CN110551669B - Near-infrared light control dynamic regulation and control system and application thereof - Google Patents
Near-infrared light control dynamic regulation and control system and application thereof Download PDFInfo
- Publication number
- CN110551669B CN110551669B CN201910836782.5A CN201910836782A CN110551669B CN 110551669 B CN110551669 B CN 110551669B CN 201910836782 A CN201910836782 A CN 201910836782A CN 110551669 B CN110551669 B CN 110551669B
- Authority
- CN
- China
- Prior art keywords
- leu
- ala
- arg
- glu
- gly
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Active
Links
Images
Classifications
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K14/00—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof
- C07K14/195—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from bacteria
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K14/00—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof
- C07K14/195—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from bacteria
- C07K14/24—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from bacteria from Enterobacteriaceae (F), e.g. Citrobacter, Serratia, Proteus, Providencia, Morganella, Yersinia
- C07K14/245—Escherichia (G)
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/63—Introduction of foreign genetic material using vectors; Vectors; Use of hosts therefor; Regulation of expression
- C12N15/70—Vectors or expression systems specially adapted for E. coli
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12P—FERMENTATION OR ENZYME-USING PROCESSES TO SYNTHESISE A DESIRED CHEMICAL COMPOUND OR COMPOSITION OR TO SEPARATE OPTICAL ISOMERS FROM A RACEMIC MIXTURE
- C12P7/00—Preparation of oxygen-containing organic compounds
- C12P7/24—Preparation of oxygen-containing organic compounds containing a carbonyl group
- C12P7/26—Ketones
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N2800/00—Nucleic acids vectors
- C12N2800/60—Vectors containing traps for, e.g. exons, promoters
Landscapes
- Chemical & Material Sciences (AREA)
- Health & Medical Sciences (AREA)
- Organic Chemistry (AREA)
- Life Sciences & Earth Sciences (AREA)
- Genetics & Genomics (AREA)
- Engineering & Computer Science (AREA)
- General Health & Medical Sciences (AREA)
- Biochemistry (AREA)
- Wood Science & Technology (AREA)
- Zoology (AREA)
- Biophysics (AREA)
- Bioinformatics & Cheminformatics (AREA)
- General Engineering & Computer Science (AREA)
- Biotechnology (AREA)
- Molecular Biology (AREA)
- Gastroenterology & Hepatology (AREA)
- Microbiology (AREA)
- Proteomics, Peptides & Aminoacids (AREA)
- Medicinal Chemistry (AREA)
- Biomedical Technology (AREA)
- General Chemical & Material Sciences (AREA)
- Plant Pathology (AREA)
- Physics & Mathematics (AREA)
- Chemical Kinetics & Catalysis (AREA)
- Preparation Of Compounds By Using Micro-Organisms (AREA)
- Micro-Organisms Or Cultivation Processes Thereof (AREA)
Abstract
本发明公开了一种近红外光控制的动态调控系统及其应用,具体公开了一种近红外光组合控制的涉及动态调控系统及其在乙偶姻生产中的应用,属于代谢工程技术领域。本发明通过分子生物学手段,选择了近红外光光敏蛋白。利用光敏蛋白特异性响应特定波长的光源特性和无需外源添加诱导剂的优势通过在工程大肠杆菌中引入靶向RpoS的动态调控基因线路,实现了在无机盐培养基中生产乙偶姻。通过本发明生产的乙偶姻产量可达66.8g/L,转化率达0.58g/g葡萄糖。
The invention discloses a near-infrared light-controlled dynamic regulation system and its application, and specifically discloses a near-infrared light combined-controlled dynamic regulation system and its application in the production of acetoin, belonging to the technical field of metabolic engineering. The present invention selects the near-infrared photosensitive protein by means of molecular biology. The production of acetoin in inorganic salt medium was achieved by introducing a dynamic regulatory gene circuit targeting RpoS in engineered E. The yield of acetoin produced by the method can reach 66.8 g/L, and the conversion rate can reach 0.58 g/g glucose.
Description
技术领域technical field
本发明涉及一种近红外光控制的动态调控系统及其应用,具体涉及一种近红外光控制的动态调控系统及其在乙偶姻生产中的应用,属于代谢工程技术领域。The invention relates to a near-infrared light-controlled dynamic regulation system and its application, in particular to a near-infrared light-controlled dynamic regulation system and its application in the production of acetoin, belonging to the technical field of metabolic engineering.
背景技术Background technique
动态调控系统是代谢工程领域中新兴的代谢流调控手段,其区别于静态调控的主要特征是在发酵过程中,工程菌株会根据发酵时间、生理状态、胞内代谢物浓度、胞外环境变化做出相应的酶活调整,进而影响代谢流分布,提高产品生产能力。动态调控系统具有发酵过程无需人为调控、不需要外源添加诱导剂的优势,在生产高附加值化合物中具有显著优势。The dynamic regulation system is an emerging metabolic flow regulation method in the field of metabolic engineering. The main feature that is different from the static regulation is that during the fermentation process, the engineered strain will act according to the fermentation time, physiological state, intracellular metabolite concentration, and changes in the extracellular environment. The corresponding enzyme activity adjustment can be made, which in turn affects the distribution of metabolic flux and improves the production capacity of the product. The dynamic regulation system has the advantages that the fermentation process does not need manual regulation and does not need to add external inducers, and has significant advantages in the production of high value-added compounds.
目前的动态调控系统,主要是依靠群体感应系统进行控制的,即系统的触发器是由本源或者异源的群体感应系统组成。当菌体浓度达到一定浓度时,群体感应系统释放积累的小分子物质会作用于调控系统受体,使调控系统做出相应动作。如2017年,ApoorvGupta等人表于Nature biotechnology的动态系统,该系统由EsaI群体感应系统和靶蛋白C末端添加降解标签组合而成,当菌体浓度达到一定阈值时,靶蛋白的被降解,胞内绝对浓度达到0,从而实现酶活关闭,调控代谢流的目的和效果。该系统具有路径独立特征,并成功应用于肌醇、葡萄糖二酸和乙偶姻的生产应用。除此之外,2017年,Xinyuan He等人发表于ACSsynthetic biology的AND逻辑门动态调控系统由群体感应系统和稳定期感应蛋白组成。当菌体浓度达到一定阈值时,启动靶蛋白的转录表达,从而实现代谢流调控目的和效果。该系统运用于PHB生产时,提高了1-2倍PHB产量。综上所述,两种动态调控系统均需要引入异源群体感应系统,然而,异源群体感应系统往往由多个转录调控蛋白组成,它的引入无疑会影响菌株的生长和发酵性能,同时限制了路径酶的表达数量。The current dynamic regulation system is mainly controlled by the quorum sensing system, that is, the trigger of the system is composed of the original or heterologous quorum sensing system. When the concentration of bacteria reaches a certain concentration, the accumulated small molecules released by the quorum sensing system will act on the receptors of the regulatory system, so that the regulatory system will act accordingly. For example, in 2017, Apoorv Gupta et al. presented the dynamic system of Nature biotechnology, which is composed of the EsaI quorum sensing system and the addition of a degradation tag to the C-terminus of the target protein. When the bacterial concentration reaches a certain threshold, the target protein is degraded and the cell The internal absolute concentration reaches 0, so as to achieve the purpose and effect of closing the enzyme activity and regulating the metabolic flow. The system is path-independent and has been successfully applied in the production of inositol, glucaric acid and acetoin. In addition, in 2017, the AND logic gate dynamic regulation system published by Xinyuan He et al. in ACSsynthetic biology consists of a quorum sensing system and a stationary phase sensing protein. When the bacterial cell concentration reaches a certain threshold, the transcription and expression of the target protein is started, so as to achieve the purpose and effect of metabolic flux regulation. When the system is used in PHB production, the PHB output is increased by 1-2 times. In summary, both dynamic regulatory systems require the introduction of a heterologous quorum sensing system. However, the heterologous quorum sensing system is often composed of multiple transcriptional regulatory proteins, and its introduction will undoubtedly affect the growth and fermentation performance of the strain, while limiting the expression of the pathway enzymes.
乙偶姻是制备香精香料的前体,在大肠杆菌中由丙酮酸路径合成。传统的生产乙偶姻的方法通过过表达路径酶,实现乙偶姻的积累。然而,乙偶姻的生产受到比表面积的限制,引起溶氧传质不均匀的问题,直接影响乙偶姻在大肠杆菌中的生长和积累。因此,提供一种在简单培养基中高产乙偶姻的方法,对于工业制备乙偶姻具有重要的意义。Acetoin is a precursor for the preparation of flavors and fragrances, which is synthesized by the pyruvate pathway in Escherichia coli. The traditional method of producing acetoin achieves the accumulation of acetoin by overexpressing the pathway enzymes. However, the production of acetoin is limited by the specific surface area, causing the problem of uneven mass transfer of dissolved oxygen, which directly affects the growth and accumulation of acetoin in E. coli. Therefore, to provide a method for high-yield acetoin in a simple medium is of great significance for the industrial preparation of acetoin.
发明内容SUMMARY OF THE INVENTION
本发明的第一个目的是提供一种工程菌株,共同表达近红外光光敏色素、近红外光光响应组分和靶蛋白,所述近红外光光敏色素包括Bphs、Bpho、Yhjh和/或Mrkh;所述近红外光光响应组分包括启动子PmrkA,所述Bphs的氨基酸序列如SEQ ID NO.14所示,所述Bpho的氨基酸序列如SEQ ID NO.15所示,所述Yhjh的氨基酸序列如SEQ ID NO.16所示,所述Mrkh的氨基酸序列如SEQ ID NO.17所示,所述PmrkA的核苷酸序列如SEQ ID NO.5所示。The first object of the present invention is to provide an engineered strain that co-expresses a near-infrared photophytochrome, a near-infrared photosensitive component and a target protein, the near-infrared photophytochrome includes Bphs, Bpho, Yhjh and/or Mrkh The near-infrared light-responsive component comprises a promoter PmrkA , the amino acid sequence of Bphs is shown in SEQ ID NO.14, the amino acid sequence of Bpho is shown in SEQ ID NO.15, and the amino acid sequence of Yhjh is shown in SEQ ID NO.15. The amino acid sequence is shown in SEQ ID NO.16, the amino acid sequence of Mrkh is shown in SEQ ID NO.17, and the nucleotide sequence of P mrkA is shown in SEQ ID NO.5.
在本发明的一种实施方式中,编码所述Bphs的核苷酸序列如SEQ ID NO.1所示,编码所述Bpho的核苷酸序列如SEQ ID NO.2所示,编码所述Yhjh的核苷酸序列如SEQ ID NO.3所示,编码所述Mrkh的核苷酸序列如SEQ ID NO.4所示。In one embodiment of the present invention, the nucleotide sequence encoding the Bphs is shown in SEQ ID NO. 1, and the nucleotide sequence encoding the Bpho is shown in SEQ ID NO. 2, encoding the Yhjh The nucleotide sequence of Mrkh is shown in SEQ ID NO.3, and the nucleotide sequence encoding the Mrkh is shown in SEQ ID NO.4.
在本发明的一种实施方式中,所述靶蛋白为RpoS,所述RpoS的氨基酸序列如SEQID NO.18所示。In one embodiment of the present invention, the target protein is RpoS, and the amino acid sequence of the RpoS is shown in SEQ ID NO.18.
在本发明的一种实施方式中,以PmrkA-RpoS和PJ23119-SO-PJ23119-Y30M-PJ23119-BudAB-NOX为表达载体。In one embodiment of the present invention, P mrkA -RpoS and P J23119 -SO-P J23119 -Y 30 MP J23119 -BudAB-NOX are used as expression vectors.
PJ23119-SO-PJ23119-Y30M的构建方法为:P J23119 -SO-P J23119 -Y 30 M is constructed as:
(1)以商业化质粒pTargetF(Addgene Plasmid#62226)为基础,采用全质粒PCR的方式,在rrnB T1终止子后插入T7Te终止子序列,以减少泄露表达;采用全质粒PCR的方式,去除sgRNA表达框,获得仅含有Pj23119组成型启动子和双终止的工程质粒PJ23119;(1) Based on the commercial plasmid pTargetF (Addgene Plasmid#62226), the whole plasmid PCR method was used to insert the T7Te terminator sequence after the rrnB T1 terminator to reduce leaked expression; the whole plasmid PCR method was used to remove the sgRNA Expression cassette, obtain the engineering plasmid P J23119 containing only Pj23119 constitutive promoter and double termination;
(2)以类球红细菌(Rhodobacter sphaeroides)基因组为模板,分别扩增得到Bphs、Bpho、Yhjh基因片段;以肺炎杆菌基因组(Klebsiella pneumoniae)为模板,扩增得到mrkH基因片段;采用同源重组的方式,将带有B0034RBS(核苷酸序列如SEQ ID NO.7所示)的Bphs和B0034RBS的Bpho片段插入PJ23119表达框中,获得PJ23119-SO质粒;以同样的方式,将带有B0030RBS(核苷酸序列如SEQ ID NO.8所示)的Yhjh和B0034RBS的mrkH片段插入PJ23119表达框中,获得PJ23119-Y30M质粒;利用同源重组的方法将PJ23119-SO和PJ23119-Y30M两个质粒整合到一起,获得PJ23119-SO-PJ23119-Y30M质粒。(2) Using the genome of Rhodobacter sphaeroides as a template, the Bphs, Bpho and Yhjh gene fragments were amplified respectively; the genome of Klebsiella pneumoniae was used as a template, and the mrkH gene fragment was obtained by amplification; homologous recombination was used In the same way, the Bphs and Bpho fragments of B0034RBS with B0034RBS (nucleotide sequence shown in SEQ ID NO. 7) were inserted into the P J23119 expression frame to obtain the P J23119- SO plasmid; The Yhjh of B0030RBS (nucleotide sequence shown in SEQ ID NO. 8) and the mrkH fragment of B0034RBS were inserted into the P J23119 expression frame to obtain the P J23119 -Y 30 M plasmid; the P J23119-SO and P J23119- SO and The two plasmids P J23119 -Y 30 M were integrated together to obtain the P J23119 -SO-P J23119 -Y 30 M plasmid.
PmrkA-BFP质粒的构建方法为:以bfp合成基因片段(如SEQ ID NO.6所示)为模板,扩增含有B0034RBS的bfp基因,同源重组方式连接到PJ23119质粒上,同时反向扩增得到PmrkA质粒,同源重组方式获得PmrkA-BFP质粒。The construction method of P mrkA- BFP plasmid is as follows: using bfp synthetic gene fragment (as shown in SEQ ID NO. 6) as a template, amplify the bfp gene containing B0034RBS, connect it to the P J23119 plasmid by homologous recombination, and reverse it at the same time. The P mrkA plasmid was obtained by amplification, and the P mrkA-BFP plasmid was obtained by homologous recombination.
PJ23119-BudAB-NOX质粒的构建方法为:以粘质沙雷氏菌(Serratia marcescens)H30基因组为模板,设计带有B0034RBS(核苷酸序列如SEQ ID NO.7所示)的引物,分别扩增获得含有B0034RBS的budA(核苷酸序列如SEQ ID NO.9所示)、budB片段(核苷酸序列如SEQID NO.10所示),将budA、budB两个片段采用多片段一步同源重组的方式,插入至PJ23119(核苷酸序列如SEQ ID NO.11所示)的表达框中,获得PJ23119-BudAB质粒;以短乳杆菌(Lactobacillus brevis)基因组为模板,扩增nox基因,将nox基因片段(如SEQ ID NO.12所示)采用多片段一步同源重组的方式,插入至PJ23119-BudAB的表达框中,获得PJ23119-BudAB-NOX质粒。The construction method of P J23119 -BudAB-NOX plasmid is as follows: using Serratia marcescens H30 genome as a template, design primers with B0034RBS (nucleotide sequence shown in SEQ ID NO. 7), respectively. Amplify to obtain budA (nucleotide sequence as shown in SEQ ID NO. 9) and budB fragments (nucleotide sequence as shown in SEQ ID NO. 10) containing B0034RBS, and the two fragments of budA and budB adopt multi-fragment one-step synchronization. The mode of source recombination was inserted into the expression frame of P J23119 (nucleotide sequence as shown in SEQ ID NO.11) to obtain the P J23119 -BudAB plasmid; the Lactobacillus brevis genome was used as a template to amplify nox gene, the nox gene fragment (as shown in SEQ ID NO. 12) was inserted into the expression frame of P J23119 -BudAB by means of multi-fragment one-step homologous recombination to obtain the P J23119 -BudAB-NOX plasmid.
所述PJ23119-SO-PJ23119-Y30M-PJ23119-BudAB-NOX质粒的构建方法为:以PJ23119-BudAB-NOX质粒为模板,通过同源重组扩增出PJ23119-BudAB-NOX片段,然后以PJ23119-SO-PJ23119-Y30M为载体,酶切,同源重组,插入PJ23119-SO-PJ23119-Y30M表达框,获得PJ23119-SO-PJ23119-Y30M-PJ23119-BudAB-NOX质粒。The construction method of the P J23119 -SO-P J23119 -Y 30 MP J23119 -BudAB-NOX plasmid is as follows: using the P J23119 -BudAB-NOX plasmid as a template, the P J23119 -BudAB-NOX fragment is amplified by homologous recombination, Then, using P J23119 -SO-P J23119 -Y 30 M as a vector, enzyme digestion, homologous recombination, and inserting the P J23119 -SO-P J23119 -Y 30 M expression cassette to obtain P J23119 -SO-P J23119 -Y 30 MP J23119 - BudAB-NOX plasmid.
所述PmrkA-RpoS质粒的构建方法为:利用同源重组的方法将PmrkA-BFP质粒中的编码BFP荧光蛋白的基因替换成编码大肠杆菌内源的RpoS蛋白的基因(核苷酸序列如SEQ IDNO.14所示)。The construction method of described PmrkA- RpoS plasmid is: utilize the method of homologous recombination to replace the gene encoding BFP fluorescent protein in the PmrkA -BFP plasmid with the gene encoding the endogenous RpoS protein of Escherichia coli (the nucleotide sequence is as shown in Fig. shown in SEQ ID NO. 14).
在本发明的一种实施方式中,以E.coli F0601为宿主细胞。In one embodiment of the present invention, E. coli F0601 is used as the host cell.
本发明的第二个目的是提供一种调控靶蛋白基因表达的方法,是在工程菌株发酵过程中补充近红外光以控制靶蛋白基因的表达,所述工程菌株共同表达近红外光光敏色素、近红外光光响应组分和靶蛋白,所述近红外光光敏色素包括Bphs、Bpho、Yhjh或Mrkh;所述近红外光光响应组分包括启动子PmrkA,所述Bphs的氨基酸序列如SEQ ID NO.14所示,所述Bpho的氨基酸序列如SEQ ID NO.15所示,所述Yhjh的氨基酸序列如SEQ ID NO.16所示,所述Mrkh的氨基酸序列如SEQ ID NO.17所示,所述PmrkA的核苷酸序列如SEQ ID NO.5所示。The second object of the present invention is to provide a method for regulating the expression of a target protein gene, which is to supplement near-infrared light in the fermentation process of an engineered strain to control the expression of the target protein gene, and the engineered strains co-express near-infrared photophytochrome, A near-infrared light-responsive component and a target protein, the near-infrared photophytochrome includes Bphs, Bpho, Yhjh or Mrkh; the near-infrared light-responsive component includes a promoter P mrkA , and the amino acid sequence of the Bphs is as shown in SEQ ID NO.14, the amino acid sequence of Bpho is shown in SEQ ID NO.15, the amino acid sequence of Yhjh is shown in SEQ ID NO.16, and the amino acid sequence of Mrkh is shown in SEQ ID NO.17 As shown, the nucleotide sequence of the P mrkA is shown in SEQ ID NO.5.
在本发明的一种实施方式中,所述近红外光的光照强度为0.2W/cm2-0.8W/cm2,波长为600-650nm,与菌株培养基的距离固定为5-15cm。In an embodiment of the present invention, the illumination intensity of the near-infrared light is 0.2W/cm 2 -0.8W/cm 2 , the wavelength is 600-650nm, and the distance from the strain medium is fixed at 5-15cm.
在本发明的一种实施方式中,波长为650nm,与菌株培养基的距离固定为10cm。In one embodiment of the present invention, the wavelength is 650 nm, and the distance from the strain medium is fixed at 10 cm.
本发明的第三个目的是提供一种生产乙偶姻的方法,将上述工程菌株单菌落接种至LB培养基中进行活化,活化后接入发酵培养基进行发酵。The third object of the present invention is to provide a method for producing acetoin, in which a single colony of the above-mentioned engineering strain is inoculated into an LB medium for activation, and then inserted into a fermentation medium for fermentation after activation.
在本发明的一种实施方式中,所述发酵的培养基包括NBS无机盐培养基。In one embodiment of the present invention, the fermentation medium comprises NBS inorganic salt medium.
在本发明的一种实施方式中,所述发酵的条件为35-38℃,200-220rpm,活化后的初始OD600为0.04-0.1,发酵70-75h;或,发酵条件为35-38℃,480-530rpm,活化后的初始OD600为0.04-0.1,接种量为5-10%,装液量为40%-60%,初始pH为6.0-7.0,通气量为1-2vvm,发酵70-80h。In one embodiment of the present invention, the fermentation conditions are 35-38°C, 200-220rpm, the initial OD 600 after activation is 0.04-0.1, and the fermentation conditions are 70-75h; or, the fermentation conditions are 35-38°C , 480-530rpm, the initial OD 600 after activation is 0.04-0.1, the inoculum volume is 5-10%, the liquid filling volume is 40%-60%, the initial pH is 6.0-7.0, the aeration volume is 1-2vvm, fermentation 70 -80h.
本发明的第四个目的是提供上述的工程菌株在制备靶蛋白或在生物、制药、食品或化工领域内的应用。The fourth object of the present invention is to provide the application of the above-mentioned engineered strain in the preparation of target protein or in the fields of biology, pharmacy, food or chemical industry.
本发明的第五个目的是提供上述的一种调控靶蛋白基因表达的方法或上述的一种生产乙偶姻的方法在制备目的蛋白或生物、制药、食品或化工领域内的应用。The fifth object of the present invention is to provide the application of the above-mentioned method for regulating the expression of target protein gene or the above-mentioned method for producing acetoin in the preparation of target protein or in the fields of biology, pharmacy, food or chemical industry.
在本发明的一种实施方式中,所述靶蛋白包括酶蛋白或非酶蛋白。In one embodiment of the present invention, the target protein includes an enzymatic protein or a non-enzymatic protein.
本发明的有益效果:Beneficial effects of the present invention:
本发明提供了一种构建动态调控基因线路的方法,具有发酵过程外源添加诱导剂的优势和时空特异性。此外,该方法设计简单,系统元件少,菌株生长负荷低。通过构建乙偶姻生产工程菌株和引入动态调控基因线路,在无机盐培养基中可以积累高达66.8g/L的乙偶姻,其转化率达0.58g/g葡萄糖,具有较好的应用前景。The invention provides a method for constructing a dynamic regulation gene circuit, which has the advantage of adding an inducer exogenously in the fermentation process and has space-time specificity. In addition, the method is simple in design, with few system components and low strain growth load. By constructing acetoin production engineering strains and introducing dynamic regulation gene circuit, acetoin can accumulate up to 66.8g/L in inorganic salt medium, and its conversion rate can reach 0.58g/g glucose, which has a good application prospect.
附图说明Description of drawings
图1:不同光照强度和光照周期下近红外光响应的荧光过程曲线,A:近红外光激活元件的可行性;B:近红外光激活的光照强度依赖性;C:近红外光激活过程的荧光曲线;D:近红外光激活的光照周期依赖性。Figure 1: Fluorescence process curves of NIR photoresponse under different illumination intensities and photoperiods, A: Feasibility of NIR photoactivated elements; B: Illumination intensity dependence of NIR photoactivation; C: NIR photoactivation process Fluorescence curves; D: photoperiod dependence of near-infrared light activation.
图2:质粒图谱,A:PJ23119-Y30M-PJ23119-BudAB-NOX;B:PmrkA-RpoS。Figure 2: Plasmid map, A: P J23119 -Y 30 MP J23119 -BudAB-NOX; B: P mrkA -RpoS.
图3:乙偶姻生产菌株的基因工程改造靶点。Figure 3: Genetically engineered targets of acetoin-producing strains.
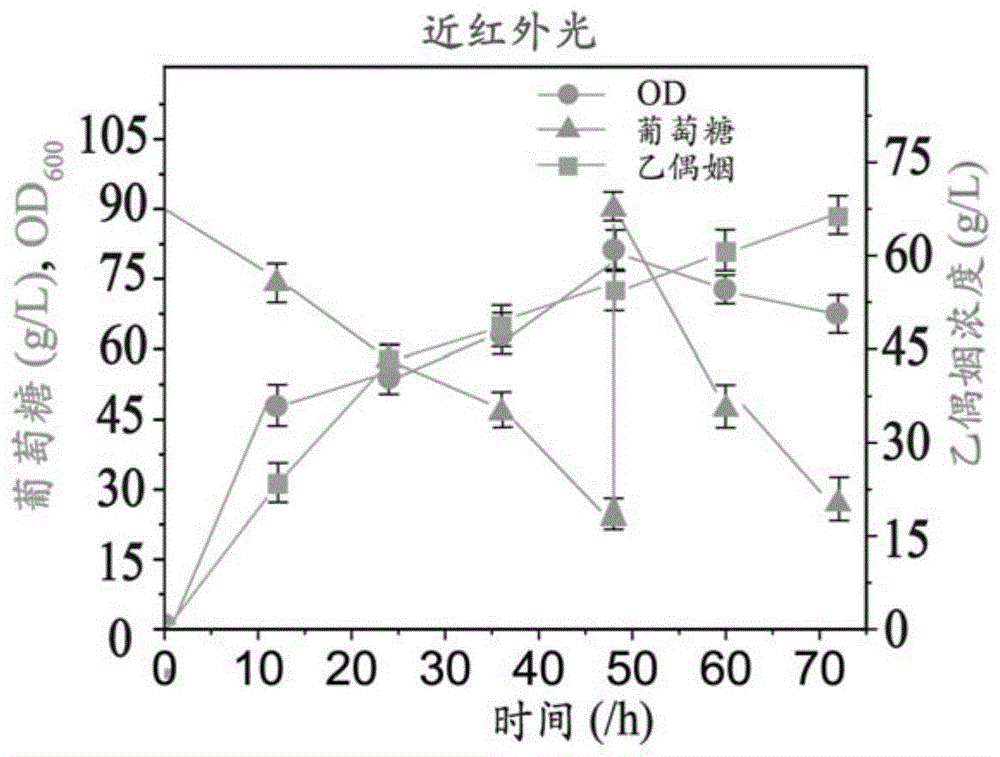
图4:乙偶姻摇瓶发酵参数。Figure 4: Acetoin shake flask fermentation parameters.
图5:乙偶姻5L发酵罐参数。Figure 5: Acetoin 5L fermenter parameters.
具体实施方式Detailed ways
材料与方法Materials and Methods
质粒构建采用经典分子生物手段进行。Plasmid construction was carried out by classical molecular biology methods.
荧光过程曲线测定采用SpectraMax M3微孔板酶标仪(Molecular Devices,Sunnyvale,CA)进行测定,控制环境温度30度。The fluorescence process curve was measured using a SpectraMax M3 microplate microplate reader (Molecular Devices, Sunnyvale, CA), and the ambient temperature was controlled at 30 degrees.
种子培养基:LB培养基,成分包含蛋白胨10g/L,酵母粉5g/L,氯化钠10g/L。Seed medium: LB medium, the ingredients include peptone 10g/L, yeast powder 5g/L, and sodium chloride 10g/L.
发酵培养基:NBS无机盐培养基,成分包含葡萄糖50g/L、CaCl2·2H2O 15mg/L、微量元素液0.667mL/L、灭菌后补加MgSO4·7H2O 0.25g/L,VB1 0.5mg/L,盐酸甜菜碱1mM。微量元素液的配置方法为FeCl3·6H2O 2.4g/L、CoCl2·6H2O 0.3g/L、CuCl2 0.15g/L、ZnCl2·4H2O0.3g/L、NaMnO4 0.3g/L、H3BO3 0.075g/L、MnCl2·4H2O 0.5g/L,溶于0.1M HCl中配制。Fermentation medium: NBS inorganic salt medium, the ingredients include glucose 50g/L, CaCl 2 ·2H 2 O 15mg/L, trace element liquid 0.667mL/L, and MgSO 4 ·7H 2 O 0.25g/L after sterilization , V B1 0.5mg/L, betaine hydrochloride 1mM. The configuration method of the trace element liquid is FeCl 3 ·6H 2 O 2.4g/L, CoCl 2 ·6H 2 O 0.3g/L, CuCl 2 0.15g/L, ZnCl 2 ·4H 2 O 0.3g/L, NaMnO 4 0.3 g/L, H 3 BO 3 0.075g/L, MnCl 2 ·4H 2 O 0.5g/L, dissolved in 0.1M HCl to prepare.
发酵样品制备:取发酵液样品,12000rpm离心5min,取上清液稀释后,过0.22μm水系膜过滤,滤液作为用来液相色谱分析。Preparation of fermentation samples: take fermentation broth samples, centrifuge at 12,000 rpm for 5 min, take the supernatant and dilute it, filter it through a 0.22 μm aqueous membrane, and use the filtrate for liquid chromatography analysis.
乙偶姻含量的测定:戴安高效液相色谱仪(配紫外可见检测器),采用伯乐AminexHPX-87H(300×7.8mm,9μm)色谱柱,流动相为浓度0.005M的H2SO4,流动相经0.22μm滤膜过滤,超声脱气,流速为0.6mL/min,柱温为35℃,在紫外检测波长210nm下检测。Determination of acetoin content: Dionex high performance liquid chromatograph (with UV-visible detector), using Biole AminexHPX-87H (300×7.8mm, 9μm) chromatographic column, the mobile phase is H 2 SO 4 with a concentration of 0.005M, The mobile phase was filtered through a 0.22 μm filter membrane, degassed by ultrasonic, the flow rate was 0.6 mL/min, the column temperature was 35 °C, and the detection was performed at an ultraviolet detection wavelength of 210 nm.
涉及的启动子、基因或相关元件等的核苷酸序列见表1。The nucleotide sequences of the involved promoters, genes or related elements are shown in Table 1.
表1核苷酸序列Table 1 Nucleotide sequences
实施例1不同光照强度和光脉冲周期下近红外光激活元件的评价Example 1 Evaluation of near-infrared light-activated elements under different light intensities and light pulse periods
将PJ23119-Y30M-PJ23119-BudAB-NOX和PmrkA-BFP质粒转入到E.coli F0601(构建方法见Dong X,Chen X,Qian Y,et al.Metabolic engineering of Escherichia coli W3110to produce L-malate[J].Biotechnology&Bioengineering,2016,114(3):656-664.)中,获得工程菌株,将其置于LB培养基中培养,使用SpectraMax M3微孔板酶标仪对上述工程菌株进行连续荧光测定。当菌株处于650nm的近红外光条件下,近红外光光敏色素(包括Bphs、Bpho、Yhjh和/或Mrkh)结合到PmrkA启动子前面,实现荧光蛋白BFP的表达。The P J23119 -Y 30 MP J23119 -BudAB-NOX and P mrkA -BFP plasmids were transferred into E. coli F0601 (see Dong X, Chen X, Qian Y, et al. Metabolic engineering of Escherichia coli W3110 to produce L- malate[J].Biotechnology&Bioengineering,2016,114(3):656-664.), obtained the engineered strain, placed it in LB medium for cultivation, and used the SpectraMax M3 microplate microplate reader to continuously carry out the above engineered strains. Fluorescence assay. When the strain is exposed to near-infrared light at 650 nm, near-infrared photophytochromes (including Bphs, Bpho, Yhjh and/or Mrkh) bind to the front of the PmrkA promoter to achieve the expression of the fluorescent protein BFP.
在黑暗或者650nm近红外光光照条件下进行实验,结果如图1所示,A图和B图表明了该近红外光系统的光照强度的剂量依赖性,当近红外光光照强度增加至0.8W/cm2时,BFP荧光蛋白表达量从142.62OD(a.u.)提高至1410.68OD(a.u.);C图表明,随着时间(0-14h)和光照强度(0.2-0.8W/cm2)的不断增加,BFP的表达量逐渐提高;D图表明,当近红外光光脉冲周期增加至100%时,mKate荧光蛋白表达量从152.69OD(a.u.)提高至1437.9OD(a.u.)。Experiments were carried out under dark or 650nm near-infrared light illumination conditions. The results are shown in Figure 1. Figures A and B show the dose dependence of the illumination intensity of the near-infrared light system. When the illumination intensity of the near-infrared light increases to 0.8W At /cm 2 , the expression of BFP fluorescent protein increased from 142.62 OD (au) to 1410.68 OD (au); Figure C shows that with time (0-14h) and light intensity (0.2-0.8W/cm 2 ) Increase, the expression of BFP gradually increased; D figure shows that when the near-infrared light pulse period increased to 100%, the mKate fluorescent protein expression increased from 152.69OD(au) to 1437.9OD(au).
实施例2动态基因线路引入乙偶姻生产菌株的摇瓶发酵性能Example 2 The shake flask fermentation performance of acetoin production strain introduced by dynamic gene line
如图3所示,以PJ23119-Y30M-PJ23119-BudAB-NOX和PmrkA-RpoS(质粒图谱见图2)为表达载体,将不同的动态基因线路引入至E.coli F0601菌株中,共获得Q1-Q7 7株工程菌株,Q1菌株表示仅表达PJ23119-Y30M-PJ23119-BudAB-NOX时获得的工程菌株,Q2菌株表示仅表达PmrkA-RpoS时获得的工程菌株,Q3-Q7分别表示近红外光光照强度分别为0.2W/cm2、0.25W/cm2、0.3W/cm2、0.4W/cm2、0.8W/cm时的工程菌株。As shown in Figure 3, using P J23119 -Y 30 MP J23119 -BudAB-NOX and P mrkA -RpoS (see Figure 2 for the plasmid map) as expression vectors, different dynamic gene lines were introduced into E.coli F0601 strain, a total of Seven engineering strains Q1-Q7 were obtained. The Q1 strain represented the engineered strain obtained when only P J23119 -Y 30 MP J23119 -BudAB-NOX was expressed, and the Q2 strain represented the engineered strain obtained when only P mrkA -RpoS was expressed. Indicates the engineering strains when the near-infrared light intensity is 0.2W/cm 2 , 0.25W/cm 2 , 0.3W/cm 2 , 0.4W/cm 2 , and 0.8W/cm , respectively.
如图4和5所示,在NBS无机盐培养基中检测菌株生长及产物合成情况。37℃,200rpm,将工程菌株单菌落在LB培养基中培养至OD600为0.05,初始接种量控制为1%,发酵70h,摇瓶发酵结果表明,Q1-Q2菌株分别表示仅表达乙偶姻生产路径、表达加速分裂基因,Q1-Q2菌株的乙偶姻产量从13.5g/L提高18.5g/L,细胞相对存活率从70.5%提高到74.6%。Q7菌株其发酵性能为乙偶姻积累量31.8g/L。As shown in Figures 4 and 5, strain growth and product synthesis were detected in NBS inorganic salt medium. 37°C, 200rpm, single colony of engineered strain was cultured in LB medium to OD 600 of 0.05, the initial inoculum was controlled to 1%, and fermented for 70h. The results of shake flask fermentation showed that strains Q1-Q2 expressed only acetoin Production path, expression of accelerated division gene, the acetoin yield of Q1-Q2 strain increased from 13.5g/L to 18.5g/L, and the relative cell survival rate increased from 70.5% to 74.6%. The fermentation performance of strain Q7 was 31.8g/L of acetoin accumulation.
实施例3动态基因线路引入乙偶姻生产菌株的发酵罐发酵性能Example 3 The fermentor fermentation performance of the acetoin-producing strain introduced into the dynamic gene line
在5L发酵罐中检测Q7菌株的发酵性能。温度恒定37℃,500rpm,将工程菌株单菌落在LB培养基中培养至OD600为0.05,初始接种量控制为5%,通气量1vvm,发酵周期为72h,装液量为3L,用4M氢氧化钠和2M盐酸控制初始pH为6.8。发酵结束时,乙偶姻积累量达66.8g/L,转化率达0.58g/g葡萄糖。The fermentation performance of the Q7 strain was tested in a 5L fermenter. The temperature was constant at 37°C, 500rpm, and the single colony of the engineered strain was cultured in LB medium to OD 600 of 0.05, the initial inoculum was controlled to 5%, the ventilation rate was 1vvm, the fermentation period was 72h, the liquid volume was 3L, and 4M hydrogen was used. Sodium oxide and 2M hydrochloric acid control the initial pH to 6.8. At the end of fermentation, the accumulation of acetoin reached 66.8 g/L, and the conversion rate reached 0.58 g/g glucose.
虽然本发明已以较佳实施例公开如上,但其并非用以限定本发明,任何熟悉此技术的人,在不脱离本发明的精神和范围内,都可做各种的改动与修饰,因此本发明的保护范围应该以权利要求书所界定的为准。Although the present invention has been disclosed above with preferred embodiments, it is not intended to limit the present invention. Anyone who is familiar with this technology can make various changes and modifications without departing from the spirit and scope of the present invention. Therefore, The protection scope of the present invention should be defined by the claims.
SEQUENCE LISTINGSEQUENCE LISTING
<110> 江南大学<110> Jiangnan University
<120> 一种近红外光控制的动态调控系统及其应用<120> A near-infrared light-controlled dynamic control system and its application
<160> 18<160> 18
<170> PatentIn version 3.3<170> PatentIn version 3.3
<210> 1<210> 1
<211> 2061<211> 2061
<212> DNA<212> DNA
<213> 人工合成<213> Synthetic
<400> 1<400> 1
atggcccgtg gttgtttaat gaccattagc ggcggcacct ttgacccgag catttgcgag 60atggcccgtg gttgtttaat gaccattagc ggcggcacct ttgacccgag catttgcgag 60
atggaaccga tcgccacccc cggtgccatt cagcctcatg gtgctttaat gacagcccgc 120atggaaccga tcgccacccc cggtgccatt cagcctcatg gtgctttaat gacagcccgc 120
gcagatagtg gccgtgtggc ccacgcaagt gtgaatctgg gcgagatttt aggtttaccc 180gcagatagtg gccgtgtggc ccacgcaagt gtgaatctgg gcgagatttt aggtttaccc 180
gctgccagtg tgttaggcgc cccgatcggt gaagttattg gccgcgtgaa cgaaatttta 240gctgccagtg tgttaggcgc cccgatcggt gaagttatg gccgcgtgaa cgaaatttta 240
ctgcgtgagg cccgccgcag cggtagtgaa acccccgaaa ccatcggtag ctttcgccgc 300ctgcgtgagg cccgccgcag cggtagtgaa acccccgaaa ccatcggtag ctttcgccgc 300
agcgacggtc agttactgca tctgcacgca ttccagagcg gtgattacat gtgtttagac 360agcgacggtc agttactgca tctgcacgca ttccagagcg gtgattacat gtgtttagac 360
attgagccgg tgcgtgatga ggatggtcgc ttacctccgg gcgcacgcca gagtgtgatc 420attgagccgg tgcgtgatga ggatggtcgc ttacctccgg gcgcacgcca gagtgtgatc 420
gaaaccttta gcagcgccat gacccaagtg gaactgtgtg aactggcagt gcacggttta 480gaaaccttta gcagcgccat gacccaagtg gaactgtgtg aactggcagt gcacggttta 480
caactggttt taggctacga ccgcgtgatg gcataccgtt ttggcgcaga tggtcatggc 540caactggttt taggctacga ccgcgtgatg gcataccgtt ttggcgcaga tggtcatggc 540
gaggtgatcg cagaacgtcg ccgccaagat ctggaaccgt atctgggttt acattatccg 600gaggtgatcg cagaacgtcg ccgccaagat ctggaaccgt atctgggttt acattatccg 600
gcaagtgaca tccctcagat cgcccgcgca ctgtatttac gtcagcgcgt tggcgcaatt 660gcaagtgaca tccctcagat cgcccgcgca ctgtatttac gtcagcgcgt tggcgcaatt 660
gcagatgcat gctatcgccc ggttccgctg ctgggtcatc cggaactgga tgatggtaag 720gcagatgcat gctatcgccc ggttccgctg ctgggtcatc cggaactgga tgatggtaag 720
ccgctggatc tgacccatag ctctttacgc agcgttagcc cggtgcatct ggactatatg 780ccgctggatc tgacccatag ctctttacgc agcgttagcc cggtgcatct ggactatatg 780
cagaatatga acaccgcagc aagcttaacc atcggtttag ccgatggcga ccgtctgtgg 840cagaatatga acaccgcagc aagcttaacc atcggtttag ccgatggcga ccgtctgtgg 840
ggtatgctgg tgtgccataa caccacaccg cgcattgccg gtccggaatg gcgtgcagca 900ggtatgctgg tgtgccataa caccacaccg cgcattgccg gtccggaatg gcgtgcagca 900
gccggtatga ttggccaagt tgtgtcttta ctgttaagtc gtttaggtga agtggaaaac 960gccggtatga ttggccaagt tgtgtcttta ctgttaagtc gtttaggtga agtggaaaac 960
gccgccgaga ctttagcccg tcaaagtact ttaagcactt tagttgaacg tctgagcact 1020gccgccgaga ctttagcccg tcaaagtact ttaagcactt tagttgaacg tctgagcact 1020
ggtgatactt tagccgcagc ctttgtggcc gcagatcaac tgattttaga tttagttggt 1080ggtgatactt tagccgcagc ctttgtggcc gcagatcaac tgattttaga tttagttggt 1080
gccagcgcag cagttgttcg tttagctggt caagaactgc attttggtcg tacaccgccg 1140gccagcgcag cagttgttcg tttagctggt caagaactgc attttggtcg tacaccgccg 1140
gtggacgcca tgcagaaggt tctggattct ttaggtcgtc cgagcccgtt agaggtgctg 1200gtggacgcca tgcagaaggt tctggattct ttaggtcgtc cgagcccgtt agaggtgctg 1200
agtctggacg atgttacctt acgccatccc gaactgccgg aactgctggc agctggtagc 1260agtctggacg atgttacctt acgccatccc gaactgccgg aactgctggc agctggtagc 1260
ggcattttac tgttaccgtt aaccagcggt gatggcgatc tgattgcttg gttccgcccc 1320ggcattttac tgttaccgtt aaccagcggt gatggcgatc tgattgcttg gttccgcccc 1320
gaacatgtgc agacaattac ttggggtggc aatccggcag aacacggcac ttggaacccg 1380gaacatgtgc agacaattac ttggggtggc aatccggcag aacacggcac ttggaacccg 1380
gcaacccagc gtatgcgtcc tcgcgcaagc tttgacgcat ggaaagaaac cgtgaccggt 1440gcaacccagc gtatgcgtcc tcgcgcaagc tttgacgcat ggaaagaaac cgtgaccggt 1440
cgctctttac cgtggacaag tgcagaacgc aattgcgccc gcgagttagg tgaagccatt 1500cgctctttac cgtggacaag tgcagaacgc aattgcgccc gcgagttagg tgaagccatt 1500
gccgccgaga tggcacagcg cacccgcgca gaagagctgg aacgcgttgc catggttgac 1560gccgccgaga tggcacagcg cacccgcgca gaagagctgg aacgcgttgc catggttgac 1560
tctttaacac gtttatggaa tcgtctgggt attgagacac tgctgaaacg cgagtgggaa 1620tctttaacac gtttatggaa tcgtctgggt attgagacac tgctgaaacg cgagtgggaa 1620
tatgccaccc gcaagaacag tccgattagc atcgtgatga ttgacttcga taatttcaaa 1680tatgccaccc gcaagaacag tccgattagc atcgtgatga ttgacttcga taatttcaaa 1680
caaatcaatg atcaacacgg ccatttagtg ggtgatgaag tgctgcaagg tagtgcacgt 1740caaatcaatg atcaacacgg ccatttagtg ggtgatgaag tgctgcaagg tagtgcacgt 1740
ctgattatta gtgtgctggc cagctatgat atcttaggtc gctggggcgg tgatgaattc 1800ctgattatta gtgtgctggc cagctatgat atcttaggtc gctggggcgg tgatgaattc 1800
atgctgattt taccgggtag tggtcgtgaa cagaccgcag ttctgctgga gcgcattcaa 1860atgctgattt taccgggtag tggtcgtgaa cagaccgcag ttctgctgga gcgcattcaa 1860
gctacaattg cacagaatcc ggtgccgaca agtgctggtc cgatggccat ctctttaagc 1920gctacaattg cacagaatcc ggtgccgaca agtgctggtc cgatggccat ctctttaagc 1920
atgggcggcg tgagtgtttt tacaaaccaa ggtgaagctt tacagtattg ggtggaacaa 1980atgggcggcg tgagtgtttt tacaaaccaa ggtgaagctt tacagtattg ggtggaacaa 1980
gctgataacc agctgatgaa agtgaagcgt ttaggcaaag gtaacttcca gctggccgag 2040gctgataacc agctgatgaa agtgaagcgt ttaggcaaag gtaacttcca gctggccgag 2040
tatcatcacc atcaccatca t 2061tatcatcacc atcaccatca t 2061
<210> 2<210> 2
<211> 594<211> 594
<212> DNA<212> DNA
<213> 人工合成<213> Synthetic
<400> 2<400> 2
atgccgctga gccgcgatct ccgcgaaaag accggtatgc tgcataatcg tgcggaaacg 60atgccgctga gccgcgatct ccgcgaaaag accggtatgc tgcataatcg tgcggaaacg 60
ctgctgggtc tgccgagcgg tatcatgggt tgggccgatt acgtggattg gctgcgccat 120ctgctgggtc tgccgagcgg tatcatgggt tgggccgatt acgtggattg gctgcgccat 120
tttctggcgc tgtacgatcc gatcgaacgc cgcatcgttg ccttcggtgg ctggagcggt 180tttctggcgc tgtacgatcc gatcgaacgc cgcatcgttg ccttcggtgg ctggagcggt 180
ctggccagct ttgatccgga tccgggccac agccgtcgtc tgatccaaga tctccatgcg 240ctggccagct ttgatccgga tccgggccac agccgtcgtc tgatccaaga tctccatgcg 240
ctgggcattg acaccgaccg tattccacgt gcgccggcgg aatactgccc accactgacg 300ctgggcattg acaccgaccg tattccacgt gcgccggcgg aatactgccc accactgacg 300
aattttgcgc gcgcgctggg tgcccgttat gttctggaag gtagcgccct cggtggccgt 360aattttgcgc gcgcgctggg tgcccgttat gttctggaag gtagcgccct cggtggccgt 360
gtgattctgc atcatctgaa gaaacgcatc ggcgatgaaa tcggcaacgc gaccgcgttc 420gtgattctgc atcatctgaa gaaacgcatc ggcgatgaaa tcggcaacgc gaccgcgttc 420
tttggtggcc caagccacgg taccgcgacc cattggcgtg cctttcaagc cgcgctggat 480tttggtggcc caagccacgg taccgcgacc cattggcgtg cctttcaagc cgcgctggat 480
cgttttggtg cggcgcatcc agataaacgt gccgatgttc tggcgggtgc ggccgccacc 540cgttttggtg cggcgcatcc agataaacgt gccgatgttc tggcgggtgc ggccgccacc 540
tttacggcgc tgctggaatg gttcacgcca ttcgttgcgg cccgccgtgt gtaa 594tttacggcgc tgctggaatg gttcacgcca ttcgttgcgg cccgccgtgt gtaa 594
<210> 3<210> 3
<211> 765<211> 765
<212> DNA<212> DNA
<213> 人工合成<213> Synthetic
<400> 3<400> 3
atgattcgcc aagttattca gcgcatcagc aacccggaag ccagcatcga aagtctgcaa 60atgattcgcc aagttattca gcgcatcagc aacccggaag ccagcatcga aagtctgcaa 60
gaacgtcgtt tttggctgca gtgcgagcgc gcctatactt ggcaaccgat ttatcagact 120gaacgtcgtt tttggctgca gtgcgagcgc gcctatactt ggcaaccgat ttatcagact 120
tgtggccgtt taatggcagt tgaactgctg accgtggtga cccatccgct gaatccgagt 180tgtggccgtt taatggcagt tgaactgctg accgtggtga cccatccgct gaatccgagt 180
cagcgtttac cgcccgatcg ctactttacc gaaatcaccg tgagccatcg catggaagtg 240cagcgtttac cgcccgatcg ctactttacc gaaatcaccg tgagccatcg catggaagtg 240
gtgaaggaac agatcgatct gctggcccag aaagccgatt ttttcatcga gcacggttta 300gtgaaggaac agatcgatct gctggcccag aaagccgatt ttttcatcga gcacggttta 300
ttagcaagcg tgaacattga cggcccgact ttaattgcac tgcgccagca accgaaaatt 360ttagcaagcg tgaacattga cggcccgact ttaattgcac tgcgccagca accgaaaatt 360
ctgcgccaga tcgaacgttt accgtggtta cgcttcgaac tggttgagca catccgtctg 420ctgcgccaga tcgaacgttt accgtggtta cgcttcgaac tggttgagca catccgtctg 420
ccgaaagaca gcaccttcgc aagcatgtgt gagttcggcc cgctgtggct ggatgatttt 480ccgaaagaca gcaccttcgc aagcatgtgt gagttcggcc cgctgtggct ggatgatttt 480
ggcaccggca tggccaattt tagcgcactg agcgaggtgc gctacgacta tatcaagatc 540ggcaccggca tggccaattt tagcgcactg agcgaggtgc gctacgacta tatcaagatc 540
gcacgcgagc tgttcgtgat gctgcgtcag agcccggaag gtcgcacttt atttagccag 600gcacgcgagc tgttcgtgat gctgcgtcag agcccggaag gtcgcacttt atttagccag 600
ctgctgcatc tgatgaaccg ctattgtcgc ggtgtgattg tggaaggtgt ggaaaccccg 660ctgctgcatc tgatgaaccg ctattgtcgc ggtgtgattg tggaaggtgt ggaaaccccg 660
gaagaatggc gtgacgtgca gaatagcccg gcctttgcag cacaaggttg gtttctgagc 720gaagaatggc gtgacgtgca gaatagcccg gcctttgcag cacaaggttg gtttctgagc 720
cgtcccgctc cgatcgaaac tttaaatacc gcagttctgg cactg 765cgtcccgctc cgatcgaaac tttaaatacc gcagttctgg cactg 765
<210> 4<210> 4
<211> 702<211> 702
<212> DNA<212> DNA
<213> 人工合成<213> Synthetic
<400> 4<400> 4
atgaccgagg gtacgatcaa gaccagcaag tacgagatca tcgccatctt ccgcgaagaa 60atgaccgagg gtacgatcaa gaccagcaag tacgagatca tcgccatctt ccgcgaagaa 60
ctccgtaagc gcaccgagat cgagatcttc ttcaacaaca cgagcatcat tacccagctg 120ctccgtaagc gcaccgagat cgagatcttc ttcaacaaca cgagcatcat tacccagctg 120
acccgcgttg attttgcgga gttccacatc cagacccacc gcaaaatccc gagcggtcac 180acccgcgttg attttgcgga gttccacatc cagacccacc gcaaaatccc gagcggtcac 180
aaaatccgct ttctgctgca cagcgatagc ggtaagatcg agttcaacgc cgccctcacg 240aaaatccgct ttctgctgca cagcgatagc ggtaagatcg agttcaacgc cgccctcacg 240
aaacatgaca acagcggcgt tgacaaaggc atccgctacg ccttcagtct gccggaatgt 300aaacatgaca acagcggcgt tgacaaaggc atccgctacg ccttcagtct gccggaatgt 300
ctgcaagttg ttcagcgtcg ccgtgatccg cgctttcgtc tgcgtcacga gcacgatttc 360ctgcaagttg ttcagcgtcg ccgtgatccg cgctttcgtc tgcgtcacga gcacgatttc 360
tactgtcgtg gccgccacaa gaacggcgaa aactatctgt tcgatatcaa ggatattagc 420tactgtcgtg gccgccacaa gaacggcgaa aactatctgt tcgatatcaa ggatattagc 420
gacggcggct gtgcgctgat gaccaagacg ccgaatctga agtttctgag ccacaatgcg 480gacggcggct gtgcgctgat gaccaagacg ccgaatctga agtttctgag ccacaatgcg 480
ctgctgaaaa acgcggttct gatgctggcc gagtacggtg agatcaccat cgatctggtg 540ctgctgaaaa acgcggttct gatgctggcc gagtacggtg agatcaccat cgatctggtg 540
gtgaagaacg tgatcgtgat cacgctggac aacgccaacg aagagagcga gagttactac 600gtgaagaacg tgatcgtgat cacgctggac aacgccaacg aagagagcga gagttactac 600
cagatcagct gccagttcaa gttccgccat ctcgacgatc agcgccgcat tgagaagatt 660cagatcagct gccagttcaa gttccgccat ctcgacgatc agcgccgcat tgagaagatt 660
ctgctggatc tgattctgga agcgaagcgc aaaaagcgta tt 702ctgctggatc tgattctgga agcgaagcgc aaaaagcgta tt 702
<210> 5<210> 5
<211> 140<211> 140
<212> DNA<212> DNA
<213> 人工合成<213> Synthetic
<400> 5<400> 5
gctgcgctgt aaacaaccac cctcgcgttt tcatctatca atggctgttt attaatagtc 60gctgcgctgt aaacaaccac cctcgcgttt tcatctatca atggctgttt attaatagtc 60
gatggttatc tgttatataa cttaatgaaa cgtgaacaaa tgtatatttg tcggcgaata 120gatggttatc tgttatataa cttaatgaaa cgtgaacaaa tgtatatttg tcggcgaata 120
aatagcattc tttgacgccg 140aatagcattc tttgacgccg 140
<210> 6<210> 6
<211> 702<211> 702
<212> DNA<212> DNA
<213> 人工合成<213> Synthetic
<400> 6<400> 6
atgagcgagc tgattaagga gaacatgcac atgaagctgt acatggaggg caccgtggac 60atgagcgagc tgattaagga gaacatgcac atgaagctgt acatggaggg caccgtggac 60
aaccatcact tcaagtgcac atccgagggc gaaggcaagc cctacgaggg cacccagacc 120aaccatcact tcaagtgcac atccgagggc gaaggcaagc cctacgaggg cacccagacc 120
atgagaatca aggtggtcga gggcggccct ctccccttcg ccttcgacat cctggctact 180atgagaatca aggtggtcga gggcggccct ctccccttcg ccttcgacat cctggctact 180
agcttcctct acggcagcaa gaccttcatc gaccacaccc agggcatccc cgacttcttc 240agcttcctct acggcagcaa gaccttcatc gaccacaccc agggcatccc cgacttcttc 240
aagcagtcct tccctgaggg cttcacatgg gagagagtca ccacatacga agacgggggc 300aagcagtcct tccctgaggg cttcacatgg gagagagtca ccacatacga agacgggggc 300
gtgctgaccg ctacccagga caccagcctc caggacggct gcctcatcta caacgtcaag 360gtgctgaccg ctacccagga caccagcctc caggacggct gcctcatcta caacgtcaag 360
atcagagggg tgaacttcac atccaacggc cctgtgatgc agaagaaaac actcggctgg 420atcagagggg tgaacttcac atccaacggc cctgtgatgc agaagaaaac actcggctgg 420
gaggccttca ccgagacgct gtaccccgct gacggcggcc tggaaggcag aaacgacatg 480gaggccttca ccgagacgct gtaccccgct gacggcggcc tggaaggcag aaacgacatg 480
gccctgaagc tcgtgggcgg gagccatctg atcgcaaaca tcaagaccac atatagatcc 540gccctgaagc tcgtgggcgg gagccatctg atcgcaaaca tcaagaccac atatagatcc 540
aagaaacccg ctaagaacct caagatgcct ggcgtctact atgtggacta cagactggaa 600aagaaacccg ctaagaacct caagatgcct ggcgtctact atgtggacta cagactggaa 600
agaatcaagg aggccaacaa cgagacctac gtcgagcagc acgaggtggc agtggccaga 660agaatcaagg aggccaacaa cgagacctac gtcgagcagc acgaggtggc agtggccaga 660
tactgcgacc tccctagcaa actggggcac aagctcaatt aa 702tactgcgacc tccctagcaa actggggcac aagctcaatt aa 702
<210> 7<210> 7
<211> 12<211> 12
<212> DNA<212> DNA
<213> 人工合成<213> Synthetic
<400> 7<400> 7
aaagaggaga aa 12
<210> 8<210> 8
<211> 16<211> 16
<212> DNA<212> DNA
<213> 人工合成<213> Synthetic
<400> 8<400> 8
gattaaagag gagaaa 16
<210> 9<210> 9
<211> 780<211> 780
<212> DNA<212> DNA
<213> 人工合成<213> Synthetic
<400> 9<400> 9
atgaacgaaa aacacgggtg ttcctgtgcg cgccatttgg cgcagggttt tgccaggcag 60atgaacgaaa aacacgggtg ttcctgtgcg cgccatttgg cgcagggttt tgccaggcag 60
tcgatcaacg ccggggaggg cgaaatctat caaatctcgc tgatgagcgc gctcatcgac 120tcgatcaacg ccggggaggg cgaaatctat caaatctcgc tgatgagcgc gctcatcgac 120
ggggtctacg agggggaaac caccattgcc gagttgctca aacacggcaa ctttggcctc 180ggggtctacg agggggaaac caccattgcc gagttgctca aacacggcaa ctttggcctc 180
ggtaccttca atcatctgga cggtgaactg atcgctttcg accaggaaat acaccagctg 240ggtaccttca atcatctgga cggtgaactg atcgctttcg accaggaaat acaccagctg 240
cgcgccgacg gcagcgcccg cccggccggc ctgcaacaga aaaccccctt cgccgtcgtc 300cgcgccgacg gcagcgcccg cccggccggc ctgcaacaga aaaccccctt cgccgtcgtc 300
acctttttcc agcccagcgt cagccagcaa ttcgatcggc cgatcaccaa ggcgcagctg 360acctttttcc agcccagcgt cagccagcaa ttcgatcggc cgatcaccaa ggcgcagctg 360
caccaatgca tcgatgaaca ggtcgcctcg ccgaacctgt tctgtgcggt gcgggtcgac 420caccaatgca tcgatgaaca ggtcgcctcg ccgaacctgt tctgtgcggt gcgggtcgac 420
ggcgagttca gccacgtgga aacccgcacc gtgccgcgcc aggagcggcc ttaccgcccg 480ggcgagttca gccacgtgga aacccgcacc gtgccgcgcc aggagcggcc ttaccgcccg 480
atgctggaag cgatagaaga gcagccgacc ttctcgttcc atcagcggcg cggcacgctg 540atgctggaag cgatagaaga gcagccgacc ttctcgttcc atcagcggcg cggcacgctg 540
gtcggcttcc gctcgccgga ttacatgcag ggcatcggcg tggccggcta tcacgaacac 600gtcggcttcc gctcgccgga ttacatgcag ggcatcggcg tggccggcta tcacgaacac 600
ttcgttaccg acgaccgcag cggcggcggc cacgtgctgg actaccagct cgatcatggc 660ttcgttaccg acgaccgcag cggcggcggc cacgtgctgg actaccagct cgatcatggc 660
cgtctgcagt tcggcgtcat cacgcgtctc aatcttcagt taccgcatga tgcggatttc 720cgtctgcagt tcggcgtcat cacgcgtctc aatcttcagt taccgcatga tgcggatttc 720
ttgcgcgcca acctctgccc agaggatttg gaccgcgcga ttcgttccgc cgagggctaa 780ttgcgcgcca acctctgccc agaggatttg gaccgcgcga ttcgttccgc cgagggctaa 780
<210> 10<210> 10
<211> 1686<211> 1686
<212> DNA<212> DNA
<213> 人工合成<213> Synthetic
<400> 10<400> 10
atggcacagg aaaaaacagg caatgactgg caatgcggcg ccgatttggt ggtgaaaaac 60atggcacagg aaaaaacagg caatgactgg caatgcggcg ccgatttggt ggtgaaaaac 60
ctggaagcgc agggcgtcaa acacgttttc ggtattccgg gcgccaagat cgaccgggtg 120ctggaagcgc agggcgtcaa acacgttttc ggtattccgg gcgccaagat cgaccgggtg 120
ttcgactcgc tggaggacgc gccgtcgatc gaaacggtgg tggtgcgcca tgaggccaac 180ttcgactcgc tggaggacgc gccgtcgatc gaaacggtgg tggtgcgcca tgaggccaac 180
gcggccttta tggcggcggc ggtcggccgc ttgaccggca aggccggggt ggcgctggtc 240gcggccttta tggcggcggc ggtcggccgc ttgaccggca aggccggggt ggcgctggtc 240
acctccgggc cgggcagctc caacctgatc accgggctgg ctaccgccac ctcggaaggg 300acctccgggc cgggcagctc caacctgatc accgggctgg ctaccgccac ctcggaaggg 300
gacgcggtgg tggccttcgg cggggcggtg aagcgcgccg acagcctgaa gcagacgcac 360gacgcggtgg tggccttcgg cggggcggtg aagcgcgccg acagcctgaa gcagacgcac 360
cagagcatgg ataccgtcag catgttccgg ccggtcacca agtattgcgc cgaagtgcat 420cagagcatgg ataccgtcag catgttccgg ccggtcacca agtattgcgc cgaagtgcat 420
gccggttcgg cgatttccga ggtgatcgcc aacgccttcc gccgcgccga gttcggccgg 480gccggttcgg cgatttccga ggtgatcgcc aacgccttcc gccgcgccga gttcggccgg 480
ccgggggcat cgttcgtcag cctgccgatg gatatcgtca atgaaccggt cagcgcgccg 540ccgggggcat cgttcgtcag cctgccgatg gatatcgtca atgaaccggt cagcgcgccg 540
gtactggcgg gctgccgctt gccgcgcatg ggggccgccg cggcagacga cattcaggcg 600gtactggcgg gctgccgctt gccgcgcatg ggggccgccg cggcagacga cattcaggcg 600
gcggtgaaac tgatccgcca ggccaaatgc ccggtgctgc tgctgggctt gcaggccagc 660gcggtgaaac tgatccgcca ggccaaatgc ccggtgctgc tgctgggctt gcaggccagc 660
cggccggaga acagcgaggc ggtgcgccat ctgctgtacc gcacccatat gccggtggtc 720cggccggaga acagcgaggc ggtgcgccat ctgctgtacc gcacccatat gccggtggtc 720
ggcacctatc aggcggcggg ggtgatcgac gtcaaccact tcgcccgttt cgccggccgc 780ggcacctatc aggcggcggg ggtgatcgac gtcaaccact tcgcccgttt cgccggccgc 780
gtcggcctgt tcaacaacca gccggccgat cagctgctgc agaaggccga tctggtggtg 840gtcggcctgt tcaacaacca gccggccgat cagctgctgc agaaggccga tctggtggtg 840
agcgtcggct atgatccgat cgaatacgat ccctgcatgt ggaacagcca cgggcgactg 900agcgtcggct atgatccgat cgaatacgat ccctgcatgt ggaacagcca cgggcgactg 900
aagctggtgc atctcgacgt gctgccggcg gatatcgata cctgctatcg gccggacgtc 960aagctggtgc atctcgacgt gctgccggcg gatatcgata cctgctatcg gccggacgtc 960
gagctggtgg gcaatatcag cgccacgctg aacatgatga ccgaagcatt cagtgaggcg 1020gagctggtgg gcaatatcag cgccacgctg aacatgatga ccgaagcatt cagtgaggcg 1020
gtctgcgtgc cgccggaggt ggagctgatc ctctccgatc tcggccgcca gcgcactgag 1080gtctgcgtgc cgccggaggt ggagctgatc ctctccgatc tcggccgcca gcgcactgag 1080
ctggcggagc gcgccgcccg ccgcggcggc atgccgatcc acccgctgcg catcgtcaag 1140ctggcggagc gcgccgcccg ccgcggcggc atgccgatcc acccgctgcg catcgtcaag 1140
gagctgcagg acatcgtcag cgacgacgtg acgctgtgtg tggacatggg cagtttccac 1200gagctgcagg acatcgtcag cgacgacgtg acgctgtgtg tggacatggg cagtttccac 1200
atctggatcg cccgttatct ttacagcttc cgtgcccgtc agctgttgat ctccaacggc 1260atctggatcg ccccgttatct ttacagcttc cgtgcccgtc agctgttgat ctccaacggc 1260
cagcaaacca tgggggtggc attgccgtgg gccatcggtg cggcgctggt gcgccccggc 1320cagcaaacca tgggggtggc attgccgtgg gccatcggtg cggcgctggt gcgccccggc 1320
gacaaggtgg tgtctatatc cggtgacggc ggcttcatgc agtcgagcat ggagctggag 1380gacaaggtgg tgtctatatc cggtgacggc ggcttcatgc agtcgagcat ggagctggag 1380
accgcggtgc ggctgaaaaa caacatcgtg cacgtgatct gggtcgataa cgcctacaac 1440accgcggtgc ggctgaaaaa caacatcgtg cacgtgatct gggtcgataa cgcctacaac 1440
atggtggaaa tgcaggaagt gaacaaatac cagcgcaagt ccggcgtgga gttcgggccg 1500atggtggaaa tgcaggaagt gaacaaatac cagcgcaagt ccggcgtgga gttcgggccg 1500
attgacttca aggcctacgc ggaatcctgc ggcgcggtcg gtttcgccgt gcagtcggtg 1560attgacttca aggcctacgc ggaatcctgc ggcgcggtcg gtttcgccgt gcagtcggtg 1560
gaagatctgc ggccgatgct gcgcaaggcg atggcgatcc aggggccggt agtggtggcc 1620gaagatctgc ggccgatgct gcgcaaggcg atggcgatcc aggggccggt agtggtggcc 1620
atcccggtcg actatgccga caactataag ctgatggcgc agatgaactt cagccagatg 1680atcccggtcg actatgccga caactataag ctgatggcgc agatgaactt cagccagatg 1680
atttaa 1686atttaa 1686
<210> 11<210> 11
<211> 29<211> 29
<212> DNA<212> DNA
<213> 人工合成<213> Synthetic
<400> 11<400> 11
ttgacagcta gctcagtcct aggtataat 29ttgacagcta gctcagtcct aggtataat 29
<210> 12<210> 12
<211> 1392<211> 1392
<212> DNA<212> DNA
<213> 人工合成<213> Synthetic
<400> 12<400> 12
atgaaaatca ttagcattaa attcgtgctc ggcggcaaca tcatgaaggt gaccgtggtt 60atgaaaatca ttagcattaa attcgtgctc ggcggcaaca tcatgaaggt gaccgtggtt 60
ggctgtaccc atgccggcac cttcgcgatc aagcagattc tggcggaaca cccagacgcc 120ggctgtaccc atgccggcac cttcgcgatc aagcagattc tggcggaaca cccagacgcc 120
gaggtgacgg tttacgagcg caacgacgtg atcagctttc tcagctgtgg catcgcgctg 180gaggtgacgg tttacgagcg caacgacgtg atcagctttc tcagctgtgg catcgcgctg 180
tatctgggtg gtaaagtggc cgacccacaa ggtctgtttt acagcagccc ggaagaactg 240tatctgggtg gtaaagtggc cgacccacaa ggtctgtttt acagcagccc ggaagaactg 240
caaaagctgg gcgcgaacgt gcagatgaac cataacgttc tggccatcga tccggaccag 300caaaagctgg gcgcgaacgt gcagatgaac cataacgttc tggccatcga tccggaccag 300
aagaccgtga ccgtggagga tctgaccagc catgcgcaga ccacggagag ctacgacaag 360aagaccgtga ccgtggagga tctgaccagc catgcgcaga ccacggagag ctacgacaag 360
ctcgttatga ccagcggtag ctggccaatc gtgccaaaga tcccgggcat tgacagcgac 420ctcgttatga ccagcggtag ctggccaatc gtgccaaaga tcccgggcat tgacagcgac 420
cgcgttaaac tgtgcaagaa ctgggcccat gcgcaagcgc tgatcgagga tgccaaggaa 480cgcgttaaac tgtgcaagaa ctgggcccat gcgcaagcgc tgatcgagga tgccaaggaa 480
gcgaaacgca tcacggtgat cggtgcgggt tacattggcg ccgaactggc cgaggcctat 540gcgaaacgca tcacggtgat cggtgcgggt tacattggcg ccgaactggc cgaggcctat 540
agcaccaccg gccatgacgt gaccctcatc gacgccatgg atcgtgtgat gccgaagtac 600agcaccaccg gccatgacgt gaccctcatc gacgccatgg atcgtgtgat gccgaagtac 600
ttcgacgccg acttcaccga cgttatcgaa caagattatc gcgaccatgg tgtgcagctc 660ttcgacgccg acttcaccga cgttatcgaa caagattatc gcgaccatgg tgtgcagctc 660
gcgctgagcg aaaccgtgga aagctttacg gacagcgcca ccggcctcac cattaagacc 720gcgctgagcg aaaccgtgga aagctttacg gacagcgcca ccggcctcac cattaagacc 720
gataagaaca gctatgagac cgatctggcg attctgtgca ttggcttccg tccgaatacc 780gataagaaca gctatgagac cgatctggcg attctgtgca ttggcttccg tccgaatacc 780
gatctgctga aaggcaaagt ggatatggcg ccaaacggcg cgatcatcac cgatgactac 840gatctgctga aaggcaaagt ggatatggcg ccaaacggcg cgatcatcac cgatgactac 840
atgcgcagca gcaacccgga catctttgcc gccggcgata gcgccgccgt gcattacaac 900atgcgcagca gcaacccgga catctttgcc gccggcgata gcgccgccgt gcattacaac 900
ccgacgcatc agaacgccta tatcccactg gccaccaatg ccgttcgcca aggcatcctc 960ccgacgcatc agaacgccta tatcccactg gccaccaatg ccgttcgcca aggcatcctc 960
gtgggcaaaa atctggttaa gccgacggtg aagtacatgg gcacgcagag cagcagtggt 1020gtgggcaaaa atctggttaa gccgacggtg aagtacatgg gcacgcagag cagcagtggt 1020
ctggcgctct acgatcgtac catcgttagt accggtctga cgctggcggc ggcgaaacag 1080ctggcgctct acgatcgtac catcgttagt accggtctga cgctggcggc ggcgaaacag 1080
caaggcgtga atgcggaaca agttatcgtg gaagacaact accgcccgga gttcatgcca 1140caaggcgtga atgcggaaca agttatcgtg gaagacaact accgcccgga gttcatgcca 1140
agcacggaac cagtgctgat gagtctggtg ttcgatccag acacccatcg cattctgggc 1200agcacggaac cagtgctgat gagtctggtg ttcgatccag acacccatcg cattctgggc 1200
ggtgcgctga tgagtaaata cgacgtgagc cagagcgcga atacgctgag tgtgtgcatc 1260ggtgcgctga tgagtaaata cgacgtgagc cagagcgcga atacgctgag tgtgtgcatc 1260
cagaacgaga atacgattga cgatctggcc atggtggata tgctgttcca gccgaacttc 1320cagaacgaga atacgattga cgatctggcc atggtggata tgctgttcca gccgaacttc 1320
gaccgcccgt tcaactatct gaacattctg gcgcaagccg cgcaagccaa agttgcgcag 1380gaccgcccgt tcaactatct gaacattctg gcgcaagccg cgcaagccaa agttgcgcag 1380
agcgtgaatg cg 1392agcgtgaatg cg 1392
<210> 13<210> 13
<211> 1029<211> 1029
<212> DNA<212> DNA
<213> 人工合成<213> Synthetic
<400> 13<400> 13
atgttccgtc aagggatcac gggtaggagc caccttatga gtcagaatac gctgaaagtt 60atgttccgtc aagggatcac gggtaggagc caccttatga gtcagaatac gctgaaagtt 60
catgatttaa atgaagatgc ggaatttgat gagaacggag ttgaggtttt tgacgaaaag 120catgatttaa atgaagatgc ggaatttgat gagaacggag ttgaggtttt tgacgaaaag 120
gccttagtag aagaggaacc cagtgataac gatttggccg aagaggaact gttatcgcag 180gccttagtag aagaggaacc cagtgataac gatttggccg aagaggaact gttatcgcag 180
ggagccacac agcgtgtgtt ggacgcgact cagctttacc ttggtgagat tggttattca 240ggagccacac agcgtgtgtt ggacgcgact cagctttacc ttggtgagat tggttattca 240
ccactgttaa cggccgaaga agaagtttat tttgcgcgtc gcgcactgcg tggagatgtc 300ccactgttaa cggccgaaga agaagtttat tttgcgcgtc gcgcactgcg tggagatgtc 300
gcctctcgcc gccggatgat cgagagtaac ttgcgtctgg tggtaaaaat tgcccgccgt 360gcctctcgcc gccggatgat cgagagtaac ttgcgtctgg tggtaaaaat tgcccgccgt 360
tatggcaatc gtggtctggc gttgctggac cttatcgaag agggcaacct ggggctgatc 420tatggcaatc gtggtctggc gttgctggac cttatcgaag agggcaacct ggggctgatc 420
cgcgcggtag agaagtttga cccggaacgt ggtttccgct tctcaacata cgcaacctgg 480cgcgcggtag agaagtttga cccggaacgt ggtttccgct tctcaacata cgcaacctgg 480
tggattcgcc agacgattga acgggcgatt atgaaccaaa cccgtactat tcgtttgccg 540tggattcgcc agacgattga acgggcgatt atgaaccaaa cccgtactat tcgtttgccg 540
attcacatcg taaaggagct gaacgtttac ctgcgaaccg cacgtgagtt gtcccataag 600attcacatcg taaaggagct gaacgtttac ctgcgaaccg cacgtgagtt gtcccataag 600
ctggaccatg aaccaagtgc ggaagagatc gcagagcaac tggataagcc agttgatgac 660ctggaccatg aaccaagtgc ggaagagatc gcagagcaac tggataagcc agttgatgac 660
gtcagccgta tgcttcgtct taacgagcgc attacctcgg tagacacccc gctgggtggt 720gtcagccgta tgcttcgtct taacgagcgc attacctcgg tagacacccc gctgggtggt 720
gattccgaaa aagcgttgct ggacatcctg gccgatgaaa aagagaacgg tccggaagat 780gattccgaaa aagcgttgct ggacatcctg gccgatgaaa aagagaacgg tccggaagat 780
accacgcaag atgacgatat gaagcagagc atcgtcaaat ggctgttcga gctgaacgcc 840accacgcaag atgacgatat gaagcagagc atcgtcaaat ggctgttcga gctgaacgcc 840
aaacagcgtg aagtgctggc acgtcgattc ggtttgctgg ggtacgaagc ggcaacactg 900aaacagcgtg aagtgctggc acgtcgattc ggtttgctgg ggtacgaagc ggcaacactg 900
gaagatgtag gtcgtgaaat tggcctcacc cgtgaacgtg ttcgccagat tcaggttgaa 960gaagatgtag gtcgtgaaat tggcctcacc cgtgaacgtg ttcgccagat tcaggttgaa 960
ggcctgcgcc gtttgcgtga aatcctgcaa acgcaggggc tgaatatcga agcgctgttc 1020ggcctgcgcc gtttgcgtga aatcctgcaa acgcaggggc tgaatatcga agcgctgttc 1020
cgcgagtaa 1029cgcgagtaa 1029
<210> 14<210> 14
<211> 687<211> 687
<212> PRT<212> PRT
<213> 人工合成<213> Synthetic
<400> 14<400> 14
Met Ala Arg Gly Cys Leu Met Thr Ile Ser Gly Gly Thr Phe Asp ProMet Ala Arg Gly Cys Leu Met Thr Ile Ser Gly Gly Thr Phe Asp Pro
1 5 10 151 5 10 15
Ser Ile Cys Glu Met Glu Pro Ile Ala Thr Pro Gly Ala Ile Gln ProSer Ile Cys Glu Met Glu Pro Ile Ala Thr Pro Gly Ala Ile Gln Pro
20 25 30 20 25 30
His Gly Ala Leu Met Thr Ala Arg Ala Asp Ser Gly Arg Val Ala HisHis Gly Ala Leu Met Thr Ala Arg Ala Asp Ser Gly Arg Val Ala His
35 40 45 35 40 45
Ala Ser Val Asn Leu Gly Glu Ile Leu Gly Leu Pro Ala Ala Ser ValAla Ser Val Asn Leu Gly Glu Ile Leu Gly Leu Pro Ala Ala Ser Val
50 55 60 50 55 60
Leu Gly Ala Pro Ile Gly Glu Val Ile Gly Arg Val Asn Glu Ile LeuLeu Gly Ala Pro Ile Gly Glu Val Ile Gly Arg Val Asn Glu Ile Leu
65 70 75 8065 70 75 80
Leu Arg Glu Ala Arg Arg Ser Gly Ser Glu Thr Pro Glu Thr Ile GlyLeu Arg Glu Ala Arg Arg Ser Gly Ser Glu Thr Pro Glu Thr Ile Gly
85 90 95 85 90 95
Ser Phe Arg Arg Ser Asp Gly Gln Leu Leu His Leu His Ala Phe GlnSer Phe Arg Arg Ser Asp Gly Gln Leu Leu His Leu His Ala Phe Gln
100 105 110 100 105 110
Ser Gly Asp Tyr Met Cys Leu Asp Ile Glu Pro Val Arg Asp Glu AspSer Gly Asp Tyr Met Cys Leu Asp Ile Glu Pro Val Arg Asp Glu Asp
115 120 125 115 120 125
Gly Arg Leu Pro Pro Gly Ala Arg Gln Ser Val Ile Glu Thr Phe SerGly Arg Leu Pro Pro Gly Ala Arg Gln Ser Val Ile Glu Thr Phe Ser
130 135 140 130 135 140
Ser Ala Met Thr Gln Val Glu Leu Cys Glu Leu Ala Val His Gly LeuSer Ala Met Thr Gln Val Glu Leu Cys Glu Leu Ala Val His Gly Leu
145 150 155 160145 150 155 160
Gln Leu Val Leu Gly Tyr Asp Arg Val Met Ala Tyr Arg Phe Gly AlaGln Leu Val Leu Gly Tyr Asp Arg Val Met Ala Tyr Arg Phe Gly Ala
165 170 175 165 170 175
Asp Gly His Gly Glu Val Ile Ala Glu Arg Arg Arg Gln Asp Leu GluAsp Gly His Gly Glu Val Ile Ala Glu Arg Arg Arg Gln Asp Leu Glu
180 185 190 180 185 190
Pro Tyr Leu Gly Leu His Tyr Pro Ala Ser Asp Ile Pro Gln Ile AlaPro Tyr Leu Gly Leu His Tyr Pro Ala Ser Asp Ile Pro Gln Ile Ala
195 200 205 195 200 205
Arg Ala Leu Tyr Leu Arg Gln Arg Val Gly Ala Ile Ala Asp Ala CysArg Ala Leu Tyr Leu Arg Gln Arg Val Gly Ala Ile Ala Asp Ala Cys
210 215 220 210 215 220
Tyr Arg Pro Val Pro Leu Leu Gly His Pro Glu Leu Asp Asp Gly LysTyr Arg Pro Val Pro Leu Leu Gly His Pro Glu Leu Asp Asp Gly Lys
225 230 235 240225 230 235 240
Pro Leu Asp Leu Thr His Ser Ser Leu Arg Ser Val Ser Pro Val HisPro Leu Asp Leu Thr His Ser Ser Leu Arg Ser Val Ser Pro Val His
245 250 255 245 250 255
Leu Asp Tyr Met Gln Asn Met Asn Thr Ala Ala Ser Leu Thr Ile GlyLeu Asp Tyr Met Gln Asn Met Asn Thr Ala Ala Ser Leu Thr Ile Gly
260 265 270 260 265 270
Leu Ala Asp Gly Asp Arg Leu Trp Gly Met Leu Val Cys His Asn ThrLeu Ala Asp Gly Asp Arg Leu Trp Gly Met Leu Val Cys His Asn Thr
275 280 285 275 280 285
Thr Pro Arg Ile Ala Gly Pro Glu Trp Arg Ala Ala Ala Gly Met IleThr Pro Arg Ile Ala Gly Pro Glu Trp Arg Ala Ala Ala Gly Met Ile
290 295 300 290 295 300
Gly Gln Val Val Ser Leu Leu Leu Ser Arg Leu Gly Glu Val Glu AsnGly Gln Val Val Ser Leu Leu Leu Ser Arg Leu Gly Glu Val Glu Asn
305 310 315 320305 310 315 320
Ala Ala Glu Thr Leu Ala Arg Gln Ser Thr Leu Ser Thr Leu Val GluAla Ala Glu Thr Leu Ala Arg Gln Ser Thr Leu Ser Thr Leu Val Glu
325 330 335 325 330 335
Arg Leu Ser Thr Gly Asp Thr Leu Ala Ala Ala Phe Val Ala Ala AspArg Leu Ser Thr Gly Asp Thr Leu Ala Ala Ala Phe Val Ala Ala Asp
340 345 350 340 345 350
Gln Leu Ile Leu Asp Leu Val Gly Ala Ser Ala Ala Val Val Arg LeuGln Leu Ile Leu Asp Leu Val Gly Ala Ser Ala Ala Val Val Arg Leu
355 360 365 355 360 365
Ala Gly Gln Glu Leu His Phe Gly Arg Thr Pro Pro Val Asp Ala MetAla Gly Gln Glu Leu His Phe Gly Arg Thr Pro Pro Val Asp Ala Met
370 375 380 370 375 380
Gln Lys Val Leu Asp Ser Leu Gly Arg Pro Ser Pro Leu Glu Val LeuGln Lys Val Leu Asp Ser Leu Gly Arg Pro Ser Pro Leu Glu Val Leu
385 390 395 400385 390 395 400
Ser Leu Asp Asp Val Thr Leu Arg His Pro Glu Leu Pro Glu Leu LeuSer Leu Asp Asp Val Thr Leu Arg His Pro Glu Leu Pro Glu Leu Leu
405 410 415 405 410 415
Ala Ala Gly Ser Gly Ile Leu Leu Leu Pro Leu Thr Ser Gly Asp GlyAla Ala Gly Ser Gly Ile Leu Leu Leu Pro Leu Thr Ser Gly Asp Gly
420 425 430 420 425 430
Asp Leu Ile Ala Trp Phe Arg Pro Glu His Val Gln Thr Ile Thr TrpAsp Leu Ile Ala Trp Phe Arg Pro Glu His Val Gln Thr Ile Thr Trp
435 440 445 435 440 445
Gly Gly Asn Pro Ala Glu His Gly Thr Trp Asn Pro Ala Thr Gln ArgGly Gly Asn Pro Ala Glu His Gly Thr Trp Asn Pro Ala Thr Gln Arg
450 455 460 450 455 460
Met Arg Pro Arg Ala Ser Phe Asp Ala Trp Lys Glu Thr Val Thr GlyMet Arg Pro Arg Ala Ser Phe Asp Ala Trp Lys Glu Thr Val Thr Gly
465 470 475 480465 470 475 480
Arg Ser Leu Pro Trp Thr Ser Ala Glu Arg Asn Cys Ala Arg Glu LeuArg Ser Leu Pro Trp Thr Ser Ala Glu Arg Asn Cys Ala Arg Glu Leu
485 490 495 485 490 495
Gly Glu Ala Ile Ala Ala Glu Met Ala Gln Arg Thr Arg Ala Glu GluGly Glu Ala Ile Ala Ala Glu Met Ala Gln Arg Thr Arg Ala Glu Glu
500 505 510 500 505 510
Leu Glu Arg Val Ala Met Val Asp Ser Leu Thr Arg Leu Trp Asn ArgLeu Glu Arg Val Ala Met Val Asp Ser Leu Thr Arg Leu Trp Asn Arg
515 520 525 515 520 525
Leu Gly Ile Glu Thr Leu Leu Lys Arg Glu Trp Glu Tyr Ala Thr ArgLeu Gly Ile Glu Thr Leu Leu Lys Arg Glu Trp Glu Tyr Ala Thr Arg
530 535 540 530 535 540
Lys Asn Ser Pro Ile Ser Ile Val Met Ile Asp Phe Asp Asn Phe LysLys Asn Ser Pro Ile Ser Ile Val Met Ile Asp Phe Asp Asn Phe Lys
545 550 555 560545 550 555 560
Gln Ile Asn Asp Gln His Gly His Leu Val Gly Asp Glu Val Leu GlnGln Ile Asn Asp Gln His Gly His Leu Val Gly Asp Glu Val Leu Gln
565 570 575 565 570 575
Gly Ser Ala Arg Leu Ile Ile Ser Val Leu Ala Ser Tyr Asp Ile LeuGly Ser Ala Arg Leu Ile Ile Ser Val Leu Ala Ser Tyr Asp Ile Leu
580 585 590 580 585 590
Gly Arg Trp Gly Gly Asp Glu Phe Met Leu Ile Leu Pro Gly Ser GlyGly Arg Trp Gly Gly Asp Glu Phe Met Leu Ile Leu Pro Gly Ser Gly
595 600 605 595 600 605
Arg Glu Gln Thr Ala Val Leu Leu Glu Arg Ile Gln Ala Thr Ile AlaArg Glu Gln Thr Ala Val Leu Leu Glu Arg Ile Gln Ala Thr Ile Ala
610 615 620 610 615 620
Gln Asn Pro Val Pro Thr Ser Ala Gly Pro Met Ala Ile Ser Leu SerGln Asn Pro Val Pro Thr Ser Ala Gly Pro Met Ala Ile Ser Leu Ser
625 630 635 640625 630 635 640
Met Gly Gly Val Ser Val Phe Thr Asn Gln Gly Glu Ala Leu Gln TyrMet Gly Gly Val Ser Val Phe Thr Asn Gln Gly Glu Ala Leu Gln Tyr
645 650 655 645 650 655
Trp Val Glu Gln Ala Asp Asn Gln Leu Met Lys Val Lys Arg Leu GlyTrp Val Glu Gln Ala Asp Asn Gln Leu Met Lys Val Lys Arg Leu Gly
660 665 670 660 665 670
Lys Gly Asn Phe Gln Leu Ala Glu Tyr His His His His His HisLys Gly Asn Phe Gln Leu Ala Glu Tyr His His His His His His
675 680 685 675 680 685
<210> 15<210> 15
<211> 197<211> 197
<212> PRT<212> PRT
<213> 人工合成<213> Synthetic
<400> 15<400> 15
Met Pro Leu Ser Arg Asp Leu Arg Glu Lys Thr Gly Met Leu His AsnMet Pro Leu Ser Arg Asp Leu Arg Glu Lys Thr Gly Met Leu His Asn
1 5 10 151 5 10 15
Arg Ala Glu Thr Leu Leu Gly Leu Pro Ser Gly Ile Met Gly Trp AlaArg Ala Glu Thr Leu Leu Gly Leu Pro Ser Gly Ile Met Gly Trp Ala
20 25 30 20 25 30
Asp Tyr Val Asp Trp Leu Arg His Phe Leu Ala Leu Tyr Asp Pro IleAsp Tyr Val Asp Trp Leu Arg His Phe Leu Ala Leu Tyr Asp Pro Ile
35 40 45 35 40 45
Glu Arg Arg Ile Val Ala Phe Gly Gly Trp Ser Gly Leu Ala Ser PheGlu Arg Arg Ile Val Ala Phe Gly Gly Trp Ser Gly Leu Ala Ser Phe
50 55 60 50 55 60
Asp Pro Asp Pro Gly His Ser Arg Arg Leu Ile Gln Asp Leu His AlaAsp Pro Asp Pro Gly His Ser Arg Arg Leu Ile Gln Asp Leu His Ala
65 70 75 8065 70 75 80
Leu Gly Ile Asp Thr Asp Arg Ile Pro Arg Ala Pro Ala Glu Tyr CysLeu Gly Ile Asp Thr Asp Arg Ile Pro Arg Ala Pro Ala Glu Tyr Cys
85 90 95 85 90 95
Pro Pro Leu Thr Asn Phe Ala Arg Ala Leu Gly Ala Arg Tyr Val LeuPro Pro Leu Thr Asn Phe Ala Arg Ala Leu Gly Ala Arg Tyr Val Leu
100 105 110 100 105 110
Glu Gly Ser Ala Leu Gly Gly Arg Val Ile Leu His His Leu Lys LysGlu Gly Ser Ala Leu Gly Gly Arg Val Ile Leu His His Leu Lys Lys
115 120 125 115 120 125
Arg Ile Gly Asp Glu Ile Gly Asn Ala Thr Ala Phe Phe Gly Gly ProArg Ile Gly Asp Glu Ile Gly Asn Ala Thr Ala Phe Phe Gly Gly Pro
130 135 140 130 135 140
Ser His Gly Thr Ala Thr His Trp Arg Ala Phe Gln Ala Ala Leu AspSer His Gly Thr Ala Thr His Trp Arg Ala Phe Gln Ala Ala Leu Asp
145 150 155 160145 150 155 160
Arg Phe Gly Ala Ala His Pro Asp Lys Arg Ala Asp Val Leu Ala GlyArg Phe Gly Ala Ala His Pro Asp Lys Arg Ala Asp Val Leu Ala Gly
165 170 175 165 170 175
Ala Ala Ala Thr Phe Thr Ala Leu Leu Glu Trp Phe Thr Pro Phe ValAla Ala Ala Thr Phe Thr Ala Leu Leu Glu Trp Phe Thr Pro Phe Val
180 185 190 180 185 190
Ala Ala Arg Arg ValAla Ala Arg Arg Val
195 195
<210> 16<210> 16
<211> 255<211> 255
<212> PRT<212> PRT
<213> 人工合成<213> Synthetic
<400> 16<400> 16
Met Ile Arg Gln Val Ile Gln Arg Ile Ser Asn Pro Glu Ala Ser IleMet Ile Arg Gln Val Ile Gln Arg Ile Ser Asn Pro Glu Ala Ser Ile
1 5 10 151 5 10 15
Glu Ser Leu Gln Glu Arg Arg Phe Trp Leu Gln Cys Glu Arg Ala TyrGlu Ser Leu Gln Glu Arg Arg Phe Trp Leu Gln Cys Glu Arg Ala Tyr
20 25 30 20 25 30
Thr Trp Gln Pro Ile Tyr Gln Thr Cys Gly Arg Leu Met Ala Val GluThr Trp Gln Pro Ile Tyr Gln Thr Cys Gly Arg Leu Met Ala Val Glu
35 40 45 35 40 45
Leu Leu Thr Val Val Thr His Pro Leu Asn Pro Ser Gln Arg Leu ProLeu Leu Thr Val Val Thr His Pro Leu Asn Pro Ser Gln Arg Leu Pro
50 55 60 50 55 60
Pro Asp Arg Tyr Phe Thr Glu Ile Thr Val Ser His Arg Met Glu ValPro Asp Arg Tyr Phe Thr Glu Ile Thr Val Ser His Arg Met Glu Val
65 70 75 8065 70 75 80
Val Lys Glu Gln Ile Asp Leu Leu Ala Gln Lys Ala Asp Phe Phe IleVal Lys Glu Gln Ile Asp Leu Leu Ala Gln Lys Ala Asp Phe Phe Ile
85 90 95 85 90 95
Glu His Gly Leu Leu Ala Ser Val Asn Ile Asp Gly Pro Thr Leu IleGlu His Gly Leu Leu Ala Ser Val Asn Ile Asp Gly Pro Thr Leu Ile
100 105 110 100 105 110
Ala Leu Arg Gln Gln Pro Lys Ile Leu Arg Gln Ile Glu Arg Leu ProAla Leu Arg Gln Gln Pro Lys Ile Leu Arg Gln Ile Glu Arg Leu Pro
115 120 125 115 120 125
Trp Leu Arg Phe Glu Leu Val Glu His Ile Arg Leu Pro Lys Asp SerTrp Leu Arg Phe Glu Leu Val Glu His Ile Arg Leu Pro Lys Asp Ser
130 135 140 130 135 140
Thr Phe Ala Ser Met Cys Glu Phe Gly Pro Leu Trp Leu Asp Asp PheThr Phe Ala Ser Met Cys Glu Phe Gly Pro Leu Trp Leu Asp Asp Phe
145 150 155 160145 150 155 160
Gly Thr Gly Met Ala Asn Phe Ser Ala Leu Ser Glu Val Arg Tyr AspGly Thr Gly Met Ala Asn Phe Ser Ala Leu Ser Glu Val Arg Tyr Asp
165 170 175 165 170 175
Tyr Ile Lys Ile Ala Arg Glu Leu Phe Val Met Leu Arg Gln Ser ProTyr Ile Lys Ile Ala Arg Glu Leu Phe Val Met Leu Arg Gln Ser Pro
180 185 190 180 185 190
Glu Gly Arg Thr Leu Phe Ser Gln Leu Leu His Leu Met Asn Arg TyrGlu Gly Arg Thr Leu Phe Ser Gln Leu Leu His Leu Met Asn Arg Tyr
195 200 205 195 200 205
Cys Arg Gly Val Ile Val Glu Gly Val Glu Thr Pro Glu Glu Trp ArgCys Arg Gly Val Ile Val Glu Gly Val Glu Thr Pro Glu Glu Trp Arg
210 215 220 210 215 220
Asp Val Gln Asn Ser Pro Ala Phe Ala Ala Gln Gly Trp Phe Leu SerAsp Val Gln Asn Ser Pro Ala Phe Ala Ala Gln Gly Trp Phe Leu Ser
225 230 235 240225 230 235 240
Arg Pro Ala Pro Ile Glu Thr Leu Asn Thr Ala Val Leu Ala LeuArg Pro Ala Pro Ile Glu Thr Leu Asn Thr Ala Val Leu Ala Leu
245 250 255 245 250 255
<210> 17<210> 17
<211> 234<211> 234
<212> PRT<212> PRT
<213> 人工合成<213> Synthetic
<400> 17<400> 17
Met Thr Glu Gly Thr Ile Lys Thr Ser Lys Tyr Glu Ile Ile Ala IleMet Thr Glu Gly Thr Ile Lys Thr Ser Lys Tyr Glu Ile Ile Ala Ile
1 5 10 151 5 10 15
Phe Arg Glu Glu Leu Arg Lys Arg Thr Glu Ile Glu Ile Phe Phe AsnPhe Arg Glu Glu Leu Arg Lys Arg Thr Glu Ile Glu Ile Phe Phe Asn
20 25 30 20 25 30
Asn Thr Ser Ile Ile Thr Gln Leu Thr Arg Val Asp Phe Ala Glu PheAsn Thr Ser Ile Ile Thr Gln Leu Thr Arg Val Asp Phe Ala Glu Phe
35 40 45 35 40 45
His Ile Gln Thr His Arg Lys Ile Pro Ser Gly His Lys Ile Arg PheHis Ile Gln Thr His Arg Lys Ile Pro Ser Gly His Lys Ile Arg Phe
50 55 60 50 55 60
Leu Leu His Ser Asp Ser Gly Lys Ile Glu Phe Asn Ala Ala Leu ThrLeu Leu His Ser Asp Ser Gly Lys Ile Glu Phe Asn Ala Ala Leu Thr
65 70 75 8065 70 75 80
Lys His Asp Asn Ser Gly Val Asp Lys Gly Ile Arg Tyr Ala Phe SerLys His Asp Asn Ser Gly Val Asp Lys Gly Ile Arg Tyr Ala Phe Ser
85 90 95 85 90 95
Leu Pro Glu Cys Leu Gln Val Val Gln Arg Arg Arg Asp Pro Arg PheLeu Pro Glu Cys Leu Gln Val Val Gln Arg Arg Arg Asp Pro Arg Phe
100 105 110 100 105 110
Arg Leu Arg His Glu His Asp Phe Tyr Cys Arg Gly Arg His Lys AsnArg Leu Arg His Glu His Asp Phe Tyr Cys Arg Gly Arg His Lys Asn
115 120 125 115 120 125
Gly Glu Asn Tyr Leu Phe Asp Ile Lys Asp Ile Ser Asp Gly Gly CysGly Glu Asn Tyr Leu Phe Asp Ile Lys Asp Ile Ser Asp Gly Gly Cys
130 135 140 130 135 140
Ala Leu Met Thr Lys Thr Pro Asn Leu Lys Phe Leu Ser His Asn AlaAla Leu Met Thr Lys Thr Pro Asn Leu Lys Phe Leu Ser His Asn Ala
145 150 155 160145 150 155 160
Leu Leu Lys Asn Ala Val Leu Met Leu Ala Glu Tyr Gly Glu Ile ThrLeu Leu Lys Asn Ala Val Leu Met Leu Ala Glu Tyr Gly Glu Ile Thr
165 170 175 165 170 175
Ile Asp Leu Val Val Lys Asn Val Ile Val Ile Thr Leu Asp Asn AlaIle Asp Leu Val Val Lys Asn Val Ile Val Ile Thr Leu Asp Asn Ala
180 185 190 180 185 190
Asn Glu Glu Ser Glu Ser Tyr Tyr Gln Ile Ser Cys Gln Phe Lys PheAsn Glu Glu Ser Glu Ser Tyr Tyr Gln Ile Ser Cys Gln Phe Lys Phe
195 200 205 195 200 205
Arg His Leu Asp Asp Gln Arg Arg Ile Glu Lys Ile Leu Leu Asp LeuArg His Leu Asp Asp Gln Arg Arg Ile Glu Lys Ile Leu Leu Asp Leu
210 215 220 210 215 220
Ile Leu Glu Ala Lys Arg Lys Lys Arg IleIle Leu Glu Ala Lys Arg Lys Lys Arg Ile
225 230225 230
<210> 18<210> 18
<211> 342<211> 342
<212> PRT<212> PRT
<213> 人工合成<213> Synthetic
<400> 18<400> 18
Met Phe Arg Gln Gly Ile Thr Gly Arg Ser His Leu Met Ser Gln AsnMet Phe Arg Gln Gly Ile Thr Gly Arg Ser His Leu Met Ser Gln Asn
1 5 10 151 5 10 15
Thr Leu Lys Val His Asp Leu Asn Glu Asp Ala Glu Phe Asp Glu AsnThr Leu Lys Val His Asp Leu Asn Glu Asp Ala Glu Phe Asp Glu Asn
20 25 30 20 25 30
Gly Val Glu Val Phe Asp Glu Lys Ala Leu Val Glu Glu Glu Pro SerGly Val Glu Val Phe Asp Glu Lys Ala Leu Val Glu Glu Glu Pro Ser
35 40 45 35 40 45
Asp Asn Asp Leu Ala Glu Glu Glu Leu Leu Ser Gln Gly Ala Thr GlnAsp Asn Asp Leu Ala Glu Glu Glu Leu Leu Ser Gln Gly Ala Thr Gln
50 55 60 50 55 60
Arg Val Leu Asp Ala Thr Gln Leu Tyr Leu Gly Glu Ile Gly Tyr SerArg Val Leu Asp Ala Thr Gln Leu Tyr Leu Gly Glu Ile Gly Tyr Ser
65 70 75 8065 70 75 80
Pro Leu Leu Thr Ala Glu Glu Glu Val Tyr Phe Ala Arg Arg Ala LeuPro Leu Leu Thr Ala Glu Glu Glu Val Tyr Phe Ala Arg Arg Ala Leu
85 90 95 85 90 95
Arg Gly Asp Val Ala Ser Arg Arg Arg Met Ile Glu Ser Asn Leu ArgArg Gly Asp Val Ala Ser Arg Arg Arg Met Ile Glu Ser Asn Leu Arg
100 105 110 100 105 110
Leu Val Val Lys Ile Ala Arg Arg Tyr Gly Asn Arg Gly Leu Ala LeuLeu Val Val Lys Ile Ala Arg Arg Tyr Gly Asn Arg Gly Leu Ala Leu
115 120 125 115 120 125
Leu Asp Leu Ile Glu Glu Gly Asn Leu Gly Leu Ile Arg Ala Val GluLeu Asp Leu Ile Glu Glu Gly Asn Leu Gly Leu Ile Arg Ala Val Glu
130 135 140 130 135 140
Lys Phe Asp Pro Glu Arg Gly Phe Arg Phe Ser Thr Tyr Ala Thr TrpLys Phe Asp Pro Glu Arg Gly Phe Arg Phe Ser Thr Tyr Ala Thr Trp
145 150 155 160145 150 155 160
Trp Ile Arg Gln Thr Ile Glu Arg Ala Ile Met Asn Gln Thr Arg ThrTrp Ile Arg Gln Thr Ile Glu Arg Ala Ile Met Asn Gln Thr Arg Thr
165 170 175 165 170 175
Ile Arg Leu Pro Ile His Ile Val Lys Glu Leu Asn Val Tyr Leu ArgIle Arg Leu Pro Ile His Ile Val Lys Glu Leu Asn Val Tyr Leu Arg
180 185 190 180 185 190
Thr Ala Arg Glu Leu Ser His Lys Leu Asp His Glu Pro Ser Ala GluThr Ala Arg Glu Leu Ser His Lys Leu Asp His Glu Pro Ser Ala Glu
195 200 205 195 200 205
Glu Ile Ala Glu Gln Leu Asp Lys Pro Val Asp Asp Val Ser Arg MetGlu Ile Ala Glu Gln Leu Asp Lys Pro Val Asp Asp Val Ser Arg Met
210 215 220 210 215 220
Leu Arg Leu Asn Glu Arg Ile Thr Ser Val Asp Thr Pro Leu Gly GlyLeu Arg Leu Asn Glu Arg Ile Thr Ser Val Asp Thr Pro Leu Gly Gly
225 230 235 240225 230 235 240
Asp Ser Glu Lys Ala Leu Leu Asp Ile Leu Ala Asp Glu Lys Glu AsnAsp Ser Glu Lys Ala Leu Leu Asp Ile Leu Ala Asp Glu Lys Glu Asn
245 250 255 245 250 255
Gly Pro Glu Asp Thr Thr Gln Asp Asp Asp Met Lys Gln Ser Ile ValGly Pro Glu Asp Thr Thr Gln Asp Asp Asp Met Lys Gln Ser Ile Val
260 265 270 260 265 270
Lys Trp Leu Phe Glu Leu Asn Ala Lys Gln Arg Glu Val Leu Ala ArgLys Trp Leu Phe Glu Leu Asn Ala Lys Gln Arg Glu Val Leu Ala Arg
275 280 285 275 280 285
Arg Phe Gly Leu Leu Gly Tyr Glu Ala Ala Thr Leu Glu Asp Val GlyArg Phe Gly Leu Leu Gly Tyr Glu Ala Ala Thr Leu Glu Asp Val Gly
290 295 300 290 295 300
Arg Glu Ile Gly Leu Thr Arg Glu Arg Val Arg Gln Ile Gln Val GluArg Glu Ile Gly Leu Thr Arg Glu Arg Val Arg Gln Ile Gln Val Glu
305 310 315 320305 310 315 320
Gly Leu Arg Arg Leu Arg Glu Ile Leu Gln Thr Gln Gly Leu Asn IleGly Leu Arg Arg Leu Arg Glu Ile Leu Gln Thr Gln Gly Leu Asn Ile
325 330 335 325 330 335
Glu Ala Leu Phe Arg GluGlu Ala Leu Phe Arg Glu
340 340
Claims (5)
Priority Applications (1)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| CN201910836782.5A CN110551669B (en) | 2019-09-05 | 2019-09-05 | Near-infrared light control dynamic regulation and control system and application thereof |
Applications Claiming Priority (1)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| CN201910836782.5A CN110551669B (en) | 2019-09-05 | 2019-09-05 | Near-infrared light control dynamic regulation and control system and application thereof |
Publications (2)
| Publication Number | Publication Date |
|---|---|
| CN110551669A CN110551669A (en) | 2019-12-10 |
| CN110551669B true CN110551669B (en) | 2022-05-06 |
Family
ID=68739188
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| CN201910836782.5A Active CN110551669B (en) | 2019-09-05 | 2019-09-05 | Near-infrared light control dynamic regulation and control system and application thereof |
Country Status (1)
| Country | Link |
|---|---|
| CN (1) | CN110551669B (en) |
Citations (7)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN1993461A (en) * | 2004-08-03 | 2007-07-04 | 德古萨股份公司 | Method for the production of L-amino acids using strains from the enterobacteraceae family |
| CN101203529A (en) * | 2005-02-18 | 2008-06-18 | 诺华疫苗和诊断公司 | Proteins and nucleic acids from meningitis/sepsis-associated Escherichia coli |
| CN103237883A (en) * | 2010-03-12 | 2013-08-07 | 布鲁克哈文科学协会有限责任公司 | Enterobacter 638 and its application method |
| CN107177621A (en) * | 2016-03-10 | 2017-09-19 | 叶海峰 | The method that far-red light gene loop expression control system carries out transgenic regulation expression |
| WO2018156646A1 (en) * | 2017-02-21 | 2018-08-30 | Duke University | Compositions and methods for robust dynamic metabolic control |
| CN110438057A (en) * | 2019-08-21 | 2019-11-12 | 江南大学 | A kind of the dynamic regulation system and its application of blue light control |
| CN110468122A (en) * | 2018-05-11 | 2019-11-19 | 苏州欣赛生物科技有限公司 | A kind of system and application of the differentiation of stem cells of far-red light regulation |
Family Cites Families (2)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US11525117B2 (en) * | 2018-04-24 | 2022-12-13 | University Of Wyoming | Microbial stem cell technology |
| US11203744B2 (en) * | 2018-06-21 | 2021-12-21 | Duke University | Compositions and methods for the production of pyruvic acid and related products using dynamic metabolic control |
-
2019
- 2019-09-05 CN CN201910836782.5A patent/CN110551669B/en active Active
Patent Citations (8)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN1993461A (en) * | 2004-08-03 | 2007-07-04 | 德古萨股份公司 | Method for the production of L-amino acids using strains from the enterobacteraceae family |
| CN101203529A (en) * | 2005-02-18 | 2008-06-18 | 诺华疫苗和诊断公司 | Proteins and nucleic acids from meningitis/sepsis-associated Escherichia coli |
| CN103237883A (en) * | 2010-03-12 | 2013-08-07 | 布鲁克哈文科学协会有限责任公司 | Enterobacter 638 and its application method |
| CN107177621A (en) * | 2016-03-10 | 2017-09-19 | 叶海峰 | The method that far-red light gene loop expression control system carries out transgenic regulation expression |
| WO2018156646A1 (en) * | 2017-02-21 | 2018-08-30 | Duke University | Compositions and methods for robust dynamic metabolic control |
| CN110536960A (en) * | 2017-02-21 | 2019-12-03 | 杜克大学 | Compositions and methods for robust dynamic metabolic control |
| CN110468122A (en) * | 2018-05-11 | 2019-11-19 | 苏州欣赛生物科技有限公司 | A kind of system and application of the differentiation of stem cells of far-red light regulation |
| CN110438057A (en) * | 2019-08-21 | 2019-11-12 | 江南大学 | A kind of the dynamic regulation system and its application of blue light control |
Non-Patent Citations (11)
| Title |
|---|
| Blue light-mediated transcriptional activation and repression of gene expression in bacteria;Premkumar Jayaraman等;《Nucleic Acids Research》;20160819;第44卷(第14期);6994-7005 * |
| c-di-GMP phosphodiesterase PdeH [Escherichia coli str. K-12 substr. MG1655];Riley,M.等;《Genbank Database》;20181011;Accession NO.NP_417982.2 * |
| heme oxygenase [Cereibacter sphaeroides];Hosoyama,A.等;《Genbank Database》;20190801;Accession NO.GEM94683.1 * |
| Light-powered Escherichia coli cell division for chemical production;Qiang Ding等;《NATURE COMMUNICATIONS》;20200508;第11卷(第1期);1-14 * |
| Metabolic engineering of Escherichia coli W3110 to produce L-malate;Xiaoxiang Dong等;《Biotechnology and Bioengineering》;20170331;第114卷(第3期);656-664 * |
| MULTISPECIES: transcriptional activator MrkH [Gammaproteobacteria];Wilksch,J.J.等;《Genbank Database》;20130829;Accession NO.WP_004152886.1 * |
| Near-infrared Light Responsive Synthetic c di-GMP Module for Optogenetic Applications;Min-Hyung Ryu;《ACS Synthetic Biology》;20141121;第3卷(第11期);第803页右栏第1段,第806页图4A,第807页右栏第1-4段,补充资料图S2 * |
| Quorum sensing and biofilm formation in mycobacteria: role of c-di-GMP and methods to study this second messenger;Indra Mani Sharma等;《IUBMB Life》;20141231;第66卷(第12期);823-834 * |
| TPA: cyclic-guanylate-specific phosphodiesterase [Shigella sp.];Parks,D.H.等;《Genbank Database》;20180904;Accession NO.HAB20624.1 * |
| 一种新型转录因子MrkH的结构与功能研究;王峰;《中国博士学位论文全文数据库(电子期刊)医药卫生科技辑》;20170216(第3期);附件补充表格4 * |
| 代谢工程改造Escherichia coli生产L-苹果酸;董晓翔等;《应用与环境生物学报》;20170425;第23卷(第2期);269-275 * |
Also Published As
| Publication number | Publication date |
|---|---|
| CN110551669A (en) | 2019-12-10 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| CN109666658B (en) | Nicotinamide phosphoribosyltransferase, encoding gene, recombinant vector and application for preparing NMN | |
| CN113186142B (en) | Escherichia coli engineering strain for efficiently producing 2' -fucosyllactose | |
| CN110438057B (en) | A dynamic control system for blue light control and its application | |
| CN110904140B (en) | A protein dynamic expression regulation system and its application in the production of shikimate | |
| CN107723307A (en) | A kind of method and its application for efficiently preparing the epimerase of D psicoses 3 | |
| CN107937361B (en) | A kind of alanine dehydrogenase mutant and its application | |
| CN106754601A (en) | A kind of application phospholipase C by intracellular protein extracellular expression method | |
| CN113583925B (en) | Method for preparing patchouli alcohol by fermenting metabolic engineering escherichia coli | |
| CN113005132B (en) | A kind of D-psicose-3-epimerase gene and application method thereof | |
| CN107338258A (en) | The method for producing the engineering bacteria structure and its production beta Alanine of beta Alanine | |
| CN113637617B (en) | Method for synthesizing methylselenocysteine by using bacillus subtilis | |
| CN104774813A (en) | Leucine dehydrogenase and preparation method and application thereof | |
| CN111621452A (en) | A Bacillus subtilis that produces D-p-hydroxyphenylglycine | |
| CN111944795A (en) | Glutamate decarboxylase mutant and its application | |
| CN110551669B (en) | Near-infrared light control dynamic regulation and control system and application thereof | |
| CN101613707B (en) | A method for producing glutathione with metabolic engineering bacteria | |
| CN109971803A (en) | A kind of L-erythrulose and erythritol production method | |
| CN110951760B (en) | Protein time-delay expression switch and application thereof in production of glucaric acid | |
| CN113929787B (en) | Strategy for regulating and controlling biofilm to improve shikimic acid production performance | |
| CN107988131B (en) | A kind of method for high-yield α-keto-γ-methylthiobutyric acid | |
| JP2024536957A (en) | Bacillus licheniformis cells that can be stably and repeatedly used for conversion synthesis of D-psicose | |
| CN116769748A (en) | A 5-aminolevulinic acid synthase mutant and E. coli producing B12 precursor ALA | |
| CN111808836B (en) | Heat-resistant mutant enzyme of pullulanase I and preparation method and application thereof | |
| CN117230099A (en) | Construction method of bacillus capable of efficiently expressing 5-aminolevulinic acid | |
| CN114806899A (en) | Trichoderma reesei engineering bacterium for producing L-malic acid and application thereof |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| PB01 | Publication | ||
| PB01 | Publication | ||
| SE01 | Entry into force of request for substantive examination | ||
| SE01 | Entry into force of request for substantive examination | ||
| GR01 | Patent grant | ||
| GR01 | Patent grant |