CN114315989A - Recombinant novel coronavirus protein vaccine, preparation method and application thereof - Google Patents
Recombinant novel coronavirus protein vaccine, preparation method and application thereof Download PDFInfo
- Publication number
- CN114315989A CN114315989A CN202210073510.6A CN202210073510A CN114315989A CN 114315989 A CN114315989 A CN 114315989A CN 202210073510 A CN202210073510 A CN 202210073510A CN 114315989 A CN114315989 A CN 114315989A
- Authority
- CN
- China
- Prior art keywords
- asn
- val
- ser
- phe
- tyr
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Pending
Links
Images
Landscapes
- Medicines Containing Antibodies Or Antigens For Use As Internal Diagnostic Agents (AREA)
- Peptides Or Proteins (AREA)
Abstract
The invention discloses a recombinant novel coronavirus protein, which covers specific mutation sites of Delta, Beta and Omicron variant RBD, and a recombinant vaccine taking the mutation sites as a target antigen has broad-spectrum protection capability of a cross-epidemic strain.
Description
Technical Field
The invention relates to the field of biomedicine, in particular to a recombinant novel coronavirus protein vaccine, a preparation method and application thereof.
Background
Novel coronaviruses (SARS-CoV-2), belonging to the order Nidovirales (Nidovirales), the family Coronaviridae (Coronaviridae), the subfamily orthocoronaviruses, the genus Betaconoviridus, the subfamily Sarbecovirus, the SARS-like virus species, single-stranded positive-strand RNA viruses, have an envelope with a genome of about 29.9kb in overall length, encode mostly non-structural proteins, are involved in functions such as viral replication and translation, and encode a small portion of the sequence of structural proteins, such as: s protein (spike protein), M protein (membrane protein), E protein (envelope protein) and N protein (nucleo protein). In addition, there are several accessory proteins: 3a,3b, p6,7a,7b,8b,9b and orf14, all of which are involved in viral assembly. S, M and the E protein constitute the viral envelope and are the major surface antigens of the virus to elicit an immune response. Wherein the S protein is a transmembrane glycoprotein with a molecular weight of about 150kDa, and forms prominent homotrimers on the surface of the virus. S consists of two functional subunits, cleaved at the boundary between the S1 and S2 subunits (S1/S2 cleavage site), which remain non-covalently associated in the pre-fusion conformation. The S2 subunit is also composed of multiple domains, whose function is primarily to mediate fusion of the virus with the host cell. The distal S1 subunit is structurally divided into four distinct domains: the vaccine is characterized by comprising an N-terminal domain (NTD), a Receptor Binding Domain (RBD), a C-terminal domain 1(CTD1) and a C-terminal domain 2(CTD2), wherein the RBD is mainly responsible for binding with a receptor angiotensin converting enzyme 2 (ACE 2) on the surface of a host cell so as to mediate virus infection on the host cell, so that the S protein and the RBD are main targets for current genetic engineering vaccine development.
Up to now, there are 8 vaccines approved worldwide for marketing, BNT162b2 and mRNA-1273 for the united states authorization for Emergency Use (EUA), AZD1222 for the british authorization for Emergency Use (EUA), 3 new crown inactivated vaccines for chinese medicine chinese (beijing and wuhan) and beijing kexing, conxino biological adenovirus vector vaccines, zhifei biological recombinant protein vaccines, and russian approved "satellite V" for marketing. In addition, there are several hundred vaccines in multiple routes in different stages of clinical research. The new coronavirus belongs to RNA virus, is easy to mutate, and has found more than seven million new coronavirus mutant strains globally, wherein the main mutant strains comprise: alpha (B.1.1.7) mutant, Beta (B.1.351) mutant, Gamma (P1) mutant, Epsilon (B.1.429) mutant, Delta (B.1.617.2) mutant, Kappa (B.1.617.1) mutant and Omicron (B.1.1.529) mutant, the occurrence of variant strains affects the protective effect of existing vaccines to different extents.
The world health organization classifies the above 5 variants as Variants (VOCs) that need attention based on the differences in the transmissibility, pathogenicity, or immune escape capabilities of the SARS-CoV-2 variants. Among these 5 VOCs, numerous studies have demonstrated that the Beta variant has a strong immune escape capacity and the Delta variant has a strong transmission capacity. Preliminary evidence shows that the Omicron variant has strong transmission capability and strong immune escape capability, and the immune escape capability of the Omicron variant is even far higher than that of a Beta strain.
The Omicron mutant is defined as a fifth 'concern variant' by the world health organization, and the mutant contains 36 amino acid mutation sites in total, wherein the mutation sites comprise K417N/T, E484K/Q/A, N501Y, D614G and P681H/R mutation sites which occur at high frequency in an dominant epidemic strain Alpha (B.1.1.7) mutant, a Beta (B.1.351) mutant, a Gamma (P1) mutant, a Delta (B.1.617.2) mutant and a Kappa (B.1.617.1) mutant. A large number of data show that the ormikrong mutant strain can improve the affinity of the virus to an ACE2 receptor, weaken the effect of a neutralizing antibody, thereby enhancing the toxicity and infectivity of the virus, accelerating the escape of the virus and reducing the protective effect of the vaccine.
The emergence of these strains with strong immune escape capacity raises a great concern for the effectiveness of the existing new corona vaccines. The development of a new generation vaccine with cross-protection capability is an effective means for dealing with these variants. Integrating multiple variants into the same immunogen to construct mosaic antigen molecules, which is a feasible strategy for realizing single-component cross-protection vaccines. Similar strategies have been applied in the design of broad-spectrum influenza vaccines. However, since there are many mutation sites in different mutants, it is urgent to find a broad-spectrum vaccine with excellent immunogenicity against many new strains of coronavirus epidemic.
Disclosure of Invention
One aspect of the present invention is to provide a single-component broad-spectrum recombinant protein vaccine capable of generating good cross-neutralization activity against a plurality of novel coronavirus epidemic strains, especially a plurality of VOC variant strains including Omicron, Beta, Delta, etc., against the shortage of broad-spectrum novel coronavirus (SARS-CoV-2) vaccines, especially against variant strains, especially against Omicron strains, in the prior art.
The technical scheme provided by the invention is as follows:
a recombinant novel coronavirus protein is a trimer, and comprises three subunits consisting of 319 th to 537 th amino acid fragments of an RBD region of a novel coronavirus S protein,
the first subunit of the recombinant novel coronavirus protein comprises an L452R and/or T478K mutation site; and
the second subunit of the recombinant novel coronavirus protein comprises one or more mutation sites selected from K417N, E484K or N501Y; and
five or more than five mutation sites of G339D, S371L, S373P, S375F, K417N, N440K, G446S, S477N, T478K, E484A, Q493R, G496S, Q498R, N501Y or Y505H are included in the third subunit of the recombinant novel coronavirus protein.
The receptor-binding domain (RBD) of the new coronavirus spike (S) protein is directly involved in the binding of host cell receptors and plays a key role in the process of virus invasion into host cells. Meanwhile, a great deal of research indicates that RBD contains a major neutralizing epitope, and thus, RBD is one of the major target antigens for development of a neocoronary vaccine. However, RBD monomers are not highly immunogenic due to their small molecular size. The natural S protein is of a trimeric structure, and the RBD also exists in a trimerized form. The trimerization RBD is constructed, the natural structural form of the RBD is simulated to the maximum extent, meanwhile, the molecular size of the antigen is increased through trimerization, the repetitive regular arrangement of the antigen is realized, the B cell receptor crosslinking is enhanced, and the immunogenicity of the RBD can be obviously improved. Trimerization of antigen is usually achieved by introducing exogenous linker arms or trimerization motifs, but the introduction of exogenous sequences may bring unexpected immune response with certain safety risk. The inventors achieved trimerization of RBD without introducing a trimerization motif by computational analysis of RBD structure. The RBD has the following structural characteristics: (1) the N-end and the C-end of the RBD are provided with longer flexible loop structures, and the loop structures of the RBD can be used as connecting arms among different RBDs in the trimerization process, so that the introduction of an external connecting arm is avoided, and the trimerization space barrier is reduced; (2) in the natural structure of the S protein, RBDs are relatively independent, strong interaction does not exist between the RBDs and other structural domains, and the folding of the RBDs does not need the assistance of the other structural domains; (3) the RBD has a compact space structure, the core structure of the RBD is composed of a plurality of beta sheet layers, 4 disulfide bonds are contained in the structure domain, the stability of the structure domain is further enhanced, meanwhile, the N-end and the C-end of the RBD are close, a large space barrier does not exist in the trimerization process, and the core structure of the RBD cannot be damaged. Based on the structural characteristics of the RBD, the invention designs an interception scheme of the RBD, namely, an amino acid fragment at 319-537 positions is intercepted, the interception scheme reserves loop structures at two ends of the RBD as far as possible, and meanwhile, the N-end and C-end interfaces are ensured to be closer. And then, connecting the three intercepted RBD zone segments in series end to form a new RBD trimerization fusion protein.
In order to realize cross protection of different variants of the new coronavirus and construct mosaic type RBD trimerization antigen, the inventor selects Delta strain with strong infectivity, and Omicron and Beta strain with strongest immune escape from VOCs variants, and intercepts and co-assembles RBDs of the three variants into mosaic type trimerization RBD protein. Compared with the prototype strain RBD, the inventors selected RBD regions of two mutation sites of L452R and T478K of Delta strain (namely, the first subunit, which can be referred to as the first subunit/Delta subunit), RBD regions of three mutation sites of K417N, E484K and N501Y of Beta strain (namely, the second subunit, which can be referred to as the second subunit/Beta subunit), and 15 mutated RBD regions of G339D, S371L, S39373 38, S375F, K417N, N440K, G446S, S477 638, T63478, E484A, Q493 686 8, G496S, Q35498, N501Y and Y505H of Omicron variant strain (namely, the third subunit, which can be referred to as the third subunit/Omicron subunit). Through screening, the inventor finds that the mutations not only appear in the three variant strains, but also frequently appear in a plurality of different virus lineages, and further verifies that the selected mutation sites have obvious selective advantages in virus evolution and also predict that the mutations can appear in future mutants independently or in combination.
In certain embodiments of the invention, a combination of portions of the above mutation sites is shown in table 1, for example.
TABLE 1
Based on the structural characteristics of the RBD region of the S protein of the novel coronavirus, the invention designs a brand-new fusion protein by utilizing a computational biology method, wherein the protein comprises three RBD structural domains, and forms a trimer form with stable antigen conformation under the condition that no exogenous connecting arm or other unrelated components can be introduced, so that the trimerization of the RBD protein is realized. The RBD tripolymer protein is recombined, expressed and purified by utilizing the genetic engineering technology, and then is mixed with an adjuvant to prepare the vaccine. After a certain dosage and dose immunization, protective neutralizing antibodies against a plurality of novel coronavirus epidemic strains can be generated, and the protective neutralizing antibodies can be used for treating and/or preventing SARS-CoV-2 infection and/or novel coronavirus diseases (COVID-19). Because the RBD region has definite function and clear structure, the gene is responsible for recognizing ACE2 receptor of host cells, and simultaneously, the Antibody generated aiming at RBD has definite function and specific target spot, thereby maximally avoiding inducing an organism to generate Antibody-Dependent Enhancement (ADE).
Preferably, in some embodiments of the present invention, the primary structure of the three subunits of the above recombinant novel coronavirus protein is such that the three subunits are connected in series in the order of N-terminus to C-terminus.
In the present invention, the first subunit, the second subunit, and the third subunit may be arranged in any suitable order. For example, as shown in table 2.
TABLE 2
| Serial number | Order of arrangement | Sequence of |
| 1 | First subunit + second subunit + third subunit | SEQ ID No.1 |
| 2 | First subunit + third subunit + second subunit | SEQ ID No.2 |
| 3 | Second subunit + first subunit + third subunit | SEQ ID No.3 |
| 4 | Second subunit + third subunit + first subunit | SEQ ID No.4 |
| 5 | Third subunit + first subunit + second subunit | SEQ ID No.5 |
| 6 | Third subunit + second subunit + first subunit | SEQ ID No.6 |
| 7 | First subunit + third subunit | SEQ ID No.7 |
| 8 | Second subunit + third subunit | SEQ ID No.8 |
| 9 | First subunit + first subunit | SEQ ID No.9 |
| 10 | Second subunit + second subunit | SEQ ID No.10 |
| 11 | Third subunit + third subunit | SEQ ID No.11 |
Preferably, in some embodiments of the present invention, the three subunits of the recombinant novel coronavirus protein are connected in series in the order of the first subunit, the second subunit, and the third subunit.
Preferably, in an embodiment of the present invention, the amino acid sequence of the above recombinant novel coronavirus protein is represented by SEQ ID nos. 1 to 11 or a sequence having 95% or more homology with the amino acid sequence thereof except the mutation site. More preferably, the amino acid sequence of the recombinant novel coronavirus protein is the amino acid sequence shown as SEQ ID No.1 or a sequence with more than 95% homology with the amino acid sequence except the mutation site.
In the above embodiments, the amino acid sequence of SEQ ID Nos. 1 to 11 excluding the mutation site may be substituted, deleted, inserted with 1 or more amino acids to obtain a new amino acid sequence, and the new protein consisting of the amino acid sequence has the same or substantially the same immunological activity as the protein consisting of the amino acid sequence of SEQ ID Nos. 1 to 11, and the new amino acid sequence is also considered to be included in the scope of the present invention.
The sequence having 95% or more homology thereto is an amino acid sequence that is 95%, 96%, 97%, 98% or 99% identical to the amino acid sequence of the recombinant novel coronavirus protein or the fusion protein except for the mutation site. The amino acid sequence of the fusion protein described in the present specification may be subjected to random or engineered point mutation in a suitable manner by those skilled in the art, for example, to obtain better affinity and/or dissociation properties, to improve expression performance, etc., and these sequences may have the same or substantially the same immunological activity as the recombinant novel coronavirus protein or the fusion protein, and these mutated amino acid sequences are included in the scope of the present invention.
The trimeric forms of the above recombinant novel coronavirus proteins may also incorporate certain exogenous trimerization motifs, e.g., T4 bacteriophage fibritin, which are also considered to be within the scope of the present invention. However, the safety of the recombinant coronavirus protein is weaker than that of the recombinant coronavirus protein of the present invention, and an unexpected immune response may occur.
In another aspect of the present invention, there is provided a fusion protein comprising the above recombinant novel coronavirus protein.
Preferably, in certain embodiments of the present invention, the fusion protein further comprises one or more selected from a signal peptide, a tag, or an immune enhancing peptide. The function of the signal peptide can be more favorable for the expression of the protein; the tag may be, for example, a Flag tag, enhanced green fluorescent protein (eGFP), glutathione mercaptotransferase (GST), etc., and the role thereof may be for detection, purification, isolation, etc. The above functional sequences may be used in any combination.
In another aspect of the present invention, there is provided a nucleic acid molecule comprising a nucleotide sequence encoding the above recombinant novel coronavirus protein, or encoding the above fusion protein.
Preferably, in one embodiment of the present invention, the codon of the trimeric protein is optimized by the inventors, and the obtained nucleotide sequence is shown in SEQ ID Nos. 12-22 or a sequence having more than 95% homology thereto.
The above-mentioned sequence having 95% or more homology thereto refers to a nucleotide sequence having 95%, 96%, 97%, 98% or 99% identity to the nucleotide sequence.
The nucleic acid molecule can be prepared by a known technique such as chemical synthesis or PCR amplification based on the nucleotide sequence. In general, the codons encoding the amino acids of the domains described above can be optimized to optimize their expression in a host cell. The information on the above-mentioned nucleotide sequence can be obtained by searching a database such as a known literature or NCBI (https:// www.ncbi.nlm.nih.gov /).
In another aspect of the present invention, there is provided a vector comprising the nucleic acid molecule described above.
In the present invention, the vector may be a linear vector or a cyclic vector. The vector may be a non-viral vector such as a plasmid, a viral vector (for example, an adenovirus vector, a measles virus vector, a mumps virus vector, a rubella virus vector, a variella virus vector, a poliovirus vector, or a yellow fever virus vector), or a vector using a transposon. The vector can contain regulatory sequences such as a promoter, a terminator and the like, and marker sequences such as a drug resistance gene, a reporter gene and the like.
Preferably, in one embodiment of the present invention, the vector is an expression vector of the nucleic acid molecule of the present invention for expressing the recombinant novel coronavirus protein of the present invention.
In another aspect of the present invention, there is provided a host cell comprising the above-described nucleic acid molecule or the above-described vector.
Preferably, in an embodiment of the present invention, the host cell is escherichia coli, a yeast cell, an insect cell, or a mammalian cell;
more preferably, in one embodiment of the present invention, the host cell is a CHO cell.
In another aspect of the present invention, there is provided a method for preparing the above recombinant novel coronavirus protein or the above fusion protein, comprising the steps of:
step A) preparing the nucleic acid molecule, constructing the expression vector, and transforming or transfecting the expression vector into the host cell;
step B) protein expression using the product of step A);
step C) purifying the expression product obtained in step B) to obtain the recombinant novel coronavirus protein or the recombinant novel coronavirus fusion protein.
Wherein, the nucleic acid molecule of the step A) comprises a nucleotide sequence which codes for the recombinant novel coronavirus protein or codes for the fusion protein.
Preferably, in one embodiment of the present invention, the above nucleotide sequence is represented by SEQ ID Nos. 12 to 22 or a sequence having 95% or more homology thereto.
The nucleic acid molecules may be prepared according to the nucleotide sequences described in the present specification using any suitable molecular biological method.
Wherein, the nucleotide sequence can be constructed in the corresponding expression vector of the host cell by any suitable method for constructing the expression vector in step A).
The expression vector is then transformed or transfected into the host cell. Preferably, in one embodiment of the present invention, the inventors constructed recombinant cell lines by transfecting a CHO cell expression vector into HEK293FT cells or CHO cells after constructing the vector.
Wherein, the protein expression in the step B) can express the recombinant protein according to different expression systems. Further, in one embodiment of the present invention, the inventors screened a cell line capable of stably secreting and expressing the recombinant novel coronavirus protein or fusion protein by limiting dilution.
Wherein the purification in step C) may be by any suitable method. For example, salting out, precipitation, dialysis or ultrafiltration, molecular sieve chromatography, ion exchange chromatography, hydrophobic chromatography, affinity chromatography, and the like. Preferably, in one embodiment of the present invention, the recombinant novel coronavirus protein or fusion protein is purified by ion exchange and hydrophobic chromatography.
Of course, according to the prior art, a collection process of the target protein should also be included before the purification step is performed, for example. Collecting cell culture supernatant rich in target protein; the host cell after the target protein has been expressed is disrupted by any suitable disruption method such as ultrasonic disruption, repeated freeze-thaw disruption, and chemical treatment. The above collection of host cells is also understood to be within the scope of such purification.
In another aspect of the present invention, there is provided a use of the above recombinant novel coronavirus protein, the above fusion protein, the above nucleic acid molecule, the above vector or the host cell for the preparation of a medicament for the treatment and/or prevention of a novel coronavirus infection and/or a disease caused by a novel coronavirus.
In another aspect, the present invention provides the use of the above recombinant novel coronavirus protein, the above fusion protein, the above nucleic acid molecule, the above vector or the above host cell for preparing a medicament for enhancing immunity of a population vaccinated with the novel coronavirus vaccine; the novel coronavirus vaccine is preferably a novel coronavirus vaccine inactivated vaccine.
The disease caused by the novel coronavirus is preferably novel coronavirus pneumonia (COVID-19).
In another aspect of the present invention, there is provided a vaccine comprising the above recombinant novel coronavirus protein or the above fusion protein, and an adjuvant.
In an embodiment of the present invention, the vaccine is a recombinant protein vaccine (or a genetically engineered subunit vaccine). Further, in other embodiments of the present invention, the vaccine may be a genetically engineered vector vaccine, or may be a nucleic acid vaccine comprising a nucleotide sequence described in the present specification or encoding an amino acid sequence described in the present specification.
Any suitable adjuvant may be included in the vaccines of the present invention. However, in a preferred embodiment of the present invention, the adjuvant is aluminum hydroxide, aluminum phosphate, MF59, or CpG. More preferably, the adjuvant is aluminum hydroxide.
In another aspect of the present invention, there is provided a method for preparing the vaccine, wherein the purified recombinant coronavirus protein or the fusion protein is mixed with the adjuvant.
In another aspect of the invention there is provided the use of a vaccine as described above in the treatment and/or prevention of a novel coronavirus infection and/or a disease caused by a novel coronavirus.
The disease caused by the novel coronavirus is preferably novel coronavirus pneumonia (COVID-19).
In another aspect of the present invention, a pharmaceutical composition is provided, which comprises the vaccine, and a pharmaceutically acceptable carrier.
The pharmaceutically acceptable carrier may be any pharmaceutically acceptable additive, for example, physiological saline, cell culture medium, glucose, water for injection, glycerol, amino acids and their compositions, stabilizers, surfactants, preservatives, isotonic agents, and the like.
The pharmaceutical composition of the present invention can also be used in combination with other drugs for treating and/or preventing a novel coronavirus infection and/or a disease caused by a novel coronavirus at an effective and safe dose.
In another aspect of the invention, there is provided a method of eliciting an immune response against a novel coronavirus or treating a novel coronavirus infection in a subject by administering an effective dose of said vaccine or said pharmaceutical composition to said subject.
The subject may be a human or other animal.
The administration may be intramuscular, intraperitoneal or subcutaneous.
The invention has the beneficial effects that:
the recombinant novel coronavirus protein provided by the invention covers specific mutation sites of Delta, Beta and Omicron variant RBD, and the recombinant vaccine taking the mutation sites as a target antigen has broad-spectrum protection capability of a cross-epidemic strain.
Drawings
FIG. 1 is a schematic diagram showing the structure of C05G11 protein in example 1 of the present invention;
FIG. 2 is a SDS-PAGE result in example 2 of the present invention, wherein lane 1 is C05G11 protein, and M is protein marker (molecular weight: kDa: 250, 130, 100, 70, 55, 35, 25, 15, 10);
FIG. 3 is a Western-blot identification result chart of the protein purified in example 2 of the present invention, wherein lane 1 is C05G11 protein, and M is protein marker (molecular weight standard: kDa: 250, 130, 100, 70, 55, 35, 25, 15, 10);
FIG. 4 is a graph showing the binding of the recombinant expressed protein to the MM43 neutralizing monoclonal antibody in example 3 of the present invention;
FIG. 5 is a graph showing the binding of the recombinant expressed protein to the MM57 neutralizing monoclonal antibody in example 3 of the present invention;
FIG. 6 is a graph showing the binding of the recombinant expressed protein to MM 117-neutralizing monoclonal antibody in example 3 of the present invention;
FIG. 7 is a graph showing the binding profile of the recombinant expression protein and the R001 neutralizing monoclonal antibody in example 3 of the present invention;
FIG. 8 is a graph showing the binding of the recombinant expression protein to the R117 neutralizing monoclonal antibody in example 3 of the present invention;
FIG. 9 is a graph showing the binding of the recombinant expression protein to the R118 neutralizing monoclonal antibody in example 3 of the present invention;
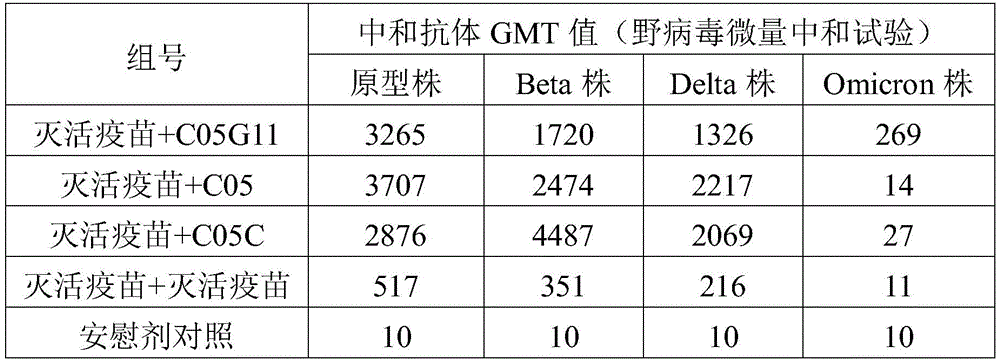
FIG. 10 is a graph showing the results of measuring the neutralizing antibody titer of mouse immune sera using the wild virus microneutralization assay in example 5 of the present invention;
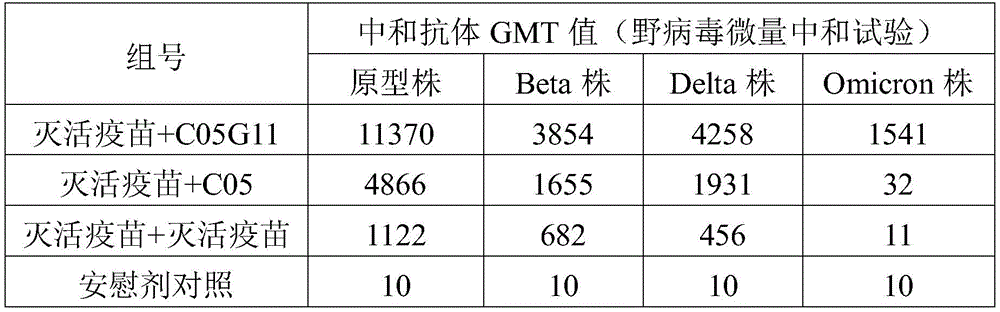
FIG. 11 is a graph showing the results of measuring the neutralizing antibody titer of rat immune serum using the wild virus microneutralization assay in example 6 of the present invention.
DESCRIPTION OF THE SEQUENCES
SEQ ID Nos. 1 to 11 are amino acid sequences of the recombinant novel coronavirus proteins according to the present invention;
SEQ ID No.12 is a nucleotide sequence which codes for the amino acid sequence shown in SEQ ID No.1, namely the nucleotide sequence which codes for the C05G11 protein;
SEQ ID Nos. 13 to 22 are nucleotide sequences encoding amino acid sequences shown in SEQ ID Nos. 2 to 11, respectively;
SEQ ID No.23 is the amino acid sequence of the C05 protein in example 3 of the present invention;
SEQ ID No.24 shows the amino acid sequence of the C05C protein in example 3 of the present invention.
Detailed Description
The invention discloses a recombinant novel coronavirus protein vaccine, a preparation method and application thereof, and can be realized by appropriately improving process parameters by referring to the contents in the text by a person skilled in the art. It is expressly intended that all such alterations and modifications which are obvious to those skilled in the art are deemed to be incorporated herein by reference, and that the techniques of the invention may be practiced and applied by those skilled in the art without departing from the spirit, scope and range of equivalents of the invention.
In the present invention, unless otherwise specified, scientific and technical terms used herein have the meanings that are commonly understood by those skilled in the art. Throughout the specification and claims, unless explicitly stated otherwise, the word "comprise", or variations such as "comprises" or "comprising", will be understood to imply the inclusion of a stated element or component but not the exclusion of any other element or component. Throughout the specification and claims, unless otherwise expressly indicated, the term "RBD" refers to the RBD domain of the spike protein of a novel coronavirus, and is understood to be interchangeable with "RBD" or the RBD domain of a novel coronavirus.
Some terms appearing in the present invention are explained below.
The term "novel coronavirus", SARS-CoV-2, belonging to the order Nedovirales (Nidovirales), Coronaviridae (Coronaviridae), the subfamily orthocoronaviruses, the genus Betaconoviridae, the subgenus Sarbecovirus, the SARS-like virus species, a single-stranded positive-strand RNA virus, having an envelope with a genome of about 29.9kb in length, encoding for the most part non-structural proteins, involved in functions such as viral replication and translation, and a small portion of the sequence encoding a structural protein, such as: s protein (spike protein), M protein (membrane protein), E protein (envelope protein) and N protein (nucleo protein), in addition to several accessory proteins: 3a,3b, p6,7a,7b,8b,9b and orf14, all of which are involved in viral assembly. S, M and the E protein constitute the viral envelope and are the major surface antigens of the virus to elicit an immune response. Wherein the S protein is a transmembrane glycoprotein with a molecular weight of about 150kDa, and forms prominent homotrimers on the surface of the virus. S consists of two functional subunits, cleaved at the boundary between the S1 and S2 subunits (S1/S2 cleavage site), which remain non-covalently associated in the pre-fusion conformation. The S2 subunit is also composed of multiple domains, whose function is primarily to mediate fusion of the virus with the host cell. The distal S1 subunit is structurally divided into four distinct domains: the vaccine is characterized by comprising an N-terminal domain (NTD), a Receptor Binding Domain (RBD), a C-terminal domain 1(CTD1) and a C-terminal domain 2(CTD2), wherein the RBD is mainly responsible for binding with a receptor angiotensin converting enzyme 2 (ACE 2) on the surface of a host cell so as to mediate virus infection on the host cell, so that the S protein and the RBD are main targets for current genetic engineering vaccine development.
The term "trimeric form" is a type of higher structure of proteins. Three protein subunits are contained in the protein, namely the trimer form.
The term "at least one" is understood to mean that two of the three amino acid sequences are identical or that the three amino acid sequences differ from one another.
The term "primary structure" is a linear sequence of amino acids in a peptide or protein. By convention, the primary structure of a protein is from the amino-terminus (N) to the carboxy-terminus (C).
The term "fusion protein" refers to an expression product of one, two or more genes obtained by recombinant DNA techniques. The fusion protein technology is a purposeful gene fusion and protein expression method for obtaining a large number of standard fusion proteins, and a novel target protein with multiple functions can be constructed and expressed by utilizing the fusion protein technology.
The term "vector" is a nucleic acid delivery vehicle into which a polynucleotide may be inserted. When a vector is capable of providing for expression of a protein encoded by an inserted polynucleotide, the vector is referred to as an expression vector. The vector may be introduced into a host cell by transformation, transduction, or transfection, and the genetic material elements carried thereby are expressed in the host cell. Vectors are well known to those skilled in the art and include, but are not limited to: a plasmid; phagemid; a cosmid; artificial chromosomes such as Yeast Artificial Chromosomes (YACs), Bacterial Artificial Chromosomes (BACs), or artificial chromosomes (PACs) derived from P1; bacteriophage such as lambda phage or M13 phage, animal virus, etc. Animal viruses that may be used as vectors include, but are not limited to, retroviruses (including lentiviruses), adenoviruses, adeno-associated viruses, herpes viruses (e.g., herpes simplex virus), poxviruses, baculoviruses, papilloma viruses, papilloma polyoma vacuolatum viruses (e.g., SV 40). A vector may contain a variety of elements that control expression, including, but not limited to, promoter sequences, transcription initiation sequences, enhancer sequences, selection elements, and reporter genes. In addition, the vector may contain a replication initiation site.
The term "host cell" is a cell into which a nucleic acid molecule has been introduced by molecular biological techniques. These techniques include transfection of viral vectors, transformation with plasmid vectors, and introduction of naked DNA by electroporation, lipofection, and particle gun acceleration.
The term "treating" refers to reducing the likelihood of disease pathology, reducing the occurrence of disease symptoms, e.g., to the extent that a subject has a longer survival period or reduced discomfort. Treatment can refer to the ability of a therapy to reduce the symptoms, signs, or causes of a disease when administered to a subject. Treating also refers to alleviating or reducing at least one clinical symptom and/or inhibiting or delaying the progression of a disorder and/or preventing or delaying the onset of a disease or disorder.
The term "subject" refers to any human or other animal, particularly other mammals, that is being prevented, treated, diagnosed. Other mammals may include, for example, dogs, cats, cows, horses, sheep, pigs, goats, rabbits, rats, guinea pigs, mice, and the like. In order to make those skilled in the art better understand the technical solution of the present invention, the following detailed description of the present invention is provided with reference to specific embodiments.
Example 1: novel recombinant coronavirus RBD trimer protein based on protein structure and computational biological design
The new coronavirus is continuously mutated, and the emergence of various variant strains causes multiple rounds of outbreaks of epidemic situations. The world health organization classifies 5 variants as those requiring attention (VOCs) including Alpha (B.1.1.7), Beta (B.1.351), Gamma (P.1), Delta (B.1.617.2) and Omicron (B.1.1.529) according to differences in the transmissibility, pathogenicity or immune escape capacity of the variants. Among these 5 VOCs, numerous studies have demonstrated that the Beta variant has a strong immune escape capacity and the Delta variant has a strong transmission capacity. Preliminary evidence shows that the Omicron variant has strong transmission capability and strong immune escape capability, and the immune escape capability of the Omicron variant is even far higher than that of a Beta strain. The emergence of these strains with a strong immune escape capacity, especially the Omicron variant strains, has raised a great concern about the effectiveness of the existing new corona vaccines. The development of a new generation vaccine with cross-protection capability is an effective means for dealing with these variants. Integrating multiple variants into the same immunogen to construct mosaic antigen molecules, which is a feasible strategy for realizing single-component cross-protection vaccines.
The receptor-binding domain (RBD) of the new coronavirus spike (S) protein is directly involved in the binding of host cell receptors and plays a key role in the process of virus invasion into host cells. Meanwhile, a great deal of research indicates that RBD contains a major neutralizing epitope, and thus, RBD is one of the major target antigens for development of a neocoronary vaccine. However, RBD monomers are not highly immunogenic due to their small molecular size. The natural S protein is of a trimeric structure, and the RBD also exists in a trimerized form. The trimerization RBD is constructed, the natural structural form of the RBD is simulated to the maximum extent, meanwhile, the molecular size of the antigen is increased through trimerization, the repetitive regular arrangement of the antigen is realized, the B cell receptor crosslinking is enhanced, and the immunogenicity of the RBD can be obviously improved. Trimerization of antigen is usually achieved by introducing exogenous linker arms or trimerization motifs, but the introduction of exogenous sequences may bring unexpected immune response with certain safety risk. We achieved trimerization of RBD without introducing a trimerization motif by computational analysis of RBD structure. The RBD has the following structural characteristics: (1) the N-end and the C-end of the RBD are provided with longer flexible loop structures, and the loop structures of the RBD can be used as connecting arms among different RBDs in the trimerization process, so that the introduction of an external connecting arm is avoided, and the trimerization space barrier is reduced; (2) in the natural structure of the S protein, RBDs are relatively independent, strong interaction does not exist between the RBDs and other structural domains, and the folding of the RBDs does not need the assistance of the other structural domains; (3) the RBD has a compact space structure, the core structure of the RBD is composed of a plurality of beta sheet layers, 4 disulfide bonds are contained in the structure domain, the stability of the structure domain is further enhanced, meanwhile, the N-end and the C-end of the RBD are close, a large space barrier does not exist in the trimerization process, and the core structure of the RBD cannot be damaged. Based on the structural characteristics of the RBD, an interception scheme of the RBD is designed, namely an amino acid fragment at 319-537 sites is intercepted, loop structures at two ends of the RBD are kept as much as possible by the interception scheme, and meanwhile, the N-end and C-end interfaces are ensured to be closer. And then, connecting the three intercepted RBD zone segments in series end to form a new RBD trimerization fusion protein.
In order to realize cross protection of different variants of a new coronavirus and construct mosaic type RBD trimerization antigen, a Delta strain with strong infectivity, and Omicron and Beta strains with strongest immune escape are selected from VOCs variants, and RBDs of the three variants are intercepted and assembled together to form mosaic type trimerization RBD protein. Compared with the prototype strain RBD, the Delta strain RBD contains two mutation sites of L452R and T478K, the Beta strain RBD contains three mutation sites of K417N, E484K and N501Y, and the Omicron variant RBD contains 15 mutations of G339D, S371L, S373P, S375F, K417N, N440K, G446S, S477N, T478K, E484A, Q493R, G496S, Q498R, N501Y and Y505H. These mutations not only occur in these three variants, but frequently in many different viral lineages, indicating that these mutations have significant selective advantages in viral evolution, and also suggesting that these mutations may occur independently or in combination again in future mutants.
In this example, RBDs of the Delta variant, the Beta variant and the Omicron variant were serially connected in this order to construct a mosaic-type trimerized RBD fusion protein (C05G11 protein), and the sequence after fusion was shown in SEQ ID No. 1.
By utilizing a homologous modeling method, a mosaic-type trimerized RBD fusion protein space structure is built, and the result is shown in figure 1. The fusion protein is shown to contain three independent RBD domains, and can form an antigen conformation stable trimer form, and the protein covers the mutation sites of Delta, Beta and Omicron variant RBD, and the mutation sites are shown in a ball stick model in figure 1. Theoretically, the recombinant vaccine taking the target antigen as the target antigen is supposed to have broad-spectrum protection capability across epidemic strains, and in addition, compared with the traditional strategy of preparing a plurality of monovalent vaccines and realizing broad-spectrum protection through multivalent combination, the multivalent broad-spectrum protection effect is realized on one antigen molecule, and the method has obvious advantages in the time cost, the economic cost and the vaccine productivity of vaccine preparation.
Example 2: recombinant protein expression, purification and identification
According to the codon preference of a CHO cell expression system, the nucleotide sequence of the coded recombinant protein C05G11 (the amino acid sequence is shown as SEQ ID NO. 1) is subjected to codon optimization, and the optimized nucleotide sequence is shown as SEQ ID NO. 2. The CHO cell expression vector is constructed and then transfected into 293FT cells or CHO cells to construct recombinant cell strains, the cell strains which can stably secrete and express the recombinant protein C05G11 are obtained by screening through a limiting dilution method, the supernatant is harvested after cell culture, and the recombinant protein C05G11 with the purity of more than or equal to 95 percent is obtained after serial chromatography purification. The SDS-PAGE detection result is shown in figure 2, the molecular weight of the protein is 70-100 kD, and meanwhile, part of product related substances such as dimeric protein, monomeric protein and the like can be seen.
The purified C05G11 protein is electrophoresed through SDS-PAGE and then electrically transferred to a PVDF membrane, and Western-blot identification is carried out by using a RBD specific antibody (manufacturer: Beijing Yiqian Shenzhou science and technology Co., Ltd.; product number: 40591-T62; dilution: 2000 times) (the result is shown in figure 3), so that the C05G11 protein can be combined with the RBD specific antibody, and has good biological activity. The purified C05G11 protein is subjected to molecular exclusion chromatography by using a TSKgel G2500PW gel chromatographic column, and the protein purity is more than 90%.
Example 3: detection of physicochemical properties and biological activity of C05G11 protein
The purified C05G11 protein, C05 protein (amino acid sequence is shown as SEQ ID No.23, which is obtained by 293FT cell or CHO cell recombinant expression and chromatography purification) and C05C protein (amino acid sequence is shown as SEQ ID No.24, which is obtained by 293FT cell or CHO cell recombinant expression and chromatography purification), prototype strain RBD protein (manufacturer: Beijing-Yi-Qian-Shenzhou technology Co., Ltd.; product No. 40592-V08B), RBD protein consistent with mutation site of Beta strain virus (K417N, E484K, N501Y; manufacturer: Beijing-Yi-Qian-Shenzhou technology Co., Ltd.; product No. 40592-V08H85), RBD protein consistent with mutation site of Delta strain virus (L R, T638; manufacturer: Beijing-Qian-Shenzhou technology Co., product No. 40592-V02H3), RBD protein consistent with mutation site of Omicron strain virus (G35339, S3576, S46478, S48, S29, S493 32, K46478, K80, K493 32, G496S, Q498R, N501Y, Y505H; the manufacturer: beijing Yi Qiao Shen Tech Co Ltd; the goods number is: 40592-V08H121), respectively diluting to 1.0000 μ g/ml, 0.3333 μ g/ml, 0.1111 μ g/ml, 0.0370 μ g/ml, 0.0123 μ g/ml, 0.0041 μ g/ml, 0.0013 μ g/ml, 0.0004 μ g/ml and 100 μ l/well by using a coating solution, coating the coating solution on a 96-well enzyme label plate, carrying out 4 ℃, 8-12H and taking a blank well as a negative control; adding a sealing liquid after washing the plate by using the PBST solution, and sealing for 2h at 37 ℃; after the PBST solution is washed, an antibody which is diluted to 1 mu g/ml is added into the PBST solution, and specifically comprises an MM43 monoclonal antibody (manufacturer: Beijing Yi Qiao Shen science and technology Limited company; goods number: 40591-MM43), an MM57 monoclonal antibody (manufacturer: Beijing Yi Qiao Shen science and technology Limited company; goods number: 40592-MM57), an MM117 monoclonal antibody (manufacturer: Beijing Yi Qiao Shen science and technology Limited company; goods number: 40592-MM117), an R001 monoclonal antibody (manufacturer: Beijing Yi Qiao Shen science and technology Limited company; goods number: 40592-R001), an R117 monoclonal antibody (manufacturer: Beijing Yi Qiao Shen science and technology Limited company; goods number: 40592-R117), an R118 monoclonal antibody (manufacturer: Beijing Yi Qiao Shen science and technology Limited company; goods number: 40592-R118), 100 mu l/well, and the temperature is 1h at 37 ℃; after washing the plate with the PBST solution, adding a diluted horseradish peroxidase-labeled goat anti-mouse or goat anti-rabbit IgG antibody, incubating at 100 mu l/hole for 1h at 37 ℃; after the PBST solution is washed, adding the developing solutions A and B in sequence, developing at room temperature for 5-10 min, and adding the stop solution C; the readings of the enzyme-linked immunosorbent assay were carried out at two wavelengths (OD450nm and 630nm), cut-off values were determined, and protein concentration-absorbance value curves were plotted.
The results of the activity of binding to MM43 monoclonal antibody are shown in FIG. 4, the results of the activity of binding to MM57 monoclonal antibody are shown in FIG. 5, the results of the activity of binding to MM117 monoclonal antibody are shown in FIG. 6, the results of the activity of binding to R001 monoclonal antibody are shown in FIG. 7, the results of the activity of binding to R117 monoclonal antibody are shown in FIG. 8, and the results of the activity of binding to R118 monoclonal antibody are shown in FIG. 9, and it was found that C05G11 protein has the biological activities of Beta strain, Delta strain and Omicron strain.
Example 4: preparation of recombinant coronavirus vaccine
Diluting the purified recombinant protein C05G11 to 2 times of target antigen concentration, mixing and adsorbing the purified recombinant protein C05G11 with 1.2mg/ml aluminum hydroxide adjuvant according to a ratio of 1:1 (w/w), stirring the mixture on a magnetic stirrer for 40-120 min at a rotating speed of 200-300 rpm to obtain a semi-finished vaccine product, wherein the content of residual protein in supernatant is less than 10% of the content of total protein, and each bottle of the semi-finished vaccine product is aseptically subpackaged according to the loading of 0.5ml to obtain the finished vaccine product.
Example 5: immunological effect evaluation of recombinant novel coronavirus vaccine in mice
The prepared recombinant novel coronavirus vaccine (wherein, the C05G11 protein vaccine is the vaccine prepared in the embodiment 4 of the invention, the C05 protein in the C05 protein vaccine is the recombinant protein in the form of homotrimer formed by 319-537 amino acid fragments of RBD regions of three novel coronavirus original strain S proteins, the C05 protein vaccine is prepared by the same method as the embodiment 4 of the invention), BALB/C mice (purchased from Beijing Wintolite laboratory animal technology Limited company, SPF grade, female, 6-8 weeks old) which are inoculated with 1-time inactivated vaccine (0.5 mu G/dose) are respectively injected into the abdominal cavity to immunize, after 1-time immunization of the inactivated vaccine at 0w, then immunizing 1 needle of recombined novel coronavirus vaccine or inactivated vaccine at the 3w, and collecting blood and separating serum at the 4 w. The test scheme is adopted for the purpose of investigating and simulating the neutralizing capacity of people who are inoculated with the inactivated vaccine and then boosted to a plurality of variant strains.
The neutralizing activity of the mouse serum after immunization against the prototype strain, the Beta strain, the Delta strain and the Omicron strain viruses is detected by using a wild virus micro-neutralization test, the result is shown in figure 10, the GMT value of a serum neutralizing antibody is shown in table 3, the C05G11 protein can generate wide neutralizing activity against various viruses, the neutralizing capacity against the prototype strain, the Beta strain and the Delta strain viruses is equivalent to that of the C05 protein, the neutralizing capacity against the Omicron virus is obviously superior to that of the C05 protein, and the neutralizing activity against the prototype strain, the Beta strain, the Delta strain and the Omicron strain viruses is obviously superior to that of an inactivated vaccine, so that the broad-spectrum protective capacity can be expected.
TABLE 3 neutralizing antibody GMT values in mouse immune sera
Example 6: immunological effect evaluation of recombinant novel coronavirus vaccine in rat body
The recombinant new coronavirus vaccine prepared (wherein, the C05G11 protein vaccine is the vaccine prepared in the embodiment 4 of the invention; the C05 protein in the C05 protein vaccine is the recombinant protein in the form of homotrimer composed of 319 th to 537 th amino acid fragments of RBD region of three new original strains S protein of coronavirus, the C05 protein vaccine is prepared by the same method as the embodiment 4 of the invention; the C05C protein in the C05C protein vaccine is the recombinant protein in the form of heterotrimer composed of 319 th to 537 th amino acid fragments of RBD region of three new strains S protein of coronavirus, wherein two subunits are respectively from Beta strain and Kappa strain, the C05C protein vaccine is prepared by the same method as the embodiment 4 of the invention), Wistar rats which have been inoculated with 1-needle inactivated vaccine (human dose) are respectively immunized by intramuscular injection (purchased from Beijing Withania technologies, SPF grade, female and male, 6-8 weeks old), 10 mu g/dose, 1 needle after the 0w immune inactivated vaccine, 1 needle of recombinant novel coronavirus vaccine or inactivated vaccine is immunized in the 3w, and blood is collected in the 4w to separate serum. The test scheme is adopted for the purpose of investigating and simulating the neutralizing capacity of people who are inoculated with the inactivated vaccine and then boosted to a plurality of variant strains.
The neutralizing activity of the immunized rat serum against the prototype strain, the Beta strain, the Delta strain and the Omicron strain viruses is detected by using a wild virus micro-neutralization test, and the result is shown in figure 11, the GMT value of a serum neutralizing antibody is shown in table 4, the neutralizing activity against various viruses can be generated by the C05G11 protein, the neutralizing capacity against the prototype strain, the Beta strain and the Delta strain viruses is equivalent to the C05 protein and the C05C protein, the neutralizing capacity against the Omicron virus is obviously superior to that against the C05 protein and the C05C protein, and the neutralizing activity against the prototype strain, the Beta strain, the Delta strain and the Omicron strain viruses is obviously superior to that of an inactivated vaccine, and the broad-spectrum protective capacity can be expected.
TABLE 4 neutralizing antibody GMT values in rat immune sera
The foregoing is only a preferred embodiment of the present invention, and it should be noted that, for those skilled in the art, various modifications and decorations can be made without departing from the principle of the present invention, and these modifications and decorations should also be regarded as the protection scope of the present invention.
Sequence listing
<110> national institute of Biotechnology, Biotechnology Limited
<120> a recombinant novel coronavirus protein vaccine, preparation method and application thereof
<130> None
<160> 24
<170> SIPOSequenceListing 1.0
<210> 1
<211> 658
<212> PRT
<213> Artificial
<400> 1
Met Arg Val Gln Pro Thr Glu Ser Ile Val Arg Phe Pro Asn Ile Thr
1 5 10 15
Asn Leu Cys Pro Phe Gly Glu Val Phe Asn Ala Thr Arg Phe Ala Ser
20 25 30
Val Tyr Ala Trp Asn Arg Lys Arg Ile Ser Asn Cys Val Ala Asp Tyr
35 40 45
Ser Val Leu Tyr Asn Ser Ala Ser Phe Ser Thr Phe Lys Cys Tyr Gly
50 55 60
Val Ser Pro Thr Lys Leu Asn Asp Leu Cys Phe Thr Asn Val Tyr Ala
65 70 75 80
Asp Ser Phe Val Ile Arg Gly Asp Glu Val Arg Gln Ile Ala Pro Gly
85 90 95
Gln Thr Gly Lys Ile Ala Asp Tyr Asn Tyr Lys Leu Pro Asp Asp Phe
100 105 110
Thr Gly Cys Val Ile Ala Trp Asn Ser Asn Asn Leu Asp Ser Lys Val
115 120 125
Gly Gly Asn Tyr Asn Tyr Arg Tyr Arg Leu Phe Arg Lys Ser Asn Leu
130 135 140
Lys Pro Phe Glu Arg Asp Ile Ser Thr Glu Ile Tyr Gln Ala Gly Ser
145 150 155 160
Lys Pro Cys Asn Gly Val Glu Gly Phe Asn Cys Tyr Phe Pro Leu Gln
165 170 175
Ser Tyr Gly Phe Gln Pro Thr Asn Gly Val Gly Tyr Gln Pro Tyr Arg
180 185 190
Val Val Val Leu Ser Phe Glu Leu Leu His Ala Pro Ala Thr Val Cys
195 200 205
Gly Pro Lys Lys Ser Thr Asn Leu Val Lys Asn Lys Arg Val Gln Pro
210 215 220
Thr Glu Ser Ile Val Arg Phe Pro Asn Ile Thr Asn Leu Cys Pro Phe
225 230 235 240
Gly Glu Val Phe Asn Ala Thr Arg Phe Ala Ser Val Tyr Ala Trp Asn
245 250 255
Arg Lys Arg Ile Ser Asn Cys Val Ala Asp Tyr Ser Val Leu Tyr Asn
260 265 270
Ser Ala Ser Phe Ser Thr Phe Lys Cys Tyr Gly Val Ser Pro Thr Lys
275 280 285
Leu Asn Asp Leu Cys Phe Thr Asn Val Tyr Ala Asp Ser Phe Val Ile
290 295 300
Arg Gly Asp Glu Val Arg Gln Ile Ala Pro Gly Gln Thr Gly Asn Ile
305 310 315 320
Ala Asp Tyr Asn Tyr Lys Leu Pro Asp Asp Phe Thr Gly Cys Val Ile
325 330 335
Ala Trp Asn Ser Asn Asn Leu Asp Ser Lys Val Gly Gly Asn Tyr Asn
340 345 350
Tyr Leu Tyr Arg Leu Phe Arg Lys Ser Asn Leu Lys Pro Phe Glu Arg
355 360 365
Asp Ile Ser Thr Glu Ile Tyr Gln Ala Gly Ser Thr Pro Cys Asn Gly
370 375 380
Val Lys Gly Phe Asn Cys Tyr Phe Pro Leu Gln Ser Tyr Gly Phe Gln
385 390 395 400
Pro Thr Tyr Gly Val Gly Tyr Gln Pro Tyr Arg Val Val Val Leu Ser
405 410 415
Phe Glu Leu Leu His Ala Pro Ala Thr Val Cys Gly Pro Lys Lys Ser
420 425 430
Thr Asn Leu Val Lys Asn Lys Arg Val Gln Pro Thr Glu Ser Ile Val
435 440 445
Arg Phe Pro Asn Ile Thr Asn Leu Cys Pro Phe Asp Glu Val Phe Asn
450 455 460
Ala Thr Arg Phe Ala Ser Val Tyr Ala Trp Asn Arg Lys Arg Ile Ser
465 470 475 480
Asn Cys Val Ala Asp Tyr Ser Val Leu Tyr Asn Leu Ala Pro Phe Phe
485 490 495
Thr Phe Lys Cys Tyr Gly Val Ser Pro Thr Lys Leu Asn Asp Leu Cys
500 505 510
Phe Thr Asn Val Tyr Ala Asp Ser Phe Val Ile Arg Gly Asp Glu Val
515 520 525
Arg Gln Ile Ala Pro Gly Gln Thr Gly Asn Ile Ala Asp Tyr Asn Tyr
530 535 540
Lys Leu Pro Asp Asp Phe Thr Gly Cys Val Ile Ala Trp Asn Ser Asn
545 550 555 560
Lys Leu Asp Ser Lys Val Ser Gly Asn Tyr Asn Tyr Leu Tyr Arg Leu
565 570 575
Phe Arg Lys Ser Asn Leu Lys Pro Phe Glu Arg Asp Ile Ser Thr Glu
580 585 590
Ile Tyr Gln Ala Gly Asn Lys Pro Cys Asn Gly Val Ala Gly Phe Asn
595 600 605
Cys Tyr Phe Pro Leu Arg Ser Tyr Ser Phe Arg Pro Thr Tyr Gly Val
610 615 620
Gly His Gln Pro Tyr Arg Val Val Val Leu Ser Phe Glu Leu Leu His
625 630 635 640
Ala Pro Ala Thr Val Cys Gly Pro Lys Lys Ser Thr Asn Leu Val Lys
645 650 655
Asn Lys
<210> 2
<211> 657
<212> PRT
<213> Artificial
<400> 2
Arg Val Gln Pro Thr Glu Ser Ile Val Arg Phe Pro Asn Ile Thr Asn
1 5 10 15
Leu Cys Pro Phe Gly Glu Val Phe Asn Ala Thr Arg Phe Ala Ser Val
20 25 30
Tyr Ala Trp Asn Arg Lys Arg Ile Ser Asn Cys Val Ala Asp Tyr Ser
35 40 45
Val Leu Tyr Asn Ser Ala Ser Phe Ser Thr Phe Lys Cys Tyr Gly Val
50 55 60
Ser Pro Thr Lys Leu Asn Asp Leu Cys Phe Thr Asn Val Tyr Ala Asp
65 70 75 80
Ser Phe Val Ile Arg Gly Asp Glu Val Arg Gln Ile Ala Pro Gly Gln
85 90 95
Thr Gly Lys Ile Ala Asp Tyr Asn Tyr Lys Leu Pro Asp Asp Phe Thr
100 105 110
Gly Cys Val Ile Ala Trp Asn Ser Asn Asn Leu Asp Ser Lys Val Gly
115 120 125
Gly Asn Tyr Asn Tyr Arg Tyr Arg Leu Phe Arg Lys Ser Asn Leu Lys
130 135 140
Pro Phe Glu Arg Asp Ile Ser Thr Glu Ile Tyr Gln Ala Gly Ser Lys
145 150 155 160
Pro Cys Asn Gly Val Glu Gly Phe Asn Cys Tyr Phe Pro Leu Gln Ser
165 170 175
Tyr Gly Phe Gln Pro Thr Asn Gly Val Gly Tyr Gln Pro Tyr Arg Val
180 185 190
Val Val Leu Ser Phe Glu Leu Leu His Ala Pro Ala Thr Val Cys Gly
195 200 205
Pro Lys Lys Ser Thr Asn Leu Val Lys Asn Lys Arg Val Gln Pro Thr
210 215 220
Glu Ser Ile Val Arg Phe Pro Asn Ile Thr Asn Leu Cys Pro Phe Asp
225 230 235 240
Glu Val Phe Asn Ala Thr Arg Phe Ala Ser Val Tyr Ala Trp Asn Arg
245 250 255
Lys Arg Ile Ser Asn Cys Val Ala Asp Tyr Ser Val Leu Tyr Asn Leu
260 265 270
Ala Pro Phe Phe Thr Phe Lys Cys Tyr Gly Val Ser Pro Thr Lys Leu
275 280 285
Asn Asp Leu Cys Phe Thr Asn Val Tyr Ala Asp Ser Phe Val Ile Arg
290 295 300
Gly Asp Glu Val Arg Gln Ile Ala Pro Gly Gln Thr Gly Asn Ile Ala
305 310 315 320
Asp Tyr Asn Tyr Lys Leu Pro Asp Asp Phe Thr Gly Cys Val Ile Ala
325 330 335
Trp Asn Ser Asn Lys Leu Asp Ser Lys Val Ser Gly Asn Tyr Asn Tyr
340 345 350
Leu Tyr Arg Leu Phe Arg Lys Ser Asn Leu Lys Pro Phe Glu Arg Asp
355 360 365
Ile Ser Thr Glu Ile Tyr Gln Ala Gly Asn Lys Pro Cys Asn Gly Val
370 375 380
Ala Gly Phe Asn Cys Tyr Phe Pro Leu Arg Ser Tyr Ser Phe Arg Pro
385 390 395 400
Thr Tyr Gly Val Gly His Gln Pro Tyr Arg Val Val Val Leu Ser Phe
405 410 415
Glu Leu Leu His Ala Pro Ala Thr Val Cys Gly Pro Lys Lys Ser Thr
420 425 430
Asn Leu Val Lys Asn Lys Arg Val Gln Pro Thr Glu Ser Ile Val Arg
435 440 445
Phe Pro Asn Ile Thr Asn Leu Cys Pro Phe Gly Glu Val Phe Asn Ala
450 455 460
Thr Arg Phe Ala Ser Val Tyr Ala Trp Asn Arg Lys Arg Ile Ser Asn
465 470 475 480
Cys Val Ala Asp Tyr Ser Val Leu Tyr Asn Ser Ala Ser Phe Ser Thr
485 490 495
Phe Lys Cys Tyr Gly Val Ser Pro Thr Lys Leu Asn Asp Leu Cys Phe
500 505 510
Thr Asn Val Tyr Ala Asp Ser Phe Val Ile Arg Gly Asp Glu Val Arg
515 520 525
Gln Ile Ala Pro Gly Gln Thr Gly Asn Ile Ala Asp Tyr Asn Tyr Lys
530 535 540
Leu Pro Asp Asp Phe Thr Gly Cys Val Ile Ala Trp Asn Ser Asn Asn
545 550 555 560
Leu Asp Ser Lys Val Gly Gly Asn Tyr Asn Tyr Leu Tyr Arg Leu Phe
565 570 575
Arg Lys Ser Asn Leu Lys Pro Phe Glu Arg Asp Ile Ser Thr Glu Ile
580 585 590
Tyr Gln Ala Gly Ser Thr Pro Cys Asn Gly Val Lys Gly Phe Asn Cys
595 600 605
Tyr Phe Pro Leu Gln Ser Tyr Gly Phe Gln Pro Thr Tyr Gly Val Gly
610 615 620
Tyr Gln Pro Tyr Arg Val Val Val Leu Ser Phe Glu Leu Leu His Ala
625 630 635 640
Pro Ala Thr Val Cys Gly Pro Lys Lys Ser Thr Asn Leu Val Lys Asn
645 650 655
Lys
<210> 3
<211> 657
<212> PRT
<213> Artificial
<400> 3
Arg Val Gln Pro Thr Glu Ser Ile Val Arg Phe Pro Asn Ile Thr Asn
1 5 10 15
Leu Cys Pro Phe Gly Glu Val Phe Asn Ala Thr Arg Phe Ala Ser Val
20 25 30
Tyr Ala Trp Asn Arg Lys Arg Ile Ser Asn Cys Val Ala Asp Tyr Ser
35 40 45
Val Leu Tyr Asn Ser Ala Ser Phe Ser Thr Phe Lys Cys Tyr Gly Val
50 55 60
Ser Pro Thr Lys Leu Asn Asp Leu Cys Phe Thr Asn Val Tyr Ala Asp
65 70 75 80
Ser Phe Val Ile Arg Gly Asp Glu Val Arg Gln Ile Ala Pro Gly Gln
85 90 95
Thr Gly Asn Ile Ala Asp Tyr Asn Tyr Lys Leu Pro Asp Asp Phe Thr
100 105 110
Gly Cys Val Ile Ala Trp Asn Ser Asn Asn Leu Asp Ser Lys Val Gly
115 120 125
Gly Asn Tyr Asn Tyr Leu Tyr Arg Leu Phe Arg Lys Ser Asn Leu Lys
130 135 140
Pro Phe Glu Arg Asp Ile Ser Thr Glu Ile Tyr Gln Ala Gly Ser Thr
145 150 155 160
Pro Cys Asn Gly Val Lys Gly Phe Asn Cys Tyr Phe Pro Leu Gln Ser
165 170 175
Tyr Gly Phe Gln Pro Thr Tyr Gly Val Gly Tyr Gln Pro Tyr Arg Val
180 185 190
Val Val Leu Ser Phe Glu Leu Leu His Ala Pro Ala Thr Val Cys Gly
195 200 205
Pro Lys Lys Ser Thr Asn Leu Val Lys Asn Lys Arg Val Gln Pro Thr
210 215 220
Glu Ser Ile Val Arg Phe Pro Asn Ile Thr Asn Leu Cys Pro Phe Gly
225 230 235 240
Glu Val Phe Asn Ala Thr Arg Phe Ala Ser Val Tyr Ala Trp Asn Arg
245 250 255
Lys Arg Ile Ser Asn Cys Val Ala Asp Tyr Ser Val Leu Tyr Asn Ser
260 265 270
Ala Ser Phe Ser Thr Phe Lys Cys Tyr Gly Val Ser Pro Thr Lys Leu
275 280 285
Asn Asp Leu Cys Phe Thr Asn Val Tyr Ala Asp Ser Phe Val Ile Arg
290 295 300
Gly Asp Glu Val Arg Gln Ile Ala Pro Gly Gln Thr Gly Lys Ile Ala
305 310 315 320
Asp Tyr Asn Tyr Lys Leu Pro Asp Asp Phe Thr Gly Cys Val Ile Ala
325 330 335
Trp Asn Ser Asn Asn Leu Asp Ser Lys Val Gly Gly Asn Tyr Asn Tyr
340 345 350
Arg Tyr Arg Leu Phe Arg Lys Ser Asn Leu Lys Pro Phe Glu Arg Asp
355 360 365
Ile Ser Thr Glu Ile Tyr Gln Ala Gly Ser Lys Pro Cys Asn Gly Val
370 375 380
Glu Gly Phe Asn Cys Tyr Phe Pro Leu Gln Ser Tyr Gly Phe Gln Pro
385 390 395 400
Thr Asn Gly Val Gly Tyr Gln Pro Tyr Arg Val Val Val Leu Ser Phe
405 410 415
Glu Leu Leu His Ala Pro Ala Thr Val Cys Gly Pro Lys Lys Ser Thr
420 425 430
Asn Leu Val Lys Asn Lys Arg Val Gln Pro Thr Glu Ser Ile Val Arg
435 440 445
Phe Pro Asn Ile Thr Asn Leu Cys Pro Phe Asp Glu Val Phe Asn Ala
450 455 460
Thr Arg Phe Ala Ser Val Tyr Ala Trp Asn Arg Lys Arg Ile Ser Asn
465 470 475 480
Cys Val Ala Asp Tyr Ser Val Leu Tyr Asn Leu Ala Pro Phe Phe Thr
485 490 495
Phe Lys Cys Tyr Gly Val Ser Pro Thr Lys Leu Asn Asp Leu Cys Phe
500 505 510
Thr Asn Val Tyr Ala Asp Ser Phe Val Ile Arg Gly Asp Glu Val Arg
515 520 525
Gln Ile Ala Pro Gly Gln Thr Gly Asn Ile Ala Asp Tyr Asn Tyr Lys
530 535 540
Leu Pro Asp Asp Phe Thr Gly Cys Val Ile Ala Trp Asn Ser Asn Lys
545 550 555 560
Leu Asp Ser Lys Val Ser Gly Asn Tyr Asn Tyr Leu Tyr Arg Leu Phe
565 570 575
Arg Lys Ser Asn Leu Lys Pro Phe Glu Arg Asp Ile Ser Thr Glu Ile
580 585 590
Tyr Gln Ala Gly Asn Lys Pro Cys Asn Gly Val Ala Gly Phe Asn Cys
595 600 605
Tyr Phe Pro Leu Arg Ser Tyr Ser Phe Arg Pro Thr Tyr Gly Val Gly
610 615 620
His Gln Pro Tyr Arg Val Val Val Leu Ser Phe Glu Leu Leu His Ala
625 630 635 640
Pro Ala Thr Val Cys Gly Pro Lys Lys Ser Thr Asn Leu Val Lys Asn
645 650 655
Lys
<210> 4
<211> 657
<212> PRT
<213> Artificial
<400> 4
Arg Val Gln Pro Thr Glu Ser Ile Val Arg Phe Pro Asn Ile Thr Asn
1 5 10 15
Leu Cys Pro Phe Gly Glu Val Phe Asn Ala Thr Arg Phe Ala Ser Val
20 25 30
Tyr Ala Trp Asn Arg Lys Arg Ile Ser Asn Cys Val Ala Asp Tyr Ser
35 40 45
Val Leu Tyr Asn Ser Ala Ser Phe Ser Thr Phe Lys Cys Tyr Gly Val
50 55 60
Ser Pro Thr Lys Leu Asn Asp Leu Cys Phe Thr Asn Val Tyr Ala Asp
65 70 75 80
Ser Phe Val Ile Arg Gly Asp Glu Val Arg Gln Ile Ala Pro Gly Gln
85 90 95
Thr Gly Asn Ile Ala Asp Tyr Asn Tyr Lys Leu Pro Asp Asp Phe Thr
100 105 110
Gly Cys Val Ile Ala Trp Asn Ser Asn Asn Leu Asp Ser Lys Val Gly
115 120 125
Gly Asn Tyr Asn Tyr Leu Tyr Arg Leu Phe Arg Lys Ser Asn Leu Lys
130 135 140
Pro Phe Glu Arg Asp Ile Ser Thr Glu Ile Tyr Gln Ala Gly Ser Thr
145 150 155 160
Pro Cys Asn Gly Val Lys Gly Phe Asn Cys Tyr Phe Pro Leu Gln Ser
165 170 175
Tyr Gly Phe Gln Pro Thr Tyr Gly Val Gly Tyr Gln Pro Tyr Arg Val
180 185 190
Val Val Leu Ser Phe Glu Leu Leu His Ala Pro Ala Thr Val Cys Gly
195 200 205
Pro Lys Lys Ser Thr Asn Leu Val Lys Asn Lys Arg Val Gln Pro Thr
210 215 220
Glu Ser Ile Val Arg Phe Pro Asn Ile Thr Asn Leu Cys Pro Phe Asp
225 230 235 240
Glu Val Phe Asn Ala Thr Arg Phe Ala Ser Val Tyr Ala Trp Asn Arg
245 250 255
Lys Arg Ile Ser Asn Cys Val Ala Asp Tyr Ser Val Leu Tyr Asn Leu
260 265 270
Ala Pro Phe Phe Thr Phe Lys Cys Tyr Gly Val Ser Pro Thr Lys Leu
275 280 285
Asn Asp Leu Cys Phe Thr Asn Val Tyr Ala Asp Ser Phe Val Ile Arg
290 295 300
Gly Asp Glu Val Arg Gln Ile Ala Pro Gly Gln Thr Gly Asn Ile Ala
305 310 315 320
Asp Tyr Asn Tyr Lys Leu Pro Asp Asp Phe Thr Gly Cys Val Ile Ala
325 330 335
Trp Asn Ser Asn Lys Leu Asp Ser Lys Val Ser Gly Asn Tyr Asn Tyr
340 345 350
Leu Tyr Arg Leu Phe Arg Lys Ser Asn Leu Lys Pro Phe Glu Arg Asp
355 360 365
Ile Ser Thr Glu Ile Tyr Gln Ala Gly Asn Lys Pro Cys Asn Gly Val
370 375 380
Ala Gly Phe Asn Cys Tyr Phe Pro Leu Arg Ser Tyr Ser Phe Arg Pro
385 390 395 400
Thr Tyr Gly Val Gly His Gln Pro Tyr Arg Val Val Val Leu Ser Phe
405 410 415
Glu Leu Leu His Ala Pro Ala Thr Val Cys Gly Pro Lys Lys Ser Thr
420 425 430
Asn Leu Val Lys Asn Lys Arg Val Gln Pro Thr Glu Ser Ile Val Arg
435 440 445
Phe Pro Asn Ile Thr Asn Leu Cys Pro Phe Gly Glu Val Phe Asn Ala
450 455 460
Thr Arg Phe Ala Ser Val Tyr Ala Trp Asn Arg Lys Arg Ile Ser Asn
465 470 475 480
Cys Val Ala Asp Tyr Ser Val Leu Tyr Asn Ser Ala Ser Phe Ser Thr
485 490 495
Phe Lys Cys Tyr Gly Val Ser Pro Thr Lys Leu Asn Asp Leu Cys Phe
500 505 510
Thr Asn Val Tyr Ala Asp Ser Phe Val Ile Arg Gly Asp Glu Val Arg
515 520 525
Gln Ile Ala Pro Gly Gln Thr Gly Lys Ile Ala Asp Tyr Asn Tyr Lys
530 535 540
Leu Pro Asp Asp Phe Thr Gly Cys Val Ile Ala Trp Asn Ser Asn Asn
545 550 555 560
Leu Asp Ser Lys Val Gly Gly Asn Tyr Asn Tyr Arg Tyr Arg Leu Phe
565 570 575
Arg Lys Ser Asn Leu Lys Pro Phe Glu Arg Asp Ile Ser Thr Glu Ile
580 585 590
Tyr Gln Ala Gly Ser Lys Pro Cys Asn Gly Val Glu Gly Phe Asn Cys
595 600 605
Tyr Phe Pro Leu Gln Ser Tyr Gly Phe Gln Pro Thr Asn Gly Val Gly
610 615 620
Tyr Gln Pro Tyr Arg Val Val Val Leu Ser Phe Glu Leu Leu His Ala
625 630 635 640
Pro Ala Thr Val Cys Gly Pro Lys Lys Ser Thr Asn Leu Val Lys Asn
645 650 655
Lys
<210> 5
<211> 657
<212> PRT
<213> Artificial
<400> 5
Arg Val Gln Pro Thr Glu Ser Ile Val Arg Phe Pro Asn Ile Thr Asn
1 5 10 15
Leu Cys Pro Phe Asp Glu Val Phe Asn Ala Thr Arg Phe Ala Ser Val
20 25 30
Tyr Ala Trp Asn Arg Lys Arg Ile Ser Asn Cys Val Ala Asp Tyr Ser
35 40 45
Val Leu Tyr Asn Leu Ala Pro Phe Phe Thr Phe Lys Cys Tyr Gly Val
50 55 60
Ser Pro Thr Lys Leu Asn Asp Leu Cys Phe Thr Asn Val Tyr Ala Asp
65 70 75 80
Ser Phe Val Ile Arg Gly Asp Glu Val Arg Gln Ile Ala Pro Gly Gln
85 90 95
Thr Gly Asn Ile Ala Asp Tyr Asn Tyr Lys Leu Pro Asp Asp Phe Thr
100 105 110
Gly Cys Val Ile Ala Trp Asn Ser Asn Lys Leu Asp Ser Lys Val Ser
115 120 125
Gly Asn Tyr Asn Tyr Leu Tyr Arg Leu Phe Arg Lys Ser Asn Leu Lys
130 135 140
Pro Phe Glu Arg Asp Ile Ser Thr Glu Ile Tyr Gln Ala Gly Asn Lys
145 150 155 160
Pro Cys Asn Gly Val Ala Gly Phe Asn Cys Tyr Phe Pro Leu Arg Ser
165 170 175
Tyr Ser Phe Arg Pro Thr Tyr Gly Val Gly His Gln Pro Tyr Arg Val
180 185 190
Val Val Leu Ser Phe Glu Leu Leu His Ala Pro Ala Thr Val Cys Gly
195 200 205
Pro Lys Lys Ser Thr Asn Leu Val Lys Asn Lys Arg Val Gln Pro Thr
210 215 220
Glu Ser Ile Val Arg Phe Pro Asn Ile Thr Asn Leu Cys Pro Phe Gly
225 230 235 240
Glu Val Phe Asn Ala Thr Arg Phe Ala Ser Val Tyr Ala Trp Asn Arg
245 250 255
Lys Arg Ile Ser Asn Cys Val Ala Asp Tyr Ser Val Leu Tyr Asn Ser
260 265 270
Ala Ser Phe Ser Thr Phe Lys Cys Tyr Gly Val Ser Pro Thr Lys Leu
275 280 285
Asn Asp Leu Cys Phe Thr Asn Val Tyr Ala Asp Ser Phe Val Ile Arg
290 295 300
Gly Asp Glu Val Arg Gln Ile Ala Pro Gly Gln Thr Gly Lys Ile Ala
305 310 315 320
Asp Tyr Asn Tyr Lys Leu Pro Asp Asp Phe Thr Gly Cys Val Ile Ala
325 330 335
Trp Asn Ser Asn Asn Leu Asp Ser Lys Val Gly Gly Asn Tyr Asn Tyr
340 345 350
Arg Tyr Arg Leu Phe Arg Lys Ser Asn Leu Lys Pro Phe Glu Arg Asp
355 360 365
Ile Ser Thr Glu Ile Tyr Gln Ala Gly Ser Lys Pro Cys Asn Gly Val
370 375 380
Glu Gly Phe Asn Cys Tyr Phe Pro Leu Gln Ser Tyr Gly Phe Gln Pro
385 390 395 400
Thr Asn Gly Val Gly Tyr Gln Pro Tyr Arg Val Val Val Leu Ser Phe
405 410 415
Glu Leu Leu His Ala Pro Ala Thr Val Cys Gly Pro Lys Lys Ser Thr
420 425 430
Asn Leu Val Lys Asn Lys Arg Val Gln Pro Thr Glu Ser Ile Val Arg
435 440 445
Phe Pro Asn Ile Thr Asn Leu Cys Pro Phe Gly Glu Val Phe Asn Ala
450 455 460
Thr Arg Phe Ala Ser Val Tyr Ala Trp Asn Arg Lys Arg Ile Ser Asn
465 470 475 480
Cys Val Ala Asp Tyr Ser Val Leu Tyr Asn Ser Ala Ser Phe Ser Thr
485 490 495
Phe Lys Cys Tyr Gly Val Ser Pro Thr Lys Leu Asn Asp Leu Cys Phe
500 505 510
Thr Asn Val Tyr Ala Asp Ser Phe Val Ile Arg Gly Asp Glu Val Arg
515 520 525
Gln Ile Ala Pro Gly Gln Thr Gly Asn Ile Ala Asp Tyr Asn Tyr Lys
530 535 540
Leu Pro Asp Asp Phe Thr Gly Cys Val Ile Ala Trp Asn Ser Asn Asn
545 550 555 560
Leu Asp Ser Lys Val Gly Gly Asn Tyr Asn Tyr Leu Tyr Arg Leu Phe
565 570 575
Arg Lys Ser Asn Leu Lys Pro Phe Glu Arg Asp Ile Ser Thr Glu Ile
580 585 590
Tyr Gln Ala Gly Ser Thr Pro Cys Asn Gly Val Lys Gly Phe Asn Cys
595 600 605
Tyr Phe Pro Leu Gln Ser Tyr Gly Phe Gln Pro Thr Tyr Gly Val Gly
610 615 620
Tyr Gln Pro Tyr Arg Val Val Val Leu Ser Phe Glu Leu Leu His Ala
625 630 635 640
Pro Ala Thr Val Cys Gly Pro Lys Lys Ser Thr Asn Leu Val Lys Asn
645 650 655
Lys
<210> 6
<211> 657
<212> PRT
<213> Artificial
<400> 6
Arg Val Gln Pro Thr Glu Ser Ile Val Arg Phe Pro Asn Ile Thr Asn
1 5 10 15
Leu Cys Pro Phe Asp Glu Val Phe Asn Ala Thr Arg Phe Ala Ser Val
20 25 30
Tyr Ala Trp Asn Arg Lys Arg Ile Ser Asn Cys Val Ala Asp Tyr Ser
35 40 45
Val Leu Tyr Asn Leu Ala Pro Phe Phe Thr Phe Lys Cys Tyr Gly Val
50 55 60
Ser Pro Thr Lys Leu Asn Asp Leu Cys Phe Thr Asn Val Tyr Ala Asp
65 70 75 80
Ser Phe Val Ile Arg Gly Asp Glu Val Arg Gln Ile Ala Pro Gly Gln
85 90 95
Thr Gly Asn Ile Ala Asp Tyr Asn Tyr Lys Leu Pro Asp Asp Phe Thr
100 105 110
Gly Cys Val Ile Ala Trp Asn Ser Asn Lys Leu Asp Ser Lys Val Ser
115 120 125
Gly Asn Tyr Asn Tyr Leu Tyr Arg Leu Phe Arg Lys Ser Asn Leu Lys
130 135 140
Pro Phe Glu Arg Asp Ile Ser Thr Glu Ile Tyr Gln Ala Gly Asn Lys
145 150 155 160
Pro Cys Asn Gly Val Ala Gly Phe Asn Cys Tyr Phe Pro Leu Arg Ser
165 170 175
Tyr Ser Phe Arg Pro Thr Tyr Gly Val Gly His Gln Pro Tyr Arg Val
180 185 190
Val Val Leu Ser Phe Glu Leu Leu His Ala Pro Ala Thr Val Cys Gly
195 200 205
Pro Lys Lys Ser Thr Asn Leu Val Lys Asn Lys Arg Val Gln Pro Thr
210 215 220
Glu Ser Ile Val Arg Phe Pro Asn Ile Thr Asn Leu Cys Pro Phe Gly
225 230 235 240
Glu Val Phe Asn Ala Thr Arg Phe Ala Ser Val Tyr Ala Trp Asn Arg
245 250 255
Lys Arg Ile Ser Asn Cys Val Ala Asp Tyr Ser Val Leu Tyr Asn Ser
260 265 270
Ala Ser Phe Ser Thr Phe Lys Cys Tyr Gly Val Ser Pro Thr Lys Leu
275 280 285
Asn Asp Leu Cys Phe Thr Asn Val Tyr Ala Asp Ser Phe Val Ile Arg
290 295 300
Gly Asp Glu Val Arg Gln Ile Ala Pro Gly Gln Thr Gly Asn Ile Ala
305 310 315 320
Asp Tyr Asn Tyr Lys Leu Pro Asp Asp Phe Thr Gly Cys Val Ile Ala
325 330 335
Trp Asn Ser Asn Asn Leu Asp Ser Lys Val Gly Gly Asn Tyr Asn Tyr
340 345 350
Leu Tyr Arg Leu Phe Arg Lys Ser Asn Leu Lys Pro Phe Glu Arg Asp
355 360 365
Ile Ser Thr Glu Ile Tyr Gln Ala Gly Ser Thr Pro Cys Asn Gly Val
370 375 380
Lys Gly Phe Asn Cys Tyr Phe Pro Leu Gln Ser Tyr Gly Phe Gln Pro
385 390 395 400
Thr Tyr Gly Val Gly Tyr Gln Pro Tyr Arg Val Val Val Leu Ser Phe
405 410 415
Glu Leu Leu His Ala Pro Ala Thr Val Cys Gly Pro Lys Lys Ser Thr
420 425 430
Asn Leu Val Lys Asn Lys Arg Val Gln Pro Thr Glu Ser Ile Val Arg
435 440 445
Phe Pro Asn Ile Thr Asn Leu Cys Pro Phe Gly Glu Val Phe Asn Ala
450 455 460
Thr Arg Phe Ala Ser Val Tyr Ala Trp Asn Arg Lys Arg Ile Ser Asn
465 470 475 480
Cys Val Ala Asp Tyr Ser Val Leu Tyr Asn Ser Ala Ser Phe Ser Thr
485 490 495
Phe Lys Cys Tyr Gly Val Ser Pro Thr Lys Leu Asn Asp Leu Cys Phe
500 505 510
Thr Asn Val Tyr Ala Asp Ser Phe Val Ile Arg Gly Asp Glu Val Arg
515 520 525
Gln Ile Ala Pro Gly Gln Thr Gly Lys Ile Ala Asp Tyr Asn Tyr Lys
530 535 540
Leu Pro Asp Asp Phe Thr Gly Cys Val Ile Ala Trp Asn Ser Asn Asn
545 550 555 560
Leu Asp Ser Lys Val Gly Gly Asn Tyr Asn Tyr Arg Tyr Arg Leu Phe
565 570 575
Arg Lys Ser Asn Leu Lys Pro Phe Glu Arg Asp Ile Ser Thr Glu Ile
580 585 590
Tyr Gln Ala Gly Ser Lys Pro Cys Asn Gly Val Glu Gly Phe Asn Cys
595 600 605
Tyr Phe Pro Leu Gln Ser Tyr Gly Phe Gln Pro Thr Asn Gly Val Gly
610 615 620
Tyr Gln Pro Tyr Arg Val Val Val Leu Ser Phe Glu Leu Leu His Ala
625 630 635 640
Pro Ala Thr Val Cys Gly Pro Lys Lys Ser Thr Asn Leu Val Lys Asn
645 650 655
Lys
<210> 7
<211> 657
<212> PRT
<213> Artificial
<400> 7
Arg Val Gln Pro Thr Glu Ser Ile Val Arg Phe Pro Asn Ile Thr Asn
1 5 10 15
Leu Cys Pro Phe Gly Glu Val Phe Asn Ala Thr Arg Phe Ala Ser Val
20 25 30
Tyr Ala Trp Asn Arg Lys Arg Ile Ser Asn Cys Val Ala Asp Tyr Ser
35 40 45
Val Leu Tyr Asn Ser Ala Ser Phe Ser Thr Phe Lys Cys Tyr Gly Val
50 55 60
Ser Pro Thr Lys Leu Asn Asp Leu Cys Phe Thr Asn Val Tyr Ala Asp
65 70 75 80
Ser Phe Val Ile Arg Gly Asp Glu Val Arg Gln Ile Ala Pro Gly Gln
85 90 95
Thr Gly Lys Ile Ala Asp Tyr Asn Tyr Lys Leu Pro Asp Asp Phe Thr
100 105 110
Gly Cys Val Ile Ala Trp Asn Ser Asn Asn Leu Asp Ser Lys Val Gly
115 120 125
Gly Asn Tyr Asn Tyr Arg Tyr Arg Leu Phe Arg Lys Ser Asn Leu Lys
130 135 140
Pro Phe Glu Arg Asp Ile Ser Thr Glu Ile Tyr Gln Ala Gly Ser Lys
145 150 155 160
Pro Cys Asn Gly Val Glu Gly Phe Asn Cys Tyr Phe Pro Leu Gln Ser
165 170 175
Tyr Gly Phe Gln Pro Thr Asn Gly Val Gly Tyr Gln Pro Tyr Arg Val
180 185 190
Val Val Leu Ser Phe Glu Leu Leu His Ala Pro Ala Thr Val Cys Gly
195 200 205
Pro Lys Lys Ser Thr Asn Leu Val Lys Asn Lys Arg Val Gln Pro Thr
210 215 220
Glu Ser Ile Val Arg Phe Pro Asn Ile Thr Asn Leu Cys Pro Phe Gly
225 230 235 240
Glu Val Phe Asn Ala Thr Arg Phe Ala Ser Val Tyr Ala Trp Asn Arg
245 250 255
Lys Arg Ile Ser Asn Cys Val Ala Asp Tyr Ser Val Leu Tyr Asn Ser
260 265 270
Ala Ser Phe Ser Thr Phe Lys Cys Tyr Gly Val Ser Pro Thr Lys Leu
275 280 285
Asn Asp Leu Cys Phe Thr Asn Val Tyr Ala Asp Ser Phe Val Ile Arg
290 295 300
Gly Asp Glu Val Arg Gln Ile Ala Pro Gly Gln Thr Gly Lys Ile Ala
305 310 315 320
Asp Tyr Asn Tyr Lys Leu Pro Asp Asp Phe Thr Gly Cys Val Ile Ala
325 330 335
Trp Asn Ser Asn Asn Leu Asp Ser Lys Val Gly Gly Asn Tyr Asn Tyr
340 345 350
Arg Tyr Arg Leu Phe Arg Lys Ser Asn Leu Lys Pro Phe Glu Arg Asp
355 360 365
Ile Ser Thr Glu Ile Tyr Gln Ala Gly Ser Lys Pro Cys Asn Gly Val
370 375 380
Glu Gly Phe Asn Cys Tyr Phe Pro Leu Gln Ser Tyr Gly Phe Gln Pro
385 390 395 400
Thr Asn Gly Val Gly Tyr Gln Pro Tyr Arg Val Val Val Leu Ser Phe
405 410 415
Glu Leu Leu His Ala Pro Ala Thr Val Cys Gly Pro Lys Lys Ser Thr
420 425 430
Asn Leu Val Lys Asn Lys Arg Val Gln Pro Thr Glu Ser Ile Val Arg
435 440 445
Phe Pro Asn Ile Thr Asn Leu Cys Pro Phe Asp Glu Val Phe Asn Ala
450 455 460
Thr Arg Phe Ala Ser Val Tyr Ala Trp Asn Arg Lys Arg Ile Ser Asn
465 470 475 480
Cys Val Ala Asp Tyr Ser Val Leu Tyr Asn Leu Ala Pro Phe Phe Thr
485 490 495
Phe Lys Cys Tyr Gly Val Ser Pro Thr Lys Leu Asn Asp Leu Cys Phe
500 505 510
Thr Asn Val Tyr Ala Asp Ser Phe Val Ile Arg Gly Asp Glu Val Arg
515 520 525
Gln Ile Ala Pro Gly Gln Thr Gly Asn Ile Ala Asp Tyr Asn Tyr Lys
530 535 540
Leu Pro Asp Asp Phe Thr Gly Cys Val Ile Ala Trp Asn Ser Asn Lys
545 550 555 560
Leu Asp Ser Lys Val Ser Gly Asn Tyr Asn Tyr Leu Tyr Arg Leu Phe
565 570 575
Arg Lys Ser Asn Leu Lys Pro Phe Glu Arg Asp Ile Ser Thr Glu Ile
580 585 590
Tyr Gln Ala Gly Asn Lys Pro Cys Asn Gly Val Ala Gly Phe Asn Cys
595 600 605
Tyr Phe Pro Leu Arg Ser Tyr Ser Phe Arg Pro Thr Tyr Gly Val Gly
610 615 620
His Gln Pro Tyr Arg Val Val Val Leu Ser Phe Glu Leu Leu His Ala
625 630 635 640
Pro Ala Thr Val Cys Gly Pro Lys Lys Ser Thr Asn Leu Val Lys Asn
645 650 655
Lys
<210> 8
<211> 657
<212> PRT
<213> Artificial
<400> 8
Arg Val Gln Pro Thr Glu Ser Ile Val Arg Phe Pro Asn Ile Thr Asn
1 5 10 15
Leu Cys Pro Phe Gly Glu Val Phe Asn Ala Thr Arg Phe Ala Ser Val
20 25 30
Tyr Ala Trp Asn Arg Lys Arg Ile Ser Asn Cys Val Ala Asp Tyr Ser
35 40 45
Val Leu Tyr Asn Ser Ala Ser Phe Ser Thr Phe Lys Cys Tyr Gly Val
50 55 60
Ser Pro Thr Lys Leu Asn Asp Leu Cys Phe Thr Asn Val Tyr Ala Asp
65 70 75 80
Ser Phe Val Ile Arg Gly Asp Glu Val Arg Gln Ile Ala Pro Gly Gln
85 90 95
Thr Gly Asn Ile Ala Asp Tyr Asn Tyr Lys Leu Pro Asp Asp Phe Thr
100 105 110
Gly Cys Val Ile Ala Trp Asn Ser Asn Asn Leu Asp Ser Lys Val Gly
115 120 125
Gly Asn Tyr Asn Tyr Leu Tyr Arg Leu Phe Arg Lys Ser Asn Leu Lys
130 135 140
Pro Phe Glu Arg Asp Ile Ser Thr Glu Ile Tyr Gln Ala Gly Ser Thr
145 150 155 160
Pro Cys Asn Gly Val Lys Gly Phe Asn Cys Tyr Phe Pro Leu Gln Ser
165 170 175
Tyr Gly Phe Gln Pro Thr Tyr Gly Val Gly Tyr Gln Pro Tyr Arg Val
180 185 190
Val Val Leu Ser Phe Glu Leu Leu His Ala Pro Ala Thr Val Cys Gly
195 200 205
Pro Lys Lys Ser Thr Asn Leu Val Lys Asn Lys Arg Val Gln Pro Thr
210 215 220
Glu Ser Ile Val Arg Phe Pro Asn Ile Thr Asn Leu Cys Pro Phe Gly
225 230 235 240
Glu Val Phe Asn Ala Thr Arg Phe Ala Ser Val Tyr Ala Trp Asn Arg
245 250 255
Lys Arg Ile Ser Asn Cys Val Ala Asp Tyr Ser Val Leu Tyr Asn Ser
260 265 270
Ala Ser Phe Ser Thr Phe Lys Cys Tyr Gly Val Ser Pro Thr Lys Leu
275 280 285
Asn Asp Leu Cys Phe Thr Asn Val Tyr Ala Asp Ser Phe Val Ile Arg
290 295 300
Gly Asp Glu Val Arg Gln Ile Ala Pro Gly Gln Thr Gly Asn Ile Ala
305 310 315 320
Asp Tyr Asn Tyr Lys Leu Pro Asp Asp Phe Thr Gly Cys Val Ile Ala
325 330 335
Trp Asn Ser Asn Asn Leu Asp Ser Lys Val Gly Gly Asn Tyr Asn Tyr
340 345 350
Leu Tyr Arg Leu Phe Arg Lys Ser Asn Leu Lys Pro Phe Glu Arg Asp
355 360 365
Ile Ser Thr Glu Ile Tyr Gln Ala Gly Ser Thr Pro Cys Asn Gly Val
370 375 380
Lys Gly Phe Asn Cys Tyr Phe Pro Leu Gln Ser Tyr Gly Phe Gln Pro
385 390 395 400
Thr Tyr Gly Val Gly Tyr Gln Pro Tyr Arg Val Val Val Leu Ser Phe
405 410 415
Glu Leu Leu His Ala Pro Ala Thr Val Cys Gly Pro Lys Lys Ser Thr
420 425 430
Asn Leu Val Lys Asn Lys Arg Val Gln Pro Thr Glu Ser Ile Val Arg
435 440 445
Phe Pro Asn Ile Thr Asn Leu Cys Pro Phe Asp Glu Val Phe Asn Ala
450 455 460
Thr Arg Phe Ala Ser Val Tyr Ala Trp Asn Arg Lys Arg Ile Ser Asn
465 470 475 480
Cys Val Ala Asp Tyr Ser Val Leu Tyr Asn Leu Ala Pro Phe Phe Thr
485 490 495
Phe Lys Cys Tyr Gly Val Ser Pro Thr Lys Leu Asn Asp Leu Cys Phe
500 505 510
Thr Asn Val Tyr Ala Asp Ser Phe Val Ile Arg Gly Asp Glu Val Arg
515 520 525
Gln Ile Ala Pro Gly Gln Thr Gly Asn Ile Ala Asp Tyr Asn Tyr Lys
530 535 540
Leu Pro Asp Asp Phe Thr Gly Cys Val Ile Ala Trp Asn Ser Asn Lys
545 550 555 560
Leu Asp Ser Lys Val Ser Gly Asn Tyr Asn Tyr Leu Tyr Arg Leu Phe
565 570 575
Arg Lys Ser Asn Leu Lys Pro Phe Glu Arg Asp Ile Ser Thr Glu Ile
580 585 590
Tyr Gln Ala Gly Asn Lys Pro Cys Asn Gly Val Ala Gly Phe Asn Cys
595 600 605
Tyr Phe Pro Leu Arg Ser Tyr Ser Phe Arg Pro Thr Tyr Gly Val Gly
610 615 620
His Gln Pro Tyr Arg Val Val Val Leu Ser Phe Glu Leu Leu His Ala
625 630 635 640
Pro Ala Thr Val Cys Gly Pro Lys Lys Ser Thr Asn Leu Val Lys Asn
645 650 655
Lys
<210> 9
<211> 657
<212> PRT
<213> Artificial
<400> 9
Arg Val Gln Pro Thr Glu Ser Ile Val Arg Phe Pro Asn Ile Thr Asn
1 5 10 15
Leu Cys Pro Phe Gly Glu Val Phe Asn Ala Thr Arg Phe Ala Ser Val
20 25 30
Tyr Ala Trp Asn Arg Lys Arg Ile Ser Asn Cys Val Ala Asp Tyr Ser
35 40 45
Val Leu Tyr Asn Ser Ala Ser Phe Ser Thr Phe Lys Cys Tyr Gly Val
50 55 60
Ser Pro Thr Lys Leu Asn Asp Leu Cys Phe Thr Asn Val Tyr Ala Asp
65 70 75 80
Ser Phe Val Ile Arg Gly Asp Glu Val Arg Gln Ile Ala Pro Gly Gln
85 90 95
Thr Gly Lys Ile Ala Asp Tyr Asn Tyr Lys Leu Pro Asp Asp Phe Thr
100 105 110
Gly Cys Val Ile Ala Trp Asn Ser Asn Asn Leu Asp Ser Lys Val Gly
115 120 125
Gly Asn Tyr Asn Tyr Arg Tyr Arg Leu Phe Arg Lys Ser Asn Leu Lys
130 135 140
Pro Phe Glu Arg Asp Ile Ser Thr Glu Ile Tyr Gln Ala Gly Ser Lys
145 150 155 160
Pro Cys Asn Gly Val Glu Gly Phe Asn Cys Tyr Phe Pro Leu Gln Ser
165 170 175
Tyr Gly Phe Gln Pro Thr Asn Gly Val Gly Tyr Gln Pro Tyr Arg Val
180 185 190
Val Val Leu Ser Phe Glu Leu Leu His Ala Pro Ala Thr Val Cys Gly
195 200 205
Pro Lys Lys Ser Thr Asn Leu Val Lys Asn Lys Arg Val Gln Pro Thr
210 215 220
Glu Ser Ile Val Arg Phe Pro Asn Ile Thr Asn Leu Cys Pro Phe Gly
225 230 235 240
Glu Val Phe Asn Ala Thr Arg Phe Ala Ser Val Tyr Ala Trp Asn Arg
245 250 255
Lys Arg Ile Ser Asn Cys Val Ala Asp Tyr Ser Val Leu Tyr Asn Ser
260 265 270
Ala Ser Phe Ser Thr Phe Lys Cys Tyr Gly Val Ser Pro Thr Lys Leu
275 280 285
Asn Asp Leu Cys Phe Thr Asn Val Tyr Ala Asp Ser Phe Val Ile Arg
290 295 300
Gly Asp Glu Val Arg Gln Ile Ala Pro Gly Gln Thr Gly Lys Ile Ala
305 310 315 320
Asp Tyr Asn Tyr Lys Leu Pro Asp Asp Phe Thr Gly Cys Val Ile Ala
325 330 335
Trp Asn Ser Asn Asn Leu Asp Ser Lys Val Gly Gly Asn Tyr Asn Tyr
340 345 350
Arg Tyr Arg Leu Phe Arg Lys Ser Asn Leu Lys Pro Phe Glu Arg Asp
355 360 365
Ile Ser Thr Glu Ile Tyr Gln Ala Gly Ser Lys Pro Cys Asn Gly Val
370 375 380
Glu Gly Phe Asn Cys Tyr Phe Pro Leu Gln Ser Tyr Gly Phe Gln Pro
385 390 395 400
Thr Asn Gly Val Gly Tyr Gln Pro Tyr Arg Val Val Val Leu Ser Phe
405 410 415
Glu Leu Leu His Ala Pro Ala Thr Val Cys Gly Pro Lys Lys Ser Thr
420 425 430
Asn Leu Val Lys Asn Lys Arg Val Gln Pro Thr Glu Ser Ile Val Arg
435 440 445
Phe Pro Asn Ile Thr Asn Leu Cys Pro Phe Gly Glu Val Phe Asn Ala
450 455 460
Thr Arg Phe Ala Ser Val Tyr Ala Trp Asn Arg Lys Arg Ile Ser Asn
465 470 475 480
Cys Val Ala Asp Tyr Ser Val Leu Tyr Asn Ser Ala Ser Phe Ser Thr
485 490 495
Phe Lys Cys Tyr Gly Val Ser Pro Thr Lys Leu Asn Asp Leu Cys Phe
500 505 510
Thr Asn Val Tyr Ala Asp Ser Phe Val Ile Arg Gly Asp Glu Val Arg
515 520 525
Gln Ile Ala Pro Gly Gln Thr Gly Lys Ile Ala Asp Tyr Asn Tyr Lys
530 535 540
Leu Pro Asp Asp Phe Thr Gly Cys Val Ile Ala Trp Asn Ser Asn Asn
545 550 555 560
Leu Asp Ser Lys Val Gly Gly Asn Tyr Asn Tyr Arg Tyr Arg Leu Phe
565 570 575
Arg Lys Ser Asn Leu Lys Pro Phe Glu Arg Asp Ile Ser Thr Glu Ile
580 585 590
Tyr Gln Ala Gly Ser Lys Pro Cys Asn Gly Val Glu Gly Phe Asn Cys
595 600 605
Tyr Phe Pro Leu Gln Ser Tyr Gly Phe Gln Pro Thr Asn Gly Val Gly
610 615 620
Tyr Gln Pro Tyr Arg Val Val Val Leu Ser Phe Glu Leu Leu His Ala
625 630 635 640
Pro Ala Thr Val Cys Gly Pro Lys Lys Ser Thr Asn Leu Val Lys Asn
645 650 655
Lys
<210> 10
<211> 657
<212> PRT
<213> Artificial
<400> 10
Arg Val Gln Pro Thr Glu Ser Ile Val Arg Phe Pro Asn Ile Thr Asn
1 5 10 15
Leu Cys Pro Phe Gly Glu Val Phe Asn Ala Thr Arg Phe Ala Ser Val
20 25 30
Tyr Ala Trp Asn Arg Lys Arg Ile Ser Asn Cys Val Ala Asp Tyr Ser
35 40 45
Val Leu Tyr Asn Ser Ala Ser Phe Ser Thr Phe Lys Cys Tyr Gly Val
50 55 60
Ser Pro Thr Lys Leu Asn Asp Leu Cys Phe Thr Asn Val Tyr Ala Asp
65 70 75 80
Ser Phe Val Ile Arg Gly Asp Glu Val Arg Gln Ile Ala Pro Gly Gln
85 90 95
Thr Gly Asn Ile Ala Asp Tyr Asn Tyr Lys Leu Pro Asp Asp Phe Thr
100 105 110
Gly Cys Val Ile Ala Trp Asn Ser Asn Asn Leu Asp Ser Lys Val Gly
115 120 125
Gly Asn Tyr Asn Tyr Leu Tyr Arg Leu Phe Arg Lys Ser Asn Leu Lys
130 135 140
Pro Phe Glu Arg Asp Ile Ser Thr Glu Ile Tyr Gln Ala Gly Ser Thr
145 150 155 160
Pro Cys Asn Gly Val Lys Gly Phe Asn Cys Tyr Phe Pro Leu Gln Ser
165 170 175
Tyr Gly Phe Gln Pro Thr Tyr Gly Val Gly Tyr Gln Pro Tyr Arg Val
180 185 190
Val Val Leu Ser Phe Glu Leu Leu His Ala Pro Ala Thr Val Cys Gly
195 200 205
Pro Lys Lys Ser Thr Asn Leu Val Lys Asn Lys Arg Val Gln Pro Thr
210 215 220
Glu Ser Ile Val Arg Phe Pro Asn Ile Thr Asn Leu Cys Pro Phe Gly
225 230 235 240
Glu Val Phe Asn Ala Thr Arg Phe Ala Ser Val Tyr Ala Trp Asn Arg
245 250 255
Lys Arg Ile Ser Asn Cys Val Ala Asp Tyr Ser Val Leu Tyr Asn Ser
260 265 270
Ala Ser Phe Ser Thr Phe Lys Cys Tyr Gly Val Ser Pro Thr Lys Leu
275 280 285
Asn Asp Leu Cys Phe Thr Asn Val Tyr Ala Asp Ser Phe Val Ile Arg
290 295 300
Gly Asp Glu Val Arg Gln Ile Ala Pro Gly Gln Thr Gly Asn Ile Ala
305 310 315 320
Asp Tyr Asn Tyr Lys Leu Pro Asp Asp Phe Thr Gly Cys Val Ile Ala
325 330 335
Trp Asn Ser Asn Asn Leu Asp Ser Lys Val Gly Gly Asn Tyr Asn Tyr
340 345 350
Leu Tyr Arg Leu Phe Arg Lys Ser Asn Leu Lys Pro Phe Glu Arg Asp
355 360 365
Ile Ser Thr Glu Ile Tyr Gln Ala Gly Ser Thr Pro Cys Asn Gly Val
370 375 380
Lys Gly Phe Asn Cys Tyr Phe Pro Leu Gln Ser Tyr Gly Phe Gln Pro
385 390 395 400
Thr Tyr Gly Val Gly Tyr Gln Pro Tyr Arg Val Val Val Leu Ser Phe
405 410 415
Glu Leu Leu His Ala Pro Ala Thr Val Cys Gly Pro Lys Lys Ser Thr
420 425 430
Asn Leu Val Lys Asn Lys Arg Val Gln Pro Thr Glu Ser Ile Val Arg
435 440 445
Phe Pro Asn Ile Thr Asn Leu Cys Pro Phe Gly Glu Val Phe Asn Ala
450 455 460
Thr Arg Phe Ala Ser Val Tyr Ala Trp Asn Arg Lys Arg Ile Ser Asn
465 470 475 480
Cys Val Ala Asp Tyr Ser Val Leu Tyr Asn Ser Ala Ser Phe Ser Thr
485 490 495
Phe Lys Cys Tyr Gly Val Ser Pro Thr Lys Leu Asn Asp Leu Cys Phe
500 505 510
Thr Asn Val Tyr Ala Asp Ser Phe Val Ile Arg Gly Asp Glu Val Arg
515 520 525
Gln Ile Ala Pro Gly Gln Thr Gly Asn Ile Ala Asp Tyr Asn Tyr Lys
530 535 540
Leu Pro Asp Asp Phe Thr Gly Cys Val Ile Ala Trp Asn Ser Asn Asn
545 550 555 560
Leu Asp Ser Lys Val Gly Gly Asn Tyr Asn Tyr Leu Tyr Arg Leu Phe
565 570 575
Arg Lys Ser Asn Leu Lys Pro Phe Glu Arg Asp Ile Ser Thr Glu Ile
580 585 590
Tyr Gln Ala Gly Ser Thr Pro Cys Asn Gly Val Lys Gly Phe Asn Cys
595 600 605
Tyr Phe Pro Leu Gln Ser Tyr Gly Phe Gln Pro Thr Tyr Gly Val Gly
610 615 620
Tyr Gln Pro Tyr Arg Val Val Val Leu Ser Phe Glu Leu Leu His Ala
625 630 635 640
Pro Ala Thr Val Cys Gly Pro Lys Lys Ser Thr Asn Leu Val Lys Asn
645 650 655
Lys
<210> 11
<211> 657
<212> PRT
<213> Artificial
<400> 11
Arg Val Gln Pro Thr Glu Ser Ile Val Arg Phe Pro Asn Ile Thr Asn
1 5 10 15
Leu Cys Pro Phe Asp Glu Val Phe Asn Ala Thr Arg Phe Ala Ser Val
20 25 30
Tyr Ala Trp Asn Arg Lys Arg Ile Ser Asn Cys Val Ala Asp Tyr Ser
35 40 45
Val Leu Tyr Asn Leu Ala Pro Phe Phe Thr Phe Lys Cys Tyr Gly Val
50 55 60
Ser Pro Thr Lys Leu Asn Asp Leu Cys Phe Thr Asn Val Tyr Ala Asp
65 70 75 80
Ser Phe Val Ile Arg Gly Asp Glu Val Arg Gln Ile Ala Pro Gly Gln
85 90 95
Thr Gly Asn Ile Ala Asp Tyr Asn Tyr Lys Leu Pro Asp Asp Phe Thr
100 105 110
Gly Cys Val Ile Ala Trp Asn Ser Asn Lys Leu Asp Ser Lys Val Ser
115 120 125
Gly Asn Tyr Asn Tyr Leu Tyr Arg Leu Phe Arg Lys Ser Asn Leu Lys
130 135 140
Pro Phe Glu Arg Asp Ile Ser Thr Glu Ile Tyr Gln Ala Gly Asn Lys
145 150 155 160
Pro Cys Asn Gly Val Ala Gly Phe Asn Cys Tyr Phe Pro Leu Arg Ser
165 170 175
Tyr Ser Phe Arg Pro Thr Tyr Gly Val Gly His Gln Pro Tyr Arg Val
180 185 190
Val Val Leu Ser Phe Glu Leu Leu His Ala Pro Ala Thr Val Cys Gly
195 200 205
Pro Lys Lys Ser Thr Asn Leu Val Lys Asn Lys Arg Val Gln Pro Thr
210 215 220
Glu Ser Ile Val Arg Phe Pro Asn Ile Thr Asn Leu Cys Pro Phe Asp
225 230 235 240
Glu Val Phe Asn Ala Thr Arg Phe Ala Ser Val Tyr Ala Trp Asn Arg
245 250 255
Lys Arg Ile Ser Asn Cys Val Ala Asp Tyr Ser Val Leu Tyr Asn Leu
260 265 270
Ala Pro Phe Phe Thr Phe Lys Cys Tyr Gly Val Ser Pro Thr Lys Leu
275 280 285
Asn Asp Leu Cys Phe Thr Asn Val Tyr Ala Asp Ser Phe Val Ile Arg
290 295 300
Gly Asp Glu Val Arg Gln Ile Ala Pro Gly Gln Thr Gly Asn Ile Ala
305 310 315 320
Asp Tyr Asn Tyr Lys Leu Pro Asp Asp Phe Thr Gly Cys Val Ile Ala
325 330 335
Trp Asn Ser Asn Lys Leu Asp Ser Lys Val Ser Gly Asn Tyr Asn Tyr
340 345 350
Leu Tyr Arg Leu Phe Arg Lys Ser Asn Leu Lys Pro Phe Glu Arg Asp
355 360 365
Ile Ser Thr Glu Ile Tyr Gln Ala Gly Asn Lys Pro Cys Asn Gly Val
370 375 380
Ala Gly Phe Asn Cys Tyr Phe Pro Leu Arg Ser Tyr Ser Phe Arg Pro
385 390 395 400
Thr Tyr Gly Val Gly His Gln Pro Tyr Arg Val Val Val Leu Ser Phe
405 410 415
Glu Leu Leu His Ala Pro Ala Thr Val Cys Gly Pro Lys Lys Ser Thr
420 425 430
Asn Leu Val Lys Asn Lys Arg Val Gln Pro Thr Glu Ser Ile Val Arg
435 440 445
Phe Pro Asn Ile Thr Asn Leu Cys Pro Phe Asp Glu Val Phe Asn Ala
450 455 460
Thr Arg Phe Ala Ser Val Tyr Ala Trp Asn Arg Lys Arg Ile Ser Asn
465 470 475 480
Cys Val Ala Asp Tyr Ser Val Leu Tyr Asn Leu Ala Pro Phe Phe Thr
485 490 495
Phe Lys Cys Tyr Gly Val Ser Pro Thr Lys Leu Asn Asp Leu Cys Phe
500 505 510
Thr Asn Val Tyr Ala Asp Ser Phe Val Ile Arg Gly Asp Glu Val Arg
515 520 525
Gln Ile Ala Pro Gly Gln Thr Gly Asn Ile Ala Asp Tyr Asn Tyr Lys
530 535 540
Leu Pro Asp Asp Phe Thr Gly Cys Val Ile Ala Trp Asn Ser Asn Lys
545 550 555 560
Leu Asp Ser Lys Val Ser Gly Asn Tyr Asn Tyr Leu Tyr Arg Leu Phe
565 570 575
Arg Lys Ser Asn Leu Lys Pro Phe Glu Arg Asp Ile Ser Thr Glu Ile
580 585 590
Tyr Gln Ala Gly Asn Lys Pro Cys Asn Gly Val Ala Gly Phe Asn Cys
595 600 605
Tyr Phe Pro Leu Arg Ser Tyr Ser Phe Arg Pro Thr Tyr Gly Val Gly
610 615 620
His Gln Pro Tyr Arg Val Val Val Leu Ser Phe Glu Leu Leu His Ala
625 630 635 640
Pro Ala Thr Val Cys Gly Pro Lys Lys Ser Thr Asn Leu Val Lys Asn
645 650 655
Lys
<210> 12
<211> 1973
<212> DNA
<213> Artificial
<400> 12
atgagggtgc agcccaccga gagcatcgtg aggttcccca acatcaccaa cctgtgcccc 60
ttcggcgagg tgttcaacgc caccaggttc gccagcgtgt acgcctggaa caggaagagg 120
atctcaaact gcgtggcaga ctacagtgtg ctgtacaact ccgcctcctt ctccacattc 180
aagtgctatg gcgtgtcccc caccaagctg aacgatctgt gctttaccaa cgtgtacgcc 240
gactccttcg tgattagggg cgacgaggtg cggcagattg cccccggtca gaccggcaag 300
atcgccgatt ataactacaa actgcccgac gacttcaccg gctgcgtgat cgcctggaac 360
tccaacaatc tggactctaa ggtggggggc aactacaact accggtacag actgttccgc 420
aaaagcaacc tgaagccctt cgaaagggac atctccacag agatctacca ggccggctcc 480
aaaccctgca acggcgtgga gggcttcaac tgctacttcc ccctgcagtc ctacggcttc 540
cagcccacca acggcgtggg ataccagccc tacagagtgg tggtgctgtc cttcgagctg 600
ctgcacgcac ctgccacagt gtgcggcccc aagaagtcta ccaacctggt gaaaaacaag 660
agagtgcagc ccacagaatc catcgtgagg tttcccaaca ttaccaatct gtgccccttt 720
ggcgaagtgt tcaacgctac caggttcgca tccgtgtacg cctggaatag aaaaaggatc 780
tccaactgcg tggccgacta ctccgtgctg tacaattccg cctccttttc caccttcaaa 840
tgctatggcg tgagccccac caagctgaat gatctgtgtt ttaccaacgt gtatgccgac 900
tcctttgtga ttaggggaga cgaggtgagg cagatcgccc ctggccagac cggcaacatc 960
gccgactata actacaagct gcccgacgat ttcaccggct gtgtgattgc ttggaactcc 1020
aataatctgg atagcaaggt ggggggaaac tacaattacc tgtacaggct gttcaggaag 1080
agtaacctga agccattcga aagggatatc tccactgaga tctaccaggc tggctccacc 1140
ccctgcaacg gagtgaaggg attcaactgc tatttccccc tgcagagcta cggcttccag 1200
ccaacctacg gcgtgggcta ccagccctat agggtggtgg tgctgtcttt cgagctgctg 1260
catgcccccg ccacagtgtg tggccccaag aaatccacca acctggtgaa gaacaaaagg 1320
gtgcagccta ccgagtccat cgtgcggttc cccaatatca caaacctgtg tcccttcgac 1380
gaggtgttta acgccacccg gttcgcctcc gtgtacgctt ggaatagaaa gagaatctct 1440
aactgcgtgg ctgactactc cgtgctgtat aacctggccc ccttctttac cttcaagtgt 1500
tatggcgtgt ctcccactaa gctgaacgac ctgtgcttca ccaacgtgta cgctgactcc 1560
ttcgtgatca ggggcgacga agtgcggcag atcgcccccg gtcagacagg caacatcgct 1620
gattacaact acaaactgcc tgacgacttc actggctgcg tgattgcctg gaacagcaac 1680
aagctggaca gcaaggtgtc cggcaactac aattatctgt acaggctgtt taggaagagt 1740
aatctgaaac ccttcgaacg ggacatctcc actgaaatct accaggcagg caacaagccc 1800
tgcaacgggg tggctggctt taactgctac tttcccctga gatcctactc cttcagaccc 1860
acctacggcg tgggacacca gccctacagg gtggtggtgc tgagcttcga gctgctgcac 1920
gcccccgcaa ccgtgtgcgg acctaagaag tccaccaacc tggtgaagaa taa 1973
<210> 13
<211> 1971
<212> DNA
<213> Artificial
<400> 13
agggtgcagc ccaccgagag catcgtgagg ttccccaaca tcaccaacct gtgccccttc 60
ggcgaggtgt tcaacgccac caggttcgcc agcgtgtacg cctggaacag gaagaggatc 120
tcaaactgcg tggcagacta cagtgtgctg tacaactccg cctccttctc cacattcaag 180
tgctatggcg tgtcccccac caagctgaac gatctgtgct ttaccaacgt gtacgccgac 240
tccttcgtga ttaggggcga cgaggtgcgg cagattgccc ccggtcagac cggcaagatc 300
gccgattata actacaaact gcccgacgac ttcaccggct gcgtgatcgc ctggaactcc 360
aacaatctgg actctaaggt ggggggcaac tacaactacc ggtacagact gttccgcaaa 420
agcaacctga agcccttcga aagggacatc tccacagaga tctaccaggc cggctccaaa 480
ccctgcaacg gcgtggaggg cttcaactgc tacttccccc tgcagtccta cggcttccag 540
cccaccaacg gcgtgggata ccagccctac agagtggtgg tgctgtcctt cgagctgctg 600
cacgcacctg ccacagtgtg cggccccaag aagtctacca acctggtgaa aaacaagagg 660
gtgcagccta ccgagtccat cgtgcggttc cccaatatca caaacctgtg tcccttcgac 720
gaggtgttta acgccacccg gttcgcctcc gtgtacgctt ggaatagaaa gagaatctct 780
aactgcgtgg ctgactactc cgtgctgtat aacctggccc ccttctttac cttcaagtgt 840
tatggcgtgt ctcccactaa gctgaacgac ctgtgcttca ccaacgtgta cgctgactcc 900
ttcgtgatca ggggcgacga agtgcggcag atcgcccccg gtcagacagg caacatcgct 960
gattacaact acaaactgcc tgacgacttc actggctgcg tgattgcctg gaacagcaac 1020
aagctggaca gcaaggtgtc cggcaactac aattatctgt acaggctgtt taggaagagt 1080
aatctgaaac ccttcgaacg ggacatctcc actgaaatct accaggcagg caacaagccc 1140
tgcaacgggg tggctggctt taactgctac tttcccctga gatcctactc cttcagaccc 1200
acctacggcg tgggacacca gccctacagg gtggtggtgc tgagcttcga gctgctgcac 1260
gcccccgcaa ccgtgtgcgg acctaagaag tccaccaacc tggtgaagaa taagagagtg 1320
cagcccacag aatccatcgt gaggtttccc aacattacca atctgtgccc ctttggcgaa 1380
gtgttcaacg ctaccaggtt cgcatccgtg tacgcctgga atagaaaaag gatctccaac 1440
tgcgtggccg actactccgt gctgtacaat tccgcctcct tttccacctt caaatgctat 1500
ggcgtgagcc ccaccaagct gaatgatctg tgttttacca acgtgtatgc cgactccttt 1560
gtgattaggg gagacgaggt gaggcagatc gcccctggcc agaccggcaa catcgccgac 1620
tataactaca agctgcccga cgatttcacc ggctgtgtga ttgcttggaa ctccaataat 1680
ctggatagca aggtgggggg aaactacaat tacctgtaca ggctgttcag gaagagtaac 1740
ctgaagccat tcgaaaggga tatctccact gagatctacc aggctggctc caccccctgc 1800
aacggagtga agggattcaa ctgctatttc cccctgcaga gctacggctt ccagccaacc 1860
tacggcgtgg gctaccagcc ctatagggtg gtggtgctgt ctttcgagct gctgcatgcc 1920
cccgccacag tgtgtggccc caagaaatcc accaacctgg tgaagaacaa a 1971
<210> 14
<211> 1971
<212> DNA
<213> Artificial
<400> 14
agagtgcagc ccacagaatc catcgtgagg tttcccaaca ttaccaatct gtgccccttt 60
ggcgaagtgt tcaacgctac caggttcgca tccgtgtacg cctggaatag aaaaaggatc 120
tccaactgcg tggccgacta ctccgtgctg tacaattccg cctccttttc caccttcaaa 180
tgctatggcg tgagccccac caagctgaat gatctgtgtt ttaccaacgt gtatgccgac 240
tcctttgtga ttaggggaga cgaggtgagg cagatcgccc ctggccagac cggcaacatc 300
gccgactata actacaagct gcccgacgat ttcaccggct gtgtgattgc ttggaactcc 360
aataatctgg atagcaaggt ggggggaaac tacaattacc tgtacaggct gttcaggaag 420
agtaacctga agccattcga aagggatatc tccactgaga tctaccaggc tggctccacc 480
ccctgcaacg gagtgaaggg attcaactgc tatttccccc tgcagagcta cggcttccag 540
ccaacctacg gcgtgggcta ccagccctat agggtggtgg tgctgtcttt cgagctgctg 600
catgcccccg ccacagtgtg tggccccaag aaatccacca acctggtgaa gaacaaaagg 660
gtgcagccca ccgagagcat cgtgaggttc cccaacatca ccaacctgtg ccccttcggc 720
gaggtgttca acgccaccag gttcgccagc gtgtacgcct ggaacaggaa gaggatctca 780
aactgcgtgg cagactacag tgtgctgtac aactccgcct ccttctccac attcaagtgc 840
tatggcgtgt cccccaccaa gctgaacgat ctgtgcttta ccaacgtgta cgccgactcc 900
ttcgtgatta ggggcgacga ggtgcggcag attgcccccg gtcagaccgg caagatcgcc 960
gattataact acaaactgcc cgacgacttc accggctgcg tgatcgcctg gaactccaac 1020
aatctggact ctaaggtggg gggcaactac aactaccggt acagactgtt ccgcaaaagc 1080
aacctgaagc ccttcgaaag ggacatctcc acagagatct accaggccgg ctccaaaccc 1140
tgcaacggcg tggagggctt caactgctac ttccccctgc agtcctacgg cttccagccc 1200
accaacggcg tgggatacca gccctacaga gtggtggtgc tgtccttcga gctgctgcac 1260
gcacctgcca cagtgtgcgg ccccaagaag tctaccaacc tggtgaaaaa caagagggtg 1320
cagcctaccg agtccatcgt gcggttcccc aatatcacaa acctgtgtcc cttcgacgag 1380
gtgtttaacg ccacccggtt cgcctccgtg tacgcttgga atagaaagag aatctctaac 1440
tgcgtggctg actactccgt gctgtataac ctggccccct tctttacctt caagtgttat 1500
ggcgtgtctc ccactaagct gaacgacctg tgcttcacca acgtgtacgc tgactccttc 1560
gtgatcaggg gcgacgaagt gcggcagatc gcccccggtc agacaggcaa catcgctgat 1620
tacaactaca aactgcctga cgacttcact ggctgcgtga ttgcctggaa cagcaacaag 1680
ctggacagca aggtgtccgg caactacaat tatctgtaca ggctgtttag gaagagtaat 1740
ctgaaaccct tcgaacggga catctccact gaaatctacc aggcaggcaa caagccctgc 1800
aacggggtgg ctggctttaa ctgctacttt cccctgagat cctactcctt cagacccacc 1860
tacggcgtgg gacaccagcc ctacagggtg gtggtgctga gcttcgagct gctgcacgcc 1920
cccgcaaccg tgtgcggacc taagaagtcc accaacctgg tgaagaataa g 1971
<210> 15
<211> 1971
<212> DNA
<213> Artificial
<400> 15
agagtgcagc ccacagaatc catcgtgagg tttcccaaca ttaccaatct gtgccccttt 60
ggcgaagtgt tcaacgctac caggttcgca tccgtgtacg cctggaatag aaaaaggatc 120
tccaactgcg tggccgacta ctccgtgctg tacaattccg cctccttttc caccttcaaa 180
tgctatggcg tgagccccac caagctgaat gatctgtgtt ttaccaacgt gtatgccgac 240
tcctttgtga ttaggggaga cgaggtgagg cagatcgccc ctggccagac cggcaacatc 300
gccgactata actacaagct gcccgacgat ttcaccggct gtgtgattgc ttggaactcc 360
aataatctgg atagcaaggt ggggggaaac tacaattacc tgtacaggct gttcaggaag 420
agtaacctga agccattcga aagggatatc tccactgaga tctaccaggc tggctccacc 480
ccctgcaacg gagtgaaggg attcaactgc tatttccccc tgcagagcta cggcttccag 540
ccaacctacg gcgtgggcta ccagccctat agggtggtgg tgctgtcttt cgagctgctg 600
catgcccccg ccacagtgtg tggccccaag aaatccacca acctggtgaa gaacaaaagg 660
gtgcagccta ccgagtccat cgtgcggttc cccaatatca caaacctgtg tcccttcgac 720
gaggtgttta acgccacccg gttcgcctcc gtgtacgctt ggaatagaaa gagaatctct 780
aactgcgtgg ctgactactc cgtgctgtat aacctggccc ccttctttac cttcaagtgt 840
tatggcgtgt ctcccactaa gctgaacgac ctgtgcttca ccaacgtgta cgctgactcc 900
ttcgtgatca ggggcgacga agtgcggcag atcgcccccg gtcagacagg caacatcgct 960
gattacaact acaaactgcc tgacgacttc actggctgcg tgattgcctg gaacagcaac 1020
aagctggaca gcaaggtgtc cggcaactac aattatctgt acaggctgtt taggaagagt 1080
aatctgaaac ccttcgaacg ggacatctcc actgaaatct accaggcagg caacaagccc 1140
tgcaacgggg tggctggctt taactgctac tttcccctga gatcctactc cttcagaccc 1200
acctacggcg tgggacacca gccctacagg gtggtggtgc tgagcttcga gctgctgcac 1260
gcccccgcaa ccgtgtgcgg acctaagaag tccaccaacc tggtgaagaa taagagggtg 1320
cagcccaccg agagcatcgt gaggttcccc aacatcacca acctgtgccc cttcggcgag 1380
gtgttcaacg ccaccaggtt cgccagcgtg tacgcctgga acaggaagag gatctcaaac 1440
tgcgtggcag actacagtgt gctgtacaac tccgcctcct tctccacatt caagtgctat 1500
ggcgtgtccc ccaccaagct gaacgatctg tgctttacca acgtgtacgc cgactccttc 1560
gtgattaggg gcgacgaggt gcggcagatt gcccccggtc agaccggcaa gatcgccgat 1620
tataactaca aactgcccga cgacttcacc ggctgcgtga tcgcctggaa ctccaacaat 1680
ctggactcta aggtgggggg caactacaac taccggtaca gactgttccg caaaagcaac 1740
ctgaagccct tcgaaaggga catctccaca gagatctacc aggccggctc caaaccctgc 1800
aacggcgtgg agggcttcaa ctgctacttc cccctgcagt cctacggctt ccagcccacc 1860
aacggcgtgg gataccagcc ctacagagtg gtggtgctgt ccttcgagct gctgcacgca 1920
cctgccacag tgtgcggccc caagaagtct accaacctgg tgaaaaacaa g 1971
<210> 16
<211> 1971
<212> DNA
<213> Artificial
<400> 16
agggtgcagc ctaccgagtc catcgtgcgg ttccccaata tcacaaacct gtgtcccttc 60
gacgaggtgt ttaacgccac ccggttcgcc tccgtgtacg cttggaatag aaagagaatc 120
tctaactgcg tggctgacta ctccgtgctg tataacctgg cccccttctt taccttcaag 180
tgttatggcg tgtctcccac taagctgaac gacctgtgct tcaccaacgt gtacgctgac 240
tccttcgtga tcaggggcga cgaagtgcgg cagatcgccc ccggtcagac aggcaacatc 300
gctgattaca actacaaact gcctgacgac ttcactggct gcgtgattgc ctggaacagc 360
aacaagctgg acagcaaggt gtccggcaac tacaattatc tgtacaggct gtttaggaag 420
agtaatctga aacccttcga acgggacatc tccactgaaa tctaccaggc aggcaacaag 480
ccctgcaacg gggtggctgg ctttaactgc tactttcccc tgagatccta ctccttcaga 540
cccacctacg gcgtgggaca ccagccctac agggtggtgg tgctgagctt cgagctgctg 600
cacgcccccg caaccgtgtg cggacctaag aagtccacca acctggtgaa gaataagagg 660
gtgcagccca ccgagagcat cgtgaggttc cccaacatca ccaacctgtg ccccttcggc 720
gaggtgttca acgccaccag gttcgccagc gtgtacgcct ggaacaggaa gaggatctca 780
aactgcgtgg cagactacag tgtgctgtac aactccgcct ccttctccac attcaagtgc 840
tatggcgtgt cccccaccaa gctgaacgat ctgtgcttta ccaacgtgta cgccgactcc 900
ttcgtgatta ggggcgacga ggtgcggcag attgcccccg gtcagaccgg caagatcgcc 960
gattataact acaaactgcc cgacgacttc accggctgcg tgatcgcctg gaactccaac 1020
aatctggact ctaaggtggg gggcaactac aactaccggt acagactgtt ccgcaaaagc 1080
aacctgaagc ccttcgaaag ggacatctcc acagagatct accaggccgg ctccaaaccc 1140
tgcaacggcg tggagggctt caactgctac ttccccctgc agtcctacgg cttccagccc 1200
accaacggcg tgggatacca gccctacaga gtggtggtgc tgtccttcga gctgctgcac 1260
gcacctgcca cagtgtgcgg ccccaagaag tctaccaacc tggtgaaaaa caagagagtg 1320
cagcccacag aatccatcgt gaggtttccc aacattacca atctgtgccc ctttggcgaa 1380
gtgttcaacg ctaccaggtt cgcatccgtg tacgcctgga atagaaaaag gatctccaac 1440
tgcgtggccg actactccgt gctgtacaat tccgcctcct tttccacctt caaatgctat 1500
ggcgtgagcc ccaccaagct gaatgatctg tgttttacca acgtgtatgc cgactccttt 1560
gtgattaggg gagacgaggt gaggcagatc gcccctggcc agaccggcaa catcgccgac 1620
tataactaca agctgcccga cgatttcacc ggctgtgtga ttgcttggaa ctccaataat 1680
ctggatagca aggtgggggg aaactacaat tacctgtaca ggctgttcag gaagagtaac 1740
ctgaagccat tcgaaaggga tatctccact gagatctacc aggctggctc caccccctgc 1800
aacggagtga agggattcaa ctgctatttc cccctgcaga gctacggctt ccagccaacc 1860
tacggcgtgg gctaccagcc ctatagggtg gtggtgctgt ctttcgagct gctgcatgcc 1920
cccgccacag tgtgtggccc caagaaatcc accaacctgg tgaagaacaa a 1971
<210> 17
<211> 1971
<212> DNA
<213> Artificial
<400> 17
agggtgcagc ctaccgagtc catcgtgcgg ttccccaata tcacaaacct gtgtcccttc 60
gacgaggtgt ttaacgccac ccggttcgcc tccgtgtacg cttggaatag aaagagaatc 120
tctaactgcg tggctgacta ctccgtgctg tataacctgg cccccttctt taccttcaag 180
tgttatggcg tgtctcccac taagctgaac gacctgtgct tcaccaacgt gtacgctgac 240
tccttcgtga tcaggggcga cgaagtgcgg cagatcgccc ccggtcagac aggcaacatc 300
gctgattaca actacaaact gcctgacgac ttcactggct gcgtgattgc ctggaacagc 360
aacaagctgg acagcaaggt gtccggcaac tacaattatc tgtacaggct gtttaggaag 420
agtaatctga aacccttcga acgggacatc tccactgaaa tctaccaggc aggcaacaag 480
ccctgcaacg gggtggctgg ctttaactgc tactttcccc tgagatccta ctccttcaga 540
cccacctacg gcgtgggaca ccagccctac agggtggtgg tgctgagctt cgagctgctg 600
cacgcccccg caaccgtgtg cggacctaag aagtccacca acctggtgaa gaataagaga 660
gtgcagccca cagaatccat cgtgaggttt cccaacatta ccaatctgtg cccctttggc 720
gaagtgttca acgctaccag gttcgcatcc gtgtacgcct ggaatagaaa aaggatctcc 780
aactgcgtgg ccgactactc cgtgctgtac aattccgcct ccttttccac cttcaaatgc 840
tatggcgtga gccccaccaa gctgaatgat ctgtgtttta ccaacgtgta tgccgactcc 900
tttgtgatta ggggagacga ggtgaggcag atcgcccctg gccagaccgg caacatcgcc 960
gactataact acaagctgcc cgacgatttc accggctgtg tgattgcttg gaactccaat 1020
aatctggata gcaaggtggg gggaaactac aattacctgt acaggctgtt caggaagagt 1080
aacctgaagc cattcgaaag ggatatctcc actgagatct accaggctgg ctccaccccc 1140
tgcaacggag tgaagggatt caactgctat ttccccctgc agagctacgg cttccagcca 1200
acctacggcg tgggctacca gccctatagg gtggtggtgc tgtctttcga gctgctgcat 1260
gcccccgcca cagtgtgtgg ccccaagaaa tccaccaacc tggtgaagaa caaaagggtg 1320
cagcccaccg agagcatcgt gaggttcccc aacatcacca acctgtgccc cttcggcgag 1380
gtgttcaacg ccaccaggtt cgccagcgtg tacgcctgga acaggaagag gatctcaaac 1440
tgcgtggcag actacagtgt gctgtacaac tccgcctcct tctccacatt caagtgctat 1500
ggcgtgtccc ccaccaagct gaacgatctg tgctttacca acgtgtacgc cgactccttc 1560
gtgattaggg gcgacgaggt gcggcagatt gcccccggtc agaccggcaa gatcgccgat 1620
tataactaca aactgcccga cgacttcacc ggctgcgtga tcgcctggaa ctccaacaat 1680
ctggactcta aggtgggggg caactacaac taccggtaca gactgttccg caaaagcaac 1740
ctgaagccct tcgaaaggga catctccaca gagatctacc aggccggctc caaaccctgc 1800
aacggcgtgg agggcttcaa ctgctacttc cccctgcagt cctacggctt ccagcccacc 1860
aacggcgtgg gataccagcc ctacagagtg gtggtgctgt ccttcgagct gctgcacgca 1920
cctgccacag tgtgcggccc caagaagtct accaacctgg tgaaaaacaa g 1971
<210> 18
<211> 1971
<212> DNA
<213> Artificial
<400> 18
agggtgcagc ccaccgagag catcgtgagg ttccccaaca tcaccaacct gtgccccttc 60
ggcgaggtgt tcaacgccac caggttcgcc agcgtgtacg cctggaacag gaagaggatc 120
tcaaactgcg tggcagacta cagtgtgctg tacaactccg cctccttctc cacattcaag 180
tgctatggcg tgtcccccac caagctgaac gatctgtgct ttaccaacgt gtacgccgac 240
tccttcgtga ttaggggcga cgaggtgcgg cagattgccc ccggtcagac cggcaagatc 300
gccgattata actacaaact gcccgacgac ttcaccggct gcgtgatcgc ctggaactcc 360
aacaatctgg actctaaggt ggggggcaac tacaactacc ggtacagact gttccgcaaa 420
agcaacctga agcccttcga aagggacatc tccacagaga tctaccaggc cggctccaaa 480
ccctgcaacg gcgtggaggg cttcaactgc tacttccccc tgcagtccta cggcttccag 540
cccaccaacg gcgtgggata ccagccctac agagtggtgg tgctgtcctt cgagctgctg 600
cacgcacctg ccacagtgtg cggccccaag aagtctacca acctggtgaa aaacaagagg 660
gtgcagccca ccgagagcat cgtgaggttc cccaacatca ccaacctgtg ccccttcggc 720
gaggtgttca acgccaccag gttcgccagc gtgtacgcct ggaacaggaa gaggatctca 780
aactgcgtgg cagactacag tgtgctgtac aactccgcct ccttctccac attcaagtgc 840
tatggcgtgt cccccaccaa gctgaacgat ctgtgcttta ccaacgtgta cgccgactcc 900
ttcgtgatta ggggcgacga ggtgcggcag attgcccccg gtcagaccgg caagatcgcc 960
gattataact acaaactgcc cgacgacttc accggctgcg tgatcgcctg gaactccaac 1020
aatctggact ctaaggtggg gggcaactac aactaccggt acagactgtt ccgcaaaagc 1080
aacctgaagc ccttcgaaag ggacatctcc acagagatct accaggccgg ctccaaaccc 1140
tgcaacggcg tggagggctt caactgctac ttccccctgc agtcctacgg cttccagccc 1200
accaacggcg tgggatacca gccctacaga gtggtggtgc tgtccttcga gctgctgcac 1260
gcacctgcca cagtgtgcgg ccccaagaag tctaccaacc tggtgaaaaa caagagggtg 1320
cagcctaccg agtccatcgt gcggttcccc aatatcacaa acctgtgtcc cttcgacgag 1380
gtgtttaacg ccacccggtt cgcctccgtg tacgcttgga atagaaagag aatctctaac 1440
tgcgtggctg actactccgt gctgtataac ctggccccct tctttacctt caagtgttat 1500
ggcgtgtctc ccactaagct gaacgacctg tgcttcacca acgtgtacgc tgactccttc 1560
gtgatcaggg gcgacgaagt gcggcagatc gcccccggtc agacaggcaa catcgctgat 1620
tacaactaca aactgcctga cgacttcact ggctgcgtga ttgcctggaa cagcaacaag 1680
ctggacagca aggtgtccgg caactacaat tatctgtaca ggctgtttag gaagagtaat 1740
ctgaaaccct tcgaacggga catctccact gaaatctacc aggcaggcaa caagccctgc 1800
aacggggtgg ctggctttaa ctgctacttt cccctgagat cctactcctt cagacccacc 1860
tacggcgtgg gacaccagcc ctacagggtg gtggtgctga gcttcgagct gctgcacgcc 1920
cccgcaaccg tgtgcggacc taagaagtcc accaacctgg tgaagaataa g 1971
<210> 19
<211> 1971
<212> DNA
<213> Artificial
<400> 19
agagtgcagc ccacagaatc catcgtgagg tttcccaaca ttaccaatct gtgccccttt 60
ggcgaagtgt tcaacgctac caggttcgca tccgtgtacg cctggaatag aaaaaggatc 120
tccaactgcg tggccgacta ctccgtgctg tacaattccg cctccttttc caccttcaaa 180
tgctatggcg tgagccccac caagctgaat gatctgtgtt ttaccaacgt gtatgccgac 240
tcctttgtga ttaggggaga cgaggtgagg cagatcgccc ctggccagac cggcaacatc 300
gccgactata actacaagct gcccgacgat ttcaccggct gtgtgattgc ttggaactcc 360
aataatctgg atagcaaggt ggggggaaac tacaattacc tgtacaggct gttcaggaag 420
agtaacctga agccattcga aagggatatc tccactgaga tctaccaggc tggctccacc 480
ccctgcaacg gagtgaaggg attcaactgc tatttccccc tgcagagcta cggcttccag 540
ccaacctacg gcgtgggcta ccagccctat agggtggtgg tgctgtcttt cgagctgctg 600
catgcccccg ccacagtgtg tggccccaag aaatccacca acctggtgaa gaacaaaaga 660
gtgcagccca cagaatccat cgtgaggttt cccaacatta ccaatctgtg cccctttggc 720
gaagtgttca acgctaccag gttcgcatcc gtgtacgcct ggaatagaaa aaggatctcc 780
aactgcgtgg ccgactactc cgtgctgtac aattccgcct ccttttccac cttcaaatgc 840
tatggcgtga gccccaccaa gctgaatgat ctgtgtttta ccaacgtgta tgccgactcc 900
tttgtgatta ggggagacga ggtgaggcag atcgcccctg gccagaccgg caacatcgcc 960
gactataact acaagctgcc cgacgatttc accggctgtg tgattgcttg gaactccaat 1020
aatctggata gcaaggtggg gggaaactac aattacctgt acaggctgtt caggaagagt 1080
aacctgaagc cattcgaaag ggatatctcc actgagatct accaggctgg ctccaccccc 1140
tgcaacggag tgaagggatt caactgctat ttccccctgc agagctacgg cttccagcca 1200
acctacggcg tgggctacca gccctatagg gtggtggtgc tgtctttcga gctgctgcat 1260
gcccccgcca cagtgtgtgg ccccaagaaa tccaccaacc tggtgaagaa caaaagggtg 1320
cagcctaccg agtccatcgt gcggttcccc aatatcacaa acctgtgtcc cttcgacgag 1380
gtgtttaacg ccacccggtt cgcctccgtg tacgcttgga atagaaagag aatctctaac 1440
tgcgtggctg actactccgt gctgtataac ctggccccct tctttacctt caagtgttat 1500
ggcgtgtctc ccactaagct gaacgacctg tgcttcacca acgtgtacgc tgactccttc 1560
gtgatcaggg gcgacgaagt gcggcagatc gcccccggtc agacaggcaa catcgctgat 1620
tacaactaca aactgcctga cgacttcact ggctgcgtga ttgcctggaa cagcaacaag 1680
ctggacagca aggtgtccgg caactacaat tatctgtaca ggctgtttag gaagagtaat 1740
ctgaaaccct tcgaacggga catctccact gaaatctacc aggcaggcaa caagccctgc 1800
aacggggtgg ctggctttaa ctgctacttt cccctgagat cctactcctt cagacccacc 1860
tacggcgtgg gacaccagcc ctacagggtg gtggtgctga gcttcgagct gctgcacgcc 1920
cccgcaaccg tgtgcggacc taagaagtcc accaacctgg tgaagaataa g 1971
<210> 20
<211> 1971
<212> DNA
<213> Artificial
<400> 20
agggtgcagc ccaccgagag catcgtgagg ttccccaaca tcaccaacct gtgccccttc 60
ggcgaggtgt tcaacgccac caggttcgcc agcgtgtacg cctggaacag gaagaggatc 120
tcaaactgcg tggcagacta cagtgtgctg tacaactccg cctccttctc cacattcaag 180
tgctatggcg tgtcccccac caagctgaac gatctgtgct ttaccaacgt gtacgccgac 240
tccttcgtga ttaggggcga cgaggtgcgg cagattgccc ccggtcagac cggcaagatc 300
gccgattata actacaaact gcccgacgac ttcaccggct gcgtgatcgc ctggaactcc 360
aacaatctgg actctaaggt ggggggcaac tacaactacc ggtacagact gttccgcaaa 420
agcaacctga agcccttcga aagggacatc tccacagaga tctaccaggc cggctccaaa 480
ccctgcaacg gcgtggaggg cttcaactgc tacttccccc tgcagtccta cggcttccag 540
cccaccaacg gcgtgggata ccagccctac agagtggtgg tgctgtcctt cgagctgctg 600
cacgcacctg ccacagtgtg cggccccaag aagtctacca acctggtgaa aaacaagagg 660
gtgcagccca ccgagagcat cgtgaggttc cccaacatca ccaacctgtg ccccttcggc 720
gaggtgttca acgccaccag gttcgccagc gtgtacgcct ggaacaggaa gaggatctca 780
aactgcgtgg cagactacag tgtgctgtac aactccgcct ccttctccac attcaagtgc 840
tatggcgtgt cccccaccaa gctgaacgat ctgtgcttta ccaacgtgta cgccgactcc 900
ttcgtgatta ggggcgacga ggtgcggcag attgcccccg gtcagaccgg caagatcgcc 960
gattataact acaaactgcc cgacgacttc accggctgcg tgatcgcctg gaactccaac 1020
aatctggact ctaaggtggg gggcaactac aactaccggt acagactgtt ccgcaaaagc 1080
aacctgaagc ccttcgaaag ggacatctcc acagagatct accaggccgg ctccaaaccc 1140
tgcaacggcg tggagggctt caactgctac ttccccctgc agtcctacgg cttccagccc 1200
accaacggcg tgggatacca gccctacaga gtggtggtgc tgtccttcga gctgctgcac 1260
gcacctgcca cagtgtgcgg ccccaagaag tctaccaacc tggtgaaaaa caagagggtg 1320
cagcccaccg agagcatcgt gaggttcccc aacatcacca acctgtgccc cttcggcgag 1380
gtgttcaacg ccaccaggtt cgccagcgtg tacgcctgga acaggaagag gatctcaaac 1440
tgcgtggcag actacagtgt gctgtacaac tccgcctcct tctccacatt caagtgctat 1500
ggcgtgtccc ccaccaagct gaacgatctg tgctttacca acgtgtacgc cgactccttc 1560
gtgattaggg gcgacgaggt gcggcagatt gcccccggtc agaccggcaa gatcgccgat 1620
tataactaca aactgcccga cgacttcacc ggctgcgtga tcgcctggaa ctccaacaat 1680
ctggactcta aggtgggggg caactacaac taccggtaca gactgttccg caaaagcaac 1740
ctgaagccct tcgaaaggga catctccaca gagatctacc aggccggctc caaaccctgc 1800
aacggcgtgg agggcttcaa ctgctacttc cccctgcagt cctacggctt ccagcccacc 1860
aacggcgtgg gataccagcc ctacagagtg gtggtgctgt ccttcgagct gctgcacgca 1920
cctgccacag tgtgcggccc caagaagtct accaacctgg tgaaaaacaa g 1971
<210> 21
<211> 1971
<212> DNA
<213> Artificial
<400> 21
agagtgcagc ccacagaatc catcgtgagg tttcccaaca ttaccaatct gtgccccttt 60
ggcgaagtgt tcaacgctac caggttcgca tccgtgtacg cctggaatag aaaaaggatc 120
tccaactgcg tggccgacta ctccgtgctg tacaattccg cctccttttc caccttcaaa 180
tgctatggcg tgagccccac caagctgaat gatctgtgtt ttaccaacgt gtatgccgac 240
tcctttgtga ttaggggaga cgaggtgagg cagatcgccc ctggccagac cggcaacatc 300
gccgactata actacaagct gcccgacgat ttcaccggct gtgtgattgc ttggaactcc 360
aataatctgg atagcaaggt ggggggaaac tacaattacc tgtacaggct gttcaggaag 420
agtaacctga agccattcga aagggatatc tccactgaga tctaccaggc tggctccacc 480
ccctgcaacg gagtgaaggg attcaactgc tatttccccc tgcagagcta cggcttccag 540
ccaacctacg gcgtgggcta ccagccctat agggtggtgg tgctgtcttt cgagctgctg 600
catgcccccg ccacagtgtg tggccccaag aaatccacca acctggtgaa gaacaaaaga 660
gtgcagccca cagaatccat cgtgaggttt cccaacatta ccaatctgtg cccctttggc 720
gaagtgttca acgctaccag gttcgcatcc gtgtacgcct ggaatagaaa aaggatctcc 780
aactgcgtgg ccgactactc cgtgctgtac aattccgcct ccttttccac cttcaaatgc 840
tatggcgtga gccccaccaa gctgaatgat ctgtgtttta ccaacgtgta tgccgactcc 900
tttgtgatta ggggagacga ggtgaggcag atcgcccctg gccagaccgg caacatcgcc 960
gactataact acaagctgcc cgacgatttc accggctgtg tgattgcttg gaactccaat 1020
aatctggata gcaaggtggg gggaaactac aattacctgt acaggctgtt caggaagagt 1080
aacctgaagc cattcgaaag ggatatctcc actgagatct accaggctgg ctccaccccc 1140
tgcaacggag tgaagggatt caactgctat ttccccctgc agagctacgg cttccagcca 1200
acctacggcg tgggctacca gccctatagg gtggtggtgc tgtctttcga gctgctgcat 1260
gcccccgcca cagtgtgtgg ccccaagaaa tccaccaacc tggtgaagaa caaaagagtg 1320
cagcccacag aatccatcgt gaggtttccc aacattacca atctgtgccc ctttggcgaa 1380
gtgttcaacg ctaccaggtt cgcatccgtg tacgcctgga atagaaaaag gatctccaac 1440
tgcgtggccg actactccgt gctgtacaat tccgcctcct tttccacctt caaatgctat 1500
ggcgtgagcc ccaccaagct gaatgatctg tgttttacca acgtgtatgc cgactccttt 1560
gtgattaggg gagacgaggt gaggcagatc gcccctggcc agaccggcaa catcgccgac 1620
tataactaca agctgcccga cgatttcacc ggctgtgtga ttgcttggaa ctccaataat 1680
ctggatagca aggtgggggg aaactacaat tacctgtaca ggctgttcag gaagagtaac 1740
ctgaagccat tcgaaaggga tatctccact gagatctacc aggctggctc caccccctgc 1800
aacggagtga agggattcaa ctgctatttc cccctgcaga gctacggctt ccagccaacc 1860
tacggcgtgg gctaccagcc ctatagggtg gtggtgctgt ctttcgagct gctgcatgcc 1920
cccgccacag tgtgtggccc caagaaatcc accaacctgg tgaagaacaa a 1971
<210> 22
<211> 1971
<212> DNA
<213> Artificial
<400> 22
agggtgcagc ctaccgagtc catcgtgcgg ttccccaata tcacaaacct gtgtcccttc 60
gacgaggtgt ttaacgccac ccggttcgcc tccgtgtacg cttggaatag aaagagaatc 120
tctaactgcg tggctgacta ctccgtgctg tataacctgg cccccttctt taccttcaag 180
tgttatggcg tgtctcccac taagctgaac gacctgtgct tcaccaacgt gtacgctgac 240
tccttcgtga tcaggggcga cgaagtgcgg cagatcgccc ccggtcagac aggcaacatc 300
gctgattaca actacaaact gcctgacgac ttcactggct gcgtgattgc ctggaacagc 360
aacaagctgg acagcaaggt gtccggcaac tacaattatc tgtacaggct gtttaggaag 420
agtaatctga aacccttcga acgggacatc tccactgaaa tctaccaggc aggcaacaag 480
ccctgcaacg gggtggctgg ctttaactgc tactttcccc tgagatccta ctccttcaga 540
cccacctacg gcgtgggaca ccagccctac agggtggtgg tgctgagctt cgagctgctg 600
cacgcccccg caaccgtgtg cggacctaag aagtccacca acctggtgaa gaataagagg 660
gtgcagccta ccgagtccat cgtgcggttc cccaatatca caaacctgtg tcccttcgac 720
gaggtgttta acgccacccg gttcgcctcc gtgtacgctt ggaatagaaa gagaatctct 780
aactgcgtgg ctgactactc cgtgctgtat aacctggccc ccttctttac cttcaagtgt 840
tatggcgtgt ctcccactaa gctgaacgac ctgtgcttca ccaacgtgta cgctgactcc 900
ttcgtgatca ggggcgacga agtgcggcag atcgcccccg gtcagacagg caacatcgct 960
gattacaact acaaactgcc tgacgacttc actggctgcg tgattgcctg gaacagcaac 1020
aagctggaca gcaaggtgtc cggcaactac aattatctgt acaggctgtt taggaagagt 1080
aatctgaaac ccttcgaacg ggacatctcc actgaaatct accaggcagg caacaagccc 1140
tgcaacgggg tggctggctt taactgctac tttcccctga gatcctactc cttcagaccc 1200
acctacggcg tgggacacca gccctacagg gtggtggtgc tgagcttcga gctgctgcac 1260
gcccccgcaa ccgtgtgcgg acctaagaag tccaccaacc tggtgaagaa taagagggtg 1320
cagcctaccg agtccatcgt gcggttcccc aatatcacaa acctgtgtcc cttcgacgag 1380
gtgtttaacg ccacccggtt cgcctccgtg tacgcttgga atagaaagag aatctctaac 1440
tgcgtggctg actactccgt gctgtataac ctggccccct tctttacctt caagtgttat 1500
ggcgtgtctc ccactaagct gaacgacctg tgcttcacca acgtgtacgc tgactccttc 1560
gtgatcaggg gcgacgaagt gcggcagatc gcccccggtc agacaggcaa catcgctgat 1620
tacaactaca aactgcctga cgacttcact ggctgcgtga ttgcctggaa cagcaacaag 1680
ctggacagca aggtgtccgg caactacaat tatctgtaca ggctgtttag gaagagtaat 1740
ctgaaaccct tcgaacggga catctccact gaaatctacc aggcaggcaa caagccctgc 1800
aacggggtgg ctggctttaa ctgctacttt cccctgagat cctactcctt cagacccacc 1860
tacggcgtgg gacaccagcc ctacagggtg gtggtgctga gcttcgagct gctgcacgcc 1920
cccgcaaccg tgtgcggacc taagaagtcc accaacctgg tgaagaataa g 1971
<210> 23
<211> 658
<212> PRT
<213> Artificial
<400> 23
Met Arg Val Gln Pro Thr Glu Ser Ile Val Arg Phe Pro Asn Ile Thr
1 5 10 15
Asn Leu Cys Pro Phe Gly Glu Val Phe Asn Ala Thr Arg Phe Ala Ser
20 25 30
Val Tyr Ala Trp Asn Arg Lys Arg Ile Ser Asn Cys Val Ala Asp Tyr
35 40 45
Ser Val Leu Tyr Asn Ser Ala Ser Phe Ser Thr Phe Lys Cys Tyr Gly
50 55 60
Val Ser Pro Thr Lys Leu Asn Asp Leu Cys Phe Thr Asn Val Tyr Ala
65 70 75 80
Asp Ser Phe Val Ile Arg Gly Asp Glu Val Arg Gln Ile Ala Pro Gly
85 90 95
Gln Thr Gly Lys Ile Ala Asp Tyr Asn Tyr Lys Leu Pro Asp Asp Phe
100 105 110
Thr Gly Cys Val Ile Ala Trp Asn Ser Asn Asn Leu Asp Ser Lys Val
115 120 125
Gly Gly Asn Tyr Asn Tyr Leu Tyr Arg Leu Phe Arg Lys Ser Asn Leu
130 135 140
Lys Pro Phe Glu Arg Asp Ile Ser Thr Glu Ile Tyr Gln Ala Gly Ser
145 150 155 160
Thr Pro Cys Asn Gly Val Glu Gly Phe Asn Cys Tyr Phe Pro Leu Gln
165 170 175
Ser Tyr Gly Phe Gln Pro Thr Asn Gly Val Gly Tyr Gln Pro Tyr Arg
180 185 190
Val Val Val Leu Ser Phe Glu Leu Leu His Ala Pro Ala Thr Val Cys
195 200 205
Gly Pro Lys Lys Ser Thr Asn Leu Val Lys Asn Lys Arg Val Gln Pro
210 215 220
Thr Glu Ser Ile Val Arg Phe Pro Asn Ile Thr Asn Leu Cys Pro Phe
225 230 235 240
Gly Glu Val Phe Asn Ala Thr Arg Phe Ala Ser Val Tyr Ala Trp Asn
245 250 255
Arg Lys Arg Ile Ser Asn Cys Val Ala Asp Tyr Ser Val Leu Tyr Asn
260 265 270
Ser Ala Ser Phe Ser Thr Phe Lys Cys Tyr Gly Val Ser Pro Thr Lys
275 280 285
Leu Asn Asp Leu Cys Phe Thr Asn Val Tyr Ala Asp Ser Phe Val Ile
290 295 300
Arg Gly Asp Glu Val Arg Gln Ile Ala Pro Gly Gln Thr Gly Lys Ile
305 310 315 320
Ala Asp Tyr Asn Tyr Lys Leu Pro Asp Asp Phe Thr Gly Cys Val Ile
325 330 335
Ala Trp Asn Ser Asn Asn Leu Asp Ser Lys Val Gly Gly Asn Tyr Asn
340 345 350
Tyr Leu Tyr Arg Leu Phe Arg Lys Ser Asn Leu Lys Pro Phe Glu Arg
355 360 365
Asp Ile Ser Thr Glu Ile Tyr Gln Ala Gly Ser Thr Pro Cys Asn Gly
370 375 380
Val Glu Gly Phe Asn Cys Tyr Phe Pro Leu Gln Ser Tyr Gly Phe Gln
385 390 395 400
Pro Thr Asn Gly Val Gly Tyr Gln Pro Tyr Arg Val Val Val Leu Ser
405 410 415
Phe Glu Leu Leu His Ala Pro Ala Thr Val Cys Gly Pro Lys Lys Ser
420 425 430
Thr Asn Leu Val Lys Asn Lys Arg Val Gln Pro Thr Glu Ser Ile Val
435 440 445
Arg Phe Pro Asn Ile Thr Asn Leu Cys Pro Phe Gly Glu Val Phe Asn
450 455 460
Ala Thr Arg Phe Ala Ser Val Tyr Ala Trp Asn Arg Lys Arg Ile Ser
465 470 475 480
Asn Cys Val Ala Asp Tyr Ser Val Leu Tyr Asn Ser Ala Ser Phe Ser
485 490 495
Thr Phe Lys Cys Tyr Gly Val Ser Pro Thr Lys Leu Asn Asp Leu Cys
500 505 510
Phe Thr Asn Val Tyr Ala Asp Ser Phe Val Ile Arg Gly Asp Glu Val
515 520 525
Arg Gln Ile Ala Pro Gly Gln Thr Gly Lys Ile Ala Asp Tyr Asn Tyr
530 535 540
Lys Leu Pro Asp Asp Phe Thr Gly Cys Val Ile Ala Trp Asn Ser Asn
545 550 555 560
Asn Leu Asp Ser Lys Val Gly Gly Asn Tyr Asn Tyr Leu Tyr Arg Leu
565 570 575
Phe Arg Lys Ser Asn Leu Lys Pro Phe Glu Arg Asp Ile Ser Thr Glu
580 585 590
Ile Tyr Gln Ala Gly Ser Thr Pro Cys Asn Gly Val Glu Gly Phe Asn
595 600 605
Cys Tyr Phe Pro Leu Gln Ser Tyr Gly Phe Gln Pro Thr Asn Gly Val
610 615 620
Gly Tyr Gln Pro Tyr Arg Val Val Val Leu Ser Phe Glu Leu Leu His
625 630 635 640
Ala Pro Ala Thr Val Cys Gly Pro Lys Lys Ser Thr Asn Leu Val Lys
645 650 655
Asn Lys
<210> 24
<211> 658
<212> PRT
<213> Artificial
<400> 24
Met Arg Val Gln Pro Thr Glu Ser Ile Val Arg Phe Pro Asn Ile Thr
1 5 10 15
Asn Leu Cys Pro Phe Gly Glu Val Phe Asn Ala Thr Arg Phe Ala Ser
20 25 30
Val Tyr Ala Trp Asn Arg Lys Arg Ile Ser Asn Cys Val Ala Asp Tyr
35 40 45
Ser Val Leu Tyr Asn Ser Ala Ser Phe Ser Thr Phe Lys Cys Tyr Gly
50 55 60
Val Ser Pro Thr Lys Leu Asn Asp Leu Cys Phe Thr Asn Val Tyr Ala
65 70 75 80
Asp Ser Phe Val Ile Arg Gly Asp Glu Val Arg Gln Ile Ala Pro Gly
85 90 95
Gln Thr Gly Lys Ile Ala Asp Tyr Asn Tyr Lys Leu Pro Asp Asp Phe
100 105 110
Thr Gly Cys Val Ile Ala Trp Asn Ser Asn Asn Leu Asp Ser Lys Val
115 120 125
Gly Gly Asn Tyr Asn Tyr Leu Tyr Arg Leu Phe Arg Lys Ser Asn Leu
130 135 140
Lys Pro Phe Glu Arg Asp Ile Ser Thr Glu Ile Tyr Gln Ala Gly Ser
145 150 155 160
Thr Pro Cys Asn Gly Val Glu Gly Phe Asn Cys Tyr Phe Pro Leu Gln
165 170 175
Ser Tyr Gly Phe Gln Pro Thr Asn Gly Val Gly Tyr Gln Pro Tyr Arg
180 185 190
Val Val Val Leu Ser Phe Glu Leu Leu His Ala Pro Ala Thr Val Cys
195 200 205
Gly Pro Lys Lys Ser Thr Asn Leu Val Lys Asn Lys Arg Val Gln Pro
210 215 220
Thr Glu Ser Ile Val Arg Phe Pro Asn Ile Thr Asn Leu Cys Pro Phe
225 230 235 240
Gly Glu Val Phe Asn Ala Thr Arg Phe Ala Ser Val Tyr Ala Trp Asn
245 250 255
Arg Lys Arg Ile Ser Asn Cys Val Ala Asp Tyr Ser Val Leu Tyr Asn
260 265 270
Ser Ala Ser Phe Ser Thr Phe Lys Cys Tyr Gly Val Ser Pro Thr Lys
275 280 285
Leu Asn Asp Leu Cys Phe Thr Asn Val Tyr Ala Asp Ser Phe Val Ile
290 295 300
Arg Gly Asp Glu Val Arg Gln Ile Ala Pro Gly Gln Thr Gly Asn Ile
305 310 315 320
Ala Asp Tyr Asn Tyr Lys Leu Pro Asp Asp Phe Thr Gly Cys Val Ile
325 330 335
Ala Trp Asn Ser Asn Asn Leu Asp Ser Lys Val Gly Gly Asn Tyr Asn
340 345 350
Tyr Leu Tyr Arg Leu Phe Arg Lys Ser Asn Leu Lys Pro Phe Glu Arg
355 360 365
Asp Ile Ser Thr Glu Ile Tyr Gln Ala Gly Ser Thr Pro Cys Asn Gly
370 375 380
Val Lys Gly Phe Asn Cys Tyr Phe Pro Leu Gln Ser Tyr Gly Phe Gln
385 390 395 400
Pro Thr Tyr Gly Val Gly Tyr Gln Pro Tyr Arg Val Val Val Leu Ser
405 410 415
Phe Glu Leu Leu His Ala Pro Ala Thr Val Cys Gly Pro Lys Lys Ser
420 425 430
Thr Asn Leu Val Lys Asn Lys Arg Val Gln Pro Thr Glu Ser Ile Val
435 440 445
Arg Phe Pro Asn Ile Thr Asn Leu Cys Pro Phe Gly Glu Val Phe Asn
450 455 460
Ala Thr Arg Phe Ala Ser Val Tyr Ala Trp Asn Arg Lys Arg Ile Ser
465 470 475 480
Asn Cys Val Ala Asp Tyr Ser Val Leu Tyr Asn Ser Ala Ser Phe Ser
485 490 495
Thr Phe Lys Cys Tyr Gly Val Ser Pro Thr Lys Leu Asn Asp Leu Cys
500 505 510
Phe Thr Asn Val Tyr Ala Asp Ser Phe Val Ile Arg Gly Asp Glu Val
515 520 525
Arg Gln Ile Ala Pro Gly Gln Thr Gly Lys Ile Ala Asp Tyr Asn Tyr
530 535 540
Lys Leu Pro Asp Asp Phe Thr Gly Cys Val Ile Ala Trp Asn Ser Asn
545 550 555 560
Asn Leu Asp Ser Lys Val Gly Gly Asn Tyr Asn Tyr Arg Tyr Arg Leu
565 570 575
Phe Arg Lys Ser Asn Leu Lys Pro Phe Glu Arg Asp Ile Ser Thr Glu
580 585 590
Ile Tyr Gln Ala Gly Ser Thr Pro Cys Asn Gly Val Gln Gly Phe Asn
595 600 605
Cys Tyr Phe Pro Leu Gln Ser Tyr Gly Phe Gln Pro Thr Asn Gly Val
610 615 620
Gly Tyr Gln Pro Tyr Arg Val Val Val Leu Ser Phe Glu Leu Leu His
625 630 635 640
Ala Pro Ala Thr Val Cys Gly Pro Lys Lys Ser Thr Asn Leu Val Lys
645 650 655
Asn Lys
Claims (23)
1. A recombinant novel coronavirus protein which is a trimer and comprises three subunits consisting of 319 th to 537 th amino acid fragments of an RBD region of a novel coronavirus S protein, and is characterized in that,
the first subunit of the recombinant novel coronavirus protein comprises an L452R and/or T478K mutation site; and
the second subunit of the recombinant novel coronavirus protein comprises one or more mutation sites selected from K417N, E484K or N501Y; and
five or more than five mutation sites selected from G339D, S371L, S373P, S375F, K417N, N440K, G446S, S477N, T478K, E484A, Q493R, G496S, Q498R, N501Y or Y505H are included in the third subunit of the recombinant novel coronavirus protein.
2. The recombinant novel coronavirus protein of claim 1, wherein the three subunits of the recombinant novel coronavirus protein have a primary structure in which the three subunits are connected in series in the order of N-terminus to C-terminus.
3. The recombinant novel coronavirus protein according to claim 1 or 2, wherein the three subunits of the recombinant novel coronavirus protein are connected in series in the order of the first subunit, the second subunit, and the third subunit.
4. The recombinant novel coronavirus protein according to claim 3, wherein the amino acid sequence of the recombinant novel coronavirus protein is represented by SEQ ID Nos. 1 to 11 or a sequence having 95% or more homology with the amino acid sequence thereof except the mutation site, preferably the amino acid sequence represented by SEQ ID No.1 or a sequence having 95% or more homology with the amino acid sequence thereof except the mutation site.
5. A fusion protein, characterized in that the amino acid sequence of the fusion protein comprises the amino acid sequence of the recombinant novel coronavirus protein as defined in any one of claims 1 to 4.
6. The fusion protein of claim 5, further comprising one or more selected from a signal peptide, a tag, and an immune enhancing peptide.
7. A nucleic acid molecule comprising a nucleotide sequence encoding a recombinant novel coronavirus protein as claimed in any one of claims 1 to 4, or encoding a fusion protein as claimed in claim 5 or 6.
8. The nucleic acid molecule of claim 7, wherein the nucleotide sequence of said nucleic acid molecule is the nucleotide sequence shown as SEQ ID nos. 12-22.
9. A vector comprising the nucleic acid molecule of claim 7.
10. A host cell comprising the nucleic acid molecule of claim 7 or the vector of claim 9.
11. The host cell of claim 10, wherein the host cell is an escherichia coli, yeast cell, insect cell, or mammalian cell.
12. The host cell of claim 11, wherein the host cell is a CHO cell.
13. A method for producing a recombinant novel coronavirus protein according to any one of claims 1 to 4 or a fusion protein according to claim 5 or 6, comprising the steps of:
step A) preparing the nucleic acid molecule of claim 7, constructing an expression vector for the nucleic acid molecule, and transforming or transfecting the expression vector into a host cell;
step B) protein expression using the product of step A);
and C) purifying the expression product obtained in the step B) to obtain the recombinant novel coronavirus protein or the recombinant novel coronavirus fusion protein.
14. Use of a recombinant novel coronavirus protein as defined in any one of claims 1 to 4, a fusion protein as defined in claim 5 or 6, a nucleic acid molecule as defined in claim 7, a vector as defined in claim 9 or a host cell as defined in claim 10 for the preparation of a medicament for the treatment and/or prevention of a novel coronavirus infection and/or a novel coronavirus-induced disease.
15. Use of a recombinant novel coronavirus protein as defined in any one of claims 1 to 4, a fusion protein as defined in claim 5 or 6, a nucleic acid molecule as defined in claim 7, a vector as defined in claim 9 or a host cell as defined in claim 10 for the preparation of a medicament for boosting the immunity of a human being vaccinated with a novel coronavirus vaccine; the novel coronavirus vaccine is preferably a novel coronavirus vaccine inactivated vaccine.
16. A recombinant protein vaccine, comprising a recombinant novel coronavirus protein according to any one of claims 1-4 or a fusion protein according to claim 5 or 6, and an adjuvant.
17. The recombinant protein vaccine of claim 15, wherein the adjuvant is aluminum hydroxide, aluminum phosphate, MF59, or CpG.
18. The recombinant protein vaccine of claim 16, wherein the adjuvant is aluminum hydroxide.
19. The method of producing a recombinant protein vaccine according to claim 16, 17 or 18, wherein the purified recombinant novel coronavirus protein or the fusion protein is mixed with the adjuvant.
20. A genetically engineered vector vaccine comprising the nucleic acid molecule of claim 7.
21. A nucleic acid vaccine comprising the nucleic acid molecule of claim 7.
22. A pharmaceutical composition comprising the vaccine of claim 16, 20 or 21, and a pharmaceutically acceptable carrier.
23. A method of eliciting an immune response against a novel coronavirus or treating a novel coronavirus infection in a subject, wherein an effective amount of the vaccine of claim 16, 20 or 21 or the pharmaceutical composition of claim 22 is administered to the subject.
Priority Applications (1)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| CN202210073510.6A CN114315989A (en) | 2022-01-21 | 2022-01-21 | Recombinant novel coronavirus protein vaccine, preparation method and application thereof |
Applications Claiming Priority (1)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| CN202210073510.6A CN114315989A (en) | 2022-01-21 | 2022-01-21 | Recombinant novel coronavirus protein vaccine, preparation method and application thereof |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| CN114315989A true CN114315989A (en) | 2022-04-12 |
Family
ID=81028990
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| CN202210073510.6A Pending CN114315989A (en) | 2022-01-21 | 2022-01-21 | Recombinant novel coronavirus protein vaccine, preparation method and application thereof |
Country Status (1)
| Country | Link |
|---|---|
| CN (1) | CN114315989A (en) |
Cited By (6)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN114573667A (en) * | 2022-05-06 | 2022-06-03 | 艾棣维欣(苏州)生物制药有限公司 | Mutant strain DNA vaccine of SARS-CoV-2 virus B.1.1.529 and application thereof |
| CN114989308A (en) * | 2022-05-12 | 2022-09-02 | 中国科学院微生物研究所 | Novel coronavirus chimeric nucleic acid vaccine and use thereof |
| CN115043915A (en) * | 2022-05-25 | 2022-09-13 | 中山大学 | A method for enhancing the immunogenicity of a new coronavirus variant strain and its application |
| CN115368468A (en) * | 2021-05-18 | 2022-11-22 | 珠海市丽珠单抗生物技术有限公司 | A kind of novel coronavirus SARS-CoV-2 mutant vaccine and its application |
| CN115678906A (en) * | 2022-05-12 | 2023-02-03 | 中国科学院微生物研究所 | Optimized novel coronavirus chimeric nucleic acid vaccine and application thereof |
| CN115724999A (en) * | 2022-05-11 | 2023-03-03 | 中国科学院微生物研究所 | Tandem hybrid trimer new crown vaccine |
Citations (2)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN113292640A (en) * | 2021-06-18 | 2021-08-24 | 国药中生生物技术研究院有限公司 | Novel recombinant coronavirus RBD trimer protein vaccine capable of generating broad-spectrum cross-neutralization activity, and preparation method and application thereof |
| CN113817029A (en) * | 2021-03-31 | 2021-12-21 | 国药中生生物技术研究院有限公司 | A novel coronavirus S-RBD trimer protein vaccine, its preparation method and application |
-
2022
- 2022-01-21 CN CN202210073510.6A patent/CN114315989A/en active Pending
Patent Citations (2)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN113817029A (en) * | 2021-03-31 | 2021-12-21 | 国药中生生物技术研究院有限公司 | A novel coronavirus S-RBD trimer protein vaccine, its preparation method and application |
| CN113292640A (en) * | 2021-06-18 | 2021-08-24 | 国药中生生物技术研究院有限公司 | Novel recombinant coronavirus RBD trimer protein vaccine capable of generating broad-spectrum cross-neutralization activity, and preparation method and application thereof |
Non-Patent Citations (2)
| Title |
|---|
| XUEMEI HE等: "SARS-CoV-2 Omicron variant: Characteristics and prevention", 《MEDCOMM》 * |
| 綦佩妍: "带你了解新冠疫苗混打及序贯免疫", 《中国国门时报》 * |
Cited By (12)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN115368468A (en) * | 2021-05-18 | 2022-11-22 | 珠海市丽珠单抗生物技术有限公司 | A kind of novel coronavirus SARS-CoV-2 mutant vaccine and its application |
| CN115368468B (en) * | 2021-05-18 | 2025-09-23 | 珠海市丽珠单抗生物技术有限公司 | A novel coronavirus SARS-CoV-2 mutant vaccine and its application |
| CN114573667A (en) * | 2022-05-06 | 2022-06-03 | 艾棣维欣(苏州)生物制药有限公司 | Mutant strain DNA vaccine of SARS-CoV-2 virus B.1.1.529 and application thereof |
| CN114573667B (en) * | 2022-05-06 | 2022-08-02 | 艾棣维欣(苏州)生物制药有限公司 | Mutant strain DNA vaccine of SARS-CoV-2 virus B.1.1.529 and application thereof |
| CN115724999A (en) * | 2022-05-11 | 2023-03-03 | 中国科学院微生物研究所 | Tandem hybrid trimer new crown vaccine |
| CN115724999B (en) * | 2022-05-11 | 2023-09-19 | 中国科学院微生物研究所 | Tandem hybrid trimer COVID-19 vaccine |
| WO2023217005A1 (en) * | 2022-05-11 | 2023-11-16 | 中国科学院微生物研究所 | Tandem-type hybrid trimeric sars-cov-2 vaccine |
| CN114989308A (en) * | 2022-05-12 | 2022-09-02 | 中国科学院微生物研究所 | Novel coronavirus chimeric nucleic acid vaccine and use thereof |
| CN115678906A (en) * | 2022-05-12 | 2023-02-03 | 中国科学院微生物研究所 | Optimized novel coronavirus chimeric nucleic acid vaccine and application thereof |
| CN115678906B (en) * | 2022-05-12 | 2023-09-19 | 中国科学院微生物研究所 | Optimized novel coronavirus chimeric nucleic acid vaccine and its use |
| CN115043915A (en) * | 2022-05-25 | 2022-09-13 | 中山大学 | A method for enhancing the immunogenicity of a new coronavirus variant strain and its application |
| CN115043915B (en) * | 2022-05-25 | 2023-10-24 | 中山大学 | Method for enhancing immunogenicity of novel coronavirus variant strain and application thereof |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| CN113292640B (en) | A recombinant novel coronavirus RBD trimeric protein vaccine producing broad-spectrum cross-neutralizing activity, its preparation method and application | |
| CN114315989A (en) | Recombinant novel coronavirus protein vaccine, preparation method and application thereof | |
| CN111560074B (en) | A novel coronavirus S protein single-domain subunit nanovaccine based on Helicobacter pylori ferritin | |
| CN114478718B (en) | Recombinant novel coronavirus protein vaccine, preparation method and application thereof | |
| CN113817029B (en) | Novel coronavirus S-RBD trimer protein vaccine, preparation method and application thereof | |
| KR102713707B1 (en) | Novel multivalent nanoparticle-based vaccine | |
| CN113461787A (en) | Recombinant novel coronavirus S-RBD trimer protein, and preparation method and application thereof | |
| CN113943373A (en) | Beta coronavirus polymer antigen, preparation method and application thereof | |
| CN116478296A (en) | Truncated respiratory syncytial virus F protein and use thereof | |
| CN114478717B (en) | Recombinant novel coronavirus protein vaccine, preparation method and application thereof | |
| HK40073048A (en) | A recombinant protein-based covid-19 vaccine as well as its preparation method and application thereof | |
| HK40073046A (en) | A recombinant protein-based covid-19 vaccine as well as its preparation method and application thereof | |
| HK40073047A (en) | A recombinant protein-based covid-19 vaccine as well as its preparation method and application thereof | |
| HK40073046B (en) | A recombinant protein-based covid-19 vaccine as well as its preparation method and application thereof | |
| CN103740736B (en) | The HSV2 virus gE glucoprotein extracellular region gene fragment of chemosynthesis and expression, application | |
| HK40073047B (en) | A recombinant protein-based covid-19 vaccine as well as its preparation method and application thereof | |
| KR101526886B1 (en) | Recombinant Protein Comprising Epitope of Fowl Adenovirus fiber 2 Protein and Antibody thereto | |
| HK40067421A (en) | Recombinant trimeric rbd protein vaccine of novel coronavirus capable of generating broad-spectrum cross-neutralization activity, and preparation method and application thereof | |
| HK40067421B (en) | Recombinant trimeric rbd protein vaccine of novel coronavirus capable of generating broad-spectrum cross-neutralization activity, and preparation method and application thereof | |
| HK40067420B (en) | Recombinant trimeric rbd protein vaccine of novel coronavirus capable of generating broad-spectrum cross-neutralization activity, and preparation method and application thereof | |
| HK40067420A (en) | Recombinant trimeric rbd protein vaccine of novel coronavirus capable of generating broad-spectrum cross-neutralization activity, and preparation method and application thereof | |
| HK40054535B (en) | Recombinant trimeric rbd protein vaccine of novel coronavirus capable of generating broad-spectrum cross-neutralization activity, and preparation method and application thereof | |
| HK40054535A (en) | Recombinant trimeric rbd protein vaccine of novel coronavirus capable of generating broad-spectrum cross-neutralization activity, and preparation method and application thereof | |
| CN113372454A (en) | Nipah virus receptor binding glycoprotein and application thereof | |
| HK40057014A (en) | A novel coronavirus s-rbd trimer protein vaccine, preparation method and use thereof |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| PB01 | Publication | ||
| PB01 | Publication | ||
| SE01 | Entry into force of request for substantive examination | ||
| SE01 | Entry into force of request for substantive examination | ||
| REG | Reference to a national code |
Ref country code: HK Ref legal event code: DE Ref document number: 40073048 Country of ref document: HK |
|
| RJ01 | Rejection of invention patent application after publication | ||
| RJ01 | Rejection of invention patent application after publication |
Application publication date: 20220412 |