδδ m 174673 177582 95%
SEQ ID No. 2210 4332.1 Contig55 P 177722 178267 98%
SEQ ID No. 2211 4333.1 Contig55 P 178294 179169 95%
SEQ ID No. 2212 4334.1 Contigδδ P 179170 180063 98%
SEQ ID No. 2213 4336.2 Contigδδ P 180168 181493 100%
SEQ ID No. 2421 4632.1 Contig55 m 181506 182885 99%
SEQ ID No. 2422 4633.2 Contigδδ P 182895 183470 100%
SEQ ID No. 2423 4634.2 Contig55 m 183488 185486 97%
SEQ ID No. 2112 4181.1 Contigδδ m 185668 186306
SEQ ID No. 903 2354.2 Contigδδ m 186307 186954 99%
SEQ ID No. 543 1748.3 Contig55 m 187301 188650 96%
SEQ ID No. 544 1749.2 Conti gδδ m 188742 189518 91 % SEQ ID No. 2110 4179.2 Conti gδδ m 189807 191867 - SEQ ID No. 252 1295.1 Conti gδδ m 191978 192574 + 96% SEQ ID No. 251 1294.2 Conti g55 m 192663 193086 - 98% SEQ ID No. 250 1293.4 Cont gδδ m 193087 193707 + 96% SEQ ID No. 2214 4337.3 Cont gδδ m 193708 194001 + 97% SEQ ID No. 2215 4338.3 Conti gδδ m 194087 194995 - 97% SEQ ID No. 2216 4339.1 Conti g55 m 194986 196857 - 97% SEQ ID No. 2218 4340.2 Cont gδδ m 196896 198794 - 97% SEQ ID No. 2859 5386.1 Conti gδδ m 198790 199812 - 97% SEQ ID No. 2532 4786.2 Conti gδδ m 199797 201254 + 98% SEQ ID No. 2531 4785.1 Conti g55 m 201664 202824 - 96% SEQ ID No. 2606 4751.2 Conti g5δ m 202977 204695 - 97% SEQ ID No. 387 1516.3 Conti gδδ m 204692 205651 - 96% SEQ ID No. 386 1515.2 Conti gδδ P 205797 206564 - 96% SEQ ID No. 385 1513.4 Conti gδδ P 206522 207880 + 98% SEQ ID No. 2679 5022.2 Conti gδδ P 207980 209053 - 99% SEQ ID No. 1967 3988.3 Conti gδδ P 209183 210673 - 96% SEQ ID No. 1966 3986.1 Conti gδδ m 210767 211888 - 97% SEQ ID No. 1965 3985.1 Conti gδδ P 212031 212642 99% SEQ ID No. 3348 856.2 Conti gδδ P 212770 213318 - 97% SEQ ID No. 3349 857.2 Conti gδδ m 213377 215725 + 98% SEQ ID No. 2612 4908.3 Conti gδδ P 215762 221632 + 98% SEQ ID No. 1974 3996.2 Cont gδδ m 219910 220161 93% + SEQ ID No. 494 1671.3 Cont gδδ m 221880 223283 + 96% + SEQ ID No. 1973 3995.5 Cont gδδ m 223444 224850 - 98% SEQ ID No. 1972 3994.5 Cont gδδ m 224983 225843 - 98% SEQ ID No. 1971 3993.2 Conti gδδ P 226063 226629 - 87% SEQ ID No. 3372 887.4 Conti gδδ m 226955 229195 - 99% SEQ ID No. 3028 5711.2 Conti gδδ P 229465 230373 - 98% SEQ ID No. 2772 6193.2 Conti gδδ P 230456 232594 - 97% SEQ ID No. 1938 3938.2 Conti gδδ m 232591 233181 - 98% SEQ ID No. 1937 3937.2 Cont gδδ m 233166 234512 - 98% SEQ ID No. 854 2272.3 Conti gδδ m 234482 235996 - 99% SEQ ID No. 855 2274.2 Conti gδδ m 236004 237023 - 99% SEQ ID No. 1936 3936.2 Conti gδδ m 236965 238269 - 99%
SEQ ID No. 684 1972.3 Contigδδ m 238224 239414 99% SEQ ID No. 1186 278.3 Cont gδδ m 239267 241183 98% SEQ ID No. 2773 5194.1 Cont gδδ m 241056 241625 SEQ ID No. 1178 277.1 Contigδδ m 241772 242491 100% SEQ ID No. 1171 276.1 Contig55 m 242556 242792 100% SEQ ID No. 547 1754.3 Contigδδ m 243052 244590 98% SEQ ID No. 548 1756.3 Contigδδ m 244529 244864 98% SEQ ID No. 549 1756.4 Contigδδ m 244943 245797 98% SEQ ID No. 1114 2669.2 Contigδδ m 246338 247676 99% SEQ ID No. 1116 2670.1 Cont gδδ m 247577 248836 99% SEQ ID No. 1117 2671.1 Cont gδδ P 248990 249631 97% SEQ ID No. 1118 2674.1 Contigδδ P 249701 252391 99% SEQ ID No. 1119 2677.2 Cont gδδ P 252653 254060 100% SEQ ID No. 3344 847.4 Cont gδδ 171 254244 257436 99% SEQ ID No. 1121 2680.1 Contigδδ P 257935 258630 99% SEQ ID No. 518 1708.4 Cont gδδ P 259050 259427 99% SEQ ID No. 519 1709.1 Cont g55 m 259455 260177 98% SEQ ID No. 520 1711.2 Contig55 m 260134 260889 99% SEQ ID No. 864 2292.2 Contigδδ P 261111 262802 95% + SEQ ID No. 1122 2682.1 Contigδδ P 262935 264578 93% + SEQ ID No. 80 1036.4 Contigδδ m 264665 266002 98% SEQ ID No. 81 1037.1 Contigδδ m 265921 266478 100% SEQ ID No. 82 1039.2 Contigδδ P 266605 267834 95% SEQ ID No. 777 2144.3 Cont gδδ P 267780 268817 91% SEQ ID No. 1123 2683.3 Contigδδ P 268818 270377 96% SEQ ID No. 2774 5195.2 Cont gδδ P 270293 271729 98% SEQ ID No. 1124 2685.2 Cont gδδ P 271681 272466 98% SEQ ID No. 2492 4732.3 Contigδδ m 272544 272921 97% SEQ ID No. 540 1743.4 Contigδδ m 273331 273900 99% SEQ ID No. 539 1742.3 Contigδδ m 273878 274645 98% SEQ ID No. 2493 4733.2 Contigδδ m 274618 275526 98% SEQ ID No. 2665 4999.2 Contigδδ m 275562 276383 99% SEQ ID No. 2668 5000.3 Contigδδ P 276648 277070 98% SEQ ID No. 2669 5001.1 Contig55 P 277565 278038 99% SEQ ID No. 2820 5307.3 Cont g5δ m 278103 279803 97% SEQ ID No. 1970 3991.3 Contiigδδ m 280182 281885 95%
SEQ ID No. 3336 837.2 Contig55 P 281957 282727 100%
SEQ ID No. 3335 835.2 Contig55 P 282705 285299 99%
SEQ ID No. 1968 3989.1 Contig55 m 286321 285770 99%
SEQ ID No. 195 1210.2 Contig55 m 285761 287344 97%
SEQ ID No. 193 1209.2 Contig55 m 287341 288849 100%
SEQ ID No. 192 1208.1 Contig55 P 288191 288577 99%
SEQ ID No. 2821 5309.1 Contig55 P 288995 289852 100%
SEQ ID No. 1016 2514.3 Contigδδ P 289828 290118 100%
SEQ ID No. 3376 893.4 Contig55 m 290434 292959 99%
SEQ ID No. 2310 4472.2 Contigδδ m 293013 293819 99%
SEQ ID No. 2501 4744.3 Contigδδ m 293795 294718 98%
SEQ ID No. 378 1503.4 Contigδδ m 294663 295850 97%
SEQ ID No. 377 1501.3 Contigδδ m 295754 296599 98%
SEQ ID No. 928 2388.3 Contigδδ m 296578 297459 97%
SEQ ID No. 927 2387.3 Contigδδ P 297507 298577 99%
SEQ ID No. 154 1144.3 Contigδδ m 298751 300040 99%
SEQ ID No. 155 1145.3 Contigδδ m 300027 302186 97%
SEQ ID No. 2823 5311.1 Contigδδ P 302427 302744 100%
SEQ ID No. 2044 4094.2 Contigδδ P 302741 304198 97%
SEQ ID No. 2043 4093.1 Contigδδ P 304199 305635 98%
SEQ ID No. 893 2343.2 Contigδδ P 305819 306838 98%
SEQ ID No. 894 2344.4 Contigδδ P 306980 309283 97%
SEQ ID No. 2906 5478.2 Contigδδ P 309284 309784 88%
SEQ ID No. 136 1115.4 Contigδδ P 309795 310592 97%
SEQ ID No. 135 1113.2 Contigδδ P 310647 311045 96%
SEQ ID No. 2610 4902.1 Contigδδ P 310700 311026
SEQ ID No. 2905 5477.1 Contigδδ 171 311164 311460 100%
SEQ ID No. 2904 5476.1 Contigδδ 171 311846 312241 98%
SEQ ID No. 1698 3556.2 Contigδδ m 312219 312647 97%
SEQ ID No. 1699 3557.1 Contigδδ m 312599 313828 98%
SEQ ID No. 2168 427.3 Contigδδ m 313737 314516 99%
SEQ ID No. 2173 428.1 Contigδδ P 314611 315411 100%
SEQ ID No. 2180 429.2 Contigδδ m 315524 316543 98%
SEQ ID No. 748 2092.2 Contigδδ P 316557 319145 99%
SEQ ID No. 747 2091.1 Contigδδ m 316811 317566 97%
SEQ ID No. 1700 3558.1 Contigδδ m 317987 318523 98%
SEQ ID No. 1701 3559.2 Contig55 m 319237 320662 97% +
SEQ ID No. 635 1886.2 Contigδδ m 320663 321231 99% +
SEQ ID No. 634 1885.3 Contigδδ m 321603 322891 99%
SEQ ID No. 2432 4649.1 Contigδδ m 322872 323477 96% +
SEQ ID No. 2434 4650.3 Contigδδ m 323644 327368 98% +
SEQ ID No. 2164 4262.2 Contigδδ m 327356 327813 98%
SEQ ID No. 1039 2560.2 Contigδδ m 327807 329195 98%
SEQ ID No. 1040 2551.2 Contigδδ m 329159 329809 97%
SEQ ID No. 1041 2553.3 Contigδδ m 329939 330940 100%
SEQ ID No. 2162 4259.2 Contigδδ m 330931 331806 97%
SEQ ID No. 1784 3674.2 Contigδδ m 331754 332569
SEQ ID No. 1785 3675.1 Contigδδ m 332739 333086 98%
SEQ ID No. 1786 3676.2 Contigδδ m 333068 334462 99%
SEQ ID No. 1787 3679.3 Contigδδ m 334405 335460 99%
SEQ ID No. 2979 561.3 Contigδδ m 335736 337042 97%
SEQ ID No. 2983 562.1 Contigδδ m 337055 337672 99%
SEQ ID No. 3130 6007.1 Contigδδ P 337580 337705 100%
SEQ ID No. 2994 564.2 Contigδδ 111 337666 340092 98%
SEQ ID No. 1789 3680.1 Contigδδ P 339922 341073 98%
SEQ ID No. 1790 3681.1 Contigδδ P 341052 341744 97%
SEQ ID No. 1791 3682.1 Contigδδ P 341754 342179 95%
SEQ ID No. 1792 3683.2 Contigδδ P 342284 342532 100%
SEQ ID No. 2817 5296.1 Contigδδ m 342696 344141 99%
SEQ ID No. 972 2448.3 Contigδδ P 344088 344876 97%
SEQ ID No. 971 2447.3 Contigδδ P 345075 346076 98%
SEQ ID No. 970 2445.4 Contigδδ P 346267 349176 99%
SEQ ID No. 2539 4797.1 Contigδδ P 349121 350293 99%
SEQ ID No. 2538 4796.1 Contigδδ P 350604 351089 96%
SEQ ID No. 2537 4795.1 Contigδδ P 350815 351099 87%
SEQ ID No. 1983 4005.2 Contigδδ m 351176 352696 99%
SEQ ID No. 1984 4009.1 Contigδδ m 352663 353715 99%
SEQ ID No. 1986 4010.1 Contigδδ m 353830 354234 100%
SEQ ID No. 1987 4012.1 Contigδδ m 354228 355055 98%
SEQ ID No. 1988 4013.1 Contigδδ m 365006 355788 100%
SEQ ID No. 1989 4014.1 Contigδδ m 355789 356577 99%
SEQ ID No. 1990 4015.1 Contigδδ m 356602 357495 99%
SEQ ID No. 1991 4016.1 Contig55 m 367458 358642 - 98%
SEQ ID No. 1992 4017.4 Contigδδ m 358638 360716 - 99%
SEQ ID No. 2848 5361.2 Contigδδ m 360713 361897 - 99%
SEQ ID No. 2847 5360.2 Contigδδ P 361044 361610 99% +
SEQ ID No. 1066 2592.4 Contigδδ P 361844 362254 97% +
SEQ ID No. 1067 2593.3 Contigδδ m 361861 362634 - 99%
SEQ ID No. 2680 5026.1 Contigδδ m 362635 362964 + 98%
SEQ ID No. 1068 2696.4 Contigδδ m 362913 363662 + 97%
SEQ ID No. 2845 5359.2 Contigδδ m 363663 364091 - 98%
SEQ ID No. 2362 4526.3 Contigδδ m 364070 364414 - 96%
SEQ ID No. 434 1586.2 Contigδδ m 364415 365494 - 95%
SEQ ID No. 435 1586.2 Contigδδ m 365921 366190 + 100%
SEQ ID No. 436 1587.4 Contigδδ P 366186 367574 97%
SEQ ID No. 2171 4275.2 Contigδδ P 367445 368293 + 97%
SEQ ID No. 2170 4273.1 Contigδδ P 368190 369320 97%
SEQ ID No. 876 2313.2 Contigδδ m 369399 370625 - 96%
SEQ ID No. 875 2312.1 Contigδδ m 370829 371116 - 98%
SEQ ID No. 874 2311.4 Contigδδ m 371117 373702 - 98%
SEQ ID No. 627 1873.2 Contigδδ m 373717 374169 - 100%
SEQ ID No. 626 1871.2 Contig55 m 374162 375211 - 100%
SEQ ID No. 2428 4643.2 Contigδδ m 375511 376461 - 99%
SEQ ID No. 2427 4642.1 Contigδδ m 376740 377243 - 98%
SEQ ID No. 3101 5910.1 Contigδδ m 377311 377523 98%
SEQ ID No. 2426 4640.2 Contigδδ P 377831 380395 - 99%
SEQ ID No. 1994 4020.2 Contigδδ P 380396 382096 99%
SEQ ID No. 3332 830.4 Contigδδ P 382039 384633 95%
SEQ ID No. 3333 832.3 Contig55 P 384667 385662 - 94%
SEQ ID No. 3041 5747.1 Contigδδ P 385787 386191 + 96%
SEQ ID No. 678 1956.3 Contigδδ m 386532 388040 + 92%
SEQ ID No. 885 2328.2 Contigδδ P 388278 390869 - 99%
SEQ ID No. 1501 3269.1 Contigδδ P 390859 394104 - 98%
SEQ. ID No. 960 2430.2 Contigδδ P 394309 395385 - 99%
SEQ ID No. 1500 3267.1 Contigδδ P 395435 396055 99%
SEQ ID No. 1499 3265.1 Contigδδ m 396103 396999 - 98%
SEQ ID No. 1498 3263.2 Contigδδ P 397202 398425 - 98%
SEQ ID No. 1497 3260.2 Contigδδ m 398427 399434 + 99%
SEQ ID No. 467 1633.2 Contig55 m 399379 SEQ ID No. 466 400365 1632.3 Contig55 99% m 400360 402146 SEQ ID No. 2811 5278.3 Contig55 99% m 402203 403054 SEQ ID No. 2810 5277.3 Contig55 99% m 403006 404235 SEQ ID No. 1371 3070.2 Contig55 100% m 404293 405312 SEQ ID No. 1369 3069.1 Contigδδ 100% m 405340 406062 SEQ ID No. 1368 3068.1 Contigδδ 99% P 406331 407521 SEQ ID No. 1031 2538.2 Contigδδ 100% P 407509 408717 SEQ ID No. 1367 3067.1 Contigδδ 100% P 408846 409169 SEQ ID No. 3015 568.2 Contigδδ 100% P 409176 410786 SEQ ID No. 3020 569.2 Contigδδ 99% P 410743 SEQ ID No. 3027 411600 571.2 Contigδδ 99% P 411503 SEQ ID No. 699 413557 2007.1 Contigδδ 97% P 413535 SEQ ID No. 145 414476 1127.3 Contigδδ 100% P 414552 416573 SEQ ID No. 1366 3065.1 Contigδδ 98% m 416733 417197 SEQ ID No. 1365 3063.2 Contigδδ 96% P 417443 SEQ ID No. 1364 418627 3061.2 Contigδδ 99% P 418602 419252 SEQ ID No. 1363 3060.1 Contigδδ 99% m 418655 419038 SEQ ID No. 1361 3059.2 Contigδδ 96% P 419253 SEQ ID No. 95 420044 1058.3 Contigδδ 99% P 420079 421542 SEQ ID No. 94 1056.1 Contigδδ 98% + P 421787 422035 SEQ ID No. 93 1055.1 Contigδδ 100% m + 422032 422802 SEQ ID No. 92 1053.2 Contigδδ 97% m 422766 SEQ ID No. 521 423317 1713.2 Contigδδ 99% m 423302 SEQ ID No. 522 425392 1714.4 Contigδδ 97% m 425368 SEQ ID No. 579 426290 1797.4 Contigδδ 100% m 426498 SEQ ID No. 582 427307 1800.4 Contigδδ 100% m 427491 SEQ ID No. 2277 429701 4419.2 Contigδδ 98% P 429883 430851 SEQ ID No. 2899 5469.1 Contigδδ 99% P 430842 431012 SEQ ID No. 2900 5470.1 Contig55 P 430963 SEQ ID No. 228 431283 1260.3 Contig55 100% P 431468 SEQ ID No. 226 432283 1258.1 100% Contig55 m 432392 433441 SEQ ID No. 225 1257.5 Contig55 97% m 433361 434646 SEQ ID No. 3129 6004.1 99% Contig55 m 434692 434850 SEQ ID No. 2901 + 5471.2 Contig55 m 434805 SEQ ID No. 3099 435017 + 5907.1 Contigδδ m 434953 435111 +
SEQ ID No. 3040 5743.1 Contig55 m 436126 435313 +
SEQ ID No. 2895 5468.1 Contig56 m 1 294 +
SEQ ID No. 1829 3744.2 Contig56 P 629 1834 + +
SEQ ID No. 1828 3743.1 Contig56 m 1999 2334 + - 96% +
SEQ ID No. 1827 3742.1 Contig56 m 2352 2624 - 92% +
SEQ ID No. 1826 3741.1 Contig56 m 2635 3855 - 98% +
SEQ ID No. 1825 3740.3 Contig56 m 4089 4625 - 100%
SEQ ID No. 1823 3739.3 Contig56 P 4848 5291 - 99%
SEQ ID No. 1822 3737.1 Contig56 P 5344 6447 + - 94%
SEQ ID No. 113 1082.3 Contig56 m 6468 8033 +
SEQ ID No. 367 1469.4 Contig56 m 8879 12292 +
SEQ ID No. 2197 4312.2 Contig56 m 12373 13725 - 98%
SEQ ID No. 2196 4311.1 Contig56 P 13958 14506 - 96%
SEQ ID No. 2195 4310.1 Contig56 m 14610 15161 - 97% +
SEQ ID No. 2193 4309.2 Contig56 m 15077 16219 . 97% +
SEQ ID No. 2010 4047.3 Contig56 m 16291 17385 + - 99%
SEQ ID No. 2011 4048.3 Contigδθ P 17493 19334 - 100%
SEQ ID No. 2013 4050.1 Contig56 P 19403 20197 + - 98%
SEQ ID No. 2014 4051.2 Contigδ6 P 20206 20601 + - 98%
SEQ ID No. 2015 4064.2 Contig56 P 20585 21265 - 99%
SEQ ID No. 2016 4066.1 Contigδδ m 21293 23185 +
SEQ ID No. 2017 4066.2 Contig56 m 23266 24495 - 99%
SEQ ID No. 2018 4068.2 Contig56 m 24583 24942 +
SEQ ID No. 472 1641.5 Contig56 P 25345 26694 - 98%
SEQ ID No. 473 1642.3 Contig56 P 26679 27341 - 100%
SEQ ID No. 2465 4699.3 Contig56 P 27452 28756 - 99%
SEQ ID No. 2340 4510.1 Contig56 P 28978 31440 - 100%
SEQ ID No. 2341 4511.1 Contigδθ P 31527 31847 - 98%
SEQ ID No. 2342 4512.4 Contig56 P 32084 33979 - 98%
SEQ ID No. 572 1787.4 Contigδθ P 34519 37686 + +
SEQ ID No. 1678 3530.1 Contigδθ m 37969 38598 + +
SEQ ID No. 1679 3532.1 Contig56 P 38652 41942 + +
SEQ ID No. 368 1487.3 Contig56 m 42394 43230 - 99%
SEQ ID No. 1680 3533.1 Contigδθ m 43296 44021 + . 99%
SEQ ID No. 1681 3534.2 Contig56 m 43987 45666 + . 99%
SEQ ID No. 1682 3536.5 Contig56 P 45982 46656 - 95%
SEQ ID No. 1683 3536.2 Contig56 m 46690 47052 98%
SEQ ID No. 2732 5098.2 Contigδδ m 47027 47311 100%
SEQ ID No. 2731 5097.1 Contigδ6 m 47312 48241 + 99%
SEQ ID No. 3128 6002.1 Contigδ6 P 48661 49746
SEQ ID No. 3125 5999.1 Contιg56 P 49760 50065 100%
SEQ ID No. 2480 4715.3 Contigδθ P 50326 50817 95%
SEQ ID No. 2479 4714.1 Contigδδ m 50839 51837 96%
SEQ ID No. 2478 4713.1 Contigδ6 P 51969 52322 93%
SEQ ID No. 2935 5532.2 Contig56 m 52635 53546 97%
SEQ ID No. 2936 5533.2 Contigδδ m 53673 53912
SEQ ID No. 499 1682.4 Contig56 P 54221 55105 94%
SEQ ID No. 500 1683.2 Contigδδ m 55230 55467 +
SEQ ID No. 501 1684.3 Contig56 P 65649 56050 96% +
SEQ ID No. 2249 4378.2 Contigδδ P 56210 56911 95%
SEQ ID No. 2250 4379.1 Contig56 P 56884 57786 94%
SEQ ID No. 2251 4381.2 Contig56 P 57812 59134 + 99%
SEQ ID No. 2252 4383.2 Contig56 P 59183 60082 + 97%
SEQ ID No. 997 2483.2 Contigδθ m 60143 60406 93%
SEQ ID No. 996 2482.1 Contig56 m 60536 61048 96%
SEQ ID No. 995 2481.4 Contigδθ m 61231 62142 97%
SEQ ID No. 2401 4600.2 Contig56 m 62433 63200 97%
SEQ ID No. 2402 4601.1 Contigδδ m 63160 63900 97%
SEQ ID No. 2403 4602.3 Contig56 P 63958 64875 98%
SEQ ID No. 3002 5652.2 Contig56 P 64957 65739 97% +
SEQ ID No. 2009 4045.2 Contigδβ P 65724 66335 +
SEQ ID No. 2008 4043.1 Contig56 m 66641 67078 98%
SEQ ID No. 2007 4042.1 Contig56 P 67106 67858 96%
SEQ ID No. 2006 4041.1 Contig56 P 68024 69148 + 96%
SEQ ID No. 2005 4040.1 Contig56 P 69187 69486 +
SEQ ID No. 2004 4039.2 Contig56 m 69539 70852 +
SEQ ID No. 2003 4037.2 Contig56 m 70837 71116 +
SEQ ID No. 2002 4036.2 Contig56 P 71116 71400 +
SEQ ID No. 2001 4035.1 Contig56 P 71831 72199 80%
SEQ ID No. 2326 4490.2 Contig56 m 72497 73351 +
SEQ ID No. 2327 4491.3 Contig56 m 73671 73943 + +
SEQ ID No. 2328 4492.3 Contig56 P 74457 75404 + +
SEQ ID No. 2329 4493.1 Contig56 P 75398 75607 - +
SEQ ID No. 967 2441.2 Contig56 P 75770 77053 - +
SEQ ID No. 965 2439.2 Contig56 P 77137 77460 - +
SEQ ID No. 964 2438.2 Contig56 P 77301 77741 - +
SEQ ID No. 2330 4496.2 Contigδδ P 77914 78756 - + +
SEQ ID No. 2975 5605.2 Contigδδ P 79041 79460 - + +
SEQ ID No. 425 1570.4 Contig56 P 79560 82762 + +
SEQ ID No. 2343 4516.3 Contigδθ P 82740 83900 + +
SEQ ID No. 306 1387.2 Contigδδ P 83888 85336 - +
SEQ ID No. 307 1388.2 Contigδθ m 85697 85953 + +
SEQ ID No. 853 2271.1 Contig56 m 85954 86205 + +
SEQ ID No. 852 2270.2 Contig56 m 86198 86578 +
SEQ ID No. 2860 5387.1 Contιg56 P 86587 87405 - +
SEQ ID No. 2861 5388.1 Contιg56 P 87345 87626 - +
SEQ ID No. 1941 3953.1 Contig56 m 87959 89164 - 99%
SEQ ID No. 506 1692.2 Contig56 P 89181 91175 - 98%
SEQ ID No. 1940 3950.2 Contig56 P 91527 93143 + 94%
SEQ ID No. 2681 5028.2 Contig56 m 93235 94818 + 97%
SEQ ID No. 2863 6390.1 Contig56 m 94884 95249 97%
SEQ ID No. 1995 4021.2 Contig56 P 95342 96935 - 98%
SEQ ID No. 1996 4023.2 Contig56 P 95838 97349 - 99%
SEQ ID No. 1997 4025.1 Contig56 P 97336 98541 - 98%
SEQ ID No. 1998 4026.2 Contig56 P 98511 99701 + 97%
SEQ ID No. 273 1334.3 Contigδ6 P 99838 100644 - 86% +
SEQ ID No. 274 1335.2 Contig56 m 100634 101995 90% +
SEQ ID No. 1999 4030.1 Contig56 P 102167 102703 - 79% +
SEQ ID No. 2000 4032.3 Contig56 m 102700 103668 - 99%
SEQ ID No. 2864 5391.2 Contig56 m 103725 105176 - 99%
SEQ ID No. 3124 6996.1 Contigδ6 m 105163 105690 + 98%
SEQ ID No. 2822 631.4 Contigδδ P 105919 107865 - 98%
SEQ ID No. 2819 630.3 Contig56 m 107831 108436 - 96%
SEQ ID No. 285 1356.2 Contigδ6 m 108648 109430 88%
SEQ ID No. 286 1357.2 Contig56 m 109467 109916 - 98%
SEQ ID No. 2635 4948.4 Contigδθ P 110012 113998 - 96%
SEQ ID No. 869 2299.3 Contigδβ P 114216 114683 - 95%
SEQ ID No. 870 2301.2 Contigδδ P 115076 117259 - 99%
SEQ ID No. 1354 3049.1 Contig56 P 117222 118001 97%
SEQ ID No. 1353 3047.2 Contig56 P 117973 118644 99%
SEQ ID No. 1352 3046.3 Contig56 m 118221 118781 97%
SEQ ID No. 1351 3045.3 Contig56 P 118645 120087 99%
SEQ ID No. 1350 3042.1 Contig56 m 120097 120294 +
SEQ ID No. 1349 3041.1 Contigδθ P 120462 121664 95% +
SEQ ID No. 2967 559.2 Contig56 P 121830 122159 97%
SEQ ID No. 2961 568.2 Contigδβ P 122262 122726 98%
SEQ ID No. 2959 557.2 Contigδδ m 122756 125467 98%
SEQ ID No. 606 1844.3 Contigδθ m 125449 126537 100%
SEQ ID No. 607 1845.3 Contigδθ P 126753 127550 100%
SEQ ID No. 608 1846.3 Contigδβ m 127547 128779 99%
SEQ ID No. 71 1022.2 Contig56 m 128901 129779 97%
SEQ ID No. 72 1023.2 Contig56 m 129880 130467 95%
SEQ ID No. 73 1024.2 Contigδ6 m 130563 131165 99%
SEQ ID No. 348 1451.4 Contig56 m 131482 132765 98%
SEQ ID No. 617 1859.2 Contig56 m 132780 133634 100%
SEQ ID No. 616 1858.1 Contigδδ m 133685 133957 100% +
SEQ ID No. 615 1866.3 Contigδδ P 134558 136363 99% +
SEQ ID No. 2383 4574.2 Contigδβ P 136288 137400 9δ%
SEQ ID No. 164 1160.2 Contig56 m 137686 138921
SEQ ID No. 162 1159.5 Contigδδ m 139130 140416 99%
SEQ ID No. 498 1681.5 Contig56 P 141000 143876 98%
SEQ ID No. 994 248.2 Contigδδ m 143990 145393 100%
SEQ ID No. 987 247.1 Contig56 m 145394 146179 99%
SEQ ID No. 973 245.1 Contigδδ m 146315 147349 100%
SEQ ID No. 966 244.1 Contig56 m 147447 148508 99%
SEQ ID No. 959 243.2 Contig56 m 148617 149470 99%
SEQ ID No. 702 2017.1 Contig56 P 149611 150183 98%
SEQ ID No. 701 2016.2 Contig56 P 150223 160675 100%
SEQ ID No. 1343 3028.1 Contig56 P 150666 150934 100%
SEQ ID No. 1342 3027.1 Contigδβ P 150941 151432 99%
SEQ ID No. 1837 376.3 Contig56 P 151338 152423 98%
SEQ ID No. 1843 377.1 Contig56 m 152601 153340 99%
SEQ ID No. 1865 381.6 Contigδδ m 153410 156961 89% +
SEQ ID No. 3090 5899.1 Contig56 P 157044 157373 100% +
SEQ ID No. 595 1823.5 Contig56 m 157377 158678 SEQ ID No. 2223 4346.2 99% Contig56 m 158924 159877 SEQ ID No. 2224 4347.2 99% Contig56 P 159936 160262 SEQ ID No. 2225 4349.2 98% Contig56 P 160489 161322 SEQ ID No. 2226 4350.2 100% Contig56 P 161323 162489 SEQ ID No. 2227 4351.2 98% Contig56 P 162510 163424 SEQ ID No. 2228 4352.3 98% Contig56 P 163425 164372 SEQ ID No. 3121 5986.2 98% Contig56 m 164438 164929 SEQ ID No. 2460 4691.2 98% Contig56 P 165075 166451 SEQ ID No. 2459 4690.2 99% Contig56 m 166544 168484 SEQ ID No. 2457 4688.2 98% Contig56 m 168556 169680 SEQ ID No. 2012 405.3 98% Contig56 m 169767 171719 SEQ ID No. 2019 406.1 98% Contig56 m 171982 172845 SEQ ID No. 2027 407.4 98% Contig56 m 172815 174182 SEQ ID No. 2874 5420.1 99% Contig56 P 174276 174809 SEQ ID No. 2875 6421.1 Contigδβ P 174790 175281 SEQ ID No. 2876 6422.1 97% Contig56 + P 175516 175929 SEQ ID No. 2877 5423.1 96% Contig56 + P 176093 176353 SEQ ID No. 825 2225.2 Contig56 P 176301 177230 SEQ ID No. 146 1128.3 97% Contigδβ m 177274 178308 SEQ ID No. 147 1132.2 99% Contigδδ P 178079 178876 SEQ ID No. 148 1133.3 97% Contig56 m 178905 180047 SEQ ID No. 149 1134.3 97% Contig56 m 180035 180313 SEQ ID No. 1861 3802.2 100% Contigδδ P 180508 181905 SEQ ID No. 1862 3803.1 100% Contig56 P 181817 182977 SEQ ID No. 1863 3804.2 100% Contigδ6 P 182964 183833 SEQ ID No. 1864 3807.2 98% Contigδδ P 183917 184408 SEQ ID No. 321 1407.2 99% Contigδδ P 184433 185311 SEQ ID No. 320 1406.2 98% Contigδδ m 185316 186611 SEQ ID No. 319 1405.2 96% Contigδβ P 186721 187620 SEQ ID No. 2878 5424.2 Contig56 98% m 187587 188366 SEQ ID No. 2749 5134.4 98% Contig56 m 188552 189529 SEQ ID No. 2748 5133.3 99% Contig56 P 189601 190266 SEQ ID No. 2747 5132.3 99% Contig56 m + 190423 190665 SEQ ID No. 1372 3072.2 Contig56 m + 190992 192803 SEQ ID No. 3203 640.2 99% Contig56 m 193129 194091 97%
SEQ ID No. 3204 641.3 Contig56 m 194043 196091 - 99%
SEQ ID No. 467 1618.3 Contig56 P 196354 196860 + - 98%
SEQ ID No. 466 1617.3 Contig56 m 196904 197545 - 98%
SEQ ID No. 785 2152.2 Contig56 m 197442 198038 - 97%
SEQ ID No. 1373 3074.1 Contig56 P 198106 199032 - 96%
SEQ ID No. 631 1880.3 Contig56 m 199130 200137 - 100%
SEQ ID No. 979 2459.1 Contig56 m 200112 200678 - 99%
SEQ ID No. 978 2458.2 Contig56 P 200733 201881 - 98%
SEQ ID No. 791 2162.2 Contig56 m 201913 203340 - 98%
SEQ ID No. 792 2164.2 Contig56 P 203649 204509 - 98%
SEQ ID No. 1374 3075.2 Contig56 m 204200 204562 - 95%
SEQ ID No. 646 1902.5 Contig56 m 204808 205812 - 98%
SEQ ID No. 2849 5369.2 Contig56 P 206011 206253 - 100%
SEQ ID No. 2851 5370.1 Contig56 P 206447 206899 - 98%
SEQ ID No. 860 2282.4 Contig56 P 206833 208641 - 99%
SEQ ID No. 169 1169.3 Contig56 P 208701 210593 - 99%
SEQ ID No. 168 1168.2 Contig56 m 209799 210206 - 94%
SEQ ID No. 3077 5871.3 Contig56 m 210934 212142 +
SEQ ID No. 1824 374.2 Contig56 m 212175 213962 - 90%
SEQ ID No. 1809 371.1 Contig56 P 214213 214563 + +
SEQ ID No. 565 1779.3 Contig56 111 214773 214958 + +
SEQ ID No. 567 1780.3 Contig56 P 215260 215511 - 100%
SEQ ID No. 568 1781.4 Contig56 P 215742 217157 - 98%
SEQ ID No. 441 1592.3 Contig56 P 217296 218225 + - 99%
SEQ ID No. 2177 4285.1 Contig56 m 218497 218913 +
SEQ ID No. 2176 4284.2 Contig56 m 218914 219987 +
SEQ ID No. 2175 4282.2 Contig56 m 220040 220858 +
SEQ ID No. 2174 4281.2 Contig56 P 220863 221168 + +
SEQ ID No. 2172 4276.2 Contig56 m 221531 223492 4-
SEQ ID No. 3078 5872.1 Contig56 m 223532 223738 + +
SEQ ID No. 2357 4532.2 Contig56 P 223941 225431 + +
SEQ ID No. 2356 4631.2 Contig56 m 225684 226010 +
SEQ ID No. 2356 4630.1 Contig56 m 226119 226424 +
SEQ ID No. 2353 4628.2 Contig56 m 226438 227223 +
SEQ ID No. 2921 5604.4 Contig56 m 227405 228223 +
SEQ ID No. 906 2357.4 Contig56 P 228590 230416 +
SEQ ID No. 3355 864.1 Contigδ6 m 230660 231064 77%
SEQ ID No. 3354 863.1 Contig56 m 231159 231653 4-
SEQ ID No. 3353 862.1 Contigδδ P 231881 232144 4-
SEQ ID No. 3352 860.2 Contig56 P 232289 234031 +
SEQ ID No. 1598 3411.2 Contig56 P 234499 235608 - +
SEQ ID No. 1599 3413.2 Contigδδ P 235844 236812 4-
SEQ ID No. 1600 3414.3 Contig56 m 237027 238826 4-
SEQ ID No. 652 1910.6 Contig56 m 239000 243505 +
SEQ ID No. 653 1911.4 Contig56 m 243805 244335 86%
SEQ ID No. 403 1536.3 Contig56 m 244316 246199 + 95%
SEQ ID No. 450 1603.3 Contig56 P 246579 248123 4-
SEQ ID No. 1902 3878.1 Contig56 m 248247 251330 4-
SEQ ID No. 1901 3876.1 Contigδδ P 251633 252130 92%
SEQ ID No. 1900 3874.1 Contig56 m 252199 252504 95% 4-
SEQ ID No. 1899 3872.1 Contig56 P 262637 252915 4- 4-
SEQ ID No. 1898 3871.1 Contig56 P 252926 253329 4- 4-
SEQ ID No. 2870 5406.1 Contig56 m 253581 255119 97%
SEQ ID No. 259 1307.4 Contig56 P 255297 256598 93%
SEQ ID No. 258 1306.5 Contigδθ m 256705 259905 + 96%
SEQ ID No. 2869 5405.2 Contigδδ m 259906 260877 + 97%
SEQ ID No. 59 1001.3 Contig56 m 260871 262217 + 95%
SEQ ID No. 60 1002.2 Contig56 m 262543 264180 80%
SEQ ID No. 2500 4742.3 Contig56 P 264520 265437 97%
SEQ ID No. 2510 4757.2 Contigδβ m 265449 266069 92%
SEQ ID No. 2511 4758.2 Contigδ6 m 266170 266652 95%
SEQ ID No. 2512 4759.2 Contig56 P 266570 267409 98%
SEQ ID No. 2513 4760.2 Contig56 m 267786 268739 95%
SEQ ID No. 2709 5065.2 Contig56 P 268942 269424 96%
SEQ ID No. 2710 5066.2 Contig56 m 269508 270020 97%
SEQ ID No. 474 1643.4 Contig56 m 269917 272451 96%
SEQ ID No. 2336 4500.1 Contig56 P 272747 273112
SEQ ID No. 2332 4498.1 Contig56 P 273538 275025 97% 4-
SEQ ID No. 2331 4497.4 Contigδθ P 275092 276381 98% 4-
SEQ ID No. 667 1937.4 Contig56 P 276466 276828 98% 4-
SEQ ID No. 668 1939.5 Contig56 P 277056 277634 95% 4-
SEQ ID No. 2942 5642.3 Contig56 P 277693 278568 98%
SEQ ID No. 1918 3904.3 35 Contig56 SEQ ID No. 2267 P 278644 440.4 280497 Contig56 97% SEQ ID No. P 280671 , 1919 285386 3908.3 Contig56 95% SEQ ID No. P 1813 285679 286721 3714.3 Contig56 97% 286765 4- SEQ ID No. P 3324 817.7 289398 Contig56 97% 4- SEQ ID No. 664 P 289615 1933.4 295440 Contig56 93% 295715 4- SEQ ID No. P 1328 3003.1 300448 Contig56 87% SEQ ID No. P 1327 300856 3002.1 301554 4- Contigδβ m 99% SEQ ID No. 963 301583 302824 2436.2 Contig56 m 96% SEQ ID No. 1326 302990 3001.1 304480 Contigδδ 96% SEQ ID No. P 1325 304644 3000.1 304823 4- Contig56 SEQ ID No. 1321 P 304917 307223 4- 2999.3 Contigδδ 98% SEQ ID No. P 307366 1320 2998.1 308247 Contigδθ m 92% SEQ ID No. 308384 1319 2996.1 308848 4- Contig56 97% SEQ ID No. P 309173 1318 309691 4- 2995.1 Contig56 98% SEQ ID No. 656 P 309823 311085 4- 1918.3 Contig56 m 97% SEQ ID No. 1026 311244 311996 + 2528.3 Contig56 m 97% SEQ ID No. 1317 311932 2994.2 312699 Contig56 88% SEQ ID No. P 312863 1316 313255 2993.2 Contig56 SEQ ID No. P 1315 313301 95% 2992.2 314026 Contigδβ 95% SEQ ID No. P 1314 314243 2991.1 315269 Contigδθ m 92% SEQ ID No. 315382 1313 316488 2990.1 Contig56 96% SEQ ID No. P 316851 1312 2988.1 317330 4- Contig56 SEQ ID No. 1311 P 317318 2987.1 317605 4- Contigδβ m SEQ ID No. 317642 1310 318247 4- 2986.1 4- Contig56 m SEQ ID No. 1309 318269 2985.1 318838 4- + Contig56 m SEQ ID No. 1308 318799 2983.1 319164 4- 4- Contigδ6 SEQ ID No. P 1307 319420 2982.1 319680 4- 4- Contigδθ m SEQ ID No. 1306 319677 2978.1 321527 4- Contigδβ m SEQ ID No. 321511 1305 2976.1 322419 4- Contig56 m SEQ ID No. 322361 1304 322648 2975.2 4- Contig56 m SEQ ID No. 1303 322563 322895 + 2973.2 4- Contig56 m SEQ ID No. 1302 322880 323161 4- 2972.3 4- Contigδδ m SEQ ID No. 323201 3013 324955 4- 5675.2 4- Contig56 m SEQ ID No. 3014 324956 5677.2 326062 4- Contig56 m SEQ ID No. 2796 326486 5242.3 327133 4- Contigδβ m 327199 327939 4-
SEQ ID No. 2797 5243.2 Contig56 m 327951 328175
SEQ ID No. 344 1446.3 Contig56 P 328546 329315 91%
SEQ ID No. 343 1444.3 Contig56 m 329451 331304 95%
SEQ ID No. 625 1870.3 Contig56 P 331500 332612 97%
SEQ ID No. 3195 630.4 Contig56 P 332655 335864 98%
SEQ ID No. 1460 3205.3 Contig56 P 335981 340345 97%
SEQ ID No. 1461 3207.5 Contig56 P 340451 342949 97%
SEQ ID No. 2563 4830.2 Contig56 P 343121 344029 99%
SEQ ID No. 2564 4833.2 Contig56 P 344290 345072 98%
SEQ ID No. 2798 5247.1 Contig56 P 345073 346131 97%
SEQ ID No. 190 1205.5 Contig56 P 346389 357734 97%
SEQ ID No. 191 1207.2 Contig56 m 357737 358141 99%
SEQ ID No. 1280 2949.2 Contig56 m 358365 359324 99%
SEQ ID No. 1282 2951.2 Contigδβ P 359726 361783 4-
SEQ ID No. 1283 2962.1 Contig56 m 361896 363770 85% 4-
SEQ ID No. 1284 2963.2 Contigδ6 P 364225 364773 96%
SEQ ID No. 813 2204.2 Contigδ6 P 364871 366475 96%
SEQ ID No. 814 2207.2 Contig56 P 366476 367105 96%
SEQ ID No. 815 2208.1 Contigδβ m 367223 367516 77%
SEQ ID No. 816 2209.4 Contig56 m 367547 368536 96%
SEQ ID No. 1043 2557.4 Contig56 m 368573 369052 96%
SEQ ID No. 1042 2555.5 Contig56 P 369375 370547 4-
SEQ ID No. 2408 4613.3 Contig56 P 370702 372222 95% 4-
SEQ ID No. 2407 4609.2 Contig56 m 372227 372832 87% 4-
SEQ ID No. 2406 4608.1 Contig56 m 373026 373588 91%
SEQ ID No. 2405 4607.2 Contig56 m 373694 374998 99%
SEQ ID No. 2981 5611.1 Contigδδ m 374824 375147
SEQ ID No. 2980 5610.1 Contigδβ m 375176 375633 92%
SEQ ID No. 2978 5609.1 Contigδβ m 375588 375899
SEQ ID No. 2723 5084.4 Contigδβ m 375956 377518 96%
SEQ ID No. 2977 5607.2 Contig56 m 377512 378960 99%
SEQ ID No. 2976 5606.2 Contig56 P 379119 379787 91% 4-
SEQ ID No. 262 1317.3 Contig56 P 379963 381123 98% 4-
SEQ ID No. 263 1318.4 Contig56 m 381171 381926 95%
SEQ ID No. 3007 5662.1 Contig56 m 381808 382452
SEQ ID No. 2366 4547.4 Contigδδ m 382744 383787 99%
SEQ ID No. 2104 4171.4 Contig56 P 384054 384602 4- 98%
SEQ ID No. 2106 4172.2 Contig56 m 384751 385848 - 97%
SEQ ID No. 2106 4173.2 Contig56 m 385948 386391 - 98%
SEQ ID No. 1025 2526.3 Contigδδ P 386531 387826 - 97%
SEQ ID No. 2107 4174.1 Contig56 P 388061 388519 4- 100%
SEQ ID No. 2108 4175.1 Contig56 m 388616 389545 - 98%
SEQ ID No. 2109 4177.2 Contig56 m 389619 391223 - 97%
SEQ ID No. 2952 5557.2 Contig56 m 391235 393079 - 95%
SEQ ID No. 2424 4636.4 Contig56 P 393601 398376 -
SEQ ID No. 647 1903.4 Contig56 m 398612 399586 - 97%
SEQ ID No. 3051 5796.3 Contig56 P 399743 400387 - 96%
SEQ ID No. 1529 3308.4 Contig56 P 400486 400734 - 4- 4-
SEQ ID No. 1528 3307.4 Contig56 P 400664 400867 - 4- 4-
SEQ ID No. 2913 549.5 Contig56 P 400978 402165 99% 4-
SEQ ID No. 2908 548.3 Contig56 m 402228 403745 4- 97% 4-
SEQ ID No. 1527 3306.4 Contig56 m 403870 404619 4- 100%
SEQ ID No. 1526 3304.2 Contig56 m 404967 406145 + 99%
SEQ ID No. 1525 3303.2 Contig56 m 406350 407627 - 95%
SEQ ID No. 115 1088.3 Contig56 m 407795 409327 - 96%
SEQ ID No. 114 1087.2 Contig56 m 409331 411073 - 96%
SEQ ID No. 1524 3301.2 Contig56 m 411031 412473 - 98%
SEQ ID No. 1521 3299.3 Contig56 m 412674 413180 - 4-
SEQ ID No. 3110 5950.1 Contig56 m 413084 413554 4-
SEQ ID No. 1520 3297.3 Contig56 P 414188 415258 - 98%
SEQ ID No. 325 1412.5 Contig56 P 415252 417015 - 98%
SEQ ID No. 326 1413.2 Contig56 P 416981 418399 - 95%
SEQ ID No. 327 1414.1 Contig56 P 418371 418622 -
SEQ ID No. 328 1415.4 Contig56 P 418607 419371 - 97% 4-
SEQ ID No. 1250 2900.2 Contig56 P 419278 420378 - 97% +
SEQ ID No. 1247 2898.4 Contig56 m 420698 420934 -
SEQ ID No. 604 1840.3 Contig56 m 420935 422323 - 99%
SEQ ID No. 602 1839.2 Contig56 P 422240 423514 - 97%
SEQ ID No. 1246 2893.2 Contig56 P 423595 426009 - 97%
SEQ ID No. 1245 2891.2 Contig56 m 426200 428203 - 98%
SEQ ID No. 1244 2889.2 Contig56 m 428194 428649 + 99%
SEQ ID No. 1034 2544.3 Contig56 P 428750 432649 - 96%
SEQ ID No. 860 2268.2 Contig56 P 432873 434093 99%
SEQ ID No. 849 2266.2 Contig56 P 434150 435448 99%
SEQ ID No. 848 2264.2 Contig56 P 435367 436682 96%
SEQ ID No. 217 1242.2 Contig56 P 436686 437339 99%
SEQ ID No. 215 1239.2 Contig56 P 437446 437651 4-
SEQ ID No. 214 1238.2 Contig56 P 437637 440546 97% +
SEQ ID No. 2769 5188.2 Contig56 m 440810 442120 98%
SEQ ID No. 2770 5189.2 Contigδ6 m 442423 442962 100%
SEQ ID No. 1478 3235.3 Contig56 P 443439 444020 95%
SEQ ID No. 3109 5947.1 Contig56 P 443846 444142
SEQ ID No. 1479 3236.3 Contig56 m 444206 446461 98%
SEQ ID No. 3401 926.4 Contig56 m 446531 447229 97%
SEQ ID No. 3402 928.1 Contig56 m 447359 448597 97%
SEQ ID No. 3403 929.2 Contigδβ m 448649 450079 97%
SEQ ID No. 573 1788.3 Contig56 m 450084 461466 98%
SEQ ID No. 57δ 1792.2 Contig56 m 451509 452316 99%
SEQ ID No. 1480 3238.1 Contig56 m 452294 453229 95%
SEQ ID No. 137 1116.2 Contig56 m 463233 453562 98%
SEQ ID No. 138 1118.2 Contig56 m 453914 455338 97%
SEQ ID No. 139 1119.2 Contigδ6 m 455395 456072 92%
SEQ ID No. 1481 3239.1 Contig56 P 456141 456452 100%
SEQ ID No. 1483 3240.3 Contigδδ m 456543 456878 100%
SEQ ID No. 2118 4192.2 Contig56 P 456954 458858 97%
SEQ ID No. 2117 4191.1 Contig56 P 458774 460276 98%
SEQ ID No. 2116 4188.1 Contig56 P 460258 461391 95%
SEQ ID No. 2114 4186.1 Contig56 P 461427 462320 98%
SEQ ID No. 2113 4184.1 Contig56 P 462305 462637 99%
SEQ ID No. 477 1647.2 Contig56 m 462634 463245 98%
SEQ ID No. 476 1645.1 Contig56 P 463276 463866 98%
SEQ ID No. 475 1644.6 Contig56 m 463899 465158 95%
SEQ ID No. 1061 2583.3 Contig56 P 465296 465898 94%
SEQ ID No. 1060 2582.3 Contig56 P 465798 468422 97%
SEQ ID No. 2549 4809.2 Contig56 P 468423 469292 98%
SEQ ID No. 2551 4810.1 Contig56 P 469441 470094 98%
SEQ ID No. 2552 4811.2 Contig56 m 470268 470777 97%
SEQ ID No. 568 1768.3 Contigδδ m 470901 472232 99%
SEQ ID No. 557 1767.2 S9 Contig56 m SEQ ID No. 472211 2562 472930 4826.2 Contig56 m 97% SEQ ID No. 2031 472935 473642 H 4073.1 Contig56 99% SEQ ID No. P 2030 474278 4072.1 475393 Contigδδ 99% SEQ ID No. P 475378 3453 476064 997.2 Contigδβ 98% SEQ ID No. P 3452 476065 477177 996.2 Contig56 99% SEQ ID No. 3451 P 477200 995.2 478231 Contigδδ 98% SEQ ID No 2029 P 478195 4071.1 478866 Contigδ6 111 91 % SEQ ID No. 2028 478863 479441 4070.3 Contig56 95% SEQ ID No. 2453 P 479536 480974 4678.3 Contig56 m 99% SEQ ID No. 2465 480971 481381 4680.2 Contig56 m 82% SEQ ID No. 2456 481393 482142 4683.3 Contig56 m 98% SEQ ID No. 482069 2914 5492.2 483433 + Contigδθ m 99% SEQ ID No. 2232 484133 4356.2 484763 + Contigδδ m 99% SEQ ID No. 484717 2231 4355.2 485379 Contigδβ m 99% SEQ ID No. 2230 485369 4354.2 486358 Contigδ6 98% SEQ ID No. P 2229 486613 4353.2 486870 Contig56 SEQ ID No. P 3303 486710 787.3 487144 + Contigδ6 100% SEQ ID No. P 3304 487168 789.4 489210 + Contigδβ m 99% SEQ ID No. 2595 489432 4880.2 489803 + Contig56 m SEQ ID No. 489804 2596 490667 4881.4 4- Contig56 m SEQ ID No. 271 490946 494197 1331.3 Contig56 m 94% SEQ ID No. 272 494404 495648 1332.3 Contig56 m 97% SEQ ID No. 1749 495853 3626.1 497343 Contig56 m 99% SEQ ID No. 497301 4- 1748 497801 3625.2 Contig56 m 99% 497609 4- SEQ ID No. 1747 498991 3623.2 Contig56 m 97% SEQ ID No. 1746 498982 499392 3622.3 Contigδδ m 100% SEQ ID No. 2653 499393 4977.3 500466 Contig56 99% SEQ ID No. P 500591 2652 4974.1 501079 Contig56 m 100% SEQ ID No. 2536 501054 4793.2 501693 4- Contigδβ 99% SEQ ID No. 2534 P 502045 4790.3 603820 Contig56 99% SEQ ID No. P 2536 503795 506126 4791.3 Contig56 .m 100% SEQ ID No. 2633 504082 504483 4788.3 Contig56 m 100% SEQ ID No. 2656 606287 506961 + 4980.3 Contigδδ m 98% SEQ ID No. 2654 506912 4979.1 506536 4- Contigδβ 99% SEQ ID No. P 1265 506583 507191 2921.3 Contigδδ 99% P 507266 510616 96%
SEQ ID No. 1502 327.2 Contig56 m 510713 512743 97%
SEQ ID No. 1510 328.1 Contig56 m 512698 513521 98%
SEQ ID No. 1523 330.2 Contig56 m 513618 515047 99%
SEQ ID No. 1264 2920.1 Contig56 P 614374 514946 100%
SEQ ID No. 1262 2919.1 Contig56 m 516128 515365
SEQ ID No. 61 1003.3 Contig56 P 515479 517950 98%
SEQ ID No. 62 1006.2 Contig56 m 518054 518786 97%
SEQ ID No. 1261 2917.1 Contig56 P 519483 520199 97%
SEQ ID No. 1260 2916.1 Contig56 m 620040 620345 97%
SEQ ID No. 909 2360.2 Contig56 P 520200 521270 98%
SEQ ID No. 910 2361.4 Contig56 m 521370 527024 91 %
SEQ ID No. 1707 3568.3 Contig56 m 527089 528615 99%
SEQ ID No. 1706 3566.1 Contig56 m 528497 529690 98% +
SEQ ID No. 3251 712.2 Contig56 m 529698 530726 99% +
SEQ ID No. 3250 711.2 Contig56 m 530771 532873 98%
SEQ ID No. 3249 709.1 Contig56 P 531595 531951 95% +
SEQ ID No. 888 2335.2 Contig56 P 533008 534369 97% +
SEQ ID No. 889 2336.2 Contig56 P 534412 536484 98%
SEQ ID No. 1705 3564.2 Contig56 m 536875 537150 100%
SEQ ID No. 1704 3563.2 Contigδδ P 537264 537611 +
SEQ ID No. 3123 5988.1 Contig56 P 537425 537688 +
ORF Best-BlastP 10.1 Best-BlastP=> >nrprot 47% Identities = 35/90 (38%), Positives = 55/90 (61 %), Gaps = 5/90 (5%) dbj|BAC94688.11 hypothetical protein (Vibrio vulnificus YJ016] Length = 343
1000.2 Best-BlastP=> >nrprot 62% Identities = 120/257 (46%), Positives = 162/257 (63%), Gaps = 3/257 (1 %) ref|ZP_00079402.1 | COG1024: Enoyl- CoA hydratase/carnithine racemase [Geobacter metallireducens] Length = 272
1001.3 Best-BlastP=> >nrprot 97% Identities = 419/437 (95%), Positives = 425/437 (97%) gb|AAM00614.11 chemiosmotic efflux system protein C-like protein [Legionella pneumophila] Length = 448
1002.2 Best-BlastP=> >nrprot 11 % Identities = 35/129 (27%), Positives = 64/129 (49%), Gaps = 6/129 (4%) ref|ZP_00108772.1 | COG0642: Signal transduction histidine kinase [Nostoc punctiforme] Length = 2053
1003.3 Best-BlastP=> >nrprot No Hits found
1005.2 Best-BlastP=> >nrprot 67% Identities = 126/243 (51%), Positives = 165/243 (67%), Gaps = 4/243 (1 %) ref|NP_820690.1 | dihydrodipicolinate reductase [Coxiella burnetii RSA 493] sp|P24703|DAPB_COXBU Dihydrodipicolinate reductase (DHPR) gb|AAO91204.11 dihydrodipicolinate reductase [Coxiella burnetii RSA 493] Length = 239
1006.3 Best-BlastP=> >nrprot 61 % Identities = 96/211 (45%), Positives = 135/211 (63%), Gaps = 18/211 (8%) ref|NP_520174.11 PROBABLE TRANSMEMBRANE NADH DEHYDROGENASE I (CHAIN J) OXIDOREDUCTASE PROTEIN [Ralstonia solanacearum] emb|CAD15760.11 PROBABLE TRANSMEMBRANE NADH DEHYDROGENASE I (CHAIN J) OXIDOREDUCTASE PROTEIN [Ralstonia solanacearum] Length = 210
1009.2 Best-BlastP=> >nrprot 79% Identities = 113/159 (71 %), Positives = 132/169 (83%) ref|NP_297604.11 NADH-ubiquinone oxidoreductase, NQ09 subunit [Xylella fastidiosa 9a5c] ref|ZP_00039600.11 hypothetical protein [Xylella fastidiosa Dixon] ref|ZP_00041894.11 hypothetical protein [Xylella fastidiosa Ann-1] ref|NP_778493.11 NADH-ubiquinone oxidoreductase NQ09 subunit [Xylella fastidiosa Temeculal] pir||C82822 NADH-ubiquinone oxidoreductase, NQ09 subunit XF0313 [imported] - Xylella fastidiosa (strain 9a5c) gb|AAF83124.1 |AE003884_9 NADH- ubiquinone oxidoreductase, NQ09 subunit [Xylella fastidiosa 9a5c] gb|AA028142.11 NADH-ubiquinone oxidoreductase NQ09 subunit [Xylella fastidiosa Temeculal] Length = 163
1010.2 Best-BlastP=> >nrprot 82% Identities = 223/332 (67%), Positives = 283/332 (85%) ref|NP_820424.1 | NADH dehydrogenase I, H subunit [Coxiella burnetii RSA 493] gb|AAO90938.1 | NADH dehydrogenase I, H subunit [Coxiella burnetii RSA 493] Length = 340
1011.5 Best-BlastP=> >nrprot 65% Identities = 374/800 (46%), Positives = 511/800 (63%), Gaps = 32/800 (4%) ref|NP_820425.11 NADH dehydrogenase I, G subunit [Coxiella burnetii RSA 493] gb|AAO90939.11 NADH dehydrogenase I, G subunit [Coxiella burnetii RSA 493] Length = 787
1013.1 Best-BlastP=> >nrprot No Hits found
1014.2 Best-BlastP=> >nrprot 55% Identities = 163/485 (33%), Positives = 254/486 (52%), Gaps = 36/485 (7%) ref|NP_711514.11 putative amidase [Leptospira interrogans serovar lai str. 56601] gb|AAN48532.1 |AE011313_8 putative amidase [Leptospira interrogans serovar lai str. 56601] Length = 500
102.1 Best-BlastP=> >nrprot 92% Identities = 811/842 (96%), Positives = 830/842 (98%) gb|AAM00631.11 putative cation efflux transporter [Legionella pneumophila] Length = 842
040.3 Best-BlastP=> >nrprot 63% Identities = 150/366 (40%), Positives = 220/366 (60%), Gaps = 25/366 (6%) r hypothetical protein [Pseudomonas syringae pv. tomato str. DC3000] gb|AA058815.1 | conserved h syringae pv. tomato str. DC3000] Length = 368
1041.3 Best-BlastP=> >nrprot 82% Identities = 557/801 (69%), Positives = 663/801 (82%), Gaps = 1/801 (0%) re [Escherichia coli CFT073] gb|AAN83054.1 |AE016769_169 DNA gyrase subunit B [Escherichia coli CFT07
1042.5 Best-BlastP=> >nrprot 76% Identities = 264/406 (65%), Positives = 314/406 (77%) ref|NP_719647.11 mal [Shewanella oneidensis MR-1] gb|AAN57091.1 |AE015843_3 malate oxidoreductase, putative [Shewanella
1044.3 Best-BlastP=> >nrprot 59% Identities = 152/394 (38%), Positives = 236/394 (59%), Gaps = 12/394 (3%) r transporter [Coxiella burnetii RSA 493] gb|AAO90673.11 major facilitator family transporter [Coxiella burnetii
1047.2 Best-BlastP=> >nrprot 59% Identities = 142/261 (54%), Positives = 179/261 (68%), Gaps = 1/261 (0%) re acetyltransferases and hydrolases with the alpha/beta hydrolase fold [Desulfitobacterium hafniense]
1048.3 Best-BlastP=> >nrprot 12% Identities = 33/98 (33%), Positives = 48/98 (48%), Gaps = 3/98 (3%) ref|NP_7 [Shewanella oneidensis MR-1] gb|AAN54755.1 |AE015616_1 hypothetical protein [Shewanella oneidensis
1049.2 Best-BlastP=> >nrprot 48% Identities = 74/232 (31 %), Positives = 112/232 (48%), Gaps = 10/232 (4%) re transporter, periplasmic glutamine-binding protein (glnH) [Archaeoglobus fulgidus DSM 4304] pir||G periplasmic glutamine-binding protein (glnH) homolog - Archaeoglobus fulgidus gb|AAB91001.11 glu glutamine-binding protein (glnH) [Archaeoglobus fulgidus DSM 4304] Length = 264
1050.4 Best-BlastP=> >nrprot 14% Identities = 96/410 (23%), Positives = 164/410 (40%), Gaps = 57/410 (13%) p (EC 3.6.3.8), plasma membrane - Paramecium tetraurelia gb|AAB81284.11 plasma membrane calci Length = 1160
1053.2 Best-BlastP=> >nrprot 64% Identities = 86/174 (49%), Positives = 114/174 (65%), Gaps = 2/174 (1%) ref| [Coxiella burnetii RSA 493] gb|AA091485.1 | phosphatase, putative [Coxiella burnetii RSA 493] Leng
1055.1 Best-BlastP=> >nrprot 59% Identities = 102/256 (39%), Positives = 144/256 (56%), Gaps = 16/256 (6%) r hypothetical protein [Coxiella burnetii RSA 493] gb|AA091367.11 conserved hypothetical protein [Coxiella b
1056.1 Best-BlastP=> >nrprot No Hits found
1058.3 Best-BlastP=> >nrprot 43% Identities = 112/611 (21 %), Positives = 204/611 (39%), Gaps = 94/611 (18%) heavy chain, non muscle pir||A2665δ myosin heavy chain [similarity] - slime mold (Dictyostelium dis heavy chain Length = 2116
1059.2 Best-BlastP=> >nrprot 85% Identities = 302/423 (71 %), Positives = 364/423 (86%), Gaps = 2/423 (0%) ref|NP_820394.11 citrate synthase [Coxiella burnetii RSA 493] sp|P18789|CISY_COXBU Citrate synthase pir||JQ1392 citrate (si)-synthase (EC 4.1.3.7) - Coxiella burnetii gb|AAA23307.11 citrate synthase (gltA) (EC 4.1.3.7) gb|AAO90908.11 citrate synthase [Coxiella burnetii RSA 493] Length = 430
1060.2 Best-BlastP=> >nrprot 57% Identities = 114/286 (39%), Positives = 168/286 (58%), Gaps = 7/286 (2%) ref|NP_813465.11 purine nucleoside phosphorylase II [Bacteroides thetaiotaomicron VPI-5482] gb|AA079659.11 purine nucleoside phosphorylase II [Bacteroides thetaiotaomicron VPI-5482] Length = 292
1063.1 Best-BlastP=> >nrprot No Hits found
1065.3 Best-BlastP=> >nrprot 48% Identities = 124/365 (33%), Positives = 187/365 (51 %), Gaps = 10/365 (2%) dbj|BAA31547.11 metal-activated pyridoxal enzyme [Arthrobacter sp.] Length = 379
1066.2 Best-BlastP=> >nrprot 53% Identities = 124/294 (42%), Positives = 160/294 (54%), Gaps = 18/294 (6%) ref|ZP_00016110.11 COG0596: Predicted hydrolases or acyltransferases (alpha/beta hydrolase superfamily) [Rhodospirillum rubrum] Length = 286
1067.1 Best-BlastP=> >nrprot 47% Identities = 28/70 (40%), Positives = 40/70 (57%), Gaps = 11/70 (15%) ref|NP_489272.11 unknown protein [Nostoc sp. PCC 7120] pir||AH2469 hypothetical protein alr5232 [imported] - Nostoc sp. (strain PCC 7120) dbj|BAB76931.11 ORF_ID:alr5232~unknown protein [Nostoc sp. PCC 7120] Length = 204
1069.2 Best-BlastP=> >nrprot 46% Identities = 103/273 (37%), Positives = 145/273 (53%), Gaps = 36/273 (12%) emb|CAA75849.11 hypothetical protein [Coxiella burnetii] Length = 309
107.1 Best-BlastP=> >nrprot 99% Identities = 1034/1047 (98%), Positives = 1041/1047 (99%) gb|AAM00628.11 chemiosmotic efflux system B protein A [Legionella pneumophila] Length = 1047
1071.3 Best-BIastP=> >nrprot 99% Identities = 411/418 (98%), Positives = 416/418 (99%) pir||T08882 proline/betaine transport protein homolog CitA - Legionella pneumophila emb|CAA75171.1 | TphA protein [Legionella pneumophila] gb|AAC38182.1 | CitA [Legionella pneumophila] emb|CAA76337.1 | TphA protein [Legionella pneumophila] Length = 418
1072.4 Best-BlastP=> >nrprot 99% Identities = 966/973 (99%), Positives = 971/973 (99%) pir||T18341 icmF protein - Legionella pneumophila emb|CAA75172.1 | IcmF protein [Legionella pneumophila] emb|CAA76338.1 | IcmF protein [Legionella pneumophila] Length = 973
1074.3 Best-BlastP=> >nrprot 67% Identities = 343/638 (53%), Positives = 456/638 (71 %), Gaps = 6/638 (0%) ref|NP_819606.11 fatty oxidation complex, alpha subunit [Coxiella burnetii RSA 493] gb|AAO90120.11 fatty oxidation complex, alpha subunit [Coxiella burnetii RSA 493] Length = 642
1075.3 Best-BlastP=> >nrprot 75% Identities = 271/422 (64%), Positives = 333/422 (78%), Gaps = 1/422 (0%) emb|CAD58320.11 beta-Subunit of fatty acid oxidation complex, 3-keto-acyl-CoA-thiolase [Azoarcus sp. EbN1] Length = 426
1076.3 Best-BlastP=> >nrprot 56% Identities = 50/1 19 (42%), Positives = 70/119 (58%) ref|NP_90201 1.1 1 conserved hypothetical protein [Chromobacterium violaceum ATCC 12472] gb|AAQ60013.1 | conserved hypothetical protein [Chromobacterium violaceum ATCC 12472] Length = 122
1077.1 Best-BlastP=> >nrprot 75% Identities = 137/226 (60%), Positives = 175/226 (77%), Gaps = 1/226 (0%) ref|NP_384333.1 | PUTATIVE TRANSCRIPTION REGULATOR PROTEIN [Sinorhizobium meliloti] emb|CAC41614.1 | PUTATIVE TRANSCRIPTION REGULATOR PROTEIN [Sinorhizobium meliloti] Length = 230
CN CN CM co CM CO CM CN co co ,- CO
CO --^ N CO σ d -- CN co CD
Is- oo co oo co CO oo σi CO CJ3 CO CO σi C35 o o o o o O o o o o o o o O
1098.3 Best-BlastP=> >nrprot 57% Identities = 136/330 (41 %), Positives = 195/330 (59%), Gaps = 3/330 (0%) ref|NP_718989.11 conserved hypothetical protein [Shewanella oneidensis MR-1] sp|Q8EBR4|TRUD_SHEON tRNA pseudouridine synthase D (Pseudouridylate synthase) (Uracil hydrolyase) gb|AAN56433.1 |AE015780_4 conserved hypothetical protein [Shewanella oneidensis MR-1] Length = 370
1099.2 Best-BlastP=> >nrprot 63% Identities = 168/328 (48%), Positives = 207/328 (63%), Gaps = 6/328 (1 %) ref|NP_403834.11 conserved hypothetical protein [Yersinia pestis] ref |NP_671254.11 hypothetical protein [Yersinia pestis KIM] pir||AH0022 conserved hypothetical protein YPO0180 [imported] - Yersinia pestis (strain C092) emb|CAC89042.1 | conserved hypothetical protein [Yersinia pestis C092] gb|AAM87505.1 |AE014000_9 hypothetical protein [Yersinia pestis KIM] Length = 326
11.2 Best-BlastP=> >nrprot 39% Identities = 77/276 (27%), Positives = 127/276 (46%), Gaps = 29/276 (10%) ref |ZP_00023112.11 hypothetical protein [Ralstonia metallidurans] Length = 348
1101.4 Best-BlastP=> >nrprot 54% Identities = 128/359 (35%), Positives = 200/359 (56%), Gaps = 24/369 (6%) ref|NP_878747.11 histidinol-phosphate aminotransferase [Candidatus Blochmannia floridanus] emb|CAD83523.1 | histidinol-phosphate aminotransferase [Candidatus Blochmannia floridanus] Length = 356
1102.2 Best-BlastP=> >nrprot 66% Identities = 203/427 (47%), Positives = 287/427 (67%), Gaps = 9/427 (2%) dbj|BAC94115.11 histidinol dehydrogenase [Vibrio vulnificus YJ016] Length = 431
1105.2 Best-BlastP=> >nrprot 36% Identities = 187/439 (42%), Positives = 279/439 (63%), Gaps = 1/439 (0%) ref|NP_715981.11 sensory box protein [Shewanella oneidensis MR-1] gb|AAN53426.1 |AE015481_9 sensory box protein [Shewanella oneidensis MR-1] Length = 1515
1106.5 Best-BlastP=> >nrprot 99% Identities = 1366/1368 (99%), Positives = 1367/1368 (99%) gb|AAC69338.11 RNA polymerase B-subunit [Legionella pneumophila] Length = 1368
1107.2 Best-BlastP=> >nrprot 66% Identities = 154/324 (47%), Positives = 219/324 (67%), Gaps = 3/324 (0%) ref|NP_657119.1| Chal_stil_syntC, Chalcone and stilbene synthases, C-terminal domain [Bacillus anthracis A2012] ref|NP_845551.11 3-oxoacyl-(acyl-carrier-protein) synthase III, putative [Bacillus anthracis str. Ames] gb|AAP27037.1 | 3-oxoacyl-(acyl-carrier-protein) synthase III, putative [Bacillus anthracis str. Ames] Length = 329
1109.3 Best-BlastP=> >nrprot 64% Identities = 150/329 (45%), Positives = 212/329 (64%), Gaps = 3/329 (0%) ref|NP_657117.11 3Beta_HSD, 3-beta hydroxysteroid dehydrogenase/isomerase family [Bacillus anthracis A2012] ref|NP_84δδ49.11 3-beta hydroxysteroid dehydrogenase/isomerase family protein [Bacillus anthracis str. Ames] gb|AAP2703δ.1| 3-beta hydroxysteroid dehydrogenase/isomerase family protein [Bacillus anthracis str. Ames] Length = 328
111.2 Best-BlastP=> >nrprot 97% Identities = 485/504 (96%), Positives = 493/504 (97%) sp|Q8RNP4|TYPH_LEGPN Putative thymidine phosphorylase (TdRPase) gb|AAM00626.1 | unknown [Legionella pneumophila] Length = 517
1111.3 Best-BlastP=> >nrprot 58% Identities = 109/270 (40%), Positives = 164/270 (60%), Gaps = 6/270 (2%) ref|NP_657116.11 hypothetical protein predicted by GeneMark [Bacillus anthracis A2012] ref|NP_84δδ48.11 conserved hypothetical protein [Bacillus anthracis str. Ames] gb|AAP27034.1| conserved hypothetical protein [Bacillus anthracis str. Ames] Length = 283
1113.2 Best-BlastP=> >nrprot 31% Identities = 37/58 (63%), Positives = 42/58 (72%), Gaps = 3/58 (5%) ref|NP_769576.11 Conserved hypothetical protein [Vibrio vulnificus CMCP6] gb|AAO09103.1|AE016799_1 Conserved hypothetical protein [Vibrio vulnificus CMCP6] Length = 116
1115.4 Best-BlastP=> >nrprot 26% Identities = 41/132 (31 %), Positives = 70/132 (53%), Gaps = 10/132 (7%) ref|NP_710625.1 | conserved hypothetical protein [Leptospira interrogans serovar lai str. 66601] gb|AAN47643.1 |AE011231_6 conserved hypothetical protein [Leptospira interrogans serovar lai str. 56601] Length = 359
1116.2 Best-BlastP=> >nrprot No Hits found
1118.2 Best-BlastP=> >nrprot 79% Identities = 277/462 (61 %), Positives = 359/462 (79%), Gaps = 1/452 (0%) gb|AAM00627. 1 unknown [Legionella pneumophila] Length = 470
1119.2 Best-BlastP=> >nrprot No Hits found
1122.1 Best-BlastP=> >nrprot 64% Identities = 142/291 (48%), Positives = 196/291 (67%), Gaps = 5/291 (1 %) ref|NP_716239.11 hflC protein [Shewanella oneidensis MR-1] gb|AAN53684.1 |AE015507_10 hflC protein [Shewanella oneidensis MR-1] Length = 297
1123.2 Best-BlastP=> >nrprot 62% Identities = 176/379 (46%), Positives = 237/379 (62%), Gaps = 24/379 (6%) ref|NP_253629.11 protease subunit HflK [Pseudomonas aeruginosa PA01] pir||B83028 proteinase subunit HflK PA4942 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG08327.11 AE004907_5 protease subunit HflK [Pseudomonas aeruginosa PA01 ] Length = 400
1124.2 Best-BlastP=> >nrprot 36% Identities = 58/152 (38%), Positives = 87/152 (57%), Gaps = 8/152 (5%) gb|AAM09314.2| similar to Mus musculus (Mouse). Uridine-cytidine kinase 2 [Dictyostelium discoideum] Length = 243
1125.2 Best-BlastP=> >nrprot 68% Identities = 193/382 (50%), Positives = 265/382 (69%) ref |NP_519762.11 PROBABLE ACETYLORNITHINE DEACETYLASE (ACETYLORNITHINASE) PROTEIN [Ralstonia solanacearum] emb|CAD15343.11 PROBABLE ACETYLORNITHINE DEACETYLASE (ACETYLORNITHINASE) PROTEIN [Ralstonia solanacearum] Length = 397
1126.3 Best-BlastP=> >nrprot 55% Identities = 163/398 (40%), Positives = 239/398 (60%), Gaps = 2/398 (0%) ref|ZP_00033611.11 COG0477: Permeases of the major facilitator superfamily [Burkholderia fungorum] Length = 470
1127.3 Best-BlastP=> >nrprot 72% Identities = 360/650 (55%), Positives = 472/650 (72%), Gaps = 4/650 (0%) ref|NP_745215.11 acetoacetyl-CoA synthetase, putative [Pseudomonas putida KT2440] gb|AAN68679.1 |AE016497_4 acetoacetyl-CoA synthetase, putative [Pseudomonas putida KT2440] Length = 650
1128.3 Best-BlastP=> >nrprot 50% Identities = 113/289 (39%), Positives = 170/289 (58%), Gaps = 13/289 (4%) ref|NP_904046.11 biotin synthesis protein [Chromobacterium violaceum ATCC 12472] gb|AAQ62035.1 | biotin synthesis protein [Chromobacterium violaceum ATCC 12472] Length = 302
1132.2 Best-BlastP=> >nrprot 57% Identities = 85/221 (38%), Positives = 135/221 (61 %), Gaps = 12/221 (5%) ref|ZP_00086846.11 COG1040: Predicted amidophosphoribosyltransferases [Pseudomonas fluorescens PfO-1] Length = 246
1133.3 Best-BlastP=> >nrprot 60% Identities = 103/346 (29%), Positives = 172/346 (49%), Gaps = 13/346 (3%) ref|ZP_00082200.1 | COG0750: Predicted membrane-associated Zn-dependent proteases 1 [Geobacter metallireducens] Length = 356
1134.3 Best-BlastP=> >nrprot No Hits found
1136.3 Best-BlastP=> >nrprot 62% Identities = 232/442 (52%), Positives = 307/442 (69%), Gaps = 1/442 (0%) ref|ZP_00043253.11 COG0790: FOG: TPR repeat, SEL1 subfamily [Magnetococcus sp. MC-1] Length = 831
1137.2 Best-BlastP=> >nrprot No Hits found
114.2 Best-BlastP=> >nrprot No Hits found
1142.4 Best-BlastP=> >nrprot 65% Identities = 222/454 (48%), Positives = 312/454 (68%), Gaps = 4/454 (0%) ref|NP_820336.11 amino acid antiporter [Coxiella burnetii RSA 493] gb|AAO90850.11 amino acid antiporter [Coxiella burnetii RSA 493] Length = 474
1144.3 Best-BlastP=> >nrprot 64% Identities = 180/379 (47%), Positives = 254/379 (67%), Gaps = 4/379 (1%) ref|NP_800676.11 conserved hypothetical protein [Vibrio parahaemolyticus RIMD 2210633] sp|Q87GZ9|CLCA_VIBPA Voltage-gated ClC-type chloride channel clcA dbj|BAC62509.11 conserved hypothetical protein [Vibrio parahaemolyticus] Length = 467
1145.3 Best-BlastP=> >nrprot 72% Identities = 391/721 (64%), Positives = 622/721 (72%), Gaps = 11/721 (1%) ref |ZP_00110122.11 COG0612: Anthranilate/para-aminobenzoate synthases component II [Nostoc punctiforme] Length = 734
1148.2 Best-BlastP=> >nrprot 66% Identities = 334/906 (36%), Positives = 512/906 (56%), Gaps = 35/906 (3%) ref|NP_289628.11 adenylylating enzyme for glutamine synthetase [Escherichia coli 0157:H7 EDL933] ref|NP_311963.11 glutamate-ammonia-ligase adenylyltransferase [Escherichia coli 0157:H7] pir||H91120 glutamate-ammonia-ligase adenylyltransferase [imported] - Escherichia coli (strain 0157:H7, substrain RIMD 0509962) pir||G85965 adenylylating enzyme for glutamine synthetase [imported] - Escherichia coli (strain 0157:H7, substrain EDL933) gb|AAG58187.1 |AE005534_9 adenylylating enzyme for glutamine synthetase [Escherichia coli 0157:H7 EDL933] dbj|BAB37359.11 glutamate-ammonia-ligase adenylyltransferase [Escherichia coli 0157:H7] Length = 946
115.2 Best-BlastP=> >nrprot 57% Identities = 53/167 (31 %), Positives = 98/167 (58%), Gaps = 5/167 (2%) ref|NP_490158.1 | probable acetyltransferase [Nostoc sp. PCC 7120] pir||AD2484 hypothetical protein alr7052 [imported] - Nostoc sp. (strain PCC 7120) plasmid pCC7120alpha dbj|BAB78136.11 ORF_ID:alr7052~probable acetyltransferase [Nostoc sp. PCC 7120] Length = 169
1152.2 Best-BlastP=> >nrprot 22% Identities = 39/137 (28%), Positives = 67/137 (48%), Gaps = 10/137 (7%) ref|NP_824638.11 putative lipase [Streptomyces avermitilis MA-4680] dbj|BAC71173.11 putative lipase [Streptomyces avermitilis MA-4680] Length = 286
1153.1 Best-BlastP=> >nrprot 46% Identities = 36/136 (26%), Positives = 67/136 (49%), Gaps = 15/136 (11 %) ref|NP_629974.11 putative integral membrane protein [Streptomyces coelicolor A3(2)] pir||T35887 hypothetical protein SC9B10.18 - Streptomyces coelicolor emb|CAA15808.11 putative integral membrane protein [Streptomyces coelicolor A3(2)] Length = 312
1156.3 Best-BlastP=> >nrprot 48% Identities = 114/437 (26%), Positives = 213/437 (48%), Gaps = 37/437 (8%) ref|ZP_00107812.11 COG4325: Predicted membrane protein [Nostoc punctiforme] Length = 449
1157.4 Best-BlastP=> >nrprot 73% Identities = 650/943 (58%), Positives = 687/943 (72%), Gaps = 15/943 (1 %) ref |ZP_00085403.11 COG0567: 2- oxoglutarate dehydrogenase complex, dehydrogenase (E1) component, and related enzymes [Pseudomonas fluorescens PfO-1] Length = 943
1159.5 Best-BlastP=> >nrprot 30% Identities = 61/269 (22%), Positives = 130/269 (48%), Gaps = 23/269 (8%) ref|NP_143635.11 chromosome assembly protein [Pyrococcus horikoshii] pir||F71190 probable chromosome assembly protein - Pyrococcus horikoshii dbj|BAA30917.11 1179aa long hypothetical chromosome assembly protein [Pyrococcus horikoshii] Length = 1179
116.1 Best-BlastP=> >nrprot 29% Identities = 42/123 (34%), Positives = 47/123 (38%), Gaps = 27/123 (21%) ref|NP_639328.11 hypothetical protein [Xanthomonas campestris pv. campestris str. ATCC 33913] gb|AAM43210.1 | hypothetical protein [Xanthomonas campestris pv. campestris str. ATCC 33913] Length = 131
1160.2 Best-BlastP=> >nrprot 51% Identities = 136/392 (34%), Positives = 206/392 (52%), Gaps = 10/392 (2%) ref|NP_085189.1 | IS10 orf [Shigella flexneri] ref |NP_858160.11 hypothetical protein [Shigella flexneri 2a] gb|AAK18345.1 |AF348706_34 IS10 orf [Shigella flexneri] gb|AAL72480.1 | hypothetical protein [Shigella flexneri 2a] Length = 407
,- m co ed σi
— •* co r-. r-- r-- TO TO
1186.2 Best-BlastP=> >nrprot 31% Identities = 52/171 (30%), Positives = 82/171 (47%), Gaps = 7/171 (4%) ref | iridescent virus 6] gb|AAK82096.1 |AF303741_235 236L [Chilo iridescent virus] Length = 265
1188.2 Best-BlastP=> >nrprot 99% Identities = 737/749 (98%), Positives = 746/749 (99%) sp|Q9WXB9|CATA_L peroxidase) dbj|BAA78342.1 | catalase-peroxidase [Legionella pneumophila] gb|AAG37106.1 |AF276752_ pneumophila] Length = 74g
119.1 Best-BlastP=> >nrprot 51 % Identities = 53/124 (42%), Positives = 77/124 (62%), Gaps = 2/124 (1 %) dbj| Neisseria meningitidis [Actinobacillus actinomycetemcomitans] Length = 255
1190.4 Best-BlastP=> >nrprot 56% Identities = 256/709 (36%), Positives = 406/709 (57%), Gaps = 25/709 (3%) Pyruvate/2-oxoglutarate dehydrogenase complex, dihydrolipoamide dehydrogenase (E3) compone [Microbulbifer degradans 2-40] Length = 704
1193.2 Best-BlastP=> >nrprot No Hits found
1194.3 Best-BlastP=> >nrprot 72% Identities = 223/383 (58%), Positives = 278/383 (72%), Gaps = 1/383 (0%) r dehydrogenase [Xanthomonas campestris pv. campestris str. ATCC 33913] gb|AAM40726.1 | gluta campestris pv. campestris str. ATCC 33913] Length = 387
1199.4 Best-BlastP=> >nrprot 80% Identities = 73/110 (66%), Positives = 90/110 (81%) ref|ZP_00067987.1 | CO [Microbulbifer degradans 2-40] Length = 110
1201.1 Best-BlastP=> >nrprot 82% Identities = 150/220 (68%), Positives = 180/220 (81%), Gaps = 1/220 (0%) r [Coxiella burnetii RSA 493] sp|085388|RS3_COXBU 30S ribosomal protein S3 gb|AAO89802.1 | ribosom Length = 227
1202.2 Best-BlastP=> >nrprot 89% Identities = 113/137 (82%), Positives = 124/137 (90%) ref|NP_742627.1| rib putida KT2440] gb|AAN66091.1 |AE016238_9 ribosomal protein L16 [Pseudomonas putida KT2440]
1205.5 Best-BlastP=> >nrprot 16% Identities = 428/1182 (36%), Positives = 630/1182 (53%), Gaps = 81/1182 ( protein [Nostoc punctiforme] Length = 2315
1207.2 Best-BlastP=> >nrprot No Hits found
1208.1 Best-BlastP=> >nrprot 42% Identities = 46/124 (36%), Positives = 56/124 (44%), Gaps = 4/124 (3%) pir|| Pyrococcus horikoshii dbj|BAA29381.1 | 215aa long hypothetical protein [Pyrococcus horikoshii] Le
1209.2 Best-BlastP=> >nrprot 84% Identities = 346/487 (71%), Positives = 417/487 (85%), Gaps = 1/487 (0%) r dehydrogenase/GMP reductase [Pseudomonas fluorescens PfO-1] Length = 506
121.1 Best-BlastP=> >nrprot 59% Identities = 58/101 (57%), Positives = 67/101 (66%), Gaps = 1/101 (0%) ref| Uncharacterized protein conserved in bacteria [Azotobacter vinelandii] Length = 160
1210.2 Best-BlastP=> >nrprot 81% Identities = 364/523 (67%), Positives = 427/523 (81%), Gaps = 1/523 (0%) r [Shigella flexneri 2a str. 301] ref |NP_838070.11 GMP synthetase (glutamine-hydrolyzing) [Shigella flexneri gb|AAN44053.1|AE015270_8 GMP synthetase [Shigella flexneri 2a str. 301] gb|AAP17880.11 GMP synth flexneri 2a str. 2457T] Length = 525
1211.3 Best-BlastP=> >nrprot 19% Identities = 47/90 (52%), Positives = 69/90 (76%), Gaps = 1/90 (1 %) ref|NP_929783.11 hypothetical protein [Photorhabdus luminescens subsp. laumondii TT01] emb|CAE14921.11 unnamed protein product [Photorhabdus luminescens subsp. laumondii TT01] Length = 94
1213.2 Best-BlastP=> >nrprot 65% Identities = 193/394 (48%), Positives = 248/394 (62%), Gaps = 22/394 (5%) ref|NP_249543.11 chitin-binding protein CbpD precursor [Pseudomonas aeruginosa PA01] pir||F83538 chitin-binding protein CbpD precursor PA0852 [imported] - Pseudomonas aeruginosa (strain PA01) gb|AAG04241.1 |AE004520_4 chitin-binding protein CbpD precursor [Pseudomonas aeruginosa PA01] Length = 389
121 δ.3 Best-BlastP=> >nrprot 64% Identities = 172/363 (47%), Positives = 233/363 (64%), Gaps = δ/363 (1 %) ref|NP_387138.11 CONSERVED HYPOTHETICAL PROTEIN [Sinorhizobium meliloti] emb|CAC47611.1 | CONSERVED HYPOTHETICAL PROTEIN [Sinorhizobium meliloti] Length = 367 122.3 Best-BlastP=> >nrprot 69% Identities = 90/169 (66%), Positives = 115/169 (72%), Gaps = 4/159 (2%) ref|NP_779515.11 chromosome partitioning protein [Xylella fastidiosa Temeculal] sp|Q87BY1 |PARB_XYLFT Probable chromosome partitioning protein parB gb|AA029164.1 | chromosome partitioning protein [Xylella fastidiosa Temeculal] Length = 310
1220.3 Best-BlastP=> >nrprot 77% Identities = 332/507 (65%), Positives = 396/507 (77%), Gaps = 3/507 (0%) dbj|BAB19801.1 | piperideine-6- carboxylate dehydrogenase ['Flavobacterium' lutescens] Length = 510
1222.5 Best-BlastP=> >nrprot No Hits found
1223.2 Best-BlastP=> >nrprot 39% Identities = 73/287 (25%), Positives = 119/287 (41 %), Gaps = 38/287 (13%) ref|NP_521275.11 PROBABLE SIGNAL PEPTIDE PROTEIN [Ralstonia solanacearum] emb|CAD16942.11 PROBABLE SIGNAL PEPTIDE PROTEIN [Ralstonia solanacearum] Length = 278 1224.2 Best-BlastP=> >nrprot 88% Identities = 212/266 (79%), Positives = 237/266 (89%) emb|CAC35728.11 OXA-29 [Fluoribacter gormanii] Length = 266 1225.2 Best-BlastP=> >nrprot 40% Identities = 90/292 (30%), Positives = 143/292 (48%), Gaps = 26/292 (8%) ref|NP_855062.11 CONSERVED HYPOTHETICAL PROTEIN [Mycobacterium bovis subsp. bovis AF2122/97] emb|CAD94271.11 CONSERVED HYPOTHETICAL PROTEIN [Mycobacterium bovis subsp. bovis AF2122/97] Length = 439
1226.2 Best-BlastP=> >nrprot 56% Identities = 129/350 (36%), Positives = 190/350 (54%), Gaps = 22/360 (6%) ref|NP_629326.11 putative sulfurylase [Streptomyces coelicolor A3(2)] emb|CAC01308.11 putative sulfurylase [Streptomyces coelicolor A3(2)] Length = 392
1227.1 Best-BlastP=> >nrprot 48% Identities = 66/202 (32%), Positives = 109/202 (53%), Gaps = 10/202 (4%) ref|ZP_00107215.11 COG0637: Predicted phosphatase/phosphohexomutase [Nostoc punctiforme] Length = 242
1229.3 Best-BlastP=> >nrprot No Hits found 123.2 Best-BlastP=> >nrprot 59% Identities = 201/462 (43%), Positives = 284/462 (61 %), Gaps = 28/462 (6%) ref|NP_253634.1 | N-acetylmuramoyl-L- alanine amidase [Pseudomonas aeruginosa PA01] pir||G83028 N-acetylmuramoyl-L-alanine amidase PA4947 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG08332.1 |AE004907_10 N-acetylmuramoyl-L-alanine amidase [Pseudomonas aeruginosa PA01] Length = 475
1230.2 Best-BlastP=> >nrprot 81% Identities = 364/516 (70%), Positives = 429/516 (83%) ref|NP_819831.1 | peptide chain release factor 3 [Coxiella burnetii RSA 493] gb|AAO90345.11 peptide chain release factor 3 [Coxiella burnetii RSA 493] Length = 525
1231.3 Best-BlastP=> >nrprot No Hits found
1235.3 Best-BlastP=> >nrprot 28% Identities = 44/137 (32%), Positives = 70/137 (51%), Gaps = 1/137 (0%) ref|NP_800044.11 putative acetyltransferase [Vibrio parahaemolyticus RIMD 2210633] dbj|BAC61877.1 | putative acetyltransferase [Vibrio parahaemolyticus] Length = 140
1236.2 Best-BlastP=> >nrprot 65% Identities = 172/176 (97%), Positives = 174/176 (98%) gb|AAM00633.1 | unknown [Legionella pneumophila] Length = 176
1237.2 Best-BlastP=> >nrprot 50% Identities = 47/132 (35%), Positives = 73/132 (55%), Gaps = 4/132 (3%) ref|NP_832446.11 Acetyltransferase [Bacillus cereus ATCC 14579] gb|AAP09647.11 Acetyltransferase [Bacillus cereus ATCC 14579] Length = 141
1238.2 Best-BlastP=> >nrprot No Hits found
1239.2 Best-BlastP=> >nrprot No Hits found
124.1 Best-BlastP=> >nrprot 63% Identities = 66/144 (45%), Positives = 95/144 (65%) ref|NP_716232.11 conserved hypothetical protein TIGR00150 [Shewanella oneidensis MR-1] gb|AAN63677.1 |AE015607_3 conserved hypothetical protein TIGR00150 [Shewanella oneidensis MR-1] Length = 152
1242.2 Best-BlastP=> >nrprot 82% Identities = 138/191 (72%), Positives = 160/191 (83%) ref|NP_353171.1 | AGR_C_216p [Agrobacterium tumefaciens] ref|NP_530843.11 uracil phosphoribosyltransferase [Agrobacterium tumefaciens str. C58 (U. Washington)] sp|Q8UJ06|UPP_AGRT5 Uracil phosphoribosyltransferase (UMP pyrophosphorylase) (UPRTase) pir||C97375 uracil phosphoribosyltransferase (UMP pyrophosphorylase) (uprtase) [imported] - Agrobacterium tumefaciens (strain C58, Cereon) pir||AI2592 uracil phosphoribosyltransferase [imported] - Agrobacterium tumefaciens (strain C58, Dupont) gb|AAK85956.11 AGR_C_216p [Agrobacterium tumefaciens str. C58 (Cereon)] gb|AAL41159.11 uracil phosphoribosyltransferase [Agrobacterium tumefaciens str. C58 (U. Washington)] Length = 209
1244.2 Best-BlastP=> >nrprot No Hits found
1245.1 Best-BlastP=> >nrprot No Hits found
1246.2 Best-BlastP=> >nrprot 22% Identities = 82/298 (27%), Positives = 130/298 (43%), Gaps = 56/298 (18%) pir||T09051 PepA protein - Pseudomonas aeruginosa gb|AAC16023.1 | ExoU [Pseudomonas aeruginosa] gb|AAC38269.1 | PepA [Pseudomonas aeruginosa] gb|AAP82959.11 type III effector protein [Pseudomonas aeruginosa] Length = 687
1249.2 Best-BlastP=> >nrprot 30% Identities = 37/119 (31%), Positives = 57/119 (47%), Gaps = 9/119 (7%) emb|CAD21525.11 hypothetical protein [Taenia solium] Length = 155
125.2 Best-BlastP=> >nrprot 59% Identities = 227/486 (46%), Positives = 293/486 (60%), Gaps = 4/486 (0%) ref|NP_820087.11 conserved hypothetical protein [Coxiella burnetii RSA 493] gb|AAOg0601.11 conserved hypothetical protein [Coxiella burnetii RSA 493] Length = 488
1250.1 Best-BlastP=> >nrprot 64% Identities = 74/160 (46%), Positives = 105/160 (65%), Gaps = 1/160 (0%) ref|NP_820968.11 dihydrofolate reductase [Coxiella burnetii RSA 493] gb|AA091482.11 dihydrofolate reductase [Coxiella burnetii RSA 493] Length = 161
1251.2 Best-BlastP=> >nrprot 68% Identities = 176/321 (54%), Positives = 222/321 (69%), Gaps = 4/321 (1 %) ref|NP_249284.11 pyridoxal phosphate biosynthetic protein PdxA [Pseudomonas aeruginosa PA01] sp|Q9l5U4|PXA1_PSEAE 4-hydroxythreonine-4-phosphate dehydrogenase 1 (4-(phosphohydroxy)-L-threonine dehydrogenase 1 ) pir||A83572 pyridoxal phosphate biosynthetic protein PdxA PA0593 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG03982.1 |AE004495_6 pyridoxal phosphate biosynthetic protein PdxA [Pseudomonas aeruginosa PA01] Length = 328
1257.6 Best-BlastP=> >nrprot 69% Identities = 242/415 (58%), Positives = 300/41 δ (72%), Gaps = δ/41 δ (1 %) ref|NP_820861.11 conserved hypothetical protein [Coxiella burnetii RSA 493] gb|AA091365.11 conserved hypothetical protein [Coxiella burnetii RSA 493] Length = 435
1258.1 Best-BlastP=> >nrprot 67% Identities = 175/312 (56%), Positives = 217/312 (69%), Gaps = 2/312 (0%) ref|ZP_00125152.11 COG0189: Glutathione synthase/Ribosomal protein S6 modification enzyme (glutaminyl transferase) [Pseudomonas syringae pv. syringae B728a] Length = 319
126.2 Best-BlastP=> >nrprot 99% Identities = 470/475 (98%), Positives = 475/475 (100%) emb|CAD42896.11 flagellin [Legionella pneumophila serogroup 1] Length = 475
1260.3 Best-BlastP=> >nrprot 48% Identities = 92/211 (43%), Positives = 126/211 (59%), Gaps = 3/211 (1 %) ref |NP_521080.11 PROBABLE LIPOPROTEIN PRECURSOR (VACJ) TRANSMEMBRANE [Ralstonia solanacearum] emb|CAD16666.11 PROBABLE LIPOPROTEIN PRECURSOR (VACJ) TRANSMEMBRANE [Ralstonia solanacearum] Length = 269
1261.2 Best-BlastP=> >nrprot 66% Identities = 286/564 (50%), Positives = 370/564 (65%), Gaps = 8/564 (1 %) gb|AAB16855.11 pyruvate decarboxylase [Arabidopsis thaliana] Length = 607
1264.3 Best-BlastP=> >nrprot 39% Identities = 97/291 (33%), Positives = 159/291 (54%), Gaps = 16/291 (5%) ref|NP_899726.11 probable aminopeptidase [Chromobacterium violaceum ATCC 12472] gb|AAQ57736.11 probable aminopeptidase [Chromobacterium violaceum ATCC 12472] Length = 415
1265.5 Best-BlastP=> >nrprot 58% Identities = 119/262 (45%), Positives = 162/262 (61 %), Gaps = 6/262 (2%) ref|ZP_00051284.11 COG0656: Aldo/keto reductases, related to diketogulonate reductase [Magnetospirillum magnetotacticum] Length = 291
1267.3 Best-BlastP=> >nrprot 20% Identities = 38/108 (35%), Positives = 56/108 (50%), Gaps = 8/108 (7%) ref|NP_616727.11 conserved hypothetical protein [Methanosarcina acetivorans str. C2A] gb|AAM05207.1 | conserved hypothetical protein [Methanosarcina acetivorans str. C2A] Length = 266
1268.2 Best-BlastP=> >nrprot 46% Identities = 82/293 (27%), Positives = 144/293 (49%), Gaps = 33/293 (11%) ref|XP_313252.11 ENSANGP00000010487 [Anopheles gambiae] gb|EAA08759.11 ENSANGP00000010487 [Anopheles gambiae str. PEST] Length = 310
127.4 Best-BlastP=> >nrprot 37% Identities = 32/94 (34%), Positives = 48/94 (51 %), Gaps = 6/94 (6%) sp|Q02910|CPN_DROME Calphotin pir||A47282 calcium-binding protein calphotin - fruit fly (Drosophila melanogaster) gb|AAA28405.11 calcium-binding protein Length = 865
1270.2 Best-BlastP=> >nrprot 70% Identities = 73/144 (50%), Positives = 102/144 (70%), Gaps = 2/144 (1 %) ref|NP_241923.1 | BH1057~unknown conserved protein [Bacillus halodurans] sp|Q9RC41 |YA57_BACHD Hypothetical protein BH1057 pir||A83782 hypothetical protein BH1057 [imported] - Bacillus halodurans (strain C-125) dbj|BAA83958.11 YHDE [Bacillus halodurans] dbj|BAB04776.11 BH1057~unknown conserved protein [Bacillus halodurans] Length = 146
1271.3 Best-BlastP=> >nrprot 61 % Identities = 35/70 (50%), Positives = 53/70 (75%) sp|P17724|GLB_TETPY Myoglobin (Hemoglobin) pir||A36270 hemoglobin - Tetrahymena pyriformis dbj|BAA03015.1 | hemoglobin [Tetrahymena pyriformis] Length = 121
1272.4 Best-BlastP=> >nrprot 46% Identities = 89/318 (27%), Positives = 147/318 (46%), Gaps = 13/318 (4%) ref|NP_253021.1 | probable ferredoxin reductase [Pseudomonas aeruginosa PA01] pir||G83104 probable ferredoxin reductase PA4331 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG07719.1 |AE004849_6 probable ferredoxin reductase [Pseudomonas aeruginosa PA01] Length = 308
1273.4 Best-BlastP=> >nrprot 40% Identities = 54/200 (27%), Positives = 95/200 (47%), Gaps = 8/200 (4%) gb|AAM88782.11 hypothetical protein [Photorhabdus luminescens] Length = 247
1275.2 Best-BlastP=> >nrprot 57% Identities = 107/291 (36%), Positives = 171/291 (58%), Gaps = 1/291 (0%) gb|AAM88781.1 | MhpE-like protein [Photorhabdus luminescens] Length = 312
1277.2 Best-BlastP=> >nrprot 48% Identities = 100/294 (34%), Positives = 162/294 (55%), Gaps = 11/294 (3%) ref|NP_624285.1 | UDP-N- acetylmuramyl tripeptide synthase [Thermoanaerobacter tengcongensis] gb|AAM25889.1 | UDP-N-acetylmuramyl tripeptide synthase [Thermoanaerobacter tengcongensis] Length = 879
1278.2 Best-BlastP=> >nrprot 43% Identities = 104/386 (26%), Positives = 176/386 (45%), Gaps = 49/386 (12%) gb|AAN83921.11 hypothetical protein [Aplysia californica] Length = 427
1279.3 Best-BlastP=> >nrprot 76% Identities = 107/173 (61 %), Positives = 136/173 (78%) ref|NP_439712.11 hypothetical protein [Haemophilus influenzae Rd] sp|P44255|YFCM_HAEIN Hypothetical protein HI1563 pir||D64036 hypothetical protein HI1563 - Haemophilus influenzae (strain Rd KW20) gb|AAC23212.11 conserved hypothetical protein [Haemophilus influenzae Rd] Length = 178
1280.3 Best-BlastP=> >nrprot 15% Identities = 110/113 (97%), Positives = 110/113 (97%) gb|AA061471.11 LidA [Legionella pneumophila] Length = 113 1282.3 Best-BlastP=> >nrprot 44% Identities = 15δ/4g3 (31 %), Positives = 230/493 (46%), Gaps = 37/493 (7%) gb|AAC35δ92.11 LphB [Legionella pneumophila] Length = 518
1283.2 Best-BlastP=> >nrprot 59% Identities = 180/429 (41 %), Positives = 260/429 (60%), Gaps = 9/429 (2%) ref |NP_716339.11 conserved hypothetical protein [Shewanella oneidensis MR-1] gb|AAN53784.1 |AE01δ516_6 conserved hypothetical protein [Shewanella oneidensis MR-1] Length = 433
1284.2 Best-BlastP=> >nrprot 45% Identities = 25/75 (33%), Positives = 43/75 (67%) ref|NP_7g6782.11 hypothetical protein VP0403 [Vibrio parahaemolyticus RIMD 2210633] dbj|BAC58666.11 hypothetical protein [Vibrio parahaemolyticus] Length = 111
1286.3 Best-BlastP=> >nrprot 19% Identities = 71/260 (27%), Positives = 123/260 (47%), Gaps = 17/260 (6%) gb|AAK19884.11 putative methoxymalonyl CoA synthase [Polyangium cellulosum] Length = 863
1287.3 Best-BlastP=> >nrprot 49% Identities = 278/933 (29%), Positives = 458/933 (49%), Gaps = 56/933 (6%) ref|ZP_00089642.11 hypothetical protein [Azotobacter vinelandii] Length = 973
1288.2 Best-BlastP=> >nrprot 14% Identities = 39/130 (30%), Positives = 67/130 (51 %), Gaps = 8/130 (6%) ref |XP_314825.1 | ENSANGP00000011098 [Anopheles gambiae] gb|EAA10144.11 ENSANGP00000011098 [Anopheles gambiae str. PEST] Length = 1842
129.3 Best-BlastP=> >nrprot No Hits found
1293.4 Best-BlastP=> >nrprot 62% Identities = 91/182 (50%), Positives = 123/182 (67%), Gaps = 2/182 (1%) ref |ZP_00013245.1 | COG2353: Uncharacterized conserved protein [Rhodospirillum rubrum] Length = 201
1294.2 Best-BlastP=> >nrprot 59% Identities = 77/177 (43%), Positives = 105/177 (59%), Gaps = 7/177 (3%) ref|NP_902948.1 | probable cytochrome b561 [Chromobacterium violaceum ATCC 12472] gb|AAQ60942.11 probable cytochrome bδ61 [Chromobacterium violaceum ATCC 12472] Length = 180
1295.1 Best-BlastP=> >nrprot 47% Identities = 64/169 (37%), Positives = 91/169 (53%), Gaps = 1/169 (0%) ref|NP_422185.1 | conserved hypothetical protein [Caulobacter crescentus CB15] pir||E87669 conserved hypothetical protein CC3391 [imported] - Caulobacter crescentus gb|AAK253δ3.11 conserved hypothetical protein [Caulobacter crescentus CB15] Length = 427
1299.3 Best-BlastP=> >nrprot 53% Identities = 150/437 (34%), Positives = 246/437 (56%), Gaps = 6/437 (1%) gb|AAD28727.1 |AF112468_6 TraH protein precursor [Salmonella typhimurium] Length = 457
13.1 Best-BlastP=> >nrprot 40% Identities = 57/260 (21%), Positives = 111/260 (42%), Gaps = 28/260 (10%) ref|NP_125735.11 hypothetical protein [Pyrococcus abyssi] pir||G75189 hypothetical protein PAB2321 - Pyrococcus abyssi (strain Orsay) emb|CAB48966.1 | Hypothetical protein [Pyrococcus abyssi] Length = 24g
130.4 Best-BlastP=> >nrprot 50% Identities = 184/664 (33%), Positives = 274/554 (49%), Gaps = 94/554 (16%) ref|NP_841634.1| Flagellar hook- associated protein 2 [Nitrosomonas europaea ATCC 19718] emb|CAD85506.11 Flagellar hook-associated protein 2 [Nitrosomonas europaea ATCC 19718] Length = 481
1302.2 Best-BlastP=> >nrprot 54% Identities = 129/348 (37%), Positives = 197/348 (66%), Gaps = 10/348 (2%) ref|ZP_00130398.11 COG2200: FOG: EAL domain [Desulfovibrio desulfuricans G20] Length = 367
1303.3 Best-BlastP=> >nrprot 80% Identities = 168/252 (66%), Positives = 207/252 (82%) ref|ZP_00125838.1| COG0024: Methionine aminopeptidase [Pseudomonas syringae pv. syringae B728a] Length = 260
1306.5 Best-BlastP=> >nrprot 98% Identities = 1033/1066 (96%), Positives = 1049/1066 (98%) gb|AAM00612.11 chemiosmotic efflux system protein A- like protein [Legionella pneumophila] Length = 1066
1307.4 Best-BlastP=> >nrprot 97% Identities = 389/418 (93%), Positives = 407/418 (97%) gb|AAM00611.11 proline/glycine betaine transporter-like protein [Legionella pneumophila] Length = 422
131.2 Best-BlastP=> >nrprot 26% Identities = 26/88 (29%), Positives = 42/88 (47%), Gaps = 15/88 (17%) ref|NP_901795.11 hypothetical protein CV2125 [Chromobacterium violaceum ATCC 12472] gb|AAQ59798.1 | hypothetical protein CV2125 [Chromobacterium violaceum ATCC 12472] Length = 202 1314.3 Best-BlastP=> >nrprot δ7% Identities = 170/474 (35%), Positives = 278/474 (58%), Gaps = 14/474 (2%) ref|ZP_00129665.11 COG1538: Outer membrane protein [Desulfovibrio desulfuricans G20] Length = 494
1317.3 Best-BlastP=> >nrprot No Hits found
1318.4 Best-BlastP=> >nrprot 64% Identities = 110/230 (47%), Positives = 162/230 (70%), Gaps = 10/230 (4%) ref|NP_819966.11 conserved hypothetical protein [Coxiella burnetii RSA 493] gb|AAO90480.11 conserved hypothetical protein [Coxiella burnetii RSA 493] Length = 414
1320.3 Best-BlastP=> >nrprot 67% Identities = 365/741 (49%), Positives = 498/741 (67%), Gaps = 11/741 (1%) ref |ZP_00010561.1| COG0068: Hydrogenase maturation factor [Rhodopseudomonas palustris] Length = 772
1322.2 Best-BlastP=> >nrprot 60% Identities = 173/452 (38%), Positives = 280/452 (61 %), Gaps = 6/452 (1 %) ref |NP_462786.11 putative MFS family tranport protein (1st mdule) [Salmonella typhimurium LT2] gb|AAL22745.11 putative MFS family tranport protein [Salmonella typhimurium LT2] Length = 47δ
1323.2 Best-BlastP=> >nrprot No Hits found
1324.4 Best-BlastP=> >nrprot 28% Identities = 76/274 (27%), Positives = 130/274 (47%), Gaps = 35/274 (12%) gb|EAA21537.11 Plasmodium falciparum CDPK2 protein [Plasmodium yoelii yoelii] Length = 565
1325.2 Best-BlastP=> >nrprot 71 % Identities = 311/589 (52%), Positives = 419/589 (71 %) ref|NP_820983.1 | arginyl-tRNA synthetase [Coxiella burnetii
RSA 493] gb|AA091497.11 arginyl-tRNA synthetase [Coxiella burnetii RSA 493] Length = 592
1328.2 Best-BlastP=> >nrprot 56% Identities = 121/344 (3δ%), Positives = 196/344 (66%), Gaps = 8/344 (2%) ref | NP_819936.1| transporter, putative [Coxiella burnetii RSA 493] gb|AAOg0450.11 transporter, putative [Coxiella burnetii RSA 4 3] Length = 376
1330.6 Best-BlastP=> >nrprot 69% Identities = 372/682 (54%), Positives = 478/682 (70%), Gaps = 4/682 (0%) ref|ZP_00092220.11 COG1200: RecG- like helicase [Azotobacter vinelandii] Length = 1006
1331.3 Best-BlastP=> >nrprot 55% Identities = 125/409 (30%), Positives = 223/409 (54%), Gaps = 18/409 (4%) ref |NP_391052.11 alternate gene name: comB, yufA [Bacillus subtilis] sp|P14203|YUXH_BACSU Hypothetical protein yuxH pir||BVBSCB competence protein ComB (yuxH) - Bacillus subtilis gb|AAA22318.1 | B competence protein emb|CAB07900.1| unknown [Bacillus subtilis] emb|CAB15162.1| yuxH [Bacillus subtilis subsp. subtilis str. 168] Length = 409
1332.3 Best-BlastP=> >nrprot 77% Identities = 308/477 (64%), Positives = 377/477 (79%) ref|NP_840694.11 Glycine cleavage system P-protein [Nitrosomonas europaea ATCC 19718] emb|CAD84521.1 | Glycine cleavage system P-protein [Nitrosomonas europaea ATCC 19718] Length = 483
1334.3 Best-BlastP=> >nrprot No Hits found
1336.2 Best-BlastP=> >nrprot No Hits found
1336.3 Best-BlastP=> >nrprot 99% Identities = 365/371 (98%), Positives = 369/371 (99%) gb|AAM00605.11 florfenicol efflux pump-like protein [Legionella pneumophila] Length = 371
1338.3 Best-BlastP=> >nrprot 76% Identities = 406/673 (60%), Positives = 510/673 (75%), Gaps = 8/673 (1%) ref|NP_837722.11 methionine tRNA synthetase [Shigella flexneri 2a str. 2457T] gb|AAP17531.11 methionine tRNA synthetase [Shigella flexneri 2a str. 2457T] Length = 677
1339.2 Best-BlastP=> >nrprot 79% Identities = 109/185 (58%), Positives = 151/185 (81%) ref |NP_770170.11 blr3530 [Bradyrhizobium japonicum] dbj|BAC48795.11 blr3530 [Bradyrhizobium japonicum USDA 110] Length = 206
1341.2 Best-BlastP=> >nrprot No Hits found
1342.3 Best-BlastP=> >nrprot 49% Identities = 73/251 (29%), Positives = 130/251 (51 %), Gaps = 16/251 (6%) gb|AAM90720.1| TraF [Salmonella typhi] Length = 259
1344.3 Best-BlastP=> >nrprot 14% Identities = 30/113 (26%), Positives = 51/113 (45%), Gaps = 15/113 (13%) ref|ZP_00101173.1| hypothetical protein [Desulfitobacterium hafniense] Length = 367
1345.3 Best-BlastP=> >nrprot 65% Identities = 173/366 (47%), Positives = 239/366 (65%), Gaps = 7/366 (1 %) ref|NP_778380.11 conserved hypothetical protein [Xylella fastidiosa Temeculal] gb|AAO28029.11 conserved hypothetical protein [Xylella fastidiosa Temeculal] Length = 381
1346.3 Best-BlastP=> >nrprot 69% Identities = 135/268 (62%), Positives = 183/268 (70%), Gaps = 5/268 (1%) ref|NP_841557.11 Uncharacterized protein family UPF0006 [Nitrosomonas europaea ATCC 19718] emb|CAD85427.11 Uncharacterized protein family UPF0006 [Nitrosomonas europaea ATCC 19718] Length = 254
1363.3 Best-BlastP=> >nrprot 43% Identities = 82/333 (24%), Positives = 152/333 (45%), Gaps = 43/333 (12%) ref|NP_819818.1 | multidrug resistance protein [Coxiella burnetii RSA 493] gb|AAO90332.11 multidrug resistance protein [Coxiella burnetii RSA 493] Length = 331
1364.2 Best-BlastP=> >nrprot No Hits found
1356.2 Best-BlastP=> >nrprot 44% Identities = 72/242 (29%), Positives = 117/242 (48%), Gaps = 8/242 (3%) ref|NP_814508.11 amino acid ABC transporter, amino acid-binding/permease protein [Enterococcus faecalis V583] gb|AAO80578.11 amino acid ABC transporter, amino acid- binding/permease protein [Enterococcus faecalis V583] Length = 722
1357.2 Best-BlastP=> >nrprot 71 % Identities = 90/146 (61%), Positives = 107/146 (73%) ref|NP_718466.1 | Yail/YqxD family protein [Shewanella oneidensis MR-1] gb|AAN55910.1 |AE015727_10 Yail/YqxD family protein [Shewanella oneidensis MR-1] Length = 151
1359.2 Best-BlastP=> >nrprot 99% Identities = 357/357 (100%), Positives = 357/357 (100%) emb|CAD43479.11 O-acetyltransferase [Legionella pneumophila] Length = 357
1360.6 Best-BlastP=> >nrprot 39% Identities = 112/443 (25%), Positives = 195/443 (44%), Gaps = 75/443 (16%) ref|NP_845547.11 conserved hypothetical protein [Bacillus anthracis str. Ames] gb|AAP27033.11 conserved hypothetical protein [Bacillus anthracis str. Ames] Length = 445
1361.3 Best-BlastP=> >nrprot 46% Identities = 99/312 (31%), Positives = 162/312 (51 %), Gaps = 25/312 (8%) ref|NP_250993.1 | hypothetical protein [Pseudomonas aeruginosa PA01] pir||A83358 hypothetical protein PA2303 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG05691.1 |AE004656_3 hypothetical protein PA2303 [Pseudomonas aeruginosa PA01] Length = 339
1362.3 Best-BlastP=> >nrprot 96% Identities = 570/607 (93%), Positives = 588/607 (96%) gb|AAK00285.1 |AF288536_7 unknown [Legionella longbeachae] Length = 607
1364.2 Best-BlastP=> >nrprot 96% Identities = 506/534 (94%), Positives = 516/534 (96%) gb|AAK00286.1 |AF288536_8 possible sensor kinase protein [Legionella longbeachae] Length = 534
1366.3 Best-BlastP=> >nrprot No Hits found
1367.3 Best-BlastP=> >nrprot 58% Identities = 55/123 (44%), Positives = 83/123 (67%) ref|ZP_00047813.11 COG2391 : Predicted transporter component [Magnetospirillum magnetotacticum] Length = 155
1368.2 Best-BlastP=> >nrprot No Hits found 1369.2 Best-BlastP=> >nrprot 63% Identities = 58/133 (43%), Positives = 92/133 (69%) ref|NP_800456.11 conserved hypothetical protein [Vibrio parahaemolyticus RIMD 2210633] dbj|BAC62289.1 | conserved hypothetical protein [Vibrio parahaemolyticus] Length = 139
1370.5 Best-BlastP=> >nrprot 56% Identities = 66/135 (48%), Positives = 87/135 (64%), Gaps = 2/135 (1 %) ref|NP_819361.11 conserved domain protein [Coxiella burnetii RSA 493] gb|AA089875.11 conserved domain protein [Coxiella burnetii RSA 493] Length = 214
1372.2 Best-BlastP=> >nrprot No Hits found
1373.1 Best-BlastP=> >nrprot No Hits found
1375.2 Best-BlastP=> >nrprot 29% Identities = 51 /102 (50%), Positives = 69/102 (67%) ref|NP_790719.11 conserved hypothetical protein [Pseudomonas syringae pv. tomato str. DC3000] gb|AA054414.11 conserved hypothetical protein [Pseudomonas syringae pv. tomato str. DC3000] Length = 249
1400.2 Best-BlastP=> >nrprot 71 % Identities = 245/484 (50%), Positives = 346/484 (71 %), Gaps = 8/484 (1 %) ref|NP_820418.11 NADH dehydrogenase I, N subunit [Coxiella burnetii RSA 493] gb|AAO90932.11 NADH dehydrogenase I, N subunit [Coxiella burnetii RSA 493] Length = 482
1402.2 Best-BlastP=> >nrprot 73% Identities = 28g/499 (57%), Positives = 367/499 (73%), Gaps = 1/499 (0%) ref|NP_820419.11 NADH dehydrogenase I, M subunit [Coxiella burnetii RSA 493] gb|AAO90933.11 NADH dehydrogenase I, M subunit [Coxiella burnetii RSA 493] Length = 506
1405.2 Best-BlastP=> >nrprot 98% Identities = 292/296 (98%), Positives = 293/296 (98%) emb|CAB43070.11 DjlA protein [Legionella pneumophila] Length = 296 1406.2 Best-BlastP=> >nrprot 97% Identities = 401/419 (95%), Positives = 408/419 (97%) emb|CAB43071.11 3-Deoxy-D-manno-oct-2-ulosonic acid transferase [Legionella pneumophila] Length = 419
1407.2 Best-BlastP=> >nrprot No Hits found
1409.3 Best-BlastP=> >nrprot 60% Identities = 102/245 (41%), Positives = 157/246 (64%), Gaps = 3/245 (1%) ref|NP_819783.11 competence lipoprotein ComL, putative [Coxiella burnetii RSA 493] gb|AAO90297.11 competence lipoprotein ComL, putative [Coxiella burnetii RSA 493] Length = 255
141.4 Best-BlastP=> >nrprot 25% Identities = 116/527 (22%), Positives = 221/527 (41 %), Gaps = 72/527 (13%) pir||T18296 myosin heavy chain - Entamoeba histolytica gb|AAB48065.11 myosin heavy chain [Entamoeba histolytica] Length = 2139
1410.2 Best-BlastP=> >nrprot 50% Identities = 94/303 (31%), Positives = 159/303 (52%), Gaps = 8/303 (2%) ref|NP_931129.11 recombinaison associated protein [Photorhabdus luminescens subsp. laumondii TT01] emb|CAE16301.1 | recombinaison associated protein [Photorhabdus luminescens subsp. laumondii TT01] Length = 303
1412.5 Best-BlastP=> >nrprot 61% Identities = 251/573 (43%), Positives = 359/673 (62%), Gaps = 17/573 (2%) ref|ZP_00110863.1 | hypothetical protein [Nostoc punctiforme] Length = 578
1413.2 Best-BlastP=> >nrprot 36% Identities = 99/349 (28%), Positives = 168/349 (48%), Gaps = 18/349 (5%) ref|NP_284204.1| putative acyl-CoA ligase [Neisseria meningitidis Z2491] pir||D81839 probable acyl-CoA ligase (EC 6.2.1.-) NMA1482 [imported] - Neisseria meningitidis (strain Z24gi serogroup A) emb|CAB84715.1 | putative acyl-CoA ligase [Neisseria meningitidis Z2491] Length = 517
1414.1 Best-BlastP=> >nrprot 52% Identities = 24/70 (34%), Positives = 41/70 (58%), Gaps = 2/70 (2%) ref|ZP_00036213.11 hypothetical protein [Enterococcus faecium] Length = 77
1415.4 Best-BlastP=> >nrprot 48% Identities = 75/253 (29%), Positives = 121/253 (47%), Gaps = 30/253 (11%) ref |ZP_00012801.1 | hypothetical protein [Rhodopseudomonas palustris] Length = 256
1417.1 Best-BlastP=> >nrprot 66% Identities = 64/129 (49%), Positives = 88/129 (68%) ref|ZP_00066647.11 hypothetical protein [Microbulbifer degradans 2-40] Length = 164
1418.3 Best-BlastP=> >nrprot 71% Identities = 295/592 (49%), Positives = 421/592 (71 %), Gaps = 2/692 (0%) ref|NP_820066.1 | transporter, putative [Coxiella burnetii RSA 493] gb|AAO90580.11 transporter, putative [Coxiella burnetii RSA 493] Length = 693
142.2 Best-BlastP=> >nrprot 62% Identities = 108/225 (48%), Positives = 144/225 (64%), Gaps = 1/225 (0%) ref|NP_465898.11 D-alanyl-D-alanine dipeptidase [Salmonella enterica subsp. enterica serovar Typhi] ref|NP_80δ294.11 D-alanyl-D-alanine dipeptidase [Salmonella enterica subsp. enterica serovar Typhi Ty 2] pir||AE0669 D-alanyl-D-alanine dipeptidase [imported] - Salmonella enterica subsp. enterica serovar Typhi (strain CT18) emb|CAD01726.11 D-alanyl-D-alanine dipeptidase [Salmonella enterica subsp. enterica serovar Typhi] gb|AA069143.11 D-alanyl-D-alanine dipeptidase [Salmonella enterica subsp. enterica serovar Typhi Ty2] Length = 256
1420.2 Best-BlastP=> >nrprot 80% Identities = 258/380 (67%), Positives = 309/380 (81 %) ref|NP_820998.11 cystathionine beta-lyase [Coxiella burnetii RSA 493] gb|AA091512.11 cystathionine beta-lyase [Coxiella burnetii RSA 493] Length = 387
1421.3 Best-BlastP=> >nrprot 55% Identities = 280/736 (38%), Positives = 432/736 (58%), Gaps = 19/736 (2%) ref|NP_900703.11 conserved hypothetical protein [Chromobacterium violaceum ATCC 12472] gb|AAQ58708.1 | conserved hypothetical protein [Chromobacterium violaceum ATCC 12472] Length = 936
1423.2 Best-BlastP=> >nrprot 66% Identities = 226/450 (50%), Positives = 315/450 (70%), Gaps = 2/450 (0%) pir||JC5798 F0F1 -ATPase (EC 3.4.-.-) beta chain - Methanosarcina barkeri gb|AAC38049.11 ATP synthase beta subunit [Methanosarcina barkeri] Length = 469
1424.2 Best-BlastP=> >nrprot 37% Identities = 27/106 (25%), Positives = 51/106 (48%) ref|ZP_00090544.11 COG0355: F0F1-type ATP synthase, epsilon subunit (mitochondrial delta subunit) [Azotobacter vinelandii] Length = 178
1425.2 Best-BlastP=> >nrprot 50% Identities = 25/88 (28%), Positives = 47/88 (53%), Gaps = 2/88 (2%) ref |NP_661922.11 ATP synthase, putative [Chlorobium tepidum TLS] gb|AAM72264.11 ATP synthase, putative [Chlorobium tepidum TLS] Length = 107
142g.4 Best-BlastP=> >nrprot 26% Identities = 68/329 (20%), Positives = 131/329 (39%), Gaps = 32/329 (9%) emb|CAD27470.11 SPAPB18E9.04c [Schizosaccharomyces pombe] Length = 800
1432.3 Best-BlastP=> >nrprot 52% Identities = 105/294 (35%), Positives = 160/294 (54%), Gaps = 6/294 (2%) ref|NP_745744.11 transcriptional regulator, LysR family [Pseudomonas putida KT2440] gb|AAN69208.1 |AE0165δδ_6 transcriptional regulator, LysR family [Pseudomonas putida KT2440] Length = 309
1434.3 Best-BlastP=> >nrprot 52% Identities = 242/765 (31 %), Positives = 388/765 (50%), Gaps = 49/765 (6%) ref|NP_842403.11 DNA internalization- related competence protein ComEC/Rec2 [Nitrosomonas europaea ATCC 19718] emb|CAD86320.1 | DNA internalization-related competence protein ComEC/Rec2 [Nitrosomonas europaea ATCC 19718] Length = 799
1438.4 Best-BlastP=> >nrprot 80% Identities = 428/631 (67%), Positives = 504/631 (79%), Gaps = 6/631 (0%) ref|NP_245307.11 ParE [Pasteurella multocida] gb|AAK02454.1 | ParE [Pasteurella multocida] Length = 632
144.3 Best-BlastP=> >nrprot 8% Identities = 21/50 (42%), Positives = 31/50 (62%), Gaps = 1/50 (2%) ref|NP_618108.11 cysteine protease (papain C1 family) [Methanosarcina acetivorans str. C2A] gb|AAM06588.1 | cysteine protease (papain C1 family) [Methanosarcina acetivorans str. C2A] Length = 340
1440.2 Best-BlastP=> >nrprot 56% Identities = 163/398 (40%), Positives = 231/398 (58%), Gaps = 45/398 (11%) ref|NP_820093.11 efflux transporter, RND family, MFP subunit [Coxiella burnetii RSA 493] gb|AAO90607.11 efflux transporter, RND family, MFP subunit [Coxiella burnetii RSA 493] Length = 380
1444.3 Best-BlastP=> >nrprot 46% Identities = 189/523 (36%), Positives = 282/523 (53%), Gaps = 48/523 (9%) ref|NP_775139.11 sphingosine-1 - phosphate lyase 1 [Rattus norvegicus] emb|CAD55407.1 | sphingosine-1 -phosphate lyase [Rattus norvegicus] Length = 568
1446.3 Best-BlastP=> >nrprot 71% Identities = 143/246 (68%), Positives = 181/246 (73%) ref|NP_793532.11 conserved hypothetical protein [Pseudomonas syringae pv. tomato str. DC3000] gb|AA067227.1 | conserved hypothetical protein [Pseudomonas syringae pv. tomato str. DC3000] Length = 249
1448.2 Best-BlastP=> >nrprot 50% Identities = 109/334 (32%), Positives = 172/334 (51 %), Gaps = 25/334 (7%) ref|NP_106951.1 | similar to oxidoreductase [Mesorhizobium loti] dbj|BAB62737.11 similar to oxidoreductase [Mesorhizobium loti] Length = 677
1449.3 Best-BlastP=> >nrprot 62% Identities = 209/736 (28%), Positives = 369/736 (50%), Gaps = 69/735 (9%) ref|ZP_00042382.11 COG2114: Adenylate cyclase, family 3 (some proteins contain HAMP domain) [Magnetococcus sp. MC-1] Length = 726
1450.2 Best-BlastP=> >nrprot 57% Identities = 168/388 (43%), Positives = 244/388 (62%), Gaps = 8/388 (2%) ref|NP_407479.11 multidrug translocase [Yersinia pestis] ref|NP_671360.11 proton motive force efflux pump protein [Yersinia pestis KIM] pir||AD0492 multidrug translocase [imported] - Yersinia pestis (strain C092) emb|CAC93504.11 multidrug translocase [Yersinia pestis C092] gb|AAM87611.1 |AE014008_5 proton motive force efflux pump protein [Yersinia pestis KIM] Length = 409
1451.4 Best-BlastP=> >nrprot 65% Identities = 188/426 (44%), Positives = 278/426 (65%), Gaps = 4/426 (0%) ref|ZP_00034777.11 COG2252: Permeases [Burkholderia fungorum] Length = 433
1454.2 Best-BlastP=> >nrprot 88% Identities = 500/638 (78%), Positives = 568/638 (89%) ref |NP_901459.11 protein kinase [Chromobacterium violaceum ATCC 12472] gb|AAQ59463.11 protein kinase [Chromobacterium violaceum ATCC 12472] Length = 642
1456.2 Best-BlastP=> >nrprot 47% Identities = 52/15δ (33%), Positives = 81/165 (52%), Gaps = 10/165 (6%) ref|NP_253150.11 conserved hypothetical protein [Pseudomonas aeruginosa PA01] pir||D83087 conserved hypothetical protein PA4460 [imported] - Pseudomonas aeruginosa (strain PA01) gb|AAG07848.1 |AE004860_4 conserved hypothetical protein [Pseudomonas aeruginosa PA01] Length = 175
1456.2 Best-BlastP=> >nrprot 80% Identities = 165/238 (69%), Positives = 194/238 (81%) ref |NP_719490.11 ABC transporter, ATP-binding protein [Shewanella oneidensis MR-1] gb|AAN56934.1 |AE015827_6 ABC transporter, ATP-binding protein [Shewanella oneidensis MR-1] Length = 243
1457.2 Best-BlastP=> >nrprot 73% Identities = 105/186 (66%), Positives = 141/185 (76%), Gaps = 1/185 (0%) ref|ZP_00079751.11 COG0512: Anthranilate/para-aminobenzoate synthases component II [Geobacter metallireducens] Length = 189
1461.3 Best-BlastP=> >nrprot 68% Identities = 167/315 (53%), Positives = 214/315 (67%), Gaps = 20/315 (6%) ref|NP_819187.1 | UDP-N- acetylenolpyruvoylglucosamine reductase [Coxiella burnetii RSA 493] gb|AAO89701.11 UDP-N-acetylenolpyruvoylglucosamine reductase [Coxiella burnetii RSA 493] Length = 316
1463.2 Best-BlastP=> >nrprot 66% Identities = 170/375 (45%), Positives = 241/375 (64%), Gaps = 16/375 (4%) ref|NP_773078.11 D-alanine-D-alanine ligase A [Bradyrhizobium japonicum] dbj|BAC51703.11 D-alanine--D-alanine ligase A [Bradyrhizobium japonicum USDA 110] Length = 373
1466.2 Best-BlastP=> >nrprot 71 % Identities = 94/205 (45%), Positives = 152/205 (74%) ref|NP_841948.1 | Prokaryotic-type carbonic anhydrase [Nitrosomonas europaea ATCC 19718] emb|CAD85837.11 Prokaryotic-type carbonic anhydrase [Nitrosomonas europaea ATCC 19718] Length = 208
1467.5 Best-BlastP=> >nrprot 65% Identities = 90/186 (48%), Positives = 126/186 (67%), Gaps = 2/186 (1 %) ref|NP_691260.11 hypothetical protein [Oceanobacillus iheyensis HTE831] dbj|BAC12295.11 hypothetical conserved protein [Oceanobacillus iheyensis HTE831] Length = 189
1469.4 Best-BlastP=> >nrprot 20% Identities = 112/493 (22%), Positives = 217/493 (44%), Gaps = 44/493 (8%) ref|NP_563924.11 nuclear matrix constituent protein -related [Arabidopsis thaliana] pir||G86266 hypothetical protein F3F19.25 - Arabidopsis thaliana gb|AAD31075.1 |AC007357_24 Similar to gb|D64087 nuclear matrix constituent protein 1 (NMCP1 ) from Daucus carota. [Arabidopsis thaliana] Length = 1128
1473.4 Best-BlastP=> >nrprot 49% Identities = 44/143 (30%), Positives = 75/143 (52%), Gaps = 4/143 (2%) ref |NP_810648.11 chromate transport protein [Bacteroides thetaiotaomicron VPI-5482] gb|AA076842.11 chromate transport protein [Bacteroides thetaiotaomicron VPI-5482] Length = 185 1474.4 Best-BlastP=> >nrprot 52% Identities = 125/314 (39%), Positives = 194/314 (61 %), Gaps = 4/314 (1 %) ref |NP_103760.1 | unknown protein [Mesorhizobium loti] dbj|BAB49546.1 | unknown protein [Mesorhizobium loti] Length = 431
1475.2 Best-BlastP=> >nrprot No Hits found
1476.2 Best-BlastP=> >nrprot 81 % Identities = 185/302 (61 %), Positives = 246/302 (81 %), Gaps = 1/302 (0%) ref|NP_762779.1 | Glutathione synthase; Ribosomal protein S6 modification enzyme [Vibrio vulnificus CMCP6] gb|AAO07769.1 |AE016811_10 Glutathione synthase; Ribosomal protein S6 modification enzyme [Vibrio vulnificus CMCP6] dbj|BAC97337.1 | ribosomal protein S6 modification protein [Vibrio vulnificus YJ016] Length = 301
1479.3 Best-BlastP=> >nrprot 69% Identities = 223/407 (54%), Positives = 287/407 (70%), Gaps = 1/407 (0%) sp|Q8GDU5|ARLY_HELMO Argininosuccinate lyase (Arginosuccinase) (ASAL) gb|AAN87483.11 Argininosuccinate lyase [Heliobacillus mobilis] Length = 458
148.2 Best-BlastP=> >nrprot 55% Identities = 186/450 (41 %), Positives = 279/450 (62%), Gaps = 14/450 (3%) ref|ZP_00060071.11 COG2200: FOG: EAL domain [Clostridium thermocellum ATCC 27405] Length = 862
1481.3 Best-BlastP=> >nrprot 51% Identities = 128/304 (42%), Positives = 191/304 (62%), Gaps = 9/304 (2%) ref |NP_126998.1 | ornithine carbamoyltransferase [Pyrococcus abyssi] sp|093656|OTC_PYRAB Ornithine carbamoyltransferase (OTCase) pir||G75041 ornithine carbamoyltransferase (argf) PAB1502 - Pyrococcus abyssi (strain Orsay) emb|CAB50228.11 argF ornithine carbamoyltransferase [Pyrococcus abyssi] Length = 317
1482.4 Best-BlastP=> >nrprot 22% Identities = 40/150 (26%), Positives = 71/150 (47%) ref |NP_819099.11 hypothetical protein [Coxiella burnetii RSA 493] gb|AA089613.1 | hypothetical protein [Coxiella burnetii RSA 493] Length = 250
1484.2 Best-BlastP=> >nrprot 72% Identities = 126/223 (56%), Positives = 164/223 (73%) ref|ZP_00053873.11 COG1 136: ABC-type antimicrobial peptide transport system, ATPase component [Magnetospirillum magnetotacticum] Length = 225
1485.4 Best-BlastP=> >nrprot 64% Identities = 171/414 (41 %), Positives = 268/414 (64%), Gaps = 4/414 (0%) ref|NP_436895.1 | CONSERVED HYPOTHETICAL PROTEIN [Sinorhizobium meliloti] pir||C95886 conserved hypothetical protein [imported] - Sinorhizobium meliloti (strain 1021) magaplasmid pSymB emb|CAC48755.1 | CONSERVED HYPOTHETICAL PROTEIN [Sinorhizobium meliloti] Length = 413
1487.3 Best-BlastP=> >nrprot 99% Identities = 266/268 (99%), Positives = 268/268 (100%) emb|CAB65183.11 enoyl reductase [Legionella pneumophila] Length = 268
1488.2 Best-BlastP=> >nrprot No Hits found
14g0.4 Best-BlastP=> >nrprot 72% Identities = 536/927 (57%), Positives = 679/927 (73%), Gaps = 9/927 (0%) ref |NP_819435.11 isoleucyl-tRNA synthetase [Coxiella burnetii RSA 493] gb|AA089949.11 isoleucyl-tRNA synthetase [Coxiella burnetii RSA 493] Length = 936
1493.6 Best-BlastP=> >nrprot 69% Identities = 822/1621 (50%), Positives = 1137/1621 (70%), Gaps = 3/1621 (0%) ref|NP_820221.11 conserved hypothetical protein [Coxiella burnetii RSA 493] gb|AAO90735.11 conserved hypothetical protein [Coxiella burnetii RSA 493] Length = 1619
1494.2 Best-BlastP=> >nrprot 33% Identities = 58/170 (34%), Positives = 89/170 (52%), Gaps = 1/170 (0%) ref|ZP_00081963.1 | COG0204: 1-acyl-sn- glycerol-3-phosphate acyltransferase [Geobacter metallireducens] Length = 234
1495.1 Best-BlastP=> >nrprot 72% Identities = 60/104 (57%), Positives = 78/104 (75%) ref|NP_458777.11 SugE protein [Salmonella enterica subsp. enterica serovar Typhi] ref|NP_463202.11 putative DMT superfamily transport protein [Salmonella typhimurium LT2] ref|NP_807981.11 SugE protein [Salmonella enterica subsp. enterica serovar Typhi Ty2] pir||AE1046 SugE protein [imported] - Salmonella enterica subsp. enterica serovar Typhi (strain CT18) gb|AAL23161.11 putative DMT superfamily transport protein [Salmonella typhimurium LT2] emb|CAD06818.1 | SugE protein [Salmonella enterica subsp. enterica serovar Typhi] gb|AA071841.1 | SugE protein [Salmonella enterica subsp. enterica serovar Typhi Ty2] Length = 105
1497.2 Best-BlastP=> >nrprot 66% Identities = 189/335 (56%), Positives = 236/335 (70%) ref|ZP_00125893.11 COG0389: Nucleotidyltransferase/DNA polymerase involved in DNA repair [Pseudomonas syringae pv. syringae B728a] Length = 354
15.1 Best-BlastP=> >nrprot 63% Identities = 181/393 (46%), Positives = 248/393 (63%), Gaps = 20/393 (5%) ref|NP_890078.11 phage integrase [Bordetella bronchiseptica] emb|CAE34037.11 phage integrase [Bordetella bronchiseptica] Length = 407
160.2 Best-BlastP=> >nrprot 52% Identities = 130/266 (48%), Positives = 172/266 (64%), Gaps = 8/266 (3%) ref|ZP_00029131.11 COG3243: Poly(3- hydroxyalkanoate) synthetase [Burkholderia fungorum] Length = 642
1501.3 Best-BlastP=> >nrprot 72% Identities = 139/276 (50%), Positives = 200/276 (72%), Gaps = 2/276 (0%) ref|NP_519387.11 PROBABLE SN- GLYCEROL-3-PHOSPHATE TRANSMEMBRANE ABC TRANSPORTER PROTEIN [Ralstonia solanacearum] emb|CAD14968.11 PROBABLE SN-GLYCEROL-3-PHOSPHATE TRANSMEMBRANE ABC TRANSPORTER PROTEIN [Ralstonia solanacearum] Length = 282
1503.4 Best-BlastP=> >nrprot 65% Identities = 183/364 (50%), Positives = 239/364 (65%), Gaps = 15/364 (4%) ref|NP_407240.11 sn-glycerol-3- phosphate transport, ATP-binding protein [Yersinia pestis] pir||AH0461 sn-glycerol-3-phosphate transport, ATP-binding protein ugpC [imported] - Yersinia pestis (strain C092) emb|CAC93260.11 sn-glycerol-3-phosphate transport, ATP-binding protein [Yersinia pestis C092] Length = 357
1505.4 Best-BlastP=> >nrprot No Hits found
1507.3 Best-BlastP=> >nrprot 64% Identities = 226/496 (45%), Positives = 318/496 (64%), Gaps = 4/496 (0%) ref|NP_231057.11 thermostable carboxypeptidase 1 [Vibrio cholerae 01 biovar eltor str. N16961] pir||B82202 thermostable carboxypeptidase 1 VC1414 [imported] - Vibrio cholerae (strain N16961 serogroup 01 ) gb|AAF94571.1 | thermostable carboxypeptidase 1 [Vibrio cholerae 01 biovar eltor str. N16961] Length = 524
1508.4 Best-BlastP=> >nrprot 38% Identities = 85/279 (30%), Positives = 141/279 (50%), Gaps = 9/279 (3%) ref|NP_622993.11 predicted nucleotide- utilizing enzyme related to molybdopterin-biosynthesis enzyme MoeA [Thermoanaerobacter tengcongensis] gb|AAM24597.1 | predicted nucleotide-utilizing enzyme related to molybdopterin-biosynthesis enzyme MoeA [Thermoanaerobacter tengcongensis] Length = 412
161.2 Best-BlastP=> >nrprot 70% Identities = 128/248 (61%), Positives = 176/248 (70%), Gaps = 4/248 (1 %) ref|NP_902034.1 | acetoacetyl-CoA reductase [Chromobacterium violaceum ATCC 12472] gb|AAQ60036.11 acetoacetyl-CoA reductase [Chromobacterium violaceum ATCC 12472] Length = 246
1510.3 Best-BlastP=> >nrprot 64% Identities = 136/259 (52%), Positives = 177/259 (68%), Gaps = 1/259 (0%) ref|ZP_00083725.1 | COG3186: Phenylalanine-4-hydroxylase [Pseudomonas fluorescens PfO-1] Length = 263
1511.3 Best-BlastP=> >nrprot 99% Identities = 217/219 (99%), Positives = 218/219 (99%) gb|AAL79360.11 GacA regulatory protein [Legionella pneumophila] Length = 219
1513.4 Best-BlastP=> >nrprot 61 % Identities = 181/448 (40%), Positives = 276/448 (61 %), Gaps = 20/448 (4%) ref|NP_820336.11 amino acid antiporter [Coxiella burnetii RSA 493] gb|AAO90850.11 amino acid antiporter [Coxiella burnetii RSA 493] Length = 474
1515.2 Best-BlastP=> >nrprot 73% Identities = 127/255 (49%), Positives = 188/255 (73%), Gaps = 1/255 (0%) ref|NP_249886.11 hypothetical protein [Pseudomonas aeruginosa PA01] pir||F83497 hypothetical protein PA1195 [imported] - Pseudomonas aeruginosa (strain PA01) gb|AAG04584.1 |AE004549_11 hypothetical protein PA1195 [Pseudomonas aeruginosa PA01] Length = 254
1516.3 Best-BlastP=> >nrprot 39% Identities = 69/269 (26%), Positives = 123/269 (45%), Gaps = 14/269 (5%) ref|NP_826905.1 | hypothetical protein [Streptomyces avermitilis MA-4680] dbj|BAC73440.11 hypothetical protein [Streptomyces avermitilis MA-4680] Length = 290
1517.3 Best-BlastP=> >nrprot 58% Identities = 164/368 (44%), Positives = 244/368 (66%), Gaps = 1/368 (0%) ref|ZP_00034488.1 | COG0156: 7-keto-8- aminopelargonate synthetase and related enzymes [Burkholderia fungorum] Length = 429
1519.2 Best-BlastP=> >nrprot 53% Identities = 126/333 (37%), Positives = 184/333 (55%), Gaps = 4/333 (1 %) ref|ZP_00034487.11 COG0784: FOG: CheY-like receiver [Burkholderia fungorum] Length = 333
152.3 Best-BlastP=> >nrprot 50% Identities = 33/120 (27%), Positives = 60/120 (50%), Gaps = 8/120 (6%) ref|NP_459344.11 putative outer membrane lipoprotein [Salmonella typhimurium LT2] gb|AAL19303.11 putative outer membrane lipoprotein [Salmonella typhimurium LT2] Length = 119
1521.2 Best-BlastP=> >nrprot 56% Identities = 69/170 (40%), Positives = 107/170 (62%) ref|NP_541215.1 | HDED PROTEIN [Brucella melitensis] ref|NP_700223.1 | conserved hypothetical protein [Brucella suis 1330] pir||AD3539 hdeD protein [imported] - Brucella melitensis (strain 16M) gb|AAL53479.1 | HDED PROTEIN [Brucella melitensis 16M] gb|AAN34228.1 |AE014598_9 conserved hypothetical protein [Brucella suis 1330] Length = 187
1522.2 Best-BlastP=> >nrprot 32% Identities = 77/231 (33%), Positives = 127/231 (54%), Gaps = 1/231 (0%) ref |NP_691224.11 hypothetical protein [Oceanobacillus iheyensis HTE831] dbj|BAC12259.1 | hypothetical conserved protein [Oceanobacillus iheyensis HTE831] Length = 243
1523.2 Best-BlastP=> >nrprot No Hits found
1524.3 Best-BlastP=> >nrprot 21 % Identities = 48/129 (37%), Positives = 76/129 (58%), Gaps = 4/129 (3%) ref|NP_561262.11 probable transcriptional regulator [Clostridium perfringens] dbj|BAB80052.1 [probable transcriptional regulator [Clostridium perfringens str. 13] Length = 249
1525.3 Best-BlastP=> >nrprot 51 % Identities = 137/384 (35%), Positives = 217/384 (56%), Gaps = 24/384 (6%) gb|AAM73854.1 |AF454865_1 putative phospholipase C [Legionella pneumophila] Length = 423
1528.4 Best-BlastP=> >nrprot 65% Identities = 214/446 (47%), Positives = 291/446 (65%), Gaps = 6/446 (1 %) sp|Q8PMU6|RUMA_XANAC 23S rRNA (Uracil-δ-)-methyltransferase ru A (23S rRNA(M-5-U1939)-methyitransferase) Length = 444
1529.2 Best-BlastP=> >nrprot 77% Identities = 263/394 (64%), Positives = 309/394 (78%) ref|NP_416880.11 putative aminotransferase [Escherichia coli K12] sp|P77434|YFDZ_ECOLI Hypothetical aminotransferase yfdZ pir||H65011 probable transaminase (EC 2.6.1.-) b2379 [similarity] - . Escherichia coli (strain K-12) gb|AAC75438.1 | putative aminotransferase [Escherichia coli K12] dbj|BAA16249.11 PROBABLE ASPARTATE AMINOTRANSFERASE (EC 2.6.1.1 ) (TRANSAMINASE A) (ASPAT). [Escherichia coli] Length = 412
153.2 Best-BlastP=> >nrprot 99% Identities = 436/437 (99%), Positives = 437/437 (100%) gb|AAM00622.1| chemiosmotic efflux system C protein C [Legionella pneumophila] Length = 445
1630.2 Best-BlastP=> >nrprot 52% Identities = 38/108 (35%), Positives = 64/108 (59%) ref|NP_819878.1 | conserved hypothetical protein [Coxiella burnetii RSA 493] gb|AAO90392.1| conserved hypothetical protein [Coxiella burnetii RSA 493] Length = 123
1531.3 Best-BlastP=> >nrprot 71 % Identities = 360/746 (48%), Positives = 523/746 (70%), Gaps = 17/746 (2%) ref|ZP_00126361.11 COG0317: Guanosine polyphosphate pyrophosphohydrolases/synthetases [Pseudomonas syringae pv. syringae B728a] Length = 747
1532.3 Best-BlastP=> >nrprot 52% Identities = 217/688 (31 %), Positives = 341/688 (49%), Gaps = 49/688 (7%) ref|NP_249777.11 flagellar hook- associated protein 1 FlgK [Pseudomonas aeruginosa PA01] pir||D83511 flagellar hook-associated protein 1 FlgK PA1086 [imported] - Pseudomonas aeruginosa (strain PA01) gb|AAG04475.1|AE004539_17 flagellar hook-associated protein 1 FlgK [Pseudomonas aeruginosa PA01] Length = 683
1533.1 Best-BlastP=> >nrprot 52% Identities = 120/413 (29%), Positives = 218/413 (52%), Gaps = 12/413 (2%) ref|NP_718794.11 flagellar hook- associated protein FlgL [Shewanella oneidensis MR-1] gb|AAN56238.1 |AE016761_8 flagellar hook-associated protein FlgL [Shewanella oneidensis MR-1] Length = 403
1536.3 Best-BlastP=> >nrprot 5g% Identities = 259/616 (42%), Positives = 374/616 (60%), Gaps = 9/616 (1 %) ref|NP_900243.11 potassium uptake protein [Chromobacterium violaceum ATCC 12472] gb|AAQ58249.1 | potassium uptake protein [Chromobacterium violaceum ATCC 12472] Length = 621
1539.2 Best-BlastP=> >nrprot No Hits found
164.1 Best-BlastP=> >nrprot 98% Identities = 312/322 (96%), Positives = 317/322 (98%) gb|AAM00621.1 | chemiosmotic efflux system C protein B [Legionella pneumophila] Length = 322
1542.2 Best-BlastP=> >nrprot 63% Identities = 161/342 (47%), Positives = 225/342 (65%), Gaps = 5/342 (1%) ref|NP_820833.1 | enoyl-CoA hydratase/isomerase family protein [Coxiella burnetii RSA 493] gb|AA091347.11 enoyl-CoA hydratase/isomerase family protein [Coxiella burnetii RSA 493] Length = 366
1546.3 Best-BlastP=> >nrprot 74% Identities = 319/546 (58%), Positives = 411/546 (75%), Gaps = 7/546 (1 %) ref|NP_745048.11 glutaminyl-tRNA synthetase [Pseudomonas putida KT2440] gb|AAN68512.1 |AE016483_3 glutaminyl-tRNA synthetase [Pseudomonas putida KT2440]
Length = 567 1547.2 Best-BlastP=> >nrprot 66% Identities = 132/266 (49%), Positives = 182/266 (68%), Gaps = 2/266 (0%) ref|NP_520102.11 PROBABLE TRYPTOPHAN SYNTHASE (ALPHA CHAIN) PROTEIN [Ralstonia solanacearum] emb|CAD15683.11 PROBABLE TRYPTOPHAN SYNTHASE (ALPHA CHAIN) PROTEIN [Ralstonia solanacearum] Length = 265
1648.2 Best-BlastP=> >nrprot 65% Identities = 83/198 (41 %), Positives = 129/198 (65%), Gaps = 1/198 (0%) ref|NP_866390.1 | ATP synthase a subunit [Pirellula sp.] emb|CAD78171.11 ATP synthase a subunit [Pirellula sp.] Length = 228
154g.1 Best-BlastP=> >nrprot 66% Identities = 46/73 (63%), Positives = 61/73 (83%) ref|ZP_00034461.11 COG0636: F0F1-type ATP synthase, subunit c/Archaeal/vacuolar-type H+-ATPase, subunit K [Burkholderia fungorum] Length = 82
1550.2 Best-BlastP=> >nrprot 50% Identities = 63/254 (24%), Positives = 125/264 (49%), Gaps = 10/254 (3%) ref|NP_617341.11 H(+)-transporting ATP synthase, subunit B [Methanosarcina acetivorans str. C2A] gb|AAM05821.11 H(+)-transporting ATP synthase, subunit B [Methanosarcina acetivorans str. C2A] Length = 329
1562.3 Best-BlastP=> >nrprot 87% Identities = 381/511 (74%), Positives = 462/511 (88%), Gaps = 1/511 (0%) ref|ZP_00065462.11 COG0056: FOFI- type ATP synthase, alpha subunit [Microbulbifer degradans 2-40] Length = 613
1δδδ.2 Best-BlastP=> >nrprot 79% Identities = 189/288 (65%), Positives = 231/288 (80%), Gaps = 2/288 (0%) ref|NP_254242.11 ATP synthase gamma chain [Pseudomonas aeruginosa PA01] pir||D82952 ATP synthase gamma chain PAδδδδ [imported] - Pseudomonas aeruginosa (strain PA01) gb|AAG08940.1 |AE004967_11 ATP synthase gamma chain [Pseudomonas aeruginosa PA01] Length = 286
1566.2 Best-BlastP=> >nrprot 92% Identities = 395/458 (86%), Positives = 423/458 (92%), Gaps = 1/468 (0%) pir||D64071 H+-transporting two-sector ATPase (EC 3.6.3.14) beta chain - Haemophilus influenzae (strain Rd KW20) Length = 468
1567.3 Best-BlastP=> >nrprot 90% Identities = 227/291 (78%), Positives = 265/291 (91 %) ref |NP_820381.11 succinyl-CoA synthetase, alpha subunit [Coxiella burnetii RSA 493] sp|P53591 |SUCD_COXBU Succinyl-CoA synthetase alpha chain (SCS-alpha) gb|AAO90895.11 succinyl-CoA synthetase, alpha subunit [Coxiella burnetii RSA 493] Length = 294
1559.2 Best-BlastP=> >nrprot 80% Identities = 269/384 (70%), Positives = 314/384 (81%) ref|NP_250279.11 succinyl-CoA synthetase beta chain [Pseudomonas aeruginosa PA01] sp|P53593|SUCC_PSEAE Succinyl-CoA synthetase beta chain (SCS-beta) pir||A83446 succinyl-CoA synthetase beta chain PA1588 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG04977.1 |AE004587_1 succinyl-CoA synthetase beta chain [Pseudomonas aeruginosa PA01] Length = 388
1660.2 Best-BlastP=> >nrprot 59% Identities = 115/277 (41 %), Positives = 166/277 (59%), Gaps = 11/277 (3%) ref|NP_442598.11 PleD gene product homologue [Synechocystis sp. PCC 6803] pir||S76977 pleD-4 protein - Synechocystis sp. (strain PCC 6803) dbj|BAA10669.1 | slr0302 [Synechocystis sp. PCC 6803] Length = 768
1562.3 Best-BlastP=> >nrprot 34% Identities = 51/213 (23%), Positives = 94/213 (44%), Gaps = 27/213 (12%) ref|NP_705294.1 | hypothetical protein [Plasmodium falciparum 3D7] emb|CAD52531.1 | hypothetical protein [Plasmodium falciparum 3D7] Length = 1936
1563.3 Best-BlastP=> >nrprot 64% Identities = 95/200 (47%), Positives = 134/200 (67%), Gaps = 1/200 (0%) ref|ZP_00087981.1 | COG1335: Amidases related to nicotinamidase [Pseudomonas fluorescens PfO-1] Length = 208
1564.3 Best-BlastP=> >nrprot 81 % Identities = 321/467 (68%), Positives = 380/467 (81 %) ref|NP_820039.11 nicotinate phosphoribosyltransferase, putative [Coxiella burnetii RSA 493] gb|AAO90553.11 nicotinate phosphoribosyltransferase, putative [Coxiella burnetii RSA 493] Length = 468
1666.3 Best-BlastP=> >nrprot 26% Identities = 34/120 (28%), Positives = 47/120 (39%), Gaps = 18/120 (15%) gb|AAQ23913.11 metallothionein HE [Crassostrea virginica] Length = 149
1567.2 Best-BlastP=> >nrprot 28% Identities = 58/293 (19%), Positives = 127/293 (43%), Gaps = 21/293 (7%) pir||T14867 interaptin - slime mold (Dictyostelium discoideum) gb|AAC34582.1 | interaptin [Dictyostelium discoideum] Length = 1738
1569.6 Best-BlastP=> >nrprot 62% Identities = 310/738 (42%), Positives = 456/738 (61 %), Gaps = 24/738 (3%) ref|NP_796632.11 primosomal protein N' [Vibrio parahaemolyticus RIMD 2210633] dbj|BAC58516.11 primosomal protein N' [Vibrio parahaemolyticus] Length = 734
157.2 Best-BlastP=> >nrprot 90% Identities = 886/1061 (83%), Positives = 973/1061 (91 %) gb|AAM00612.1 | chemiosmotic efflux system protein A-like protein [Legionella pneumophila] Length = 1066
1570.4 Best-BlastP=> >nrprot 74% Identities = 612/1047 (58%), Positives = 788/1047 (75%), Gaps = 9/1047 (0%) ref|ZP_00034374.1 | COG0841 : Cation/multidrug efflux pump [Burkholderia fungorum] Length = 1098
1575.2 Best-BlastP=> >nrprot 50% Identities = 46/122 (37%), Positives = 71/122 (58%), Gaps = 7/122 (5%) ref|NP_925923.11 MarR family transcriptional regulatory protein [Gloeobacter violaceus] dbj|BAC90918.11 MarR family transcriptional regulatory protein [Gloeobacter violaceus] Length = 143
1577.5 Best-BlastP=> >nrprot 63% Identities = 425/983 (43%), Positives = 617/983 (62%), Gaps = 36/983 (3%) ref|NP_288121.11 putative oxidase [Escherichia coli 01δ7:H7 EDL933] ref|NP_310421.11 putative oxidase [Escherichia coli 0157:H7] pir||B90928 probable oxidase [imported] - Escherichia coli (strain 01 δ7:H7, substrain RIMD 0609952) pir||F85776 probable oxidase ydiJ [imported] - Escherichia coli (strain 0157:H7, substrain EDL933) gb|AAG66674.1 |AE005391_11 putative oxidase [Escherichia coli 0167:H7 EDL933] dbj|BAB35817.1 | putative oxidase [Escherichia coli 0167:H7] Length = 1018
1678.3 Best-BlastP=> >nrprot 67% Identities = 135/246 (54%), Positives = 173/246 (70%), Gaps = 3/246 (1 %) ref|NP_436290.11 Hypothetical protein [Sinorhizobium meliloti] pir||D95392 protein [imported] - Sinorhizobium meliloti (strain 1021 ) magaplasmid pSymA gb|AAK65702.1 | Hypothetical protein [Sinorhizobium meliloti] Length = 408
1579.4 Best-BlastP=> >nrprot 44% Identities = 36/124 (29%), Positives = 68/124 (54%), Gaps = 5/124 (4%) ref|NP_635420.11 conserved hypothetical protein [Xanthomonas campestris pv. campestris str. ATCC 33913] gb|AAM39344.1 | conserved hypothetical protein [Xanthomonas campestris pv. campestris str. ATCC 33913] Length = 143
158.2 Best-BlastP=> >nrprot 74% Identities = 139/205 (67%), Positives = 165/206 (80%), Gaps = 1/205 (0%) ref|NP_478237.1 | ORF_ID:all7590~unknown protein [Nostoc sp. PCC 7120] pir||AC2538 hypothetical protein all7590 [imported] - Nostoc sp. (strain PCC 7120) plasmid pCC7120beta dbj|BAB77233.11 ORF_ID:all7590~unknown protein [Nostoc sp. PCC 7120] Length = 217
1581.4 Best-BlastP=> >nrprot No Hits found
1582.2 Best-BlastP=> >nrprot 76% Identities = 99/138 (71 %), Positives = 115/138 (83%), Gaps = 2/138 (1 %) ref|NP_233004.11 peptide methionine sulfoxide reductase [Vibrio cholerae 01 biovar eltor str. N16961] pir||C82439 peptide methionine sulfoxide reductase VCA0615 [imported] - Vibrio cholerae (strain N16961 serogroup 01 ) gb|AAF96616.1 | peptide methionine sulfoxide reductase [Vibrio cholerae 01 biovar eltor str. N 16961] Length = 394
1584.3 Best-BlastP=> >nrprot 79% Identities = 574/901 (63%), Positives = 711/901 (78%), Gaps = 7/901 (0%) ref|NP_285794.11 preprotein translocase; secretion protein [Escherichia coli 0167:H7 EDL933] ref |NP_308129.11 preprotein translocase SecA [Escherichia coli 0167:H7] ref|NP_7δ2070.11 Preprotein translocase secA subunit [Escherichia coli CFT073] pir||F90641 preprotein translocase SecA [imported] - Escherichia coli (strain 0157:H7, substrain RIMD 0609952) pir||F85492 preprotein translocase, secretion protein [imported] - Escherichia coli (strain 0157:H7, substrain EDL933) gb|AAG64402.1 |AE005186_8 preprotein translocase; secretion protein [Escherichia coli 0157:H7 EDL933] dbj|BAB33625.1 | preprotein translocase SecA [Escherichia coli 01δ7:H7] gb|AAN78614.1 |AE01675δ_114 Preprotein translocase secA subunit [Escherichia coli CFT073] Length = 901
1585.2 Best-BlastP=> >nrprot 54% Identities = 11 δ/350 (32%), Positives = 194/360 (55%), Gaps = 31/350 (8%) ref|NP_642273.11 flagellar protein [Xanthomonas axonopodis pv. citri str. 306] gb|AAM36809.11 flagellar protein [Xanthomonas axonopodis pv. citri str. 306] Length = 337
1586.2 Best-BlastP=> >nrprot No Hits found
1587.4 Best-BlastP=> >nrprot 43% Identities = 139/387 (35%), Positives = 203/387 (52%), Gaps = 19/387 (4%) ref|NP_643897.11 oxidoreductase [Xanthomonas axonopodis pv. citri str. 306] gb|AAM38433.1 | oxidoreductase [Xanthomonas axonopodis pv. citri str. 306] Length = 443
1589.3 Best-BlastP=> >nrprot 79% Identities = 138/220 (62%), Positives = 179/220 (81 %), Gaps = 2/220 (0%) ref|NP_820222.11 DNA-binding response regulator [Coxiella burnetii RSA 4 3] gb|AAO90736.11 DNA-binding response regulator [Coxiella burnetii RSA 493] Length = 223
159.1 Best-BlastP=> >nrprot 54% Identities = 44/97 (45%), Positives = 57/97 (58%), Gaps = 14/g7 (14%) ref|NP_747491.1 | hypothetical protein [Pseudomonas putida KT2440] gb|AAN7095δ.1 |AE016739_8 hypothetical protein [Pseudomonas putida KT2440] Length = 91
1590.1 Best-BlastP=> >nrprot 69% Identities = 92/172 (53%), Positives = 126/172 (73%) ref|NP_819923.11 intracellular septation protein A [Coxiella burnetii RSA 493] gb|AAO90437.11 intracellular septation protein A [Coxiella burnetii RSA 493] Length = 181
1591.4 Best-BlastP=> >nrprot 56% Identities = 183/441 (41 %), Positives = 275/441 (62%), Gaps = 13/441 (2%) ref|ZP_00065233.11 COG0741 : Soluble lytic murein transglycosylase and related regulatory proteins (some contain LysM/invasin domains) [Microbulbifer degradans 2-40] Length = 543
1592.3 Best-BlastP=> >nrprot 62% Identities = 130/295 (44%), Positives = 191/295 (64%) ref|NP_406039.11 putative membrane protein [Yersinia pestis] ref|NP_669001.11 hypothetical [Yersinia pestis KIM] pir||AB0306 probable membrane protein YPO2505 [imported] - Yersinia pestis (strain C092) emb|CAC91310.1 | putative membrane protein [Yersinia pestis C092] gb|AAM85252.1 |AE013771_7 hypothetical [Yersinia pestis KIM] Length = 312
1594.2 Best-BlastP=> >nrprot 57% Identities = 83/248 (33%), Positives = 138/248 (65%), Gaps = 14/248 (5%) ref|NP_842295.11 possible BioH, catalyzes some early step in biotin biosynthesis [Nitrosomonas europaea ATCC 19718] emb|CAD86210.11 possible BioH, catalyzes some early step in biotin biosynthesis [Nitrosomonas europaea ATCC 19718] Length = 259
1595.3 Best-BlastP=> >nrprot 52% Identities = 130/329 (39%), Positives = 199/329 (60%), Gaps = 2/329 (0%) ref|NP_790344.11 8-amino-7- oxononanoate synthase [Pseudomonas syringae pv. tomato str. DC3000] gb|AAO54039.1 | 8-amino-7-oxononanoate synthase [Pseudomonas syringae pv. tomato str. DC3000] Length = 396
1598.6 Best-BlastP=> >nrprot 75% Identities = 189/307 (61 %), Positives = 240/307 (78%) gb|AAG47791.1 |AF311738_7 BioB [Mesorhizobium loti] emb|CAD31399.11 BIOTIN SYNTHASE PROTEIN [Mesorhizobium loti] Length = 331
1599.4 Best-BlastP=> >nrprot 48% Identities = 481/1746 (27%), Positives = 836/1746 (47%), Gaps = 149/1746 (8%) ref|NP_932226.11 putative conjugative transfer protein Tral [Vibrio vulnificus YJ016] dbj|BAC97749.11 putative conjugative transfer protein Tral [Vibrio vulnificus YJ016] Length = 1924 .
16.1 Best-BlastP=> >nrprot 60% Identities = 34/55 (61 %), Positives = 42/55 (76%) ref|NP_890077.11 putative phage excisionase [Bordetella bronchiseptica] emb|CAE34036.1 1 putative phage excisionase [Bordetella bronchiseptica] Length = 84
160.1 Best-BlastP=> >nrprot 35% Identities = 36/95 (37%), Positives = 43/95 (45%), Gaps = 7/95 (7%) gb|AAF86199.1 |AF238885_2 VrrB [Bacillus anthracis] Length = 265
1601.3 Best-BlastP=> >nrprot 61 % Identities = 166/370 (44%), Positives = 236/370 (63%), Gaps = 17/370 (4%) ref|NP_660g96.11 glycosyl hydrolase, family 3 [Chlorobium tepidum TLS] gb|AAM71338.11 glycosyl hydrolase, family 3 [Chlorobium tepidum TLS] Length = 372
1602.2 Best-BlastP=> >nrprot 52% Identities = 133/394 (33%), Positives = 207/394 (52%), Gaps = 17/394 (4%) ref|NP_798008.11 putative SAM- dependent methyltransferase [Vibrio parahaemolyticus RIMD 2210633] dbj|BAC59892.11 putative SAM-dependent methyltransferase [Vibrio parahaemolyticus] Length = 418
1603.3 Best-BlastP=> >nrprot 10% Identities = 40/105 (38%), Positives = 55/105 (52%), Gaps = 5/105 (4%) pdb| 1 N0R| A Chain A, 4ank: A Designed Ankyrin Repeat Protein With Four Identical Consensus Repeats Length = 126
1606.2 Best-BlastP=> >nrprot 64% Identities = 136/270 (50%), Positives = 178/270 (65%) ref|ZP_00065849.1 | COG0266: Formamidopyrimidine-DNA glycosylase [Microbulbifer degradans 2-40] Length = 271
1607.2 Best-BlastP=> >nrprot 55% Identities = 24/78 (30%), Positives = 43/78 (55%), Gaps = 1/78 (1 %) ref|NP_820114.1 | hypothetical protein [Coxiella burnetii RSA 493] gb|AAO90628.11 hypothetical protein [Coxiella burnetii RSA 493] Length = 105
1608.2 Best-BlastP=> >nrprot 70% Identities = 201/383 (52%), Positives = 279/383 (72%) ref|NP_903067.11 probable stearoyl-CoA 9-desaturase [Chromobacterium violaceum ATCC 12472] gb|AAQ61061.11 probable stearoyl-CoA 9-desaturase [Chromobacterium violaceum ATCC 12472] Length = 405
1611.5 Best-BlastP=> >nrprot 75% Identities = 206/338 (60%), Positives = 260/338 (76%), Gaps = 1/338 (0%) ref|ZP_00085994.11 COG0128: 5- enolpyruvylshikimate-3-phosphate synthase [Pseudomonas fluorescens PfO-1] Length = 679
1615.2 Best-BlastP=> >nrprot 64% Identities = 291/607 (47%), Positives = 389/607 (64%), Gaps = 17/607 (2%) ref|NP_826054.11 putative oxidoreductase [Streptomyces avermitilis MA-4680] dbj|BAC72589.1 | putative oxidoreductase [Streptomyces avermitilis MA-4680] Length = 642
1617.3 Best-BlastP=> >nrprot 62% Identities = 77/167 (46%), Positives = 107/167 (64%), Gaps = 3/167 (1%) ref|NP_820461.1 | alkylhydroperoxidase AhpD family core domain protein [Coxiella burnetii RSA 493] gb|AAO90975.11 alkylhydroperoxidase AhpD family core domain protein [Coxiella burnetii RSA 493] Length = 177
1618.3 Best-BlastP=> >nrprot 97% Identities = 159/162 (98%), Positives = 159/162 (98%) sp|P53637|SODC_LEGPN Superoxide dismutase [Cu-Zn] precursor dbj|BAA06223.1 | [Cu,Zn]-superoxide dismutase [Legionella pneumophila] gb|AAB36467.1 | periplasmic copper-zinc superoxide dismutas; CuZnSOD [Legionella pneumophila] Length = 162
1623.4 Best-BlastP=> >nrprot 46% Identities = 116/438 (26%), Positives = 202/438 (46%), Gaps = 28/438 (6%) ref |NP_873416.11 conserved hypothetical protein [Haemophilus ducreyi 35000HP] gb|AAP9680δ.1 | conserved hypothetical protein [Haemophilus ducreyi 36000HP] Length = 666
1624.4 Best-BlastP=> >nrprot 42% Identities = 121 /δ43 (22%), Positives = 226/643 (41 %), Gaps = 73/543 (13%) ref|ZP_00089633.11 hypothetical protein [Azotobacter vinelandii] Length = 642
1625.3 Best-BlastP=> >nrprot 90% Identities = 317/379 (83%), Positives = 345/379 (91 %) gb|AAP88975.11 S-adenosylmethionine synthetase [Amoeba proteus symbiotic bacterium] Length = 381
CO CO ,- c CO CM t CN CM '^ m CD CO CO
τ~ --- CM CO m CD CO (33 -— : CM CM CN CO co co co co CO CO CO CD co co CD co CD CD CO CD CD CD CD
1642.3 Best-BlastP=> >nrprot 82% Identities = 150/203 (73%), Positives = 178/203 (87%), Gaps = 1/203 (0%) ref| protease proteolytic subunit [Salmonella enterica subsp. enterica serovar Typhi] ref|NP_469444.11 proteolyt serine protease, heat shock protein F21.5 [Salmonella typhimurium LT2] ref |NP_806142.11 ATP-dependent [Salmonella enterica subsp. enterica serovar Typhi Ty2] sp|Q9LC07|CLPP_SALTY ATP-dependent Clp pr (Endopeptidase Clp) pir||AC0568 ATP-dependent dp protease proteolytic chain [imported] - Salmonella ent (strain CT18) dbj|BAA94668.11 serine protease subunit [Salmonella typhimurium] gb|AAL19403.11 proteolyt serine protease [Salmonella typhimurium LT2] emb|CAD08907.11 ATP-dependent dp protease proteolytic s enterica serovar Typhi] gb|AAO70002.11 ATP-dependent clp protease proteolytic subunit [Salmonella enteri Length = 207
1643.4 Best-BlastP=> >nrprot 97% Identities = 791/816 (96%), Positives = 800/816 (98%) gb|AAM0061δ.11 respo [Legionella pneumophila] Length = 816
1644.6 Best-BlastP=> >nrprot 59% Identities = 171/417 (41 %), Positives = 246/417 (58%), Gaps = 9/417 (2%) ref | [Bradyrhizobium japonicum] dbj|BAC50759.1 | bll5494 [Bradyrhizobium japonicum USDA 110] Length
1645 1 Best-BlastP=> >nrprot 59% Identities = 76/172 (44%), Positives = 110/172 (63%) ref|NP_717325.11 conse oneidensis MR-1] gb|AAN54769.1 |AE015617_4 conserved hypothetical protein [Shewanella oneidensis MR
1647.2 Best-BiastP=> >nrprot 67% Identities = 101/180 (56%), Positives = 127/180 (70%), Gaps = 1/180 (0%) ref| protein [Shewanella oneidensis MR-1] gb|AAN53210.1 |AE01δ463_7 acyltransferase family protein [Shewan 186
1649.3 Best-BlastP=> >nrprot 53% Identities = 276/732 (37%), Positives = 397/732 (54%), Gaps = 63/732 (8%) r [Bacillus cereus ATCC 14579] gb|AAP10083.11 hypothetical protein [Bacillus cereus ATCC 14579] Le
1650.2 Best-BlastP=> >nrprot 35% Identities = 119/317 (37%), Positives = 181/317 (57%), Gaps = 6/317 (1%) ref| oxidoreductase chain L [Bacillus cereus ATCC 14579] gb|AAP10084.11 NADH-quinone oxidoreductase chai Length = 510
1652.4 Best-BlastP=> >nrprot 35% Identities = 37/135 (27%), Positives = 67/135 (49%), Gaps = 11/135 (8%) pir||l sordellii emb|CAA57959.11 cytotoxin L [Clostridium sordellii] Length = 2364
1653.3 Best-BlastP=> >nrprot 63% Identities = 142/281 (50%), Positives = 196/281 (69%), Gaps = 1/281 (0%) pir| precursor - Yersinia pestis Length = 312
1656.2 Best-BlastP=> >nrprot 81% Identities = 41/60 (68%), Positives = 49/60 (81%), Gaps = 2/60 (3%) ref|NP_8 kinase [Nitrosomonas europaea ATCC 19718] emb|CAD86075.1 | Tetraacyldisaccharide-1-P 4'-kinas 19718] Length = 396
1657.1 Best-BlastP=> >nrprot 64% Identities = 124/250 (49%), Positives = 163/250 (65%), Gaps = 7/250 (2%) ref| octulosonate cytidylyltransferase [Haemophilus ducreyi 35000HP] gb|AAP95311.11 3-deoxy-manno- [Haemophilus ducreyi 35000HP] Length = 253
1668.2 Best-BlastP=> >nrprot 33% Identities = 96/392 (24%), Positives = 148/392 (37%), Gaps = 116/392 (29%) Endoglucanase [Microbulbifer degradans 2-40] Length = 725
1659.4 Best-BlastP=> >nrprot 25% Identities = 156/726 (21 %), Positives = 334/726 (46%), Gaps = 107/726 (14%) B, nonmuscle - African clawed frog gb|AAA49915.11 nonmuscle myosin heavy chain b Length = 1992
166.2 Best-BlastP=> >nrprot 96% Identities = 1010/1064 (94%), Positives = 1030/1064 (96%), Gaps = 4/1064 (0 efflux ATPase [Legionella pneumophila] Length = 1060
1660.2 Best-BlastP=> >nrprot No Hits found
1661.3 Best-BlastP=> >nrprot 61 % Identities = 110/213 (51%), Positives = 140/213 (65%), Gaps = 1/213 (0%) sp| protein radC homolog dbj|BAC93049.11 DNA repair protein [Vibrio vulnificus YJ016] Length = 224
1663.6 Best-BlastP=> >nrprot 86% Identities = 1036/1406 (73%), Positives = 1216/1406 (86%), Gaps = 14/1406 ( RNA polymerase, beta' subunit [Pseudomonas putida KT2440] gb|AAN66078.1 |AE016237_2 DNA-d subunit [Pseudomonas putida KT2440] Length = 1399
1664.2 Best-BlastP=> >nrprot 29% Identities = 47/193 (24%), Positives = 80/193 (41 %), Gaps = 13/193 (6%) ref| protein involved in colicin uptake [Lactobacillus gasseri] Length = 962
1666.3 Best-BlastP=> >nrprot No Hits found 1669.3 Best-BlastP=> >nrprot 68% Identities = 177/313 (56%), Positives = 220/313 (70%) ref|NP_617527.11 triac acetivorans str. C2A] gb|AAM06007.1 | triacylglycerol lipase [Methanosarcina acetivorans str. C2A] Le
1670.3 Best-BlastP=> >nrprot 16% Identities = 43/147 (29%), Positives = 74/147 (50%), Gaps = 6/147 (4%) ref|X [Neurospora crassa] gb|EAA27811.11 predicted protein [Neurospora crassa] Length = 554
1671.3 Best-BlastP=> >nrprot No Hits found
1673.5 Best-BlastP=> >nrprot 31 % Identities = 97/388 (25%), Positives = 174/388 (44%), Gaps = 42/388 (10%) pi [imported] - Arabidopsis thaliana gb|AAF82144.1 |AC034256_8 Contains similarity to F-box protein FBL2 fro and contains multiple Leucine Rich PF|00560 repeats. ESTs gb|Z34572, gb|Z34571 , gb|AI100681 , gb|AI100674, gb|AA651378, gb|AA007067, gb|T46145, gb|T22090, gb|AI995016, gb|H36884, come from this gene. [Arabidopsis thaliana] Length = 568
1675.3 Best-BlastP=> >nrprot 39% Identities = 20/67 (29%), Positives = 42/67 (62%) ref|NP_439878.11 hypotheti Rd] sp|P44300|YH36_HAEIN Hypothetical protein HI1736 pir||D64041 hypothetical protein HI1736 - Haem KW20) gb|AAC23384.11 H. influenzae predicted coding region HI1736 [Haemophilus influenzae Rd]
168.1 Best-BlastP=> >nrprot 42% Identities = 43/160 (26%), Positives = 75/160 (46%), Gaps = 25/160 (15%) ref| pprrootteeiinn [[SSiinnoorrhhiizzoobbiiuumm mmeelliilloottii]] ppiirr||||EE9955995588 hhyyppootthheettiiccaall mmeemmbbrraannee pprrootteeiinn [[iimmppoorrtteedd]] -- SSiinnoorrhhiizzoobbiiuumm melil pSymB emb|CAC49333.1 | hypothetical membrane protein [Sinorhizobium meliloti] Length = 172
1681.5 Best-BlastP=> >nrprot 65% Identities = 432/984 (43%), Positives = 629/984 (63%), Gaps = 47/984 (4%) r associated protein (ATP-dependent helicase) [Photorhabdus luminescens subsp. laumondii TT01] e associated protein (ATP-dependent helicase) [Photorhabdus luminescens subsp. laumondii TT01]
1682.4 Best-BlastP=> >nrprot 35% Identities = 63/190 (33%), Positives = 105/190 (56%), Gaps = 8/190 (4%) ref| desaturase [Microbulbifer degradans 2-40] Length = 273
1683.2 Best-BlastP=> >nrprot No Hits found
1684.3 Best-BlastP=> >nrprot No Hits found 1685.3 Best-BlastP=> >nrprot 72% Identities = 199/372 (53%), Positives = 270/372 (72%), Gaps = 3/372 (0%) ref|ZP_00016064.11 COG0842: ABC-type multidrug transport system, permease component [Rhodospirillum rubrum] Length = 376
1689.3 Best-BlastP=> >nrprot 68% Identities = 207/377 (54%), Positives = 275/377 (72%), Gaps = 9/377 (2%) ref |NP_720122.11 cytochrome c oxidase, subunit II [Shewanella oneidensis MR-1] gb|AAN57566.1 |AE015892_6 cytochrome c oxidase, subunit II [Shewanella oneidensis MR-1] Length = 513 169.1 Best-BlastP=> >nrprot 86% Identities = 55/68 (80%), Positives = 60/68 (88%) gb|AAM00618.11 unknown [Legionella pneumophila] Length = 68
1691.3 Best-BlastP=> >nrprot 84% Identities = 388/528 (73%), Positives = 458/528 (86%), Gaps = 8/528 (1%) ref| P_720123.1 | cytochrome c oxidase, subunit I [Shewanella oneidensis MR-1] gb|AAN57567.1 |AE015892_7 cytochrome c oxidase, subunit I [Shewanella oneidensis MR-1] Length = 530 1692.2 Best-BlastP=> >nrprot 86% Identities = 473/671 (70%), Positives = 577/671 (85%), Gaps = 7/671 (1%) ref|NP_819550.11 excinuclease ABC, B subunit [Coxiella burnetii RSA 493] gb|AAO90064.11 excinuclease ABC, B subunit [Coxiella burnetii RSA 493] Length = 672
1693.2 Best-BlastP=> >nrprot 56% Identities = 27/75 (36%), Positives = 45/76 (60%) ref|NP_754358.11 Hypothetical protein [Escherichia coli CFT073] gb|AAN80925.1 |AE016762_178 Hypothetical protein [Escherichia coli CFT073] Length = 82
1695.2 Best-BlastP=> >nrprot 56% Identities = 118/355 (33%), Positives = 189/355 (53%), Gaps = 47/355 (13%) ref|NP_884030.11 conserved hypothetical protein [Bordetella parapertussis] emb|CAE37058.1 | conserved hypothetical protein [Bordetella parapertussis] Length = 392
1697.2 Best-BlastP=> >nrprot 56% Identities = 66/174 (37%), Positives = 104/174 (59%), Gaps = 2/174 (1 %) ref |ZP_00013244.1 | COG3038: Cytochrome B561 [Rhodospirillum rubrum] Length = 188
1698.2 Best-BlastP=> >nrprot 35% Identities = 84/273 (30%), Positives = 130/273 (47%), Gaps = 17/273 (6%) ref|ZP_00085491.11 COG0354: Predicted aminomethyltransferase related to GcvT [Pseudomonas fluorescens PfO-1] Length = 313
1699.4 Best-BlastP=> >nrprot 58% Identities = 130/277 (46%), Positives = 176/277 (63%), Gaps = 6/277 (2%) ref|ZP_00090118.1 | COG0771 : UDP-N- ' acetylmuramoylalanine-D-glutamate ligase [Azotobacter vinelandii] Length = 448
1701.5 Best-BlastP=> >nrprot 66% Identities = 192/359 (53%), Positives = 262/359 (72%) ref|NP_819182.1 | cell division protein FtsW [Coxiella burnetii RSA 493] gb|AAD39750.1 |AF123260_1 FtsW [Coxiella burnetii] gb|AA089696.11 cell division protein FtsW [Coxiella burnetii RSA 493] Length = 372
1703.1 Best-BlastP=> >nrprot 36% Identities = 57/215 (26%), Positives = 93 215 (43%), Gaps = 32/215 (14%) ref|NP_780971.1 | DNA helicase [Clostridium tetani E88] gb|AAO34g08.11 DNA helicase [Clostridium tetani E88] Length = 1352
1704.4 Best-BlastP=> >nrprot δ% Identities = 33/108 (30%), Positives = 55/108 (50%), Gaps = 5/108 (4%) ref|NP_266277.1 | hypothetical protein [Lactococcus lactis subsp. lactis] pir||A86640 hypothetical protein ybcH [imported] - Lactococcus lactis subsp. lactis (strain IL1403) gb|AAK04219.1 |AE006250_6 HYPOTHETICAL PROTEIN [Lactococcus lactis subsp. lactis] Length = 311
1705.2 Best-BlastP=> >nrprot No Hits found
CM CN rt ι- CM CN rt O CM m
CD Is-! CO (33 co rt O CM O O O O O CM CM CM CN Is- Is- Is- Is- Is- Is-
1724.3 Best-BlastP=> >nrprot 35% Identities = 62/241 (25%), Positives = 104/241 (43%), Gaps = 20/241 (8%) ref| [Mesorhizobium loti] dbj|BAB48103.11 probable hydrolase [Mesorhizobium loti] Length = 248
1725.3 Best-BlastP=> >nrprot 74% Identities = 172/279 (61 %), Positives = 212/279 (75%) ref|NP_747209.11 RNA [Pseudomonas putida KT2440] gb|AAF80334.1 |AF157048_1 heat shock sigma factor RpoH [Pseudomonas RNA polymerase sigma-32 factor [Pseudomonas putida KT2440] Length = 284
1726.4 Best-BlastP=> >nrprot 14% Identities = 50/174 (28%), Positives = 82/174 (47%), Gaps = 18/174 (10%) ref| [Plasmodium falciparum 3D7] pir||T18459 hypothetical protein C051 δc - malaria parasite (Plasmodium hypothetical protein [Plasmodium falciparum 3D7] Length = 1236
1730.3 Best-BlastP=> >nrprot 72% Identities = 498/899 (55%), Positives = 632/899 (70%), Gaps = 39/899 (4%) r factor IF-2 [Yersinia pestis KIM] sp|Q8ZBC2|IF2_YERPE Translation initiation factor IF-2 gb|AAM84276.1 | factor IF-2 [Yersinia pestis KIM] Length = 892
1733.3 Best-BlastP=> >nrprot 70% Identities = 225/399 (56%), Positives = 293/399 (73%), Gaps = 1/399 (0%) ref| Sodium:dicarboxylate symporter family [Bacillus anthracis A2012] ref|NP_847618.1 | proton/glutamat [Bacillus anthracis str. Ames] gb|AAP29104.11 proton/glutamate symporter family protein, putative [B Length = 405
1734.2 Best-BlastP=> >nrprot No Hits found
1735.1 Best-BlastP=> >nrprot No Hits found
1737.2 Best-BlastP=> >nrprot 34% Identities = 88/258 (34%), Positives = 129/258 (50%), Gaps = 30/258 (11 %) r [Coxiella burnetii] gb|AAD33500.1 |AF131076_26 hypothetical protein [Coxiella burnetii] Length = 361
1738.3 Best-BlastP=> >nrprot 84% Identities = 284/394 (72%), Positives = 339/394 (86%) ref|ZP_00065178.1 1 C chain [Microbulbifer degradans 2-40] Length = 403
1739.2 Best-BlastP=> >nrprot 65% Identities = 95/197 (48%), Positives = 136/197 (69%) ref|NP_902433.11 phosp [Chromobacterium violaceum ATCC 12472] gb|AAQ60431 .1 | phosphoribosylanthranilate isomerase ATCC 12472] Length = 206
174.1 Best-BlastP=> >nrprot 71 % Identities = 165/284 (58%), Positives = 206/284 (72%) ref|NP_820298.1 1 pepti [Coxiella burnetii RSA 493] gb|AAO90812.11 peptide methionine sulfoxide reductase [Coxiella burnetii RSA
1740.3 Best-BlastP=> >nrprot 73% Identities = 148/259 (57%), Positives = 193/259 (74%), Gaps = 1/259 (0%) ref| Pseudouridylate synthase [Pseudomonas fluorescens PfO-1] Length = 313
1742.3 Best-BlastP=> >nrprot 79% Identities = 163/239 (68%), Positives = 198/239 (82%), Gaps = 1/239 (0%) ref| kinase [Azotobacter vinelahdii] Length = 24g
1743.4 Best-BlastP=> >nrprot 84% Identities = 118/184 (64%), Positives = 158/184 (85%) ref|NP_928020.11 ribos luminescens subsp. laumondii TT01] emb|CAE12970.1 | ribosome releasing factor [Photorhabdus lu TT01] Length = 186
1746.2 Best-BlastP=> >nrprot 47% Identities = 71/209 (33%), Positives = 100/209 (47%), Gaps = 8/209 (3%) ref|NP_23314δ.11 arginine ABC transporter, periplasmic arginine-binding protein [Vibrio cholerae 01 biovar eltor str. N16961] pir||H82420 arginine ABC transporter, periplasmic arginine-binding protein VCA0759 [imported] - Vibrio cholerae (strain N 16961 serogroup 01 ) gb|AAF96657.1 | arginine ABC transporter, periplasmic arginine-binding protein [Vibrio cholerae 01 biovar eltor str. N16961] Length = 243
1747.2 Best-BlastP=> >nrprot No Hits found
1748.3 Best-BlastP=> >nrprot 57% Identities = 160/430 (37%), Positives = 244/430 (56%), Gaps = 14/430 (3%) ref|NP_662910.1 | threonine synthase [Chlorobium tepidum TLS] gb|AAM73252.11 threonine synthase [Chlorobium tepidum TLS] Length = 441
1749.2 Best-BlastP=> >nrprot 29% Identities = 48/192 (26%), Positives = 77/192 (40%), Gaps = 21/192 (10%) ref|NP_698612.11 outer membrane protein, 31 kDa [Brucella suis 1330] gb|AAN30527.1 |AE014465_11 outer membrane protein, 31 kDa [Brucella suis 1330] Length = 261
1750.3 Best-BlastP=> >nrprot 69% Identities = 162/295 (54%), Positives = 201/295 (68%), Gaps = 6/295 (2%) ref|NP_742276.11 cytochrome c oxidase, subunit III [Pseudomonas putida KT2440] gb|AAN65740.1 |AE016200_4 cytochrome c oxidase, subunit III [Pseudomonas putida KT2440] Length = 295 752.2 Best-BlastP=> >nrprot 66% Identities = 83/173 (47%), Positives = 120/173 (69%), Gaps = 2/173 (1 %) ref|NP_720124.11 cytochrome c oxidase assembly protein coxG [Shewanella oneidensis MR-1] gb|AAN57568.1 |AE015892_8 cytochrome c oxidase assembly protein coxG [Shewanella oneidensis MR-1] Length = 193
1754.3 Best-BlastP=> >nrprot 49% Identities = 174/422 (41 %), Positives = 252/422 (59%), Gaps = 9/422 (2%) ref|NP_820704.11 thiol:disulfide interchange protein DsbD [Coxiella burnetii RSA 493] gb|AA091218.11 thiol:disulfide interchange protein DsbD [Coxiella burnetii RSA 493] Length = 584
1755.3 Best-BlastP=> >nrprot No Hits found
1756.4 Best-BlastP=> >nrprot 63% Identities = 133/271 (4g%), Positives = 181/271 (66%) ref|NP_902359.11 probable oxidoreductase, short-chain dehydrogenase/reductase family [Chromobacterium violaceum ATCC 12472] gb|AAQ60359.11 probable oxidoreductase, short-chain dehydrogenase/reductase family [Chromobacterium violaceum ATCC 12472] Length = 278
1759.4 Best-BlastP=> >nrprot 43% Identities = 55/224 (24%), Positives = 102/224 (45%), Gaps = 3/224 (1 %) ref|NP_746234.11 conserved hypothetical protein [Pseudomonas putida KT2440] gb|AAN69698.1 |AE016606_1 conserved hypothetical protein [Pseudomonas putida KT2440] Length = 267
176.1 Best-BlastP=> >nrprot No Hits found
1760.4 Best-BlastP=> >nrprot 75% Identities = 116/223 (52%), Positives = 170/223 (76%) ref|XP_306643.11 ENSANGP00000000843 [Anopheles gambiae] gb|EAA02110.11 ENSANGP00000000843 [Anopheles gambiae str. PEST] Length = 228
1761.4 Best-BlastP=> >nrprot 9% Identities = 54/209 (25%), Positives = 100/209 (47%), Gaps = 38/209 (18%) ref|ZP_00010059.11 COG1680: Beta- lactamase class C and other penicillin binding proteins [Rhodopseudomonas palustris] Length = 395
1762.2 Best-BlastP=> >nrprot 39% Identities = 66/245 (26%), Positives = 125/245 (51 %), Gaps = 9/245 (3%) ref|NP_834835.11 Transcriptional regulator, MerR family [Bacillus cereus ATCC 14579] gb|AAP12036.11 Transcriptional regulator, MerR family [Bacillus cereus ATCC 14579] Length = 254
1764.3 Best-BlastP=> >nrprot 99% Identities = 377/377 (100%), Positives = 377/377 (100%) emb|CAB65211.1 | N-acylglucosamine 2-epimerase [Legionella pneumophila] Length = 377
1765.4 Best-BlastP=> >nrprot 99% Identities = 201/201 (100%), Positives = 201/201 (100%) emb|CAB65210.1 | putative acetyl transferase [Legionella pneumophila] Length = 419
1767.2 Best-BlastP=> >nrprot 56% Identities = 103/234 (44%), Positives = 135/234 (57%), Gaps = 1/234 (0%) prf||1712315B glycerophosphoryl diester esterase Length = 247
1768.3 Best-BlastP=> >nrprot 52% Identities = 150/405 (37%), Positives = 216/405 (53%), Gaps = 12/405 (2%) ref|NP_5i 385.1| PROBABLE GLYCEROL-3-PHOSPHATE-BINDING PERIPLASMIC LIPOPROTEIN SIGNAL PEPTIDE [Ralstonia solanacearum] emb|CAD14g66.11 PROBABLE GLYCEROL-3-PHOSPHATE-BINDING PERIPLASMIC LIPOPROTEIN SIGNAL PEPTIDE [Ralstonia solanacearum] Length = 438
1770.2 Best-BlastP=> >nrprot 99% Identities = 1034/1035 (99%), Positives = 1034/1035 (99%) gb|AAF32510.1 |AF095231_1 defect in organelle trafficking protein [Legionella pneumophila] Length = 1035
1772.1 Best-BlastP=> >nrprot 94% Identities = 136/161 (89%), Positives = 144/151 (95%) gb|AAC35591.1| IcmV [Legionella pneumophila] Length = 161
1773.1 Best-BlastP=> >nrprot 99% Identities = 150/161 (99%), Positives = 151/151 (100%) pir||S61384 icmW protein - Legionella pneumophila gb|AAC35589.11 IcmW [Legionella pneumophila] Length = 151
1774.2 Best-BlastP=> >nrprot 90% Identities = 411/472 (87%), Positives = 426/472 (90%), Gaps = 6/472 (1 %) gb|AAC35590.11 IcmX [Legionella pneumophila] Length = 466
1775.3 Best-BlastP=> >nrprot No Hits found
1778.5 Best-BlastP=> >nrprot 62% Identities = 63/115 (64%), Positives = 82/11 δ (71 %), Gaps = 4/116 (3%) ref|NP_215343.11 hypothetical protein Rv0828c [Mycobacterium tuberculosis H37Rv] ref|NP_854509.1 | POSSIBLE DEAMINASE [Mycobacterium bovis subsp. bovis AF2122/97] pir||D70811 hypothetical protein Rv0828c - Mycobacterium tuberculosis (strain H37RV) emb|CAA17634.11 hypothetical protein Rv0828c [Mycobacterium tuberculosis H37Rv] emb|CAD93713.11 POSSIBLE DEAMINASE [Mycobacterium bovis subsp. bovis AF2122/97] Length = 140
1779.3 Best-BlastP=> >nrprot No Hits found 178.1 Best-BlastP=> >nrprot No Hits found
1780.3 Best-BlastP=> >nrprot 80% Identities = 51/69 (73%), Positives = 56/69 (81%) ref|ZP_00067276.1 | COG1278: Cold shock proteins [Microbulbifer degradans 2-40] Length = 71
1781.4 Best-BlastP=> >nrprot 72% Identities = 245/436 (56%), Positives = 318/436 (72%), Gaps = 1/436 (0%) ref|ZP_00031144.11 COG0513: Superfamily II DNA and RNA helicases [Burkholderia fungorum] Length = 534
1782.3 Best-BlastP=> >nrprot 51 % Identities = 134/404 (33%), Positives = 216/404 (53%), Gaps = 3/404 (0%) ref|NP_819936.11 major facilitator family transporter [Coxiella burnetii RSA 493] gb|AAO90449.11 major facilitator family transporter [Coxiella burnetii RSA 493] Length = 437
1784.2 Best-BlastP=> >nrprot No Hits found
1786.2 Best-BlastP=> >nrprot 99% Identities = 123/123 (100%), Positives = 123/123 (100%) gb|AAN08839.1 | transmission trait enhancer protein LetE [Legionella pneumophila] Length = 123
1787.4 Best-BlastP=> >nrprot 26% Identities = 131/667 (19%), Positives = 275/667 (41 %), Gaps = 111/667 (16%) ref|NP_082559.1 | nuclear membrane binding protein NUCLING [Mus musculus] gb|AAH42415.1 | Nuclear membrane binding protein NUCLING [Mus musculus] Length = 1413
1788.3 Best-BlastP=> >nrprot 62% Identities = 208/445 (46%), Positives = 285/445 (64%), Gaps = 6/445 (1 %) ref|ZP_00118630.11 hypothetical protein [Cytophaga hutchinsonii] Length = 452
179.1 Best-BlastP=> >nrprot 97% Identities = 85/89 (95%), Positives = 88/89 (98%) emb|CAC34415.11 putative TatB protein [Legionella pneumophila] Length = 89
1792.2 Best-BlastP=> >nrprot 46% Identities = 70/254 (27%), Positives = 121/254 (47%), Gaps = 13/254 (5%) ref|NP_421699.11 hypothetical protein [Caulobacter crescentus CB15] pir||G87608 hypothetical protein CC2905 [imported] - Caulobacter crescentus gb|AAK24867.1 | hypothetical protein [Caulobacter crescentus CB15] Length = 261
1793.3 Best-BlastP=> >nrprot 65% Identities = 133/262 (50%), Positives = 182/262 (69%), Gaps = 1/262 (0%) ref |NP_819072.11 xanthosine phosphorylase [Coxiella burnetii RSA 493] gb|AA089586.11 xanthosine phosphorylase [Coxiella burnetii RSA 493] Length = 273
17g4.3 Best-BlastP=> >nrprot 51 % Identities = 88/257 (34%), Positives = 132/257 (51 %), Gaps = 21/257 (8%) ref|NP_671037.11 2-deoxyribose-5- phosphate aldolase [Yersinia pestis KIM] gb|AAM87288.1 |AE013977_7 2-deoxyribose-5-phosphate aldolase [Yersinia pestis KIM] Length = 270
1795.3 Best-BlastP=> >nrprot 19% Identities = 101/385 (26%), Positives = 172/385 (44%), Gaps = 54/385 (14%) ref|NP_220907.11 unknown [Rickettsia prowazekii] pir||E71657 hypothetical protein RP534 - Rickettsia prowazekii emb|CAA14983.11 unknown [Rickettsia prowazekii] Length = 598
1797.4 Best-BlastP=> >nrprot 66% Identities = 123/252 (48%), Positives = 174/252 (69%), Gaps = 8/252 (3%) dbj|BAA20497.1 | 27kDa outer membrane protein [Coxiella burnetii] Length = 262
18.1 Best-BlastP=> >nrprot No Hits found
180.1 Best-BlastP=> >nrprot 98% Identities = 60/61 (98%), Positives = 61/61 (100%) emb|CAC34414.11 putative TatA protein [Legionella pneumophila] Length = 61 1800.4 Best-BlastP=> >nrprot 74% Identities = 417/723 (57%), Positives = 543/723 (75%), Gaps = 7/723 (0%) ref|NP_407289.11 DNA helicase II [Yersinia pestis] pir||AI0467 DNA helicase II (EC 3.6.1.-) [imported] - Yersinia pestis (strain C092) emb|CAC93309.11 DNA helicase II [Yersinia pestis C092] Length = 720
1803.2 Best-BlastP=> >nrprot 45% Identities = 30/59 (60%), Positives = 40/69 (67%) ref|ZP_00021376.11 COG0477: Permeases of the major facilitator superfamily [Ralstonia metallidurans] Length = 120
1804.2 Best-BlastP=> >nrprot 88% Identities = 320/414 (77%), Positives = 367/414 (88%) gb|AAA92282.1 | Hel Length = 414
1805.3 Best-BlastP=> >nrprot 27% Identities = 39/113 (34%), Positives = 59/113 (52%), Gaps = 6/113 (4%) ref|NP_811310.11 conserved hypothetical protein [Bacteroides thetaiotaomicron VPI-5482] gb|AAO77504.1 | conserved hypothetical protein [Bacteroides thetaiotaomicron VPI- 5482] Length = 207
1806.2 Best-BlastP=> >nrprot 80% Identities = 90/151 (59%), Positives = 121/151 (80%), Gaps = 3/151 (1 %) ref|NP_820500.1 | RNA methyltransferase, TrmH family, group 2 [Coxiella burnetii RSA 493] gb|AAO91014.11 RNA methyltransferase, TrmH family, group 2 [Coxiella burnetii RSA 493] Length = 152
1807.3 Best-BlastP=> >nrprot 65% Identities = 221/460 (48%), Positives = 309/460 (67%), Gaps = 2/460 (0%) ref |NP_841539.11 putative homospermidine synthase protein [Nitrosomonas europaea ATCC 19718] emb|CAD85409.11 putative homospermidine synthase protein [Nitrosomonas europaea ATCC 19718] Length = 472
1809.4 Best-BlastP=> >nrprot 42% Identities = 190/672 (33%), Positives = 326/672 (66%), Gaps = 4/572 (0%) ref|NP_902623.11 conserved hypothetical protein [Chromobacterium violaceum ATCC 12472] gb|AAQ60621.1 | conserved hypothetical protein [Chromobacterium violaceum ATCC 12472] Length = 1390
1812.4 Best-BlastP=> >nrprot 42% Identities = 88/335 (26%), Positives = 168/335 (50%), Gaps = 2/335 (0%) ref|ZP_00092491.1 | COG0477: Permeases of the major facilitator superfamily [Azotobacter vinelandii] Length = 432
1815.2 Best-BlastP=> >nrprot 45% Identities = 49/148 (33%), Positives = 80/148 (54%) ref|ZP_00087676.11 COG0454: Histone acetyltransferase HPA2 and related acetyltransferases [Pseudomonas fluorescens PfO-1] Length = 167
1817.2 Best-BlastP=> >nrprot No Hits found 1818.4 Best-BlastP=> >nrprot 65% Identities = 280/280 (100%), Positives = 280/280 (100%) emb|CAB65209.11 hypothetical protein [Legionella pneumophila] Length = 280
1819.6 Best-BlastP=> >nrprot 58% Identities = 819/859 (95%), Positives = 836/859 (97%), Gaps = 1/859 (0%) gb|AAD47371.11 LigA [Legionella pneumophila] Length = 869
182.3 Best-BlastP=> >nrprot 66% Identities = 272/471 (57%), Positives = 367/471 (77%), Gaps = 2/471 (0%) ref|NP_820993.11 ubiquinone biosynthesis protein AarF, putative [Coxiella burnetii RSA 493] gb|AA091507.11 ubiquinone biosynthesis protein AarF, putative [Coxiella . burnetii RSA 493] Length = 541
1823.6 Best-BlastP=> >nrprot 66% Identities = 206/419 (49%), Positives = 287/419 (68%) ref|NP_262387.11 hypothetical protein [Pseudomonas aeruginosa PA01] pir||A83183 hypothetical protein PA3697 [imported] - Pseudomonas aeruginosa (strain PA01 ) . gb|AAG07085.1 |AE004789_δ hypothetical protein PA3697 [Pseudomonas aeruginosa PA01] Length = 431
1826.2 Best-BlastP=> >nrprot No Hits found
1827.3 Best-BlastP=> >nrprot No Hits found
1831.3 Best-BlastP=> >nrprot 73% Identities = 235/399 (58%), Positives = 297/399 (74%) ref|NP_819677.11 riboflavin biosynthesis protein RibA [Coxiella burnetii RSA 493] gb|AAO90191.11 riboflavin biosynthesis protein RibA [Coxiella burnetii RSA 493] Length = 406
1832.4 Best-BlastP=> >nrprot 58% Identities = 82/191 (42%), Positives = 120/191 (62%) ref|NP_819676.11 riboflavin synthase, alpha subunit [Coxiella burnetii RSA 493] gb|AAO90190.11 riboflavin synthase, alpha subunit [Coxiella burnetii RSA 493] Length = 202
1834.4 Best-BlastP=> >nrprot 49% Identities = 144/439 (32%), Positives = 228/439 (51%), Gaps = 14/439 (3%) ref|NP_484368.11 hypothetical protein [Nostoc sp. PCC 7120] pir||AC1847 hypothetical protein all0324 [imported] - Nostoc sp. (strain PCC 7120) dbj|BAB72282.11 ORF_ID:all0324~hypothetical protein [Nostoc sp. PCC 7120] Length = 447
1838.2 Best-BlastP=> >nrprot 52% Identities = 170/486 (34%), Positives = 256/486 (52%), Gaps = 27/486 (5%) ref|ZP_00110911.11 COG1816: Adenosine deaminase [Nostoc punctiforme] Length = 523
1839.2 Best-BlastP=> >nrprot 57% Identities = 157/414 (37%), Positives = 230/414 (55%), Gaps = 33/414 (7%) ref|NP_711587.11 sterol desaturase- related protein [Leptospira interrogans serovar lai str. 56601] gb|AAN48605.1 |AE01 320_5 sterol desaturase-related protein [Leptospira interrogans serovar lai str. 56601] Length = 442
184.2 Best-BlastP=> >nrprot No Hits found
1840.3 Best-BlastP=> >nrprot 53% Identities = 138/380 (36%), Positives = 220/380 (57%), Gaps = 7/380 (1 %) ref|NP_820736.11 drug resistance transporter, Bcr/CflA family [Coxiella burnetii RSA 493] gb|AA091250.11 drug resistance transporter, Bcr/CflA family [Coxiella burnetii RSA 493] Length = 393
1841.2 Best-BlastP=> >nrprot 58% Identities = 215/536 (40%), Positives = 327/536 (61 %), Gaps = 9/536 (1 %) ref|NP_715995.11 AMP-binding protein [Shewanella oneidensis MR-1] gb|AAN53440.1 |AE015483_6 AMP-binding protein [Shewanella oneidensis MR-1] Length = 554
1844.3 Best-BlastP=> >nrprot 54% Identities = 120/330 (36%), Positives = 193/330 (58%), Gaps = 3/330 (0%) ref|NP_925659.11 UDP-3-0-[3- hydroxymyristoyl] glucosamine N-acyltransferase [Gloeobacter violaceus] dbj|BAC90654.1 | UDP-3-0-[3-hydroxymyristoyl] glucosamine N- acyltransferase [Gloeobacter violaceus] Length = 346
1845.3 Best-BlastP=> >nrprot No Hits found 1846.3 Best-BlastP=> >nrprot 62% Identities = 181/411 (44%), Positives = 255/411 (62%), Gaps = 4/411 (0%) gb|AAN87389.11 3-oxoacyl-[acyl-carrier- protein] synthase [Heliobacillus mobilis] Length = 415
1847.3 Best-BlastP=> >nrprot 19% Identities = 77/190 (40%), Positives = 110/190 (57%), Gaps = δ/190 (2%) ref|NP_253486.11 hypothetical protein [Pseudomonas aeruginosa PA01] sp|Q9HV11 |YBJ8_PSEAE Hypothetical protein PA4798 pir||C83045 hypothetical protein PA4798 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG08184.1 |AE004893_2 hypothetical protein PA4798 [Pseudomonas aeruginosa PA01] Length = 242
1848.4 Best-BlastP=> >nrprot 31% Identities = 77/268 (29%), Positives = 129/258 (50%), Gaps = 21/258 (8%) ref|NP_832609.11 Methyltransferase [Bacillus cereus ATCC 14679] gb|AAP09810.11 Methyltransferase [Bacillus cereus ATCC 14679] Length = 263
1849.3 Best-BlastP=> >nrprot 40% Identities = 46/134 (33%), Positives = 73/134 (54%), Gaps = 4/134 (2%) ref|NP_636084.1 | acetyltransferase [Xanthomonas campestris pv. campestris str. ATCC 33913] gb|AAM40008.1 | acetyltransferase [Xanthomonas campestris pv. campestris str. ATCC 33913] Length = 157
185.1 Best-BlastP=> >nrprot 69% Identities = 141/253 (56%), Positives = 182/253 (71 %), Gaps = 1/263 (0%) ref|NP_484519.1 | probable short-chain dehydrogenase [Nostoc sp. PCC 7120] pir||AB1866 hypothetical protein all0475 [imported] - Nostoc sp. (strain PCC 7120) dbj|BAB72433.11 ORF_ID:all047δ~probable short-chain dehydrogenase [Nostoc sp. PCC 7120] Length = 257
1852.2 Best-BlastP=> >nrprot 54% Identities = 43/110 (39%), Positives = 65/110 (59%), Gaps = 6/110 (5%) ref|NP_900345.1 | conserved hypothetical protein [Chromobacterium violaceum ATCC 12472] gb|AAQ58351.11 conserved hypothetical protein [Chromobacterium violaceum ATCC 12472] Length = 128
1853.2 Best-BlastP=> >nrprot 58% Identities = 65/127 (51 %), Positives = 79/127 (62%), Gaps = 9/127 (7%) ref|NP_459344.11 putative outer membrane lipoprotein [Salmonella typhimurium LT2] gb|AAL19303.11 putative outer membrane lipoprotein [Salmonella typhimurium LT2] Length = 119
1856.3 Best-BlastP=> >nrprot No Hits found
1858.1 Best-BlastP=> >nrprot No Hits found
1869.2 Best-BlastP=> >nrprot 51% Identities = 85/271 (31 %), Positives = 139/271 (51 %), Gaps = 21/271 (7%) ref|NP_219985.11 hypothetical protein [Chlamydia trachomatis] pir||E71509 hypothetical protein CT472 - Chlamydia trachomatis (serotype D, strain UW3/Cx) gb|AAC68072.11 hypothetical protein [Chlamydia trachomatis] Length = 264
1860.2 Best-BlastP=> >nrprot 47% Identities = 104/344 (30%), Positives = 174/344 (60%), Gaps = 38/344 (11 %) gb|AAF86695.1 |AF180966_1 AMPC cephalosporinase precursor protein ACC-3a [Hafnia alvei] Length = 377
1861.2 Best-BlastP=> >nrprot No Hits found
1862.2 Best-BlastP=> >nrprot 63% Identities = 206/470 (43%), Positives = 279/470 (59%), Gaps = 41/470 (8%) ref|NP_249771.11 flagellar hook protein FlgE [Pseudomonas aeruginosa PA01] pir||F83510 flagellar hook protein FlgE PA1080 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG04469.1 |AE004539_11 flagellar hook protein FlgE [Pseudomonas aeruginosa PA01] Length = 462
1863.2 Best-BlastP=> >nrprot 64% Identities = 97/223 (43%), Positives = 145/223 (65%), Gaps = 2/223 (0%) ref |ZP_00138665.11 COG1843: Flagellar hook capping protein [Pseudomonas aeruginosa UCBPP-PA14] Length = 237
1864.3 Best-BlastP=> >nrprot 73% Identities = 273/454 (60%), Positives = 346/454 (76%), Gaps = 5/454 (1%) ref |NP_638130.11 succinyl- diaminopimelate desuccinylase [Xanthomonas campestris pv. campestris str. ATCC 33913] gb|AAM42054.11 succinyl-diaminopimelate desuccinylase [Xanthomonas campestris pv. campestris str. ATCC 33913] Length = 4 7
1865.3 Best-BlastP=> >nrprot 72% Identities = 175/353 (49%), Positives = 263/353 (74%), Gaps = 1/353 (0%) ref|NP_562979.11 UDP-N- acetylglucosamine-N-acetylmuramyl-(pentape ptide) pyrophosphoryl N-acetylglucosamine transferase [Clostridium perfringens] sp|Q8XIQ1 |MURG_CLOPE UDP-N-acetylglucosamine-N-acetylmuramyl-(pentapeptide) pyrophosphoryl-undecaprenol N- acetylglucosamine transferase (Undecaprenyl-PP-MurNAc-pentapeptide-UDPGIcNAc GlcNAc transferase) dbj|BAB81769.11 UDP-N-acetylglucosamine-N-acetylmuramyl-(pentape ptide) pyrophosphoryl N-acetylglucosamine transferase [Clostridium perfringens str. 13] Length = 357
187.1 Best-BlastP=> >nrprot No Hits found
1870.3 Best-BlastP=> >nrprot 60% Identities = 152/349 (43%), Positives = 223/349 (63%), Gaps = 19/349 (5%) ref|NP_767636.11 HlyD family secretion protein [Bradyrhizobium japonicum] dbj|BAC46261.11 HlyD family secretion protein [Bradyrhizobium japonicum USDA 110] Length = 410
1871.2 Best-BlastP=> >nrprot 99% Identities = 348/348 (100%), Positives = 348/348 (100%) emb|CAC33484.11 RecA protein [Legionella pneumophila] Length = 348
1873.2 Best-BlastP=> >nrprot 99% Identities = 150/160 (100%), Positives = 150/160 (100%) sp|P37864|RECX_LEGPN Regulatory protein recX emb|CAC33485.11 RecX protein [Legionella pneumophila] Length = 150
1875.2 Best-BlastP=> >nrprot 7δ% Identities = 234/379 (61 %), Positives = 294/379 (77%) ref|NP_842481.11 General substrate transporters [Nitrosomonas europaea ATCC 19718] emb|CAD86404.11 General substrate transporters [Nitrosomonas europaea ATCC 19718] Length = 391
1878.3 Best-BlastP=> >nrprot 63% Identities = 130/311 (41 %), Positives = 199/311 (63%), Gaps = 7/311 (2%) ref|NP_520407.11 HYPOTHETICAL PROTEIN [Ralstonia solanacearum] emb|CAD15993.11 HYPOTHETICAL PROTEIN [Ralstonia solanacearum] Length = 317
188.1 Best-BlastP=> >nrprot No Hits found
1880.3 Best-BlastP=> >nrprot 79% Identities = 208/324 (64%), Positives = 262/324 (80%) ref|NP_820236.11 malate dehydrogenase [Coxiella burnetii RSA 493] gb|AAO90750.11 malate dehydrogenase [Coxiella burnetii RSA 493] Length = 328
1882.2 Best-BlastP=> >nrprot 52% Identities = 116/255 (45%), Positives = 165/256 (64%), Gaps = 12/255 (4%) ref | NP_819457.1| polysaccharide deacetylase-related protein [Coxiella burnetii RSA 493] gb|AA089971.1 | polysaccharide deacetylase-related protein [Coxiella burnetii RSA 493] Length = 276
1883.4 Best-BlastP=> >nrprot No Hits found
1885.3 Best-BlastP=> >nrprot 67% Identities = 221/446 (49%), Positives = 307/446 (68%), Gaps = 5/446 (1%) ref|NP_252709.11 UDP-N- acetylmuramate:L-alanyl-gamma-D-glutamyl-meso-diaminopimelate ligase [Pseudomonas aeruginosa PA01] pir||A83145 UDP-N- acetylmuramate-L-alanyl-gamma-D-glutamyl-meso-diaminop imelate ligase (EC 6.3.2.-) PA4020 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG07407.1 |AE004818_13 UDP-N-acetylmuramate:L-alanyl-gamma-D-glutamyl-meso-diaminopimelate ligase [Pseudomonas aeruginosa PA01] Length = 451
1886.2 Best-BlastP=> >nrprot No Hits found 1887.2 Best-BlastP=> >nrprot 37% Identities = 126/504 (25%), Positives = 206/504 (40%), Gaps = 128/504 (25%) ref|NP_905954.11 leucine-rich protein [Porphyromonas gingivalis W83] gb|AAQ66853.11 leucine-rich protein [Porphyromonas gingivalis W83] Length = 1266
1889.2 Best-BlastP=> >nrprot 30% Identities = 64/297 (21 %), Positives = 122/297 (41 %), Gaps = 33/297 (11 %) ref|NP_764683.11 ebhA protein [Staphylococcus epidermidis ATCC 12228] gb|AAO04725.1 |AE016747_222 ebhA protein [Staphylococcus epidermidis ATCC 12228] Length = 9439 189.3 Best-BlastP=> >nrprot 81 % Identities = 272/394 (69%), Positives = 330/394 (83%) dbj|BAB56449.11 NAD+-dependent formate dehydrogenase [Hyphomicrobium sp. JC17] Length = 399
1891.2 Best-BlastP=> >nrprot 47% Identities = 72/239 (30%), Positives = 123/239 (51 %) ref|NP_812276.11 hypothetical protein [Bacteroides thetaiotaomicron VPI-5482] gb|AAO78470.11 hypothetical protein [Bacteroides thetaiotaomicron VPI-5482] Length = 247
1893.2 Best-BlastP=> >nrprot 64% Identities = 98/218 (44%), Positives = 142/218 (65%), Gaps = 2/218 (0%) ref |NP_820819.11 ribosomal 5S rRNA E- loop binding protein Ctc/L2δ/TLδ [Coxiella burnetii RSA 493] gb|AA091333.11 ribosomal 6S rRNA E-loop binding protein Ctc/L25/TLδ [Coxiella burnetii RSA 4 3] Length = 244
1894.2 Best-BlastP=> >nrprot 73% Identities = 63/101 (62%), Positives = 76/101 (75%) ref|ZP_00086202.1 | COG0261 : Ribosomal protein L21 [Pseudomonas fluorescens PfO-1] Length = 103
1895.6 Best-BlastP=> >nrprot 47% Identities = 59/173 (34%), Positives = 100/173 (57%), Gaps = 14/173 (8%) ref|NP_842310.1 | putative type 4 fimbrial biogenesis protein PilP [Nitrosomonas europaea ATCC ig718] emb|CAD86225.11 putative type 4 fimbrial biogenesis protein PilP [Nitrosomonas europaea ATCC 19718] Length = 176
1896.5 Best-BlastP=> >nrprot 61 % Identities = 282/695 (40%), Positives = 430/695 (61 %), Gaps = 40/695 (5%) ref|NP_715925.11 type IV pilus biogenesis protein PilQ [Shewanella oneidensis MR-1] gb|AAN53370.1 |AE016476 11 type IV pilus biogenesis protein PilQ [Shewanella oneidensis MR-1] Length = 684
19*1 Best-BlastP=> >nrprot 37% Identities = 48/174 (27%), Positives = 80/174 (45%), Gaps = 14/174 (8%) ref|NP_820400.1 | peptidase, family S24 [Coxiella burnetii RSA 493] gb|AAO90914.11 peptidase, family S24 [Coxiella burnetii RSA 493] Length = 216
190.3 Best-BlastP=> >nrprot No Hits found
1902.5 Best-BlastP=> >nrprot 79% Identities = 205/338 (60%), Positives = 264/338 (78%), Gaps = 5/338 (1 %) ref|ZP_00086819.11 COG0533: Metal- dependent proteases with possible chaperone activity [Pseudomonas fluorescens PfO-1] Length = 341
1903.4 Best-BlastP=> >nrprot 68% Identities = 119/287 (41 %), Positives = 161/287 (56%), Gaps = 14/287 (4%) ref|NP_821016.11 chitinase domain protein [Coxiella burnetii RSA 493] gb|AA091530.11 chitinase domain protein [Coxiella burnetii RSA 493] Length = 593
1906.2 Best-BlastP=> >nrprot 77% Identities = 248/422 (68%), Positives = 328/422 (77%), Gaps = 1/422 (0%) ref|NP_718465.11 conserved hypothetical protein [Shewanella oneidensis MR-1] sp|P59352|YS83_SHEON Hypothetical UPF0229 protein S02883 gb|AAN55899.1 |AE015726_7 conserved hypothetical protein [Shewanella oneidensis MR-1] Length = 422 '
1906.2 Best-BlastP=> >nrprot 83% Identities = 347/504 (68%), Positives = 425/604 (84%), Gaps = 1/504 (0%) ref|NP_790392.11 SpoVR like family protein [Pseudomonas syringae pv. tomato str. DC3000] gb|AAO54087.11 SpoVR like family protein [Pseudomonas syringae pv. tomato str. DC3000] Length = 520
1908.4 Best-BlastP=> >nrprot 60% Identities = 348/811 (42%), Positives = 495/811 (61 %), Gaps = 14/811 (1 %) emb|CAD58321.1 | Long chain acyl-CoA dehydrogenase [Azoarcus sp. EbN1] Length = 829
191.1 Best-BlastP=> >nrprot 50% Identities = 76/297 (25%), Positives = 144/297 (48%), Gaps = 33/297 (11 %) ref|NP_832129.11 Transcriptional regulators, LysR family [Bacillus cereus ATCC 14579] gb|AAP09330.11 Transcriptional regulators, LysR family [Bacillus cereus ATCC 1457g] Length = 300
1910.6 Best-BlastP=> >nrprot 12% Identities = 92/441 (20%), Positives = 185/441 (41 %), Gaps = 64/441 (14%) gb|AAB70839.1 | ZipA [Dictyostelium discoideum] Length = 924
1911.4 Best-BlastP=> >nrprot 41 % Identities = 37/110 (33%), Positives = 64/110 (58%), Gaps = 6/110 (5%) ref|ZP_00054083.11 COG0664: cAMP- binding proteins - catabolite gene activator and regulatory subunit of cAMP-dependent protein kinases [Magnetospirillum magnetotacticum] Length = 282
1913.2 Best-BlastP=> >nrprot 61% Identities = 41/91 (45%), Positives = 61/91 (67%) ref|NP_819252.11 hypothetical protein [Coxiella burnetii RSA 493] gb|AA089766.11 hypothetical protein [Coxiella burnetii RSA 493] Length = 130
1915.3 Best-BlastP=> >nrprot 56% Identities = 237/625 (37%), Positives = 364/625 (58%), Gaps = 18/625 (2%) ref|NP_518200.11 PROBABLE TRANSMEMBRANE PROTEIN [Ralstonia solanacearum] emb|CAD13607.11 PROBABLE TRANSMEMBRANE PROTEIN [Ralstonia solanacearum] Length = 795
1918.3 Best-BlastP=> >nrprot 74% Identities = 142/226 (62%), Positives = 176/226 (77%), Gaps = 1/226 (0%) ref|NP_384333.1 | PUTATIVE TRANSCRIPTION REGULATOR PROTEIN [Sinorhizobium meliloti] emb|CAC41614.1 | PUTATIVE TRANSCRIPTION REGULATOR PROTEIN [Sinorhizobium meliloti] Length = 230
1920.3 Best-BlastP=> >nrprot 66% Identities = 198/345 (57%), Positives = 239/346 (69%), Gaps = 9/345 (2%) emb|CAB82454.11 CnrT protein [Ralstonia metallidurans] Length = 361
1923.2 Best-BlastP=> >nrprot No Hits found
1924.4 Best-BlastP=> >nrprot 62% Identities = 75/166 (45%), Positives = 106/165 (64%), Gaps = 6/165 (3%) ref|NP_840350.11 putative antirestriction protein [Nitrosomonas europaea ATCC 19718] emb|CAD84171.11 putative antirestriction protein [Nitrosomonas europaea ATCC 19718] Length = 171
1926.2 Best-BlastP=> >nrprot 71 % Identities = 121/188 (64%), Positives = 150/188 (79%), Gaps = 1/188 (0%) ref|NP_405720.1 | thymidine kinase [Yersinia pestis] ref|NP_669456.11 thymidine kinase [Yersinia pestis KIM] sp|Q8ZEJ1 |KITH_YERPE Thymidine kinase pir||AD0265 thymidine kinase (EC 2.7.1.21 ) [similarity] - Yersinia pestis (strain C092) emb|CAC90984.11 thymidine kinase [Yersinia pestis C092] gb|AAM85707.1 |AE013818_1 thymidine kinase [Yersinia pestis KIM] Length = 196
1928.2 Best-BlastP=> >nrprot 60% Identities = 181/408 (44%), Positives = 254/408 (62%), Gaps = 8/408 (1 %) ref|NP_821038.11 major facilitator family transporter [Coxiella burnetii RSA 493] gb|AA091552.11 major facilitator family transporter [Coxiella burnetii RSA 493] Length = 446
193.3 Best-BlastP=> >nrprot 56% Identities = 132/320 (41 %), Positives = 199/320 (62%), Gaps = 17/320 (5%) gb|AAC21671.1 | PvcA [Pseudomonas aeruginosa] Length = 327
1930.3 Best-BlastP=> >nrprot No Hits found
1933.4 Best-BlastP=> >nrprot 14% Identities = 109/524 (20%), Positives = 228/524 (43%), Gaps = 74/524 (14%) ref|NP_010225.11 involved intracellular protein transport, coiled-coil protein necessary for protein transport from ER to Golgi; Uso1 p [Saccharomyces cerevisiae] pir||S67593 transport protein US01 - yeast (Saccharomyces cerevisiae) emb|CAA98621.1 | US01 [Saccharomyces cerevisiae] Length = 1790
1934.4 Best-BlastP=> >nrprot 97% Identities = 310/333 (93%), Positives = 324/333 (97%) emb|CAB65198.11 hypothetical protein [Legionella pneumophila] Length = 333
1935.4 Best-BlastP=> >nrprot 96% Identities = 348/372 (93%), Positives = 360/372 (96%), Gaps = 3/372 (0%) emb|CAB65199.11 hypothetical protein [Legionella pneumophila] Length = 369
1937.4 Best-BlastP=> >nrprot No Hits found
1939.5 Best-BlastP=> >nrprot No Hits found 1940.2 Best-BlastP=> >nrprot 53% Identities = 153/524 (29%), Positives = 286/524 (54%), Gaps = 30/524 (5%) ref|NP_922967.11 HlyB/MsbA family ABC transporter [Gloeobacter violaceus] dbj|BAC87962.11 HlyB/MsbA family ABC transporter [Gloeobacter violaceus] Length = 605
1943.3 Best-BlastP=> >nrprot 55% Identities = 136/303 (44%), Positives = 183/303 (60%), Gaps = 3/303 (0%) ref|ZP_00095364.11 COG0845: Membrane-fusion protein [Novosphingobium aromaticivorans] Length = 371
1945.4 Best-BlastP=> >nrprot 80% Identities = 408/591 (69%), Positives = 473/591 (80%), Gaps = 5/591 (0%) ref|NP_706511.11 succinate dehydrogenase flavoprotein subunit [Shigella flexneri 2a str. 301] gb|AAN42218.1 |AE015088_δ succinate dehydrogenase flavoprotein subunit [Shigella flexneri 2a str. 301] Length = 692
1946.2 Best-BlastP=> >nrprot 78% Identities = 156/227 (68%), Positives = 188/227 (82%), Gaps = 1/227 (0%) emb|CAA74088.1 | succinate dehydrogenase putative iron sulphur subunit [Shewanella frigidimarina] Length = 235
1947.3 Best-BlastP=> >nrprot 45% Identities = 76/274 (27%), Positives = 133/274 (48%), Gaps = 12/274 (4%) ref|ZP_00069289.11 COG1577: Mevalonate kinase [Oenococcus oeni MCW] Length = 306
1949.4 Best-BlastP=> >nrprot 67% Identities = 38/83 (45%), Positives = 61/83 (73%) ref|NP_252324.11 conserved hypothetical protein [Pseudomonas aeruginosa PA01] pir||G83191 conserved hypothetical protein PA3634 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG07022.1 |AE004783_7 conserved hypothetical protein [Pseudomonas aeruginosa PA01] Length = 4
195.3 Best-BlastP=> >nrprot 79% Identities = 181/278 (65%), Positives = 223/278 (80%) ref|NP_231578.11 PvcB protein [Vibrio cholerae 01 biovar eltor str. N 16961] pir||B82137 PvcB protein VC1944 [imported] - Vibrio cholerae (strain N16961 serogroup 01 ) gb|AAF95092.1 | PvcB protein [Vibrio cholerae 01 biovar eltor str. N16961] Length = 287
1950.2 Best-BlastP=> >nrprot 55% Identities = 159/405 (39%), Positives = 239/405 (69%), Gaps = 13/405 (3%) ref|NP_820347.11 conserved hypothetical protein [Coxiella burnetii RSA 493] gb|AAO90861.1 | conserved hypothetical protein [Coxiella burnetii RSA 493] Length = 426
1951.4 Best-BlastP=> >nrprot 81 % Identities = 170/245 (69%), Positives = 205/245 (83%) ref|NP_924317.11 probable ABC transporter ATP-binding protein [Gloeobacter violaceus] dbj|BAC89312.1 | girl 371 [Gloeobacter violaceus] Length = 252
1965.3 Best-BlastP=> >nrprot 46% Identities = 141/475 (29%), Positives = 221/475 (46%), Gaps = 64/475 (13%) ref |ZP_00015335.11 COG2067: Long- chain fatty acid transport protein [Rhodospirillum rubrum] Length = 436
1960.2 Best-BlastP=> >nrprot No Hits found
1961.3 Best-BlastP=> >nrprot 96% Identities = 283/293 (96%), Positives = 284/293 (96%) gb|AAM00623.11 unknown [Legionella pneumophila] Length = 293
1966.2 Best-BlastP=> >nrprot 84% Identities = 131/188 (69%), Positives = 160/188 (85%) ref|NP_820795.11 translation elongation factor P [Coxiella burnetii RSA 493] gb|AA091309.11 translation elongation factor P [Coxiella burnetii RSA 4 3] Length = 188
1968.1 Best-BlastP=> >nrprot 41 % Identities = 25/64 (39%), Positives = 32/64 (50%), Gaps = 13/64 (20%) ref|ZP_00011706.11 hypothetical protein [Rhodopseudomonas palustris] Length = 150
197.3 Best-BlastP=> >nrprot 80% Identities = 324/474 (68%), Positives = 389/474 (82%) ref|NP_231579.11 FAD monooxygenase, PheA/TfdB family [Vibrio cholerae 01 biovar eltor str. N16961] pir||C82137 FAD monooxygenase, PheA/TfdB family VC1945 [imported] - Vibrio cholerae (strain N16961 serogroup 01 ) gb|AAF95093.11 FAD monooxygenase, PheATfdB family [Vibrio cholerae 01 biovar eltor str. N16961] Length = 487
1972.3 Best-BlastP=> >nrprot 62% Identities = 158/342 (46%), Positives = 216/342 (63%), Gaps = 3/342 (0%) gb|AAP58486.11 putative phosphoribosylformylglycinamidine cyclo ligase [uncultured Acidobacteria bacterium] Length = 343
1976.2 ■ Best-BlastP=> >nrprot 79% Identities = 427/626 (68%), Positives = 497/626 (79%), Gaps = 6/626 (0%) ref|NP_720274.11 glucose-inhibited division protein A [Shewanella oneidensis MR-1] gb|AAN57717.1 |AE016908_2 glucose-inhibited division protein A [Shewanella oneidensis MR-1] Length = 629 1978.1 Best-BlastP=> >nrprot 63% Identities = 94/201 (46%), Positives = 133/201 (66%), Gaps = 6/201 (2%) ref|NP_246425.1 | GidB [Pasteurella multocida] sp|P67946|GIDB_PASMU Methyltransferase gidB (Glucose inhibited division protein B) gb|AAK03570.1 | GidB [Pasteurella multocida] Length = 210
1979.1 Best-BlastP=> >nrprot 79% Identities = 163/254 (64%), Positives = 204/254 (80%) ref|NP_820903.11 sporulation initiation inhibitor protein soj [Coxiella burnetii RSA 493] gb|AA091417.11 sporulation initiation inhibitor protein soj [Coxiella burnetii RSA 493] Length = 256
1983.1 Best-BlastP=> >nrprot 63% Identities = 138/265 (52%), Positives = 175/265 (66%), Gaps = 1/265 (0%) ref|NP_820962.1 | bis(5'-nucleosyl)- tetraphosphatase, symmetrical [Coxiella burnetii RSA 493] sp|Q83AB7|APAH_COXBU Bis(δ'-nucleosyl)-tetraphosphatase, symmetrical (Diadenosine tetraphosphatase) (Ap4A hydrolase) (Diadenosine δ',δ'"-P1 ,P4-tetraphosphate pyrophosphohydrolase) gb|AA091476.1 | bis(5'-nucleosyl)-tetraphosphatase, symmetrical [Coxiella burnetii RSA 493] Length = 291
1986.2 Best-BlastP=> >nrprot 24% Identities = 56/235 (23%), Positives = 111/235 (47%), Gaps = 11/235 (4%) gb|AAP84130.11 putative pathogenesis- related protein [Pseudomonas aeruginosa] Length = 639
1989.2 Best-BlastP=> >nrprot 70% Identities = 263/481 (54%), Positives = 348/481 (72%), Gaps = 1/481 (0%) sp|P37986|G6PD_ERWCH Glucose-6- phosphate 1 -dehydrogenase (G6PD) pir||S37053 glucose-6-phosphate 1 -dehydrogenase (EC 1.1.1.49) - Erwinia chrysanthemi emb|CAA52858.11 glucose-6-phosphate 1 -dehydrogenase [Erwinia chrysanthemi] Length = 491
199.1 Best-BlastP=> >nrprot 45% Identities = 105/371 (28%), Positives = 173/371 (46%), Gaps = 13/371 (3%) ref|ZP_00014611.1 | COG2814: Arabinose efflux permease [Rhodospirillum rubrum] Length = 411
1990.1 Best-BlastP=> >nrprot 55% Identities = 102/228 (44%), Positives = 136/228 (59%), Gaps = 2/228 (0%) gb|AAL76390.1 | 6- phosphogluconolactonase [uncultured proteobacterium] Length = 226
1992.2 Best-BlastP=> >nrprot No Hits found
1998.3 Best-BlastP=> >nrprot 51% Identities = 119/265 (44%), Positives = 170/265 (64%), Gaps = 9/265 (3%) ref|ZP_00089281.1 | COG1295: Predicted membrane protein [Azotobacter vinelandii] Length = 408
2.1 Best-BlastP=> >nrprot 59% Identities = 70/170 (41 %), Positives = 102/170 (60%), Gaps = 2/170 (1 %) ref|NP_907750.1 | HYPOTHETICAL PROTEIN-RecB family exonuclease [Wolinella succinogenes] emb|CAE10650.11 HYPOTHETICAL PROTEIN-RecB family exonuclease [Wolinella succinogenes] Length = i 3
20.1 Best-BlastP=> >nrprot No Hits found
2000.1 Best-BlastP=> >nrprot 99% Identities = 149/149 (100%), Positives = 149/149 (100%) gb|AAC38305.11 type IV pilin; competence and adherence associated pilin; CAP [Legionella pneumophila] Length = 149
2006.2 Best-BlastP=> >nrprot 46% Identities = 64/229 (27%), Positives = 102/229 (44%), Gaps = 27/229 (11 %) ref|NP_762597.11 Conserved hypothetical protein [Vibrio vulnificus CMCP6] gb|AAO07587.1 |AE016810_90 Conserved hypothetical protein [Vibrio vulnificus CMCP6] Length = 232
2007.1 Best-BlastP=> >nrprot 72% Identities = 173/298 (58%), Positives = 220/298 (73%) ref|NP_792548.11 hydroxymethylglutaryl-CoA lyase [Pseudomonas syringae pv. tomato str. DC3000] gb|AA056243.11 hydroxymethylglutaryl-CoA lyase [Pseudomonas syringae pv. tomato str.-DC3000] Length = 2g9
201.2 Best-BlastP=> >nrprot 50% Identities = 139/407 (34%), Positives = 209/407 (51 %), Gaps = 9/407 (2%) dbj|BAB69410.11 hypothetical protein [Streptomyces avermitilis] Length = 468
2016.2 Best-BlastP=> >nrprot 64% Identities = 54/141 (38%), Positives = 90/141 (63%), Gaps = 4/141 (2%) ref|NP_820504.11 rhodanese domain protein [Coxiella burnetii RSA 493] gb|AAO91018.11 rhodanese domain protein [Coxiella burnetii RSA 493] Length = 144
2017.1 Best-BlastP=> >nrprot 52% Identities = 70/179 (39%), Positives = 99/179 (55%), Gaps = 7/179 (3%) ref|ZP_00124407.11 COG2840: Uncharacterized protein conserved in bacteria [Pseudomonas syringae pv. syringae B728a] Length = 185
2019.2 Best-BlastP=> >nrprot 61% Identities = 112/266 (42%), Positives = 153/266 (57%), Gaps = 28/266 (10%) ref |NP_253119.11 probable cytochrome d precursor [Pseudomonas aeruginosa PA01] pir||E83092 probable cytochrome d precursor PA4429 [imported] - Pseudomonas aeruginosa (strain PA01) gb|AAG07817.1 |AE004857_8 probable cytochrome d precursor [Pseudomonas aeruginosa PA01] Length = 260
202.3 Best-BlastP=> >nrprot 57% Identities = 299/765 (39%), Positives = 444/765 (58%), Gaps = 17/765 (2%) ref|NP_519533.11 PUTATIVE OUTER MEMBRANE SIGNAL PEPTIDE PROTEIN [Ralstonia solanacearum] emb|CAD15114.1 | PUTATIVE OUTER MEMBRANE SIGNAL PEPTIDE PROTEIN [Ralstonia solanacearum] Length = 765
T-; T-; - CN -; CN C CM
O CN t O r^ OO O -i-1 ■^ r-' oo '^ -i-^ rt
CN CN CN CN CM CM CN CO CO CO CO CO rt rt rt
O O O O O O O O O O O O O O O
CM CM CN CN CM CM CN CM CN CN CN C CM CN CM
2049.2 Best-BlastP=> >nrprot 26% Identities = 57/288 (19%), Positives = 112/288 (38%), Gaps = 32/288 (11 %) pir||T13030 microtubule binding protein D-CLIP-190 - fruit fly (Drosophila melanogaster) gb|AAB96783.1 | microtubule binding protein D-CLIP-190 [Drosophila melanogaster] Length = 1690 205.1 Best-BlastP=> >nrprot 59% Identities = 103/267 (38%), Positives = 158/267 (59%), Gaps = 15/267 (5%) ref|NP_820370.1 | phosphatidate cytidylyltransferase [Coxiella burnetii RSA 493] gb|AAO90884.11 phosphatidate cytidylyltransferase [Coxiella burnetii RSA 493] Length = 272
2051.2 Best-BlastP=> >nrprot 37% Identities = 34/108 (31 %), Positives = 63/108 (58%), Gaps = 2/108 (1 %) ref|NP_932218.11 putative conjugative transfer protein TrbB [Vibrio vulnificus YJ016] dbj|BAC97741.11 putative conjugative transfer protein TrbB [Vibrio vulnificus YJ016] Length = 137 2053.2 Best-BlastP=> >nrprot 65% Identities = 146/331 (44%), Positives = 180/331 (54%), Gaps = 57/331 (17%) gb|AAC83331.11 major outer membrane protein precursor [Legionella pneumophila] Length = 288
2054.2 Best-BlastP=> >nrprot 54% Identities = 128/2g6 (43%), Positives = 184/296 (62%), Gaps = 17/296 (5%) ref|NP_442648.11 unknown protein [Synechocystis sp. PCC 6803] pir||S76674 hypothetical protein - Synechocystis sp. (strain PCC 6803) dbj|BAA10618.1 | slr0619 [Synechocystis sp. PCC 6803] Length = 348
2066.1 Best-BlastP=> >nrprot 56% Identities = 109/292 (37%), Positives = 165/292 (56%), Gaps = 8/292 (2%) ref|ZP_00134420.11 COG0500: SAM- dependent methyltransferases [Actinobacillus pleuropneumoniae serovar 1 str. 4074] Length = 290
2057.3 Best-BlastP=> >nrprot 59% Identities = 145/360 (41 %), Positives = 208/350 (59%), Gaps = 17/350 (4%) ref|ZP_00097544.11 COG0722: 3-deoxy- D-arabino-heptulosonate 7-phosphate (DAHP) synthase [Desulfitobacterium hafniense] Length = 342
206.3 Best-BlastP=> >nrprot 66% Identities = 120/227 (52%), Positives = 150/227 (66%) ref|NP_252342.11 undecaprenyl pyrophosphate synthetase [Pseudomonas aeruginosa PA01] pir||G83188 undecaprenyl pyrophosphate synthetase PA3652 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG07040.1 |AE004785_4 undecaprenyl pyrophosphate synthetase [Pseudomonas aeruginosa PA01] Length = 251
2060.3 Best-BlastP=> >nrprot 32% Identities = 71/221 (32%), Positives = 123/221 (55%), Gaps = 24/221 (10%) ref|ZP_00082359.11 COG219g: FOG: GGDEF domain [Geobacter metallireducens] Length = 363
2062.2 Best-BlastP=> >nrprot No Hits found
2064.3 Best-BlastP=> >nrprot No Hits found 2065.2 Best-BlastP=> >nrprot No Hits found 2066.5 Best-BlastP=> >nrprot No Hits found 2067.5 Best-BlastP=> >nrprot 61 % Identities = 151/349 (43%), Positives = 215/349 (61 %), Gaps = 19/349 (5%) ref|NP_660996.11 glycosyl hydrolase, family 3 [Chlorobium tepidum TLS] gb|AAM71338.11 glycosyl hydrolase, family 3 [Chlorobium tepidum TLS] Length = 372 2068.3 Best-BlastP=> >nrprot 27% Identities = 71/336 (21%), Positives = 133/336 (39%), Gaps = 67/336 (19%) emb|CAE02882.1 | OSJNBb0022F23.19 [Oryza sativa (japonica cultivar-group)] Length = 2391
207.3 Best-BlastP=> >nrprot 98% Identities = 418/423 (98%), Positives = 419/423 (99%) gb|AAM73854.1 |AF454865_1 putative phospholipase C [Legionella pneumophila] Length = 423
2070.4 Best-BlastP=> >nrprot 49% Identities = 166/568 (29%), Positives = 274/568 (48%), Gaps = 46/568 (8%) ref|NP_251765.11 hypothetical protein [Pseudomonas aeruginosa PA01] ref|ZP_00136432.11 hypothetical protein [Pseudomonas aeruginosa UCBPP-PA14] pir||D83262 hypothetical protein PA3076 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG06463.1 |AE004731_11 hypothetical protein PA3076 [Pseudomonas aeruginosa PA01] Length = 543
2073.4 Best-BlastP=> >nrprot 37% Identities = 49/192 (25%), Positives = 86/192 (44%), Gaps = 23/192 (11 %) ref|NP_929249.1 | hypothetical protein [Photorhabdus luminescens subsp. laumondii TT01] emb|CAE14276.1 | unnamed protein product [Photorhabdus luminescens subsp. laumondii TT01] Length = 231
2075.1 Best-BlastP=> >nrprot No Hits found
2078.2 Best-BlastP=> >nrprot No Hits found
2079.2 Best-BlastP=> >nrprot No Hits found
2080.3 Best-BlastP=> >nrprot 57% Identities = 88/221 (39%), Positives = 128/221 (57%), Gaps = 1/221 (0%) ref|NP_925444.11 hypothetical protein gll2498 [Gloeobacter violaceus] dbj|BAC90439.11 gll2498 [Gloeobacter violaceus] Length = 222
2082.2 Best-BlastP=> >nrprot 76% Identities = 34/66 (51 %), Positives = 51/66 (77%) ref|NP_768003.11 bsM 363 [Bradyrhizobium japonicum] dbj|BAC46628.1 | bsl1363 [Bradyrhizobium japonicum USDA 110] Length = 73
2083.2 Best-BlastP=> >nrprot 98% Identities = 224/227 (98%), Positives = 226/227 (99%) gb|AAM00399.1 |AF386079_9 CcmH [Legionella pneumophila] Length = 360
2085.2 Best-BlastP=> >nrprot 99% Identities = 132/133 (99%), Positives = 133/133 (100%) gb|AAM00399.1 |AF386079_9 CcmH [Legionella pneumophila] Length = 360
2087.2 Best-BlastP=> >nrprot 36% Identities = 88/166 (53%), Positives = 116/166 (69%) ref|NP_819420.11 conserved hypothetical protein [Coxiella burnetii RSA 493] gb|AA089934.11 conserved hypothetical protein [Coxiella burnetii RSA 493] Length = 176
209.2 Best-BlastP=> >nrprot 24% Identities = 41/135 (30%), Positives = 61/135 (45%), Gaps = 36/135 (26%) gb|AAO49307.11 outer surface protein precursor [Wolbachia pipientis] Length = 186
2091.1 Best-BlastP=> >nrprot 39% Identities = 60/226 (26%), Positives = 99/226 (43%), Gaps = 8/226 (3%) gb|AAK31375.1 |AC084329_1 ppg3 [Leishmania major] Length = 1325
2092.2 Best-BlastP=> >nrprot 86% Identities = 634/850 (74%), Positives = 741/850 (87%) ref|NP_457131.11 ClpB protein (heat shock protein f84.1 ) [Salmonella enterica subsp. enterica serovar Typhi] ref|NP_461591.1 | ATP-dependent protease, Hsp 100, part of novel multi-chaperone system with DnaK, DnaJ, and GrpE [Salmonella typhimurium LT2] ref|NP_806327.1 | ClpB protein [Salmonella enterica subsp. enterica serovar Typhi Ty2] pir||AI0831 ClpB protein (heat shock protein f84.1 ) [imported] - Salmonella enterica subsp. enterica serovar Typhi (strain CT18) gb|AAL21560.1 j ATP-dependent protease [Salmonella typhimurium LT2] emb|CAD05840.1 | ClpB protein (heat shock protein f84.1 ) [Salmonella enterica subsp. enterica serovar Typhi] gb|AAO70187.11 ClpB protein [Salmonella enterica subsp. enterica serovar Typhi Ty2] Length = 867
2096.2 Best-BlastP=> >nrprot 72% Identities = 218/393 (65%), Positives = 285/393 (72%), Gaps = 1/393 (0%) ref|ZP_00125180.11 COG2081 : Predicted flavoproteins [Pseudomonas syringae pv. syringae B728a] Length = 392
2098.2 Best-BlastP=> >nrprot No Hits found
2103.2 Best-BlastP=> >nrprot 69% Identities = 424/791 (53%), Positives = 552/791 (69%), Gaps = 3/791 (0%) ref|NP_637344.11 3-hydroxyacyl-CoA dehydrogenase [Xanthomonas campestris pv. campestris str. ATCC 33913] gb|AAM41268.1 | 3-hydroxyacyl-CoA dehydrogenase [Xanthomonas campestris pv. campestris str. ATCC 33913] Length = 7 o
2104.1 Best-BlastP=> >nrprot 85% Identities = 91/127 (71 %), Positives = 108/127 (85%), Gaps = 1/127 (0%) ref|NP_878931.1 | 50S ribosomal protein L7/L12 [Bordetella pertussis] ref|NP_882379.1 | 50S ribosomal protein L7/L12 [Bordetella parapertussis] ref|NP_886566.1 | 50S ribosomal protein L7/L12 [Bordetella bronchiseptica] emb|CAE39754.1 | 60S ribosomal protein L7/L12 [Bordetella parapertussis] emb|CAE40393.1 | 50S ribosomal protein L7/L12 [Bordetella pertussis] emb|CAE3051 δ.1 | 50S ribosomal protein L7/L12 [Bordetella bronchiseptica] Length = 127
2106.2 Best-BlastP=> >nrprot 73% Identities = 89/173 (51 %), Positives = 131/173 (75%) ref | NP_819272.1 | ribosomal protein L10 [Coxiella burnetii RSA 493] gb|AA089786.11 ribosomal protein L10 [Coxiella burnetii RSA 493] Length = 174
2108.2 Best-BlastP=> >nrprot 45% Identities = 81/300 (27%), Positives = 142/300 (47%), Gaps = 14/300 (4%) ref|NP_561674.11 conserved hypothetical protein [Clostridium perfringens] dbj|BAB80464.1 | conserved hypothetical protein [Clostridium perfringens str. 13] Length = 308
211.1 Best-BlastP=> >nrprot 58% Identities = 118/341 (34%), Positives = 204/341 (59%), Gaps = 13/341 (3%) ref|NP_882114.1 | putative membrane protein [Bordetella pertussis] ref|NP_882789.1 | putative membrane protein [Bordetella parapertussis] ref|NP_886988.1 | putative membrane protein [Bordetella bronchiseptica] emb|CAE43862.1 | putative membrane protein [Bordetella pertussis] emb|CAE36021.1 | putative membrane protein [Bordetella parapertussis] emb|CAE30937.1 | putative membrane protein [Bordetella bronchiseptica] Length = 367
2112.2 Best-BlastP=> >nrprot 28% Identities = 83/302 (27%), Positives = 137/302 (45%), Gaps = 15/302 (4%) ref|NP_772278.11 bll5638 [Bradyrhizobium japonicum] dbj|BAC50903.11 bllδ638 [Bradyrhizobium japonicum USDA 1 10] Length = 500
21 16.2 Best-BlastP=> >nrprot 26% Identities = 23/73 (31 %), Positives = 42/73 (57%), Gaps = 1/73 (1 %) ref|NP_391095.1 1 transcriptional regulator [Bacillus subtilis] sp|P21340|PAIA_BACSU Protease synthase and sporulation negative regulatory protein PA1 1 emb|CAB15205.11 transcriptional regulator [Bacillus subtilis subsp. subtilis str. 168] Length = 172
2119.2 Best-BlastP=> >nrprot 47% Identities = 175/527 (33%), Positives = 263/527 (49%), Gaps = 77/527 (14%) dbj|BAB86344.1 | metalloprotease [Vibrio fluvialis] Length = 610
212.1 Best-BlastP=> >nrprot 68% Identities = 83/152 (54%), Positives = 110/152 (72%), Gaps = 2/152 (1 %) ref|NP_742683.1 | phosphatidylglycerophosphatase A [Pseudomonas putida KT2440] gb|AAN66147.1 |AE016242_15 phosphatidylglycerophosphatase A [Pseudomonas putida KT2440] Length = 167
2120.2 Best-BlastP=> >nrprot No Hits found
2121.2 Best-BlastP=> >nrprot 60% Identities = 108/221 (48%), Positives = 148/221 (66%), Gaps = 1/221 (0%) ref|NP_251544.11 conserved hypothetical protein [Pseudomonas aeruginosa PA01] pir||B83288 conserved hypothetical protein PA2854 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG06242.1 |AE004712_2 conserved hypothetical protein [Pseudomonas aeruginosa PA01] Length = 323
2122.2 Best-BlastP=> >nrprot 43% Identities = 100/377 (26%), Positives = 171/377 (45%), Gaps = 29/377 (7%) ref|NP_820237.11 membrane protein, putative [Coxiella burnetii RSA 493] gb|AAO90751.11 membrane protein, putative [Coxiella burnetii RSA 493] Length = 408
2127.2 Best-BlastP=> >nrprot 62% Identities = 56/112 (50%), Positives = 74/112 (66%), Gaps = 3/112 (2%) ref|NP_840790.11 Uncharacterised protein family UPF0102 [Nitrosomonas europaea ATCC 19718] emb|CAD84622.1 | Uncharacterised protein family UPF0102 [Nitrosomonas europaea ATCC 19718] Length = 118
2129.1 Best-BlastP=> >nrprot 70% Identities = 98/192 (51%), Positives = 141/192 (73%) ref|NP_743483.11 phosphoheptose isomerase [Pseudomonas putida KT2440] gb|AAN66947.1 |AE016322_14 phosphoheptose isomerase [Pseudomonas putida KT2440] Length = 195
213.1 Best-BlastP=> >nrprot 56% Identities = 141/268 (52%), Positives = 180/268 (67%), Gaps = 4/268 (1 %) ref|NP_406655.11 thiamine- monophosphate kinase [Yersinia pestis] ref|NP_668333.1 | thiamin-monophosphate kinase [Yersinia pestis KIM] pir||AD0386 thiamine-phosphate kinase (EC 2.7.4.16) [imported] - Yersinia pestis (strain C092) emb|CAC92415.11 thiamine-monophosphate kinase [Yersinia pestis C092] gb|AAM84584.1 |AE013704_1 thiamin-monophosphate kinase [Yersinia pestis KIM] Length = 329
2130.1 Best-BlastP=> >nrprot 62% Identities = 88/188 (46%), Positives = 120/188 (63%), Gaps = 7/188 (3%) ref|NP_820724.11 conserved hypothetical protein [Coxiella burnetii RSA 493] gb|AA091238.11 conserved hypothetical protein [Coxiella burnetii RSA 493] Length = 191
2132.2 Best-BlastP=> >nrprot 47% Identities = 63/181 (29%), Positives = 96/181 (53%), Gaps = 7/181 (3%) ref|NP_715937.11 lipoprotein, putative [Shewanella oneidensis MR-1] gb|AAN53382.1 |AE015477_12 lipoprotein, putative [Shewanella oneidensis MR-1] Length = 188
2133.1 Best-BlastP=> >nrprot No Hits found
2135.1 Best-BlastP=> >nrprot 63% Identities = 55 118 (46%), Positives = 70/118 (59%), Gaps = 1/118 (0%) ref|NP_254248.11 ATP synthase protein I [Pseudomonas aeruginosa PA01] pir||B82953 ATP synthase protein I PA6561 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG08946.1 |AE004967_17 ATP synthase protein I [Pseudomonas aeruginosa PA01] Length = 126
2136.1 Best-BlastP=> >nrprot 75% Identities = 176/277 (63%), Positives = 210/277 (75%), Gaps = 19/277 (6%) ref|NP_720269.11 ATP synthase F0, A subunit [Shewanella oneidensis MR-1] gb|AAN57712.1 |AE015907_10 ATP synthase F0, A subunit [Shewanella oneidensis MR-1] Length = 264
2137.1 Best-BlastP=> >nrprot 81% Identities = 72/80 (90%), Positives = 75/80 (93%) ref |ZP_00124678.11 COG0636: F0F1-tyρe ATP synthase, subunit c/Archaeal/vacuolar-type H+-ATPase, subunit K [Pseudomonas syringae pv. syringae B728a] ref|NP_795322.1 | ATP synthase F0, C subunit [Pseudomonas syringae pv. tomato str. DC3000] gb|AAO59017.11 ATP synthase F0, C subunit [Pseudomonas syringae pv. tomato str. DC3000] Length = 85
2138.2 Best-BlastP=> >nrprot 72% Identities = 8-5 156 (54%), Positives = 114/156 (73%) ref|NP_820917.11 ATP synthase F0, B subunit [Coxiella burnetii RSA 493] gb|AA091431.1 | ATP synthase F0, B subunit [Coxiella burnetii RSA 493] Length = 156
214.1 Best-BlastP=> >nrprot 66% Identities = 61/140 (43%), Positives = 99/140 (70%) ref|NP_252741.11 NusB protein [Pseudomonas aeruginosa PA01] sp|Q9HWX6|NUSB_PSEAE N utilization substance protein B homolog (NusB protein) pir||G83140 NusB protein PA4052 [imported] - Pseudomonas aeruginosa (strain PA01) gb|AAG07439.1 |AE004821_12 NusB protein [Pseudomonas aeruginosa PA01] Length = 159
2140.2 Best-BlastP=> >nrprot 64% Identities = 86/178 (48%), Positives = 116/178 (65%), Gaps = 2/178 (1 %) ref|NP_254244.11 ATP synthase delta chain [Pseudomonas aeruginosa PA01] pir||F82952 ATP synthase delta chain PA5557 [imported] - Pseudomonas aeruginosa (strain PA01) gb|AAG08942.1 |AE004967_13 ATP synthase delta chain [Pseudomonas aeruginosa PA01] Length = 178
2141.2 Best-BlastP=> >nrprot No Hits found 2143.5 Best-BlastP=> >nrprot 21 % Identities = 66/242 (27%), Positives = 110/242 (45%), Gaps = 32/242 (13%) ref|NP_819452.11 hypothetical protein [Coxiella burnetii RSA 493] gb|AA089966.11 hypothetical protein [Coxiella burnetii RSA 493] Length = 262
2144.3 Best-BlastP=> >nrprot 59% Identities = 133/308 (43%), Positives = 196/308 (63%), Gaps = 1/308 (0%) ref|NP_716251.1 | arginine N- succinyltransferase [Shewanella oneidensis MR-1] gb|AAN53696.1 |AE015509_1 arginine N-succinyltransferase [Shewanella oneidensis MR-1] Length = 339
2146.2 Best-BlastP=> >nrprot 60% Identities = 151/363 (41 %), Positives = 224/363 (61 %), Gaps = 5/363 (1 %) ref|NP_251855.11 histidinol-phosphate aminotransferase [Pseudomonas aeruginosa PA01] sp|Q9HZ68|HI82_PSEAE Histidinol-phosphate aminotransferase 2 (Imidazole acetol- phosphate transaminase 2) pir||F83250 histidinol-phosphate aminotransferase PA3165 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG06563.1 |AE004740_6 histidinol-phosphate aminotransferase [Pseudomonas aeruginosa PA01] Length = 369
2147.1 Best-BlastP=> >nrprot 70% Identities = 58/107 (54%), Positives = 80/107 (74%), Gaps = 2/107 (1 %) ref |NP_840178.11 Pterin 4 alpha carbinolamine dehydratase [Nitrosomonas europaea ATCC 19718] emb|CAD83988.11 Pterin 4 alpha carbinolamine dehydratase [Nitrosomonas europaea ATCC 19718] Length = 113
2148.1 Best-BlastP=> >nrprot 7δ% Identities = 172/305 (66%), Positives = 230/305 (75%), Gaps = 1/306 (0%) ref |NP_820140.11 protein-export membrane protein SecF [Coxiella burnetii RSA 493] gb|AAO90654.11 protein-export membrane protein SecF [Coxiella burnetii RSA 493] Length = 304
2149.4 Best-BlastP=> >nrprot 99% Identities = 427/431 (99%), Positives = 430/431 (99%) sp|Q8RNM2|PURA_LEGPN Adenylosuccinate synthetase (IMP-aspartate ligase) (AdSS) (AMPSase) gb|AAM00648.11 adenylosuccinate synthetase [Legionella pneumophila] Length = 431
215.1 Best-BlastP=> >nrprot 7g% Identities = 97/150 (64%), Positives = 124/150 (82%) ref|ZP_00126026.11 COG1327: Predicted transcriptional regulator, consists of a Zn-ribbon and ATP-cone domains [Pseudomonas syringae pv. syringae B728a] Length = 165
2150.1 Best-BlastP=> >nrprot 39% Identities = 37/148 (25%), Positives = 64/148 (43%), Gaps = 19/148 (12%) ref|NP_799791.11 hypothetical protein VPA0281 [Vibrio parahaemolyticus RIMD 2210633] dbj|BAC61624.11 hypothetical protein [Vibrio parahaemolyticus] Length = 133
2151.2 Best-BlastP=> >nrprot 14% Identities = 39/144 (27%), Positives = 67/144 (46%), Gaps = 24/144 (16%) sp|Q9U7E0|ATRX_CAEEL Transcriptional regulator ATRX homolog (X-linked nuclear protein-1 ) gb|AAD55361.1 |AF134186_1 XNP-1 [Caenorhabditis elegans] Length = 1359
2152.2 Best-BlastP=> >nrprot 83% Identities = 127/176 (72%), Positives = 150/176 (85%) ref|NP_820460.11 antioxidant, AhpC/TSA family [Coxiella burnetii RSA 493] gb|AAO90974.11 antioxidant, AhpC/TSA family [Coxiella burnetii RSA 493] Length = 179
2153.3 Best-BlastP=> >nrprot No Hits found
2156.3 Best-BlastP=> >nrprot 45% Identities = 126/444 (28%), Positives = 197/444 (44%), Gaps = 30/444 (6%) ref|NP_436941.11 putative oxidoreductase protein [Sinorhizobium meliloti] pir||A95892 probable oxidoreductase protein SMb20415 [imported] - Sinorhizobium meliloti (strain 1021 ) magaplasmid pSymB emb|CAC48801.11 putative oxidoreductase protein [Sinorhizobium meliloti] Length = 429
2169.2 Best-BlastP=> >nrprot 76% Identities = 163/238 (68%), Positives = 191/238 (80%) ref|NP_249464.11 pyridoxal phosphate biosynthetic protein PdxJ [Pseudomonas aeruginosa PA01] sp|Q9l5Gδ|PDXJ_PSEAE Pyridoxal phosphate biosynthetic protein pdxJ (PNP synthase) pir||H83548 pyridoxal phosphate biosynthetic protein PdxJ PA0773 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG04162.1 |AE004512_δ pyridoxal phosphate biosynthetic protein PdxJ [Pseudomonas aeruginosa PA01] Length = 248
216.2 Best-BlastP=> >nrprot 83% Identities = 296/416 (71 %), Positives = 361/416 (84%), Gaps = 1/416 (0%) ref|ZP_00138159.1 | COG0112: Glycine/serine hydroxymethyltransferase [Pseudomonas aeruginosa UCBPP-PA14] Length = 421
2160.2 Best-BlastP=> >nrprot 70% Identities = 166/32g (50%), Positives = 227/329 (68%), Gaps = 15/329 (4%) ref|NP_867540.11 2-oxoglutarate ferredoxin oxidoreductase beta subunit [Pirellula sp.] emb|CAD75087.11 2-oxoglutarate ferredoxin oxidoreductase beta subunit [Pirellula sp.] Length = 353
2162.2 Best-BlastP=> >nrprot 52% Identities = 153/496 (30%), Positives = 247/496 (49%), Gaps = 41/496 (8%) ref |NP_478172.11 amidase [Nostoc sp. PCC 7120] pir||AB2530 amidase [imported] - Nostoc sp. (strain PCC 7120) plasmid pCC7120beta dbj|BAB77168.1 | amidase [Nostoc sp. PCC 7120] Length = 607
2164.2 Best-BlastP=> >nrprot 48% Identities = 91/191 (47%), Positives = 135/191 (70%), Gaps = 1/191 (0%) ref|NP_820234.11 membrane protein, putative [Coxiella burnetii RSA 493] sp|Q83C89|YC39_COXBU Hypothetical UPF0078 protein CBU1239 gb|AAO90748.1 | membrane protein, putative [Coxiella burnetii RSA 493] Length = 193
2166.2 Best-BlastP=> >nrprot No Hits found
2167.2 Best-BlastP=> >nrprot 65% Identities = 230/469 (49%), Positives = 315/469 (67%), Gaps = 5/469 (1 %) ref |NP_819701.1 | mannose-1 -phosphate guanylyltransferase/mannose-6-phosphate isomerase [Coxiella burnetii RSA 493] gb|AAO90215.1 | mannose-1 -phosphate guanylyltransferase/mannose-6-phosphate isomerase [Coxiella burnetii RSA 493] Length = 477
2169.2 Best-BlastP=> >nrprot 69% Identities = 77/136 (56%), Positives = 95/136 (69%), Gaps = 4/136 (2%) ref|ZP_00067440.1 | COG4969: Tfp pilus assembly protein, major pilin PilA [Microbulbifer degradans 2-40] Length = 164
2170.1 Best-BlastP=> >nrprot 60% Identities = 67/136 (49%), Positives = 84/136 (61 %), Gaps = 5/136 (3%) ref|ZP_00067440.11 COG4969: Tfp pilus assembly protein, major pilin PilA [Microbulbifer degradans 2-40] Length = 164
2172.2 Best-BlastP=> >nrprot 52% Identities = 47/147 (31 %), Positives = 81/147 (55%), Gaps = 1/147 (0%) ref|NP_717367.11 conserved hypothetical protein [Shewanella oneidensis MR-1] gb|AAN54811.1 |AE015620_3 conserved hypothetical protein [Shewanella oneidensis MR-1] Length = 159
2173.2 Best-BlastP=> >nrprot 52% Identities = 59/125 (47%), Positives = 75/125 (60%), Gaps = 1/125 (0%) sp|Q45292|YOUG_BACLI Hypothetical 17.3 kDa protein in GNTR 5'region pir||JC2302 oug protein - Bacillus licheniformis dbj|BAA06500.1 | hypothetical protein [Bacillus licheniformis] Length = 147
2174.1 Best-BlastP=> >nrprot 50% Identities = 37/71 (52%), Positives = 50/71 (70%) ref|NP_840482.11 DUF167 [Nitrosomonas europaea ATCC 19718] emb|CAD84306.11 DUF167 [Nitrosomonas europaea ATCC 19718] Length = 100
2175.3 Best-BlastP=> >nrprot No Hits found 2176.3 Best-BlastP=> >nrprot 52% Identities = 35/119 (29%), Positives = 64/119 (53%), Gaps = 6/1 19 (5%) ref|NP_784901.1 | unknown [Lactobacillus plantarum WCFS1] emb|CAD63748.11 unknown [Lactobacillus plantarum WCFS1] Length = 269
221.1 Best-BlastP=> >nrprot 70% Identities = 158/286 (65%), Positives = 212/285 (74%) ref|ZP_00033588.11 COG0329: Dihydrodipicolinate synthase/N-acetylneuraminate lyase [Burkholderia fungorum] Length = 2 7
2212.4 Best-BiastP=> >nrprot 51% Identities = 232/616 (37%), Positives = 344/616 (55%), Gaps = 48/616 (7%) ref|NP_762611.1 | Type IV secretory pathway, VirD4 component [Vibrio vulnificus CMCP6] gb|AAO07601.1 |AE016810_104 Type IV secretory pathway, VirD4 component [Vibrio vulnificus CMCP6] Length = 697
2213.2 Best-BlastP=> >nrprot No Hits found
2217.3 Best-BlastP=> >nrprot 66% Identities = 302/609 (49%), Positives = 411/609 (67%), Gaps = 13/609 (2%) sp|P32966|UVRC_PSEFL Excinuclease ABC subunit C gb|AAA98758.11 UVR excinuclease subunit C Length = 607
2219.2 Best-BlastP=> >nrprot 34% Identities = 41/91 (45%), Positives = 48/91 (52%) gb|AAA73346.11 [Mycobacterium tuberculosis DNA sequence, complete eds.], gene products Length = 152
222.1 Best-BlastP=> >nrprot 63% Identities = 73/149 (48%), Positives = 102/149 (68%), Gaps = 5/149 (3%) ref|NP_928633.11 hypothetical protein [Photorhabdus luminescens subsp. laumondii TT01] emb|CAE13616.1 | unnamed protein product [Photorhabdus luminescens subsp. laumondii TT01] Length = 149
2220.2 Best-BlastP=> >nrprot 71% Identities = 110/213 (51%), Positives = 156/213 (73%) ref|NP_486035.11 hypothetical protein [Nostoc sp. PCC 7120] pir||AE2055 hypothetical protein all1995 [imported] - Nostoc sp. (strain PCC 7120) dbj|BAB73694.11 ORF_ID:all1995~hypothetical protein [Nostoc sp. PCC 7120] Length = 221
2221.3 Best-BlastP=> >nrprot 57% Identities = 58/132 (43%), Positives = 84/132 (63%), Gaps = 4/132 (3%) gb|EAA20351.11 cytosol aminopeptidase [Plasmodium yoelii yoelii] Length = 612
2225.2 Best-BlastP=> >nrprot 5δ% Identities = 116/284 (40%), Positives = 167/284 (58%), Gaps = 6/284 (2%) ref|NP_g02361.11 geranyltranstransferase [Chromobacterium violaceum ATCC 12472] gb|AAQ60361.1 | geranyltranstransferase [Chromobacterium violaceum ATCC 12472] Length = 298
223.1 Best-BlastP=> >nrprot 51 % Identities = 97/266 (36%), Positives = 151/266 (56%), Gaps = 9/266 (3%) ref|ZP_00020713.11 hypothetical protein [Chloroflexus aurantiacus] Length = 278
2230.2 Best-BlastP=> >nrprot 58% Identities = 68/196 (34%), Positives = 117/196 (69%), Gaps = 2/196 (1 %) ref|NP_767278.11 Maf-like protein [Bradyrhizobium japonicum] dbj|BAC45903.11 Maf-like protein [Bradyrhizobium japonicum USDA 110] Length = 202
2231.3 Best-BlastP=> >nrprot 76% Identities = 254/407 (62%), Positives = 318/407 (78%), Gaps = 1/407 (0%) ref|ZP_00079853.11 COG0148: Enolase [Geobacter metallireducens] Length = 429
2232.1 Best-BlastP=> >nrprot 32% Identities = 41/123 (33%), Positives = 61/123 (49%), Gaps = 5/123 (4%) ref |NP_899813.11 2-dehydro-3-deoxy- phosphogluconate aldolase [Chromobacterium violaceum ATCC 12472] gb|AAQ57822.1 | 2-dehydro-3-deoxy-phosphogluconate aldolase [Chromobacterium violaceum ATCC 12472] Length = 208
2233.3 Best-BlastP=> >nrprot 63% Identities = 64/119 (53%), Positives = 83/119 (69%) ref|NP_819436.11 lipoprotein signal peptidase [Coxiella burnetii RSA 493] gb|AAO89950.11 lipoprotein signal peptidase [Coxiella burnetii RSA 493] Length = 163
2235.2 Best-BlastP=> >nrprot 57% Identities = 220/636 (34%), Positives = 371/636 (58%), Gaps = 2g/636 (4%) ref|NP_819137.11 sulfatase domain protein [Coxiella burnetii RSA 493] gb|AA089651.11 sulfatase domain protein [Coxiella burnetii RSA 493] Length = 638
2239.3 Best-BlastP=> >nrprot 45% Identities = 96/277 (34%), Positives = 149/277 (53%), Gaps = 40/277 (14%) emb|CAA75849.11 hypothetical protein [Coxiella burnetii] Length = 309
2240.2 Best-BlastP=> >nrprot 66% Identities = 75/155 (48%), Positives = 105/165 (67%) ref|ZP_00067293.11 COG1576: Uncharacterized conserved protein [Microbulbifer degradans 2-40] Length = 155
2241.2 Best-BlastP=> >nrprot 67% Identities = 51/105 (48%), Positives = 76/105 (72%) ref|ZP_00090593.11 COG0799: Uncharacterized homolog of plant lojap protein [Azotobacter vinelandii] Length = 117
2242.1 Best-BlastP=> >nrprot 47% Identities = 46/187 (24%), Positives = 89/187 (47%), Gaps = 16/187 (8%) ref|NP_820065.11 conserved hypothetical protein [Coxiella burnetii RSA 493] gb|AAO90579.11 conserved hypothetical protein [Coxiella burnetii RSA 493] Length = 184
2244.2 Best-BlastP=> >nrprot 77% Identities = 129/202 (63%), Positives = 156/202 (77%) gb|AAC33273.11 TnpR [Pseudomonas alcaligenes] Length = 309 2245.2 Best-BlastP=> >nrprot 65% Identities = 178/331 (53%), Positives = 236/331 (71 %), Gaps = 6/331 (1 %) ref|NP_842305.11 conserved hypothetical protein [Nitrosomonas europaea ATCC 19718] emb|CAD86220.1 | conserved hypothetical protein [Nitrosomonas europaea ATCC 19718] Length = 346 2247.5 Best-BlastP=> >nrprot 61 % Identities = 233/561 (42%), Positives = 342/551 (62%), Gaps = 4/561 (0%) ref|NP_369923.11 multidrug resistance ABC transporter ATP-binding protein [Rickettsia conorii] pir||F97735 hypothetical protein abcT3 [imported] - Rickettsia conorii (strain Malish 7) gb|AAL02824.11 multidrug resistance ABC transporter ATP-binding protein [Rickettsia conorii] Length = 689
2250.2 Best-BlastP=> >nrprot No Hits found
2251.3 Best-BlastP=> >nrprot No Hits found
2252.1 Best-BlastP=> >nrprot 83% Identities = 254/363 (69%), Positives = 303/363 (83%) ref|NP_231816.11 GTP-binding protein [Vibrio cholerae 01 biovar eltor str. N16961] pir||D82107 GTP-binding protein VC2185 [imported] - Vibrio cholerae (strain N16961 serogroup 01 ) gb|AAF95330.11 GTP-binding protein [Vibrio cholerae 01 biovar eltor str. N16961] Length = 383
2253.2 Best-BlastP=> >nrprot 66% Identities = 96/181 (53%), Positives = 127/181 (70%) ref|NP_283782.11 putative peptidyl-tRNA hydrolase [Neisseria meningitidis Z24gi] sp|Q9JV42|PTH_NEIMA Peptidyl-tRNA hydrolase (PTH) pir||B81948 probable aminoacyl-tRNA hydrolase (EC 3.1.1.29) NMA1004 [imported] - Neisseria meningitidis (strain Z2491 serogroup A) emb|CAB84273.11 putative peptidyl-tRNA hydrolase [Neisseria meningitidis Z24gi] Length = 192
2255 2 Best-BlastP=> >nrprot 67% Identities = 158/294 (53%), Positives = 199/294 (67%), Gaps = 4/294 (1 %) ref|NP_461016.11 ATP phosphoribosyltransferase [Salmonella typhimurium LT2] sp|P00499|HIS1_SALTY ATP phosphoribosyltransferase pir||XREBT ATP phosphoribosyltransferase (EC 2.4.2.17) [validated] - Salmonella typhimurium emb|CAA31822.1 | unnamed protein product [Salmonella typhimurium] gb|AAA27142.1 | hisG gb|AAA88614.1 | ATP phosphoribosyltransferase gb|AAA80244.1 | ATP phosphoribosyltransferase gb|AAA80247.11 ATP phosphoribosyltransferase gb|AAA80249.11 ATP phosphoribosyltransferase gb|AAA80252.11 ATP phosphoribosyltransferase gb|AAA80254.1 | ATP phosphoribosyltransferase gb|AAA80257.1 | ATP phosphoribosyltransferase gb|AAA80259.1 | ATP phosphoribosyltransferase gb|AAA80262.1 | ATP phosphoribosyltransferase gb|AAL20975.1 | ATP phosphoribosyltransferase [Salmonella typhimurium LT2] Length = 299
2268.2 Best-BlastP=> >nrprot No Hits found
226.4 Best-BlastP=> >nrprot No Hits found
2260.2 Best-BlastP=> >nrprot 65% Identities = 115/209 (55%), Positives = 147/209 (70%), Gaps = 1/209 (0%) ref|ZP_00013996.1 | COG2872: Predicted metal-dependent hydrolases related to alanyl-tRNA synthetase HxxxH domain [Rhodospirillum rubrum] Length = 214
2261.3 Best-BlastP=> >nrprot No Hits found 2264.2 Best-BlastP=> >nrprot 59% Identities = 172/411 (41 %), Positives = 247/411 (60%), Gaps = 6/411 (1 %) ref|NP_773878.11 blr7238 [Bradyrhizobium japonicum] dbj|BAC52503.11 blr7238 [Bradyrhizobium japonicum USDA 110] Length = 412
2266.2 Best-BlastP=> >nrprot 76% Identities = 282/410 (68%), Positives = 322/410 (78%), Gaps = 1/410 (0%) ref|ZP_00021755.11 hypothetical protein [Ralstonia metallidurans] Length = 412
2268.2 Best-BlastP=> >nrprot 48% Identities = 131/405 (32%), Positives = 197/405 (48%), Gaps = 29/405 (7%) ref|NP_616925.11 conserved hypothetical protein [Methanosarcina acetivorans str. C2A] gb|AAM0540δ.1 | conserved hypothetical protein [Methanosarcina acetivorans str. C2A] Length = 417
227.2 Best-BlastP=> >nrprot 96% Identities = 281/303 (92%), Positives = 291/303 (96%) gb|AAM00625.11 unknown [Legionella pneumophila] Length = 303
2270.2 Best-BlastP=> >nrprot 45% Identities = 38/120 (31 %), Positives = 58/120 (48%), Gaps = 22/120 (18%) gb|AAA89101.1 | protein kinase Length = 379
2271.1 Best-BlastP=> >nrprot 41 % Identities = 2δ/δ7 (43%), Positives = 35/67 (61 %), Gaps = 4/57 (7%) ref|NP_012348.11 Delays the onset of mitosis by phosphorylation and inactivation of the cyclin-dependent kinase Cdc28, thereby relaying the morphogenetic signal to the cell cycle. S. pombe wee1+ homolog; Swelp [Saccharomyces cerevisiae] sp|P32944|SWE1_YEAST Mitosis inhibitor protein kinase SWE1 pir||S40400 protein kinase SWE1 (EC 2.7.1.-) - yeast (Saccharomyces cerevisiae) emb|CAA52150.1 | SWE1 [Saccharomyces cerevisiae] emb|CAA89482.11 SWE1 [Saccharomyces cerevisiae] Length = 819
2272.3 Best-BlastP=> >nrprot 61 % Identities = 228/487 (46%), Positives = 308/487 (63%), Gaps = 8/487 (1 %) ref|NP_744150.11 amidophosphoribosyltransferase [Pseudomonas putida KT2440] gb|AAN67614.1 |AE016391_5 amidophosphoribosyltransferase [Pseudomonas putida KT2440] Length = 501
2274.2 Best-BlastP=> >nrprot 74% Identities = 194/313 (61 %), Positives = 233/313 (74%), Gaps = 5/313 (1 %) gb|AAL85973.11 putative phosphoribosyamidoimidazole-succinocarboxamide synthase [Arabidopsis thaliana] Length = 374
2275.3 Best-BlastP=> >nrprot 34% Identities = 75/185 (40%), Positives = 107/185 (57%), Gaps = 19/185 (10%) ref|NP_437235.11 putative protein, similar to gene related to biosynthesis of peptide antibiotic trifolitoxin [Sinorhizobium meliloti] pir||G95928 hypothetical protein SMb21116 [imported] - Sinorhizobium meliloti (strain 1021 ) magaplasmid pSymB emb|CAC4909δ.1 | putative protein, similar to gene related to biosynthesis of peptide antibiotic trifolitoxin [Sinorhizobium meliloti] Length = 243
2277.2 Best-BlastP=> >nrprot 47% Identities = 27/65 (49%), Positives = 41/55 (74%), Gaps = 1/55 (1 %) ref|NP_820051.11 carbon storage regulator [Coxiella burnetii RSA 493] gb|AAO90δ65.11 carbon storage regulator [Coxiella burnetii RSA 493] Length = 70
2279.2 Best-BlastP=> >nrprot No Hits found
228.1 Best-BlastP=> >nrprot No Hits found
2282.4 Best-BlastP=> >nrprot 98% Identities = 669/577 (98%), Positives = 571/577 (98%), Gaps = 1/577 (0%) sp|P71481 |PRIM_LEGPN DNA primase gb|AAB09542.11 LpdnaG Length = 676
2289.2 Best-BlastP=> >nrprot 50% Identities = 50/159 (31 %), Positives = 81/159 (50%), Gaps = 13/159 (8%) ref |NP_718655.11 hypothetical protein [Shewanella oneidensis MR-1] gb|AAN56099.1 |AE015746_3 hypothetical protein [Shewanella oneidensis MR-1] Length = 197
2290.1 Best-BlastP=> >nrprot 53% Identities = 106/293 (36%), Positives = 164/293 (55%), Gaps = 2/293 (0%) ref|NP_744181.11 conserved hypothetical protein [Pseudomonas putida KT2440] gb|AAN67645.1 |AE016394_6 conserved hypothetical protein [Pseudomonas putida KT2440] Length = 318
2291.2 Best-BlastP=> >nrprot 81 % Identities = 208/316 (66%), Positives = 262/316 (82%), Gaps = 1/316 (0%) ref|ZP_00067583.1 | COG0714: MoxR- like ATPases [Microbulbifer degradans 2-40] Length = 321
2292.2 Best-BlastP=> >nrprot 28% Identities = 81/371 (21 %), Positives = 168/371 (42%), Gaps = 34/371 (9%) prf||2210342A myosin:SUBUNIT=heavy chain Length = 2241
2293.5 Best-BlastP=> >nrprot 14% Identities = 74/228 (32%), Positives = 119/228 (52%), Gaps = 19/228 (8%) ref|NP_844638.11 conserved hypothetical protein [Bacillus anthracis str. Ames] gb|AAP26124.11 conserved hypothetical protein [Bacillus anthracis str. Ames] Length = 324
2295.3 Best-BlastP=> >nrprot 72% Identities = 93/152 (61 %), Positives = 113/152 (74%) ref|NP_249699.11 bacterioferritin comigratory protein [Pseudomonas aeruginosa PA01] pir||A83520 bacterioferritin comigratory protein PA1008 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG04397.1 |AE004533_8 bacterioferritin comigratory protein [Pseudomonas aeruginosa PA01] Length = 157
2297.3 Best-BlastP=> >nrprot 79% Identities = 98/156 (62%), Positives = 125/156 (80%), Gaps = 3/156 (1 %) ref|NP_311509.11 small protein B [Escherichia coli 01δ7:H7] ref|NP_417110.11 small protein B [Escherichia coli K12] ref|NP_708467.1 | small protein B [Shigella flexneri 2a str. 301] ref|NP_75δ024.1 | SsrA-binding protein [Escherichia coli CFT073] ref|NP_838189.1 | ssrA(tmRNA)-binding protein [Shigella flexneri 2a str. 2457T] sp|P320δ2|SSRP_ECOLI SsrA-binding protein (Small protein B) pir||JS0701 small protein B, smpB - Escherichia coli (strain K-12) pir||B91064 small protein B [imported] - Escherichia coli (strain 0157:H7, substrain RIMD 0609952) dbj|BAA02062.1 | small protein [Escherichia coli] gb|AAA79790.1 | smpB gene product gb|AAC75669.1 | small protein B [Escherichia coli K12] dbj|BAB36905.1 | small protein B [Escherichia coli 0157:H7] gb|AAN44174.1 |AE015283_δ small protein B [Shigella flexneri 2a str. 301] gb|AAN81592.1 |AE016764_274 SsrA- binding protein [Escherichia coli CFT073] gb|AAP17999.11 ssrA(tmRNA)-binding protein [Shigella flexneri 2a str. 2457T] Length = 160
22g8.2 Best-BlastP=> >nrprot 16% Identities = 40/142 (28%), Positives = 73/142 (51 %), Gaps = 8/142 (5%) ref|NP_038716.1 | t-complex-associated testis expressed 1 [Mus musculus] pir||A45841 T-complex-associated-testes-expressed-1 protein - mouse gb|AAA40406.1| Tcte-1 peptide Length = 506
2299.3 Best-BlastP=> >nrprot 37% Identities = 22/88 (25%), Positives = 41/88 (46%), Gaps = 5/88 (5%) ref|NP_819930.11 conserved domain protein [Coxiella burnetii RSA 493] gb|AAO90444.11 conserved domain protein [Coxiella burnetii RSA 493] Length = 169
2301.2 Best-BlastP=> >nrprot 70% Identities = 397/715 (56%), Positives = 512/716 (71 %), Gaps = 4/715 (0%) ref|NP_820090.1 | ribonuclease R [Coxiella burnetii RSA 493] gb|AAO90604.1 | ribonuclease R [Coxiella burnetii RSA 493] Length = 736
2302.2 Best-BlastP=> >nrprot 72% Identities = 64/108 (59%), Positives = 81/108 (75%) ref|NP_355166.11 AGR_C_4014p [Agrobacterium tumefaciens] ref|NP_532881.11 secretion chaperone [Agrobacterium tumefaciens str. C68 (U. Washington)] pir||F97624 csaa protein [imported] - Agrobacterium tumefaciens (strain C58, Cereon) pir||AG2847 secretion chaperone [imported] - Agrobacterium tumefaciens (strain C58, Dupont) gb|AAK87951.1 | AGR_C_4014p [Agrobacterium tumefaciens str. C58 (Cereon)] gb|AAL43197.1 | secretion chaperone [Agrobacterium tumefaciens str. C68 (U. Washington)] Length = 113
2308.2 Best-BlastP=> >nrprot No Hits found
2309.2 Best-BlastP=> >nrprot 45% Identities = 104/406 (25%), Positives = 173/406 (42%), Gaps = 63/406 (15%) sp|Q9MYU4|ENP1_PIG Ectonucleoside triphosphate diphosphohydrolase 1 (NTPDasel ) (Ecto-ATP diphosphohydrolase) (ATPDase) (Lymphoid cell activation antigen) (Ecto-apyrase) (CD39 antigen) emb|CAB95871.11 ATP-diphosphohydrolase [Sus scrofa] Length = 510
2311.4 Best-BlastP=> >nrprot 75% Identities = 490/868 (56%), Positives = 648/868 (74%), Gaps = 13/868 (1 %) ref|NP_928561.11 alanyl-tRNA synthetase (alanine-tRNA ligase) [Photorhabdus luminescens subsp. laumondii TT01] emb|CAE13544.11 alanyl-tRNA synthetase (alanine- -tRNA ligase) [Photorhabdus luminescens subsp. laumondii TT01] Length = 876
2312.1 Best-BlastP=> >nrprot No Hits found 2313.2 Best-BlastP=> >nrprot 19% Identities = 40/156 (25%), Positives = 78/156 (50%), Gaps = 13/156 (8%) ref|NP_614055.11 Uncharacterized protein [Methanopyrus kandleri AV19] gb|AAM01985.11 Uncharacterized protein [Methanopyrus kandleri AV19] Length = 609
2315.3 Best-BlastP=> >nrprot 56% Identities = 93/262 (35%), Positives = 149/262 (56%), Gaps = 2/262 (0%) ref|ZP_00108734.11 COG0596: Predicted hydrolases or acyltransferases (alpha/beta hydrolase superfamily) [Nostoc punctiforme] Length = 270
2317.2 Best-BlastP=> >nrprot 71 % Identities = 266/484 (54%), Positives = 351/484 (72%), Gaps = 1/484 (0%) ref|NP_390984.11 glycine betaine aldehyde dehydrogenase [Bacillus subtilis] sp|P71016|DHAB_BACSU Betaine aldehyde dehydrogenase (BADH) pir||A69629 glycine betaine aldehyde dehydrogenase gbsA - Bacillus subtilis gb|AAC44364.1 1 GbsA emb|CAB15084.11 glycine betaine aldehyde dehydrogenase [Bacillus subtilis subsp. subtilis str. 168] Length = 4gθ
2319.3 Best-BlastP=> >nrprot 72% Identities = 263/462 (56%), Positives = 331/462 (71 %), Gaps = 6/462 (1 %) ref|NP_638201.1 1 L-serine dehydratase [Xanthomonas campestris pv. campestris str. ATCC 33913] gb|AAM42125.1 1 L-serine dehydratase [Xanthomonas campestris pv. campestris str. ATCC 33913] Length = 460
2320.3 Best-BlastP=> >nrprot 50% Identities = 163/400 (40%), Positives = 243/400 (60%), Gaps = 17/400 (4%) ref|NP_865739.11 alginate o- acetyltransferase algl [Pirellula sp.] emb|CAD73424.1 | alginate o-acetyltransferase algl [Pirellula sp.] Length = 470
2321.3 Best-BlastP=> >nrprot 65% Identities = 90/172 (52%), Positives = 120/172 (69%) ref|NP_884513.11 putative chromate reductase [Bordetella parapertussis] ref|NP_888264.1 1 putative chromate reductase [Bordetella bronchiseptica] emb|CAE32216.1 | putative chromate reductase [Bordetella bronchiseptica] emb|CAE37565.1 | putative chromate reductase [Bordetella parapertussis] Length = 184
2322.2 Best-BlastP=> >nrprot 65% Identities = 1 13/231 (48%), Positives = 152/231 (65%), Gaps = 3/231 (1 %) ref|NP_820714.11 conserved hypothetical protein [Coxiella burnetii RSA 493] gb|AA091228.1 1 conserved hypothetical protein [Coxiella burnetii RSA 493] Length = 237
2323.2 Best-BlastP=> >nrprot 34% Identities = 47/220 (21 %), Positives = 91/220 (41 %), Gaps = 16/220 (7%) ref|NP_902123.11 hypothetical protein CV2453 [Chromobacterium violaceum ATCC 12472] gb|AAQ60124.11 hypothetical protein CV2453 [Chromobacterium violaceum ATCC 12472] Length = 268
2324.4 Best-BlastP=> >nrprot No Hits found 2328.2 Best-BlastP=> >nrprot 61 % Identities = 376/865 (43%), Positives = 527/865 (60%), Gaps = 23/865 (2%) ref|NP_778639.11 diaminopimelate decarboxylase; aspartate kinase [Xylella fastidiosa Temeculal] gb|AA028288.11 diaminopimelate decarboxylase; aspartate kinase [Xylella fastidiosa Temeculal] Length = 868
2330.2 Best-BlastP=> >nrprot 72% Identities = 380/665 (57%), Positives = 488/665 (73%), Gaps = 1/665 (0%) ref|NP_246655.11 Lig [Pasteurella multocida] gb|AAK03800.1 | Lig [Pasteurella multocida] Length = 673
2333.3 Best-BlastP=> >nrprot 70% Identities = 381/700 (54%), Positives = 501/700 (71%), Gaps = 10/700 (1%) ref |NP_841209.11 Bacterial extracellular solute-binding protein, family 5 [Nitrosomonas europaea ATCC 19718] emb|CAD85063.11 Bacterial extracellular solute-binding protein, family 5 [Nitrosomonas europaea ATCC 19718] Length = 746
2335.2 Best-BlastP=> >nrprot No Hits found 2336.2 Best-BlastP=> >nrprot 70% Identities = 353/672 (52%), Positives = 479/672 (71 %), Gaps = 4/672 (0%) ref|NP_796449.11 oligopeptidase A [Vibrio parahaemolyticus RIMD 2210633] dbj|BAC58333.11 oligopeptidase A [Vibrio parahaemolyticus] Length = 680
2337.3 Best-BlastP=> >nrprot 76% Identities = 135/215 (62%), Positives = 170/215 (79%) gb|AAK20881.1 |AF334761_2 cell division ATP-binding protein [Aeromonas hydrophila] Length = 222
2339.4 Best-BlastP=> >nrprot 74% Identities = 210/345 (60%), Positives = 264/345 (76%), Gaps = 6/345 (1 %) ref|ZP_00126801.11 COG0562: Signal recognition particle GTPase [Pseudomonas syringae pv. syringae B728a] Length = 506
234.2 Best-BlastP=> >nrprot 99% Identities = 731/736 (99%), Positives = 734/736 (99%) gb|AAM00624.11 putative copper efflux ATPase [Legionella pneumophila] Length = 736
2343.2 Best-BlastP=> >nrprot 33% Identities = 67/259 (25%), Positives = 108/259 (41 %), Gaps = 34/259 (13%) ref |NP_359656.11 cell surface antigen [Rickettsia conorii] pir||C97702 cell surface antigen [imported] - Rickettsia conorii (strain Malish 7) gb|AAL02557.11 cell surface antigen [Rickettsia conorii] Length = 1902
2344.4 Best-BlastP=> >nrprot 66% Identities = 270/726 (37%), Positives = 405/726 (55%), Gaps = 16/726 (2%) ref|ZP_00043557.11 COG2114: Adenylate cyclase, family 3 (some proteins contain HAMP domain) [Magnetococcus sp. MC-1] Length = 734
2345.2 Best-BlastP=> >nrprot No Hits found
2346.1 Best-BlastP=> >nrprot No Hits found
2347.2 Best-BlastP=> >nrprot No Hits found
235.2 Best-BlastP=> >nrprot 50% Identities = 145/450 (32%), Positives = 218/450 (48%), Gaps = 60/450 (13%) sp|P42042| AMYG_ARXAD Glucoamylase precursor (Glucan 1 ,4-alpha-glucosidase) (1 ,4-alpha-D-glucan glucohydrolase) emb|CAA86997.11 glucoamylase precursor [Arxula adeninivorans] Length = 624
2350.2 Best-BlastP=> >nrprot No Hits found
2351.2 Best-BlastP=> >nrprot 59% Identities = 114/258 (44%), Positives = 164/258 (63%) ref|ZP_00122702.11 COG1043: Acyl-[acyl carrier protein]- UDP-N-acetylglucosamine O-acyltransferase [Haemophilus somnus 129PT] Length = 262
2352.2 Best-BlastP=> >nrprot 66% Identities = 160/338 (47%), Positives = 229/338 (67%) ref|ZP_00052962.11 COG1044: UDP-3-0-[3-hydroxymyristoyl] glucosamine N-acyltransferase [Magnetospirillum magnetotacticum] Length = 339
2353.3 Best-BlastP=> >nrprot 65% Identities = 174/375 (46%), Positives = 251/375 (66%), Gaps = 3/375 (0%) ref|NP_252333.11 lipid A-disaccharide synthase [Pseudomonas aeruginosa PA01] sp|Q9HXY8|LPXB_PSEAE Lipid-A-disaccharide synthase pir||C83190 lipid A-disaccharide synthase PA3643 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG07031.1 |AE004784_4 lipid A-disaccharide synthase [Pseudomonas aeruginosa PA01] Length = 378
2354.2 Best-BlastP=> >nrprot 23% Identities = 27/92 (29%), Positives = 49/92 (53%), Gaps = 2/92 (2%) dbj|BAC97056.11 RTX (repeat in toxin) cytotoxin [Vibrio vulnificus YJ016] Length = 5206
2355.2 Best-BlastP=> >nrprot 45% Identities = 83/256 (32%), Positives = 134/256 (52%), Gaps = 18/256 (7%) ref|NP_832533.11 Probable short-chain type dehydrogenase/reductase vdlC [Bacillus cereus ATCC 14579] gb|AAP09734.1 | Probable short-chain type dehydrogenase/reductase vdlC [Bacillus cereus ATCC 14579] Length = 281
2356.2 Best-BlastP=> >nrprot 65% Identities = 106/213 (49%), Positives = 141/213 (66%), Gaps = 2/213 (0%) ref |ZP_00138632.1 | COG0259: Pyridoxamine-phosphate oxidase [Pseudomonas aeruginosa UCBPP-PA14] Length = 215
2357.4 Best-BlastP=> >nrprot 6% Identities = 26/77 (33%), Positives = 38/77 (49%), Gaps = 2/77 (2%) ref|NP_752812.11 Putative conserved protein [Escherichia coli CFT073] gb|AAN79355.1 |AE016757_2δ9 Putative conserved protein [Escherichia coli CFT073] Length = 101
2368.3 Best-BlastP=> >nrprot 73% Identities = 272/470 (57%), Positives = 346/470 (73%), Gaps = 3/470 (0%) ref|ZP_00082833.11 COG0773: UDP-N- acetylmuramate-alanine ligase [Pseudomonas fluorescens PfO-1] Length = 486
236.1 Best-BlastP=> >nrprot No Hits found
2360.2 Best-BlastP=> >nrprot 52% Identities = 115/337 (34%), Positives = 189/337 (66%), Gaps = 40/337 (11 %) ref|NP_486436.11 unknown protein [Nostoc sp. PCC 7120] pir||AE2105 hypothetical protein all2396 [imported] - Nostoc sp. (strain PCC 7120) dbj|BAB7409δ.1 | ORF_ID:all2396~unknown protein [Nostoc sp. PCC 7120] Length = 454
2361.4 Best-BlastP=> >nrprot 21 % Identities = 193/1010 (19%), Positives = 405/1010 (40%), Gaps = 174/1010 (17%) pir||T14867 interaptin - slime mold (Dictyostelium discoideum) gb|AAC34582.1 | interaptin [Dictyostelium discoideum] Length = 1738
2362.3 Best-BlastP=> >nrprot 99% Identities = 302/303 (99%), Positives = 303/303 (100%) gb|AAC38180.11 DotC [Legionella pneumophila] Length = 303
2364.1 Best-BlastP=> >nrprot 99% Identities = 376/377 (99%), Positives = 377/377 (100%) gb|AAC38181.1 | DotB [Legionella pneumophila] Length = 377
2365.2 Best-BlastP=> >nrprot 52% Identities = 197/528 (37%), Positives = 302/528 (57%), Gaps = 37/528 (7%) ref|NP_658139.11 5_nucleotidase, 5'- nucleotidase, catalytic domain [Bacillus anthracis A2012] ref|NP_846555.11 5'-nucleotidase family protein [Bacillus anthracis str. Ames] gb|AAP28041.11 5'-nucleotidase family protein [Bacillus anthracis str. Ames] Length = 529
2367.2 Best-BlastP=> >nrprot No Hits found
2368.5 Best-BlastP=> >nrprot 46% Identities = 147/450 (32%), Positives = 228/450 (50%), Gaps = 1 δ/460 (3%) ref|NP_821561.11 putative acyl-CoA synthetase, long-chain fatty acid:CoA ligase [Streptomyces avermitilis MA-4680] dbj|BAC68086.11 putative acyl-CoA synthetase, long-chain fatty acid:CoA ligase [Streptomyces avermitilis MA-4680] Length = 4 5
2369.3 Best-BlastP=> >nrprot 54% Identities = 71/194 (36%), Positives = 111/194 (67%), Gaps = 1/194 (0%) ref|NP_743077.11 transporter, LysE family [Pseudomonas putida KT2440] gb|AAN66541.1 |AE016282_9 transporter, LysE family [Pseudomonas putida KT2440] Length = 204
2371.1 Best-BlastP=> >nrprot 61 % Identities = 118/343 (34%), Positives = 176/343 (61%), Gaps = 17/343 (4%) ref|NP_360212.1 | capM protein [Rickettsia conorii] pir||G97771 capM protein [imported] - Rickettsia conorii (strain Malish 7) gb|AAL03113.1 | capM protein [Rickettsia conorii] Length = 338
2372.3 Best-BlastP=> >nrprot 66% Identities = 53/118 (44%), Positives = 79/118 (66%) ref|ZP_00111665.11 C0G2146: Ferredoxin subunits of nitrite reductase and ring-hydroxylating dioxygenases [Nostoc punctiforme] Length = 119
2373.2 Best-BlastP=> >nrprot 73% Identities = 185/309 (59%), Positives = 235/309 (76%), Gaps = 2/309 (0%) ref |NP_819938.11 lytic murein transglycosylase, putative [Coxiella burnetii RSA 493] gb|AAO90452.11 lytic murein transglycosylase, putative [Coxiella burnetii RSA 493] Length = 333
2374.2 Best-BlastP=> >nrprot No Hits found
2376.3 Best-BlastP=> >nrprot 75% Identities = 115/187 (61 %), Positives = 144/187 (77%), Gaps = 2/187 (1%) ref|ZP_00091537.1 | COG0164: Ribonuclease Hll [Azotobacter vinelandii] Length = 236
2377.2 Best-BlastP=> >nrprot 73% Identities = 181/309 (58%), Positives = 236/309 (76%) ref|NP_819651.11 oxidoreductase family protein [Coxiella burnetii RSA 493] gb|AAO90165.11 oxidoreductase family protein [Coxiella burnetii RSA 493] Length = 327
238.1 Best-BlastP=> >nrprot 57% Identities = 185/432 (42%), Positives = 272/432 (62%), Gaps = 10/432 (2%) ref|NP_391464.11 similar to metabolite transport protein [Bacillus subtilis] pir||E70070 metabolite transport protein homolog ywtG - Bacillus subtilis emb|CAB07473.1 | ywtG [Bacillus subtilis] emb|CAB15600.11 ywtG [Bacillus subtilis subsp. subtilis str. 168] Length = 457
2381.2 Best-BlastP=> >nrprot 58% Identities = 148/381 (38%), Positives = 225/381 (69%), Gaps = 2/381 (0%) ref|ZP_0007987δ.11 COG0763: Lipid A disaccharide synthetase [Geobacter metallireducens] Length = 400
2382.2 Best-BlastP=> >nrprot 54% Identities = 39/79 (49%), Positives = 54/79 (68%) ref|NP_716031.11 DNA-binding protein Fis [Shewanella oneidensis MR-1] gb|AAN53476.1 |AE015487_10 DNA-binding protein Fis [Shewanella oneidensis MR-1] Length = 101
2383.4 Best-BlastP=> >nrprot 37% Identities = 21/45 (46%), Positives = 26/45 (57%), Gaps = 2/45 (4%) gb|AAQ17065.11 nucleolin 3 [Cyprinus carpio] gb|AAQ55855.11 nucleolin [Cyprinus carpio] Length = 637
2387.3 Best-BlastP=> >nrprot 55% Identities = 130/320 (40%), Positives = 187/320 (58%), Gaps = 4/320 (1 %) ref|NP_768143.1 | quinone oxidoreductase [Bradyrhizobium japonicum] dbj|BAC46768.11 quinone oxidoreductase [Bradyrhizobium japonicum USDA 110] Length = 332
2388.3 Best-BlastP=> >nrprot 72% Identities = 147/280 (52%), Positives = 212/280 (75%) ref|ZP_00021514.11 COG1175: ABC-type sugar transport systems, permease components [Ralstonia metallidurans] Length = 2 3
239.1 Best-BlastP=> >nrprot 65% Identities = 106/203 (52%), Positives = 145/203 (71 %) ref|NP_718073.11 2-deydro-3-deoxyphosphogluconate aldolase/4-hydroxy-2-oxoglutarate aldolase [Shewanella oneidensis MR-1] gb|AAN5δ517.1 |AE016690_5 2-deydro-3- deoxyphosphogluconate aldolase/4-hydroxy-2-oxoglutarate aldolase [Shewanella oneidensis MR-1] Length = 213
2391.2 Best-BlastP=> >nrprot No Hits found
2392.2 Best-BlastP=> >nrprot No Hits found
2396.3 Best-BlastP=> >nrprot 63% Identities = 174/360 (48%), Positives = 236/360 (65%), Gaps = 1/360 (0%) ref|NP_819557.11 phosphoserine aminotransferase [Coxiella burnetii RSA 493] gb|AAO90071.11 phosphoserine aminotransferase [Coxiella burnetii RSA 493] Length = 360
24.1 Best-BlastP=> >nrprot 23% Identities = 65/242 (26%), Positives = 117/242 (48%), Gaps = 20/242 (8%) ref|ZP_00096911.11 COG1738: Uncharacterized conserved protein [Novosphingobium aromaticivorans] Length = 243
240.1 Best-BlastP=> >nrprot 57% Identities = 144/316 (45%), Positives = 194/316 (61 %), Gaps = 1/316 (0%) ref|NP_928703.1 | Glucokinase (Glucose kinase) [Photorhabdus luminescens subsp. laumondii TT01] emb|CAE13698.11 Glucokinase (Glucose kinase) [Photorhabdus luminescens subsp. laumondii TT01] Length = 321
2400.2 Best-BlastP=> >nrprot 76% Identities = 181/285 (63%), Positives = 226/285 (79%), Gaps = 4/285 (1 %) ref|NP_248803.11 probable cytochrome c oxidase assembly factor [Pseudomonas aeruginosa PA01] ref |ZP_00140528.11 COG0109: Polyprenyltransferase (cytochrome oxidase assembly factor) [Pseudomonas aeruginosa UCBPP-PA14] pir||F83632 probable cytochrome c oxidase assembly factor PA0113 [imported] Pseudomonas aeruginosa (strain PA01 ) gb|AAG03503.1 |AE004449_12 probable cytochrome c oxidase assembly factor [Pseudomonas aeruginosa PA01] Length = 304
2401.2 Best-BlastP=> >nrprot 44% Identities = 57/167 (34%), Positives = 96/167 (57%), Gaps = 5/167 (2%) ref|ZP_00065551.11 COG1999: Uncharacterized protein SC01/SenC/PrrC, involved in biogenesis of respiratory and photosynthetic systems [Microbulbifer degradans 2-40] Length = 219
2402.3 Best-BlastP=> >nrprot 81 % Identities = 316/450 (70%), Positives = 370/450 (82%), Gaps = 5/450 (1 %) ref|NP_819057.11 chromosomal replication initiator protein DnaA [Coxiella burnetii RSA 493] gb|AA089571.11 chromosomal replication initiator protein DnaA [Coxiella burnetii RSA 493] Length = 451
2404.2 Best-BlastP=> >nrprot 67% Identities = 145/366 (39%), Positives = 247/366 (67%), Gaps = 2/366 (0%) ref|NP_796391.11 DNA polymerase III, beta chain [Vibrio parahaemolyticus RIMD 2210633] dbj|BAC5827δ.1 | DNA polymerase III, beta chain [Vibrio parahaemolyticus] Length = 366
2406.3 Best-BlastP=> >nrprot 57% Identities = 133/360 (36%), Positives = 205/360 (56%), Gaps = 12/360 (3%) ref|NP_759959.11 Recombinational DNA repair ATPase [Vibrio vulnificus CMCP6] sp|Q8DDJ1 |RECF_VIBVU DNA replication and repair protein recF gb|AAO09486.1 |AE016800_91 Recombinational DNA repair ATPase [Vibrio vulnificus CMCP6] Length = 359
2407.3 Best-BlastP=> >nrprot 67% Identities = 370/794 (46%), Positives = 536/794 (67%), Gaps = 5/794 (0%) ref|NP_820311.11 phenylalanyl-tRNA synthetase, beta subunit [Coxiella burnetii RSA 493] gb|AAO90825.11 phenylalanyl-tRNA synthetase, beta subunit [Coxiella burnetii RSA 493] Length = 792
2409.3 Best-BlastP=> >nrprot 100% Identities = 233/234 (99%), Positives = 234/234 (100%) emb|CAD42890.11 macrophage infectivity potentiator [Legionella pneumophila serogroup 8] Length = 236
241.2 Best-BlastP=> >nrprot 76% Identities = 386/608 (63%), Positives = 470/608 (77%) ref|NP_718074.11 6-phosphogluconate dehydratase [Shewanella oneidensis MR-1] gb|AAN5δδ18.1 |AE016690_6 6-phosphogluconate dehydratase [Shewanella oneidensis MR-1] Length = 608
2410.2 Best-BlastP=> >nrprot 56% Identities = 157/390 (40%), Positives = 240/390 (61 %), Gaps = 7/390 (1 %) ref|NP_668353.11 ampG protein [Yersinia pestis KIM] gb|AAM84604.1 |AE013705_7 ampG protein [Yersinia pestis KIM] Length = 510
2412.2 Best-BlastP=> >nrprot 62% Identities = 245/553 (44%), Positives = 350/653 (63%), Gaps = 5/553 (0%) ref|NP_719011.11 DNA repair protein RecN [Shewanella oneidensis MR-1] gb|AAN564δδ.1 |AE01 δ782_7 DNA repair protein RecN [Shewanella oneidensis MR-1] Length = 552
2413.1 Best-BlastP=> >nrprot 73% Identities = 46/67 (68%), Positives = 51/67 (76%), Gaps = 1/67 (1 %) ref|ZP_00065318.1 | COG1278: Cold shock proteins [Microbulbifer degradans 2-40] Length = 69
2414.2 Best-BlastP=> >nrprot 46% Identities = 86/333 (25%), Positives = 166/333 (49%), Gaps = 15/333 (4%) ref|NP_819780.1 | efflux transporter, RND family, MFP subunit [Coxiella burnetii RSA 493] gb|AAO90294.11 efflux transporter, RND family, MFP subunit [Coxiella burnetii RSA 493] Length = 348
2415.2 Best-BlastP=> >nrprot 37% Identities = 69/273 (25%), Positives = 121/273 (44%), Gaps = 25/273 (9%) ref|ZP_00087134.1 | COG2162: Arylamine N-acetyltransferase [Pseudomonas fluorescens PfO-1] Length = 292
2418.2 Best-BlastP=> >nrprot 63% Identities = 106/234 (45%), Positives = 154/234 (65%) ref|NP_821026.1 | membrane protein, putative [Coxiella burnetii RSA 493] gb|AA091540.11 membrane protein, putative [Coxiella burnetii RSA 493] Length = 237
241 g.3 Best-BlastP=> >nrprot 48% Identities = 74/222 (33%), Positives = 110/222 (49%), Gaps = 10/222 (4%) ref|NP_626575.11 putative dipeptidase [Streptomyces coelicolor A3(2)] emb|CAB93448.11 putative dipeptidase [Streptomyces coelicolor A3(2)] Length = 218
2421.2 Best-BlastP=> >nrprot 68% Identities = 266/502 (52%), Positives = 345/502 (68%), Gaps = 12/502 (2%) ref |NP_639214.11 competence related protein [Xanthomonas campestris pv. campestris str. ATCC 33913] gb|AAM43105.1 | competence related protein [Xanthomonas campestris pv. campestris str. ATCC 33913] Length = 506
2422.3 Best-BlastP=> >nrprot 49% Identities = 31/123 (25%), Positives = 64/123 (52%), Gaps = 3/123 (2%) ref|NP_703938.11 6-pyruvoyl tetrahydropterin synthase, putative [Plasmodium falciparum 3D7] emb|CAD50550.1 | 6-pyruvoyl tetrahydropterin synthase, putative [Plasmodium falciparum 3D7] Length = 173
2423.2 Best-BlastP=> >nrprot 63% Identities = 163/360 (45%), Positives = 237/360 (65%), Gaps = 3/360 (0%) ref|NP_820807.11 conserved hypothetical protein [Coxiella burnetii RSA 493] gb|AA091321.11 conserved hypothetical protein [Coxiella burnetii RSA 493] Length = 388
2424.3 Best-BlastP=> >nrprot 63% Identities = 92/176 (52%), Positives = 120/176 (68%), Gaps = 1/176 (0%) ref|NP_407270.1 | putative membrane protein [Yersinia pestis] pir||AF0465 probable membrane protein YP03822 [imported] - Yersinia pestis (strain C092) emb|CAC93290.11 putative membrane protein [Yersinia pestis C092] Length = 222
2425.2 Best-BlastP=> >nrprot 51% Identities = 72/187 (38%), Positives = 110/187 (58%), Gaps = 1/187 (0%) ref|NP_419508.11 conserved hypothetical . protein [Caulobacter crescentus CB15] pir||H87334 conserved hypothetical protein CC0691 [imported] - Caulobacter crescentus gb|AAK22676.11 conserved hypothetical protein [Caulobacter crescentus CB15] Length = 208
2426.2 Best-BlastP=> >nrprot No Hits found
2427.4 Best-BlastP=> >nrprot 61 % Identities = 37/65 (56%), Positives = 44/65 (67%) ref|ZP_00091135.11 COG2852: Uncharacterized protein conserved in bacteria [Azotobacter vinelandii] Length = 150
2428.2 Best-BlastP=> >nrprot 58% Identities = 36/85 (42%), Positives = 55/85 (64%), Gaps = 7/85 (8%) ref|NP_681031.11 ORF_ID:tH0240~putative transposase [Thermosynechococcus elongatus BP-1] ref|NP_681232.11 ORF_ID:tH0442~putative transposase [Thermosynechococcus elongatus BP-1] ref|NP_681335.11 ORF_ID:tH054δ~putative transposase [Thermosynechococcus elongatus BP-1] ref|NP_681541.11 ORF_ID:tlr0752~putative transposase [Thermosynechococcus elongatus BP-1] ref|NP_681563.1 | ORF_ID:tlr0774~putative transposase [Thermosynechococcus elongatus BP-1] ref|NP_681789.11 ORF_ID:tlr0999~putative transposase [Thermosynechococcus elongatus BP-1] ref|NP_682035.11 ORF_ID:tll124δ~putative transposase [Thermosynechococcus elongatus BP-1] ref|NP_682721.1 | ORF_ID:tlr1931 -putative transposase [Thermosynechococcus elongatus BP-1] dbj|BAC07793.1 | ORF_ID:tll0240~putative transposase [Thermosynechococcus elongatus BP-1] dbj|BAC07994.1 | ORF_ID:tH0442~putative transposase [Thermosynechococcus elongatus BP-1] dbj|BAC08097.1 | ORF_ID:tll0δ4δ~putative transposase [Thermosynechococcus elongatus BP-1] dbj|BAC08303.1 | ORF_ID:tlr0752~p
2429.2 Best-BlastP=> >nrprot 69% Identities = 159/294 (54%), Positives = 211/294 (71 %), Gaps = 1/294 (0%) ref|NP_792069.11 moxR protein, putative [Pseudomonas syringae pv. tomato str. DC3000] gb|AA055764.11 moxR protein, putative [Pseudomonas syringae pv. tomato str. DC3000] Length = 305
243.2 Best-BlastP=> >nrprot 67% Identities = 163/295 (55%), Positives = 210/295 (71 %), Gaps = 1/295 (0%) dbj|BAC95199.1 | putative adenine- specific methylase [Vibrio vulnificus YJ016] Length = 310
2430.2 Best-BlastP=> >nrprot 63% Identities = 172/365 (47%), Positives = 229/365 (62%), Gaps = 12/365 (3%) ref|NP_840708.11 Domain of unknown function DUF59 [Nitrosomonas europaea ATCC 19718] emb|CAD84535.11 Domain of unknown function DUF59 [Nitrosomonas europaea ATCC 19718] Length = 361
2432.2 Best-BlastP=> >nrprot 70% Identities = 233/378 (61%), Positives = 292/378 (77%) ref|NP_719320.11 ATP-dependent RNA helicase, DEAD box family [Shewanella oneidensis MR-1] gb|AAN56764.1 |AE015812_3 ATP-dependent RNA helicase, DEAD box family [Shewanella oneidensis MR-1] Length = 635
2434.2 Best-BlastP=> >nrprot 70% Identities = 184/388 (47%), Positives = 270/388 (69%), Gaps = 8/388 (2%) ref|NP_903067.11 probable stearoyl-CoA 9-desaturase [Chromobacterium violaceum ATCC 12472] gb|AAQ61061.11 probable stearoyl-CoA 9-desaturase [Chromobacterium violaceum ATCC 12472] Length = 405
2436.2 Best-BlastP=> >nrprot No Hits found 2438.2 Best-BlastP=> >nrprot 79% Identities = 59/92 (64%), Positives = 75/92 (81 %) gb|AAL59720.11 unknown [Vibrio cholerae] Length = 92
2439.2 Best-BlastP=> >nrprot 86% Identities = 81/107 (75%), Positives = 93/107 (86%), Gaps = 1/107 (0%) gb|AAL59719.1 | unknown [Vibrio cholerae] Length = 107 244.1 Best-BlastP=> >nrprot 80% Identities = 230/350 (65%), Positives = 285/350 (81 %) dbj|BAC95198.11 chorismate synthase [Vibrio vulnificus YJ016] Length = 377
2441.2 Best-BlastP=> >nrprot 51 % Identities = 134/398 (33%), Positives = 209/398 (52%), Gaps = 16/398 (4%) ref|NP_779202.1 | phage-related integrase [Xylella fastidiosa Temeculal] gb|AA028851.11 phage-related integrase [Xylella fastidiosa Temeculal] Length = 410
2442.2 Best-BlastP=> >nrprot 36% Identities = 93/315 (29%), Positives = 168/315 (60%), Gaps = 21/316 (6%) ref|NP_435846.11 Probable adenylate cyclase [Sinorhizobium meliloti] pir||H95336 probable adenylate cyclase (EC 4.6.1.1 ) [imported] - Sinorhizobium meliloti (strain 1021 ) magaplasmid pSymA gb|AAK65268.1 | Probable adenylate cyclase [Sinorhizobium meliloti] Length = 584
2443.2 Best-BlastP=> >nrprot 36% Identities = 55/232 (23%), Positives = 100/232 (43%), Gaps = 10/232 (4%) ref|ZP_00068000.11 hypothetical protein [Microbulbifer degradans 2-40] Length = 260
2445.4 Best-BlastP=> >nrprot 79% Identities = 635/926 (68%), Positives = 746/925 (80%), Gaps = 6/925 (0%) ref|NP_820536.11 ribonucleoside- diphosphate reductase, alpha subunit [Coxiella burnetii RSA 493] gb|AA091050.11 ribonucleoside-diphosphate reductase, alpha subunit [Coxiella burnetii RSA 493] Length = 941
2447.3 Best-BlastP=> >nrprot 19% Identities = 35/121 (28%), Positives = 62/121 (51 %), Gaps = 6/121 (4%) ref|NP_599246.11 protein associating with small stress protein PASS1 [Rattus norvegicus] gb|AAD48846.1 |AF168362_1 protein associating with small stress protein PASS1 [Rattus norvegicus] Length = 428
2448.3 Best-BlastP=> >nrprot 54% Identities = 94/226 (41 %), Positives = 142/226 (62%), Gaps = 8/226 (3%) ref|NP_821052.11 conserved hypothetical protein [Coxiella burnetii RSA 493] gb|AA091566.11 conserved hypothetical protein [Coxiella burnetii RSA 493] Length = 268
245.1 Best-BlastP=> >nrprot 97% Identities = 331/340 (97%), Positives = 332/340 (97%) sp|031219|DHAS_LEGPN Aspartate-semialdehyde dehydrogenase (ASA dehydrogenase) (ASADH) gb|AAC46292.11 aspartate-B-semialdehyde dehydrogenase [Legionella pneumophila] Length = 347
2450.3 Best-BlastP=> >nrprot 67% Identities = 299/616 (48%), Positives = 409/616 (66%), Gaps = 24/616 (3%) ref|NP_842471.11 ATPase component ABC-type dipeptide/oligopeptide/nickel transport system [Nitrosomonas europaea ATCC 19718] emb|CAD86392.11 ATPase component ABC-type dipeptide/oligopeptide/nickel transport system [Nitrosomonas europaea ATCC 19718] Length = 693
2453.4 Best-BlastP=> >nrprot 75% Identities = 554/897 (61 %), Positives = 689/897 (76%), Gaps = 3/897 (0%) ref|ZP_00096570.11 COG0474: Cation transport ATPase [Novosphingobium aromaticivorans] Length = 911
2456.2 Best-BlastP=> >nrprot 78% Identities = 201/314 (64%), Positives = 251/314 (79%) ref|NP_820493.11 acetyl-CoA carboxylase, carboxyl transferase, alpha subunit [Coxiella burnetii RSA 493] gb|AA091007.11 acetyl-CoA carboxylase, carboxyl transferase, alpha subunit [Coxiella burnetii RSA 493] Length = 316
2457.2 Best-BlastP=> >nrprot 46% Identities = 75/215 (34%), Positives = 114/215 (53%), Gaps = 16/215 (7%) ref|ZP_00107102.1 | COG2091 : Phosphopantetheinyl transferase [Nostoc punctiforme] Length = 239
2458.2 Best-BlastP=> >nrprot 68% Identities = 190/371 (51%), Positives = 259/371 (69%) ref|NP_819627.11 oxygen-independent coproporphyrinogen III oxidase, putative [Coxiella burnetii RSA 493] gb|AAO90141.1 | oxygen-independent coproporphyrinogen III oxidase, putative [Coxiella burnetii RSA 493] Length = 375
2459.1 Best-BlastP=> >nrprot 69% Identities = 94/176 (53%), Positives = 131/176 (74%), Gaps = 1/176 (0%) ref|NP_715832.11 MutT/nudix family protein [Shewanella oneidensis MR-1] gb|AAN53277.1 |AE015469_1 MutT/nudix family protein [Shewanella oneidensis MR-1] Length = 183
2460.3 Best-BlastP=> >nrprot 62% Identities = 230/569 (40%), Positives = 358/569 (62%), Gaps = 12/569 (2%) ref |NP_820129.11 oligopeptide transporter, OPT family [Coxiella burnetii RSA 493] gb|AAO90643.11 oligopeptide transporter, OPT family [Coxiella burnetii RSA 4 3] Length = 669
2461.2 Best-BlastP=> >nrprot 57% Identities = 33/49 (67%), Positives = 41/49 (83%) ref|NP_051689.11 integrase/recombinase XerD, putative [Deinococcus radiodurans] pir||G75636 probable integrase/recombinase XerD - Deinococcus radiodurans (strain R1 ) gb|AAF12667.1 |AE001827_5 integrase/recombinase XerD, putative [Deinococcus radiodurans] Length = 236
2462.2 Best-BlastP=> >nrprot 33% Identities = 20/38 (52%), Positives = 25/38 (65%) ref|ZP_00111545.11 COG4644: Transposase and inactivated derivatives, TnpA family [Nostoc punctiforme] Length = 1014
2464.2 Best-BlastP=> >nrprot 67% Identities = 61/108 (56%), Positives = 80/108 (74%) ref|NP_841626.11 transposase [Nitrosomonas europaea ATCC 19718] ref|NP_842205.11 transposase [Nitrosomonas europaea ATCC 19718] ref|NP_842439.1 | transposase [Nitrosomonas europaea ATCC 19718] emb|CAD85497.11 transposase [Nitrosomonas europaea ATCC 19718] emb|CAD86112.11 transposase [Nitrosomonas europaea ATCC 19718] emb|CAD86359.11 transposase [Nitrosomonas europaea ATCC 19718] Length = 122
2466.3 Best-BlastP=> >nrprot 52% Identities = 316/879 (35%), Positives = 473/879 (53%), Gaps = 49/879 (5%) ref|NP_819380.11 aminopeptidase N [Coxiella burnetii RSA 493] gb|AA089894.1 | aminopeptidase N [Coxiella burnetii RSA 493] Length = 878
2468.2 Best-BlastP=> >nrprot 39% Identities = 76/253 (30%), Positives = 122/253 (48%), Gaps = 18/253 (7%) ref|NP_634577.11 putative hydrolase [Methanosarcina mazei Goel] gb|AAM32249.1 | putative hydrolase [Methanosarcina mazei Goel] Length = 279
2469.2 Best-BlastP=> >nrρrot 44% Identities = 138/423 (32%), Positives = 216/423 (51 %), Gaps = 7/423 (1 %) ref|NP_798657.11 conserved hypothetical protein [Vibrio parahaemolyticus RIMD 2210633] dbj|BAC60541.11 conserved hypothetical protein [Vibrio parahaemolyticus] Length = 483 247.1 Best-BlastP=> >nrprot 2g% Identities = 49/120 (40%), Positives = 76/120 (63%) ref|ZP_00066693.11 COG2840: Uncharacterized protein conserved in bacteria [Microbulbifer degradans 2-40] Length = 190
2471.3 Best-BlastP=> >nrprot 56% Identities = 129/344 (37%), Positives = 199/344 (57%), Gaps = 6/344 (1 %) ref|NP_819673.11 riboflavin biosynthesis protein RibD [Coxiella burnetii RSA 493] gb|AAO90187.11 riboflavin biosynthesis protein RibD [Coxiella burnetii RSA 493] Length = 354
2472.2 Best-BlastP=> >nrprot 61 % Identities = 87/219 (39%), Positives = 131/219 (59%), Gaps = 12/219 (5%) ref|NP_820010.1 | dethiobiotin synthetase [Coxiella burnetii RSA 493] gb|AAO90524.11 dethiobiotin synthetase [Coxiella burnetii RSA 493] Length = 242
2473.1 Best-BlastP=> >nrprot 54% Identities = 36/70 (60%), Positives = 48/70 (68%), Gaps = 3/70 (4%) ref|NP_820641.11 conserved hypothetical protein [Coxiella burnetii RSA 493] gb|AAO910δδ.11 conserved hypothetical protein [Coxiella burnetii RSA 493] Length = 91
2474.1 Best-BlastP=> >nrprot 72% Identities = 105/192 (54%), Positives = 152/192 (79%), Gaps = 2/192 (1%) ref|NP_820542.1 | conserved hypothetical protein [Coxiella burnetii RSA 493] gb|AA091056.11 conserved hypothetical protein [Coxiella burnetii RSA 493] Length = 209
2477.1 Best-BlastP=> >nrprot 52% Identities = 68/115 (59%), Positives = 85/115 (73%), Gaps = 2/115 (1 %) ref|NP_819994.11 rare lipoprotein A family protein [Coxiella burnetii RSA 493] gb|AAO90508.11 rare lipoprotein A family protein [Coxiella burnetii RSA 493] Length = 261
2479.2 Best-BlastP=> >nrprot No Hits found
248.2 Best-BlastP=> >nrprot 59% Identities = 191/456 (41%), Positives = 278/456 (60%), Gaps = 7/456 (1%) ref|NP_820284.1 | ankyrin repeat domain protein [Coxiella burnetii RSA 493] gb|AAO90798.11 ankyrin repeat domain protein [Coxiella burnetii RSA 493] Length = 465
2481.4 Best-BlastP=> >nrprot 77% Identities = 185/291 (63%), Positives = 231/291 (79%), Gaps = 1/291 (0%) ref|NP_286072.11 putative phosphonomutase 2 [Escherichia coli 0157:H7 EDL933] ref|NP_308412.1 | putative phosphonomutase 2 [Escherichia coli 0157:H7] pir||A90677 probable phosphonomutase 2 [imported] - Escherichia coli (strain 0157:H7, substrain RIMD 0509952) pir||D85527 probable phosphonomutase 2 [imported] - Escherichia coli (strain 0157:H7, substrain EDL933) gb|AAG54680.1 |AE005212_δ putative phosphonomutase 2 [Escherichia coli 0157:H7 EDLg33] dbj|BAB33808.11 putative phosphonomutase 2 [Escherichia coli 01δ7:H7] Length = 296
2482.1 Best-BlastP=> >nrprot 59% Identities = 75/126 (60%), Positives = 101/125 (80%) ref|NP_820989.1 | conserved domain protein [Coxiella burnetii RSA 493] gb|AAO91503.1 | conserved domain protein [Coxiella burnetii RSA 493] Length = 127
2483.2 Best-BlastP=> >nrprot No Hits found
2485.3 Best-BlastP=> >nrprot No Hits found "
2486.3 Best-BlastP=> >nrprot 55% Identities = 188/514 (36%), Positives = 280/514 (54%), Gaps = 43/514 (8%) ref|ZP_00087809.1 | COG2202: FOG: PAS/PAC domain [Pseudomonas fluorescens PfO-1] Length = 757
2488.2 Best-BlastP=> >nrprot No Hits found
2489.2 Best-BlastP=> >nrprot No Hits found '
249.4 Best-BlastP=> >nrprot 97% Identities = 558/575 (97%), Positives = 662/576 (97%), Gaps = 1/575 (0%) gb|AAC12716.1 | pilus assembly protein PilB [Legionella pneumophila] Length = 576
2490.2 Best-BlastP=> >nrprot No Hits found
2492.2 Best-BlastP=> >nrprot No Hits found
2493.2 Best-BlastP=> >nrprot 67% Identities = 99/203 (48%), Positives = 135/203 (66%), Gaps = 10/203 (4%) ref|NP_929371.11 Holliday junction DNA helicase [Photorhabdus luminescens subsp. laumondii TT01] emb|CAE14404.11 Holliday junction DNA helicase [Photorhabdus luminescens subsp. laumondii TT01] Length = 205
2495.2 Best-BlastP=> >nrprot 69% Identities = 93/170 (54%), Positives = 121/170 (71 %), Gaps = 1/170 (0%) ref|NP_231481.1 | crossover junction endodeoxyribonuclease RuvC [Vibrio cholerae 01 biovar eltor str. N16961] sp|Q9KR00|RUVC_VIBCH Crossover junction endodeoxyribonuclease ruvC (Holliday junction nuclease ruvC) (Holliday juction resolvase ruvC) pir||H82149 crossover junction endodeoxyribonuclease RuvC VC1847 [imported] - Vibrio cholerae (strain N16961 serogroup 01 ) gb|AAFg4g95.1 | crossover junction endodeoxyribonuclease RuvC [Vibrio cholerae 01 biovar eltor str. N 16961] Length = 173
25.1 Best-BlastP=> >nrprot 46% Identities = 46/124 (37%), Positives = 72/124 (58%), Gaps = 3/124 (2%) ref|NP_488674.11 probable cytosine deaminase [Nostoc sp. PCC 7120] pir||AB2385 hypothetical protein alr4634 [imported] - Nostoc sp. (strain PCC 7120) dbj|BAB76333.1 | ORF_ID:alr4634~probable cytosine deaminase [Nostoc sp. PCC 7120] Length = 140
250.1 Best-BlastP=> >nrprot 98% Identities = 398/406 (98%), Positives = 400/406 (98%) gb|AAC12717.11 pilus assembly protein PilC [Legionella pneumophila] Length = 406
2501.2 Best-BlastP=> >nrprot 70% Identities = 213/412 (51%), Positives = 294/412 (71 %), Gaps = 12/412 (2%) ref|NP_819142.1 | tolB protein [Coxiella burnetii RSA 493] sp|Q83F59|TOLB_COXBU TolB protein precursor gb|AA089666.11 tolB protein [Coxiella burnetii RSA 493] Length = 437
2604.4 Best-BlastP=> >nrprot 51 % Identities = 106/301 (35%), Positives = 164/301 (54%), Gaps = 34/301 (11 %) ref|NP_718333.11 tolA protein [Shewanella oneidensis MR-1] gb|AAN55777.1 |AE015714_4 tolA protein [Shewanella oneidensis MR-1] Length = 345
2506.2 Best-BlastP=> >nrprot 83% Identities = 238/338 (70%), Positives = 282/338 (83%) gb|AAN87043.11 HypE [Thiocapsa roseopersicina] Length = 360
2508.3 Best-BlastP=> >nrprot 77% Identities = 234/375 (62%), Positives = 285/375 (76%), Gaps = 6/375 (1 %) ref|ZP_00021585.11 COG0409: Hydrogenase maturation factor [Ralstonia metallidurans] Length = 380
2509.3 Best-BlastP=> >nrprot No Hits found
251.2 Best-BlastP=> >nrprot 97% Identities = 277/287 (96%), Positives = 281/287 (97%),sp|068433|LEP4_LEGPN Type 4 prepilin-like proteins leader peptide processing enzyme [Includes: Leader peptidase (Prepilin peptidase); N-methyltransferase ] gb|AAC12718.11 type IV prepilin- like protein specific leader peptidase PilD [Legionella pneumophila] Length = 287
2510.2 Best-BlastP=> >nrprot 69% Identities = 310/637 (48%), Positives = 427/637 (67%), Gaps = 19/637 (2%) ref|ZP_00067611.11 COG0488: ATPase components of ABC transporters with duplicated ATPase domains [Microbulbifer degradans 2-40] Length = 637
2514.3 Best-BlastP=> >nrprot 74% Identities = 52/84 (61 %), Positives = 67/84 (79%), Gaps = 2/84 (2%) ref|ZP_00128272.11 COG0851 : Septum formation topological specificity factor [Pseudomonas syringae pv. syringae B728a] Length = 84
2517.2 Best-BlastP=> >nrprot 65% Identities = 197/336 (58%), Positives = 240/336 (71 %), Gaps = 2/336 (0%) ref|NP_634897.11 L-sorbosone dehydrogenase [Methanosarcina mazei Goel] gb|AAM32569.1 | L-sorbosone dehydrogenase [Methanosarcina mazei Goel] Length = 381
2518.4 Best-BlastP=> >nrprot 86% Identities = 123/179 (68%), Positives = 156/179 (87%) ref|NP_250365.11 GTP cyclohydrolase I precursor [Pseudomonas aeruginosa PA01] sp|Q9l351 |GC12_PSEAE GTP cyclohydrolase I 2 (GTP-CH-I 2) pir||C83435 GTP cyclohydrolase I precursor PA1674 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG05063.1 |AE00459δ_2 GTP cyclohydrolase I precursor [Pseudomonas aeruginosa PA01] Length = 181
252.2 Best-BlastP=> >nrprot 40% Identities = 86/303 (28%), Positives = 161/303 (53%), Gaps = 13/303 (4%) ref |NP_819235.11 CAAX amino terminal protease family protein [Coxiella burnetii RSA 493] gb|AA089749.1 | CAAX amino terminal protease family protein [Coxiella burnetii RSA 493] Length = 297
2520.4 Best-BlastP=> >nrprot 73% Identities = 64/111 (57%), Positives = 84/111 (75%), Gaps = 1/111 (0%) ref|NP_819816.11 HIT family protein [Coxiella burnetii RSA 493] gb|AAO90330.11 HIT family protein [Coxiella burnetii RSA 493] Length = 113
2521.4 Best-BlastP=> >nrprot 71% Identities = 206/348 (68%), Positives = 251/348 (72%), Gaps = 1/348 (0%) ref|ZP_00028857.11 COG1064: Zn- dependent alcohol dehydrogenases [Burkholderia fungorum] Length = 377
2522.1 Best-BlastP=> >nrprot No Hits found
2523.1 Best-BlastP=> >nrprot 54% Identities = 46/128 (35%), Positives = 73/128 (57%) ref |ZP_00011417.11 hypothetical protein [Rhodopseudomonas palustris] Length = 136
2624.2 Best-BlastP=> >nrprot 51 % Identities = 71/192 (36%), Positives = 105/192 (54%), Gaps = 7/192 (3%) sp|Q92JI7|DEF2_RICCN Peptide deformylase 2 (PDF 2) (Polypeptide deformylase 2) Length = 202
2526.3 Best-BlastP=> >nrprot 13% Identities = 40/132 (30%), Positives = 59/132 (44%), Gaps = 16/132 (12%) ref|NP_440029.11 acetylpolyamine aminohydolase [Synechocystis sp. PCC 6803] sp|P72702|Y245_SYNY3 Hypothetical protein slr0245 pir||S74557 acetylpolyamine aminohydrolase - Synechocystis sp. (strain PCC 6803) dbj|BAA16709.11 acetylpolyamine aminohydolase [Synechocystis sp. PCC 6803] Length = 304
2528.3 Best-BlastP=> >nrprot 41 % Identities = 53/236 (22%), Positives = 105/236 (44%), Gaps = 7/236 (2%) ref|NP_845027.11 acetyltransferase, GNAT family [Bacillus anthracis str. Ames] gb|AAP26513.1 | acetyltransferase, GNAT family [Bacillus anthracis str. Ames] Length = 266 2630.3 Best-BlastP=> >nrprot 63% Identities = 266/626 (42%), Positives = 401/626 (64%), Gaps = 8/626 (1 %) ref|NP_900243.11 potassium uptake protein [Chromobacterium violaceum ATCC 12472] gb|AAQ58249.1 | potassium uptake protein [Chromobacterium violaceum ATCC 12472] Length = 621
2533.3 Best-BlastP=> >nrprot 52% Identities = 88/286 (30%), Positives = 158/286 (55%), Gaps = 4/286 (1 %) ref|NP_763522.11 Transcriptional regulator [Vibrio vulnificus CMCP6] gb|AAO08512.1 |AE016813_264 Transcriptional regulator [Vibrio vulnificus CMCP6] Length = 307
2536.2 Best-BlastP=> >nrprot 70% Identities = 200/351 (56%), Positives = 253/351 (72%) ref|NP_519918.11 PROBABLE PYRUVATE DEHYDROGENASE E1 COMPONENT (ALPHA SUBUNIT) OXIDOREDUCTASE PROTEIN [Ralstonia solanacearum] emb|CAD15499.11 PROBABLE PYRUVATE DEHYDROGENASE E1 COMPONENT (ALPHA SUBUNIT) OXIDOREDUCTASE PROTEIN [Ralstonia solanacearum] Length = 363
2536.2 Best-BlastP=> >nrprot 57% Identities = 180/451 (39%), Positives = 255/451 (56%), Gaps = 36/451 (7%) ref|NP_798254.11 para-aminobenzoate synthase, component I [Vibrio parahaemolyticus RIMD 2210633] dbj|BAC60138.1 | para-aminobenzoate synthase, component I [Vibrio parahaemolyticus] Length = 454
2538.2 Best-BlastP=> >nrprot 78% Identities = 245/394 (62%), Positives = 309/394 (78%) ref|NP_901758.11 acetyl-CoA C-acetyltransferase [Chromobacterium violaceum ATCC 12472] gb|AAQ59760.11 acetyl-CoA C-acetyltransferase [Chromobacterium violaceum ATCC 12472] Length = 394
254.2 Best-BlastP=> >nrprot 49% Identities = 275/861 (32%), Positives = 428/851 (50%), Gaps = 29/851 (3%) ref|NP_490573.11 ATP-binding protein [Salmonella typhimurium LT2] gb|AAL23492.11 conjugative transfer: assembly [Salmonella typhimurium LT2] Length = 882
2642.2 Best-BlastP=> >nrprot 61% Identities = 214/406 (62%), Positives = 282/405 (69%), Gaps = 8/405 (1%) ref|ZP_00014043.1 | COG0260: Leucyl aminopeptidase [Rhodospirillum rubrum] Length = 444
2544.3 Best-BlastP=> >nrprot 5% Identities = 36/130 (27%), Positives = 68/130 (52%), Gaps = 7/130 (5%) ref|NP_711281.11 outer membrane efflux protein [Leptospira interrogans serovar lai str. 66601] gb|AAN48299.1 |AE011292_12 outer membrane efflux protein [Leptospira interrogans serovar lai str. 56601] Length = 533
2546.2 Best-BlastP=> >nrprot No Hits found
2546.4 Best-BlastP=> >nrprot 47% Identities = 155/432 (35%), Positives = 239/432 (65%), Gaps = 21/432 (4%) emb|CAE02834.11 OSJNBa0043A12.39 [Oryza sativa (japonica cultivar-group)] Length = 487
2549.3 Best-BlastP=> >nrprot No Hits found
255.1 Best-BlastP=> >nrprot 39% Identities = 23/83 (27%), Positives = 44/83 (53%), Gaps = 1/83 (1 %) ref|NP_762592.11 Unknown [Vibrio vulnificus CMCP6] gb|AAO07582.1 |AE016810_85 Unknown [Vibrio vulnificus CMCP6] Length = 114
2560.2 Best-BlastP=> >nrprot 99% Identities = 443/449 (98%), Positives = 446/449 (99%) gb|AAB52239.11 nucleotide binding protein Flil [Legionella pneumophila] Length = 449
2551.2 Best-BlastP=> >nrprot 34% Identities = 74/75 (98%), Positives = 74/75 (98%) gb|AAB52238.11 FliH [Legionella pneumophila] Length = 75
2553.3 Best-BlastP=> >nrprot 76% Identities = 168/326 (51 %), Positives = 251/326 (76%) ref|NP_791782.11 flagellar motor switch protein FliG [Pseudomonas syringae pv. tomato str. DC3000] gb|AA055477.11 flagellar motor switch protein FliG [Pseudomonas syringae pv. tomato str. DC3000] Length = 333
2555.5 Best-BlastP=> >nrprot No Hits found
2567.4 Best-BlastP=> >nrprot 61 % Identities = 70/135 (51 %), Positives = 91/135 (67%) ref|ZP_00085924.11 hypothetical protein [Pseudomonas fluorescens PfO-1] Length = 141
2559.3 Best-BlastP=> >nrprot 85% Identities = 243/333 (72%), Positives = 289/333 (86%) ref|NP_438478.11 Holliday junction DNA helicase [Haemophilus influenzae Rd] sp|P44631|RUVB_HAEIN Holliday junction DNA helicase ruvB pir||B64061 DNA-binding protein ruvB - Haemophilus influenzae (strain Rd KW20) gb|AAC21975.11 Holliday junction DNA helicase (ruvB) [Haemophilus influenzae Rd] Length = 335
256.1 Best-BlastP=> >nrprot 62% Identities = 79/194 (40%), Positives = 109/194 (56%), Gaps = 14/194 (7%) ref|NP_762593.11 Conserved hypothetical protein [Vibrio vulnificus CMCP6] gb|AAO07583.1 |AE016810_86 Conserved hypothetical protein [Vibrio vulnificus CMCP6] Length = 226
2560.3 Best-BlastP=> >nrprot 98% Identities = 186/187 (99%), Positives = 186/187 (99%) gb|AAQ18125.11 RpoE [Legionella pneumophila] Length = 187
2662.2 Best-BlastP=> >nrprot No Hits found
2565.4 Best-BlastP=> >nrprot 55% Identities = 88/226 (38%), Positives = 129/226 (57%), Gaps = 17/226 (7%) ref|NP_651138.11 CG6763-PA [Drosophila melanogaster] gb|AAF56122.1 | CG6763-PA [Drosophila melanogaster] gb|AAL68281.1 | RE28575p [Drosophila melanogaster] Length = 354
2567.2 Best-BlastP=> >nrprot 42% Identities = 53/201 (26%), Positives = 92/201 (45%), Gaps = 3/201 (1 %) ref|NP_683243.11 ORF_ID:tll2454~unknown protein [Thermosynechococcus elongatus BP-1] dbj|BAC10005.1 | ORF_ID:tll2454~unknown protein [Thermosynechococcus elongatus BP-1] Length = 253
2568.1 Best-BlastP=> >nrprot No Hits found
2569.3 Best-BlastP=> >nrprot 56% Identities = 80/207 (38%), Positives = 122/207 (58%), Gaps = 18/207 (8%) ref|NP_840655.11 conserved hypothetical protein [Nitrosomonas europaea ATCC 19718] emb|CAD84482.1 | conserved hypothetical protein [Nitrosomonas europaea ATCC 19718] Length = 253
2570.2 Best-BlastP=> >nrprot 64% Identities = 144/374 (38%), Positives = 243/374 (64%), Gaps = 4/374 (1 %) ref |NP_518286.11 PROBABLE TRANSMEMBRANE PROTEIN [Ralstonia solanacearum] emb|CAD13693.11 PROBABLE TRANSMEMBRANE PROTEIN [Ralstonia solanacearum] Length = 381
2571.2 Best-BlastP=> >nrprot 64% Identities = 135/292 (46%), Positives = 199/292 (68%), Gaps = 3/292 (1 %) reflZP_00056519.11 COG1131 : ABC-type multidrug transport system, ATPase component [Magnetospirillum magnetotacticum] Length = 308
2573.3 Best-BlastP=> >nrprot 98% Identities = 188/192 (97%), Positives = 190/192 (98%) gb|AAK00281.1 |AF288536_3 unknown [Legionella longbeachae] Length = 192
2576.2 Best-BlastP=> >nrprot 99% Identities = 319/319 (100%), Positives = 319/319 (100%) gb|AAM00641.11 putative Na/Ca antiporter [Legionella pneumophila] Length = 319
2578.2 Best-BlastP=> >nrprot 99% Identities = 234/234 (100%), Positives = 234/234 (100%) gb|AAM00640.11 unknown [Legionella pneumophila] Length = 234
2579.2 Best-BlastP=> >nrprot 62% Identities = 54/135 (40%), Positives = 89/135 (65%) ref|NP_903458.11 conserved hypothetical protein [Chromobacterium violaceum ATCC 12472] gb|AAQ61460.1 | conserved hypothetical protein [Chromobacterium violaceum ATCC 12472] Length = 159
258.2 Best-BlastP=> >nrprot 56% Identities = 161/309 (52%), Positives = 213/309 (68%), Gaps = 4/309 (1%) gb|AAM90716.11 TraU [Salmonella typhi] Length = 331
2580.2 Best-BlastP=> >nrprot 78% Identities = 155/249 (62%), Positives = 198/249 (79%) ref|ZP_00133889.11 COG2226: Methylase involved in ubiquinone/menaquinone biosynthesis [Actinobacillus pleuropneumoniae serovar 1 str. 4074] Length = 258
2582.3 Best-BlastP=> >nrprot 47% Identities = 242/883 (27%), Positives = 412/883 (46%), Gaps = 56/883 (6%) ref |NP_842182.11 conserved hypothetical protein [Nitrosomonas europaea ATCC 19718] emb|CAD86089.1 | conserved hypothetical protein [Nitrosomonas europaea ATCC 19718] Length = 909
2583.3 Best-BlastP=> >nrprot 38% Identities = 40/156 (25%), Positives = 72/156 (46%), Gaps = 23/156 (14%) ref|NP_764254.1 | Na+/H+ antiporter-like protein [Staphylococcus epidermidis ATCC 12228] gb|AAO04296.1 |AE016746_86 Na+/H+ antiporter-like protein [Staphylococcus epidermidis ATCC 12228] Length = 614
2586.2 Best-BlastP=> >nrprot 13% Identities = 45/193 (23%), Positives = 77/193 (39%), Gaps = 30/193 (15%) ref |NP_866179.11 hypothetical protein- transmembrane region and signal peptide prediction [Pirellula sp.] emb|CAD73865.1 | hypothetical protein-transmembrane region and signal peptide prediction [Pirellula sp.] Length = 500
2587.2 Best-BlastP=> >nrprot 59% Identities = 114/283 (40%), Positives = 169/283 (59%), Gaps = 4/283 (1 %) ref|ZP_00094776.11 COG0583: Transcriptional regulator [Novosphingobium aromaticivorans] Length = 290
2590.3 Best-BlastP=> >nrprot 60% Identities = 125/288 (43%), Positives = 184/288 (63%), Gaps = 3/288 (1 %) ref|ZP_00096382.11 COG0121 : Predicted glutamine amidotransferase [Novosphingobium aromaticivorans] Length = 444
2591.3 Best-BlastP=> >nrprot 66% Identities = 182/348 (52%), Positives = 234/348 (67%), Gaps = 2/348 (0%) ref|NP_233382.11 NADH-dependent flavin oxidoreductase, Oye family [Vibrio cholerae - 01 biovar eltor str. N16961] pir||H82391 NADH-dependent flavin oxidoreductase, Oye family VCA0998 [imported] - Vibrio cholerae (strain N 16961 serogroup 01 ) gb|AAF96894.11 NADH-dependent flavin oxidoreductase, Oye family [Vibrio cholerae 01 biovar eltor str. N16961] Length = 347
2592.4 Best-BlastP=> >nrprot No Hits found 2593.3 Best-BlastP=> >nrprot 62% Identities = 98/248 (39%), Positives = 161/248 (64%), Gaps = 7/248 (2%) ref|NP_250139.11 flagellar biosynthetic protein FliR [Pseudomonas aeruginosa PA01] pir||B83465 flagellar biosynthetic protein FliR PA1448 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG04837.1 |AE004574_8 flagellar biosynthetic protein FliR [Pseudomonas aeruginosa PA01] Length = 258
2595.4 Best-BlastP=> >nrprot 77% Identities = 142/241 (58%), Positives = 193/241 (80%), Gaps = 1/241 (0%) ref|NP_746469.11 flagellar biosynthetic protein FliP [Pseudomonas putida KT2440] gb|AAD01927.2| FliP [Pseudomonas putida] gb|AAN69933.1 |AE016632_4 flagellar biosynthetic protein FliP [Pseudomonas putida KT2440] Length = 251
2597.4 Best-BlastP=> >nrprot 99% Identities = 225/226 (99%), Positives = 226/226 (100%) gb|AAM00392.1 |AF386079_2 CcmB [Legionella pneumophila] Length = 226
2598.3 Best-BlastP=> >nrprot 59% Identities = 76/216 (35%), Positives = 130/216 (60%), Gaps = 6/216 (2%) ref|NP_799389.11 conserved hypothetical protein [Vibrio parahaemolyticus RIMD 2210633] dbj|BAC61273.11 conserved hypothetical protein [Vibrio parahaemolyticus] Length = 219
2599.4 Best-BlastP=> >nrprot 82% Identities = 380/556 (68%), Positives = 461/566 (82%), Gaps = 2/556 (0%) ref|NP_927905.11 ATP-binding protein YjjK [Photorhabdus luminescens subsp. laumondii TT01] emb|CAE12850.11 ATP-binding protein YjjK [Photorhabdus luminescens subsp. laumondii TT01] Length = 565
26.1 Best-BlastP=> >nrprot 30% Identities = 57/132 (43%), Positives = 81/132 (61 %), Gaps = 6/132 (4%) ref|ZP_00031526.11 hypothetical protein [Burkholderia fungorum] Length = 163
2600.4 Best-BlastP=> >nrprot 84% Identities = 239/341 (70%), Positives = 287/341 (84%) ref |NP_931999.11 threonine 3-dehydrogenase [Photorhabdus luminescens subsp. laumondii TT01] emb|CAE17217.1 | threonine 3-dehydrogenase [Photorhabdus luminescens subsp. laumondii TT01] Length = 341
2602.2 Best-BlastP=> >nrprot 63% Identities = 144/273 (62%), Positives = 188/273 (68%), Gaps = 2/273 (0%) ref|ZP_00067856.11 COG0061 : Predicted sugar kinase [Microbulbifer degradans 2-40] Length = 294
2604.3 Best-BlastP=> >nrprot 83% Identities = 437/603 (72%), Positives = 511/603 (84%) ref|ZP_00043195.1 | COG1217: Predicted membrane GTPase involved in stress response [Magnetococcus sp. MC-1] Length = 611
2605.3 Best-BlastP=> >nrprot 53% Identities = 300/814 (36%), Positives = 445/814 (54%), Gaps = 23/814 (2%) ref|NP_111982.11 Type III restriction- modification enzyme, helicase subunit [Thermoplasma volcanium] dbj|BAB60631.1 | TVG1539639 [Therrnoplasrna volcanium] Length = 843
2607.2 Best-BlastP=> >nrprot 34% Identities = 87/295 (29%), Positives = 141/295 (47%), Gaps = 10/296 (3%) ref|NP_827021.11 hypothetical protein [Streptomyces avermitilis MA-4680] dbj|BAC73566.1 | hypothetical protein [Streptomyces avermitilis MA-4680] Length = 416
2609.2 Best-BlastP=> >nrprot No Hits found
2611.1 Best-BlastP=> >nrprot No Hits found
2616.1 Best-BlastP=> >nrprot 35% Identities = 58/180 (32%), Positives = 106/180 (58%), Gaps = 1/180 (0%) ref|NP_624053.11 predicted transposase [Thermoanaerobacter tengcongensis] gb|AAM25657.11 predicted transposase [Thermoanaerobacter tengcongensis] Length = 267
2619.1 Best-BlastP=> >nrprot 40% Identities = 37/137 (27%), Positives = 62/137 (45%), Gaps = 39/137 (28%) emb|CAB46580.1 | IS1400 transposase B [Yersinia enterocolitica] Length = 294
262.3 Best-BlastP=> >nrprot 41% Identities = 123/341 (36%), Positives = 184/341 (53%), Gaps = 21/341 (6%) ref|NP_827217.11 putative quinolinate synthetase [Streptomyces avermitilis MA-4680] dbj|BAC73752.1 | putative quinolinate synthetase [Streptomyces avermitilis MA-4680] Length = 414
2620.1 Best-BlastP=> >nrprot 91 % Identities = 65/88 (73%), Positives = 81/88 (92%) ref|NP_395197.11 putative transposase ORFA [Yersinia pestis C092] ref|NP_857719.11 low calcium response locus protein S homolog [Yersinia pestis KIM] ref|NP_857914.1 | putative IS element protein [Yersinia pestis KIM] sp|Q00931 |LCRS_YERPE Low calcium response locus protein S pir||T43562 probable IS element protein - Yersinia pestis plasmid pCD1 gb|AAA27655.1 | IcrS gb|AAC62579.1 | low calcium response locus protein S homolog [Yersinia pestis KIM] gb|AAC69827.1 | putative IS element protein [Yersinia pestis KIM] emb|CAB54940.11 putative transposase ORFA [Yersinia pestis] Length = 88
2622.1 Best-BlastP=> >nrprot 58% Identities = 267/639 (41 %), Positives = 378/639 (59%), Gaps = 54/639 (8%) ref|NP_111983.11 Adenine specific DNA methylase (Mod-related) [Thermoplasma volcanium] dbj|BAB60632.1 | modification methylase [Thermoplasma volcanium] Length = 616
2624.1 Best-BlastP=> >nrprot 38% Identities = 44/156 (28%), Positives = 71/156 (45%), Gaps = 20/156 (12%) ref |ZP_00023112.11 hypothetical protein [Ralstonia metallidurans] Length = 348
2626.2 Best-BlastP=> >nrprot 58% Identities = 129/264 (48%), Positives = 170/264 (64%), Gaps = 2/264 (0%) gb|AAM08235.11 LvrA [Legionella pneumophila] Length = 289
2626.1 Best-BlastP=> >nrprot 56% Identities = 80/221 (36%), Positives = 124/221 (56%), Gaps = 16/221 (7%) gb|AAM08234.11 putative phage repressor [Legionella pneumophila] Length = 227
2627.1 Best-BlastP=> >nrprot 79% Identities = 300/397 (75%), Positives = 334/397 (84%), Gaps = 5/397 (1 %) gb|AAB05678.11 HelB Length = 400
263.1 Best-BlastP=> >nrprot 61 % Identities = 259/628 (49%), Positives = 339/528 (64%), Gaps = 12/528 (2%) ref|NP_903600.11 L-aspartate oxidase [Chromobacterium violaceum ATCC 12472] gb|AAQ61592.11 L-aspartate oxidase [Chromobacterium violaceum ATCC 12472] Length = 529
2631.4 Best-BlastP=> >nrprot 94% Identities = 875/974 (89%), Positives = 928/974 (95%) sp|Q48815|HELA_LEGPN Protein helA gb|AAB06679.11 HelA Length = 1052
2637.1 Best-BlastP=> >nrprot No Hits found
2639.1 Best-BlastP=> >nrprot 53% Identities = 52/149 (34%), Positives = 80/149 (53%), Gaps = 14/149 (9%) ref|NP_747492.11 hypothetical protein [Pseudomonas putida KT2440] gb|AAN70956.1 |AE016739_9 hypothetical protein [Pseudomonas putida KT2440] Length = 193
264.2 Best-BlastP=> >nrprot 72% Identities = 264/467 (67%), Positives = 333/467 (72%), Gaps = 1/457 (0%) ref|ZP_00066450.11 COG0015: Adenylosuccinate lyase [Microbulbifer degradans 2-40] Length = 459
2643.1 Best-BlastP=> >nrprot 44% Identities = 119/434 (27%), Positives = 193/434 (44%), Gaps = 26/434 (5%) ref|NP_564271.11 expressed protein [Arabidopsis thaliana] gb|AAF79860.1 |AC000348_13 T7N9.21 [Arabidopsis thaliana] Length = 468
2644.1 Best-BlastP=> >nrprot No Hits found 2645.1 Best-BlastP=> >nrprot 24% Identities = 53/135 (39%), Positives = 80/135 (59%), Gaps = 12/135 (8%) ref|ZP_00014821.1 | hypothetical protein [Rhodospirillum rubrum] Length = 149
2646.1 Best-BlastP=> >nrprot 49% Identities = 45/117 (38%), Positives = 63/117 (53%), Gaps = 1/117 (0%) ref|NP_386943.11 CONSERVED HYPOTHETICAL PROTEIN [Sinorhizobium meliloti] emb|CAC47416.1 | CONSERVED HYPOTHETICAL PROTEIN [Sinorhizobium meliloti] Length = 253
2647.2 Best-BlastP=> >nrprot 72% Identities = 102/185 (55%), Positives = 138/185 (74%), Gaps = 1/185 (0%) ref|NP_621791.1 | 3-Methyladenine DNA glycosylase [Thermoanaerobacter tengcongensis] gb|AAM23395.11 3-Methyladenine DNA glycosylase [Thermoanaerobacter tengcongensis] Length = 188
2648.2 Best-BlastP=> >nrprot 40% Identities = 98/337 (29%), Positives = 148/337 (43%), Gaps = 67/337 (19%) pir||A42596 major outer membrane protein - Legionella pneumophila gb|AAA25300.11 major outer membrane protein Length = 2 7
2649.1 Best-BlastP=> >nrprot 45% Identities = 99/375 (26%), Positives = 182/375 (48%), Gaps = 2/375 (0%) ref |ZP_00026377.11 COG0475: Kef-type K+ transport systems, membrane components [Ralstonia metallidurans] Length = 406
2660.1 Best-BlastP=> >nrprot 64% Identities = 199/424 (46%), Positives = 284/424 (66%), Gaps = 14/424 (3%) ref|ZP_00031775.1 | COG1253: Hemolysins and related proteins containing CBS domains [Burkholderia fungorum] Length = 427
2652.2 Best-BlastP=> >nrprot 46% Identities = 119/354 (33%), Positives = 193/354 (54%), Gaps = 6/354 (1 %) gb|AAM00606.11 unknown [Legionella pneumophila] Length = 421
2653.1 Best-BlastP=> >nrprot No Hits found
2654.1 Best-BlastP=> >nrprot No Hits found
2655.1 Best-BlastP=> >nrprot 50% Identities = 36/105 (33%), Positives = 56/105 (53%), Gaps = 6/105 (5%) dbj|BAC93400.11 conserved hypothetical protein [Vibrio vulnificus YJ016] Length = 142
2657.3 Best-BlastP=> >nrprot 61 % Identities = 324/677 (47%), Positives = 456/677 (67%), Gaps = 12/677 (1 %) ref|ZP_00054877.11 COG0204: 1 -acyl- sn-glycerol-3-phosphate acyltransferase [Magnetospirillum magnetotacticum] Length = 1158
2659.2 Best-BlastP=> >nrprot 32% Identities = 34/116 (29%), Positives = 54/116 (46%), Gaps = 12/116 (10%) ref|NP_052947.1 | prepropilin [Plasmid R100] sp|P14494|PILδ_ECOLI FIMBRIAL PROTEIN PRECURSOR (PILIN) pir||YQECR1 fimbrial protein precursor - Escherichia coli plasmid R100-1 gb|AAA92754.1 | pilin dbj|BAA78861.1 | prepropilin [Plasmid R100] Length = 119
2660.1 Best-BlastP=> >nrprot 42% Identities = 24/86 (27%), Positives = 41/86 (47%) sp|P12058|TRAL_SALTI TRAL PROTEIN pir||C25161 traL protein - Salmonella typhimurium plasmid pED208 gb|AAA25608.1 | TraL protein [Plasmid pED208] gb|AAMg0704.1 | TraL [Salmonella typhi] Length = 101
2661.2 Best-BlastP=> >nrprot 45% Identities = 44/183 (24%), Positives = 85/183 (46%) ref|NP_932206.11 putative conjugative transfer protein TraE [Vibrio vulnificus YJ016] dbj|BAC97729.11 putative conjugative transfer protein TraE [Vibrio vulnificus YJ016] Length = 200
2662.2 Best-BlastP=> >nrprot 45% Identities = 72/242 (29%), Positives = 109/242 (45%), Gaps = 19/242 (7%) ref|NP_762δ87.11 Unknown [Vibrio vulnificus CMCP6] gb|AAO07577.1 |AE016810_80 Unknown [Vibrio vulnificus CMCP6] Length = 247
2666.1 Best-BlastP=> >nrprot No Hits found
2666.1 Best-BlastP=> >nrprot No Hits found
2667.1 Best-BlastP=> >nrprot No Hits found
2669.2 Best-BlastP=> >nrprot 20% Identities = 36/164 (23%), Positives = 71/164 (46%), Gaps = 6/154 (3%) ref|XP_223341.2| similar to KIAA0635 gene product [Rattus norvegicus] Length = 1266
267.3 Best-BlastP=> >nrprot 68% Identities = 167/307 (54%), Positives = 216/307 (70%), Gaps = 5/307 (1 %) ref|NP_820081.11 tRNA delta(2)- isopentenylpyrophosphate transferase [Coxiella burnetii RSA 493] gb|AAO90595.11 tRNA delta(2)-isopentenylpyrophosphate transferase [Coxiella burnetii RSA 493] Length = 311
2670.1 Best-BlastP=> >nrprot 16% Identities = 43/143 (30%), Positives = 70/143 (48%), Gaps = 6/143 (4%) ref|NP_842098.11 possible flagellar hook- ' length control protein [Nitrosomonas europaea ATCC 19718] emb|CAD85999.11 possible flagellar hook-length control protein [Nitrosomonas europaea ATCC 19718] Length = 381
2671.1 Best-BlastP=> >nrprot 98% Identities = 203/208 (97%), Positives = 205/208 (98%) sp|P37033|YAC1_LEGPN Hypothetical 23.7 kDa protein in ACN 5'region pir||A48642 hypothetical protein (acn 5' region) - Legionella pneumophila gb|AAA26294.1 | putative Length = 208
2674.1 Best-BlastP=> >nrprot 99% Identities = 885/891 (99%), Positives = 889/891 (99%) sp|P37032|ACON_LEGPN Aconitate hydratase (Citrate hydr lyase) (Aconitase) (Major iron-containing protein) (MICP) (IP210) pir||B48642 aconitate hydratase (EC 4.2.1.3) - Legionella pneumophila gb|AAA2529δ.11 aconitase Length = 891
2677.2 Best-BlastP=> >nrprot 70% Identities = 231/451 (51 %), Positives = 328/461 (72%), Gaps = 1/451 (0%) ref|NP_820994.11 amino acid antiporter [Coxiella burnetii RSA 493] gb|AA091508.11 amino acid antiporter [Coxiella burnetii RSA 493] Length = 476
268.1 Best-BlastP=> >nrprot 63% Identities = 39/66 (59%), Positives = 45/66 (68%), Gaps = 2/66 (3%) ref|NP_643775.11 conserved hypothetical protein [Xanthomonas axonopodis pv. citri str. 306] gb|AAM38311.11 conserved hypothetical protein [Xanthomonas axonopodis pv. citri str. 306] Length = 72
2680.1 Best-BlastP=> >nrprot 56% Identities = 93/186 (50%), Positives = 128/186 (68%) ref |NP_819966.11 conserved hypothetical protein [Coxiella burnetii RSA 493] gb|AAO90480.11 conserved hypothetical protein [Coxiella burnetii RSA 493] Length = 414
2682.1 Best-BlastP=> >nrprot 30% Identities = 76/390 (19%), Positives = 169/390 (43%), Gaps = 77/390 (19%) ref|NP_703923.11 hypothetical protein [Plasmodium falciparum 3D7] emb|CAD50δ35.11 hypothetical protein [Plasmodium falciparum 3D7] Length = 947
2683.3 Best-BlastP=> >nrprot 64% Identities = 253/483 (52%), Positives = 321/483 (66%), Gaps = 1/483 (0%) ref |NP_791669.11 succinylglutamic semialdehyde dehydrogenase [Pseudomonas syringae pv. tomato str. DC3000] gb|AA05δ354.11 succinylglutamic semialdehyde dehydrogenase [Pseudomonas syringae pv. tomato str. DC3000] Length = 488
2686.2 Best-BlastP=> >nrprot 32% Identities = 47/192 (24%), Positives = 86/192 (44%), Gaps = 4/192 (2%) ref|ZP_00031024.1 | COG0421 : Spermidine synthase [Burkholderia fungorum] Length = 252
2688.2 Best-BlastP=> >nrprot No Hits found
269.1 Best-BlastP=> >nrprot 63% Identities = 159/347 (45%), Positives = 224/347 (64%), Gaps = 2/347 (0%) ref|NP_820979.11 heptosyl transferase, glycosyltransferase family 9 protein [Coxiella burnetii RSA 493] gb|AA091493.11 heptosyl transferase, glycosyltransferase family 9 protein [Coxiella burnetii RSA 493] Length = 351
2690.2 Best-BlastP=> >nrprot No Hits found
2695.2 Best-BlastP=> >nrprot 45% Identities = 55/141 (39%), Positives = 89/141 (63%), Gaps = 6/141 (4%) gb|AAN86353.11 unknown [Listonella pelagia] Length = 195
2696.2 Best-BlastP=> >nrprot 70% Identities = 241/483 (49%), Positives = 342/483 (70%), Gaps = 10/483 (2%) ref|ZP_00117991.11 COG0433: Predicted ATPase [Cytophaga hutchinsonii] Length = 517
2698.1 Best-BlastP=> >nrprot 69% Identities = 64/127 (50%), Positives = 89/127 (70%) ref |NP_522810.11 CONSERVED HYPOTHETICAL PROTEIN [Ralstonia solanacearum] emb|CAD18400.11 CONSERVED HYPOTHETICAL PROTEIN [Ralstonia solanacearum] Length = 127
27.1 Best-BlastP=> >nrprot 37% Identities = 41/149 (27%), Positives = 82/149 (55%), Gaps = 2/149 (1 %) ref|NP_644250.11 hypothetical protein [Xanthomonas axonopodis pv. citri str. 306] gb|AAM38786.11 hypothetical protein [Xanthomonas axonopodis pv. citri str. 306] Length = 211
270.1 Best-BlastP=> >nrprot 61 % Identities = 125/278 (44%), Positives = 170/278 (61%), Gaps = 14/278 (5%) gb|AAH53853.1 | MGC16638 protein [Homo sapiens] Length = 291
2700.2 Best-BlastP=> >nrprot 45% Identities = 112/367 (30%), Positives = 187/367 (60%), Gaps = 6/367 (1 %) ref|NP_907216.1 | PUTATIVE EFFLUX PROTEIN [Wolinella succinogenes] emb|CAE10116.1 | PUTATIVE EFFLUX PROTEIN [Wolinella succinogenes] Length = 397
2702.2 Best-BlastP=> >nrprot 66% Identities = 103/291 (35%), Positives = 165/291 (56%), Gaps = 7/291 (2%) ref|NP_799583.11 putative transcriptional regulator [Vibrio parahaemolyticus RIMD 2210633] dbj|BAC61416.11 putative transcriptional regulator [Vibrio parahaemolyticus] Length = 289
2704.4 Best-BlastP=> >nrprot 45% Identities = 135/395 (34%), Positives = 213/395 (53%), Gaps = 17/395 (4%) ref|NP_865607.11 probable sensor protein fixL [Pirellula sp.] emb|CAD73291.11 probable sensor protein fixL [Pirellula sp.] Length = 651
2705.3 Best-BlastP=> >nrprot 61 % Identities = 173/388 (44%), Positives = 239/388 (61 %), Gaps = 16/388 (4%) ref|ZP_00054896.11 COG3287: Uncharacterized conserved protein [Magnetospirillum magnetotacticum] Length = 376
2706.2 Best-BlastP=> >nrprot 24% Identities = 72/75 (96%), Positives = 72/75 (96%) gb|AA061480.11 unknown [Legionella pneumophila] Length = 77
2709.1 Best-BlastP=> >nrprot 64% Identities = 158/320 (49%), Positives = 212/320 (66%), Gaps = 2/320 (0%) ref|NP_245036.11 unknown [Pasteurella multocida] gb|AAK02183.1 | unknown [Pasteurella multocida] Length = 337
271.1 Best-BlastP=> >nrprot 2g% Identities = 46/133 (33%), Positives = 79/133 (69%), Gaps = 3/133 (2%) ref|NP_149698.11 235L [Invertebrate iridescent virus 6] gb|AAK82096.1 |AF303741_235 236L [Chilo iridescent virus] Length = 265
2712.1 Best-BlastP=> >nrprot 29% Identities = 57/175 (32%), Positives = 81/175 (46%), Gaps = 13/175 (7%) ref|NP_231966.11 hypothetical protein VC2335 [Vibrio cholerae 01 biovar eltor str. N 16961] pir||E82090 hypothetical protein VC2335 [imported] - Vibrio cholerae (strain N16961 serogroup 01 ) gb|AAF95479.1 | hypothetical protein [Vibrio cholerae 01 biovar eltor str. N16961] Length = 195
2713.2 Best-BlastP=> >nrprot 52% Identities = 107/319 (33%), Positives = 178/319 (56%), Gaps = 14/319 (4%) ref|NP_441143.11 unknown protein [Synechocystis sp. PCC 6803] sp|P73771 |YB64_SYNY3 Hypothetical transport protein sll1164 pir||S74862 hypothetical protein sll1164 - Synechocystis sp. (strain PCC 6803) dbj|BAA17823.11 ORF_ID:sll1164~unknown protein [Synechocystis sp. PCC 6803] Length = 349
2717.1 Best-BlastP=> >nrprot No Hits found
2718.1 Best-BlastP=> >nrprot 71 % Identities = 254/438 (57%), Positives = 324/438 (73%), Gaps = 10/438 (2%) ref|NP_220059.11 Hexosphosphate Transport [Chlamydia trachomatis] sp|084548|UHPT_CHLTR Probable hexose phosphate transport protein pir||A71501 probable hexosphosphate transport - Chlamydia trachomatis (serotype D, strain UW3/Cx) gb|AAC68146.11 Hexosphosphate Transport [Chlamydia trachomatis] Length = 456
2721.3 Best-BlastP=> >nrprot 67% Identities = 383/736 (52%), Positives = 502/736 (68%), Gaps = 10/736 (1 %) ref|ZP_00095016.1 | COG0022: Pyruvate/2-oxoglutarate dehydrogenase complex, dehydrogenase (E1 ) component, eukaryotic type, beta subunit [Novosphingobium aromaticivorans] Length = 738
2724.3 Best-BlastP=> >nrprot 45% Identities = 96/404 (23%), Positives = 177/404 (43%), Gaps = 76/404 (18%) ref|ZP_00083983.11 hypothetical protein [Pseudomonas fluorescens PfO-1] Length = 524
2725.1 Best-BlastP=> >nrprot 62% Identities = 39/75 (52%), Positives = 56/7δ (74%) ref|NP_420714.11 flhB-related protein [Caulobacter crescentus CB1δ] pir||F8748δ flhB-related protein [imported] - Caulobacter crescentus gb|AAK23882.1 | flhB-related protein [Caulobacter crescentus CB16] Length = 87
2726.1 Best-BlastP=> >nrprot No Hits found
2727.2 Best-BlastP=> >nrprot 16% Identities = 70/336 (20%), Positives = 147/336 (43%), Gaps = 50/336 (14%) gb|AAL99918.1 |AF432211_1 CLL- associated antigen KW-11 [Homo sapiens] Length = 460
2728.1 Best-BlastP=> >nrprot 61% Identities = 49/1U3 (47%), Positives = 65/103 (b37o) ret|NP_903992.1 | conserved hypothetical protein [Chromobacterium violaceum ATCC 12472] gb|AAQ61981.11 conserved hypothetical protein [Chromobacterium violaceum ATCC 12472] Length = 117
273.1 Best-BlastP=> >nrprot No Hits found
2730.1 Best-BlastP=> >nrprot No Hits found
2732.1 Best-BlastP=> >nrprot No Hits found
2733.1 Best-BlastP=> >nrprot 58% Identities = 133/345 (38%), Positives = 206/345 (59%), Gaps = 12/345 (3%) ref|NP_437388.11 putative conserved membrane-anchored protein [Sinorhizobium meliloti] pir||H95947 probable conserved membrane-anchored protein SMb21182 [imported] - Sinorhizobium meliloti (strain 1021 ) magaplasmid pSymB emb|CAC49248.1 | putative conserved membrane-anchored protein [Sinorhizobium meliloti] Length = 394
2737.2 Best-BlastP=> >nrprot No Hits found
2738.2 Best-BlastP=> >nrprot 43% Identities = δδ/255 (21 %), Positives = 112/255 (43%), Gaps = 16/255 (6%) ref|NP_900118.11 probable ABC transport system permease protein [Chromobacterium violaceum ATCC 12472] gb|AAQ58126.11 probable ABC transport system permease protein [Chromobacterium violaceum ATCC 12472] Length = 267
2739 3 Best-BlastP=> >nrprot 50% Identities = 67/256 (26%), Positives = 115/256 (44%), Gaps = 51/256 (19%) ret|NP_518722.11 PROBABLE TRANSMEMBRANE PROTEIN [Ralstonia solanacearum] emb|CAD14131.1 | PROBABLE TRANSMEMBRANE PROTEIN [Ralstonia solanacearum] Length = 547
2741.2 Best-BlastP=> >nrprot 52% Identities = 20/40 (50%), Positives = 24/40 (60%) ref|NP_907600.11 hypothetical protein WS1440 [Wolinella succinogenes] emb|CAE10600.1 | hypothetical protein [Wolinella succinogenes] Length = 42
2742.2 Best-BlastP=> >nrprot 77% Identities = 34/42 (80%), Positives = 34/42 (80%) ref|NP_769472.1 | Conserved hypothetical protein [Vibrio vulnificus CMCP6] ref|NP_75989δ.1 | Conserved hypothetical protein [Vibrio vulnificus CMCP6] ref|NP_759901.1 | Conserved hypothetical protein [Vibrio vulnificus CMCP6] ref |NP_760122.11 Conserved hypothetical protein [Vibrio vulnificus CMCP6] ref|NP_760328.1 | Conserved hypothetical protein [Vibrio vulnificus CMCP6] ref|NP_763326.11 Conserved hypothetical protein [Vibrio vulnificus CMCP6] gb|AAO08316.1 |AE016813_68 Conserved hypothetical protein [Vibrio vulnificus CMCP6] gb|AAO08g99.1 |AE016798_159 Conserved hypothetical protein [Vibrio vulnificus CMCP6] gb|AAO09422.1 |AE016800_27 Conserved hypothetical protein [Vibrio vulnificus CMCP6] gb|AAO09428.1 |AE016800_33 Conserved hypothetical protein [Vibrio vulnificus CMCP6] gb|AAO09649.1 |AE016800_254 Conserved hypothetical protein [Vibrio vulnificus CMCP6] gb|AAO0985δ.1 |AE016801_174 Conserved hypothetical protein [Vibrio vulnificus CMCP6] Length = 43
2743.2 Best-BlastP=> >nrprot 82% Identities = 62/66 (78%), Positives = 66/66 (84%) ref|NP_769471.11 Conserved hypothetical protein [Vibrio vulnificus CMCP6] ref|NP_759894.1 | Conserved hypothetical protein [Vibrio vulnificus CMCP6] ref|NP_759900.1 | Conserved hypothetical protein [Vibrio vulnificus CMCP6] ref|NP_759939.11 Conserved hypothetical protein [Vibrio vulnificus CMCP6] ref|NP_760021.11 Conserved hypothetical protein [Vibrio vulnificus CMCP6] ref|NP_760123.11 Conserved hypothetical protein [Vibrio vulnificus CMCP6] ref|NP_760329.11 Conserved hypothetical protein [Vibrio vulnificus CMCP6] ref|NP_760403.11 Conserved hypothetical protein [Vibrio vulnificus CMCP6] ref|NP_763327.11 Conserved hypothetical protein [Vibrio vulnificus CMCP6] gb|AAO08317.1 |AE016813_69 Conserved hypothetical protein [Vibrio vulnificus CMCP6] gb|AAO08998.1 |AE016798_158 Conserved hypothetical protein [Vibrio vulnificus CMCP6] gb|AAO09421.1 |AE016800_26 Conserved hypothetical protein [Vibrio vulnificus CMCP6] gb|AAO09427.1 |AE016800_32 Conserved hypothetical protein [Vibrio vulnificus CMCP6] gb|AAO09466.1 |AE016800_71 Conserved hypothetical protein [Vibrio vulnificus CMCP6] gb|AAO09548.1 |AE016800_1δ3 Conserved hypothetical
2746.1 Best-BlastP=> >nrprot No Hits found
2747.1 Best-BlastP=> >nrprot 62% Identities = 129/323 (39%), Positives = 203/323 (62%), Gaps = 4/323 (1 %) ref|NP_384238.11 PUTATIVE ADENOSINE DEAMINASE PROTEIN [Sinorhizobium meliloti] sp|Q92T48|ADD_RHIME Adenosine deaminase (Adenosine aminohydrolase) emb|CAC41519.1 | PUTATIVE ADENOSINE DEAMINASE PROTEIN [Sinorhizobium meliloti] Length = 324
2748.1 Best-BlastP=> >nrprot No Hits found
2749.4 Best-BlastP=> >nrprot 67% Identities = 204/41 δ (49%), Positives = 283/41 δ (68%) ref|NP_819070.11 phosphate transporter family protein [Coxiella burnetii RSA 493] gb|AA089584.11 phosphate transporter family protein [Coxiella burnetii RSA 493] Length = 417
2761.1 Best-BlastP=> >nrprot 48% Identities = 83/226 (36%), Positives = 126/226 (55%), Gaps = 5/226 (2%) ref|NP_212383.11 phosphatidyltransferase [Borrelia burgdorferi] pir||A70131 phosphatidyltransferase homolog - Lyme disease spirochete gb|AAB91497.11 phosphatidyltransferase [Borrelia burgdorferi B31] Length = 234
2752.1 Best-BlastP=> >nrprot 49% Identities = 53/167 (31 %), Positives = 81/167 (48%), Gaps = 14/167 (8%) ref|NP_819930.11 conserved domain protein [Coxiella burnetii RSA 493] gb|AAO90444.11 conserved domain protein [Coxiella burnetii RSA 493] Length = 169
2753.1 Best-BlastP=> >nrprot No Hits found
2754.2 Best-BlastP=> >nrprot 72% Identities = 109/184 (59%), Positives = 134/184 (72%) ref |NP_820271.1 | conserved hypothetical protein [Coxiella burnetii RSA 493] gb|AAO90785.11 conserved hypothetical protein [Coxiella burnetii RSA 493] Length = 186
2765.1 Best-BlastP=> >nrprot 29% Identities = 102/461 (22%), Positives = 198/461 (42%), Gaps = 56/461 (12%) ref|NP_788908.11 CG33206-PB [Drosophila melanogaster] gb|AAF48467.2| CG33206-PB [Drosophila melanogaster] Length = 1208
2757.1 Best-BlastP=> >nrprot 73% Identities = 78/148 (52%), Positives = 110/148 (74%) ref|ZP_00125260.11 COG0359: Ribosomal protein L9 [Pseudomonas syringae pv. syringae B728a] ref|NP_7g4663.1 | ribosomal protein L9 [Pseudomonas syringae pv. tomato str. DC3000] gb|AA0583δ8.1 | ribosomal protein L9 [Pseudomonas syringae pv. tomato str. DC3000] Length = 148
2769.1 Best-BlastP=> >nrprot 40% Identities = 74/283 (26%), Positives = 130/283 (46%), Gaps = 14/283 (4%) ref|NP_819886.11 membrane protein, putative [Coxiella burnetii RSA 493] gb|AAO90399.11 membrane protein, putative [Coxiella burnetii RSA 493] Length = 299
276.1 Best-BlastP=> >nrprot 26% Identities = 20/26 (80%), Positives = 20/25 (80%), Gaps = 1/25 (4%) ref|NP_084281.11 RIKEN cDNA A030006K14 [Mus musculus] dbj|BAB32176.1 | unnamed protein product [Mus musculus] Length = 122
2760.1 Best-BlastP=> >nrprot 85% Identities = 53/76 (70%), Positives = 6δ/7δ (86%) ref|NP_240367.11 30S ribosomal protein S18 [Buchnera aphidicola str. APS (Acyrthosiphon pisum)] sp|Pδ7626|RS18_BUCAI 30S ribosomal protein S18 pir||E84995 30S ribosomal protein S18 [imported] - Buchnera sp. (strain APS) dbj|BAB13253.11 30S ribosomal protein S18 [Buchnera aphidicola str. APS (Acyrthosiphon pisum)] Length = 75
2761.1 Best-BlastP=> >nrprot 75% Identities = 70/107 (65%), Positives = 86/107 (79%) ref|NP_903310.11 30S ribosomal protein S6 [Chromobacterium violaceum ATCC 12472] gb|AAQ61302.11 30S ribosomal protein S6 [Chromobacterium violaceum ATCC 12472] Length = 124 2763.2 Best-BlastP=> >nrprot No Hits found
2765.2 Best-BlastP=> >nrprot 33% Identities = 48/222 (21 %), Positives = 99/222 (44%), Gaps = 21/222 (9%) ref|NP_106991.1 | unknown protein [Mesorhizobium loti] dbj|BAB62777.1 | unknown protein [Mesorhizobium loti] Length = 328
2767.1 Best-BlastP=> >nrprot No Hits found
2769.2 Best-BlastP=> >nrprot 53% Identities = 203/694 (34%), Positives = 317/594 (53%), Gaps = 26/594 (4%) ref|NP_266737.11 hypothetical protein [Lactococcus lactis subsp. lactis] pir||E86697 conserved hypothetical protein yfhG [imported] - Lactococcus lactis subsp. lactis (strain IL1403) gb|AAK04670.1 |AE006291_13 conserved hypothetical protein [Lactococcus lactis subsp. lactis] Length = 598
277.1 Best-BlastP=> >nrprot 41 % Identities = 60/196 (30%), Positives = 99/196 (50%), Gaps = 18/196 (9%) ref|NP_436484.11 hypothetical protein [Sinorhizobium meliloti] pir||F95416 hypothetical protein SMa2299 [imported] - Sinorhizobium meliloti (strain 1021 ) magaplasmid pSymA gb|AAK65896.11 hypothetical protein [Sinorhizobium meliloti] Length = 308
2770.1 Best-BlastP=> >nrprot 65% Identities = 114/240 (47%), Positives = 163/240 (63%), Gaps = 14/240 (6%) ref|NP_867184.1 | short chain alcohol dehydrogenase-like [Pirellula sp.] emb|CAD74729.11 short chain alcohol dehydrogenase-like [Pirellula sp.] Length = 247
2771 -2 Best-BlastP=> >nrprot 66% Identities = 206/386 (53%), Positives = 260/386 (67%), Gaps = 1/386 (0%) ref|NP_355900.11 AGR_L_236p [Agrobacterium tumefaciens] ref|NP_536243.1 | conserved hypothetical protein [Agrobacterium tumefaciens str. C68 (U. Washington)] pir||C9814δ hypothetical protein AGR_L_236 [imported] - Agrobacterium tumefaciens (strain Cδ8, Cereon) pir||AI3142 conserved hypothetical protein Atu476δ [imported] - Agrobacterium tumefaciens (strain Cδ8, Dupont) gb|AAK8868δ.11 AGR_L_236p [Agrobacterium tumefaciens str. C58 (Cereon)] gb|AAL45559.11 conserved hypothetical protein [Agrobacterium tumefaciens str. C58 (U. Washington)] Length = 507
2772.1 Best-BlastP=> >nrprot 51 % Identities = 40/131 (30%), Positives = 71/131 (54%), Gaps = 12/131 (9%) ref|NP_469941.1 | Iin0598 [Listeria innocua] pir| |AF1607 hypothetical protein Iin0598 [imported] - Listeria innocua (strain Clip11262) emb|CAC95830.1 | Iin0598 [Listeria innocua] Length = 138
2774.1 Best-BlastP=> >nrprot 69% Identities = 152/286 (53%), Positives = 204/286 (71 %) ref|NP_697358.1 | transcriptional regulator, LysR family [Brucella suis 1330] gb|AAN29273.1 |AE014344_8 transcriptional regulator, LysR family [Brucella suis 1330] Length = 296
2775.1 Best-BlastP=> >nrprot 50% Identities = 31/52 (59%), Positives = 36/52 (69%), Gaps = 3/52 (5%) ref |ZP_00021959.11 COG3024: Uncharacterized protein conserved in bacteria [Ralstonia metallidurans] Length = 63
2777.1 Best-BlastP=> >nrprot 64% Identities = 174/343 (50%), Positives = 232/343 (67%), Gaps = 5/343 (1 %) ref|NP_780210.11 phosphoribosylaminoimidazole carboxylase, ATPase subunit [Xylella fastidiosa Temeculal] gb|AA029859.11 phosphoribosylaminoimidazole carboxylase, ATPase subunit [Xylella fastidiosa Temeculal] Length = 394
2778.2 Best-BlastP=> >nrprot 82% Identities = 113/160 (70%), Positives = 138/160 (86%) ref|ZP_00066669.11 COG0041 : Phosphoribosylcarboxyaminoimidazole (NCAIR) mutase [Microbulbifer degradans 2-40] Length = 168
278.3 Best-BlastP=> >nrprot 51% Identities = 218/623 (34%), Positives = 331/623 (53%), Gaps = 47/623 (7%) ref|NP_070765.11 phosphoribosylformylglycinamidine synthase II (purL) [Archaeoglobus fulgidus DSM 4304] sp|028339|PURL_ARCFU Phosphoribosylformylglycinamidine synthase II (FGAM synthase II) pir||C69492 phosphoribosylformylglycinamidine synthase (EC 6.3.5.3) component II - Archaeoglobus fulgidus gb|AAB89311.11 phosphoribosylformylglycinamidine synthase II (purL) [Archaeoglobus fulgidus DSM 4304] Length = 765
2780.2 Best-BlastP=> >nrprot 57% Identities = 116/273 (42%), Positives = 162/273 (59%), Gaps = 3/273 (1 %) ref|NP_925031.11 hypothetical protein glr208δ [Gloeobacter violaceus] dbj|BAC90026.11 glr2085 [Gloeobacter violaceus] Length = 288
2781.1 Best-BlastP=> >nrprot 70% Identities = 49/103 (47%), Positives = 76/103 (72%) ref|NP_213586.11 putative protein [Aquifex aeolicus] pir||F70374 hypothetical protein aq_862 - Aquifex aeolicus gb|AAC06993.11 putative protein [Aquifex aeolicus VF5] Length = 109
2784.3 Best-BlastP=> >nrprot 49% Identities = 36/80 (45%), Positives = 52/80 (65%) ref|NP_931069.11 hypothetical protein [Photorhabdus luminescens subsp. laumondii TT01] emb|CAE16236.1 | unnamed protein product [Photorhabdus luminescens subsp. laumondii TT01] Length = 124
2785.1 Best-BlastP=> >nrprot 71 % Identities = 163/265 (57%), Positives = 202/266 (76%), Gaps = 1/265 (0%) ref|NP_819150.11 nicotinate-nucleotide pyrophosphorylase [Coxiella burnetii RSA 493] gb|AA089664.11 nicotinate-nucleotide pyrophosphorylase [Coxiella burnetii RSA 493] Length = 274
2788.1 Best-BlastP=> >nrprot 54% Identities = 168/473 (33%), Positives = 263/473 (56%), Gaps = 39/473 (8%) ref|NP_485346.11 Na+/H+ antiporter [Nostoc sp. PCC 7120] pir||AD1969 Na+/H+ antiporter [imported] - Nostoc sp. (strain PCC 7120) dbj|BAB73260.1 | Na+/H+ antiporter [Nostoc sp. PCC 7120] Length = 470
2791.1 Best-BlastP=> >nrprot 82% Identities = 637/779 (68%), Positives = 666/779 (84%), Gaps = δ/779 (0%) ref|NP_310436.11 phosphoenolpyruvate synthase [Escherichia coli 0157:H7] pir||A90930 phosphoenolpyruvate synthase [imported] - Escherichia coli (strain 01 δ7:H7, substrain RIMD 0609962) dbj|BAB36832.11 phosphoenolpyruvate synthase [Escherichia coli 0167:H7] Length = 792
2792.1 Best-BlastP=> >nrprot No Hits found
2793.1 Best-BlastP=> >nrprot 60% Identities = 100/343 (29%), Positives = 170/343 (49%), Gaps = 28/343 (8%) ref|NP_233263.11 hydrolase, putative [Vibrio cholerae 01 biovar eltor str. N16961] pir||E82406 probable hydrolase VCA0877 [imported] - Vibrio cholerae (strain N16961 serogroup 01 ) gb|AAF96775.11 hydrolase, putative [Vibrio cholerae 01 biovar eltor str. N16961] Length = 358
2794.2 Best-BlastP=> >nrprot 44% Identities = 152/632 (28%), Positives = 250/532 (46%), Gaps = 31/532 (5%) ref|ZP_00029700.11 COG1960: Acyl- CoA dehydrogenases [Burkholderia fungorum] Length = 587
2795.1 Best-BlastP=> >nrprot 87% Identities = 60/88 (68%), Positives = 78/88 (88%) ref|NP_395197.11 putative transposase ORFA [Yersinia pestis C092] ref|NP_867719.11 low calcium response locus protein S homolog [Yersinia pestis KIM] ref|NP_867914.11 putative IS element protein [Yersinia pestis KIM] sp|Q00931 |LCRS_YERPE Low calcium response locus protein S pir||T43662 probable IS element protein - Yersinia pestis plasmid pCD1 gb|AAA2765δ.11 IcrS gb|AAC62δ7g.11 low calcium response locus protein S homolog [Yersinia pestis KIM] gb|AAC69827.11 putative IS element protein [Yersinia pestis KIM] emb|CAB64940.11 putative transposase ORFA [Yersinia pestis] Length = 88
2799.1 Best-BlastP=> >nrprot 67% Identities = 71/126 (56%), Positives = 91/126 (72%) ref|NP_867058.11 aspartate 1 -decarboxylase [Pirellula sp.] emb|CAD74603.1 | aspartate 1 -decarboxylase [Pirellula sp.] Length = 156
2800.1 Best-BlastP=> >nrprot 66% Identities = 136/258 (52%), Positives = 172/258 (66%), Gaps = 3/258 (1 %) ref|NP_719181.11 dimethyladenosine transferase [Shewanella oneidensis MR-1] sp|Q8EB93|KSGA_SHEON Dimethyladenosine transferase (S-adenosylmethionine-6-N', N'- adenosyl(rRNA) dimethyltransferase) (16S rRNA dimethylase) (High level kasugamycin resistance protein ksgA) (Kasugamycin dimethyltransferase) gb|AAN5662δ.1 |AE016799_12 dimethyladenosine transferase [Shewanella oneidensis MR-1] Length = 268
2804.1 Best-BlastP=> >nrprot 63% Identities = 232/443 (52%), Positives = 286/443 (64%), Gaps = 2/443 (0%) ref|NP_629805.11 putative 4- aminobutyrate aminotransferase [Streptomyces coelicolor A3(2)] pir||T3δ794 probable 4-aminobutyrate aminotransferase - Streptomyces coelicolor emb|CAA20213.1 | putative 4-aminobutyrate aminotransferase [Streptomyces coelicolor A3(2)] Length = 444
2806.1 Best-BiastP=> >nrprot 76% Identities = 118/196 (60%), Positives = 153/196 (78%) ref|NP_249757.11 probable short-chain dehydrogenase [Pseudomonas aeruginosa PA01] pir||H83612 probable short-chain dehydrogenase PA1066 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG0445δ.1 |AE004538_7 probable short-chain dehydrogenase [Pseudomonas aeruginosa PA01] Length = 218
2807.1 Best-BlastP=> >nrprot 71% Identities = 64/113 (47%), Positives = 81/113 (71 %), Gaps = 6/113 (4%) ref|NP_798658.1 | arsenate reductase [Vibrio parahaemolyticus RIMD 2210633] dbj|BAC60642.11 arsenate reductase [Vibrio parahaemolyticus] Length = 116
2808.1 Best-BlastP=> >nrprot 60% Identities = 39/83 (46%), Positives = 56/83 (67%), Gaps = 1/83 (1%) gb|AAL07519.11 RNA-binding protein precursor [Solanum tuberosum] Length = 339
2810.1 Best-BlastP=> >nrprot 48% Identities = 130/457 (28%), Positives = 218/457 (47%), Gaps = 24/457 (5%) emb|CAD48863.1 | EefC outer membrane protein [Enterobacter aerogenes] Length = 454
2811.2 Best-BlastP=> >nrprot 25% Identities = 43/179 (24%), Positives = 89/179 (49%), Gaps = 4/179 (2%) ref|NP_820323.11 conserved domain protein [Coxiella burnetii RSA 493] gb|AAO90837.11 conserved domain protein [Coxiella burnetii RSA 493] Length = 377
2815.2 Best-BlastP=> >nrprot 9% Identities = 31/89 (34%), Positives = 64/89 (60%) ref |NP_819452.1 1 hypothetical protein [Coxiella burnetii RSA 493] gb|AA089966.11 hypothetical protein [Coxiella burnetii RSA 493] Length = 262
2816.2 Best-BlastP=> >nrprot 50% Identities = 191/730 (26%), Positives = 361/730 (49%), Gaps = 43/730 (5%) ref|ZP_00055987.11 COG0659: Sulfate permease and related transporters (MFS superfamily) [Magnetospirillum magnetotacticum] Length = 733
2818.1 Best-BlastP=> >nrprot 61 % Identities = 156/372 (41 %), Positives = 219/372 (58%), Gaps = 37/372 (9%) gb|AAK97454.1 |AF388182_2 alkane-1 - monooxygenase [Rhodococcus sp. Q15] Length = 408
282.2 Best-BlastP=> >nrprot 53% Identities = 58/179 (32%), Positives = 103/179 (57%), Gaps = 12/179 (6%) ref|NP_900154.11 probable type IV prepilin [Chromobacterium violaceum ATCC 12472] gb|AAQ68161.1 | probable type IV prepilin [Chromobacterium violaceum ATCC 12472] Length = 18δ
2820.1 Best-BlastP=> >nrprot 64% Identities = 146/273 (63%), Positives = 183/273 (67%), Gaps = 4/273 (1 %) ref|NP_439796.1 | hypothetical protein [Haemophilus influenzae Rd] sp|P46298|YRAL_HAEIN Hypothetical protein HI1654 pir||A64174 hypothetical protein HI1654 - Haemophilus influenzae (strain Rd KW20) gb|AAC23298.1 | conserved hypothetical protein [Haemophilus influenzae Rd] Length = 283
2822.1 Best-BlastP=> >nrprot 62% Identities = 202/610 (33%), Positives = 316/610 (61 %), Gaps = 29/610 (4%) ref|ZP_00137912.11 COG3107: Putative lipoprotein [Pseudomonas aeruginosa UCBPP-PA14] Length = 604
2824.1 Best-BlastP=> >nrprot 72% Identities = 83/138 (60%), Positives = 102/138 (73%) ref|NP_254240.1 1 ATP synthase epsilon chain [Pseudomonas aeruginosa PA01] sp|Q9HT21 |ATPE_PSEAE ATP synthase epsilon chain (ATP synthase F1 sector epsilon subunit) pir||B82962 ATP synthase epsilon chain PA6653 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG08938.1 |AE004967_9 ATP synthase epsilon chain [Pseudomonas aeruginosa PA01 ] Length = 141
2826.1 Best-BlastP=> >nrprot 42% Identities = 110/640 (20%), Positives = 217/640 (40%), Gaps = 94/640 (17%) ref |ZP_00144200.1 | EXONUCLEASE SBCC [Fusobacterium nucleatum subsp. vincentii ATCC 49256] gb|EAA24200.1 | EXONUCLEASE SBCC [Fusobacterium nucleatum subsp. vincentii ATCC 49256] Length = 921
2827.3 Best-BlastP=> >nrprot No Hits found
2828.3 Best-BlastP=> >nrprot 72% Identities = 29/52 (55%), Positives = 36/52 (69%), Gaps = 2/52 (3%) ref|NP_885117.11 phosphafidylserine decarboxylase proenzyme [Bordetella parapertussis] emb|CAE38217.1 | phosphatidylserine decarboxylase proenzyme [Bordetella parapertussis] Length = 328
283.1 Best-BlastP=> >nrprot 47% Identities = 52/142 (36%), Positives = 86/142 (60%), Gaps = 8/142 (5%) ref |NP_900149.11 probable type-4 fimbrial biogenesis protein [Chromobacterium violaceum ATCC 12472] gb|AAQ58156.1 | probable type-4 fimbrial biogenesis protein [Chromobacterium violaceum ATCC 12472] Length = 159
2830.3 Best-BlastP=> >nrprot No Hits found
2831.1 Best-BlastP=> >nrprot 47% Identities = 71/219 (32%), Positives = 106/219 (48%), Gaps = 10/219 (4%) ref|NP_801239.11 transcriptional regulator, LuxR family [Vibrio parahaemolyticus RIMD 2210633] dbj|BAC63072.11 transcriptional regulator, LuxR family [Vibrio parahaemolyticus] Length = 251
2833.1 Best-BlastP=> >nrprot 56% Identities = 125/332 (37%), Positives = 186/332 (56%), Gaps = 22/332 (6%) sp|Q56686|CPDP_VIBFI S'.δ'-CYCLIC- NUCLEOTIDE PHOSPHODIESTERASE PRECURSOR (PDEASE) (3':δ'-CNP) pir||A40602 3',5'-cyclic-nucleotide phosphodiesterase (EC 3.1.4.17) - Vibrio fischeri gb|AAA27613.1 | cpdP Length = 330
2836.1 Best-BlastP=> >nrprot 70% Identities = 161/281 (53%), Positives = 200/281 (71%), Gaps = 1/281 (0%) ref|NP_754369.11 Putative 3-hydroxyacyl- CoA dehydrogenase [Escherichia coli CFT073] gb|AAN80926.1 |AE016762_179 Putative 3-hydroxyacyl-CoA dehydrogenase [Escherichia coli CFT073] Length = 289
2838.1 Best-BlastP=> >nrprot 67% Identities = 176/365 (48%), Positives = 253/365 (69%) ref|ZP_00110259.11 COG1960: Acyl-CoA dehydrogenases [Nostoc punctiforme] Length = 395
284.2 Best-BlastP=> >nrprot 57% Identities = 128/361 (35%), Positives = 203/361 (56%), Gaps = 24/361 (6%) ref |NP_900150.11 conserved hypothetical protein [Chromobacterium violaceum ATCC 12472] gb|AAQ58157.1 | conserved hypothetical protein [Chromobacterium violaceum ATCC 12472] Length = 354
2840.1 Best-BlastP=> >nrprot 62% Identities = 225/602 (44%), Positives = 316/502 (62%), Gaps = g/502 (1 %) pir||T44808 mycosubtilin synthetase mycC [imported] - Bacillus subtilis gb|AAF08797.1 |AF184956_4 MycC [Bacillus subtilis] Length = 2609
2841.4 Best-BlastP=> >nrprot 58% Identities = 176/439 (39%), Positives = 259/439 (58%), Gaps = 14/439 (3%) ref|ZP_00111186.11 COG3321 : Polyketide synthase modules and related proteins [Nostoc punctiforme] Length = 1853
2847.4 Best-BlastP=> >nrprot No Hits found
2848.4 Best-BlastP=> >nrprot No Hits found
2849.1 Best-BlastP=> >nrprot No Hits found
2858.1 Best-BlastP=> >nrprot 28% Identities = 23/33 (69%), Positives = 24/33 (72%) pir||A44803 pG1 protein - human (fragment) Length = 75
2859.2 Best-BlastP=> >nrprot 76% Identities = 254/393 (64%), Positives = 308/393 (78%) ref|NP_716935.11 tyrosyl-tRNA synthetase [Shewanella oneidensis MR-1] gb|AAN54380.1 |AE01δδ7δ_6 tyrosyl-tRNA synthetase [Shewanella oneidensis MR-1] Length = 398
286.2 Best-BlastP=> >nrprot 48% Identities = 48/165 (29%), Positives = 83/165 (50%), Gaps = 18/165 (10%) ref|NP_900151.11 hypothetical protein CV0481 [Chromobacterium violaceum ATCC 12472] gb|AAQ58168.1 | hypothetical protein CV0481 [Chromobacterium violaceum ATCC 12472] Length = 168
2861.1 Best-BlastP=> >nrprot 67% Identities = 131/262 (50%), Positives = 185/262 (70%), Gaps = 1/262 (0%) ref|ZP_00068001.11 COG3220: Uncharacterized protein conserved in bacteria [Microbulbifer degradans 2-40] Length = 290
2862.1 Best-BlastP=> >nrprot 52% Identities = 142/396 (35%), Positives = 210/396 (53%), Gaps = 7/396 (1 %) ref|NP_349992.1 | Dipeptidyl aminopeptidase/acylaminoacyl-peptidase related protein [Clostridium acetobutylicum] pir||A97318 dipeptidyl aminopeptidase/acylaminoacyl- peptidase related protein [imported] - Clostridium acetobutylicum gb|AAK81332.1 |AE007837_10 Dipeptidyl aminopeptidase/acylaminoacyl- peptidase related protein [Clostridium acetobutylicum] Length = 400
2863.1 Best-BlastP=> >nrprot 35% Identities = 46/117 (39%), Positives = 68/117 (58%), Gaps = 4/117 (3%) ref|ZP_00008611.11 COG0845: Membrane- fusion protein [Rhodopseudomonas palustris] Length = 267
2866.1 Best-BlastP=> >nrprot 78% Identities = 276/431 (64%), Positives = 347/431 (80%), Gaps = 1/431 (0%) ref|NP_901043.11 probable oxidoreductase [Chromobacterium violaceum ATCC 12472] gb|AAQ69048.1 | probable oxidoreductase [Chromobacterium violaceum ATCC 12472] Length = 438
2868.1 Best-BlastP=> >nrprot 60% Identities = 247/686 (42%), Positives = 357/685 (61 %), Gaps = 15/685 (2%) ref|NP_820704.11 thiohdisulfide interchange protein DsbD [Coxiella burnetii RSA 493] gb|AA091218.11 thiohdisulfide interchange protein DsbD [Coxiella burnetii RSA 493] Length = 584
2869.1 ' Best-BlastP=> >nrprot 98% Identities = 96/96 (100%), Positives = 96/96 (100%) sp|P26879|CH10_LEGPN 10 kDa chaperonin (Protein Cpn10) (groES protein) (Heat shock protein A) pir||B41468 heat shock protein groES - Legionella pneumophila gb|AAA25297.1 | htpA Length = 96 .
2873.1 Best-BlastP=> >nrprot 40% Identities = 66/274 (24%), Positives = 130/274 (47%), Gaps = 16/274 (5%) ref|NP_692096.11 hypothetical protein [Oceanobacillus iheyensis HTE831] dbj|BAC13131.1 | hypothetical conserved protein [Oceanobacillus iheyensis HTE831] Length = 314
2874.1 Best-BlastP=> >nrprot 59% Identities = 111/225 (49%), Positives = 165/226 (68%), Gaps = 1/225 (0%) ref |NP_662190.11 hypothetical protein [Chlorobium tepidum TLS] gb|AAM72532.11 hypothetical protein [Chlorobium tepidum TLS] Length = 232
2877.2 Best-BlastP=> >nrprot 49% Identities = 71/206 (34%), Positives = 109/206 (52%), Gaps = 3/206 (1 %) ref|NP_902629.11 hypothetical protein CV2959 [Chromobacterium violaceum ATCC 12472] gb|AAQ60627.1| hypothetical protein CV2959 [Chromobacterium violaceum ATCC 12472] Length = 214
2878.2 Best-BlastP=> >nrprot 98% Identities = 226/230 (98%), Positives = 228/230 (99%) gb|AAK00284.1 |AF288536_6 possible transcriptional regulatory protein [Legionella longbeachae] Length = 230
2879.3 Best-BlastP=> >nrprot 96% Identities = 419/444 (94%), Positives = 430/444 (96%), Gaps = 2/444 (0%) gb|AAK00283.1 |AF288536_5 unknown [Legionella longbeachae] Length = 444
2881.2 Best-BlastP=> >nrprot 93% Identities = 270/302 (89%), Positives = 281/302 (93%), Gaps = 1/302 (0%) gb|AAK00282.1 |AF288δ36_4 unknown [Legionella longbeachae] Length = 302
2882.2 Best-BlastP=> >nrprot 76% Identities = 146/221 (66%), Positives = 175/221 (79%), Gaps = 1/221 (0%) ref|NP_249667.11 conserved hypothetical protein [Pseudomonas aeruginosa PA01] ref|ZP_00138566.11 COG0603: Predicted PP-loop superfamily ATPase [Pseudomonas aeruginosa UCBPP-PA14] pir||E83522 conserved hypothetical protein PA0g76 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG04365.1 |AE004531_2 conserved hypothetical protein [Pseudomonas aeruginosa PA01] gb|AAP82946.1 | conserved hypothetical protein [Pseudomonas aeruginosa] Length = 224
2884.1 Best-BlastP=> >nrprot No Hits found 2889.2 Best-BlastP=> >nrprot 69% Identities = 72/124 (68%), Positives = 90/124 (72%) ref |NP_719248.11 membrane protein, putative [Shewanella oneidensis MR-1] gb|AANδ6692.1 |AE01580δ_1 membrane protein, putative [Shewanella oneidensis MR-1] Length = 127
2891.2 Best-BlastP=> >nrprot 68% Identities = 236/584 (40%), Positives = 358/584 (61 %), Gaps = 4/584 (0%) ref|NP_359923.11 multidrug resistance ABC transporter ATP-binding protein [Rickettsia conorii] pir||F97735 hypothetical protein abcT3 [imported] - Rickettsia conorii (strain Malish 7) gb|AAL02824.11 multidrug resistance ABC transporter ATP-binding protein [Rickettsia conorii] Length = 689
2893.2 Best-BlastP=> >nrprot 51 % Identities = 263/741 (35%), Positives = 402/741 (54%), Gaps = 32/741 (4%) ref|ZP_00080393.11 hypothetical protein [Geobacter metallireducens] Length = 768
2898.4 Best-BlastP=> >nrprot 61 % Identities = 27/72 (37%), Positives = 47/72 (65%) ref|NP_660683.11 acyl-carrier protein [Buchnera aphidicola str. Sg (Schizaphis graminum)] sp|Q8K9J4|ACP_BUCAP Acyl carrier protein (ACP) gb|AAM67894.1 | acyl-carrier protein [Buchnera aphidicola str. Sg (Schizaphis graminum)] Length = 79
29.1 Best-BlastP=> >nrprot 39% Identities = 26/62 (41 %), Positives = 38/62 (61 %) gb|AA043539.11 probable conjugal transfer protein TraD [Rhizobium etii] Length = 71
290.2 Best-BlastP=> >nrprot 58% Identities = 501/1060 (47%), Positives = 679/1050 (64%), Gaps = 57/1060 (5%) ref|NP_900152.1 | conserved hypothetical protein [Chromobacterium violaceum ATCC 12472] gb|AAQ58169.2| conserved hypothetical protein [Chromobacterium violaceum ATCC 12472] Length = 1040
2900.2 Best-BlastP=> >nrprot 60% Identities = 121/308 (39%), Positives = 170/308 (56%), Gaps = 4/308 (1 %) ref|NP_829350.1 | 3-oxoacyl-(acyl-carrier- protein) synthase III [Chlamydophila caviae GPIC] gb|AAP05228.11 3-oxoacyl-(acyl-carrier-protein) synthase III [Chlamydophila caviae GPIC] Length = 336
2905.2 Best-BlastP=> >nrprot No Hits found 2906.2 Best-BlastP=> >nrprot 62% Identities = 95/201 (47%), Positives = 127/201 (63%), Gaps = 1/201 (0%) ref|NP_929829.11 Glutathione S- transferase [Photorhabdus luminescens subsp. laumondii TT01] emb|CAE14968.11 Glutathione S-transferase [Photorhabdus luminescens subsp. laumondii TT01] Length = 201
2908.2 Best-BlastP=> >nrprot No Hits found 2909.1 Best-BlastP=> >nrprot 54% Identities = 137/387 (35%), Positives = 210/387 (54%), Gaps = 25/387 (6%) ref|NP_441897.11 D-alanyl-D-alanine carboxypeptidase [Synechocystis sp. PCC 6803] pir||S76446 hypothetical protein - Synechocystis sp. (strain PCC 6803) dbj|BAA18576.11 D- alanyl-D-alanine carboxypeptidase [Synechocystis sp. PCC 6803] Length = 400
291.4 Best-BlastP=> >nrprot 79% Identities = 283/419 (67%), Positives = 342/419 (81 %) ref|NP_820426.11 NADH dehydrogenase I, F subunit [Coxiella burnetii RSA 493] gb|AAO90940.11 NADH dehydrogenase I, F subunit [Coxiella burnetii RSA 493] Length = 422
2910.1 Best-BlastP=> >nrprot 73% Identities = 272/466 (68%), Positives = 348/465 (74%), Gaps = 1/465 (0%) ref|NP_928701.11 glutamyl-tRNA synthetase, catalytic subunit [Photorhabdus luminescens subsp. laumondii TT01] emb|CAE13694.11 glutamyl-tRNA synthetase, catalytic subunit [Photorhabdus luminescens subsp. laumondii TT01] Length = 472
2911.3 Best-BlastP=> >nrprot 58% Identities = 47/95 (49%), Positives = 62/95 (65%) ref |NP_931076.11 BolA protein [Photorhabdus luminescens subsp. laumondii TT01] emb|CAE16243.1 | BolA protein [Photorhabdus luminescens subsp. laumondii TT01] Length = 104
2914.1 Best-BlastP=> >nrprot 60% Identities = 90/225 (40%), Positives = 135/225 (60%), Gaps = 15/226 (6%) ref|NP_759161.1 | Inactive homolog of metal-dependent proteases [Vibrio vulnificus CMCP6] gb|AAO08678.1 |AE016797_133 Inactive homolog of metal-dependent proteases [Vibrio vulnificus CMCP6] Length = 233
2915.2 Best-BlastP=> >nrprot 8% Identities = 45/199 (22%), Positives = 77/199 (38%), Gaps = 4/199 (2%) ref|NP_497967.1 | cyclin-like F-box (3F797) [Caenorhabditis elegans] pir||T24435 hypothetical protein T04A8.13 - Caenorhabditis elegans emb|CAA84732.2| Hypothetical protein T04A8.13 [Caenorhabditis elegans] Length = 7 i
2916.1 Best-BlastP=> >nrprot No Hits found
2917.1 ,Best-BlastP=> >nrprot 30% Identities = 44/132 (33%), Positives = 71/132 (53%), Gaps = 11/132 (8%) ref|NP_707288.1 | Activator of ProP osmoprotectant transporter [Shigella flexneri 2a str. 301] gb|AAN42995.1 |AE015164_2 Activator of ProP osmoprotectant transporter [Shigella flexneri 2a str. 301] Length = 232
2919.1 Best-BlastP=> >nrprot δδ% Identities = 23/65 (41 %), Positives = 33/56 (60%), Gaps = 4/56 (7%) ref|NP_768003.1 | bsl1363 [Bradyrhizobium japonicum] dbj|BAC46628.11 bsl1363 [Bradyrhizobium japonicum USDA 110] Length = 73
292.1 Best-BlastP=> >nrprot 69% Identities = 75/152 (49%), Positives = 108/152 (71 %), Gaps = 1/152 (0%) dbj|BAA2δ988.11 24-kDa subunit of complex I [Homo sapiens] Length = 231
2920.1 Best-BlastP=> >nrprot No Hits found
2921.3 Best-BlastP=> >nrprot 17% Identities = 81/447 (18%), Positives = 193/447 (43%), Gaps = 56/447 (12%) pir||E71606 hypothetical protein PFB0765w - malaria parasite (Plasmodium falciparum) Length = 980
2924.3 Best-BlastP=> >nrprot No Hits found
2925.1 Best-BlastP=> >nrprot 61 % Identities = 385/477 (80%), Positives = 412/477 (86%), Gaps = 24/477 (5%) gb|AAD60296.1 |AF173009 rep helicase [Legionella pneumophila] Length = 467
2926.1 Best-BlastP=> >nrprot No Hits found
2928.2 Best-BlastP=> >nrprot 63% Identities = 116/298 (38%), Positives = 178/298 (59%), Gaps = 22/298 (7%) ref|ZP_00031876.11 COG1281 : Disulfide bond chaperones of the HSP33 family [Burkholderia fungorum] Length = 316
2929.1 Best-BlastP=> >nrprot 46% Identities = 50/130 (38%), Positives = 77/130 (59%), Gaps = 1/130 (0%) ref|NP_635963.11 conserved hypothetical protein [Xanthomonas campestris pv. campestris str. ATCC 33913] gb|AAM39887.1 | conserved hypothetical protein [Xanthomonas campestris pv. campestris str. ATCC 33913] Length = 171
293.2 Best-BlastP=> >nrprot 89% Identities = 326/417 (78%), Positives = 374/417 (89%) ref|NP_820428.11 NADH dehydrogenase I, D subunit [Coxiella burnetii RSA 493] gb|AAO90942.1 | NADH dehydrogenase I, D subunit [Coxiella burnetii RSA 493] Length = 417
2932.2 Best-BlastP=> >nrprot 60% Identities = 309/814 (37%), Positives = 491/814 (60%), Gaps = 18/814 (2%) ref|NP_890811.11 probable membrane protein [Bordetella bronchiseptica] emb|CAE34640.1 | probable membrane protein [Bordetella bronchiseptica] Length = 1028
2933.2 Best-BlastP=> >nrprot 60% Identities = 122/303 (40%), Positives = 184/303 (60%), Gaps = 14/303 (4%) ref|NP_603145.11 Hypothetical protein [Fusobacterium nucleatum subsp. nucleatum ATCC 25586] gb|AAL94444.11 Hypothetical protein [Fusobacterium nucleatum subsp. nucleatum ATCC 25586] Length = 310
2934.1 Best-BlastP=> >nrprot No Hits found
2938.3 Best-BlastP=> >nrprot 41 % Identities = 250/1286 (19%), Positives = 507/1286 (39%), Gaps = 229/1286 (17%) ref|NP_212646.11 B. burgdorferi predicted coding region BB0512 [Borrelia burgdorferi] pir||G70163 hypothetical protein BB0512 - Lyme disease spirochete gb|AAC66876.1 | B. burgdorferi predicted coding region BB0512 [Borrelia burgdorferi B31] Length = 2166
294.1 Best-BlastP=> >nrprot 67% Identities = 121/219 (56%), Positives = 153/219 (69%), Gaps = 1/219 (0%) ref|NP_820429.1 | NADH dehydrogenase I, C subunit [Coxiella burnetii RSA 493] gb|AAO90943.11 NADH dehydrogenase I, C subunit [Coxiella burnetii RSA 493] Length = 227
2941.1 Best-BlastP=> >nrprot 49% Identities = 173/480 (36%), Positives = 267/480 (55%), Gaps = 7/480 (1 %) ref|NP_819594.11 apolipoprotein N- acyltransferase [Coxiella burnetii RSA 493] gb|AAO90108.11 apolipoprotein N-acyltransferase [Coxiella burnetii RSA 493] Length = 485
2942.1 Best-BlastP=> >nrprot 76% Identities = 502/825 (60%), Positives = 628/825 (76%), Gaps = 8/825 (0%) ref |NP_819590.11 leucyl-tRNA synthetase [Coxiella burnetii RSA 493] gb|AAO90104.11 leucyl-tRNA synthetase [Coxiella burnetii RSA 493] Length = 820
2943.1 Best-BlastP=> >nrprot 42% Identities = 48/160 (30%), Positives = 76/160 (47%), Gaps = 4/160 (2%) ref|NP_252677.11 hypothetical protein [Pseudomonas aeruginosa PA01] ref|ZP_00137430.11 COG2980: Rare lipoprotein B [Pseudomonas aeruginosa UCBPP-PA14] pir||F83148 hypothetical protein PA3988 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG07375.1 |AE004816_11 hypothetical protein PA3988 [Pseudomonas aeruginosa PA01] Length = 207
2949.2 Best-BlastP=> >nrprot 74% Identities = 185/307 (60%), Positives = 238/307 (77%) ref|ZP_00087322.11 hypothetical protein [Pseudomonas fluorescens PfO-1] Length = 325
295.1 Best-BlastP=> >nrprot 88% Identities = 128/152 (84%), Positives = 141/152 (92%) ref|ZP_00023480.1 | COG0377: NADH:ubiquinone oxidoreductase 20 kD subunit and related Fe-S oxidoreductases [Ralstonia metallidurans] Length = 160
2951.2 Best-BlastP=> >nrprot 26% Identities = 100/453 (22%), Positives = 179/453 (39%), Gaps = 86/453 (18%) ref|NP_703829.11 hypothetical protein [Plasmodium falciparum 3D7] emb|CAD50441.11 hypothetical protein [Plasmodium falciparum 3D7] Length = 1321
2952.1 Best-BlastP=> >nrprot 8% Identities = 32/93 (34%), Positives = 53/93 (56%), Gaps = 4/93 (4%) gb|AAL78307.1 |AF288617_4 Dotl [Legionella longbeachae] Length = 212
2953.2 Best-BlastP=> >nrprot 97% Identities = 161/166 (96%), Positives = 163/166 (98%) pir||S49042 global stress protein gspA - Legionella pneumophila Length = 166
2954.3 Best-BlastP=> >nrprot 74% Identities = 198/312 (63%), Positives = 247/312 (79%), Gaps = 5/312 (1 %) ref|NP_903586.11 probable electron- transferring-flavoprotein dehydrogenase [Chromobacterium violaceum ATCC 12472] gb|AAQ61577.1 | probable electron-transferring- flavoprotein dehydrogenase [Chromobacterium violaceum ATCC 12472] Length = 539
2955.2 Best-BlastP=> >nrprot 99% Identities = 126/127 (99%), Positives = 127/127 (100%) gb|AAD51393.1 |AF117715_2 unknown [Legionella pneumophila] Length = 127
2958.2 Best-BlastP=> >nrprot 98% Identities = 248/252 (98%), Positives = 250/262 (99%), Gaps = 1/252 (0%) sp|Q9S4T3|SURE_LEGPN Acid phosphatase surE gb|AAD51394.1 |AF11771 δ_3 survival protein homolog [Legionella pneumophila] Length = 261
2969.1 Best-BlastP=> >nrprot 97% Identities = 242/247 (97%), Positives = 243/247 (98%) gb|AAD61396.1 |AF11771 δ_4 novel lipoprotein homolog [Legionella pneumophila] Length = 247
296.1 Best-BlastP=> >nrprot 78% Identities = 76/118 (64%), Positives = 93/118 (78%) ref|NP_820431.11 NADH dehydrogenase I, A subunit [Coxiella burnetii RSA 493] gb|AAO90945.11 NADH dehydrogenase I, A subunit [Coxiella burnetii RSA 493] Length = 118
2960.1 Best-BlastP=> >nrprot 99% Identities = 337/341 (98%), Positives = 340/341 (99%) gb|AAD51396.1 |AF117715_δ stationary phase specific sigma factor homolog [Legionella pneumophila] Length = 341
2961.1 Best-BlastP=> >nrprot 99% Identities = 410/416 (98%), Positives = 414/416 (99%) sp|Q9S4T0|HGD_LEGPN Homogentisate 1 ,2-dioxygenase (Homogentisicase) (Homogentisate oxygenase) (Homogentisic acid oxidase) Length = 416
2962.1 Best-BlastP=> >nrprot 95% Identities = 235/238 (98%), Positives = 236/238 (99%) gb|AAD51398.1 |AF11771 δ_7 YebC [Legionella pneumophila] Length = 238
2963.2 Best-BlastP=> >nrprot 69% Identities = 110/267 (42%), Positives = 161/267 (58%), Gaps = 10/257 (3%) ref|NP_231867.11 hydroxyacylglutathione hydrolase GloB, putative [Vibrio cholerae 01 biovar eltor str. N16961] pir||A82102 probable hydroxyacylglutathione hydrolase GloB VC2236 [imported] - Vibrio cholerae (strain N16961 serogroup 01 ) gb|AAF96380.1 | hydroxyacylglutathione hydrolase GloB, putative [Vibrio cholerae 01 biovar eltor str. N 16961] Length = 262
2964.1 Best-BlastP=> >nrprot 62% Identities = 121/275 (44%), Positives = 177/275 (64%), Gaps = 3/275 (1 %) ref|NP_229844.11 conserved hypothetical protein [Vibrio cholerae 01 biovar eltor str. N16961 ] pir||B82353 conserved hypothetical protein VC0187 [imported] - Vibrio cholerae (strain N16961 serogroup 01 ) gb|AAF93363.1 | conserved hypothetical protein [Vibrio cholerae 01 biovar eltor str. N16961] Length = 27g 2965.1 Best-BlastP=> >nrprot No Hits found
2966.1 Best-BlastP=> >nrprot 73% Identities = 164/276 (59%), Positives = 209/276 (75%), Gaps = 1/276 (0%) ref|NP_438767.11 methylenetetrahydrofolate dehydrogenase/methenyltetrahydrofolate cyclohydrolase [Haemophilus influenzae Rd] sp|P44313|FOLD_HAEIN FolD bifunctional protein [Includes: Methylenetetrahydrofolate dehydrogenase ; Methenyltetrahydrofolate cyclohydrolase ] pir||A64081 methylenetetrahydrofolate dehydrogenase (NADP) (EC 1.5.1.5) / methenyltetrahydrofolate cyclohydrolase (EC 3.5.4.9) - Haemophilus influenzae (strain Rd KW20) gb|AAC22268.1 | methylenetetrahydrofolate dehydrogenase/methenyltetrahydrofolate cyclohydrolase (folD) [Haemophilus influenzae Rd] Length = 282
2967.1 Best-BlastP=> >nrprot No Hits found
2968.3 Best-BlastP=> >nrprot 13% Identities = 115/119 (96%), Positives = 117/119 (98%) gb|AAM21056.11 FimV [Legionella pneumophila] Length = 119
2969.1 Best-BlastP=> >nrprot 81 % Identities = 50/79 (63%), Positives = 64/79 (81 %) ref|NP_389755.11 similar to transcriptional regulator [Bacillus subtilis] pir||C69931 transcription regulator homolog yozG - Bacillus subtilis emb|CAB13766.11 yozG [Bacillus subtilis subsp. subtilis str. 168] Length = 84
297.2 Best-BlastP=> >nrprot 63% Identities = 69/101 (68%), Positives = 69/101 (68%), Gaps = 8/101 (7%) ref|NP_820432.11 preprotein translocase, SecG subunit [Coxiella burnetii RSA 493] gb|AAO90946.11 preprotein translocase, SecG subunit [Coxiella burnetii RSA 493] Length = 98
2970.2 Best-BlastP=> >nrprot 32% Identities = 34/103 (33%), Positives = 56/103 (54%) ref|ZP_00136103.1 | hypothetical protein [Pseudomonas aeruginosa UCBPP-PA14] Length = 185
2972.3 Best-BlastP=> >nrprot 10% Identities = 43/138 (31 %), Positives = 63/138 (45%), Gaps = 15/138 (10%) ref|NP_704193.1 | ubiquifin carboxyl- terminal hydrolase, putative [Plasmodium falciparum 3D7] emb|CAD51009.11 ubiquitin carboxyl-terminal hydrolase, putative [Plasmodium falciparum 3D7] Length = 3183
2973.2 Best-BlastP=> >nrprot No Hits found 2975.2 Best-BlastP=> >nrprot No Hits found 2976.1 Best-BlastP=> >nrprot No Hits found 2978.1 Best-BlastP=> >nrprot 22% Identities = 41/121 (33%), Positives = 64/121 (52%), Gaps = 11/121 (9%) ref |NP_818058.11 gp8δ [Mycobacteriophage Che9d] gb|AAN08003.11 gp85 [Mycobacteriophage Che9d] Length = 301
2982.1 Best-BlastP=> >nrprot 38% Identities = 174/388 (44%), Positives = 236/388 (60%), Gaps = 17/388 (4%) ref|NP_706610.11 orf, partial conserved hypothetical protein [Shigella flexneri 2a str. 301] ref|NP_836390.11 putative bacteriophage protein [Shigella flexneri 2a str. 2457T] gb|AAN42317.1 |AE015098_7 orf, partial conserved hypothetical protein [Shigella flexneri 2a str. 301] gb|AAP16196.1 | putative bacteriophage protein [Shigella flexneri 2a str. 2457T] Length = 619
2983.1 Best-BlastP=> >nrprot 54% Identities = 33/66 (50%), Positives = 47/66 (71 %) ref|NP_478090.11 hypothetical protein [Corynebacterium glutamicum] emb|CAD12221.1 | hypothetical protein [Corynebacterium glutamicum] Length = 81
2985.1 Best-BlastP=> >nrprot No Hits found
2986.1 Best-BlastP=> >nrprot No Hits found 2987.1 Best-BlastP=> >nrprot No Hits found 2988.1 Best-BlastP=> >nrprot No Hits found 2990.1 Best-BlastP=> >nrprot 44% Identities = 34/97 (35%), Positives = 49/97 (50%), Gaps = 1/97 (1 %) ref|NP_771656.11 bll-5016 [Bradyrhizobium japonicum] dbj|BAC50281.1 | bll5016 [Bradyrhizobium japonicum USDA 110] Length = 198
2991.1 Best-BlastP=> >nrprot 43% Identities = 90/344 (26%), Positives = 145/344 (42%), Gaps = 68/344 (19%) gb|AAC01562.11 S adenosylhomocysteine hydrolase [Thermotoga maritima] Length = 404
2992.2 Best-BlastP=> >nrprot 14% Identities = 38/68 (55%), Positives = 50/68 (73%), Gaps = 1/68 (1 %) gb|EAA26507.11 unknown [Rickettsia sibirica] Length = 73
2993.2 Best-BlastP=> >nrprot 31% Identities = 40/160 (25%), Positives = 76/160 (47%), Gaps = 11/160 (6%) dbj|BAA00448.11 open reading frame (196 AA) [Mus musculus] Length = 196
2994.2 Best-BlastP=> >nrprot 65% Identities = 56/118 (46%), Positives = 82/118 (69%) ref|NP_353891.11 AGR_C_1587p [Agrobacterium tumefaciens] ref|NP_531567.11 conserved hypothetical protein [Agrobacterium tumefaciens str. C58 (U. Washington)] pir||C97465 hypothetical protein AGR_C_1δ87 [imported] - Agrobacterium tumefaciens (strain C58, Cereon) pir||AE2683 conserved hypothetical protein Atu0869 [imported] - Agrobacterium tumefaciens (strain C58, Dupont) gb|AAK86676.11 AGR_C_1587p [Agrobacterium tumefaciens str. C68 (Cereon)] gb|AAL41883.1 | conserved hypothetical protein [Agrobacterium tumefaciens str. C68 (U. Washington)] Length = 122
2995.1 Best-BlastP=> >nrprot 32% Identities = 68/290 (23%), Positives = 135/290 (46%), Gaps = 39/290 (13%) ref|NP_473345.2| hypothefical protein [Plasmodium falciparum 3D7] emb|CAB39052.2| hypothetical protein [Plasmodium falciparum 3D7] Length = 670
2996.1 Best-BlastP=> >nrprot No Hits found
2998.1 Best-BlastP=> >nrprot 61 % Identities = 44/134 (32%), Positives = 73/134 (64%), Gaps = 1/134 (0%) ref|NP_82092δ.11 hypothetical protein [Coxiella burnetii RSA 493] gb|AA091439.11 hypothetical protein [Coxiella burnetii RSA 493] Length = 148
2999.3 Best-BlastP=> >nrprot 24% Identities = 39/156 (25%), Positives = 69/156 (44%), Gaps = 33/156 (21 %) ref|ZP_00047224.11 COG3774: Mannosyltransferase OCH1 and related enzymes [Lactobacillus gasseri] Length = 233
3.1 Best-BlastP=> >nrprot 49% Identities = 36/79 (45%), Positives = 50/79 (63%) ref|NP_907749.11 hypothetical protein WS1616 [Wolinella succinogenes] emb|CAE10649.11 hypothetical protein [Wolinella succinogenes] Length = 96
30.1 Best-BlastP=> >nrprot No Hits found 300.2 Best-BlastP=> >nrprot 75% Identities = 346/570 (60%), Positives = 431/570 (75%), Gaps = 3/570 (0%) ref|NP_752183.11 Prolyl-tRNA synthetase [Escherichia coli CFT073] gb|AAN78727.11 AE01675δ_227 Prolyl-tRNA synthetase [Escherichia coli CFT073] Length = 590
3000.1 Best-BlastP=> >nrprot 58% Identities = 312/711 (43%), Positives = 440/711 (61 %), Gaps = 35/711 (4%) ref|ZP_00090403.11 COG3243: Poly(3- hydroxyalkanoate) synthetase [Azotobacter vinelandii] Length = 824
3001.1 Best-BlastP=> >nrprot No Hits found 3002.1 Best-BlastP=> >nrprot 36% Identities = 96/343 (27%), Positives = 152/343 (44%), Gaps = 29/343 (8%) ref|ZP_00019713.11 hypothetical protein [Chloroflexus aurantiacus] Length = 360
3003.1 Best-BlastP=> >nrprot 57% Identities = 77/202 (38%), Posifives = 119/202 (58%), Gaps = 3/202 (1 %) ref|ZP_00008122.1 | COGOδOO: SAM- dependent methyltransferases [Rhodobacter sphaeroides] Length = 204
3006.2 Best-BlastP=> >nrprot 69% Identities = 74/164 (45%), Positives = 116/164 (70%), Gaps = 2/164 (1 %) ref|NP_924145.11 hypothetical protein gll1199 [Gloeobacter violaceus] dbj|BAC89140.1 | gill 199 [Gloeobacter violaceus] Length = 175
3008.1 Best-BlastP=> >nrprot 56% Identities = 65/195 (33%), Positives = 114/195 (58%), Gaps = 1/195 (0%) ref|NP_820992.11 conserved hypothetical protein [Coxiella burnetii RSA 493] gb|AA091506.11 conserved hypothetical protein [Coxiella burnetii RSA 493] Length = 203
3009.2 Best-BlastP=> >nrprot No Hits found
3011.2 Best-BlastP=> >nrprot 49% Identities = 88/315 (27%), Positives = 164/315 (62%), Gaps = 27/31 δ (8%) ref|NP_819925.11 hypothetical protein [Coxiella burnetii RSA 493] gb|AAO90439.11 hypothetical protein [Coxiella burnetii RSA 493] Length = 324
3013.2 Best-BlastP=> >nrprot 48% Identities = 48/152 (31%), Positives = 76/152 (49%), Gaps = 5/152 (3%) ref|NP_720010.1 | acetyltransferase, GNAT family [Shewanella oneidensis MR-1] gb|AAN57454.1 |AE01 δ881_1 acetyltransferase, GNAT family [Shewanella oneidensis MR-1] Length = 164
3014.1 Best-BlastP=> >nrprot 48% Identities = 67/192 (29%), Positives = 98/192 (51 %), Gaps = 22/192 (11%) ref|NP_656418.1| APSJcinase, Adenylylsulfate kinase [Bacillus anthracis A2012] ref|NP_844924.11 hypothetical protein [Bacillus anthracis str. Ames] gb|AAP26410.11 hypothetical protein BA2566 [Bacillus anthracis str. Ames] Length = 186
301 δ.1 Best-BlastP=> >nrprot 60% Identities = 83/260 (33%), Positives = 124/260 (49%), Gaps = 7/260 (2%) ref|NP_656739.11 Ubie_methyltran, ubiE/COQδ methyltransferase family [Bacillus anthracis A2012] ref|NP_84δ200.11 conserved hypothetical protein [Bacillus anthracis str. Ames] gb|AAP26686.1 | conserved hypothetical protein [Bacillus anthracis str. Ames] Length = 254
3016.1 Best-BlastP=> >nrprot No Hits found
3017.1 Best-BlastP=> >nrprot No Hits found 3018.1 Best-BlastP=> >nrprot No Hits found 3022.1 Best-BlastP=> >nrprot No Hits found 3023.1 Best-BlastP=> >nrprot 68% Identities = 26/56 (46%), Positives = 41/56 (73%) ref|NP_832415.11 Glutamate-rich protein grpB [Bacillus cereus ATCC 14579] gb|AAP09616.11 Glutamate-rich protein grpB [Bacillus cereus ATCC 14579] Length = 168
3024.1 Best-BlastP=> >nrprot 76% Identities = 253/425 (59%), Positives = 326/425 (76%), Gaps = 2/425 (0%) ref|ZP_00091804.11 COG0172: Seryl- tRNA synthetase [Azotobacter vinelandii] Length = 607
3027.1 Best-BlastP=> >nrprot 68% Identities = 86/149 (57%), Positives = 111/149 (74%) gb|AAM89273.1 |AF528189_2 SecB [Serratia marcescens] Length = 156
3028.1 Best-BlastP=> >nrprot 75% Identities = 48/83 (57%), Positives = 64/83 (77%) ref|ZP_00089760.11 COG0695: Glutaredoxin and related proteins [Azotobacter vinelandii] Length = 84
3030.2 Best-BlastP=> >nrprot 63% Identities = 70/163 (42%), Positives = 104/163 (63%), Gaps = 2/163 (1 %) ref|ZP_00021668.11 COG2606: Uncharacterized conserved protein [Ralstonia metallidurans] Length = 165
3031.2 Best-BlastP=> >nrprot 42% Identities = 37/136 (27%), Positives = 60/136 (44%), Gaps = 2/136 (1 %) ref|NP_347707.11 Hypothetical protein [Clostridium acetobutylicum] pir||D97032 hypothetical protein CAC1073 [imported] - Clostridium acetobutylicum gb|AAK79047.1 |AE007622_9 Hypothetical protein [Clostridium acetobutylicum] Length = 152
3036.2 Best-BlastP=> >nrprot 62% Identities = 89/185 (48%), Positives = 120/185 (64%), Gaps = 9/185 (4%) ref|NP_753769.11 Hypothetical protein ydcN [Escherichia coli CFT073] gb|AAN80321.1 |AE016760_180 Hypothetical protein ydcN [Escherichia coli CFT073] Length = 178
3036.2 Best-BlastP=> >nrprot 46% Identities = 54/146 (36%), Positives = 85/146 (58%), Gaps = 5/146 (3%) ref|NP_34g095.11 Predicted kinase from adenilate kinase family, FLAR-like protein [Clostridium acetobutylicum] pir||H97205 probable kinase from adenilate kinase family, FLAR-like protein [imported] - Clostridium acetobutylicum gb|AAK8043δ.1 |AE007747_4 Predicted kinase from adenilate kinase family, FLAR-like protein [Clostridium acetobutylicum] Length = 177
304.2 Best-BlastP=> >nrprot 21 % Identities = 122/500 (24%), Positives = 202/500 (40%), Gaps = 87/600 (17%) gb|AAH16985.2| Unknown (protein for MGC:21968) [Homo sapiens] Length = 579
3041.1 Best-BlastP=> >nrprot 16% Identities = 33/146 (22%), Positives = 67/146 (45%), Gaps = 20/146 (13%) gb|AAQ5δ479.1 | hypothetical protein [Methanococcus voltae] Length = 178
3042.1 Best-BlastP=> >nrprot No Hits found
3045.3 Best-BlastP=> >nrprot 58% Identities = 162/484 (33%), Positives = 261/484 (53%), Gaps = 46/484 (9%) gb|AAP68896.11 putative N5'- nucleotidase [Oryza sativa (japonica cultivar-group)] Length = 569
3046.3 Best-BlastP=> >nrprot 26% Identities = 39/103 (37%), Positives = 50/103 (48%), Gaps = 10/103 (9%) pir||F72654 hypothetical protein APE0666 - Aeropyrum pernix (strain K1 ) dbj|BAA79638.1 | 102aa long hypothetical protein [Aeropyrum pernix] Length = 102
3047.2 Best-BlastP=> >nrprot 73% Identities = 131/210 (62%), Positives = 159/210 (75%) ref|NP_716775.11 ribose 5-phosphate isomerase [Shewanella oneidensis MR-1] sp|Q8EHR7|RPIA_SHEON Ribose 5-phosphate isomerase A (Phosphoriboisomerase A) (PRI) gb|AAN54220.1 |AE016569_4 ribose 5-phosphate isomerase [Shewanella oneidensis MR-1] Length = 219
3049.1 Best-BlastP=> >nrprot 69% Identities = 133/239 (55%), Positives = 174/239 (72%), Gaps = 1/239 (0%) ref|NP_746983.11 RNA methyltransferase, TrmH family, group 3 [Pseudomonas putida KT2440] gb|AAN70447.1 |AE016686_1 RNA methyltransferase, TrmH family, group 3 [Pseudomonas putida KT2440] Length = 248
305.2 Best-BlastP=> >nrprot 69% Identities = 124/236 (52%), Positives = 167/236 (70%) ref|NP_820118.11 dienelactone hydrolase family protein [Coxiella burnetii RSA 493] gb|AAO90632.11 dienelactone hydrolase family protein [Coxiella burnetii RSA 493] Length = 237
3053.3 Best-BlastP=> >nrprot 54% Identifies = 40/122 (32%), Positives = 80/122 (65%), Gaps = 4/122 (3%) gb|AAA29909.11 ORF 3 Length = 393 3064.3 Best-BlastP=> >nrprot 59% Identities = 117/304 (38%), Positives = 170/304 (56%), Gaps = 27/304 (8%) ref|ZP_00067579.11 COG2304: Uncharacterized protein containing a von Willebrand factor type A (vWA) domain [Microbulbifer degradans 2-40] Length = 658
3056.1 Best-BlastP=> >nrprot 52% Identities = 114/330 (34%), Positives = 164/330 (49%), Gaps = 35/330 (10%) ref|NP_718657.11 TPR domain protein [Shewanella oneidensis MR-1] gb|AAN56101.1 |AE015746_δ TPR domain protein [Shewanella oneidensis MR-1] Length = 679
3057.1 Best-BlastP=> >nrprot 61% Identities = 150/319 (47%), Positives = 213/319 (66%), Gaps = 9/319 (2%) ref|ZP_00087368.1 | COG2304: Uncharacterized protein containing a von Willebrand factor type A (vWA) domain [Pseudomonas fluorescens PfO-1] Length = 359
3058.1 Best-BlastP=> >nrprot No Hits found
3059.2 Best-BlastP=> >nrprot 99% Identities = 258/259 (99%), Positives = 259/259 (100%) emb|CAA06664.11 29 kDa immunogenic protein [Legionella pneumophila] Length = 259
306.1 Best-BlastP=> >nrprot 75% Identities = 290/467 (62%), Positives = 356/467 (76%), Gaps = 1/467 (0%) ref|NP_254123.11 probable biotin carboxylase subunit of a transcarboxylase [Pseudomonas aeruginosa PA01] pir||G82966 probable biotin carboxylase subunit of a transcarboxylase PA5436 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG08821.1 |AE004956_5 probable biotin carboxylase subunit of a transcarboxylase [Pseudomonas aeruginosa PA01] Length = 471
3060.1 Best-BlastP=> >nrprot No Hits found
3061.2 Best-BlastP=> >nrprot 72% Identities = 119/204 (58%), Positives = 156/204 (76%), Gaps = 2/204 (0%) ref|ZP_00084510.11 COG2011 : ABC-type metal ion transport system, permease component [Pseudomonas fluorescens PfO-1] Length = 224
3063.2 Best-BlastP=> >nrprot 69% Identities = 174/343 (50%), Positives = 238/343 (69%), Gaps = 7/343 (2%) ref|NP_87357δ.11 D-methionine transport ATP-binding protein MetN [Haemophilus ducreyi 35000HP] gb|AAP95964.1 | D-methionine transport ATP-binding protein MetN [Haemophilus ducreyi 35000HP] Length = 344
3066.1 Best-BlastP=> >nrprot 40% Identities = 39/138 (28%), Positives = 63/138 (45%), Gaps = 15/138 (10%) ref|NP_520406.1 | PUTATIVE SIGNAL PEPTIDE PROTEIN [Ralstonia solanacearum] emb|CAD15992.11 PUTATIVE SIGNAL PEPTIDE PROTEIN [Ralstonia solanacearum] Length = 143
3067.1 Best-BlastP=> >nrprot 54% Identities = 27/100 (27%), Positives = 59/100 (59%), Gaps = 6/100 (6%) ref|NP_820531.11 hypothetical protein [Coxiella burnetii RSA 493] gb|AA091045.11 hypothetical protein [Coxiella burnetii RSA 493] Length = 124
3068.1 Best-BlastP=> >nrprot 84% Identities = 290/388 (74%), Positives = 331/388 (85%) ref|NP_800128.11 putative acyl-CoA dehydrogenase [Vibrio parahaemolyticus RIMD 2210633] dbj|BAC61961.11 putative acyl-CoA dehydrogenase [Vibrio parahaemolyticus] Length = 389
3069.1 Best-BlastP=> >nrprot 55% Identities = 83/222 (37%), Positives = 126/222 (56%), Gaps = 4/222 (1 %) ref|NP_819980.11 conserved hypothetical protein [Coxiella burnetii RSA 493] gb|AAO90494.11 conserved hypothetical protein [Coxiella burnetii RSA 493] Length = 230
307.2 Best-BlastP=> >nrprot 47% Identities = 54/168 (32%), Positives = 84/168 (50%), Gaps = 8/168 (4%) ref |NP_796502.11 hypothetical protein VP0123 [Vibrio parahaemolyticus RIMD 2210633] dbj|BAC58386.11 hypothetical protein [Vibrio parahaemolyticus] Length = 180
3070.2 Best-BlastP=> >nrprot No Hits found 3072.2 Best-BlastP=> >nrprot 67% Identities = 300/598 (50%), Positives = 400/598 (66%), Gaps = 9/598 (1 %) ref|NP_637990.11 ATP-dependent RNA helicase [Xanthomonas campestris pv. campestris str. ATCC 33913] gb|AAM41914.11 ATP-dependent RNA helicase [Xanthomonas campestris pv. campestris str. ATCC 33913] Length = 642
3074.1 Best-BlastP=> >nrprot No Hits found
3075.2 Best-BlastP=> >nrprot No Hits found
3077.1 Best-BlastP=> >nrprot 42% Identities = 66/214 (30%), Positives = 103/214 (48%), Gaps = 18/214 (8%) ref |NP_35δ 106.11 AGR_C_3887p [Agrobacterium tumefaciens] ref|NP_532818.1 | conserved hypothetical protein [Agrobacterium tumefaciens str. C68 (U. Washington)] pir||AH2839 conserved hypothetical protein Atu2144 [imported] - Agrobacterium tumefaciens (strain C58, Dupont) pir||B97617 similar to orf3 gene in methylobacterium extorquenS [imported] - Agrobacterium tumefaciens (strain C58, Cereon) gb|AAK87891.11 AGR_C_3887p [Agrobacterium tumefaciens str. Cδ8 (Cereon)] gb|AAL43134.11 conserved hypothetical protein [Agrobacterium tumefaciens str. C58 (U. Washington)] Length = 267
3078.2 Best-BlastP=> >nrprot 53% Identities = 128/399 (32%), Positives = 219/399 (54%), Gaps = 38/399 (9%) ref|NP_071132.11 hypothetical protein [Archaeoglobus fulgidus DSM 4304] sp|O27g77|YN07_ARCFU Hypothetical protein AF2307 pir||C69538 hypothetical protein AF2307 - Archaeoglobus fulgidus gb|AAB88967.11 A. fulgidus predicted coding region AF2307 [Archaeoglobus fulgidus DSM 4304] Length = 365
3080.2 Best-BlastP=> >nrprot 64% Identities = 210/4gθ (42%), Positives = 312/490 (63%), Gaps = 12/490 (2%) ref|ZP_00139612.11 COG0189: Glutathione synthase/Ribosomal protein S6 modification enzyme (glutaminyl transferase) [Pseudomonas aeruginosa UCBPP-PA14] Length = 495
3081.2 Best-BlastP=> >nrprot No Hits found
3082.1 Best-BlastP=> >nrprot 67% Identities = 75/167 (47%), Positives = 106/157 (67%) ref|NP_231147.11 hypothetical protein VC1506 [Vibrio cholerae 01 biovar eltor str. N16961] pir||B82191 hypothetical protein VC1506 [imported] - Vibrio cholerae (strain N16961 serogroup 01 ) gb|AAF94661.11 hypothetical protein [Vibrio cholerae 01 biovar eltor str. N16961] Length = 158
3083.1 Best-BlastP=> >nrprot 48% Identities = 52/171 (30%), Positives = 90/171 (52%), Gaps = 1/171 (0%) ref|NP_404077.11 osmotically inducible protein Y [Yersinia pestis] ref|NP_671042.1 | hyperosmotically inducible periplasmic protein [Yersinia pestis KIM] pir||AF0053 osmotically inducible protein Y [imported] - Yersinia pestis (strain C092) emb|CAC89289.11 osmotically inducible protein Y [Yersinia pestis C092] gb|AAM87293.1 |AE013978_5 hyperosmotically inducible periplasmic protein [Yersinia pestis KIM] Length = 204
3085.2 Best-BlastP=> >nrprot 84% Identities = 693/944 (73%), Positives = 805/944 (85%), Gaps = 4/944 (0%) ref|NP_819318.11 excinuclease ABC, A subunit [Coxiella burnetii RSA 493] gb|AA089832.11 excinuclease ABC, A subunit [Coxiella burnetii RSA 493] Length = 954
3086.1 Best-BlastP=> >nrprot 78% Identities = 133/191 (69%), Positives = 152/191 (79%) ref|NP_819577.11 lemA protein [Coxiella burnetii RSA 493] gb|AAO90091.11 lemA protein [Coxiella burnetii RSA 493] Length = 192
3087.1 Best-BlastP=> >nrprot 77% Identities = 198/344 (57%), Positives = 265/344 (77%), Gaps = 6/344 (1 %) ref|NP_819578.11 heat shock protein HtpX [Coxiella burnetii RSA 493] gb|AAO90092.11 heat shock protein HtpX [Coxiella burnetii RSA 493] Length = 348
3088.1 Best-BlastP=> >nrprot 71% Identities = 141/251 (56%), Positives = 185/251 (73%) ref|ZP_001282gi .11 COG0842: ABC-type multidrug transport system, permease component [Pseudomonas syringae pv. syringae B728a] Length = 262
309.2 Best-BlastP=> >nrprot 67% Identities = 545/1113 (48%), Positives = 752/1113 (67%), Gaps = 2/1113 (0%) ref|NP_820221.11 conserved hypothetical protein [Coxiella burnetii RSA 493] gb|AAO90735.11 conserved hypothetical protein [Coxiella burnetii RSA 493] Length = 1619
3091.1 Best-BlastP=> >nrprot No Hits found
3093.1 Best-BlastP=> >nrprot 73% Idenfities = 139/243 (57%), Positives = 185/243 (76%), Gaps = 2/243 (0%) ref|NP_928570.11 tRNA (guanine-N1 -)- methyltransferase (M1 G-methyltransferase) [Photorhabdus luminescens subsp. laumondii TT01] emb|CAE13553.1 | tRNA (guanine-N1-)- methyltransferase (M1G-methyltransferase) [Photorhabdus luminescens subsp. laumondii TT01] Length = 250
3095.1 Best-BlastP=> >nrprot 61% Identities = 65/169 (38%), Positives = 105/169 (62%), Gaps = 7/169 (4%) ref |NP_716978.11 16S rRNA processing protein RimM [Shewanella oneidensis MR-1] gb|AAN54423.1 |AE015579_12 16S rRNA processing protein RimM [Shewanella oneidensis MR-1] Length = 177
3096.1 Best-BlastP=> >nrprot 74% Identities = 51/85 (60%), Posifives = 65/8δ (76%) ref|NP_841705.11 Ribosomal protein S16 [Nitrosomonas europaea ATCC 19718] emb|CAD85582.11 Ribosomal protein S16 [Nitrosomonas europaea ATCC 19718] Length = 89
3097.3 Best-BlastP=> >nrprot 79% Identities = 299/447 (66%), Positives = 365/447 (81 %), Gaps = 1/447 (0%) ref|NP_716976.11 signal recognition particle protein Ffh [Shewanella oneidensis MR-1] gb|AAN54421.1 |AE015579_10 signal recognition particle protein Ffh [Shewanella oneidensis MR-1] Length = 457
310.1 Best-BlastP=> >nrprot No Hits found
3101.1 Best-BlastP=> >nrprot 12% Identities = 46/178 (25%), Positives = 78/178 (43%), Gaps = 12/178 (6%) gb|AAQ73211.11 M protein [Streptococcus pyogenes] Length = 243
3102.2 Best-BlastP=> >nrprot 50% Identities = 61/141 (43%), Positives = 80/141 (56%), Gaps = 3/141 (2%) ref|ZP_00084504.11 COG0735: Fe2+/Zn2+ uptake regulation proteins [Pseudomonas fluorescens PfO-1] Length = 160
3104.2 Best-BlastP=> >nrprot 38% Identities = 208/652 (31%), Positives = 329/652 (50%), Gaps = 68/652 (10%) ref|NP_717300.11 cation transport ATPase, E1-E2 family [Shewanella oneidensis MR-1] gb|AAN54744.1 |AE015614_11 cation transport ATPase, E1-E2 family [Shewanella oneidensis MR-1] Length = 753
3105.2 Best-BlastP=> >nrprot 41 % Identities = 26/69 (37%), Positives = 42/69 (60%) ref|ZP_00117353.11 COG0642: Signal transduction histidine kinase [Cytophaga hutchinsonii] Length = 856
3106.2 Best-BlastP=> >nrprot 36% Identities = 149/353 (42%), Positives = 220/353 (62%), Gaps = 3/353 (0%) ref|NP_442598.11 PleD gene product homologue [Synechocystis sp. PCC 6803] pir||S76977 pleD-4 protein - Synechocystis sp. (strain PCC 6803) dbj|BAA10669.1 | slr0302 [Synechocystis sp. PCC 6803] Length = 768
3107.2 Best-BlastP=> >nrprot No Hits found 310g.2 Best-BlastP=> >nrprot 55% Identities = 39/100 (39%), Positives = 63/100 (63%), Gaps = 1/100 (1 %) ref|NP_455037.11 cytochrome o ubiquinol oxidase C subunit [Salmonella enterica subsp. enterica serovar Typhi] ref|NP_459436.11 cytochrome o ubiquinol oxidase subunit IV [Salmonella typhimurium LT2] ref|NP_806150.11 cytochrome o ubiquinol oxidase C subunit [Salmonella enterica subsp. enterica serovar Typhi Ty2] pir||AC0567 cytochrome o ubiquinol oxidase C chain [imported] - Salmonella enterica subsp. enterica serovar Typhi (strain CT18) gb|AAL1939δ.1 | cytochrome o ubiquinol oxidase subunit IV [Salmonella typhimurium LT2] emb|CAD08899.1 | cytochrome o ubiquinol oxidase C subunit [Salmonella enterica subsp. enterica serovar Typhi] gb|AAO70010.11 cytochrome o ubiquinol oxidase C subunit [Salmonella enterica subsp. enterica serovar Typhi Ty2] Length = 109
3110.1 Best-BlastP=> >nrprot 76% Identities = 128/188 (68%), Positives = 153/188 (81%) ref|NP_820040.1 | cytochrome o ubiquinol oxidase, subunit III [Coxiella burnetii RSA 493] gb|AAO905δ4.1 | cytochrome o ubiquinol oxidase, subunit III [Coxiella burnetii RSA 493] Length = 198
3113.1 Best-BlastP=> >nrprot 86% Identities = 482/665 (72%), Positives = 568/665 (85%), Gaps = 3/665 (0%) ref|NP_820041.11 cytochrome o ubiquinol oxidase, subunit I [Coxiella burnetii RSA 493] gb|AAO9055δ.11 cytochrome o ubiquinol oxidase, subunit I [Coxiella burnetii RSA 493] Length = 668
3114.2 Best-BlastP=> >nrprot 69% Identities = 173/292 (59%), Positives = 222/292 (76%), Gaps = 10/292 (3%) ref|NP_820042.1 | cytochrome o ubiquinol oxidase, subunit II [Coxiella burnetii RSA 493] gb|AAO90556.11 cytochrome o ubiquinol oxidase, subunit II [Coxiella burnetii RSA 493] Length = 298
3116.2 Best-BlastP=> >nrprot 80% Identities = 82/130 (63%), Positives = 106/130 (81 %) ref|NP_820044.11 hypothetical protein [Coxiella burnetii RSA 493] gb|AAO90558.11 hypothetical protein [Coxiella burnetii RSA 493] Length = 133
3117.2 Best-BlastP=> >nrprot 68% Identities = 116/211 (54%), Positives = 152/211 (72%), Gaps = 2/211 (0%) ref|ZP_0008566δ.11 COG0323: DNA mismatch repair enzyme (predicted ATPase) [Pseudomonas fluorescens PfO-1] Length = 262
3118.1 Best-BlastP=> >nrprot No Hits found
3119.1 Best-BlastP=> >nrprot 61% Identities = 111/264 (42%), Positives = 162/264 (61 %), Gaps = 7/264 (2%) ref|NP_794780.1 | pyrroline-5-carboxylate reductase [Pseudomonas syringae pv. tomato str. DC3000] gb|AAOδ8475.11 pyrroline-δ-carboxylate reductase [Pseudomonas syringae pv. tomato str. DC3000] Length = 272
312.3 Best-BlastP=> >nrprot No Hits found
3121.2 Best-BlastP=> >nrprot 67% Identities = 119/225 (52%), Positives = 156/225 (69%) ref|ZP_00089980.11 COG0325: Predicted enzyme with a TIM- barrel fold [Azotobacter vinelandii] Length = 234
3124.2 Best-BlastP=> >nrprot 88% Identities = 258/345 (74%), Positives = 305/345 (88%) ref|NP_638103.11 twitching motility protein [Xanthomonas campestris pv. campestris str. ATCC 33913] ref|NP_643233.11 twitching motility protein [Xanthomonas axonopodis pv. citri str. 306] gb|AAM37769.11 twitching motility protein [Xanthomonas axonopodis pv. citri str. 306] gb|AAM42027.11 twitching motility protein [Xanthomonas campestris pv. campestris str. ATCC 33913] Length = 345
3127.1 Best-BlastP=> >nrprot 99% Identities = 235/235 (100%), Positives = 235/235 (100%) sp|Q9Xδ28|RNPH_LEGPN Ribonuclease PH (RNase PH) (tRNA nucleotidyltransferase) gb|AAD28218.1 |AF120720_1 ribonuclease PH [Legionella pneumophila] Length = 236
3128.1 Best-BlastP=> >nrprot 67% Identities = 62/172 (36%), Positives = 95/172 (56%), Gaps = 13/172 (7%) ref|NP_747016.11 conserved hypothetical protein [Pseudomonas putida KT2440] gb|AAN70480.1 |AE016689_8 conserved hypothetical protein [Pseudomonas putida KT2440] Length = 172
3129.2 Best-BlastP=> >nrprot 73% Identities = 195/336 (58%), Positives = 247/336 (73%) ref|NP_252613.11 S-adenosylmethionine:trna ribosyltransferase-isomerase [Pseudomonas aeruginosa PA01] sp|Q9HXH8|QUEA_PSEAE S-adenosylmethionine:tRNA ribosyltransferase isomerase (Queuosine biosynthesis protein queA) pir||A83170 S-adenosylmethionine-tRNA ribosyltransferase-isomerase (EC 5.4.99.-) queA PA3824 [similarity] - Pseudomonas aeruginosa (strain PA01) gb|AAG07211.1 |AE004799_17 S-adenosylmethionine:trna ribosyltransferase-isomerase [Pseudomonas aeruginosa PA01] Length = 347
3130.1 Best-BlastP=> >nrprot 18% Identities = 31/100 (31 %), Positives = 52/100 (52%), Gaps = 3/100 (3%) ref|NP_562861.1 | queuine tRNA- ribosyltransferase [Clostridium perfringens] sp|Q8XJ16|TGT_CLOPE Queuine tRNA-ribosyltransferase (tRNA-guanine transglycosylase) (Guanine insertion enzyme) dbj|BAB81651.1 | queuine tRNA-ribosyltransferase [Clostridium perfringens str. 13] Length = 380
3131.1 Best-BlastP=> >nrprot 68% Identities = 53/111 (47%), Positives = 77/111 (69%) ref|ZP_00087740.11 COG1862: Preprotein translocase subunit YajC [Pseudomonas fluorescens PfO-1] Length = 111
3132.1 Best-BlastP=> >nrprot No Hits found
3133.2 Best-BlastP=> >nrprot No Hits found
3134.1 Best-BlastP=> >nrprot 57% Identities = 65/150 (43%), Positives = 91/150 (60%), Gaps = 6/150 (4%) ref|NP_419685.1 | hypothetical protein [Caulobacter crescentus CB15] pir||A87357 hypothetical protein CC0868 [imported] - Caulobacter crescentus gb|AAK22853.1 | hypothetical protein [Caulobacter crescentus CB15] Length = 196
3135.1 Best-BlastP=> >nrprot 69% Identities = 43/63 (68%), Positives = 49/63 (77%) ref|ZP_00067639.11 COG4728: Uncharacterized protein conserved in bacteria [Microbulbifer degradans 2-40] Length = 93
3136.1 Best-BlastP=> >nrprot 54% Idenfities = 38/105 (36%), Positives = 60/105 (57%), Gaps = 2/105 (1 %) gb|AAN62313.11 AF440524 00 conserved hypothetical protein [Pseudomonas aeruginosa] Length = 130
3137.1 Best-BlastP=> >nrprot No Hits found
3138.2 Best-BlastP=> >nrprot No Hits found 3142.2 Best-BlastP=> >nrprot 53% Identities = 106/281 (37%), Positives = 158/281 (56%), Gaps = 19/281 (6%) ref|ZP_00065061.11 COG3961 : Rod binding protein [Microbulbifer degradans 2-40] Length = 318
3146.1 Best-BlastP=> >nrprot No Hits found
3146.2 Best-BlastP=> >nrprot 48% Identities = 74/241 (30%), Positives = 122/241 (50%), Gaps = 7/241 (2%) ref|NP_928923.11 hypothetical protein [Photorhabdus luminescens subsp. laumondii TT01] emb|CAE13928.11 unnamed protein product [Photorhabdus luminescens subsp. laumondii TT01] Length = 279
3147.1 Best-BlastP=> >nrprot 78% Identities = 107/158 (67%), Positives = 127/158 (80%) ref|NP_439483.11 transcription elongation factor [Haemophilus influenzae Rd] sp|P43881 |GREA_HAEIN Transcription elongation factor greA (Transcript cleavage factor greA) pir||B64117 transcription elongation factor greA - Haemophilus influenzae (strain Rd KW20) gb|AAC22976.11 transcription elongation factor (greA) [Haemophilus influenzae Rd] Length = 158
3148.1 Best-BlastP=> >nrprot 56% Identities = 101/297 (34%), Positives = 150/297 (50%), Gaps = 36/297 (12%) ref|NP_842462.11 conserved hypothetical protein [Nitrosomonas europaea ATCC 19718] emb|CAD86383.11 conserved hypothetical protein [Nitrosomonas europaea ATCC 19718] Length = 304
314g.1 Best-BlastP=> >nrprot 36% Identities = 69/341 (20%), Positives = 132/341 (38%), Gaps = 53/341 (1 δ%) ref|NP_616447.11 hypothetical protein (multi-domain) [Methanosarcina acetivorans str. C2A] gb|AAM04927.11 hypothetical protein (multi-domain) [Methanosarcina acetivorans str. C2A] Length = 684
316.3 Best-BlastP=> >nrprot 37% Identities = 97/212 (46%), Positives = 127/212 (59%) gb|AAO50865.1 | similar to Leishmania major. Ppg3 ■ [Dictyostelium discoideum] Length = 374
3160.1 Best-BlastP=> >nrprot 60% Identities = 94/212 (44%), Positives = 138/212 (65%), Gaps = 3/212 (1 %) ref|NP_819377.11 acid phosphatase, class B [Coxiella burnetii RSA 493] gb|AA089891.11 acid phosphatase, class B [Coxiella burnetii RSA 493] Length = 221
3152.1 Best-BlastP=> >nrprot 98% Identities = 174/177 (98%), Positives = 176/177 (99%) gb|AAM00398.1 |AF386079_8 CcmG [Legionella pneumophila] Length = 177
3153.3 Best-BlastP=> >nrprot 99% Identities = 643/650 (98%), Positives = 649/650 (99%) gb|AAM00397.1 |AF386079_7 CcmF [Legionella pneumophila] Length = 650
3157.1 Best-BlastP=> >nrprot 98% Identities = 142/143 (99%), Positives = 142/143 (99%) gb|AAM00396.1 |AF386079_6 CcmE [Legionella pneumophila] Length = 143
3158.1 Best-BlastP=> >nrprot 97% Identities = 72/73 (98%), Positives = 72/73 (98%) gb|AAM00395.1 |AF386079_5 CcmD [Legionella pneumophila] Length = 73
3169.2 Best-BlastP=> >nrprot 98% Identities = 247/251 (98%), Positives = 249/251 (99%) gb|AAM00394.1 |AF386079_4 CcmC [Legionella pneumophila] Length = 251
3160.1 Best-BlastP=> >nrprot No Hits found
3161.1 Best-BlastP=> >nrprot 44% Identifies = 60/224 (26%), Posifives = 103/224 (45%), Gaps = 13/224 (5%) gb|AAK18828.1 |AF327739_5 Peb1 [Streptococcus thermophilus] Length = 277
3163.1 Best-BlastP=> >nrprot 69% Identities = 49/91 (53%), Positives = 68/91 (74%) gb|AAP83334.1 |AF469614_2 unknown [Francisella tularensis subsp. tularensis] Length = 94
3165.1 Best-BlastP=> >nrprot No Hits found
3166.1 Best-BlastP=> >nrprot 65% Identities = 99/176 (56%), Positives = 127/176 (72%), Gaps = 2/176 (1 %) ref|NP_903124.1 | probable DNA-directed DNA polymerase, bacteriophage-type [Chromobacterium violaceum ATCC 12472] gb|AAQ61115.1 | probable DNA-directed DNA polymerase, bacteriophage-type [Chromobacterium violaceum ATCC 12472] Length = 280
3167.1 Best-BlastP=> >nrprot 63% Identities = 199/41 δ (47%), Positives = 270/41 δ (65%), Gaps = 4/415 (0%) ref|NP_820906.11 transporter, putative [Coxiella burnetii RSA 493] gb|AA091420.11 transporter, putative [Coxiella burnetii RSA 493] Length = 435
3169.1 Best-BlastP=> >nrprot 50% Identities = 90/186 (48%), Positives = 122/186 (65%), Gaps = 3/186 (1 %) ref|NP_926264.11 unknown protein [Gloeobacter violaceus] dbj|BAC91259.11 glr3318 [Gloeobacter violaceus] Length = 228
317.1 Best-BlastP=> >nrprot 59% Identities = 142/362 (39%), Positives = 202/362 (55%), Gaps = 33/362 (9%) ref|ZP_00069313.1 | COG0517: FOG: CBS domain [Oenococcus oeni MCW] Length = 382
3172.2 Best-BlastP=> >nrprot 68% Identities = 118/270 (43%), Positives = 188/270 (69%), Gaps = 5/270 (1 %) ref|ZP_00014387.11 COG1752: Predicted esterase of the alpha-beta hydrolase superfamily [Rhodospirillum rubrum] Length = 369
3173.1 Best-BlastP=> >nrprot 63% Identities = 38/88 (43%), Positives = 54/88 (61 %), Gaps = 3/88 (3%) ref|ZP_00068002.1 | hypothetical protein [Microbulbifer degradans 2-40] Length = 128
3174.1 Best-BlastP=> >nrprot 53% Identities = 100/270 (37%), Positives = 155/270 (67%), Gaps = 1/270 (0%) ref|NP_903183.1 | conserved hypothetical protein [Chromobacterium violaceum ATCC 12472] gb|AAQ61174.1 | conserved hypothetical protein [Chromobacterium violaceum ATCC 12472] Length = 276
3175.1 Best-BlastP=> >nrprot 35% Identities = 51/201 (25%), Positives = 92/201 (46%), Gaps = 3/201 (1 %) ref|NP_903184.11 conserved hypothetical protein [Chromobacterium violaceum ATCC 12472] gb|AAQ61175.11 conserved hypothetical protein [Chromobacterium violaceum ATCC 12472] Length = 256
3177.1 Best-BlastP=> >nrprot 54% Identities = 64/138 (46%), Positives = 84/138 (60%), Gaps = 14/138 (10%) ref|NP_903185.11 conserved hypothetical protein [Chromobacterium violaceum ATCC 12472] gb|AAQ61176.11 conserved hypothefical protein [Chromobacterium violaceum ATCC 12472] Length = 145
3178.1 Best-BlastP=> >nrprot 72% Identities = 124/219 (56%), Positives = 157/219 (71 %), Gaps = 4/219 (1 %) ref|ZP_00066950.11 COG0036: Pentose- 5-phosphate-3-epimerase [Microbulbifer degradans 2-40] Length = 228
3179.1 Best-BlastP=> >nrprot 47% Identities = 179/574 (31 %), Positives = 288/674 (60%), Gaps = 25/674 (4%) ref|ZP_00067610.1 | COG0741 : Soluble lytic murein transglycosylase and related regulatory proteins (some contain LysM/invasin domains) [Microbulbifer degradans 2-40] Length = 669
318.2 Best-BlastP=> >nrprot 76% Identities = 266/465 (57%), Positives = 361/465 (77%), Gaps = 7/465 (1 %) ref|NP_902486.1 | probable PhoH-related protein [Chromobacterium violaceum ATCC 12472] gb|AAQ60484.1 | probable PhoH-related protein [Chromobacterium violaceum ATCC 12472] Length = 467
3181.2 Best-BlastP=> >nrprot 66% Identities = 154/327 (47%), Positives = 218/327 (66%) ref|ZP_00016063.11 COG0842: ABC-type multidrug transport system, permease component [Rhodospirillum rubrum] Length = 371
3184.3 Best-BlastP=> >nrprot 71 % Identities = 221/407 (54%), Positives = 299/407 (73%), Gaps = 7/407 (1 %) ref|NP_820219.1 | conserved hypothetical protein [Coxiella burnetii RSA 493] gb|AAO90733.11 conserved hypothetical protein [Coxiella burnetii RSA 493] Length = 445
3186.2 Best-BlastP=> >nrprot 71% Identities = 622/1146 (64%), Positives = 830/1145 (72%), Gaps = 9/1145 (0%) ref|NP_791924.11 transcription-repair coupling factor [Pseudomonas syringae pv. tomato str. DC3000] gb|AAOδδ619.11 transcription-repair coupling factor [Pseudomonas syringae pv. tomato str. DC3000] Length = 1160
3190.1 Best-BlastP=> >nrprot 42% Identities = 89/233 (38%), Positives = 127/233 (64%), Gaps = 2/233 (0%) ref|NP_δ19922.11 GALA PROTEIN 3 [Ralstonia solanacearum] Length = 622
3191.1 Best-BlastP=> >nrprot No Hits found
3193.2 Best-BlastP=> >nrprot 50% Identities = 77/223 (34%), Positives = 116/223 (52%), Gaps = 7/223 (3%) sp|P57974|RECO_PASMU DNA repair protein recO (Recombination protein O) Length = 240
3196.3 Best-BlastP=> >nrprot 65% Identities = 225/478 (47%), Positives = 330/478 (69%), Gaps = 2/478 (0%) ref|NP_462493.11 putative POT family, peptide transport protein [Salmonella typhimurium LT2] gb|AAL22452.11 putative POT family peptide transport protein [Salmonella typhimurium LT2] Length = 489
3198.1 Best-BlastP=> >nrprot 54% Identities = 72/174 (41 %), Positives = 101/174 (58%), Gaps = 1/174 (0%) ref|ZP_00081025.1 | COG0568: Phosphatidylglycerophosphate synthase [Geobacter metallireducens] Length = 198
3199.1 ' Best-BlastP=> >nrprot 98% Identities = 661/576 (97%), Positives = 566/575 (98%) gb|AAC44717.1 | FrgA [Legionella pneumophila] Length = 575
3201.1 Best-BlastP=> >nrprot 81 % Identities = 451/609 (74%), Positives = 520/609 (85%), Gaps = 3/609 (0%) ref|NP_820341.11 ATP-dependent metalloprotease FtsH [Coxiella burnetii RSA 493] gb|AAO90855.11 ATP-dependent metalloprotease FtsH [Coxiella burnetii RSA 493] Length = 647
3205.3 Best-BlastP=> >nrprot 46% Identities = 466/1207 (38%), Positives = 682/1207 (56%), Gaps = 45/1207 (3%) emb|CAC01603.11 peptide synthetase [Anabaena sp. 90] Length = 2258
3207.5 Best-BlastP=> >nrprot 50% Identities = 247/840 (29%), Positives = 419/840 (49%), Gaps = 91/840 (10%) ref|NP_819809.11 sensory box histidine kinase/response regulator [Coxiella burnetii RSA 493] gb|AAO90323.11 sensory box histidine kinase/response regulator [Coxiella burnetii RSA 493] Length = 808
321.3 Best-BlastP=> >nrprot 22% Identities = 44/165 (26%), Positives = 67/165 (40%), Gaps = 22/165 (13%) pir||T18253 probable mitochondrial carrier protein - yeast (Candida albicans) emb|CAA22027.11 putative mitochondrial carrier protein [Candida albicans] Length = 284
3212.2 Best-BlastP=> >nrprot 45% Identities = 64/206 (31 %), Positives = 105/206 (50%), Gaps = 11/206 (5%) ref|NP_389092.1 | similar to endo-1 ,4- beta-xylanase [Bacillus subtilis] sp|034798|YJEA_BACSU Hypothetical protein yjeA precursor pir||G69849 endo-1 ,4-beta-xylanase homolog yjeA Bacillus subtilis gb|AAC46306.11 NodB-like protein [Bacillus subtilis] emb|CAB13067.11 yjeA [Bacillus subtilis subsp. subtilis str. 168] Length = 467
3217.3 Best-BlastP=> >nrprot 79% Identities = 261/396 (65%), Positives = 314/396 (79%), Gaps = 3/396 (0%) ref|NP_819161.1 | 2-amino-3-ketobutyrate coenzyme A ligase [Coxiella burnetii RSA 4 3] gb|AA089675.1 | 2-amino-3-ketobutyrate coenzyme A ligase [Coxiella burnetii RSA 493] Length = 396
3218.1 Best-BlastP=> >nrprot 59% Identities = 94/211 (44%), Positives = 129/211 (61 %) ref|NP_840138.11 possible pcm; protein-L-isoaspartate o- methyltransferase [Nitrosomonas europaea ATCC 19718] emb|CAD83948.1 | possible pcm; protein-L-isoaspartate o-methyltransferase [Nitrosomonas europaea ATCC 19718] Length = 218
322.3 Best-BlastP=> >nrprot No Hits found
3221.1 Best-BlastP=> >nrprot 62% Identities = 187/450 (41 %), Positives = 275/450 (61%), Gaps = 16/450 (3%) ref|NP_819109.1 | outer membrane protein TolC, putative [Coxiella burnetii RSA 493] gb|AA089623.1 | outer membrane protein TolC, putative [Coxiella burnetii RSA 493] Length = 616
3223.1 Best-BlastP=> >nrprot 74% Identities = 182/248 (73%), Positives = 212/248 (85%) ref|NP_820067.11 conserved hypothetical protein [Coxiella
burnetii RSA 493] gb|AAO90581.11 conserved hypothetical protein [Coxiella burnetii RSA 493] Length = 256
3224.3 Best-BlastP=> >nrprot 62% Identities = 316/713 (44%), Positives = 464/713 (65%), Gaps = 14/713 (1%) ref|NP_869762.1 | probable sulfate transporter [Pirellula sp.] emb|CAD77140.11 probable sulfate transporter [Pirellula sp.] Length = 768
3226.2 Best-BlastP=> >nrprot 50% Identities = 28/64 (43%), Positives = 44/64 (68%) gb|AAP78483.11 C.Ahdl [Aeromonas hydrophila] Length = 74
3228.1 Best-BlastP=> >nrprot 70% Identities = 40/66 (60%), Positives = 49/66 (74%), Gaps = 1/66 (1 %) ref|ZP_00067276.1 | COG1278: Cold shock proteins [Microbulbifer degradans 2-40] Length = 71
323.2 Best-BlastP=> >nrprot No Hits found 3230.1 Best-BlastP=> >nrprot No Hits found 3231.1 Best-BlastP=> >nrprot 66% Identities = 25/62 (48%), Positives = 31/52 (59%) ref|NP_143190.11 hypothetical protein PH1305 [Pyrococcus horikoshii] pir||A71001 hypothetical protein PH1306 - Pyrococcus horikoshii dbj|BAA30409.1 | 252aa long hypothetical protein [Pyrococcus horikoshii] Length = 262
3232.1 Best-BlastP=> >nrprot No Hits found 3233.1 Best-BlastP=> >nrprot 19% Identities = 34/98 (34%), Positives = 52/98 (53%), Gaps = 1/98 (1 %) ref|ZP_00106280.1 | COG2197: Response regulator containing a CheY-like receiver domain and an HTH DNA-binding domain [Nostoc punctiforme] Length = 210
3234.3 Best-BlastP=> >nrprot 53% Identities = 71/228 (31 %), Positives = 125/228 (54%), Gaps = 1/228 (0%) ref|NP_928824.11 hypothetical protein [Photorhabdus luminescens subsp. laumondii TT01] emb|CAE13825.11 unnamed protein product [Photorhabdus luminescens subsp. laumondii TT01] Length = 231
3235.3 Best-BlastP=> >nrprot 66% Identities = 89/162 (54%), Positives = 120/162 (74%) ref|NP_311228.11 hypothetical protein [Escherichia coli 0157:H7] ref|NP_416820.1 | orf, hypothetical protein [Escherichia coli K12] sp|P09548|DEDA_ECOLI DedA protein (DSG-1 protein) pir||XMECAD dedA protein - Escherichia coli (strain K-12) pir||A98029 hypothetical protein ECs3201 [imported] - Escherichia coli (strain 01δ7:H7, substrain RIMD 0609952) gb|AAA23964.1 | dedA gb|AAC75377.1 | orf, hypothetical protein [Escherichia coli K12] dbj|BAA16174.1 | dedA protein [Escherichia coli] dbj|BAB36624.1 | hypothetical protein [Escherichia coli 0157:H7] Length = 219
3236.3 Best-BlastP=> >nrprot No Hits found
3238.1 Best-BlastP=> >nrprot 67% Identities = 134/263 (50%), Positives = 195/263 (74%), Gaps = 1/263 (0%) ref|NP_421700.11 conserved hypothetical protein [Caulobacter crescentus CB15] pir||H87608 conserved hypothetical protein CC2906 [imported] - Caulobacter crescentus gb|AAK24868.1 | conserved hypothetical protein [Caulobacter crescentus CB1δ] Length = 289
3239.1 Best-BlastP=> >nrprot 68% Identities = 31/85 (36%), Positives = 57/85 (67%), Gaps = 2/85 (2%) ref|NP_820672.11 hypothefical protein [Coxiella burnetii RSA 493] gb|AA091086.11 hypothetical protein [Coxiella burnetii RSA 493] Length = 102
324.3 Best-BlastP=> >nrprot 72% Identities = 106/224 (47%), Positives = 160/224 (71 %), Gaps = 5/224 (2%) ref|ZP_00045186.11 COG2200: FOG: EAL domain [Magnetococcus sp. MC-1] Length = 577
3240.3 Best-BlastP=> >nrprot 52% Identities = 24/64 (37%), Positives = 42/64 (65%) ref|NP_458973.11 Putative periplasmic protein [Salmonella enterica subsp. enterica serovar Typhi] ref|NP_463417.1 | hyperosmotically inducible periplasmic protein, RpoS-dependent stationary phase gene [Salmonella typhimurium LT2] ref|NP_808176.11 Putative periplasmic protein [Salmonella enterica subsp. enterica serovar Typhi Ty2]
pir||AE1072 Putative periplasmic protein [imported] - Salmonella enterica subsp. enterica serovar Typhi (strain CT18) gb|AAL23376.11 hyperosmotically inducible periplasmic protein [Salmonella typhimurium LT2] emb|CAD03396.1 | Putative periplasmic protein [Salmonella enterica subsp. enterica serovar Typhi] gb|AAO72036.1 | Putative periplasmic protein [Salmonella enterica subsp. enterica serovar Typhi Ty2] Length = 205
3242.3 Best-BlastP=> >nrprot 14% Identities = 72/325 (22%), Positives = 126/325 (38%), Gaps = 37/325 (11 %) gb|AAB96623.11 spheroidin [Heliothis armigera entomopoxvirus] Length = 1007
3243.1 Best-BlastP=> >nrprot No Hits found
3244.2 Best-BlastP=> >nrprot 60% Identities = 102/241 (42%), Positives = 153/241 (63%), Gaps = 2/241 (0%) ref|NP_465213.11 similar to glucose 1- dehydrogenase [Listeria monocytogenes EGD-e] pir||AH128δ glucose 1 -dehydrogenase homolog lmo1688 [imported] - Listeria monocytogenes (strain EGD-e) emb|CAC99766.1 | lmo1688 [Listeria monocytogenes] Length = 248
3246.2 Best-BlastP=> >nrprot 24% Identities = 100/360 (27%), Posifives = 160/360 (44%), Gaps = 67/360 (18%) ref|NP_g05966.11 conserved hypothetical protein [Porphyromonas gingivalis W83] gb|AAQ66865.1 | conserved hypothetical protein [Porphyromonas gingivalis W83] Length = 33g
3248.1 Best-BlastP=> >nrprot 38% Identities = 87/154 (56%), Positives = 114/154 (74%) ref|ZP_00111795.11 COG0784: FOG: CheY-like receiver [Nostoc punctiforme] Length = 557
3249.2 Best-BlastP=> >nrprot 13% Identities = 22/55 (40%), Positives = 29/55 (52%) ref|NP_798062.11 hypothetical protein VP1683 [Vibrio parahaemolyticus RIMD 2210633] dbj|BAC59946.11 hypothetical protein [Vibrio parahaemolyticus] Length = 325
325.1 Best-BlastP=> >nrprot No Hits found 3250.1 Best-BlastP=> >nrprot No Hits found
3251.1 Best-BlastP=> >nrprot 48% Identities = 39/131 (29%), Positives = 65/131 (49%), Gaps = 3/131 (2%) emb|CAC05487.11 YcfB protein [Erwinia amylovora] Length = 132
3252.1 Best-BlastP=> >nrprot No Hits found
3257.2 Best-BlastP=> >nrprot 46% Identities = 40/122 (32%), Positives = 70/122 (57%), Gaps = 4/122 (3%) ref|NP_75δδδ6.11 Hypothetical protein [Escherichia coli CFT073] gb|AAN82129.1 |AE016766_217 Hypothetical protein [Escherichia coli CFT073] Length = 131
3258.2 Best-BlastP=> >nrprot 35% Identities = 70/225 (31 %), Positives = 111/225 (49%), Gaps = 27/225 (12%) ref|NP_756567.1 | Hypothetical protein [Escherichia coli CFT073] gb|AAN82130.1 |AE016766_218 Hypothetical protein [Escherichia coli CFT073] Length = 305
326.2 Best-BlastP=> >nrp rrot No Hits found
3260.2 Best-BlastP=> >nrprot No Hits found
3263.2 Best-BlastP=> >nrprot 61 % Identities = 128/368 (34%), Positives = 227/368 (61 %), Gaps = 10/368 (2%) ref|NP_819469.11 major facilitator family transporter [Coxiella burnetii RSA 493] gb|AA089983.11 major facilitator family transporter [Coxiella burnetii RSA 493] Length = 428
3265.1 Best-BlastP=> >nrprot 65% Identifies = 140/293 (47%), Positives = 195/293 (66%), Gaps = 2/293 (0%) ref|NP_820459.11 hydrogen peroxide- inducible genes activator OxyR [Coxiella burnetii RSA 493] gb|AAO90973.1 1 hydrogen peroxide-inducible genes activator OxyR [Coxiella burnetii RSA 493] Length = 311
3267.1 Best-BlastP=> >nrprot 50% Identities = 57/258 (22%), Positives = 104/258 (40%), Gaps = 75/258 (29%) ref|NP_797353.11 hypothetical protein VP0974 [Vibrio parahaemolyticus RIMD 2210633] dbj|BAC59237.11 hypothetical protein [Vibrio parahaemolyticus] Length = 269 .
3269.1 Best-BlastP=> >nrprot 56% Identities = 400/11 13 (35%), Positives = 600/1113 (53%), Gaps = 64/1113 (5%) ref|NP_82022δ.11 UvrD/REP helicase family protein [Coxiella burnetii RSA 493] gb|AAO90739.11 UvrD/REP helicase family protein [Coxiella burnetii RSA 493] Length = 1110
327.2 Best-BlastP=> >nrprot 70% Identities = 364/616 (59%), Positives = 458/616 (74%), Gaps = 7/616 (1 %) ref|ZP_00141174.1 | hypothetical protein [Pseudomonas aeruginosa UCBPP-PA14] Length = 645
3271.3 Best-BlastP=> >nrprot 46% Identities = 187/674 (27%), Positives = 308/674 (45%), Gaps = 55/674 (8%) ref|NP_437090.11 putative membrane- located cell surface saccharide saccharide acetylase protein [Sinorhizobium meliloti] pir||F95910 probable membrane-located cell surface saccharide saccharide acetylase protein [imported] - Sinorhizobium meliloti (strain 1021 ) magaplasmid pSymB emb|CAC48950.1 | putative membrane-located cell surface saccharide saccharide acetylase protein [Sinorhizobium meliloti] Length = 677
3272.1 Best-BlastP=> >nrprot 55% Identities = 49/148 (33%), Positives. = 82/148 (55%), Gaps = 14/148 (9%) ref|NP_820842.11 conserved hypothetical protein [Coxiella burnetii RSA 493] gb|AA091356.11 conserved hypothetical protein [Coxiella burnetii RSA 493] Length = 175
3273.1 Best-BlastP=> >nrprot 40% Identities = 34/95 (35%), Positives = 50/96 (52%), Gaps = 4/95 (4%) ref|NP_93085δ.11 hypothetical protein [Photorhabdus luminescens subsp. laumondii TT01] emb|CAE16020.1 | unnamed protein product [Photorhabdus luminescens subsp. laumondii TT01] Length = 175
3275.1 Best-BlastP=> >nrprot No Hits found
3276.2 Best-BlastP=> >nrprot No Hits found
3277.2 Best-BlastP=> >nrprot 61 % Identities = 109/264 (41 %), Positives = 165/264 (62%), Gaps = 5/264 (1 %) ref|ZP_00055240.11 COG1073: Hydrolases of the alpha/beta superfamily [Magnetospirillum magnetotacticum] Length = 297
3278.1 Best-BlastP=> >nrprot 28% Identities = 19/43 (44%), Positives = 28/43 (65%) ref|NP_523079.1 1 CONSERVED HYPOTHETICAL PROTEIN [Ralstonia solanacearum] emb|CAD18671.1 1 CONSERVED HYPOTHETICAL PROTEIN [Ralstonia solanacearum] Length = 232
328.1 Best-BlastP=> >nrprot 68% Identities = 161/292 (55%), Positives = 206/292 (70%), Gaps = 3/292 (1 %) ref|NP_770597.1 1 3-hydroxyisobutyrate dehydrogenase [Bradyrhizobium japonicum] dbj|BAC49222.11 3-hydroxyisobutyrate dehydrogenase [Bradyrhizobium japonicum USDA 110] Length = 29δ
3280.1 Best-BlastP=> >nrprot 54% Identities = 101/267 (37%), Positives = 156/267 (58%), Gaps = 18/267 (6%) ref|NP_631554.1 | conserved hypothetical protein [Streptomyces coelicolor A3(2)] emb|CAC44686.1 | conserved hypothetical protein [Streptomyces coelicolor A3(2)] . Length = 277 3281.1 Best-BlastP=> >nrprot No Hits found
3282.3 Best-BlastP=> >nrprot 82% Identities = 289/413 (69%), Positives = 341/413 (82%), Gaps = 4/413 (0%) ref|NP_819190.11 cell division protein FtsA [Coxiella burnetii RSA 493] gb|AAO89704.11 cell division protein FtsA [Coxiella burnetii RSA 493] Length = 410
3283.1 Best-BlastP=> >nrprot 60% Identities = 72/220 (32%), Positives = 120/220 (64%), Gaps = 1/220 (0%) ref|NP_253099.11 cell division protein FtsQ [Pseudomonas aeruginosa PA01] pir||B83094 cell division protein FtsQ PA4409 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAF26457.1 | FtsQ [Pseudomonas aeruginosa] gb|AAG07797.1 |AE004866_8 cell division protein FtsQ [Pseudomonas aeruginosa PA01] Length = 287
3286.3 Best-BlastP=> >nrprot 48% Identities = 106/303 (34%), Positives = 174/303 (57%), Gaps = 1/303 (0%) ref|ZP_00032705.11 COG0642: Signal transducfion histidine kinase [Burkholderia fungorum] Length = 536
3288.3 Best-BlastP=> >nrprot 71 % Identities = 120/221 (54%), Positives = 161 /221 (72%) ref|ZP_00080602.11 COG0745: Response regulators consisting of a CheY-like receiver domain and a winged-helix DNA-binding domain [Geobacter metallireducens] Length = 306
3290.1 Best-BlastP=> >nrprot No Hits found
3291.2 Best-BlastP=> >nrprot No Hits found
3295.2 Best-BlastP=> >nrprot 77% Identities = 563/923 (60%), Positives = 715/923 (77%), Gaps = 5/923 (0%) ref|ZP_00066862.11 COG0525: Valyl- tRNA synthetase [Microbulbifer degradans 2-40] Length = 922
3297.3 Best-BlastP=> >nrprot 36% Identities = 75/282 (26%), Positives = 129/282 (45%), Gaps = 11/282 (3%) ref|NP_603055.1 | 3-oxoacyl-[acyl-car er- protein] synthase III [Fusobacterium nucleatum subsp. nucleatum ATCC 25586] gb|AAL94354.1 | 3-oxoacyl-[acyl-carrier-protein] synthase Ill [Fusobacterium nucleatum subsp. nucleatum ATCC 26586] Length = 328
3299.3 Best-BlastP=> >nrprot 36% Identities = 36/137 (26%), Positives = 51/137 (37%), Gaps = 37/137 (27%) ref|ZP_00014317.1 | hypothetical protein [Rhodospirillum rubrum] Length = 417
33.1 Best-BlastP=> >nrprot 87% Identities = 711/878 (80%), Positives = 774/878 (88%), Gaps = 16/878 (1%) gb|AAG45149.11 TraA-like protein [Legionella pneumophila] Length = 883
330.2 Best-BlastP=> >nrprot 70% Identities = 267/493 (54%), Positives = 353/493 (71 %), Gaps = 1/493 (0%) ref |NP_819940.11 methylmalonate- semialdehyde dehydrogenase [Coxiella burnetii RSA 493] gb|AAO90454.1 | methylmalonate-semialdehyde dehydrogenase [Coxiella burnefii RSA 493] Length = 498
3301.2 Best-BlastP=> >nrprot 55% Identities = 160/488 (32%), Positives = 261/488 (53%), Gaps = 38/488 (7%) ref|NP_787151.1 | propionyl-CoA carboxylase beta chain [Tropheryma whipplei str. Twist] ref|NP_788980.11 propionyl-CoA carboxylase beta chain [Tropheryma whipplei TW08/27] emb|CAD66717.11 propionyl-CoA carboxylase beta chain [Tropheryma whipplei TW08/27] gb|AA044120.11 propionyl-CoA carboxylase beta chain [Tropheryma whipplei str. Twist] Length = 525
3303.2 Best-BlastP=> >nrprot No Hits found
3304.2 Best-BlastP=> >nrprot 50% Identities = 133/302 (44%), Positives = 189/302 (62%), Gaps = 7/302 (2%) ref|ZP_00043253.11 hypothetical protein [Magnetococcus sp. MC-1] Length = 831
3306.4 Best-BlastP=> >nrprot 29% Identities = 48/138 (34%), Positives = 71/138 (61 %), Gaps = 13/138 (9%) ref|ZP_000δ254δ.1 | COG1020: Non- ribosomal peptide synthetase modules and related proteins [Magnetospirillum magnetotacticum] Length = 676
3307.4 Best-BlastP=> >nrprot No Hits found 3308.4 Best-BlastP=> >nrprot No Hits found
331.2 Best-BlastP=> >nrprot 69% Identities = 272/538 (50%), Positives = 371/538 (68%), Gaps = 5/538 (0%) ref |NP_406414.11 putative glutamine- dependent NAD [Yersinia pestis] ref|NP_668639.11 putative NH3-dependent NAD(+) synthetase [Yersinia pestis KIM] pir||AG0354 NAD synthase (glutamine-hydrolysing) (EC 6.3.5.1 ) - Yersinia pestis (strain C092) emb|CAC92162.11 putative glutamine-dependent NAD [Yersinia pestis C092] gb|AAM84890.1 |AE013734_5 putative NH3-dependent NAD(+) synthetase [Yersinia pestis KIM] Length = 640
3311.1 Best-BlastP=> >nrprot 74% Identities = 275/469 (59%), Positives = 341/459 (74%), Gaps = 9/459 (1 %) ref|NP_260486.11 cysteinyl-tRNA synthetase [Pseudomonas aeruginosa PA01] sp|Q9l2U7|SYC_PSEAE Cysteinyl-tRNA synthetase (Cysteine-tRNA ligase) (CysRS) pir||G83421 cysteinyl-tRNA synthetase PA1795 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG05184.1 |AE00460δ_6 cysteinyl-tRNA synthetase [Pseudomonas aeruginosa PA01] Length = 460
3312.1 Best-BlastP=> >nrprot 32% Identities = 20/66 (30%), Positives = 34/66 (51 %) ref|NP_759δ13.11 Unknown [Vibrio vulnificus CMCP6] gb|AAO09040.1 |AE016798_200 Unknown [Vibrio vulnificus CMCP6] dbj|BAC93437.11 hypothetical protein [Vibrio vulnificus YJ016] Length = 103
3314.2 Best-BlastP=> >nrprot No Hits found
3316.1 Best-BlastP=> >nrprot 26% Identifies = 80/363 (22%), Positives = 152/363 (41 %), Gaps = 41/363 (11 %) ref|NP_659473.11 hypothetical protein MGC33887 [Homo sapiens] gb|AAM49719.1 |AF458591_1 hypothetical protein [Homo sapiens] Length = 590
3319.1 Best-BlastP=> >nrprot No Hits found
3322.2 Best-BlastP=> >nrprot 98% Identities = 265/269 (98%), Positives = 267/269 (99%) pir||T18335 icmG protein - Legionella pneumophila emb|CAA75166.1 | IcmG protein [Legionella pneumophila] gb|AAC38187.1 | DotF [Legionella pneumophila] emb|CAA75332.1 | IcmG protein [Legionella pneumophila] Length = 269
3323.1 Best-BlastP=> >nrprot 99% Identities = 194/194 (100%), Positives = 194/194 (100%) pir||T18336 icmC protein - Legionella pneumophila emb|CAA75167.1 | IcmC protein [Legionella pneumophila] gb|AAC38186.1 | DotE [Legionella pneumophila] emb|CAA75333.1 | IcmC protein [Legionella pneumophila] Length = 194
3324.1 Best-BlastP=> >nrprot 98% Identities = 130/132 (98%), Positives = 131/132 (99%) pir||T18337 icmD protein - Legionella pneumophila emb|CAA75168.1 | IcmD protein [Legionella pneumophila] emb|CAA75334.1 | icmD protein [Legionella pneumophila] Length = 132
3326.2 Best-BlastP=> >nrprot 99% Identities = 261/261 (100%), Positives = 261/261 (100%) gb|AAQ10306.11 DotU [Legionella pneumophila] Length = 261
3327.1 Best-BlastP=> >nrprot 46% Identities = 53/219 (24%), Positives = 99/219 (45%), Gaps = 19/219 (8%) ref|NP_840938.1 | conserved hypothetical protein [Nitrosomonas europaea ATCC 19718] emb|CAD8477δ.1 | conserved hypothetical protein [Nitrosomonas europaea ATCC 19718] Length = 343
3328.1 Best-BlastP=> >nrprot 70% Identities = 316/576 (54%), Posifives = 407/576 (70%), Gaps = 5/675 (0%) ref |NP_252414.11 single-stranded-DNA- specific exonuclease RecJ [Pseudomonas aeruginosa PAOt] pir||A83181 single-stranded-DNA-specific exonuclease RecJ PA3725 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG07112.1 |AE004791_9 single-stranded-DNA-specific exonuclease RecJ [Pseudomonas aeruginosa PA01 ] Length = 571
333.3 Best-BlastP=> >nrprot 57% Identities = 426/1088 (39%), Positives = 613/1088 (56%), Gaps = 70/1088 (6%) gb|AAP85938.1 | putative helicase, superfamily II [Ralstonia eutropha] Length = 1106
3331.1 Best-BlastP=> >nrprot 61 % Identities = 145/291 (49%), Positives = 203/291 (69%), Gaps = 1/291 (0%) ref|NP_719443.11 TIM-barrel protein, yjbN family [Shewanella oneidensis MR-1] gb|AAN56887.1 |AE015823_9 TIM-barrel protein, yjbN family [Shewanella oneidensis MR-1] Length = 335
3333.1 Best-BlastP=> >nrprot 58% Identities = 52/122 (42%), Positives = 79/122 (64%), Gaps = 1/122 (0%) ref|NP_930852.11 Mutator mutT protein (7,8- dihydro-8-oxoguanine-triphosphatase) (8-oxo-dGTPase) (dGTP pyrophosphohydrolase) [Photorhabdus luminescens subsp. laumondii TT01] emb|CAE16017.11 Mutator mutT protein (7,8-dihydro-8-oxoguanine-triphosphatase) (8-oxo-dGTPase) (dGTP pyrophosphohydrolase) [Photorhabdus luminescens subsp. laumondii TT01] Length = 130
3334.2 Best-BlastP=> >nrprot No Hits found 3336.2 Best-BlastP=> >nrprot No Hits found 3337.2 Best-BlastP=> >nrprot No Hits found
3338.1 Best-BlastP=> >nrprot No Hits found
334.5 Best-BlastP=> >nrprot 42% Identities = 272/697 (39%), Positives = 416/697 (59%), Gaps = 23/697 (3%) ref|NP_440178.11 regulatory components of sensory transduction system [Synechocystis sp. PCC 6803] pir||S74707 nitrogen fixation positive acfivator protein - Synechocystis sp. (strain PCC 6803) dbj|BAA16858.11 regulatory components of sensory transduction system [Synechocystis sp. PCC 6803] Length = 840
3341.2 Best-BlastP=> >nrprot No Hits found
3343.1 Best-BlastP=> >nrprot No Hits found 3345.2 Best-BlastP=> >nrprot 17% Identities = 53/272 (19%), Positives = 137/272 (50%), Gaps = 28/272 (10%) ref|NP_701067.11 hypothetical protein [Plasmodium falciparum 3D7] gb|AAN35791.1 |AE014838_69 hypothetical protein [Plasmodium falciparum 3D7] Length = 964
3346.2 Best-BlastP=> >nrprot 28% Identities = 105/343 (30%), Positives = 168/343 (48%), Gaps = 60/343 (17%) ref|NP_623492.11 Cell division protein Ftsl/penicillin-binding protein 2 [Thermoanaerobacter tengcongensis] gb|AAM25096.11 Cell division protein Ftsl/penicillin-binding protein 2 [Thermoanaerobacter tengcongensis] Length = 563
3347.1 Best-BlastP=> >nrprot 94% Identities = 239/266 (89%), Positives = 251/266 (94%) emb|CAC35728.11 OXA-29 [Fluoribacter gormanii] Length = 266
3348.2 Best-BlastP=> >nrprot 31 % Identities = 115/296 (38%), Positives = 170/295 (57%), Gaps = 15/295 (5%) ref|ZP_00119503.11 COG0726: Predicted xylanase/chitin deacetylase [Cytophaga hutchinsonii] Length = 314
3349.3 Best-BlastP=> >nrprot 45% Identities = 72/238 (30%), Positives = 130/238 (54%), Gaps = 8/238 (3%) ref|NP_638901.11 conserved hypothetical protein [Xanthomonas campestris pv. campestris str. ATCC 33913] gb|AAM42825.1 | conserved hypothefical protein [Xanthomonas campestris pv. campestris str. ATCC 33913] Length = 314
336.1 Best-BlastP=> >nrprot 67% Identities = 115/207 (55%), Positives = 145/207 (70%), Gaps = 2/207 (0%) ref|NP_903041.11 conserved hypothetical protein [Chromobacterium violaceum ATCC 12472] gb|AAQ61035.11 conserved hypothetical protein [Chromobacterium violaceum ATCC 12472] Length = 206
3350.3 Best-BlastP=> >nrprot 41 % Identities = 33/101 (32%), Positives = 50/101 (49%), Gaps = 3/101 (2%) ref|ZP_00116659.11 COG0346: Lactoylglutathione lyase and related lyases [Cytophaga hutchinsonii] Length = 150
3351.3 Best-BlastP=> >nrprot 30% Identities = 88/194 (45%), Posifives = 121/194 (62%), Gaps = 1/194 (0%) ref|NP_531634.11 methyltransferase [Agrobacterium tumefaciens str. C58 (U. Washington)] pir||AH2691 methyltransferase Atu0936 [imported] - Agrobacterium tumefaciens (strain C58, Dupont) gb|AAL41950.1 | methyltransferase [Agrobacterium tumefaciens str. C58 (U. Washington)] Length = 202
3352.1 Best-BlastP=> >nrprot 42% Identities = 52/109 (47%), Positives = 67/109 (61 %), Gaps = 6/109 (5%) ref|NP_819444.11 acetyltransferase, GNAT family [Coxiella burnetii RSA 493] gb|AA089958.11 acetyltransferase, GNAT family [Coxiella burnetii RSA 493] Length = 118
3353.1 Best-BlastP=> >nrprot 48% Identities = 157/467 (33%), Positives = 245/467 (52%), Gaps = 16/467 (3%) ref|NP_417114.11 hypothetical protein [Escherichia coli K12] sp|P52124|YFJI_ECOLI Hypothetical protein yfjl pir||T08637 hypothetical protein b2625 - Escherichia coli (strain K-12) gb|AAA79794.11 ORF_o469 gb|AAC75673.11 orf, hypothetical protein [Escherichia coli K12] Length = 469
3354.1 Best-BlastP=> >nrprot 47% Identities = 58/190 (30%), Positives = 91/190 (47%), Gaps = 2/190 (1 %) gb|AAD53919.1 |AF179611_3 pyridoxamine δ'-phosphate oxidase [Zymomonas mobilis] Length = 192
3356.2 Best-BlastP=> >nrprot 72% Identifies = 53/94 (66%), Positives = 70/94 (74%), Gaps = 1/94 (1 %) gb|AAL56698.1 |AF246719_2 hypothetical protein [Escherichia coli] Length = δ
3366.2 Best-BlastP=> >nrprot 70% Identities = 44/83 (53%), Positives = 59/83 (71 %) ref|NP_232869.11 conserved hypothetical protein [Vibrio cholerae 01 biovar eltor str. N 16961] pir||E82456 conserved hypothetical protein VCA0477 [imported] - Vibrio cholerae (strain N16961 serogroup 01 ) gb|AAF96381.11 conserved hypothetical protein [Vibrio cholerae 01 biovar eltor str. N16961] Length = 90
3357.2 Best-BlastP=> >nrprot No Hits found
3359.2 Best-BlastP=> >nrprot 48% Identities = 221/706 (31 %), Positives = 362/706 (51 %), Gaps = 53/706 (7%) ref|NP_820588.11 membrane protein, putative [Coxiella burnetii RSA 493] gb|AA091102.11 membrane protein, putative [Coxiella burnetii RSA 493] Length = 698
336.1 Best-BlastP=> >nrprot 69% Identities = 186/324 (57%), Positives = 230/324 (70%), Gaps = 2/324 (0%) ref|NP_384292.11 PHOSPHATE TRANSPORT TRANSMEMBRANE PROTEIN [Sinorhizobium meliloti] sp|O30499|PIT_RHIME Probable low-affinity inorganic phosphate transporter gb|AAB70171.1 | phosphate transport protein [Sinorhizobium meliloti] emb|CAC41573.11 PHOSPHATE TRANSPORT TRANSMEMBRANE PROTEIN [Sinorhizobium meliloti] Length = 334
3360.1 Best-BlastP=> >nrprot No Hits found
3362.1 Best-BlastP=> >nrprot 43% Identities = 68/209 (32%), Positives = 106/209 (50%), Gaps = 5/209 (2%) ref|NP_178286.11 membrane protein - related [Arabidopsis thaliana] pir||H84428 probable membrane protein [imported] - Arabidopsis thaliana gb|AAD12695.11 putative membrane protein [Arabidopsis thaliana] Length = 250
3363.2 Best-BlastP=> >nrprot 64% Identities = 148/300 (4g%), Positives = 204/300 (68%), Gaps = 11/300 (3%) gb|AAC64375.11 thiamine synthase homolog [Botryotinia fuckeliana] Length = 342
3365.2 Best-BlastP=> >nrprot 66% Identities = 173/347 (49%), Positives = 238/347 (68%), Gaps = 13/347 (3%) ref|NP_819373.11 thiamine biosynthesis oxidoreductase ThiO, putative [Coxiella burnetii RSA 493] gb|AA089887.11 thiamine biosynthesis oxidoreductase ThiO, putative [Coxiella burnetii RSA 493] Length = 338
3367.1 Best-BlastP=> >nrprot 41 % Identities = 81/269 (30%), Positives = 123/269 (45%), Gaps = 20/269 (7%) emb|CAC17409.11 3'- nucleotidase/nuclease [Leishmania mexicana] Length = 378
3368.2 Best-BlastP=> >nrprot No Hits found 337.2 Best-BlastP=> >nrprot 67% Identities = 241/457 (52%), Positives = 311/457 (68%), Gaps = 4/457 (0%) ref|NP_830123.11 Amino acid permease [Bacillus cereus ATCC 14579] gb|AAP07324.1 | Amino acid permease [Bacillus cereus ATCC 14579] Length = 471
3371.2 Best-BlastP=> >nrprot 83% Identities = 321/438 (73%), Positives = 372/438 (84%) ref|NP_819δ9g.11 conserved hypothetical protein [Coxiella burnetii RSA 493] gb|AAO90113.11 conserved hypothefical protein [Coxiella burnetii RSA 493] Length = 439
3374.1 Best-BlastP=> >nrprot 2δ% Identities = 60/222 (27%), Posifives = 94/222 (42%), Gaps = 21/222 (9%) ref|NP_774046.11 bll7406 [Bradyrhizobium japonicum] dbj|BAC52671.1 | bll7406 [Bradyrhizobium japonicum USDA 110] Length = 300
3376.1 Best-BlastP=> >nrprot 30% Identities = 62/236 (26%), Positives = 106/236 (44%), Gaps = 15/236 (6%) ref|NP_819950.11 conserved hypothetical protein [Coxiella burnetii RSA 493] gb|AAO90464.11 conserved hypothetical protein [Coxiella burnetii RSA 493] Length = 465
3378.2 Best-BlastP=> >nrprot 56% Identities = 210/580 (36%), Positives = 331/580 (57%), Gaps = 9/580 (1 %) gb|AAC44717.1 | FrgA [Legionella pneumophila] Length = 67δ
3386.1 Best-BlastP=> >nrprot 74% Identities = 252/264 (99%), Positives = 254/254 (100%) gb|AAM00642.11 essential conserved GTPase [Legionella pneumophila] Length = 254
3386.1 Best-BlastP=> >nrprot 77% Identities = 62/85 (72%), Positives = 72/85 (84%) ref|NP_636525.11 50S ribosomal protein L27 [Xanthomonas campestris pv. campestris str. ATCC 33913] sp|Q8PBH1 |RL27_XANCP 50S ribosomal protein L27 gb|AAM40449.11 60S ribosomal protein L27 [Xanthomonas campestris pv. campestris str. ATCC 33913] Length = 86
3388.1 Best-BlastP=> >nrprot 28% Identities = 65/168 (38%), Positives = 105/168 (62%), Gaps = 1/168 (0%) ref|ZP_00125980.11 COG2199: FOG: GGDEF domain [Pseudomonas syringae pv. syringae B728a] Length = 972
3389.1 Best-BlastP=> >nrprot 68% Idenfities = 149/315 (47%), Positives = 222/315 (70%) ref|ZP_00068004.11 COG0142: Geranylgeranyl pyrophosphate synthase [Microbulbifer degradans 2-40] Length = 320
339.4 Best-BlastP=> >nrprot 89% Identities = 168/194 (86%), Positives = 174/194 (89%), Gaps = 8/194 (4%) gb|AAK52070.11 Rep [Legionella pneumophila] Length = 186
3390.1 Best-BlastP=> >nrprot 37% Identities = 21 /61 (34%), Positives = 32/61 (52%) ref|NP_820745.11 conserved hypothetical protein [Coxiella burnetii RSA 493] gb|AA091259.11 conserved hypothetical protein [Coxiella burnefii RSA 493] Length = 72
3391.2 Best-BlastP=> >nrprot 99% Identities = 748/751 (99%), Positives = 749/751 (99%) gb|AAN17185.1 |AF492466_3 ferrous iron transporter B [Legionella pneumophila] Length = 751
3394.1 Best-BlastP=> >nrprot No Hits found
3395.2 Best-BlastP=> >nrprot No Hits found 3396.1 Best-BlastP=> >nrprot No Hits found 3397.1 Best-BlastP=> >nrprot 62% Identities = 72/167 (43%), Positives = 106/167 (63%), Gaps = 6/167 (3%) ref|NP_840350.11 putative antirestriction protein [Nitrosomonas europaea ATCC 19718] emb|CAD84171.11 putative antirestriction protein [Nitrosomonas europaea ATCC 19718] Length = 171
3398.1 Best-BlastP=> >nrprot 61 % Identities = 58/146 (40%), Positives = 87/145 (60%), Gaps = 7/145 (4%) ref|NP_252922.1 | single-stranded DNA- binding protein [Pseudomonas aeruginosa PA01] sp|P40947|SSB_PSEAE Single-strand binding protein (SSB) (Helix-destabilizing protein) pir||S44302 single-stranded DNA-binding protein - Pseudomonas aeruginosa emb|CAA83688.1 | single-stranded DNA binding protein [Pseudomonas aeruginosa] gb|AAG07620.1 |AE004840_6 single-stranded DNA-binding protein [Pseudomonas aeruginosa PA01] Length = 166
3399.1 Best-BlastP=> >nrprot No Hits found 34.1 Best-BlastP=> >nrprot No Hits found
3401.2 Best-BlastP=> >nrprot No Hits found 3402.2 Best-BlastP=> >nrprot 53% Identities = 108/298 (36%), Positives = 160/298 (53%), Gaps = 14/298 (4%) ref|NP_862370.11 unknown [Francisella tularensis subsp. novicida] gb|AAD17308.11 unknown [Francisella tularensis subsp. novicida] Length = 292
3404.2 Best-BlastP=> >nrprot 31 % Identities = 51/187 (27%), Positives = 86/187 (45%), Gaps = 9/187 (4%) ref|ZP_00091084.11 COG0582: Integrase [Azotobacter vinelandii] Length = 287
3406.2 Best-BlastP=> >nrprot 34% Identities = 46/197 (23%), Positives = 76/197 (38%), Gaps = 13/197 (6%) ref|NP_466297.11 hypothefical membrane protein [Listeria monocytogenes EGD-e] pir||AF1421 hypothetical membrane protein lmo2775 [imported] - Listeria monocytogenes (strain EGD-e) emb|CAD00988.11 lmo2775 [Listeria monocytogenes] Length = 722
341.6 Best-BlastP=> >nrprot 99% Identities = 141/141 (100%), Positives = 141/141 (100%) gb|AAC44222.1 | hemin binding protein Hbp [Legionella pneumophila] Length = 141
3411.2 Best-BlastP=> >nrprot 20% Identities = 42/162 (25%), Posifives = 75/162 (46%), Gaps = 10/162 (6%) ref|NP_871522.1 | rplA [Wigglesworthia glossinidia endosymbiont of Glossina brevipalpis] sp|Q8D236|RL1_WIGBR 50S ribosomal protein L1 dbj|BAC24665.11 rplA [Wigglesworthia brevipalpis] Length = 242
3413.2 Best-BlastP=> >nrprot 49% Identities = 76/296 (25%), Positives = 150/296 (50%), Gaps = 16/296 (5%) ref|NP_832129.11 Transcriptional regulators, LysR family [Bacillus cereus ATCC 14679] gb|AAP09330.11 Transcriptional regulators, LysR family [Bacillus cereus ATCC 14679] Length = 300
3414.3 Best-BlastP=> >nrprot 33% Identities = 116/406 (28%), Positives = 184/406 (46%), Gaps = 47/406 (11 %) ref|ZP_00112010.11 COG0666: FOG: Ankyrin repeat [Nostoc punctiforme] Length = 427
3415.2 Best-BlastP=> >nrprot 38% Identities = 48/154 (31 %), Positives = 69/154 (44%), Gaps = 14/154 (9%) ref|NP_740808.1 | dotll Histone Methyltransferase (1C952) [Caenorhabditis elegans] gb|AAK39620.2| Hypothetical protein Y39G10AR.18a [Caenorhabditis elegans] Length = 1016
3416.1 Best-BlastP=> >nrprot 73% Identities = 65/99 (65%), Positives = 76/99 (76%) ref|NP_820718.11 integration host factor, beta subunit [Coxiella burnetii RSA 493] gb|AA091232.11 integration host factor, beta subunit [Coxiella burnetii RSA 493] Length = 112
3417.1 Best-BlastP=> >nrprot 91 % Identities = 166/191 (81 %), Positives = 172/191 (90%), Gaps = 3/191 (1 %) ref|NP_780077.1 | deoxycytidine triphosphate deaminase [Xylella fastidiosa Temeculal] sp|Q87AD1 |DCD_XYLFT Deoxycytidine triphosphate deaminase (dCTP deaminase) gb|AA029726.11 deoxycytidine triphosphate deaminase [Xylella fastidiosa Temeculal] Length = 191
3418.1 Best-BlastP=> >nrprot 99% Identities = 283/283 (100%), Positives = 283/283 (100%) gb|AAC83338.1 | major outer membrane protein precursor [Legionella pneumophila] gb|AAC83342.11 major outer membrane protein precursor [Legionella pneumophila] Length = 289
3419.3 Best-BlastP=> >nrprot 36% Identities = 47/225 (20%), Positives = 93/225 (41 %), Gaps = 37/225 (16%) ref|NP_703974.1 | erythrocyte membrane protein 1 (PfEMPI) [Plasmodium falciparum 3D7] emb|CAD50587.1 | erythrocyte membrane protein 1 (PfEMPI ) [Plasmodium falciparum 3D7] Length = 2879
342.2 Best-BlastP=> >nrprot 74% Identities = 237/407 (58%), Positives = 310/407 (76%) ref|NP_924319.11 probable aminotransferase [Gloeobacter violaceus] dbj|BAC89314.11 girl 373 [Gloeobacter violaceus] Length = 417
3421.2 Best-BlastP=> >nrprot 27% Idenfities = 56/257 (21 %), Positives = 108/257 (42%), Gaps = 32/257 (12%) ref|NP_705457.11 hypothetical protein [Plasmodium falciparum 3D7] emb|CAD52694.1 | hypothetical protein [Plasmodium falciparum 3D7] Length = 1179
3423.3 Best-BlastP=> >nrprot 86% Identities = 335/366 (91 %), Positives = 346/366 (94%) gb|AAL23711.11 RalF [Legionella pneumophila] Length = 374
3426.1 Best-BlastP=> >nrprot 33% Identities = 56/142 (39%), Positives = 78/142 (54%), Gaps = 2/142 (1%) ref|NP_108258.11 hypothetical protein [Mesorhizobium loti] dbj|BAB53719.1 | hypothetical protein [Mesorhizobium loti] Length = 285
3428.1 Best-BlastP=> >nrprot 47% Identities = 72/233 (30%), Positives = 129/233 (55%), Gaps = 2/233 (0%) ref|ZP_00006345.11 COG1409: Predicted phosphohydrolases [Rhodobacter sphaeroides] Length = 273
3429.1 Best-BlastP=> >nrprot 33% Identities = 126/638 (19%), Positives = 247/638 (38%), Gaps = 85/638 (13%) gb|EAA15312.1 | hypothetical protein [Plasmodium yoelii yoelii] Length = 1627
343.1 Best-BlastP=> >nrprot 70% Identities = 78/139 (56%), Positives = 106/139 (76%) ref|NP_924320.11 hypothetical protein glr1374 [Gloeobacter violaceus] dbj|BAC89315.11 girl 374 [Gloeobacter violaceus] Length = 148
3431.3 Best-BlastP=> >nrprot 39% Identities = 109/279 (39%), Positives = 169/279 (60%), Gaps = 8/279 (2%) ref|ZP_00023400.11 COG2056: Predicted permease [Ralstonia metallidurans] Length = 325
3432.2 Best-BlastP=> >nrprot 63% Identities = 161/334 (48%), Positives = 218/334 (65%) ref|NP_346834.11 Similar to chloromuconate cycloisomerase [Clostridium acetobutylicum] pir||C96923 similar to chloromuconate cycloisomerase [imported] - Clostridium acetobutylicum gb|AAK78174.1 |AE007532_7 Similar to chloromuconate cycloisomerase [Clostridium acetobutylicum] Length = 358
3437.1 Best-BlastP=> >nrprot 32% Identities = 79/336 (23%), Positives = 125/336 (37%), Gaps = 58/336 (17%) ref|NP_523168.1 | PROBABLE ACID SPHINGOMYELINASE-LIKE PHOSPHODIESTERASE TRANSMEMBRANE PROTEIN [Ralstonia solanacearum] emb|CAD18760.11 PROBABLE ACID SPHINGOMYELINASE-LIKE PHOSPHODIESTERASE TRANSMEMBRANE PROTEIN [Ralstonia solanacearum] Length = 476
344.1 Best-BlastP=> >nrprot 67% Identities = 58/107 (54%), Posifives = 76/107 (71 %) ref|ZP_00096500.11 COG2151 : Predicted metal-sulfur cluster biosynthetic enzyme [Novosphingobium aromaticivorans] Length = 160
3442.3 Best-BlastP=> >nrprot 41 % Identities = 160/450 (35%), Positives = 247/460 (54%), Gaps = 6/450 (1 %) ref|NP_819065.11 D-alanyl-D-alanine carboxypeptidase/D-alanyl-D-alanine-endopeptidase [Coxiella burnetii RSA 493] gb|AA089579.1 | D-alanyl-D-alanine carboxypeptidase/D-alanyl-D-alanine-endopeptidase [Coxiella burnetii RSA 493] Length = 477
3443.1 Best-BlastP=> >nrprot 64% Identities = 1 12/252 (44%), Posifives = 170/252 (67%) ref|ZP_00091089.1 | COG1475: Predicted transcriptional regulators [Azotobacter vinelandii] Length = 294
3444.1 Best-BlastP=> >nrprot No Hits found
3446.3 Best-BlastP=> >nrprot 53% Identifies = 141/332 (42%), Positives = 185/332 (55%), Gaps = 16/332 (4%) ref |ZP_00140422.1 | COG1652: Uncharacterized protein containing LysM domain [Pseudomonas aeruginosa UCBPP-PA14] Length = 371
3446.1 Best-BlastP=> >nrprot 56% Identities = 158/2gi (54%), Positives = 205/291 (70%), Gaps = 3/291 (1 %) ref|NP_820973.11 DNA processing protein DprA, putative [Coxiella burnetii RSA 493] gb|AA091487.11 DNA processing protein DprA, putative [Coxiella burnetii RSA 493] Length = 308
3447.1 Best-BlastP=> >nrprot 43% Identities = 30/121 (24%), Positives = 64/121 (52%), Gaps = 5/121 (4%) ref|NP_484103.11 hypothetical protein [Nostoc sp. PCC 7120] pir||AC1814 hypothetical protein all0059 [imported] - Nostoc sp. (strain PCC 7120) dbj|BAB77δ83.11 ORF_ID:all00δ9~hypothetical protein [Nostoc sp. PCC 7120] Length = 727
3450.2 Best-BlastP=> >nrprot 73% Identities = 202/344 (58%), Positives = 264/344 (76%), Gaps = 1/344 (0%) ref|NP_469414.11 similar to E. coli Ada protein (06-methylguanine-DNA methyltransferase) [Listeria innocua] pir||AE1441 E. coli Ada protein (06-methylguanine-DNA methyltransferase) homolog Iin0068 [imported] - Listeria innocua (strain Clip11262) emb|CAC95301.11 Iin0068 [Listeria innocua] Length = 350
3451.1 Best-BlastP=> >nrprot No Hits found 3453.1. Best-BlastP=> >nrprot 10% Identities = 40/127 (31 %), Positives = 53/127 (41 %), Gaps = 19/127 (14%) ref|NP_356305.11 AGR_L_1030p [Agrobacterium tumefaciens] ref|NP_534828.1 | hypothetical protein [Agrobacterium tumefaciens str. C58 (U. Washington)] pir||AB3091 hypothetical protein Atu4350 [imported] - Agrobacterium tumefaciens (strain C58, Dupont) pir||H98195 hypothetical protein AGR_L_1030 [imported] - Agrobacterium tumefaciens (strain C58, Cereon) gb|AAK89090.1 | AGR_L_1030p [Agrobacterium tumefaciens str. C58 (Cereon)] gb|AAL45144.1 | hypothetical protein [Agrobacterium tumefaciens str. C58 (U. Washington)] Length = 278
3454.1 Best-BlastP=> >nrprot 42% Idenfities = 31/137 (22%), Positives = 62/137 (45%), Gaps = 21/137 (15%) ref|NP_832446.1 | Acetyltransferase [Bacillus cereus ATCC 14579] gb|AAP09647.11 Acetyltransferase [Bacillus cereus ATCC 14579] Length = 141
3455.3 Best-BlastP=> >nrprot No Hits found
3456.4 Best-BlastP=> >nrprot 70% Identities = 83/166 (50%), Positives = 119/166 (71 %), Gaps = 7/166 (4%) ref|ZP_00012554.11 COG3158: K+ transporter [Rhodopseudomonas palustris] Length = 620
3459.3 Best-BlastP=> >nrprot 57% Identities = 189/476 (39%), Positives = 283/475 (59%), Gaps = 7/476 (1 %) gb|AAP86869.11 putative potassium uptake protein [Ralstonia eutropha] Length = 632
346.2 Best-BlastP=> >nrprot 67% Identities = 160/326 (49%), Positives = 216/326 (66%), Gaps = 11/326 (3%) sp|Q9KNS6|SYK3_VIBCH Putative lysyl- tRNA synthetase (Lysine-tRNA ligase) (LysRS) (GX) Length = 324
3461.4 Best-BlastP=> >nrprot 18% Identities = 67/286 (23%), Positives = 116/286 (40%), Gaps = 45/286 (15%) gb|EAAig312.1 | heat shock protein hsp70 homologue Pfhsp70-3 [Plasmodium yoelii ψ yoelii] Length = 663
3463.1 Best-BlastP=> >nrprot 84% Identities = 199/269 (73%), Positives = 233/269 (86%) ref|ZP_00103515.11 COG2877: 3-deoxy-D-manno- octulosonic acid (KDO) 8-phosphate synthase [Desulfitobacterium hafniense] Length = 277
3465.2 Best-BlastP=> >nrprot 84% Identities = 371/544 (68%), Positives = 460/544 (84%), Gaps = 1/544 (0%) ref|ZP_00067534.11 COG0504: CTP synthase (UTP-ammonia lyase) [Microbulbifer degradans 2-40] Length = 643
3466.1 Best-BlastP=> >nrprot 61% Identities = 28/60 (46%), Positives = 41/60 (68%), Gaps = 2/60 (3%) ref|NP_90803δ.11 HYPOTHETICAL PROTEIN- Putative Site-specific recombinase [Wolinella succinogenes] emb|CAE10935.11 HYPOTHETICAL PROTEIN-Putative Site-specific recombinase [Wolinella succinogenes] Length = 207
3468.2 Best-BlastP=> >nrprot 78% Identities = 236/412 (57%), Positives = 320/412 (77%), Gaps = 14/412 (3%) pir||S42875 dihydrolipoamide S- succinyltransferase (EC 2.3.1.61 ) - Coxiella burnetii emb|CAA54875.11 putative dihydrolipoamide succinyltransferase [Coxiella burnefii] Length = 405
347.1 Best-BlastP=> >nrprot 60% Identities = 119/292 (40%), Positives = 182/292 (62%), Gaps = 1/292 (0%) ref|ZP_00055839.11 COG2933: Predicted SAM-dependent methyltransferase [Magnetospirillum magnetotacticum] Length = 324
3472.1 Best-BlastP=> >nrprot 27% Identities = 26/50 (52%), Positives = 28/50 (56%), Gaps = 10/50 (20%) ref|NP_773097.11 bll6457 [Bradyrhizobium japonicum] dbj|BAC61722.11 bll6457 [Bradyrhizobium japonicum USDA 110] Length = 264
3473.1 . Best-BlastP=> >nrprot No Hits found
3476.1 Best-BlastP=> >nrprot 63% Identities = 160/356 (44%), Positives = 238/356 (66%) ref|NP_819753.1 | membrane protein, putative [Coxiella burnefii RSA 493] gb|AAO90267.11 membrane protein, putative [Coxiella burnetii RSA 493] Length = 377
3477.2 Best-BlastP=> >nrprot 76% Identities = 140/241 (58%), Positives = 187/241 (77%) ref|NP_819764.11 ABC transporter, ATP-binding protein [Coxiella burnetii RSA 493] gb|AAO90268.11 ABC transporter, ATP-binding protein [Coxiella burnetii RSA 493] Length = 268
3480.4 Best-BlastP=> >nrprot 56% Identities = 185/561 (33%), Positives = 314/561 (56%), Gaps = 12/551 (2%) ref |NP_441242.11 hypothetical protein [Synechocystis sp. PCC 6803] pir||S75060 conserved hypothetical protein sll159δ - Synechocystis sp. (strain PCC 6803) dbj|BAA17922.11 ORF_ID:sll159δ~hypothetical protein [Synechocystis sp. PCC 6803] Length = 568
3481.2 Best-BlastP=> >nrprot 67% Identities = 38/89 (42%), Positives = 61/89 (68%), Gaps = 1/89 (1 %) ref|NP_442504.11 PCC7942 clock gene...ORFE [Synechocystis sp. PCC 6803] pir||S76630 hypothetical protein - Synechocystis sp. (strain PCC 6803) dbj|BAA10574.1 | sll0486 [Synechocystis sp. PCC 6803] Length = 102
3483.3 Best-BlastP=> >nrprot 52% Identities = 71/196 (36%), Positives = 106/196 (54%), Gaps = 7/196 (3%) ref |NP_216630.11 hypothetical protein Rv2114 [Mycobacterium tuberculosis H37Rv] ref|NP_855787.1 | HYPOTHETICAL PROTEIN [Mycobacterium bovis subsp. bovis AF2122/97] pir||E70612 hypothetical protein Rv2114 - Mycobacterium tuberculosis (strain H37RV) emb|CAB1070δ.11 hypothetical protein Rv2114 [Mycobacterium tuberculosis H37Rv] emb|CAD96991.11 HYPOTHETICAL PROTEIN [Mycobacterium bovis subsp. bovis AF2122/97] Length = 207
3484.1 Best-BlastP=> >nrprot No Hits found
3485.1 Best-BlastP=> >nrprot No Hits found
3486.1 Best-BlastP=> >nrprot No Hits found
3488.1 Best-BlastP=> >nrprot 16% Identities = 52/260 (20%), Positives = 116/260 (44%), Gaps = 26/260 (10%) pir||T14867 interaptin - slime mold (Dictyostelium discoideum) gb|AAC34582.1 | interaptin [Dictyostelium discoideum] Length = 1738
3489.2 Best-BlastP=> >nrprot 22% Identities = 37/178 (20%), Positives = 74/178 (41 %), Gaps = 13/178 (7%) ref|NP_624872.11 hypothetical protein SCF73.06c [Streptomyces coelicolor A3(2)] emb|CAB57411.11 hypothefical protein SCF73.06c [Streptomyces coelicolor A3(2)] Length = 333
349.2 Best-BlastP=> >nrprot 66% Identities = 109/211 (51 %), Positives = 143/211 (67%), Gaps = 1/211 (0%) ref|ZP_00013996.11 COG2872: Predicted metal-dependent hydrolases related to alanyl-tRNA synthetase HxxxH domain [Rhodospirillum rubrum] Length = 214
3494.2 Best-BlastP=> >nrprot 55% Identities = 41/85 (48%), Positives = 58/85 (68%) ref|ZP_00015213.11 COG2823: Predicted periplasmic or secreted lipoprotein [Rhodospirillum rubrum] Length = 104
3496.2 Best-BlastP=> >nrprot 30% Identities = 37/120 (30%), Positives = 50/120 (41 %), Gaps = 24/120 (20%) ref|NP_8277δδ.11 hypothetical protein [Streptomyces avermitilis MA-4680] dbj|BAC74290.1 | hypothetical protein [Streptomyces avermitilis MA-4680] Length = 574
3498.1 Best-BlastP=> >nrprot No Hits found
3499.1 Best-BlastP=> >nrprot No Hits found
35.1 Best-BlastP=> >nrprot 98% Identities = 262/265 (98%), Posifives = 262/265 (98%) emb|CAB60063.11 IvrE [Legionella pneumophila] Length = 265
350.3 Best-BlastP=> >nrprot 25% Identities = 29/124 (23%), Positives = 50/124 (40%), Gaps = 12/124 (9%) gb|AAD40638.1 |AF128842_1 extracellular calcium-sensing receptor [Mus musculus] Length = 1079
3500.3 Best-BlastP=> >nrprot 30% Identities = 61/283 (21 %), Positives = 120/283 (42%), Gaps = 16/283 (5%) ref|NP_928738.1 | hypothetical protein [Photorhabdus luminescens subsp. laumondii TT01] emb|CAE13733.1 | unnamed protein product [Photorhabdus luminescens subsp. laumondii TT01] Length = 417
3502.3 Best-BlastP=> >nrprot 55% Identifies = 57/145 (39%), Positives = 91/145 (62%), Gaps = 5/145 (3%) pir||JQ0144 probable protein-disulfide oxidoreductase (EC 1.8.4.-) - Pseudomonas aeruginosa Length = 163
3503.2 Best-BlastP=> >nrprot 74% Identities = 229/383 (59%), Positives = 287/383 (74%) ref|NP_820071.11 queuine tRNA-ribosyltransferase [Coxiella burnetii RSA 493] gb|AAO9058δ.11 queuine tRNA-ribosyltransferase [Coxiella burnetii RSA 493] Length = 386
3504.1 Best-BlastP=> >nrprot 98% Identifies = 163/164 (99%), Positives = 163/164 (99%) pir||S49314 peptidylprolyl isomerase (EC 5.2.1.8) - Legionella pneumophila emb|CAA58722.1 | cyclophilin [Legionella pneumophila] Length = 164
3506.1 Best-BlastP=> >nrprot 69% Identities = 116/212 (54%), Positives = 146/212 (68%), Gaps = 4/212 (1 %) ref|NP_245872.11 unknown [Pasteurella multocida] gb|AAK03019.11 unknown [Pasteurella multocida] Length = 226
3507.1 Best-BlastP=> >nrprot 52% Identities = 29/123 (23%), Positives = 60/123 (48%), Gaps = 8/123 (6%) gb|EAA21917.11 hypothetical protein [Plasmodium yoelii yoelii] Length = 2 6
3508.2 Best-BlastP=> >nrprot 61 % Identities = 43/94 (45%), Positives = 61/94 (64%) ref|ZP_00088933.11 COG0640: Predicted transcriptional regulators [Azotobacter vinelandii] Length = 107
3509.2 Best-BlastP=> >nrprot 15% Identities = 31/117 (26%), Positives = 55/117 (47%) ref|XP_306772.11 ENSANGP00000000282 [Anopheles gambiae] gb|EAA02010.2| ENSANGP00000000282 [Anopheles gambiae str. PEST] Length = 210
351.1 Best-BlastP=> >nrprot 59% Identities = 52 141 (36%), Positives = 89/141 (63%), Gaps = 6/141 (4%) ref|NP_562901.11 conserved hypothetical protein [Clostridium perfringens] d bj | BAB81691.11 conserved hypothetical protein [Clostridium perfringens str. 13] Length = 168
3511.2 Best-BlastP=> >nrprot 61 % Identities = 160/368 (43%), Positives = 237/368 (64%) ref|NP_902574.11 conserved hypothetical protein [Chromobacterium violaceum ATCC 12472] gb|AAQ60572.11 conserved hypothetical protein [Chromobacterium violaceum ATCC 12472] Length = 377
3612.1 Best-BlastP=> >nrprot 13% Identities = 37/145 (25%), Positives = 72/145 (49%), Gaps = 9/145 (6%) dbj|BAB62990.11 hypothetical protein [Macaca fascicularis] Length = 533
3514.1 Best-BlastP=> >nrprot No Hits found
3515.2 Best-BlastP=> >nrprot 25% Identities = 28/86 (32%), Positives = 42/86 (49%), Gaps = 4/86 (4%) ref|NP_74δ177.11 pyocin R2_PP, transcriptional repressor, CI/C2 family [Pseudomonas putida KT2440] gb|AAN68641.1 |AE016494_1 pyocin R2_PP, transcriptional repressor, CI/C2 family [Pseudomonas putida KT2440] Length = 243
3516.2 Best-BlastP=> >nrprot 53% Identifies = 25/66 (44%), Positives = 38/56 (67%) ref|NP_246251.11 unknown [Pasteurella multocida] sp|Q9CLC6|YD13_PASMU Hypothefical UPF0235 protein PM1313 gb|AAK03397.1 | unknown [Pasteurella multocida] Length = 99
3517.3 Best-BlastP=> >nrprot 56% Identities = 135/383 (35%), Positives = 219/383 (57%), Gaps = 2/383 (0%) ref|NP_820467.11 major facilitator family transporter [Coxiella burnetii RSA 493] gb|AAO90981.11 major facilitator family transporter [Coxiella burnetii RSA 493] Length = 428
3519.1 Best-BlastP=> >nrprot 31 % Identities = 80/231 (34%), Positives = 124/231 (53%), Gaps = 7/231 (3%) ref|NP_828099.11 putative glycosyl transferase [Streptomyces avermitilis MA-4680] dbj|BAC74634.1 | putative glycosyl transferase [Streptomyces avermitilis MA-4680] Length = 265
3520.1 Best-BlastP=> >nrprot 33% Identities = 90/291 (30%), Positives = 153/291 (52%), Gaps = 5/291 (1 %) ref|ZP_00056311.1 | COG1807: 4-amino-4- deoxy-L-arabinose transferase and related glycosyltransferases of PMT family [Magnetospirillum magnetotacticum] Length = 500
3521.2 Best-BlastP=> >nrprot 38% Identities = 88/281 (31 %), Positives = 147/281 (52%), Gaps = 16/281 (5%) ref|NP_561543.11 probable choline kinase [Clostridium perfringens] dbj|BAB80333.11 probable choline kinase [Clostridium perfringens str. 13] Length = 622
3522.1 Best-BlastP=> >nrprot 70% Identities = 143/261 (54%), Posifives = 180/261 (68%), Gaps = 11/261 (4%) ref|ZP_00022284.11 COG3298: Predicted 3'-5' exonuclease related to the exonuclease domain of PolB [Ralstonia metallidurans] Length = 278
3524.1 Best-BlastP=> >nrprot 67% Identities = 76/142 (53%), Positives = 97/142 (68%) ref|ZP_00065344.11 COG4764: Uncharacterized protein conserved in bacteria [Microbulbifer degradans 2-40] Length = 204
3526.1 Best-BlastP=> >nrprot 78% Identities = 137/218 (62%), Positives = 171/218 (78%) ref|NP_888560.1 | adenylate kinase [Bordetella bronchiseptica] emb|CAE32602.1 | adenylate kinase [Bordetella bronchiseptica] Length = 218
3627.1 Best-BlastP=> >nrprot 63% Idenfities = 50/115 (43%), Posifives = 77/11 δ (66%) ref|ZP_00042420.11 COG0526: Thiol-disulfide isomerase and thioredoxins [Magnetococcus sp. MC-1] Length = 124
3530.1 Best-BlastP=> >nrprot No Hits found 3532.1 Best-BlastP=> >nrprot 25% Identities = 141/689 (20%), Positives = 276/689 (40%), Gaps = 124/689 (17%) ref|NP_048227.1 | ORF MSV156 hypothetical protein [Melanoplus sanguinipes entomopoxvirus] pir||T28317 ORF MSV156 hypothetical protein - Melanoplus sanguinipes entomopoxvirus gb|AAC97677.1 | ORF MSV156 hypothetical protein [Melanoplus sanguinipes entomopoxvirus] Length = 1127
3554.4 Best-BlastP=> >nrprot 64% Identifies = 92/193 (47%), Positives = 129/193 (66%), Gaps = 5/193 (2%) ref| protein PilO [Pseudomonas syringae pv. tomato str. DC3000] gb|AA058567.11 type IV pilus biogene syringae pv. tomato str. DC3000] Length = 207
3565.2 Best-BlastP=> >nrprot 60% Identities = 72/175 (41 %), Positives = 111/175 (63%) gb|AAA93087.1 | membr
3566.2 Best-BlastP=> >nrprot 44% Identities = 36/118 (29%), Positives = 63/118 (53%), Gaps = 1/118 (0%) ref |N [Coxiella burnetii RSA 493] gb|AAO90641.11 NifU family protein [Coxiella burnetii RSA 493] Length =
3557.1 Best-BlastP=> >nrprot 78% Identities = 236/383 (61 %), Positives = 303/383 (79%) ref|NP_717860.11 cyst oneidensis MR-1 ] gb|AAN55304.1 |AE016668_5 cysteine desulfurase [Shewanella oneidensis MR-1 ]
3658.1 Best-BlastP=> >nrprot No Hits found
3559.2 Best-BlastP=> >nrprot No Hits found
3561.2 Best-BlastP=> >nrprot 99% Identities = 782/783 (99%), Positives = 783/783 (100%) pir||T18329 icmO prot emb|CAA73241.1 | IcmO protein [Legionella pneumophila] gb|AAC38193.1 | DotL [Legionella pneumophila] [Legionella pneumophila] Length = 783
3562.3 Best-BlastP=> >nrprot 99% Identities = 374/376 (99%), Positives = 376/376 (100%) pir||T18328 icmP prot emb|CAA73240.1 | IcmP protein [Legionella pneumophila] gb|AAC38194.1 | DotM [Legionella pneumophila] [Legionella pneumophila] Length = 376
3563.2 Best-BlastP=> >nrprot 47% Identities = 35/90 (38%), Positives = 55/90 (61 %), Gaps = 5/90 (5%) dbj|BAC vulnificus YJ016] Length = 343
3564.2 Best-BlastP=> >nrprot No Hits found
3566.1 Best-BlastP=> >nrprot 76% Identities = 246/389 (63%), Positives = 305/389 (78%) ref|NP_716561.11 phos oneidensis MR-1] sp|Q8EIB1 |PGK_SHEON Phosphoglycerate kinase gb|AAN54006.1 |AE015538_3 phosp oneidensis MR-1] Length = 391
3568.3 Best-BlastP=> >nrprot 66% Identities = 231/477 (48%), Positives = 318/477 (66%), Gaps = 6/477 (1 %) ref [Pseudomonas putida KT2440] gb|AAN66985.1 |AE016326_11 pyruvate kinase II [Pseudomonas putida KT
357.2 Best-BlastP=> >nrprot 75% Identities = 138/234 (58%), Positives = 177/234 (75%) ref|NP_540δ13.11 LrgB pir||AF3461 IrgB protein [imported] - Brucella melitensis (strain 16M) gb|AAL52777.1 | murein hydrolase exp Length = 235
3570.2 Best-BlastP=> >nrprot 67% Identities = 33/70 (47%), Positives = 48/70 (68%) ref|NP_902003.11 conserve [Chromobacterium violaceum ATCC 12472] gb|AAQ60005.1 | conserved hypothetical protein [Chrom 12472] Length = 75
3572.1 Best-BlastP=> >nrprot 88% Identities = 109/139 (78%), Positives = 124/139 (89%) ref |ZP_00012735.1 | C protein [Rhodopseudomonas palustris] Length = 139
3574.1 Best-BlastP=> >nrprot 40% Identities = 83/316 (26%), Positives = 130/316 (41 %), Gaps = 44/316 (13%) r hypothetical protein [Pseudomonas syringae pv. tomato str. DC3000] gb|AA054384.1 | conserved h syringae pv. tomato str. DC3000] Length = 312
3575.2 Best-BlastP=> >nrprot 42% Identifies = 112/408 (27%), Positives = 192/408 (47%), Gaps = 20/408 (4%) r hypothetical protein [Xanthomonas campestris pv. campestris str. ATCC 33913] gb|AAM42387.1 | co
[Xanthomonas campestris pv. campestris str. ATCC 33913] Length = 414
3578.2 Best-BlastP=> >nrprot 91 % Identities = 465/524 (88%), Positives = 480/524 (91 %), Gaps = 6/524 (1 %) gb pneumophila] Length = 518
3581.1 Best-BlastP=> >nrprot 64% Identities = 148/160 (92%), Positives = 153/160 (95%) pir||S61389 small basi pneumophila gb|AAA79903.1 | SbpA Length = 161
3582.2 Best-BlastP=> >nrprot 64% Idenfities = 134/282 (47%), Positives = 186/282 (65%), Gaps = 1/282 (0%) ref [Pseudomonas syringae pv. syringae B728a] Length = 288
3584.1 Best-BlastP=> >nrprot 60% Identities = 26/60 (43%), Positives = 39/60 (65%), Gaps = 2/60 (3%) ref|NP_8 [Nitrosomonas europaea ATCC 19718] ref|NP_841665.11 possible transposase [Nitrosomonas europaea A possible transposase [Nitrosomonas europaea ATCC 19718] emb|CAD85541.1 | possible transposase [Nitr Length = 136
3586.2 Best-BlastP=> >nrprot 67% Identities = 186/367 (50%), Positives = 251/367 (68%) ref|ZP_00012g35.1 | C protein [Rhodopseudomonas palustris] Length = 367
3588.2 Best-BlastP=> >nrprot 69% Identities = 83/161 (51 %), Positives = 115/161 (71 %) ref|NP_531634.1 | meth tumefaciens str. Cδ8 (U. Washington)] pir||AH2691 methyltransferase Atu0936 [imported] - Agrobact Dupont) gb|AAL4i δ0.11 methyltransferase [Agrobacterium tumefaciens str. C58 (U. Washington)]
369.2 Best-BlastP=> >nrprot 41 % Identities = 67/120 (56%), Positives = 85/120 (70%) ref|NP_697359.11 conser 1330] gb|AAN2g274.1 |AE014344_9 conserved hypothetical protein [Brucella suis 1330] Length = 13
3591.2 Best-BlastP=> >nrprot 65% Identities = 83/182 (45%), Positives = 124/182 (68%), Gaps = 1/182 (0%) ref| transcriptional regulator [Microbulbifer degradans 2-40] Length = 203
3592.1 Best-BlastP=> >nrprot 65% Identities = 73/134 (54%), Positives = 94/134 (70%) gb|AAA69116.11 ORF_o1 Length = 180
3593.1 Best-BlastP=> >nrprot 62% Identities = 133/297 (44%), Positives = 193/297 (64%), Gaps = 10/297 (3%) r carbamoyltransferase [Coxiella burnetii RSA 493] gb|AA091579.11 aspartate carbamoyltransferase [Coxiell 310
3594.1 Best-BlastP=> >nrprot No Hits found
3596.1 Best-BlastP=> >nrprot 64% Identities = 39/75 (52%), Positives = 56/75 (74%) ref|NP_820996.11 conserve RSA 493] gb|AAO91510.11 conserved hypothetical protein [Coxiella burnetii RSA 493] Length = 81
3596.1 Best-BlastP=> >nrprot 79% Identities = 82/112 (73%), Positives = 99/112 (88%) ref|ZP_00025955.11 CO [Ralstonia metallidurans] Length = 112
3597.3 Best-BlastP=> >nrprot 56% Identities = 73/189 (38%), Positives = 109/189 (57%), Gaps = 7/189 (3%) ref|NP_819119.11 5-formyltetrahydrofolate cyclo-ligase family protein [Coxiella burnetii RSA 493] gb|AA089633.11 5-formyltetrahydrofolate cyclo-ligase family protein [Coxiella burnetii RSA 493] Length = 197
3598.3 Best-BlastP=> >nrprot No Hits found
3599.4 Best-BlastP=> >nrprot 48% Identities = 78/229 (34%), Positives = 131/229 (57%), Gaps = 3/229 (1 %) ref|NP_819456.11 4-amino-4- deoxychorismate lyase, putative [Coxiella burnetii RSA 493] gb|AAO89970.1 | 4-amino-4-deoxychorismate lyase, putative [Coxiella burnetii RSA 493] Length = 281
36.1 Best-BlastP=> >nrprot 98% Identities = 606/613 (98%), Positives = 607/613 (99%) emb|CAB60062.1 | lvhD4 [Legionella pneumophila] Length = 691 360.3 Best-BlastP=> >nrprot 7% Identities = 39/94 (41 %), Positives = 47/94 (50%), Gaps = 1/94 (1 %) ref|NP_900714.11 hypothetical protein CV1044 [Chromobacterium violaceum ATCC 12472] gb|AAQ68719.1 | hypothetical protein CV1044 [Chromobacterium violaceum ATCC 12472] Length = 499 3600.2 Best-BlastP=> >nrprot 66% Identities = 147/311 (47%), Positives = 207/311 (66%), Gaps = 1/311 (0%) ref|NP_2325δ9.11 conserved hypothetical protein [Vibrio cholerae 01 biovar eltor str. N16961] pir||H82491 conserved hypothetical protein VCA0159 [imported] - Vibrio cholerae (strain N16961 serogroup OI ) gb|AAF96072.1 | conserved hypothetical protein [Vibrio cholerae 01 biovar eltor str. N16961] Length = 313
3601.1 Best-BlastP=> >nrprot 68% Identities = 183/386 (47%), Positives = 262/385 (68%), Gaps = 8/385 (2%) ref|ZP_00065865.11 COG3004: Na+/H+ antiporter [Microbulbifer degradans 2-40] Length = 401
3602.2 Best-BlastP=> >nrprot 20% Identities = 73/282 (25%), Positives = 120/282 (42%), Gaps = 50/282 (17%) gb|EAA19568.11 hypothetical protein [Plasmodium yoelii yoelii] Length = 5074
3607.2 Best-BlastP=> >nrprot 56% Identifies = 158/379 (41 %), Positives = 224/379 (59%), Gaps = 12/379 (3%) ref|ZP_00128328.11 COG2850: Uncharacterized conserved protein [Pseudomonas syringae pv. syringae B728a] Length = 388
3608.1 Best-BlastP=> >nrprot No Hits found
3609.1 Best-BlastP=> >nrprot 99% Identities = 191/192 (99%), Positives = 192/192 (100%) sp|P31108|SODF_LEGPN Superoxide dismutase [Fe] pir||JS0749 superoxide dismutase (EC 1.15.1.1 ) (Fe) - Legionella pneumophila dbj|BAA02306.1 | iron superoxide dismutase (Fe-SOD) [Legionella pneumophila] gb|AAM00603.1 | superoxide dismutase [Legionella pneumophila] prf||2014300A Fe superoxide dismutase Length = i 2
3610.1 Best-BlastP=> >nrprot 72% Identities = 202/384 (52%), Positives = 284/384 (73%) ref |NP_841480.11 argD; acetylomithine aminotransferase [Nitrosomonas europaea ATCC 19718] emb|CAD85350.1 | argD; acetylomithine aminotransferase [Nitrosomonas europaea ATCC 19718] Length = 393
3611.2 Best-BlastP=> >nrprot 66% Idenfities = 119/254 (46%), Positives = 169/254 (66%), Gaps = 2/254 (0%) ref|NP_440288.11 unknown protein [Synechocystis sp. PCC 6803] pir||S74928 hypothetical protein sll0647 - Synechocystis sp. (strain PCC 6803) dbj|BAA16968.11 ORF_ID:sll0647~unknown protein [Synechocystis sp. PCC 6803] Length = 256
3612.2 Best-BlastP=> >nrprot 40% Identities = 68/222 (30%), Positives = 106/222 (47%), Gaps = 16/222 (7%) ref|NP_229421.11 hypothetical protein [Thermotoga maritima] pir||G72232 hypothetical protein - Thermotoga maritima (strain MSB8) gb|AAD36688.1 |AE001805_13 hypothetical protein [Thermotoga maritima] Length = 222
3613.3 Best-BlastP=> >nrprot 76% Identities = 246/422 (68%), Positives = 326/422 (77%), Gaps = 2/422 (0%) ref|NP_40δ097.11 NAD-dependent malic enzyme [Yersinia pestis] ref|NP_669960.11 NAD-linked malate dehydrogenase (malic enzyme) [Yersinia pestis KIM] pir||AC0184 malate dehydrogenase (oxaloacetate-decarboxylating) (EC 1.1.1.38) [imported] - Yersinia pestis (strain C092) emb|CAC90334.11 NAD-dependent malic enzyme [Yersinia pesfis C092] gb|AAM86211.1 |AE013868_5 NAD-linked malate dehydrogenase (malic enzyme) [Yersinia pestis KIM] Length = 665
3614.3 Best-BlastP=> >nrprot 99% Identities = 353/354 (99%), Positives = 354/354 (100%) gb|AAM00637.11 putative cobalt/magnesium uptake transporter [Legionella pneumophila] Length = 354
3617.2 Best-BlastP=> >nrprot 83% Identities = 68/101 (67%), Positives = 85/101 (84%) rβf|NP_820421.1 | NADH dehydrogenase I, K subunit [Coxiella burnetii RSA 493] gb|AAO90935.11 NADH dehydrogenase I, K subunit [Coxiella burnetii RSA 4g3] Length = 101
3618.2 Best-BlastP=> >nrprot 72% Identities = 388/654 (59%), Positives = 480/654 (73%), Gaps = 4/654 (0%) ref|NP_820420.11 NADH dehydrogenase ■ I, L subunit [Coxiella burnetii RSA 493] gb|AAO90934.11 NADH dehydrogenase I, L subunit [Coxiella burnetii RSA 493] Length = 653
3619.1 Best-BlastP=> >nrprot 70% Identities = 63/137 (45%), Positives = 100/137 (72%) ref|NP_253434.1 | conserved hypothetical protein [Pseudomonas aeruginosa PA01] sp|Q9HV53|YBE6_PSEAE Hypothetical UPF0090 protein PA4746 pir||A83053 conserved hypothetical protein PA4746 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG08132.1 |AE004888_7 conserved hypothetical protein [Pseudomonas aeruginosa PA01] Length = 152
3621.2 Best-BlastP=> >nrprot 78% Identities = 307/4gi (62%), Positives = 385/491 (78%), Gaps = 2/491 (0%) ref|ZP_00066603.11 COG0195: Transcription elongation factor [Microbulbifer degradans 2-40] Length = 493
3622.3 Best-BlastP=> >nrprot 71 % Identities = 168/313 (53%), Positives = 226/313 (72%), Gaps = 1/313 (0%) ref|ZP_00092328.11 hypothetical protein [Azotobacter vinelandii] Length = 379
3623.2 Best-BlastP=> >nrprot 73% Identities = 74/123 (60%), Positives = 92/123 (74%), Gaps = 1/123 (0%) ref|NP_716411.11 glycine cleavage system H protein [Shewanella oneidensis MR-1] gb|AAN53856.1 |AE015522_11 glycine cleavage system H protein [Shewanella oneidensis MR-1] Length = 129 3625.2 Best-BlastP=> >nrprot 73% Identities = 270/452 (59%), Positives = 338/452 (74%), Gaps = 2/452 (0%) ref|NP_840693.11 Glycine cleavage system P-protein [Nitrosomonas europaea ATCC 19718] emb|CAD84520.11 Glycine cleavage system P-protein [Nitrosomonas europaea ATCC 19718] Length = 453
3626.1 Best-BlastP=> >nrprot 59% Identities = 44/90 (48%), Positives = 61/90 (67%) ref|NP_713147.11 conserved hypothetical protein [Leptospira interrogans serovar lai str. 56601] gb|AAN50165.1 |AE011460_7 conserved hypothetical protein [Leptospira interrogans serovar lai . str. 56601] Length = 104
3629.2 Best-BlastP=> >nrprot 75% Identities = 170/291 (58%), Positives = 223/291 (76%) ref|ZP_00065024.1 | COG1159: GTPase [Microbulbifer degradans 2-40] Length = 298
3631.4 Best-BlastP=> >nrprot No Hits found
3632.3 Best-BlastP=> >nrprot 49% Idenfities = 273/1022 (26%), Positives = 507/1022 (49%), Gaps = 42/1022 (4%) gb|AAP44228.11 transposase TnpA [Pseudomonas sp. ND6] Length = 1009
3633.3 Best-BlastP=> >nrprot 63% Identities = 122/307 (39%), Positives = 196/307 (63%) ref|NP_819755.11 conserved hypothetical protein [Coxiella burnetii RSA 493] gb|AAO90269.11 conserved hypothetical protein [Coxiella burnetii RSA 493] Length = 308
3634.1 Best-BlastP=> >nrprot 56% Idenfities = 103/103 (100%), Positives = 103/103 (100%) sp|P26880|YPA1_LEGPN Hypothetical protein in PAL δ'region (ORFU) Length = 103
3636.1 Best-BlastP=> >nrprot 99% Identities = 176/176 (100%), Positives = 176/176 (100%) sp|P26493|PAL_LEGPN Peptidoglycan-associated lipoprotein precursor (19 kDa surface antigen) (PPL) pir||A60337 outer membrane protein pplA, peptidoglycan-associated, precursor - Legionella pneumophila emb|CAA43033.1 | lipoprotein antigen [Legionella pneumophila] Length = 176
3637.1 Best-BlastP=> >nrprot 69% Identities = 188/201 (93%), Positives = 193/201 (96%), Gaps = 4/201 (1 %) sp|P26881 |YPA2_LEGPN Hypothefical protein in PAL 3'region (ORFD) Length = 201
3638.2 Best-BlastP=> >nrprot 78% Identities = 141/217 (64%), Positives = 171/217 (78%), Gaps = 1/217 (0%) ref|ZP_0008381 1 .1 | COG0602: Organic radical activating enzymes [Pseudomonas fluorescens PfO-1] Length = 218
364.2 Best-BlastP=> >nrprot 5% Identities = 31/70 (44%), Positives = 38/70 (64%), Gaps = 3/70 (4%) ref|NP_900714.11 hypothetical protein CV1044 [Chromobacterium violaceum ATCC 12472] gb|AAQ68719.1 | hypothetical protein CV1044 [Chromobacterium violaceum ATCC 12472] Length = 499
3640.2 Best-BlastP=> >nrprot 61 % Identities = 94/313 (30%), Positives = 162/313 (51 %), Gaps = 21/313 (6%) emb|CAC51371 .1 | mevalonate diphosphate decarboxylase [Lactobacillus helveticus] Length = 320
3641.1 Best-BlastP=> >nrprot 42% Identities = 65/228 (28%), Positives = 108/228 (47%), Gaps = 12/228 (5%) ref|NP_820494.11 transporter, ZIP family [Coxiella burnetii RSA 493] gb|AA091008.1 1 transporter, ZIP family [Coxiella burnetii RSA 493] Length = 261
3642.2 Best-BlastP=> >nrprot 57% Identities = 154/329 (46%), Positives = 220/329 (66%), Gaps = 3/329 (0%) ref|NP_820351.11 cation-efflux family protein [Coxiella burnefii RSA 493] gb|AAO90865.11 cation-efflux family protein [Coxiella burnetii RSA 493] Length = 378
3645.3 Best-BlastP=> >nrprot 67% Identities = 132/255 (51 %), Positives = 171/255 (67%), Gaps = 15/255 (5%) ref|ZP_00016164.1 | COG0378: Ni2+- binding GTPase involved in regulation of expression and maturation of urease and hydrogenase [Rhodospirillum rubrum] Length = 268
3647.2 Best-BlastP=> >nrprot 71 % Identities = 37/67 (56%), Positives = 64/67 (80%) ref |ZP_00021586.11 COG0298: Hydrogenase maturation factor [Ralstonia metallidurans] Length = 103
365.4 Best-BlastP=> >nrprot 49% Identities = 88/327 (26%), Positives = 165/327 (50%), Gaps = 15/327 (4%) ref|NP_819818.1 | multidrug resistance protein [Coxiella burnetii RSA 493] gb|AAO90332.11 multidrug resistance protein [Coxiella burnetii RSA 4g3] Length = 331
3650.1 Best-BlastP=> >nrprot 52% Identities = 171/363 (47%), Positives = 221/363 (60%), Gaps = 8/363 (2%) ref|ZP_00089787.11 COG1 145: Ferredoxin [Azotobacter vinelandii] Length = 382
3651.1 Best-BlastP=> >nrprot 56% Identities = 114/269 (42%), Positives = 158/269 (58%), Gaps = 3/269 (1 %) ref|ZP_00089785.1 1 COG0543: 2- polyprenylphenol hydroxylase and related flavodoxin oxidoreductases [Azotobacter vinelandii] Length = 283
3652.2 Best-BlastP=> >nrprot 64% Identifies = 132/249 (53%), Positives = 169/249 (67%), Gaps = 1/249 (0%) ref|ZP_00089784.11 COG1941 : Coenzyme F420-reducing hydrogenase, gamma subunit [Azotobacter vinelandii] Length = 256
3653.3 Best-BlastP=> >nrprot 56% Identities = 26/40 (65%), Positives = 33/40 (82%), Gaps = 3/40 (7%) gb|AAG45149.11 TraA-like protein [Legionella pneumophila] Length = 883
3655.1 Best-BlastP=> >nrprot No Hits found
3657.1 Best-BlastP=> >nrprot 48% Identities = 65/171 (38%), Positives = 103/171 (60%), Gaps = 2/171 (1%) ref|NP_174343.2| expressed protein [Arabidopsis thaliana] Length = 391
3658.1 Best-BlastP=> >nrprot No Hits found
3659.2 Best-BlastP=> >nrprot No Hits found
366.2 Best-BlastP=> >nrprot No Hits found
3661.4 Best-BlastP=> >nrprot 32% Identities = 71/191 (37%), Positives = 108/191 (56%), Gaps = 22/191 (11 %) ref |NP_901795.11 hypothetical protein CV2125 [Chromobacterium violaceum ATCC 12472] gb|AAQ69798.1 | hypothetical protein CV2126 [Chromobacterium violaceum ATCC 12472] Length = 202 3663.1 Best-BlastP=> >nrprot No Hits found 3664.1 Best-BlastP=> >nrprot 52% Identities = 142/332 (42%), Positives = 203/332 (61 %), Gaps = 5/332 (1 %) gb|AAK49795.11 WcbB [Burkholderia pseudomallei] Length = 365
3665.1 Best-BlastP=> >nrprot 52% Identities = 109/243 (44%), Positives = 162/243 (66%) ref|ZP_00125435.11 COG0500: SAM-dependent methyltransferases [Pseudomonas syringae pv. syringae B728a] Length = 646
3666.2 Best-BlastP=> >nrprot 44% Identities = 121/287 (42%), Positives = 173/287 (60%), Gaps = 2/287 (0%) gb|AAK49795.1 | WcbB [Burkholderia pseudomallei] Length = 365
3667.1 Best-BlastP=> >nrprot No Hits found 3669.1 Best-BlastP=> >nrprot 83% Idenfities = 227/316 (71 %), Positives = 264/316 (83%) ref|NP_820997.11 cystathionine beta-synthase, putative [Coxiella burnetii RSA 493] gb|AA091511.11 cystathionine beta-synthase, putative [Coxiella burnetii RSA 493] Length = 316
3670.2 Best-BlastP=> >nrprot No Hits found
3671.2 Best-BlastP=> >nrprot 24% Identities = 34/134 (25%), Positives = 59/134 (44%), Gaps = 7/134 (5%) ref|XP_283791.11 RIKEN cDNA B230210E21 gene [Mus musculus] Length = 484
3673.2 Best-BlastP=> >nrprot 51 % Identities = 137/403 (33%), Posifives = 211/403 (52%), Gaps = 9/403 (2%) ref|NP_793575.11 major facilitator family transporter [Pseudomonas syringae pv. tomato str. DC3000] gb|AAO57270.11 major facilitator family transporter [Pseudomonas syringae pv. tomato str. DC3000] Length = 445
3674.2 Best-BlastP=> >nrprot 63% Identities = 123/273 (45%), Positives = 173/273 (63%), Gaps = 17/273 (6%) ref|ZP_00084952.11 COG1766: Flagellar biosynthesis/type III secretory pathway lipoprotein [Pseudomonas fluorescens PfO-1] Length = 596
3675.1 Best-BlastP=> >nrprot 60% Identities = 45/87 (51 %), Positives = 64/87 (73%), Gaps = 5/87 (5%) dbj|BAC95246.11 flagellar hook-basal body protein FliE [Vibrio vulnificus YJ016] Length = 122
3676.2 Best-BlastP=> >nrprot 60% Identities = 261/267 (97%), Positives = 263/267 (98%) gb|AAG45148.11 putative response regulator [Legionella pneumophila] Length = 267
3679.3 Best-BlastP=> >nrprot 56% Identities = 129/324 (39%), Positives = 194/324 (59%), Gaps = 5/324 (1 %) ref|NP_746486.11 sensory box histidine kinase FleS [Pseudomonas putida KT2440] gb|AAN69950.1 |AE016633_8 sensory box histidine kinase FleS [Pseudomonas putida KT2440] Length = 405 368.2 Best-BlastP=> >nrprot 22% Identities = 123/487 (25%), Positives = 188/487 (38%), Gaps = 62/487 (12%) emb|CAD27470.11 SPAPB18E9.04c [Schizosaccharomyces pombe] Length = 800
3701.1 Best-BlastP=> >nrprot 31 % Identities = 57/156 (36%), Positives = 84/156 (53%), Gaps = 8/156 (5%) ref|NP_522544.11 PROBABLE TRANSCRIPTION REGULATOR PROTEIN [Ralstonia solanacearum] emb|CAD18134.1 | PROBABLE TRANSCRIPTION REGULATOR PROTEIN [Ralstonia solanacearum] Length = 227
3702.1 Best-BlastP=> >nrprot 69% Identities = 131/242 (54%), Positives = 172/242 (71 %), Gaps = 4/242 (1 %) ref|NP_284288.11 putative pseudouridine synthase [Neisseria meningitidis Z2491] pir||H81849 probable pseudouridine synthase NMA1573 [imported] - Neisseria meningitidis (strain Z2491 serogroup A) emb|CAB84800.11 putative pseudouridine synthase [Neisseria meningitidis Z2491] Length = 256
3703.1 Best-BlastP=> >nrprot 67% Identities = 90/173 (52%), Positives = 130/173 (75%) ref|ZP_00127917.1 1 COG1386: Predicted transcriptional regulator containing the HTH domain [Pseudomonas syringae pv. syringae B728a] Length = 255
3705.1 Best-BlastP=> >nrprot 70% Identities = 140/260 (56%), Positives = 186/250 (74%) ref|NP_642633.11 conserved hypothetical protein [Xanthomonas axonopodis pv. citri str. 306] gb|AAM37169.1 1 conserved hypothetical protein [Xanthomonas axonopodis pv. citri str. 306] Length = 279
3706.2 Best-BlastP=> >nrprot 77% Identities = 254/396 (64%), Positives = 314/396 (79%) ref|NP_519264.11 PROBABLE TRYPTOPHANYL-TRNA SYNTHETASE (SYW PROTEIN) [Ralstonia solanacearum] sp|Q8Y0A1 |SYW_RALSO Tryptophanyl-tRNA synthetase (Tryptophan-tRNA ligase) (TrpRS) emb|CAD14845.11 PROBABLE TRYPTOPHANYL-TRNA SYNTHETASE (SYW PROTEIN) [Ralstonia solanacearum] Length = 400
3708.1 Best-BlastP=> >nrprot 75% Identities = 130/206 (63%), Positives = 167/205 (81 %) ref|ZP_00067870.11 COG0009: Putative translation factor (SUA5) [Microbulbifer degradans 2-40] Length = 207
3709.1 Best-BlastP=> >nrprot 58% Idenfities = 1 19/275 (43%), Positives = 165/275 (60%), Gaps = 2/275 (0%) ref |NP_841756.11 PHP domain N- terminal region:PHP domain C-terminal region [Nitrosomonas europaea ATCC 19718] emb|CAD85635.11 PHP domain N-terminal region:PHP domain C-terminal region [Nitrosomonas europaea ATCC 19718] Length = 316
371.1 Best-BlastP=> >nrprot No Hits found 3710.1 Best-BlastP=> >nrprot 67% Identities = 152/329 (46%), Positives = 214/329 (65%), Gaps = 25/329 (7%) ref|NP_245983.1 | PhoH [Pasteurella multocida] gb|AAK03130.1 | PhoH [Pasteurella multocida] Length = 372
371 1.1 Best-BlastP=> >nrprot 66% Identities = 78/152 (51 %), Positives = 105/152 (69%), Gaps = 2/152 (1 %) ref|NP_746893.11 conserved hypothetical protein TIGR00043 [Pseudomonas putida KT2440] gb|AAN70357.1 |AE016677_8 conserved hypothetical protein TIGR00043 [Pseudomonas putida KT2440] Length = 157
3712.2 Best-BlastP=> >nrprot 65% Identities = 139/261 (53%), Positives = 189/261 (72%) dbj|BAC93678.11 putative hemolysin [Vibrio vulnificus YJ016] Length = 306
3714.3 Best-BlastP=> >nrprot No Hits found 3720.2 Best-BlastP=> >nrprot 86% Identities = 192/264 (72%), Positives = 228/264 (86%) ref|ZP_00024661.11 COG0207: Thymidylate synthase [Ralstonia metallidurans] Length = 264
3721.1 Best-BlastP=> >nrprot 64% Idenfities = 79/143 (55%), Positives = 101/143 (70%) ref |NP_819678.11 riboflavin synthase, beta subunit [Coxiella burnetii RSA 493] gb|AAO90192.1 | riboflavin synthase, beta subunit [Coxiella burnetii RSA 493] Length = 151
3729.1 Best-BlastP=> >nrprot 53% Identities = 70/188 (37%), Positives = 101/188 (53%), Gaps = 14/188 (7%) ref|NP_636438.11 conserved hypothetical protein [Xanthomonas campestris pv. campestris str. ATCC 33913] gb|AAM40362.1 | conserved hypothetical protein [Xanthomonas campestris pv. campestris str. ATCC 33913] Length = 176
3730.1 Best-BlastP=> >nrprot No Hits found
3731.1 Best-BlastP=> >nrprot 63% Identities = 56/114 (49%), Positives = 77/114 (67%), Gaps = 1/114 (0%) ref|NP_890078.1 | phage integrase
[Bordetella bronchiseptica] emb|CAE34037.1 | phage integrase [Bordetella bronchiseptica] Length = 407
3732.1 Best-BlastP=> >nrprot 36% Identities = 58/180 (32%), Positives = 107/180 (59%), Gaps = 1/180 (0%) ref|NP_624053.11 predicted transposase [Thermoanaerobacter tengcongensis] gb|AAM25657.11 predicted transposase [Thermoanaerobacter tengcongensis] Length = 267
3734.1 Best-BlastP=> >nrprot 37% Identities = 86/323 (26%), Positives = 140/323 (43%), Gaps = 53/323 (16%) ref|NP_563745.11 expressed protein [Arabidopsis thaliana] gb|AAM65464.1 | unknown [Arabidopsis thaliana] gb|AAN72060.1 | expressed protein [Arabidopsis thaliana] gb| AAP42733.11 At1 g05620 [Arabidopsis thaliana] Length = 322
3735.2 Best-BlastP=> >nrprot 36% Identities = 106/496 (21 %), Positives = 199/496 (40%), Gaps = 61/496 (12%) gb|EAA16521.1 | 235 kDa rhoptry protein [Plasmodium yoelii yoelii] Length = 2740
3737.1 Best-BlastP=> >nrprot No Hits found
3739.3 Best-BlastP=> >nrprot 79% Identities = 100/146 (68%), Positives = 117/146 (80%) ref|NP_539647.1 | LACTOYLGLUTATHIONE LYASE [Brucella melitensis] pir||AD3343 lactoylglutathione lyase (EC 4.4.1.5) [imported] - Brucella melitensis (strain 16M) gb|AAL51911.1 | LACTOYLGLUTATHIONE LYASE [Brucella melitensis 16M] Length = 173
374.2 Best-BlastP=> >nrprot 66% Identities = 302/5g5 (50%), Positives = 393/595 (66%), Gaps = 13/595 (2%) ref|ZP_00029131.11 COG3243: Poly(3- hydroxyalkanoate) synthetase [Burkholderia fungorum] Length = 642
3740.3 Best-BlastP=> >nrprot 72% Identities = 95/159 (59%), Positives = 128/159 (80%) ref|ZP_00021733.11 COG2862: Predicted membrane protein [Ralstonia metallidurans] Length = 206
3741.1 Best-BlastP=> >nrprot 32% Identities = 61/271 (22%), Positives = 133/271 (49%), Gaps = 18/271 (6%) ref|XP_230851.2| similar to hypothetical protein [Rattus norvegicus] Length = 396
3742.1 Best-BlastP=> >nrprot No Hits found
3743.1 Best-BlastP=> >nrprot No Hits found
3744.2 Best-BlastP=> >nrprot No Hits found 3747.2 Best-BlastP=> >nrprot 49% Identities = 99/264 (37%), Positives = 149/264 (56%), Gaps = 7/264 (2%) ref|NP_743187.11 peptidase, M23/M37 family [Pseudomonas putida KT2440] gb|AAN66651.1 |AE016293_1 peptidase, M23/M37 family [Pseudomonas putida KT2440] Length = 275
3748.1 Best-BlastP=> >nrprot 63% Identities = 192/445 (43%), Positives = 280/445 (62%), Gaps = 5/445 (1 %) ref|NP_461447.11 exonuclease VII, large subunit [Salmonella typhimurium LT2] sp|Q8ZN58|EX7L_SALTY Probable exodeoxyribonuclease VII large subunit (Exonuclease VII large subunit) gb|AAL21406.11 exonuclease VII, large subunit [Salmonella typhimurium LT2] Length = 449
3750.2 Best-BlastP=> >nrprot 42% Identities = 156/424 (36%), Positives = 245/424 (57%), Gaps = 2/424 (0%) pir||S27611 agglutination protein Pseudomonas putida gb|AAA25695.1 | agglutination protein Length = 462
3752.2 Best-BlastP=> >nrprot 54% Identities = 82/187 (43%), Posifives = 116/187 (62%), Gaps = 2/187 (1 %) ref |ZP_00065146.11 COG3672: Predicted periplasmic protein [Microbulbifer degradans 2-40] Length = 241
3753.2 Best-BlastP=> >nrprot 56% Identities = 197/655 (30%), Positives = 352/656 (53%), Gaps = 41/655 (6%) ref|ZP_00086698.11 COG2200: FOG: EAL domain [Pseudomonas fluorescens PfO-1] Length = 648
3754.2 Best-BlastP=> >nrprot 36% Identities = 48/170 (28%), Positives = 88/170 (51 %), Gaps = 11/170 (6%) ref|NP_773217.1 | bll6577 [Bradyrhizobium japonicum] dbj|BAC51842.1 | bll6577 [Bradyrhizobium japonicum USDA 110] Length = 237
3756.2 Best-BlastP=> >nrprot 67% Identities = 234/439 (53%), Positives = 301/439 (68%), Gaps = 7/439 (1 %) ref|NP_713336.11 putative flavin- containing monooxygenase [Leptospira interrogans serovar lai str. 56601] gb|AAN50354.1 |AE011478_5 putative flavin-containing monooxygenase [Leptospira interrogans serovar lai str. 66601] Length = 468
376.3 Best-BlastP=> >nrprot 69% Identities = 1 δδ/329 (47%), Positives = 230/329 (69%), Gaps = 4/329 (1 %) ref|NP_931992.11 glycerol-3-phosphate dehydrogenase [NAD(P)+] (NAD(P)H-dependent glycerol-3-phosphate dehydrogenase) [Photorhabdus luminescens subsp. laumondii TTO1] emb|CAE17210.1 | glycerol-3-phosphate dehydrogenase [NAD(P)+] (NAD(P)H-dependent glycerol-3-phosphate dehydrogenase) [Photorhabdus luminescens subsp. laumondii TT01] Length = 340
3760.3 Best-BlastP=> >nrprot 69% Identities = 292/614 (47%), Positives = 418/614 (68%), Gaps = 13/614 (2%) ref|NP_924289.11 glutathione-regulated potassium efflux system protein KefC homolog [Gloeobacter violaceus] dbj|BAC89284.11 glr1343 [Gloeobacter violaceus] Length = 634
3761.1 Best-BlastP=> >nrprot No Hits found
3763.1 Best-BlastP=> >nrprot 44% Identities = 59/195 (30%), Positives = 90/195 (46%), Gaps = 34/195 (17%) ref|NP_052362.11 unnamed protein product [Coxiella burnetii] ref|NP_819025.1 | hypothetical protein [Coxiella burnetii RSA 493] pir||S38244 hypothetical protein - Coxiella burnetii emb|CAA53132.11 unnamed protein product [Coxiella burnetii] gb|AA091585.11 hypothetical protein [Coxiella burnetii RSA 493] Length = 361
3764.3 Best-BlastP=> >nrprot No Hits found
3765.1 Best-BlastP=> >nrprot 46% Identities = 80/315 (25%), Positives = 147/315 (46%), Gaps = 20/315 (6%) ref |ZP_00124222.11 COG1721 : Uncharacterized conserved protein (some members contain a von Willebrand factor type A (vWA) domain) [Pseudomonas syringae pv. syringae B728a] Length = 327
377.1 Best-BlastP=> >nrprot 66% Identities = 137/265 (51 %), Positives = 185/265 (69%), Gaps = 6/265 (2%) ref|NP_819341.11 glutamate racemase [Coxiella burnetii RSA 493] gb|AA089855.1 | glutamate racemase [Coxiella burnetii RSA 493] Length = 280
3771.2 Best-BlastP=> >nrprot 66% Identities = 105/197 (53%), Positives = 138/197 (70%), Gaps = 2/197 (1 %) ref|NP_717670.11 phosphoribosyl-ATP pyrophosphatase/phosphoribosyl-AMP cyclohydrolase [Shewanella oneidensis MR-1] gb|AAN55114.1 |AE015648_7 phosphoribosyl-ATP pyrophosphatase/phosphoribosyl-AMP cyclohydrolase [Shewanella oneidensis MR-1] Length = 211
3772.2 Best-BlastP=> >nrprot 69% Identities = 185/353 (52%), Positives = 245/353 (69%), Gaps = 2/353 (0%) ref|NP_405132.11 histidinol-phosphatase and imidazoleglycerol-phosphate dehydratase [Yersinia pestis] ref|NP_669926.1 | imidazoleglycerolphosphate dehydratase; histidinol- phosphate phosphatase [Yersinia pestis KIM] sp|Q8ZFX7|HIS7_YERPE Histidine biosynthesis bifunctional protein hisB [Includes: Histidinol-phosphatase ; Imidazoleglycerol-phosphate dehydratase (IGPD)] pir||AF0188 imidazoleglycerol-phosphate dehydratase (EC 4.2.1.19) [imported] - Yersinia pestis (strain C092) emb|CAC90369.1 | histidinol-phosphatase and imidazoleglycerol-phosphate dehydratase [Yersinia pestis C092] gb|AAM86177.1 |AE013864_4 imidazoleglycerolphosphate dehydratase; histidinol-phosphate phosphatase [Yersinia pestis KIM] Length = 356
3778.2 Best-BlastP=> >nrprot 62% Identities = 50/84 (59%), Positives = 62/84 (73%) ref|ZP_00038938.11 COG4496: Uncharacterized protein conserved in bacteria [Xylella fastidiosa Dixon] Length = 112
3780.2 Best-BlastP=> >nrprot 23% Identities = 30/60 (50%), Positives = 37/60 (61 %), Gaps = 5/60 (8%) ref|NP_700818.11 merozoite surface protein 3 [Plasmodium falciparum 3D7] gb|AAC09377.1 | antigen [Plasmodium falciparum] gb|AAN35δ42.1 |AE014834_39 merozoite surface protein 3 [Plasmodium falciparum 3D7] Length = 364
3783.1 Best-BlastP=> >nrprot 75% Identities = 56/95 (57%), Positives = 74/95 (77%) ref|ZP_00096296.11 COG2827: Predicted endonuclease containing a URI domain [Novosphingobium aromaticivorans] Length = 111
3784.2 Best-BlastP=> >nrprot 72% Identities = 190/364 (52%), Positives = 266/364 (73%), Gaps = 3/364 (0%) ref|ZP_00122463.11 COG3842: ABC-type spermidine/putrescine transport systems, ATPase components [Haemophilus somnus 129PT] Length = 372
3785.2 Best-BlastP=> >nrprot 71 % Identities = 124/276 (44%), Positives = 203/276 (73%), Gaps = 2/276 (0%) ref|NP_439497.11 spermidine/putrescine ABC transporter permease protein [Haemophilus influenzae Rd] sp|P45170|POTB_HAEIN Spermidine/putrescine transport system permease protein potB pir||A64118 spermidine/putrescine transport system permease potB - Haemophilus influenzae (strain Rd KW20) gb|AAC22990.1 | spermidine/putrescine ABC transporter, permease protein (potB) [Haemophilus influenzae Rd] Length = 286
3788.1 Best-BlastP=> >nrprot 68% Idenfities = 115/251 (45%), Positives = 176/251 (70%) ref|ZP_00128580.11 COG1177: ABC-type spermidine/putrescine transport system, permease component II [Desulfovibrio desulfuricans G20] Length = 257
3789.2 Best-BlastP=> >nrprot 65% Identities = 122/283 (43%), Positives = 187/283 (66%), Gaps = 3/283 (1 %) ref|NP_231067.1 | spermidine/putrescine ABC transporter, periplasmic spermidine/putrescine-binding protein [Vibrio cholerae 01 biovar eltor str. N 16961] pir||B82201 spermidine/putrescine ABC transporter, periplasmic spermidine/putrescine-binding protein VC1424 [imported] - Vibrio cholerae (strain N16961 serogroup 01 ) gb|AAF94581.1 | spermidine/putrescine ABC transporter, periplasmic spermidine/putrescine-binding protein [Vibrio cholerae 01 biovar eltor str. N16961] Length = 345
3791.1 Best-BlastP=> >nrprot No Hits found
3792.1 Best-BlastP=> >nrprot No Hits found
3793.3 Best-BlastP=> >nrprot 43% Identifies = 78/264 (29%), Positives = 132/264 (50%), Gaps = 10/264 (3%) ref|ZP_00018971.11 hypothetical protein [Chloroflexus aurantiacus] Length = 303
3796.3 Best-BlastP=> >nrprot 58% Identities = 87/178 (48%), Positives = 120/178 (67%) ref|NP_840387.11 Bacterial regulatory proteins, TetR family [Nitrosomonas europaea ATCC 19718] emb|CAD84211.1 | Bacterial regulatory proteins, TetR family [Nitrosomonas europaea ATCC 19718] Length = 213
3797.1 Best-BlastP=> >nrprot 68% Identities = 167/346 (48%), Positives = 239/346 (69%), Gaps = 6/346 (1 %) ref|NP_831149.1 | Nitropropane dioxygenase / Trans-enoyl-CoA reductase family [Bacillus cereus ATCC 14δ7g] gb|AAP083δ0.11 Nitropropane dioxygenase / Trans-enoyl- CoA reductase family [Bacillus cereus ATCC 14579] Length = 363
37g8.1 Best-BlastP=> >nrprot No Hits found
3800.4 Best-BlastP=> >nrprot No Hits found
3801.2 Best-BlastP=> >nrprot 99% Identities = 477/483 (98%), Positives = 481/483 (99%) gb|AAM00644.11 adenylate cyclase [Legionella pneumophila] Length = 483
3802.2 Best-BlastP=> >nrprot 63% Identities = 189/413 (45%), Positives = 270/413 (65%), Gaps = 2/413 (0%) ref|NP_742893.11 glutamyl-tRNA reductase [Pseudomonas putida KT2440] gb|AAN66357.1 |AE016264_1 glutamyl-tRNA reductase [Pseudomonas putida KT2440] Length = 425
3803.1 Best-BlastP=> >nrprot 84% Identities = 235/358 (65%), Positives = 306/358 (85%) ref|NP_820940.11 peptide chain release factor 1 [Coxiella .' burnetii RSA 493] sp|P47849|RF1_COXBU Peptide chain release factor 1 (RF-1 ) gb|AA091454.1 | peptide chain release factor 1 [Coxiella burnetii RSA 493] Length = 361
3804.2 Best-BlastP=> >nrprot 66% Identities = 134/281 (47%), Positives = 192/281 (68%), Gaps = 6/281 (2%) ref|ZP_00066170.11 COG2890: Methylase of polypeptide chain release factors [Microbulbifer degradans 2-40] Length = 288
3807.2 Best-BlastP=> >nrprot 84% Identities = 97/133 (72%), Positives = 115/133 (86%) ref|NP_706093.11 dnaK suppressor protein [Shigella flexneri 2a str. 301] ref|NP_752128.11 DnaK suppressor protein [Escherichia coli CFT073] gb|AAN41800.1 |AE015050_16 dnaK suppressor protein [Shigella flexneri 2a str. 301] gb|AAN78672.1 |AE016755_172 DnaK suppressor protein [Escherichia coli CFT073] Length = 157
381.6 Best-BlastP=> >nrprot 24% Identities = 141/721 (19%), Positives = 282/721 (39%), Gaps = 94/721 (13%) ref |NP_010225.11 involved intracellular protein transport, coiled-coil protein necessary for protein transport from ER to Golgi; Uso1 p [Saccharomyces cerevisiae] pir||S67593 transport protein US01 - yeast (Saccharomyces cerevisiae) emb|CAA98621.11 US01 [Saccharomyces cerevisiae] Length = 1790
3810.1 Best-BlastP=> >nrprot 53% Identities = 78/237 (32%), Positives = 123/237 (51 %), Gaps = 22/237 (9%) ref|NP_721657.11 conserved hypothetical protein [Streptococcus mutans UA159] gb|AAN58863.1 |AE014954_2 conserved hypothetical protein [Streptococcus mutans UA159] Length = 246
3811.1 Best-BlastP=> >nrprot 48% Identities = 43/154 (27%), Positives = 73/154 (47%), Gaps = 35/154 (22%) ref|ZP_00074g07.11 COG0534: Na+- driven multidrug efflux pump [Trichodesmium erythraeum IMS101] Length = 931
3814.1 Best-BlastP=> >nrprot 98% Identities = 278/282 (98%), Positives = 280/282 (99%) gb|AAM73852.1 |AF454863_1 putative lipase LipA [Legionella pneumophila] Length = 282
3815.2 Best-BlastP=> >nrprot 65% Identities = 41/81 (50%), Positives = 62/81 (76%), Gaps = 2/81 (2%) ref |NP_901660.11 conserved hypothetical protein [Chromobacterium violaceum ATCC 12472] gb|AAQ59662.11 conserved hypothetical protein [Chromobacterium violaceum ATCC 12472] Length = 104
3816.2 Best-BlastP=> >nrprot 69% Identities = 121/232 (52%), Positives = 170/232 (73%) ref|NP_719120.1 | CDP-diacylglycerol-serine O- phosphatidyltransferase, putative [Shewanella oneidensis MR-1] gb|AAN56564.1 |AE015794_2 CDP-diacylglycerol-serine O- phosphatidyltransferase, putative [Shewanella oneidensis MR-1] Length = 26
3818.1 Best-BlastP=> >nrprot 81 % Identities = 50/89 (56%), Positives = 73/89 (82%) ref|NP_900485.11 sugar transport PTS system phosphocarrier protein HPR [Chromobacterium violaceum ATCC 12472] gb|AAQ58490.11 sugar transport PTS system phosphocarrier protein HPR [Chromobacterium violaceum ATCC 12472] Length = 89
3819.2 Best-BlastP=> >nrprot 76% Identities = 61/95 (64%), Positives = 76/95 (80%) dbj|BAC9321 1.11 putative sigma-54 modulation protein [Vibrio vulnificus YJ016] Length = 95
3821.1 Best-BlastP=> >nrprot No Hits found
3822.2 Best-BlastP=> >nrprot 41 % Identities = 44/167 (28%), Positives = 74/157 (47%), Gaps = 16/157 (10%) ref|NP_863847.11 hypothefical protein [Pirellula sp.] emb|CAD71520.11 hypothefical protein [Pirellula sp.] Length = 171
3824.2 Best-BlastP=> >nrprot 56% Identities = 166/428 (38%), Positives = 244/428 (57%), Gaps = 12/428 (2%) ref|NP_820492.11 mesJ protein [Coxiella burnetii RSA 493] gb|AA091006.11 mesJ protein [Coxiella burnetii RSA 493] Length = 44g
3826.1 Best-BlastP=> >nrprot 57% Identifies = 129/323 (39%), Positives = 190/323 (58%), Gaps = 9/323 (2%) ref|NP_820009.11 birA bifunctional protein [Coxiella burnetii RSA 493] gb|AAO90523.11 birA bifunctional protein [Coxiella burnetii RSA 493] Length = 323
3827.2 Best-BlastP=> >nrprot 49% Identities = 224/663 (33%), Positives = 330/663 (49%), Gaps = 66/663 (9%) ref|NP_106287.11 O-antigen acetylase [Mesorhizobium loti] dbj|BAB52073.1 | O-antigen acetylase [Mesorhizobium loti] Length = 628
3830.2 Best-BlastP=> >nrprot No Hits found
3832.1 Best-BlastP=> >nrprot 41 % Identities = 107/294 (36%), Positives = 168/294 (57%), Gaps = 16/294 (5%) ref|NP_899726.1 | probable aminopeptidase [Chromobacterium violaceum ATCC 12472] gb|AAQ57736.1 | probable aminopeptidase [Chromobacterium violaceum ATCC 12472] Length = 415
3834.1 Best-BlastP=> >nrprot No Hits found
3835.2 Best-BlastP=> >nrprot 47% Identities = 66/245 (26%), Positives = 118/245 (48%), Gaps = 10/245 (4%) emb|CAA60105.11 artJ [Escherichia coli] Length = 243
3837.3 Best-BlastP=> >nrprot 12% Identifies = 45/120 (37%), Positives = 61/120 (50%), Gaps = 8/120 (6%) gb|AAH52346.1 | 4g21520G13Rik protein [Mus musculus] Length = 379
3838.3 Best-BlastP=> >nrprot 99% Identities = 710/718 (98%), Positives = 716/718 (99%) emb|CAD90951.1 | LssB protein [Legionella pneumophila] Length = 718 384.3 Best-BlastP=> >nrprot 30% Identifies = 78/169 (46%), Positives = 11 1/169 (65%), Gaps = 2/169 (1 %) gb|AAN34371.11 ORF1 transposase [Acinetobacter baumannii] Length = 180
3840.1 Best-BlastP=> >nrprot 99% Identities = 352/355 (99%), Positives = 356/355 (100%) emb|CAD90958.11 LssD protein [Legionella pneumophila] Length = 378
3841.2 Best-BlastP=> >nrprot 91 % Identities = 719/842 (8δ%), Positives = 774/842 (91 %) emb|CAD90957.11 LssE protein [Legionella pneumophila] Length = 842
3846.2 Best-BlastP=> >nrprot 41 % Identities = 205/677 (30%), Positives = 350/677 (51 %), Gaps = 31/677 (4%) gb|AAM82673.11 PacS [Synechococcus sp. PCC 7942] Length = 747
3849.1 Best-BlastP=> >nrprot 69% Identities = 117/228 (51 %), Positives = 169/228 (74%), Gaps = 2/228 (0%) ref|ZP_00067594.11 COG0861 : Membrane protein TerC, possibly involved in tellurium resistance [Microbulbifer degradans 2-40] Length = 244
CJ -— ' CO m r-1 σs' o co r-1 CM rt cb od rt Is-! oό m m CD CD CD Is- Is- r-- Is- Is- CO TO CO
∞ TO co TO TO TO 00 TO oo co co oo co ∞ 00 00 TO co CO co co co co co CO CO co co co co CO CO CO co
3889.1 Best-BlastP=> >nrprot 48% Identities = 75/257 (29%), Positives = 126/257 (49%), Gaps = 23/257 (8%) ref| gll0032 [Gloeobacter violaceus] dbj|BAC87973.11 gll0032 [Gloeobacter violaceus] Length = 267
389.3 Best-BlastP=> >nrprot 61 % Identities = 283/637 (44%), Positives = 401/637 (62%), Gaps = 10/637 (1 %) r dependent DNA helicase-related protein [Nitrosomonas europaea ATCC 19718] emb|CAD84791.11 helicase-related protein [Nitrosomonas europaea ATCC 19718] Length = 646
3890.2 Best-BlastP=> >nrprot No Hits found
3891.2 Best-BlastP=> >nrprot 56% Identities = 85/194 (43%), Positives = 120/194 (61 %), Gaps = 7/194 (3%) ref| protein [Brucella suis 1330] gb|AAN29988.1 |AE014408_2 conserved hypothetical protein [Brucella suis 133
3892.3 Best-BlastP=> >nrprot 55% Identities = 149/349 (42%), Positives = 207/349 (59%), Gaps = 3/349 (0%) ref| esterase of the alpha-beta hydrolase superfamily [Rhodopseudomonas palustris] Length = 379
3895.2 Best-BlastP=> >nrprot 48% Identities = 92/324 (28%), Positives = 160/324 (49%), Gaps = 11/324 (3%) ref protein [Coxiella burnetii RSA 4g3] gb|AAO90332.11 multidrug resistance protein [Coxiella burnetii RSA 493
3898.2 Best-BlastP=> >nrprot No Hits found 3899.2 Best-BlastP=> >nrprot No Hits found 39.1 Best-BlastP=> >nrprot 96% Identities = 340/363 (93%), Positives = 352/363 (96%) emb|CAB60060.1 | IvhB Length = 363 390.2 Best-BlastP=> >nrprot 86% Identities = 80/106 (75%), Positives = 97/106 (91%) sp|P08811 |FER_PSEST [3Fe-4S][4Fe-4S] - Pseudomonas stutzeri prf||1410240A ferredoxin Length = 106
3901.2 Best-BlastP=> >nrprot 75% Identities = 152/272 (55%), Positives = 204/272 (75%) ref|NP_250460.11 cons [Pseudomonas aeruginosa PA01] sp|Q9l2X0|YH69_PSEAE Hypothetical UPF0085 protein PA1769 pir||D8 PA1769 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG05158.1 |AE004602_9 conserv aeruginosa PA01] Length = 274
3902.3 Best-BlastP=> >nrprot 79% Identities = 312/481 (64%), Positives = 382/481 (79%) ref |ZP_00081898.11 C involved in propionate catabolism [Geobacter metallireducens] Length = 481
3904.3 Best-BlastP=> >nrprot 54% Identities = 222/592 (37%), Positives = 332/592 (56%), Gaps = 16/592 (2%) r ABC transporter ATP-binding protein [Rickettsia conorii] pir||F97735 hypothetical protein abcT3 [imp Malish 7) gb|AAL02824.1 | multidrug resistance ABC transporter ATP-binding protein [Rickettsia con
3908.3 Best-BlastP=> >nrprot No Hits found
3909.3 Best-BlastP=> >nrprot No Hits found
3911.2 Best-BlastP=> >nrprot 78% Identities = 384/621 (61 %), Positives = 489/621 (78%), Gaps = 3/621 (0%) sp| protein htpG (Heat shock protein htpG) (High temperature protein G) Length = 624
3913.2 Best-BlastP=> >nrprot 71 % Identities = 194/356 (54%), Positives = 267/356 (75%) ref|NP_819581.11 rod s [Coxiella burnetii RSA 493] gb|AAO90095.11 rod shape-determining protein RodA [Coxiella burnetii RSA 49
co co CM CM CN CM co rt cb ro o c rt in r-1 ∞ oo co CM CN CM CM CM CM CM co co ra ro oo ro ro ro (33 ro ro ro OO ro ra co co co co co co co co co co co oo co
3936.2 Best-BlastP=> >nrprot 31% Identities = 95/227 (41 %), Positives = 131/227 (57%), Gaps = 29/227 (12%) s Phosphoribosylformylglycinamidine synthase I (FGAM synthase I) pir||T51700 phosphoribosylformylglycina component I [similarity] - Lactococcus lactis gb|AAD12625.11 phosphoribosylformylglycinamidine syn Length = 226 3937.2 Best-BlastP=> >nrprot 65% Identities = 216/436 (49%), Positives = 286/436 (65%), Gaps = 4/436 (0%) ref|
Phosphoribosylamine-glycine ligase [Methanosarcina barkeri] Length = 433
3938.2 Best-BlastP=> >nrprot 65% Identities = 75/186 (40%), Positives = 127/186 (68%), Gaps = 4/186 (2%) ref| Phosphoribosylglycinamide formyltransferase [Methanosarcina mazei Goel] gb|AAM30139.1 | Phos formyltransferase [Methanosarcina mazei Goel] Length = 202
3942.2 Best-BlastP=> >nrprot 76% Identities = 186/311 (59%), Positives = 241/311 (77%), Gaps = 2/311 (0%) ref| transfer flavoprotein, alpha subunit [Microbulbifer degradans 2-40] Length = 312
3950.2 Best-BlastP=> >nrprot 46% Identities = 148/462 (32%), Positives = 248/462 (53%), Gaps = 23/462 (4%) r [Nostoc sp. PCC 7120] pir||AB2334 hypothetical protein all4225 [imported] - Nostoc sp. (strain PCC ORF_ID:all4225-hypothetical protein [Nostoc sp. PCC 7120] Length = 565
3953.1 Best-BlastP=> >nrprot 74% Identities = 231/395 (58%), Positives = 292/395 (73%), Gaps = 7/395 (1%) ref| class-l [Nitrosomonas europaea ATCC 19718] emb|CAD84697.11 Aminotransferases class-l [Nitrosomonas = 397
3956.2 Best-BlastP=> >nrprot 50% Identities = 71/224 (31 %), Positives = 124/224 (55%), Gaps = 11/224 (4%) ref| [Nitrosomonas europaea ATCC 19718] emb|CAD84923.11 SURF1 family [Nitrosomonas europaea ATCC i
3957.1 Best-BlastP=> >nrprot 67% Identities = 28/65 (43%), Positives = 46/65 (70%) ref|NP_518489.11 PROBAB [Ralstonia solanacearum] emb|CAD13896.11 PROBABLE TRANSMEMBRANE PROTEIN [Ralstonia solana
3958.1 Best-BlastP=> >nrprot 42% Identities = 76/80 (95%), Positives = 77/80 (96%) gb|AA061477.11 unknown [ = 80
3959.2 Best-BlastP=> >nrprot 18% Identities = 60/197 (30%), Positives = 98/197 (49%), Gaps = 18/197 (9%) ref| 1700007B22 [Homo sapiens] gb|AAH24189.2| Similar to RIKEN cDNA 1700007B22 [Homo sapiens]
396.4 Best-BlastP=> >nrprot 68% Identities = 152/296 (51 %), Positives = 203/296 (68%), Gaps = 2/296 (0%) ref| [Vibrio parahaemolyticus RIMD 2210633] dbj|BAC60416.11 L-asparaginase I [Vibrio parahaemolyticus]
3960.2 Best-BlastP=> >nrprot No Hits found
3962.2 Best-BlastP=> >nrprot 57% Identities = 37/110 (33%), Positives = 65/110 (59%), Gaps = 1/110 (0%) ref|N [Coxiella burnetii RSA 493] gb|AA089681.11 cell division protein FtsL [Coxiella burnetii RSA 493] Len
3964.1 Best-BlastP=> >nrprot 72% Identities = 166/311 (53%), Positives = 224/311 (72%), Gaps = 7/311 (2%) ref| methyltransferase MraW [Photorhabdus luminescens subsp. laumondii TT01] emb|CAE16035.11 S- [Photorhabdus luminescens subsp. laumondii TT01] Length = 314
T- CN CN CM CM CM m co h-. ∞ ro CN! ,O— ; -<-
' rCMi t r+t r--.! rroή CM
Λi o
CD CD CD CD CD r^ Is- |v- f— Is- Is- Is- 00 00 ro ro ro ro ro ro ro ro ro ro ro ra ro ra co oo co co co co co co co oo co co co co
3982.2 Best-BlastP=> >nrprot 55% Identities = 62/185 (33%), Positives = 110/185 (59%), Gaps = 4/185 (2%) ref| lipoprotein LolB, putative [Coxiella burnetii RSA 493] gb|AA091322.11 outer membrane lipoprotein L 493] Length = 210
3985.1 Best-BlastP=> >nrprot 34% Identities = 40/148 (27%), Positives = 70/148 (47%) gb|AAC01725.11 rifamyci mediterranei] Length = 522
3986.1 Best-BlastP=> >nrprot 59% Identities = 155/374 (41 %), Positives = 222/374 (59%), Gaps = 11/374 (2%) r [Nostoc sp. PCC 7120] pir||AI2041 hypothetical protein all1887 [imported] - Nostoc sp. (strain PCC 7 ORF_ID:all1887-hypothetical protein [Nostoc sp. PCC 7120] Length = 375
3988.3 Best-BlastP=> >nrprot 21 % Identities = 60/228 (26%), Positives = 106/228 (46%), Gaps = 16/228 (7%) dbj [Homo sapiens] Length = 499
3989.1 Best-BlastP=> >nrprot 70% Identities = 84/147 (57%), Positives = 106/147 (72%), Gaps = 3/147 (2%) ref| protein [Salmonella enterica subsp. enterica serovar Typhi] ref|NP_461503.11 putative Cytosine/ade typhimurium LT2] ref|NP_804165.1 | conserved hypothetical protein [Salmonella enterica subsp. enterica conserved hypothetical protein STΥ2814 [imported] - Salmonella enterica subsp. enterica serovar Ty putative cytosine/adenosine deaminase [Salmonella typhimurium LT2] emb|CAD02770.1 | conserved hypot subsp. enterica serovar Typhi] gb|AAO68014.11 conserved hypothetical protein [Salmonella enterica Ty2] Length = 183
399.2 Best-BlastP=> >nrprot 20% Identities = 79/403 (19%), Positives = 171/403 (42%), Gaps = 54/403 (13%) g protein-related [Plasmodium yoelii yoelii] Length = 1441
3991.3 Best-BlastP=> >nrprot 10% Identities = 35/108 (32%), Positives = 60/108 (55%), Gaps = 4/108 (3%) ref |X [Anopheles gambiae] gb|EAA11974.11 ENSANGP00000016119 [Anopheles gambiae str. PEST] Len
3993.2 Best-BlastP=> >nrprot No Hits found
3994.5 Best-BlastP=> >nrprot 29% Identities = 41/133 (30%), Positives = 65/133 (48%), Gaps = 27/133 (20%) dbj [Chryseobacterium proteolyticum] Length = 320
3995.5 Best-BlastP=> >nrprot 29% Identities = 85/264 (32%), Positives = 125/264 (47%), Gaps = 26/264 (9%) ref| protein [Bacteroides thetaiotaomicron VPI-5482] gb|AA077157.1 | conserved hypothetical protein [Ba 5482] Length = 425
3996.2 Best-BlastP=> >nrprot No Hits found
3998.3 Best-BlastP=> >nrprot 42% Identities = 36/142 (25%), Positives = 66/142 (46%), Gaps = 10/142 (7%) ref| dehydrogenase [Escherichia coli CFT073] gb|AAN78518.1 |AE016755_18 Putative glutamate dehydrogena Length = 678
3999.3 Best-BlastP=> >nrprot 99% Identities = 641/644 (99%), Positives = 642/644 (99%) sp|032482|DNAKJ_E shock protein 70) (Heat shock 70 kDa protein) (HSP70) dbj|BAA22783.11 DnaK [Legionella pneumo
4.1 Best-BlastP=> >nrprot 59% Identities = 120/298 (40%), Positives = 181/298 (60%), Gaps = 1/298 (0%) ref protein [Wolinella succinogenes] emb|CAE10648.1 | conserved hypothetical protein [Wolinella succinogene
400.1 Best-BlastP=> >nrprot 30% Identities = 45/177 (25%), Positives = 80/177 (45%), Gaps = 25/177 (14%) ref protein [Schizosaccharomyces pombe] pir||T41077 hypothetical structural protein - fission yeast (Sc
emb|CAB53076.1 | SPCC16A11.04 [Schizosaccharomyces pombe] Length = 1010
4000.2 Best-BlastP=> >nrprot 45% Identities = 42/110 (38%), Positives = 64/110 (58%), Gaps = 5/110 (4%) ref|N transcriptional regulatory protein [Gloeobacter violaceus] dbj|BAC90918.11 MarR family transcription violaceus] Length = 143
4002.2 Best-BlastP=> >nrprot 24% Identities = 99/340 (29%), Positives = 154/340 (45%), Gaps = 53/340 (15%) r Ftsl/penicillin-binding protein 2 [Thermoanaerobacter tengcongensis] gb|AAM24853.11 Cell division [Thermoanaerobacter tengcongensis] Length = 678
4003.1 Best-BlastP=> >nrprot 50% Identities = 151/498 (30%), Positives = 248/498 (49%), Gaps = 51/498 (10%) ATPase components of ABC transporters with duplicated ATPase domains [Nostoc punctiforme]
4004.2 Best-BlastP=> >nrprot 61 % Identities = 244/575 (42%), Positives = 366/575 (63%), Gaps = 8/575 (1 %) ref ABC transporter ATP-binding protein [Rickettsia conorii] pir||F97735 hypothetical protein abcT3 [imp Malish 7) gb|AAL02824.11 multidrug resistance ABC transporter ATP-binding protein [Rickettsia con
4005.2 Best-BlastP=> >nrprot 76% Identities = 309/502 (61 %), Positives = 381/502 (75%), Gaps = 7/502 (1 %) gb [Citrobacter freundii] Length = 505
4009.1 Best-BlastP=> >nrprot 85% Identities = 254/332 (76%), Positives = 288/332 (86%) ref|NP_457439.11 pepti [Salmonella enterica subsp. enterica serovar Typhi] ref|NP_806651.11 peptide chain release factor 2 serovar Typhi Ty2] sp|P28353|RF2_SALTY Peptide chain release factor 2 (RF-2) pir||A36480 translation r typhimurium pir||AG0871 peptide chain release factor 2 (RF-2) [imported] - Salmonella enterica subs gb|AAA72914.11 peptide chain release factor 2 emb|CAD02871.11 peptide chain release factor 2 (RF-2) [Sa enterica serovar Typhi] gb|AAO70511.11 peptide chain release factor 2 [Salmonella enterica subsp. enteric Length = 365
401.2 Best-BlastP=> >nrprot 44% Identities = 128/335 (38%), Positives = 180/335 (53%), Gaps = 12/335 (3%) r [Rhodobacter sphaeroides] Length = 458
4010.1 Best-BlastP=> >nrprot 42% Identities = 26/118 (22%), Positives = 58/118 (49%), Gaps = 13/118 (11 %) ref CV3873 [Chromobacterium violaceum ATCC 12472] gb|AAQ61535.11 hypothetical protein CV3873 [Chrom Length = 117
4012.1 Best-BlastP=> >nrprot 54% Identities = 98/248 (39%), Positives = 150/248 (60%), Gaps = 14/248 (5%) ref|NP_250152.11 probable chemotaxis protein [Pseudomonas aeruginosa PA01] ref|ZP_00139088.1 | COG1360: Flagellar motor protein [Pseudomonas aeruginosa UCBPP- PA14] pir||T46617 probable chemotaxis protein PA1461 [imported] - Pseudomonas aeruginosa (strain PA01 ) dbj|BAA33552.1 | ORF2 [Pseudomonas aeruginosa] gb|AAG04850.1 |AE004575_9 probable chemotaxis protein [Pseudomonas aeruginosa PA01] Length = 296
4013.1 Best-BlastP=> >nrprot 67% Identities = 129/244 (52%), Positives = 175/244 (71 %) ref|NP_746451.11 flagellar motor protein MotA [Pseudomonas putida KT2440] gb|AAN69915.1 |AE016630_6 flagellar motor protein MotA [Pseudomonas putida KT2440] Length = 246
4014.1 Best-BlastP=> >nrprot 97% Identities = 231/238 (97%), Positives = 234/238 (98%) emb|CAA67397.11 signma factor 28 [Legionella pneumophila] Length = 238 4015.1 Best-BlastP=> >nrprot 76% Identities = 119/229 (51 %), Positives = 178/229 (77%) gb|AAC62540.2| MotR [Pseudomonas aeruginosa] Length = 275
4016.1 Best-BlastP=> >nrprot 49% Identities = 124/284 (43%), Positives = 187/284 (65%), Gaps = 5/284 (1 %) gb|AAF32412.1 | flagellar biosynthesis protein FlhF [Vibrio parahaemolyticus] Length = 503
4017.4 Best-BlastP=> >nrprot 80% Identities = 433/701 (61 %), Positives = 560/701 (79%), Gaps = 9/701 (1 %) ref|NP_250143.11 flagellar biosynthesis protein FlhA [Pseudomonas aeruginosa PA01] ref |ZP_00139079.11 C0G1298: Flagellar biosynthesis pathway, component FlhA [Pseudomonas aeruginosa UCBPP-PA14] pir||F83465 flagellar biosynthesis protein FlhA PA1452 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG04841.1 |AE004574_12 flagellar biosynthesis protein FlhA [Pseudomonas aeruginosa PA01] Length = 707
402.2 Best-BlastP=> >nrprot No Hits found
4020.2 Best-BlastP=> >nrprot 48% Identities = 177/535 (33%), Positives = 277/535 (51 %), Gaps = 2g/535 (5%) ref|NP_819244.11 conserved hypothetical protein [Coxiella burnetii RSA 493] gb|AA089758.11 conserved hypothetical protein [Coxiella burnetii RSA 493] Length = 578
4021 .2 Best-BlastP=> >nrprot 41 % Identities = 53/162 (32%), Positives = 81/162 (50%), Gaps = 3/162 (1 %) ref|NP_643713.11 conserved hypothetical protein [Xanthomonas axonopodis pv. citri str. 306] sp|Q8PH54|YY06_XANAC Hypothetical UPF014g protein XAC3406 gb|AAM38249.1 | conserved hypothetical protein [Xanthomonas axonopodis pv. citri str. 306] Length = 180
4023.2 Best-BlastP=> >nrprot 69% Identities = 221/433 (51 %), Positives = 306/433 (70%), Gaps = 1/433 (0%) ref|ZP_00092323. 1 COG0006: Xaa-Pro aminopeptidase [Azotobacter vinelandii] Length = 537
4025.1 Best-BlastP=> >nrprot 56% Identities = 161/389 (41 %), Positives = 227/389 (58%), Gaps = 12/389 (3%) ref|NP_253910.1 | ubiH protein [Pseudomonas aeruginosa PA01] pir||G82992 ubiH protein PA5223 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG08608.1 |AE004g35_5 ubiH protein [Pseudomonas aeruginosa PA01] Length = 394
4026.2 Best-BlastP=> >nrprot 59% Identities = 171/394 (43%), Positives = 229/394 (58%), Gaps = 9/394 (2%) ref | NP_716409.1 | oxidoreductase, FAD- binding, UbiH/Coq6 family [Shewanella oneidensis MR-1] gb|AAN53854.1 |AE015522_9 oxidoreductase, FAD-binding, UbiH/Coq6 family [Shewanella oneidensis MR-1] Length = 407
4030.1 Best-BlastP=> >nrprot 35% Identities = 33/123 (26%), Positives = 54/123 (43%), Gaps = 30/123 (24%) ref|NP_604443.11 dystonin isoform b; bullous pemphigoid antigen 1 ; dystonia musculorum [Mus musculus] sp|Q91ZU6|BPA1_MOUSE Bullous pemphigoid antigen 1 , isoforms 1/2/3/4 (BPA) (Hemidesmosomal plaque protein) (Dystonia musculorum protein) (Dystonin) gb|AAK83384.1 |AF396879_1 bullous pemphigoid antigen 1-b [Mus musculus] Length = 7389
4032.3 Best-BlastP=> >nrprot 76% Identities = 179/322 (55%), Positives = 242/322 (75%), Gaps = 7/322 (2%) ref|ZP_00090005.11 hypothetical protein [Azotobacter vinelandii] Length = 328
4035.1 Best-BlastP=> >nrprot 55% Identities = 47/121 (38%), Positives = 68/121 (56%), Gaps = 7/121 (5%) gb|AAL25256.11 TraK [Legionella pneumophila] Length = 114
4036.2 Best-BlastP=> >nrprot 50% Identities = 30/99 (30%), Positives = 48/99 (48%), Gaps = 6/99 (6%) ref|NP_903527.11 conserved hypothetical protein [Chromobacterium violaceum ATCC 12472] gb|AAQ61519.2| conserved hypothetical protein [Chromobacterium violaceum ATCC 12472] Length = 576
4037.2 Best-BlastP=> >nrprot 75% Identities = 42/78 (53%), Positives = 60/78 (76%) ref|NP_759375.11 Predicted transcriptional regulator [Vibrio vulnificus CMCP6] gb|AAO08902.1 |AE016798_62 Predicted transcriptional regulator [Vibrio vulnificus CMCP6] Length = 85
4039.2 Best-BlastP=> >nrprot 62% Identities = 206/434 (47%), Positives = 273/434 (62%), Gaps = 5/434 (1 %) ref|NP_932051.11 HipA protein [Photorhabdus luminescens subsp. laumondii TT01] emb|CAE17272.11 HipA protein [Photorhabdus luminescens subsp. laumondii TT01] Length = 439 4040.1 Best-BlastP=> >nrprot 32% Identities = 19/53 (35%), Positives = 32/53 (60%), Gaps = 2/53 (3%) ref|NP_297g21.11 phage-related integrase [Xylella fastidiosa 9a5c] pir||E82782 phage-related integrase XF0631 [imported] - Xylella fastidiosa (strain 9a5c) gb|AAF83441.1 |AE003908_9 phage-related integrase [Xylella fastidiosa 9a5c] Length = 413
4041.1 Best-BlastP=> >nrprot No Hits found 4042.1 Best-BlastP=> >nrprot 84% Identities = 169/231 (73%), Positives = 198/231 (85%) ref|NP_435396.11 hypothetical protein [Sinorhizobium meliloti] pir||F95280 hypothetical protein SMa0280 [imported] - Sinorhizobium meliloti (strain 1021 ) magaplasmid pSymA gb|AAK64808.1 | hypothetical protein [Sinorhizobium meliloti] Length = 262
4043.1 Best-BlastP=> >nrprot 65% Identities = 63/126 (50%), Positives = 88/126 (69%), Gaps = 5/126 (3%) ref|NP_435397.11 putative regulator, MerR family [Sinorhizobium meliloti] pir||G95280 probable regulator, MerR family [imported] - Sinorhizobium meliloti (strain 1021 ) magaplasmid pSymA gb|AAK64809.1 | putative regulator, MerR family [Sinorhizobium meliloti] Length = 134
4045.2 Best-BlastP=> >nrprot No Hits found 4047.3 Best-BlastP=> >nrprot 57% Identities = 142/327 (43%), Positives = ig5/327 (59%), Gaps = 7/327 (2%) ref |NP_819580.11 lytic murein transglycosylase [Coxiella burnetii RSA 493] gb|AAO90094.11 lytic murein transglycosylase [Coxiella burnetii RSA 493] Length = 334
4048.3 Best-BlastP=> >nrprot 86% Identities = 443/598 (74%), Positives = 520/598 (86%) ref|ZP_00090179.11 COG0481 : Membrane GTPase LepA [Azotobacter vinelandii] Length = 599
405.3 Best-BlastP=> >nrprot 11 % Identities = 44/140 (31 %), Positives = 73/140 (52%), Gaps = 4/140 (2%) pir||OXRTGU L-gulonolactone oxidase (EC 1.1.3.8) - rat dbj|BAA02232.11 L-gulono-gamma-lactone oxidase [Rattus norvegicus] Length = 440
4050.1 Best-BlastP=> >nrprot 73% Identities = 133/254 (52%), Positives = 184/254 (72%), Gaps = 9/254 (3%) ref|NP_820098.11 signal peptidase I [Coxiella burnetii RSA 493] gb|AAO90612.1 | signal peptidase I [Coxiella burnetii RSA 493] Length = 259
4051.2 Best-BlastP=> >nrprot 51% Identities = 37/118 (31 %), Positives = 67/118 (56%), Gaps = 6/118 (5%) ref|NP_842322.1 | possible transmembrane protein [Nitrosomonas europaea ATCC 19718] emb|CAD86237.1 | possible transmembrane protein [Nitrosomonas europaea ATCC 19718] Length = 126 4054.2 Best-BlastP=> >nrprot 68% Identities = 122/222 (54%), Positives = 155/222 (69%), Gaps = 4/222 (1 %) ref |NP_716968.11 ribonuclease III [Shewanella oneidensis MR-1] gb|AAN54413.1 |AE015579_2 ribonuclease III [Shewanella oneidensis MR-1] Length = 226
4055.1 Best-BlastP=> >nrprot 67% Identities = 309/628 (49%), Positives = 418/628 (66%), Gaps = 14/628 (2%) ref|ZP_00092302.11 COG0488: ATPase components of ABC transporters with duplicated ATPase domains [Azotobacter vinelandii] Length = 830
4056.2 Best-BlastP=> >nrprot 75% Identities = 247/407 (60%), Positives = 308/407 (75%) ref|NP_052356.11 unnamed protein product [Coxiella burnetii] pir||S38238 hypothetical protein - Coxiella burnetii emb|CAA53126.1 | unnamed protein product [Coxiella burnetii] emb|CAA63678.11 orf 410 [Coxiella burnetii] Length = 410
4058.2 Best-BlastP=> >nrprot No Hits found 406.1 Best-BlastP=> >nrprot 51 % Identities = 95/241 (39%), Positives = 143/241 (59%), Gaps = 6/241 (2%) ref|NP_107761.1 | unknown protein [Mesorhizobium loti] dbj|BAB53547.1 | unknown protein [Mesorhizobium loti] Length = 273
4060.1 Best-BlastP=> >nrprot 66% Identities = 208/400 (52%), Positives = 273/400 (68%), Gaps = 5/400 (1 %) ref|NP_747386.11 phosphopantothenoylcysteine decarboxylase/phosphopantothenate-cysteine ligase [Pseudomonas putida KT2440] gb|AAN70850.1 |AE016729_8 phosphopantothenoylcysteine decarboxylase/phosphopantothenate-cysteine ligase [Pseudomonas putida KT2440] Length = 403
4061.1 Best-BlastP=> >nrprot 79% Identities = 108/147 (73%), Positives = 121/147 (82%) ref |ZP_00134300.11 COG0756: dUTPase [Actinobacillus pleuropneumoniae serovar 1 str. 4074] Length = 151
4063.1 Best-BlastP=> >nrprot 66% Identities = 229/455 (50%), Positives = 310/455 (68%) ref|NP_747389.11 phosphomannomutase [Pseudomonas putida KT2440] sp|Q88C93|ALGC_PSEPK Phosphomannomutase/phosphoglucomutase (PMM / PGM) gb|AAN70853.1 |AE016729_11 phosphomannomutase [Pseudomonas putida KT2440] Length = 463
4065.3 Best-BlastP=> >nrprot 50% Identities = 181/557 (32%), Positives = 290/557 (52%), Gaps = 17/557 (3%) ref|NP_819579.11 TPR domain protein [Coxiella burnetii RSA 493] gb|AAO90093.11 TPR domain protein [Coxiella burnetii RSA 493] Length = 561
4066.2 Best-BlastP=> >nrprot 69% Identities = 102/190 (53%), Positives = 136/190 (71 %), Gaps = 2/190 (1 %) ref|NP_928489.11 hypothetical protein [Photorhabdus luminescens subsp. laumondii TT01] emb|CAE13471.1 | unnamed protein product [Photorhabdus luminescens subsp. laumondii TT01] Length = 197
4067.2 Best-BlastP=> >nrprot No Hits found
4068.2 Best-BlastP=> >nrprot 56% Identities = 174/381 (45%), Positives = 240/381 (62%), Gaps = 6/381 (1 %) ref|NP_296878.11 sodium:dicarboxylate symporter family protein [Chlamydia muridarum] gb|AAF73565.11 sodium :dicarboxylate symporter family protein [Chlamydia muridarum] Length = 415
407.4 Best-BlastP=> >nrprot 78% Identities = 2g5/44g (65%), Positives = 356/449 (79%) ref|NP_253299.11 DNA repair protein RadA [Pseudomonas aeruginosa PA01] sp|P96963|RADA_PSEAE DNA repair protein radA homolog (DNA repair protein sms homolog) pir||A83069 DNA repair protein RadA PA4609 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG07997.1 |AE004875_3 DNA repair protein RadA [Pseudomonas aeruginosa PA01] Length = 453
4070.3 Best-BlastP=> >nrprot 82% Identities = 322/464 (69%), Positives = 387/464 (83%) ref|NP_415450.11 asparagine tRNA synthetase [Escherichia coli K12] ref|NP_752997.11 Asparaginyl-tRNA synthetase [Escherichia coli CFT073] ref|NP_836636.11 asparagine tRNA synthetase [Shigella flexneri 2a str. 2457T] sp|P17242|SYN_ECOLI Asparaginyl-tRNA synthetase (Asparagine-tRNA ligase) (AsnRS) pir||SYECNT asparagine-tRNA ligase (EC 6.1.1.22) - Escherichia coli (strain K-12) emb|CAA48274.1 | Asparaginyl-tRNA synthetase [Escherichia coli] gb|AAA24666.11 asparaginyl-tRNA synthetase (asnS) dbj|BAA35682.11 Asparaginyl-tRNA synthetase (EC 6.1.1.22) (asparagine-tRNA ligase) (asnRS). [Escherichia coli K12] gb|AAC74016.1 | asparagine tRNA synthetase [Escherichia coli K12] gb|AAN79540.1 |AE016758_144 Asparaginyl-tRNA synthetase [Escherichia coli CFT073] gb|AAP16442.11 asparagine tRNA synthetase [Shigella flexneri 2a str. 2457T] Length = 466
4071.1 Best-BlastP=> >nrprot No Hits found
4072.1 Best-BlastP=> >nrprot 98% Identities = 215/218 (98%), Positives = 216/218 (99%) gb|AAC32842.11 unknown [Legionella pneumophila] Length = 218
4073.1 Best-BlastP=> >nrprot 99% Identities = 355/357 (99%), Positives = 356/357 (99%) gb|AAC32841.11 unknown [Legionella pneumophila] Length = 357
4075.3 Best-BlastP=> >nrprot 58% Identities = 69/160 (43%), Positives = 93/160 (58%), Gaps = 4/160 (2%) ref|ZP_00051893.11 COG3012: . Uncharacterized protein conserved in bacteria [Magnetospirillum magnetotacticum] Length = 163
4076.1 Best-BlastP=> >nrprot No Hits found
4078.1 Best-BlastP=> >nrprot 70% Identities = 227/395 (57%), Positives = 303/395 (76%), Gaps = 1/395 (0%) ref|ZP_00031357.11 COG0038: Chloride channel protein EriC [Burkholderia fungorum] Length = 443
408.3 Best-BlastP=> >nrprot 47% Identities = 152/478 (31 %), Positives = 258/478 (53%), Gaps = 28/478 (5%) ref |NP_820790.11 outer membrane efflux family protein [Coxiella burnetii RSA 493] gb|AA091304.11 outer membrane efflux family protein [Coxiella burnetii RSA 493] Length = 507
4080.2 Best-BlastP=> >nrprot 89% Identities = 377/385 (97%), Positives = 381/385 (98%) gb|AAD47247.11 putative transport protein [Legionella pneumophila] Length = 387
4082.2 Best-BlastP=> >nrprot 34% Identities = 78/288 (27%), Positives = 138/288 (47%), Gaps = 2/288 (0%) ref|NP_931673.11 hypothetical protein [Photorhabdus luminescens subsp. laumondii TT01] emb|CAE16879.11 unnamed protein product [Photorhabdus luminescens subsp. laumondii TT01] Length = 833
4083.3 Best-BlastP=> >nrprot No Hits found 4084.2 Best-BlastP=> >nrprot 55% Identities = 143/383 (37%), Positives = 217/383 (56%), Gaps = 1/383 (0%) ref|ZP_00058706.11 COG0438: Glycosyltransferase [Thermobifida fusca] Length = 426
4087.2 Best-BlastP=> >nrprot 61 % Identities = 175/436 (40%), Positives = 262/436 (60%), Gaps = 25/436 (5%) ref |NP_845547.11 conserved hypothetical protein [Bacillus anthracis str. Ames] gb|AAP27033.1 | conserved hypothetical protein [Bacillus anthracis str. Ames] Length = 445 4088.2 Best-BlastP=> >nrprot 75% Identities = 182/288 (63%), Positives = 224/288 (77%) gb|AAM00632.11 unknown [Legionella pneumophila] Length = 294 409.2 Best-BlastP=> >nrprot 56% Identities = 123/352 (34%), Positives = 210/352 (59%), Gaps = 8/352 (2%) ref|NP_820789.11 lipoprotein, putative [Coxiella burnetii RSA 493] gb|AA091303.11 lipoprotein, putative [Coxiella burnetii RSA 493] Length = 391
4093.1 Best-BlastP=> >nrprot 72% Identities = 255/481 (53%), Positives = 346/481 (71 %), Gaps = 9/481 (1 %) ref|NP_820458.11 glutamyl-tRNA(Gln) amidotransferase, B subunit [Coxiella burnetii RSA 493] gb|AAO90972.11 glutamyl-tRNA(Gln) amidotransferase, B subunit [Coxiella burnetii RSA 4 3] Length = 477
4094.2 Best-BlastP=> >nrprot 75% Identities = 284/480 (59%), Positives = 366/480 (76%), Gaps = 1/480 (0%) ref|NP_820457.11 glutamyl-tRNA(Gln) amidotransferase, A subunit [Coxiella burnetii RSA 493] gb|AAO90971.11 glutamyl-tRNA(Gln) amidotransferase, A subunit [Coxiella burnetii RSA 4 3] Length = 483
4096.1 Best-BlastP=> >nrprot 70% Identities = 86/155 (55%), Positives = 113/155 (72%) ref |NP_841624.11 Transposase IS4 family [Nitrosomonas europaea ATCC 19718] ref|NP_841817.1 | Transposase IS4 family [Nitrosomonas europaea ATCC 19718] ref|NP_842206.1 | Transposase IS4 family [Nitrosomonas europaea ATCC 19718] ref|NP_842438.11 Transposase IS4 family [Nitrosomonas europaea ATCC 19718] emb|CAD85496.11 Transposase IS4 family [Nitrosomonas europaea ATCC 19718] emb|CAD86113.11 Transposase IS4 family [Nitrosomonas europaea ATCC 19718] emb|CAD85700.11 Transposase IS4 family [Nitrosomonas europaea ATCC 19718] emb|CAD86358.11 Transposase IS4 family [Nitrosomonas europaea ATCC 19718] Length = 191
4097.1 Best-BlastP=> >nrprot 62% Identities = 26/34 (76%), Positives = 31/34 (91 %) ref|ZP_00111545.11 COG4644: Transposase and inactivated derivatives, TnpA family [Nostoc punctiforme] Length = 1014
4098.1 Best-BlastP=> >nrprot 62% Identities = 33/89 (37%), Positives = 57/89 (64%) gb|AA092366.11 transposase [Listonella anguillarum] Length = 980
410.1 Best-BlastP=> >nrprot 70% Identities = 125/226 (55%), Positives = 166/226 (73%) ref|NP_721272.11 putative ABC transporter, ATP-binding protein [Streptococcus mutans UA159] gb|AAN58578.1 |AE014927_7 putative ABC transporter, ATP-binding protein [Streptococcus mutans UA159] Length = 235
4101.1 Best-BlastP=> >nrprot 50% Identities = 22/49 (44%), Positives = 34/49 (69%) ref|NP_277100.11 putative transposase [Deinococcus radiodurans] Length = 828
4104.2 Best-BlastP=> >nrprot 44% Identities = 121/430 (28%), Positives = 195/430 (45%), Gaps = 67/430 (15%) ref|NP_873428.1 | conserved hypothetical protein [Haemophilus ducreyi 35000HP] gb|AAP95817.1 | conserved hypothetical protein [Haemophilus ducreyi 35000HP] Length = 481
4105.1 Best-BlastP=> >nrprot 48% Identities = 74/241 (30%), Positives = 126/241 (52%), Gaps = 19/241 (7%) ref|ZP_00132729.11 hypothetical protein [Haemophilus somnus 2336] Length = 270
4106.1 Best-BlastP=> >nrprot 39% Identities = 47/197 (23%), Positives = 88/197 (44%), Gaps = 18/197 (9%) ref|ZP_00123126.11 hypothetical protein [Haemophilus somnus 129PT] Length = 212
4107.1 Best-BlastP=> >nrprot No Hits found
4108.1 Best-BlastP=> >nrprot 34% Identities = 61/215 (28%), Positives = 97/215 (45%), Gaps = 29/215 (13%) ref|NP_928397.1 | hypothetical protein [Photorhabdus luminescens subsp. laumondii TT01] emb|CAE13363.1 | unnamed protein product [Photorhabdus luminescens subsp. laumondii TT01] Length = 245
4109.2 Best-BlastP=> >nrprot 34% Identities = 36/145 (24%), Positives = 67/145 (46%), Gaps = 16/145 (11%) ref|XP_323218.11 hypothetical protein ( (AL356172) conserved hypothetical protein [Neurospora crassa] ) emb|CAD11783.11 conserved hypothetical protein [Neurospora crassa] gb|EAA28302.11 hypothetical protein ( (AL356172) conserved hypothetical protein [Neurospora crassa] ) Length = 743
4111.2 Best-BlastP=> >nrprot 97% Identities = 350/359 (97%), Positives = 350/359 (97%), Gaps = 8/359 (2%) emb|CAB65206.11 RmlB protein [Legionella pneumophila] Length = 351
4112.1 Best-BlastP=> >nrprot 99% Identities = 294/294 (100%), Positives = 294/294 (100%) emb|CAB65207.11 RmlD protein [Legionella pneumophila] Length = 2 4 4113.1 Best-BlastP=> >nrprot 94% Identities = 172/188 (91%), Positives = 177/188 (94%), Gaps = 1/188 (0%) emb|CAB65208.1 | RmlC protein [Legionella pneumophila] Length = 188
4115.1 Best-BlastP=> >nrprot 7% Identities = 331/334 (g9%), Positives = 332/334 (99%) emb|CAB65212.11 N-acetylneuraminic acid condensing enzyme [Legionella pneumophila] Length = 338
4116.3 Best-BlastP=> >nrprot 99% Identities = 232/232 (100%), Positives = 232/232 (100%) emb|CAB65213.11 CMP-N-acetlyneuraminic acid synthetase [Legionella pneumophila] Length = 232
4118.3 Best-BlastP=> >nrprot 99% Identities = 213/213 (100%), Positives = 213/213 (100%) sp|Q9RDX3|HIS5_LEGPN Imidazole glycerol phosphate synthase subunit hisH (IGP synthase glutamine amidotransferase subunit) (IGP synthase subunit hisH) (ImGP synthase subunit hisH) (IGPS subunit hisH) emb|CAB65214.1 | glutamine amidotransferase [Legionella pneumophila] Length = 213
4119.2 Best-BlastP=> >nrprot 82% Identities = 207/210 (98%), Positives = 210/210 (100%) sp|Q9RDX2|HIS6_LEGPN Imidazole glycerol phosphate synthase subunit hisF (IGP synthase cyclase subunit) (IGP synthase subunit hisF) (ImGP synthase subunit hisF) (IGPS subunit hisF) emb|CAB65215.1 | HisF protein [Legionella pneumophila] Length = 212
412.3 Best-BlastP=> >nrprot 66% Identities = 189/397 (47%), Positives = 263/397 (66%) ref|NP_820787.11 ABC transporter, permease protein [Coxiella burnetii RSA 493] gb|AA091301.11 ABC transporter, permease protein [Coxiella burnetii RSA 493] Length = 404
4123.2 Best-BlastP=> >nrprot 63% Identities = 39/74 (52%), Positives = 54/74 (72%), Gaps = 1/74 (1 %) gb|AAM08234.11 putative phage repressor [Legionella pneumophila] Length = 227
4126.2 Best-BlastP=> >nrprot 54% Identities = 118/267 (44%), Positives = 164/267 (61 %), Gaps = 2/267 (0%) gb|AAM08235.11 LvrA [Legionella pneumophila] Length = 289
4127.1 Best-BlastP=> >nrprot 51 % Identities = 47/155 (30%), Positives = 77/155 (49%), Gaps = 20/155 (12%) gb|AAM08236.11 LvrB [Legionella pneumophila] Length = 150
4128.1 Best-BlastP=> >nrprot 61 % Identities = 25/62 (40%), Positives = 45/62 (72%), Gaps = 2/62 (3%) emb|CAB60050.11 IvrC [Legionella pneumophila] Length = 67
4129.1 Best-BlastP=> >nrprot 40% Identities = 37/114 (32%), Positives = 62/114 (54%), Gaps = 4/114 (3%) gb|AAL05416.11 PilL [Yersinia pseudotuberculosis] Length = 356
413.5 Best-BlastP=> >nrprot 23% Identities = 104/399 (26%), Positives = 187/399 (46%), Gaps = 28/399 (7%) ref|NP_486788.11 hypothetical protein [Nostoc sp. PCC 7120] pir||AE2149 hypothetical protein all2748 [imported] - Nostoc sp. (strain PCC 7120) dbj|BAB74447.1 | ORF_ID:all2748~hypothetical protein [Nostoc sp. PCC 7120] Length = 426
4130.2 Best-BlastP=> >nrprot 45% Identities = 33/104 (31 %), Positives = 60/104 (57%), Gaps = 6/104 (5%) ref |NP_819572.11 SMC family protein [Coxiella burnetii RSA 493] emb|CAD66594.1 | SMC protein [Coxiella burnetii] gb|AAO90086.1 | SMC family protein [Coxiella burnetii RSA 493] Length = 1169
4131.3 Best-BlastP=> >nrprot 31 % Identities = 60/185 (32%), Positives = 87/185 (47%), Gaps = 8/185 (4%) gb|AAN62293.1 |AF440524_80 hypothetical protein [Pseudomonas aeruginosa] Length = 241
4132.3 Best-BlastP=> >nrprot 45% Identities = 42/99 (42%), Positives = 58/99 (58%), Gaps = 2/99 (2%) ref|ZP_00123136.11 hypothetical protein [Haemophilus somnus 129PT] Length = 170
4133.1 Best-BlastP=> >nrprot 75% Identities = 39/61 (63%), Positives = 49/61 (80%) ref|NP_403868.11 50S ribosomal protein L29 [Yersinia pestis] ref|NP_671291.1 | 50S ribosomal subunit protein L29 [Yersinia pestis KIM] sp|Q8ZJA4|RL29_YERPE 50S ribosomal protein L29 pir||AB0027 50S ribosomal protein L29 [imported] - Yersinia pestis (strain C092) emb|CAC89077.1 | 50S ribosomal protein L29 [Yersinia pestis C092] gb|AAM87542.1 |AE014002_15 50S ribosomal subunit protein L29 [Yersinia pestis KIM] Length = 63
4134.1 Best-BlastP=> >nrprot 77% Identities = 54/79 (68%), Positives = 66/79 (83%) ref|ZP_00090912.1 | COG0186: Ribosomal protein S17 [Azotobacter vinelandii] Length = 90
4135.1 Best-BlastP=> >nrprot 90% Identities = 100/122 (81 %), Positives = 110/122 (90%), Gaps = 1/122 (0%) ref|ZP_00067983.11 COG0093: Ribosomal protein L14 [Microbulbifer degradans 2-40] Length = 122
4136.1 Best-BlastP=> >nrprot 70% Identities = 58/106 (54%), Positives = 77/106 (72%), Gaps = 1/106 (0%) ref|NP_273211.1 | 50S ribosomal protein L24 [Neisseria meningitidis MC58] ref|NP_282968.1 | 50S ribosomal protein L24 [Neisseria meningitidis Z2491] pir||C81232 50S ribosomal protein L24 NMB0153 [imported] - Neisseria meningitidis (strain MC58 serogroup B, strain Z2491 serogroup A) gb|AAF40611.11 50S ribosomal protein L24 [Neisseria meningitidis MC58] emb|CAB83433.11 50S ribosomal protein L24 [Neisseria meningitidis Z2491] Length = 107
4138.1 Best-BlastP=> >nrprot 83% Identities = 128/178 (71 %), Positives = 153/178 (85%) ref|NP_273212.11 50S ribosomal protein L5 [Neisseria meningitidis MC58] pir||D81232 50S ribosomal protein L5 NMB0154 [imported] - Neisseria meningitidis (strain MC58 serogroup B) gb|AAF40612.11 50S ribosomal protein L5 [Neisseria meningitidis MC58] Length = 17
4139.1 Best-BlastP=> >nrprot 65% Identities = 49/101 (48%), Positives = 66/101 (65%), Gaps = 1/101 (0%) ref|ZP_00004335.1 | COG0199: Ribosomal protein S14 [Rhodobacter sphaeroides] Length = 101
4140.1 Best-BlastP=> >nrprot 78% Identities = 79/133 (59%), Positives = 103/133 (77%), Gaps = 5/133 (3%) ref|NP_273214.1 | 30S ribosomal protein S8 [Neisseria meningitidis MC58] ref|NP_282965.11 30S ribosomal protein S8 [Neisseria meningitidis Z2491] sp|Q9JR58|RS8_NEIMA 30S ribosomal protein S8 pir||F81232 30S ribosomal protein S8 NMB0156 [imported] - Neisseria meningitidis (strain MC58 serogroup B, strain Z2491 serogroup A) gb|AAF40614.11 30S ribosomal protein S8 [Neisseria meningitidis MC58] emb|CAB83430.11 30S ribosomal protein S8 [Neisseria meningitidis Z2491] Length = 130
4141.1 Best-BlastP=> >nrprot 71 % Identities = 102/179 (56%), Positives = 129/179 (72%), Gaps = 2/179 (1 %) ref|NP_289866.11 50S ribosomal subunit protein L6 [Escherichia coli 0157:H7 EDL933] ref|NP_312197.1 | 50S ribosomal subunit protein L6 [Escherichia coli 0157:H7] ref | NP_417764.1 | 50S ribosomal subunit protein L6 [Escherichia coli K12] ref|NP_709093.1 [ 50S ribosomal subunit protein L6 [Shigella flexneri 2a str. 301] ref|NP_755932.11 50S ribosomal protein L6 [Escherichia coli CFT073] ref|NP_839565.11 50S ribosomal subunit protein L6 [Shigella flexneri 2a str. 2457T] sp|P02390|RL6_ECOLI 50S ribosomal protein L6 pir||R5EC6 ribosomal protein L6 [validated] - cherichia coli (strain K-12) pir||B91150 50S ribosomal subunit protein L6 [imported] - Escherichia coli (strain 0157:H7, substrain RIMD 0509952) 995 50S ribosomal subunit protein L6 [imported] - Escherichia coli (strain 0157:H7, substrain EDL933) gb|AAA58102.11 50S ribosomal subunit protein L6 [Escherichia coli] gb|AAC76330.1 | 50S ribosomal subunit protein L6 [Escherichia coli K12] E005556_19 50S ribosomal subunit protein L6 [Escherichia coli 0157:H7 EDL933] dbj|BAB37593.11 50S ribosomal subunit protein L6 [Escherichia coli 0157:H7] E015343_ 6 50S ribosomal subunit protein L6 [Shigella fie
4142.3 Best-BlastP=> >nrprot 71 % Identities = 65/117 (55%), Positives = 86/117 (73%), Gaps = 4/117 (3%) ref|NP_252937.11 50S ribosomal protein L18 [Pseudomonas aeruginosa PA01] ref |ZP_00137735.1 | COG0256: Ribosomal protein L18 [Pseudomonas aeruginosa UCBPP-PA14] pir||E83114 50S ribosomal protein L18 PA4247 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG07635.1 |AE004841_13 50S ribosomal protein L18 [Pseudomonas aeruginosa PA01] Length = 116
4143.3 Best-BlastP=> >nrprot 75% Identities = 105/158 (66%), Positives = 127/158 (80%) ref|ZP_00067978.11 COG0098: Ribosomal protein S5 [Microbulbifer degradans 2-40] Length = 170
4144.3 Best-BlastP=> >nrprot 66% Identities = 31/56 (55%), Positives = 41/56 (73%) ref|NP_563303.11 50S ribosomal protein L30 [Clostridium perfringens] dbj|BAB82093.11 50S ribosomal protein L30 [Clostridium perfringens str. 13] Length = 57
4145.3 Best-BlastP=> >nrprot 76% Identities = 92/144 (63%), Positives = 111/144 (77%) ref|ZP_00067977.11 COG0200: Ribosomal protein L15 [Microbulbifer degradans 2-40] Length = 144
4148 3 Best-BlastP=> >nrprot 81% Identities = 2g3/437 (67%), Positives = 364/437 (83%) ref|NP_819302.11 preprotein translocase, SecY subunit [Coxiella burnetii RSA 493] gb|AA089816.11 preprotein translocase, SecY subunit [Coxiella burnetii RSA 493] Length = 442
4149.1 Best-BlastP=> >nrprot 81 % Identities = 83/114 (72%), Positives = 97/114 (85%) ref|ZP_00125957.11 COG0099: Ribosomal protein S13 [Pseudomonas syringae pv. syringae B728a] ref|NP_790495.1 | ribosomal protein S13 [Pseudomonas syringae pv. tomato str. DC3000] sp|Q889U9|RS13_PSESM 30S ribosomal protein S13 gb|AAO54190.1 | ribosomal protein S13 [Pseudomonas syringae pv. tomato str. DC3000] Length = 118
415.2 Best-BlastP=> >nrprot 57% Identities = 176/280 (62%), Positives = 213/280 (76%), Gaps = 2/280 (0%) ref|NP_838055.11 putative transposase [Shigella flexneri 2a str. 2457T] gb|AAK64580.11 putative transposase [Vibrio cholerae] dbj|BAB79611.11 orf6 [Salmonella enterica subsp. enterica serovar Choleraesuis] gb|AAL59686.1 | putative transposase [Vibrio cholerae] gb|AAP17865.1 | putative transposase [Shigella flexneri 2a str. 2457T] dbj|BAC79056.11 putative transposase [Vibrio cholerae] Length = 497
4151.1 Best-BlastP=> >nrprot 80% Identities = 91/130 (70%), Positives = 107/130 (82%) ref |NP_660814.11 30S ribosomal protein S11 [Buchnera aphidicola str. Sg (Schizaphis graminum)] sp|Q8K972|RS11_BUCAP 30S ribosomal protein S11 gb|AAM68025.11 30S ribosomal protein S11 [Buchnera aphidicola str. Sg (Schizaphis graminum)] Length = 130
4153.1 Best-BlastP=> >nrprot 81 % Identities = 139/206 (67%), Positives = 169/206 (82%) ref|NP_289857.1 | 30S ribosomal subunit protein S4 [Escherichia coli 0157:H7 EDL933] ref|NP_312188.1 | 30S ribosomal subunit protein S4 [Escherichia coli 0157:H7] ref|NP_417755.11 30S ribosomal subunit protein S4 [Escherichia coli K12] ref|NP_755921.11 30S ribosomal protein S4 [Escherichia coli CFT073] 54|RS4_ECOLI 30S ribosomal protein S4 pir||R3EC4 ribosomal protein S4 [validated] - Escherichia coli (strain K-12) pir||A91149 30S ribosomal subunit protein S4 [imported] - Escherichia coli (strain 0157:H7, substrain RIMD 0509952) pir||E85994 30S ribosomal subunit protein S4 [imported] - Escherichia coli (strain 0157:H7, substrain EDL933) emb|CAA26394.1 | unnamed protein product [Escherichia coli] gb|AAA58094.1 | 30S ribosomal subunit protein S4 [Escherichia coli] gb|AAC76321.1 | 30S ribosomal subunit protein S4 [Escherichia coli K12] .1 |AE005556_10 30S ribosomal subunit protein S4 [Escherichia coli 0157:H7 EDL933] dbj|BAB37584.1 | 30S ribosomal subunit protein S4 [Escherichia coli 0157:H7] gb|AAN82495.1 |AE016767_255 30S ribosomal protein S4 [Escherichia coli CFT073] Length = 206
4154.2 Best-BlastP=> >nrprot 79% Identities = 216/328 (65%), Positives = 264/328 (80%), Gaps = 3/328 (0%) gb|AAM33636.1 |AF506984_1 RpoA [Pseudomonas putida] Length = 333
4156.3 Best-BlastP=> >nrprot 99% Identities = 283/283 (100%), Positives = 283/283 (100%) emb|CAB65193.11 Wzm protein [Legionella pneumophila] Length = 283
4157.2 Best-BlastP=> >nrprot 99% Identities = 473/474 (99%), Positives = 473/474 (99%) emb|CAB65192.11 Wzt protein [Legionella pneumophila] . Length = 474
4158.1 Best-BlastP=> >nrprot 98% Identities = 379/382 (99%), Positives = 379/382 (99%) emb|CAB65191.11 hypothetical protein [Legionella pneumophila] Length = 382
4159.1 Best-:BlastP=> >nrprot 91% Identities = 273/279 (97%), Positives = 275/279 (98%) emb|CAB65190.11 putative glycosyl transferase [Legionella pneumophila] Length = 297
4160.3 Best-BlastP=> >nrprot 97% Identities = 281/282 (99%), Positives = 281/282 (99%) emb|CAD43478.11 putative glycosyltransferase [Legionella pneumophila] Length = 2 7
4161 '3 Best-BlastP=> >nrprot 99% Identities = 336/339 (99%), Positives = 338/339 (99%) emb|CAB65189.11 putative glycosyl transferase [Legionella pneumophila] Length = 339
4162.2 Best-BlastP=> >nrprot 37% Identities = 188/579 (32%), Positives = 316/579 (54%), Gaps = 35/579 (6%) ref|NP_834965.11 Sensory box/GGDEF family protein [Bacillus cereus ATCC 14579] gb|AAP12166.11 Sensory box/GGDEF family protein [Bacillus cereus ATCC 1457g] Length = 909
4164.1 Best-BlastP=> >nrprot 57% Identities = 138/138 (100%), Positives = 138/138 (100%) gb|AAD41585.1 |AF057704_1 EnhA [Legionella pneumophila] Length = 164
4165.1 Best-BlastP=> >nrprot 68% Identities = 127/133 (95%), Positives = 130/133 (97%) gb|AAD41586.1 |AF057704_2 EnhB [Legionella pneumophila] Length = 142
4167.4 Best-BlastP=> >nrprot 99% Identities = 1193/1201 (99%), Positives = 1198/1201 (99%), Gaps = 1/1201 (0%) gb|AAD41587.1 |AF057704_3 enhanced entry protein EnhC [Legionella pneumophila] Length = 1201
4168.2 Best-BlastP=> >nrprot No Hits found
417.3 Best-BlastP=> >nrprot 30% Identities = 45/190 (23%), Positives = 83/190 (43%), Gaps = 2/190 (1 %) ref|ZP_00091084.11 COG0582: Integrase [Azotobacter vinelandii] Length = 287
4170.1 Best-BlastP=> >nrprot 6% Identities = 36/1 17 (30%), Positives = 57/117 (48%), Gaps = 10/1 17 (8%) ref|NP_701057.11 hypothetical protein [Plasmodium falciparum 3D7] gb|AAN35781.1 |AE014838_59 hypothetical protein [Plasmodium falciparum 3D7] Length = 371
4171.4 Best-BlastP=> >nrprot 52% Identities = 69/147 (46%), Positives = 93/147 (63%), Gaps = 1/147 (0%) ref|ZP_00080184.1 1 COG1881 : Phospholipid-binding protein [Geobacter metallireducens] Length = 176
4172.2 Best-BlastP=> >nrprot 11 % Identities = 22/63 (34%), Positives = 39/63 (61 %), Gaps = 1/63 (1 %) gb|EAA24489.1 | hypothetical protein [Fusobacterium nucleatum subsp. vincentii ATCC 49256] Length = 265
4173.2 Best-BlastP=> >nrprot 62% Identities = 68/129 (52%), Positives = 84/129 (65%), Gaps = 4/129 (3%) ref|NP_230027.11 conserved hypothetical protein [Vibrio cholerae] pir||A82330 conserved hypothetical protein VC0373 [imported] - Vibrio cholerae (strain N16961 serogroup 01 ) gb|AAF93546.11 conserved hypothetical protein [Vibrio cholerae] Length = 139
4174.1 Best-BlastP=> >nrprot 66% Identities = 57/143 (39%), Positives = 93/143 (65%), Gaps = 7/143 (4%) gb|AAC44222.11 hemin binding protein Hbp [Legionella pneumophila] Length = 141
4175.1 Best-BlastP=> >nrprot 54% Identities = 115/302 (38%), Positives = 169/302 (55%), Gaps = 11/302 (3%) ref |ZP_00086085.1 1 hypothetical protein [Pseudomonas fluorescens PfO-1] Length = 300
4177.2 Best-BlastP=> >nrprot 13% Identities = 48/172 (27%), Positives = 70/172 (40%), Gaps = 25/172 (14%) dbj|BAC27865.11 unnamed protein product [Mus musculus] Length = 531
417g.2 Best-BlastP=> >nrprot 27% Identities = 86/423 (20%), Positives = 183/423 (43%), Gaps = 69/423 (16%) gb|EAA18183.1 | hypothetical protein [Plasmodium yoelii yoelii] Length = 1154
418*4 Best-BlastP=> >nrprot 69% Identities = 119/215 (55%), Positives = 156/215 (72%), Gaps = 3/215 (1 %) ref|NP_286845.11 putative carrier/transport protein [Escherichia coli 0157:H7 EDL933] ref|NP_309081 .1 | putative carrier/transport protein [Escherichia coli 0157:H7] pir||D85624 probable carrier/transport protein yccA [imported] - Escherichia coli (strain 0157:H7, substrain EDL933) pir||F90760 probable carrier/transport protein ECs1054 [imported] - Escherichia coli (strain 0157:H7, substrain RIMD 0509952) gb|AAG55456.1 |AE005287_3 putative carrier/transport protein [Escherichia coli 0157:H7 EDL933] dbj|BAB34477.11 putative carrier/transport protein [Escherichia coli 0157:H7] Length = 219
4181.1 Best-BlastP=> >nrprot No Hits found
4184.1 Best-BlastP=> >nrprot 53% Identities = 34/97 (35%), Positives = 56/97 (57%) ref|NP_820123.11 conserved hypothetical protein [Coxiella burnetii RSA 493] gb|AAO90637.1 1 conserved hypothetical protein [Coxiella burnetii RSA 493] Length = 106
4186.1 Best-BlastP=> >nrprot 70% Identities = 155/310 (50%), Positives = 210/310 (67%), Gaps = 16/310 (5%) ref|NP_841212.1 | DnaJ N-terminal domain:DnaJ C terminal domain [Nitrosomonas europaea ATCC 19718] emb|CAD85066.1 | DnaJ N-terminal domain:DnaJ C terminal domain [Nitrosomonas europaea ATCC 19718] Length = 314
4188.1 Best-BlastP=> >nrprot 65% Identities = 171/397 (43%), Positives = 244/397 (61 %), Gaps = 36/397 (9%) ref|ZP_00016589.11 hypothetical protein [Rhodospirillum rubrum] Length = 402
419.2 Best-BlastP=> >nrprot 67% Identities = 136/265 (51 %), Positives = 182/265 (68%), Gaps = 4/265 (1 %) ref|NP_253165.1 1 conserved hypothetical protein [Pseudomonas aeruginosa PA01] pir||E83086 conserved hypothetical protein PA4475 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG07863.1 |AE004861_4 conserved hypothetical protein [Pseudomonas aeruginosa PA01] Length = 282
4191.1 Best-BlastP=> >nrprot 71 % Identities = 258/457 (56%), Positives = 333/457 (72%), Gaps = 3/457 (0%) ref|ZP_00009339.11 hypothetical protein [Rhodopseudomonas palustris] Length = 471 4192.2 Best-BlastP=> >nrprot 66% Identities = 305/605 (50%), Positives = 398/605 (65%), Gaps = 21/605 (3%) ref|ZP_00029131.11 hypothetical protein [Burkholderia fungorum] Length = 642 4194.3 Best-BlastP=> >nrprot 99% Identities = 428/429 (99%), Positives = 428/429 (99%) gb|AAM00638.11 unknown [Legionella pneumophila] Length = 429 4195.3 Best-BlastP=> >nrρrot 46% Identities = 100/388 (25%), Positives = 180/388 (46%), Gaps = 7/388 (1 %) ref|NP_820942.11 drug resistance transporter, Bcr/CflA family [Coxiella burnetii RSA 493] gb|AA091456.11 drug resistance transporter, Bcr/CflA family [Coxiella burnetii RSA 493] Length = 409
4197.2 Best-BlastP=> >nrprot 91% Identities = 725/730 (99%), Positives = 727/730 (99%) gb|AAK35045.2|AF330136_1 type II protein secretion LspD [Legionella pneumophila] Length = 730
42.1 Best-BlastP=> >nrprot 98% Identities = 243/250 (97%), Positives = 247/250 (98%) gb|AAM08246.11 probable conjugal transfer protein [Legionella pneumophila] Length = 250
4200.2 Best-BlastP=> >nrprot No Hits found 4203.2 Best-BlastP=> >nrprot 47% Identities = 121/412 (29%), Positives = 202/412 (49%), Gaps = 29/412 (7%) ref|ZP_00034486.1 | COG0642: Signal transduction histidine kinase [Burkholderia fungorum] Length = 479
4205.2 Best-BlastP=> >nrprot No Hits found 4206.2 Best-BlastP=> >nrρrot 33% Identities = 156/825 (18%), Positives = 318/825 (38%), Gaps = 118/825 (14%) gb|EAA20682.1 | rhoptry protein [Plasmodium yoelii yoelii] Length = 2719
4209.2 Best-BlastP=> >nrprot 42% Identities = 72/314 (22%), Positives = 139/314 (44%), Gaps = 28/314 (8%) gb|AAB03184.1 | TnpA [Pseudomonas putida] Length = 584
421.2 Best-BlastP=> >nrprot 78% Identities = 307/479 (64%), Positives = 377/479 (78%), Gaps = 4/479 (0%) ref|NP_520780.11 PROBABLE TLDD PROTEIN [Ralstonia solanacearum] emb|CAD16366.11 PROBABLE TLDD PROTEIN [Ralstonia solanacearum] Length = 486
4211.2 Best-BlastP=> >nrprot 61 % Identities = 182/449 (40%), Positives = 276/449 (61 %), Gaps = 11/449 (2%) ref|ZP_00056132.1 | COG1355: Predicted dioxygenase [Magnetospirillum magnetotacticum] Length = 468 4212.1 Best-BlastP=> >nrprot 72% Identities = 22g/348 (65%), Positives = 263/348 (75%) ref|NP_217654.11 pflA [Mycobacterium tuberculosis H37Rv] ref|NP_337751.1 | pyruvate formate lyase-activating enzyme, putative [Mycobacterium tuberculosis CDC1551] pir||C70646 probable pflA protein - Mycobacterium tuberculosis (strain H37RV) emb|CAB062g2.1 | pflA [Mycobacterium tuberculosis H37Rv] gb|AAK47565.1 | pyruvate formate lyase-activating enzyme, putative [Mycobacterium tuberculosis CDC1551] Length = 362
4214.3 Best-BlastP=> >nrprot 54% Identities = 176/517 (34%), Positives = 286/517 (55%), Gaps = 21/517 (4%) ref |NP_790671.1 | sensor protein PilS [Pseudomonas syringae pv. tomato str. DC3000] gb|AA054366.1 | sensor protein PilS [Pseudomonas syringae pv. tomato str. DC3000] Length = 531
422.3 Best-BlastP=> >nrprot 63% Identities = 127/295 (43%), Positives = 187/295 (63%), Gaps = 14/295 (4%) ref|NP_460713.11 putative phosphoesterase [Salmonella typhimurium LT2] gb|AAL20672.11 putative phosphoesterase [Salmonella typhimurium LT2] Length = 301
4220.2 Best-BlastP=> >nrprot 69% Identities = 49/91 (53%), Positives = 68/91 (74%) gb|AAP83334.1 |AF469614_2 unknown [Francisella tularensis subsp. tularensis] Length = 94
4221.3 Best-BlastP=> >nrprot 58% Identities = 38/86 (44%), Positives = 54/86 (62%) dbj|BAC94314.11 acylphosphatase [Vibrio vulnificus YJ016] Length = 90
4222.3 Best-BlastP=> >nrprot 45% Identities = 33/83 (39%), Positives = 48/83 (57%) gb|EAA16908.11 Drosophila melanogaster CG8797 gene product- related [Plasmodium yoelii yoelii] Length = 2198
4225.1 Best-BlastP=> >nrprot No Hits found
4226.2 Best-BlastP=> >nrprot 82% Identities = 578/861 (67%), Positives = 714/861 (82%), Gaps = 5/861 (0%) gb|AAB95117.11 DNA gyrase [Serratia marcescens] Length = 880
4227.1 Best-BlastP=> >nrprot 65% Identities = 238/489 (48%), Positives = 322/489 (65%), Gaps = 4/489 (0%) ref|NP_899921.11 glycerol kinase [Chromobacterium violaceum ATCC 12472] gb|AAQ57930.11 glycerol kinase [Chromobacterium violaceum ATCC 12472] Length = 500
4228.2 Best-BlastP=> >nrprot 64% Identities = 229/504 (45%), Positives = 327/504 (64%), Gaps = 15/504 (2%) ref |ZP_00091277.11 COG0578: Glycerol- 3-phosphate dehydrogenase [Azotobacter vinelandii] Length = 510
423.2 Best-BlastP=> >nrprot 47% Identities = 139/430 (32%), Positives = 204/430 (47%), Gaps = 28/430 (6%) ref|NP_407457.11 hypothetical protein [Yersinia pestis] pir||AD0489 hypothetical protein YPO4021 [imported] - Yersinia pestis (strain C092) emb|CAC93480.1 | hypothetical protein [Yersinia pestis C092] Length = 414
4231.2 Best-BlastP=> >nrprot 16% Identities = 49/205 (23%), Positives = 81/205 (39%), Gaps = 31/205 (15%) emb|CAE02991.1 | OSJNBa0043L09.10 [Oryza sativa (japonica cultivar-group)] Length = 687
4232.3 Best-BlastP=> >nrprot 42% Identities = 73/161 (45%), Positives = 103/161 (63%), Gaps = 1/161 (0%) ref|ZP_00088088.1 | COG0745: Response regulators consisting of a CheY-like receiver domain and a winged-helix DNA-binding domain [Pseudomonas fluorescens PfO-1] Length = 241
4233.3 Best-BlastP=> >nrprot No Hits found 4234.3 Best-BlastP=> >nrprot No Hits found
4236.3 Best-BlastP=> >nrprot 58% Identities = 257/678 (37%), Positives = 391/678 (57%), Gaps = 37/678 (5%) ref|ZP_00087890.11 hypothetical protein [Pseudomonas fluorescens PfO-1] Length = 701
4238.1 Best-BlastP=> >nrprot No Hits found
4239.4 Best-BlastP=> >nrprot No Hits found 424.1 Best-BlastP=> >nrprot 73% Identities = 63/103 (61 %), Positives = 76/103 (73%) ref|ZP_00082900.11 COG0662: Mannose-6-phosphate isomerase [Pseudomonas fluorescens PfO-1] Length = 103
4242.2 Best-BlastP=> >nrprot 34% Identities = 67/250 (26%), Positives = 112/250 (44%), Gaps = 18/250 (7%) ref|XP_340725.11 iron/ascorbate oxidoreductase family protein, putative [Trypanosoma brucei] gb|AAQ16084.11 iron/ascorbate oxidoreductase family protein, putative [Trypanosoma brucei] Length = 319
4243.2 Best-BlastP=> >nrprot 67% Identities = 76/154 (49%), Positives = 101/154 (65%), Gaps = 8/154 (5%) ref|NP_360925.1 | unknown [Rickettsia conorii] pir||H97860 hypothetical protein RC1288 [imported] - Rickettsia conorii (strain Malish 7) gb|AAL03826.11 unknown [Rickettsia conorii] Length = 154
4244.1 Best-BlastP=> >nrprot No Hits found
4246.2 Best-BlastP=> >nrprot 54% Identities = 95/280 (33%), Positives = 154/280 (55%), Gaps = 7/280 (2%) gb|AAM73852.1 |AF454863_1 putative lipase LipA [Legionella pneumophila] Length = 282
4247.2 Best-BlastP=> >nrprot 40% Identities = 63/264 (23%), Positives = 117/264 (44%), Gaps = 9/264 (3%) ref|ZP_00118032.11 COG1560: Lauroyl/myristoyl acyltransferase [Cytophaga hutchinsonii] Length = 307
4248.1 Best-BlastP=> >nrprot 52% Identities = 31/74 (41 %), Positives = 47/74 (63%), Gaps = 6/74 (8%) ref|ZP_00066809.11 COG1748: Saccharopine dehydrogenase and related proteins [Microbulbifer degradans 2-40] Length = 371
4249.1 Best-BlastP=> >nrprot 36% Identities = 89/354 (25%), Positives = 144/354 (40%), Gaps = 39/354 (11 %) gb|EAA22829.11 hypothetical protein [Plasmodium yoelii yoelii] Length = 2694
425.1 Best-BlastP=> >nrprot No Hits found
4250.1 Best-BlastP=> >nrprot No Hits found
4251.2 Best-BlastP=> >nrprot No Hits found
4253.1 Best-BlastP=> >nrprot 55% Identities = 173/457 (37%), Positives = 272/457 (59%), Gaps = 19/457 (4%) ref|NP_654426.11 aa_permeases, Amino acid permease [Bacillus anthracis A2012] ref|NP_843029.11 amino acid permease family protein [Bacillus anthracis str. Ames] gb|AAP24515.1 | amino acid permease family protein [Bacillus anthracis str. Ames] Length = 473
4254.2 Best-BlastP=> >nrprot No Hits found
4255.1 Best-BlastP=> >nrprot No Hits found
4259.2 Best-BlastP=> >nrprot 57% Identities = 104/292 (35%), Positives = 167/292 (57%), Gaps = 31/292 (10%) ref|ZP_00065076.1 | COG1766: Flagellar biosynthesis/type III secretory pathway lipoprotein [Microbulbifer degradans 2-40] Length = 556
426.3 Best-BlastP=> >nrprot No Hits found
4262.2 Best-BlastP=> >nrprot 47% Identities = 42/141 (29%), Positives = 72/141 (51%) ref|NP_249796.11 flagellar protein FliJ [Pseudomonas aeruginosa PA01] ref |ZP_00138693.11 COG2882: Flagellar biosynthesis chaperone [Pseudomonas aeruginosa UCBPP-PA14] pir||B83509 flagellar protein FliJ PA110'5 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG04494.1 |AE004540_14 flagellar protein FliJ [Pseudomonas aeruginosa PA01] Length = 147
4266.3 Best-BlastP=> >nrprot 35% Identities = 82/312 (26%), Positives = 145/312 (46%), Gaps = 51/312 (16%) gb|AAN63820.11 lysophospholipase A [Legionella pneumophila] Length = 309
4267.5 Best-BlastP=> >nrprot 68% Identities = 360/694 (51 %), Positives = 475/694 (68%), Gaps = 15/694 (2%) emb|CAA86935.11 polyphosphate kinase [Acinetobacter sp. ADP1] Length = 691
4269.3 Best-BlastP=> >nrprot No Hits found
427.3 Best-BlastP=> >nrprot 67% Identities = 127/239 (53%), Positives = 174/239 (72%), Gaps = 5/239 (2%) ref|NP_743000.11 RNA methyltransferase, TrmH family, group 1 [Pseudomonas putida KT2440] gb|AAN66464.1 |AE016275_9 RNA methyltransferase, TrmH family, group 1 [Pseudomonas putida KT2440] Length = 251
4272.3 Best-BlastP=> >nrprot No Hits found
4273.1 Best-BlastP=> >nrprot 35% Identities = 83/286 (29%), Positives = 133/286 (46%), Gaps = 2/286 (0%) ref|ZP_00065171.11 COG0583: Transcriptional regulator [Microbulbifer degradans 2-40] Length = 299
4275.2 Best-BlastP=> >nrprot 49% Identities = 77/244 (31 %), Positives = 120/244 (49%), Gaps = 19/244 (7%) ref|NP_661075.11 oxidoreductase, short- chain dehydrogenase/reductase family [Chlorobium tepidum TLS] gb|AAM71417.1 | oxidoreductase, short-chain dehydrogenase/reductase family [Chlorobium tepidum TLS] Length = 246
4276.2 Best-BlastP=> >nrprot 11 % Identities = 46/172 (26%), Positives = 73/172 (42%), Gaps = 36/172 (20%) dbj|BAC96628.11 conserved hypothetical protein [Vibrio vulnificus YJ016] Length = 442
428.1 Best-BlastP=> >nrprot 75% Identities = 142/257 (55%), Positives = 197/257 (76%) ref |NP_820132.11 inositol-1-monophosphatase [Coxiella burnetii RSA 493] gb|AAO90646.11 inositol-1-monophosphatase [Coxiella burnetii RSA 493] Length = 266
4281.2 Best-BlastP=> >nrprot 50% Identities = 32/88 (36%), Positives = 45/88 (51 %), Gaps = 4/88 (4%) ref|NP_441652.11 unknown protein [Synechocystis sp. PCC 6803] pir||S75873 hypothetical protein slr1163 - Synechocystis sp. (strain PCC 6803) dbj|BAA18332.11 ORF_ID:slr1163~unknown protein [Synechocystis sp. PCC 6803] Length = 556
4282.2 Best-BlastP=> >nrprot 67% Identities = 126/249 (50%), Positives = 169/249 (67%), Gaps = 2/249 (0%) gb|AAM51645.11 putative transposase [Francisella tularensis subsp. tularensis] Length = 247
4284.2 Best-BlastP=> >nrprot 41 % Identities = 100/266 (37%), Positives = 142/266 (53%), Gaps = 35/266 (13%) ref|NP_769986.11 bll3346 [Bradyrhizobium japonicum] dbj|BAC48611.11 bll3346 [Bradyrhizobium japonicum USDA 110] Length = 314
4285.1 Best-BlastP=> >nrprot No Hits found
4288.2 Best-BlastP=> >nrprot 98% Identities = 314/322 (97%), Positives = 318/322 (98%) gb| AAD43224.11 AF111940_6 LspK precursor [Legionella pneumophila] Length = 322
4289.2 Best-BlastP=> >nrprot 99% Identities = 203/205 (99%), Positives = 204/205 (99%) gb|AAD43223.1 |AF111940_5 LspJ precursor [Legionella pneumophila] Length = 205
429.2 Best-BlastP=> >nrprot 60% Identities = 149/315 (47%), Positives = 207/315 (65%), Gaps = 4/315 (1 %) ref|NP_793611.11 signal peptide peptidase SppA, 36K type [Pseudomonas syringae pv. tomato str. DC3000] gb|AAO57306.11 signal peptide peptidase SppA, 36K type [Pseudomonas syringae pv. tomato str. DC3000] Length = 332
4291.1 Best-BlastP=> >nrprot 98% Identities = 123/125 (98%), Positives = 124/125 (99%) gb|AAD43222.1 |AF111940_4 Lspl precursor [Legionella pneumophila] Length = 125
4293.2 Best-BlastP=> >nrprot 67% Identities = 116/253 (45%), Positives = 173/253 (68%), Gaps = 1/253 (0%) ref|NP_927916.11 hypothetical protein [Photorhabdus luminescens subsp. laumondii TT01] emb|CAE12861.1 | unnamed protein product [Photorhabdus luminescens subsp. laumondii TT01] Length = 258
4294.3 Best-BlastP=> >nrprot 57% Identities = 107/276 (38%), Positives = 161/276 (58%) ref|NP_720100.11 cell division ABC transporter, permease protein FtsX [Shewanella oneidensis MR-1] gb|AAN57544.1 |AE015890_5 cell division ABC transporter, permease protein FtsX [Shewanella oneidensis MR-1] Length = 321
4295.3 Best-BlastP=> >nrprot 69% Identities = 210/424 (49%), Positives = 301/424 (70%) ref|NP_820879.11 peptidase, M16 family [Coxiella burnetii RSA 493] gb|AA091393.11 peptidase, M16 family [Coxiella burnetii RSA 493] Length = 459
4296.1 Best-BlastP=> >nrprot 63% Identities = 164/388 (42%), Positives = 248/388 (63%) ref|NP_820878.11 peptidase, M16 family [Coxiella burnetii RSA 493] gb|AA091392.11 peptidase, M16 family [Coxiella burnetii RSA 493] Length = 443
4297.2 Best-BlastP=> >nrprot 64% Identities = 85/182 (46%), Positives = 117/182 (64%), Gaps = 6/182 (3%) ref|ZP_00134417.11 COG0742: N6- adenine-specific methylase [Actinobacillus pleuropneumoniae serovar 1 str. 4074] Length = 197
43.1 Best-BlastP=> >nrprot 97% Identities = 228/238 (95%), Positives = 233/238 (97%) gb|AAM08245.11 probable conjugal transfer protein [Legionella pneumophila] Length = 238
4301.1 Best-BlastP=> >nrprot 63% Identities = 130/287 (45%), Positives = 188/287 (65%), Gaps = 2/287 (0%) ref|XP_306575.11 ENSANGP00000014633 [Anopheles gambiae] gb|EAA02168.11 ENSANGP00000014633 [Anopheles gambiae str. PEST] Length = 304
4302.1 Best-BlastP=> >nrprot 81 % Identities = 190/271 (70%), Positives = 227/271 (83%) ref|NP_459218.11 2,3,4,5-tetrahydropyridine-2-carboxylate N- succinyltransferase [Salmonella typhimurium LT2] gb|AAL19177.1 | 2,3,4,5-tetrahydropyridine-2-carboxylate N-succinyltransferase [Salmonella typhimurium LT2] Length = 274
4303.1 Best-BlastP=> >nrprot 68% Identities = 203/374 (54%), Positives = 258/374 (68%), Gaps = 1/374 (0%) ref|ZP_00064957.11 COG0624: Acetylomithine deacetylase/Succinyl-diaminopimelate desuccinylase and related deacylases [Microbulbifer degradans 2-40] Length = 382
4305.2 Best-BlastP=> >nrprot No Hits found 4307.2 Best-BlastP=> >nrprot No Hits found 4309.2 Best-BlastP=> >nrprot 37% Identities = 79/320 (24%), Positives = 143/320 (44%), Gaps = 16/320 (5%) gb|AAC83363.11 outer membrane secretion protein Y [Pseudomonas alcaligenes] Length = 381
431.1 Best-BlastP=> >nrprot No Hits found 4310.1 Best-BlastP=> >nrprot 45% Identities = 34/139 (24%), Positives = 71/139 (51%), Gaps = 3/139 (2%) ref|NP_755574.1 | Putative general secretion pathway protein M-type yghD [Escherichia coli CFT073] gb|AAN82147.1 |AE016766_235 Putative general secretion pathway protein M-type yghD [Escherichia coli CFT073] Length = 178
4311.1 Best-BlastP=> >nrprot 41 % Identities = 47/173 (27%), Positives = 74/173 (42%), Gaps = 14/173 (8%) ref|NP_232963.11 DamX-related protein [Vibrio cholerae 01 biovar eltor str. N16961] pir||B82443 DamX-related protein VCA0573 [imported] - Vibrio cholerae (strain N16961 serogroup 01 ) gb|AAF96475.1 | DamX-related protein [Vibrio cholerae 01 biovar eltor str. N16961] Length = 195
4312.2 Best-BlastP=> >nrprot 57% Identities = 186/432 (43%), Positives = 253/432 (58%), Gaps = 8/432 (1 %) ref|NP_231613.11 deoxyguanosinetriphosphate triphosphohydrolase [Vibrio cholerae 01 biovar eltor str. N16961 ] sp|Q9KQL9|DG1A_VIBCH Deoxyguanosinetriphosphate triphosphohydrolase-like protein 1 pir||B82132 deoxyguanosinetriphosphate triphosphohydrolase VC1979 [imported] Vibrio cholerae (strain N 16961 serogroup 01 ) gb|AAF95127.1 | deoxyguanosinetriphosphate triphosphohydrolase [Vibrio cholerae 01 biovar eltor str. N16 61 ] Length = 441
4316.4 Best-BlastP=> >nrprot 23% Identities = 87/391 (22%), Positives = 154/391 (39%), Gaps = 61/391 (15%) gb|AAC21558.11 paramyosin related protein [Echinococcus granulosus] Length = 601
4319.2 Best-BlastP=> >nrprot 38% Identities = 56/226 (24%), Positives = 95/226 (42%), Gaps = 24/226 (10%) ref|NP_820254.11 ompA-like transmembrane domain protein [Coxiella burnetii RSA 493] gb|AAO90768.1 | ompA-like transmembrane domain protein [Coxiella burnetii RSA 493] Length = 248
432.1 Best-BlastP=> >nrprot No Hits found
4320.1 Best-BlastP=> >nrprot 61 % Identities = 163/420 (38%), Positives = 265/420 (63%), Gaps = 4/420 (0%) ref|NP_249285.11 peptidyl-prolyl cis-trans isomerase SurA [Pseudomonas aeruginosa PA01] pir||B83572 peptidyl-prolyl cis-trans isomerase SurA PA0594 [imported] - Pseudomonas aeruginosa (strain PA01) gb|AAG03983.1 |AE004495_7 peptidyl-prolyl cis-trans isomerase SurA [Pseudomonas aeruginosa PA01] Length = 430
4321.2 Best-BlastP=> >nrprot 48% Identities = 274/816 (33%), Positives = 410/816 (50%), Gaps = 61/816 (7%) ref|NP_820953.11 organic solvent tolerance protein [Coxiella burnetii RSA 493] gb|AA091467.11 organic solvent tolerance protein [Coxiella burnetii RSA 493] Length = 870
4322.1 Best-BlastP=> >nrprot 57% Identities = 133/323 (41 %), Positives = 187/323 (57%), Gaps = 9/323 (2%) ref|NP_840285.11 Domain of unknown function DUF227 [Nitrosomonas europaea ATCC 19718] emb|CAD84102.11 Domain of unknown function DUF227 [Nitrosomonas europaea ATCC 19718] Length = 332
4323.1 Best-BlastP=> >nrprot 65% Identities = 105/218 (48%), Positives = 145/218 (66%), Gaps = 4/218 (1 %) ref|ZP_00126866.11 COG1208: Nucleoside-diphosphate-sugar pyrophosphorylase involved in lipopolysaccharide biosynthesis/translation initiation factor 2B, gamma/epsilon subunits (elF-2Bgamma/elF-2Bepsilon) [Pseudomonas syringae pv. syringae B728a] Length = 223
4325.3 Best-BlastP=> >nrprot 66% Identities = 154/307 (50%), Positives = 205/307 (66%), Gaps = 1/307 (0%) ref|NP_462824.1 | porphobilinogen deaminase (hydroxymethylbilane synthase) [Salmonella typhimurium LT2] gb|AAF33453.11 89% identity with E. coli porphobilinogen deaminase (HEMC) (SP.P06983); contains similarity to Pfam family PF01379 (Porphobilinogen deaminase), score=627.8, E=6.2e- 185, N=1 [Salmonella typhimurium LT2] gb|AAL22783.1 | porphobilinogen deaminase [Salmonella typhimurium LT2] Length = 318
4326.1 Best-BlastP=> >nrprot 50% Identities = 75/233 (32%), Positives = 127/233 (54%), Gaps = 3/233 (1 %) dbj|BAC92844.1 | uroporphyrinogen-lll synthase [Vibrio vulnificus YJ016] Length = 260
4327.1 Best-BlastP=> >nrprot 40% Identities = 86/318 (27%), Positives = 151/318 (47%), Gaps = 26/318 (8%) ref|ZP_00065835.11 COG2g59: Uncharacterized enzyme of heme biosynthesis [Microbulbifer degradans 2-40] Length = 494
4328.1 Best-BlastP=> >nrprot 56% Identities = 139/389 (35%), Positives = 224/389 (57%), Gaps = 3/389 (0%) ref|NP_821051.11 hemY protein [Coxiella burnetii RSA 493] gb|AA091565.11 hemY protein [Coxiella burnetii RSA 493] Length = 392
433.1 Best-BlastP=> >nrprot 99% Identities = 1007/1009 (99%), Positives = 1008/1009 (99%) pir||T18339 icmB protein - Legionella pneumophila emb|CAA75170.1 | IcmB protein [Legionella pneumophila] gb|AAC38183.1 | DotO [Legionella pneumophila] emb|CAA75336.1 | IcmB protein [Legionella pneumophila] Length = 1009
4332.1 Best-BlastP=> >nrprot 59% Identities = 82/151 (54%), Positives = 107/151 (70%) ref|ZP_00024252.11 COG0412: Dienelactone hydrolase and related enzymes [Ralstonia metallidurans] Length = 435
4333.1 Best-BlastP=> >nrprot 61 % Identities = 128/282 (45%), Positives = 174/282 (61%), Gaps = 4/282 (1 %) ref|ZP_00077190.11 COG0454: Histone acetyltransferase HPA2 and related acetyltransferases [Methanosarcina barkeri] Length = 286
4334.1 Best-BlastP=> >nrprot 62% Identities = 135/293 (46%), Positives = 187/293 (63%), Gaps = 6/293 (2%) ref|NP_421406.1 | conserved hypothetical protein [Caulobacter crescentus CB15] pir||B87572 conserved hypothetical protein CC2605 [imported] - Caulobacter crescentus gb|AAK24574.11 conserved hypothetical protein [Caulobacter crescentus CB15] Length = 304
4336.2 Best-BlastP=> >nrprot 59% Identities = 163/401 (40%), Positives = 242/401 (60%), Gaps = 1/401 (0%) ref|NP_626512.1 | putative integral membrane protein. [Streptomyces coelicolor A3(2)] pir||T50573 probable integral membrane protein [imported] - Streptomyces coelicolor emb|CAB61710.11 putative integral membrane protein. [Streptomyces coelicolor A3(2)] Length = 431
4337.3 Best-BlastP=> >nrprot 57% Identities = 36/63 (57%), Positives = 48/63 (76%), Gaps = 2/63 (3%) ref|NP_799393.11 putative signal peptide protein [Vibrio parahaemolyticus RIMD 2210633] dbj|BAC61277.11 putative signal peptide protein [Vibrio parahaemolyticus] Length = 86
4338.3 Best-BlastP=> >nrprot 68% Identities = 147/290 (50%), Positives = 206/290 (71%), Gaps = 1/290 (0%) ref|NP_781198.1 | myo-inositol catabolism protein iolE [Clostridium tetani E88] gb|AA035135.1 | myo-inositol catabolism protein iolE [Clostridium tetani E88] Length = 298
4339.1 Best-BlastP=> >nrprot 72% Identities = 347/644 (53%), Positives = 451/644 (70%), Gaps = 25/644 (3%) ref|ZP_00131855.11 COG3962: Acetolactate synthase [Haemophilus somnus 2336] Length = 645
434.3 Best-BlastP=> >nrprot 99% Identities = 207/208 (99%), Positives = 207/208 (99%) pir||T18338 icmJ protein - Legionella pneumophila emb|CAA75169.1 | IcmJ protein [Legionella pneumophila] gb|AAC38184.1 | DotN [Legionella pneumophila] emb|CAA75335.1 | IcmJ protein [Legionella pneumophila] Length = 208
4340.2 Best-BlastP=> >nrprot 36% Identities = 179/356 (50%), Positives = 232/356 (65%), Gaps = 4/356 (1 %) ref|ZP_00122182.11 COG3892: Uncharacterized protein conserved in bacteria [Haemophilus somnus 129PT] Length = 636
4341.3 Best-BlastP=> >nrprot No Hits found 4342.1 Best-BlastP=> >nrprot 53% Identities = 29/77 (37%), Positives = 44/77 (57%) ref|NP_289764.11 orf, hypothetical protein [Escherichia coli 0157:H7 EDL933] ref|NP_312096.11 hypothetical protein [Escherichia coli 0157:H7] ref|NP_417657.11 hypothetical protein [Escherichia coli K12] ref|NP_708989.1 | orf, conserved hypothetical protein [Shigella flexneri 2a str. 301] ref|NP_755814.1 | Protein yrbA [Escherichia coli CFT073] ref|NP_838699.11 hypothetical protein [Shigella flexneri 2a str. 2457T] pir||H65109 hypothetical 9.5 kD protein in murZ-rpoN intergenic region - Escherichia coli (strain K-12) pir||E91137 hypothetical protein ECs4069 [imported] - Escherichia coli (strain 0157:H7, substrain RIMD 0509952) pir||H85982 hypothetical protein yrbA [imported] - Escherichia coli (strain 0157:H7, substrain EDL933) gb|AAA57991.1 | ORF 89 [Escherichia coli] gb|AAC76222.11 orf, hypothetical protein [Escherichia coli K12] gb|AAF21251.1 |AF053073_4 YrbA [Shigella flexneri] gb|AAG58324.1 |AE005547_10 orf, hypothetical protein [Escherichia coli 0157:H7 EDL933] dbj|BAB37492.1 | hypothetical protein [Escherichia coli 0157:H7] gb|AAN44696.1 |AE015334_10 orf, conserved hypothetical protein [Shigella flexneri 2a str. 301] gb|AAN82388.1 |AE016767_148 F
4344.1 Best-BlastP=> >nrprot 80% Identities = 272/419 (64%), Positives = 339/419 (80%) ref|NP_794195.11 UDP-N-acetylglucosamine 1 - carboxyvinyltransferase [Pseudomonas syringae pv. tomato str. DC3000] gb|AAO57890.1 | UDP-N-acetylglucosamine 1-carboxyvinyltransferase [Pseudomonas syringae pv. tomato str. DC3000] Length = 421
4345.1 Best-BlastP=> >nrprot 71% Identities = 136/252 (53%), Positives = 182/252 (72%), Gaps = 3/252 (1 %) ref|NP_718206.11 conserved hypothetical protein TIGR00486 [Shewanella oneidensis MR-1] gb|AAN55650.1 |AE015704_1 conserved hypothetical protein TIGR00486 [Shewanella oneidensis MR-1] Length = 250
4346.2 Best-BlastP=> >nrprot 71% Identities = 153/286 (53%), Positives = 207/286 (72%), Gaps = 1/286 (0%) ref|NP_249701.11 dihydrodipicolinate synthase [Pseudomonas aeruginosa PA01] sp|Q9l4W3|DAPA_PSEAE Dihydrodipicolinate synthase (DHDPS) pir||C83520 dihydrodipicolinate synthase PA1010 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG04399.1 |AE004533_10 dihydrodipicolinate synthase [Pseudomonas aeruginosa PA01] Length = 292
4347.2 Best-BlastP=> >nrprot 44% Identities = 31/68 (45%), Positives = 38/68 (55%), Gaps = 7/68 (10%) ref|ZP_00035058.11 COG2885: Outer membrane protein and related peptidoglycan-associated (lipo)proteins [Burkholderia fungorum] Length = 237
4349.2 Best-BlastP=> >nrprot 81 % Identities = 180/260 (69%), Positives = 213/260 (81 %) ref|NP_900365.11 3-hydroxybutyrate dehydrogenase [Chromobacterium violaceum ATCC 12472] gb|AAQ58371.11 3-hydroxybutyrate dehydrogenase [Chromobacterium violaceum ATCC 12472] Length = 260
4350.2 Best-BlastP=> >nrprot 59% Identities = 152/375 (40%), Positives = 230/375 (61 %), Gaps = 4/375 (1 %) ref|ZP_00008996.11 COG1752: Predicted esterase of the alpha-beta hydrolase superfamily [Rhodopseudomonas palustris] Length = 37g
4351.2 Best-BlastP=> >nrprot 73% Identities = 161/286 (56%), Positives = 221/286 (77%) ref|NP_522970.11 PROBABLE CHEMOTAXIS (MOTILITY PROTEIN A) TRANSMEMBRANE [Ralstonia solanacearum] emb|CAD18562.11 PROBABLE CHEMOTAXIS (MOTILITY PROTEIN A) TRANSMEMBRANE [Ralstonia solanacearum] Length = 286
4352.3 Best-BlastP=> >nrprot 61 % Identities = 140/284 (49%), Positives = 191/284 (67%), Gaps = 1/284 (0%) ref|NP_840146.1 | Bacterial outer membrane protein [Nitrosomonas europaea ATCC 19718] emb|CAD83956.11 Bacterial outer membrane protein [Nitrosomonas europaea ATCC 19718] Length = 307
4353.2 Best-BlastP=> >nrprot No Hits found
4354.2 Best-BlastP=> >nrprot No Hits found
4355.2 Best-BlastP=> >nrprot 64% Identities = 161/310 (51 %), Positives = 204/310 (65%), Gaps = 5/310 (1 %) ref|NP_438574.11 hypothetical protein [Haemophilus influenzae Rd] sp|P44433|RLUC_HAEIN Ribosomal large subunit pseudouridine synthase C (Pseudouridylate synthase) (Uracil hydrolyase) pir||G64151 hypothetical protein HI0412 - Haemophilus influenzae (strain Rd KW20) gb|AAC22071.1 | conserved hypothetical protein [Haemophilus influenzae Rd] Length = 322
4356.2 Best-BlastP=> >nrprot 52% Identities = 77/215 (35%), Positives = 116/215 (53%), Gaps = 8/215 (3%) ref|ZP_00013688.11 hypothetical protein [Rhodospirillum rubrum] Length = 236
4357.3 Best-BlastP=> >nrprot 98% Identities = 414/424 (97%), Positives = 420/424 (99%) gb|AAM00604.11 putative histidine kinase [Legionella pneumophila] Length = 424
4358.2 Best-BlastP=> >nrprot 49% Identities = 64/187 (34%), Positives = 91 /187 (48%), Gaps = 7/187 (3%) ref|NP_761114.11 Guanylate cyclase-related protein [Vibrio vulnificus CMCP6] gb|AAO10641.1 |AE016804_151 Guanylate cyclase-related protein [Vibrio vulnificus CMCP6] dbj|BAC94846.1 | guanylate cyclase-related protein [Vibrio vulnificus YJ016] Length = 185
436.4 Best-BlastP=> >nrprot No Hits found
4363.2 Best-BlastP=> >nrprot 43% Identities = 68/183 (37%), Positives = 106/183 (57%), Gaps = 2/183 (1 %) ref|ZP_00073054.11 COG3555: Aspartyl/asparaginyl beta-hydroxylase and related dioxygenases [Trichodesmium erythraeum IMS101] Length = 283
4364.2 Best-BlastP=> >nrprot 41 % Identities = 36/77 (46%), Positives = 44/77 (57%) pir||D72548 hypothetical protein APE1672 - Aeropyrum pernix (strain K1 ) dbj|BAA80673.11 113aa long hypothetical protein [Aeropyrum pernix] Length = 113
4365.4 Best-BlastP=> >nrprot 63% Identities = 78/175 (44%), Positives = 115/175 (65%), Gaps = 6/175 (3%) ref|ZP_00021197.11 COG0262: Dihydrofolate reductase [Ralstonia metallidurans] Length = 177
4366.1 Best-BlastP=> >nrprot 47% Identities = 37/123 (30%), Positives = 66/123 (53%), Gaps = 4/123 (3%) ref|NP_800262.11 glutathione S-transfersae- related protein [Vibrio parahaemolyticus RIMD 2210633] dbj|BAC62095.1 | glutathione S-transfersae-related protein [Vibrio parahaemolyticus] Length = 128
4367.1 Best-BlastP=> >nrprot No Hits found
4369.1 Best-BlastP=> >nrprot 35% Identities = 53/153 (34%), Positives = 79/153 (51 %), Gaps = 2/153 (1 %) ref|ZP_00026335.1 | COG3714: Predicted membrane protein [Ralstonia metallidurans] Length = 233
4370.1 Best-BlastP=> >nrprot No Hits found
4371.1 Best-BlastP=> >nrprot 57% Identities = 122/252 (48%), Positives = 171/252 (67%), Gaps = 4/252 (1%) ref|NP_819457.1 | polysaccharide deacetylase-related protein [Coxiella burnetii RSA 493] gb|AA089971.1 | polysaccharide deacetylase-related protein [Coxiella burnetii RSA 493] Length = 276
4372.3 Best-BlastP=> >nrprot 61% Identities = 167/391 (42%), Positives = 247/391 (63%), Gaps = 5/391 (1 %) ref|NP_820110.11 membrane-bound lytic murein transglycosylase family protein [Coxiella burnetii RSA 493] gb|AAO90624.11 membrane-bound lytic murein transglycosylase family protein [Coxiella burnetii RSA 493] Length = 400
4373.3 Best-BlastP=> >nrprot 64% Identities = 45/105 (42%), Positives = 68/105 (64%), Gaps = 7/105 (6%) ref|NP_800642.11 hypothetical protein VPA1132 [Vibrio parahaemolyticus RIMD 2210633] dbj|BAC62475.11 hypothetical protein [Vibrio parahaemolyticus] Length = 108
4374.4 Best-BlastP=> >nrprot 68% Identities = 262/502 (52%), Positives = 358/502 (71 %) ref |NP_819429.11 integral membrane protein MviN [Coxiella burnetii RSA 493] gb|AA089943.11 integral membrane protein MviN [Coxiella burnetii RSA 493] Length = 515
4376.3 Best-BlastP=> >nrprot 63% Identities = 105/215 (48%), Positives = 143/215 (66%) pdb|1AZO| Dna Mismatch Repair Protein Muth From E. Coli Length = 232
4377.4 Best-BlastP=> >nrprot No Hits found
4378.2 Best-BlastP=> >nrprot 75% Identities = 135/205 (65%), Positives = 167/205 (81%), Gaps = 1/205 (0%) ref|NP_799538.1 | antibiotic acetyltransferase [Vibrio parahaemolyticus RIMD 2210633] dbj|BAC61371.11 antibiotic acetyltransferase [Vibrio parahaemolyticus] Length = 212
4379.1 Best-BlastP=> >nrprot 40% Identities = 85/221 (38%), Positives = 120/221 (54%), Gaps = 8/221 (3%) ref|ZP_00108724.11 COG3393: Predicted acetyltransferase [Nostoc punctiforme] Length = 222
4381 .2 Best-BlastP=> >nrprot 79% Identities = 276/420 (65%), Positives = 340/420 (80%) ref|NP_819962.11 amino acid permease family protein [Coxiella burnetii RSA 493] gb|AAO90476.11 amino acid permease family protein [Coxiella burnetii RSA 493] Length = 437
4383.2 Best-BlastP=> >nrprot 37% Identities = 83/190 (43%), Positives = 109/190 (57%), Gaps = 3/190 (1 %) ref|NP_435788.11 hypothetical protein [Sinorhizobium meliloti] pir||F95329 hypothetical protein SMa1005 [imported] - Sinorhizobium meliloti (strain 1021 ) magaplasmid pSymA gb|AAK65200.11 hypothetical protein [Sinorhizobium meliloti] Length = 266
4384.2 Best-BlastP=> >nrprot 55% Identities = 171/477 (35%), Positives = 267/477 (55%), Gaps = 9/477 (1 %) ref |NP_819538.11 proton/peptide symporter family protein [Coxiella burnetii RSA 493] gb|AAO90052.11 proton/peptide symporter family protein [Coxiella burnetii RSA 493] Length = 492
4385.1 Best-BlastP=> >nrprot 85% Identities = 411/546 (75%), Positives = 474/546 (86%), Gaps = 3/546 (0%) ref|NP_230848.11 urocanate hydratase [Vibrio cholerae 01 biovar eltor str. N16961 ] sp|Q9KSQ3|HUTU_VIBCH Urocanate hydratase (Urocanase) (Imidazolonepropionate hydrolase) pir||F82228 urocanate hydratase VC1203 [imported] - Vibrio cholerae (strain N16961 serogroup 01 ) gb|AAF94362.11 urocanate hydratase [Vibrio cholerae 01 biovar eltor str. N16961] Length = 565
4386.2 Best-BlastP=> >nrprot 73% Identities = 296/486 (60%), Positives = 375/486 (77%), Gaps = 1/486 (0%) ref|NP_230847.11 histidine ammonia- lyase [Vibrio cholerae 01 biovar eltor str. N16961] sp|Q9KSQ4|HUTH_VIBCH Histidine ammonia-lyase (Histidase) pir||E82228 histidine ammonia-lyase (EC 4.3.1.3) [similarity] - Vibrio cholerae (strain N16961 serogroup 01 ) gb|AAF94361.1 | histidine ammonia-lyase [Vibrio cholerae 01 biovar eltor str. N16961 ] Length = 511
4387.3 Best-BlastP=> >nrprot 62% Identities = 96/190 (50%), Positives = 128/190 (67%), Gaps = 1/190 (0%) ref |NP_246159.11 Dam [Pasteurella multocida] gb|AAK03306.1 | Dam [Pasteurella multocida] gb|AAL05864.1 |AF411317_2 DNA adenine methylase [Pasteurella multocida] Length = 301
4388.1 Best-BlastP=> >nrprot 54% Identities = 151/402 (37%), Positives = 232/402 (57%), Gaps = 5/402 (1%) ref|NP_819596.11 major facilitator family transporter [Coxiella burnetii RSA 493] gb|AAO90110.11 major facilitator family transporter [Coxiella burnetii RSA 493] Length = 421
4389.2 Best-BlastP=> >nrprot 62% Identities = 206/405 (50%), Positives = 270/405 (66%), Gaps = 8/405 (1 %) ref|NP_820255.11 D-alanyl-D-alanine carboxypeptidase [Coxiella burnetii RSA 493] gb|AAO90769.11 D-alanyl-D-alanine carboxypeptidase [Coxiella burnetii RSA 493] Length = 418 4391.1 Best-BlastP=> >nrprot 98% Identities = 270/278 (97%), Positives = 275/278 (98%) emb|CAD90964.11 putative D-Ala-amino transferase [Legionella pneumophila] Length = 278
4392.1 Best-BlastP=> >nrprot 60% Identities = 35/87 (40%), Positives = 53/87 (60%) ref|NP_841528.11 conserved hypothetical protein [Nitrosomonas europaea ATCC 19718] sp|Q82UJ7|YE87_NITEU Hypothetical UPF0250 protein NE1487 emb|CAD85398.1 | conserved hypothetical protein [Nitrosomonas europaea ATCC 19718] Length = 87
4393.1 Best-BlastP=> >nrprot 98% Identities = 192/199 (96%), Positives = 196/199 (98%) emb|CAD90955.11 LssX protein [Legionella pneumophila] Length = 199
4394.4 Best-BlastP=> >nrprot 98% Identities = 666/678 (98%), Positives = 672/678 (99%) emb|CAD90962.11 LssY protein [Legionella pneumophila] Length = 678
4395.2 Best-BlastP=> >nrprot 63% Identities = 337/723 (46%), Positives = 453/723 (62%), Gaps = 27/723 (3%) ref|NP_251738.11 conserved hypothetical protein [Pseudomonas aeruginosa PA01] pir||A83266 conserved hypothetical protein PA3048 [imported] - Pseudomonas aeruginosa (strain PA01) gb|AAG06436.1 |AE004729_10 conserved hypothetical protein [Pseudomonas aeruginosa PA01] Length = 725
4396.1 Best-BlastP=> >nrprot 59% Identities = 66/157 (42%), Positives = 95/157 (60%) ref|ZP_00090069.11 COG3028: Uncharacterized protein conserved in bacteria [Azotobacter vinelandii] Length = 172
4398.2 Best-BlastP=> >nrprot 48% Identities = 70/252 (27%), Positives = 135/252 (53%), Gaps = 4/252 (1 %) ref|NP_784615.11 integral membrane protein [Lactobacillus plantarum WCFS1] emb|CAD63460.11 integral membrane protein [Lactobacillus plantarum WCFS1] Length = 293
44.1 Best-BlastP=> >nrprot 97% Identities = 335/346 (96%), Positives = 339/346 (97%) gb|AAM08244.11 probable conjugal transfer protein [Legionella pneumophila] Length = 346
440.4 Best-BlastP=> >nrprot 14% Identities = 134/534 (25%), Positives = 215/534 (40%), Gaps = 106/534 (19%) pir||JC6009 surface-located membrane protein Imp3 precursor - Mycoplasma hominis emb|CAA64858.1 | Lmp3 protein [Mycoplasma hominis] Length = 1302
4402.2 Best-BlastP=> >nrprot 65% Identities = 83/174 (47%), Positives = 119/174 (68%), Gaps = 2/174 (1 %) ref|NP_930722.11 hypothetical protein [Photorhabdus luminescens subsp. laumondii TT01] emb|CAE15878.11 unnamed protein product [Photorhabdus luminescens subsp. laumondii TT01] Length = 176
4403.2 Best-BlastP=> >nrprot 60% Identities = 124/258 (48%), Positives = 160/258 (62%) ref|NP_252268.11 conserved hypothetical protein [Pseudomonas aeruginosa PA01] sp|Q9HY42|YZ78_PSEAE Hypothetical protein PA3578 pir||E83199 conserved hypothetical protein PA3578 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG06966.1 |AE004778_10 conserved hypothetical protein [Pseudomonas aeruginosa PA01] Length = 261
4404.1 Best-BlastP=> >nrprot 49% Identities = 51/124 (41 %), Positives = 69/124 (55%) ref| NP_105863.1 | transcriptional regulator [Mesorhizobium loti] dbj|BAB5164g.11 transcriptional regulator [Mesorhizobium loti] Length = 143
4405.1 Best-BlastP=> >nrprot 73% Identities = 77/148 (52%), Positives = 112/148 (75%) ref|NP_662760.11 conserved hypothetical protein [Chlorobium tepidum TLS] gb|AAM73102.1 | conserved hypothetical protein [Chlorobium tepidum TLS] Length = 155
4406.1 Best-BlastP=> >nrprot 63% Identities = 78/187 (41 %), Positives = 121/187 (64%), Gaps = 1/187 (0%) ref|ZP_00092584.1 | COG1278: Cold shock proteins [Azotobacter vinelandii] Length = 333
4407.1 Best-BlastP=> >nrprot 65% Identities = 39/69 (56%), Positives = 51/69 (73%) ref|NP_743260.11 cold-shock domain family protein [Pseudomonas putida KT2440] gb|AAN66724.1 |AE016300_9 cold-shock domain family protein [Pseudomonas putida KT2440] Length = 69
4408.1 Best-BlastP=> >nrprot No Hits found
4409.1 Best-BlastP=> >nrprot 71 % Identities = 120/212 (56%), Positives = 150/212 (70%), Gaps = 3/212 (1 %) ref|NP_403711.11 orotate phosphoribosyltransferase [Yersinia pestis] ref|NP_667439.1 | orotate phosphoribosyltransferase [Yersinia pestis KIM] sp|Q8ZJP7|PYRE_YERPE Orotate phosphoribosyltransferase (OPRT) (OPRTase) pir||AF0006 orotate phosphoribosyltransferase (EC 2.4.2.10) [imported] - Yersinia pestis (strain C092) emb|CAC88912.11 orotate phosphoribosyltransferase [Yersinia pestis C092] gb|AAM83690.1 |AE013610_2 orotate phosphoribosyltransferase [Yersinia pestis KIM] Length = 215
4415.3 Best-BlastP=> >nrprot 55% Identities = 157/417 (37%), Positives = 239/417 (57%), Gaps = 8/417 (1 %) ref|ZP_00034486.11 COG0642: Signal transduction histidine kinase [Burkholderia fungorum] Length = 479
4419.2 Best-BlastP=> >nrprot 72% Identities = 161/317 (50%), Positives = 233/317 (73%) ref|NP_794895.1 | proline iminopeptidase [Pseudomonas syringae pv. tomato str. DC3000] gb|AAO58590.11 proline iminopeptidase [Pseudomonas syringae pv. tomato str. DC3000] Length = 323
4421.2 Best-BlastP=> >nrprot 41% Identities = 77/254 (30%), Positives = 111/254 (43%), Gaps = 51/254 (20%) ref|NP_052363.11 unnamed protein product [Coxiella burnetii] ref|NP_819024.11 hypothetical protein [Coxiella burnetii RSA 493] pir||S38245 hypothetical protein - Coxiella burnetii emb|CAA53133.1 | unnamed protein product [Coxiella burnetii] gb|AA091584.11 hypothetical protein [Coxiella burnetii RSA 493] Length = 341
4422.1 Best-BiastP=> >nrprot No Hits found
4424.1 Best-BlastP=> >nrprot No Hits found
4425.1 Best-BlastP=> >nrprot 46% Identities = 57/160 (35%), Positives = 80/160 (50%), Gaps = 3/160 (1 %) ref|NP_105070.11 hypothetical protein, acetyltransferase, putative [Mesorhizobium loti] dbj|BAB50856.1 | hypothetical protein [Mesorhizobium loti] Length = 168
4427.1 Best-BlastP=> >nrprot No Hits found
4428.1 Best-BlastP=> >nrprot No Hits found
4429.2 Best-BlastP=> >nrprot No Hits found 443.3 Best-BlastP=> >nrprot 99% Identities = 190/191 (99%), Positives = 191/191 (100%) pir||T18327 icmQ protein - Legionella pneumophila emb|CAA73239.1 | IcmQ protein [Legionella pneumophila] emb|CAA75324.1 | IcmQ protein [Legionella pneumophila] Length = 191
4431.2 Best-BlastP=> >nrprot 14% Identities = 61/293 (20%), Positives = 128/293 (43%), Gaps = 33/293 (11%) dbj|BAC86266.11 unnamed protein product [Homo sapiens] Length = 486
4432.2 Best-BlastP=> >nrprot No Hits found 4433.1 Best-BlastP=> >nrprot 42% Identities = 64/200 (32%), Positives = 86/200 (43%), Gaps = 12/200 (6%) ref|NP_799850.11 putative tellurite resistance protein-related protein [Vibrio parahaemolyticus RIMD 2210633] dbj|BAC61683.11 putative tellurite resistance protein-related protein [Vibrio parahaemolyticus] Length = 195
4434.1 Best-BlastP=> >nrprot 34% Identities = 52/215 (24%), Positives = 91/215 (42%), Gaps = 23/215 (10%) ref|NP_788733.1 | multiple ankyrin repeats single KH domain CG33106-PA [Drosophila melanogaster] ref|NP_788734.11 multiple ankyrin repeats single KH domain CG33106- PB [Drosophila melanogaster] gb|AAO41600.11 CG33106-PA [Drosophila melanogaster] gb|AAO41601.11 CG33106-PB [Drosophila melanogaster] Length = 4001
4435.4 Best-BlastP=> >nrprot 13% Identities = 50/196 (25%), Positives = 92/196 (46%), Gaps = 32/196 (16%) ref |NP_001804.11 centromere protein E; Centromere autoantigen E (312kD); centromere protein E (312kD); kinesin family member 10 [Homo sapiens] sp|Q02224|CENE_HUMAN Centromeric protein E (CENP-E protein) pir||S28261 centromere protein E - human emb|CAA78727.1 | CENP-E [Homo sapiens] Length = 2663
444.2 Best-BlastP=> >nrprot 96% Identities = 116/120 (96%), Positives = 117/120 (97%) pir||T18326 icmR protein - Legionella pneumophila emb|CAA73238.11 IcmR protein [Legionella pneumophila] emb|CAA75323.11 IcmR protein [Legionella pneumophila] Length = 120
4440.1 Best-BlastP=> >nrprot 73% Identities = 71/144 (49%), Positives = 104/144 (72%), Gaps = 4/144 (2%) ref|ZP_00065054.11 COG1558: Flagellar basal body rod protein [Microbulbifer degradans 2-40] Length = 148
4441.1 Best-BlastP=> >nrprot 61 % Identities = 61/132 (46%), Positives = 80/132 (60%), Gaps = 1/132 (0%) gb|AAN08637.11 FlgB [Aeromonas hydrophila] Length = 132
4442.1 Best-BlastP=> >nrprot 75% Identities = 202/297 (68%), Positives = 235/297 (79%) ref|NP_928683.11 oxygen-dependent coproporphyrinogen III oxidase [Photorhabdus luminescens subsp. laumondii TT01] emb|CAE13676.11 oxygen-dependent coproporphyrinogen III oxidase [Photorhabdus luminescens subsp. laumondii TT01] Length = 302
4444.2 Best-BlastP=> >nrprot 57% Identities = 168/420 (40%), Positives = 249/420 (59%), Gaps = 12/420 (2%) ref|ZP_00030428.11 COG0477: Permeases of the major facilitator superfamily [Burkholderia fungorum] Length = 645
4452.2 Best-BlastP=> >nrprot 34% Identities = 127/238 (53%), Positives = 161/238 (67%) ref |ZP_00112372.11 COG2114: Adenylate cyclase, family 3 (some proteins contain HAMP domain) [Nostoc punctiforme] Length = 616
4453.2 Best-BlastP=> >nrprot 41 % Identities = 97/469 (20%), Positives = 194/469 (41 %), Gaps = 60/469 (12%) gb|EAA15516.11 hypothetical protein [Plasmodium yoelii yoelii] Length = 585
4454.4 Best-BlastP=> >nrprot 87% Identities = 258/329 (78%), Positives = 290/329 (88%) gb|AAB58447.11 spectinomycin phosphotransferase [Legionella pneumophila] Length = 331
4457.2 Best-BlastP=> >nrprot 51 % Identities = 102/334 (30%), Positives = 174/334 (52%), Gaps = 14/334 (4%) ref|NP_249522.1 | transcriptional regulator OruR [Pseudomonas aeruginosa PA01] ref|ZP_00138425.11 COG2207: AraC-type DNA-binding domain-containing proteins [Pseudomonas aeruginosa UCBPP-PA14] sp|P72171 |ORUR_PSEAE Ornithine utilization regulator pir||G83540 transcription regulator OruR PA0831 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAB94774.1 | OruR [Pseudomonas aeruginosa]
gb|AAG04220.1 |AE004518_1 transcriptional regulator OruR [Pseudomonas aeruginosa PA01] Length = 339
4462.3 Best-BlastP=> >nrprot 71 % Identities = 115/192 (59%), Positives = 145/192 (75%) ref|ZP_00067991.1 | COG0088: Ribosomal protein L4 [Microbulbifer degradans 2-40] Length = 204
4463.2 Best-BlastP=> >nrprot 80% Identities = 138/209 (66%), Positives = 175/209 (83%), Gaps = 1/209 (0%) ref|NP_403860.11 50S ribosomal protein L3 [Yersinia pestis] ref | N P_671283.11 50S ribosomal subunit protein L3 [Yersinia pestis KIM] pir||AB0026 50S ribosomal protein L3 [imported] - Yersinia pestis (strain C092) emb|CAC89069.11 50S ribosomal protein L3 [Yersinia pestis C092] gb|AAM87534.1 |AE014002_7 50S ribosomal subunit protein L3 [Yersinia pestis KIM] Length = 209
4464.1 Best-BlastP=> >nrprot 87% Identities = 86/103 (83%), Positives = 93/103 (90%) ref|NP_778069.11 30S ribosomal protein S10 [Buchnera aphidicola (Baizongia pistaciae)] sp|Q89A67|RS10_BUCBP 30S ribosomal protein S10 gb|AA027174.1 | 30S ribosomal protein S10 [Buchnera aphidicola str. Bp (Baizongia pistaciae)] Length = 104
4465.3 Best-BlastP=> >nrprot 37% Identities = 43/118 (36%), Positives = 61/118 (51 %), Gaps = 10/118 (8%) pir||H71023 hypothetical protein PH1485 - Pyrococcus horikoshii dbj|BAA30592.1 | 156aa long hypothetical protein [Pyrococcus horikoshii] Length = 156
4466.3 Best-BlastP=> >nrprot 93% Identities = 338/397 (85%), Positives = 370/397 (93%), Gaps = 1/397 (0%) ref|ZP_00090901.11 COG0050: GTPases - translation elongation factors [Azotobacter vinelandii] Length = 397
4468.2 Best-BlastP=> >nrprot 89% Identities = 527/691 (76%), Positives = 621/691 (89%) ref|NP_819279.11 translation elongation factor G [Coxiella burnetii RSA 493] sp|Q83ES7|EFG_COXBU Elongation factor G (EF-G) gb|AA089793.11 translation elongation factor G [Coxiella burnetii RSA 493] Length = 699
4469.2 Best-BlastP=> >nrprot 48% Identities = 86/277 (31 %), Positives = 138/277 (49%), Gaps = 23/277 (8%) ref|NP_519890.11 PROBABLE OXIDOREDUCTASE DEHYDROGENASE SIGNAL PEPTIDE PROTEIN [Ralstonia solanacearum] emb|CAD15471.11 PROBABLE OXIDOREDUCTASE DEHYDROGENASE SIGNAL PEPTIDE PROTEIN [Ralstonia solanacearum] Length = 300
447.2 Best-BlastP=> >nrprot 99% Identities = 114/114 (100%), Positives = 114/114 (100%) pir||T18325 icmS protein - Legionella pneumophila emb|CAA73237.1 | IcmS protein [Legionella pneumophila] emb|CAA75322.1 | IcmS protein [Legionella pneumophila] Length = 114
4470.1 Best-BlastP=> >nrprot 64% Identities = 156/329 (47%), Positives = 223/329 (67%), Gaps = 6/329 (1 %) ref|NP_742587.11 anthranilate phosphoribosyltransferase [Pseudomonas putida KT2440] sp|Q88QR7|TRPD_PSEPK Anthranilate phosphoribosyltransferase gb|AAN66051.1 |AE016234_4 anthranilate phosphoribosyltransferase [Pseudomonas putida KT2440] Length = 349
4471.3 Best-BlastP=> >nrprot 43% Identities = 47/160 (29%), Positives = 83/160 (51 %), Gaps = 2/160 (1 %) ref|NP_819774.11 conserved hypothetical protein [Coxiella burnetii RSA 493] gb|AAO90288.11 conserved hypothetical protein [Coxiella burnetii RSA 493] Length = 195
4472.2 Best-BlastP=> >nrprot 59% Identities = 116/247 (46%), Positives = 155/247 (62%), Gaps = 1/247 (0%) ref|NP_902024.1 1 probable hydrolase [Chromobacterium violaceum ATCC 12472] gb|AAQ60026.11 probable hydrolase [Chromobacterium violaceum ATCC 12472] Length = 267
4473.4 Best-BlastP=> >nrprot 59% Identities = 89/180 (49%), Positives = 122/180 (67%), Gaps = 1/180 (0%) ref|ZP_00092748.11 hypothetical protein [Azotobacter vinelandii] Length = 397
4474.1 Best-BlastP=> >nrprot 52% Identities = 45/1 14 (39%), Positives = 65/114 (57%), Gaps = 4/1 14 (3%) ref|ZP_00092749.11 hypothetical protein [Azotobacter vinelandii] Length = 135
4475.1 Best-BlastP=> >nrprot 42% Identities = 51/183 (27%), Positives = 96/183 (52%), Gaps = 14/183 (7%) ref|ZP_00047462.11 COG4123: Predicted O-methyltransferase [Lactobacillus gasseri] Length = 338
4477.2 Best-BlastP=> >nrprot 46% Identities = 204/834 (24%), Positives = 357/834 (42%), Gaps = 142/834 (17%) ref|NP_907987.11 PHOSPHOENOLPYRUVATE CARBOXYLASE PEPCASE PEPC [Wolinella succinogenes] emb|CAE10887.1 1 PHOSPHOENOLPYRUVATE CARBOXYLASE PEPCASE PEPC [Wolinella succinogenes] Length = 885
4479.2 Best-BlastP=> >nrprot 60% Identities = 46/119 (38%), Positives = 71/1 19 (59%), Gaps = 3/119 (2%) ref|NP_769098.11 blr2458 [Bradyrhizobium japonicum] dbj|BAC47723.1 | blr2458 [Bradyrhizobium japonicum USDA 1 10] Length = 163
448.1 Best-BlastP=> >nrprot 98% Identities = 86/86 (100%), Positives = 86/86 (100%) pir||T18324 icmT protein - Legionella pneumophila emb|CAA73236.1 1 IcmT protein [Legionella pneumophila] emb|CAA75321.11 IcmT protein [Legionella pneumophila] Length = 86
4480.4 Best-BlastP=> >nrprot 69% Identities = 234/416 (56%), Positives = 291/416 (69%) ref |NP_812629.11 gamma-glutamyl phosphate reductase [Bacteroides thetaiotaomicron VPI-5482] gb|AA078823.11 gamma-glutamyl phosphate reductase [Bacteroides thetaiotaomicron VPI- 5482] Length = 417
4481.1 Best-BlastP=> >nrprot 70% Identities = 91/172 (52%), Positives = 124/172 (72%), Gaps = 3/172 (1 %) ref|NP_177142.11 expressed protein [Arabidopsis thaliana] ref|NP_849870.1 | expressed protein [Arabidopsis thaliana] pir||F96720 unknown protein, 58197-59415 [imported] - Arabidopsis thaliana gb|AAG52556.1 |AC010675_4 unknown protein; 58197-59415 [Arabidopsis thaliana] gb|AAM20691.1 | unknown protein [Arabidopsis thaliana] gb|AAN15655.11 unknown protein [Arabidopsis thaliana] Length = 286
4482.1 Best-BlastP=> >nrprot 56% Identities =Λ 19/320 (37%), Positives = 168/320 (52%), Gaps = 33/320 (10%) ref|ZP_00119712.11 COG4823: Abortive infection bacteriophage resistance protein [Cytophaga hutchinsonii] Length = 331
4483.2 Best-BlastP=> >nrprot No Hits found
4484.1 Best-BlastP=> >nrprot No Hits found '
4485.2 Best-BlastP=> >nrprot 98% Identities = 252/254 (99%), Positives = 252/254 (99%) gb|AAM73853.1 |AF454864_1 putative lipase LipB [Legionella pneumophila] Length = 254
4487.1 Best-BlastP=> >nrprot 70% Identities = 133/223 (59%), Positives = 163/223 (73%) ref|NP_231545.11 orotidine 5'-phosphate decarboxylase [Vibrio cholerae 01 biovar eltor str. N16961] sp|Q9KQT7|PYRF_VIBCH Orotidine 5'-phosphate decarboxylase (OMP decarboxylase) (OMPDCase) (OMPdecase) pir||A82143 orotidine-5'-phosphate decarboxylase (EC 4.1.1.23) VC191 1 [similarity] - Vibrio cholerae (strain N16961 serogroup 01 ) gb|AAF95059.1 | orotidine 5%-phosphate decarboxylase [Vibrio cholerae 01 biovar eltor str. N 16961] Length = 231
4488.1 Best-BlastP=> >nrprot 72% Identities = 213/362 (58%), Positives = 270/362 (74%) ref|NP_903794.11 probable aminotransferase [Chromobacterium violaceum ATCC 12472] gb|AAQ61785.11 probable aminotransferase [Chromobacterium violaceum ATCC 12472] Length = 365
4489.3 Best-BlastP=> >nrprot 67% Identities = 195/383 (50%), Positives = 263/383 (68%), Gaps = 2/383 (0%) ref |NP_819562.11 TPR domain protein [Coxiella burnetii RSA 493] gb|AAO90076.11 TPR domain protein [Coxiella burnetii RSA 493] Length = 388
4490.2 Best-BlastP=> >nrprot 38% Identities = 65/177 (36%), Positives = 109/177 (61 %), Gaps = 12/177 (6%) ref|NP_808996.11 conserved hypothetical protein [Bacteroides thetaiotaomicron VPI-5482] gb|AA075i θ.1 | conserved hypothetical protein [Bacteroides thetaiotaomicron VPI- 5482] Length = 288
4491.3 Best-BlastP=> >nrprot No Hits found
4492.3 Best-BlastP=> >nrprot No Hits found
4493.1 Best-BlastP=> >nrprot No Hits found
4496.2 Best-BlastP=> >nrprot 51 % Identities = 106/240 (44%), Positives = 141/240 (58%), Gaps = 3/240 (1 %) emb|CAB60048.1 | IvrA [Legionella pneumophila] Length = 288
4497.4 Best-BlastP=> >nrprot No Hits found 4498.1 Best-BlastP=> >nrprot No Hits found
45.1 Best-BlastP=> >nrprot 94% Identities = 115/131 (87%), Positives = 125/131 (95%) gb|AAM08243.1 | LvrD [Legionella pneumophila] Length = 131
450.1 Best-BlastP=> >nrprot 39% Identities = 22/50 (44%), Positives = 28/50 (56%) gb|AAF87782.1 |AF279293_1 p76 membrane protein precursor [Mycoplasma hyopneumoniae] Length = 1427
4500.1 Best-BlastP=> >nrprot 82% Identities = 69/116 (59%), Positives = 97/116 (83%) ref|ZP_00067079.11 hypothetical protein [Microbulbifer degradans 2-40] Length = 117
4503.1 Best-BlastP=> >nrprot 77% Identities = 209/334 (62%), Positives = 266/334 (79%) ref|NP_744617.11 phenylalanyl-tRNA synthetase, alpha subunit [Pseudomonas putida KT2440] gb|AAN68081.1 |AE016440_1 phenylalanyl-tRNA synthetase, alpha subunit [Pseudomonas putida KT2440] Length = 338
4505.3 Best-BlastP=> >nrprot 88% Identities = 98/118 (83%), Positives = 106/118 (89%) ref|NP_929901.11 50S ribosomal protein L20 [Photorhabdus luminescens subsp. laumondii TT01] emb|CAE15040.11 50S ribosomal protein L20 [Photorhabdus luminescens subsp. laumondii TT01] Length = 118
4508.4 Best-BlastP=> >nrprot 77% Identities = 90/161 (55%), Positives = 126/161 (78%) ref|NP_719351.11 conserved hypothetical protein [Shewanella oneidensis MR-1] sp|Q8EAS7|Y2B5_SHEON Hypothetical UPF0234 protein S03815 gb|AAN56795.1 |AE015815_1 conserved hypothetical protein [Shewanella oneidensis MR-1] Length = 161
451.3 Best-BlastP=> >nrprot No Hits found 4510.1 Best-BlastP=> >nrprot 83% Identities = 545/812 (67%), Positives = 672/812 (82%), Gaps = 10/812 (1 %) ref|NP_819766.11 ATP-dependent protease La [Coxiella burnetii RSA 493] gb|AAO90280.11 ATP-dependent protease La [Coxiella burnetii RSA 493] Length = 817
4511.1 Best-BlastP=> >nrprot 70% Identities = 53/90 (58%), Positives = 66/90 (73%) ref|ZP_00119213.11 COG0776: Bacterial nucleoid DNA-binding protein [Cytophaga hutchinsonii] Length = 90
4512.4 Best-BlastP=> >nrprot 49% Identities = 181/604 (29%), Positives = 311 /604 (51 %), Gaps = 28/604 (4%) ref|ZP_00139461.11 COG0760: Parvulin- like peptidyl-prolyl isomerase [Pseudomonas aeruginosa UCBPP-PA14] Length = 621
4516.3 Best-BlastP=> >nrprot 56% Identities = 157/337 (46%), Positives = 214/337 (63%), Gaps = 3/337 (0%) ref |ZP_00030194.11 COG0845: Membrane-fusion protein [Burkholderia fungorum] Length = 513
4517.2 Best-BlastP=> >nrprot 51 % Identities = 97/274 (35%), Positives = 145/274 (52%), Gaps = 21/274 (7%) gb|AAK81664.11 MdcB [Burkholderia cepacia] Length = 290
4519.3 Best-BlastP=> >nrprot 78% Identities = 341/546 (62%), Positives = 431/546 (78%), Gaps = 2/546 (0%) ref|NP_640913.11 alpha subunit of malonate decarboxylase [Xanthomonas axonopodis pv. citri str. 306] gb|AAM35449.11 alpha subunit of malonate decarboxylase [Xanthomonas axonopodis pv. citri str. 306] Length = 548
4520.1 Best-BlastP=> >nrprot 73% Identities = 225/396 (56%), Positives = 299/396 (75%), Gaps = 2/396 (0%) sp|Q8GDU2|ASSY_HELMO Argininosuccinate synthase (Citrulline-aspartate ligase) gb|AAN87486.11 Argininosuccinate synthase [Heliobacillus mobilis] Length = 408
4521.1 Best-BlastP=> >nrprot 60% Identities = 96/207 (46%), Positives = 135/207 (65%), Gaps = 15/207 (7%) ref|NP_465264.11 similar to amino acid (glutamine) ABC transporter (ATP-binding protein) [Listeria monocytogenes EGD-e] pir||AC1292 amino acid (glutamine) ABC transporter (ATP-binding protein) homolog lmo173g [imported] - Listeria monocytogenes (strain EGD-e) emb|CAC99817.1 | lmo1739 [Listeria monocytogenes] Length = 215
4522.4 Best-BlastP=> >nrprot 66% Identities = 94/203 (46%), Positives = 144/203 (70%), Gaps = 2/203 (0%) ref|NP_359808.11 amino acid ABC transporter permease protein [Rickettsia conorii] ref |ZP_00142355.1 | amino acid ABC transporter permease protein [Rickettsia sibirica] pir||C97721 amino acid ABC transporter permease protein yqiY [imported] - Rickettsia conorii (strain Malish 7) gb|AAL02709.11 amino acid ABC transporter permease protein [Rickettsia conorii] gb|EAA25764.11 amino acid ABC transporter permease protein [Rickettsia sibirica] Length = 218 4523.1 Best-BlastP=> >nrprot 49% Identities = 56/159 (35%), Positives = 89/159 (55%), Gaps = 4/159 (2%) ref|NP_714273.1 | MutT/nudix family protein [Leptospira interrogans serovar lai str. 56601] gb|AAN51291.1 |AE011563_7 MutT/nudix family protein [Leptospira interrogans serovar lai str. 56601] Length = 223
4524.1 Best-BlastP=> >nrprot 52% Identities = 79/246 (32%), Positives = 125/246 (50%), Gaps = 11/246 (4%) ref|NP_670144.11 arginine 3rd transport system periplasmic binding protein [Yersinia pestis KIM] gb|AAM86395.1 |AE013887_2 arginine 3rd transport system periplasmic binding protein [Yersinia pestis KIM] Length = 252
4525.2 Best-BlastP=> >nrprot 73% Identities = 317/556 (57%), Positives = 418/556 (75%), Gaps = 2/556 (0%) ref|NP_718167.11 long-chain-fatty-acid- CoA ligase [Shewanella oneidensis MR-1] gb|AAN55611.1 |AE015699_9 long-chain-fatty-acid-CoA ligase [Shewanella oneidensis MR-1] Length = 557 4526.3 Best-BlastP=> >nrprot 83% Identities = 64/111 (57%), Positives = 93/111 (83%), Gaps = 2/111 (1 %) ref|ZP_00065090.11 COG1886: Flagellar motor switch/type III secretory pathway protein [Microbulbifer degradans 2-40] Length = 141
4528.2 Best-BlastP=> >nrprot 21 % Identities = 42/73 (57%), Positives = 56/73 (76%) gb|AAN34372.11 ORF2 transposase [Acinetobacter baumannii] Length = 76 453.3 Best-BlastP=> >nrprot 87% Identities = 181/244 (74%), Positives = 220/244 (90%), Gaps = 6/244 (2%) ref|ZP_00023826.11 COG0330: Membrane protease subunits, stomatin/prohibitin homologs [Ralstonia metallidurans] Length = 251
4530.1 Best-BlastP=> >nrprot 69% Identities = 49/91 (53%), Positives = 68/91 (74%) gb|AAP83334.1 |AF469614_2 unknown [Francisella tularensis subsp. tularensis] Length = 94
4531.2 Best-BlastP=> >nrprot 53% Identities = 34/77 (44%), Positives = 47/77 (61 %), Gaps = 5/77 (6%) gb|AAG53985.1 |AF327444_1 putative transposase A [Pantoea agglomerans] Length = 149
4532.2 Best-BlastP=> >nrprot 30% Identities = 127/185 (68%), Positives = 148/185 (80%), Gaps = 11/185 (5%) emb|CAC33489.1 | hypothetical protein [Legionella pneumophila] Length = 189
4533.2 Best-BlastP=> >nrprot 50% Identities = 643/2158 (29%), Positives = 955/2158 (44%), Gaps = 397/2158 (18%) ref|NP_758987.11 unknown [Zymomonas mobilis] gb|AAL36122.11 unknown [Zymomonas mobilis] Length = 2201
4535.2 Best-BlastP=> >nrprot 69% Identities = 226/438 (51%), Positives = 312/438 (71 %), Gaps = 18/438 (4%) ref|ZP_00089764.1 | COG0793: Periplasmic protease [Azotobacter vinelandii] Length = 456
4536.1 Best-BlastP=> >nrprot 49% Identities = 114/349 (32%), Positives = 189/349 (54%), Gaps = 12/349 (3%) ref|ZP_00065627.11 COG4942: Membrane-bound metallopeptidase [Microbulbifer degradans 2-40] Length = 402
4538.2 Best-BlastP=> >nrprot 68% Identities = 273/506 (53%), Positives = 353/506 (69%), Gaps = 4/506 (0%) ref|ZP_00141602.11 COG0696: Phosphoglyceromutase [Pseudomonas aeruginosa UCBPP-PA14] Length = 515
454.1 Best-BlastP=> >nrprot 59% Identities = 215/454 (47%), Positives = 301/454 (66%), Gaps = 31/454 (6%) ref|NP_820466.11 membrane protein, putative [Coxiella burnetii RSA 493] gb|AAO90980.11 membrane protein, putative [Coxiella burnetii RSA 493] Length = 458
4540.2 Best-BlastP=> >nrprot No Hits found
4542.4 Best-BlastP=> >nrprot 35% Identities = 136/313 (43%), Positives = 191/313 (61%), Gaps = 22/313 (7%) ref|ZP_00033133.1 | COG2010: Cytochrome c, mono- and diheme variants [Burkholderia fungorum] Length = 432
4546.3 Best-BlastP=> >nrprot 21 % Identities = 47/163 (28%), Positives = 74/163 (45%), Gaps = 18/163 (11 %) ref|ZP_00011332.11 COG0183: Acetyl- CoA acetyltransferase [Rhodopseudomonas palustris] Length = 504
4547.4 Best-BlastP=> >nrprot 76% Identities = 206/337 (61 %), Positives = 260/337 (77%), Gaps = 3/337 (0%) ref|ZP_00125317.11 COG0604: NADPH:quinone reductase and related Zn-dependent oxidoreductases [Pseudomonas syringae pv. syringae B728a] Length = 337
4549.2 Best-BlastP=> >nrprot 21 % Identities = 73/109 (66%), Positives = 94/109 (86%) ref|NP_820449.11 hypothetical protein [Coxiella burnetii RSA 493] gb|AAO90963.11 hypothetical protein [Coxiella burnetii RSA 493] Length = 113
4551.2 Best-BlastP=> >nrprot 55% Identities = 31/78 (39%), Positives = 49/78 (62%) ref|NP_875585.11 Uncharacterized conserved membrane protein [Prochlorococcus marinus subsp. marinus str. CCMP1375] gb|AAQ00238.1 | Uncharacterized conserved membrane protein [Prochlorococcus marinus subsp. marinus str. CCMP1375] Length = 91
4552.4 Best-BlastP=> >nrprot No Hits found
4553.2 Best-BlastP=> >nrprot 66% Identities = 107/240 (44%), Positives = 156/240 (65%), Gaps = 14/240 (5%) ref|NP_867184.1 1 short chain alcohol dehydrogenase-like [Pirellula sp.] emb|CAD74729.11 short chain alcohol dehydrogenase-like [Pirellula sp.] Length = 247
4554.2 Best-BlastP=> >nrprot 35% Identities = 27/4g (55%), Positives = 31/49 (63%), Gaps = 1/49 (2%) ref|ZP_00031568.1 | COG1051 : ADP-ribose pyrophosphatase [Burkholderia fungorum] Length = 181
4558.1 Best-BlastP=> >nrprot 29% Identities = 43/182 (23%), Positives = 82/182 (45%), Gaps = 9/182 (4%) gb|AAK27486.1 |AF343323_1 DNA gyrase B [Cycloclasticus sp. NOP-122A] gb|AAK27491.1 |AF343328_1 DNA gyrase B [Cycloclasticus sp. N-221 A] gb|AAK27492.1 |AF343329_1 DNA gyrase B [Cycloclasticus sp. N-231 B] gb|AAK27493.1 |AF343330_1 DNA gyrase B [Cycloclasticus sp. P-211 A2] Length = 362
4559.1 Best-BlastP=> >nrprot 59% Identities = 206/502 (41 %), Positives = 308/502 (61 %), Gaps = 6/502 (1%) ref|NP_819464.11 amino acid permease family protein [Coxiella burnetii RSA 493] gb|AA089978.11 amino acid permease family protein [Coxiella burnetii RSA 493] Length = 531
4560.2 Best-BlastP=> >nrprot 26% Identities = 49/217 (22%), Positives = 98/217 (45%), Gaps = 30/217 (13%) ref|NP_702678.11 hypothetical protein [Plasmodium falciparum 3D7] emb|CAD49112.11 hypothetical protein [Plasmodium falciparum 3D7] Length = 1248
4561.2 Best-BlastP=> >nrprot 48% Identities = 140/428 (32%), Positives = 222/428 (51 %), Gaps = 3/428 (0%) ref|NP_636850.11 outer membrane component of multidrug efflux pump [Xanthomonas campestris pv. campestris str. ATCC 33913] gb|AAM40774.11 outer membrane component of multidrug efflux pump [Xanthomonas campestris pv. campestris str. ATCC 33913] Length = 467
4564.2 Best-BlastP=> >nrprot 99% Identities = 198/200 (99%), Positives = 199/200 (99%) gb|AAM00391.1 |AF386079_1 CcmA [Legionella pneumophila] Length = 200
4566.1 Best-BlastP=> >nrprot 62% Identities = 93/192 (48%), Positives = 124/192 (64%), Gaps = 5/192 (2%) ref|ZP_00071947.11 COG0693: Putative intracellular protease/amidase [Trichodesmium erythraeum IMS101] Length = 191
4567.2 Best-BlastP=> >nrprot 10% Identities = 44/175 (25%), Positives = 69/175 (39%), Gaps = 36/175 (20%) gb|AAK20704.1 |AF316641_10 WciT [Streptococcus pneumoniae] Length = 242
457.2 Best-BlastP=> >nrprot 50% Identities = 180/528 (34%), Positives = 271/528 (51 %), Gaps = 58/528 (10%) ref|NP_820680.1 | conserved hypothetical protein [Coxiella burnetii RSA 493] gb|AA091194.11 conserved hypothetical protein [Coxiella burnetii RSA 493] Length = 518
4571.2 Best-BlastP=> >nrprot 34% Identities = 29/58 (50%), Positives = 38/58 (65%) ref|NP_819567.11 lipoprotein, putative [Coxiella burnetii RSA 493] gb|AAO90081.11 lipoprotein, putative [Coxiella burnetii RSA 493] Length = 323
4572.2 Best-BlastP=> >nrprot 63% Identities = 175/348 (50%), Positives = 227/348 (65%), Gaps = 2/348 (0%) ref|ZP_00068140.11 COG1194: A/G- specific DNA glycosylase [Microbulbifer degradans 2-40] Length = 355
4573.4 Best-BlastP=> >nrprot 43% Identities = 123/542 (22%), Positives = 226/542 (41%), Gaps = 78/542 (14%) ref|NP_819951.11 conserved hypothetical protein [Coxiella burnetii RSA 493] gb|AAO90465.11 conserved hypothetical protein [Coxiella burnetii RSA 493] Length = 502
4574.2 Best-BlastP=> >nrprot 44% Identities = 95/367 (25%), Positives = 164/367 (44%), Gaps = 55/367 (14%) ref|ZP_00089878.11 COG1562: Phytoene/squalene synthetase [Azotobacter vinelandii] Length = 377
4577.2 Best-BlastP=> >nrprot 48% Identities = 101/231 (43%), Positives = 135/231 (58%), Gaps = 19/231 (8%) ref|NP_819994.11 rare lipoprotein A family protein [Coxiella burnetii RSA 493] gb|AAO90508.11 rare lipoprotein A family protein [Coxiella burnetii RSA 493] Length = 261
4578.1 Best-BlastP=> >nrprot 80% Identities = 224/390 (57%), Positives = 314/390 (80%), Gaps = 1/390 (0%) ref|NP_819495.11 sodium/hydrogen antiporter family protein [Coxiella burnetii RSA 493] gb|AAO90009.11 sodium/hydrogen antiporter family protein [Coxiella burnetii RSA 493] Length = 389
458.3 Best-BlastP=> >nrprot 47% Identities = 93/334 (27%), Positives = 164/334 (49%), Gaps = 18/334 (5%) ref|NP_819588.11 DNA polymerase III,
delta subunit [Coxiella burnetii RSA 493] gb|AAO90102.11 DNA polymerase III, delta subunit [Coxiella burnetii RSA 493] Length = 339
4580.1 Best-BlastP=> >nrprot 34% Identities = 41/110 (37%), Positives = 58/110 (52%), Gaps = 3/110 (2%) ref|NP_742951.11 inner membrane protein AmpE [Pseudomonas putida KT2440] gb|AAN66415.1 |AE016269_2 inner membrane protein AmpE [Pseudomonas putida KT2440] Length = 276
4581.2 Best-BlastP=> >nrprot 29% Identities = 22/97 (22%), Positives = 39/97 (40%), Gaps = 1/97 (1 %) ref|NP_820299.11 conserved hypothetical protein [Coxiella burnetii RSA 493] gb|AAO90813.11 conserved hypothetical protein [Coxiella burnetii RSA 493] Length = 155
4584.3 Best-BlastP=> >nrprot No Hits found
4587.2 Best-BlastP=> >nrprot No Hits found 4588.2 Best-BlastP=> >nrprot No Hits found 4589.1 Best-BlastP=> >nrprot No Hits found 459.3 Best-BlastP=> >nrprot 56% Identities = 76/212 (35%), Positives = 119/212 (56%), Gaps = 2/212 (0%) ref |NP_406133.11 putative nicotinate- nucleotide adenylyltransferase [Yersinia pestis] sp|Q8ZDG1 |NADD_YERPE Probable nicotinate-nucleotide adenylyltransferase (Deamido- NAD(+) pyrophosphorylase) (Deamido-NAD(+) diphosphorylase) (Nicotinate mononucleotide adenylyltransferase) (NaMN adenylyltransferase) pir||AC0318 probable nicotinate-nucleotide adenylyltransferase (EC 2.7.7.18) [imported] - Yersinia pestis (strain C092) emb|CAC92850.11 putative nicotinate-nucleotide adenylyltransferase [Yersinia pestis C092] Length = 220
4590.1 Best-BlastP=> >nrprot 56% Identities = 57/130 (43%), Positives = 77/130 (59%), Gaps = 6/130 (4%) ref|ZP_00086142.11 COG2764: Uncharacterized protein conserved in bacteria [Pseudomonas fluorescens PfO-1] Length = 137
4591.2 Best-BlastP=> >nrprot 71 % Identities = 86/155 (55%), Positives = 113/155 (72%), Gaps = 4/155 (2%) ref|ZP_00109160.11 COG3865: Uncharacterized protein conserved in bacteria [Nostoc punctiforme] Length = 165
4597.1 Best-BlastP=> >nrprot No Hits found
4598.2 Best-BlastP=> >nrprot 71 % Identities = 132/248 (53%), Positives = 176/248 (70%), Gaps = 5/248 (2%) ref|NP_902034.11 acetoacetyl-CoA reductase [Chromobacterium violaceum ATCC 12472] gb|AAQ60036.11 acetoacetyl-CoA reductase [Chromobacterium violaceum ATCC 12472] Length = 246
4599.2 Best-BlastP=> >nrprot 78% Identities = 135/206 (65%), Positives = 169/206 (82%) ref|ZP_00024696.11 COG1484: DNA replication protein [Ralstonia metallidurans] Length = 268
46.1 Best-BlastP=> >nrprot 97% Identities = 220/236 (93%), Positives = 230/236 (97%) gb|AAM08241.1 1 putative TraC protein [Legionella pneumophila] Length = 236
460.2 Best-BlastP=> >nrprot 53% Identities = 59/158 (37%), Positives = 95/158 (60%) ref|NP_819911.11 colicin V production protein [Coxiella burnetii RSA 493] gb|AAO90425.11 colicin V production protein [Coxiella burnetii RSA 493] Length = 184
4600.2 Best-BlastP=> >nrprot 52% Identities = 83/238 (34%), Positives = 128/238 (53%), Gaps = 27/238 (11 %) ref|NP_932037.11 hypothetical protein [Photorhabdus luminescens subsp. laumondii TT01] emb|CAE17256.11 unnamed protein product [Photorhabdus luminescens subsp. laumondii TT01] Length = 255
4601.1 Best-BlastP=> >nrprot No Hits found
4602.3 Best-BlastP=> >nrprot 99% Identities = 300/302 (99%), Positives = 302/302 (100%) emb|CAC14311.11 putative transcriptional regulator [Legionella pneumophila] Length = 302
4604.3 Best-BlastP=> >nrprot 48% Identities = 48/182 (26%), Positives = 91/182 (50%), Gaps = 12/182 (6%) ref|NP_251679.1 | hypothetical protein [Pseudomonas aeruginosa PA01] pir||F83271 hypothetical protein PA2989 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG06377.1 |AE004724_6 hypothetical protein PA2989 [Pseudomonas aeruginosa PA01] Length = 254
4607.2 Best-BlastP=> >nrprot 79% Identities = 265/421 (62%), Positives = 335/421 (79%) ref|ZP_00092427.11 hypothetical protein [Azotobacter vinelandii] Length = 838
4608.1 Best-BlastP=> >nrprot 33% Identities = 34/111 (30%), Positives = 60/111 (54%), Gaps = 3/111 (2%) ref|NP_700535.11 hypothetical protein [Plasmodium falciparum 3D7] gb|AAN35259.1 |AE014830_3 hypothetical protein [Plasmodium falciparum 3D7] Length = 426
4609.2 Best-BlastP=> >nrprot No Hits found
4613.3 Best-BlastP=> >nrprot 48% Identities = 121/487 (24%), Positives = 240/487 (49%), Gaps = 29/487 (5%) ref|NP_753921.1 1 Hypothetical transporter ydgR [Escherichia coli CFT073] gb|AAN80486.1 |AE016761_61 Hypothetical transporter ydgR [Escherichia coli CFT073] Length = 500
4615.3 Best-BlastP=> >nrprot No Hits found
4616.2 Best-BlastP=> >nrprot 48% Identities = 93/315 (29%), Positives = 150/315 (47%), Gaps = 35/315 (11 %) ref|NP_820641.1 | hypothetical protein [Coxiella burnetii RSA 493] gb|AA091155.11 hypothetical protein [Coxiella burnetii RSA 493] Length = 429
4618.1 Best-BlastP=> >nrprot 58% Identities = 110/250 (44%), Positives = 151/250 (60%), Gaps = 7/250 (2%) dbj|BAB72031.1 | lipopolysaccharide biosynthesis glycosyltransferase [Photobacterium damselae subsp. piscicida] Length = 255
462.2 Best-BlastP=> >nrprot 44% Identities = 71/278 (25%), Positives = 116/278 (41 %), Gaps = 35/278 (12%) ref|NP_718631 .11 conserved hypothetical protein [Shewanella oneidensis MR-1] gb|AAN56075.1 |AE015743_3 conserved hypothetical protein [Shewanella oneidensis MR-1] Length = 281 4620.1 Best-BlastP=> >nrprot 69% Identities = 49/91 (53%), Positives = 68/91 (74%) gb|AAP83334.1 |AF469614_2 unknown [Francisella tularensis subsp. tularensis] Length = 94
4623.2 Best-BlastP=> >nrprot 52% Identities = 66/169 (39%), Positives = 93/169 (55%), Gaps = 5/169 (2%) ref|NP_814807.1 | acetyltransferase, GNAT family [Enterococcus faecalis V583] gb|AAO80877.11 acetyltransferase, GNAT family [Enterococcus faecalis V583] Length = 168
4625.2 Best-BlastP=> >nrprot 68% Identities = 162/307 (52%), Positives = 215/307 (70%), Gaps = 1/307 (0%) ref|NP_761810.11 S-malonyltransferase [Vibrio vulnificus CMCP6] gb|AA011337.1 |AE016807_56 S-malonyltransferase [Vibrio vulnificus CMCP6] Length = 307
4626.1 Best-BlastP=> >nrprot 72% Identities = 172/314 (54%), Positives = 231/314 (73%), Gaps = 3/314 (0%) ref|NP_930069.1 | 3-oxoacyl-[acyl carrier- protein] synthase III (beta-ketoacyl-ACP synthase III) (KAS III) [Photorhabdus luminescens subsp. laumondii TT01] emb|CAE15209.11 3-oxoacyl-[acyl carrier-protein] synthase III (beta-ketoacyl-ACP synthase III) (KAS III) [Photorhabdus luminescens subsp. laumondii TT01 ] Length = 317
4627.2 Best-BlastP=> >nrprot 72% Identities = 183/330 (55%), Positives = 241/330 (73%), Gaps = 2/330 (0%) ref|NP_819526.11 fatty acid/phospholipid synthesis protein PlsX [Coxiella burnetii RSA 493] sp|Q83E40|PLSX_COXBU Fatty acid/phospholipid synthesis protein plsX gb|AAO90040.11 fatty acid/phospholipid synthesis protein PlsX [Coxiella burnetii RSA 493] Length = 343
4628.2 Best-BlastP=> >nrprot 78% Identities = 45/55 (81 %), Positives = 50/55 (90%) ref|ZP_00135404.11 COG0333: Ribosomal protein L32 [Actinobacillus pleuropneumoniae serovar 1 str. 4074] ref|NP_873288.1 | 50S ribosomal protein L32 [Haemophilus ducreyi 35000HP] gb|AAP95677.11 50S ribosomal protein L32 [Haemophilus ducreyi 35000HP] Length = 56
4629.2 Best-BlastP=> >nrprot 33% Identities = 23/98 (23%), Positives = 48/98 (48%), Gaps = 1/98 (1%) ref|ZP_00136314.1 | COG1399: Predicted metal- binding, possibly nucleic acid-binding protein [Pseudomonas aeruginosa UCBPP-PA14] Length = 148 463.2 Best-BlastP=> >nrprot No Hits found
4632.1 Best-BlastP=> >nrprot No Hits found
4633.2 Best-BlastP=> >nrprot 82% identities = 121/184 (65%), Positives = 154/184 (83%) ref|NP_842482.11 Uncharacterized protein family UPF0016 [Nitrosomonas europaea ATCC 19718] emb|CAD86405.11 Uncharacterized protein family UPF0016 [Nitrosomonas europaea ATCC 19718] Length = 192
4634.2 Best-BlastP=> >nrprot 52% Identities = 222/716 (31 %), Positives = 348/716 (48%), Gaps = 80/716 (11%) ref|NP_924649.1 | probable peptidase [Gloeobacter violaceus] dbj|BAC89644.11 gill 703 [Gloeobacter violaceus] Length = 730
4636.4 Best-BlastP=> >nrprot 4% Identities = 49/164 (29%), Positives = 74/164 (45%), Gaps = 12/164 (7%) ref|ZP_00009072.11 hypothetical protein [Rhodopseudomonas palustris] Length = 275
464.2 Best-BlastP=> >nrprot 60% Identities = 167/405 (41 %), Positives = 258/405 (63%), Gaps = 4/405 (0%) ref|NP_819909.11 FolC bifunctional protein [Coxiella burnetii RSA 493] gb|AAO90423.11 FolC bifunctional protein [Coxiella burnetii RSA 493] Length = 416
4640.2 Best-BlastP=> >nrprot 74% Identities = 466/845 (55%), Positives = 627/845 (74%), Gaps = 12/845 (1 %) ref|NP_820057.11 DNA mismatch repair protein MutS [Coxiella burnetii RSA 493] gb|AAO90571.11 DNA mismatch repair protein MutS [Coxiella burnetii RSA 493] Length = 859
4642.1 Best-BlastP=> >nrprot 63% Identities = 79/154 (51%), Positives = 107/154 (69%), Gaps = 1/154 (0%) ref|ZP_00025457.1 | COG1546: Uncharacterized protein (competence- and mitomycin-induced) [Ralstonia metallidurans] Length = 200
4643.2 Best-BlastP=> >nrprot 56% Identities = 176/176 (100%), Positives = 176/176 (100%) emb|CAC33483.11 hypothetical protein [Legionella pneumophila] Length = 176
4644.3 Best-BlastP=> >nrprot 41% Identities = 95/419 (22%), Positives = 174/419 (41%), Gaps = 31/419 (7%) ref |ZP_00071833.11 hypothetical protein [Trichodesmium erythraeum IMS101] Length = 424
4645.2 Best-BlastP=> >nrprot 44% Identities = 96/328 (29%), Positives = 162/328 (49%), Gaps = 10/328 (3%) ref|ZP_00056223.11 COG0477: Permeases of the major facilitator superfamily [Magnetospirillum magnetotacticum] Length = 407
4647.2 Best-BlastP=> >nrprot 25% Identities = 54/234 (23%), Positives = 94/234 (40%), Gaps = 8/234 (3%) ref|ZP_00034418.11 COG0683: ABC-type branched-chain amino acid transport systems, periplasmic component [Burkholderia fungorum] Length = 451
4649.1 Best-BlastP=> >nrprot No Hits found 465.2 Best-BlastP=> >nrprot 75% Identities = 186/283 (65%), Positives = 222/283 (78%), Gaps = 3/283 (1 %) ref|NP_819908.11 acetyl-CoA carboxylase, carboxyl transferase, beta subunit [Coxiella burnetii RSA 493] gb|AAO90422.11 acetyl-CoA carboxylase, carboxyl transferase, beta subunit [Coxiella burnetii RSA 493] Length = 291
4650.3 Best-BlastP=> >nrprot 45% Identities = 301/1302 (23%), Positives = 574/1302 (44%), Gaps = 91/1302 (6%) ref |NP_901766.11 probable transmembrane protein [Chromobacterium violaceum ATCC 12472] gb|AAQ59768.1 | probable transmembrane protein [Chromobacterium violaceum ATCC 12472] Length = 1272
4652.2 Best-BlastP=> >nrprot 62% Identities = 51/115 (44%), Positives = 79/115 (68%) ref|NP_353968.1 | AGR_C_1731 p [Agrobacterium tumefaciens] pir||H97474 hypothetical 14.1K protein in rpli-cpdb intergenic region [imported] - Agrobacterium tumefaciens (strain C58, Cereon) gb|AAK86753.11 AGR_C_1731 p [Agrobacterium tumefaciens str. C58 (Cereon)] Length = 190
4653.2 Best-BlastP=> >nrprot 26% Identities = 50/154 (32%), Positives = 79/154 (51 %), Gaps = 17/154 (11 %) ref |NP_440166.11 hypothetical protein [Synechocystis sp. PCC 6803] sp|P72831 |YC98_SYNY3 Hypothetical protein slr1298 pir||S74695 hypothetical protein slr1298 - Synechocystis sp. (strain PCC 6803) dbj|BAA16846.11 ORF_ID:slr1298~hypothetical protein [Synechocystis sp. PCC 6803] Length = 755
4655.3 Best-BlastP=> >nrprot 47% Identities = 130/460 (28%), Positives = 216/460 (46%), Gaps = 31/460 (6%) ref|NP_840264.11 Outer membrane efflux protein [Nitrosomonas europaea ATCC 19718] emb|CAD84081.1 | Outer membrane efflux protein [Nitrosomonas europaea ATCC 19718] Length = 516 4656.2 Best-BlastP=> >nrprot 26% Identities = 69/366 (18%), Positives = 161/366 (43%), Gaps = 26/366 (7%) ref|XP_230851.2| similar to hypothetical protein [Rattus norvegicus] Length = 396
4657.1 Best-BlastP=> >nrprot 62% Identities = 94/202 (46%), Positives = 131/202 (64%), Gaps = 7/202 (3%) ref |NP_520142.1 1 PUTATIVE GST- RELATED PROTEIN [Ralstonia solanacearum] emb|CAD15723.11 PUTATIVE GST-RELATED PROTEIN [Ralstonia solanacearum] Length = 230
4658.2 Best-BlastP=> >nrprot 75% Identities = 94/134 (70%), Positives = 105/134 (78%), Gaps = 1/134 (0%) ref|NP_105189.11 organic hydroperoxide resistance protein [Mesorhizobium loti] dbj|BAB50975.1 1 organic hydroperoxide resistance protein [Mesorhizobium loti] Length = 137
4659.2 Best-BlastP=> >nrprot No Hits found
4660.2 Best-BlastP=> >nrprot No Hits found 4662.1 Best-BlastP=> >nrprot No Hits found
4663.3 Best-BlastP=> >nrprot 98% Identities = 288/297 (96%), Positives = 294/297 (98%) pir||A42596 major outer membrane protein - Legionella pneumophila gb|AAA25300.11 major outer membrane protein Length = 297
4665.2 Best-BlastP=> >nrprot 45% Identities = 244/604 (40%), Positives = 363/604 (60%), Gaps = 38/604 (6%) ref|NP_715981.11 sensory box protein [Shewanella oneidensis MR-1] gb|AAN53426.1 |AE015481_9 sensory box protein [Shewanella oneidensis MR-1] Length = 1515
4666.2 Best-BlastP=> >nrprot 53% Identities = 118/339 (34%), Positives = 174/339 (51 %), Gaps = 44/339 (12%) ref|NP_720115.11 ribonuclease, T2 family [Shewanella oneidensis MR-1] gb|AAN57559.1 |AE015891_10 ribonuclease, T2 family [Shewanella oneidensis MR-1] Length = 384
4668.1 Best-BlastP=> >nrprot 25% Identities = 75/198 (37%), Positives = 112/198 (56%), Gaps = 1/198 (0%) ref|ZP_00054876.11 hypothetical protein [Magnetospirillum magnetotacticum] Length = 626
4669.2 Best-BlastP=> >nrprot 76% Identities = 48/71 (67%), Positives = 55/71 (77%), Gaps = 1/71 (1 %) ref|NP_232566.11 cold shock transcriptional regulator CspA [Vibrio cholerae 01 biovar eltor str. N16961] sp|Q9KN00|CSPA_VIBCH Cold shock-like protein cspA pir||G82492 cold shock transcription regulator CspA VCA0166 [imported] - Vibrio cholerae (strain N16961 serogroup 01) gb|AAF96079.1 | cold shock transcriptional regulator CspA [Vibrio cholerae 01 biovar eltor str. N 16961] Length = 70
4671.3 Best-BlastP=> >nrprot 58% Identities = 131/353 (37%), Positives = 208/353 (58%), Gaps = 5/353 (1 %) ref|NP_243998.1 | endo-1 ,4-beta- glucanase [Bacillus halodurans] pir||D84041 endo-1 ,4-beta-glucanase BH3132 [imported] - Bacillus halodurans (strain C-125) dbj|BAB06851.11 endo-1 ,4-beta-glucanase [Bacillus halodurans] Length = 361
4672.1 Best-BlastP=> >nrprot 67% Identities = 119/224 (53%), Positives = 161 /224 (71 %), Gaps = 2/224 (0%) ref|ZP_00131766.11 COG0220: Predicted S-adenosylmethionine-dependent methyltransferase [Haemophilus somnus 2336] Length = 251
4673.1 Best-BlastP=> >nrprot 83% Identities = 59/78 (75%), Positives = 66/78 (84%) ref|NP_439112.11 ribosomal protein L28 [Haemophilus influenzae Rd] sp|P44364|RL28_HAEIN 50S ribosomal protein L28 pir||E64104 ribosomal protein L28 - Haemophilus influenzae (strain Rd KW20) gb|AAC22612.11 ribosomal protein L28 (rpL28) [Haemophilus influenzae Rd] Length = 78
4676.2 Best-BlastP=> >nrprot 73% Identities = 262/488 (53%), Positives = 342/488 (70%), Gaps = 32/488 (6%) dbj|BAC93212.11 DNA-directed RNA polymerase specialized sigma subunit [Vibrio vulnificus YJ016] Length = 487
4678.3 Best-BlastP=> >nrprot No Hits found
468.3 Best-BlastP=> >nrprot 76% Identities = 81/121 (66%), Positives = 99/121 (81 %) ref|ZP_00089152.11 COG0251 : Putative translation initiation inhibitor, yjgF family [Azotobacter vinelandii] Length = 234
4680.2 Best-BlastP=> >nrprot 74% Identities = 129/219 (58%), Positives = 169/219 (77%) ref|NP_642403.11 ABC transporter ATP-binding protein [Xanthomonas axonopodis pv. citri str. 306] gb|AAM36939.11 ABC transporter ATP-binding protein [Xanthomonas axonopodis pv. citri str. 306] Length = 227
4683.3 Best-BlastP=> >nrprot 73% Identities = 215/417 (51 %), Positives = 305/417 (73%), Gaps = 5/417 (1 %) ref|NP_820008.11 lipoprotein ABC transporter, permease protein, putative [Coxiella burnetii RSA 493] gb|AAO90522.11 lipoprotein ABC transporter, permease protein, putative [Coxiella burnetii RSA 493] Length = 414
4688.2 Best-BlastP=> >nrprot 73% Identities = 199/352 (56%), Positives = 275/352 (78%) ref |NP_643914.11 type II secretion system protein-like protein [Xanthomonas axonopodis pv. citri str. 306] gb|AAM38450.1 | type II secretion system protein-like protein [Xanthomonas axonopodis pv. citri str. 306] Length = 377
469.3 Best-BlastP=> >nrprot 74% Identities = 411/703 (58%), Positives = 530/703 (75%), Gaps = 5/703 (0%) ref|NP_819346.11 guanosine-3,5- bis(diphosphate) 3-pyrophosphohydrolase [Coxiella burnetii RSA 493] gb|AAO89860.1 | guanosine-3,5-bis(diphosphate) 3- pyrophosphohydrolase [Coxiella burnetii RSA 493] Length = 707
4690.2 Best-BlastP=> >nrprot 4% Identities = 21/50 (42%), Positives = 31/50 (62%) ref|NP_055206.11 cardiac ankyrin repeat protein; cytokine inducible nuclear protein [Homo sapiens] pir||A57291 cytokine inducible nuclear protein C193 - human emb|CAA58676.1 | nuclear protein [Homo sapiens] Length = 319
4691.2 Best-BlastP=> >nrprot 82% Identities = 279/413 (67%), Positives = 356/413 (86%), Gaps = 3/413 (0%) ref|NP_670162.11 probable serine transporter [Yersinia pestis KIM] gb|AAM86413.1 |AE013888_9 probable serine transporter [Yersinia pestis KIM] Length = 440
4692.5 Best-BlastP=> >nrprot 71 % Identities = 232/427 (54%), Positives = 316/427 (74%), Gaps = 3/427 (0%) ref|NP_820015.11 adenosylmethionine-8- amino-7-oxononanoate aminotransferase [Coxiella burnetii RSA 493] gb|AAO90529.11 adenosylmethionine-8-amino-7-oxononanoate aminotransferase [Coxiella burnetii RSA 493] Length = 442
4694.1 Best-BlastP=> >nrprot 59% Identities = 187/507 (36%), Positives = 297/507 (58%), Gaps = 30/507 (5%) ref|NP_616892.11 serine-type D-Ala-D- Ala carboxypeptidase [Methanosarcina acetivorans str. C2A] gb|AAM05372.11 serine-type D-Ala-D-Ala carboxypeptidase [Methanosarcina acetivorans str. C2A] Length = 568
4695.1 Best-BlastP=> >nrprot No Hits found
4696.3 Best-BlastP=> >nrprot 31 % Identities = 102/309 (33%), Positives = 170/309 (55%), Gaps = 8/309 (2%) ref|NP_461159.11 putative membrane protein involved in resistance to lambda and N4 phages [Salmonella typhimurium LT2] gb|AAL21118.11 putative membrane protein involved in resistance to lambda and N4 phages [Salmonella typhimurium LT2] Length = 518
4699.3 Best-BlastP=> >nrprot 85% Identities = 315/417 (75%), Positives = 364/417 (87%), Gaps = 5/417 (1 %) ref|NP_793499.11 ATP-dependent Clp protease, ATP-binding subunit ClpX [Pseudomonas syringae pv. tomato str. DC3000] gb|AA057194.1 | ATP-dependent Clp protease, ATP- binding subunit ClpX [Pseudomonas syringae pv. tomato str. DC3000] Length = 427
47.1 Best-BlastP=> >nrprot 97% Identities = 784/826 (94%), Positives = 806/826 (97%) gb|AAM08240.11 putative type IV secretion protein B4 [Legionella pneumophila] Length = 826
470.3 Best-BlastP=> >nrprot 88% Identities = 48/68 (70%), Positives = 60/68 (88%) ref|ZP_00085276.11 COG1758: DNA-directed RNA polymerase, subunit K/omega [Pseudomonas fluorescens PfO-1] Length = 87
4701.2 Best-BlastP=> >nrprot 58% Identities = 118/269 (43%), Positives = 163/269 (60%), Gaps = 14/269 (5%) ref|NP_819610.11 conserved hypothetical protein [Coxiella burnetii RSA 493] gb|AAO90124.11 conserved hypothetical protein [Coxiella burnetii RSA 493] Length = 273
4702.1 Best-BlastP=> >nrprot 65% Identities = 96/188 (51%), Positives = 131/188 (69%) ref |NP_819608.11 glycosyl transferase, group 2 family protein [Coxiella burnetii RSA 493] gb|AAO90122.11 glycosyl transferase, group 2 family protein [Coxiella burnetii RSA 493] Length = 324
4703.2 Best-BlastP=> >nrprot 48% Identities = 53/118 (44%), Positives = 79/118 (66%) ref|NP_900830.11 probable glycosyl transferase [Chromobacterium violaceum ATCC 12472] gb|AAQ58835.11 probable glycosyl transferase [Chromobacterium violaceum ATCC 12472] Length = 335
4704.2 Best-BlastP=> >nrprot 56% Identities = 81/220 (36%), Positives = 127/220 (57%), Gaps = 3/220 (1 %) ref|ZP_00021591.11 COG3376: High- affinity nickel permease [Ralstonia metallidurans] Length = 278
4705.3 Best-BlastP=> >nrprot 31% Identities = 55/205 (26%), Positives = 94/205 (45%), Gaps = 29/205 (14%) ref|NP_052786.11 pXO1-90 [Bacillus anthracis] ref|NP_652888.11 S-layer protein, (pXO1-90) [Bacillus anthracis str. A2012] pir||B59102 hypothetical protein pXO1-90 - Bacillus anthracis virulence plasmid pXOI gb|AAD32394.11 pXO1-90 [Bacillus anthracis] gb|AAM26077.11 S-layer protein, (pXO1-90) [Bacillus anthracis str. A2012] Length = 652
4706.3 Best-BlastP=> >nrprot 86% Identities = 106/141 (75%), Positives = 123/141 (87%) dbj|BAC55152.1 | nucleoside diphosphate kinase [Halomonas sp. #593] Length = 141
4709.1 Best-BlastP=> >nrprot 73% Identities = 213/358 (59%), Positives = 281/358 (78%), Gaps = 1/358 (0%) ref|NP_274327.11 conserved hypothetical protein [Neisseria meningitidis MC58] pir||B81098 conserved hypothetical protein NMB1308 [imported] - Neisseria meningitidis (strain MC58 serogroup B) gb|AAF41683.11 conserved hypothetical protein [Neisseria meningitidis MC58] Length = 364
4710.1 Best-BlastP=> >nrprot 40% Identities = 70/205 (34%), Positives = 107/205 (52%) ref |NP_231252.11 fimbrial biogenesis and twitching motility protein, putative [Vibrio cholerae 01 biovar eltor str. N16961] pir||F82178 probable fimbrial biogenesis and twitching motility protein VC1612 [imported] - Vibrio cholerae (strain N16961 serogroup 01 ) gb|AAF94766.11 fimbrial biogenesis and twitching motility protein, putative [Vibrio cholerae 01 biovar eltor str. N 16961 ] Length = 237
4711.1 Best-BlastP=> >nrprot 41 % Identities = 42/159 (26%), Positives = 78/159 (49%), Gaps = 21/159 (13%) ref|NP_820244.11 DNA-binding protein, putative [Coxiella burnetii RSA 493] gb|AAO90758.11 DNA-binding protein, putative [Coxiella burnetii RSA 493] Length = 203
4712.2 Best-BlastP=> >nrprot 71 % Identities = 250/418 (59%), Positives = 306/418 (73%), Gaps = 2/418 (0%) ref|ZP_00091666.1 | COG0124: Histidyl- tRNA synthetase [Azotobacter vinelandii] Length = 428
4713.1 Best-BlastP=> >nrprot No Hits found
4714.1 Best-BlastP=> >nrprot 47% Identities = 102/310 (32%), Positives = 159/310 (51 %), Gaps = 5/310 (1%) ref|NP_800300.1 | putative YhfP protein [Vibrio parahaemolyticus RIMD 2210633] dbj|BAC62133.11 putative YhfP protein [Vibrio parahaemolyticus] Length = 334
4715.3 Best-BlastP=> >nrprot 48% Identities = 43/158 (27%), Positives = 80/158 (50%), Gaps = 7/158 (4%) ref|ZP_00084605.11 hypothetical protein [Pseudomonas fluorescens PfO-1] Length = 171
4716.2 Best-BlastP=> >nrprot 48% Identities = 88/323 (27%), Positives = 162/323 (50%), Gaps = 34/323 (10%) ref |ZP_00144869.11 Magnesium and cobalt transport protein corA [Fusobacterium nucleatum subsp. vincentii ATCC 49256] gb|EAA23536.11 Magnesium and cobalt transport protein corA [Fusobacterium nucleatum subsp. vincentii ATCC 49256] Length = 351
4718.1 Best-BlastP=> >nrprot No Hits found 4719.2 Best-BlastP=> >nrprot 42% Identities = 149/405 (36%), Positives = 217/405 (53%), Gaps = 35/405 (8%) pir||T03487 potential multicopper oxidase - Rhodobacter capsulatus gb|AAC16140.11 potential multicopper oxidase [Rhodobacter capsulatus] Length = 491
472.1 Best-BlastP=> >nrprot 73% Identities = 107/205 (52%), Positives = 155/205 (75%) ref|NP_819344.11 guanylate kinase [Coxiella burnetii RSA 493] gb|AA089858.11 guanylate kinase [Coxiella burnetii RSA 493] Length = 206
4721.2 Best-BlastP=> >nrprot 70% Identities = 91/161 (56%), Positives = 125/161 (77%), Gaps = 1/161 (0%) ref|NP_440118.11 spore maturation protein A [Synechocystis sp. PCC 6803] pir||S74646 spore maturation protein A - Synechocystis sp. (strain PCC 6803) dbj|BAA16798.11 spore maturation protein A [Synechocystis sp. PCC 6803] Length = 182
4724.2 Best-BlastP=> >nrprot 70% Identities = 98/199 (49%), Positives = 142/199 (71%), Gaps = 5/199 (2%) ref|NP_440119.1 | spore maturation protein B [Synechocystis sp. PCC 6803] pir||S74647 spore maturation protein B - Synechocystis sp. (strain PCC 6803) dbj|BAA16799.11 spore maturation protein B [Synechocystis sp. PCC 6803] Length = 217
4725.2 Best-BlastP=> >nrprot 31% Identities = 73/267 (27%), Positives = 111/267 (41 %), Gaps = 62/267 (23%) ref|NP_230974.1 | hypothetical protein VC1330 [Vibrio cholerae 01 biovar eltor str. N16961] pir||B82212 hypothetical protein VC1330 [imported] - Vibrio cholerae (strain N16961 serogroup 01 ) gb|AAF94488.11 hypothetical protein [Vibrio cholerae 01 biovar eltor str. N16961] Length = 275
4726.1 Best-BlastP=> >nrprot No Hits found
4727.1 Best-BlastP=> >nrprot 42% Identities = 43/76 (56%), Positives = 57/76 (75%) ref|ZP_00101887.11 COG5394: Uncharacterized protein conserved in bacteria [Desulfitobacterium hafniense] Length = 107
4729.3 Best-BlastP=> >nrprot 64% Identities = 111/246 (45%), Positives = 161/246 (65%), Gaps = 1/246 (0%) ref|NP_902034.11 acetoacetyl-CoA reductase [Chromobacterium violaceum ATCC 12472] gb|AAQ60036.1 | acetoacetyl-CoA reductase [Chromobacterium violaceum ATCC 12472] Length = 246
473.2 Best-BlastP=> >nrprot 63% Identities = 122/288 (42%), Positives = 183/288 (63%) ref|NP_22g866.11 conserved hypothetical protein [Vibrio cholerae 01 biovar eltor str. N 16961] pir||C82350 conserved hypothetical protein VC0209 [imported] - Vibrio cholerae (strain N16961 serogroup 01) gb|AAF93385.1 | conserved hypothetical protein [Vibrio cholerae 01 biovar eltor str. N16961] Length = 288
4732.3 Best-BlastP=> >nrprot 33% Identities = 30/96 (31 %), Positives = 42/96 (43%), Gaps = 17/96 (17%) dbj|BAA89216.11 soluble cytochrome cA [Shewanella violacea] Length = 85
4733.2 Best-BlastP=> >nrprot 71 % Identities = 154/2g6 (52%), Positives = 210/296 (70%), Gaps = 8/296 (2%) ref|NP_820374.11 translation elongation factor Ts [Coxiella burnetii RSA 4 3] sp|Q9X5U9|EFTS_COXBU Elongation factor Ts (EF-Ts) gb|AAD33343.1 |AF127534_2 elongation factor Ts [Coxiella burnetii] gb|AAO90888.11 translation elongation factor Ts [Coxiella burnetii RSA 493] Length = 2 6
4734.2 Best-BlastP=> >nrprot 65% Identities = 134/341 (39%), Positives = 213/341 (62%), Gaps = 16/341 (4%) ref|NP_439856.11 hypothetical protein [Haemophilus influenzae Rd] sp|P45339|YJEQ_HAEIN Hypothetical protein HI1714 pir||B64176 hypothetical protein HI1714 - Haemophilus influenzae (strain Rd KW20) gb|AAC23359.1 | conserved hypothetical protein [Haemophilus influenzae Rd] Length = 346
4736.2 Best-BlastP=> >nrprot 70% Identities = 219/403 (54%), Positives = 289/403 (71 %), Gaps = 2/403 (0%) ref|NP_232075.11 tRNA nucleotidyltransferase [Vibrio cholerae 01 biovar eltor str. N16961] pir||D82076 tRNA nucleotidyltransferase VC2446 [imported] - Vibrio cholerae (strain N16961 serogroup 01 ) gb|AAF95588.1 | tRNA nucleotidyltransferase [Vibrio cholerae 01 biovar eltor str. N16961] Length = 403
4737.1 Best-BlastP=> >nrprot 7g% Identities = 122/178 (68%), Positives = 149/178 (83%) ref|ZP_00066572.1 | COG1949: Oligoribonuclease (3'->5' exoribonuclease) [Microbulbifer degradans 2-40] Length = 185
4739.1 Best-BlastP=> >nrprot 81 % Identities = 156/208 (75%), Positives = 180/208 (86%) ref|NP_841903.11 DUF208 [Nitrosomonas europaea ATCC 19718] emb|CAD85792.11 DUF208 [Nitrosomonas europaea ATCC 19718] Length = 215
474.2 Best-BlastP=> >nrprot 44% Identities = 24/107 (22%), Positives = 49/107 (45%) ref|NP_878011.11 Tn1546 transposase [Staphylococcus aureus] sp|Q06238|TNP6_ENTFC Transposase for transposon Tn1546 pir||A40628 probable transposase - Enterococcus faecium transposon Tn1546 gb|AAA65951.1 | transposase [Enterococcus faecium] gb|AAQ17155.1 | Tn1546 transposase [Staphylococcus aureus] Length = 988
4740.2 Best-BlastP=> >nrprot 62% Identities = 137/287 (47%), Positives = 193/287 (67%), Gaps = 1/287 (0%) ref|NP_478257.11 cation efflux system protein [Nostoc sp. PCC 7120] pir||AG2540 cation efflux system protein [imported] - Nostoc sp. (strain PCC 7120) plasmid pCC7120beta dbj|BAB77253.11 cation efflux system protein [Nostoc sp. PCC 7120] Length = 304
4742.3 Best-BlastP=> >nrprot 48% Identities = 85/305 (27%), Positives = 148/305 (48%), Gaps = 14/305 (4%) ref|NP_759924.11 Transcriptional regulator [Vibrio vulnificus CMCP6] gb|AAO09451.1 |AE016800_56 Transcriptional regulator [Vibrio vulnificus CMCP6] dbj|BAC95964.11 transcriptional regulator [Vibrio vulnificus YJ016] Length = 313
4744.3 Best-BlastP=> >nrprot 40% Identities = 66/257 (25%), Positives = 119/257 (46%), Gaps = 35/257 (13%) ref|ZP_00143092.11 hypothetical protein [Rickettsia sibirica] gb|EAA26501.11 unknown [Rickettsia sibirica] Length = 290
4746.1 Best-BlastP=> >nrprot 52% Identities = 34/80 (42%), Positives = 48/80 (60%), Gaps = 3/80 (3%) ref|NP_799393.11 putative signal peptide protein [Vibrio parahaemolyticus RIMD 2210633] dbj|BAC61277.1 | putative signal peptide protein [Vibrio parahaemolyticus] Length = 86
4748.1 Best-BlastP=> >nrprot 71 % Identities = 145/246 (58%), Positives = 182/246 (73%), Gaps = 2/246 (0%) ref|ZP_00028198.1 | COG4689: Acetoacetate decarboxylase [Burkholderia fungorum] Length = 442
4749.2 Best-BlastP=> >nrprot 39% Identities = 206/416 (49%), Positives = 284/416 (68%), Gaps = 3/416 (0%) ref|NP_386185.11 PUTATIVE NADH DEHYDROGENASE TRANSMEMBRANE PROTEIN [Sinorhizobium meliloti] emb|CAC46658.1 | PUTATIVE NADH DEHYDROGENASE TRANSMEMBRANE PROTEIN [Sinorhizobium meliloti] Length = 422
475.1 Best-BlastP=> >nrprot No Hits found
4751.2 Best-BlastP=> >nrprot 44% Identities = 161/483 (33%), Positives = 245/483 (50%), Gaps = 72/483 (14%) gb|AAN78225.11 class 4 metalloprotease [Chromobacterium violaceum] Length = 489
4752.2 Best-BlastP=> >nrprot 60% Identities = 120/275 (43%), Positives = 180/275 (65%) ref|ZP_00029901.11 COG0697: Permeases of the drug/metabolite transporter (DMT) superfamily [Burkholderia fungorum] Length = 341
4754.1 Best-BlastP=> >nrprot No Hits found
4755.2 Best-BlastP=> >nrprot 58% Identities = 74/170 (43%), Positives = 109/170 (64%) ref|NP_310486.11 hypothetical protein [Escherichia coli 0157:H7] pir||C90936 hypothetical protein ECs2459 [imported] - Escherichia coli (strain 0157:H7, substrain RIMD 0509952) dbj|BAB35882.1 | hypothetical protein [Escherichia coli 0157:H7] Length = 182
4757.2 Best-BlastP=> >nrprot 53% Identities = 60/192 (31 %), Positives = 105/192 (54%), Gaps = 3/192 (1 %) ref|ZP_00111879.11 COG0259: Pyridoxamine-phosphate oxidase [Nostoc punctiforme] Length = 214
4758.2 Best-BlastP=> >nrprot 51 % Identities = 44/g2 (47%), Positives = 66/92 (71 %) ref|NP_926536.11 unknown protein [Gloeobacter violaceus] dbj|BAC91531.11 gsl3590 [Gloeobacter violaceus] Length = 98
4759.2 Best-BlastP=> >nrprot 70% Identities = 142/258 (55%), Positives = 181/258 (70%), Gaps = 5/258 (1 %) ref|NP_863091.1 | putative oxidoreductase [Pseudomonas putida] gb|AA064293.1 | putative oxidoreductase [Pseudomonas putida] gb|AAP44207.1 | putative dehydrogenase [Pseudomonas sp. ND6] Length = 257
4760.2 Best-BlastP=> >nrprot No Hits found
4763.2 Best-BlastP=> >nrprot 52% Identities = 133/408 (32%), Positives = 219/408 (53%), Gaps = 23/408 (5%) ref|NP_903521.11 probable prophage integrase [Chromobacterium violaceum ATCC 12472] gb|AAQ61513.11 probable prophage integrase [Chromobacterium violaceum ATCC 12472] Length = 423
4764.2 Best-BlastP=> >nrprot No Hits found
4766.2 Best-BlastP=> >nrprot 74% Identities = 80/144 (55%), Positives = 108/144 (75%), Gaps = 1/144 (0%) ref|NP_820296.11 conserved hypothetical protein [Coxiella burnetii RSA 493] gb|AAO90810.11 conserved hypothetical protein [Coxiella burnetii RSA 493] Length = 146 4767.2 Best-BlastP=> >nrprot 62% Identities = 36/93 (38%), Positives = 57/93 (61%), Gaps = 3/93 (3%) ref|NP_273838.11 conserved hypothetical protein [Neisseria meningitidis MC58] ref|NP_283783.1 | hypothetical protein NMA1005 [Neisseria meningitidis Z2491] sp|Q9JRC2|YA05_NEIMA Hypothetical protein NMA1005/NMB0796 pir||C81157 conserved hypothetical protein NMB0796 [imported] - Neisseria meningitidis (strain MC58 serogroup B, strain Z2491 serogroup A) gb|AAF41209.11 conserved hypothetical protein [Neisseria meningitidis MC58] emb|CAB84274.11 hypothetical protein NMA1005 [Neisseria meningitidis Z2491] Length = 92
4768.1 Best-BlastP=> >nrprot 52% Identities = 35/101 (34%), Positives = 62/101 (61%), Gaps = 12/101 (11%) ref|ZP_00022930.11 COG2913: Small protein A (tmRNA-binding) [Ralstonia metallidurans] Length = 159
4769.1 Best-BlastP=> >nrprot 99% Identities = 135/136 (99%), Positives = 136/136 (100%) sp|Q48835|FUR_LEGPN Ferric uptake regulation protein (Ferric uptake regulator) gb|AAA19656.11 Fur Length = 136
4770.1 Best-BlastP=> >nrprot 68% Identities = 25/46 (54%), Positives = 33/46 (71 %), Gaps = 3/46 (6%) emb|CAB87569.11 FldC protein [Sphingomonas sp. LB126] Length = 533
4771.2 Best-BlastP=> >nrprot 26% Identities = 65/166 (39%), Positives = 94/166 (56%), Gaps = 7/166 (4%) ref|NP_907052.11 COMPONENTS OF SENSORY TRANSDUCTION SYSTEM [Wolinella succinogenes] emb|CAE09952.11 COMPONENTS OF SENSORY TRANSDUCTION SYSTEM [Wolinella succinogenes] Length = 407
4773.3 Best-BlastP=> >nrprot 77% Identities = 292/461 (63%), Positives = 362/461 (78%) ref|ZP_00015785.11 hypothetical protein [Rhodospirillum rubrum] sp|Q59765|PNTB_RHORU NAD(P) transhydrogenase subunit beta (Pyridine nucleotide transhydrogenase subunit beta) (Nicotinamide nucleotide transhydrogenase subunit beta) (Proton-translocating transhydrogenase NADP(H)-binding component) (dill) gb|AAC43257.1 | nicotinamide nucleotide transhydrogenase, subunit beta gb|AAA62495.1 | proton-translocating nicotinamide nucleotide transhydrogenase subunit PntB prf||2102322C energy-transducing nicotinamide nucleotide transhydrogenase: Length = 464
4774.1 Best-BlastP=> >nrprot 74% Identities = 61/91 (67%), Positives = 74/91 (81 %) ref|NP_840934.11 probable transmembrane NAD(P) transhydrogenase (alpha subunit part 2) [Nitrosomonas europaea ATCC 19718] emb|CAD84771.1 | probable transmembrane NAD(P) transhydrogenase (alpha subunit part 2) [Nitrosomonas europaea ATCC 19718] Length = 102
4775.2 Best-BlastP=> >nrprot 66% Identities = 179/361 (49%), Positives = 250/361 (69%), Gaps = 9/361 (2%) ref|NP_840933.11 Alanine dehydrogenase and pyridine nucleotide transhydrogenase [Nitrosomonas europaea ATCC 19718] emb|CAD84770.11 Alanine dehydrogenase and pyridine nucleotide transhydrogenase [Nitrosomonas europaea ATCC 19718] Length = 375
4776.3 Best-BlastP=> >nrprot 51 % Identities = 59/146 (40%), Positives = 89/146 (60%), Gaps = 8/146 (5%) ref|ZP_00111831.11 COG2340: Uncharacterized protein with SCP/PR1 domains [Nostoc punctiforme] Length = 182
4777.2 Best-BlastP=> >nrprot No Hits found
4778.1 Best-BlastP=> >nrprot No Hits found
4779.1 Best-BlastP=> >nrprot 62% Identities = 68/152 (44%), Positives = 93/152 (61 %), Gaps = 4/152 (2%) ref|ZP_00125177.11 COG2153: Predicted acyltransferase [Pseudomonas syringae pv. syringae B728a] Length = 152
4780.2 Best-BlastP=> >nrprot No Hits found
4782.3 Best-BlastP=> >nrprot 99% Identities = 359/363 (98%), Positives = 362/363 (99%) emb|CAB65197.11 hypothetical protein [Legionella pneumophila] Length = 363
4785.1 Best-BlastP=> >nrprot No Hits found
4786.2 Best-BlastP=> >nrprot 52% Identities = 159/409 (38%), Positives = 250/409 (61 %), Gaps = 8/409 (1 %) ref|NP_819388.11 d-xylose-proton symporter, putative [Coxiella burnetii RSA 493] gb|AAO89902.11 d-xylose-proton symporter, putative [Coxiella burnetii RSA 493] Length = 409
4788.3 Best-BlastP=> >nrprot 98% Identities = 200/204 (98%), Positives = 201/204 (98%) sp|P50024|DSBA_LEGPN THIOLDISULFIDE INTERCHANGE PROTEIN DSBA PRECURSOR gb|AAA67725.11 disulfide bond forming protein Length = 204
4790.3 Best-BlastP=> >nrprot 77% Identities = 278/425 (65%), Positives = 336/425 (79%), Gaps = 1/425 (0%) ref |NP_820750.11 ABC transporter, ATP- binding protein [Coxiella burnetii RSA 493] gb|AA091264.11 ABC transporter, ATP-binding protein [Coxiella burnetii RSA 493] Length = 432
4791.3 Best-BlastP=> >nrprot 35% Identities = 37/91 (40%), Positives = 47/91 (51%) pir||A71007 hypothetical protein PH1351 - Pyrococcus horikoshii dbj|BAA30457.1 | 101aa long hypothetical protein [Pyrococcus horikoshii] Length = 101
4793.2 Best-BlastP=> >nrprot 75% Identities = 306/545 (56%), Positives = 412/545 (75%) ref|NP_820751.11 ABC transporter, permease protein [Coxiella burnetii RSA 493] gb|AA091265.11 ABC transporter, permease protein [Coxiella burnetii RSA 493] Length = 581
4795.1 Best-BlastP=> >nrprot No Hits found
4796.1 Best-BlastP=> >nrprot 25% Identities = 24/70 (34%), Positives = 36/70 (51 %), Gaps = 12/70 (17%) ref|NP_650587.11 CG5225-PA [Drosophila melanogaster] gb|AAF55377.2| CG5225-PA [Drosophila melanogaster] Length = 594
4797.1 Best-BlastP=> >nrprot 91 % Identities = 293/351 (83%), Positives = 324/351 (92%) ref|NP_820537.11 ribonucleoside-diphosphate reductase, beta subunit [Coxiella burnetii RSA 493] gb|AA091051.11 ribonucleoside-diphosphate reductase, beta subunit [Coxiella burnetii RSA 493] Length = 401
48.1 Best-BlastP=> >nrprot 98% Identities = 90/93 (96%), Positives = 93/93 (100%) emb|CAB60052.11 lvhB3 [Legionella pneumophila] gb|AAM08239.11 putative type IV secretion protein B3 [Legionella pneumophila] Length = 93
480.3 Best-BlastP=> >nrprot 88% Identities = 373/475 (78%), Positives = 428/475 (90%), Gaps = 1/475 (0%) ref|NP_820349.11 conserved hypothetical protein [Coxiella burnetii RSA 493] gb|AAO90863.11 conserved hypothetical protein [Coxiella burnetii RSA 493] Length = 480
4800.2 Best-BlastP=> >nrprot 61% Identities = 102/252 (40%), Positives = 160/252 (63%), Gaps = 1/252 (0%) ref |ZP_00110131.11 COG0300: Short- chain dehydrogenases of various substrate specificities [Nostoc punctiforme] Length = 270
4801.1 Best-BlastP=> >nrprot 33% Identities = 31/122 (25%), Positives = 56/122 (45%), Gaps = 24/122 (19%) ref|ZP_00026008.1 | COG4970: Tfp pilus assembly protein FimT [Ralstonia metallidurans] Length = 222
4802.2 Best-BlastP=> >nrprot 31 % Identities = 34/110 (30%), Positives = 56/110 (50%), Gaps = 7/110 (6%) ref|ZP_00030895.11 COG3161 : 4- hydroxy benzoate synthetase (chorismate lyase) [Burkholderia fungorum] Length = 227
4803.2 Best-BlastP=> >nrprot 79% Identities = 148/247 (59%), Positives = 197/247 (79%) ref|NP_793605.11 3-oxoacyl-(acyl-carrier-protein) reductase [Pseudomonas syringae pv. tomato str. DC3000] gb|AAO57300.11 3-oxoacyl-(acyl-carrier-protein) reductase [Pseudomonas syringae pv. tomato str. DC3000] Length = 247
4804.1 Best-BlastP=> >nrprot 84% Identities = 65/73 (89%), Positives = 70/73 (95%) pir||T12021 acyl carrier protein - Pseudomonas aeruginosa gb|AAB94392.11 acyl carrier protein [Pseudomonas aeruginosa] Length = 78
4805.1 Best-BlastP=> >nrprot 78% Identities = 258/411 (62%), Positives = 325/411 (79%) ref|NP_251655.11 beta-ketoacyl-acyl carrier protein synthase II [Pseudomonas aeruginosa PA01] pir||T12022 3-oxoacyl-[acyl-carher-protein] synthase (EC 2.3.1.41 ) II - Pseudomonas aeruginosa gb|AAB94396.1 | 3-oxoacyl-acyl carrier protein synthase II [Pseudomonas aeruginosa] gb|AAG06353.1 |AE004722_9 beta-ketoacyl- acyl carrier protein synthase II [Pseudomonas aeruginosa PA01] Length = 414 4806.4 Best-BlastP=> >nrprot 60% Identities = 127/330 (38%), Positives = 200/330 (60%), Gaps = 14/330 (4%) ref |NP_819532.11 conserved hypothetical protein TIGR00247 [Coxiella burnetii RSA 493] gb|AAO90046.11 conserved hypothetical protein TIGR00247 [Coxiella burnetii RSA 493] Length = 370
4809.2 Best-BlastP=> >nrprot 46% Identities = 85/287 (29%), Positives = 130/287 (45%), Gaps = 15/287 (5%) ref|ZP_00013543.11 hypothetical protein [Rhodospirillum rubrum] Length = 352
481.1 Best-BlastP=> >nrprot 57% Identities = 62/150 (41 %), Positives = 88/150 (58%) ref|NP_298766.11 conserved hypothetical protein [Xylella fastidiosa 9a5c] pir||H82676 conserved hypothetical protein XF1477 [imported] - Xylella fastidiosa (strain 9a5c) gb|AAF84286.1 |AE003977_9 conserved hypothetical protein [Xylella fastidiosa 9a5c] Length = 153
4810.1 Best-BlastP=> >nrprot No Hits found
4811.2 Best-BlastP=> >nrprot 67% Identities = 89/163 (54%), Positives = 110/163 (67%), Gaps = 1/163 (0%) ref|NP_404518.11 putative membrane protein [Yersinia pestis] pir||AE01 10 probable membrane protein YPO0899 [imported] - Yersinia pestis (strain C092) emb|CAC89744.11 putative membrane protein [Yersinia pestis C092] Length = 165
4813.2 Best-BlastP=> >nrprot 63% Identities = 161/389 (41 %), Positives = 246/389 (63%), Gaps = 4/389 (1 %) gb|AAD47247.11 putative transport protein [Legionella pneumophila] Length = 387
4814.2 Best-BlastP=> >nrprot 65% Identities = 101/199 (50%), Positives = 138/199 (69%), Gaps = 5/199 (2%) ref |NP_820137.11 enhanced entry protein EnhA, putative [Coxiella burnetii RSA 493] gb|AAO90651.11 enhanced entry protein EnhA, putative [Coxiella burnetii RSA 493] Length = 248 4815.2 Best-BlastP=> >nrprot 59% Identities = 56/115 (48%), Positives = 82/115 (71 %), Gaps = 1/115 (0%) ref|NP_883806.11 flagellar protein FliS [Bordetella parapertussis] emb|CAE36818.1 | flagellar protein FliS [Bordetella parapertussis] Length = 142
4817.4 Best-BlastP=> >nrprot No Hits found 4818.3 Best-BlastP=> >nrprot No Hits found
rt rt ,- rt co ^ d rt m cb -~: f-1 oo co co co rt rt TO TO TO TO co co TO rt rt rt rt rt
485.3 Best-BlastP=> >nrprot 41% Identities = 106/439 (24%), Positives = 199/439 (45%), Gaps = 41/439 (9%) ref|NP_932208.11 putative conjugative transfer protein TraB [Vibrio vulnificus YJ016] dbj|BAC97731.11 putative conjugative transfer protein TraB [Vibrio vulnificus YJ016] Length = 603
4850.2 Best-BlastP=> >nrprot No Hits found
4851.2 Best-BlastP=> >nrprot 67% Identities = 89/161 (55%), Positives = 109/161 (67%), Gaps = 2/161 (1%) ref|ZP_00054128.1 | COG3837: Uncharacterized conserved protein, contains double-stranded beta-helix domain [Magnetospirillum magnetotacticum] Length = 180 4855.2 Best-BlastP=> >nrprot 80% Identities = 67/91 (73%), Positives = 79/91 (86%) ref|NP_717690.11 integration host factor, alpha subunit [Shewanella oneidensis MR-1] gb|AAN55134.1 |AE015650_4 integration host factor, alpha subunit [Shewanella oneidensis MR-1] Length = 98 4856.2 Best-BlastP=> >nrprot 67% Identities = 51/100 (51 %), Positives = 72/100 (72%) ref |NP_841896.11 putative ferredoxin 2fe-2s protein [Nitrosomonas europaea ATCC 19718] emb|CAD85785.11 putative ferredoxin 2fe-2s protein [Nitrosomonas europaea ATCC 19718] Length = 102 4859.2 Best-BlastP=> >nrprot No Hits found 486.2 Best-BlastP=> >nrprot No Hits found
4860.1 Best-BlastP=> >nrprot No Hits found 4863.4 Best-BlastP=> >nrprot 69% Identities = 220/430 (51 %), Positives = 294/430 (68%), Gaps = 13/430 (3%) ref|NP_406861.11 poly(A) polymerase [Yersinia pestis] pir||AI0412 polynucleotide adenylyltransferase (EC 2.7.7.19) [imported] - Yersinia pestis (strain C092) emb|CAC92629.11 poly(A) polymerase [Yersinia pestis C092] Length = 440
4864.2 Best-BlastP=> >nrprot 66% Identities = 62/131 (47%), Positives = 88/131 (67%) ref|NP_735626.11 Unknown [Streptococcus agalactiae NEM316] emb|CAD46839.11 Unknown [Streptococcus agalactiae NEM316] Length = 162
4865.2 Best-BlastP=> >nrprot 58% Identities = 48/146 (32%), Positives = 83/146 (56%), Gaps = 12/146 (8%) ref|NP_841101.11 Universal stress protein (Usp) [Nitrosomonas europaea ATCC 19718] emb|CAD84939.1 | Universal stress protein (Usp) [Nitrosomonas europaea ATCC 19718] Length = 148
4867.2 Best-BlastP=> >nrprot 62% Identities = 209/445 (46%), Positives = 293/445 (65%), Gaps = 3/445 (0%) ref|ZP_00025176.1 | COG1012: NAD- dependent aldehyde dehydrogenases [Ralstonia metallidurans] Length = 520
4868.2 Best-BlastP=> >nrprot 72% Identities = 155/269 (57%), Positives = 197/269 (73%), Gaps = 1/269 (0%) pir||T34105 hypothetical protein C17G10.8 - Caenorhabditis elegans Length = 938
4869.3 Best-BlastP=> >nrprot 75% Identities = 90/139 (64%), Positives = 108/139 (77%) ref|NP_540374.1 | RIBONUCLEASE HI [Brucella melitensis] ref|NP_697505.11 ribonuclease H [Brucella suis 1330] sp|Q8YFR3|RNH_BRUME Ribonuclease H (RNase H) pir||AC3434 calf thymus ribonuclease H (EC 3.1.26.4) [imported] - Brucella melitensis (strain 16M) gb|AAL52638.11 RIBONUCLEASE HI [Brucella melitensis 16M] gb|AAN29420.1 |AE014357_5 ribonuclease H [Brucella suis 1330] Length = 154
4872.3 Best-BlastP=> >nrprot 98% Identities = 511/520 (98%), Positives = 515/520 (99%) gb|AAM00608.11 unknown [Legionella pneumophila] Length = 520 4873.3 Best-BlastP=> >nrprot 99% Identities = 383/387 (98%), Positives = 385/387 (99%) gb|AAM00609.11 unknown [Legionella pneumophila] Length = 388
4874.2 Best-BlastP=> >nrprot 41% Identities = 108/445 (24%), Positives = 193/445 (43%), Gaps = 79/445 (17%) ref|NP_621958.1 | ATPase involved in DNA repair [Thermoanaerobacter tengcongensis] gb|AAM23562.11 ATPase involved in DNA repair [Thermoanaerobacter tengcongensis] Length = 1177
4875.1 Best-BlastP=> >nrprot 69% Identities = 73/119 (61 %), Positives = 91/119 (76%), Gaps = 6/119 (5%) ref|ZP_00033092.1 | COG0316: Uncharacterized conserved protein [Burkholderia fungorum] Length = 121
4877.2 Best-BlastP=> >nrprot 74% Identities = 202/356 (56%), Positives = 269/356 (75%), Gaps = 1/356 (0%) ref |NP_746144.11 tRNA (5- methylaminomethyl-2-thiouridylate)-methyltransferase [Pseudomonas putida KT2440] gb|AAN69608.1 |AE016594_5 tRNA (5- methylaminomethyl-2-thiouridylate)-methyltransferase [Pseudomonas putida KT2440] Length = 374
4878.2 Best-BlastP=> >nrprot 54% Identities = 161/418 (38%), Positives = 242/418 (57%), Gaps = 6/418 (1 %) ref|ZP_00086772.11 COG0739: Membrane proteins related to metalloendopeptidases [Pseudomonas fluorescens PfO-1] Length = 471
488.2 Best-BlastP=> >nrprot 75% Identities = 181/298 (60%), Positives = 230/298 (77%) ref|ZP_00068320.11 COG1131 : ABC-type multidrug transport system, ATPase component [Microbulbifer degradans 2-40] Length = 318
4880.2 Best-BlastP=> >nrprot 49% Identities = 92/270 (34%), Positives = 136/270 (50%), Gaps = 9/270 (3%) ref|NP_085189.11 IS10 orf [Shigella flexneri] ref |NP_858160.11 hypothetical protein [Shigella flexneri 2a] gb|AAK18345.1 |AF348706_34 IS10 orf [Shigella flexneri] gb|AAL72480.1 | hypothetical protein [Shigella flexneri 2a] Length = 407
4881.4 Best-BlastP=> >nrprot 11% Identities = 72/284 (25%), Positives = 121/284 (42%), Gaps = 19/284 (6%) ref|ZP_00112010.11 hypothetical protein [Nostoc punctiforme] Length = 427
4883.2 Best-BlastP=> >nrprot 50% Identities = 45/105 (42%), Positives = 67/105 (63%), Gaps = 1/105 (0%) ref|NP_819268.11 preprotein translocase, SecE subunit [Coxiella burnetii RSA 493] gb|AA089782.11 preprotein translocase, SecE subunit [Coxiella burnetii RSA 493] Length = 127
4884.1 Best-BlastP=> >nrprot 80% Identities = 125/175 (71 %), Positives = 148/175 (84%) ref|ZP_00123792.11 COG0250: Transcription antiterminator [Pseudomonas syringae pv. syringae B728a] ref|NP_790461.11 transcription antitermination protein NusG [Pseudomonas syringae pv. tomato str. DC3000] gb|AA054156.11 transcription antitermination protein NusG [Pseudomonas syringae pv. tomato str. DC3000] Length = 177
4885.2 Best-BlastP=> >nrprot 87% Identities = 112/143 (78%), Positives = 127/143 (88%) ref|NP_282gg6.1 1 50S ribosomal protein L11 [Neisseria meningitidis Z2491] pir||H82007 50S ribosomal protein L11 NMA0146 [imported] - Neisseria meningitidis (strain Z2491 serogroup A) emb|CAB83461 .11 50S ribosomal protein L11 [Neisseria meningitidis Z2491] Length = 144
4887.3 Best-BlastP=> >nrprot 57% Identities = 72/176 (40%), Positives = 102/176 (57%), Gaps = 6/176 (3%) ref|NP_419908.11 acetyltransferase, GNAT family [Caulobacter crescentus CB15] pir||H87384 acetyltransferase, GNAT family [imported] - Caulobacter crescentus gb|AAK23076.1 | acetyltransferase, GNAT family [Caulobacter crescentus CB15] Length = 181
4888.1 Best-BlastP=> >nrprot 59% Identities = 63/151 (41 %), Positives = 103/151 (68%) emb|CAD31280.11 PUTATIVE TRANSCRIPTION REGULATOR PROTEIN [Mesorhizobium loti] Length = 151
4889.1 Best-BlastP=> >nrprot 49% Identities = 44/120 (36%), Positives = 66/120 (55%), Gaps = 3/120 (2%) ref|NP_819252.11 hypothetical protein [Coxiella burnetii RSA 493] gb|AA089766.11 hypothetical protein [Coxiella burnetii RSA 493] Length = 130
489.1 Best-BlastP=> >nrprot No Hits found
4891.2 Best-BlastP=> >nrprot 26% Identities = 39/138 (28%), Positives = 67/138 (48%), Gaps = 19/138 (13%) pir||A59234 slow myosin heavy chain 3 - quail gb| AAC59911.11 slow myosin heavy chain 3 gb|AAC59912.11 slow myosin heavy chain 3 Length = 1931
4894.1 Best-BlastP=> >nrprot 69% Identities = 52/121 (42%), Positives = 81/121 (66%), Gaps = 7/121 (5%) ref|NP_746309.11 succinate dehydrogenase, hydrophobic membrane anchor protein [Pseudomonas putida KT2440] gb|AAN69773.1 |AE016613_8 succinate dehydrogenase, hydrophobic membrane anchor protein [Pseudomonas putida KT2440] Length = 122
4895.1 Best-BlastP=> >nrprot 69% Identities = 60/124 (48%), Positives = 87/124 (70%) ref|NP_250272.11 succinate dehydrogenase (C subunit) [Pseudomonas aeruginosa PA01] pir||C83448 succinate dehydrogenase (C subunit) PA1581 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG04970.1 |AE004586_8 succinate dehydrogenase (C subunit) [Pseudomonas aeruginosa PA01] Length = 128
4897.2 Best-BlastP=> >nrprot 47% Identities = 103/370 (27%), Positives = 179/370 (48%), Gaps = 22/370 (5%) ref|NP_761717.11 Predicted signal transduction protein [Vibrio vulnificus CMCP6] gb|AA011244.1 |AE016806_234 Predicted signal transduction protein [Vibrio vulnificus CMCP6] Length = 404
4900.3 Best-BlastP=> >nrprot 41 % Identities = 205/972 (21 %), Positives = 412/972 (42%), Gaps = 110/972 (11 %) ref|NP_245295.11 unknown [Pasteurella multocida] gb|AAK02442.1 | unknown [Pasteurella multocida] Length = 1113
4901.3 Best-BlastP=> >nrprot 76% Identities = 357/592 (60%), Positives = 454/592 (76%), Gaps = 1/592 (0%) ref|NP_842254.11 aspartyl-tRNA synthetase [Nitrosomonas europaea ATCC 19718] emb|CAD86164.11 aspartyl-tRNA synthetase [Nitrosomonas europaea ATCC 19718] Length = 593
4902.1 Best-BlastP=> >nrprot 25% Identities = 23/43 (53%), Positives = 28/43 (65%) ref|NP_275122.11 hypothetical protein [Neisseria meningitidis MC58] pir||G81001 hypothetical protein NMB2137 [imported] - Neisseria meningitidis (strain MC58 serogroup B) gb|AAF42445.1 | hypothetical protein [Neisseria meningitidis MC58] Length = 70
4907.2 Best-BlastP=> >nrprot 72% Identities = 134/257 (52%), Positives = 186/257 (72%), Gaps = 9/257 (3%) ref|ZP_00067917.11 COG1054: Predicted sulfurtransferase [Microbulbifer degradans 2-40] Length = 309
4908.3 Best-BlastP=> >nrprot 58% Identities = 715/1981 (36%), Positives = 1108/1981 (55%), Gaps = 111/1981 (5%) ref |NP_406102.11 putative membrane protein [Yersinia pestis] ref|NP_668470.1 | conserved hypothetical protein [Yersinia pestis KIM] sp|Q8ZDJ2|YP73_YERPE Hypothetical UPF0192 protein YP02573/Y1143 precursor pir||AC0314 probable membrane protein YP02573 [imported] - Yersinia pestis (strain C092) emb|CAC91375.1 | putative membrane protein [Yersinia pestis C092] gb|AAM84721.1 |AE013717_3 conserved hypothetical protein [Yersinia pestis KIM] Length = 2004
491.1 Best-BlastP=> >nrprot 34% Identities = 62/271 (22%), Positives = 116/271 (42%), Gaps = 20/271 (7%) sp|O60610|DIA1_HUMAN Diaphanous protein homolog 1 (Diaphanous-related formin 1 ) (DRF1) gb|AAC05373.1 | diaphanous 1 [Homo sapiens] Length = 1248
4913.2 Best-BlastP=> >nrprot No Hits found
4914.2 Best-BlastP=> >nrprot No Hits found
4916.2 Best-BlastP=> >nrprot 30% Identities = 53/186 (28%), Positives = 93/186 (50%), Gaps = 11/186 (5%) ref|NP_179611.11 leucine rich repeat protein family [Arabidopsis thaliana] pir||D84586 hypothetical protein At2g20210 [imported] - Arabidopsis thaliana gb|AAD21766.1 | hypothetical protein [Arabidopsis thaliana] Length = 271
4917.2 Best-BlastP=> >nrprot No Hits found
4918.2 Best-BlastP=> >nrprot 17% Identities = 31/114 (27%), Positives = 52/114 (45%), Gaps = 4/114 (3%) ref|NP_281741.1 | putative integral membrane protein [Campylobacter jejuni] pir||E81402 probable integral membrane protein Cj0557c [imported] - Campylobacter jejuni (strain NCTC 11168) emb|CAB75193.11 putative integral membrane protein [Campylobacter jejuni subsp. jejuni NCTC 11168] Length = 361
4919.1 Best-BlastP=> >nrprot 44% Identities = 109/421 (25%), Positives = 192/421 (45%), Gaps = 24/421 (5%) ref |NP_900103.11 outer membrane efflux protein [Chromobacterium violaceum ATCC 12472] gb|AAQ58111.11 outer membrane efflux protein [Chromobacterium violaceum ATCC 12472] Length = 466
492.4 Best-BlastP=> >nrprot 59% Identities = 48/101 (47%), Positives = 70/101 (69%), Gaps = 1/101 (0%) ref|NP_761137.11 SM-20-related protein [Vibrio vulnificus CMCP6] gb|AAO10664.1 |AE016804_174 SM-20-related protein [Vibrio vulnificus CMCP6] dbj|BAC94823.1 | SM-20-related protein [Vibrio vulnificus YJ016] Length = 200
4920.4 Best-BlastP=> >nrprot 56% Identities = 88/284 (30%), Positives = 143/284 (50%), Gaps = 38/284 (13%) ref|NP_791175.11 sensory box/GGDEF domain/EAL domain protein [Pseudomonas syringae pv. tomato str. DC3000] gb|AAO54870.11 sensory box/GGDEF domain/EAL domain protein [Pseudomonas syringae pv. tomato str. DC3000] Length = 763
4923.2 Best-BlastP=> >nrprot 79% Identities = 433/634 (68%), Positives = 507/634 (79%) ref|ZP_00066531.11 COG0441 : Threonyl-tRNA synthetase [Microbulbifer degradans 2-40] Length = 636
4927.1 Best-BlastP=> >nrprot No Hits found
4929.2 Best-BlastP=> >nrprot 72% Identities = 164/298 (55%), Positives = 223/298 (74%), Gaps = 2/298 (0%) ref|NP_622015.11 uncharacterized enzyme involved in pigment biosynthesis [Thermoanaerobacter tengcongensis] gb|AAM23619.1 | uncharacterized enzyme involved in pigment biosynthesis [Thermoanaerobacter tengcongensis] Length = 307
493.4 Best-BlastP=> >nrprot 55% Identities = 83/216 (38%), Positives = 131/216 (60%) ref|NP_634315.11 Zinc metalloprotease [Methanosarcina mazei Goel] gb|AAM31987.1 | Zinc metalloprotease [Methanosarcina mazei Goel] Length = 238
4930.2 Best-BlastP=> >nrprot 78% Identities = 51/90 (56%), Positives = 71/90 (78%) ref|NP_819953.11 conserved hypothetical protein [Coxiella burnetii RSA 493] sp|Q83D06|Y941_COXBU Hypothetical UPF0269 protein CBU0941 gb|AAO90467.1 | conserved hypothetical protein [Coxiella burnetii RSA 493] Length = 90
4934.3 Best-BlastP=> >nrprot 26% Identities = 59/198 (29%), Positives = 101/198 (51%), Gaps = 27/198 (13%) ref|NP_628263.11 possible secreted peptidase [Streptomyces coelicolor A3(2)] emb|CAB56362.11 possible secreted peptidase [Streptomyces coelicolor A3(2)] Length = 279
4935.2 Best-BlastP=> >nrprot 63% Identities = 168/345 (48%), Positives = 225/345 (65%) ref|NP_216774.11 hypothetical protein Rv2258c [Mycobacterium tuberculosis H37Rv] ref|NP_336787.1 | methyltransferase-related protein [Mycobacterium tuberculosis CDC1551] ref|NP_855931.11 Possible transcriptional regulatory protein [Mycobacterium bovis subsp. bovis AF2122/97] pir||F70862 probable helix-turn helix motif at aa 47-68 - Mycobacterium tuberculosis (strain H37RV) emb|CAA17295.11 hypothetical protein Rv2258c [Mycobacterium tuberculosis H37Rv] gb|AAK46601.1 | methyltransferase-related protein [Mycobacterium tuberculosis CDC1551] emb|CAD97135.1 | Possible transcriptional regulatory protein [Mycobacterium bovis subsp. bovis AF2122/97] Length = 353
494.2 Best-BlastP=> >nrprot 75% Identities = 160/283 (56%), Positives = 212/283 (74%), Gaps = 5/283 (1 %) ref|NP_518195.11 PROBABLE METALLOPROTEASE ZINC TRANSMEMBRANE PROTEIN [Ralstonia solanacearum] sp|Q8Y3A6|HTPX_RALSO Probable protease htpX homolog emb|CAD13602.11 PROBABLE METALLOPROTEASE ZINC TRANSMEMBRANE PROTEIN [Ralstonia solanacearum] Length = 286
4941.2 Best-BlastP=> >nrprot No Hits found
4942.1 Best-BlastP=> >nrprot No Hits found
4943.2 Best-BlastP=> >nrprot No Hits found
4945.2 Best-BlastP=> >nrprot 79% Identities = 214/340 (62%), Positives = 268/340 (78%), Gaps = 11/340 (3%) ref|NP_052843.11 hypothetical protein [Coxiella burnetii] ref|NP_819053.1 | repB protein, putative [Coxiella burnetii RSA 493] pir||S52723 qsopB protein - Coxiella burnetii plasmid QpH1 gb|AAA69865.1 | qsopB gene product emb|CAA59788.1 | orf 334 [Coxiella burnetii] emb|CAA75818.1 | hypothetical protein [Coxiella burnetii] gb|AAD33475.1 |AF131076_1 hypothetical protein [Coxiella burnetii] gb|AA091613.1 | repB protein, putative [Coxiella burnetii RSA 493] prf||2117254B trans-acting factor Length = 334
4947.2 Best-BlastP=> >nrprot 93% Identities = 340/402 (84%), Positives = 377/402 (93%) ref|NP_052336.11 unnamed protein product [Coxiella burnetii] ref|NP_052844.11 hypothetical protein [Coxiella burnetii] ref |NP_819052.11 parA protein, putative [Coxiella burnetii RSA 493] pir||S68866 qsopA protein - Coxiella burnetii plasmid QpH1 emb|CAA53106.1 | unnamed protein product [Coxiella burnetii] gb|AAA69864.1 | qsopA gene product emb|CAA59789.1 | orf 406 [Coxiella burnetii] emb|CAA75819.1 | putative SopA protein (protein a) [Coxiella burnetii] gb|AAD33476.1 |AF131076_2 hypothetical protein [Coxiella burnetii] gb|AA091612.11 parA protein, putative [Coxiella burnetii RSA 493] prf||2117254A trans-acting factor Length = 406
4948.4 Best-BlastP=> >nrprot No Hits found
4951.2 Best-BlastP=> >nrprot 24% Identities = 25/79 (31 %), Positives = 40/79 (50%), Gaps = 4/79 (5%) ref|ZP_00103190.11 COG1396: Predicted transcriptional regulators [Desulfitobacterium hafniense] Length = 123
4952.1 Best-BlastP=> >nrprot No Hits found
4954.2 Best-BlastP=> >nrprot 53% Identities = 62/155 (40%), Positives = 92/155 (59%) ref|NP_746668.11 polypeptide deformylase [Pseudomonas putida KT2440] sp|Q88EA7|DEF2_PSEPK Peptide deformylase 2 (PDF 2) (Polypeptide deformylase 2) gb|AAN70132.1 |AE016653_3 polypeptide deformylase [Pseudomonas putida KT2440] Length = 178
4955.4 Best-BlastP=> >nrprot 50% Identities = 132/417 (31%), Positives = 218/417 (52%), Gaps = 9/417 (2%) ref |NP_221200.11 PROLINE/BETAINE TRANSPORTER (proP6) [Rickettsia prowazekii] pir||D71647 proline/betaine transporter (proP6) RP852 - Rickettsia prowazekii emb|CAA15276.11 PROLINE/BETAINE TRANSPORTER (proP6) [Rickettsia prowazekii] Length = 415
4957.2 Best-BlastP=> >nrprot 62% Identities = 53/108 (49%), Positives = 74/108 (68%), Gaps = 3/108 (2%) ref|NP_251650.11 type 4 fimbrial biogenesis • protein PilZ [Pseudomonas aeruginosa PA01] pir||B59241 type 4 fimbriae biogenesis protein [imported] - Pseudomonas aeruginosa gb|AAA93519.11 involved in biogenesis of type 4 fimbriae gb|AAG06348.1 |AE004722_4 type 4 fimbrial biogenesis protein PilZ [Pseudomonas aeruginosa PA01] Length = 118
4958.3 Best-BlastP=> >nrprot 47% Identities = 90/313 (28%), Positives = 142/313 (45%), Gaps = 20/313 (6%) ref|NP_819534.11 DNA polymerase III, delta prime subunit [Coxiella burnetii RSA 493] gb|AAO90048.11 DNA polymerase III, delta prime subunit [Coxiella burnetii RSA 493] Length = 319 496.1 Best-BlastP=> >nrprot No Hits found
4960.2 Best-BlastP=> >nrprot No Hits found
4961.1 Best-BlastP=> >nrprot 77% Identities = 70/127 (55%), Positives = 99/127 (77%), Gaps = 4/127 (3%) ref|NP_391517.11 similar to large conductance mechanosensitive channel protein [Bacillus subtilis] sp|P94585|MSCL_BACSU Large-conductance mechanosensitive channel pir||E70065 large conductance mechanosensitive channel homolog ywpC - Bacillus subtilis emb|CAB05944.11 ywpC [Bacillus subtilis] emb|CAB15653.11 large conductance mechanosensitive channel protein [Bacillus subtilis subsp. subtilis str. 168] Length = 130
4962.2 Best-BlastP=> >nrprot No Hits found 4965.2 Best-BlastP=> >nrprot 61 % Identities = 99/105 (94%), Positives = 102/105 (97%) emb|CAB65201.1 | hypothetical protein [Legionella pneumophila] Length = 356
4966.2 Best-BlastP=> >nrprot 98% Identities = 586/598 (97%), Positives = 590/598 (98%) emb|CAB65200.11 hypothetical protein [Legionella pneumophila] Length = 598
4968.2 Best-BlastP=> >nrprot 63% Identities = 60/128 (46%), Positives = 83/128 (64%) ref | N P_439721.11 alanine racemase biosynthetic [Haemophilus influenzae Rd] sp|P45257|ALR_HAEIN Alanine racemase pir||E64130 alanine racemase (EC 5.1.1.1 ), biosynthetic - Haemophilus influenzae (strain Rd KW20) gb|AAC23218.1 | alanine racemase, biosynthetic (air) [Haemophilus influenzae Rd] Length = 360
4969.3 Best-BlastP=> >nrprot 81% Identities = 117/163 (71%), Positives = 141/163 (86%), Gaps = 1/163 (0%) ref |NP_719448.11 replicative DNA helicase [Shewanella oneidensis MR-1] gb|AAN56892.1 |AE015824_3 replicative DNA helicase [Shewanella oneidensis MR-1] Length = 468
497.1 Best-BlastP=> >nrprot No Hits found 4972.3 Best-BlastP=> >nrprot 68% Identities = 111/195 (56%), Positives = 146/195 (74%) ref|ZP_00091033.11 COG0305: Replicative DNA helicase [Azotobacter vinelandii] Length = 463
4974.1 Best-BlastP=> >nrprot 40% Identities = 43/138 (31 %), Positives = 71/138 (51%), Gaps = 6/138 (4%) pir||T18332 icmL protein - Legionella pneumophila gb|AAC38190.1 | Dotl [Legionella pneumophila] emb|CAA75329.1 | IcmL protein [Legionella pneumophila] emb|CAD43145.1 | Dotl protein [Legionella pneumophila serogroup 6] Length = 212
4977.3 Best-BlastP=> >nrprot No Hits found
4979.1 Best-BlastP=> >nrprot 71 % Identities = 1 15/192 (59%), Positives = 144/192 (75%) ref|NP_246450.1 1 unknown [Pasteurella multocida] sp|P57947|ENGB_PASMU Probable GTP-binding protein engB gb|AAK03595.1 1 unknown [Pasteurella multocida] Length = 205
498.4 Best-BlastP=> >nrprot 37% Identities = 78/305 (25%), Positives = 125/305 (40%), Gaps = 56/305 (18%) ref|NP_421401.11 amine oxidase, flavin- containing [Caulobacter crescentus CB15] pir||E87571 amine oxidase, flavin-containing [imported] - Caulobacter crescentus gb|AAK24569.11 amine oxidase, flavin-containing [Caulobacter crescentus CB15] Length = 454
4980.3 Best-BlastP=> >nrprot 69% Identities = 102/206 (49%), Positives = 139/206 (67%), Gaps = 7/206 (3%) ref|NP_759874.11 Cytochrome c4 [Vibrio vulnificus CMCP6] gb|AAO09401.1 |AE016800_6 Cytochrome c4 [Vibrio vulnificus CMCP6] Length = 205
4984.2 Best-BlastP=> >nrprot 73% Identities = 109/166 (65%), Positives = 126/166 (75%), Gaps = 1/166 (0%) gb|AA038281.1 | Lfe115p1 [Leptospirillum ferrooxidans] Length = 178
4987.2 Best-BlastP=> >nrprot 77% Identities = 274/490 (55%), Positives = 360/490 (73%), Gaps = 28/490 (5%) ref|NP_457051.11 putative GTP-binding protein [Salmonella enterica subsp. enterica serovar Typhi] ref|NP_804211.11 putative GTP-binding protein [Salmonella enterica subsp. enterica serovar Typhi Ty2] sp|Q8Z4P6|ENGA_SALTI Probable GTP-binding protein engA pir||AF0821 probable GTP-binding protein STY2764 [imported] - Salmonella enterica subsp. enterica serovar Typhi. (strain CT18) emb|CAD02722.1 | putative GTP-binding protein [Salmonella enterica subsp. enterica serovar Typhi] gb|AAO68060.11 putative GTP-binding protein [Salmonella enterica subsp. enterica serovar Typhi Ty2] Length = 490
499.2 Best-BlastP=> >nrprot 51% Identities = 129/343 (37%), Positives = 196/343 (57%), Gaps = 10/343 (2%) ref|ZP_00106589.11 COG1680: Beta- lactamase class C and other penicillin binding proteins [Nostoc punctiforme] Length = 393
4990.3 Best-BlastP=> >nrprot No Hits found 4992.2 Best-BlastP=> >nrprot No Hits found
4993.2 Best-BlastP=> >nrprot 36% Identities = 44/89 (49%), Positives = 63/89 (70%) ref|ZP_00102874.11 hypothetical protein [Desulfitobacterium hafniense] Length = 106
4995.1 Best-BlastP=> >nrprot 33% Identities = 42/179 (23%), Positives = 73/179 (40%), Gaps = 36/179 (20%) ref|NP_229450.11 alpha-amylase, putative [Thermotoga maritima] pir||G72227 hypothetical protein TM1650 - Thermotoga maritima (strain MSB8) gb|AAD36717.1 |AE001807_8 alpha-amylase, putative [Thermotoga maritima] Length = 422
4998.2 Best-BlastP=> >nrprot 16% Identities = 47/200 (23%), Positives = 87/200 (43%), Gaps = 16/200 (8%) ref|NP_036450.11 leucine zipper-EF-hand containing transmembrane protein 1 [Homo sapiens] gb|AAD13138.11 leucine zipper-EF-hand containing transmembrane protein 1 [Homo sapiens] gb|AAH14500.1 | Leucine zipper-EF-hand containing transmembrane protein 1 [Homo sapiens] gb|AAH21208.1 | Leucine zipper- EF-hand containing transmembrane protein 1 [Homo sapiens] Length = 739
4999.2 Best-BlastP=> >nrprot 77% Identities = 157/241 (65%), Positives = 197/241 (81 %), Gaps = 4/241 (1 %) ref|NP_252346.11 30S ribosomal protein S2 [Pseudomonas aeruginosa PA01] sp|O82850|RS2_PSEAE 30S ribosomal protein S2 pir||C83189 30S ribosomal protein S2 PA3656 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG07044.1 |AE004785_8 30S ribosomal protein S2 [Pseudomonas aeruginosa PA01] Length = 246
50.1 Best-BlastP=> >nrprot 84% Identities = 76/92 (82%), Positives = 80/92 (86%) emb|CAB60051.11 lvhB2 [Legionella pneumophila] gb|AAM08238.11 putative pilin subunit [Legionella pneumophila] Length = 96
500.2 Best-BlastP=> >nrprot 97% Identities = 341/352 (96%), Positives = 346/352 (98%) gb|AAG59860.1 |AF299349_1 major acid phosphatase [Legionella pneumophila] Length = 352 -
5000.3 Best-BlastP=> >nrprot 97% Identities = 131/136 (96%), Positives = 134/136 (98%) emb|CAB09802.11 16 kD immunogenic protein [Legionella pneumophila] Length = 136
5001.1 Best-BlastP=> >nrprot No Hits found
5003.2 Best-BlastP=> >nrprot No Hits found
5005.3 Best-BlastP=> >nrprot 99% Identities = 454/455 (99%), Positives = 455/455 (100%) gb|AAQ18124.11 CpxA [Legionella pneumophila] Length = 455
501.2 Best-BlastP=> >nrprot 30% Identities = 40/167 (23%), Positives = 75/167 (44%), Gaps = 13/167 (7%) dbj|BAC45194.1 | kinesin-like protein [Oryza sativa (japonica cultivar-group)] Length = 1967
5010.2 Best-BlastP=> >nrprot 18% Identities = 34/117 (29%), Positives = 54/117 (46%), Gaps = 12/117 (10%) emb|CAD90592.1 | C3L protein [Cowpox virus] Length = 833
5015.2 Best-BlastP=> >nrprot 58% Identities = 43/110 (39%), Positives = 66/110 (60%), Gaps = 3/110 (2%) ref|ZP_00056081.11 hypothetical protein [Magnetospirillum magnetotacticum] Length = 164
5018.2 Best-BlastP=> >nrprot No Hits found
5019.1 Best-BlastP=> >nrprot 61% Identities = 87/190 (45%), Positives = 125/190 (65%) gb|AAG10504.1 |AF279106_66 predicted YacE family of P-loop kinases [uncultured marine gamma proteobacterium EBAC31A08] Length = 197
502.3 Best-BlastP=> >nrprot No Hits found
5021.3 Best-BlastP=> >nrprot 48% Identities = 64/247 (25%), Positives = 120/247 (48%), Gaps = 8/247 (3%) ref|NP_903494.11 conserved hypothetical protein [Chromobacterium violaceum ATCC 12472] gb|AAQ61486.11 conserved hypothetical protein [Chromobacterium violaceum ATCC 12472] Length = 252
5022.2 Best-BlastP=> >nrprot 58% Identities = 124/336 (36%), Positives = 204/336 (60%), Gaps = 5/336 (1 %) ref|NP_742592.11 membrane protein, putative [Pseudomonas putida KT2440] gb|AAN66056.1 |AE016234_9 membrane protein, putative [Pseudomonas putida KT2440] Length = 346
5026.1 Best-BlastP=> >nrprot 77% Identities = 49/89 (55%), Positives = 70/89 (78%) ref|NP_250138.11 flagellar biosynthetic protein FliQ [Pseudomonas aeruginosa PA01] ref |ZP_00139064.1 | COG1987: Flagellar biosynthesis pathway, component FliQ [Pseudomonas aeruginosa UCBPP- PA14] pir||A83465 flagellar biosynthetic protein FliQ PA1447 [imported] - Pseudomonas aeruginosa (strain PA01) gb|AAG04836.1 |AE004574_7 flagellar biosynthetic protein FliQ [Pseudomonas aeruginosa PA01] Length = 89
5028.2 Best-BlastP=> >nrprot 70% Identities = 270/528 (51 %), Positives = 367/528 (69%), Gaps = 5/528 (0%) ref|NP_820198.11 conserved hypothetical protein [Coxiella burnetii RSA 493] gb|AAO90712.11 conserved hypothetical protein [Coxiella burnetii RSA 493] Length = 550
503.3 Best-BlastP=> >nrprot 55% Identities = 162/399 (40%), Positives = 247/399 (61 %), Gaps = 3/399 (0%) ref|ZP_00065233.11 COG0741 : Soluble lytic murein transglycosylase and related regulatory proteins (some contain LysM/invasin domains) [Microbulbifer degradans 2-40] Length = 543
5030.2 Best-BlastP=> >nrprot 84% Identities = 225/326 (69%), Positives = 276/326 (84%) ref|NP_820836.11 peptide ABC transporter, permease protein [Coxiella burnetii RSA 493] gb|AA091350.11 peptide ABC transporter, permease protein [Coxiella burnetii RSA 493] Length = 327
5031.2 Best-BiastP=> >nrprot 40% Identities = 42/173 (24%), Positives = 75/173 (43%), Gaps = 12/173 (6%) ref|NP_542876.11 hypothetical protein [Pseudomonas putida] emb|CAC86816.1 | hypothetical protein [Pseudomonas putida] Length = 267
5032.1 Best-BlastP=> >nrprot No Hits found 5033.1 Best-BlastP=> >nrprot No Hits found 5037.1 Best-BlastP=> >nrprot 84% Identities = 35/43 (81 %), Positives = 38/43 (88%) ref|NP_30005g.11 50S ribosomal protein L34 [Xylella fastidiosa 9a5c] ref|NP_780293.1 | 50S ribosomal protein L34 [Xylella fastidiosa Temeculal] sp|Q9P9T9|RL34_XYLFA 50S ribosomal protein L34 pir||B82517 50S ribosomal protein L34 XF2782 [imported] - Xylella fastidiosa (strain 9a5c) gb|AAF85567.1 |AE004083_6 50S ribosomal protein L34 [Xylella fastidiosa 9a5c] gb|AA029942.11 50S ribosomal protein L34 [Xylella fastidiosa Temeculal] Length = 46
5038-1 Best-BlastP=> >nrprot 57% Identities = 45/107 (42%), Positives = 66/107 (61%), Gaps = 2/107 (1%) ref|NP_290337.1 | RNase P, protein component; protein C5; processes tRNA, 4.5S RNA [Escherichia coli 0157:H7 EDL933] ref|NP_312666.11 ribonuclease P protein component [Escherichia coli 0157:H7] sp|Q8XB43|RNPA_EC057 Ribonuclease P protein component (RNaseP protein) (RNase P protein) (Protein C5) pir||G91208 ribonuclease P protein component [imported] - Escherichia coli (strain 0157:H7, substrain RIMD 0509952) pir||A86055 hypothetical protein rnpA [imported] - Escherichia coli (strain 0157:H7, substrain EDL933) gb|AAG58901.1 |AE005601_7 RNase P, protein component; protein C5; processes tRNA, 4.5S RNA [Escherichia coli 0157:H7 EDL933] dbj|BAB38062.11 ribonuclease P protein component [Escherichia coli 0157:H7] Length = 119
5039.2 Best-BlastP=> >nrprot 62% Identities = 38/68 (55%), Positives = 51/68 (75%) ref|ZP_00134326.11 COG0759: Uncharacterized conserved protein [Actinobacillus pleuropneumoniae serovar 1 str. 4074] Length = 100
5040.2 Best-BlastP=> >nrprot 68% Identities = 268/565 (47%), Positives = 381/565 (67%), Gaps = 23/565 (4%) ref|NP_820897.11 inner-membrane protein, 60kDa [Coxiella burnetii RSA 493] sp|P45650|60IM_COXBU 60 kDa inner-membrane protein homolog gb|AA091411.1 | inner-membrane protein, 60kDa [Coxiella burnetii RSA 493] Length = 566
5041.2 Best-BlastP=> >nrprot 98% Identities = 176/178 (98%), Positives = 176/178 (98%) sp|034955|IPYR_LEGPN Inorganic pyrophosphatase (Pyrophosphate phospho-hydrolase) (PPase) gb|AAB84257.1 | inorganic pyrophosphatase [Legionella pneumophila] gb|AAC02428.1 | inorganic pyrophosphatase [Legionella pneumophila] Length = 178
5042.1 Best-BlastP=> >nrprot No Hits found
5043.3 Best-BlastP=> >nrprot 70% Identities = 210/415 (50%), Positives = 298/415 (71 %), Gaps = 2/415 (0%) ref|NP_760503.1 | Putative Mg2+ and Co2+ transporter CorB [Vibrio vulnificus CMCP6] gb|AA010030.1 |AE016802_73 Putative Mg2+ and Co2+ transporter CorB [Vibrio vulnificus CMCP6] Length = 425
5044.2 Best-BlastP=> >nrprot 73% Identities = 166/299 (55%), Positives = 217/299 (72%), Gaps = 13/299 (4%) ref|NP_763465.11 Conserved hypothetical protein [Vibrio vulnificus CMCP6] gb|AAO08455.1 |AE016813_207 Conserved hypothetical protein [Vibrio vulnificus CMCP6] Length = 304
5046.2 Best-BlastP=> >nrprot 53% Identities = 45/118 (38%), Positives = 73/118 (61 %), Gaps = 5/118 (4%) ref|XP_132330.11 RIKEN cDNA 2810006K23 [Mus musculus] gb|AAH46909.11 Similar to RIKEN cDNA 2810006K23 gene [Mus musculus] Length = 184
5047.3 Best-BlastP=> >nrprot 41 % Identities = 43/167 (25%), Positives = 80/167 (47%), Gaps = 6/167 (3%) ref|ZP_00067126.11 COG3009: Uncharacterized protein conserved in bacteria [Microbulbifer degradans 2-40] Length = 217
505.3 Best-BlastP=> >nrprot No Hits found
5050.2 Best-BlastP=> >nrprot 99% Identities = 241/243 (99%), Positives = 242/243 (99%) emb|CAD90952.11 LssA protein [Legionella pneumophila] Length = 243
5052.4 Best-BlastP=> >nrprot 97% Identities = 194/201 (96%), Positives = 197/201 (98%) emb|CAD90953.11 LssZ protein [Legionella pneumophila] Length = 204
5054.2 Best-BlastP=> >nrprot 33% Identities = 77/161 (47%), Positives = 108/161 (67%) ref|NP_390248.11 yqkA [Bacillus subtilis] sp|P54564|YQKA_BACSU Hypothetical protein yqkA pir||C69966 hypothetical protein yqkA - Bacillus subtilis dbj|BAA12633.11 YqkA [Bacillus subtilis] emb|CAB14299.11 yqkA [Bacillus subtilis subsp. subtilis str. 168] Length = 343
5056.2 Best-BlastP=> >nrprot 59% Identities = 57/101 (56%), Positives = 67/101 (66%), Gaps = 1/101 (0%) ref |ZP_00091135.11 COG2852: Uncharacterized protein conserved in bacteria [Azotobacter vinelandii] Length = 150
5058.2 Best-BlastP=> >nrprot 57% Identities = 29/66 (43%), Positives = 46/66 (69%), Gaps = 1/66 (1 %) dbj|BAA75251.11 Similar to IS1301 of Neisseria meningitidis [Actinobacillus actinomycetemcomitans] Length = 255
5059.3 Best-BlastP=> >nrprot 44% Identities = 33/70 (47%), Positives = 46/70 (65%) pir||S61903 hypothetical protein 1 - Neisseria meningitidis emb|CAA88914.11 orfl [Neisseria meningitidis] Length = 151
506.3 Best-BlastP=> >nrprot No Hits found
5060.2 Best-BlastP=> >nrprot 99% Identities = 504/505 (99%), Positives = 504/505 (99%) emb|CAB65195.11 hypothetical protein [Legionella pneumophila] Length = 505
5061.4 Best-BlastP=> >nrprot 87% Identities = 468/562 (83%), Positives = 492/562 (87%), Gaps = 16/562 (2%) emb|CAB65194.11 hypothetical protein [Legionella pneumophila] Length = 548
5062.3 Best-BlastP=> >nrprot 18% Identities = 24/71 (33%), Positives = 39/71 (54%) gb|AAA21525.11 meiotin-1 Length = 259 5064.2 Best-BlastP=> >nrprot No Hits found
5065.2 Best-BlastP=> >nrprot 34% Identities = 30/100 (30%), Positives = 54/100 (54%), Gaps = 11/100 (11 %) ref|NP_593310.1 | F-box protein [Schizosaccharomyces pombe] sp|P87053|POF1_SCHPO F-box/WD-repeat protein pofl (Skp1 -binding protein 1 ) pir||T38932 probable sulfur metabolite control protein - fission yeast (Schizosaccharomyces pombe) emb|CAB08168.11 SPAC57A10.05c [Schizosaccharomyces pombe] dbj|BAA84528.1 | Pof1 [Schizosaccharomyces pombe] Length = 605
5066.2 Best-BlastP=> >nrprot 57% Identities = 50/121 (41 %), Positives = 80/121 (66%), Gaps = 5/121 (4%) ref|NP_487806.11 two-component response regulator [Nostoc sp. PCC 7120] pir||AG2276 two-component response regulator all3766 [imported] - Nostoc sp. (strain PCC 7120) dbj|BAB75465.11 two-component response regulator [Nostoc sp. PCC 7120] Length = 143
5068.3 Best-BlastP=> >nrprot 51% Identities = 52/194 (26%), Positives = 98/194 (50%), Gaps = 8/194 (4%) ref |ZP_00110196.11 hypothetical protein [Nostoc punctiforme] Length = 223
5069.3 Best-BlastP=> >nrprot 53% Identities = 48/149 (32%), Positives = 74/149 (49%), Gaps = 25/149 (16%) ref|NP_812673.1 | arginine repressor, transcriptional regulator of arginine metabolism [Bacteroides thetaiotaomicron VPI-5482] gb|AA078867.11 arginine repressor, transcriptional regulator of arginine metabolism [Bacteroides thetaiotaomicron VPI-5482] Length = 157
507.3 Best-BlastP=> >nrprot 46% Identities = 18/32 (56%), Positives = 22/32 (68%) gb|AAN04217.11 putative transposase Tnp [Aeromonas salmonicida] Length = 383
5071.2 Best-BlastP=> >nrprot No Hits found 5072.2 Best-BlastP=> >nrprot 26% Identities = 30/85 (35%), Positives = 41/85 (48%) ref|NP_473229.2| putative protein kinase [Plasmodium falciparum 3D7] emb|CAA15620.3| putative protein kinase [Plasmodium falciparum 3D7] Length = 2515
5075.4 Best-BlastP=> >nrprot 11 % Identities = 62/172 (36%), Positives = 102/172 (59%), Gaps = 18/172 (10%) ref|NP_703336.1 | P. falciparum RESA- like protein with DnaJ domain [Plasmodium falciparum 3D7] emb|CAD48951.1 | P. falciparum RESA-like protein with DnaJ domain [Plasmodium falciparum 3D7] Length = 1451
5076.2 Best-BlastP=> >nrprot 54% Identities = 50/158 (31 %), Positives = 87/158 (55%), Gaps = 10/158 (6%) ref|NP_6341 g6.11 Hydrogenase expression/formation protein [Methanosarcina mazei Goel] emb|CAA62962.1 | F420-nonreducing hydrogenase II [Methanosarcina mazei] gb|AAM31868.1 | Hydrogenase expression/formation protein [Methanosarcina mazei Goel] Length = 161
5077.3 Best-BlastP=> >nrprot 60% Identities = 198/424 (46%), Positives = 262/424 (61 %), Gaps = 2/424 (0%) ref|ZP_00089783.11 COG3259: Coenzyme F420-reducing hydrogenase, alpha subunit [Azotobacter vinelandii] Length = 442
5078.2 Best-BlastP=> >nrprot 76% Identities = 250/432 (57%), Positives = 329/432 (76%), Gaps = 3/432 (0%) ref|NP_052842.11 hypothetical protein [Coxiella burnetii] gb|AAD33508.1 |AF131076_34 hypothetical protein [Coxiella burnetii] Length = 433
5080.4 Best-BlastP=> >nrprot 44% Identities = 121/443 (27%), Positives = 217/443 (48%), Gaps = 26/443 (5%) ref|NP_903425.1 1 probable peptide transporter protein [Chromobacterium violaceum ATCC 12472] gb|AAQ61417.1 | probable peptide transporter protein [Chromobacterium violaceum ATCC 12472] Length = 495
5081.3 Best-BlastP=> >nrprot 50% Identities = 62/173 (35%), Positives = 96/173 (55%) ref|NP_520208.11 CONSERVED HYPOTHETICAL PROTEIN [Ralstonia solanacearum] emb|CAD15794.1 1 CONSERVED HYPOTHETICAL PROTEIN [Ralstonia solanacearum] Length = 194
5082.3 Best-BlastP=> >nrprot No Hits found
5084.4 Best-BlastP=> >nrprot 30% Identities = 94/351 (26%), Positives = 160/351 (45%), Gaps = 28/351 (7%) ref|ZP_00087727.11 hypothetical protein [Pseudomonas fluorescens PfO-1] Length = 375
5087.2 Best-BlastP=> >nrprot 19% Identities = 37/151 (24%), Positives = 69/151 (45%), Gaps = 1 1/151 (7%) ref|NP_70541 1.11 hypothetical protein, conserved [Plasmodium falciparum 3D7] emb|CAD52648.11 hypothetical protein, conserved [Plasmodium falciparum 3D7] Length = 2533
5088.2 Best-BlastP=> >nrprot 53% Identities = 49/131 (37%), Positives = 76/131 (58%), Gaps = 4/131 (3%) ref|NP_832446.11 Acetyltransferase [Bacillus cereus ATCC 14579] gb|AAP09647.11 Acetyltransferase [Bacillus cereus ATCC 14579] Length = 141
509.1 Best-BlastP=> >nrprot 50% Identities = 69/172 (40%), Positives = 99/172 (57%), Gaps = 10/172 (5%) ref|NP_903527.11 conserved hypothetical protein [Chromobacterium violaceum ATCC 12472] gb|AAQ61519.2| conserved hypothetical protein [Chromobacterium violaceum ATCC 12472] Length = 576
5090.2 Best-BlastP=> >nrprot 59% Identities = 148/359 (41 %), Positives = 211/359 (58%), Gaps = 12/359 (3%) ref|NP_662343.1 | glutamate 5-kinase [Chlorobium tepidum TLS] sp|Q8KCG4|PROB_CHLTE Glutamate 5-kinase (Gamma-glutamyl kinase) (GK) gb|AAM72685.1 | glutamate 5-kinase [Chlorobium tepidum TLS] Length = 361
5092.3 Best-BlastP=> >nrprot No Hits found
5093.3 Best-BlastP=> >nrprot 52% Identities = 97/243 (39%), Positives = 128/243 (52%), Gaps = 2/243 (0%) ref|NP_820472.11 UDP-2,3- diacylglucosamine hydrolase [Coxiella burnetii RSA 493] gb|AAO90986.1 | UDP-2,3-diacylglucosamine hydrolase [Coxiella burnetii RSA 493] Length = 243
5094.2 Best-BlastP=> >nrprot 55% Identities = 93/209 (44%), Positives = 129/209 (61 %), Gaps = 13/209 (6%) ref|ZP_00085068.11 COG0850: Septum formation inhibitor [Pseudomonas fluorescens PfO-1] Length = 245
5097.1 Best-BlastP=> >nrprot 46% Identities = 76/295 (25%), Positives = 139/295 (47%), Gaps = 7/295 (2%) ref|NP_355292.11 AGR_C_4240p [Agrobacterium tumefaciens] ref|NP_533007.1 | conserved hypothetical protein [Agrobacterium tumefaciens str. C58 (U. Washington)] pir||D97640 hypothetical protein AGR_C_4240 [imported] - Agrobacterium tumefaciens (strain C58, Cereon) pir||AE2863 conserved hypothetical protein Atu2334 [imported] - Agrobacterium tumefaciens (strain C58, Dupont) gb|AAK88077.11 AGR_C_4240p [Agrobacterium tumefaciens str. C58 (Cereon)] gb|AAL43323.1 | conserved hypothetical protein [Agrobacterium tumefaciens str. C58 (U. Washington)] Length = 302
5098.2 Best-BlastP=> >nrprot No Hits found
510.2 Best-BlastP=> >nrprot No Hits found
5100.2 Best-BlastP=> >nrprot 42% Identities = 101/389 (25%), Positives = 173/389 (44%), Gaps = 16/389 (4%) ref |NP_444149.11 Y4xM [Rhizobium sp. NGR234] sp|P55705|Y4XM_RHISN HYPOTHETICAL TRANSPORT PROTEIN Y4XM gb|AAB91936.11 Y4xM [Rhizobium sp. NGR234] Length = 404
5103.4 Best-BlastP=> >nrprot 42% Identities = 25/61 (40%), Positives = 37/61 (60%), Gaps = 1/61 (1%) gb|AAG10082.1 |AF295331_2 outer membrane lipoprotein Pep [Edwardsiella tarda] Length = 155
5104.3 Best-BlastP=> >nrprot 48% Identities = 203/509 (39%), Positives = 293/509 (57%), Gaps = 9/509 (1 %) ref|NP_896011.11 FAD linked oxidase, N- terminal [Prochlorococcus marinus str. MIT 9313] emb|CAE22361.11 FAD linked oxidase, N-terminal [Prochlorococcus marinus str. MIT 9313] Length = 571
5106.3 Best-BlastP=> >nrprot 32% Identities = 54/242 (22%), Positives = 111/242 (45%), Gaps = 24/242 (9%) gb|EAA15516.11 hypothetical protein [Plasmodium yoelii yoelii] Length = 585
5108.2 Best-BlastP=> >nrprot No Hits found
5113.3 Best-BlastP=> >nrprot No Hits found 5114.3 Best-BlastP=> >nrprot No Hits found
5115.2 Best-BlastP=> >nrprot 44% Identities = 77/284 (27%), Positives = 141/284 (49%), Gaps = 19/284 (6%) ref |ZP_00128740.1 | COG0454: Histone acetyltransferase HPA2 and related acetyltransferases [Desulfovibrio desulfuricans G20] Length = 326
5116.3 Best-BlastP=> >nrprot 56% Identities = 71/173 (41%), Positives = 107/173 (61%) ref|NP_800289.1 | hypothetical protein VPA0779 [Vibrio parahaemolyticus RIMD 2210633] dbj|BAC62122.11 hypothetical protein [Vibrio parahaemolyticus] Length = 203
5123.3 Best-BlastP=> >nrprot 57% Identities = 44/78 (56%), Positives = 54/78 (69%) ref|NP_819379.11 DNA-binding protein Fis [Coxiella burnetii RSA 493] gb|AA089893.11 DNA-binding protein Fis [Coxiella burnetii RSA 493] Length = 103
5124.3 Best-BlastP=> >nrprot 78% Identities = 96/151 (63%), Positives = 120/151 (79%) ref|NP_820809.11 ribose-phosphate pyrophosphokinase [Coxiella burnetii RSA 493] gb|AA091323.11 ribose-phosphate pyrophosphokinase [Coxiella burnetii RSA 493] Length = 319
5127.4 Best-BlastP=> >nrprot 77% Identities = 117/162 (72%), Positives = 141/162 (87%) ref|ZP_00068148.1 | COG0462: Phosphoribosylpyrophosphate synthetase [Microbulbifer degradans 2-40] Length = 316
5129.4 Best-BlastP=> >nrprot 76% Identities = 607/1050 (57%), Positives = 799/1050 (76%), Gaps = 12/1050 (1 %) ref|NP_773366.11 AcrB/AcrD/AcrF family protein [Bradyrhizobium japonicum] dbj|BAC51991.11 AcrB/AcrD/AcrF family protein [Bradyrhizobium japonicum USDA 110] Length = 1052
5132.3 Best-BlastP=> >nrprot No Hits found
5133.3 Best-BlastP=> >nrprot No Hits found
5134.4 Best-BlastP=> >nrprot 99% Identities = 308/309 (99%), Positives = 309/309 (100%) gb|AAN63820.11 lysophospholipase A [Legionella pneumophila] Length = 309
5135.2 Best-BlastP=> >nrprot No Hits found
514.5 Best-BlastP=> >nrprot 16% Identities = 124/559 (22%), Positives = 243/559 (43%), Gaps = 100/559 (17%) gb|AAB00143.11 putative Length = 1015 5146.2 Best-BlastP=> >nrprot 55% Identities = 154/418 (36%), Positives = 238/418 (56%), Gaps = 13/418 (3%) gb|AAC44538.11 ProP [Escherichia coli] Length = 500
5147.1 Best-BlastP=> >nrprot No Hits found
5151.1 Best-BlastP=> >nrprot 56% Identities = 109/295 (36%), Positives = 167/295 (56%), Gaps = 9/295 (3%) ref|NP_346934.11 MccF-like protein [Clostridium acetobutylicum] pir||G96935 mccF-like protein [imported] - Clostridium acetobutylicum gb|AAK78274.1 |AE007544_3 MccF-like protein [Clostridium acetobutylicum] Length = 306
5152.1 Best-BlastP=> >nrprot 55% Identities = 56/126 (44%), Positives = 72/126 (57%), Gaps = 2/126 (1 %) ref|NP_107051.1 | unknown protein [Mesorhizobium loti] dbj|BAB52837.1 | unknown protein [Mesorhizobium loti] Length = 274
5153.1 Best-BlastP=> >nrprot 97% Identities = 141/145 (97%), Positives = 142/145 (97%) gb|AAK00280.1 |AF288536_2 unknown [Legionella longbeachae] Length = 145
5154.2 Best-BlastP=> >nrprot 93% Identities = 253/273 (92%), Positives = 259/273 (94%) gb|AAK00279.1 |AF288536_1 spectinomycin 3' adenylyltransferase [Legionella longbeachae] Length = 274
5156.1 Best-BlastP=> >nrprot No Hits found
5159.2 Best-BlastP=> >nrprot 62% Identities = 240/471 (50%), Positives = 313/471 (66%), Gaps = 7/471 (1 %) ref|NP_900831 .11 probable melitin resistence protein [Chromobacterium violaceum ATCC 12472] gb|AAQ58836.1 | probable melitin resistence protein [Chromobacterium violaceum ATCC 12472] Length = 495
5162.1 Best-BlastP=> >nrprot No Hits found 5164.1 Best-BlastP=> >nrprot No Hits found
5167.4 Best-BlastP=> >nrprot 69% Identities = 49/91 (53%), Positives = 68/91 (74%) gb|AAP83334.1 |AF469614_2 unknown [Francisella tularensis subsp. tularensis] Length = 94
5173.1 Best-BlastP=> >nrprot 41 % Identities = 195/643 (30%), Positives = 282/643 (43%), Gaps = 44/643 (6%) ref|NP_251563.11 hypothetical protein [Pseudomonas aeruginosa PA01] pir||F83287 hypothetical protein PA2873 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG06261.1 |AE004713_10 hypothetical protein PA2873 [Pseudomonas aeruginosa PA01] Length = 668
5174.2 Best-BlastP=> >nrprot No Hits found
5176.1 Best-BlastP=> >nrprot No Hits found
5177.2 Best-BlastP=> >nrprot 41 % Identities = 61/168 (36%), Positives = 95/168 (56%), Gaps = 1/168 (0%) ref |ZP_00144063.1 | Outer membrane protein [Fusobacterium nucleatum subsp. vincentii ATCC 49256] gb|EAA24326.11 Outer membrane protein [Fusobacterium nucleatum subsp. vincentii ATCC 49256] Length = 202
5178.1 Best-BlastP=> >nrprot 98% Identities = 162/163 (99%), Positives = 162/163 (99%) gb|AAC38179.1 | DotD [Legionella pneumophila] Length = 163
5180.1 Best-BlastP=> >nrprot 98% Identities = 74/75 (98%), Positives = 75/75 (100%) gb|AAN17184.1 |AF492466_2 ferrous iron transporter A [Legionella pneumophila] Length = 75
5188.2 Best-BlastP=> >nrprot 70% Identities = 205/420 (48%), Positives = 305/420 (72%), Gaps = 2/420 (0%) ref|ZP_00025967.11 COG1301 : Na+/H+- dicarboxylate symporters [Ralstonia metallidurans] Length = 467
5189.2 Best-BlastP=> >nrprot No Hits found
519.3 Best-BlastP=> >nrprot 99% Identities = 549/550 (99%), Positives = 549/550 (99%) pir||A41468 60K heat shock protein htpB - Legionella pneumophila Length = 550
5193.2 Best-BlastP=> >nrprot 10% Identities = 49/180 (27%), Positives = 75/180 (41 %), Gaps = 40/180 (22%) ref|NP_229450.1 | alpha-amylase, putative [Thermotoga maritima] pir||G72227 hypothetical protein TM 1650 - Thermotoga maritima (strain MSB8) gb|AAD36717.1 |AE001807_8 alpha-amylase, putative [Thermotoga maritima] Length = 422
5194.1 Best-BlastP=> >nrprot 52% Identities = 74/157 (47%), Positives = 98/157 (62%), Gaps = 5/157 (3%) ref|ZP_00036504.11 COG0046: Phosphoribosylformylglycinamidine (FGAM) synthase, synthetase domain [Enterococcus faecium] Length = 738
5195.2 Best-BlastP=> >nrprot 69% Identities = 237/447 (53%), Positives = 312/447 (69%), Gaps = 4/447 (0%) ref|NP_718290.11 succinylarginine dihydrolase [Shewanella oneidensis MR-1] gb|AAN55734.1 |AE015710_2 succinylarginine dihydrolase [Shewanella oneidensis MR-1] Length = 444 52.1 Best-BlastP=> >nrprot 88% Identities = 52/66 (78%), Positives = 60/66 (90%) emb|CAB60050.11 IvrC [Legionella pneumophila] Length = 67
520.1 Best-BlastP=> >nrprot 62% Identities = 67/139 (48%), Positives = 92/139 (66%) ref|NP_440670.11 hypothetical protein [Synechocystis sp. PCC 6803] sp|P73321 |YI94_SYNY3 Hypothetical protein slr1894 pir||S77503 hypothetical protein slr1894 - Synechocystis sp. (strain PCC 6803) dbj|BAA17350.11 ORF_ID:slr1894~hypothetical protein [Synechocystis sp. PCC 6803] Length = 156
5200.2 Best-BlastP=> >nrprot 77% Identities = 489/756 (64%), Positives = 587/756 (77%), Gaps = 5/756 (0%) ref|NP_820975.11 DNA topoisomerase I [Coxiella burnetii RSA 493] gb|AA091489.11 DNA topoisomerase I [Coxiella burnetii RSA 493] Length = 765
5201.2 Best-BlastP=> >nrprot 73% Identities = 122/217 (56%), Positives = 165/217 (76%), Gaps = 2/217 (0%) sp|066188|SCNC_THITI Thiocyanate hydrolase gamma subunit dbj|BAA28288.11 thiocyanate hydrolase gamma subunit [Thiobacillus thioparus] Length = 243
5202.2 Best-BlastP=> >nrprot 55% Identities = 46/92 (50%), Positives = 57/92 (61 %), Gaps = 2/92 (2%) sp|066187|SCNA_THITI Thiocyanate hydrolase alpha subunit dbj|BAA28287.11 thiocyanate hydrolase alpha subunit [Thiobacillus thioparus] Length = 126
5204.2 Best-BlastP=> >nrprot 46% Identities = 56/110 (50%), Positives = 74/110 (67%), Gaps = 1/110 (0%) sp|066186|SCNB_THITI Thiocyanate hydrolase beta subunit dbj|BAA28286.11 thiocyanate hydrolase beta subunit [Thiobacillus thioparus] Length = 157
5206.1 Best-BlastP=> >nrprot 73% Identities = 212/360 (58%), Positives = 272/360 (75%), Gaps = 2/360 (0%) ref|ZP_00138670.11 COG1706: Flagellar basal-body P-ring protein [Pseudomonas aeruginosa UCBPP-PA14] Length = 440
5208.1 Best-BlastP=> >nrprot 55% Identities = 34/86 (39%), Positives = 57/86 (66%), Gaps = 1/86 (1 %) ref |NP_924146.11 hypothetical protein gsl1200 [Gloeobacter violaceus] dbj|BAC89141.11 gsl1200 [Gloeobacter violaceus] Length = 96
521.2 Best-BlastP=> >nrprot 55% Identities = 239/722 (33%), Positives = 382/722 (52%), Gaps = 53/722 (7%) ref|NP_773225.11 bll6585 [Bradyrhizobium japonicum] dbj|BAC51850.11 bll6585 [Bradyrhizobium japonicum USDA 110] Length = 861
5216.2 Best-BlastP=> >nrprot 64% Identities = 127/282 (45%), Positives = 181/282 (64%), Gaps = 9/282 (3%) ref|NP_643189.11 pirin [Xanthomonas axonopodis pv. citri str. 306] gb|AAM37725.1 | pirin [Xanthomonas axonopodis pv. citri str. 306] Length = 285
5217.1 Best-BlastP=> >nrprot 76% Identities = 78/118 (66%), Positives = 95/118 (80%) ref|ZP_00024696.1 | COG1484: DNA replication protein [Ralstonia metallidurans] Length = 268
5219.1 Best-BlastP=> >nrprot 50% Identities = 98/254 (38%), Positives = 135/254 (53%), Gaps = 11/254 (4%) gb|AAM90719.11 TraN [Salmonella typhi] Length = 617
5220.1 Best-BlastP=> >nrprot 26% Identities = 41/119 (34%), Positives = 58/119 (48%), Gaps = 16/119 (13%) ref|NP_052852.11 hypothetical protein [Coxiella burnetii] pir||S52231 hypothetical protein 160 - Coxiella burnetii emb|CAA59944.1 | orf 160 [Coxiella burnetii] gb|AAD33484.1 |AF131076_10 hypothetical protein [Coxiella burnetii] Length = 160
5224.2 Best-BlastP=> >nrprot No Hits found 5226.2 Best-BlastP=> >nrprot 76% Identities = 117/173 (67%), Positives = 138/173 (79%) ref|NP_744614.1 | translation initiation factor IF-3 [Pseudomonas putida KT2440] gb|AAN68078.1 |AE016439_13 translation initiation factor IF-3 [Pseudomonas putida KT2440] Length = 177
5227.2 Best-BlastP=> >nrprot 68% Identities = 34/63 (53%), Positives = 46/63 (73%), Gaps = 1/63 (1 %) sp|P13069|RL35_BACST 50S ribosomal protein L35 pir||R5BS35 ribosomal protein L35 - Bacillus stearothermophilus emb|CAA34313.1 | unnamed protein product [Geobacillus stearothermophilus] Length = 66
5229.1 Best-BlastP=> >nrprot 81 % Identities = 108/142 (76%), Positives = 118/142 (83%) ref|NP_230221.1 | ribosomal protein L13 [Vibrio cholerae 01 biovar eltor str. N16961] pir||B82308 ribosomal protein L13 VC0570 [imported] - Vibrio cholerae (strain N16961 serogroup 01 ) gb|AAF93738.1 | ribosomal protein L13 [Vibrio cholerae 01 biovar eltor str. N16961] Length = 142
523.2 Best-BlastP=> >nrprot 48% Identities = 87/267 (32%), Positives = 144/267 (53%), Gaps = 1/267 (0%) ref |NP_252216.11 probable outer membrane protein [Pseudomonas aeruginosa PA01] pir||D83204 probable outer membrane protein PA3526 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG06914.1 |AE004773_3 probable outer membrane protein precursor [Pseudomonas aeruginosa PA01] Length = 321 J
5232.1 Best-BiastP=> >nrprot 39% Identities = 94/302 (31 %), Positives = 135/302 (44%), Gaps = 38/302 (12%) ref|ZP_00065012.11 COG0323: DNA mismatch repair enzyme (predicted ATPase) [Microbulbifer degradans 2-40] Length = 630
5238.1 Best-BlastP=> >nrprot 71 % Identities = 266/495 (53%), Positives = 354/495 (71 %), Gaps = 5/495 (1 %) ref|ZP_00067387.11 COG0138: AICAR transformylase/IMP cyclohydrolase PurH (only IMP cyclohydrolase domain in Aful) [Microbulbifer degradans 2-40] Length = 526
524.2 Best-BlastP=> >nrprot 69% Identities = 194/342 (56%), Positives = 241/342 (70%), Gaps = 2/342 (0%) ref|NP_820684.1 | dihydroorotase, homodimeric type [Coxiella burnetii RSA 493] gb|AA091 198.11 dihydroorotase, homodimeric type [Coxiella burnetii RSA 493] Length = 351
5242.3 Best-BlastP=> >nrprot No Hits found
5243.2 Best-BlastP=> >nrprot No Hits found
5247.1 Best-BlastP=> >nrprot 40% Identities = 80/312 (25%), Positives = 141/312 (45%), Gaps = 26/312 (8%) ref|ZP_00110262.1 | hypothetical protein [Nostoc punctiforme] Length = 348
5250.2 Best-BlastP=> >nrprot 86% Identities = 234/324 (72%), Positives = 280/324 (86%) ref|NP_819669.11 dehydrogenase, E1 component, beta subunit, putative [Coxiella burnetii RSA 493] gb|AAO90183.1 | dehydrogenase, E1 component, beta subunit, putative [Coxiella burnetii RSA 493] Length = 326
5253.1 Best-BlastP=> >nrprot 62% Identities = 92/196 (46%), Positives = 132/196 (67%), Gaps = 3/196 (1 %) ref|ZP_00090036.1 | COG4445: Hydroxylase for synthesis of 2-methylthio-cis-ribozeatin in tRNA [Azotobacter vinelandii] Length = 200
5254.1 Best-BlastP=> >nrprot 53% Identities = 72/206 (34%), Positives = 121/206 (58%) ref|NP_820242.11 conserved hypothetical protein [Coxiella burnetii RSA 493] gb|AAO90756.11 conserved hypothetical protein [Coxiella burnetii RSA 493] Length = 206
5255.1 Best-BlastP=> >nrprot 54% Identities = 132/378 (34%), Positives = 210/378 (55%), Gaps = 8/378 (2%) ref|ZP_00087763.1 | COG1520: FOG: WD40-like repeat [Pseudomonas fluorescens PfO-1] Length = 440
5256.2 Best-BlastP=> >nrprot 98% Identities = 64/64 (100%), Positives = 64/64 (100%) gb|AAG40471.11 global regulator [Legionella pneumophila] Length = 64
526.2 Best-BlastP=> >nrprot 97% Identities = 202/207 (97%), Positives = 203/207 (98%) gb|AAM00600.11 Rnase T [Legionella pneumophila] Length = 207
5266.2 Best-BlastP=> >nrprot 39% Identities = 212/1088 (19%), Positives = 439/1088 (40%), Gaps = 185/1088 (17%) pir||T14867 interaptin - slime mold (Dictyostelium discoideum) gb|AAC34582.1 | interaptin [Dictyostelium discoideum] Length = 1738
5268.2 Best-BlastP=> >nrprot 54% Identities = 56/183 (30%), Positives = 107/183 (58%), Gaps = 6/183 (3%) ref|NP_821059.11 conserved hypothetical protein [Coxiella burnetii RSA 493] gb|AA091573.11 conserved hypothetical protein [Coxiella burnetii RSA 493] Length = 192
5269.1 Best-BlastP=> >nrprot No Hits found
5270.2 Best-BlastP=> >nrprot No Hits found 5273.1 Best-BlastP=> >nrprot 55% Identities = 42/101 (41 %), Positives = 61/101 (60%), Gaps = 3/101 (2%) ref |NP_820129.11 oligopeptide transporter, OPT family [Coxiella burnetii RSA 493] gb|AAO90643.11 oligopeptide transporter, OPT family [Coxiella burnetii RSA 493] Length = 669
5277.3 Best-BlastP=> >nrprot 60% Identities = 186/328 (56%), Positives = 235/328 (71 %), Gaps = 5/328 (1 %) ref |NP_901482.11 dihydroorotate oxidase [Chromobacterium violaceum ATCC 12472] gb|AAQ59486.11 dihydroorotate oxidase [Chromobacterium violaceum ATCC 12472] Length = 344 5278.3 Best-BlastP=> >nrprot No Hits found 528.2 Best-BlastP=> >nrprot 99% Identities = 200/201 (99%), Positives = 201/201 (100%) gb|AAM00601.11 peroxynitrite reductase [Legionella pneumophila] Length = 201
5282.3 Best-BlastP=> >nrprot 44% Identities = 20/54 (37%), Positives = 32/54 (59%) ref|ZP_00077653.11 COG0693: Putative intracellular protease/amidase [Methanosarcina barkeri] Length = 209
5288.1 Best-BlastP=> >nrprot 60% Identities = 60/130 (46%), Positives = 83/130 (63%), Gaps = 4/130 (3%) pir||A60635 glutathione transferase (EC 2.5.1.18), fosfomycin-modifying - Escherichia coli plasmid pSU961 transposon Tn2921 gb|AAA98399.11 fosfomycin-resistance protein [Serratia marcescens] Length = 141
5289.2 Best-BlastP=> >nrprot 32% Identities = 49/169 (28%), Positives = 73/169 (43%), Gaps = 19/169 (11 %) ref|NP_819837.11 aminoglycoside N(6')- acetyltransferase [Coxiella burnetii RSA 493] gb|AAO90351.11 aminoglycoside N(6')-acetyltransferase [Coxiella burnetii RSA 493] Length = 188
529.2 Best-BlastP=> >nrprot 99% Identities = 105/105 (100%), Positives = 105/105 (100%) gb|AAM00602.11 glutaredoxin-like protein [Legionella pneumophila] Length = 115
5295.1 Best-BlastP=> >nrprot 68% Identities = 218/435 (50%), Positives = 309/435 (71 %) ref|NP_253162.11 PmbA protein [Pseudomonas aeruginosa PA01] pir||B83086 PmbA protein PA4472 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG07860.1 |AE004861_1 PmbA protein [Pseudomonas aeruginosa PA01] Length = 449
5297.2 Best-BlastP=> >nrprot No Hits found
530.3 Best-BlastP=> >nrprot 50% Identities = 75/162 (46%), Positives = 100/162 (61 %), Gaps = 3/162 (1%) ref|ZP_00129339.11 hypothetical protein [Desulfovibrio desulfuricans G20] Length = 231
5307.3 Best-BlastP=> >nrprot 19% Identities = 64/234 (27%), Positives = 111/234 (47%), Gaps = 23/234 (9%) ref|NP_820063.11 hypothetical protein [Coxiella burnetii RSA 493] gb|AAO90577.11 hypothetical protein [Coxiella burnetii RSA 493] Length = 468
5309.1 Best-BlastP=> >nrprot 84% Identities = 196/267 (73%), Positives = 234/267 (87%) ref|NP_743889.11 septum site-determining protein MinD [Pseudomonas putida KT2440] gb|AAN67353.1 |AE016361_7 septum site-determining protein MinD [Pseudomonas putida KT2440] Length = 270
531.4 Best-BlastP=> >nrprot No Hits found
5311.1 Best-BlastP=> >nrprot 58% Identities = 43/94 (45%), Positives = 58/94 (61 %) ref|ZP_00090060.11 COG0721 : Asp-tRNAAsn/Glu-tRNAGIn amidotransferase C subunit [Azotobacter vinelandii] Length = 146
5312.2 Best-BlastP=> >nrprot 99% Identities = 206/208 (99%), Positives = 207/208 (99%) gb|AAF05324.2| unknown virulence protein [Legionella pneumophila] Length = 208
5313.1 Best-BlastP=> >nrprot 32% Identities = 28/91 (30%), Positives = 46/91 (50%), Gaps = 6/91 (6%) ref|NP_520202.11 PROBABLE TRANSMEMBRANE PROTEIN [Ralstonia solanacearum] emb|CAD15788.11 PROBABLE TRANSMEMBRANE PROTEIN [Ralstonia solanacearum] Length = 176
5316.1 Best-BlastP=> >nrprot 41% Identities = 200/203 (98%), Positives = 203/203 (100%) gb|AAF05325.11 unknown virulence protein [Legionella pneumophila] Length = 205
5317.2 Best-BlastP=> >nrprot 65% Identities = 94/204 (46%), Positives = 136/204 (66%), Gaps = 1/204 (0%) ref|ZP_00067804.11 hypothetical protein [Microbulbifer degradans 2-40] Length = 206
5318.2 Best-BlastP=> >nrprot 34% Identities = 47/115 (40%), Positives = 72/115 (62%) ref|NP_790719.11 conserved hypothetical protein [Pseudomonas syringae pv. tomato str. DC3000] gb|AA054414.1 | conserved hypothetical protein [Pseudomonas syringae pv. tomato str. DC3000] Length = 249 5319.1 Best-BlastP=> >nrprot No Hits found 532.2 Best-BlastP=> >nrprot 57% Identities = 132/383 (34%), Positives = 215/383 (56%), Gaps = 24/383 (6%) ref|NP_718711.11 conserved hypothetical protein [Shewanella oneidensis MR-1] gb|AAN56155.1 |AE015753_1 conserved hypothetical protein [Shewanella oneidensis MR-1] Length = 374 5321 .1 Best-BlastP=> >nrprot 43% Identities = 27/53 (50%), Positives = 38/53 (71 %), Gaps = 2/53 (3%) gb|AAP84173.11 conserved hypothetical protein [Pseudomonas aeruginosa] Length = 744
5322.1 Best-BlastP=> >nrprot 58% Identities = 36/102 (35%), Positives = 61/102 (59%), Gaps = 3/102 (2%) ref |NP_816469.11 conserved domain protein [Enterococcus faecalis V583] gb|AA082539.1 1 conserved domain protein [Enterococcus faecalis V583] Length = 104
5324.1 Best-BlastP=> >nrprot 51 % Identities = 72/194 (37%), Positives = 113/194 (58%), Gaps = 2/194 (1 %) ref |NP_719033.11 AcrB/AcrD/AcrF family protein [Shewanella oneidensis MR-1] gb|AAN56477.1 |AE015784_10 AcrB/AcrD/AcrF family protein [Shewanella oneidensis MR-1] Length = 1046
5328.1 Best-BlastP=> >nrprot 41 % Identities = 34/123 (27%), Positives = 62/123 (50%), Gaps = 4/123 (3%) ref|NP_692567.11 transposase for IS652 [Oceanobacillus iheyensis HTE831] dbj|BAC13602.11 transposase for IS652 [Oceanobacillus iheyensis HTE831] Length = 402
533.2 Best-BlastP=> >nrprot 64% Identities = 122/264 (46%), Positives = 172/264 (65%), Gaps = 8/264 (3%) ref |NP_246208.11 AroE [Pasteurella multocida] sp|P57932|AROE_PASMU Shikimate 5-dehydrogenase gb|AAK03355.1 | AroE [Pasteurella multocida] Length = 269
5334.1 Best-BlastP=> >nrprot 32% Identities = 70/121 (57%), Positives = 83/121 (68%), Gaps = 9/121 (7%) gb|AAQ82687.11 Epaδp [Candida glabrata] Length = 1218
5337.1 Best-BlastP=> >nrprot 69% Identities = 49/91 (53%), Positives = 68/91 (74%) gb|AAP83334.1 |AF469614_2 unknown [Francisella tularensis subsp. tularensis] Length = 94
5338.2 Best-BlastP=> >nrprot 30% Identities = 78/169 (46%), Positives = 111/169 (65%), Gaps = 2/169 (1%) gb|AAN34371.11 ORF1 transposase [Acinetobacter baumannii] Length = 180
5340.1 Best-BlastP=> >nrprot 14% Identities = 40/136 (29%), Positives = 73/136 (53%), Gaps = 19/136 (13%) gb|AAA67447.11 P120 Length = 232
5341.1 Best-BlastP=> >nrprot No Hits found
5342.2 Best-BlastP=> >nrprot 60% Identities = 87/178 (48%), Positives = 113/178 (63%), Gaps = 5/178 (2%) ref|NP_819838.11 transcriptional regulator, TetR family [Coxiella burnetii RSA 493] gb|AAO90352.11 transcriptional regulator, TetR family [Coxiella burnetii RSA 493] Length = 193
5344.2 Best-BlastP=> >nrprot 21 % Identities = 36/77 (46%), Positives = 48/77 (62%), Gaps = 1/77 (1 %) ref|ZP_00028865.11 hypothetical protein [Burkholderia fungorum] Length = 123
5349.3 Best-BlastP=> >nrprot 15% Identities = 65/306 (21 %), Positives = 140/306 (45%), Gaps = 40/306 (13%) dbj|BAC86266.11 unnamed protein product [Homo sapiens] Length = 486
5354.3 Best-BlastP=> >nrprot No Hits found
5359.2 Best-BlastP=> >nrprot 43% Identities = 22/59 (37%), Positives = 36/59 (61 %) ref|NP_637282.11 flagellar protein [Xanthomonas campestris pv. campestris str. ATCC 33913] gb|AAM41206.1 | flagellar protein [Xanthomonas campestris pv. campestris str. ATCC 33913] Length = 135
536.3 Best-BlastP=> >nrprot 69% Identities = 459/876 (52%), Positives = 600/876 (68%), Gaps = 23/876 (2%) ref|NP_744167.11 aminopeptidase N [Pseudomonas putida KT2440] gb|AAN67631.1 |AE016392_12 aminopeptidase N [Pseudomonas putida KT2440] Length = 885
5360.2 Best-BlastP=> >nrprot No Hits found
5361.2 Best-BlastP=> >nrprot 68% Identities = 179/380 (47%), Positives = 261/380 (68%), Gaps = 3/380 (0%) ref|NP_746466.11 flagellar biosynthetic protein FlhB [Pseudomonas putida KT2440] gb|AAN69930.1 |AE016632_1 flagellar biosynthetic protein FlhB [Pseudomonas putida KT2440] Length = 380
5369.2 Best-BlastP=> >nrprot 72% Identities = 47/68 (69%), Positives = 58/68 (85%) ref |NP_639162.11 30S ribosomal protein S21 [Xanthomonas campestris pv. campestris str. ATCC 33913] ref |NP_644178.11 30S ribosomal protein S21 [Xanthomonas axonopodis pv. citri str. 306] sp|Q8NL04|RS21_XANAC 30S ribosomal protein S21 gb|AAM38714.11 30S ribosomal protein S21 [Xanthomonas axonopodis pv. citri str. 306] gb|AAM43491.11 30S ribosomal protein S21 [Xanthomonas campestris pv. campestris str. ATCC 33913] Length = 71
537.2 Best-BlastP=> >nrprot No Hits found 5370.1 Best-BlastP=> >nrprot 97% Identities = 142/147 (96%), Positives = 144/147 (97%), Gaps = 1/147 (0%) gb|AAB09541.1 | LporfX Length = 146
5373.1 Best-BlastP=> >nrprot No Hits found
5376.2 Best-BlastP=> >nrprot 25% Identities = 185/917 (20%), Positives = 352/917 (38%), Gaps = 160/917 (17%) gb|EAA15312.11 hypothetical protein [Plasmodium yoelii yoelii] Length = 1527
5377.2 Best-BlastP=> >nrprot No Hits found 538.1 Best-BlastP=> >nrprot No Hits found
5380.3 Best-BlastP=> >nrprot 49% Identities = 52/160 (32%), Positives = 91/160 (56%), Gaps = 5/160 (3%) ref|NP_248967.11 hypothetical protein [Pseudomonas aeruginosa PA01] pir||D83612 hypothetical protein PA0276 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG03665.1 |AE004465_11 hypothetical protein PA0276 [Pseudomonas aeruginosa PA01] Length = 171
5381.1 Best-BlastP=> >nrprot No Hits found
5382.1 Best-BlastP=> >nrprot No Hits found
5386.1 Best-BlastP=> >nrprot 67% Identities = 170/327 (51 %), Positives = 225/327 (68%), Gaps = 3/327 (0%) ref|NP_844883.11 oxidoreductase, NAD- binding [Bacillus anthracis str. Ames] gb|AAP26369.1 | oxidoreductase, NAD-binding [Bacillus anthracis str. Ames] Length = 341
5387.1 Best-BlastP=> >nrprot 67% Identities = 128/249 (51 %), Positives = 169/249 (67%), Gaps = 2/249 (0%) gb|AAM51645.11 putative transposase [Francisella tularensis subsp. tularensis] Length = 247
5388.1 Best-BlastP=> >nrprot No Hits found
539.3 Best-BlastP=> >nrprot 61% Identities = 107/144 (74%), Positives = 123/144 (85%) emb|CAB46580.11 IS1400 transposase B [Yersinia enterocolitica] Length = 294
5390.1 Best-BlastP=> >nrprot 41 % Identities = 30/89 (33%), Positives = 51/89 (57%) ref|NP_716406.1 1 conserved hypothetical protein [Shewanella oneidensis MR-1] gb|AAN53851 .1 |AE015522_6 conserved hypothetical protein [Shewanella oneidensis MR-1 ] Length = 100
5391.2 Best-BlastP=> >nrprot 58% Identities = 192/471 (40%), Positives = 283/471 (60%), Gaps = 5/471 (1 %) ref|NP_840383.11 conserved hypothetical protein [Nitrosomonas europaea ATCC 19718] emb|CAD84207.1 | conserved hypothetical protein [Nitrosomonas europaea ATCC 19718] Length = 510
5398.3 Best-BlastP=> >nrprot 66% Identities = 173/351 (49%), Positives = 237/351 (67%) ref|ZP_00084264.11 COG4972: Tfp pilus assembly protein, ATPase PilM [Pseudomonas fluorescens PfO-1] Length = 354
540.3 Best-BlastP=> >nrprot 72% Identities = 58/101 (57%), Positives = 78/101 (77%) emb|CAB46580.11 IS1400 transposase B [Yersinia enterocolitica] Length = 294
5402.1 Best-BlastP=> >nrprot No Hits found
5404.2 Best-BlastP=> >nrprot No Hits found 5405.2 Best-BlastP=> >nrprot 98% Identities = 314/322 (97%), Positives = 319/322 (99%) gb|AAM00613.11 chemiosmotic efflux system protein B-like protein [Legionella pneumophila] Length = 322
5406.1 Best-BlastP=> >nrprot 34% Identities = 109/300 (36%), Positives = 175/300 (58%), Gaps = 15/300 (5%) ref|NP_902623.11 conserved hypothetical protein [Chromobacterium violaceum ATCC 12472] gb|AAQ60621.1 | conserved hypothetical protein [Chromobacterium violaceum ATCC 12472] Length = 1390
5412.1 Best-BlastP=> >nrprot 72% Identities = 190/322 (59%), Positives = 232/322 (72%), Gaps = 6/322 (1 %) ref|NP_638907.11 conserved hypothetical protein [Xanthomonas campestris pv. campestris str. ATCC 33913] gb|AAM42831.1 | conserved hypothetical protein [Xanthomonas campestris pv. campestris str. ATCC 33913] Length = 323
5413.1 Best-BlastP=> >nrprot No Hits found 5417.1 Best-BlastP=> >nrprot 59% Identities = 50/137 (36%), Positives = 82/137 (59%), Gaps = 9/137 (6%) ref|NP_487806.11 two-component response regulator [Nostoc sp. PCC 7120] pir||AG2276 two-component response regulator all3766 [imported] - Nostoc sp. (strain PCC 7120) dbj|BAB75465.11 two-component response regulator [Nostoc sp. PCC 7120] Length = 143
5420.1 Best-BlastP=> >nrprot 49% Identities = 42/134 (31 %), Positives = 67/134 (50%), Gaps = 20/134 (14%) gb|AAH41716.11 Similar to myosin, heavy polypeptide 4, skeletal muscle [Xenopus laevis] Length = 1170
5421.1 Best-BlastP=> >nrprot No Hits found
5422.1 Best-BlastP=> >nrprot No Hits found
5423.1 Best-BlastP=> >nrprot 64% Identities = 36/75 (48%), Positives = 50/75 (66%) ref|NP_252731.11 exodeoxyribonuclease VII small subunit [Pseudomonas aeruginosa PA01] ref|ZP_00137487.11 COG1722: Exonuclease VII small subunit [Pseudomonas aeruginosa UCBPP- PA14] sp|Q9HWY5|EX7S_PSEAE Probable exodeoxyribonuclease VII small subunit (Exonuclease VII small subunit) pir||E83139 exodeoxyribonuclease VII small subunit PA4042 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG07429.1 |AE004821_2 exodeoxyribonuclease VII small subunit [Pseudomonas aeruginosa PA01] Length = 80
5424.2 Best-BlastP=> >nrprot No Hits found
5426.2 Best-BlastP=> >nrprot 71 % Identities = 86/142 (60%), Positives = 109/142 (76%), Gaps = 2/142 (1 %) ref|NP_820558.1 | tolR protein [Coxiella burnetii RSA 493] gb|AAO91072.11 tolR protein [Coxiella burnetii RSA 493] Length = 147
5427.1 Best-BlastP=> >nrprot 68% Identities = 1 14/225 (50%), Positives = 154/225 (68%), Gaps = 4/225 (1 %) ref|ZP_00082839.11 COG081 1 : Biopolymer transport proteins [Pseudomonas fluorescens PfO-1] Length = 231
5428.1 Best-BlastP=> >nrprot 53% Identities = 48/116 (41 %), Positives = 70/116 (60%), Gaps = 1/116 (0%) ref|NP_404733.1 | conserved hypothetical protein [Yersinia pestis] ref|NP_670358.1 | hypothetical protein [Yersinia pestis KIM] pir||AH0137 conserved hypothetical protein YPO1120 [imported] - Yersinia pestis (strain C092) emb|CAC89963.1 1 conserved hypothetical protein [Yersinia pestis C092] gb|AAM86609.1 |AE013907_3 hypothetical protein [Yersinia pestis KIM] Length = 133
5429.1 Best-BlastP=> >nrprot 56% Identities = 33/73 (45%), Positives = 45/73 (61 %), Gaps = 2/73 (2%) ref|NP_819889.11 conserved hypothetical protein [Coxiella burnetii RSA 493] gb|AAO90403.11 conserved hypothetical protein [Coxiella burnetii RSA 493] Length = 82
543.1 Best-BlastP=> >nrprot No Hits found
5437.2 Best-BlastP=> >nrprot No Hits found 5438.1 Best-BlastP=> >nrprot 73% Identities = 114/187 (60%), Positives = 144/187 (77%) ref|ZP_00065453.11 COG1207: N-acetylglucosamine-1- phosphate uridyltransferase (contains nucleotidyltransferase and l-patch acetyltransferase domains) [Microbulbifer degradans 2-40] Length = 451
544.2 Best-BlastP=> >nrprot 76% Identities = 265/446 (59%), Positives = 337/446 (75%), Gaps = 5/446 (1 %) ref|ZP_00138105.11 COG2204: Response regulator containing CheY-like receiver, AAA-type ATPase, and DNA-binding domains [Pseudomonas aeruginosa UCBPP- PA14] Length = 597
5440.1 Best-BlastP=> >nrprot No Hits found
5446.3 Best-BlastP=> >nrprot 62% Identities = 188/449 (41 %), Positives = 279/449 (62%), Gaps = 12/449 (2%) gb|AAP74578.11 kynurenine 3- monooxygenase [Polaribacter filamentus] Length = 469
5447.1 Best-BlastP=> >nrprot 51 % Identities = 88/224 (39%), Positives = 129/224 (57%), Gaps = 8/224 (3%) ref|NP_484370.11 unknown protein [Nostoc sp. PCC 7120] pir||AE1847 hypothetical protein all0326 [imported] - Nostoc sp. (strain PCC 7120) dbj|BAB72284.11 ORF_ID:all0326~unknown protein [Nostoc sp. PCC 7120] Length = 224
5448.1 Best-BlastP=> >nrprot 35% Identities = 36/120 (30%), Positives = 61/120 (50%), Gaps = 7/120 (5%) sp|P45790|GSPC_AERHY GENERAL SECRETION PATHWAY PROTEIN C emb|CAA47125.11 ExeC [Aeromonas hydrophila] Length = 290
5453.2 Best-BlastP=> >nrprot 68% Identities = 171/344 (49%), Positives = 242/344 (70%), Gaps = 1/344 (0%) ref|NP_819316.1 | major facilitator family transporter [Coxiella burnetii RSA 493] gb|AAO89830.11 major facilitator family transporter [Coxiella burnetii RSA 493] Length = 458
5455.1 Best-BlastP=> >nrprot 64% Identities = 71/152 (46%), Positives = 101/152 (66%) ref|ZP_00140766.11 COG5528: Predicted integral membrane protein [Pseudomonas aeruginosa UCBPP-PA14] Length = 155
5456.1 Best-BlastP=> >nrprot 99% Identities = 225/226 (99%), Positives = 226/226 (100%) gb|AAQ18123.11 CpxR [Legionella pneumophila] Length = 226 5457.1 Best-BlastP=> >nrprot 57% Identities = 52/117 (44%), Positives = 77/117 (65%) ref|NP_384234.11 PUTATIVE CYTIDINE DEAMINASE PROTEIN [Sinorhizobium meliloti] emb|CAC41515.11 PUTATIVE CYTIDINE DEAMINASE PROTEIN [Sinorhizobium meliloti] Length = 152
5458.1 Best-BlastP=> >nrprot 53% Identities = 32/96 (33%), Positives = 52/96 (54%), Gaps = 2/96 (2%) gb|AAN34371.11 ORF1 transposase [Acinetobacter baumannii] Length = 180
5460.1 Best-BlastP=> >nrprot 56% Identities = 147/325 (45%), Positives = 206/325 (63%), Gaps = 10/325 (3%) ref|NP_189431.2| DegP protease [Arabidopsis thaliana] gb|AAK62640.1 | K16N12.18/K16N12.18 [Arabidopsis thaliana] gb|AAM47381.1 | At3g27g25/K16N12.18 [Arabidopsis thaliana] Length = 439
5462.2 Best-BlastP=> >nrprot 66% Identities = 270/541 (49%), Positives = 367/541 (67%), Gaps = 6/541 (1 %) ref|NP_819168.11 penicillin-binding protein 3 [Coxiella burnetii RSA 493] gb|AA089682.11 penicillin-binding protein 3 [Coxiella burnetii RSA 493] Length = 548
5467.1 Best-BiastP=> >nrprot No Hits found
5469.1 Best-BlastP=> >nrprot 70% Identities = 24/55 (43%), Positives = 40/55 (72%) ref|NP_438830.11 hypothetical protein [Haemophilus influenzae Rd] sp|P44814|DTD_HAEIN D-tyrosyl-tRNA(Tyr) deacylase pir||E64156 hypothetical protein HI0670 - Haemophilus influenzae (strain Rd KW20) pdb|1J7G|A Chain A, Structure Of Yihz From Haemophilus Influenzae (Hi0670), A D- Tyr-Trna(Tyr) Deacylase gb|AAC22330.1 | conserved hypothetical protein [Haemophilus influenzae Rd] Length = 144
5470.1 Best-BlastP=> >nrprot 76% Identities = 58/83 (69%), Positives = 65/83 (78%) ref|ZP_00083364.11 COG1490: D-Tyr-tRNAtyr deacylase
[Pseudomonas fluorescens PfO-1] Length = 145
5471.2 Best-BlastP=> >nrprot 59% Identities = 30/57 (52%), Positives = 41/57 (71 %) ref|NP_773232.11 bsr6592 [Bradyrhizobium japonicum] dbj|BAC51857.1 | bsr6592 [Bradyrhizobium japonicum USDA 110] Length = 95
5473.1 Best-BlastP=> >nrprot 73% Identities = 154/270 (57%), Positives = 197/270 (72%), Gaps = 5/270 (1 %) ref|ZP_00029131.11 COG3243: Poly(3- hydroxyalkanoate) synthetase [Burkholderia fungorum] Length = 642
5474.2 Best-BlastP=> >nrprot 42% Identities = 44/134 (32%), Positives = 73/134 (54%), Gaps = 2/134 (1 %) ref |ZP_00027817.11 COG0454: Histone acetyltransferase HPA2 and related acetyltransferases [Burkholderia fungorum] Length = 174
5476.1 Best-BlastP=> >nrprot 54% Identities = 48/106 (45%), Positives = 67/106 (63%), Gaps = 1/106 (0%) ref|NP_759432.11 HesB family protein [Vibrio vulnificus CMCP6] gb|AAO08959.1 |AE016798_119 HesB family protein [Vibrio vulnificus CMCP6] Length = 107
5477.1 Best-BlastP=> >nrprot 57% Identities = 36/84 (42%), Positives = 55/84 (65%), Gaps = 1/84 (1 %) ref|NP_873022.11 DNA-binding protein [Haemophilus ducreyi 35000HP] gb|AAP95411.11 DNA-binding protein [Haemophilus ducreyi 35000HP] Length = 98
5478.2 Best-BlastP=> >nrprot 28% Identities = 30/102 (29%), Positives = 47/102 (46%), Gaps = 13/102 (12%) sp|Q9W751 |PIX1_XENLA Pituitary homeobox 1 (X-PITX-1 ) (xPitxl ) gb|AAD45292.1 |AF155206_1 homeodomain transcription factor Pitx-1 [Xenopus laevis] Length = 305
5479.2 Best-BlastP=> >nrprot 51% Identities = 62/169 (36%), Positives = 102/169 (60%), Gaps = 2/169 (1 %) gb|AAN46162.1 | unknown protein [Synechococcus sp. PCC 7942] Length = 208
548.3 Best-BlastP=> >nrprot No Hits found
5481.1 Best-BlastP=> >nrprot 50% Identities = 143/306 (46%), Positives = 192/306 (62%), Gaps = 1/306 (0%) ref|ZP_00043253.11 COG0790: FOG: TPR repeat, SEL1 subfamily [Magnetococcus sp. MC-1] Length = 831
5482.2 Best-BlastP=> >nrprot 41 % Identities = 75/284 (26%), Positives = 124/284 (43%), Gaps = 17/284 (5%) sp|O57809|1A1 D_PYRHO Putative 1- aminocyclopropane-1-carboxylate deaminase (ACC deaminase) pdb|1 J0A|A Chain A, Crystal Structure Analysis Of The Ace Deaminase Homologue pdb|1 J0A|B Chain B, Crystal Structure Analysis Of The Ace Deaminase Homologue pdb|1 J0A|C Chain C, Crystal Structure Analysis Of The Ace Deaminase Homologue pdb|1 J0B|A Chain A, Crystal Structure Analysis Of The Ace Deaminase Homologue Complexed With Inhiitor pdb|1J0B|B Chain B, Crystal Structure Analysis Of The Ace Deaminase Homologue Complexed With Inhiitor pdb|1J0B|C Chain C, Crystal Structure Analysis Of The Ace Deaminase Homologue Complexed With Inhiitor pdb|1 J0B|D Chain D, Crystal Structure Analysis Of The Ace Deaminase Homologue Complexed With Inhiitor pdb|1 J0B|E Chain E, Crystal Structure Analysis Of The Ace Deaminase Homologue Complexed With Inhiitor pdb|1 J0B|F Chain F, Crystal Structure Analysis Of The Ace Deaminase Homologue Complexed With Inhiitor pdb|1 J0B|G Chain G, Crystal Structure Analysis Of The Ace Deaminase Homologue Complexed With Inhiitor pdb|'
5484.4 Best-BlastP=> >nrprot No Hits found
5489.2 Best-BlastP=> >nrprot 52% Identities = 64/189 (33%), Positives = 107/189 (56%), Gaps = 8/189 (4%) ref|NP_882264.11 putative exported protein [Bordetella pertussis] ref|NP_886390.1 | putative exported protein [Bordetella parapertussis] ref|NP_891381.1 | putative exported protein [Bordetella bronchiseptica] emb|CAE44018.1 | putative exported protein [Bordetella pertussis] emb|CAE39540.1 | putative exported protein [Bordetella parapertussis] emb|CAE3521 1 .11 putative exported protein [Bordetella bronchiseptica] Length = 207
549.5 Best-BlastP=> >nrprot 27% Identities = 60/253 (23%), Positives = 108/253 (42%), Gaps = 12/253 (4%) ref|NP_638097.11 conserved hypothetical protein [Xanthomonas campestris pv. campestris str. ATCC 33913] gb|AAM42021.1 | conserved hypothetical protein [Xanthomonas campestris pv. campestris str. ATCC 33913] Length = 618
5492.2 Best-BlastP=> >nrprot 77% Identities = 105/178 (58%), Positives = 145/178 (81 %) ref|NP_251274.11 CDP-diacylglycerol-glycerol-3-phosphate 3 phosphatidyltransferase [Pseudomonas aeruginosa PA01 ] pir||B83322 CDP-diacylglycerol-glycerol-3-phosphate 3-phosphatidyltransferase PA2584 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG05972.1 |AE004687_1 CDP-diacylglycerol-glycerol-3-phosphate 3- phosphatidyltransferase [Pseudomonas aeruginosa PA01] Length = 186
5496.2 Best-BlastP=> >nrprot 95% Identities = 168/175 (96%), Positives = 171/175 (97%) emb|CAB65201.1 | hypothetical protein [Legionella pneumophila] Length = 356
5497.1 Best-BlastP=> >nrprot 98% Identities = 328/334 (98%), Positives = 330/334 (98%) emb|CAB65202.11 WecA protein [Legionella pneumophila] Length = 334 5498.1 Best-BlastP=> >nrprot 99% Identities = 317/318 (99%), Positives = 318/318 (100%) emb|CAB65203.1 | hypothetical protein [Legionella pneumophila] Length = 318
5499.1 Best-BlastP=> >nrprot 99% Identities = 291/291 (100%), Positives = 291/291 (100%) emb|CAB65204.11 RmlA protein [Legionella pneumophila] Length = 291 55.1 Best-BlastP=> >nrprot 98% Identities = 125/128 (97%), Positives = 127/128 (99%) gb|AAM08236.1 | LvrB [Legionella pneumophila] Length = 150 5500.1 Best-BlastP=> >nrprot 98% Identities = 484/494 (97%), Positives = 486/494 (98%), Gaps = 2/494 (0%) sp|Q9RDY2|G6PI_LEGPN Glucose-6- phosphate isomerase (GPI) (Phosphoglucose isomerase) (PGI) (Phosphohexose isomerase) (PHI) emb|CAB65205.1 | Gpi protein [Legionella pneumophila] Length = 497
5504.4 Best-BlastP=> >nrprot No Hits found 551.2 Best-BlastP=> >nrprot 64% Identities = 122/243 (50%), Positives = 163/243 (67%), Gaps = 3/243 (1 %) ref|ZP_00039313.1 | COG1028: Dehydrogenases with different specificities (related to short-chain alcohol dehydrogenases) [Xylella fastidiosa Dixon] Length = 245 5514.2 Best-BlastP=> >nrprot 32% Identities = 56/241 (23%), Positives = 110/241 (45%), Gaps = 36/241 (14%) gb|EAA16038.11 repeat organellar protein-related [Plasmodium yoelii yoelii] Length = 1441
5515.2 Best-BlastP=> >nrprot 78% Identities = 149/249 (59%), Positives = 197/249 (79%) ref|NP_251642.11 electron transfer flavoprotein beta-subunit [Pseudomonas aeruginosa PA01] pir||C83277 electron transfer flavoprotein beta-subunit PA2952 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG06340.1 |AE004721_8 electron transfer flavoprotein beta-subunit [Pseudomonas aeruginosa PA01] Length = 249
5517.1 Best-BlastP=> >nrprot 50%. Identities = 47/123 (38%), Positives = 65/123 (52%), Gaps = 14/123 (11 %) ref|ZP_00087881.1 | COG0357: Predicted S-adenosylmethionine-dependent methyltransferase involved in bacterial cell division [Pseudomonas fluorescens PfO-1] Length = 138
5520.1 Best-BlastP=> >nrprot 38% Identities = 31/125 (24%), Positives = 57/125 (45%), Gaps = 4/125 (3%) ref|NP_716604.11 hypothetical protein [Shewanella oneidensis MR-1] gb|AAN54049.1 |AE015542_5 hypothetical protein [Shewanella oneidensis MR-1] Length = 474
5521.1 Best-BlastP=> >nrprot 61 % Identities = 47/133 (35%), Positives = 83/133 (62%), Gaps = 1/133 (0%) ref|NP_903590.11 conserved hypothetical protein [Chromobacterium violaceum ATCC 12472] gb|AAQ61581.1 | conserved hypothetical protein [Chromobacterium violaceum ATCC 12472] Length = 144
5523.2 Best-BlastP=> >nrprot 60% Identities = 157/327 (48%), Positives = 203/327 (62%), Gaps = 9/327 (2%) ref|ZP_00086640.11 COG1612: Uncharacterized protein required for cytochrome oxidase assembly [Pseudomonas fluorescens PfO-1] Length = 359
5524.1 Best-BlastP=> >nrprot 43% Identities = 46/180 (25%), Positives = 76/180 (42%), Gaps = 25/180 (13%) ref|NP_800052.11 hypothetical protein VPA0542 [Vibrio parahaemolyticus RIMD 2210633] dbj|BAC61885.11 hypothetical protein [Vibrio parahaemolyticus] Length = 178
5526.2 Best-BlastP=> >nrprot 30% Identities = 42/141 (29%), Positives = 64/141 (45%), Gaps = 2/141 (1 %) ref|ZP_00081004.11 COG3637: Opacity protein and related surface antigens [Geobacter metallireducens] Length = 219
5527.2 Best-BlastP=> >nrprot 69% Identities = 91/170 (53%), Positives = 120/170 (70%) ref|NP_743494.11 UDP-N-acetylmuramoylalanine-D-glutamate ligase [Pseudomonas putida KT2440] gb|AAN66958.1 |AE016324_8 UDP-N-acetylmuramoylalanine-D-glutamate ligase [Pseudomonas putida KT2440] Length = 450
5528.2 Best-BlastP=> >nrprot 99% Identities = 239/239 (100%), Positives = 239/239 (100%) emb|CAB65196.11 hypothetical protein [Legionella pneumophila] Length = 239
553.1 Best-BlastP=> >nrprot No Hits found 5530.2 Best-BlastP=> >nrprot 68% Identities = 119/231 (51 %), Positives = 161/231 (69%) ref|NP_767647.1 | bll1007 [Bradyrhizobium japonicum] dbj|BAC46272.11 bll1007 [Bradyrhizobium japonicum USDA 110] Length = 345
5532.2 Best-BlastP=> >nrprot 50% Identities = 86/270 (31 %), Positives = 146/270 (54%), Gaps = 11 /270 (4%) ref|ZP_00117263.11 COG3781 : Predicted membrane protein [Cytophaga hutchinsonii] Length = 290
5533.2 Best-BlastP=> >nrprot 43% Identities = 28/84 (33%), Positives = 35/84 (41 %), Gaps = 16/84 (19%) gb|AAO52009.11 similar to exonuclease ii [Schizosaccharomyces pombe] [Dictyostelium discoideum] Length = 1749
5534.2 Best-BlastP=> >nrprot 98% Identities = 486/494 (98%), Positives = 489/494 (98%) gb|AAK35046.1 |AF330136_2 type II protein secretion ATPase LspE [Legionella pneumophila] Length = 494
5535.1 Best-BlastP=> >nrprot No Hits found
5538.2 Best-BlastP=> >nrprot 75% Identities = 95/159 (59%), Positives = 121/159 (76%), Gaps = 1/159 (0%) ref|NP_819315.11 single-strand binding protein [Coxiella burnetii RSA 493] gb|AA089829.11 single-strand binding protein [Coxiella burnetii RSA 4 3] Length = 158
5539.1 Best-BlastP=> >nrprot No Hits found
5540.1 Best-BlastP=> >nrprot 81 % Identities = 90/126 (71 %), Positives = 104/126 (82%) ref|NP_252927.11 50S ribosomal protein L17 [Pseudomonas aeruginosa PA01] sp|052761 |RL17_PSEAE 50S ribosomal protein L17 pir||C83113 50S ribosomal protein L17 PA4237 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAC03117.11 ribosomal large subunit protein L17 [Pseudomonas aeruginosa] gb|AAG07625.1 |AE004841_3 50S ribosomal protein L17 [Pseudomonas aeruginosa PA01] Length = 129
5542.3 Best-BlastP=> >nrprot 70% Identities = 120/255 (47%), Positives = 183/255 (71%), Gaps = 1/255 (0%) ref|NP_831729.11 Aminoglycoside 6- adenylyltransferase [Bacillus cereus ATCC 14579] gb|AAP08930.11 Aminoglycoside 6-adenylyltransferase [Bacillus cereus ATCC 14579] Length = 290
5546.1 Best-BlastP=> >nrprot 73% Identities = 65/107 (60%), Positives = 79/107 (73%), Gaps = 1/107 (0%) ref|NP_820379.11 ErfK/YbiS/YcfS/YnhG family protein [Coxiella burnetii RSA 493] gb|AAO90893.11 ErfK/YbiS/YcfS/YnhG family protein [Coxiella burnetii RSA 493] Length = 175
5548.1 Best-BlastP=> >nrprot No Hits found
5549.2 Best-BlastP=> >nrprot 73% Identities = 134/229 (58%), Positives = 172/229 (75%), Gaps = 3/229 (1 %) ref|ZP_00065236.11 COG0847: DNA polymerase III, epsilon subunit and related 3'-5' exonucleases [Microbulbifer degradans 2-40] Length = 238
555.2 Best-BlastP=> >nrprot 53% Identities = 110/317 (34%), Positives = 171/317 (53%), Gaps = 11/317 (3%) ref|NP_442295.11 hypothetical protein [Synechocystis sp. PCC 6803] sp|Q55724|Y644_SYNY3 Hypothetical protein slr0644 pir||S76519 hypothetical protein - Synechocystis sp. (strain PCC 6803) dbj|BAA10365.11 ORF_ID:slr0644~hypothetical protein [Synechocystis sp. PCC 6803] Length = 355
5550.3 Best-BlastP=> >nrprot 80% Identities = 334/503 (66%), Positives = 412/503 (81 %), Gaps = 2/503 (0%) ref|NP_819973.11 cytochrome d ubiquinol oxidase, subunit I [Coxiella burnetii RSA 493] gb|AAO90487.1 | cytochrome d ubiquinol oxidase, subunit I [Coxiella burnetii RSA 493] Length = 521 >
5552.2 Best-BlastP=> >nrprot 75% Identities = 232/378 (61 %), Positives = 287/378 (75%), Gaps = 2/378 (0%) gb|AAG01153.1 |AF284438_4 cytochrome d oxidase subunit [Brucella melitensis biovar Abortus] Length = 384
5554.2 Best-BlastP=> >nrprot No Hits found 5555.2 Best-BlastP=> >nrprot No Hits found
5556.1 Best-BlastP=> >nrprot No Hits found
5557.2 Best-BlastP=> >nrprot 42% Identities = 113/561 (20%), Positives = 251/561 (44%), Gaps = 65/561 (11%) dbj|BAB40921.2| myosin heavy chain 2x [Bos taurus] Length = 1938
5559.3 Best-BlastP=> >nrprot No Hits found 5560.3 Best-BlastP=> >nrprot 54% Identities = 68/232 (29%), Positives = 116/232 (50%), Gaps = 21/232 (9%) ref]NP_923239.11 probable carbonyl reductase [Gloeobacter violaceus] dbj|BAC88234.11 glr0293 [Gloeobacter violaceus] Length = 243
5563.1 Best-BlastP=> >nrprot 54% Identities = 57/145 (39%), Positives = 84/145 (57%), Gaps = 3/145 (2%) ref|NP_841440.11 GCN5-related N- acetyltransferase [Nitrosomonas europaea ATCC 19718] emb|CAD85310.11 GCN5-related N-acetyltransferase [Nitrosomonas europaea ATCC 19718] Length = 157
5564.1 Best-BlastP=> >nrprot 81 % Identities = 322/475 (67%), Positives = 387/475 (81 %), Gaps = 2/475 (0%) ref|NP_819499.11 dihydrolipoamide dehydrogenase [Coxiella burnetii RSA 493] gb|AAO90013.11 dihydrolipoamide dehydrogenase [Coxiella burnetii RSA 493] Length = 474
5566.1 Best-BlastP=> >nrprot 75% Identities = 38/55 (69%), Positives = 45/55 (81 %) ref|NP_523225.11 PROBABLE TRANSMEMBRANE PROTEIN [Ralstonia solanacearum] emb|CAD18817.11 PROBABLE TRANSMEMBRANE PROTEIN [Ralstonia solanacearum] Length = 72
5569.1 Best-BlastP=> >nrprot No Hits found 557.2 Best-BlastP=> >nrprot 75% Identities = 517/898 (57%), Positives = 675/898 (75%), Gaps = 15/898 (1 %) ref|NP_820774.11 DNA polymerase I [Coxiella burnetii RSA 493] gb|AA091288.11 DNA polymerase I [Coxiella burnetii RSA 493] Length = 895
5576.3 Best-BlastP=> >nrprot 26% Identities = 61/216 (28%), Positives = 87/216 (40%), Gaps = 25/216 (11 %) ref|NP_662329.1 | hypothetical protein [Chlorobium tepidum TLS] gb|AAM72671.11 hypothetical protein [Chlorobium tepidum TLS] Length = 325
558.2 Best-BlastP=> >nrprot 34% Identities = 31/95 (32%), Positives = 53/95 (55%), Gaps = 9/95 (9%) gb|AAM15532.1 |AF482691_1 probable sensor/response regulator hybrid [Pseudomonas aeruginosa] Length = 469
5582.4 Best-BlastP=> >nrprot 88% Identities = 584/737 (79%), Positives = 653/737 (88%), Gaps = 3/737 (0%) gb|AAM00624.11 putative copper efflux ATPase [Legionella pneumophila] Length = 736
5584.2 Best-BlastP=> >nrprot 52% Identities = 188/189 (99%), Positives = 188/189 (99%) emb|CAC33489.11 hypothetical protein [Legionella pneumophila] Length = 189
5586.1 Best-BlastP=> >nrprot 37% Identities = 32/76 (42%), Positives = 48/76 (63%) ref|NP_217688.11 hypothetical protein Rv3172c [Mycobacterium tuberculosis H37Rv] ref|NP_337786.1 | hypothetical protein [Mycobacterium tuberculosis CDC1551 ] ref|NP_856842.1 | HYPOTHETICAL PROTEIN [Mycobacterium bovis subsp. bovis AF2122/97] pir||B70948 hypothetical protein Rv3172c - Mycobacterium tuberculosis (strain H37RV) emb|CAA16637.11 hypothetical protein Rv3172c [Mycobacterium tuberculosis H37Rv] gb|AAK47600.1 | hypothetical protein [Mycobacterium tuberculosis CDC1551] emb|CAD95289.1 | HYPOTHETICAL PROTEIN [Mycobacterium bovis subsp. bovis AF2122/97] Length = 160
5587.3 Best-BlastP=> >nrprot 61 % Identities = 132/276 (47%), Positives = 173/276 (62%) ref|NP_229753.11 4-hydroxybenzoate octaprenyltransferase [Vibrio cholerae 01 biovar eltor str. N16961 ] pir||C82365 4-hydroxybenzoate octaprenyltransferase VC0094 [imported] - Vibrio cholerae (strain N16961 serogroup 01 ) gb|AAF93272.11 4-hydroxybenzoate octaprenyltransferase [Vibrio cholerae 01 biovar eltor str. N16961] Length = 284
5588.2 Best-BlastP=> >nrprot 67% Identities = 33/98 (33%), Positives = 65/98 (66%), Gaps = 4/98 (4%) ref|NP_819561 .11 hypothetical protein [Coxiella burnetii RSA 493] gb|AAO90075.1 1 hypothetical protein [Coxiella burnetii RSA 493] Length = 96
559.2 Best-BlastP=> >nrprot 83% Identities = 78/107 (72%), Positives = 92/107 (85%) ref|NP_862496.11 hypothetical protein [Pseudomonas sp. ADP] gb|AAK50291.1 |U66917_59 hypothetical protein [Pseudomonas sp. ADP] Length = 152
5592.2 Best-BlastP=> >nrprot 81 % Identities = 150/229 (65%), Positives = 188/229 (82%) ref|ZP_00134926.1 1 COG0081 : Ribosomal protein L1 [Actinobacillus pleuropneumoniae serovar 1 str. 4074] Length = 229
5593.1 Best-BlastP=> >nrprot 40% Identities = 30/79 (37%), Positives = 45/79 (56%) ref|NP_755977.11 Hypothetical protein [Escherichia coli CFT073] gb|AAN82551 .1 |AE016767_311 Hypothetical protein [Escherichia coli CFT073] Length = 144
5595.2 Best-BlastP=> >nrprot 72% Identities = 112/174 (64%), Positives = 127/174 (72%), Gaps = 18/174 (10%) ref|ZP_00123239.1 1 COG0049: Ribosomal protein S7 [Haemophilus somnus 129PT] ref|ZP_00131785.1 | hypothetical protein [Haemophilus somnus 2336] Length = 156
5596.3 Best-BlastP=> >nrprot 90% Identities = 109/123 (88%), Positives = 115/123 (93%) ref|NP_799152.11 ribosomal protein S12 [Vibrio parahaemolyticus RIMD 2210633] sp|Q87L43|RS12_VIBPA 30S ribosomal protein S12 dbj|BAC61036.11 ribosomal protein S12 [Vibrio parahaemolyticus] Length = 124
5598.2 Best-BlastP=> >nrprot 66% Identities = 189/394 (47%), Positives = 264/394 (67%), Gaps = 2/394 (0%) ref|NP_820522.11 tryptophan/tyrosine permease family protein [Coxiella burnetii RSA 493] gb|AA091036.11 tryptophan/tyrosine permease family protein [Coxiella burnetii RSA 493] Length = 426
5600.2 Best-BlastP=> >nrprot 72% Identities = 45/66 (68%), Positives = 55/66 (83%) ref|NP_747188.11 ribosomal protein L31 [Pseudomonas putida KT2440] gb|AAN70652.1 |AE016709_4 ribosomal protein L31 [Pseudomonas putida KT2440] Length = 100
5602.3 Best-BlastP=> >nrprot 65% Identities = 60/155 (38%), Positives = 102/155 (65%), Gaps = 7/155 (4%) ref|NP_069835.11 conserved hypothetical protein [Archaeoglobus fulgidus DSM 4304] pir||B69375 phosphohistidine phosphatase (EC3.1.3.-) sixA-related [similarity] - Archaeoglobus fulgidus gb|AAB90241.11 conserved hypothetical protein [Archaeoglobus fulgidus DSM 4304] Length = 151
5605.2 Best-BlastP=> >nrprot No Hits found
5606.2 Best-BlastP=> >nrprot No Hits found 5607.2 Best-BlastP=> >nrprot 64% Identities = 221/475 (46%), Positives = 306/475 (64%), Gaps = 6/475 (1 %) ref|NP_654832.11 hypothetical protein predicted by GeneMark [Bacillus anthracis A2012] ref|NP_843400.11 alginate O-acetyltransferase, putative [Bacillus anthracis str. Ames] gb|AAP24886.11 alginate O-acetyltransferase, putative [Bacillus anthracis str. Ames] Length = 471
5609.1 Best-BlastP=> >nrprot 70% Identities = 60/84 (71 %), Positives = 67/84 (79%) ref|NP_819512.1 | hypothetical protein [Coxiella burnetii RSA 493] gb|AAO90026.11 hypothetical protein [Coxiella burnetii RSA 493] Length = 92
561.3 Best-BlastP=> >nrprot 80% Identities = 305/432 (70%), Positives = 352/432 (81 %), Gaps = 2/432 (0%) ref |NP_901046.11 probable ATPase associated with chromosome architecture [Chromobacterium violaceum ATCC 12472] gb|AAQ59051.11 probable ATPase associated with chromosome architecture [Chromobacterium violaceum ATCC 12472] Length = 443
5610.1 Best-BlastP=> >nrprot 60% Identities = 47/109 (43%), Positives = 67/109 (61 %), Gaps = 1/109 (0%) ref|ZP_00092417.11 hypothetical protein [Azotobacter vinelandii] Length = 137
5611.1 Best-BlastP=> >nrprot 43% Identities = 35/61 (57%), Positives = 47/61 (77%) ref|ZP_00092427.11 hypothetical protein [Azotobacter vinelandii] Length = 838 5615.1 Best-BlastP=> >nrprot 67% Identities = 115/242 (47%), Positives = 150/242 (61 %), Gaps = 31/242 (12%) ref|ZP_00090468.1 | COG0582: Integrase [Azotobacter vinelandii] Length = 399
562.1 Best-BlastP=> >nrprot 59% Identities = 78/206 (37%), Positives = 120/206 (58%), Gaps = 6/206 (2%) ref|ZP_00091807.1 1 COG2834: Outer membrane lipoprotein-sorting protein [Azotobacter vinelandii] Length = 207
5620.3 Best-BlastP=> >nrprot 51 % Identities = 28/61 (45%), Positives = 39/61 (63%) ref|NP_600458.11 predicted transcriptional regulator [Corynebacterium glutamicum ATCC 13032] dbj|BAB98628.1 | Predicted transcriptional regulators [Corynebacterium glutamicum ATCC 13032] Length = 75
5621.1 Best-BlastP=> >nrprot 35% Identities = 72/345 (20%), Positives = 148/345 (42%), Gaps = 30/345 (8%) ref|ZP_00118987.11 COG0477: Permeases of the major facilitator superfamily [Cytophaga hutchinsonii] Length = 441
5623.2 Best-BlastP=> >nrprot 24% Identities = 57/252 (22%), Positives = 114/252 (45%), Gaps = 15/252 (5%) ref|NP_484762.11 unknown protein [Nostoc sp. PCC 7120] pir||AE1896 hypothetical protein alr0719 [imported] - Nostoc sp. (strain PCC 7120) dbj|BAB72676.11 ORF_ID:alr0719~unknown protein [Nostoc sp. PCC 7120] Length = 393
Best-BlastP=> >nrprot 42% Identities = 47/154 (30%), Positives = 72/154 (46%), Gaps = 24/154 (15%) ref |NP_519620.11 HYPOTHETICAL PROTEIN [Ralstonia solanacearum] emb|CAD15201.11 HYPOTHETICAL PROTEIN [Ralstonia solanacearum] Length = 233
5633.2 Best-BlastP=> >nrprot 68% Identities = 143/273 (52%), Positives = 190/273 (69%), Gaps = 1/273 (0%) ref|NP_790075.11 diaminopimelate epimerase [Pseudomonas syringae pv. tomato str. DC3000] sp|Q88B09|DAPF_PSESM Diaminopimelate epimerase (DAP epimerase) gb|AAO53770.1 | diaminopimelate epimerase [Pseudomonas syringae pv. tomato str. DC3000] Length = 276
5634.2 Best-BlastP=> >nrprot 58% Identities = 16/32 (50%), Positives = 25/32 (78%) ref|ZP_00068204.11 hypothetical protein [Microbulbifer degradans 2-40] Length = 57
5636.1 Best-BlastP=> >nrprot No Hits found
5637.1 Best-BlastP=> >nrprot 58% Identities = 86/198 (43%), Positives = 126/198 (63%), Gaps = 4/198 (2%) ref|NP_840924.11 Phospholipase/Carboxylesterase [Nitrosomonas europaea ATCC 19718] emb|CAD84761.11 Phospholipase/Carboxylesterase [Nitrosomonas europaea ATCC 19718] Length = 224
5638.2 Best-BlastP=> >nrprot No Hits found
564.2 Best-BlastP=> >nrprot 68% Identities = 417/792 (52%), Positives = 546/792 (68%), Gaps = 28/792 (3%) ref|NP_251305.11 cell division protein FtsK [Pseudomonas aeruginosa PA01] sp|Q9IOM3|FTSK_PSEAE DNA translocase ftsK pir||E83318 cell division protein FtsK PA2615 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG06003.1 |AE004690_3 cell division protein FtsK [Pseudomonas aeruginosa PA01] Length = 811
5642.1 Best-BlastP=> >nrprot No Hits found 5644.3 Best-BlastP=> >nrprot 47% Identities = 193/498 (38%), Positives = 268/498 (53%), Gaps = 78/498 (15%) ref|NP_233251.11 hemagglutinin/protease [Vibrio cholerae 01 biovar eltor str. N16961] sp|P24153|HAPT_VIBCH Hemagglutinin/proteinase precursor (HA/protease) (Vibriolysin) pir||A42358 vibriolysin (EC 3.4.24.-) precursor [validated] - Vibrio cholerae (strain N16961 serogroup 01 ) gb|AAA27579.11 HA/protease gb|AAF96763.11 hemagglutinin/protease [Vibrio cholerae 01 biovar eltor str. N16961] Length = 609
5645.2 Best-BlastP=> >nrprot 82% Identities = 520/752 (69%), Positives = 622/752 (82%), Gaps = 9/752 (1 %) ref|NP_251310.11 ATP-binding protease component ClpA [Pseudomonas aeruginosa PA01] pir||B83319 ATP-binding proteinase component ClpA PA2620 [imported] - Pseudomonas aeruginosa (strain PA01) gb|AAG06008.1 |AE004690_8 ATP-binding protease component ClpA [Pseudomonas aeruginosa PA01] Length = 758
5647.1 Best-BlastP=> >nrprot 62% Identities = 55/80 (68%), Positives = 70/80 (87%) ref|NP_642326.11 conserved hypothetical protein [Xanthomonas axonopodis pv. citri str. 306] sp|Q8PL06|CLPS_XANAC ATP-dependent Clp protease adaptor protein clpS gb|AAM36862.11 conserved hypothetical protein [Xanthomonas axonopodis pv. citri str. 306] Length = 106
5648.1 Best-BlastP=> >nrprot 60% Identities = 42/112 (37%), Positives = 68/112 (60%) ref|NP_753425.11 Hypothetical protein [Escherichia coli CFT073] gb|AAN79985.1 |AE016759_259 Hypothetical protein [Escherichia coli CFT073] Length = 393
5649.1 Best-BlastP=> >nrprot 88% Identities = 327/418 (78%), Positives = 371/418 (88%) ref|NP_746141.11 isocitrate dehydrogenase, NADP- dependent, prokaryotic-type [Pseudomonas putida KT2440] gb|AAN69605.1 |AE016594_2 isocitrate dehydrogenase, NADP-dependent, prokaryotic-type [Pseudomonas putida KT2440] Length = 418
5650.3 Best-BlastP=> >nrprot 45% Identities = 67/239 (28%), Positives = 107/239 (44%), Gaps = 43/239 (17%) ref|NP_924301.11 unknown protein [Gloeobacter violaceus] dbj|BAC89296.11 glM 355 [Gloeobacter violaceus] Length = 278
5652.2 Best-BlastP=> >nrprot No Hits found 5654.2 Best-BlastP=> >nrprot No Hits found
5655.2 Best-BlastP=> >nrprot 58% Identities = 73/131 (55%), Positives = 94/131 (71 %) ref|ZP_00108601.11 COG2193: Bacterioferritin (cytochrome b1 ) [Nostoc punctiforme] Length = 144
5656.2 Best-BlastP=> >nrprot 15% Identities = 23/70 (32%), Positives = 37/70 (52%), Gaps = 1/70 (1 %) ref|NP_171674.2| expressed protein [Arabidopsis thaliana] pir||F86147 hypothetical protein T1 N6.5 [imported] - Arabidopsis thaliana gb|AAF78398.1 |AC009273_4 Contains similarity to a putative protein T2J13.100 gi|6522560 from Arabidopsis thaliana BAC T2J13 gb|AL132967 Length = 308 •
5660.1 Best-BlastP=> >nrprot 37% Identities = 69/275 (25%), Positives = 124/275 (45%), Gaps = 24/275 (8%) ref|NP_692096.11 hypothetical protein [Oceanobacillus iheyensis HTE831] dbj|BAC13131.1 | hypothetical conserved protein [Oceanobacillus iheyensis HTE831] Length = 314
5662.1 Best-BlastP=> >nrprot 65% Identities = 92/150 (61 %), Positives = 116/150 (77%) ref | NP_819966.1 | conserved hypothetical protein [Coxiella burnetii RSA 493] gb|AAO90480.11 conserved hypothetical protein [Coxiella burnetii RSA 493] Length = 414
5668.2 Best-BlastP=> >nrprot 28% Identities = 100/406 (24%), Positives = 167/406 (41 %), Gaps = 82/406 (20%) ref|NP_220907.11 unknown [Rickettsia prowazekii] pir||E71657 hypothetical protein RP534 - Rickettsia prowazekii emb|CAA14983.1 1 unknown [Rickettsia prowazekii] Length = 598
5669.2 Best-BlastP=> >nrprot 43% Identities = 125/540 (23%), Positives = 240/540 (44%), Gaps = 44/540 (8%) ref|ZP_00044012.11 hypothetical protein [Magnetococcus sp. MC-1] Length = 725
567.5 Best-BlastP=> >nrprot 46% Identities = 236/953 (24%), Positives = 425/953 (44%), Gaps = 81/953 (8%) ref|NP_052976.11 93% identical to sp:TRG1_ECOLI,gp:FPLTRAH_3[TraG of plasmid F, responsible for pilus biogenesis and stabilization of mating pairs] [Plasmid R100] gb|AAD28728.1 |AF112468_7 inner membrane and periplasmic mating pair stabilization protein TraG [Salmonella typhimurium] dbj|BAA78880.11 93% identical to sp:TRG1_ECOLI,gp:FPLTRAH_3[TraG of plasmid F, responsible for pilus biogenesis and stabilization of mating pairs] [Plasmid R100] Length = 940
5671.2 Best-BlastP=> >nrprot 74% Identities = 73/123 (59%), Positives = 94/123 (76%), Gaps = 3/123 (2%) ref|ZP_00122793.11 COG1607: Acyl-CoA hydrolase [Haemophilus somnus 129PT] ref|ZP_00133225.11 hypothetical protein [Haemophilus somnus 2336] Length = 154
5672.1 Best-BlastP=> >nrprot No Hits found
5675.2 Best-BlastP=> >nrprot 52% Identities = 113/338 (33%), Positives = 186/338 (55%), Gaps = 23/338 (6%) ref|NP_767510.11 bll0870 [Bradyrhizobium japonicum] dbj|BAC46135.1 | bll0870 [Bradyrhizobium japonicum USDA 110] Length = 329
5677.2 Best-BlastP=> >nrprot 34% Identities = 42/164 (25%), Positives = 70/164 (42%), Gaps = 21/164 (12%) gb|EAA22028.1 | putative yir4 protein [Plasmodium yoelii yoelii] Length = 299
568.2 Best-BlastP=> >nrprot 83% Identities = 380/535 (71 %), Positives = 448/535 (83%) ref |ZP_00082147.11 COG4799: Acetyl-CoA carboxylase, carboxyltransferase component (subunits alpha and beta) [Geobacter metallireducens] Length = 535
5685.2 Best-BlastP=> >nrprot 99% Identities = 405/410 (98%), Positives = 408/410 (99%) gb|AAM00606.11 unknown [Legionella pneumophila] Length = 421
5686.2 Best-BlastP=> >nrprot 70% Identities = 85/154 (55%), Positives = 112/154 (72%) ref|NP_252654.11 probable transcriptional regulator [Pseudomonas aeruginosa PA01] ref|ZP_00137402.11 COG1522: Transcriptional regulators [Pseudomonas aeruginosa UCBPP-PA14] pir||F83150 probable transcription regulator PA3965 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG07352.1 |AE004814_7 probable transcriptional regulator [Pseudomonas aeruginosa PA01] Length = 169
5687.1 Best-BlastP=> >nrprot No Hits found
5688.2 Best-BlastP=> >nrprot 39% Identities = 38/170 (22%), Positives = 75/170 (44%), Gaps = 33/170 (19%) gb|EAA16038.11 repeat organellar protein- related [Plasmodium yoelii yoelii] Length = 1441
569.2 Best-BlastP=> >nrprot 64% Identities = 136/258 (52%), Positives = 170/258 (65%), Gaps = 1/258 (0%) ref|ZP_00082146.1 | COG1024: Enoyl- CoA hydratase/carnithine racemase [Geobacter metallireducens] Length = 263
5690.2 Best-BlastP=> >nrprot 70% Identities = 75/146 (51 %), Positives = 99/146 (67%), Gaps = 9/146 (6%) ref|NP_641722.11 conserved hypothetical protein [Xanthomonas axonopodis pv. citri str. 306] gb|AAM36258.1 | conserved hypothetical protein [Xanthomonas axonopodis pv. citri str. 306] Length = 149
5694.2 Best-BlastP=> >nrprot 64% Identities = 70/152 (46%), Positives = 103/152 (67%), Gaps = 3/152 (1 %) ref |NP_719482.11 mce-related protein [Shewanella oneidensis MR-1] gb|AAN56926.1 |AE015826_11 mce-related protein [Shewanella oneidensis MR-1] Length = 157
5695.1 Best-BlastP=> >nrprot 77% Identities = 152/259 (58%), Positives = 202/259 (77%) ref|NP_931230.11 hypothetical protein [Photorhabdus luminescens subsp. laumondii TT01] emb|CAE16405.11 unnamed protein product [Photorhabdus luminescens subsp. laumondii TTO1] Length = 260
5696.2 Best-BlastP=> >nrprot 77% Identities = 168/258 (65%), Positives = 205/258 (79%) ref|NP_794201.11 toluene tolerance ABC transporter, ATP- binding protein, putative [Pseudomonas syringae pv. tomato str. DC3000] gb|AA057896.11 toluene tolerance ABC transporter, ATP-binding protein, putative [Pseudomonas syringae pv. tomato str. DC3000] Length = 269
57.1 Best-BlastP=> >nrprot 96% Identities = 272/289 (94%), Positives = 279/289 (96%) gb|AAM08235.11 LvrA [Legionella pneumophila] Length = 289 5701.2 Best-BlastP=> >nrprot 67% Identities = 179/375 (47%), Positives = 250/375 (66%), Gaps = 9/375 (2%) ref|NP_519920.11 PUTATIVE DIHYDROLIPOAMIDE ACETYLTRANSFERASE (COMPONENT E2 OF PYRUVATE DEHYDROGENASE COMPLEX) PROTEIN [Ralstonia solanacearum] emb|CAD15501.11 PUTATIVE DIHYDROLIPOAMIDE ACETYLTRANSFERASE (COMPONENT E2 OF PYRUVATE DEHYDROGENASE COMPLEX) PROTEIN [Ralstonia solanacearum] Length = 372
571.2 Best-BlastP=> >nrprot 67% Identities = 336/671 (50%), Positives = 440/671 (65%), Gaps = 23/671 (3%) ref |ZP_00082145.11 COG4770: Acetyl/propionyl-CoA carboxylase, alpha subunit [Geobacter metallireducens] Length = 668
5711.2 Best-BlastP=> >nrprot No Hits found
5719.2 Best-BlastP=> >nrprot 66% Identities = 116/246 (47%), Positives = 167/246 (67%), Gaps = 1/246 (0%) ref|ZP_00123187.11 COG0149: Triosephosphate isomerase [Haemophilus somnus 129PT] Length = 255
572.2 Best-BlastP=> >nrprot 38% Identities = 161/480 (33%), Positives = 219/480 (45%), Gaps = 84/480 (17%) pir||JC4908 alkaline serine proteinase (EC 3.4.-.-) I precursor - Alteromonas sp Length = 715
5720.1 Best-BlastP=> >nrprot No Hits found
5723.1 Best-BlastP=> >nrprot 69% Identities = 49/91 (53%), Positives = 68/91 (74%) gb|AAP83334.1 |AF469614_2 unknown [Francisella tularensis subsp. tularensis] Length = 94
5724.2 Best-BlastP=> >nrprot No Hits found
573.1 Best-BlastP=> >nrprot 79% Identities = 52/94 (55%), Positives = 75/94 (79%) ref|NP_820064.11 hypothetical protein [Coxiella burnetii RSA 493] gb|AAO90578.11 hypothetical protein [Coxiella burnetii RSA 493] Length = 94
5730.1 Best-BlastP=> >nrprot No Hits found
5735.2 Best-BlastP=> >nrprot 63% Identities = 181/388 (46%), Positives = 251/388 (64%), Gaps = 2/388 (0%) ref|NP_820522.11 tryptophan/tyrosine permease family protein [Coxiella burnetii RSA 493] gb|AA091036.11 tryptophan/tyrosine permease family protein [Coxiella burnetii RSA 493] Length = 426
5739.3 Best-BlastP=> >nrprot 31 % Identities = 357/1425 (25%), Positives = 551/1425 (38%), Gaps = 320/1425 (22%) ref|NP_772111.11 bll-5471 [Bradyrhizobium japonicum] dbj|BAC50736.1 | bll5471 [Bradyrhizobium japonicum USDA 110] Length = 4210
574.2 Best-BlastP=> >nrprot No Hits found 5741.1 Best-BlastP=> >nrprot 60% Identities = 76/174 (43%), Positives = 116/174 (66%) ref|ZP_00043325.11 COG0398: Uncharacterized conserved protein [Magnetococcus sp. MC-1] Length = 222
5743.1 Best-BlastP=> >nrprot No Hits found 5747.1 Best-BlastP=> >nrprot No Hits found 5761 .1 Best-BlastP=> >nrprot 25% Identities = 26/72 (36%), Positives = 38/72 (52%), Gaps = 1/72 (1 %) ref|NP_902894.11 probable dehydrogenase [Chromobacterium violaceum ATCC 12472] gb|AAQ60890.1 | probable dehydrogenase [Chromobacterium violaceum ATCC 12472] Length = 417
5765.1 Best-BlastP=> >nrprot 53% Identities = 39/100 (39%), Positives = 62/100 (62%) ref|NP_795325.11 ParB family protein [Pseudomonas syringae pv. tomato str. DC3000] gb|AAO59020.11 ParB family protein [Pseudomonas syringae pv. tomato str. DC3000] Length = 290
5766.1 Best-BlastP=> >nrprot 34% Identities = 39/142 (27%), Positives = 62/142 (43%), Gaps = 18/142 (12%) ref |NP_464180.11 Imo0653 [Listeria monocytogenes EGD-e] pir||AE1 156 hypothetical protein Imo0653 [imported] - Listeria monocytogenes (strain EGD-e) emb|CAC98731.1 | Imo0653 [Listeria monocytogenes] Length = 306
5767.1 Best-BlastP=> >nrprot No Hits found
5769.2 Best-BlastP=> >nrprot 58% Identities = 52/126 (41 %), Positives = 77/126 (61 %), Gaps = 4/126 (3%) ref|NP_251 91.1 ) hypothetical protein [Pseudomonas aeruginosa PA01] pir||A83296 hypothetical protein PA2801 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG06189.1 |AE004707_8 hypothetical protein PA2801 [Pseudomonas aeruginosa PA01] Length = 134
577.2 Best-BlastP=> >nrprot 37% Identities = 58/180 (32%), Positives = 107/180 (59%), Gaps = 1/180 (0%) ref|NP_624053.11 predicted transposase [Thermoanaerobacter tengcongensis] gb|AAM25657.11 predicted transposase [Thermoanaerobacter tengcongensis] Length = 267
5774.2 Best-BlastP=> >nrprot No Hits found 5776.1 Best-BlastP=> >nrprot No Hits found 5780.1 Best-BlastP=> >nrprot 61 % Identities = 56/79 (70%), Positives = 63/79 (79%) ref|NP_246141.11 HisF [Pasteurella multocida] sp|Q9CLM0|HIS6_PASMU Imidazole glycerol phosphate synthase subunit hisF (IGP synthase cyclase subunit) (IGP synthase subunit hisF) (ImGP synthase subunit hisF) (IGPS subunit hisF) gb|AAK03288.11 HisF [Pasteurella multocida] Length = 256
5796.3 Best-BlastP=> >nrprot 46% Identities = 61/195 (31 %), Positives = 92/195 (47%), Gaps = 26/195 (13%) ref|NP_890659.1 | putative ring- hydroxylating dioxygenase large subunit [Bordetella bronchiseptica] emb|CAE34488.11 putative ring-hydroxylating dioxygenase large subunit [Bordetella bronchiseptica] Length = 438
58.1 Best-BlastP=> >nrprot 95% Identities = 211/227 (92%), Positives = 217/227 (95%) gb|AAM08234.11 putative phage repressor [Legionella pneumophila] Length = 227
5804.1 Best-BlastP=> >nrprot 53% Identities = 55/137 (40%), Positives = 76/137 (55%), Gaps = 12/137 (8%) ref|NP_229825.11 cytochrome c5 [Vibrio cholerae 01 biovar eltor str. N16961] pir||F82355 cytochrome cδ VC0168 [imported] - Vibrio cholerae (strain N16961 serogroup 01 ) gb|AAF93344.11 cytochrome c5 [Vibrio cholerae 01 biovar eltor str. N16961] Length = 135
5806.2 Best-BlastP=> >nrprot 75% Identities = 54/87 (62%), Positives = 67/87 (77%) ref|ZP_00125359.11 COG0268: Ribosomal protein S20 [Pseudomonas syringae pv. syringae B728a] ref|NP_790649.11 ribosomal protein S20 [Pseudomonas syringae pv. tomato str. DC3000] gb|AA054344.11 ribosomal protein S20 [Pseudomonas syringae pv. tomato str. DC3000] Length = 92
5807.1 Best-BlastP=> >nrprot 66% Identities = 48/1 13 (42%), Positives = 76/113 (67%) sp|Q43948|HYPA_AZOCH Hydrogenase nickel incorporation protein hypA (Protein hupA) pir||JN0646 hydrogenase expression/formation protein HupA - Azotobacter chroococcum gb|AAA22132.11 hydrogenase accessory protein HupA Length = 1 13
5808.1 Best-BlastP=> >nrprot 48% Identities = 59/180 (32%), Positives = 87/180 (48%), Gaps = 13/180 (7%) gb|AAA25680.11 aminoglycoside 6'-N- acetyltransferase Length = 180
5809.1 Best-BlastP=> >nrprot No Hits found 5812.1 Best-BlastP=> >nrprot 60% Identities = 57/154 (37%), Positives = 95/154 (61 %), Gaps = 6/154 (3%) emb|CAB60049.1 | IvrB [Legionella pneumophila] Length = 157
5813.1 Best-BlastP=> >nrprot 65% Identities = 30/57 (52%), Positives = 43/57 (75%) emb|CAB60050.11 IvrC [Legionella pneumophila] Length = 67
5814.1 Best-BlastP=> >nrprot 45% Identities = 38/121 (31 %), Positives = 66/121 (54%), Gaps = 6/121 (4%) gb|AAL05416.1 | PilL [Yersinia pseudotuberculosis] Length = 356
5816.1 Best-BlastP=> >nrprot 69% Identities = 52/96 (54%), Positives = 79/96 (82%) ref|ZP_00122751.11 COG0718: Uncharacterized protein conserved in bacteria [Haemophilus somnus 129PTJ ref |ZP_00132634.1 | hypothetical protein [Haemophilus somnus 2336] Length = 109
5818.1 Best-BlastP=> >nrprot No Hits found
5821.1 Best-BlastP=> >nrprot No H its fou nd
5822.1 Best-BlastP=> >nrprot No Hits found
5823.2 Best-BlastP=> >nrprot 49% Identities = 125/369 (33%), Positives = 198/369 (53%), Gaps = 33/369 (8%) ref|NP_899726.11 probable aminopeptidase [Chromobacterium violaceum ATCC 12472] gb|AAQ57736.11 probable aminopeptidase [Chromobacterium violaceum ATCC 12472] Length = 415
5824.1 Best-BlastP=> >nrprot 56% Identities = 43/67 (64%), Positives = 53/67 (79%) ref|NP_773113.11 bll6473 [Bradyrhizobium japonicum] dbj|BAC51738.1 | bll6473 [Bradyrhizobium japonicum USDA 110] Length = 345
5829.2 Best-BlastP=> >nrprot 50% Identities = 67/218 (30%), Positives = 124/218 (56%), Gaps = 1/218 (0%) ref|NP_880554.11 putative glutamine- binding periplasmic protein precursor [Bordetella pertussis] emb|CAE42138.11 putative glutamine-binding periplasmic protein precursor [Bordetella pertussis] Length = 247
583.2 Best-BlastP=> >nrprot 82% Identities = 128/198 (64%), Positives = 164/198 (82%), Gaps = 1/198 (0%) ref|NP_743801.11 trp repressor binding protein [Pseudomonas putida KT2440] gb|AAN67265.1 |AE016353_5 trp repressor binding protein [Pseudomonas putida KT2440] Length = 201
5844.1 Best-BlastP=> >nrprot 65% Identities = 66/135 (48%), Positives = 88/135 (65%), Gaps = 5/135 (3%) ref|NP_044227.11 KlcA [Enterobacter aerogenes] ref|NP_862440.1 | KlcA protein [Pseudomonas sp. ADP] sp|P52602|KLA1_ECOLI Antirestriction protein klcA pir||T08486 probable aniti-restriction protein klcA - Enterobacter aerogenes plasmid R751 gb|AAC64430.11 KlcA [Enterobacter aerogenes] gb|AAK50236.1 |U66917_3 KlcA protein [Pseudomonas sp. ADP] Length = 142
5846.2 Best-BlastP=> >nrprot No Hits found 5848.1 Best-BlastP=> >nrprot No Hits found 585.2 Best-BlastP=> >nrprot 72% Identities = 62/118 (52%), Positives = 85/118 (72%), Gaps = 1/118 (0%) ref|NP_274848.11 conserved hypothetical protein [Neisseria meningitidis MC58] pir||A81035 conserved hypothetical protein NMB1852 [imported] - Neisseria meningitidis (strain MC58 serogroup B) gb|AAF42186.1 | conserved hypothetical protein [Neisseria meningitidis MC58] Length = 129
5856.1 Best-BlastP=> >nrprot 42% Identities = 20/43 (46%), Positives = 27/43 (62%), Gaps = 1/43 (2%) ref|ZP_00141162.11 COG0617: tRNA nucleotidyltransferase/poly(A) polymerase [Pseudomonas aeruginosa UCBPP-PA14] Length = 467
5857.1 Best-BlastP=> >nrprot 64% Identities = 65/139 (46%), Positives = 92/139 (66%) ref|NP_735626.11 Unknown [Streptococcus agalactiae NEM316] emb|CAD46839.11 Unknown [Streptococcus agalactiae NEM316] Length = 162
5867.1 Best-BlastP=> >nrprot No Hits found
587.2 Best-BlastP=> >nrprot 74% Identities = 14g/254 (58%), Positives = 194/254 (76%), Gaps = 1/254 (0%) gb|EAA20230.11 exodeoxyribonuclease III [Plasmodium yoelii yoelii] Length = 271
5871.3 Best-BlastP=> >nrprot 46% Identities = 94/364 (25%), Positives = 174/364 (47%), Gaps = 21/364 (5%) ref |NP_691531.1 | transposase for IS652 [Oceanobacillus iheyensis HTE831] ref|NP_692364.1 | hypothetical protein [Oceanobacillus iheyensis HTE831] ref|NP_693263.1 | transposase for IS652 [Oceanobacillus iheyensis HTE831] dbj|BAC12566.11 transposase for IS652 [Oceanobacillus iheyensis HTE831] dbj|BAC13399.11 hypothetical conserved protein [Oceanobacillus iheyensis HTE831] dbj|BAC14298.1 | transposase for IS652 [Oceanobacillus iheyensis HTE831] Length = 402 5872.1 Best-BlastP=> >nrprot No Hits found
5874.1 Best-BlastP=> >nrprot 71 % Identities = 96/178 (53%), Positives = 128/178 (71 %), Gaps = 1/178 (0%) ref|ZP_00024697.11 COG4584: Transposase and inactivated derivatives [Ralstonia metallidurans] Length = 343
5875.1 Best-BlastP=> >nrprot 43% Identities = 28/90 (31 %), Positives = 44/90 (48%) gb|AAN34371.11 ORF1 transposase [Acinetobacter baumannii] Length = 180
5876.2 Best-BlastP=> >nrprot 71% Identities = 121/219 (55%), Positives = 165/219 (75%), Gaps = 1/219 (0%) ref|NP_791574.11 cytidylate kinase [Pseudomonas syringae pv. tomato str. DC3000] gb|AA055269.11 cytidylate kinase [Pseudomonas syringae pv. tomato str. DC3000] Length = 229
588.3 Best-BlastP=> >nrprot 72% Identities = 146/252 (57%), Positives = 186/252 (73%) ref|NP_840124.11 Exodeoxyribonuclease llhExodeoxyribonuclease III xth [Nitrosomonas europaea ATCC 19718] emb|CAD83934.1 | Exodeoxyribonuclease MhExodeoxyribonuclease III xth [Nitrosomonas europaea ATCC 19718] Length = 254
58gθ.1 Best-BlastP=> >nrprot No Hits found
5892.1 Best-BlastP=> >nrprot 73% Identities = 59/99 (59%), Positives = 73/99 (73%) ref|NP_742622.11 ribosomal protein L23 [Pseudomonas putida KT2440] gb|AAN66086.1 |AE016238_4 ribosomal protein L23 [Pseudomonas putida KT2440] Length = 99
5894.3 Best-BlastP=> >nrprot 82% Identities = 202/275 (73%), Positives = 229/275 (83%) ref|ZP_00067989.11 COG0090: Ribosomal protein L2 [Microbulbifer degradans 2-40] Length = 275
5895.3 Best-BlastP=> >nrprot 86% Identities = 65/90 (72%), Positives = 80/90 (88%) ref|NP_882398.11 30S ribosomal protein S19 [Bordetella parapertussis] ref|NP_886586.11 30S ribosomal protein S19 [Bordetella bronchiseptica] emb|CAE39774.11 30S ribosomal protein S19 [Bordetella parapertussis] emb|CAE30535.11 30S ribosomal protein S19 [Bordetella bronchiseptica] Length = 91
5896.1 Best-BlastP=> >nrprot 62% Identities = 42/115 (36%), Positives = 75/115 (65%) ref|NP_286169.11 putative transport protein [Escherichia coli 0157:H7 EDL933] ref|NP_308508.1 | putative transport protein [Escherichia coli 0157:H7] pir||A90689 probable transport protein ECs0481 [imported] - Escherichia coli (strain 0157:H7, substrain RIMD 0509952) pir||E85539 probable transport protein yajR [imported] - Escherichia coli (strain 0157:H7, substrain EDL933) gb|AAG54777.1 |AE005222_2 putative transport protein [Escherichia coli 0157:H7 EDL933] dbj|BAB33904.11 putative transport protein [Escherichia coli 0157:H7] Length = 456
5897.1 Best-BlastP=> >nrprot 55% Identities = 41/73 (56%), Positives = 51/73 (69%) ref|NP_931085.11 hypothetical protein [Photorhabdus luminescens subsp. laumondii TT01] emb|CAE16252.11 unnamed protein product [Photorhabdus luminescens subsp. laumondii TT01] Length = 459
5898.2 Best-BlastP=> >nrprot 97% Identities = 266/271 (98%), Positives = 268/271 (98%) gb|AAC83338.11 major outer membrane protein precursor [Legionella pneumophila] gb|AAC83342.11 major outer membrane protein precursor [Legionella pneumophila] Length = 289
5899.1 Best-BlastP=> >nrprot No Hits found
59.1 Best-BlastP=> >nrprot 95% Identities = 137/145 (94%), Positives = 142/145 (97%) gb|AAM08233.1 | unknown [Legionella pneumophila] Length = 243
590.4 Best-BlastP=> >nrprot 17% Identities = 93/427 (21 %), Positives = 182/427 (42%), Gaps = 50/427 (11 %) ref|NP_603419.11 Exonuclease SBCC [Fusobacterium nucleatum subsp. nucleatum ATCC 25586] gb|AAL94718.1 | Exonuclease SBCC [Fusobacterium nucleatum subsp. nucleatum ATCC 25586] Length = 921
5900.3 Best-BlastP=> >nrprot No Hits found
5901.3 Best-BlastP=> >nrprot 17% Identities = 49/177 (27%), Positives = 85/177 (48%), Gaps = 27/177 (15%) ref|NP_001259.11 RCC1-like G exchanging factor RLG [Homo sapiens] pir||T50663 RCC1-like G exchanging factor RLG [imported] - human gb|AAC79987.11 RCC1-like G exchanging factor RLG [Homo sapiens] gb|AAH29052.1 | RCC1-like G exchanging factor RLG [Homo sapiens] gb|AAP88928.1 | chromosome condensation 1 -like [Homo sapiens] Length = 551
5903.2 Best-BlastP=> >nrprot 76% Identities = 191/318 (60%), Positives = 247/318 (77%) ref|NP_819776.11 KpsF/GutQ family protein [Coxiella burnetii RSA 493] gb|AAO90290.11 KpsF/GutQ family protein [Coxiella burnetii RSA 493] Length = 324
5904.2 Best-BlastP=> >nrprot 21 % Identities = 25/100 (25%), Positives = 46/100 (46%), Gaps = 6/100 (6%) ref|NP_283688.1 | hypothetical protein NMA0899 [Neisseria meningitidis Z2491] pir||D81936 hypothetical protein NMA0899 [imported] - Neisseria meningitidis (strain Z2491 serogroup A) emb|CAB84177.1 | hypothetical protein NMA0899 [Neisseria meningitidis Z2491] Length = 124
5905.1 Best-BlastP=> >nrprot 53% Identities = 118/304 (38%), Positives = 165/304 (54%), Gaps = 7/304 (2%) ref|NP_902888.11 probable translation initiation protein, Suaδ/YciO/YrdC family [Chromobacterium violaceum ATCC 12472] gb|AAQ60884.11 probable translation initiation protein, Sua5/YciO/YrdC family [Chromobacterium violaceum ATCC 12472] Length = 321
5906.1 Best-BlastP=> >nrprot 100% Identities = 138/138 (100%), Positives = 138/138 (100%) emb|CAA67994.1 | oxaloacetate decarboxylase alpha- chain [Legionella pneumophila] Length = 596
5907.1 Best-BlastP=> >nrprot 56% Identities = 20/31 (64%), Positives = 30/31 (96%) ref|NP_283131.11 hypothetical protein NMA0292 [Neisseria meningitidis Z2491] pir||G82024 hypothetical protein NMA0292 [imported] - Neisseria meningitidis (strain Z2491 serogroup A) emb|CAB83599.11 hypothetical protein NMA0292 [Neisseria meningitidis Z2491] Length = 94
591.3 Best-BlastP=> >nrprot 74% Identities = 156/258 (60%), Positives = 198/258 (76%) ref|ZP_00024696.11 COG1484: DNA replication protein [Ralstonia metallidurans] Length = 268
5910.1 Best-BlastP=> >nrprot No Hits found
592.3 Best-BlastP=> >nrprot 96% Identities = 531/559 (94%), Positives = 540/559 (96%) gb|AAM00619.11 unknown [Legionella pneumophila] Length = 559
5920.2 Best-BlastP=> >nrprot 46% Identities = 94/364 (25%), Positives = 174/364 (47%), Gaps = 21/364 (5%) ref|NP_691531.11 transposase for IS652 [Oceanobacillus iheyensis HTE831] ref|NP_692364.1 | hypothetical protein [Oceanobacillus iheyensis HTE831] ref|NP_693263.1 | transposase for IS652 [Oceanobacillus iheyensis HTE831] dbj|BAC12566.11 transposase for IS652 [Oceanobacillus iheyensis HTE831] dbj|BAC13399.1 | hypothetical conserved protein [Oceanobacillus iheyensis HTE831] dbj(BAC14298.1 | transposase for IS652 [Oceanobacillus iheyensis HTE831] Length = 402
5926.1 Best-BlastP=> >nrprot 59% Identities = 36/74 (48%), Positives = 47/74 (63%) ref|NP_355900.11 AGR_L_236p [Agrobacterium tumefaciens] ref|NP_535243.1 | conserved hypothetical protein [Agrobacterium tumefaciens str. C58 (U. Washington)] pir||C98145 hypothetical protein AGR_L_236 [imported] - Agrobacterium tumefaciens (strain C58, Cereon) pir||AI3142 conserved hypothetical protein Atu4765 [imported] - Agrobacterium tumefaciens (strain C58, Dupont) gb|AAK88685.11 AGR_L_236p [Agrobacterium tumefaciens str. C58 (Cereon)] gb|AAL45559.1 | conserved hypothetical protein [Agrobacterium tumefaciens str. C58 (U. Washington)] Length = 507
593.2 Best-BlastP=> >nrprot No Hits found
5935.1 Best-BlastP=> >nrprot No Hits found
594.1 Best-BlastP=> >nrprot 97% Identities = 145/150 (96%), Positives = 147/150 (98%) gb|AAM00620.1 | chemiosmotic efflux system C protein A [Legionella pneumophila] Length = 150
5944.1 Best-BlastP=> >nrprot No Hits found
5947.1 Best-BlastP=> >nrprot 65% Identities = 17/39 (43%), Positives = 30/39 (76%) ref|NP_469382.11 similar to E. coli DedA protein [Listeria innocua] pir||AD1437 E. coli DedA protein homolog Iin0035 [imported] - Listeria innocua (strain Clipl 1262) emb|CAC95268.11 Iin0035 [Listeria innocua] Length = 219
5950.1 Best-BlastP=> >nrprot No Hits found 5952.1 Best-BlastP=> >nrprot 79% Identities = 49/52 (94%), Positives = 49/52 (94%) gb|AAM00633.11 unknown [Legionella pneumophila] Length = 176 5953.1 Best-BlastP=> >nrprot 87% Identities = 48/57 (84%), Positives = 50/57 (87%) sp|Q48815|HELA_LEGPN Protein helA gb|AAB05679.11 HelA Length = 1052
5957.1 Best-BlastP=> >nrprot 68% Identities = 162/270 (60%), Positives = 214/270 (79%) ref|ZP_00122424.11 COG1185: Polyribonucleotide nucleotidyltransferase (polynucleotide phosphorylase) [Haemophilus somnus 129PT] Length = 713
596.3 Best-BlastP=> >nrprot 73% Identities = 314/570 (55%), Positives = 419/570 (73%), Gaps = 5/570 (0%) ref|NP_819843.11 malate oxidoreductase [Coxiella burnetii RSA 493] gb|AAO90357.11 malate oxidoreductase [Coxiella burnetii RSA 493] Length = 565
597.2 Best-BlastP=> >nrprot 37% Identities = 29/80 (36%), Positives = 46/80 (57%), Gaps = 1/80 (1 %) ref|NP_820267.11 hypothetical protein [Coxiella burnetii RSA 493] gb|AAO90781.11 hypothetical protein [Coxiella burnetii RSA 493] Length = 160
5978.2 Best-BlastP=> >nrprot 68% Identities = 92/166 (55%), Positives = 126/166 (75%) ref|NP_931234.11 hypothetical protein [Photorhabdus luminescens subsp. laumondii TT01] emb|CAE16409.1 | unnamed protein product [Photorhabdus luminescens subsp. laumondii TT01] Length = 187
5979.1 Best-BlastP=> >nrprot No Hits found
598.2 Best-BlastP=> >nrprot 46% Identities = 36/101 (35%), Positives = 49/101 (48%), Gaps = 15/101 (14%) ref|NP_249404.1 | hypothetical protein [Pseudomonas aeruginosa PA01] pir||A83555 hypothetical protein PA0713 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG04102.1 |AE004507_1 hypothetical protein PA0713 [Pseudomonas aeruginosa PA01] Length = 97
5983.1 Best-BlastP=> >nrprot No Hits found 5985.1 Best-BlastP=> >nrprot 44% Identities = 29/106 (27%), Positives = 56/106 (52%), Gaps = 4/106 (3%) ref|NP_692567.11 transposase for IS652 [Oceanobacillus iheyensis HTE831] dbj|BAC13602.11 transposase for IS652 [Oceanobacillus iheyensis HTE831] Length = 402
5986.2 Best-BlastP=> >nrprot No Hits found 5987.1 Best-BlastP=> >nrprot No Hits found 5988.1 Best-BlastP=> >nrprot 35% Identities = 27/55 (49%), Positives = 31/55 (56%), Gaps = 3/55 (5%) dbj|BAC94688.11 hypothetical protein [Vibrio vulnificus YJ016] Length = 343
5996.1 Best-BlastP=> >nrprot 51 % Identities = 67/159 (42%), Positives = 90/159 (56%), Gaps = 3/159 (1 %) ref|NP_742500.11 conserved hypothetical protein [Pseudomonas putida KT2440] gb|AAN65964.1 |AE016224_8 conserved hypothetical protein [Pseudomonas putida KT2440] Length = 166
5999.1 Best-BlastP=> >nrprot 69% Identities = 49/91 (53%), Positives = 68/91 (74%) gb|AAP83334.1 |AF469614_2 unknown [Francisella tularensis subsp. tularensis] Length = 94
60.1 Best-BlastP=> >nrprot No Hits found
600.3 Best-BlastP=> >nrprot 73% Identities = 117/172 (68%), Positives = 137/172 (79%) ref|NP_220600.1 | ABC TRANSPORTER ATP-BINDING PROTEIN (abcT3) [Rickettsia prowazekii] pir||F71732 ABC transporter ATP-binding protein (abcT3) RP214 - Rickettsia prowazekii emb|CAA14677.11 ABC TRANSPORTER ATP-BINDING PROTEIN (abcT3) [Rickettsia prowazekii] Length = 548
6002.1 Best-BlastP=> >nrprot 30% Identities = 78/169 (46%), Positives = 111/169 (65%), Gaps = 2/169 (1 %) gb|AAN34371.1 | ORF1 transposase [Acinetobacter baumannii] Length = 180
6004.1 Best-BlastP=> >nrprot No Hits found 6007.1 Best-BlastP=> >nrprot No Hits found 601.4 Best-BlastP=> >nrprot 45% Identities = 97/361 (26%), Positives = 180/361 (49%), Gaps = 13/361 (3%) ref|NP_359923.11 multidrug resistance ABC transporter ATP-binding protein [Rickettsia conorii] pir||F97735 hypothetical protein abcT3 [imported] - Rickettsia conorii (strain Malish 7) gb|AAL02824.11 multidrug resistance ABC transporter ATP-binding protein [Rickettsia conorii] Length = 589
6015.1 Best-BlastP=> >nrprot No Hits found
6016.1 Best-BlastP=> >nrprot 40% Identities = 31/71 (43%), Positives = 42/71 (59%), Gaps = 1/71 (1 %) ref|NP_774998.1 | putative protein [Citrobacter freundii] gb|AAN87662.11 putative protein [Citrobacter freundii] Length = 112
6018.1 Best-BlastP=> >nrprot No Hits found
6019.1 Best-BlastP=> >nrprot 56% Identities = 67/145 (46%), Positives = 100/145 (68%) ref|NP_635334.11 transcriptional regulator, AraC family [Methanosarcina mazei Goel] gb|AAM33006.11 transcriptional regulator, AraC family [Methanosarcina mazei Goel] Length = 209
602.4 Best-BlastP=> >nrprot 39% Identities = 64/253 (25%), Positives = 125/253 (49%), Gaps = 21/253 (8%) ref|NP_834835.11 Transcriptional regulator, MerR family [Bacillus cereus ATCC 14579] gb|AAP12036.11 Transcriptional regulator, MerR family [Bacillus cereus ATCC 14579] Length = 254
6029.1 Best-BlastP=> >nrprot 70% Identities = 31/48 (64%), Positives = 39/48 (81 %) ref |NP_520567.11 PROBABLE 50S RIBOSOMAL SUBUNIT PROTEIN L33 [Ralstonia solanacearum] sp|Q8XWM8|RL33_RALSO 50S ribosomal protein L33 emb|CAD16153.1 | PROBABLE 50S RIBOSOMAL SUBUNIT PROTEIN L33 [Ralstonia solanacearum] Length = 56
6036.1 Best-BlastP=> >nrprot 99% Identities = 110/110 (100%), Positives = 110/110 (100%) emb|CAC33488.11 hypothetical protein [Legionella pneumophila] Length = 110
6037.1 Best-BlastP=> >nrprot 47% Identities = 33/78 (42%), Positives = 54/78 (69%) sp|P04928|SANT_PLAFN S-ANTIGEN PROTEIN PRECURSOR pir||YAZQN7 S-antigen precursor - malaria parasite (Plasmodium falciparum) (strain NF7/Ghana) gb|AAA29758.11 S antigen precursor Length = 309
6038.1 Best-BlastP=> >nrprot No Hits found
6039.1 Best-BlastP=> >nrprot 82% Identities = 366/558 (65%), Positives = 460/558 (82%), Gaps = 3/558 (0%) ref|NP_251852.11 30S ribosomal protein S1 [Pseudomonas aeruginosa PA01] pir||C83250 30S ribosomal protein S1 PA3162 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG06550.1 |AE004740_3 30S ribosomal protein S1 [Pseudomonas aeruginosa PA01] Length = 559
604*3 Best-BlastP=> >nrprot 55% Identities = 104/249 (41 %), Positives = 138/249 (55%), Gaps = 6/249 (2%) ref|NP_419511.11 transcriptional regulator SkgA [Caulobacter crescentus CB15] sp|Q9RP67|SKGA_CAUCR Transcriptional regulator skgA (Stationary-phase regulation of katG protein) pir||C87335 transcription regulator SkgA [imported] - Caulobacter crescentus gb|AAF01797.1 |AF170912_1 putative helix-turn-helix transcriptional regulator SkgA [Caulobacter crescentus] gb|AAK22679.11 transcriptional regulator SkgA [Caulobacter crescentus CB15] Length = 255
6040.1 Best-BlastP=> >nrprot 71 % Identities = 54/92 (58%), Positives = 72/92 (78%) ref |NP_819558.11 3-phosphoshikimate 1-carboxyvinyltransferase [Coxiella burnetii RSA 493] gb|AAO90072.1 | 3-phosphoshikimate 1-carboxyvinyltransferase [Coxiella burnetii RSA 493] Length = 438
6042.1 Best-BlastP=> >nrprot 69% Identities = 105/209 (50%), Positives = 149/209 (71 %), Gaps = 2/209 (0%) ref |ZP_00122964.1 | COG0125: Thymidylate kinase [Haemophilus somnus 129PT] Length = 210
6044.1 Best-BlastP=> >nrprot No Hits found
6049.1 Best-BlastP=> >nrprot 74% Identities = 123/208 (59%), Positives = 155/208 (74%), Gaps = 2/208 (0%) ref|NP_903586.11 probable electron- transferring-flavoprotein dehydrogenase [Chromobacterium violaceum ATCC 12472] gb|AAQ61577.1 | probable electron-transferring- flavoprotein dehydrogenase [Chromobacterium violaceum ATCC 12472] Length = 539
6052.1 Best-BlastP=> >nrprot 47% Identities = 55/162 (33%), Positives = 91/162 (56%), Gaps = 2/162 (1%) ref |NP_719829.11 conserved hypothetical protein [Shewanella oneidensis MR-1] gb|AAN57273.1 |AE015863_2 conserved hypothetical protein [Shewanella oneidensis MR-1] Length = 191
606.3 Best-BlastP=> >nrprot 56% Identities = 199/513 (38%), Positives = 301/513 (58%), Gaps = 6/513 (1 %) ref|NP_819600.11 amino acid permease family protein [Coxiella burnetii RSA 493] gb|AAO90114.11 amino acid permease family protein [Coxiella burnetii RSA 493] Length = 560
6061.1 Best-BlastP=> >nrprot No Hits found
6062.1 Best-BlastP=> >nrprot 62% Identities = 75/166 (45%), Positives = 108/166 (65%), Gaps = 2/166 (1 %) gb|AAN34371.1 | ORF1 transposase [Acinetobacter baumannii] Length = 180
6064.1 Best-BlastP=> >nrprot No Hits found
6066.1 Best-BlastP=> >nrprot 35% Identities = 34/122 (27%), Positives = 58/122 (47%), Gaps = 10/122 (8%) ref|NP_242278.11 transposase (01 ) [Bacillus halodurans] pir||D83826 transposase (01 ) BH1412 [imported] - Bacillus halodurans (strain C-125) dbj|BAB05131.1 | transposase (01 ) [Bacillus halodurans] Length = 405
6070.1 Best-BlastP=> >nrprot 65% Identities = 59/121 (48%), Positives = 85/121 (70%) ref|NP_781505.11 2-hydroxyacid dehydrogenase [Clostridium tetani E88] gb|AA035442.11 2-hydroxyacid dehydrogenase [Clostridium tetani E88] Length = 357
6071.1 Best-BlastP=> >nrprot 43% Identities = 23/67 (34%), Positives = 31/67 (46%) gb|AAK61303.11 putative transposase [Xanthomonas oryzae pv. oryzae] Length = 344
6072.1 Best-BlastP=> >nrprot 45% Identities = 18/43 (41 %), Positives = 29/43 (67%) gb|AAB38861.11 putative transposase [Burkholderia cepacia] Length = 345
6074.1 Best-BlastP=> >nrprot 49% Identities = 48/175 (27%), Positives = 84/175 (48%), Gaps = 14/175 (8%) ref|NP_692337.11 transposase for IS652 [Oceanobacillus iheyensis HTE831] ref|NP_693308.1 | transposase for IS652 [Oceanobacillus iheyensis HTE831] dbj|BAC13372.11 transposase for IS652 [Oceanobacillus iheyensis HTE831] dbj|BAC14343.11 transposase for IS652 [Oceanobacillus iheyensis HTE831] Length = 397
6079.1 Best-BlastP=> >nrprot 44% Identities = 35/96 (36%), Positives = 49/96 (51 %) gb|AAB97874.11 surface antigen [Trypanosoma cruzi] Length = 722
6081.1 Best-BlastP=> >nrprot 45% Identities = 24/44 (54%), Positives = 32/44 (72%) dbj|BAC93915.11 L-asparaginase I [Vibrio vulnificus YJ016] Length = 337
6083.1 Best-BlastP=> >nrprot No Hits found
6086.1 Best-BlastP=> >nrprot 98% Identities = 102/104 (98%), Positives = 103/104 (99%) gb|AAM00607.11 unknown [Legionella pneumophila] Length = 121
6087.1 Best-BlastP=> >nrprot No Hits found
6089.1 Best-BlastP=> >nrprot 49% Identities = 48/175 (27%), Positives = 84/175 (48%), Gaps = 14/175 (8%) ref|NP_692337.11 transposase for IS652 [Oceanobacillus iheyensis HTE831] ref|NP_693308.11 transposase for IS652 [Oceanobacillus iheyensis HTE831] dbj|BAC13372.11 transposase for IS652 [Oceanobacillus iheyensis HTE831 ] dbj|BAC14343.1 1 transposase for IS652 [Oceanobacillus iheyensis HTE831 ] Length = 397
6093.1 Best-BlastP=> >nrprot No Hits found
6094.1 Best-BlastP=> >nrprot No Hits found
6097.1 Best-BlastP=> >nrprot 41 % Identities = 1 129/1221 (92%), Positives = 1163/1221 (95%), Gaps = 16/1221 (1 %) gb|AAD41583.1 |AF057703_1 structural toxin protein RtxA [Legionella pneumophila] Length = 1208
6098.1 Best-BlastP=> >nrprot 50% Identities = 41/74 (55%), Positives = 53/74 (71 %) ref|NP_927457.11 DNA adenine methylase (Deoxyadenosyl- methyltransferase) [Photorhabdus luminescens subsp. laumondii TT01] emb|CAE12382.11 DNA adenine methylase (Deoxyadenosyl- methyltransferase) [Photorhabdus luminescens subsp. laumondii TT01 ] Length = 270
61.1 Best-BlastP=> >nrprot No Hits found
6106.1 Best-BlastP=> >nrprot 37% Identities = 20/56 (35%), Positives = 32/56 (57%) ref|NP_051664.11 transposase, putative [Deinococcus radiodurans] pir||A75633 probable transposase - Deinococcus radiodurans (strain R1 ) gb|AAF12606.1 |AE001826_75 transposase, putative [Deinococcus radiodurans] Length = 327
6107.1 Best-BlastP=> >nrprot No Hits found
6108.1 Best-BlastP=> >nrprot 63% Identities = 76/169 (44%), Positives = 110/169 (65%), Gaps = 2/169 (1 %) gb|AAN34371.1 | ORF1 transposase [Acinetobacter baumannii] Length = 180
61 10.1 Best-BlastP=> >nrprot 85% Identities = 74/77 (96%), Positives = 74/77 (96%) emb|CAB65201.11 hypothetical protein [Legionella pneumophila] Length = 356
6117.1 Best-BlastP=> >nrprot 11 % Identities = 83/277 (29%), Positives = 126/277 (45%), Gaps = 51/277 (18%) ref|XP_300615.1 | similar to hypothetical protein [Homo sapiens] Length = 423
6119.1 Best-BlastP=> >nrprot 41 % Identities = 34/123 (27%), Positives = 62/123 (50%), Gaps = 4/123 (3%) ref|NP_692567.11 transposase for IS652 [Oceanobacillus iheyensis HTE831] dbj|BAC13602.11 transposase for IS652 [Oceanobacillus iheyensis HTE831] Length = 402
612.2 Best-BlastP=> >nrprot 97% Identities = 696/721 (96%), Positives = 706/721 (97%) gb|AAC64361.11 catalase-peroxidase KatB [Legionella pneumophila] Length = 721
6121.1 Best-BlastP=> >nrprot 52% Identities = 51/199 (25%), Positives = 100/199 (50%), Gaps = 21/199 (10%) gb|AAB05879.11 putative transposase Length = 424 6122.1 Best-BlastP=> >nrprot No Hits found 6123.1 Best-BlastP=> >nrprot No Hits found 6125.1 Best-BlastP=> >nrprot 98% Identities = 51/51 (100%), Positives = 51/51 (100%) pir||T42167 Rep protein - Escherichia coli plasmid p0157 gb|AAC70135.11 Rep protein E1 [Escherichia coli 0157:H7] Length = 51
6128.1 Best-BlastP=> >nrprot 99% Identities = 388/388 (100%), Positives = 388/388 (100%) ref|NP_061425.11 100 pet identical to gp:FPLMCG_6[SopA of plasmid F] [Plasmid F] gb|AAA24902.11 Protein A dbj|BAA97916.11 100 pet identical to gp:FPLMCG_6[SopA of plasmid F] [Plasmid F] emb|CAC79981.1 | orf1176 [Escherichia coli] gb|AA061293.11 ParA [BAC cloning vector pEBAC190G] emb|CAD50597.11 SopA protein [Cloning vector pUvBBAC] Length = 391
6130-1 Best-BlastP=> >nrprot 99% Identities = 323/323 (100%), Positives = 323/323 (100%) ref|NP_061426.11 100 pet identical to sp:SOPB_ECOLI[SopB of plasmid F] [Plasmid F] ref|NP_052641.11 SopB protein [Escherichia coli 0157:H7] sp|P08867|SOPB_ECOLI SopB protein (Plasmid partition protein B) pir||T00244 sopB protein - Escherichia coli plasmids p0157 and F emb|CAA28296.1 | unnamed protein product [Escherichia coli] gb|AAA24903.1 | Protein B gb|AAC53637.1 | SopB dbj|BAA31791.1 | SopB protein [Escherichia coli] gb|AAC70137.1 | plasmid partitioning protein [Escherichia coli 0157:H7] dbj|BAA97917.1 | 100 pet identical to sp:SOPB_ECOLI[SopB of plasmid F] [Plasmid F] emb|CAC79980.1 | orf972 [Escherichia coli] gb|AA061294.11 ParB [BAC cloning vector pEBAC190G] emb|CAD50598.1 | SopB protein [Cloning vector pUvBBAC] Length = 323
6131.1 Best-BlastP=> >nrprot 47% Identities = 239/927 (25%), Positives = 368/927 (39%), Gaps = 188/927 (20%) ref|NP_772111.11 bll5471 [Bradyrhizobium japonicum] dbj|BAC50736.11 bll5471 [Bradyrhizobium japonicum USDA 110] Length = 4210
6132.1 Best-BlastP=> >nrprot 37% Identities = 21/57 (36%), Positives = 33/57 (57%) ref|NP_051664.11 transposase, putative [Deinococcus radiodurans] pir||A75633 probable transposase - Deinococcus radiodurans (strain R1 ) gb|AAF12606.1 |AE001826_75 transposase, putative [Deinococcus radiodurans] Length = 327
6133.1 Best-BlastP=> >nrprot 67% Identities = 44/80 (55%), Positives = 59/80 (73%) ref|NP_273113.11 conserved hypothetical protein [Neisseria meningitidis MC58] pir||B81244 conserved hypothetical protein NMB0047 [imported] - Neisseria meningitidis (strain MC58 serogroup B) gb|AAF40518.1 | conserved hypothetical protein [Neisseria meningitidis MC58] Length = 94
6134.1 Best-BlastP=> >nrprot 54% Identities = 78/169 (46%), Positives = 111/169 (65%), Gaps = 2/169 (1 %) gb|AAN34371.1 | ORF1 transposase [Acinetobacter baumannii] Length = 180
615.5 Best-BlastP=> >nrprot 53% Identities = 58/230 (25%), Positives = 114/230 (49%), Gaps = 26/230 (11%) ref|NP_603496.1 | Transposase [Fusobacterium nucleatum subsp. nucleatum ATCC 25586] gb|AAL94795.1 | Transposase [Fusobacterium nucleatum subsp. nucleatum ATCC 25586] Length = 428
616.4 Best-BlastP=> >nrprot 67% Identities = 179/333 (53%), Positives = 231/333 (69%), Gaps = 4/333 (1%) ref|ZP_00024697.1 | COG4584: Transposase and inactivated derivatives [Ralstonia metallidurans] Length = 343
618.3 Best-BlastP=> >nrprot 55% Identities = 48/125 (38%), Positives = 77/125 (61 %), Gaps = 2/125 (1 %) ref|NP_867668.11 probable acyl carrier protein [Pirellula sp.] emb|CAD75215.11 probable acyl carrier protein [Pirellula sp.] Length = 129
619.1 Best-BlastP=> >nrprot 19% Identities = 32/118 (27%), Positives = 58/118 (49%), Gaps = 2/118 (1 %) ref|NP_785252.11 (3R)-hydroxymyristoyl- [acyl carrier protein] dehydratase [Lactobacillus plantarum WCFS1] sp|Q88WG9|FABZ_LACPL (3R)-hydroxymyristoyl-[acyl carrier protein] dehydratase ((3R)-hydroxymyristoyl ACP dehydrase) emb|CAD64100.11 (3R)-hydroxymyristoyl-[acyl carrier protein] dehydratase [Lactobacillus plantarum WCFS1] Length = 147
621.5 Best-BlastP=> >nrprot 62% Identities = 195/428 (45%), Positives = 269/428 (62%), Gaps = 17/428 (3%) ref|NP_214178.11 3-oxoacyl-[acyl-carrier protein] synthase II [Aquifex aeolicus] pir||B70448 3-oxoacyl-[acyl-carrier-protein] synthase (EC 2.3.1.41 ) II - Aquifex aeolicus gb|AAC07574.11 3-oxoacyl-[acyl-carrier-protein] synthase II [Aquifex aeolicus VF5] Length = 415
622.3 Best-BlastP=> >nrprot 69% Identities = 412/793 (51 %), Positives = 556/793 (70%), Gaps = 12/793 (1 %) ref|NP_820864.1 | penicillin-binding protein 1 A [Coxiella burnetii RSA 4g3] gb|AA091378.11 penicillin-binding protein 1 A [Coxiella burnetii RSA 493] Length = 793
623.2 Best-BlastP=> >nrprot 77% Identities = 253/370 (68%), Positives = 289/370 (78%) ref|ZP_00013240.11 COG0686: Alanine dehydrogenase [Rhodospirillum rubrum] Length = 372
627.1 Best-BlastP=> >nrprot 39% Identities = 110/214 (51 %), Positives = 138/214 (64%), Gaps = 14/214 (6%) ref |NP_612899.11 hypothetical protein [Stx2 converting bacteriophage I] dbj|BAB87868.1 | hypothetical protein [Stx2 converting bacteriophage I] Length = 678
628.3 Best-BlastP=> >nrprot No Hits found 63.1 Best-BlastP=> >nrprot No Hits found
630.4 Best-BlastP=> >nrprot 73% Identities = 577/1040 (55%), Positives = 773/1040 (74%), Gaps = 1/1040 (0%) ref|ZP_00055701.11 hypothetical protein [Magnetospirillum magnetotacticum] Length = 1059
632.2 Best-BlastP=> >nrprot 71 % Identities = 136/258 (52%), Positives = 189/258 (73%), Gaps = 1/258 (0%) ref|NP_760580.11 Dihydropteroate synthase [Vibrio vulnificus CMCP6] gb|AA010107.1 |AE016802_150 Dihydropteroate synthase [Vibrio vulnificus CMCP6] Length = 259
633.3 Best-BlastP=> >nrprot 75% Identities = 282/442 (63%), Positives = 344/442 (77%), Gaps = 1/442 (0%) ref|NP_406959.11 probable phosphoglucomutase/phosphomannomutase [Yersinia pestis] ref|NP_668021.11 mrsA protein [Yersinia pestis KIM] pir||AE0425 probable phosphoglucomutase/phosphomannomutase [imported] - Yersinia pestis (strain C092) emb|CAC92729.1 | probable phosphoglucomutase/phosphomannomutase [Yersinia pestis C092] gb|AAM84272.1 |AE013670_9 mrsA protein [Yersinia pestis KIM] Length = 446
634.4 Best-BlastP=> >nrprot 51 % Identities = 85/166 (51 %), Positives = 93/166 (56%) emb|CAB44711.11 hypothetical protein (P4(21 )n) [Mus musculus] Length = 400
635.5 Best-BlastP=> >nrprot 83% Identities = 278/398 (69%), Positives = 332/398 (83%), Gaps = 14/398 (3%) ref|NP_819191.1 | cell division protein FtsZ [Coxiella burnetii RSA 493] gb|AAO89705.11 cell division protein FtsZ [Coxiella burnetii RSA 493] Length = 386
636.2 Best-BlastP=> >nrprot 77% Identities = 198/287 (68%), Positives = 236/287 (82%) ref )ZP_00090126.11 COG0774: UDP-3-O-acyl-N- acetylglucosamine deacetylase [Azotobacter vinelandii] Length = 303
637.3 Best-BlastP=> >nrprot 62% Identities = 261/623 (41%), Positives = 396/623 (63%), Gaps = 5/623 (0%) ref |NP_616924.11 endothelin converting enzyme homolog PepO [Methanosarcina acetivorans str. C2A] gb|AAM05404.11 endothelin converting enzyme homolog PepO [Methanosarcina acetivorans str. C2A] Length = 665
64.3 Best-BlastP=> >nrprot No Hits found
640.2 Best-BlastP=> >nrprot 51 % Identities = 102/309 (33%), Positives = 157/309 (50%), Gaps = 8/309 (2%) ref|NP_103911.1 ) hypothetical protein [Mesorhizobium loti] dbj|BAB49697.1 | hypothetical protein [Mesorhizobium loti] Length = 345
641.3 Best-BlastP=> >nrprot 74% Identities = 416/663 (62%), Positives = 506/663 (76%), Gaps = 5/663 (0%) ref|NP_931190.11 2,4-dienoyl-CoA reductase [NADPH] (2,4-dienoyl coenzyme A reductase) [Photorhabdus luminescens subsp. laumondii TT01] emb|CAE16362.11 2,4-dienoyl-CoA reductase [NADPH] (2,4-dienoyl coenzyme A reductase) [Photorhabdus luminescens subsp. laumondii TT01] Length = 673 43*3 Best-BlastP=> >nrprot 59% Identities = 208/462 (45%), Positives = 287/462 (62%), Gaps = 10/462 (2%) ref|NP_284769.11 UDP-N- acetylmuramyl-tripeptide synthetase [Neisseria meningitidis Z2491] sp|Q9JSZ0|MURE_NEIMA UDP-N-acetylmuramoylalanyl-D-glutamate- 2,6-diaminopimelate ligase (UDP-N-acetylmuramyl-tripeptide synthetase) (Meso-diaminopimelate-adding enzyme) (UDP- MurNAc-tripeptide synthetase) pir||A81778 UDP-N-acetylmuramoylalanyl-D-glutamate-2,6-diamino-pimelate ligase (EC 6.3.2.13) NMA2071 [similarity] - Neisseria meningitidis (strain Z2491 serogroup A) emb|CAB85288.1 | UDP-N-acetylmuramyl-tripeptide synthetase [Neisseria meningitidis Z2491] Length = 492
644.2 Best-BlastP=> >nrprot 52% Identities = 113/344 (32%), Positives = 185/344 (53%), Gaps = 10/344 (2%) ref|NP_820791.11 erythronate-4- phosphate dehydrogenase, putative [Coxiella burnetii RSA 493] gb|AA091305.11 erythronate-4-phosphate dehydrogenase, putative [Coxiella burnetii RSA 493] Length = 375
645.4 Best-BlastP=> >nrprot 66% Identities = 107/219 (48%), Positives = 147/219 (67%), Gaps = 5/219 (2%) ref|NP_820725.11 membrane protein, putative [Coxiella burnetii RSA 493] gb|AA091239.11 membrane protein, putative [Coxiella burnetii RSA 493] Length = 230
647.5 Best-BlastP=> >nrprot 49% Identities = 55/179 (30%), Positives = 107/179 (59%), Gaps = 1/179 (0%) gb|AAQ75156.1 | Pap2 superfamily protein [Alvinella pompejana epibiont 7G3] Length = 202
65.1 Best-BlastP=> >nrprot No Hits found
650.3 Best-BlastP=> >nrprot 72% Identities = 211/372 (56%), Positives = 272/372 (73%), Gaps = 2/372 (0%) ref|ZP_00029745.11 COG2170: Uncharacterized conserved protein [Burkholderia fungorum] Length = 388
651.2 Best-BlastP=> >nrprot 73% Identities = 70/130 (53%), Positives = 98/130 (75%) ref|ZP_00079460.11 COG1832: Predicted CoA-binding protein [Geobacter metallireducens] Length = 136
652.1 Best-BlastP=> >nrprot 62% Identities = 144/267 (53%), Positives = 179/267 (67%), Gaps = 5/267 (1 %) ref|NP_404647.11 conserved hypothetical protein [Yersinia pestis] ref|NP_670446.1 | hypothetical protein [Yersinia pestis KIM] pir||AI0126 conserved hypothetical protein YPO1034 [imported] - Yersinia pestis (strain C092) emb|CAC89876.1 | conserved hypothetical protein [Yersinia pestis C092] gb|AAM86697.1 |AE013915_6 hypothetical protein [Yersinia pestis KIM] Length = 281
656.3 Best-BlastP=> >nrprot No Hits found
657.3 Best-BlastP=> >nrprot 6% Identities = 39/123 (31 %), Positives = 64/123 (52%), Gaps = 11/123 (8%) ref|NP_828913.11 serine/threonine-protein kinase [Chlamydophila caviae GPIC] gb|AAP04791.11 serine/threonine-protein kinase [Chlamydophila caviae GPIC] Length = 501
659.2 Best-BlastP=> >nrprot 31 % Identities = 56/209 (26%), Positives = 100/209 (47%), Gaps = 11/209 (5%) ref|NP_843168.11 conserved hypothetical
protein [Bacillus anthracis str. Ames] gb|AAP24654.11 conserved hypothetical protein [Bacillus anthracis str. Ames] Length = 323
660.1 Best-BlastP=> >nrprot 61 % Identities = 83/178 (46%), Positives = 117/178 (65%) ref|NP_794412.11 acetyltransferase, GNAT family [Pseudomonas syringae pv. tomato str. DC3000] gb|AAO58107.11 acetyltransferase, GNAT family [Pseudomonas syringae pv. tomato str. DC3000] Length = 189
661.3 Best-BlastP=> >nrprot 20% Identities = 51/148 (34%), Positives = 76/148 (51%), Gaps = 10/148 (6%) ref|ZP_00109305.11 COG1670: Acetyltransferases, including N-acetylases of ribosomal proteins [Nostoc punctiforme] Length = 191
663.2 Best-BlastP=> >nrprot 49% Identities = 193/671 (28%), Positives = 325/671 (48%), Gaps = 62/671 (9%) gb|AAL77346.1 |AF443847_2 putative O- acetyltransferase WavN [Vibrio cholerae] Length = 663
665.2 Best-BlastP=> >nrprot 69% Identities = 43/78 (55%), Positives = 62/78 (79%) ref|NP_273842.11 conserved hypothetical protein [Neisseria meningitidis MC58] pir||G81157 conserved hypothetical protein NMB0800 [imported] - Neisseria meningitidis (strain MC58 serogroup B) gb|AAF41213.11 conserved hypothetical protein [Neisseria meningitidis MC58] Length = 94
666.2 Best-BlastP=> >nrprot 76% Identities = 120/204 (58%), Positives = 158/204 (77%) ref|ZP_00126281.11 COG0293: 23S rRNA methylase [Pseudomonas syringae pv. syringae B728a] Length = 216
667.3 Best-BlastP=> >nrprot 32% Identities = 68/178 (38%), Positives = 71/178 (39%) pir||A40215 TcD antigen - Trypanosoma cruzi Length = 207
668.4 Best-BlastP=> >nrprot 99% Identities = 1026/1048 (97%), Positives = 1040/1048 (99%) pir||T18334 icmE protein - Legionella pneumophila emb|CAA75331.11 IcmE protein [Legionella pneumophila] Length = 1048
670.1 Best-BlastP=> >nrprot 66% Identities = 213/471 (45%), Positives = 308/471 (65%), Gaps = 18/471 (3%) ref |ZP_00107537.11 COG1249: Pyruvate/2-oxoglutarate dehydrogenase complex, dihydrolipoamide dehydrogenase (E3) component, and related enzymes [Nostoc punctiforme] Length = 472
673.3 Best-BlastP=> >nrprot 42% Identities = 123/548 (22%), Positives = 218/548 (39%), Gaps = 102/548 (18%) ref|NP_458664.11 putative membrane protein [Salmonella enterica subsp. enterica serovar Typhi] ref|NP_807874.11 putative membrane protein [Salmonella enterica subsp. enterica serovar Typhi Ty2] pir||AI1031 probable membrane protein STY4579 [imported] - Salmonella enterica subsp. enterica serovar Typhi (strain CT18) emb|CAD09354.11 putative membrane protein [Salmonella enterica subsp. enterica serovar Typhi] gb|AA071734.11 putative membrane protein [Salmonella enterica subsp. enterica serovar Typhi Ty2] Length = 501
675.6 Best-BlastP=> >nrprot No Hits found
677.6 Best-BlastP=> >nrprot 58% Identities = 258/683 (37%), Positives = 389/683 (56%), Gaps = 39/683 (5%) gb|AAN62290.1 |AF440524_77 conserved hypothetical protein [Pseudomonas aeruginosa] Length = 724
678.3 Best-BlastP=> >nrprot No Hits found
680.3 Best-BlastP=> >nrprot 53% Identities = 57/132 (43%), Positives = 83/132 (62%), Gaps = 2/132 (1 %) ref|NP_283256.11 patch repair protein [Neisseria meningitidis Z2491] sp|Q9JWD6|VSR_NEIMA Putative very-short-patch-repair endonuclease pir||H81959 patch repair protein (EC 3.1.- .-) NMA0429 [imported] - Neisseria meningitidis (strain Z2491 serogroup A) emb|CAB83728.11 patch repair protein [Neisseria meningitidis Z2491] Length = 140
681.3 Best-BlastP=> >nrprot No Hits found
682.2 Best-BlastP=> >nrprot 76% Identities = 191/295 (64%), Positives = 229/295 (77%), Gaps = 7/295 (2%) sp|Q59603|MTB1_NEIGO Modification methylase NgoBI (Cytosine-specific methyltransferase NgoBI) (M.NgoBI) (M.Ngol) gb|AAB03206.2| cytosine DNA methylase M.Ngol [Neisseria gonorrhoeae] Length = 317
683.2 Best-BlastP=> >nrprot 64% Identities = 200/443 (45%), Positives = 289/443 (65%), Gaps = 11/443 (2%) ref|NP_819174.11 UDP-N- acetylmuramoylalanyl-D-glutamyl-2, 6-diaminopimelate-D-alanyl-D-alanyl ligase [Coxiella burnetii RSA 493] gb|AA089688.1 | UDP- N-acetylmuramoylalanyl-D-glutamyl-2, 6-diaminopimelate-D-alanyl-D-alanyl ligase [Coxiella burnetii RSA 493] Length = 446
684.5 Best-BlastP=> >nrprot 48% Identities = 73/253 (28%), Positives = 127/253 (50%), Gaps = 40/253 (15%) ref|NP_819573.11 cell division protein ZipA, putative [Coxiella burnetii RSA 493] gb|AAO90087.11 cell division protein ZipA, putative [Coxiella burnetii RSA 493] Length = 225
687.5 Best-BlastP=> >nrprot 47% Identities = 120/426 (28%), Positives = 204/426 (47%), Gaps = 29/426 (6%) ref|ZP_00082430.11 COG0304: 3- oxoacyl-(acyl-carrier-protein) synthase [Geobacter metallireducens] Length = 410
688.1 Best-BlastP=> >nrprot 48% Identities = 71/266 (26%), Positives = 138/266 (51 %), Gaps = 11/266 (4%) ref|NP_908017.11 LIPID A BIOSYNTHESIS ACYLTRANSFERASE [Wolinella succinogenes] emb|CAE10917.11 LIPID A BIOSYNTHESIS ACYLTRANSFERASE [Wolinella succinogenes] Length = 300
692.1 Best-BlastP=> >nrprot No Hits found
693.1 Best-BlastP=> >nrprot 71 % Identities = 96/172 (55%), Positives = 128/172 (74%) ref|NP_299266.11 conserved hypothetical protein [Xylella fastidiosa 9a5c] pir||C82613 conserved hypothetical protein XF1984 [imported] - Xylella fastidiosa (strain 9a5c) gb|AAF84786.1 |AE004018_2 conserved hypothetical protein [Xylella fastidiosa 9a5c] Length = 187
694.3 Best-BlastP=> >nrprot 72% Identities = 239/453 (52%), Positives = 330/453 (72%), Gaps = 9/453 (1 %) dbj|BAB33285.11 glutathione reductase [Acinetobacter sp. M-1] Length = 450
696.2 Best-BlastP=> >nrprot 74% Identities = 92/143 (64%), Positives = 107/143 (74%), Gaps = 13/143 (9%) ref|NP_230222.11 ribosomal protein S9 [Vibrio cholerae 01 biovar eltor str. N16961] sp|Q9KUF0|RS9_VIBCH 30S ribosomal protein S9 pir||C82308 ribosomal protein S9 VC0571 [imported] - Vibrio cholerae (strain N16961 serogroup 01 ) gb|AAF93739.1 | ribosomal protein S9 [Vibrio cholerae 01 biovar eltor str. N16961] Length = 130
697.1 Best-BlastP=> >nrprot 74% Identities = 111/188 (59%), Positives = 142/188 (75%), Gaps = 3/188 (1 %) ref|NP_840883.1 | Rieske iron-sulfur protein 2Fe-2S subunit [Nitrosomonas europaea ATCC 19718] emb|CAD84720.11 Rieske iron-sulfur protein 2Fe-2S subunit [Nitrosomonas europaea ATCC 19718] Length = 201
698.2 Best-BlastP=> >nrprot 81 % Identities = 273/401 (68%), Positives = 332/401 (82%) ref|NP_743478.11 ubiquinol-cytochrome c reductase, cytochrome b [Pseudomonas putida KT2440] gb|AAN66942.1 |AE016322_9 ubiquinol-cytochrome c reductase, cytochrome b [Pseudomonas putida KT2440] Length = 403
699.3 Best-BlastP=> >nrprot 59% Identities = 157/374 (41 %), Positives = 226/374 (60%), Gaps = 4/374 (1 %) ref|ZP_00089161.11 COG0607: Rhodanese-related sulfurtransferase [Azotobacter vinelandii] Length = 377
7.1 Best-BlastP=> >nrprot 43% Identities = 394/1345 (29%), Positives = 596/1345 (44%), Gaps = 225/1345 (16%) ref|NP_907747.11 hypothetical protein WS1613 [Wolinella succinogenes] emb|CAE10647.1 | hypothetical protein [Wolinella succinogenes] Length = 1409
701.2 Best-BlastP=> >nrprot 62% Identities = 231/482 (47%), Positives = 307/482 (63%), Gaps = 6/482 (1 %) ref |NP_819376.11 phosphomethylpyrimidine kinase/thiamin-phosphate pyrophosphorylase [Coxiella burnetii RSA 493] gb|AAO89890.1 | phosphomethylpy midine kinase/thiamin-phosphate pyrophosphorylase [Coxiella burnetii RSA 493] Length = 479
702.2 Best-BlastP=> >nrprot 73% Identities = 153/259 (59%), Positives = 193/259 (74%) ref|ZP_00067659.11 COG2022: Uncharacterized enzyme of thiazole biosynthesis [Microbulbifer degradans 2-40] Length = 269
704.4 Best-BlastP=> >nrprot No Hits found
705.1 Best-BlastP=> >nrprot 60% Identities = 179/427 (41 %), Positives = 263/427 (61 %), Gaps = 17/427 (3%) ref|NP_691145.11 hydroxymethylglutaryl- CoA reductase [Oceanobacillus iheyensis HTE831] dbj|BAC12180.11 hydroxymethylglutaryl-CoA reductase [Oceanobacillus iheyensis HTE831] Length = 429
706.2 Best-BlastP=> >nrprot 27% Identities = 40/115 (34%), Positives = 49/115 (42%), Gaps = 1/115 (0%) pir||S75426 hypothetical protein c05009 - Sulfolobus solfataricus emb|CAA69540.11 orf c05009 [Sulfolobus solfataricus] Length = 115
707.2 Best-BlastP=> >nrprot 57% Identities = 141/338 (41 %), Positives = 196/338 (57%), Gaps = 4/338 (1 %) ref|ZP_00072976.11 COG1304: L-lactate dehydrogenase (FMN-dependent) and related alpha-hydroxy acid dehydrogenases [Trichodesmium erythraeum IMS101] Length = 345
709.1 Best-BlastP=> >nrprot No Hits found
711.2 Best-BlastP=> >nrprot 74% Identities = 396/663 (59%), Positives = 497/663 (74%), Gaps = 2/663 (0%) ref|NP_249239.11 transketolase [Pseudomonas aeruginosa PA01] pir||B83577 transketolase PA0548 [imported] - Pseudomonas aeruginosa (strain PA01) gb|AAG03937.1 |AE004491_4 transketolase [Pseudomonas aeruginosa PA01] Length = 665
712.2 Best-BlastP=> >nrprot 79% Identities = 212/329 (64%), Positives = 263/329 (79%), Gaps = 5/329 (1 %) ref|ZP_00033578.11 hypothetical protein [Burkholderia fungorum] Length = 336
713.2 Best-BlastP=> >nrprot No Hits found 715.1 Best-BlastP=> >nrprot No Hits found
716.1 Best-BlastP=> >nrprot No Hits found
717.2 Best-BlastP=> >nrprot No Hits found 719.4 Best-BlastP=> >nrprot 75% Identities = 484/781 (61 %), Positives = 592/781 (75%), Gaps = 24/781 (3%) ref|NP_901207.11 probable transcriptional accessory protein [Chromobacterium violaceum ATCC 12472] gb|AAQ59212.11 probable transcriptional accessory protein [Chromobacterium violaceum ATCC 12472] Length = 770
722.3 Best-BlastP=> >nrprot 58% Identities = 79/215 (36%), Positives = 130/215 (60%), Gaps = 3/215 (1 %) ref|NP_716217.1| thiopurine S- methyltransferase [Shewanella oneidensis MR-1] gb|AAN53662.1 |AE015505_5 thiopurine S-methyltransferase [Shewanella oneidensis MR-1] Length = 218
723.3 Best-BlastP=> >nrprot 75% Identities = 353/611 (57%), Positives = 454/611 (74%), Gaps = 7/611 (1 %) reflNP_820767.11 glucosamine-fructose- 6-phosphate aminotransferase, isomerizing [Coxiella burnetii RSA 493] gb|AA091281.1 | glucosamine-fructose-6-phosphate aminotransferase, isomerizing [Coxiella burnetii RSA 493] Length = 611
727.3 Best-BlastP=> >nrprot 82% Identities = 756/1070 (70%), Positives = 881/1070 (82%), Gaps = 5/1070 (0%) ref|NP_794254.11 carbamoyl- phosphate synthase, large subunit [Pseudomonas syringae pv. tomato str. DC3000] gb|AA057949.11 carbamoyl-phosphate synthase, large subunit [Pseudomonas syringae pv. tomato str. DC3000] Length = 1073
728.2 Best-BlastP=> >nrprot 53% Identities = 49/111 (44%), Positives = 72/111 (64%) ref|NP_799407.11 CrcB protein [Vibrio parahaemolyticus RIMD 2210633] sp|Q87KE9|CRCB_VIBPA Protein crcB homolog dbj|BAC61291.11 CrcB protein [Vibrio parahaemolyticus] Length = 127
729.1 Best-BlastP=> >nrprot 73% Identities = 135/256 (52%), Positives = 190/256 (74%) ref|NP_252334.11 UDP-N-acetylglucosamine acyltransferase [Pseudomonas aeruginosa PA01] sp|Q9X6P4|LPXA_PSEAE Acyl-[acyl-carrier-protein]~UDP-N-acetylglucosamine O- acyltransferase (UDP-N-acetylglucosamine acyltransferase) pir||D83190 UDP-N-acetylglucosamine acyltransferase PA3644 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAD30149.1 |AF142597_1 hydroxydecanoyl-acyl carrier protein-dependent UDP-N- acetylglucosamine-3-O-acyltransferase [Pseudomonas aeruginosa] gb|AAG07032.1 |AE004784_5 UDP-N-acetylglucosamine acyltransferase [Pseudomonas aeruginosa PA01] Length = 258
730.1 Best-BlastP=> >nrprot 68% Identities = 79/142 (55%), Positives = 104/142 (73%), Gaps = 1/142 (0%) ref|NP_873649.11 (3R)-hydroxymyristoyl- acyl carrier protein dehydratase [Haemophilus ducreyi 35000HP] gb|AAPg6038.1 | (3R)-hydroxymyristoyl-acyl carrier protein dehydratase [Haemophilus ducreyi 35000HP] Length = 154
731.2 Best-B!astP=> >nrprot 57% Identities = 132/334 (39%), Positives = 193/334 (57%) ref|NP_717250.11 UDP-3-0-(3-hydroxymyristoyl) glucosamine n-acyltransferase [Shewanella oneidensis MR-1] gb|AAN54694.1 |AE015609_13 UDP-3-0-(3-hydroxymyristoyl) glucosamine n- acyltransferase [Shewanella oneidensis MR-1] Length = 341
732.3 Best-BlastP=> >nrprot 53% Identities = 50/139 (35%), Positives = 89/139 (64%) ref |NP_819642.11 outer membrane protein OmpH, putative [Coxiella burnetii RSA 493] gb|AAO90156.11 outer membrane protein OmpH, putative [Coxiella burnetii RSA 493] Length = 165
737.1 Best-BlastP=> >nrprot 45% Identities = 101/299 (33%), Positives = 156/299 (52%), Gaps = 15/299 (5%) ref|NP_812239.11 ATP-dependent DNA helicase recG [Bacteroides thetaiotaomicron VPI-5482] gb|AA078433.1 1 ATP-dependent DNA helicase recG [Bacteroides thetaiotaomicron VPI-5482] Length = 485
738.2 Best-BlastP=> >nrprot 46% Identities = 48/153 (31 %), Positives = 81/153 (52%), Gaps = 2/153 (1 %) ref|NP_617319.1 | ATP-dependent DNA helicase [Methanosarcina acetivorans str. C2A] gb|AAM05799.11 ATP-dependent DNA helicase [Methanosarcina acetivorans str. C2A] Length = 514
739.4 Best-BlastP=> >nrprot 69% Identities = 146/295 (49%), Positives = 202/295 (68%), Gaps = 10/295 (3%) ref|NP_052845.11 hypothetical protein [Coxiella burnetii] ref|NP_819051.11 parB protein, putative [Coxiella burnetii RSA 493] gb|AAD33477.1 |AF131076_3 hypothetical protein [Coxiella burnetii] gb|AA091611.11 parB protein, putative [Coxiella burnetii RSA 493] Length = 289
740.2 Best-BlastP=> >nrprot 81 % Identities = 72/114 (63%), Positives = 93/114 (81%) ref|NP_772410.1 | bll5770 [Bradyrhizobium japonicum] dbj|BAC51035.1 | bll5770 [Bradyrhizobium japonicum USDA 110] Length = 118
741.2 Best-BlastP=> >nrprot No Hits found
742.3 Best-BlastP=> >nrprot 76% Identities = 280/467 (59%), Positives = 357/467 (76%), Gaps = 3/467 (0%) ref|NP_820336.11 amino acid antiporter [Coxiella burnetii RSA 493] gb|AAO90850.11 amino acid antiporter [Coxiella burnetii RSA 493] Length = 474
743.3 Best-BlastP=> >nrprot 94% Identities = 124/131 (94%), Positives = 125/131 (95%) gb|AAM00634.1 | unknown [Legionella pneumophila] Length = 131
744.3 Best-BlastP=> >nrprot 97% Identities = 163/168 (97%), Positives = 164/168 (97%) gb|AAM00635.11 putative cadmium efflux ATPase [Legionella pneumophila] Length = 635
747.3 Best-BlastP=> >nrprot 99% Identities = 723/729 (99%), Positives = 725/729 (99%) gb|AAM00636.11 unknown [Legionella pneumophila] Length = 729 748.3 Best-BlastP=> >nrprot 30% Identities = 99/321 (30%), Positives = 137/321 (42%), Gaps = 72/321 (22%) ref|ZP_00129242.11 COG2821 : Membrane-bound lytic murein transglycosylase [Desulfovibrio desulfuricans G20] Length = 442
749.1 Best-BlastP=> >nrprot No Hits found
750.2 Best-BlastP=> >nrprot No Hits found 751.2 Best-BlastP=> >nrprot No Hits found 752.2 Best-BlastP=> >nrprot No Hits found
753.2 Best-BlastP=> >nrprot 51 % Identities = 92/311 (29%), Positives = 164/311 (52%), Gaps = 28/311 (9%) ref|NP_782216.11 conserved protein [Clostridium tetani E88] gb|AA036153.11 conserved protein [Clostridium tetani E88] Length = 307
754.2 Best-BlastP=> >nrprot 62% Identities = 160/323 (49%), Positives = 209/323 (64%), Gaps = 6/323 (1 %) ref|ZP_00107451.11 COG0667: Predicted oxidoreductases (related to aryl-alcohol dehydrogenases) [Nostoc punctiforme] Length = 326
755.3 Best-BlastP=> >nrprot 46% Identities = 72/227 (31 %), Positives = 126/227 (55%), Gaps = 12/227 (5%) ref|ZP_00065015.1 | COG0062: Uncharacterized conserved protein [Microbulbifer degradans 2-40] Length = 500
757.1 Best-BlastP=> >nrprot 64% Identities = 95/203 (46%), Positives = 132/203 (65%), Gaps = 9/203 (4%) ref|NP_820677.11 4Fe-4S binding domain protein [Coxiella burnetii RSA 4 3] gb|AA091191.11 4Fe-4S binding domain protein [Coxiella burnetii RSA 493] Length = 213
758.1 Best-BlastP=> >nrprot 82% Identities = 155/208 (74%), Positives = 174/208 (83%) ref|ZP_00122593.11 COG0177: Predicted Endolll-related endonuclease [Haemophilus somnus 129PT] Length = 211
760.2 Best-BlastP=> >nrprot No Hits found
761.2 Best-BlastP=> >nrprot 22% Identities = 41/124 (33%), Positives = 70/124 (56%), Gaps = 8/124 (6%) ref|NP_656804.11 Acetyltransf, Acetyltransferase (GNAT) family [Bacillus anthracis A2012] ref|NP_845271.11 acetyltransferase, GNAT family [Bacillus anthracis str. Ames] gb|AAP26757.11 acetyltransferase, GNAT family [Bacillus anthracis str. Ames] Length = 135
762.2 Best-BlastP=> >nrprot 56% Identities = 80/200 (40%), Positives = 123/200 (61 %), Gaps = 1/200 (0%) ref|ZP_00086065.11 COG1280: Putative threonine efflux protein [Pseudomonas fluorescens PfO-1] Length = 209
764.1 Best-BlastP=> >nrprot 52% Identities = 53/167 (31 %), Positives = 90/167 (53%), Gaps = 4/167 (2%) ref|NP_490158.11 probable acetyltransferase [Nostoc sp. PCC 7120] pir||AD2484 hypothetical protein alr7052 [imported] - Nostoc sp. (strain PCC 7120) plasmid pCC7120alpha dbj|BAB78136.11 ORF_ID:alr7052~probable acetyltransferase [Nostoc sp. PCC 7120] Length = 169
765.1 Best-BlastP=> >nrprot 61 % Identities = 99/234 (42%), Positives = 148/234 (63%), Gaps = 8/234 (3%) ref|NP_217318.1 | hypothetical protein Rv2802c [Mycobacterium tuberculosis H37Rv] ref|NP_856471.1 | HYPOTHETICAL ARGININE AND ALANINE RICH PROTEIN [Mycobacterium bovis subsp. bovis AF2122/97] pir|lE70689 hypothetical protein Rv2802c - Mycobacterium tuberculosis (strain H37RV) emb|CAB03674.1 | hypothetical protein Rv2802c [Mycobacterium tuberculosis H37Rv] emb|CAD95010.1 | HYPOTHETICAL ARGININE AND ALANINE RICH PROTEIN [Mycobacterium bovis subsp. bovis AF2122/97] Length = 347
766.3 Best-BlastP=> >nrprot 36% Identities = 99/398 (24%), Positives = 173/398 (43%), Gaps = 71/398 (17%) ref|NP_845051.11 esterase, putative [Bacillus anthracis str. Ames] gb|AAP26537.1 | esterase, putative [Bacillus anthracis str. Ames] Length = 382
767.3 Best-BlastP=> >nrprot 64% Identities = 206/430 (47%), Positives = 274/430 (63%), Gaps = 8/430 (1%) ref|NP_248707.11 conserved hypothetical protein [Pseudomonas aeruginosa PA01] pir||F83643 conserved hypothetical protein PA0017 [imported] - Pseudomonas aeruginosa (strain PA01) gb|AAG03407.1 |AE004441_8 conserved hypothetical protein [Pseudomonas aeruginosa PA01] Length = 434
769.2 Best-BlastP=> >nrprot 71 % Identities = 167/311 (53%), Positives = 225/311 (72%), Gaps = 4/311 (1 %) ref|ZP_00082909.11 COG0223: Methionyl- tRNA formyltransferase [Pseudomonas fluorescens PfO-1] Length = 319
77.1 Best-BlastP=> >nrprot No Hits found
770.3 Best-BlastP=> >nrprot 98% Identities = 157/161 (97%), Positives = 160/161 (99%) gb|AAD43221.1 |AF111940_3 LspH precursor [Legionella pneumophila] Length = 161
771.4 Best-BlastP=> >nrprot 99% Identities = 140/140 (100%), Positives = 140/140 (100%) gb| AAD43220.1 |AF111 g40_2 LspG precursor [Legionella pneumophila] gb|AAP69529.1 | type II protein secretion LspG pseudopilin [Legionella pneumophila] Length = 140
772.4 Best-BlastP=> >nrprot 99% Identities = 393/399 (98%), Positives = 396/399 (99%) gb|AAK35047.2|AF330137_1 type II protein secretion LspF [Legionella pneumophila] Length = 399
774.3 Best-BlastP=> >nrprot 82% Identities = 325/469 (69%), Positives = 386/469 (82%) ref |NP_719933.11 glutamine synthetase, type I [Shewanella oneidensis MR-1] gb|AAN57377.1 |AE015874_4 glutamine synthetase, type I [Shewanella oneidensis MR-1] Length = 469
775.2 Best-BlastP=> >nrprot 51 % Identities = 62/171 (36%), Positives = 96/171 (56%), Gaps = 11/171 (6%) ref|NP_796499.11 hypothetical protein VP0120 [Vibrio parahaemolyticus RIMD 2210633] dbj|BAC58383.11 hypothetical protein [Vibrio parahaemolyticus] Length = 193
776.3 Best-BlastP=> >nrprot 99% Identities = 497/502 (99%), Positives = 500/502 (99%), Gaps = 1/502 (0%) gb|AAF34822.1 |AF167992_2 IraB [Legionella pneumophila] Length = 501
778.2 Best-BlastP=> >nrprot 99% Identities = 272/272 (100%), Positives = 272/272 (100%) emb|CAB65217.11 hypothetical protein [Legionella pneumophila] gb|AAF34821.1 |AF167992_1 small-molecule methyltransferase IraA [Legionella pneumophila] Length = 272
779.4 Best-BlastP=> >nrprot 99% Identities = 459/460 (99%), Positives = 460/460 (100%) emb|CAB65216.11 hypothetical protein [Legionella pneumophila] Length = 460
783.2 Best-BlastP=> >nrprot 62% Identities = 51/105 (48%), Positives = 75/105 (71 %), Gaps = 1/105 (0%) ref|NP_359553.1 | Conserved hypothetical protein [Streptococcus pneumoniae R6] pir||G98116 conserved hypothetical protein spr1962 [imported] - Streptococcus pneumoniae (strain R6) gb|AAL00764.1 | Conserved hypothetical protein [Streptococcus pneumoniae R6] Length = 299
785.4 Best-BlastP=> >nrprot No Hits found
787.3 Best-BlastP=> >nrprot 68% Identities = 344/658 (52%), Positives = 452/658 (68%), Gaps = 23/658 (3%) ref|NP_718364.11 ribonuclease E [Shewanella oneidensis MR-1] gb|AAN55808.1 |AE015717_9 ribonuclease E [Shewanella oneidensis MR-1] Length = 1088
789.4 Best-BlastP=> >nrprot 51 % Identities = 25/77 (32%), Positives = 43/77 (55%) reflNP_085189.11 IS10 orf [Shigella flexneri] ref|NP_858160.1 | . hypothetical protein [Shigella flexneri 2a] gb|AAK18345.1 |AF348706_34 IS10 orf [Shigella flexneri] gb|AAL72480.11 hypothetical protein [Shigella flexneri 2a] Length = 407
'y π CMCP6] ref|NP_759894.1 | Conserved hypothetical protein [Vibrio vulnificus CMCP6] ref|NP_759900.1 | Conserved hypothetical protein [Vibrio vulnificus CMCP6] ref|NP_759939.1 | Conserved hypothetical protein [Vibrio vulnificus CMCP6] ref|NP_760021.1 ) Conserved hypothetical protein [Vibrio vulnificus CMCP6] ref|NP_760123.1 | Conserved hypothetical protein [Vibrio vulnificus CMCP6] ref|NP_760329.1 | Conserved hypothetical protein [Vibrio vulnificus CMCP6] ref|NP_760403.11 Conserved hypothetical protein [Vibrio vulnificus CMCP6] ref|NP_763327.1 | Conserved hypothetical protein [Vibrio vulnificus CMCP6] gb|AAO08317.1 |AE016813_69 Conserved hypothetical protein [Vibrio vulnificus CMCP6] gb|AAO08998.1 |AE016798_158 Conserved hypothetical protein [Vibrio vulnificus CMCP6] gb|AAO09421.1 |AE016800_26 Conserved hypothetical protein [Vibrio vulnificus CMCP6] gb|AAO09427.1 |AE016800_32 Conserved hypothetical protein [Vibrio vulnificus CMCP6] gb|AAO09466.1 |AE016800_71 Conserved hypothetical protein [Vibrio vulnificus CMCP6] gb|AAO09548.1 |AE016800_153 Conserved hypothetical
790.3 Best-BlastP=> >nrprot No Hits found
791.5 Best-BlastP=> >nrprot 98% Identities = 367/375 (97%), Positives = 372/375 (99%) gb|AAM00643.11 cyclopropane fatty acyl phospholipid synthase [Legionella pneumophila] Length = 375
793.2 Best-BlastP=> >nrprot No Hits found 794.2 Best-BlastP=> >nrprot No Hits found 795.2 Best-BlastP=> >nrprot 69% Identities = 134/260 (51 %), Positives = 181/260 (69%), Gaps = 5/260 (1 %) ref|NP_635864.11 indole-3-glycerol phosphate synthase [Xanthomonas campestris pv. campestris str. ATCC 33913] sp|Q8PD70|TRPC_XANCP lndole-3-glycerol phosphate synthase (IGPS) gb|AAM39788.1 | indole-3-glycerol phosphate synthase [Xanthomonas campestris pv. campestris str. ATCC 33913] Length = 265 796.2 Best-BlastP=> >nrprot 39% Identities = 60/227 (26%), Positives = 98/227 (43%), Gaps = 25/227 (11 %) reflNP_783943.11 adenylyl transferase (putative) [Lactobacillus plantarum WCFS1] emb|CAD62779.11 adenylyl transferase (putative) [Lactobacillus plantarum WCFS1] Length = 259
798.2 Best-BlastP=> >nrprot 60% Identities = 242/575 (42%), Positives = 364/575 (63%), Gaps = 8/575 (1 %) ref|NP_359923.11 multidrug resistance ABC transporter ATP-binding protein [Rickettsia conorii] pir||F97735 hypothetical protein abcT3 [imported] - Rickettsia conorii (strain Malish 7) gb|AAL02824.11 multidrug resistance ABC transporter ATP-binding protein [Rickettsia conorii] Length = 589
799.3 Best-BlastP=> >nrprot 50% Identities = 156/500 (31 %), Positives = 249/500 (49%), Gaps = 60/500 (12%) ref|ZP_00112433.11 COG0488: ATPase components of ABC transporters with duplicated ATPase domains [Nostoc punctiforme] Length = 544
80.3 Best-BlastP=> >nrprot 77% Identities = 34/42 (80%), Positives = 34/42 (80%) ref|NP_759472.11 Conserved hypothetical protein [Vibrio vulnificus CMCP6] ref|NP_759895.11 Conserved hypothetical protein [Vibrio vulnificus CMCP6] ref|NP_759901.1 | Conserved hypothetical protein [Vibrio vulnificus CMCP6] ref|NP_760122.11 Conserved hypothetical protein [Vibrio vulnificus CMCP6] ref|NP_760328.11 Conserved hypothetical protein [Vibrio vulnificus CMCP6] ref|NP_763326.11 Conserved hypothetical protein [Vibrio vulnificus CMCP6] gb|AAO08316.1 |AE016813_68 Conserved hypothetical protein [Vibrio vulnificus CMCP6] gb|AAO08999.1 |AE016798_159 Conserved hypothetical protein [Vibrio vulnificus CMCP6] gb|AAO09422.1 |AE016800_27 Conserved hypothetical protein [Vibrio vulnificus CMCP6] gb|AAO09428.1 |AE016800_33 Conserved hypothetical protein [Vibrio vulnificus CMCP6] gb|AAO09649.1 |AE016800_254 Conserved hypothetical protein [Vibrio vulnificus CMCP6] gb|AAO09855.1 |AE016801_174 Conserved hypothetical protein [Vibrio vulnificus CMCP6] Length = 43
801.3 Best-BlastP=> >nrprot 61 % Identities = 104/249 (41 %), Positives = 152/249 (61 %), Gaps = 3/249 (1 %) ref|ZP_00032174.11 COG4787: Flagellar basal body rod protein [Burkholderia fungorum] Length = 461
802.3 Best-BlastP=> >nrprot 74% Identities = 158/261 (60%), Positives = 196/261 (75%) ref|NP_249773.11 flagellar basal-body rod protein FlgG [Pseudomonas aeruginosa PA01] ref|ZP_00138668.11 COG4786: Flagellar basal body rod protein [Pseudomonas aeruginosa UCBPP- PA14] pir||H83510 flagellar basal-body rod protein FlgG PA1082 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG04471.1 |AE004539_13 flagellar basal-body rod protein FlgG [Pseudomonas aeruginosa PA01] Length = 261
803.3 Best-BlastP=> >nrprot 58% Identities = 84/223 (37%), Positives = 136/223 (60%), Gaps = 3/223 (1 %) gb|AAN08643.11 FlgH [Aeromonas hydrophila] Length = 223
806.2 Best-BlastP=> >nrprot 41 % Identities = 54/206 (26%), Positives = 102/206 (49%), Gaps = 11/206 (5%) sp|Q00924lHYUE_PSESN HYDANTOIN RACEMASE pir||B41895 hydantoin racemase HyuE - Pseudomonas sp gb|AAA25843.1 | 5-substituted hydantoin racemase Length = 249
807.2 Best-BlastP=> >nrprot 46% Identities = 135/390 (34%), Positives = 205/390 (52%), Gaps = 14/390 (3%) ref|ZP_00090340.11 COG0402: Cytosine deaminase and related metal-dependent hydrolases [Azotobacter vinelandii] Length = 463
808.4 Best-BlastP=> >nrprot 20% Identities = 110/492 (22%), Positives = 193/492 (39%), Gaps = 57/492 (11 %) ref|NP_213438.11 chromosome assembly protein homolog [Aquifex aeolicus] pir||B70356 chromosome assembly protein homolog - Aquifex aeolicus gb|AAC06839.11 chromosome assembly protein homolog [Aquifex aeolicus VF5] Length = 1156
81.3 Best-BlastP=> >nrprot 52% Identities = 20/40 (50%), Positives = 24/40 (60%) ref|NP_907600.11 hypothetical protein WS1440 [Wolinella succinogenes] emb|CAE10500.11 hypothetical protein [Wolinella succinogenes] Length = 42
813.2 Best-BlastP=> >nrprot 60% Identities = 178/412 (43%), Positives = 259/412 (62%), Gaps = 8/412 (1 %) ref|NP_821038.11 major facilitator family transporter [Coxiella burnetii RSA 493] gb|AA091552.11 major facilitator family transporter [Coxiella burnetii RSA 493] Length = 446
815.2 Best-BlastP=> >nrprot 75% Identities = 251/408 (61%), Positives = 306/408 (75%), Gaps = 7/408 (1 %) ref|NP_671036.1 | phosphopentomutase [Yersinia pestis KIM] gb|AAM87287.1 |AE013977_6 phosphopentomutase [Yersinia pestis KIM] Length = 429
817.7 Best-BlastP=> >nrprot 21% Identities = 198/995 (19%), Positives = 410/995 (41 %), Gaps = 142/995 (14%) dbj|BAA34954.1 | myosin heavy chain [Dugesia japonica] Length = 1958
rt co **^* σi m I
s-! cd co co co TO TO TO TO TO TO TO TO eo TO
84.4 Best-BlastP=> >nrprot 59% Identities = 243/551 (44%), Positives = 334/551 (60%), Gaps = 49/551 (8%) ref|NP_819689.11 DNA polymerase III, gamma and tau subunits [Coxiella burnetii RSA 493] gb|AAO90203.1 | DNA polymerase III, gamma and tau subunits [Coxiella burnetii RSA 493] Length = 509
840.2 Best-BlastP=> >nrprot 63% Identities = 124/242 (51 %), Positives = 155/242 (64%) reflNP_404551.11 conserved hypothetical protein [Yersinia pestis] ref|NP_670619.1 | hypothetical protein [Yersinia pestis KIM] pir||AF0114 conserved hypothetical protein YPO0934 [imported] - Yersinia pestis (strain C092) emb|CAC89777.1 | conserved hypothetical protein [Yersinia pestis C092] gb|AAM86870.1 |AE013933_7 hypothetical protein [Yersinia pestis KIM] Length = 243
841.3 Best-BlastP=> >nrprot 78% Identities = 70/108 (64%), Positives = 86/108 (79%) ref|ZP_00023150.11 COG0526: Thiol-disulfide isomerase and thioredoxins [Ralstonia metallidurans] Length = 108
843.2 Best-BlastP=> >nrprot 99% Identities = 188/189 (99%), Positives = 189/189 (100%) emb|CAD90948.11 putative pyrimidine phosphoribosyl transferase [Legionella pneumophila] Length = 189
844.2 Best-BlastP=> >nrprot 70% Identities = 61/101 (60%), Positives = 77/101 (76%), Gaps = 2/101 (1%) ref|ZP_00068029.11 COG4517: Uncharacterized protein conserved in bacteria [Microbulbifer degradans 2-40] Length = 189
845.4 Best-BlastP=> >nrprot 38% Identities = 54/212 (25%), Positives = 97/212 (45%), Gaps = 6/212 (2%) ref|NP_819358.11 conserved hypothetical protein [Coxiella burnetii RSA 493] gb|AA089872.1 | conserved hypothetical protein [Coxiella burnetii RSA 493] Length = 243
847.4 Best-BlastP=> >nrprot 68% Identities = 562/1047 (53%), Positives = 719/1047 (68%), Gaps = 18/1047 (1 %) ref|NP_639180.1 | bifunctional PutA protein [Xanthomonas campestris pv. campestris str. ATCC 33913] gb|AAM43071.1 | bifunctional PutA protein [Xanthomonas campestris pv. campestris str. ATCC 33913] Length = 1066
852.3 Best-BlastP=> >nrprot 58% Identities = 92/194 (47%), Positives = 116/194 (59%) ref|NP_642158.11 amidotransferase [Xanthomonas axonopodis pv. citri str. 306] gb|AAM36694.11 amidotransferase [Xanthomonas axonopodis pv. citri str. 306] Length = 200
853.1 Best-BlastP=> >nrprot 62% Identities = 99/239 (41 %), Positives = 149/239 (62%), Gaps = 2/239 (0%) ref|NP_928857.11 phosphoribosylformimino-5-aminoimidazole carboxamide ribotide isomerase [Photorhabdus luminescens subsp. laumondii TT01] emb|CAE13859.11 phosphoribosylformimino-5-aminoimidazole carboxamide ribotide isomerase [Photorhabdus luminescens subsp. laumondii TT01] Length = 245
854.3 Best-BlastP=> >nrprot 62% Identities = 97/179 (54%), Positives = 132/179 (73%), Gaps = 2/179 (1 %) ref|NP_797522.11 imidazoleglycerol- phosphate synthase, cyclase subunit [Vibrio parahaemolyticus RIMD 2210633] sp|Q87QK6|HIS6_VIBPA Imidazole glycerol phosphate synthase subunit hisF (IGP synthase cyclase subunit) (IGP synthase subunit hisF) (ImGP synthase subunit hisF) (IGPS subunit hisF) dbj|BAC59406.11 imidazoleglycerol-phosphate synthase, cyclase subunit [Vibrio parahaemolyticus] Length = 257
856.2 Best-BlastP=> >nrprot No Hits found 857.2 Best-BlastP=> >nrprot 64% Identities = 349/779 (44%), Positives = 498/779 (63%), Gaps = 20/779 (2%) ref|NP_668468.11 putative penicillin- binding protein [Yersinia pestis KIM] gb|AAM84719.1 |AE013717_1 putative penicillin-binding protein [Yersinia pestis KIM] Length = 845
858.2 Best-BlastP=> >nrprot 60% Identities = 71/157 (45%), Positives = 99/157 (63%) ref|ZP_00089699.1 | COG1376: Uncharacterized protein conserved in bacteria [Azotobacter vinelandii] Length = 174
859.2 Best-BlastP=> >nrprot 69% Identities = 240/454 (52%), Positives = 324/454 (71 %) ref|NP_820199.11 aldehyde dehydrogenase family protein [Coxiella burnetii RSA 493] gb|AAO90713.11 aldehyde dehydrogenase family protein [Coxiella burnetii RSA 493] Length = 455
860.2 Best-BlastP=> >nrprot 28% Identities = 112/372 (30%), Positives = 167/372 (44%), Gaps = 46/372 (12%) ref|XP_313257.11
ENSANGP00000010409 [Anopheles gambiae] gblEAA08897.1 | ENSANGP00000010409 [Anopheles gambiae str. PEST] Length = 2206
862.1 Best-BlastP=> >nrprot No Hits found 863.1 Best-BlastP=> >nrprot 72% Identities = 70/114 (61%), Positives = 90/114 (78%) ref)ZP_00092569.11 COG3293: Transposase and inactivated derivatives [Azotobacter vinelandii] Length = 249
864.1 Best-BlastP=> >nrprot 85% Identities = 70/95 (73%), Positives = 83/95 (87%) ref|NP_820693.11 conserved hypothetical protein [Coxiella burnetii RSA 493] gb|AA091207.11 conserved hypothetical protein [Coxiella burnetii RSA 493] Length = 96
865.2 Best-BlastP=> >nrprot 75% Identities = 227/379 (59%), Positives = 280/379 (73%), Gaps = 12/379 (3%) ref|NP_232020.11 carbamoyl-phosphate synthase, small subunit [Vibrio cholerae 01 biovar eltor str. N16961] sp|Q9KPH8|CARA_VIBCH Carbamoyl-phosphate synthase small chain (Carbamoyl-phosphate synthetase glutamine chain) pir||E82083 carbamoyl-phosphate synthase, small chain VC2390 [imported] - Vibrio cholerae (strain N16961 serogroup 01) gb|AAF95533.11 carbamoyl-phosphate synthase, small subunit [Vibrio cholerae 01 biovar eltor str. N16961] Length = 379
866.2 Best-BlastP=> >nrprot 99% Identities = 379/379 (100%), Positives = 379/379 (100%) sp|P50025|DNAJ_LEGPN Chaperone protein dnaJ gb|AAA80278.11 heat-shock protein Length = 37g
869.2 Best-BlastP=> >nrprot No Hits found
870.3 Best-BlastP=> >nrprot 98% Identities = 355/360 (98%), Positives = 357/360 (99%) pir||T18333 icmK protein - Legionella pneumophila gb|AAC38189.1 | DotH [Legionella pneumophila] emb|CAA75330.1 | IcmK protein [Legionella pneumophila] Length = 360
872.2 Best-BlastP=> >nrprot 99% Identities = 212/212 (100%), Positives = 212/212 (100%) pir||T18332 icmL protein - Legionella pneumophila gb|AAC38190.1 | Dotl [Legionella pneumophila] emb|CAA75329.1 | IcmL protein [Legionella pneumophila] emb|CAD43145.1 | Dotl protein [Legionella pneumophila serogroup 6] Length = 212
873.2 Best-BlastP=> >nrprot 96% Identities = 91/94 (96%), Positives = 92/94 (97%) pir||T18331 icmM protein - Legionella pneumophila gb|AAC38191.1 | DotJ [Legionella pneumophila] emb|CAA75328.1 | IcmM protein [Legionella pneumophila] Length = 94
874.2 Best-BlastP=> >nrprot 99% Identities = 187/189 (98%), Positives = 189/189 (100%) pir||T18330 IphA protein - Legionella pneumophila emb|CAA75327.11 LphA protein [Legionella pneumophila] Length = 189
875.5 Best-BlastP=> >nrprot 53% Identities = 133/408 (32%), Positives = 227/408 (55%), Gaps = 10/408 (2%) ref|NP_827016.11 putative integral membrane transport protein [Streptomyces avermitilis MA-4680] dbj|BAC73551.11 putative integral membrane transport protein [Streptomyces avermitilis MA-4680] Length = 490
876.1 Best-BlastP=> >nrprot 23% Identities = 23/67 (34%), Positives = 37/67 (55%), Gaps = 2/67 (2%) ref|NP_355599.11 AGR_C_4826p [Agrobacterium tumefaciens] ref|NP_533327.11 acetyltransferase [Agrobacterium tumefaciens str. C58 (U. Washington)] pir||G97678 hypothetical protein AGR_C_4826 [imported] - Agrobacterium tumefaciens (strain C58, Cereon) pir||AE2903 acetyltransferase [imported] - Agrobacterium tumefaciens (strain C58, Dupont) gb|AAK88384.1 | AGR_C_4826p [Agrobacterium tumefaciens str. C58 (Cereon)] gb|AAL43643.11 acetyltransferase [Agrobacterium tumefaciens str. C58 (U. Washington)] Length = 168
879.2 Best-BlastP=> >nrprot 60% Identities = 65/153 (42%), Positives = 94/153 (61 %), Gaps = 10/153 (6%) ref|ZP_00090468.11 COG0582: Integrase [Azotobacter vinelandii] Length = 399
880.3 Best-BlastP=> >nrprot 60% Identities = 137/352 (38%), Positives = 215/352 (61 %) ref |NP_819830.11 membrane protein, putative [Coxiella burnetii RSA 493] gb|AAO90344.1 | membrane protein, putative [Coxiella burnetii RSA 493] Length = 355
881.2 Best-BlastP=> >nrprot 61% Identities = 141/359 (39%), Positives = 221/359 (61%), Gaps = 7/359 (1 %) ref |NP_819829.1 ] membrane protein, putative [Coxiella burnetii RSA 493] gb|AAO90343.11 membrane protein, putative [Coxiella burnetii RSA 493] Length = 370
882.3 Best-BlastP=> >nrprot 63% Identities = 218/487 (44%), Positives = 308/487 (63%), Gaps = 12/487 (2%) ref|NP_439847.11 aminopeptidase A/I [Haemophilus influenzae Rd] sp|P45334|AMPA_HAEIN Cytosol aminopeptidase (Leucine aminopeptidase) (LAP) (Leucyl aminopeptidase) pir||C64137 leucyl aminopeptidase (EC 3.4.11.1) A - Haemophilus influenzae (strain Rd KW20) gb|AAC23351.1 | aminopeptidase A/I (pepA) [Haemophilus influenzae Rd] Length = 491
884.3 Best-BlastP=> >nrprot 23% Identities = 48/193 (24%), Positives = 77/193 (39%), Gaps = 9/193 (4%) ref |NP_624872.11 hypothetical protein SCF73.06c [Streptomyces coelicolor A3(2)] emb|CAB57411.11 hypothetical protein SCF73.06c [Streptomyces coelicolor A3(2)] Length = 333
885.2 Best-BlastP=> >nrprot 38% Identities = 160/289 (55%), Positives = 204/289 (70%) ref|NP_386181.1 | HYPOTHETICAL PROTEIN [Sinorhizobium meliloti] emb|CAC46654.11 HYPOTHETICAL PROTEIN [Sinorhizobium meliloti] Length = 299
886.3 Best-BlastP=> >nrprot 31 % Identities = 50/153 (32%), Positives = 83/153 (54%), Gaps = 2/153 (1 %) ref|NP_799124.11 shikimate kinase [Vibrio parahaemolyticus RIMD 2210633] sp|Q87L67|AROK_VIBPA Shikimate kinase (SK) dbj|BAC61008.11 shikimate kinase [Vibrio parahaemolyticus] Length = 172
887.4 Best-BlastP=> >nrprot 18% Identities = 82/315 (26%), Positives = 132/315 (41 %), Gaps = 67/315 (21 %) ref|NP_229450.11 alpha-amylase, putative [Thermotoga maritima] pir||G72227 hypothetical protein TM1650 - Thermotoga maritima (strain MSB8) gb|AAD36717.1 |AE001807_8 alpha-amylase, putative [Thermotoga maritima] Length = 422
888.3 Best-BlastP=> >nrprot 73% Identities = 240/415 (57%), Positives = 303/415 (73%), Gaps = 2/415 (0%) ref|NP_794674.11 GTP-binding protein HflX [Pseudomonas syringae pv. tomato str. DC3000] gb|AA058369.11 GTP-binding protein HflX [Pseudomonas syringae pv. tomato str. DC3000] Length = 433
889.1 Best-BlastP=> >nrprot 45% Identities = 45/128 (35%), Positives = 71/128 (55%), Gaps = 1/128 (0%) sp|P35160|RESA_BACSU Thiol-disulfide oxidoreductase resA Length = 179
891.3 Best-BlastP=> >nrprot 29% Identities = 84/420 (20%), Positives = 157/420 (37%), Gaps = 115/420 (27%) emb|CAC18568.11 phosphatidylcholine- hydrolyzing phospholipase C [Pseudomonas fluorescens] Length = 385
893.4 Best-BlastP=> >nrprot 67% Identities = 430/811 (53%), Positives = 561/811 (69%), Gaps = 6/811 (0%) dbj|BAA77891.11 Acyl-CoA dehydrogenase (EC 1.3.99.-). [Escherichia coli] Length = 840
894.3 Best-BlastP=> >nrprot No Hits found 895.3 Best-BlastP=> >nrprot 54% Identities = 102/300 (34%), Positives = 159/300 (53%), Gaps = 25/300 (8%) ref|ZP_00107238.11 COG0697: Permeases of the drug/metabolite transporter (DMT) superfamily [Nostoc punctiforme] Length = 369
897.2 Best-BlastP=> >nrprot 99% Identities = 764/764 (100%), Positives = 764/764 (100%) gb|AAF05327.2l phosphoenolpyruvate-protein phosphotransferase PtsP [Legionella pneumophila] Length = 764
898.2 Best-BlastP=> >nrprot 57% Identities = 96/257 (37%), Positives = 156/257 (60%), Gaps = 1/257 (0%) ref|ZP_00130066.1 | COG0730: Predicted permeases [Desulfovibrio desulfuricans G20] Length = 265
899.3 Best-BlastP=> >nrprot 77% Identities = 163/257 (63%), Positives = 200/257 (77%), Gaps = 1/257 (0%) ref|NP_820532.11 prolipoprotein diacylglyceryl transferase [Coxiella burnetii RSA 493] gb|AA091046.11 prolipoprotein diacylglyceryl transferase [Coxiella burnetii RSA 493] Length = 261
9.1 Best-BlastP=> >nrprot 44% Identities = 57/140 (40%), Positives = 75/140 (53%), Gaps = 4/140 (2%) dbj|BAC94688.11 hypothetical protein [Vibrio vulnificus YJ016] Length = 343
901.2 Best-BlastP=> >nrprot No Hits found 903.2 Best-BlastP=> >nrprot No Hits found 905.2 Best-BlastP=> >nrprot 78% Identities = 176/285 (61 %), Positives = 225/285 (78%), Gaps = 2/285 (0%) ref|NP_773784.11 oxidoreductase [Bradyrhizobium japonicum] dbj|BAC52409.11 oxidoreductase [Bradyrhizobium japonicum USDA 110] Length = 308
907.4 Best-BlastP=> >nrprot 81% Identities = 571/879 (64%), Positives = 727/879 (82%), Gaps = 2/879 (0%) ref |ZP_00091354.11 COG2609: Pyruvate dehydrogenase complex, dehydrogenase (E1 ) component [Azotobacter vinelandii] Length = 893
908.4 Best-BlastP=> >nrprot 69% Identities = 273/562 (48%), Positives = 378/562 (67%), Gaps = 26/562 (4%) ref |NP_716062.11 pyruvate dehydrogenase complex, E2 component, dihydrolipoamide acetyltransferase [Shewanella oneidensis MR-1] gb|AAN53507.1 |AE015490_8 pyruvate dehydrogenase complex, E2 component, dihydrolipoamide acetyltransferase [Shewanella oneidensis MR-1] Length = 677
909.2 Best-BlastP=> >nrprot 54% Identities = 107/278 (38%), Positives = 156/278 (56%) ref|NP_820642.11 conserved hypothetical protein [Coxiella burnetii RSA 493] gb|AA091156.11 conserved hypothetical protein [Coxiella burnetii RSA 493] Length = 294
910.4 Best-BlastP=> >nrprot 78% Identities = 476/739 (64%), Positives = 590/739 (79%) ref|NP_253651.11 topoisomerase IV subunit A [Pseudomonas aeruginosa PA01] pir||G83025 topoisomerase IV subunit A PA4964 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG08349.1 |AE004909_5 topoisomerase IV subunit A [Pseudomonas aeruginosa PA01] Length = 754
914.3 Best-BlastP=> >nrprot 70% Identities = 338/573 (58%), Positives = 411/573 (71 %), Gaps = 6/573 (1%) gb|AAL27383.1 |AF426171_14 putative ABC transporter ElsE [Yersinia pestis] Length = 580
915 3 Best-BlastP=> >nrprot 64% Identities = 132/332 (39%), Positives = 183/332 (55%), Gaps = 54/332 (16%) ref |NP_455356.11 HlyD-family secretion protein [Salmonella enterica subsp. enterica serovar Typhi] ref|NP_805830.11 HlyD-family secretion protein [Salmonella enterica subsp. enterica serovar Typhi Ty2] sp|Q8Z879|YBHG_SALTI Hypothetical UPF0194 membrane protein ybhG pir||AG0599 HlyD-family secretion protein [imported] - Salmonella enterica subsp. enterica serovar Typhi (strain CT18) emb|CAD05265.11 HlyD-family secretion protein [Salmonella enterica subsp. enterica serovar Typhi] gb|AAO69690.11 HlyD-family secretion protein [Salmonella enterica subsp.
enterica serovar Typhi Ty2] Length = 331
916.3 Best-BlastP=> >nrprot 69% Identities = 162/336 (48%), Positives = 232/336 (69%), Gaps = 11/336 (3%) ref|NP_819097.11 ferrochelatase [Coxiella burnetii RSA 493] gb|AA089611.11 ferrochelatase [Coxiella burnetii RSA 493] Length = 343
917.3 Best-BlastP=> >nrprot 65% Identities = 43/71 (60%), Positives = 51/71 (71 %) ref|NP_404959.11 cold shock-like protein [Yersinia pestis] pir||AH0166 cold shock-like protein [imported] - Yersinia pestis (strain C092) emb|CAC90195.11 cold shock-like protein [Yersinia pestis C092] Length = 87
918.3 Best-BlastP=> >nrprot No Hits found
919.1 Best-BlastP=> >nrprot 62% Identities = 51/125 (40%), Positives = 86/125 (68%), Gaps = 1/125 (0%) ref|NP_249650.11 hypothetical protein [Pseudomonas aeruginosa PA01] pir||A83524 hypothetical protein PA0959 [imported] - Pseudomonas aeruginosa (strain PA01 ) gb|AAG04348.1 |AE004530_1 hypothetical protein PA0959 [Pseudomonas aeruginosa PA01] Length = 209
920.2 Best-BlastP=> >nrprot 43% Identities = 28/116 (24%), Positives = 52/116 (44%), Gaps = 4/116 (3%) ref|ZP_00082077.11 hypothetical protein [Geobacter metallireducens] Length = 130
921.3 Best-BlastP=> >nrprot 71 % Identities = 107/194 (55%), Positives = 141/194 (72%), Gaps = 1/194 (0%) ref|NP_760875.1 | Recombinational DNA repair protein [Vibrio vulnificus CMCP6] sp|Q8DB22|RECR_VIBVU Recombination protein recR gb|AA010402.1 |AE016803_189 Recombinational DNA repair protein [Vibrio vulnificus CMCP6] dbj|BAC95173.11 Recombinational DNA repair protein [Vibrio vulnificus YJ016] Length = 200
922.2 Best-BlastP=> >nrprot 54% Identities = 101/244 (41 %), Positives = 150/244 (61 %), Gaps = 10/244 (4%) ref|NP_232123.11 hypothetical protein VC24g4 [Vibrio cholerae 01 biovar eltor str. N16961] pir||A82069 hypothetical protein VC2494 [imported] - Vibrio cholerae (strain N16961 serogroup 01 ) gb|AAF95636.1 | hypothetical protein [Vibrio cholerae 01 biovar eltor str. N16961] Length = 263
923.3 Best-BlastP=> >nrprot 26% Identities = 104/453 (22%), Positives = 180/453 (39%), Gaps = 103/453 (22%) ref |NP_126866.1 | hypothetical protein [Pyrococcus abyssi] pir||C75099 hypothetical protein PAB0793 - Pyrococcus abyssi (strain Orsay) emb|CAB50096.1 | Sulfatase [Pyrococcus abyssi] Length = 479
925.3 Best-BlastP=> >nrprot 63% Identities = 282/620 (45%), Positives = 401/620 (64%), Gaps = 9/620 (1 %) ref|NP_246321.11 SpeA [Pasteurella multocida] gb|AAK03466.11 SpeA [Pasteurella multocida] Length = 644
926.4 Best-BlastP=> >nrprot 55% Identities = 85/232 (36%), Positives = 130/232 (56%), Gaps = 1/232 (0%) ref|NP_743997.11 glutamine amidotransferase, class I [Pseudomonas putida KT2440] gb|AAN67461.1 |AE016373_3 glutamine amidotransferase, class I [Pseudomonas putida KT2440] Length = 244
928.1 Best-BlastP=> >nrprot 72% Identities = 217/380 (57%), Positives = 279/380 (73%) ref|NP_386317.11 PUTATIVE ALCOHOL DEHYDROGENASE PROTEIN [Sinorhizobium meliloti] emb|CAC46790.11 PUTATIVE ALCOHOL DEHYDROGENASE PROTEIN [Sinorhizobium meliloti] Length = 381
929.2 Best-BlastP=> >nrprot 72% Identities = 255/464 (54%), Positives = 339/464 (73%), Gaps = 3/464 (0%) ref|NP_104264.11 dehydrogenase, (succinatesemialdehyde dehydrogenase, aldehyde dehydrogenase, aldehyde dehydrogenase) [Mesorhizobium loti] dbj|BAB50050.1 | dehydrogenase; succinatesemialdehyde dehydrogenase; aldehyde dehydrogenase; aldehyde dehydrogenase [Mesorhizobium loti] Length = 462
930.3 Best-BlastP=> >nrprot 67% Identities = 226/441 (51 %), Positives = 309/441 (70%), Gaps = 3/441 (0%) ref|NP_220602.11 CYTOCHROME D UBIQUINOL OXIDASE SUBUNIT I (cydA) [Rickettsia prowazekii] pir||H71732 cytochrome D ubiquinol oxidase chain I (cydA) RP216 - Rickettsia prowazekii emb|CAA14679.11 CYTOCHROME D UBIQUINOL OXIDASE SUBUNIT I (cydA) [Rickettsia prowazekii] Length = 453
932.1 Best-BlastP=> >nrprot 69% Identities = 157/331 (47%), Positives = 229/331 (69%), Gaps = 3/331 (0%) ref|NP_794398.11 cyanide-insensitive terminal oxidase CioB [Pseudomonas syringae pv. tomato str. DC3000] gb|AAO58093.11 cyanide-insensitive terminal oxidase CioB [Pseudomonas syringae pv. tomato str. DC3000] Length = 335
935.2 Best-BlastP=> >nrprot 50% Identities = 81/260 (31%), Positives = 132/260 (50%), Gaps = 6/260 (2%) ref|NP_532654.1 | conserved hypothetical protein [Agrobacterium tumefaciens str. C58 (U. Washington)] pir||AD2819 conserved hypothetical protein Atu1975 [imported] - Agrobacterium tumefaciens (strain C58, Dupont) gb|AAL42970.1 | conserved hypothetical protein [Agrobacterium tumefaciens str. C58 (U. Washington)] Length = 262
936.3 Best-BlastP=> >nrprot 35% Identities = 38/156 (24%), Positives = 65/156 (41%), Gaps = 22/156 (14%) ref |NP_718932.11 conserved hypothetical protein [Shewanella oneidensis MR-1] gb|AAN56376.1 |AE015774_11 conserved hypothetical protein [Shewanella oneidensis MR-1] Length = 190
937.3 Best-BlastP=> >nrprot 67% Identities = 237/470 (50%), Positives = 320/470 (68%), Gaps = 8/470 (1 %) ref|NP_820223.11 sensor histidine kinase [Coxiella burnetii RSA 493] gb|AAO90737.11 sensor histidine kinase [Coxiella burnetii RSA 493] Length = 478
938.3 Best-BlastP=> >nrprot 6% Identities = 22/49 (44%), Positives = 36/49 (73%) gb|EAA16201.11 ERYTHROCYTE MEMBRANE PROTEIN PFEMP3 [Plasmodium yoelii yoelii] Length = 643
94.7 Best-BlastP=> >nrprot 23% Identities = 354/1425 (24%), Positives = 554/1425 (38%), Gaps = 309/1425 (21%) ref|NP_772111.11 bll-5471 [Bradyrhizobium japonicum] dbj|BAC50736.11 bll5471 [Bradyrhizobium japonicum USDA 110] Length = 4210
940.4 Best-BlastP=> >nrprot 60% Identities = 134/293 (45%), Positives = 176/293 (60%), Gaps = 23/293 (7%) ref|NP_931279.11 ribosomal protein L11 methyltransferase [Photorhabdus luminescens subsp. laumondii TT01] emb|CAE16459.1 | ribosomal protein L11 methyltransferase [Photorhabdus luminescens subsp. laumondii TT01] Length = 296
941.3 Best-BlastP=> >nrprot 85% Identities = 333/444 (75%), Positives = 381/444 (85%) ref|NP_931267.11 biotin carboxylase (A subunit of acetyl-CoA carboxylase) (ACC) [Photorhabdus luminescens subsp. laumondii TT01] emb|CAE16447.11 biotin carboxylase (A subunit of acetyl-CoA carboxylase) (ACC) [Photorhabdus luminescens subsp. laumondii TT01] Length = 449
943.3 Best-BlastP=> >nrprot 72% Identities = 86/160 (53%), Positives = 117/160 (73%), Gaps = 6/160 (3%) ref |NP_760165.11 Biotin carboxyl carrier protein [Vibrio vulnificus CMCP6] gb|AAO09692.1 |AE016801_11 Biotin carboxyl carrier protein [Vibrio vulnificus CMCP6] dbj|BAC95899.11 biotin carboxyl carrier protein [Vibrio vulnificus YJ016] Length = 154
944.3 Best-BlastP=> >nrprot 78% Identities = 93/145 (64%), Positives = 114/145 (78%) reflZP_00090970.1 1 COG0757: 3-dehydroquinate dehydratase II [Azotobacter vinelandii] Length = 147
946.3 Best-BlastP=> >nrprot 62% Identities = 80/201 (39%), Positives = 126/201 (62%) ref|NP_768568.11 bll1928 [Bradyrhizobium japonicum] gb|AAG60905.1 |AF322013_24 ID481 [Bradyrhizobium japonicum] dbj|BAC47193.1 | bll1928 [Bradyrhizobium japonicum USDA 110] Length = 223
947.3 Best-BlastP=> >nrprot 43% Identities = 42/78 (53%), Positives = 53/78 (67%) ref|NP_768567.11 bll1927 [Bradyrhizobium japonicum] gb|AAG60904.1 |AF322013_23 ID479 [Bradyrhizobium japonicum] dbj|BAC47192.1 | bll1927 [Bradyrhizobium japonicum USDA 110] Length = 374
948.3 Best-BlastP=> >nrprot 72% Identities = 88/155 (56%), Positives = 118/155 (76%) ref|ZP_00092102.11 COG1943: Transposase and inactivated derivatives [Azotobacter vinelandii] Length = 270
949.3 Best-BlastP=> >nrprot 70% Identities = 322/616 (52%), Positives = 436/616 (70%), Gaps = 4/616 (0%) ref|NP_820141.1 | protein-export membrane protein SecD [Coxiella burnetii RSA 493] gb|AAO90655.11 protein-export membrane protein SecD [Coxiella burnetii RSA 493] Length = 622
950.2 Best-BlastP=> >nrprot 15% Identities = 49/165 (29%), Positives = 77/165 (46%), Gaps = 13/165 (7%) emb|CAA70596.1 | cinnamate 4- hydroxylase [Phaseolus vulgaris] Length = 355
952.1 Best-BlastP=> >nrprot 46% Identities = 87/296 (29%), Positives = 142/296 (47%), Gaps = 12/296 (4%) gb|AAD02145.11 regulatory protein [Pseudomonas stutzeri] Length = 300
953.2 Best-BlastP=> >nrprot 12% Identities = 30/1 18 (25%), Positives = 62/118 (52%), Gaps = 19/118 (16%) ref|NP_785847.11 esterase (putative) [Lactobacillus plantarum WCFS1] emb|CAD64698.11 esterase (putative) [Lactobacillus plantarum WCFS1] Length = 339
956.2 Best-BlastP=> >nrprot 56% Identities = 102/249 (40%), Positives = 143/249 (57%), Gaps = 3/249 (1 %) ref|NP_419511 .11 transcriptional regulator SkgA [Caulobacter crescentus CB15] sp|Q9RP67|SKGA_CAUCR Transcriptional regulator skgA (Stationary-phase regulation of katG protein) pir||C87335 transcription regulator SkgA [imported] - Caulobacter crescentus gb|AAF01797.1 |AF170912_1 putative helix-tum-helix transcriptional regulator SkgA [Caulobacter crescentus] gb|AAK22679.11 transcriptional regulator SkgA [Caulobacter crescentus CB15] Length = 255
957.2 Best-BlastP=> >nrprot 44% Identities = 89/269 (33%), Positives = 129/269 (47%), Gaps = 3/269 (1 %) ref|NP_337940.11 hydrolase, alpha/beta hydrolase fold family [Mycobacterium tuberculosis CDC1551] gb|AAK47754.11 hydrolase, alpha/beta hydrolase fold family [Mycobacterium tuberculosis CDC1551 ] Length = 310
959.2 Best-BlastP=> >nrprot No Hits found
96.6 Best-BlastP=> >nrprot 99% Identities = 276/277 (99%), Positives = 277/277 (100%) gb|AAM00632.11 unknown [Legionella pneumophila] Length = 2g4
960.2 Best-BlastP=> >nrprot 82% Identities = 79/102 (77%), Positives = 86/102 (84%) gb|AAM00607.11 unknown [Legionella pneumophila] Length = 121
964.3 Best-BlastP=> >nrprot 24% Identities = 61/202 (30%), Positives = 103/202 (50%), Gaps = 18/202 (8%) gb|AAF05325.11 unknown virulence protein [Legionella pneumophila] Length = 205
965.3 BBeesstt--BBllaassttPP==>> >nrprot 99% Identities = 118/118 (100%), Positives = 118/118 (100%) gb|AAQ10305.11 DotV [Legionella pneumophila] Length = 180
966.2 Best-BlastP=> >nrprot 53% Identities = 95/244 (38%), Positives = 133/244 (54%), Gaps = 7/244 (2%) ref|NP_639315.11 phenol hydroxylase [Xanthomonas campestris pv. campestris str. ATCC 33913] gb|AAM43197.11 phenol hydroxylase [Xanthomonas campestris pv. campestris str. ATCC 33913] Length = 307
967.2 Best-BlastP=> >nrprot 71 % Identities = 184/333 (55%), Positives = 242/333 (72%) ref|ZP_00032409.11 COG3588: Fructose-1 ,6-bisphosphate aldolase [Burkholderia fungorum] Length = 370
969.2 Best-BlastP=> >nrprot 67% Identities = 116/243 (47%), Positives = 168/243 (69%), Gaps = 1/243 (0%) ref|NP_819931.11 endonuclease/exonuclease/phosphatase family [Coxiella burnetii RSA 493] gb|AAO90445.1 | endonuclease/exonuclease/phosphatase family [Coxiella burnetii RSA 493] Length = 255
97.2 Best-BlastP=> >nrprot No Hits found
971.2 Best-BlastP=> >nrprot No Hits found
972.3 Best-BlastP=> >nrprot 24% Identities = 82/304 (26%), Positives = 130/304 (42%), Gaps = 29/304 (9%) ref|NP_477218.11 loki CG10895-PB [Drosophila melanogaster] ref|NP_724241.11 loki CG10895-PC [Drosophila melanogaster] dbj|BAA28756.11 short form of nuclear kinase [Drosophila melanogaster] gb|AAL48020.1 | LD27857p [Drosophila melanogaster] gb|AAN11062.11 CG10895-PB [Drosophila melanogaster] gb|AAN11063.11 CG10895-PC [Drosophila melanogaster] Length = 459
974.2 Best-BlastP=> >nrprot 58% Identities = 151/387 (39%), Positives = 226/387 (58%), Gaps = 8/387 (2%) ref|NP_925546.11 aspartate aminotransferase [Gloeobacter violaceus] dbj|BAC90541.11 aspartate aminotransferase [Gloeobacter violaceus] Length = 395
976.3 Best-BlastP=> >nrprot 58% Identities = 31/82 (37%), Positives = 51/82 (62%) ref|NP_378270.11 341aa long conserved hypothetical protein [Sulfolobus tokodaii] dbj|BAB67379.1 | 341aa long conserved hypothetical protein [Sulfolobus tokodaii] Length = 341
977.3 Best-BlastP=> >nrprot 39% Identities = 69/181 (38%), Positives = 88/181 (48%), Gaps = 6/181 (3%) ref|NP_632394.11 chorismate mutase; prephenate dehydratase [Methanosarcina mazei Goel] gb|AAM30066.11 chorismate mutase; prephenate dehydratase [Methanosarcina mazei Goel] Length = 354
979.3 Best-BlastP=> >nrprot 40% Identities = 124/252 (49%), Positives = 162/252 (64%), Gaps = 3/252 (1 %) gb|AAB97629.11 malonate decarboxylase beta subunit [Acinetobacter calcoaceticus] Length = 295
980.1 Best-BlastP=> >nrprot 58% Identities = 89/229 (38%), Positives = 140/229 (61 %), Gaps = 3/229 (1 %) ref|NP_640916.11 malonate decarboxylase gamma subunit [Xanthomonas axonopodis pv. citri str. 306] gb|AAM35452.11 malonate decarboxylase gamma subunit [Xanthomonas axonopodis pv. citri str. 306] Length = 234
981 .2 Best-BlastP=> >nrprot 52% Identities = 60/217 (27%), Positives = 113/217 (52%), Gaps = 29/217 (13%) gb| AAF20287.1 |AF121266_9 MdcE [Acinetobacter calcoaceticus] Length = 226
982.3 Best-BlastP=> >nrprot 64% Identities = 196/465 (42%), Positives = 301/465 (64%), Gaps = 21/465 (4%) ref|NP_819781.1 | protease DO [Coxiella burnetii RSA 493] gb|AAO90295.1 | protease DO [Coxiella burnetii RSA 493] Length = 451
983.1 Best-BlastP=> >nrprot 64% Identities = 112/239 (46%), Positives = 156/239 (65%), Gaps = 7/239 (2%) ref|NP_230359.11 conserved hypothetical protein [Vibrio cholerae 01 biovar eltor str. N16961] pir||H82289 conserved hypothetical protein VC0710 [imported] - Vibrio cholerae (strain N 16961 serogroup 01 ) gb|AAF93875.1 | conserved hypothetical protein [Vibrio cholerae 01 biovar eltor str. N 16961] Length = 240
985.2 Best-BlastP=> >nrprot 66% Identities = 148/309 (47%), Positives = 215/309 (69%), Gaps = 4/309 (1 %) ref|NP_779940.11 ribosomal large subunit pseudouridine synthase D [Xylella fastidiosa Temeculal] sp|Q87AR7|RLUD_XYLFT Ribosomal large subunit pseudouridine synthase D (Pseudouridylate synthase) (Uracil hydrolyase) gb|AA029589.11 ribosomal large subunit pseudouridine synthase D [Xylella fastidiosa Temeculal] Length = 331
986.3 Best-BlastP=> >nrprot 83% Identities = 55/83 (66%), Positives = 72/83 (86%), Gaps = 3/83 (3%) ref|ZP_00065010.11 COG1923: Uncharacterized host factor I protein [Microbulbifer degradans 2-40] Length = 86
989.3 Best-BlastP=> >nrprot No Hits found 99.1 Best-BlastP=> >nrprot No Hits found 991.2 Best-BlastP=> >nrprot 53% Identities = 104/276 (37%), Positives = 152/276 (55%), Gaps = 3/276 (1 %) ref|NP_680903.11 ORF_ID:tlr0112~probable hydrolase [Thermosynechococcus elongatus BP-1] dbj|BAC07665.11 ORF_ID:tlr0112~probable hydrolase [Thermosynechococcus elongatus BP-1] Length = 285
992.2 Best-BlastP=> >nrprot No Hits found 993.2 Best-BlastP=> >nrprot 80% Identities = 285/416 (68%), Positives = 332/416 (79%), Gaps = 3/416 (0%) ref|NP_820266.11 phosphofructokinase, putative [Coxiella burnetii RSA 493] gb|AAO90780.11 phosphofructokinase, putative [Coxiella burnetii RSA 493] Length = 420
994.2 Best-BlastP=> >nrprot 53% Identities = 284/930 (30%), Positives = 483/930 (51 %), Gaps = 57/930 (6%) ref|ZP_00126364.11 COG0784: FOG: CheY-like receiver [Pseudomonas syringae pv. syringae B728a] Length = 917
995.2 Best-BlastP=> >nrprot 70% Identities = 109/212 (51%), Positives = 151/212 (71%), Gaps = 6/212 (2%) ref|NP_717282.11 glutathione S- transferase family protein [Shewanella oneidensis MR-1] gb|AAN54726.1 |AE015613_3 glutathione S-transferase family protein [Shewanella oneidensis MR-1] Length = 216
996.2 Best-BlastP=> >nrprot 79% Identities = 221/328 (67%), Positives = 266/328 (81%), Gaps = 1/328 (0%) ref|NP_717281.11 fumarylacetoacetate hydrolase family protein [Shewanella oneidensis MR-1 ] gb|AAN54725.1 |AE015613_2 fumarylacetoacetate hydrolase family protein [Shewanella oneidensis MR-1] Length = 328
997.2 Best-BlastP=> >nrprot 99% Identities = 345/348 (99%), Positives = 347/348 (99%) sp|Q53407|LLY_LEGPN 4-HYDROXYPHENYLPYRUVATE DIOXYGENASE (4HPPD) (HPD) (LEGIOLYSIN) gb|AAC32843.11 legiolysin [Legionella pneumophila] Length = 348
998.3 Best-BlastP=> >nrprot 69% Identities = 337/597 (56%), Positives = 425/597 (71%), Gaps = 4/597 (0%) ref |NP_819507.11 ATP-dependent DNA helicase RecQ [Coxiella burnetii RSA 493] gb|AAO90021.11 ATP-dependent DNA helicase RecQ [Coxiella burnetii RSA 493] Length = 601
999.2 Best-BlastP=> >nrprot 75% Identities = 225/375 (60%), Positives = 290/375 (77%) ref|ZP_00056039.11 COG1960: Acyl-CoA dehydrogenases [Magnetospirillum magnetotacticum] Length = 408
Table XIV EMBL Name ORF SEQ ID .. , „_ Posit°l Posit°2 Sens SignalP of the Product of the gene NAME Note Class gene Chromosomal replication 2402.4 4329 IppOOOl 204 1562 dnaA initiator protein DnaA 3.1 DNA polymerase III, beta 2404.2 4330 Ipp0002 1576 2679 P dnaN chain 3.1 RecF recombinational DNA 2406.3 4331 Ipp0003 2676 3737 P recF repair ATPase 3.3 DNA gyrase, subunit B 1041.3 3535 Ipp0004 4034 6451 P gyrB (type II topoisomerase) 3.4 1040.3 3534 Ipp0005 6821 7861 Similar to peptidylarginine m unknown deiminase and related enzymes 2.2 925.3 6541 Ipp0006 7864 9756 similar to biosynthetic arginine m speA unknown decarboxylase 2.2 Putative carbon-nitrogen hydrolase 991.2 6583 Ipp0007 9776 10621 m unknown family protein 2.1.1 989.3 6581 IppOOOδ 11162 12421 m unknown 6 986.3 6580 Ipp0009 12609 12866 P unknown Similar to host factor- 1 protein 4.4 888.3 6512 IppOOlO 12894 14138 P unknown Similar to GTP-binding protein HflX 4.6 889.1 6513 IppOOll 14243 14713 m unknown Similar to thioredoxin 1.4 891.3 6514 Ipp0012 14880 16457 m unknown 6 5216.2 6044 Ipp0013 16603 17442 P unknown Similar to other protein 5.2 3892.3 5223 Similar to conserved hypothetical Ipp0014 17975 19090 P unknown protein 5.2 Similar to multidrug resistance 3895.2 5224 Ipp0015 19192 20265 unknown efflux pump 1.2 5642.1 6229 Ipp0016 20240 21172 P unknown 6 4918.2 5900 Ipp0017 21157 22062 P unknown 6 4919.1 similar to outer membrane efflux 5901 Ipp0018 22046 23431 P unknown proteins 1.2 5644.3 6230 Similar to Legionella zinc Ipp0019 23666 25342 P unknown metalloproteinase precursor 2.2 4398.2 5567 Ipp0020 25563 26474 P unknown Putative integral membrane protein 5.2
able XIV similar to conserved hypothetical
4396.1 5566 Ipp0021 26665 27144 P unknown protein 5.2
4395.2 Similar to conserved hypothetical 5565 Ipp0022 27141 29264 P unknown protein 5.2
5380.3 6106 Ipp0023 29334 29882 m unknown Putative membrane protein 5.2 341.6 4921 Ipp0024 30029 30454 m + hbp hemin binding protein 4.1 Rep protein, confers resistance to cationic 339.5 4907 Ipp0025 30594 31166 m + rep antimicrobial peptides and promotes intracellular infection 337.3 4898 Ipp0026 31326 32717 P unknown similar to amino acid permease 1 similar to low-affinity inorganic 336.1 4891 Ipp0027 32727 33722 m unknown phosphate transporter 1 335.1 4889 similar to ubiquinone biosynthesis Ipp0028 33739 34380 m unknown protein 2 334.5 4885 Ipp0029 34603 37533 P Unknown 1
3830.2 5191 Ipp0030 37645 38535 m unknown 6
3832.1 5192 Ipp0031 38594 39787 m + unknown Similar to aminopeptidase 2.2
3834.1 5193 Ipp0032 39849 40874 m unknown 6
2567.2 4437 lpp0033 41138 41899 P unknown 6
2568.1 4438 Ipp0034 42156 42512 P + unknown 6 Similar to conserved hypothetical
2569.3 4439 Ipp0035 42575 43297 m unknown protein 5.2 Similar to arginine transport
3835.2 5194 Ipp0036 43511 44254 m unknown system periplasmic binding protein 1.2
3837.4 5195 Ipp0037 44415 45932 m unknown Ankyrin repeat protein 4.6
6151.1 6338 Ipp0038 46159 47244 P Unknown Similar to transposase (IS4 family) 4.5
5999.1 6317 Ipp0039 47270 47563 P unknown Similar to unknown proteins 5.2
4715.3 5769 Ipp0040 47833 48315 P + unknown Similar to hypothetical protein 5.2 Similar to conserved hypothetical
4714.1 5768 Ipp0041 48337 49332 m unknown protein 5.2
4713.1 5767 Ipp0042 49497 49820 P unknown 6 Similar to conserved hypothetical
5532.2 6175 Ipp0043 50133 50993 m unknown protein 5.2
5533.2 6176 Ipp0044 51171 51407 m unknown 6
en c TO c c c C c C C c c C C C C C C C c c 5 5 5 5 5 5 5 5 5 5 5 5 5 5 ^ 2 5 5 5 5 O O Oo O o o o o o o o o o o o o o o ro o o o o o o CZ Z cZ c c c c c c c c c c c c c c c - -5 c c c c c c _--: .^ -^ ..- .-i __- _*: -i. -: -V. --£ ... .: .-- ..- c c c c c c c c c c c c c c c c c c c c c c 3 3 3 3 3 3 3 3 3 3 3 3 3 3 3 3 § § 3U 3 3 o E 3 3 3 3 < Q.
< -O Q. Q.
CL ε C ε ε ε ε ε CL CL CL E E L ro O 00 n n- CM 0 ro vo in ro o Is- ro ro VD VD rr rr 00 Is-. VD (N O rr rr o 00 ro 00 co (N 10 in Is- ro ro rf ιn rr co rr Is- 0s. 0s. ro 0 10 0s. D rr CM D 00 VD D ro o (M co in ro VO 0s. ro Lθ OO VD co ro cn CM CM ro rf in VD Is- IV 00 ro o 1-1 CM o ro rf in vD D 00 OO O cn o (M LO IS) T) LO LO in in vo o VD VD vo vD VD vD vD vD vo VD vo Is- Is-
00 CO o r- vo I
s- rH rr cn no in co ro cn ro in VD I
s- in CN in cn CM CM in π- 0 ro rr ro CM CO in VD I
s- ro IH in VD ro o ro (Tι D VD I
s- P- rH r-. rf ro r-. VD 0 I
s- 0
s. VD in rr LO ΓM vo in I
s- O ro VD co 0
s. in 1-H CM 00 ro VD r 00 00 0
s. o IN ro rf rf m VD I
s- co 00 cn in in in in in in in in in in in vo vo VD vo vD vo vo vo VD vo VD vo D I
s- I
s- in VD r 00 cn o (N ro rf in VD I
s- 00 cn O -t CM CO rr in VO r- 00 cn o rr rf rr rr rf in m in m in in in m in in VD VD VD VD VD VD VD vD vD vo I
s- 0 0 0 o O o o o o o o o o o o O O O O O O O O 0 0 0 0 0 0 o O o o o o o o o o o o O O O O O O O O 0 0 0 CL C CL CL O. α. Q. CL CL CL O- L CL CL CL Q. Q. CL CL CL Q. Q. CL CL CL CL α Q. α 0. O- CL CL Q. Q. L o. o. CL CL CL CL CL CL CL co
¬ CL CL CL CL CL CL CM ro rr (N ro rr 0
s. CO r rr m VD in 0 cr> CO r- vD in rr ro (N (N ro n- CM IN (N LO m in m CO CO 00 cn cn n ro CM fN CM (N cn cn cn LO in in LO O ro CO vD D VD (M ro ro ro m ro ro ro ro CO VD VD VD O ro CO in in in in rr rr rf m in in VD in in in in in in in in in in m in IN (N (N (N ro ro ro CM TH o cn I
s- V0 in O rH (N rf r rr r ro ro 00 ro cn cn cn O O O O o o o O rf rf rr rr rr rr rf rr rr rf n- r n- rr
00 c 3 JO 3 cn c c c c c c c c c c c c c , , c c c c c c 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 ^ 5 5 5 5 5 5 5 o o o o o o o o o o o o o o o ? o o o o o o o c c c c c c c c c c c c c *
•£ C C C C C Z CZ c c C C C C C Z Z Z Z Z CZ CZ 5) c c C C CZ Z __3 3 3 3 3 3 3 3 3 3 3 3 3 3 3 (0 3 3 3 3 3 3 3 LJ U 3 C υ x Φ O 0_ ι_ CL — > φ JO 3 CL 3
CL CL CL CL CL CL L CL CL ε E ε CL CL E CL CL ε ε CL CL CL
LO TH co cn rr co o co rr oo oo oo oo rr in oo rH r vo oo Is- <o> cn o in in ro m in vD cn OO O CO ΓM O ΓM OO IS- rr o cn oo rr ro cn TH m n CM iN cn iN oo oo rr VD cn cn -Ή vo VD VD OO VD rr co o -rπ oo r rr in VD VD O -M (N oo oo ro rr m vo oo O ΓM IN OO rr vo r- rv r rv r rv rv oo co oo co co oo oo co oo oo cn cn cn n n cn cn
TH TH in in rH O r oo in Ln rM vO rr -Η rv ΓM CO OO ΓM CM rv rv vo rr ro oo in (M vD co m cn cn in cn Ln cn in oo OM OO OO in ro m oo Cn ro VD 0s. rr in O (N OO O rr vD -rH CO r)- VD o rv ro co in oo o CM ro rr rr LO VD ΓV O i-i oo oo co rr rr in vo cn o IN CN ΓO rr vo rv rv rv rv rv r rv co co co co oo co co oo oo co cn cn cn cn cn cn TH ΓM OO rr Lo vo rv co cn o TH IN ro rr in vo rv co cn o TH ΓM ΓO rv rv r rv rv rv rv ι— rv co co co oo co oo co co to co cn cn cn cn o o o o o o o o o o o o o o o o o o o o o o o o o o o o o o o o o o o o o o o o o o o o o o CL CL CL CL CL CL CL CL l CL CL CL CL CL CL L CL CL CL CL L CL CL CL CL CL CL CL CL CL CL CL CL CL CL CL CL CL CL CL CL CL CL CL CL CL in 0s. rv VD VD o cn in VD n CO cn o n- cn 00 o (N VD Is- CO cn ( in in in (N in OO 00 ro rr rT o rH m (M in VD rH o o o o vn oo ro OO VD ΓM CO vD Is- Is- ΓM ΓM iH ΓM cn (N n OO 00 00 00 in rr rT rr in VD ro LO ro oo r r VD VO m oo O in VO in LO m in
H -rH ΓM (N ΓM ΓM ΓM rf 00 ΓM ΓM rH (N rH rH rH CM ΓM ro rH ΓM ΓM rH ΓM oj o TH n co vo m o VD rv co rH o rv co oo ΓM o co o rn ro LO VO
-S en r oo oo cn o rv rn co co rv rv co co Lo CJJ Ln rM cn ΓM IN IM IN
»Q rf rr rr rr rr vo in in ro oo rM rM oo oo cn vo cn o oo o o o o eβ rr ΓM ΓM ΓM rj- in H rr rH rH rM rM in m ro -ΓH oo in m rr rr rr rr
H
Table XIV 1334.3 3707 Ipp0094 97375 98142 P unknown 6 1335.2 3708 Ipp0095 98132 99484 m unknown 6 4030.1 5310 Ipp0096 99743 100201 P unknown 6 4032.4 5311 Ipp0097 100198 101145 m unknown similar to unknown proteins 5.2 5391.3 6113 Ipp0098 101142 102665 m unknown similar to unknown proteins 5.2 5996.1 6316 Ipp0099 102655 103182 m unknown Putative secreted protein 5.2 531.4 6077 IppOlOO 103432 105357 P unknown Putative membrane protein 6 530.3 6074 IppOlOl 105323 105913 m unknown 6 Similar to arginine-binding 1356.2 3717 Ipp0102 106140 106868 m unknown periplasmic protein 1.2 1357.2 3718 Ipp0103 106959 107405 m unknown conserved hypothetical protein 5.2 4948.4 5915 Ipp0104 107516 111490 P unknown 6 2299.3 4263 Ipp0105 111711 112175 P unknown 6 2301.2 4264 Ipp0106 112571 114751 P rnr/lidP unknown Similar to exoribonuclease RNase R 2.3 3049.1 4708 Ipp0107 114744 115493 unknown similar to RNA methyltransferase 3.6 Highly similar to ribose 5- 3047.2 4707 Ipp0108 115486 116136 P unknown phosphate isomerase RpiA 2.1.1 3045.3 4706 Ipp0109 116200 117579 P unknown Similar to 5 -nucleotidase 2.3 3041.1 4705 IppOllO 117963 119156 P unknown 6 similar to unknown protein, 559.2 6204 IppOlll 119322 119651 P unknown truncated 5.2 Weakly similar to putative 558.2 6198 Ipp0112 119766 120218 P unknown response regulator 3.5.2 557.2 6196 Ipp0113 120248 122938 m polA DNA polymerase I 3.1 similar to UDP-3-0-(R-3- 1844.3 4026 Ipp0114 122941 123996 m IpxD unknown hydroxymyristoyl)-glucosamine N- acyltransferase 1.1 1845.3 4027 Ipp0115 124260 125042 P + unknown 6 3-oxoacyl-[acyl-carrier-protein] 1846.3 4028 Ipp0116 125039 126268 m fabF unknown synthase 2.4 1022.2 3521 Ipp0117 126393 127253 m unknown similar to acetyltransferase 2.1.1 similar to peptide methionine 1023.2 3522 Ipp0118 127372 127947 m unknown sulfoxide reductase msrA 4.2 Similar to conserved hypothetical 1024.2 3523 Ipp0119 128055 128651 m unknown protein, predicted membrane protein 5.2
Table XIV similar to putative xanthine/uracil 1451.4 3777 Ipp0120 128974 130254 m unknown permeases 1.2 Similar to conserved hypothetical 1859.2 4034 Ipp0121 130272 131078 m unknown protein 5.2 1858.1 4033 Ipp0122 131177 131446 m unknown 6 1856.3 4032 Ipp0123 132053 133855 P unknown 6 Similar to farnesyl-diphosphate 4574.2 5677 Ipp0124 133855 134892 p unknown famesyltransferase (Squalene synthetase) 2.4 1160.3 3610 Ipp0125 135178 136374 m Unknown Similar to transposase (IS4 family) 4.5 4881.4 5879 Ipp0126 136692 139913 m unknown Ankyrin repeat protein 4.6 Similar to predicted signal 1331.3 3705 Ipp0127 140150 141358 m unknown transduction protein. Contains EAL motifs 5.2 glycine dehydrogenase [decarboxylating] subunit 2 1332.3 3706 Ipp0128 141599 143053 m gcsB unknown (glycine decarboxylase) (glycine cleavage system P-protein) 2.2 Similar to conserved hypothetical 3626.1 5066 Ipp0129 143047 143355 m unknown protein 5.2 glycine dehydrogenase [decarboxylating] subunit 1 3625.2 5065 Ipp0130 143355 144725 m gcsA unknown (glycine decarboxylase) (glycine cleavage system P-protein) 2.2 Similar to glycine cleavage system 3623.2 5064 Ipp0131 144728 145105 m gcvH unknown H protein 2.2 Similar to glycine cleavage system 3622.4 5063 Ipp0132 145139 146221 m gcvT unknown protein T 2.2 4977.4 5932 Ipp0133 146344 146823 P unknown 6 Some similarity with L. 4974.1 5931 Ipp0134 146798 147328 m unknown pneumophila IcmL/Dotl 5.1 Similar to ABC transporter, 4793.2 5824 Ipp0135 147825 149564 P unknown permease protein 1.2 Similar to ABC transporter, ATP- 4790.3 5823 Ipp0136 149572 150870 P unknown binding protein 1.2
Table XIV thio disulfide interchange 4788.3 5822 Ipp0137 151031 151645 m + dsbA protein precursor DsbA 3.9 4980.3 5935 Ipp0138 151656 152258 m + Unknown similar to cytochrome c4 1.4 4979.1 5933 Ipp0139 152333 152935 P - unknown Similar to GTP-binding protein 4.6 2921.3 4626 Ipp0140 153049 156360 P - unknown 6 Highly similar to acetyl-CoA 327.2 4843 Ipp0141 156457 158415 m - acsB unknown synthetase 2.1.1 Similar to 3-hydroxyisobutyrate 328.1 4850 Ipp0142 158342 159238 m - unknown dehydrogenase 2.1.1 Similar to C. burnetii 330.2 4862 Ipp0143 159262 160761 m unknown methylmalonate-semialdehyde dehydrogenase MmsA 2.1.1 Weakly similar to conserved 2919.1 4624 Ipp0144 160872 161048 m unknown hypothetical protein 5.2 Some similarity with eukaryotic 1003.3 3512 Ipp0145 161226 163694 p unknown proteins, putative coiled-coil protein 6 similar to dihydrodipicolinate 1005.2 3513 Ipp0146 163798 164529 m dapB unknown reductase proteins 2.2 hypothetical, similar to 6425.1 6380 Ipp0147 164709 164894 m Unknown hypothetical proteins 5.2 Similar to N-terminal part of ProQ, 2917.1 4623 Ipp0148 165251 165943 p unknown activator of ProP osmoprotectant transporter (truncated?) 3.5.2 2360.2 4301 Ipp0149 165944 167014 p unknown Similar to hypothetical protein 5.2 SdhB protein, substrate of 2361.4 4302 Ipp0150 167114 172741 m sdhB the Dot/Icm system 5.1 Similar to pyruvate kinase II PykA, 3568.3 5025 Ipp0151 172833 174257 m unknown glucose stimulated 2.1.2 3566.1 5024 Ipp0152 174241 175431 m pgk phosphoglycerate kinase 2.1.2 glyceraldehyde 3- 712.2 6419 Ipp0153 175442 176434 m gap phosphate dehydrogenase 2.1.2 711.2 6418 Ipp0154 176515 178521 m tkt unknown Similar to transketolase 2.1.2 2335.2 4282 Ipp0155 178779 180113 P unknown 6 2336.2 4283 Ipp0156 180177 182228 P prlC Oligopeptidase A 2.2
Table XIV 3563.3 5023 Ipp0158 183008 183355 P unknown 5.2 9.2 6521 Ipp0159 183169 183672 P unknown 5.2 Similar to Wolinella succinogenes 7.1 6412 Ipp0160 183831 187949 unknown hypothetical protein 5.2 Similar to Wolinella succinogenes 4.1 5288 Ipp0161 187946 188938 unknown hypothetical protein 5.2 Similar to Wolinella succinogenes 3.1 4683 Ipp0162 188945 189244 unknown hypothetical protein 5.2 Similar to Wolinella succinogenes 2.1 4112 Ipp0163 189225 189818 unknown hypothetical protein -RecB family eλui iuv.it: d-ae 5.2 59.1 6298 Ipp0164 192785 193228 P unknown similar to unknown proteins 5.2 58.1 6273 Ipp0165 193218 193901 m prpA unknown Similar to putative phage repressor 3.5.2 Legionella vir region 57.1 6255 Ipp0166 194054 194923 P IvrA protein 5.1 Legionella vir region 55.1 6161 Ipp0167 194865 195251 P IvrB protein 5.1 Legionella vir region similar to carbon storage regulator 52.1 6035 Ipp0168 195264 195467 P IvrC protein gene csrA 3.5.2 Legionella vir homologue 50.1 5945 Ipp0169 195464 195766 P lvhB2 protein 1.8 Legionella vir homologue 48.1 5827 Ipp0170 195776 196057 P lvhB3 protein 1.6 Legionella vir homologue 47.1 5756 Ipp0171 196044 198524 P lvhB4 protein 1.6 Legionella vir homologue 46.1 5692 Ipp0172 198521 199231 P lvhB5 putative traC protein protein B5 4.5 Legionella vir homologue putative component of the type IV 9001.1 6522 Ipp0173 199246 199392 P lvhB7 protein secretion system 1.6 Legionella vir region 45.1 5629 Ipp0174 199389 199784 P IvrD protein 5.1 Legionella vir homologue 44.1 5568 Ipp0175 199788 200828 P lvhB6 protein 1.6 Legionella vir homologue 43.1 5494 Ipp0176 200825 201541 P lvhB8 protein 1.6
Table XIV 42.1 5432 Ipp0177 Legionella vir homologue 201538 202290 P lvhB9 protein 1.6 Legionella vir homologue 39.1 5227 Ipp0178 202287 203378 P IvhBlO protein 1.6 37.1 5115 Legionella vir homologue Ipp0179 203380 204384 P IvhBll protein 1.6 Legionella vir homologue putative component of type IV 36.1 5046 IppOlδO 204377 206278 P lvhD4 protein secretion system 1.6 4975 Legionella vir region 35.1 IppOlδl 206376 207173 P IvrE protein 5.1 34.1 4916 Ipp0182 207419 208168 P unknown 6 33.1 4861 Ipp0183 208292 Highly similar to L.pneumophila 210931 m unknown TraA-like protein 4.5 30.1 4684 Ipp0184 211180 211386 P unknown 6 29.1 4611 Ipp0185 211373 211642 P unknown some similarity with TraD protein 4.5 27.1 4506 Ipp0186 211736 212383 P unknown similar to unknown protein 5.2 26.1 4457 Ipp0187 212387 213196 P unknown similar to unknown proteins 5.2 25.1 4399 Ipp0188 Similar to cytosine/adenosine 213223 213684 m unknown deaminases 2.3 similar to putative membrane 24.1 4325 Ipp0189 213677 215104 m unknown proteins 5.2 20.1 4113 Ipp0190 215097 215456 m unknown 6 Similar to transcriptional 19.1 4061 pp0191 215473 216120 m repressors (RecA-mediated unknown autopeptidases) and prophage repressors 3.5.2 18.1 4000 pp0192 216289 216528 unknown 6 3869 Similar to putative phage 16.1 pp0193 216559 216765 unknown excisionase 4.4 15.1 3803 pp0194 216714 217922 unknown Similar to phage integrase 4.4 13.1 3688 pp0195 218149 218973 unknown Similar to hypothetical protein 5.2 11.2 3573 pp0196 218960 219922 unknown Similar to hypothetical protein 5.2 Similar to adenine specific DNA 2622.1 4468 pp0197 220253 222193 unknown methylase (Mod-related) 3.2 similar to Type III restriction- 2605.3 4461 pp0198 222190 224688 unknown modification enzyme, helicase subunit 3.2 2620.1 4467 pp0199 224863 225129 unknown Similar to transposase 4.5
Table XIV 2619.1 4465 Ipp0200 225090 225545 P unknown similar to transposase 4.5 2616.1 4464 Ipp0201 225539 226405 P unknown Similar to unknown proteins 5.2 6117.2 6335 Ipp0202 226596 229781 P unknown Ankyrin repeat protein 6 577.2 6270 Ipp0203 230009 230875 P Unknown Similar to unknown proteins 5.2 Similar to conserved hypothetical 3729.1 5130 Ipp0204 230880 231446 m unknown proteins 5.2 3730.1 5131 Ipp0205 231770 232150 m unknown 6 Similar to C-terminal part of phage 3731.1 5132 Ipp0206 232229 232591 m unknown integrase 4.4 Similar to conserved hypothetical 3732.1 5133 Ipp0207 233168 234034 P unknown protein 5.2 Some similarity with nucleoside 3734.1 5134 Ipp0208 234326 235429 P unknown hydrolase 2.3 3735.2 5135 Ipp0209 235729 237378 P unknown Similar to N-terminal part of sidC 5.1 6178.1 Similar to transposase (ISL3 6343 Ipp0210 237675 238718 P Unknown family) 4.5 similar to unknown proteins, 4421.2 5579 Ipp0211 238709 239518 unknown possibly interrupted by an IS element 5.2 4422.1 5580 Ipp0212 239615 239896 P unknown hypothetical gene 6 4424.1 5581 Ipp0213 240007 240228 P unknown 6 4425.1 5582 Ipp0214 240269 240784 P unknown Similar to hypothetical protein 5.2 4427.1 5583 Ipp0215 241233 241457 m unknown 6 4429.2 5584 Ipp0216 241674 241988 m unknown hypothetical protein 6 1827.3 4017 Ipp0217 242283 243719 P unknown 6 1826.2 4016 Ipp0218 243980 244726 P unknown 6 4920.4 5903 Ipp0219 245074 245952 m Unknown regulatory protein (GGDEF domain) 1.3 1302.2 3690 Ipp0220 245982 247064 m Unknown regulatory protein (EAL domain) 1.3 Similar to type I methionine 1303.3 3691 Ipp0221 247179 247940 m unknown aminopeptidase proteins 3.8 Similar to conserved hypothetical 3570.2 5027 Ipp0222 247921 248133 m unknown protein 5.2 2392.2 4323 Ipp0223 248310 248789 m unknown Predicted membrane protein 6 2391.2 4322 Ipp0224 249025 250002 P unknown 6 Similar to conserved hypothetical 3572.1 5028 Ipp0225 250131 250550 m unknown protein 5.2
Table XIV Similar to conserved hypothetical 1578.3 3852 Ipp0226 250975 251748 P unknown protein 5.2 Similar to conserved hypothetical 1579.4 3853 Ipp0227 251830 252285 m unknown protein 5.2 similar to conserved hypothetical 5044.2 5973 Ipp0228 252453 253337 m unknown protein 5.2 5046.2 5974 Ipp0229 253604 254014 m unknown similar to unknown proteins - 5.2 5047.3 5975 Ipp0230 254115 254840 m unknown Similar to hypothetical protein 5.2 Similar to C-terminal part of 2739.3 4531 Ipp0231 254837 255523 m unknown paraquat-inducible protein 5.2 Similar to hypothetical ABC 2738.2 4530 Ipp0232 255591 256346 m unknown transporter (permease) 1.2 2737.2 4529 Ipp0233 256612 257178 m unknown Protein with a F-box domain 6 190.3 4062 Ipp0234 257395 258093 P unknown 6 Similar to transcriptional regulator 191.2 4068 Ipp0235 258223 259083 m unknown (LysR family) 3.5.2 Similar to pyoverdine biosynthesis 193.4 4079 Ipp0236 259194 260243 P unknown protein PvcA 2.1.1 Similar to pyoverdine biosynthesis 195.3 4092 Ipp0237 260233 261069 P unknown protein PvcB 2.1.1 similar to FAD monooxygenase, 197.3 4100 Ipp0238 261066 262508 P unknown PheA/TfdB family 2.1.1 199.1 4108 Ipp0239 262505 263653 P unknown some similarity with transporters 1.2 201.2 4116 Ipp0240 263650 264882 P unknown similar to hypothetical protein 5.2 2030.2 4127 Ipp0241 265040 266182 m unknown 6 some similarity with 2041.2 4133 Ipp0242 266605 267696 P unknown methyltransferases 4.6 2039.1 4131 Ipp0243 267801 268619 P unknown 6 Similar to protease heat shock 494.2 5911 Ipp0244 268794 269636 m htpX unknown protein 4.1 496.1 5922 Ipp0245 269845 270756 P unknown 6 497.1 5929 Ipp0246 270803 271255 m unknown 6 Similar to conserved hypothetical 498.5 5934 Ipp0247 271666 273144 P unknown protein 5.2 2037.3 4130 Ipp0248 273404 274813 P unknown Similar to Zn metalloprotein 2.2 Similar to predicted acyl-CoA 2733.1 4528 Ipp0249 274940 275992 P unknown transferases 2.4 2732.1 4527 Ipp0250 276189 277055 m + unknown 6
Table XIVr 2730.1 4526 Ipp0251 277434 278387 P - unknown 6 1188.2 3624 Ipp0252 278622 280871 m + katG catalase-peroxidase 4.2 1930.3 4080 Ipp0253 280991 281995 m - unknown 6 2049.2 4135 Ipp0254 282282 283535 m - unknown 6 similar to conserved hypothetical 2728.1 4524 Ipp0255 283927 284244 P - unknown protein 5.2 2727.2 4523 Ipp0256 284496 287144 P - unknown 6 Similar to chitin-binding protein 1213.2 3638 Ipp0257 287559 288695 P + unknown CbpD 2.1.1 1211.3 3637 Ipp0258 289073 290107 P + unknown 6 similar to cytochrome d ubiquinol 930.3 6545 Ipp0259 290220 291590 P - qxtA unknown oxidase subunit I 1.4 similar to cytochrome d ubiquinol 932.1 6546 Ipp0260 291590 292579 P - qxtB unknown oxidase subunit II 1.4 similar to conserved hypothetical 935.2 6547 Ipp0261 292580 293365 m unknown protein 5.2 936.3 6548 Ipp0262 293362 293898 m unknown Putative membrane protein 6 2769.2 4548 Ipp0263 293888 295675 m unknown Similar to hypothetical protein 5.2 2770.1 4550 Ipp0264 295758 296456 P unknown similar to oxydoreductase 4.6 similar to unknown protein (N- 2771.2 4551 Ipp0265a 296473 297642 P unknown terminal part) 5.2 similar to unknown protein (C- 5926.1 6307 Ipp0265b 297776 298012 P unknown terminal part) 5.2 2772.1 4552 Ipp0266 298017 298430 m unknown 5.2 similar to serine/threonine-protein 657.3 6389 Ipp0267 298794 301679 unknown kinase (conserved domain) 3.8 656.3 6388 Ipp0268 301698 303668 P unknown 6 350.3 4976 Ipp0269 303748 304335 P unknown 6 Similar to sensory protein 351.1 4984 Ipp0270 304477 304923 m unknown (eukaryotic) 4.6 similar to deoxyribodipyrimidine 354.1 5005 Ipp0271 304984 306399 m unknown photolyase phrB 3.2 357.2 5026 Ipp0272 306494 307201 m unknown similar to membrane protein LrgB 5.2 Predicted membrane protein, 359.2 5037 Ipp0273 307179 307568 m unknown similar to conserved hypothetical protein LrgA 5.2
Table XIV similar to transcriptional regulator, 2774.1 4553 Ipp0274 307674 308558 p unknown lysR family 3.5.2 Similar to conserved hypothetical 2775.1 4554 Ipp0275 308681 308893 p unknown protein 5.2 Phosphoribosylaminoimidaz 2777.1 4555 Ipp0276 308979 310058 m purK ole carboxylase, ATPase subunit 2.3 Phosphoribosylaminoimidaz 2778.2 4556 Ipp0277 310055 310555 m purE ole carboxylase catalytic subunit 2.3 2780.3 4558 Ipp0278 310615 311454 m - unknown similar to unknown protein 5.2 similar to conserved hypothetical 2781.1 4559 Ipp0279 311479 311832 m - unknown protein 5.2 1649.4 3901 Ipp0280 311846 314134 m - unknown similar to unknown protein 5.2 Similar to NADH-ubiquinone 1650.2 3902 Ipp0281 314149 315669 m - unknown oxidoreductase chain 5 1.4 similar to transcriptional regulator, 2587.2 4449 Ipp0282 315771 316622 P - unknown lysR family 3.5.2 Similar to toxin secretion ABC 1940.2 4086 Ipp0283 316664 318253 P + unknown transporter ATP-binding protein 1.2 Similar to RND efflux membrane 1943.3 4087 Ipp0284 318250 319236 P + unknown fusion proteins 1.2 similar to outer membrane 4561.2 5669 Ipp0285 319233 320630 unknown component of multidrug efflux pump 1.2 4560.2 5668 Ipp0286 320813 321919 P - unknown 6 4559.1 5667 Ipp0287 322090 323655 m + unknown Similar to amino acid transporters 1.2 4558.1 5666 Ipp0288 323803 324636 m - unknown 6 6072.1 6330 Ipp0289 324863 324988 P - unknown hypothetical gene 6 6071.1 6329 Ipp0290 324918 325130 P unknown similar to unknown protein 5.2 Similar to 3-oxacyl-(acyl-carrier- 4553.2 5665 Ipp0291 325750 326448 P + unknown protein) reductase 2.4 Similar to proline/betaine 5146.2 6022 Ipp0292 326541 327836 P + unknown transporter ProP 1.2 Similar to conserved hypothetical - 3116.2 4752 Ipp0293 327811 328203 m - unknown protein 5.2 cytochrome o ubiquinol 3114.2 4751 Ipp0294 328441 329259 P - cyoA oxidase subunit II 1.4
Table XIV cytochrome o ubiquinol 3113.1 4750 Ipp0295 329262 331256 P cyoB oxidase subunit I 1.4 Cytochrome o ubiquinol 3110.1 4749 Ipp0296 331256 331852 P cyoC oxidase subunit III cyoC 1.4 3109.2 Cytochrome o ubiquinol 4748 Ipp0297 331854 332192 P cyoD oxidase subunit IV 1.4 6263.1 6357 Ipp0298 332287 332469 m unknown Hypothetical gene 6 3106.2 4747 Ipp0299 332712 334541 m Unknown regulatory protein (GGDEF domain) 1.3 3105.2 4746 Ipp0300 334546 334842 m unknown 6 3104.2 4745 Ipp0301 334988 337546 m unknown similar to cation transport ATPase 1.2 Similar to Fe2+/Zn2+ uptake 3102.2 4744 Ipp0302 337878 338324 P unknown regulation proteins 3.5.2 3101.1 4743 Ipp0303 338633 340444 P unknown 6 m SidE protein, substrate of 514.5 6021 Ipp0304 340555 345045 sidE the Dot/Icm system 5.1 Similar to conserved hypothetical 2150.1 4193 Ipp0305 345463 345945 m unknown protein 5.2 1222.5 3642 Ipp0306 346040 348019 m unknown 6 2315.3 4271 Ipp0307 348301 349095 P unknown Similar to hydrolase 2.1.1 similar to betaine aldehyde 2317.2 4272 Ipp0308 349112 350578 P unknown dehydrogenase BetB 4.1 similar to 4-aminobutyrate 2804.1 4569 Ipp0309 350553 351905 P gabT unknown aminotransferase 2.2 2279.2 4252 Ipp0310 351994 352752 P unknown 6 1722.2 3948 Ipp0311 353016 353948 P unknown similar to glutaminase 2.2 Similar to D-3 phosphoglycerate 1723.6 3949 Ipp0312 354072 354959 P unknown dehydrogenase SerA 2.2 2806.1 4570 Ipp0313 355096 355695 m unknown Similar to dehydrogenase 4.6 670.1 6394 Ipp0314 356025 357419 m unknown similar to oxydoreductase 4.6 similar to C-terminal part of 309.2 4736 Ipp0315 357471 360830 m unknown conserved hypothetical protein 5.2 310.1 4742 Ipp0316 361030 361755 P unknown 6 312.3 4754 Ipp0317 361996 362949 P unknown 6 2807.1 4571 Ipp0318 363109 363450 P unknown similar to arsenate reductase 4.2 2434.2 4354 Ipp0319 363537 364691 m unknown similar to fatty acid desaturase 2.4
Table XIV similar to ATP-dependent RNA 2432.2 4353 Ipp0320 364810 366054 m rhlE unknown helicase RhlE 3.6 Similar to N-terminal part of 2808.1 4572 Ipp0321 366385 366660 p unknown eukaryotic RNA-binding protein precursor 4.6 Similar to unknown protein; 2424.4 4345 Ipp0322 366836 367513 p unknown putative membrane protein 5.2 similar to putative 2425.3 4346 Ipp0323 367551 368189 P unknown acetyltransferase 2.1.1 2426.2 4347 Ipp0324 368323 368850 P unknown 6 2810.1 4573 Ipp0325 369005 370351 P unknown similar to outer membrane protein 1.1 N-terminal part similar to N- 2811.2 4574 Ipp0326 370341 371378 unknown terminal part of conserved hypothetical protein 5.2 similar to multidrug resistance 365.4 5079 Ipp0327 371435 372427 unknown proteins 1.2 364.2 5073 Ipp0328 372604 374547 P unknown 6 360.3 5047 Ipp0329 374566 376350 P unknown 6 2031.2 4128 Ipp0330 376713 377027 P Unknown Similar to transposase (IS4 family) 4.5 similar to conserved hypothetical 3163.1 4778 Ipp0331 377053 377346 P unknown protein 5.2 3165.1 4779 Ipp0332 377546 377944 P unknown 6 Similar to C-terminal part of DNA 3166.1 4780 Ipp0333 378030 378632 m unknown polymerase, bacteriophage-type 3.1 3167.1 4781 Ipp0334 378927 380201 unknown Similar to transporter, MFS family 1.2 3169.1 4782 Ipp0335 380322 380978 P + unknown weakly similar to amidase 1.1 2488.2 4392 Ipp0336 381115 382095 m + unknown 6 2489.2 4393 Ipp0337 382142 382390 m + unknown 6 2490.2 4395 Ipp0338 382554 383003 P + unknown 6 4719.2 5772 Ipp0339 383108 384655 m + unknown Similar to multicopper oxidase 4.7 4718.1 5771 Ipp0340 384883 385317 P - unknown 6 Similar to magnesium and cobalt 4716.2 5770 Ipp0341 385332 386324 m - unknown transport proteins 1.2 2695.2 4503 Ipp0342 386589 387176 P - unknown Similar to predicted hydrolase 2.1.1
Table XIV similar to conserved hypothetical 2696.2 4504 Ipp0343 387311 388810 p unknown protein 5.2 Nicotinate 1564.3 3845 Ipp0344 388891 390294 p pncB phosphoribosyltransferase 2.5 nicotinamidase/ 1563.3 3844 Ipp0345 390299 390919 p pncA pyrazinamidase 2.5 similar to conserved hypothetical 2698.1 4505 Ipp0346 391002 391385 m unknown protein 5.2 Similar to transporter of the major 2700.2 4508 Ipp0347 391385 392599 m unknown facilitator superfamily (MFS) 1.2 2702.2 4509 Ipp0348 392717 393586 p unknown Similar to transcriptional regulators 3.5.2 SdbA protein, putative 326.2 4837 Ipp0349 393998 397348 p sdbA substrate of the Dot/Icm system 5.1 325.1 4831 Ipp0350 397428 398981 P Unknown Unknown 6 324.3 4823 Ipp0351 399047 399712 Unknown regulatory protein (EAL domain) 1.3 2486.3 4391 Ipp0352 399749 401272 m Unknown regulatory protein (GGDEF domain) 1.3 Similar to two-component sensor 2704.4 4510 Ipp0353 401275 402690 m unknown histidine kinase 1.3 Similar to conserved hypothetical 2705.3 4511 Ipp0354 402713 403876 m unknown protein 5.2 similar to transcriptional regulator 2706.2 4512 Ipp0355 404049 404930 m unknown lysR family 3.5.2 1183.4 3621 Ipp0356 405129 407048 P unknown protein with ankyrin motif 6 1184.2 3622 Ipp0357 407059 408243 m unknown similar to amino acid transporter 1.2 Some similarity with eukaryotic 1186.2 3623 Ipp0358 408311 409087 m unknown proteins 5.2 Similar to NAD+-dependent 189.3 4055 Ipp0359 409295 410506 P unknown formate dehydrogenase 1.4 188.1 4047 Ipp0360 410611 411735 m unknown 6 187.1 4041 Ipp0361 411972 412655 P unknown 6
Table XIV similar to oxidoreductase, short 185.1 4030 Ipp0362 412760 413533 P + unknown chain dehydrogenase/reductase family 2.1.1 184.3 4023 Ipp0363 413777 414136 P unknown hypothetical gene 6 1968.1 4099 Ipp0364 414759 414989 P + unknown signal peptide predicted 6 1966.2 4098 Ipp0365 415044 415613 m - efp unknown Similar to elongation factor P 3.7.4 similar to conserved hypothetical 2709.1 4513 Ipp0366 415685 416665 P - unknown protein 5.2 4267.5 5473 Ipp0367 416734 418806 m - ppk Polyphosphate kinase 2.6 1474.4 Similar to conserved hypothetical 3788 Ipp0368 418847 420052 m - unknown protein 5.2 Weakly similar to chromate 1473.4 3787 Ipp0369 420128 420661 m - unknown transport protein 1.2 4269.3 5474 Ipp0370 420658 421278 m - unknown 6 1908.4 4067 Ipp0371 421454 423892 P - unknown similar to acyl-CoA dehydrogenase 2.4 4272.3 5476 Ipp0372 424006 424698 P - unknown 6 similar to nucleotidyltransferase 4323.1 5509 Ipp0373 424735 425397 m - unknown family protein 1.1 Similar to conserved hypothetical 4322.1 5508 Ipp0374 425394 426371 m - unknown protein 5.2 Similar to organic solvent tolerance 4321.2 5507 Ipp0375 426614 429133 P - unknown protein 4.2 Similar to peptidyl-prolyl cis-trans 4320.1 5506 Ipp0376 429309 430598 P + surA unknown isomerase SurA 3.8 4-hydroxythreonine-4- 1251.2 3659 Ipp0377 430595 431569 P ~ pdxA phosphate dehydrogenase PdxA 2.5 dihydrofolate reductase 1250.1 3658 Ipp0378 431571 432062 P - folA FolA 2.5 1249.3 3656 Ipp0379 432069 432614 m - unknown similar to eukaryotic proteins 5.2 6247.1 6356 Ipp0380 438476 439666 P - tufA elongation factor Tu 3.7.4 Preprotein translocase secE 4883.3 5880 Ipp0381 439817 440188 P - secE subunit 1.6 transcription 4884.1 5881 Ipp0382 440191 440739 P ~ nusG antitermination protein NusG 3.5.4
Table XIV 4885.2 5882 Ipp0383 440849 441283 P rplK 50S ribosomal protein Ll l 3.7.1 5592.2 6205 Ipp0384 441293 441988 P rplA 50S ribosomal protein Ll 3.7.1 2106.2 4164 Ipp0385 442161 50S ribosomal subunit 442694 P rplJ protein L10 3.7.1 50S ribosomal subunit 2104.1 4163 Ipp0386 442725 443105 P rplL protein L7/L12 3.7.1 1106.5 3577 Ipp0387 443197 447303 P rpoB RNA polymerase B-subunit 3.5.3 RNA polymerase beta 1663.6 3912 Ipp0388 447392 451597 P rpoC subunit 3.5.3 5596.3 6207 Ipp0389 451711 452091 P rpsL 30S ribosomal protein S12 3.7.1 5595.2 6206 Ipp0390 452112 452639 P rpsG 30S ribosomal protein S7 3.7.1 4468.2 translation elongation 5604 Ipp0391 452654 454738 P fusA factor G 3.7.4 translation elongation 4466.3 5603 Ipp0392 454759 455949 P tufA factor Tu 3.7.4 4464.1 5602 30S ribosomal subunit Ipp0393 455955 456272 P rpsJ protein S10 3.7.1 50S ribosomal subunit 4463.2 5601 Ipp0394 456307 456957 P rplC protein L3 3.7.1 50S ribosomal subunit 4462.3 5600 Ipp0395 456957 457562 P rplD protein L4 3.7.1 5892.1 6293 Ipp0396 50S ribosomal subunit 457559 457855 P rplW protein L23 3.7.1 457867 50S ribosomal subunit 5894.3 6294 Ipp0397 458694 P rplB protein L2 3.7.1 30S ribosomal subunit 5895.3 6295 Ipp0398 458713 458991 P rpsS protein S19 3.7.1 1199.4 50S ribosomal subunit 3629 Ipp0399 459001 459336 P rplV protein L22 3.7.1 1201.1 3630 Ipp0400 459339 459995 P rpsC 30S ribosomal protein S3 3.7.1 1202.2 3631 Ipp0401 460012 460425 P rplP 50S ribosomal protein L16 3.7.1 50S ribosomal subunit 4133.1 5383 Ipp0402 460425 460619 P rpmC protein L29 3.7.1.
Table XIV r 4134.1 5384 Ipp0403 460621 460875 P rpsQ 30S ribosomal protein S17 3.7.1 4135.1 5385 Ipp0404 460964 461329 P rplN 50S ribosomal protein L14 3.7.1 4136.1 5386 Ipp0405 461342 461671 P rplX 50S ribosomal protein L24 3.7.1 4138.1 5387 Ipp0406 461687 462238 P rplE 50S ribosomal protein L5 3.7.1 4139.1 5388 Ipp0407 462251 462553 P rpsN 30S ribosomal protein S14 3.7.1 4140.1 5389 Ipp0408 462576 462971 P rpsH 30S ribosomal protein S8 3.7.1 50S ribosomal subunit 4141.1 5390 Ipp0409 462989 463528 P rplF protein L6 3.7.1 50S ribosomal subunit 4142.3 5391 Ipp0410 463539 463898 P rplR protein L18 3.7.1 Ipp0411 30S ribosomal subunit 4143.3 5392 463908 464414 P rpsE protein S5 3.7.1 50S ribosomal subunit 4144.3 5393 Ipp0412 464417 464602 P rpmD protein L30 3.7.1 50S ribosomal subunit 4145.3 5394 Ipp0413 464602 465036 P rplO protein L15 3.7.1 preprotein translocase. 4148.3 5395 Ipp0414 465033 466367 P secY SecY subunit 1.6 Similar to 50S ribosomal protein 9003.1 6524 Ipp0415 466384 466497 P unknown βf. 3 1 4149.1 5396 Ipp0416 466575 466931 P rpsM 30S ribosomal protein S13 3.7.1 4151.1 5398 Ipp0417 466955 467353 P rpsK 30S ribosomal protein Sl l 3.7.1 30S ribosomal subunit 4153.1 5399 Ipp0418 467370 467990 P rpsD protein S4 3.7.1 4154.2 DNA-directed RNA 5400 Ipp0419 468009 469001 P rpoA polymerase alpha chain 3.5.3 5540.1 6179 Ipp0420 469020 469403 P rplQ 50S ribosomal protein L17 3.7.1 Single-strand binding 5538.3 6178 Ipp0421 469470 469949 m ssb protein (SSB) (Helix- destabilizing protein)
Table XIV putative transport protein, MFS 5453.3 6138 Ipp0422 470033 471400 m Unknown family 1.2 similar to 3-oxoacyl-[acyl-carrier 3244.2 4827 Ipp0423 471562 472320 p Unknown protein] reductase 2.4 618.3 6344 Ipp0424 472391 472804 p Unknown similar to acyl carrier proteins 2.4 similar to hydroxymyristoyl-(acyl 619.1 6348 Ipp0425 472804 473700 p I Inknnwn carrier protein) dehydratase 2.4 621.5 6351 Ipp0426 473705 474997 P I Ink nnwn similar to 3-oxoacyl-[acyl-carrier- protein]synthase II 2.4 687.5 6404 Ipp0427 474998 476275 P 1 Inknnwn similar to 3-oxoacyl-[acyl-carrier- protein] synthase beta chain 2.4 similar to lipid A biosynthesis 688.1 6405 Ipp0428 476268 477113 P
acyltransferase 2.4 3243.1 4826 Ipp0429 477223 477528 m Unknown 6. 3242.4 4825 Ipp0430 477704 480394 m unknown 6 Diaminopimelate 5633.3 6223 Ipp0431 480705 481538 m dapF epimerase 2.2 5634.2 6224 Ipp0432 481548 481676 m unknown putative lipopeptide 1.2 5636.1 6225 Ipp0433 481835 482212 m Unknown Unknown 6. similar to 5637.1 6226 Ipp0434 482375 483034 Unknown phospholipase/carboxylesterase 2.4 similar to conserved hypothetical 4766.2 5805 Ipp0435 483031 483465 Unknown proteins 5.2 similar to conserved hypothetical 4767.2 5806 Ipp0436 483452 483724 P Unknown proteins 5.2 4768.1 5807 Ipp0437 483812 484126 m Unknown similar to other protein 5.2 Ferric uptake regulation 4769.1 5808 Ipp0438 484354 484764 P fur protein 3.5.2 similar to choline dehydrogenase 4770.1 5809 Ipp0439 484902 485045 unknown (N-terminal part) 5.2 4771.2 5810 Ipp0440 485188 486258 P Unknown regulatory protein (GGDEF domain) 1.3 5638.2 6227 Ipp0441 486569 486955 m + Unknown 6. 5377.2 6104 Ipp0442 487212 487811 P Unknown 6.
Table XIVr .. . SdhA, substrate of the 5376.2 6103 Ipp0443 487869 492140 m sdhA ' τ Dot/Icm system 5.1 similar to conserved hypothetical 3077.1 4726 Ipp0444 492303 493046 m Unknown proteins 5.2 similar to conserved hypothetical 3078.2 4727 Ipp0445 493027 494253 m Unknown proteins 5.2 similar to hypothetical protein 3080.3 4728 Ipp0446 494240 495694 m unknown modification enzyme 3.8 3081.3 4729 Ipp0447 495698 496411 m unknown 6 3082.1 4730 Ipp0448 496606 497079 m Unknown similar to hypothetical proteins 5.2 Similar to putative 3083.1 4731 Ipp0449 497076 497627 m Unknown hyperosmotically inducible periplasmic proteins 5.2 2251.3 4238 Ipp0450 497886 498344 m Unknown 6. 3085.2 4732 Ipp0451 498439 501294 P uvrA excinuclease ABC subunit A 3.2 similar to LemA from Coxiella 3086.1 4733 Ipp0452 501477 502058 P Unknown burnetii 5.2 Highly similar to C. burnetii heat 3087.1 4734 Ipp0453 502069 503088 P Unknown shock protein HtpX 4.1 Highly similar to ABC-type 3088.1 4735 Ipp0454 503115 503888 m unknown multidrug transport system, permease component 1.2 Highly similar to ATP-binding 488.2 5878 Ipp0455 503878 504792 m unknown component of ABC transporter 1.2 489.1 5886 Ipp0456 504874 504975 m Unknown 6 491.1 5895 Ipp0457 505023 506042 m Unknown 6 492.5 5902 Ipp0458 506168 506806 m unknown similar to prolyl/lysyl hydroxylase 3.8 Weakly similar to Zinc 493.4 5907 Ipp0459 506892 507599 m Unknown metalloprotease 2.2 3091.1 4737 Ipp0460 507786 508067 m Unknown 6. 1705.2 3938 Ipp0461 508216 509079 P Unknown 6 similar to methylated-DNA-protein- 1706.2 3939 Ipp0462 509180 509647 m Unknown cysteine S-methyltransferase 3.2 1707.2 3940 Ipp0463 509651 510016 m rplS 50S ribosomal protein L19 3.7.1
Table XIVr Highly similar to tRNA (guanine- 3093.1 4738 Ipp0464 510043 510795 m - trmD unknown Nl)-methyltransferase 3.6 similar to 16S rRNA processing 3095.1 4739 Ipp0465 510795 511304 m - rimM unknown protein RimM 3.6 Highly similar to 30S ribosomal 3096.1 4740 Ipp0466 511310 511570 m - rpsP unknown protein S16 3.7.1 similar to signal recognition particle 3097.3 4741 Ipp0467 511657 513033 m - ffh unknown protein Ffh 1.6 3960.2 5260 Ipp0468 513290 513952 m - Unknown 6. 3959.2 5258 Ipp0469 514226 515734 P - Unknown Ankyrin repeat protein 6. 742.3 6431 Ipp0470 516071 517474 P + Unknown similar to amino acid antiporter 1.2 741.2 6430 Ipp0471 517464 518054 P - Unknown 6. similar to conserved hypothetical 740.2 6429 Ipp0472 518258 518599 P - Unknown proteins 5.2 2213.2 4221 Ipp0473 518813 519256 m + Unknown 6. Predicted membrane protein, 3958.1 5257 Ipp0474 519275 519823 m + lidJ Unknown similar to hypothetical proteins 5.1 similar to putative transmembrane 3957.1 5256 Ipp0475 520240 520443 m + unknown proteins 5.2 similar to conserved hypothetical 3956.2 5255 Ipp0476 520511 521239 P + Unknown proteins 5.2 5524.1 6170 Ipp0477 521223 521768 P - Unknown similar to hypothetical proteins 5.2 similar to conserved hypothetical 5523.2 6169 Ipp0478 521773 522780 Unknown proteins, hypothetical cytochrome oxidase assembly protein 1.4 similar to Polyprenyltransferase 2400.2 4327 Ipp0479 522770 523654 Unknown (cytochrome oxidase assembly factor) 1.4 2401.2 4328 Ipp0480 523664 524305 P + Unknown similar to hypothetical proteins 5.2 similar to ribosomal protein S6 1476.2 3790 Ipp0481 524671 525579 P Unknown modification enzyme 3.8 1475.2 3789 Ipp0482 525583 525828 m Unknown similar to hypothetical proteins 5.2 similar to Glucose-6-phosphate 1- 1989.2 4107 Ipp0483 526144 527634 P zwf Unknown dehydrogenase 2.1.1 similar to 6- 1990.1 4109 Ipp0484 527603 528334 P pgi Unknown phosphogluconolactonase 2.1.1 similar to 6-phosphogluconate 241.2 4334 Ipp0485 528322 530160 P edd Unknown dehydratase 2.1.1
Table XIV 240.1 4326 Ipp0486 530135 531142 p glk Unknown similar to glucokinase 2.1.1 similar to 2-deydro-3-
■ 239.1 4321 Ipp0487 531129 531791 p eda Unknown deoxyphosphogluconate aldolase/4- hydroxy-2-oxoglutarate aldolase 2.1.1

238.1 4315 Ipp0488 531905 533326 p Unknown similar to sugar transport protein 1.2 similar to eukaryotic glucoamylase 235.2 4292 Ipp0489 533416 534723 m Unknown precursor (Glucan 1,4-alpha- glucosidase) 2.1.1 similar to transcriptional regulator 2969.1 4660 Ipp0490 534891 535127 m Unknown (XRE-family) 3.5.2 2970.2 4662 Ipp0491 535214 535753 m Unknown similar to hypothetical proteins 5.2 similar to Protoheme ferro- 916.3 6534 Ipp0492 535881 536876 m hemZ Unknown lyase(ferrochelatase) 2.5 917.3 6535 Ipp0493 536987 537220 m cspD unknown 4.1 similar to N-terminus of 919.1 6536 Ipp0494 537457 537870 P Unknown Diadenosine tetraphosphate (Ap4A) hydrolase 920.2 6537 Ipp0495 537973 538332 m Unknown similar to hypothetical proteins 4655.3 5728 Ipp0496 538419 539771 P Unknown similar to outer membrane proteins similar to Multidrug resistance 1353.3 3715 Ipp0497 539911 540948 P Unknown efflux pump 1354.2 3716 Ipp0498 540926 541858 P Unknown Unknown 4653.2 5727 Ipp0499 541825 542724 P Unknown similar to hypothetical proteins 4652.2 5726 Ipp0500 542746 543126 P Unknown putative transcriptional regulator 3.5.2 4432.2 5587 Ipp0501 543128 543496 m Unknown 6. 4433.1 5588 Ipp0502 543821 544432 P Unknown similar to hypothetical protein 5.2 4434.1 5589 Ipp0503 544586 545386 m Unknown ankyrin repeat protein 5.2 4435.4 5590 Ipp0504 545657 547657 P Unknown 6 451.3 5635 Ipp0505 548479 549255 m Unknown 6. 450.1 5630 Ipp0506 549621 549833 P Unknown 6. 448.1 5615 Ipp0507 550004 550264 P icmT Unknown 1.6 447.2 5606 Ipp0508 550265 550609 P icmS Unknown 1.6 444.2 5591 Ipp0509 550709 551071 P icmR Unknown 1.6 443.3 5585 IppOSlO 551162 551737 P icmQ Unknown 1.6
OO CO rr rr (N ro (M
+ + + + α α α α α α α α α α E E r ΓM m in o in rv in o rr oo ro rv 00 00 00 rr vo ro in o I
s- r- ΓM VD CO rv O (N σi rr vo m ΓM VD vo o o o rr 00 VD ro rT in 00 cn rr (N (N vo m oo ΓM rr cn r rr VD oo m vD VD I
s- 00 ΓM (N 00 r rv co TH ΓM rf LO vD vo rv cn m m in Ln m Ln vD vo vD vo vo vD vo I
s- I
s- pv I
s- rv I
s- rv rv CO Ln in m in m in in m in m in in in m in in m ιn m ιn in in r -rH o rv rv oo in vo r o-i c TH co I
s- oo co rr in rv vo CM rM Ln o oo oo rM rH rv cn o cn vD co rr o oo in IN CO r- I
s- cn o co oo vo oo rr in ro o Ln rM ro VD in vo rT cn VD σ> rH oo Ln vD vD rv θθ τH (N ro ro rr I
s- CO rH IN rT m VD vo I
s- cn m in Lri Ln m in Ln vD vD vD vD vD VD VD I
s- rv rv r- I
s- rv rv I
s- in m in Ln m in m in Ln in in in in m in in m m in in LO ιn TH rM o r in vD rv co cn o rH rM ro rr in vo r- 00 cn o τH *rH τH τH τH rH rH rH τH rM (N rM (N IN IN ΓM (N ro
J OO oo m Ln Ln Ln m Ln in in in m m Ln in in in in m in in in in in o o o o o o o o o o o o o o o o o o o o o o CL Cα. O- f-L D- CL CL CL CL θ. O- 0. CL O- L o. CL o. α. CL 0- 0- θ. CL α. o. Q. o. CL CL L CL o. CL CL Q. CL rM rH rr ro rM rH oo rM ro rr rM rr rr m (N ro r in vD (N rM o o o o cn rv rv rv rM -rH in I
s- in m in in rr rr o o Ln Ln in in ro oo oo co in Ln in LO CO o in LO in in (N fM in in vD vD vD vD vD rr rr rr Ln m ro ro rT vo VD vo vo in m
R n ^ rM rM r o r ^ ^ l ro rr ΓM oo TH ΓM
CU (N r-S VD VD rr oo ΓM ΓM ro rr rr oo IN VO 00 CO cn Is- Is- rv ΓM ΓM ΓM rv Is- IN ro O 00 rr rT rr IN ΓM
S LO Ln - S S S - r ro r 00 00 (N rr rr
03 oo oo rr rr o O OO 00 m cn cn σ> cn cn cn oo oo
Table XIV Putative lipase LipA 3814.1 5180 Ipp0533 581605 582453 m Unknown (L.pneumophila) 2.4 similar to conserved hypothetical 969.2 6567 Ipp0534 582903 583676 P Unknown proteins 5.2 Similar to fructose-bisphosphate 967.2 6566 Ipp0535 583855 584865 m unknown aldolase 2.1.2 Similar to ferredoxin— NADP 966.2 6565 Ipp0536 585015 585749 m unknown reductase 1.4 965.3 6564 Ipp0537 585833 586375 P dotV Unknown 1.6 3815.2 5181 Ipp0538 586519 586815 m Unknown conserved hypothetical protein 5.2 similar to CDP-diacylglycerol-serine 3816.2 5182 Ipp0539 587033 587770 p pssA unknown O-phosphatidyltransferase (Phosphatidylserine synthase) 2.4 similar to sugar transport PTS 3818.1 5183 Ipp0540 587999 588268 m unknown phosphocarrier protein Hpr 1.2 similar to putative sigma-54 3819.2 5184 Ipp0541 588459 588758 m Unknown modulation protein 3.5.2 RNA polymerase sigma-54 4676.2 5743 Ipp0542 588785 590179 m rpoN factor (sigma-L) 3.5.1 50S ribosomal subunit 6029.1 6321 Ipp0543 590387 590551 m rpmG protein L33 3.7.1 4673.1 5742 Ipp0544 590566 590802 m rpmB 50S ribosomal protein L28 3.7.1 similar to S-adenosylmethionine- 4672.1 5741 Ipp0545 591071 591784 p Unknown dependent methyltransferase 3.6 similar to endo-l,4-beta-glucanase 4671.3 5740 Ipp0546 591759 592883 p Unknown (hypothetical) 2.1.1 5082.3 5997 Ipp0547 592961 594448 P Unknown Protein with ankyrin domain 6. 1123.2 3587 Ipp0548 594657 595799 P hflK protease subunit HflK 2.2 membrane protease 1122.1 3586 Ipp0549 595802 596716 P hflC subunit HflC 2.2 Adenylosuccinate 2149.4 4191 Ipp0550 596851 598146 purA synthetase (IMP— aspartate ligase) (AdSS) (AMPSase) 2.3 5068.3 5990 Ipp0551 598367 598936 Unknown similar to hypothetial proteins 5.2
Table XIV similar to transcriptional regulator 5069.3 5991 Ipp0552 599136 599594 m Unknown of arginine metabolism 3.5.2 similar to putative glutamine- 5829.2 6284 Ipp0553 599725 600459 p Unknown binding periplasmic protein precursor 2.2 similar to amino acid ABC 4522.4 5644 Ipp0554 600456 601103 p Unknown transporter permease 1.2 similar to amino acid (glutamine) 4521.1 5643 Ipp0555 601087 601755 p unknown ABC transporter (ATP-binding protein) 1.2 4520.1 5642 Ipp0556 601752 602969 p argG Argininosuccinate synthase 1479.3 3791 Ipp0557 602962 604197 p argH Argininosuccinate lyase 2.2 Ornithine 1481.3 3793 Ipp0558 604200 605315 p argF carbamoyltransferase 2.2 1838.2 4021 Ipp0559 605404 606879 m Unknown similar to adenosine deaminase 2.3 similar to ABC-type branched-chain 4647.2 5722 Ipp0560 607054 608169 p Unknown amino acid transport systems, periplasmic component 1.2 similar to carboxy-terminal 4535.2 5655 Ipp0561 608242 609579 m + unknown protease family protein similar to membrane-bound 4536.1 5656 Ipp0562 609660 610802 m + Unknown metallopeptidase 2.2 Highly similar to phosphoglycerate 4538.2 5657 Ipp0563 610789 612333 m gpml unknown mutase proteins 2.1.2 4540.2 5659 Ipp0564 612436 612732 m + Unknown 6 207.3 4148 Ipp0565 612809 614080 m + Unknown putative phospholipase C Undecaprenyl 206.3 4141 Ipp0566 614430 615146 P - uppS pyrophosphate synthetase 1.1 phosphatidate 205.1 4136 Ipp0567 615157 615954 P + cdsA cytidylyltransferase 2.4 similar to putative membrane- 204.1 4132 Ipp0568 615987 617339 P _ ecfE Unknown associated Zn-dependent protease EcfE 2.2
Table XIV 202.3 4120 Ipp0569 617510 619822 + similar to protective surface Unknown antigen 5.2 732.3 6428 Ipp0570 619936 620436 + similar to putative outer membrane Unknown proteins 1.1 UDP-3-0-[3- hydroxymyristoyl] 731.2 6427 Ipp0571 620450 621460 IpxD glucosamine N- acyltransferase 1.1 (3R)-hydroxymyristoyl- 730.1 6426 Ipp0572 621588 622040 fabZ [acyl carrier protein]dehydratase 1.1 UDP-N-acetylglucosamine 729.1 6425 Ipp0573 622037 622807 IpxA acyltransferase 1.1 Similar to integral membrane 728.2 6424 Ipp0574 622811 623215 Unknown protein possibly involved in chromosome condensation 3024.1 4701 Ipp0575 623237 624517 P serS unknown Seryl-tRNA synthetase 3.7.2 3023.1 4700 Ipp0576 624685 624879 P Unknown 6. 3022.1 4699 Ipp0577 625050 625313 P Unknown 6 Weakly similar to eukaryotic 1268.2 3668 Ipp0578 625653 626585 P Unknown phytanoyl coA dioxygenase 5.2 1267.3 3667 Ipp0579 626621 627433 P Unknown 6 similar to oxidoreductase,aldo/keto 1265.5 3666 Ipp0580 627423 628253 m Unknown reductase family,related to diketogulonate reductase 2.1.1 3018.1 4698 Ipp0581 628692 629540 m Unknown 6 3017.1 4697 Ipp0582 629527 629817 m Unknown 6. 3016.1 4696 Ipp0583 629878 630369 m Unknown 6. 3015.1 4695 Ipp0584 630329 631087 m Unknown similar to methyltransferase 3.2 3014.1 4694 Similar to adenosine Ipp0585 631053 631661 m Unknown phosphosulfate (APS) kinase 2.3 similar to acetyltransferase, GNAT 3013.2 4693 Ipp0586 631658 632125 m Unknown family 2.1.1 weakly similar to C.bumetii 3011.2 4692 Ipp0587 632122 633120 m Unknown hypothetical protein 5.2
Table XIV similar to P.aeruginosa probable 5288.1 6069 Ipp0588 633117 633527 m unknown fosfomycin resistance protein 4.2 similar to aminoglycoside 6 -N- 5289.2 6070 Ipp0589 633524 634087 m Unknown acetyltransferase 4.2 5312.2 6079 Ipp0590 634498 635124 P IvgA Unknown Unknown virulence protein 5.1 5313.1 6080 Ipp0591 635154 635531 m Unknown 6. similar to conserved hypothetical 4897.2 5890 Ipp0592 635681 636820 m Unknown protein 5.2
4895.1 5889 Ipp0593 637141 637515 p + succinate dehydrogenase, sdhC cytochrome b556 subunit succinate dehydrogenase, 4894.1 5888 Ipp0594 637509 637856 p sdhD hydrophobic membrane anchor protein succinate dehydrogenase 1945.4 4088 Ipp0595 637858 639627 p sdhA flavoprotein subunit .3 succinate dehydrogenase, 1946.2 4089 Ipp0596 639640 640362 p sdhB iron sulfur protein 2-oxoglutarate 1157.4 3607 Ipp0597 640420 643230 p sucA dehydrogenase, El subunit dihydrolipoamide 3468.2 4958 Ipp0598 643266 644495 p sucB succinyltransferase, E2 subunit .3 succinyl-CoA synthetase, 1559.2 3841 Ipp0599 644558 645721 p sucC beta subunit succinyl-CoA synthetase, 1557.3 3840 Ipp0600 645796 646671 p sucD alpha subunit Similar to pyridoxine 5 -phosphate 2356.2 4298 Ipp0601 647094 647741 p pdxH unknown oxidase 2.5 transmission trait enhancer 1786.2 3991 Ipp0602 648354 648725 p letE protein LetE 4.6 1784.2 3990 Ipp0603 648796 649185 m Unknown 6
Table XIV similar to putative transport 1782.3 3989 Ipp0604 649849 651117 P Unknown proteins 1.2 C-terminal part similar to unknown 5316.1 6081 Ipp0605 651345 652802 P Unknown virulence protein 5.2 5123.3 6014 Ipp0606 652992 653273 P Unknown similar to DNA-binding proteins Fis 3.5.2 Ribose-phosphate 5127.5 6015 Ipp0607 653335 654282 m prs pyrophosphokinase 2.3 similar to putative outer membrane 3982.2 5277 Ipp0608 654664 655263 m Unknown I ipo proteins 1.2 similar to phosphopantetheine 3980.1 5276 Ipp0609 655326 655832 P Unknown adenylyltransferase 2.5 similar to Gamma- 3979.3 5274 Ipp0610 655822 657546 P unknown glutamyltranspeptidase 4.1 weakly similar to D-amino acid 3977.2 5273 Ipp0611 657644 659860 m Unknown dehydrogenase, C-terminal cAMP binding motif 2.2 similar to l-acyl-sn-glycerol-3- 1494.2 3800 Ipp0612 660069 660800 m Unknown phosphate acyltransferase 2.4 1495.1 3801 Ipp0613 660869 661189 m sugE unknown similar to suppressor of groEL 3.5.2 Similar to DNA damage inducible 1497.2 3802 Ipp0614 661528 662595 m dinP unknown protein P 3.2 5317.2 6082 Ipp0615 663035 663655 m Unknown similar to hypothetical proteins 5.2 similar to formamidopyrimidine- 1606.2 3874 Ipp0616 663730 664554 m mutM unknown DNA glycosylase MutM 3.2 1607.2 3875 Ipp0617 664554 664784 m Unknown Hypothetical protein 6. 1608.2 3876 Ipp0618 664965 666152 P unknown similar to fatty acid desaturase 2.4 4597.1 5690 Ipp0619 666193 666591 m Unknown 6. similar to acetoacetyl-CoA 4598.2 5691 Ipp0620 666857 667597 P Unknown reductase similar to acetoacetyl-CoA 4729.3 5779 Ipp0621 667776 668522 P Unknown reductase 4727.1 5778 Ipp0622 668652 669050 P Unknown similar to hypothetical proteins 4726.1 5777 Ipp0623 669257 669550 P Unknown 4725.2 5776 Ipp0624 669796 670866 m Unknown similar to hypothetical proteins similar to Bacillus subtilis spore 4724.2 5775 Ipp0625 670936 671541 P Unknown maturation protein A
Table XIV similar to uncharacterized membrane protein, similar to 4721.2 5774 Ipp0626 671538 672071 Unknown Bacillus subtilis spore maturation protein B 1.1 similar to membrane proteins 4878.2 5877 Ipp0627 672371 673804 m Unknown related to metalloendopeptidases 2.2 2859.2 4597 Ipp0628 674061 675266 P tyrS tyrosyl-tRNA synthetase 3.7.2 Similar to conserved hypothetical 693.1 6406 Ipp0629 680614 681150 m unknown protein 5.2 694.3 6407 Ipp0630 681157 682515 m gor Glutathione reductase 4.1 similar to adenosine deaminase 2747.1 4532 Ipp0631 682746 683726 m add unknown protein 2.3 2748.1 4533 Ipp0632 684108 684554 P unknown 6 2749.4 4534 Ipp0633 684595 685848 m unknown Similar to phosphate permease 1.2 Similar to conserved hypothetical 1760.4 3974 Ipp0634 685858 686529 m unknown protein 5.2 1759.4 3972 Ipp0635 686720 687484 P unknown Similar to hypothetical protein 5.2 Similar to conserved hypothetical 3591.2 5038 Ipp0636 687786 688349 P unknown protein 5.2 Similar to conserved hypothetical 3592.1 5039 Ipp0637 688342 688761 P unknown protein 5.2 Similar to aspartate 3593.1 5040 Ipp0638 688811 689704 P pyrB unknown carbamoyltransferase 2.3 3594.1 5041 Ipp0639 689818 690201 m unknown 6 Similar to competence protein 2421.2 4342 Ipp0640 690313 691824 m unknown comM 5.2 Similar to conserved hypothetical 3595.1 5042 Ipp0641 691889 692149 m unknown protein 5.2 3596.1 5043 Ipp0642 692294 692632 P glnB Nitrogen regulatory protein 2.2 similar to 5-formyltetrahydrofolate 3597.4 5044 Ipp0643 692642 693223 m unknown cyclo-ligase 2.5 6242.1 6355 Ipp0644 693470 693655 p unknown similar to unknown protein 5.2 similar to aminodeoxychorismate 3599.5 5045 Ipp0645 693669 694484 p unknown lyase (PabC) 2.5
Table XIV Similar to conserved hypothetical 5741.1 6266 Ipp0646 694511 695194 m unknown protein. Predicted membrane protein. 5.2 Similar to l-acyl-sn-glycerol-3- 2657.3 4488 Ipp0647 695376 697550 p unknown phosphate acyltransferase 2.4 2655.2 4487 Ipp0648 697551 697970 P unknown similar to unknown protein 5.2 2653.2 4486 Ipp0649 698057 698440 m unknown 6 similar to poly-beta- 482.2 5846 Ipp0650 698489 700231 m unknown hydroxybutyrate synthase 2.4 Similar to conserved hypothetical 481.1 5837 Ipp0651 700672 701133 P unknown protein 5.2 Similar to ABC transporter, 480.3 5828 Ipp0652 701126 702574 P unknown permease component 1.2 Similar to ABC transporter ATP- 1951.4 4094 Ipp0653 702576 703328 P unknown binding protein 1.2 Similar ABC transporter, permease 1950.2 4093 Ipp0654 703325 704611 P unknown component 1.2 similar to cysteine desulfurase and 342.4 4930 Ipp0655 704601 705845 P unknown to selenocysteine lyase 2.2 NifU protein family, possibly 343.3 4936 Ipp0656 705842 706291 unknown involved in the formation or repair of [Fe-Sl clusters 2.5 Similar to conserved hypothetical 344.1 4940 Ipp0657 706363 706698 unknown protein 5.2 similar to putative lysyl-tRNA 346.2 4953 Ipp0658 706701 707654 P unknown synthetase 3.6 347.1 4959 Ipp0659 707727 708641 m unknown Similar to methyltransferase 2.1.1 similar to a domain of alanyl-tRNA 349.2 4972 Ipp0660 708888 709532 m unknown synthetase 3.7.2 Similar to major facilitator family 2652.2 4485 Ipp0661 709535 710830 m unknown transporter 1.2 2650.1 4484 Ipp0662 711484 712797 P unknown similar to unknown protein 5.2 2521.4 4413 Ipp0663 712857 713903 m unknown similar to alcohol dehydrogenase 2.1.1 2522.1 4414 Ipp0664 714104 714448 m unknown 6 2523.1 4415 Ipp0665 714635 715039 P unknown similar to hypothetical protein 5.2 2524.2 4416 Ipp0666 715075 715764 m unknown Similar to polypeptide deformylase 2.2 2649.1 4483 Ipp0667 716007 717194 P unknown Similar to Na+/H+ antiporters 1.2
Table XIV 650.4 6385 Ipp0668 717196 718317 P unknown similar to unknown protein 5.2 651.3 6386 Ipp0669 718382 718783 m unknown similar to unknown protein 5.2 similar to conserved hypothetical 652.1 6387 Ipp0670 718931 719788 P unknown protein 5.2 2648.2 4482 Ipp0671 Similar to L.pneumophila major 720181 721272 P unknown outer membrane protein 1.1 2647.2 4481 Similar to 3-methyladenine-DNA Ipp0672 721432 722004 m tag unknown glycosylase I 3.2 2646.1 4480 Ipp0673 722321 722701 P unknown 6 2645.1 4479 Ipp0674 722802 723755 m unknown 6 SidA protein, substrate of 1288.2 3683 Ipp0675 723866 725290 m sidA the Dot/Icm transport system 5.1 similar to probable transmembrane 1915.3 4072 Ipp0676 725451 727379 m unknown protein 5.2 1913.2 4071 Similar to hypothetical protein, Ipp0677 727505 727897 m - unknown predicted membrane protein 5.2 2644.1 4478 Ipp0678 728034 728411 P + Unknown 6 2643.1 Similar to unknown eukaryotic 4477 Ipp0679 728612 729919 P - unknown proteins 5.2 1434.3 weakly similar with DNA 3768 Ipp0680 729945 732152 m - unknown uptake/competence proteins 1.2 2000.1 4114 competence and adherence Ipp0681 732230 732679 m - pilE type-IV pilin associated protein -CAP- 1.8 290.2 4612 weakly similar to type 4 fimbrial Ipp0682 732691 736200 m + unknown biogenesis protein PilYl 1.8 286.2 Similar to Tfp pilus assembly 4598 Ipp0683 736213 736725 m + unknown protein PilX 1.8 284.2 4591 Similar to Tfp pilus assembly Ipp0684 736722 737789 m - unknown protein PilW 1.8 weakly similar with pre-pilin leader 283.1 4585 Ipp0685 737786 738325 m - unknown sequence 1.8 282.2 similar to type-4 fimbrial pilin 4578 Ipp0686 738338 738916 m - unknown related protein 1.8 1882.2 Similar to peptidoglycan GlcNAc 4049 Ipp0687 739267 740127 m + unknown deacetylase proteins 2.1.1 1883.4 4050 Ipp0688 740304 741641 m - unknown 6 5159.2 6023 Ipp0689 741941 743422 P - Unknown similar to hypothetical proteins 5.2 1926.2 4077 Ipp0690 743613 744242 P - tdk unknown similar to thymidine kinase 2.3
Table XIV Similar to major facilitator family 1928.2 4078 Ipp0691 744290 745561 unknown transporter 1.2 Similar to major facilitator family 813.2 6466 Ipp0692 745574 746857 unknown transporter 1.2 815.2 6467 Ipp0693 746854 748077 deoB Phosphopentomutase 2.3 Peptidase component of 3860.2 5208 Ipp0694 748281 748829 hslV the HslUV protease (Heat shock protein) 4.1 ATP-dependent hsl 3859.3 5207 Ipp0695 748831 750156 hslU protease ATP-binding subunit HslU 4.1 2690.2 4502 Ipp0696 750356 751495 P - unknown 6 2688.2 4500 Ipp0697 751592 751930 P - unknown 6 similar to tRNA processing 1998.4 4111 Ipp0698 752080 753318 m - unknown ribonuclease BN 3.6 structural toxin protein 6097.4 6333 Ipp0699 753773 776812 P - rtxA-1 RtxA 4.3 Similar to flavoprotein WrbA (Trp 583.2 6285 Ipp0700 776991 777590 P - wrbA unknown repressor binding protein) 3.5.2 Similar to conserved hypothetical 585.2 6286 Ipp0701 778084 778434 P - unknown protein 5.2 587.2 6287 Ipp0702 778490 779272 P - exoA unknown Similar to exodeoxyribonuclease 3.2 Similar to exodeoxyribonuclease III 588.3 6291 Ipp0703 779265 780035 P - xthA unknown XthA 3.2 5600.2 6209 Ipp0704 780172 780399 P - rpmE 50S ribosomal protein L31 3.7.1 Similar to NADP-dependent malic 1042.5 3536 Ipp0705 780526 781761 P - Unknown enzyme 2.1.1 Similar to major facilitator family 1044.3 3537 Ipp0706 781973 783265 P - unknown transporter .2 Similar to major facilitator family 4388.1 5559 Ipp0707 783246 784523 P - unknown transporter 2 4387.4 5558 Ipp0708 784554 785372 m - dam DNA adenine methylase 2 Similar to tyrosine-specific 5598.2 6208 Ipp0709 785543 786739 m + unknown transport protein 2 Similar to tyrosine-specific 5735.2 6264 Ipp0710 786940 788130 m + unknown transport protein 2 Putative lipoprotein, similar to 5177.2 6027 Ipp0711 788141 788833 m - Unknown other proteins 2
Table XIV 5176.1 6026 Ipp0712 789064 789264 P unknown 6 915.3 6533 Ipp0713 789717 790562 P unknown similar to membrane protein 1.2 Similar to ABC-type multidrug 914.3 6532 Ipp0714 790565 792307 P unknown transport, ATP-binding protein 1.2 Similar to ABC transporter 1685.5 3925 Ipp0715 792325 793446 P unknown (permease) 1.2 384.5 5197 Ipp0716 793632 794717 P Unknown Similar to transposase (IS4 family) 4.5 6195.2 6350 Ipp0717 794743 795036 P unknown similar to unknow proteins 5.2 Similar to putative ABC transporter 3181.2 4792 Ipp0718 795137 796207 unknown permease protein, hypothetical start codon 1.2 similar to outer membrane efflux 1314.3 3695 Ipp0719 796216 797655 P + unknown protein 1.2 + Similar to soluble lytic murein 3179.1 4790 Ipp0720 797980 799761 m unknown transglycosylase 1.1 ribulose-phosphate 3- 3178.1 4789 Ipp0721 799803 800456 m - rpe epimerase 2.1.1 3177.1 4788 Ipp0722 800670 801134 m - unknown Predicted membrane protein 5.2 Similar to conserved hypothetical 3175.1 4787 Ipp0723 801131 801910 m - unknown protein 5.2 Similar to conserved hypothetical 3174.1 4786 Ipp0724 801903 802772 m - unknown protein 5.2 3173.1 4785 Ipp0725 802778 803032 m + unknown Similar to hypothetical protein 5.2 3172.3 4784 Ipp0726 803369 804394 P - unknown Similar to predicted esterae 2.4 similar to NADH-ubiquinone 4749.2 5793 Ipp0727 804510 806726 unknown oxidoreductase 1.4 similar to acetoacetate 4748.1 5792 Ipp0728 806768 807532 unknown decarboxylase 2.1.1 4746.1 5791 Ipp0729 807569 807841 P + unknown Similar to unknown protein 5.2 2442.2 4360 Ipp0730 807834 809147 P Unknown similar to adenylate cyclase 2.3 Similar to conserved hypothetical 2443.2 4361 Ipp0731 809167 809982 m unknown protein 5.2 similar to conserved hypothetical 2861.1 4599 Ipp0732 809975 810802 m unknown protein 5.2 1747.2 3964 Ipp0733 811100 811366 P unknown 6 similar to amino acid ABC 1746.2 3963 Ipp0734 811621 812373 m unknown transporter, periplasmic binding protein. 1.2
Table XIV ar to glutamine synthetase 1148.2 3602 Ipp0735 812511 815252 glnE i n simil U in 1 IMkn lU VVn 1 1 adenylyltransferase 2.2 Similar to conserved hypothetical 2862.1 4600 Ipp0736 815538 816728 unknown protein 5.2 2863.1 4601 Ipp0737 816796 817368 unknown similar to unknown protein 5.2 similar to ABCrtype multidrug 2571.2 4441 Ipp0738 817361 818287 unknown transport system, ATPase component 1.2 similar to ABC transporter, 2570.2 4440 Ipp0739 818284 819420 p unknown permease component 1.2 Similar to conserved hypothetical 2866.1 4602 Ipp0740 819490 820809 m unknown protein 5.2 Similar to thiol :disulfιde 2868.1 4603 Ipp0741 820899 822659 m + dsbD/lidC unknown interchange protein DsbD 3.9 10 kDa chaperonin (Protein 2869.1 4604 Ipp0742 822845 823135 p groES CpnlO) (groES protein) (Heat shock protein A) 3.9 60 kDa chaperonin (Protein Cpn60)(groEL 519.3 6032 Ipp0743 823163 824809 p htpB protein)(Heat shock protein B). 3.9 Weakly similar to DNA-binding 520.1 6036 Ipp0744 824932 825372 p unknown ferritin-like protein (oxidative damage protectant) 5.2 Similar to conserved hypothetical 521.2 6043 Ipp0745 825629 827746 p unknown protein, predicted membrane protein 5.2 Topoisomerase IV subunit 1438.4 3769 Ipp0746 828080 829960 m parE B 3.4 Similar to ABC transporter ATP- 2450.3 4366 Ipp0747 830094 831902 p unknown binding protein 1.2 LigA protein(Legionella Infectivity 1819.6 4013 Ipp0748 831991 836274 p unknown Gene A) 6 Prolyl-tRNA synthetase (Proline— tRNA 300.2 4685 Ipp0749 836632 838341 m proS Mgase)(ProRS)(Global RNA synthesis factor) 3.7.2 304.2 4704 Ipp0750 838515 841352 P unknown Ankyrin repeat protein 4.6 1666.3 3914 Ipp0751 841359 843017 m unknown 6
Table XIV Similar to inorganic transporter and 3224.3 4810 Ipp0752 843158 845464 m + unknown to carbonic anhydrase (bifunctional) 4.7 similar to conserved hypothetical 3223.1 4809 Ipp0753 845654 846505 m - unknown protein 5.2 similar to outer membrane protein 3221.1 4808 Ipp0754 846523 847890 m - tolC unknown TolC 1.2 Similar to L-isoaspartate 3218.1 4807 Ipp0755 847902 848552 m - unknown carboxylmethyltransferase protein pcm 3.8 2-amino-3-ketobutyrate 3217.3 4806 Ipp0756 848733 849917 P - kbl coenzyme A ligase 2.2 2600.4 4458 Ipp0757 850004 851026 P - tdh threonine dehydrogenase 2.2 Similar to ABC transporter, ATP- 2599.4 4456 Ipp0758 851069 852742 P - unknown binding protein 1.2 Similar to enhanced entry protein 1370.5 3725 Ipp0759 852784 853395 P - unknown EnhA 5.2 Similar to conserved hypothetical 1369.2 3724 Ipp0760 853505 853942 P - unknown protein, predicted membrane protein 5.2 Similar to conserved hypothetical 1367.3 3723 Ipp0761 853942 854367 P + unknown protein, predicted membrane protein 2122.2 4173 Ipp0762 854407 855579 m + unknown probable outer membrane protein weakly similar to L. pneumophila 1034.3 3528 Ipp0763 855636 856610 m + unknown IcmL protein 1032.3 3527 Ipp0764 856623 857582 m - hutG unknown Similar to formimidoylglutamase 1030.3 3526 Ipp0765 857920 858681 P - unknown Similar to oxidoreductase similar to imidazolone-5-propionate 1028.3 3524 Ipp0766 858681 859892 P - hutl unknown hydrolase Hutl Similar to conserved hypothetical 3212.2 4805 Ipp0767 859919 860614 P + unknown protein 2460.4 4372 Ipp0768 860614 862614 P + unknown Similar to transporters 2461.2 4373 Ipp0769 862645 862857 m unknown similar to putative integrase 2462.2 4374 Ipp0770 862999 863223 m - unknown similar to transposase, partial 2464.2 4375 Ipp0771 863397 863732 P - Unknown Similar to transposase (IS5 family)
Table XIV 4096.1 5355 Ipp0772 863645 864127 P Unknown Similar to transposase (IS5 family) 4097.1 5356 Ipp0773 864103 864252 m unknown similar to transposase, partial 4098.1 5357 Ipp0774 864361 864633 m unknown similar to tranposase, partial 4101.1 5359 Ipp0775 865026 865229 m unknown similar to transposase 6404.1 6378 Ipp0776 865196 865414 m Unknown Similar to transposase, partial 474.2 5787 Ipp0777 865372 865704 m unknown similar to tranposase 475.1 5794 Ipp0778 865705 865884 m unknown hypothetical gene Repeats in the N-terminal domain, 4533.2 5654 Ipp0779 866241 871886 P + unknown similar to autotransporter 4.6 Similar to two-component sensor 3285.3 4854 Ipp0780 872079 873152 m unknown histidine kinase 1.3 Similar to two component 3288.3 4855 Ipp0781 873133 873804 m unknown transcriptional regulator 3.5.2 3290.1 4856 Ipp0782 873953 874948 P - unknown 6 3291.2 4857 Ipp0783 875049 875510 m + unknown 6 Similar to proton/sodium- 1733.3 3954 Ipp0784 875599 876846 m + unknown glutamate symport protein 1.2 3295.2 4858 Ipp0785 876968 879733 m - valS valyl-tRNA synthetase 3.7.2 Highly similar to multidrug efflux 1021.3 3520 Ipp0786 879992 883039 m - unknown transporter 1.2 + Similar to efflux transporter, RND 1440.4 3771 Ipp0787 883156 884418 m unknown family 1.2 6191.2 6349 Ipp0788 884659 885063 P unknown 6 3496.4 4974 Ipp0789 885129 885566 P unknown 6 Similar to predicted periplasmic or 3494.2 4973 Ipp0790 885691 886002 P + unknown secreted lipoprotein 5.2 Similar to serine 216.2 4199 Ipp0791 886030 887283 P - glyA unknown hydroxymethyltransferase 2.2 215.1 4192 Ipp0792 887286 887753 P - unknown similar to unknown proteins 5.2 Similar to N utilization substance 214.1 4184 Ipp0793 887811 888254 P - nusB unknown protein B homolog 3.5.4 Thiamine-monophosphate 213.1 4176 Ipp0794 888247 889203 P - thiL kinase 2.5 Phosphatidylglycerophosph 212.1 4170 Ipp0795 889293 889775 P - pgpA atase A 2.4 211.1 4166 Ipp0796 889775 890827 P - unknown Similar to predicted permease 1.2
Table XIV Some similarity with outer surface 209.2 4158 Ipp0797 890619 891556 m + unknown protein 1.8 weakly similar to outer membrane 5526.2 6171 Ipp0798 891566 892204 m + unknown protein 1.8 2549.3 4428 Ipp0799 892394 893818 P - unknown 6 Similar to glutamine-dependent 331.2 4867 Ipp0800 893870 895480 m - unknown NAD(+) synthetase 2.5 Similar to DNA/RNA helicases, 333.3 4878 IppOδOl 895764 899030 P - unknown superfamily II, SNF2 family 3.2 Similar to conserved hypothetical 5690.2 6251 Ipp0802 899289 899723 P + unknown protein 5.2 4972.4 5930 Ipp0δ03 899901 901283 P - dnaB Replicative DNA helicase 3.1 4968.4 5928 Ipp0δ04 901285 902358 P - unknown similar to alanine racemase 1 2.2 similar to surface antigens (17 6188.2 6347 Ipp0805 902431 902880 P + unknown kDa) 5.2 Similar to conserved hypothetical 5521.2 6168 Ipp0806 903188 903622 P - unknown protein 5.2 3684.3 5107 Ipp0807 903731 904918 P + unknown similar to unknown proteins 5.2 3685.1 5108 IppOδOδ 905030 905326 P - unknown similar to unknown proteins 5.2 3686.1 5109 Ipp0809 905323 906873 P - Unknown regulatory protein (GGDEF domain) 1.3 3687.1 5110 IppOδlO 907025 908014 P - lipA Lipoic acid synthetase LipA 2.5 similar to conserved hypothetical 3688.1 5111 IppOδl l 908326 908607 m - unknown protein 5.2 di/tripeptide transporter 776.3 6452 Ipp0812 909490 910998 m - iraB homolog IraB 1.2 small-molecule 778.2 6453 Ipp0813 911037 911855 m - iraA methyltransferase IraA 4.6 779.4 6454 Ipp0814 912015 913397 P - unknown Similar to LPS biosynthesis protein 5.1 Similar to imidazole glycerol 4119.2 5372 Ipp0815 913397 914158 hisF Unknown phosphate synthase subunit HisF 2.2 similar to Imidazole glycerol 4118.3 5371 Ipp0816 914155 914796 hisH unknown phosphate synthase subunit HisH 2.2 CMP-N-acetlyneuraminic 4116.3 5370 Ipp0817 914809 915507 neuA acid synthetase 1.1
Table XIV N-acetylneuraminic acid 4115.1 5369 Ipp0818 915504 916526 p neuB condensing enzyme 1.1 N-acylglucosamine 2- 1764.3 3975 Ipp0819 916526 917659 p neuC epimerase 1.1 1765.4 3976 Ipp0820 917649 918257 p unknown Similar to acetyl transferase 2.1.1 similar to polysaccharide 1818.4 4012 Ipp0821 918268 919770 p unknown biosynthesis protein 1.1 dTDP-4-dehydrorhamnose 4113.1 5368 Ipp0822 919860 920423 p rmlC

dTDP-4-keto-L-rhamnose 4112.1 5367 Ipp0823 920424 921308 p rmlD reductase 2.1.1 dTDP-D-glucose 4,6- 4111.2 5366 Ipp0824 921268 922347 p rmlB dehydratase 2.1.1 Glucose-6-phosphate 5500.1 6162 Ipp0825 922386 923885 p gpi isomerase 2.1.2 glucose-1-phosphate 5499.1 6160 Ipp0826 924040 924915 p rmlA thymidylyltransferase 2.1.1 similar to NAD dependent 5498.1 6159 Ipp0827 924936 925892 p unknown epimerase/dehydratase family protein 2.1.1 alpha-N- 5497.1 6158 Ipp0828 925876 926880 p wecA acetylglucosaminyltransfer ase 1.1 Similar to N-terminal part of 6110.1 6334 Ipp0829a 926995 927237 P unknown Legionella hypothetical protein 5.1 Similar to central part of Legionella 5496.2 6157 Ipp0829b 927224 927763 P unknown hypothetical protein 5.1 Similar to C-terminal part of 4965.2 5926 Ipp0829c 927739 928236 P unknown Legionella hypothetical protein 5.1 4966.2 5927 Ipp0830 928337 930133 m unknown 5.1 1935.4 4083 Ipp0831 930136 931254 m unknown 5.1 1934.4 4082 Ipp0832 931265 932266 m unknown 5.1 4782.3 5819 Ipp0833 932256 933347 m unknown Similar to sialic acid synthase 1.1 5528.2 6172 Ipp0834 933360 934079 m unknown 5.1 5060.2 5984 Ipp0835 934395 935912 P unknown 5.1 5061.4 5985 Ipp0836 936145 937827 P unknown 5.1 ABC transporter of LPS O- 4156.3 5401 Ipp0837 937793 938644 wzm antigen, Wzm 1.2
Table XIV ABC transporter of LPS 0- 4157.2 5402 Ipp0838 938641 940065 P - wzt antigen, Wzt 1.2 4158.1 5403 Ipp0839 940071 941219 P + unknown 5.1 Similar to putative 4159.1 5404 Ipp0840 941233 942123 m - unknown glycosyltransferase 1.1 1359.2 3719 Ipp0841 942904 943977 P - lag-1 O-acetyltransferase 1.1 4160.4 5405 Ipp0842 944090 944983 m - unknown similar to glycosyltransferase 1.1 4161.4 5406 Ipp0843 944977 945996 m - unknown similar to glycosyl transferase 1.1 5530.2 6174 Ipp0844 946208 946915 P - unknown similar to glyoxalase II 4.1 5256.2 6060 Ipp0845 947474 947668 P - csrA global regulator CsrA 3.5.2 3827.2 5190 Ipp0846 947794 949776 m unknown Similar to O-antigen acetylase 1.1 biotin-[acetylCoA carboxylase] holoenzyme 3826.1 5189 Ipp0847 949939 950934 m - birA synthetase and biotin operon repressor 3.5.2 Similar to phosphopantetheinyl 2457.2 4369 Ipp0848 950938 951675 m - unknown transferase 2.4 acetyl-CoA carboxylase 2456.2 4368 Ipp0849 951757 952710 P " accA carboxyl transferase subunit alpha 2.4 3824.2 5188 Ipp0850 952703 953998 P - unknown Similar to cell cycle protein MesJ 1.7 3822.2 5187 Ipp0851 954348 954884 P - unknown 6 3821.1 5186 Ipp0852 955223 956389 P - unknown 6 Similar to alginate o- 2320.3 4274 Ipp0853 956386 957828 P - unknown acetyltransferase Algl 1.1 2319.3 4273 Ipp0854 957927 959303 m - sdhL unknown Similar to L-serine dehydratase 2.2 macrophage infectivity 2409.3 4333 Ipp0855 959508 960209 P + mip potentiator 3.9 2410.2 4335 Ipp0856 960375 961634 P + unknown Similar to AmpG protein 1.2 2795.1 4566 Ipp0857 961746 962012 P - unknown Similar to transposase 4.5 540.3 6115 Ipp0858a 962033 962356 P - unknown similar to transposase, partial 4.5 539.3 6111 Ipp0858b 962379 962978 P - unknown similar to transposase, partial 4.5 538.1 6105 Ipp0859 963489 964139 P - unknown 6 537.3 6101 Ipp0860 964419 964805 P + unknown similar to unknown proteins 5.2 262.4 4466 Ipp0861 964909 966252 m - nadA unknown similar to quinolinate synthetase A 2.5 263.1 4472 Ipp0862 966327 967976 m - nadB L-aspartate oxidase 2.5
Table XIV 264.2 4476 Ipp0863 967973 969343 m purB adenylosuccinate lyase 2.3 829.2 6474 Ipp0864 969496 971199 P unknown Similar to sulfate transporter 1.2 2794.2 4565 Ipp0865 971402 973105 m unknown Similar to acyl-CoA dehydrogenase 2.4 2793.1 4564 Ipp0866 973335 974333 m unknown Similar to hydrolase 4.6 similar to phosphoenolpyruvate 2791.1 4563 Ipp0867 974558 976945 P ppsA unknown synthase 2.1.2 2788.1 4562 Ipp0868 977187 978626 P unknown Similar to Na(+)/H(+) antiporter 1.2 nicotinate-nucleotide 2785.1 4561 Ipp0869 978623 979465 P nadC pyrophosphorylase 2.5 UDP-N-acetylglucosamine- N-acetylmuramyl- (pentapeptide)pyrophosph 1865.3 4040 Ipp0870 979462 980553 murG oryl-undecaprenol N- acetylglucosamine transferase Similar to succinyl-diaminopimelate 1864.3 4039 Ipp0871 980528 981943 P unknown desuccinylase 2784.3 4560 Ipp0872 982035 982352 P unknown Similar to other proteins Rod shape-determining 4841.2 5855 Ipp0873 982548 983585 P mreB protein MreB Rod shape-determining 4837.1 5854 Ipp0874 983563 984471 P mreC protein MreC Rod shape-determining 4836.1 5853 Ipp0875 984468 984947 P mreD protein MreD Similar to ribosomal large subunit 4835.4 5852 Ipp0876 984883 985485 m unknown pseudouridine synthase (Pseudouridylate synthase) 3.6 5650.3 6234 Ipp0877 985746 986444 P unknown 6 5649.1 6233 Ipp0878 986554 987810 P icd isocitrate dehydrogenase 2.1.3 Similar to ATP-dependent Clp 5647.1 6232 Ipp0879 988212 988547 unknown protease adaptor protein ClpS 4.1 ATP-dependent Clp 5645.2 6231 Ipp0880 988579 990846 clpA protease ATP-binding subunit ClpA 4.1 6002.2 6318 Ipp0881 991147 992232 Unknown Similar to transposase (IS4 family) 4.5
Table XIV 6186.1 6346 Ipp0882 992258 992551 P ~ unknown Similar to unknown proteins 5.2 Similar to lipopolysaccharide 4618.1 5704 Ipp0883 992707 993486 P - unknown biosynthesis glycosyltransferase 1.1 Similar to putative O-antigen 4616.3 5703 Ipp0884 993489 994739 P + unknown biosynthesis protein 1.1 4615.3 5702 Ipp0885 994740 995075 m + unknown hypothetical gene 6 Similar to uncharacterized 5479.2 6149 Ipp0886 995245 995844 m - unknown membrane protein 5.2 3747.2 5143 Ipp0887 995841 996743 m + unknown similar to peptidase proteins 2.2 Exonuclease VII, large 3748.1 5144 Ipp0888 996752 998083 m - xseA subunit 2.3 3750.2 5145 Ipp0889 998233 999957 P + unknown similar to outer membrane protein 1.1 3752.2 5146 Ipp0890 999908 1000642 P + unknown similar to unknown proteins 5.2 regulatory protein (GGDEF and EAL 3753.2 5147 Ipp0891 1000658 1002577 P + Unknown domains) 1.3 3754.2 5148 Ipp0892 1002713 1003438 P + unknown 6 Similar to flavin-containing 3756.2 5149 Ipp0893 1003416 1004756 P - unknown monooxygenases 2.1.1 similar to oxidoreductases, short- 4469.2 5605 Ipp0894 1004849 1005697 m unknown chain dehydrogenase/reductase . family 2.1.1 indole-3-glycerol 795.2 6459 Ipp0895 1006173 1006949 m trpC phosphate synthases 2.2 anthranilate 4470.1 5607 Ipp0896 1006942 1007976 m trpD phosphoribosyltransferase Anthranilate synthase 1457.2 3781 Ipp0897 1007954 1008532 m trpG component II Similar to ABC transporter, ATP- 1456.2 3780 Ipp0898 1008566 1009291 m unknown binding protein Similar to conserved hypothetical 1455.2 3779 Ipp0899 1009288 1009797 m + unknown protein Similar to conserved hypothetical 4471.3 5608 Ipp0900 1009778 1010347 m + unknown protein Similar to conserved hypothetical 5978.2 6312 Ipp0901 1010344 1010871 m unknown protein
Table XIV Similar to arabinose 5-phosphate 5903.2 6301 Ipp0902 1010885 1011847 m - kdsD unknown isomerase 1.1 Similar to ABC transporter, ATP- 5696.2 6254 Ipp0903 1012215 1013012 P - unknown binding protein 1.2 Similar to permease of ABC 5695.1 6253 Ipp0904 1013005 1013787 P - unknown transporter 1.2 5694.2 6252 Ipp0905 1013788 1014264 P + unknown similar to unknown protein 5.2 5489.2 6154 Ipp0906 1014272 1014880 P + unknown similar to unknown protein 5.2 weakly similar to anti-anti-sigma 4341.3 5524 Ipp0907 1014883 1015164 P - unknown factor 3.5.1 Similar to conserved hypothetical 4342.1 5525 Ipp0908 1015554 1015799 P _ unknown protein 5.2 UDP-N-acetylglucosamine 4344.1 5526 Ipp0909 1015792 1017060 murA 1-carboxyvinyltransferase 1.1 Similar to conserved hypothetical 4345.1 5527 Ipp0910 1017057 1017815 unknown protein 5.2 Highly similar to ABC transporter, 1484.2 3795 Ipp0911 1018004 1018681 m unknown ATP-binding protein 1.2 Similar to conserved hypothetical 1485.4 3796 Ipp0912 1018678 1019928 m + unknown protein 5.2 4848.3 5859 Ipp0913 1019928 1020944 m + unknown similar to membrane-fusion protein 1.2 4847.2 5858 Ipp0914 1021334 1021927 P + unknown 6 transcriptional regulator 1096.3 3570 Ipp0915 1022123 1023538 P - fleQ FleQ 3.5.2 1094.1 3569 Ipp0916 1023582 1023863 m - unknown 6 1093.3 3568 Ipp0917 1024086 1025024 m - unknown Similar to protease 2.2 heme exporter protein 4564.2 5670 Ipp0918 1025169 1025771 P - ccmA CcmA heme exporter protein 2597.4 4454 Ipp0919 1025768 1026448 P - ccmB CcmB heme exporter protein 3159.2 4775 Ipp0920 1026662 1027417 P + ccmC CcmC heme exporter protein 3158.1 4774 Ipp0921 1027338 1027559 P - ccmD CcmD cytochrome c-type 3157.1 4773 Ipp0922 1027565 1027996 ccmE biogenesis protein CcmE 1.4
Table XIV 3153.3 4772 Ipp0923 1027993 1029945 p + cytochrome C-type ccmF biogenesis protein CcmF 1.4 3152.1 4771 Ipp0924 1029942 1030475 p + cytochrome C biogenesis ccmG protein 1.4 Cytochrome c-type 2085.2 4157 Ipp0925 1030475 1030876 p ccmH biogenesis protein CcmH similar to cytochrome c-type 2083.2 4156 Ipp0926 1030869 1031555 p Unknown biogenesis protein Similar to conserved hypothetical 2082.2 4155 Ipp0927 1031545 1031748 p unknown protein Similar to outer membrane 1853.2 4031 Ipp0928 1031804 1032208 p unknown lipoprotein Similar to methyladenine DNA 9002.1 6523 Ipp0929 1032403 1032954 m unknown glycosylase similar to ATP-dependent DNA 998.3 6589 Ipp0930 1032964 1034790 m unknown helicase RecQ 999.2 6590 Ipp0931 1034904 1036058 p unknown similar to acyl-CoA dehydrogenase similar to enoyl-CoA 1000.2 3509 Ipp0932 1036075 1036851 p Unknown hydratase/carnithine racemase similar to enoyl-CoA 1542.2 3831 Ipp0933 1036854 1037912 p unknown hydratase/isomerase family protein 2.4 1539.2 3829 Ipp0934 1038078 1038944 m unknown 6 peptide chain release 1230.2 3648 Ipp0935 1039168 1040748 p prfC factor 3 3.7.5 1231.3 3649 Ipp0936 1040814 1041197 m unknown 6 NAD(P) transhydrogenase subunit beta (Pyridine 4773.3 5811 Ipp0937 1041332 1042729 m pntB nucleotide transhydrogenase subunit beta) 1.4 NAD(P) transhydrogenase subunit beta (Pyridine 4774.1 5812 Ipp0938 1042753 1043049 m pnaB nucleotide transhydrogenase subunit alpha II) 1.4
Table XIV pyridine nucleotide 4775.2 5813 Ipp0939 1043042 1044175 m pntA transhydrogenase, alpha subunit 2.3 4776.3 5814 Ipp0940 1044569 1045123 m unknown Similar to uncharacterized proteins 5.2 6093.1 6332 Ipp0941 1045377 1045652 m unknown 6 2060.3 4142 Ipp0942 1045949 1047082 P Unknown regulatory protein (GGDEF domain) 1.3 2062.2 4143 Ipp0943 1047144 1047785 m unknown 6 2065.2 4144 Ipp0944 1048268 1048702 P unknown predicted transmembrane protein 6 2066.5 4145 Ipp0945 1048826 1049272 P unknown 6 2067.5 4146 Ipp0946 1049399 1050475 P unknown similar to glycosyl hydrolase 1.1 4068.2 5336 Ipp0947 1050990 1052270 P unknown Similar to amino acid transporter 1.2 Succinyl-diaminopimelate 4303.1 5497 Ipp0948 1052594 1053727 m dapE desuccinylase 2.2 2,3,4,5-tetrahydropyridine- 4302.1 5496 Ipp0949 1053720 1054550 m dapD 2,6-dicarboxylate N- succinyltransferase 2.2 4301.1 5495 Ipp0950 1054554 1055447 m unknown Similar to acetyltransferase 4.6 1420.2 3761 Ipp0951 1055514 1056665 m metB unknown Similar to cystathionine beta-lyase 2.2 regulatory protein (GGDEF and EAL 1421.3 3762 Ipp0952 1056814 1059129 P Unknown domains) 1.3 Similar to kynurenine 3- 5446.3 6135 Ipp0953 1059431 1060780 P unknown monooxygenase 2.2 5447.1 6136 Ipp0954 1060998 1061750 P unknown Similar to unknown proteins 5.2 1834.4 Similar to eukaryotic cytokinin 4020 Ipp0955 1061707 1063077 P unknown oxidase 4.6 5448.1 6137 Ipp0956 1063427 1063936 P unknown 6 4571.2 5674 Ipp0957 1063942 1064319 P unknown 6 2576.2 4442 Similar to putative sodium/calcium Ipp0958 1064401 1065360 m unknown antiporter 1.2 2578.2 4443 Ipp0959 1065633 1066337 m unknown 5.1 4572.2 Similar to A/G-specific adenine 5675 Ipp0960 1066576 1067643 m mutY unknown glycosylase 3.2
Table XIV Similar to conserved hypothetical 4573.4 5676 Ipp0961 1067640 1069181 m + unknown protein 5.2 5904.2 6302 Ipp0962 1069595 1070251 P - unknown 6 5010.2 5952 Ipp0963 1070262 1071158 P + unknown 6 5015.2 5953 Ipp0964 1071374 1071712 P - unknown Similar to hypothetical protein 5.2 5460.1 6142 Ipp0965 1071687 1072778 P + unknown Similar to protease 3974.2 5272 Ipp0966 1072898 1073443 P - unknown Similar to hypothetical protein Similar to 3-hydroxyacyl-CoA 3972.2 5271 Ipp0967 1073653 1074396 P - unknown dehydrogenase type II 3971.2 5270 Ipp0968 1074453 1074950 m - unknown Similar to unknown protein Similar to negative regulator of 3970.1 5269 Ipp0969 1074967 1075287 m _ flg unknown flagellin synthesis (Anti-sigma-28 factor) 3.5.2 flagellar basal body P-ring 3969.1 5267 Ipp0970 1075378 1076079 m flgA unknown biosynthesis protein FlgA 1.5 similar to cytochrome c-type 3968.1 5266 Ipp0971 1076166 1076573 p unknown protein 1.4 Similar to enhanced entry protein 3967.1 5265 Ipp0972 1076650 1077207 p unknown EnhA 5.1 Similar to putative transcriptional 3966.1 5264 Ipp0973 1077494 1078264 p unknown regulator, homolog of Bvg accessory factor 3.5.2 similar to conserved hypothetical 3965.1 5263 Ipp0974 1078882 1079340 p unknown protein 5.2 Similar to S-adenosyl- 3964.1 5262 Ipp0975 1079352 1080278 p unknown methyltransferase MraW 2.1.1 3962.2 5261 Ipp0976 1080275 1080613 p unknown similar to cell division protein FtsL 1.7 Peptidoglycan synthetase 5462.3 6143 Ipp0977 1080994 1082655 p ftsl Ftsl precursor 1.1 Similar to UDP-N- 643.3 6381 Ipp0978 1082676 1084127 p murE unknown acetylmuramoylalanyl-D-glutamate- 2,6-diaminopimelate ligase 1.1 similar to erythronate-4-phosphate 644.2 6382 Ipp0979 1084172 1085224 m pdxB unknown dehydrogenase 2.5 645.4 6383 Ipp0980 1085221 1085880 m unknown 5.2
Table XIV 647.5 6384 Ipp0981 1086129 1086776 P unknown similar to unknown protein 5.2 4859.2 5864 Ipp0982 1086816 1088060 P unknown 6 4860.1 5865 Ipp0983 1088145 1088216 P unknown hypothetical gene 6 Electron transfer 5515.3 6166 Ipp0984 1088469 1089218 etfB flavoprotein beta-subunit (Beta-ETF) 1.4 Electron transfer 3942.2 5252 Ipp0985 1089232 1090170 P ~ etfA ffllaavvoopprrootteeiinn,, aallpphhaa ssuubbuunniitt 623.2 6354 Ipp0986 1090182 1091303 P - aid unknown Similar to alanine dehydrogenase 2.2 Similar to peptidoglycan 622.3 6353 Ipp0987 1091375 1093759 m + mrcA unknown synthetase; penicillin-binding protein 1A 1.1 5514.2 6165 Ipp0988 1093909 1094916 m - unknown 6 Tfp pilus assembly protein, 5398.3 6114 Ipp0989 1095333 1096397 P - pilM ATPase PilM 1.8 Tfp pilus assembly protein 3555.2 5017 Ipp0990 1096401 1096949 P - pilN PilN 1.8 Tfp pilus assembly protein 3554.4 5016 Ipp0991 1096963 1097562 P - pilO PilO 1.8 Tfp pilus assembly protein 1895.6 4059 Ipp0992 1097559 1098143 P + pilP PilP 1.8 type IV pilus (Tfp) 1896.5 4060 Ipp0993 1098147 1100243 P + pilQ assembly protein PilQ 1.8 3551.1 5015 Ipp0994 1100689 1101216 P + aroK shikimate kinase I 2.2 3550.1 5014 Ipp0995 1101203 1102312 P - aroB 3-dehydroquinate synthase 2.2 3548.2 5013 Ipp0996 1102309 1103760 P - unknown similar to unknown protein 5.2 Similar to universal stress protein 3547.1 5012 Ipp0997 1103869 1104300 P - unknown A 4.1 Riboflavin biosynthesis protein RibF (Riboflavin 3545.1 5011 Ipp0998 1104424 1105410 p ribF kinase/FMN adenylyltransferase) 2.5 1490.4 3798 Ipp0999 1105523 1108318 p ileS Isoleucyl-tRNA synthetase 3.7.2 Lipoprotein signal 2233.3 4231 IpplOOO 1108315 1108779 p IspA peptidase 1.6
O Q X <I J X φ XJ XJ Q.
+ + +
E E L Q. E ε ε ε C rv rT rr 00 rr ro in Is- σ> rr CO ΓM vO VO in rv 00 VD in in rT o (N O 00 00 ΓM IN Is- 00 VD rr ro rv cn cn ro (N cn VD VD o in CN rT rr in OO ΓM 00 oo 00 rv rT CO (N 00 ΓM in 00 rv cn CN in VD rv 00 cn O IN ro rT in VO rv cn ΓM rr in vo CO o ro o H ΓM IN (N ΓM ΓM IN (N ΓM ro ro ro ro ro rr r rr •H CO CO cn cn rr cn VD in 00 cn in CO VD rf cn in 00 cn rr in in 00 rv cn rr CO LO cn Is- o o VD in Is- VD VD in cn VD o 00 (N in in IN in rv in 00 VD m 00 00 rv rr cn ΓM 00 in VD 00 CO n cn ΓM in VD Is- 00 cn o ΓM ro rr in VO rv cn ΓM rr LO VO 00 O o o ΓM ΓM CM (N (N ΓM (N ΓM 00 ro 00 00 ro rT rT
ΓM 00 rr in VD Is- 00 cn o ΓM ro r in VD Is- 00 cn o (N 00 o o o o o o o O o (N IN (N ΓM o o o o o o o O o o o o o o O o o o O o O o o CL CL Q. CL o. O. Q. O- L CL CL Q. Q. CL Q. α. CL Q. Q. Q. Q. CL Q. CL CL Q. Q. CL CL CL Q. Q. Q. CL C Q. Q. Q. Q. Q. Q. CL Q. o rv CM Is- Is- VD in co 00 O VD 00 ΓM ιn rr 00 cn OO O ΓM rr m rv (N VD cn cn cn cn rv O ΓM ΓM ro rr ΓM cn ro ro 0 rv rv v Isn 0 00 vD rr IN VD P O Is- 00 ΓM rr rr rr rr o r - in LO ΓM ΓM cn rM rr 00 ro in rr rr rr rT ro rT m ιn in in rr rr rr ro ro r rr in in
> P-H
X 00 ro CM ΓM (N ΓM ΓM rT ΓM ΓM ro ΓM Γ 00 00 ΓM (N ro ΓM (N ϋ IN o VD 0s. 00 o cn in d 00 ΓM VD rf cd Γ-1 O ro C3 r-l
2 ro 00 VD (N σi cn cn in vD cn o 00 ΓM IN (N 00 VD VD ΓM ΓM rr CO 00 rr rr ro r rr rr rr rr 00 o o oo oo o cn a ΓM m 00 O OO (N (N 00 ro (N (N in 00
H
Table XIV DNA polymerase III, alpha 1636.2 3892 ppl024 1143240 1146686 P dnaE chain 3.1 3925.2 5243 ppl025 1146772 1148013 m unknown 6 Predicted membrane protein, similar to putative 4- 3924.2 5242 ppl026 1148382 1149227 unknown hydroxybenzoate octaprenyltranferase 2.1.1 1282.3 3678 ppl027 1149188 1150735 P unknown Similar to LphB protein 5.1 similar to nucleoside-diphosphate 3923.2 5241 ppl028 1150802 1152667 P unknown sugar epimerases 1.1 4844.3 5856 ppl029 1152813 1153364 m unknown 6 4845.6 5857 ppl030 1153690 1155384 m unknown 6 4254.3 5468 ppl031 1155852 1156592 P unknown 6 Similar to amino acid (lysine) 4253.1 5467 ppl032 1156889 1158352 m unknown permease 1.2 Similar to eukaryotic 2309.2 4267 ppl033 1158733 1159878 unknown ectonucleoside triphosphate diphosphohydrolase (apyrase) 2.3 2308.2 4266 ppl034 1160427 1160738 P unknown 6 Similar to transposase (ISL3 6180.1 6345 ppl035 1160719 1161894 m Unknown family) 4.5 5630.1 6222 ppl036 1162018 1162479 P unknown hypothetical protein 6 5774.2 6271 ppl037 1162551 1163618 P unknown 6 Similar to conserved hypothetical 2080.3 4154 ppl038 1164101 1164769 P unknown protein 5.2 397.2 5268 ppl039 1164905 1165411 P unknown Similar to UmuD protein 3.2 similar to DNA repair proteins 398.2 5275 ppl040 1165398 1166675 P unknown UmuC 3.2 3394.1 4910 ppl041 1167580 1167714 P unknown hypothetical gene 6 3395.3 4911 ppl042 1167841 1168683 m unknown 6 3396.1 4912 ppl043 1169145 1169642 P unknown hypothetical gene 6 Similar to putative antirestriction 3397.1 4913 ppl044 1169805 1170311 m unknown protein 3.2 similar to single-stranded DNA- 3398.1 4914 ppl045 1170377 1170799 m unknown binding protein (ssb) 3.1 3399.1 4915 ppl046 1170899 1171093 m unknown 6 3401.2 4917 ppl047 1171610 1171903 P unknown 6 3402.2 4918 ppl048 1171958 1172857 m unknown Weakly similar to integrase 4.5
Table XIV 3404.2 4919 Ippl049 1172850 1173656 m - unknown 6 3406.2 4920 Ippl050 1174294 f 1174938 P + unknown 6 5322.2 6087 Ippl051 1174951 1175262 m - unknown similar to unknown proteins 5.2 5321.2 6086 Ippl052 1175314 1175574 m - unknown similar to unknown protein 5.2 4914.3 5897 Ippl053 1175704 1175988 P + unknown 6 4913.2 5896 Ippl054 1176324 1177259 m - unknown 6 Similar to conserved hypothetical 677.6 6397 Ippl055 1177462 1179441 m + unknown protein 5.2 675.6 6396 Ippl056 1179438 1179767 m + unknown 6 Putative membrane protein similar 673.3 6395 ppl057 1179961 1181514 m unknown to conserved hypothetical protein 5.2 Similar to conserved hypothetical 3575.2 5030 ppl058 1181517 1182884 m unknown protein; signal peptide predicted 5.2 Similar to conserved hypothetical 3574.1 5029 ppl059 1182894 1183868 m unknown protein 5.2 1089.2 3563 ppl060 1183861 1186611 m unknown similar to unknown proteins 5.2 1090.1 3565 ppl061 1186621 1186968 m + unknown similar to unknown proteins 5.2 Similar to conserved hypothetical 1091.2 3566 ppl062 1186925 1188184 m + unknown protein 5.2 Similar to conserved hypothetical 1092.3 3567 ppl063 1188181 1188957 m unknown protein 5.2 Similar to conserved hypothetical 5031.2 5963 ppl064 1188944 1189600 m unknown protein 5.2 5032.1 5964 ppl065 1189593 1189955 m Unknown 6. 5033.1 5965 ppl066 1189967 1190332 m unknown 6 5034.2 5966 ppl067 1190329 1190595 m unknown 6 5319.1 6084 ppl068 1190595 1190921 m unknown 6 C-terminal part similar to C- 5318.2 6083 ppl069 1190931 1191563 m unknown terminal part of conserved hypothetical protein 5.2 Similar to conserved hypothetical 4132.3 5382 ppl070 1191554 1191985 m unknown protein 5.2 4131.4 5381 ppl071 1191985 1192806 m + unknown similar to unknown protein 5.2 4130.3 5380 ppl072 1192799 1193383 m + unknown 6 4129.1 5378 ppl073 1193380 1193793 m + unknown similar to PilL protein 1.8
Table XIV 4128.1 5377 Ippl074 1193786 1194004 m unknown similar to carbon storage regulator 3.5.2 4127.1 5376 Ippl075 1193998 1194510 m unknown Similar to Legionella LvrB protein 5.1 4126.2 5375 Ippl076 1194362 1195264 m unknown Similar to LvrA protein 5.1 4123.2 5374 Ippl077 1195431 1195685 P unknown Similar to phage repressor 4.4 similar to DNA modification 682.3 6401 Ippl078 1195761 1196723 P unknown methylase 3.2 681.4 6400 Ippl079 1196727 1197470 m unknown 6 Similar to very-short-patch-repair 680.3 6399 Ippl080 1197572 1198027 m unknown endonuclease vsr 3.2 678.3 6398 IpplOδl 1198125 1199183 m unknown 6 Similar to major facilitator family 4955.4 5919 Ippl082 1199195 1200490 m unknown transporter 1.2 4954.2 5918 Ippl083 1200495 1201013 m unknown Similar to peptide deformylase 3.8 4952.1 5917 Ippl084 1200994 1201569 m unknown 6 4951.2 5916 Ippl085 1201598 1202089 m unknown 6 5382.1 6107 Ippl086 1202296 1202541 P unknown 6 Similar to putative transcriptional 4764.2 5804 Ippl087 1202564 1202782 P unknown regulator 3.5.2 4763.2 5803 Ippl088 1203051 1204295 m unknown Similar to prophage integrase 4.4 1847.3 4029 Ippl089 1204670 1206475 P unknown 6 3588.2 5036 Ippl090 1206588 1207193 P unknown Similar to methyltransferase 2.1.1 similar to conserved hypothetical 3586.2 5035 Ippl091 1207525 1208634 P unknown protein 5.2 1229.3 3646 Ippl092 1208718 1209173 P unknown 6 Similar to beta- 1227.1 3645 Ippl093 1209729 1210397 P unknown phosphoglucomutase 2.1.1 2383.4 4318 Ippl094 1210551 1210757 P unknown 6 1390.5 3738 Ippl095 1210905 1211363 P unknown 6 1391.1 3739 Ippl096 1211486 1212088 P unknown Similar to other protein 5.2 1392.2 3740 Ippl097 1212164 1212487 m Unknown Similar to transposase (IS5 family) 4.5 3584.1 5034 Ippl098 1212577 1212771 m Unknown Similar to transposase (IS5 family) 4.5 131.2 3694 Ippl099 1213052 1213525 m unknown 6 139.6 3737 IppllOO 1214119 1218525 P Unknown Ankyrin repeat protein 6. 2609.2 4463 IppllOl 1219613 1220581 P unknown 6
Table XIV 2607.2 4462 ppl 02 1220717 1221931 P + unknown Putative secreted protein 6 3489.2 4971 ppl 03 1221940 1222932 m unknown 6 similar to Peptide methionine 1582.2 3856 ppl 04 1223130 1223633 P unknown sulfoxide reductase msrB 3.8 1581.4 3855 ppl 05 1223732 1224994 m unknown 6 1156.3 3606 ppl 06 1225330 1226628 m unknown Putative membrane protein 5.2 1153.1 3605 ppl 07 1226823 1227242 m unknown Putative membrane protein 5.2 1152.2 3604 ppl 08 1227429 1228310 m unknown 6 Some similarity with eukaryotic 3488.1 4970 ppl 09 1228551 1230611 m unknown protein 6 3486.1 4969 ppl 10 1230904 1231038 m unknown hypothetical protein 6 3485.1 4968 ppl 11 1231171 1231866 P unknown 6 3484.1 4967 ppl 12 1231960 1232598 m unknown 6 3483.3 4966 ppl 13 1232814 1233419 m unknown Similar to hypothetical proteins 5.2 regulatory protein (GGDEF and EAL 1809.4 4008 ppl 14 1233896 1236223 m Unknown domains) 1.3 3481.2 4965 ppl .15 1236289 1236561 m unknown similar to other proteins 5.2 similar to other protein, ATP 3480.4 4964 ppl 16 1236566 1238218 m unknown binding site 5.2 833.3 6477 ppl 17 1238638 1240986 P unknown similar to chitinase 2.1.1 Similar to B. subtilis PaiA 2116.2 4168 ppl 18 1241095 1241568 m unknown transcriptional repressor of sporulation 3.5.2 similar to D-alanyl-D-alanine 499.2 5938 ppl 19 1241660 1242775 m unknown carboxypeptidase 1.1 Major acid phosphatase PilD-dependent secreted protein, 500.2 5946 ppl 20 1243043 1244107 p map Map (histidine-acid tartrate-sensitive acid phosphatase phosphatase) 2.6 501.2 5951 ppl 21 1244305 1245039 p unknown 6 502.3 5956 ppl 22 1245241 1245621 m unknown 6 similar to membrane-bound lytic 503.3 5961 ppl 23 1245903 1247231 p unknown murein transglycosylase 1.1 Similar to amino acid ABC 3161.1 4777 lppl 24 1247357 1248118 unknown transporter (amino acid binding protein) 1.2 3160.1 4776 lppl 25 1248354 1248944 m unknown 6 Similar to amino acid ABC 1049.2 3540 lppl 26 1249226 1249921 p unknown transporter 1.2
Table XIV Some similarity with eukaryotic 1050.4 3541 Ippl l27 1250105 1253536 unknown proteins 5.2 Similar to long-chain acyl-CoA 2368.5 4307 Ippll28 1253631 1255088 m unknown synthetase 2.4 1778.5 3984 Ippl l29 1255222 1255686 P unknown Similar to guanine deaminase 2.3 1775.3 3983 Ippl l30 1256114 1257673 P unknown 6 3801.2 5172 Ippl l31 1257838 1259289 m ladC adenylate cyclase 2.3 3800.4 5171 Ippl l32 1259451 1259777 m unknown 6 cyclopropane fatty acyl phospholipid synthase 791.5 6458 Ippl l33 1259942 1261069 m cfa (Cyclopropane fatty acid synthase) (CFA synthase) 2.4 3798.1 5170 Ippll34 1261123 1261503 m unknown 6 Similar to 2-nitropropane 3797.1 5169 Ippl l35 1261720 1262772 m unknown dioxygenase 4.2 Similar to transcriptional regulator, 3796.3 5168 Ippl l36 1262750 1263361 m unknown (TetR family?) 3.5.2 3793.3 5167 Ippll37 1263531 1264433 p + unknown Similar to hypothetical protein 5.2 3792.1 5166 Ippl l38 1264540 1264761 m + unknown 6 3791.1 5165 Ippl l39 1264987 1265940 m unknown 6 Similar to spermidine/putrescine- 3789.2 5164 Ippl l40 1266221 1267243 m potD unknown binding periplasmic protein precursor potD (SPBP) 1.2 Similar to spermidine/putrescine transport system permease protein 3788.1 5163 Ippl l41 1267240 1268007 m potC unknown potC. Putative integral membrane protein 1.2 Similar to spermidine/putrescine 3785.2 5162 Ippl l42 1268004 1268852 m potB unknown transport system permease protein PotB 1.2 Similar to spermidine/putrescine 3784.2 5161 Ippl l43 1268818 1269912 m potA unknown transport system ATP-binding protein PotA 1.2
Table XIV Hypothetical protein, similar to 3783.1 5160 ppl 44 1270081 1270374 m - unknown endonuclease 3.2 2355.2 4297 ppl 45 1270574 1271458 m - unknown Similar to dehydrogenase 2.1.1 3780.2 5159 ppl 46 1271801 1272271 m - unknown 6 1505.4 3807 ppl 47 1272563 1274872 P - unknown 6 Similar to thermostable 1507.3 3808 ppl 48 1275002 1276483 P - unknown carboxypeptidase 1 2.2 3454.1 4950 ppl .49 1276607 1277047 P unknown Similar to acetyltransferase 2.1.1 3453.1 4949 ppl .50 1277368 1278852 P - unknown Putative coiled-coil protein 6 3451.1 4948 ppl .51 1279030 1279812 P - unknown 6 similar to E. coli Ada protein (06- 3450.2 4947 ppl .52 1279909 1280979 P - unknown methylguanine-DNA methyltransferase) 3.2 5135.2 6020 ppl .53 1281056 1282528 m - unknown 6 2545.2 4426 ppl .54 1282739 1283563 m - unknown 6 2546.4 4427 ppl 55 1283589 1285076 m + unknown Similar to amine oxidase 2.2 3933.3 5248 ppl 56 1285562 1286644 P - unknown 6 Similar to eukaryotic pyruvate 1261.2 3664 ppl 57 1286884 1288563 P - unknown decarboxylase 2.1.1 1264.3 3665 ppl 58 1288621 1289817 m + unknown Similar to aminopeptidase 2.2 Similar to L.pneumophila putative 4485.2 5618 ppl .59 1290224 1290988 P + unknown lipase LipB 2.4 Some similarity with eukaryotic 4484.1 5617 ppl .60 1291137 1291871 P _ unknown proteins similar to putative drug metabolite 895.3 6517 ppl 61 1292240 1293277 p unknown transport protein (DMT family) 1.2 894.3 6516 lppl 62 1293379 1293624 m unknown 6 Similar to predicted phosphoribosyl 2220.2 4224 1lppl 63 1293956 1294606 p unknown transferase 2.3 4319.2 5505 lppl 64 1294673 1295404 m unknown similar to other proteins 5.2 Similar to permeases of the major 1126.3 3590 lppl 65 1295575 1296861 p unknown facilitator superfamily (MFS) 1.2 Similar to acetylomithine 1125.2 3589 lppl 66 1296873 1298027 p unknown deacetylase 2.2 1124.2 3588 lppl 67 1298353 1299069 p unknown Weakly similar to uridine kinase 2.3 Some similarity with eukaryotic 4316.4 5504 lppl 68 1299134 1301122 m unknown proteins 5.2
able XIV Similar to conserved hypothetical
4508.4 5634 Ippl l69 1301406 1301891 p unknown protein 5.2 regulatory protein (GGDEF and EAL
1105.2 33557766 Ippl l70 1302012 1304318 m Unknown domains) 1.3
4211.2 5439 Ippl l71 1304470 1305813 m unknown Similar to unknown protein 5.2 similar to Pyruvate formate-lyase
4212.1 55444400 Ippl l72 1305885 1306961 p unknown activating enzyme 2.1.1
1137.2 3597 Ippl l73 1307020 1307439 p unknown 6 Similar to conserved hypothetical
1136.3 3596 Ippl l74 1307601 1309073 p NdL unknown protein, similar to C-terminal part of EnhC protein 5.2 Similar to Pseudomonas sensor protein PilS (member of the 2 component response regulator
4214.3 5441 Ippl l75 1309172 1310758 pilS unknown PilS/PilR involved in the regulation of the expression of the type 4 fimbriae) 1.3 Similar to type 4 fimbriae expression regulatory protein PilR 544.2 6133 Ippl l76 1310755 1312083 p pilR unknown (two-component response requlator) 3.5.2 543.1 6131 Ippl l77 1312513 1312758 p unknown 6
2468.2 4377 Ippl l78 1313196 1314131 m unknown similar to putative hydrolase 2.1.1
2469.2 4378 Ippl l79 1314410 1315846 m unknown similar to putative protease 2.2 Riboflavin biosynthesis
2471.3 4380 Ippl lδO 1315909 1316982 p ribD protein RibD 2.5 Riboflavin synthase alpha
1832.4 4019 Ippl lδl 1316967 1317581 p ribE chain 2.5 Riboflavin biosynthesis 1831.3 4018 Ippll82 1317578 1318786 p ribA protein RibA 2.5 riboflavin synthase beta
3721.1 5129 Ippl l83 1318794 1319261 p ribH chain (6,7-dimethyl-8- ribityllumazine synthase) 2.5
3465.2 4956 Ippl l84 1319387 1321024 p pyrG CTP synthase 2.3
Table XIV 2-dehydro-3- deoxyphosphooctonate aldolase (Phospho-2- dehydro-3-deoxyoctonate aldolase) (3-deoxy-D- 3463.1 4955 Ippl l85 1321021 1321845 kdsA manno-octulosonic acid 8- phosphate synthetase) (KDO-8-phosphate synthetase) (KDO 8-P synthase) 2.4 3461.4 4954 Ippllδδ 1322050 1323909 P unknown 6 2293.5 4259 Ippll87 1324001 1326388 P unknown 6 Similar to competence lipoprotein 1409.3 3751 Ippll88 1326772 1327545 P unknown comL precursor 1.2 Similar to recombinaison 1410.2 3753 Ippll89 1327693 1328631 m unknown associated protein RdgC 3.3 Similar to low affinity potassium 3459.4 4952 Ippll90 1328749 1330623 m unknown transport system protein Kup 1.2 3455.3 4951 Ippll91 1330651 1331634 m unknown 6 1602.2 3872 Ippll92 1331712 1332884 P unknown Similar to unknown protein 5.2 Similar to N-acetyl-beta- 1601.3 3871 Ippll93 1333009 1334157 P unknown glucosaminidase 2.1.1 5282.4 6068 Ippll94 1334297 1335022 P unknown similar to unknown proteins 5.2 phosphoribosyl-AMP cyclohydrolase (PRA-CH) / 3771.2 5156 Ippl l95 1335111 1335734 m hisl phosphoribosyl-ATP pyrophosphatase (PRA-PH) 2.2 Imidazole glycerol phosphate synthase 854.4 6490 Ippl l96 1335731 1336498 m hisF subunit HisF (IGP synthase cyclase subunit) 2.2 phosphoribosylformimino-5- aminoimidazole 853.1 6489 Ippl l97 1336492 1337211 m hisA carboxamide ribotide isomerase 2.2
Table XIV Imidazole glycerol phosphate synthase 852.3 6488 Ippl l98 1337205 1337804 m hisH subunit HisH (IGP synthase glutamine amidotransferase subunit) 2.2 histidinol- phosphatase/imisazoleglyc 3772.2 5157 Ippl l99 1337801 1338859 m hisB erol-phosphate dehydratase 2.2 Histidinol-phosphate 1101.4 3574 Ippl200 1338837 1339931 aminotransferase similar to histidinol-phosphate m hisC (Imidazole acetol- aminotransferase phosphate transaminase) 2.2 1102.2 3575 Ippl201 1339932 1341227 m hisD histidinol dehydrogenase 2.2 ATP 2255.2 4241 Ippl202 1341233 1342114 m hisG phosphoribosyltransferase 2.2 Weakly similar to E. coli Trp operon 3778.2 5158 Ippl203 1342111 1342407 m unknown repressor 3.5.2 cytochrome d ubiquinol 5550.3 6185 Ippl204 1342784 1344316 p cydA oxidase subunit I 1.4 cytochrome d ubiquinol 5552.2 6186 Ippl205 1344332 1345468 p cydB oxidase subunit II 1.4 orotate 4409.1 5576 Ippl206 1345694 1346326 m pyrE phosphoribosyltransferase 2.3 4407.1 5575 Ippl207 1346575 1346808 P unknown Similar to cold shock proteins 4.1 4406.1 5574 Ippl208 1347319 1347891 P unknown similar to unknown protein 5.2 4405.1 5573 Ippl209 Similar to conserved hypothetical 1348051 1348506 P unknown protein 5.2 4404.1 5572 Similar to transcriptional regulator, Ippl210 1348536 1348955 m unknown MarR family 3.5.2 4403.2 5571 Ippl211 Similar to conserved hypothetical 1349033 1349821 P unknown protein 5.2 4402.2 5570 Ippl212 1349818 1350366 P unknown Similar to acetyltransferase 2.1.1 2598.3 similar to putative amino acid 4455 Ippl213 1350490 1351149 m + unknown (threonine) efflux protein 1.2
Table XIV Similar to transcriptional regulator, 5115.2 6012 ppl214 1351439 1352398 P - unknown MarR family 3.5.2 Similar to conserved hypothetical 5116.3 6013 ppl215 1352456 1353019 P - unknown protein 5.2 5554.3 6187 ppl216 1353408 1353659 P + unknown hypothetical gene 6 5555.3 6188 ppl217 1353674 1354309 P - unknown 6 6052.1 6326 ppl218 1354320 1354946 m - unknown similar to unknown protein 5.2 Similar to thiocyanate hydrolase 5201.2 6038 Ippl219 1354960 1355628 m - unknown gamma subunit 4.2 Simimlar to thiocyanate hydrolase 5202.2 6039 Ippl220 1355628 1355936 m - unknown alpha subunit 4.2 Similar to thiocyanate hydrolase 5204.2 6040 ppl221 1355937 1356362 m - unknown beta subunit 4.2 4444.2 5595 ppl222 1356712 1358019 P - unknown similar to unknown protein 5.2 oxygen-dependent 4442.1 5594 ppl223 1358164 1359099 m - hemF coproporphyrinogen III oxidase 2.5 Flagellar basal-body rod 4441.1 5593 ppl224 1359315 1359707 P - flgB protein FlgB 1.5 Flagellar basal-body rod 4440.1 5592 ppl225 1359710 1360132 P - figc protein FlgC 1.5 Flagellar basal-body rod 1863.2 4038 ppl226 1360143 1360820 P - flgD modification protein FlgD 1.5 1862.2 4037 ppl227 1360934 1362247 P - flgE Flagellar hook protein FlgE 1.5 flagellar biosynthesis 801.3 6460 ppl228 1362258 1363004 P - flgF protein FlgF 1.5 flagellar biosynthesis 802.3 6461 ppl229 1363155 1363940 P - flgG protein FlgG 1.5 flagellar L-ring protein 803.3 6462 ppl230 1363953 1364645 P + flgH precursor FlgH 1.5 flagellar P-ring protein 5206.1 6041 ppl231 1364684 1365787 P + flgl precursor Flgl 1.5 flagellar biosynthesis 3142.2 4764 ppl232 1365799 1366674 P - flgj protein FlgJ 1.5 flagellar hook-associated 1532.3 3826 ppl233 1366724 1368673 P - flgK protein 1 1.5 flagellar hook-associated 1533.1 3827 ppl234 1368677 1369912 P - flgL protein FlgL 1.5
Table XIV 1246.2 3655 lppl235 1370107 1371846 unknown 6 1244.2 3654 Ippl236 1372499 1373563 unknown 6 Similar to conserved hypothetical 1417.1 3758 Ippl237 1373611 1374066 unknown protein 5.2 similar to putative transport 1418.3 3759 Ippl238 1374169 1375941 m unknown protein 1.2 Similar to electron transfer 2954.4 4646 Ippl239 1376030 1377661 m unknown flavoprotein-ubiquinone oxidoreductase 1.4 601.5 6319 Ippl240 1377897 1379708 i inknn n Similar to multidrug resistance ABC transporter ATP-binding protein 1.2 Similar to conserved hypothetical 598.2 6313 Ippl241 1379791 1380108 + unknown protein 5.2 597.2 6311 Ippl242 1380144 1380515 P + unknown 6 596.3 6310 Ippl243 1380572 1382287 m sfcA malate oxidoreductase 2.1.1 2955.2 4647 Ippl244 1382367 1382750 P unknown 5.1 Acid phosphatase SurE 2958.2 4648 Ippl245 1382753 1383511 surE (Stationary phase survival protein ) 2.6 novel lipoprotein homolog 2959.1 4649 Ippl246 1383530 1384273 p nlpD NlpD 1.1 RNA polymerase sigma 2960.1 4651 Ippl247 1384359 1385384 p rpoS factor RpoS 3.5.1 Homogen 2961.1 4652 Ippl248 1385487 1386737 m
imπΔ tisate 1,2- dioxygenase 2.2 Similar to conserved hypothetical 2962.1 4653 Ippl249 1386884 1387627 p unknown protein 5.2 Crossover junction 2495.2 4398 Ippl250 1387627 1388151 p ruvC endodeoxyribonuclease ruvC 3.3 holliday junction DNA 2493.2 4397 Ippl251 1388148 1388747 p ruvA helicase 3.3 2492.2 4396 Ippl252 1388797 1389117 p unknown 6 938.3 6550 Ippl253 1389158 1390762 m unknown 6 937.3 6549 Ippl254 1391044 1392459 m unknown similar to sensor histidine kinase 1.3 similar to two component response 1589.3 3861 Ippl255 1392456 1393133 m unknown regulator 3.5.2
Table XIV similar to intracellular septation 1590.1 3863 ppl256 1393194 1393739 m + unknown protein 1.7 Similar to membrane bound lytic 1591.4 3864 ppl257 1393982 1395421 m + unknown murein transglycosylase 1.1 hydroxyacylglutathione 2963.2 4654 ppl258 1395535 1396299 m gloB hydrolase (glyoxalase II) 2.5 Similar to conserved hypothetical 2964.1 4655 ppl259 1396303 1397151 m unknown protein 5.2 2965.1 4656 ppl260 1397437 1397646 p unknown 6 methylenetetra hyd rofolate dehydrogenase/methenylte 2966.1 4657 ppl261 1397649 1398503 m FolD trahydrofolate cyclohydrolase 2.5 2967.1 4658 ppl262 1398590 1398790 m unknown 6 2968.3 4659 ppl263 1399059 1401752 P fimV FimV protein 1.8 Similar to conserved hypothetical 5455.1 6139 ppl264 1401804 1402271 m unknown protein 5.2 Similar to conserved hypothetical 2156.3 4197 ppl265 1402271 1403569 m unknown protein 5.2 tRNA-pseudouridine 1740.3 3960 ppl266 1403813 1404601 truA synthase I 3.6 N-(5 - 1739.2 3958 ppl267 1404602 1405225 trpF phosphoribosyl)anthranilat e isomerase 2.2 tryptophan synthase beta 1738.3 3957 ppl268 1405227 1406426 P trpB subunit 2.2 tryptophan synthase, alpha 1547.2 3833 ppl269 1406452 1407270 P trpA subunit 2.2 1545.3 3832 ppl270 1407267 1408922 P glnS glutamine tRNA synthetase 3.7.2 3311.1 4868 ppl271 1408932 1410302 P cysS cysteine tRNA synthetase 3.7.2 2235.2 4232 ppl272 1410481 1412394 P unknown Similar to hypothetical sulfatase 4.7 3312.1 4869 ppl273 1412524 1412838 P unknown 6 type II protein secretion 5534.3 6177 ppl274 1412939 1414423 m IspE ATPase LspE 1.6
Table XIV type II protein secretion 4197.2 5431 ppl275 1414427 1416802 m IspD LspD 1.6 1448.2 3774 ppl276 1416975 1418003 p unknown Similar to oxydoreductase 4.6 1449.3 3775 ppl277 1418024 1420129 m + unknown Similar to adenylate cyclase 2.3 Similar to drug resistance 4195.3 5430 ppl278 1420273 1421433 m + unknown transporter, MFS superfamily 1.2 Similar to multidrug resistance 5100.2 6009 ppl279 1421430 1422644 m unknown efflux pump protein 1.2 similar to FrgA (Iron- and Fur- regulated gene), iron repressed 3378.2 4902 ppl280 1422637 1424379 m unknown gene; promotes intracellular infection; similar to aerobactin svnthetases 4.1 3376.1 4901 ppl281 1424512 1425546 m unknown similar to unknown protein 5.2 366.2 5088 ppl282 1425986 1426300 p unknown 6 Some similarities with eukaryotic 368.2 5102 ppl283 1426380 1428782 m unknown protein 6 3374.1 4900 ppl284 1428910 1430046 m unknown 6 periplasmic serine protease 982.3 6577 ppl285 1430644 1432044 m htrA Do; heat shock protein HtrA 4.1 Similar to conserved hypothetical 983.1 6578 ppl286 1432037 1432762 m unknown protein 5.2 Ribosomal large subunit pseudouridine synthase D 985.2 6579 ppl287 1432752 1433717 m rluD (Pseudouridylate synthase) (Uracil hydrolyase) Similar to 2-methylthioadenine 3371.2 4899 ppl288 1433717 1435060 m unknown synthetase Similar to major facilitator 4813.2 5840 ppl289 1435097 1436395 m unknown superfamily (MFS) transporter Similar to enhanced entry protein 4814.2 5841 ppl290 1436636 1437382 m unknown EnhA 4815.2 5842 ppl291 1437705 1438115 m fliS unknown Similar to flagellar protein FliS Similar to flagellar hook-associated 130.4 3689 ppl292 1438125 1439750 m fliD Unknown protein 2 (flagellar capping protein) 1.5 129.3 3684 ppl293 1439813 1440094 m unknown 6
Table XIV 126.2 3662 ppl294 1440177 1441604 m flaA flagelline 1.5 Similar to acetyl-CoA carboxylase 465.2 5724 ppl295 1442014 1442898 p accD unknown beta subunit 2.4 Similar to 464.2 5716 ppl296 1442879 1444165 p folC unknown dihydrofolate :folylpolyglutamate synthetase FolC 2.5 Similar to conserved hypothetical 462.2 5705 ppl297 1444162 1444950 p unknown protein 5.2 Similar to colicin V production 460.2 5693 ppl298 1444931 1445464 p unknown protein DedE 4.3 Similar to nicotinate-nucleotide 459.3 5687 ppl299 1445439 1446074 m nadD unknown adenylyltransferase NadD 2.5 Similar to DNA polymerase III, 458.3 5680 ppl300 1446062 1447087 m unknown delta subunit HolA 3.1 2943.1 4640 ppl301 1447089 1447580 m unknown Similar to rare lipoprotein B RlpB 1.2 2942.1 4639 ppl302 1447681 1450152 m leuS leucyl-tRNA synthetase 3.7.2 Similar to apolipoprotein N- acyltransferase (ALP N- 2941.1 4638 ppl303 1450218 1451753 m unknown acyltransferase) (copper homeostasis protein CutE) 3.8 1215.3 3639 ppl304 1451907 1452995 p unknown Similar to dehydrogenase 4.6 1220.3 3641 ppl305 1453005 1454525 p unknown Similar to aldehyde dehydrogenase 2.1.1 Similar to 3-hydroxyacyl-CoA 2103.2 4162 ppl306 1454645 1457014 p unknown dehydrogenase 2.4 Similar to 3-ketoacyl-CoA thiolase 1638.2 3893 ppl307 1457030 1458214 p fadA unknown (thiolase I, acetyl-CoA transferase) 2.4 1639.4 3894 ppl308 1458327 1459502 m unknown 6 SidG protein, substrate of 1635.4 3891 ppl309 1459664 1462561 m sidG the Dot/Icm system 4.6 5481.1 6151 ppl310 1462790 1463920 p unknown similar to unknown protein 5.2 regulatory protein (GGDEF and EAL 4665.2 5736 ppl311 1464059 1466437 m Unknown domains) 1 type II secretory pathway 4288.2 5485 ppl312 1466752 1467720 m IspK protein 1 type II secretory pathway 4289.2 5486 ppl313 1467707 1468324 m IspJ protein LspJ 1
Table XIV type II secretory pathway 4291.1 5488 ppl314 1468321 1468698 m + Ispl protein Lspl 1.6 type II secretory pathway 770.3 6447 ppl315 1468695 1469180 m - IspH protein LspH 1.6 type II secretory pathway 771.4 6448 Ippl316 1469167 1469589 m - IspG protein LspG 1.6 type II secretory pathway 772.4 6449 Ippl317 1469693 1470892 m - IspF protein LspF 1.6 774.3 6450 Ippl318 1470989 1472398 P - glnA glutamine synthetase 2.2 Similar to conserved hypothetical 775.2 6451 Ippl319 1472411 1472971 P + unknown protein 5.2 Similar to conserved hypothetical 4293.2 5489 ppl320 1472984 1473748 P - unknown protein 5.2 similar to 1-aminocyclopropane-l- 5482.2 6152 ppl321 1473778 1474677 P - unknown carboxylate deaminase 2.2 3909.3 5233 ppl322 1474889 1476466 P - unknown 6 Class III heat-shock 3911.2 5234 ppl323 1476668 1478539 P ~ htpG protein HtpG(molecular chaperone) 4.1 2382.2 4317 ppl324 1478651 1478947 m - unknown Similar to DNA-binding protein Fis 3.5.2 2381.2 4316 Similar to lipid-A-disaccharide ppl325 1479063 1480217 m - IpxB unknown synthase 1.1 2377.2 4314 ppl326 1480204 1481166 m - unknown Similar to oxidoreductase 2.1.1 2375.3 4313 ppl327 1481387 1481962 P - rnhB unknown Similar to ribonuclease HII 3.1 -determining 3913.2 5235 ppl328 1482074 1483192 m + Rod shape rodA protein rodA 1.7 3914.3 5236 ppl329 1483189 1485042 m + mrdA penicillin-binding protein 2 1.1 2239.3 4233 ppl330 1485268 1486245 P - unknown Similar to hypothetical protein 5.2 Similar to conserved hypothetical 2240.2 4234 ppl331 1486508 1486978 m - unknown protein 5.2 Similar to conserved hypothetical 2241.2 4235 ppl332 1486987 1487325 m - unknown protein 5.2 Similar to proton/peptide 4384.2 5555 ppl333 1487430 1488872 m - unknown symporter family protein 1.2 Similar to urocanate hydratase 4385.1 5556 ppl334 1489095 1490759 P hutU unknown (urocanase) (imidazolonepropionate hydrolase) 2.2
Table XIV 4386.2 5557 Similar to histidine ammonia-lyase ppl335 1490773 1492293 P hutH unknown (Histidase) 2.2 4867.2 5869 ppl336 1492286 1493695 P unknown Similar to aldehyde dehydrogenase 2.1.1 4868.2 5870 ppl337 Similar to short chain 1493707 1494516 P unknown dehydrogenase 2.1.1 4869.3 5871 ppl338 1494513 1494944 P rnhA unknown Similar to ribonuclease HI 3.1 5549.2 Similar to DNA polymerase III, 6183 ppl339 1494949 1495650 P dnaQ unknown epsilon chain 3.1 5548.1 6182 ppl340 1495857 1496234 m unknown 6 5546.1 6181 ppl341 1496526 1496999 P unknown Some similarity with EnhA protein 5.1 4874.2 5874 ppl342 1497118 1498521 P unknown 6 Similar to conserved hypothetical 4875.1 5875 ppl343 1498587 1498979 m unknown protein 5.2 Similar to tRNA (5- 4877.2 5876 ppl344 1499114 1500199 m trmU unknown methylaminomethyl-2- thiouridylate)-methyltransferase 3.6 4629.2 5711 ppl345 1500469 1500897 P unknown 6 50S ribosomal subunit 4628.2 5710 ppl346 1500983 1501174 P rpmF protein L32 3.7.1 4627.2 Fatty acid/phospholipid 5709 ppl347 1501180 1502208 P plsX synthesis protein 2.4 3-oxoacyl-[acyl-carrier- 4626.1 5708 ppl348 1502205 1503158 P fabH protein] synthase III 2.4 4625.2 Malonyl CoA-acyl carrier 5707 ppl349 1503185 1504132 P fabD protein transacylase 2.4 3-oxoacyl-[acyl-carrier 4803.2 protein] reductase (3- 5832 ppl350 1504144 1504890 P fabG ketoacyl-acyl carrier protein reductase) 2.4 4804.1 5833 ppl351 1504975 1505223 P acp Acyl carrier protein (ACP) 2.4 3-oxoacyl-[acyl-carrier- 4805.1 5834 ppl352 1505243 1506481 P fabF protein] synthase II (Beta- ketoacyl-ACP synthase II) 2.4 4806.4 Similar to conserved hypothetical 5835 ppl353 1506482 1507480 P unknown protein 5.2
Table XIV 6042.1 6325 ppl354 1507480 1508118 P tmk unknown Similar to thymidylate kinase 2.3 Similar to DNA polymerase III, 4958.3 5921 ppl355 1508115 1509020 P unknown delta subunit 3.1 Similar to type 4 fimbrial 4957.2 5920 ppl356 1509131 1509487 pilZ unknown biogenesis protein PilZ 1.8 Similar to putative 1346.3 3714 ppl357 1509547 1510335 lidH unknown deoxyribonuclease belonging to the TatD DNAse family 5.2 Similar to conserved hypothetical 1345.3 3713 ppl358 1510447 1511547 P unknown proteins 5.2 Similar to major facilitator 3517.3 4990 ppl359 1511603 1512895 P unknown membrane proteins 1.2 1450.2 3776 ppl360 1513413 1514693 m unknown similar to multidrug translocase 1.2 Similar to dolichol-phosphate 3519.1 4991 ppl361 1514924 1516090 P unknown mannosyltransferase 3.8 3520.1 4992 ppl362 1516178 1517563 m unknown similar to glycosyl transferase 3.8 3521.2 4993 ppl363 1517653 1518804 P unknown similar to putative choline kinase 2.4 Similar to conserved hypothetical 3522.1 4994 ppl364 1518857 1519627 m unknown proteins 5.2 3524.1 4995 ppl365 1519873 1520469 P unknown similar to unknown protein 5.2 3525.1 4996 ppl366 1520874 1521530 P adk adenylate kinase 2.3 3527.1 4997 ppl367 1521539 1521901 P unknown Similar to thioredoxin proteins 1.4 glycerol-3-phosphate 4228.2 5448 ppl368 1521861 1523381 m glpD dehydrogenase 2.1.1 4227.1 5447 ppl369 1523443 1524918 P glpK Glycerol kinase 2.1.1 1059.2 3546 ppl370 1525012 1526283 m gltA citrate synthase 2.1.3 Similar to purine nucleoside 1060.2 3547 ppl371 1526509 1527390 P unknown phosphorylase proteins 2.3 DNA gyrase, subunit A, 4226.2 5446 ppl372 1527474 1530065 P gyrA type II topoisomerase 3.4 similar to phosphoserine 2395.3 4324 ppl373 1530058 1531146 P serC unknown aminotransferase 2.2 3-phosphoshikimate 1- 1611.6 3877 ppl374 1531152 1532453 P aroA carboxyvinyltransferase 2.2 5876.3 6290 ppl375 1532453 1533145 P key Cytidylate kinase 2.3 6039.1 6323 ppl376 1533211 1534887 P rpsA 30S ribosomal protein SI 3.7.1
Table XIV Similar to conserved hypothetical 5588.2 6203 Ippl377 1535001 1535291 P + unknown protein 5.2 4489.3 5621 Ippl378 1535338 1536507 P - unknown similar to unknown protein 5.2 similar to polysaccharide 4488.1 5620 Ippl379 1536771 1537886 P - unknown biosynthesis protein 1.1 orotidine 5' -phosphate 4487.1 5619 Ippl380 1537887 1538576 P - pyrF decarboxylase 2.3 1795.3 3998 Ippl381 1538766 1541417 P - unknown 6 Similar to short-chain 4800.2 5829 Ippl382 1541362 1542141 P - unknown dehydrogenase 5.2 4801.1 5830 Ippl383 1542147 1542629 P + unknown 6 4802.2 5831 Ippl384 1542746 1543282 P - unknown 6 Similar to 4-hydroxybenzoate- 5587.3 6202 Ippl385 1543279 1544127 P - ubiA unknown octaprenyltransferase 2.5 5103.4 6010 Ippl386 1544261 1544659 m - unknown 6 5104.3 6011 Ippl387 1544739 1546511 m + unknown Similar to oxidase 2.1.1 Similar to 2-deoxyribose-5- 1794.3 3997 Ippl388 1546565 1547329 m - deoC unknown phosphate aldolase 2.3 Similar to xanthosine 1793.3 3996 Ippl389 1547326 1548165 m - xapA unknown phosphorylase 2.3 Similar to cytidine/deoxycytidine 5457.1 6141 Ippl390 1548178 1548573 m - unknown deaminase Cdd 2.3 5003.2 5949 Ippl391 1548612 1549415 m - unknown 6 5005.3 5950 Ippl392 1549592 1550959 m + cpxA Sensor histidine kinase 1.3 transcriptional regulatory 5456.1 6140 Ippl393 1550952 1551632 m - cpxR protein CpxR 3.5.2 Similar to magnesium and cobalt 3712.2 5126 Ippl394 1551640 1552518 m - unknown efflux protein CorC 1.2 similar to conserved hypothetical 3711.1 5125 Ippl395 1552490 1552966 m - unknown protein 5.2 similar to phosphate starvation- 3710.1 5124 Ippl396 1552963 1553913 m - unknown inducible protein PhoH 2.6 3709.1 5122 Ippl397 1554192 1555031 P - unknown similar to TrpH protein 5.2 Similar to putative translation 3708.1 5121 Ippl398 1555028 1555648 P - unknown factor 3.5.1 tryptophanyl-tRNA 3706.2 5120 Ippl399 1555894 1557108 P - trpS synthetase TrpS 3.7.2
Table XIV similar to segregation and 3705.1 5119 ppl400 1557108 1557899 p scpA unknown condensation protein A 3.4 similar to segregation and 3703.1 5118 ppl401 1557901 1558482 p scpB unknown condensation protein B 3.4 Similar to ribosomal large subunit 3702.1 5117 ppl402 1558472 1559218 p rluB unknown pseudouridine synthase B (Pseudouridylate synthase) 3.6 Similar to transcriptional regulator, 3701.1 5116 ppl403 1559481 1560239 p Unknown LuxR family 3.5.2 3699.3 5114 Ippl404 1560625 1563237 P unknown 6 4242.3 5458 Ippl405 1563333 1564316 P unknown similar to oxidoreductase 2.1.1 4243.3 5459 Ippl406 1564441 1564890 m unknown similar to unknown protein 5.2 4244.1 5460 Ippl407 1565108 1565473 P unknown 6 4246.2 5461 Ippl408 1565512 1566363 P unknown Similar to hydrolases 2.1.1 1323.2 3700 Ippl409 1566490 1567008 P unknown 6 Similar to probable multidrug efflux 1322.2 3699 ppl410 1567130 1568515 m + unknown protein 1.2 Similar to Legionella pneumophila 1525.3 3820 ppl411 1568544 1569800 m unknown putative phospholipase C Similar to 23S rRNA (Uracil-5-)- 1528.4 3821 ppl412 1569921 1571255 P rumA unknown methyltransferase RumA 3.6 1531.3 3825 ppl413 1571261 1573465 P relA GTP pyrophosphokinase 2.3 Similar to conserved hypothetical 1530.2 3824 ppl414 1573506 1573871 m unknown protein 5 Similar to putative 1529.2 3822 ppl415 1573930 1575123 unknown aminotransferase 3327.1 4876 ppl416 1575128 1575937 unknown similar to unknown protein Singli e-stranded-DNA- 3328.1 4877 ppl417 1575927 1577666 recJ specific exonuclease RecJ 3.3 Similar to tRNA-dihydrouridine 3331.1 4879 ppl418 1577745 1578737 P unknown synthase A DusA 3.6 Preprotein translocase, 1584.3 3857 ppl419 1578838 1581528 P ~ secA secretion protein SecA subunit 1.6 3333.1 4880 ppl420 • 1581525 1581929 P - mutT Mutator protein MutT 3.2 3334.2 4881 ppl421 1582158 1582976 P + unknown 6
Table XIV similar to conserved hypothetical 5021.3 5957 ppl422 1583021 1583767 m unknown protein 5.2 5019.1 5955 ppl423 1584005 1584610 m unknown Similar to dephospho-CoA kinase 2.5 5018.2 5954 ppl424 1584582 1585055 m + unknown 6 4696.3 5754 ppl425 1585052 1586650 m + Unknown regulatory protein (EAL domain) 1.3 similar to serine-type D-Ala-D-Ala 4694.1 5753 ppl426 1586789 1588435 p unknown carboxypeptidase 1.1 adenosylmethionine-8- 4692.5 5752 Ippl427 1588756 1590078 m bioA amino-7-oxononanoate aminotransferase 2.5 1598.6 3868 Ippl428 1590151 1591098 P bioB Biotin synthase 2.5 8-amino-7-oxononanoate 1595.3 3867 Ippl429 1591099 1592241 P bioF synthase 2.5 Biotin biosynthesis protein 1594.2 3866 Ippl430 1592225 1592944 P bioH BioH 2.5 2472.2 4381 Ippl431 1592941 1593579 P bioD Dethiobiotin synthetase 2.5 Similar to conserved hypothetical 2473.1 4382 Ippl432 1593714 1594028 P unknown protein 5.2 Similar to conserved hypothetical 2474.1 4383 Ippl433 1594025 1594654 P unknown protein 5.2 4901.3 5892 Ippl434 1594673 1596454 P aspS Aspartyl-tRNA synthetase 3.7.2 Similar to potassium efflux system 4900.3 5891 Ippl435 1596429 1599380 P unknown kefA 1.2 similar to DNA mismatch repair 4376.3 5549 Ippl436 1599343 1600017 P mutH unknown protein mutH 3.2 4377.4 5550 Ippl437 1600090 1601130 m unknown 6 971.2 6569 Ippl438 1601329 1602165 m Unknown 6 similar to eukaryotic serine 972.3 6570 Ippl439 1602311 1603900 P Unknown threonin protein kinase 3.8 4891.2 5887 Ippl440 1603954 1604703 m Unknown weak similarity to myosin 6 4889.1 5885 Ippl441 1604962 1605363 m Unknown similar to unknown protein 5.2 Similar to transcriptional regulator 4888.1 5884 Ippl442 1605449 1605970 m unknown (Lrp family) 3.5.2 Similar to acetyltransferase, GNAT 4887.3 5883 Ippl443 1606090 1606620 P unknown family 2.1.1 5075.4 5994 Ippl444 1606872 1609445 P unknown 6
Table XIV 5071.2 5993 Ippl445 1609655 1610680 P unknown 6 similar to adenylate cyclase, family 4452.2 5596 Ippl446 1610846 1612171 P unknown 3 2.3 Some similarity with eukaryotic 4453.2 5597 Ippl447 1612347 1613732 unknown proteins 5.2 spectinomycin 4454.4 5598 Ippl448 1613921 1614919 m aph phosphotransferase 4.2 6036.1 6322 Ippl449 1615172 1615504 P unknown Similar to hypothetical protein 5.1 5584.2 6200 Ippl450 1615585 1616649 m Unknown 6. 4993.2 5941 Ippl451 1616882 1617259 m unknown 6 4992.2 5940 Ippl452 1617348 1617641 P unknown 6 4990.3 5939 Ippl453 1618016 1619824 P unknown Some similarities with sidE protein 5.1 2466.3 4376 Ippl454 1620044 1622725 P unknown Similar to aminopeptidase N 2.2 4584.3 5683 Ippl455 1622860 1623783 m unknown 6 5559.3 6190 Ippl456 1623870 1624412 m unknown 6 5560.3 6191 Ippl457 1624461 1625102 m unknown Similar to oxidoreductase 2.1.1 5563.1 6192 Ippl458 1625664 1626128 m unknown Similar to acetyltransferase 2.1.1 5564.1 6193 Ippl459 1626462 1627886 m Ipd Lipoamide dehydrogenase 2.1.1 Pyruvate dehydrogenase 908.4 6529 Ippl460 1627975 1629609 m aceF (dihydrolipoyltransacetylas e component) E2p 2.1.2 Pyruvate dehydrogenase 907.4 6528 Ippl461 1629667 1632330 m aceE (decarboxylase component) Elp 2.1.2 4581.3 5682 Ippl462 1632358 1632747 m unknown Cystein rich protein 5.2 4580.2 5681 Ippl463 1632786 1633538 m unknown putative membrane protein 5.2 Similar to sodium/hydrogen 4578.1 5679 Ippl464 1633543 1634715 m unknown antiporter family protein 1.2 4577.2 5678 Ippl465 1635034 1635855 m unknown Similar to rare lipoprotein A RlpA 1.2 Similar to penicillin-binding protein precursor (D-alanyl-D- 4389.2 5560 Ippl466 1635982 1637274 + unknown alaninecarboxypeptidase fraction C) 1.1
Table XIV Similar to D-alanine 4391.1 5561 ppl467 1637287 1638123 P ala unknown aminotransferase 2.2 Similar to conserved hypothetical 4392.1 5562 ppl468 1638127 1638390 P unknown protein 5.2 Legionella secretion system member of a putative type I 4393.1 5563 ppl469 1638390 1638989 P IssX protein X secretion system 1.6 Legionella secretion system member of a putative type I 4394.4 5564 ppl470 1638999 1641035 P IssY protein Y secretion system 1.6 Legionella secretion system member of a putative type I 5052.4 5978 ppl471 1641162 1641767 P IssZ protein Z secretion system 1.6 Legionella secretion system member of a putative type I 5050.2 5977 ppl472 1641822 1642553 P IssA protein A secretion system 1.6 Legionella secretion system member of a putative type I 3838.3 5196 ppl473 1642643 1644799 P IssB protein B secretion system 1.6 Legionella secretion system member of a putative type I 3840.1 5198 ppl474 1644803 1645939 P IssD protein D secretion system 1.6 regulatory protein (GGDEF and EAL 3841.2 5199 ppl475 1646075 1648603 P IssE unknown domains) 1.3 putative pyrimidine phosphoribosyl 843.2 6484 ppl476 1648605 1649174 m ppt unknown transferase 2.3 844.2 6485 ppl477 1649217 1649543 m unknown Similar to hypothetical protein 5.2 Similar to conserved hypothetical 845.4 6486 ppl478 1649726 1650469 m unknown protein 5.2 249.4 4394 ppl479 1650728 1652452 P pilB pilus assembly protein PilB 1.8 250.1 4400 ppl480 1652458 1653678 P pilC pilus assembly protein PilC 1.8 Type 4 prepilin-like 251.2 4406 ppl481 1653724 1654587 pilD proteins leader peptide processing enzyme 1.8 putative cAMP/cGMP binding 252.2 4411 ppl482 1654601 1655806 m protein 4.6 3898.2 5225 ppl483 1655914 1656507 m unknown 6 3899.2 5226 ppl484 1656523 1656978 m unknown 6 similar to conserved hypothetical 3901.2 5229 ppl485 1657112 1657927 unknown protein 5.2 2-methylcitrate 3902.3 5230 ppl486 1658047 1659495 m prpD dehydratase 2.1.1 3857.3 5206 ppl487 1659497 1660615 m prpC 2thylcitrate synthase 2.1.1
Table XIV 3855.1 5205 ppl488 1660831 1661616 m unknown similar to unknown protein 5.2 1398.2 3744 ppl489 1661702 1663702 m unknown Similar to unknown protein 5.2 similar to peptide transport 1399.3 3745 ppl490 1663695 1665182 m unknown proteins 1.2 Similar to glutamate-1- 3853.2 5204 ppl491 1665620 1666906 m hemL unknown sci i iiαiuci ιyuc-<-, -.-αi i in IUI i ιu Lose 2.5 3851.1 5203 ppl492 1667294 1667470 p unknown Similar to rubredoxin protein 1.4 similar to conserved hypothetical 3850.1 5202 ppl493 1667467 1667889 m unknown protein 5.2 similar to putative transport 3849.1 5201 ppl494 1667892 1668623 m unknown proteins 1.2 Poly(A) polymerase (PAP) 4863.4 5866 Ippl495 1668930 1670201 P pcnB (Plasmid copy number protein) 3.6 Similar to 2-amino-4-hydroxy-6- 4864.3 5867 Ippl496 1670198 1670623 P folK unknown hydroxymethyldihydropteridine pyrophosphokinase 2.5 similar to universal stress protein A

4865.2 5868 Ippl497 1670678 1671100 P Unknown UspA 4.1 Highly similar to GTP-binding 4987.2 5937 Ippl498 1671177 1672565 m Unknown proteins 4.6 5255.1 6059 Ippl499 1672570 1673724 m Unknown Conserved hypothetical protein 5.2 similar to conserved hypothetical 5254.1 6058 Ippl500 1673749 1674423 m Unknown protein 5.2 4712.2 5766 Ippl501 1674441 1675721 m hisS histidyl-tRNA synthetase 3.7.2 4711.1 5765 Ippl502 1675711 1676280 m Unknown hypothetical protein 5.2 similar to fimbrial biogenesis and 4710.1 5764 Ippl503 1676255 1677037 m Unknown twitching motility protein (type 4) 1.5 similar to conserved hypothetical 4709.1 5763 ppl504 1677218 1678366 m Unknown protein 5.2 similar to nucleoside diphosphate 4706.3 5762 ppl505 1678387 1678812 m ndk Unknown kinase 2.3 4705.3 5761 ppl506 1678947 1679831 m Unknown 6. 5253.1 6057 ppl507 1679905 1680528 P Unknown conserved hypothetical protein 5.2 5092.3 6003 ppl508 1680617 1681348 m Unknown 6. similar to UDP-2,3- 5093.3 6004 ppl509 1681360 1682088 m Unknown diacylglucosamine hydrolase 1.1
Table XIV similar to cell division inhibitor 5094.2 6005 Ippl510 1682085 1682795 m Unknown MinC (septum placement) 1.7 Highly similar to acyl-CoA 4525.2 5647 Ippl511 1682852 1684561 p fadD/lidS unknown synthetase, long-chain-fatty-acid— Co A ligase 2.4 Similar to arginine 3rd transport 4524.1 5646 Ippl512 1684609 1685325 m unknown system periplasmic binding protein 1.2 4523.1 5645 Ippl513 1685486 1686049 m Unknown similar to unknown protein 5.2 similar to para-aminobenzoate 2536.2 4422 Ippl514 1685976 1687298 m pabB unknown synthase, component I 2.5 similar to pyruvate dehydrogenase, 2535.2 4421 Ippl515 1687479 1688552 p unknown (El alpha subunit) 2.1.2 similar to pyruvate dehydrogenase 5250.2 6056 Ippl516 1688545 1689519 p Unknown El (beta subunit) 2.1.2 pyruvate dehydrogenase E2 5701.2 6256 Ippl517 1689522 1690634 p Unknown (dihydrolipoamide acetyltransferase) 2.1.2 3359.2 4890 Ippl518 1690827 1693025 p Unknown putative transmembrane protein 5.2 putative Pyruvate/2-oxoglutarate dehydrogenase complex, 1190.4 3626 Ippl519 1693198 1695342 p Unknown dihydrolipoamide dehydrogenase (E3) component 2.1.2 3360.1 4892 Ippl520 1695398 1696108 m + Unknown 6 3362.1 4893 Ippl521 1696186 1696911 m Unknown 5.2 similar to eukaryotic thiamine 3363.2 4894 Ippl522 1697183 1698127 p Unknown biosynthesis protein NMT-1 2.3 similar to C. burnetii thiamine 3365.2 4895 Ippl523 1698130 1699200 p unknown biosynthesis oxidoreductase ThiO 2.5 Similar to thiamine biosynthesis 3366.1 4896 Ippl524 1699193 1699396 p thiS unknown protein This 2.5 similar to thiamine biosynthesis 702.2 6414 Ippl525 1699399 1700187 p thiG 1 unknown protein ThiG 2.5
Table XIV similar to phosphomethylpyrimidine 701.2 6413 ppl526 1700184 1701650 P - thiDE unknown kinase/thiamin-phosphate pyrophosphorylase 2.5 similar to molybdopterin 699.3 6411 ppl527 1701631 1702770 P - Unknown biosynthesis protein 2.5 some similarities to 3 - 3367.1 4897 ppl528 1702799 1703632 m + Unknown nucleotidase/nuclease 2.3 2501.2 4401 ppl529 1703635 1704894 m + tolB TolB protein 1.2 2504.4 4402 ppl530 1704900 1705898 m - Unknown weakly similar to TolA protein 1.2 5426.2 6127 ppl531 1705895 1706350 m - tolR unknown Similar to TolR proteins 1.2 Similar to TolQ, involved in the 5427.1 6128 ppl532 1706359 1707033 m - tolQ unknown tonB-independent uptake of proteins 1.2 5428.1 6129 ppl533 1707061 1707456 m - Unknown hypothetical protein 5.2 Highly similar to Holliday junction 2559.3 4434 ppl534 1707460 1708470 m - ruvB unknown DNA helicase RuvB 3.3 Sigma factor RpoE (sigma 2560.3 4435 ppl535 1708617 1709180 m - rpoE 24) 3.5.1 5429.1 6130 ppl536 1709666 1709905 m - Unknown Conserved hypothetical protein 5.2 some similarities to 1698.2 3933 ppl537 1709962 1711020 P - Unknown aminomethyltransferases 2.2 some similarities to cytochrome 1697.2 3932 ppl538 1711051 1711599 m - Unknown B561 1.4 weakly similar to NADP-specific 1493.6 3799 ppl539 1711620 1716497 m - Unknown glutamate dehydrogenase 2.2 858.2 6493 ppl540 1716549 1717037 m - unknown similar to unknown proteins 5.2 859.2 6494 ppl541 1717024 1718412 m - unknown similar to aldehyde dehydrogenase 2.1.1 similar to phosphatidylcholine 2751.1 4535 ppl542 1718755 1719522 P - lidK Unknown synthase 2.4 2752.1 4536 ppl543 1719845 1720333 P + Unknown 6 2753.1 4537 ppl544 1720337 1720837 P + unknown 6 2754.2 4538 ppl545 1720862 1721419 P - Unknown similar to unknown protein 5.2 some similarity with Legionella 33 2755.1 4539 ppl546 1721515 1723533 m - Unknown kDa polypeptide 5.1 2757.1 4540 ppl547 1723741 1724190 m - rpll 50S ribosomal protein L9 3.7.1 2759.1 4541 ppl548 1724203 1725162 m - Unknown similar to unknown protein 5.2
Table XIV 30S ribosomal subunit 2760.1 4543 ppl549 1725116 1725343 m - rpsR protein S18 3.7.1 2761.1 4544 ppl550 1725357 1725695 m - rpsF 30S ribosomal protein S6 3.7.1 2763.2 4545 ppl551 1726004 1726201 p - Unknown similar to carbon storage regulator 3.5.2 2765.2 4546 ppl552 1726212 1727102 p + Unknown 6 similar to unknown protein, 2121.2 4172 ppl553 1727139 1727873 m + unknown possibly truncated 5.2 similar to alpha subunit of fatty- 1074.3 3556 ppl554 1727933 1729951 m unknown acid oxidation complex, 3- hydroxyacyl-CoA dehydrogenase 2.4 similar to beta-Subunit of fatty acid 1075.3 3557 ppl555 1729944 1731263 m unknown oxidation complex, 3-keto-acyl- CoA-thiolase 2.4 2120.2 4171 Ippl556 1731428 1732489 m unknown 6 402.2 5304 Ippl557 1733202 1734773 m unknown 6 Similar to transposase (ISNCY 401.2 5296 Ippl558 1735049 1736254 m unknown family) 4.5 400.1 5289 Ippl559 1736387 1737181 P Unknown 6 399.2 5282 Ippl560 1737211 1739634 P Unknown 5.1 2767.1 4547 Ippl561 1739721 1741109 P unknown 5.1 615.5 6337 Ippl562 1741778 1742419 m Unknown similar to transposase 4.5 Similar to transposase (IS21 5874.2 6289 Ippl563 1742601 1743623 P Unknown family) 4.5 Similar to transposase (IS21 6217.2 6352 Ippl564 1743620 1744414 P Unknown family) 4.5 Similar to transposase (ISL3 5328.1 6088 Ippl565 1744456 1744953 m Unknown family) 4.5 Similar to transposase (ISL3 4209.2 5437 Ippl566 1744987 1746135 m Unknown family) 4.5 Some similarities to eukaryotic 2298.2 4262 Ippl567 1746200 1747495 m Unknown proteins 6. 1035.3 3529 Ippl568 1747969 1749393 P plaB phospholipase 2.4 4473.4 5610 Ippl569 1749465 1750142 m Unknown similar to hypothetical proteins 5.2 4474.1 5611 Ippl570 1750144 1750518 m Unknown Similar to hypothetical proteins 5.2 similar to conserved hypothetical 4475.1 5612 ppl571 1750710 1751390 Unknown proteins 5.2
Table XIV similar to phosphoenolpyruvate 4477.2 5613 Ippl572 1751628 1753943 P - Unknown carboxylase 2.1.2 similar to MutT/nudix family 4479.2 5614 Ippl573 1754096 1754578 P - Unknown protein 3.2 gamma-glutamyl 4480.4 5616 Ippl574 1754629 1755882 m - pro A phosphate reductase 2.2 5090.2 6002 Ippl575 1755891 1756961 m - proB glutamate 5-kinase Similar to acetyltransferase, GNAT 5088.2 6000 Ippl576 1757251 1757676 m - Unknown family 2.1.1 5660.1 6239 Ippl577 1757883 1758863 P - Unknown similar to hypothetical proteins 5.2 5056.3 5980 Ippl578 1759030 1759365 P - unknown similar to unknown protein 5.2 5058.2 5981 Ippl579 1759597 1759836 m - Unknown Similar to transposase (IS5 family) 4.5 5059.3 5982 Ippl580 1759829 1760140 m - Unknown Similar to transposase (IS5 family) 4.5 Similar to transcription regulators 602.4 6320 Ippl581 1760325 1761272 m - Unknown (MerR Family) 3.5.2 similar to transcription regulator ' 604.3 6324 Ippl582 1761480 1762229 m - Unknown (MerR family) 3.5.2 Similar to acetyltransferase, GNAT 4623.2 5706 Ippl583 1762295 1762822 m - Unknown family 2.1.1 5054.2 5979 Ippl584 1762819 1763775 m - Unknown similar to hypothetical proteins 5.2 similar to multidrug resistance ABC 4004.2 5293 Ippl585 1763779 1765560 m - unknown transporter ATP-binding protein 1.2 similar to ATPase components of 4003.1 5292 Ippl586 1765553 1767013 m - Unknown ABC transporters with duplicated ATPase domains 1.2 Weakly similar to cytochrome C 1223.2 3643 Ippl587 1767380 1768255 P - Unknown family proteins 1.4 1224.2 3644 Ippl588 1768315 1769115 m + Unknown Similar to class-D beta-lactamase 4.2 similar to Cell division protein 4002.2 5291 Ippl589 1769165 1771075 m + Unknown Ftsl/penicillin-binding protein 2 1.7 Similar to transcriptional regulator 4000.2 5290 Ippl590 1771075 1771515 m - Unknown (MarR family) 3.5.2 4307.2 5499 Ippl591 1771686 1773005 m - Unknown 6 2261.3 4243 Ippl592 1773175 1773933 m - Unknown 6. similar to conserved hypothetical 1098.3 3571 Ippl593 1774067 1775083 m - Unknown proteins 5.2
Table XIV 1099.2 3572 ppl594 1775376 1776359 P unknown similar to unknown protein 5.2 4305.2 5498 ppl595 1776384 1776779 m Unknown 6 5582.4 6199 ppl596 1777014 1779221 m Unknown putative copper efflux ATPase 1.2 4088.2 5351 ppl597 1779397 1780290 P Unknown Similar to unknown protein 5.1 similar to conserved hypothetical 4087.2 5350 ppl598 1780384 1781658 m Unknown proteins 5.2 similar to conserved hypothetical 1111.3 3581 ppl599 1781652 1782488 m Unknown protein 5.2 similar to 3-beta hydroxysteroid 1109.3 3579 ppl600 1782473 1783453 m Unknown dehydrogenase/isomerase 2.4 Similar to 3-oxoacyl-[acyl-carrier- 1107.2 3578 Ippl601 1783455 1784438 m unknown protein] synthase III 2.4 4084.3 5349 Ippl602 1784588 1785754 P Unknown similar to glycosyl transferase 1.1 4083.4 5348 Ippl603 1785751 1786407 P unknown 6 1577.5 3851 Ippl604 1786587 1789487 m unknown Similar to oxidoreductase proteins 2.1.1 4332.1 5515 Ippl605 1789642 1790181 P Unknown similar to hypothetical proteins 5.2 similar to acetyltransferase, GNAT 4333.1 5516 Ippl606 1790235 1791083 P unknown family 2.1.1 4334.1 5517 Ippl607 1791087 1791977 P unknown similar to unknown protein 5.2 similar to permease of the major 4336.2 5518 Ippl608 1792193 1793407 P Unknown facilitator superfamily 1.2 4632.1 5712 Ippl609 1793420 1794679 m Unknown 6 similar to conserved hypothetical 4633.2 5713 Ippl610 1794827 1795384 P Unknown proteins 5.2 similar to acylaminoacyl-peptidase 4634.2 5714 Ipplδl l 1795402 1797372 m unknown proteins 2.2 C-terminal part of L. pneumophila 4181.1 5422 Ippl612b 1797582 1798151 m unknown sidB protein 6 Similar to N-terminal part of sidB 2354.2 4296 Ippl612a 1798221 1798835 m unknown protein 6 1748.3 3965 Ippl613 1799215 1800489 m thrC unknown similar to threonine synthase ThrC 2.2 similar to putative outer membrane 1749.2 3966 Ippl614 1800656 1801420 m Unknown proteins 5.2 Some similarities with L. 4179.2 5420 Ippl615 1801721 1803745 m Unknown pneumophila SidE protein 5.1
Table XIV similar to conserved hypothetical 1295.1 3687 ppl616 1803892 1804467 m + Unknown proteins 5.2 1294.2 3686 ppl617 1804467 1804997 m - Unknown similar to Cytochrome B561 1.4 similar to conserved hypothetical 1293.4 3685 ppl618 1805001 1805594 m + Unknown proteins 5.2 4337.3 5519 ppl619 1805622 1805879 m + Unknown putative secreted protein 5.2 similar to myo-inositol catabolism 4338.3 5520 ppl620 1806001 1806897 m - iolE Unknown protein iolE 2.1.1 similar to malonic semialdehyde 4339.1 5521 ppl621 1806900 1808771 m - iolD Unknown oxidative decarboxylase 2.1.1 Bifuncional protein similar to IolC 4340.2 5523 ppl622 1808810 1810696 m - iolCB Unknown (sugar kinase) and IolB similar to myo-inositol 2- 5386.1 6108 ppl623 1810704 1811654 m - iolG Unknown dehydrogenase 2.1.1 4786.2 5821 ppl624 1811711 1813126 m + Unknown similar to sugar-proton symporter 1.2 4785.1 5820 ppl625 1813578 1814693 m - Unknown 6. 4751.2 5795 ppl626 1814891 1816564 m Unknown similar to metalloprotease 2.2 1516.3 3815 ppl627 1816606 1817529 m - Unknown similar to hyothetical proteins 5.2 similar to dimethylarginine 1515.2 3814 ppl628 1817711 1818478 P - Unknown dimethylaminohydrolase 2.2 1513.4 3813 ppl629 1818457 1819794 P + Unknown similar to amino acid transporter 1.2 5022.2 5958 ppl630 1819930 1820967 P - Unknown similar to hypothetical proteins 5.2 3988.3 5280 ppl631 1821106 1822587 P - Unknown 6. similar to conserved hypothetical 3986.1 5279 ppl632 1822681 1823799 m - Unknown proteins 5.2 similar to putative transport 3985.1 5278 ppl633 1823960 1824556 P - Unknown proteins, MFS family 1.2 856.2 6491 ppl634 1824720 1825232 P - Unknown 6. 857.2 6492 ppl635 1825291 1827618 m + Unknown 2.2 similar to putative membrane 4908.3 5894 Ippl636 1827781 1833546 P + Unknown proteins 5.2 1671.3 3917 ppl637 1833794 1835197 m + Unknown 6. 3995.5 5286 Ippl638 1835358 1836725 m -. Unknown 6. 3994.5 5285 Ippl639 1836897 1837721 m - Unknown 6. 3993.2 5284 Ippl640 1837986 1838543 P - Unknown 6. 887.4 6511 Ippl641 1838869 1840971 m - Unknown 5.1
■ + l +
E E ε ε ε ε CL CL CL rv oo 00 cn in VD co rH 00 cn rr m o CO rr in in m rr CO o CO cn σ> m 00 CO (N 00 LO o in cn rr ΓM OO cn in ΓM in o 00 00 CO rH rH in ro VD ro rv VD rr rv in ΓM cn <N rr m VD rv CO o *?H oo rr rT VD VD Is- cn o rH rr m rr rr rr rr rT rr m in LO in ιn in m ιn in VD VD VD VD CO CO 00 00 00 00 00 00 00 00 00 00 00 CO 00 00 00 00 00 P rH in o VD 00 cn CO rH VO o o 00 r ΓM rH cn o o cn cn o 00 cn rH rv 00 00 rv vD in 00 rr rr 00 o ΓM VD ro oo in o 00 Ol co rH rH VD rr cn rr 00 ΓM rT cn vo m rH ΓM rr in VD rv CO o rH ro rr rr vo VD 00 cn o rH rr rr rr rT rr rr rr rT in in in in m tn in in in VO VO vo 00 00 00 CO 00 00 00 00 CO 00 00 00 CO 00 CO CO 00 00 CO rH rH rH rH rH rH H H rH rH rH rH rH rH rH rH rH rH (N 00 rr in VD Is- CO cn o ΓM OO rr in VD rv CO cn o rr rr rT rr r rr rr rr in in in in in in in m m in vo VD VD VD VD VD VD VD vo D VD VD vo vo vo VD VD VD vD vD rH rH rH rH rH rH rH H rH rH rH rH rH H rH rH rH rH CL Q. Q. Q. Q. Q. CL Q. CL Q. Q. Q. CL Q. Q. Q. Q. Q. O. Q. CL CL Q. Q. Q. CL CL CL Q. Q. Q. Q. CL CL CL Q. CL CL 00 00 rH o o rH cn rH Is- cn (M cn O rH cn rH (N 00 rr in oo in in m in rT O m rr rr VD Is- rv 00 cn cn cn cn ΓM O ΓM ΓM ΓM ΓM ΓM rH m in in cn CJl cn rr rr rr rr rr VD VD m in rr rr in rr rr rr rr 00 ro 00 rr rr rr rr rr
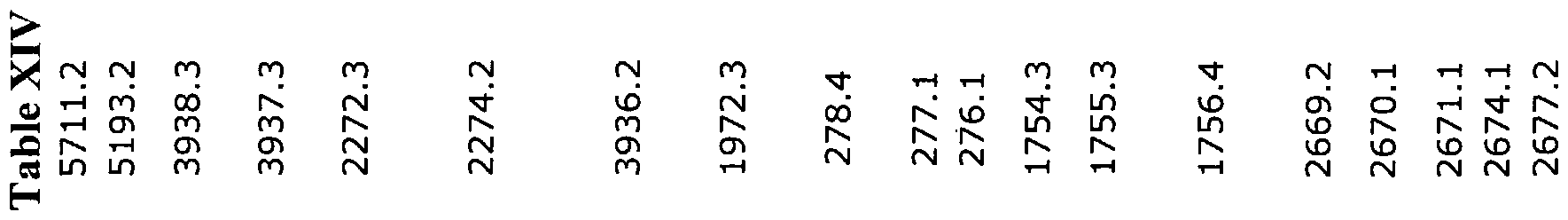
Table XIV similar to bifunctional PutA protein (proline dehydrogenase/ delta-1- 847.4 6487 Ippl661 1866148 1869300 m putA unknown pyrroline-5-carboxylate dehydrogenase) 2.2 2680.1 4496 Ippl662 1869857 1870534 p unknown conserved hypothetical protein 5.2 similar to activator of 1708.4 3941 Ippl663 1870966 1871331 p Unknown osmoprotectant transporter ProP (N-terminal part) 4.1 3-demethylubiquinone-9 3- 1709.1 3942 Ippl664 1871359 1872051 m ubiG methyltransferase 2.5 1711.2 3943 Ippl665 1872038 1872757 m unknown similar to Uracil-DNA glycosylase 3.2 2292.2 4258 Ippl666 1873027 1874706 p unknown 6. 2682.1 4497 Ippl667 1874842 1876482 p Unknown 6. similar to error-prone repair 1036.4 3530 Ippl668 1876569 1877825 m umuC Unknown protein 3.2 similar to error-prone repair: SOS- response transcriptional repressors 1037.1 3531 Ippl669 1877825 1878241 m umuD Unknown (LexA homologs, RecA-mediated autopeptidases) 3.2 similar to carboxypeptidase G2 and to acetylomithine 1039.2 3532 Ippl670 1878515 1879738 p Unknown deacetylase/Succinyl- diaminopimelate desuccinylase 2.2 arginine N- 2144.4 4187 Ippl671 1879738 1880778 p a st A succinyltransferase, beta chain 2.2 Succinylglutamic 2683.4 4498 Ippl672 1880792 1882279 p astD semialdehyde dehydrogenase 2.2 succinylarginine 5195.2 6034 Ippl673 1882285 1883631 p astB dihydrolase 2.2 2685.2 4499 Ippl674 1883643 1884368 p unknown similar to unknown proteins 5.2 4732.3 5781 Ippl675 1884446 1884811 m Unknown Weakly similar to cytochrome c5 1.4 1743.4 3962 Ippl676 1885233 1885790 m rrf Ribosome recycling factor 3.7.5
Table XIV Uridylate kinase (UK) 1742.3 3961 Ippl677 1885780 1886523 m - pyrH (Uridine monophosphate kinase) 2.3 Elongation factor Ts (EF- 4733.2 5782 Ippl678 1886520 1887398 m tsf Ts) 3.7.4 4999.2 5944 Ippl679 1887464 1888228 m - rpsB 30S ribosomal protein S2 3.7.1 5000.3 5947 Ippl680 1888562 1888972 P + Unknown 16 kD immunogenic protein 5.1 5001.1 5948 Ippl681 1889494 1889940 P - Unknown 6. 5307.3 6075 Ippl682 1890005 1891693 m - Unknown 6. 3991.3 5283 Ippl683 1892084 1893769 m - Unknown ankyrin repeat protein 6. similar to methionine 837.2 6479 Ippl684 1893865 1894629 P - Unknown aminopeptidase, type I 2.2 835.2 6478 Ippl685 1894616 1897201 P - glnD unknown similar to PII uridylyl-transferase 3.8 3989.1 5281 Ippl686 1897223 1897672 m - Unknown similar to hypothetical proteins 5.2 similar to GMP synthetase 1210.2 3636 Ippl687 1897663 1899240 m - guaA Unknown (glutamine-hydrolyzing) 2.3 similar to IMP dehydrogenase/GMP 1209.2 3634 Ippl688 1899243 1900715 m - guaB Unknown reductase 2.3 Septum site-determining 5309.1 6076 Ippl689 1900924 1901754 P - minD protein (Cell division inhibitor) 1.7 Septum formation 2514.3 4408 Ippl690 1901751 1902020 P minE topological specificity factor 1.7 893.4 6515 Ippl691 1902336 1904822 m + Unknown similar to acyl-CoA dehydrogenase 2.4 similar to putative hydrolases or 4472.2 5609 Ippl692 1904915 1905700 m - Unknown acyltransferases (alpha/beta hydrolase superfamily) 2.1.1 4744.3 5790 Ippl693 1905697 1906605 m - Unknown similar to hypothetical proteins 5.2 similar to sn-glycerol-3-phosphate 1503.4 3806 Ippl694 1906565 1907656 m - Unknown transport ATP-binding protein 1.2 similar to glycerol-3-phosphate 1501.3 3805 Ippl695 1907656 1908483 m + Unknown ABC transporter, permease component 1.2
Table XIV similar to glycerol-3-phosphate 2388.3 4320 Ippl696 1908480 1909358 m - Unknown ABC transporter, permease components 1.2 similar to quinone oxidoreductase 2387.3 4319 Ippl697 1909478 1910479 P - Unknown (NADPH :quinone reductase) 1.4 similar to Voltage-gated ClC-type 1144.3 3600 Ippl698 1910653 1911936 m - clcA unknown chloride channel 1.2 1145.3 3601 Ippl699 1911929 1914085 m - trpE Anthranilate synthase 2.2 Glutamyl-tRNA(Gln) 5311.1 6078 Ippl700 1914350 1914646 P - gate amidotransferase (subunit C) 3.7.2 Glutamyl-tRNA(Gln) 4094.2 5354 Ippl701 1914649 1916100 P - gatA amidotransferase (subunit A) Glutamyl-tRNA(Gln) 4093.1 5353 Ippl702 1916104 1917537 P - gatB amidotransferase (subunit B) 2343.2 4287 Ippl703 1917751 1918740 P - Unknown 6. similar to Adenylate cyclase 1(ATP 2344.4 4288 Ippl704 1918909 1921185 P - Unknown pyrophosphate-lyase 1; Adenylylcyclase 1) 2.3 5478.2 6148 Ippl705 1921225 1921686 P + Unknown 6 1115.4 3582 Ippl706 1921706 1922494 P + Unknown 6 5477.1 6147 Ippl707 1923066 1923353 m - Unknown similar to DNA-binding protein fis 3.5.2 similar to HesB/YadR/YfhF family 5476.1 6146 Ippl708 1923748 1924116 m - Unknown proteins 5.2 similar to iron-sulpher cluster 3556.2 5018 Ippl709 1924121 1924504 m - unknown proteins NifU 2.5 3557.1 5019 Ippl710 1924501 1925664 m - IscS unknown similar cysteine desulfurase 3.8 o putative tRNA/rRNA 427.3 5475 Ippl711 1925639 1926412 m - 1 U Iln 1k Mri similar t ιoi VΛV/n 1 1 methyltransferase 3.6 similar to inositol-1- 428.1 5480 Ippl712 1926528 1927313 P - Unknown monophosphatase 2.1.1 similar to putative signal peptide 429.2 5487 Ippl713 1927426 1928382 m - Unknown peptidases 1.6 endopeptidase Clp ATP- 2092.2 4159 Ippl714 1928471 1931047 P - clpB binding chain B (ClpB) 4.1 3559.2 5020 Ippl715 1931139 1932452 m - unknown 6.
Table XIV 1886.2 4052 ppl716 1932465 1933073 m Unknown similar to UDP-N-acetylmuramate: 1885.3 4051 ppl717 1933405 1934772 m mpl Unknown L-alanyl-gamma-D-glutamyl-meso- diaminopimelate ligase 1.1 4649.2 5723 ppl718 1934774 1935394 m unknown hypothetical gene 6 4650.4 5725 ppl719 1935447 1939268 m + unknown similar to unknown protein 5.2 4262.2 5471 ppl720 1939258 1939713 m fliJ unknown similar to flagellar protein FliJ 1.5 flagellum-specific ATP 2550.2 4429 Ippl721 1939710 1941059 m flil synthase Flil 1.5 Polar flagellar assembly 2551.2 4430 Ippl722 1941062 1941700 m fliH protein FliH 1.5 Flagellar motor switch 2553.3 4431 Ippl723 1941842 1942831 m fliG protein 1.5 4259.3 5469 ppl724 1942834 1944468 m fliF Flagellar M-ring protein 1.5 Flagellar hook-basal body 3675.1 5099 ppl725 1944641 1944955 m fliE complex protein 1.5 similar to two-component response 3676.2 5100 ppl726 1944970 1946283 m fleR unknown regulator 3.5.2 3679.3 5101 ppl727 1946307 1947338 m fleS unknown similar to sensor histidine kinase 3.5.2 561.3 6215 Ippl728 1947637 1948941 m unknown conserved hypothetical protein 5.2 similar to periplasmic chaperone 562.1 6217 ppl729 1948957 1949565 m unknown LolA 3.9 564.2 6228 ppl730 1949568 1951952 m ftsK unknown similar to cell division protein FtsK 1.7 3680.1 5103 ppl731 1952025 1952975 P trxB Thioredoxin reductase 1.4 similar to leucyl/phenylalanyl-tRNA- 3681.1 5104 ppl732 1952978 1953646 P aat unknown protein transferase 3.8 similar to rhodanese domain 3682.1 5105 ppl733 1953710 1954081 P Unknown protein 5.2 Translation initiation factor 3683.2 5106 ppl734 1954213 1954434 P infA IF-1 3.5.1 5295.1 6072 ppl735 1954598 1955956 m pmbA unknown 1.2 Similar to conserved hypothetical 2448.3 4364 Ippl736 1956002 1956778 P unknown protein 5.2 2447.3 4363 Ippl737 1957028 1957978 P Unknown 6.
Table XIV similar to ribonucleoside- 2445.4 4362 ppl738 1958250 1961078 P - rirl unknown diphosphate reductase, alpha subunit 2.3 Similar to ribonucleoside- 4797.1 5826 ppl739 1961092 1962195 P - rir2 unknown diphosphate reductase, beta subunit 2.3 4796.1 5825 ppl740 1962509 1962991 P + unknown 6 Highly similar to lysyl-tRNA 4005.2 5294 ppl741 1963078 1964568 m - lysS unknown synthetase 3.7.2 Highly similar to peptide chain 4009.1 5295 ppl742 1964565 1965572 m - prfB unknown release factor 2 3.5.4 4010.1 5297 ppl743 1965732 1966133 m - Unknown similar to hypothetical poteins 5.2 4012.1 5298 ppl744 1966130 1966951 m - motB unknown Similar to chemotaxis MotB protein 1.5 flagellar motor protein 4013.1 5299 ppl745 1966908 1967681 m - motA MotA 1.5 4014.1 5300 ppl746 1967691 1968407 m - fliA sigma factor 28 3.5.1 similar to flagellar synthesis 4015.1 5301 ppl747 1968504 1969373 m - fleN unknown regulator 1.5 Flagellar biosynthesis 4016.1 5302 ppl748 1969360 1970499 m - flhF protein FlhF 1.5 Flagellar biosynthesis 4017.4 5303 ppl749 1970540 1972618 m - flhA protein flhA 1.5 Flagellar biosynthetic 5361.2 6099 ppl750 1972615 1973763 m - flhB protein FlhB 1.5 Flagellar biosynthetic 2593.3 4452 ppl751 1973763 1974533 m - fliR protein fliR 1.5 Flagellar biosynthetic 5026.1 5959 ppl752 1974537 1974806 m + fliQ protein FliQ 1.5 Flagellar biosynthetic 2595.4 4453 ppl753 1974815 1975564 m + fliP protein FliP 1.5 5359.2 6097 ppl754 1975565 1975975 m - fliO Flagellar protein fliO 1.5 Flagellar motor switch 4526.3 5648 ppl755 1975972 1976307 m - fliN protein FliN 1.5 Flagellar motor switch 1585.2 3858 ppl756 1976317 1977378 m - fliM protein FliM 1.5 1586.2 3859 ppl757 1977823 1978062 m + Unknown 6. 1587.4 3860 ppl758 1978181 1979476 P - Unknown similar to oxidoreductase 2.1.1 similar to hypothetical 4275.2 5478 ppl759 1979473 1980195 P + Unknown oxidoreductase 2.1.1
Table XIVr similar to transcriptional regulator 4273.1 5477 ppl760 1980314 1981222 Unknown (LysR family) 3.5.2 2313.2 4270 ppl761 1981301 1982494 m Unknown 6. 2312.1 4269 ppl762 1982731 1982940 m Unknown 6. 2311.4 4268 ppl763 1983019 1985601 m alaS alanyl-tRNA synthetase 3.7.2 Regulatory protein 1873.2 4044 ppl764 1985619 1986071 m recX RecX 3.5.2 1871.2 4043 ppl765 1986064 1987110 m recA RecA protein 3.2 4643.2 5719 ppl766 1987413 1988348 m Unknown 6. 4642.1 5718 ppl767 1988642 1989136 m Unknown similar to hypothetical proteins 5.2 4640.2 DNA mismatch repair 5717 ppl768 1989757 1992297 mutS protein MutS 3.2 4020.2 5305 ppl769 1992310 1993998 P + Unknown similar to hypothtical proteins 5.2 830.4 6475 ppl770 1993995 1996535 P + Unknown similar to hypothetical proteins 5.2 similar to delta-aminolevulinic acid 832.3 6476 ppl771 1996569 1997564 hemB Unknown dehydratases (porphobilinogen synthase) 5747.1 6267 ppl772 1997701 1998093 P + unknown 6 Similar to long-chain fatty acid 1955.3 4095 ppl773 1998434 1999870 m + unknown transport protein 1.2 Similar to diaminopimelate 2328.2 4279 ppl774 2000192 2002771 p lysAC unknown decarboxylase, aspartate kinase (fusion of lysA and lysC) 2.2 Similar to UvrD/REP helicase family 3269.1 4842 ppl775 2002776 2006006 p unknown protein 3.1 2430.2 4352 ppl776 2006214 2007287 p unknown similar to unknown protein 5.2 Similar to conserved hypothetical 3267.1 4841 ppl777 2007403 2007957 p unknown protein 5.2 3265.1 Hydrogen peroxide- 4840 ppl778 2008005 2008895 m oxyR inducible genes activator 3.5.2 3263.2 Similar to major facilitator family 4839 ppl779 2009227 2010327 p unknown transporter 1.2 3260.2 4838 ppl780 2010329 2011276 m unknown 6 1633.2 Similar to tetraacyldisaccharide 4 - 3890 ppl781 2011281 2012252 m IpxK unknown kinase 2.4 Lipid A export ATP- 1632.3 3889 Ippl782 2012252 2014018 m msbA binding/permease protein MsbA 1.2
Table XIV 5278.3 6066 ppl783 2014105 2014887 m unknown 5277.3 Dihydroorotate 6065 ppl784 2014908 2015909 m pyrD dehydrogenase 2.3 3070.2 4723 ppl785 2016195 2017193 m unknown Predicted transmembrane protein 6 Similar to conserved hypothetical 3069.1 4721 ppl786 2017242 2017928 m unknown protein 5C2 3068.1 4720 ppl787 2018254 2019423 P unknown Similar to acyl-CoA dehydrogenase 2.4 Similar to acetyl-CoA 2538.2 4423 ppl788 2019435 2020619 P unknown acetyltransferase 2.4 3067.1 4719 ppl789 2020751 2021071 P Unknown hypothetical gene 6 similar to Acetyl/propionyl-CoA 568.2 6245 ppl790 2021081 2022688 P unknown carboxylase, beta subunit 2.2 similar to enoyl-CoA 569.2 6250 ppl791 2022672 2023502 P Unknown hydratase/isomerase 2.2 similar to Acetyl/propionyl-CoA 571.2 6257 ppl792 2023495 2025459 P Unknown carboxylase, alpha subunit 2.2 Similar to hydroxymethylglutaryl- 2007.1 4115 ppl793 2025470 2026378 P unknown CoA lyase 2.2 Similar to acetyl-coenzyme A 1127.3 3591 ppl794 2026535 2028475 P unknown synthetase 2.1.1 3065.1 4718 ppl795 2028635 2029051 m unknown similar to unknown proteins 5.2 Similar to ABC transporter, ATP- 3063.2 4717 ppl796 2029504 2030529 P unknown binding protein 1.2 Similar to ABC transporter 3061.2 4716 ppl797 2030507 2031154 P unknown permease protein 1.2 3059.2 4714 ppl798 2031167 2031946 P Unknown 29 kDa immunogenic protein 4.6 Some similarity with eukaryotic 1058.3 3545 ppl799 2032026 2033444 P unknown protein 6 1056.1 3544 pplδOO 2033770 2033937 P unknown 6 Similar to conserved hypothetical 1055.1 3543 pplδOl 2033934 2034665 m unknown protein 5.2 1053.2 3542 ppl802 2034668 2035198 m unknown Similar to phosphatase 2.6 glycyl-tRNA synthetase 1713.2 3944 Ippl803 2035204 2037270 m glyS beta chain 3.7.2 glycyl-tRNA synthetase 1714.4 3945 ppl804 2037260 2038165 m giyQ alpha chain 3.7.2
CO m C > C 0 Ul XI O J r 3 o 3 σι 3 o
+ '
E ε L ε ε ε ε CL E m 00 00 cn n rv ΓM rr in cn rr cn VD IN cn O (N ro rv rT rH Is- VD CO in 00 VD VD ro oo 00 H cn in rr ΓM VD VD VD vo VD (N 00 o in 00 VD r in rv rH iH ΓM in cn o r rT Is- o rv Is- rr 00 O Is- ΓM vo rH Is- cn rH ΓM 00 rT m VD vo 00 00 0s. cn o O ΓM oo rr 00 co ΓM rr in rv oo rr rr rr rr rr rr rr rr rr rr rr m in in in m in m VD VD VD VD o o O o o o o o o o o o o O o o o o o o o o o ΓM ΓM ΓM (N (N ΓM ΓM ΓM IN ΓM ΓM (N IN (N ΓM ΓM (N ΓM ΓM ΓM ΓM (N ΓM o ro cn IN rv CO rv rH o rv CO CO Is- o (N CM VD rT o Is- cn VD o cn oo 00 rv oo cn rH CO rr rr VO o 00 VD m rH VD rv Is- v o rH rT 00 r Is- ro ΓM IN VD o rH in rr rv rH rH Is- in 00 rH o Is- n 00 oo cn ΓM 00 rr in VD Is- 00 00 n cn o rH (N ro rr 00 cn ΓM rr in 00 00 rr rr rr rr rr rr rT rT rr rr rr LO in LO in in in in VD VD vo o o O o o O o o o o o o o O o o o o o o O o o ΓM ΓM IN (N ΓM ΓM (N (N ΓM ΓM IN IN ΓM (N (N ΓM ΓM ΓM (N ΓM ΓM ΓM (N in vD rv oo cn o rH ΓM 00 rr in VD rv 00 cn o rH ΓM ro H rH rr in VD Is- o o o o o H rH rH rH rH rH rH ΓM ΓM ΓM ΓM IN CM (N ΓM CO 00 CO 00 00 CO 00 CO CO 00 00 CO 00 00 00 00 00 00 00 CO CO 00 00 rH rH rH rH rH rH rH rH rH rH rH rH rH rH rH rH rH rH rH rH rH rH rH CL CL Q. CL Q. Q. Q. Q. Q. CL Q. CL Q. Q. CL α. O- Q. Q. CL Q. Q. Q. CL CL o. Q. Q. Q. α. Q. Q. Q. O. Q. Q. Q. Q. CL CL CL Q. Q. Q. Q. Q. cn (N 00 rr ro τ-1 o cn 00 ιn VD rv 00 IN rH o Is- cn 00 ΓM CO rv VD cn o Is- rr vo VD VD VD VD CO o o O O O o cn n cn cn 00 00 00 cn o m rH vo VO VD oo ro rH O o O O O o Is- σi cn cn VD VD VD 00 rT in VD ro OO oo VO VD m VO VO m in m in 00 r rr ro in in in
>
X rr in ro (N rr rH VD rH rH VD rH IN (N m ΓM rH oo H rH rr r rH rH
Q rv o cn o o 00 rv ΓM o ΓM o rv (N rH o cn o rH rv vo in in ΓM ΓM ΓM Is- CO VD in rr OO Is- M ΓM ro 00 00 σ> cn 00 ro 00 oo CO oo ro CO rH rH rH
Έ Is- 00 rr rr (N Γ ct r-H rH rr m rH tH rH VD ro o o in in in LO rr in in rv m in in VD in in 00 ro 00 OO rH 00 ro rH rr rf rr
H
Table XIV ATP-dependent Clp 4699.3 5755 ppl828 2068000 2069274 m clpX protease ATP-binding subunit ClpX 4.1 ATP-dependent Clp 1642.3 3896 ppl829 2069415 2070059 m clpP protease proteolytic subunit 4.1 peptidyl-prolyl cis-trans 1641.5 3895 ppl830 2070062 2071393 m tig isomerase (trigger factor) 3.9 4058.2 5329 ppl831 2071883 2072173 p Unknown 6. Similar to site specific 4056.2 5328 ppl832 2072264 2073490 p unknown recombinase, phage integrase family 4.4 Similar to ABC transporter, ATP- 4055.1 5327 ppl833 2073613 2075463 p unknown binding protein 1.2 4054.2 5326 ppl834 2075491 2076165 m rnc unknown Similar to ribonuclease III 2.3 4051.2 5325 ppl835 2076155 2076547 m + Unknown similar to unknown protein 5.2 4050.1 5324 ppl836 2076559 2077314 m + lepB Signal peptidase I 1.6 Similar to GTP-binding elongation 4048.3 5322 ppl837 2077422 2079254 m unknown factor 3.7.4 similar to membrane-bound lytic 4047.4 5321 ppl838 2079449 2080465 unknown murein transglycosylase B precursor similar to general secretion 4309.3 5500 lppl839 2080540 2081679. P unknown pathway protein L Similar to putative general 4310.1 5501 Ippl840 2081676 2082146 P Unknown secretion pathway protein 4311.1 5502 Ippl841 2082250 2082783 m unknown similar to unknown protein Similar to dGTP 4312.2 5503 Ippl842 2083058 2084383 P unknown triphosphohydrolase 1469.4 some similarity with eukaryotic 3786 Ippl843 2084470 2087877 P Unknown proteins 1082.3 3560 Ippl844 2088756 2090288 P Unknown Similar to alpha, alpha-trehalase 2.1.1 3737.1 5136 Ippl845 2090309 2091394 m Unknown 6. 3739.3 5137 Ippl846 2091465 2091905 m unknown Similar to lactoylglutathione lyase 2.1.1 3740.3 5139 Ippl847 2092140 2092667 P unknown putative membrane protein 5.2
Table XIV Some similarity with eukaryotic 3741.1 5140 Ippl848 2092904 2094121 P - Unknown proteins 5.2 3743.1 5141 Ippl849 2094443 2094757 P + Unknown 6. 3744.2 5142 Ippl850 2094922 2096031 m + Unknown 6 6318.1 6366 ppl851 2096462 2097547 Unknown Similar to transposase (IS4 family) 4.5 similar to conserved hypothetical 4220.2 5443 ppl852 2097573 2097866 unknown protein 5.2 4221.3 5444 Ippl853 2098044 2098319 m - unknown Similar to acylphosphatase 2.4 4222.3 5445 Ippl854 2098476 2098829 P - unknown Q-rich protein 6 1670.3 3916 Ippl855 2099011 2100342 m - unknown 6 1669.3 3915 Ippl856 2100571 2101536 P - unknown Similar to esterase/lipase 2.4 1673.5 3918 Ippl857 2101600 2103261 m - unknown 6 1675.3 3919 Ippl858 2103397 2103699 P - unknown Similar to hypothetical protein 5.2 5174.2 6025 Ippl859 2103753 2104172 P + unknown signal peptide predicted 6 Similar to major facilitator family 4080.2 5346 Ippl860 2104182 2105462 m + unknown transporter (MFS) 1.2 4078.1 5344 Ippl861 2105939 2107219 P - unknown similar to chloride channel protein 1.2 1735.1 3955 Ippl862 2107418 2107564 m - unknown 6 1737.2 3956 Ippl863 2107802 2108908 m - unknown Similar to hypothetical protein 5.2 4076.1 5343 Ippl864 2109110 2109613 P - unknown 6 4075.3 5342 Ippl865 2109877 2110356 P - unknown Similar to hypothetical protein 5.2 Similar to glutathione-regulated 3760.3 5151 ppl866 2110376 2112181 p unknown potassium-efflux system protein KefC (K(+)/H(+)antiporter) 1.2 3763.1 5152 ppl867 2112357 2112968 m unknown Similar to hypothetical protein 5.2 3764.3 5153 ppl868 2113061 2113363 m unknown 6 2427.4 4348 ppl869 2113758 2114093 p Unknown similar to unknown protein 5.2 2428.2 4349 ppl870 2114117 2114389 m Unknown Similar to transposase (IS5 family) 4.5 Similar to MoxR-like ATPases, 2429.2 4350 ppl871 2114625 2115539 p unknown putative regulator 3.5.2 Similar to conserved hypothetical 3765.1 5154 ppl872 2115536 2116486 p unknown protein 5.2 5173.1 6024 ppl873 2116486 2118474 p unknown Similar to unknown protein 5.2
Table XIV 3607.2 similar to conserved hypothetical 5051 ppl874 2118556 2119743 P - unknown protein 5.2 3608.1 5052 ppl875 2119851 2120177 P - unknown 6 1066.2 3550 ppl876 2120613 2121515 P - unknown similar to hypothetical protein 5.2 1067.1 3551 ppl877 2121681 2121923 m - unknown 6 1069.2 Similar to conserved hypothetical 3552 ppl878 2122150 2123079 m - unknown protein 5.2 2108.2 4165 ppl879 2123354 2124268 m - unknown similar to hypothetical transporter 1.2 similar to eukaryotic 1278.2 3675 ppl880 2124526 2125665 P + unknown ectonucleoside triphosphate diphosphohydrolase (apyrase) 2.3 Similar to conserved hypothetical 1279.3 3676 pplδδl 2125670 2126203 m - unknown protein 5.2 2905.2 4614 ppl882 2126214 2128019 m - unknown 6 2906.2 4615 ppl883 2128346 2128951 m - gst glutathione S-transferase 4.1 2908.2 4616 ppl884 2129148 2130143 m - unknown 6 2909.1 Similar to D-alanyl-D-alanine 4617 ppl885 2130184 2131446 m - unknown carboxypeptidase 1.1 Glutamyl-tRNA synthetase, 2910.1 4619 ppl886 2131693 2133105 m - gltX catalytic subunit 3.7.2 Legionella transmission 994.2 6585 ppl887 2133204 2135936 P + gacS/letS sensor Lets 1.3 993.2 6584 similar to PPi dependent ppl888 2136034 2137278 P - pfp Unknown phosphofructokinase 2.1.2 2170.1 4206 ppl889 2137275 2137688 m + unknown similar to type IV pilin PilA 1.8 2169.2 4205 ppl890 2137882 2138292 m + unknown similar to type IV pilin PilA 1.8 2911.3 4620 ppl891 2138595 2138912 P - unknown Similar to BolA protein 4.1 1142.4 3599 ppl892 2138987 2140408 P + unknown Similar to amino acid antiporter 1.2 1658.2 3907 ppl893 2140514 2141944 m - unknown weakly similar to endoglucanase 2.1.1 3-deoxy-manno- 1657.1 3906 ppl894 2142192 2142944 m kdsB octulosonate cytidylyltransferase 1.1 1655.2 3905 ppl895 2142941 2143120 m unknown Similar to hypothetical protein 5.2 Similar to conserved hypothetical 2914.1 4621 ppl896 2143113 2143784 m unknown protein 5.2
Table XIV Similar to ATP-dependent DNA 389.3 5220 Ippl897 2143762 2145720 m unknown helicase 3.2 390.2 5228 Ippl898 2145708 2146043 m unknown Similar to ferredoxin 1.4 2915.2 4622 Ippl899 2146325 2149117 P unknown 6 790.3 6457 Ippl900 2149421 2152066 P unknown 6 Similar to conserved hypothetical 4604.3 5697 ppl901 2152142 2152699 unknown protein 5.2 Similar to conserved hypothetical 4930.2 5908 ppl902 2152775 2153044 unknown protein 5.2 Similar to conserved hypothetical 4929.2 5906 ppl903 2153056 2153982 unknown protein 5.2 4927.1 5905 ppl904 2154197 2154922 P unknown 6 413.5 5379 ppl905 2155355 2157721 P unknown ankyrin repeat protein 6 similar to N-terminal part of 415.2 5397 ppl906 2158068 2159177 m Unknown putative transposase (IS91 family) 4.5 417.3 5412 Ippl907 2159832 2160641 P Unknown 6. 6314.1 6365 Ippl908 2160593 2160793 P Unknown 6. 2026.1 4124 Ippl909 2160742 2161014 m unknown 6 2027.2 4125 Ippl910 2161597 2162877 P unknown conserved hypothetical protein 5.2 2029.2 4126 Ippl911 2163280 2164302 P unknown 6 2849.1 4596 Ippl912 2165206 2165949 m unknown 6 2848.4 4595 Ippl913 2166319 2166573 P unknown 6 2847.4 4594 Ippl914 2166631 2167245 m unknown 6 1283.2 3679 Ippl915 2167395 2168690 m unknown similar to unknown protein 5.2 1284.2 3680 Ippl916 2168677 2168958 m unknown similar to transcription regulators 3.5.2 1286.3 3681 Ippl917 2169198 2171090 m unknown similar to methyl transferase 4.6 Similar to conserved hypothetical 1361.3 3721 Ippl918 2171105 2172148 m unknown protein 5.2 Similar to capsular polysaccharide 1360.6 3720 Ippl919 2172809 2174290 P unknown biosynthesis protein 1.1 Similar to N-terminal part of 2841.4 4593 Ippl920 2174603 2175919 m unknown polyketide synthase 4.6 similar to peptide antibiotic 2840.1 4592 Ippl921 2176089 2177612 m unknown synthetase 4.6
Table XIV 2838.1 4590 ppl922 2177629 2178756 m unknown similar to acyl-coA dehydrogenase 2.4 similar to 3-hydroxybutyryl-CoA 2835.1 4589 ppl923 2178826 2179680 m unknown dehydrogenase 2.4 1693.2 3930 ppl924 2179789 2180028 m unknown Similar to hypothetical protein 5.2 Similar to conserved hypothetical 1695.2 3931 ppl925 2180406 2181425 p unknown protein 5.2 similar to 3 ,5 -cyclic-nucleotide 2833.1 4588 ppl926 2181607 2182590 m unknown phosphodiesterase precursor (CpdP) 2.3 Similar to transcriptional regulator, 2831.1 4587 Ippl927 2182695 2183366 m unknown LuxR family 3.5.2 2830.3 4586 Ippl928 2183632 2184222 m unknown 6 5224.2 6045 Ippl929 2184210 2184572 m unknown 6 3419.3 4929 Ippl930 2184863 2185621 P unknown 6 3421.2 4931 Ippl931 2185721 2187055 m unknown 6 RalF protein, translocated 3423.3 4932 ppl932 2187286 2188482 ralF into host cells by the Dot/Icm system 4.6 Weakly similar to sphingosine 3426.1 4933 Ippl933 2188978 2189658 P unknown kinase 5.2 Similar to predicted 3428.1 4934 Ippl934 2189837 2190649 P unknown phosphohydrolases 2.3 3429.1 4935 Ippl935 2190832 2193066 m unknown 6 2250.2 4237 , Ippl936 2193506 2194144 m unknown 6 3431.4 4937 Ippl937 2194390 2195679 m unknown similar to other protein 5.2 similar to chloromuconate 3432.3 4938 Ippl938 2195679 2196782 m unknown cycloisomerase 2.1.1 1889.2 4054 Ippl939 2197085 2198338 P unknown 6 Some similarity with eukaryotic 1887.2 4053 Ippl940 2198679 2200283 P unknown proteins 5.2 1704.4 3937 Ippl941 2200547 2202541 P unknown 6 1703.1 3936 Ippl942 2202725 2203498 P unknown 6 5724.2 6262 Ippl943 2203535 2205046 m unknown 6 5723.2 6261 Ippl944 2205207 2205500 m unknown similar to unknown protein 5.2 6309.1 6364 Ippl945 2205526 2206611 m Unknown Similar to transposase (IS4 family) 4.5
Table XIV similar to peptidyl-prolyl cis-trans 4984.2 5936 Ippl946 2206829 2207395 P unknown isomerase proteins 3.9 5340.1 6091 Ippl947 2207576 2209132 m unknown 6 5341.1 6092 Ippl948 2209468 2209695 m unknown 6 Similar to transcriptional regulator, 5342.2 6093 Ippl949 2209957 2210517 P unknown TetR family 3.5.2 5344.2 6094 Ippl950 2210666 2211325 P unknown 6 6064.1 6328 Ippl951 2211382 2211624 m unknown hypothetical gene 6 4656.4 5729 Ippl952 2212041 2213834 m unknown 6. Similar to glutathione S- 4657.1 5730 Ippl953 2214269 2214892 m unknown transferase 4.1 Similar to organic hydroperoxide 4658.2 5731 ppl954 2214927 2215346 m unknown resistance protein 4.2 4659.2 5732 ppl955 2215573 2215950 m unknown 6 4660.2 5733 ppl956 2216182 2216343 P unknown 6 4662.1 5734 ppl957 2216796 2217038 P unknown 5.1 major outer membrane 4663.3 5735 ppl958 2217507 2218400 protein 1.1 Some similarity with eukaryotic 5901.3 6300 ppl959 2218765 2220192 unknown proteins. Pattern of chromosome condensation regulator conserved. 5.2 Similar to putative intracellular 4566.1 5671 ppl960 2220369 2220965 uι i rv.ι iv wi i protease/amidase 2.2 4567.2 5672 Ippl961 2221035 2222930 m unknown 6 5484.4 6153 Ippl962 2223205 2224569 P unknown 6 Some similarity with eukaryotic 1726.4 3952 Ippl963 2224659 2226323 P unknown proteins 5.2 4082.2 5347 Ippl964 2226507 2227706 P unknown 6 806.2 6463 Ippl965 2227726 2228463 P unknown Similar to hydantoin-racemase 2.2 807.2 6464 Ippl966 2228503 2229816 m unknown Similar to guanine deaminase 2.3 808.4 6465 Ippl967 2229998 2232826 m unknown 6 Similar to phosphohistidine 5602.3 6210 Ippl968 2233042 2233509 m unknown phosphatase SixA 3.8 Similar to conserved hypothetical 4365.4 5540 Ippl969 2233584 2234129 P unknown protein 5.2 4366.1 5541 Ippl970 2234228 2234644 P unknown putative membrane protein 6
Table XIV 4367.1 5542 Some similarity with eukaryotic ppl971 2234931 2235569 m - unknown proteins 5.2 4369.1 5543 ppl972 2235773 2236423 m - unknown predicted membrane protein 5.2 4370.1 5544 ppl973 2236503 2237003 m - unknown 6 Similar to putative polysaccharide 4371.1 5545 ppl974 2237495 2238355 p + unknown deacetylase-related protein 2.1.1 4372.3 5546 Similar to membrane-bound lytic ppl975 2238473 2239669 m + mltA unknown murein transglycosylase 1.1 5270.2 6064 ppl976 2239778 2240335 m + unknown 6 3133.2 4763 ppl977 2240332 2241138 m - unknown 6 3132.1 4762 ppl978 2241077 2241391 m - unknown 6 2146.2 4188 similar to histidinol-phosphate ppl979 2241551 2242660 p - hisC unknown aminotransferase 2.2 similar to pterin-4-alpha- 2147.1 4189 ppl980 2242697 2243038 p - unknown carbinolamine dehydratase phhB 2.5 Similar to protein-export 2148.1 4190 ppl981 2243216 2244133 m - secF unknown membrane protein SecF 1.6 Similar to protein-export 949.3 6558 ppl982 2244146 2246002 m + secD unknown membrane protein SecD 1.6 3131.1 4761 ppl983 2246024 2246359 m + unknown Similar to preprotein translocase 1.6 3130.1 4760 ppl984 2246505 2247338 m - unknown 6 S- adenosylmethionine:tRNA 3129.2 4759 ppl985 2247385 2248398 m - queA ribosyltransferase- isomerase 3.6 2509.3 4405 ppl986 2248513 2248926 m + unknown putative membrane protein 6 similar to ABC transporter ATP- 2510.2 4407 ppl987 2249008 2250861 m - unknown binding protein 1.2 Similar to conserved hypothetical 3128.1 4758 ppl988 2250922 2251413 m - unknown protein 5.2 Similar to putative 468.3 5745 ppl989 2251416 2251802 m - unknown endoribonuclease L-PSP 3.6 guanosine-3 ,5 - 469.3 5749 Ippl990 2251802 2253925 m ~ spoT bis(diphosphate) 3 - pyrophosphohydrolase 2.3
Table XIV RNA polymerase omega 470.3 5757 Ippl991 2254093 2254296 m rpoZ subunit 3.5.3 472.1 5773 Ippl992 2254347 2254976 -m gmk guanylate kinase 2.3 Similar to conserved hypothetical 473.2 5780 Ippl993 2254982 2255848 m unknown protein 5.2 Ribonuclease PH (RNase 3127.1 4757 Ippl994 2256150 2256857 p rph PH) (tRNA nucleotidyltransferase) Similar to twitching motility protein 3124.2 4756 Ippl995 2257186 2258220 m unknown Pi IT Similar to conserved hypothetical 3121.2 4755 Ippl996 2258384 2259076 P unknown protein similar to pyrroline-5-carboxylate 3119.1 4753 Ippl997 2259076 2259864 P proC unknown reductase Predicted integral membrane 5268.2 6063 Ippl998 2260208 2260783 P unknown protein 574.2 6265 Ippl999 2260831 2261445 P unknown Similar to conserved hypothetical 573.1 6263 Ipp2000 2261459 2261740 P unknown protein 5.2 572.2 6260 Ipp2001 2261894 2263582 m + unknown Similar to serine protease 2.2 Similar to transcriptional regulator, 4457.2 5599 Ipp2002 2263714 2264721 m unknown AraC family 3.5.2 Adenosylhomocysteinase 1626.3 3884 Ipp2003 2264845 2266170 m sahH (S-adenosyl-L- homocysteinehydrolase) 2.2 S-adenosylmethionine 1625.3 3883 Ipp2004 2266189 2267337 m metK synthetase 2.2 carbamoyl-phosphate 865.2 6499 Ipp2005 2267546 2268658 m carA synthetase, glutamine (small subunit) 2.3 chaperone protein DnaJ 866.2 ' 6500 Ipp2006 2268754 2269893 m dnaJ (heat shock protein) 4.1 Chaperone protein DnaK (Heat shock protein 70) 3999.3 5287 Ipp2007 2270082 2272016 m dnaK (Heat shock 70 kDa protein) (HSP70) 4.1 Heat-shock protein 3863.2 5209 Ipp2008 2272161 2272760 m grpE GrpE(HSP-70 cofactor) 4.1
Table XIV Similar to 3-deoxy-D-arabino- 3867.2 5210 Ipp2009 2273040 2274377 p unknown heptulosonate 7-phosphate (DAHP) synthase 2.2 Uroporphyrinogen 1161.2 3611 Ipp2010 2274675 2275733 m hemE decarboxylase 2.5 Similar to dihydroneopterin 1162.1 3612 Ipp2011 2275973 2276311 P folB unknown aldolase 2.5 1163.2 3613 Ipp2012 2276351 2277046 m unknown Similar to cell division protein 1.7 1325.2 3702 Ipp2013 2277046 2278815 m argS arginine tRNA synthetase 3.7.2 Similar to putative transport 1328.2 3703 Ipp2014 2278919 2279977 m unknown protein 1.2 ATP-dependent DNA 1330.5 3704 Ipp2015 2280124 2282196 P recG helicase RecG 3.3 5087.2 5999 Ipp2016 2282718 2283791 m unknown 6 Similar to cation-efflux family 3642.2 5076 Ipp2017 2284140 2285282 P unknown protein 1.2 3641.1 5075 Ipp2018 2285398 2286156 P unknown Similar to transporters 1.2 Similar to septum formation 2230.2 4228 Ipp2019 2286149 2286751 P unknown protein Maf 1.7 2231.3 4229 Ipp2020 2286783 2288051 P eno enolase 2.1.2 Similar to cell division protein ftsB 1949.4 4091 Ipp2021 2288273 2288542 P unknown homolog 1.7 1947.3 4090 Ipp2022 2288555 2289436 m unknown Similar to mevalonate kinase 2.4 Similar to diphosphomevalonate 3640.2 5074 Ipp2023 2289423 2290370 m unknown decarboxylase 2.4 Similar to organic radical activating 3638.2 5072 Ipp2024 2290388 2291041 m unknown enzymes 4.2 Similar to conserved hypothetical 3637.1 5071 Ipp2025 2291038 2292006 m unknown protein 5.2 Peptidoglycan-associated 3636.1 5070 Ipp2026 2292003 2292533 m pal lipoprotein precursor (19 kDa surface antigen) (PPL) 1.1 Similar to conserved hypothetical 3634.1 5069 Ipp2027 2292631 2293233 m unknown protein 5.2 Similar to ABC-type transport 3633.3 5068 Ipp2028 2293237 2294160 m unknown system involved in resistance to organic solvents 1.2
Table XIV Similar to ABC transporter ATP- 3477.2 4963 Ipp2029 2294190 2294927 m unknown binding protein 1.2 Similar to ABC transporter 3476.1 4962 Ipp2030 2294928 2296052 m unknown permease protein 1.2 3473.1 4961 Ipp2031 2296146 2296997 P unknown 6 3472.1 4960 Ipp2032 2297124 2297423 P unknown 6 1992.2 4110 Ipp2033 2297567 2298625 m unknown 6 Similar to isopentenyl-diphosphate 707.2 6417 Ipp2034 2298793 2299821 unknown delta-isomerase 2.4 Similar to 3-hydroxy-3- 705.1 6416 Ipp2035 2299812 2301110 unknown methylglutaryl-coenzyme A reductase 2.4 Weakly similar to mevalonate 704.4 6415 Ipp2036 2301107 2301985 P - unknown kinase 5.2 Similar to transposase (ISL3 5871.4 6288 Ipp2037 2302584 2303759 m - Unknown family) 4.5 Similar to poly(3- 374.2 5138 Ipp2038 2303825 2305591 m - unknown hydroxyalkanoate) synthetase 2.1.1 371.1 5123 Ipp2039 2305902 2306213 P - unknown 6 Similar to transposase (ISL3 1779.3 3985 Ipp2040 2306423 2306608 m Unknown family) 4.5 1780.3 3987 Ipp2041 2306952 2307161 P - unknown Similar to cold shock protein 4.1 Similar to ATP-dependent RNA 1781.4 3988 Ipp2042 2307401 2308807 P - dbpA unknown helicase 3.6 1592.3 3865 Ipp2043 2308964 2309875 P + unknown Similar to cation efflux protein 1.2 4285.1 5484 Ipp2044 2310147 2310548 m + unknown 6 4284.3 5483 Ipp2045 2310564 2311646 m - unknown similar to amidase 2.2 4282.3 5482 Ipp2046 2311680 2312429 m - Unknown Similar to transposase (IS5 family) 4.5 4281.2 5481 Ipp2047 2312539 2312808 P + unknown 6 4276.2 5479 Ipp2048 2313171 2315123 m - unknown 6 4532.2 5653 Ipp2049 2315629 2317071 P - unknown 5.1 4531.2 5652 Ipp2050 2317324 2317548 m - Unknown Similar to transposase (IS5 family) 4.5 Similar to conserved hypothetical 4530.1 5651 Ipp2051 2317759 2318052 m - unknown protein 5.2 C-terminal part similar to 4528.2 5649 Ipp2052 2318078 2318863 m - Unknown transposase (IS4 family) 4.5 5504.4 6163 Ipp2053 2319045 2319797 m - unknown 5.1
Table XIV 2357.4 4299 Ipp2054 2320236 2322056 P unknown 6 864.1 Similar to conserved hypothetical 6498 Ipp2055 2322300 2322590 m unknown protein 5.2 863.1 6497 Ipp2056 2322799 2323173 m Unknown Similar to transposase (IS5 family) 4.5 862.1 6496 Ipp2057 2323533 2323784 P Unknown 6. 860.2 6495 Ipp2058 2323929 2325671 P unknown Ankyrin repeat protein 5.2 3411.2 4922 Ipp2059 2326169 2327248 P unknown 6 3413.2 Similar to transcriptional regulator, 4923 Ipp2060 2327547 2328452 P unknown LysR family 3.5.2 3414.3 4924 Ipp2061 2328667 2330421 m unknown Ankyrin repeat protein 5.2 1910.6 4069 Ipp2062 2330640 2335121 m unknown 6 putative cAMP/cGMP binding 1911.4 4070 Ipp2063 2335445 2335906 m unknown protein 4.6 Similar to potassium uptake 1536.3 3828 Ipp2064 2335956 2337833 m unknown protein 1.2 1603.3 3873 Ipp2065 2338237 2339763 P unknown Putative ankyrin repeat protein 5.2 3878.1 5215 Ipp2066 2339887 2342958 m unknown 6 3876.1 5214 Similar to conserved hypothetical Ipp2067 2343282 2343770 P + unknown proteins 5.2 3874.1 5213 Ipp2068 2343839 2344093 m unknown 6 3872.1 5212 Ipp2069 2344289 2344555 P unknown 6 3871.1 5211 Some similarities with eukaryotic Ipp2070 2344580 2344969 P unknown proteins 6 5406.1 6119 Ipp2071 2345221 2346759 m Unknown regulatory protein (GGDEF domain) 1.3 proline/glycine betaine 1307.4 3693 Ipp2072 2346970 2348238 P tphB transporter-like protein 1.2 Chemiosmotic efflux 1306.5 3692 Ipp2073 2348345 2351545 m ceaA system protein A-like protein 1.2 Chemiosmotic efflux 5405.2 6118 Ipp2074 2351546 2352514 m ceaB system protein B-like protein 1.2 Chemiosmotic efflux 1001.3 3510 Ipp2075 2352511 2353824 m ceaC system protein C-like protein 1.2 1002.2 3511 Ipp2076 2354183 2355811 m unknown Similar to protein kinase 3.8
Table XIV r Similar to transcriptional regulator, 4742.3 5789 Ipp2077 2356163 2357077 P unknown LysR family 3.5.2 similar to pyridoxamine 5 - 4757.2 5799 Ipp2078 2357089 2357676 m unknown phosphate oxidase 2.5 4758.2 5800 Ipp2079 2357810 2358190 m unknown similar to unknown proteins 5.2 Similar to 3-oxoacyl-[acyl-carrier 4759.2 5801 Ipp2080 2358285 2359049 P unknown protein] reductase 2.4 4760.2 5802 Ipp2081 2359426 2360367 m unknown 6 Protein with ankyrin repeat and a F- 5065.2 5988 Ipp2082 2360600 2361064 P unknown box domain 4.6 Similar to two-component response 5066.2 5989 Ipp2083 2361148 2361567 m unknown regulator 3.5.2 1643.4 3897 Ipp2084 2361557 2364007 m stuC sensor histidine kinase 1.3 Similar to conserved hypothetical 4500.1 5631 Ipp2085 2364399 2364752 P unknown proteins 5.2 4498.1 5628 Ipp2086 2365217 2366665 P unknown 6 4497.4 5627 Ipp2087 2366762 2368021 P unknown 5.1 1937.4 4084 Ipp2088 2368109 2368468 P unknown 6 1939.5 4085 Ipp2089 2368702 2369274 P unknown 6 Similar to aminoglycoside 6- 5542.3 6180 Ipp2090 2369426 2370208 P Unknown adenylyltransferase 4.2 Similar to multidrug resistance ABC 3904.3 5231 Ipp2091 2370341 2372137 unknown transporter ATP-binding protein 1.2 SdeC protein, substrate of 440.4 5569 Ipp2092 2372425 2377026 P sdeC the Dot/Icm system 5.1 similar to L. pneumophila SdeA 3908.3 5232 Ipp2093 2377240 2378361 P unknown protein 5.1 3714.3 5127 Ipp2094 2378414 2381038 P unknown 6 substrates of the Legionella 817.7 6468 Ipp2095 2381318 2387080 P sdeB pneumophila Dot/Icm system SdeB 5.1 1933.4 4081 Ipp2096 2387451 2392088 P sdeA unknown 5.1 3003.1 4688 Ipp2097 2392571 2393194 P unknown Similar to methyltransferase 2.1.1 similar to (hydroxyindole) O- 3002.1 4687 Ipp2098 2393223 2394344 m unknown methyltransferase 2.2 2436.2 4355 Ipp2099 2394630 2395997 m unknown 5.1 similar to conserved hypothetical 3000.1 4686 Ipp2100 2396605 2398863 P unknown protein 5.2
vD VO VD VD
CZ CZ CZ CZ CZ CZ CZ CZ Z Z Z Z CZ Z CZ CZ Z CZ Z CZ CZ c c c c c 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 O O O O O O O O O O O O O O O O O O O O O O O O O O O CZ CZ CZ CZ C C CZ Z Z Z CZ CZ CZ Z CZ CZ Z Z Z CZ CZ CZ c c c c c .- -S- -S- - --- .Ϊ*- --_ .S(: - -V. ^. --; -S- -^ -Ϊ: --J _* .-•. __••: .•*: .-- -: .: _^ ^. --- s; c c c c c c c c c c c c c c c c c c c c c c c c c c 3 3 3 3 3 3 3 3 3 3 3 3 3 3 3 3 3 3 3 3 3 3 3 3 3 3 3
+ +
CL E CL CL E ε L CL ε ε ε ε ε ε E ε E ε ε ε Is- IN m in vo cn in o in o o rr cn rr VD vo ΓM rv TH oo co in CO m 00 ΓM Is- 00 cn VD cn ΓM Is- rr 00 00 o ΓM rr ΓM cn in OO rv in in CO rr 00 Is- in 00 00 vD 00 cn (N CO rr CO 00 O ΓM LO Is- LO VO Is- in Is- cn σi o ΓM ro rr rT in VD 00 CO n cn o o 00 rT rf rr rr VD rv 00 0s. cn o cn o o o o o o o o o o o o ΓM 00 rr rr rr rr rr rT rr rr rT rr rr rT rr rr rT rr rr rr rr rT rT rT rT rT rr rT IN ΓM ΓM ΓM ΓM ΓM ΓM ΓM IN ΓM ΓM (N ΓM ΓM (N CM ΓM ΓM ΓM ΓM ΓM ΓM (N OM ΓM (N ΓM cn rr cn VD rr ΓM (N cn ΓM ro (N ΓM cn cn o rv 00 o VD VD Is- cn rr 00 IN VD 00 (N in CO ΓM o CO 00 o oo VD LO O o ΓM rT cn ΓM ro cn cn o o CO rr CO m in o 00 o in cn ΓM cn rr O ro O (N in CO m CO in σ> o o IN ro rr in in r CO CO cn cn o ro rT rT rr rr vo 00 CO n o n o o o o o o o o o o o o o (N ro rr rr rr rr rr rT rr rr rr rr rT rr rr rT rT rT rT rT rT rT rr rT rT rr rr rr ΓM ΓM (N (M (N ΓM ΓM ΓM ΓM (N IN ΓM ΓM (N CM ΓM (N ΓM ΓM (N ΓM (M ΓM IN ΓM ΓM ΓM (N ro r m vo Is- 00 cn o ΓM 00 rT in vD Is- 00 n o (N ro rr in vo Is- o o o o o o o o o ΓM ΓM ΓM (N ΓM ΓM ΓM ΓM ΓM (N ΓM IN (N (N IN ΓM IN (N IN (N (N ΓM ΓM (N IN (N (N ΓM (N IN (N ΓM ΓM (N ΓM L Q. CL CL CL Q. CL Q. CL CL CL CL Q. CL L Q. Q. CL Q. α. Q. CL Q. CL α. CL Q. o. CL CL CL CO¬ o. α. Q. Q. Q. CL Q. α. CL CL Q. CL CL Q. Q. CL Q. CL Q. Q. CL L (N o cn CO 00 CO rv VD ιn s- rr ro ΓM 00 rv VD in rv rT r oo s- o cn o rr rr ιn ro CO CO CO I Is- rv I Is- Is- Is- Is- rv rv VD VO VD VD vo VO VD rr rT in rv rv VD VD VD vo o rr VD VD VD VD VD VD D VD D VD VO VD VD vo VO VD IN ΓM o ro r rT rr rr rr rr rr rT rr rT rT rr rr rT rr rr rr rr rT rT rr rr rT VD VD VD VD 00
X OO rH rH rH OO 00 fM (N (N r*J oi n ΓM ro rn oo oj n oo VD in co 00 rr ro IN o oo r VD in ro ΓM 00 VO in ro IN m r- ΓM Is- VD
-S cn cn cn cn -Ή ΓM cn cn cn cn cn oo oo oo oo 00 CO rv rv Is- P Is- rv Is- rr Is- rr
_θ n cn cn cn cn in cn cn cn cn cn cn cn n cn cn cn cn cn cn cn 0s. VD VO ΓM 00 rr
CJ fM IN (M (N rH ΓM (N ΓM (N ΓM ΓM IN ΓM ΓM ΓM ΓM ΓM IN (N ΓM (N IN m in in VD
Table XIV Similar to eukaryotic sphingosine-1- 1444.3 3772 Ipp2128 2421091 2422908 m unknown phosphate lyase 1 2.4 Similar to multidrug efflux RND 1870.3 4042 Ipp2129 2423146 2424252 P + unknown membrane fusion protein MexE 1.2 Similar to RND multidrug efflux 630.4 6363 Ipp2130 2424352 2427504 P unknown transporter 1.2 3205.3 4802 Ipp2131 2427666 2431985 P unknown similar to peptide synthase 4.6 Similar to two component sensor 3207.5 4803 Ipp2132 2432106 2434589 P unknown histidine kinase 1.3 4830.2 5850 Ipp2133 2434761 2435669 P unknown Similar to response regulator 3.5.2 Similar to eukaryotic SAM 4833.2 5851 Ipp2134 2435951 2436712 P unknown dependent methyltransferase 4.6 Similar to conserved hypothetical 5247.1 6055 Ipp2135 2436737 2437771 unknown protein 5.2 similar to polyketide synthase of 1205.5 3632 Ipp2136 2438029 2449374 P unknown type I 4.6 1207.2 3633 Ipp2137 2449377 2449778 m unknown 6 Similar to putative transport 2949.2 4641 Ipp2138 2450005 2450958 m unknown proteins 1.2 Some similarity with eukaryotic 2951.2 4643 Ipp2139 2451420 2453423 P unknown proteins 6 Present a domain similar to IcmL 2952.1 4644 Ipp2140 2453536 2455377 m unknown prtein 5.2 2953.2 4645 Ipp2141 2455913 2456413 P gspA global stress protein GspA 4.1 2204.2 4216 Ipp2142 2456595 2458115 P unknown Similar to sulfate transporters 1.2 Similar to carbonic anhydrase 2207.2 4217 Ipp2143 2458119 2458745 P unknown proteins 4.7 2208.1 4218 Ipp2144 2458863 2459156 m unknown hypothetical gene 6 2209.4 4219 Ipp2145 2459187 2460158 m unknown similar to ornithine cyclodeaminase 2.2 Similar to ketosteroid isomerase 2557.4 4433 Ipp2146 2460213 2460659 m unknown homolog 2.4 2555.5 4432 Ipp2147 2461033 2462187 P unknown 6 4613.3 5701 Ipp2148 2462381 2463862 P unknown Similar to transporters 1.2 4609.2 5700 Ipp2149 2463867 2464448 m unknown 6 4608.1 5699 Ipp2150 2464665 2465201 m unknown 6 4607.3 5698 Ipp2151 2465334 2466764 m unknown similar to unknown proteins 5.2
Table XIV 5610.1 6216 pp2152 2466798 2467241 m - unknown similar to unknown protein 5.2 Similar to conserved hypothetical 5609.1 6214 pp2153 2467211 2467495 m - unknown protein 5.2 Periplamic protein weakly similar to 5084.4 5998 pp2154 2467579 2469132 m - unknown AlgJ 5.2 Similar to alginate o- 5607.2 6213 pp2155 2469135 2470556 m - unknown acetyltransferase Algl 2.1.1 Predicted integral membrane 5606.2 6212 pp2156 2470772 2471410 P - unknown protein 6 1317.3 3696 pp2157 2471646 2472746 P + unknown 6 1318.5 3697 pp2158 2472794 2474017 m - unknown similar to unknown protein 5.2 4547.4 5662 pp2159 2474360 2475373 m - unknown Similar to oxidoreductase 2.1.1 similar to conserved hypothetical 4171.4 5414 pp2160 2475691 2476218 P + unknown protein 5.2 4172.2 5415 pp2161 2476367 2477428 m - unknown 6 similar to conserved hypothetical 4173.2 5416 pp2162 2477564 2477968 m - unknown protein 5.2 2526.3 4417 pp2163 2478168 2479442 P - unknown 6 Similar to hemin binding protein 4174.1 5417 pp2164 2479719 2480135 P + unknown Hbp 5.1 Similar to hypothetical sugar 4175.1 5418 pp2165 2480232 2481155 m - unknown nucleotide epimerase 2.1.1 4177.2 5419 pp2166 2481235 2482830 m - unknown 6 Some similarity with eukaryotic 5557.2 6189 pp2167 2482851 2484623 m - unknown protein 5.2 Some similarity with eukaryotic 4636.4 5715 pp2168 2485250 2489992 P - unknown protein 5.2 1903.4 4064 pp2169 2490228 2491202 m - unknown similar to chitinase 2.1.1 5796.5 6272 pp2170 2491410 2492495 P - unknown Similar to choline monooxygenase 2.1.1 549.6 6155 pp2171 2492656 2493777 P - unknown similar to unknown proteins 5.2 548.3 6150 pp2172 2493840 2495339 m + unknown 6 3306.4 4866 pp2173 2495482 2496201 m + unknown 6 3304.2 4865 pp2174 2496579 2497700 m + Unknown similar to unknown protein 5.2 3303.3 4864 pp2175 2497962 2499185 m - unknown 6 1088.4 3562 pp2176 2499394 2500923 m - unknown similar to other proteins 5.2 1087.2 3561 pp2177 2500930 2502621 m - unknown similar to acyl-CoA dehydrogenase 2.4
Table XIV similar to propionyl-CoA 3301.2 4863 Ipp2178 2502630 2504051 m unknown carboxylase beta chain 2.4 3299.4 4860 Ipp2179 2504273 2505046 m unknown similar to outer membrane protein 1.1 Similar to 3-oxoacyl-[acyl-carrier- 3297.3 4859 Ipp2180 2505791 2506852 P unknown protein] synthase III 2.4 1412.5 3754 Ipp2181 2506864 2508609 P unknown Similar to acyl-CoA synthetase 2.4 1413.2 3755 Ipp2182 2508614 2509993 P unknown Similar to acyl-CoA synthetase 2.4 1414.1 3756 Ipp2183 2509983 2510216 P unknown similar to acyl carrier protein (ACP) 2.4 Similar to 3-oxoacyl-[acyl-carrier 1415.4 3757 Ipp2184 2510213 2510965 P unknown protein] reductase 2.4 Similar to 3-oxoacyl-(acyl-carrier- 2900.2 4613 Ipp2185 2510962 2511972 P unknown protein) synthase III 2.4 2898.4 4610 Ipp2186 2512292 2512519 m unknown Similar to acyl-carrier protein 2.4 Similar to multidrug resistance 1840.3 4024 Ipp2187 2512529 2513752 m unknown protein 1.2 similar to sterol desaturase-related 1839.2 4022 Ipp2188 2513906 2515108 P unknown protein 2.4 2893.2 4609 Ipp2189 2515261 2517603 P unknown similar to nitric oxide reductase 4.2 similar to multidrug resistance ABC 2891.2 4608 Ipp2190 2517794 2519626 m unknown transporter ATP-binding protein 1.2 2889.2 4607 Ipp2191 2519788 2520174 m + unknown Putative membrane protein 5.2 2544.3 4425 Ipp2192 2520386 2524243 P unknown 6 Similar to conserved hypothetical 2268.2 4246 Ipp2193 2524479 2525687 P unknown protein 5.2 2266.2 4245 Ipp2194 2525774 2527042 P unknown Similar to GTP cyclohydrolase II 2.5 2264.2 4244 Ipp2195 2526993 2528276 P unknown similar to hypothetical protein 5.2 1239.2 3651 Ipp2197 2528262 2529245 P Unknown 6. Similar to uracil 1242.2 3653 pp2196 2528289 2528933 upp unknown phosphoribosyltransferase (upp) 2.3 1238.2 3650 Ipp2198 2529239 2532139 P unknown 6 Similar to C4-dicarboxylate 5188.2 6030 Ipp2199 2532404 2533696 m unknown transport protein 1.2 5189.2 6031 Ipp2200 2534017 2534526 m + unknown Similar to hypothetical protein 3235.4 4819 Ipp2201 2535075 2535731 P unknown DedA 5.2
CZ Z Z cz cz cz cz z cz cz cz cz cz c ra 5 5 5 5 5 5 5 5 5 5 5 5 5 5 c c cz cz z cz cz cz cz sz cz O O O O O O O O O O O O O o Is*. 5 5 5 5 5 5 5 5 5 5 o o o o o o o o o o CZ CZ Z z z cz z cz c cz cz cz z c --: ._. --- - -* .s<-: -¥ - --*- __- --_ ._<-: __<: ^ Φ C C C C C CZ CZ SZ SZ SZ Z Z CZ CZ CZ C CZ CZ CZ CZ CZ CZ CZ ro C C C C C CZ CZ CZ Z SZ 3 3 3 3 3 3 3 3 3 3 3 3 3 3 3 3 3 3 3 3 3 3 3 3 o ro
O --- 'S
ε ε ε ε ε ε ε ε E E o- ε CL E o. E cn 00 CO in vo VD VD ro rH rT rH ro rv in o cn VD D in VD Is- rH rH ro in ΓM rH o ro rr 00 rr Is- VD 00 rT rr rr vo 00 o CM CO m (N 00 rH 00 00 00 r-l VD o 00 rv O 00 VD O rr 00 cn ΓM Is- rr - rT o rH rr cn Is rr O CO vD ro CO 00 o OO 00 rr in VD rv CO 00 o rH IN oo rT rT in VD rv O O rH IN 00 00 rT rT rr rr rr rr rT rr rT rr in in in in in in in in in VD VD VD VO m LO ιn ιn in in in in LO in in in in in LO LO in in in in in in in LO LO CM IN ΓM (N (N (N ΓM (N IN ΓM ΓM ΓM (N IN IN ΓM ΓM ΓM ΓM ΓM ΓM ΓM ΓM (N IN m o 00 00 ro 00 00 (N 00 rr rH ΓM o cn ΓM cn IN oo rr 00 rv in o cn 00 Is- cn (N rr ro rv cn 00 CM 00 in 00 VD m vo rH rH IN o 00 ΓM ro oo 00 LO Is- rH cn IN VO o 00 CO in cn rv rH VD rT CO o cn IN cn rr cn rT o o 00 in 00 00 o rH ro ro rr in VD rv CO 00 o rH OO ro rT rr LO VD rv o rH rH 00 OO 00 rr rr rr rr rr rr rr rr rr rr LO in in in in m in in m VD VD VD in in m in LO in in in ιn in in in in in in in in in in in in m in in in ΓM ΓM (N IN IN IN IN ΓM ΓM ΓM ΓM ΓM ΓM CM ΓM (N (N ΓM ΓM ΓM ΓM (N (N (N ΓM ΓM 00 rr in VD Is- 00 σ> o rH (N oo rr in vo Is- 00 cn o rH ΓM ro o o o o o o o o rH rH rH rH rH rH H rH rH N (N ΓM (N n in ΓM- VD (N ΓM IN IN ΓM ΓM (N ΓM ΓM IN IN ΓM (N ΓM ΓM (N (N ΓM ΓM (N ΓM (N ΓM (N ΓM ΓM ΓM IN ΓM (N ΓM (N ΓM CM ΓM IN ΓM (N ΓM ΓM (N IN IN ΓM ΓM ΓM CM ΓM IN ΓM ΓM (N Q. o. O- CL Q. Q. CL CL CL Q. Q. Q. CL Q. CL CL Q. Q. Q. CL CL CL CL CL Q. Q. L CL Q. Q. Q. CL Q. Q. CL Q. CL CL Q. Q. Q. o. Q. Q. CL Q. CL L L o ΓM ro rr ro in rH ΓO rT in ΓM rr CO ΓM IN c (No Is- m rr 00 o cn 00 Is- VD VD 00 (M cn (N rr rT rr cn cn ΓM CO 00 ΓM ΓM ΓM o cn CO in in LO cn cn 00 in in in 00 en rr rr 00 ro ro OO rr rT rr rr rr cn 00 00 rr rr 00 CO 00 rT VD D vD 00 00 rr ro ro ro rr rr in in in in in oo ro 00 rr rr in in in
> ro rr rH (N 00 ΓM rH (N (N CM rH ro ΓM VD ro oo (N IN CO oi CO ΓM CO vo CO cn cn O (N rH
— vo VD 00 VD rr Is- in rr ro IN cn o rH
-5 oo ΓM ΓM (N CO n ro rH rH rH ro rr cn cn CO 00 oo rr rr rr 00 CO o rH rH IN n n cn rv Is- ΓM rH rH rH (N ΓM rH rH rH rH rH VD VD VD in in 00 00 CO
93 oo rH rH ro rH rH rH ro ro rT rT rT rr rT rH rH rH ΓM ΓM rr rr rT H
Table XIV Similar to glycerol-3-phosphate- 1768.3 3978 Ipp2227 2562490 2563803 m ugpB unknown binding periplasmic protein precursor 1.2 Similar to glycerophosphodiester 1767.2 3977 Ipp2228 2563800 2564519 m unknown phosphodiesterase 2.4 4826.2 5849 Ipp2229 2564524 2565219 m unknown Similar to hypothetical protein 5.2 4073.1 5341 Ipp2230 2565909 2566982 p unknown Similar to leucine dehydrogenase 2.2 4072.1 5340 Ipp2231 2566997 2567653 p unknown Similar to O-methyltransferase 2.1.1 4-hydroxyphenylpyruvate 997.2 6588 Ipp2232 2567720 2568766 p lly dioxygenase (legiolysin) 2.1.1 996.2 6587 Ipp2233 2568822 2569820 p unknown similar to unknown protein 5.2 Similar to glutathione S- 995.2 6586 Ipp2234 2569817 2570455 p unknown transferase (maleylacetoacetate
4071.1 5339 Ipp2235 2570452 2571012 m unknown
asparagine tRNA 4070.3 5338 Ipp2236 2571160 2572563 p asnS synthetase 3.7.2 4678.3 5744 Ipp2237 2572560 2572958 m unknown 6 similar to ATP-binding component 4680.2 5746 Ipp2238 2572982 2573665 m unknown of ABC transporter 1.2 Similar to ABC transporter, 4683.3 5747 Ipp2239 2573658 2574905 m unknown permease component 1.2 Similar to CDP-diacylglycerol— 5492.2 6156 Ipp2240 2575722 2576285 m pgsA unknown glycerol-3-phosphate 3- phosphatidyltransferase 2.4 4356.2 5536 Ipp2241 2576306 2576965 m unknown Similar to phosphatase 2.6 Ribosomal large subunit 4355.3 5535 Ipp2242 2576958 2577905 m rluC pseudouridine synthase 3.6 4354.3 5534 Ipp2243 2578258 2578455 P unknown 6 787.3 6455 Ipp2244 2578804 2580795 P rne Ribonuclease E 3.6 789.4 6456 Ipp2245b 2581017 2581268 m Unknown Similar to transposase (IS4 family) 4.5 6165.1 6342 Ipp2245a 2581389 2582213 m Unknown Similar to transposase (IS4 family) 4.5 1159.5 3608 Ipp2246 2582461 2583738 m unknown 6
Table XIV RNA polymerase- 1681.5 3921 Ipp2247 2584331 2587207 P - hepA associated protein HepA 3.5.3 similar to ankyrin repeat domain 248.2 4386 Ipp2248 2587321 2588724 m - unknown protein 5.2 247.1 4379 Ipp2249 2588725 2589498 m - unknown 6 Aspartate-semialdehyde 245.1 4365 Ipp2250 2589646 2590668 m - asd dehydrogenase 2.2 244.1 4358 Ipp2251 2590778 2591836 m - aroC chorismate synthase 2.2 ;imilar to putative adenine-specific 243.2 4351 Ipp2252 2591848 2592780 m - unknown methylase 3.2 similar to conserved hypothetical 2017.1 4118 Ipp2253 2592945 2593514 P - unknown protein 5.2 similar to conserved hypothetical 2016.2 4117 Ipp2254 2593587 2594006 P - unknown protein 5.2 3028.1 4703 Ipp2255 2594011 2594265 P - grx unknown Similar to glutaredoxin Grx 4.6 similar to protein-export protein 3027.1 4702 Ipp2256 2594275 2594763 P - secB unknown SecB 1.6 similar to glycerol-3-phosphate 376.3 5150 Ipp2257 2594765 2595754 P + gpsA unknown dehydrogenase (NAD+) 2.1.1 377.1 5155 Ipp2258 2595832 2596671 m - murl unknown similar to glutamate racemase 2.2 381.6 5177 Ipp2259 2596741 2600256 m - unknown 6 5899.1 6297 Ipp2260 2600402 2600704 P + unknown 6 Similar to conserved hypothetical 1823.5 4015 Ipp2261 2600708 2602000 m - unknown protein 5.2 dihydrodipicolinate 4346.2 5528 Ipp2262 2602255 2603127 m - dapA synthase 2.2 4347.2 5529 Ipp2263 2603339 2603593 P + unknown 6 Similar to 3-hydroxybutyrate 4349.2 5530 Ipp2264 2603871 2604653 P + unknown dehydrogenase 2.4 Similar to conserved hypothetical 4350.2 5531 Ipp2265 2604660 2605820 P - unknown protein 5.2 similar to proton conductor 4351.2 5532 Ipp2266 2605850 2606755 p motA2 unknown component of motor, chemotaxis and motility protein 1.5 4352.3 5533 Ipp2267 2606765 2607703 p motB2 unknown similar to flagellar motor protein 1.5 5986.2 6314 Ipp2268 2607769 2608242 m unknown 6 4691.2 5751 Ipp2269 2608487 2609782 p sdaC unknown Similar to serine transporter 1.2 4690.2 5750 Ipp2270 2609875 2611800 m unknown ankyrin repeat protein 6
Table XIV similar to type II secretion system 4688.2 5748 Ipp2271 2611887 2613008 m unknown protein-like protein and twitching mobility protein 1.6 405.3 5323 Ipp2272 2613098 2615026 m unknown 6 406.1 55333300 Ipp2273 2615313 2616149 m unknown Similar to hypothetical protein 5.2 407.4 5337 Ipp2274 2616146 2617498 m radA unknown Similar to DNA repair protein RadA 3.2 5420.2 6123 Ipp2275 2617733 2618611 p unknown 6 Unknown, N-terminal similar to 5422.1 6124 Ipp2276 2618876 2619259 p unknown Legionella 33 kDa polypeptide 5.1 Similar to exodeoxyribonuclease 5423.1 6125 Ipp2277 2619453 2619683 p xseB unknown VII, small subunit 2.3 Similar to geranyltranstransferase; 2225.2 4226 Ipp2278 2619664 2620560 p unknown farnesyl-diphosphate synthase 2.4 1128.3 3592 Ipp2279 2620604 2621458 m unknown Similar to biotin synthesis protein 2.5 Similar to competence protein 1132.2 3593 Ipp2280 2621502 2622206 p unknown ComF 1.2 Similar to membrane-associated 1133.3 3594 Ipp2281 2622235 2623302 m unknown metalloprotease proteins 2.2 1134.3 3595 Ipp2282 2623365 2623643 m unknown 6 3802.2 5173 Ipp2283 2623961 2625235 p hemA glutamyl tRNA reductase 2.5 peptide chain release 3803.1 5174 Ipp2284 2625219 2626307 p prfA factor 1 3.7.5 3804.2 5175 Ipp2285 2626300 2627163 p hemK unknown Similar to methyltransferase hemK 3.8 3807.2 5176 Ipp2286 2627307 2627738 p dksA unknown Similar to DnaK suppressor protein 4.1 1407.2 3750 Ipp2287 2627808 2628641 p unknown 6 3-Deoxy-D-manno-oct-2- 1406.2 3749 Ipp2288 2628646 2629905 m kdtA ulosonic acid transferase 1.1 1405.2 3748 Ipp2289 2630060 2630950 p djlA unknown Similar to DnaJ-like protein 3.9 5424.2 6126 Ipp2290 2630917 2631690 m + unknown 6 5134.4 6019 Ipp2291 2631882 2632811 m + plaA lysophospholipase A 2.4 5133.3 6018 Ipp2292 2633081 2633596 p unknown 6
Table XIV 5132.3 6017 Ipp2293 2633753 2633971 m unknown Similar to ATP-dependent RNA 3072.2 4724 Ipp2294 2634322 2636091 m deaD unknown helicase deaD (cold-shock DEAD- box protein A) Similar to conserved hypothetical 640.2 6377 Ipp2295 2636459 2637376 m unknown protein Similar to 2,4-dienoyl-CoA 641.3 6379 Ipp2296 2637373 2639397 m fadH unknown reductase 1618.3 3880 Ipp2297 2639702 2640190 P + Superoxide dismutase [Cu- sodC Zn] precursor Similar to alkyl hydroperoxide 1617.3 3879 Ipp2298 2640234 2640749 m unknown reductase AhpD Similar to alkyl hydroperoxide 2152.2 4195 Ipp2299 2640772 2641311 m unknown reductase AhpC 3074.1 4725 Ipp2300 2641511 2642362 P unknown 6 1880.3 4048 Ipp2301 2642460 2643452 m mdh Malate dehydrogenase 2.1.3 Similar to MutT/nudix family 2459.1 4371 Ipp2302 2643442 2644002 m unknown protein 2.3 Similar to oxygen-independent 2458.2 4370 Ipp2303 2644084 2645211 P hemN unknown coproporphyrinogen III oxidase 2.5 2162.2 4201 Ipp2304 2645243 2646652 m Unknown Similar to amidase 2.2 2164.2 4202 Ipp2305 2647006 2647839 P unknown Predicted membrane protein 5.2 Similar to O-sialoglycoprotein 1902.5 4063 Ipp2306 2648138 2649139 m gcp unknown endopeptidase 2.2 5369.2 6100 Ipp2307 2649344 2649583 P rpsU 30S ribosomal protein S21 3.7.1 Similar to conserved hypothetical 5370.1 6102 Ipp2308 2649786 2650229 P IporfX unknown protein 5.2 2282.4 4254 Ipp2309 2650238 2651971 P dnaG DNA primase 3.1 RNA polymerase sigma 1169.3 3614 Ipp2310 2652058 2653923 P rpoD factor rpoD (Sigma-70) 3.5.1 5920.3 6306 Ipp2311 2654264 2655439 m Unknown Similar to transposase 4.5 879.3 •6507 Ipp2312 2655505 2656599 m unknown Similar to integrase 4.4 876.1 6506 Ipp2313 2657086 2657556 m unknown 6 Similar to L.pneumophila 875.5 6505 Ipp2314 2657573 2658847 m unknown proline/betaine transport protein homolog CitA (TphA) 1.2 5474.2 6145 Ipp2315 2658825 2659337 m unknown Similar to acetyltransferase 2.1.1 3234.3 4818 Ipp2316 2659379 2660062 P unknown Similar to hydrolase 2.1.1
C C C C C C C C C C C C C C C C C Z SZ SZ Z Z z cz 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 o o o o o o O O O O O O O O O O O o o o o o o o o Z Z Z Z SZ CZ SZ C C CZ Z SZ CZ CZ CZ SZ Z SZ SZ SZ SZ Z SZ Z SZ -_ -s- --<; ^ --- --i ._<-: .-- _*: _>. -- ->*: _-<•: .__ .__ .__: _•-_: __-: ^ st: ^. ^ ^ ^ ^ C C C C C C C C C C C C C C C Z SZ SZ SZ SZ Z SZ 3 3 3 3 3 3 3 3 3 3 3 3 3 3 3 3 3 3 3 3 3 3 3 3 3
Q. Q. CL CL ε C CL ε ε IN LO rT IN VD in in VO VD in ro oo o cn oo in rH IN ro co VD cn rH o cn cn ΓM ΓM H (N 00 rr o ΓM cn CO VD o VD 00 cn O IN rv 00 CO vo o ΓM cn (N o cn o Is- 00 CO (N VO rH rr ro rr rr ro 00 o Is- VD CO O rH rH CM ΓM ro in Is- Is- 00 cn rH ΓM ΓM (N ro rT LO vD VD D VO vD Is- Is- rv rv rv rv Is- rv rv crnv crvn O rH rH VO VD VD VD D VD vo o V 00 00 CO VO VO VD VD VD VD vo O VD VO VD VD VD VD VD vo vo VD vD VD vD vo VD VD VD ΓM ΓM (N (N CM ΓM IN ΓM (N ΓM ΓM ΓM ΓM ΓM (N IN ΓM ΓM IN (N ΓM (N IN (N ΓM cn VD o rH rv ΓM oo in rH 00 LO ΓM LO in VD in vo vo vD rv cn ΓM O rT rH 00 Is- rr VO co 00 rH cn 00 rv cn CO rH rH cn CO o cn ΓM ΓM rr 00 cn Is- Is- o cn rT rv ΓM in VD vo rH o 00 rH VO o IN cn IN cn m rr rH o Is- VD rH o o rH ΓM ΓM 00 m Is- 00 cn o rH (N IN (N ro oo in VD Is- cn cn o rH vD VD VD VD VD VD VD vo VD VD VD rv rv rv Is- rv Is- Is- rv Is- Is- Is- Is- 00 00 VD VD VD VD VD o VD VO VD VD VD VD vo VD Vθ VD vo VD VD VD VD VD D VD VD ΓM ΓM fM ΓM ΓM ΓM (N ΓM ΓM ΓM ΓM ΓM (N ΓM IN (N ΓM IN ΓM ΓM (N ΓM ΓM ΓM ΓM rv CO cn o H ΓM o rr in VD ΓV 00 cn o rH (N ro rr LO VD Is- 00 rH rH rH IN ΓM (N ΓM ΓM ΓM ΓM (N ΓM ΓM ro 00 ro 00 oo 00 00 00 cn o rH 00 00 rr rT OO 00 00 00 ro oo ro 00 ro ro ro ro ro oo 00 ro oo 00 oo 00 00 00 ro ro 00 ΓM (N IN IN ΓM (N (N ΓM ΓM (N ΓM ΓM (N (N IN N ΓM ΓM (N ΓM (N IN (N ΓM ΓM CL D- CL Q. CL CL CL CL o. Q. CL CL CL Q. Q. C Q. Q. Q. CL Q. CL Q. L Q. CL CL Q. Q. α. CL Q. CO¬ CL CL Q. Q. Q. Q. Q. Q. L CL CL o. Q. Q. CL CL CL Is- VO in T (N o rr IN ro rH LO 00 rr in rT LO vD o rr CO V0 m VD r rH rH rH rH rH rH O cn (N rH Is- VD VD VD 00 00 00 Is- c Vn T D r- CO 00 CO cn in 00 CO 00 00 00 OO 00 rv rH CO rr Is- rv Is- CO 00 00 rr rr rr CO 00 CO o CO rT rr rr rT rT 00 ro ro rr rr rr 00 ro 00 ro 00 00 VD VD rr ro 00 oo rr 00
>
X ΓM 00 (N ΓM ΓM rH (N ΓM ΓM ΓM rH ΓM Γ ΓM rH 00 00 ΓM (N U ro ΓM rH o CO o O TH o' CO rr vo cn ro rr in 00 cn o ΓM (N ro ΓM ΓM ΓM rr rr LO c Is- H cn CO o ro (N ro 00 ΓM (N vo oό
IS ro o c (N (N ΓM (N (N LO in rr rH rH o (N vo rf rr rr LO in LO rHn rHό vo VD VD VD cn LO ro ro ro ro ro ΓM ro ΓM rH rH rH rH rH rH 00 00 ΓM rH rH rH rH rH
Table XIV 159.1 3862 Ipp2342 2681842 2682135 m unknown similar to unknown protein 5.2 160.1 3870 lpp'2343 2682331 2682654 P unknown 6 Similar to glutamate 163.1 3887 Ipp2344 2682882 2684180 m gdhA unknown dehydrogenase 166.2 3909 Ipp2345 2684358 2687552 m copA2 unknown Similar to copper efflux ATPase 168.1 3920 Ipp2346 2687471 2687998 m unknown predicted transmembrane protein 169.1 3927 Ipp2347 2688144 2688350 P unknown 592.3 6305 Ipp2348 2688363 2690042 m unknown 593.2 6308 Ipp2349 2690279 2690590 P unknown 594.1 Chemiosmotic efflux 6309 Ipp2350 2690729 2691181 m cecA system C protein A 1.2 Chemiosmotic efflux 157.2 3849 Ipp2351 2691178 2694393 m cecA system protein A-like protein 2 Chemiosmotic efflux 154.1 3830 Ipp2352 2694394 2695362 m + cecB system C protein B 2 Chemiosmotic efflux 153.2 3823 Ipp2353 2695359 2696672 m + cecC system C protein C 2 Similar to outer membrane 152.3 3818 Ipp2354 2696697 2697056 m + unknown lipoprotein 1 1961.3 4097 Ipp2355 2697208 2698089 m Unknown regulatory protein (GGDEF domain) 3 234.2 4286 Ipp2356 2698228 2700438 m copAl unknown similar to copper efflux ATPase 2 228.1 4253 Ipp2357 2700502 2700669 m unknown 6 Similar to ribose-phosphate 227.2 4247 Ipp2358 2700694 2701605 m unknown pyrophosphokinase 2.3 similar to thymidine/pyrimidine- 111.2 3580 Ipp2359 2701602 2703155 m unknown nucleoside phosphorylase Similar to metallo-beta-lactamase 109.1 3564 Ipp2360 2703113 2704525 m unknown superfamily proteins 107.1 Chemiosmotic efflux 3553 Ipp2361 2704471 2707614 m cebA system B protein A Chemiosmotic efflux 104.1 3533 Ipp2362 2707689 2708822 m cebB system B protein B Chemiosmotic efflux 103.1 3525 Ipp2363 2708819 2710078 m cebC system B protein C 102.1 3519 Ipp2364 2710065 2712755 m ctpA unknown similar to cation transport ATPase
Table XIV 99.1 6582 Ipp2365 2712946 2713077 p unknown hypothetical gene 6 97.2 6568 Ipp2366 2713176 2713448 p unknown hypothetical gene 6 96.6 6562 Ipp2367 2713576 2714433 p unknown 5.1 743.4 6432 Ipp2368 2715263 2715658 p unknown 5.2 similar to cadmium-transporting 744.4 6433 Ipp2369 2715673 2716179 m cadAl unknown ATPase (C-terminal part) 4 747.4 6434 Ipp2370 2717023 2719158 m cadA2 unknown similar to cation transport ATPase 2631.5 4473 Ipp2371 2719234 2722383 m helA HelA protein Similar to cation efflux system 2627.1 4471 Ipp2372 2722393 2723649 m + helB HelB protein protein CzcB Similar to cation efflux system 1804.2 4004 Ipp2373 2723646 2724890 m + helC HelC protein protein CzcC 1803.2 4003 Ipp2374 2725059 2725175 m unknown similar to unknown protein Similar to Legionella pneumophila 2626.1 4470 Ipp2375 2725344 2726006 m unknown putative phage repressor, PrpA 4.4 Similar to Legionella vir region 2625.2 4469 Ipp2376 2726191 2727063 unknown protein LvrA 5.1 Similar to Legionella vir region 5812.1 6278 Ipp2377 2726921 2727391 unknown protein LvrB 5.1 Similar to Legionella vir region 5813.1 6279 Ipp2378 2727411 2727608 unknown protein LvrC 5.1 5814.1 6280 Ipp2379 2727605 2728039 P + Unknown similar to other proteins 5.2 4109.2 5365 Ipp2380 2728036 2728626 P + unknown 6 4108.1 5364 Ipp2381 2728637 2729473 P + unknown Similar to hypothetical protein 5.2 1376.2 3729 Ipp2382 2729475 2729933 P - unknown Similar to hypothetical protein 5.2 1375.2 3728 Ipp2383 2729924 2730622 P + unknown similar to unknown protein 5.2 1373.1 3727 Ipp2384 2730632 2730958 P - unknown 6 1372.2 3726 Ipp2385 2730961 2731224 P - unknown 6 2367.2 4306 Ipp2386 2731227 2731613 P + unknown 6 4107.1 5363 Ipp2387 2731622 2731972 P - unknown 6 4106.1 5362 Ipp2388 2731977 2732639 P + unknown putative secreted protein 5.2 Similar to conserved hypothetical 4105.1 5361 Ipp2389 2732626 2733408 P + unknown protein, putative secreted protein 5.2 Similar to conserved hypothetical 4104.2 5360 Ipp2390 2733405 2734718 unknown protein 5.2
Table XIV Similar to conserved hypothetical 5517.1 6167 Ipp2391 2734675 2735064 p unknown protein, putative secreted protein 5.2 Similar to type IV secretory 1287.3 3682 Ipp2392 2735075 2737849 P - unknown pathway, VirB4 components 1.6 Similar to conserved hypothetical 4822.3 5848 Ipp2393 2737758 2738819 P - unknown protein 5.2 Similar to conserved hypothetical 1623.4 3881 Ipp2394 2738829 2740211 P - unknown protein 5.2 Similar to conserved hypothetical 1624.5 3882 Ipp2395 2740213 2741781 P - unknown protein 5.2 4233.4 5452 Ipp2396 2741784 2741993 m + unknown 6 4234.4 5453 Ipp2397 2742007 2742339 P + unknown hypothetical gene 6 Similar to conserved hypothetical 4236.4 5454 Ipp2398 2742343 2744349 P - unknown protein 5.2 4238.1 5455 Ipp2399 2744351 2744833 P - unknown 6 4239.4 5456 Ipp2400 2744922 2745209 m - unknown hypothetical gene 6 5576.3 6197 Ipp2401 2745277 2746245 m - Unknown 6. Similar to transposase (ISL3 6160.1 6341 Ipp2402 2746636 2747811 m - Unknown family) 4.5 5987.2 6315 Ipp2403 2748197 2749000 P - unknown 6 5626.3 6221 Ipp2404 2748993 2749895 P - unknown Weakly similar to integrase 4.5 Similar to cobalt-nickel resistance 1920.3 4074 Ipp2405 2750070 2751128 P + unknown system protein 1.2 1923.2 4075 Ipp2406 2751783 2751974 P - unknown 6 1924.4 4076 Ipp2407 2752031 2752537 P - unknown Similar to antirestriction protein 3.2 Similar to conserved hypothetical 3258.2 4836 Ipp2408 2752559 2753485 m - unknown protein 5.2 3257.2 4835 Ipp2409 2753475 2753927 m - unknown Similar to hypothetical protein 5.2 Similar to transposase (ISL3 6159.1 6339 Ipp2410 2754296 2755276 m - Unknown family) 4.5 Similar to transposase (ISL3 5890.1 6292 Ipp2411 2755537 2755635 m - Unknown family) - partial 4.5 3653.3 5083 Ipp2412 2755742 2755915 m - unknown 6 weakly similar to serine/threonine 3655.1 5084 Ipp2413 2756221 2756919 m - unknown protein kinase 3.8 510.2 6008 Ipp2414 2756912 2757601 m - unknown 6 509.1 6001 Ipp2415 2757715 2758308 m unknown similar to unknown proteins 5.2 507.3 5992 Ipp2416 2758338 2758478 m - Unknown Similar to transposase - partial 4.5
Table XIV 506.3 5983 Ipp2417 2759103 2759459 P - unknown 6 505.3 5976 Ipp2418 2759500 2760432 m - unknown 6 3657.1 5085 Ipp2419 2760752 2761390 m - Unknown 6. 3658.1 5086 Ipp2420 2761865 2762041 P - unknown hypothetical gene 6 948.3 6557 Ipp2421 2762052 2762537 P - Unknown similar to transposase 4.5 947.3 6556 Ipp2422 2762506 2762868 m unknown similar to unknown protein 5.2 946.3 6555 Ipp2423 2762906 2763514 m - Unknown Similar to unknown protein 5.2 3659.2 5087 Ipp2424 2763698 2764150 m - Unknown 6. 3661.4 5089 Ipp2425 2764204 2765190 m + Unknown 5.2 5620.3 6218 Ipp2426 2765844 2766071 P - unknown Similar to transcriptional regulator 3.5.2 2153.3 4196 Ipp2427 2766068 2766304 P - unknown hypothetical gene 6 426.3 5470 Ipp2428 2766613 2767311 P - unknown 6 6369.1 6373 Ipp2429 2767308 2767526 P - unknown similar to transposase, partial 4.5 425.1 5465 Ipp2430 2767602 2767829 m - unknown 6 Similar to conserved hypothetical 424.1 5457 Ipp2431 2767876 2768187 m - unknown protein 5.2 423.2 5449 Ipp2432 2768203 2769495 m + unknown Similar to hypothetical protein 5.2 5621.1 6219 Ipp2433 2769530 2770738 m + unknown Similar to transporters 1.2 5623.2 6220 Ipp2434 2771034 2772428 m - unknown 6 Similar to conserved hypothetical 3891.2 5222 Ipp2435 2772433 2773068 m - unknown protein 5.2 3890.2 5221 Ipp2436 2773237 2774583 P - unknown 6 3889.1 5219 Ipp2437 2774573 2775328 m - unknown Similar to hypothetical protein 5.2 3888.1 5218 Ipp2438 2775331 2775528 m - unknown hypothetical gene 6 Similar to multidrug resistance 3887.1 5217 Ipp2439 2775548 2776774 m - unknown proteins 1.2 3884.2 5216 Ipp2440 2777410 2777883 P - unknown 6 similar to transcriptional regulator, 1432.3 3767 Ipp2441 2778073 2778990 P - unknown LysR family 3.5.2 1429.4 3766 Ipp2442 2779173 2780660 m + unknown 6 5404.2 6117 Ipp2443 2781074 2781973 m - unknown 6 5402.1 6116 Ipp2444 2782135 2783574 P - unknown 6 similar to transcriptional regulator, 2533.3 4420 Ipp2445 2783820 2784728 P - unknown LysR family 3.5.2 similar to potassium uptake protein 2530.3 4419 Ipp2446 2784866 2786755 P _ kup unknown Kup 1.2
Table XIV Some similarity with eukaryotic 3602.2 5050 Ipp2447 2786973 2788736 p unknown proteins 6 3601.1 5049 Ipp2448 2788923 2790071 p unknown similar to sodium-proton antiporter 1.2 conserved hypothetical protein, 3600.2 5048 Ipp2449 2790295 2791233 p unknown some similarity with stress proteins 5.2 2453.4 4367 Ipp2450 2791251 2793971 p unknown similar to cation transport ATPase 1.2 1652.4 3903 Ipp2451 2794065 2794628 p unknown 6 similar to plasminogen activator 1653.3 3904 Ipp2452 2794743 2795669 m unknown precursor from Yersinia 2.2 Similar to amino acid transport 606.3 6327 Ipp2453 2795797 2797404 m unknown protein 1.2 612.2 6336 Ipp2454 2797735 2799900 katB catalase-peroxidase KatB 4.2 2079.2 4153 Ipp2455 2800211 2801488 m unknown 6 2078.2 4152 Ipp2456 2801622 2801834 m unknown hypothetical gene 6 Similar to major facilitator family 3673.2 5098 Ipp2457 2802048 2803280 m unknown transporter 1.2 SdbC proteins, putative 4917.2 5899 Ipp2458 2803321 2804625 m sdbC substrate of the Dot/Icm system 5.1 Some similarity with eukaryotic 4916.2 5898 Ipp2459 2804894 2805778 P unknown proteins 5.2 5655.2 6237 Ipp2460 2805899 2806363 m unknown Similar to bacterioferritin 4.2 5654.2 6236 Ipp2461 2806763 2807122 P unknown 6 Similar to conserved hypothetical 4935.2 5910 Ipp2462 2807250 2808320 m unknown protein 5.2 4934.3 5909 Ipp2463 2808690 2809889 m unknown similar to hypothetical peptidases 2.2 Similar to aminoglycoside 6 -N- 5808.1 6277 Ipp2464 2809944 2810516 m unknown acetyltransferase 4.2 C-terminal part similar to 964.3 6563 Ipp2465 2810706 2812052 m unknown Legionella unknown virulence protein 5.1 953.2 6561 Ipp2466 2812652 2814166 p unknown 6 Similar to transcriptional regulator, 952.1 6560 Ipp2467 2814143 2815063 m Unknown LysR family 3.5.2
5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 5 o o o o o o o o o o o o o o o o o o o o o o o o o o o cz cz sz cz sz cz cz cz cz cz c cz cz c cz cz cz cz c c c c c c c c c c c c c c c c c c c c c c c c c c c c c 3 3 3 3 3 3 3 3 3 3 3 3 3 3 3 3 3 3 3 3 3 3 3 3 3 3 3
LU XI
ε α. Q. E D* E E n o* ε ε CL ε CL CL ε CL rH ro rH cn cn vo ro Is- ΓM 00 o oo ro OO rH rv in rH 00 VD in Is- VD 00 o vo rH VD rv rv o ro 00 rH ro 00 IN o rr Is- CO Is- rv cn 00 00 LO r- o cn 00 rH IN ro rv cn 00 VD o cn in rr in in r Is- VD IN cn rr o 00 rH VD VD rH cn o CO 00 O VD Is- 00 cn rH rH ΓM ro rr in VO 00 cn rH H ΓM oo VD Γ- cn rH ro 00 in VD 00 o rH rH rH rH ΓM (N ΓM IN ΓM ΓM (N ΓM IN 00 00 00 00 00 00 ro rT rr rr rr rT rr in 00 CO 00 CO 00 00 00 00 00 oo CO 00 00 CO 00 00 CO CO 00 00 00 00 CO 00 co CO 00 IN ΓM ΓM ΓM ΓM ΓM ΓM IN ΓM ΓM (N ΓM IN ΓM ΓM (N ΓM ΓM IN ΓM ΓM ΓM ΓM ΓM CM ΓM ΓM Is- cn o in ΓM vD o 00 rr ΓM rH 00 VD rT rr rr rr rv in in m rT ΓM 00 (N o in rT in Is- in rr (N in Is- o o o in r o 00 00 rH rH rr cn rT vD rr CO rH CO ro IN o cn rr cn rH ΓM Is- rv r- LO rv CO ΓM m H 00 00 in 00 H cn Is- rl- ΓM in rH 00 in rv Is- 00 cn r ΓM (N 00 rr in vo CO o rH (N ΓM OO vo rv cn rH ro rr m Is- 00 rH H rH rH rH IN ΓM ΓM ΓM (N ΓM ΓM ΓM ro 00 oo 00 OO ro ro 00 rT rr r rr rT rr 00 00 00 00 00 00 00 CO 00 00 00 00 00 00 00 00 00 00 CO 00 00 CO 00 00 00 CO 00 (N ΓM ΓM ΓM ΓM (N ΓM ΓM N (N ΓM ΓM ΓM ΓM ΓM ΓM (N ΓM ΓM ΓM (N ΓM ΓM (N ΓM ΓM IN CO cn o rH ΓM ro rT in vo rv CO cn o H (N ro rr in VD rv 00 cn o rH ΓM ro rr VD VD Is- rv r- Is- Is- rv rv rv rv Is- 00 00 CO 00 00 00 00 00 00 CO n cn cn cn cn rT rT r rr rr rr rT rr rr rr rr rr rr rr rr rr rr rr rr rr rr rr rr T rT rr rr ΓM ΓM (M (N (N ΓM ΓM ΓM ΓM ΓM (N (N ΓM (N IN ΓM ΓM ΓM ΓM ΓM ΓM ΓM ΓM (N ΓM ΓM ΓM CL O- CL Q. CL CL Q. o. Q. CL CL CL CL Q. CL L Q. Q. CL Q. o. Q. CL CL CL CL CL Q. CL Q. CL α. CL CL Q. Q. CL CL Q. CL C Q. Q. CL α. CL α. Q. Q. CL Q. CL Q. CL cn rr 00 ιn VD r (N o ro ΓM cn 00 VD m ΓM 00 rr vo Is- 00 00 Is- VD ro 00 rH rr in ro 00 (N ΓM ΓM ro 00 rT rr Γ ΓM ro ro CO 00 00 00 00 00 rr rr rr rH Is- rT rv in CO 00 in in in CO CO co 00 CO 00 O o CO 00 00 00 00 00 rr 00 co cn o VD m vo rr rT VD vD o rr rr 00 ro r rr rr rr rr rr rr rr rr rr rr 00 00 ro vo m D
> 1—1
X rH rH (N (M (N IN rH ΓM oo IN ΓM ro oo IN CM H ΓM rH ΓM ΓM ΓM ro ΓM ΓM ro ro o -i to CM r rH ro in o cn ΓM o 00 VD rH o VD rv 00 H ro LO in r vD rr Is- cn
2 in in in o o o in rr VD VD rr rr VD vo 00 ΓO 00 rT rT rr 00 VD vD vo cn rH oi cn (N (N cn cn n ΓM (N in in (N ΓM 00 00 00 ro 00 00 ro ro in LO in Is- rH rH vo ΓM LO oo ro ro 00 00 ro rH rH 00 ro 00 00 ro 00 ΓM rH rH rH in rr cn
H
Table XIV Similar to malonate decarboxylase, 980.1 6575 Ipp2495 2850032 2850751 p unknown gamma subunit 2.1.1 Similar to phosphoribosyl- 981.2 6576 Ipp2496 2850748 2851371 p unknown dephospho-CoA transferase 3.8 Similar to 2-(5 -triphosphoribosyl)- 4517.2 5640 Ipp2497 2851343 2852185 p unknown 3 -dephosphocoenzyme-A synthase 2.1.1 Similar to malonyl-CoA acyl-carrier- 4819.2 5845 Ipp2498 2852182 2853129 P - unknown protein transacylase 2.1.1 4821.2 5847 Ipp2499 2853263 2854156 P - unknown Similar to phosphatase 2.6 4850.2 5860 Ipp2500 2854334 2856118 P - unknown 6 Similar to conserved hypothetical 4851.2 5861 Ipp2501 2856394 2856879 P - unknown protein 5.2 2347.2 4291 Ipp2502 2856906 2857949 P + unknown 6 2346.1 4290 Ipp2503 2857986 2858366 P + unknown putative membrane protein 6 2345.2 4289 Ipp2504 2858473 2858901 m + unknown 6 Similar to florfenicol efflux pump1336.3 3709 Ipp2505 2859132 2860247 m - sflR unknown like protein 1.2 similar to conserved hypothetical 2321.3 4275 Ipp2506 2860589 2861140 m - unknown protein 5.2 Similar to conserved hypothetical 5412.1 6120 Ipp2507 2861289 2862248 P - unknown protein 5.2 4591.2 5689 Ipp2508 2862286 2862759 P - unknown similar to hypothetical proteins 5.2 4590.1 5688 Ipp2509 2862859 2863269 P - unknown 5.2 4589.1 5686 Ipp2510 2863393 2863932 m - unknown 6 4588.2 5685 Ipp2511 2864177 2864728 m - unknown 6 4587.2 5684 Ipp2512 2864938 2865423 P + unknown 6 5413.1 6121 Ipp2513 2865508 2865786 P - unknown 6 Similar to D-alanyl-D-alanine 2419.3 4341 Ipp2514 2865969 2866649 P - unknown dipeptidase 1.1 predicted integral membrane 2418.2 4340 Ipp2515 2866732 2867460 unknown protein 5.2 Similar to N-hydroxyarylamine O- 2415.2 4339 Ipp2516 2867540 2868508 m unknown acetyltransferase 2.1.1 4170.1 5413 Ipp2517 2868667 2871432 m unknown Ankyrin repeat protein 6 4168.2 5411 Ipp2518 2871656 2872150 P unknown 6 4780.2 5818 Ipp2519 2872198 2872647 m unknown 6
Table XIV 4779.1 5817 Ipp2520 2872970 2873416 p unknown Similar to unknown protein 5.2 4778.1 5816 Ipp2521 2873577 2874206 p unknown 6 4777.2 5815 Ipp2522 2874405 2875712 p unknown ankyrin repeat protein 6 Similar to two-component response 5417.1 6122 Ipp2523 2875837 2876244 m unknown regulator 3.5.2 Similar to two-component sensor 4357.3 5537 Ipp2524 2876308 2877585 m shkA unknown histidine kinase 1.3 Similar to guanylate cyclase- 4358.2 5538 Ipp2525 2877578 2878132 m unknown related protein 2.3 5586.1 6201 Ipp2526 2878357 2878740 m unknown 6 2485.3 4390 Ipp2527 2878902 2879519 P unknown 6 1878.3 4046 Ipp2528 2879724 2880656 P unknown Similar to hypothetical protein 5.2 Similar to major facilitator 1875.2 4045 Ipp2529 2880843 2882006 P unknown superfamily transporters 1.2 Similar to aspartyl/asparaginyl 4363.2 5539 Ipp2530 2882145 2882864 P unknown beta-hydroxylase 3.8 Similar to hydrogenase 1 5076.2 5995 Ipp2531 2882924 2883403 m unknown maturation protease 3.8 Similar to coenzyme F420-reducing 5077.3 5996 Ipp2532 2883400 2884692 m unknown hydrogenase, alpha subunit 1.4 Similar to coenzyme F420-reducing 3652.2 5082 Ipp2533 2884685 2885470 m unknown hydrogenase, gamma subunit 1.4 Similar to 2-polyprenylphenol 3651.1 5081 Ipp2534 2885488 2886333 m unknown hydroxylase and related flavodoxin oxidoreductases 1.4 3650.1 5080 Ipp2535 2886326 2887465 m unknown Similar to hydrogenase subunit 1.4 Hydrogenase 2506.2 4403 Ipp2536 2887501 2888559 m hypE expression/formation protein HypE 1.4 Hydrogenase 2508.3 4404 Ipp2537 2888556 2889662 m hypD expression/formation protein HypD 1.4 Hydrogenase 3647.2 5078 Ipp2538 2889659 2889886 m hypC expression/formation protein HypC 1.4 Hydrogenase maturation 1320.3 3698 Ipp2539 2889896 2892121 m hypF protein HypF 1.4
Table XIV hydrogenase nickel 3645.3 5077 Ipp2540 2892135 2892893 m hypB incorporation protein HypB 1.4 Hydrogenase nickel 5807.1 6276 Ipp2541 2892893 2893234 m hypA incorporation protein HypA 1.4 weakly similar to high-affinity 4704.3 ' 5760 Ipp2542 2893246 2893962 m unknown nickel-transport protein NixA 1.2 4702.2 5759 Ipp2543 2894069 2895028 p unknown similar to glycosyl transferase 1.1 Similar to conserved hypothetical 4701.2 5758 Ipp2544 2895025 2895855 m unknown protein 5.2 Integral membrane protein, similar 4752.2 5796 Ipp2545 2896022 2896912 m unknown to metabolite efflux pump 1.2 SdbB protein (putative 4754.1 5797 Ipp2546 2897193 2898539 p sdbB substrate of the Dot/Icm system) 5.1 4755.2 5798 Ipp2547 2898566 2899123 m unknown Similar to hypothetical protein 5.2 Similar to conserved hypothetical 4066.2 5335 Ipp2548 2899172 2899756 p unknown protein 5.2 Protein with TPR motifs (protein- 4065.3 5334 Ipp2549 2899758 2901491 p unknown protein interaction motif) 5.2 4063.1 5333 Ipp2550 2901597 2902985 m algC phosphomannomutase 2.1.1 Deoxyuridine 5 - triphosphate 4061.1 5332 Ipp2551 2902986 2903444 m dut nucleotidohydrolase (dUTPase) (dUTP pyrophosphatase) 2.3 similar to DNA/pantothenate 4060.1 5331 Ipp2552 2903471 2904667 m dfp Unknown metabolism flavoprotein 2.5 1661.3 3911 Ipp2553 2904744 2905427 P radC DNA repair protein RadC 3.2 1660.2 3910 Ipp2554 2905463 2905699 P unknown hypothetical gene 6 effector protein B, 1659.4 3908 Ipp2555 2905779 2909663 lepB substrate of the Dot/Icm secretion system 5.1 Similar to conserved hypothetical 2242.1 4236 Ipp2556 2909787 2910350 m unknown protein 5.2
Table XIV similar to hypothetical Inosine-5 - 1394.2 3741 Ipp2557 2910430 2910879 m unknown monophosphate dehydrogenase 2.3 1396.3 3742 Ipp2558 2911010 2911996 P unknown Similar to alcohol dehydrogenase 2.1.1 1397.2 3743 Ipp2559 2912258 2912752 P unknown Similar to small heat shock protein 4.1 3811.1 5179 Ipp2560 2912801 2913304 m unknown Predicted membrane protein 6 Similar to conserved hypothetical 3810.1 5178 Ipp2561 2913574 2914266 P unknown protein 5.2 Similar to homospermidine 1807.3 4007 Ipp2562 2914317 2915735 m unknown synthase 2.2 1806.2 4006 Ipp2563 2916013 2916462 P unknown Similar to rRNA methylase 3.6 Similar to thiamin 1805.3 4005 Ipp2564 2916620 2917273 m unknown pyrophosphokinase 2.5 Similar to conserved hypothetical 4551.2 5663 Ipp2565 2917254 2917520 m unknown protein 5.2 4552.4 5664 Ipp2566 2917788 2918696 P unknown 6 5354.3 6096 Ipp2567 2919033 2919257 P unknown 6 Similar to conserved hypothetical 1467.5 3785 Ipp2568 2919387 2919959 P unknown protein 5.2 1466.2 3784 Ipp2569 2919999 2920739 P unknown similar to carbonic anhydrase 4.7 similar to multidrug resistance 4645.2 5721 Ipp2570 2920824 2922056 P unknown protein, MFS superfamily 1.2 4644.3 5720 Ipp2571 2922053 2923318 m unknown Similar to hypothetical protein 5.2 4206.2 5436 Ipp2572 2923626 2926454 P unknown 6 4205.2 5435 - Ipp2573 2926474 2927346 P unknown 6 similar to sensor histidine 4203.2 5434 Ipp2574 2927752 2929014 P unknown kinase/response regulator 1.3 4200.2 5433 Ipp2575 2929062 2929757 m unknown 6 1377.5 3730 Ipp2576 2930154 2932577 m unknown 6 SdeD protein (substrate of 1378.5 3731 Ipp2577 2932649 2933797 m sdeD the Dot/Icm system) 5.1 SdeA protein, paralog of 5349.3 6095 ' Ipp2578 2934089 2936779 p sdeA SidC (substrate of the Dot/Icm system) 5.1 SidC protein (substrate of 4431.2 5586 Ipp2579 2936942 2939656 p sidC the Dot/Icm system) 5.1 5129.4 6016 Ipp2580 2939975 2943127 m ImxF unknown Similar to multidrug efflux pump 1.2
Table XIV 4873.3 5873 Ipp2581 2943187 2944341 m - ImxE unknown Similar to efflux protein 1.2 Similar to outer membrane efflux 4872.3 5872 Ipp2582 2944347 2945909 m + IprN unknown protein 1.2 6086.1 6331 Ipp2583 2946091 2946405 m - unknown 5.1 Major facilitator superfamily (MFS) 5685.2 6246 Ipp2584 2946460 2947692 m - smlA unknown transporter 5.1 Similar to transcriptional regulator 5686.2 6247 Ipp2585 2947892 2948365 P - unknown (Lrp family) 3.5.2 5687.1 6248 Ipp2586 2948562 2948903 P - unknown 6 5688.2 6249 Ipp2587 2949093 2949653 P - unknown 6 5569.1 6195 Ipp2588 2949746 2950114 m + Unknown 6 5566.1 6194 Ipp2589 2950336 2950515 P + Unknown 6 1065.3 3549 Ipp2590 2950556 2951686 m - unknown similar to other protein 5.2 1063.1 3548 Ipp2591 2952111 2953355 m - unknown 6 4998.2 5943 Ipp2592 2953818 2955437 m - unknown 6 4995.2 5942 Ipp2593 2955658 2957208 P - unknown weakly similar to alpha-glucosidase 2.1.1 3316.1 4870 Ipp2594 2957609 2959303 m - unknown 6 phospho-2-dehydro-3- 2057.3 4140 Ipp2595 2960106 2961149 p aroF deoxyheptonate aldolase 2.2 similar to chorismate mutase (N- 977.3 6573 Ipp2596 2961146 2961730 p unknown terminal part) 2.2 similar to chorismate mutase (C- 976.3 6572 Ipp2597 2961764 2962024 p unknown terminal part) 2.2 Similar to aspartate 974.2 6571 Ipp2598 2962024 2963190 p aspB unknown aminotransferase 2.2 Similar to tellurite resistance 2056.1 4139 Ipp2599 2963368 2964264 p unknown protein TehB 4.2 1270.3 3669 Ipp2600 2964297 2964728 m unknown similar to unknown protein 5.2 similar to hemoglobin 1271.4 3670 Ipp2601 2964808 2965182 p unknown (protozoan/cyanobacterial globin family) 1.4 1272.5 3671 Ipp2602 2965179 2966135 p unknown similar to xylene monooxygenase 2.1.1 Similar to conserved hypothetical 2054.2 4138 Ipp2603 2966360 2967379 p unknown protein 5.2 3319.1 4871 Ipp2604 2967503 2968849 m unknown 6 4962.2 5925 Ipp2605 2969502 2969909 m unknown 6
Table XIV Similar to large conductance 4961.1 5924 Ipp2606 2970108 2970491 m unknown mechanosensitive channel protein 1.2 4960.2 5923 Ipp2607 2970606 2971433 m unknown 6 4943.2 5914 Ipp2608 2971714 2972139 m unknown 6 4942.1 5913 Ipp2609 2972273 2972482 m unknown 6 4941.3 5912 Ipp2610 2972499 2972939 m unknown 6 similar to membrane-bound lytic 748.4 6435 Ipp2611 2973586 2974959 P unknown murein transglycosylase A 1.1 749.1 6436 Ipp2612 2975335 2975625 P unknown 6 750.2 6437 Ipp2613 2975856 2976002 m unknown hypothetical gene 6 752.2 6438 Ipp2614 2976101 2976256 m unknown hypothetical gene 6 2143.5 4186 Ipp2615 2976501 2977898 m unknown 6 similar to protein secretion 2302.2 4265 Ipp2616 2978085 2978420 m unknown chaperonin CsaA 1.6 1182.3 3620 Ipp2617 2978430 2979344 m unknown Predicted transmembrane protein 5.2 Similar to trancriptional regulator, 1180.2 3619 Ipp2618 2979410 2980165 m unknown AraC family 3.5.2 1179.1 3617 Ipp2619 2980423 2981019 m + unknown 6 2712.1 4515 Ipp2620 2981450 2982223 m unknown 6 Similar to permeases of the 2713.2 4516 Ipp2621 2982430 2983425 m unknown drug/metabolite transporter (DMT) superfamily 1.2 2717.1 4517 Ipp2622 2983872 2985539 m unknown putative membrane protein 6 Similar to hexose phosphate 2718.1 4518 Ipp2623 2985717 2987084 P unknown transport protein 1.2 Similar to N-terminal part of rare 2477.1 4384 Ipp2624 2987347 2987829 P unknown lipoprotein A 1.2 2479.2 4385 Ipp2625 2988299 2989141 m unknown 6 1324.4 similar to eukaryotic serine 3701 Ipp2626 2989724 2991112 P Unknown threonin protein kinase 3.8 Similar to Legionella putative 2073.4 4150 Ipp2627 2991137 2991820 m unknown transcriptionnal regulator 3.5.2 Similar to conserved hypothetical 1386.3 3734 Ipp2628 2992329 2992706 m unknown protein 5.2 1385.2 3733 Ipp2629 2993009 2993734 m unknown 6 2075.1 4151 Ipp2630 2993857 2994111 m unknown 6
E υ o c Q c c c c c c c c c c c c c 5 5 5 5 5 5 Φ E 5 5 5 5 5 5 5 5 5 5 5 5 o o o o o o JZ Φ _ c- c _ c_. c c c c c c c c c c c c c ,-i - c^ ->. Ul c c c c c O c >- c c c c c c c c c c c c 3 3 Φ Ul ro
m ro o.
+ ε ε ε ε E
rr ro co vo in o rv O CO rr o o rH Pv in VD cn ro oo VD CO rr o o oo ro LO LO 00 cn o rv rr o o n oo rr ΓM in r o rT rT o oo cn o o oo rr LO I
s- Cn O rH ΓM ro rr in VD cn cn cn cn cn o o o o o o O rH rH TH cn cn cn cn cn o o o o o o o o o o O o ΓM IN (N ΓM ΓM 00 ro ro oo oo oo o o 00 oo ro 00 oo IN ro rr LO VD rv co cn o (N ro rT in VD rv co cn 00 00 ro 00 oo ro ro oo 00 T rT rT rr rT rr rr rr rT rr VD o vo VD vo VD vD VO vo VD VD vo VD VO VD VD vo VD VD IN ΓM ΓM ΓM IN ΓM ΓM ΓM (N N ΓM (N ΓM (N (N ΓM IN ΓM ΓM Q. Q. CL CL CL Q. CL Q. CL Q. Q. Q. CL Q. o. Q. Q. Q. CL C CL Q. Q. CL Q. Q. O. CL CL Q.

Table XIV 3446.1 4945 Ipp2650 3017091 3018176 p unknown similar to E. coli Smf protein 5.2 3447.1 4946 Ipp2651 3018166 3018603 p unknown similar to unknown protein 5.2 5200.2 6037 Ipp2652 3018683 3020962 p topA DNA topoisomerase I 3.4 3271.3 4844 similar to membrane bound Ipp2653 3021173 3023146 m unknown acyltransferase 2.1.1 Similar to conserved hypothetical 3272.1 4845 Ipp2654 3023282 3023722 m unknown protein 5.2 3273.1 4846 Ipp2655 3023853 3024275 P unknown Similar to hypothetical protein 5.2 3275.1 4847 Ipp2656 3024356 3025597 P unknown 6 3277.2 4848 Ipp2657 3025898 3026695 P unknown Similar to unknown protein 5.2 3278.1 4849 Ipp2658 3026750 3027166 m unknown 6 3280.1 4851 Ipp2659 3027281 3028132 m unknown Similar to unknown protein 5.2 637.3 6374 Ipp2660 3028286 3030322 m unknown Similar to peptidase 2.2 UDP-3-0-[3- hydroxymyristoyl] N- 636.2 6372 Ipp2661 3030466 3031380 m IpxC acetylglucosamine deacetylase 2.4 635.5 6371 Ipp2662 3031628 3032824 m ftsZ Cell division protein FtsZ 1.7 ATP-binding cell division 3282.3 4852 Ipp2663 3033019 3034257 m ftsA protein FtsA 1.7 3283.1 4853 Ipp2664 3034257 3034976 m ftsQ Cell division protein FtsQ .7 D-alanine~D-alanine ligase 1463.2 3783 Ipp2665 3034992 3036086 m ddlA A .1 UDP-N- 1461.3 3782 Ipp2666 3036070 3036981 m murB acetylenolpyruvoylglucosa mine reductase .1 UDP-N-acetylmuramate— L- 2358.3 4300 Ipp2667 3037005 3038414 m murC alanine ligase .1 1701.6 3935 Ipp2668 3038424 3039599 m + ftsW Cell division protein ftsW .7 UDP-N- 1699.7 3934 Ipp2669 3039605 3040948 m murD acetylmuramoylalanine— D- glutamate ligase .1 Phospho-N-acetylmuramoyl 3694.3 5112 Ipp2670 3040962 3042047 m mraY pentapeptide-transferase
Table XP V UDP-N-acetylmuramoyl- 683.2 6402 Ipp2671 3042162 3043499 m + murF tripeptide—D-alanyl-D- alanine ligase 1.1 684.5 6403 Ipp2672 3043740 3044519 m unknown Similar to cell division protein 1.7 similar to chromosome partition 3697.5 5113 Ipp2673 3044524 3048018 m - smc unknown protein smc 3.4 Similar to acid phosphatase, class 3150.1 4770 Ipp2674 3048193 3048873 P + unknown B 2.6 3149.1 4768 Ipp2675 3048975 3050036 m + unknown Weakly similar to cysteine protease 2.2 3148.1 4767 Ipp2676 3050346 3051158 P + unknown Similar to hypothtical protein 5.2 transcription elongation 3147.1 4766 Ipp2677 3051232 3051714 m - greA factor GreA 3.5.3 carbamoyl-phosphate 727.3 6423 Ipp2678 3051723 3054926 m + carB synthase large subunit 2.3 2098.2 4161 Ipp2679 3055055 3055327 m unknown 6 2095.2 Similar to conserved hypothetical 4160 Ipp2680 3055440 3056621 P - unknown protein 5.2 3146.2 4765 Ipp2681 3056661 3057413 m unknown Similar to hypothetical protein 5.2 880.3 6508 Ipp2682 3057413 3058483 m - unknown Putative membrane protein 5.2 Predicted membrane protein, 881.2 6509 Ipp2683 3058480 3059559 m + unknown similar to putative permease 5.2 882.3 6510 Ipp2684 3059763 3061214 P - pepA unknown Similar to leucine aminopeptidase 3.8 Similar to DNA polymerase III, chi 2221.3 4225 Ipp2685 3061195 3061629 P - holC unknown subunit 3.1 4373.3 5547 Ipp2686 3061776 3062093 m unknown Similar to hypothetical protein 5.2 2542.2 4424 Ipp2687 3062116 3063480 m _ unknown Similar to aminopeptidase 2.2 similar to integral membrane 4374.4 5548 Ipp2688 3063485 3065056 m - unknown protein MviN 5.2 30S ribosomal subunit 5806.2 6275 Ipp2689 3065414 3065680 P rpsT protein S20 3.7.1 4818.3 5844 Ipp2690 3065741 3066952 m unknown 6 4817.4 5843 Ipp2691 3067304 3068686 m unknown 6 enhanced entry protein 4167.4 5410 Ipp2692 3069093 3072695 m + enhC EnhC 4.6 enhanced entry protein 4165.1 5409 Ipp2693 3072722 3073288 m + enhB EnhB 4.6
Table XIV enhanced entry protein 4164.1 5408 Ipp2694 3073285 3074007 m + enhA EnhA 4.6 regulatory protein (GGDEF and EAL 4162.2 5407 Ipp2695 3074180 3076687 P - Unknown domains) 1.3 628.3 6361 Ipp2696 3076956 3077615 P + unknown 6 similar to putative protein from 627.1 6358 Ipp2697 3077783 3078931 P - Unknown Stx2 converting bacteriophage I 4.5 similar to excinuclease ABC subunit 2217.3 4222 Ipp2698 3078955 3080811 m uvrC unknown C 3.2 Legionella transmission 1511.3 3812 Ipp2699 3080823 3081482 m gacA/letA activator LetA 3.5.2 similar to phenylalanine-4- 1510.3 3811 Ipp2700 3081607 3082425 m phhA unknown hydroxylase 2.2 1508.4 3809 Ipp2701 3082865 3083962 P - unknown belong to the CinA protein family 5.2 Similar to Legionella essential 3385.1 4903 Ipp2702 3083921 3084946 m obgL unknown GTPase 2.3 3386.1 4904 Ipp2703 3085160 3085438 m rpmA 50S ribosomal protein L27 3.7.1 1894.2 4058 Ipp2704 3085451 3085762 m rplU 50S ribosomal protein L21 3.7.1 similar to 50S ribosomal subunit 1893.2 4057 Ipp2705 3086005 3086664 P - unknown protein L25, RplY 3.7.1 2253.2 4240 Ipp2706 3086781 3087350 P pth unknown similar to peptidyl-tRNA hydrolase 3.7.5 2252.1 4239 Ipp2707 3087369 3088460 P - unknown Similar to GTP-binding protein 4.6 3388.1 4905 Ipp2708 3088822 3089937 P - Unknown regulatory protein (GGDEF domain) 1.3 similar to octaprenyl-diphosphate 3389.1 4906 Ipp2709 3090071 3091039 P ispB unknown synthase 2.5 3390.1 4908 Ipp2710 3091066 3091281 m - unknown hypothetical gene 6 3391.2 4909 Ipp2711 3091262 3093517 m feoB ferrous iron transporter B 1.2 5180.1 6029 Ipp2712 3093514 3093741 m feoA ferrous iron transporter A 1.2 3544.2 5010 Ipp2713 3093830 3094912 m - unknown similar to predicted ATPase 5.2 3543.1 5009 Ipp2714 3094909 3095571 m - unknown similar to unknown protein 5.2
Table XIV similar to 3-methyl-2- 3542.1 5008 Ipp2715 3095808 3096596 p panB unknown oxobutanoatehydroxymethyltransfe rase 2.5 similar to pantothenate 3541.1 5007 Ipp2716 3096608 3097366 p panC unknown synthetases 2.5 Similar to conserved hypothetical 307.2 4722 Ipp2717 3097363 3097896 p unknown proteins 5.2 Similar to biotin carboxylase (A 306.1 4715 Ipp2718 3097910 3099325 m unknown subunit of acetyl-CoAcarboxylase) 2.4 Similar to dienelactone hydrolase 305.2 4709 Ipp2719 3099318 3100034 m unknown family protein 2.1.1 1724.4 3950 Ipp2720 3100316 3101182 p unknown similar to unknown protein 5.2 RNA polymerase sigma-32 1725.4 3951 Ipp2721 3101317 3102171 m rpoH factor 3.7.3 Similar to cell division ABC 4294.3 5490 Ipp2722 3102467 3103396 m ftsX unknown transporter, permease protein FtsX 1.7 Highly similar to cell division ABC 2337.3 4284 Ipp2723 3103390 3104040 m ftsE unknown transporter, ATP-binding protein FtsE 1.7 Similar to C-terminal part of signal 2339.4 4285 Ipp2724 3104027 3105094 m unknown recognition particle GTPase, FtsY 1.7 4295.3 5491 Ipp2725 3105157 3106482 p unknown Similar to zinc protease 2.2 4296.1 5492 Ipp2726 3106479 3107783 p unknown Similar to zinc protease 2.2 Similar to methyltransferase 4297.2 5493 Ipp2727 3107780 3108325 p unknown proteins 3.2 5178.1 6028 Ipp2728 3108400 3108891 p dotD lipoprotein DotD 1.6 defect in organelle 2362.3 4303 Ipp2729 3108872 3109783 p dotC trafficking lipoprotein DotC 1.6 defect in organelle 2364.1 4304 Ipp2730 3109783 3110916 p dotB trafficking protein DotB (ATPase) 1.6 Similar to 5 -nucleotidase, catalytic 2365.2 4305 Ipp2731 3110998 3112725 p unknown domain 2.3 1720.3 3947 Ipp2732 3112767 3113564 m unknown similar to unknown protein 5.2 3538.1 5004 Ipp2733 3113594 3114538 m unknown similar to oxidoreductase 2.1.1
Table XIV similar to D-alanine-D-alanine 1277.2 3674 Ipp2734 3114775 3115797 unknown ligase (N-terminal part) 2.2 1275.2 3673 Ipp2735 3116071 3116958 P unknown similar to aldolase 2.1.1 1273.4 3672 Ipp2736 3117097 3117804 P unknown Similar to hypothetical protein 5.2 DlpA protein (isocitrate and isopropylmalate

3540.2 5006 Ipp2737 3117791 3119638 dlpA dehydrogenases family protein) 2.1.1 3582.2 5033 Ipp2738 3119638 3120498 p unknown similar to hypothetical protein 5.2 3581.1 5032 Ipp2739 3120759 3121475 m sbpA unknown small basic protein SbpA 5.2 defect in organelle 1770.2 3979 Ipp2740 3121606 3124713 m dotA trafficking protein DotA 1.6 intracellular multiplication 1772.1 3980 Ipp2741 3124710 3125165 m icmV protein IcmV 1.6 Intracellular multiplication 1773.1 3981 Ipp2742 3125278 3125733 icmW protein IcmW 1.6 Intracellular multiplication 1774.2 3982 Ipp2743 3125733 3127151 P icmX protein IcmX 1.6 3578.3 5031 Ipp2744 3127158 3128798 m IphB unknown, LphB 5.2 3846.2 5200 Ipp2745 3129165 3131708 P unknown similar to cation transport ATPase 1.2 273.1 4525 Ipp2746 3131959 3132489 P unknown 6 m Similar to SAM dependent 271.1 4514 Ipp2747 3132561 3133361 Unknown methyltransferase 4.6 Similar to eukaryotic phytanoyl- 270.1 4507 Ipp2748 3133485 3134315 m unknown CoA dioxygenase 2.4 269.1 4501 Ipp2749 3134430 3135479 m unknown similar to glycosyltransferase 1.1 268.1 4495 Ipp2750 3135460 3135654 m unknown similar to unknown proteins 5.2 similar to tRNA delta(2)- 267.3 4490 Ipp2751 3135677 3136618 m miaA unknown isopentenyl pyrophosphate transferase 3.6 DNA mismatch repair 5232.2 6051 Ipp2752 3136611 3138341 m mutL protein MutL 3.2 similar to N-acetylmuramoyl-L- 123.2 3647 Ipp2753 3138332 3139762 m unknown alanine amidase 1.1
Table XIV similar to conserved hypothetical 124.1 3652 Ipp2754 3139759 3140241 m unknown protein 5.2 similar to conserved hypothetical 125.2 3657 Ipp2755 3140367 3141848 m unknown protein 5.2 similar to stringent starvation 2021.1 4122 Ipp2756 3141976 3142371 m sspB unknown protein B 4.1 similar to stringent starvation 2020.1 4121 Ipp2757 3142374 3142994 m sspA unknown protein A 4.1 similar to ubiquinol-cytochrome c 2019.2 4119 Ipp2758 3143242 3143988 m petC unknown oxydo reductase, cytochrome cl 1.4 Similar to ubiquinol-cytochrome c 698.2 6410 Ipp2759 3143985 3145199 m petB unknown reductase, cytochrome b 1.4 Similar to ubiquinol-cytochrome c 697.1 6409 Ipp2760 3145210 3145827 m petA unknown reductase, iron-sulfur subunit 1.4 30S ribosomal subunit 696.2 6408 Ipp2761 3146001 3146432 m rpsl protein S9 3.7.1 50S ribosomal subunit 5229.1 6049 Ipp2762 3146438 3146872 m rplM protein L13 3.7.1 similar to putative ferredoxin 2fe- 4856.2 5863 Ipp2763 3147212 3147529 m unknown 2s protein 1.4 Similar to integration host factor, 4855.2 5862 Ipp2764 3147531 3147824 m himA unknown alpha subunit 3.4 Phenylalanyl-tRNA 2407.3 4332 Ipp2765 3147827 3150208 m pheT 3.7.2 Phenylalanyl-tRNA 4503.1 5632 Ipp2766 3150466 3151482 m pheS synthetase, alpha subunit 3.7 4505.3 5633 Ipp2767 3151619 3151978 m rplT 50S ribosomal protein L20 3.7 5227.2 6048 Ipp2768 3151994 3152194 m rpml 50S ribosomal protein L35 3.7 Translation initiation factor 5226.2 6047 Ipp2769 3152215 3152751 m infC ATT start codon IF-3 3.7 4923.2 5904 Ipp2770 3152776 3154689 m thrS Threonyl tRNA synthetase 3.7 5225.1 6046 Ipp2771 3154938 3155150 m unknown hypothetical gene 6
Table XIV Similar to conserved hypothetical 3516.2 4989 Ipp2772 3155205 3155492 P unknown protein 5.2 3515.2 4988 Ipp2773 3155827 3156312 P unknown 6 3514.1 4987 Ipp2774 3156327 3156569 P unknown hypothetical gene 6 3512.1 4986 Ipp2775 3156958 3158520 m unknown 6 similar to conserved hypothetical 3511.2 4985 Ipp2776 3158801 3159952 P unknown protein 5.2 Putative cAMP/cGMP binding 3509.2 4983 Ipp2777 3159931 3160962 P unknown protein 5.2 2590.3 4450 Ipp2778 3160969 3161886 P unknown Similar to unknown protein 5.2 Similar to NADH-dependent flavin 2591.3 4451 Ipp2779 3162014 3163069 m unknown oxidoreductase (Oye family) 1.4 Similar to transcriptional regulator, 3508.2 4982 Ipp2780 3163141 3163440 m unknown ArsR family 3.5.2 Some simillarity with eukaryotic 3507.1 4981 Ipp2781 3163693 3164037 m unknown proteins 6 3506.1 4980 Ipp2782 3164154 3164783 p unknown similar to membrane protein 5.2 Peptidyl-prolyl cis-trans 3504.1 4979 Ipp2783 3164993 3165487 m ppiB isomerase B (cyclophilin- type PPIase family) 3.8 Similar to queuine tRNA- 3503.2 4978 Ipp2784 3165514 3166665 m - tgt unknown ribosyltransferase 3.6 m Similar to disulfide bond formation 3502.3 4977 Ipp2785 3166727 3167218 + unknown protein DsbB 3.9 5804.1 6274 Ipp2786 3167215 3167643 m + unknown Similar to cytochrome c5 1.4 similar to 2-amino-3-ketobutyrate 1517.3 3816 Ipp2787 3167832 3169082 P - unknown coenzyme A ligase 2.2 1519.2 3817 Ipp2788 3169163 3170188 m - unknown Putative response regulator 3.5.2 1521.2 3819 Ipp2789 3170469 3171032 P - unknown Putative membrane protein 5.2 Similar to signal transduction 4415.3 5577 Ipp2790 3171120 3172409 m - unknown histidine kinase 1.3 . „ Porphobilinogen deaminase 4325.3 5510 Ipp2791 3172800 3173729 P - HemC 2.5 Uroporphyrinogen-III 4326.1 5511 Ipp2792 3173726 3174478 P - hemD synthetase 2.5 4327.1 5512 Similar to uroporphyrinogen III Ipp2793 3174482 3175606 P - hemX unknown methylase HemX 2.5
Table XIV protoporphyrinogen IX and 4328.1 5513 Ipp2794 3175609 3176796 + hemY coproporphyrinogen III oxidase HemY 2.5 Similar to cation-efflux system 4740.2 5788 Ipp2795 3176932 3177855 m unknown membrane protein 1.2 Similar to conserved hypothetical 4739.1 5786 Ipp2796 3177852 3178514 m unknown protein 5.2 4737.1 5785 Ipp2797 3178504 3179067 m unknown Similar to oligoribonuclease 2.3 4736.2 5784 Ipp2798 3179057 3180289 m cca tRNA nucleotidyltransferase 3.6 4734.2 5783 Ipp2799 3180352 3181329 P unknown Similar to putative GTPase 4.6 5440.1 6134 Ipp2800 3181364 3182533 m unknown 6 64.3 6376 Ipp2801 3182694 3184613 m unknown 6 DNA polymerase III, 84.4 6481 Ipp2802 3190550 3192220 dnaX gamma and tau subunits 3.1 5816.1 6281 Ipp2803 3192254 3192592 P - unknown Similar to hypothetical protein 5.2 Recombination and repair 921.3 6538 Ipp2804 3192601 3193194 P - recR protein recR 3.2 922.2 6539 Ipp2805 3193187 3194047 P + unknown Similar to hypothetical protein 5.2 923.3 6540 Ipp2806 3194044 3196041 P + unknown Similar to unknown protein 5.2 4231.2 5450 Ipp2807 3196289 3197767 m - unknown 6 4232.3 5451 Ipp2808 3197990 3198715 P - unknown Similar to response regulator 3.5.2 5062.3 5986 Ipp2809 3199108 3199749 m - unknown 6 5064.2 5987 Ipp2810 3200014 3200676 m - unknown 6 5043.3 5972 Ipp2811 3201045 3202310 m + unknown similar to unknown protein 5.2 5041.2 5971 Ipp2812 3202460 3202996 m - ppa Inorganic pyrophosphatase 2.6 Similar to HIT( Histidine triad 2520.4 4412 Ipp2813 3203126 3203467 p unknown nucleotide-binding protein) family protein 5.2 2518.4 4410 Ipp2814 3203479 3204018 p folE GTP cyclohydrolase I 2.5 Similar to L-sorbosone 2517.2 4409 Ipp2815 3204077 3205168 m unknown dehydrogenase 2.1.1 Polynucleotide 3916.4 5237 Ipp2816 3205303 3207492 m pnp phosphorylase (PNPase) 2.3
Table XIV 3917.1 5238 Ipp2817 3207607 3207882 m - rpsO 30S ribosomal protein S15 3.7.1 tRNA pseudouridine 3919.1 5239 Ipp2818 3208030 3208941 m - truB synthase B 3.6 3920.1 5240 Ipp2819 3208928 3209299 m - rbfA Ribosome-binding factor A 3.7.3 Translation initiation factor 1730.3 3953 Ipp2820 3209303 3211909 m - infB IF-2 3.7.3 Transcription elongation 3621.2 5062 Ipp2821 3211998 3213476 m - nusA protein nusA 3.5.3 Similar to conserved hypothetical 3619.1 5061 Ipp2822 3213488 3213946 m - unknown protein 5.2 NADH dehydrogenase I 1400.2 3746 Ipp2823 3214187 3215626 m - nuoN chain N .4 3747 - NADH-quinone 1402.2 ' Ipp2824 3215683 3217188 m nuoM oxidoreductase chain M 4 NADH-quinone 3618.2 5060 Ipp2825 3217204 3219177 m + nuoL oxidoreductase chain L 4 NADH-quinone 3617.2 5059 Ipp2826 3219182 3219487 m - nuoK oxidoreductase chain K 4 NADH-quinone 1006.3 3514 Ipp2827 3219505 3220164 m + nuoJ oxidoreductase chain J 4 NADH-quinone 1009.2 3515 Ipp2828 3220176 3220676 m - nuol oxidoreductase chain I 4 NADH-quinone 1010.2 3516 Ipp2829 3220695 3221717 m - nuoH oxidoreductase chain H 4 NADH dehydrogenase I 1011.5 3517 Ipp2830 3221730 3224081 m - nuoG chain G 4 NADH dehydrogenase I 291.4 4618 Ipp2831 3224098 3225375 m - nuoF chain F 4 NADH dehydrogenase I 292.1 4625 Ipp2832 3225394 3225861 m - nuoE chain E 4 NADH dehydrogenase I 293.2 4632 Ipp2833 3225958 3227211 m - nuoD chain D 4 NADH dehydrogenase I 294.1 4637 Ipp2834 3227204 3227887 m - nuoC chain C 4 NADH dehydrogenase I 295.1 4642 Ipp2835 3227910 3228386 m - nuoB chain B 4 NADH dehydrogenase I 296.1 4650 Ipp2836 3228633 3228989 m + nuoA chain A 4
Table XIV Protein-export membrane 297.2 4661 Ipp2837 3229169 3229492 m secG protein secG (Preprotein translocase subunit) 1.6 5719.2 6259 Ipp2838 3229480 3230229 m tpi triosephosphate isomerase 2.1.2 effector protein A, 5266.2 6062 Ipp2839 3230473 3233832 p lepA substrate of the Dot/Icm secretion system 5.1 633.3 6370 Ipp2840 3233902 3235269 m msrA unknown Similar to phosphoglucomutase 2.1.1 632.2 6367 Ipp2841 3235333 3236127 m folP Dihydropteroate synthase 2.5 3201.1 4801 Ipp2842 3236241 3238160 m ftsH Cell division protease ftsH 1.7 Ribosomal RNA large 666.2 6392 Ipp2843 3238274 3238894 m rrmJ subunit methyltransferase (Cell division protein FtsJ)

Similar to predicted RNA-binding 665.2 6391 Ipp2844 3238997 3239263 p unknown protein 4.6 663.2 6390 Ipp2845 3239314 3241290 m unknown Similar to O-acetyltransferase 1.1 3199.1 4800 Ipp2846 3241478 3243205 p frgA FrgA protein 5.2 Similar to 3198.1 4799 Ipp2847 3243465 3244025 p unknown phosphatidylglycerophosphate synthase 2.4 Similar to DnaA, ATPase involved in 2322.2 4276 Ipp2848 3244022 3244714 p unknown DNA replication initiation 3.1 Similar to putative 2323.2 4277 Ipp2849 3244997 3245797 P unknown coproporphyrinogen oxidase A 5.1 2324.5 4278 Ipp2850 3245682 3246995 m unknown 6 Predicted membrane protein, 3196.3 4798 Ipp2851 3247256 3248758 m unknown similar to transporters 1.2 2151.2 4194 Ipp2852 3248826 3250208 m unknown 6 Similar to conserved hypothetical 532.2 6085 Ipp2853 3250346 3251458 P unknown protein 5.2 shikimate 5- 533.2 6089 Ipp2854 3251508 3252305 m aroE dehydrogenase 2.2 536.3 6098 Ipp2855 3252529 3255126 m pepN aminopeptidase N 2.2
Table XIV Similar to conserved hypothetical 1454.2 3778 Ipp2856 3255293 3257224 P - unknown protein 5.2 Similar to conserved hypothetical 1905.2 4065 Ipp2857 3257378 3258643 P - unknown protein 5.2 1906.2 4066 Ipp2858 3258640 3260163 P - unknown similar to unknown protein 5.2 Similar to conserved hypothetical 220.1 4215 Ipp2859 3260558 3261250 m - unknown protein 5.2 Similar to dihydrodipicolinate 221.1 4220 Ipp2860 3261250 3262149 m - unknown synthase 2.2 222.1 4223 Ipp2861 3262142 3262621 m - unknown Similar to hypothetical protein 5.2 223.1 4227 Ipp2862 3262734 3263612 P - unknown Similar to transcriptional regulator 3.5.2 5822.1 6282 Ipp2863 3263771 3263935 m - unknown 6 3466.1 4957 Ipp2864 3263922 3264122 m - unknown similar to resolvase (partial) 4.5 226.4 4242 Ipp2865 3264230 3265090 P - unknown 6 5823.2 6283 Ipp2866 3265171 3266376 m + unknown Similar to aminopeptidase 2.2 2924.3 4627 Ipp2867 3266557 3268539 P + unknown Putative membrane protein 6 ATP-dependent DNA 2925.1 4628 Ipp2868 3268623 3270620 m - rep helicase Rep 3.1 2926.1 4629 Ipp2869 3270679 3270828 m - unknown hypothetical gene 6 Similar to disulfide bond 2928.2 4630 Ipp2870 3271036 3271881 P - unknown chaperones of the HSP33 family 3.9 2929.1 4631 Ipp2871 3271895 3272392 m - unknown Similar to hypothetical protein 5.2 Similar to conserved hypothetical 753.2 6439 Ipp2872 3272659 3273618 m + unknown protein 5.2 754.2 6440 Ipp2873 3273796 3274806 P - unknown Similar to predicted oxidoreductase 2.1.1 755.3 6441 Ipp2874 3274914 3275726 P - unknown similar to unknown protein 5.2 Similar to GTP-binding protein 2604.3 4460 Ipp2875 3275838 3277664 P - typA unknown TypA/BipA 3.7.4 Similar to probable inorganic 2602.2 4459 Ipp2876 3277671 3278558 P - unknown polyphosphate/ATP-NAD kinase 2.5 2412.2 4336 Ipp2877 3278562 3280229 P - recN DNA repair protein recN 3.2 2413.1 4337 Ipp2878 3280385 3280591 P - unknown Similar to cold shock protein CspC 4.1 2414.2 4338 Ipp2879 3280794 3281861 P - unknown Similar to efflux protein 1.2 2932.3 4633 Ipp2880 3281866 3284910 P - unknown similar to unknown protein 5.2
Table XIV Similar to conserved hypothetical 2933.2 4634 Ipp2881 3284999 3285910 p unknown protein 5.2 2934.1 4635 Ipp2882 3286057 3287313 p unknown 6 substrate of the Legionella pneumophila Dot/Icm 2938.4 4636 Ipp2883 3287380 3291033 m sidH system, truncated C_terminal part 5.2 Similar to transposase (IS21 616.5 6340 Ipp2884 3291076 3292122 p Unknown family) 4.5 Similar to transposase (IS21 591.4 6304 Ipp2885 3292119 3292913 p Unknown family) 4.5 substrates of the Legionella pneumophila Dot/Icm 590.4 6299 Ipp2886 3292962 3296060 m sidH system, truncated N_terminal part 5.2 conserved ubiquitin conjugation 5656.2 6238 Ipp2887 3296181 3296903 m unknown factor E4 family domain (U-box) 5.2 5668.2 6240 Ipp2888 3297191 3299038 p unknown 6 5669.2 6241 Ipp2889 3299095 3300735 m unknown 6 5671.2 6242 Ipp2890 3300732 3301112 m unknown Similar to acyl-CoA hydrolase 2.4 Similar to transcriptional accessory 719.4 66442200 Ipp2891 3301115 3303484 m unknown protein 3.5.2 Similar to thiopurine S- 722.3 6421 Ipp2892 3303608 3304273 m unknown methyltransferase 2.3 Glucosamine— fructose-6- 723.3 6422 Ipp2893 3304270 3306084 m glmS phosphate aminotransferase 2.1.1 4266.3 5472 Ipp2894 3306449 3307750 m unknown Similar to lysophospholipase A 2.4 6285.1 6362 Ipp2895 3307993 3309078 P Unknown Similar to transposase (IS4 family) 4.5 similar to conserved hypothetical 5337.1 6090 Ipp2896 3309104 3309397 P unknown protein 5.2 4907.2 5893 Ipp2897 3309640 3310404 P unknown similar to unknown protein 5.2 2297.3 4261 Ipp2898 3310516 3310986 P unknown similar to SsrA-binding protein 3.6 Highly similar to bacterioferritin 2295.3 4260 Ipp2899 3310998 3311462 P unknown comigratory protein 5.2
Table XIV 1812.4 4009 Ipp2900 3311586 3312785 unknown similar to efflux protein 1.2 318.2 4791 Ipp2901 3312940 3314346 unknown similar to PhoH protein 5.2 similar to guanosine 317.1 4783 Ipp2902 3314369 3315382 unknown monophosphate reductase GuaC 2.3 315.4 4769 Ipp2903 3315416 3316705 m - unknown protein with repetitions 6 2034.2 4129 Ipp2904 3317220 3317981 P - unknown similar to esterase 2.4 1482.4 3794 Ipp2905 3318617 3319579 m - unknown 6 4666.2 5737 Ipp2906 3319718 3320698 m + unknown Similar to ribonuclease 3.6 4668.1 5738 Ipp2907 3320861 3322165 P - unknown similar to unknown proteins 5.2 4669.2 5739 Ipp2908 3322266 3322481 m - unknown Similar to cold shock protein 4.1 2068.3 4147 Ipp2909 3322720 3324177 m + unknown 6 1014.2 3518 Ipp2910 3324458 3325840 m - unknown similar to amidase 2.1.1 2070.5 4149 Ipp2911 3326077 3327726 m + unknown similar to unknown protein 5.2 hypothetical tetratricopeptide 3054.4 4710 Ipp2912 3327723 3328604 m - unknown repeat protein 6 3055.1 4711 Ipp2913 3328601 3329533 m - unknown similar to unknwon proteins 5.2 3057.1 4712 Ipp2914 3329523 3330557 m - unknown similar to unknow protein 5.2 2289.2 4255 Ipp2915 3330545 3331030 m - unknown Similar to hypothetical protein 5.2 2290.1 4256 Ipp2916 3331023 3331934 m - unknown similar to unknown protein 5.2 Similar to MoxR-like ATPases, 2291.2 4257 Ipp2917 3331944 3332939 m - unknown putative regulator 3.5.2 3058.1 4713 Ipp2918 3333116 3333559 m - unknown 6 Similar to tRNA-dihydrouridine 555.2 6184 Ipp2919 3333641 3334627 P - unknown synthase B 3.6 553.1 6173 Ipp2920 3334686 3335045 m - Unknown 6 551.2 6164 Ipp2921 3335102 3335881 m - Unknown 2.5 Similar to conserved hypothetical 2423.2 4344 Ipp2922 3336034 3337146 P - unknown protein 5.2 2422.3 4343 Ipp2923 3337149 3337643 P - unknown Similar to unknown protein 5.2 Primosomal protein N 1569.8 3848 Ipp2924 3337644 3339821 P - priA (replication factor Y) 3.1 5769.4 6269 Ipp2925 3339818 3340210 P - unknown similar to unknown protein 5.2 5437.3 6132 Ipp2926 3340328 3340597 m + Unknown 6. 3720.2 5128 Ipp2927 3340673 3341467 m - thyA Thymidylate synthase 2.3
Table XIV prolipoprotein 899.3 6520 Ipp2928 3341464 3342234 m Igt diacylglyceryl transferase 3.8 898.2 6519 Ipp2929 3342252 3343058 m unknown similar to predicted permeases 1.2 phosphoenolpyruvate- 897.2 6518 Ipp2930 3343055 3345349 m ptsP protein phosphotransferase PtsP 1.2 Similar to probable (di)nucleoside 393.1 5247 Ipp2931 3345350 3345877 m unknown polyphosphate hydrolase NudH 2.3 396.5 5259 Ipp2932 3346230 3347240 p asg unknown Similar to L-asparaginase I 2.2 821.2 6471 Ipp2933 3347338 3348222 m unknown 6 Bifunctional GlmU protein, UDP-N-acetylglucosamine 823.5 6472 Ipp2934 3348344 3349729 glmU pyrophosphorylase and Glucosamine-1-phosphate N-acetyltransferase 1.1 arsenite efflux membrane 825.4 6473 Ipp2935 3349746 3350987 IrsB component-like protein 1.2 4194.3 5429 Ipp2936 3351014 3352303 m Unknown 5.1 putative cobalt/magnesium 3614.3 5058 Ipp2937 3352573 3353637 corA uptake transporter 1.2 760.2 6444 Ipp2938 3353750 3355483 m unknown 6 758.1 6443 Ipp2939 3355742 3356377 m nth Endonuclease III 3.2 Similar to electron transport 757.1 6442 Ipp2940 3356370 3356984 m unknown complex protein rnfB 1.4 1338.3 3710 Ipp2941 3356984 3358996 m metG methionyl-tRNA synthetase 3.7.2 Belongs to the polyprenyl P- 1339.2 3711 Ipp2942 3359265 3359834 p unknown hydroxybenzoate / phenylacrylic acid decarboxylases family. 2.1.1 1341.2 3712 Ipp2943 3359890 3360627 m unknown 6 5766.1 6268 Ipp2944 3360929 3361471 m unknown Putative secreted protein 6
Table XIV Highly similar to transcriptional 2882.2 4605 Ipp2945 3361526 3362212 m + unknown regulator ExsB 3.5.2 Highly similar to mannose-1- phosphate 2167.2 4204 Ipp2946 3362302 3363738 P - unknown guanylyltransferase/mannose-6- phosphate isomerase 1.1 2166.2 4203 Ipp2947 3363966 3365879 P - unknown 6 Highly simislar to glucose-inhibited 1976.2 4102 Ipp2948 3366010 3367884 P - gidA unknown division protein A GidA Similar to glucose inhibited division 1978.1 4103 Ipp2949 3367881 3368507 P - gidB unknown protein B GidB Highly similar to chromosome 1979.1 4104 Ipp2950 3368510 3369280 P - parA unknown partitioning protein ParA family similar to chromosome partitioning 122.4 3640 Ipp2951 3369294 3370199 P + parB unknown protein parB 121.1 3635 Ipp2952 3370307 3370642 P - unknown Similar to hypothetical protein 119.1 3625 Ipp2953 3370963 3371412 m - Unknown Similar to transposase (IS5 family) 118.1 3618 Ipp2954 3371800 3372387 m - unknown 6 116.1 3609 Ipp2955 3373298 3373855 m - unknown 6 115.2 3603 Ipp2956 3374194 3374700 m - unknown Similar to acetyltransferases 2.1.1 114.2 3598 Ipp2957 3374783 3374926 m - unknown hypothetical gene 6 2884.1 4606 Ipp2958 3375531 3375803 P - unknown 6 cytochrome c oxidase, 1750.3 3967 Ipp2959 3375826 3376695 m - coxC subunit III + cytochrome c oxidase 1752.2 3968 Ipp2960 3376854 3377396 m ctaG assembly protein cytochrome c oxidase, 1691.3 3928 Ipp2961 3377409 3379025 m - coxA subunit I cytochrome c oxidase, 1689.3 3926 Ipp2962 3379022 3380227 m + coxB subunit II 4542.4 5660 Ipp2963 3380309 3381922 m + unknown similar to cytochrome c similar to ferredoxin component of 2372.3 4310 Ipp2964 3382304 3382657 P - Unknown dioxygenase 2371.1 4309 Ipp2965 3382723 3383751 P - unknown similar to glycosyltransferases 2369.3 4308 Ipp2966 3383810 3384415 P + unknown Similar to transporter, LysE family 4546.3 5661 Ipp2967 3384600 3385652 m - unknown 6 3009.2 4691 Ipp2968 3386173 3387123 P + unknown 6
Table XIV Similar to conserved hypothetical 2579.2 4444 Ipp2969 3387424 3387849 p unknown protein 5.2 Ubiquinone/menaquinone 2580.2 4445 Ipp2970 3387974 3388726 p ubiE biosynthesis methyltransferase 2.5 similar to conserved hypothetical 3008.1 4690 Ipp2971 3388734 3389351 p unknown protein 5.2 similar to P.aeruginosa probable 182.3 4014 Ipp2972 3389441 3391090 p ubiB unknown ubiquinone biosynthesis protein ubiB 2.5 Putative TatA protein(twin 180.1 4001 Ipp2973 3391092 3391277 p tatA arginine translocation) 1.6 Putative TatB protein (twin 179.1 3994 Ipp2974 3391274 3391543 p tatB arginine translocation) 1.6 178.1 3986 Ipp2975 3391586 3391846 m - unknown 6 176.1 3973 Ipp2976 3392008 3392991 P - unknown 6 Highly similar to peptide 174.1 3959 Ipp2977 3393015 3393884 m - unknown methionine sulfoxide reductase 4.2 172.1 3946 Ipp2978 3394545 3394733 P - unknown similar to hypothetical protein 5.2 2044.2 4134 Ipp2979 3394781 3396715 m - unknown similar to copper amine oxidase 2.2 323.2 4813 Ipp2980 3397955 3399442 P - unknown 6 321.3 4804 Ipp2981 3399827 3400720 m - unknown 6 6277.1 6360 Ipp2982 3400988 3401179 P - unknown Hypothetical gene 6 Weakly similar to 3005.2 4689 Ipp2983 3401767 3402270 m - unknown acetyltransferases 2.1.1 Weakly similar to conserved 5208.1 6042 Ipp2984 3402282 3402566 m - unknown hypothetical protein 5.2 1817.2 4011 Ipp2985 3403078 3403581 m - unknown 6 1815.2 4010 Ipp2986 3403971 3404498 m - unknown Similar to acetyltransferase 2.1.1 2374.2 4312 Ipp2987 3404826 3405722 m - unknown 6 Highly similar to putative lytic 2373.2 4311 Ipp2988 3406047 3407009 m + unknown murein transglycosylase 1.1 similar to ABC transporter 412.3 5373 Ipp2989 3407344 3408537 m + unknown permease protein 1.2 •imilar to ATP-binding protein of 410.1 5358 Ipp2990 3408534 3409223 m - unknown ABC transporter 1.2 409.2 5352 Ipp2991 3409216 3410361 m - unknown conserved lipoprotein 1.2
Table XIV Similar to outer membrane efflux 408.3 5345 Ipp2992 3410358 3411986 m unknown protein 1.2 similar to bis(5 -nucleosyl)- 1983.1 4105 Ipp2993 3412030 3412860 m apaH unknown tetraphosphatase ApaH 2.3 1986.2 4106 Ipp2994 3412853 3414190 m unknown similar to unknown protein 5.2 similar to dimethyladenosine 2800.1 4568 Ipp2995 3414227 3414997 m ksgA unknown transferase ( 16S rRNA dimethylase) 3.6 similar to aspartate 1- 2799.1 4567 Ipp2996 3415333 3415734 p panD unknown decarboxylase 2.5 Similar to Sec-independent protein 2199.2 4214 Ipp2997 3415840 3416565 p tatC unknown translocase TatC 1.6 similar to conserved hypothetical 2198.1 4213 Ipp2998 3416627 3416938 p unknown protein 5.2 2197.1 4212 Ipp2999 3416976 3417170 m Unknown 6. Similar to C-terminal part of 2196.1 4211 Ipp3000 3417188 3417901 m unknown NAD (P)H -flavin reductase 2.1.1 Highly similar to 3-polyprenyl-4- 2195.1 4210 Ipp3001 3417901 3419367 m ubiD unknown hydroxybenzoate decarboxylase and related decarboxylases 2.5 transcription termination 2193.4 4209 Ipp3002 3419513 3420775 m rho factor Rho 3.5.4 841.3 6483 Ipp3003 3421098 3421424 m trxA unknown Highly similar to thioredoxin 1.4 Similar to conserved hypothetical 840.2 6482 Ipp3004 3421637 3422371 m unknown protein 5.2 fumarate hydratase, class 838.3 6480 Ipp3005 3422616 3424010 fumC II 2.1.3 2112.2 4167 Ipp3006 3424223 3425650 m Unknown similar to unknown protein 5.2 1891.2 4056 Ipp3007 3425932 3426693 P unknown Similar to hypothetical protein 4251.2 5466 Ipp3008 3426719 3428029 P unknown 6 4249.1 5464 Ipp3009 3428505 3429689 m unknown 6 4248.1 5463 Ipp3010 3429757 3430020 m unknown similar to unknown protein 5.2 1081.2 3559 Ipp3011 3430495 3431640 P unknown 6 similar to lipid A biosynthesis 1080.2 3558 Ipp3012 3431846 3432682 m unknown lauroyi acyltransferase 2.4 4247.2 5462 Ipp3013 3432675 3433523 m unknown similar to acyl transferase 2.4 Similar to lipid A-disaccharide 2353.3 4295 Ipp3014 3433682 3434839 P IpxB unknown synthase 1.1
α < Q XI U X X (J Q. Q. E ( >J- lc J >- Q.
+ + + ε ε ε ε ε Q* E U α ε ε E E
cn rT rv rr rv in m rr vO in cn rr ro ro rv rr in ro CO in 00 CO 00 cn cn rv in O 00 rr o cn rr ro rr rr rT in vo 00 n o m rv rv co o ΓM ro rr in ro ro rr rT rT T rr rr rr rT rT r in in in in in rT rr rr rT rr rr r rT rr rT rr rr rT rT r rr rr rT rr rr rr oo ro ro ro ro ro ro ro ro ro ro ro ro oo ro ro ro in vD rv CO cn o ΓM rT m rv 00 cn ro rr in ΓM ΓM ΓM (N ΓM ΓM ΓM ro ro ro ro oo o o o o o o o o o o o o O o o o o o o o ro ro ro oo ro ro ro ro ro ro ro oo ro oo o. α. o. o. o. rr o ΓM rr in in rv O C0 oo rv vo o cn cn cn σ> cn cn cn cn m cn ΓM ΓM m I
s- ro σ> in ΓM ΓM o o o o o o o cn cn cn vD vD cn o rr rr in m in in m in in in rr rT rr in in in rT rr VD vo

Table XIV 526.2 6061 Ipp3036 3457628 3458251 p rnsT ribonuclease T 2.3 alkyl hydroperoxide 528.2 6067 Ipp3037 3458377 3458982 m ahpC reductase 4.2 529.2 6071 Ipp3038 3459035 3459382 m grlA glutaredoxin-like protein 4.2 3609.1 5053 Ipp3039 3459683 3460261 p sodB superoxide dismutase, iron 4.2 Similar to omithine/acetylornithine 3610.1 5054 Ipp3040 3460321 3461490 p argD unknown aminotransferase 2.2 Similar to conserved hypothetical 3611.2 5055 Ipp3041 3461619 3462386 m unknown protein 5.2 Similar to glycerophosphoryl 3612.2 5056 Ipp3042 3462542 3463324 m unknown diester phosphodiesterase (ATA start codon) 2.4 Similar to NAD-linked malate 3613.4 5057 Ipp3043 3463576 3465246 p unknown dehydrogenase (malic enzyme) 2.1.1 Similar to putative translation 5905.3 6303 Ipp3044 3465476 3466444 P - unknown initiation protein 3.7.3 6273.1 6359 Ipp3045 3466542 3466625 P - unknown hypothetical gene 5.2 Phosphatidylserine 2828.5 4584 Ipp3046 3466618 3467469 m - psd decarboxylase 2.4 2827.3 4583 Ipp3047 3467562 3470177 m - unknown 6 2826.1 4582 Ipp3048 3470440 3471969 m - unknown 6 2119.2 4169 Ipp3049 3472423 3474084 m + unknown similar to protease 2.2 909.2 6530 Ipp3050 3474336 3475190 P - unknown hypothetical protein 5.2 DNA topoisomerase IV, A 910.4 6531 Ipp3051 3475297 3477552 m - parC subunit 3.4 Highly similar to H+-transporting 2824.1 4581 Ipp3052 3478181 3478603 m - atpC unknown ATP synthase epsilon chain 1.4 Highly similar to H+- transporting 1556.2 3839 Ipp3053 3478615 3479991 m atpD unknown ATP synthase beta chain 1.4 Highly similar to H+- transporting 1555.2 3838 Ipp3054 3480004 3480870 m atpG unknown ATP synthase chain gamma 1.4 Highly similar to H+-transporting. 1552.3 3837 Ipp3055 3480960 3482513 m atpA unknown ATP synthase chain alpha 1.4
Table XIV Highly similar to H+-transporting 2140.2 4185 Ipp3056 3482531 3483073 m ~ atpH unknown ATP synthase chain delta 1.4 Highly similar to H+-transporting 2138.2 4183 Ipp3057 3483075 3483545 m - atpF unknown ATP synthase chain b 1.4 Highly similar to H+-transporting 2137.1 4182 Ipp3058 3483597 3483872 m - atpE unknown ATP synthase chain c 1.4 Highly similar to H+- transporting 2136.1 4181 Ipp3059 3483913 3484749 m - atpB unknown ATP synthase chain a 1.4 2135.1 4180 Ipp3060 3484742 3485137 m + atpl unknown Similar to ATP synthase subunit i 1.4 2133.1 4179 Ipp3061 3485291 3485536 P - Unknown 6. 2132.2 4178 Ipp3062 3485606 3486193 m + unknown Similar to lipoproteins 1.2 2130.1 4177 Ipp3063 3486195 3486767 m + unknown similar to lipoproteins 1.2 similar to phosphoheptose 2129.1 4175 Ipp3064 3486771 3487370 m - unknown isomerase 2.1.1 similar to conserved hypothetical 2127.2 4174 Ipp3065 3487397 3487753 m - unknown protein 5.2 conserved hypothetical protein, 2822.1 4580 Ipp3066 3487775 3489586 m + unknown putative lipoprotein 5.2 conserved hypothetical protein, 2820.1 4579 Ipp3067 3489626 3490477 P - unknown putative methyltransferase 5.2 2818.1 4577 Ipp3068 3490607 3491674 P - unknown similar to alkane monooxygenase 2.1.1 Similar to sulfate permease and related transporters (MFS 2816.2 4576 Ipp3069 3491725 3493890 m unknown superfamily) with a C-terminal cAMP bindinq motif. 1.2 2815.2 4575 Ipp3070 3494454 3496214 p unknown 6 Similar to eukaryotic zinc 2565.4 4436 Ipp3071 3496419 3497219 p unknown metalloproteinase 2.2 1176.5 3616 Ipp3072 3497216 3499054 m unknown 6 Similar to GTPase for tRNA 1174.2 3615 Ipp3073 3499350 3500690 m unknown modification trmE 3.6 5040.2 5970 Ipp3074 3500691 3502361 m + unknown putative inner membrane protein 5.2
Table XIVr Similar to conserved hypothetical 5039.2 5969 Ipp3075 3502371 3502616 m unknown protein 5.2 similar to ribonuclease P protein 5038.1 5968 Ipp3076 3502583 3502927 m rnpA unknown component (RNase P) 3.6 5037.2 5967 Ipp3077 3502928 3503062 m rpmH 50S ribosomal protein L34 3.7.1 highly similar to partition protein 739.5 6715 plppOOOl 66 941 m parB unknown ParB 3.4 Highly similar to partition protein 4947.2 6684 plpp0002 1297 2505 P unknown SopA/ParA 3.4 similar to hypothetical replication 4945.2 6683 plpp0003 2507 3514 P unknown protein RepB 3.1 similar to hypothetical replicative 5078.2 6685 plpp0004 3552 4841 P unknown protein RepA 3.1 5081.5 6687 plpp0005 4992 5558 P + unknown similar to unknown proteins 5.2 similar to major facilitator 5080.6 6686 plpp0006 5591 7045 P unknown superfamily (MFS) transporter 1.2 5106.3 6688 plpp0007 7047 8069 m unknown 6 6227.1 6702 plppOOOδ 8091 8285 m Unknown 6. 5147.1 6691 plpp0009 8469 9731 P unknown 6 3500.3 6674 plppOOlO 9688 10866 P Unknown similar to unknown protein 5.2 similar to molybdopterin converting 3499.1 6673 plppOOll 11016 11267 Unknown factor 2 (subunit 1) similar to molybdenum cofactor 1226.2 6597 plpp0012 11272 12294 Unknown synthesis protein 3 Weakly similar to conserved 1225.2 6596 plpp0013 12291 13352 unknown hypothetical proteins 2275.4 6632 plpp0014 13345 14379 P Unknown 6229.1 6703 plpp0015 14492 14677 P Unknown 6. Weakly similar to carbon storage 6230.1 6704 plpp0016 14712 14972 unknown regulator CsrA 3.5.2 Similar to E.coli TraT complement 1171.4 6594 plpp0017 14997 15743 + unknown resistance protein 4.5 1172.2 6595 plpp0018 15837 16181 P unknown 6 3498.1 6672 plpp0019 16301 16657 P + unknown 6 6231.3 6705 plpp0020 16712 17758 P unknown similar to transposase 4.5 similar to plasmid DNA replication 6224.3 6701 plpp0021 17755 18549 unknown protein 3.1
Table XIV similar to fimbrial protein precursor 2659.3 6638 plpp0022 18713 19036 P + unknown (Pilin) 1.8 2660.1 6639 weakly similar to conjugative plpp0023 19026 19316 P - unknown transfer protein TraL 4.5 2661.2 6640 plpp0024 19321 19878 Weakly similar to conjugative P - unknown transfer protein TraE 4.5 2662.2 6641 Weakly similar to conjugative plpp0025 19875 20594 P + unknown transfer protein TraK 4.5 485.3 weakly similar to conjugative 6681 plpp0026 20609 22063 P + unknown transfer protein TraB 4.5 486.2 6682 plpp0027 22056 22283 P - unknown hypothetical gene 6 weakly similar to conjugative 254.2 6633 plpp0028 22313 24889 P - unknown transfer protein TraC 4.5 255.1 6634 plpp0029 24879 25214 P + unknown Weakly similar to Trbl protein 4.5 weakly similar to conjugative 256.1 6635 plpp0030 25211 25837 P - unknown transfer protein TraW 4.5 258.2 6637 similar to conjugative transfer plpp0031 25849 26844 P + unknown protein TraU 4.5 2006.2 6616 plpp0032 Weakly similar to conjugative 26861 27523 P + unknown transfer protein TrbC 4.5 1344.4 Similar to putative conjugative 6603 plpp0033 27516 29315 P + Unknown transfer protein TraN 4.5 Similar to putative conjugative 1342.3 6602 plpp0034 29312 30100 P + unknown transfer protein TraF 4.5 Weakly similar to putative 2051.2 6617 plpp0035 30097 30594 unknown conjugative transfer protein TrbB Similar to conjugative transfer 1299.3 6601 plpp0036 30612 31988 P unknown protein TraH similar to sex pilus assembly and 567.6 6696 plpp0037 31985 34744 P unknown mating pair protein TraG Highly similar to conjugative 2212.4 6626 plpp0038 34823 36826 P unknown transfer protein TraD similar to conjugative transfer 1599.5 6610 plpp0039 36848 42715 P unknown protein Tral 2665.1 6642 plpp0040 42895 43176 P unknown 6 2666.1 6643 plpp0041 43309 43629 m unknown 6 2667.2 6644 plpp0042 43620 44069 m Unknown 6 717.3 6713 plpp0043 44445 44891 P unknown 6
Table XIV 716.1 6712 plpp0044 44933 45136 P unknown 6 715.1 6711 plpp0045 45305 45622 P unknown 6 713.2 6710 plpp0046 45695 46039 P unknown 6 similar to putative anti restriction 5844.1 6697 plpp0047 46327 46731 m unknown protein KlcA 3.2

4483.2 6679 plpp0048 46854 47477 m unknown 6 Similar to abortive infection 4482.1 6678 plpp0049 47722 48621 unknown bacteriophage resistance protein 4.4 Similar to eukaryotic 4481.1 6677 plpp0050 48703 49233 m unknown hypersensitive-induced response protein 3632.3 6676 plpp0051 49324 52371 m Unknown Similar to transposase (Tn3 family) 2244.2 6627 plpp0052 52521 53126 P unknown Similar to resolvase TnpR Similar to conserved hypothetical 2245.2 6628 plpp0053 53129 54202 P unknown protein Similar to ABC transporter (ATP- 2247.7 6629 plpp0054 54192 55985 P unknown binding protein) 1 783.2 6721 plpp0055 56024 56380 unknown similar to unknown proteins 5 785.4 6722 plpp0056 56410 57264 m unknown 6 5846.2 6698 plpp0057 57242 57415 m Unknown 6. 3631.4 6675 plpp0058 57472 57702 m unknown 6 6016.1 6700 plpp0059 57783 58091 m unknown 6 6015.1 6699 plpp0060 58262 58546 P unknown hypothetical gene 6 5113.3 6689 plpp0061 58548 58988 P unknown 6 5114.3 6690 plpp0062 59104 59916 P unknown 6 3138.2 6659 plpp0063 59972 60844 P unknown 6 3137.1 6658 plpp0064 60896 61381 P Unknown 6. Weakly similar to conserved 3136.1 6657 plpp0065 61462 61791 unknown hypothetical protein 5.2 Similar to ATPase components of 799.3 6725 plpp0066 62005 63480 unknown ABC transporters 1.2 Similar to multidrug resistance ABC 798.2 6724 plpp0067 63461 65260 unknown transporter ATP-binding protein 1.2
Table XIV Weakly similar to Legionella 796.2 6723 plpp0068 65253 66035 unknown longbeachae spectinomycin 3 adenylyltransferase 5.1 N-terminal part similar to 2087.2 6618 plpp0069 66036 66995 P - unknown conserved hypothetical protein 5.2 2180.2 6625 plpp0070 66992 68476 P - unknown Fusion of three acetyltransferase 2.1.1 Similar to conserved hypothetical 3135.1 6656 plpp0071 68473 68685 P - unknown protein 5.2 3134.1 6655 plpp0072 68678 69250 P - unknown Similar to hypothetical protein 5.2 Weakly similar to MerR family 1762.2 6612 plpp0073 69327 70271 P - unknown transcriptional regulators 3.5.2 1761.4 6611 plpp0074 70485 73598 P - unknown penicilin binding protein domain 1.1 1575.2 6609 plpp0075 73929 74351 P - unknown Similar to transcriptional regulators 3.5.2 similar to penicillin-binding protein 3346.2 6660 plpp0076 74354 76276 P - unknown 2 (Ftsl) 1.1 3347.1 6661 plpp0077 76344 77144 P + unknown Similar to beta-lactamase 4.2 hypothetical polysaccharide 3348.3 6662 plpp0078 77148 78728 P + Unknown deacetylase-related, fusion protein 2.1.1 Similar to conserved hypothetical 3349.3 6663 plpp0079 78732 79628 P - unknown protein 5.2 3350.3 6664 plpp0080 79771 80133 P - unknown Similar to unknown protein 5.2 Similar to dihydrofolate reductase 3351.3 6665 plppOOδl 80153 81325 unknown and methyltransferase 2.5 Similar to acetyltransferase, GNAT 3352.1 6666 plpp0082 81354 81824 m unknown family 2.1.1 3353.1 6667 plpp0083 81973 83481 P unknown Similar to hypothetical protein 5.2 Similar to pyridoxamine 5 - 3354.1 6668 plpp0084 83532 84107 P unknown phosphate oxidase 2.5 960.2 6732 plpp0085 84116 84427 P unknown similar to unknown protein 5.2 959.2 6731 plpp0086 84483 84665 P unknown hypothetical gene 6 Weakly similar to alpha/beta 957.2 6730 plpp0087 84759 85634 m unknown hydrolase fold family protein 2.1.1 956.2 6729 plpp0088 85692 86453 m unknown Similar to transcriptional regulator 3.5.2
Table XIV Weakly similar to stability protein 3355.2 6669 plpp0089 86666 86953 m unknown StbE 4.5 Weakly similar to stability protein 3356.2 6670 plpp0090 86943 87194 m unknown StbD 4.5 3357.2 6671 plpp0091 87443 87910 m unknown 6 Highly similar to spectinomycin 3 5154.2 6695 plpp0092 88529 89350 m unknown adenylyltransferase 4.2 Highly similar to Legionella 5153.1 6694 plpp0093 89334 89771 m unknown longbeachae unknown protein 5.1 Highly similar to Legionella 2573.3 6636 plpp0094 89758 90336 m unknown longbeachae unknown protein 5.1 2881.3 6650 plpp0095 90624 91529 m unknown 6 2879.4 6649 plpp0096 91618 92943 m unknown similar to unknown protein 5.2 Highly similar to Legionella 2878.2 6648 plpp0097 93135 93827 IrpR unknown longbeachae putative transcriptional regulatory protein 3.5.2 Highly similar to Legionella 1362.3 6604 plpp0098 94067 95890 unknown longbeachae unknown protein, ankyrin repeat containing protein 5.1 Highly similar to L.longbeachae 1364.2 6605 plpp0099 95894 97498 m IskS unknown sensor histidine kinase protein 1.3 2877.2 6647 plppOlOO 97597 98253 m unknown Similar to hypothetical protein 5.2 2175.3 6622 plppOlOl 98341 98559 m unknown 6 2174.1 6621 plpp0102 98563 98859 m unknown conserved hypothetical protein 5.2 1078.2 6593 plpp0103 99016 99993 P unknown 6 Similar to hypothetical 1077.1 6592 plpp0104 100101 100793 P unknown transcriptional regulator 3.5.2 1076.3 6591 plpp0105 100869 101237 P unknown conserved hypothetical protein 5.2 2173.2 6620 plpp0106 101240 101668 P unknown similar to hypothetical protein 5.2 Weakly similar to probable 2172.2 6619 plpp0107 101692 102153 P unknown transcriptional regulator 3.5.2 Weakly similar to DNA alkylation 2874.1 6646 plpp0108 102176 102919 P unknown repair enzyme 3.2 2873.1 6645 plpp0109 102950 103921 m unknown Similar to hypothetical protein 5.2 C-terminal part similar to 761.2 6716 plppOHO 103938 104891 m unknown acetyltransferase 2.1.1
able XIV weakly similar to lysE family 762.2 6717 plppOlll 105014 105661 P unknown transporter protein 1.2 764.1 6718 plpp0112 105738 106253 P unknown similar to acetyltransferase protein 2.1.1 Similar to N-terminal part of 765.1 6719 plpp0113 106325 107044 m unknown hypothetical protein 5.2 766.4 6720 plpp0114 107216 108649 m unknown similar to esterase 4.1 886.4 6728 plpp0115 108721 109491 m unknown similar to shikimate kinase (AroK) 2.2 similar to aminoglycoside 3 - 885.3 6727 plpp0116 109545 111149 P unknown phosphotransferase 4.2 884.3 6726 plpp0117 111146 112111 P unknown 6 Putative adenine-specific DNA
4549.2 6680 plppOllδ 112324 113640 P unknown methylase 3.2
1237.2 6600 plpp0119 113656 114093 P unknown Weakly similar to acetyltransferase 2.1.1
1236.2 6599 plpp0120 114212 115009 m unknown 5.1
1235.3 6598 plpp0121 115070 115798 m unknown Weakly similar to acetyltransferase 2.1.1
5152.1 6693 plpp0122 115604 116190 m unknown Similar to unknown protein 5.2
5151.1 6692 plpp0123 116238 117131 m unknown similar to unknown protein 5.2 bifunctional protein, similar to lδ4δ.5 6613 plpp0124 117144 116394 m unknown acetyl transferase and to methyl transferase 4.6 similar to acetyltransferase, GNAT
1849.4 6614 plpp0125 118391 118855 m unknown family 2.1.1 Similar to conserved hypothetical
1852.2 6615 plpp0126 118878 119234 m unknown protein 5.2 similar to acetyltransferase (C-
1522.2 6606 plpp0127 119231 120406 m unknown terminal part) 4.6
1523.2 6607 plpp0128 120457 120729 m unknown 6 Some similarity with transcriptional
1524.3 6608 plpp0129 120790 121833 m unknown regulator, MerR family 3.5.2
2258.2 6630 plpp0130 122086 123333 m unknown 6
2260.2 6631 plpp0131 123311 123985 m unknown Similar to alanyl tRNA synthetase 3.7.2
3030.2 6651 plpp0132 123989 124480 m unknown Similar to hypothetical protein 5.2
3031.2 6652 plpp0133 124497 124919 m unknown 6
Table XIV 3035.2 6653 plpp0134 125023 125598 P unknown Weakly similar to transcriptional regulator 3.5.2 659.2 6707 plpp0135 125603 126556 m unknown Similar to conserved hypothetical protein 5.2 660.1 6708 plpp0136 126553 127119 m unknown Weakly similar to acetyltransferase 2.1.1 661.3 6709 plpp0137 127106 128239 m C-terminal part similar to unknown acetyltransferase 2.1.1 3036.2 6654 plpp0138 128246 128788 m Weakly similar to DNA topology unknown modulation protein 3.4 2176.3 6623 plpp0139 128871 129329 m weakly similar to regulatory protein unknown recX 3.3 2177.2 6624 plpp0140 129429 130013 P unknown Similar to integrase proteins 4.5 737.2 6714 olpp0141t 130347 131369 m similar to ATP-dependent DNA unknown helicase RecG(C-terminal part) 3.3 6429.1 6706 plpp0141c 131326 131850 m unknown Similar to N-terminal part of ATP- dependent DNA helicase recG 3.3
Table XVI: SEQ EMBL Name of Product of ORF Posit°l Posit°2 Sens SignalP .. Note Class ID NAME " the gene the gene 1955.1 6810 lpl0904 1015112 1016491 m unknown 6 2100.1 6811 lpllOOό 1140978 1141112 P unknown hypothetical gene 6 Some similarity with RNA-directed
2110.1 6812 lpll015 1147633 1149258 m unknown 3.1 DNA polymerase
2134.1 6813 lpll034 1164732 1165319 m unknown 6 2135.1 6814 lpll035 1165316 1165729 m + unknown Similar to PilL protein 1.8 2136.1 6815 lpll036 1165738 1165938 m unknown Similar to carbon storage regulator 3.5.2 2137.1 6816 lpll037 1165932 1166450 m unknown Similar to Legionella LvrB protein 5.1 2139.1 6817 lpll039 1167370 1167624 P unknown Similar to phage repressor 4.4 2141.1 6818 Ipl 1041 1168453 1168914 m + unknown Similar to putative lipoprotein 5.1 Similar to conserved hypothetical
2142.2 6819 Ipl1042 1169480 1170091 m unknown 5.2 protein Similar to C-terminal part of
2144.1 6820 lpll043b 1170400 1170588 m unknown 1.2 arginine-binding periplasmic protein Similar to central part of arginine-
2145.1 6821 lpll043a 1170581 1170910 m unknown 1.2 binding periplasmic protein
2151.1 6822 lpll047 1176999 1177199 m + unknown Hypothetical protein 6 Weakly similar to transcriptional
2152.2 6823 lpll048 1177363 1177659 p unknown 3.5.2 regulator
2155.1 6824 lpll050 1180093 1180326 m Unknown 6
2157.1 6825 lpllOSl 1180809 1180973 p unknown Hypothetical protein 6 Similar to C-terminal part of
2168.1 6826 Ipl1058 1186757 1186903 m unknown 5.1 unknown protein Similar to EnhC protein, contains 7
2170.1 6827 lpll059 1187369 1188397 p unknown 4.6 sel-1 domains
2180.1 6828 lpll067 1194223 1195173 m unknown 6
Some similarity with helicase
2181.1 6829 Ipl1068 1195308 1198694 m unknown 3.2 proteins
2182.1 6830 Ipl1069 1198678 1200126 m unknown 6 Weakly similar to chemotaxis motB
2184.1 6831 Ipl1070 1200113 1200817 m unknown 1.5 protein
2186.2 6832 Ipl1071 1200795 1202768 m Unknown Hypothetical protein 6 Similar to conserved hypothetical
2189.1 6833 Ipl1073 1203936 1205711 P unknown 5.2 protein
2192.1 6834 Ipl1075 1207092 1207256 P unknown Hypothetical protein 6
2194.1 6835 Ipl1076 1207510 1208286 P unknown Similar to hypothetical protein 5.2
2195.1 6836 Ipl1077 1208304 1208747 m Unknown hypothetical gene 6
2198.1 6837 Ipl1080 1209864 1210679 m unknown 6
2199.1 6838 Ipl1081 1210788 1211342 P unknown 6
2200.1 6839 Ipl1082 1211557 1211928 P unknown 6 Similar to conserved hypothetical
2201.1 6840 Ipl1083 1211925 1212284 P unknown 5.2 protein Similar to DNA-damage-inducible
2202.1 6841 Ipl1084 1212352 1212606 P unknown 3.2 protein J Similar to phage-related integrase
2203.1 6842 Ipl1085 1212983 1214224 m unknown 4.4 proteins
2205.1 6843 Ipl1086 1214553 1214804 m unknown 6 Similar to conserved hypothetical
2206.1 6844 Ipl1087 1214933 1216825 m unknown 5.2 protein
2207.1 6845 lpll088 1216892 1217056 m unknown Hypothetical protein 6
2208.1 6846 Ipl1089 1217691 1218077 P Unknown 6
2209.1 6847 Ipl1090 1218170 1218307 P unknown Hypothetical protein 6 Weakly similar to DNA-binding
2210.1 6848 Ipl1091 1218451 1218768 m unknown 3.5.2 protein Similar to plasmid maintenance
2211.1 6849 Ipl1092 1219050 1219331 P unknown 4.5 system killer protein
Similar to plasmid maintenance
2213.1 6850 lpll093 1219338 1219661 p unknown 4 5 system antidote protein
2224.1 6851 lpll lOl 1225340 1226203 m unknown 2238.1 6852 Ipll l lO 1235461 1235754 p unknown Similar to transposase 4.5 4011.1 7007 lpll l l3 1237989 1238123 m unknown hypothetical protein 6 4007.1 7006 lpll 116 1239963 1240346 m unknown 6 3995.1 7005 lpll 125 1251579 1251812 m unknown 6 Similar to glutathione-regulated
3987.1 7004 lpll 132 1261359 1263053 p + unknown potassium-efflux system protein 1.2 kefC (K(+)/H(+) antiporter)
3976.1 7003 lpll 138 1269104 1270132 m unknown Similar to transposase, IS630 family 4.5 Some similarity with eukaryotic
3949.1 7002 lpll 158 1291134 1292444 m unknown 6 proteins
3937.1 7001 lpll 165 1301149 1301286 P unknown Hypothetical protein 6
4229.2 7027 Ipl 1393 1555031 1555522 m unknown 6 hypothetical, similar to hypothetical 175.1 6802 lpl0132 157452 157604 m unknown 6 proteins
4203.1 7026 lpll412 1575836 1576864 P unknown Similar to transposase, IS630 family 4.5 Similar to major facilitator family
4197.1 7025 lpll416 1581011 1582300 m unknown 1.2 (MFS) transporter Similar to conserved hypothetical
4196.1 7024 lpll417 1582309 1583310 m unknown 5.2 protein
4189.2 7023 lpll424 1590828 1592246 p unknown Similar to transposase 4.5 898.1 7057 lpll425 1592331 1592816 p unknown Similar to transposase (truncated) 4.5 761.1 7056 lpll536 1711241 1711603 m unknown 6 Weakly similar to mitomycin 760.1 7055 lpll537 1711726 1711977 m unknown 4.1 resistance protein 711.1 7054 lpll574 1755441 1755842 m unknown Hypothetical protein 6 709.1 7053 lpll575 1755944 1756645 m unknown Similar to replication protein 3.1 707.1 7052 lpll577 1757623 1757916 m unknown Similar to transposase 4.5
Similar to N-terminal part of
705.1 7051 lpll578 1757963 1758193 m unknown 3.1 replication protein
703.1 7050 lpll579 1758826 1760220 P unknown Similar to other proteins 6
159.1 6799 lpl0143 175902 177617 P unknown 6
700.1 7049 lpll580 1760423 1761040 m unknown 6 Similar to part of conjugal transfer
699.1 7048 lpllSδl 1761052 1762173 m unknown 4.5 protein traA
697.1 7047 lpll582 1762266 1762577 m unknown Similar to transposase 4.5
694.1 7046 Ipl1584 1763137 1763430 m unknown Similar to tranposase 4.5 Similar to N-terminal part of
692.1 7045 lρll585 1763491 1764834 m unknown 4.5 conjugal transfer protein TraA
690.1 7044 lpll586 1765077 1765298 P unknown Hypothetical protein 6
689.1 7043 Ipl1587 1765353 1765601 P unknown Similar to hypothetical protein 5.2 Weakly similar to stability protein
688.1 7042 Ipl1588 1765591 1765848 P unknown 4.5 StbE
157.1 6798 lpl0144 177672 179090 m unknown Similar to hypothetical protein 5.2 Similar to cytosine-specific DNA
156.1 6797 lpl0145 179293 180546 P unknown 3.2 methylase Similar to DNA mismatch repair
154.1 6796 lpl0146 180543 180968 P unknown 3.2 endonuclease Legionella vir similar to carbon storage regulator
148.1 6794 lpl0150 183077 183280 P lvrC 5.1 region protein gene csrA
566.1 7041 Ipl1676 1869326 1869454 m unknown Hypothetical protein 6
564.1 7040 Ipl1678 1870005 1870523 P unknown 6
562.1 7039 Ipl1679 1871002 1871832 m unknown 6
561.1 7038 lpll680 1872034 1873806 P unknown 6
560.1 7037 Ipl1681 1873869 1874516 P unknown Ankyrin repeat protein 4.6
127.1 6778 lpl0164 195255 195590 P + unknown 6
126.1 6777 lpl0165 195737 196612 m unknown Similar to transposase 4.5
125.1 6772 lpl0166 196771 196944 m unknown Hypothetical protein 6
124.1 6771 lpl0167 197056 197223 P + unknown Hypothetical protein 6
442.1 7036 Ipl 1768 1971829 1972026 m + Unknown Unknown 6 117.1 6762 lρl0171 201954 202217 P unknown some similarity with TraD protein 4.5 116.1 6761 lpl0172 202239 202805 P unknown 6 115.1 6760 lpl0173 202924 204147 P unknown 6 113.1 6758 lρl0174 204158 204652 P unknown 6 Similar to conserved hypothetical 112.1 6757 lpl0175 204671 205426 P unknown 5.2 protein 111.1 6756 lpl0176 205401 206120 m unknown Similar to other proteins 5.2 110.1 6754 lpl0177 206286 206567 P unknown 6
322.1 6941 lpll 857 2078673 2078882 m unknown Hypothetical protein 6 106.1 6752 lpl0180 208124 209491 m unknown Similar to predicted ATPase 5.2 Similar to predicted Rossmann fold 105.1 6749 lpioiβi 210028 210912 P unknown nucleotide-binding protein involved 1.2 in DNA uptake
293.1 6924 lpl 1879 2104753 2105166 m unknown similar to type IN pilin PilA 1.8 103.1 6740 lpl0182 210914 211552 P unknown 6 102.1 6738 lpl0183 211654 211914 P unknown Similar to hypothetical protein 5.2
273.2 6907 lpll 893 2121129 2122547 P unknown Similar to transposase 4.5
1048.1 6747 lpl 1894 2122544 2123488 P unknown Similar to transposase 4.5
1047.1 6746 lpll 895 2124195 2125265 m unknown Putative carbonic anhydrase 4.7
1045.1 6745 lpl 1896 2125661 2126002 P unknown 6
1043.1 6744 lpll 897 2126564 2126761 m unknown Hypothetical protein 6 Some similarity with hypothetical
1041.1 6743 lpl 1899 2128985 2131282 m unknown 5.2 protein
1040.1 6742 lpl 1900 2131700 2132722 m unknown 6
1038.1 6741 lpl 1901 2132987 2133136 P unknown Hypothetical protein 6
4188.1 7022 lpioiδs 213723 214016 P unknown Similar to transposase 4.5 Some similarity with eukaryotic
1020.1 6739 lpll915 2147637 2150381 P unknown 5.2 proteins Similar to C-terminal part of type I
4185.1 7021 lpl0187 214968 217403 P unknown 3.2 restπction enzyme
Weakly similar to putative
1003.1 6737 lpll926 2164000 2164812 p unknown 3.5.2 transcriptional regulator
1002.1 6736 lpl1927 2164842 2165246 m unknown 6 1001.1 6735 lpll928 2165432 2166199 p unknown 6 Some similarity with eukaryotic
997.1 7060 lpll931 2169048 2170490 p unknown 5.2 proteins
995.2 7059 lpll933 2171360 2172388 p unknown Similar to transposase 4.5 4237.1 7029 lpl1934 2172470 2172721 p unknown 6 Similar to C-terminal part of
4182.1 7020 lpl0188 217615 218598 m unknown 5.2 predicted nuclease Similar to major outer membrane 4248.1 7030 lpl1942 2179995 2180972 p unknown 1.1 protein momp Similar to capsular polysaccharide
4250.1 7031 lpl1943 2181017 2182399 P unknown 1.1 biosynthesis protein 4251.1 7032 lpl1944 2182621 2183325 P unknown Similar to transcriptional regulator 3.5.2 Similar to methionyl-tRNA 4252.1 7033 lpl1945 2183318 2184877 P unknown 3.7.2 synthetase Similar to siderophore biosynthesis
4253.1 7034 lpl1946 2184855 2186192 p unknown 2.5 protein IucD Some similarity with hypothetical 4254.1 7035 lpll947 2186176 2186526 p unknown 5.2 protein
4181.1 7019 lpl0189 218758 220989 P unknown Some similarity with SidC 5.1 4236.2 7028 lpl 1965 2207266 2208294 P unknown Similar to transposase, IS630 family 4.5
4177.1 7018 lpl0190 221031 221240 m Unknown 6 4175.1 7017 lpl0191 221304 221948 m unknown 6 4174.1 7016 lpl0192 222102 223523 P unknown Similar to transposase 4.5 Similar to predicted nuclease,
4173.1 7015 lpl0193 223610 224104 m unknown 5.2 truncated
4172.1 7014 lpl0194 224432 225391 P unknown Similar to virulence protein 5.2 4171.2 7013 lpl0195 225848 227266 P unknown Similar to transposase 4.5
Similar to transposase and
4170.1 7012 lpl0196 227299 228132 m unknown 4.5 inactivated derivatives
899.2 7058 lpl2032 2276023 2277441 P unknown Similar to transposase 4.5
3162.1 6940 lpl2033 2277621 2278655 m unknown Similar to transposase 4.5
3161.1 6939 lpl2034 2278869 2279909 m unknown Similar to threonine dehydrogenase 2.2 Similar to conserved hypothetical
3160.1 6938 lpl2035 2280074 2280520 m unknown 5.2 protein Similar to putative DNA-3-
3158.1 6937 lpl2036 2280636 2281013 m unknown 3.2 methyladenine glycosylase II
4169.1 7011 lpl0197 228170 228463 P unknown Similar to transposase 4.5
3156.1 6936 lpl2037 2282022 2283050 P unknown 6
3155.1 6935 lpl2038 2283257 2283496 m unknown Hypothetical protein 6
3151.1 6934 lpl2042 2286768 2286950 P unknown Hypothetical protein 6
2241.1 6853 lpl0199 229638 230066 P unknown Similar to hypothetical protein 5.2
3140.1 6933 lpl2049 2297022 2298080 P unknown 6
2242.1 6854 lpl0200 230056 231003 P unknown Similar to hypothetical protein 5.2
2244.1 6855 lpl0201 231474 231704 P Unknown Tral gene fragment 4.5
2245.1 6856 lpl0202 231520 231810 P Unknown Tral gene fragment 4.5
2246.1 6857 lpl0203 231882 232007 P Unknown Tral gene fragment 4.5
2247.1 6858 lpl0204 232253 232501 m unknown Hypothetical protein 6
2248.1 6859 lpl0205 232524 232730 m unknown Similar to transposase, truncated 4.5 Similar to conserved hypothetical
2253.1 6860 lpl0207 234172 235023 P unknown 5.2 protein
1105.2 6755 lpl0019 23436 24410 m unknown 6 Similar to conserved hypothetical
2259.1 6861 lplO211 237456 238250 P unknown 5.2 protein Weakly similar to transcriptional
3066.1 6932 lpl2098 2378077 2378541 m unknown 3.5.2 regulator, LuxR family
3065.1 6931 lpl2099 2378599 2378952 P unknown 6
3064.1 6930 lpl2100 2379202 2380512 P unknown 6
2260.1 6862 lpl0212 238252 239340 P unknown Similar to putative aminotransferase 2.2
contains an HPT Histidine-
3057.1 6929 lpl2105 2388174 2388623 p unknown containing phosphotransfer (HPt) 1.3 domain
3056.1 6928 lpl2106 2388928 2395878 p
2261.1 6863 lpl0213 239330 239494 p unknown Hypothetical protein 6 Similar to two component sensor
3054.1 6927 lpl2107 2395875 2397905 p unknown 1.3 histidine kinase ilarity with eukaryotic
3042.1 6926 lpl2114 2414580 2416295 p n m Some sim tllnUlVllnWw Wn 11 5.2 proteins Similar to ABC-type antimicrobial
3417.1 6942 lpl0216 242653 243363 p unknown peptide transport system, ATPase 1.2 component Similar to ABC-type antimicrobial
3420.1 6943 lpl0217 243357 245723 p + unknown peptide transport system, permease 1.2 component Similar to major outer membrane
3002.1 6925 lpl2148 2454186 2455163 P + unknown 1.1 protein
3422.1 6944 lpl0218 245723 246901 P + unknown Similar to membrane protein 1.2
3435.1 6945 lpl0226 253065 253781 P + unknown Similar to putative outer membrane Ull^ll W 11 1.1 protein Similar to C-terminal part of
2810.1 6923 lpl2284 2603747 2604793 m unknown 4.5 transposase Similar to N-terminal part of
2809.1 6922 lpl2285 2604794 2604934 m - unknown 4.5 transposase
2808.1 6921 lpl2286 2604936 2605313 m - unknown Hypothetical protein 6
2807.1 6920 lpl2287 2605322 2605540 m unknown Similar to hypothetical protein 6 ar to conserved hypothetical
2806.1 6919 lpl2288 2605737 2607245 P - liπVtiπwn Simil 5.2 protein
2805.1 6918 lpl2289 2607284 2607952 P - unknown 6
Similar to conserved hypothetical
2802.1 6917 lpl2291 2608606 2608914 P unknown 5.2 protein Similar to conserved hypothetical
2801.1 6916 lpl2292 2608911 2609849 P unknown 5.2 protein
2798.1 6915 lpl2293 2610374 2611462 m unknown Similar to transposase 4.5 Similar to predicted ATPase (AAA+
2796.1 6914 lpl2294 2611818 2613191 P unknown 5.2 superfamily)
2795.1 6913 lpl2295 2613390 2614295 P unknown Similar to restriction endonuclease 3.2 Similar to acetoacetyl-CoA
2782.1 6912 lpl2305 2625469 2626209 P unknown 2.4 reductase
2777.1 6911 lpl2308 2629983 2630168 m unknown Hypothetical protein 6
2775.1 6910 lpl2309 2630203 2631090 m unknown Similar to universal stress protein 4.1 Some similarity with eukaryotic
2749.1 6909 lpl2323 2647610 2648674 P unknown 5.2 proteins Some similarity with eukaryotic
2738.1 6908 lpl2330 2656109 2657023 P unknown 5.2 proteins
2723.1 6906 lpl2340 2669004 2669732 P + unknown 6
2722.1 6905 lpl2341 2669753 2671468 P unknown Similar to beta-lactamase precursor 4.2
2720.1 6904 lpl2342 2671598 2672392 m + unknown 6
2719.1 6902 lpl2343 2672575 2673126 P unknown 6
2717.1 6901 lpl2344 2673318 2675249 m unknown Ankyrin repeat protein 6
2708.1 6900 lpl2350 2682950 2683255 P Unknown 6
2706.1 6899 lpl2351 2683321 2683521 P unknown Hypothetical protein 6
2701.1 6898 lpl2354 2686212 2687375 m unknown 6
2696.1 6896 lpl2358 2689651 2689839 m unknown Hypothetical protein 6 Some similarity with Legionella
2660.1 6893 lpl2384 2713562 2715481 m unknown 5.1 effector protein B
2658.1 6891 lpl2385 2715575 2717098 P unknown Similar to SidD protein 5.1
2640.1 6890 lpl2399 2729218 2729607 P unknown Similar to hypothetical protein 5.2
2587.1 6889 lpl2443 2784183 2784362 P + Unknown 6
Some similarity with transcriptional
2584.1 6888 lpl2445 2785779 2786138 P unknown 3.5.2 regulators
2558.1 6887 lpl2465 2806208 2806354 m unknown Hypothetical protein 6
2547.1 6886 lpl2472 2811871 2812713 P unknown 6
2541.1 6885 lpl2476 2816937 2818298 m unknown Similar to protein kinase 3.8 Some similarity with eukaryotic
2540.1 6884 lpl2477 2818401 2819000 m unknown proteins
2539.1 6883 lpl2478 2819044 2819529 m unknown Similar to transposase 4.5
2534.1 6882 lpl2483 2824095 2824961 P unknown 6
2533.1 6881 lpl2484 2824980 2825720 P unknown Similar to heat shock protein DnaJ 4.1
2532.1 6880 lpl2485 2825862 2826545 P unknown 6
2531.1 6879 lpl2486 2826766 2828007 P unknown Similar to phage-related integrase 4.4
2530.1 6878 lpl2487 2828021 2829058 P unknown Similar to putative virulence proteins 5.2
2529.1 6877 lpl2488 2829356 2832133 m unknown Similar to helicase 3.7.4 Similar to conserved hypothetical
2527.1 6876 lpl2489 2832440 2833039 P unknown 5.2 protein Similar to conserved hypothetical
2526.1 6875 lpl2490 2833042 2834082 P unknown 5.2 protein
2525.1 6874 lpl2491 2834127 2834498 P unknown 6
2524.1 6873 lpl2492 2835267 2835971 P unknown Similar to Legionella LvrA protein 5.1 Similar to conserved hypothetical
2523.1 6872 lpl2493 2836365 2839757 P unknown 5.2 protein
2521.1 6871 lpl2494 2839769 2846089 P unknown Similar to other protein 5.2
2518.1 6870 lpl2495 2846774 2847583 P unknown 6
2517.1 6869 lpl2496 2847573 2848469 P unknown Similar to integrase 4.5
2516.1 6868 lpl2497 2848638 2849378 m unknown Similar to hypothetical protein 5.2
2441.1 6867 lpl2545 2905983 2906834 m + unknown 6
2440.1 6866 lpl2546 2906910 2908442 m unknown 6
2422.1 6865 lpl2558 2921462 2921779 m unknown Similar to hypothetical protein 5.2
2392.1 6864 lpl2580 2948197 2948331 m unknown Hypothetical protein 6
1434.1 6793 lpl2732 3125135 3125284 m unknown hypothetical gene 6
1422.1 6792 lpl2741 3134300 3136033 m unknown 6
1321.1 6791 lpl2806 3208466 3210055 m unknown 6
3536.1 6946 lpl0286 321867 322067 m unknown Hypothetical protein 6
3537.1 6947 lpl0287 322127. 322402 m unknown Hypothetical protein 6 Similar to regulatory protein
1297.1 6790 lpl2826 3226572 3227516 P unknown 1.3 (GGDEF domains)
1296.1 6789 lpl2827 3227525 3227884 m Unknown 6 Similar to putative transcriptional
1285.1 6788 lpl2833 3233251 3233541 m unknown 3.5.2 regulator
1284.1 6787 lpl2834 3233591 3233854 m unknown Similar to hypothetical protein 5.2
1283.1 6786 lpl2835 3234032 3234310 P unknown Putative transposase 4.5
1282.1 6785 lpl2836 3234439 3234627 P unknown Similar to transposase, truncated 4.5
1280.1 6784 lpl2837 3234734 3235702 P unknown Similar to unknown protein 5.2
1279.1 6783 lpl2838 3235699 3239094 P unknown Similar to predicted helicase 3.7.4
Similar to conserved hypothetical
1278.1 6782 lpl2839 3239087 3240352 P unknown 5.2 protein Similar to conserved hypothetical
1276.1 6781 lpl2840 3240349 3241290 p unknown 5.2 protein Similar to conserved hypothetical
1275.1 6780 lpl2841 3241306 3242331 p unknown 5.2 protein Similar to conserved hypothetical
1274.1 6779 lpl2842 3242341 3242895 p unknown 5.2 protein
1258.1 6776 lpl2843 3247702 3248046 m unknown Similar to transposase 4.5 Similar to conserved hypothetical
1255.2 6775 lpl2845 3248864 3249910 m unknown 5.2 protein Similar to conserved hypothetical
1252.1 6774 lpl2847 3251745 3252164 p unknown 5.2 protein
1251.1 6773 lpl2848 3252157 3252483 p unknown Similar to hypothetical protein 5.2
1237.2 6770 lpl2857 3261124 3263307 m unknown 6 1221.1 6769 lpl2867 3272087 3272719 p unknown Similar to hypothetical protein 5.2 1219.1 6768 lpl2868 3273424 3274155 p unknown Similar to transposase 4.5
+ + +
Some similarity with eukaryotic
1050.1 6750 lpl0063 68898 70691 unknown 6 protein
1049.2 6748 lpl0064 70860 72278 P - unknown Similar to transposase 4.5
272.1 6903 lpl0065 72587 73840 P - unknown Similar to phage-related integrase 4.4
270.1 6897 lpl0066 74228 74809 P - unknown Similar to hypothetical protein 5.2
269.1 6895 lpl0067 74915 75700 P - unknown Similar to hypothetical protein 5.2
267.1 6894 lpl0068 75697 76572 P - unknown Similar to hypothetical protein 5.2
266.1 6892 lpl0069 77003 77278 P - unknown 6
1697.1 6800 lpl0718 806040 806339 P - Unknown 6
1718.1 6801 lpl0729 820733 821518 P + unknown 6
1824.1 6803 lpl0801 907894 908922 P - unknown Similar to transposase 4.5
1825.1 6804 lpl0802 908919 909056 m - unknown Similar to transposase, truncated 4.5
1826.1 6805 lpl0803 909248 910531 P + unknown Similar to hypothetical protein 5.1 Similar to molybdopterin cofactor
1827.1 6806 lpl0804 910565 911593 P - unknown ..1.1 synthesis protein A
1828.1 6807 lpl0805 912012 912104 P - Unknown Similar to transposase, truncated 4.5
1829.1 6808 lpl0806 912186 912323 P - unknown Similar to transposase, truncated 4.5
1834.1 6809 lpl0811 916637 918157 P - unknown Similar to hypothetical protein 5.1
3859.1 6975 plplOOl l 11934 12119 P - Unknown Unknown 6 Weakly similar to carbon storage
3861.1 6976 plpl0012 12176 12334 P - unknown 6 regulator Similar to E.coli TraT complement
3862.1 6977 plpl0013 12468 13214 P + unknown 4.5 resistance protein
3863.1 6978 plpl0014 13334 13621 P + unknown 6 Some similarity with eukaryotic
3850.1 6974 plpl0002 1350 2747 P - unknown 6 proteins Similar to putative conjugative
3865.1 6979 plpl0015 13634 13930 P - unknown 4.5 transfer protein TraL Similar to putative conjugative
3866.1 6980 plplOOlό 13940 14509 P + unknown 4.5 transfer protein TraE
m in in in in m in m in m in m in rf in VO • t f rf rf f rf rr' r rf rf rf rf rf* rf
+ + + + + + + + ft
OO rf σs m Is- O VO CO σs ro CN 00 OO r- in 00 —H σs CN rf rf CO VO rf CN m 00 rf Is- rj- CO 00 m CN r- σs ro CN m vo vo σs σs © CO rf rf VO 00 ro in rf σs CN CN CN CN CN CN CN CN CN in CN © CN Is- r- σs ro CO σs m rf © © VO m 00 © rf © rf rf CN m ro in CN VD 00 rf Is- rf CO 00 m CN r- r- σs VO rf m VO vo σs σs O ro rf rf VO CN 00 σs CN CN CN CN CN CN CN CN CN 00 σs o CN ro rf m vo r- 00 σs © CN CN CN CN CN CN CN N CN CN © ro ro O o © © © © © © © © © © © © © © o o © O O © © © o © © O O o © © "ft "ft "ft "ft "ft "ft "ft "ft "ft "ft "ft "ft Έ "ft "ft "ft "ft "ft "ft "ft "ft "ft "ft "ft "ft "ft CN ro rf m vo r- 00 σs © CN ro ro rf m 00 0O 00 00 00 00 00 00 00 σs σs σs σs r- σs σs σs σs σs σs σs os σs σs σs σs σs σs σs σs σs σs vo vo vo vo vo vo vo vo vo vo vo vo vo vo vo vo CN 00 © ro rf m r- 00 © CN rf VO 00 00 © vo r- r- r- r- r- t-- r- 00 00 00 00 00 rf 00 O 00 00 00 00 00 00 00 00 00 00 00 00 00 00 00 00 ro ro ro ro ro ro CO ro ro ro ro ro ro ro ro ro
3891.2 6996 plpl0032 31463 34411 p unknown 4.5 protein Tral Some similarity with eukaryotic
3893.1 6997 plpl0033 34428 34751 p unknown 6 proteins
Similar to conserved hypothetical
3894.1 6998 plpl0034 35323 35727 m unknown 5.2 protein Similar to conserved hypothetical
3895.1 6999 plpl0035 35724 35921 m unknown 5.2 protein
3896.1 7000 plpl0036 36007 36225 m Unknown 6
3815.2 6957 plpl0037 36793 37656 m unknown Similar to antirestriction protein 3.2
3816.1 6958 plpl0038 37966 38127 m unknown Hypothetical protein 6
3818.1 6959 plpl0039 38700 39233 m unknown 6
3820.1 6960 plpl0040 39807 40790 P + unknown Similar to hypothetical protein 5.2 Similar to conserved hypothetical
3822.1 6961 plpl0042 41534 41965 P unknown 5.2 protein
3823.1 6962 plpl0043 42194 42760 P unknown Similar to site-specific recombinases 3.3
3827.1 6963 plpl0044 45122 45241 m unknown Hypothetical protein 6
3828.1 6964 plpl0045 45347 45472 m unknown Hypothetical protein 6
3831.1 6965 plpl0046 46027 46149 P unknown Hypothetical protein 5.2 Similar to conserved hypothetical
3835.1 6966 plpl0047 46715 47269 m unknown 5.2 protein Similar to conserved hypothetical
3836.1 6967 plpl0048 47280 48305 m unknown 5.2 protein
3837.1 6968 plpl0049 48321 49262 m + unknown Similar to hypothetical protein 5.2 Similar to conserved hypothetical
3839.1 6969 plpl0050 49259 50524 m unknown 5.2 protein
3840.1 6970 plpl0051 50517 53912 m unknown Similar to helicases 3.3 Similar to uncharacterized protein
3841.1 6971 plpl0052 53909 54877 m unknown predicted to be involved in DNA 3.2 repair 3843.1 6972 plpl0053 55064 55336 m unknown Similar to transposase 4.5