CN115044665A - Application of ARG1 in the preparation of sepsis diagnosis, severity judgment or prognosis assessment reagents or kits - Google Patents
Application of ARG1 in the preparation of sepsis diagnosis, severity judgment or prognosis assessment reagents or kits Download PDFInfo
- Publication number
- CN115044665A CN115044665A CN202210661083.3A CN202210661083A CN115044665A CN 115044665 A CN115044665 A CN 115044665A CN 202210661083 A CN202210661083 A CN 202210661083A CN 115044665 A CN115044665 A CN 115044665A
- Authority
- CN
- China
- Prior art keywords
- sepsis
- arg1
- patients
- peripheral blood
- reagent
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Pending
Links
- 206010040047 Sepsis Diseases 0.000 title claims abstract description 113
- 102100021723 Arginase-1 Human genes 0.000 title claims abstract description 63
- 101000752037 Homo sapiens Arginase-1 Proteins 0.000 title claims abstract description 61
- 101000800287 Homo sapiens Tubulointerstitial nephritis antigen-like Proteins 0.000 title claims abstract description 61
- 239000003153 chemical reaction reagent Substances 0.000 title claims abstract description 23
- 238000003745 diagnosis Methods 0.000 title claims abstract description 16
- 238000004393 prognosis Methods 0.000 title claims abstract description 14
- 210000005259 peripheral blood Anatomy 0.000 claims abstract description 40
- 239000011886 peripheral blood Substances 0.000 claims abstract description 40
- 230000014509 gene expression Effects 0.000 claims abstract description 33
- 206010040070 Septic Shock Diseases 0.000 claims abstract description 29
- 230000036303 septic shock Effects 0.000 claims abstract description 29
- 238000001514 detection method Methods 0.000 claims abstract description 12
- 231100000518 lethal Toxicity 0.000 claims abstract description 3
- 230000001665 lethal effect Effects 0.000 claims abstract description 3
- 101150088826 arg1 gene Proteins 0.000 claims description 14
- 239000012472 biological sample Substances 0.000 claims description 10
- 230000003321 amplification Effects 0.000 claims description 4
- 238000003199 nucleic acid amplification method Methods 0.000 claims description 4
- 238000012360 testing method Methods 0.000 claims description 4
- 238000010839 reverse transcription Methods 0.000 claims description 3
- 239000000090 biomarker Substances 0.000 abstract description 35
- 230000003319 supportive effect Effects 0.000 abstract description 7
- 230000035939 shock Effects 0.000 abstract description 3
- 238000011156 evaluation Methods 0.000 abstract description 2
- 108090000623 proteins and genes Proteins 0.000 description 65
- 102000004169 proteins and genes Human genes 0.000 description 15
- 238000000034 method Methods 0.000 description 13
- 241000699670 Mus sp. Species 0.000 description 9
- 238000004458 analytical method Methods 0.000 description 7
- 230000006870 function Effects 0.000 description 7
- 230000004850 protein–protein interaction Effects 0.000 description 7
- 230000001105 regulatory effect Effects 0.000 description 7
- 238000012216 screening Methods 0.000 description 7
- 210000004369 blood Anatomy 0.000 description 6
- 239000008280 blood Substances 0.000 description 6
- 201000010099 disease Diseases 0.000 description 6
- 208000037265 diseases, disorders, signs and symptoms Diseases 0.000 description 6
- 208000024891 symptom Diseases 0.000 description 6
- 230000002103 transcriptional effect Effects 0.000 description 6
- 230000003993 interaction Effects 0.000 description 5
- 108020004414 DNA Proteins 0.000 description 4
- 238000011161 development Methods 0.000 description 4
- 238000002474 experimental method Methods 0.000 description 4
- 238000003012 network analysis Methods 0.000 description 4
- 238000003753 real-time PCR Methods 0.000 description 4
- 230000004044 response Effects 0.000 description 4
- 238000003766 bioinformatics method Methods 0.000 description 3
- 238000003759 clinical diagnosis Methods 0.000 description 3
- 230000004186 co-expression Effects 0.000 description 3
- 238000005516 engineering process Methods 0.000 description 3
- 208000015181 infectious disease Diseases 0.000 description 3
- 239000011159 matrix material Substances 0.000 description 3
- 210000000056 organ Anatomy 0.000 description 3
- 102000004452 Arginase Human genes 0.000 description 2
- 101710129000 Arginase-1 Proteins 0.000 description 2
- 108700024123 Arginases Proteins 0.000 description 2
- 238000011740 C57BL/6 mouse Methods 0.000 description 2
- 102000001708 Protein Isoforms Human genes 0.000 description 2
- 108010029485 Protein Isoforms Proteins 0.000 description 2
- 238000011529 RT qPCR Methods 0.000 description 2
- 150000001413 amino acids Chemical group 0.000 description 2
- 238000009534 blood test Methods 0.000 description 2
- 210000004534 cecum Anatomy 0.000 description 2
- 210000003850 cellular structure Anatomy 0.000 description 2
- 238000010586 diagram Methods 0.000 description 2
- 238000010195 expression analysis Methods 0.000 description 2
- 238000011223 gene expression profiling Methods 0.000 description 2
- 238000012986 modification Methods 0.000 description 2
- 230000004048 modification Effects 0.000 description 2
- 230000035945 sensitivity Effects 0.000 description 2
- 238000013517 stratification Methods 0.000 description 2
- 210000004895 subcellular structure Anatomy 0.000 description 2
- 238000006467 substitution reaction Methods 0.000 description 2
- 210000001519 tissue Anatomy 0.000 description 2
- 239000004475 Arginine Substances 0.000 description 1
- 206010009944 Colon cancer Diseases 0.000 description 1
- 208000001333 Colorectal Neoplasms Diseases 0.000 description 1
- 238000000018 DNA microarray Methods 0.000 description 1
- 108700024394 Exon Proteins 0.000 description 1
- 102100031181 Glyceraldehyde-3-phosphate dehydrogenase Human genes 0.000 description 1
- 101100216634 Homo sapiens ARG1 gene Proteins 0.000 description 1
- 208000002260 Keloid Diseases 0.000 description 1
- 206010023330 Keloid scar Diseases 0.000 description 1
- AHLPHDHHMVZTML-BYPYZUCNSA-N L-Ornithine Chemical compound NCCC[C@H](N)C(O)=O AHLPHDHHMVZTML-BYPYZUCNSA-N 0.000 description 1
- 241001465754 Metazoa Species 0.000 description 1
- 208000034486 Multi-organ failure Diseases 0.000 description 1
- AHLPHDHHMVZTML-UHFFFAOYSA-N Orn-delta-NH2 Natural products NCCCC(N)C(O)=O AHLPHDHHMVZTML-UHFFFAOYSA-N 0.000 description 1
- UTJLXEIPEHZYQJ-UHFFFAOYSA-N Ornithine Natural products OC(=O)C(C)CCCN UTJLXEIPEHZYQJ-UHFFFAOYSA-N 0.000 description 1
- 238000003559 RNA-seq method Methods 0.000 description 1
- XSQUKJJJFZCRTK-UHFFFAOYSA-N Urea Chemical compound NC(N)=O XSQUKJJJFZCRTK-UHFFFAOYSA-N 0.000 description 1
- 230000001154 acute effect Effects 0.000 description 1
- 230000033289 adaptive immune response Effects 0.000 description 1
- ODKSFYDXXFIFQN-UHFFFAOYSA-N arginine Natural products OC(=O)C(N)CCCNC(N)=N ODKSFYDXXFIFQN-UHFFFAOYSA-N 0.000 description 1
- 230000009286 beneficial effect Effects 0.000 description 1
- 239000004202 carbamide Substances 0.000 description 1
- 230000008859 change Effects 0.000 description 1
- 239000002299 complementary DNA Substances 0.000 description 1
- 238000010276 construction Methods 0.000 description 1
- 230000000694 effects Effects 0.000 description 1
- 230000000855 fungicidal effect Effects 0.000 description 1
- 108020004445 glyceraldehyde-3-phosphate dehydrogenase Proteins 0.000 description 1
- 210000003714 granulocyte Anatomy 0.000 description 1
- 238000007417 hierarchical cluster analysis Methods 0.000 description 1
- 210000003917 human chromosome Anatomy 0.000 description 1
- 230000007062 hydrolysis Effects 0.000 description 1
- 238000006460 hydrolysis reaction Methods 0.000 description 1
- 208000026278 immune system disease Diseases 0.000 description 1
- 230000008676 import Effects 0.000 description 1
- 230000006872 improvement Effects 0.000 description 1
- 230000002757 inflammatory effect Effects 0.000 description 1
- 230000015788 innate immune response Effects 0.000 description 1
- 238000009434 installation Methods 0.000 description 1
- 210000001117 keloid Anatomy 0.000 description 1
- 210000003734 kidney Anatomy 0.000 description 1
- 210000000265 leukocyte Anatomy 0.000 description 1
- 210000004185 liver Anatomy 0.000 description 1
- 210000004072 lung Anatomy 0.000 description 1
- 210000004698 lymphocyte Anatomy 0.000 description 1
- 239000003550 marker Substances 0.000 description 1
- 230000007246 mechanism Effects 0.000 description 1
- 238000010197 meta-analysis Methods 0.000 description 1
- 230000004060 metabolic process Effects 0.000 description 1
- 239000002679 microRNA Substances 0.000 description 1
- 239000000203 mixture Substances 0.000 description 1
- 238000010172 mouse model Methods 0.000 description 1
- 230000008383 multiple organ dysfunction Effects 0.000 description 1
- 208000029744 multiple organ dysfunction syndrome Diseases 0.000 description 1
- 108020004707 nucleic acids Proteins 0.000 description 1
- 102000039446 nucleic acids Human genes 0.000 description 1
- 150000007523 nucleic acids Chemical class 0.000 description 1
- 229960003104 ornithine Drugs 0.000 description 1
- 244000052769 pathogen Species 0.000 description 1
- 230000008506 pathogenesis Effects 0.000 description 1
- 230000001717 pathogenic effect Effects 0.000 description 1
- 230000008569 process Effects 0.000 description 1
- 230000005180 public health Effects 0.000 description 1
- 238000011002 quantification Methods 0.000 description 1
- 239000002994 raw material Substances 0.000 description 1
- 238000011160 research Methods 0.000 description 1
- 238000012552 review Methods 0.000 description 1
- 239000000523 sample Substances 0.000 description 1
- 238000011896 sensitive detection Methods 0.000 description 1
- 238000012163 sequencing technique Methods 0.000 description 1
- 238000009120 supportive therapy Methods 0.000 description 1
- 230000004083 survival effect Effects 0.000 description 1
- 230000009897 systematic effect Effects 0.000 description 1
- 108700026220 vif Genes Proteins 0.000 description 1
Images
Classifications
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q1/00—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions
- C12Q1/68—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions involving nucleic acids
- C12Q1/6876—Nucleic acid products used in the analysis of nucleic acids, e.g. primers or probes
- C12Q1/6883—Nucleic acid products used in the analysis of nucleic acids, e.g. primers or probes for diseases caused by alterations of genetic material
-
- G—PHYSICS
- G01—MEASURING; TESTING
- G01N—INVESTIGATING OR ANALYSING MATERIALS BY DETERMINING THEIR CHEMICAL OR PHYSICAL PROPERTIES
- G01N33/00—Investigating or analysing materials by specific methods not covered by groups G01N1/00 - G01N31/00
- G01N33/48—Biological material, e.g. blood, urine; Haemocytometers
- G01N33/50—Chemical analysis of biological material, e.g. blood, urine; Testing involving biospecific ligand binding methods; Immunological testing
- G01N33/53—Immunoassay; Biospecific binding assay; Materials therefor
- G01N33/573—Immunoassay; Biospecific binding assay; Materials therefor for enzymes or isoenzymes
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q2600/00—Oligonucleotides characterized by their use
- C12Q2600/118—Prognosis of disease development
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q2600/00—Oligonucleotides characterized by their use
- C12Q2600/158—Expression markers
-
- G—PHYSICS
- G01—MEASURING; TESTING
- G01N—INVESTIGATING OR ANALYSING MATERIALS BY DETERMINING THEIR CHEMICAL OR PHYSICAL PROPERTIES
- G01N2333/00—Assays involving biological materials from specific organisms or of a specific nature
- G01N2333/90—Enzymes; Proenzymes
- G01N2333/914—Hydrolases (3)
- G01N2333/978—Hydrolases (3) acting on carbon to nitrogen bonds other than peptide bonds (3.5)
-
- G—PHYSICS
- G01—MEASURING; TESTING
- G01N—INVESTIGATING OR ANALYSING MATERIALS BY DETERMINING THEIR CHEMICAL OR PHYSICAL PROPERTIES
- G01N2800/00—Detection or diagnosis of diseases
- G01N2800/52—Predicting or monitoring the response to treatment, e.g. for selection of therapy based on assay results in personalised medicine; Prognosis
-
- Y—GENERAL TAGGING OF NEW TECHNOLOGICAL DEVELOPMENTS; GENERAL TAGGING OF CROSS-SECTIONAL TECHNOLOGIES SPANNING OVER SEVERAL SECTIONS OF THE IPC; TECHNICAL SUBJECTS COVERED BY FORMER USPC CROSS-REFERENCE ART COLLECTIONS [XRACs] AND DIGESTS
- Y02—TECHNOLOGIES OR APPLICATIONS FOR MITIGATION OR ADAPTATION AGAINST CLIMATE CHANGE
- Y02A—TECHNOLOGIES FOR ADAPTATION TO CLIMATE CHANGE
- Y02A90/00—Technologies having an indirect contribution to adaptation to climate change
- Y02A90/10—Information and communication technologies [ICT] supporting adaptation to climate change, e.g. for weather forecasting or climate simulation
Landscapes
- Life Sciences & Earth Sciences (AREA)
- Health & Medical Sciences (AREA)
- Chemical & Material Sciences (AREA)
- Engineering & Computer Science (AREA)
- Immunology (AREA)
- Organic Chemistry (AREA)
- Molecular Biology (AREA)
- Analytical Chemistry (AREA)
- Proteomics, Peptides & Aminoacids (AREA)
- Physics & Mathematics (AREA)
- General Health & Medical Sciences (AREA)
- Urology & Nephrology (AREA)
- Microbiology (AREA)
- Pathology (AREA)
- Genetics & Genomics (AREA)
- Wood Science & Technology (AREA)
- Zoology (AREA)
- Biochemistry (AREA)
- Hematology (AREA)
- Biomedical Technology (AREA)
- Biotechnology (AREA)
- General Engineering & Computer Science (AREA)
- Cell Biology (AREA)
- Bioinformatics & Cheminformatics (AREA)
- Biophysics (AREA)
- Food Science & Technology (AREA)
- Medicinal Chemistry (AREA)
- General Physics & Mathematics (AREA)
- Measuring Or Testing Involving Enzymes Or Micro-Organisms (AREA)
- Investigating Or Analysing Biological Materials (AREA)
Abstract
Description
技术领域technical field
本发明涉及生物检测技术领域,涉及ARG1作为脓毒症生物标志物的应用,具体涉及ARG1在制备脓毒症诊断、严重程度判断或预后评估试剂或试剂盒中的应用。The invention relates to the technical field of biological detection, relates to the application of ARG1 as a sepsis biomarker, and particularly relates to the application of ARG1 in the preparation of a sepsis diagnosis, severity judgment or prognosis assessment reagent or kit.
背景技术Background technique
脓毒症在临床上非常常见,但是其机制极其复杂。脓毒症随着宿主对病原体感染的反应失调而发展,并进一步导致肺脏、肝脏、肾脏等急性多器官衰竭以及患者死亡。脓毒症的发病率非常高,在2006年~2016年,全球累计约有3150万脓毒症病例,以及1940万严重脓毒症病例(C Fleischmann,A Scherag and N K Adhikari,et al.Assessment ofGlobal Incidence and Mortality of Hospital-Treated Sepsis.)。脓毒症的死亡率也非常高,其30天平均死亡率为24.4%,90天的平均死亡率为32.2%,是全球最主要的死亡原因之一(M Bauer,H Gerlach and T Vogelmann,et al.Mortality in Sepsis and SepticShock in Europe,North America and Australia Between 2009 and 2019-Resultsfrom a Systematic Review and Meta-Analysis.Crit Care,2020,24(1):p.239.)。所以,脓毒症是一个非常关键重要的公共卫生问题,它极大的影响着住院患者尤其是重症监护室内患者的生存质量以及生存期。Sepsis is very common clinically, but its mechanism is extremely complex. Sepsis develops as a dysregulated host response to pathogen infection and further leads to acute multiorgan failure of the lungs, liver, kidneys, etc., as well as patient death. The incidence of sepsis is very high. From 2006 to 2016, there were about 31.5 million cases of sepsis and 19.4 million cases of severe sepsis in the world (C Fleischmann, A Scherag and N K Adhikari, et al. Assessment of Global Incidence and Mortality of Hospital-Treated Sepsis.). Sepsis also has a very high mortality rate, with an average 30-day mortality rate of 24.4% and a 90-day average mortality rate of 32.2%, making it one of the leading causes of death worldwide (M Bauer, H Gerlach and T Vogelmann, et al. al. Mortality in Sepsis and SepticShock in Europe, North America and Australia Between 2009 and 2019-Results from a Systematic Review and Meta-Analysis. Crit Care, 2020, 24(1):p.239.). Therefore, sepsis is a very critical and important public health problem, which greatly affects the quality of life and survival of hospitalized patients, especially those in intensive care units.
生物标志物是指标记系统、器官、组织、细胞及亚细胞结构或功能的改变的生化指标,可用于疾病诊断以及判断疾病分期等方面。分子层面的生物标志物可以更快速、更好地对脓毒症患者进行分层,从而有助于脓毒症患者得到及时的治疗(S Gibot,M C Bene andR Noel,et al.Combination Biomarkers to Diagnose Sepsis in the Critically IllPatient.Am J Respir Crit Care Med,2012,186(1):p.65-71.)。Biomarkers refer to biochemical indicators that mark changes in system, organ, tissue, cell and subcellular structure or function, and can be used for disease diagnosis and disease staging. Molecular-level biomarkers can allow for faster and better stratification of patients with sepsis, thereby helping patients with sepsis receive timely treatment (S Gibot, M C Bene and R Noel, et al. Combination Biomarkers to Diagnose Sepsis in the Critically IllPatient. Am J Respir Crit Care Med, 2012, 186(1):p.65-71.).
生物信息学的出现和发展为寻找脓毒症生物标志物提供了新的途径。最近几十年基因芯片(DNAMicroarray)以及RNA高通量测序(RNA-Sequencing)等技术的不断发展,使生物信息学不断发展出新的领域,并产生了海量的数据(Z Chen,Y Lin and J Gao,etal.Identification of Key Candidate Genes for Colorectal Cancer byBioinformatics Analysis.Oncol Lett,2019,18(6):p.6583-6593.)。这些数据促使了基因加权共表达网络分析(Weighted Correlation Network Analysis,WGCNA)、构建蛋白-蛋白相互作用网络(Protein-Protein Interaction Network,PPI Network)模型以及构建mRNA-miRNA网络等方法的出现。这些方法为寻找生物标志物提供了非常有价值的线索。例如,中国专利申请CN202011059286.2,发明名称为“一种利用生物信息学筛选疾病标志物的方法及其应用”,申请公布号:CN114333991A,公开了一种在瘢痕疙瘩等疾病上,利用生物信息学筛选生物标志物的方法及其应用。The emergence and development of bioinformatics provides new avenues for finding biomarkers of sepsis. The continuous development of technologies such as DNA Microarray and RNA-Sequencing in recent decades has led to the continuous development of new fields in bioinformatics and generated massive amounts of data (Z Chen, Y Lin and J Gao, et al. Identification of Key Candidate Genes for Colorectal Cancer by Bioinformatics Analysis. Oncol Lett, 2019, 18(6): p. 6583-6593.). These data prompted the emergence of methods such as Weighted Correlation Network Analysis (WGCNA), constructing Protein-Protein Interaction Network (PPI Network) models, and constructing mRNA-miRNA networks. These methods provide very valuable clues for finding biomarkers. For example, Chinese patent application CN202011059286.2, the name of the invention is "a method for screening disease markers using bioinformatics and its application", application publication number: CN114333991A, discloses a kind of keloid and other diseases, using biological information Methods for screening biomarkers and their applications.
ARG1基因是编码精氨酸酶1(Arginase 1)的基因,该基因所编码的蛋白催化精氨酸水解为鸟氨酸和尿素。免疫功能紊乱是脓毒症重要的特征之一,而精氨酸酶相关的代谢是先天性和适应性免疫反应的关键调节剂(Markus Munder,Faustino Mollinedo,JeroCalafat et al.Arginase I is constitutively expressed in human granulocytesand participates in fungicidal activity.Blood.2005 Mar 15;105(6):2549-56.),这提示了ARG1可能与脓毒症相关。人ARG1基因的Ensembl ID为ENSG00000118520,其蛋白的三种亚型的氨基酸序列见NP_001231367.1、NP_000036.2和NP_001355949.1。The ARG1 gene is a gene encoding arginase 1 (Arginase 1), and the protein encoded by this gene catalyzes the hydrolysis of arginine into ornithine and urea. Immune dysfunction is one of the important features of sepsis, and arginase-related metabolism is a key regulator of innate and adaptive immune responses (Markus Munder, Faustino Mollinedo, JeroCalafat et al. Arginase I is constitutively expressed in human granulocytes and participates in fungicidal activity. Blood. 2005 Mar 15; 105(6): 2549-56.), suggesting that ARG1 may be associated with sepsis. The Ensembl ID of the human ARG1 gene is ENSG00000118520, and the amino acid sequences of the three isoforms of the protein are shown in NP_001231367.1, NP_000036.2 and NP_001355949.1.
目前国内外尚无有关ARG1作为脓毒症生物标志物方面的文献报道。At present, there is no literature report on ARG1 as a sepsis biomarker at home and abroad.
发明内容SUMMARY OF THE INVENTION
本发明依托上述研究进行,目的在于提供脓毒症的诊断及预后标志物,本发明的目的也在于提供ARG1的新用途,即在制备脓毒症诊断、严重程度判断或预后评估试剂盒中的应用。The present invention relies on the above-mentioned research to provide diagnostic and prognostic markers for sepsis, and the present invention also aims to provide new uses of ARG1, that is, in the preparation of sepsis diagnosis, severity judgment or prognosis assessment kits application.
为了寻找脓毒症患者外周血中特征性的基因作为生物标志物。本发明通过下载并分析脓毒症患者和健康对照的数据集,进行差异表达基因的筛选,然后对差异表达基因分别使用PPI网络分析以及WGCNA分析,将两种分析结果取交集,得到了ARG1。然后分析ARG1在脓毒症精确诊断、严重程度判断以及预后预测等方面的价值,从而发现ARG1适合做脓毒症诊断和治疗的生物标志物。In order to find the characteristic genes in the peripheral blood of sepsis patients as biomarkers. In the present invention, the data sets of sepsis patients and healthy controls are downloaded and analyzed, and the differentially expressed genes are screened, and then PPI network analysis and WGCNA analysis are respectively used for the differentially expressed genes, and the two analysis results are obtained by intersecting the two analysis results to obtain ARG1. Then, the value of ARG1 in the precise diagnosis, severity judgment and prognosis prediction of sepsis was analyzed, and it was found that ARG1 was suitable as a biomarker for the diagnosis and treatment of sepsis.
具体地,本发明提供了如下技术方案:Specifically, the present invention provides the following technical solutions:
本发明的第一方面,提供了一种寻找生物标志物的方法,具体包括以下步骤:A first aspect of the present invention provides a method for finding biomarkers, which specifically includes the following steps:
(1)从Gene Expression Omnibus数据库(在线地址为:https://www.ncbi.nlm.nih.gov/geo/)中下载脓毒症患者和健康对照的基因表达谱数据。(1) Download the gene expression profiling data of sepsis patients and healthy controls from the Gene Expression Omnibus database (online address: https://www.ncbi.nlm.nih.gov/geo/).
(2)将这些表达数据使用limmaR包进行基因的差异表达分析。(2) The differential expression analysis of genes was performed using the limmaR package for these expression data.
(3)根据每个差异表达基因的表达谱,使用基因加权共表达网络分析(Weightedcorrelation network analysis,WGCNA)算法,鉴定出显著性程度较高(即:与临床特征相关性较为密切)的基因。(3) According to the expression profile of each differentially expressed gene, the weighted co-expression network analysis (WGCNA) algorithm was used to identify genes with a high degree of significance (ie, closely related to clinical characteristics).
(4)构建蛋白-蛋白相互作用网络(Protein-Protein Interaction Network)模型,在差异表达基因所对应的蛋白中,筛选出与其他蛋白存在较多相互作用的蛋白,其基因被称为关键基因。(4) Constructing a protein-protein interaction network model. Among the proteins corresponding to the differentially expressed genes, the proteins that have more interactions with other proteins are screened out, and their genes are called key genes.
(5)将步骤(4)中PPI网络筛选出的基因,与步骤(3)中WGCNA的鉴定出的显著性较高的基因取交集,得到关键的生物标志物。(5) The genes screened by the PPI network in step (4) are intersected with the genes with higher significance identified by WGCNA in step (3) to obtain key biomarkers.
本发明的第二方面,提供了一种考察脓毒症外周血生物标志物性能的方案,具体包括以下步骤:The second aspect of the present invention provides a scheme for investigating the performance of peripheral blood biomarkers of sepsis, which specifically includes the following steps:
(1)考察该生物标志物在脓毒症患者外周血中的表达水平是否要显著高于健康对照组。(1) To investigate whether the expression level of this biomarker in peripheral blood of patients with sepsis is significantly higher than that of healthy controls.
(2)考察该生物标志物在严重脓毒症或致死性脓毒症患者外周血中的表达水平要是否显著高于普通的脓毒症患者。(2) To investigate whether the expression level of this biomarker in the peripheral blood of patients with severe sepsis or fatal sepsis is significantly higher than that of ordinary patients with sepsis.
(3)考察该生物标志物在脓毒性休克患者外周血中的表达水平要显著高于其他类型休克的患者。(3) The expression level of this biomarker in peripheral blood of patients with septic shock was significantly higher than that of patients with other types of shock.
(4)考察该生物标志物在在对后续支持性治疗无响应的脓毒症患者外周血中的表达水平是否要显著高于有相应患者。(4) To investigate whether the expression level of this biomarker in the peripheral blood of sepsis patients who did not respond to follow-up supportive treatment was significantly higher than that of the corresponding patients.
(5)考察该生物标志物在脓毒症造模小鼠外周血中的表达是否要显著高于假手术组小鼠。(5) To investigate whether the expression of this biomarker in the peripheral blood of the sepsis model mice is significantly higher than that of the sham-operated mice.
本发明的第三方面,提供了ARG1作为脓毒症标志物的应用,具体是在精确诊断、严重程度判断以及预后评估等方面作为生物标志物的应用,更具体是在制备脓毒症诊断、严重程度判断或预后评估试剂或试剂盒中的应用。The third aspect of the present invention provides the application of ARG1 as a marker of sepsis, specifically the application of ARG1 as a biomarker in the aspects of accurate diagnosis, severity judgment and prognosis evaluation, more specifically in the preparation of sepsis diagnosis, Severity judgment or the use of prognostic assessment reagents or kits.
在本发明中,所述的ARG1指的是一个基因及其对应的蛋白。该基因位于人第6号染色体上,具体位置为6q23.2,外显子数为8,Ensembl ID为ENSG00000118520。其对应的蛋白三种亚型的氨基酸序列见NP_001231367.1、NP_000036.2和NP_001355949.1。In the present invention, the ARG1 refers to a gene and its corresponding protein. The gene is located on
在本发明中,所述的脓毒症是由于宿主对感染的反应失调而导致的危及生命的多器官功能障碍疾病。所述的生物标志物是指可以标记系统、器官、组织、细胞及亚细胞结构或功能的改变或可能发生的改变的生化指标。In the present invention, sepsis is a life-threatening multiple organ dysfunction disease caused by a dysregulated host response to infection. The biomarkers refer to biochemical indicators that can mark changes in system, organ, tissue, cell and subcellular structure or function or possible changes.
优选的,上述严重程度判断试剂或试剂盒为判断是否为严重或致死的脓毒症的试剂或试剂盒,或者为进一步判断是否为脓毒性休克的试剂或试剂盒。Preferably, the above-mentioned severity judging reagent or kit is a reagent or kit for judging whether it is severe or fatal sepsis, or a reagent or kit for further judging whether it is septic shock.
实验结果显示,相较于健康人群,ARG1在脓毒症患者外周血中显著高表达;在严重或致死的脓毒症患者外周血中转录丰度显著高于普通脓毒症患者,通过ARG1基因的表达量有助于判断脓毒症病情的严重程度;在脓毒性休克(一种严重的脓毒症)中的表达水平显著高于非脓毒性休克。鉴于脓毒性休克与非脓毒性休克具有相似的症状,在临床实践中,ARG1作为生物标志物对这两类病情的区分具有重要价值。The experimental results showed that compared with healthy people, ARG1 was significantly expressed in the peripheral blood of patients with sepsis; the transcriptional abundance in the peripheral blood of patients with severe or fatal sepsis was significantly higher than that of patients with ordinary sepsis. The expression level of s is helpful to judge the severity of sepsis; the expression level in septic shock (a severe form of sepsis) is significantly higher than that in non-septic shock. Given that septic shock and non-septic shock have similar symptoms, in clinical practice, ARG1 is of great value as a biomarker to differentiate these two conditions.
优选的,诊断或预后评估试剂为检测生物样品中ARG1表达量的试剂;所述试剂盒包含了检测生物样品中ARG1含量的试剂。Preferably, the diagnostic or prognostic assessment reagent is a reagent for detecting ARG1 expression in a biological sample; the kit includes a reagent for detecting ARG1 content in a biological sample.
进一步,所述的检测生物样品中ARG1含量的试剂选自:对ARG1基因具有检测特异性的PCR引物,或者与ARG1蛋白特异性结合的抗体。其中,对ARG1基因具有检测特异性的PCR引物如SEQ ID NO.1~4所示。Further, the reagent for detecting ARG1 content in the biological sample is selected from: PCR primers with detection specificity for ARG1 gene, or antibodies that specifically bind to ARG1 protein. Among them, PCR primers with detection specificity for ARG1 gene are shown in SEQ ID NO. 1-4.
ARG1-F引物:TCACCTGAGCTTTGATGTCGA(SEQ ID NO.1);ARG1-F primer: TCACCTGAGCTTTGATGTCGA (SEQ ID NO. 1);
ARG1-R引物:TGAAAGGAGCCCTGTCTTGTA(SEQ ID NO.2);ARG1-R primer: TGAAAGGAGCCCTGTCTTGTA (SEQ ID NO. 2);
GAPDH-F引物:TCACCATCTTCCAGGAGCGAGAC;(SEQ ID NO.3);GAPDH-F primer: TCACCATCTTCCAGGAGCGAGAC; (SEQ ID NO. 3);
GAPDH-R引物:AGACACCAGTAGACTCCACGACATAC(SEQ ID NO.4)。GAPDH-R primer: AGACACCAGTAGACTCCACGACATAC (SEQ ID NO. 4).
进一步,所述的生物样品获自对象的外周血。Further, the biological sample is obtained from the peripheral blood of the subject.
本发明的第四方面,提供了一种脓毒症诊断、严重程度判断或预后评估试剂盒,该试剂盒包含了检测生物样品中ARG1含量的试剂。A fourth aspect of the present invention provides a sepsis diagnosis, severity judgment or prognosis assessment kit, which contains a reagent for detecting ARG1 content in a biological sample.
优选的,该试剂盒是由逆转录系统、引物系统和扩增系统组成,所述的引物系统包括如SEQ ID NO.1~4所示的PCR引物。Preferably, the kit is composed of a reverse transcription system, a primer system and an amplification system, and the primer system includes PCR primers as shown in SEQ ID NO. 1-4.
本发明的第五方面,提供了一种利用上述试剂盒进行脓毒症诊断、严重程度判断或预后评估的方法,如下:A fifth aspect of the present invention provides a method for diagnosing sepsis, judging the severity or evaluating the prognosis using the above-mentioned kit, as follows:
A.将待检血液样本常温离心,获取血浆,提取血浆中的总RNA并测定血浆纯度;A. Centrifuge the blood sample to be tested at room temperature to obtain plasma, extract the total RNA in the plasma and determine the plasma purity;
B.对步骤A中的总RNA进行逆转录,得到cDNA;B. The total RNA in step A is reverse transcribed to obtain cDNA;
C.采用实时荧光定量PCR对ARG1的拷贝数进行定量检测,检测过程中所用引物如SEQ ID NO.1~4所示。C. Quantitatively detect the copy number of ARG1 by real-time fluorescence quantitative PCR, and the primers used in the detection process are shown in SEQ ID NO. 1-4.
数据均采用SPSS16.0处理,采用平均值±标准差的方式表示。All data were processed by SPSS 16.0 and expressed as mean ± standard deviation.
相比于现有技术,本发明的有益效果如下:Compared with the prior art, the beneficial effects of the present invention are as follows:
首先,就方法而言,本发明提供了一种生物标志物的鉴定方法,能够利用公开、免费的数据集,通过生物信息学分析,快速寻找到生物标志物。同时,本发明提供了一种考察脓毒症外周血生物标志物性能的方案,能够利用生物信息学数据集和实验小鼠,较为快速地考察和验证一个生物标志物在脓毒症精确诊断、严重程度判断以及预后预测等方面的性能。此外,本发明发现ARG1可以作为脓毒症的一个“多功能”生物标志物,这有利于脓毒症患者病情的精确诊断以及预后的快速预测,从而有利于脓毒症患者的快速分层和及时治疗。First, in terms of methods, the present invention provides a method for identifying biomarkers, which can quickly find biomarkers through bioinformatics analysis using open and free data sets. At the same time, the present invention provides a scheme for examining the performance of biomarkers in peripheral blood of sepsis, which can use bioinformatics data sets and experimental mice to relatively quickly examine and verify the use of a biomarker in accurate diagnosis of sepsis, Severity judgment and prognosis prediction. In addition, the present invention finds that ARG1 can be used as a "multifunctional" biomarker of sepsis, which is conducive to the accurate diagnosis of the condition of patients with sepsis and the rapid prediction of prognosis, thereby facilitating the rapid stratification and rapid prediction of patients with sepsis. Treat in time.
其次,就技术而言,ARG1的检测本质上是一种血液基因组的定量PCR检测,具有操作简便、检测灵敏、特异性好、重复性高等特点,现今已越来越多地被应用于临床检验技术中。本发明所采用基本检测方法是定量PCR,这一技术在现代实验诊断学中已被证实是高灵敏度、高准确度的检测方法,试验技术已经十分成熟,并且我们采用的是这一技术中的标准曲线定量法,可以准确地对各种样品中核酸分子做精确定量。Secondly, in terms of technology, the detection of ARG1 is essentially a quantitative PCR detection of blood genome, which has the characteristics of simple operation, sensitive detection, good specificity and high repeatability, and has been increasingly used in clinical testing. in technology. The basic detection method adopted in the present invention is quantitative PCR, which has been proved to be a high-sensitivity and high-accuracy detection method in modern laboratory diagnostics. The standard curve quantification method can accurately quantify nucleic acid molecules in various samples.
第三,就效果而言,本发明所涉及的指标ARG1在脓毒症患者外周血中显著高表达,在严重或致死的脓毒症患者外周血中转录丰度显著高于普通脓毒症患者,在脓毒性休克中的表达水平显著高于非脓毒性休克,且差异具有统计学意义(P<0.05),其临床参考价值和可信度较高。不仅可以通过ARG1基因的表达量有助于判断脓毒症病情的严重程度,而且对于症状相似的脓毒性休克与非脓毒性休克的区分具有重要价值。Third, in terms of effect, the index ARG1 involved in the present invention is significantly highly expressed in the peripheral blood of patients with sepsis, and the transcriptional abundance in the peripheral blood of patients with severe or fatal sepsis is significantly higher than that in patients with ordinary sepsis , the expression level in septic shock was significantly higher than that in non-septic shock, and the difference was statistically significant (P<0.05), and its clinical reference value and reliability were high. The expression of ARG1 gene can not only help to judge the severity of sepsis, but also has important value in distinguishing septic shock and non-septic shock with similar symptoms.
第四,就检测方式而言,只需要采集检测者的血液即可获得检测结果,操作简单、无创,患者的接受程度高。Fourth, as far as the detection method is concerned, the test results can be obtained only by collecting the blood of the tester. The operation is simple, non-invasive and highly acceptable to patients.
附图说明Description of drawings
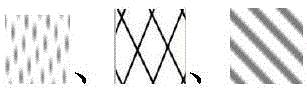
图1是使用WGCNA算法,根据基因的表达模式,将脓毒症患者外周血中的差异表达基因划分为4个模块(基因群),分别以纹理以及表示,灰色是没能划分进任何模块的散在基因。Figure 1 uses the WGCNA algorithm to divide the differentially expressed genes in the peripheral blood of patients with sepsis into 4 modules (gene groups) according to their gene expression patterns. as well as Indicates that the gray are scattered genes that could not be classified into any modules.
图2是每个基因模块与临床诊断(脓毒症或对照)的相关系数,其中,相关系数在方格中展示,相关系数的绝对值越大,则模块颜色越深。除了模块1(Module 1)是负相关外,其余都是正相关。Figure 2 is the correlation coefficient between each gene module and clinical diagnosis (sepsis or control), wherein the correlation coefficient is displayed in the square, and the greater the absolute value of the correlation coefficient, the darker the color of the module. Except for
图3是脓毒症差异表达基因所对应的蛋白构成的蛋白-蛋白相互作用网络,其中深灰色代表上调基因,浅灰色代表下调基因。Figure 3 is a protein-protein interaction network composed of proteins corresponding to differentially expressed genes in sepsis, in which dark gray represents up-regulated genes and light gray represents down-regulated genes.
图4是寻找PPI网络筛选与WGCNA分析结果中的共有基因。结果显示,ARG1是两个结果中唯一的共有基因。Figure 4 is to find common genes in the results of PPI network screening and WGCNA analysis. The results showed that ARG1 was the only shared gene in both results.
图5显示了ARG1在脓毒症患者外周血中高表达:分别使用了GSE95233、GSE134347、GSE154918、GSE13015、GSE60424、GSE131761这六个成人数据集以及GSE8121、GSE26378、GSE26440、GSE145227这四个儿童数据集来说明ARG1在脓毒症患者外周血中的高表达。每个数据集对应的箱式图显示无论是成人患者还是儿童患者,ARG1在外周血中都出现了显著的高表达。每个箱式图下方都有ROC曲线,ROC曲线的曲线下面积也等于或接近于1,证明ARG1具有较为良好的灵敏度和特异性,有作为生物标志物的潜力。***表示脓毒症组和对照组相比较,有统计学差异(P<0.001)。Figure 5 shows that ARG1 is highly expressed in the peripheral blood of patients with sepsis: six adult datasets GSE95233, GSE134347, GSE154918, GSE13015, GSE60424, GSE131761 and four pediatric datasets GSE8121, GSE26378, GSE26440, GSE145227 were used, respectively. This indicates that ARG1 is highly expressed in the peripheral blood of patients with sepsis. The boxplots corresponding to each dataset show that ARG1 is significantly overexpressed in peripheral blood in both adult and pediatric patients. There is a ROC curve below each box plot, and the area under the curve of the ROC curve is also equal to or close to 1, which proves that ARG1 has relatively good sensitivity and specificity, and has the potential as a biomarker. *** indicates that there is a statistical difference between the sepsis group and the control group (P<0.001).
图6显示了ARG1在脓毒性休克中的表达水平要显著高于非脓毒性休克:分别使用GSE131411和GSE131761两个数据集通过绘制箱式图证实了ARG1在脓毒性休克(一种严重的脓毒症)中的表达水平要显著高于非脓毒性休克。鉴于脓毒性休克与非脓毒性休克具有相似的症状,在临床实践中,ARG1作为生物标志物对这两类病情的区分具有重要价值。***表示脓毒性休克组和非脓毒性休克组相比较,有统计学差异(P<0.001);*表示脓毒性休克组和非脓毒性休克组相比较,有统计学差异(P<0.05)。Figure 6 shows that the expression level of ARG1 in septic shock is significantly higher than that in non-septic shock: using two datasets GSE131411 and GSE131761, respectively Symptoms) were significantly higher than those in non-septic shock. Given that septic shock and non-septic shock have similar symptoms, in clinical practice, ARG1 is of great value as a biomarker to differentiate these two conditions. *** means there is a statistical difference between the septic shock group and the non-septic shock group (P<0.001); * means there is a statistical difference between the septic shock group and the non-septic shock group (P<0.05) ).
图7显示了ARG1基因在严重或致死的脓毒症患者外周血中转录丰度要显著高于普通脓毒症患者:分别使用了GSE63042和GSE154918两个数据集通过绘制箱式图证实了ARG1基因在严重或致死的脓毒症患者外周血中转录丰度要显著高于普通脓毒症患者。这说明,ARG1基因的表达量有助于判断脓毒症病情的严重程度。***表示普通脓毒症组和脓毒性休克组相比较,有统计学差异(P<0.001);*表示普通脓毒症和严重或致死性脓毒症组相比较,有统计学差异(P<0.05)。Figure 7 shows that the transcriptional abundance of ARG1 gene in the peripheral blood of patients with severe or lethal sepsis is significantly higher than that of patients with common sepsis: using two datasets GSE63042 and GSE154918, respectively, the ARG1 gene was confirmed by drawing box plots The transcript abundance in peripheral blood of severe or fatal sepsis patients was significantly higher than that of common sepsis patients. This shows that the expression of ARG1 gene is helpful to judge the severity of sepsis. *** indicates that there is a statistical difference between the common sepsis group and the septic shock group (P<0.001); * indicates that there is a statistical difference between the common sepsis group and the severe or fatal sepsis group ( P<0.05).
图8显示了ARG1在对后续治疗无响应的脓毒症患者外周血中显著高于有响应的患者:使用GSE110487这个数据集,通过绘制箱式图证实了ARG1在对后续治疗无响应的脓毒症患者外周血中显著高于有响应的患者,说明ARG1的表达水平有助于预测脓毒症患者对后续的支持性治疗是否会有响应,体现出其作为生物标志物的良好性能。**表示对后续治疗无响应的脓毒症患者和对后续治疗有响应的脓毒症患者相比较,有统计学差异(P<0.01)。Figure 8 shows that ARG1 is significantly higher in the peripheral blood of septic patients who do not respond to subsequent treatment than in patients who do respond: Using the GSE110487 dataset, it was confirmed that ARG1 was not involved in septic patients who did not respond to subsequent treatment by drawing a box plot. The peripheral blood of patients with sepsis was significantly higher than that of patients who responded, indicating that the expression level of ARG1 can help predict whether patients with sepsis will respond to subsequent supportive treatment, reflecting its good performance as a biomarker. ** Indicates that there is a statistical difference between sepsis patients who do not respond to subsequent treatment and sepsis patients who respond to subsequent treatment (P<0.01).
图9是使用实时荧光定量PCR的方法,从实验层面确证了ARG1在脓毒症小鼠外周血中的表达水平要显著高于假手术组小鼠。***表示对脓毒症小鼠和假手术组小鼠相比较,有统计学差异(P<0.01)。Figure 9 shows the method using real-time fluorescence quantitative PCR, which confirms from the experimental level that the expression level of ARG1 in the peripheral blood of septic mice is significantly higher than that of sham-operated mice. *** indicates that there is a statistical difference between the septic mice and the sham-operated mice (P<0.01).
具体实施方式Detailed ways
现结合实施例和附图,对本发明作详细描述,但本发明的实施不仅限于此。The present invention will now be described in detail with reference to the embodiments and the accompanying drawings, but the implementation of the present invention is not limited thereto.
本发明所用C57BL/6小鼠购自上海斯莱克实验动物有限公司。The C57BL/6 mice used in the present invention were purchased from Shanghai Slack Laboratory Animal Co., Ltd.
本发明所用试剂和原料均市售可得或可按文献方法制备。下列实施例中未注明具体条件的分析方法和实验方法,通常按照常规条件,或按照制造厂商所建议的条件。The reagents and raw materials used in the present invention are commercially available or can be prepared according to literature methods. In the following examples, the analytical methods and experimental methods without specific conditions are usually in accordance with conventional conditions, or in accordance with the conditions suggested by the manufacturer.
实施例1:脓毒症患者外周血差异表达基因的筛选Example 1: Screening of differentially expressed genes in peripheral blood of patients with sepsis
本发明在GEO数据库(在线地址:https://www.ncbi.nlm.nih.gov/geo/)中,找到了四个基因表达谱数据集,编号分别是GSE28750、GSE57065、GSE65682、GSE69528,每个数据集内都包含脓毒症患者外周血样本(实验组)以及健康人外周血样本(对照组)。下载每个数据集的基因表达谱。The present invention has found four gene expression profile data sets in the GEO database (online address: https://www.ncbi.nlm.nih.gov/geo/), the numbers are GSE28750, GSE57065, GSE65682, GSE69528, each Each dataset contains peripheral blood samples from patients with sepsis (experimental group) and peripheral blood samples from healthy people (control group). Download gene expression profiles for each dataset.
在Microsoft Windows 1064位操作系统上,从R语言主页(在线地址为:www.r-project.org)下载R 4.1.2版本后,按照提示进行默认安装。在集成开发环境RStudio主页(在线地址为:www.rstudio.com/)下载RStudio 1.3.1073版本后,按照默认提示进行安装。使用BiocManager::install(“limma”)命令安装limmaR模块。然后使用该模块分别对上述四个基因表达谱数据集进行差异表达分析,以log Fold Change(FC)>1或<-1且adjustedPvalue<0.05为筛选标准,获得每个数据集的上调和下调的差异表达基因。On Microsoft Windows 1064-bit operating system, after downloading the R 4.1.2 version from the R language homepage (online address: www.r-project.org), follow the prompts for default installation. After downloading the RStudio 1.3.1073 version from the RStudio homepage of the integrated development environment (online address: www.rstudio.com/), follow the default prompts to install it. Install the limmaR module using the BiocManager::install("limma") command. Then use this module to perform differential expression analysis on the above four gene expression profiling datasets respectively, with log Fold Change (FC)>1 or <-1 and adjustedPvalue<0.05 as the screening criteria to obtain the up-regulated and down-regulated values of each dataset. differentially expressed genes.
使用在线工具Venny 2.1.0(在线地址:https://bioinfogp.cnb.csic.es/tools/venny/index.ht ml)生成韦恩图,以查找四个数据集中共有的上调和下调差异表达基因,差异表达基因的详细列表如表1所示:Venn diagrams were generated using the online tool Venny 2.1.0 (online address: https://bioinfogp.cnb.csic.es/tools/venny/index.html) to find up- and down-regulated differential expressions common to the four datasets Genes, a detailed list of differentially expressed genes is shown in Table 1:
表1 脓毒症上调和下调差异表达基因列表Table 1 List of up-regulated and down-regulated differentially expressed genes in sepsis
实施例2:脓毒症患者差异表达基因的基因加权共表达网络分析Example 2: Gene-weighted co-expression network analysis of differentially expressed genes in patients with sepsis
在RStudio集成开发环境中,使用BiocManager::install("WGCNA")命令安装WGCNA的R包。对于每个数据集,使用goodSamplesGenesMS函数移除样本数量缺失较多的基因以及基因数量确实较多的样本。接着,使用hclust函数对样本进行聚类分析,并移除过于离群的脓毒症或健康对照样本。下一步,使用pickSoftThreshold函数和softConnectivity函数调整软阈值,使后续网络中基因与基因之间的连接近似服从无尺度分布,从而使网络更具有生物学意义。然后生成基因表达相关性矩阵,再转化为邻接矩阵,再进一步转化为拓扑重叠矩阵而得到共表达网络。然后进行基因的层次聚类分析,使用动态切割树算法,将表达模式相近的基因划分到同一个模块,计算每个模块与临床诊断(脓毒症或健康对照)之间的皮尔逊相关系数(Pearson correlation coefficient)。最后,计算在最相关的模块中,每个基因的表达模式与临床诊断之间的皮尔逊相关系数作为基因的显著性程度(GeneSignificance,GS)。In the RStudio IDE, use the BiocManager::install("WGCNA") command to install the WGCNA R package. For each dataset, use the goodSamplesGenesMS function to remove genes with a large number of missing samples and samples with a really large number of genes. Next, the samples were clustered using the hclust function and the sepsis or healthy control samples that were too outliers were removed. Next, use the pickSoftThreshold function and the softConnectivity function to adjust the soft threshold, so that the connections between genes in the subsequent network approximately obey a scale-free distribution, thus making the network more biologically meaningful. Then a gene expression correlation matrix is generated, converted into an adjacency matrix, and further converted into a topological overlap matrix to obtain a co-expression network. Then perform hierarchical clustering analysis of genes, use dynamic cutting tree algorithm, divide genes with similar expression patterns into the same module, and calculate the Pearson correlation coefficient between each module and clinical diagnosis (sepsis or healthy controls) ( Pearson correlation coefficient). Finally, the Pearson correlation coefficient between the expression pattern of each gene and the clinical diagnosis in the most relevant module was calculated as the degree of significance (GeneSignificance, GS) of the gene.
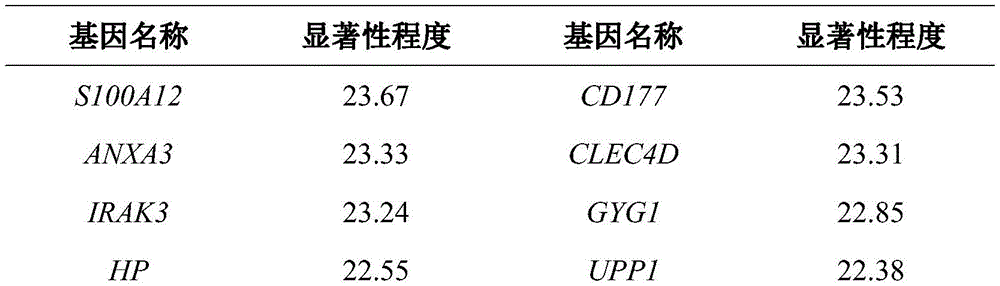
基因的模块划分如图1所示,每个模块与临床特征的皮尔逊相关系数如图2所示。显著性程度较高的基因如表2所示。The module division of genes is shown in Figure 1, and the Pearson correlation coefficient between each module and clinical characteristics is shown in Figure 2. The genes with a higher degree of significance are shown in Table 2.
表2 使用WGCNA分析所得到的与临床特征关系最密切(显著性程度最高)的15个基因Table 2 The 15 genes most closely related to clinical characteristics (highest degree of significance) obtained by WGCNA analysis
实施例3:脓毒症患者差异表达基因的蛋白-蛋白相互作用网络的构建Example 3: Construction of protein-protein interaction network of differentially expressed genes in sepsis patients
打开STRING数据库(在线地址:https://string-db.org/),将上述筛选出的脓毒症患者的差异表达基因列表导入至该数据库中,构建蛋白-蛋白相互作用网络(Protein-Protein Interaction Network,PPI Network)。使用Cytoscape软件(版本:3.8.2)将该网络可视化。然后使用Cytoscape中的MCODE插件识别与其他蛋白存在较多相互作用的蛋白。这些蛋白更有可能处在脓毒症致病过程中的核心位置,因而其对应的基因更加关键。Open the STRING database (online address: https://string-db.org/), import the differentially expressed gene list of the patients with sepsis screened above into the database, and construct a protein-protein interaction network (Protein-Protein Interaction Network). Interaction Network, PPI Network). The network was visualized using Cytoscape software (version: 3.8.2). Proteins with more interactions with other proteins were then identified using the MCODE plugin in Cytoscape. These proteins are more likely to be at the core of the pathogenesis of sepsis, so their corresponding genes are more critical.
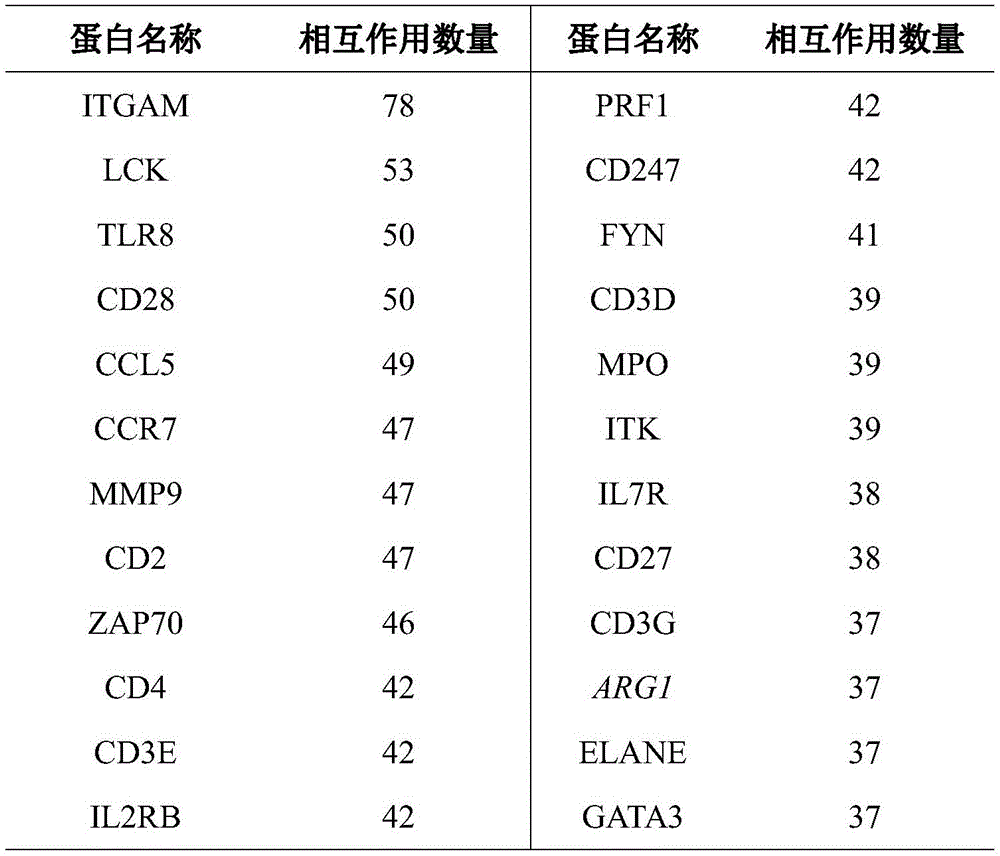
蛋白-蛋白相互作用网络如图3所示,与其他蛋白存在较多相互作用的蛋白其对应的基因名称列表如表3所示。The protein-protein interaction network is shown in Figure 3, and the corresponding gene names of proteins that have more interactions with other proteins are shown in Table 3.
表3 PPI网络模型鉴定出来的相互作用较多的蛋白汇总Table 3 Summary of proteins with more interactions identified by the PPI network model
实施例4:PPI网络筛选与WGCNA分析结果中共有基因的寻找Example 4: Search for common genes in PPI network screening and WGCNA analysis results
使用在线工具Venny 2.1.0(在线地址:https://bioinfogp.cnb.csic.es/tools/venny/index.ht ml)生成韦恩图,以查找在PPI网络筛选所得蛋白其对应的基因,与WGCNA分析所得的基因中,是否存在交集。Use the online tool Venny 2.1.0 (online address: https://bioinfogp.cnb.csic.es/tools/venny/index.html) to generate a Venn diagram to find the corresponding genes of the proteins screened in the PPI network, Whether there is an intersection with the genes analyzed by WGCNA.
结果如图4所示,表明ARG1基因是两种生物信息学方法所得结果中的共有基因。The results are shown in Figure 4, indicating that the ARG1 gene is a common gene in the results obtained by the two bioinformatics methods.
实施例5:ARG1在区分脓毒症与健康对照方面的性能考察Example 5: Performance of ARG1 in distinguishing sepsis from healthy controls
使用BiocManager::install("ggplot2")命令安装ggplot2的R包。使用该R包来生成数据集中的基因相对表达量的柱状图。Use the BiocManager::install("ggplot2") command to install the R package for ggplot2. Use this R package to generate a histogram of the relative expression of genes in the dataset.
使用BiocManager::install("pROC")命令安装pROC的R包。使用该R包来生成每个数据集中受试者工作(Receiver Operating Characteristic,ROC)曲线图,并计算曲线下面积(Area Under Curve)。Use the BiocManager::install("pROC") command to install the pROC R package. This R package was used to generate Receiver Operating Characteristic (ROC) plots for each dataset and to calculate the Area Under Curve.
上述所使用的数据集包括了成人数据集GSE95233、GSE134347、GSE154918、GSE13015、GSE60424、GSE131761以及儿童数据集GSE8121、GSE26378、GSE26440、GSE145227。每个数据集中都包含了脓毒症患者以及健康人对照或对照组病人。The datasets used above include adult datasets GSE95233, GSE134347, GSE154918, GSE13015, GSE60424, GSE131761, and children's datasets GSE8121, GSE26378, GSE26440, and GSE145227. Patients with sepsis and healthy controls or control patients were included in each dataset.
结果如图5所示,显示ARG1基因在脓毒症患者的外周血中转录丰度都显著高于对照组,其ROC曲线的曲线下面积也等于或接近于1,证明ARG1具有较为良好的灵敏度和特异性,有作为生物标志物的潜力。The results are shown in Figure 5, showing that the ARG1 gene transcriptional abundance in the peripheral blood of sepsis patients was significantly higher than that of the control group, and the area under the curve of the ROC curve was also equal to or close to 1, proving that ARG1 has relatively good sensitivity and specificity, have potential as biomarkers.
实施例6:ARG1在区分脓毒症和症状类似疾病方面的性能考察Example 6: Performance of ARG1 in distinguishing sepsis from symptom-like diseases
GSE131411和GSE131761两个数据集都包含脓毒性休克以及非脓毒性休克患者的外周血样本。本发明继续使用limma R包来生成这两个数据集中的ARG1相对表达量的柱状图。Both datasets GSE131411 and GSE131761 contain peripheral blood samples from septic shock as well as non-septic shock patients. The present invention continues to use the limma R package to generate a histogram of the relative expression levels of ARG1 in these two data sets.
结果如图6所示,显示ARG1基因在脓毒性休克患者外周血中转录丰度显著大于非脓毒性休克。由于脓毒性休克是脓毒症的一种严重形式,并且与非脓毒性休克具有相似的症状,在临床实践中,ARG1作为生物标志物对这两类病情的区分具有重要价值。The results are shown in Fig. 6, showing that the transcriptional abundance of ARG1 gene in peripheral blood of patients with septic shock is significantly greater than that in non-septic shock. Since septic shock is a severe form of sepsis and has similar symptoms to non-septic shock, ARG1 is of great value as a biomarker to differentiate these two conditions in clinical practice.
实施例7:ARG1在区分脓毒症严重程度方面的性能考察Example 7: Performance of ARG1 in differentiating sepsis severity
GSE63042和GSE154918两个数据集都包含普通脓毒症患者以及严重或致死性脓毒症患者的外周血样本。本发明继续使用limmaR包来生成这两个数据集中的ARG1相对表达量的柱状图。Both datasets GSE63042 and GSE154918 contain peripheral blood samples from patients with common sepsis as well as patients with severe or fatal sepsis. The present invention continues to use the limmaR package to generate a histogram of the relative expression levels of ARG1 in these two datasets.
结果如图7所示,显示ARG1基因在严重或致死的脓毒症患者外周血中转录丰度显著大于普通脓毒症患者。这说明,外周血中ARG1的水平有助于判断脓毒症病情的严重程度,从而方便筛选出严重脓毒症的患者给予更具针对性的治疗。The results are shown in Figure 7, showing that the transcriptional abundance of ARG1 gene in the peripheral blood of severe or fatal sepsis patients was significantly greater than that of common sepsis patients. This shows that the level of ARG1 in peripheral blood can help to judge the severity of sepsis, so as to facilitate the screening of patients with severe sepsis and give more targeted treatment.
实施例8:ARG1在预测对脓毒症支持性治疗是否有相应方面的性能考察Example 8: The performance of ARG1 in predicting whether there is a response to supportive treatment of sepsis
GSE110487是唯一能找到的分析对脓毒症支持性治疗是否有相应的数据集。在这个数据集采样时,首先患者在进入重症监护室时接受了血检,同时将其血液样本送去测序。然后接下来的几天记录了每位脓毒症患者对后续支持性治疗是否存在响应。响应者和非响应者在进入ICU接受治疗时,在感染类型、炎症循环标志物或白细胞和淋巴细胞计数等血检结果方面并不存在显著差异(Barcella M,Bollen P B,Braga D,et al.Identificationof a transcriptome profile associated with improvement of organ function inseptic shock patients after early supportive therapy[J].Crit Care,2018,22(1):312.)。而如图8所示,与有响应的脓毒症患者相比,对后续治疗无响应的脓毒症患者全血中ARG1的水平显著较高。这说明脓毒症患者外周血中ARG1的水平有助于预测脓毒症患者对后续的支持性治疗是否会有响应,体现出其作为生物标志物的良好潜质。GSE110487 is the only dataset that could be found to analyze whether there is a corresponding dataset for sepsis supportive care. When this dataset was sampled, patients first underwent blood tests upon admission to the intensive care unit, and their blood samples were sent for sequencing. Each septic patient's response to follow-up supportive care was then recorded over the next few days. Responders and non-responders did not differ significantly in type of infection, inflammatory cycle markers, or blood test results such as white blood cell and lymphocyte counts when admitted to the ICU (Barcella M, Bollen P B, Braga D, et al. Identification of a transcriptome profile associated with improvement of organ function inseptic shock patients after early supportive therapy[J].Crit Care,2018,22(1):312.). However, as shown in Figure 8, ARG1 levels in whole blood were significantly higher in sepsis patients who did not respond to subsequent treatment compared with responsive sepsis patients. This indicates that the level of ARG1 in peripheral blood of patients with sepsis can help predict whether patients with sepsis will respond to subsequent supportive treatment, reflecting its good potential as a biomarker.
实施例9:动物实验确证ARG1在脓毒症外周血中高表达Example 9: Animal experiments confirm that ARG1 is highly expressed in the peripheral blood of sepsis
使用C57BL/6小鼠,实验组通过盲肠结扎穿刺(Cecum Ligation and Puncture,CLP)实验构建脓毒症模型,以假手术(Sham)为对照组。在手术进行24小时后,取小鼠外周血,使用RNAFast200试剂盒(上海飞捷生物技术有限公司)抽提RNA后,使用逆转录酶混合物(翌圣生物科技(上海)股份有限公司)按说明书进行逆转录,然后使用qPCR SYBRGreen Master Mix(No Rox)(翌圣生物科技(上海)股份有限公司)以GAPDH为内参基因进行实时荧光定量PCR实验。引物序列为:Using C57BL/6 mice, the experimental group established a sepsis model by cecum ligation and puncture (Cecum Ligation and Puncture, CLP) experiment, and the sham operation (Sham) was used as the control group. Twenty-four hours after the operation, the peripheral blood of mice was collected, RNA was extracted with RNAFast200 kit (Shanghai Feijie Biotechnology Co., Ltd.) perform reverse transcription, then use qPCR SYBRGreen Master Mix (No Rox) (Yisheng Biotechnology (Shanghai) Co., Ltd.) used GAPDH as the internal reference gene for real-time quantitative PCR experiments. The primer sequences are:
ARG1-F引物:TCACCTGAGCTTTGATGTCGA(SEQ ID NO.1)ARG1-F primer: TCACCTGAGCTTTGATGTCGA (SEQ ID NO. 1)
ARG1-R引物:TGAAAGGAGCCCTGTCTTGTA(SEQ ID NO.2)ARG1-R primer: TGAAAGGAGCCCTGTCTTGTA (SEQ ID NO. 2)
GAPDH-F引物:TCACCATCTTCCAGGAGCGAGAC(SEQ ID NO.3)GAPDH-F primer: TCACCATCTTCCAGGAGCGAGAC (SEQ ID NO. 3)
GAPDH-R引物:AGACACCAGTAGACTCCACGACATAC(SEQ ID NO.4)。GAPDH-R primer: AGACACCAGTAGACTCCACGACATAC (SEQ ID NO. 4).
扩增条件参见表4:The amplification conditions are shown in Table 4:
表4 扩增条件汇总Table 4 Summary of amplification conditions
结果如图9所示,显示ARG1在通过CLP造模的脓毒症小鼠的外周血中基因表达量要显著高于假手术组(Sham组)小鼠。这从实验层面证实了ARG1作为脓毒症生物标志物的性能。The results are shown in FIG. 9 , showing that the gene expression level of ARG1 in the peripheral blood of the septic mice modeled by CLP was significantly higher than that of the sham-operated group (Sham group) mice. This experimentally confirms the performance of ARG1 as a sepsis biomarker.
以上已对本发明创造的较佳实施例进行了具体说明,但本发明创造并不限于所述实施例,熟悉本领域的技术人员在不违背本发明创造精神的前提下还可作出种种的等同的变型或替换,这些等同的变型或替换均包含在本申请权利要求所限定的范围内。The preferred embodiments of the present invention have been specifically described above, but the present invention is not limited to the embodiments. Those skilled in the art can make various equivalents without departing from the spirit of the present invention. Modifications or substitutions, these equivalent modifications or substitutions are all included within the scope defined by the claims of the present application.
序列表sequence listing
<110> 中国人民解放军海军军医大学<110> Chinese People's Liberation Army Naval Medical University
<120> ARG1在制备脓毒症诊断、严重程度判断或预后评估试剂或试剂盒中的应用<120> Application of ARG1 in the preparation of sepsis diagnosis, severity judgment or prognosis assessment reagents or kits
<130> 权利要求书 说明书<130> Claims Specification
<160> 4<160> 4
<170> SIPOSequenceListing 1.0<170> SIPOSequenceListing 1.0
<210> 1<210> 1
<211> 21<211> 21
<212> DNA<212> DNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence
<400> 1<400> 1
tcacctgagc tttgatgtcg a 21tcacctgagc tttgatgtcg a 21
<210> 2<210> 2
<211> 21<211> 21
<212> DNA<212> DNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence
<400> 2<400> 2
tgaaaggagc cctgtcttgt a 21tgaaaggagc cctgtcttgt a 21
<210> 3<210> 3
<211> 23<211> 23
<212> DNA<212> DNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence
<400> 3<400> 3
tcaccatctt ccaggagcga gac 23tcaccatctt ccaggagcga gac 23
<210> 4<210> 4
<211> 26<211> 26
<212> DNA<212> DNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence
<400> 4<400> 4
agacaccagt agactccacg acatac 26agacaccagt agactccacg acatac 26
Claims (9)
Applications Claiming Priority (2)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| CN2022106397635 | 2022-06-08 | ||
| CN202210639763 | 2022-06-08 |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| CN115044665A true CN115044665A (en) | 2022-09-13 |
Family
ID=83162240
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| CN202210661083.3A Pending CN115044665A (en) | 2022-06-08 | 2022-06-13 | Application of ARG1 in the preparation of sepsis diagnosis, severity judgment or prognosis assessment reagents or kits |
Country Status (1)
| Country | Link |
|---|---|
| CN (1) | CN115044665A (en) |
Cited By (2)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN115678973A (en) * | 2022-11-03 | 2023-02-03 | 中国人民解放军海军军医大学 | Combined biomarker detection method for evaluating health state of organism |
| CN116994653A (en) * | 2023-09-26 | 2023-11-03 | 南方医科大学南方医院 | Sepsis diagnostic model construction method, compound screening method and electronic equipment |
Citations (3)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US20160055295A1 (en) * | 2013-06-20 | 2016-02-25 | Immunexpress Pty Ltd | Biomarker identification |
| WO2019168622A1 (en) * | 2018-02-27 | 2019-09-06 | The Board Of Trustees Of The Leland Stanford Junior University | Classifier for identification of robust sepsis subtypes |
| CN114113631A (en) * | 2021-11-26 | 2022-03-01 | 北京医院 | Sepsis test kit |
-
2022
- 2022-06-13 CN CN202210661083.3A patent/CN115044665A/en active Pending
Patent Citations (4)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US20160055295A1 (en) * | 2013-06-20 | 2016-02-25 | Immunexpress Pty Ltd | Biomarker identification |
| WO2019168622A1 (en) * | 2018-02-27 | 2019-09-06 | The Board Of Trustees Of The Leland Stanford Junior University | Classifier for identification of robust sepsis subtypes |
| CN111836906A (en) * | 2018-02-27 | 2020-10-27 | 斯坦福大学托管董事会 | A classifier for the identification of robust sepsis subtypes |
| CN114113631A (en) * | 2021-11-26 | 2022-03-01 | 北京医院 | Sepsis test kit |
Non-Patent Citations (2)
| Title |
|---|
| DINGQIAO XU等: "Metabolomics Coupled with Transcriptomics Approach Deciphering Age Relevance in Sepsis" * |
| 沈灵芝;李莉;严静;: "巨噬细胞极化在脓毒症免疫机制中的作用" * |
Cited By (2)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN115678973A (en) * | 2022-11-03 | 2023-02-03 | 中国人民解放军海军军医大学 | Combined biomarker detection method for evaluating health state of organism |
| CN116994653A (en) * | 2023-09-26 | 2023-11-03 | 南方医科大学南方医院 | Sepsis diagnostic model construction method, compound screening method and electronic equipment |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| US9238841B2 (en) | Multi-biomarker-based outcome risk stratification model for pediatric septic shock | |
| JP2023178345A (en) | Methods and systems for determining risk of autism spectrum disorder | |
| US20180004895A1 (en) | COPD Biomarker Signatures | |
| Humerfelt et al. | Effectiveness of postal smoking cessation advice: a randomized controlled trial in young men with reduced FEV1 and asbestos exposure | |
| EP2861989A2 (en) | Biomarker test for prediction or early detection of preeclampsia and/or hellp syndrome | |
| CN114875149A (en) | Application of reagent for detecting biomarkers in preparation of product for predicting gastric cancer prognosis | |
| TW202409297A (en) | Molecular biomarkers and methods of analysis for acute diagnosis of kawasaki disease | |
| Hancock et al. | Deciphering sepsis: transforming diagnosis and treatment through systems immunology | |
| CN115044665A (en) | Application of ARG1 in the preparation of sepsis diagnosis, severity judgment or prognosis assessment reagents or kits | |
| CN113970640B (en) | Biomarkers for ICH prognosis evaluation and uses thereof | |
| KR102745283B1 (en) | Biomarker for diagnosing atopic dermatitis and use thereof | |
| CN118222713A (en) | Application of biomarkers in detecting TLS associated with brain glioma | |
| WO2016123058A1 (en) | Biomarkers for detection of tuberculosis risk | |
| EP4555327A1 (en) | Methods of assessing dementia risk | |
| CN109813912B (en) | Application of group of serum differential protein combinations in preparation of reagent for detecting autism | |
| CN117233389A (en) | Markers for rapid identification of CEBPA double mutations in acute myeloid leukemia | |
| WO2023278502A1 (en) | Renal health determination and uses thereof | |
| CN118127149B (en) | Biomarker, model and kit for assessing risk of sepsis and infection in a subject | |
| CN116246710B (en) | A cluster-based colorectal cancer prediction model and its application | |
| US20260009802A1 (en) | Biomarkers | |
| CN116804676A (en) | Biomarker related to asthma and application thereof | |
| JP2025534223A (en) | Methods for assessing tobacco use status | |
| CN121023007A (en) | A biomarker for the diagnosis of blood stasis syndrome in ischemic heart failure and its application | |
| CN120249460A (en) | Intestinal microbial markers of long-lived people and their applications | |
| CN119351550A (en) | Prognostic biomarkers and their applications in retroperitoneal liposarcoma |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| PB01 | Publication | ||
| PB01 | Publication | ||
| SE01 | Entry into force of request for substantive examination | ||
| SE01 | Entry into force of request for substantive examination | ||
| RJ01 | Rejection of invention patent application after publication |
Application publication date: 20220913 |
|
| RJ01 | Rejection of invention patent application after publication |