Detailed Description
The invention provides a chimeric antigen receptor CAR, which comprises the following structures from N end to C end: shRNA sequence targeting Blimp-1 or shRNA negative control sequence, single chain variable region sequence of CD19 antibody, CD8 alpha finger, CD8 alpha TM,4-1BB and CD3 ζ.
The Blimp-1 is B lymphocyte induced mature protein 1 (B lymphocyte induced maturation protein 1, blimp-1), and the Blimp-1 promotes the terminal differentiation of T cells, is highly expressed in Teff and Tex, and inhibits the proliferation of the T cells and the formation of memory T cells; blimp-1 regulates secretion of the immunocompetent molecule Granzyme B. In addition, blimp-1 is also involved in regulating the expression of T-bet, eomes, bcl-2, bcl-2, IL-10, IL-35 and other genes. Thus, downregulating Blimp-1 expression in CAR-T cells is presumed to improve CAR-T cell depletion status, reversing CAR-T cell effector function.
The present invention designs 3 RNAi sequences based on the published Blimp-1 (PRDM 1) gene sequence (Genebank number: NM-001198) as shown in Table 1.
TABLE 1 Easy-siRNA design
The nucleotide sequence of the single-chain variable region sequence of the CD19 antibody is preferably shown as SEQ ID NO.5, the nucleotide sequence of CD8 alpha finger is shown as SEQ ID NO.6, the nucleotide sequence of CD8 alpha TM is shown as SEQ ID NO.7, the nucleotide sequence of 4-1BB is shown as SEQ ID NO.8, and the nucleotide sequence of CD3 zeta is shown as SEQ ID NO. 9.
According to the invention, the CARs are formed after the sequences are sequentially connected, and because the target Blimp-1 sequences in the CAR structure are different, the two CARs are respectively named as sh Blimp-1CAR and sh NT CAR, the amino acid sequence of the sh Blimp-1CAR is preferably shown as SEQ ID NO.10, and the nucleotide sequence is preferably shown as SEQ ID NO. 12; the amino acid sequence of sh NT CAR is preferably set forth in SEQ ID NO.11; the nucleotide sequence is preferably as shown in SEQ ID NO.13.
In the embodiment of the present invention, the best PRDM1-2# is preferably used as the best shRNA for constructing the CAR, but it cannot be considered as the full scope of the present invention. The optimal shRNA is coupled with a basic CAR frame sequence, and U6 is used as a promoter.
The invention also provides application of the CAR in constructing lentiviral vectors and CAR-T cells.
The present invention preferably inserts the nucleotide sequences of the above-mentioned CARs into lentiviral vectors, respectively, and packages the lentiviruses. The base vector of the lentiviral vector of the invention preferably comprises a GV401 vector, a pH elper 1.0 vector or a pH elper 2.0 vector, and the site of insertion is preferablyPacI/PacI. The vectors used in the packaging lentiviruses of the present invention are preferably all available from Shanghai Ji Kai company.
After the lentiviral vector corresponding to the CAR is constructed, the T cells are transfected by the lentiviral vector, and two CAR-T cells, namely the sh Blimp-1CAR-T cells and the sh NT CAR-T cells, are constructed.
The invention also provides a lentiviral vector constructed based on the CAR.
The invention also provides a CAR-T cell comprising the CAR described above.
The CAR-T cell exhaustion degree is improved, memory phenotype groups are taken as the main, the proliferation capacity is high, the immune check point molecule expression is down-regulated, and the expression level of genes related to the T cell exhaustion is correspondingly changed.
The invention also provides application of the CAR or the lentiviral vector or the CAR-T cell in preparing a medicament for treating hematological tumors.
The hematological neoplasm of the present invention preferably comprises a CD19 positive neoplasm.
For further explanation of the present invention, the construction and application of a sh Blimp-1CAR-T cell provided by the present invention will be described in detail with reference to the accompanying drawings and examples, which should not be construed as limiting the scope of the present invention.
Example 1
Bioinformatics analysis of the role of Blimp-1 in tumor immunotherapy
1.1 Differential analysis of Blimp-1 in cancer tissue and paracancerous tissue
The expression of Blimp-1 (PRDM 1) in various tumor (TCGA database) cancer tissues and paracancerous tissues was analyzed using TIMER 2.0.
As shown in FIG. 1, blimp-1 is differentially expressed in various cancerous and paracancerous tissues.
1.2 Expression of Blimp-1 in tumor-infiltrating immune cells
The expression of Blimp-1 (PRDM 1) in tumor infiltrating immune cells, including innate immune cells and acquired immune cells, was analyzed using TIMER 2.0.
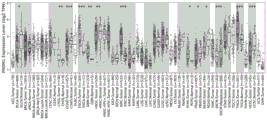
As shown in FIG. 2, blimp-1 was distributed among tumor immunoinfiltrating cells, where CD8+ T cells were highly expressed and CD8+ T was the most predominant immune cell involved in tumor immunity.
1.3 Clinical relevance analysis of Blimp-1
The correlation of Blimp-1 (PRDM 1) with the survival time of tumor patients was analyzed by TIMER2.0, and predictive analysis was performed on whether it could be used as a tumor risk assessment factor.
As shown in FIG. 3, blimp-1 expression is correlated with prognosis of tumor patients, and can be used as one of risk assessment factors, namely, the survival quality of patients with high Blimp-1 expression is lower than that of patients with low Blimp-1 expression.
1.4 Correlation analysis of Blimp-1 and T cell depletion related genes
The TISIDB is used for analyzing the relativity between Blimp-1 (PRDM 1) and the gene which has definitely regulated and controlled the depletion of T cells, and the relativity mainly comprises three major classes of immune check points, transcription factors and Blimp-1 target genes.
As shown in FIG. 4, blimp-1 is positively correlated with immune checkpoint molecules such as PDCD1, CTLA-4, LAG-3, HAVCR2, TIGIT, etc., has a certain correlation with transcription factors TOX, TOX2, NR4A family, etc., which have been reported to be involved in cell depletion, and moreover, has a tendency to eliminate each other with its regulatory target genes BCL-2, BCL-6, TBX21, ID3, etc.
Example 2
shRNA sequence for targeting Blimp-1 based on shRNA gene editing technology
2.1 shRNA sequence design
(1) Based on the published Blimp-1 (PRDM 1) gene sequence, 3 RNAi sequences shown in Table 1 were designed. (2) The obtained siRNA sequences were further designed as shRNA sequences (table 2).
TABLE 2 Synthesis of oligo information
2.2 RNAi lentiviral vector construction and packaging
The designed shRNA sequence is further synthesized and annealed to form double-stranded DNA, and the linearization vector and the double-stranded DNA are connected through T4 DNAligenase overnight; taking 10 mu L of overnight connection product, and performing transformation; culturing transformed bacterial liquid, and then carrying out colony PCR (polymerase chain reaction) to identify the connection efficiency; sequencing is then performed. Comparing the sequencing result with a target gene sequence for analysis; and (5) taking the bacterial liquid with the correct sequence identification, carrying out shake culture overnight, and extracting plasmids.
2.3 lentivirus preparation
The invention adopts a three-plasmid packaging system to package lentiviruses, and mainly comprises a lentivirus vector containing a target gene and 2 auxiliary packaging plasmids. (the carriers used are Shanghai Ji Kai company)
(1) Preparing the plasmids required for virus packaging, and extracting and purifying the plasmids required according to the instructions of a plasmid extraction kit. Ensuring that high-purity plasmid (A260/A280 is 1.8-2.0) is obtained; (2) 293T cells (density about 70-80%) cultured overnight with plasmid transfection as above were transfected for 48 hours and cell culture supernatants were collected; (3) concentrating the collected culture supernatant by high-speed centrifugation at 50 mu L/branch, subpackaging, and preserving at-80deg.C; (4) and taking a proper amount of concentrated virus for sterile and fluorescent titer determination. Viral titer = number of fluorescent cells/amount of viral stock
Through detection, each group of lentiviruses has a titer of more than 10 8 TUmL, specific titers of each group of lentiviruses are shown in Table 3.
TABLE 3 shRNA lentiviral titres
2.4 optimal shRNA sequence selection
In the invention, the human leukemia cell strain K562 cells and the human primary T cells are simultaneously used as tool cells for screening the best shRNA sequence of the target Blimp-1.
(1) Lentiviruses infect K562 cells. The well-conditioned K562 cell density was adjusted to 1X 10 6 And (3) adding shRNA lentivirus with corresponding volume according to MOI=20, gently mixing, centrifuging by a horizontal centrifuge (1000 rpm,30 min), culturing at 37 ℃ for 18 hours, adding fresh complete culture medium with equivalent amount, continuously culturing, and observing virus infection efficiency under a mirror after 72 hours. On day 3, day 5 and day 7, cells were collected, total cellular RNA was extracted, and the effect on Blimp-1 mRNA expression levels was detected by RT-qPCR using different shRNA sequences. Meanwhile, whole protein of the 7d cell is collected, and the change of the Blimp-1 protein level is detected.
(2) Lentiviruses infect primary T cells.
Day0, preparing T cell activation culture plates, namely diluting Anti-human CD3/CD28 into sterile PBS (phosphate buffered saline) one Day before T cells are separated, wherein the final concentration is 1 mug/mL, a 24-pore plate is used, and 400 mu L of coating liquid is added into each pore;
day1 separating and purifying human peripheral blood CD3+ T cells, and preparing culture plates required by the next Day of virus infection of the T cells, namely 48-hole cell culture plates
(20. Mu.g/mL), anti-human CD3/CD28 (5. Mu.g/mL), 200. Mu.L of the system;
day2 lentivirus infects T cells.
The next day, after PBS washes virus infects the culture plate 1 time, according to MOI value, add slow virus of corresponding volume in each hole, 2000g,2h horizontal centrifugation; t cells were resuspended at 1X 10 in complete medium 6 Adding 200 mu L of cell suspension into a virus infection plate per mL, and centrifuging at 1000g and 1h level; after culturing for 24 hours in an incubator, adding an equal volume of complete T cell culture medium into each hole;
after the lentivirus infects the T cells for 72 hours, preliminarily observing the positive rate of the lentivirus infects the T cells under a fluorescence microscope; cells were collected on day 7, total cellular RNA and total cellular protein were extracted, and the effect of Blimp-1 mRNA expression level was detected by RT-qPCR and Western Blot.
The results are shown in fig. 5 and 6, where the lentivirus successfully infects K562 cells and T cells. Out of the three shRNA sequences designed, sh PRDM 12 # down-regulates Blimp-1 expression by about 50%, with a trend consistent in K562 and T cells. In addition, RT-qPCR results at different time points show that Blimp-1 expression can be stably down-regulated by using the sequence.
Example 3
Preparation of sh NT CAR-T, sh Blimp-1CAR-T cells
After the best target Blimp-1 shRNA sequence was obtained, it was coupled to the basal CAR frame sequence, with U6 as promoter. Lentiviruses and infected T cells were prepared using the procedure in example 2. The amino acid sequences of the sh Blimp-1CAR and the sh NT CAR are respectively SEQ ID NO.10 and SEQ ID NO.11; the nucleotide sequences are SEQ ID NO.12 and SEQ ID NO.13 respectively.
The shNT CAR, sh Blimp-1CAR framework design is shown in FIG. 7A. The CAR-T cell infection is shown in FIG. 7B. Flow cytometry detection showed that each group had a CAR-T cell positive rate above 70% (C and D in fig. 7).
Example 4
Detection of memory phenotype, cell viability and proliferation Capacity of sh NT, sh Blimp-1CAR-T cells
4.1 memory phenotype detection:
(1) collecting CAR-T cells of each group after 4d of spermidine treatment with different concentrations, and centrifugally washing; (2) adjustment ofCell density of 1X 10 6 /mL; (3) 100. Mu.L of the cell suspension was aspirated, and 5. Mu.L of BB700 Rat Anti-Human CCR7 (CD 197) and 5. Mu.L of APC Mouse Anti-Human CD45RA were added to each tube and mixed gently; (4) incubating for 20-30min at room temperature in a dark place, and adding 500 mu L of PBS to wash the unbound antibody; (5) the 300. Mu.L PBS resuspended cells were assayed on-line within 1 h.
The results are shown in FIG. 8, where after Blimp-1 down-regulation, the proportion of TCM, TEM cell populations in CAR-T cells increased, while the effector cell phenotype decreased. The increase in the memory cell population facilitates the clearance of CAR-T cells from tumor cells.
4.2 cell viability assay:
(1) collecting each group of T cells treated by spermidine with different concentrations for 4 days, and centrifugally washing; (2) cell pellet was resuspended in 1×binding buffer and the cell density was adjusted to 1×10 6 /mL; (3) dyeing according to the reagent instruction; (4) incubation for 20-30min at room temperature in dark place, adding 400. Mu.L of 1 Xbinding buffer, and performing on-line analysis within 1 h.
By apoptosis detection of the CAR-T cells, the result is shown in figure 9, a large number of apoptotic cells are not found in the sh Blimp-1CAR-T group cells, and the fact that the downregulation of Blimp-1 has no obvious influence on T cell survival is shown, so that the subsequent experiments can be continuously carried out.
4.3 proliferation potency assay:
the proliferation potency was measured by the fluorescent dye method of Cell Trace Far Red, and the tumor cells were treated with mitomycin C prior to the cell proliferation potency measurement, so that they were not proliferation potency while ensuring viability.
The cell trace specific staining procedure is as follows:
(1) diluting 20 mu L of mother solution to a working concentration (1 mu M) by adopting PBS, adding a corresponding volume of dye solution according to a cell counting result, re-suspending and precipitating, and incubating for 20min at 37 ℃ in a dark place; (2) adding 4-5 times of serum-containing culture medium of original staining system to stop staining (37 deg.C, 5 min); centrifuging to remove the supernatant, and culturing in a cell culture box for at least 1h to fully combine the dye liquor and the cells; (3) according to the effective target ratio of 20:1, each group of CAR-T cells were co-cultured with mitomycin C-treated tumor cells for 24 hours and then examined for cell proliferation by flow cytometry.
The proliferation capacity of the shBlimp-1CAR-T cells is increased compared with that of shNTCAR-T cells under the stimulation of Daudi cells. When K562 cells are used for stimulation, no obvious proliferation of sh NTCAR-T cells and sh Blimp-1CAR-T cells occurs, which indicates that the CAR-T cells show antigen-dependent proliferation, namely only positive target cells are identified, and no off-target side effect exists. The results are shown in FIG. 10.
Example 5
Detection of sh NT CAR-T, sh Blimp-1CAR-T cell immune checkpoint expression
(1) Collecting each group of CAR-T cells which are subjected to in vitro amplification culture for about 7 days; (2) cell density was adjusted to 1X 10 6 /mL; (3) 100. Mu.L of cell suspension was aspirated, 5. Mu.L of APC Mouse Anti-Human CD279 (PD-1), BV421 Mouse Anti-Human CD152 (CTLA 4), PE Mouse Anti-Human TIGIT flow antibody was added to each tube, and mixed gently; (4) incubating for 20-30min at room temperature in dark, adding 500 μl PBS, and washing to remove unbound antibody; (5) cell pellet was resuspended in 300 μl PBS and assayed on-press within 1 h.
The results are shown in FIG. 11, in which there is some reduction in PD-1, CTLA-4 and TIGIT expression after down-regulating Blimp-1 expression, suggesting that Blimp-1 low expression may increase the killing of tumor cells by CAR-T cells.
Example 6
Detection of sh NT CAR-T, sh Blimp-1CAR-T cytokine secretion levels
sh NT CAR-T, sh Blimp-1CAR-T cells were co-cultured with tumor cells (K562, daudi) at a 10:1 effective target ratio for 24h, and cell supernatants were collected and assayed for IL-2, IFN-gamma levels by ELISA. According to LEGEND MAXTM Human IFN-gamma, the IL-2 ELISA Kit reagent instructions were run and all required reagents were pre-equilibrated to room temperature prior to performing the official experiments. As shown in FIGS. 12-13, after Blimp-1 down-regulation, the IL-2 secretion level of CAR-T cells was significantly increased compared to shNT CAR-T cells after 24h of Daudi cell stimulation. Similarly, IFN-gamma secretion capacity is also increased. However, there was no significant difference between IL-2 and IFN-gamma expression upon stimulation of K562 cells.
Example 7
Detection of sh NT CAR-T, sh Blimp-1CAR-T cell cytotoxicity in vitro using LDH assay
Grouping is set up and operated according to the reagent instructions. Cytotoxicity:
Cytolisis%=(Experiment-Effector Spontaneous-Target Spontaneous)/(Target Maxium-Target Spontaneous)×100
the results are shown in FIG. 14, the ability of sh Blimp-1CAR-T to kill tumor cells is superior to sh NT CAR-T cells, and similar results exist at different target ratios. With the increase of the effective target ratio, the killing rate of shNT and sh Blimp-1CAR-T cells on tumor cells can be increased by 90%.
Example 8
Detecting the expression condition of T cell depletion related genes after Blimp-1 down regulation
Extracting total RNA of cells by using a Trizol method; cDNA library synthesis; the RNA was reverse transcribed to cDNA (RevertAid Fisrt St ran d cDN A Synthesis kit).
The qRT-PCR primer sequences contained in the invention are shown in Table 4.
TABLE 4 qRT-PCR primer sequences
qRT-PCR reaction System (20. Mu.L): cDNA 1. Mu.L, upstream primer 1.4. Mu.L, downstream primer 1.4. Mu. L, mix 12. Mu. L, DEPC water 4.2. Mu.L. qRT-PCR procedure: 95 ℃ for 2min; circulation is carried out at 95 ℃ for 5s and 60 ℃ for 30s and 45; 95℃15s,60℃1min,95℃15s.
The results are shown in FIG. 15, where the level of immune checkpoint molecule expression was reduced following Blimp-1 down-regulation, with TIGIT down-regulation being most pronounced; secondly, BCL-2, BCL-6 and ID-3 expression are increased after Blimp-1 is down-regulated, and the genes mainly participate in resisting apoptosis, so that the in vitro proliferation advantage of shBlimp-1CAR-T cells is explained; finally, the TOX, TOX2, NR4A family did not show significant downregulation after Blimp-1 downregulation, suggesting that Blimp-1 may regulate T cell depletion status through other genes.
Example 9
sh NT CAR-T, sh Blimp-1CAR-T cell in vivo experiments
The animal experiments related to the invention are carried out after the welfare ethical examination of the experimental animal. The animal experiment certificate number is: GDY2104005. NCG mice used in the experiments were purchased from Guangdong Kangdong Biotechnology Co., ltd (license number: SYXK (Guangdong) 2019-0213).
9.1 construction of mouse blood tumor model
After the mice are adaptively raised in the barrier for one week, diet, activity and the like of the mice are observed, so that the mice are in an optimal state, and modeling is started. (1) Resuscitating Daudi-luc cells, and carrying out passage for 2-3 times to ensure the activity of tumor cells in a proliferation stage; (2) tumor cell resuspension count was collected at 5×10 6 /mL; (3) the mice were intravenously injected after the tail vein was disinfected, 200 μl/mouse; (4) after one week of molding, live imaging was performed to determine whether the mouse molding was successful.
9.2 injection of CAR-T cells
After the success of the pre-experiment verification of the modeling of the mice, the formal experiment is carried out, the steps are the same as 9.1, after one week, the tail vein injection is used for in vitro expansion of each group of CAR-T cells with good states for about 10 days, and meanwhile, the equal volume PBS is injected as a control group.
9.3 Living animal imaging
(1) The mice were removed and body weights were weighed to determine the volume of D-fluorescein potassium salt injected; (2) detecting fluorescence value after substrate injection for about 10min and inhalation anesthesia for 3min by isoflurane; (3) the image acquisition process continuously anesthetizing the mice, setting parameters as automatic exposure, and storing the images after the exposure is finished.
9.4 detection of CAR-T cell and tumor cell persistence in mouse peripheral blood
(1) Collecting peripheral blood of a mouse by adopting an anticoagulated micro blood taking needle through an orbital vein, and collecting the peripheral blood in a 1.5mL centrifuge tube pretreated by 0.1% heparin; (2) lysing erythrocytes in peripheral blood; (3) resuspended cell density was 1X 106/mL, stained with CD3, CD20 flow antibody and control tubes were set; incubate at room temperature for 25min under dark conditions and check on a machine.
A blood tumor model was constructed and an in vivo flow chart is shown in fig. 16. Surprisingly, in vivo imaging of small animals after 1 week of CAR-T cell injection compared to in vitro results showed that the tumor burden was significantly higher in the sh Blimp-1CAR-T cell group than in the sh NT CAR-T cell group, although CAR-T groups were effective in delaying tumor progression compared to PBS groups, and the results are shown in fig. 17-18. Considering the in vitro experimental results, the shBlimp-1CAR-T cells show a memory-like phenotype and have increased proliferation and IL-2 secretion capacity, and these factors are involved in T cell self-renewal and survival, therefore, the invention predicts that the shBlimp-1CAR-T cells may last longer in mice. The results showed that there was no significant difference in the ratio of CAR-T cells in peripheral blood between the shNT CAR-T group and the sh Blimp-1CAR-T group at week 4 (p=0.223), but the ratio of T cells remained significantly higher in the sh Blimp-1CAR-T group than in the shNT CAR-T group at week 8 (p=0.071), as shown in fig. 19. Furthermore, in the continuous observations of mice, while shBlimp-1CAR-T group failed to control tumor cells at the initial time with shNT CAR-T group, it can extend the survival time of mice, whereas sh NT CAR-T group mice succumbed to death several weeks after treatment, although no significant tumor burden was observed upon in vivo imaging (fig. 19). Meanwhile, in observing tumor progression, the body weight of the mice was not significantly changed (fig. 20). The above results indicate that the Blimp-1 low expressing group CAR-T cells may extend mouse residence time in a more "gentle" manner.
Example 10
sh NT CAR-T, sh Blimp-1CAR-T cell expression profile characteristics
10.1 RNA-seq sample preparation
Each group of CAR-T cells expanded and cultured for 7 days was collected, and after resuspension, GFP positive cell proportion was analyzed by a flow cytometer and CAR-T cells were collected by sorting according to this signal. The collected cells were used to extract total RNA from the cells by Trizol method.
10.2 RNA-seq data analysis
In order to ensure high quality of data for subsequent analysis, the data obtained by sequencing is subjected to filterability processing, which mainly comprises removing reads with joints, removing reads with base information which cannot be determined, removing low-quality reads, and simultaneously, performing Q20, Q30 and GC content calculation on clean data.
10.3 differential Gene expression analysis
Differential expression analysis between two comparison combinations was performed using DESeq2 software (1.20.0); the resulting P value was adjusted using the methods Benjamini and Hochberg to control the false discovery rate.
10.4 differential Gene enrichment analysis and Gene set enrichment analysis
GO enrichment of the differentially expressed genes was analyzed using clusterif iotaler (3.8.1) software; statistical analysis of KEGG enrichment of differentially expressed genes using clusterif iotaler (3.8.1) software; GO, KEGG datasets of sequencing results were analyzed using GSEA (http:// www.broadinstit. Mu. Te. Org/GSEA/index. Jsp).
The RNA-seq results indicated that the two sets of CAR-T cells exhibited different expression profiles, the 5214 genes of the sh Blimp-1CAR-T cell set were up-regulated, and the 9685 genes were down-regulated (P < 0.05). The results are shown in FIG. 21. The differential gene analysis shows that sh Blimp-1CAR-T cell groups, such as TCF7, ID3, TCF12, bcl2, bcl6, PO M6F 1 and the like are involved in the up-regulation of cell memory gene expression; up-regulating expression of T cell proliferation related genes MKI67, EBI3, CD3EAP, TNFSF10 and TNFRSF9 (4-1 BB); genes involved in proliferation and recruitment of other immune cells such as IL2RB, IL15, IL12RB2, IL18, CCL22, CCL4, CCL20, XCL1, CCR10, etc. are up-regulated; meanwhile, the expression of the genes related to the exhaustion such as HAVCR2, tbx21, EOMES and the like is not obviously up-regulated, and the expression of TOX genes is up-regulated to a certain extent; furthermore, similar to the results of the related study, in shBlimp-1CAR-T group, BATF was up-regulated and SOX4 was down-regulated, indicating that SOX4 was down-regulated by up-regulating BATF to reverse CAR-T cell failure; at the same time, the gene related to FAO is up-regulated, which indicates that the energy uptake and metabolism modes of sh Blimp-1CAR-T cells and sh NT CAR-T are different, and the metabolic pathway also affects the memory capacity and effector function of the cells. The results are shown in FIG. 22.
Enrichment analysis was performed on the differentially expressed genes. The GO analysis result shows that the differential genes in sh Blimp-1CAR-T cells are mainly involved in the biological processes of cytokine production/secretion, T cell migration, T cell proliferation, T cell receptor signals and the like. The KEGG analysis results show that the differential genes are mainly enriched in important pathways such as cytokine-cytokine receptor, glycine metabolism and the like. The GSEA gene set enrichment analysis shows that in the sh Blimp-1CAR-T group, the enriched gene set is mainly involved in the biological processes of memory, proliferation, survival and the like. The results are shown in FIGS. 23 to 27.
The results show that the down-regulating Blimp-1 promotes the memory phenotype formation of the CAR-T cells, increases the proliferation capacity, improves the anti-apoptosis level and cytokine secretion capacity of the CAR-T cells, and prolongs the in-vivo residence time of the CAR-T cells, thereby effectively controlling the blood tumor progression of mice. The enhanced anti-tumor capacity of the CAR-T cells after Blimp-1 down-regulation revealed that Blimp-1 can be used as one of potential research targets for future CAR-T cell treatment, and the action mechanism is shown in FIG. 28.
Although the foregoing embodiments have been described in some, but not all, embodiments of the invention, it should be understood that other embodiments may be devised in accordance with the present embodiments without departing from the spirit and scope of the invention.