CN114457157A - PDE1A for tumor treatment target and diagnosis biomarker, kit and application - Google Patents
PDE1A for tumor treatment target and diagnosis biomarker, kit and application Download PDFInfo
- Publication number
- CN114457157A CN114457157A CN202210161665.5A CN202210161665A CN114457157A CN 114457157 A CN114457157 A CN 114457157A CN 202210161665 A CN202210161665 A CN 202210161665A CN 114457157 A CN114457157 A CN 114457157A
- Authority
- CN
- China
- Prior art keywords
- pde1a
- seq
- base sequence
- cells
- tumor
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Granted
Links
Images
Classifications
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q1/00—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions
- C12Q1/68—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions involving nucleic acids
- C12Q1/6876—Nucleic acid products used in the analysis of nucleic acids, e.g. primers or probes
- C12Q1/6883—Nucleic acid products used in the analysis of nucleic acids, e.g. primers or probes for diseases caused by alterations of genetic material
- C12Q1/6886—Nucleic acid products used in the analysis of nucleic acids, e.g. primers or probes for diseases caused by alterations of genetic material for cancer
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K38/00—Medicinal preparations containing peptides
- A61K38/16—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof
- A61K38/17—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans
- A61K38/1703—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans from vertebrates
- A61K38/1709—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans from vertebrates from mammals
- A61K38/1738—Calcium binding proteins, e.g. calmodulin
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P11/00—Drugs for disorders of the respiratory system
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P35/00—Antineoplastic agents
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P35/00—Antineoplastic agents
- A61P35/04—Antineoplastic agents specific for metastasis
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q2600/00—Oligonucleotides characterized by their use
- C12Q2600/106—Pharmacogenomics, i.e. genetic variability in individual responses to drugs and drug metabolism
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q2600/00—Oligonucleotides characterized by their use
- C12Q2600/118—Prognosis of disease development
Landscapes
- Health & Medical Sciences (AREA)
- Life Sciences & Earth Sciences (AREA)
- Chemical & Material Sciences (AREA)
- Organic Chemistry (AREA)
- General Health & Medical Sciences (AREA)
- Animal Behavior & Ethology (AREA)
- Engineering & Computer Science (AREA)
- Proteomics, Peptides & Aminoacids (AREA)
- Veterinary Medicine (AREA)
- Public Health (AREA)
- Medicinal Chemistry (AREA)
- Pharmacology & Pharmacy (AREA)
- Zoology (AREA)
- Nuclear Medicine, Radiotherapy & Molecular Imaging (AREA)
- General Chemical & Material Sciences (AREA)
- Chemical Kinetics & Catalysis (AREA)
- Immunology (AREA)
- Bioinformatics & Cheminformatics (AREA)
- Oncology (AREA)
- Wood Science & Technology (AREA)
- Analytical Chemistry (AREA)
- Genetics & Genomics (AREA)
- Pathology (AREA)
- Hospice & Palliative Care (AREA)
- Epidemiology (AREA)
- Gastroenterology & Hepatology (AREA)
- Pulmonology (AREA)
- Physics & Mathematics (AREA)
- Biophysics (AREA)
- Biotechnology (AREA)
- Microbiology (AREA)
- Molecular Biology (AREA)
- Marine Sciences & Fisheries (AREA)
- Biochemistry (AREA)
- General Engineering & Computer Science (AREA)
- Measuring Or Testing Involving Enzymes Or Micro-Organisms (AREA)
Abstract
Description
技术领域technical field
本申请涉及肿瘤诊断技术领域,尤其涉及一种用于肿瘤治疗靶点和诊断生 物标志物的PDE1A及试剂盒和应用。The present application relates to the technical field of tumor diagnosis, and in particular, to a PDE1A, a kit and application for tumor therapy targets and diagnostic biomarkers.
背景技术Background technique
肺癌在中国以及全球的发病率位于恶性肿瘤首位,也是造成所有癌症死亡 率高的主要原因。中国每年有70多万的肺癌确诊患者和60万死亡病例,其中 近90%的肺癌患者属于非小细胞肺癌患者。近年来,针对肺癌的手术、放化疗 等医疗技术不断优化,包括分子靶向和免疫治疗等在内的治疗手段也在不断突 破取得新进展,但是,大部分中晚期肺癌患者的治疗率并没有提高很多,5年生 存率仅有16%,几乎没有提升,肺癌也成为当下危害国民生命健康安全的最主 要恶性肿瘤之一。转移被认为是肺癌死亡率居高不下的主要原因。一项研究表 明,近80%的肺癌患者在就诊时就出现了转移的II,III,IV期病情,针对肿瘤转移的患者,传统手术、放化疗法以及新兴生物疗法带来的获益很小。对于确 诊较早的肺癌患者而言,虽然治疗有效缓解了病情,但后期复发以及出现肿瘤 转移的可能性依旧较高,降低肿瘤转移发生率是降低NSCLC患者死亡率的关键, 因此,深入研究NSCLC发生转移的分子机制,研发可靠的预防及治疗手段是目 前肺癌研究最为迫切的难题。The incidence of lung cancer in China and the world ranks first among malignant tumors, and it is also the main reason for the high mortality of all cancers. There are more than 700,000 lung cancer patients diagnosed and 600,000 deaths in China every year, of which nearly 90% of lung cancer patients belong to non-small cell lung cancer patients. In recent years, medical technologies such as surgery, radiotherapy and chemotherapy for lung cancer have been continuously optimized, and treatment methods including molecular targeting and immunotherapy have also made breakthroughs and made new progress. However, the treatment rate of most patients with advanced lung cancer has not The 5-year survival rate is only 16%, which is almost no improvement. Lung cancer has also become one of the most important malignant tumors that endanger people's lives, health and safety. Metastasis is considered to be the main reason for the high mortality rate of lung cancer. A study shows that nearly 80% of lung cancer patients have metastatic stage II, III, and IV disease at the time of diagnosis, and for patients with tumor metastasis, traditional surgery, radiotherapy and chemotherapy and emerging biological therapy bring little benefit. . For lung cancer patients diagnosed earlier, although the treatment has effectively alleviated the disease, the possibility of late recurrence and tumor metastasis is still high. Reducing the incidence of tumor metastasis is the key to reducing the mortality rate of NSCLC patients. Therefore, in-depth research on NSCLC The molecular mechanism of metastasis and the development of reliable prevention and treatment methods are the most urgent problems in lung cancer research.
PDE1A(phosphodiesterase 1A)是PDE家族成员PDE1的亚型,此外还有 PDE1B和PDE1C。PDE1A主要是由Ca2+和钙调蛋白(CaM)组成,发挥作用 主要是通过钙-钙调蛋白结合到PDE1的CaM结构域,缓解催化位点的N端自 动抑制,从而促进酶活性。PDE1作为最早发现的同工酶,能够同时水解cGMP 和cAMP。PDE1A常见分布在血管平滑肌系统,具有紧张调节功能,而PDE1B 主要是在神经系统中,与记忆,免疫调节等有关。PDE1C则在两种系统中都有分布,共同参与血管平滑肌系统的生物学功能。PDE1A (phosphodiesterase 1A) is a subtype of PDE1, a member of the PDE family, in addition to PDE1B and PDE1C. PDE1A is mainly composed of Ca2+ and calmodulin (CaM), and its function is mainly through the binding of calcium-calmodulin to the CaM domain of PDE1 to relieve the N-terminal auto-inhibition of the catalytic site, thereby promoting enzyme activity. As the earliest discovered isoenzyme, PDE1 can simultaneously hydrolyze cGMP and cAMP. PDE1A is commonly distributed in the vascular smooth muscle system and has a tonic regulation function, while PDE1B is mainly found in the nervous system and is related to memory and immune regulation. PDE1C is distributed in both systems and participates in the biological function of the vascular smooth muscle system.
PDE1A可将cAMP水解,降低其在胞内表达量,从而调控人体内各种生理 过程。降低PDE1A表达能减弱白血病细胞的增殖,影响细胞周期,诱导白血病 细胞发生凋亡。Tehranolide是一种具有过氧化物基团的天然倍半萜内酯,Noori 等研究发现它能够抑制CaM的活性,进一步抑制PDE1A活化,导致细胞内cAMP 积累和PKA激活,最终抑制癌细胞增殖。PDE1A can hydrolyze cAMP and reduce its intracellular expression, thereby regulating various physiological processes in the human body. Reducing the expression of PDE1A can attenuate the proliferation of leukemia cells, affect the cell cycle, and induce apoptosis of leukemia cells. Tehranolide is a natural sesquiterpene lactone with a peroxide group. Noori et al. found that it can inhibit the activity of CaM, further inhibit the activation of PDE1A, lead to the accumulation of intracellular cAMP and activation of PKA, and finally inhibit the proliferation of cancer cells.
因此,探索PDE1A在非小细胞肺癌发生发展中的具体作用和分子机制,开 发新型早期诊断标志物和可行治疗靶标,对非小细胞肺癌患者的诊断和治疗提 供有潜力的临床诊断和治疗背景。Therefore, to explore the specific role and molecular mechanism of PDE1A in the occurrence and development of non-small cell lung cancer, develop new early diagnostic markers and feasible therapeutic targets, and provide a potential clinical diagnosis and treatment background for the diagnosis and treatment of patients with non-small cell lung cancer.
发明内容SUMMARY OF THE INVENTION
有鉴于此,本发明的目的在于提供一种用于肿瘤治疗靶点和诊断生物标志 物的PDE1A及试剂盒和应用。In view of this, the purpose of the present invention is to provide a PDE1A, a kit and application for tumor therapy target and diagnostic biomarker.
根据本发明实施例的第一方面,提供了一种用于肿瘤治疗靶点和诊断生物 标志物的PDE1A,其特征在于,所述PDE1A的碱基序列如SEQ ID NO.1所示。According to the first aspect of the embodiments of the present invention, there is provided a PDE1A used as a tumor therapy target and a diagnostic biomarker, characterized in that the base sequence of the PDE1A is shown in SEQ ID NO.1.
根据本发明实施例的第二方面,提供了一种检测样本中肿瘤标志物表达水 平的试剂盒,包括与权利要求1所述的PDE1A特异性互补配对的分子探针对和 /或用于扩增权权利要求1所述的PDE1A的引物对。According to a second aspect of the embodiments of the present invention, there is provided a kit for detecting the expression level of tumor markers in a sample, comprising a molecular probe pair specifically complementary to PDE1A according to
进一步地,所述引物对选自以下引物对1~9中的至少一种;Further, the primer pair is selected from at least one of the following primer pairs 1-9;
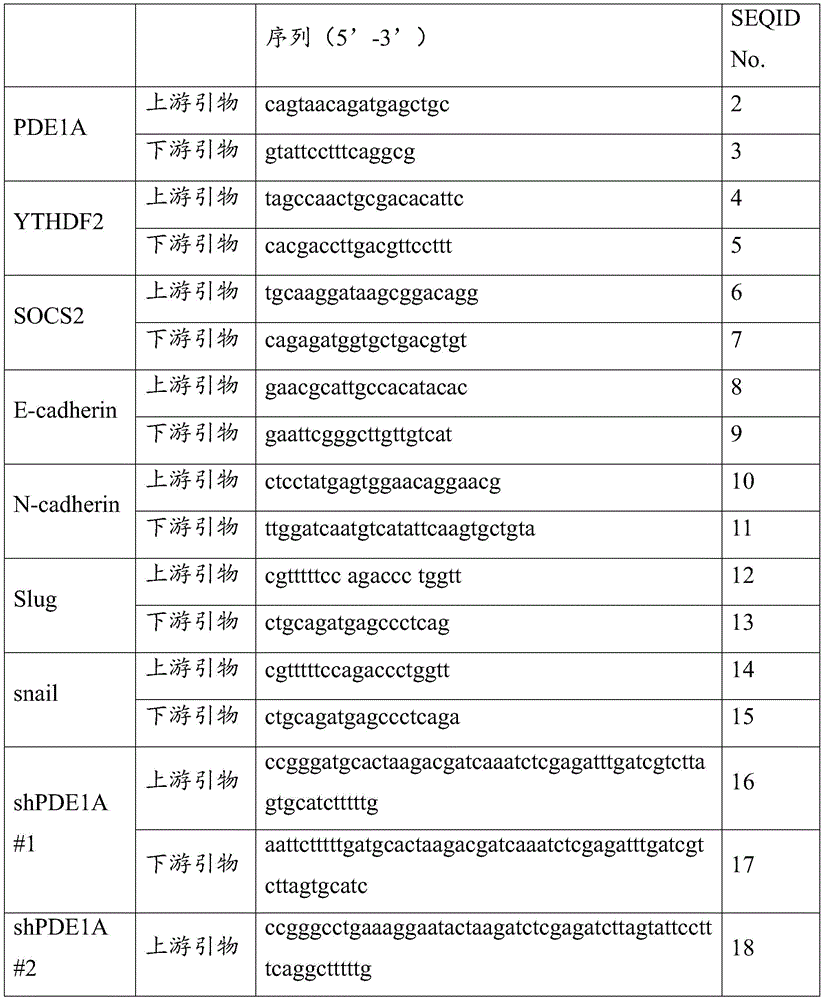
其中,引物对1的碱基序列如SEQ ID NO.2~3所示;引物对2的碱基序列 如SEQ IDNO.4~5所示;引物对3的碱基序列如SEQ ID NO.6~7所示;引物 对4的碱基序列如SEQ IDNO.8~9所示;引物对5的碱基序列如SEQ ID NO.10~11所示;引物对6的碱基序列如SEQ IDNO.12~13所示;引物对7 的碱基序列如SEQ ID NO.14~15所示;引物对8的碱基序列如SEQID NO.16~17所示;引物对9的碱基序列如SEQ ID NO.18~19所示。The base sequence of
进一步地,所述分子探针对选自以下分子探针对1~7中的至少一种;Further, the molecular probe pair is selected from at least one of the following molecular probe pairs 1-7;
其中,分子探针对1的碱基序列如SEQ ID NO.20~21所示;分子探针对2 的碱基序列如SEQ ID NO.22~23所示;分子探针对3的碱基序列如SEQ ID NO.24~25所示;分子探针对4的碱基序列如SEQ ID NO.26~27所示;分子探 针对5的碱基序列如SEQ ID NO.28~29所示;分子探针对6的碱基序列如SEQ ID NO.30~31所示;分子探针对7的碱基序列如SEQID NO.32~33所示。The base sequence of
进一步地,所述检测样本包括非小细胞肺癌组织和非小细胞肺癌癌旁组织 中的至少一种。Further, the detection sample includes at least one of non-small cell lung cancer tissue and non-small cell lung cancer adjacent tissue.
根据本发明实施例的第三方面,提供了第一方面所述的PDE1A在制备预防 和/或治疗肿瘤的药物中的应用。According to the third aspect of the embodiments of the present invention, there is provided the use of the PDE1A described in the first aspect in the preparation of a medicament for preventing and/or treating tumors.
进一步地,所述肿瘤包括肺癌,优选的,所述肿瘤包括非小细胞肺癌。Further, the tumor includes lung cancer, preferably, the tumor includes non-small cell lung cancer.
进一步地,所述肿瘤包括原发瘤和转移瘤。Further, the tumors include primary tumors and metastases.
根据本发明实施例的第四方面,提供了一种用于检测样本中肿瘤标志物表 达水平的试剂在制备用于检测非小细胞肺癌的试剂盒中的应用,所述肿瘤标志 物PDE1A碱基序列如SEQ ID No.1所示。According to a fourth aspect of the embodiments of the present invention, there is provided an application of a reagent for detecting the expression level of a tumor marker in a sample in preparing a kit for detecting non-small cell lung cancer, the tumor marker PDE1A base The sequence is shown in SEQ ID No.1.
根据本发明实施例的第五方面,提供了一种用于预防和/或治疗肿瘤的药物, 所述药物以第一方面所述PDE1A为活性成分。According to a fifth aspect of the embodiments of the present invention, there is provided a medicament for preventing and/or treating tumors, wherein the medicament uses the PDE1A of the first aspect as an active ingredient.
本申请的实施例提供的技术方案可以包括以下有益效果:The technical solutions provided by the embodiments of the present application may include the following beneficial effects:
本发明通过对PDE1A这一种标志物表达水平的检测,利用这一标志物在非 小细胞肺癌癌组织及其癌旁组织中的差异化表达,可用于非小细胞肺癌的诊断、 预后评估或筛药。The present invention detects the expression level of PDE1A, a marker, and utilizes the differential expression of this marker in non-small cell lung cancer tissue and its adjacent tissue, which can be used for the diagnosis, prognosis evaluation or evaluation of non-small cell lung cancer. Screen medicine.
应当理解的是,以上的一般描述和后文的细节描述仅是示例性和解释性的, 并不能限制本申请。It is to be understood that the foregoing general description and the following detailed description are exemplary and explanatory only and are not limiting of the present application.
附图说明Description of drawings
此处的附图被并入说明书中并构成本说明书的一部分,示出了符合本申请 的实施例,并与说明书一起用于解释本申请的原理。应当理解,以下附图仅示 出了本发明的某些实施例,因此不应被看作是对范围的限定,对于本领域普通 技术人员来讲,在不付出创造性劳动的前提下,还可以根据这些附图获得其他 相关的附图。The accompanying drawings, which are incorporated in and constitute a part of this specification, illustrate embodiments consistent with the application and together with the description serve to explain the principles of the application. It should be understood that the following drawings only show some embodiments of the present invention, and therefore should not be regarded as a limitation of the scope. Other related figures are obtained from these figures.
图1为本发明实施例提供的PDE1A对NSCLC患者总体生存的影响结果图; A为PDE1A表达与NSCLC患者风险预后的关系图;B为PDE1A表达对于 NSCLC患者生存期的影响图;Fig. 1 is a graph showing the effect of PDE1A on the overall survival of NSCLC patients provided in the embodiment of the present invention; A is a graph showing the relationship between PDE1A expression and risk prognosis of NSCLC patients; B is a graph showing the influence of PDE1A expression on the survival period of NSCLC patients;
图2为本发明实施例提供的PDE1参与调控NSCLC细胞迁移运动能力及 EMT进程,A为Transwell实验检测NCI-H1299细胞沉默PDE1A,PDE1B,PDE1C 后迁移运动能力变化结果图;B为A实验结果的统计分析图;Figure 2 is a diagram showing the PDE1 involved in regulating the migration and motility of NSCLC cells and the process of EMT provided by the embodiment of the present invention, A is the result of the Transwell experiment to detect the changes in the migration and motility of NCI-H1299 cells after silencing PDE1A, PDE1B, and PDE1C; B is the result of the A experiment result Statistical analysis chart;
图3为本发明实施例提供的Western blot检测A549细胞沉默PDE1后转移 抑制蛋白E-cadherin的蛋白变化结果图;3 is a graph showing the results of the protein change of the transfer inhibitor E-cadherin after A549 cells silenced PDE1 by Western blot detection provided in the embodiment of the present invention;
图4为本发明实施例提供的亲本细胞T0与高转移性的NSCLC细胞T3的 迁移运动能力图;A为Transwell小室构建高转移性细胞T3方法示意图;B为 NCI-H1299,A549细胞T0代与T3代细胞的迁移能力实验代表图;Figure 4 is a graph of the migration and motility of parental cells T0 and highly metastatic NSCLC cells T3 provided in the example of the present invention; A is a schematic diagram of the method for constructing highly metastatic T3 cells in a Transwell chamber; B is NCI-H1299, A549 cells T0 generation and Representative graph of the migration ability experiment of T3 cells;
图5为本发明实施例提供的Western blot检测NCI-H1299和A549亲本细胞 T0与高转移性细胞T3的PDE1A及转移相关蛋白表达结果图;Figure 5 is a graph showing the results of Western blot detection of the expression of PDE1A and metastasis-related proteins in NCI-H1299 and A549 parental cells T0 and highly metastatic cells T3 according to the embodiment of the present invention;
图6为本发明实施例提供的qRT-PCR检测亲本细胞T0与高转移性细胞T3 的转移相关蛋白的mRNA表达图;ANCI-H1299细胞T0及T3中E-cadherin, N-cadherin以及PDE1A的mRNA水平;B为A549细胞T0及T3中E-cadherin, 转录因子Slug以及PDE1A的mRNA水平结果图;Figure 6 is the qRT-PCR detection of the mRNA expression of the transfer-related proteins in parental cells T0 and highly metastatic cells T3 provided in the embodiment of the present invention; mRNA expression of E-cadherin, N-cadherin and PDE1A in T0 and T3 of ANCI-H1299 cells level; B is the mRNA level result of E-cadherin, transcription factor Slug and PDE1A in T0 and T3 of A549 cells;
图7为本发明实施例提供的PDE1A在HELF和NSCLC细胞中的表达结果 图;Figure 7 is a graph showing the expression results of PDE1A in HELF and NSCLC cells provided in the embodiment of the present invention;
图8为本发明实施例提供的沉默PDE1A后NSCLC细胞迁移能力减弱结果 图;A为A549,NCI-H1299细胞PDE1A的siRNA沉默效率检测;B为Transwell 实验检测沉默PDE1A对A549,NCI-H1299细胞迁移能力的影响结果图;C为B 实验结果的统计分析图;D为划痕实验检测沉默PDE1A对A549,NCI-H1299 伤口愈合能力影响结果图;E为D实验结果的统计分析图;Figure 8 is a graph showing the result of the reduced migration ability of NSCLC cells after silencing PDE1A provided by the examples of the present invention; A is the detection of siRNA silencing efficiency of PDE1A in A549, NCI-H1299 cells; B is Transwell assay to detect the effect of silencing PDE1A on the migration of A549, NCI-H1299 cells The graph of the effect of ability; C is the statistical analysis graph of the experimental results of B; D is the graph of the effect of the scratch test to detect the effect of silencing PDE1A on the wound healing ability of A549 and NCI-H1299; E is the statistical analysis graph of the experimental results of D;
图9为本发明实施例提供沉默PDE1A后NSCLC细胞侵袭能力减弱结果图; A为沉默PDE1A后A549,NCI-H1299细胞的侵袭代表图;B为A实验结果的 统计分析图Fig. 9 is a graph showing the result of the reduced invasive ability of NSCLC cells after silencing PDE1A in the example of the present invention; A is a representative graph of the invasion of A549 and NCI-H1299 cells after silencing PDE1A; B is a statistical analysis graph of the experimental results of A
图10为本发明实施例提供的沉默PDE1A后Western blot检测NSCLC细胞 中EMT相关蛋白变化结果图;Figure 10 is a diagram showing the results of Western blot detection of changes in EMT-related proteins in NSCLC cells after silencing PDE1A provided in the embodiment of the present invention;
图11为本发明实施例提供的沉默PDE1A后qRT-PCR检测PDE1A、 N-cadherin、Snail的mRNA变化结果图;Figure 11 is a graph showing the results of qRT-PCR detection of the mRNA changes of PDE1A, N-cadherin, and Snail after silencing PDE1A provided in the embodiment of the present invention;
图12为本发明实施例提供的PDE1A促进NSCLC细胞迁移能力图;A为 Western blot检测A549、NCI-H1299和NCI-H460细胞PDE1A过表达效率,B 为三株NSCLC细胞PDE1A过表达迁移结果图;C为B实验结果的统计分析图;Figure 12 is a graph showing the ability of PDE1A to promote the migration of NSCLC cells provided by the examples of the present invention; A shows the overexpression efficiency of PDE1A in A549, NCI-H1299 and NCI-H460 cells detected by Western blot, and B shows the migration results of PDE1A overexpression in three NSCLC cells; C is the statistical analysis diagram of the experimental results of B;
图13为本发明实施例提供的PDE1A过表达伤口愈合实验结果图和实验统 计图;Fig. 13 is the PDE1A overexpression wound healing experiment result graph and experiment statistical graph provided in the embodiment of the present invention;
图14为本发明实施例提供的PDE1A过表达侵袭实验结果图和实验统计图;FIG. 14 is a result diagram and an experimental statistical diagram of the PDE1A overexpression invasion experiment provided in the embodiment of the present invention;
图15为本发明实施例提供的Western blot检测过表达PDE1A后A549、 NCI-H1299、NCI-H460细胞EMT及转移相关蛋白变化结果图;Figure 15 is a diagram showing the results of Western blot detection of changes in EMT and metastasis-related proteins of A549, NCI-H1299, and NCI-H460 cells after PDE1A overexpression provided in the embodiment of the present invention;
图16为本发明实施例提供的过表达PDE1A促进NSCLC细胞的EMT过程 结果图;A为qRT-PCR检测NCI-H1299细胞过表达PDE1A后PDE1A、N-cadherin、 Slug的mRNA变化结果图;B为qRT-PCR检测NCI-H460细胞过表达PDE1A 后PDE1A、N-cadherin、Slug的mRNA变化结果图;Figure 16 is the result of overexpression of PDE1A to promote the EMT process of NSCLC cells provided in the example of the present invention; A is the result of qRT-PCR detection of the mRNA changes of PDE1A, N-cadherin and Slug after NCI-H1299 cells overexpressed PDE1A; B is qRT-PCR detection of the mRNA changes of PDE1A, N-cadherin and Slug after NCI-H460 cells overexpressed PDE1A;
图17为本发明实施例提供的Western blot检测PDE1A的蛋白表达水平,以 及Transwell实验检测PDE1A在体外对NCI-H1299细胞迁移运动能力的影响结 果图;Figure 17 is the Western blot that provides the embodiment of the present invention to detect the protein expression level of PDE1A, and Transwell experiment detects the effect of PDE1A on NCI-H1299 cell migration and motility in vitro results figure;
图18为本发明实施例提供的PDE1A在裸鼠体内促进NCI-H1299细胞转移 结果图;A为通过尾静脉注射的方式在每只裸鼠体内注入200×104个 NCI-H1299细胞,60天后,解剖小鼠,分出小鼠肺,并用包氏固定液固定,观 察并统计每组的转移灶个数图;B为H&E染色检测小鼠肺组织的转移灶图;C 为统计学分析对照组以及过表达PDE1A组裸鼠肺部转移灶结果图;Figure 18 is a graph showing the results of PDE1A promoting the transfer of NCI-H1299 cells in nude mice provided in the example of the present invention; A is the injection of 200×104 NCI-H1299 cells into each nude mouse by tail vein injection. After 60 days, The mice were dissected, the lungs of the mice were separated, and fixed with Bauer's fixative, and the number of metastases in each group was observed and counted; B is the map of the metastases in the lung tissue of the mice detected by H&E staining; C is the statistical analysis control group And the results of lung metastases in nude mice overexpressing PDE1A group;
图19为本发明实施例提供的shPDE1A在裸鼠体内抑制NCI-H1299细胞转 移图;A为通过尾静脉注射的方式在每只裸鼠体内注入200×104个NCI-H1299 细胞,60天后,解剖小鼠,分出小鼠肺,并用包氏固定液固定,观察并统计每 组的转移灶个数图,B为A统计学分析结果图;Figure 19 is a graph showing the inhibition of NCI-H1299 cell migration in nude mice by shPDE1A provided in the example of the present invention; A is the injection of 200×104 NCI-H1299 cells into each nude mouse by tail vein injection, and 60 days later, dissection For mice, the lungs of mice were separated and fixed with Bauer's fixative, and the number of metastases in each group was observed and counted. B is the result of statistical analysis of A;
图20本发明实施例提供的为H&E染色检测小鼠肺组织的转移灶结果图;Figure 20 is a graph of the results of H&E staining detection of metastases in mouse lung tissue provided by the embodiment of the present invention;
图21为本发明实施例提供的Gene Ontology富集分析靶向蛋白的分子功能 结果图;Fig. 21 is the molecular function result diagram of Gene Ontology enrichment analysis target protein provided in the embodiment of the present invention;
图22为本发明实施例提供的银染实验检测与PDE1A存在结合的蛋白图;Figure 22 is a map of the protein that is bound to PDE1A detected by the silver staining experiment provided in the embodiment of the present invention;
图23为本发明实施例提供的免疫沉淀实验检测PDE1A与YTHDF2的蛋白 作用图;Figure 23 is an immunoprecipitation experiment provided in the embodiment of the present invention to detect the protein interaction diagram of PDE1A and YTHDF2;
图24为本发明实施例提供的YTHDF2沉默效率结果图;A为Western blot 检测A549和NCI-H1299细胞YTHDF2的siRNA沉默效率;B为qRT-PCR检测 siYTHDF2的沉默效率结果图;Figure 24 is a graph showing the silencing efficiency of YTHDF2 provided in the embodiment of the present invention; A is the siRNA silencing efficiency of YTHDF2 detected by Western blot in A549 and NCI-H1299 cells; B is the result graph of the silencing efficiency of siYTHDF2 detected by qRT-PCR;
图25为本发明实施例提供的YTHDF2逆转PDE1A促进的NCI-H1299细胞 转移结果图;A为沉默YTHDF2对PDE1A促进NCI-H1299细胞迁移结果图;B 为A统计分析图;C为沉默YTHDF2对PDE1A促进NCI-H1299细胞侵袭能力 影响结果图;D为C统计分析图;Figure 25 is a graph of the results of YTHDF2 reversing the migration of NCI-H1299 cells promoted by PDE1A provided in the example of the present invention; A is a graph of the results of silencing YTHDF2 on PDE1A to promote the migration of NCI-H1299 cells; B is a statistical analysis graph of A; C is a graph of silencing YTHDF2 on PDE1A The graph of the effect of promoting the invasion ability of NCI-H1299 cells; D is the statistical analysis graph of C;
图26为本发明实施例提供的RNA免疫沉淀-qPCR实验检测SCOS2 mRNA 在抗PDE1A沉淀物和抗YTHDF2沉淀物中的富集结果图;Figure 26 is a graph showing the results of the RNA immunoprecipitation-qPCR assay provided in the embodiment of the present invention to detect the enrichment of SCOS2 mRNA in anti-PDE1A precipitates and anti-YTHDF2 precipitates;
图27为本发明实施例提供的在NCI-H1299细胞中转染METTL3的siRNA 后过表达PDE1A和YTHDF2,转染48h后收集mRNA,qRT-PCR检测SOCS2 的mRNA变化结果图;Figure 27 is a graph of the results of overexpressing PDE1A and YTHDF2 after transfecting METTL3 siRNA in NCI-H1299 cells provided in the embodiment of the present invention, collecting
图28为本发明实施例提供的在NCI-H1299细胞中过表达PDE1A/YTHDF2 后再转染YTHDF2/PDE1A的siRNA或同时转染YTHDF2和PDE1A的siRNA, 转染48h后收集mRNA,qRT-PCR检测SOCS2的mRNA变化结果图;Figure 28 shows the overexpression of PDE1A/YTHDF2 in NCI-H1299 cells and then transfected siRNA of YTHDF2/PDE1A or the siRNA of YTHDF2 and PDE1A simultaneously transfected in NCI-H1299 cells provided in the example of the present invention, mRNA was collected 48 hours after transfection, and detected by qRT-PCR The result of mRNA change of SOCS2;
图29为本发明实施例提供的PDE1A影响SOCS2 mRNA降解结果图;A为 在A549细胞中沉默PDE1A的后,在0h,3h和6h进行放线菌素D(5μg/ml)处 理分析SOCS2的mRNA半衰期结果图;B为A549细胞中沉默PDE1A的后, 在0h,3h和6h进行放线菌素D(5μg/ml)处理分析SOCS2 mRNA表达结果图;Figure 29 shows the results of the degradation of SOCS2 mRNA affected by PDE1A provided in the example of the present invention; A is the analysis of SOCS2 mRNA after silencing PDE1A in A549 cells after treatment with actinomycin D (5 μg/ml) at 0h, 3h and 6h Half-life result graph; B is the result graph of SOCS2 mRNA expression analysis after silencing PDE1A in A549 cells, treated with actinomycin D (5μg/ml) at 0h, 3h and 6h;
图30为本发明实施例提供的PDE1A影响SOCS2 mRNA降解结果图;A为 在NCI-H1299细胞中沉默PDE1A的后,在0h,3h和6h进行放线菌素D(5μg/ml) 处理分析SOCS2的mRNA半衰期;B为在NCI-H1299细胞中沉默PDE1A的后, 在0h,3h和6h进行放线菌素D(5μg/ml)处理分析SOCS2 mRNA表达结果图。Figure 30 shows the results of the degradation of SOCS2 mRNA affected by PDE1A provided in the example of the present invention; A is the analysis of SOCS2 after silencing PDE1A in NCI-H1299 cells at 0h, 3h and 6h with actinomycin D (5μg/ml) treatment The mRNA half-life of ; B is the result of analysis of SOCS2 mRNA expression after silencing PDE1A in NCI-H1299 cells at 0 h, 3 h and 6 h with actinomycin D (5 μg/ml) treatment.
具体实施方式Detailed ways
这里将详细地对示例性实施例进行说明,其示例表示在附图中。下面的描 述涉及附图时,除非另有表示,不同附图中的相同数字表示相同或相似的要素。 以下示例性实施例中所描述的实施方式并不代表与本申请相一致的所有实施方 式。相反,它们仅是与如所附权利要求书中所详述的、本申请的一些方面相一 致的装置和方法的例子。Exemplary embodiments will be described in detail herein, examples of which are illustrated in the accompanying drawings. When the following description refers to the drawings, the same numbers in different drawings refer to the same or similar elements unless otherwise indicated. The implementations described in the following exemplary examples are not intended to represent all implementations consistent with this application. Rather, they are merely examples of apparatus and methods consistent with some aspects of the present application as recited in the appended claims.
在本申请使用的术语是仅仅出于描述特定实施例的目的,而非旨在限制本 申请。在本申请和所附权利要求书中所使用的单数形式的“一种”、“所述”和“该” 也旨在包括多数形式,除非上下文清楚地表示其他含义。还应当理解,本文中 使用的术语“和/或”是指并包含一个或多个相关联的列出项目的任何或所有可能 组合。The terminology used in this application is for the purpose of describing particular embodiments only and is not intended to limit the application. As used in this application and the appended claims, the singular forms "a," "the," and "the" are intended to include the plural forms as well, unless the context clearly dictates otherwise. It will also be understood that the term "and/or" as used herein refers to and includes any and all possible combinations of one or more of the associated listed items.
本发明提供了检测样本中PDE1A表达水平的试剂在制备用于非小细胞肺癌 检测的试剂盒中的应用。The present invention provides the application of a reagent for detecting the expression level of PDE1A in a sample in preparing a kit for detecting non-small cell lung cancer.
上述肿瘤标志物PDE1A属于RNA肿瘤标志物。The above-mentioned tumor marker PDE1A belongs to RNA tumor markers.
本文中的“检测”可以指对样品进行非小细胞肺癌的诊断、辅助诊断或预 后评估的检测。As used herein, "detection" may refer to detection of a sample for the diagnosis, auxiliary diagnosis, or prognostic assessment of non-small cell lung cancer.
具体地,PDE1A的碱基序列如SEQ ID NO.1所示。Specifically, the base sequence of PDE1A is shown in SEQ ID NO.1.
在一些实施例中,所述试剂包括与第一方面所述PDE1A特异性互补配对的 分子探针或用于扩增第一方面所述的PDE1A的引物对。In some embodiments, the reagents include molecular probes that specifically pair complementary to the PDE1A of the first aspect or a primer pair for amplifying the PDE1A of the first aspect.
优选地,所述试剂包括:用于检测PDE1A的引物对1。Preferably, the reagents include:
其中,引物对1的碱基序列如SEQ ID NO.2~3所示。The base sequences of
需要说明的是,引物对通常包括上游引物和下游引物,本文中“引物对1 的碱基序列如SEQ ID NO.2~3所示”是指引物对中上游引物的碱基序列如SEQ ID NO.2所示,下游引物的碱基序列如SEQ ID NO.3所示。It should be noted that a primer pair usually includes an upstream primer and a downstream primer, and "the base sequence of
优选地,所述样本包括生物组织样本。Preferably, the sample comprises a biological tissue sample.
优选地,所述生物组织样本包括非小细胞肺癌癌组织和非小细胞肺癌癌旁 组织中的至少一种。Preferably, the biological tissue sample includes at least one of non-small cell lung cancer tissue and non-small cell lung cancer adjacent tissue.
本发明实施例还提供了检测样本中肿瘤标志物表达水平的试剂在筛选用于 治疗非小细胞肺癌药物中的应用,所述肿瘤标志物包括碱基序列依次如SEQID No.1所示的PDE1A,所述应用不以疾病的诊断或治疗为直接目的。The embodiment of the present invention also provides an application of a reagent for detecting the expression level of a tumor marker in a sample in screening a drug for the treatment of non-small cell lung cancer, where the tumor marker includes PDE1A whose base sequence is shown in SEQID No.1 in sequence , the application is not directly aimed at the diagnosis or treatment of the disease.
发明人发现,上述肿瘤标志物PDE1A在非小细胞肺癌患者的癌组织和癌旁 组织中的表达具有差异化,通过对患者癌组织和癌旁组织中的标志物进行检测, 能够对非小细胞肺癌进行诊断和预后评估,为非小细胞肺癌的药物开发和延长 预后具有重要作用。The inventors found that the expression of the above-mentioned tumor marker PDE1A in the cancer tissue and paracancerous tissue of patients with non-small cell lung cancer is differentiated. The diagnosis and prognostic assessment of lung cancer play an important role in drug development and prolonged prognosis of non-small cell lung cancer.
上述标志物能用于非小细胞肺癌的诊断和预后评估,因此,通过对上述标 志物的检测,能够加快开发针对非小细胞肺癌患者特异性的靶向药物。下面结 合实施例对本发明提供的一种用于肿瘤治疗靶点和诊断生物标志物的PDE1A 及试剂盒和应用进行详细的说明,但是不能把它们理解为对本发明保护范围的 限定。The above-mentioned markers can be used for the diagnosis and prognosis evaluation of non-small cell lung cancer. Therefore, through the detection of the above-mentioned markers, the development of specific targeted drugs for patients with non-small cell lung cancer can be accelerated. The PDE1A, the kit and the application for tumor therapy targets and diagnostic biomarkers provided by the present invention are described in detail below with reference to the examples, but they should not be construed as limiting the protection scope of the present invention.
实施例1Example 1
PDE1A表达与肺癌患者预后相关:PDE1A expression is associated with prognosis in lung cancer patients:
首先,本实施例通过对TCGA数据库的检索分析发现,肺癌患者中PDE1A 表达高的患者的风险预后要高于PDE1A表达低的患者,而且PDE1A高表达的 肺癌患者的生存期显著缩短(参见图1)。可见,本实施例成功预测PDE1A可 能参与肺癌的发生发展,PDE1A高表达与肺癌患者的不良预后密切相关。First of all, through the search and analysis of the TCGA database in this example, it is found that the risk prognosis of patients with high PDE1A expression in lung cancer patients is higher than that of patients with low PDE1A expression, and the survival time of lung cancer patients with high PDE1A expression is significantly shortened (see Figure 1 ). ). It can be seen that this example successfully predicts that PDE1A may be involved in the occurrence and development of lung cancer, and the high expression of PDE1A is closely related to the poor prognosis of lung cancer patients.
实施例2Example 2
PDE1A与NSCLC细胞的迁移与上皮间质转化相关:Migration of PDE1A and NSCLC cells is associated with epithelial-mesenchymal transition:
本实施例于实验前在NSCLC细胞NCI-H1299上对PDE1的3个亚型, PDE1A、PDE1B和PDE1C分别进行了siRNA基因沉默实验,然后采用Transwell 方法检测了细胞的迁移能力。结果发现,沉默PDE1均能抑制NCI-H1299细胞 的迁移运动能力,其中,沉默PDE1A后,NCI-H1299细胞的迁移运动能力抑制 最为显著(参见图2)。此外,本实施例也检测了A549细胞沉默PDE1后,转 移抑制蛋白E-cadherin的变化,发现沉默PDE1A组E-cadherin的蛋白增加最为 明显(参见图3)。提示,PDE1A表达可能与NSCLC的转移有关。In this example, siRNA gene silencing experiments were performed on three subtypes of PDE1, PDE1A, PDE1B and PDE1C, respectively, on NSCLC cells NCI-H1299 before the experiment, and then the migration ability of the cells was detected by Transwell method. The results showed that silencing of PDE1 could inhibit the migration and motility of NCI-H1299 cells, among which, after silencing PDE1A, the migration and motility of NCI-H1299 cells was inhibited most significantly (see Figure 2). In addition, this Example also detected the changes of the transfer inhibitor E-cadherin after A549 cells silenced PDE1, and found that the protein of E-cadherin increased most obviously in the silenced PDE1A group (see Figure 3). It is suggested that the expression of PDE1A may be related to the metastasis of NSCLC.
本实施例前期通过3次Transwell小室富集的方法构建了NSCLC/T3细胞系, 研究不同转移性细胞系的迁移运动能力,结果如图所示,与亲本细胞 (NCI-H1299/T0,A549/T0)相比,高转移性的细胞(NCI-H1299/T3,A549/T3) 具有更强的迁移运动能力(参见图4)。为了进一步研究PDE1A与NSCLC细 胞转移相关性,本实施例从蛋白和mRNA两个层面对不同迁移能力的细胞进行 分析。结果发现,A549,NCI-H1299的高迁移能力细胞的促转移相关蛋白N-cadherin、Vimentin和Snail的蛋白表达更高,而转移抑制蛋白E-cadherin表达 更低。有意思的是,与低转移能力的NSCLC相比,本实施例在高转移能力的细 胞中还观察到了PDE1A与转录因子smad 3的磷酸化蛋白水平的增加(参见图5)。 同时,我们也从mRNA水平进一步验证了这一结果(参见图6),以上数据提 示,PDE1A很可能具有潜在的促进转移作用。In the early stage of this example, NSCLC/T3 cell line was constructed by 3 times of Transwell chamber enrichment method, and the migration and motility of different metastatic cell lines were studied. Compared with T0), the highly metastatic cells (NCI-H1299/T3, A549/T3) had stronger migratory motility (see Figure 4). In order to further study the correlation between PDE1A and the metastasis of NSCLC cells, this example analyzes cells with different migration abilities from two levels of protein and mRNA. The results showed that the high migration ability cells of A549 and NCI-H1299 had higher protein expressions of the pro-metastasis-related proteins N-cadherin, Vimentin and Snail, but lower expression of the metastasis inhibitory protein E-cadherin. Interestingly, increased levels of phosphorylated proteins of PDE1A and
实施例3Example 3
PDE1A在HELF和NSCLC细胞中的表达差异:为了评估PDE1A在NSCLC 细胞中的表达,本实施例选取了正常的人肺成纤维细胞与不同的NSCLC细胞系 进行研究。Western blot实验结果表明,与正常人肺成纤维细胞HELF相比, PDE1A在NSCLC细胞A549的表达较高,NCI-H1299的表达中等,NCI-H460 中表达较低,而在HELF中几乎不表达(参见图7)。这提示PDE1A蛋白的表 达程度与NSCLC细胞的恶性化相关。Differences in the expression of PDE1A in HELF and NSCLC cells: In order to evaluate the expression of PDE1A in NSCLC cells, normal human lung fibroblasts and different NSCLC cell lines were selected for research in this example. Western blot experiments showed that compared with normal human lung fibroblasts HELF, PDE1A was highly expressed in NSCLC cells A549, moderately expressed in NCI-H1299, lower in NCI-H460, and almost not expressed in HELF ( See Figure 7). This suggests that the expression level of PDE1A protein is related to the malignancy of NSCLC cells.
实施例4Example 4
PDE1A促进NSCLC细胞的迁移,侵袭能力:PDE1A promotes the migration, invasion ability of NSCLC cells:
首先,通过Transwell实验检测了沉默PDE1A前后NSCLC细胞的迁移能力 变化,发现沉默PDE1A后,A549,NCI-H1299细胞的迁移细胞数显著少于阴性 对照组(参见图8);同时通过划痕实验检测沉默PDE1A对NSCLC细胞伤口 愈合能力的影响。结果表明,沉默PDE1A能降低A549,NCI-H1299细胞的伤 口愈合能力(参见图9)。采用Matrigel基质胶铺小室进行侵袭实验发现,沉默 PDE1A后,细胞穿过Matrigel基质胶的数目显著低于阴性对照组(参见图10), 以上结果说明沉默PDE1A能抑制NSCLC细胞的迁移,侵袭能力。First, the migration ability of NSCLC cells before and after silencing PDE1A was detected by Transwell assay, and it was found that after silencing PDE1A, the number of migrating cells in A549 and NCI-H1299 cells was significantly less than that in the negative control group (see Figure 8); Effects of silencing PDE1A on the wound healing ability of NSCLC cells. The results showed that silencing of PDE1A reduced the wound healing ability of A549, NCI-H1299 cells (see Figure 9). The Matrigel matrigel-coated chamber was used for the invasion assay, and it was found that after silencing PDE1A, the number of cells passing through Matrigel was significantly lower than that of the negative control group (see Figure 10). The above results indicate that silencing PDE1A can inhibit the migration and invasion of NSCLC cells.
肿瘤在转移的过程中,肿瘤细胞的骨架或运动相关蛋白会发生变化。肿瘤 转移中的一个重要过程是上皮间质转化(EMT),此过程中会伴有相关蛋白变 化,如转移抑制蛋白E-cadherin和促转移蛋白Fibronectin,N-cadherin等,因此 我们进一步研究了沉默PDE1A对这些蛋白的影响。结果发现,沉默PDE1A后, 转录因子Snail,Slug的蛋白水平下降,转录因子抑制E-cadherin的启动子活性, 激活其蛋白转录,降低N-cadherin蛋白的表达,进而抑制肿瘤转移(参见图11)。 另外从mRNA水平进行了检测,qRT-PCR实验结果表明,沉默PDE1A抑制转 录因子Snail表达,抑制促转移蛋白N-cadherin的mRNA表达(参见图12), 以上蛋白表达和mRNA表达水平与本实施例在前面观察到的转移运动现象一致。During the process of tumor metastasis, the cytoskeleton or movement-related proteins of tumor cells will change. An important process in tumor metastasis is epithelial-mesenchymal transition (EMT), which is accompanied by changes in related proteins, such as metastasis suppressor protein E-cadherin and pro-metastatic proteins Fibronectin, N-cadherin, etc. Therefore, we further studied the silencing of Effects of PDE1A on these proteins. The results showed that after silencing PDE1A, the protein levels of the transcription factors Snail and Slug decreased, and the transcription factors inhibited the promoter activity of E-cadherin, activated its protein transcription, and reduced the expression of N-cadherin protein, thereby inhibiting tumor metastasis (see Figure 11). . In addition, the mRNA level was detected. The results of qRT-PCR experiments showed that silencing PDE1A inhibited the expression of the transcription factor Snail and the mRNA expression of the pro-transfer protein N-cadherin (see Figure 12). The above protein expression and mRNA expression levels were the same as those in this example. The transfer motion phenomenon observed earlier is consistent.
为了进一步研究PDE1A对NSCLC细胞转移侵袭运动能力的影响,本实施 例在A549、NCI-H1299和NCI-H460细胞中过表达PDE1A,检测了细胞的迁移 和侵袭能力。结果如图所示,过表达PDE1A显著增加了细胞的迁移数和迁移距 离,即细胞迁移能力增强(参见图13-14),Matrigel基质胶铺小室Transwell 实验也进一步表明过表达PDE1A能显著增强细胞的侵袭能力(Figure 10(参见 图15)。以上结果说明,PDE1A能促进NSCLC细胞的迁移,侵袭运动能力。In order to further study the effect of PDE1A on the ability of metastasis and invasion of NSCLC cells, in this example, PDE1A was overexpressed in A549, NCI-H1299 and NCI-H460 cells, and the migration and invasion abilities of the cells were detected. As shown in the figure, overexpression of PDE1A significantly increased the number and distance of cell migration, that is, the cell migration ability was enhanced (see Figure 13-14). Matrigel Matrigel-coated chamber Transwell experiments further showed that overexpression of PDE1A could significantly enhance cell migration. The invasion ability of NSCLC cells (Figure 10 (see Figure 15). The above results show that PDE1A can promote the migration and invasion ability of NSCLC cells.
接着,本实施例通过Western blot检测了过表达PDE1A后转移相关蛋白的 变化情况。结果发现,PDE1A过表达会引起转录因子STAT3和smad3的磷酸化 水平增加,转移抑制蛋白E-cadherin表达降低,促转移蛋白N-cadherin的表达升 高(参见图16);另外选取了PDE1A本底蛋白表达较低的NCI-H1299和NCI-H460 细胞,进行PDE1A过表达实验,然后检测了PDE1A与EMT相关蛋白及转录因 子的mRNA变化,结果显示,过表达PDE1A能够上调N-cadherin和Slug的mRNA (参见图17)。以上提示,PDE1A参与NSCLC细胞的EMT过程,在其转移过程中具有重要作用。Next, in this example, the changes of transfer-related proteins after overexpression of PDE1A were detected by Western blot. The results showed that the overexpression of PDE1A would increase the phosphorylation levels of transcription factors STAT3 and smad3, decrease the expression of the metastasis suppressor protein E-cadherin, and increase the expression of the transfer-promoting protein N-cadherin (see Figure 16). In addition, the PDE1A background was selected. In NCI-H1299 and NCI-H460 cells with low protein expression, PDE1A overexpression experiment was performed, and then the mRNA changes of PDE1A and EMT-related proteins and transcription factors were detected. The results showed that overexpression of PDE1A can upregulate the mRNA of N-cadherin and Slug (See Figure 17). The above suggests that PDE1A is involved in the EMT process of NSCLC cells and plays an important role in its metastasis.
实施例5Example 5
PDE1A在裸鼠体内对NCI-H1299细胞转移的影响:The effect of PDE1A on the metastasis of NCI-H1299 cells in nude mice:
通过前面的研究,本实施例发现PDE1A对NSCLC细胞的迁移和侵袭能力 具有显著促进作用。然而,肿瘤的发生发展由肿瘤细胞本身以及肿瘤微环境共 同决定,为了进一步研究PDE1A在体内对NSCLC细胞转移和定植能力的影响, 本实施例构建了稳定过表达PDE1A的NCI-H1299细胞系(PDE1A: pLenti-CMV-PDE1A-Flag)以及对照(vect:pLenti-CMV-PCIG3-Flag)。结果表 明,稳定过表达PDE1A的NCI-H1299细胞的迁移能力更强(参见图18),即 稳定过表达模型构建成功。另外,通过尾静脉注射的方式,构建了裸小鼠模型, 进一步确证了PDE1A在体内的促转移作用。解剖出小鼠的各个器官后,对裸鼠 肺部进行包式液浸染,发现稳定过表达PDE1A组细胞的裸鼠肺部灶点数多于对 照组,对裸鼠肺组织石蜡包埋后进行H&E染色,结果表明,过表达PDE1A组 裸鼠肺部存在明显的转移灶(参见图19)。因此,PDE1A能够在裸鼠体内促进 NSCLC细胞转移。Through the previous studies, this example found that PDE1A has a significant promoting effect on the migration and invasion ability of NSCLC cells. However, the occurrence and development of tumors are determined by the tumor cells themselves and the tumor microenvironment. In order to further study the effect of PDE1A on the metastasis and colonization ability of NSCLC cells in vivo, the NCI-H1299 cell line (PDE1A stably overexpressing PDE1A) was constructed in this example. : pLenti-CMV-PDE1A-Flag) and control (vect: pLenti-CMV-PCIG3-Flag). The results showed that the NCI-H1299 cells stably overexpressing PDE1A had stronger migration ability (see Figure 18), that is, the stable overexpression model was successfully constructed. In addition, a nude mouse model was constructed by means of tail vein injection, which further confirmed the pro-metastatic effect of PDE1A in vivo. After dissecting the various organs of the mice, the lungs of nude mice were stained with encapsulated liquid, and it was found that the number of foci in the lungs of nude mice with stable overexpression of PDE1A cells was more than that of the control group. The results showed that there were obvious metastases in the lungs of the nude mice in the PDE1A overexpression group (see Figure 19). Therefore, PDE1A can promote NSCLC cell metastasis in nude mice.
为了进一步研究PDE1A在裸鼠体内对NCI-H1299细胞转移的影响,本实施 例也构建了沉默PDE1A的shPDE1A模型,将稳定转染shPDE1A和对照plko.1 的NCI-H1299细胞系通过尾静脉注射的方式接种在裸鼠体内。结果发现, shPDE1A在裸鼠体内抑制NCI-H1299细胞转移。shPDE1A组裸鼠肺部的转移灶 点数明显少于对照plko.1组,且H&E染色结果显示,shPDE1A组无明显转移灶 (参见图20-21)。以上结果说明,shPDE1A在裸鼠体内抑制NCI-H1299细胞 转移。In order to further study the effect of PDE1A on the transfer of NCI-H1299 cells in nude mice, the shPDE1A model of silencing PDE1A was also constructed in this example, and the NCI-H1299 cell lines stably transfected with shPDE1A and control plko. inoculated into nude mice. The results showed that shPDE1A inhibited the metastasis of NCI-H1299 cells in nude mice. The number of metastases in the lungs of nude mice in the shPDE1A group was significantly less than that in the control plko.1 group, and the results of H&E staining showed that there were no obvious metastases in the shPDE1A group (see Figure 20-21). The above results indicated that shPDE1A inhibited the metastasis of NCI-H1299 cells in nude mice.
实施例6Example 6
PDE1A与YTHDF2发生结合,并逆转PDE1A促进NSCLC细胞迁移侵袭:PDE1A binds to YTHDF2 and reverses PDE1A to promote NSCLC cell migration and invasion:
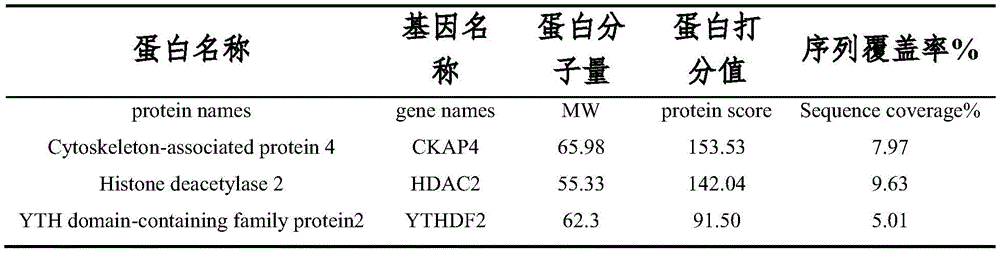
为了进一步寻找参与调控PDE1A促进非小细胞肺癌细胞转移的下游靶点分 子,本实施例对A549细胞进行质粒转染过表达PDE1A后,收集样本进行免疫 沉淀实验,Westernblot跑胶后,将胶用考马斯亮蓝染色,送样进行蛋白质谱分 析。根据质谱分析结果中Score得分情况(参见表1)和Gene Ontology富集分 析(参见图22-23),我们发现m6A结合蛋白(GO:1990247)富集倍数较大,结合蛋白的富集最为显著,综合银染结果,最终选择YTHDF2作为下一步 的研究对象。In order to further search for the downstream target molecules involved in regulating PDE1A to promote the metastasis of non-small cell lung cancer cells, in this example, after plasmid transfection of A549 cells to overexpress PDE1A, samples were collected for immunoprecipitation experiments. The samples were stained with Maas Brilliant Blue and sent for protein profiling. According to the Score in the mass spectrometry results (see Table 1) and the Gene Ontology enrichment analysis (see Figures 22-23), we found that the m6A-binding protein (GO: 1990247) had a larger enrichment fold, and the binding protein had the most significant enrichment. , comprehensive silver staining results, and finally selected YTHDF2 as the next research object.
表1蛋白质谱分析与PDE1A表达相关度较高蛋白Table 1 Protein profiling analysis is highly correlated with PDE1A expression
PDE1A主要分布在细胞胞浆内,一般胞浆内的蛋白往往是通过与蛋白质的 相互结合来实现其生物学功能的,因此采用免疫沉淀实验检测PDE1A是否与 YTHDF2存在蛋白结合作用。结果发现,PDE1A能够与YTHDF2产生相互结合 (参见图24)。然后我们又检测了沉默YTHDF2对PDE1A促进的NSCLC细 胞迁移,侵袭能力的影响,结果发现,YTHDF2能够逆转PDE1A促进NSCLC 细胞迁移,侵袭能力增强(参见图25-26)。PDE1A is mainly distributed in the cytoplasm of cells. Generally, the proteins in the cytoplasm often realize their biological functions by combining with proteins. Therefore, immunoprecipitation experiments were used to detect whether PDE1A has protein binding effect with YTHDF2. As a result, it was found that PDE1A was able to interact with YTHDF2 (see Fig. 24). Then we examined the effect of silencing YTHDF2 on PDE1A-promoted NSCLC cell migration and invasion, and found that YTHDF2 could reverse PDE1A-promoted NSCLC cell migration and enhanced invasion (see Figures 25-26).
YTHDF2是目前已知的m6A修饰过程中的阅读蛋白之一。YTHDF2识别 mRNA上的m6A修饰会引导CCR4-NOT蛋白复合体降解所识别的mRNA。有 文献表明,SOCS2作为YTHDF2的下游靶点,可被YTHDF2降解其mRNA, 于是首先利用RNA免疫沉淀-qPCR技术验证PDE1A以及YTHDF2与SOCS2 是否存在RNA结合,实验结果发现,与IgG组比较,分别由PDE1A抗体和YTHDF2抗体所富集的SOCS2 mRNA明显增多,这一结果说明PDE1A和 YTHDF2与SOCS2 mRNA存在相互结合(参见图27)。本实施例还利用荧光 定量PCR技术检测METTL3、YTHDF2以及PDE1A三者表达改变引起的SOCS2 表达水平的变化,结果发现影响METTL3、YTHDF2以及PDE1A三者表达可以 引起SOCS2的mRNA水平的变化(参见图28-29)。此外,mRNA稳定性测定 显示,在A549和NCI-H1299细胞中,通过PDE1A沉默,SOCS2的mRNA表 达升高,并且SOCS2的mRNA半衰期持续延长(参见图30)。YTHDF2 is one of the currently known readers during m6A modification. YTHDF2 recognizes m6A modifications on mRNAs that direct the CCR4-NOT protein complex to degrade the recognized mRNAs. Some literatures show that SOCS2, as a downstream target of YTHDF2, can be degraded by YTHDF2 for its mRNA. Therefore, RNA immunoprecipitation-qPCR technology was used to verify whether there is RNA binding between PDE1A and YTHDF2 and SOCS2. The SOCS2 mRNA enriched by PDE1A antibody and YTHDF2 antibody was significantly increased, which indicated that PDE1A and YTHDF2 were combined with SOCS2 mRNA (see Figure 27). In this example, fluorescence quantitative PCR technology was used to detect changes in the expression levels of SOCS2 caused by changes in the expressions of METTL3, YTHDF2 and PDE1A, and it was found that affecting the expressions of METTL3, YTHDF2 and PDE1A could cause changes in the mRNA levels of SOCS2 (see Figure 28 ). -29). In addition, mRNA stability assays showed that SOCS2 mRNA expression was elevated and SOCS2 mRNA half-life continued to be prolonged by PDE1A silencing in A549 and NCI-H1299 cells (see Figure 30).
上述实施例中所述引物对序列及探针碱基对序列,具体如表2所示。The primer pair sequences and probe base pair sequences described in the above embodiments are shown in Table 2 in detail.
表2:Table 2:
实验方法:experimental method:
1.细胞培养1. Cell Culture
人非小细胞肺癌细胞A549,NCI-H1299,NCI-H460和人脐静脉内皮细胞 HUVEC培养于具有10%FBS和90%RPMI 1640培养基中。所有细胞在5%CO2, 37℃的细胞培养箱中培养。Human non-small cell lung cancer cells A549, NCI-H1299, NCI-H460 and human umbilical vein endothelial cells HUVEC were cultured in medium with 10% FBS and 90% RPMI 1640. All cells were cultured in a cell incubator at 37°C with 5% CO 2 .
2.Transwell实验2. Transwell experiment
(1)细胞迁移实验(1) Cell Migration Experiment
将高低表达circPOLK的肺癌细胞血清饥饿过夜,消化计数,接种到transwell 小室中,密度为2×104/孔,每孔无血清培液200μl。相对应的24孔板中加入600 μl含有20%血清的培养液。细胞孵育24小时后,用1%结晶紫对小室进行避光 染色30min,接着用1×PBS清洗小室内部的细胞,然后用显微镜拍摄迁移到小 室下表面的细胞并计数。Lung cancer cells with high and low expression of circPOLK were serum-starved overnight, digested and counted, and inoculated into transwell chambers at a density of 2×10 4 /well, with 200 μl of serum-free culture medium per well. 600 μl of culture medium containing 20% serum was added to the corresponding 24-well plate. After the cells were incubated for 24 hours, the chamber was stained with 1% crystal violet for 30 min in the dark, then the cells inside the chamber were washed with 1×PBS, and the cells that migrated to the lower surface of the chamber were photographed and counted under a microscope.
(2)细胞侵袭实验(2) Cell invasion assay
Matrigel提前放置4℃解冻,小室,24孔板和所用枪头放置-20℃预冷。用 细胞培养液将Matrigel按1:20稀释,每个小室中加入100μl的Matrigel,将小 室放在24孔板中置于37℃孵箱30min,取出24孔板,弃去小室中未凝固的 Matrigel,并用培养液润洗一遍。与此同时,高低表达circPOLK的非小细胞肺 癌细胞血清饥饿过夜,消化计数,接种到有Matrigel胶的transwell小室中,密 度为3×104/孔,每孔无血清培养液200μl。相对应的24孔板中加入600μl含有 20%血清的培养液。细胞孵育24小时后,用1%结晶紫对小室进行避光染色30 min,接着用1×PBS清洗小室内部的细胞,然后用显微镜拍摄侵袭到小室下表 面的细胞并计数。Matrigel was thawed at 4°C in advance, and the chamber, 24-well plate and pipette tips used were pre-cooled at -20°C. Dilute Matrigel 1:20 with cell culture medium, add 100 μl of Matrigel to each chamber, place the chamber in a 24-well plate and place it in a 37°C incubator for 30 minutes, take out the 24-well plate, and discard the uncoagulated Matrigel in the chamber. , and rinsed with culture medium. At the same time, non-small cell lung cancer cells with high and low expression of circPOLK were serum starved overnight, digested and counted, and inoculated into a transwell chamber with Matrigel gel at a density of 3×10 4 /well, and each well of serum-free medium was 200 μl. 600 μl of culture medium containing 20% serum was added to the corresponding 24-well plate. After the cells were incubated for 24 hours, the chamber was stained with 1% crystal violet for 30 min in the dark, and then the cells inside the chamber were washed with 1×PBS, and the cells that invaded the lower surface of the chamber were photographed and counted with a microscope.
3.伤口愈合实验3. Wound Healing Experiment
将细胞以2×105/孔的密度接种在六孔板中,待细胞贴壁后,转染所需质粒 或siRNA。待细胞汇合度达到80-100%,用移液器在细胞表面划痕,用PBS洗 去表面掉落的细胞,换上新的培养液,显微镜拍下0h的伤口距离,24h后,用 PBS清洗六孔板以除去非贴壁细胞,显微镜拍下24h的伤口距离,计算伤口愈 合情况。The cells were seeded in a six-well plate at a density of 2×10 5 /well, and after the cells adhered, the desired plasmids or siRNA were transfected. When the cell confluence reaches 80-100%, scratch the cell surface with a pipette, wash off the cells dropped on the surface with PBS, replace with a new medium, and take a microscope to take a picture of the wound distance for 0h. After 24h, use PBS The six-well plate was washed to remove non-adherent cells, and the wound distance for 24 hours was photographed with a microscope to calculate the wound healing.
4.Western blot法4. Western blot method
(1)蛋白样本提取和定量(1) Extraction and quantification of protein samples
①提取:收集提前处理过的细胞,2000rpm离心4min,弃上清,将沉淀 转移至EP管中,2000rpm离心4min,弃上清。根据细胞数量加入全细胞裂解 液,每隔10min涡旋一次,①Extraction: Collect the pretreated cells, centrifuge at 2000rpm for 4min, discard the supernatant, transfer the pellet to an EP tube, centrifuge at 2000rpm for 4min, and discard the supernatant. Add whole cell lysate according to the number of cells, vortex every 10min,
进行3次,充分将细胞和裂解液混合均匀,12000rpm,4℃离心30min, 收集上清,上清即蛋白样本。
②定量:BSA(2mg/ml)用去离子水倍半稀释作标准曲线,样本蛋白用去 离子水稀释10倍。不同浓度的BSA溶液和样本蛋白分别取5μl,加入200μl 现配的BCA试剂混合液(A液:B液=50:1),充分混匀后置于37℃,30min, 用酶标仪读取波长为562nm处的吸光度。根据所得的标准曲线的公式,计算出 样本蛋白的浓度,按一定量标定蛋白。②Quantitative: BSA (2mg/ml) was diluted by half and half with deionized water as the standard curve, and the sample protein was diluted 10 times with deionized water. Take 5 μl of BSA solution and sample protein with different concentrations respectively, add 200 μl of BCA reagent mixture (solution A: solution B = 50:1), mix well and place at 37°C for 30 min, read with a microplate reader Absorbance at wavelength 562 nm. According to the formula of the obtained standard curve, the concentration of the sample protein was calculated, and the protein was calibrated according to a certain amount.
③直接loading法③Direct loading method
将预处理的细胞上清弃去,用PBS清洗两遍,根据细胞的密度加入相应量 的2.5×loading直接搅拌收细胞,离心,沸水煮10min使蛋白失活,冷却离心, -20℃保存。The pretreated cell supernatant was discarded, washed twice with PBS, and the corresponding amount of 2.5 × loading was added according to the cell density, and the cells were directly stirred and harvested, centrifuged, boiled in boiling water for 10 min to inactivate the protein, cooled and centrifuged, and stored at -20°C.
(2)蛋白质免疫印迹(2) Western blot
①凝胶电泳:配制500ml的1×Running Buffer,蛋白样本按一定顺序和一 定量上样。先调节电压70V,待样本跑至分离胶时,可将电压增加至110V,样 本离胶底端1cm时,停止电泳。①Gel electrophoresis: Prepare 500ml of 1×Running Buffer, and load protein samples in a certain order and in a certain amount. First adjust the voltage to 70V. When the sample runs to the separation gel, increase the voltage to 110V. When the sample is 1cm away from the bottom of the gel, stop electrophoresis.
②转膜:配制900ml的1×Transfer Buffer,将大小合适的PVDF膜用100 ml的无水乙醇激活1min,加入配制好的Transfer Buffer。根据实验所需的蛋白 大小,将胶切成相应大小,按黑胶白膜方式转膜,330mA横定电流,冰水浴转 膜90min。②Membrane transfer: Prepare 900ml of 1×Transfer Buffer, activate the PVDF membrane of suitable size with 100ml of absolute ethanol for 1min, and add the prepared Transfer Buffer. According to the protein size required for the experiment, cut the glue into the corresponding size, transfer the membrane in the way of black glue and white film, 330mA horizontal current, and transfer the membrane in an ice-water bath for 90min.
③牛奶封闭及孵育一抗:转膜时间到后,将PVDF膜放入5%脱脂牛奶 (T-PBS配制)中室温封闭1h。弃去牛奶,用T-PBS清洗PVDF膜三遍,加入 实验所需的一抗,摇床4℃孵育过夜。③ Milk blocking and primary antibody incubation: After the transfer time is up, put the PVDF membrane in 5% skim milk (prepared with T-PBS) for 1 h at room temperature. Discard the milk, wash the PVDF membrane three times with T-PBS, add the primary antibody required for the experiment, and incubate at 4°C on a shaker overnight.
④孵育二抗:回收一抗,用T-PBS洗条带3次,每次10min。加入一抗对 应的辣根过氧化物酶标记的二抗,室温摇床孵育90min,弃去二抗,T-PBS洗3 次。④Incubate the secondary antibody: Recover the primary antibody and wash the strip with T-
⑤曝光:提前将曝光的东西准备好,条带用ECL显色液(ECL试剂盒中A 液与B液1:1)暗处孵育1min,转移至曝光盒中,暗房中X光胶片压片,短 时间5-20s(根据条带荧光强度而定),长时间30min,将胶片取出,显影液中 放置1.5min,自来水冲洗,置于定影液中1.5min,成像。⑤Exposure: Prepare the exposed things in advance, incubate the strips with ECL chromogenic solution (1:1 solution A and solution B in the ECL kit) in the dark for 1 min, transfer to the exposure box, and press X-ray film in the darkroom , a short time of 5-20s (depending on the fluorescence intensity of the strip), a long time of 30min, the film is taken out, placed in the developer solution for 1.5min, rinsed with tap water, placed in the fixer solution for 1.5min, and imaged.
5.RNA提取和逆转录5. RNA extraction and reverse transcription
将预先处理过的六孔板细胞弃去上清,用预冷的PBS漂洗两次,每孔加入 1ml的Trizol,反复吹打至溶液透明。将细胞转移至1.5ml RNA酶free的EP 管中,加入200μl的氯仿,剧烈震荡30s,4℃静置2min,12000rpm,4℃ 离心10min。吸取450μl的上清至新的EP管中,加入等量异丙醇,轻柔地上下 颠倒5次,冰上静置10min,12000rpm,4℃离心15min。弃去上清,沉淀用 500μl预冷的75%乙醇轻柔清洗,8000rpm,4℃离心5min。重复清洗一遍, 放置通风处,等乙醇挥干后,加入20μl的DEPC水溶解mRNA,测定mRNA 浓度。Discard the supernatant from the pre-treated six-well plate cells, rinse twice with pre-cooled PBS, add 1 ml of Trizol to each well, and repeatedly pipet until the solution is transparent. The cells were transferred to a 1.5 ml RNase-free EP tube, 200 μl of chloroform was added, vigorously shaken for 30 s, stood at 4° C. for 2 min, and centrifuged at 12,000 rpm and 4° C. for 10 min. Pipette 450 μl of the supernatant into a new EP tube, add an equal amount of isopropanol, gently invert 5 times, stand on ice for 10 min, centrifuge at 12000 rpm for 15 min at 4°C. The supernatant was discarded, the pellet was gently washed with 500 μl of pre-chilled 75% ethanol, and centrifuged at 8000 rpm for 5 min at 4°C. Repeat the washing, place in a ventilated place, and after the ethanol evaporates, add 20 μl of DEPC water to dissolve the mRNA, and measure the mRNA concentration.
使用全式金的TransStart Top Green qPCR SuperMix(Lot#40426)试剂盒将mRNA逆转录为互补DNA(cDNA)。The mRNA was reverse transcribed to complementary DNA (cDNA) using the TransStart Top Green qPCR SuperMix (Lot#40426) kit in full gold.
6.qRT-PCR6. qRT-PCR
qRT-PCR体系为10μl,分别是5μl 2×QuantiTect SYBR Green PCR Master Mix,1μl模板cDNA,目标正反向引物各0.6μl,DEPC水2.8μl。其反应条件 为95℃预变性15min,94℃变性30s,63℃退火30s,72℃延伸30s, 39个循环。The qRT-PCR system was 10 μl, 5 μl of 2×QuantiTect SYBR Green PCR Master Mix, 1 μl of template cDNA, 0.6 μl of target forward and reverse primers, and 2.8 μl of DEPC water. The reaction conditions were pre-denaturation at 95 °C for 15 min, denaturation at 94 °C for 30 s, annealing at 63 °C for 30 s, and extension at 72 °C for 30 s, 39 cycles.
7.统计学分析7. Statistical analysis
使用t检验来检验统计分析,p<0.05被认为实验结果具有显著性差异(*P <0.05,**P<0.01和***P<0.001)。每个实验至少重复三次。Statistical analysis was tested using t-test and p<0.05 was considered to be significantly different (*P<0.05, **P<0.01 and ***P<0.001). Each experiment was repeated at least three times.
本领域技术人员在考虑说明书及实践这里公开的内容后,将容易想到本申 请的其它实施方案。本申请旨在涵盖本申请的任何变型、用途或者适应性变化, 这些变型、用途或者适应性变化遵循本申请的一般性原理并包括本申请未公开 的本技术领域中的公知常识或惯用技术手段。说明书和实施例仅被视为示例性 的,本申请的真正范围和精神由权利要求指出。Other embodiments of the present application will readily occur to those skilled in the art upon consideration of the specification and practice of what is disclosed herein. This application is intended to cover any variations, uses or adaptations of this application that follow the general principles of this application and include common knowledge or conventional techniques in the technical field not disclosed in this application . The specification and examples are to be regarded as exemplary only, with the true scope and spirit of the application being indicated by the claims.
应当理解的是,本申请并不局限于上面已经描述并在附图中示出的精确结 构,并且可以在不脱离其范围进行各种修改和改变。本申请的范围仅由所附的 权利要求来限制。It is to be understood that the present application is not limited to the precise structures described above and illustrated in the accompanying drawings, and that various modifications and changes may be made without departing from the scope thereof. The scope of the application is limited only by the appended claims.
序列表sequence listing
<110> 浙大城市学院<110> Zhejiang University City College
<120> 一种用于肿瘤治疗靶点和诊断生物标志物的PDE1A及试剂盒和应用<120> A PDE1A and kit and application for tumor therapy target and diagnostic biomarker
<160> 33<160> 33
<170> SIPOSequenceListing 1.0<170> SIPOSequenceListing 1.0
<210> 1<210> 1
<211> 2136<211> 2136
<212> DNA<212> DNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence
<400> 1<400> 1
agagaggaat tcagcttctt ctggagcgcg aaagtcattc acgtttctct tgtgcataat 60agagaggaat tcagcttctt ctggagcgcg aaagtcattc acgtttctct tgtgcataat 60
agagctcgta aactgtagga attctgatgt gcttcagtgc acagaacagt aacagatgag 120agagctcgta aactgtagga attctgatgt gcttcagtgc acagaacagt aacagatgag 120
ctgcttttgg ggagagcttg agtactcagt cggagcatca tcatggggtc tagtgccaca 180ctgcttttgg ggagagcttg agtactcagt cggagcatca tcatggggtc tagtgccaca 180
gagattgaag aattggaaaa caccactttt aagtatctta caggagaaca gactgaaaaa 240gagattgaag aattggaaaa caccactttt aagtatctta caggagaaca gactgaaaaa 240
atgtggcagc gcctgaaagg aatactaaga tgcttggtga agcagctgga aagaggtgat 300atgtggcagc gcctgaaagg aatactaaga tgcttggtga agcagctgga aagaggtgat 300
gttaacgtcg tcgacttaaa gaagaatatt gaatatgcgg catctgtgct ggaagcagtt 360gttaacgtcg tcgacttaaa gaagaatatt gaatatgcgg catctgtgct ggaagcagtt 360
tatatcgatg aaacaagaag acttctggat actgaagatg agctcagtga cattcagact 420tatatcgatg aaacaagaag acttctggat actgaagatg agctcagtga cattcagact 420
gactcagtcc catctgaagt ccgggactgg ttggcttcta cctttacacg gaaaatgggg 480gactcagtcc catctgaagt ccgggactgg ttggcttcta cctttacacg gaaaatgggg 480
atgacaaaaa agaaacctga ggaaaaacca aaatttcgga gcattgtgca tgctgttcaa 540atgacaaaaa agaaacctga ggaaaaacca aaatttcgga gcattgtgca tgctgttcaa 540
gctggaattt ttgtggaaag aatgtaccga aaaacatatc atatggttgg tttggcatat 600gctggaattt ttgtggaaag aatgtaccga aaaacatatc atatggttgg tttggcatat 600
ccagcagctg tcatcgtaac attaaaggat gttgataaat ggtctttcga tgtatttgcc 660ccagcagctg tcatcgtaac attaaaggat gttgataaat ggtctttcga tgtatttgcc 660
ctaaatgaag caagtggaga gcatagtctg aagtttatga tttatgaact gtttaccaga 720ctaaatgaag caagtggaga gcatagtctg aagtttatga tttatgaact gtttaccaga 720
tatgatctta tcaaccgttt caagattcct gtttcttgcc taatcacctt tgcagaagct 780tatgatctta tcaaccgttt caagattcct gtttcttgcc taatcacctt tgcagaagct 780
ttagaagttg gttacagcaa gtacaaaaat ccatatcaca atttgattca tgcagctgat 840ttagaagttg gttacagcaa gtacaaaaat ccatatcaca atttgattca tgcagctgat 840
gtcactcaaa ctgtgcatta cataatgctt catacaggta tcatgcactg gctcactgaa 900gtcactcaaa ctgtgcatta cataatgctt catacaggta tcatgcactg gctcactgaa 900
ctggaaattt tagcaatggt ctttgctgct gccattcatg attatgagca tacagggaca 960ctggaaattt tagcaatggt ctttgctgct gccattcatg attatgagca tacagggaca 960
acaaacaact ttcacattca gacaaggtca gatgttgcca ttttgtataa tgatcgctct 1020acaaacaact ttcacattca gacaaggtca gatgttgcca ttttgtataa tgatcgctct 1020
gtccttgaga atcaccacgt gagtgcagct tatcgactta tgcaagaaga agaaatgaat 1080gtccttgaga atcaccacgt gagtgcagct tatcgactta tgcaagaaga agaaatgaat 1080
atcttgataa atttatccaa agatgactgg agggatcttc ggaacctagt gattgaaatg 1140atcttgataa atttatccaa agatgactgg agggatcttc ggaacctagt gattgaaatg 1140
gttttatcta cagacatgtc aggtcacttc cagcaaatta aaaatataag aaacagtttg 1200gttttatcta cagacatgtc aggtcacttc cagcaaatta aaaatataag aaacagtttg 1200
cagcagcctg aagggattga cagagccaaa accatgtccc tgattctcca cgcagcagac 1260cagcagcctg aagggattga cagagccaaa accatgtccc tgattctcca cgcagcagac 1260
atcagccacc cagccaaatc ctggaagctg cattatcggt ggaccatggc cctaatggag 1320atcagccacc cagccaaatc ctggaagctg cattatcggt ggaccatggc cctaatggag 1320
gagtttttcc tgcagggaga taaagaagct gaattagggc ttccattttc cccactttgt 1380gagtttttcc tgcagggaga taaagaagct gaattagggc ttccattttc cccactttgt 1380
gatcggaagt caaccatggt ggcccagtca caaataggtt tcatcgattt catagtagag 1440gatcggaagt caaccatggt ggcccagtca caaataggtt tcatcgattt catagtagag 1440
ccaacatttt ctcttctgac agactcaaca gagaaaattg ttattcctct tatagaggaa 1500ccaacatttt ctcttctgac agactcaaca gagaaaattg ttattcctct tatagaggaa 1500
gcctcaaaag ccgaaacttc ttcctatgtg gcaagcagct caaccaccat tgtggggtta 1560gcctcaaaag ccgaaacttc ttcctatgtg gcaagcagct caaccaccat tgtggggtta 1560
cacattgctg atgcactaag acgatcaaat acaaaaggct ccatgagtga tgggtcctat 1620ccattgctg atgcactaag acgatcaaat acaaaaggct ccatgagtga tgggtcctat 1620
tccccagact actcccttgc agcagtggac ctgaagagtt tcaagaacaa cctggtggac 1680tccccagact actcccttgc agcagtggac ctgaagagtt tcaagaacaa cctggtggac 1680
atcattcagc agaacaaaga gaggtggaaa gagttagctg cacaagaagc aagaaccagt 1740atcattcagc agaacaaaga gaggtggaaa gagttagctg cacaagaagc aagaaccagt 1740
tcacagaagt gtgagtttat tcatcagtaa acacctttaa gtaaaacctc gtgcatggtg 1800tcacagaagt gtgagtttat tcatcagtaa acacctttaa gtaaaacctc gtgcatggtg 1800
gcagctctaa tttgaccaaa agacttggag attttgatta tgcttgctgg aaatctaccc 1860gcagctctaa tttgaccaaa agacttggag attttgatta tgcttgctgg aaatctaccc 1860
tgtcctgtgt gagacaggaa atctattttt gcagattgct caataagcat catgagccac 1920tgtcctgtgt gagacaggaa atctattttt gcagattgct caataagcat catgagccac 1920
ataaataaca gctgtaaact ccttaattca ccgggctcaa ctgctaccga acagattcat 1980ataaataaca gctgtaaact ccttaattca ccgggctcaa ctgctaccga acagattcat 1980
ctagtggcta catcagcacc ttgtgctttc agatatctgt ttcaatggca ttttgtggca 2040ctagtggcta catcagcacc ttgtgctttc agatatctgt ttcaatggca ttttgtggca 2040
tttgtcttta ccgagtgcca ataaattttc tttgagcagc taattgctaa ttttgtcatt 2100tttgtcttta ccgagtgcca ataaattttc tttgagcagc taattgctaa ttttgtcatt 2100
tctacaataa agcttggtcc acctgttttc acctta 2136tctacaataa agcttggtcc acctgttttc acctta 2136
<210> 2<210> 2
<211> 18<211> 18
<212> DNA<212> DNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence
<400> 2<400> 2
cagtaacaga tgagctgc 18cagtaacaga tgagctgc 18
<210> 3<210> 3
<211> 16<211> 16
<212> DNA<212> DNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence
<400> 3<400> 3
gtattccttt caggcg 16gtattccttt caggcg 16
<210> 4<210> 4
<211> 20<211> 20
<212> DNA<212> DNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence
<400> 4<400> 4
tagccaactg cgacacattc 20tagccaactg cgacacattc 20
<210> 5<210> 5
<211> 20<211> 20
<212> DNA<212> DNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence
<400> 5<400> 5
cacgaccttg acgttccttt 20
<210> 6<210> 6
<211> 20<211> 20
<212> DNA<212> DNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence
<400> 6<400> 6
tgcaaggata agcggacagg 20
<210> 7<210> 7
<211> 20<211> 20
<212> DNA<212> DNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence
<400> 7<400> 7
cagagatggt gctgacgtgt 20
<210> 8<210> 8
<211> 20<211> 20
<212> DNA<212> DNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence
<400> 8<400> 8
gaacgcattg ccacatacac 20
<210> 9<210> 9
<211> 20<211> 20
<212> DNA<212> DNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence
<400> 9<400> 9
gaattcgggc ttgttgtcat 20
<210> 10<210> 10
<211> 23<211> 23
<212> DNA<212> DNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence
<400> 10<400> 10
ctcctatgag tggaacagga acg 23ctcctatgag tggaacagga acg 23
<210> 11<210> 11
<211> 29<211> 29
<212> DNA<212> DNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence
<400> 11<400> 11
ttggatcaat gtcatattca agtgctgta 29ttggatcaat gtcatattca agtgctgta 29
<210> 12<210> 12
<211> 20<211> 20
<212> DNA<212> DNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence
<400> 12<400> 12
cgtttttcca gaccctggtt 20
<210> 13<210> 13
<211> 18<211> 18
<212> DNA<212> DNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence
<400> 13<400> 13
ctgcagatga gccctcag 18ctgcagatga gccctcag 18
<210> 14<210> 14
<211> 20<211> 20
<212> DNA<212> DNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence
<400> 14<400> 14
cgtttttcca gaccctggtt 20
<210> 15<210> 15
<211> 19<211> 19
<212> DNA<212> DNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence
<400> 15<400> 15
ctgcagatga gccctcaga 19ctgcagatga gccctcaga 19
<210> 16<210> 16
<211> 58<211> 58
<212> DNA<212> DNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence
<400> 16<400> 16
ccgggatgca ctaagacgat caaatctcga gatttgatcg tcttagtgca tctttttg 58ccgggatgca ctaagacgat caaatctcga gatttgatcg tcttagtgca tctttttg 58
<210> 17<210> 17
<211> 58<211> 58
<212> DNA<212> DNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence
<400> 17<400> 17
aattcaaaaa gatgcactaa gacgatcaaa tctcgagatt tgatcgtctt agtgcatc 58aattcaaaaa gatgcactaa gacgatcaaa tctcgagatt tgatcgtctt agtgcatc 58
<210> 18<210> 18
<211> 58<211> 58
<212> DNA<212> DNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence
<400> 18<400> 18
ccgggcctga aaggaatact aagatctcga gatcttagta ttcctttcag gctttttg 58ccgggcctga aaggaatact aagatctcga gatcttagta ttcctttcag gctttttg 58
<210> 19<210> 19
<211> 58<211> 58
<212> DNA<212> DNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence
<400> 19<400> 19
aattcaaaaa gcctgaaagg aatactaaga tctcgagatc ttagtattcc tttcaggc 58aattcaaaaa gcctgaaagg aatactaaga tctcgagatc ttagtattcc tttcaggc 58
<210> 20<210> 20
<211> 21<211> 21
<212> DNA/RNA<212> DNA/RNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence
<400> 20<400> 20
ccacgugagu gcagcuuaut t 21ccacgugagu gcagcuuaut
<210> 21<210> 21
<211> 21<211> 21
<212> DNA/RNA<212> DNA/RNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence
<400> 21<400> 21
auaagcugca cucacguggt t 21auaagcugca cucacguggt
<210> 22<210> 22
<211> 21<211> 21
<212> DNA/RNA<212> DNA/RNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence
<400> 22<400> 22
ccagcagcug ucaucguaat t 21ccagcagcug ucaucguaat
<210> 23<210> 23
<211> 21<211> 21
<212> DNA/RNA<212> DNA/RNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence
<400> 23<400> 23
uuacgaugac agcugcuggt t 21uuacgaugac agcugcuggt
<210> 24<210> 24
<211> 21<211> 21
<212> DNA/RNA<212> DNA/RNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence
<400> 24<400> 24
gcuggaagcc gucuacauat t 21gcuggaagcc gucuacauat
<210> 25<210> 25
<211> 21<211> 21
<212> DNA/RNA<212> DNA/RNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence
<400> 25<400> 25
uauguagacg gcuuccagct t 21uauguagacg gcuuccagct
<210> 26<210> 26
<211> 21<211> 21
<212> DNA/RNA<212> DNA/RNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence
<400> 26<400> 26
gcugcagcua uccaugauut t 21gcugcagcua uccagauut
<210> 27<210> 27
<211> 21<211> 21
<212> DNA/RNA<212> DNA/RNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence
<400> 27<400> 27
aaucauggau agcugcagct t 21aaucauggau agcugcagct
<210> 28<210> 28
<211> 21<211> 21
<212> DNA/RNA<212> DNA/RNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence
<400> 28<400> 28
gcuucagugg uagaucuuat t 21gcuucagugg uagaucuuat
<210> 29<210> 29
<211> 21<211> 21
<212> DNA/RNA<212> DNA/RNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence
<400> 29<400> 29
uaagaucuac cacugaagct t 21uaagaucuac cacugaagct
<210> 30<210> 30
<211> 21<211> 21
<212> DNA/RNA<212> DNA/RNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence
<400> 30<400> 30
ccagcuguua uugaggcaut t 21ccagcuguua uugaggcaut
<210> 31<210> 31
<211> 21<211> 21
<212> DNA/RNA<212> DNA/RNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence
<400> 31<400> 31
augccucaau aacagcuggt t 21augccucaau aacagcuggt
<210> 32<210> 32
<211> 21<211> 21
<212> DNA/RNA<212> DNA/RNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence
<400> 32<400> 32
aaggacguuc ccaauagcca a 21aaggacguuc ccaauagcca a 21
<210> 33<210> 33
<211> 21<211> 21
<212> DNA/RNA<212> DNA/RNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence
<400> 33<400> 33
uuggcuauug ggaacguccu u 21uuggcuauug ggaacguccu
Claims (10)
Priority Applications (1)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| CN202210161665.5A CN114457157B (en) | 2022-02-22 | 2022-02-22 | PDE1A, a tumor treatment target and diagnostic biomarker, and a kit and application thereof |
Applications Claiming Priority (1)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| CN202210161665.5A CN114457157B (en) | 2022-02-22 | 2022-02-22 | PDE1A, a tumor treatment target and diagnostic biomarker, and a kit and application thereof |
Publications (2)
| Publication Number | Publication Date |
|---|---|
| CN114457157A true CN114457157A (en) | 2022-05-10 |
| CN114457157B CN114457157B (en) | 2025-09-12 |
Family
ID=81414724
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| CN202210161665.5A Active CN114457157B (en) | 2022-02-22 | 2022-02-22 | PDE1A, a tumor treatment target and diagnostic biomarker, and a kit and application thereof |
Country Status (1)
| Country | Link |
|---|---|
| CN (1) | CN114457157B (en) |
Citations (5)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN1703522A (en) * | 2002-09-30 | 2005-11-30 | 肿瘤疗法科学股份有限公司 | Methods of diagnosing testicular seminoma |
| US20060019256A1 (en) * | 2003-06-09 | 2006-01-26 | The Regents Of The University Of Michigan | Compositions and methods for treating and diagnosing cancer |
| CN107312824A (en) * | 2016-04-26 | 2017-11-03 | 中国科学院上海药物研究所 | Applications of the PDE3A in anagrelide treatment tumor effect is judged |
| US20190127805A1 (en) * | 2016-03-15 | 2019-05-02 | Almac Diagnostics Limited | Gene signatures for cancer detection and treatment |
| CN112180088A (en) * | 2020-04-30 | 2021-01-05 | 郑州大学第一附属医院 | Test paper strip for early screening of esophageal cancer |
-
2022
- 2022-02-22 CN CN202210161665.5A patent/CN114457157B/en active Active
Patent Citations (5)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN1703522A (en) * | 2002-09-30 | 2005-11-30 | 肿瘤疗法科学股份有限公司 | Methods of diagnosing testicular seminoma |
| US20060019256A1 (en) * | 2003-06-09 | 2006-01-26 | The Regents Of The University Of Michigan | Compositions and methods for treating and diagnosing cancer |
| US20190127805A1 (en) * | 2016-03-15 | 2019-05-02 | Almac Diagnostics Limited | Gene signatures for cancer detection and treatment |
| CN107312824A (en) * | 2016-04-26 | 2017-11-03 | 中国科学院上海药物研究所 | Applications of the PDE3A in anagrelide treatment tumor effect is judged |
| CN112180088A (en) * | 2020-04-30 | 2021-01-05 | 郑州大学第一附属医院 | Test paper strip for early screening of esophageal cancer |
Non-Patent Citations (2)
| Title |
|---|
| ABDURAZZAG ABUSNINA 等: ""Anti-proliferative effect of curcumin on melanoma cells is mediated by PDE1A inhibition that regulates the epigenetic integrator UHRF1"", 《MOL. NUTR. FOOD RES》, vol. 55, 31 December 2011 (2011-12-31), pages 1677 * |
| TAM, C.H.T. 等: ""Homo sapiens phosphodiesterase 1A (PDE1A), transcript variant 2, mRNA,NCBI Reference Sequence: NM_001003683.3"", 《GENBANK》, 9 June 2021 (2021-06-09), pages 1 - 2 * |
Also Published As
| Publication number | Publication date |
|---|---|
| CN114457157B (en) | 2025-09-12 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| Wu et al. | Retracted: effects of Glut1 gene silencing on proliferation, differentiation, and apoptosis of colorectal cancer cells by targeting the TGF‐β/PI3K‐AKT‐mTOR signaling pathway | |
| Li et al. | Circular RNA hsa_circ_0000073 contributes to osteosarcoma cell proliferation, migration, invasion and methotrexate resistance by sponging miR-145-5p and miR-151-3p and upregulating NRAS | |
| Zhao et al. | Long noncoding RNA CASC2 regulates hepatocellular carcinoma cell oncogenesis through miR‐362‐5p/Nf‐κB axis | |
| Gong et al. | LncRNA HAND2‐AS1 represses cervical cancer progression by interaction with transcription factor E2F4 at the promoter of C16orf74 | |
| CN105435228B (en) | New anti-tumor application of arsenic trioxide and anti-tumor preparation | |
| Liu et al. | IGF2BP2 promotes gastric cancer progression by regulating the IGF1R-RhoA-ROCK signaling pathway | |
| Wang et al. | Down‐regulation of long non‐coding RNA HOTAIR inhibits invasion and migration of oesophageal cancer cells via up‐regulation of microRNA‐204 | |
| Gao et al. | BZW2 gene knockdown induces cell growth inhibition, G1 arrest and apoptosis in muscle‐invasive bladder cancers: A microarray pathway analysis | |
| Zhuo et al. | Exosomal linc-FAM138B from cancer cells alleviates hepatocellular carcinoma progression via regulating miR-765 | |
| CN114934048A (en) | circMTMR14 for tumor treatment target and diagnosis biomarker, kit and application | |
| Liu et al. | RETRACTED ARTICLE: Long Noncoding RNA VPS9D1-AS1 Sequesters microRNA-525-5p to Promote the Oncogenicity of Colorectal Cancer Cells by Upregulating HMGA1 | |
| Yu et al. | Clinical significance of lncRNA BCYRN1 in colorectal cancer and its role in cell metastasis. | |
| Huang et al. | Metabolites of intestinal microflora upregulate miR-192-5p to suppress proliferation of colon cancer cells via RhoA-ROCK-LIMK2 pathway. | |
| Feng et al. | Effect of solanum lyratum polysaccharide on malignant behaviors of lung cancer cells by regulating the Circ_UHRF1/miR-513b-5p axis | |
| CN111778333B (en) | Application of reagent for determining EDAR expression level and kit | |
| Gao et al. | miRNA-217 inhibits proliferation of hepatocellular carcinoma cells by regulating KLF5. | |
| Ye et al. | Circular RNA circSEMA5A facilitates colorectal cancer development by regulating microRNA-195-5p to target CCNE1 axis | |
| Zhang et al. | Folic acid supplementation acts as a chemopreventive factor in tumorigenesis of hepatocellular carcinoma by inducing H3K9Me2-dependent transcriptional repression of LCN2 | |
| Wang et al. | SKA2 promotes proliferation and invasion of hepatocellular carcinoma cells via activating the β-catenin signaling pathway. | |
| CN114457157A (en) | PDE1A for tumor treatment target and diagnosis biomarker, kit and application | |
| Wei et al. | MicroRNA-30a-3p inhibits malignant progression of hepatocellular carcinoma through regulating IGF1. | |
| WO2023155857A1 (en) | Circpolk as therapeutic target and diagnostic biomarker for tumors and use thereof | |
| Rao et al. | MicroRNA-150-5p-mediated inhibition of cell proliferation, G1/S transition, and migration in bladder cancer through targeting NEDD4-binding protein 2-like 1 gene | |
| Yan et al. | E2F1-activated NRSN2 promotes esophageal squamous cell carcinoma progression through AKT/mTOR pathway | |
| Chen et al. | Tumor suppressor miR-449a inhibits the development of gastric cancer via down-regulation of SGPL1 |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| PB01 | Publication | ||
| PB01 | Publication | ||
| SE01 | Entry into force of request for substantive examination | ||
| SE01 | Entry into force of request for substantive examination | ||
| GR01 | Patent grant | ||
| GR01 | Patent grant |