CN111662903B - Logarithmic phase specific promoter and application thereof - Google Patents
Logarithmic phase specific promoter and application thereof Download PDFInfo
- Publication number
- CN111662903B CN111662903B CN201910228659.5A CN201910228659A CN111662903B CN 111662903 B CN111662903 B CN 111662903B CN 201910228659 A CN201910228659 A CN 201910228659A CN 111662903 B CN111662903 B CN 111662903B
- Authority
- CN
- China
- Prior art keywords
- lysine
- promoter
- lysine decarboxylase
- leu
- glu
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Active
Links
Classifications
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N9/00—Enzymes; Proenzymes; Compositions thereof; Processes for preparing, activating, inhibiting, separating or purifying enzymes
- C12N9/88—Lyases (4.)
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12P—FERMENTATION OR ENZYME-USING PROCESSES TO SYNTHESISE A DESIRED CHEMICAL COMPOUND OR COMPOSITION OR TO SEPARATE OPTICAL ISOMERS FROM A RACEMIC MIXTURE
- C12P13/00—Preparation of nitrogen-containing organic compounds
- C12P13/001—Amines; Imines
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Y—ENZYMES
- C12Y401/00—Carbon-carbon lyases (4.1)
- C12Y401/01—Carboxy-lyases (4.1.1)
- C12Y401/01018—Lysine decarboxylase (4.1.1.18)
Landscapes
- Chemical & Material Sciences (AREA)
- Organic Chemistry (AREA)
- Life Sciences & Earth Sciences (AREA)
- Engineering & Computer Science (AREA)
- Zoology (AREA)
- Health & Medical Sciences (AREA)
- Wood Science & Technology (AREA)
- Bioinformatics & Cheminformatics (AREA)
- Genetics & Genomics (AREA)
- General Health & Medical Sciences (AREA)
- General Engineering & Computer Science (AREA)
- Biochemistry (AREA)
- Microbiology (AREA)
- Biotechnology (AREA)
- Biomedical Technology (AREA)
- Molecular Biology (AREA)
- Medicinal Chemistry (AREA)
- Proteomics, Peptides & Aminoacids (AREA)
- Chemical Kinetics & Catalysis (AREA)
- General Chemical & Material Sciences (AREA)
- Micro-Organisms Or Cultivation Processes Thereof (AREA)
- Preparation Of Compounds By Using Micro-Organisms (AREA)
Abstract
本发明提供了对数期特异性启动子(序列如SEQ ID NO:1‑10所示)及其应用。本发明提供的对数期特异性诱导的启动子主要集中在菌体生长的对数期启动重组表达,这对于发酵生产赖氨酸脱羧酶具有很重要的意义,本发明保护的对数期启动子可以在菌株的生长度过停滞期后开始诱导赖氨酸脱羧酶基因的表达,因此避免了菌体在停滞期就开始表达基因给菌体适应发酵环境带来的压力,可以在同样的发酵时间内表达更多的赖氨酸脱羧酶。本发明为利用微生物表达赖氨酸脱羧酶并利用赖氨酸脱羧酶作为体外酶催化L‑赖氨酸生产1,5‑戊二胺提供有力的技术支持。The present invention provides a logarithmic phase-specific promoter (sequence shown in SEQ ID NO: 1‑10) and applications thereof. The logarithmic phase-specific induction promoter provided by the present invention is mainly focused on the logarithmic phase of bacterial growth to initiate recombinant expression, which is of great significance for the fermentation and production of lysine decarboxylase. The logarithmic phase of the protection of the present invention starts The seed can induce the expression of the lysine decarboxylase gene after the growth of the strain passes through the stagnation period, thus avoiding the pressure that the bacterium begins to express the gene during the stagnation period and brings the pressure on the bacterium to adapt to the fermentation environment, and can be used in the same fermentation Express more lysine decarboxylase over time. The invention provides powerful technical support for expressing lysine decarboxylase by microorganisms and using lysine decarboxylase as an in vitro enzyme to catalyze L-lysine to produce 1,5-pentanediamine.
Description
技术领域technical field
本发明属于生物技术领域,具体地说,涉及一种对数期特异性启动子及其应用。The invention belongs to the field of biotechnology, and in particular relates to a logarithmic phase-specific promoter and its application.
背景技术Background technique
赖氨酸脱羧酶(lysine decarboxylase,简称LDC,分类号为EC 4.1.1.18)广泛存在于微生物、动物和高等植物中,其可以将L-赖氨酸脱去一个羧基生成1,5-戊二胺(尸胺)和CO2。1,5-戊二胺的用途相当广泛,例如可以和二元酸进行聚合反应合成新型聚酰胺,在工业生产中具有很高的应用价值。目前,微生物法戊二胺生产主要采用微生物发酵生产和微生物体外酶催化生产戊二胺。Lysine decarboxylase (LDC for short, classification number EC 4.1.1.18) widely exists in microorganisms, animals and higher plants, which can remove a carboxyl group from L-lysine to generate 1,5-pentanedi Amines (cadaverine) and CO 2 . 1,5-Pentanediamine has a wide range of uses, for example, it can be polymerized with dibasic acids to synthesize new polyamides, which has high application value in industrial production. At present, the microbial production of pentamethylenediamine mainly adopts microbial fermentation production and microbial in vitro enzyme catalysis to produce pentamethylenediamine.
利用微生物体外酶催化生产1,5-戊二胺时,使用较多的是根据乳糖操纵子来构建赖氨酸脱羧酶基因表达盒,利用lacI功能缺失的乳糖操纵子,lac启动子可以组成型表达,不依赖于诱导剂的添加与否,实现异源基因伴随着菌体生长的大量表达。When catalyzing the production of 1,5-pentanediamine by microorganisms in vitro, the lysine decarboxylase gene expression cassette is often constructed based on the lactose operon, and the lactose operon with the lack of function of lacI is used, and the lac promoter can be constitutive Expression does not depend on the addition of inducers, and realizes the massive expression of heterologous genes accompanied by bacterial growth.
除了依赖于乳糖操纵子的lac启动子之外,还希望获得一些新的于菌体生长对数期就开始诱导下游基因转录的启动子,并且这些启动子的诱导不需要添加诱导剂。In addition to the lac promoter that depends on the lactose operon, it is also hoped to obtain some new promoters that can induce the transcription of downstream genes in the logarithmic phase of bacterial growth, and the induction of these promoters does not require the addition of inducers.
发明内容Contents of the invention
本发明的目的是提供对数期特异性启动子及其应用。The purpose of the present invention is to provide a logarithmic phase-specific promoter and its application.
本发明的另一目的是提供一种全细胞催化生产1,5-戊二胺的方法。Another object of the present invention is to provide a method for whole-cell catalytic production of 1,5-pentanediamine.
为了实现本发明目的,本发明人提供的对数期诱导的启动子能够在对数期利用微生物大量表达异源蛋白,为利用微生物表达对菌体生长产生毒性的蛋白提供技术支持。进一步地,为了实现全细胞催化生产1,5-戊二胺,将编码赖氨酸脱羧酶的基因构建至对数期特异性启动子之后,并转化至细菌菌株中,得到的重组菌株可以用于全细胞催化生产1,5-戊二胺。In order to achieve the purpose of the present invention, the logarithmic phase-induced promoter provided by the inventors can use microorganisms to express a large amount of heterologous proteins in logarithmic phases, and provide technical support for using microorganisms to express proteins that are toxic to bacterial growth. Further, in order to realize whole-cell catalytic production of 1,5-pentanediamine, the gene encoding lysine decarboxylase was constructed behind a logarithmic phase-specific promoter, and transformed into a bacterial strain, and the resulting recombinant strain could be used Catalyzes the production of 1,5-pentanediamine in whole cells.
第一方面,本发明提供的对数期特异性启动子,所述启动子的大小为37-42bp,至少包括-10区、RNA聚合酶бs特异性识别位点和-35区;In the first aspect, the logarithmic phase-specific promoter provided by the present invention has a size of 37-42 bp, and at least includes the -10 region, the RNA polymerase бs specific recognition site and the -35 region;
其中,将所述启动子序列的3’末端碱基对应于下游基因转录起始位点的位置记为-1位,所述-10区位于-12~-7位,碱基组成为5’-TATACT-3’;Wherein, the position of the 3' end base of the promoter sequence corresponding to the transcription start site of the downstream gene is recorded as position -1, the -10 region is located at positions -12 to -7, and the base composition is 5' -TATACT-3';
所述启动子序列的-13位碱基为C;The -13 base of the promoter sequence is C;
所述RNA聚合酶бs特异性识别位点位于-18~-14位,碱基组成为5’-TTGTT-3’;The RNA polymerase б s specific recognition site is located at positions -18 to -14, and the base composition is 5'-TTGTT-3';
所述-35区的碱基组成为5’-T(C或T)(C或T)(C或T)G(C或T或A)(T或C)-3’,但 5’-TCCCGCC-3’除外,且所述-35区和-10区之间间隔的碱基数为15-20bp。The base composition of the -35 region is 5'-T(C or T)(C or T)(C or T)G(C or T or A)(T or C)-3', but 5'- Except for TCCCGCC-3', and the number of bases between the -35 region and the -10 region is 15-20bp.
进一步地,所述对数期特异性启动子的序列为:Further, the sequence of the logarithmic phase-specific promoter is:
i)SEQ ID NO:1-10任一所示的核苷酸序列;或i) the nucleotide sequence shown in any one of SEQ ID NO: 1-10; or
ii)SEQ ID NO:1-10任一所示的核苷酸序列经取代、缺失和/或增加一个或多个核苷酸且以对数期特异性方式显示出启动子活性的核苷酸序列;或ii) The nucleotide sequence shown in any one of SEQ ID NO: 1-10 is substituted, deleted and/or increased by one or more nucleotides and exhibits promoter activity in a logarithmic phase-specific manner sequence; or
iii)在严格条件下与SEQ ID NO:1-10任一所示序列杂交且以对数期特异性方式显示出启动子活性的核苷酸序列,所述严格条件为在含0.1%SDS的0.1×SSPE或含 0.1%SDS的0.1×SSC溶液中,在65℃下杂交,并用该溶液洗膜;或iii) a nucleotide sequence that hybridizes to any of the sequences shown in SEQ ID NO: 1-10 and exhibits promoter activity in a logarithmic phase-specific manner under stringent conditions, and the stringent conditions are in the presence of 0.1% SDS Hybridize in 0.1×SSPE or 0.1×SSC solution containing 0.1% SDS at 65°C, and wash the membrane with this solution; or
iv)与i)、ii)或iii)的核苷酸序列具有90%以上同源性且以对数期特异性方式显示出启动子活性的核苷酸序列。iv) A nucleotide sequence that has more than 90% homology with the nucleotide sequence of i), ii) or iii) and exhibits promoter activity in a log-phase specific manner.
第二方面,本发明提供含有所述对数期启动子的生物材料,所述生物材料包括但不限于重组DNA、表达盒、转座子、质粒载体、噬菌体载体、病毒载体或工程菌。In a second aspect, the present invention provides biological materials containing the log-phase promoter, including but not limited to recombinant DNA, expression cassettes, transposons, plasmid vectors, phage vectors, viral vectors or engineering bacteria.
第三方面,本发明提供所述对数期启动子的以下任一应用:In a third aspect, the present invention provides any of the following applications of the logarithmic phase promoter:
1)作为原核生物的对数期特异性启动子的应用;1) Application as a logarithmic phase-specific promoter of prokaryotes;
2)在构建重组DNA、表达盒、转座子、质粒载体、噬菌体载体、病毒载体或工程菌中的应用。2) Application in the construction of recombinant DNA, expression cassettes, transposons, plasmid vectors, phage vectors, viral vectors or engineering bacteria.
所述原核生物为细菌,例如埃希氏菌属(Escherichia)、棒杆菌属(Corynebacterium)、短杆菌属(Brevibacterium)、链霉菌属(Streptomyces)、哈夫尼菌属(Hafnia)中的细菌;优选地,所述细菌选自大肠杆菌(E.coli)、枯草芽孢杆菌(B.subtilis)、天蓝色链霉菌(S.coelicolor)、蜂房哈夫尼菌(H.alvei)、谷氨酸棒状杆菌(C.glutamicum),或经过诱变或随机突变之后的菌株或基因工程菌。The prokaryote is a bacterium, such as bacteria in the genus Escherichia, Corynebacterium, Brevibacterium, Streptomyces, Hafnia; Preferably, the bacterium is selected from the group consisting of Escherichia coli (E.coli), Bacillus subtilis (B.subtilis), Streptomyces coelicolor (S.coelicolor), Hafnia alvei (H.alvei), Corynebacterium glutamicum Bacillus (C. glutamicum), or strains or genetically engineered bacteria after mutagenesis or random mutation.
进一步地,本发明提供所述对数期启动子作为大肠杆菌(E.coli)、枯草芽孢杆菌(B.subtilis)、天蓝色链霉菌(S.coelicolor)、蜂房哈夫尼菌(H.alvei)或谷氨酸棒状杆菌(C.glutamicum)在发酵生产赖氨酸脱羧酶中调控赖氨酸脱羧酶基因表达的对数期特异性启动子的用途。Further, the present invention provides the log phase promoter as Escherichia coli (E.coli), Bacillus subtilis (B.subtilis), Streptomyces coelicolor (S.coelicolor), Hafnia alvei (H.alvei ) or Corynebacterium glutamicum (C.glutamicum) in the production of lysine decarboxylase in the fermentative production of lysine decarboxylase gene expression of the use of logarithmic phase-specific promoter regulation.
第四方面,本发明提供一种重组DNA,由所述启动子与下游目的基因可操作连接而成。In the fourth aspect, the present invention provides a recombinant DNA, which is formed by operably linking the promoter with a downstream target gene.
本发明中,所述目的基因选自编码蛋白质的核酸、编码核酶的核酸、编码反义 RNA的核酸;优选地,所述蛋白质为酶、激素、抗体或生长因子;更优选地,所述酶选自氧化还原酶、转移酶、水解酶、裂合酶、异构酶、连接酶。进一步地,所述裂合酶为脱羧酶;所述脱羧酶为氨基酸脱羧酶,如赖氨酸脱羧酶、酪氨酸脱羧酶、精氨酸脱羧酶、鸟氨酸脱羧酶或谷氨酸脱羧酶。又进一步地,所述赖氨酸脱羧酶来源于大肠杆菌(Escherichia coli)、枯草芽孢杆菌(Bacillus subtilis)、嗜碱芽孢杆菌 (Bacillus halodurans)、天蓝色链霉菌(Streptomyces coelicolor)、蜂房哈夫尼菌(Hafnia alvei)、谷氨酸棒状杆菌(Corynebacterium glutamicum)或奥克西托克雷白杆菌 (Klebsiella oxytoca)。更进一步地,所述赖氨酸脱羧酶由cadA基因、ldcC基因、haldc 基因、cadA基因的片段、ldcC基因的片段或haldc基因的片段所编码。更进一步优选由cadA基因编码的赖氨酸脱羧酶。In the present invention, the target gene is selected from nucleic acids encoding proteins, nucleic acids encoding ribozymes, and nucleic acids encoding antisense RNAs; preferably, the proteins are enzymes, hormones, antibodies or growth factors; more preferably, the The enzyme is selected from the group consisting of oxidoreductases, transferases, hydrolases, lyases, isomerases, ligases. Further, the lyase is a decarboxylase; the decarboxylase is an amino acid decarboxylase, such as lysine decarboxylase, tyrosine decarboxylase, arginine decarboxylase, ornithine decarboxylase or glutamic acid decarboxylase enzyme. Still further, the lysine decarboxylase is derived from Escherichia coli, Bacillus subtilis, Bacillus halodurans, Streptomyces coelicolor, hive hafni Hafnia alvei, Corynebacterium glutamicum or Klebsiella oxytoca. Furthermore, the lysine decarboxylase is encoded by cadA gene, ldcC gene, haldc gene, a fragment of cadA gene, a fragment of ldcC gene or a fragment of haldc gene. Even more preferred is lysine decarboxylase encoded by the cadA gene.
第五方面,本发明提供一种表达载体,所述表达载体包含所述的重组DNA。In the fifth aspect, the present invention provides an expression vector, which contains the recombinant DNA.
在一些实施例中,所述表达载体携带含有赖氨酸脱羧酶基因的表达盒,且所述表达盒由所述对数期特异性启动子驱动赖氨酸脱羧酶基因的表达。In some embodiments, the expression vector carries an expression cassette containing a lysine decarboxylase gene, and the expression cassette is driven by the log-phase specific promoter to express the lysine decarboxylase gene.
在一些实施例中,所述表达载体还含有能够在宿主细胞中自主复制的骨架质粒;优选地,所述骨架质粒选自pUC18、pUC19、pBR322、pACYC、pET、pSC101和它们的衍生质粒。In some embodiments, the expression vector also contains a backbone plasmid capable of autonomous replication in the host cell; preferably, the backbone plasmid is selected from pUC18, pUC19, pBR322, pACYC, pET, pSC101 and their derivative plasmids.
第六方面,本发明提供一种转化体,所述转化体为携带所述表达载体的宿主菌。In the sixth aspect, the present invention provides a transformant, which is a host bacterium carrying the expression vector.
所述宿主菌选自埃希氏菌属(Escherichia)、棒杆菌属(Corynebacterium)、短杆菌属(Brevibacterium)、链霉菌属(Streptomyces)、哈夫尼菌属(Hafnia)中的菌种;优选地,所述宿主菌选自大肠杆菌(E.coli)、枯草芽孢杆菌(B.subtilis)、天蓝色链霉菌(S.coelicolor)、蜂房哈夫尼菌(H.alvei)、谷氨酸棒状杆菌(C.glutamicum),或经过诱变或随机突变之后的菌株或基因工程菌;更优选地,所述宿主菌为大肠杆菌(E.coli) 或蜂房哈夫尼菌(H.alvei)。The host bacterium is selected from bacterial species in Escherichia (Escherichia), Corynebacterium (Corynebacterium), Brevibacterium (Brevibacterium), Streptomyces (Streptomyces), Hafnia (Hafnia); preferably Preferably, the host bacterium is selected from the group consisting of Escherichia coli (E.coli), Bacillus subtilis (B.subtilis), Streptomyces coelicolor (S.coelicolor), Hafnia hive (H.alvei), rod-shaped glutamic acid Bacillus (C. glutamicum), or strains or genetically engineered bacteria after mutagenesis or random mutation; more preferably, the host bacteria is Escherichia coli (E.coli) or Hafnia alvei (H.alvei).
第七方面,本发明提供所述转化体在发酵生产氨基酸、多肽或蛋白质中的应用。In the seventh aspect, the present invention provides the use of the transformant in the fermentative production of amino acids, polypeptides or proteins.
第八方面,本发明提供一种产赖氨酸脱羧酶的工程菌,所述工程菌的出发菌株为大肠杆菌(E.coli)或蜂房哈夫尼菌(H.alvei),并携带含有赖氨酸脱羧酶基因表达盒的质粒,所述赖氨酸脱羧酶基因表达盒由所述对数期特异性启动子驱动赖氨酸脱羧酶基因的表达。In the eighth aspect, the present invention provides an engineering bacterium producing lysine decarboxylase, the starting strain of the engineering bacterium is Escherichia coli (E.coli) or haffney bacteria (H.alvei), and carries A plasmid for an amino acid decarboxylase gene expression cassette, the expression of the lysine decarboxylase gene is driven by the log phase specific promoter.
优选地,所述赖氨酸脱羧酶基因是来自大肠杆菌内源的cadA基因(SEQ ID NO:11),其编码大肠杆菌诱导型赖氨酸脱羧酶CadA(SEQ ID NO:12)。Preferably, the lysine decarboxylase gene is an endogenous cadA gene (SEQ ID NO: 11) from Escherichia coli, which encodes an Escherichia coli inducible lysine decarboxylase CadA (SEQ ID NO: 12).
优选地,所述对数期特异性启动子为SEQ ID NO:1、2、3、4、6任一所示的启动子。Preferably, the log-phase-specific promoter is the promoter shown in any one of SEQ ID NO: 1, 2, 3, 4, or 6.
进一步地,本发明提供的产赖氨酸脱羧酶的工程菌,所述工程菌含有所述对数期特异性启动子,或者携带含有赖氨酸脱羧酶基因表达盒的质粒,所述赖氨酸脱羧酶基因表达盒由所述对数期特异性启动子驱动赖氨酸脱羧酶基因的表达。Further, the engineered bacterium producing lysine decarboxylase provided by the present invention contains the logarithmic phase-specific promoter, or carries a plasmid containing a lysine decarboxylase gene expression cassette, and the lysine Acid decarboxylase gene expression cassette Expression of the lysine decarboxylase gene is driven by the log-phase specific promoter.
第九方面,本发明提供一种发酵生产赖氨酸脱羧酶的方法,包括在发酵培养基中对所述产赖氨酸脱羧酶的工程菌进行培养。In the ninth aspect, the present invention provides a method for fermentatively producing lysine decarboxylase, comprising culturing the engineering bacteria producing lysine decarboxylase in a fermentation medium.
第十方面,本发明提供一种发酵生产1,5-戊二胺的方法,包括利用上述第九方面得到的工程菌发酵生产赖氨酸脱羧酶,再利用获得的赖氨酸脱羧酶催化L-赖氨酸转化生成1,5-戊二胺。In the tenth aspect, the present invention provides a method for fermentatively producing 1,5-pentanediamine, comprising using the engineering bacteria obtained in the ninth aspect to ferment and produce lysine decarboxylase, and then using the obtained lysine decarboxylase to catalyze L - Conversion of lysine to 1,5-pentanediamine.
优选地,所述工程菌经发酵获得酶发酵液,将酶发酵液与L-赖氨酸、L-赖氨酸盐溶液或L-赖氨酸发酵液混合,催化L-赖氨酸转化生成1,5-戊二胺。Preferably, the engineered bacteria are fermented to obtain an enzyme fermentation liquid, and the enzyme fermentation liquid is mixed with L-lysine, L-lysine salt solution or L-lysine fermentation liquid to catalyze the conversion of L-lysine into 1,5-Pentanediamine.
优选地,所述L-赖氨酸发酵液选自L-赖氨酸发酵原液,L-赖氨酸发酵原液的浓缩液或稀释液,L-赖氨酸发酵原液去除菌体后的除菌L-赖氨酸发酵液和除菌L-赖氨酸发酵液的浓缩液或稀释液。所述L-赖氨酸发酵液可通过现有技术制备,例如在发酵培养基中培养产L-赖氨酸的工程菌获得L-赖氨酸发酵液,可参见CN104762336A,或者所述L-赖氨酸发酵液通过市售获得。Preferably, the L-lysine fermentation liquid is selected from the L-lysine fermentation stock solution, the concentrate or dilution of the L-lysine fermentation stock solution, and the sterilizing solution after the L-lysine fermentation stock solution removes bacteria. L-lysine fermented liquid and concentrate or diluted liquid of sterilized L-lysine fermented liquid. The L-lysine fermentation broth can be prepared by existing techniques, for example, cultivating L-lysine-producing engineering bacteria in a fermentation medium to obtain an L-lysine fermentation broth, see CN104762336A, or the L-lysine fermentation broth. Lysine fermentation broth was obtained commercially.
优选地,所述方法还包括向所述L-赖氨酸发酵液中添加辅酶,所述辅酶选自吡哆醛、磷酸吡哆醛、吡哆醇、吡哆胺中的一种或多种,更优选5′-磷酸吡哆醛。Preferably, the method also includes adding a coenzyme to the L-lysine fermentation broth, and the coenzyme is selected from one or more of pyridoxal, pyridoxal phosphate, pyridoxine, and pyridoxamine , more preferably pyridoxal 5'-phosphate.
优选地,催化L-赖氨酸转化生成1,5-戊二胺时,反应体系的温度为25℃-55℃,更优选28℃-40℃。Preferably, when catalyzing the conversion of L-lysine to 1,5-pentanediamine, the temperature of the reaction system is 25°C-55°C, more preferably 28°C-40°C.
借由上述技术方案,本发明至少具有下列优点及有益效果:By virtue of the above technical solutions, the present invention has at least the following advantages and beneficial effects:
本发明提供的对数期特异性诱导的启动子主要集中在菌体生长的对数期启动重组表达,这对于发酵生产赖氨酸脱羧酶具有很重要的意义,与组成型Plac启动子相比较,本发明保护的对数期启动子可以在菌株的生长度过停滞期后开始诱导赖氨酸脱羧酶基因的表达,避免了菌体在停滞期就开始表达基因给菌体适应发酵环境带来的压力,可以在同样的发酵时间内表达更多的赖氨酸脱羧酶。本发明为利用微生物表达赖氨酸脱羧酶并利用赖氨酸脱羧酶催化L-赖氨酸生产1,5-戊二胺提供有力的技术支持。The logarithmic phase-specific induction promoter provided by the present invention mainly focuses on the logarithmic phase of bacterial growth to initiate recombinant expression, which is of great significance for the fermentation and production of lysine decarboxylase, compared with the constitutive Plac promoter , the logarithmic phase promoter protected by the present invention can induce the expression of lysine decarboxylase gene after the growth of the bacterial strain crosses the stagnation period, avoiding that the bacterium begins to express the gene in the stagnation period and brings the bacterium to adapt to the fermentation environment. The pressure can express more lysine decarboxylase in the same fermentation time. The invention provides powerful technical support for expressing lysine decarboxylase by microorganism and catalyzing L-lysine to produce 1,5-pentanediamine by using lysine decarboxylase.
具体实施方式detailed description
以下实施例用于说明本发明,但不用来限制本发明的范围。若未特别指明,实施例均按照常规实验条件,如Sambrook等分子克隆实验手册(Sambrook J&Russell DW,Molecular Cloning:a Laboratory Manual,2001),或按照制造厂商说明书建议的条件。The following examples are used to illustrate the present invention, but are not intended to limit the scope of the present invention. Unless otherwise specified, the examples are all in accordance with conventional experimental conditions, such as Sambrook et al. Molecular Cloning Experiment Manual (Sambrook J & Russell DW, Molecular Cloning: a Laboratory Manual, 2001), or in accordance with the conditions suggested by the manufacturer's instructions.
以下实施例中述及的PCR扩增、质粒提取、酶切、酶切产物连接、转化的具体步骤、条件参数等均按所购相关酶和试剂的说明书建议的条件进行。其中PCR扩增所用的DNA聚合酶、酶切所用的限制性内切酶、酶切产物连接所用的连接酶、大肠杆菌 E.coli BL21感受态细胞均购自宝生物工程(大连)有限公司。质粒提取试剂盒、DNA 胶回收试剂盒、PCR纯化试剂盒均购自康宁生命科学(吴江)有限公司,商标Axygen,引物均购自赛默飞世尔科技(中国)有限公司,商标INVITROGEN。The specific steps of PCR amplification, plasmid extraction, enzyme digestion, enzyme digestion product ligation, transformation, and condition parameters mentioned in the following examples were all carried out according to the conditions suggested in the instructions of the purchased relevant enzymes and reagents. The DNA polymerase used for PCR amplification, the restriction endonuclease used for digestion, the ligase used for ligation of the digested product, and Escherichia coli E. coli BL21 competent cells were all purchased from Treasure Bioengineering (Dalian) Co., Ltd. Plasmid extraction kits, DNA gel recovery kits, and PCR purification kits were purchased from Corning Life Sciences (Wujiang) Co., Ltd. with the trademark Axygen, and primers were purchased from Thermo Fisher Scientific (China) Co., Ltd. with the trademark INVITROGEN.
以下实施例中述及的蜂房哈夫尼菌Am607(Hafnia alvei Am607)现已保藏于中国典型培养物保藏中心,地址:中国武汉,武汉大学,邮编430072,保藏编号CCTCC NO:M2018737,保藏日期2018年11月1日。The Hafnia alvei Am607 (Hafnia alvei Am607) mentioned in the following examples has been preserved in the China Center for Type Culture Collection, address: Wuhan, China, Wuhan University, zip code 430072, preservation number CCTCC NO: M2018737, preservation date 2018 November 1st.
以下实施例反应体系的L-赖氨酸-1,5-戊二胺转化率即1,5-戊二胺与初始L-赖氨酸的摩尔比的百分数,其中L-赖氨酸和1,5-戊二胺的检测使用核磁共振(BRUKERULTRASHIEDTM400PLUS,Beckman NMR)检测。The conversion rate of L-lysine-1,5-pentanediamine in the reaction system of the following examples is the percentage of the molar ratio of 1,5-pentanediamine to the initial L-lysine, wherein L-lysine and 1 , The detection of 5-pentanediamine was detected by nuclear magnetic resonance (BRUKERULTRASHIEDTM400PLUS, Beckman NMR).
以下实施例中述及的质粒转化方法如下:将连接产物加入至100μl的大肠杆菌E.coli BL21(DE3)感受态细胞中,冰浴20min后在42℃中热激90s。冰上孵育5min后加入1ml的LB。涂布至相对应的抗性平板上。The plasmid transformation method described in the following examples is as follows: the ligation product was added to 100 μl of E. coli E. coli BL21 (DE3) competent cells, and then heat-shocked at 42° C. for 90 seconds after 20 minutes of ice bathing. Add 1ml of LB after incubation on ice for 5min. Apply to the corresponding resistant plates.
以下实施例中所用的引物如下:The primers used in the following examples are as follows:
实施例1 对数期表达的启动子序列Example 1 Promoter sequence expressed in logarithmic phase
本发明设计并合成了10条启动子序列,所述启动子的长度为37-42bp,分别如下所示(5’-3’,启动子序列中3’端最后一个碱基对应下游基因转录起始位点的位置为-1 位),其中双下划线标注的为-10区(-12~-7),-10区碱基组成为5’-TATACT-3’;延伸的-10区为(-13位)保守碱基C;-18~-14位碱基组成为5’-TTGTT-3’,该区域为RNA 聚合酶бs的特异性识别所必须;单下划线标注为可能的-35区,为本发明发现的保守区域所在,调控启动子的启动时间,其碱基序列可以是 5’-T(C/T)(C/T)(C/T)G(C/T/A)(T/C)-3’,但其序列不可以是5’-TCCCGCC-3’,所述启动子-35区和-10区之间的间隔区碱基数可以为15-20bp。The present invention designs and synthesizes 10 promoter sequences, the length of the promoter is 37-42bp, respectively as follows (5'-3', the last base at the 3' end of the promoter sequence corresponds to the start of transcription of the downstream gene The position of the starting site is -1 position), in which the double underline marks the -10 region (-12~-7), and the base composition of the -10 region is 5'-TATACT-3'; the extended -10 region is ( -13 position) Conserved base C; -18~-14 base composition is 5'-TTGTT-3', this region is necessary for the specific recognition of RNA polymerase б s ; the single underline is marked as possible -35 Region, where the conserved region found in the present invention, regulates the start time of the promoter, and its base sequence can be 5'-T(C/T)(C/T)(C/T)G(C/T/A )(T/C)-3', but its sequence cannot be 5'-TCCCGCC-3', and the base number of the spacer between the -35 region and the -10 region of the promoter can be 15-20 bp.
1.CAGTCTTGTCAAATTTGTAAAATTGTTCGTATTG(SEQ ID NO:1);1. CAG TCTTGTC AAATTTGTAAAAATTGTTC GTATTG (SEQ ID NO: 1);
2.CCATTCCTGACAAATTTTTAATATTGTTCGTATTG(SEQ ID NO:2);2. CCAT TCCTGAC AAATTTTTAATATTGTTC GTATTG (SEQ ID NO: 2);
3.AATCTCGCCAAATTTCAATTTGTTCGTATTG(SEQ ID NO:3);3. AA TCTCGCC AAATTTCAATTTGTTC GTATTG (SEQ ID NO: 3);
4.TAAGTCTTGCCAAATCCTTATTGTTCGTATTG(SEQ ID NO:4);4. TAAG TCTTGCC AAATCCTTATTGTTC GTATTG (SEQ ID NO: 4);
5.TCCTGACAAATTTACAATATTGTTCGTATTG(SEQ ID NO:5);5. TCCTGAC AAATTTACAATATTGTTC GTATTG (SEQ ID NO: 5);
6.TCCTGCCAAATCCGTAAATTTTGTTCGTATTG(SEQ ID NO:6);6. TCCTGCC AAATCCGTAAATTTTGTTC GTATTG (SEQ ID NO: 6);
7.CACTTTCGTCAAATTTGTAATTATTGTTCGTATTG(SEQ ID NO:7);7. CAC TTTCGTC AAATTTGTAATTATTGTTC GTATTG (SEQ ID NO: 7);
8.TCTTGTTAAATTTGTAATTATTGTTCGTATTG(SEQ ID NO:8);8. TCTTGTT AAATTTGTAATTATTGTTC GTATTG (SEQ ID NO: 8);
9.TTTTGCTAAATTTGTAAATTATTGTTCGTATTG(SEQ ID NO:9);9. TTTTGCT AAATTTGTAAATTATTGTTC GTATTG (SEQ ID NO: 9);
10.CGGAGTCTTGACAATTGTAAATTATTGTTCGTATTG(SEQ ID NO:10)。10. CGGAG TCTTGAC AATTGTAAATTATTGTTC GTATTG (SEQ ID NO: 10).
实施例2 构建含有不同启动子的红色荧光蛋白表达质粒Example 2 Construction of red fluorescent protein expression plasmids containing different promoters
mCherry红色荧光蛋白的编码基因均为利用基因组装的方法合成。红色荧光蛋白mCherry的氨基酸序列为SEQ ID NO:14,经过密码子优化后得到其核苷酸序列 SEQ ID NO:15。密码子优化及其DNA组装的操作参考Hoover DM&Lubkowski J, Nucleic AcidsResearch 30:10,2002。The coding genes of mCherry red fluorescent protein are all synthesized by the method of gene assembly. The amino acid sequence of red fluorescent protein mCherry is SEQ ID NO:14, and its nucleotide sequence is SEQ ID NO:15 after codon optimization. Codon optimization and its DNA assembly operations refer to Hoover DM & Lubkowski J, Nucleic Acids Research 30:10, 2002.
利用引物mCherry-F(SEQ ID NO:16)和mCherry-R(SEQ ID NO:17)对密码子优化后的mCherry基因进行PCR扩增,并在其基因起始密码子上游引入核糖结合位点(RBS,SEQID NO:18);切胶回收目的大小的DNA片段后,进行加A反应,然后与pMD18-T(购自宝生物工程大连有限公司)载体进行连接,连接产物转化至 E.coli BL21感受态细胞中,在含有终浓度为100μg/ml的氨苄青霉素抗性的平板中进行筛选,37℃培养过夜(16h,下同),分别多挑取几个克隆摇菌后送样测序,选取测序序列正确且mCherry基因反向连入乳糖启动子plac后的克隆抽提质粒,得到含有mCherry基因的质粒T-mCherry。在测序过程中发现,mCherry基因测序验证正确且正向插入至plac后的克隆,在平板上可以表现为红色。由此,所克隆的mCherry基因可以在大肠杆菌中顺利表达且翻译产物均为有功能的蛋白。而mCherry 基因反向插入至plac后的克隆,虽然测序验证正确,克隆也没有红色出现,说明质粒T-mCherry可以用于后续验证启动子的启动时间。The codon-optimized mCherry gene was PCR amplified using primers mCherry-F (SEQ ID NO: 16) and mCherry-R (SEQ ID NO: 17), and a ribose binding site was introduced upstream of the gene start codon (RBS, SEQID NO: 18); after cutting the gel to recover the DNA fragment of the target size, carry out the A reaction, and then connect it with the pMD18-T (purchased from Treasure Bioengineering Dalian Co., Ltd.) vector, and transform the ligated product into E.coli BL21 competent cells were screened on a plate containing ampicillin resistance at a final concentration of 100 μg/ml, cultured overnight at 37°C (16h, the same below), and several clones were picked and shaken and sent for sequencing. Select the clone with the correct sequencing sequence and the reverse connection of the mCherry gene into the lactose promoter plac to extract the plasmid to obtain the plasmid T-mCherry containing the mCherry gene. During the sequencing process, it was found that the clones whose mCherry gene sequencing was verified to be correct and positively inserted into plac could appear red on the plate. Thus, the cloned mCherry gene can be successfully expressed in Escherichia coli and the translation products are all functional proteins. However, the clone after the mCherry gene was reversely inserted into plac, although the sequencing verification was correct, the clone did not appear red, indicating that the plasmid T-mCherry can be used for subsequent verification of the start time of the promoter.
以质粒T-mCherry为模板,首先利用引物T-mC-F1(SEQ ID NO:19)和引物 T-mC-R1(SEQ ID NO:20)进行质粒扩增,利用PCR纯化试剂盒对PCR产物进行纯化后,用限制性内切酶DpnI将模板消化掉后,转化至E.coli BL21细胞中,在含有终浓度为100μg/ml的氨苄青霉素的抗性平板上进行筛选,37℃培养过夜。随机挑取三个单克隆进行测序鉴定。选择测序正确的克隆抽提质粒,以该质粒为模板,利用引物T-mC-F2(SEQ ID NO:21)和引物T-mC-R2(SEQ ID NO:22)对质粒进行PCR 扩增,利用PCR纯化试剂盒对PCR产物进行纯化后,利用限制性内切酶DpnI将质粒模板消化掉后,转化至大肠杆菌E.coli BL21感受态细胞中,在含有终浓度为100 μg/ml的氨苄青霉素的抗性平板上进行筛选,37℃培养过夜。随机挑取三个单克隆进行测序鉴定。选择测序正确的克隆抽提质粒,得到在mCherry基因之前插入了RBS 序列及多克隆位点的质粒T-Psyn-mCherry。Using the plasmid T-mCherry as a template, first use primer T-mC-F1 (SEQ ID NO: 19) and primer T-mC-R1 (SEQ ID NO: 20) to carry out plasmid amplification, and use the PCR purification kit to purify the PCR product After purification, the template was digested with restriction endonuclease DpnI, transformed into E.coli BL21 cells, screened on a resistance plate containing ampicillin at a final concentration of 100 μg/ml, and cultured overnight at 37°C. Three single clones were randomly selected for sequencing identification. Select the correct clone to extract the plasmid, use the plasmid as a template, and use primer T-mC-F2 (SEQ ID NO:21) and primer T-mC-R2 (SEQ ID NO:22) to carry out PCR amplification of the plasmid, After the PCR product was purified using the PCR purification kit, the plasmid template was digested with the restriction endonuclease DpnI, and then transformed into E. coli E. Screened on penicillin resistance plates and cultured overnight at 37°C. Three single clones were randomly selected for sequencing identification. The clones with correct sequencing were selected to extract the plasmids to obtain the plasmid T-Psyn-mCherry with the RBS sequence and multiple cloning sites inserted before the mCherry gene.
使用本领域常用基因序列合成方法分别合成含有实施例1所列的10条启动子的双链DNA序列,并在序列5’和3’端分别连接有KpnI和ClaI的酶切位点,利用KpnI 和ClaI双酶切后,分别连接至KpnI和ClaI双酶切的质粒T-Psyn-mCherry中。得到含有不同启动子的质粒。分别将这10个质粒转化至大肠杆菌E.coli BL21感受态细胞中,得到分别含有这10个质粒的菌株,并分别命名为BL21-1~BL21-10。Double-stranded DNA sequences containing the 10 promoters listed in Example 1 were respectively synthesized using gene sequence synthesis methods commonly used in the art, and enzyme cleavage sites of KpnI and ClaI were respectively connected to the 5' and 3' ends of the sequence, and KpnI was used to After double digestion with KpnI and ClaI, they were ligated into the plasmid T-Psyn-mCherry which was digested with KpnI and ClaI, respectively. Plasmids containing different promoters were obtained. These 10 plasmids were respectively transformed into E. coli E. coli BL21 competent cells to obtain strains containing these 10 plasmids, which were named BL21-1-BL21-10 respectively.
实施例3 利用红色荧光蛋白验证启动子的启动时间Example 3 Using red fluorescent protein to verify the start time of the promoter
分别从10个质粒转化的平板上长出的克隆中各挑取4个于96孔深孔板中的600 μl含有终浓度为100μg/ml的氨苄青霉素的LB液体培养基中,于37℃、250rpm的摇床中培养过夜后,分别吸取20μl菌液转接至新鲜的600μl含有终浓度为100μg/ml 的氨苄青霉素的LB液体培养基中,于37℃、250rpm的摇床中培养2hrs,再转接 8μl菌液转接至酶标板中新鲜的200μl含有终浓度为100μg/ml的氨苄青霉素的LB 液体培养基中(设置四个重复),置于酶标仪中。设置确定不含有红色荧光蛋白的菌株的菌液为阴性对照,阴性对照菌体正常生长,但无荧光检测到。设置温度为37℃,设置工作流程:首先摇菌15s,测定菌体的OD600nm的值,然后测定菌体在575nm 的激发光、610nm的发射光条件下的荧光值,最后摇菌600s,该流程循环,每隔15min 测定一次,全长时间为24hrs。测定结束后,分别绘制菌体的生长曲线和单位OD的荧光曲线(菌体某个时间点的荧光值于该时间点测定的OD的比值)曲线。根据两条曲线的对应关系获得该菌体荧光显著增强的时间点为菌体生长的哪个阶段。当菌体生长的对数期后荧光值就开始显著提高的克隆中所含有的启动子即确定为对数期特异性诱导的启动子。Pick 4 clones from the 10 plasmid-transformed plates and place them in 600 μl LB liquid medium containing ampicillin at a final concentration of 100 μg/ml in a 96-well deep-well plate, and store them at 37°C, After culturing overnight in a shaker at 250 rpm, transfer 20 μl of the bacterial solution to fresh 600 μl LB liquid medium containing ampicillin at a final concentration of 100 μg/ml, and culture in a shaker at 37°C and 250 rpm for 2 hrs, and then Transfer 8 μl of the bacteria solution to the fresh 200 μl LB liquid medium containing ampicillin at a final concentration of 100 μg/ml in the microplate plate (set up four replicates), and place it in a microplate reader. The bacteria solution of the bacterial strain that does not contain red fluorescent protein is set as a negative control, and the negative control bacteria grow normally, but no fluorescence is detected. Set the temperature to 37°C and set the workflow: first, shake the bacteria for 15s, measure the OD 600nm value of the bacteria, then measure the fluorescence value of the bacteria under the conditions of 575nm excitation light and 610nm emission light, and finally shake the bacteria for 600s, the The process cycle is measured every 15 minutes, and the total time is 24hrs. After the measurement, the growth curve of the bacteria and the fluorescence curve of the unit OD (the ratio of the fluorescence value of the bacteria at a certain time point to the OD measured at this time point) curves were drawn respectively. According to the corresponding relationship between the two curves, it is obtained which stage of the growth of the bacteria is the time point when the fluorescence of the bacteria is significantly enhanced. The promoter contained in the clone whose fluorescence value began to increase significantly after the logarithmic phase of cell growth was determined to be a logarithmic phase-specifically induced promoter.
从表1可以看出,含有本发明启动子的重组菌株,培养时间为360~480min时,菌体OD增长迅速,从约0.2增加至约0.4,伴随着菌体的荧光值/OD也出现了显著增加,可以从约80单位增加至约120单位;随着菌体的培养时间继续延长,菌体的 OD和荧光值/OD都无显著增加,说明这些启动子都是在菌体生长的对数期开始启动下游基因转录的。As can be seen from Table 1, when the recombinant bacterial strain containing the promoter of the present invention is cultivated for 360-480 minutes, the OD of the thalline grows rapidly, from about 0.2 to about 0.4, and the fluorescence value/OD of the thalline also increases. significantly increased, from about 80 units to about 120 units; as the culture time of the bacteria continued to prolong, the OD and fluorescence value/OD of the bacteria did not increase significantly, indicating that these promoters are all on the right side of the growth of the bacteria. Several phases begin to initiate transcription of downstream genes.
表1 利用酶标仪检测的各菌株不同培养时间的OD值和荧光值/ODTable 1 OD value and fluorescence value/OD of each strain detected by microplate reader at different culture time
实施例4 对数期启动子用于表达赖氨酸脱羧酶进行体外酶催化生产1,5-戊二胺Example 4 The logarithmic phase promoter is used to express lysine decarboxylase for in vitro enzymatic production of 1,5-pentanediamine
1、构建含有对数期启动子的诱导表达赖氨酸脱羧酶的质粒及重组菌株1. Construction of plasmids and recombinant strains containing logarithmic promoters for inducible expression of lysine decarboxylase
以大肠杆菌MG1655 K12的基因组为模板,利用引物cadA-F(SEQ ID NO:23)和cadA-R(SEQ ID NO:24)进行第一轮扩增,对目的片段切胶回收后,作为模板,再利用引物cadA-F2(SEQ ID NO:25)和cadA-R(SEQ ID NO:24)进行第二轮扩增,切胶回收后得到5’端带有HindIII酶切位点以及核糖体结合位点的cadA基因(SEQ ID NO: 13);抽提实施例2中得到的含有10种启动子的质粒,利用HindIII和XbaI双酶切,与同样双酶切的cadA基因(SEQID NO:13)进行连接,得到T-1-cadA,T-2-cadA,T-3-cadA, T-4-cadA,T-5-cadA,T-6-cadA,T-7-cadA,T-8-cadA,T-9-cadA,T-10-cadA这10个 cadA表达质粒,将这10个质粒分别转化至蜂房哈夫尼菌株Am607中,得到10个重组菌株,HA-1-cadA,HA-2-cadA,HA-3-cadA,HA-4-cadA,HA-5-cadA,HA-5-cadA, HA-6-cadA,HA-7-cadA,HA-8-cadA,HA-9-cadA,HA-10-cadA。Using the genome of Escherichia coli MG1655 K12 as a template, the primers cadA-F (SEQ ID NO:23) and cadA-R (SEQ ID NO:24) were used for the first round of amplification, and the target fragment was gel-cut and recovered as a template , and then use primers cadA-F2 (SEQ ID NO: 25) and cadA-R (SEQ ID NO: 24) for the second round of amplification, after gel cutting and recovery, the 5' end with HindIII restriction site and ribosome The cadA gene (SEQ ID NO: 13) of the binding site; the plasmid containing 10 kinds of promoters obtained in the extraction example 2 was digested with HindIII and XbaI, and the cadA gene (SEQ ID NO: 13) Connect to get T-1-cadA, T-2-cadA, T-3-cadA, T-4-cadA, T-5-cadA, T-6-cadA, T-7-cadA, T- 8-cadA, T-9-cadA, T-10-cadA these 10 cadA expression plasmids, these 10 plasmids were respectively transformed into hive Hafney strain Am607, and 10 recombinant strains were obtained, HA-1-cadA, HA-2-cadA, HA-3-cadA, HA-4-cadA, HA-5-cadA, HA-5-cadA, HA-6-cadA, HA-7-cadA, HA-8-cadA, HA- 9-cadA, HA-10-cadA.
已知在没有LacI阻遏蛋白表达的情况下,plac启动子可以为组成型启动子,其下游基因是随着菌体生长而启动表达的,为了比较plac和本发明对数期诱导的启动子在表达异源蛋白方面的优劣。同时,以pUC18质粒为模板,利用引物plac-F(SEQ ID NO:26)和plac-R(SEQ ID NO:27)扩增得到plac组成型启动子(SEQ ID NO:28), KpnI、ClaI双酶切后,与同样双酶切的质粒T-1-cadA连接,得到质粒T-plac-cadA。同样将其转化至蜂房哈夫尼菌株Am607中,得到HA-plac-cadA重组菌株。It is known that in the absence of LacI repressor protein expression, the plac promoter can be a constitutive promoter, and its downstream genes are expressed along with the growth of the bacterium. In order to compare plac and the logarithmic phase-induced promoter of the present invention in Pros and cons in expressing heterologous proteins. At the same time, using the pUC18 plasmid as a template, primers plac-F (SEQ ID NO: 26) and plac-R (SEQ ID NO: 27) were used to amplify the plac constitutive promoter (SEQ ID NO: 28), KpnI, ClaI After double digestion, it was ligated with the same double digestion plasmid T-1-cadA to obtain plasmid T-plac-cadA. It was also transformed into Haffney strain Am607 to obtain HA-plac-cadA recombinant strain.
2、发酵培养重组菌株2. Fermentation and culture of recombinant strains
上述重组菌株分别挑取三个单克隆至5mL的LB液体培养基中,所述LB液体培养基含有胰蛋白胨(tryptone)1%(w/v)、酵母提取物(yeast extract)0.5%(w/v)、NaCl 1%(w/v)、氨苄青霉素100μg/mL,于37℃培养24h,获得酶发酵液,分别测定各菌株酶发酵液的OD560nm。Three single clones of the above-mentioned recombinant strains were respectively picked into 5 mL of LB liquid medium, and the LB liquid medium contained tryptone (tryptone) 1% (w/v), yeast extract (yeast extract) 0.5% (w /v), NaCl 1% (w/v), and ampicillin 100 μg/mL, cultured at 37° C. for 24 hours to obtain enzyme fermentation liquid, and measure the OD 560nm of each strain’s enzyme fermentation liquid.
3、催化L-赖氨酸转化生成1,5-戊二胺3. Catalyze the conversion of L-lysine into 1,5-pentanediamine
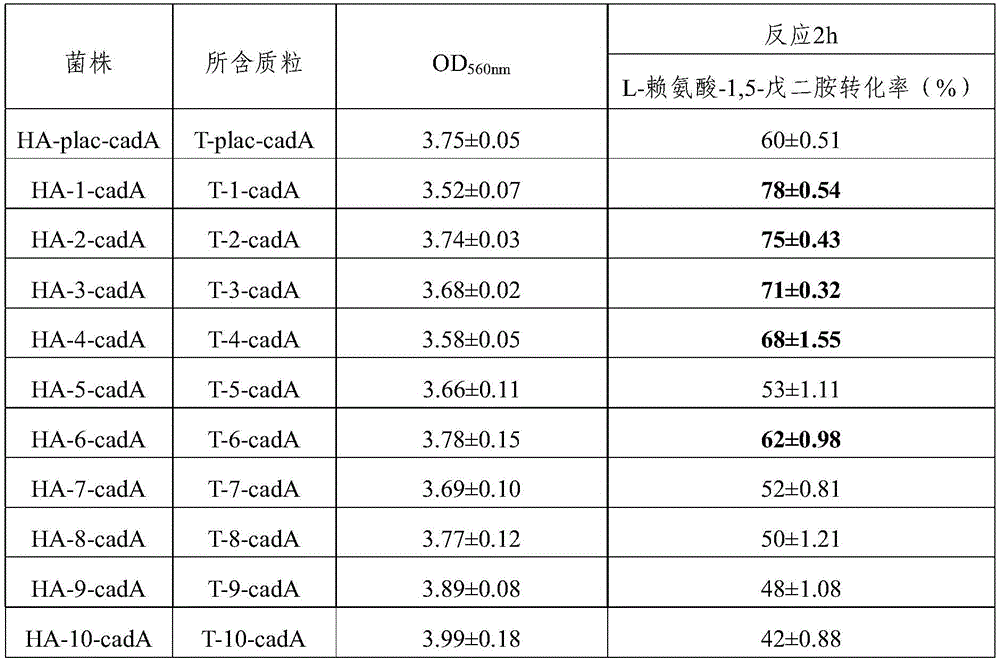
取上述步骤获得的每个重组菌株的酶发酵液500μl,加入500μl L-赖氨酸盐酸盐水溶液(L-赖氨酸浓度为400g/L),终浓度为0.05mM的5′-磷酸吡哆醛,pH自然,反应温度37℃,转速250rpm,反应2h,反应结束后对反应液进行离心(12000rpm,3 min),取500μl的反应液进行检测并计算各反应体系的L-赖氨酸-1,5-戊二胺转化率 (表2)。Get 500 μl of enzyme fermentation broth of each recombinant strain obtained in the above steps, add 500 μl of L-lysine hydrochloride aqueous solution (L-lysine concentration is 400g/L), and the final concentration is 0.05mM 5'-pyridine phosphate Pyridoxal, pH natural, reaction temperature 37°C, rotation speed 250rpm, react for 2h, after the reaction, centrifuge the reaction solution (12000rpm, 3 min), take 500μl of the reaction solution for detection and calculate the L-lysine of each reaction system - 1,5-pentanediamine conversion (Table 2).
表2 检测各反应体系中剩余的L-赖氨酸和转化生成的1,5-戊二胺的水平Table 2 Detection of the levels of remaining L-lysine and converted 1,5-pentanediamine in each reaction system
从表2可以看出,通过比较本发明中的10个对数期启动子与组成型的plac启动子诱导表达赖氨酸脱羧酶,结果表明本发明筛选到的对数期启动子1-10均能够催化L- 赖氨酸转化为1,5-戊二胺,而且对数期启动子1、2、3、4、6诱导表达赖氨酸脱羧酶获得的转化率比组成型的plac启动子更有优势,转化率分别为78%、75%、71%、68%和62%。As can be seen from Table 2, by comparing 10 logarithmic phase promoters in the present invention with constitutive plac promoters to induce expression of lysine decarboxylase, the results show that the present invention screened logarithmic phase promoters 1-10 Both can catalyze the conversion of L-lysine into 1,5-pentanediamine, and the conversion rate of lysine decarboxylase induced by logarithmic promoters 1, 2, 3, 4, and 6 is higher than that of constitutive plac Sons are more dominant, with conversion rates of 78%, 75%, 71%, 68% and 62%, respectively.
实施例5 对数期启动子用于表达赖氨酸脱羧酶进行体外酶催化生产1,5-戊二胺Example 5 The logarithmic phase promoter is used to express lysine decarboxylase for in vitro enzymatic production of 1,5-pentanediamine
构建含有对数期启动子的诱导表达赖氨酸脱羧酶的质粒及重组菌株的方法、发酵培养重组菌株的方法与实施例4相同,区别在于,催化L-赖氨酸转化生成1,5-戊二胺:将获得的重组菌株HA-1-cadA,HA-2-cadA,HA-3-cadA,HA-4-cadA,HA-6-cadA 和HA-plac-cadA的酶发酵液与L-赖氨酸发酵液混合,转化生成1,5-戊二胺,具体包括如下步骤:The method of constructing a plasmid containing a logarithmic phase promoter to induce expression of lysine decarboxylase and the recombinant strain, and the method of fermenting and cultivating the recombinant strain are the same as in Example 4, the difference is that the conversion of L-lysine is catalyzed to generate 1,5- Pentylenediamine: The obtained recombinant strains HA-1-cadA, HA-2-cadA, HA-3-cadA, HA-4-cadA, HA-6-cadA and HA-plac-cadA were mixed with L -Lysine fermentation broth is mixed and converted into 1,5-pentanediamine, which specifically includes the following steps:
取600μl赖氨酸发酵原液(购自凯赛(金乡)生物材料有限公司),赖氨酸含量为11%。向此赖氨酸发酵原液中分别加入上述酶发酵液100μl和终浓度为0.05mM的 5`-磷酸吡哆醛,pH自然,反应温度38℃,转速250rpm,反应2h,反应结束后对反应液进行离心(12000rpm,3min),取500μl的反应液进行检测并计算各反应体系的L- 赖氨酸-1,5-戊二胺转化率(表3)。Take 600 μl of lysine fermentation stock solution (purchased from Kaiser (Jinxiang) Biomaterials Co., Ltd.), with a lysine content of 11%. Add 100 μl of the above-mentioned enzyme fermentation broth and 5`-pyridoxal phosphate with a final concentration of 0.05 mM to this lysine fermentation stock solution, the pH is natural, the reaction temperature is 38 ° C, the rotation speed is 250 rpm, and the reaction is 2 hours. After the reaction, the reaction solution is After centrifugation (12000 rpm, 3 min), 500 μl of the reaction solution was taken for detection and the conversion rate of L-lysine-1,5-pentanediamine in each reaction system was calculated (Table 3).
表3 检测各反应体系中剩余的L-赖氨酸和转化生成的1,5-戊二胺的水平Table 3 Detection of the levels of remaining L-lysine and converted 1,5-pentanediamine in each reaction system
从表3可以看出,本发明筛选的对数期启动子1、2、3、4、6诱导表达的赖氨酸脱羧酶在对赖氨酸发酵原液进行催化转化时,同样获得了较高的L-赖氨酸-1,5-戊二胺转化率,本发明筛选的对数期启动子促进简化了L-赖氨酸催化转化1,5-戊二胺的工艺。As can be seen from Table 3, the lysine decarboxylase induced and expressed by the logarithmic phase promoters 1, 2, 3, 4, and 6 screened by the present invention also obtained higher The L-lysine-1,5-pentanediamine conversion rate, the logarithmic phase promoter screened by the present invention promotes and simplifies the process of catalytic conversion of L-lysine to 1,5-pentanediamine.
虽然,上文中已经用一般性说明及具体实施方式对本发明作了详尽的描述,但在本发明基础上,可以对之做一些修改或改进,这对本领域技术人员而言是显而易见的。因此,在不偏离本发明精神的基础上所做的这些修改或改进,均属于本发明要求保护的范围。Although the present invention has been described in detail with general descriptions and specific implementations above, it is obvious to those skilled in the art that some modifications or improvements can be made on the basis of the present invention. Therefore, the modifications or improvements made on the basis of not departing from the spirit of the present invention all belong to the protection scope of the present invention.
序列表sequence listing
<110> 上海凯赛生物技术研发中心有限公司 CIBT美国公司<110> Shanghai Cathay Biotechnology R&D Center Co., Ltd. CIBT America Inc.
<120> 对数期特异性启动子及其应用<120> Log-phase specific promoters and applications thereof
<130> KHP181115191.8<130> KHP181115191.8
<160> 28<160> 28
<170> SIPOSequenceListing 1.0<170> SIPOSequenceListing 1.0
<210> 1<210> 1
<211> 40<211> 40
<212> DNA<212> DNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence (Artificial Sequence)
<400> 1<400> 1
cagtcttgtc aaatttgtaa aattgttcta tactgtattg 40cagtcttgtc aaatttgtaa aattgttcta tactgtattg 40
<210> 2<210> 2
<211> 41<211> 41
<212> DNA<212> DNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence (Artificial Sequence)
<400> 2<400> 2
ccattcctga caaattttta atattgttct atactgtatt g 41ccattcctga caaattttta atattgttct atactgtatt g 41
<210> 3<210> 3
<211> 37<211> 37
<212> DNA<212> DNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence (Artificial Sequence)
<400> 3<400> 3
aatctcgcca aatttcaatt tgttctatac tgtattg 37aatctcgcca aatttcaatt tgttctatac tgtattg 37
<210> 4<210> 4
<211> 38<211> 38
<212> DNA<212> DNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence (Artificial Sequence)
<400> 4<400> 4
taagtcttgc caaatcctta ttgttctata ctgtattg 38taagtcttgc caaatcctta ttgttctata ctgtattg 38
<210> 5<210> 5
<211> 37<211> 37
<212> DNA<212> DNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence (Artificial Sequence)
<400> 5<400> 5
tcctgacaaa tttacaatat tgttctatac tgtattg 37tcctgacaaa tttacaatat tgttctatac tgtattg 37
<210> 6<210> 6
<211> 38<211> 38
<212> DNA<212> DNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence (Artificial Sequence)
<400> 6<400> 6
tcctgccaaa tccgtaaatt ttgttctata ctgtattg 38tcctgccaaa tccgtaaatt ttgttctata ctgtattg 38
<210> 7<210> 7
<211> 41<211> 41
<212> DNA<212> DNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence (Artificial Sequence)
<400> 7<400> 7
cactttcgtc aaatttgtaa ttattgttct atactgtatt g 41cactttcgtc aaatttgtaa ttattgttct atactgtatt g 41
<210> 8<210> 8
<211> 38<211> 38
<212> DNA<212> DNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence (Artificial Sequence)
<400> 8<400> 8
tcttgttaaa tttgtaatta ttgttctata ctgtattg 38tcttgttaaa tttgtaatta ttgttctata ctgtattg 38
<210> 9<210> 9
<211> 39<211> 39
<212> DNA<212> DNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence (Artificial Sequence)
<400> 9<400> 9
ttttgctaaa tttgtaaatt attgttctat actgtattg 39ttttgctaaa tttgtaaatt attgttctat actgtattg 39
<210> 10<210> 10
<211> 42<211> 42
<212> DNA<212> DNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence (Artificial Sequence)
<400> 10<400> 10
cggagtcttg acaattgtaa attattgttc tatactgtat tg 42cggagtcttg acaattgtaa attattgttc tatactgtat tg 42
<210> 11<210> 11
<211> 2148<211> 2148
<212> DNA<212> DNA
<213> 大肠杆菌(Escherichia coli)<213> Escherichia coli
<400> 11<400> 11
atgaacgtta ttgcaatatt gaatcacatg ggggtttatt ttaaagaaga acccatccgt 60atgaacgtta ttgcaatatt gaatcacatg ggggtttatt ttaaagaaga acccatccgt 60
gaacttcatc gcgcgcttga acgtctgaac ttccagattg tttacccgaa cgaccgtgac 120gaacttcatc gcgcgcttga acgtctgaac ttccagattg tttacccgaa cgaccgtgac 120
gacttattaa aactgatcga aaacaatgcg cgtctgtgcg gcgttatttt tgactgggat 180gacttattaa aactgatcga aaacaatgcg cgtctgtgcg gcgttatttt tgactgggat 180
aaatataatc tcgagctgtg cgaagaaatt agcaaaatga acgagaacct gccgttgtac 240aaatataatc tcgagctgtg cgaagaaatt agcaaaatga acgagaacct gccgttgtac 240
gcgttcgcta atacgtattc cactctcgat gtaagcctga atgacctgcg tttacagatt 300gcgttcgcta atacgtattc cactctcgat gtaagcctga atgacctgcg tttacagatt 300
agcttctttg aatatgcgct gggtgctgct gaagatattg ctaataagat caagcagacc 360agcttctttg aatatgcgct gggtgctgct gaagatattg ctaataagat caagcagacc 360
actgacgaat atatcaacac tattctgcct ccgctgacta aagcactgtt taaatatgtt 420actgacgaat atatcaacac tattctgcct ccgctgacta aagcactgtt taaatatgtt 420
cgtgaaggta aatatacttt ctgtactcct ggtcacatgg gcggtactgc attccagaaa 480cgtgaaggta aatatacttt ctgtactcct ggtcacatgg gcggtactgc attccagaaa 480
agcccggtag gtagcctgtt ctatgatttc tttggtccga ataccatgaa atctgatatt 540agcccggtag gtagcctgtt ctatgatttc tttggtccga ataccatgaa atctgatatt 540
tccatttcag tatctgaact gggttctctg ctggatcaca gtggtccaca caaagaagca 600tccatttcag tatctgaact gggttctctg ctggatcaca gtggtccaca caaagaagca 600
gaacagtata tcgctcgcgt ctttaacgca gaccgcagct acatggtgac caacggtact 660gaacagtata tcgctcgcgt ctttaacgca gaccgcagct acatggtgac caacggtact 660
tccactgcga acaaaattgt tggtatgtac tctgctccag caggcagcac cattctgatt 720tccactgcga acaaaattgt tggtatgtac tctgctccag caggcagcac cattctgatt 720
gaccgtaact gccacaaatc gctgacccac ctgatgatga tgagcgatgt tacgccaatc 780gaccgtaact gccacaaatc gctgacccac ctgatgatga tgagcgatgt tacgccaatc 780
tatttccgcc cgacccgtaa cgcttacggt attcttggtg gtatcccaca gagtgaattc 840tatttccgcc cgacccgtaa cgcttacggt attcttggtg gtatcccaca gagtgaattc 840
cagcacgcta ccattgctaa gcgcgtgaaa gaaacaccaa acgcaacctg gccggtacat 900cagcacgcta ccattgctaa gcgcgtgaaa gaaacaccaa acgcaacctg gccggtacat 900
gctgtaatta ccaactctac ctatgatggt ctgctgtaca acaccgactt catcaagaaa 960gctgtaatta ccaactctac ctatgatggt ctgctgtaca acaccgactt catcaagaaa 960
acactggatg tgaaatccat ccactttgac tccgcgtggg tgccttacac caacttctca 1020acactggatg tgaaatccat ccactttgac tccgcgtggg tgccttacac caacttctca 1020
ccgatttacg aaggtaaatg cggtatgagc ggtggccgtg tagaagggaa agtgatttac 1080ccgattacg aaggtaaatg cggtatgagc ggtggccgtg tagaagggaa agtgattac 1080
gaaacccagt ccactcacaa actgctggcg gcgttctctc aggcttccat gatccacgtt 1140gaaacccagt ccactcacaa actgctggcg gcgttctctc aggcttccat gatccacgtt 1140
aaaggtgacg taaacgaaga aacctttaac gaagcctaca tgatgcacac caccacttct 1200aaaggtgacg taaacgaaga aacctttaac gaagcctaca tgatgcacac caccacttct 1200
ccgcactacg gtatcgtggc gtccactgaa accgctgcgg cgatgatgaa aggcaatgca 1260ccgcactacg gtatcgtggc gtccactgaa accgctgcgg cgatgatgaa aggcaatgca 1260
ggtaagcgtc tgatcaacgg ttctattgaa cgtgcgatca aattccgtaa agagatcaaa 1320ggtaagcgtc tgatcaacgg ttctattgaa cgtgcgatca aattccgtaa agagatcaaa 1320
cgtctgagaa cggaatctga tggctggttc tttgatgtat ggcagccgga tcatatcgat 1380cgtctgagaa cggaatctga tggctggttc tttgatgtat ggcagccgga tcatatcgat 1380
acgactgaat gctggccgct gcgttctgac agcacctggc acggcttcaa aaacatcgat 1440acgactgaat gctggccgct gcgttctgac agcacctggc acggcttcaa aaacatcgat 1440
aacgagcaca tgtatcttga cccgatcaaa gtcaccctgc tgactccggg gatggaaaaa 1500aacgagcaca tgtatcttga cccgatcaaa gtcaccctgc tgactccggg gatggaaaaa 1500
gacggcacca tgagcgactt tggtattccg gccagcatcg tggcgaaata cctcgacgaa 1560gacggcacca tgagcgactt tggtattccg gccagcatcg tggcgaaata cctcgacgaa 1560
catggcatcg ttgttgagaa aaccggtccg tataacctgc tgttcctgtt cagcatcggt 1620catggcatcg ttgttgagaa aaccggtccg tataacctgc tgttcctgtt cagcatcggt 1620
atcgataaga ccaaagcact gagcctgctg cgtgctctga ctgactttaa acgtgcgttc 1680atcgataaga ccaaagcact gagcctgctg cgtgctctga ctgactttaa acgtgcgttc 1680
gacctgaacc tgcgtgtgaa aaacatgctg ccgtctctgt atcgtgaaga tcctgaattc 1740gacctgaacc tgcgtgtgaa aaacatgctg ccgtctctgt atcgtgaaga tcctgaattc 1740
tatgaaaaca tgcgtattca ggaactggct cagaatatcc acaaactgat tgttcaccac 1800tatgaaaaca tgcgtattca ggaactggct cagaatatcc acaaactgat tgttcaccac 1800
aatctgccgg atctgatgta tcgcgcattt gaagtgctgc cgacgatggt aatgactccg 1860aatctgccgg atctgatgta tcgcgcattt gaagtgctgc cgacgatggt aatgactccg 1860
tatgctgcat tccagaaaga gctgcacggt atgaccgaag aagtttacct cgacgaaatg 1920tatgctgcat tccagaaaga gctgcacggt atgaccgaag aagtttacct cgacgaaatg 1920
gtaggtcgta ttaacgccaa tatgatcctt ccgtacccgc cgggagttcc tctggtaatg 1980gtaggtcgta ttaacgccaa tatgatccctt ccgtacccgc cgggagttcc tctggtaatg 1980
ccgggtgaaa tgatcaccga agaaagccgt ccggttctgg agttcctgca gatgctgtgt 2040ccgggtgaaa tgatcaccga agaaagccgt ccggttctgg agttcctgca gatgctgtgt 2040
gaaatcggcg ctcactatcc gggctttgaa accgatattc acggtgcata ccgtcaggct 2100gaaatcggcg ctcactatcc gggctttgaa accgatattc acggtgcata ccgtcaggct 2100
gatggccgct ataccgttaa ggtattgaaa gaagaaagca aaaaataa 2148gatggccgct ataccgttaa ggtattgaaa gaagaaagca aaaaataa 2148
<210> 12<210> 12
<211> 715<211> 715
<212> PRT<212> PRT
<213> 大肠杆菌(Escherichia coli)<213> Escherichia coli
<400> 12<400> 12
Met Asn Val Ile Ala Ile Leu Asn His Met Gly Val Tyr Phe Lys GluMet Asn Val Ile Ala Ile Leu Asn His Met Gly Val Tyr Phe Lys Glu
1 5 10 151 5 10 15
Glu Pro Ile Arg Glu Leu His Arg Ala Leu Glu Arg Leu Asn Phe GlnGlu Pro Ile Arg Glu Leu His Arg Ala Leu Glu Arg Leu Asn Phe Gln
20 25 30 20 25 30
Ile Val Tyr Pro Asn Asp Arg Asp Asp Leu Leu Lys Leu Ile Glu AsnIle Val Tyr Pro Asn Asp Arg Asp Asp Leu Leu Lys Leu Ile Glu Asn
35 40 45 35 40 45
Asn Ala Arg Leu Cys Gly Val Ile Phe Asp Trp Asp Lys Tyr Asn LeuAsn Ala Arg Leu Cys Gly Val Ile Phe Asp Trp Asp Lys Tyr Asn Leu
50 55 60 50 55 60
Glu Leu Cys Glu Glu Ile Ser Lys Met Asn Glu Asn Leu Pro Leu TyrGlu Leu Cys Glu Glu Ile Ser Lys Met Asn Glu Asn Leu Pro Leu Tyr
65 70 75 8065 70 75 80
Ala Phe Ala Asn Thr Tyr Ser Thr Leu Asp Val Ser Leu Asn Asp LeuAla Phe Ala Asn Thr Tyr Ser Thr Leu Asp Val Ser Leu Asn Asp Leu
85 90 95 85 90 95
Arg Leu Gln Ile Ser Phe Phe Glu Tyr Ala Leu Gly Ala Ala Glu AspArg Leu Gln Ile Ser Phe Phe Glu Tyr Ala Leu Gly Ala Ala Glu Asp
100 105 110 100 105 110
Ile Ala Asn Lys Ile Lys Gln Thr Thr Asp Glu Tyr Ile Asn Thr IleIle Ala Asn Lys Ile Lys Gln Thr Thr Asp Glu Tyr Ile Asn Thr Ile
115 120 125 115 120 125
Leu Pro Pro Leu Thr Lys Ala Leu Phe Lys Tyr Val Arg Glu Gly LysLeu Pro Pro Leu Thr Lys Ala Leu Phe Lys Tyr Val Arg Glu Gly Lys
130 135 140 130 135 140
Tyr Thr Phe Cys Thr Pro Gly His Met Gly Gly Thr Ala Phe Gln LysTyr Thr Phe Cys Thr Pro Gly His Met Gly Gly Thr Ala Phe Gln Lys
145 150 155 160145 150 155 160
Ser Pro Val Gly Ser Leu Phe Tyr Asp Phe Phe Gly Pro Asn Thr MetSer Pro Val Gly Ser Leu Phe Tyr Asp Phe Phe Gly Pro Asn Thr Met
165 170 175 165 170 175
Lys Ser Asp Ile Ser Ile Ser Val Ser Glu Leu Gly Ser Leu Leu AspLys Ser Asp Ile Ser Ile Ser Val Ser Glu Leu Gly Ser Leu Leu Asp
180 185 190 180 185 190
His Ser Gly Pro His Lys Glu Ala Glu Gln Tyr Ile Ala Arg Val PheHis Ser Gly Pro His Lys Glu Ala Glu Gln Tyr Ile Ala Arg Val Phe
195 200 205 195 200 205
Asn Ala Asp Arg Ser Tyr Met Val Thr Asn Gly Thr Ser Thr Ala AsnAsn Ala Asp Arg Ser Tyr Met Val Thr Asn Gly Thr Ser Thr Ala Asn
210 215 220 210 215 220
Lys Ile Val Gly Met Tyr Ser Ala Pro Ala Gly Ser Thr Ile Leu IleLys Ile Val Gly Met Tyr Ser Ala Pro Ala Gly Ser Thr Ile Leu Ile
225 230 235 240225 230 235 240
Asp Arg Asn Cys His Lys Ser Leu Thr His Leu Met Met Met Ser AspAsp Arg Asn Cys His Lys Ser Leu Thr His Leu Met Met Met Ser Asp
245 250 255 245 250 255
Val Thr Pro Ile Tyr Phe Arg Pro Thr Arg Asn Ala Tyr Gly Ile LeuVal Thr Pro Ile Tyr Phe Arg Pro Thr Arg Asn Ala Tyr Gly Ile Leu
260 265 270 260 265 270
Gly Gly Ile Pro Gln Ser Glu Phe Gln His Ala Thr Ile Ala Lys ArgGly Gly Ile Pro Gln Ser Glu Phe Gln His Ala Thr Ile Ala Lys Arg
275 280 285 275 280 285
Val Lys Glu Thr Pro Asn Ala Thr Trp Pro Val His Ala Val Ile ThrVal Lys Glu Thr Pro Asn Ala Thr Trp Pro Val His Ala Val Ile Thr
290 295 300 290 295 300
Asn Ser Thr Tyr Asp Gly Leu Leu Tyr Asn Thr Asp Phe Ile Lys LysAsn Ser Thr Tyr Asp Gly Leu Leu Tyr Asn Thr Asp Phe Ile Lys Lys
305 310 315 320305 310 315 320
Thr Leu Asp Val Lys Ser Ile His Phe Asp Ser Ala Trp Val Pro TyrThr Leu Asp Val Lys Ser Ile His Phe Asp Ser Ala Trp Val Pro Tyr
325 330 335 325 330 335
Thr Asn Phe Ser Pro Ile Tyr Glu Gly Lys Cys Gly Met Ser Gly GlyThr Asn Phe Ser Pro Ile Tyr Glu Gly Lys Cys Gly Met Ser Gly Gly
340 345 350 340 345 350
Arg Val Glu Gly Lys Val Ile Tyr Glu Thr Gln Ser Thr His Lys LeuArg Val Glu Gly Lys Val Ile Tyr Glu Thr Gln Ser Thr His Lys Leu
355 360 365 355 360 365
Leu Ala Ala Phe Ser Gln Ala Ser Met Ile His Val Lys Gly Asp ValLeu Ala Ala Phe Ser Gln Ala Ser Met Ile His Val Lys Gly Asp Val
370 375 380 370 375 380
Asn Glu Glu Thr Phe Asn Glu Ala Tyr Met Met His Thr Thr Thr SerAsn Glu Glu Thr Phe Asn Glu Ala Tyr Met Met His Thr Thr Thr Ser
385 390 395 400385 390 395 400
Pro His Tyr Gly Ile Val Ala Ser Thr Glu Thr Ala Ala Ala Met MetPro His Tyr Gly Ile Val Ala Ser Thr Glu Thr Ala Ala Ala Met Met
405 410 415 405 410 415
Lys Gly Asn Ala Gly Lys Arg Leu Ile Asn Gly Ser Ile Glu Arg AlaLys Gly Asn Ala Gly Lys Arg Leu Ile Asn Gly Ser Ile Glu Arg Ala
420 425 430 420 425 430
Ile Lys Phe Arg Lys Glu Ile Lys Arg Leu Arg Thr Glu Ser Asp GlyIle Lys Phe Arg Lys Glu Ile Lys Arg Leu Arg Thr Glu Ser Asp Gly
435 440 445 435 440 445
Trp Phe Phe Asp Val Trp Gln Pro Asp His Ile Asp Thr Thr Glu CysTrp Phe Phe Asp Val Trp Gln Pro Asp His Ile Asp Thr Thr Glu Cys
450 455 460 450 455 460
Trp Pro Leu Arg Ser Asp Ser Thr Trp His Gly Phe Lys Asn Ile AspTrp Pro Leu Arg Ser Asp Ser Thr Trp His Gly Phe Lys Asn Ile Asp
465 470 475 480465 470 475 480
Asn Glu His Met Tyr Leu Asp Pro Ile Lys Val Thr Leu Leu Thr ProAsn Glu His Met Tyr Leu Asp Pro Ile Lys Val Thr Leu Leu Thr Pro
485 490 495 485 490 495
Gly Met Glu Lys Asp Gly Thr Met Ser Asp Phe Gly Ile Pro Ala SerGly Met Glu Lys Asp Gly Thr Met Ser Asp Phe Gly Ile Pro Ala Ser
500 505 510 500 505 510
Ile Val Ala Lys Tyr Leu Asp Glu His Gly Ile Val Val Glu Lys ThrIle Val Ala Lys Tyr Leu Asp Glu His Gly Ile Val Val Glu Lys Thr
515 520 525 515 520 525
Gly Pro Tyr Asn Leu Leu Phe Leu Phe Ser Ile Gly Ile Asp Lys ThrGly Pro Tyr Asn Leu Leu Phe Leu Phe Ser Ile Gly Ile Asp Lys Thr
530 535 540 530 535 540
Lys Ala Leu Ser Leu Leu Arg Ala Leu Thr Asp Phe Lys Arg Ala PheLys Ala Leu Ser Leu Leu Arg Ala Leu Thr Asp Phe Lys Arg Ala Phe
545 550 555 560545 550 555 560
Asp Leu Asn Leu Arg Val Lys Asn Met Leu Pro Ser Leu Tyr Arg GluAsp Leu Asn Leu Arg Val Lys Asn Met Leu Pro Ser Leu Tyr Arg Glu
565 570 575 565 570 575
Asp Pro Glu Phe Tyr Glu Asn Met Arg Ile Gln Glu Leu Ala Gln AsnAsp Pro Glu Phe Tyr Glu Asn Met Arg Ile Gln Glu Leu Ala Gln Asn
580 585 590 580 585 590
Ile His Lys Leu Ile Val His His Asn Leu Pro Asp Leu Met Tyr ArgIle His Lys Leu Ile Val His His Asn Leu Pro Asp Leu Met Tyr Arg
595 600 605 595 600 605
Ala Phe Glu Val Leu Pro Thr Met Val Met Thr Pro Tyr Ala Ala PheAla Phe Glu Val Leu Pro Thr Met Val Met Thr Pro Tyr Ala Ala Phe
610 615 620 610 615 620
Gln Lys Glu Leu His Gly Met Thr Glu Glu Val Tyr Leu Asp Glu MetGln Lys Glu Leu His Gly Met Thr Glu Glu Val Tyr Leu Asp Glu Met
625 630 635 640625 630 635 640
Val Gly Arg Ile Asn Ala Asn Met Ile Leu Pro Tyr Pro Pro Gly ValVal Gly Arg Ile Asn Ala Asn Met Ile Leu Pro Tyr Pro Pro Gly Val
645 650 655 645 650 655
Pro Leu Val Met Pro Gly Glu Met Ile Thr Glu Glu Ser Arg Pro ValPro Leu Val Met Pro Gly Glu Met Ile Thr Glu Glu Ser Arg Pro Val
660 665 670 660 665 670
Leu Glu Phe Leu Gln Met Leu Cys Glu Ile Gly Ala His Tyr Pro GlyLeu Glu Phe Leu Gln Met Leu Cys Glu Ile Gly Ala His Tyr Pro Gly
675 680 685 675 680 685
Phe Glu Thr Asp Ile His Gly Ala Tyr Arg Gln Ala Asp Gly Arg TyrPhe Glu Thr Asp Ile His Gly Ala Tyr Arg Gln Ala Asp Gly Arg Tyr
690 695 700 690 695 700
Thr Val Lys Val Leu Lys Glu Glu Ser Lys LysThr Val Lys Val Leu Lys Glu Glu Ser Lys Lys
705 710 715705 710 715
<210> 13<210> 13
<211> 1642<211> 1642
<212> DNA<212> DNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence (Artificial Sequence)
<400> 13<400> 13
aagcgaacga acaacaagaa ggaaaacaag aacgagcaaa gaacacaggg ggaaaagaag 60aagcgaacga acaacaagaa ggaaaacaag aacgagcaaa gaacacaggg ggaaaagaag 60
aacccaccgg aaccacgcgc gcgaacgcga acccagagac ccgaacgacc ggacgacaaa 120aacccaccgg aaccacgcgc gcgaacgcga accgagac ccgaacgacc ggacgacaaa 120
aacgacgaaa acaagcgcgc ggcggcgaga cgggaaaaaa accgagcggc gaagaaaagc 180aacgacgaaa acaagcgcgc ggcggcgaga cgggaaaaaa accgagcggc gaagaaaagc 180
aaaagaacga gaaccgccgg acgcgcgcaa acgaccaccc gagaagccga agaccgcgac 240aaaagaacga gaaccgccgg acgcgcgcaa acgaccacccc gagaagccga agaccgcgac 240
agaagccgaa agcgcggggc gcgaagaagc aaaagacaag cagaccacga cgaaaacaac 300agaagccgaa agcgcggggc gcgaagaagc aaaagacaag cagaccacga cgaaaacaac 300
acacgccccg cgacaaagca cgaaaagcgg aaggaaaaac cgacccggca cagggcggac 360acacgccccg cgacaaagca cgaaaagcgg aaggaaaaac cgacccggca cagggcggac 360
gcaccagaaa agcccggagg agccgcagac ggccgaaacc agaaacgaac cacagacgaa 420gcaccagaaa agcccggagg agccgcagac ggccgaaacc agaaacgaac cacagacgaa 420
cgggccgcgg acacagggcc acacaaagaa gcagaacaga acgccgcgca acgcagaccg 480cgggccgcgg acacagggcc acacaaagaa gcagaacaga acgccgcgca acgcagaccg 480
cagcacaggg accaacggac ccacgcgaac aaaagggaga ccgcccagca ggcagcacca 540cagcacagggg accaacggac ccacgcgaac aaaagggaga ccgcccagca ggcagcacca 540
cgagaccgaa cgccacaaac gcgacccacc gagagagagc gagacgccaa caccgcccga 600cgagaccgaa cgccacaaac gcgacccacc gagagagagc gagacgccaa caccgcccga 600
cccgaacgca cggacgggga cccacagagg aaccagcacg caccagcaag cgcggaaaga 660cccgaacgca cggacgggga cccacagagg aaccagcacg caccagcaag cgcggaaaga 660
aacaccaaac gcaaccggcc ggacagcgaa accaaccacc agaggcgcga caacaccgac 720aacaccaaac gcaaccggcc ggacagcgaa accaaccacc agaggcgcga caacaccgac 720
cacaagaaaa cacggaggaa accaccacga cccgcggggg ccacaccaac ccaccgaacg 780cacaagaaaa cacggaggaa accacacga cccgcggggg ccacaccaac ccaccgaacg 780
aaggaaagcg gagagcgggg ccggagaagg gaaaggaacg aaacccagcc accacaaacg 840aaggaaagcg gagagcgggg ccggagaagg gaaaggaacg aaacccagcc accacaaacg 840
cggcggcgcc caggcccaga ccacgaaagg gacgaaacga agaaaccaac gaagccacag 900cggcggcgcc caggcccaga ccacgaaagg gacgaaacga agaaaccaac gaagccacag 900
agcacaccac caccccgcac acggacgggc gccacgaaac cgcgcggcga gagaaaggca 960agcacaccac caccccgcac acggacgggc gccacgaaac cgcgcggcga gagaaaggca 960
agcaggaagc gcgacaacgg cagaacggcg acaaaccgaa agagacaaac gcgagaacgg 1020agcaggaagc gcgacaacgg cagaacggcg acaaaccgaa agagacaaac gcgagaacgg 1020
aacgaggcgg cgagaggcag ccggacaacg aacgacgaag cggccgcgcg cgacagcacc 1080aacgaggcgg cgagaggcag ccggacaacg aacgacgaag cggccgcgcg cgacagcacc 1080
ggcacggcca aaaacacgaa acgagcacag acgacccgac aaagcacccg cgacccgggg 1140ggcacggcca aaaacacgaa acgagcacag acgacccgac aaagcacccg cgacccgggg 1140
aggaaaaaga cggcaccaga gcgacggacc ggccagcacg ggcgaaaacc cgacgaacag 1200aggaaaaaga cggcaccaga gcgacggacc ggccagcacg ggcgaaaacc cgacgaacag 1200
gcacgggaga aaaccggccg aaaccgcgcc gcagcacgga cgaaagacca aagcacgagc 1260gcacgggaga aaaccggccg aaaccgcgcc gcagcacgga cgaaagacca aagcacgagc 1260
cgcgcggccg acgacaaacg gcgcgaccga accgcgggaa aaacagcgcc gccgacggaa 1320cgcgcggccg acgacaaacg gcgcgaccga accgcgggaa aaacagcgcc gccgacggaa 1320
gaccgaacag aaaacagcga caggaacggc cagaaaccac aaacgagcac cacaacgccg 1380gaccgaacag aaaacagcga caggaacggc cagaaaccac aaacgagcac cacaacgccg 1380
gacgagacgc gcagaaggcg ccgacgagga agacccgagc gcaccagaaa gagcgcacgg 1440gacgagacgc gcagaaggcg ccgacgagga agacccgagc gcaccagaaa gagcgcacgg 1440
agaccgaaga agacccgacg aaaggaggcg aaacgccaaa gaccccgacc cgccgggagc 1500agaccgaaga agaccgacg aaaggaggcg aaacgccaaa gaccccgacc cgccgggagc 1500
ccggaagccg gggaaagaca ccgaagaaag ccgccggcgg agccgcagag cgggaaacgg 1560ccggaagccg gggaaagaca ccgaagaaag ccgccggcgg agccgcagag cgggaaacgg 1560
cgccacaccg ggcgaaaccg aacacgggca accgcaggcg aggccgcaac cgaaggagaa 1620cgccacaccg ggcgaaaccg aacacgggca accgcaggcg aggccgcaac cgaaggagaa 1620
agaagaaagc aaaaaaaaca ga 1642agaagaaagc aaaaaaaaca ga 1642
<210> 14<210> 14
<211> 236<211> 236
<212> PRT<212> PRT
<213> 蘑菇珊瑚(mushroom coral)<213> mushroom coral (mushroom coral)
<400> 14<400> 14
Met Val Ser Lys Gly Glu Glu Asp Asn Met Ala Ile Ile Lys Glu PheMet Val Ser Lys Gly Glu Glu Asp Asn Met Ala Ile Ile Lys Glu Phe
1 5 10 151 5 10 15
Met Arg Phe Lys Val His Met Glu Gly Ser Val Asn Gly His Glu PheMet Arg Phe Lys Val His Met Glu Gly Ser Val Asn Gly His Glu Phe
20 25 30 20 25 30
Glu Ile Glu Gly Glu Gly Glu Gly Arg Pro Tyr Glu Gly Thr Gln ThrGlu Ile Glu Gly Glu Gly Glu Gly Arg Pro Tyr Glu Gly Thr Gln Thr
35 40 45 35 40 45
Ala Lys Leu Lys Val Thr Lys Gly Gly Pro Leu Pro Phe Ala Trp AspAla Lys Leu Lys Val Thr Lys Gly Gly Pro Leu Pro Phe Ala Trp Asp
50 55 60 50 55 60
Ile Leu Ser Pro Gln Phe Met Tyr Gly Ser Lys Ala Tyr Val Lys HisIle Leu Ser Pro Gln Phe Met Tyr Gly Ser Lys Ala Tyr Val Lys His
65 70 75 8065 70 75 80
Pro Ala Asp Ile Pro Asp Tyr Leu Lys Leu Ser Phe Pro Glu Gly PhePro Ala Asp Ile Pro Asp Tyr Leu Lys Leu Ser Phe Pro Glu Gly Phe
85 90 95 85 90 95
Lys Trp Glu Arg Val Met Asn Phe Glu Asp Gly Gly Val Val Thr ValLys Trp Glu Arg Val Met Asn Phe Glu Asp Gly Gly Val Val Thr Val
100 105 110 100 105 110
Thr Gln Asp Ser Ser Leu Gln Asp Gly Glu Phe Ile Tyr Lys Val LysThr Gln Asp Ser Ser Leu Gln Asp Gly Glu Phe Ile Tyr Lys Val Lys
115 120 125 115 120 125
Leu Arg Gly Thr Asn Phe Pro Ser Asp Gly Pro Val Met Gln Lys LysLeu Arg Gly Thr Asn Phe Pro Ser Asp Gly Pro Val Met Gln Lys Lys
130 135 140 130 135 140
Thr Met Gly Trp Glu Ala Ser Ser Glu Arg Met Tyr Pro Glu Asp GlyThr Met Gly Trp Glu Ala Ser Ser Glu Arg Met Tyr Pro Glu Asp Gly
145 150 155 160145 150 155 160
Ala Leu Lys Gly Glu Ile Lys Gln Arg Leu Lys Leu Lys Asp Gly GlyAla Leu Lys Gly Glu Ile Lys Gln Arg Leu Lys Leu Lys Asp Gly Gly
165 170 175 165 170 175
His Tyr Asp Ala Glu Val Lys Thr Thr Tyr Lys Ala Lys Lys Pro ValHis Tyr Asp Ala Glu Val Lys Thr Thr Tyr Lys Ala Lys Lys Pro Val
180 185 190 180 185 190
Gln Leu Pro Gly Ala Tyr Asn Val Asn Ile Lys Leu Asp Ile Thr SerGln Leu Pro Gly Ala Tyr Asn Val Asn Ile Lys Leu Asp Ile Thr Ser
195 200 205 195 200 205
His Asn Glu Asp Tyr Thr Ile Val Glu Gln Tyr Glu Arg Ala Glu GlyHis Asn Glu Asp Tyr Thr Ile Val Glu Gln Tyr Glu Arg Ala Glu Gly
210 215 220 210 215 220
Arg His Ser Thr Gly Gly Met Asp Glu Leu Tyr LysArg His Ser Thr Gly Gly Met Asp Glu Leu Tyr Lys
225 230 235225 230 235
<210> 15<210> 15
<211> 711<211> 711
<212> DNA<212> DNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence (Artificial Sequence)
<400> 15<400> 15
atggtgtcta aaggcgagga agataatatg gcgattatca aagaatttat gcgttttaaa 60atggtgtcta aaggcgagga agataatatg gcgattatca aagaatttat gcgttttaaa 60
gtgcatatgg aaggcagcgt gaatgggcat gagtttgaaa ttgaaggcga aggagaaggc 120gtgcatatgg aaggcagcgt gaatgggcat gagtttgaaa ttgaaggcga aggagaaggc 120
cgtccgtatg aaggcaccca gaccgctaaa ctgaaagtga ccaaaggcgg accactgccg 180cgtccgtatg aaggcaccca gaccgctaaa ctgaaagtga ccaaaggcgg accactgccg 180
tttgcgtggg acattctgag cccgcagttt atgtatggca gcaaagcgta tgtgaaacat 240tttgcgtggg attctgag cccgcagttt atgtatggca gcaaagcgta tgtgaaacat 240
ccggcggata ttccggatta tctgaaactg agctttccgg agggcttcaa atgggaacgt 300ccggcggata ttccggatta tctgaaactg agctttccgg agggcttcaa atgggaacgt 300
gtgatgaatt ttgaagatgg cggcgtggtg accgtgaccc aggatagcag cctgcaagac 360gtgatgaatt ttgaagatgg cggcgtggtg accgtgaccc aggatagcag cctgcaagac 360
ggcgaattca tttacaaggt gaagctgcgt ggcaccaact ttcccagcga tggcccggtg 420ggcgaattca tttacaaggt gaagctgcgt ggcaccaact ttcccagcga tggcccggtg 420
atgcagaaaa agaccatggg ctgggaggcg agcagcgaac gtatgtaccc ggaggatggc 480atgcagaaaa agaccatggg ctgggaggcg agcagcgaac gtatgtaccc ggaggatggc 480
gcgctgaagg gcgaaattaa gcagcgtctg aagttaaaag atggtgggca ctatgatgcg 540gcgctgaagg gcgaaattaa gcagcgtctg aagttaaaag atggtgggca ctatgatgcg 540
gaagtgaaaa ccacctataa agcgaaaaaa ccggtgcagt taccaggcgc ttataatgtg 600gaagtgaaaa ccacctataa agcgaaaaaa ccggtgcagt taccaggcgc ttataatgtg 600
aacattaagc tggatattac cagccataat gaagattata ccattgtgga acagtatgag 660aacattaagc tggatattac cagccataat gaagattata ccattgtgga acagtatgag 660
cgtgcggagg gacggcatag cacgggcgga atggatgaac tgtataaata a 711cgtgcggagg gacggcatag cacgggcgga atggatgaac tgtataaata a 711
<210> 16<210> 16
<211> 59<211> 59
<212> DNA<212> DNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence (Artificial Sequence)
<400> 16<400> 16
gctctagagt ttttttggga attcacacac aggaggagct gatggtgtct aaaggcgag 59gctctagagt ttttttggga attcacacac aggagagct gatggtgtct aaaggcgag 59
<210> 17<210> 17
<211> 37<211> 37
<212> DNA<212> DNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence (Artificial Sequence)
<400> 17<400> 17
gctctagatt atttatacag ttcatccatt ccgcccg 37gctctagatt atttatacag ttcatccatt ccgcccg 37
<210> 18<210> 18
<211> 744<211> 744
<212> DNA<212> DNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence (Artificial Sequence)
<400> 18<400> 18
gtttttttgg gaattcacac acaggaggag ctgatggtgt ctaaaggcga ggaagataat 60gtttttttgg gaattcacac acaggagggag ctgatggtgt ctaaaggcga ggaagataat 60
atggcgatta tcaaagaatt tatgcgtttt aaagtgcata tggaaggcag cgtgaatggg 120atggcgatta tcaaagaatt tatgcgtttt aaagtgcata tggaaggcag cgtgaatggg 120
catgagtttg aaattgaagg cgaaggagaa ggccgtccgt atgaaggcac ccagaccgct 180catgagtttg aaattgaagg cgaaggagaa ggccgtccgt atgaaggcac ccagaccgct 180
aaactgaaag tgaccaaagg cggaccactg ccgtttgcgt gggacattct gagcccgcag 240aaactgaaag tgaccaaagg cggaccactg ccgtttgcgt gggacattct gagcccgcag 240
tttatgtatg gcagcaaagc gtatgtgaaa catccggcgg atattccgga ttatctgaaa 300tttatgtatg gcagcaaagc gtatgtgaaa catccggcgg atattccgga ttatctgaaa 300
ctgagctttc cggagggctt caaatgggaa cgtgtgatga attttgaaga tggcggcgtg 360ctgagctttc cggagggctt caaatgggaa cgtgtgatga attttgaaga tggcggcgtg 360
gtgaccgtga cccaggatag cagcctgcaa gacggcgaat tcatttacaa ggtgaagctg 420gtgaccgtga cccaggatag cagcctgcaa gacggcgaat tcatttacaa ggtgaagctg 420
cgtggcacca actttcccag cgatggcccg gtgatgcaga aaaagaccat gggctgggag 480cgtggcacca actttcccag cgatggcccg gtgatgcaga aaaagaccat gggctgggag 480
gcgagcagcg aacgtatgta cccggaggat ggcgcgctga agggcgaaat taagcagcgt 540gcgagcagcg aacgtatgta cccggaggat ggcgcgctga agggcgaaat taagcagcgt 540
ctgaagttaa aagatggtgg gcactatgat gcggaagtga aaaccaccta taaagcgaaa 600ctgaagttaa aagatggtgg gcactatgat gcggaagtga aaaccaccta taaagcgaaa 600
aaaccggtgc agttaccagg cgcttataat gtgaacatta agctggatat taccagccat 660aaaccggtgc agttaccagg cgcttataat gtgaacatta agctggatat taccagccat 660
aatgaagatt ataccattgt ggaacagtat gagcgtgcgg agggacggca tagcacgggc 720aatgaagatt ataccattgt ggaacagtat gagcgtgcgg agggacggca tagcacgggc 720
ggaatggatg aactgtataa ataa 744ggaatggatg aactgtataa ataa 744
<210> 19<210> 19
<211> 57<211> 57
<212> DNA<212> DNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence (Artificial Sequence)
<400> 19<400> 19
gatatcgaat tcttaacttt aagaaggaat atacatatgg tgtctaaagg cgaggaa 57gatatcgaat tcttaacttt aagaaggaat atacatatgg tgtctaaagg cgaggaa 57
<210> 20<210> 20
<211> 40<211> 40
<212> DNA<212> DNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence (Artificial Sequence)
<400> 20<400> 20
aaagttaaga attcgatatc ggcactggcc gtcgttttac 40aaagttaaga attcgatatc ggcactggcc gtcgttttac 40
<210> 21<210> 21
<211> 42<211> 42
<212> DNA<212> DNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence (Artificial Sequence)
<400> 21<400> 21
gaattcgagc tcggtaccat cgataagctt gatatcgaat tc 42gaattcgagc tcggtaccat cgataagctt gatatcgaat tc 42
<210> 22<210> 22
<211> 41<211> 41
<212> DNA<212> DNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence (Artificial Sequence)
<400> 22<400> 22
gatggtaccg agctcgaatt cggcactggc cgtcgtttta c 41gatggtaccg agctcgaatt cggcactggc cgtcgtttta c 41
<210> 23<210> 23
<211> 51<211> 51
<212> DNA<212> DNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence (Artificial Sequence)
<400> 23<400> 23
gaattcttaa ctttaagaag gaatatacat atgaacgtta ttgcaatatt g 51gaattcttaa ctttaagaag gaatatacat atgaacgtta ttgcaatatt g 51
<210> 24<210> 24
<211> 28<211> 28
<212> DNA<212> DNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence (Artificial Sequence)
<400> 24<400> 24
gcctctagac cacttccctt gtacgagc 28gcctctagac cacttccctt gtacgagc 28
<210> 25<210> 25
<211> 35<211> 35
<212> DNA<212> DNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence (Artificial Sequence)
<400> 25<400> 25
gccaagcttg atatcgaatt cttaacttta agaag 35gccaagcttg atatcgaatt ctaacttta agaag 35
<210> 26<210> 26
<211> 31<211> 31
<212> DNA<212> DNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence (Artificial Sequence)
<400> 26<400> 26
ggggtaccga gtgagctgat accgctcgcc g 31ggggtaccga gtgagctgat accgctcgcc g 31
<210> 27<210> 27
<211> 30<211> 30
<212> DNA<212> DNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence (Artificial Sequence)
<400> 27<400> 27
ggatcgatag ctgtttcctg tgtgaaattg 30ggatcgatag ctgtttcctg tgtgaaattg 30
<210> 28<210> 28
<211> 286<211> 286
<212> DNA<212> DNA
<213> 人工序列(Artificial Sequence)<213> Artificial Sequence (Artificial Sequence)
<400> 28<400> 28
gagtgagctg ataccgctcg ccgcagccga acgaccgagc gcagcgagtc agtgagcgag 60gagtgagctg ataccgctcg ccgcagccga acgaccgagc gcagcgagtc agtgagcgag 60
gaagcggaag agcgcccaat acgcaaaccg cctctccccg cgcgttggcc gattcattaa 120gaagcggaag agcgcccaat acgcaaaccg cctctccccg cgcgttggcc gattcattaa 120
tgcagctggc acgacaggtt tcccgactgg aaagcgggca gtgagcgcaa cgcaattaat 180tgcagctggc acgacaggtt tcccgactgg aaagcgggca gtgagcgcaa cgcaattaat 180
gtgagttagc tcactcatta ggcaccccag gctttacact ttatgcttcc ggctcgtatg 240gtgagttagc tcactcatta ggcaccccag gctttacact ttatgcttcc ggctcgtatg 240
ttgtgtggaa ttgtgagcgg ataacaattt cacacaggaa acagct 286ttgtgtggaa ttgtgagcgg ataacaattt cacacaggaa acagct 286
Claims (15)
Applications Claiming Priority (2)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| CN2019101770711 | 2019-03-08 | ||
| CN201910177071 | 2019-03-08 |
Publications (2)
| Publication Number | Publication Date |
|---|---|
| CN111662903A CN111662903A (en) | 2020-09-15 |
| CN111662903B true CN111662903B (en) | 2022-12-27 |
Family
ID=72382185
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| CN201910228659.5A Active CN111662903B (en) | 2019-03-08 | 2019-03-25 | Logarithmic phase specific promoter and application thereof |
Country Status (1)
| Country | Link |
|---|---|
| CN (1) | CN111662903B (en) |
Families Citing this family (1)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN118813659A (en) * | 2021-04-14 | 2024-10-22 | 上海凯赛生物技术股份有限公司 | Recombinant DNA, strain and use thereof for fermentation production of 1,5-pentanediamine |
Citations (4)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN104278031A (en) * | 2013-07-10 | 2015-01-14 | 中国科学院过程工程研究所 | Promoter A regulated by xanthine as well as recombinant expression vector and application of promoter A |
| CN106222190A (en) * | 2016-08-25 | 2016-12-14 | 江南大学 | A kind of bacillus subtilis logarithmic (log) phase expression system |
| CN108118059A (en) * | 2017-12-30 | 2018-06-05 | 苏州金唯智生物科技有限公司 | An improved promoter and its composition vector and application |
| CN108165551A (en) * | 2017-12-30 | 2018-06-15 | 苏州金唯智生物科技有限公司 | A kind of improved promoter and its composition T carrier and application |
Family Cites Families (2)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US20020164706A1 (en) * | 2001-01-30 | 2002-11-07 | Huang Lisa L. | High level promoters from cyanobacteria |
| MX2007015355A (en) * | 2005-06-06 | 2008-02-19 | Dow Global Technologies Inc | Mannitol induced promoter systems in bacterial host cells. |
-
2019
- 2019-03-25 CN CN201910228659.5A patent/CN111662903B/en active Active
Patent Citations (4)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN104278031A (en) * | 2013-07-10 | 2015-01-14 | 中国科学院过程工程研究所 | Promoter A regulated by xanthine as well as recombinant expression vector and application of promoter A |
| CN106222190A (en) * | 2016-08-25 | 2016-12-14 | 江南大学 | A kind of bacillus subtilis logarithmic (log) phase expression system |
| CN108118059A (en) * | 2017-12-30 | 2018-06-05 | 苏州金唯智生物科技有限公司 | An improved promoter and its composition vector and application |
| CN108165551A (en) * | 2017-12-30 | 2018-06-15 | 苏州金唯智生物科技有限公司 | A kind of improved promoter and its composition T carrier and application |
Non-Patent Citations (2)
| Title |
|---|
| "Unprecedented High-Resolution View of Bacterial Operon Architecture Revealed by RNA Sequencing";Tyrrell Conway et al.;《mBio》;20140708;第5卷(第4期);第1-12页 * |
| "诱导型启动子在谷氨酸棒杆菌ATCC13032中调控强度的分析";陈晓雪 等;《沈阳药科大学学报》;20170228;第34卷(第2期);第169-175页 * |
Also Published As
| Publication number | Publication date |
|---|---|
| CN111662903A (en) | 2020-09-15 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| US8067210B2 (en) | Method of producing lysine by culturing a host cell expressing a polynucleotide encoding a feedback resistant aspartokinase from corynebacterium | |
| US20030017553A1 (en) | Nucleotide sequences for transcriptional regulation in corynebacterium glutamicum | |
| CN103215291B (en) | For the production of the carrier of C4H9NO2, engineering strain and method | |
| CN101849002A (en) | Method for maintaining foreign gene in cell stably | |
| CN110592036A (en) | A kind of glufosinate-ammonium dehydrogenase mutant and its application in oxidative-reduction multi-enzyme coupling production of L-glufosinate-ammonium | |
| CN111321141B (en) | Stationary phase specific promoter and application thereof | |
| CN113969269A (en) | D-amino acid oxidase mutant and its application in the preparation of L-glufosinate | |
| CN111978407B (en) | Heterologous expression method and application of lysine decarboxylase derived from thermophilic bacteria | |
| CN117904020A (en) | Construction of E.coli acetylation mutant strain and application of E.coli acetylation mutant strain in production of 3-hydroxy propionic acid | |
| CN114058560B (en) | Process for the production of glycine | |
| CN113151270A (en) | Promoter for efficiently expressing alkaline protease and application thereof | |
| CN111662903B (en) | Logarithmic phase specific promoter and application thereof | |
| CN113201514B (en) | Polypeptides having aspartokinase activity and their use for producing amino acids | |
| CN115873852A (en) | Recombinant nucleic acid sequence, genetically engineered bacteria and method for producing 1,5-pentanediamine | |
| CN111979257B (en) | A kind of recombinant DNA and its application | |
| WO2000063354A1 (en) | Novel amidase gene | |
| US7049097B2 (en) | Antibiotics-independent vector for constant high-expression and method for gene expression using the same | |
| CN118667786A (en) | Genetically engineered bacteria expressing ectoine synthase and construction method and application thereof | |
| CN110964682B (en) | L-lysine-tolerant bacteria, producing bacteria and use thereof | |
| CN115873880B (en) | Recombinant nucleic acid sequence, recombinant expression vector and genetically engineered bacteria | |
| CN114181288A (en) | Method for preparing L-valine and gene used therefor and protein encoded by the gene | |
| CN108456669B (en) | A kind of ribosome binding site, recombinant expression plasmid, transformant and application thereof | |
| JP2002153286A (en) | Enzyme purification method | |
| CN115678863A (en) | Succinate dehydrogenase mutant, recombinant microorganism, preparation method and application thereof | |
| WO2011125467A1 (en) | Plasmid vector |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| PB01 | Publication | ||
| PB01 | Publication | ||
| SE01 | Entry into force of request for substantive examination | ||
| SE01 | Entry into force of request for substantive examination | ||
| TA01 | Transfer of patent application right |
Effective date of registration: 20221205 Address after: 201203 Shanghai Zhangjiang High Tech Park of Pudong New Area Cailun Road No. 5 No. 1690 Applicant after: CATHAY R&D CENTER Co.,Ltd. Applicant after: CIBT USA Applicant after: Kasai (Shanghai) Biotechnology Co.,Ltd. Address before: 201203 Shanghai Zhangjiang High Tech Park of Pudong New Area Cailun Road No. 5 No. 1690 Applicant before: CATHAY R&D CENTER Co.,Ltd. Applicant before: CIBT USA |
|
| TA01 | Transfer of patent application right | ||
| GR01 | Patent grant | ||
| GR01 | Patent grant |