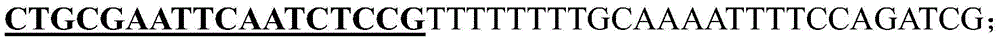CN114438056A - CasF2 protein, CRISPR/Cas gene editing system and its application in plant gene editing - Google Patents
CasF2 protein, CRISPR/Cas gene editing system and its application in plant gene editing Download PDFInfo
- Publication number
- CN114438056A CN114438056A CN202210204370.1A CN202210204370A CN114438056A CN 114438056 A CN114438056 A CN 114438056A CN 202210204370 A CN202210204370 A CN 202210204370A CN 114438056 A CN114438056 A CN 114438056A
- Authority
- CN
- China
- Prior art keywords
- casf2
- gene editing
- protein
- gene
- plant
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Granted
Links
Images
Classifications
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N9/00—Enzymes; Proenzymes; Compositions thereof; Processes for preparing, activating, inhibiting, separating or purifying enzymes
- C12N9/14—Hydrolases (3)
- C12N9/16—Hydrolases (3) acting on ester bonds (3.1)
- C12N9/22—Ribonucleases [RNase]; Deoxyribonucleases [DNase]
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K14/00—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof
- C07K14/415—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from plants
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/11—DNA or RNA fragments; Modified forms thereof; Non-coding nucleic acids having a biological activity
- C12N15/113—Non-coding nucleic acids modulating the expression of genes, e.g. antisense oligonucleotides; Antisense DNA or RNA; Triplex- forming oligonucleotides; Catalytic nucleic acids, e.g. ribozymes; Nucleic acids used in co-suppression or gene silencing
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/63—Introduction of foreign genetic material using vectors; Vectors; Use of hosts therefor; Regulation of expression
- C12N15/79—Vectors or expression systems specially adapted for eukaryotic hosts
- C12N15/82—Vectors or expression systems specially adapted for eukaryotic hosts for plant cells, e.g. plant artificial chromosomes (PACs)
- C12N15/8201—Methods for introducing genetic material into plant cells, e.g. DNA, RNA, stable or transient incorporation, tissue culture methods adapted for transformation
- C12N15/8202—Methods for introducing genetic material into plant cells, e.g. DNA, RNA, stable or transient incorporation, tissue culture methods adapted for transformation by biological means, e.g. cell mediated or natural vector
- C12N15/8205—Agrobacterium mediated transformation
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/63—Introduction of foreign genetic material using vectors; Vectors; Use of hosts therefor; Regulation of expression
- C12N15/79—Vectors or expression systems specially adapted for eukaryotic hosts
- C12N15/82—Vectors or expression systems specially adapted for eukaryotic hosts for plant cells, e.g. plant artificial chromosomes (PACs)
- C12N15/8216—Methods for controlling, regulating or enhancing expression of transgenes in plant cells
- C12N15/8218—Antisense, co-suppression, viral induced gene silencing [VIGS], post-transcriptional induced gene silencing [PTGS]
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N9/00—Enzymes; Proenzymes; Compositions thereof; Processes for preparing, activating, inhibiting, separating or purifying enzymes
- C12N9/0004—Oxidoreductases (1.)
- C12N9/001—Oxidoreductases (1.) acting on the CH-CH group of donors (1.3)
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Y—ENZYMES
- C12Y103/00—Oxidoreductases acting on the CH-CH group of donors (1.3)
- C12Y103/99—Oxidoreductases acting on the CH-CH group of donors (1.3) with other acceptors (1.3.99)
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N2310/00—Structure or type of the nucleic acid
- C12N2310/10—Type of nucleic acid
- C12N2310/20—Type of nucleic acid involving clustered regularly interspaced short palindromic repeats [CRISPR]
Landscapes
- Health & Medical Sciences (AREA)
- Life Sciences & Earth Sciences (AREA)
- Genetics & Genomics (AREA)
- Chemical & Material Sciences (AREA)
- Engineering & Computer Science (AREA)
- Organic Chemistry (AREA)
- Zoology (AREA)
- Bioinformatics & Cheminformatics (AREA)
- Wood Science & Technology (AREA)
- Biomedical Technology (AREA)
- Molecular Biology (AREA)
- General Engineering & Computer Science (AREA)
- Biotechnology (AREA)
- General Health & Medical Sciences (AREA)
- Biochemistry (AREA)
- Microbiology (AREA)
- Biophysics (AREA)
- Medicinal Chemistry (AREA)
- Physics & Mathematics (AREA)
- Plant Pathology (AREA)
- Cell Biology (AREA)
- Virology (AREA)
- Botany (AREA)
- Gastroenterology & Hepatology (AREA)
- Proteomics, Peptides & Aminoacids (AREA)
- Breeding Of Plants And Reproduction By Means Of Culturing (AREA)
Abstract
Description
技术领域technical field
本发明涉及基因编辑技术领域,尤其是涉及CasF2蛋白、CRISPR/Cas基因编辑系统及其在植物基因编辑中的应用。The invention relates to the technical field of gene editing, in particular to a CasF2 protein, a CRISPR/Cas gene editing system and its application in plant gene editing.
背景技术Background technique
基因编辑技术是近年来新兴的一种科学技术,可用于对基因进行靶向沉默和改造,创制非转基因新材料。CRISPR/Cas(clustered regularly interspaced shortpalindromic repeatsand CRISPR-associated protein)系统构建简单、花费少和精确度高,而且能够实现同时多个基因进行编辑,效率高,是一项具有良好应用前景的基因靶向修饰技术。Gene editing technology is an emerging scientific technology in recent years, which can be used for targeted silencing and modification of genes to create new non-transgenic materials. CRISPR/Cas (clustered regularly interspaced shortpalindromic repeats and CRISPR-associated protein) system is simple to construct, low cost and high precision, and can achieve multiple gene editing at the same time with high efficiency. It is a gene targeting modification with good application prospects. technology.
目前,CRISPR/Cas系统中主要是Cas9、Cas12和Cas13。然而,这些Cas基因的分子量都很大,且PAM识别序列范围很窄。因此,开发新的Cas基因应用于植物中的基因编辑迫在眉睫。研发新的基因编辑系统可以为作物基因编辑育种提供有效的工具,以实现人们预期的目标。At present, Cas9, Cas12 and Cas13 are mainly in the CRISPR/Cas system. However, the molecular weights of these Cas genes are large, and the range of PAM recognition sequences is very narrow. Therefore, it is urgent to develop new Cas genes for gene editing in plants. The development of new gene editing systems can provide effective tools for crop gene editing breeding to achieve the desired goals.
Cas基因家族中,目前还没有详细地关于利用CasF2基因构建CRISPR基因编辑载体在作物例如拟南芥、烟草、大豆和玉米中进行应用的相关报道。In the Cas gene family, there is no detailed report on the use of CasF2 gene to construct CRISPR gene editing vector for application in crops such as Arabidopsis, tobacco, soybean and maize.
鉴于此,特提出本发明。In view of this, the present invention is proposed.
发明内容SUMMARY OF THE INVENTION
本发明的目的在于提供CasF2蛋白、CRISPR/Cas基因编辑系统及其在植物基因编辑中的应用;本发明提供了一种新的高效、稳定的基因编辑系统,可使用更短的sgRNA引导CasF2进行靶点序列的编辑,产生DNA序列的插入或缺失。The purpose of the present invention is to provide CasF2 protein, CRISPR/Cas gene editing system and its application in plant gene editing; the present invention provides a new efficient and stable gene editing system, which can use shorter sgRNA to guide CasF2 to perform Editing of target sequences, resulting in insertions or deletions of DNA sequences.
本发明提供的技术方案如下:The technical scheme provided by the present invention is as follows:
在一个方面,本发明提供了CasF2蛋白,所述CasF2蛋白的氨基酸序列如SEQ IDNo.2所示;或者所述CasF2蛋白的氨基酸序列为与SEQ ID NO.2所示的序列相比具有一个或多个氨基酸的置换、缺失或添加,且具有相同或相似生物学功能。In one aspect, the present invention provides a CasF2 protein, the amino acid sequence of the CasF2 protein is shown in SEQ ID No. 2; or the amino acid sequence of the CasF2 protein is compared with the sequence shown in SEQ ID NO. Substitution, deletion or addition of multiple amino acids with the same or similar biological function.
SEQ ID No.2所示的氨基酸序列如下所示:The amino acid sequence shown in SEQ ID No.2 is as follows:
MDYKDHDGDYKDHDIDYKDDDDKMAPKKKRKVGIHGVPAAPKPAVESEFSKVLKKHFPGERFRSSYMKRGGKILAAQGEEAVVAYLQGKSEEEPPNFQPPAKCHVVTKSRDFAEWPIMKASEAIQRYIYALSTTERAACKPGKSSESHAAWFAATGVSNHGYSHVQGLNLIFDHTLGRYDGVLKKVQLRNEKARARLESINASRADEGLPEIKAEEEEVATNETGHLLQPPGINPSFYVYQTISPQAYRPRDEIVLPPEYAGYVRDPNAPIPLGVVRNRCDIQKGCPGYIPEWQREAGTAISPKTGKAVTVPGLSPKKNKRMRRYWRSEKEKAQDALLVTVRIGTDWVVIDVRGLLRNARWRTIAPKDISLNALLDLFTGDPVIDVRRNIVTFTYTLDACGTYARKWTLKGKQTKATLDKLTATQTVALVAIDLGQTNPISAGISRVTQENGALQCEPLDRFTLPDDLLKDISAYRIAWDRNEEELRARSVEALPEAQQAEVRALDGVSKETARTQLCADFGLDPKRLPWDKMSSNTTFISEALLSNSVSRDQVFFTPAPKKGAKKKAPVEVMRKDRTWARAYKPRLSVEAQKLKNEALWALKRTSPEYLKLSRRKEELCRRSINYVIEKTRRRTQCQIVIPVIEDLNVRFFHGSGKRLPGWDNFFTAKKENRWFIQGLHKAFSDLRTHRSFYVFEVRPERTSITCPKCGHCEVGNRDGEAFQCLSCGKTCNADLDVATHNLTQVALTGKTMPKREEPRDAQGTAPARKTKKASKSKAPPAEREDQTPAQEPSQTSKRPAATKKAGQAKKKK。MDYKDHDGDYKDHDIDYKDDDDKMAPKKKRKVGIHGVPAAPKPAVESEFSKVLKKHFPGERFRSSYMKRGGKILAAQGEEAVVAYLQGKSEEEPPNFQPPAKCHVVTKSRDFAEWPIMKASEAIQRYIYALSTTERAACKPGKSSESHAAWFAATGVSNHGYSHVQGLNLIFDHTLGRYDGVLKKVQLRNEKARARLESINASRADEGLPEIKAEEEEVATNETGHLLQPPGINPSFYVYQTISPQAYRPRDEIVLPPEYAGYVRDPNAPIPLGVVRNRCDIQKGCPGYIPEWQREAGTAISPKTGKAVTVPGLSPKKNKRMRRYWRSEKEKAQDALLVTVRIGTDWVVIDVRGLLRNARWRTIAPKDISLNALLDLFTGDPVIDVRRNIVTFTYTLDACGTYARKWTLKGKQTKATLDKLTATQTVALVAIDLGQTNPISAGISRVTQENGALQCEPLDRFTLPDDLLKDISAYRIAWDRNEEELRARSVEALPEAQQAEVRALDGVSKETARTQLCADFGLDPKRLPWDKMSSNTTFISEALLSNSVSRDQVFFTPAPKKGAKKKAPVEVMRKDRTWARAYKPRLSVEAQKLKNEALWALKRTSPEYLKLSRRKEELCRRSINYVIEKTRRRTQCQIVIPVIEDLNVRFFHGSGKRLPGWDNFFTAKKENRWFIQGLHKAFSDLRTHRSFYVFEVRPERTSITCPKCGHCEVGNRDGEAFQCLSCGKTCNADLDVATHNLTQVALTGKTMPKREEPRDAQGTAPARKTKKASKSKAPPAEREDQTPAQEPSQTSKRPAATKKAGQAKKKK。
所述CasF2蛋白为RNA核酸酶,具有以下活性:与靶序列特定位点结合并切割核酸的活性或识别PAM位点以及核酸内切酶活性。在一个实施方案中,所述CasF2蛋白的氨基酸序列可以为与SEQ ID No.2所示的氨基酸序列相比,其序列相似性在95%以上,优选在97%以上,更优选在98%以上,最优选在99%以上,只要与SEQ ID No.2所示的氨基酸序列具有相同的功能即可。The CasF2 protein is an RNA nuclease, and has the following activities: the activity of binding to a specific site of the target sequence and cleaving nucleic acid, or the activity of recognizing the PAM site and the endonuclease activity. In one embodiment, the amino acid sequence of the CasF2 protein can be compared with the amino acid sequence shown in SEQ ID No. 2, and its sequence similarity is more than 95%, preferably more than 97%, more preferably more than 98% , most preferably 99% or more, as long as it has the same function as the amino acid sequence shown in SEQ ID No. 2.
本发明还提供了所述CasF2蛋白的编码基因,所述编码基因的核苷酸序列如以下SEQ ID No.1所示:The present invention also provides the encoding gene of the CasF2 protein, and the nucleotide sequence of the encoding gene is shown in the following SEQ ID No.1:
ATGGATTACAAGGATCACGACGGTGATTACAAGGACCATGATATCGACTACAAGGATGACGATGACAAGATGGCGCCTAAGAAGAAGAGAAAAGTGGGAATTCACGGTGTTCCTGCTGCCCCAAAGCCGGCCGTGGAGTCAGAGTTCTCTAAGGTTCTTAAGAAGCATTTCCCGGGCGAGCGCTTCAGATCTTCCTACATGAAGAGGGGCGGAAAGATCCTGGCGGCTCAGGGAGAGGAGGCTGTGGTTGCCTACCTCCAGGGCAAGAGCGAGGAGGAGCCTCCAAACTTCCAGCCGCCTGCTAAGTGCCACGTCGTGACTAAGTCAAGAGATTTCGCCGAGTGGCCAATTATGAAGGCGTCAGAGGCTATCCAGCGCTACATTTACGCTCTGTCTACTACCGAGAGAGCCGCGTGCAAGCCGGGCAAGTCTTCTGAGAGCCACGCTGCTTGGTTCGCTGCTACAGGCGTCTCAAACCACGGATACTCTCATGTGCAGGGACTCAATCTTATCTTCGATCATACTCTCGGAAGATACGACGGCGTTCTGAAGAAGGTCCAGCTCCGCAACGAGAAGGCTAGGGCCCGCCTTGAGTCAATCAATGCTTCTAGGGCTGACGAGGGCCTGCCTGAGATTAAGGCTGAGGAGGAGGAGGTCGCCACCAACGAGACAGGTCATCTCCTTCAGCCACCGGGCATCAATCCATCCTTCTACGTGTACCAGACAATTAGCCCTCAGGCTTACAGACCAAGGGATGAGATCGTGCTCCCTCCAGAGTACGCGGGTTACGTTAGAGATCCTAACGCTCCTATTCCACTTGGCGTTGTCCGCAATAGATGCGATATCCAGAAGGGCTGCCCAGGATACATTCCGGAGTGGCAGAGGGAGGCTGGAACTGCCATTTCCCCGAAGACTGGAAAGGCCGTCACCGTGCCTGGCCTCAGCCCAAAGAAGAACAAGAGGATGAGAAGGTACTGGCGCTCAGAGAAGGAGAAGGCGCAGGACGCTCTGCTCGTTACAGTCAGAATCGGAACTGATTGGGTGGTTATTGACGTGCGCGGTCTTCTGAGAAACGCGAGGTGGCGCACTATCGCTCCAAAGGATATTTCTCTTAATGCCCTCCTTGACCTGTTCACCGGAGATCCGGTCATCGACGTGCGCAGAAATATTGTTACCTTCACATACACTCTCGATGCTTGCGGCACCTACGCCAGGAAGTGGACACTTAAGGGAAAGCAGACTAAGGCTACCCTTGATAAGCTGACCGCCACACAGACTGTTGCCCTTGTCGCGATCGACCTGGGCCAGACCAACCCTATCTCCGCCGGAATTAGCCGCGTCACACAGGAGAATGGTGCGCTCCAGTGCGAGCCACTTGATAGATTCACTCTGCCGGATGACCTGCTCAAGGATATCTCTGCGTACAGAATTGCTTGGGACAGGAACGAGGAGGAGCTTAGAGCGAGGTCCGTTGAGGCTCTGCCAGAGGCTCAGCAGGCCGAAGTTAGGGCTCTCGATGGCGTCAGCAAGGAGACAGCCAGGACTCAGCTCTGCGCGGATTTCGGACTCGACCCGAAGAGGCTTCCTTGGGATAAGATGTCTTCCAACACAACTTTCATCTCAGAGGCTCTTCTGAGCAATTCAGTCTCTCGCGACCAGGTGTTCTTCACCCCGGCCCCTAAGAAGGGTGCTAAGAAGAAGGCGCCTGTGGAGGTTATGCGCAAGGATAGAACATGGGCGCGCGCTTACAAGCCAAGACTCTCTGTGGAGGCCCAGAAGCTTAAGAATGAGGCGCTCTGGGCTCTTAAGAGAACCTCCCCAGAGTACCTGAAGCTCAGCAGGCGCAAGGAGGAGCTTTGCAGAAGGTCTATCAACTACGTTATTGAGAAGACACGCAGAAGGACTCAGTGCCAGATCGTGATTCCGGTTATTGAGGATCTTAATGTCAGATTCTTCCACGGTTCCGGCAAGAGGCTGCCTGGTTGGGACAACTTCTTCACTGCCAAGAAGGAGAATAGATGGTTCATCCAGGGCCTGCACAAGGCGTTCTCAGATCTCAGGACCCATCGCTCTTTCTACGTCTTCGAAGTGAGGCCTGAGAGGACCTCTATTACATGCCCAAAGTGCGGACACTGCGAGGTTGGAAACAGGGACGGAGAGGCTTTCCAGTGCCTCTCCTGCGGAAAGACCTGCAACGCCGATCTTGACGTTGCGACCCATAATCTGACACAGGTCGCCCTCACTGGAAAGACCATGCCGAAGAGGGAGGAGCCTAGGGATGCTCAGGGTACTGCTCCAGCTAGGAAGACCAAGAAGGCTTCCAAGAGCAAGGCTCCACCTGCTGAGAGGGAGGACCAGACACCAGCTCAGGAGCCGTCCCAGACAAGCAAGAGGCCTGCCGCGACTAAGAAGGCTGGACAGGCTAAGAAGAAGAAGTGA。ATGGATTACAAGGATCACGACGGTGATTACAAGGACCATGATATCGACTACAAGGATGACGATGACAAGATGGCGCCTAAGAAGAAGAGAAAAGTGGGAATTCACGGTGTTCCTGCTGCCCCAAAGCCGGCCGTGGAGTCAGAGTTCTCTAAGGTTCTTAAGAAGCATTTCCCGGGCGAGCGCTTCAGATCTTCCTACATGAAGAGGGGCGGAAAGATCCTGGCGGCTCAGGGAGAGGAGGCTGTGGTTGCCTACCTCCAGGGCAAGAGCGAGGAGGAGCCTCCAAACTTCCAGCCGCCTGCTAAGTGCCACGTCGTGACTAAGTCAAGAGATTTCGCCGAGTGGCCAATTATGAAGGCGTCAGAGGCTATCCAGCGCTACATTTACGCTCTGTCTACTACCGAGAGAGCCGCGTGCAAGCCGGGCAAGTCTTCTGAGAGCCACGCTGCTTGGTTCGCTGCTACAGGCGTCTCAAACCACGGATACTCTCATGTGCAGGGACTCAATCTTATCTTCGATCATACTCTCGGAAGATACGACGGCGTTCTGAAGAAGGTCCAGCTCCGCAACGAGAAGGCTAGGGCCCGCCTTGAGTCAATCAATGCTTCTAGGGCTGACGAGGGCCTGCCTGAGATTAAGGCTGAGGAGGAGGAGGTCGCCACCAACGAGACAGGTCATCTCCTTCAGCCACCGGGCATCAATCCATCCTTCTACGTGTACCAGACAATTAGCCCTCAGGCTTACAGACCAAGGGATGAGATCGTGCTCCCTCCAGAGTACGCGGGTTACGTTAGAGATCCTAACGCTCCTATTCCACTTGGCGTTGTCCGCAATAGATGCGATATCCAGAAGGGCTGCCCAGGATACATTCCGGAGTGGCAGAGGGAGGCTGGAACTGCCATTTCCCCGAAGACTGGAAAGGCCGTCACCGTGCCTGGCCTCAGCCCAAAGAAGAACAAGAGGATGAGAAGGTACTGGCGCTCAGAGAAGGAGAAGGCGC AGGACGCTCTGCTCGTTACAGTCAGAATCGGAACTGATTGGGTGGTTATTGACGTGCGCGGTCTTCTGAGAAACGCGAGGTGGCGCACTATCGCTCCAAAGGATATTTCTCTTAATGCCCTCCTTGACCTGTTCACCGGAGATCCGGTCATCGACGTGCGCAGAAATATTGTTACCTTCACATACACTCTCGATGCTTGCGGCACCTACGCCAGGAAGTGGACACTTAAGGGAAAGCAGACTAAGGCTACCCTTGATAAGCTGACCGCCACACAGACTGTTGCCCTTGTCGCGATCGACCTGGGCCAGACCAACCCTATCTCCGCCGGAATTAGCCGCGTCACACAGGAGAATGGTGCGCTCCAGTGCGAGCCACTTGATAGATTCACTCTGCCGGATGACCTGCTCAAGGATATCTCTGCGTACAGAATTGCTTGGGACAGGAACGAGGAGGAGCTTAGAGCGAGGTCCGTTGAGGCTCTGCCAGAGGCTCAGCAGGCCGAAGTTAGGGCTCTCGATGGCGTCAGCAAGGAGACAGCCAGGACTCAGCTCTGCGCGGATTTCGGACTCGACCCGAAGAGGCTTCCTTGGGATAAGATGTCTTCCAACACAACTTTCATCTCAGAGGCTCTTCTGAGCAATTCAGTCTCTCGCGACCAGGTGTTCTTCACCCCGGCCCCTAAGAAGGGTGCTAAGAAGAAGGCGCCTGTGGAGGTTATGCGCAAGGATAGAACATGGGCGCGCGCTTACAAGCCAAGACTCTCTGTGGAGGCCCAGAAGCTTAAGAATGAGGCGCTCTGGGCTCTTAAGAGAACCTCCCCAGAGTACCTGAAGCTCAGCAGGCGCAAGGAGGAGCTTTGCAGAAGGTCTATCAACTACGTTATTGAGAAGACACGCAGAAGGACTCAGTGCCAGATCGTGATTCCGGTTATTGAGGATCTTAATGTCAGATTCTTCCACGGTTCCGGCAAGAGGCTGCCTGGTTGGGACAACTTCTTCAC TGCCAAGAAGGAGAATAGATGGTTCATCCAGGGCCTGCACAAGGCGTTCTCAGATCTCAGGACCCATCGCTCTTTCTACGTCTTCGAAGTGAGGCCTGAGAGGACCTCTATTACATGCCCAAAGTGCGGACACTGCGAGGTTGGAAACAGGGACGGAGAGGCTTTCCAGTGCCTCTCCTGCGGAAAGACCTGCAACGCCGATCTTGACGTTGCGACCCATAATCTGACACAGGTCGCCCTCACTGGAAAGACCATGCCGAAGAGGGAGGAGCCTAGGGATGCTCAGGGTACTGCTCCAGCTAGGAAGACCAAGAAGGCTTCCAAGAGCAAGGCTCCACCTGCTGAGAGGGAGGACCAGACACCAGCTCAGGAGCCGTCCCAGACAAGCAAGAGGCCTGCCGCGACTAAGAAGGCTGGACAGGCTAAGAAGAAGAAGTGA。
CasF2基因是Cas12j类型的Cas基因,具有长片段DNA序列删除的潜力。CasF2基因的PAM识别序列为:5’-TBN-3’(B=G/T/C),并且CasF2对识别的靶序列位点特异性很高。CasF2编辑系统的靶序列为18bp。CasF2在基因组DNA上可编辑的位点更多和具有更加广泛的潜在目标靶点。The CasF2 gene is a Cas12j-type Cas gene with the potential to delete long DNA sequences. The PAM recognition sequence of CasF2 gene is: 5'-TBN-3' (B=G/T/C), and CasF2 is highly specific to the recognized target sequence site. The target sequence of the CasF2 editing system is 18 bp. CasF2 has more editable sites on genomic DNA and has a wider range of potential targets.
本发明还可包括与SEQ ID No.1所示的CasF2蛋白的核苷酸序列反向互补的核苷酸序列。The present invention may also include a nucleotide sequence that is reverse complementary to the nucleotide sequence of the CasF2 protein shown in SEQ ID No. 1.
在另一个方面,本发明还提供了一种表达载体,所述表达载体包含前述的核苷酸序列。表达载体可以在目标细胞中表达CasF2蛋白,从而在目标细胞中进行相应的基因编辑。常用的载体可以是例如pBI121载体和PCAMBIA1300载体等。In another aspect, the present invention also provides an expression vector comprising the aforementioned nucleotide sequence. The expression vector can express the CasF2 protein in the target cells, so that the corresponding gene editing can be performed in the target cells. Commonly used vectors may be, for example, the pBI121 vector, the PCAMBIA1300 vector, and the like.
本发明也可包括含有所述表达载体的重组细胞,优选地,所述重组细胞为真核细胞,更优选地,所述重组细胞为植物细胞。The present invention may also include recombinant cells containing the expression vector, preferably, the recombinant cells are eukaryotic cells, more preferably, the recombinant cells are plant cells.
在另一个方面,本发明还提供了所述CasF2蛋白、所述CasF2蛋白的编码基因或权所述表达载体在基因编辑中的应用。In another aspect, the present invention also provides the application of the CasF2 protein, the gene encoding the CasF2 protein or the expression vector in gene editing.
在另一个方面,本发明还提供了一种CRISPR/Cas基因编辑系统,所述基因编辑系统包含前述的CasF2蛋白或表达所述CasF2蛋白的质粒。In another aspect, the present invention also provides a CRISPR/Cas gene editing system, the gene editing system comprising the aforementioned CasF2 protein or a plasmid expressing the CasF2 protein.
在一个实施方案中,所述基因编辑系统还包含sgRNA或表达sgRNA的质粒,例如所述表达质粒含有所述sgRNA的编码基因。In one embodiment, the gene editing system further comprises an sgRNA or a plasmid expressing the sgRNA, eg, the expression plasmid contains the gene encoding the sgRNA.
上述两个sgRNA都能用于引导CasF2蛋白在受体材料中靶基因序列位置进行基因编辑,当使用SEQ ID No.3所示的sgRNA3序列时,CasF2蛋白具有sgRNA序列加工的功能,会把sgRNA3剪切成dgRNA1序列,从而引导CasF2在靶点位点处进行基因编辑。所述sgRNA可以为一个或多个sgRNA,用于靶向一个或多个靶基因。所述sgRNA包括含有其对应识别区序列的核苷酸。The above two sgRNAs can be used to guide the CasF2 protein to perform gene editing at the target gene sequence position in the receptor material. When the sgRNA3 sequence shown in SEQ ID No. 3 is used, the CasF2 protein has the function of sgRNA sequence processing, and the sgRNA3 Cut into a dgRNA1 sequence to guide CasF2 for gene editing at the target site. The sgRNA can be one or more sgRNAs for targeting one or more target genes. The sgRNA includes nucleotides containing its corresponding recognition region sequence.
在另一个方面,本发明提供了所述CRISPR/Cas基因编辑系统在植物基因编辑中的应用;优选地,所述植物为单子叶植物或双子叶植物;更优选地,所述植物选自拟南芥、烟草、大豆和玉米中的任一种。In another aspect, the present invention provides the application of the CRISPR/Cas gene editing system in plant gene editing; preferably, the plant is a monocotyledonous plant or a dicotyledonous plant; more preferably, the plant is selected from the group consisting of Any of Arabidopsis, tobacco, soybean, and corn.
在一个实施方案中,所述应用包括以下步骤:In one embodiment, the application comprises the steps of:
(a)根据所述CasF2蛋白的PAM识别位点确定待编辑基因的sgRNA;(a) determining the sgRNA of the gene to be edited according to the PAM recognition site of the CasF2 protein;
(b)将所述CasF2蛋白的基因序列和相应的sgRNA克隆到表达载体中;(b) cloning the gene sequence of the CasF2 protein and the corresponding sgRNA into an expression vector;
(c)将所述表达载体转化到农杆菌中,通过农杆菌介导的遗传转化方法,将所述表达载体导入目标植物,获得定向基因编辑植株;(c) transforming the expression vector into Agrobacterium, and introducing the expression vector into a target plant through an Agrobacterium-mediated genetic transformation method to obtain a targeted gene editing plant;
优选地,在所述遗传转化中,使用MgCl2激活基因编辑活性。Preferably, in the genetic transformation, MgCl2 is used to activate gene editing activity.
CasF2基因的编辑活性需要一定浓度的Mg2+才能激活。CasF2在植物体内存在时,没有额外添加一定浓度的Mg2+,CasF2不对细胞中的靶位点进行编辑。The editing activity of the CasF2 gene requires a certain concentration of Mg 2+ to activate. When CasF2 exists in plants, without adding a certain concentration of Mg 2+ , CasF2 does not edit the target site in cells.
在一个实施方案中,所述植物为拟南芥,所述sgRNA对应的核苷酸序列(sgRNA3序列)为SEQ ID No.3所示。具体地,SEQ ID No.3:In one embodiment, the plant is Arabidopsis thaliana, and the nucleotide sequence (sgRNA3 sequence) corresponding to the sgRNA is shown in SEQ ID No.3. Specifically, SEQ ID No. 3:
TAATGTCGGAACGCTCAACGATTGCCCCTCACGAGGGGAC-N18(靶点序列)。TAATGTCGGAACGCTCAACGATTGCCCCCTCACGAGGGGAC-N 18 (target sequence).
在一个实施方案中,所述植物为烟草,所述sgRNA对应的核苷酸序列(sgRNA1序列)为如SEQ ID No.4所示。具体地,SEQ ID No.4:In one embodiment, the plant is tobacco, and the nucleotide sequence (sgRNA1 sequence) corresponding to the sgRNA is as shown in SEQ ID No.4. Specifically, SEQ ID No. 4:
CGCTCAACGATTGCCCCTCACGAGGGGAC-N18(靶点序列)。CGCTCAACGATTGCCCCCTCACGAGGGGAC-N 18 (target sequence).
在一个实施方案中,本发明提供了植物拟南芥BRI1基因的sgRNA靶点序列;所述sgRNA靶点序列的核苷酸序列如SEQ ID No.5所示。In one embodiment, the present invention provides the sgRNA target sequence of plant Arabidopsis thaliana BRI1 gene; the nucleotide sequence of the sgRNA target sequence is shown in SEQ ID No.5.
具体地,SEQ ID No.5:CTGCGAATTCAATCTCCG。Specifically, SEQ ID No. 5: CTGCGAATTCAATCTCCG.
在一个实施方案中,本发明提供了植物烟草PDS基因的sgRNA靶点序列;所述sgRNA靶点序列的核苷酸序列如SEQ ID No.6所示。In one embodiment, the present invention provides the sgRNA target sequence of the plant tobacco PDS gene; the nucleotide sequence of the sgRNA target sequence is shown in SEQ ID No.6.
具体地,SEQ ID No.6:GTAGTAGCGACTCCATGG。Specifically, SEQ ID No. 6: GTAGTAGCGACTCCATGG.
基于本发明所述CRISPR/Cas基因编辑系统在植物基因编辑中的应用,本发明同时也提供了一种利用本发明所述的基因编辑载体系统进行对植物进行基因编辑的方法。Based on the application of the CRISPR/Cas gene editing system of the present invention in plant gene editing, the present invention also provides a method for gene editing of plants using the gene editing vector system of the present invention.
在本发明中,CRISPR/Cas系统中的CasF2蛋白以及sgRNA,二者可以在同一表达载体中也可以在不同的表达载体中。In the present invention, the CasF2 protein and sgRNA in the CRISPR/Cas system can be in the same expression vector or in different expression vectors.
在本发明中,简写“CRISPR”或者“crispr”均指规律成簇间隔短回文重复,即Clustered regularly interspaced short palindromic repeats的首字母缩写。在核苷酸序列中,表示碱基时,如无特别说明,字母N所代表的碱基具有本领域通常的含义,即N代表随机或者任意碱基A、T、C或者G。In the present invention, the abbreviation "CRISPR" or "crispr" both refer to the acronym of Clustered regularly interspaced short palindromic repeats, namely Clustered regularly interspaced short palindromic repeats. In a nucleotide sequence, when expressing a base, unless otherwise specified, the base represented by the letter N has the usual meaning in the art, that is, N represents random or any base A, T, C or G.
本文中使用的“含有”、“具有”或“包括”包括了“包含”、“主要由……构成”、“基本上由……构成”和“由……构成”。As used herein, "comprising," "having," or "including" includes "comprising," "consisting essentially of," "consisting essentially of," and "consisting of."
有益效果:Beneficial effects:
本发明提供了一种新的CasF2蛋白和一种高效、稳定的Cas介导的基因编辑载体,可以在植物细胞中稳定、高效的表达,从而更好的调控靶基因的表达。CasF2蛋白氨基酸数目少,CasF2体积小为传递进入植物细胞中提供了优势(便于包装和递送),且CasF2使用单一活性位点进行crRNA处理和对DNA序列的切割。CasF2具有广泛的PAM识别位点,可以构建多个基因编辑位点载体,实现对更多的基因进行敲除。本发明可用于拟南芥、烟草、大豆和玉米中DNA序列的基因编辑。The present invention provides a novel CasF2 protein and an efficient and stable Cas-mediated gene editing vector, which can be stably and efficiently expressed in plant cells, thereby better regulating the expression of target genes. CasF2 protein has a small number of amino acids, the small size of CasF2 provides advantages for delivery into plant cells (easy packaging and delivery), and CasF2 uses a single active site for crRNA processing and cleavage of DNA sequences. CasF2 has a wide range of PAM recognition sites, and can construct multiple gene editing site vectors to achieve knockout of more genes. The present invention can be used for gene editing of DNA sequences in Arabidopsis, tobacco, soybean and maize.
附图说明Description of drawings
为了更清楚地说明本发明具体实施方式或现有技术中的技术方案,下面将对具体实施方式或现有技术描述中所需要使用的附图作简单地介绍,显而易见地,下面描述中的附图是本发明的一些实施方式,对于本领域普通技术人员来讲,在不付出创造性劳动的前提下,还可以根据这些附图获得其他的附图。In order to more clearly illustrate the specific embodiments of the present invention or the technical solutions in the prior art, the following briefly introduces the accompanying drawings required in the description of the specific embodiments or the prior art. Obviously, the accompanying drawings in the following description The drawings are some embodiments of the present invention. For those of ordinary skill in the art, other drawings can also be obtained based on these drawings without creative efforts.
图1为本发明实施例提供的PCR测序结果;Fig. 1 is the PCR sequencing result provided in the embodiment of the present invention;
图2为本发明实施例提供通过CRISPR/CasF2基因编辑系统获得的拟南芥BRI1基因编辑植株的表型图;2 is a phenotypic diagram of an Arabidopsis BRI1 gene-edited plant obtained by the CRISPR/CasF2 gene-editing system according to an embodiment of the present invention;
图3为本发明实施例提供的sgRNA3 T质粒载体的示意图;Fig. 3 is the schematic diagram of the sgRNA3 T plasmid vector provided in the embodiment of the present invention;
图4为本发明实施例提供的pHCasF2载体的示意图;Fig. 4 is the schematic diagram of pHCasF2 carrier provided in the embodiment of the present invention;
图5为本发明实施例提供的sgRNA1T质粒的示意图;Fig. 5 is the schematic diagram of the sgRNA1T plasmid provided in the embodiment of the present invention;
图6为本发明实施例提供的烟草组培结果图;Fig. 6 is the tobacco tissue culture result graph that the embodiment of the present invention provides;
图7为本发明实施例提供的烟草PDS靶点序列测序图。FIG. 7 is a sequence diagram of the tobacco PDS target sequence provided by the embodiment of the present invention.
具体实施方式Detailed ways
下面将结合实施例对本发明的技术方案进行清楚、完整地描述,显然,所描述的实施例是本发明一部分实施例,而不是全部的实施例。基于本发明中的实施例,本领域普通技术人员在没有做出创造性劳动前提下所获得的所有其他实施例,都属于本发明保护的范围。The technical solutions of the present invention will be clearly and completely described below with reference to the embodiments. Obviously, the described embodiments are part of the embodiments of the present invention, but not all of the embodiments. Based on the embodiments of the present invention, all other embodiments obtained by those of ordinary skill in the art without creative efforts shall fall within the protection scope of the present invention.
实施例1Example 1
1.拟南芥靶基因BRI1打靶策略:1. Targeting strategy of Arabidopsis target gene BRI1:
以拟南芥(Arabidopsis thaliana)油菜素内酯受体BRI1为目的基因,利用CRISPR/CasF2定向编辑拟南芥BRI1。Using the brassinosteroid receptor BRI1 of Arabidopsis thaliana as the target gene, CRISPR/CasF2 was used to directionally edit Arabidopsis BRI1.
CasF2/sgRNA设计:CasF2/sgRNA design:
根据CasF2识别的PAM序列,选择拟南芥BRI1的sgRNA靶点的核苷酸序列(靶点序列SEQ ID No.5)(5’-3’):CTGCGAATTCAATCTCCG。According to the PAM sequence recognized by CasF2, the nucleotide sequence of the sgRNA target of Arabidopsis BRI1 was selected (target sequence SEQ ID No. 5) (5'-3'): CTGCGAATTCAATCTCCG.
2.拟南芥BRI1基因的编辑2. Editing of the Arabidopsis BRI1 gene
构建拟南芥BRI1基因的sgRNA,选择拟南芥AtU6-26启动子驱动sgRNA序列的表达;同时,在sgRNA序列后面添加AtU6-26终止子序列。以拟南芥U6启动子驱动sgRNA的CRISPR/CasF2基因编辑载体。The sgRNA of the Arabidopsis BRI1 gene was constructed, and the Arabidopsis AtU6-26 promoter was selected to drive the expression of the sgRNA sequence; at the same time, the AtU6-26 terminator sequence was added after the sgRNA sequence. CRISPR/CasF2 gene editing vector driven by sgRNA driven by Arabidopsis U6 promoter.
利用KOD-plus neo,以sgRNA3 T质粒(图3为质粒载体的示意图)作为模板扩增打靶sgRNA序列。第1轮PCR,扩增体系(20μL):KOD-plus neo was used to amplify the targeted sgRNA sequence with the sgRNA3 T plasmid (Fig. 3 is a schematic diagram of the plasmid vector) as a template. The first round of PCR, amplification system (20μL):
引物对:U6-26p-F+F2BRI1-gRNA-R扩增1管,引物对:F2BRI1-U6-26t-F+U6-26t-R扩增1管。Primer pair: U6-26p-F+F2BRI1-gRNA-R to amplify 1 tube, primer pair: F2BRI1-U6-26t-F+U6-26t-R to amplify 1 tube.
U6-26p-F(SEQ ID No.7):U6-26p-F (SEQ ID No. 7):
TTCAGAggtctcTtagtCGACTTGCCTTCCGCACAATAC;TTCAGAggtctcTtagtCGACTTGCCTTCCGCACAATAC;
U6-26t-R(SEQ ID No.8):U6-26t-R (SEQ ID No. 8):
AGCGTGggtctcGcgccTATTGGTTTATCTCATCGGAAC;AGCGTGggtctcGcgccTATTGGTTTATCTCATCGGAAC;
F2BRI1-U6-26t-F(SEQ ID No.9):F2BRI1-U6-26t-F (SEQ ID No. 9):
(其中,加粗下划线部分为靶点序列)(The bold underlined part is the target sequence)
F2BRI1-gRNA-R(SEQ ID No.10):F2BRI1-gRNA-R (SEQ ID No. 10):
CGGAGATTGAATTCGCAGGTCCCCTCGTGAGGGGCAATC。CGGAGATTGAATTCGCAGGTCCCCTCGTGAGGGGCAATC.
PCR反应程序:PCR reaction program:
PCR扩增结束后,分别取2管的PCR产物各2μL,作为进行第2轮PCR的模板。After PCR amplification, 2 μL of PCR products in 2 tubes were taken as templates for the second round of PCR.
扩增体系:Amplification system:
反应程序:Reaction program:
第2轮PCR结束后,取PCR产物1μL作为第3轮PCR的模板。反应体系和扩增体系同第1轮。扩增结束后,进行琼脂糖凝胶电泳。After the second round of PCR, take 1 μL of the PCR product as the template for the third round of PCR. The reaction system and amplification system were the same as in the first round. After amplification, agarose gel electrophoresis was performed.
PCR产物进行胶回收后,与pHCasF2载体(图4为载体示意图)连接。After the PCR product was recovered by gel, it was ligated with the pHCasF2 vector (Fig. 4 is a schematic diagram of the vector).
连接体系:Connection system:
连接反应:37℃5min,10℃5min;20℃5min;15个循环,最后37℃5min。Ligation reaction: 37°C for 5 min, 10°C for 5 min; 20°C for 5 min; 15 cycles, and finally 37°C for 5 min.
连接产物转化E.coli DH5α感受态,感受态涂布在LB(Kan+)平板上。从平板上挑选菌落送公司测序验证。The ligation product was transformed into E. coli DH5α competent, and the competent was spread on LB (Kan + ) plates. Pick colonies from the plate and send them to the company for sequencing verification.
构建的BRI1基因打靶表达载体pHCasF2-BRI1转化农杆菌GV3101。The constructed BRI1 gene targeting expression vector pHCasF2-BRI1 was transformed into Agrobacterium GV3101.
挑取农杆菌对处于盛花期的拟南芥进行侵染。农杆菌法侵染拟南芥的步骤如下:农杆菌菌种吸取20μL加于3mL酵母液体培养基(Yeast Extract Broth,YEB)(kan+、Rif+)中;28℃,200rpm,摇24h;Agrobacterium was picked to infect Arabidopsis thaliana in full bloom. The steps of infecting Arabidopsis thaliana by Agrobacterium method are as follows: 20 μL of Agrobacterium strain is drawn and added to 3 mL of yeast liquid medium (Yeast Extract Broth, YEB) (kan + , Rif + ); 28°C, 200rpm, shake for 24h;
取摇好的菌,12,000rpm,1min离心收集菌株,去上清,将沉淀用1mL5%蔗糖1/2MS溶液重悬,再次收集菌株;最后仍用1mL 5%蔗糖1/2MS溶液重悬,同时加入0.2μL Silwet77和终浓度为10mM MgCl2,充分混匀后,即可侵染拟南芥。Take the shaken bacteria, centrifuge at 12,000 rpm for 1 min to collect the strains, remove the supernatant, resuspend the pellet with 1 mL of 5
筛选抗性苗并进行测序:Screen and sequence resistant shoots:
待拟南芥种子成熟后,收获种子。种子播种在含有50mg/L潮霉素的1/2MS培养基平板上,挑取抗性苗。After the Arabidopsis seeds are mature, the seeds are harvested. Seeds were sown on 1/2 MS medium plates containing 50 mg/L hygromycin, and resistant shoots were picked.
PCR扩增打靶位点附近的DNA序列,PCR产物送公司测序。对BRI1基因被编辑的拟南芥植株进行拍照。PCR amplifies the DNA sequence near the target site, and the PCR product is sent to the company for sequencing. Photographs of Arabidopsis plants with the BRI1 gene edited.
用于靶点序列附近DNA扩增的PCR扩增引物:PCR amplification primers for DNA amplification near the target sequence:
F2BRI1crispr-identify-F(SEQ ID No.11):F2BRI1crispr-identify-F (SEQ ID No. 11):
CTGGCTTCAAGTGCTCTGCTTC;CTGGCTTCAAGTGCTCTGCTTC;
F2BRI1crispr-identify-R(SEQ ID No.12):F2BRI1crispr-identify-R (SEQ ID No. 12):
GTAATTCGCCGGAAAACTCG。GTAATTCGCCGGAAAACTCG.
扩增体系同第1轮PCR。The amplification system was the same as the first round PCR.
实验结果分析:Analysis of results:
(1)在侵染拟南芥的侵染液中不加入终浓度为10mM MgCl2没有发现BRI1基因被编辑的植株,只有在加入了10mM MgCl2的侵染液侵染的拟南芥植株,才出现BRI1基因被编辑的植株。(1) No BRI1 gene-edited plants were found in the infection solution of Arabidopsis thaliana without adding a final concentration of 10 mM MgCl 2 , but only Arabidopsis plants infected with the infection solution with 10 mM MgCl 2 added, Only plants with the BRI1 gene edited have emerged.
(2)扩增BRI1靶点附近的DNA序列,通过测序发现,靶点出现了套峰(图1中示出);并且拟南芥植株出现了不育表型,与报道的文献结果一致(图2中示出)。(2) Amplify the DNA sequence near the BRI1 target. It is found by sequencing that the target has a set of peaks (shown in Figure 1); and the Arabidopsis plant has a sterile phenotype, which is consistent with the reported literature results ( shown in Figure 2).
结果表明:CasF2蛋白在拟南芥中进行基因编辑需要MgCl2的存在,并且能对拟南芥基因进行基因编辑,产生DNA序列的插入、缺失。The results showed that the gene editing of the CasF2 protein in Arabidopsis requires the presence of MgCl 2 , and the gene editing of the Arabidopsis thaliana gene can produce insertion and deletion of DNA sequences.
实施例2Example 2
1.本氏烟PDS基因打靶策略:1. N. benthamiana PDS gene targeting strategy:
以本氏烟草(Nicotiana benthamiana)八氢番茄红素脱氢酶PDS为目的基因,利用CRISPR/CasF2定向编辑本氏烟PDS。The phytoene dehydrogenase PDS of Nicotiana benthamiana was used as the target gene, and the PDS of Nicotiana benthamiana was edited using CRISPR/CasF2.
CasF2/sgRNA设计:CasF2/sgRNA design:
根据CasF2识别的PAM序列,选择本氏烟PDS的sgRNA靶点的核苷酸序列(靶点序列SEQ ID No.6)(5’-3’):GTAGTAGCGACTCCATGG。According to the PAM sequence recognized by CasF2, the nucleotide sequence of the sgRNA target of N. benthamiana PDS was selected (target sequence SEQ ID No. 6) (5'-3'): GTAGTAGCGACTCCATGG.
2.拟南芥BRI1基因的编辑2. Editing of the Arabidopsis BRI1 gene
本氏烟PDS基因的编辑Editing of N. benthamiana PDS gene
构建烟草PDS基因的sgRNA,构建方法同拟南芥一样,把拟南芥BRI1靶点引物更换成烟草的PDS靶点引物,扩增sgRNA序列的模板换成sgRNA1T质粒(图5为载体示意图)。烟草PDS基因的靶点引物如下:To construct the sgRNA of the tobacco PDS gene, the construction method is the same as that of Arabidopsis. The Arabidopsis thaliana BRI1 target primer is replaced with the tobacco PDS target primer, and the template for amplifying the sgRNA sequence is replaced by the sgRNA1T plasmid (Figure 5 is a schematic diagram of the vector). The target primers for the tobacco PDS gene are as follows:
F2NtPDS-U6-26t-F(SEQ ID No.13):F2NtPDS-U6-26t-F (SEQ ID No. 13):
(其中,加粗下划线部分为靶点序列)(The bold underlined part is the target sequence)
F2NtPDS-gRNA-R(SEQ ID No.14):F2NtPDS-gRNA-R (SEQ ID No. 14):
CCATGGAGTCGCTACTACGTCCCCTCGTGAGGGGCAATC。CCATGGAGTCGCTACTACGTCCCCTCGTGAGGGGCAATC.
构建的烟草打靶PDS基因质粒pHCasF2-PDS转化农杆菌GV3101,叶盘法转化烟草,具体步骤如下:The constructed tobacco targeting PDS gene plasmid pHCasF2-PDS was transformed into Agrobacterium GV3101, and the leaf disc method was used to transform tobacco. The specific steps were as follows:
1、叶片消毒:1. Leaf disinfection:
1)选生长1-2月本氏烟的叶片放入一个烧杯中,用无菌水浸泡20-30min;1) Select the leaves of Nicotiana benthamiana that grow from 1-2 months into a beaker, soak in sterile water for 20-30min;
2)倒弃无菌水,用0.6%次氯酸钠浸泡叶片(每50mL溶液中加入1滴Tween-20),让叶片在水中浸泡10min;2) Discard sterile water, soak the leaves with 0.6% sodium hypochlorite (add 1 drop of Tween-20 to every 50 mL of solution), and let the leaves soak in water for 10 minutes;
3)用无菌水冲洗5次,3) Rinse 5 times with sterile water,
2、叶片侵染:2. Leaf infection:
1)用灭菌剪子或者打孔器把叶片边缘切掉,把叶片剪成1cm2大小,避开叶片主脉,将叶片放到液体共培养培养基中;1) Cut off the edge of the leaf with sterilized scissors or a hole punch, cut the leaf into 1cm 2 size, avoid the main vein of the leaf, and put the leaf into the liquid co-cultivation medium;
2)把所有的剪好的叶片转入到侵染液中,侵染8min;然后用无菌滤纸吸去叶片表面的侵染液;2) transfer all the cut leaves into the infection solution, infect 8min; then use sterile filter paper to absorb the infection solution on the leaf surface;
3)将叶片迅速放到固体共培养基上,叶片背面朝下,25℃暗培养48h;3) Put the leaves on the solid co-medium quickly, with the back of the leaves facing down, and cultivate in the dark at 25°C for 48h;
4)将叶片转移到筛选培养基上,25℃,16h光照/8h黑暗培养,每10天更换一次新鲜的筛选培养基,直至转化细胞分化出不定芽。4) The leaves were transferred to the selection medium, and cultured at 25°C under 16h light/8h dark, and the selection medium was replaced every 10 days until the transformed cells differentiated into adventitious buds.
5)选择白化苗进行照相,并提取白化苗基因组DNA,用引物扩增靶点附近的DNA序列,PCR产物送公司测序。5) Select the albino seedlings for photography, extract the genomic DNA of the albino seedlings, use primers to amplify the DNA sequence near the target, and send the PCR product to the company for sequencing.
用于靶点序列附近DNA扩增的PCR扩增引物:PCR amplification primers for DNA amplification near the target sequence:
NtPDS-seq-F(SEQ ID No.15):ATGCCCCAAATTGGACTTGTTTC;NtPDS-seq-F (SEQ ID No. 15): ATGCCCCAAATTGGACTTGTTTC;
NtPDS1-23-R(SEQ ID No.16):CTGGAGCGTGGACAACATAAAT。NtPDS1-23-R (SEQ ID No. 16): CTGGAGCGTGGACAACATAAAT.
构建烟草sgRNA靶点的方法与构建拟南芥体系相同。The method for constructing tobacco sgRNA targets is the same as that for constructing the Arabidopsis system.
实验结果分析:Analysis of results:
根据拟南芥的基因编辑结果,CasF2蛋白发挥基因编辑需要MgCl2的存在,才会具有活性,在sgRNA的引导,对靶点DNA序列进行编辑。因此,在烟草PDS基因组培中,筛选培养基中加入终浓度为100nM MgCl2;通过农杆菌侵染后,经过2轮的筛选培养基后,产生了白化苗(见图6)。在没有加入MgCl2的筛选培养基中,产生的是绿色分化苗。对白化苗的靶点序列进行测序发现,在靶点序列发生了碱基缺失。烟草PDS靶点序列测序图7。According to the gene editing results of Arabidopsis thaliana, the CasF2 protein needs the presence of MgCl 2 for gene editing to be active, and the target DNA sequence is edited under the guidance of sgRNA. Therefore, in tobacco PDS gene tissue culture, the final concentration of 100nM MgCl 2 was added to the screening medium; after Agrobacterium infection, after 2 rounds of screening medium, albino seedlings were produced (see Figure 6). In selection medium without the addition of MgCl2 , green differentiated shoots were produced. The target sequence of the albino seedlings was sequenced, and it was found that there was a base deletion in the target sequence. Figure 7. Tobacco PDS target sequence sequencing.
同时,在拟南芥中使用sgRNA3序列作为sgRNA序列,在烟草中使用sgRNA1序列作为sgRNA序列。通过对构建的拟南芥sgRNA3引导序列和烟草中sgRNA1引导序列分析发现,CasF2蛋白能对pre-crRNA进行处理,可以使用序列更短的sgRNA1引导CasF2进行靶点序列的切割。这样可以同时构建多个sgRNA进行多基因的打靶。Meanwhile, the sgRNA3 sequence was used as the sgRNA sequence in Arabidopsis, and the sgRNA1 sequence was used as the sgRNA sequence in tobacco. Through the analysis of the constructed Arabidopsis sgRNA3 guide sequence and tobacco sgRNA1 guide sequence, it was found that the CasF2 protein can process pre-crRNA, and the sgRNA1 with a shorter sequence can be used to guide CasF2 to cut the target sequence. In this way, multiple sgRNAs can be constructed simultaneously for multi-gene targeting.
最后应说明的是:以上各实施例仅用以说明本发明的技术方案,而非对其限制;尽管参照前述各实施例对本发明进行了详细的说明,本领域的普通技术人员应当理解:其依然可以对前述各实施例所记载的技术方案进行修改,或者对其中部分或者全部技术特征进行等同替换;而这些修改或者替换,并不使相应技术方案的本质脱离本发明各实施例技术方案的范围。Finally, it should be noted that the above embodiments are only used to illustrate the technical solutions of the present invention, but not to limit them; although the present invention has been described in detail with reference to the foregoing embodiments, those of ordinary skill in the art should understand that: The technical solutions described in the foregoing embodiments can still be modified, or some or all of the technical features thereof can be equivalently replaced; and these modifications or replacements do not make the essence of the corresponding technical solutions deviate from the technical solutions of the embodiments of the present invention. scope.
SEQUENCE LISTINGSEQUENCE LISTING
<110> 吉林省农业科学院<110> Jilin Academy of Agricultural Sciences
<120> CasF2蛋白、CRISPR/Cas基因编辑系统及其在植物基因编辑中的应用<120> CasF2 protein, CRISPR/Cas gene editing system and its application in plant gene editing
<130> PA20037888<130> PA20037888
<160> 16<160> 16
<170> PatentIn version 3.3<170> PatentIn version 3.3
<210> 1<210> 1
<211> 2439<211> 2439
<212> DNA<212> DNA
<213> CasF2基因<213> CasF2 gene
<400> 1<400> 1
atggattaca aggatcacga cggtgattac aaggaccatg atatcgacta caaggatgac 60atggattaca aggatcacga cggtgattac aaggaccatg atatcgacta caaggatgac 60
gatgacaaga tggcgcctaa gaagaagaga aaagtgggaa ttcacggtgt tcctgctgcc 120gatgacaaga tggcgcctaa gaagaagaga aaagtgggaa ttcacggtgt tcctgctgcc 120
ccaaagccgg ccgtggagtc agagttctct aaggttctta agaagcattt cccgggcgag 180ccaaagccgg ccgtggagtc agagttctct aaggttctta agaagcattt cccgggcgag 180
cgcttcagat cttcctacat gaagaggggc ggaaagatcc tggcggctca gggagaggag 240cgcttcagat cttcctacat gaagaggggc ggaaagatcc tggcggctca gggagaggag 240
gctgtggttg cctacctcca gggcaagagc gaggaggagc ctccaaactt ccagccgcct 300gctgtggttg cctacctcca gggcaagagc gaggaggagc ctccaaactt ccagccgcct 300
gctaagtgcc acgtcgtgac taagtcaaga gatttcgccg agtggccaat tatgaaggcg 360gctaagtgcc acgtcgtgac taagtcaaga gatttcgccg agtggccaat tatgaaggcg 360
tcagaggcta tccagcgcta catttacgct ctgtctacta ccgagagagc cgcgtgcaag 420tcagaggcta tccagcgcta catttacgct ctgtctacta ccgagagagc cgcgtgcaag 420
ccgggcaagt cttctgagag ccacgctgct tggttcgctg ctacaggcgt ctcaaaccac 480ccgggcaagt cttctgagag ccacgctgct tggttcgctg ctacaggcgt ctcaaaccac 480
ggatactctc atgtgcaggg actcaatctt atcttcgatc atactctcgg aagatacgac 540ggatactctc atgtgcaggg actcaatctt atcttcgatc atactctcgg aagatacgac 540
ggcgttctga agaaggtcca gctccgcaac gagaaggcta gggcccgcct tgagtcaatc 600ggcgttctga agaaggtcca gctccgcaac gagaaggcta gggcccgcct tgagtcaatc 600
aatgcttcta gggctgacga gggcctgcct gagattaagg ctgaggagga ggaggtcgcc 660aatgcttcta gggctgacga gggcctgcct gagattaagg ctgaggagga ggaggtcgcc 660
accaacgaga caggtcatct ccttcagcca ccgggcatca atccatcctt ctacgtgtac 720accaacgaga caggtcatct ccttcagcca ccgggcatca atccatcctt ctacgtgtac 720
cagacaatta gccctcaggc ttacagacca agggatgaga tcgtgctccc tccagagtac 780cagacaatta gccctcaggc ttacagacca agggatgaga tcgtgctccc tccagagtac 780
gcgggttacg ttagagatcc taacgctcct attccacttg gcgttgtccg caatagatgc 840gcgggttacg ttagagatcc taacgctcct attccacttg gcgttgtccg caatagatgc 840
gatatccaga agggctgccc aggatacatt ccggagtggc agagggaggc tggaactgcc 900gatatccaga agggctgccc aggatacatt ccggagtggc agagggaggc tggaactgcc 900
atttccccga agactggaaa ggccgtcacc gtgcctggcc tcagcccaaa gaagaacaag 960atttccccga agactggaaa ggccgtcacc gtgcctggcc tcagcccaaa gaagaacaag 960
aggatgagaa ggtactggcg ctcagagaag gagaaggcgc aggacgctct gctcgttaca 1020aggatgagaa ggtactggcg ctcagagaag gagaaggcgc aggacgctct gctcgttaca 1020
gtcagaatcg gaactgattg ggtggttatt gacgtgcgcg gtcttctgag aaacgcgagg 1080gtcagaatcg gaactgattg ggtggttatt gacgtgcgcg gtcttctgag aaacgcgagg 1080
tggcgcacta tcgctccaaa ggatatttct cttaatgccc tccttgacct gttcaccgga 1140tggcgcacta tcgctccaaa ggatatttct cttaatgccc tccttgacct gttcaccgga 1140
gatccggtca tcgacgtgcg cagaaatatt gttaccttca catacactct cgatgcttgc 1200gatccggtca tcgacgtgcg cagaaatatt gttaccttca catacactct cgatgcttgc 1200
ggcacctacg ccaggaagtg gacacttaag ggaaagcaga ctaaggctac ccttgataag 1260ggcacctacg ccaggaagtg gacacttaag ggaaagcaga ctaaggctac ccttgataag 1260
ctgaccgcca cacagactgt tgcccttgtc gcgatcgacc tgggccagac caaccctatc 1320ctgaccgcca cacagactgt tgcccttgtc gcgatcgacc tgggccagac caaccctatc 1320
tccgccggaa ttagccgcgt cacacaggag aatggtgcgc tccagtgcga gccacttgat 1380tccgccggaa ttagccgcgt cacacaggag aatggtgcgc tccagtgcga gccacttgat 1380
agattcactc tgccggatga cctgctcaag gatatctctg cgtacagaat tgcttgggac 1440agattcactc tgccggatga cctgctcaag gatatctctg cgtacagaat tgcttgggac 1440
aggaacgagg aggagcttag agcgaggtcc gttgaggctc tgccagaggc tcagcaggcc 1500aggaacgagg aggagcttag agcgaggtcc gttgaggctc tgccagaggc tcagcaggcc 1500
gaagttaggg ctctcgatgg cgtcagcaag gagacagcca ggactcagct ctgcgcggat 1560gaagttaggg ctctcgatgg cgtcagcaag gagacagcca ggactcagct ctgcgcggat 1560
ttcggactcg acccgaagag gcttccttgg gataagatgt cttccaacac aactttcatc 1620ttcggactcg acccgaagag gcttccttgg gataagatgt cttccaacac aactttcatc 1620
tcagaggctc ttctgagcaa ttcagtctct cgcgaccagg tgttcttcac cccggcccct 1680tcagaggctc ttctgagcaa ttcagtctct cgcgaccagg tgttcttcac cccggcccct 1680
aagaagggtg ctaagaagaa ggcgcctgtg gaggttatgc gcaaggatag aacatgggcg 1740aagaagggtg ctaagaagaa ggcgcctgtg gaggttatgc gcaaggatag aacatgggcg 1740
cgcgcttaca agccaagact ctctgtggag gcccagaagc ttaagaatga ggcgctctgg 1800cgcgcttaca agccaagact ctctgtggag gcccagaagc ttaagaatga ggcgctctgg 1800
gctcttaaga gaacctcccc agagtacctg aagctcagca ggcgcaagga ggagctttgc 1860gctcttaaga gaacctcccc agagtacctg aagctcagca ggcgcaagga ggagctttgc 1860
agaaggtcta tcaactacgt tattgagaag acacgcagaa ggactcagtg ccagatcgtg 1920agaaggtcta tcaactacgt tattgagaag acacgcagaa ggactcagtg ccagatcgtg 1920
attccggtta ttgaggatct taatgtcaga ttcttccacg gttccggcaa gaggctgcct 1980attccggtta ttgaggatct taatgtcaga ttcttccacg gttccggcaa gaggctgcct 1980
ggttgggaca acttcttcac tgccaagaag gagaatagat ggttcatcca gggcctgcac 2040ggttgggaca acttcttcac tgccaagaag gagaatagat ggttcatcca gggcctgcac 2040
aaggcgttct cagatctcag gacccatcgc tctttctacg tcttcgaagt gaggcctgag 2100aaggcgttct cagatctcag gacccatcgc tctttctacg tcttcgaagt gaggcctgag 2100
aggacctcta ttacatgccc aaagtgcgga cactgcgagg ttggaaacag ggacggagag 2160aggacctcta ttacatgccc aaagtgcgga cactgcgagg ttggaaacag ggacggagag 2160
gctttccagt gcctctcctg cggaaagacc tgcaacgccg atcttgacgt tgcgacccat 2220gctttccagt gcctctcctg cggaaagacc tgcaacgccg atcttgacgt tgcgacccat 2220
aatctgacac aggtcgccct cactggaaag accatgccga agagggagga gcctagggat 2280aatctgacac aggtcgccct cactggaaag accatgccga agagggagga gcctagggat 2280
gctcagggta ctgctccagc taggaagacc aagaaggctt ccaagagcaa ggctccacct 2340gctcagggta ctgctccagc taggaagacc aagaaggctt ccaagagcaa ggctccacct 2340
gctgagaggg aggaccagac accagctcag gagccgtccc agacaagcaa gaggcctgcc 2400gctgagaggg aggaccagac accagctcag gagccgtccc agacaagcaa gaggcctgcc 2400
gcgactaaga aggctggaca ggctaagaag aagaagtga 2439gcgactaaga aggctggaca ggctaagaag aagaagtga 2439
<210> 2<210> 2
<211> 812<211> 812
<212> PRT<212> PRT
<213> CasF2蛋白的氨基酸序列<213> Amino acid sequence of CasF2 protein
<400> 2<400> 2
Met Asp Tyr Lys Asp His Asp Gly Asp Tyr Lys Asp His Asp Ile AspMet Asp Tyr Lys Asp His Asp Gly Asp Tyr Lys Asp His Asp Ile Asp
1 5 10 151 5 10 15
Tyr Lys Asp Asp Asp Asp Lys Met Ala Pro Lys Lys Lys Arg Lys ValTyr Lys Asp Asp Asp Asp Lys Met Ala Pro Lys Lys Lys Arg Lys Val
20 25 30 20 25 30
Gly Ile His Gly Val Pro Ala Ala Pro Lys Pro Ala Val Glu Ser GluGly Ile His Gly Val Pro Ala Ala Pro Lys Pro Ala Val Glu Ser Glu
35 40 45 35 40 45
Phe Ser Lys Val Leu Lys Lys His Phe Pro Gly Glu Arg Phe Arg SerPhe Ser Lys Val Leu Lys Lys His Phe Pro Gly Glu Arg Phe Arg Ser
50 55 60 50 55 60
Ser Tyr Met Lys Arg Gly Gly Lys Ile Leu Ala Ala Gln Gly Glu GluSer Tyr Met Lys Arg Gly Gly Lys Ile Leu Ala Ala Gln Gly Glu Glu
65 70 75 8065 70 75 80
Ala Val Val Ala Tyr Leu Gln Gly Lys Ser Glu Glu Glu Pro Pro AsnAla Val Val Ala Tyr Leu Gln Gly Lys Ser Glu Glu Glu Pro Pro Asn
85 90 95 85 90 95
Phe Gln Pro Pro Ala Lys Cys His Val Val Thr Lys Ser Arg Asp PhePhe Gln Pro Pro Ala Lys Cys His Val Val Thr Lys Ser Arg Asp Phe
100 105 110 100 105 110
Ala Glu Trp Pro Ile Met Lys Ala Ser Glu Ala Ile Gln Arg Tyr IleAla Glu Trp Pro Ile Met Lys Ala Ser Glu Ala Ile Gln Arg Tyr Ile
115 120 125 115 120 125
Tyr Ala Leu Ser Thr Thr Glu Arg Ala Ala Cys Lys Pro Gly Lys SerTyr Ala Leu Ser Thr Thr Glu Arg Ala Ala Cys Lys Pro Gly Lys Ser
130 135 140 130 135 140
Ser Glu Ser His Ala Ala Trp Phe Ala Ala Thr Gly Val Ser Asn HisSer Glu Ser His Ala Ala Trp Phe Ala Ala Thr Gly Val Ser Asn His
145 150 155 160145 150 155 160
Gly Tyr Ser His Val Gln Gly Leu Asn Leu Ile Phe Asp His Thr LeuGly Tyr Ser His Val Gln Gly Leu Asn Leu Ile Phe Asp His Thr Leu
165 170 175 165 170 175
Gly Arg Tyr Asp Gly Val Leu Lys Lys Val Gln Leu Arg Asn Glu LysGly Arg Tyr Asp Gly Val Leu Lys Lys Val Gln Leu Arg Asn Glu Lys
180 185 190 180 185 190
Ala Arg Ala Arg Leu Glu Ser Ile Asn Ala Ser Arg Ala Asp Glu GlyAla Arg Ala Arg Leu Glu Ser Ile Asn Ala Ser Arg Ala Asp Glu Gly
195 200 205 195 200 205
Leu Pro Glu Ile Lys Ala Glu Glu Glu Glu Val Ala Thr Asn Glu ThrLeu Pro Glu Ile Lys Ala Glu Glu Glu Glu Val Ala Thr Asn Glu Thr
210 215 220 210 215 220
Gly His Leu Leu Gln Pro Pro Gly Ile Asn Pro Ser Phe Tyr Val TyrGly His Leu Leu Gln Pro Pro Gly Ile Asn Pro Ser Phe Tyr Val Tyr
225 230 235 240225 230 235 240
Gln Thr Ile Ser Pro Gln Ala Tyr Arg Pro Arg Asp Glu Ile Val LeuGln Thr Ile Ser Pro Gln Ala Tyr Arg Pro Arg Asp Glu Ile Val Leu
245 250 255 245 250 255
Pro Pro Glu Tyr Ala Gly Tyr Val Arg Asp Pro Asn Ala Pro Ile ProPro Pro Glu Tyr Ala Gly Tyr Val Arg Asp Pro Asn Ala Pro Ile Pro
260 265 270 260 265 270
Leu Gly Val Val Arg Asn Arg Cys Asp Ile Gln Lys Gly Cys Pro GlyLeu Gly Val Val Arg Asn Arg Cys Asp Ile Gln Lys Gly Cys Pro Gly
275 280 285 275 280 285
Tyr Ile Pro Glu Trp Gln Arg Glu Ala Gly Thr Ala Ile Ser Pro LysTyr Ile Pro Glu Trp Gln Arg Glu Ala Gly Thr Ala Ile Ser Pro Lys
290 295 300 290 295 300
Thr Gly Lys Ala Val Thr Val Pro Gly Leu Ser Pro Lys Lys Asn LysThr Gly Lys Ala Val Thr Val Pro Gly Leu Ser Pro Lys Lys Asn Lys
305 310 315 320305 310 315 320
Arg Met Arg Arg Tyr Trp Arg Ser Glu Lys Glu Lys Ala Gln Asp AlaArg Met Arg Arg Tyr Trp Arg Ser Glu Lys Glu Lys Ala Gln Asp Ala
325 330 335 325 330 335
Leu Leu Val Thr Val Arg Ile Gly Thr Asp Trp Val Val Ile Asp ValLeu Leu Val Thr Val Arg Ile Gly Thr Asp Trp Val Val Ile Asp Val
340 345 350 340 345 350
Arg Gly Leu Leu Arg Asn Ala Arg Trp Arg Thr Ile Ala Pro Lys AspArg Gly Leu Leu Arg Asn Ala Arg Trp Arg Thr Ile Ala Pro Lys Asp
355 360 365 355 360 365
Ile Ser Leu Asn Ala Leu Leu Asp Leu Phe Thr Gly Asp Pro Val IleIle Ser Leu Asn Ala Leu Leu Asp Leu Phe Thr Gly Asp Pro Val Ile
370 375 380 370 375 380
Asp Val Arg Arg Asn Ile Val Thr Phe Thr Tyr Thr Leu Asp Ala CysAsp Val Arg Arg Asn Ile Val Thr Phe Thr Tyr Thr Leu Asp Ala Cys
385 390 395 400385 390 395 400
Gly Thr Tyr Ala Arg Lys Trp Thr Leu Lys Gly Lys Gln Thr Lys AlaGly Thr Tyr Ala Arg Lys Trp Thr Leu Lys Gly Lys Gln Thr Lys Ala
405 410 415 405 410 415
Thr Leu Asp Lys Leu Thr Ala Thr Gln Thr Val Ala Leu Val Ala IleThr Leu Asp Lys Leu Thr Ala Thr Gln Thr Val Ala Leu Val Ala Ile
420 425 430 420 425 430
Asp Leu Gly Gln Thr Asn Pro Ile Ser Ala Gly Ile Ser Arg Val ThrAsp Leu Gly Gln Thr Asn Pro Ile Ser Ala Gly Ile Ser Arg Val Thr
435 440 445 435 440 445
Gln Glu Asn Gly Ala Leu Gln Cys Glu Pro Leu Asp Arg Phe Thr LeuGln Glu Asn Gly Ala Leu Gln Cys Glu Pro Leu Asp Arg Phe Thr Leu
450 455 460 450 455 460
Pro Asp Asp Leu Leu Lys Asp Ile Ser Ala Tyr Arg Ile Ala Trp AspPro Asp Asp Leu Leu Lys Asp Ile Ser Ala Tyr Arg Ile Ala Trp Asp
465 470 475 480465 470 475 480
Arg Asn Glu Glu Glu Leu Arg Ala Arg Ser Val Glu Ala Leu Pro GluArg Asn Glu Glu Glu Leu Arg Ala Arg Ser Val Glu Ala Leu Pro Glu
485 490 495 485 490 495
Ala Gln Gln Ala Glu Val Arg Ala Leu Asp Gly Val Ser Lys Glu ThrAla Gln Gln Ala Glu Val Arg Ala Leu Asp Gly Val Ser Lys Glu Thr
500 505 510 500 505 510
Ala Arg Thr Gln Leu Cys Ala Asp Phe Gly Leu Asp Pro Lys Arg LeuAla Arg Thr Gln Leu Cys Ala Asp Phe Gly Leu Asp Pro Lys Arg Leu
515 520 525 515 520 525
Pro Trp Asp Lys Met Ser Ser Asn Thr Thr Phe Ile Ser Glu Ala LeuPro Trp Asp Lys Met Ser Ser Asn Thr Thr Phe Ile Ser Glu Ala Leu
530 535 540 530 535 540
Leu Ser Asn Ser Val Ser Arg Asp Gln Val Phe Phe Thr Pro Ala ProLeu Ser Asn Ser Val Ser Arg Asp Gln Val Phe Phe Thr Pro Ala Pro
545 550 555 560545 550 555 560
Lys Lys Gly Ala Lys Lys Lys Ala Pro Val Glu Val Met Arg Lys AspLys Lys Gly Ala Lys Lys Lys Lys Ala Pro Val Glu Val Met Arg Lys Asp
565 570 575 565 570 575
Arg Thr Trp Ala Arg Ala Tyr Lys Pro Arg Leu Ser Val Glu Ala GlnArg Thr Trp Ala Arg Ala Tyr Lys Pro Arg Leu Ser Val Glu Ala Gln
580 585 590 580 585 590
Lys Leu Lys Asn Glu Ala Leu Trp Ala Leu Lys Arg Thr Ser Pro GluLys Leu Lys Asn Glu Ala Leu Trp Ala Leu Lys Arg Thr Ser Pro Glu
595 600 605 595 600 605
Tyr Leu Lys Leu Ser Arg Arg Lys Glu Glu Leu Cys Arg Arg Ser IleTyr Leu Lys Leu Ser Arg Arg Lys Glu Glu Leu Cys Arg Arg Ser Ile
610 615 620 610 615 620
Asn Tyr Val Ile Glu Lys Thr Arg Arg Arg Thr Gln Cys Gln Ile ValAsn Tyr Val Ile Glu Lys Thr Arg Arg Arg Thr Gln Cys Gln Ile Val
625 630 635 640625 630 635 640
Ile Pro Val Ile Glu Asp Leu Asn Val Arg Phe Phe His Gly Ser GlyIle Pro Val Ile Glu Asp Leu Asn Val Arg Phe Phe His Gly Ser Gly
645 650 655 645 650 655
Lys Arg Leu Pro Gly Trp Asp Asn Phe Phe Thr Ala Lys Lys Glu AsnLys Arg Leu Pro Gly Trp Asp Asn Phe Phe Thr Ala Lys Lys Glu Asn
660 665 670 660 665 670
Arg Trp Phe Ile Gln Gly Leu His Lys Ala Phe Ser Asp Leu Arg ThrArg Trp Phe Ile Gln Gly Leu His Lys Ala Phe Ser Asp Leu Arg Thr
675 680 685 675 680 685
His Arg Ser Phe Tyr Val Phe Glu Val Arg Pro Glu Arg Thr Ser IleHis Arg Ser Phe Tyr Val Phe Glu Val Arg Pro Glu Arg Thr Ser Ile
690 695 700 690 695 700
Thr Cys Pro Lys Cys Gly His Cys Glu Val Gly Asn Arg Asp Gly GluThr Cys Pro Lys Cys Gly His Cys Glu Val Gly Asn Arg Asp Gly Glu
705 710 715 720705 710 715 720
Ala Phe Gln Cys Leu Ser Cys Gly Lys Thr Cys Asn Ala Asp Leu AspAla Phe Gln Cys Leu Ser Cys Gly Lys Thr Cys Asn Ala Asp Leu Asp
725 730 735 725 730 735
Val Ala Thr His Asn Leu Thr Gln Val Ala Leu Thr Gly Lys Thr MetVal Ala Thr His Asn Leu Thr Gln Val Ala Leu Thr Gly Lys Thr Met
740 745 750 740 745 750
Pro Lys Arg Glu Glu Pro Arg Asp Ala Gln Gly Thr Ala Pro Ala ArgPro Lys Arg Glu Glu Pro Arg Asp Ala Gln Gly Thr Ala Pro Ala Arg
755 760 765 755 760 765
Lys Thr Lys Lys Ala Ser Lys Ser Lys Ala Pro Pro Ala Glu Arg GluLys Thr Lys Lys Ala Ser Lys Ser Lys Ala Pro Pro Ala Glu Arg Glu
770 775 780 770 775 780
Asp Gln Thr Pro Ala Gln Glu Pro Ser Gln Thr Ser Lys Arg Pro AlaAsp Gln Thr Pro Ala Gln Glu Pro Ser Gln Thr Ser Lys Arg Pro Ala
785 790 795 800785 790 795 800
Ala Thr Lys Lys Ala Gly Gln Ala Lys Lys Lys LysAla Thr Lys Lys Ala Gly Gln Ala Lys Lys Lys Lys
805 810 805 810
<210> 3<210> 3
<211> 40<211> 40
<212> DNA<212> DNA
<213> sgRNA3序列<213> sgRNA3 sequence
<400> 3<400> 3
taatgtcgga acgctcaacg attgcccctc acgaggggac 40taatgtcgga acgctcaacg attgcccctc acgaggggac 40
<210> 4<210> 4
<211> 29<211> 29
<212> DNA<212> DNA
<213> sgRNA1序列<213> sgRNA1 sequence
<400> 4<400> 4
cgctcaacga ttgcccctca cgaggggac 29cgctcaacga ttgcccctca cgaggggac 29
<210> 5<210> 5
<211> 18<211> 18
<212> DNA<212> DNA
<213> 拟南芥BRI1基因的sgRNA靶点序列<213> sgRNA target sequence of Arabidopsis BRI1 gene
<400> 5<400> 5
ctgcgaattc aatctccg 18ctgcgaattc aatctccg 18
<210> 6<210> 6
<211> 18<211> 18
<212> DNA<212> DNA
<213> 烟草PDS基因的sgRNA靶点序列<213> sgRNA target sequence of tobacco PDS gene
<400> 6<400> 6
gtagtagcga ctccatgg 18gtagtagcga ctccatgg 18
<210> 7<210> 7
<211> 39<211> 39
<212> DNA<212> DNA
<213> 人工序列 U6-26p-F<213> Artificial sequence U6-26p-F
<400> 7<400> 7
ttcagaggtc tcttagtcga cttgccttcc gcacaatac 39ttcagaggtc tcttagtcga cttgccttcc gcacaatac 39
<210> 8<210> 8
<211> 39<211> 39
<212> DNA<212> DNA
<213> 人工序列 U6-26t-R<213> Artificial sequence U6-26t-R
<400> 8<400> 8
agcgtgggtc tcgcgcctat tggtttatct catcggaac 39agcgtgggtc tcgcgcctat tggtttatct catcggaac 39
<210> 9<210> 9
<211> 44<211> 44
<212> DNA<212> DNA
<213> 人工序列 F2BRI1-U6-26t-F<213> Artificial sequence F2BRI1-U6-26t-F
<400> 9<400> 9
ctgcgaattc aatctccgtt ttttttgcaa aattttccag atcg 44ctgcgaattc aatctccgtt ttttttgcaa aattttccag atcg 44
<210> 10<210> 10
<211> 39<211> 39
<212> DNA<212> DNA
<213> 人工序列<213> Artificial sequences
<400> 10<400> 10
cggagattga attcgcaggt cccctcgtga ggggcaatc 39cggagattga attcgcaggt cccctcgtga ggggcaatc 39
<210> 11<210> 11
<211> 22<211> 22
<212> DNA<212> DNA
<213> 人工序列 F2BRI1crispr-identify-F<213> Artificial sequence F2BRI1crispr-identify-F
<400> 11<400> 11
ctggcttcaa gtgctctgct tc 22ctggcttcaa gtgctctgct tc 22
<210> 12<210> 12
<211> 20<211> 20
<212> DNA<212> DNA
<213> 人工序列 F2BRI1crispr-identify-R<213> Artificial sequence F2BRI1crispr-identify-R
<400> 12<400> 12
gtaattcgcc ggaaaactcg 20gtaattcgcc ggaaaactcg 20
<210> 13<210> 13
<211> 44<211> 44
<212> DNA<212> DNA
<213> 人工序列 F2NtPDS-U6-26t-F<213> Artificial sequence F2NtPDS-U6-26t-F
<400> 13<400> 13
gtagtagcga ctccatggtt ttttttgcaa aattttccag atcg 44gtagtagcga ctccatggtt ttttttgcaa aattttccag atcg 44
<210> 14<210> 14
<211> 39<211> 39
<212> DNA<212> DNA
<213> 人工序列 F2NtPDS-gRNA-R<213> Artificial sequence F2NtPDS-gRNA-R
<400> 14<400> 14
ccatggagtc gctactacgt cccctcgtga ggggcaatc 39ccatggagtc gctactacgt cccctcgtga ggggcaatc 39
<210> 15<210> 15
<211> 23<211> 23
<212> DNA<212> DNA
<213> 人工序列 NtPDS-seq-F<213> Artificial sequence NtPDS-seq-F
<400> 15<400> 15
atgccccaaa ttggacttgt ttc 23atgccccaaa ttggacttgt ttc 23
<210> 16<210> 16
<211> 22<211> 22
<212> DNA<212> DNA
<213> 人工序列 NtPDS1-23-R<213> Artificial sequence NtPDS1-23-R
<400> 16<400> 16
ctggagcgtg gacaacataa at 22ctggagcgtg gacaacataa at 22
Claims (10)
Priority Applications (2)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| CN202310514396.0A CN116555226A (en) | 2022-03-03 | 2022-03-03 | CasF2 protein, CRISPR/Cas gene editing system and its application in plant gene editing |
| CN202210204370.1A CN114438056B (en) | 2022-03-03 | 2022-03-03 | CasF2 protein, CRISPR/Cas gene editing system and application thereof in plant gene editing |
Applications Claiming Priority (1)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| CN202210204370.1A CN114438056B (en) | 2022-03-03 | 2022-03-03 | CasF2 protein, CRISPR/Cas gene editing system and application thereof in plant gene editing |
Related Child Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| CN202310514396.0A Division CN116555226A (en) | 2022-03-03 | 2022-03-03 | CasF2 protein, CRISPR/Cas gene editing system and its application in plant gene editing |
Publications (2)
| Publication Number | Publication Date |
|---|---|
| CN114438056A true CN114438056A (en) | 2022-05-06 |
| CN114438056B CN114438056B (en) | 2023-11-21 |
Family
ID=81358995
Family Applications (2)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| CN202210204370.1A Active CN114438056B (en) | 2022-03-03 | 2022-03-03 | CasF2 protein, CRISPR/Cas gene editing system and application thereof in plant gene editing |
| CN202310514396.0A Pending CN116555226A (en) | 2022-03-03 | 2022-03-03 | CasF2 protein, CRISPR/Cas gene editing system and its application in plant gene editing |
Family Applications After (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| CN202310514396.0A Pending CN116555226A (en) | 2022-03-03 | 2022-03-03 | CasF2 protein, CRISPR/Cas gene editing system and its application in plant gene editing |
Country Status (1)
| Country | Link |
|---|---|
| CN (2) | CN114438056B (en) |
Families Citing this family (1)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN117402855B (en) * | 2023-12-14 | 2024-03-19 | 中国农业科学院植物保护研究所 | Cas protein, gene editing system and application |
Citations (6)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN113308451A (en) * | 2020-12-07 | 2021-08-27 | 中国科学院动物研究所 | Engineered Cas effector proteins and methods of use thereof |
| CN113373130A (en) * | 2021-05-31 | 2021-09-10 | 复旦大学 | Cas12 protein, gene editing system containing Cas12 protein and application |
| WO2021216512A1 (en) * | 2020-04-20 | 2021-10-28 | The Regents Of The University Of California | Crispr systems in plants |
| CN113717962A (en) * | 2021-09-10 | 2021-11-30 | 武汉艾迪晶生物科技有限公司 | Cas phi-2 protein for rice gene editing and expression cassette and expression vector thereof |
| CN113913499A (en) * | 2020-12-25 | 2022-01-11 | 山东舜丰生物科技有限公司 | Method for detecting target mutation by using Cas12j effector protein |
| WO2022026403A2 (en) * | 2020-07-31 | 2022-02-03 | Inari Agriculture Technology, Inc. | Removable plant transgenic loci with cognate guide rna recognition sites |
-
2022
- 2022-03-03 CN CN202210204370.1A patent/CN114438056B/en active Active
- 2022-03-03 CN CN202310514396.0A patent/CN116555226A/en active Pending
Patent Citations (6)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO2021216512A1 (en) * | 2020-04-20 | 2021-10-28 | The Regents Of The University Of California | Crispr systems in plants |
| WO2022026403A2 (en) * | 2020-07-31 | 2022-02-03 | Inari Agriculture Technology, Inc. | Removable plant transgenic loci with cognate guide rna recognition sites |
| CN113308451A (en) * | 2020-12-07 | 2021-08-27 | 中国科学院动物研究所 | Engineered Cas effector proteins and methods of use thereof |
| CN113913499A (en) * | 2020-12-25 | 2022-01-11 | 山东舜丰生物科技有限公司 | Method for detecting target mutation by using Cas12j effector protein |
| CN113373130A (en) * | 2021-05-31 | 2021-09-10 | 复旦大学 | Cas12 protein, gene editing system containing Cas12 protein and application |
| CN113717962A (en) * | 2021-09-10 | 2021-11-30 | 武汉艾迪晶生物科技有限公司 | Cas phi-2 protein for rice gene editing and expression cassette and expression vector thereof |
Non-Patent Citations (3)
| Title |
|---|
| PATRICK PAUSCH ET AL.: "CRISPR-CasF from huge phages is a hypercompact genome editor", 《SCIENCE》 * |
| PAUSCH,P. ET AL.: "PDB:7LYS_A", 《NCBI》 * |
| 刘海英: "基因编辑工具再添超紧凑型系统——CRISPR-CasΦ 具有更广泛靶标识别能力", 《JOURNAL OF FOOD SCIENCE AND BIOTECHNOLOGY》 * |
Also Published As
| Publication number | Publication date |
|---|---|
| CN116555226A (en) | 2023-08-08 |
| CN114438056B (en) | 2023-11-21 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| CN107988229B (en) | A method for obtaining tiller-altered rice by modifying the OsTAC1 gene using CRISPR-Cas | |
| CN109652422B (en) | Efficient single base editing system OsSpCas9-eCDA and its application | |
| CN106467909A (en) | A kind of method that acquisition glyphosate-resistant rice is replaced by nucleotide fixed point | |
| CN106957855A (en) | Use CRISPR/Cas9 technology targeting knock out paddy rice Dwarfing genes SD1 method | |
| Ding et al. | Effective reduction in chimeric mutants of poplar trees produced by CRISPR/Cas9 through a second round of shoot regeneration | |
| CN113801891B (en) | Construction method and application of sugar beet BvCENH3 gene haploid induction line | |
| CN116286742B (en) | CasD protein, CRISPR/CasD gene editing system and application thereof in plant gene editing | |
| WO2023092731A1 (en) | Mad7-nls fusion protein, and nucleic acid construct for site-directed editing of plant genome and application thereof | |
| CN103981199A (en) | Glyphosate resistance gene-containing expression vector and application thereof | |
| CN110951743B (en) | Method for improving plant gene replacement efficiency | |
| CN117402874A (en) | Potato U6 gene promoter StU6 8-1 and its cloning and application | |
| CN102943091A (en) | Method for cultivating tobacco capable of resisting various viruses by adopting RNAi (RNA interference) technique | |
| CN114438056B (en) | CasF2 protein, CRISPR/Cas gene editing system and application thereof in plant gene editing | |
| WO2001096583A2 (en) | Removal of selectable markers from transformed cells | |
| CN115197332A (en) | Plant expression vector for creating high-zinc low-cadmium rice and application thereof | |
| CN118853732A (en) | Application of TB1 in Leymus chinensis production and breeding | |
| CN117625682A (en) | Methods to improve potato PSTVd resistance based on CRISPR-RfxCas13d system and its products and applications | |
| CN116751272A (en) | Application of NAC079 gene in regulation and control of rice sheath blight resistance | |
| CN115960948A (en) | Application of ZmCP03 gene in corn | |
| CN113897372A (en) | Application of OsFWL7 gene in increasing content of metal trace elements in rice grains | |
| CN119285795B (en) | Cas9-GR fusion protein and application thereof in gene editing | |
| CN120944847B (en) | Wheat TaPPKL1 mutant and application | |
| CN105087591B (en) | Paddy rice root tip specific expression promoter POsRo3 | |
| KR102921480B1 (en) | NML1 gene controlling Cucumber Mosaic Virus resistance of plant and method for producing tomato plant having edited NML1 gene using CRISPR/Cas system | |
| CN117024547B (en) | GsSYP51b protein and application of encoding gene thereof in regulation and control of plant stress tolerance |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| PB01 | Publication | ||
| PB01 | Publication | ||
| SE01 | Entry into force of request for substantive examination | ||
| SE01 | Entry into force of request for substantive examination | ||
| GR01 | Patent grant | ||
| GR01 | Patent grant | ||
| CP03 | Change of name, title or address |
Address after: No.130033 ecological street, Changchun City, Jilin Province Patentee after: Jilin Academy of Agricultural Sciences (China Agricultural Science and Technology Northeast Innovation Center) Country or region after: China Address before: No. 1363 Ecological Street, Jingyue High tech Development Zone, Changchun City, Jilin Province Patentee before: Jilin Academy of Agricultural Sciences Country or region before: China |
|
| CP03 | Change of name, title or address |