CN110845862B - Preparation method of fluorescent dye - Google Patents
Preparation method of fluorescent dye Download PDFInfo
- Publication number
- CN110845862B CN110845862B CN201911185975.5A CN201911185975A CN110845862B CN 110845862 B CN110845862 B CN 110845862B CN 201911185975 A CN201911185975 A CN 201911185975A CN 110845862 B CN110845862 B CN 110845862B
- Authority
- CN
- China
- Prior art keywords
- fluorescent dye
- preparing
- bacteria
- structural formula
- reaction temperature
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Active
Links
- 239000007850 fluorescent dye Substances 0.000 title claims abstract description 24
- 238000002360 preparation method Methods 0.000 title claims abstract description 8
- 238000006243 chemical reaction Methods 0.000 claims description 26
- 239000000126 substance Substances 0.000 claims description 21
- 238000000034 method Methods 0.000 claims description 19
- 239000002994 raw material Substances 0.000 claims description 11
- QWXYZCJEXYQNEI-OSZHWHEXSA-N intermediate I Chemical compound COC(=O)[C@@]1(C=O)[C@H]2CC=[N+](C\C2=C\C)CCc2c1[nH]c1ccccc21 QWXYZCJEXYQNEI-OSZHWHEXSA-N 0.000 claims description 9
- -1 amine compound Chemical class 0.000 claims description 7
- 150000001450 anions Chemical group 0.000 claims description 7
- 238000010992 reflux Methods 0.000 claims description 7
- VEXZGXHMUGYJMC-UHFFFAOYSA-N Hydrochloric acid Chemical compound Cl VEXZGXHMUGYJMC-UHFFFAOYSA-N 0.000 claims description 6
- PXIPVTKHYLBLMZ-UHFFFAOYSA-N Sodium azide Chemical compound [Na+].[N-]=[N+]=[N-] PXIPVTKHYLBLMZ-UHFFFAOYSA-N 0.000 claims description 6
- LPXPTNMVRIOKMN-UHFFFAOYSA-M sodium nitrite Chemical compound [Na+].[O-]N=O LPXPTNMVRIOKMN-UHFFFAOYSA-M 0.000 claims description 6
- 238000003756 stirring Methods 0.000 claims description 6
- VEXZGXHMUGYJMC-UHFFFAOYSA-M Chloride anion Chemical compound [Cl-] VEXZGXHMUGYJMC-UHFFFAOYSA-M 0.000 claims description 4
- XMBWDFGMSWQBCA-UHFFFAOYSA-N hydrogen iodide Chemical compound I XMBWDFGMSWQBCA-UHFFFAOYSA-N 0.000 claims description 4
- 150000002497 iodine compounds Chemical class 0.000 claims description 4
- BMYNFMYTOJXKLE-UHFFFAOYSA-N 3-azaniumyl-2-hydroxypropanoate Chemical compound NCC(O)C(O)=O BMYNFMYTOJXKLE-UHFFFAOYSA-N 0.000 claims description 3
- CPNAVTYCORRLMH-UHFFFAOYSA-N 6-phenylphenanthridine-3,8-diamine Chemical compound C=1C(N)=CC=C(C2=CC=C(N)C=C22)C=1N=C2C1=CC=CC=C1 CPNAVTYCORRLMH-UHFFFAOYSA-N 0.000 claims description 3
- 125000000217 alkyl group Chemical group 0.000 claims description 3
- 235000010288 sodium nitrite Nutrition 0.000 claims description 3
- RIFGWPKJUGCATF-UHFFFAOYSA-N ethyl chloroformate Chemical compound CCOC(Cl)=O RIFGWPKJUGCATF-UHFFFAOYSA-N 0.000 claims description 2
- 238000002156 mixing Methods 0.000 claims description 2
- 239000002253 acid Substances 0.000 claims 2
- 241000894006 Bacteria Species 0.000 abstract description 45
- 238000001514 detection method Methods 0.000 abstract description 20
- 210000004027 cell Anatomy 0.000 description 31
- 239000000975 dye Substances 0.000 description 28
- 108020004414 DNA Proteins 0.000 description 26
- XEKOWRVHYACXOJ-UHFFFAOYSA-N Ethyl acetate Chemical compound CCOC(C)=O XEKOWRVHYACXOJ-UHFFFAOYSA-N 0.000 description 18
- 239000000047 product Substances 0.000 description 13
- OKKJLVBELUTLKV-UHFFFAOYSA-N Methanol Chemical compound OC OKKJLVBELUTLKV-UHFFFAOYSA-N 0.000 description 12
- WEVYAHXRMPXWCK-UHFFFAOYSA-N Acetonitrile Chemical compound CC#N WEVYAHXRMPXWCK-UHFFFAOYSA-N 0.000 description 9
- 210000000170 cell membrane Anatomy 0.000 description 8
- 238000011529 RT qPCR Methods 0.000 description 7
- XLYOFNOQVPJJNP-UHFFFAOYSA-N water Substances O XLYOFNOQVPJJNP-UHFFFAOYSA-N 0.000 description 7
- QTBSBXVTEAMEQO-UHFFFAOYSA-N Acetic acid Chemical compound CC(O)=O QTBSBXVTEAMEQO-UHFFFAOYSA-N 0.000 description 6
- YMWUJEATGCHHMB-UHFFFAOYSA-N Dichloromethane Chemical compound ClCCl YMWUJEATGCHHMB-UHFFFAOYSA-N 0.000 description 6
- 230000001580 bacterial effect Effects 0.000 description 6
- 239000002904 solvent Substances 0.000 description 6
- 239000012065 filter cake Substances 0.000 description 5
- 238000002372 labelling Methods 0.000 description 5
- 239000000243 solution Substances 0.000 description 5
- 241000191967 Staphylococcus aureus Species 0.000 description 4
- HEMHJVSKTPXQMS-UHFFFAOYSA-M Sodium hydroxide Chemical compound [OH-].[Na+] HEMHJVSKTPXQMS-UHFFFAOYSA-M 0.000 description 3
- PBCJIPOGFJYBJE-UHFFFAOYSA-N acetonitrile;hydrate Chemical compound O.CC#N PBCJIPOGFJYBJE-UHFFFAOYSA-N 0.000 description 3
- 239000000872 buffer Substances 0.000 description 3
- 238000001816 cooling Methods 0.000 description 3
- 230000007547 defect Effects 0.000 description 3
- 238000000605 extraction Methods 0.000 description 3
- 238000010438 heat treatment Methods 0.000 description 3
- 108020004999 messenger RNA Proteins 0.000 description 3
- 108020004707 nucleic acids Proteins 0.000 description 3
- 102000039446 nucleic acids Human genes 0.000 description 3
- 150000007523 nucleic acids Chemical class 0.000 description 3
- 230000001717 pathogenic effect Effects 0.000 description 3
- 108090000623 proteins and genes Proteins 0.000 description 3
- 239000007787 solid Substances 0.000 description 3
- 238000000967 suction filtration Methods 0.000 description 3
- 231100000419 toxicity Toxicity 0.000 description 3
- 230000001988 toxicity Effects 0.000 description 3
- 244000063299 Bacillus subtilis Species 0.000 description 2
- 235000014469 Bacillus subtilis Nutrition 0.000 description 2
- 102000053602 DNA Human genes 0.000 description 2
- 241000588724 Escherichia coli Species 0.000 description 2
- 241000186781 Listeria Species 0.000 description 2
- JUJWROOIHBZHMG-UHFFFAOYSA-N Pyridine Chemical compound C1=CC=NC=C1 JUJWROOIHBZHMG-UHFFFAOYSA-N 0.000 description 2
- 241000607142 Salmonella Species 0.000 description 2
- 229910052782 aluminium Inorganic materials 0.000 description 2
- 230000003321 amplification Effects 0.000 description 2
- 230000004071 biological effect Effects 0.000 description 2
- 239000007853 buffer solution Substances 0.000 description 2
- 210000002421 cell wall Anatomy 0.000 description 2
- 239000003153 chemical reaction reagent Substances 0.000 description 2
- 150000001875 compounds Chemical class 0.000 description 2
- 238000012136 culture method Methods 0.000 description 2
- 230000007812 deficiency Effects 0.000 description 2
- 239000003814 drug Substances 0.000 description 2
- 229940079593 drug Drugs 0.000 description 2
- 230000000694 effects Effects 0.000 description 2
- 238000001914 filtration Methods 0.000 description 2
- 238000001819 mass spectrum Methods 0.000 description 2
- 239000012528 membrane Substances 0.000 description 2
- 230000002503 metabolic effect Effects 0.000 description 2
- 238000003199 nucleic acid amplification method Methods 0.000 description 2
- 238000011002 quantification Methods 0.000 description 2
- 238000011160 research Methods 0.000 description 2
- 238000012360 testing method Methods 0.000 description 2
- AAAXMNYUNVCMCJ-UHFFFAOYSA-N 1,3-diiodopropane Chemical compound ICCCI AAAXMNYUNVCMCJ-UHFFFAOYSA-N 0.000 description 1
- QLIMAARBRDAYGQ-UHFFFAOYSA-N 1,6-diiodohexane Chemical compound ICCCCCCI QLIMAARBRDAYGQ-UHFFFAOYSA-N 0.000 description 1
- SKHUPKIELOSDON-UHFFFAOYSA-N 1-phenylphenanthridine Chemical compound C1=CC=CC=C1C1=CC=CC2=NC=C(C=CC=C3)C3=C12 SKHUPKIELOSDON-UHFFFAOYSA-N 0.000 description 1
- 241000193830 Bacillus <bacterium> Species 0.000 description 1
- ZXFDSSZCWQFXAT-UHFFFAOYSA-N Br.[N-]=[N+]=[N-] Chemical compound Br.[N-]=[N+]=[N-] ZXFDSSZCWQFXAT-UHFFFAOYSA-N 0.000 description 1
- 208000035473 Communicable disease Diseases 0.000 description 1
- 230000008836 DNA modification Effects 0.000 description 1
- 241000233866 Fungi Species 0.000 description 1
- 241000186779 Listeria monocytogenes Species 0.000 description 1
- 241000187479 Mycobacterium tuberculosis Species 0.000 description 1
- ZMXDDKWLCZADIW-UHFFFAOYSA-N N,N-Dimethylformamide Chemical compound CN(C)C=O ZMXDDKWLCZADIW-UHFFFAOYSA-N 0.000 description 1
- 238000012408 PCR amplification Methods 0.000 description 1
- 241000700605 Viruses Species 0.000 description 1
- 244000052616 bacterial pathogen Species 0.000 description 1
- 230000015556 catabolic process Effects 0.000 description 1
- 231100000135 cytotoxicity Toxicity 0.000 description 1
- 230000003013 cytotoxicity Effects 0.000 description 1
- 238000006731 degradation reaction Methods 0.000 description 1
- 230000014670 detection of bacterium Effects 0.000 description 1
- 230000006806 disease prevention Effects 0.000 description 1
- BFMYDTVEBKDAKJ-UHFFFAOYSA-L disodium;(2',7'-dibromo-3',6'-dioxido-3-oxospiro[2-benzofuran-1,9'-xanthene]-4'-yl)mercury;hydrate Chemical compound O.[Na+].[Na+].O1C(=O)C2=CC=CC=C2C21C1=CC(Br)=C([O-])C([Hg])=C1OC1=C2C=C(Br)C([O-])=C1 BFMYDTVEBKDAKJ-UHFFFAOYSA-L 0.000 description 1
- 238000001035 drying Methods 0.000 description 1
- 230000008030 elimination Effects 0.000 description 1
- 238000003379 elimination reaction Methods 0.000 description 1
- 230000002708 enhancing effect Effects 0.000 description 1
- 230000007613 environmental effect Effects 0.000 description 1
- ZMMJGEGLRURXTF-UHFFFAOYSA-N ethidium bromide Chemical compound [Br-].C12=CC(N)=CC=C2C2=CC=C(N)C=C2[N+](CC)=C1C1=CC=CC=C1 ZMMJGEGLRURXTF-UHFFFAOYSA-N 0.000 description 1
- 229960005542 ethidium bromide Drugs 0.000 description 1
- 230000005284 excitation Effects 0.000 description 1
- 238000002474 experimental method Methods 0.000 description 1
- 239000012467 final product Substances 0.000 description 1
- 235000013305 food Nutrition 0.000 description 1
- 239000012634 fragment Substances 0.000 description 1
- 238000004108 freeze drying Methods 0.000 description 1
- 230000014509 gene expression Effects 0.000 description 1
- 239000005457 ice water Substances 0.000 description 1
- 208000015181 infectious disease Diseases 0.000 description 1
- 230000005764 inhibitory process Effects 0.000 description 1
- 231100000518 lethal Toxicity 0.000 description 1
- 230000001665 lethal effect Effects 0.000 description 1
- 231100000053 low toxicity Toxicity 0.000 description 1
- 238000010907 mechanical stirring Methods 0.000 description 1
- QABLOFMHHSOFRJ-UHFFFAOYSA-N methyl 2-chloroacetate Chemical compound COC(=O)CCl QABLOFMHHSOFRJ-UHFFFAOYSA-N 0.000 description 1
- 239000000203 mixture Substances 0.000 description 1
- 238000006386 neutralization reaction Methods 0.000 description 1
- 229910052757 nitrogen Inorganic materials 0.000 description 1
- 125000004433 nitrogen atom Chemical group N* 0.000 description 1
- 244000052769 pathogen Species 0.000 description 1
- 230000007918 pathogenicity Effects 0.000 description 1
- 230000035699 permeability Effects 0.000 description 1
- 230000008569 process Effects 0.000 description 1
- ZDWVWKDAWBGPDN-UHFFFAOYSA-O propidium Chemical compound C12=CC(N)=CC=C2C2=CC=C(N)C=C2[N+](CCC[N+](C)(CC)CC)=C1C1=CC=CC=C1 ZDWVWKDAWBGPDN-UHFFFAOYSA-O 0.000 description 1
- 230000005180 public health Effects 0.000 description 1
- 238000000746 purification Methods 0.000 description 1
- UMJSCPRVCHMLSP-UHFFFAOYSA-N pyridine Natural products COC1=CC=CN=C1 UMJSCPRVCHMLSP-UHFFFAOYSA-N 0.000 description 1
- 239000011541 reaction mixture Substances 0.000 description 1
- 238000009877 rendering Methods 0.000 description 1
- 230000000241 respiratory effect Effects 0.000 description 1
- 230000000717 retained effect Effects 0.000 description 1
- 230000035945 sensitivity Effects 0.000 description 1
- 230000001954 sterilising effect Effects 0.000 description 1
- 238000004659 sterilization and disinfection Methods 0.000 description 1
- 125000001424 substituent group Chemical group 0.000 description 1
- 230000002194 synthesizing effect Effects 0.000 description 1
- 231100000331 toxic Toxicity 0.000 description 1
- 230000002588 toxic effect Effects 0.000 description 1
- 238000005406 washing Methods 0.000 description 1
Images
Classifications
-
- C—CHEMISTRY; METALLURGY
- C09—DYES; PAINTS; POLISHES; NATURAL RESINS; ADHESIVES; COMPOSITIONS NOT OTHERWISE PROVIDED FOR; APPLICATIONS OF MATERIALS NOT OTHERWISE PROVIDED FOR
- C09B—ORGANIC DYES OR CLOSELY-RELATED COMPOUNDS FOR PRODUCING DYES, e.g. PIGMENTS; MORDANTS; LAKES
- C09B57/00—Other synthetic dyes of known constitution
-
- C—CHEMISTRY; METALLURGY
- C09—DYES; PAINTS; POLISHES; NATURAL RESINS; ADHESIVES; COMPOSITIONS NOT OTHERWISE PROVIDED FOR; APPLICATIONS OF MATERIALS NOT OTHERWISE PROVIDED FOR
- C09K—MATERIALS FOR MISCELLANEOUS APPLICATIONS, NOT PROVIDED FOR ELSEWHERE
- C09K11/00—Luminescent, e.g. electroluminescent, chemiluminescent materials
- C09K11/06—Luminescent, e.g. electroluminescent, chemiluminescent materials containing organic luminescent materials
-
- C—CHEMISTRY; METALLURGY
- C09—DYES; PAINTS; POLISHES; NATURAL RESINS; ADHESIVES; COMPOSITIONS NOT OTHERWISE PROVIDED FOR; APPLICATIONS OF MATERIALS NOT OTHERWISE PROVIDED FOR
- C09K—MATERIALS FOR MISCELLANEOUS APPLICATIONS, NOT PROVIDED FOR ELSEWHERE
- C09K2211/00—Chemical nature of organic luminescent or tenebrescent compounds
- C09K2211/10—Non-macromolecular compounds
- C09K2211/1003—Carbocyclic compounds
- C09K2211/1007—Non-condensed systems
-
- C—CHEMISTRY; METALLURGY
- C09—DYES; PAINTS; POLISHES; NATURAL RESINS; ADHESIVES; COMPOSITIONS NOT OTHERWISE PROVIDED FOR; APPLICATIONS OF MATERIALS NOT OTHERWISE PROVIDED FOR
- C09K—MATERIALS FOR MISCELLANEOUS APPLICATIONS, NOT PROVIDED FOR ELSEWHERE
- C09K2211/00—Chemical nature of organic luminescent or tenebrescent compounds
- C09K2211/10—Non-macromolecular compounds
- C09K2211/1018—Heterocyclic compounds
- C09K2211/1025—Heterocyclic compounds characterised by ligands
- C09K2211/1029—Heterocyclic compounds characterised by ligands containing one nitrogen atom as the heteroatom
Landscapes
- Chemical & Material Sciences (AREA)
- Organic Chemistry (AREA)
- Engineering & Computer Science (AREA)
- Materials Engineering (AREA)
- Measuring Or Testing Involving Enzymes Or Micro-Organisms (AREA)
Abstract
The invention relates to a preparation method of a fluorescent dye, which can be applied to bacteria detection.
Description
The invention belongs to a fluorescent dye, a preparation method thereof and application thereof in bacterial detection, and relates to divisional application of invention application with application date of 2017, 12 and 29 and application number of 2017114799437, which is part of a preparation method of a product.
Technical Field
The invention belongs to the technical field of dye compounds, and particularly relates to a fluorescent dye, a preparation method thereof and application thereof in bacterial detection.
Background
Conventional bacterial culture methods are the most common methods of bacterial detection. However, some bacteria are in a special Viable state, i.e., in a Viable but non-culturable state (VBNC), characterized by biological activity and pathogenicity (i.e., invisible sources of infection), but do not grow and multiply on the medium when cultured under normal conditions, rendering conventional bacterial culture methods ineffective. The detection of bacteria in such a state is particularly important in the fields of food quarantine, water quality detection, disease prevention and control, and the like. For example, food-borne pathogenic bacteria often cause missed detection once entering the VBNC state, thereby causing serious public health crisis and even outbreak of infectious diseases. There are many methods for detecting bacteria in a viable, non-culturable state, one of which is a nucleic acid (DNA) molecule-based PCR detection method for VBNC bacteria. The method has great advantages, because the VBNC bacterial genome DNA is easy to extract, the PCR technology is very quick, sensitive and specific, and the operation is simple and convenient. However, in practical application, the method has the biggest defect that false positive interference is difficult to eliminate. Since the DNA in dead bacteria is still amplified if the dead bacteria and live bacteria cannot be distinguished during the extraction of genomic DNA, the extraction of DNA in dead bacteria will undoubtedly result in false positive results in the subsequent PCR.
The viable, non-culturable state of bacteria was proposed in 1982. Bacteria in the VBNC state no longer divide and proliferate but retain their activity, i.e., remain potentially pathogenic and have biological properties such as metabolic activity and gene expression. The live bacteria have the following characteristics: the cell membrane is complete, and the cell wall structure is changed; have metabolic activity (e.g., respiratory movement, uptake of biochemical substances, etc.) or are capable of synthesizing DNA, the presence of mRNA molecules, and VBNC-specific genes. Therefore, detection methods based on detection of nucleic acids are one of the important means for detecting VNBC bacteria. Among these methods, the detection techniques based on mRNA molecules have the characteristic of low false positives, but because mRNA molecules are low in abundance and unstable, sensitivity and specificity are low. The DNA detection-based method, namely the VBNC bacteria PCR detection method based on DNA molecules, has high specificity, is simple and saves time. However, since the DNA molecule is stable, even though the bacteria are lysed or even die, the DNA can be retained for a long time without degradation, the conventional PCR method cannot distinguish the dead and live states of the bacteria, and dead cells or free DNA can be detected as the target gene, but the target gene has lost pathogenic ability, so that the false positive rate of detection is very high, and therefore, a new detection method needs to be established to make up for the deficiency.
In order to make up for the deficiency of the PCR detection method of VBNC bacteria based on DNA molecules, a dye is applied to the detection method to eliminate possible false positive; such as ethidium bromide (EMA), propidium azide bromide (PMA), PMAxx dyes, PEMAX dyes, and the like. There have been many research papers on the above dyes. Research shows that aiming at heat lethal bacteria, if the temperature is not high enough, the loss of cell membranes is insufficient, then both EMA and PMA can not enter cells, and experiments prove that the bacteria can generate obvious membrane damage level at 60 ℃ or higher, and in this case, EMA penetrates the heat damaged cells more effectively than PMA; EMA has great defects compared with PMA, for example, not only does EMA have much greater permeability to living cell membranes than PMA, resulting in significant loss of genomic DNA of living cells; however, some studies have also found that the PMA-PCR method is not completely effective in removing dead cell signals; kralik et al reported that no more than 2 log of PCR signal could be obtained using membrane permeable cells of Mycobacteria; pan and Breedt also indicate that PMA-PCR does not always eliminate the signal of heat-killed Listeria monocytogenes; EMA-PCR also has similar problems, namely, the incomplete inhibition of the signal of dead cells; PMAxx developed by biotium. inc. is a new and improved feasible PCR dye that functions very similar to PMA, has greater activity and the ability to distinguish between live and dead bacteria, but is also more toxic to cells. This dye is more effective in eliminating PCR amplification of dead cell DNA, but many times it is required to be used with an enhancement buffer provided by Biotium inc. for example when amplifying sequences from gram negative bacteria by PCR, samples pretreated with dye and enhancement buffer show reduced signal from dead cells, while the signal from live cells is unchanged, but such enhancement buffer is necessary for samples using mild methods such as low heat sterilization of bacteria.
Disclosure of Invention
Aiming at the problem that the extraction of DNA in the non-viable bacteria generates false positive on the result of the subsequent PCR, the invention provides a novel compound which can mark dead bacteria in the mixed population of the live bacteria and the dead bacteria, and the DNA in the marked dead bacteria cannot be amplified any more even if being extracted, thereby avoiding the false positive problem caused by the DNA in the dead bacteria; compared with the existing dyes, the reagent is more sensitive, has low metering and far lower toxicity to bacteria than the existing several reagents, is quicker and more economical than the existing products in the aspect of quick detection, has wide application range, and is very effective in PMA-qPCR tests of various bacteria, protozoa, viruses and fungi (including pathogens and environmental strains).
The invention adopts the following technical scheme:
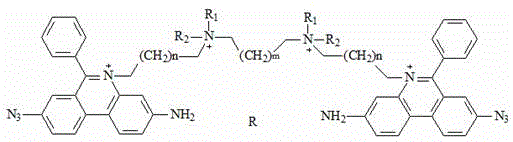
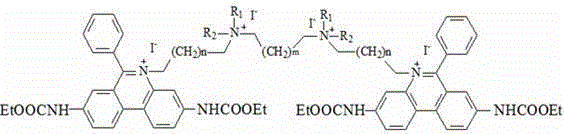
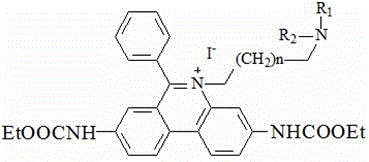
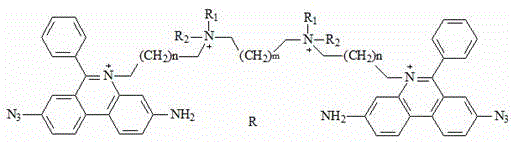
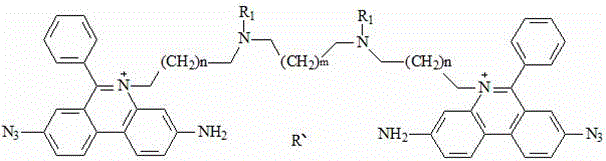
a fluorescent dye having the chemical formula:
wherein R is1、R2Independently selected from linear alkyl of C1-C3 or H; n is 1-4; m is 1-4; r is an anion; r 'is an anion, preferably, R' is two chloride ions; r is four chloride ions; the invention can only have one substituent group on the nitrogen atom of two phenyl phenanthridine connectors, and the anion can be a multi-valence anion or a plurality of single-valence anions, such as 2Cl-Or 4Cl-。
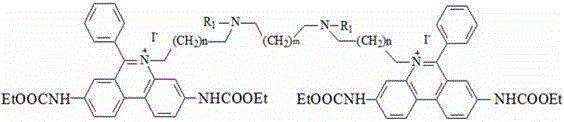
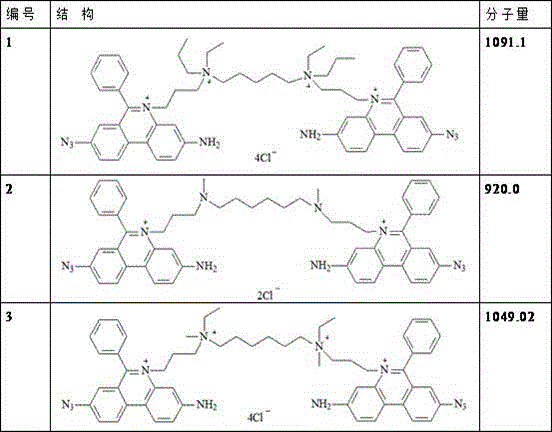
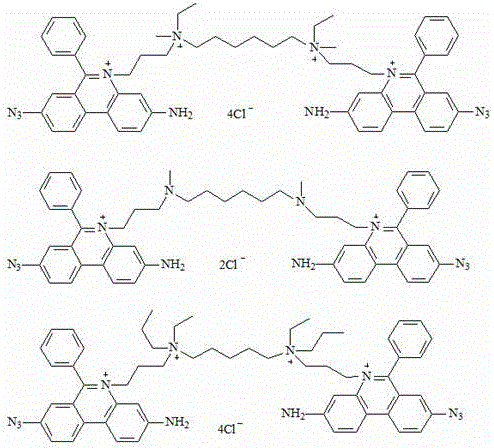
Preferably, the chemical structural formula of the fluorescent dye is as follows:
the dyes of the invention are all photoreactive dyes which are combined with dsDNA with high affinity, the fluorescence of the dyes is very weak, but the fluorescence signals are very strong after being combined with nucleic acid, the dyes are impermeable to cell membranes and can only enter damaged cell membranes to be combined with DNA through covalent bonds so as to cause permanent DNA modification, and the modified DNA can not be amplified any more, so the dyes can well make up the defects of the conventional detection method when being combined with a PCR method. Or that these dyes can selectively modify dead cell "exposed" DNA (i.e., cells with damaged cell walls and cell membranes) in a mixed population of cells and dead cells without destroying the integrity of living cells, which feature allows the dyes to selectively detect living pathogenic cells by quantitative real-time PCR in the presence of dead cells that have been modified by the dyes and thus cannot be amplified, thereby solving the prior art problem of false positives due to dead cells.
The invention also discloses a preparation method of the fluorescent dye, which comprises the following steps:
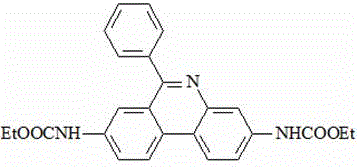
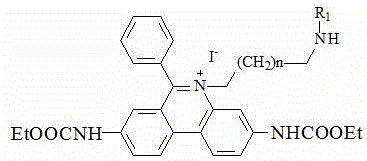
(1) preparing an intermediate I by using 3, 8-diamino-6-phenylphenanthridine and ethyl chloroformate as raw materials;
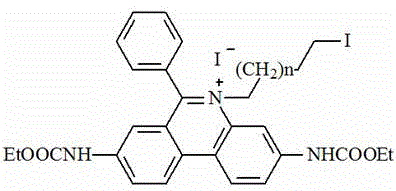
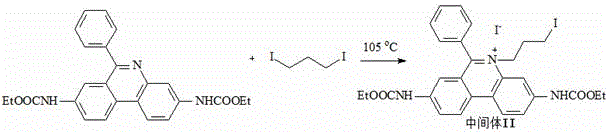
(2) preparing an intermediate II by taking the intermediate I and iodide as raw materials; the chemical structural formula of the iodide is I (CH)2)nI;
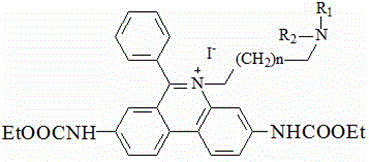
(3) Preparing an intermediate III by taking the intermediate II and an amine compound as raw materials; the chemical structure of the amine compound is R1NHR2Or R1NH2;
(4) Preparing an intermediate IV by taking the intermediate III and an iodine compound as raw materials; the chemical structural formula of the iodine compound is I (CH)2)mI;
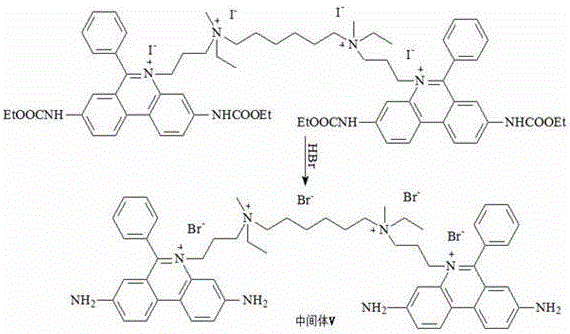
(5) Preparing an intermediate V by taking the intermediate IV and hydrobromic acid as raw materials;
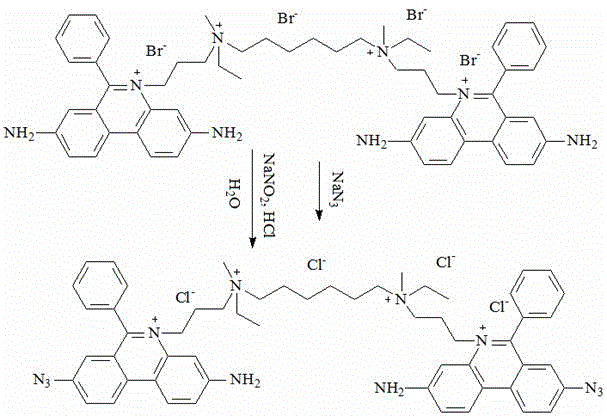
(6) and mixing the intermediate V and hydrochloric acid, then dropwise adding a sodium nitrite solution, stirring, dropwise adding a sodium azide solution, and reacting to obtain the fluorescent dye.
In the technical proposal, the device comprises a base,
the chemical structural formula of the intermediate I is as follows:
the chemical structural formula of the intermediate II is as follows:
the chemical structural formula of the intermediate III is one of the following structural formulas:
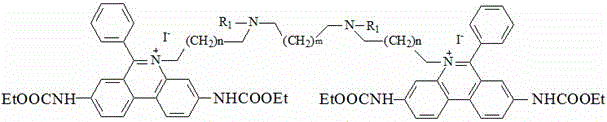
the chemical structural formula of the intermediate IV is one of the following structural formulas:
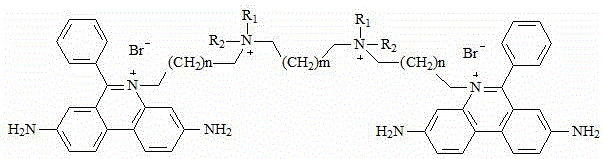
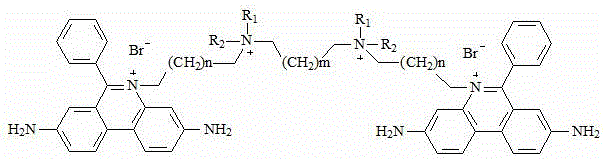
the chemical structural formula of the intermediate V is one of the following structural formulas:
in the technical scheme, in the step (1), when the intermediate I is prepared, the reaction temperature is 0-5 ℃; in the step (2), when the intermediate II is prepared, the reaction temperature is 105-115 ℃; in the step (3), when the intermediate III is prepared, the reaction temperature is reflux; in the step (4), when the intermediate IV is prepared, the reaction temperature is reflux; in the step (5), when the intermediate V is prepared, the reaction temperature is 105-115 ℃; in the step (6), the reaction temperature is less than 0 ℃.
The invention also discloses application of the fluorescent dye in VBNC bacteria PCR detection and application of the fluorescent dye in bacteria detection.
The invention also discloses the application of the fluorescent dye in marking non-viable bacteria; a method for labeling non-viable bacteria, comprising the following steps of incubating the fluorochrome and bacteria in claim 1 for 5-50 minutes to complete labeling of the non-viable bacteria, wherein the labeled dead bacteria can emit strong fluorescence, especially can not participate in amplification reaction, and the problem of false positive caused by the dead bacteria is solved.
The product of the invention not only has little toxicity to cells, namely, the impermeability to cell membranes is greatly increased, but also has more thorough elimination of PCR products in dead cell DNA compared with the prior product, does not need additional enhanced buffer solution, has very wide application range, and can be used for treating salmonella, staphylococcus aureus, drug-resistant staphylococcus aureus, escherichia coli, mycobacterium tuberculosis, listeria and the like.
Drawings
FIG. 1 is a mass spectrum of structure number 1 of Table 1 of the present invention;
FIG. 2 is a mass spectrum of structure number 2 of Table 1 of the present invention;
FIG. 3 is a fluorescence plot of the dye number 3 of Table 1 of the present invention and the conventional dye PMAxx at 5 minutes after labeling the bacteria;
FIG. 4 is a fluorescence plot of the dye number 3 of Table 1 of the present invention and the conventional dye PMAxx at 10 minutes after labeling bacteria;
FIG. 5 is a fluorescence plot of the dye number 3 of Table 1 of the present invention and the conventional dye PMAxx at 30 minutes after labeling bacteria;
FIG. 6 is a graph comparing the qPCR results of the dye number 3 of Table 1 of the present invention with the prior dye PMAxX treated bacteria.
Detailed Description
Example one
Putting 60.0 g of 3, 8-diamino-6-phenylphenanthridine, 360 mL of DMF (dimethyl formamide) and 37.2 mL of pyridine into a 1000 mL three-neck round-bottom flask, mechanically stirring, dropwise adding 42 mL of methyl chloroacetate in an ice water bath (0-5 ℃), and keeping the temperature of the dropwise adding process at 0-5 ℃; after the dropwise addition is finished, continuing the reaction for 10 hours until the reaction is finished; performing suction filtration, adding the filter cake into about 2L of pure water, mechanically stirring for 30 min, and performing suction filtration; washing the filter cake with 2L of pure water again, and performing suction filtration until the filter cake is dry; drying under reduced pressure to constant weight to obtain 48 g of intermediate I.
Example two
In a 1L three-necked flask equipped with a temperature controller and mechanical stirring, the intermediate 20 g I and 80 mL of 1, 3-diiodopropane were added, mixed and stirred, reacted at 105 ℃ to 115 ℃ for 3 days, followed by TLC, with a developing solvent of dichloromethane: methanol: ethyl acetate: acetic acid = 75: 10: 10: 5, according to volume ratio;
after the reaction was completed, the reaction was stopped and cooled to room temperature. Adding about 150 mL of ethyl acetate into the reaction system, heating and refluxing for 1 hour, cooling to room temperature, and filtering; the solid obtained by filtration was washed with ethyl acetate. The collected solid was dried in a vacuum oven to give 22 g of intermediate II.
EXAMPLE III
To a 250 mL egg-shaped flask was added 7.25 g of intermediate II, 60 mL MeOH, and 0.65 gNHeating to reflux state, reacting overnight, tracking by TLC, and developing by dichloromethane: methanol: ethyl acetate: acetic acid = 75: 10: 10: 5, according to volume ratio;
after the reaction is finished, the solvent is removed by rotation, 40 mL of ethyl acetate is added, the mixture is refluxed for 30 minutes, then cooled to the normal temperature, filtered, the filter cake is washed by ethyl acetate, and after the filter cake is dried in vacuum for 12 hours, 5.5 g of intermediate III is obtained.
Example four
A100 mL reaction flask was charged with 2.62 g of intermediate III, 676 mg of 1, 6-diiodohexane, 40 mL of methanol, stirred, heated to reflux for overnight reaction, followed by TLC with the following developer: acetonitrile: water = 5: 1, according to volume ratio;
after the reaction was completed, the solvent was removed by rotation, purified by an aluminum peroxide column, and eluted with water/acetonitrile (2% -8%) to obtain 1.46 g of intermediate IV.
EXAMPLE five
Adding 1.46 g of intermediate and 20 mL of hydrobromic acid into a 100 mL reaction bottle, stirring, and heating to 110 ℃ for reacting overnight; TLC tracking, the developing solvent was: acetonitrile water = 10: 1, according to volume ratio;
after the reaction was complete, the reaction mixture was cooled to room temperature, filtered, washed with 100 mL of water, and the solid was dried to give 0.7 g of intermediate V.
EXAMPLE six
Adding 0.6 g of intermediate V and 30 mL of hydrochloric acid (1M) into a 100 mL three-necked bottle, stirring, and cooling to-5 ℃; slowly dripping 2 mL of sodium nitrite solution (142 mg) and keeping the temperature less than zero; TLC tracking, the developing solvent was: acetonitrile water = 10: 3, according to volume ratio;
after the reaction is finished, slowly dropwise adding 2 mL of sodium azide solution (75 mg), keeping the temperature less than zero, and reacting for 10 hours; TLC tracking, the developing solvent was: acetonitrile water = 10: 3, according to volume ratio;
after the reaction is finished, cooling the system to zero degree, dropwise adding a sodium hydroxide solution for neutralization to neutrality, and freeze-drying; purifying with aluminum peroxide column, eluting with water/acetonitrile (5% -10%) to obtain 0.35 g final product.
EXAMPLE seven
According to the method, various products can be obtained by replacing the raw material amine compound, and particularly, the mass spectrograms of the structures of the numbers 1 and 2 in the table 1 are shown in the attached figures 1 and 2 respectively, and are shown as a half of the obtained molecular weight because of the charge relation.
TABLE 1 structural formula of the high efficiency fluorescent dye of the present invention
Example eight cytotoxicity comparison
The experimental steps are as follows: coli cells are cultured in a 24-well plate, then PMAxx and the fluorescent dye with the number 3 in the table 1 are respectively added into the cells, the final concentration is 50 μ M, the cells are incubated at 37 ℃ for a period of time, for example, the cells are respectively incubated for 5, 15 and 30 min, pictures are taken at each time point, fluorescence pictures of different time points are respectively collected by using a Cy3 excitation light source, and the fluorescence pictures of the existing PMAxx and the fluorescent dye with the number 3 in the invention at different time points are respectively shown in the attached figures 3, 4 and 5.
EXAMPLE nine test results for live-dead bacteria
The procedure was carried out using the method published by the company Biotium, the live or heat-inactivated Bacillus subtilis being treated with 50. mu.M of the fluorescent dye numbered 3 in Table 1 or PMAxx and then exposed by purification with PMA-Lite and DNA from the company. Using the UEIris fluorescent quantitation kit produced by Suzhou constant Biotechnology, Inc. to amplify a 500-bp fragment of Bacillus subtilis DNA, treatment of bacteria with such dyes did not affect amplification of viable cell DNA, but resulted in a decrease in the number of fluorescent quantitation (qPCR) cycles (Ct) in dead cells. The ordinate of fig. 6 is the fluorescence quantification (qPCR) cycle (Ct) by treatment of the sample with the dye minus the fluorescence quantification (qPCR) cycle (Ct) without treatment with the dye, and it can be seen from fig. 6 that when live bacteria are treated under the same conditions, the PMAxx by the false positive Biotium company is significantly higher than the product of invention No. 3 due to the toxicity of the product itself, and when samples containing dead bacteria are treated, the fluorescent dye of the invention is significantly better than the PMAxx by the Biotium company, and qPCR of dead cells treated with the fluorescent dye of the invention shows a significantly further delay (> 6 Ct) compared to dead cells treated with Biotium PMAxx.
Compared with the existing products, the product of the invention has low toxicity to cells, greatly increases the impermeability to cell membranes, is more thorough in eliminating PCR products in dead cell DNA than the existing products, and does not need additional enhancing buffer solution. The application range is wide, and salmonella, staphylococcus aureus, drug-resistant staphylococcus aureus, escherichia coli, tubercle bacillus, listeria and the like can be treated by the product.
Claims (5)
1. A preparation method of a fluorescent dye comprises the following steps:
(1) preparing an intermediate I by using 3, 8-diamino-6-phenylphenanthridine and ethyl chloroformate as raw materials;
(2) preparing an intermediate II by taking the intermediate I and iodide as raw materials;
(3) preparing an intermediate III by taking the intermediate II and an amine compound as raw materials;
(4) preparing an intermediate IV by taking the intermediate III and an iodine compound as raw materials;
(5) preparing an intermediate V by taking the intermediate IV and hydrobromic acid as raw materials;
(6) mixing the intermediate V and acid, then dropwise adding a sodium nitrite solution, stirring, and dropwise adding a sodium azide solution to react to obtain a fluorescent dye;
the chemical structural formula of the intermediate I is as follows:
the chemical structural formula of the intermediate II is as follows:
the chemical structural formula of the intermediate III is one of the following structural formulas:
the chemical structural formula of the intermediate IV is one of the following structural formulas:
the chemical structural formula of the intermediate V is one of the following structural formulas:
the chemical structural formula of the iodide in the step (2) is I (CH)2)nI; the chemical structure of the amine compound is R1NHR2Or R1NH2(ii) a The chemical structural formula of the iodine compound in the step (4) is I (CH)2)mI; the acid is hydrochloric acid; r1、R2Independently selected from linear alkyl of C1-C3; n is 1-4; m is 1 to 4.
3. The method for preparing a fluorescent dye according to claim 2, wherein R' is two chloride ions; r is four chloride ions.
5. the method for preparing fluorescent dye according to claim 1, wherein in the step (1), the reaction temperature is 0-5 ℃ when preparing the intermediate I; in the step (2), when the intermediate II is prepared, the reaction temperature is 105-115 ℃; in the step (3), when the intermediate III is prepared, the reaction temperature is reflux; in the step (4), when the intermediate IV is prepared, the reaction temperature is reflux; in the step (5), when the intermediate V is prepared, the reaction temperature is 105-115 ℃; in the step (6), the reaction temperature is less than 0 ℃.
Priority Applications (1)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| CN201911185975.5A CN110845862B (en) | 2017-12-29 | 2017-12-29 | Preparation method of fluorescent dye |
Applications Claiming Priority (2)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| CN201711479943.7A CN108047135B (en) | 2017-12-29 | 2017-12-29 | A kind of fluorescent dye and its preparation method and application in bacteria detection |
| CN201911185975.5A CN110845862B (en) | 2017-12-29 | 2017-12-29 | Preparation method of fluorescent dye |
Related Parent Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| CN201711479943.7A Division CN108047135B (en) | 2017-12-29 | 2017-12-29 | A kind of fluorescent dye and its preparation method and application in bacteria detection |
Publications (2)
| Publication Number | Publication Date |
|---|---|
| CN110845862A CN110845862A (en) | 2020-02-28 |
| CN110845862B true CN110845862B (en) | 2021-03-19 |
Family
ID=62129101
Family Applications (2)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| CN201911185975.5A Active CN110845862B (en) | 2017-12-29 | 2017-12-29 | Preparation method of fluorescent dye |
| CN201711479943.7A Active CN108047135B (en) | 2017-12-29 | 2017-12-29 | A kind of fluorescent dye and its preparation method and application in bacteria detection |
Family Applications After (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| CN201711479943.7A Active CN108047135B (en) | 2017-12-29 | 2017-12-29 | A kind of fluorescent dye and its preparation method and application in bacteria detection |
Country Status (1)
| Country | Link |
|---|---|
| CN (2) | CN110845862B (en) |
Families Citing this family (4)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN111363377B (en) * | 2020-03-13 | 2021-11-30 | 苏州优逸兰迪生物科技有限公司 | Nucleic acid dye and preparation method and application thereof |
| CN114891903A (en) * | 2022-04-25 | 2022-08-12 | 华南农业大学 | Kit and method for rapidly detecting salmonella live bacteria based on PMAxx-LAMP-NALFA |
| CN119604588A (en) * | 2022-07-29 | 2025-03-11 | 松下知识产权经营株式会社 | Organic pigment composition and spray containing the same |
| CN119462681B (en) * | 2024-10-16 | 2025-09-30 | 聊城柏莱斯生物科技有限公司 | A fluorescent quantitative PCR reference dye and its preparation method and application |
Citations (3)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US5545535A (en) * | 1993-04-13 | 1996-08-13 | Molecular Probes, Inc. | Fluorescent assay for bacterial gram reaction |
| EP1348713A2 (en) * | 2002-03-12 | 2003-10-01 | Enzo Life Sciences, Inc. | Labeling reagents and labeled tabeled targets, target labeling processes and other processes for using same in nucleic acid determinations and analyses |
| CN100560733C (en) * | 2001-07-20 | 2009-11-18 | 英詹尼斯公司 | Parallel, antiparallel, homologous or complementary binding of nucleic acids |
-
2017
- 2017-12-29 CN CN201911185975.5A patent/CN110845862B/en active Active
- 2017-12-29 CN CN201711479943.7A patent/CN108047135B/en active Active
Patent Citations (3)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US5545535A (en) * | 1993-04-13 | 1996-08-13 | Molecular Probes, Inc. | Fluorescent assay for bacterial gram reaction |
| CN100560733C (en) * | 2001-07-20 | 2009-11-18 | 英詹尼斯公司 | Parallel, antiparallel, homologous or complementary binding of nucleic acids |
| EP1348713A2 (en) * | 2002-03-12 | 2003-10-01 | Enzo Life Sciences, Inc. | Labeling reagents and labeled tabeled targets, target labeling processes and other processes for using same in nucleic acid determinations and analyses |
Also Published As
| Publication number | Publication date |
|---|---|
| CN110845862A (en) | 2020-02-28 |
| CN108047135B (en) | 2019-12-17 |
| CN108047135A (en) | 2018-05-18 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| CN110845862B (en) | Preparation method of fluorescent dye | |
| CN108138217B (en) | Method for specifically labeling living microorganisms comprising the use of modified monosaccharide compounds | |
| AU2019320825B2 (en) | Methods and compositions for nucleic acid isolation | |
| ES2846884T3 (en) | A method of specifically marking live bacteria comprising the use of modified non-endogenous monosaccharide compounds | |
| Ye et al. | Molecular methods for rapid detection and identification of foodborne pathogenic bacteria | |
| CN104194379B (en) | A kind of thiazole orange class Molecule of Cyanine Dyes and application thereof | |
| Váradi et al. | β-Alanyl aminopeptidase-activated fluorogenic probes for the rapid identification of Pseudomonas aeruginosa in clinical samples | |
| CN105906817A (en) | Amphoteric carboxylic acid hexanuclear copper metal coordination polymer and preparation method thereof | |
| WO1999050438A2 (en) | Chromogenic indole derivatives | |
| KR102243449B1 (en) | A nucleic acid aptamer specifically binding to Enterobacter sakazakii and the use thereof | |
| CN107383927A (en) | A kind of nitrine indoles dimethime Molecule of Cyanine Dyes and application | |
| JP3607327B2 (en) | Rapid detection of single microbial cells | |
| Csuk et al. | Synthesis of pathogen inactivating nucleic acid intercalators | |
| JP5134071B2 (en) | Method for avoiding inhibition in nucleic acid amplification reaction | |
| RU2764021C1 (en) | METHOD FOR DETECTING SARS-CoV-2 VIRUS RNA IN ULTRA-LOW CONCENTRATIONS AND SPECIFIC OLIGONUCLEOTIDES FOR USE IN THE METHOD | |
| RU2764023C1 (en) | CRISPR-Cas 12 SYSTEM FOR DETECTING SARS-CoV-2 VIRUS RNA IN ULTRA-LOW CONCENTRATIONS | |
| RU2764020C1 (en) | CRISPR-CAS14 SYSTEM FOR DETECTING SARS-CoV-2 VIRUS RNA AT ULTRA-LOW CONCENTRATIONS | |
| CN116004872B (en) | A primer combination for detecting drug-resistant genes and its application | |
| RU2747820C1 (en) | Crispr-cas system for detection of john cunningham virus (jcpyv) dna at ultra-low concentrations | |
| RU2747880C1 (en) | Method for detecting dna of john cunningham virus (jcpyv) at ultra-low concentrations and specific oligonucleotides for use in method | |
| RU2747819C1 (en) | Method for obtaining preparation of ribonucleoprotein complex crispr / cas and preparation for detecting dna of john cunningham virus (jcpyv) in ultra-low concentrations | |
| CN113004886B (en) | Preparation method and application of benzoquinolines ratio type near-infrared fluorescent molecular probe for fluorine ion detection | |
| CN111455021A (en) | Method and kit for removing host DNA in metagenome | |
| Csuk et al. | Synthesis of monomeric acridine derived nucleic acid intercalators | |
| Ishizuka et al. | Click chemistry takes a snapshot of DNA-RNA hybrid G-quadruplex in living cells |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| PB01 | Publication | ||
| PB01 | Publication | ||
| SE01 | Entry into force of request for substantive examination | ||
| SE01 | Entry into force of request for substantive examination | ||
| TA01 | Transfer of patent application right | ||
| TA01 | Transfer of patent application right |
Effective date of registration: 20201218 Address after: Room 320-1, building 1, 18 Chunyao Road, Caohu Industrial Park, Xiangcheng Economic Development Zone, Suzhou City, Jiangsu Province Applicant after: Suzhou Youyi landi Biotechnology Co.,Ltd. Address before: 215100 room 1001, No.1, Guantang Road, economic development zone, Xiangcheng District, Suzhou City, Jiangsu Province Applicant before: Suzhou Yuheng Biological Technology Co.,Ltd. Applicant before: SUQIAN AIRUISI BIOTECHNOLOGY Co.,Ltd. |
|
| GR01 | Patent grant | ||
| GR01 | Patent grant |