CN116244602A - Sample pollution detection and model training method, device, equipment and medium - Google Patents
Sample pollution detection and model training method, device, equipment and medium Download PDFInfo
- Publication number
- CN116244602A CN116244602A CN202310273973.1A CN202310273973A CN116244602A CN 116244602 A CN116244602 A CN 116244602A CN 202310273973 A CN202310273973 A CN 202310273973A CN 116244602 A CN116244602 A CN 116244602A
- Authority
- CN
- China
- Prior art keywords
- sample
- sample data
- data
- detected
- target site
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Pending
Links
Images
Classifications
-
- G—PHYSICS
- G06—COMPUTING OR CALCULATING; COUNTING
- G06N—COMPUTING ARRANGEMENTS BASED ON SPECIFIC COMPUTATIONAL MODELS
- G06N20/00—Machine learning
Landscapes
- Engineering & Computer Science (AREA)
- Theoretical Computer Science (AREA)
- Software Systems (AREA)
- Data Mining & Analysis (AREA)
- Evolutionary Computation (AREA)
- Medical Informatics (AREA)
- Computer Vision & Pattern Recognition (AREA)
- Physics & Mathematics (AREA)
- Computing Systems (AREA)
- General Engineering & Computer Science (AREA)
- General Physics & Mathematics (AREA)
- Mathematical Physics (AREA)
- Artificial Intelligence (AREA)
- Measuring Or Testing Involving Enzymes Or Micro-Organisms (AREA)
Abstract
The invention discloses a sample pollution detection and model training method, device, equipment and medium. The sample pollution detection method comprises the following steps: acquiring allelic variation abundance of a target site of sample data to be detected; inputting the allelic variation abundance of the target site of the sample data to be detected into a pre-trained sample pollution detection model to obtain a sample detection result and sample pollution probability of the sample to be detected. The technical scheme of the embodiment of the invention improves the efficiency and accuracy of sample pollution detection.
Description
Technical Field
The invention relates to the field of biotechnology, in particular to a sample pollution detection and model training method, device, equipment and medium.
Background
Sample contamination is an important factor affecting laboratory data analysis results. Sample contamination can be generally classified into reagent contamination, human contamination, environmental contamination, and the like.
At present, whether the sample is polluted or not is judged by manually checking a sample analysis mass spectrogram.
However, the detection mode of manual identification is time-consuming and laborious, and meanwhile, due to the difference of identification standards among different manual works, the problems of low sample pollution detection efficiency and poor accuracy exist.
Disclosure of Invention
The invention provides a sample pollution detection and model training method, device, equipment and medium, which improve the efficiency and accuracy of sample pollution detection.
According to an aspect of the present invention, there is provided a sample contamination detection method comprising:
acquiring allelic variation abundance of a target site of sample data to be detected;
inputting the allelic variation abundance of the target site of the sample data to be detected into a pre-trained sample pollution detection model to obtain a sample detection result and sample pollution probability of the sample to be detected.
According to another aspect of the present invention, there is provided a model training method including:
obtaining allelic variation abundance of a target site of reference sample data;
inputting the allelic variation abundance of the target site of the reference sample data into a sample pollution detection model to obtain a reference detection result and a reference pollution probability of a sample to be detected;
obtaining a standard detection result of reference sample data;
training a sample pollution detection model according to the difference between the reference detection result and the standard detection result of the reference sample data;
the sample pollution detection model can realize the sample pollution detection method of any embodiment of the invention.
According to an aspect of the present invention, there is provided a sample contamination detection apparatus comprising:
the sample data to be detected acquisition module is used for acquiring the allelic variation abundance of the target site of the sample data to be detected;
the sample detection result determining module is used for inputting the allelic variation abundance of the target site of the sample data to be detected into a pre-trained sample pollution detection model to obtain a sample detection result and a sample pollution probability of the sample to be detected.
According to another aspect of the present invention, there is provided a model training apparatus comprising:
the sample data to be detected acquisition module is used for acquiring the allelic variation abundance of the target site of the sample data to be detected;
the sample detection result determining module is used for inputting the allelic variation abundance of the target site of the sample data to be detected into a pre-trained sample pollution detection model to obtain a sample detection result and a sample pollution probability of the sample to be detected.
According to another aspect of the present invention, there is provided an electronic apparatus including:
at least one processor; and
a memory communicatively coupled to the at least one processor; wherein,,
The memory stores a computer program executable by the at least one processor to enable the at least one processor to perform the sample contamination detection method or the model training method of any one of the embodiments of the present invention.
According to another aspect of the present invention, there is provided a computer readable storage medium storing computer instructions for causing a processor to implement the sample contamination detection method or the model training method of any of the embodiments of the present invention when executed.
According to the technical scheme, the allelic variation abundance of the target site of the sample data to be detected is obtained; the allele variation abundance of the target site of the sample data to be detected is input into a pre-trained sample pollution detection model, so that a sample detection result and sample pollution probability of the sample to be detected are obtained, the problems of time and labor waste in a manual identification detection mode, and meanwhile, due to the difference of identification standards among different manual works, the problems of low sample pollution detection efficiency and poor sample pollution detection accuracy exist, and the efficiency and the sample pollution detection accuracy are improved.
It should be understood that the description in this section is not intended to identify key or critical features of the embodiments of the invention or to delineate the scope of the invention. Other features of the present invention will become apparent from the description that follows.
Drawings
In order to more clearly illustrate the technical solutions of the embodiments of the present invention, the drawings required for the description of the embodiments will be briefly described below, and it is apparent that the drawings in the following description are only some embodiments of the present invention, and other drawings may be obtained according to these drawings without inventive effort for a person skilled in the art.
FIG. 1 is a flow chart of a sample contamination detection method according to a first embodiment of the present invention;
FIG. 2 is a flow chart of a model training method according to a second embodiment of the present invention;
FIG. 3 is a distribution of reference sample data 01-AA allele variation abundance according to a second embodiment of the invention;
FIG. 4 is a distribution of abundance of variation of the 02-BB allele in reference sample data provided in accordance with a second embodiment of the present invention;
FIG. 5 is a flow chart of a model training method according to a second embodiment of the present invention;
fig. 6 is a schematic structural diagram of a sample contamination detection apparatus according to a third embodiment of the present invention;
FIG. 7 is a schematic structural diagram of a model training apparatus according to a third embodiment of the present invention;
fig. 8 is a schematic structural diagram of an electronic device implementing a sample contamination detection method or a model training method according to an embodiment of the present invention.
Detailed Description
In order that those skilled in the art will better understand the present invention, a technical solution in the embodiments of the present invention will be clearly and completely described below with reference to the accompanying drawings in which it is apparent that the described embodiments are only some embodiments of the present invention, not all embodiments. All other embodiments, which can be made by those skilled in the art based on the embodiments of the present invention without making any inventive effort, shall fall within the scope of the present invention.
It should be noted that the terms "first," "second," and the like in the description and the claims of the present invention and the above figures are used for distinguishing between similar objects and not necessarily for describing a particular sequential or chronological order. It is to be understood that the data so used may be interchanged where appropriate such that the embodiments of the invention described herein may be implemented in sequences other than those illustrated or otherwise described herein. Furthermore, the terms "comprises," "comprising," and "having," and any variations thereof, are intended to cover a non-exclusive inclusion, such that a process, method, system, article, or apparatus that comprises a list of steps or elements is not necessarily limited to those steps or elements expressly listed but may include other steps or elements not expressly listed or inherent to such process, method, article, or apparatus.
Example 1
Fig. 1 is a flowchart of a sample contamination detection method according to an embodiment of the present invention. The embodiment of the invention is applicable to the detection of sample pollution, the method can be executed by a sample pollution detection device, the sample pollution detection device can be realized in the form of hardware and/or software, and the sample pollution detection device can be configured in an electronic device carrying a sample pollution detection function.
Referring to the sample contamination detection method shown in fig. 1, the method includes:
s110, acquiring the allelic variation abundance of the target site of the sample data to be detected.
Sample contamination is an important factor affecting laboratory data analysis results. Sample contamination can be categorized from sources into reagent contamination, human contamination, environmental contamination, and the like. Sample data to be tested may be acquired and analyzed in order to determine if the sample is contaminated. Further, it can be determined whether the sample to be detected is contaminated by analyzing mutation data in the sample data to be detected. The sample data to be detected may be genetic sequencing sample data. Alternatively, the mutation data in the sample data to be detected may include the site where the mutation occurs, the type of mutation, the ratio of mutation, the abundance of allelic variation, and the like. The target site may be a site where sample contamination detection is performed. The target site may be a part of the typical site where the mutation occurs. Optionally, a preset number of target sites can be selected for the sites with mutation, and the target sites can be screened according to the allelic variation abundance of the sites with mutation. Alleles are used to describe genes that control different morphologies of the same trait at the same position on a pair of homologous chromosomes. Allelic variation abundance, also known as allele frequency (Variant Allele Frequency, VAF), is used to indicate the diversity of genes, or the abundance of gene pools, in a population. The allelic variation abundance may be the ratio of the number of mutated alleles to the number of total alleles.
Optionally, the sample data to be detected may be analyzed to determine a target site of the sample data to be detected, and further determine the allelic variation abundance of the target site of the sample data to be detected. Alternatively, the allelic variation abundance of each site of the sample data to be detected may be calculated, and then the target site is determined in each site, thereby determining the allelic variation abundance of the target site of the sample data to be detected.
S120, inputting the allelic variation abundance of the target site of the sample data to be detected into a pre-trained sample pollution detection model, and obtaining a sample detection result and a sample pollution probability of the sample to be detected.
The sample test results may include "contaminated" and "uncontaminated". The sample contamination probability may be a probability that the sample to be detected is contaminated. Alternatively, the sample contamination probability may be determined based on a confidence of a sample detection result of the sample contamination detection model. For example, if the sample contamination detection result is "uncontaminated", the sample contamination probability may be 0%; if the sample pollution detection result is pollution, the confidence level of the pollution result in the sample pollution detection model can be determined as the sample pollution probability.
Specifically, the allelic variation abundance of the target site of the sample data to be detected can be input into a pre-trained sample pollution detection model, and the sample pollution detection model outputs a sample detection result representing whether the sample to be detected is polluted or not and a sample pollution probability.
According to the technical scheme, the allele variation abundance of the target site of the sample data to be detected is obtained and is input into the sample pollution detection model trained in advance, so that the sample detection result and the sample pollution probability of the sample to be detected are obtained, the problems of time and labor waste in manually identifying sample pollution are avoided, high-flux detection of sample pollution is realized, the sample pollution detection efficiency is improved, meanwhile, the difference of sample pollution detection standards among different workers is avoided, and the sample pollution detection accuracy is improved.
In an alternative embodiment of the present invention, the allelic variation abundance of the target site for obtaining the sample data to be detected is specified as follows: carrying out gene sequencing on sample data to be detected to obtain crowd frequency and genome components of alternative sites of the sample data to be detected; screening target sites of crowd frequency in a preset crowd frequency range from a plurality of candidate sites of sample data to be detected; and calculating the allelic variation abundance of the target site according to the gene components of the target site.
The candidate sites may be all sites in the sample data to be detected. Optionally, the number of candidate sites is greater than the target site. Illustratively, the number of candidate sites may be 1000 and the number of target sites may be 61. The population frequency may be the frequency with which a locus occurs in the genome of a population. The predetermined crowd frequency range may be a predetermined crowd frequency range. The frequency range of the preset crowd can be set and adjusted according to the experience of technicians. Illustratively, the predetermined crowd frequency range may be 0.4-0.6. A genome component may be a constituent of an allele at a locus. Illustratively, the allele at a locus may be AABB and the constituent components of the allele may be a and B.
Specifically, the gene sequencing tool can be used for carrying out gene sequencing on the sample data to be detected, so that the crowd frequency and the gene components of the candidate sites of the sample data to be detected are obtained. Among other things, gene sequencing tools may include NGS (Next Generation Sequencing, second generation/high throughput sequencing) techniques, sanger (Sanger) sequencing, long read long sequencing, and the like. Among a plurality of candidate sites of the sample data to be detected, candidate sites with crowd frequency within a preset crowd frequency range can be screened and used as target sites. The ratio of the genome component of the mutated allele to the total genome component of the allele can be calculated from the genome components of the target site. Illustratively, if the genome of the mutated allele is a and the genome of the other allele is B, then the allelic variation abundance=a/(a+b). If an allele at a certain site is expressed as AAAA, the allelic variation abundance of the site is 100%; if an allele at a certain site is expressed as AAAB, the allelic variation abundance at that site is 25%; if an allele at a certain site is expressed as ABAB, the allelic variation abundance at that site is 50%; if the allele at a certain site is expressed as ABBB, the allelic variation abundance of the site is 75%; if an allele at a certain site is expressed as BBBB, the abundance of the allelic variation at that site is 100%.
According to the method, the crowd frequency and the genome components of the candidate sites of the sample data to be detected are obtained by carrying out gene sequencing on the sample data to be detected, the target sites of the crowd frequency in a preset crowd frequency range are screened from the candidate sites of the sample data to be detected, the allele variation abundance of the target sites is obtained through calculation according to the genome components of the target sites, the high heterozygosity of the target sites is ensured through crowd frequency, the problem that the allele variation abundance is calculated for the candidate sites first, then the allele variation abundance of the target sites is obtained, the calculated amount is large is solved, the calculated amount of the allele variation abundance is reduced, the calculation efficiency of the allele variation abundance is improved, and the detection efficiency of sample pollution is further improved.
In an alternative embodiment of the present invention, after obtaining the allelic variation abundance of the target site of the sample data to be detected, further comprising: carrying out sample pollution correlation analysis on the allelic variation abundance of each target site; and removing target sites with sample pollution correlation analysis results lower than a preset correlation threshold value from each target site.
The target sites of the sample data to be detected are more, and the allelic variation abundance of part of the target sites has weaker correlation with whether the sample is polluted or not. Therefore, the correlation analysis can be carried out on each target site of the sample data to be detected, and the target sites with weak correlation with sample pollution can be removed.
Specifically, sample pollution correlation analysis can be performed on the allelic variation abundance of each target site by adopting a characteristic engineering method, the target site with obvious influence on pollution is screened, and if the sample pollution correlation analysis result of the allelic variation abundance of the target site is lower than a preset correlation threshold, it can be understood that the target site has weaker correlation with sample pollution, and then the target site corresponding to the position is eliminated. Among them, feature engineering methods include Filter, wrapper, embedded, and the like. The preset correlation threshold may be a minimum value of a correlation analysis result set in advance. The preset correlation threshold can be set and adjusted according to the experience of the skilled person and the characteristic engineering method. For example, the preset correlation threshold may be 0.03.
According to the scheme, after the allelic variation abundance of the target sites of the sample data to be detected is obtained, sample pollution correlation analysis is carried out on the allelic variation abundance of each target site; and removing target sites with sample pollution correlation analysis results lower than a preset correlation threshold value from the target sites, removing target sites with weak pollution correlation with the sample from the target sites, realizing characteristic engineering of the target sites, further reducing the quantity of input data of a sample pollution detection model, improving the operation efficiency of the sample pollution detection model, and further improving the detection efficiency of sample pollution.
Example two
Fig. 2 is a flowchart of a model training method according to a second embodiment of the present invention. The embodiment of the invention is applicable to the training of the sample pollution detection model, the method can be executed by a model training device, the model training device can be realized in a form of hardware and/or software, and the model training device can be configured in electronic equipment for carrying model training.
Referring to the model training method shown in fig. 2, it includes:
s210, acquiring allelic variation abundance of a target site of reference sample data.
The reference sample data may be sample data employed for model training. The reference sample data may also be genetic sequencing sample data. The reference sample data may be acquired and analyzed in order to determine whether the sample is contaminated. Further, it may be determined whether the sample to be detected is contaminated by analyzing mutation data in the reference sample data. Alternatively, the mutation data in the reference sample data may include the site where the mutation occurred, the type of mutation, the ratio of mutation, the abundance of allelic variation, etc. The target site may be a site where sample contamination detection is performed. The target site may be a part of the typical site where the mutation occurs. Optionally, a preset number of target sites can be selected for the sites with mutation, and the target sites can be screened according to the allelic variation abundance of the sites with mutation. Alleles are used to describe genes that control different morphologies of the same trait at the same position on a pair of homologous chromosomes. The allelic variation abundance may be the ratio of the number of mutated alleles to the number of total alleles.
Optionally, the reference sample data may be analyzed to determine a target site of the reference sample data, and thus the allelic variation abundance of the target site of the reference sample data.
Alternatively, the allelic variation abundance of each site of the reference sample data may be calculated, and then the target site may be determined in each site, thereby determining the allelic variation abundance of the target site of the reference sample data.
Alternatively, the reference sample data may be subjected to genetic sequencing using a genetic sequencing tool to obtain the population frequency and genetic composition of the candidate sites of the reference sample data. Among a plurality of candidate sites of the reference sample data, candidate sites of the crowd frequency within a preset crowd frequency range can be screened as target sites. The ratio of the genome component of the mutated allele to the total genome component of the allele can be calculated from the genome components of the target site.
S220, inputting the allelic variation abundance of the target site of the reference sample data into a sample pollution detection model to obtain a reference detection result and a reference pollution probability of the sample to be detected.
The sample pollution detection model of the step is an untrained sample pollution detection model. Alternatively, the sample contamination detection model may be constructed by logistic regression (Logistic Regression), decision Tree (Decision Tree), support vector machine (Support Vector Machines, SVM), XGB (eXtreme Gradient Boosting, extreme gradient lifting algorithm), GMM (Gaussian Mixture Model ), random Forest (Random Forest), naive Bayes, light GBM (Light Gradient Boosting Machine, regression model classification model), and the like. The reference test results may include "contaminated" and "uncontaminated". The reference contamination probability may be a probability that the reference sample is contaminated. Alternatively, the sample contamination detection probability may be determined according to the confidence of the reference detection result of the sample contamination detection model. The sample contamination detection probability may be displayed as a percentage value.
Specifically, the allelic variation abundance of the target site of the reference sample data may be input into an untrained sample contamination detection model, which outputs a reference detection result that characterizes whether the reference sample is contaminated and a reference contamination probability.
Optionally, after obtaining the allelic variation abundance of the target site of the reference sample data, further comprising: carrying out sample pollution correlation analysis on the allelic variation abundance of each target site; and removing target sites with sample pollution correlation analysis results lower than a preset correlation threshold value from each target site.
The allele variation abundance of each target site is used for further processing each target site, so that target sites which have weak pollution correlation with the sample in each target site are eliminated, the characteristic engineering of each target site is realized, the quantity of input data of a sample pollution detection model is further reduced, the operation efficiency of the sample pollution detection model is improved, and the detection efficiency of sample pollution is further improved.
S230, standard detection results of the reference sample data are obtained.
The standard test result may be a known test result of the reference sample data.
Specifically, standard detection results known from reference sample data may be obtained.
S240, training the sample pollution detection model according to the difference between the reference detection result and the standard detection result of the reference sample data.
Specifically, the reference detection result output by the sample pollution detection model is compared with the standard detection result of the known reference sample data, and if the probability of the comparison result being consistent is greater than or equal to a preset accuracy threshold, the sample pollution detection model is trained; if the probability of the consistent comparison result is smaller than the preset accuracy threshold, the parameter adjusting method is adopted to adjust parameters of the sample pollution detection model, and training of the sample pollution detection model is continued until the probability of the consistent comparison result is larger than or equal to the preset accuracy threshold.
The parameter tuning method can comprise a grid searching method, a random searching method, a traditional manual parameter tuning method, a Bayesian searching method and the like. The preset accuracy threshold may be a minimum value of a probability that the preset comparison results agree. The preset accuracy threshold may be set and adjusted according to the experience of the technician.
According to the technical scheme, the allele variation abundance of the target site of the reference sample data is obtained, the allele variation abundance of the target site of the reference sample data is input into the sample pollution detection model, the reference detection result and the reference pollution probability of the sample to be detected are obtained, the standard detection result of the reference sample data is obtained, the sample pollution detection model is trained according to the difference between the reference detection result and the standard detection result of the reference sample data, the sample pollution detection model is trained, the problem that sample pollution is time-consuming and labor-consuming in manual identification is avoided, high-throughput detection of sample pollution is realized, the sample pollution detection efficiency is improved, meanwhile, the difference of sample pollution detection standards among different workers is avoided, and the sample pollution detection accuracy is improved.
In an alternative embodiment of the present invention, the standard detection result of acquiring the reference sample data is embodied as: and obtaining standard detection results of the reference sample data marked in advance in the experiment.
The reference sample data may be experimentally pre-generated sample data. At the same time as the reference sample data is generated, whether the reference sample data is actually contaminated, i.e., a standard detection result of the reference sample data, may be marked in advance.
Specifically, different leukocyte data can be directly used as reference sample data, and standard detection results of the reference sample data are marked as pollution-free in advance; different white blood cell data or tissue data can be mixed in different proportions through experiments, and standard detection results of reference sample data are marked as pollution in advance. Standard detection results of experimentally pre-labeled reference sample data can be directly obtained. White blood cells have the advantage of being relatively genome stable relative to other cellular data.
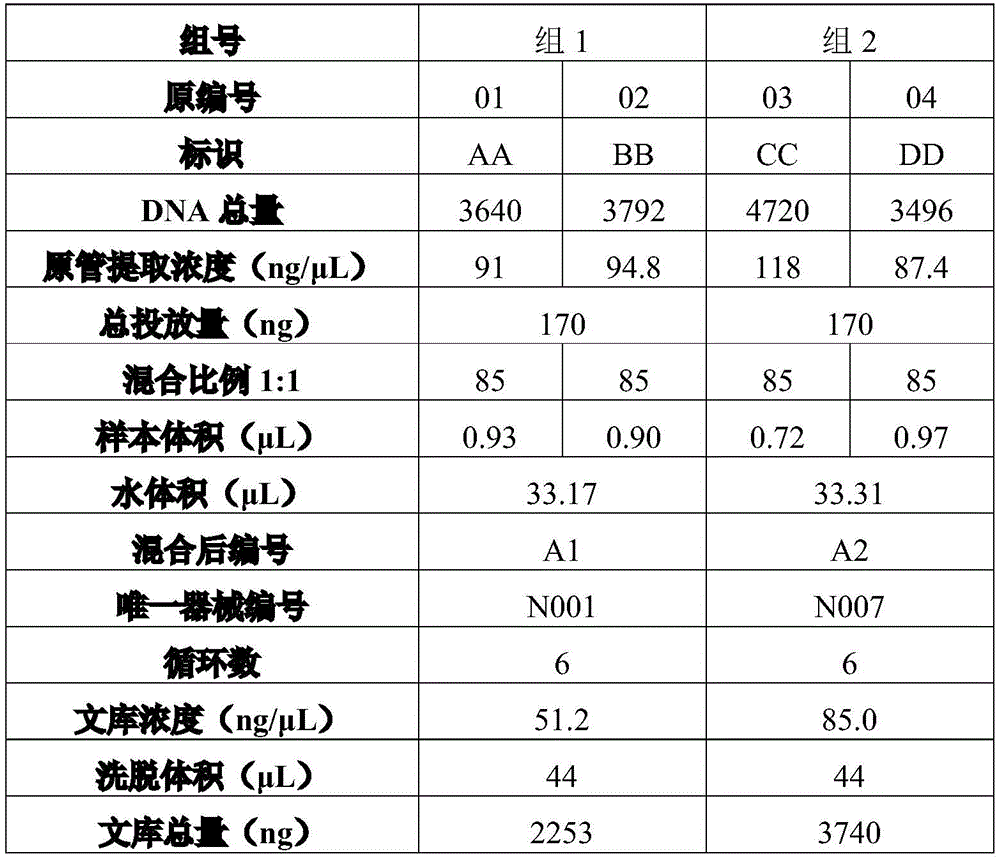
For example, table 1 is reference sample data for a blend ratio of 1:1.
As shown in table 1, two sets of reference sample data are shown. Wherein the reference sample data in group 1 identifies the group of AA by number 01 The organization data of the organization data and the identification BB of the number 02 are mixed according to the mixing proportion of 1:1 to generate. The reference sample data in the group 2 is generated by mixing the organization data of the identification CC of the number 03 and the organization data of the identification DD of the number 04 according to the mixing ratio of 1:1. The total amount of DNA (deoxynucleotides), the original tube extraction concentration, the mixing ratio, and the sample volume of the tissue data of AA (or the tissue data of BB) are identified by number 01, see table 1. The water volume, post-mix number, unique instrument number, number of cycles, library concentration, elution volume, and library total of the reference sample data for group 1 are shown in table 1. The total amount of DNA may be the total amount of DNA extracted by an instrument by making the surgically excised tissue into wax blocks. The original tube extraction concentration may be the concentration of DNA extracted by the instrument. The total dose may be the total dose of DNA for sample contamination detection. The number of cycles may be a power of 2 times the number of exponentiations of the disrupted DNA fragment. For example, if the number of cycles is 6, 2 are amplified for the broken DNA fragment 6 . The library concentration may be the concentration of DNA after amplification of the disrupted DNA fragment. The elution volume may be the volume of the library after eluting other solutions or enzymes or the like contained in the library than the DNA. The total amount of library may be the total amount of DNA after amplification of the disrupted DNA fragment. Group 2 is similar to group 1 and will not be described in detail herein.
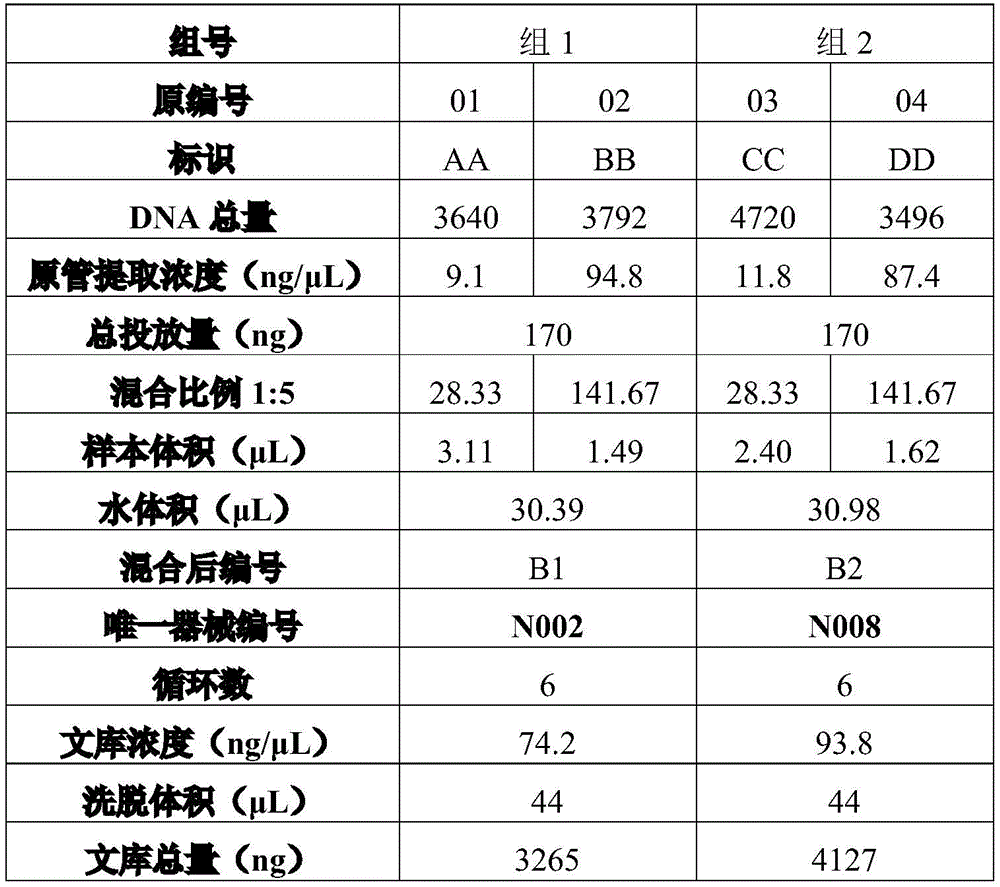
Exemplary, table 2 is reference sample data for a blend ratio of 1:5.
Exemplary, table 3 is reference sample data for a blend ratio of 1:10.
Exemplary, table 4 is reference sample data for a blend ratio of 1:99.
Exemplary, table 5 is reference sample data for a blend ratio of 3:97.
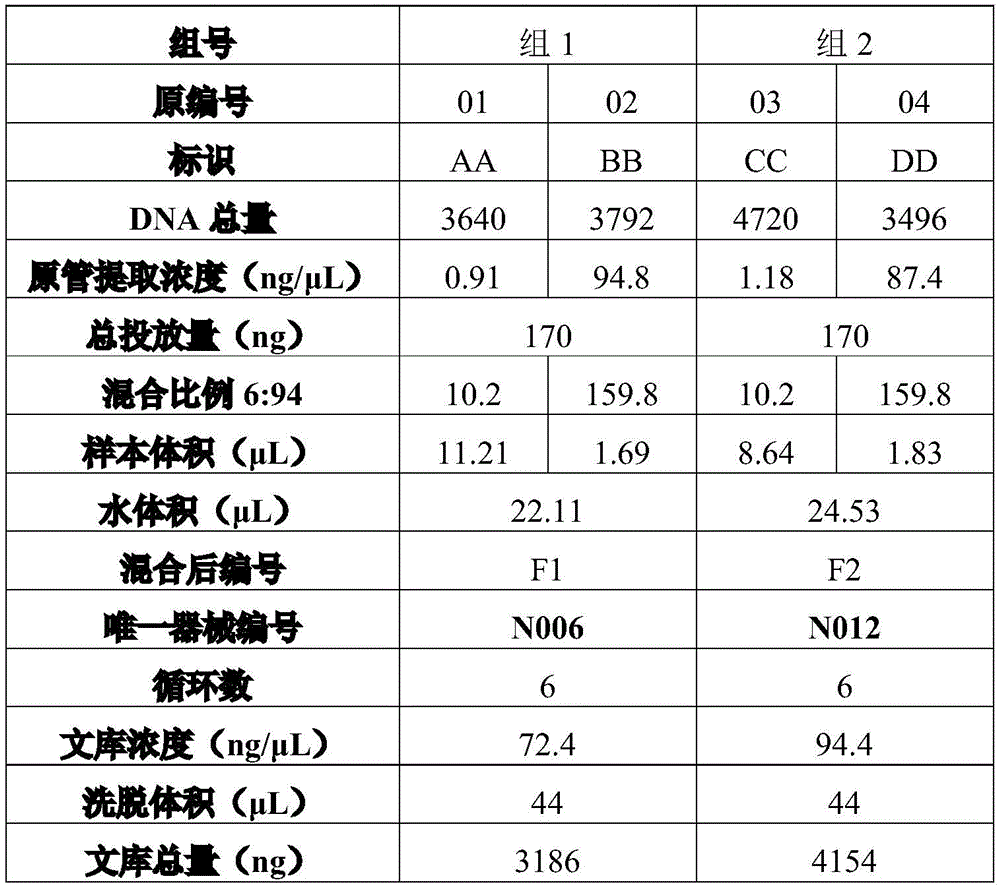
For example, table 6 is reference sample data for a blend ratio of 6:94.
The contents of tables 2 to 6 are similar to those of table 1, and are not repeated here.
Alternatively, after the genes are performed on each reference sample data, the number of gene sequencing data may also be subjected to standardized detection. Specifically, the number of the gene sequencing data generated after the gene sequencing of the single reference sample data can be compared with the amount of the data to be generated, and when the number of the gene sequencing data generated after the gene sequencing of the single reference sample data is greater than or equal to the amount of the data to be generated, the sample pollution detection can be performed. The amount of data to be produced may be a minimum value of the amount of gene sequencing data produced after gene sequencing of a single reference sample data set in advance. For example, if the sample type of the reference sample data is tissue data, the amount of the expected data may be 0.6G. Specifically, the sum of the amounts of the genetic test data generated after the genetic test of each reference sample data for single inspection can be compared with the amount of the data to be generated at this time, and when the sum of the amounts of the genetic test data generated after the genetic test of each reference sample data for single inspection is greater than or equal to the amount of the data to be generated at this time, the sample pollution detection can be performed. The data amount to be produced at this time may be the minimum value of the sum of the numbers of the genetic test data produced after the genetic test of each reference sample data of the preset single inspection. Alternatively, gene sequencing may be performed using a small panel (probe that simultaneously detects multiple sites of multiple genes) probe.
According to the scheme, the standard detection result of the reference sample data marked in advance in the experiment is obtained, so that the efficiency and accuracy of sample pollution detection model training are improved.
In an alternative embodiment of the present invention, the standard detection result of acquiring the reference sample data is embodied as: analyzing the distribution of the allelic variation abundance of the target site of the reference sample data, and determining a pollution mark of the reference sample data, wherein the pollution mark is used for distinguishing whether the reference sample data is polluted or not; and determining the pollution mark of the reference sample data as a standard detection result.
The contamination flag is used to distinguish whether the reference sample data is contaminated. For example, if the reference sample data is contaminated, the contamination label may be "contaminated"; if the reference sample data is not contaminated, the contamination marker may be "no contamination". By way of example, a "1" may be used to indicate contamination and a "0" may be used to indicate no contamination.
Specifically, the distribution of the allelic variation abundance of the target sites of the reference sample data can be analyzed, and if the distribution of the allelic variation abundance of each target site of the reference sample data is within the range of at most three target abundances, the pollution mark of the reference sample data is determined to be pollution-free; the contamination flag of the remaining reference sample data is determined to be contaminated. And determining the pollution mark of the reference sample data as a standard detection result. Wherein the target abundance may include 0%, 50% and 100%. The target abundance range may be a range equal to or greater than a difference between the target abundance and a preset abundance threshold, and equal to or less than a sum of the target wind power and the preset threshold. Illustratively, the preset abundance threshold may be 5%.
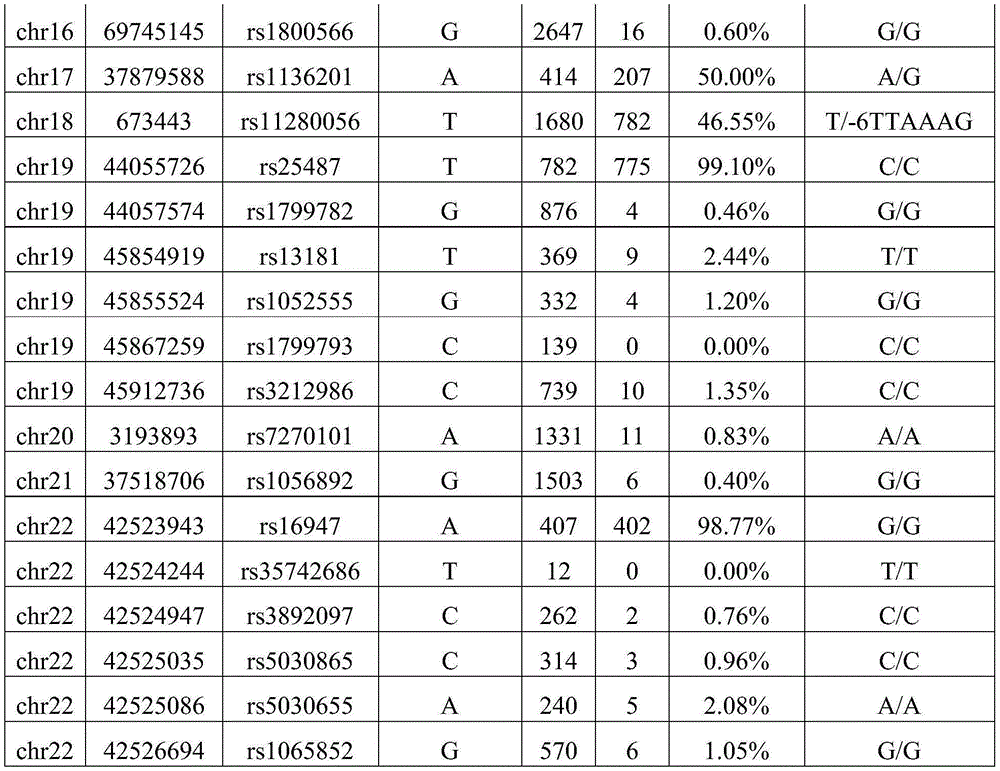
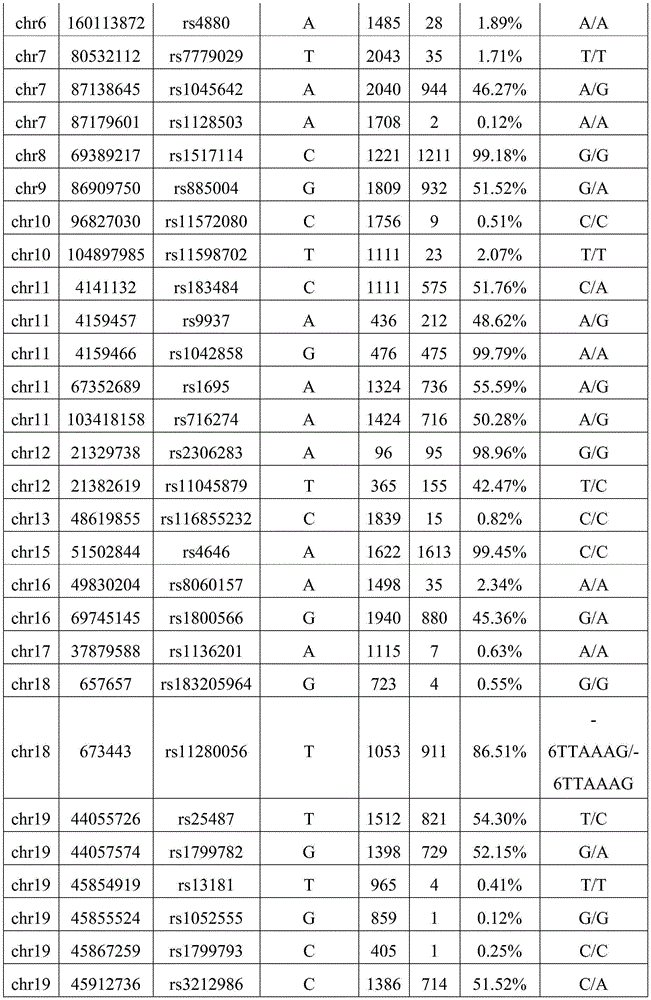
Illustratively, table 7 is a table of mutation data for the target site of reference sample data 01-AA.
Table 7 is reference sample data for a standard test result of "no contamination". The chromosome number may be the number of the chromosome where the target site is located. The chromosomal location may be the location number of the chromosome where the target site is located. The chromosome location identity may be identity information of the location of the chromosome where the target site is located. The reference base may be the base type of the human reference genome. The sequencing depth may be the number of repeated sequencing of the target site. The mutation number may be the number of data for detecting mutation at the target site. The genotype may be a comparison of the reference base of the human reference genome to the actual assay determined by the gene sequencing assay. By way of example, the genotype may be G/A. Wherein the reference base is G; the actual base is A. The results of the detection of the abundance of allelic variation are shown in Table 7. FIG. 3 is a graph showing the distribution of the abundance of 01-AA allelic variation in reference sample data provided according to the second embodiment of the invention. As shown in fig. 3, the allelic variation abundance distribution of the reference sample data is within three target abundance ranges of 0%, 50% and 100%.
Illustratively, table 8 is a table of mutation data for the target sites of reference sample data 02-BB.
Table 8 also shows reference sample data for "no contamination" for standard test results. The details are similar to those of table 7, and will not be described here again. FIG. 4 is a distribution of the abundance of variation of the 02-BB allele in reference sample data provided in accordance with a second embodiment of the present invention. As shown in fig. 4, the allelic variation abundance distribution of the reference sample data is within three target abundance ranges of 0%, 50% and 100%.
According to the method, the distribution of the allelic variation abundance of the target site of the reference sample data is analyzed, the reference sample data is determined to carry out pollution marking, the pollution marking of the reference sample data is determined to be a standard detection result, the pollution marking is determined in a targeted manner, the efficiency and the accuracy of pollution marking determination are further guaranteed, the pollution marking of the reference sample is determined to be a standard detection result, the accuracy of the output result of the sample pollution detection model is verified, and the training efficiency and the accuracy of the sample pollution detection model are further improved.
Fig. 5 is a flowchart of a model training method according to a second embodiment of the present invention. Referring to the model training method shown in fig. 5, it includes:
S510, carrying out gene sequencing on the reference sample data to obtain gene sequencing data.
The genetic sequencing data may include Fastp (filtered second generation sequencing) data.
S520, performing data quality control on the gene sequencing data to obtain data quality control data.
The data quality control data may be data obtained after the data quality control of the gene sequencing data. The data quality control is used for controlling the data quality of the gene sequencing data. Optionally, the data quality control may include deduplication correction, alignment correction, and the like. Wherein, the deduplication correction may be to discard duplicate data in the gene sequencing data. The comparison correction can be to compare the gene sequencing data with human reference genome data, and if the ratio of the number of the data which can be compared to the number of all the data is greater than or equal to a preset comparison ratio, the comparison correction of the data quality of the gene sequencing data is passed; if the ratio of the number of data which can be compared to the number of all data is smaller than the preset comparison example, the comparison correction of the data quality of the gene sequencing data is not passed, and the gene sequencing data is removed. Alternatively, the data quality control tools may include a Fastp (filtered second generation sequencing) tool, a Fastqc (data quality control) tool, a UMI (Unique Molecular Identifiers, unique molecular tag technology) tool, a Trimmomatic (high throughput data deduplication) tool, and the like.
Specifically, a data quality control tool can be used for carrying out data quality control on the gene sequencing data, so that data quality control data after the data quality control is obtained.
S530, data comparison is carried out on the data quality control data and the reference genome, and mutation data with mutation in the data quality control data is determined.
Specifically, a data comparison tool may be used to compare the data quality control data with the reference genome (i.e., the human reference genome) to determine mutation data in the data quality control data. Among them, the data Alignment tools include BWA (Burows-Wheeler-Alignment Tool), bowtie2 (sequencing sequence and reference sequence Alignment Tool), minimap2 (third generation sequencing data Alignment Tool), and the like.
S540, according to mutation data of mutation in the data quality control data, the allelic variation abundance of the alternative sites of the reference sample data is obtained.
The mutation data of the mutation in the data quality control data comprise the gene components of each alternative site.
Specifically, the allelic variation abundance of each candidate site of the reference sample data can be calculated according to the mutation data of the mutation in the data quality control. The specific calculation process can be seen from the technical scheme.
S550, screening target sites according to the crowd frequency of each candidate site.
Specifically, among a plurality of candidate sites of the reference sample data, candidate sites with crowd frequency within a preset crowd frequency range can be screened as target sites.
S560, carrying out characteristic engineering on each target site according to the allelic variation abundance of each target site of the reference sample data.
S570, constructing a sample pollution detection model.
S580, parameter adjustment is carried out on the sample pollution detection model.
S590, obtaining a trained sample pollution detection model.
The method comprises the steps of carrying out gene sequencing on reference sample data to obtain gene sequencing data, carrying out data quality control on the gene sequencing data to obtain data quality control data, carrying out data comparison on the data quality control data and a reference genome, determining mutation data with mutation in the data quality control data, obtaining allelic variation abundance of candidate sites of the reference sample data according to the mutation data with mutation in the data quality control data, screening target sites according to crowd frequency of each candidate site, carrying out characteristic engineering on each target site according to allelic variation abundance of each target site of the reference sample data, constructing a sample pollution detection model, carrying out parameter adjustment on the sample pollution detection model to obtain a trained sample pollution detection model, avoiding the problems of time and labor waste in identifying sample pollution manually, realizing high-throughput detection of sample pollution, improving sample pollution detection efficiency, simultaneously avoiding sample pollution detection standard variability among different manual works, and improving sample pollution detection accuracy.
Example III
Fig. 6 is a schematic structural diagram of a sample contamination detection apparatus according to a third embodiment of the present invention. The embodiment of the invention is applicable to the detection of sample pollution, the device can execute a sample pollution detection method, the device can be realized in a form of hardware and/or software, and the device can be configured in electronic equipment carrying a sample pollution detection function.
Referring to the sample contamination detection apparatus shown in fig. 6, comprising: a sample data acquisition module to be detected 610 and a sample detection result determination module 620. The sample data to be detected obtaining module 610 is configured to obtain the allelic variation abundance of the target site of the sample data to be detected; the sample detection result determining module 620 is configured to input the allelic variation abundance of the target site of the sample data to be detected into a sample pollution detection model trained in advance, so as to obtain a sample detection result and a sample pollution probability of the sample to be detected.
According to the technical scheme, the allele variation abundance of the target site of the sample data to be detected is obtained and is input into the sample pollution detection model trained in advance, so that the sample detection result and the sample pollution probability of the sample to be detected are obtained, the problems of time and labor waste in manually identifying sample pollution are avoided, high-flux detection of sample pollution is realized, the sample pollution detection efficiency is improved, meanwhile, the difference of sample pollution detection standards among different workers is avoided, and the sample pollution detection accuracy is improved.
In an alternative embodiment of the present invention, the sample data to be detected acquisition module 610 includes: the sample data gene sequencing unit is used for carrying out gene sequencing on the sample data to be detected to obtain crowd frequency and genome components of alternative sites of the sample data to be detected; the target site screening unit is used for screening target sites with crowd frequency in a preset crowd frequency range from a plurality of candidate sites of sample data to be detected; the allelic variation abundance calculating unit is used for calculating the allelic variation abundance of the target site according to the gene component of the target site.
In an alternative embodiment of the present invention, after the sample data to be detected acquisition module 610 acquires the allelic variation abundance of the target site of the sample data to be detected, the sample contamination detection apparatus further includes: the sample pollution correlation analysis module is used for carrying out sample pollution correlation analysis on the allelic variation abundance of each target site; and the target site eliminating module is used for eliminating target sites of which the sample pollution correlation analysis results are lower than a preset correlation threshold value from all target sites.
The sample pollution detection device provided by the embodiment of the invention can execute the sample pollution detection method provided by any embodiment of the invention, and has the corresponding functional modules and beneficial effects of the execution method.
In the technical scheme of the embodiment of the invention, the acquisition, storage, application and the like of the allelic variation abundance and the like of the target site of the related sample data to be detected accord with the regulations of related laws and regulations, and the public sequence is not violated.
Example IV
Fig. 7 is a schematic structural diagram of a model training device according to a fourth embodiment of the present invention. The embodiment of the invention is applicable to the training of the sample pollution detection model, the device can execute a model training method, the device can be realized in a hardware and/or software mode, and the device can be configured in electronic equipment carrying the model training function.
Referring to fig. 7, the model training apparatus includes: a reference sample data acquisition module 710, a reference detection result determination module 720, a standard detection result acquisition module 730, and a sample contamination detection model training module 740. Wherein, the reference sample data acquisition module 710 is configured to acquire allele variation abundance of a target site of the reference sample data; the reference detection result determining module 720 is configured to input the allelic variation abundance of the target site of the reference sample data into the sample pollution detection model, so as to obtain a reference detection result and a reference pollution probability of the sample to be detected; a standard detection result obtaining module 730, configured to obtain a standard detection result of the reference sample data; the sample pollution detection model training module 740 is configured to train the sample pollution detection model according to a difference between the reference detection result and a standard detection result of the reference sample data; the sample pollution detection model can realize the sample pollution detection method of any embodiment of the invention.
According to the technical scheme, the allele variation abundance of the target site of the reference sample data is obtained, the allele variation abundance of the target site of the reference sample data is input into the sample pollution detection model, the reference detection result and the reference pollution probability of the sample to be detected are obtained, the standard detection result of the reference sample data is obtained, the sample pollution detection model is trained according to the difference between the reference detection result and the standard detection result of the reference sample data, the sample pollution detection model is trained, the problem that sample pollution is time-consuming and labor-consuming in manual identification is avoided, high-throughput detection of sample pollution is realized, the sample pollution detection efficiency is improved, meanwhile, the difference of sample pollution detection standards among different workers is avoided, and the sample pollution detection accuracy is improved.
In an alternative embodiment of the present invention, the standard detection result obtaining module 730 includes: a first standard detection result acquisition unit for acquiring standard detection results of experimentally pre-labeled reference sample data. Sample contamination detection model training module 740, comprising: and the first sample pollution detection model training unit is used for training the sample pollution detection model according to the difference between the reference detection result and the standard detection result of the reference sample data marked in advance in the experiment.
In an alternative embodiment of the present invention, the reference sample data acquisition module 710 includes: the pollution mark determining unit is used for analyzing the distribution of the allelic variation abundance of the target site of the reference sample data, determining the pollution mark of the reference sample data, and distinguishing whether the reference sample data is polluted or not; and the second standard detection result determining unit is used for determining the pollution mark of the reference sample data as the standard detection result of the reference sample data.
The model training device provided by the embodiment of the invention can execute the model training method provided by any embodiment of the invention, and has the corresponding functional modules and beneficial effects of the execution method.
In the technical scheme of the embodiment of the invention, the acquisition, storage, application and the like of the allelic variation abundance of the target site of the reference sample data, the standard detection result of the reference sample data marked in advance by experiments and the like all meet the regulations of related laws and regulations, and the public welfare is not violated.
Example five
Fig. 8 shows a schematic structural diagram of an electronic device 800 that may be used to implement an embodiment of the invention. Electronic devices are intended to represent various forms of digital computers, such as laptops, desktops, workstations, personal digital assistants, servers, blade servers, mainframes, and other appropriate computers. Electronic equipment may also represent various forms of mobile devices, such as personal digital processing, cellular telephones, smartphones, wearable devices (e.g., helmets, glasses, watches, etc.), and other similar computing devices. The components shown herein, their connections and relationships, and their functions, are meant to be exemplary only, and are not meant to limit implementations of the inventions described and/or claimed herein.
As shown in fig. 8, the electronic device 800 includes at least one processor 801, and a memory such as a Read Only Memory (ROM) 802, a Random Access Memory (RAM) 803, etc., communicatively connected to the at least one processor 801, wherein the memory stores a computer program executable by the at least one processor, and the processor 801 may perform various appropriate actions and processes according to the computer program stored in the Read Only Memory (ROM) 802 or the computer program loaded from the storage unit 808 into the Random Access Memory (RAM) 803. In the RAM 803, various programs and data required for the operation of the electronic device 800 can also be stored. The processor 801, the ROM 802, and the RAM 803 are connected to each other by a bus 804. An input/output (I/O) interface 805 is also connected to the bus 804.
Various components in electronic device 800 are connected to I/O interface 805, including: an input unit 806 such as a keyboard, mouse, etc.; an output unit 807 such as various types of displays, speakers, and the like; a storage unit 808, such as a magnetic disk, optical disk, etc.; and a communication unit 809, such as a network card, modem, wireless communication transceiver, or the like. The communication unit 809 allows the electronic device 800 to exchange information/data with other devices through a computer network such as the internet and/or various telecommunication networks.
The processor 801 may be a variety of general and/or special purpose processing components with processing and computing capabilities. Some examples of processor 801 include, but are not limited to, a Central Processing Unit (CPU), a Graphics Processing Unit (GPU), various specialized Artificial Intelligence (AI) computing chips, various processors running machine learning model algorithms, digital Signal Processors (DSPs), and any suitable processor, controller, microcontroller, etc. The processor 801 performs the various methods and processes described above, such as a sample contamination detection method or a model training method.
In some embodiments, the sample contamination detection method or model training method may be implemented as a computer program tangibly embodied on a computer-readable storage medium, such as storage unit 808. In some embodiments, part or all of the computer program may be loaded and/or installed onto the electronic device 800 via the ROM 802 and/or the communication unit 809. When the computer program is loaded into RAM 803 and executed by processor 801, one or more of the steps of the sample contamination detection method or model training method described above may be performed. Alternatively, in other embodiments, the processor 801 may be configured to perform the sample contamination detection method or the model training method in any other suitable manner (e.g., by means of firmware).
Various implementations of the systems and techniques described here above can be implemented in digital electronic circuitry, integrated circuit systems, field Programmable Gate Arrays (FPGAs), application Specific Integrated Circuits (ASICs), application Specific Standard Products (ASSPs), systems On Chip (SOCs), complex Programmable Logic Devices (CPLDs), computer hardware, firmware, software, and/or combinations thereof. These various embodiments may include: implemented in one or more computer programs, the one or more computer programs may be executed and/or interpreted on a programmable system including at least one programmable processor, which may be a special purpose or general-purpose programmable processor, that may receive data and instructions from, and transmit data and instructions to, a storage system, at least one input device, and at least one output device.
A computer program for carrying out methods of the present invention may be written in any combination of one or more programming languages. These computer programs may be provided to a processor of a general purpose computer, special purpose computer, or other programmable data processing apparatus, such that the computer programs, when executed by the processor, cause the functions/acts specified in the flowchart and/or block diagram block or blocks to be implemented. The computer program may execute entirely on the machine, partly on the machine, as a stand-alone software package, partly on the machine and partly on a remote machine or entirely on the remote machine or server.
In the context of the present invention, a computer-readable storage medium may be a tangible medium that can contain, or store a computer program for use by or in connection with an instruction execution system, apparatus, or device. The computer readable storage medium may include, but is not limited to, an electronic, magnetic, optical, electromagnetic, infrared, or semiconductor system, apparatus, or device, or any suitable combination of the foregoing. Alternatively, the computer readable storage medium may be a machine readable signal medium. More specific examples of a machine-readable storage medium would include an electrical connection based on one or more wires, a portable computer diskette, a hard disk, a Random Access Memory (RAM), a read-only memory (ROM), an erasable programmable read-only memory (EPROM or flash memory), an optical fiber, a portable compact disc read-only memory (CD-ROM), an optical storage device, a magnetic storage device, or any suitable combination of the foregoing.
To provide for interaction with a user, the systems and techniques described here can be implemented on an electronic device having: a display device (e.g., a CRT (cathode ray tube) or LCD (liquid crystal display) monitor) for displaying information to a user; and a keyboard and a pointing device (e.g., a mouse or a trackball) through which a user can provide input to the electronic device. Other kinds of devices may also be used to provide for interaction with a user; for example, feedback provided to the user may be any form of sensory feedback (e.g., visual feedback, auditory feedback, or tactile feedback); and input from the user may be received in any form, including acoustic input, speech input, or tactile input.
The systems and techniques described here can be implemented in a computing system that includes a background component (e.g., as a data server), or that includes a middleware component (e.g., an application server), or that includes a front-end component (e.g., a user computer having a graphical user interface or a web browser through which a user can interact with an implementation of the systems and techniques described here), or any combination of such background, middleware, or front-end components. The components of the system can be interconnected by any form or medium of digital data communication (e.g., a communication network). Examples of communication networks include: local Area Networks (LANs), wide Area Networks (WANs), blockchain networks, and the internet.
The computing system may include clients and servers. The client and server are typically remote from each other and typically interact through a communication network. The relationship of client and server arises by virtue of computer programs running on the respective computers and having a client-server relationship to each other. The server can be a cloud server, also called a cloud computing server or a cloud host, and is a host product in a cloud computing service system, so that the defects of high management difficulty and weak service expansibility in the traditional physical host and VPS (Virtual Private Server ) service are overcome.
It should be appreciated that various forms of the flows shown above may be used to reorder, add, or delete steps. For example, the steps described in the present invention may be performed in parallel, sequentially, or in a different order, so long as the desired results of the technical solution of the present invention are achieved, and the present invention is not limited herein.
The above embodiments do not limit the scope of the present invention. It will be apparent to those skilled in the art that various modifications, combinations, sub-combinations and alternatives are possible, depending on design requirements and other factors. Any modifications, equivalent substitutions and improvements made within the spirit and principles of the present invention should be included in the scope of the present invention.
Claims (10)
Priority Applications (1)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| CN202310273973.1A CN116244602A (en) | 2023-03-20 | 2023-03-20 | Sample pollution detection and model training method, device, equipment and medium |
Applications Claiming Priority (1)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| CN202310273973.1A CN116244602A (en) | 2023-03-20 | 2023-03-20 | Sample pollution detection and model training method, device, equipment and medium |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| CN116244602A true CN116244602A (en) | 2023-06-09 |
Family
ID=86635005
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| CN202310273973.1A Pending CN116244602A (en) | 2023-03-20 | 2023-03-20 | Sample pollution detection and model training method, device, equipment and medium |
Country Status (1)
| Country | Link |
|---|---|
| CN (1) | CN116244602A (en) |
Cited By (1)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN116612814A (en) * | 2023-06-14 | 2023-08-18 | 上海睿璟生物科技有限公司 | Regression model-based batch detection method, device, equipment and medium for gene sample pollution |
Citations (4)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US20180373832A1 (en) * | 2017-06-27 | 2018-12-27 | Grail, Inc. | Detecting cross-contamination in sequencing data |
| US20190065674A1 (en) * | 2017-08-22 | 2019-02-28 | Noblis, Inc. | Nucleic acid sample analysis |
| CN112669903A (en) * | 2020-12-29 | 2021-04-16 | 北京旌准医疗科技有限公司 | HLA typing method and device based on Sanger sequencing |
| CN114530198A (en) * | 2020-11-23 | 2022-05-24 | 福建和瑞基因科技有限公司 | Screening method of SNP (single nucleotide polymorphism) sites for detecting sample pollution level and detection method of sample pollution level |
-
2023
- 2023-03-20 CN CN202310273973.1A patent/CN116244602A/en active Pending
Patent Citations (4)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US20180373832A1 (en) * | 2017-06-27 | 2018-12-27 | Grail, Inc. | Detecting cross-contamination in sequencing data |
| US20190065674A1 (en) * | 2017-08-22 | 2019-02-28 | Noblis, Inc. | Nucleic acid sample analysis |
| CN114530198A (en) * | 2020-11-23 | 2022-05-24 | 福建和瑞基因科技有限公司 | Screening method of SNP (single nucleotide polymorphism) sites for detecting sample pollution level and detection method of sample pollution level |
| CN112669903A (en) * | 2020-12-29 | 2021-04-16 | 北京旌准医疗科技有限公司 | HLA typing method and device based on Sanger sequencing |
Non-Patent Citations (1)
| Title |
|---|
| EVAN MCCARTNEY-MELSTAD: "VCFcontam: A Machine Learning Approach to Estimate Cross-Sample Contamination from Variant Call Data", 《HTTPS://WWW.RESEARCHGATE.NET/PUBLICATION/350039917_VCFCONTAM_A_MACHINE_LEARNING_APPROACH_TO_ESTIMATE_CROSS-SAMPLE_CONTAMINATION_FROM_VARIANT_CALL_DATA》, 12 March 2012 (2012-03-12), pages 1 - 13 * |
Cited By (2)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN116612814A (en) * | 2023-06-14 | 2023-08-18 | 上海睿璟生物科技有限公司 | Regression model-based batch detection method, device, equipment and medium for gene sample pollution |
| CN116612814B (en) * | 2023-06-14 | 2024-08-27 | 上海睿璟生物科技有限公司 | Regression model-based batch detection method, device, equipment and medium for gene sample pollution |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| CN115035950B (en) | Genotype detection method, sample pollution detection method, device, equipment and medium | |
| Daber et al. | Understanding the limitations of next generation sequencing informatics, an approach to clinical pipeline validation using artificial data sets | |
| CN107229841B (en) | A kind of genetic mutation appraisal procedure and system | |
| CN110070915A (en) | The next generation utilizes the Prognosis in Breast Cancer prediction technique and forecasting system based on machine learning of base sequence analysis | |
| Keightley et al. | Inferring the frequency spectrum of derived variants to quantify adaptive molecular evolution in protein-coding genes of Drosophila melanogaster | |
| CN108763859B (en) | Method for establishing analog data set required for providing CNV detection based on unknown CNV sample | |
| WO2024140368A1 (en) | Sample cross contamination detection method and device | |
| Marciano et al. | Developmental validation of PACE™: Automated artifact identification and contributor estimation for use with GlobalFiler™ and PowerPlex® fusion 6c generated data | |
| CN116244602A (en) | Sample pollution detection and model training method, device, equipment and medium | |
| Min et al. | Spatial structure alters the site frequency spectrum produced by hitchhiking | |
| CN116665773B (en) | Method and device for evaluating pathogenicity of mutant genes based on Bayesian algorithm | |
| KR101770962B1 (en) | A method and apparatus of providing information on a genomic sequence based personal marker | |
| CN111370065B (en) | Method and device for detecting cross-sample contamination rate of RNA | |
| CN110444253B (en) | Method and system suitable for mixed pool gene positioning | |
| Wiehe et al. | Identification of selective sweeps using a dynamically adjusted number of linked microsatellites | |
| Kuo et al. | Illuminating the dark side of the human transcriptome with TAMA Iso-Seq analysis | |
| Weiner et al. | Inferring replication timing and proliferation dynamics from single-cell DNA sequencing data | |
| CN118866116B (en) | A method, device, system and storage medium for analyzing contamination of sequencing samples | |
| CN119724337A (en) | Cross-contamination prediction methods and related products | |
| Braichenko et al. | Polymorphism-aware models in RevBayes: species trees, disentangling balancing selection, and GC-biased gene conversion | |
| CN119418764A (en) | Gene single nucleotide polymorphism site analysis method based on SNaPshot | |
| CN121175754A (en) | Prediction model of gRNA HDR potential based on indel spectrum | |
| CN117253540A (en) | Single cell characterization methods, devices, equipment and storage media | |
| CN113981070B (en) | Method, device, equipment and storage medium for detecting embryo chromosome microdeletion | |
| CN115985399A (en) | HRD panel site selection optimization method and system for high-throughput sequencing |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| PB01 | Publication | ||
| PB01 | Publication | ||
| SE01 | Entry into force of request for substantive examination | ||
| SE01 | Entry into force of request for substantive examination |