US20090082577A1 - Fluorescent probes for biological studies - Google Patents
Fluorescent probes for biological studies Download PDFInfo
- Publication number
- US20090082577A1 US20090082577A1 US12/235,454 US23545408A US2009082577A1 US 20090082577 A1 US20090082577 A1 US 20090082577A1 US 23545408 A US23545408 A US 23545408A US 2009082577 A1 US2009082577 A1 US 2009082577A1
- Authority
- US
- United States
- Prior art keywords
- fluorescence
- gst
- compound
- crk
- abl
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Abandoned
Links
- 239000007850 fluorescent dye Substances 0.000 title description 3
- 150000001875 compounds Chemical class 0.000 claims abstract description 93
- IJGRMHOSHXDMSA-UHFFFAOYSA-N nitrogen Substances N#N IJGRMHOSHXDMSA-UHFFFAOYSA-N 0.000 claims description 28
- 229910052739 hydrogen Inorganic materials 0.000 claims description 21
- 239000001257 hydrogen Substances 0.000 claims description 21
- 229910052757 nitrogen Inorganic materials 0.000 claims description 20
- 125000000217 alkyl group Chemical group 0.000 claims description 17
- 125000004435 hydrogen atom Chemical group [H]* 0.000 claims description 12
- 229910052736 halogen Inorganic materials 0.000 claims description 11
- 150000002367 halogens Chemical group 0.000 claims description 11
- QJGQUHMNIGDVPM-UHFFFAOYSA-N nitrogen group Chemical group [N] QJGQUHMNIGDVPM-UHFFFAOYSA-N 0.000 claims description 7
- 125000003545 alkoxy group Chemical group 0.000 claims description 6
- 229910052731 fluorine Inorganic materials 0.000 claims description 6
- PXGOKWXKJXAPGV-UHFFFAOYSA-N Fluorine Chemical group FF PXGOKWXKJXAPGV-UHFFFAOYSA-N 0.000 claims description 5
- 239000011737 fluorine Chemical group 0.000 claims description 5
- 150000002431 hydrogen Chemical group 0.000 claims 4
- 125000002887 hydroxy group Chemical group [H]O* 0.000 claims 1
- 238000000034 method Methods 0.000 abstract description 30
- 238000012544 monitoring process Methods 0.000 abstract description 7
- 230000004850 protein–protein interaction Effects 0.000 abstract description 6
- 108090000765 processed proteins & peptides Proteins 0.000 description 59
- OKKJLVBELUTLKV-UHFFFAOYSA-N Methanol Chemical compound OC OKKJLVBELUTLKV-UHFFFAOYSA-N 0.000 description 45
- 108090000623 proteins and genes Proteins 0.000 description 37
- XEKOWRVHYACXOJ-UHFFFAOYSA-N Ethyl acetate Chemical compound CCOC(C)=O XEKOWRVHYACXOJ-UHFFFAOYSA-N 0.000 description 36
- 102000004169 proteins and genes Human genes 0.000 description 35
- WEVYAHXRMPXWCK-UHFFFAOYSA-N Acetonitrile Chemical compound CC#N WEVYAHXRMPXWCK-UHFFFAOYSA-N 0.000 description 33
- HEDRZPFGACZZDS-MICDWDOJSA-N Trichloro(2H)methane Chemical class [2H]C(Cl)(Cl)Cl HEDRZPFGACZZDS-MICDWDOJSA-N 0.000 description 31
- 102000014400 SH2 domains Human genes 0.000 description 26
- 108050003452 SH2 domains Proteins 0.000 description 26
- 239000002904 solvent Substances 0.000 description 26
- -1 (cyclohexyl)methyl Chemical group 0.000 description 25
- YMWUJEATGCHHMB-UHFFFAOYSA-N Dichloromethane Chemical compound ClCCl YMWUJEATGCHHMB-UHFFFAOYSA-N 0.000 description 23
- 239000000203 mixture Substances 0.000 description 23
- VLKZOEOYAKHREP-UHFFFAOYSA-N n-Hexane Chemical class CCCCCC VLKZOEOYAKHREP-UHFFFAOYSA-N 0.000 description 22
- 238000006862 quantum yield reaction Methods 0.000 description 21
- 102000004196 processed proteins & peptides Human genes 0.000 description 19
- 239000000523 sample Substances 0.000 description 19
- 235000019439 ethyl acetate Nutrition 0.000 description 18
- 0 [4*]C1=C2C(=O)N(C[C@H](C)C(=O)O)C(=O)C2=C([9*])C2=C([8*])C([7*])=C([6*])C([5*])=C12 Chemical compound [4*]C1=C2C(=O)N(C[C@H](C)C(=O)O)C(=O)C2=C([9*])C2=C([8*])C([7*])=C([6*])C([5*])=C12 0.000 description 16
- 239000000243 solution Substances 0.000 description 16
- YXFVVABEGXRONW-UHFFFAOYSA-N Toluene Chemical compound CC1=CC=CC=C1 YXFVVABEGXRONW-UHFFFAOYSA-N 0.000 description 15
- 238000002835 absorbance Methods 0.000 description 15
- 238000006243 chemical reaction Methods 0.000 description 14
- 230000005284 excitation Effects 0.000 description 14
- 238000004448 titration Methods 0.000 description 14
- 125000000623 heterocyclic group Chemical group 0.000 description 13
- 108020005497 Nuclear hormone receptor Proteins 0.000 description 12
- 102000007399 Nuclear hormone receptor Human genes 0.000 description 12
- 125000001072 heteroaryl group Chemical group 0.000 description 12
- 230000003993 interaction Effects 0.000 description 12
- 102000005962 receptors Human genes 0.000 description 12
- 108020003175 receptors Proteins 0.000 description 12
- 230000005281 excited state Effects 0.000 description 11
- 125000006239 protecting group Chemical group 0.000 description 11
- 229920006395 saturated elastomer Polymers 0.000 description 11
- XLYOFNOQVPJJNP-UHFFFAOYSA-N water Substances O XLYOFNOQVPJJNP-UHFFFAOYSA-N 0.000 description 11
- 150000001413 amino acids Chemical group 0.000 description 10
- 239000000872 buffer Substances 0.000 description 10
- 125000001424 substituent group Chemical group 0.000 description 10
- 238000005160 1H NMR spectroscopy Methods 0.000 description 9
- LFQSCWFLJHTTHZ-UHFFFAOYSA-N Ethanol Chemical compound CCO LFQSCWFLJHTTHZ-UHFFFAOYSA-N 0.000 description 9
- PEDCQBHIVMGVHV-UHFFFAOYSA-N Glycerine Chemical compound OCC(O)CO PEDCQBHIVMGVHV-UHFFFAOYSA-N 0.000 description 9
- 150000008064 anhydrides Chemical class 0.000 description 9
- 125000003118 aryl group Chemical group 0.000 description 9
- 230000009141 biological interaction Effects 0.000 description 9
- 125000004209 (C1-C8) alkyl group Chemical group 0.000 description 8
- JGFZNNIVVJXRND-UHFFFAOYSA-N N,N-Diisopropylethylamine (DIPEA) Chemical compound CCN(C(C)C)C(C)C JGFZNNIVVJXRND-UHFFFAOYSA-N 0.000 description 8
- 125000004432 carbon atom Chemical group C* 0.000 description 8
- 230000008859 change Effects 0.000 description 8
- 125000005842 heteroatom Chemical group 0.000 description 8
- 108020004707 nucleic acids Proteins 0.000 description 8
- 150000007523 nucleic acids Chemical class 0.000 description 8
- 102000039446 nucleic acids Human genes 0.000 description 8
- 125000003088 (fluoren-9-ylmethoxy)carbonyl group Chemical group 0.000 description 7
- 238000001644 13C nuclear magnetic resonance spectroscopy Methods 0.000 description 7
- LRAMQBKQEYONOM-UHFFFAOYSA-N 9h-fluoren-9-ylmethyl n-(3-aminopropyl)carbamate Chemical compound C1=CC=C2C(COC(=O)NCCCN)C3=CC=CC=C3C2=C1 LRAMQBKQEYONOM-UHFFFAOYSA-N 0.000 description 7
- KFZMGEQAYNKOFK-UHFFFAOYSA-N Isopropanol Chemical compound CC(C)O KFZMGEQAYNKOFK-UHFFFAOYSA-N 0.000 description 7
- 239000007832 Na2SO4 Substances 0.000 description 7
- VYPSYNLAJGMNEJ-UHFFFAOYSA-N Silicium dioxide Chemical compound O=[Si]=O VYPSYNLAJGMNEJ-UHFFFAOYSA-N 0.000 description 7
- PMZURENOXWZQFD-UHFFFAOYSA-L Sodium Sulfate Chemical compound [Na+].[Na+].[O-]S([O-])(=O)=O PMZURENOXWZQFD-UHFFFAOYSA-L 0.000 description 7
- UIIMBOGNXHQVGW-UHFFFAOYSA-M Sodium bicarbonate Chemical class [Na+].OC([O-])=O UIIMBOGNXHQVGW-UHFFFAOYSA-M 0.000 description 7
- 238000010790 dilution Methods 0.000 description 7
- 239000012895 dilution Substances 0.000 description 7
- 230000005283 ground state Effects 0.000 description 7
- 239000012044 organic layer Substances 0.000 description 7
- 239000011541 reaction mixture Substances 0.000 description 7
- 229910052702 rhenium Inorganic materials 0.000 description 7
- 229910052938 sodium sulfate Inorganic materials 0.000 description 7
- 239000007787 solid Substances 0.000 description 7
- 102000004190 Enzymes Human genes 0.000 description 6
- 108090000790 Enzymes Proteins 0.000 description 6
- NQRYJNQNLNOLGT-UHFFFAOYSA-N Piperidine Chemical compound C1CCNCC1 NQRYJNQNLNOLGT-UHFFFAOYSA-N 0.000 description 6
- FAPWRFPIFSIZLT-UHFFFAOYSA-M Sodium chloride Chemical compound [Na+].[Cl-] FAPWRFPIFSIZLT-UHFFFAOYSA-M 0.000 description 6
- WYURNTSHIVDZCO-UHFFFAOYSA-N Tetrahydrofuran Chemical compound C1CCOC1 WYURNTSHIVDZCO-UHFFFAOYSA-N 0.000 description 6
- 238000010521 absorption reaction Methods 0.000 description 6
- 229940088598 enzyme Drugs 0.000 description 6
- 238000002189 fluorescence spectrum Methods 0.000 description 6
- NPZTUJOABDZTLV-UHFFFAOYSA-N hydroxybenzotriazole Substances O=C1C=CC=C2NNN=C12 NPZTUJOABDZTLV-UHFFFAOYSA-N 0.000 description 6
- 230000009021 linear effect Effects 0.000 description 6
- 125000004433 nitrogen atom Chemical group N* 0.000 description 6
- 230000008569 process Effects 0.000 description 6
- 230000019491 signal transduction Effects 0.000 description 6
- RYHBNJHYFVUHQT-UHFFFAOYSA-N 1,4-Dioxane Chemical group C1COCCO1 RYHBNJHYFVUHQT-UHFFFAOYSA-N 0.000 description 5
- 239000001576 FEMA 2977 Substances 0.000 description 5
- UFHFLCQGNIYNRP-UHFFFAOYSA-N Hydrogen Chemical compound [H][H] UFHFLCQGNIYNRP-UHFFFAOYSA-N 0.000 description 5
- 239000006180 TBST buffer Substances 0.000 description 5
- 125000003342 alkenyl group Chemical group 0.000 description 5
- 125000000304 alkynyl group Chemical group 0.000 description 5
- 239000012472 biological sample Substances 0.000 description 5
- 230000015572 biosynthetic process Effects 0.000 description 5
- 125000003917 carbamoyl group Chemical group [H]N([H])C(*)=O 0.000 description 5
- 210000004027 cell Anatomy 0.000 description 5
- 239000003153 chemical reaction reagent Substances 0.000 description 5
- LOUPRKONTZGTKE-UHFFFAOYSA-N cinchonine Natural products C1C(C(C2)C=C)CCN2C1C(O)C1=CC=NC2=CC=C(OC)C=C21 LOUPRKONTZGTKE-UHFFFAOYSA-N 0.000 description 5
- 125000004122 cyclic group Chemical group 0.000 description 5
- 238000003818 flash chromatography Methods 0.000 description 5
- 102000037865 fusion proteins Human genes 0.000 description 5
- 108020001507 fusion proteins Proteins 0.000 description 5
- RWSXRVCMGQZWBV-WDSKDSINSA-N glutathione Chemical compound OC(=O)[C@@H](N)CCC(=O)N[C@@H](CS)C(=O)NCC(O)=O RWSXRVCMGQZWBV-WDSKDSINSA-N 0.000 description 5
- 230000007246 mechanism Effects 0.000 description 5
- 239000012528 membrane Substances 0.000 description 5
- 125000000449 nitro group Chemical group [O-][N+](*)=O 0.000 description 5
- 229910052760 oxygen Inorganic materials 0.000 description 5
- 238000010647 peptide synthesis reaction Methods 0.000 description 5
- 229960003110 quinine sulfate Drugs 0.000 description 5
- AKYHKWQPZHDOBW-UHFFFAOYSA-N (5-ethenyl-1-azabicyclo[2.2.2]octan-7-yl)-(6-methoxyquinolin-4-yl)methanol Chemical compound OS(O)(=O)=O.C1C(C(C2)C=C)CCN2C1C(O)C1=CC=NC2=CC=C(OC)C=C21 AKYHKWQPZHDOBW-UHFFFAOYSA-N 0.000 description 4
- 125000003903 2-propenyl group Chemical group [H]C([*])([H])C([H])=C([H])[H] 0.000 description 4
- ZZVAOWRXBFJIJR-UHFFFAOYSA-N 6-(dimethylamino)benzo[f][2]benzofuran-1,3-dione Chemical compound C1=C2C(=O)OC(=O)C2=CC2=CC(N(C)C)=CC=C21 ZZVAOWRXBFJIJR-UHFFFAOYSA-N 0.000 description 4
- 241000894006 Bacteria Species 0.000 description 4
- HEDRZPFGACZZDS-UHFFFAOYSA-N Chloroform Chemical compound ClC(Cl)Cl HEDRZPFGACZZDS-UHFFFAOYSA-N 0.000 description 4
- 108010024636 Glutathione Proteins 0.000 description 4
- QAOWNCQODCNURD-UHFFFAOYSA-N Sulfuric acid Chemical compound OS(O)(=O)=O QAOWNCQODCNURD-UHFFFAOYSA-N 0.000 description 4
- QTBSBXVTEAMEQO-UHFFFAOYSA-N acetic acid Substances CC(O)=O QTBSBXVTEAMEQO-UHFFFAOYSA-N 0.000 description 4
- 150000001412 amines Chemical class 0.000 description 4
- QVGXLLKOCUKJST-UHFFFAOYSA-N atomic oxygen Chemical compound [O] QVGXLLKOCUKJST-UHFFFAOYSA-N 0.000 description 4
- 125000003178 carboxy group Chemical group [H]OC(*)=O 0.000 description 4
- 150000004985 diamines Chemical class 0.000 description 4
- 239000000975 dye Substances 0.000 description 4
- 150000002500 ions Chemical class 0.000 description 4
- 125000002950 monocyclic group Chemical group 0.000 description 4
- 239000001301 oxygen Substances 0.000 description 4
- 125000004430 oxygen atom Chemical group O* 0.000 description 4
- DCWXELXMIBXGTH-UHFFFAOYSA-N phosphotyrosine Chemical compound OC(=O)C(N)CC1=CC=C(OP(O)(O)=O)C=C1 DCWXELXMIBXGTH-UHFFFAOYSA-N 0.000 description 4
- 239000000047 product Substances 0.000 description 4
- 239000012460 protein solution Substances 0.000 description 4
- 239000003586 protic polar solvent Substances 0.000 description 4
- 230000035945 sensitivity Effects 0.000 description 4
- 238000001228 spectrum Methods 0.000 description 4
- 238000003756 stirring Methods 0.000 description 4
- 239000011550 stock solution Substances 0.000 description 4
- 239000000126 substance Substances 0.000 description 4
- 229910052717 sulfur Inorganic materials 0.000 description 4
- 125000004434 sulfur atom Chemical group 0.000 description 4
- 238000003786 synthesis reaction Methods 0.000 description 4
- 238000012360 testing method Methods 0.000 description 4
- 238000012546 transfer Methods 0.000 description 4
- 230000032258 transport Effects 0.000 description 4
- 150000000094 1,4-dioxanes Chemical class 0.000 description 3
- MPPQGYCZBNURDG-UHFFFAOYSA-N 2-propionyl-6-dimethylaminonaphthalene Chemical compound C1=C(N(C)C)C=CC2=CC(C(=O)CC)=CC=C21 MPPQGYCZBNURDG-UHFFFAOYSA-N 0.000 description 3
- CSCPPACGZOOCGX-UHFFFAOYSA-N Acetone Chemical compound CC(C)=O CSCPPACGZOOCGX-UHFFFAOYSA-N 0.000 description 3
- 125000004648 C2-C8 alkenyl group Chemical group 0.000 description 3
- 125000004649 C2-C8 alkynyl group Chemical group 0.000 description 3
- LNLZDKYLUVIXEF-UHFFFAOYSA-N COC(=O)CN1C(=O)C2=CC3=CC=C(N(C)C)C=C3C=C2C1=O Chemical compound COC(=O)CN1C(=O)C2=CC3=CC=C(N(C)C)C=C3C=C2C1=O LNLZDKYLUVIXEF-UHFFFAOYSA-N 0.000 description 3
- GDBQQVLCIARPGH-UHFFFAOYSA-N Leupeptin Natural products CC(C)CC(NC(C)=O)C(=O)NC(CC(C)C)C(=O)NC(C=O)CCCN=C(N)N GDBQQVLCIARPGH-UHFFFAOYSA-N 0.000 description 3
- 229910003844 NSO2 Inorganic materials 0.000 description 3
- RWRDLPDLKQPQOW-UHFFFAOYSA-N Pyrrolidine Chemical compound C1CCNC1 RWRDLPDLKQPQOW-UHFFFAOYSA-N 0.000 description 3
- NINIDFKCEFEMDL-UHFFFAOYSA-N Sulfur Chemical compound [S] NINIDFKCEFEMDL-UHFFFAOYSA-N 0.000 description 3
- 238000000862 absorption spectrum Methods 0.000 description 3
- MGSKVZWGBWPBTF-UHFFFAOYSA-N aebsf Chemical compound NCCC1=CC=C(S(F)(=O)=O)C=C1 MGSKVZWGBWPBTF-UHFFFAOYSA-N 0.000 description 3
- 239000007864 aqueous solution Substances 0.000 description 3
- 125000001584 benzyloxycarbonyl group Chemical group C(=O)(OCC1=CC=CC=C1)* 0.000 description 3
- 125000002619 bicyclic group Chemical group 0.000 description 3
- 239000012148 binding buffer Substances 0.000 description 3
- 230000033077 cellular process Effects 0.000 description 3
- 230000001419 dependent effect Effects 0.000 description 3
- 238000004128 high performance liquid chromatography Methods 0.000 description 3
- 150000002430 hydrocarbons Chemical group 0.000 description 3
- 125000003453 indazolyl group Chemical group N1N=C(C2=C1C=CC=C2)* 0.000 description 3
- GDBQQVLCIARPGH-ULQDDVLXSA-N leupeptin Chemical compound CC(C)C[C@H](NC(C)=O)C(=O)N[C@@H](CC(C)C)C(=O)N[C@H](C=O)CCCN=C(N)N GDBQQVLCIARPGH-ULQDDVLXSA-N 0.000 description 3
- 108010052968 leupeptin Proteins 0.000 description 3
- 150000002632 lipids Chemical class 0.000 description 3
- 238000005259 measurement Methods 0.000 description 3
- 239000003921 oil Substances 0.000 description 3
- 239000012071 phase Substances 0.000 description 3
- 108020001580 protein domains Proteins 0.000 description 3
- 238000000746 purification Methods 0.000 description 3
- 238000010791 quenching Methods 0.000 description 3
- 230000000171 quenching effect Effects 0.000 description 3
- 238000011160 research Methods 0.000 description 3
- 229910000030 sodium bicarbonate Inorganic materials 0.000 description 3
- 239000011780 sodium chloride Substances 0.000 description 3
- 125000000547 substituted alkyl group Chemical group 0.000 description 3
- 239000011593 sulfur Substances 0.000 description 3
- YLQBMQCUIZJEEH-UHFFFAOYSA-N tetrahydrofuran Natural products C=1C=COC=1 YLQBMQCUIZJEEH-UHFFFAOYSA-N 0.000 description 3
- 238000004809 thin layer chromatography Methods 0.000 description 3
- 239000003643 water by type Substances 0.000 description 3
- NHJVRSWLHSJWIN-UHFFFAOYSA-N 2,4,6-trinitrobenzenesulfonic acid Chemical compound OS(=O)(=O)C1=C([N+]([O-])=O)C=C([N+]([O-])=O)C=C1[N+]([O-])=O NHJVRSWLHSJWIN-UHFFFAOYSA-N 0.000 description 2
- YOGPCAYIEHLIMG-UHFFFAOYSA-N 2-(3-nitrophenyl)-1,3-dioxolane Chemical compound [O-][N+](=O)C1=CC=CC(C2OCCO2)=C1 YOGPCAYIEHLIMG-UHFFFAOYSA-N 0.000 description 2
- LXFWRZZEYKVCIX-UHFFFAOYSA-N 2-[2-(benzenesulfonylmethyl)-5-nitrophenyl]-1,3-dioxolane Chemical compound O1CCOC1C1=CC([N+](=O)[O-])=CC=C1CS(=O)(=O)C1=CC=CC=C1 LXFWRZZEYKVCIX-UHFFFAOYSA-N 0.000 description 2
- QKNYBSVHEMOAJP-UHFFFAOYSA-N 2-amino-2-(hydroxymethyl)propane-1,3-diol;hydron;chloride Chemical compound Cl.OCC(N)(CO)CO QKNYBSVHEMOAJP-UHFFFAOYSA-N 0.000 description 2
- PUXKVTSJYZJFRZ-UHFFFAOYSA-N 5-(dimethylamino)isoindole-1,3-dione Chemical compound CN(C)C1=CC=C2C(=O)NC(=O)C2=C1 PUXKVTSJYZJFRZ-UHFFFAOYSA-N 0.000 description 2
- XKRFYHLGVUSROY-UHFFFAOYSA-N Argon Chemical compound [Ar] XKRFYHLGVUSROY-UHFFFAOYSA-N 0.000 description 2
- 125000001313 C5-C10 heteroaryl group Chemical group 0.000 description 2
- 125000000041 C6-C10 aryl group Chemical group 0.000 description 2
- OKTJSMMVPCPJKN-UHFFFAOYSA-N Carbon Chemical compound [C] OKTJSMMVPCPJKN-UHFFFAOYSA-N 0.000 description 2
- IAZDPXIOMUYVGZ-UHFFFAOYSA-N Dimethylsulphoxide Chemical compound CS(C)=O IAZDPXIOMUYVGZ-UHFFFAOYSA-N 0.000 description 2
- KCXVZYZYPLLWCC-UHFFFAOYSA-N EDTA Chemical compound OC(=O)CN(CC(O)=O)CCN(CC(O)=O)CC(O)=O KCXVZYZYPLLWCC-UHFFFAOYSA-N 0.000 description 2
- WSFSSNUMVMOOMR-UHFFFAOYSA-N Formaldehyde Chemical compound O=C WSFSSNUMVMOOMR-UHFFFAOYSA-N 0.000 description 2
- 102000004882 Lipase Human genes 0.000 description 2
- 108090001060 Lipase Proteins 0.000 description 2
- 239000004367 Lipase Substances 0.000 description 2
- YNAVUWVOSKDBBP-UHFFFAOYSA-N Morpholine Chemical compound C1COCCN1 YNAVUWVOSKDBBP-UHFFFAOYSA-N 0.000 description 2
- 102000016943 Muramidase Human genes 0.000 description 2
- 108010014251 Muramidase Proteins 0.000 description 2
- 108010062010 N-Acetylmuramoyl-L-alanine Amidase Proteins 0.000 description 2
- 238000005481 NMR spectroscopy Methods 0.000 description 2
- 206010028980 Neoplasm Diseases 0.000 description 2
- 108091034117 Oligonucleotide Proteins 0.000 description 2
- 102000038030 PI3Ks Human genes 0.000 description 2
- 108091007960 PI3Ks Proteins 0.000 description 2
- KDLHZDBZIXYQEI-UHFFFAOYSA-N Palladium Chemical compound [Pd] KDLHZDBZIXYQEI-UHFFFAOYSA-N 0.000 description 2
- 108091000080 Phosphotransferase Proteins 0.000 description 2
- GLUUGHFHXGJENI-UHFFFAOYSA-N Piperazine Chemical compound C1CNCCN1 GLUUGHFHXGJENI-UHFFFAOYSA-N 0.000 description 2
- 102000001253 Protein Kinase Human genes 0.000 description 2
- 229910006074 SO2NH2 Inorganic materials 0.000 description 2
- 229920004890 Triton X-100 Polymers 0.000 description 2
- 239000013504 Triton X-100 Substances 0.000 description 2
- 238000011481 absorbance measurement Methods 0.000 description 2
- 125000001931 aliphatic group Chemical group 0.000 description 2
- 125000003710 aryl alkyl group Chemical group 0.000 description 2
- 125000004429 atom Chemical group 0.000 description 2
- 239000011324 bead Substances 0.000 description 2
- 125000003785 benzimidazolyl group Chemical group N1=C(NC2=C1C=CC=C2)* 0.000 description 2
- 125000001164 benzothiazolyl group Chemical group S1C(=NC2=C1C=CC=C2)* 0.000 description 2
- 125000003354 benzotriazolyl group Chemical group N1N=NC2=C1C=CC=C2* 0.000 description 2
- GDTBXPJZTBHREO-UHFFFAOYSA-N bromine Chemical compound BrBr GDTBXPJZTBHREO-UHFFFAOYSA-N 0.000 description 2
- FJDQFPXHSGXQBY-UHFFFAOYSA-L caesium carbonate Chemical compound [Cs+].[Cs+].[O-]C([O-])=O FJDQFPXHSGXQBY-UHFFFAOYSA-L 0.000 description 2
- 125000000837 carbohydrate group Chemical group 0.000 description 2
- 229910052799 carbon Inorganic materials 0.000 description 2
- 239000011203 carbon fibre reinforced carbon Substances 0.000 description 2
- 125000002915 carbonyl group Chemical group [*:2]C([*:1])=O 0.000 description 2
- 238000005119 centrifugation Methods 0.000 description 2
- 239000005515 coenzyme Substances 0.000 description 2
- 229940126214 compound 3 Drugs 0.000 description 2
- 238000010511 deprotection reaction Methods 0.000 description 2
- UGDQYRNNGLTGMU-UHFFFAOYSA-N diethyl 6-(dimethylamino)naphthalene-2,3-dicarboxylate Chemical compound C1=C(N(C)C)C=C2C=C(C(=O)OCC)C(C(=O)OCC)=CC2=C1 UGDQYRNNGLTGMU-UHFFFAOYSA-N 0.000 description 2
- IOJXFSPZXAMVHW-UHFFFAOYSA-N diethyl 6-nitronaphthalene-2,3-dicarboxylate Chemical compound C1=C([N+]([O-])=O)C=C2C=C(C(=O)OCC)C(C(=O)OCC)=CC2=C1 IOJXFSPZXAMVHW-UHFFFAOYSA-N 0.000 description 2
- ZUOUZKKEUPVFJK-UHFFFAOYSA-N diphenyl Chemical group C1=CC=CC=C1C1=CC=CC=C1 ZUOUZKKEUPVFJK-UHFFFAOYSA-N 0.000 description 2
- 201000010099 disease Diseases 0.000 description 2
- 208000037265 diseases, disorders, signs and symptoms Diseases 0.000 description 2
- 239000012153 distilled water Substances 0.000 description 2
- 230000000694 effects Effects 0.000 description 2
- 238000002330 electrospray ionisation mass spectrometry Methods 0.000 description 2
- 239000012149 elution buffer Substances 0.000 description 2
- 125000001495 ethyl group Chemical group [H]C([H])([H])C([H])([H])* 0.000 description 2
- 238000000198 fluorescence anisotropy Methods 0.000 description 2
- 238000002866 fluorescence resonance energy transfer Methods 0.000 description 2
- 125000002541 furyl group Chemical group 0.000 description 2
- 230000004927 fusion Effects 0.000 description 2
- 125000001188 haloalkyl group Chemical group 0.000 description 2
- 125000005843 halogen group Chemical group 0.000 description 2
- 239000005556 hormone Substances 0.000 description 2
- 229940088597 hormone Drugs 0.000 description 2
- 230000002209 hydrophobic effect Effects 0.000 description 2
- 125000002883 imidazolyl group Chemical group 0.000 description 2
- 210000003000 inclusion body Anatomy 0.000 description 2
- 125000001041 indolyl group Chemical group 0.000 description 2
- 239000003112 inhibitor Substances 0.000 description 2
- BPHPUYQFMNQIOC-NXRLNHOXSA-N isopropyl beta-D-thiogalactopyranoside Chemical compound CC(C)S[C@@H]1O[C@H](CO)[C@H](O)[C@H](O)[C@H]1O BPHPUYQFMNQIOC-NXRLNHOXSA-N 0.000 description 2
- 125000005956 isoquinolyl group Chemical group 0.000 description 2
- 125000001786 isothiazolyl group Chemical group 0.000 description 2
- 125000000842 isoxazolyl group Chemical group 0.000 description 2
- 235000019421 lipase Nutrition 0.000 description 2
- 210000004698 lymphocyte Anatomy 0.000 description 2
- 239000012139 lysis buffer Substances 0.000 description 2
- 239000004325 lysozyme Substances 0.000 description 2
- 229960000274 lysozyme Drugs 0.000 description 2
- 235000010335 lysozyme Nutrition 0.000 description 2
- 229910021645 metal ion Inorganic materials 0.000 description 2
- 125000002496 methyl group Chemical group [H]C([H])([H])* 0.000 description 2
- 238000002156 mixing Methods 0.000 description 2
- 230000009456 molecular mechanism Effects 0.000 description 2
- 239000012454 non-polar solvent Substances 0.000 description 2
- 230000009022 nonlinear effect Effects 0.000 description 2
- 125000001715 oxadiazolyl group Chemical group 0.000 description 2
- 125000002971 oxazolyl group Chemical group 0.000 description 2
- 238000005897 peptide coupling reaction Methods 0.000 description 2
- 239000008363 phosphate buffer Substances 0.000 description 2
- DCWXELXMIBXGTH-QMMMGPOBSA-N phosphonotyrosine Chemical group OC(=O)[C@@H](N)CC1=CC=C(OP(O)(O)=O)C=C1 DCWXELXMIBXGTH-QMMMGPOBSA-N 0.000 description 2
- 238000006366 phosphorylation reaction Methods 0.000 description 2
- 102000020233 phosphotransferase Human genes 0.000 description 2
- 230000010287 polarization Effects 0.000 description 2
- BWHMMNNQKKPAPP-UHFFFAOYSA-L potassium carbonate Chemical compound [K+].[K+].[O-]C([O-])=O BWHMMNNQKKPAPP-UHFFFAOYSA-L 0.000 description 2
- 239000000843 powder Substances 0.000 description 2
- 238000002731 protein assay Methods 0.000 description 2
- 108060006633 protein kinase Proteins 0.000 description 2
- 230000009822 protein phosphorylation Effects 0.000 description 2
- 125000000561 purinyl group Chemical group N1=C(N=C2N=CNC2=C1)* 0.000 description 2
- 125000003373 pyrazinyl group Chemical group 0.000 description 2
- 125000003226 pyrazolyl group Chemical group 0.000 description 2
- 125000002098 pyridazinyl group Chemical group 0.000 description 2
- 125000004076 pyridyl group Chemical group 0.000 description 2
- 125000000714 pyrimidinyl group Chemical group 0.000 description 2
- 125000000168 pyrrolyl group Chemical group 0.000 description 2
- 125000002943 quinolinyl group Chemical group N1=C(C=CC2=CC=CC=C12)* 0.000 description 2
- 125000005493 quinolyl group Chemical group 0.000 description 2
- 125000001567 quinoxalinyl group Chemical group N1=C(C=NC2=CC=CC=C12)* 0.000 description 2
- 230000002285 radioactive effect Effects 0.000 description 2
- 230000022983 regulation of cell cycle Effects 0.000 description 2
- 239000011347 resin Substances 0.000 description 2
- 229920005989 resin Polymers 0.000 description 2
- 150000003839 salts Chemical class 0.000 description 2
- 238000000926 separation method Methods 0.000 description 2
- 230000011664 signaling Effects 0.000 description 2
- 239000000377 silicon dioxide Substances 0.000 description 2
- 239000007790 solid phase Substances 0.000 description 2
- 238000000527 sonication Methods 0.000 description 2
- 238000010561 standard procedure Methods 0.000 description 2
- 238000000859 sublimation Methods 0.000 description 2
- 230000008022 sublimation Effects 0.000 description 2
- 238000006467 substitution reaction Methods 0.000 description 2
- 125000003831 tetrazolyl group Chemical group 0.000 description 2
- 125000001113 thiadiazolyl group Chemical group 0.000 description 2
- 125000000335 thiazolyl group Chemical group 0.000 description 2
- 125000001544 thienyl group Chemical group 0.000 description 2
- 125000004306 triazinyl group Chemical group 0.000 description 2
- 125000001425 triazolyl group Chemical group 0.000 description 2
- 229930195735 unsaturated hydrocarbon Natural products 0.000 description 2
- 238000001262 western blot Methods 0.000 description 2
- REITVGIIZHFVGU-IBGZPJMESA-N (2s)-2-(9h-fluoren-9-ylmethoxycarbonylamino)-3-[(2-methylpropan-2-yl)oxy]propanoic acid Chemical compound C1=CC=C2C(COC(=O)N[C@@H](COC(C)(C)C)C(O)=O)C3=CC=CC=C3C2=C1 REITVGIIZHFVGU-IBGZPJMESA-N 0.000 description 1
- PKAUMAVONPSDRW-IBGZPJMESA-N (2s)-2-(9h-fluoren-9-ylmethoxycarbonylamino)-3-[(2-methylpropan-2-yl)oxycarbonylamino]propanoic acid Chemical compound C1=CC=C2C(COC(=O)N[C@@H](CNC(=O)OC(C)(C)C)C(O)=O)C3=CC=CC=C3C2=C1 PKAUMAVONPSDRW-IBGZPJMESA-N 0.000 description 1
- JAUKCFULLJFBFN-VWLOTQADSA-N (2s)-2-(9h-fluoren-9-ylmethoxycarbonylamino)-3-[4-[(2-methylpropan-2-yl)oxy]phenyl]propanoic acid Chemical compound C1=CC(OC(C)(C)C)=CC=C1C[C@@H](C(O)=O)NC(=O)OCC1C2=CC=CC=C2C2=CC=CC=C21 JAUKCFULLJFBFN-VWLOTQADSA-N 0.000 description 1
- WNFKGEXQEWKECX-LJAQVGFWSA-N (2s)-2-(9h-fluoren-9-ylmethoxycarbonylamino)-3-[4-[hydroxy(phenylmethoxy)phosphoryl]oxyphenyl]propanoic acid Chemical compound C([C@@H](C(=O)O)NC(=O)OCC1C2=CC=CC=C2C2=CC=CC=C21)C(C=C1)=CC=C1OP(O)(=O)OCC1=CC=CC=C1 WNFKGEXQEWKECX-LJAQVGFWSA-N 0.000 description 1
- UGNIYGNGCNXHTR-SFHVURJKSA-N (2s)-2-(9h-fluoren-9-ylmethoxycarbonylamino)-3-methylbutanoic acid Chemical compound C1=CC=C2C(COC(=O)N[C@@H](C(C)C)C(O)=O)C3=CC=CC=C3C2=C1 UGNIYGNGCNXHTR-SFHVURJKSA-N 0.000 description 1
- FODJWPHPWBKDON-IBGZPJMESA-N (2s)-2-(9h-fluoren-9-ylmethoxycarbonylamino)-4-[(2-methylpropan-2-yl)oxy]-4-oxobutanoic acid Chemical compound C1=CC=C2C(COC(=O)N[C@@H](CC(=O)OC(C)(C)C)C(O)=O)C3=CC=CC=C3C2=C1 FODJWPHPWBKDON-IBGZPJMESA-N 0.000 description 1
- OTKXCALUHMPIGM-FQEVSTJZSA-N (2s)-2-(9h-fluoren-9-ylmethoxycarbonylamino)-5-[(2-methylpropan-2-yl)oxy]-5-oxopentanoic acid Chemical compound C1=CC=C2C(COC(=O)N[C@@H](CCC(=O)OC(C)(C)C)C(O)=O)C3=CC=CC=C3C2=C1 OTKXCALUHMPIGM-FQEVSTJZSA-N 0.000 description 1
- QWXZOFZKSQXPDC-NSHDSACASA-N (2s)-2-(9h-fluoren-9-ylmethoxycarbonylamino)propanoic acid Chemical compound C1=CC=C2C(COC(=O)N[C@@H](C)C(O)=O)C3=CC=CC=C3C2=C1 QWXZOFZKSQXPDC-NSHDSACASA-N 0.000 description 1
- HNICLNKVURBTKV-NDEPHWFRSA-N (2s)-5-[[amino-[(2,2,4,6,7-pentamethyl-3h-1-benzofuran-5-yl)sulfonylamino]methylidene]amino]-2-(9h-fluoren-9-ylmethoxycarbonylamino)pentanoic acid Chemical compound C12=CC=CC=C2C2=CC=CC=C2C1COC(=O)N[C@H](C(O)=O)CCCN=C(N)NS(=O)(=O)C1=C(C)C(C)=C2OC(C)(C)CC2=C1C HNICLNKVURBTKV-NDEPHWFRSA-N 0.000 description 1
- LZOLWEQBVPVDPR-VLIAUNLRSA-N (2s,3r)-2-(9h-fluoren-9-ylmethoxycarbonylamino)-3-[(2-methylpropan-2-yl)oxy]butanoic acid Chemical compound C1=CC=C2C(COC(=O)N[C@@H]([C@H](OC(C)(C)C)C)C(O)=O)C3=CC=CC=C3C2=C1 LZOLWEQBVPVDPR-VLIAUNLRSA-N 0.000 description 1
- QXVFEIPAZSXRGM-DJJJIMSYSA-N (2s,3s)-2-(9h-fluoren-9-ylmethoxycarbonylamino)-3-methylpentanoic acid Chemical compound C1=CC=C2C(COC(=O)N[C@@H]([C@@H](C)CC)C(O)=O)C3=CC=CC=C3C2=C1 QXVFEIPAZSXRGM-DJJJIMSYSA-N 0.000 description 1
- 125000004178 (C1-C4) alkyl group Chemical group 0.000 description 1
- WNXJIVFYUVYPPR-UHFFFAOYSA-N 1,3-dioxolane Chemical compound C1COCO1 WNXJIVFYUVYPPR-UHFFFAOYSA-N 0.000 description 1
- NDOVLWQBFFJETK-UHFFFAOYSA-N 1,4-thiazinane 1,1-dioxide Chemical compound O=S1(=O)CCNCC1 NDOVLWQBFFJETK-UHFFFAOYSA-N 0.000 description 1
- YHIIJNLSGULWAA-UHFFFAOYSA-N 1,4-thiazinane 1-oxide Chemical compound O=S1CCNCC1 YHIIJNLSGULWAA-UHFFFAOYSA-N 0.000 description 1
- XGVAEAVGTOZSNS-UHFFFAOYSA-N 1-[(4-methoxyphenoxy)methyl]-4-nitrobenzene Chemical compound C1=CC(OC)=CC=C1OCC1=CC=C([N+]([O-])=O)C=C1 XGVAEAVGTOZSNS-UHFFFAOYSA-N 0.000 description 1
- 125000001478 1-chloroethyl group Chemical group [H]C([H])([H])C([H])(Cl)* 0.000 description 1
- 125000004214 1-pyrrolidinyl group Chemical group [H]C1([H])N(*)C([H])([H])C([H])([H])C1([H])[H] 0.000 description 1
- XEZNGIUYQVAUSS-UHFFFAOYSA-N 18-crown-6 Chemical compound C1COCCOCCOCCOCCOCCO1 XEZNGIUYQVAUSS-UHFFFAOYSA-N 0.000 description 1
- UTQNKKSJPHTPBS-UHFFFAOYSA-N 2,2,2-trichloroethanone Chemical group ClC(Cl)(Cl)[C]=O UTQNKKSJPHTPBS-UHFFFAOYSA-N 0.000 description 1
- MEKOFIRRDATTAG-UHFFFAOYSA-N 2,2,5,8-tetramethyl-3,4-dihydrochromen-6-ol Chemical compound C1CC(C)(C)OC2=C1C(C)=C(O)C=C2C MEKOFIRRDATTAG-UHFFFAOYSA-N 0.000 description 1
- YQTCQNIPQMJNTI-UHFFFAOYSA-N 2,2-dimethylpropan-1-one Chemical group CC(C)(C)[C]=O YQTCQNIPQMJNTI-UHFFFAOYSA-N 0.000 description 1
- OWQJSRUZDKLDPT-UHFFFAOYSA-N 2-(benzenesulfonylmethyl)-5-nitrobenzaldehyde Chemical compound O=CC1=CC([N+](=O)[O-])=CC=C1CS(=O)(=O)C1=CC=CC=C1 OWQJSRUZDKLDPT-UHFFFAOYSA-N 0.000 description 1
- MGADZUXDNSDTHW-UHFFFAOYSA-N 2H-pyran Chemical compound C1OC=CC=C1 MGADZUXDNSDTHW-UHFFFAOYSA-N 0.000 description 1
- ZPSJGADGUYYRKE-UHFFFAOYSA-N 2H-pyran-2-one Chemical compound O=C1C=CC=CO1 ZPSJGADGUYYRKE-UHFFFAOYSA-N 0.000 description 1
- LLPBUXODFQZPFH-UHFFFAOYSA-N 3,3-dimethyl-1h-benzo[g]indole-2,4,5-trione Chemical compound O=C1C(=O)C2=CC=CC=C2C2=C1C(C)(C)C(=O)N2 LLPBUXODFQZPFH-UHFFFAOYSA-N 0.000 description 1
- ZETIVVHRRQLWFW-UHFFFAOYSA-N 3-nitrobenzaldehyde Chemical compound [O-][N+](=O)C1=CC=CC(C=O)=C1 ZETIVVHRRQLWFW-UHFFFAOYSA-N 0.000 description 1
- JVQIKJMSUIMUDI-UHFFFAOYSA-N 3-pyrroline Chemical compound C1NCC=C1 JVQIKJMSUIMUDI-UHFFFAOYSA-N 0.000 description 1
- CSDQQAQKBAQLLE-UHFFFAOYSA-N 4-(4-chlorophenyl)-4,5,6,7-tetrahydrothieno[3,2-c]pyridine Chemical compound C1=CC(Cl)=CC=C1C1C(C=CS2)=C2CCN1 CSDQQAQKBAQLLE-UHFFFAOYSA-N 0.000 description 1
- SSMIFVHARFVINF-UHFFFAOYSA-N 4-amino-1,8-naphthalimide Chemical class O=C1NC(=O)C2=CC=CC3=C2C1=CC=C3N SSMIFVHARFVINF-UHFFFAOYSA-N 0.000 description 1
- QRXMUCSWCMTJGU-UHFFFAOYSA-N 5-bromo-4-chloro-3-indolyl phosphate Chemical compound C1=C(Br)C(Cl)=C2C(OP(O)(=O)O)=CNC2=C1 QRXMUCSWCMTJGU-UHFFFAOYSA-N 0.000 description 1
- HBAQYPYDRFILMT-UHFFFAOYSA-N 8-[3-(1-cyclopropylpyrazol-4-yl)-1H-pyrazolo[4,3-d]pyrimidin-5-yl]-3-methyl-3,8-diazabicyclo[3.2.1]octan-2-one Chemical class C1(CC1)N1N=CC(=C1)C1=NNC2=C1N=C(N=C2)N1C2C(N(CC1CC2)C)=O HBAQYPYDRFILMT-UHFFFAOYSA-N 0.000 description 1
- 229920000936 Agarose Polymers 0.000 description 1
- 102000002260 Alkaline Phosphatase Human genes 0.000 description 1
- 108020004774 Alkaline Phosphatase Proteins 0.000 description 1
- 208000024827 Alzheimer disease Diseases 0.000 description 1
- 102000007592 Apolipoproteins Human genes 0.000 description 1
- 108010071619 Apolipoproteins Proteins 0.000 description 1
- 238000009020 BCA Protein Assay Kit Methods 0.000 description 1
- XCRHIFZSKWEBPF-UHFFFAOYSA-N C1CCC1.CC(C)C Chemical compound C1CCC1.CC(C)C XCRHIFZSKWEBPF-UHFFFAOYSA-N 0.000 description 1
- SVXGUXAJMSTRSL-DRLLIAJESA-N C=CCOC(=O)[C@H](CN)NC(=O)OCC1C2=C(C=CC=C2)C2=C1C=CC=C2.C=CCOC(=O)[C@H](CN1C(=O)C2=CC3=CC=C(N(C)C)C=C3C=C2C1=O)NC(=O)OCC1C2=C(C=CC=C2)C2=C1C=CC=C2.C=CCOC(=O)[C@H](CNC(=O)OC(C)(C)C)NC(=O)OCC1C2=C(C=CC=C2)C2=C1C=CC=C2.CC(C)(C)OC(=O)NC[C@H](NC(=O)OCC1C2=C(C=CC=C2)C2=C1C=CC=C2)C(=O)O.CN(C)C1=CC=C2C=C3C(=O)N(C[C@H](NC(=O)OCC4C5=C(C=CC=C5)C5=C4C=CC=C5)C(=O)O)C(=O)C3=CC2=C1 Chemical compound C=CCOC(=O)[C@H](CN)NC(=O)OCC1C2=C(C=CC=C2)C2=C1C=CC=C2.C=CCOC(=O)[C@H](CN1C(=O)C2=CC3=CC=C(N(C)C)C=C3C=C2C1=O)NC(=O)OCC1C2=C(C=CC=C2)C2=C1C=CC=C2.C=CCOC(=O)[C@H](CNC(=O)OC(C)(C)C)NC(=O)OCC1C2=C(C=CC=C2)C2=C1C=CC=C2.CC(C)(C)OC(=O)NC[C@H](NC(=O)OCC1C2=C(C=CC=C2)C2=C1C=CC=C2)C(=O)O.CN(C)C1=CC=C2C=C3C(=O)N(C[C@H](NC(=O)OCC4C5=C(C=CC=C5)C5=C4C=CC=C5)C(=O)O)C(=O)C3=CC2=C1 SVXGUXAJMSTRSL-DRLLIAJESA-N 0.000 description 1
- GXQCYIDNIOBARY-UHFFFAOYSA-N CCOC(=O)C1=CC2=CC=C(N(C)C)C=C2C=C1C(=O)OCC.CCOC(=O)C1=CC2=CC=C([N+](=O)[O-])C=C2C=C1C(=O)OCC.CN(C)C1=CC=C2C=C3C(=O)OC(=O)C3=CC2=C1.O=CC1=CC([N+](=O)[O-])=CC=C1.O=[N+]([O-])C1=CC=C(CS(=O)(=O)C2=CC=CC=C2)C(C2OCCO2)=C1.O=[N+]([O-])C1=CC=CC(C2OCCO2)=C1 Chemical compound CCOC(=O)C1=CC2=CC=C(N(C)C)C=C2C=C1C(=O)OCC.CCOC(=O)C1=CC2=CC=C([N+](=O)[O-])C=C2C=C1C(=O)OCC.CN(C)C1=CC=C2C=C3C(=O)OC(=O)C3=CC2=C1.O=CC1=CC([N+](=O)[O-])=CC=C1.O=[N+]([O-])C1=CC=C(CS(=O)(=O)C2=CC=CC=C2)C(C2OCCO2)=C1.O=[N+]([O-])C1=CC=CC(C2OCCO2)=C1 GXQCYIDNIOBARY-UHFFFAOYSA-N 0.000 description 1
- MVUUQAKIQCXRQZ-JTQLQIEISA-N C[C@@H](CN1C(=O)C2=CC3=CC=C(N(C)C)C=C3C=C2C1=O)C(=O)O Chemical compound C[C@@H](CN1C(=O)C2=CC3=CC=C(N(C)C)C=C3C=C2C1=O)C(=O)O MVUUQAKIQCXRQZ-JTQLQIEISA-N 0.000 description 1
- 108090000565 Capsid Proteins Proteins 0.000 description 1
- OKTJSMMVPCPJKN-NJFSPNSNSA-N Carbon-14 Chemical compound [14C] OKTJSMMVPCPJKN-NJFSPNSNSA-N 0.000 description 1
- 108050001186 Chaperonin Cpn60 Proteins 0.000 description 1
- 102000052603 Chaperonins Human genes 0.000 description 1
- KZBUYRJDOAKODT-UHFFFAOYSA-N Chlorine Chemical compound ClCl KZBUYRJDOAKODT-UHFFFAOYSA-N 0.000 description 1
- 238000004057 DFT-B3LYP calculation Methods 0.000 description 1
- 102000053602 DNA Human genes 0.000 description 1
- 238000003775 Density Functional Theory Methods 0.000 description 1
- 238000005698 Diels-Alder reaction Methods 0.000 description 1
- 241000588724 Escherichia coli Species 0.000 description 1
- 241000620209 Escherichia coli DH5[alpha] Species 0.000 description 1
- LYCAIKOWRPUZTN-UHFFFAOYSA-N Ethylene glycol Chemical compound OCCO LYCAIKOWRPUZTN-UHFFFAOYSA-N 0.000 description 1
- 102000010834 Extracellular Matrix Proteins Human genes 0.000 description 1
- 108010037362 Extracellular Matrix Proteins Proteins 0.000 description 1
- 102000003688 G-Protein-Coupled Receptors Human genes 0.000 description 1
- 108090000045 G-Protein-Coupled Receptors Proteins 0.000 description 1
- 102000003886 Glycoproteins Human genes 0.000 description 1
- 108090000288 Glycoproteins Proteins 0.000 description 1
- 102000004157 Hydrolases Human genes 0.000 description 1
- 108090000604 Hydrolases Proteins 0.000 description 1
- DGAQECJNVWCQMB-PUAWFVPOSA-M Ilexoside XXIX Chemical compound C[C@@H]1CC[C@@]2(CC[C@@]3(C(=CC[C@H]4[C@]3(CC[C@@H]5[C@@]4(CC[C@@H](C5(C)C)OS(=O)(=O)[O-])C)C)[C@@H]2[C@]1(C)O)C)C(=O)O[C@H]6[C@@H]([C@H]([C@@H]([C@H](O6)CO)O)O)O.[Na+] DGAQECJNVWCQMB-PUAWFVPOSA-M 0.000 description 1
- WRYCSMQKUKOKBP-UHFFFAOYSA-N Imidazolidine Chemical compound C1CNCN1 WRYCSMQKUKOKBP-UHFFFAOYSA-N 0.000 description 1
- 206010061218 Inflammation Diseases 0.000 description 1
- 102000004895 Lipoproteins Human genes 0.000 description 1
- 108090001030 Lipoproteins Proteins 0.000 description 1
- 239000006137 Luria-Bertani broth Substances 0.000 description 1
- 102000018697 Membrane Proteins Human genes 0.000 description 1
- 108010052285 Membrane Proteins Proteins 0.000 description 1
- 102000029749 Microtubule Human genes 0.000 description 1
- 108091022875 Microtubule Proteins 0.000 description 1
- 150000001204 N-oxides Chemical class 0.000 description 1
- 239000000020 Nitrocellulose Substances 0.000 description 1
- BZQFBWGGLXLEPQ-UHFFFAOYSA-N O-phosphoryl-L-serine Natural products OC(=O)C(N)COP(O)(O)=O BZQFBWGGLXLEPQ-UHFFFAOYSA-N 0.000 description 1
- 229910019142 PO4 Inorganic materials 0.000 description 1
- NFHFRUOZVGFOOS-UHFFFAOYSA-N Pd(PPh3)4 Substances [Pd].C1=CC=CC=C1P(C=1C=CC=CC=1)C1=CC=CC=C1.C1=CC=CC=C1P(C=1C=CC=CC=1)C1=CC=CC=C1.C1=CC=CC=C1P(C=1C=CC=CC=1)C1=CC=CC=C1.C1=CC=CC=C1P(C=1C=CC=CC=1)C1=CC=CC=C1 NFHFRUOZVGFOOS-UHFFFAOYSA-N 0.000 description 1
- 108091005804 Peptidases Proteins 0.000 description 1
- 102000035195 Peptidases Human genes 0.000 description 1
- 102000045595 Phosphoprotein Phosphatases Human genes 0.000 description 1
- 108700019535 Phosphoprotein Phosphatases Proteins 0.000 description 1
- 102000004160 Phosphoric Monoester Hydrolases Human genes 0.000 description 1
- 108090000608 Phosphoric Monoester Hydrolases Proteins 0.000 description 1
- 229920000361 Poly(styrene)-block-poly(ethylene glycol) Polymers 0.000 description 1
- 239000004365 Protease Substances 0.000 description 1
- 102000004022 Protein-Tyrosine Kinases Human genes 0.000 description 1
- 108090000412 Protein-Tyrosine Kinases Proteins 0.000 description 1
- 108020004511 Recombinant DNA Proteins 0.000 description 1
- 229920002684 Sepharose Polymers 0.000 description 1
- XUIMIQQOPSSXEZ-UHFFFAOYSA-N Silicon Chemical compound [Si] XUIMIQQOPSSXEZ-UHFFFAOYSA-N 0.000 description 1
- 206010041316 Solvent sensitivity Diseases 0.000 description 1
- 108091023040 Transcription factor Proteins 0.000 description 1
- 102000040945 Transcription factor Human genes 0.000 description 1
- YZCKVEUIGOORGS-NJFSPNSNSA-N Tritium Chemical compound [3H] YZCKVEUIGOORGS-NJFSPNSNSA-N 0.000 description 1
- 108010067390 Viral Proteins Proteins 0.000 description 1
- 241000700605 Viruses Species 0.000 description 1
- PNDPGZBMCMUPRI-XXSWNUTMSA-N [125I][125I] Chemical compound [125I][125I] PNDPGZBMCMUPRI-XXSWNUTMSA-N 0.000 description 1
- JLCPHMBAVCMARE-UHFFFAOYSA-N [3-[[3-[[3-[[3-[[3-[[3-[[3-[[3-[[3-[[3-[[3-[[5-(2-amino-6-oxo-1H-purin-9-yl)-3-[[3-[[3-[[3-[[3-[[3-[[5-(2-amino-6-oxo-1H-purin-9-yl)-3-[[5-(2-amino-6-oxo-1H-purin-9-yl)-3-hydroxyoxolan-2-yl]methoxy-hydroxyphosphoryl]oxyoxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(5-methyl-2,4-dioxopyrimidin-1-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(6-aminopurin-9-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(6-aminopurin-9-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(6-aminopurin-9-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(6-aminopurin-9-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxyoxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(5-methyl-2,4-dioxopyrimidin-1-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(4-amino-2-oxopyrimidin-1-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(5-methyl-2,4-dioxopyrimidin-1-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(5-methyl-2,4-dioxopyrimidin-1-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(6-aminopurin-9-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(6-aminopurin-9-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(4-amino-2-oxopyrimidin-1-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(4-amino-2-oxopyrimidin-1-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(4-amino-2-oxopyrimidin-1-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(6-aminopurin-9-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(4-amino-2-oxopyrimidin-1-yl)oxolan-2-yl]methyl [5-(6-aminopurin-9-yl)-2-(hydroxymethyl)oxolan-3-yl] hydrogen phosphate Polymers Cc1cn(C2CC(OP(O)(=O)OCC3OC(CC3OP(O)(=O)OCC3OC(CC3O)n3cnc4c3nc(N)[nH]c4=O)n3cnc4c3nc(N)[nH]c4=O)C(COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3CO)n3cnc4c(N)ncnc34)n3ccc(N)nc3=O)n3cnc4c(N)ncnc34)n3ccc(N)nc3=O)n3ccc(N)nc3=O)n3ccc(N)nc3=O)n3cnc4c(N)ncnc34)n3cnc4c(N)ncnc34)n3cc(C)c(=O)[nH]c3=O)n3cc(C)c(=O)[nH]c3=O)n3ccc(N)nc3=O)n3cc(C)c(=O)[nH]c3=O)n3cnc4c3nc(N)[nH]c4=O)n3cnc4c(N)ncnc34)n3cnc4c(N)ncnc34)n3cnc4c(N)ncnc34)n3cnc4c(N)ncnc34)O2)c(=O)[nH]c1=O JLCPHMBAVCMARE-UHFFFAOYSA-N 0.000 description 1
- 230000001594 aberrant effect Effects 0.000 description 1
- 230000002159 abnormal effect Effects 0.000 description 1
- 230000005856 abnormality Effects 0.000 description 1
- 125000002777 acetyl group Chemical group [H]C([H])([H])C(*)=O 0.000 description 1
- 239000002253 acid Substances 0.000 description 1
- 230000002378 acidificating effect Effects 0.000 description 1
- 230000009471 action Effects 0.000 description 1
- 230000003213 activating effect Effects 0.000 description 1
- 230000004913 activation Effects 0.000 description 1
- 239000012190 activator Substances 0.000 description 1
- 125000002252 acyl group Chemical group 0.000 description 1
- 150000001266 acyl halides Chemical class 0.000 description 1
- 125000004423 acyloxy group Chemical group 0.000 description 1
- 150000001299 aldehydes Chemical class 0.000 description 1
- 125000004450 alkenylene group Chemical group 0.000 description 1
- 125000002947 alkylene group Chemical group 0.000 description 1
- 125000004419 alkynylene group Chemical group 0.000 description 1
- BHELZAPQIKSEDF-UHFFFAOYSA-N allyl bromide Chemical compound BrCC=C BHELZAPQIKSEDF-UHFFFAOYSA-N 0.000 description 1
- HSFWRNGVRCDJHI-UHFFFAOYSA-N alpha-acetylene Natural products C#C HSFWRNGVRCDJHI-UHFFFAOYSA-N 0.000 description 1
- 150000003862 amino acid derivatives Chemical class 0.000 description 1
- 125000003277 amino group Chemical group 0.000 description 1
- 125000004103 aminoalkyl group Chemical group 0.000 description 1
- 238000004458 analytical method Methods 0.000 description 1
- 238000010719 annulation reaction Methods 0.000 description 1
- 230000009830 antibody antigen interaction Effects 0.000 description 1
- 229910052786 argon Inorganic materials 0.000 description 1
- 125000002029 aromatic hydrocarbon group Chemical group 0.000 description 1
- 125000005160 aryl oxy alkyl group Chemical group 0.000 description 1
- 238000003556 assay Methods 0.000 description 1
- 125000004603 benzisoxazolyl group Chemical group O1N=C(C2=C1C=CC=C2)* 0.000 description 1
- 125000000499 benzofuranyl group Chemical group O1C(=CC2=C1C=CC=C2)* 0.000 description 1
- 125000004196 benzothienyl group Chemical group S1C(=CC2=C1C=CC=C2)* 0.000 description 1
- 125000003236 benzoyl group Chemical group [H]C1=C([H])C([H])=C(C([H])=C1[H])C(*)=O 0.000 description 1
- 125000001797 benzyl group Chemical group [H]C1=C([H])C([H])=C(C([H])=C1[H])C([H])([H])* 0.000 description 1
- GPRLTFBKWDERLU-UHFFFAOYSA-N bicyclo[2.2.2]octane Chemical compound C1CC2CCC1CC2 GPRLTFBKWDERLU-UHFFFAOYSA-N 0.000 description 1
- 230000003851 biochemical process Effects 0.000 description 1
- 238000005842 biochemical reaction Methods 0.000 description 1
- 230000008827 biological function Effects 0.000 description 1
- 230000031018 biological processes and functions Effects 0.000 description 1
- 230000033228 biological regulation Effects 0.000 description 1
- 239000004305 biphenyl Chemical group 0.000 description 1
- 235000010290 biphenyl Nutrition 0.000 description 1
- 229910052794 bromium Inorganic materials 0.000 description 1
- 210000004899 c-terminal region Anatomy 0.000 description 1
- 229910000024 caesium carbonate Inorganic materials 0.000 description 1
- 201000011510 cancer Diseases 0.000 description 1
- 238000001460 carbon-13 nuclear magnetic resonance spectrum Methods 0.000 description 1
- 150000001732 carboxylic acid derivatives Chemical class 0.000 description 1
- 230000003915 cell function Effects 0.000 description 1
- 230000012292 cell migration Effects 0.000 description 1
- 230000009134 cell regulation Effects 0.000 description 1
- 230000010267 cellular communication Effects 0.000 description 1
- 230000007248 cellular mechanism Effects 0.000 description 1
- 239000001913 cellulose Substances 0.000 description 1
- 229920002678 cellulose Polymers 0.000 description 1
- 239000002738 chelating agent Substances 0.000 description 1
- 239000003638 chemical reducing agent Substances 0.000 description 1
- 239000003795 chemical substances by application Substances 0.000 description 1
- 239000000460 chlorine Substances 0.000 description 1
- 229910052801 chlorine Inorganic materials 0.000 description 1
- NXAIQSVCXQZNRY-UHFFFAOYSA-N chloromethylsulfonylbenzene Chemical compound ClCS(=O)(=O)C1=CC=CC=C1 NXAIQSVCXQZNRY-UHFFFAOYSA-N 0.000 description 1
- 239000013611 chromosomal DNA Substances 0.000 description 1
- 125000000259 cinnolinyl group Chemical group N1=NC(=CC2=CC=CC=C12)* 0.000 description 1
- 238000003776 cleavage reaction Methods 0.000 description 1
- 230000004154 complement system Effects 0.000 description 1
- 229940125782 compound 2 Drugs 0.000 description 1
- 229940125898 compound 5 Drugs 0.000 description 1
- 239000000356 contaminant Substances 0.000 description 1
- 230000008878 coupling Effects 0.000 description 1
- 238000010168 coupling process Methods 0.000 description 1
- 238000005859 coupling reaction Methods 0.000 description 1
- 125000000596 cyclohexenyl group Chemical group C1(=CCCCC1)* 0.000 description 1
- 125000000113 cyclohexyl group Chemical group [H]C1([H])C([H])([H])C([H])([H])C([H])(*)C([H])([H])C1([H])[H] 0.000 description 1
- 125000002433 cyclopentenyl group Chemical group C1(=CCCC1)* 0.000 description 1
- 125000001511 cyclopentyl group Chemical group [H]C1([H])C([H])([H])C([H])([H])C([H])(*)C1([H])[H] 0.000 description 1
- 125000004186 cyclopropylmethyl group Chemical group [H]C([H])(*)C1([H])C([H])([H])C1([H])[H] 0.000 description 1
- 230000009089 cytolysis Effects 0.000 description 1
- 238000013461 design Methods 0.000 description 1
- 238000001514 detection method Methods 0.000 description 1
- 239000003599 detergent Substances 0.000 description 1
- 238000011161 development Methods 0.000 description 1
- 229950006137 dexfosfoserine Drugs 0.000 description 1
- 150000005690 diesters Chemical class 0.000 description 1
- VKNUORWMCINMRB-UHFFFAOYSA-N diethyl malate Chemical compound CCOC(=O)CC(O)C(=O)OCC VKNUORWMCINMRB-UHFFFAOYSA-N 0.000 description 1
- 238000007865 diluting Methods 0.000 description 1
- BNIILDVGGAEEIG-UHFFFAOYSA-L disodium hydrogen phosphate Chemical compound [Na+].[Na+].OP([O-])([O-])=O BNIILDVGGAEEIG-UHFFFAOYSA-L 0.000 description 1
- 229910000397 disodium phosphate Inorganic materials 0.000 description 1
- 239000012154 double-distilled water Substances 0.000 description 1
- 239000003814 drug Substances 0.000 description 1
- 230000009977 dual effect Effects 0.000 description 1
- 239000012143 dye reagent concentrate Substances 0.000 description 1
- 238000005516 engineering process Methods 0.000 description 1
- 238000011067 equilibration Methods 0.000 description 1
- 230000032050 esterification Effects 0.000 description 1
- 238000005886 esterification reaction Methods 0.000 description 1
- 150000002148 esters Chemical class 0.000 description 1
- 125000003754 ethoxycarbonyl group Chemical group C(=O)(OCC)* 0.000 description 1
- 125000002534 ethynyl group Chemical group [H]C#C* 0.000 description 1
- 210000002744 extracellular matrix Anatomy 0.000 description 1
- 230000008622 extracellular signaling Effects 0.000 description 1
- 238000002875 fluorescence polarization Methods 0.000 description 1
- 238000001506 fluorescence spectroscopy Methods 0.000 description 1
- 125000001153 fluoro group Chemical group F* 0.000 description 1
- 125000002485 formyl group Chemical group [H]C(*)=O 0.000 description 1
- 230000006870 function Effects 0.000 description 1
- 239000000499 gel Substances 0.000 description 1
- 238000001502 gel electrophoresis Methods 0.000 description 1
- 102000034356 gene-regulatory proteins Human genes 0.000 description 1
- 108091006104 gene-regulatory proteins Proteins 0.000 description 1
- 229960003180 glutathione Drugs 0.000 description 1
- 230000012010 growth Effects 0.000 description 1
- 239000003102 growth factor Substances 0.000 description 1
- 238000010438 heat treatment Methods 0.000 description 1
- WJRBRSLFGCUECM-UHFFFAOYSA-N hydantoin Chemical compound O=C1CNC(=O)N1 WJRBRSLFGCUECM-UHFFFAOYSA-N 0.000 description 1
- 229940091173 hydantoin Drugs 0.000 description 1
- COQRGFWWJBEXRC-UHFFFAOYSA-N hydron;methyl 2-aminoacetate;chloride Chemical compound Cl.COC(=O)CN COQRGFWWJBEXRC-UHFFFAOYSA-N 0.000 description 1
- 125000004356 hydroxy functional group Chemical group O* 0.000 description 1
- YAMHXTCMCPHKLN-UHFFFAOYSA-N imidazolidin-2-one Chemical compound O=C1NCCN1 YAMHXTCMCPHKLN-UHFFFAOYSA-N 0.000 description 1
- 125000004857 imidazopyridinyl group Chemical class N1C(=NC2=C1C=CC=N2)* 0.000 description 1
- 230000003832 immune regulation Effects 0.000 description 1
- 238000011065 in-situ storage Methods 0.000 description 1
- 125000003406 indolizinyl group Chemical group C=1(C=CN2C=CC=CC12)* 0.000 description 1
- 208000015181 infectious disease Diseases 0.000 description 1
- 210000004969 inflammatory cell Anatomy 0.000 description 1
- 230000004054 inflammatory process Effects 0.000 description 1
- 102000006495 integrins Human genes 0.000 description 1
- 108010044426 integrins Proteins 0.000 description 1
- 230000003834 intracellular effect Effects 0.000 description 1
- 230000004068 intracellular signaling Effects 0.000 description 1
- PNDPGZBMCMUPRI-UHFFFAOYSA-N iodine Chemical compound II PNDPGZBMCMUPRI-UHFFFAOYSA-N 0.000 description 1
- 229940044173 iodine-125 Drugs 0.000 description 1
- 125000000959 isobutyl group Chemical group [H]C([H])([H])C([H])(C([H])([H])[H])C([H])([H])* 0.000 description 1
- 125000000904 isoindolyl group Chemical group C=1(NC=C2C=CC=CC12)* 0.000 description 1
- 125000000555 isopropenyl group Chemical group [H]\C([H])=C(\*)C([H])([H])[H] 0.000 description 1
- 125000001449 isopropyl group Chemical group [H]C([H])([H])C([H])(*)C([H])([H])[H] 0.000 description 1
- 230000000155 isotopic effect Effects 0.000 description 1
- 239000003446 ligand Substances 0.000 description 1
- 239000007788 liquid Substances 0.000 description 1
- 239000004973 liquid crystal related substance Substances 0.000 description 1
- 238000011068 loading method Methods 0.000 description 1
- 239000006166 lysate Substances 0.000 description 1
- 230000001404 mediated effect Effects 0.000 description 1
- LVWZTYCIRDMTEY-UHFFFAOYSA-N metamizole Chemical compound O=C1C(N(CS(O)(=O)=O)C)=C(C)N(C)N1C1=CC=CC=C1 LVWZTYCIRDMTEY-UHFFFAOYSA-N 0.000 description 1
- 125000001160 methoxycarbonyl group Chemical group [H]C([H])([H])OC(*)=O 0.000 description 1
- 210000004688 microtubule Anatomy 0.000 description 1
- 230000004048 modification Effects 0.000 description 1
- 238000012986 modification Methods 0.000 description 1
- 125000006682 monohaloalkyl group Chemical group 0.000 description 1
- 125000004573 morpholin-4-yl group Chemical group N1(CCOCC1)* 0.000 description 1
- 125000002757 morpholinyl group Chemical group 0.000 description 1
- 125000004108 n-butyl group Chemical group [H]C([H])([H])C([H])([H])C([H])([H])C([H])([H])* 0.000 description 1
- 125000004123 n-propyl group Chemical group [H]C([H])([H])C([H])([H])C([H])([H])* 0.000 description 1
- 230000001537 neural effect Effects 0.000 description 1
- 229920001220 nitrocellulos Polymers 0.000 description 1
- 125000006574 non-aromatic ring group Chemical group 0.000 description 1
- UMRZSTCPUPJPOJ-KNVOCYPGSA-N norbornane Chemical compound C1C[C@H]2CC[C@@H]1C2 UMRZSTCPUPJPOJ-KNVOCYPGSA-N 0.000 description 1
- 238000000655 nuclear magnetic resonance spectrum Methods 0.000 description 1
- 239000002773 nucleotide Substances 0.000 description 1
- 125000003729 nucleotide group Chemical group 0.000 description 1
- 229920001542 oligosaccharide Polymers 0.000 description 1
- 150000002482 oligosaccharides Chemical class 0.000 description 1
- 230000003287 optical effect Effects 0.000 description 1
- 239000007800 oxidant agent Substances 0.000 description 1
- 239000008188 pellet Substances 0.000 description 1
- 108010091212 pepstatin Proteins 0.000 description 1
- FAXGPCHRFPCXOO-LXTPJMTPSA-N pepstatin A Chemical compound OC(=O)C[C@H](O)[C@H](CC(C)C)NC(=O)[C@H](C)NC(=O)C[C@H](O)[C@H](CC(C)C)NC(=O)[C@H](C(C)C)NC(=O)[C@H](C(C)C)NC(=O)CC(C)C FAXGPCHRFPCXOO-LXTPJMTPSA-N 0.000 description 1
- 230000000144 pharmacologic effect Effects 0.000 description 1
- 125000001997 phenyl group Chemical group [H]C1=C([H])C([H])=C(*)C([H])=C1[H] 0.000 description 1
- UYWQUFXKFGHYNT-UHFFFAOYSA-N phenylmethyl ester of formic acid Natural products O=COCC1=CC=CC=C1 UYWQUFXKFGHYNT-UHFFFAOYSA-N 0.000 description 1
- PARWUHTVGZSQPD-UHFFFAOYSA-N phenylsilane Chemical compound [SiH3]C1=CC=CC=C1 PARWUHTVGZSQPD-UHFFFAOYSA-N 0.000 description 1
- 125000003170 phenylsulfonyl group Chemical group C1(=CC=CC=C1)S(=O)(=O)* 0.000 description 1
- NBIIXXVUZAFLBC-UHFFFAOYSA-K phosphate Chemical compound [O-]P([O-])([O-])=O NBIIXXVUZAFLBC-UHFFFAOYSA-K 0.000 description 1
- 239000010452 phosphate Substances 0.000 description 1
- 125000002467 phosphate group Chemical group [H]OP(=O)(O[H])O[*] 0.000 description 1
- 230000026731 phosphorylation Effects 0.000 description 1
- BZQFBWGGLXLEPQ-REOHCLBHSA-N phosphoserine Chemical compound OC(=O)[C@@H](N)COP(O)(O)=O BZQFBWGGLXLEPQ-REOHCLBHSA-N 0.000 description 1
- 125000004592 phthalazinyl group Chemical group C1(=NN=CC2=CC=CC=C12)* 0.000 description 1
- XKJCHHZQLQNZHY-UHFFFAOYSA-N phthalimide Chemical compound C1=CC=C2C(=O)NC(=O)C2=C1 XKJCHHZQLQNZHY-UHFFFAOYSA-N 0.000 description 1
- XUWHAWMETYGRKB-UHFFFAOYSA-N piperidin-2-one Chemical compound O=C1CCCCN1 XUWHAWMETYGRKB-UHFFFAOYSA-N 0.000 description 1
- 125000003386 piperidinyl group Chemical group 0.000 description 1
- 239000013612 plasmid Substances 0.000 description 1
- 230000010118 platelet activation Effects 0.000 description 1
- 125000006684 polyhaloalkyl group Polymers 0.000 description 1
- 230000004481 post-translational protein modification Effects 0.000 description 1
- 229910000027 potassium carbonate Inorganic materials 0.000 description 1
- 235000008476 powdered milk Nutrition 0.000 description 1
- 239000002243 precursor Substances 0.000 description 1
- 238000002360 preparation method Methods 0.000 description 1
- 238000002953 preparative HPLC Methods 0.000 description 1
- 125000001501 propionyl group Chemical group O=C([*])C([H])([H])C([H])([H])[H] 0.000 description 1
- 125000001436 propyl group Chemical group [H]C([*])([H])C([H])([H])C([H])([H])[H] 0.000 description 1
- 230000012846 protein folding Effects 0.000 description 1
- 230000006916 protein interaction Effects 0.000 description 1
- 230000009145 protein modification Effects 0.000 description 1
- 230000006432 protein unfolding Effects 0.000 description 1
- 238000000425 proton nuclear magnetic resonance spectrum Methods 0.000 description 1
- 125000001042 pteridinyl group Chemical group N1=C(N=CC2=NC=CN=C12)* 0.000 description 1
- USPWKWBDZOARPV-UHFFFAOYSA-N pyrazolidine Chemical compound C1CNNC1 USPWKWBDZOARPV-UHFFFAOYSA-N 0.000 description 1
- HNJBEVLQSNELDL-UHFFFAOYSA-N pyrrolidin-2-one Chemical compound O=C1CCCN1 HNJBEVLQSNELDL-UHFFFAOYSA-N 0.000 description 1
- 125000000719 pyrrolidinyl group Chemical group 0.000 description 1
- 125000002294 quinazolinyl group Chemical group N1=C(N=CC2=CC=CC=C12)* 0.000 description 1
- SBYHFKPVCBCYGV-UHFFFAOYSA-N quinuclidine Chemical compound C1CC2CCN1CC2 SBYHFKPVCBCYGV-UHFFFAOYSA-N 0.000 description 1
- 150000003254 radicals Chemical class 0.000 description 1
- 238000010992 reflux Methods 0.000 description 1
- 230000026267 regulation of growth Effects 0.000 description 1
- 230000004044 response Effects 0.000 description 1
- 229910052703 rhodium Inorganic materials 0.000 description 1
- 210000003705 ribosome Anatomy 0.000 description 1
- 230000007017 scission Effects 0.000 description 1
- 125000002914 sec-butyl group Chemical group [H]C([H])([H])C([H])([H])C([H])(*)C([H])([H])[H] 0.000 description 1
- 239000000741 silica gel Substances 0.000 description 1
- 229910002027 silica gel Inorganic materials 0.000 description 1
- 229910052710 silicon Inorganic materials 0.000 description 1
- 239000010703 silicon Substances 0.000 description 1
- 125000003808 silyl group Chemical group [H][Si]([H])([H])[*] 0.000 description 1
- 239000002002 slurry Substances 0.000 description 1
- 102000030938 small GTPase Human genes 0.000 description 1
- 108060007624 small GTPase Proteins 0.000 description 1
- 150000003384 small molecules Chemical class 0.000 description 1
- 239000011734 sodium Substances 0.000 description 1
- 229910052708 sodium Inorganic materials 0.000 description 1
- 238000002415 sodium dodecyl sulfate polyacrylamide gel electrophoresis Methods 0.000 description 1
- 239000001488 sodium phosphate Substances 0.000 description 1
- 229910000162 sodium phosphate Inorganic materials 0.000 description 1
- 230000003595 spectral effect Effects 0.000 description 1
- 238000004611 spectroscopical analysis Methods 0.000 description 1
- 230000000087 stabilizing effect Effects 0.000 description 1
- 238000010186 staining Methods 0.000 description 1
- 239000012086 standard solution Substances 0.000 description 1
- 239000007858 starting material Substances 0.000 description 1
- 239000000758 substrate Substances 0.000 description 1
- 125000000472 sulfonyl group Chemical group *S(*)(=O)=O 0.000 description 1
- 239000006228 supernatant Substances 0.000 description 1
- 125000000999 tert-butyl group Chemical group [H]C([H])([H])C(*)(C([H])([H])[H])C([H])([H])[H] 0.000 description 1
- ILMRJRBKQSSXGY-UHFFFAOYSA-N tert-butyl(dimethyl)silicon Chemical group C[Si](C)C(C)(C)C ILMRJRBKQSSXGY-UHFFFAOYSA-N 0.000 description 1
- 239000012085 test solution Substances 0.000 description 1
- RAOIDOHSFRTOEL-UHFFFAOYSA-N tetrahydrothiophene Chemical compound C1CCSC1 RAOIDOHSFRTOEL-UHFFFAOYSA-N 0.000 description 1
- CZDYPVPMEAXLPK-UHFFFAOYSA-N tetramethylsilane Chemical compound C[Si](C)(C)C CZDYPVPMEAXLPK-UHFFFAOYSA-N 0.000 description 1
- 125000004001 thioalkyl group Chemical group 0.000 description 1
- BRNULMACUQOKMR-UHFFFAOYSA-N thiomorpholine Chemical compound C1CSCCN1 BRNULMACUQOKMR-UHFFFAOYSA-N 0.000 description 1
- IBBLKSWSCDAPIF-UHFFFAOYSA-N thiopyran Chemical compound S1C=CC=C=C1 IBBLKSWSCDAPIF-UHFFFAOYSA-N 0.000 description 1
- 230000007704 transition Effects 0.000 description 1
- 238000002054 transplantation Methods 0.000 description 1
- 125000004044 trifluoroacetyl group Chemical group FC(C(=O)*)(F)F 0.000 description 1
- 125000002023 trifluoromethyl group Chemical group FC(F)(F)* 0.000 description 1
- 125000000026 trimethylsilyl group Chemical group [H]C([H])([H])[Si]([*])(C([H])([H])[H])C([H])([H])[H] 0.000 description 1
- RYFMWSXOAZQYPI-UHFFFAOYSA-K trisodium phosphate Chemical compound [Na+].[Na+].[Na+].[O-]P([O-])([O-])=O RYFMWSXOAZQYPI-UHFFFAOYSA-K 0.000 description 1
- 229910052722 tritium Inorganic materials 0.000 description 1
- 125000002221 trityl group Chemical group [H]C1=C([H])C([H])=C([H])C([H])=C1C([*])(C1=C(C(=C(C(=C1[H])[H])[H])[H])[H])C1=C([H])C([H])=C([H])C([H])=C1[H] 0.000 description 1
- 238000002211 ultraviolet spectrum Methods 0.000 description 1
- 125000000391 vinyl group Chemical group [H]C([*])=C([H])[H] 0.000 description 1
Images
Classifications
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07D—HETEROCYCLIC COMPOUNDS
- C07D209/00—Heterocyclic compounds containing five-membered rings, condensed with other rings, with one nitrogen atom as the only ring hetero atom
- C07D209/56—Ring systems containing three or more rings
- C07D209/58—[b]- or [c]-condensed
- C07D209/62—Naphtho [c] pyrroles; Hydrogenated naphtho [c] pyrroles
- C07D209/66—Naphtho [c] pyrroles; Hydrogenated naphtho [c] pyrroles with oxygen atoms in positions 1 and 3
-
- C—CHEMISTRY; METALLURGY
- C09—DYES; PAINTS; POLISHES; NATURAL RESINS; ADHESIVES; COMPOSITIONS NOT OTHERWISE PROVIDED FOR; APPLICATIONS OF MATERIALS NOT OTHERWISE PROVIDED FOR
- C09B—ORGANIC DYES OR CLOSELY-RELATED COMPOUNDS FOR PRODUCING DYES, e.g. PIGMENTS; MORDANTS; LAKES
- C09B57/00—Other synthetic dyes of known constitution
- C09B57/04—Isoindoline dyes
-
- G—PHYSICS
- G01—MEASURING; TESTING
- G01N—INVESTIGATING OR ANALYSING MATERIALS BY DETERMINING THEIR CHEMICAL OR PHYSICAL PROPERTIES
- G01N33/00—Investigating or analysing materials by specific methods not covered by groups G01N1/00 - G01N31/00
- G01N33/48—Biological material, e.g. blood, urine; Haemocytometers
- G01N33/50—Chemical analysis of biological material, e.g. blood, urine; Testing involving biospecific ligand binding methods; Immunological testing
- G01N33/53—Immunoassay; Biospecific binding assay; Materials therefor
- G01N33/531—Production of immunochemical test materials
- G01N33/532—Production of labelled immunochemicals
- G01N33/533—Production of labelled immunochemicals with fluorescent label
Definitions
- Fluorescence is the result of a three-stage process that occurs when certain molecules absorb energy.
- the three stages comprise: 1) excitation; 2) excited-state lifetime; and 3) fluorescence emission.
- excitation a photon of a certain energy is absorbed by the fluorophore.
- the fluorophore is initially in its ground state (S 0 ). Absorption of the photon causes that fluorophore to become excited. The energy of the absorbed photon is transferred to an electron. The electron is transferred to a higher energy state.
- S 1′ excited electronic singlet state
- the excited state of the fluorophore exists for a finite time, typically 10 ⁇ 8 to 10 ⁇ 9 seconds.
- the fluorophore changes in its translational, vibrational, and electronic energy states, and is subject to interactions with its molecular environment.
- the excited fluorophore releases energy and returns to the ground state, S 0 , by fluorescence emission.
- Other processes such as fluorescence energy transfer, intersystem crossing, and collisional quenching may also depopulate S 1 .
- the quantum yield is a measure of the efficiency of fluorescence in competition with other processes such as fluorescence energy transfer, intersystem crossing, and collisional quenching.
- a photon of energy hv (where h is Planck's constant and v is the frequency of the photon) is emitted, returning the fluorophore to its ground state S 0 .
- the energy of the emitted photon is lower than the energy of the photon absorbed during the excitation stage.
- the difference in energy can be attributed to dissipation through processes during the excited-state lifetime, such processes include fluorescence energy transfer, intersystem crossing, and collisional quenching.
- the difference in energy of the absorbed photon and the emitted photon is called the Stokes shift.
- the Stokes shift is fundamental to the sensitivity of fluorescence techniques because it allows emission photons to be detected against a low background, and at a different wavelength than the excitation photons.
- Fluorescent molecules can be used in single molecule spectroscopy, liquid crystal displays, light emitting diodes, solar energy collectors, and laser active media. Fluorescent molecules whose spectra or quantum yields are sensitive to their environments are valuable as fluorescent dyes and in the study of heterogeneous media, organized media, and biological media.
- the present invention provides compounds that can be used to monitor biological interactions continuously with a fluorescence readout.
- the compounds of the present invention are environment-sensitive fluorophores which have spectroscopic behavior that is dependent on the physicochemical properties of the surrounding environment.
- the compounds of the present invention can be used in biochemical research to monitor ions, small molecules, and biological processes such as protein folding, protein-protein interactions and phosphorylation events.
- Environment-sensitive fluorophores are a special class of chromophores that have spectroscopic behavior that is dependent on the physicochemical properties of the surrounding environment.
- Solvatochromic fluorophores display sensitivity to the polarity of the local environment. These molecules exhibit a low quantum yield in aqueous solution, but become highly fluorescent in nonpolar solvents or when bound to hydrophobic sites in proteins or membranes. Examples of solvatochromic fluorophores include 2-propionyl-6-dimethylaminonaphthalene (PRODAN) (Weber et al. Biochemistry 1979, 18, 3075-3078; Cohen et al.
- PRODAN 2-propionyl-6-dimethylaminonaphthalene
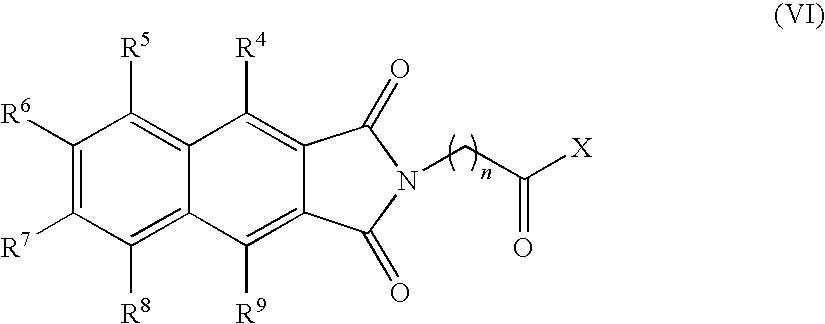
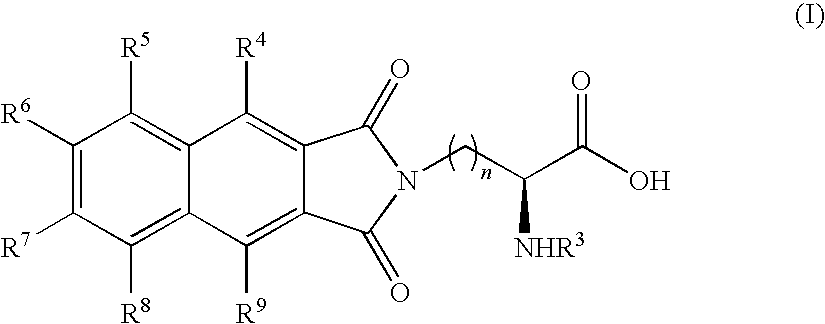
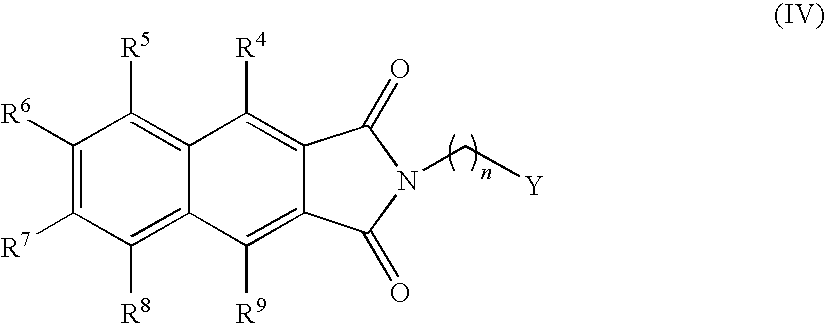
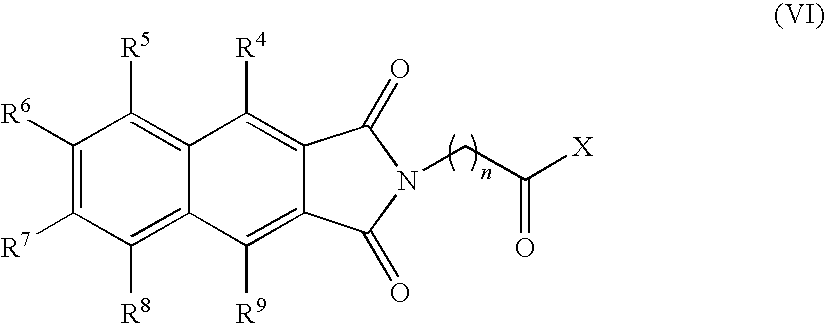
- the present invention provides novel compounds of the formula (I):
- n, R 3 , R 4 , R 5 , R 6 , R 7 , R 8 and R 9 are as defined below.
- One especially preferred compound (III) referred to as Dap(6DMN) is disclosed.
- the present invention also provides peptides containing the compound (I) of the present invention.
- the present invention also provides a method for probing biological interactions using peptides containing the compound (I).
- FIG. 1 are fluorescence titrations of peptides (Crk-bp and Crk-bp2) binding to GST-Crk SH2 domain.
- FIG. 2 are fluorescence titrations of peptide Abl-bp with various GST SH2 domains.
- FIG. 3 are fluorescence titrations of peptide Abl-bp2 with various GST SH2 domains.
- Abl-bp2 (20 ⁇ M in PBS buffer, pH 7.5) with GST-Abl SH2.
- FIG. 4 are graphs depicting fluorescence titrations of peptides with non-target SH2 domains. a) Crk-bp with GST-Abl SH2, b) Crk-bp2 with GST-Abl SH2, c) Abl-bp with GST-Crk SH2, d) Abl-bp2 with GST-Crk SH2.
- FIG. 5 are plots of integrated fluorescence vs. absorbance used to calculate the quantum yield ( ⁇ ) of Ac-Gly-Dap(6DMN)-Gln-pTyr-Glu-Asn-Val-Gln-Ser-(CONH 2 ).
- the quantum yield is proportional to the gradient of the plot. The absolute value is calculated using a standard sample, quinine sulfate.
- FIG. 6 are UV absorption and fluorescence emission spectra of compound VIII, 6DMN-GlyOMe. a) UV absorption spectra (40 ⁇ M) in methanol. b) Fluorescence emission spectra in different solvents. Labels show maximum emission wavelengths for selected solvents (toluene, chloroform and methanol).
- FIG. 7 are Lippert-Mataga plots for compound VIII, 6DMN-GlyOMe.
- ⁇ A ⁇ F ( ⁇ ⁇ ) is the difference in the maximum absorption and emission wavelengths, expressed in wavenumbers.
- ⁇ A ⁇ F ( ⁇ ⁇ ) was measured from a series of 1,4-dioxane/acetonitrile mixtures.
- ⁇ f was calculated based upon the composition of the 1,4-dioxane/acetonitrile mixture using the dielectric constant and refraction index of 1,4-dioxane and acetonitrile.
- the slope corresponds to ⁇ e ⁇ g ( ⁇ ) which is the difference between the dipole moments of the excited and ground states for 6DMN-GlyOMe. The greater the change in the dipole moment, the more sensitive the fluorophore.
- ⁇ f was calculated for each solvent using the dielectric constant and refraction index of the corresponding solvent.
- the line corresponds to the best linear fit to the data, excluding the values obtained for protic solvents (methanol, isopropanol and ethanol, closed circles), indicating that the Stokes shift is proportional to the orientation polarizability.
- Protic solvents methanol, isopropanol and ethanol
- the present invention is directed to compounds and salts thereof, compositions and methods useful in monitoring biological interactions continuously with a fluorescence readout.
- Alkyl by itself or as part of another substituent refers to a hydrocarbon group which may be linear, cyclic, or branched or a combination thereof having the number of carbon atoms designated (i.e., C 1-8 means one to eight carbon atoms).
- alkyl groups include methyl, ethyl, n-propyl, isopropyl, n-butyl, t-butyl, isobutyl, sec-butyl, cyclohexyl, cyclopentyl, (cyclohexyl)methyl, cyclopropylmethyl, bicyclo[2.2.1]heptane, bicyclo[2.2.2]octane, etc.
- Alkyl groups can be substituted or unsubstituted, unless otherwise indicated. Examples of substituted alkyl include haloalkyl, thioalkyl, aminoalkyl, and the like.
- Alkoxy refers to —O-alkyl. Examples of an alkoxy group include methoxy, ethoxy, n-propoxy, etc.
- Alkenyl refers to an unsaturated hydrocarbon group which may be linear, cyclic or branched or a combination thereof. Alkenyl groups with 2-8 carbon atoms are preferred. The alkenyl group may contain 1, 2 or 3 carbon-carbon double bonds. Examples of alkenyl groups include ethenyl, n-propenyl, isopropenyl, n-but-2-enyl, n-hex-3-enyl, cyclohexenyl, cyclopentenyl and the like. Alkenyl groups can be substituted or unsubstituted, unless otherwise indicated.
- Alkynyl refers to an unsaturated hydrocarbon group which may be linear, cyclic or branched or a combination thereof. Alkynyl groups with 2-8 carbon atoms are preferred. The alkynyl group may contain 1, 2 or 3 carbon-carbon triple bonds. Examples of alkynyl groups include ethynyl, n-propynyl, n-but-2-ynyl, n-hex-3-ynyl and the like. Alkynyl groups can be substituted or unsubstituted, unless otherwise indicated.
- Aryl refers to a polyunsaturated, aromatic hydrocarbon group having a single ring (monocyclic) or multiple rings (bicyclic), which can be fused together or linked covalently.
- Aryl groups with 6-10 carbon atoms are preferred, where this number of carbon atoms can be designated by C 6-10 , for example.
- Examples of aryl groups include phenyl and naphthalene-1-yl, naphthalene-2-yl, biphenyl and the like.
- Aryl groups can be substituted or unsubstituted, unless otherwise indicated.
- Fluorescence encompasses the release of fluorescent energy. Less broadly, the term “fluorescence” refers to fluorescent emission, the rate of change of fluorescence over time (i.e., fluorescence lifetime), fluorescence polarization, fluorescence anisotropy, and fluorescence resonance energy transfer. See Eftink, M. R., Biophysical J. 66:482-501 (1994).
- Fluorescence probe molecule refers to a compound of the present invention.
- the compound after excitement by light of a defined wavelength or defined range of wavelengths, is capable of emitting fluorescent energy.
- the fluorescent molecule or a compound may be capable of binding to a peptide, protein, membrane or receptor.
- Halo or “halogen”, by itself or as part of a substituent refers to a chlorine, bromine, iodine, or fluorine atom.
- Haloalkyl refers to a monohaloalkyl or polyhaloalkyl group, most typically substituted with from 1-3 halogen atoms. Examples include 1-chloroethyl, 3-bromopropyl, trifluoromethyl and the like.
- Heterocyclyl refers to a saturated or unsaturated non-aromatic ring containing at least one heteroatom (typically 1 to 5 heteroatoms) selected from nitrogen, oxygen or sulfur.
- the heterocyclyl ring may be monocyclic or bicyclic.
- these groups contain 0-5 nitrogen atoms, 0-2 sulfur atoms and 0-2 oxygen atoms. More preferably, these groups contain 0-3 nitrogen atoms, 0-1 sulfur atoms and 0-1 oxygen atoms.
- heterocycle groups include pyrrolidine, piperidine, imidazolidine, pyrazolidine, butyrolactam, valerolactam, imidazolidinone, hydantoin, dioxolane, phthalimide, piperidine, 1,4-dioxane, morpholine, thiomorpholine, thiomorpholine-S-oxide, thiomorpholine-S,S-dioxide, piperazine, pyran, pyridone, 3-pyrroline, thiopyran, pyrone, tetrahydrofuran, tetrahydrothiophene, quinuclidine and the like.
- Preferred heterocyclic groups are monocyclic, though they may be fused or linked covalently to an aryl or heteroaryl ring system.
- heterocyclic groups may be represented by formula (AA) below:
- R a , R b , R c , R d , R e , R f , and R g are independently selected from the group consisting of hydrogen, halogen, unsubstituted or substituted C 1-8 alkyl, unsubstituted or substituted C 2-8 alkenyl, unsubstituted or substituted C 2-8 alkynyl, —COR h , —CO 2 R h , —CONR h R i , —NR h COR i , —SO 2
- R a +R b +R c +R d groups that are other than hydrogen is 0, 1 or 2.
- R a , R b , R c , R d , R e , R f , and R g are independently selected from the group consisting of hydrogen, halogen, unsubstituted or substituted C 1-8 alkyl, —COR h , —CO 2 R h , —CONR h R h , —NR h COR h , —SO 2 R h , —SO 2 NR h R i , NSO 2 R h R i , —NR h R i , and —OR h , wherein R h and R i are independently selected from the group consisting of hydrogen and unsubstituted C 1-8 alkyl and wherein the aliphatic portions of each of the R a , R b , R c
- R a , R b , R c , R d , R e , R f , and R g are independently hydrogen or C 1-4 alkyl. In another preferred embodiment, at least three of R a , R b , R c , R d , R e , R f , and R g are hydrogen.
- Heteroaryl refers to an aromatic group containing at least one heteroatom, where the heteroaryl group may be monocyclic or bicyclic. Examples include pyridyl, pyridazinyl, pyrazinyl, pyrimidinyl, triazinyl, quinolinyl, quinoxalinyl, quinazolinyl, cinnolinyl, phthalazinyl, benzotriazinyl, purinyl, benzimidazolyl, benzopyrazolyl, benzotriazolyl, benzisoxazolyl, isobenzofuryl, isoindolyl, indolizinyl, benzotriazinyl, thienopyridinyl, thienopyrimidinyl, pyrazolopyrimidinyl, imidazopyridines, benzothiazolyl, benzofuranyl, benzothienyl, indolyl, quinolyl, is
- Preferred heteroaryl groups are those having at least one aryl ring nitrogen atom, such as quinolinyl, quinoxalinyl, purinyl, benzimidazolyl, benzopyrazolyl, benzotriazolyl, benzothiazolyl, indolyl, quinolyl, isoquinolyl and the like.
- Preferred 6-ring heteroaryl systems include pyridyl, pyridazinyl, pyrazinyl, pyrimidinyl, triazinyl and the like.
- Preferred 5-ring heteroaryl systems include isothiazolyl, pyrazolyl, imidazolyl, thienyl, furyl, triazolyl, tetrazolyl, oxazolyl, isoxazolyl, oxadiazolyl, thiadiazolyl, pyrrolyl, thiazolyl and the like.
- Heterocyclyl and heteroaryl can be attached at any available ring carbon or heteroatom.
- Each heterocyclyl and heteroaryl may have one or more rings. When multiple rings are present, they can be fused together or linked covalently.
- Each heterocyclyl and heteroaryl must contain at least one heteroatom (typically 1 to 5 heteroatoms) selected from nitrogen, oxygen or sulfur. Preferably, these groups contain 0-5 nitrogen atoms, 0-2 sulfur atoms and 0-2 oxygen atoms. More preferably, these groups contain 0-3 nitrogen atoms, 0-1 sulfur atoms and 0-1 oxygen atoms. Heterocyclyl and heteroaryl groups can be substituted or unsubstituted, unless otherwise indicated.
- the substitution may be on a carbon or heteroatom.
- the substitution is oxo ( ⁇ O or —O ⁇ )
- the resulting group may have either a carbonyl (—C(O)—) or a N-oxide (—N + —O ⁇ ).
- Suitable substituents for substituted alkyl include halogen, —CN, —CO 2 R′, —C(O)R′, —C(O)NR′R′′, oxo ( ⁇ O or —O ⁇ ), —OR′, —OC(O)R′, —OC(O)NR′R′′, —NO 2 , —NR′C(O)R′′, —NR′′′C(O)NR′R′′, —NR′R′′, —NR′CO 2 R′′, —NR′S(O)R′′, —NR′S(O) 2 R′′′, NR′′′S(O)NR′R′′, —NR′′′S(O) 2 NR′R′′, —SR′, —S(O)R′, —S(O) 2 R′, —S(O) 2 NR′R′′, —NR′—C(NHR′′) ⁇ NR′′′, —SiR′R′′R′′′, —
- Suitable substituents for substituted ring include halogen, —CN, —CO 2 R′, —C(O)R′, —C(O)NR′R′′, oxo ( ⁇ O or —O ⁇ ), —OR′, —OC(O)R′, —OC(O)NR′R′′, —NO 2 , —NR′C(O)R′′, —NR′′′C(O)NR′R′′, —NR′R′′, —NR′CO 2 R′′, —NR′S(O)R′′, —NR′S(O) 2 R′′, —NR′′′S(O)NR′R′′, —NR′′′S(O) 2 NR′R′′, —SR′, —S(O)R′, —S(O) 2 R′, —S(O) 2 NR′R′′, —NR′—C(NHR′′) ⁇ NR′′′, —SiR′R′′R′′′, —N
- R′, R′′ and R′′′ each independently refer to a variety of groups including hydrogen, substituted or unsubstituted C 1-8 alkyl, substituted or unsubstituted C 2-8 alkenyl, substituted or unsubstituted C 2 -8 alkynyl, substituted or unsubstituted aryl, substituted or unsubstituted heteroaryl, substituted or unsubstituted heterocyclyl, substituted or unsubstituted arylalkyl, substituted or unsubstituted aryloxyalkyl.
- R′ and R′′ When R′ and R′′ are attached to the same nitrogen atom, they can be combined with the nitrogen atom to form a 3-, 4-, 5-, 6-, or 7-membered ring (for example, —NR′R′′ includes 1-pyrrolidinyl and 4-morpholinyl). Furthermore, R′ and R′′, R′′ and R′′′, or R′ and R′′′ may together with the atom(s) to which they are attached, form a substituted or unsubstituted 5-, 6- or 7-membered ring.
- Heteroatom is meant to include oxygen (O), nitrogen (N), sulfur (S) and silicon (Si).
- N-protecting group refers to an easily removable group which is known in the art to protect an amino group against undesirable reaction during synthetic procedures and to be selectively removable. The use of N-protecting groups is well known in the art for protecting groups against undesirable reactions during a synthetic procedure and many such protecting groups are known. Commonly used N-protecting groups are known to those skilled in the art, examples of which are disclosed in Greene and Wuts, Protective Groups in Organic Synthesis, 2nd ed.; John Wiley & Sons, New York, 1991).
- N-protecting groups include, but are not limited to, acyl groups including formyl, acetyl (Ac), trifluoroacetyl, trichloroacetyl, propionyl, pivaloyl, t-butylacetyl, acylisothiocyanate, aminocaproyl, benzoyl and the like; acyloxy groups, including t-butyloxycarbonyl (Boc), benzyloxycarbonyl (Cbz), 9-fluorenylmethoxycarbonyl (Fmoc), p-methoxybenzyloxycarbonyl, methoxycarbonyl, ethoxycarbonyl, allyloxycarbonyl and the like; sulfonyl groups such as benzenesulfonyl, p-toluenesulfonyl and the like; arylalkyl groups such as benzyl, triphenylmethyl, benzyloxymethyl
- alkyl e.g., “alkyl,” “aryl,” “heteroaryl” etc.
- substituents include alkyl, aryl, heteroaryl, heterocyclyl, halogen, alkoxy, oxygen, and nitrogen.
- Certain compounds of the present invention can exist in unsolvated forms as well as solvated forms, including hydrated forms. In general, both solvated forms and unsolvated forms are intended to be encompassed within the scope of the present invention. Certain compounds of the present invention may exist in multiple crystalline or amorphous forms (i.e., as polymorphs). In general, all physical forms are equivalent for the uses contemplated by the present invention and are intended to be within the scope of the present invention.
- Certain compounds of the present invention possess asymmetric carbon atoms (optical centers) or double bonds; the racemates, diastereomers, geometric isomers and individual isomers (e.g., separate enantiomers) are all intended to be encompassed within the scope of the present invention.
- the compounds of the present invention may also contain unnatural proportions of atomic isotopes at one or more of the atoms that constitute such compounds.
- the compounds may be radiolabeled with radioactive isotopes, such as for example tritium ( 3 H), iodine-125 ( 125 I) or carbon-14 ( 14 C). All isotopic variations of the compounds of the present invention, whether radioactive or not, are intended to be encompassed within the scope of the present invention.
- biological interactions encompasses the interaction of a compound or molecule with a target molecule.
- Protein and “peptide”, as used herein, are synonymous.
- the term “unfolding” encompasses any change in structure due to heating.
- the term “unfolding” refers to the transition of from the liquid crystalline state to the molten globule state. In the molten globule state, tertiary and quaternary structure has been altered, relative to the native state of the protein, and at least some secondary structure remains intact.
- the term “unfolding” also encompasses loss of crystalline ordering of amino acid side-chains, secondary, tertiary or quaternary structure.
- the term “unfolding” also encompasses formation of a random coil.
- “Folding” and “refolding,” and “renaturing” refer to the acquisition of the correct amino acid side-chain ordering, secondary, tertiary, or quaternary structure, of a protein or a nucleic acid, which affords the full chemical and biological function of the biomolecule.
- target molecule encompasses peptides, proteins, nucleic acids, ions, and other receptors.
- the term encompasses both enzymes, and proteins which are not enzymes.
- the term encompasses monomeric and multimeric proteins. Multimeric proteins may be homomeric or heteromeric.
- the term encompasses nucleic acids comprising at least two nucleotides, such as oligonucleotides. Nucleic acids can be single-stranded, double-stranded, or triple-stranded.
- the term encompasses a nucleic acid which is a synthetic oligonucleotide, a portion of a recombinant DNA molecule, or a portion of chromosomal DNA.
- target molecule also encompasses portions of peptides, proteins, and other receptors which are capable of acquiring secondary, tertiary, or quaternary structure through folding, coiling or twisting.
- the target molecule may be substituted with substituents including, but not limited to, cofactors, coenzymes, prosthetic groups, lipids, oligosaccharides, or phosphate groups.
- target molecule and “receptor” are synonymous.
- target molecules examples include those disclosed in Faisst, S. et al., Nucleic Acids Research 20:3-26 (1992); Pimentel, E., Handbook of Growth Factors , Volumes I-III, CRC Press, (1994); Gilman, A. G. et al., The Pharmacological Basis of Therapeutics , Pergamon Press (1990); Lewin, B., Genes V, Oxford University Press (1994); Roitt, I., Essential Immunology , Blackwell Scientific Publ. (1994); Shimizu, Y., Lymphocyte Adhesion Molecules , R G Austin (1993); Hyams, J. S.
- contacting a target molecule refers broadly to placing the target molecule in solution with the molecule to be screened for binding or with the condition(s) to be tested for stabilizing the target molecule. Less broadly, contacting refers to the turning, swirling, shaking or vibrating of a solution of the target molecule and the molecule to be screened for binding. More specifically, contacting refers to the mixing of the target molecule with the molecule to be tested for binding. Mixing can be accomplished, for example, by repeated uptake and discharge through a pipette tip, either manually or using an automated pipetting device. Preferably, contacting refers to the equilibration of binding between the target molecule and the molecule to be tested for binding. Contacting can occur in the container, infra, or before the target molecule and the molecule to be screened are placed in the container.
- the target molecule may be contacted with a nucleic acid prior to being contacted with the molecule to be screened for binding.
- the target molecule may be complexed with a peptide prior to being contacted with the molecule to be screened for binding.
- the target molecule may be phosphorylated or dephosphorylated prior to being contacted with the molecule to be screened for binding.
- a carbohydrate moiety may be added to the target molecule before the target molecule is contacted with the molecule to be screened for binding.
- a carbohydrate moiety may be removed from the target molecule before the target molecule is contacted with the molecule to be screened for binding.
- tainer refers to any vessel or chamber in which the receptor and molecule to be tested for binding can be placed.
- the term “container” encompasses reaction tubes (e.g., test tubes, microtubes, vials, etc.).
- biological sample refers to the contents of a container.
- “Spectral emission,” “thermal change,” and “physical change” encompass the release of energy in the form of light or heat, the absorption of energy in the form or light or heat, changes in turbidity and changes in the polar properties of light.
- the terms refer to fluorescent emission, fluorescent energy transfer, absorption of ultraviolet or visible light, changes in the polarization properties of light, changes in the polarization properties of fluorescent emission, changes in the rate of change of fluorescence over time (i.e., fluorescence lifetime), changes in fluorescence anisotropy, changes in fluorescence resonance energy transfer, changes in turbidity, and changes in enzyme activity.
- the terms refer to fluorescence, and more preferably to fluorescence emission.
- Fluorescence emission can be intrinsic to a protein or can be due to a fluorescence reporter molecule.
- fluorescence techniques to monitor protein unfolding is well known to those of ordinary skill in the art. For example, see Eftink, M. R., Biophysical J. 66:482-501 (1994).
- Biochemical conditions encompass any component of a physical, chemical, or biochemical reaction. Specifically, the term refers to conditions of temperature, pressure, protein concentration, pH, ionic strength, salt concentration, time, electric current, potential difference, concentrations of cofactor, coenzyme, oxidizing agents, reducing agents, detergents, metal ion, ligands, or glycerol.
- the compounds of the present invention undergo enhanced fluorescence in nonpolar environments as compared to polar environments.
- nonpolar environments include nonpolar solvents and hydrophobic proteins or membranes.
- R 4 , R 5 , R 6 , R 7 , R 8 , and R 9 are each independently hydrogen, fluorine, or alkyl, with the proviso that one of R 6 or R 7 is —NR 1 R 2 , —OR 1 , or —SR 1 ;
- R 1 and R 2 are each independently hydrogen or alkyl
- R 1 and R 2 together with the nitrogen to which they are attached, may form a substituted or unsubstituted 5- or 6-membered ring; wherein said nitrogen is not conjugated with said 5- or 6-membered ring;
- R 3 is hydrogen or a N-protecting group
- n 1, 2, 3 or 4.
- R 6 or R 7 is —NR 1 R 2 . More preferably, R 7 is —NR 1 R 2 .
- R 1 or R 2 is alkyl. More preferably, R 1 and R 2 are both alkyl. Even more preferably, R 1 and R 2 are both methyl, ethyl or propyl.
- R 1 and R 2 together with the nitrogen to which they are attached are pyrrolidinyl, piperidinyl, or morpholinyl.
- R 3 is a N-protecting group. More preferably R 3 is Boc, Cbz, or Fmoc. Even more preferably R 3 is Fmoc.
- the compound of the present invention can have the formula (II):
- the compound of the present invention can have the formula (IV):
- Y is NH 2 or C(O)X
- the compound of the present invention can have the formula (V):
- the compound of the present invention can have the formula (VI):
- the compound of the present invention can have the formula (VII):
- the compound can have the formula (VIII):
- the compound in another embodiment, can be a peptide containing the formula (I):
- the compound can be a peptide containing the formula (II):
- R 1 , R 2 , and R 3 are each described above for formula (I).
- the compound can be a peptide containing the formula (III):
- R 3 is described above for formula (I).
- compound (III) is referred to herein as Dap(6DMN).
- compound (III) is herein referred to as Fmoc-Dap(6DMN).
- Compounds of the present invention are useful as fluorescence probe molecules in applications wherein fluorescence probes are known to be useful.
- a compound of the present invention is added to a sample to be probed.
- the sample comprising the compound is then exposed to a light source.
- the light source produces light that is limited to a range of wavelengths.
- the range of wavelengths is between about 360 and about 410 nanometers (nm), preferably between about 370 and about 390 nm, most preferably between about 375 and about 380 nm.
- the compound of the present invention is fluorescent and emits fluorescent energy.
- the emitted fluorescent energy is detected using methods well known in the art.
- the intensity and wavelength of the emitted fluorescent energy provides information about the sample.
- the emitted fluorescent energy preferably has a range of wavelengths between about 480 and about 640 nm, preferably between about 560 and about 590 nm.
- the fluorescence of a molecule is defined by the quantum yield.
- the quantum yield is the ratio of the photons absorbed by the compound to the photons emitted through fluorescence by the compound.
- Compounds of the present invention have quantum yields that are preferably low in aqueous solutions and high in non-polar environments. Quantum yields range from about 0.001 and about 0.1, preferably between about 0.001 and about 0.005 for aqueous solutions, preferably between about 0.2 and about 0.7 for non-polar environment.
- Fluorescence can also be evaluated by determining the dipole moment change between the ground and excited state.
- the change in the dipole moment can be estimated from a plot of the Stokes shift vs. the orientation polarizability, known as a Lippert-Mataga plot.
- a Lippert-Mataga plot (Lippert, V. E. Z. Elektrochem. 1957, 61, 962-975; Mataga, N.; Kaifu, Y.; Koizumi, M. Bull. Chem. Soc. Jpn. 1956, 29, 465-470.)
- the relationship between the Stokes shift ( ⁇ ⁇ ) and the orientation polarizability ( ⁇ f) is expressed by the following equation:
- ⁇ is the velocity of light
- h Plank's constant
- c is the solvent dielectric constant
- n is the solvent refraction index
- ⁇ e ⁇ g is the difference between the dipole moments of the excited and the ground states respectively ( ⁇ )
- a 0 is the radius of the Onsager cavity around the compound.
- the magnitude of ⁇ correlates to the sensitivity of the fluorescent probe, where larger changes in the dipole moment correlate to greater sensitivity.
- Fluorescence is sensitive to the pH of the surrounding environment.
- Compounds of the present invention are useful as fluorescence probes in the pH range from about 4 to about 8.
- Bio interactions play important roles in the sequence and mechanisms of action of various cellular processes and signal pathways.
- the time course, nature, and sequence of the different cellular processes can be elucidated by in situ observation using the compounds of the present invention.
- Specific inhibitors and/or activators of the cellular processes and signal pathways being studied may optionally be used in addition to compounds of the present invention.
- Bio interactions comprise the interaction of a compound or molecule with a target molecule.
- target molecules include peptides, proteins, enzymes, nucleic acids, ions, and other receptors; preferably metal ion chelators, proteases, polymerases, hydrolases, phosphatases, and kinases; more preferably protein domains, and protein domains of phosphatases and kinases.
- Proteins and protein-protein interactions play a central role in the various essential biochemical processes. For example, these interactions are evident in the interaction of hormones with their respective receptors, in the intracellular and extracellular signaling events mediated by proteins, in enzyme substrate interactions, in intracellular protein trafficking, in the formation of complex structures like ribosomes, viral coat proteins, and filaments, and in antigen-antibody interactions. These interactions are usually facilitated by the interaction of small regions within the proteins that can fold independently of the rest of the protein. These independent units are called protein domains. Abnormal or disease states can be the direct result of aberrant protein-protein interactions. Protein-protein interactions are also central to the mechanism of a virus recognizing its receptor on the cell surface as a prelude to infection. Identification of domains that interact with each other not only leads to a broader understanding of protein-protein interactions, but also aids in the design of inhibitors of these interactions.
- Phosphorylation-dependent peptide-protein interactions include phosphoserine peptides with 14-3-3, which is a protein involved in cell cycle control (Muslin, A. J., Tanner, J. W., Allen, P. M., Shaw, A. S. Cell 1996, 84, 889-897), and phosphotyrosine peptides with SH2 domains.
- SH2 domains are binding modules that are involved in tyrosine kinase signaling networks and recognize phosphotyrosine-containing peptide sequences. The phosphotyrosine binding is complemented by simultaneous peptide-protein interactions on the protein surface.
- SH2 domains include Abl SH2, Crk SH2, and C-terminal PI3K SH2 which can be expressed in bacteria as GST fusion proteins, which are referred to as GST-Abl SH2, GST-Crk SH2, and GST-PI3K SH2.
- Recognition sequences for SH2 domains comprise a phosphotyrosine residue and other amino acids.
- the recognition sequence is different for different SH2 domains.
- Amino acid recognition sequences for binding members of the SH2 domain family are disclosed in Songyang, Z. et al.; Cell 1993, 72, 767-778.
- the recognition sequence is pTyr-Asp-His-Pro.
- the recognition sequence is pTyr-Glu-Asn-Val.
- Compounds of the formula (I) are useful for studying the peptide-protein interactions on the protein surface of the SH2 domain.
- Compounds of formula (I) can be incorporated into peptides containing the desired SH2 recognition sequence.
- Table 1 shows peptides incorporating the Crk SH2 or Abl SH2 recognition sequences and Dap(6DMN) into the (+2) position relative to the phosphotyrosine residue.
- the peptides of Table 1 were incubated with targeted and nontargeted SH2 domains.
- the binding of peptides Crk-bp, Crk-bp2, Abl-bp, and Abl-bp2 to SH2 target domains can be studied by fluorescence titration as shown in FIGS. 1-4 .
- FIG. 1 ( a ) and ( b ) Fluorescence titrations of Crk-bp and Crk-bp2 with targeted GST-Crk SH2 are shown in FIG. 1 ( a ) and ( b ), respectively. Fluorescence titrations of Crk-bp and Crk-bp2 with non-targeted GST-Abl SH2 are shown in FIG. 4 ( a ) and ( b ), respectively.
- FIG. 1 ( c ) shows relative fluorescence emission intensities for Crk-bp and Crk-bp2 peptides, Crk-bp and Crk-bp2 bound to GST-Crk SH2 domain, and Crk-bp2 bound to non-target domains GST-Abl SH2 and GST-PKI3 SH2.
- the relative fluorescence emission intensities for non-target SH2 domains is notably less than for the targeted GST-Crk SH2 domain.
- Fluorescence titrations of Abl-bp with targeted GST-Abl SH2 is shown in FIG. 2 ( a ). Fluorescence titrations of Abl-bp with non-targeted GST-PI3K SH2 and GST-Abl SH2 are shown in FIG. 2 ( b ) and 4 ( c ), respectively. The relative fluorescence emission intensities for the Abl-bp peptide and aforementioned SH2 domain titrations are shown in FIG. 2 ( c ).
- FIG. 3 ( a ) Fluorescence titrations of Abl-bp2 with targeted GST-Abl SH2 is shown in FIG. 3 ( a ). Fluorescence titrations Abl-bp2 with non-targeted GST-PI3K SH2 and GST-Crk SH2 are shown in FIGS. 3 ( b ) and 4 ( d ), respectively. The relative fluorescence emission intensities for the Abl-bp2 peptide and aforementioned SH2 domain titrations are shown in FIG. 3 ( c ).
- Compounds of the present invention are also useful in a method of monitoring biological interactions, comprising providing a compound of the present invention, contacting a target molecule with the compound to form a biological sample, and monitoring the fluorescence of the biological sample.
- the compound is a peptide. More preferably, the compound is a peptide containing at least one of formula (I), (II), and (III).
- the monitoring step comprise contacting the compound with the one or more target molecules or different biochemical conditions, wherein the measuring step comprises exciting the compound of the present invention with light, and measuring the fluorescence.
- the concentration used will depend on the detection equipment. Typically, the concentration of the compound is from greater than about 0.1 nM.
- the compounds of formula (I) can be prepared according to the reaction as shown in Scheme 1, wherein, n, R 3 , R 4 , R 5 , R 6 , R 7 , R 8 and R 9 are as defined above.
- Anhydride (1) is stirred with an appropriate diamine (2) in a suitable solvent to form compound 3.
- the allyl protecting group of compound 3 can be removed under standard conditions known to those skilled in the art, examples of which are disclosed in Greene and Wuts, Protective Groups in Organic Synthesis” 2nd ed.; John Wiley & Sons, New York, 1991) to afford a compound of Formula I (4).
- the diamine compound 2 can be prepared from the corresponding diamino acid using standard conditions for protecting a carboxylic acid with an allyl protecting group known to those skilled in the art, examples of which are provided in Greene and Wuts.
- Anhydride 1 can be prepared by stirring diester 5 with an appropriate base in a suitable solvent.
- Compound 5 can be assembled by methods known to one skilled in the art such ring annulation methods or Diels-Alder reaction.
- the compounds of formula (IV) can be prepared by stirring anhydride (1) with an appropriate amine in a suitable solvent to form compounds of formula (IV), wherein, n, R 4 , R 5 , R 6 , R 7 , R 8 , R 9 and Y are as defined above.
- Y is NH 2
- the amine is diamino compound (6) as shown in Scheme 3 which is stirred with anhydride (1) to for compound (7).
- Diamines are commercially available from vendors such as Aldrich, for example.
- compound 7 may be prepared by stirring anhydride (1) with a diamine where one of the nitrogens is protected with a suitable N-protecting group. After formation of the imide bond, the N-protecting group can be removed to afford compound (7).
- Methods for introduction and removal of N-protecting groups are known to those skilled in the art, examples of which are disclosed in Greene and Wuts, Protective Groups in Organic Synthesis, 2nd ed.; John Wiley & Sons, New York, 1991
- Compound (10) where X is halogen or alkoxy can be prepared under standard conditions acyl halide ( Org. Syn. Coll. Vol. 1 1941, 12, 147 ; Org. Syn. Coll. Vol. 3 1955, 169, 490, 547, 555, 712 ; Org. Syn. Coll. Vol 4 1963, 154, 263, 339, 715, 739) or esterification ( Tetrahedron. 1980, 36, 2409) conditions.
- Peptides containing the formula (I) can be prepared using standard peptide coupling techniques known to those skilled in the art. Examples are provided in Bodansky, M. Peptide Chemistry a Practical Textbook, 2nd ed.; Springer-Verlag, Berlin, 1993. Dap(6DMN), can incorporated into peptides using Fmoc-Dap(6DMN) and standard Fmoc solid-phase peptide synthesis.
- the sensor of the present invention can be used in a method for detecting biological interactions.
- the method of the present invention comprises providing a peptide incorporating an amino acid of the formula (I), contacting a target molecule with the peptide to form a biological sample, and monitoring the fluorescence of the biological sample.
- Electrospray Ionization Mass Spectrometry was performed on a PerSeptive Biosystems MarinerTM Biospectrometry Workstation (Turbo Ion Source). Fluorescence spectroscopy measurements were made using a Fluoromax-P spectrofluorimeter controlled by the DataMax 2.20 software, and coupled to a NesLab RTE-111 water bath for temperature control. All measurements were made at 20° C.
- Peptide synthesis was carried using standard Fmoc-based solid phase peptide synthesis (SPPS) protocols on a 0.05 to 0.1 mmol scale using a 0.21 mmol/g loading PAL-PEG-PS solid support.
- Amino acids were coupled in three-fold excess using a mixture of 0.2 M HBTU/0.2 M HOBt in DMF as activating agents. Each amino acid was activated for two minutes with the HBTU/HOBt mixture (1 eq.) and diisopropylethylamine (DIPEA), 0.195 M in DMF (1.5 eq.) before being added to the resin.
- Peptide coupling was monitored using the 2,4,6-trinitrobenzenesulphonic acid (TNBS) test.
- TNBS 2,4,6-trinitrobenzenesulphonic acid
- Fmoc-amino acids were used as protected Fmoc-amino acids with the standard side chain protecting groups: Fmoc-Ala-OH, Fmoc-Arg(Pbf)-OH, Fmoc-Asp(OtBu)-OH, Fmoc-Glu(OtBu)-OH, Fmoc-Ile-OH, Fmoc-Ser(tBu)-OH, Fmoc-Thr(tBu)-OH, Fmoc-Tyr(tBu)-OH, Fmoc-Val-OH and Fmoc-Dap(6DMN).
- Phosphotyrosine was introduced as the monobenzyl ester Fmoc-Tyr(PO(OBzl)OH)—OH.
- High-performance liquid chromatography HPLC was performed using a Waters 600E HPLC fitted with a Waters 600 automated control module and a Waters 2487 dual wavelength absorbance detector recording at 228 and 280 nm.
- analytical HPLC a Beckman Ultrasphere C 18 , 5 ⁇ m, 4.6 ⁇ 150 mm reverse-phase column was used.
- For preparative separations a YMC-pack, C 18 , 250 ⁇ 20 mm reversed phase column was used.
- the standard gradient for analytical and preparative HPLC used was 93:7 to 5:95 over 35 minutes (water:acetonitrile, 0.1% TFA).
- the 6DMN side chain proved resistant to the standard mildly basic amino acid coupling conditions (0.12 M diisopropylethylamine), the Fmoc deprotection conditions (20% piperidine), and the acidic resin cleavage and deprotection cocktail (95% TFA).
- ⁇ x ⁇ std ⁇ ( grad x grad std ) ⁇ ( ⁇ x 2 ⁇ std 2 ) ( 1 )
- ⁇ is the fluorescence quantum yield
- grad is the gradient (grad) from the plot of integrated fluorescence intensity vs. absorbance
- 77 the refractive index of the solvent.
- the UV absorbance spectra (40 ⁇ M in methanol) and fluorescence emission spectra of compound (VIII), 6DMN-GlyOMe are shown in FIG. 6 and tabulated in Table 4.
- the UV spectrum of 6DMN-GlyOMe has an intense absorption band at 378 nm which is not found in unsubstituted 2,3-naphthalimides.
- the maximum excitation wavelength of 6DMN-GlyOMe is at 375 nm.
- the data were fitted to a linear equation with gradient (grad), and intercept 0.
- the gradient (grad) is proportional to the quantum yield of the sample. Absolute values were calculated using the standard sample, which has a fixed and known fluorescence quantum yield value, according to the equation above.
- ⁇ ⁇ The difference ( ⁇ ⁇ ) in the maximum absorption ( ⁇ A ) and emission wavelengths ( ⁇ F ), known as the Stokes shift and expressed in wavenumbers, is plotted vs. ⁇ f.
- ⁇ f The term ⁇ f is called the orientation polarizability and can be calculated according to the following equation:
- ⁇ ⁇ ⁇ f ( ⁇ - 1 2 ⁇ ⁇ ⁇ + 1 - n 2 - 1 2 ⁇ ⁇ n 2 + 1 ) ( 2 )
- FIG. 7( a ) shows the plot of ( ⁇ ⁇ ) for compound (VIII) vs. ⁇ f for a series of 1,4-dioxane/acetonitrile mixtures.
- ⁇ f is obtained for each mixture using ⁇ and n values calculated using the molar fractions ( ⁇ ) in the equation as follow:
- n ( ⁇ acetonitrile )(1.3442)+( ⁇ diox )(1.4224) (4)
- c is the velocity of light
- h Plank's constant
- ⁇ e ⁇ g is the difference between the dipole moments of the excited and the ground states respectively ( ⁇ )
- a 0 is the radius of the Onsager cavity around the compound.
- the Onsager radius for compound VIII was calculated from the optimized structure obtained with a DFT minimization using the Gaussian program (Pople, J. A. et al. Gaussian 98 (Gaussian, Inc., Pittsburgh, Pa., 1998), (B3LYP functional using 6-31G(d) orbital base).
- the Onsager radius (4.19 ⁇ ) was taken as half of the average distance between nitrogen of the amine donor and the two carbonyl oxygens, which corresponds to the longest distance across the molecule where charge separation can take place.
- the Onsager radius (4.19 ⁇ ) was taken as half of the average distance between nitrogen of the amine donor and the two carbonyl oxygens, which corresponds to the longest distance across the molecule where charge separation can take place.
- FIG. 7( a ) As can be seen in FIG. 7( a ), the Stokes shift of compound VIII in a series of mixtures of 1,4-dioxane/acetonitrile changes linearly in response to the solvent polarity, which correlates with an increase in the dipole moment ( ⁇ ⁇ ) of 5.5 D.
- Peptide stock solutions of known concentration were used to prepare 5 ⁇ M peptide concentration samples in buffered water (150 ⁇ L total volume, 100 mM NaCl, 10 mM sodium phosphate, pH 7.5).
- 2 ⁇ L aliquots of the corresponding stock protein solution (GST-Crk SH2, GST-Abl SH2 or GST-PI3K SH2, concentration determined by Bio-Rad Protein Assay) were consecutively added, and fluorescence emission spectra were recorded 1 minute after each addition (excitation 395 nm, 2 nm excitation and emission slit width). The increase in emission intensity at the maximum emission wavelength was plotted against total protein concentration and fitted by least-square analysis assuming a 1 to 1 binding mode to obtain the apparent Kd.
- Lysis buffer PBS with 100 mM EDTA, 1% Triton X-100, 10% glycerol, 1 mM DTT, AEBSF, leupeptin and PepA. (add 100 ⁇ L AEBSF+25 ⁇ L Leupeptin+12.5 ⁇ L Pep A+to 10 mL total buffer).
- the sample was recentrifuged to remove particulate debris (5000 g for 30 min at 4° C.). Then the supernatant was incubated for at least 1 hour with 10 mL of glutathione Sepharose 4 Fast Flow (Amersham Biosciences). The column was equilibrated with 5 volumes of binding buffer at 4° C. The sample was the loaded using a syringe and low flow rate (0.2 to 0.5 mL/min) at 4° C. The column was then washed with 5 volumes of binding buffer (higher rate of 1 mL/min) room temperature. The GST fusion protein was eluted with elution buffer at room temperature.
- Binding buffer PBS pH 7.4 (140 mM NaCl, 2.7 mM KCl, 10 mM Na 2 HPO 4 , 1.8 mM NaHPO 4 ).
- Elution buffer 50 mM Tris-HCl, 10 mM reduced glutathione, pH 8.
- the gel was transferred to a nitrocellulose membrane (1 h, 100 V), and probed using the standard procedure: blocked overnight with TBST+powder milk; washed with TBST (2 ⁇ 10 mL ⁇ 10 min.); incubated with primary antibody (anti-GST, dilution 1:10,000 in TBST) for 30 min; washed with TBST (3 ⁇ 10 mL ⁇ 10 min.); incubated for 30 min with secondary antibody (anti-goat conjugated with alkaline phosphatase, 1:1,000 dilution in TBS); washed with TBST (2 ⁇ 10 mL ⁇ 10 min.); washed with TBS (1 ⁇ 10 mL ⁇ 10 min.); One Step NBT/BCIP added and incubated until the bands appeared; then quenched the reaction with ddH 2 O.
- Bio-Rad Protein Assay was used to quantify the isolated following the standard procedure.
- Dye reagent was prepared by diluting 1 part Dye Reagent Concentrate with 4 parts distilled water.
- Stock BSA dilutions were prepared corresponding to 0.1, 0.2, 0.3, 0.4, 0.5 0.6, 0.7, 0.8 and 0.9 mg/mL as follows. 20 ⁇ L of each stock dilution were added to 980 ⁇ L dye reagent and incubated at room temperature for 5 min. The absorbance was measured at 595 nm using a blank of 20 ⁇ L of distilled water and 980 ⁇ L dye reagent prepared at the same time as the samples. The observed absorbance was plotted against BSA concentration.
- Abl-SH2 was received as a GST fusion in a pGEX-KT vector, amp-resistant, IPTG-inducible.
- the SH2 domain was cloned using BamH1 (5′) EcoR1 (3′) sites.
- Src and PI3K SH2 domains were expressed and purified as GST fusion following reported procedures. (Smith, D. B.; Johnson, K. S. Gene, 1988, 67, 31-40) In short, plasmids including GST-fusion proteins were transformed into bacteria ( E. coli DH5 ⁇ chemically competent cells, Invitrogen). Bacteria were grown to mid log phase, induced at 37° C.
- Proteins were eluted from the beads (50 mM Tris-HCl, 10 mM reduced glutathione, pH 8), and finally concentrated by centrifugation through a cellulose membrane with a 10,000 Da cutoff (Millipore). Proteins were quantified using the Micro BCA protein Assay Kit (Pierce) relative to a BSA standard. All fusion proteins were analyzed by Coomasie staining and Western analysis with anti-GST antibodies.
Landscapes
- Health & Medical Sciences (AREA)
- Chemical & Material Sciences (AREA)
- Immunology (AREA)
- Life Sciences & Earth Sciences (AREA)
- Organic Chemistry (AREA)
- Engineering & Computer Science (AREA)
- Urology & Nephrology (AREA)
- Biomedical Technology (AREA)
- Hematology (AREA)
- Molecular Biology (AREA)
- Biotechnology (AREA)
- Microbiology (AREA)
- Food Science & Technology (AREA)
- Medicinal Chemistry (AREA)
- Physics & Mathematics (AREA)
- Analytical Chemistry (AREA)
- Biochemistry (AREA)
- General Health & Medical Sciences (AREA)
- General Physics & Mathematics (AREA)
- Pathology (AREA)
- Cell Biology (AREA)
- Plural Heterocyclic Compounds (AREA)
Abstract
Description
- The subject matter of this application was developed in part with funding from the Cell Migration Consortium (grant application no. GM64346) and National Science Foundation (grant application no. CHE-0414243). The government may have certain rights in this technology.
- Fluorescence is the result of a three-stage process that occurs when certain molecules absorb energy. The three stages comprise: 1) excitation; 2) excited-state lifetime; and 3) fluorescence emission. During
stage 1, excitation, a photon of a certain energy is absorbed by the fluorophore. The fluorophore is initially in its ground state (S0). Absorption of the photon causes that fluorophore to become excited. The energy of the absorbed photon is transferred to an electron. The electron is transferred to a higher energy state. The fluorophore exists in an excited electronic singlet state (S1′), also called an excited state. The excited state of the fluorophore exists for a finite time, typically 10−8 to 10−9 seconds. During the excited state, the fluorophore changes in its translational, vibrational, and electronic energy states, and is subject to interactions with its molecular environment. The excited fluorophore releases energy and returns to the ground state, S0, by fluorescence emission. Other processes such as fluorescence energy transfer, intersystem crossing, and collisional quenching may also depopulate S1. The ratio of the number of fluorescence photons emitted, during the emission stage, to the number of photons absorbed, during the excitation stage, is termed the quantum yield. The quantum yield is a measure of the efficiency of fluorescence in competition with other processes such as fluorescence energy transfer, intersystem crossing, and collisional quenching. - During the third stage, fluorescence emission, a photon of energy hv (where h is Planck's constant and v is the frequency of the photon) is emitted, returning the fluorophore to its ground state S0. The energy of the emitted photon is lower than the energy of the photon absorbed during the excitation stage. The difference in energy can be attributed to dissipation through processes during the excited-state lifetime, such processes include fluorescence energy transfer, intersystem crossing, and collisional quenching. The difference in energy of the absorbed photon and the emitted photon is called the Stokes shift. The Stokes shift is fundamental to the sensitivity of fluorescence techniques because it allows emission photons to be detected against a low background, and at a different wavelength than the excitation photons.
- Compounds that have fluorescent properties have numerous uses. Fluorescent molecules can be used in single molecule spectroscopy, liquid crystal displays, light emitting diodes, solar energy collectors, and laser active media. Fluorescent molecules whose spectra or quantum yields are sensitive to their environments are valuable as fluorescent dyes and in the study of heterogeneous media, organized media, and biological media.
- The present invention provides compounds that can be used to monitor biological interactions continuously with a fluorescence readout. The compounds of the present invention are environment-sensitive fluorophores which have spectroscopic behavior that is dependent on the physicochemical properties of the surrounding environment. The compounds of the present invention can be used in biochemical research to monitor ions, small molecules, and biological processes such as protein folding, protein-protein interactions and phosphorylation events.
- Environment-sensitive fluorophores are a special class of chromophores that have spectroscopic behavior that is dependent on the physicochemical properties of the surrounding environment. Solvatochromic fluorophores display sensitivity to the polarity of the local environment. These molecules exhibit a low quantum yield in aqueous solution, but become highly fluorescent in nonpolar solvents or when bound to hydrophobic sites in proteins or membranes. Examples of solvatochromic fluorophores include 2-propionyl-6-dimethylaminonaphthalene (PRODAN) (Weber et al. Biochemistry 1979, 18, 3075-3078; Cohen et al. Science 2002, 296, 1700-1703), 4-dimethylamino phthalimide (4-DMAP) (Saroja et al. J. Fluoresc. 1998, 8, 405-410), and 4-amino-1,8-naphthalimide derivatives (Grabchev et al. J. Photochem. Photobiol., A 2003, 158, 37-43; Martin et al. J. Lumin. 1996, 68, 157-146). Although PRODAN and derivatives are widely used, these probes have limitations resulting from the relatively intense fluorescence even in aqueous environments. Thus, there is a need for alternate compounds.
- The present invention provides novel compounds of the formula (I):
- where n, R3, R4, R5, R6, R7, R8 and R9 are as defined below. One especially preferred compound (III) referred to as Dap(6DMN) is disclosed.
- The present invention also provides peptides containing the compound (I) of the present invention.
- The present invention also provides a method for probing biological interactions using peptides containing the compound (I).
-
FIG. 1 are fluorescence titrations of peptides (Crk-bp and Crk-bp2) binding to GST-Crk SH2 domain. (a) Peptide Crk-bp (20 μM in PBS buffer, pH 7.5) with GST-Crk SH2, and (b) Peptide Crk-bp2 (20 μM in PBS buffer, pH 7.5) with GST-Crk SH2. Inserts show plots of the fluorescence emission intensity with the best fitting binding curves. Spectra were corrected for the dilution upon addition of the protein solution. (c) Relative fluorescence emission intensities at 565 nm for: 1) Peptides Crk-bp and Crk-bp2 in buffer. 2) Peptide Crk-bp saturated with GST-Crk SH2. 3) Peptide Crk-bp2 saturated with GST-Crk SH2. 4) Peptide Crk-bp2 saturated with GST-Abl SH2. 5) Peptide Crk-bp2 saturated with GST-PI3K SH2. -
FIG. 2 are fluorescence titrations of peptide Abl-bp with various GST SH2 domains. (a) Abl-bp (20 μM in PBS buffer, pH 7.5) with GST-Abl SH2. (b) Abl-bp (20 μM in PBS buffer, pH 7.5) with GST-PI3K SH2. Inserts show plots of the fluorescence emission intensity at 599 nm, with the best fitting binding curves. Spectra were corrected for the dilution upon addition of the protein solution. (c) Relative fluorescence emission intensities at 590 nm for: 1) Peptide Abl-bp. 2) Peptide Abl-bp saturated with GST-Abl SH2 and 3) Peptide Abl-bp saturated with GST-PI3K SH2. 4) Peptide Abl-bp saturated with GST-Crk SH2. (GST-PI3K SH2 and GST-Crk SH2 are not-target SH2 domains for the peptide Abl-bp.) -
FIG. 3 are fluorescence titrations of peptide Abl-bp2 with various GST SH2 domains. (a) Abl-bp2 (20 μM in PBS buffer, pH 7.5) with GST-Abl SH2. (b) Abl-bp2 (20 μM in PBS buffer, pH 7.5) with GST-PI3K SH2. Inserts show plots of the fluorescence emission intensity at 599 nm, with the best fitting binding curves. Spectra were corrected for the dilution upon addition of the protein solution. (c) Relative fluorescence emission intensities at 599 nm for: 1) Peptide Abl-bp2. 2) Peptide Abl-bp2 saturated with GST-Abl SH2 and 3) Peptide Abl-bp2 saturated with GST-PI3K SH2. 4) Peptide Abl-bp2 saturated with GST-Crk SH2. (GST-PI3K SH2 and GST-Crk SH2 are not-target SH2 domains for the peptide Abl-bp.2) -
FIG. 4 are graphs depicting fluorescence titrations of peptides with non-target SH2 domains. a) Crk-bp with GST-Abl SH2, b) Crk-bp2 with GST-Abl SH2, c) Abl-bp with GST-Crk SH2, d) Abl-bp2 with GST-Crk SH2. -
FIG. 5 are plots of integrated fluorescence vs. absorbance used to calculate the quantum yield (Φ) of Ac-Gly-Dap(6DMN)-Gln-pTyr-Glu-Asn-Val-Gln-Ser-(CONH2). The quantum yield is proportional to the gradient of the plot. The absolute value is calculated using a standard sample, quinine sulfate. a) Integrated fluorescence vs. absorbance for standard solution: quinine sulfate in 0.1 M H2SO4. b) Integrated fluorescence vs. absorbance for test solution: Ac-Gly-Dap(6DMN)-Gln-pTyr-Glu-Asn-Val-Gln-Ser-(CONH2) in 10 mM phosphate buffer, pH 7.5. -
FIG. 6 are UV absorption and fluorescence emission spectra of compound VIII, 6DMN-GlyOMe. a) UV absorption spectra (40 μM) in methanol. b) Fluorescence emission spectra in different solvents. Labels show maximum emission wavelengths for selected solvents (toluene, chloroform and methanol). -
FIG. 7 are Lippert-Mataga plots for compound VIII, 6DMN-GlyOMe. (a) Plot of νA−νF (Stokes shift) vs. Δf (orientation polarizability of the solvent) for 6DMN-GlyOMe in a series of 1,4-dioxane/acetonitrile mixtures. νA−νF (Δν ) is the difference in the maximum absorption and emission wavelengths, expressed in wavenumbers.ν A−νF (Δν ) was measured from a series of 1,4-dioxane/acetonitrile mixtures. Δf was calculated based upon the composition of the 1,4-dioxane/acetonitrile mixture using the dielectric constant and refraction index of 1,4-dioxane and acetonitrile. The slope corresponds to μe−μg (Δμ) which is the difference between the dipole moments of the excited and ground states for 6DMN-GlyOMe. The greater the change in the dipole moment, the more sensitive the fluorophore. (b) Plot of νA−ν F (Stokes shift) vs. Δf (orientation polarizability of the solvent) for 6DMN-GlyOMe in a series of different solvents. Δf was calculated for each solvent using the dielectric constant and refraction index of the corresponding solvent. The line corresponds to the best linear fit to the data, excluding the values obtained for protic solvents (methanol, isopropanol and ethanol, closed circles), indicating that the Stokes shift is proportional to the orientation polarizability. Protic solvents (methanol, isopropanol and ethanol) show disproportionately large Stokes shifts which may be due to two different excited states. (Kosower, E. M.; Dodiuk, H. J. Am. Chem. Soc. 1974, 96, 6195-6196; Kosower, E. M. Acc. Chem. Res. 1982, 15, 259-266; Saha, S.; Samanta, A. J. Phys. Chem. A 2002, 106, 4763-4771) - The present invention is directed to compounds and salts thereof, compositions and methods useful in monitoring biological interactions continuously with a fluorescence readout.
- When describing the compounds, compositions, methods and processes of this invention, the following terms have the following meanings, unless otherwise indicated.
- “Alkyl” by itself or as part of another substituent refers to a hydrocarbon group which may be linear, cyclic, or branched or a combination thereof having the number of carbon atoms designated (i.e., C1-8 means one to eight carbon atoms). Examples of alkyl groups include methyl, ethyl, n-propyl, isopropyl, n-butyl, t-butyl, isobutyl, sec-butyl, cyclohexyl, cyclopentyl, (cyclohexyl)methyl, cyclopropylmethyl, bicyclo[2.2.1]heptane, bicyclo[2.2.2]octane, etc. Alkyl groups can be substituted or unsubstituted, unless otherwise indicated. Examples of substituted alkyl include haloalkyl, thioalkyl, aminoalkyl, and the like.
- “Alkoxy” refers to —O-alkyl. Examples of an alkoxy group include methoxy, ethoxy, n-propoxy, etc.
- “Alkenyl” refers to an unsaturated hydrocarbon group which may be linear, cyclic or branched or a combination thereof. Alkenyl groups with 2-8 carbon atoms are preferred. The alkenyl group may contain 1, 2 or 3 carbon-carbon double bonds. Examples of alkenyl groups include ethenyl, n-propenyl, isopropenyl, n-but-2-enyl, n-hex-3-enyl, cyclohexenyl, cyclopentenyl and the like. Alkenyl groups can be substituted or unsubstituted, unless otherwise indicated.
- “Alkynyl” refers to an unsaturated hydrocarbon group which may be linear, cyclic or branched or a combination thereof. Alkynyl groups with 2-8 carbon atoms are preferred. The alkynyl group may contain 1, 2 or 3 carbon-carbon triple bonds. Examples of alkynyl groups include ethynyl, n-propynyl, n-but-2-ynyl, n-hex-3-ynyl and the like. Alkynyl groups can be substituted or unsubstituted, unless otherwise indicated.
- “Aryl” refers to a polyunsaturated, aromatic hydrocarbon group having a single ring (monocyclic) or multiple rings (bicyclic), which can be fused together or linked covalently. Aryl groups with 6-10 carbon atoms are preferred, where this number of carbon atoms can be designated by C6-10, for example. Examples of aryl groups include phenyl and naphthalene-1-yl, naphthalene-2-yl, biphenyl and the like. Aryl groups can be substituted or unsubstituted, unless otherwise indicated.
- “Fluorescence” encompasses the release of fluorescent energy. Less broadly, the term “fluorescence” refers to fluorescent emission, the rate of change of fluorescence over time (i.e., fluorescence lifetime), fluorescence polarization, fluorescence anisotropy, and fluorescence resonance energy transfer. See Eftink, M. R., Biophysical J. 66:482-501 (1994).
- “Fluorescence probe molecule” refers to a compound of the present invention. The compound, after excitement by light of a defined wavelength or defined range of wavelengths, is capable of emitting fluorescent energy. The fluorescent molecule or a compound may be capable of binding to a peptide, protein, membrane or receptor.
- “Halo” or “halogen”, by itself or as part of a substituent refers to a chlorine, bromine, iodine, or fluorine atom.
- “Haloalkyl”, as a substituted alkyl group, refers to a monohaloalkyl or polyhaloalkyl group, most typically substituted with from 1-3 halogen atoms. Examples include 1-chloroethyl, 3-bromopropyl, trifluoromethyl and the like.
- “Heterocyclyl” refers to a saturated or unsaturated non-aromatic ring containing at least one heteroatom (typically 1 to 5 heteroatoms) selected from nitrogen, oxygen or sulfur. The heterocyclyl ring may be monocyclic or bicyclic. Preferably, these groups contain 0-5 nitrogen atoms, 0-2 sulfur atoms and 0-2 oxygen atoms. More preferably, these groups contain 0-3 nitrogen atoms, 0-1 sulfur atoms and 0-1 oxygen atoms. Examples of heterocycle groups include pyrrolidine, piperidine, imidazolidine, pyrazolidine, butyrolactam, valerolactam, imidazolidinone, hydantoin, dioxolane, phthalimide, piperidine, 1,4-dioxane, morpholine, thiomorpholine, thiomorpholine-S-oxide, thiomorpholine-S,S-dioxide, piperazine, pyran, pyridone, 3-pyrroline, thiopyran, pyrone, tetrahydrofuran, tetrahydrothiophene, quinuclidine and the like. Preferred heterocyclic groups are monocyclic, though they may be fused or linked covalently to an aryl or heteroaryl ring system.
- In one preferred embodiment, heterocyclic groups may be represented by formula (AA) below:
- where formula (AA) is attached via a tree valence on either M1 or M2; M1 represents O, NRe, or S(O)l; M2 represents CRfRg, O, S(O)l, or NRe; l is 0, 1 or 2; j is 1, 2 or 3 and k is 1, 2 or 3, with the proviso that j+k is 3, 4, or 5; and Ra, Rb, Rc, Rd, Re, Rf, and Rg are independently selected from the group consisting of hydrogen, halogen, unsubstituted or substituted C1-8 alkyl, unsubstituted or substituted C2-8 alkenyl, unsubstituted or substituted C2-8 alkynyl, —CORh, —CO2Rh, —CONRhRi, —NRhCORi, —SO2Rh, SO2NRhRi, NSO2Rh Ri, NRhRi, —ORh, -Q1CORh, -Q1CO2Rh, -Q1CONRhRi, -Q1NRhCORi, -Q1SO2R28, -Q1SO2NRhRi, -Q1NSO2RhRi, -Q1NRhRi, -Q1ORh, wherein Q1 is a member selected from the group consisting of C1-4 alkylene, C2-4 alkenylene and C2-4 alkynylene, and Rh and Ri are independently selected from the group consisting of hydrogen and C1-8 alkyl, and wherein the aliphatic portions of each of the Ra, Rb, Rc, Rd, Re, Rf, Rg, Rh and Ri substituents are optionally substituted with from one to three members selected from the group consisting of halogen, —OH, —ORn, —OC(O)NHRn, —OC(O)NRnRo, —SH, —SRn, —S(O)Rn, —S(O)2Rn, —SO2NH2, —S(O)2NHRn, —S(O)2NRnRo, —NHS(O)2Rn, —NRnS(O)2Ro, —C(O)NH2, —C(O)NHRn, —C(O)NRnRo, —C(O)Rn, —NHC(O)Ro, —NRnC(O)Ro, —NHC(O)NH2, —NRnC(O)NH2, —NRnC(O)NHRo, —NHC(O)NHRn, —NRnC(O)NRo Rp, —NHC(O)NRnRo, —CO2H, —CO2Rn, —NHCO2Rn, —NRnCO2Ro, —ON, —NO2, —NH2—NHRn, —NRnRo, —NRnS(O)NH2 and —NRnS(O)2NHRo, wherein Rn, Ro and Rp are independently an unsubstituted C1-8 alkyl. Additionally, any two of Ra, Rb, Rc, Rd, Re, Rf and Rg may be combined to form a bridged or spirocyclic ring system.
- In one preferred embodiment, the number of Ra+Rb+Rc+Rd groups that are other than hydrogen is 0, 1 or 2. In a more preferred embodiment, Ra, Rb, Rc, Rd, Re, Rf, and Rg are independently selected from the group consisting of hydrogen, halogen, unsubstituted or substituted C1-8 alkyl, —CORh, —CO2Rh, —CONRhRh, —NRhCORh, —SO2Rh, —SO2NRhRi, NSO2RhRi, —NRhRi, and —ORh, wherein Rh and Ri are independently selected from the group consisting of hydrogen and unsubstituted C1-8 alkyl and wherein the aliphatic portions of each of the Ra, Rb, Rc, Rd, Re, Rf and Rg substituents are optionally substituted with from one to three members selected from the group consisting of halogen, —OH, —ORe, —OC(O)NHRn, —OC(O)NRnRo, —SH, —SRn, —S(O)Ro, —S(O)2Rn, —SO2NH2, —S(O)2NHRn, —S(O)2NRnRo, —NHS(O)2Rn, —NRnS(O)2Ro, —C(O)NH2, —C(O)NHRn, —C(O)NRnRo, —C(O)Rn, —NHC(O)Rn, —NRnC(O)Ro, —NHC(O)NH2, —NRnC(O)NH2, —NRnC(O)NHRo, —NHC(O)NHRn, —NRnC(O)NRo Rp, —NHC(O)NRnRo, —CO2H, —CO2Rn, —NHCO2Rn, —NRnCO2Ro, —CN, —NO2, —NH2—, NHRn, —NRnRo, —NRnS(O)NH2 and —NR S(O)2NHRo, wherein Rn, Ro and Rp are independently an unsubstituted C1-8 alkyl.
- In a more preferred embodiment, Ra, Rb, Rc, Rd, Re, Rf, and Rg are independently hydrogen or C1-4 alkyl. In another preferred embodiment, at least three of Ra, Rb, Rc, Rd, Re, Rf, and Rg are hydrogen.
- “Heteroaryl” refers to an aromatic group containing at least one heteroatom, where the heteroaryl group may be monocyclic or bicyclic. Examples include pyridyl, pyridazinyl, pyrazinyl, pyrimidinyl, triazinyl, quinolinyl, quinoxalinyl, quinazolinyl, cinnolinyl, phthalazinyl, benzotriazinyl, purinyl, benzimidazolyl, benzopyrazolyl, benzotriazolyl, benzisoxazolyl, isobenzofuryl, isoindolyl, indolizinyl, benzotriazinyl, thienopyridinyl, thienopyrimidinyl, pyrazolopyrimidinyl, imidazopyridines, benzothiazolyl, benzofuranyl, benzothienyl, indolyl, quinolyl, isoquinolyl, isothiazolyl, pyrazolyl, indazolyl, pteridinyl, imidazolyl, triazolyl, tetrazolyl, oxazolyl, isoxazolyl, oxadiazolyl, thiadiazolyl, pyrrolyl, thiazolyl, furyl or thienyl. Preferred heteroaryl groups are those having at least one aryl ring nitrogen atom, such as quinolinyl, quinoxalinyl, purinyl, benzimidazolyl, benzopyrazolyl, benzotriazolyl, benzothiazolyl, indolyl, quinolyl, isoquinolyl and the like. Preferred 6-ring heteroaryl systems include pyridyl, pyridazinyl, pyrazinyl, pyrimidinyl, triazinyl and the like. Preferred 5-ring heteroaryl systems include isothiazolyl, pyrazolyl, imidazolyl, thienyl, furyl, triazolyl, tetrazolyl, oxazolyl, isoxazolyl, oxadiazolyl, thiadiazolyl, pyrrolyl, thiazolyl and the like.
- Heterocyclyl and heteroaryl can be attached at any available ring carbon or heteroatom. Each heterocyclyl and heteroaryl may have one or more rings. When multiple rings are present, they can be fused together or linked covalently. Each heterocyclyl and heteroaryl must contain at least one heteroatom (typically 1 to 5 heteroatoms) selected from nitrogen, oxygen or sulfur. Preferably, these groups contain 0-5 nitrogen atoms, 0-2 sulfur atoms and 0-2 oxygen atoms. More preferably, these groups contain 0-3 nitrogen atoms, 0-1 sulfur atoms and 0-1 oxygen atoms. Heterocyclyl and heteroaryl groups can be substituted or unsubstituted, unless otherwise indicated. For substituted groups, the substitution may be on a carbon or heteroatom. For example, when the substitution is oxo (═O or —O−), the resulting group may have either a carbonyl (—C(O)—) or a N-oxide (—N+—O−).
- Suitable substituents for substituted alkyl include halogen, —CN, —CO2R′, —C(O)R′, —C(O)NR′R″, oxo (═O or —O−), —OR′, —OC(O)R′, —OC(O)NR′R″, —NO2, —NR′C(O)R″, —NR′″C(O)NR′R″, —NR′R″, —NR′CO2R″, —NR′S(O)R″, —NR′S(O)2R′″, NR′″S(O)NR′R″, —NR′″S(O)2NR′R″, —SR′, —S(O)R′, —S(O)2R′, —S(O)2NR′R″, —NR′—C(NHR″)═NR′″, —SiR′R″R′″, —N3, substituted or unsubstituted C6-10 aryl, substituted or unsubstituted 5- to 10-membered heteroaryl, and substituted or unsubstituted 3- to 10-membered heterocyclyl. The number of possible substituents range from zero to (2m′+1), where m′ is the total number of carbon atoms in such radical.
- Suitable substituents for substituted ring include halogen, —CN, —CO2R′, —C(O)R′, —C(O)NR′R″, oxo (═O or —O−), —OR′, —OC(O)R′, —OC(O)NR′R″, —NO2, —NR′C(O)R″, —NR′″C(O)NR′R″, —NR′R″, —NR′CO2R″, —NR′S(O)R″, —NR′S(O)2R″, —NR′″S(O)NR′R″, —NR′″S(O)2NR′R″, —SR′, —S(O)R′, —S(O)2R′, —S(O)2NR′R″, —NR′—C(NHR″)═NR′″, —SiR′R″R′″, —N3, substituted or unsubstituted C1-8 alkyl, substituted or unsubstituted C2-8 alkenyl, substituted or unsubstituted C2-8 alkynyl, substituted or unsubstituted C6-10 aryl, substituted or unsubstituted 5- to 10-membered heteroaryl, and substituted or unsubstituted 3- to 10 membered heterocyclyl. The number of possible substituents range from zero to the total number of open valences on the ring system.
- As used above, R′, R″ and R′″ each independently refer to a variety of groups including hydrogen, substituted or unsubstituted C1-8 alkyl, substituted or unsubstituted C2-8 alkenyl, substituted or unsubstituted C2-8 alkynyl, substituted or unsubstituted aryl, substituted or unsubstituted heteroaryl, substituted or unsubstituted heterocyclyl, substituted or unsubstituted arylalkyl, substituted or unsubstituted aryloxyalkyl. When R′ and R″ are attached to the same nitrogen atom, they can be combined with the nitrogen atom to form a 3-, 4-, 5-, 6-, or 7-membered ring (for example, —NR′R″ includes 1-pyrrolidinyl and 4-morpholinyl). Furthermore, R′ and R″, R″ and R′″, or R′ and R′″ may together with the atom(s) to which they are attached, form a substituted or unsubstituted 5-, 6- or 7-membered ring.
- “Heteroatom” is meant to include oxygen (O), nitrogen (N), sulfur (S) and silicon (Si).
- “N-protecting group” refers to an easily removable group which is known in the art to protect an amino group against undesirable reaction during synthetic procedures and to be selectively removable. The use of N-protecting groups is well known in the art for protecting groups against undesirable reactions during a synthetic procedure and many such protecting groups are known. Commonly used N-protecting groups are known to those skilled in the art, examples of which are disclosed in Greene and Wuts, Protective Groups in Organic Synthesis, 2nd ed.; John Wiley & Sons, New York, 1991). Examples of N-protecting groups include, but are not limited to, acyl groups including formyl, acetyl (Ac), trifluoroacetyl, trichloroacetyl, propionyl, pivaloyl, t-butylacetyl, acylisothiocyanate, aminocaproyl, benzoyl and the like; acyloxy groups, including t-butyloxycarbonyl (Boc), benzyloxycarbonyl (Cbz), 9-fluorenylmethoxycarbonyl (Fmoc), p-methoxybenzyloxycarbonyl, methoxycarbonyl, ethoxycarbonyl, allyloxycarbonyl and the like; sulfonyl groups such as benzenesulfonyl, p-toluenesulfonyl and the like; arylalkyl groups such as benzyl, triphenylmethyl, benzyloxymethyl and the like; and silyl groups such as trimethylsilyl, t-butyldimethylsilyl and the like.
- All of the above terms (e.g., “alkyl,” “aryl,” “heteroaryl” etc.), in some embodiments, include both substituted and unsubstituted forms of the indicated groups. These groups may be substituted 1 to 10 times, as chemically allowed. Suitable substituents include alkyl, aryl, heteroaryl, heterocyclyl, halogen, alkoxy, oxygen, and nitrogen.
- Certain compounds of the present invention can exist in unsolvated forms as well as solvated forms, including hydrated forms. In general, both solvated forms and unsolvated forms are intended to be encompassed within the scope of the present invention. Certain compounds of the present invention may exist in multiple crystalline or amorphous forms (i.e., as polymorphs). In general, all physical forms are equivalent for the uses contemplated by the present invention and are intended to be within the scope of the present invention.
- Certain compounds of the present invention possess asymmetric carbon atoms (optical centers) or double bonds; the racemates, diastereomers, geometric isomers and individual isomers (e.g., separate enantiomers) are all intended to be encompassed within the scope of the present invention. The compounds of the present invention may also contain unnatural proportions of atomic isotopes at one or more of the atoms that constitute such compounds. For example, the compounds may be radiolabeled with radioactive isotopes, such as for example tritium (3H), iodine-125 (125I) or carbon-14 (14C). All isotopic variations of the compounds of the present invention, whether radioactive or not, are intended to be encompassed within the scope of the present invention.
- The term “biological interactions” encompasses the interaction of a compound or molecule with a target molecule.
- “Protein” and “peptide”, as used herein, are synonymous. For proteins or peptides, the term “unfolding” encompasses any change in structure due to heating. For example, the term “unfolding” refers to the transition of from the liquid crystalline state to the molten globule state. In the molten globule state, tertiary and quaternary structure has been altered, relative to the native state of the protein, and at least some secondary structure remains intact. The term “unfolding” also encompasses loss of crystalline ordering of amino acid side-chains, secondary, tertiary or quaternary structure. The term “unfolding” also encompasses formation of a random coil.
- “Folding” and “refolding,” and “renaturing” refer to the acquisition of the correct amino acid side-chain ordering, secondary, tertiary, or quaternary structure, of a protein or a nucleic acid, which affords the full chemical and biological function of the biomolecule.
- The term “target molecule” encompasses peptides, proteins, nucleic acids, ions, and other receptors. The term encompasses both enzymes, and proteins which are not enzymes. The term encompasses monomeric and multimeric proteins. Multimeric proteins may be homomeric or heteromeric. The term encompasses nucleic acids comprising at least two nucleotides, such as oligonucleotides. Nucleic acids can be single-stranded, double-stranded, or triple-stranded. The term encompasses a nucleic acid which is a synthetic oligonucleotide, a portion of a recombinant DNA molecule, or a portion of chromosomal DNA. The term target molecule also encompasses portions of peptides, proteins, and other receptors which are capable of acquiring secondary, tertiary, or quaternary structure through folding, coiling or twisting. The target molecule may be substituted with substituents including, but not limited to, cofactors, coenzymes, prosthetic groups, lipids, oligosaccharides, or phosphate groups.
- The terms “target molecule” and “receptor” are synonymous.
- Examples of target molecules are included, but not limited to those disclosed in Faisst, S. et al., Nucleic Acids Research 20:3-26 (1992); Pimentel, E., Handbook of Growth Factors, Volumes I-III, CRC Press, (1994); Gilman, A. G. et al., The Pharmacological Basis of Therapeutics, Pergamon Press (1990); Lewin, B., Genes V, Oxford University Press (1994); Roitt, I., Essential Immunology, Blackwell Scientific Publ. (1994); Shimizu, Y., Lymphocyte Adhesion Molecules, R G Landes (1993); Hyams, J. S. et al., Microtubules, Wiley-Liss (1995); Montreuil, J. et al., Glycoproteins, Elsevier (1995); Woolley, P., Lipases: their Structure Biochemistry and Applications, Cambridge University Press (1994); Kurjan, J., Signal Transduction: Prokaryotic and Simple Eukaryotic Systems, Academic Press (1993); Kreis, T., et al., Guide Book to the Extra Cellular Matrix and Adhesion Proteins, Oxford University Press (1993); Schlesinger, M. J., Lipid Modifications of Proteins, CRC Press (1992); Conn, P. M., Receptors: Model Systems and Specific Receptors, Oxford University Press (1993); Lauffenberger, D. A. et al, Receptors. Models For Binding Trafficking and Signaling, Oxford University Press (1993); Webb, E. C., Enzyme Nomenclature, Academic Press (1992); Parker, M. G., Nuclear Hormone Receptors; Molecular Mechanisms, Cellular Functions Clinical Abnormalities, Academic Press Ltd. (1991); Woodgett, J. R., Protein Kinases, Oxford University Press (1995); Balch, W. E. et al., Methods in Enzymology, Vol. 257, Pt. C: “Small GTPases and Their Regulators: Proteins Involved in Transport,” Academic Press (1995); The Chaperonins, Academic Press (1996); Pelech, L., Protein Kinase Circuitry in Cell Cycle Control, R G Landes (1996); Atkinson, Regulatory Proteins of the Complement System, Franklin Press (1992); Cooke, D. T. et al., Transport and Receptor Proteins of Plant Membranes: Molecular Structure and Function, Plenum Press (1992); Schumaker, V. N., Advances in Protein Chemistry: Lipoproteins, Apolipoproteins, and Lipases, Academic Press (1994); Brann, M., Molecular Biology of G-Protein-Coupled Receptors: Applications of Molecular Genetics to Pharmacology, Birkhauser (1992); Konig, W., Peptide and Protein Hormones: Structure, Regulations, Activity—A Reference Manual, VCH Publ. (1992); Tuboi, S. et al., Post-Translational Modification of Proteins, CRC Press (1992); Heilmeyer, L. M., Cellular Regulation by Protein Phosphorylation, Springer-Verlag (1991); Takada, Y., Integrin: The Biological Problem, CRC Press (1994); Ludlow, J. W., Tumor Suppressors: Involvement in Human Disease, Viral Protein Interactions, and Growth Regulation, R G Landes (1994); Schlesinger, M. J., Lipid Modification of Proteins, CRC Press (1992); Nitsch, R. M., Alzheimer's Disease. Amyloid Precursor Proteins, Signal Transduction, and Neuronal Transplantation, New York Academy of Sciences (1993); Cochrane, C. G., et al., Cellular and Molecular Mechanisms of Inflammation, Vol. 3: Signal Transduction in Inflammatory Cells, Part A, Academic Press (1992); Gupta, S. et al., Mechanisms of Lymphocyte Activation and Immune Regulation IV: Cellular Communications, Plenum Press (1992); Authi, K. S. et al., Mechanisms of Platelet Activation and Control, Plenum Press (1994); Grunicke, H., Signal Transduction Mechanisms in Cancer, R G Landes (1995); Latchman, D. S., Eukaryotic Transcription Factors, Academic Press (1995).
- The term “contacting a target molecule” refers broadly to placing the target molecule in solution with the molecule to be screened for binding or with the condition(s) to be tested for stabilizing the target molecule. Less broadly, contacting refers to the turning, swirling, shaking or vibrating of a solution of the target molecule and the molecule to be screened for binding. More specifically, contacting refers to the mixing of the target molecule with the molecule to be tested for binding. Mixing can be accomplished, for example, by repeated uptake and discharge through a pipette tip, either manually or using an automated pipetting device. Preferably, contacting refers to the equilibration of binding between the target molecule and the molecule to be tested for binding. Contacting can occur in the container, infra, or before the target molecule and the molecule to be screened are placed in the container.
- The target molecule may be contacted with a nucleic acid prior to being contacted with the molecule to be screened for binding. The target molecule may be complexed with a peptide prior to being contacted with the molecule to be screened for binding. The target molecule may be phosphorylated or dephosphorylated prior to being contacted with the molecule to be screened for binding.
- A carbohydrate moiety may be added to the target molecule before the target molecule is contacted with the molecule to be screened for binding. Alternatively, a carbohydrate moiety may be removed from the target molecule before the target molecule is contacted with the molecule to be screened for binding.
- The term “container” refers to any vessel or chamber in which the receptor and molecule to be tested for binding can be placed. The term “container” encompasses reaction tubes (e.g., test tubes, microtubes, vials, etc.).
- The term “biological sample” refers to the contents of a container.
- “Spectral emission,” “thermal change,” and “physical change” encompass the release of energy in the form of light or heat, the absorption of energy in the form or light or heat, changes in turbidity and changes in the polar properties of light. Specifically, the terms refer to fluorescent emission, fluorescent energy transfer, absorption of ultraviolet or visible light, changes in the polarization properties of light, changes in the polarization properties of fluorescent emission, changes in the rate of change of fluorescence over time (i.e., fluorescence lifetime), changes in fluorescence anisotropy, changes in fluorescence resonance energy transfer, changes in turbidity, and changes in enzyme activity. Preferably, the terms refer to fluorescence, and more preferably to fluorescence emission. Fluorescence emission can be intrinsic to a protein or can be due to a fluorescence reporter molecule. The use of fluorescence techniques to monitor protein unfolding is well known to those of ordinary skill in the art. For example, see Eftink, M. R., Biophysical J. 66:482-501 (1994).
- “Biochemical conditions” encompass any component of a physical, chemical, or biochemical reaction. Specifically, the term refers to conditions of temperature, pressure, protein concentration, pH, ionic strength, salt concentration, time, electric current, potential difference, concentrations of cofactor, coenzyme, oxidizing agents, reducing agents, detergents, metal ion, ligands, or glycerol.
- It is therefore intended that the foregoing detailed description be regarded as illustrative rather than limiting, and that it be understood that it is the following claims, including all equivalents, that are intended to define the spirit and scope of this invention.
- Compounds that have a Fluorescent Readout
- The compounds of the present invention undergo enhanced fluorescence in nonpolar environments as compared to polar environments. Examples of nonpolar environments include nonpolar solvents and hydrophobic proteins or membranes.
- Compounds in accordance with the present invention are of the formula (I):
- where R4, R5, R6, R7, R8, and R9 are each independently hydrogen, fluorine, or alkyl, with the proviso that one of R6 or R7 is —NR1R2, —OR1, or —SR1;
- R1 and R2 are each independently hydrogen or alkyl; or
- R1 and R2 together with the nitrogen to which they are attached, may form a substituted or unsubstituted 5- or 6-membered ring; wherein said nitrogen is not conjugated with said 5- or 6-membered ring;
- R3 is hydrogen or a N-protecting group;
- n is 1, 2, 3 or 4.
- Preferably one of R6 or R7 is —NR1R2. More preferably, R7 is —NR1R2. Preferably, R1 or R2 is alkyl. More preferably, R1 and R2 are both alkyl. Even more preferably, R1 and R2 are both methyl, ethyl or propyl. Preferably, R1 and R2 together with the nitrogen to which they are attached are pyrrolidinyl, piperidinyl, or morpholinyl. Preferably R3 is a N-protecting group. More preferably R3 is Boc, Cbz, or Fmoc. Even more preferably R3 is Fmoc.
- In a preferred embodiment, the compound of the present invention can have the formula (II):
- where R1, R2, and R3 are described above for formula (I).
- In another embodiment, the compound of the present invention can have the formula (IV):
- where n, R4, R5, R6, R7, R8, and R9 are described above for formula (I);
- Y is NH2 or C(O)X; and
-
- X is halogen, hydroxy or alkoxy.
- In another embodiment, the compound of the present invention can have the formula (V):
- where n, R4, R5, R6, R7, R8, and R9 are described above for formula (I).
- In another embodiment, the compound of the present invention can have the formula (VI):
- where n, R4, R5, R6, R7, R8, and R9 are described above for formula (I) and X is described in formula (IV).
- In another embodiment, the compound of the present invention can have the formula (VII):
- where n, R4, R5, R6, R7, R8, and R9 are described above for formula (I).
- In another embodiment, the compound can have the formula (VIII):
- In another embodiment, the compound can be a peptide containing the formula (I):
- where n, R1, R2, R3, R4, R5, R6, R7, R8, and R9 are described above for formula (I).
- In another preferred embodiment, the compound can be a peptide containing the formula (II):
- where R1, R2, and R3 are each described above for formula (I).
- In another preferred embodiment, the compound can be a peptide containing the formula (III):
- where R3 is described above for formula (I). When R3 is hydrogen, compound (III) is referred to herein as Dap(6DMN). When R3 is Fmoc, compound (III) is herein referred to as Fmoc-Dap(6DMN).
- Compounds of the present invention are useful as fluorescence probe molecules in applications wherein fluorescence probes are known to be useful. In using a compound of the present invention as a fluorescence probe molecule, the compound is added to a sample to be probed. The sample comprising the compound is then exposed to a light source. The light source produces light that is limited to a range of wavelengths. The range of wavelengths is between about 360 and about 410 nanometers (nm), preferably between about 370 and about 390 nm, most preferably between about 375 and about 380 nm. Upon exposure to a light source, the compound of the present invention is fluorescent and emits fluorescent energy. The emitted fluorescent energy is detected using methods well known in the art. The intensity and wavelength of the emitted fluorescent energy provides information about the sample. The emitted fluorescent energy preferably has a range of wavelengths between about 480 and about 640 nm, preferably between about 560 and about 590 nm.
- The fluorescence of a molecule is defined by the quantum yield. The quantum yield is the ratio of the photons absorbed by the compound to the photons emitted through fluorescence by the compound. Compounds of the present invention have quantum yields that are preferably low in aqueous solutions and high in non-polar environments. Quantum yields range from about 0.001 and about 0.1, preferably between about 0.001 and about 0.005 for aqueous solutions, preferably between about 0.2 and about 0.7 for non-polar environment.
- Fluorescence can also be evaluated by determining the dipole moment change between the ground and excited state. The change in the dipole moment can be estimated from a plot of the Stokes shift vs. the orientation polarizability, known as a Lippert-Mataga plot. (Lippert, V. E. Z. Elektrochem. 1957, 61, 962-975; Mataga, N.; Kaifu, Y.; Koizumi, M. Bull. Chem. Soc. Jpn. 1956, 29, 465-470.) The relationship between the Stokes shift (Δ
ν ) and the orientation polarizability (Δf) and is expressed by the following equation: -
- where c is the velocity of light, h is Plank's constant, c is the solvent dielectric constant, n is the solvent refraction index, μe−μg is the difference between the dipole moments of the excited and the ground states respectively (Δμ), and a0 is the radius of the Onsager cavity around the compound. The magnitude of Δμ correlates to the sensitivity of the fluorescent probe, where larger changes in the dipole moment correlate to greater sensitivity.
- Fluorescence is sensitive to the pH of the surrounding environment. Compounds of the present invention are useful as fluorescence probes in the pH range from about 4 to about 8.
- Compounds of the present invention are useful in monitoring biological interactions. Biological interactions play important roles in the sequence and mechanisms of action of various cellular processes and signal pathways. The time course, nature, and sequence of the different cellular processes can be elucidated by in situ observation using the compounds of the present invention. Specific inhibitors and/or activators of the cellular processes and signal pathways being studied may optionally be used in addition to compounds of the present invention.
- Biological interactions, as defined herein, comprise the interaction of a compound or molecule with a target molecule. Examples of target molecules include peptides, proteins, enzymes, nucleic acids, ions, and other receptors; preferably metal ion chelators, proteases, polymerases, hydrolases, phosphatases, and kinases; more preferably protein domains, and protein domains of phosphatases and kinases.
- Proteins and protein-protein interactions play a central role in the various essential biochemical processes. For example, these interactions are evident in the interaction of hormones with their respective receptors, in the intracellular and extracellular signaling events mediated by proteins, in enzyme substrate interactions, in intracellular protein trafficking, in the formation of complex structures like ribosomes, viral coat proteins, and filaments, and in antigen-antibody interactions. These interactions are usually facilitated by the interaction of small regions within the proteins that can fold independently of the rest of the protein. These independent units are called protein domains. Abnormal or disease states can be the direct result of aberrant protein-protein interactions. Protein-protein interactions are also central to the mechanism of a virus recognizing its receptor on the cell surface as a prelude to infection. Identification of domains that interact with each other not only leads to a broader understanding of protein-protein interactions, but also aids in the design of inhibitors of these interactions.
- Phosphorylation-dependent peptide-protein interactions include phosphoserine peptides with 14-3-3, which is a protein involved in cell cycle control (Muslin, A. J., Tanner, J. W., Allen, P. M., Shaw, A. S. Cell 1996, 84, 889-897), and phosphotyrosine peptides with SH2 domains. SH2 domains are binding modules that are involved in tyrosine kinase signaling networks and recognize phosphotyrosine-containing peptide sequences. The phosphotyrosine binding is complemented by simultaneous peptide-protein interactions on the protein surface. Examples of SH2 domains include Abl SH2, Crk SH2, and C-terminal PI3K SH2 which can be expressed in bacteria as GST fusion proteins, which are referred to as GST-Abl SH2, GST-Crk SH2, and GST-PI3K SH2.
- Recognition sequences for SH2 domains comprise a phosphotyrosine residue and other amino acids. The recognition sequence is different for different SH2 domains. Amino acid recognition sequences for binding members of the SH2 domain family are disclosed in Songyang, Z. et al.; Cell 1993, 72, 767-778. For the Crk SH2 domain, the recognition sequence is pTyr-Asp-His-Pro. For the Abl SH2 domain, the recognition sequence is pTyr-Glu-Asn-Val.
- Compounds of the formula (I) are useful for studying the peptide-protein interactions on the protein surface of the SH2 domain. Compounds of formula (I) can be incorporated into peptides containing the desired SH2 recognition sequence. Table 1 shows peptides incorporating the Crk SH2 or Abl SH2 recognition sequences and Dap(6DMN) into the (+2) position relative to the phosphotyrosine residue.
-
TABLE 1 Peptide Sequences and Corresponding SH2 Domain Target Target Peptide SH2 Peptide sequence Crk-bp Crk Ac-Glu-Dap(6DMN)-Gln-pTyr-Asp-His-Pro-Asn- Ile-(CONH2) Crk-bp2 Crk Ac-Glu-Dap(6DMN)-Gly-pTyr-Asp-His-Pro-Asn- Ile-(CONH2) Abl-bp Abl Ac-Glu-Dap(6DMN)-Gly-pTyr-Glu-Asn-Val-Gln- Ser-(CONH2) Abl-bp2 Abl Ac-Glu-Dap(6DMN)-pTyr-Glu-Asn-Val-Gln-Ser- (CONH2) - The peptides of Table 1 were incubated with targeted and nontargeted SH2 domains. The binding of peptides Crk-bp, Crk-bp2, Abl-bp, and Abl-bp2 to SH2 target domains can be studied by fluorescence titration as shown in
FIGS. 1-4 . - Fluorescence titrations of Crk-bp and Crk-bp2 with targeted GST-Crk SH2 are shown in
FIG. 1 (a) and (b), respectively. Fluorescence titrations of Crk-bp and Crk-bp2 with non-targeted GST-Abl SH2 are shown inFIG. 4 (a) and (b), respectively.FIG. 1 (c) shows relative fluorescence emission intensities for Crk-bp and Crk-bp2 peptides, Crk-bp and Crk-bp2 bound to GST-Crk SH2 domain, and Crk-bp2 bound to non-target domains GST-Abl SH2 and GST-PKI3 SH2. The relative fluorescence emission intensities for non-target SH2 domains is notably less than for the targeted GST-Crk SH2 domain. - Fluorescence titrations of Abl-bp with targeted GST-Abl SH2 is shown in
FIG. 2 (a). Fluorescence titrations of Abl-bp with non-targeted GST-PI3K SH2 and GST-Abl SH2 are shown inFIG. 2 (b) and 4 (c), respectively. The relative fluorescence emission intensities for the Abl-bp peptide and aforementioned SH2 domain titrations are shown inFIG. 2 (c). - Fluorescence titrations of Abl-bp2 with targeted GST-Abl SH2 is shown in
FIG. 3 (a). Fluorescence titrations Abl-bp2 with non-targeted GST-PI3K SH2 and GST-Crk SH2 are shown inFIGS. 3 (b) and 4 (d), respectively. The relative fluorescence emission intensities for the Abl-bp2 peptide and aforementioned SH2 domain titrations are shown inFIG. 3 (c). -
TABLE 2 Kinetic and Fluorescence Properties of Peptides containing Dap(6DMN) Target Peptide Fluorescence Peptide SH2 Sequence Kd(μM) increase Crk-bp Crk Ac-Glu-Dap(6DMN)-Gln- 4.8 ± 2 a 500% b pTyr-Asp-His-Pro-Asn-Ile- (CONH2) Crk-bp2 Crk Ac-Gly-Dap(6DMN)-Gln- 2.4 ± 2 a 1100% b pTyr-Asp-His-Pro-Asn-Ile- (CONH2) Abl-bp Abl Ac-Gly-Dap(6DMN)-Gln- 13 ± 2 a 400% c pTyr-Glu-Asn-Val-Gln-Ser- (CONH2) Abl-bp2 Abl Ac-Dap(6DMN)-Gln-pTyr- 12 ± 2 a — Glu-Asn-Val-Gln-Ser- (CONH2) a Assay conditions: 20 μM in PBS buffer, pH 7.5. b Excitation wavelength: 375 nm, Emission wavelength, 565 nm. c Excitation wavelength: 375 nm, Emission wavelength, 599 nm. - Compounds of the present invention are also useful in a method of monitoring biological interactions, comprising providing a compound of the present invention, contacting a target molecule with the compound to form a biological sample, and monitoring the fluorescence of the biological sample. Preferably, the compound is a peptide. More preferably, the compound is a peptide containing at least one of formula (I), (II), and (III).
- In the methods of the present invention, the monitoring step comprise contacting the compound with the one or more target molecules or different biochemical conditions, wherein the measuring step comprises exciting the compound of the present invention with light, and measuring the fluorescence.
- In all methods of using a compound of the present invention, the concentration used will depend on the detection equipment. Typically, the concentration of the compound is from greater than about 0.1 nM.
- The compounds of formula (I) can be prepared according to the reaction as shown in
Scheme 1, wherein, n, R3, R4, R5, R6, R7, R8 and R9 are as defined above. Anhydride (1) is stirred with an appropriate diamine (2) in a suitable solvent to formcompound 3. The allyl protecting group ofcompound 3 can be removed under standard conditions known to those skilled in the art, examples of which are disclosed in Greene and Wuts, Protective Groups in Organic Synthesis” 2nd ed.; John Wiley & Sons, New York, 1991) to afford a compound of Formula I (4). Thediamine compound 2 can be prepared from the corresponding diamino acid using standard conditions for protecting a carboxylic acid with an allyl protecting group known to those skilled in the art, examples of which are provided in Greene and Wuts. -
Anhydride 1 can be prepared by stirringdiester 5 with an appropriate base in a suitable solvent.Compound 5 can be assembled by methods known to one skilled in the art such ring annulation methods or Diels-Alder reaction. - The compounds of formula (IV) can be prepared by stirring anhydride (1) with an appropriate amine in a suitable solvent to form compounds of formula (IV), wherein, n, R4, R5, R6, R7, R8, R9 and Y are as defined above. In the case where Y is NH2, the amine is diamino compound (6) as shown in
Scheme 3 which is stirred with anhydride (1) to for compound (7). Diamines are commercially available from vendors such as Aldrich, for example. Alternatively,compound 7 may be prepared by stirring anhydride (1) with a diamine where one of the nitrogens is protected with a suitable N-protecting group. After formation of the imide bond, the N-protecting group can be removed to afford compound (7). Methods for introduction and removal of N-protecting groups are known to those skilled in the art, examples of which are disclosed in Greene and Wuts, Protective Groups in Organic Synthesis, 2nd ed.; John Wiley & Sons, New York, 1991. - The compounds of formula (IV) wherein, n, R4, R5, R6, R7, R8, R9 are as defined above as Y is C(O)X, can be prepared as shown in
scheme 4.Amino acid 8 can be carboxyl protected with a suitable protecting group, then stirred with anhydride (1) in a suitable solvent. Removal of the carboxyl protecting group under standard conditions afford compound (9). Methods for introduction and removal of N-protecting groups are known to those skilled in the art, examples of which are disclosed in Greene and Wuts, Protective Groups in Organic Synthesis, 2nd ed.; John Wiley & Sons, New York, 1991. Compound (10) where X is halogen or alkoxy can be prepared under standard conditions acyl halide (Org. Syn. Coll. Vol. 1 1941, 12, 147; Org. Syn. Coll. Vol. 3 1955, 169, 490, 547, 555, 712; Org. Syn. Coll.Vol 4 1963, 154, 263, 339, 715, 739) or esterification (Tetrahedron. 1980, 36, 2409) conditions. - Peptides containing the formula (I) can be prepared using standard peptide coupling techniques known to those skilled in the art. Examples are provided in Bodansky, M. Peptide Chemistry a Practical Textbook, 2nd ed.; Springer-Verlag, Berlin, 1993. Dap(6DMN), can incorporated into peptides using Fmoc-Dap(6DMN) and standard Fmoc solid-phase peptide synthesis.
- The sensor of the present invention can be used in a method for detecting biological interactions. The method of the present invention comprises providing a peptide incorporating an amino acid of the formula (I), contacting a target molecule with the peptide to form a biological sample, and monitoring the fluorescence of the biological sample.
- It is therefore intended that the foregoing detailed description be regarded as illustrative rather than limiting, and that it be understood that it is the following claims, including all equivalents, that are intended to define the spirit and scope of this invention.
- All peptide synthesis reagents and amino acid derivatives were purchased from Applied Biosystems or Novabiochem. All other chemicals were purchased from Aldrich. Dichloromethane was distilled from calcium hydride under nitrogen, and tetrahydrofuran was distilled from sodium under argon. Analytical thin-layer chromatography (TLC) was carried out on F254 250-μm silica gel plates, and visualized by UV.
- 1H and 13C NMR spectra were acquired on a 400 MHz Bruker spectrometer. NMR samples were prepared in deuterated chloroform (CDCl3) unless otherwise stated. Chemical shifts are reported in ppm relative to the internal standard (tetramethyl silane for 1H), and J values are reported in Hz. Electrospray Ionization Mass Spectrometry (ESIMS) was performed on a PerSeptive Biosystems Mariner™ Biospectrometry Workstation (Turbo Ion Source). Fluorescence spectroscopy measurements were made using a Fluoromax-P spectrofluorimeter controlled by the DataMax 2.20 software, and coupled to a NesLab RTE-111 water bath for temperature control. All measurements were made at 20° C.
-
- Protection of the aldehyde was performed following reported procedures. (Borchardt, R. T.; Huber, J. A.; Houston, M. J. Med. Chem. 1982, 25, 258; Brasili, L.; Sorbi, C.; Franchini, S.; Manicardi, M.; Angeli, P.; Marucci, G.; Leonarke, A.; Poggesi, E. J. Med. Chem. 2003, 46, 1504) 3-Nitrobenzaldehyde (4.0 g, 26.4 mmol) was dissolved in toluene (200 mL). Following this, ethanediol (3.0 mL, 53 mmol, 2.0 eq) was added to the solution and the resulting mixture was refluxed overnight with a Dean-Stark trap. The reflux mixture was then extracted with saturated NaHCO3 (2×75 mL). The organic layer was dried with Na2SO4 and concentrated under reduced pressure. The residue was then purified by flash column chromatography (30% EtOAc/hexanes) to afford 3-nitrophenyl-1,3-dioxolane as a clear yellow oil that solidified upon standing. (5.09 g, 98%, Rf=0.7 40% EtOAc/hexanes). 1H-NMR (400 MHz, CDCl3, δ): 4.0-4.2 (m, 4H), 5.87 (s, 1H), 7.55 (t, 1H, J=7.93 Hz), 7.79 (d, 1H, J=7.7 Hz), 8.21 (dd, 1H, J1=1.2 Hz, J2=8.2 Hz), 8.34 (s, 1H). 13C-NMR (100 MHz, CDCl3, δ): 148.4, 140.5, 132.8, 129.6, 124.1, 121.8, 102.3, 65.6.
- 3-Nitrophenyl-1,3-dioxolane (1.0 g, 5.1 mmol) and chloromethyl-phenyl sulfone (975 mg, 5.1 mmol) were dissolved in dry dimethyl sulfoxide. Then, KOH powder (2.00 g) was added to the solution, which immediately turned dark purple. The resulting mixture was stirred overnight at room temperature, then poured over 75
mL 2% aqueous HCl and extracted with EtOAc (3×50 mL). The combined organic layers were dried with Na2SO4 and concentrated under reduced pressure to give a dark brown oil that was purified by flash column chromatography (30% EtOAc/hexanes). Two regioisomeric products were isolated. The combined yield of the reaction is 1.23 g (35% desired product, Rf=0.4 40% EtOAc/hexanes). Compounds ortho- and para- could be identified by the 1H-NMR signals of the protons ortho- to the nitro group. In the case of the desired compound, 2-(2-phenylsulfonylmethyl-5-nitrophenyl)-1,3-dioxolane, both signals are present in the NMR spectra. 1H-NMR (400 MHz, CDCl3, δ): 4.01 (s, 4H), 4.73 (s, 2H), 5.93 (s, 1H), 7.32 (d, 1H, J=8.3 Hz), 7.53 (t, 2H, J=7.6 Hz), 7.67 (t, 1H, J=7.7 Hz), 7.72 (d, 2H, J=7.6 Hz), 8.10 (d, 1H, J=6.6 Hz), 8.45 (s, 1H). 13C-NMR (100 MHz, CDCl3, δ): 148.1, 140.7, 138.2, 134.5, 133.7, 129.5, 128.6, 123.6, 121.7, 100.2, 65.5, 59.0. HRMS-ESI (m/z): [M+H+] calcd for C16H15NO6S 350.0693, found, 350.0677. - 2-(2-Phenylsulfonylmethyl-5-nitrophenyl)-1,3-dioxolane (400 mg, 1.14 mmol) was dissolved in AcOH (8 mL) and water (2 mL). The solution was refluxed for 5 h and then concentrated under reduced pressure. The resulting solid was confirmed as the desired product by NMR and used in the next step without further purification. In a 25 mL round bottom flask, 5-nitro-2-phenylsulfonylmethylbenzaldehyde (300 mg, 1.00 mmol), diethylmalate (195 μL, 207 mg, 1.20 mmol) and 18-crown-6 (13 mg, 0.05 mmol) were dissolved in acetonitrile (10 mL). K2CO3 (700 mg) was added and the resulting mixture was stirred for 30 min. The reaction was refluxed for 3 h, then filtered over celite and concentrated under reduced pressure. The resulting dark oily residue was purified by flash chromatography (30% EtOAc/hexanes) to give diethyl 6-nitro-2,3-naphthalenedicarboxylate (231 mg, 64%, Rf=0.35 20% EtOAc/hexanes). 1H-NMR (400 MHz, CDCl3, δ): 1.42 (t, 3H, J=7.1 Hz), 1.43 (t, 3H, J=7.1 Hz), 4.45 (c, 4H, J=7.1 Hz), 8.07 (d, 1H, J=9.0 Hz), 8.31 (s, 1H), 8.37 (dd, 1H, J1=9.0 Hz, J2=2.1 Hz), 8.47 (s, 1H), 8.87 (d, 1H, J=1.8 Hz). 13C-NMR (100 MHz, CDCl3, δ): 167.1, 166.7, 147.2, 135.9, 132.8, 132.3, 131.9, 130.9, 130.5, 129.9, 129.7, 125.13, 121.9, 62.36, 14.33. HRMS-ESI (m/z): [M+H+] calcd for C16H15NO6 318.0972, found, 318.0986.
- Diethyl 6-nitro-2,3-naphthalenedicarboxylate (350 mg, 1.11 mmol) was dissolved in MeOH (50 mL). Formalin (7 mL) was added and Pd/
C 10% (100 mg) was added to the solution. The resulting mixture was stirred under hydrogen for 2.5 h until TLC showed that the starting material has been consumed. The reaction mixture was then filtered through celite and partly concentrated to approximately 20 mL. Following this, the mixture was poured over 2% aqueous NaHCO3 (150 mL) and extracted (3×50 mL EtOAc). The combined organic layers were dried with Na2SO4 and concentrated under reduced pressure to give diethyl 6-(dimethylamino)-2,3-naphthalenedicarboxylate (295 mg, 84%, Rf=0.5 30% EtOAc/hexanes). 1H-NMR (400 MHz, CDCl3, δ): 1.38 (t, 3H, J=7.1 Hz), 1.39 (t, 3H, J=7.1 Hz), 3.03 (s, 6H), 4.37 (c, 2H, J=7.1 Hz), 4.40 (c, 2H, J=7.1 Hz), 6.84 (s, 1H), 7.15 (dd, 1H, J1=9.1 Hz, J2=2.5 Hz), 7.7 (d, 1H, J=9.1 Hz), 7.88 (s, 1H), 8.17 (s, 1H). 13C-NMR (100 MHz, CDCl3, δ): 169.1, 167.6, 150.16, 135.7, 130.7, 130.4, 129.8, 127.4, 125.7, 123.2, 117.6, 105.8, 61.5, 61.3, 40.4, 14.4, 14.3. HRMS-ESI (m/z): [M+H+] calcd for C18H21NO4 316.1549, found, 316.1556. - Diethyl 6-(dimethylamino)-2,3-naphthalenedicarboxylate (373 mg, 1.2 mmol), was refluxed in 1:5 MeOH: 40% KOH (25 mL) for 3 h. The reaction mixture was then poured over water (150 mL), acidified with 6 M HCl to
pH 2 and extracted with EtOAc (5×30 mL). The combined organic layers were dried with Na2SO4 and concentrated under reduced pressure. The resulting solid residue, corresponding to the diacid intermediate, was used in the next step without purification (228 mg, 85%). The diacid was placed in a sublimation apparatus and heated at 175° C. under vacuum for 4 h. 6-(Dimethylamino)-2,3-naphthalenedicarboxylic anhydride was recovered after sublimation as a bright yellow solid (213 mg, 77% overall yield, Rf=0.6 40% EtOAc/hexanes). 1H-NMR (400 MHz, CDCl3, δ): 3.19 (s, 6H), 7.02 (d, 1H, J=2.4 Hz), 7.34 (dd, 1H, J1=9.2 Hz, J2=2.6 Hz), 7.92 (d, 1H, J=9.2 Hz), 8.20 (s, 1H), 8.30 (s, 1H). HRMS-ESI (m/z): [M+H+] calcd for C14H11NO3 242.0812, found, 242.0805. For diacid intermediate: HRMS-ESI (m/z): [M+H+] calcd for C14H13NO4 258.0772, found, 252.0764. -
- Fmoc-Dap(Boc)-OH (450 mg, 1.05 mmol) was dissolved in MeOH (15 mL). Cs2CO3 (172 mg, 0.53 mmol) was added and the resulting solution was stirred at room temperature for 10 minutes. The reaction mixture was concentrated under reduced pressure and redissolved in DMF (30 mL). Allyl bromide (280 μL, 3.3 mmol) was added and the mixture was stirred for 1 h at room temperature. The reaction was poured over 2% aqueous NaHCO3 (150 mL) and extracted with EtOAc (4×50 mL). The combined organic layers were dried with Na2SO4 and concentrated under reduced pressure. The resulting brown oily residue was purified by flash column chromatography (1% MeOH/CH2Cl2, Rf=0.6 5% MeOH/CH2Cl2) to give allyl N-α-Fmoc-N-β-Boc-L-diaminopropionate as a white powder (516 mg, 78%). Rf=0.7 (1% MeOH/CH2Cl2). 1H NMR (400 MHz, CDCl3, δ): 1.34 (s, 9H), 3.48 (br s, 2H), 4.12 (br s, 1H), 4.2-4.4 (m, 2H), 4.55 (br s, 2H), 4.9 (br s, 1H), 5.14 (d, 1H, J=10.3 Hz), 5.22 (d, 1H, J=17.1 Hz), 5.8 (br s, 1H), 5.9 (d, 1H, 5.8 Hz), 7.18 (t, 2H, J=7.3 Hz), 7.27 (t, 2H, J=7.4 Hz), 7.5 (br s, 2H), 7.65 (d, 2H, J=7.4 Hz). 13C NMR (400 MHz, CDCl3, δ): 170.4, 156.6, 156.2, 143.8, 141.4, 131.6, 127.8, 127.2, 125.3, 120.9, 119.1, 80.1, 67.3, 66.5, 55.3, 47.2, 42.2, 28.4. HRMS-ESI (m/z): [M+H+] calcd for C26H30N2O6 467.2177, found, 467.2177.
- Allyl N-α-Fmoc-N-β-Boc-L-diaminopropionate (175 mg, 0.38 mmol) was dissolved in dry dichloromethane (10 mL), and the solution was cooled to 0° C. Then, TFA (10 mL) was added dropwise and the resulting mixture was stirred at 0° C. for 30 minutes and then at room temperature for 1 h. The reaction was concentrated under reduced pressure, redissolved in toluene and concentrated again. The resulting amine was placed under vacuum for 20 minutes and dissolved in dry DMF (3 mL). DIEA (360 μL, 2.6 mmol) and 6-(dimethylamino)-2,3-naphthalenedicarboxylic anhydride were added, and the mixture was stirred at room temperature for 30 minutes. HOBt/HBTU (2 mL 0.2 M solution in DMF) were added and the reaction was stirred at room temperature for 12 h. HOBt/HBTU (1 mL 0.2 M solution in DMF, 0.5 eq.) was added and the reaction mixture was allowed to stir for another 1 hour. The crude reaction was added to 150
mL 2% aqueous NaHCO3 and extracted with EtOAc (3×30 mL). The combined organic layers were dried with Na2SO4 and concentrated under reduced pressure. The residue was loaded in 20% EtOAc/toluene onto a silica column and purified (gradient 20 to 50% EtOAc/hexanes) to afford Fmoc-Dap(6DMN)—O-Allyl as a bright yellow oil (220 mg, >95% yield, Rf=0.8 10% MeOH/CH2Cl2). 1H-NMR (400 MHz, CDCl3, δ): 3.12 (s, 6H), 4.1-4.4 (m, 5H), 4.65-4.83 (m, 3H), 5.26 (d, 1H, J=10.3 Hz), 5.38 (d, 1H, J=17.1 Hz), 5.9-6.1 (m, 1H), 7.20 (dd, 1H, J1=2.0 Hz, J2=9.1 Hz), 7.29 (c, 2H, J=7.4 Hz), 7.29 (td, 2H, J1=1.7 Hz, J2=5.2 Hz), 7.57 (d, 1H, J=7.6 Hz), 7.60 (d, 1H, J=7.5 Hz), 7.73 (d, 2H, J=7.5 Hz), 7.80 (d, 1H, J=9.1 Hz), 8.03 (s, 1H), 8.10 (s, 1H). 13C-NMR (100 MHz, CDCl3, δ): 169.9, 168.6, 168.5, 156.0, 150.5, 144.1, 143.8, 141.3, 137.7, 131.5, 128.2, 127.7, 127.4, 127.2, 125.2, 125.5, 123.1, 122.5, 120.0, 119.3, 117.9, 107.9, 67.5, 66.8, 53.8, 47.1, 40.4, 39.2. HRMS-ESI (m/z): [M+H+] calcd for C35H31N3O6 590.2291, found, 590.2270. - Fmoc-Dap(6DMN)-O-allyl (226 mg, 0.383 mmol) was dissolved in dry CH2Cl2 (12 mL). Phenyl silane (1.2 mL, 9.6 mmol) and Pd(PPh3)4 (20 mg, 0.014 mmol) were added and the reaction mixture was stirred at room temperature for 30 minutes. The reaction mixture was then directly loaded onto a silica flash column and the desired product purified (gradient, 3% to 5% MeOH/CH2Cl2). Fmoc-Dap(6DMN) was isolated as a yellow solid (210 mg, >95%, Rf=0.4 10% MeOH/CH2Cl2). MS-ESI: m/z: 550.1 (MH+), 572 (MNa+), 1099.3 (M2H+). HRMS-ESI (m/z): [M+H+] calcd for C32H27N3O6 550.1973, found, 550.1982.
-
- Glycine methyl ester hydrochloride (21 mg, 0.164 mmol) was dissolved in DMF (2 mL), and DIEA (450 μL, 3.28 mmol, 10 eq.) and 6-(dimethylamino)-2,3-naphthalenedicarboxylic anhydride (80 mg, 0.328 mmol) were added. The resulting mixture was stirred at room temperature for 10 minutes. HOBt/HBTU (1 mL, 0.2 M in DMF) was added and the reaction mixture was stirred for 12 h. Then for an additional 0.5 mL of the HOBt/HBTU mixture were added, and the reaction was stirred for another hour. The reaction was poured over 100
mL 1% aqueous HCl and extracted with EtOAc (2×50 mL). The combined organic layers were dried with Na2SO4 and concentrated under reduced pressure. The solid residue was purified by flash column chromatography (EtOAc/hexanes 35%) to afford 6DMN-GlyOMe as a bright yellow solid in quantitative yield (Rf=0.5 40% EtOAc/hexanes). 1H-NMR (400 MHz, CDCl3, δ): 3.12 (s, 6H), 3.76 (s, 3H), 4.47 (s, 2H), 6.96 (d, 1H, J=2.0 Hz), 7.21 (dd, 1H, J=9.1 Hz, J2=2.5 Hz), 7.81 (d, 1H, J=9.1 Hz), 8.06 (s, 1H), 8.13 (s, 1H). 13C-NMR (100 MHz, CDCl3, δ): 168.1, 167.9, 167.8, 150.6, 137.8, 131.4, 128.5, 127.5, 125.2, 123.1, 122.8, 117.9, 107.9, 52.8, 40.4, 38.9. HRMS-ESI (m/z): [M+H+] calcd for C17H16N2O4 313.1188, found, 313.1201. UV: ε (388 nm, M−1 cm−1)=7980. - Peptide synthesis was carried using standard Fmoc-based solid phase peptide synthesis (SPPS) protocols on a 0.05 to 0.1 mmol scale using a 0.21 mmol/g loading PAL-PEG-PS solid support. Amino acids were coupled in three-fold excess using a mixture of 0.2 M HBTU/0.2 M HOBt in DMF as activating agents. Each amino acid was activated for two minutes with the HBTU/HOBt mixture (1 eq.) and diisopropylethylamine (DIPEA), 0.195 M in DMF (1.5 eq.) before being added to the resin. Peptide coupling was monitored using the 2,4,6-trinitrobenzenesulphonic acid (TNBS) test. (Hancock, W. S.; Battersby, J. E. Anal. Biochem. 1976, 71, 260-264) Amino acids were used as protected Fmoc-amino acids with the standard side chain protecting groups: Fmoc-Ala-OH, Fmoc-Arg(Pbf)-OH, Fmoc-Asp(OtBu)-OH, Fmoc-Glu(OtBu)-OH, Fmoc-Ile-OH, Fmoc-Ser(tBu)-OH, Fmoc-Thr(tBu)-OH, Fmoc-Tyr(tBu)-OH, Fmoc-Val-OH and Fmoc-Dap(6DMN). Phosphotyrosine was introduced as the monobenzyl ester Fmoc-Tyr(PO(OBzl)OH)—OH. High-performance liquid chromatography (HPLC) was performed using a Waters 600E HPLC fitted with a
Waters 600 automated control module and a Waters 2487 dual wavelength absorbance detector recording at 228 and 280 nm. For analytical HPLC a Beckman Ultrasphere C18, 5 μm, 4.6×150 mm reverse-phase column was used. For preparative separations a YMC-pack, C18, 250×20 mm reversed phase column was used. The standard gradient for analytical and preparative HPLC used was 93:7 to 5:95 over 35 minutes (water:acetonitrile, 0.1% TFA). The 6DMN side chain proved resistant to the standard mildly basic amino acid coupling conditions (0.12 M diisopropylethylamine), the Fmoc deprotection conditions (20% piperidine), and the acidic resin cleavage and deprotection cocktail (95% TFA). - Quantum yields (Φ) were measured by comparison with a standard, which emits in a similar region to the test sample. From the available standard compounds quinine sulfate in 0.1 M H2SO4 (Φ=0.54) was selected. (Melhuish, W. B. J. Phys. Chem., 1961, 65, 229-235) Absorbance of the measured solutions did not exceed 0.1 at and above the excitation wavelength to avoid non-linear effects.
- UV absorbance at the excitation wavelength (375 nm), and fluorescence emission spectra of a series of solutions of the standard and compound were recorded. Solutions were prepared with different absorbances, ranging from 0 (solvent blank) to 0.1. A series of graphs of integrated fluorescence vs. absorbance were obtained. The data were fitted to a linear equation with gradient (grad), and intercept=0, See
FIG. 5 as an example of the plots obtained. - The gradients of graphs in
FIG. 5 are proportional to the quantum yield of the different samples. Absolute values were calculated using the standard sample, which has a fixed and known fluorescence quantum yield value, according to the following equation: -
- Where the subscripts std and x denote standard and test respectively. The value Φ is the fluorescence quantum yield, grad is the gradient (grad) from the plot of integrated fluorescence intensity vs. absorbance, and 77 the refractive index of the solvent.
- The UV absorbance spectra (40 μM in methanol) and fluorescence emission spectra of compound (VIII), 6DMN-GlyOMe are shown in
FIG. 6 and tabulated in Table 4. The UV spectrum of 6DMN-GlyOMe has an intense absorption band at 378 nm which is not found inunsubstituted 2,3-naphthalimides. The maximum excitation wavelength of 6DMN-GlyOMe is at 375 nm. -
TABLE 4 Photophysical Properties of 6DMN-GlyOMe in Different Solvents: absorbance emission quantum maximum maximum yield solvent (nm) (nm) (Φ)b watera 388 592 0.002 methanol 382 589 0.012 2-propanol 385 589 0.018 ethanol 379 584 0.027 acetonitrile 380 549 0.135 DMF 380 545 0.155 acetone 375 532 0.148 tetrahydrofuran 373 510 0.147 1,4-dioxane 372 498 0.220 dichloromethane 379 517 0.210 chloroform 380 509 0.225 toluene 373 491 0.208 aBecause of solubility limitations, the quantum yield in water was calculated using peptide Abl-bp. bQuantum yields (Φ) were measured in comparison to quinine sulfate in 0.1 M H2SO4 (Φ = 0.54) as the standard. - Quantum yields (Φ) were measured in the solvents listed in Table 2 by comparison with a standard, which emits in a similar region to 6DMN-GlyOMe. Quinine sulfate in 0.1 M H2SO4 (Φ=0.54) was used as the standard. (Melhuish, W. B. J. Phys. Chem., 1961, 65, 229-235). Solutions were prepared at different concentrations such that the absorbance measurements ranged between 0 and 0.1 at and above the excitation wavelength to avoid non-linear effects.
- UV absorbance at the excitation wavelength (375 nm), and fluorescence emission spectra of a series of solutions of the standard and 6DMN-GlyOMe were recorded. A series of graphs of integrated fluorescence vs. absorbance were obtained. The data were fitted to a linear equation with gradient (grad), and intercept=0. The gradient (grad) is proportional to the quantum yield of the sample. Absolute values were calculated using the standard sample, which has a fixed and known fluorescence quantum yield value, according to the equation above.
- It is known that the solvent sensitivity to polarity can be analyzed in terms of difference in the dipole moments in the ground and the excited state and it has been found that the most sensitive fluorophores are those with the largest changes in the dipole moment. This can be estimated from a Lippert-Mataga plot, which is essentially a plot of the Stokes shift of the fluorescence emission versus the solvent polarity. (Lippert, V. E. Z. Elektrochem. 1957, 61, 962-975; Mataga, N.; Kaifu, Y.; Koizumi, M. Bull. Chem. Soc. Jpn. 1956, 29, 465-470.)
- The difference (Δ
ν ) in the maximum absorption (ν A) and emission wavelengths (ν F), known as the Stokes shift and expressed in wavenumbers, is plotted vs. Δf. The term Δf is called the orientation polarizability and can be calculated according to the following equation: -
- where ε is the solvent dielectric constant and n is the solvent refraction index. The plot of (Δ
ν ) for compound (VIII) vs. Δf for the solvents listed in Table 2 is shown inFIG. 7( b). - The term Δf can also be calculated for a mixture of solvents.
FIG. 7( a) shows the plot of (Δν ) for compound (VIII) vs. Δf for a series of 1,4-dioxane/acetonitrile mixtures. In this case, Δf is obtained for each mixture using ε and n values calculated using the molar fractions (χ) in the equation as follow: -
ε=(χacetonitrile)(38.8)+(χdiox)(2.218) (3) -
n=(χacetonitrile)(1.3442)+(χdiox)(1.4224) (4) - where χacetonitrile+χdiox=1.
- The relationship between Δ
ν and Δf is expressed by the following equation: -
- where c is the velocity of light, h is Plank's constant, μe−μg is the difference between the dipole moments of the excited and the ground states respectively (Δμ), and a0 is the radius of the Onsager cavity around the compound. The Onsager radius for compound VIII was calculated from the optimized structure obtained with a DFT minimization using the Gaussian program (Pople, J. A. et al. Gaussian 98 (Gaussian, Inc., Pittsburgh, Pa., 1998), (B3LYP functional using 6-31G(d) orbital base). The Onsager radius (4.19 Å) was taken as half of the average distance between nitrogen of the amine donor and the two carbonyl oxygens, which corresponds to the longest distance across the molecule where charge separation can take place. (Mukherjee, S., Chattopdhyay, A., Samanta, A., Soujanya, T. J. Phys. Chem. 1994, 98, 2809-2812.) As can be seen in
FIG. 7( a), the Stokes shift of compound VIII in a series of mixtures of 1,4-dioxane/acetonitrile changes linearly in response to the solvent polarity, which correlates with an increase in the dipole moment (Δν ) of 5.5 D. - Lippert plots also give evidence of specific solvent effects for protic solvents. As can be seen in
FIG. 7( b), in most solvents the Stokes shift is proportional to the orientation polarizability. However, protic solvents induce a disproportionately large Stokes shift. This phenomenon has also been observed for other environment-sensitive fluorophores, and has been explained as the result of emission from two different excited states: one with higher dipole moment that is stabilized in polar environments, and another less polar excited state responsible for the emission in less polar media. (Kosower, E. M., Dodiuk, H. J. Am. Chem. Soc. 1974, 96, 6195-6196; Kosower, E. M. Acc. Chem. Res. 1982, 15,259-266; Saha, S., Samanta, A. J. Phys. Chem. A 2002, 106, 4763-4771.) - Peptide stock solutions of known concentration (determined by UV absorbance at 388 nm, E=7980 M-1 cm-1) were used to prepare 5 μM peptide concentration samples in buffered water (150 μL total volume, 100 mM NaCl, 10 mM sodium phosphate, pH 7.5). 2 μL aliquots of the corresponding stock protein solution (GST-Crk SH2, GST-Abl SH2 or GST-PI3K SH2, concentration determined by Bio-Rad Protein Assay) were consecutively added, and fluorescence emission spectra were recorded 1 minute after each addition (excitation 395 nm, 2 nm excitation and emission slit width). The increase in emission intensity at the maximum emission wavelength was plotted against total protein concentration and fitted by least-square analysis assuming a 1 to 1 binding mode to obtain the apparent Kd.
- Human Crk SH2 domain was expressed as a GST fusion protein transformed into E. coli NB42 strain. Expression of SH2 domains was carried out following reported conditions. The culture was inoculated intro prewarmed LB media with carbenicilin 50 μg/mL. The sample was shaken at 37° C. for at least 2 h (passed semi-logarithmic growth A600=0.5)). IPTG was added to 0.1 mM; (4) the sample was shaken at 25° C. for four hours. Using lower temperature than previously reported conditions minimizes inclusion bodies formation. The sample was then centrifuged (5000 g for 30 min at 4° C.) and the pellet was resuspended in ice-cold lysis buffer (see below). The slurry was then sonicated until cells were disrupted (20 sec sonication and 30 seconds in ice for 4 times). Note: it was observed that treatment of the resuspended bacteria with lysozyme (1 mg/mL) for 30 min at 4° C. improved the lysis process and resulted in higher yields.
- Lysis buffer: PBS with 100 mM EDTA, 1% Triton X-100, 10% glycerol, 1 mM DTT, AEBSF, leupeptin and PepA. (add 100 μL AEBSF+25 μL Leupeptin+12.5 μL Pep A+to 10 mL total buffer).
- Purification: The sample was recentrifuged to remove particulate debris (5000 g for 30 min at 4° C.). Then the supernatant was incubated for at least 1 hour with 10 mL of
glutathione Sepharose 4 Fast Flow (Amersham Biosciences). The column was equilibrated with 5 volumes of binding buffer at 4° C. The sample was the loaded using a syringe and low flow rate (0.2 to 0.5 mL/min) at 4° C. The column was then washed with 5 volumes of binding buffer (higher rate of 1 mL/min) room temperature. The GST fusion protein was eluted with elution buffer at room temperature. The pooled fractions were then collected and analyzed using gel electrophoresis to confirm purity using 10% SDS-PAGE. Binding buffer: PBS pH 7.4 (140 mM NaCl, 2.7 mM KCl, 10 mM Na2HPO4, 1.8 mM NaHPO4). Elution buffer: 50 mM Tris-HCl, 10 mM reduced glutathione,pH 8. - The gel was transferred to a nitrocellulose membrane (1 h, 100 V), and probed using the standard procedure: blocked overnight with TBST+powder milk; washed with TBST (2×10 mL×10 min.); incubated with primary antibody (anti-GST, dilution 1:10,000 in TBST) for 30 min; washed with TBST (3×10 mL×10 min.); incubated for 30 min with secondary antibody (anti-goat conjugated with alkaline phosphatase, 1:1,000 dilution in TBS); washed with TBST (2×10 mL×10 min.); washed with TBS (1×10 mL×10 min.); One Step NBT/BCIP added and incubated until the bands appeared; then quenched the reaction with ddH2O. Once the protein was isolated, it was separated from the reduced glutathione using the Ultrafree-15 centrifugal filter (Biomax-5). The stock solution of Crk-SH2-GST was concentrated at 2000 g until the volume was approximately 2 mL. Phosphate buffer (10 mM phosphate, pH 7.5 100 mM NaCl) was added to a final volume of 15 mL and centrifuged again to a final volume of 2 mL. This procedure was repeated three times to ensure that the concentration of the contaminant solutes was minimal. The final volume was 1.2 mL.
- Bio-Rad Protein Assay was used to quantify the isolated following the standard procedure. Dye reagent was prepared by diluting 1 part Dye Reagent Concentrate with 4 parts distilled water. Stock BSA dilutions were prepared corresponding to 0.1, 0.2, 0.3, 0.4, 0.5 0.6, 0.7, 0.8 and 0.9 mg/mL as follows. 20 μL of each stock dilution were added to 980 μL dye reagent and incubated at room temperature for 5 min. The absorbance was measured at 595 nm using a blank of 20 μL of distilled water and 980 μL dye reagent prepared at the same time as the samples. The observed absorbance was plotted against BSA concentration.
- Three samples of increasing concentration of Crk-SH2-GST stock solution were prepared. Then, 10, 15 and 20 μL of stock solution with water up to 20 μL as necessary were added to 980 μL dye reagent. Absorbance measurements of these solutions in comparison with the standard curve afforded a final concentration of 11 mg/mL of Crk-SH2-GST.
- Abl-SH2 was received as a GST fusion in a pGEX-KT vector, amp-resistant, IPTG-inducible. The SH2 domain was cloned using BamH1 (5′) EcoR1 (3′) sites.
- The expression of GST-Abl SH2 GST-Src SH2 and GST-PI3K SH2 was carried out using the same protocol and conditions as the GST-Crk SH2 mutant, but in these cases it was found that expression at 37° C. resulted in increased yields of protein with no inclusion bodies formation.
- Src and PI3K SH2 domains were expressed and purified as GST fusion following reported procedures. (Smith, D. B.; Johnson, K. S. Gene, 1988, 67, 31-40) In short, plasmids including GST-fusion proteins were transformed into bacteria (E. coli DH5α chemically competent cells, Invitrogen). Bacteria were grown to mid log phase, induced at 37° C. for 3-4 hours with isopropyl-1-thio-β-D-galactopyranoside, and lysed by treatment with lysozyme (1 mg/mL, 30 minutes) followed by sonication (PBS buffer pH 7.4 with 100 mM EDTA, 1% Triton X-100, 10% glycerol, 1 mM DTT, 0.1 mM AEBSF, 30 μg/mL Leupeptin and 0.5 μg/mL Pepstatin A). The lysates were clarified by centrifugation, and fusion proteins were purified by binding to glutathione-agarose beads (Amersham Biosciences). Proteins were eluted from the beads (50 mM Tris-HCl, 10 mM reduced glutathione, pH 8), and finally concentrated by centrifugation through a cellulose membrane with a 10,000 Da cutoff (Millipore). Proteins were quantified using the Micro BCA protein Assay Kit (Pierce) relative to a BSA standard. All fusion proteins were analyzed by Coomasie staining and Western analysis with anti-GST antibodies.
Claims (7)
1-5. (canceled)
6. A compound of the formula (IV):
wherein R4, R5, R6, R7, R8, and R9 are each independently hydrogen, fluorine, or alkyl, with the proviso that one of R6 or R7 is —NR1R2, —OR1, or —SR1;
R1 and R2 are each independently hydrogen or alkyl; or
R1 and R2 together with the nitrogen to which they are attached, may form a substituted or unsubstituted 5- or 6-membered ring; wherein said nitrogen is not conjugated with said 5- or 6-membered ring;
Y is NH2 or C(O)X;
X is halogen, hydroxy or alkoxy; and
n is 1, 2, 3, or 4.
7. The compound of claim 6 , which is of the formula (V):
wherein R4, R5, R6, R7, R8, and R9 are each independently hydrogen, fluorine, or alkyl, with the proviso that one of R6 or R7 is —NR1R2, —OR1, or —SR1;
R1 and R2 are each independently hydrogen or alkyl; or
R1 and R2 together with the nitrogen to which they are attached, may form a substituted or unsubstituted 5- or 6-membered ring; wherein said nitrogen is not conjugated with said 5- or 6-membered ring.
8. The compound of claim 6 , which is of the formula (VI):
wherein R4, R5, R6, R7, R8, and R9 are each independently hydrogen, fluorine, or alkyl, with the proviso that one of R6 or R7 is —NR1R2, —OR1, or —SR1;
R1 and R2 are each independently hydrogen or alkyl; or
R1 and R2 together with the nitrogen to which they are attached, may form a substituted or unsubstituted 5- or 6-membered ring; wherein said nitrogen is not conjugated with said 5- or 6-membered ring.
9. The compound of claim 6 , which is of the formula (VII):
wherein R4, R5, R6, R7, R8, and R9 are each independently hydrogen, fluorine, or alkyl, with the proviso that one of R6 or R7 is —NR1, —OR1, or —SR1;
R1 and R2 are each independently hydrogen or alkyl; or
R1 and R2 together with the nitrogen to which they are attached, may form a substituted or unsubstituted 5- or 6-membered ring; wherein said nitrogen is not conjugated with said 5- or 6-membered ring.
11-20. (canceled)
Priority Applications (1)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| US12/235,454 US20090082577A1 (en) | 2005-04-13 | 2008-09-22 | Fluorescent probes for biological studies |
Applications Claiming Priority (2)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| US11/106,349 US7442529B2 (en) | 2005-04-13 | 2005-04-13 | Fluorescent probes for biological studies |
| US12/235,454 US20090082577A1 (en) | 2005-04-13 | 2008-09-22 | Fluorescent probes for biological studies |
Related Parent Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| US11/106,349 Division US7442529B2 (en) | 2005-04-13 | 2005-04-13 | Fluorescent probes for biological studies |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| US20090082577A1 true US20090082577A1 (en) | 2009-03-26 |
Family
ID=37108913
Family Applications (2)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| US11/106,349 Expired - Lifetime US7442529B2 (en) | 2005-04-13 | 2005-04-13 | Fluorescent probes for biological studies |
| US12/235,454 Abandoned US20090082577A1 (en) | 2005-04-13 | 2008-09-22 | Fluorescent probes for biological studies |
Family Applications Before (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| US11/106,349 Expired - Lifetime US7442529B2 (en) | 2005-04-13 | 2005-04-13 | Fluorescent probes for biological studies |
Country Status (1)
| Country | Link |
|---|---|
| US (2) | US7442529B2 (en) |
Cited By (4)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US20100168428A1 (en) * | 2007-02-26 | 2010-07-01 | Massachusetts Institute Of Technology | Environmentally sensitive fluorophores |
| US20110053180A1 (en) * | 2009-08-31 | 2011-03-03 | Massachusetts Institute Of Technology | Kinase sensors |
| US8586570B2 (en) | 2006-08-28 | 2013-11-19 | Massachusetts Institute Of Technology | Sox-based kinase sensor |
| US10799603B2 (en) | 2018-03-12 | 2020-10-13 | Massachusetts Institute Of Technology | Kinase and/or phosphatase sensing via hydroxyquinoline-sensitized chelates |
Families Citing this family (2)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US20100248975A1 (en) * | 2006-12-29 | 2010-09-30 | Gunjan Tiwari | Fluorogenic peptide substrate arrays for highly multiplexed, real-time monitoring of kinase activities |
| US20080206885A1 (en) * | 2007-02-26 | 2008-08-28 | Massachusetts Institute Of Technology | Environmentally sensitive fluorophores |
Family Cites Families (1)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US5854275A (en) * | 1996-05-16 | 1998-12-29 | Pfizer Inc. | Cyclic imide derivatives |
-
2005
- 2005-04-13 US US11/106,349 patent/US7442529B2/en not_active Expired - Lifetime
-
2008
- 2008-09-22 US US12/235,454 patent/US20090082577A1/en not_active Abandoned
Cited By (6)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US8586570B2 (en) | 2006-08-28 | 2013-11-19 | Massachusetts Institute Of Technology | Sox-based kinase sensor |
| US20100168428A1 (en) * | 2007-02-26 | 2010-07-01 | Massachusetts Institute Of Technology | Environmentally sensitive fluorophores |
| US8440835B2 (en) | 2007-02-26 | 2013-05-14 | Massachusetts Institute Of Technology | Environmentally sensitive fluorophores |
| US20110053180A1 (en) * | 2009-08-31 | 2011-03-03 | Massachusetts Institute Of Technology | Kinase sensors |
| US8409820B2 (en) | 2009-08-31 | 2013-04-02 | Massachusetts Institute Of Technology | Kinase sensors |
| US10799603B2 (en) | 2018-03-12 | 2020-10-13 | Massachusetts Institute Of Technology | Kinase and/or phosphatase sensing via hydroxyquinoline-sensitized chelates |
Also Published As
| Publication number | Publication date |
|---|---|
| US20060234206A1 (en) | 2006-10-19 |
| US7442529B2 (en) | 2008-10-28 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| Zheng et al. | Rational design of fluorogenic and spontaneously blinking labels for super-resolution imaging | |
| JP7159233B2 (en) | Quinone Masked Probes as Labeling Reagents for Measuring Uptake by Cells | |
| ES2211650T3 (en) | IMIDAZOQUINOLINE COMPOUNDS MARKED WITH COLOR. | |
| US9006437B2 (en) | Xanthene dyes comprising a sulfonamide group | |
| US20120115128A1 (en) | Selective protein labeling | |
| US20090082577A1 (en) | Fluorescent probes for biological studies | |
| US20080206885A1 (en) | Environmentally sensitive fluorophores | |
| CA3091814A1 (en) | Dyes for analysis of soluble protein aggregates or misfolded protein oligomers | |
| JP6862368B2 (en) | Use as an inhibitor of thienopyrol compounds and their Oplophorus-derived luciferase | |
| US8440835B2 (en) | Environmentally sensitive fluorophores | |
| US8409820B2 (en) | Kinase sensors | |
| Zhang et al. | Synthesis and application of the blue fluorescent amino acid l-4-cyanotryptophan to assess peptide–membrane interactions | |
| US9701667B2 (en) | Coumarin-based fluorogenic agents and uses thereof for specific protein labelling | |
| CN107098923A (en) | One class feux rouges targets fluorescent dye and preparation method thereof and purposes near infrared emission lysosome | |
| JP2604993B2 (en) | New biotinylation reagent | |
| Wurnig et al. | Development of the first geldanamycin-based HSP90 degraders | |
| Takimoto et al. | Esterification of an unnatural amino acid structurally deviating from canonical amino acids promotes its uptake and incorporation into proteins in mammalian cells | |
| KR20050049513A (en) | Protein labelling with o6-alkylguanine-dna alkyltransferase | |
| Clavé et al. | A universal and ready-to-use heterotrifunctional cross-linking reagent for facile synthetic access to sophisticated bioconjugates | |
| US20230257587A1 (en) | Fluorenyl cyanine dyes | |
| Amina et al. | Biophysical characterization of the interaction of bovine serum albumin with anticancer sipholane triterpenoid from the Red Sea sponge | |
| JPWO2015125892A1 (en) | Protein labeling compounds | |
| Feng et al. | Several Golgi targeting fluorescence markers based on 1, 8-naphthalimide derivatives with amide and long carbon chain | |
| WO2017154997A1 (en) | Indole structure-selective crosslinking agent and composite in which same is used | |
| US8715529B1 (en) | Synthesis and applications of triazaborolopyridinium compounds and substituted triazaborolopyridinium compounds and methods of use |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| STCB | Information on status: application discontinuation |
Free format text: ABANDONED -- FAILURE TO RESPOND TO AN OFFICE ACTION |