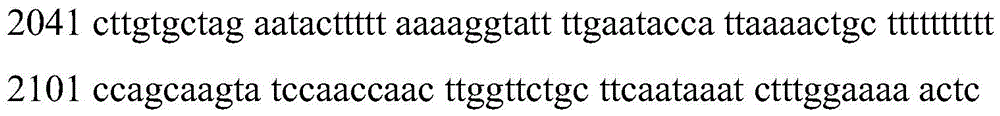CN114107502A - A diagnostic marker for upper urothelial carcinoma and its application - Google Patents
A diagnostic marker for upper urothelial carcinoma and its application Download PDFInfo
- Publication number
- CN114107502A CN114107502A CN202111431742.6A CN202111431742A CN114107502A CN 114107502 A CN114107502 A CN 114107502A CN 202111431742 A CN202111431742 A CN 202111431742A CN 114107502 A CN114107502 A CN 114107502A
- Authority
- CN
- China
- Prior art keywords
- cancer
- utuc
- biomarkers
- diagnostic marker
- urinary tract
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Pending
Links
- 206010044412 transitional cell carcinoma Diseases 0.000 title claims abstract description 25
- 239000003550 marker Substances 0.000 title claims abstract description 22
- 208000023747 urothelial carcinoma Diseases 0.000 title 1
- 239000000090 biomarker Substances 0.000 claims abstract description 35
- 102100040896 Growth/differentiation factor 15 Human genes 0.000 claims abstract description 26
- 101000893549 Homo sapiens Growth/differentiation factor 15 Proteins 0.000 claims abstract description 25
- 101000834948 Homo sapiens Tomoregulin-2 Proteins 0.000 claims abstract description 23
- 102100026160 Tomoregulin-2 Human genes 0.000 claims abstract description 23
- 101000803403 Homo sapiens Vimentin Proteins 0.000 claims abstract description 21
- 102100035071 Vimentin Human genes 0.000 claims abstract description 19
- 201000009030 Carcinoma Diseases 0.000 claims abstract description 17
- 238000000034 method Methods 0.000 claims abstract description 16
- 210000001635 urinary tract Anatomy 0.000 claims abstract description 16
- 108020004999 messenger RNA Proteins 0.000 claims abstract description 15
- 108090000623 proteins and genes Proteins 0.000 claims abstract description 15
- 230000007067 DNA methylation Effects 0.000 claims abstract description 8
- FWMNVWWHGCHHJJ-SKKKGAJSSA-N 4-amino-1-[(2r)-6-amino-2-[[(2r)-2-[[(2r)-2-[[(2r)-2-amino-3-phenylpropanoyl]amino]-3-phenylpropanoyl]amino]-4-methylpentanoyl]amino]hexanoyl]piperidine-4-carboxylic acid Chemical compound C([C@H](C(=O)N[C@H](CC(C)C)C(=O)N[C@H](CCCCN)C(=O)N1CCC(N)(CC1)C(O)=O)NC(=O)[C@H](N)CC=1C=CC=CC=1)C1=CC=CC=C1 FWMNVWWHGCHHJJ-SKKKGAJSSA-N 0.000 claims abstract description 4
- 238000011282 treatment Methods 0.000 claims description 20
- 238000013399 early diagnosis Methods 0.000 claims description 16
- 238000004393 prognosis Methods 0.000 claims description 10
- 238000011161 development Methods 0.000 claims description 6
- 230000014509 gene expression Effects 0.000 claims description 5
- 238000012795 verification Methods 0.000 claims description 5
- 230000008827 biological function Effects 0.000 claims description 4
- 230000003950 pathogenic mechanism Effects 0.000 claims description 4
- 239000012472 biological sample Substances 0.000 claims description 3
- 238000011156 evaluation Methods 0.000 claims description 3
- 230000006870 function Effects 0.000 claims description 3
- 230000001717 pathogenic effect Effects 0.000 claims description 3
- 230000001105 regulatory effect Effects 0.000 claims description 3
- 239000000126 substance Substances 0.000 claims description 3
- 230000001276 controlling effect Effects 0.000 claims description 2
- 238000002360 preparation method Methods 0.000 claims description 2
- 208000031128 Upper tract urothelial carcinoma Diseases 0.000 abstract description 58
- 238000003745 diagnosis Methods 0.000 abstract description 25
- 230000035945 sensitivity Effects 0.000 abstract description 8
- 238000012351 Integrated analysis Methods 0.000 abstract 1
- 206010028980 Neoplasm Diseases 0.000 description 18
- 210000002700 urine Anatomy 0.000 description 13
- 230000004083 survival effect Effects 0.000 description 12
- 201000011510 cancer Diseases 0.000 description 10
- 201000010099 disease Diseases 0.000 description 9
- 208000037265 diseases, disorders, signs and symptoms Diseases 0.000 description 9
- 238000011160 research Methods 0.000 description 8
- 238000001514 detection method Methods 0.000 description 7
- 230000011987 methylation Effects 0.000 description 7
- 238000007069 methylation reaction Methods 0.000 description 7
- 238000012544 monitoring process Methods 0.000 description 6
- 206010027476 Metastases Diseases 0.000 description 5
- 230000009401 metastasis Effects 0.000 description 5
- 230000000694 effects Effects 0.000 description 4
- 230000008569 process Effects 0.000 description 4
- 210000001519 tissue Anatomy 0.000 description 4
- 108091007854 Cdh1/Fizzy-related Proteins 0.000 description 3
- 102000038594 Cdh1/Fizzy-related Human genes 0.000 description 3
- 108091026890 Coding region Proteins 0.000 description 3
- 108020004414 DNA Proteins 0.000 description 3
- 102100028515 Heat shock-related 70 kDa protein 2 Human genes 0.000 description 3
- 101000985806 Homo sapiens Heat shock-related 70 kDa protein 2 Proteins 0.000 description 3
- -1 RASSF1A Proteins 0.000 description 3
- 210000004027 cell Anatomy 0.000 description 3
- 230000007547 defect Effects 0.000 description 3
- 230000001973 epigenetic effect Effects 0.000 description 3
- 238000002474 experimental method Methods 0.000 description 3
- 238000003384 imaging method Methods 0.000 description 3
- 230000036210 malignancy Effects 0.000 description 3
- 238000011470 radical surgery Methods 0.000 description 3
- 210000003932 urinary bladder Anatomy 0.000 description 3
- 102100028187 ATP-binding cassette sub-family C member 6 Human genes 0.000 description 2
- 108700020463 BRCA1 Proteins 0.000 description 2
- 102000036365 BRCA1 Human genes 0.000 description 2
- 101150072950 BRCA1 gene Proteins 0.000 description 2
- 101000986621 Homo sapiens ATP-binding cassette sub-family C member 6 Proteins 0.000 description 2
- 101000740180 Homo sapiens Sal-like protein 3 Proteins 0.000 description 2
- 101000659879 Homo sapiens Thrombospondin-1 Proteins 0.000 description 2
- 206010061309 Neoplasm progression Diseases 0.000 description 2
- 208000006265 Renal cell carcinoma Diseases 0.000 description 2
- 102100037191 Sal-like protein 3 Human genes 0.000 description 2
- 102100036034 Thrombospondin-1 Human genes 0.000 description 2
- 230000008901 benefit Effects 0.000 description 2
- 238000002512 chemotherapy Methods 0.000 description 2
- 229940044683 chemotherapy drug Drugs 0.000 description 2
- 238000002405 diagnostic procedure Methods 0.000 description 2
- 238000010586 diagram Methods 0.000 description 2
- 238000005516 engineering process Methods 0.000 description 2
- 230000007717 exclusion Effects 0.000 description 2
- 230000002068 genetic effect Effects 0.000 description 2
- 238000011337 individualized treatment Methods 0.000 description 2
- 210000005259 peripheral blood Anatomy 0.000 description 2
- 239000011886 peripheral blood Substances 0.000 description 2
- 238000002271 resection Methods 0.000 description 2
- 230000005751 tumor progression Effects 0.000 description 2
- 230000002485 urinary effect Effects 0.000 description 2
- 210000003741 urothelium Anatomy 0.000 description 2
- 206010002091 Anaesthesia Diseases 0.000 description 1
- 101100380241 Caenorhabditis elegans arx-2 gene Proteins 0.000 description 1
- 108010041834 Growth Differentiation Factor 15 Proteins 0.000 description 1
- 238000010824 Kaplan-Meier survival analysis Methods 0.000 description 1
- 206010060862 Prostate cancer Diseases 0.000 description 1
- 208000000236 Prostatic Neoplasms Diseases 0.000 description 1
- 108700005075 Regulator Genes Proteins 0.000 description 1
- 101710175558 Tomoregulin-2 Proteins 0.000 description 1
- 208000023915 Ureteral Neoplasms Diseases 0.000 description 1
- 206010046392 Ureteric cancer Diseases 0.000 description 1
- 108010065472 Vimentin Proteins 0.000 description 1
- 101150092805 actc1 gene Proteins 0.000 description 1
- 230000009471 action Effects 0.000 description 1
- 230000037005 anaesthesia Effects 0.000 description 1
- 210000003484 anatomy Anatomy 0.000 description 1
- 230000033115 angiogenesis Effects 0.000 description 1
- 230000006907 apoptotic process Effects 0.000 description 1
- 238000013459 approach Methods 0.000 description 1
- BBFQZRXNYIEMAW-UHFFFAOYSA-N aristolochic acid I Chemical compound C1=C([N+]([O-])=O)C2=C(C(O)=O)C=C3OCOC3=C2C2=C1C(OC)=CC=C2 BBFQZRXNYIEMAW-UHFFFAOYSA-N 0.000 description 1
- 210000004369 blood Anatomy 0.000 description 1
- 239000008280 blood Substances 0.000 description 1
- DQLATGHUWYMOKM-UHFFFAOYSA-L cisplatin Chemical compound N[Pt](N)(Cl)Cl DQLATGHUWYMOKM-UHFFFAOYSA-L 0.000 description 1
- 229960004316 cisplatin Drugs 0.000 description 1
- 238000003759 clinical diagnosis Methods 0.000 description 1
- 230000000120 cytopathologic effect Effects 0.000 description 1
- 230000034994 death Effects 0.000 description 1
- 238000013461 design Methods 0.000 description 1
- 238000000502 dialysis Methods 0.000 description 1
- 239000003814 drug Substances 0.000 description 1
- 238000012143 endoscopic resection Methods 0.000 description 1
- 230000007613 environmental effect Effects 0.000 description 1
- 230000002349 favourable effect Effects 0.000 description 1
- SDUQYLNIPVEERB-QPPQHZFASA-N gemcitabine Chemical compound O=C1N=C(N)C=CN1[C@H]1C(F)(F)[C@H](O)[C@@H](CO)O1 SDUQYLNIPVEERB-QPPQHZFASA-N 0.000 description 1
- 229960005277 gemcitabine Drugs 0.000 description 1
- 108091006104 gene-regulatory proteins Proteins 0.000 description 1
- PCHJSUWPFVWCPO-UHFFFAOYSA-N gold Chemical compound [Au] PCHJSUWPFVWCPO-UHFFFAOYSA-N 0.000 description 1
- 210000000987 immune system Anatomy 0.000 description 1
- 230000006872 improvement Effects 0.000 description 1
- 238000007901 in situ hybridization Methods 0.000 description 1
- 230000008595 infiltration Effects 0.000 description 1
- 238000001764 infiltration Methods 0.000 description 1
- 238000007689 inspection Methods 0.000 description 1
- 230000003993 interaction Effects 0.000 description 1
- 230000009545 invasion Effects 0.000 description 1
- 210000003734 kidney Anatomy 0.000 description 1
- 210000001165 lymph node Anatomy 0.000 description 1
- 230000003211 malignant effect Effects 0.000 description 1
- 208000020984 malignant renal pelvis neoplasm Diseases 0.000 description 1
- 238000010197 meta-analysis Methods 0.000 description 1
- 230000004048 modification Effects 0.000 description 1
- 238000012986 modification Methods 0.000 description 1
- 210000003205 muscle Anatomy 0.000 description 1
- 230000000683 nonmetastatic effect Effects 0.000 description 1
- 238000010827 pathological analysis Methods 0.000 description 1
- 230000002093 peripheral effect Effects 0.000 description 1
- 230000004983 pleiotropic effect Effects 0.000 description 1
- 238000010837 poor prognosis Methods 0.000 description 1
- 230000008092 positive effect Effects 0.000 description 1
- 102000004169 proteins and genes Human genes 0.000 description 1
- 230000009467 reduction Effects 0.000 description 1
- 102000037983 regulatory factors Human genes 0.000 description 1
- 230000008844 regulatory mechanism Effects 0.000 description 1
- 201000007444 renal pelvis carcinoma Diseases 0.000 description 1
- 239000000523 sample Substances 0.000 description 1
- 238000012216 screening Methods 0.000 description 1
- 239000013049 sediment Substances 0.000 description 1
- 230000011664 signaling Effects 0.000 description 1
- 239000007787 solid Substances 0.000 description 1
- 230000009897 systematic effect Effects 0.000 description 1
- 230000001225 therapeutic effect Effects 0.000 description 1
- 230000001988 toxicity Effects 0.000 description 1
- 231100000419 toxicity Toxicity 0.000 description 1
- 230000009466 transformation Effects 0.000 description 1
- 239000000439 tumor marker Substances 0.000 description 1
- 208000026517 ureter neoplasm Diseases 0.000 description 1
- 201000011476 ureteral benign neoplasm Diseases 0.000 description 1
- 208000029584 urinary system neoplasm Diseases 0.000 description 1
- 210000005048 vimentin Anatomy 0.000 description 1
Images
Classifications
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q1/00—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions
- C12Q1/68—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions involving nucleic acids
- C12Q1/6876—Nucleic acid products used in the analysis of nucleic acids, e.g. primers or probes
- C12Q1/6883—Nucleic acid products used in the analysis of nucleic acids, e.g. primers or probes for diseases caused by alterations of genetic material
- C12Q1/6886—Nucleic acid products used in the analysis of nucleic acids, e.g. primers or probes for diseases caused by alterations of genetic material for cancer
-
- G—PHYSICS
- G16—INFORMATION AND COMMUNICATION TECHNOLOGY [ICT] SPECIALLY ADAPTED FOR SPECIFIC APPLICATION FIELDS
- G16B—BIOINFORMATICS, i.e. INFORMATION AND COMMUNICATION TECHNOLOGY [ICT] SPECIALLY ADAPTED FOR GENETIC OR PROTEIN-RELATED DATA PROCESSING IN COMPUTATIONAL MOLECULAR BIOLOGY
- G16B5/00—ICT specially adapted for modelling or simulations in systems biology, e.g. gene-regulatory networks, protein interaction networks or metabolic networks
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q2600/00—Oligonucleotides characterized by their use
- C12Q2600/118—Prognosis of disease development
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q2600/00—Oligonucleotides characterized by their use
- C12Q2600/154—Methylation markers
Landscapes
- Health & Medical Sciences (AREA)
- Life Sciences & Earth Sciences (AREA)
- Chemical & Material Sciences (AREA)
- Engineering & Computer Science (AREA)
- Organic Chemistry (AREA)
- Proteomics, Peptides & Aminoacids (AREA)
- Bioinformatics & Cheminformatics (AREA)
- Physics & Mathematics (AREA)
- General Health & Medical Sciences (AREA)
- Biotechnology (AREA)
- Wood Science & Technology (AREA)
- Biophysics (AREA)
- Pathology (AREA)
- Zoology (AREA)
- Analytical Chemistry (AREA)
- Molecular Biology (AREA)
- Immunology (AREA)
- Genetics & Genomics (AREA)
- Physiology (AREA)
- Oncology (AREA)
- Hospice & Palliative Care (AREA)
- Theoretical Computer Science (AREA)
- Spectroscopy & Molecular Physics (AREA)
- Microbiology (AREA)
- Medical Informatics (AREA)
- Evolutionary Biology (AREA)
- Bioinformatics & Computational Biology (AREA)
- Biochemistry (AREA)
- General Engineering & Computer Science (AREA)
- Measuring Or Testing Involving Enzymes Or Micro-Organisms (AREA)
Abstract
The invention belongs to the technical field of biomarkers, and discloses an Upper urinary tract epithelial cancer (UTUC) diagnosis marker and application thereof, wherein the Upper urinary tract epithelial cancer diagnosis marker is DNA methylation biomarkers GDF15, TMEFF2 and VIM; the mRNA sequence NM-004864 of GDF15 is SEQ ID NO: 1; the mRNA sequence NM-001305134 of the TMEF 2 is SEQ ID NO: 2; the mRNA sequence NM-003380 of VIM is SEQ ID NO: 3. according to the invention, the data of the upper urothelial cancer is comprehensively mined from a multiomic layer, and the biomarker related to the upper urothelial cancer is searched, so that the limitation of the traditional candidate gene method can be overcome, and the result accuracy is improved; the clinical data are combined for integrated analysis, multiple biomarkers are jointly detected, a multi-index combined diagnosis model is established, and the diagnosis sensitivity and specificity are improved.
Description
Technical Field
The invention belongs to the technical field of biomarkers, and particularly relates to an upper urinary tract epithelial cancer diagnosis marker and application thereof.
Background
Epithelial cancer (UTUC) including renal pelvis cancer and ureteral cancer is a complex urinary system tumor that is commonly caused by environmental factors and genetic factors, and accounts for 5-10% of the urinary epithelial cancer (UC) in the european and american countries, with an annual incidence rate of about 2 cases/10 million residents. The incidence rate of UTUC in China is higher and accounts for about 20-30% of Transitional Cell Carcinomas (TCCs). The history of possible contact with aristolochic acid medicaments is a unique cause of UTUC patients in China. Compared with European and American countries, the proportion of women in UTUC patients in China is higher than that of men, namely about 1.3:1, and the proportion of metastasis of lymph nodes is higher because the male UTUC patients have later stages, larger tumors and higher.
Due to the special anatomical structure of the affected part of the UTUC and the complexity of the influencing factors, the UTUC is hidden in the onset and difficult to diagnose in early stage. The delay in diagnosis results in a higher proportion of late stage tumors at the time of diagnosis (56% muscle layer infiltration or local invasion), making their 5-year survival rate much lower than that of urinary bladder epithelial cancer (UCB). The research proves that: 5-year specific survival rate of the pT2/pT3 staged patient is lower than 50%, and 5-year specific survival rate of the pT4 staged patient is lower than 10%; the 5-year specific survival rate was also lower for patients graded G3-G4 compared to G1-G2. Meanwhile, the UTUC has high invasive ability and high recurrence rate, the recurrence rate can reach 16-58%, wherein the recurrence rate of the bladder is about 22-47%.
The life threat and the life quality reduction caused by UTUC can be effectively reduced by early diagnosis and early treatment, and the search of effective characteristic targets of early diagnosis and treatment gateways is a key point and a hot point of research. In recent years, clinicians and researchers have tried to find effective prediction methods from multiple dimensions, but the existing findings are not sufficient to meet the requirements of accurate diagnosis and treatment. The detection rate of the ureteral tumor by imaging examination such as CT, MRI and the like is relatively low, the positive rate of urine cast-off cytology detection is lower than 50%, urine FISH detection is also limited by specimen conditions and the tumor obstruction level, and ureteroscope/soft-scope examination is invasive and can increase the risk of bladder metastasis.
In addition, there are studies focused on epigenetic features of the UTUC. Although several susceptibility genes or biomarkers have been discovered that are associated with the development, progression and prognosis of UTUC, because of numerous molecular events accompanying the development of UTUC, the process of the related genes for exerting biological functions and the regulation mechanism are complex and are not caused by single factors or simple sets of single factors, but is formed by the interaction of various biomolecular approaches such as genes, regulatory factors, proteins and the like, and the candidate gene method is mostly adopted in the past research, meanwhile, the research result is influenced by factors such as research design, inclusion and exclusion standards, sample size, individual difference of patients (genetic heterogeneity and different environments), different disease grading stages and the like, the types of the discovered biomarkers are different, even the expression trends are opposite, therefore, the reliability of evaluating the disease state is poor, the sensitivity and specificity are not high when the method is used for diagnosis, and the method is difficult to be applied to clinical practice.
Therefore, the method finds the biomarkers with high sensitivity and strong specificity, establishes a multi-index combined diagnosis model, establishes a rapid, simple and noninvasive early diagnosis technology, and has important theoretical significance and potential transformation application value for the early diagnosis and prognosis judgment of the UTUC.
Through the above analysis, the problems and defects of the prior art are as follows:
(1) UTUC diagnosis relies primarily on imaging examinations, ureteroscopy, and urine cytology examinations. These diagnostic methods have limited sensitivity and specificity and high rates of misdiagnosis or missed diagnosis.
(2) The existing UTUC biomarker has single research level, rare types and poor repeatability, and can not completely meet the clinical diagnosis and treatment requirements only by a single-factor prediction model.
(3) Early diagnosis is difficult, and effective molecular diagnosis markers suitable for Chinese population are lacked.
(4) The prior art is lack of individual treatment means, and the prognosis of patients in middle and later stages is extremely poor.
The difficulty in solving the above problems and defects is:
(1) the gene expression between single cells has heterogeneity, which is not favorable for the tracing of cytopathic process and the research of biodiversity.
(2) How to correlate multiple sets of mathematical data with clinical data to jointly detect multiple biomarkers.
The significance of solving the problems and the defects is as follows:
(1) an efficient and accurate early diagnosis and early warning system is established, especially noninvasive detection is used for early diagnosis and disease state assessment, and the survival rate and the survival quality of patients can be greatly improved by treating antedisplacement of the gateway.
(2) The UTUC related biomarkers are searched and the pathogenic mechanism of related genes is deeply researched, so that a target is provided for individualized treatment.
(3) If a highly sensitive and highly specific biomarker combination can be found in urine or peripheral blood, the kit has great significance for UTUC prognosis monitoring and recurrence metastasis monitoring.
Disclosure of Invention
Aiming at the problems in the prior art, the invention provides a diagnosis marker for upper urothelial cancer and application thereof.
The invention is realized by the implementation of the method, and the upper urinary tract epithelial cancer diagnostic marker is DNA methylation biomarkers GDF15, TMEFF2 and VIM.
Further, the mRNA sequence NM _004864 of GDF15 is SEQ ID NO: 1.
further, the mRNA sequence NM-001305134 of TMEFF2 is SEQ ID NO: 2.
further, the mRNA sequence NM-003380 of VIM is SEQ ID NO: 3.
another object of the present invention is to provide a method for identifying a diagnostic marker for upper urothelial cancer using the diagnostic marker for upper urothelial cancer, comprising the steps of:
step one, performing joint analysis on multiple groups of chemical data by comprehensively using a bioinformatics means through a biological sample library, and discovering potential pathogenic sites/genes of UTUC and various biomarkers;
establishing a model for the early diagnosis and prognosis evaluation of the UTUC, and deeply determining the clinical application value of the early diagnosis of the related biomarkers for multiple verification;
and step three, the function of the relevant biomarkers for regulating and controlling the UTUC is determined, the relationship between the differential expression level and the occurrence and development of the UTUC is clarified, and the biological functions and pathogenic mechanisms of the differential sites/genes are clarified and potential treatment targets are discovered through multi-level verification of molecular biology, cell biology and the like.
The invention also aims to provide application of the diagnosis marker for the upper urinary tract epithelial cancer in preparation of a kit for diagnosing the upper urinary tract epithelial cancer.
By combining all the technical schemes, the invention has the advantages and positive effects that: the development of UTUC is a dynamic process, and the biomarkers and their sensitivity and specificity are different for the assessment of different stages of the disease, so it is important to use the most suitable biomarkers in different stages of the disease. Due to its own characteristics, the application of the biomarkers in the disease diagnosis and treatment process is particularly important:
(1) early diagnosis, in particular non-invasive detection for early diagnosis and disease state assessment
UTUC diagnosis relies primarily on imaging examinations, ureteroscopy, and urine cytology examinations. These diagnostic methods have limited sensitivity and specificity and high rates of misdiagnosis or missed diagnosis. And ureteroscope examination belongs to the operation, needs to carry out under anesthesia, requires to the operator highly, has subjective difference, has the wound to the patient simultaneously, causes the misery easily, also has the probability to take place the complication. Most patients with UTUC are diagnosed at a middle or advanced stage, resulting in a very poor prognosis. If the UTUC high-risk population can be effectively early diagnosed in a non-invasive or minimally invasive mode, the risk of suffering from cancer is detected as early as possible, a high-efficiency and accurate early diagnosis and early warning system is established, the treatment gateway is moved forward, and the survival rate and the survival quality of patients can be greatly improved.
(2) Searching treatment target points and providing targets for individualized treatment
The current gold standard for treatment of non-metastatic UTUC is radical nephro-uretero-vesical intramural resection. However, there is an overtreatment risk in radical surgery for patients with early or low grade malignancy UTUC. For advanced or highly malignant patients, even if radical surgery is performed, tumors still have a high probability of relapse, the treatment means for patients with side urinary tract system relapse after radical surgery is more limited, if the urinary tract system is preserved, the treatment effect is not good, and if the kidney cannot be preserved, dialysis is needed after resection. In the aspect of comprehensive treatment, chemotherapy mainly comprising gemcitabine and cisplatin is developed in many centers, and an auxiliary chemotherapy study after the UTUC operation is carried out, but the curative effect of the chemotherapy drugs is still controversial because of the toxicity of the chemotherapy drugs. The efficacy of other treatment methods (such as endoscopic resection) is not exact, and high-quality evidence of different types of patients and disease subtypes is lacking. The targeted biomarkers can modulate angiogenesis, signaling, immune system, and apoptosis. At present, the available UTUC treatment target is lacking clinically, and individualized medical treatment cannot be realized, so that the search of related biomarkers lays a solid foundation for screening the UTUC treatment target.
(3) Prognostic monitoring, UTUC recurrence metastasis monitoring
Because UTUC mostly develops from the affected side and multiple centers and seldom affects the clinical characteristics of the opposite side, after pure nephroureterectomy, the peripheral residual urothelium is easy to recur again. Even though patients with UTUC still have a greater possibility of relapse or progression after radical treatment, a monitoring means capable of accurately assessing the possibility of relapse and metastasis of patients is urgently needed to obtain a more practical treatment scheme and follow-up strategy. If a highly sensitive and highly specific biomarker combination can be found in urine or peripheral blood, the significance of the UTUC prognosis monitoring is great.
The upper urothelial cancer diagnosis marker provided by the invention is DNA methylation biomarkers GDF15, TMEFF2 and VIM which can be used for combined inspection and diagnosis of the upper urothelial cancer; the methylation biomarkers GDF15, TMEFF2 and VIM were all associated with prognosis of upper urothelial Cancer, including Tumor recurrence (Tumor recurrence), Tumor progression (Tumor progression) and Cancer-specific mortality (Cancer-specific mortality).
Drawings
In order to more clearly illustrate the technical solutions of the embodiments of the present invention, the drawings needed to be used in the embodiments of the present invention will be briefly described below, and it is obvious that the drawings described below are only some embodiments of the present invention, and it is obvious for those skilled in the art that other drawings can be obtained according to the drawings without creative efforts.
Fig. 1 is a flowchart of a method for identifying a marker for diagnosing upper urothelial cancer according to an embodiment of the present invention.
FIG. 2 is a schematic diagram showing the pleiotropic effects of the methylated GDF15, TMEFF2 and VIM genes in different prognostic results provided by the examples of the present invention.
FIG. 3 is a schematic diagram of the survival analysis of the methylated GDF15 and TMEFF2 genes provided in the examples of the present invention.
Detailed Description
In order to make the objects, technical solutions and advantages of the present invention more apparent, the present invention is further described in detail with reference to the following embodiments. It should be understood that the specific embodiments described herein are merely illustrative of the invention and are not intended to limit the invention.
In view of the problems in the prior art, the present invention provides a diagnostic marker for upper urothelial cancer and the application thereof, and the present invention is described in detail below with reference to the accompanying drawings.
The upper urinary tract epithelial cancer diagnosis markers provided by the embodiment of the invention are DNA methylation biomarkers GDF15, TMEFF2 and VIM.
As shown in fig. 1, the method for identifying a diagnosis marker for upper urothelial cancer provided by the embodiment of the present invention includes the following steps:
s101, performing joint analysis on multiple groups of chemical data by comprehensively using a bioinformatics means through a biological sample library, and discovering potential pathogenic sites/genes of UTUC and various biomarkers;
s102, establishing a model for early diagnosis and prognosis evaluation of UTUC, and deeply determining the clinical application value of early diagnosis of the related biomarkers and multiple verifications;
s103, the function of the relevant biomarkers for regulating UTUC is determined, the relationship between differential expression level and UTUC occurrence and development is clarified, the biological functions and pathogenic mechanisms of differential sites/genes are clarified through multi-level verification of molecular biology, cell biology and the like, and potential therapeutic targets are discovered.
The technical solution of the present invention is further described below with reference to specific examples.
Example 1
1. Applicants followed The requirements of The project report of first choice (PRISMA) and tumor marker prognostic research report Specification (REMARK) for systematic summary and meta analysis and searched thoroughly for evidence related to UTUC epigenetic inheritance, including DNA methylation, published in authoritative databases (PubMed, The Cochrane Library, EMBASE, Scopus) from 1/2004 to 12/2020/31. Based on inclusion exclusion criteria, a total of 11 papers related to DNA methylation were included, of which 8 reported the epigenetic status of 10 genes (GDF15, TMEFF2, VIM, RASSF1A, BRCA1, ABCC6, SALL3, HSPA2, CDH1, THBS1) in UTUC and their relationship to clinical outcome.
(see Table 1).
2. Role of methylated genes in UTUC
(ii) diagnostic action
Of the 8 studies described above, 2 reported that methylated genes were associated with the diagnosis of UTUC.
In one study, tissue samples of 57 cases of UTUC (G1/> G1 ═ 15/42) and 36 cases of control (control being normal upper urothelium from renal cell carcinoma patients); 22 cases of UTUC (G1/> G1 ═ 3/19) were compared with 20 urine samples from a control group (renal cell carcinoma: n ═ 10; prostate cancer: n ═ 7; healthy blood donors without personal or family history of cancer: n ═ 3).
Specific methylation PCR experiments found that the methylation levels of GDF15, TMEFF2, and VIM were higher in UTUC tissue and urine samples than in the control group (P ═ 0.022; P < 0.001; P < 0.001). The combined examination of tissue and urine with methylated GDF15, TMEFF2 and VIM accurately identified UTUC with sensitivity of 100% and 91%, respectively, corresponding to areas under the tissue and urine curves of 1.000 and 0.923, respectively, and specificity of 100%.
One study collected 98 samples of UTUC (G1/> G1 ═ 26/63) and 113 control groups of urine (control groups were benign urological patients, no cystoscopic or pathological diagnosis of urothelial cancer, and no history of urothelial cancer or other genitourinary malignancies).
Experiments show that the sensitivity of methylated GDF15, TMEFF2, VIM, CDH1, RASSF1A, HSPA2 and joint detection UTUC is 0.82 (0.72-0.88), the specificity is 0.68 (0.59-0.76), and the area under the curve is 0.836 (0.782-0.891); compared with urine cytology examination and fluorescence in situ hybridization technology, the detection accuracy of UTUC in urinary sediment is highest.
In summary, the following steps: the urine/tissue-DNA methylation biomarkers GDF15, TMEFF2 and VIM can be tested in combination to diagnose UTUC.
② prognostic effect
Methylation of 10 genes (GDF15, TMEFF2, VIM, RASSF1A, BRCA1, ABCC6, SALL3, HSPA2, CDH1, THBS1) was associated with relapse, progression and cancer-specific mortality of UTUC. (see Table 2)
3. Summary of the invention
(1) The DNA methylation biomarkers GDF15, TMEFF2 and VIM can be tested in combination to diagnose upper urothelial cancer.
(2) The methylation biomarkers GDF15, TMEFF2 and VIM were associated with prognosis of upper urothelial cancer, including tumor recurrence, progression and cancer-specific mortality.
Example 2
GDF15:growth differentiation factor 15
mRNA sequence:
(r) NM-0048641200 bp mRNA (see SEQ ID NO: 1)
Coding region (CDS): 33-959bp
TMEFF2:transmembrane protein with EGF like and two follistatin like domains
mRNA sequence:
(r) NM-0013051342856 bp mRNA (see SEQ ID NO: 2)
ORIGIN
Coding region (CDS): 410-1450bp
VIM:vimentin
mRNA sequence:
(r) NM-0033802154 bp mRNA (see SEQ ID NO: 3)
ORIGIN
Coding region (CDS): 432-1832 bp-
ORIGIN
The technical effects of the present invention will be described in detail with reference to experiments.
(meta) -merging the analytical results
GDF15, TMEFF2, and VIM methylation were all associated with relapse, progression, and cancer-specific mortality of UTUC. (see Table 3)
NA:notapplicable.
As shown in fig. 2, Meta pooled results show that methylated GDF15 is able to significantly reduce the risk of UTUC relapse (Z2.82; P0.005) and progression (Z4.17; P < 0.0001); methylated GDF15 and TMEFF2 were able to significantly increase the risk of cancer specific death (Z ═ 2.10; P ═ 0.04, Z ═ 4.71; P < 0.00001); methylated VIM can significantly increase the risk of UTUC (Z ═ 2.10; P ═ 0.04) progression.
(ii) survival analysis result
And (4) carrying out Kaplan-Meier survival analysis by using a UCB data set of a TCGA data base. UTUC is a urothelial malignancy, which has a more aggressive clinical phenotype than UCB. Although the UTUC data was not included in the TCGA database, the results of the analysis of the UCB dataset (434 patients) had important reference values for UTUC clinical care. As shown in FIG. 3, the Overall survival rate (OS) and Disease specific survival rate (DSS) of patients hypermethylated with GDF15 were significantly lower than those of patients hypomethylated with GDF15 (CpG cg12008047, P. 0.0052; cg12008047, P. 0.0323) among UCB patients. Patients with TMEFF2 hypermethylated UCB had significantly lower OS and DSS than patients with TMEFF2 hypomethylated (cg15003763, P ═ 0.0112; cg15003763, P ═ 0.0166).
The above description is only for the purpose of illustrating the present invention and the appended claims are not to be construed as limiting the scope of the invention, which is intended to cover all modifications, equivalents and improvements that are within the spirit and scope of the invention as defined by the appended claims.
<110> Sichuan university Hospital in western China
<120> diagnosis marker for upper urinary tract epithelial cancer and application thereof
<160> 3
<210> 1
<211> 1200
<212> DNA
<213> Artificial Sequence (Artificial Sequence)
<400>1
agtcccagct cagagccgca acctgcacag ccatgcccgg gcaagaactc aggacggtga
atggctctca gatgctcctg gtgttgctgg tgctctcgtg gctgccgcat gggggcgccc
tgtctctggc cgaggcgagc cgcgcaagtt tcccgggacc ctcagagttg cactccgaag
actccagatt ccgagagttg cggaaacgct acgaggacct gctaaccagg ctgcgggcca
accagagctg ggaagattcg aacaccgacc tcgtcccggc ccctgcagtc cggatactca
cgccagaagt gcggctggga tccggcggcc acctgcacct gcgtatctct cgggccgccc
ttcccgaggg gctccccgag gcctcccgcc ttcaccgggc tctgttccgg ctgtccccga
cggcgtcaag gtcgtgggac gtgacacgac cgctgcggcg tcagctcagc cttgcaagac
cccaggcgcc cgcgctgcac ctgcgactgt cgccgccgcc gtcgcagtcg gaccaactgc
tggcagaatc ttcgtccgca cggccccagc tggagttgca cttgcggccg caagccgcca
gggggcgccg cagagcgcgt gcgcgcaacg gggaccactg tccgctcggg cccgggcgtt
gctgccgtct gcacacggtc cgcgcgtcgc tggaagacct gggctgggcc gattgggtgc
tgtcgccacg ggaggtgcaa gtgaccatgt gcatcggcgc gtgcccgagc cagttccggg
cggcaaacat gcacgcgcag atcaagacga gcctgcaccg cctgaagccc gacacggtgc
cagcgccctg ctgcgtgccc gccagctaca atcccatggt gctcattcaa aagaccgaca
ccggggtgtc gctccagacc tatgatgact tgttagccaa agactgccac tgcatatgag
cagtcctggt ccttccactg tgcacctgcg cggaggacgc gacctcagtt gtcctgccct
gtggaatggg ctcaaggttc ctgagacacc cgattcctgc ccaaacagct gtatttatat
aagtctgtta tttattatta atttattggg gtgaccttct tggggactcg ggggctggtc
tgatggaact gtgtatttat ttaaaactct ggtgataaaa ataaagctgt ctgaactgtt
<210> 2
<211> 2856
<212> DNA
<213> Artificial Sequence (Artificial Sequence)
<400> 2
agagggatgc gggcggcaga gctcgagagg cggctgccgg gctgcggggc gccttgactc
tccctccacc ctgcctcctc gggctccact cgtctgcccc tggactcccg tctcctcctg
tcctccggct tcccagagct ccctccttat ggcagcagct tcccgcgtct ccggcgcagc
ttctcagcgg acgaccctct cgctccgggg ctgagcccag tccctggatg ttgctgaaac
tctcgagatc atgcgcgggt ttggctgctg cttccccgcc gggtgccact gccaccgccg
ccgcctctgc tgccgccgtc cgcgggatgc tcagtagccc gctgcccggc ccccgcgatc
ctgtgttcct cggaagccgt ttgctgctgc agagttgcac gaactagtca tggtgctgtg
ggagtccccg cggcagtgca gcagctggac actttgcgag ggcttttgct ggctgctgct
gctgcccgtc atgctactca tcgtagcccg cccggtgaag ctcgctgctt tccctacctc
cttaagtgac tgccaaacgc ccaccggctg gaattgctct ggttatgatg acagagaaaa
tgatctcttc ctctgtgaca ccaacacctg taaatttgat ggggaatgtt taagaattgg
agacactgtg acttgcgtct gtcagttcaa gtgcaacaat gactatgtgc ctgtgtgtgg
ctccaatggg gagagctacc agaatgagtg ttacctgcga caggctgcat gcaaacagca
gagtgagata cttgtggtgt cagaaggatc atgtgccaca gatgcaggat caggatctgg
agatggagtc catgaaggct ctggagaaac tagtcaaaag gagacatcca cctgtgatat
ttgccagttt ggtgcagaat gtgacgaaga tgccgaggat gtctggtgtg tgtgtaatat
tgactgttct caaaccaact tcaatcccct ctgcgcttct gatgggaaat cttatgataa
tgcatgccaa atcaaagaag catcgtgtca gaaacaggag aaaattgaag tcatgtcttt
gggtcgatgt caagataaca caactacaac tactaagtct gaagatgggc attatgcaag
aacagattat gcagagaatg ctaacaaatt agaagaaagt gccagagaac accacatacc
ttgtccggaa cattacaatg gcttctgcat gcatgggaag tgtgagcatt ctatcaatat
gcaggagcca tcttgcaggt gtgatgctgg ttatactgga caacactgtg aaaaaaagga
ctacagtgtt ctatacgttg ttcccggtcc tgtacgattt cagtatgtct taatcgcagc
tgtgattgga acaattcaga ttgctgtcat ctgtgtggtg gtcctctgca tcacaagggc
caaactttag gtaatagcat tggactgaga tttgtaaact ttccaacctt ccaggaaatg
ccccagaagc aacagaattc acagacagaa gcaaaataca gggcactaca gttcagacaa
tacaacaaga gcgtccacga ggttaatcta aagggagcat gtttcacagt ggctggacta
ccgagagctt ggactacaca atacagtatt atagacaaaa gaataagaca agagatctac
acatgttgcc ttgcatttgt ggtaatctac accaatgaaa acatgtacta cagctatatt
tgattatgta tggatatatt tgaaatagta tacattgtct tgatgttttt tctgtaatgt
aaataaacta tttatatcac acaatatagt tttttctttc ccatgtattt gttatatata
ataaatactc agtgatgaga aaaaattggc attcttaaat ttgcggtatc tcataactgt
aaatataatc agactagtac aatctgtaca gctaccaata tttcatgttt cttctcatct
tgagacagca cattagttcg tacaggactc agtggctagg ttttgaatga ttccaagatc
aagggaaatg atggttattg gaaaagagaa aaaataattt actttatatc gagtgaggat
aaaatatttc cgatctttga atcatctcta tttcatcaac tttcttccct ggtcttccat
tttcatccct agagcagaaa aatctctggc atataaacta aataaaagaa gaaggggagg
gaaagtgttt tataactcat aaaggagagg gaaagaaaat attggttttt attggggaag
tagcttagaa tcccccagtt aagtgcatat atctgaactt actgaacaag ttacatacta
ggtatacaca gagtggcaaa atatattcca tttaggtggg tggaattacc aggggaaaaa
tgtaataaca ccactagatg tgaaacacca aaatcgtgaa ttctcaaaag caccatacaa
tatgtatagt atatagttct ttgaaaagaa gttagaatca caaccaatac cccatgaata
gctttgtggc taatgcagca ccataatttg taatggaact aagatgatga tgacgatatt
tcatgaaaac agagagatgt tttgagcata tttatgtggt gaggtaagaa agaaaattaa
tcctatagca tctgaaagac ctcactggga agttggtatg gatttttgtt tgatttgtgc
atacaaatag gtatcacaac ttgatctgga aaaaataagc tgtgaaaatt ctcaaggaat
aagatgaaaa taaatcaatt attatcattt agctctgcaa agctttccat ggctaacaca
gtaaatttaa ataaagctct ctttgtctcc ttcaaa
<210> 3
<211> 2154
<212> DNA
<213> Artificial Sequence (Artificial Sequence)
<400> 3
gggaggccca cgtatggcgc ctctccaaag gctgcagaag tttcttgcta acaaaaagtc
cgcacattcg agcaaagaca ggctttagcg agttattaaa aacttagggg cgctcttgtc
ccccacaggg cccgaccgca cacagcaagg cgatggccca gctgtaagtt ggtagcactg
agaactagca gcgcgcgcgg agcccgctga gacttgaatc aatctggtct aacggtttcc
cctaaaccgc taggagccct caatcggcgg gacagcaggg cgcgtcctct gccactctcg
ctccgaggtc cccgcgccag agacgcagcc gcgctcccac cacccacacc caccgcgccc
tcgttcgcct cttctccggg agccagtccg cgccaccgcc gccgcccagg ccatcgccac
cctccgcagc catgtccacc aggtccgtgt cctcgtcctc ctaccgcagg atgttcggcg
gcccgggcac cgcgagccgg ccgagctcca gccggagcta cgtgactacg tccacccgca
cctacagcct gggcagcgcg ctgcgcccca gcaccagccg cagcctctac gcctcgtccc
cgggcggcgt gtatgccacg cgctcctctg ccgtgcgcct gcggagcagc gtgcccgggg
tgcggctcct gcaggactcg gtggacttct cgctggccga cgccatcaac accgagttca
agaacacccg caccaacgag aaggtggagc tgcaggagct gaatgaccgc ttcgccaact
acatcgacaa ggtgcgcttc ctggagcagc agaataagat cctgctggcc gagctcgagc
agctcaaggg ccaaggcaag tcgcgcctgg gggacctcta cgaggaggag atgcgggagc
tgcgccggca ggtggaccag ctaaccaacg acaaagcccg cgtcgaggtg gagcgcgaca
acctggccga ggacatcatg cgcctccggg agaaattgca ggaggagatg cttcagagag
aggaagccga aaacaccctg caatctttca gacaggatgt tgacaatgcg tctctggcac
gtcttgacct tgaacgcaaa gtggaatctt tgcaagaaga gattgccttt ttgaagaaac
tccacgaaga ggaaatccag gagctgcagg ctcagattca ggaacagcat gtccaaatcg
atgtggatgt ttccaagcct gacctcacgg ctgccctgcg tgacgtacgt cagcaatatg
aaagtgtggc tgccaagaac ctgcaggagg cagaagaatg gtacaaatcc aagtttgctg
acctctctga ggctgccaac cggaacaatg acgccctgcg ccaggcaaag caggagtcca
ctgagtaccg gagacaggtg cagtccctca cctgtgaagt ggatgccctt aaaggaacca
atgagtccct ggaacgccag atgcgtgaaa tggaagagaa ctttgccgtt gaagctgcta
actaccaaga cactattggc cgcctgcagg atgagattca gaatatgaag gaggaaatgg
ctcgtcacct tcgtgaatac caagacctgc tcaatgttaa gatggccctt gacattgaga
ttgccaccta caggaagctg ctggaaggcg aggagagcag gatttctctg cctcttccaa
acttttcctc cctgaacctg agggaaacta atctggattc actccctctg gttgataccc
actcaaaaag gacacttctg attaagacgg ttgaaactag agatggacag gttatcaacg
aaacttctca gcatcacgat gaccttgaat aaaaattgca cacactcagt gcagcaatat
attaccagca agaataaaaa agaaatccat atcttaaaga aacagctttc aagtgccttt
ctgcagtttt tcaggagcgc aagatagatt tggaatagga ataagctcta gttcttaaca
accgacactc ctacaagatt tagaaaaaag tttacaacat aatctagttt acagaaaaat
cttgtgctag aatacttttt aaaaggtatt ttgaatacca ttaaaactgc tttttttttt
ccagcaagta tccaaccaac ttggttctgc ttcaataaat ctttggaaaa actc
Claims (6)
1. A diagnostic marker for upper urothelial cancer, which is DNA methylation biomarkers GDF15, TMEFF2 and VIM.
2. The diagnostic marker of upper urothelial cancer of claim 1, wherein the mRNA sequence NM _004864 of GDF15 is SEQ ID NO: 1.
3. the diagnostic marker of upper urothelial cancer of claim 1, wherein the mRNA sequence NM _001305134 of TMEFF2 is SEQ ID NO: 2.
4. the diagnostic marker of upper urothelial cancer according to claim 1, wherein the mRNA sequence NM _003380 of VIM is SEQ ID NO: 3.
5. a method for identifying a diagnostic marker for upper urothelial cancer using the diagnostic marker for upper urothelial cancer according to any one of claims 1 to 4, comprising the steps of:
step one, performing joint analysis on multiple groups of chemical data by comprehensively using a bioinformatics means through a biological sample library, and discovering potential pathogenic sites/genes and various biomarkers of the upper urothelial cancer;
establishing a model for early diagnosis and prognosis evaluation of the upper urinary tract epithelial cancer, and deeply determining and multiply verifying the clinical application value of early diagnosis of related biomarkers;
and step three, the function of regulating and controlling the upper urinary tract epithelial cancer by the related biomarkers is determined, the relationship between the differential expression level and the occurrence and development of the upper urinary tract epithelial cancer is clarified, the biological function and pathogenic mechanism of the differential sites/genes are clarified through multi-level verification in molecular biology, cell biology and the like, and the potential treatment targets are discovered.
6. Use of the upper urinary tract epithelial cancer diagnostic marker according to any one of claims 1 to 4 in the preparation of a kit for diagnosing upper urinary tract epithelial cancer.
Priority Applications (1)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| CN202111431742.6A CN114107502A (en) | 2021-11-29 | 2021-11-29 | A diagnostic marker for upper urothelial carcinoma and its application |
Applications Claiming Priority (1)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| CN202111431742.6A CN114107502A (en) | 2021-11-29 | 2021-11-29 | A diagnostic marker for upper urothelial carcinoma and its application |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| CN114107502A true CN114107502A (en) | 2022-03-01 |
Family
ID=80371229
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| CN202111431742.6A Pending CN114107502A (en) | 2021-11-29 | 2021-11-29 | A diagnostic marker for upper urothelial carcinoma and its application |
Country Status (1)
| Country | Link |
|---|---|
| CN (1) | CN114107502A (en) |
Cited By (2)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN117079710A (en) * | 2023-08-18 | 2023-11-17 | 上海爱谱蒂康生物科技有限公司 | Biomarkers and their use in predicting and/or diagnosing UTUC muscle infiltration |
| CN117568404A (en) * | 2024-01-16 | 2024-02-20 | 四川大学华西医院 | Polydactyly-related gene fragments and animal model construction methods and applications |
Citations (5)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN102080123A (en) * | 2009-12-01 | 2011-06-01 | 四川大学华西医院 | Sexually transmitted disease detection kit |
| CN110198734A (en) * | 2017-01-27 | 2019-09-03 | 伊玛提克斯生物技术有限公司 | New type of peptides and peptide combinations for oophoroma and other cancer immunotherapies |
| CN111051889A (en) * | 2017-08-30 | 2020-04-21 | 东曹株式会社 | Methods and detection reagents for detecting cancer |
| CN111868260A (en) * | 2017-08-07 | 2020-10-30 | 约翰斯霍普金斯大学 | Methods and materials for the assessment and treatment of cancer |
| CN113444796A (en) * | 2021-06-29 | 2021-09-28 | 浙江医院 | Biomarkers associated with lung cancer and their use in diagnosing cancer |
-
2021
- 2021-11-29 CN CN202111431742.6A patent/CN114107502A/en active Pending
Patent Citations (5)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN102080123A (en) * | 2009-12-01 | 2011-06-01 | 四川大学华西医院 | Sexually transmitted disease detection kit |
| CN110198734A (en) * | 2017-01-27 | 2019-09-03 | 伊玛提克斯生物技术有限公司 | New type of peptides and peptide combinations for oophoroma and other cancer immunotherapies |
| CN111868260A (en) * | 2017-08-07 | 2020-10-30 | 约翰斯霍普金斯大学 | Methods and materials for the assessment and treatment of cancer |
| CN111051889A (en) * | 2017-08-30 | 2020-04-21 | 东曹株式会社 | Methods and detection reagents for detecting cancer |
| CN113444796A (en) * | 2021-06-29 | 2021-09-28 | 浙江医院 | Biomarkers associated with lung cancer and their use in diagnosing cancer |
Non-Patent Citations (7)
| Title |
|---|
| CRUICKSHANK T等: "Homo sapiens growth differentiation factor 15 (GDF15), mRNA" * |
| DING Y,等: "Homo sapiens vimentin (VIM), mRNA" * |
| ELAHI A等: "Homo sapiens transmembrane protein with EGF like and two follistatin like domains 2 (TMEFF2), transcript variant 2, mRNA" * |
| SARA MONTEIRO-REIS,等: "Accurate detection of upper tract urothelial carcinoma in tissue and urine by means of quantitative GDF15, TMEFF2 and VIM promoter methylation" * |
| YIFEI LIN,等: "DNA Methylation Architecture Provides Insight into the Pathogenesis of Upper Tract Urothelial Carcinoma: A Systematic Review and Meta-Analysis" * |
| 张德莲,等: "基于生物信息学分析筛选肾上腺皮质癌诊断及预后相关生物标记物" * |
| 熊萱;李一;喻冬柯;张远;: "利用TCGA公共数据库挖掘乳腺癌预后相关长链非编码RNA生物标志物" * |
Cited By (4)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN117079710A (en) * | 2023-08-18 | 2023-11-17 | 上海爱谱蒂康生物科技有限公司 | Biomarkers and their use in predicting and/or diagnosing UTUC muscle infiltration |
| CN117079710B (en) * | 2023-08-18 | 2024-05-31 | 上海爱谱蒂康生物科技有限公司 | Biomarkers and their use in predicting and/or diagnosing UTUC muscle infiltrates |
| CN117568404A (en) * | 2024-01-16 | 2024-02-20 | 四川大学华西医院 | Polydactyly-related gene fragments and animal model construction methods and applications |
| CN117568404B (en) * | 2024-01-16 | 2024-03-15 | 四川大学华西医院 | Multi-finger (toe) related gene segment, animal model construction method and application |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| Laudadio et al. | Fluorescence in situ hybridization for detecting transitional cell carcinoma: implications for clinical practice | |
| Bubendorf et al. | Multiprobe FISH for enhanced detection of bladder cancer in voided urine specimens and bladder washings | |
| US20030190602A1 (en) | Cell-based detection and differentiation of disease states | |
| Okada et al. | Digital PCR-based plasma cell-free DNA mutation analysis for early-stage pancreatic tumor diagnosis and surveillance | |
| Herrero et al. | Evaluation of nuclear matrix protein 22 as a tumour marker in the detection of transitional cell carcinoma of the bladder | |
| CA2534633A1 (en) | Biomarker panel for colorectal cancer | |
| Chen et al. | Is there a role for FISH in the management and surveillance of patients with upper tract transitional-cell carcinoma? | |
| Dunn et al. | Molecular markers for early detection | |
| JP2024105502A (en) | DNA methylation markers for non-invasive detection of cancer and their uses - Patents.com | |
| US20120135877A1 (en) | DNA Methylation Markers For Prostate Cancer Field Defect | |
| CN115851951A (en) | Construction of early liver cancer detection model containing multiple groups of chemical marker compositions and kit | |
| Tong et al. | A potential novel biomarker in predicting lymph node metastasis of gastric signet ring cell carcinoma: a derived monocyte to lymphocyte ratio | |
| CN117165688A (en) | Marker for urothelial cancer and application thereof | |
| Zhu et al. | Risk factor analysis and construction of prediction models of gallbladder carcinoma in patients with gallstones | |
| CN114107502A (en) | A diagnostic marker for upper urothelial carcinoma and its application | |
| Jiang et al. | Combined genetic analysis of sputum and computed tomography for noninvasive diagnosis of non-small-cell lung cancer | |
| US20210404018A1 (en) | Unbiased dna methylation markers define an extensive field defect in histologically normal prostate tissues associated with prostate cancer: new biomarkers for men with prostate cancer | |
| Murphy et al. | Patented prostate cancer biomarkers | |
| Kipp et al. | Assessing the value of reflex fluorescence in situ hybridization testing in the diagnosis of bladder cancer when routine urine cytological examination is equivocal | |
| Dai et al. | Combining methylated RNF180 and SFRP2 plasma biomarkers for noninvasive diagnosis of gastric cancer | |
| Wu et al. | A novel dual-target Septin9 methylation assay for improved detection of early-stage colorectal cancer and high-grade intraepithelial neoplasia | |
| CN116144782A (en) | A combination marker for lung cancer detection and its application | |
| JP7463357B2 (en) | Preoperative risk stratification based on PDE4D7 and DHX9 expression | |
| CN117106918A (en) | Method for differential diagnosis of benign lung nodules and malignant tumors by gene methylation and kit thereof | |
| Wei et al. | A urine-based liquid biopsy for detection of upper tract urothelial carcinoma: a self-matched study |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| PB01 | Publication | ||
| PB01 | Publication | ||
| SE01 | Entry into force of request for substantive examination | ||
| SE01 | Entry into force of request for substantive examination | ||
| RJ01 | Rejection of invention patent application after publication |
Application publication date: 20220301 |
|
| RJ01 | Rejection of invention patent application after publication |