CN112410331A - Linker with molecular label and sample label and single-chain library building method thereof - Google Patents
Linker with molecular label and sample label and single-chain library building method thereof Download PDFInfo
- Publication number
- CN112410331A CN112410331A CN202011174182.6A CN202011174182A CN112410331A CN 112410331 A CN112410331 A CN 112410331A CN 202011174182 A CN202011174182 A CN 202011174182A CN 112410331 A CN112410331 A CN 112410331A
- Authority
- CN
- China
- Prior art keywords
- sequence
- seq
- sequencing
- optionally
- dna
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Pending
Links
- 238000000034 method Methods 0.000 title claims abstract description 30
- 238000012163 sequencing technique Methods 0.000 claims abstract description 146
- 238000010276 construction Methods 0.000 claims abstract description 49
- 230000008569 process Effects 0.000 claims abstract description 5
- 108020004414 DNA Proteins 0.000 claims description 275
- 102000053602 DNA Human genes 0.000 claims description 51
- 108020004682 Single-Stranded DNA Proteins 0.000 claims description 31
- 230000000295 complement effect Effects 0.000 claims description 31
- 238000006243 chemical reaction Methods 0.000 claims description 29
- 238000002156 mixing Methods 0.000 claims description 28
- 239000011324 bead Substances 0.000 claims description 20
- 102000012410 DNA Ligases Human genes 0.000 claims description 17
- 108010061982 DNA Ligases Proteins 0.000 claims description 17
- 238000006209 dephosphorylation reaction Methods 0.000 claims description 12
- 238000004925 denaturation Methods 0.000 claims description 11
- 230000036425 denaturation Effects 0.000 claims description 11
- YBJHBAHKTGYVGT-ZKWXMUAHSA-N (+)-Biotin Chemical compound N1C(=O)N[C@@H]2[C@H](CCCCC(=O)O)SC[C@@H]21 YBJHBAHKTGYVGT-ZKWXMUAHSA-N 0.000 claims description 10
- 230000030609 dephosphorylation Effects 0.000 claims description 10
- 239000003550 marker Substances 0.000 claims description 10
- 238000002844 melting Methods 0.000 claims description 10
- 230000008018 melting Effects 0.000 claims description 10
- 230000015572 biosynthetic process Effects 0.000 claims description 9
- 238000003786 synthesis reaction Methods 0.000 claims description 9
- 102000004594 DNA Polymerase I Human genes 0.000 claims description 7
- 108010017826 DNA Polymerase I Proteins 0.000 claims description 7
- 229960002685 biotin Drugs 0.000 claims description 6
- 239000011616 biotin Substances 0.000 claims description 6
- 238000010438 heat treatment Methods 0.000 claims description 6
- 102000016928 DNA-directed DNA polymerase Human genes 0.000 claims description 5
- 108010014303 DNA-directed DNA polymerase Proteins 0.000 claims description 5
- 108010090804 Streptavidin Proteins 0.000 claims description 5
- 235000020958 biotin Nutrition 0.000 claims description 5
- 102000004190 Enzymes Human genes 0.000 claims description 4
- 108090000790 Enzymes Proteins 0.000 claims description 4
- LSNNMFCWUKXFEE-UHFFFAOYSA-N Sulfurous acid Chemical compound OS(O)=O LSNNMFCWUKXFEE-UHFFFAOYSA-N 0.000 claims description 4
- 238000002372 labelling Methods 0.000 claims description 4
- 238000010309 melting process Methods 0.000 claims description 4
- 238000003505 heat denaturation Methods 0.000 claims description 3
- 125000002467 phosphate group Chemical group [H]OP(=O)(O[H])O[*] 0.000 claims description 3
- 239000002773 nucleotide Substances 0.000 abstract description 6
- 125000003729 nucleotide group Chemical group 0.000 abstract description 6
- 206010034944 Photokeratitis Diseases 0.000 abstract description 4
- 239000000523 sample Substances 0.000 description 88
- 239000000047 product Substances 0.000 description 21
- 238000005516 engineering process Methods 0.000 description 17
- 239000006228 supernatant Substances 0.000 description 17
- 238000002474 experimental method Methods 0.000 description 16
- 206010028980 Neoplasm Diseases 0.000 description 10
- 238000004458 analytical method Methods 0.000 description 9
- XLYOFNOQVPJJNP-UHFFFAOYSA-N water Chemical compound O XLYOFNOQVPJJNP-UHFFFAOYSA-N 0.000 description 9
- 108091092584 GDNA Proteins 0.000 description 8
- 206010058467 Lung neoplasm malignant Diseases 0.000 description 8
- 201000005202 lung cancer Diseases 0.000 description 8
- 208000020816 lung neoplasm Diseases 0.000 description 8
- 238000003908 quality control method Methods 0.000 description 8
- 238000010828 elution Methods 0.000 description 7
- 239000000203 mixture Substances 0.000 description 7
- 238000002360 preparation method Methods 0.000 description 7
- 238000000746 purification Methods 0.000 description 7
- LFQSCWFLJHTTHZ-UHFFFAOYSA-N Ethanol Chemical compound CCO LFQSCWFLJHTTHZ-UHFFFAOYSA-N 0.000 description 5
- 238000000137 annealing Methods 0.000 description 5
- 101710086015 RNA ligase Proteins 0.000 description 4
- 230000003321 amplification Effects 0.000 description 4
- 238000003199 nucleic acid amplification method Methods 0.000 description 4
- 102000039446 nucleic acids Human genes 0.000 description 4
- 108020004707 nucleic acids Proteins 0.000 description 4
- 150000007523 nucleic acids Chemical class 0.000 description 4
- 102000040430 polynucleotide Human genes 0.000 description 4
- 108091033319 polynucleotide Proteins 0.000 description 4
- 239000002157 polynucleotide Substances 0.000 description 4
- 229910001868 water Inorganic materials 0.000 description 4
- 239000007795 chemical reaction product Substances 0.000 description 3
- 238000004128 high performance liquid chromatography Methods 0.000 description 3
- 238000009396 hybridization Methods 0.000 description 3
- 108090000623 proteins and genes Proteins 0.000 description 3
- OKTJSMMVPCPJKN-UHFFFAOYSA-N Carbon Chemical group [C] OKTJSMMVPCPJKN-UHFFFAOYSA-N 0.000 description 2
- 238000001712 DNA sequencing Methods 0.000 description 2
- 108091034117 Oligonucleotide Proteins 0.000 description 2
- 239000007984 Tris EDTA buffer Substances 0.000 description 2
- 230000027455 binding Effects 0.000 description 2
- 239000000090 biomarker Substances 0.000 description 2
- 238000007664 blowing Methods 0.000 description 2
- 238000001514 detection method Methods 0.000 description 2
- 238000003745 diagnosis Methods 0.000 description 2
- 238000010586 diagram Methods 0.000 description 2
- 239000012634 fragment Substances 0.000 description 2
- 230000008676 import Effects 0.000 description 2
- 238000011534 incubation Methods 0.000 description 2
- 230000004048 modification Effects 0.000 description 2
- 238000012986 modification Methods 0.000 description 2
- 230000035772 mutation Effects 0.000 description 2
- 230000008520 organization Effects 0.000 description 2
- 238000009609 prenatal screening Methods 0.000 description 2
- 230000008439 repair process Effects 0.000 description 2
- 230000000717 retained effect Effects 0.000 description 2
- 239000000243 solution Substances 0.000 description 2
- 125000006850 spacer group Chemical group 0.000 description 2
- 239000000126 substance Substances 0.000 description 2
- 239000000725 suspension Substances 0.000 description 2
- LSNNMFCWUKXFEE-UHFFFAOYSA-M Bisulfite Chemical compound OS([O-])=O LSNNMFCWUKXFEE-UHFFFAOYSA-M 0.000 description 1
- MVXXEGDUIIWXFP-UFLZEWODSA-N OCC(O)CO.OC(=O)CCCC[C@@H]1SC[C@@H]2NC(=O)N[C@H]12 Chemical compound OCC(O)CO.OC(=O)CCCC[C@@H]1SC[C@@H]2NC(=O)N[C@H]12 MVXXEGDUIIWXFP-UFLZEWODSA-N 0.000 description 1
- 229920002594 Polyethylene Glycol 8000 Polymers 0.000 description 1
- 229920001213 Polysorbate 20 Polymers 0.000 description 1
- 108020004518 RNA Probes Proteins 0.000 description 1
- 239000003391 RNA probe Substances 0.000 description 1
- 206010041662 Splinter Diseases 0.000 description 1
- JLCPHMBAVCMARE-UHFFFAOYSA-N [3-[[3-[[3-[[3-[[3-[[3-[[3-[[3-[[3-[[3-[[3-[[5-(2-amino-6-oxo-1H-purin-9-yl)-3-[[3-[[3-[[3-[[3-[[3-[[5-(2-amino-6-oxo-1H-purin-9-yl)-3-[[5-(2-amino-6-oxo-1H-purin-9-yl)-3-hydroxyoxolan-2-yl]methoxy-hydroxyphosphoryl]oxyoxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(5-methyl-2,4-dioxopyrimidin-1-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(6-aminopurin-9-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(6-aminopurin-9-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(6-aminopurin-9-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(6-aminopurin-9-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxyoxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(5-methyl-2,4-dioxopyrimidin-1-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(4-amino-2-oxopyrimidin-1-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(5-methyl-2,4-dioxopyrimidin-1-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(5-methyl-2,4-dioxopyrimidin-1-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(6-aminopurin-9-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(6-aminopurin-9-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(4-amino-2-oxopyrimidin-1-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(4-amino-2-oxopyrimidin-1-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(4-amino-2-oxopyrimidin-1-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(6-aminopurin-9-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(4-amino-2-oxopyrimidin-1-yl)oxolan-2-yl]methyl [5-(6-aminopurin-9-yl)-2-(hydroxymethyl)oxolan-3-yl] hydrogen phosphate Polymers Cc1cn(C2CC(OP(O)(=O)OCC3OC(CC3OP(O)(=O)OCC3OC(CC3O)n3cnc4c3nc(N)[nH]c4=O)n3cnc4c3nc(N)[nH]c4=O)C(COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3CO)n3cnc4c(N)ncnc34)n3ccc(N)nc3=O)n3cnc4c(N)ncnc34)n3ccc(N)nc3=O)n3ccc(N)nc3=O)n3ccc(N)nc3=O)n3cnc4c(N)ncnc34)n3cnc4c(N)ncnc34)n3cc(C)c(=O)[nH]c3=O)n3cc(C)c(=O)[nH]c3=O)n3ccc(N)nc3=O)n3cc(C)c(=O)[nH]c3=O)n3cnc4c3nc(N)[nH]c4=O)n3cnc4c(N)ncnc34)n3cnc4c(N)ncnc34)n3cnc4c(N)ncnc34)n3cnc4c(N)ncnc34)O2)c(=O)[nH]c1=O JLCPHMBAVCMARE-UHFFFAOYSA-N 0.000 description 1
- 238000007259 addition reaction Methods 0.000 description 1
- 239000007864 aqueous solution Substances 0.000 description 1
- 230000009286 beneficial effect Effects 0.000 description 1
- 230000000035 biogenic effect Effects 0.000 description 1
- 239000000872 buffer Substances 0.000 description 1
- 229910052799 carbon Inorganic materials 0.000 description 1
- 230000015556 catabolic process Effects 0.000 description 1
- 238000005119 centrifugation Methods 0.000 description 1
- 239000003153 chemical reaction reagent Substances 0.000 description 1
- 238000001816 cooling Methods 0.000 description 1
- 238000012937 correction Methods 0.000 description 1
- 230000008878 coupling Effects 0.000 description 1
- 238000010168 coupling process Methods 0.000 description 1
- 238000005859 coupling reaction Methods 0.000 description 1
- 238000006731 degradation reaction Methods 0.000 description 1
- 238000012217 deletion Methods 0.000 description 1
- 230000037430 deletion Effects 0.000 description 1
- 239000012153 distilled water Substances 0.000 description 1
- 230000009977 dual effect Effects 0.000 description 1
- 230000000694 effects Effects 0.000 description 1
- 230000002255 enzymatic effect Effects 0.000 description 1
- 230000004907 flux Effects 0.000 description 1
- 230000001788 irregular Effects 0.000 description 1
- 239000007788 liquid Substances 0.000 description 1
- 238000004519 manufacturing process Methods 0.000 description 1
- 238000013507 mapping Methods 0.000 description 1
- 239000000463 material Substances 0.000 description 1
- 238000012164 methylation sequencing Methods 0.000 description 1
- 230000036438 mutation frequency Effects 0.000 description 1
- 230000009871 nonspecific binding Effects 0.000 description 1
- 238000006366 phosphorylation reaction Methods 0.000 description 1
- 239000000256 polyoxyethylene sorbitan monolaurate Substances 0.000 description 1
- 235000010486 polyoxyethylene sorbitan monolaurate Nutrition 0.000 description 1
- 238000011176 pooling Methods 0.000 description 1
- 125000002924 primary amino group Chemical group [H]N([H])* 0.000 description 1
- 238000012545 processing Methods 0.000 description 1
- 238000012797 qualification Methods 0.000 description 1
- 239000002096 quantum dot Substances 0.000 description 1
- 239000002994 raw material Substances 0.000 description 1
- 239000011535 reaction buffer Substances 0.000 description 1
- 239000011435 rock Substances 0.000 description 1
- 230000005783 single-strand break Effects 0.000 description 1
- 238000006467 substitution reaction Methods 0.000 description 1
- 229910021642 ultra pure water Inorganic materials 0.000 description 1
- 239000012498 ultrapure water Substances 0.000 description 1
- 238000005406 washing Methods 0.000 description 1
Images
Classifications
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/11—DNA or RNA fragments; Modified forms thereof; Non-coding nucleic acids having a biological activity
-
- C—CHEMISTRY; METALLURGY
- C40—COMBINATORIAL TECHNOLOGY
- C40B—COMBINATORIAL CHEMISTRY; LIBRARIES, e.g. CHEMICAL LIBRARIES
- C40B50/00—Methods of creating libraries, e.g. combinatorial synthesis
- C40B50/06—Biochemical methods, e.g. using enzymes or whole viable microorganisms
Landscapes
- Life Sciences & Earth Sciences (AREA)
- Chemical & Material Sciences (AREA)
- Health & Medical Sciences (AREA)
- Engineering & Computer Science (AREA)
- Organic Chemistry (AREA)
- Genetics & Genomics (AREA)
- Molecular Biology (AREA)
- Biochemistry (AREA)
- Biomedical Technology (AREA)
- Biotechnology (AREA)
- General Engineering & Computer Science (AREA)
- Bioinformatics & Cheminformatics (AREA)
- Microbiology (AREA)
- Wood Science & Technology (AREA)
- Zoology (AREA)
- Physics & Mathematics (AREA)
- Biophysics (AREA)
- General Health & Medical Sciences (AREA)
- Plant Pathology (AREA)
- Chemical Kinetics & Catalysis (AREA)
- General Chemical & Material Sciences (AREA)
- Medicinal Chemistry (AREA)
- Measuring Or Testing Involving Enzymes Or Micro-Organisms (AREA)
Abstract
In the adaptor, the 3 'end of a first molecular label is connected to the 5' end of a first sequencing adaptor, the 5 'end of the first molecular label is connected to the 3' end of a forward chain of the first sample label, the 5 'end of the forward chain of the first sample label is used for being connected to a single-chain template molecule to be subjected to library construction, and the 3' end of a reverse chain of the first sample label is connected with a random sequence. The molecular label is added on the hairpin type single-chain library building joint to correct random errors in the PCR and sequencing processes; the sample tag is added beside the molecular tag, so that the problem of sequencing error caused by signal snow blindness caused by the same nucleotide at the specific base position of a plurality of molecules in a library to be tested by a camera system of a sequencer is effectively avoided, and the inter-sample tag crosstalk in multi-sample mixed sequencing is greatly reduced through the direct connection of the sample tag and the warehousing template molecule.
Description
Technical Field
The invention relates to the technical field of high-throughput DNA sequencing library construction, in particular to a joint with a molecular label and a sample label and a single-strand library construction method thereof.
Background
The existing high-throughput DNA sequencing library is generally characterized in that a double-stranded sequencing joint is connected to a double-stranded DNA template, in many severely degraded DNA samples, DNA molecules exist in a form of single strand and double strand mixture, or one strand of the double-stranded DNA molecules has discontinuous deletion (single strand break), such as extracellular circulating DNA samples (also called cfDNA samples, which are common biomarkers in the fields of prenatal screening, tumor diagnosis, allograft and the like), or such as formalin-fixed and paraffin-embedded biological tissue samples, forensic samples and DNA samples extracted from ancient biogenic rocks, and in addition, in methylation sequencing, DNA after bisulfite conversion is broken into short pieces and single-stranded. If the conventional library construction method of double-link joint is still adopted for constructing the library of the sample, all template molecules except the regular double-stranded DNA molecules can not be or rarely be built into the library, so that a great amount of original template molecules are lost; if the single-chain library construction technology is adopted, various template molecules can be constructed into a library, the loss of the irregular double-chain template molecules is avoided to the maximum extent, and the library construction efficiency and the quality of downstream sequencing data are greatly improved.
The existing single-chain database building technology mainly comprises the following two types:
1. t4 RNA ligase-based single-strand library construction method: according to the method, a single-stranded sequencing joint is directly connected to a single-stranded DNA template molecule, so that a molecular label can be easily added on the single-stranded joint, but the method depends on T4 RNA ligase, is high in price, and does not realize stable quality-guaranteeing production at home, so that the method completely depends on import.
2. T4DNA ligase-based single-strand library construction method: the joint of the hairpin structure designed by the method realizes the connection of the single-strand sequencing joint and the single-strand template molecule by using T4DNA ligase, the adopted T4DNA ligase is cheap and easy to obtain, and has domestic raw material suppliers with reliable quality, thereby being beneficial to large-scale popularization and industrialization. However, the unique structure of the hairpin sequencing linker results in its inability to directly attach molecular tags to the 5' end of the single-stranded linker (i.e., directly contact the DNA template) as in the direct single-stranded ligation method (i.e., the first method described above). There is no effective method in the prior art for attaching molecular tags to hairpin linkers.
Disclosure of Invention
According to a first aspect, an embodiment provides a linker with a molecular tag and a sample tag, comprising a first molecular tag, a first sample tag, and a first sequencing linker, wherein the first sample tag has a reverse complementary forward strand and a reverse strand, the first molecular tag is a single-stranded DNA molecule, the 3 'end of the first molecular tag is connected to the 5' end of the first sequencing linker, the 5 'end of the first molecular tag is connected to the 3' end of the forward strand of the first sample tag, the 5 'end of the forward strand of the first sample tag is used for connecting to a single-stranded template molecule to be pooled, and the 3' end of the reverse strand of the first sample tag is connected with a random sequence.
According to a second aspect, there is provided in one embodiment a kit for library construction comprising a linker as described in the first aspect.
According to a third aspect, one embodiment provides a method of constructing a single-stranded library with a molecular tag, comprising:
dephosphorizing, namely dephosphorizing the template molecules of the library to be built;
a first sequencing joint connection step, which comprises connecting the joint of the first aspect to a single-stranded template molecule to be built by DNA ligase, wherein the 3' end of the first sequencing joint is modified with a marker molecule;
a first melting step comprising denaturing the product of the first sequencing linker ligation step, allowing the reverse strand of the first sample tag, to which the random sequence is ligated, to dissociate from the linker sequence, and collecting the intermediate library molecules modified with the tag molecule;
a primary double-stranded synthesis step, comprising mixing a first primer, a DNA polymerase and the intermediate library molecule modified with a marker molecule, allowing the first primer to anneal and extend a first sequencing linker part on the intermediate library molecule, and reacting to obtain a double-stranded blunt-ended DNA molecule, wherein the first primer contains a sequence complementary to the first sequencing linker;
a second sequencing adaptor connection step, which comprises mixing the double-stranded blunt-end DNA molecules with a second sequencing adaptor and DNA ligase, and reacting to obtain double-stranded DNA library molecules connected with the second sequencing adaptor, wherein the second sequencing adaptor is a double-stranded sequencing adaptor;
a secondary unzipping step comprising denaturing the double-stranded DNA library molecules such that the double-stranded DNA library molecules dissociate into single-stranded DNA molecules, collecting single-stranded DNA library molecules that are not modified with a marker molecule;
and a second double-stranded synthesis step, which comprises adding a second primer and a third primer, and amplifying to obtain a double-stranded DNA library for sequencing, wherein the second primer contains a sequence of a first sequencing joint complementary to the single-stranded DNA library molecule without the modified labeled molecule, and the third primer contains a sequence of a second sequencing joint complementary to the single-stranded DNA library molecule without the modified labeled molecule.
According to the adaptor with the molecular tag and the sample tag and the single-chain library building method thereof, the molecular tag is added on the hairpin type single-chain library building adaptor so as to correct random errors in the PCR and sequencing processes; meanwhile, by adding the combined sample label beside the molecular label, the problem of sequencing error caused by signal snow blindness caused by the same nucleotide at the specific base position of a plurality of molecules in a library to be tested in a camera system of a sequencer is effectively avoided, and by directly connecting the sample label with the warehousing template molecule, the label crosstalk among samples in multi-sample mixed sequencing is greatly reduced.
Drawings
FIG. 1 is a flow chart illustrating single-stranded library construction based on single-ended molecular tag adapters in an embodiment.
FIG. 2 is a flow chart of single-stranded library construction based on a double-ended molecular tag linker in an embodiment.
Fig. 3 and 4 are diagrams illustrating an exemplary label assembly in an embodiment.
FIG. 5 is a schematic diagram of the structure of the molecular tag and the sample tag in one embodiment.
Detailed Description
The present invention will be described in further detail with reference to the following detailed description and accompanying drawings. Wherein like elements in different embodiments are numbered with like associated elements. In the following description, numerous details are set forth in order to provide a better understanding of the present application. However, those skilled in the art will readily recognize that some of the features may be omitted or replaced with other elements, materials, methods in different instances. In some instances, certain operations related to the present application have not been shown or described in detail in order to avoid obscuring the core of the present application from excessive description, and it is not necessary for those skilled in the art to describe these operations in detail, so that they may be fully understood from the description in the specification and the general knowledge in the art.
Furthermore, the features, operations, or characteristics described in the specification may be combined in any suitable manner to form various embodiments. Also, the various steps or actions in the method descriptions may be transposed or transposed in order, as will be apparent to one of ordinary skill in the art. Thus, the various sequences in the specification and drawings are for the purpose of describing certain embodiments only and are not intended to imply a required sequence unless otherwise indicated where such sequence must be followed.
The numbering of the components as such, e.g., "first", "second", etc., is used herein only to distinguish the objects as described, and does not have any sequential or technical meaning. The term "connected" and "coupled" when used in this application, unless otherwise indicated, includes both direct and indirect connections (couplings).
Herein, the hairpin structure (which may also be expressed as hairpin structure) means: a specific spatial structure formed by a pair of inverted repeat folding pairs.
Herein, the forward strand of the first sample tag is the strand that is co-stranded with and 5' to the first molecular tag, as shown in FIG. 5. The strand that is reverse complementary paired to the forward strand is the reverse strand of the first sample tag.
In a first aspect, in an embodiment, there is provided a linker, including a first molecular tag, a first sample tag, and a first sequencing linker, the first sample tag has a reverse complementary forward strand and a reverse strand, the first molecular tag is a single-stranded DNA molecule, the 3 'end of the first molecular tag is connected to the 5' end of the first sequencing linker, the 5 'end of the first molecular tag is connected to the 3' end of the forward strand of the first sample tag, the 5 'end of the forward strand of the first sample tag is used for connecting to a single-stranded template molecule to be pooled, and the 3' end of the reverse strand of the first sample tag is connected with a random sequence.
In one embodiment, the molecular tag is added on the hairpin type single-chain library building joint, so that random errors in the PCR and sequencing processes are avoided; meanwhile, by adding the combined sample label beside the molecular label, the problem of sequencing error caused by signal snow blindness caused by the same nucleotide when a camera system of a sequencer reads the specific base position of a plurality of molecules in a library to be tested is effectively avoided, and label crosstalk among samples subjected to hybrid sequencing is greatly reduced by the direct connection of the sample label and the template molecule.
In one embodiment, after the single-link tag is added with the molecular tag, error correction can be better performed, and ultra-low frequency mutation detection can be achieved, and generally can be achieved by 0.05%.
In one embodiment, the linker can effectively prevent the camera system for detecting the fluorescence signal from generating a snow-blind effect when all the DNA molecules to be detected are the same nucleotide (A, T, C or G) at a certain base site when a high-throughput sequencer reads the sequence of the detected DNA molecules, so that the sequencer cannot normally interpret the nucleotide type of the site.
In one embodiment, the adaptor can be used as a phase-variable sample tag, and a total of 192 sample tags (hairpin adaptors) are divided into 24 groups, 8 groups are used as one group, each group can be used for the library construction of one sample independently, and can be mixed with any group in the other 23 groups for use, namely, at most 24 libraries constructed by the method of the invention can be mixed and sequenced in the same lane (lane) during sample mixing sequencing, and at least one library (i.e. one group of sample tags is used) can be tested, and the flux is flexible.
In one embodiment, as shown in FIG. 5, the first sample tag is modified at the 5' end with a phosphate group.
In one embodiment, the first molecular tag is a random sequence.
In one embodiment, the first molecular tag is 4-19nt in length. The length of the first molecular tag may be 4nt, 5nt, 6nt, 7nt, 8nt, 9nt, 10nt, 11nt, 12nt, 13nt, 14nt, 15nt, 16nt, 17nt, 18nt, 19nt, and so on.
In one embodiment, the 3' end of the first sequencing linker is modified with a labeling molecule.
In one embodiment, the labeling molecule is selected from at least one of biotin, TEG biotin (or biotin-glycerol). The tag molecule can be captured by a streptavidin molecule.
In one embodiment, the first sequencing linker is a single stranded DNA.
In one embodiment, the library building template molecules to be built are single-stranded DNA molecules. If the original sample contains double-stranded DNA, the double-stranded DNA can be dissociated into single-stranded DNA by means of denaturation treatment and the like, and the single-stranded DNA is used as a library building template molecule to be built after dephosphorylation treatment.
In one embodiment, the forward strand of the first sample tag is selected from at least one of the sequences set forth in SEQ ID No.1-SEQ ID No. 192.
In one embodiment, the forward strand of the first sample tag is selected from any one of the sequences shown in SEQ ID No.1-SEQ ID No. 192.
In one embodiment, the random sequence attached to the 3' end of the reverse strand of the first sample tag is 5-20nt in length. Specifically, the number of the first and second groups may be 5nt, 6nt, 7nt, 8nt, 9nt, 10nt, 11nt, 12nt, 13nt, 14nt, 15nt, 16nt, 17nt, 18nt, 19nt, 20nt, and the like.
In one embodiment, the first sequencing linker is selected from any one of a P7-terminal sequencing linker of the Illumina sequencing platform, a P1-terminal sequencing linker of the MGI sequencing platform.
In one embodiment, the template molecules to be pooled are from severely degraded samples and/or sulfite treated samples. Specifically, the sample may include, but is not limited to, at least one of extracellular circulating DNA samples (also called cfDNA samples, common biomarkers in the fields of prenatal screening, tumor diagnosis, allograft, etc.), formalin-fixed and paraffin-embedded biological tissue samples, forensic samples, and DNA samples extracted from ancient fossils, and the like. The severely degraded sample and/or sulfite-treated sample may be subjected to a melting treatment (e.g., thermal denaturation treatment) and a dephosphorylation treatment in sequence to obtain a template molecule to be pooled.
In a second aspect, in one embodiment, there is provided a kit for library construction comprising the linker of the first aspect.
In one embodiment, a second sequencing linker for attaching to the intermediate library molecule is also included.
In one embodiment, the second sequencing adapter may or may not have a second molecular tag attached to its end for attachment to the intermediate library molecule, i.e., the second sequencing adapter may or may not have a second molecular tag attached to its end for attachment to the intermediate library molecule.
In one embodiment, the second molecular tag is a random sequence.
In one embodiment, the second molecular tag is 4-19nt in length. Specifically, the number of the first and second groups may be 4nt, 5nt, 6nt, 7nt, 8nt, 9nt, 10nt, 11nt, 12nt, 13nt, 14nt, 15nt, 16nt, 17nt, 18nt, 19nt, and the like. The length of the first molecular label and the length of the second molecular label can be the same or different.
In one embodiment, the second sequencing linker is selected from any one of a P5-terminal sequencing linker of the Illumina sequencing platform, a P2-terminal sequencing linker of the MGI sequencing platform.
In one embodiment, a second primer and/or a third primer is further included.
In one embodiment, the second primer contains or does not contain a second sample tag.
In one embodiment, when the second primer does not contain the second sample tag, the second primer comprises an inner adaptor sequence, an outer adaptor sequence, wherein the 5 'end of the inner adaptor sequence is linked to the 3' end of the outer adaptor sequence, and wherein the inner adaptor sequence is reverse complementary to the first sequencing adaptor.
In one embodiment, when the second primer comprises the second sample tag, the second primer further comprises an inner adaptor sequence and an outer adaptor sequence, wherein the 5 'end of the inner adaptor sequence is linked to the 3' end of the outer adaptor sequence, the inner adaptor sequence is reverse complementary to the first sequencing adaptor, and the second sample tag is located between the inner adaptor sequence and the outer adaptor sequence.
In one embodiment, the third primer contains or does not contain a third sample tag.
In one embodiment, when the third primer does not contain the second sample tag, the third primer comprises an inner adaptor sequence, an outer adaptor sequence, wherein the 5 'end of the inner adaptor sequence is linked to the 3' end of the outer adaptor sequence, and wherein the inner adaptor sequence is reverse complementary to the second sequencing adaptor.
In one embodiment, when the third primer comprises a third sample tag, the second primer comprises an inner adaptor sequence and an outer adaptor sequence, the 5 'end of the inner adaptor sequence is linked to the 3' end of the outer adaptor sequence, the inner adaptor sequence is reverse complementary mateable to the second sequencing adaptor, and the third sample tag is located between the inner adaptor sequence and the outer adaptor sequence.
In a third aspect, in one embodiment, a method for single-stranded library construction with a molecular tag and a sample tag is provided, comprising:
dephosphorizing, namely dephosphorizing the template molecules of the library to be built;
a first sequencing joint connection step, which comprises connecting the joint of the first aspect to a single-stranded template molecule to be built by DNA ligase, wherein the 3' end of the first sequencing joint is modified with a marker molecule;
a first melting step comprising denaturing the product of the first sequencing linker ligation step, allowing the reverse strand of the first sample tag to dissociate from the linker, and collecting the intermediate library molecules modified with the tag molecules;
a primary double-stranded synthesis step, comprising mixing a first primer, a DNA polymerase and the intermediate library molecule modified with a marker molecule, allowing the first primer to anneal and extend a first sequencing linker part on the intermediate library molecule, and reacting to obtain a double-stranded blunt-ended DNA molecule, wherein the first primer contains a sequence complementary to the first sequencing linker;
a second sequencing adaptor connection step, which comprises mixing the double-stranded blunt-end DNA molecules with a second sequencing adaptor and DNA ligase, and reacting to obtain double-stranded DNA library molecules connected with the second sequencing adaptor, wherein the second sequencing adaptor is a double-stranded sequencing adaptor;
a secondary unzipping step comprising denaturing the double-stranded DNA library molecules such that the double-stranded DNA library molecules dissociate into single-stranded DNA molecules, collecting single-stranded DNA library molecules that are not modified with a marker molecule;
and a second double-stranded synthesis step, which comprises adding a second primer and a third primer, and amplifying to obtain a double-stranded DNA library for sequencing, wherein the second primer contains a sequence of a first sequencing joint complementary to the single-stranded DNA library molecule without the modified labeled molecule, and the third primer contains a sequence of a second sequencing joint complementary to the single-stranded DNA library molecule without the modified labeled molecule.
In one embodiment, the forward strand of the first sample tag is selected from at least one of the sequences set forth in SEQ ID No.1-SEQ ID No. 192.
In one embodiment, the forward strand of the first sample tag is selected from at least one of the following 24 sets of sequences:
1) a sequence shown as SEQ ID NO.1-SEQ ID NO. 8;
2) a sequence shown as SEQ ID NO.9-SEQ ID NO. 16;
3) a sequence shown as SEQ ID NO.17-SEQ ID NO. 24;
4) a sequence shown as SEQ ID NO.25-SEQ ID NO. 32;
5) a sequence shown as SEQ ID NO.33-SEQ ID NO. 40;
6) a sequence shown as SEQ ID NO.41-SEQ ID NO. 48;
7) a sequence shown as SEQ ID NO.49-SEQ ID NO. 56;
8) a sequence shown as SEQ ID NO.57-SEQ ID NO. 64;
9) a sequence shown as SEQ ID NO.65-SEQ ID NO. 72;
10) a sequence shown as SEQ ID NO.73-SEQ ID NO. 80;
11) a sequence shown as SEQ ID NO.81-SEQ ID NO. 88;
12) a sequence shown as SEQ ID NO.89-SEQ ID NO. 96;
13) a sequence shown as SEQ ID NO.97-SEQ ID NO. 104;
14) a sequence shown as SEQ ID NO.105-SEQ ID NO. 112;
15) a sequence shown as SEQ ID NO.113-SEQ ID NO. 120;
16) a sequence shown as SEQ ID NO.121-SEQ ID NO. 128;
17) a sequence shown as SEQ ID NO.129-SEQ ID NO. 136;
18) a sequence shown as SEQ ID NO.137-SEQ ID NO. 144;
19) a sequence shown as SEQ ID NO.145-SEQ ID NO. 152;
20) a sequence shown as SEQ ID NO.153-SEQ ID NO. 160;
21) a sequence shown as SEQ ID NO.161-SEQ ID NO. 168;
22) a sequence shown as SEQ ID NO.169-SEQ ID NO. 176;
23) a sequence shown as SEQ ID NO.177-SEQ ID NO. 184;
24) the sequence shown in SEQ ID NO.185-SEQ ID NO. 192.
In one embodiment, the first sample tag can be selected from any 1, any 2, any 3, any 4, any 5, any 6, any 7, any 8, any 9, any 10, any 11, any 12, any 13, any 14, any 15, any 16, any 17, any 18, any 19, any 20, any 21, any 22, any 23 of the above 24 sequences, or the first sample tag can comprise any 24 of the above 24 sequences.
In one example, 8 sequences of each set are mixed in equimolar ratios for preparing linkers containing the forward strand of the first sample tag.
In one embodiment, the template molecule to be pooled is a single stranded DNA template molecule.
In one embodiment, if the original template molecule is in a non-single-stranded state, the method further comprises, before the dephosphorylation step, performing a melting process on the original template molecule to obtain a single-stranded DNA template molecule.
In one embodiment, the melting process is a denaturation process.
In one embodiment, the primary melting step, and/or the secondary melting step, and/or the denaturation treatment of the original template molecule, the denaturation treatment is a thermal denaturation treatment, and specifically, the original template molecule may be heated to at least 80 ℃ for at least 1 min.
In one embodiment, the heat denaturation treatment is carried out by heating the molecules to be denatured to at least 80 ℃ for at least 1 min.
In one embodiment, the heat denaturation treatment is carried out by heating the molecules to be denatured to 80-98 deg.C for 1-30 min.
In one embodiment, the dephosphorylation treatment step comprises adding a dephosphorylating enzyme to perform dephosphorylation treatment under the following conditions: at 20-30 ℃ for 5-30min, then carrying out denaturation treatment on dephosphorylation enzyme, wherein the reaction conditions are as follows: heating to at least 80 deg.C for at least 1 min.
In one embodiment, in the single melting step, the reverse strand of the first sample tag with the attached random sequence is dissociated from the linker sequence, streptavidin-coated magnetic beads are added, and the intermediate library molecules modified with the tag molecules are collected using a magnet.
In one embodiment, the DNA polymerase is selected from the group consisting of the Klenow fragment of DNA polymerase I, also denoted as DNA polymerase I Klenow fragment. The fragments may be purchased commercially.
In one embodiment, in the first sequencing linker ligation step and/or the second sequencing linker ligation step, the DNA ligase is selected from T4DNA ligase. The T4DNA ligase can be purchased from the market, and the cost of the T4DNA ligase is obviously lower than that of the T4 RNA ligase, so that the import is not relied on.
In one embodiment, the second sequencing linker includes, but is not limited to, any one of a P5-terminal sequencing linker of the Illumina sequencing platform, a P2-terminal sequencing linker of the MGI sequencing platform.
In one embodiment, the second sequencing adapter contains or does not contain a second molecular tag.
In one embodiment, FIG. 1 shows a single-stranded library construction method in which the second sequencing adapter does not comprise the second molecular-tagged single-stranded linker, i.e., a single-stranded library construction method based on a single-ended molecular-tagged adapter.
In one embodiment, FIG. 2 shows a single-stranded library construction method in which the second sequencing adapter comprises a second molecular tag, i.e., a single-stranded library construction method based on a double-ended molecular tag adapter.
In one embodiment, the second molecular tag is a random sequence.
In one embodiment, the second molecular tag is 4-19nt in length. Specifically, the number of the first and second groups may be 4nt, 5nt, 6nt, 7nt, 8nt, 9nt, 10nt, 11nt, 12nt, 13nt, 14nt, 15nt, 16nt, 17nt, 18nt, 19nt, and the like.
In one embodiment, in the step of primary double-strand synthesis, the reaction conditions are 65-75 ℃ and 1-3min in sequence; 30-37 deg.C, 5-30 min.
In one embodiment, in the second sequencing linker connection step, the reaction conditions are 20-30 ℃ and 15-60 min.
In one embodiment, the second primer contains or does not contain a second sample tag.
In one embodiment, the second exemplar label is a random sequence.
In one embodiment, the length of the second sample label is 4-10nt, and specifically may be 4nt, 5nt, 6nt, 7nt, 8nt, 9nt, 10nt, and so on.
In one embodiment, when the second primer does not contain the second sample tag, the second primer comprises an inner adaptor sequence, an outer adaptor sequence, wherein the 5 'end of the inner adaptor sequence is linked to the 3' end of the outer adaptor sequence, and wherein the inner adaptor sequence is reverse complementary to the first sequencing adaptor.
In one embodiment, when the second primer comprises the second sample tag, the second primer comprises an inner adaptor sequence and an outer adaptor sequence, wherein the 5 'end of the inner adaptor sequence is linked to the 3' end of the outer adaptor sequence, the inner adaptor sequence is reverse complementary to the first sequencing adaptor, and the second sample tag is located between the inner adaptor sequence and the outer adaptor sequence.
In one embodiment, the third primer contains or does not contain a third sample tag.
In one embodiment, the third sample label is a random sequence.
In one embodiment, the length of the third sample label is 4-10nt, and specifically may be 4nt, 5nt, 6nt, 7nt, 8nt, 9nt, 10nt, and so on.
In one embodiment, when the third primer does not contain the second sample tag, the third primer comprises an inner adaptor sequence, an outer adaptor sequence, wherein the 5 'end of the inner adaptor sequence is linked to the 3' end of the outer adaptor sequence, and wherein the inner adaptor sequence is reverse complementary to the second sequencing adaptor.
In one embodiment, when the third primer comprises a third sample tag, the second primer comprises an inner adaptor sequence and an outer adaptor sequence, the 5 'end of the inner adaptor sequence is linked to the 3' end of the outer adaptor sequence, the inner adaptor sequence is reverse complementary mateable to the second sequencing adaptor, and the third sample tag is located between the inner adaptor sequence and the outer adaptor sequence.
In one embodiment, the enzymatic purification steps required to prepare the hairpin linkers of the prior art are omitted and HPLC purification is used at the time of synthesis.
In one example, the prior art 9-step elution was reduced to a 3-step elution.
In one embodiment, the primer extension steps in the prior art are reduced from 25 ℃ for 5 minutes and 35 ℃ for 25 minutes to 30-37 ℃ for 5-30 min.
In one embodiment, as shown in fig. 3 and 4, 192 sample tags are provided, and 8 sequences in the same column can be mixed for the pooling of one sample, so as to effectively avoid snow blindness of a sequencer due to too single base type at the same sequencing reaction site.
The length of each sequence is 20nt, the 8 sample label sequences in the same column form a group (8 in the same column), and the group can be totally used for 24 groups.
Example 1
In this example, one example of the DNA sample is lung cancer ctDNA standard set 1% AF, 20 ng/. mu.L (Shenzhen JingyangGeneGen science and technology Limited, GW-OCTM 009). Wherein, AF refers to mutation frequency.
Another example of the DNA sample is a tumor SNV 2.5% gDNA standard, 50 ng/. mu.L (Shenzhen JingyangGen technology Limited, GW-OCTM 002).
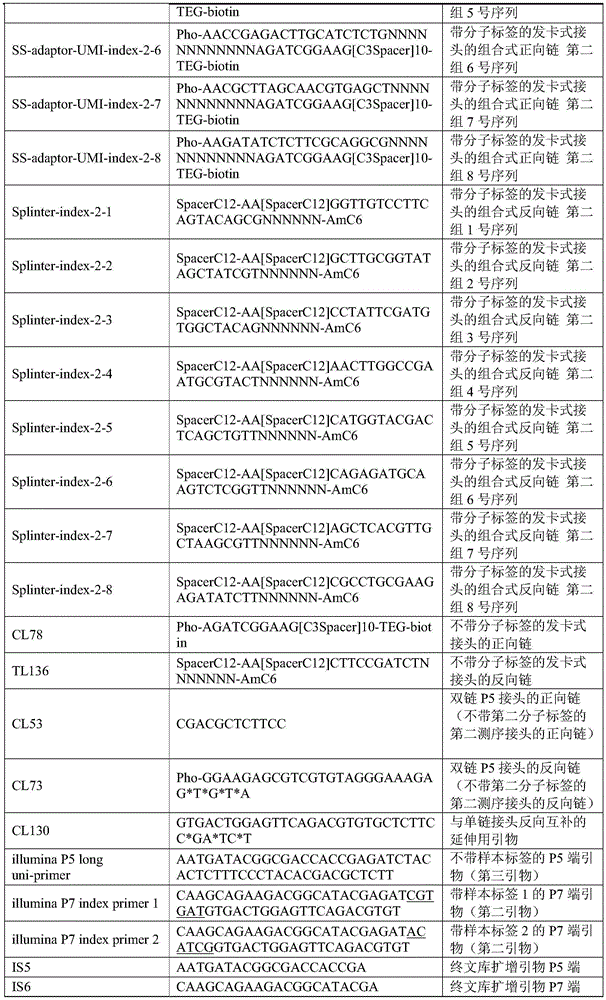
The padlock probes (synthesized by Kinsley Biotechnology Ltd., HPLC purification) are shown in Table 1.
TABLE 1
Remarking: 1. CL78, TL136, CL53, CL73, CL130, IS5, IS6 in the table are all cited from reference 1 and experiments were performed exactly as in this reference as experimental group 1 of this example (single-stranded building library without molecular tag); other oligonucleotides (oligos) were designed autonomously by the inventors for the single-stranded library building panel with molecular and sample tags, and CL53, CL73, CL130, IS5, IS6 were also used for this panel 1.
Reference 1: Marie-Theres Gansuge TG, Glocke I, Korlevic' P, Lippik L, Sarah Nagel LR, Schmidt A, Meyer M (2016) Single-stranded DNA library preparation from high stranded DNA using T4DNA library nucleic Acids Res.45(10): e79.
2. CL73, CL130, "+" indicates a disulfide bond to strengthen the linkage between nucleotides and prevent degradation of the polynucleotide.
3. "AmC 6" in Splint-index-x-y indicates an amino modification at the carbon position 6 to block the 3' end of the polynucleotide.
4. "Spacer C12" and "C3 Spacer" represent 12 and 3 empty carbon backbones, respectively, to prevent non-specific binding of primers.
5. "-TEG-biotin" indicates that TEG and biotin molecules are attached to the 3' end of the polynucleotide.
6. "Pho-" indicates that the 5' end of the polynucleotide carries a phosphate group.
7. in the illumina P7 index primer 1 and the illumina P7 index primer 2, the underlined sequences are sample labels.
T4DNA fragment (Rapid) (Cat: N103-01) (available from NyVo Nuo Zan Biotech Co., Ltd.) was used for each ligation reaction.
The library Amplification reaction was performed using VAHTS HiFi Amplification Mix (cat # N616-01) (available from Nanjing NuoZan Biotechnology Ltd.).
PCR product purification magnetic Beads VAHTS DNA Clean Beads (cat # N411-01) (available from Nanjing NuoWei Zan Biotech Co., Ltd.).
Library construction of conventional library construction technology group with molecular tag Using NanoPrepTMDNA library construction kit (for illumina) (available from Shanghai Naon Biotech Co., Ltd., cat. No. 1002101C 1).
The primer extension complementary to the single-stranded linker in the reverse direction was performed using DNA polymerase I Klenow fragment (cat # N104-01) (available from Nanjing Novowed Biotech Co., Ltd.).
T4 RNA ligase buffer (10X) and FastAP (1U/. mu.L) required for DNA sample dephosphorylation reaction were respectively adopted from NEB Limited (cat # B0216L) and from EnWeijie Jie (Shanghai) trade Limited (cat # EF 0651).
Dynabeads from streptavidin magnetic beads for binding single stranded ligation productsTMMyOneTMStreptavidin C1 (Yinxie Jie (Shanghai) trade Co., Ltd., Cat number 65001).
Ultra pure water of molecular biology grade used in each step of experiment is adoptedTMDNase/RNase-Free Distilled Water (Yinxie Jie Co., Ltd., Cat. 10977023).
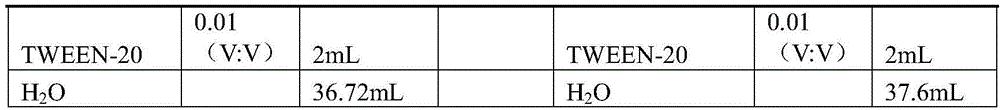
Other solutions used in the experiment were formulated as shown in Table 2 below (in a total volume of 40 mL).
TABLE 2
In Table 2, V represents the volume concentration, i.e., the volume concentration of a corresponding substance in an aqueous solution obtained after dissolving the substance in water, for example, a SDS0.2 (V: V) solution prepared by mixing a liquid pure SDS and pure water in a volume ratio of 2: 8 are mixed and prepared.
The "concentrations" in Table 2 each refer to the initial concentration of each component.
The instrument is ABI veriti96 type PCR instrument (product of Yinxie Jie based (Shanghai) trade Co., Ltd.); a constant temperature blending instrument (Hangzhou Yongning instruments Co., Ltd., Cat number HC-100); a four-dimensional rotary mixer (manufactured by shobel instruments ltd, haman, cat # BE-1100); magnetic force rack (Wuxi Baige Biotechnology Co., Ltd., product No. BMB 16-1.5-2); qubitTM4Fluorometer, with WiFi (England Weiji trading Limited, Cat # Q33238); a Bioptic full-automatic multiple nucleic acid detection system (Hangzhou Kyoze Biotech Co., Ltd., product number Qsep-100); an Eppendorf-brand pipette (available from Eppendort, Germany), a 1000. mu.L range, a 100. mu.L range and a 10. mu.L range.
The single-ended molecular tag library construction method of this example is shown in FIG. 1.
The experimental procedure was as follows:
1. single-chain library construction experimental group with molecular label (marked as experimental group 1)
1.1 preparation of hairpin linkers with molecular and sample labels
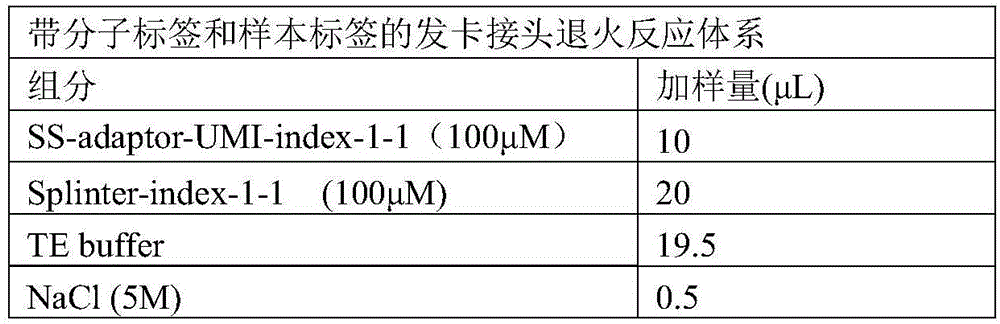
1.1.1 the following reaction system was placed in a 200. mu.L PCR tube.
TABLE 3
The combined forward chain SS-adaptor-UMI-index-x-y of the hairpin type joint with the molecular label and the combined reverse chain Splint-index-x-y of the hairpin type joint with the molecular label are respectively added into the reaction system in a one-to-one correspondence mode according to the group number (x) and the sequence number (y), and each pair is respectively provided with a reaction system, and the two groups are 16 pairs in total, so that the reaction system is a 16 hairpin joint annealing reaction system with the molecular label and the sample label.
1.1.2 annealing reaction conditions were as follows: 95 ℃ for 10 seconds; RAMP 4% was then added so that the reaction temperature was reduced to 14 ℃ at a rate of 0.1 ℃/sec.
1.1.3 to the above reaction product (50. mu.L), 50. mu.L of TE buffer was added to obtain a final concentration of SS-adapter-UMI-index-x-y/Splint-index-x-y of 10/20. mu.M, i.e., 0.5. mu.M.
The obtained product can be stored at-20 deg.C for a long time, or at 4 deg.C for 8 hr.
1.2 preparation of double-stranded ligation Tab (CL53/CL73)
1.2.1 the following reaction system was placed in a 200. mu.L PCR tube.
TABLE 4
1.2.2 annealing reaction conditions: 95 ℃ for 10 seconds; then adding RAMP 4%, and cooling to 14 deg.C at 0.1 deg.C/s.
1.2.3 to the above reaction product (50. mu.L) was added 50. mu.L of TE buffer to obtain a double-stranded linker (denoted as CL53/CL73) at a final concentration of 100. mu.M.
The obtained product can be stored at-20 deg.C for a long time, or at 4 deg.C for 8 hr.
1.3 Single-stranded DNA template preparation and splint linker ligation
1.3.1 the following phosphorylation reaction system was placed in a 200. mu.L PCR tube.
TABLE 5
Two DNA samples were prepared, namely 0.5. mu.L lung cancer ctDNA standard set 1% AF, 20 ng/. mu.L (GW-OCTM009) and 0.5. mu.L tumor SNV 2.5% gDNA standard, 50 ng/. mu.L (GW-OCTM 002).
1.3.2 pipette, blow, mix, centrifuge briefly.
1.3.3 dephosphorylation reaction conditions: 10 minutes at 37 ℃; at 95 ℃ for 2 minutes; immediately placed in an ice bath for 5 minutes.
1.3.4 to the above reaction product, the following components were added as a hairpin linker ligation reaction system.
TABLE 6
| Components | Sample addition amount (μ L) |
| PEG 8000(50%) | 32 |
| ATP(100mM) | 0.4 |
| SS-adaptor-UMI-index-x-y/Splinter-index- |
1 |
| T4 DNA ligase(600U/μL) | 1 |
| Total volume | 34.4 |
1.3.5 pipette, blow, mix well, and centrifuge briefly.
1.3.6 hairpin linker ligation reaction conditions: 1h at 37 ℃; 1 minute at 95 ℃; removed quickly and placed in an ice bath for at least 5 minutes.
1.4 streptavidin magnetic bead binding single-stranded ligation products
1.4.1 washing magnetic beads (each reaction was prepared separately), the specific steps were as follows:
A) the beads were removed from the freezer and allowed to equilibrate for 30 minutes at room temperature.
B) After mixing, the magnetic beads (20. mu.L/rxn) were put into a 1.5mL tube and placed on a magnetic stand.
C) The supernatant was discarded and washed twice with 500. mu.L of 1xBWT + SDS.
D) Add 1xBWT + SDS to dissolve back (250. mu.L/rxn).
1.4.2 adding the single-chain ligation product stored in the ice bath of 1.3 into the magnetic bead suspension of (A), blowing and uniformly mixing, incubating for 20-60 minutes at room temperature, and simultaneously rotating and uniformly mixing.
1.4.3 the product was centrifuged briefly, placed on a magnetic stand, allowed to stand for 3 minutes, the supernatant was discarded, 200. mu.L of 0.1xBWT + SDS was added, and centrifuged briefly.
1.4.4 Place 1.5ml tube in the thermostatic mixer, shake for 8 seconds, 1500rpm, 25 ℃.
1.4.5 the 1.5mL tube was removed from the thermostatic mixer and placed in a magnetic stand, allowed to stand for 3 minutes, the supernatant was aspirated off, and the agglomerated magnetic beads were retained.
1.5 stringent elution
1.5.1 Add 100. mu.L of SWB and vortex to mix.
1.5.2 placing the mixture in a constant temperature mixing instrument (preheating to 45 ℃), and incubating (SW).
1.5.3 constant temperature mixer program as follows: time (total length): 3 minutes; temperature: 45 ℃; mixing time (Mixing time per time): 15 s; pause time): 45 s.
1.5.4 the 1.5mL tube was removed from the thermostatic mixer and placed in a magnetic stand, and left to stand for 3 minutes, the supernatant was aspirated off, and the agglomerated magnetic beads were retained. The thermostatic mixer was adjusted to 25 ℃.
1.6 magnetic bead elution
Add 200. mu.L of 0.1xBWT and place the 1.5mL tube on a thermostatic mixer and shake for 8 seconds at 1500 rpm.
1.7 primer extension reverse complementary to Single Strand
1.7.1 the following primer annealing reaction system was placed in a 200. mu.L PCR tube.
TABLE 7
| Components | Sample addition amount (μ L) |
| H2O | 28 |
| Klenow reaction buffer(10X) | 5 |
| dNTP(10mM each) | 5 |
| TWEEN-20(1%) | 2.5 |
| CL130(100μM) | 5.5 |
| Total amount of | 46 |
1.7.2 in 1.5mL tube for short centrifugation, placed on the magnetic frame, and left for 3 minutes, discard the supernatant.
1.7.3 adding the primer annealing reaction system, uniformly mixing by vortex, placing on a constant-temperature mixing instrument (preheating to 65 ℃), and incubating for 2 minutes.
1.7.4 quickly place 1.5mL tube in an ice bath for 5 minutes.
1.7.5 mu.L of Klenow fragment (5U/. mu.L) was added to the tube.
1.7.6 incubation in a constant temperature mixer, the procedure of the constant temperature mixer is as follows: time (total length): 15 minutes; temperature: 35 ℃; mixing time (Mixing time per time): 15 s; pause time): 45 s.
1.8 elution after extension
1.8.1 Place 1.5mL tube on magnetic stand, let stand for 3 minutes, discard supernatant, add 100. mu.L SWB.
1.8.2 putting the mixture into a constant-temperature blending instrument for incubation (SW); the constant temperature mixer program was as follows: time (total length): 3 minutes; temperature: 45 ℃; mixing time (Mixing time per time): 15 s; pause time): 45 s.
1.8.3 placing 1.5mL tube on magnetic frame, standing for 3min, discarding supernatant, and adding 200 μ L0.1 xBWT;
1.8.4 is put into a constant temperature mixing instrument (preheated to 25 ℃), is at 1500rpm and is vibrated for 8 seconds;
1.8.5 centrifuging for a short time, placing on a magnetic frame, standing for 3min, and discarding the supernatant.
1.9 Dual Link connection
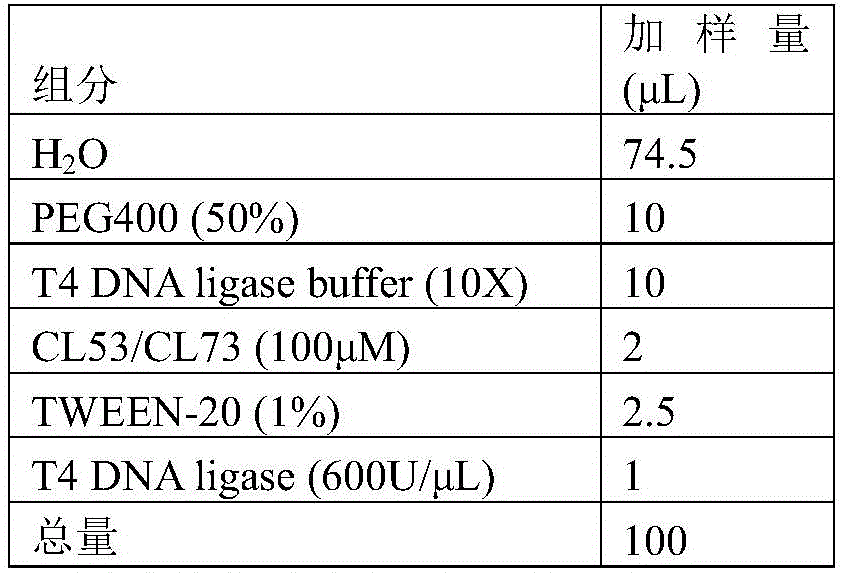
1.9.1 the following ligation system was placed in a 200. mu.L PCR tube.
TABLE 8
1.9.2 the beads from the supernatant from the previous step were resuspended in the ligation batch and vortexed.
1.9.3 was placed in a thermostatic mixer (pre-heated to 22 ℃ C.), at 1500rpm, for 3 seconds, incubated at 22 ℃ C., and the following program was started: time (total length): 1 h; temperature: 22 ℃; mixing time (Mixing time per time): 15 s; pause time): 45 s.
1.10 elution after ligation
1.10.1 the 1.5mL tube was placed on a magnetic stand and allowed to stand for 3 minutes, the supernatant was discarded, 100. mu.L of SWB was added, vortexed and mixed, and placed in a thermostatic mixer (preheated to 45 ℃).
1.10.2 SW program was started and the program of the constant temperature mixer was as follows: time (total length): 3 minutes; temperature: 45 ℃; mixing time (Mixing time per time): 15 s; pause time): 45 s.
1.10.3 taking out, centrifuging for a short time, placing on a magnetic frame, standing for 3 minutes, removing supernatant, and adjusting to 25 ℃ by a constant-temperature mixer.
1.10.4 mu.L of 0.1xBWT was added, resuspended, and placed on a homomixer at 1500rpm for 8 seconds.
1.11 Single Strand library elution
1.11.1 taking out from the constant-temperature mixer, centrifuging for a short time, placing on a magnetic frame, standing for 3 minutes, and discarding the supernatant.
1.11.2 mu.L of EBT was added, and the mixture was put into a homomixer (preheated to 25 ℃ C.), at 1500rpm for 8 seconds.
1.11.3, the magnetic bead suspension was removed and transferred to 8 PCR tubes.
1.11.4 was placed in a PCR machine (95 ℃ C., 1 minute), removed, centrifuged briefly, placed on a magnetic stand, and allowed to stand for 3 minutes.
1.11.57 minutes later, the supernatant was transferred to a new eight-row tube.
1.12illumina index PCR
1.12.1 the following reaction system was placed in a 200. mu.L PCR tube.
TABLE 9
1.12.2 the following program was run on the PCR machine:
watch 10
| Hold (Hold) | 95℃ | 2min |
| 7cycles (cycle) | 98℃ | 20 |
| 60℃ | 45 |
|
| 72℃ | 30 seconds | |
| Hold (Hold) | 72℃ | 3min |
| Hold (Hold) | 4℃ | - |
1.13PCR product purification
1.13.1 mu.L of magnetic Beads of VAHTS DNA Clean Beads were added to the PCR product and mixed by pipetting.
1.13.2 and standing for 3min, discarding the supernatant, adding 200 μ L of 75% ethanol, and standing for 30 s.
1.13.3 discard the supernatant, add 200. mu.L of 80% ethanol, and let stand for 30 seconds.
1.13.4 discard the supernatant and aspirate residual alcohol with a small pipette.
And (3) incubating at 1.13.537 ℃ for 2-3min until the surface of the magnetic beads is matt (the magnetic beads should not be cracked).
1.13.6 adding 25 μ L water, blowing, mixing, and standing at room temperature for 5 min.
1.13.7 were allowed to stand on a magnetic rack for 3 minutes and the supernatant was transferred to a new PCR tube.
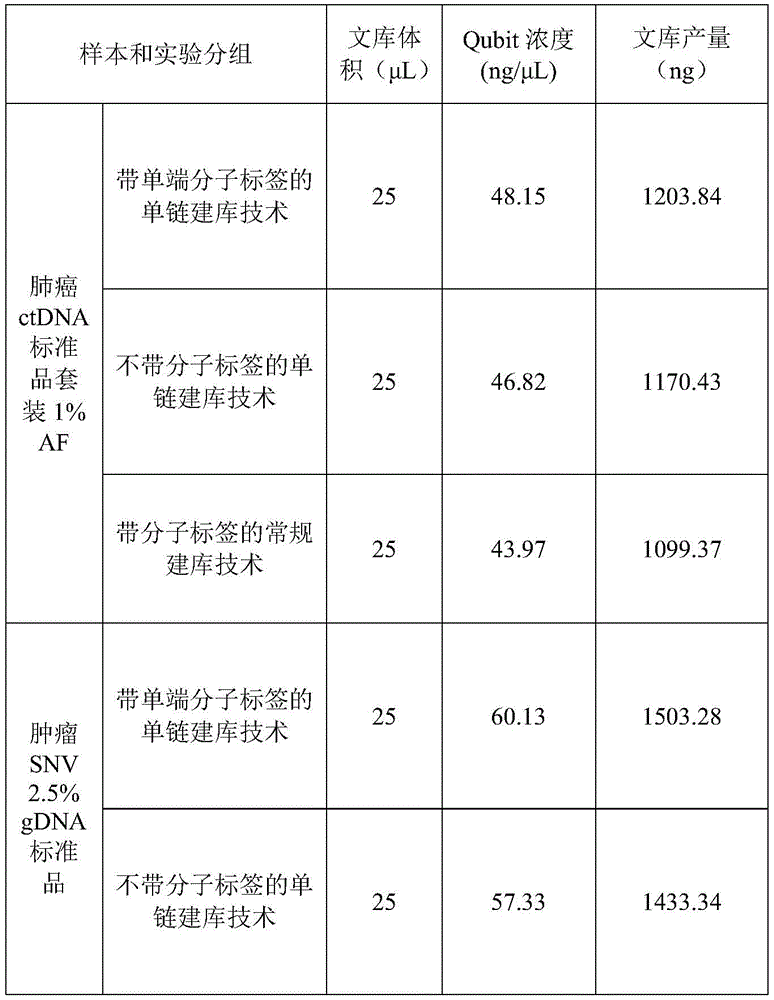
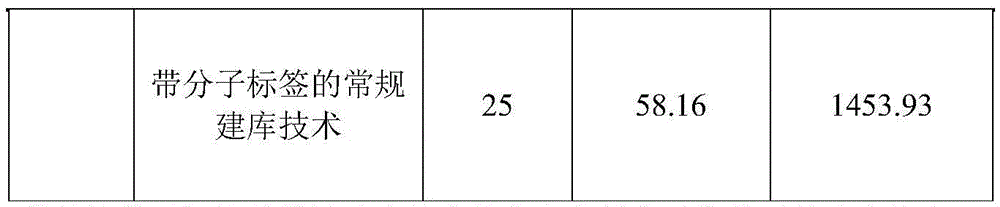
1.14 quality control of the library
Quibit 4.0HS was quantified and the concentrations recorded are as follows:
TABLE 11
As can be seen from the table above, the library yield of the single-chain library construction technology with the single-ended molecular tag is higher than that of the other two library construction technologies, which reflects that the number of the original template molecules put in the library is more and the library construction efficiency is higher.
1.15 on-machine sequencing
The model number of the instrument is illumina Hiseq 4000, the strategy is PE150, and the data volume is 3Gb per sample.
2. Single-chain library construction control group without molecular label (marked as control group 1)
The samples in this control group were identical to those in group 1 (single-stranded library construction group with molecular tag, i.e., group 1), and two DNA samples were taken, 2.5. mu.L lung cancer ctDNA standard set 1% AF, 20 ng/. mu.L (GW-OCTM009), 1. mu.L tumor SNV 2.5% gDNA standard, and 50 ng/. mu.L (GW-OCTM 002). Experiments were performed using CL78, TL136, CL53, CL73, CL130, illumina P5long uni-primer, illumina P7 index primer 1 and 2 (all cited in reference 1 below) described in table 1, and strictly following reference 1 as a control (the main steps include hairpin ligation without molecular tag, primer extension, double-stranded ligation, library index PCR and quality control), and designated as control 1.
Reference 1: Marie-Theres Gansuge TG, Glocke I, Korlevic' P, Lippik L, Sarah Nagel LR, Schmidt A, Meyer M (2016) Single-stranded DNA library preparation from high stranded DNA using T4DNA library nucleic Acids Res.45(10): e79.
3. Conventional library construction with molecular label (record as contrast group 2)
The samples of the control group are the same as the experiment of the group 1 (the single-chain library building experiment group with molecular tags, namely the experiment group 1), two DNA samples are taken, and the samples are respectively 2.5 mu L lung cancer ctDNA standard set 1% AF, 20 ng/mu L (GW-OCTM009) and 1 mu L tumor SNV 2.5% gDNA standard and 50 ng/mu L (GW-OCTM 002); using NanoPrepTMA DNA library construction kit (for illumina) (available from Shanghai Naon Biotechnology Ltd., cat # 1002101C1) was used as a control group 2 by performing library construction (including end repair and A-addition reaction, analysis of tag y-type linker ligation and library amplification) strictly in compliance with the product instructions.
4. Analysis of results
4.1 sequencing data quality control and analysis procedure
The original treatment was performed using fastp software, the genome alignment was performed using bwa-mem software, the reference genome was performed using GRCh38/hg38 (International Universal human genome reference sequence), and the markdup was performed using sambamba software.
4.2 results of analysis
TABLE 12
In Table 12, the Duplicate rate indicates the redundancy rate of the sequencing results.
Genome coverage by > -1 read indicates the percentage of the Genome that was covered by at least one sequencing.
QC pass ratio represents data quality control qualification rate.
The alignnemnt ratio represents the proportion of the sequencing results that is available for alignment analysis.
index-false assignment represents the inter-sample label crosstalk ratio.
As can be seen from Table 12, the method of this example can achieve higher average sequencing depth and lower cross-talk rate of sample-to-sample tags in the whole genome range with the same amount of sequencing data, compared to the single-strand library construction without molecular tags and the conventional library construction with molecular tags.
Example 2
In the same manner as in example 1, one example of the DNA sample is lung cancer ctDNA standard set 1% AF, 20 ng/. mu.L (Shenzhen Jingyan gene science and technology Limited, GW-OCTM 009).
Another example of the DNA sample is a tumor SNV 2.5% gDNA standard, 50 ng/. mu.L (Shenzhen JingyangGen technology Limited, GW-OCTM 002).
The padlock probes (synthesized by Kinsley Biotechnology Ltd., HPLC purification) are shown in Table 1.
Unless otherwise stated, the apparatus and reagents used in this example were the same as those used in example 1.
In this example, the whole of the hybridization capture experiment was performed by using SureSelectXT human whole exon V6 Plus2, 16 times of reactions (Agilent technologies, Inc., cat # 5190-8869).
The experimental procedure was as follows:
1. single-chain library construction experimental group with molecular label (marked as experimental group 1)
Step 1.1 to step 1.13 were carried out with reference to example 1.
In step 1.3, the system is as follows.
Two DNA samples, 10 μ L lung cancer ctDNA standard set 1% AF, 20ng/μ L (GW-OCTM 009); and 5. mu.L of tumor SNV 2.5% gDNA standard, 50 ng/. mu.L (GW-OCTM 002).
1.14 Pre-Capture library quality control
Quibit 4.0HS was quantified and the concentrations recorded as follows.
TABLE 14
As can be seen from the table above, the library yield of the single-chain library construction technology with the single-ended molecular tag is higher than that of the other two library construction technologies, which reflects that the number of the original template molecules put in the library is more and the library construction efficiency is higher.
1.15 hybrid Capture
500ng of the library from the previous experiment was used for hybrid capture, and the experimental module used the whole set of SureSelectXT whole exon V6 Plus2, 16 reactions (Agilent technologies, Inc.: 5190-8869). The library is hybridized with probes following exactly the instructions of the product, the captured products are PCR amplified by using magnetic beads and washed, and the PCR products are purified (the purification step is the same as the step 1.13).
1.16 Final library quality control after Capture
Quibit 4.0HS was quantified and the concentrations recorded as follows.
As can be seen from the table above, the library yield of the single-chain library construction technology with the single-ended molecular tag is higher than that of the other two library construction technologies, which reflects that the number of the original template molecules put in the library is more and the library construction efficiency is higher.
1.17 sequencing on machine
The model number of the instrument is illumina Hiseq 4000, the strategy is PE150, and the data volume is 25Gb per sample.
2. Single-chain library construction experimental group without molecular label
As with the experiment of group 1 (single-stranded library with molecular tag, i.e., experiment group 1), two DNA samples were taken, 10. mu.L lung cancer ctDNA standard set 1% AF, 20 ng/. mu.L (GW-OCTM009) and 5. mu.L tumor SNV 2.5% gDNA standard, 50 ng/. mu.L (GW-OCTM 002). CL78, TL136, CL53, CL73, CL130, illumina P5long uni-primer, illumina P7 index primer 1 and 2 described in Table 1 were all cited from reference 1 and experiments were performed exactly as in reference 1 as controls for this example (the main steps included hairpin ligation without molecular tag, primer extension, double-stranded ligation, library index PCR and quality control). The hybridization capture module was performed in its entirety using SureSelectXT human full exon V6 Plus2, 16 reactions (Agilent technologies, Inc., cat # 5190-8869) and following the procedures of the product instructions.
3. Conventional library construction with molecular tags
As with the experiment of group 1 (single-stranded library with molecular tag, i.e., experiment group 1), two DNA samples were taken, 10. mu.L lung cancer ctDNA standard set 1% AF, 20 ng/. mu.L (GW-OCTM009) and 5. mu.L tumor SNV 2.5% gDNA standard, 50 ng/. mu.L (GW-OCTM 002). Using NanoPrepTMDNA library construction kit (for illumina) (available from Shanghai Naon Biotechnology Ltd., cat. No. 1002101C1) was subjected to library construction (including end repair and addition of A, analytical tag y-type linker ligation and library amplification) strictly according to the product instructions. The hybridization capture module was performed in its entirety using SureSelectXT human full exon V6 Plus2, 16 reactions (Agilent technologies, Inc., cat # 5190-8869) and following the procedures of the product instructions.
4, analysis of results
4.1 sequencing data quality control and analysis procedure
The original processing was performed using fastp software, the genome alignment was performed using bwa-mem software, the reference genome was performed using GRCh38 (also denoted hg38, international universal human genome reference sequence), and the markdup was performed using sambamba software.
4.2 results of analysis
TABLE 16
In table 16, Q30_ rate represents the proportion of sequencing data that achieves 99.9% sequencing accuracy.
gc _ content represents the sum of the G and C base content in the sequencing results.
Mapping _ rate represents the proportion of sequencing results that can be used for alignment analysis.
The duration _ rate represents the redundancy rate of the sequencing results.
The capture _ rate represents the efficiency of target gene region enrichment.
Depth _ in _ target represents the average sequencing Depth of the target gene region.
index false-assignment indicates the sample label crosstalk ratio between samples.
As can be seen from Table 16, the method of this example can achieve higher sequencing depth and lower sample-to-sample tag crosstalk ratio in the target region under the same amount of sequencing data compared to the single-strand library without molecular tag and the conventional library construction with molecular tag.
Reference documents:
[1]Marie-Theres Gansauge TG,Glocke I,Korlevic′P,Lippik L,Sarah Nagel LR,Schmidt A,Meyer M(2016)Single-stranded DNA library preparation from highly degraded DNA using T4 DNA ligase.Nucleic Acids Res.45(10):e79.
[2].Gansauge,M.T.and Meyer,M.(2013)Single-stranded DNA library preparation for the sequencing of ancient or damaged DNA.Nat.
Protoc.,8,737–748.
[3].Qing Wang et al.Targeted sequencing of both DNA strands barcoded and captured individually by RNA probes to identify genome-wide ultra-rare mutations.Scientific Reports.2017,7:3356.DOI:10.1038/s41598-017-03448-8.
the present invention has been described in terms of specific examples, which are provided to aid understanding of the invention and are not intended to be limiting. For a person skilled in the art to which the invention pertains, several simple deductions, modifications or substitutions may be made according to the idea of the invention.
Organization Applicant
----------------------
Street :
City :
State :
Country :
PostalCode :
PhoneNumber :
FaxNumber :
EmailAddress :
<110> organization Name Shenzhen Rui method Biotech limited
Application Project
-------------------
<120> Title, adaptor with molecular label and sample label and single-chain library construction method thereof
<130> AppFileReference : 20I30444
<140> CurrentAppNumber :
<141> CurrentFilingDate : ____-__-__
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 1
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 2
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 3
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
agtgtcattc cagagtcagg 20
<212> Type : DNA
<211> Length : 20
SequenceName : 4
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 5
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 6
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 7
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 8
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 9
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 10
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 11
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 12
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 13
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 14
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 15
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
aagatatctc ttcgcaggcg 20
<212> Type : DNA
<211> Length : 20
SequenceName : 16
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 17
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 18
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 19
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 20
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 21
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
acgatgcttg acgagatctc 20
<212> Type : DNA
<211> Length : 20
SequenceName : 22
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 23
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 24
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
agatccagca cgtatgagtc 20
<212> Type : DNA
<211> Length : 20
SequenceName : 25
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 26
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 27
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 28
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 29
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
caaccacatc gagctgtgtt 20
<212> Type : DNA
<211> Length : 20
SequenceName : 30
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 31
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
caatgttatc ggaaccggtc 20
<212> Type : DNA
<211> Length : 20
SequenceName : 32
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 33
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 34
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 35
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 36
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 37
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 38
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 39
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 40
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 41
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 42
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 43
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 44
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 45
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 46
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 47
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
gagcgacact actgtgaatc 20
<212> Type : DNA
<211> Length : 20
SequenceName : 48
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 49
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 50
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
gcacgaattc tcgaacatgg 20
<212> Type : DNA
<211> Length : 20
SequenceName : 51
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 52
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 53
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
ggaagaacgt acgtgtcttc 20
<212> Type : DNA
<211> Length : 20
SequenceName : 54
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
ggtgcgaacc ttgatcaatc 20
<212> Type : DNA
<211> Length : 20
SequenceName : 55
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 56
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 57
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 58
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 59
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 60
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 61
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 62
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 63
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
tgaagagagg tctcacactc 20
<212> Type : DNA
<211> Length : 20
SequenceName : 64
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 65
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 66
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 67
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 68
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 69
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
acggatctct aacgaggatc 20
<212> Type : DNA
<211> Length : 20
SequenceName : 70
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 71
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
aatccacgtc tcgcaatagg 20
<212> Type : DNA
<211> Length : 20
SequenceName : 72
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 73
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 74
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 75
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 76
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 77
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 78
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 79
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
atcaacagcg agtagcgatc 20
<212> Type : DNA
<211> Length : 20
SequenceName : 80
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 81
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 82
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 83
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 84
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 85
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 86
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 87
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 88
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 89
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 90
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 91
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 92
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
gactacggtc catacgatgt 20
<212> Type : DNA
<211> Length : 20
SequenceName : 93
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 94
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 95
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
gcgagtcgtc agcatcttac 20
<212> Type : DNA
<211> Length : 20
SequenceName : 96
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
tctgcactac ggcagctatc 20
<212> Type : DNA
<211> Length : 20
SequenceName : 97
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 98
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 99
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 100
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 101
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 102
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 103
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
gcaggtattc tcgatcagac 20
<212> Type : DNA
<211> Length : 20
SequenceName : 104
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 105
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
agatagcgat tgatgccgtc 20
<212> Type : DNA
<211> Length : 20
SequenceName : 106
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 107
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 108
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 109
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
ttcgcaattc acagcgcagt 20
<212> Type : DNA
<211> Length : 20
SequenceName : 110
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
aacagttcta ggtcgcagat 20
<212> Type : DNA
<211> Length : 20
SequenceName : 111
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
gtgaatgatg cggtaccatc 20
<212> Type : DNA
<211> Length : 20
SequenceName : 112
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 113
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 114
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 115
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 116
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 117
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 118
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 119
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 120
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 121
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 122
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 123
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 124
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 125
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 126
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 127
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 128
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 129
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
gtccaacatc acttagtgcg 20
<212> Type : DNA
<211> Length : 20
SequenceName : 130
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 131
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 132
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 133
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 134
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 135
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 136
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 137
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 138
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 139
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
gcacgatatc tgcggtaatc 20
<212> Type : DNA
<211> Length : 20
SequenceName : 140
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 141
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 142
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 143
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 144
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 145
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 146
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 147
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 148
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 149
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 150
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 151
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 152
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
gaacactgca ggatgatctc 20
<212> Type : DNA
<211> Length : 20
SequenceName : 153
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 154
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 155
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 156
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 157
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 158
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 159
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 160
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 161
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 162
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 163
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 164
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
cggtctcctc atcgtgttaa 20
<212> Type : DNA
<211> Length : 20
SequenceName : 165
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 166
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 167
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
cgagcctaat tggaagtgtc 20
<212> Type : DNA
<211> Length : 20
SequenceName : 168
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 169
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 170
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
aagtgaacgc tgatgacctc 20
<212> Type : DNA
<211> Length : 20
SequenceName : 171
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 172
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 173
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 174
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 175
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 176
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 177
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 178
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 179
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 180
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 181
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 182
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 183
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 184
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 185
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 186
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 187
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 188
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 189
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 190
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
<212> Type : DNA
<211> Length : 20
SequenceName : 191
SequenceDescription :
Sequence
--------
<213> OrganismName : Artificial Sequence
<400> PreSequenceString :
gagtgatgaa tcgctacgtc 20
<212> Type : DNA
<211> Length : 20
SequenceName : 192
SequenceDescription :
Claims (10)
1. The linker with the molecular tag and the sample tag is characterized by comprising a first molecular tag, a first sample tag and a first sequencing linker, wherein the first sample tag is provided with a reverse complementary forward strand and a reverse strand, the first molecular tag is a single-stranded DNA molecule, the 3 'end of the first molecular tag is connected to the 5' end of the first sequencing linker, the 5 'end of the first molecular tag is connected to the 3' end of the forward strand of the first sample tag, the 5 'end of the forward strand of the first sample tag is used for being connected to a single-stranded template molecule to be built, and the 3' end of the reverse strand of the first sample tag is connected with a random sequence.
2. The linker of claim 1 wherein the 5' end of the forward strand of the first sample tag is modified with a phosphate group.
3. The linker of claim 1 wherein the first molecular tag is a random sequence;
optionally, the first molecular tag is 4-19nt in length.
4. The linker of claim 1 wherein the 3' end of the first sequencing linker is modified with a labeling molecule;
optionally, the marker molecule is selected from at least one of biotin and TEG biotin;
optionally, the first sequencing linker is a single-stranded DNA molecule;
optionally, the library building template molecules to be built are single-stranded DNA molecules.
5. The linker of claim 1 wherein the forward strand of the first sample tag is selected from at least one of the sequences set forth in SEQ ID No.1 to SEQ ID No. 192;
optionally, the forward strand of the first sample tag is selected from any one of the sequences shown in SEQ ID NO.1-SEQ ID NO. 192.
6. The linker of claim 1 wherein the random sequence attached to the 3' end of the reverse strand of the first sample tag is 5-20nt in length;
optionally, the first sequencing linker is selected from any one of a P7 terminal sequencing linker of Illumina sequencing platform, a P1 terminal sequencing linker of MGI sequencing platform;
optionally, the template molecules to be pooled are from severely degraded samples and/or sulfite treated samples.
7. A kit for library construction comprising the linker of any one of claims 1 to 6.
8. The kit of claim 7, further comprising a second sequencing linker for linking to an intermediate library molecule;
optionally, a second molecular tag is or is not attached to one end of the second sequencing adapter for attachment to the intermediate library molecule;
optionally, the second molecular tag is a random sequence;
optionally, the second molecular tag is 4-19nt in length;
optionally, the second sequencing linker is selected from any one of a P5 terminal sequencing linker of Illumina sequencing platform, a P2 terminal sequencing linker of MGI sequencing platform;
optionally, a second primer and/or a third primer is also included;
optionally, the second primer contains or does not contain a second sample tag;
optionally, when the second primer does not contain a second sample tag, the second primer comprises an inner adaptor sequence, an outer adaptor sequence, the 5 'end of the inner adaptor sequence being linked to the 3' end of the outer adaptor sequence, the inner adaptor sequence being reverse complementary to the first sequencing adaptor;
optionally, when the second primer comprises a second sample tag, the second primer further comprises an inner adaptor sequence and an outer adaptor sequence, wherein the 5 'end of the inner adaptor sequence is linked to the 3' end of the outer adaptor sequence, the inner adaptor sequence is reverse complementary to the first sequencing adaptor, and the second sample tag is located between the inner adaptor sequence and the outer adaptor sequence;
optionally, the third primer contains or does not contain a third sample tag;
optionally, when the third primer does not contain a second sample tag, the third primer comprises an inner adaptor sequence, an outer adaptor sequence, the 5 'end of the inner adaptor sequence being linked to the 3' end of the outer adaptor sequence, the inner adaptor sequence being reverse complementary to the second sequencing adaptor;
optionally, when the third primer comprises a third sample tag, the second primer comprises an inner adaptor sequence and an outer adaptor sequence, the 5 'end of the inner adaptor sequence is linked to the 3' end of the outer adaptor sequence, the inner adaptor sequence is reverse complementary mateable to the second sequencing adaptor, and the third sample tag is located between the inner adaptor sequence and the outer adaptor sequence.
9. A single-strand library construction method with a molecular tag and a sample tag is characterized by comprising the following steps:
dephosphorizing, namely dephosphorizing the template molecules of the library to be built;
a first sequencing adaptor connection step, which comprises connecting the adaptor of any one of claims 1-6 to a single-stranded template molecule to be built by DNA ligase, wherein the 3' end of the first sequencing adaptor is modified with a marker molecule;
a first melting step comprising denaturing the product of the first sequencing linker ligation step, allowing the reverse strand of the first sample tag to dissociate from the linker, and collecting the intermediate library molecules modified with the tag molecules;
a primary double-stranded synthesis step, comprising mixing a first primer, a DNA polymerase and the intermediate library molecule modified with a marker molecule, allowing the first primer to anneal and extend a first sequencing linker part on the intermediate library molecule, and reacting to obtain a double-stranded blunt-ended DNA molecule, wherein the first primer contains a sequence complementary to the first sequencing linker;
a second sequencing adaptor connection step, which comprises mixing the double-stranded blunt-end DNA molecules with a second sequencing adaptor and DNA ligase, and reacting to obtain double-stranded DNA library molecules connected with the second sequencing adaptor, wherein the second sequencing adaptor is a double-stranded sequencing adaptor;
a secondary unzipping step comprising denaturing the double-stranded DNA library molecules such that the double-stranded DNA library molecules dissociate into single-stranded DNA molecules, collecting single-stranded DNA library molecules that are not modified with a marker molecule;
and a second double-stranded synthesis step, which comprises adding a second primer and a third primer, and amplifying to obtain a double-stranded DNA library for sequencing, wherein the second primer contains a sequence of a first sequencing joint complementary to the single-stranded DNA library molecule without the modified labeled molecule, and the third primer contains a sequence of a second sequencing joint complementary to the single-stranded DNA library molecule without the modified labeled molecule.
10. The single-strand library construction method of claim 9, wherein the forward strand of the first sample tag is selected from at least one of the sequences shown in SEQ ID No.1 to SEQ ID No. 192;
optionally, the forward strand of the first sample tag is selected from at least one of the following sets of 24 sequences:
1) a sequence shown as SEQ ID NO.1-SEQ ID NO. 8;
2) a sequence shown as SEQ ID NO.9-SEQ ID NO. 16;
3) a sequence shown as SEQ ID NO.17-SEQ ID NO. 24;
4) a sequence shown as SEQ ID NO.25-SEQ ID NO. 32;
5) a sequence shown as SEQ ID NO.33-SEQ ID NO. 40;
6) a sequence shown as SEQ ID NO.41-SEQ ID NO. 48;
7) a sequence shown as SEQ ID NO.49-SEQ ID NO. 56;
8) a sequence shown as SEQ ID NO.57-SEQ ID NO. 64;
9) a sequence shown as SEQ ID NO.65-SEQ ID NO. 72;
10) a sequence shown as SEQ ID NO.73-SEQ ID NO. 80;
11) a sequence shown as SEQ ID NO.81-SEQ ID NO. 88;
12) a sequence shown as SEQ ID NO.89-SEQ ID NO. 96;
13) a sequence shown as SEQ ID NO.97-SEQ ID NO. 104;
14) a sequence shown as SEQ ID NO.105-SEQ ID NO. 112;
15) a sequence shown as SEQ ID NO.113-SEQ ID NO. 120;
16) a sequence shown as SEQ ID NO.121-SEQ ID NO. 128;
17) a sequence shown as SEQ ID NO.129-SEQ ID NO. 136;
18) a sequence shown as SEQ ID NO.137-SEQ ID NO. 144;
19) a sequence shown as SEQ ID NO.145-SEQ ID NO. 152;
20) a sequence shown as SEQ ID NO.153-SEQ ID NO. 160;
21) a sequence shown as SEQ ID NO.161-SEQ ID NO. 168;
22) a sequence shown as SEQ ID NO.169-SEQ ID NO. 176;
23) a sequence shown as SEQ ID NO.177-SEQ ID NO. 184;
24) a sequence shown as SEQ ID NO.185-SEQ ID NO. 192;
optionally, 8 sequences of each set are mixed in equimolar ratios for preparing linkers containing the forward strand of the first sample tag;
optionally, the template molecules to be pooled are from severely degraded samples and/or sulfite-treated samples;
optionally, the template molecule of the library to be built is a single-stranded DNA template molecule;
optionally, if the original template molecule is in a non-single-stranded state, the method further comprises, before the dephosphorylation step, performing a melting process on the original template molecule to obtain a single-stranded DNA template molecule;
optionally, the melting process is a denaturation process;
optionally, in the primary melting step, and/or in the secondary melting step, and/or when the original template molecule is subjected to a denaturation treatment, the denaturation treatment is a heat denaturation treatment;
optionally, heat-denaturing, in particular heating the product to at least 80 ℃ for at least 1min, preferably heating the molecule to be treated to 80-98 ℃ for 1-30 min;
optionally, the dephosphorylation treatment step comprises adding a dephosphorylating enzyme to perform dephosphorylation treatment under the following conditions: at 20-30 ℃ for 5-30min, then carrying out denaturation treatment on dephosphorylation enzyme, wherein the reaction conditions are as follows: heating to at least 80 deg.C for at least 1 min;
optionally, in the first melting step, after the reverse strand of the first sample tag connected with the random sequence is dissociated from the linker sequence, streptavidin-coated magnetic beads are added, and the intermediate library molecules modified with the labeling molecules are collected by using a magnet;
optionally, the DNA polymerase is selected from the Klenow fragment of DNA polymerase I;
optionally, in the first sequencing linker ligation step and/or the second sequencing linker ligation step, the DNA ligase is selected from T4DNA ligase;
optionally, the second sequencing linker is selected from any one of a P5 terminal sequencing linker of Illumina sequencing platform, a P2 terminal sequencing linker of MGI sequencing platform;
optionally, the second sequencing adapter contains or does not contain a second molecular tag;
optionally, the second molecular tag is a random sequence;
optionally, the second molecular tag is 4-19nt in length;
optionally, in the step of primary double-chain synthesis, the reaction conditions are 65-75 ℃ in sequence and 1-3min in sequence; 30-37 deg.C, 5-30 min;
optionally, in the second sequencing linker connection step, the reaction conditions are 20-30 ℃ and 15-60 min;
optionally, the second primer contains or does not contain a second sample tag;
optionally, the second sample tag is a random sequence;
optionally, the second sample label is 4-10nt in length;
optionally, when the second primer does not contain a second sample tag, the second primer comprises an inner adaptor sequence, an outer adaptor sequence, the 5 'end of the inner adaptor sequence being linked to the 3' end of the outer adaptor sequence, the inner adaptor sequence being reverse complementary to the first sequencing adaptor;
optionally, when the second primer comprises a second sample tag, the second primer comprises an inner adaptor sequence and an outer adaptor sequence, wherein the 5 'end of the inner adaptor sequence is linked to the 3' end of the outer adaptor sequence, the inner adaptor sequence is reverse complementary to the first sequencing adaptor, and the second sample tag is located between the inner adaptor sequence and the outer adaptor sequence;
optionally, the third primer contains or does not contain a third sample tag;
optionally, the third sample tag is a random sequence;
optionally, the third sample label is 4-10nt in length;
optionally, when the third primer does not contain a second sample tag, the third primer comprises an inner adaptor sequence, an outer adaptor sequence, the 5 'end of the inner adaptor sequence being linked to the 3' end of the outer adaptor sequence, the inner adaptor sequence being reverse complementary to the second sequencing adaptor;
optionally, when the third primer comprises a third sample tag, the second primer comprises an inner adaptor sequence and an outer adaptor sequence, the 5 'end of the inner adaptor sequence is linked to the 3' end of the outer adaptor sequence, the inner adaptor sequence is reverse complementary mateable to the second sequencing adaptor, and the third sample tag is located between the inner adaptor sequence and the outer adaptor sequence.
Priority Applications (1)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| CN202011174182.6A CN112410331A (en) | 2020-10-28 | 2020-10-28 | Linker with molecular label and sample label and single-chain library building method thereof |
Applications Claiming Priority (1)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| CN202011174182.6A CN112410331A (en) | 2020-10-28 | 2020-10-28 | Linker with molecular label and sample label and single-chain library building method thereof |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| CN112410331A true CN112410331A (en) | 2021-02-26 |
Family
ID=74841507
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| CN202011174182.6A Pending CN112410331A (en) | 2020-10-28 | 2020-10-28 | Linker with molecular label and sample label and single-chain library building method thereof |
Country Status (1)
| Country | Link |
|---|---|
| CN (1) | CN112410331A (en) |
Cited By (4)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN112941073A (en) * | 2021-03-29 | 2021-06-11 | 武汉伯远生物科技有限公司 | Single-stranded DNA (deoxyribonucleic acid) joint and preparation and application thereof |
| CN116200478A (en) * | 2023-03-07 | 2023-06-02 | 安徽安龙基因科技有限公司 | Capturing library building method based on single-chain connection and application |
| US20230392201A1 (en) * | 2022-06-06 | 2023-12-07 | Element Biosciences, Inc. | Methods for assembling and reading nucleic acid sequences from mixed populations |
| CN117701679A (en) * | 2024-02-06 | 2024-03-15 | 中国医学科学院基础医学研究所 | Single-stranded DNA specific high-throughput sequencing method based on 5' connection |
Citations (12)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN105506125A (en) * | 2016-01-12 | 2016-04-20 | 上海美吉生物医药科技有限公司 | DNA sequencing method and next generation sequencing library |
| CN106867995A (en) * | 2017-03-01 | 2017-06-20 | 安徽安科生物工程(集团)股份有限公司 | CfDNA builds joint, primer sets, kit and the banking process in storehouse |
| CN107829146A (en) * | 2017-11-29 | 2018-03-23 | 广州赛哲生物科技股份有限公司 | Primer group for constructing 16SrRNA gene amplicon sequencing library and construction method |
| CN108004301A (en) * | 2017-12-15 | 2018-05-08 | 格诺思博生物科技南通有限公司 | Gene target region enrichment method and build storehouse kit |
| CN108148900A (en) * | 2018-01-24 | 2018-06-12 | 深圳因合生物科技有限公司 | Sequencing approach, kit and its application of sequencing mistake are reduced based on molecular label and the sequencing of two generations |
| CN108300716A (en) * | 2018-01-05 | 2018-07-20 | 武汉康测科技有限公司 | Joint component, its application and the method that targeting sequencing library structure is carried out based on asymmetric multiplex PCR |
| CN108949941A (en) * | 2018-06-25 | 2018-12-07 | 北京莲和医学检验所有限公司 | Low-frequency mutation detection method, kit and device |
| CN109706219A (en) * | 2018-12-20 | 2019-05-03 | 臻和(北京)科技有限公司 | Construct the method for splitting of the method for sequencing library, kit, upper machine method and sequencing data |
| CN110904512A (en) * | 2018-09-14 | 2020-03-24 | 广州华大基因医学检验所有限公司 | A high-throughput sequencing library construction method suitable for single-stranded DNA |
| CN111020711A (en) * | 2019-10-07 | 2020-04-17 | 深圳易倍科华生物科技有限公司 | Single-chain library construction method with molecular label, joint combination and kit |
| CN111321208A (en) * | 2020-02-14 | 2020-06-23 | 上海厦维生物技术有限公司 | Database building method based on high-throughput sequencing |
| CN111748613A (en) * | 2019-03-27 | 2020-10-09 | 华大数极生物科技(深圳)有限公司 | Design method and preparation method of double-label joint |
-
2020
- 2020-10-28 CN CN202011174182.6A patent/CN112410331A/en active Pending
Patent Citations (12)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN105506125A (en) * | 2016-01-12 | 2016-04-20 | 上海美吉生物医药科技有限公司 | DNA sequencing method and next generation sequencing library |
| CN106867995A (en) * | 2017-03-01 | 2017-06-20 | 安徽安科生物工程(集团)股份有限公司 | CfDNA builds joint, primer sets, kit and the banking process in storehouse |
| CN107829146A (en) * | 2017-11-29 | 2018-03-23 | 广州赛哲生物科技股份有限公司 | Primer group for constructing 16SrRNA gene amplicon sequencing library and construction method |
| CN108004301A (en) * | 2017-12-15 | 2018-05-08 | 格诺思博生物科技南通有限公司 | Gene target region enrichment method and build storehouse kit |
| CN108300716A (en) * | 2018-01-05 | 2018-07-20 | 武汉康测科技有限公司 | Joint component, its application and the method that targeting sequencing library structure is carried out based on asymmetric multiplex PCR |
| CN108148900A (en) * | 2018-01-24 | 2018-06-12 | 深圳因合生物科技有限公司 | Sequencing approach, kit and its application of sequencing mistake are reduced based on molecular label and the sequencing of two generations |
| CN108949941A (en) * | 2018-06-25 | 2018-12-07 | 北京莲和医学检验所有限公司 | Low-frequency mutation detection method, kit and device |
| CN110904512A (en) * | 2018-09-14 | 2020-03-24 | 广州华大基因医学检验所有限公司 | A high-throughput sequencing library construction method suitable for single-stranded DNA |
| CN109706219A (en) * | 2018-12-20 | 2019-05-03 | 臻和(北京)科技有限公司 | Construct the method for splitting of the method for sequencing library, kit, upper machine method and sequencing data |
| CN111748613A (en) * | 2019-03-27 | 2020-10-09 | 华大数极生物科技(深圳)有限公司 | Design method and preparation method of double-label joint |
| CN111020711A (en) * | 2019-10-07 | 2020-04-17 | 深圳易倍科华生物科技有限公司 | Single-chain library construction method with molecular label, joint combination and kit |
| CN111321208A (en) * | 2020-02-14 | 2020-06-23 | 上海厦维生物技术有限公司 | Database building method based on high-throughput sequencing |
Non-Patent Citations (1)
| Title |
|---|
| H. CHRISTINA FAN等: "Combinatorial labeling of single cells for gene expression cytometry", 《SCIENCE》 * |
Cited By (6)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN112941073A (en) * | 2021-03-29 | 2021-06-11 | 武汉伯远生物科技有限公司 | Single-stranded DNA (deoxyribonucleic acid) joint and preparation and application thereof |
| CN112941073B (en) * | 2021-03-29 | 2023-03-14 | 武汉伯远生物科技有限公司 | A kind of single-stranded DNA linker and its preparation and application |
| US20230392201A1 (en) * | 2022-06-06 | 2023-12-07 | Element Biosciences, Inc. | Methods for assembling and reading nucleic acid sequences from mixed populations |
| CN116200478A (en) * | 2023-03-07 | 2023-06-02 | 安徽安龙基因科技有限公司 | Capturing library building method based on single-chain connection and application |
| CN117701679A (en) * | 2024-02-06 | 2024-03-15 | 中国医学科学院基础医学研究所 | Single-stranded DNA specific high-throughput sequencing method based on 5' connection |
| US12534720B2 (en) | 2024-02-06 | 2026-01-27 | Institute Of Basic Medical Sciences Chinese Academy Of Medical Sciences | Methods for preparing 5′-end ligation-based ssDNA-specific sequencing libraries |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| CN110734908B (en) | Construction method of high-throughput sequencing library and kit for library construction | |
| EP3555305B1 (en) | Method for increasing throughput of single molecule sequencing by concatenating short dna fragments | |
| US10400279B2 (en) | Method for constructing a sequencing library based on a single-stranded DNA molecule and application thereof | |
| CN109593757B (en) | Probe and method for enriching target region by using same and applicable to high-throughput sequencing | |
| CN112410331A (en) | Linker with molecular label and sample label and single-chain library building method thereof | |
| CN110079592B (en) | High throughput sequencing-targeted capture of target regions for detection of genetic mutations and known, unknown gene fusion types | |
| TW201321518A (en) | Method of micro-scale nucleic acid library construction and application thereof | |
| CN109576346B (en) | Construction method and application of high-throughput sequencing library | |
| US20210214783A1 (en) | Method for constructing sequencing library, obtained sequencing library and sequencing method | |
| CN110904512A (en) | A high-throughput sequencing library construction method suitable for single-stranded DNA | |
| CN107604046B (en) | Next-generation sequencing method for bimolecular self-checking library preparation and hybrid capture for ultra-low frequency mutation detection of trace DNA | |
| CN114096678A (en) | Multiple nucleic acid co-labeling support, and preparation method and application thereof | |
| CN114729349A (en) | Method for detecting and sequencing barcode nucleic acid | |
| CN114555831B (en) | Method for preparing dual-index methylation sequence library | |
| CN115873922B (en) | A single-cell full-length transcript library construction and sequencing method | |
| CN112680796A (en) | Target gene enrichment and library construction method | |
| WO2022007863A1 (en) | Method for rapidly enriching target gene region | |
| CN112941147A (en) | High-fidelity target gene library building method and kit thereof | |
| WO2025000136A1 (en) | Method for preparing strand-specific library for rapid detection of various types of rnas, and high-throughput sequencing technique | |
| CN116287124A (en) | Single-stranded linker pre-ligation method, high-throughput sequencing library construction method and kit | |
| CN120344725A (en) | Methods for reducing free adapters in sequencing libraries | |
| CN114214734A (en) | A single-molecule target gene library building method and kit thereof | |
| CN118086457B (en) | Construction and application of DNA library | |
| CN106283198B (en) | Library construction method for single cell whole genome bisulfite sequencing | |
| CN114507903A (en) | Plasmid sequencing method |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| PB01 | Publication | ||
| PB01 | Publication | ||
| SE01 | Entry into force of request for substantive examination | ||
| SE01 | Entry into force of request for substantive examination |