CN112430657B - Methylation marker related to colorectal cancer and kit for detecting colorectal cancer - Google Patents
Methylation marker related to colorectal cancer and kit for detecting colorectal cancer Download PDFInfo
- Publication number
- CN112430657B CN112430657B CN202011165628.9A CN202011165628A CN112430657B CN 112430657 B CN112430657 B CN 112430657B CN 202011165628 A CN202011165628 A CN 202011165628A CN 112430657 B CN112430657 B CN 112430657B
- Authority
- CN
- China
- Prior art keywords
- methylation
- znf304
- lifr
- colorectal cancer
- gene
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Active
Links
- 238000007069 methylation reaction Methods 0.000 title claims abstract description 144
- 230000011987 methylation Effects 0.000 title claims abstract description 143
- 206010009944 Colon cancer Diseases 0.000 title claims abstract description 71
- 208000001333 Colorectal Neoplasms Diseases 0.000 title claims abstract description 71
- 239000003550 marker Substances 0.000 title claims abstract description 12
- 238000001514 detection method Methods 0.000 claims abstract description 57
- 101001042362 Homo sapiens Leukemia inhibitory factor receptor Proteins 0.000 claims abstract description 53
- 101000723909 Homo sapiens Zinc finger protein 304 Proteins 0.000 claims abstract description 53
- 102100028422 Zinc finger protein 304 Human genes 0.000 claims abstract description 45
- 102100021747 Leukemia inhibitory factor receptor Human genes 0.000 claims abstract description 40
- 108091029430 CpG site Proteins 0.000 claims abstract description 25
- 238000007847 digital PCR Methods 0.000 claims abstract description 23
- 101000869152 Homo sapiens Dynein axonemal assembly factor 9 Proteins 0.000 claims abstract description 15
- 239000000523 sample Substances 0.000 claims description 98
- 210000000349 chromosome Anatomy 0.000 claims description 21
- 239000002773 nucleotide Substances 0.000 claims description 21
- 125000003729 nucleotide group Chemical group 0.000 claims description 21
- 108090000623 proteins and genes Proteins 0.000 claims description 21
- 238000012360 testing method Methods 0.000 claims description 15
- 239000012634 fragment Substances 0.000 claims description 12
- 101150101871 ZNF304 gene Proteins 0.000 claims description 11
- 101150071228 Lifr gene Proteins 0.000 claims description 10
- 238000010791 quenching Methods 0.000 claims description 9
- 230000000171 quenching effect Effects 0.000 claims description 9
- 125000006853 reporter group Chemical group 0.000 claims description 6
- UDGUGZTYGWUUSG-UHFFFAOYSA-N 4-[4-[[2,5-dimethoxy-4-[(4-nitrophenyl)diazenyl]phenyl]diazenyl]-n-methylanilino]butanoic acid Chemical compound COC=1C=C(N=NC=2C=CC(=CC=2)N(C)CCCC(O)=O)C(OC)=CC=1N=NC1=CC=C([N+]([O-])=O)C=C1 UDGUGZTYGWUUSG-UHFFFAOYSA-N 0.000 claims description 3
- -1 CY3 Chemical compound 0.000 claims description 3
- ABZLKHKQJHEPAX-UHFFFAOYSA-N tetramethylrhodamine Chemical compound C=12C=CC(N(C)C)=CC2=[O+]C2=CC(N(C)C)=CC=C2C=1C1=CC=CC=C1C([O-])=O ABZLKHKQJHEPAX-UHFFFAOYSA-N 0.000 claims description 3
- 108020004414 DNA Proteins 0.000 abstract description 92
- 238000006243 chemical reaction Methods 0.000 abstract description 28
- 238000005516 engineering process Methods 0.000 abstract description 12
- 238000000034 method Methods 0.000 abstract description 12
- 102000056874 human ZNF304 Human genes 0.000 abstract description 8
- 238000003745 diagnosis Methods 0.000 abstract description 6
- 210000004369 blood Anatomy 0.000 abstract description 5
- 239000008280 blood Substances 0.000 abstract description 5
- 102100032237 Dynein axonemal assembly factor 9 Human genes 0.000 abstract description 4
- 238000011529 RT qPCR Methods 0.000 abstract description 3
- 101150106774 9 gene Proteins 0.000 abstract description 2
- 206010028980 Neoplasm Diseases 0.000 description 54
- 230000003321 amplification Effects 0.000 description 36
- 201000011510 cancer Diseases 0.000 description 36
- 238000003199 nucleic acid amplification method Methods 0.000 description 36
- 210000001519 tissue Anatomy 0.000 description 32
- 238000003908 quality control method Methods 0.000 description 20
- 210000005259 peripheral blood Anatomy 0.000 description 13
- 239000011886 peripheral blood Substances 0.000 description 13
- 210000002381 plasma Anatomy 0.000 description 13
- 238000012216 screening Methods 0.000 description 13
- 238000012163 sequencing technique Methods 0.000 description 13
- XLYOFNOQVPJJNP-UHFFFAOYSA-N water Substances O XLYOFNOQVPJJNP-UHFFFAOYSA-N 0.000 description 12
- LSNNMFCWUKXFEE-UHFFFAOYSA-M Bisulfite Chemical compound OS([O-])=O LSNNMFCWUKXFEE-UHFFFAOYSA-M 0.000 description 11
- 210000000265 leukocyte Anatomy 0.000 description 10
- 239000013612 plasmid Substances 0.000 description 10
- 230000007067 DNA methylation Effects 0.000 description 9
- 238000007403 mPCR Methods 0.000 description 9
- 239000000126 substance Substances 0.000 description 9
- 238000012408 PCR amplification Methods 0.000 description 8
- 238000004458 analytical method Methods 0.000 description 8
- 238000000137 annealing Methods 0.000 description 8
- 230000035945 sensitivity Effects 0.000 description 8
- 238000010790 dilution Methods 0.000 description 7
- 239000012895 dilution Substances 0.000 description 7
- 238000000605 extraction Methods 0.000 description 7
- 239000000090 biomarker Substances 0.000 description 6
- OPTASPLRGRRNAP-UHFFFAOYSA-N cytosine Chemical compound NC=1C=CNC(=O)N=1 OPTASPLRGRRNAP-UHFFFAOYSA-N 0.000 description 6
- 238000013461 design Methods 0.000 description 6
- 239000000203 mixture Substances 0.000 description 6
- 208000005443 Circulating Neoplastic Cells Diseases 0.000 description 5
- 101000632056 Homo sapiens Septin-9 Proteins 0.000 description 5
- 239000000463 material Substances 0.000 description 5
- 238000005987 sulfurization reaction Methods 0.000 description 5
- 108700028369 Alleles Proteins 0.000 description 4
- 101000756632 Homo sapiens Actin, cytoplasmic 1 Proteins 0.000 description 4
- 102100028024 Septin-9 Human genes 0.000 description 4
- 230000008901 benefit Effects 0.000 description 4
- 238000002052 colonoscopy Methods 0.000 description 4
- 238000004925 denaturation Methods 0.000 description 4
- 230000036425 denaturation Effects 0.000 description 4
- 238000011304 droplet digital PCR Methods 0.000 description 4
- 230000036541 health Effects 0.000 description 4
- 208000020816 lung neoplasm Diseases 0.000 description 4
- 108020004707 nucleic acids Proteins 0.000 description 4
- 102000039446 nucleic acids Human genes 0.000 description 4
- 150000007523 nucleic acids Chemical class 0.000 description 4
- 238000012257 pre-denaturation Methods 0.000 description 4
- 238000011002 quantification Methods 0.000 description 4
- 238000003753 real-time PCR Methods 0.000 description 4
- 210000002966 serum Anatomy 0.000 description 4
- 238000005486 sulfidation Methods 0.000 description 4
- 102000012406 Carcinoembryonic Antigen Human genes 0.000 description 3
- 108010022366 Carcinoembryonic Antigen Proteins 0.000 description 3
- 206010058467 Lung neoplasm malignant Diseases 0.000 description 3
- 108091028043 Nucleic acid sequence Proteins 0.000 description 3
- 230000000875 corresponding effect Effects 0.000 description 3
- 229940104302 cytosine Drugs 0.000 description 3
- 239000007850 fluorescent dye Substances 0.000 description 3
- 102000046686 human LIFR Human genes 0.000 description 3
- 201000005202 lung cancer Diseases 0.000 description 3
- 230000035772 mutation Effects 0.000 description 3
- 238000002360 preparation method Methods 0.000 description 3
- 230000004083 survival effect Effects 0.000 description 3
- 208000003200 Adenoma Diseases 0.000 description 2
- 206010001233 Adenoma benign Diseases 0.000 description 2
- 208000005623 Carcinogenesis Diseases 0.000 description 2
- 102000004190 Enzymes Human genes 0.000 description 2
- 108090000790 Enzymes Proteins 0.000 description 2
- 206010064571 Gene mutation Diseases 0.000 description 2
- 208000037062 Polyps Diseases 0.000 description 2
- 230000006907 apoptotic process Effects 0.000 description 2
- 238000003556 assay Methods 0.000 description 2
- 238000009534 blood test Methods 0.000 description 2
- 230000036952 cancer formation Effects 0.000 description 2
- 231100000504 carcinogenesis Toxicity 0.000 description 2
- 108091092240 circulating cell-free DNA Proteins 0.000 description 2
- 230000000295 complement effect Effects 0.000 description 2
- 238000001816 cooling Methods 0.000 description 2
- 238000007405 data analysis Methods 0.000 description 2
- 230000034994 death Effects 0.000 description 2
- 231100000517 death Toxicity 0.000 description 2
- 238000009826 distribution Methods 0.000 description 2
- 239000012154 double-distilled water Substances 0.000 description 2
- 238000013399 early diagnosis Methods 0.000 description 2
- 239000003480 eluent Substances 0.000 description 2
- 238000002474 experimental method Methods 0.000 description 2
- 230000002550 fecal effect Effects 0.000 description 2
- 230000006607 hypermethylation Effects 0.000 description 2
- 239000003112 inhibitor Substances 0.000 description 2
- 238000007855 methylation-specific PCR Methods 0.000 description 2
- 238000005457 optimization Methods 0.000 description 2
- 238000011895 specific detection Methods 0.000 description 2
- 238000012546 transfer Methods 0.000 description 2
- 238000012795 verification Methods 0.000 description 2
- 101150087690 ACTB gene Proteins 0.000 description 1
- 208000004804 Adenomatous Polyps Diseases 0.000 description 1
- 206010067484 Adverse reaction Diseases 0.000 description 1
- 206010055113 Breast cancer metastatic Diseases 0.000 description 1
- 108091029523 CpG island Proteins 0.000 description 1
- 108700039691 Genetic Promoter Regions Proteins 0.000 description 1
- 101100215371 Homo sapiens ACTB gene Proteins 0.000 description 1
- 101100343323 Homo sapiens LIFR gene Proteins 0.000 description 1
- 206010061902 Pancreatic neoplasm Diseases 0.000 description 1
- 208000005718 Stomach Neoplasms Diseases 0.000 description 1
- NINIDFKCEFEMDL-UHFFFAOYSA-N Sulfur Chemical compound [S] NINIDFKCEFEMDL-UHFFFAOYSA-N 0.000 description 1
- 108700025716 Tumor Suppressor Genes Proteins 0.000 description 1
- 102000044209 Tumor Suppressor Genes Human genes 0.000 description 1
- 230000001594 aberrant effect Effects 0.000 description 1
- 238000010521 absorption reaction Methods 0.000 description 1
- 239000002253 acid Substances 0.000 description 1
- 230000006838 adverse reaction Effects 0.000 description 1
- 230000032683 aging Effects 0.000 description 1
- 230000004075 alteration Effects 0.000 description 1
- 230000009286 beneficial effect Effects 0.000 description 1
- 238000005251 capillar electrophoresis Methods 0.000 description 1
- 230000030833 cell death Effects 0.000 description 1
- 208000029742 colonic neoplasm Diseases 0.000 description 1
- 230000002596 correlated effect Effects 0.000 description 1
- 230000001186 cumulative effect Effects 0.000 description 1
- 230000007423 decrease Effects 0.000 description 1
- 230000002950 deficient Effects 0.000 description 1
- 238000011161 development Methods 0.000 description 1
- 230000018109 developmental process Effects 0.000 description 1
- 235000008242 dietary patterns Nutrition 0.000 description 1
- 230000004069 differentiation Effects 0.000 description 1
- 201000010099 disease Diseases 0.000 description 1
- 208000037265 diseases, disorders, signs and symptoms Diseases 0.000 description 1
- 239000003814 drug Substances 0.000 description 1
- 229940079593 drug Drugs 0.000 description 1
- 230000000694 effects Effects 0.000 description 1
- 230000008995 epigenetic change Effects 0.000 description 1
- 230000004049 epigenetic modification Effects 0.000 description 1
- 238000009472 formulation Methods 0.000 description 1
- 206010017758 gastric cancer Diseases 0.000 description 1
- 210000001035 gastrointestinal tract Anatomy 0.000 description 1
- 230000014509 gene expression Effects 0.000 description 1
- 230000002068 genetic effect Effects 0.000 description 1
- 201000010536 head and neck cancer Diseases 0.000 description 1
- 208000014829 head and neck neoplasm Diseases 0.000 description 1
- 238000012203 high throughput assay Methods 0.000 description 1
- 238000012165 high-throughput sequencing Methods 0.000 description 1
- 230000000415 inactivating effect Effects 0.000 description 1
- 230000002779 inactivation Effects 0.000 description 1
- 230000003902 lesion Effects 0.000 description 1
- 238000011528 liquid biopsy Methods 0.000 description 1
- 230000003211 malignant effect Effects 0.000 description 1
- 208000015486 malignant pancreatic neoplasm Diseases 0.000 description 1
- 230000007246 mechanism Effects 0.000 description 1
- 239000012528 membrane Substances 0.000 description 1
- 238000012544 monitoring process Methods 0.000 description 1
- 230000017074 necrotic cell death Effects 0.000 description 1
- 239000013642 negative control Substances 0.000 description 1
- 238000007481 next generation sequencing Methods 0.000 description 1
- 238000010606 normalization Methods 0.000 description 1
- 239000002777 nucleoside Substances 0.000 description 1
- 125000003835 nucleoside group Chemical group 0.000 description 1
- 210000004940 nucleus Anatomy 0.000 description 1
- 201000002528 pancreatic cancer Diseases 0.000 description 1
- 208000008443 pancreatic carcinoma Diseases 0.000 description 1
- 238000002205 phenol-chloroform extraction Methods 0.000 description 1
- 230000035755 proliferation Effects 0.000 description 1
- 238000012552 review Methods 0.000 description 1
- 238000010079 rubber tapping Methods 0.000 description 1
- 238000007789 sealing Methods 0.000 description 1
- 238000010206 sensitivity analysis Methods 0.000 description 1
- 230000003068 static effect Effects 0.000 description 1
- 201000011549 stomach cancer Diseases 0.000 description 1
- 230000019635 sulfation Effects 0.000 description 1
- 238000005670 sulfation reaction Methods 0.000 description 1
- 229910052717 sulfur Inorganic materials 0.000 description 1
- 239000011593 sulfur Substances 0.000 description 1
- 238000002560 therapeutic procedure Methods 0.000 description 1
- 239000000107 tumor biomarker Substances 0.000 description 1
- 210000004881 tumor cell Anatomy 0.000 description 1
Images
Classifications
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q1/00—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions
- C12Q1/68—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions involving nucleic acids
- C12Q1/6876—Nucleic acid products used in the analysis of nucleic acids, e.g. primers or probes
- C12Q1/6883—Nucleic acid products used in the analysis of nucleic acids, e.g. primers or probes for diseases caused by alterations of genetic material
- C12Q1/6886—Nucleic acid products used in the analysis of nucleic acids, e.g. primers or probes for diseases caused by alterations of genetic material for cancer
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q1/00—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions
- C12Q1/68—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions involving nucleic acids
- C12Q1/6844—Nucleic acid amplification reactions
- C12Q1/6858—Allele-specific amplification
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q2600/00—Oligonucleotides characterized by their use
- C12Q2600/154—Methylation markers
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q2600/00—Oligonucleotides characterized by their use
- C12Q2600/16—Primer sets for multiplex assays
Landscapes
- Chemical & Material Sciences (AREA)
- Life Sciences & Earth Sciences (AREA)
- Health & Medical Sciences (AREA)
- Organic Chemistry (AREA)
- Proteomics, Peptides & Aminoacids (AREA)
- Zoology (AREA)
- Wood Science & Technology (AREA)
- Engineering & Computer Science (AREA)
- Genetics & Genomics (AREA)
- Analytical Chemistry (AREA)
- Immunology (AREA)
- Molecular Biology (AREA)
- General Engineering & Computer Science (AREA)
- Biotechnology (AREA)
- Biophysics (AREA)
- Physics & Mathematics (AREA)
- Biochemistry (AREA)
- Bioinformatics & Cheminformatics (AREA)
- Microbiology (AREA)
- General Health & Medical Sciences (AREA)
- Pathology (AREA)
- Hospice & Palliative Care (AREA)
- Oncology (AREA)
- Chemical Kinetics & Catalysis (AREA)
- Measuring Or Testing Involving Enzymes Or Micro-Organisms (AREA)
Abstract
本发明公开了一种结直肠癌相关的甲基化标志物以及用于检测结直肠癌的试剂盒。所述的甲基化标志物为分布在ZNF304,LIFR及C20orf194基因上的10个CpG位点。本发明首次在血液和组织DNA中检测了C20orf194、ZNF304、LIFR基因的甲基化水平,并首次使用qMSP以及数字PCR和MethyLight相结合的检测方法,通过优化PCR引物和探针,优化反应体系和反应条件,成功建立了基于qMSP以及ddPCR技术检测人ZNF304、LIFR基因甲基化的检测方法,和已有的基于qPCR技术的人SEPT 9基因甲基化检测方法相比,本发明的检测方法具有更高的特异性。ddPCR方法通过计算甲基化阳性液滴数和甲基化阳性液滴及非甲基化阳性液滴的比值进而对结直肠癌患者ZNF304、LIFR基因甲基化进行定性和定量检测,从而实现对结直肠癌的辅助诊断。The invention discloses a colorectal cancer-related methylation marker and a kit for detecting colorectal cancer. The methylation markers are 10 CpG sites distributed on ZNF304, LIFR and C20orf194 genes. The present invention detects the methylation levels of C20orf194, ZNF304 and LIFR genes in blood and tissue DNA for the first time, and uses the detection method combining qMSP and digital PCR and MethyLight for the first time. The reaction conditions have successfully established a detection method based on qMSP and ddPCR technology to detect the methylation of human ZNF304 and LIFR genes. Compared with the existing human SEPT 9 gene methylation detection method based on qPCR technology, the detection method of the present invention has higher specificity. The ddPCR method can qualitatively and quantitatively detect the methylation of ZNF304 and LIFR genes in colorectal cancer patients by calculating the number of methylation-positive droplets and the ratio of methylation-positive droplets to non-methylation-positive droplets. Auxiliary diagnosis of colorectal cancer.
Description
技术领域technical field
本发明涉及一种结直肠癌相关的甲基化标志物,还涉及一种用于检测结直肠癌的试剂盒,本发明属于生物医药技术领域。The invention relates to a colorectal cancer-related methylation marker, and also relates to a kit for detecting colorectal cancer, which belongs to the technical field of biomedicine.
背景技术Background technique
结直肠癌(colorectal cancer)是常见的消化道恶性疾病,据GLOBCAN 2018预计新发病例超过180万,约占每年癌症新发病例的10%,居世界第三位,而死亡率仅次于肺癌[1]。据估计2019年美国将有超过14万新发病例及5万死亡病例,约占全部癌症发病和死亡的十分之一[2]。结直肠癌的5年生存率可达到65%以上,早期诊断并治疗的患者5年生存率可达到80%以上,而Ⅳ期患者的5年生存率只有12%[3]。据国家癌症中心统计2000年到2011年间的数据显示,我国结直肠癌的发病率和死亡率均排在第五位,但发病率呈显著上升趋势,而男性增长速度一直高于女性[4]。与我国情况相反,自1990年美国的结直肠癌发病率和死亡率呈显著下降趋势,这与美国生活方式和饮食模式的转变尤其是早期筛查策略的制定密切相关[5],早期筛查能够早期发现并且清除处于癌变前期或早期的结直肠病变,进而避免部分肿瘤的发生与进展[6],而近年来发病率的迅速下降主要归功于结肠镜的使用,美国50岁及以上成年人的结肠镜检查使用率从2000年的21%增至2015年的60%,增长了近3倍;而50岁以下的成年人的发病率仍是呈现2%年增长率[7]。因此早期筛查是控制结直肠癌的有效措施。Colorectal cancer is a common malignant disease of the digestive tract. According to GLOBCAN 2018, it is estimated that there will be more than 1.8 million new cases, accounting for about 10% of new cancer cases each year, ranking third in the world, and the mortality rate is second only to lung cancer. [1] . It is estimated that there will be more than 140,000 new cases and 50,000 deaths in the United States in 2019, accounting for about one-tenth of all cancer incidence and deaths [2] . The 5-year survival rate of colorectal cancer can reach more than 65%, and the 5-year survival rate of patients with early diagnosis and treatment can reach more than 80%, while the 5-year survival rate of stage IV patients is only 12% [3] . According to the statistics of the National Cancer Center from 2000 to 2011, the incidence and mortality of colorectal cancer in China ranked fifth, but the incidence showed a significant upward trend, and the growth rate of men has been higher than that of women [4] . Contrary to the situation in China, the incidence and mortality of colorectal cancer in the United States have shown a significant downward trend since 1990, which is closely related to the changes in American lifestyles and dietary patterns, especially the formulation of early screening strategies [5] . It can early detect and remove colorectal lesions in the precancerous or early stage, thereby avoiding the occurrence and progression of some tumors [6] , and the rapid decline in the incidence in recent years is mainly due to the use of colonoscopy. Adults aged 50 and above in the United States The use of colonoscopy in 2000 increased from 21% in 2000 to 60% in 2015, an increase of nearly three-fold; while the incidence of adults under the age of 50 is still showing a 2% annual growth rate [7] . Therefore, early screening is an effective measure to control colorectal cancer.
结直肠癌的早期筛查主要包括粪便隐血试验(FOBT)和结肠镜检查等,粪便隐血试验具有简便无创等优点,但其灵敏度较低,假阳性率高,患者的依从性差[8]。结肠镜作为目前筛查,诊断和治疗结直肠癌的主流方式已经显著降低了美国欧洲等的发病及死亡率,但其有创性和不良反应,耗时,以及费用高可能会在一定程度上限制其的普及率[9]。并且只能提供一种静态的检查结果[10]。血清癌胚抗原(CEA)作为一种非侵入性生物标志物具有良好的特异性,可识别隐匿性CRC,但由于其敏感性低(40%至60%)而受到限制[11]。The early screening of colorectal cancer mainly includes fecal occult blood test (FOBT) and colonoscopy. The fecal occult blood test has the advantages of simplicity and non-invasiveness, but its sensitivity is low, the false positive rate is high, and the patient's compliance is poor [8] . Colonoscopy, as the current mainstream method for screening, diagnosing and treating colorectal cancer, has significantly reduced morbidity and mortality in the United States and Europe, but its invasiveness and adverse reactions, time-consuming, and high cost may be to some extent Limit its popularity [9] . And can only provide a static check result [10] . Serum carcinoembryonic antigen (CEA) has good specificity as a non-invasive biomarker to identify occult CRC, but is limited by its low sensitivity (40% to 60%) [11] .
循环肿瘤DNA(Circulating tumor DNA,ctDNA)主要由原发性肿瘤,循环肿瘤细胞(CTC),微小或明显转移到癌症患者血液中的DNA片段组成,ctDNA大部分来源于肿瘤细胞的凋亡和坏死[12]。Mandelp于1948年首次在正常人外周血发现游离DNA(cfDNA)[13]。1977年,Leon SA等再次发现肿瘤患者血浆cfDNA含量显著高于健康人,且晚期肿瘤患者的cfDNA水平更高[14]。直到1994年,Sorenson,G.D首次在胰腺癌患者血液中发现RAS突变,并且和组织中保持一致[15]。随后有研究表明ctDNA携带肿瘤特异性遗传改变,包括突变[16],拷贝数变异[17],同时发现ctDNA还携带有癌症组织特异的表观遗传改变,肿瘤组织的特异性甲基化模式与ctDNA高度相关[18]。因此通过对ctDNA的生物学特性检测可以达到微创快速的筛查癌症并能对癌症进行实时纵向监测。Circulating tumor DNA (ctDNA) is mainly composed of primary tumors, circulating tumor cells (CTCs), and DNA fragments that have been minutely or significantly transferred to the blood of cancer patients. Most of ctDNA is derived from apoptosis and necrosis of tumor cells. [12] . Mandelp first discovered cell-free DNA (cfDNA) in normal human peripheral blood in 1948 [13] . In 1977, Leon SA et al again found that the plasma cfDNA content of tumor patients was significantly higher than that of healthy people, and the cfDNA level of advanced tumor patients was higher [14] . Until 1994, Sorenson, GD found RAS mutation in the blood of pancreatic cancer patients for the first time, and it was consistent with the tissue [15] . Subsequent studies have shown that ctDNA carries tumor-specific genetic changes, including mutation [16] , copy number variation [17] , and found that ctDNA also carries cancer tissue-specific epigenetic changes, and tumor tissue-specific methylation patterns are related to ctDNA is highly correlated [18] . Therefore, the detection of biological characteristics of ctDNA can achieve minimally invasive and rapid cancer screening and real-time longitudinal monitoring of cancer.
DNA甲基化是表观遗传学修饰中的一种,涉及分化和发育,衰老,肿瘤发生等。全基因组低甲基化和启动子区CpG岛高甲基化是肿瘤组织中最常见的生物学现象[19]。异常甲基化通常会导致基因表达缺陷,抑癌基因的高甲基化是许多肿瘤的早期事件,并且可能也是与肿瘤发生有关的最早可检测到的肿瘤改变之一[20,21]。相比于突变检测,甲基化在癌症筛查及诊断方面更具优势,包括更高的临床敏感性和动态范围,多个可检测到的甲基化目标区域,以及每个目标基因组区域内多个改变的CpG位点[22]。可以将每个选定区域中的多个CpG基因座组合检测,以获得“癌症特异性甲基化模式”。此外,已经有研究表明甲基化分析可用于区分组织起源和癌症亚型[23]。因此,甲基化作为肿瘤诊断标志物要比基因突变更具优势,是应用潜力巨大的肿瘤特异诊断的生物标志物之一。DNA methylation is one of epigenetic modifications involved in differentiation and development, aging, tumorigenesis, etc. Genome-wide hypomethylation and CpG island hypermethylation in promoter regions are the most common biological phenomena in tumor tissues [19] . Aberrant methylation often results in defective gene expression, and hypermethylation of tumor suppressor genes is an early event in many tumors and may be one of the earliest detectable tumor alterations associated with tumorigenesis [20,21] . Compared with mutation detection, methylation has advantages in cancer screening and diagnosis, including higher clinical sensitivity and dynamic range, multiple detectable methylation target regions, and within each target genomic region Multiple altered CpG sites [22] . Multiple CpG loci in each selected region can be detected in combination to obtain "cancer-specific methylation patterns." In addition, it has been shown that methylation analysis can be used to distinguish tissue origin and cancer subtype [23] . Therefore, methylation has more advantages as a tumor diagnostic marker than gene mutation, and is one of the biomarkers with great potential for tumor-specific diagnosis.
血浆SEPT9启动子区甲基化已在结直肠癌中进行了广泛的研究,研究表明SEPT9甲基化用于诊断结直肠癌的灵敏度在48.2%到95.6%之间,特异度在69%到97.1%之间[24]。已经被证实是结直肠癌血浆检测的高度灵敏和特异性的生物标志物,并且于2016年获得了美国食品药品监督管理局(FDA)的批准[25,26]。但是SEPT9也在其他癌症血浆中例如肺癌[27],头颈部癌症[28],胃癌[29]中均呈现不同程度的高甲基化。因此,用于结直肠癌早期诊断的ctDNA特异性甲基化生物标记物仍有待进一步研究。Plasma SEPT9 promoter methylation has been extensively studied in colorectal cancer, and studies have shown that SEPT9 methylation has a sensitivity of 48.2% to 95.6% and a specificity of 69% to 97.1 for the diagnosis of colorectal cancer. % between [24] . It has been proven to be a highly sensitive and specific biomarker for colorectal cancer plasma detection and was approved by the US Food and Drug Administration (FDA) in 2016 [25,26] . However, SEPT9 is also hypermethylated to varying degrees in the plasma of other cancers such as lung cancer [27] , head and neck cancer [28] , and gastric cancer [29] . Therefore, ctDNA-specific methylation biomarkers for early diagnosis of colorectal cancer remain to be further investigated.
ctDNA在cfDNA中的占比在0.01%到60%不等,其浓度和肿瘤体积,分期,血管化,增殖率和细胞死亡率相关[30]。因此要对ctDNA甲基化水平实现灵敏可重复的检测是一项巨大的挑战。目前SEPT9检测试剂盒使用的是将甲基化特异性PCR(MSP)与TaqMan探针结合的检测方法,称MethyLight,其可以在校正DNA载量和标准化甲基化DNA标准品后,以相对方式定量甲基化等位基因的量,大大提高了检测和定量甲基化等位基因的能力[31]。然而,MethyLight仍然容易受到PCR抑制剂的影响,在未甲基化等位基因背景下检测稀有甲基化等位基因的灵敏度有限,并可能在标准化过程中出现偏差,这可能导致结果不一致,尤其是在检测低水平的DNA甲基化方面(包括低浓度和低分子量)。因此,常规的甲基化特异性定量PCR通常不能达到准确检测低水平DNA甲基化并以此作为癌症生物标志物的灵敏度[32]。数字PCR(droplet digital PCR)是一种核酸分子绝对定量技术,其最初被定义为单分子模板PCR扩增,其将样品分离成单个分子并通过PCR扩增;通过终点信号的存在来判断阴性和阳性,结合泊松分布来计算原始样品中的拷贝数[33];数字PCR不受扩增效率及PCR抑制物的影响[34]。有研究显示数字PCR可以从背景DNA中检测到0.001%的靶基因突变[35]。Ming等人证明相比于常规MethyLight PCR,MethyLight ddPCR的定量下限(LOQ)降低25倍,检测限(LOD)降低20倍[36]。因此MethyLight ddPCR是一种更适合于检测ctDNA甲基化水平的技术。The proportion of ctDNA in cfDNA varies from 0.01% to 60%, and its concentration correlates with tumor volume, stage, vascularization, proliferation rate and cell death rate [30] . Therefore, it is a great challenge to achieve sensitive and reproducible detection of ctDNA methylation levels. The current SEPT9 detection kit uses a detection method that combines methylation-specific PCR (MSP) with TaqMan probes, called MethyLight, which can be used in a relative manner after correcting for DNA load and normalizing methylated DNA standards. Quantifying the amount of methylated alleles greatly improves the ability to detect and quantify methylated alleles [31] . However, MethyLight is still susceptible to PCR inhibitors, has limited sensitivity to detect rare methylated alleles in a background of unmethylated alleles, and can be biased during normalization, which can lead to inconsistent results, especially is in the detection of low levels of DNA methylation (including low concentrations and low molecular weight). Therefore, conventional methylation-specific quantitative PCR usually cannot achieve the sensitivity to accurately detect low-level DNA methylation as a cancer biomarker [32] . Digital PCR (droplet digital PCR) is a technique for absolute quantification of nucleic acid molecules, which was originally defined as single-molecule template PCR amplification, which separates a sample into individual molecules and amplifies them by PCR; negative and Positive, combined with Poisson distribution to calculate the copy number in the original sample [33] ; digital PCR is not affected by amplification efficiency and PCR inhibitors [34] . Studies have shown that digital PCR can detect 0.001% of target gene mutations from background DNA [35] . Ming et al. demonstrated that MethyLight ddPCR has a 25-fold lower limit of quantitation (LOQ) and a 20-fold lower limit of detection (LOD) compared to conventional MethyLight PCR [36] . Therefore, MethyLight ddPCR is a more suitable technique for detecting the methylation level of ctDNA.
参考文献:references:
1.Bray,F.,J.Ferlay,I.Soerjomataram,R.L.Siegel,L.A.Torre,andA.Jemal.Global cancer statistics 2018:GLOBOCAN estimates of incidence andmortality worldwide for 36cancers in 185countries.CA Cancer J Clin.2018;68(6):394-424.1. Bray, F., J. Ferlay, I. Soerjomataram, R.L. Siegel, L.A. Torre, and A. Jemal. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36cancers in 185countries.CA Cancer J Clin.2018;68( 6): 394-424.
2.Siegel,R.L.,K.D.Miller,and A.Jemal.Cancer statistics,2019.CA CancerJ Clin.2019;69(1):7-34.2. Siegel, R.L., K.D. Miller, and A. Jemal. Cancer statistics, 2019. CA Cancer J Clin. 2019;69(1):7-34.
3.Miller,K.D.,L.Nogueira,A.B.Mariotto,J.H.Rowland,K.R.Yabroff,C.M.Alfano,A.Jemal,J.L.Kramer,and R.L.Siegel.Cancer treatment andsurvivorship statistics,2019.CA Cancer J Clin.2019;69(5):363-385.3. Miller, K.D., L.Nogueira, A.B.Marioto, J.H.Rowland, K.R.Yabroff, C.M.Alfano, A.Jemal, J.L.Kramer, and R.L.Siegel.Cancer treatment and survival statistics, 2019.CA Cancer J Clin.2019;69(5 ): 363-385.
4.Chen,W.,R.Zheng,P.D.Baade,S.Zhang,H.Zeng,F.Bray,A.Jemal,X.Q.Yu,andJ.He.Cancer statistics in China,2015.CA Cancer J Clin.2016;66(2):115-32.4.Chen,W.,R.Zheng,P.D.Baade,S.Zhang,H.Zeng,F.Bray,A.Jemal,X.Q.Yu,andJ.He.Cancer statistics in China,2015.CA Cancer J Clin.2016 ;66(2):115-32.
5.Singh,G.K.,B.A.Miller,and B.F.Hankey.Changing area socioeconomicpatterns in U.S.cancer mortality,1950-1998:Part II--Lung and colorectalcancers.J Natl Cancer Inst.2002;94(12):916-25.5. Singh, G.K., B.A. Miller, and B.F. Hankey. Changing area socioeconomic patterns in U.S. cancer mortality, 1950-1998:Part II--Lung and colorectalcancers.J Natl Cancer Inst.2002;94(12):916-25.
6.Edwards,B.K.,E.Ward,B.A.Kohler,C.Eheman,A.G.Zauber,R.N.Anderson,A.Jemal,M.J.Schymura,I.Lansdorp-Vogelaar,L.C.Seeff,M.van Ballegooijen,S.L.Goede,and L.A.Ries.Annual report to the nation on the status of cancer,1975-2006,featuring colorectal cancer trends and impact of interventions(riskfactors,screening,and treatment)to reduce future rates.Cancer.2010;116(3):544-73.6. Edwards, B.K., E.Ward, B.A. Kohler, C.Eheman, A.G.Zauber, R.N.Anderson, A.Jemal, M.J.Schymura, I.Lansdorp-Vogelaar, L.C.Seeff, M.van Ballegooijen, S.L.Goede, and L.A.Ries .Annual report to the nation on the status of cancer, 1975-2006, featuring colorectal cancer trends and impact of interventions (riskfactors, screening, and treatment) to reduce future rates. Cancer. 2010;116(3):544-73.
7.National Center for Health,S.,Health,United States,in Health,UnitedStates,2015:With Special Feature on Racial and Ethnic HealthDisparities.2016,National Center for Health Statistics(US):Hyattsville(MD).7.National Center for Health,S.,Health,United States,in Health,UnitedStates,2015:With Special Feature on Racial and Ethnic HealthDisparities.2016,National Center for Health Statistics(US):Hyattsville(MD).
8.Ibanez-Sanz,G.,M.Garcia,N.Mila,R.A.Hubbard,C.Vidal,G.Binefa,L.Benito,and V.Moreno.False-Positive Results in a Population-Based ColorectalScreening Program:Cumulative Risk from 2000 to 2017 with BiennialScreening.Cancer Epidemiol Biomarkers Prev.2019;28(11):1909-1916.8.Ibanez-Sanz,G.,M.Garcia,N.Mila,R.A.Hubbard,C.Vidal,G.Binefa,L.Benito,and V.Moreno.False-Positive Results in a Population-Based ColorectalScreening Program:Cumulative Risk from 2000 to 2017 with Biennial Screening. Cancer Epidemiol Biomarkers Prev. 2019;28(11):1909-1916.
9.Nee,J.,R.Z.Chippendale,and J.D.Feuerstein.Screening for ColonCancer in Older Adults:Risks,Benefits,and When to Stop.Mayo Clin Proc.2020;95(1):184-196.9. Nee, J., R.Z. Chippendale, and J.D. Feuerstein. Screening for Colon Cancer in Older Adults: Risks, Benefits, and When to Stop. Mayo Clin Proc. 2020;95(1):184-196.
10.Dominguez-Vigil,I.G.,A.K.Moreno-Martinez,J.Y.Wang,M.H.A.Roehrl,andH.A.Barrera-Saldana.The dawn of the liquid biopsy in the fight againstcancer.Oncotarget.2018;9(2):2912-2922.10. Dominguez-Vigil, I.G., A.K. Moreno-Martinez, J.Y. Wang, M.H.A. Roehrl, and H.A. Barrera-Saldana. The dawn of the liquid biopsy in the fight against cancer. Oncotarget. 2018;9(2):2912-2922 .
11.Macdonald,J.S.Carcinoembryonic antigen screening:pros andcons.Semin Oncol.1999;26(5):556-60.11. Macdonald, J.S. Carcinoembryonic antigen screening: pros and cons. Semin Oncol. 1999;26(5):556-60.
12.Stroun,M.,J.Lyautey,C.Lederrey,A.Olson-Sand,and P.Anker.About thepossible origin and mechanism of circulating DNA apoptosis and active DNArelease.Clin Chim Acta.2001;313(1-2):139-42.12. Stroun, M., J. Lyautey, C. Lederrey, A. Olson-Sand, and P. Anker. About the possible origin and mechanism of circulating DNA apoptosis and active DNA release. Clin Chim Acta. 2001; 313(1-2 ): 139-42.
13.Mandel,P.and P.Metais.Les acides nucléiques du plasma sanguin chezl'homme.C R Seances Soc Biol Fil.1948;142(3-4):241-3.13. Mandel, P. and P. Metais. Les acides nucléiques du plasma sanguin chezl'homme. C R Seances Soc Biol Fil. 1948;142(3-4):241-3.
14.Leon,S.A.,B.Shapiro,D.M.Sklaroff,and M.J.Yaros.Free DNA in theserum of cancer patients and the effect of therapy.Cancer Res.1977;37(3):646-50.14. Leon, S.A., B. Shapiro, D.M. Sklaroff, and M.J. Yaros. Free DNA in theserum of cancer patients and the effect of therapy. Cancer Res. 1977;37(3):646-50.
15.Sorenson,G.D.,D.M.Pribish,F.H.Valone,V.A.Memoli,D.J.Bzik,andS.L.Yao.Soluble normal and mutated DNA sequences from single-copy genes inhuman blood.Cancer Epidemiol Biomarkers Prev.1994;3(1):67-71.15. Sorenson, G.D., D.M.Pribish, F.H.Valone, V.A.Memoli, D.J.Bzik, and S.L.Yao.Soluble normal and mutated DNA sequences from single-copy genes inhuman blood.Cancer Epidemiol Biomarkers Prev.1994;3(1): 67-71.
16.Medford,A.J.,R.N.Gillani,and B.H.Park.Detection of Cancer DNA inEarly Stage and Metastatic Breast Cancer Patients.Methods Mol Biol.2018;1768:209-227.16. Medford, A.J., R.N.Gillani, and B.H.Park. Detection of Cancer DNA in Early Stage and Metastatic Breast Cancer Patients. Methods Mol Biol. 2018;1768:209-227.
17.Whale,A.S.,J.F.Huggett,S.Cowen,V.Speirs,J.Shaw,S.Ellison,C.A.Foy,and D.J.Scott.Comparison of microfluidic digital PCR and conventionalquantitative PCR for measuring copy number variation.Nucleic Acids Res.2012;40(11):e82.17.Whale,A.S.,J.F.Huggett,S.Cowen,V.Speirs,J.Shaw,S.Ellison,C.A.Foy,and D.J.Scott.Comparison of microfluidic digital PCR and conventional quantitative PCR for measuring copy number variation.Nucleic Acids Res. 2012;40(11):e82.
18.Xu,R.H.,W.Wei,M.Krawczyk,W.Wang,H.Luo,K.Flagg,S.Yi,W.Shi,Q.Quan,K.Li,L.Zheng,H.Zhang,B.A.Caughey,Q.Zhao,J.Hou,R.Zhang,Y.Xu,H.Cai,G.Li,R.Hou,Z.Zhong,D.Lin,X.Fu,J.Zhu,Y.Duan,M.Yu,B.Ying,W.Zhang,J.Wang,E.Zhang,C.Zhang,O.Li,R.Guo,H.Carter,J.K.Zhu,X.Hao,and K.Zhang.Circulating tumour DNAmethylation markers for diagnosis and prognosis of hepatocellularcarcinoma.Nat Mater.2017;16(11):1155-1161.18.Xu,R.H.,W.Wei,M.Krawczyk,W.Wang,H.Luo,K.Flagg,S.Yi,W.Shi,Q.Quan,K.Li,L.Zheng,H.Zhang, B.A.Caughey,Q.Zhao,J.Hou,R.Zhang,Y.Xu,H.Cai,G.Li,R.Hou,Z.Zhong,D.Lin,X.Fu,J.Zhu,Y.Duan ,M.Yu,B.Ying,W.Zhang,J.Wang,E.Zhang,C.Zhang,O.Li,R.Guo,H.Carter,J.K.Zhu,X.Hao,and K.Zhang.Circulating tumour DNAmethylation markers for diagnosis and prognosis of hepatocellular carcinoma. Nat Mater. 2017;16(11):1155-1161.
19.Suzuki,M.M.and A.Bird.DNA methylation landscapes:provocativeinsights from epigenomics.Nat Rev Genet.2008;9(6):465-76.19. Suzuki, M.M. and A. Bird. DNA methylation landscapes: provocativeinsights from epigenomics. Nat Rev Genet. 2008;9(6):465-76.
20.Lund,A.H.and M.van Lohuizen.Epigenetics and cancer.Genes Dev.2004;18(19):2315-35.20. Lund, A.H. and M. van Lohuizen. Epigenetics and cancer. Genes Dev. 2004;18(19):2315-35.
21.Baylin,S.B.and P.A.Jones.A decade of exploring the cancerepigenome-biological and translational implications.Nat Rev Cancer.2011;11(10):726-34.21. Baylin, S.B. and P.A. Jones. A decade of exploring the cancer epigenome-biological and translational implications. Nat Rev Cancer. 2011;11(10):726-34.
22.Laird,P.W.The power and the promise of DNA methylation markers.NatRev Cancer.2003;3(4):253-66.22. Laird, P.W. The power and the promise of DNA methylation markers. NatRev Cancer. 2003;3(4):253-66.
23.Moran,S.,A.Martinez-Cardus,S.Sayols,E.Musulen,C.Balana,A.Estival-Gonzalez,C.Moutinho,H.Heyn,A.Diaz-Lagares,M.C.de Moura,G.M.Stella,P.M.Comoglio,M.Ruiz-Miro,X.Matias-Guiu,R.Pazo-Cid,A.Anton,R.Lopez-Lopez,G.Soler,F.Longo,I.Guerra,S.Fernandez,Y.Assenov,C.Plass,R.Morales,J.Carles,D.Bowtell,L.Mileshkin,D.Sia,R.Tothill,J.Tabernero,J.M.Llovet,andM.Esteller.Epigenetic profiling to classify cancer of unknown primary:amulticentre,retrospective analysis.Lancet Oncol.2016;17(10):1386-1395.23.Moran,S.,A.Martinez-Cardus,S.Sayols,E.Musulen,C.Balana,A.Estival-Gonzalez,C.Moutinho,H.Heyn,A.Diaz-Lagares,M.C.de Moura,G.M. Stella,P.M.Comoglio,M.Ruiz-Miro,X.Matias-Guiu,R.Pazo-Cid,A.Anton,R.Lopez-Lopez,G.Soler,F.Longo,I.Guerra,S.Fernandez,Y .Assenov,C.Plass,R.Morales,J.Carles,D.Bowtell,L.Mileshkin,D.Sia,R.Tothill,J.Tabernero,J.M.Llovet,andM.Esteller.Epigenetic profiling to classify cancer of unknown primary : amulticentre, retrospective analysis. Lancet Oncol. 2016;17(10):1386-1395.
24.Koch,A.,S.C.Joosten,Z.Feng,T.C.de Ruijter,M.X.Draht,V.Melotte,K.M.Smits,J.Veeck,J.G.Herman,L.Van Neste,W.Van Criekinge,T.De Meyer,and M.vanEngeland.Analysis of DNA methylation in cancer:location revisited.Nat RevClin Oncol.2018;15(7):459-466.24.Koch,A.,S.C.Joosten,Z.Feng,T.C.de Ruijter,M.X.Draht,V.Melotte,K.M.Smits,J.Veeck,J.G.Herman,L.Van Neste,W.Van Criekinge,T.De Meyer, and M. van Engeland. Analysis of DNA methylation in cancer: location revisited. Nat RevClin Oncol. 2018;15(7):459-466.
25.Church,T.R.,M.Wandell,C.Lofton-Day,S.J.Mongin,M.Burger,S.R.Payne,E.Castanos-Velez,B.A.Blumenstein,T.Rosch,N.Osborn,D.Snover,R.W.Day,andD.F.Ransohoff.Prospective evaluation of methylated SEPT9 in plasma fordetection of asymptomatic colorectal cancer.Gut.2014;63(2):317-25.25.Church,T.R.,M.Wandell,C.Lofton-Day,S.J.Mongin,M.Burger,S.R.Payne,E.Castanos-Velez,B.A.Blumenstein,T.Rosch,N.Osborn,D.Snover,R.W.Day, and D.F.Ransohoff.Prospective evaluation of methylated SEPT9 in plasma for detection of asymptomatic colorectal cancer.Gut.2014;63(2):317-25.
26.Issa,I.A.and M.Noureddine.Colorectal cancer screening:An updatedreview of the available options.World J Gastroenterol.2017;23(28):5086-5096.26. Issa, I.A. and M. Noureddine. Colorectal cancer screening: An updated review of the available options. World J Gastroenterol. 2017;23(28):5086-5096.
27.Powrozek,T.,P.Krawczyk,T.Kucharczyk,and J.Milanowski.Septin9promoter region methylation in free circulating DNA-potential role innoninvasive diagnosis of lung cancer:preliminary report.Med Oncol.2014;31(4):917.27. Powrozek, T., P. Krawczyk, T. Kucharczyk, and J. Milanowski. Septin9promoter region methylation in free circulating DNA-potential role innoninvasive diagnosis of lung cancer:preliminary report.Med Oncol.2014;31(4):917 .
28.Schrock,A.,A.Leisse,L.de Vos,H.Gevensleben,F.Droge,A.Franzen,M.Wachendorfer,F.Schrock,J.Ellinger,M.Teschke,T.Wilhelm-Buchstab,J.Landsberg,S.Holdenrieder,G.Hartmann,J.K.Field,F.Bootz,G.Kristiansen,andD.Dietrich.Free-Circulating Methylated DNA in Blood for Diagnosis,Staging,Prognosis,and Monitoring of Head and Neck Squamous Cell Carcinoma Patients:AnObservational Prospective Cohort Study.Clin Chem.2017;63(7):1288-1296.28.Schrock,A.,A.Leisse,L.de Vos,H.Gevensleben,F.Droge,A.Franzen,M.Wachendorfer,F.Schrock,J.Ellinger,M.Teschke,T.Wilhelm-Buchstab, J. Landsberg, S. Holdenrieder, G. Hartmann, J. K. Field, F. Bootz, G. Kristiansen, and D. Dietrich. Free-Circulating Methylated DNA in Blood for Diagnosis, Staging, Prognosis, and Monitoring of Head and Neck Squamous Cell Carcinoma Patients: An Observational Prospective Cohort Study. Clin Chem. 2017;63(7):1288-1296.
29.Lee,H.S.,S.M.Hwang,T.S.Kim,D.W.Kim,D.J.Park,S.B.Kang,H.H.Kim,andK.U.Park.Circulating methylated septin 9 nucleic Acid in the plasma ofpatients with gastrointestinal cancer in the stomach and colon.TranslOncol.2013;6(3):290-6.29. Lee, H.S., S.M.Hwang, T.S.Kim, D.W.Kim, D.J.Park, S.B.Kang, H.H.Kim, and K.U.Park.Circulating methylated septin 9 nucleic Acid in the plasma of patients with gastrointestinal cancer in the stomach and colon.TranslOncol .2013;6(3):290-6.
30.Bettegowda,C.,M.Sausen,R.J.Leary,I.Kinde,Y.Wang,N.Agrawal,B.R.Bartlett,H.Wang,B.Luber,R.M.Alani,E.S.Antonarakis,N.S.Azad,A.Bardelli,H.Brem,J.L.Cameron,C.C.Lee,L.A.Fecher,G.L.Gallia,P.Gibbs,D.Le,R.L.Giuntoli,M.Goggins,M.D.Hogarty,M.Holdhoff,S.M.Hong,Y.Jiao,H.H.Juhl,J.J.Kim,G.Siravegna,D.A.Laheru,C.Lauricella,M.Lim,E.J.Lipson,S.K.Marie,G.J.Netto,K.S.Oliner,A.Olivi,L.Olsson,G.J.Riggins,A.Sartore-Bianchi,K.Schmidt,M.Shih l,S.M.Oba-Shinjo,S.Siena,D.Theodorescu,J.Tie,T.T.Harkins,S.Veronese,T.L.Wang,J.D.Weingart,C.L.Wolfgang,L.D.Wood,D.Xing,R.H.Hruban,J.Wu,P.J.Allen,C.M.Schmidt,M.A.Choti,V.E.Velculescu,K.W.Kinzler,B.Vogelstein,N.Papadopoulos,and L.A.Diaz,Jr.Detection of circulating tumor DNA in early-and late-stagehuman malignancies.Sci Transl Med.2014;6(224):224ra24.30.Bettegowda,C.,M.Sausen,R.J.Leary,I.Kinde,Y.Wang,N.Agrawal,B.R.Bartlett,H.Wang,B.Luber,R.M.Alani,E.S.Antonarakis,N.S.Azad,A.Bardelli, H.Brem,J.L.Cameron,C.C.Lee,L.A.Fecher,G.L.Gallia,P.Gibbs,D.Le,R.L.Giuntoli,M.Goggins,M.D.Hogarty,M.Holdhoff,S.M.Hong,Y.Jiao,H.H.Juhl,J.J.Kim , G.Siravegna,D.A.Laheru,C.Lauricella,M.Lim,E.J.Lipson,S.K.Marie,G.J.Netto,K.S.Oliner,A.Olivi,L.Olsson,G.J.Riggins,A.Sartore-Bianchi,K.Schmidt,M .Shih l,S.M.Oba-Shinjo,S.Siena,D.Theodorescu,J.Tie,T.T.Harkins,S.Veronese,T.L.Wang,J.D.Weingart,C.L.Wolfgang,L.D.Wood,D.Xing,R.H.Hruban,J.Wu , P.J.Allen,C.M.Schmidt,M.A.Choti,V.E.Velculescu,K.W.Kinzler,B.Vogelstein,N.Papadopoulos,and L.A.Diaz,Jr.Detection of circulating tumor DNA in early-and late-stagehuman malignancies.Sci Transl Med.2014; 6(224):224ra24.
31.Eads,C.A.,K.D.Danenberg,K.Kawakami,L.B.Saltz,C.Blake,D.Shibata,P.V.Danenberg,and P.W.Laird.MethyLight:a high-throughput assay to measure DNAmethylation.Nucleic Acids Res.2000;28(8):E32.31. Eads, C.A., K.D. Danenberg, K. Kawakami, L.B. Saltz, C. Blake, D. Shibata, P.V. Danenberg, and P.W. Laird. MethyLight: a high-throughput assay to measure DNA methylation. Nucleic Acids Res. 8):E32.
32.Yu,M.,T.J.Heinzerling,and W.M.Grady.DNA Methylation Analysis UsingDroplet Digital PCR.Methods Mol Biol.2018;1768:363-383.32. Yu, M., T.J. Heinzerling, and W.M.Grady. DNA Methylation Analysis Using Droplet Digital PCR. Methods Mol Biol. 2018;1768:363-383.
33.Vogelstein,B.and K.W.Kinzler.Digital PCR.Proc Natl Acad Sci U SA.1999;96(16):9236-41.33. Vogelstein, B. and K.W. Kinzler. Digital PCR. Proc Natl Acad Sci U SA. 1999;96(16):9236-41.
34.Van Wesenbeeck,L.,L.Janssens,H.Meeuws,O.Lagatie,andL.Stuyver.Droplet digital PCR is an accurate method to assess methylationstatus on FFPE samples.Epigenetics.2018;13(3):207-213.34. Van Wesenbeeck, L., L. Janssens, H. Meeuws, O. Lagatie, and L. Stuyver. Droplet digital PCR is an accurate method to assess methylationstatus on FFPE samples. Epigenetics. 2018;13(3):207-213 .
35.Hindson,B.J.,K.D.Ness,D.A.Masquelier,P.Belgrader,N.J.Heredia,A.J.Makarewicz,I.J.Bright,M.Y.Lucero,A.L.Hiddessen,T.C.Legler,T.K.Kitano,M.R.Hodel,J.F.Petersen,P.W.Wyatt,E.R.Steenblock,P.H.Shah,L.J.Bousse,C.B.Troup,J.C.Mellen,D.K.Wittmann,N.G.Erndt,T.H.Cauley,R.T.Koehler,A.P.So,S.Dube,K.A.Rose,L.Montesclaros,S.Wang,D.P.Stumbo,S.P.Hodges,S.Romine,F.P.Milanovich,H.E.White,J.F.Regan,G.A.Karlin-Neumann,C.M.Hindson,S.Saxonov,and B.W.Colston.High-throughput droplet digital PCR system for absolutequantitation of DNA copy number.Anal Chem.2011;83(22):8604-10.35.Hindson,B.J.,K.D.Ness,D.A.Masquelier,P.Belgrader,N.J.Heredia,A.J.Makarewicz,I.J.Bright,M.Y.Lucero,A.L.Hiddessen,T.C.Legler,T.K.Kitano,M.R.Hodel,J.F.Petersen,P.W.Wyatt,E.R.Steenblock,P.H.H. Shah,L.J.Bousse,C.B.Troup,J.C.Mellen,D.K.Wittmann,N.G.Erndt,T.H.Cauley,R.T.Koehler,A.P.So,S.Dube,K.A.Rose,L.Montesclaros,S.Wang,D.P.Stumbo,S.P.Hodges,S.Romine , F.P.Milanovich, H.E.White, J.F.Regan, G.A.Karlin-Neumann, C.M.Hindson, S.Saxonov, and B.W.Colston.High-throughput droplet digital PCR system for absolutequantitation of DNA copy number.Anal Chem.2011;83(22): 8604-10.
36.Yu,M.,K.T.Carter,K.W.Makar,K.Vickers,C.M.Ulrich,R.E.Schoen,D.Brenner,S.D.Markowitz,and W.M.Grady.MethyLight droplet digital PCR fordetection and absolute quantification of infrequently methylatedalleles.Epigenetics.2015;10(9):803-9.36. Yu,M.,K.T.Carter,K.W.Makar,K.Vickers,C.M.Ulrich,R.E.Schoen,D.Brenner,S.D.Markowitz,and W.M.Grady.MethyLight droplet digital PCR fordetection and absolute quantification of infrequently methylatedalleles.Epigenetics.2015; 10(9):803-9.
发明内容SUMMARY OF THE INVENTION
为了填补目前并没有一种能够实现特异性检测结直肠癌的甲基化标志物的空白,本发明的目的之一即在于提供可用于特异性检测结直肠癌的甲基化标志物。In order to fill the gap that there is currently no methylation marker that can specifically detect colorectal cancer, one of the objectives of the present invention is to provide a methylation marker that can be used to specifically detect colorectal cancer.
本发明的目的之二在于提供一种基于上述甲基化标志物建立的可用于检测结直肠癌的试剂盒。The second purpose of the present invention is to provide a kit for detecting colorectal cancer based on the above methylation markers.
为了达到上述目的,本发明采用了以下技术手段:In order to achieve the above object, the present invention has adopted the following technical means:
本发明利用TCGA等数据库分析结直肠癌组织特异性甲基化谱,筛选出肿瘤甲基化特异性标志物,主要筛选出分布在ZNF304,LIFR及C20orf194基因上的10个CpG位点。然后使用靶向测序MethylTarget技术在结直肠癌和癌旁正常组织中进行差异甲基化验证,并在组织结果的基础上提出了一种基于DNA甲基化特异性定量检测技术(QuantitativeMethylation Specific PCR,qMSP)和基于数字PCR技术检测人组织/血浆DNA中ZNF304和LIFR基因甲基化的方法及所用探针与引物,和已有的基于qPCR技术的人SEPT 9基因甲基化检测方法相比,本发明的检测方法具有更高的特异性。此外,数字PCR能够对待检测目的片段进行绝对定量,通过计算甲基化和非甲基化阳性液滴或者拷贝数的比值而得到待测样品中目的片段的甲基化水平,能够实现高灵敏度的检测。The invention utilizes databases such as TCGA to analyze the tissue-specific methylation profile of colorectal cancer, and screens out tumor methylation-specific markers, mainly screening out 10 CpG sites distributed on ZNF304, LIFR and C20orf194 genes. Then, the targeted sequencing MethylTarget technology was used to verify the differential methylation in colorectal cancer and adjacent normal tissues, and based on the tissue results, a DNA methylation-specific quantitative detection technology (QuantitativeMethylation Specific PCR, qMSP) and digital PCR-based methods for detecting ZNF304 and LIFR gene methylation in human tissue/plasma DNA and the probes and primers used, compared with the existing human SEPT 9 gene methylation detection methods based on qPCR technology, The detection method of the present invention has higher specificity. In addition, digital PCR can perform absolute quantification of the target fragment to be detected, and obtain the methylation level of the target fragment in the sample to be tested by calculating the ratio of methylation and non-methylation positive droplets or copy numbers, which can achieve high-sensitivity analysis. detection.
因此,本发明首先提出了一种与结直肠癌相关的甲基化标志物,所述的甲基化标志物为含有至少一个CpG甲基化位点的ZNF304,LIFR或C20orf194基因序列或其片段,或是所述的基因序列或其片段的任意组合;Therefore, the present invention first proposes a methylation marker related to colorectal cancer, the methylation marker is a ZNF304, LIFR or C20orf194 gene sequence or a fragment thereof containing at least one CpG methylation site , or any combination of described gene sequences or fragments thereof;
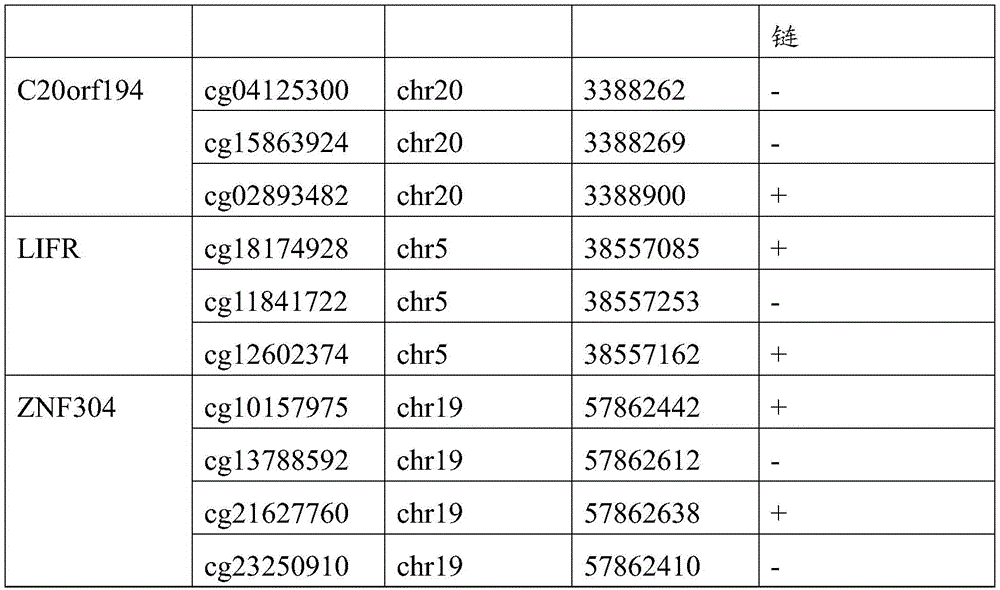
根据450K芯片的基因组注释信息,其中在C20orf194基因上存在3个CpG位点,分别为cg04125300、cg15863924以及cg02893482,cg04125300位于第20号染色体反链的第3388262核苷酸处,cg15863924位于第20号染色体反链的第3388269核苷酸处,cg02893482位于第20号染色体正链的第3388900核苷酸处;According to the genome annotation information of the 450K chip, there are 3 CpG sites on the C20orf194 gene, cg04125300, cg15863924 and cg02893482, cg04125300 is located at the 3388262 nucleotide of the anti-strand of
其中在LIFR基因上存在3个CpG位点,分别为cg18174928、cg11841722以及cg12602374,cg18174928位于第5号染色体正链的第38557085核苷酸处,cg11841722位于第5号染色体反链的第38557253核苷酸处,cg12602374位于第5号染色体正链的第38557162核苷酸处;Among them, there are three CpG sites on the LIFR gene, cg18174928, cg11841722 and cg12602374, cg18174928 is located at
其中在ZNF304基因上存在4个CpG位点,分别为cg10157975、cg13788592、cg21627760以及cg23250910,cg10157975位于第19号染色体正链的第57862442核苷酸处,cg13788592位于第19号染色体反链的第57862612核苷酸处,cg21627760位于第19号染色体正链的第57862638核苷酸处,cg23250910位于第19号染色体反链的第57862410核苷酸处。Among them, there are 4 CpG sites in ZNF304 gene, namely cg10157975, cg13788592, cg21627760 and cg23250910, cg10157975 is located at the 57862442 nucleotide of the positive strand of
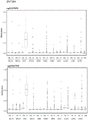
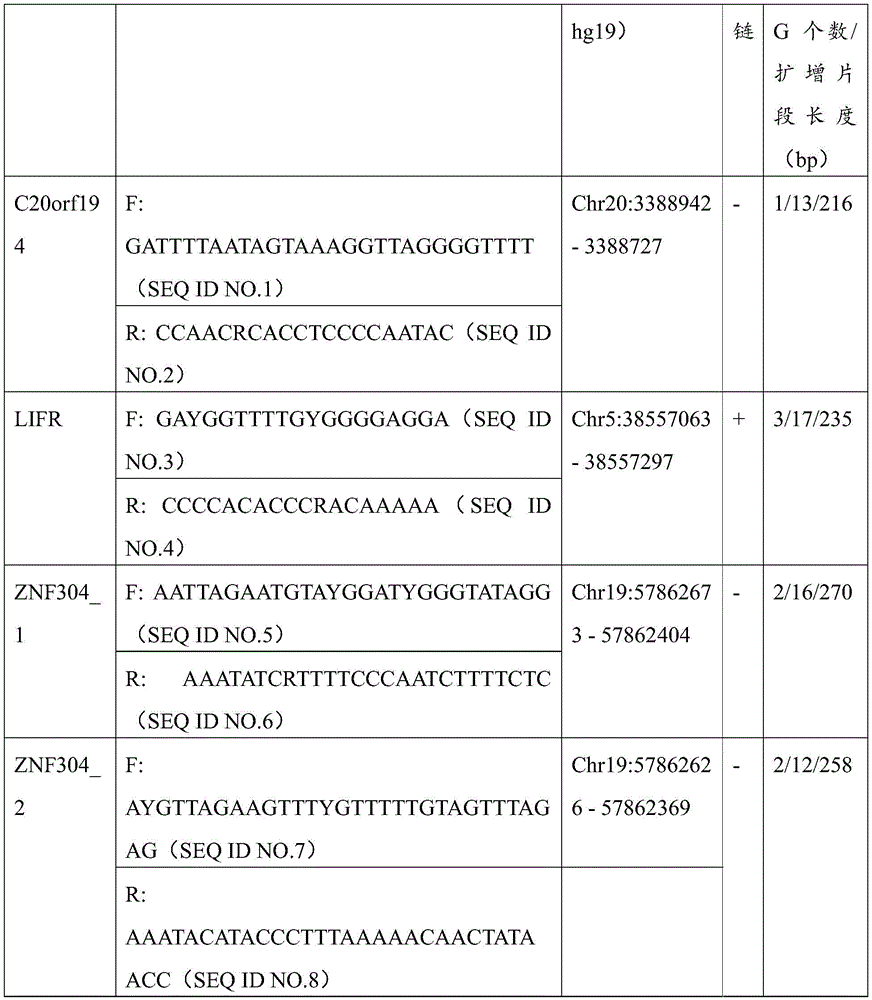
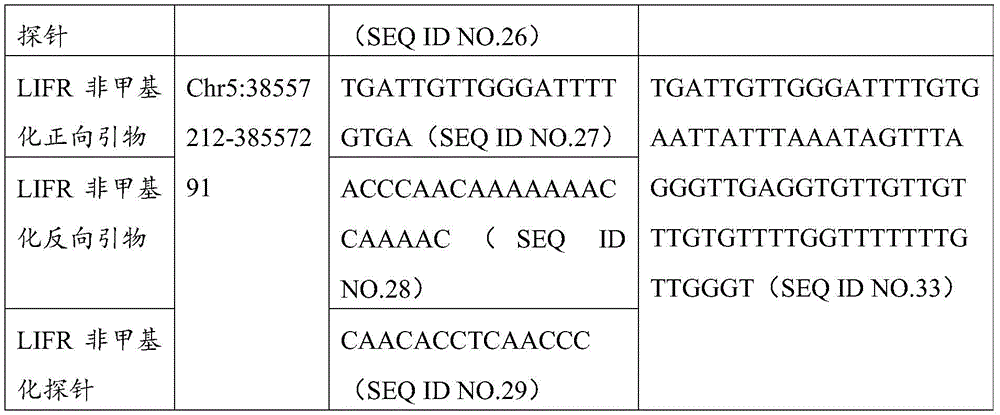
进一步的,本发明还提出了用于扩增所述的结直肠癌的甲基化标志物的引物组,所述的引物组由分别用于扩增含有所述CpG位点的ZNF304,LIFR及C20orf194基因片段的引物对组成,所述的引物序列如下所示:Further, the present invention also provides a primer set for amplifying the methylation marker of colorectal cancer, the primer set is respectively used to amplify ZNF304, LIFR and LIFR containing the CpG site. The primer pair composition of the C20orf194 gene fragment is as follows:
其中,ZNF304_1引物对用于扩增ZNF304基因上存在的2个CpG位点,分别为cg13788592以及cg21627760,cg13788592位于第19号染色体反链的第57862612核苷酸处,cg21627760位于第19号染色体正链的第57862638核苷酸处;ZNF304_2引物对用于扩增ZNF304基因上存在的另外2个CpG位点,分别为cg10157975以及cg23250910,cg10157975位于第19号染色体正链的第57862442核苷酸处,cg23250910位于第19号染色体反链的第57862410核苷酸处。Among them, the ZNF304_1 primer pair is used to amplify the two CpG sites present on the ZNF304 gene, cg13788592 and cg21627760 respectively, cg13788592 is located at the 57862612 nucleotide of the reverse strand of
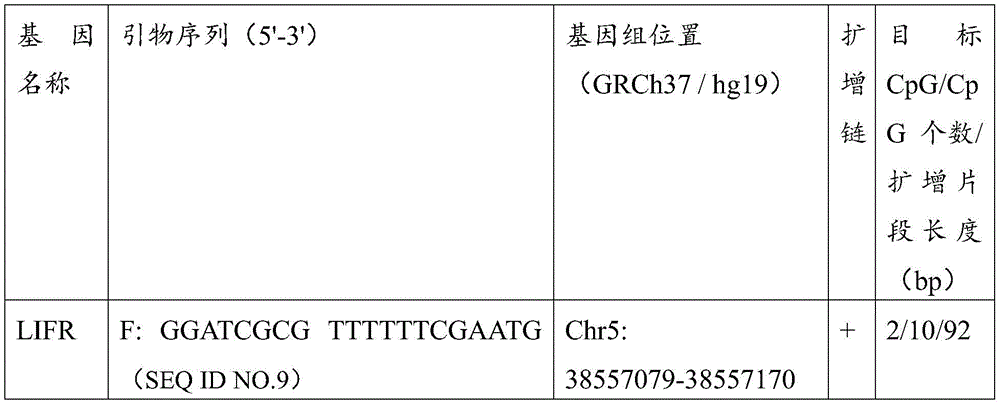
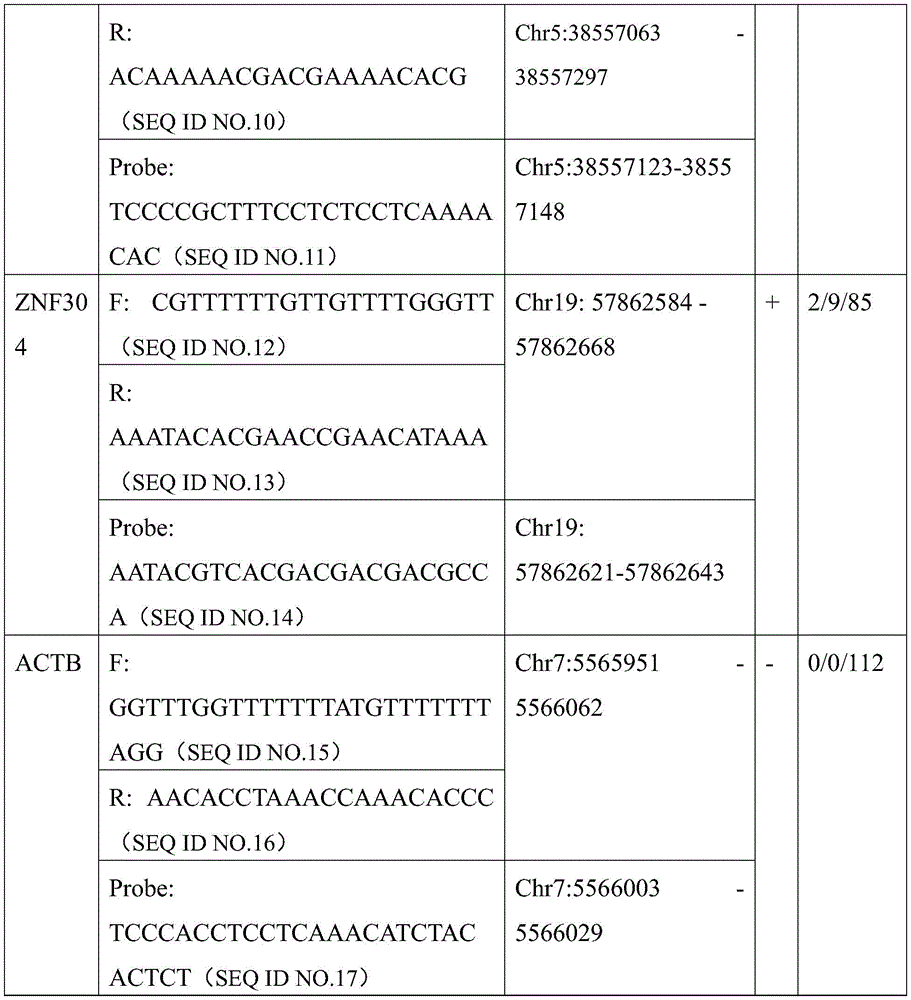
更进一步的,本发明还提出了一种用于结直肠癌相关基因甲基化检测的数字PCR试剂盒,所述试剂盒包括用于结直肠癌相关基因ZNF304以及LIFR基因甲基化检测的ddPCR引物探针组合,所述的ddPCR引物探针组合如下1)和/或2)所示:Further, the present invention also provides a digital PCR kit for colorectal cancer-related gene methylation detection, the kit includes ddPCR for colorectal cancer-related gene ZNF304 and LIFR gene methylation detection Primer-probe combination, described ddPCR primer-probe combination is shown in 1) and/or 2) below:
1)ZNF304甲基化检测的ddPCR引物及探针以及ZNF304非甲基化检测的ddPCR引物及探针,所述的引物及探针序列如下所示:1) ddPCR primers and probes for ZNF304 methylation detection and ddPCR primers and probes for ZNF304 non-methylation detection, described primers and probe sequences are as follows:
2)LIFR甲基化检测的ddPCR引物及探针以及LIFR非甲基化检测的PCR引物及探针,所述的引物及探针序列如下所示:2) ddPCR primers and probes for LIFR methylation detection and PCR primers and probes for LIFR non-methylation detection, described primers and probe sequences are as follows:
其中,优选的,所述探针的5’端包含有荧光报告基团,所述荧光报告基团包括FAM、HEX、NED、ROX、TET、JOE、TAMRA、CY3、CY5中的任意一种,且甲基化探针和非甲基化探针使用不同的荧光基团,所述荧光探针的3’端包含有荧光淬灭基团,所述荧光淬灭基团包括MGB、BHQ-1、BHQ-2、BHQ-3中的任意一种。Wherein, preferably, the 5' end of the probe contains a fluorescent reporter group, and the fluorescent reporter group includes any one of FAM, HEX, NED, ROX, TET, JOE, TAMRA, CY3, CY5, And methylated probe and non-methylated probe use different fluorescent groups, the 3' end of the fluorescent probe contains a fluorescence quenching group, and the fluorescence quenching group includes MGB, BHQ-1 , any one of BHQ-2 and BHQ-3.
其中,优选的,所述试剂盒的数字PCR反应体系和条件如下:Wherein, preferably, the digital PCR reaction system and conditions of the kit are as follows:
数字PCR反应体系:2×ddPCR Supermix for Probes 10uL,加入引物及探针使其终浓度分别达到引物终浓度为0.9uM以及探针终浓度为0.25uM,5-6uL浓度为0.002-70ng/uL的模板DNA,补足ddH2O至21uL;Digital PCR reaction system: 2×ddPCR Supermix for Probes 10uL, add primers and probes to make the final concentrations of primers and probes respectively reach the final concentration of 0.9uM and the final concentration of probes of 0.25uM, 5-6uL of 0.002-70ng/uL Template DNA, make up ddH2O to 21uL;
数字PCR反应条件如下:The digital PCR reaction conditions are as follows:
95℃预变性10分钟,94℃变性30秒,60℃退火60秒,共进行40个循环,98℃灭活酶10分钟,之后4℃保持;升降温速度≤2.5℃/s。Pre-denaturation at 95°C for 10 minutes, denaturation at 94°C for 30 seconds, annealing at 60°C for 60 seconds, a total of 40 cycles, inactivating the enzyme at 98°C for 10 minutes, and then holding at 4°C;
其中,优选的,所述的模板DNA为待测者血液中的DNA。Wherein, preferably, the template DNA is DNA in the blood of the test subject.
其中,优选的,所述的模板DNA为外周血血浆或血清中循环游离DNA(circulatingcellfreetumor DNA,cfDNA)、循环肿瘤DNA(circulating tumor DNA,ctDNA)或循环肿瘤细胞(circulating tumor cells,CTCs)DNA。Wherein, preferably, the template DNA is circulating cell free DNA (cfDNA), circulating tumor DNA (ctDNA) or circulating tumor cells (CTCs) DNA in peripheral blood plasma or serum.
更进一步的,本发明还提出了一种用于结直肠癌相关基因甲基化检测的qMSP试剂盒,所述试剂盒包括用于结直肠癌相关基因ZNF304以及LIFR基因甲基化检测的qMSP引物探针组合,所述的qMSP引物探针组合如下1)和/或2)所示:Further, the present invention also provides a qMSP kit for colorectal cancer-related gene methylation detection, the kit includes qMSP primers for colorectal cancer-related gene ZNF304 and LIFR gene methylation detection Probe combinations, the qMSP primer probe combinations are shown in the following 1) and/or 2):
1)ZNF304甲基化检测的qMSP引物及探针,所述的引物及探针序列如下所示:1) qMSP primers and probes for ZNF304 methylation detection, described primers and probe sequences are as follows:
ZNF304甲基化正向引物 5’-CGTTTTTTGTTGTTTTGGGTT-3’ZNF304 methylation forward primer 5’-CGTTTTTTGTTGTTTTGGGTT-3’
ZNF304甲基化反向引物 5’-AAATACACGAACCGAACATAAA-3’ZNF304 methylation reverse primer 5'-AAATACACGAACCGAACATAAA-3'
ZNF304甲基化探针 5’-AATACGTCACGACGACGACGCCA-3’ZNF304 methylation probe 5'-AATACGTCACGACGACGACGACCA-3'
2)LIFR甲基化检测的qMSP引物及探针,所述的引物及探针序列如下所示:2) qMSP primers and probes for LIFR methylation detection, described primers and probe sequences are as follows:
LIFR甲基化正向引物 5’-GGATCGCG TTTTTTCGAATG-3’LIFR methylation forward primer 5’-GGATCGCG TTTTTTCGAATG-3’
LIFR甲基化反向引物 5’-ACAAAAACGACGAAAACACG-3’LIFR methylation reverse primer 5'-ACAAAAACGACGAAAACACG-3'
LIFR甲基化探针 5’-TCCCCGCTTTCCTCTCCTCAAAACAC-3’LIFR methylation probe 5’-TCCCCGCTTTCCTCTCCTCAAAACAC-3’
其中,优选的,所述探针的5’端包含有荧光报告基团,所述荧光报告基团包括FAM、HEX、NED、ROX、TET、JOE、TAMRA、CY3、CY5中的任意一种,所述荧光探针的3’端包含有荧光淬灭基团,所述荧光淬灭基团包括MGB、BHQ-1、BHQ-2、BHQ-3中的任意一种。Wherein, preferably, the 5' end of the probe contains a fluorescent reporter group, and the fluorescent reporter group includes any one of FAM, HEX, NED, ROX, TET, JOE, TAMRA, CY3, CY5, The 3' end of the fluorescent probe contains a fluorescence quenching group, and the fluorescence quenching group includes any one of MGB, BHQ-1, BHQ-2, and BHQ-3.
其中,优选的,所述试剂盒的PCR反应体系和条件如下:Wherein, preferably, the PCR reaction system and conditions of the kit are as follows:
PCR反应扩增条件:95℃预变性10分钟,95℃变性10秒,60℃退火30秒,共进行45个循环,40℃冷却30秒,之后4℃保持。PCR amplification conditions: pre-denaturation at 95°C for 10 minutes, denaturation at 95°C for 10 seconds, annealing at 60°C for 30 seconds, a total of 45 cycles, cooling at 40°C for 30 seconds, and then maintained at 4°C.
相较于现有技术,本发明的有益效果是:Compared with the prior art, the beneficial effects of the present invention are:
1、本发明通过分析甲基化450K高通量芯片数据首次发现了分布在C20orf194、ZNF304、LIFR基因上的10个结直肠癌特异性的CpG位点,并使用靶向测序的方法在组织中对其中8个位点进行了验证;并成功建立了基于qMSP技术检测人组织ZNF304、LIFR基因甲基化的检测方法。1. The present invention discovered 10 colorectal cancer-specific CpG sites distributed on C20orf194, ZNF304, and LIFR genes for the first time by analyzing the
2、本发明首次在外周血DNA中检测了C20orf194、ZNF304、LIFR基因的甲基化水平,并首次使用数字PCR和MethyLight相结合的检测方法,通过优化数字PCR引物和探针,优化反应体系和反应条件,成功建立了基于ddPCR技术检测人ZNF304、LIFR基因甲基化的检测方法。和已有的基于qPCR技术的人SEPT9基因甲基化检测方法相比,本发明的检测方法具有更高的特异度,通过计算甲基化阳性液滴数和甲基化阳性液滴及非甲基化阳性液滴的比值进而实现绝对定量,可以对结直肠癌患者ZNF304、LIFR基因甲基化进行特异性定性和定量检测,从而实现对结直肠癌的辅助诊断。2. The present invention detects the methylation levels of C20orf194, ZNF304 and LIFR genes in peripheral blood DNA for the first time, and uses the detection method combining digital PCR and MethyLight for the first time. According to the reaction conditions, a detection method based on ddPCR technology to detect the methylation of human ZNF304 and LIFR genes was successfully established. Compared with the existing human SEPT9 gene methylation detection method based on qPCR technology, the detection method of the present invention has higher specificity. The ratio of the methylation-positive droplets can then achieve absolute quantification, which can specifically qualitatively and quantitatively detect the methylation of ZNF304 and LIFR genes in patients with colorectal cancer, thereby realizing the auxiliary diagnosis of colorectal cancer.
附图说明Description of drawings
图1为11种来源于TCGA数据库的癌症组织及来自于GEO的健康人外周血白细胞数据的平均甲基化水平;Figure 1 shows the average methylation levels of 11 cancer tissues from the TCGA database and the data of healthy human peripheral blood leukocytes from GEO;
图2A以及图2B为C20orf194基因所包含的位点甲基化水平箱式图;Figure 2A and Figure 2B are box plots of site methylation levels contained in the C20orf194 gene;
图3A以及图3B为LIFR基因所包含的位点甲基化水平箱式图;Figure 3A and Figure 3B are box plots of site methylation levels contained in the LIFR gene;
图4A以及图4B为ZNF304基因所包含的位点甲基化水平箱式图;Figure 4A and Figure 4B are box plots of site methylation levels contained in the ZNF304 gene;
图5为CRC癌和癌旁组织靶向测序差异性甲基化分析结果;Figure 5 shows the results of differential methylation analysis by targeted sequencing of CRC cancer and adjacent tissues;
图6为CRC癌、腺瘤、息肉组织和癌旁组织差异性甲基化分析结果;Figure 6 shows the results of differential methylation analysis of CRC cancer, adenoma, polyp tissue and paracancerous tissue;
图7A和图7B分别为扩增ZNF304基因以及基因LIFR的引物位置图;Fig. 7A and Fig. 7B are respectively the primer position map of amplifying ZNF304 gene and gene LIFR;
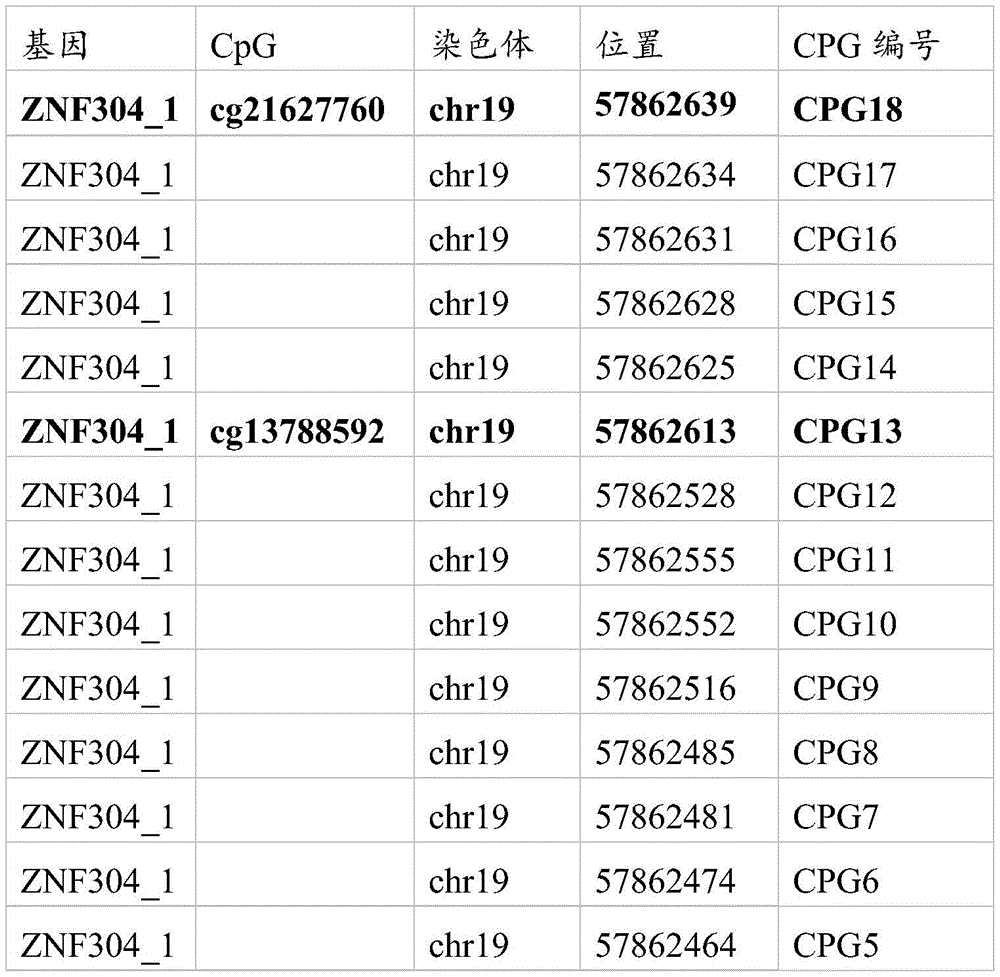
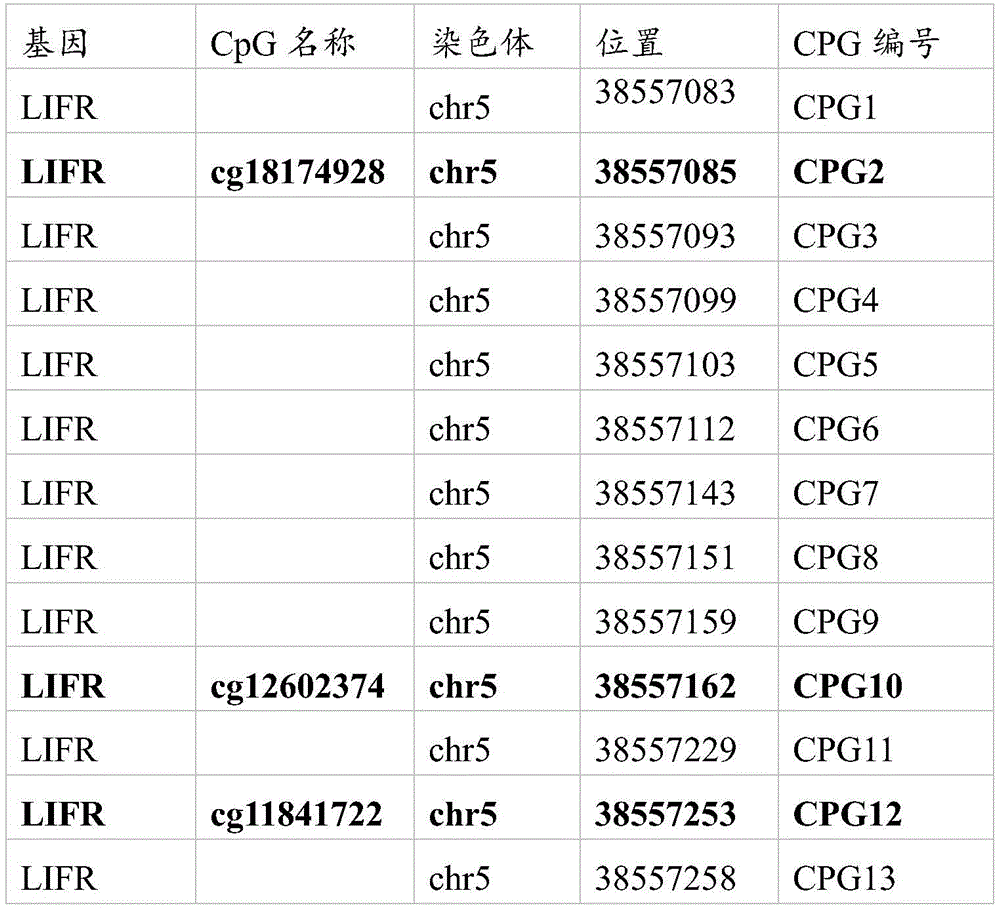
其中,图7A中各CPG的位置以及所对应的编号如下所示:Wherein, the position of each CPG in FIG. 7A and the corresponding number are as follows:
其中,图7B中各CPG的位置以及所对应的编号如下所示:Wherein, the position of each CPG in FIG. 7B and the corresponding number are as follows:
图8为ZNF304 qMSP扩增标准曲线;Figure 8 is the ZNF304 qMSP amplification standard curve;
图9为LIFR qMSP扩增标准曲线;Fig. 9 is LIFR qMSP amplification standard curve;
图10为ACTB qMSP扩增标准曲线;Figure 10 is the standard curve of ACTB qMSP amplification;
图11为qMSP扩增ZNF304甲基化水平的箱式散点图;Figure 11 is a box plot of qMSP amplification of ZNF304 methylation levels;
图12为qMSP扩增LIFR甲基化水平的箱式散点图;Figure 12 is a box plot of qMSP amplified LIFR methylation levels;
图13为ZNF304数字PCR质控品检测结果;Figure 13 is the test result of ZNF304 digital PCR quality control product;
图14为LIFR数字PCR质控品检测结果;Figure 14 shows the detection results of LIFR digital PCR quality control products;
图15为结直肠癌患者及健康对照血浆中ZNF304的甲基化水平箱式散点图;Figure 15 is a box plot of the methylation levels of ZNF304 in the plasma of colorectal cancer patients and healthy controls;
图16为结直肠癌患者及健康对照血浆中LIFR的甲基化水平箱式散点图。Figure 16 is a box plot of LIFR methylation levels in the plasma of colorectal cancer patients and healthy controls.
具体实施方式Detailed ways
以下通过具体实施例对本发明进行详细描述,以使本领域技术人员能够容易地根据本说明书的公开内容实施本发明。以下所描述的实施例仅是本发明的一部分实施例,而不是全部。基于本发明中的实施例,本领域普通技术人员在没有做出创造性劳动前提下所获得的所有其它实施例,都属于本发明保护的范围。The present invention will be described in detail below through specific embodiments, so that those skilled in the art can easily implement the present invention according to the disclosure of the present specification. The embodiments described below are only some, but not all, embodiments of the present invention. Based on the embodiments of the present invention, all other embodiments obtained by those of ordinary skill in the art without creative efforts shall fall within the protection scope of the present invention.
实施例1结直肠癌的cfDNA甲基化标志物的筛选及验证Example 1 Screening and verification of cfDNA methylation markers in colorectal cancer
1、结直肠癌组织特异性1. Colorectal cancer tissue specificity
本发明利用TCGA等数据库分析结直肠癌组织特异性甲基化谱,筛选出肿瘤甲基化特异性标志物,筛选出主要分布在ZNF304,LIFR及C20orf194基因上的10个CpG位点。基于TCGA数据库中包括结直肠癌在内的11种癌症组织及来自于GEO的健康人外周血白细胞的450K甲基化数据,表明10个CpG的结直肠癌组织特异性,平均甲基化水平见图1,三个基因所包含的位点甲基化水平箱式图见图2、图3、图4。10个CpG位点基因组信息见表1,癌症类型见表2。The present invention utilizes databases such as TCGA to analyze colorectal cancer tissue-specific methylation profiles, screen out tumor methylation-specific markers, and screen out 10 CpG sites mainly distributed on ZNF304, LIFR and C20orf194 genes. Based on the 450K methylation data of 11 cancer tissues including colorectal cancer in the TCGA database and healthy human peripheral blood leukocytes from GEO, the 10 CpGs were shown to be tissue-specific for colorectal cancer. The average methylation levels are shown in Figure 1. Box plots of site methylation levels included in the three genes are shown in Figure 2, Figure 3, and Figure 4. The genome information of the 10 CpG sites is shown in Table 1, and the cancer types are shown in Table 2.
表1Table 1
注:基因组注释信息取自450K芯片(GRCh37/hg19)Note: The genome annotation information is taken from the 450K chip (GRCh37/hg19)
表2Table 2
2、甲基化靶向测序2. Methylation-targeted sequencing
组织甲基化检测方法是结合目标区域富集和二代测序技术MethylTarget,凭借高深的测序深度实现多重PCR,能够准确计算每个CpG位点的甲基化水平。便于评价本发明所选取的10个CpG位点的甲基化水平,及差异甲基化水平。该技术准确性高,灵活性强,性价比优。The tissue methylation detection method is a combination of target region enrichment and next-generation sequencing technology MethylTarget, which realizes multiplex PCR with high sequencing depth and can accurately calculate the methylation level of each CpG site. It is convenient to evaluate the methylation level and differential methylation level of the 10 CpG sites selected in the present invention. The technology is highly accurate, flexible, and cost-effective.
本发明共纳入经病理组织学确诊原发结直肠癌患者276例,腺瘤组织8例,息肉组织10例,部分匹配的癌旁组织23例;使用酚-氯仿法和DNA Mini Kit(Qiagen,51306,德国)提取组织中的DNA,具体操作按照试剂盒操作手册进行。用Thermo ScientificNanoDrop 2000c分光光度计对提取的DNA进行定性和定量检测(A260/A280在1.7-1.9之间,A260/A230>2)。提取后DNA分装于-80℃保存。A total of 276 patients with primary colorectal cancer confirmed by histopathology were included in the present invention, including 8 adenoma tissues, 10 polyp tissues, and 23 partially matched adjacent tissues; the phenol-chloroform method and DNA Mini Kit (Qiagen, 51306, Germany) was used to extract DNA from tissues, and the specific operations were carried out according to the kit operation manual. The extracted DNA was qualitatively and quantitatively detected with a Thermo Scientific NanoDrop 2000c spectrophotometer (A260/A280 between 1.7-1.9, A260/A230>2). After extraction, the DNA was aliquoted and stored at -80°C.
使用Fast DNA Bisulfite Kit(Qiagen,59826,德国)对1-2ug原始DNA进行亚硫酸氢盐处理,具体操作按照试剂盒操作手册进行。硫化后DNA用NanoDrop 2000c进行浓度(ng/ul)和纯度测定,硫化后DNA序列中非甲基化的C被转化为U,因此大多数序列不再互补而成为单链,与RNA较为相似。因此测定模式为RNA,检测结果A260/280在2.0-2.4之间且A260/A230>1提示硫化后DNA质量合格。硫化后DNA稀释至20-25ng/uL分装后于-20℃保存。use The Fast DNA Bisulfite Kit (Qiagen, 59826, Germany) was subjected to bisulfite treatment on 1-2ug of the original DNA, and the specific operation was carried out according to the operation manual of the kit. The concentration (ng/ul) and purity of DNA after sulfidation were determined with NanoDrop 2000c. The unmethylated C in the sulfidized DNA sequence was converted to U, so most sequences were no longer complementary and became single-stranded, which was similar to RNA. Therefore, the assay mode is RNA, the detection result A260/280 is between 2.0-2.4 and A260/A230>1 indicates that the quality of DNA after sulfidation is qualified. After sulfuration, the DNA was diluted to 20-25ng/uL and stored at -20℃.
甲基化靶向测序实验步骤:Methylation-targeted sequencing experimental steps:
1)样品质控1) Sample quality control
硫化后DNA 浓度在20-25ng/uL之间,总量≥500ng,OD260/280=2.0-2.4,OD260/230≥1。The concentration of DNA after sulfuration is between 20-25ng/uL, the total amount is ≥500ng, OD260/280=2.0-2.4, OD260/230≥1.
2)引物设计与单位点PCR条件优化2) Primer design and optimization of single-site PCR conditions
使用primer3或自行调整针对目标CpG位点所在区域设计高质量的测序引物。在将序列导入软件之前要将目标序列手动转换为硫化后序列,即保持目标CpG的胞嘧啶不改变而其余CpG的C转换为U;挑选能够以经重亚硫酸盐处理后的人基因组为模板,扩增获得清晰单一条带的引物用于后续实验。C20orf194基因共包括3个CpG位点,引物预试成功一个位点(3388900);LIFR包含的3个CpG位点均预试成功;ZNF304包含4个CpG位点,分别设计在两条引物上。引物序列见表3。Use primer3 or self-adjust to design high-quality sequencing primers for the region where the target CpG site is located. Before importing the sequence into the software, the target sequence must be manually converted to the post-sulfuration sequence, that is, the cytosine of the target CpG is kept unchanged and the C of the remaining CpG is converted to U; the human genome after bisulfite treatment can be selected as a template. , and the primers that amplified a clear single band were used for subsequent experiments. The C20orf194 gene contains a total of 3 CpG sites, and one site (3388900) was successfully pre-tested by primers; the three CpG sites contained in LIFR were pre-tested successfully; ZNF304 contained four CpG sites, which were designed on two primers respectively. The primer sequences are shown in Table 3.
表3table 3
3)多重PCR引物panel优化3) Multiplex PCR primer panel optimization
将经步骤2)优化后的引物混合为多重PCR引物panel,以标准人基因组为模板进行扩增。基于毛细管电泳结果判断多重体系中每对引物是否高效、特异地进行扩增,并以此调整,优化多重PCR panel中的引物组成及浓度。The primers optimized in step 2) are mixed into a multiplex PCR primer panel, and the standard human genome is used as a template for amplification. Based on the results of capillary electrophoresis, determine whether each pair of primers in the multiplex system can efficiently and specifically amplify, and adjust accordingly to optimize the composition and concentration of primers in the multiplex PCR panel.
4)样本目标片段多重PCR反应4) Multiplex PCR reaction of sample target fragment
使用优化后的多重PCR引物panel,以硫化后DNA为模板,进行多重PCR扩增。经质控后,将以同一个样品基因组DNA为模板的所有多重PCR引物panel的扩增产物混合,并确保每个位点引物扩增产物的量相当,将混合后的多重PCR产物,各取2μL,稀释到60μL,作为后续Index PCR步骤的模板。多重PCR反应体系见表4和扩增条件见表5,总反应体系为10uL,余下用水补齐。The optimized multiplex PCR primer panel was used to carry out multiplex PCR amplification with the sulfurized DNA as the template. After quality control, mix the amplification products of all multiplex PCR primer panels using the same sample genomic DNA as the template, and ensure that the amount of each site primer amplification product is the same. 2 μL, diluted to 60 μL, as a template for the subsequent Index PCR step. The multiplex PCR reaction system is shown in Table 4 and the amplification conditions are shown in Table 5. The total reaction system is 10uL, and the rest is filled with water.
表4多重反应体系Table 4 Multiple reaction systems
表5多重反应条件Table 5 Multiple reaction conditions
5)样本添加特异性标签序列5) Add a specific tag sequence to the sample
利用带有Index序列的引物,通过PCR扩增向上述步骤扩增后建立的文库末端引入和illumina平台兼容的特异性标签序列。反应采用12个循环数的PCR程序,尽可能降低PCR偏倚。IndexPCR体系及扩增条件见表6,表7。总反应体系为10uL,余下用水补齐。Using primers with Index sequences, specific tag sequences compatible with the illumina platform were introduced into the end of the library established after the amplification in the above steps by PCR amplification. The reaction used a PCR program of 12 cycles to minimize PCR bias. The IndexPCR system and amplification conditions are shown in Table 6 and Table 7. The total reaction system is 10uL, and the rest is made up with water.
表6Table 6
表7Table 7
6)定量后上机测序6) On-board sequencing after quantification
将所有样品Index PCR扩增产物等量混合,并经割胶回收获得最终的MethylTarget测序文库,文库的片段长度分布经Agilent 2100Bioanalyzer验证。文库摩尔浓度精确定量后,最终于Illumina Hiseq平台,以2×150bp的双端测序模式进行高通量测序,获得FastQ数据。The Index PCR amplification products of all samples were mixed in equal amounts, and recovered by gel tapping to obtain the final MethylTarget sequencing library. The fragment length distribution of the library was verified by the Agilent 2100 Bioanalyzer. After the library molar concentration was accurately quantified, the Illumina Hiseq platform was finally used for high-throughput sequencing in a 2×150bp paired-end sequencing mode to obtain FastQ data.
3、甲基化靶向测序数据分析3. Methylation-targeted sequencing data analysis
首先根据目标区域的基因组位置核对引物扩增的区域在参考基因组上的注释信息。其次对测序原始数据整理及质量评估,及对胞嘧啶转化效率统计以评价硫化转化效率。最后计算各CpG位点甲基化水平。Firstly, the annotation information of the region amplified by the primers on the reference genome is checked according to the genomic location of the target region. Secondly, the sequencing raw data is sorted and the quality is evaluated, and the cytosine conversion efficiency is counted to evaluate the sulfur conversion efficiency. Finally, the methylation level of each CpG site was calculated.
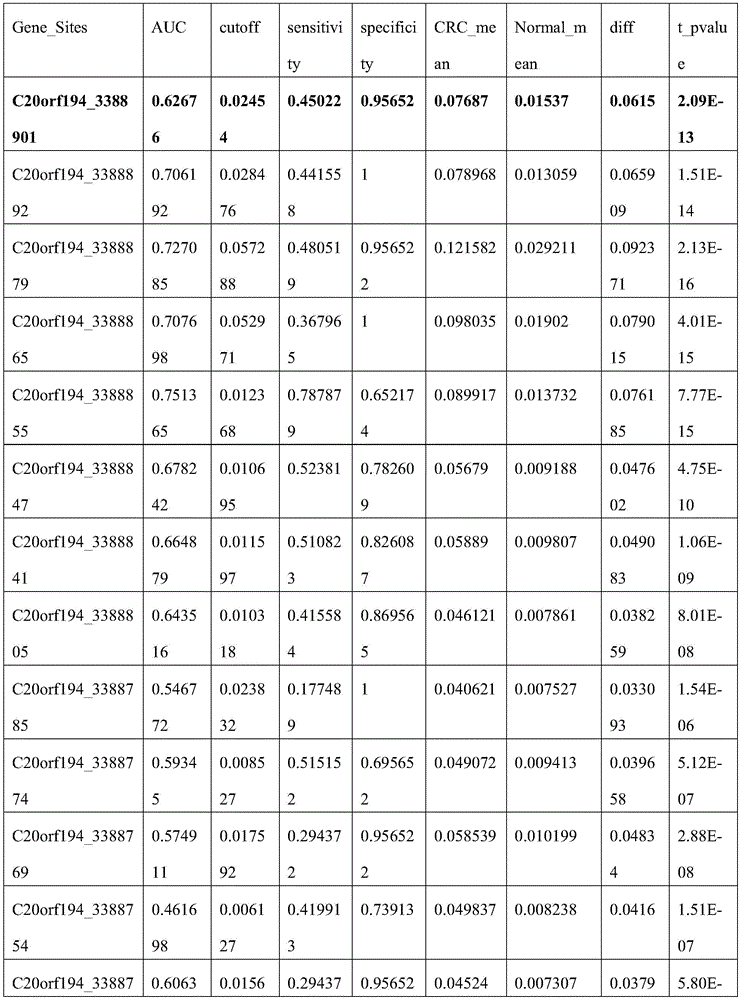
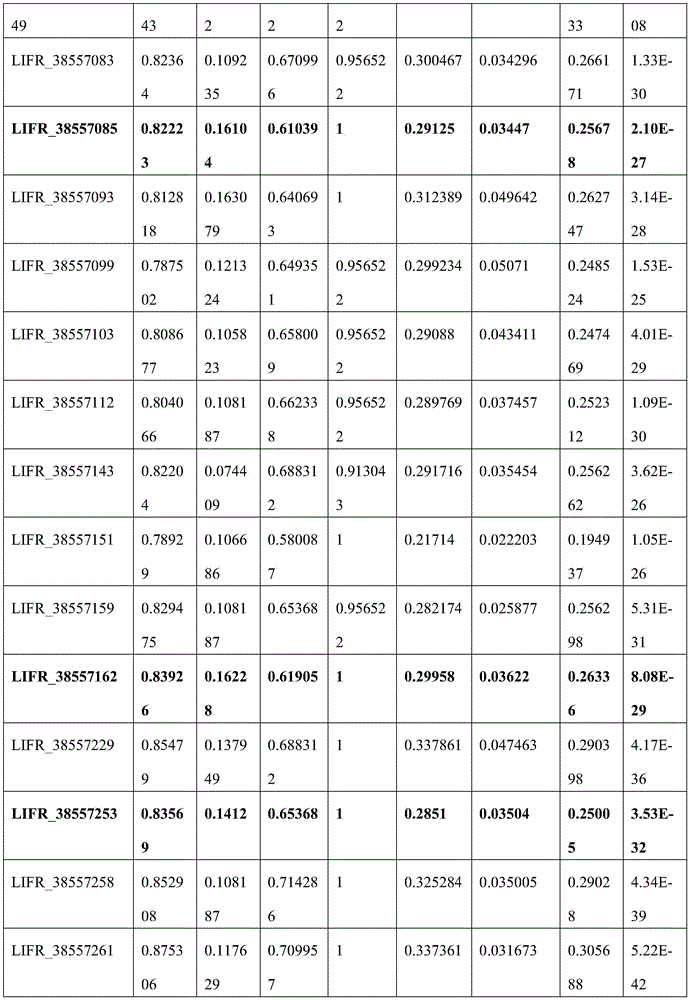
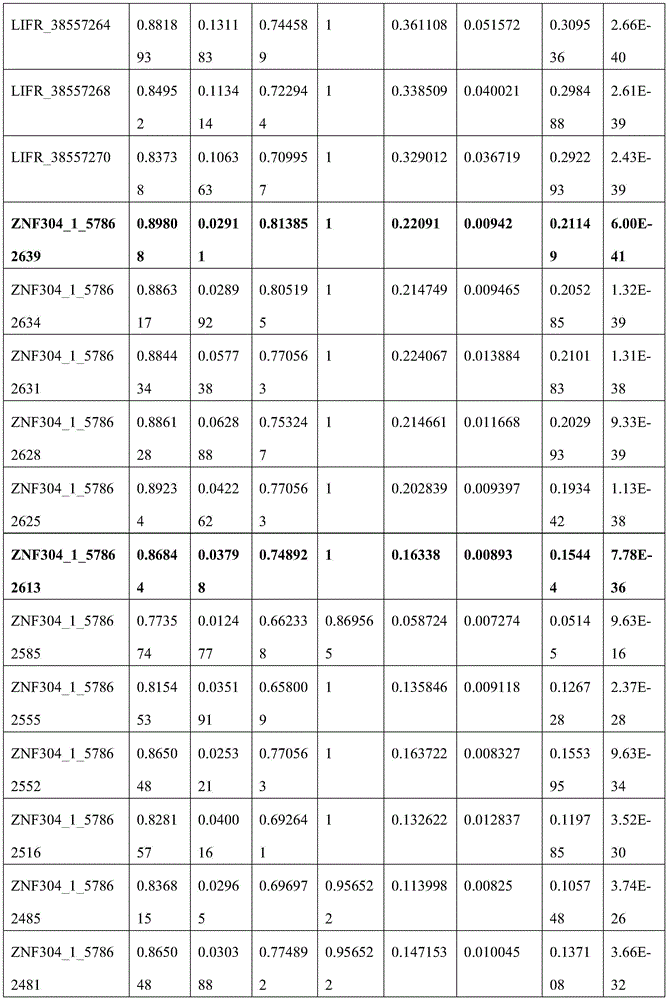
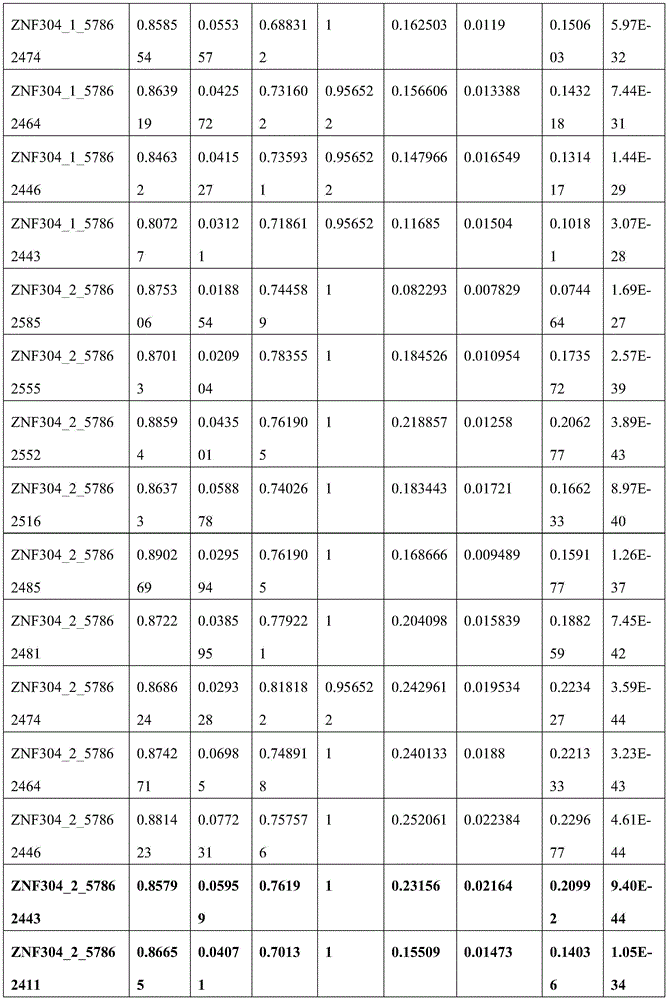
C20orf194,ZNF304_1,ZNF304_2均设计在反链上,因此基因位点的位置相比450K注释信息增加一位。剔除胞嘧啶转化效率<98%的3个样本以保证完全硫化转化,剔除测序深度低于10×的42个样本以保证数据质量,剩余272例样本数据。CRC癌和癌旁差异性甲基化分析结果见图5,所筛选的位点及其周围临近位点均显示癌组织甲基化水平显著高于癌旁组织,AUC的范围为46.17%-89.81%,结果见表8(目标位点加粗);且数据结果表明所筛选的基因可能在腺瘤性息肉阶段甲基化水平就有升高趋势(见图6或表9)。基于外部数据和组织验证结果,说明本发明筛选的得到的10个CpG甲基化位点能够作为特异性检测结直肠癌的甲基化位点,而含有上述CpG甲基化位点的核苷酸序列可以作为特异性检测结直肠癌的甲基化标志物。C20orf194, ZNF304_1, ZNF304_2 are all designed on the reverse strand, so the position of the gene locus is increased by one bit compared to the 450K annotation information. Three samples with cytosine conversion efficiency <98% were excluded to ensure complete sulfuration conversion, 42 samples with sequencing depth less than 10× were excluded to ensure data quality, and the remaining 272 samples were data. The results of differential methylation analysis between CRC cancer and adjacent cancer are shown in Figure 5. The selected sites and their surrounding adjacent sites all show that the methylation level of cancer tissue is significantly higher than that of adjacent cancer tissue, and the AUC range is 46.17%-89.81 %, the results are shown in Table 8 (target sites are in bold); and the data results indicate that the screened genes may have an increasing trend in the methylation level at the stage of adenomatous polyps (see Figure 6 or Table 9). Based on external data and tissue verification results, it shows that the 10 CpG methylation sites screened in the present invention can be used as methylation sites for the specific detection of colorectal cancer, and the nucleosides containing the above CpG methylation sites Acid sequences can be used as methylation markers for the specific detection of colorectal cancer.
表8 MethylTarget_AUCTable 8 MethylTarget_AUC
表9 MethylTarget_methylationTable 9 MethylTarget_methylation
实施例2甲基化特异性荧光定量PCR(qMSP)Example 2 Methylation-specific fluorescence quantitative PCR (qMSP)
使用qMSP探针法对结直肠癌和癌旁组织及外周血白细胞DNA进行甲基化检测,在靶向测序结果的基础上设计引物。以经过亚硫酸氢盐处理过的全甲基化人类基因组DNA为模板构建目标片段质粒,梯度稀释构建甲基化标准曲线并建立方程,以ACTB上无CG位点序列为内参基因,样本目的基因扩增拷贝数和内参基因扩增拷贝数的比值作为目标基因相对甲基化水平。The qMSP probe method was used for methylation detection of colorectal cancer, adjacent tissue and peripheral blood leukocyte DNA, and primers were designed based on the results of targeted sequencing. The target fragment plasmid was constructed with the fully methylated human genomic DNA treated with bisulfite as the template, and the methylation standard curve was constructed by gradient dilution and the equation was established. The ratio of the amplified copy number to that of the reference gene was used as the relative methylation level of the target gene.
1.引物及探针设计1. Primer and probe design
针对所筛选位点设计ZNF304、LIFR及内参基因ACTB的qMSP引物和探针,扩增引物包括甲基化扩增引物及内参扩增引物,qMSP探针包括甲基化探针及内参探针;内参引物设计在ACTB基因上无CG序列,防止对甲基化检测有干扰。探针的荧光基团如FAM、VIC等具体参照荧光定量PCR仪器荧光范围,猝灭基团可以选择能够增加探针退火温度的MGB基团或其他;挑选能够在阳性质控品即经过亚硫酸氢盐处理过的全甲基化人类基因组DNA为模板的体系中扩增获得清晰单一光滑的“S”型扩增曲线的引物探针组合用于后续实验,而在去RNA酶水或空白对照NTC中无任何荧光信号。反应引物及探针序列见表10。引物位置如图7所示。Design qMSP primers and probes for ZNF304, LIFR and internal reference gene ACTB for the screened sites, amplification primers include methylation amplification primers and internal reference amplification primers, qMSP probes include methylation probes and internal reference probes; The internal reference primer was designed without CG sequence on the ACTB gene to prevent interference with methylation detection. The fluorescent groups of the probe, such as FAM, VIC, etc., refer to the fluorescence range of the fluorescence quantitative PCR instrument. The quenching group can be selected as the MGB group or others that can increase the annealing temperature of the probe; The primer-probe combination that obtained a clear, single and smooth "S"-shaped amplification curve was amplified in a system using hydrogen-salt-treated fully methylated human genomic DNA as a template for subsequent experiments. There was no fluorescent signal in NTC. The reaction primers and probe sequences are shown in Table 10. The primer positions are shown in Figure 7.
表10引物及探针序列Table 10 Primer and probe sequences
2.DNA样本的提取和制备2. Extraction and Preparation of DNA Samples
本发明共纳入经病理组织学确诊原发结直肠癌(CRC)患者90例,部分匹配的癌旁组织(Normal)41例,结直肠癌患者外周血白细胞(CRC_WB)52例,健康对照外周血白细胞(WB)50例;使用DNA Mini Kit(Qiagen,#51306,德国)提取样本中的DNA,具体操作按照试剂盒操作手册进行。用Thermo Scientific NanoDrop 2000c分光光度计对提取的DNA进行定性和定量检测(A260/A280在1.7-1.9之间,A260/A230>2)。提取后DNA分装于-80℃保存。A total of 90 cases of primary colorectal cancer (CRC) patients confirmed by histopathology, 41 cases of partially matched adjacent tissue (Normal), 52 cases of peripheral blood white blood cells (CRC_WB) of colorectal cancer patients, and peripheral blood of healthy controls were included in the present invention. White blood cells (WB) 50 cases; use DNA Mini Kit (Qiagen, #51306, Germany) was used to extract DNA from samples, and the specific operations were carried out according to the kit's operating manual. The extracted DNA was qualitatively and quantitatively detected with a Thermo Scientific NanoDrop 2000c spectrophotometer (A260/A280 between 1.7-1.9, A260/A230>2). After extraction, the DNA was aliquoted and stored at -80°C.
使用Fast DNA Bisulfite Kit(Qiagen,#59826,德国)对1-2ug原始DNA进行亚硫酸氢盐处理,具体操作按照试剂盒操作手册进行。硫化后DNA用NanoDrop2000c进行浓度(ng/ul)和纯度测定,硫化后DNA序列中非甲基化的C被转化为U,因此大多数序列不再互补而成为单链,与RNA较为相似。因此测定模式为RNA,检测结果A260/280在2.0-2.4之间且A260/A230>1提示硫化后DNA质量合格。硫化后DNA稀释至20-25ng/uL分装后于-20℃保存。use The Fast DNA Bisulfite Kit (Qiagen, #59826, Germany) bisulfite 1-2ug of the original DNA, and the specific operation was carried out according to the kit's operating manual. The concentration (ng/ul) and purity of DNA after sulfidation were determined with NanoDrop2000c. The unmethylated C in the sulfidized DNA sequence was converted to U, so most sequences were no longer complementary and became single-stranded, which was similar to RNA. Therefore, the assay mode is RNA, the detection result A260/280 is between 2.0-2.4 and A260/A230>1 indicates that the quality of DNA after sulfidation is qualified. After sulfuration, the DNA was diluted to 20-25ng/uL and stored at -20℃.
3、人ZNF304、LIFR甲基化荧光定量PCR检测体系3. Human ZNF304, LIFR methylation fluorescent quantitative PCR detection system
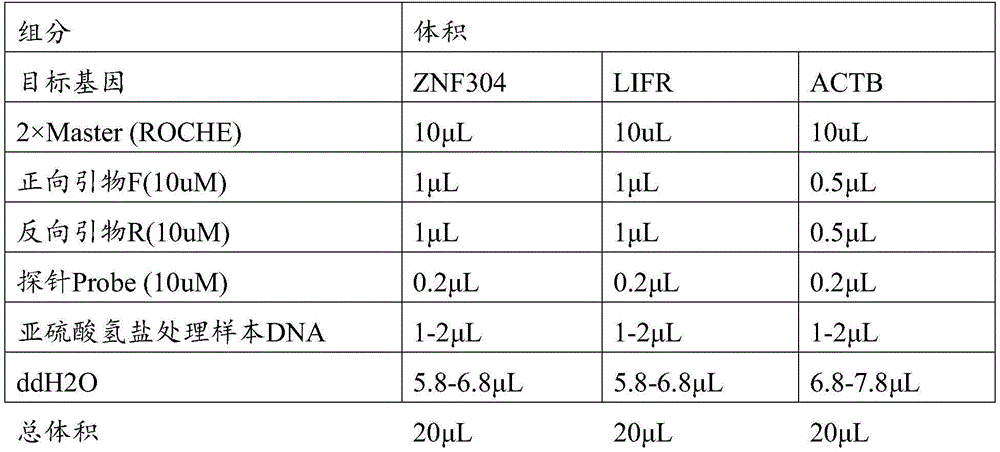
反应体系包括480Probes Master PCR预混液及目的基因引物探针组合和模板见表11。使用480Software release 1.5.0设定PCR反应扩增条件:95℃预变性10分钟,95℃变性10秒,60℃退火30秒,共进行45个循环,40℃冷却30秒,之后4℃保持。PCR完成后使用Abs Quant/2nd Derivative Max模块进行分析,将样本Cp值带入目标基因和内参基因标准曲线,计算相对甲基化水平。The reaction system includes The 480Probes Master PCR master mix and target gene primer probe combinations and templates are shown in Table 11. use 480Software release 1.5.0 sets PCR amplification conditions: 95°C pre-denaturation for 10 minutes, 95°C denaturation for 10 seconds, 60°C annealing for 30 seconds, a total of 45 cycles, 40°C cooling for 30 seconds, and then 4°C holding. After the PCR is completed, the Abs Quant/2nd Derivative Max module is used for analysis, and the Cp value of the sample is brought into the standard curve of the target gene and the internal reference gene to calculate the relative methylation level.
表11Table 11
4、检测体系有效性及结果的判断标准4. Judgment criteria for the effectiveness of the testing system and results
(1)ZNF304检测体系有效性及结果的判断:(1) Judgment of the effectiveness and results of the ZNF304 testing system:
a)有效性判定:梯度稀释质粒浓度范围为5E7-5E1 copies/ul,均有扩增信号且扩增曲线为光滑的“S”型,去RNA酶水/NTC无扩增信号或无“S”型,判断有效。否则重新配置。a) Judgment of validity: the gradient dilution plasmid concentration range is 5E7-5E1 copies/ul, there are amplification signals and the amplification curve is smooth "S" shape, RNase-free water/NTC has no amplification signal or no "S" "type, the judgment is valid. Otherwise reconfigure.
b)结果判定:使用标准梯度稀释质粒的检测Cp值和经对数转换的浓度构建标准曲线,标准曲线的扩增效率在1.9-2.1之间(90%-110%)。样本Cp值在标准曲线范围内可纳入标准曲线计算拷贝数;样本Cp大于最低浓度质粒Cp值,提示人ZNF304基因甲基化阴性或甲基化程度低于最低检出限。ZNF304扩增标准曲线见图8。b) Result judgment: The standard curve was constructed using the detection Cp value of the standard gradient dilution plasmid and the log-transformed concentration, and the amplification efficiency of the standard curve was between 1.9-2.1 (90%-110%). If the Cp value of the sample is within the range of the standard curve, it can be included in the standard curve to calculate the copy number; if the Cp value of the sample is greater than the Cp value of the lowest concentration plasmid, it indicates that the methylation of the human ZNF304 gene is negative or the degree of methylation is lower than the minimum detection limit. The ZNF304 amplification standard curve is shown in Figure 8.
(2)LIFR检测体系有效性及结果的判断:(2) Judgment of the effectiveness and results of the LIFR testing system:
a)有效性判定:梯度稀释质粒浓度范围为5E7-5E1 copies/ul,均有扩增信号且扩增曲线为光滑的“S”型,去RNA酶水/NTC无扩增信号或无“S”型,判断有效。否则重新配置。a) Judgment of validity: the gradient dilution plasmid concentration range is 5E7-5E1 copies/ul, there are amplification signals and the amplification curve is smooth "S" shape, RNase-free water/NTC has no amplification signal or no "S" "type, the judgment is valid. Otherwise reconfigure.
b)结果判定:使用标准梯度稀释质粒的检测Cp值和经对数转换的浓度构建标准曲线,标准曲线的扩增效率在1.9-2.1之间(90%-110%)。样本Cp值在标准曲线范围内可纳入标准曲线计算拷贝数;样本Cp大于最低浓度质粒Cp值,提示人LIFR基因甲基化阴性或甲基化程度低于最低检出限。LIFR扩增标准曲线见图9。b) Result judgment: The standard curve was constructed using the detection Cp value of the standard gradient dilution plasmid and the log-transformed concentration, and the amplification efficiency of the standard curve was between 1.9-2.1 (90%-110%). The sample Cp value within the range of the standard curve can be included in the standard curve to calculate the copy number; the sample Cp value is greater than the lowest concentration plasmid Cp value, indicating that the human LIFR gene methylation is negative or the methylation degree is lower than the minimum detection limit. The LIFR amplification standard curve is shown in Figure 9.
(3)ACTB检测体系有效性及结果的判断:(3) Judgment of the validity and results of the ACTB testing system:
a)有效性判定:梯度稀释质粒浓度范围为5E7-5E1 copies/ul,其中5个连续浓度有扩增信号且扩增曲线为光滑的“S”型,去RNA酶水/NTC无扩增信号或无“S”型,判断有效。否则重新配置。a) Validity determination: The concentration range of the gradient dilution plasmid is 5E7-5E1 copies/ul, of which 5 consecutive concentrations have amplification signals and the amplification curve is a smooth "S" shape, and there is no amplification signal in RNase-free water/NTC If there is no "S" type, the judgment is valid. Otherwise reconfigure.
b)结果判定:使用标准梯度稀释质粒的检测Cp值和经对数转换的浓度构建标准曲线,标准曲线的扩增效率在1.9-2.1之间(90%-110%)。样本Cp值在标准曲线范围内可纳入标准曲线计算拷贝数;样本Cp大于最低浓度质粒Cp值,提示人ACTB基因阴性或低于最低检出限。ACTB扩增标准曲线见图10。b) Result judgment: The standard curve was constructed using the detection Cp value of the standard gradient dilution plasmid and the log-transformed concentration, and the amplification efficiency of the standard curve was between 1.9-2.1 (90%-110%). If the Cp value of the sample is within the range of the standard curve, it can be included in the standard curve to calculate the copy number; if the Cp value of the sample is greater than the Cp value of the lowest concentration plasmid, it indicates that the human ACTB gene is negative or lower than the minimum detection limit. The ACTB amplification standard curve is shown in Figure 10.
5、检测结果及分析5. Test results and analysis
本发明共检测原发性结直肠癌90例,部分匹配的癌旁组织41例,结直肠癌患者外周血白细胞52例,健康对照外周血白细胞50例;将样本Cp值带入标准曲线计算相对甲基化水平。结果显示ZNF304、LIFR在结直肠癌组织中呈现高甲基化状态而在癌旁及外周血白细胞中呈较低甲基化水平,表明外周血白细胞DNA甲基化水平不会对后续cfDNA中两基因检测有干扰,ZNF304、LIFR箱式散点图分别见图11、图12。ZNF304甲基化水平检测结直肠癌组织的灵敏度为77.8%,特异度为97.6%,受试者曲线下面积为0.812;LIFR甲基化水平检测结直肠癌组织的灵敏度为73.3%,特异度为90.2%,受试者曲线下面积为0.806。使用ROC曲线计算两基因最佳cut-off值,分别在不同分期中计算两基因及其联合阳性率,结果见表12、表13、表14。The invention detects a total of 90 cases of primary colorectal cancer, 41 cases of partially matched adjacent tissues, 52 cases of peripheral blood leukocytes of colorectal cancer patients, and 50 cases of peripheral blood leukocytes of healthy controls; the sample Cp value is brought into the standard curve to calculate the relative methylation level. The results showed that ZNF304 and LIFR were hypermethylated in colorectal cancer tissues, but showed lower methylation levels in paracancerous and peripheral blood leukocytes, indicating that peripheral blood leukocyte DNA methylation levels did not affect the subsequent detection of the two genes in cfDNA. Interference, ZNF304 and LIFR box scatter plots are shown in Figure 11 and Figure 12 respectively. The sensitivity of ZNF304 methylation level in detecting colorectal cancer tissue was 77.8%, the specificity was 97.6%, and the area under the subject curve was 0.812; the sensitivity of LIFR methylation level in detecting colorectal cancer tissue was 73.3%, the specificity was 0.812 90.2% and the subject area under the curve was 0.806. The ROC curve was used to calculate the best cut-off value of the two genes, and the two genes and their combined positive rates were calculated in different stages. The results are shown in Table 12, Table 13, and Table 14.
表12Table 12
表13Table 13
表14Table 14
实施例3 MethyLight ddPCR检测方法以及试剂盒的建立Example 3 MethyLight ddPCR detection method and establishment of the kit
本发明所采用的技术是数字PCR技术结合MethyLight法,通过设计特异性的人ZNF304、LIFR候选甲基化区域的引物和探针,扩增经亚硫酸氢盐修饰并经微滴化制备的待测DNA样本,根据PCR扩增的阳性微滴数及与总阳性液滴数的比值来确定待测样本中人ZNF304、LIFR候选甲基化区域是否发生了甲基化以及甲基化水平,从而判断待测样本是否发生结直肠癌。The technology used in the present invention is digital PCR technology combined with the MethyLight method. By designing specific primers and probes of the candidate methylation regions of human ZNF304 and LIFR, amplifying the bisulfite modified and prepared by microdroplet DNA samples were tested, and the ratio of the number of positive droplets amplified by PCR to the total number of positive droplets was used to determine whether the methylation and methylation levels of the candidate methylation regions of human ZNF304 and LIFR in the sample to be tested had occurred. Determine whether the sample to be tested has colorectal cancer.
1、引物及探针设计1. Primer and probe design
针对所筛选位点设计ZNF304和LIFR的ddPCR引物和探针,扩增引物包括甲基化序列扩增引物及非甲基化序列扩增引物,TaqMan探针包括甲基化探针和非甲基化探针,为连接报告荧光基团和淬灭基团的核苷酸,甲基化探针和非甲基化探针应使用不同的荧光基团如FAM、VIC等具体参照数字PCR机器荧光范围,猝灭基团可以选择能够增加探针退火温度的MGB基团,或者使用锁式探针;ddPCR反应引物及探针序列见表15。Design ddPCR primers and probes for ZNF304 and LIFR for the screened sites, amplification primers include methylated sequence amplification primers and non-methylated sequence amplification primers, TaqMan probes include methylated probes and non-methylated sequence amplification primers The methylated probe, in order to connect the nucleotides of the reporter fluorescent group and the quenching group, the methylated probe and the unmethylated probe should use different fluorescent groups such as FAM, VIC, etc. Refer to the digital PCR machine for specific fluorescence The quenching group can choose the MGB group that can increase the annealing temperature of the probe, or use a padlock probe; the ddPCR reaction primers and probe sequences are shown in Table 15.
表15Table 15
在本发明的示例性实施例中,数字PCR采用微滴PCR技术(droplet digital,PCR)。其主要特征是将数字PCR混合液加入微滴发生器,生成10000~20000个微反应液滴后进行PCR扩增。在每次的PCR布板中要同时包括阳性质控品(M)、阴性质控品(U)、去RNA酶水和空白对照NTC,所谓的阳性质控品可以是经过亚硫酸氢盐处理过的全甲基化人类基因组DNA,而阴性质控品可以是经过硫化的非甲基化人类基因组DNA。根据质控品和空白对照设定阈值判断待测样品中是否含有发生甲基化的人ZNF304、LIFR基因的DNA目标分子,同时确定发生甲基化的人ZNF304、LIFR的目标DNA分子的数量。通过相应的已经商品化的软件(如QuantaSoft,Bio-Rad)进行数据分析。In an exemplary embodiment of the present invention, the digital PCR employs droplet digital PCR (droplet digital, PCR). The main feature is that the digital PCR mixture is added to the droplet generator to generate 10,000 to 20,000 micro-reaction droplets for PCR amplification. In each PCR layout, the positive quality control substance (M), negative quality control substance (U), RNase-free water and blank control NTC should be included at the same time. The so-called positive quality control substance can be treated with bisulfite. fully methylated human genomic DNA, while the negative control can be sulfurized unmethylated human genomic DNA. According to the quality control material and blank control, the threshold is set to determine whether the sample to be tested contains the DNA target molecules of the methylated human ZNF304 and LIFR genes, and at the same time, the quantity of the methylated human ZNF304 and LIFR target DNA molecules is determined. Data analysis is performed by corresponding software already commercially available (eg QuantaSoft, Bio-Rad).
2、DNA样本的提取和制备2. DNA sample extraction and preparation
本发明检测的DNA模板可以来源于提取待测者外周血血浆/血清中的循环游离DNA(cfDNA)、循环肿瘤DNA(ctDNA)或通过提取循环肿瘤细胞的DNA;具体操作详述如下。The DNA template detected by the present invention can be derived from the extraction of circulating cell-free DNA (cfDNA), circulating tumor DNA (ctDNA) in the peripheral blood plasma/serum of the subject or by extracting the DNA of circulating tumor cells; the specific operation is detailed as follows.
1)待测样本cfDNA/ctDNA的提取1) Extraction of cfDNA/ctDNA from the sample to be tested
取40例经病理组织学诊断的结直肠癌患者或13例健康对照的3-5mL血浆或血清,使用商品化的游离核酸提取试剂盒如QIAamp Circulating Nucleic Acid Kit制备DNA样本,具体操作按照试剂盒操作手册进行。用50uL洗脱液将待测DNA从离心柱上洗脱下来,用Qubit3.0荧光定量仪对样本浓度进行测定,此系统采用专门研制的荧光染料,比传统的紫外吸光法更加灵敏,尤其适用于本发明中珍贵的微量样品。Take 3-5 mL of plasma or serum from 40 patients with colorectal cancer diagnosed by histopathology or 13 healthy controls, and prepare DNA samples using a commercial free nucleic acid extraction kit such as QIAamp Circulating Nucleic Acid Kit. The specific operation is in accordance with the kit operation manual. Use 50uL of eluent to elute the DNA to be tested from the spin column, and use the Qubit3.0 fluorescence quantitative analyzer to measure the sample concentration. This system uses a specially developed fluorescent dye, which is more sensitive than the traditional UV absorption method, especially suitable for precious trace samples in the present invention.
2)待测样本DNA的硫化2) Sulfation of the DNA of the sample to be tested
使用市面上常用的商业化试剂盒例如Fast DNA Bisulfite Kit对提取的待测DNA进行亚硫酸氢盐处理,具体操作按照试剂盒操作手册进行。用20ul洗脱液洗脱待测DNA,用于数字PCR检测。Use commercially available kits such as The Fast DNA Bisulfite Kit performs bisulfite treatment on the extracted DNA to be tested. The specific operation is carried out according to the kit operation manual. The DNA to be tested was eluted with 20ul of eluent for digital PCR detection.
3、人ZNF304、LIFR基因甲基化数字PCR检测体系3. Human ZNF304, LIFR gene methylation digital PCR detection system
(1)数字PCR反应体系:将硫化后样本分为两份,分别配置两基因的反应体系,在20uL PCR反应体系中,2×ddPCR Supermix for Probes 10uL,加入引物及探针使其终浓度分别达到引物终浓度为0.9uM以及探针终浓度为0.25uM,5-6uL浓度为0.002-70ng/uL的模板DNA,补足ddH2O至21uL。(1) Digital PCR reaction system: Divide the sulfurized sample into two parts, and configure the reaction systems of the two genes respectively. In a 20uL PCR reaction system, 2×ddPCR Supermix for Probes 10uL, add primers and probes to make their final concentrations respectively To achieve a final primer concentration of 0.9uM and a final probe concentration of 0.25uM, 5-6uL of template DNA with a concentration of 0.002-70ng/uL, make up ddH2O to 21uL.
(2)微滴PCR反应体系的制备与扩增:在配置好的体系中抽取20ul(上步配多体系以免产生气泡)加入微滴发生卡(Bio-Rad)中间一排的8个孔内,在下方的空孔中加入70uL微滴发生油,垫上胶垫,缓慢转移到微滴生成仪(Bio-Rad)中生成微滴,约在2分钟内生成10000-20000个微滴。将制备好的微滴吸取40uL缓慢转移到96孔板中,封好膜之后应该在30分钟内进行PCR反应,或者放于4℃冰箱4小时之内进行PCR,可在任意一台96孔PCR仪中按照设定好的反应条件进行PCR反应。(2) Preparation and amplification of the droplet PCR reaction system: extract 20ul from the configured system (multiple systems were prepared in the previous step to avoid air bubbles) and add it to the 8 holes in the middle row of the droplet generating card (Bio-Rad). , add 70uL of droplet generating oil to the empty hole below, put a rubber pad on it, and slowly transfer it to a droplet generator (Bio-Rad) to generate droplets, and generate 10,000-20,000 droplets in about 2 minutes. Pipette 40uL of the prepared droplets and slowly transfer them to a 96-well plate. After sealing the membrane, the PCR reaction should be performed within 30 minutes, or placed in a refrigerator at 4°C for PCR within 4 hours. The PCR reaction was carried out in the instrument according to the set reaction conditions.
上述PCR反应扩增条件为95℃预变性10分钟,94℃变性30秒,60℃退火60秒,共进行40个循环,98℃灭活酶10分钟,之后4℃保持;升降温速度≤2.5℃/s。The above PCR amplification conditions were pre-denaturation at 95°C for 10 minutes, denaturation at 94°C for 30 seconds, annealing at 60°C for 60 seconds, a total of 40 cycles, inactivation of the enzyme at 98°C for 10 minutes, and then maintained at 4°C; °C/s.
(3)信号检测和结果判定:PCR扩增结束后,将96孔板放入微滴读取仪,按照设定好的顺序吸取每个样品的微滴并随微滴阅读油通过双色检测器(Bio-Rad),通过QuantaSoft(version 1.7)软件分析计算出待测样本中ZNF304、LIFR的液滴数。(3) Signal detection and result judgment: After PCR amplification, put the 96-well plate into the microdroplet reader, draw the microdroplets of each sample according to the set order, and pass the two-color detector with the microdroplet reading oil. (Bio-Rad), the number of droplets of ZNF304 and LIFR in the sample to be tested was calculated and analyzed by QuantaSoft (version 1.7) software.
4、检测体系有效性及结果的判断标准4. Judgment criteria for the effectiveness of the testing system and results
(1)ZNF304检测体系有效性及结果的判断:(1) Judgment of the effectiveness and results of the ZNF304 testing system:
a)阳性质控品(M):在FAM(Ch1)通道有信号,在VIC(Ch2)通道、去RNA酶水/NTC无信号,则判断为阳性质控品有效。否则重新配置。a) Positive quality control material (M): if there is a signal in the FAM (Ch1) channel, but no signal in the VIC (Ch2) channel and RNase-free water/NTC, the positive quality control material is judged to be valid. Otherwise reconfigure.
b)阴性质控品(U):在FAM(Ch1)通道、去RNA酶水/NTC无信号,在VIC(Ch2)通道有信号,则判断阴性质控品有效。否则重新配置。b) Negative quality control substance (U): There is no signal in the FAM (Ch1) channel, RNase-free water/NTC, and there is a signal in the VIC (Ch2) channel, and the negative quality control substance is judged to be effective. Otherwise reconfigure.
c)结果判定:根据阳性、阴性质控品、去RNA酶水/NTC划定阈值线,FAM(Ch1)通道阳性液滴数2个及以上判定为甲基化阳性;VIC(Ch2)通道阳性液滴数3个及以上判定为非甲基化阳性。在本检测体系中非甲基化阳性液滴要远远多于甲基化阳性液滴,质控品检测结果见图13。c) Determination of results: The threshold line is defined according to the positive and negative quality control substances, RNase-free water/NTC, and the number of positive droplets in the FAM (Ch1) channel is 2 or more positive for methylation; the VIC (Ch2) channel is positive Three or more droplets were judged as non-methylation positive. In this detection system, there are far more non-methylation positive droplets than methylation positive droplets. The test results of the quality control products are shown in Figure 13.
(2)LIFR检测体系有效性及结果的判断:(2) Judgment of the effectiveness and results of the LIFR testing system:
a)阳性质控品(M):在FAM(Ch1)通道有信号,在VIC(Ch2)通道、去RNA酶水/NTC无信号,则判断为阳性质控品有效。否则重新配置。a) Positive quality control material (M): if there is a signal in the FAM (Ch1) channel, but no signal in the VIC (Ch2) channel and RNase-free water/NTC, the positive quality control material is judged to be valid. Otherwise reconfigure.
b)阴性质控品(U):在FAM(Ch1)通道、去RNA酶水/NTC无信号,在VIC(Ch2)通道有信号,则判断阴性质控品有效。否则重新配置。b) Negative quality control substance (U): There is no signal in the FAM (Ch1) channel, RNase-free water/NTC, and there is a signal in the VIC (Ch2) channel, and the negative quality control substance is judged to be effective. Otherwise reconfigure.
c)结果判定:根据阳性、阴性质控品、去RNA酶水/NTC划定阈值线,FAM(Ch1)通道阳性液滴数2个及以上判定为甲基化阳性;VIC(Ch2)通道阳性液滴数2个及以上判定为非甲基化阳性,质控品检测结果见图14。c) Determination of results: The threshold line is defined according to the positive and negative quality control substances, RNase-free water/NTC, and the number of positive droplets in the FAM (Ch1) channel is 2 or more positive for methylation; the VIC (Ch2) channel is positive The number of droplets of 2 or more is judged as non-methylation positive, and the test results of the quality control products are shown in Figure 14.
5.检测结果及分析5. Test results and analysis
本发明共检测40例CRC患者和13例健康对照血浆游离DNA,根据阈值线计算每个样品的甲基化阳性液滴数和非甲基化阳性液滴数,甲基化阳性液滴数在总阳性液滴数中占的比例作为样品甲基化水平,结直肠癌患者血浆中ZNF304、LIFR均呈现高甲基化状态见图15、图16。使用ROC曲线计算两基因最佳cut-off值分别为0.003和0.002,分别在不同分期中计算两基因及其联合阳性率,结果见表16、表17、表18。The invention detects a total of 40 CRC patients and 13 healthy controls plasma free DNA, and calculates the number of methylation positive droplets and the number of non-methylation positive droplets for each sample according to the threshold line. The number of methylation positive droplets is in The proportion of the total number of positive droplets was taken as the methylation level of the sample. Both ZNF304 and LIFR in the plasma of colorectal cancer patients showed a hypermethylated state, as shown in Figure 15 and Figure 16 . The ROC curve was used to calculate the best cut-off values of the two genes, which were 0.003 and 0.002, respectively. The two genes and their combined positive rates were calculated in different stages. The results are shown in Table 16, Table 17, and Table 18.
表16Table 16
表17Table 17
表18Table 18
序列表sequence listing
<110> 哈尔滨医科大学<110> Harbin Medical University
<120> 结直肠癌相关的甲基化标志物以及用于检测结直肠癌的试剂盒<120> Colorectal cancer-related methylation markers and kits for detecting colorectal cancer
<141> 2022-06-13<141> 2022-06-13
<160> 33<160> 33
<170> SIPOSequenceListing 1.0<170> SIPOSequenceListing 1.0
<210> 1<210> 1
<211> 28<211> 28
<212> DNA<212> DNA
<213> artificial sequence<213> artificial sequence
<400> 1<400> 1
gattttaata gtaaaggtta ggggtttt 28gattttaata gtaaaggtta ggggtttt 28
<210> 2<210> 2
<211> 20<211> 20
<212> DNA<212> DNA
<213> artificial sequence<213> artificial sequence
<400> 2<400> 2
ccaacrcacc tccccaatac 20
<210> 3<210> 3
<211> 19<211> 19
<212> DNA<212> DNA
<213> artificial sequence<213> artificial sequence
<400> 3<400> 3
gayggttttg yggggagga 19
<210> 4<210> 4
<211> 18<211> 18
<212> DNA<212> DNA
<213> artificial sequence<213> artificial sequence
<400> 4<400> 4
ccccacaccc racaaaaa 18ccccacaccc racaaaaa 18
<210> 5<210> 5
<211> 27<211> 27
<212> DNA<212> DNA
<213> artificial sequence<213> artificial sequence
<400> 5<400> 5
aattagaatg tayggatygg gtatagg 27aattagaatg tayggatygg gtatagg 27
<210> 6<210> 6
<211> 26<211> 26
<212> DNA<212> DNA
<213> artificial sequence<213> artificial sequence
<400> 6<400> 6
aaatatcrtt ttcccaatct tttctc 26aaatatcrtt ttcccaatct tttctc 26
<210> 7<210> 7
<211> 31<211> 31
<212> DNA<212> DNA
<213> artificial sequence<213> artificial sequence
<400> 7<400> 7
aygttagaag tttygttttt gtagtttaga g 31aygttagaag tttygttttt gtagtttaga g 31
<210> 8<210> 8
<211> 31<211> 31
<212> DNA<212> DNA
<213> artificial sequence<213> artificial sequence
<400> 8<400> 8
aaatacatac cctttaaaaa caactataac c 31aaatacatac cctttaaaaa caactataac c 31
<210> 9<210> 9
<211> 20<211> 20
<212> DNA<212> DNA
<213> artificial sequence<213> artificial sequence
<400> 9<400> 9
ggatcgcgtt ttttcgaatg 20
<210> 10<210> 10
<211> 20<211> 20
<212> DNA<212> DNA
<213> artificial sequence<213> artificial sequence
<400> 10<400> 10
acaaaaacga cgaaaacacg 20
<210> 11<210> 11
<211> 26<211> 26
<212> DNA<212> DNA
<213> artificial sequence<213> artificial sequence
<400> 11<400> 11
tccccgcttt cctctcctca aaacac 26tccccgcttt cctctcctca aaacac 26
<210> 12<210> 12
<211> 21<211> 21
<212> DNA<212> DNA
<213> artificial sequence<213> artificial sequence
<400> 12<400> 12
cgttttttgt tgttttgggt t 21cgtttttttgt tgttttgggt t 21
<210> 13<210> 13
<211> 22<211> 22
<212> DNA<212> DNA
<213> artificial sequence<213> artificial sequence
<400> 13<400> 13
aaatacacga accgaacata aa 22aaatacacga accgaacata
<210> 14<210> 14
<211> 23<211> 23
<212> DNA<212> DNA
<213> artificial sequence<213> artificial sequence
<400> 14<400> 14
aatacgtcac gacgacgacg cca 23aatacgtcac gacgacgacg cca 23
<210> 15<210> 15
<211> 27<211> 27
<212> DNA<212> DNA
<213> artificial sequence<213> artificial sequence
<400> 15<400> 15
ggtttggttt ttttatgttt ttttagg 27ggtttggttt ttttatgttt ttttagg 27
<210> 16<210> 16
<211> 20<211> 20
<212> DNA<212> DNA
<213> artificial sequence<213> artificial sequence
<400> 16<400> 16
aacacctaaa ccaaacaccc 20
<210> 17<210> 17
<211> 27<211> 27
<212> DNA<212> DNA
<213> artificial sequence<213> artificial sequence
<400> 17<400> 17
tcccacctcc tcaaacatct acactct 27tcccacctcc tcaaacatct acactct 27
<210> 18<210> 18
<211> 18<211> 18
<212> DNA<212> DNA
<213> artificial sequence<213> artificial sequence
<400> 18<400> 18
gttgtagggg cgagattt 18gttgtaggggg cgagattt 18
<210> 19<210> 19
<211> 22<211> 22
<212> DNA<212> DNA
<213> artificial sequence<213> artificial sequence
<400> 19<400> 19
caaaatacac gaaccgaaca ta 22caaaatacac gaaccgaaca ta 22
<210> 20<210> 20
<211> 14<211> 14
<212> DNA<212> DNA
<213> artificial sequence<213> artificial sequence
<400> 20<400> 20
tcacgacgac gacg 14tcacgacgac gacg 14
<210> 21<210> 21
<211> 19<211> 19
<212> DNA<212> DNA
<213> artificial sequence<213> artificial sequence
<400> 21<400> 21
gttgttttgg gttgtaggg 19
<210> 22<210> 22
<211> 21<211> 21
<212> DNA<212> DNA
<213> artificial sequence<213> artificial sequence
<400> 22<400> 22
acaaccaaaa tacacaaacc a 21acaaccaaaa tacacaaacc a 21
<210> 23<210> 23
<211> 16<211> 16
<212> DNA<212> DNA
<213> artificial sequence<213> artificial sequence
<400> 23<400> 23
catcacaaca acaaca 16catcacaaca acaaca 16
<210> 24<210> 24
<211> 25<211> 25
<212> DNA<212> DNA
<213> artificial sequence<213> artificial sequence
<400> 24<400> 24
gggattttgc gaattattta aatag 25gggattttgc gaattattta aatag 25
<210> 25<210> 25
<211> 20<211> 20
<212> DNA<212> DNA
<213> artificial sequence<213> artificial sequence
<400> 25<400> 25
caaaaaaacc aaaacgcgaa 20
<210> 26<210> 26
<211> 13<211> 13
<212> DNA<212> DNA
<213> artificial sequence<213> artificial sequence
<400> 26<400> 26
acgacgcctc gac 13acgacgcctc gac 13
<210> 27<210> 27
<211> 20<211> 20
<212> DNA<212> DNA
<213> artificial sequence<213> artificial sequence
<400> 27<400> 27
tgattgttgg gattttgtga 20
<210> 28<210> 28
<211> 21<211> 21
<212> DNA<212> DNA
<213> artificial sequence<213> artificial sequence
<400> 28<400> 28
acccaacaaa aaaaccaaaa c 21acccaacaaa aaaaccaaaa c 21
<210> 29<210> 29
<211> 14<211> 14
<212> DNA<212> DNA
<213> artificial sequence<213> artificial sequence
<400> 29<400> 29
caacacctca accc 14caacacctca accc 14
<210> 30<210> 30
<211> 69<211> 69
<212> DNA<212> DNA
<213> artificial sequence<213> artificial sequence
<400> 30<400> 30
gttgtagggg cgagattttt ggcgtcgtcg tcgtgacgta ttttttttat gttcggttcg 60gttgtaggggg cgagatttttt ggcgtcgtcg tcgtgacgta ttttttttat gttcggttcg 60
tgtattttg 69tgtattttg 69
<210> 31<210> 31
<211> 84<211> 84
<212> DNA<212> DNA
<213> artificial sequence<213> artificial sequence
<400> 31<400> 31
gttgttttgg gttgtagggg tgagattttt ggtgttgttg ttgtgatgta ttttttttat 60gttgttttgg gttgtaggggg tgagattttt ggtgttgttg ttgtgatgta ttttttttat 60
gtttggtttg tgtattttgg ttgt 84gtttggtttg tgtattttgg ttgt 84
<210> 32<210> 32
<211> 66<211> 66
<212> DNA<212> DNA
<213> artificial sequence<213> artificial sequence
<400> 32<400> 32
gggattttgc gaattattta aatagtttag ggtcgaggcg tcgtcgttcg cgttttggtt 60gggattttgc gaattattta aatagtttag ggtcgaggcg tcgtcgttcg cgttttggtt 60
tttttg 66tttttg 66
<210> 33<210> 33
<211> 80<211> 80
<212> DNA<212> DNA
<213> artificial sequence<213> artificial sequence
<400> 33<400> 33
tgattgttgg gattttgtga attatttaaa tagtttaggg ttgaggtgtt gttgtttgtg 60tgattgttgg gattttgtga attatttaaa tagtttaggg ttgaggtgtt gttgtttgtg 60
ttttggtttt tttgttgggt 80ttttggtttt ttttgttgggt 80
Claims (5)
Priority Applications (1)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| CN202011165628.9A CN112430657B (en) | 2020-10-27 | 2020-10-27 | Methylation marker related to colorectal cancer and kit for detecting colorectal cancer |
Applications Claiming Priority (1)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| CN202011165628.9A CN112430657B (en) | 2020-10-27 | 2020-10-27 | Methylation marker related to colorectal cancer and kit for detecting colorectal cancer |
Publications (2)
| Publication Number | Publication Date |
|---|---|
| CN112430657A CN112430657A (en) | 2021-03-02 |
| CN112430657B true CN112430657B (en) | 2022-09-09 |
Family
ID=74696199
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| CN202011165628.9A Active CN112430657B (en) | 2020-10-27 | 2020-10-27 | Methylation marker related to colorectal cancer and kit for detecting colorectal cancer |
Country Status (1)
| Country | Link |
|---|---|
| CN (1) | CN112430657B (en) |
Families Citing this family (3)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN113981046A (en) * | 2021-11-05 | 2022-01-28 | 朱运峰 | DNA methylation detection method based on quantitative PCR technology and kit thereof |
| CN114908166B (en) * | 2022-04-27 | 2025-04-22 | 哈尔滨医科大学 | A set of specific methylation markers related to colorectal cancer and detection kits thereof |
| CN116144775A (en) * | 2022-12-13 | 2023-05-23 | 重庆伯豪诊断技术有限公司 | Methylation marker for colorectal cancer detection, primer probe combination, kit and application of methylation marker |
Family Cites Families (7)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| JP6269491B2 (en) * | 2012-09-19 | 2018-01-31 | シスメックス株式会社 | Method for obtaining information on colorectal cancer, and marker and kit for obtaining information on colorectal cancer |
| WO2016060278A1 (en) * | 2014-10-17 | 2016-04-21 | 国立大学法人東北大学 | Method for estimating sensitivity to drug therapy for colorectal cancer |
| CN109072300B (en) * | 2015-12-17 | 2023-01-31 | 伊路敏纳公司 | Differentiating methylation levels in complex biological samples |
| CN105543379B (en) * | 2016-01-27 | 2019-02-19 | 哈尔滨医科大学 | A primer, kit and application for early screening of colorectal cancer-related IGF2 gene methylation phenotype |
| WO2018140781A1 (en) * | 2017-01-27 | 2018-08-02 | Exact Sciences Development Company, Llc | Detection of colon neoplasia by analysis of methylated dna |
| EP3692164B1 (en) * | 2017-10-06 | 2025-10-15 | Helio Health Inc. | Methylation markers for diagnosing cancer |
| JP2021532735A (en) * | 2018-07-09 | 2021-12-02 | エイチケージー エピセラピューティクス リミテッドHkg Epitherapeutics Limited | DNA Methylation Markers and Their Use for Non-Invasive Detection of Cancer |
-
2020
- 2020-10-27 CN CN202011165628.9A patent/CN112430657B/en active Active
Also Published As
| Publication number | Publication date |
|---|---|
| CN112430657A (en) | 2021-03-02 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| EP2891720B1 (en) | Method for screening cancer | |
| US20100068720A1 (en) | Method and kit for detection of early cancer or pre-cancer using blood and body fluids | |
| JP2022530920A (en) | Markers for identifying and quantifying mutations, expression, splice variants, translocations, copy counts, or methylation changes in nucleic acid sequences | |
| CN112430657B (en) | Methylation marker related to colorectal cancer and kit for detecting colorectal cancer | |
| US9328379B2 (en) | Hypermethylation biomarkers for detection of head and neck squamous cell cancer | |
| CN104513851B (en) | Novel epigenetic biomarkers and methods for detecting bladder cancer | |
| US10017824B2 (en) | Method for determining nucleic acid composition of nucleic acid mixture | |
| CN116219020B (en) | Methylation reference gene and application thereof | |
| EP4592392A1 (en) | Novel tumor detection marker tagme and use thereof | |
| CN113215264A (en) | Detection kit for early screening of TMEM101 gene methylation in human peripheral blood circulating tumor DNA (deoxyribonucleic acid) of endometrial cancer | |
| WO2021169875A1 (en) | Cancer gene methylation measuring system and cancer in vitro detection method executed in same | |
| WO2021180106A1 (en) | Probe composition for detecting five tumors of digestive tract | |
| CN116254344A (en) | Composition for detecting bladder cancer, kit and application thereof | |
| WO2021180105A1 (en) | Probe composition for detecting common cancers of both sexes | |
| CN111440863B (en) | Application of KAZN gene methylation detection reagent in preparation of colorectal cancer prognosis diagnosis reagent | |
| WO2021185274A1 (en) | Probe composition for detecting 6 cancers with high incidence in china | |
| US20220033911A1 (en) | Method for determining prognosis of endometrial cancer | |
| CN110117660A (en) | It can be used for the marker STAMP-EP11 and its detection reagent of tumour identification | |
| WO2021169874A1 (en) | Probe composition for detecting three lumen organ tumors | |
| WO2021175284A1 (en) | Probe composition for detecting three types of solid organ tumors | |
| CN116064788B (en) | Multiplex gene methylation detection fluorescence quantitative PCR kit for early screening of breast cancer | |
| CN118421801A (en) | Colorectal cancer hypermethylation target and application thereof | |
| CN114908166A (en) | A group of specific methylation markers associated with colorectal cancer and their detection kits | |
| CN111440866B (en) | Application of DUSP3 gene methylation detection reagent in preparation of colorectal cancer prognostic diagnostic reagent | |
| WO2021185275A1 (en) | Probe composition for detecting 11 cancers |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| PB01 | Publication | ||
| PB01 | Publication | ||
| SE01 | Entry into force of request for substantive examination | ||
| SE01 | Entry into force of request for substantive examination | ||
| GR01 | Patent grant | ||
| GR01 | Patent grant |