CN111218513B - Peripheral blood extracellular vesicle microRNA biomarker for early diagnosis of lung cancer and application thereof - Google Patents
Peripheral blood extracellular vesicle microRNA biomarker for early diagnosis of lung cancer and application thereof Download PDFInfo
- Publication number
- CN111218513B CN111218513B CN202010329484.XA CN202010329484A CN111218513B CN 111218513 B CN111218513 B CN 111218513B CN 202010329484 A CN202010329484 A CN 202010329484A CN 111218513 B CN111218513 B CN 111218513B
- Authority
- CN
- China
- Prior art keywords
- hsa
- peripheral blood
- mirna
- mir
- lung cancer
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Active
Links
- 239000002679 microRNA Substances 0.000 title claims abstract description 82
- 239000000090 biomarker Substances 0.000 title claims abstract description 39
- 210000005259 peripheral blood Anatomy 0.000 title claims abstract description 39
- 239000011886 peripheral blood Substances 0.000 title claims abstract description 39
- 206010058467 Lung neoplasm malignant Diseases 0.000 title claims abstract description 38
- 201000005202 lung cancer Diseases 0.000 title claims abstract description 38
- 208000020816 lung neoplasm Diseases 0.000 title claims abstract description 38
- 238000013399 early diagnosis Methods 0.000 title claims abstract description 16
- 108700011259 MicroRNAs Proteins 0.000 title description 4
- 108091070501 miRNA Proteins 0.000 claims abstract description 87
- 206010056342 Pulmonary mass Diseases 0.000 claims abstract description 36
- 108091070514 Homo sapiens let-7b stem-loop Proteins 0.000 claims abstract description 10
- 108091068840 Homo sapiens miR-101-1 stem-loop Proteins 0.000 claims abstract description 10
- 108091065458 Homo sapiens miR-101-2 stem-loop Proteins 0.000 claims abstract description 10
- 108091072926 Homo sapiens miR-3168 stem-loop Proteins 0.000 claims abstract description 10
- 108091069006 Homo sapiens miR-125b-1 stem-loop Proteins 0.000 claims abstract description 9
- 108091069087 Homo sapiens miR-125b-2 stem-loop Proteins 0.000 claims abstract description 9
- 108091069088 Homo sapiens miR-150 stem-loop Proteins 0.000 claims abstract description 9
- 238000001514 detection method Methods 0.000 claims description 28
- 230000003211 malignant effect Effects 0.000 claims description 28
- 239000000523 sample Substances 0.000 claims description 24
- 238000000034 method Methods 0.000 claims description 18
- 239000003153 chemical reaction reagent Substances 0.000 claims description 9
- 239000003550 marker Substances 0.000 claims description 7
- 238000012167 Small RNA sequencing Methods 0.000 claims description 6
- 239000013068 control sample Substances 0.000 claims description 3
- 238000012937 correction Methods 0.000 claims description 3
- 238000013178 mathematical model Methods 0.000 claims description 2
- 238000002360 preparation method Methods 0.000 claims description 2
- 238000011160 research Methods 0.000 abstract description 8
- 238000012549 training Methods 0.000 description 21
- 206010054107 Nodule Diseases 0.000 description 11
- 238000004458 analytical method Methods 0.000 description 9
- 238000003745 diagnosis Methods 0.000 description 9
- 230000001575 pathological effect Effects 0.000 description 9
- 238000005516 engineering process Methods 0.000 description 8
- 238000010200 validation analysis Methods 0.000 description 8
- 210000004369 blood Anatomy 0.000 description 6
- 239000008280 blood Substances 0.000 description 6
- 108091070519 Homo sapiens miR-19b-1 stem-loop Proteins 0.000 description 5
- 108091070495 Homo sapiens miR-19b-2 stem-loop Proteins 0.000 description 5
- 239000000091 biomarker candidate Substances 0.000 description 5
- 210000004027 cell Anatomy 0.000 description 5
- 238000003384 imaging method Methods 0.000 description 5
- 108090000623 proteins and genes Proteins 0.000 description 5
- 230000002685 pulmonary effect Effects 0.000 description 5
- 238000012163 sequencing technique Methods 0.000 description 5
- 238000012795 verification Methods 0.000 description 5
- 206010028980 Neoplasm Diseases 0.000 description 4
- 238000011161 development Methods 0.000 description 4
- 230000018109 developmental process Effects 0.000 description 4
- 201000010099 disease Diseases 0.000 description 4
- 208000037265 diseases, disorders, signs and symptoms Diseases 0.000 description 4
- 238000011156 evaluation Methods 0.000 description 4
- 102000004169 proteins and genes Human genes 0.000 description 4
- 230000035945 sensitivity Effects 0.000 description 4
- 239000006228 supernatant Substances 0.000 description 4
- 108091070512 Homo sapiens let-7d stem-loop Proteins 0.000 description 3
- 108091070491 Homo sapiens miR-16-1 stem-loop Proteins 0.000 description 3
- 108091068927 Homo sapiens miR-16-2 stem-loop Proteins 0.000 description 3
- 108091070489 Homo sapiens miR-17 stem-loop Proteins 0.000 description 3
- 108091064515 Homo sapiens miR-503 stem-loop Proteins 0.000 description 3
- 108091063565 Homo sapiens miR-532 stem-loop Proteins 0.000 description 3
- 108091068856 Homo sapiens miR-98 stem-loop Proteins 0.000 description 3
- 238000010276 construction Methods 0.000 description 3
- 238000013211 curve analysis Methods 0.000 description 3
- 230000000694 effects Effects 0.000 description 3
- 239000012528 membrane Substances 0.000 description 3
- 239000000047 product Substances 0.000 description 3
- 238000001356 surgical procedure Methods 0.000 description 3
- 102100025222 CD63 antigen Human genes 0.000 description 2
- 102100037904 CD9 antigen Human genes 0.000 description 2
- 101000836492 Dictyostelium discoideum ALG-2 interacting protein X Proteins 0.000 description 2
- LFQSCWFLJHTTHZ-UHFFFAOYSA-N Ethanol Chemical compound CCO LFQSCWFLJHTTHZ-UHFFFAOYSA-N 0.000 description 2
- 101000934368 Homo sapiens CD63 antigen Proteins 0.000 description 2
- 101000738354 Homo sapiens CD9 antigen Proteins 0.000 description 2
- 101001134621 Homo sapiens Programmed cell death 6-interacting protein Proteins 0.000 description 2
- 101000613251 Homo sapiens Tumor susceptibility gene 101 protein Proteins 0.000 description 2
- 206010036790 Productive cough Diseases 0.000 description 2
- 102100033344 Programmed cell death 6-interacting protein Human genes 0.000 description 2
- 102000006276 Syntenins Human genes 0.000 description 2
- 108010083130 Syntenins Proteins 0.000 description 2
- 239000006180 TBST buffer Substances 0.000 description 2
- 102100040879 Tumor susceptibility gene 101 protein Human genes 0.000 description 2
- 238000013276 bronchoscopy Methods 0.000 description 2
- 201000011510 cancer Diseases 0.000 description 2
- 210000000038 chest Anatomy 0.000 description 2
- 230000012202 endocytosis Effects 0.000 description 2
- 210000001808 exosome Anatomy 0.000 description 2
- 238000000605 extraction Methods 0.000 description 2
- 239000000706 filtrate Substances 0.000 description 2
- 239000012634 fragment Substances 0.000 description 2
- 238000011534 incubation Methods 0.000 description 2
- 210000004072 lung Anatomy 0.000 description 2
- 108020004999 messenger RNA Proteins 0.000 description 2
- 210000002487 multivesicular body Anatomy 0.000 description 2
- 108020004707 nucleic acids Proteins 0.000 description 2
- 102000039446 nucleic acids Human genes 0.000 description 2
- 150000007523 nucleic acids Chemical class 0.000 description 2
- 230000002093 peripheral effect Effects 0.000 description 2
- 238000001556 precipitation Methods 0.000 description 2
- 238000011002 quantification Methods 0.000 description 2
- 238000002271 resection Methods 0.000 description 2
- 230000000717 retained effect Effects 0.000 description 2
- 238000010839 reverse transcription Methods 0.000 description 2
- 238000012502 risk assessment Methods 0.000 description 2
- 238000013058 risk prediction model Methods 0.000 description 2
- 238000012216 screening Methods 0.000 description 2
- 230000000391 smoking effect Effects 0.000 description 2
- 210000003802 sputum Anatomy 0.000 description 2
- 208000024794 sputum Diseases 0.000 description 2
- 230000004083 survival effect Effects 0.000 description 2
- 238000012360 testing method Methods 0.000 description 2
- 108091032973 (ribonucleotides)n+m Proteins 0.000 description 1
- 102000040650 (ribonucleotides)n+m Human genes 0.000 description 1
- 108020005345 3' Untranslated Regions Proteins 0.000 description 1
- 108091032955 Bacterial small RNA Proteins 0.000 description 1
- 108020004414 DNA Proteins 0.000 description 1
- 238000013382 DNA quantification Methods 0.000 description 1
- 102000010834 Extracellular Matrix Proteins Human genes 0.000 description 1
- 108010037362 Extracellular Matrix Proteins Proteins 0.000 description 1
- SXRSQZLOMIGNAQ-UHFFFAOYSA-N Glutaraldehyde Chemical compound O=CCCCC=O SXRSQZLOMIGNAQ-UHFFFAOYSA-N 0.000 description 1
- 206010018691 Granuloma Diseases 0.000 description 1
- 101710088172 HTH-type transcriptional regulator RipA Proteins 0.000 description 1
- 206010018910 Haemolysis Diseases 0.000 description 1
- 208000002927 Hamartoma Diseases 0.000 description 1
- 108091030146 MiRBase Proteins 0.000 description 1
- 108700020796 Oncogene Proteins 0.000 description 1
- 241000283973 Oryctolagus cuniculus Species 0.000 description 1
- 238000012408 PCR amplification Methods 0.000 description 1
- 229910019142 PO4 Inorganic materials 0.000 description 1
- 239000002033 PVDF binder Substances 0.000 description 1
- XUIMIQQOPSSXEZ-UHFFFAOYSA-N Silicon Chemical compound [Si] XUIMIQQOPSSXEZ-UHFFFAOYSA-N 0.000 description 1
- 108700025716 Tumor Suppressor Genes Proteins 0.000 description 1
- 102000044209 Tumor Suppressor Genes Human genes 0.000 description 1
- 239000002253 acid Substances 0.000 description 1
- 208000009956 adenocarcinoma Diseases 0.000 description 1
- 230000006907 apoptotic process Effects 0.000 description 1
- 230000000903 blocking effect Effects 0.000 description 1
- 210000000481 breast Anatomy 0.000 description 1
- 210000000170 cell membrane Anatomy 0.000 description 1
- 230000005754 cellular signaling Effects 0.000 description 1
- 238000005119 centrifugation Methods 0.000 description 1
- 238000012512 characterization method Methods 0.000 description 1
- 230000008602 contraction Effects 0.000 description 1
- 238000002790 cross-validation Methods 0.000 description 1
- 238000007405 data analysis Methods 0.000 description 1
- 239000000032 diagnostic agent Substances 0.000 description 1
- 229940039227 diagnostic agent Drugs 0.000 description 1
- 238000003748 differential diagnosis Methods 0.000 description 1
- 230000004069 differentiation Effects 0.000 description 1
- 229940079593 drug Drugs 0.000 description 1
- 239000003814 drug Substances 0.000 description 1
- 238000001493 electron microscopy Methods 0.000 description 1
- 238000001962 electrophoresis Methods 0.000 description 1
- 238000002474 experimental method Methods 0.000 description 1
- 210000002744 extracellular matrix Anatomy 0.000 description 1
- 238000001914 filtration Methods 0.000 description 1
- 230000008588 hemolysis Effects 0.000 description 1
- 238000009396 hybridization Methods 0.000 description 1
- 208000026278 immune system disease Diseases 0.000 description 1
- 230000002757 inflammatory effect Effects 0.000 description 1
- 230000003902 lesion Effects 0.000 description 1
- 150000002632 lipids Chemical class 0.000 description 1
- 238000011528 liquid biopsy Methods 0.000 description 1
- 238000011068 loading method Methods 0.000 description 1
- 239000006166 lysate Substances 0.000 description 1
- 230000036210 malignancy Effects 0.000 description 1
- 239000002184 metal Substances 0.000 description 1
- 238000002156 mixing Methods 0.000 description 1
- 239000003147 molecular marker Substances 0.000 description 1
- 230000000877 morphologic effect Effects 0.000 description 1
- 239000002773 nucleotide Substances 0.000 description 1
- 125000003729 nucleotide group Chemical group 0.000 description 1
- 238000010827 pathological analysis Methods 0.000 description 1
- 230000007170 pathology Effects 0.000 description 1
- NBIIXXVUZAFLBC-UHFFFAOYSA-K phosphate Chemical compound [O-]P([O-])([O-])=O NBIIXXVUZAFLBC-UHFFFAOYSA-K 0.000 description 1
- 239000010452 phosphate Substances 0.000 description 1
- 238000007747 plating Methods 0.000 description 1
- 229920002981 polyvinylidene fluoride Polymers 0.000 description 1
- 230000002980 postoperative effect Effects 0.000 description 1
- 238000012545 processing Methods 0.000 description 1
- 238000004393 prognosis Methods 0.000 description 1
- 230000035755 proliferation Effects 0.000 description 1
- -1 rRNA Proteins 0.000 description 1
- 238000002601 radiography Methods 0.000 description 1
- 238000010814 radioimmunoprecipitation assay Methods 0.000 description 1
- 210000002966 serum Anatomy 0.000 description 1
- 235000020183 skimmed milk Nutrition 0.000 description 1
- 150000003384 small molecules Chemical class 0.000 description 1
- 238000002415 sodium dodecyl sulfate polyacrylamide gel electrophoresis Methods 0.000 description 1
- 206010041823 squamous cell carcinoma Diseases 0.000 description 1
- 238000007619 statistical method Methods 0.000 description 1
- 208000024891 symptom Diseases 0.000 description 1
- 230000001225 therapeutic effect Effects 0.000 description 1
- 210000001519 tissue Anatomy 0.000 description 1
- 238000012546 transfer Methods 0.000 description 1
- 230000009466 transformation Effects 0.000 description 1
- 238000013519 translation Methods 0.000 description 1
- XLYOFNOQVPJJNP-UHFFFAOYSA-N water Substances O XLYOFNOQVPJJNP-UHFFFAOYSA-N 0.000 description 1
Images
Classifications
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q1/00—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions
- C12Q1/68—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions involving nucleic acids
- C12Q1/6876—Nucleic acid products used in the analysis of nucleic acids, e.g. primers or probes
- C12Q1/6883—Nucleic acid products used in the analysis of nucleic acids, e.g. primers or probes for diseases caused by alterations of genetic material
- C12Q1/6886—Nucleic acid products used in the analysis of nucleic acids, e.g. primers or probes for diseases caused by alterations of genetic material for cancer
-
- G—PHYSICS
- G16—INFORMATION AND COMMUNICATION TECHNOLOGY [ICT] SPECIALLY ADAPTED FOR SPECIFIC APPLICATION FIELDS
- G16B—BIOINFORMATICS, i.e. INFORMATION AND COMMUNICATION TECHNOLOGY [ICT] SPECIALLY ADAPTED FOR GENETIC OR PROTEIN-RELATED DATA PROCESSING IN COMPUTATIONAL MOLECULAR BIOLOGY
- G16B40/00—ICT specially adapted for biostatistics; ICT specially adapted for bioinformatics-related machine learning or data mining, e.g. knowledge discovery or pattern finding
-
- G—PHYSICS
- G16—INFORMATION AND COMMUNICATION TECHNOLOGY [ICT] SPECIALLY ADAPTED FOR SPECIFIC APPLICATION FIELDS
- G16B—BIOINFORMATICS, i.e. INFORMATION AND COMMUNICATION TECHNOLOGY [ICT] SPECIALLY ADAPTED FOR GENETIC OR PROTEIN-RELATED DATA PROCESSING IN COMPUTATIONAL MOLECULAR BIOLOGY
- G16B5/00—ICT specially adapted for modelling or simulations in systems biology, e.g. gene-regulatory networks, protein interaction networks or metabolic networks
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q2600/00—Oligonucleotides characterized by their use
- C12Q2600/158—Expression markers
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q2600/00—Oligonucleotides characterized by their use
- C12Q2600/178—Oligonucleotides characterized by their use miRNA, siRNA or ncRNA
Landscapes
- Health & Medical Sciences (AREA)
- Life Sciences & Earth Sciences (AREA)
- Engineering & Computer Science (AREA)
- Chemical & Material Sciences (AREA)
- Physics & Mathematics (AREA)
- Bioinformatics & Cheminformatics (AREA)
- Medical Informatics (AREA)
- Biophysics (AREA)
- Organic Chemistry (AREA)
- Proteomics, Peptides & Aminoacids (AREA)
- Biotechnology (AREA)
- General Health & Medical Sciences (AREA)
- Pathology (AREA)
- Immunology (AREA)
- Theoretical Computer Science (AREA)
- Zoology (AREA)
- Wood Science & Technology (AREA)
- Molecular Biology (AREA)
- Analytical Chemistry (AREA)
- Spectroscopy & Molecular Physics (AREA)
- Bioinformatics & Computational Biology (AREA)
- Genetics & Genomics (AREA)
- Evolutionary Biology (AREA)
- Databases & Information Systems (AREA)
- Data Mining & Analysis (AREA)
- Software Systems (AREA)
- Oncology (AREA)
- Public Health (AREA)
- Evolutionary Computation (AREA)
- Epidemiology (AREA)
- Hospice & Palliative Care (AREA)
- Physiology (AREA)
- Microbiology (AREA)
- Computer Vision & Pattern Recognition (AREA)
- Bioethics (AREA)
- Artificial Intelligence (AREA)
- Biochemistry (AREA)
- General Engineering & Computer Science (AREA)
- Measuring Or Testing Involving Enzymes Or Micro-Organisms (AREA)
Abstract
The invention discloses a peripheral blood extracellular vesicle miRNA biomarker for early diagnosis of lung cancer, which is characterized by comprising hsa-let-7b-3p, hsa-miR-125b-5p, hsa-miR-150-5p, hsa-miR-101-3p and hsa-miR-3168. The invention is based on the peripheral blood of the lung nodule patients detected by the LDCT in clinic as a research sample, has higher specificity and is more reliable, and the research result has important significance for accurately identifying the false-positive lung cancer patients from the lung nodule patients detected by the LDCT.
Description
Technical Field
The invention relates to the technical field of early detection of diseases, in particular to a peripheral blood extracellular vesicle microRNA biomarker for early diagnosis of lung cancer.
Background
Lung cancer is the most common malignancy with the highest incidence and mortality worldwide. Early stage lung cancer is insidious and usually has no symptoms, but most patients are in the middle and late stage at the initial diagnosis and lose the chance of surgical resection. The five-year survival rate of patients with advanced lung cancer is less than 5 percent, and the five-year survival rate of patients with early lung cancer can reach more than 90 percent. Therefore, early diagnosis is an important opportunity for a good prognosis for lung cancer patients.
Lung cancer usually occurs in early stages in the form of lung nodules. The rapid development of imaging techniques has increased the detection rate of lung nodules. However, the pulmonary nodule disease is complex, and the clinical manifestation lacks specificity, so that the differential diagnosis of benign and malignant pulmonary nodules has certain difficulty. Generally, 40-50% of nodules with a diameter >3 cm are malignant, 25% -30% of nodules with a diameter between 0.5-1 cm are malignant, and only 1% of nodules with a diameter <0.5 cm are malignant. Studies have shown that a significant proportion of lung nodule patients exhibit post-operative pathology as benign disease. Surgical resection of such benign lung nodules can result in over-treatment and increase pain for this portion of the patient. Therefore, how to accurately diagnose lung cancer patients carrying malignant nodules from patients carrying different lung nodules is clinically significant.
At present, early diagnosis methods of lung cancer include breast imaging, bronchoscopy technology, sputum cast-off cytology detection and the like, but the detection effects of the methods are not ideal. The sensitivity of sputum examination to central lung cancer is about 50%, while it is less than 20% for peripheral lung cancer. The bronchoscopy technique has a diagnosis rate of about 90% for central lung cancer, but less than 30% for peripheral lung cancer, especially for precancerous lesions. Chest imaging examination methods include X-ray chest radiography (CxR), low dose helical CT (LDCT), and PET-CT. CxR the misdiagnosis rate is as high as 50-90%, and the specificity of detecting the nodules in the lung by LDCT and PET-CT is poor. It is reported that the detection of LDCT technology widely used in early screening and diagnosis of lung cancer can generate over 21% of false positives. Therefore, it is difficult to accurately diagnose early lung cancer by using imaging method alone, and there is no effective biomarker combined with imaging technology (LDCT) to improve the specificity of early lung cancer diagnosis.
Extracellular Vesicles (EVs) are vesicles produced by different cells and released to the outside of the cells, and can be classified into various types. Among them, small extracellular vesicles, also known as exosomes, are membrane vesicles of about 30-150 nm in diameter. It is derived from vesicles of late endocytosis bodies (MVB), which are formed by the inward recession of endocytosis vesicle membranes of cells to form a multivesicular body containing a plurality of vesicles, and the multivesicular body is fused with cell membranes and released into extracellular matrix. The extracellular vesicle-carried contents include proteins, lipids, mRNA, rRNA, miRNA, etc. Cells secrete extracellular vesicles under both normal and pathological conditions, which can participate in the transfer of information between cells. The content of the extracellular vesicles can represent certain physiological and pathological conditions. Among various diseases (e.g., malignant tumor, immune disease, etc.), extracellular vesicles (exosomes) free in peripheral blood have been widely noticed and studied as an important form of liquid biopsy.
Micro nucleic acids (microRNAs, miRNAs) are a class of non-coding small-molecule RNAs with the length of 19-25 nucleotides discovered in recent years. It mainly degrades the target gene mRNA or inhibits its translation through the complete or incomplete pairing with the target gene 3' UTR, thereby participating in the regulation of individual development, apoptosis, proliferation and differentiation and other life activities, and playing a role similar to oncogene or cancer suppressor gene in the process of generating and developing tumors. The expression profile of miRNA has obvious tissue specificity and specific expression pattern in different tumors. These characteristics make mirnas possible to become new biological markers and therapeutic targets for tumor diagnosis. Recently, there are studies suggesting that some differences exist between blood extracellular vesicle mirnas in healthy people and lung cancer patients, lung cancer adenocarcinoma and lung squamous carcinoma patients. However, the blood extracellular vesicle miRNA is rarely studied as a biomarker in identifying good and malignant lung nodules, and no biomarker capable of improving the specificity of early diagnosis of lung cancer is found and verified.
According to the invention, based on the research, peripheral blood of a lung nodule patient detected by clinical LDCT is taken as a research sample, peripheral blood extracellular vesicles are extracted, the expression of peripheral blood extracellular vesicle miRNA of a benign patient and a malignant patient is further detected by small RNA sequencing technology, and a high-specificity peripheral blood extracellular vesicle miRNA biomarker diagnosis model for early diagnosis of lung cancer is found and verified in two independent groups.
Disclosure of Invention
The invention aims to provide a peripheral blood extracellular vesicle miRNA marker for early diagnosis of lung cancer, based on a large amount of sample verification, 5 specific diagnosis markers suitable for early diagnosis of lung cancer are determined, and compared with other reported miRNA markers, the sample provided by the invention is based on clinical LDCT to detect blood of a lung nodule patient as a research sample, so that the sample has higher specificity; the 5 miRNAs are provided for the first time as early diagnosis markers of lung cancer, and are more reliable compared with other miRNA molecular markers. The method has important significance for further identifying the false positive patients from the lung nodule patients detected by LDCT.
The technical scheme adopted by the invention for solving the technical problems is as follows:
a peripheral blood extracellular vesicle miRNA marker for early diagnosis of lung cancer comprises hsa-let-7b-3p, hsa-miR-101-3p and hsa-miR-3168.
Further, the miRNA biomarker also comprises one or more miRNAs selected from hsa-miR-125b-5p, hsa-miR-150-5p, hsa-let-7d-3p, hsa-miR-503-5p, hsa-miR-16-5p, hsa-miR-17-5p, hsa-miR-19b-3p, hsa-miR-532-5p and hsa-miR-98-5 p.
Preferably, the miRNA biomarker can be a combination of hsa-let-7b-3p, hsa-miR-125b-5p, hsa-miR-150-5p, hsa-miR-101-3p and hsa-miR-3168.
Preferably, the miRNA biomarker can be a combination of hsa-let-7b-3p, hsa-miR-125b-5p, hsa-miR-150-5p, hsa-miR-101-3p, hsa-miR-3168, hsa-let-7d-3p, hsa-miR-19b-3p and hsa-miR-503-5 p.
Preferably, the miRNA biomarker can also be a combination of hsa-let-7b-3p, hsa-miR-125b-5p, hsa-miR-150-5p, hsa-miR-101-3p, hsa-miR-3168, hsa-miR-16-5p, hsa-miR-17-5p, hsa-miR-19b-3p, hsa-miR-532-5p and hsa-miR-98-5 p.
Further, the expression values of the peripheral blood extracellular vesicle miRNA markers in the peripheral blood extracellular vesicles of patients diagnosed with lung cancer are statistically significantly different from the expression values in the control samples.
Further, the control sample is a benign lung nodule patient, not the general normal population.
A kit for early diagnosis of lung cancer, which comprises the detection reagent of the peripheral blood extracellular vesicle miRNA marker.
Use of a peripheral blood extracellular vesicle miRNA marker for the preparation of a diagnostic agent for predicting the early stage lung cancer of a subject by a method comprising:
-collecting a sample of peripheral blood extracellular vesicles, detecting the presence of miRNA in peripheral blood extracellular vesicles obtained from the subject;
-measuring the expression level of said miRNA biomarker in said peripheral blood extracellular vesicle sample;
-using a risk scoring model based on previously measured expression levels of mirnas to predict the likelihood of the subject having early stage lung cancer.
Further, the expression level of the miRNA is obtained by quantification based on the result of the small RNA sequencing detection technology.
Further, the risk scoring model includes a mathematical model formulated as follows:
wherein Risk-Score is a lung nodule benign and malignant Risk prediction value, Gi represents an expression value of an ith miRNA, β i represents a Risk Score parameter value of the ith miRNA, i = 1, 2 … n, n is the total number of lung nodule benign and malignant biomarkers, and α represents a model correction parameter.
Drawings
FIG. 1 shows the results of electron microscopy of extracellular vesicles.
FIG. 2 protein expression characteristic of extracellular vesicles.
FIG. 3. training queue ROC curve.
FIG. 4 is a validation queue ROC curve.
FIG. 5. construction of a risk scoring model with 3 miRNAs as biomarkers, ROC curve analysis in training trains.
FIG. 6. construction of a risk scoring model with 8 miRNAs as markers, ROC curve analysis in training trains.
FIG. 7. construction of a risk scoring model using 10 miRNAs as markers, ROC curve analysis in training trains.
Detailed Description
All of the miRNAs involved in the present invention are mature miRNAs annotated in the miRBase database (http:// www.mirbase.org /).
Screening process of peripheral blood extracellular vesicle miRNA marker
(1) Research cohort and clinical information
The study included a total of 109 patients with LDCT as lung nodules from a total of two study cohorts, and plasma samples were taken from the patients prior to surgery. Each patient in the group gives an accurate diagnosis after surgery based on the results of the pathological examination. The training cohort was enrolled into 47 patients with a cohort time of 2018 months 9 to 2018 months 10, including 17 patients with benign lung nodules and 30 patients with malignant lung nodules (table 1). The validation cohort was 62 patients with a cohort time of 4 months to 5 months 2019, including 24 patients with benign lung nodules and 38 patients with malignant lung nodules (table 1). The benign specimen is classified into granuloma, inflammatory nodule, hamartoma, etc. Table 1 shows clinical information of patients, including sex, age, smoking history, and pathological diagnosis results. The analysis result shows that the sex, the age and the smoking history of the two groups of patients have no significant difference with the good and the malignant patient proportion.
TABLE 1.109 patient clinical information
(2) Extraction and characterization of peripheral blood extracellular vesicles
Collecting peripheral blood and extracting extracellular vesicles
Peripheral blood samples were collected in anticoagulated vacuum blood collection tubes (REF367863, BD, USA) prior to surgery or drug treatment and delivered at 4 ℃ within one hour after collection. And (3) centrifuging at 4 ℃ for 10min at 1600g after receiving the blood sample, judging the hemolysis grade of the blood sample after centrifuging, transferring the supernatant into a 1.5ml centrifuge tube for centrifuging 16000g again, centrifuging at 4 ℃ for 15min, subpackaging the supernatant into 1ml per tube, and storing in a refrigerator at minus 80 ℃. The plasma sample is taken out and placed in a metal bath at 37 ℃ for incubation until the plasma sample is completely melted, 12000g is centrifuged at 4 ℃ for 10min, 500 mul of supernatant is transferred to 0.45 mul of tube filter (Costar, CLS8163-100EA, Corning, USA), 12000g is centrifuged at 4 ℃ for 5min, and filtrate is transferred to 0.22 mul of tube filter (Costar, CLS8161-100EA, USA) for 12000g and centrifuged at 4 ℃ for 5 min. The filtrate was transferred to a 1.5ml centrifuge tube and 1/4 volumes of extracellular vesicle precipitant (cat # REX015S, 3D Med, Shanghai, China) were added, after mixing, incubation was carried out at 4 ℃ for 30min, 4700g was carried out, centrifugation was carried out at 4 ℃ for 30min, and 200. mu.l of PBS (phosphate buffersaline) was added to discard the supernatant to suspend extracellular vesicle precipitation.
② the characteristics of the extracellular vesicles of peripheral blood
In order to detect the characteristics of the extracellular vesicles of the peripheral blood, the method adopts a scanning electron microscope to detect the morphology of the extracellular vesicles, and further utilizes an immune hybridization experiment to detect the expression level of characteristic proteins of the extracellular vesicles. And (3) identifying the morphological characteristics of the extracellular vesicles: after precipitation, it was resuspended in PBS, then fixed with 5% glutaraldehyde, washed with PBS for 5min, then fixed with 1% osmic acid, then dehydrated with ethanol of various concentration gradients of 40%, 60%, 80%, 96-98%, and dried for 24 hours at room temperature after plating a silicon metal film on the sample surface for analysis by scanning electron microscope (SU8020, High-Technologies, Japan). The detection result shows that the extracellular vesicle has a typical horseshoe-shaped form (see figure 1).
Detecting extracellular vesicle characteristic protein: the extracellular vesicles were precipitated with extracellular vesicle precipitant (cat # REX015S, 3DMEd, Shanghai, China) and then lysed with RIPA lysate (P0013B, Beyotime, Shanghai, China) on ice for 30 min. Electrophoresis was performed at 4% -20% SDS-PAGE gel (#4561095, Bio-Rad, USA) for about 1h at constant pressure, followed by constant current transformation on PVDF membrane (Millipore) for 45min and blocking with 5% skim milk overnight. First-order information: TSG101 (1:1000diluted, ab125011, Abcam, England), CD63 (1:1000diluted, ab216130, Abcam, England), CD9(1:1000 diluted, ab92726, Abcam, England), Alix (1:1000diluted, 2171, Cell Signaling Technology, Danvers, MA, USA) and Syntenin (1:1000diluted, ab19903, Abcam, England) antibodies were incubated for 2h at room temperature, TBST was washed for 10min four times, rabbit secondary antibody (A0208, Biyun day) or mouse secondary antibody (A0216, Biyun day) was incubated for 1h at room temperature, TBST was washed for 10min four times, and developed with chemiluminescence system (Tanon-5200Multi, Shanghai, China). The detection result shows that the extracellular vesicle characteristic proteins TSG101, CD63, CD9, Alix and Syntenin are all expressed in the representative sample extracted in the patent (see figure 2).
(3) Extraction and expression quantity of extracellular vesicle miRNA
Extracting miRNA from extracellular vesicle of peripheral blood
MiRNeasy Serum/Plasma Kit (217184, QIAGEN, Shanghai, China) is selected for separating the miRNA of the extracellular vesicle of the peripheral blood, and the specific operation flow is according to the product instruction. miRNA quantification and fragment distribution detection are carried out by using an Agilent 2100 analyzer matched with a corresponding chip (5067-.
② expression detection of peripheral blood extracellular vesicles
This patent adopts small RNA sequencing to detect the expression level of peripheral blood extracellular vesicle miRNA. The Library was constructed using NEBNext, Multiplex SmallRNA Library Prep Set for Illumina (E7300L, NEB, USA) kit, and the specific procedures were according to the product instructions. The loading amount of miRNA in each plasma sample is 100ng, the total volume is not more than 6 mu l, and the miRNA is connected with a 3 'joint, hybridized reverse transcription primers, connected with a 5' joint, reverse transcription and added with Illumina indexprimers for PCR amplification for 18 cycles. The PCR product was purified using a NucleoSpin Gel and PCR Clean-up (740609.50, QIAGEN, Shanghai, China) kit, and DNA was eluted with 30. mu.l of enzyme-free water. DNA quantification and fragment distribution detection were performed using LabChip GX Touch HT nucleic acid analyzer and its associated chip (CLS138948, PerkinElmer, USA) and reagents (CLS760672, PerkinElmer, USA). Typically 20-25 libraries were sequenced at equimolar ratios mixed melane, using Illumina HiSeq PE150 analyzer.
(4) Sequencing data analysis process
Based on small RNA sequencing detection technology, the expression quantity of miRNA in the patient peripheral blood extracellular vesicles is obtained. The analysis flow of the sequencing data is as follows:
comparing sequencing data. After removing the sequencing joint of the small RNA sequencing data, BWA software (version: 0.7.12-r1039) is used to align the sequencing data to the human reference genome hg19 (genome download link: http:// hgdownload. soe. ucsc. edu/goldenPath/hg19/bigZips /), and the number of reads aligned to miRNA is counted.
② annotation of miRNA. Mirnas were annotated using the Gencodev25 and miRBasev21 databases, and the annotated mature mirnas were retained for subsequent analysis.
Filtering miRNA. For the training cohort, keeping mature miRNAs with the length less than or equal to 30nt and at least covering 2 reads in each sample in the training cohort data for subsequent analysis; for validation cohorts, mirnas screened by the training cohort and covering at least 2 reads of mature mirnas per sample in validation cohort data were retained for subsequent analysis.
And normalizing the miRNA expression quantity. And respectively carrying out miRNA expression quantity standardization processing on the training queue sample and the verification queue sample by using an M value weighted mean of M-values (TMM) and a limma-voom method in a limma analysis packet in the R language.
(5) Discovery of biomarkers
Based on the expression quantity of miRNAs in the training queue, grouping samples according to pathological detection results, and using a statistical method to discover miRNA which can be used for distinguishing benign and malignant lung nodules as biomarkers. The process is as follows:
training the queue grouping. According to the lung nodule pathological detection result, the patients in the training queue are divided into two groups, namely a benign lung nodule group and a malignant lung nodule group.
② candidate biomarkers. And analyzing the miRNAs with expression quantities different in two groups of benign and malignant by using a fitting linear model (limma-voom) method in an R language limma analysis package, wherein the expression quantities are more than 50 reads (CPM, Counts Per Million) mapped and read Per Million, and the miRNAs with the variation quantity of more than 1.5 times between two groups and the test result P of less than or equal to 0.01 are used as candidate biomarkers, and the candidate biomarkers comprise hsa-let-7b-3P, hsa-miR-125b-5P, hsa-miR-150-5P, hsa-miR-101-3P, hsa-miR-3168 and has-miR-144-3P, and the total number of 6 miRNAs are used for subsequent analysis.
③ biomarkers. In The training queue, The contribution degree of 6 candidate biomarkers to The prediction of The benign and malignant lung nodules is calculated by using a least absolute contraction and selection operator (LASSO) model, and The candidate biomarker has-miR-144-3p with The contribution degree of 0 is removed. Finally, 5 miRNAs are selected as biomarkers including hsa-let-7b-3p, hsa-miR-125b-5p, hsa-miR-150-5p, hsa-miR-101-3p and hsa-miR-3168, and are used for constructing a lung nodule benign and malignant risk score model in a follow-up manner.
Example 1: establishment and verification of risk scoring model with 5 miRNAs as biomarkers
(1) Lung nodule benign and malignant risk scoring model
And (3) constructing a benign and malignant risk scoring model of the pulmonary nodules by using the miRNA expression data of the training queue and 5 biomarkers as variables and combining pathological detection results. The model consists of three parts of parameters, a model formula and a reference value. The process is as follows:
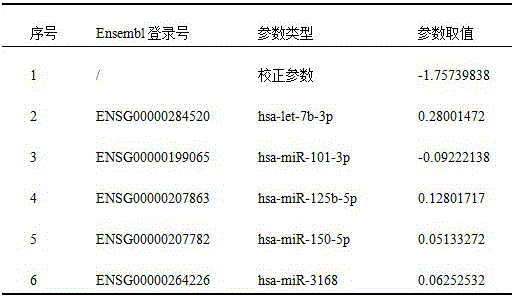
model parameters. In the training cohort, model calibration parameters and model coefficients for the biomarkers were obtained using 100 replicates of the 10-fold cross-validation method with 5 biomarkers as variables (table 2).
TABLE 2 parameters of risk scoring model constructed with 5 miRNAs as biomarkers
And ② a risk scoring model. The risk scoring model formula is as follows:
wherein Risk-Score is a lung nodule benign and malignant Risk prediction value, Gi represents an expression value of an ith miRNA, β i represents a Risk Score model parameter of the ith miRNA, i = 1, 2 … n, n is the total number of lung nodule benign and malignant biomarkers, and α represents a model correction value. Using the risk scoring model and the amount of expression of the biomarkers for each sample, a risk value for each sample can be obtained.
③ value of reference. When the risk value is less than or equal to the reference value, the sample is predicted to be a benign lung nodule; otherwise, a malignant lung nodule is predicted. And drawing a receiver operating characteristic curve (ROC curve) of the training queue according to the risk value and the pathological detection result of each patient in the training queue. In clinical application, the high-specificity risk assessment model can be used for assisting in reducing the false positive rate of LDCT detection, reducing the operation open rate of clinicians and improving the specificity of early lung cancer diagnosis. Therefore, the present model focuses on the high specificity of the model. The reference value is calculated according to the condition that the specificity value is more than 0.9 and the sensitivity value is more than 0.5 in the ROC curve. In this example, the reference value is 0.79.
And fourthly, evaluating the efficiency of the model. The training cohort samples were divided into low risk groups (i.e. predicted to be benign nodules) and high risk groups (i.e. predicted to be malignant nodules) based on the reference value of 0.79. And evaluating the prediction efficiency of the model by taking the pathological detection result as a true value. The model prediction efficiency evaluation method comprises the following steps of Area Under the ROC Curve (AUC, Area Under Curve, Value range 0-1), Positive Predictive Value (PPV, Value range 0-1), specificity (Value range 0-1) and sensitivity (Value range 0-1), wherein the higher the Value is, the better the effect is. 0.920 (FIG. 3), 93.8%, 94.1% and 50.0%, respectively (Table 3). The results show that: in the training queue, the risk prediction model has higher AUC, PPV and specificity, and the model has better prediction efficiency on benign pulmonary nodules.
(2) Verification of prediction efficiency of risk scoring model
And in the verification queue, verifying the efficiency of predicting the benign and malignant pulmonary nodules of the model according to the risk scoring model and the reference value determined in the training queue. The process is as follows:
(ii) a risk value. In the validation queue, a risk value is calculated for each sample.
And secondly, verifying the efficiency of the model. Patients in the validation cohort were divided into low risk groups and high risk groups (cohort training) based on a reference value of 0.79. With the pathological test results as the true value, ROC curves were plotted (fig. 4) and the model predictive potency, including AUC, PPV, specificity and sensitivity, were evaluated as 0.763, 91.3%, 91.7% and 55.3%, respectively (table 3). The results show that: in the validation cohort, this risk prediction model also has a higher AUC, PPV and specificity, i.e. the model prediction performance is validated in the validation cohort.
TABLE 3 evaluation of potency of five molecular marker models
Example 2: development of risk scoring model research by taking 3 miRNAs as biomarkers
The risk score model development research shows that when the biomarker takes 5 miRNAs (hsa-let-7b-3p, hsa-miR-125b-5p, hsa-miR-150-5p, hsa-miR-101-3p and hsa-miR-3168) as the core, the number of the miRNAs is reduced or increased, and higher prediction efficiency can be obtained.
By using 3 miRNAs as markers and hsa-let-7b-3p, hsa-miR-101-3p and hsa-miR-3168 as biomarkers, a risk scoring model (shown in the table 4) is constructed in the same way, and by using 0.77 as a reference value, higher AUC (0.914) and specificity (94.1%) can be obtained (shown in figure 5).
TABLE 4 parameters of risk scoring model constructed with 3 miRNAs as markers
Example 3 expansion of Risk Scoring model Studies-8 miRNAs as markers
Based on 5 core biomarkers, hsa-let-7d-3p, hsa-miR-19b-3p and hsa-miR-503-5p are newly added to be 8-variable biomarkers, a risk score model (Table 5) is constructed, and a reference value of 0.79 is used, so that higher AUC (0.920) and specificity (94.1%) can be obtained (FIG. 6).
TABLE 5 parameters of risk scoring model constructed with 8 miRNAs as markers
Example 4 expansion of Risk scoring model Studies-10 miRNAs as markers
Based on 5 core biomarkers, hsa-miR-16-5p, hsa-miR-17-5p, hsa-miR-19b-3p, hsa-miR-532-5p and hsa-miR-98-5p are newly added as 10-variable biomarkers, a risk score model (Table 6) is constructed, and higher AUC (0.961) and specificity (94.1%) can be obtained by taking 0.68 as a reference value (figure 7).
TABLE 6 parameters of risk scoring model constructed with 10 miRNAs as markers
According to the invention, a model can be established by using fewer miRNA locus combinations for risk evaluation, and more miRNA locus combinations can be used for risk evaluation; when the number of miRNA sites used is increased, benign and malignant lung nodules can be detected and distinguished from each other from more dimensions, and therefore a better detection result is obtained. Based on the miRNA locus or locus combination of the present invention, one skilled in the art can establish other risk assessment models and select corresponding parameters, and is not limited to the risk scoring models and parameter selections used in the embodiments of the present invention.
Claims (7)
1. A detection reagent of a peripheral blood extracellular vesicle miRNA biomarker for early diagnosis of lung cancer is characterized in that the peripheral blood extracellular vesicle miRNA biomarker for targeted detection by the detection reagent is a combination of hsa-let-7b-3p, hsa-miR-125b-5p, hsa-miR-150-5p, hsa-miR-101-3p and hsa-miR-3168.
2. The detection reagent of claim 1, wherein the expression value of the miRNA marker of the peripheral blood extracellular vesicle in the peripheral blood extracellular vesicle of the lung cancer patient is statistically significantly different from the expression value in the control sample.
3. The detection reagent of claim 2, wherein the control sample is a benign lung nodule patient.
4. A kit for early diagnosis of lung cancer, comprising: the kit comprises a detection reagent of the peripheral blood extracellular vesicle miRNA marker of claim 1.
5. Use of the detection reagent for the peripheral blood extracellular vesicle miRNA biomarker for early diagnosis of lung cancer according to claim 1 in the preparation of a diagnostic reagent for predicting early lung cancer by the method comprising:
(1) aiming at a subject with lung nodules detected by LDCT in clinic, collecting a peripheral blood sample, and detecting the existence of miRNA in the extracellular vesicles of peripheral blood;
(2) measuring the expression level of the miRNA biomarker of claim 1 in the peripheral blood extracellular vesicle sample;
(3) a risk scoring model based on previously measured expression levels of mirnas is used to predict the likelihood of LDCT detecting a malignant nodule carried by a subject with existing lung nodules.
6. The use according to claim 5, wherein the expression level of the miRNA is quantified based on small RNAseq detection results.
7. The use of claim 6, wherein the risk scoring model comprises a mathematical model formulated as:
wherein Risk-Score is a lung nodule benign and malignant Risk prediction value, Gi represents an expression value of an ith miRNA, β i represents a Risk Score parameter value of the ith miRNA, i = 1, 2 … n, n is the total number of lung nodule benign and malignant biomarkers, and α represents a model correction parameter.
Priority Applications (1)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| CN202010329484.XA CN111218513B (en) | 2020-04-24 | 2020-04-24 | Peripheral blood extracellular vesicle microRNA biomarker for early diagnosis of lung cancer and application thereof |
Applications Claiming Priority (1)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| CN202010329484.XA CN111218513B (en) | 2020-04-24 | 2020-04-24 | Peripheral blood extracellular vesicle microRNA biomarker for early diagnosis of lung cancer and application thereof |
Publications (2)
| Publication Number | Publication Date |
|---|---|
| CN111218513A CN111218513A (en) | 2020-06-02 |
| CN111218513B true CN111218513B (en) | 2020-08-14 |
Family
ID=70808074
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| CN202010329484.XA Active CN111218513B (en) | 2020-04-24 | 2020-04-24 | Peripheral blood extracellular vesicle microRNA biomarker for early diagnosis of lung cancer and application thereof |
Country Status (1)
| Country | Link |
|---|---|
| CN (1) | CN111218513B (en) |
Families Citing this family (10)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN111808950B (en) * | 2020-06-02 | 2023-11-14 | 中南大学湘雅医院 | A kind of miRNA marker for papillary thyroid cancer and its application |
| CN111676291B (en) * | 2020-07-14 | 2021-04-13 | 徐州医科大学 | A miRNA marker for risk assessment of lung cancer |
| CN111996260A (en) * | 2020-09-17 | 2020-11-27 | 上海思路迪医学检验所有限公司 | Extracellular vesicle microRNA biomarker for early diagnosis of liver cancer and application thereof |
| CN112301130B (en) * | 2020-11-12 | 2021-11-30 | 苏州京脉生物科技有限公司 | Marker, kit and method for early detection of lung cancer |
| CN112899359B (en) * | 2021-01-27 | 2023-06-23 | 广州市基准医疗有限责任公司 | Methylation marker for benign and malignant lung nodule detection or combination and application thereof |
| CN114369652B (en) * | 2021-11-19 | 2024-01-19 | 中山大学 | Application of exosome miR-17-5p in preparation of detection product for severe pneumonia |
| CN117448320A (en) * | 2022-07-25 | 2024-01-26 | 上海思路迪生物医学科技有限公司 | PCR internal reference, preparation method and application of blood small extracellular vesicle miRNA |
| CN117133467B (en) * | 2023-10-27 | 2023-12-22 | 上海晟燃生物科技有限公司 | System for early screening, auxiliary diagnosis and/or prognosis monitoring of lung cancer |
| CN119307612B (en) * | 2024-10-05 | 2025-08-19 | 江西省肿瘤医院(江西省第二人民医院、江西省癌症中心) | Application of blood extracellular vesicle miRNA combination as biomarker in early lung cancer diagnosis |
| CN120138157A (en) * | 2025-04-14 | 2025-06-13 | 杭州市第一人民医院(西湖大学附属杭州市第一人民医院) | Application and kit of mRNAs markers in early diagnosis of pulmonary signet ring cell carcinoma |
Citations (7)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN101400361A (en) * | 2006-01-05 | 2009-04-01 | 俄亥俄州立大学研究基金会 | micro-RNA based methods and compositions for diagnosis, prognosis and treatment of lung cancer |
| CN101918594A (en) * | 2007-11-30 | 2010-12-15 | 俄亥俄州立大学研究基金会 | MicroRNA expression profiling and targeting in peripheral blood in lung cancer |
| CN102892897A (en) * | 2009-12-24 | 2013-01-23 | 复旦大学 | Compositions and methods for microrna expression profiling of lung cancer |
| US9056135B2 (en) * | 2006-04-03 | 2015-06-16 | Trustees Of Dartmouth College | MicroRNA biomarkers for human breast and lung cancer |
| CN106520929A (en) * | 2016-10-17 | 2017-03-22 | 上海赛安生物医药科技有限公司 | Non-small cell lung cancer detection primer and probe and kit |
| CN106755544A (en) * | 2017-03-10 | 2017-05-31 | 大连医科大学附属第医院 | MiRNA marker and application in a kind of serum excretion body related to adenocarcinoma of lung early diagnosis |
| CN108841962A (en) * | 2018-08-01 | 2018-11-20 | 博奥生物集团有限公司 | A kind of non-small cell lung cancer detection kit and its application |
-
2020
- 2020-04-24 CN CN202010329484.XA patent/CN111218513B/en active Active
Patent Citations (7)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN101400361A (en) * | 2006-01-05 | 2009-04-01 | 俄亥俄州立大学研究基金会 | micro-RNA based methods and compositions for diagnosis, prognosis and treatment of lung cancer |
| US9056135B2 (en) * | 2006-04-03 | 2015-06-16 | Trustees Of Dartmouth College | MicroRNA biomarkers for human breast and lung cancer |
| CN101918594A (en) * | 2007-11-30 | 2010-12-15 | 俄亥俄州立大学研究基金会 | MicroRNA expression profiling and targeting in peripheral blood in lung cancer |
| CN102892897A (en) * | 2009-12-24 | 2013-01-23 | 复旦大学 | Compositions and methods for microrna expression profiling of lung cancer |
| CN106520929A (en) * | 2016-10-17 | 2017-03-22 | 上海赛安生物医药科技有限公司 | Non-small cell lung cancer detection primer and probe and kit |
| CN106755544A (en) * | 2017-03-10 | 2017-05-31 | 大连医科大学附属第医院 | MiRNA marker and application in a kind of serum excretion body related to adenocarcinoma of lung early diagnosis |
| CN108841962A (en) * | 2018-08-01 | 2018-11-20 | 博奥生物集团有限公司 | A kind of non-small cell lung cancer detection kit and its application |
Non-Patent Citations (5)
| Title |
|---|
| Diagnostic Value of MiR-125b as a Potential Biomarker for Stage I Lung Adenocarcinoma;Zeybec等;《Current Molecular Medicine》;20190404;第19卷(第3期);第216-227页 * |
| Exosomal miRNAs species in the blood of small cell and non small cell lung cancer patients;Valeriy Poroyko等;《Oncotarget》;20181231;第9卷(第28期);第19793-19806页 * |
| MicroRNAs as Liquid Biopsy Biomarkers for Early Detection in Lung Cancer;P.Reis等;《Journal of Thoracic Oncology》;20181030;第13卷(第10S期);第S785-S786页 * |
| Tissue and exosomal miRNA editing in Non-Small Cell Lung Cancer;Giovanni Nigita等;《Scientific Reports》;20180705;第1-8页 * |
| 肺癌相关非编码RNA的筛选及功能研究;张艳秋;《中国博士学位论文全文数据库医药卫生科技辑》;20180215(第2期);E072-30 * |
Also Published As
| Publication number | Publication date |
|---|---|
| CN111218513A (en) | 2020-06-02 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| CN111218513B (en) | Peripheral blood extracellular vesicle microRNA biomarker for early diagnosis of lung cancer and application thereof | |
| CN109777874B (en) | Plasma exosome miRNA marker suitable for diagnosis and prognosis of pancreatic ductal adenocarcinoma and application thereof | |
| CN105067822B (en) | Marker for diagnosing esophagus cancer | |
| CN112301130B (en) | Marker, kit and method for early detection of lung cancer | |
| CN111748629A (en) | Detection reagent of biomarker for early diagnosis of pancreatic cancer | |
| CN105473739A (en) | Biomarkers for colorectal cancer | |
| CN115803448A (en) | Micronucleus DNA from peripheral red blood cells and uses thereof | |
| CN111424093A (en) | Kit, device and method for lung cancer diagnosis | |
| CN106897579A (en) | New early tumor markers based on chromosomal variation index and its application | |
| CN118406765A (en) | MiRNA marker for detecting non-small cell lung cancer, detection kit and detection method | |
| CN117316278A (en) | Cancer noninvasive early screening method and system based on cfDNA fragment length distribution characteristics | |
| CN119307611A (en) | Blood extracellular vesicle miRNA combination and its application in early diagnosis of lung cancer | |
| CN111996260A (en) | Extracellular vesicle microRNA biomarker for early diagnosis of liver cancer and application thereof | |
| CN119491050B (en) | Application of blood small extracellular vesicle microRNA combination in lung cancer diagnosis and differential diagnosis | |
| WO2017156739A1 (en) | Isolated nucleic acid application thereof | |
| CN114150063A (en) | Urine miRNA marker for bladder cancer diagnosis, diagnostic reagent and kit | |
| CN119351556A (en) | A platelet marker and detection kit for detecting lung cancer | |
| CN116254332B (en) | A urine PCR kit for predicting minimal change disease in adults | |
| CN119351549A (en) | Small extracellular vesicle miRNA and its application in early detection of lung cancer | |
| CN107904310B (en) | Urine microRNA biomarker for colorectal cancer diagnosis, kit and application thereof | |
| CN109266750A (en) | Biomarker for nasopharyngeal carcinoma diagnosis and application | |
| CN117448320A (en) | PCR internal reference, preparation method and application of blood small extracellular vesicle miRNA | |
| CN118547069B (en) | A gastric cancer diagnostic marker based on extracellular vesicle miRNA and its application | |
| CN115948546B (en) | Exosome miRNA biomarker for breast cancer and application thereof | |
| CN109182520A (en) | A kind of cervical carcinoma and its precancerous lesion detection kit and its application |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| PB01 | Publication | ||
| PB01 | Publication | ||
| SE01 | Entry into force of request for substantive examination | ||
| SE01 | Entry into force of request for substantive examination | ||
| GR01 | Patent grant | ||
| GR01 | Patent grant |