CN113943805A - Gene mutation detection primer set and its use - Google Patents
Gene mutation detection primer set and its use Download PDFInfo
- Publication number
- CN113943805A CN113943805A CN202111286587.3A CN202111286587A CN113943805A CN 113943805 A CN113943805 A CN 113943805A CN 202111286587 A CN202111286587 A CN 202111286587A CN 113943805 A CN113943805 A CN 113943805A
- Authority
- CN
- China
- Prior art keywords
- seq
- dna
- artificial sequence
- breast cancer
- odd numbers
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Pending
Links
Classifications
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q1/00—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions
- C12Q1/68—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions involving nucleic acids
- C12Q1/6876—Nucleic acid products used in the analysis of nucleic acids, e.g. primers or probes
- C12Q1/6883—Nucleic acid products used in the analysis of nucleic acids, e.g. primers or probes for diseases caused by alterations of genetic material
- C12Q1/6886—Nucleic acid products used in the analysis of nucleic acids, e.g. primers or probes for diseases caused by alterations of genetic material for cancer
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K45/00—Medicinal preparations containing active ingredients not provided for in groups A61K31/00 - A61K41/00
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P35/00—Antineoplastic agents
-
- G—PHYSICS
- G16—INFORMATION AND COMMUNICATION TECHNOLOGY [ICT] SPECIALLY ADAPTED FOR SPECIFIC APPLICATION FIELDS
- G16B—BIOINFORMATICS, i.e. INFORMATION AND COMMUNICATION TECHNOLOGY [ICT] SPECIALLY ADAPTED FOR GENETIC OR PROTEIN-RELATED DATA PROCESSING IN COMPUTATIONAL MOLECULAR BIOLOGY
- G16B20/00—ICT specially adapted for functional genomics or proteomics, e.g. genotype-phenotype associations
- G16B20/50—Mutagenesis
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q2600/00—Oligonucleotides characterized by their use
- C12Q2600/136—Screening for pharmacological compounds
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q2600/00—Oligonucleotides characterized by their use
- C12Q2600/156—Polymorphic or mutational markers
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q2600/00—Oligonucleotides characterized by their use
- C12Q2600/16—Primer sets for multiplex assays
Landscapes
- Health & Medical Sciences (AREA)
- Life Sciences & Earth Sciences (AREA)
- Chemical & Material Sciences (AREA)
- Proteomics, Peptides & Aminoacids (AREA)
- Engineering & Computer Science (AREA)
- Organic Chemistry (AREA)
- General Health & Medical Sciences (AREA)
- Analytical Chemistry (AREA)
- Genetics & Genomics (AREA)
- Physics & Mathematics (AREA)
- Bioinformatics & Cheminformatics (AREA)
- Zoology (AREA)
- Biotechnology (AREA)
- Wood Science & Technology (AREA)
- Biophysics (AREA)
- Pathology (AREA)
- Immunology (AREA)
- Veterinary Medicine (AREA)
- Molecular Biology (AREA)
- Public Health (AREA)
- Medicinal Chemistry (AREA)
- Animal Behavior & Ethology (AREA)
- Pharmacology & Pharmacy (AREA)
- Chemical Kinetics & Catalysis (AREA)
- Nuclear Medicine, Radiotherapy & Molecular Imaging (AREA)
- General Chemical & Material Sciences (AREA)
- Theoretical Computer Science (AREA)
- Epidemiology (AREA)
- Hospice & Palliative Care (AREA)
- Spectroscopy & Molecular Physics (AREA)
- Oncology (AREA)
- Medical Informatics (AREA)
- Microbiology (AREA)
- Evolutionary Biology (AREA)
- Bioinformatics & Computational Biology (AREA)
- Biochemistry (AREA)
- General Engineering & Computer Science (AREA)
- Measuring Or Testing Involving Enzymes Or Micro-Organisms (AREA)
Abstract
The invention relates to a gene mutation detection primer group and application thereof. Specifically, the present invention provides a primer composition comprising: (1) 1-260, or (2) a primer pair consisting of SEQ ID NO: n and n +1, wherein n is any one or more or all odd numbers of 1-259. The primer group is used for detecting gene mutation, so that the library construction success rate and the gene average coverage multiple can be obviously improved.
Description
Technical Field
The invention relates to the field of gene mutation detection, in particular to a gene mutation detection primer group and application thereof.
Background
Breast cancer is one of the most common malignant tumors of women in the world, accounting for 23 percent of the total cancer morbidity of women and accounting for 14 percent of the cancer mortality of women. Recurrence of breast cancer metastasis is an important cause of death for patients, with median survival time after metastasis being less than 2 years. In China, the incidence rate of breast cancer is on the trend of rising year by year in recent years.
The causative factors of breast cancer are not fully understood to date. Family history of breast cancer, history of benign breast disease, age of menstrual onset later, higher mental stress, higher body mass index, family history of tumors, and the like are considered as important causes of breast cancer. The etiology of breast cancer also has more breakthroughs from the genetic point of view, and a series of breast cancer susceptibility gene mutations are proved to be related to the breast cancer. Existing studies have clearly demonstrated that tumor cell gene mutations are closely associated with the development and metastasis of breast cancer. Tumor cells carrying different genetic mutations also have different sensitivities to various drugs. For example, for tumors with PIK3CA gene mutation, Alpelisib, Buparlisb + Fulvestrant, Copanlisb, Everolimus, GDC-0077, Serabelisib, Taselisib or Taselisib + Fulvestrant are used for better prognosis of curative effect; the tumor with AKT1 gene mutation has better curative effect prognosis by using AZD5363 or Everolimus; and the tumor with ERBB2 gene mutation is more suitable for Apatinib, Lapatinib or Neratinib. Different drugs and treatment regimens are directed to individuals with different mutations, both in terms of efficacy and in terms of prognostic survival.
At present, the tumor mutant gene detection has a plurality of primer sets and different schemes. More genes detected means higher sample quality requirements, lower second generation sequencing pooling power and lower mean gene coverage. The first two problems may directly lead to the result of a detection failure. The reduction of the mean fold coverage of the gene means that the detection of the critical mutated DNA may be missed and the opportunity for accurate treatment may be missed. There is still a need in the art for a method for detecting breast cancer mutations with high library formation power and high gene mean fold coverage.
Disclosure of Invention
The invention realizes the fun detection of the mutation of the three genes through the specially designed primer composition, has high library construction power and high gene average coverage factor, and can be used for breast cancer classification and treatment scheme selection.
In a first aspect, the present invention provides a primer composition comprising:
(1) any one or more of SEQ ID NO 1-SEQ ID NO 260, or
(2) N and n +1, where n is any one or more or all odd numbers from 1 to 259.
In one or more embodiments, the primer composition comprises one, two, or all three groups selected from:
(1) any one or more of SEQ ID NO 1-26, or a primer pair consisting of SEQ ID NO n and n +1, wherein n is any one or more or all odd numbers of 1-25, and/or any one or more of SEQ ID NO 27-50, or a primer pair consisting of SEQ ID NO n and n +1, wherein n is any one or more or all odd numbers of 27-49,
(2) any one or more of SEQ ID NO 51-106, or a primer pair consisting of SEQ ID NO n and n +1, wherein n is any one or more or all odd numbers of 51-105, and/or any one or more of SEQ ID NO 107-160, or a primer pair consisting of SEQ ID NO n and n +1, wherein n is any one or more or all odd numbers of 51-105,
(3) any one or more of SEQ ID NO 161-210, or a primer pair consisting of SEQ ID NO n and n +1, wherein n is any one or more or all odd numbers of 161-209, and/or any one or more of SEQ ID NO 211-260, or a primer pair consisting of SEQ ID NO n and n +1, wherein n is any one or more or all odd numbers of 211-259.
In one or more embodiments, the primer composition comprises one, two, or all three groups selected from:
(1) any one or more of SEQ ID NOs 1-50, or a primer pair consisting of SEQ ID NOs n and n +1, wherein n is any one or more or all odd numbers from 1 to 49, (2) any one or more of SEQ ID NOs 51-160, or a primer pair consisting of SEQ ID NOs n and n +1, wherein n is any one or more or all odd numbers from 51 to 159,
(3) any one or more of SEQ ID NO 161-260, or a primer pair consisting of SEQ ID NO n and n +1, wherein n is any one or more or all odd numbers of 161-259.
The present invention also provides a method for detecting gene mutation, comprising the step of amplifying a genome of a subject using the primer composition according to any one of the embodiments of the first aspect of the present invention.
In one or more embodiments, the method is a non-therapeutic or diagnostic method.
In one or more embodiments, the subject is a breast cancer patient.
In one or more embodiments, the method further comprises the step of determining the sequence of the amplification product.
In one or more embodiments, the method further comprises the step of determining the sequence of the portion of the gene that it covers based on the sequence of the amplification product.
In one or more embodiments, the method further comprises the step of comparing the sequence of the amplification product or the portion of the gene covered thereby to the corresponding wild-type sequence to determine the genetic mutation.
The invention also includes the use of a primer composition according to any one of the embodiments of the first aspect of the invention in the preparation of a kit for detecting a gene mutation.
In one or more embodiments, the gene comprises one or two or three selected from the group consisting of: AKT1, ERBB2, PIK3 CA.
In one or more embodiments, the gene comprises AKT1, and the primer composition comprises: 1-50 or a primer pair consisting of SEQ ID NO n and n +1, wherein n is any one or more or all odd numbers from 1 to 49.
In one or more embodiments, the gene comprises ERBB2, and the primer composition comprises: any one or more of SEQ ID NO 51-160, or a primer pair consisting of SEQ ID NO n and n +1, wherein n is any one or more or all odd numbers of 51-159.
In one or more embodiments, the gene comprises PIK3CA, and the primer composition comprises: any one or more of SEQ ID NO 161-260, or a primer pair consisting of SEQ ID NO n and n +1, wherein n is any one or more or all odd numbers of 161-259.
In one or more embodiments, the mutation of AKT1 includes a mutation at position E17.
In one or more embodiments, the mutation of ERBB2 comprises a mutation at the S310 and/or D769 site.
In one or more embodiments, the mutation of PIK3CA includes a mutation at one or more positions selected from E542, P366, H1047.
In one or more embodiments, the mutation comprises one or more of the following: E17K of AKT1, E17K of AKT1, S310F of ERBB2, D769Y of ERBB2, H1047L of PIK3CA, E542K of PIK3CA, P366R of PIK3 CA.
Use of a primer composition according to any one of the embodiments of the first aspect of the invention in the preparation of a breast cancer identification kit or a kit for selecting a breast cancer treatment regimen.
In one or more embodiments, the primer composition is used to detect mutations in a gene comprising one or two or three selected from the group consisting of: AKT1, ERBB2, PIK3 CA.
In one or more embodiments, the genetic mutation comprises a mutation to AKT1, and the primer composition comprises: 1-50 or a primer pair consisting of SEQ ID NO n and n +1, wherein n is any one or more or all odd numbers from 1 to 49. In certain embodiments, the breast cancer is AKT1 mutant breast cancer, and the treatment regimen comprises administering an agent that treats AKT1 mutant breast cancer, such as AZD5363 and/or Everolimus.
In one or more embodiments, the gene comprises ERBB2, and the primer composition comprises: any one or more of SEQ ID NO 51-160, or a primer pair consisting of SEQ ID NO n and n +1, wherein n is any one or more or all odd numbers of 51-159. In certain embodiments, the breast cancer is an ERBB2 mutant breast cancer, and the treatment regimen comprises administering an agent that treats ERBB2 mutant breast cancer, such as Apatinib, Lapatinib, and/or Neratinib.
In one or more embodiments, the gene comprises PIK3CA, and the primer composition comprises: any one or more of SEQ ID NO 161-260, or a primer pair consisting of SEQ ID NO n and n +1, wherein n is any one or more or all odd numbers of 161-259. In certain embodiments, the breast cancer is PIK3CA mutant breast cancer, and the treatment regimen comprises administering an agent that treats PIK3CA mutant breast cancer, such as Alpelisib and/or Fulvestrant.
The invention also provides the use of a primer composition according to any one of the embodiments of the first aspect of the invention in the preparation of a kit for the treatment of breast cancer.
In one or more embodiments, the primer composition comprises: (1) any one or more of SEQ ID NOs 1-26, or a primer pair consisting of SEQ ID NOs n and n +1, wherein n is any one or more or all odd numbers from 1 to 25, and/or (2) any one or more of SEQ ID NOs 27-50, or a primer pair consisting of SEQ ID NOs n and n +1, wherein n is any one or more or all odd numbers from 27 to 49. Preferably, the kit further comprises a medicament for treating AKT1 mutant breast cancer, such as AZD5363 and/or Everolimus.
In one or more embodiments, the primer composition comprises: (3) any one or more of SEQ ID NOS 51-106, or a primer pair consisting of SEQ ID NOS n and n +1, wherein n is any one or more or all of the odd numbers of 51-105, and/or (4) any one or more of SEQ ID NOS 107-160, or a primer pair consisting of SEQ ID NOS n and n +1, wherein n is any one or more or all of the odd numbers of 51-105. Preferably, the kit further comprises a medicament for treating ERBB2 mutant breast cancer, such as one, two or all three of Apatinib, Lapatinib and Neratinib.
In one or more embodiments, the primer composition comprises: (5) any one or more of SEQ ID NO:161-210, or a primer pair consisting of SEQ ID NO: n and n +1, wherein n is any one or more or all odd numbers of 161-209, and/or (6) any one or more of SEQ ID NO:211-260, or a primer pair consisting of SEQ ID NO: n and n +1, wherein n is any one or more or all odd numbers of 211-250. Preferably, the kit further comprises a medicament for treating PIK3CA mutant breast cancer, such as Alpelisib and/or Fulvestrant.
The invention also provides a kit comprising a primer composition according to any one of the embodiments of the first aspect of the invention and optionally a medicament.
In one or more embodiments, the primer composition comprises: (1) any one or more of SEQ ID NOs 1-26, or a primer pair consisting of SEQ ID NOs n and n +1, wherein n is any one or more or all odd numbers from 1 to 25, and/or (2) any one or more of SEQ ID NOs 27-50, or a primer pair consisting of SEQ ID NOs n and n +1, wherein n is any one or more or all odd numbers from 27 to 49. Preferably, the medicament comprises a medicament for treating AKT1 mutant breast cancer, such as AZD5363 and/or Everolimus.
In one or more embodiments, the primer composition comprises: (3) any one or more of SEQ ID NOS 51-106, or a primer pair consisting of SEQ ID NOS n and n +1, wherein n is any one or more or all of the odd numbers of 51-105, and/or (4) any one or more of SEQ ID NOS 107-160, or a primer pair consisting of SEQ ID NOS n and n +1, wherein n is any one or more or all of the odd numbers of 51-105. Preferably, the medicament comprises a medicament for treating ERBB2 mutant breast cancer, such as one, two or all three of Apatinib, Lapatinib and Neratinib.
In one or more embodiments, the primer composition comprises: (5) any one or more of SEQ ID NO:161-210, or a primer pair consisting of SEQ ID NO: n and n +1, wherein n is any one or more or all odd numbers of 161-209, and/or (6) any one or more of SEQ ID NO:211-260, or a primer pair consisting of SEQ ID NO: n and n +1, wherein n is any one or more or all odd numbers of 211-250. Preferably, the medicament comprises a medicament for treating PIK3CA mutant breast cancer, such as Alpelisib and/or Fulvestrant.
The invention also provides an apparatus for providing a treatment plan, the apparatus comprising a memory, a processor and a computer program stored on the memory and executable on the processor, wherein the processor when executing the program performs the steps of:
(1) identifying whether a breast cancer tumor cell of a subject has a mutation in one, two or more genes selected from AKT1, ERBB2, PIK3CA using a primer composition according to any of the embodiments of the first aspect of the invention,
(2) providing a treatment regimen according to the results of (1).
In one or more embodiments, the gene includes AKT1 and the treatment regimen includes determining whether to administer a drug to treat AKT1 mutant breast cancer, such as AZD5363 and/or Everolimus. AZD5363 and/or Everolimus are administered if AKT1 has a mutation. In one or more embodiments, the primer composition comprises: (a) 1-26 or a primer pair consisting of SEQ ID NOs n and n +1, wherein n is any one or more or all odd numbers from 1 to 25, and/or (b) any one or more of SEQ ID NOs 27 to 50 or a primer pair consisting of SEQ ID NOs n and n +1, wherein n is any one or more or all odd numbers from 27 to 49.
In one or more embodiments, the gene comprises ERBB2 and the treatment regimen comprises determining whether to administer an agent that treats ERBB2 mutant breast cancer, such as one or more of Apatinib, Lapatinib, Neratinib. AZD5363 and/or Everolimus are administered if ERBB2 has a mutation. In one or more embodiments, the primer composition comprises: (c) any one or more of SEQ ID NOS 51-106, or a primer pair consisting of SEQ ID NOS n and n +1, wherein n is any one or more or all of the odd numbers of 51-105, and/or (d) any one or more of SEQ ID NOS 107-160, or a primer pair consisting of SEQ ID NOS n and n +1, wherein n is any one or more or all of the odd numbers of 51-105.
In one or more embodiments, the gene comprises PIK3CA and the treatment regimen comprises determining whether to administer a drug, such as Alpelisib and/or Fulvestrant, to treat PIK3CA mutant breast cancer. If PIK3CA has a mutation, Alpelisib and/or Fulvestrant is administered. In one or more embodiments, the primer composition comprises: (e) any one or more of SEQ ID NO:161-210, or a primer pair consisting of SEQ ID NO: n and n +1, wherein n is any one or more or all odd numbers of 161-209, and/or (f) any one or more of SEQ ID NO:211-260, or a primer pair consisting of SEQ ID NO: n and n +1, wherein n is any one or more or all odd numbers of 211-250.
Detailed Description
The inventor finds that, according to actual work, the tumor cell mutations (somatic mutation and non-germ line mutation) which can provide important information for treatment scheme selection in Chinese human breast cancer are concentrated on the genes PIK3CA, AKT1 and ERBBB2, and the mutations of the three genes account for 87 percent of all targeted drug mutation sites. Only the tumor cell mutation of the three genes needs to be detected, and almost 90% of the accurate medical requirements can be met.
The inventors have repeatedly designed and developed a primer set for detecting the mutation of the three genes. When the detection object is an FFPE sample, compared with about 50% of library construction success rate of 500 gene primer groups, the success rate of using the primer to construct the library is 100%, and the improvement effect is very obvious. Particularly, on the aspect of gene coverage, under the condition of the same data volume, the average coverage multiple of the primer group is 170 times of the average coverage multiple of a 500 genome, so that the capability of detecting mutant cells which do not have the advantage of simple quantity in tumors is greatly improved, and more opportunities for precise medical treatment are strived for breast cancer patients.
The invention provides a primer composition, which comprises: (1) 1 to 260, or (2) a primer pair consisting of SEQ ID NO n and n +1, wherein n is any one or more or all odd numbers of 1 to 259,
wherein, the primer composition for detecting AKT1 comprises: (1) any one or more of SEQ ID NOs 1-26, or a primer pair consisting of SEQ ID NOs n and n +1, wherein n is any one or more or all odd numbers from 1 to 25, and/or (2) any one or more of SEQ ID NOs 27-50, or a primer pair consisting of SEQ ID NOs n and n +1, wherein n is any one or more or all odd numbers from 27 to 49. The primer composition for detecting ERBBB2 comprises: (1) any one or more of SEQ ID NO 51-106, or a primer pair consisting of SEQ ID NO n and n +1, wherein n is any one or more or all odd numbers of 51-105, and/or any one or more of SEQ ID NO 107-160, or a primer pair consisting of SEQ ID NO n and n +1, wherein n is any one or more or all odd numbers of 51-105. The primer composition for detecting PIK3CA comprises: (1) any one or more of SEQ ID NO:161-210, or a primer pair consisting of SEQ ID NO: n and n +1, wherein n is any one or more or all odd numbers of 161-209, and/or (2) any one or more of SEQ ID NO:211-260, or a primer pair consisting of SEQ ID NO: n and n +1, wherein n is any one or more or all odd numbers of 211-259.
Methods for detecting gene mutations can be obtained by amplifying (e.g., PCR) a subject genome using the primer compositions described herein. The subject may be a breast cancer patient or a non-breast cancer patient. Experimental methods for obtaining gene mutation detection results from amplification products are well known in the art, for example, by second generation sequencing. Thus, the method further comprises the step of determining the sequence of the amplification product, and optionally further comprises the step of determining the sequence of the portion of the gene covered by the amplification product based on the sequence of the amplification product. Further, the method may further comprise the step of comparing the sequence of the amplification product or the sequence of the gene portion covered thereby with the corresponding wild type sequence to thereby determine the gene mutation.
The primer composition described herein can be used for preparing a kit for detecting gene mutation. The gene comprises one or two or three selected from the group consisting of: AKT1, ERBB2, PIK3 CA. Wherein, when the gene to be detected comprises AKT1, the primer composition comprises: 1-50 or a primer pair consisting of SEQ ID NO n and n +1, wherein n is any one or more or all odd numbers from 1 to 49. When the gene to be tested comprises ERBB2, the primer composition comprises: any one or more of SEQ ID NO 51-160, or a primer pair consisting of SEQ ID NO n and n +1, wherein n is any one or more or all odd numbers of 51-159. When the gene to be tested comprises PIK3CA, the primer composition comprises: any one or more of SEQ ID NO 161-260, or a primer pair consisting of SEQ ID NO n and n +1, wherein n is any one or more or all odd numbers of 161-259.
Based on the results of the detected gene mutations, breast cancer can be identified and a treatment regimen for breast cancer can be selected with specificity. Herein, breast cancers are classified into at least AKT1 mutant breast cancers, ERBB2 mutant breast cancers, and PIK3CA mutant breast cancers, which respectively represent AKT1, ERBB2, and PIK3CA mutant breast cancers. The above mentioned types of mutations do not usually occur simultaneously. Mutations in AKT1 include mutations at the E17 site. Mutations in ERBB2 include mutations at the S310 and/or D769 sites. Mutations in PIK3CA include mutations at one or more positions selected from E542, P366, H1047. In exemplary embodiments, the mutation at the above-mentioned site specifically includes one or more selected from the group consisting of: E17K of AKT1, E17K of AKT1, S310F of ERBB2, D769Y of ERBB2, H1047L of PIK3CA, E542K of PIK3CA, P366R of PIK3 CA.
For example, agents for treating AKT1 mutant breast cancer include, but are not limited to, AZD5363 and/or Everolimus, agents for treating ERBB2 mutant breast cancer include, but are not limited to, Apatinib, Lapatinib, and/or Neratinib, and agents for treating PIK3CA mutant breast cancer include, but are not limited to, Alpelisib and/or Fulvestrant.
The primer compositions described herein can also be prepared as a therapeutic kit (kit) with a breast cancer treatment drug, comprising: the primer composition for detecting AKT1 and a medicament for treating AKT1 mutant breast cancer, and/or the primer composition for detecting ERBB2 and a medicament for treating ERBB2 mutant breast cancer, and/or the primer composition for detecting PIK3CA and a medicament for treating PIK3CA mutant breast cancer. In particular embodiments, the kit comprises: (1) any one or more of SEQ ID NOs 1-26, or a primer pair consisting of SEQ ID NOs n and n +1, wherein n is any one or more or all odd numbered members of 1-25, (2) any one or more of SEQ ID NOs 27-50, or a primer pair consisting of SEQ ID NOs n and n +1, wherein n is any one or more or all odd numbered members of 27-49, and (3) a medicament for treating AKT1 mutant breast cancer. Alternatively or additionally, the kit comprises: (1) any one or more of SEQ ID NOS 51-106, or a primer pair consisting of SEQ ID NOS n and n +1, wherein n is any one or more or all odd numbers of 51-105, (2) any one or more of SEQ ID NOS 107-160, or a primer pair consisting of SEQ ID NOS n and n +1, wherein n is any one or more or all odd numbers of 51-105, and (3) a drug for treating ERBB2 mutant breast cancer. Alternatively or additionally, the kit comprises: (1) any one or more of SEQ ID NO:161-210, or a primer pair consisting of SEQ ID NO: n and n +1, wherein n is any one or more or all odd numbers of 161-209, (2) any one or more of SEQ ID NO:211-260, or a primer pair consisting of SEQ ID NO: n and n +1, wherein n is any one or more or all odd numbers of 211-250, and (3) a drug for treating PIK3CA mutant breast cancer.
Also disclosed is a computer readable storage medium storing a computer program, the computer program stored on the storage medium when executed performs the method of identifying breast cancer and selecting a treatment for breast cancer as described herein. The steps of a method or algorithm described in connection with the embodiments disclosed herein may be embodied directly in hardware, in a software module executed by a processor, or in a combination of the two. If implemented in software as a computer program product, the functions may be stored on or transmitted over as one or more instructions or code on a computer-readable medium. Computer-readable media includes both computer storage media and communication media including any medium that facilitates transfer of a computer program from one place to another. A storage media may be any available media that can be accessed by a computer. By way of example, and not limitation, such computer-readable media can comprise RAM, ROM, EEPROM, CD-ROM or other optical disk storage, magnetic disk storage or other magnetic storage devices, or any other medium that can be used to carry or store desired program code in the form of instructions or data structures and that can be accessed by a computer. An exemplary storage medium is coupled to the processor such the processor can read information from, and write information to, the storage medium. In the alternative, the storage medium may be integral to the processor. The processor and the storage medium may reside in an ASIC. The ASIC may reside in a user terminal. In the alternative, the processor and the storage medium may reside as discrete components in a user terminal.
Specifically, the apparatus for providing a treatment plan of the present invention comprises a memory, a processor and a computer program stored on the memory and executable on the processor, wherein the processor executes the program to realize the following steps: (1) identifying whether a breast cancer tumor cell of a subject has a mutation in one, two or more genes selected from AKT1, ERBB2, PIK3CA using a primer composition according to any one of the embodiments of the first aspect of the invention, (2) providing a treatment regimen based on the results of (1). The gene comprises AKT1, the treatment regimen comprising determining whether to administer AZD5363 and/or Everolimus, wherein AZD5363 and/or Everolimus is administered if AKT1 has a mutation; the gene includes ERBB2, and the treatment regimen includes determining whether Apatinib, Lapatinib, and/or Neratinib is administered, wherein AZD5363 and/or Everolimus is administered if ERBB2 has a mutation. The gene comprises PIK3CA, and the treatment regimen comprises determining whether to administer Alpelisib and/or Fulvestrant, wherein Alpelisib and/or Fulvestrant is administered if PIK3CA has a mutation.
Unless defined otherwise, all technical and scientific terms used herein have the same meaning as commonly understood by one of ordinary skill in the art to which this invention belongs. Although any methods and materials similar or equivalent to those described herein can be used in the practice or testing of the present invention, the preferred methods and materials are now described. All publications and patents specifically mentioned herein are incorporated herein by reference in their entirety for all purposes including describing and disclosing the chemicals, equipment, statistical analyses and methods reported in the publications that could be used in connection with the invention. All references cited in this specification are to be considered as indicative of the level of skill in the art. Nothing herein is to be construed as an admission that the invention is not entitled to antedate such disclosure by virtue of prior invention.
The invention will be further illustrated with reference to the following specific examples. It should be understood that these examples are for illustrative purposes only and are not intended to limit the scope of the present invention. The experimental procedures, for which specific conditions are not noted in the following examples, are generally performed according to conventional conditions such as those described in J. SammBruk et al, molecular cloning protocols, third edition, scientific Press, 2002, or according to the manufacturer's recommendations.
Examples
Example 1
Patient information
Breast cancer patient 001, clinically diagnosed luminel type B (HER2 negative);
breast cancer patient 002, clinically diagnosed luminel type B (HER2 positive);
breast cancer patient 003, clinically diagnosed as triple negative breast cancer;
breast cancer patient 004, clinically diagnosed luminel type B (HER2 negative);
a breast cancer patient 005 clinically diagnosed with triple negative breast cancer;
breast cancer patient 006, clinically diagnosed lumineal type B (HER2 negative).
Breast cancer patient 007 with clinical diagnosis of triple negative breast cancer.
Sample acquisition and processing
After paraffin is dissolved by using dewaxing liquid, cell walls are damaged by proteinase K, and DNA is enriched and fragmented. Samples were processed according to kit instructions.
Main reagents and devices:
GeneRead DNA FFPE Kit(QINGEN);
breaking the tube (Covaris microTUBE 130, Gene Co.);
High sensitivity cartridge kit(N1)(Bioptic);
Agencourt AMPure XPkit(Beckman Coulter)。
second generation sequencing
The detection experiment was performed by dividing the primers into two groups (Pool _1 and Pool _2, as shown in tables 1-3) for PCR, followed by pooling for second-generation sequencing and data analysis as follows:
PCR products were ligated to DNA linkers and specific index fragments, and all pools were mixed in equal amounts for high throughput sequencing. Sequencing was performed on a NovaSeq 6000sequence (Illumina, Inc, San Diego, CA) single channel, as per the manufacturer's instructions. Sequencing reads were processed using CASAVA 1.8 software (Illumina, San Diego, Calif.) followed by quality testing using the FastQC algorithm (http:// www.bioinformatics.babraham.ac.uk/projects/FastQC /). Variants were identified using the genome analysis kit (GATK) and VarScan software. Coverage analysis was performed using Picard software, scale HsMetrics. SNP and insertion/deletion (indels) analyses were performed using different filtration steps. Annovar is used to annotate the variant list.
TABLE 1 detection primer sequence number of AKT1 (SEQ ID NO:)
| Group 1 | Forward primer | Reverse primer | Group 2 | Forward primer | Reverse primer |
| Pool_1 | 1 | 2 | Pool_2 | 27 | 28 |
| Pool_1 | 3 | 4 | Pool_2 | 29 | 30 |
| Pool_1 | 5 | 6 | Pool_2 | 31 | 32 |
| Pool_1 | 7 | 8 | Pool_2 | 33 | 34 |
| Pool_1 | 9 | 10 | Pool_2 | 35 | 36 |
| Pool_1 | 11 | 12 | Pool_2 | 37 | 38 |
| Pool_1 | 13 | 14 | Pool_2 | 39 | 40 |
| Pool_1 | 15 | 16 | Pool_2 | 41 | 42 |
| Pool_1 | 17 | 18 | Pool_2 | 43 | 44 |
| Pool_1 | 19 | 20 | Pool_2 | 45 | 46 |
| Pool_1 | 21 | 22 | Pool_2 | 47 | 48 |
| Pool_1 | 23 | 24 | Pool_2 | 49 | 50 |
| Pool_1 | 25 | 26 |
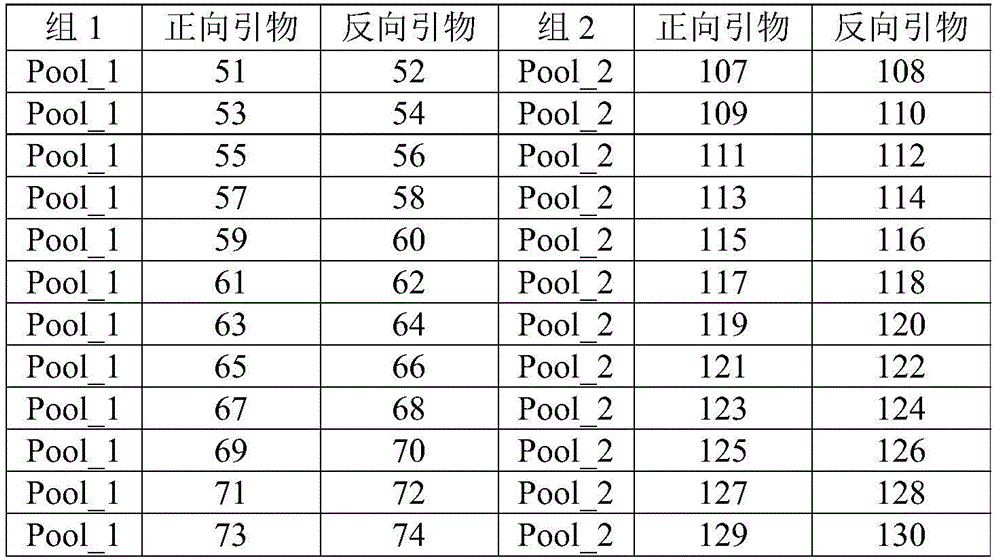
TABLE 2 ERBB2 detection primer sequence number (SEQ ID NO:)
TABLE 3 PIK3CA detection primer sequence number (SEQ ID NO:)
| Group 1 | Forward primer | Reverse primer | Group 2 | Forward primer | Reverse primer |
| Pool_1 | 161 | 162 | Pool_2 | 211 | 212 |
| Pool_1 | 163 | 164 | Pool_2 | 213 | 214 |
| Pool_1 | 165 | 166 | Pool_2 | 215 | 216 |
| Pool_1 | 167 | 168 | Pool_2 | 217 | 218 |
| Pool_1 | 169 | 170 | Pool_2 | 219 | 220 |
| Pool_1 | 171 | 172 | Pool_2 | 221 | 222 |
| Pool_1 | 173 | 174 | Pool_2 | 223 | 224 |
| Pool_1 | 175 | 176 | Pool_2 | 225 | 226 |
| Pool_1 | 177 | 178 | Pool_2 | 227 | 228 |
| Pool_1 | 179 | 180 | Pool_2 | 229 | 230 |
| Pool_1 | 181 | 182 | Pool_2 | 231 | 232 |
| Pool_1 | 183 | 184 | Pool_2 | 233 | 234 |
| Pool_1 | 185 | 186 | Pool_2 | 235 | 236 |
| Pool_1 | 187 | 188 | Pool_2 | 237 | 238 |
| Pool_1 | 189 | 190 | Pool_2 | 239 | 240 |
| Pool_1 | 191 | 192 | Pool_2 | 241 | 242 |
| Pool_1 | 193 | 194 | Pool_2 | 243 | 244 |
| Pool_1 | 195 | 196 | Pool_2 | 245 | 246 |
| Pool_1 | 197 | 198 | Pool_2 | 247 | 248 |
| Pool_1 | 199 | 200 | Pool_2 | 249 | 250 |
| Pool_1 | 201 | 202 | Pool_2 | 251 | 252 |
| Pool_1 | 203 | 204 | Pool_2 | 253 | 254 |
| Pool_1 | 205 | 206 | Pool_2 | 255 | 256 |
| Pool_1 | 207 | 208 | Pool_2 | 257 | 258 |
| Pool_1 | 209 | 210 | Pool_2 | 259 | 260 |
Results of the experiment
In the case of the breast cancer patient 001, when the tumor cell mutation PIK3CA p.H1047L was detected, it was found that Alpelisib + Fulvestrant was used as the treatment.
In the breast cancer patient 002, the tumor cell mutation PIK3CA p.E542K is detected, and the applicable treatment scheme Alpelisib + Fulvestrant is known.
In a breast cancer patient 003, a tumor cell mutation PIK3CA p.P366R is detected, and an applicable treatment scheme of Alpelisib + Fulvestrant is known.
In a breast cancer patient 004, tumor cell mutation AKT1 p.E17K is detected, and AZD5363 or Everolimus is known to be applicable to a treatment scheme.
In the breast cancer patient 005, the tumor cell mutation AKT1 p.E17K is detected, and the AZD5363 or Everolimus can be used as the treatment scheme.
In the breast cancer patient 006, the mutation of the tumor cell ERBB2 p.S310F is detected, and Apatinib, Lapatinib or Neratinib is applied as the treatment scheme.
In a breast cancer patient 007, tumor cell mutation ERBB2 p.D769Y is detected, and Apatinib, Lapatinib or Neratinib is applicable to a treatment scheme.
When the detection object is an FFPE sample, compared with about 50% of database construction success rate of 500 gene primer groups, the success rate of three-gene database construction is 100%, and the improvement effect is very obvious. Particularly, on the aspect of gene coverage, under the condition of the same data volume, the average coverage multiple of the primer group is 170 times that of the 500 genome primers, so that the capability of detecting mutant cells which do not have the advantage of simple quantity in the tumor is greatly improved, and more opportunities for precise medical treatment are obtained for breast cancer patients.
Sequence listing
<110> Shanghai human genome research center
<120> gene mutation detection primer set and use thereof
<130> 217887
<160> 260
<170> SIPOSequenceListing 1.0
<210> 1
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 1
cctctccatc cctccaagct at 22
<210> 2
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 2
tgagctgtct acacccacag at 22
<210> 3
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 3
gcctctctga gtgtggagag aa 22
<210> 4
<211> 26
<212> DNA
<213> Artificial Sequence
<400> 4
agactgacac caggtatttt gatgag 26
<210> 5
<211> 20
<212> DNA
<213> Artificial Sequence
<400> 5
ggcgagtgtg tgggaaatct 20
<210> 6
<211> 21
<212> DNA
<213> Artificial Sequence
<400> 6
gaggacgcca aggagatcat g 21
<210> 7
<211> 25
<212> DNA
<213> Artificial Sequence
<400> 7
aagctttggc tatcagtgta gtctg 25
<210> 8
<211> 19
<212> DNA
<213> Artificial Sequence
<400> 8
ttggtcccga ggccaagtc 19
<210> 9
<211> 24
<212> DNA
<213> Artificial Sequence
<400> 9
gagctcaaaa agcttctcat ggtc 24
<210> 10
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 10
aggtgctgga ggacaatgac ta 22
<210> 11
<211> 18
<212> DNA
<213> Artificial Sequence
<400> 11
gacgcagcaa cgcgtatg 18
<210> 12
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 12
ggagaacctc atgctggaca ag 22
<210> 13
<211> 20
<212> DNA
<213> Artificial Sequence
<400> 13
gtagtccagg gctgacacaa 20
<210> 14
<211> 19
<212> DNA
<213> Artificial Sequence
<400> 14
gacccggtct gagaaaccc 19
<210> 15
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 15
gcgtactcca tgacaaagca ga 22
<210> 16
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 16
gtatcaggcg acgtggtatc aa 22
<210> 17
<211> 19
<212> DNA
<213> Artificial Sequence
<400> 17
cggttctcgg tgagtgtgt 19
<210> 18
<211> 27
<212> DNA
<213> Artificial Sequence
<400> 18
catgaagatc ctcaagaagg aagtcat 27
<210> 19
<211> 19
<212> DNA
<213> Artificial Sequence
<400> 19
gcccagcagc ttcaggtac 19
<210> 20
<211> 19
<212> DNA
<213> Artificial Sequence
<400> 20
gtaggccacc aggtgtgaa 19
<210> 21
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 21
agggacacct ccatctcttc ag 22
<210> 22
<211> 20
<212> DNA
<213> Artificial Sequence
<400> 22
tcctgatctg gtacaggcct 20
<210> 23
<211> 19
<212> DNA
<213> Artificial Sequence
<400> 23
acgtaccgct cctcaggag 19
<210> 24
<211> 21
<212> DNA
<213> Artificial Sequence
<400> 24
atgcacgcag acagaggctc t 21
<210> 25
<211> 18
<212> DNA
<213> Artificial Sequence
<400> 25
cctggcagcg ggtactaa 18
<210> 26
<211> 24
<212> DNA
<213> Artificial Sequence
<400> 26
cacattcagc ttcctttgct tctc 24
<210> 27
<211> 19
<212> DNA
<213> Artificial Sequence
<400> 27
cgctgtccac acactccat 19
<210> 28
<211> 20
<212> DNA
<213> Artificial Sequence
<400> 28
ctggccgtgc tttaagaggt 20
<210> 29
<211> 23
<212> DNA
<213> Artificial Sequence
<400> 29
ggtggtgtga tggtgatcat ctg 23
<210> 30
<211> 23
<212> DNA
<213> Artificial Sequence
<400> 30
tgaaggtctt gagcacactt gag 23
<210> 31
<211> 21
<212> DNA
<213> Artificial Sequence
<400> 31
ctgccacacg ataccggcaa a 21
<210> 32
<211> 21
<212> DNA
<213> Artificial Sequence
<400> 32
ggccctacat cacaggagga a 21
<210> 33
<211> 21
<212> DNA
<213> Artificial Sequence
<400> 33
ccttcttgag cagccctgaa a 21
<210> 34
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 34
catgtacgag atgatgtgcg gt 22
<210> 35
<211> 20
<212> DNA
<213> Artificial Sequence
<400> 35
ggccgtagtc attgtcctcc 20
<210> 36
<211> 18
<212> DNA
<213> Artificial Sequence
<400> 36
cgtgcatacg cgttgctg 18
<210> 37
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 37
cagcccgaag tctgtgatct ta 22
<210> 38
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 38
actcggagaa gaacgtggtg ta 22
<210> 39
<211> 21
<212> DNA
<213> Artificial Sequence
<400> 39
gtctggtgcc atggagagta g 21
<210> 40
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 40
gccctgaagt actctttcca ga 22
<210> 41
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 41
gtgcccaaga agacaggaca tc 22
<210> 42
<211> 20
<212> DNA
<213> Artificial Sequence
<400> 42
gccctgcttt acaggacgag 20
<210> 43
<211> 25
<212> DNA
<213> Artificial Sequence
<400> 43
cttgaggatc ttcatggcgt agtag 25
<210> 44
<211> 19
<212> DNA
<213> Artificial Sequence
<400> 44
gtgcctgccc atagaccat 19
<210> 45
<211> 20
<212> DNA
<213> Artificial Sequence
<400> 45
gtggagtgct gagtgtctcc 20
<210> 46
<211> 20
<212> DNA
<213> Artificial Sequence
<400> 46
gctcacccag tgacaactca 20
<210> 47
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 47
cagtttccaa actgggctct ga 22
<210> 48
<211> 21
<212> DNA
<213> Artificial Sequence
<400> 48
tccatgtgga gactcctgag g 21
<210> 49
<211> 20
<212> DNA
<213> Artificial Sequence
<400> 49
gcgccacaga gaagttgttg 20
<210> 50
<211> 19
<212> DNA
<213> Artificial Sequence
<400> 50
acgggtagag tgtgcgtgg 19
<210> 51
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 51
gatgtgactg tctcctccca aa 22
<210> 52
<211> 25
<212> DNA
<213> Artificial Sequence
<400> 52
aataatggtt ttcccaccac atcct 25
<210> 53
<211> 20
<212> DNA
<213> Artificial Sequence
<400> 53
gcagggaaac ctggaactca 20
<210> 54
<211> 21
<212> DNA
<213> Artificial Sequence
<400> 54
gccacccacc tgtaaacaga g 21
<210> 55
<211> 20
<212> DNA
<213> Artificial Sequence
<400> 55
ctacgtgctc atcgctcaca 20
<210> 56
<211> 21
<212> DNA
<213> Artificial Sequence
<400> 56
tcagcgggtc tccattgtct a 21
<210> 57
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 57
ctctgctacc aggacacgat tt 22
<210> 58
<211> 24
<212> DNA
<213> Artificial Sequence
<400> 58
actgggttgt aagttgggag tttg 24
<210> 59
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 59
ccctgttctc cgatgtgtaa gg 22
<210> 60
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 60
cctgtatgac acctgcattc ca 22
<210> 61
<211> 21
<212> DNA
<213> Artificial Sequence
<400> 61
ctgcctggta ctgccctatt g 21
<210> 62
<211> 21
<212> DNA
<213> Artificial Sequence
<400> 62
gggattgggc atggactcaa a 21
<210> 63
<211> 19
<212> DNA
<213> Artificial Sequence
<400> 63
ggcacggtaa tgctgctca 19
<210> 64
<211> 20
<212> DNA
<213> Artificial Sequence
<400> 64
acttctcaca ccgctgtgtt 20
<210> 65
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 65
gttgacctgt cccggtatga ag 22
<210> 66
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 66
aggctcccaa agatcttctt gc 22
<210> 67
<211> 20
<212> DNA
<213> Artificial Sequence
<400> 67
ccagtgctgg gagtgatgtc 20
<210> 68
<211> 20
<212> DNA
<213> Artificial Sequence
<400> 68
cctgcagaac aggatgcaga 20
<210> 69
<211> 20
<212> DNA
<213> Artificial Sequence
<400> 69
gcgtcttcca gaacctgcaa 20
<210> 70
<211> 20
<212> DNA
<213> Artificial Sequence
<400> 70
agtactccct caggcccaaa 20
<210> 71
<211> 21
<212> DNA
<213> Artificial Sequence
<400> 71
gtgcccacct ttctcccata g 21
<210> 72
<211> 20
<212> DNA
<213> Artificial Sequence
<400> 72
gggttccgaa agagctggtc 20
<210> 73
<211> 20
<212> DNA
<213> Artificial Sequence
<400> 73
gccagttccc tggttcactt 20
<210> 74
<211> 20
<212> DNA
<213> Artificial Sequence
<400> 74
cgaaggaact ggctgcagtt 20
<210> 75
<211> 19
<212> DNA
<213> Artificial Sequence
<400> 75
caggagtgcg tggaggaat 19
<210> 76
<211> 18
<212> DNA
<213> Artificial Sequence
<400> 76
gtggcacggc aaacagtg 18
<210> 77
<211> 21
<212> DNA
<213> Artificial Sequence
<400> 77
gcctgtgccc actataagga c 21
<210> 78
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 78
tcctttctgc agaaaagacc gt 22
<210> 79
<211> 21
<212> DNA
<213> Artificial Sequence
<400> 79
cccaaactag ccctcaatcc c 21
<210> 80
<211> 19
<212> DNA
<213> Artificial Sequence
<400> 80
gcagtctccg catcgtgta 19
<210> 81
<211> 21
<212> DNA
<213> Artificial Sequence
<400> 81
ctgcaggaaa cggaggtgag g 21
<210> 82
<211> 21
<212> DNA
<213> Artificial Sequence
<400> 82
tcctcagctc cgtctctttc a 21
<210> 83
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 83
cccacgctct tctcactcat at 22
<210> 84
<211> 27
<212> DNA
<213> Artificial Sequence
<400> 84
acgtctaaga tttctttgtt ggctttg 27
<210> 85
<211> 21
<212> DNA
<213> Artificial Sequence
<400> 85
gtggtctccc ataccctctc a 21
<210> 86
<211> 24
<212> DNA
<213> Artificial Sequence
<400> 86
gttttcccgg acatggtcta agag 24
<210> 87
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 87
cagaaggtct acatgggtgc tt 22
<210> 88
<211> 27
<212> DNA
<213> Artificial Sequence
<400> 88
tgtaattttg acatggttgg gactctt 27
<210> 89
<211> 24
<212> DNA
<213> Artificial Sequence
<400> 89
gacattgacg agacagagta ccat 24
<210> 90
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 90
agtcctccaa ctgtgtgttg tg 22
<210> 91
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 91
gctacctgcc atgatgctag ac 22
<210> 92
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 92
gggatcccat cgtaaggttt gg 22
<210> 93
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 93
cccatctgca ccattgatgt ct 22
<210> 94
<211> 23
<212> DNA
<213> Artificial Sequence
<400> 94
cctgggtcta catacatcct ggt 23
<210> 95
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 95
gactctgaat gtcggccaag at 22
<210> 96
<211> 24
<212> DNA
<213> Artificial Sequence
<400> 96
gcatactgga ctcatctctc cttc 24
<210> 97
<211> 23
<212> DNA
<213> Artificial Sequence
<400> 97
ctctccttcc tccacagaat gag 23
<210> 98
<211> 23
<212> DNA
<213> Artificial Sequence
<400> 98
gggtaccaga tactcctcag cat 23
<210> 99
<211> 20
<212> DNA
<213> Artificial Sequence
<400> 99
cccatcccag atccgtgagt 20
<210> 100
<211> 24
<212> DNA
<213> Artificial Sequence
<400> 100
caggtcacca tcaaatacat cgga 24
<210> 101
<211> 21
<212> DNA
<213> Artificial Sequence
<400> 101
tctacagcgg tacagtgagg a 21
<210> 102
<211> 23
<212> DNA
<213> Artificial Sequence
<400> 102
gcccatcagt ctctctgctt aga 23
<210> 103
<211> 24
<212> DNA
<213> Artificial Sequence
<400> 103
gtcaccttct cttgaccttt caga 24
<210> 104
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 104
cccaaaggca aaaacgtctt tg 22
<210> 105
<211> 21
<212> DNA
<213> Artificial Sequence
<400> 105
ttcagcccag ccttcgacaa c 21
<210> 106
<211> 21
<212> DNA
<213> Artificial Sequence
<400> 106
tgaggacaca tcagggcttc t 21
<210> 107
<211> 21
<212> DNA
<213> Artificial Sequence
<400> 107
ccagtgtcct ctgacccatc t 21
<210> 108
<211> 20
<212> DNA
<213> Artificial Sequence
<400> 108
gtgggcaggt aggtgagttc 20
<210> 109
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 109
gggtggcagt gttcctattt ca 22
<210> 110
<211> 23
<212> DNA
<213> Artificial Sequence
<400> 110
ggcatagttg tcctcaaaga gct 23
<210> 111
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 111
gggaaagggt cctctgatca tt 22
<210> 112
<211> 23
<212> DNA
<213> Artificial Sequence
<400> 112
agctggttgt tcttgtggaa gat 23
<210> 113
<211> 23
<212> DNA
<213> Artificial Sequence
<400> 113
gctgtttgtg cctctctctg tta 23
<210> 114
<211> 25
<212> DNA
<213> Artificial Sequence
<400> 114
ctctgacaat cctcagaact ctctc 25
<210> 115
<211> 20
<212> DNA
<213> Artificial Sequence
<400> 115
gcactctgac tgcctggtat 20
<210> 116
<211> 21
<212> DNA
<213> Artificial Sequence
<400> 116
gctttcactg tgctgttttg c 21
<210> 117
<211> 21
<212> DNA
<213> Artificial Sequence
<400> 117
ccctggtcac ctacaacaca g 21
<210> 118
<211> 28
<212> DNA
<213> Artificial Sequence
<400> 118
ccaccaaaat gagaaaactg tgtttctc 28
<210> 119
<211> 21
<212> DNA
<213> Artificial Sequence
<400> 119
ctgcacaacc aagaggtgac a 21
<210> 120
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 120
gggtctgagg aaggatagga ca 22
<210> 121
<211> 23
<212> DNA
<213> Artificial Sequence
<400> 121
gttaccagtg ccaatatcca gga 23
<210> 122
<211> 23
<212> DNA
<213> Artificial Sequence
<400> 122
catctggatc ctcaggactc tgt 23
<210> 123
<211> 21
<212> DNA
<213> Artificial Sequence
<400> 123
gtctctgcat cctgttctgc a 21
<210> 124
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 124
tgctcacttg tgcagaattc gt 22
<210> 125
<211> 21
<212> DNA
<213> Artificial Sequence
<400> 125
ggtaaggtgc ccacctttct c 21
<210> 126
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 126
gaggtgggtg ttatggtgga tg 22
<210> 127
<211> 20
<212> DNA
<213> Artificial Sequence
<400> 127
gggaccagct ctttcggaac 20
<210> 128
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 128
cagtctgcac aagtccaaga ac 22
<210> 129
<211> 20
<212> DNA
<213> Artificial Sequence
<400> 129
gtgtcaactg cagccagttc 20
<210> 130
<211> 21
<212> DNA
<213> Artificial Sequence
<400> 130
ctggcattca catactccct g 21
<210> 131
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 131
cccaactaag ggcctgatcc ta 22
<210> 132
<211> 23
<212> DNA
<213> Artificial Sequence
<400> 132
gtaggagagg tcaggtttca cac 23
<210> 133
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 133
caagagggtg gttcccagaa tt 22
<210> 134
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 134
gagtgaaggg caatgaaggg ta 22
<210> 135
<211> 21
<212> DNA
<213> Artificial Sequence
<400> 135
gcagcagaag atccggaagt a 21
<210> 136
<211> 19
<212> DNA
<213> Artificial Sequence
<400> 136
tcgctccgct aggtgtcag 19
<210> 137
<211> 20
<212> DNA
<213> Artificial Sequence
<400> 137
cgcagatgcg gatcctgaaa 20
<210> 138
<211> 20
<212> DNA
<213> Artificial Sequence
<400> 138
catcagaact gccgaccaca 20
<210> 139
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 139
ggccatcaaa gtgttgaggg aa 22
<210> 140
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 140
ggtttgcggg agtcatatct cc 22
<210> 141
<211> 20
<212> DNA
<213> Artificial Sequence
<400> 141
gctggtgaca cagcttatgc 20
<210> 142
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 142
ctccggagag acctgcaaag ag 22
<210> 143
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 143
ggatgagcta cctggaggat gt 22
<210> 144
<211> 23
<212> DNA
<213> Artificial Sequence
<400> 144
ccttggtcct tcacctaacc ttg 23
<210> 145
<211> 21
<212> DNA
<213> Artificial Sequence
<400> 145
tgtctagccc atgggagaac t 21
<210> 146
<211> 21
<212> DNA
<213> Artificial Sequence
<400> 146
cccatcacac accataactc c 21
<210> 147
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 147
tctctgcctt aggtgtgact gt 22
<210> 148
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 148
ccacgcacat ttgaccatga tc 22
<210> 149
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 149
ctgtcccttg ggactgtcta ga 22
<210> 151
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 151
atgcgggaga attcagacac ca 22
<210> 151
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 151
gggaaggaga gatgagtcca gt 22
<210> 152
<211> 20
<212> DNA
<213> Artificial Sequence
<400> 152
cggtagaagg tgctgtccaa 20
<210> 153
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 153
cctggtggat gctgaggagt at 22
<210> 154
<211> 23
<212> DNA
<213> Artificial Sequence
<400> 154
gtaagcagac agccacacag tgt 23
<210> 155
<211> 20
<212> DNA
<213> Artificial Sequence
<400> 155
acactagggc tggagccctc 20
<210> 156
<211> 20
<212> DNA
<213> Artificial Sequence
<400> 156
taccgctgta gagggctggg 20
<210> 157
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 157
gcctggtatg gagtccagtc ta 22
<210> 158
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 158
cgaacatctg gctggttcac at 22
<210> 159
<211> 21
<212> DNA
<213> Artificial Sequence
<400> 159
gggtcgtcaa agacgttttt g 21
<210> 160
<211> 19
<212> DNA
<213> Artificial Sequence
<400> 160
tgggtcctgg tcccagtaa 19
<210> 161
<211> 30
<212> DNA
<213> Artificial Sequence
<400> 161
tttccttctt tgatttaggt ttctgctttg 30
<210> 162
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 162
ccacagttca cctgatgatg gt 22
<210> 163
<211> 30
<212> DNA
<213> Artificial Sequence
<400> 163
cccaagaatc ctagtagaat gtttactacc 30
<210> 164
<211> 30
<212> DNA
<213> Artificial Sequence
<400> 164
aagaagattc atcttgaaga agttgatgga 30
<210> 165
<211> 25
<212> DNA
<213> Artificial Sequence
<400> 165
tgaaacaaga cgactttgtg acctt 25
<210> 166
<211> 33
<212> DNA
<213> Artificial Sequence
<400> 166
gaaatatttt agaaagggac aacagttaag ctt 33
<210> 167
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 167
ggcatgccag tgtgtgaatt tg 22
<210> 168
<211> 30
<212> DNA
<213> Artificial Sequence
<400> 168
catttggagg atagacatac attgctctac 30
<210> 169
<211> 27
<212> DNA
<213> Artificial Sequence
<400> 169
tcccttgaaa aatgaaagag agatggt 27
<210> 170
<211> 27
<212> DNA
<213> Artificial Sequence
<400> 170
tgattgcttc agcaattact tgttctg 27
<210> 171
<211> 28
<212> DNA
<213> Artificial Sequence
<400> 171
ctctgtgttt tagaatatca gggcaagt 28
<210> 172
<211> 30
<212> DNA
<213> Artificial Sequence
<400> 172
aaaaatacag atactcatcc tcaatgtgat 30
<210> 173
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 173
ttgggaggat gcccaatttg at 22
<210> 174
<211> 29
<212> DNA
<213> Artificial Sequence
<400> 174
gtttctccat tcatatatgg tgtagctgt 29
<210> 175
<211> 30
<212> DNA
<213> Artificial Sequence
<400> 175
cagaataaaa attctttgtg caacctacgt 30
<210> 176
<211> 30
<212> DNA
<213> Artificial Sequence
<400> 176
tctgaaaaca tactacaggt caacagatta 30
<210> 177
<211> 32
<212> DNA
<213> Artificial Sequence
<400> 177
tttttctttt agatctatgt tcgaacaggt at 32
<210> 178
<211> 33
<212> DNA
<213> Artificial Sequence
<400> 178
actgctaaac actaatataa cctttggaaa tat 33
<210> 179
<211> 27
<212> DNA
<213> Artificial Sequence
<400> 179
tttacatagg tggaatgaat ggctgaa 27
<210> 180
<211> 30
<212> DNA
<213> Artificial Sequence
<400> 180
agcggtataa tcaggagttt ttaaaggtaa 30
<210> 181
<211> 30
<212> DNA
<213> Artificial Sequence
<400> 181
tttgattaca cagacactct agtatctgga 30
<210> 182
<211> 27
<212> DNA
<213> Artificial Sequence
<400> 182
aagagagaag gtttgactgc cataaaa 27
<210> 183
<211> 26
<212> DNA
<213> Artificial Sequence
<400> 183
gtggtaaagt tcccagatat gtcagt 26
<210> 184
<211> 30
<212> DNA
<213> Artificial Sequence
<400> 184
ggatcctttt ccatagagaa agtatctacc 30
<210> 185
<211> 33
<212> DNA
<213> Artificial Sequence
<400> 185
gcaagaatgt ttatgtttat tttgtttctc cca 33
<210> 186
<211> 33
<212> DNA
<213> Artificial Sequence
<400> 186
gtgacatacc aatttgtaca acagttatct agt 33
<210> 187
<211> 32
<212> DNA
<213> Artificial Sequence
<400> 187
catacacaga tgtattgctt ggtaaaagat tg 32
<210> 188
<211> 33
<212> DNA
<213> Artificial Sequence
<400> 188
cctgtactag ctgaattaaa tactgagaaa gtt 33
<210> 189
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 189
ccatgcagaa actgaccctg at 22
<210> 190
<211> 33
<212> DNA
<213> Artificial Sequence
<400> 190
gggaaaataa ttagacttac tttaaatgcc aaa 33
<210> 191
<211> 33
<212> DNA
<213> Artificial Sequence
<400> 191
ttcaaaaatg agtgtttaaa ttgtttagca aag 33
<210> 192
<211> 23
<212> DNA
<213> Artificial Sequence
<400> 192
ctccaaaagc aggccaaacc tct 23
<210> 193
<211> 27
<212> DNA
<213> Artificial Sequence
<400> 193
gggatgtatt tgaagcacct gaatagg 27
<210> 194
<211> 29
<212> DNA
<213> Artificial Sequence
<400> 194
aaaagagtct caaacacaaa ctagagtca 29
<210> 195
<211> 33
<212> DNA
<213> Artificial Sequence
<400> 195
tttttaatca ggtacagatg aagtttttag ttg 33
<210> 196
<211> 26
<212> DNA
<213> Artificial Sequence
<400> 196
ccaagaaagt acctgaggtt tcctag 26
<210> 197
<211> 33
<212> DNA
<213> Artificial Sequence
<400> 197
gaaagttgta aatctttgta acacttcaaa aag 33
<210> 198
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 198
caattcaacc acagtggcct tt 22
<210> 199
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 199
cccaaatttg catctgtggc at 22
<210> 200
<211> 28
<212> DNA
<213> Artificial Sequence
<400> 200
ggataacttt caacatacag gttgcctt 28
<210> 201
<211> 28
<212> DNA
<213> Artificial Sequence
<400> 201
cagaatgtta ccttatggtt gtctgtca 28
<210> 202
<211> 28
<212> DNA
<213> Artificial Sequence
<400> 202
catttctcct ttgttcttgt ctttgagc 28
<210> 203
<211> 28
<212> DNA
<213> Artificial Sequence
<400> 203
attcgttgtc agtgattgtt ttcattgt 28
<210> 204
<211> 25
<212> DNA
<213> Artificial Sequence
<400> 204
gctacacagt atccagcaca tgaac 25
<210> 205
<211> 24
<212> DNA
<213> Artificial Sequence
<400> 205
aacatcatgg tgaaagacga tgga 24
<210> 206
<211> 30
<212> DNA
<213> Artificial Sequence
<400> 206
cttgtgatcc aaaaagtgtc caaaatctat 30
<210> 207
<211> 30
<212> DNA
<213> Artificial Sequence
<400> 207
aatagtgatt agtaaaggag cccaagaatg 30
<210> 208
<211> 30
<212> DNA
<213> Artificial Sequence
<400> 208
ccagccaaca tttattattt tgaaattgct 30
<210> 209
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 209
cgacagcatg ccaatctctt ca 22
<210> 210
<211> 29
<212> DNA
<213> Artificial Sequence
<400> 210
ccatgatgtg catcattcat ttgtttcat 29
<210> 211
<211> 29
<212> DNA
<213> Artificial Sequence
<400> 211
gtaaaacttg caaagaatca gaacaatgc 29
<210> 212
<211> 24
<212> DNA
<213> Artificial Sequence
<400> 212
cacggaggca ttctaaagtc acta 24
<210> 213
<211> 30
<212> DNA
<213> Artificial Sequence
<400> 213
tgaggctaca ttaataacca taaagcatga 30
<210> 214
<211> 28
<212> DNA
<213> Artificial Sequence
<400> 214
ccctttctgc ttcttgagta acacttac 28
<210> 215
<211> 26
<212> DNA
<213> Artificial Sequence
<400> 215
cttccaaatc tacagagttc cctgtt 26
<210> 216
<211> 23
<212> DNA
<213> Artificial Sequence
<400> 216
cggaagtcct gtacttctgg atc 23
<210> 217
<211> 23
<212> DNA
<213> Artificial Sequence
<400> 217
gctgtggatc ttagggacct caa 23
<210> 218
<211> 30
<212> DNA
<213> Artificial Sequence
<400> 218
acaggtagaa gactgcacta tagtaatgat 30
<210> 219
<211> 29
<212> DNA
<213> Artificial Sequence
<400> 219
aagcagaagt atactctgaa aatcaacca 29
<210> 220
<211> 26
<212> DNA
<213> Artificial Sequence
<400> 220
ggaagtattc atcacatcca cacact 26
<210> 221
<211> 25
<212> DNA
<213> Artificial Sequence
<400> 221
atgaaaaacc ttacaggaaa tggct 25
<210> 222
<211> 26
<212> DNA
<213> Artificial Sequence
<400> 222
tggcagttga gaataaaggc tttctt 26
<210> 223
<211> 28
<212> DNA
<213> Artificial Sequence
<400> 223
gactgtttta caatgccatc ttattcca 28
<210> 224
<211> 30
<212> DNA
<213> Artificial Sequence
<400> 224
catttgactt taccttatca atgtctcgaa 30
<210> 225
<211> 30
<212> DNA
<213> Artificial Sequence
<400> 225
gaaccaagct atatctgaac aaaaattccg 30
<210> 226
<211> 27
<212> DNA
<213> Artificial Sequence
<400> 226
tgttcacatt gtcacataag ggttctc 27
<210> 227
<211> 27
<212> DNA
<213> Artificial Sequence
<400> 227
ccaatcaatc tctttcctgt ttttcgt 27
<210> 228
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 228
gcacgaggaa gatcaggaat gt 22
<210> 229
<211> 28
<212> DNA
<213> Artificial Sequence
<400> 229
gggaagaaaa gtgttttgaa atgtgttt 28
<210> 230
<211> 24
<212> DNA
<213> Artificial Sequence
<400> 230
ccatgaggta ctggccaaag attc 24
<210> 231
<211> 27
<212> DNA
<213> Artificial Sequence
<400> 231
tttttatggc agtcaaacct tctctct 27
<210> 232
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 232
cgggatacag accaattggc at 22
<210> 233
<211> 31
<212> DNA
<213> Artificial Sequence
<400> 233
attttacaga gtaacagact agctagagac a 31
<210> 234
<211> 31
<212> DNA
<213> Artificial Sequence
<400> 234
gaaaaagaaa cagagaatct ccattttagc a 31
<210> 235
<211> 28
<212> DNA
<213> Artificial Sequence
<400> 235
tcagaagtta aggcagtgtt ttagatgg 28
<210> 236
<211> 24
<212> DNA
<213> Artificial Sequence
<400> 236
gaagttccat agcctgttca ggtt 24
<210> 237
<211> 26
<212> DNA
<213> Artificial Sequence
<400> 237
gctgttcggt gcttggaaaa atattt 26
<210> 238
<211> 30
<212> DNA
<213> Artificial Sequence
<400> 238
agcataaaac tagttagtgc agtaggtttt 30
<210> 239
<211> 30
<212> DNA
<213> Artificial Sequence
<400> 239
gaagaaagca ttgactaatc aaaggattgg 30
<210> 240
<211> 33
<212> DNA
<213> Artificial Sequence
<400> 240
actcttccag ccaaacataa acaaaagtat ata 33
<210> 241
<211> 31
<212> DNA
<213> Artificial Sequence
<400> 241
acgattcttt tagatctgag atgcacaata a 31
<210> 242
<211> 32
<212> DNA
<213> Artificial Sequence
<400> 242
gaatgtcagt taagttaatg agcttttcca tt 32
<210> 243
<211> 29
<212> DNA
<213> Artificial Sequence
<400> 243
tgtagattca gagactgtac agtactgag 29
<210> 244
<211> 25
<212> DNA
<213> Artificial Sequence
<400> 244
tgtagagcat ccatgaaatc tggtc 25
<210> 245
<211> 23
<212> DNA
<213> Artificial Sequence
<400> 245
gggctttctg tctcctctaa acc 23
<210> 246
<211> 33
<212> DNA
<213> Artificial Sequence
<400> 246
ggagaatgag agagagaagc ataaattatt gaa 33
<210> 247
<211> 26
<212> DNA
<213> Artificial Sequence
<400> 247
aaggcttgaa gagtgtcgaa ttatgt 26
<210> 248
<211> 33
<212> DNA
<213> Artificial Sequence
<400> 248
agatagctaa attcatgcat cataagctca tta 33
<210> 249
<211> 23
<212> DNA
<213> Artificial Sequence
<400> 249
gcagtaaagg tcatgcatga caa 23
<210> 250
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 250
caccacctca ataagtccca ca 22
<210> 251
<211> 23
<212> DNA
<213> Artificial Sequence
<400> 251
ctgcagttca acagccacac act 23
<210> 252
<211> 28
<212> DNA
<213> Artificial Sequence
<400> 252
ttctaactca gaggaataca caaacacc 28
<210> 253
<211> 28
<212> DNA
<213> Artificial Sequence
<400> 253
ttctttgtag atatgatgca gccattga 28
<210> 254
<211> 30
<212> DNA
<213> Artificial Sequence
<400> 254
acaccaaaac attttaaaca gagaaaacca 30
<210> 255
<211> 29
<212> DNA
<213> Artificial Sequence
<400> 255
gattttggac actttttgga tcacaagaa 29
<210> 256
<211> 24
<212> DNA
<213> Artificial Sequence
<400> 256
ctcgagctca cctctcaaat tctc 24
<210> 257
<211> 27
<212> DNA
<213> Artificial Sequence
<400> 257
acatttgagc aaagacctga aggtatt 27
<210> 258
<211> 22
<212> DNA
<213> Artificial Sequence
<400> 258
ttccagagcc aagcatcatt ga 22
<210> 259
<211> 27
<212> DNA
<213> Artificial Sequence
<400> 259
ccttagataa aactgagcaa gaggctt 27
<210> 260
<211> 28
<212> DNA
<213> Artificial Sequence
<400> 260
ccagagtgag ctttcatttt ctcagtta 28
Claims (10)
1. A primer composition comprising:
(1) any one or more of SEQ ID NO 1-SEQ ID NO 260, or
(2) N and n +1, wherein n is any one or more or all odd numbers of 1-259,
preferably, the primer composition comprises one, two or all three groups selected from:
(1) any one or more of SEQ ID NO 1-26, or a primer pair consisting of SEQ ID NO n and n +1, wherein n is any one or more or all odd numbers of 1-25, and/or any one or more of SEQ ID NO 27-50, or a primer pair consisting of SEQ ID NO n and n +1, wherein n is any one or more or all odd numbers of 27-49,
(2) any one or more of SEQ ID NO 51-106, or a primer pair consisting of SEQ ID NO n and n +1, wherein n is any one or more or all odd numbers of 51-105, and/or any one or more of SEQ ID NO 107-160, or a primer pair consisting of SEQ ID NO n and n +1, wherein n is any one or more or all odd numbers of 51-105,
(3) any one or more of SEQ ID NO 161-210, or a primer pair consisting of SEQ ID NO n and n +1, wherein n is any one or more or all odd numbers of 161-209, and/or any one or more of SEQ ID NO 211-260, or a primer pair consisting of SEQ ID NO n and n +1, wherein n is any one or more or all odd numbers of 211-259.
2. The primer composition of claim 1, comprising one, two, or all three groups selected from:
(1) any one or more of SEQ ID NO 1-50, or a primer pair consisting of SEQ ID NO n and n +1, wherein n is any one or more or all odd numbers of 1-49,
(2) any one or more of SEQ ID NO 51-160, or a primer pair consisting of SEQ ID NO n and n +1, wherein n is any one or more or all odd numbers of 51-159,
(3) any one or more of SEQ ID NO 161-260, or a primer pair consisting of SEQ ID NO n and n +1, wherein n is any one or more or all odd numbers of 161-259.
3. A method for detecting gene mutation, comprising the step of amplifying a genome of a subject using the primer composition according to claim 1 or 2,
preferably, the first and second electrodes are formed of a metal,
the method further comprises the step of determining the sequence of the amplification product, and/or
The method further comprises the step of determining the sequence of the portion of the gene covered by the amplification product based on the sequence of the amplification product, and/or
The method further comprises the step of comparing the sequence of the amplification product or the portion of the gene covered thereby with the corresponding wild type sequence to thereby determine the gene mutation.
4. Use of the primer composition of claim 1 or 2 for the preparation of a breast cancer identification kit or a kit for selecting a breast cancer treatment regimen,
preferably, the primer composition is used for detecting a mutation in a gene comprising one or two or three selected from the group consisting of: AKT1, ERBB2, PIK3 CA.
5. The use according to claim 4,
the gene mutation comprises a mutation of AKT1, and the primer composition comprises: 1 to 50 or a primer pair consisting of SEQ ID NO n and n +1, wherein n is any one or more or all odd numbers of 1 to 49; preferably, the breast cancer is AKT1 mutant breast cancer and the treatment regimen comprises administering an agent that treats AKT1 mutant breast cancer, or
The gene mutation comprises a mutation of ERBB2, and the primer composition comprises: any one or more of SEQ ID NO 51-160, or a primer pair consisting of SEQ ID NO n and n +1, wherein n is any one or more or all odd numbers of 51-159; preferably, the breast cancer is an ERBB2 mutant breast cancer, the treatment regimen comprising administering an agent that treats ERBB2 mutant breast cancer, or
The gene mutation comprises a mutation of PIK3CA, and the primer composition comprises: any one or more of SEQ ID NO 161-260, or a primer pair consisting of SEQ ID NO n and n +1, wherein n is any one or more or all odd numbers of 161-259; preferably, the breast cancer is PIK3CA mutant breast cancer and the treatment regimen comprises administering an agent that treats PIK3CA mutant breast cancer.
6. Use of the primer composition of claim 1 or 2 for the preparation of a kit for breast cancer treatment.
7. The use according to claim 6,
the primer composition comprises: (1) 1-26 or a primer pair consisting of SEQ ID NOs n and n +1, wherein n is any one or more or all odd numbers from 1 to 25, and/or (2) any one or more of SEQ ID NOs 27 to 50 or a primer pair consisting of SEQ ID NOs n and n +1, wherein n is any one or more or all odd numbers from 27 to 49; preferably, the kit further comprises a medicament for treating AKT1 mutant breast cancer, and/or
The primer composition comprises: (1) any one or more of SEQ ID NO 51-106, or a primer pair consisting of SEQ ID NO n and n +1, wherein n is any one or more or all odd numbers of 51-105, and/or (2) any one or more of SEQ ID NO 107-160, or a primer pair consisting of SEQ ID NO n and n +1, wherein n is any one or more or all odd numbers of 51-105; preferably, the kit further comprises a medicament for treating ERBB2 mutant breast cancer, and/or
The primer composition comprises: (1) any one or more of SEQ ID NO:161-210, or a primer pair consisting of SEQ ID NO: n and n +1, wherein n is any one or more or all odd numbers of 161-209, and/or (2) any one or more of SEQ ID NO:211-260, or a primer pair consisting of SEQ ID NO: n and n +1, wherein n is any one or more or all odd numbers of 211-250; preferably, the kit further comprises a medicament for treating PIK3CA mutant breast cancer.
8. A kit comprising the primer composition of claim 1 or 2, optionally further comprising a drug for the treatment of breast cancer.
9. The kit of claim 8,
the primer composition comprises: (1) 1-26 or a primer pair consisting of SEQ ID NOs n and n +1, wherein n is any one or more or all odd numbers from 1 to 25, and/or (2) any one or more of SEQ ID NOs 27 to 50 or a primer pair consisting of SEQ ID NOs n and n +1, wherein n is any one or more or all odd numbers from 27 to 49; preferably, the medicament comprises a medicament for treating AKT1 mutant breast cancer, and/or
The primer composition comprises: (1) any one or more of SEQ ID NO 51-106, or a primer pair consisting of SEQ ID NO n and n +1, wherein n is any one or more or all odd numbers of 51-105, and/or (2) any one or more of SEQ ID NO 107-160, or a primer pair consisting of SEQ ID NO n and n +1, wherein n is any one or more or all odd numbers of 51-105; preferably, the medicament comprises a medicament for treating ERBB2 mutant breast cancer, and/or
The primer composition comprises: (1) any one or more of SEQ ID NO:161-210, or a primer pair consisting of SEQ ID NO: n and n +1, wherein n is any one or more or all odd numbers of 161-209, and/or (2) any one or more of SEQ ID NO:211-260, or a primer pair consisting of SEQ ID NO: n and n +1, wherein n is any one or more or all odd numbers of 211-250; preferably, the medicament comprises a medicament for treating PIK3CA mutant breast cancer.
10. An apparatus for providing a treatment plan comprising a processor and a memory storing a program, the processor when executing the program performing the steps of:
(1) identifying whether a breast cancer tumor cell of a subject has a mutation in one, two or more genes selected from AKT1, ERBB2, PIK3CA using a primer composition according to any of the embodiments of the first aspect of the invention,
(2) providing a treatment regimen according to the results of (1),
preferably, the first and second electrodes are formed of a metal,
the gene comprises AKT1, the treatment regimen comprises determining whether to treat a drug of AKT1 mutant breast cancer, and/or
The gene comprises ERBB2, the treatment regimen comprises determining whether to administer an agent to treat ERBB2 mutant breast cancer, and/or
The gene includes PIK3CA, and the treatment regimen includes determining whether to administer an agent to treat PIK3CA mutant breast cancer.
Priority Applications (1)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| CN202111286587.3A CN113943805A (en) | 2021-11-02 | 2021-11-02 | Gene mutation detection primer set and its use |
Applications Claiming Priority (1)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| CN202111286587.3A CN113943805A (en) | 2021-11-02 | 2021-11-02 | Gene mutation detection primer set and its use |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| CN113943805A true CN113943805A (en) | 2022-01-18 |
Family
ID=79337321
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| CN202111286587.3A Pending CN113943805A (en) | 2021-11-02 | 2021-11-02 | Gene mutation detection primer set and its use |
Country Status (1)
| Country | Link |
|---|---|
| CN (1) | CN113943805A (en) |
Citations (6)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN102994620A (en) * | 2011-09-13 | 2013-03-27 | 广州益善生物技术有限公司 | AKT1 gene mutation detection specificity primer and liquid chip thereof |
| CN106939337A (en) * | 2017-03-21 | 2017-07-11 | 重庆市肿瘤研究所 | The construction method of hormone-receptor-positive breast cancer recurrence monitoring gene mutation library |
| CN108004317A (en) * | 2017-11-09 | 2018-05-08 | 上海赛安生物医药科技股份有限公司 | PIK3CA detection in gene mutation system and its kit |
| CN109022573A (en) * | 2017-06-12 | 2018-12-18 | 深圳华大基因研究院 | The combination of breast cancer PIK3CA hot spot mutation detection probe primer sequence and kit |
| CN109715805A (en) * | 2016-04-01 | 2019-05-03 | 艾维迪提生物科学有限责任公司 | Nucleic acid-polypeptide compositions and uses thereof |
| CN110337450A (en) * | 2017-03-02 | 2019-10-15 | 基因泰克公司 | Adjunctive treatment for HER2 positive breast cancer |
-
2021
- 2021-11-02 CN CN202111286587.3A patent/CN113943805A/en active Pending
Patent Citations (6)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN102994620A (en) * | 2011-09-13 | 2013-03-27 | 广州益善生物技术有限公司 | AKT1 gene mutation detection specificity primer and liquid chip thereof |
| CN109715805A (en) * | 2016-04-01 | 2019-05-03 | 艾维迪提生物科学有限责任公司 | Nucleic acid-polypeptide compositions and uses thereof |
| CN110337450A (en) * | 2017-03-02 | 2019-10-15 | 基因泰克公司 | Adjunctive treatment for HER2 positive breast cancer |
| CN106939337A (en) * | 2017-03-21 | 2017-07-11 | 重庆市肿瘤研究所 | The construction method of hormone-receptor-positive breast cancer recurrence monitoring gene mutation library |
| CN109022573A (en) * | 2017-06-12 | 2018-12-18 | 深圳华大基因研究院 | The combination of breast cancer PIK3CA hot spot mutation detection probe primer sequence and kit |
| CN108004317A (en) * | 2017-11-09 | 2018-05-08 | 上海赛安生物医药科技股份有限公司 | PIK3CA detection in gene mutation system and its kit |
Non-Patent Citations (1)
| Title |
|---|
| 王帅: "极年轻乳腺癌的临床病理特征及全外显子 测序研究", 《中国博士学位论文全文数据库 医药卫生科技辑》, no. 6, 15 June 2021 (2021-06-15), pages 066 - 37 * |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| Xiao et al. | The long noncoding RNA TTTY15, which is located on the Y chromosome, promotes prostate cancer progression by sponging let-7 | |
| KR102622309B1 (en) | Detection of chromosomal interactions | |
| EP2536854B1 (en) | Personalized tumor biomarkers | |
| CN105671187B (en) | Group of genes for molecular typing of head and neck squamous cell carcinoma and application thereof | |
| CN107723352A (en) | A kind of Circulating tumor DNA liver cancer drives gene high-flux detection method | |
| CN104508149A (en) | Prediction of treatment response to JAK/STAT inhibitor | |
| CN104328164A (en) | Kit for detecting human EGFR gene mutation by using fluorescence probe hybridization method | |
| CN110055331B (en) | Kit for bladder cancer auxiliary diagnosis or screening and application thereof | |
| CN113646443A (en) | Compositions for diagnosing gliomas or predicting prognosis and methods of providing information about the same | |
| CN105463111A (en) | Primers and probes and kit for detecting five mutation types of human PIK3CA gene | |
| CN104845970A (en) | Gene relevant to papillary thyroid tumors | |
| CN110093423A (en) | Purposes of the gene marker in adenocarcinoma of lung diagnosis and treatment | |
| CN109735622A (en) | LncRNA relevant to colorectal cancer and its application | |
| CN106337058B (en) | CRYL1-IFT88 fusion gene and application thereof in diagnosis and treatment of primary hepatocellular carcinoma | |
| CN113943805A (en) | Gene mutation detection primer set and its use | |
| CN110714077B (en) | Primers and applications for simultaneous detection of BRCA1/2 exon sequences and chemotherapy drug sites | |
| CN104762399A (en) | Tumor circulating DNA KRAS mutation detection method | |
| CN108753981B (en) | Application of quantitative detection of HOXB8 gene in colorectal cancer prognosis judgment | |
| CN109457031B (en) | BRCA2 gene g.32338309A>G mutant and its application in auxiliary diagnosis of breast cancer | |
| CN112609000A (en) | Application of scar NA9 gene in early judgment of acute myeloid leukemia prognosis | |
| CN109897900B (en) | Application of EPB42 gene in liver cancer SBRT curative effect evaluation | |
| CN107365864A (en) | Detect BRCA1 genes, BRCA2 genes, the kit of PALB2 gene mutation sites and method | |
| CN107354197A (en) | A kind of kit for detecting mankind's NRAS gene mutations | |
| CN107723351A (en) | A kind of high-flux detection method of Circulating tumor DNA lung cancer driving gene | |
| CN105255869A (en) | Application of ABCC4 gene polymorphic site rs3742106 as well as detection primer and kit thereof |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| PB01 | Publication | ||
| PB01 | Publication | ||
| SE01 | Entry into force of request for substantive examination | ||
| SE01 | Entry into force of request for substantive examination |