CN109929894B - A kind of preparation and activity identification method of porcine second messenger molecule 2'3'-cGAMP - Google Patents
A kind of preparation and activity identification method of porcine second messenger molecule 2'3'-cGAMP Download PDFInfo
- Publication number
- CN109929894B CN109929894B CN201910307619.XA CN201910307619A CN109929894B CN 109929894 B CN109929894 B CN 109929894B CN 201910307619 A CN201910307619 A CN 201910307619A CN 109929894 B CN109929894 B CN 109929894B
- Authority
- CN
- China
- Prior art keywords
- cgamp
- concentration
- activity
- synthetase
- cell
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Active
Links
- XRILCFTWUCUKJR-INFSMZHSSA-N 2'-3'-cGAMP Chemical compound C([C@H]([C@H]1O)O2)OP(O)(=O)O[C@H]3[C@@H](O)[C@H](N4C5=NC=NC(N)=C5N=C4)O[C@@H]3COP(O)(=O)O[C@H]1[C@@H]2N1C=NC2=C1NC(N)=NC2=O XRILCFTWUCUKJR-INFSMZHSSA-N 0.000 title claims abstract description 73
- 230000000694 effects Effects 0.000 title claims abstract description 32
- 238000000034 method Methods 0.000 title claims abstract description 23
- 238000002360 preparation method Methods 0.000 title abstract description 12
- 102000003960 Ligases Human genes 0.000 claims abstract description 41
- 108090000364 Ligases Proteins 0.000 claims abstract description 41
- NYHBQMYGNKIUIF-UUOKFMHZSA-N Guanosine Chemical compound C1=NC=2C(=O)NC(N)=NC=2N1[C@@H]1O[C@H](CO)[C@@H](O)[C@H]1O NYHBQMYGNKIUIF-UUOKFMHZSA-N 0.000 claims description 46
- 210000004027 cell Anatomy 0.000 claims description 46
- 125000004122 cyclic group Chemical group 0.000 claims description 36
- 239000007795 chemical reaction product Substances 0.000 claims description 28
- MIKUYHXYGGJMLM-GIMIYPNGSA-N Crotonoside Natural products C1=NC2=C(N)NC(=O)N=C2N1[C@H]1O[C@@H](CO)[C@H](O)[C@@H]1O MIKUYHXYGGJMLM-GIMIYPNGSA-N 0.000 claims description 23
- NYHBQMYGNKIUIF-UHFFFAOYSA-N D-guanosine Natural products C1=2NC(N)=NC(=O)C=2N=CN1C1OC(CO)C(O)C1O NYHBQMYGNKIUIF-UHFFFAOYSA-N 0.000 claims description 23
- 238000006911 enzymatic reaction Methods 0.000 claims description 23
- 229940029575 guanosine Drugs 0.000 claims description 23
- 239000000243 solution Substances 0.000 claims description 22
- 238000004113 cell culture Methods 0.000 claims description 21
- 239000000047 product Substances 0.000 claims description 20
- 238000012258 culturing Methods 0.000 claims description 11
- 239000006228 supernatant Substances 0.000 claims description 11
- JKMHFZQWWAIEOD-UHFFFAOYSA-N 2-[4-(2-hydroxyethyl)piperazin-1-yl]ethanesulfonic acid Chemical compound OCC[NH+]1CCN(CCS([O-])(=O)=O)CC1 JKMHFZQWWAIEOD-UHFFFAOYSA-N 0.000 claims description 10
- 239000007995 HEPES buffer Substances 0.000 claims description 10
- TWRXJAOTZQYOKJ-UHFFFAOYSA-L Magnesium chloride Chemical compound [Mg+2].[Cl-].[Cl-] TWRXJAOTZQYOKJ-UHFFFAOYSA-L 0.000 claims description 10
- 210000000170 cell membrane Anatomy 0.000 claims description 8
- 239000003795 chemical substances by application Substances 0.000 claims description 8
- 229910001629 magnesium chloride Inorganic materials 0.000 claims description 8
- 239000000203 mixture Substances 0.000 claims description 8
- 230000000638 stimulation Effects 0.000 claims description 8
- 108060001084 Luciferase Proteins 0.000 claims description 7
- 238000002156 mixing Methods 0.000 claims description 6
- HBZBAMXERPYTFS-SECBINFHSA-N (4S)-2-(6,7-dihydro-5H-pyrrolo[3,2-f][1,3]benzothiazol-2-yl)-4,5-dihydro-1,3-thiazole-4-carboxylic acid Chemical compound OC(=O)[C@H]1CSC(=N1)c1nc2cc3CCNc3cc2s1 HBZBAMXERPYTFS-SECBINFHSA-N 0.000 claims description 5
- 239000005089 Luciferase Substances 0.000 claims description 5
- 230000004913 activation Effects 0.000 claims description 5
- 229950006790 adenosine phosphate Drugs 0.000 claims description 5
- 239000000758 substrate Substances 0.000 claims description 4
- 238000005406 washing Methods 0.000 claims description 4
- QRLVDLBMBULFAL-UHFFFAOYSA-N Digitonin Natural products CC1CCC2(OC1)OC3C(O)C4C5CCC6CC(OC7OC(CO)C(OC8OC(CO)C(O)C(OC9OCC(O)C(O)C9OC%10OC(CO)C(O)C(OC%11OC(CO)C(O)C(O)C%11O)C%10O)C8O)C(O)C7O)C(O)CC6(C)C5CCC4(C)C3C2C QRLVDLBMBULFAL-UHFFFAOYSA-N 0.000 claims description 3
- 108091054729 IRF family Proteins 0.000 claims description 3
- 229930006000 Sucrose Natural products 0.000 claims description 3
- CZMRCDWAGMRECN-UGDNZRGBSA-N Sucrose Chemical compound O[C@H]1[C@H](O)[C@@H](CO)O[C@@]1(CO)O[C@@H]1[C@H](O)[C@@H](O)[C@H](O)[C@@H](CO)O1 CZMRCDWAGMRECN-UGDNZRGBSA-N 0.000 claims description 3
- 239000003054 catalyst Substances 0.000 claims description 3
- UVYVLBIGDKGWPX-KUAJCENISA-N digitonin Chemical compound O([C@@H]1[C@@H]([C@]2(CC[C@@H]3[C@@]4(C)C[C@@H](O)[C@H](O[C@H]5[C@@H]([C@@H](O)[C@@H](O[C@H]6[C@@H]([C@@H](O[C@H]7[C@@H]([C@@H](O)[C@H](O)CO7)O)[C@H](O)[C@@H](CO)O6)O[C@H]6[C@@H]([C@@H](O[C@H]7[C@@H]([C@@H](O)[C@H](O)[C@@H](CO)O7)O)[C@@H](O)[C@@H](CO)O6)O)[C@@H](CO)O5)O)C[C@@H]4CC[C@H]3[C@@H]2[C@@H]1O)C)[C@@H]1C)[C@]11CC[C@@H](C)CO1 UVYVLBIGDKGWPX-KUAJCENISA-N 0.000 claims description 3
- UVYVLBIGDKGWPX-UHFFFAOYSA-N digitonine Natural products CC1C(C2(CCC3C4(C)CC(O)C(OC5C(C(O)C(OC6C(C(OC7C(C(O)C(O)CO7)O)C(O)C(CO)O6)OC6C(C(OC7C(C(O)C(O)C(CO)O7)O)C(O)C(CO)O6)O)C(CO)O5)O)CC4CCC3C2C2O)C)C2OC11CCC(C)CO1 UVYVLBIGDKGWPX-UHFFFAOYSA-N 0.000 claims description 3
- 239000005720 sucrose Substances 0.000 claims description 3
- -1 DTT Chemical compound 0.000 claims description 2
- 230000000149 penetrating effect Effects 0.000 claims description 2
- ZDXPYRJPNDTMRX-VKHMYHEASA-N L-glutamine Chemical compound OC(=O)[C@@H](N)CCC(N)=O ZDXPYRJPNDTMRX-VKHMYHEASA-N 0.000 claims 2
- 229930182816 L-glutamine Natural products 0.000 claims 2
- 150000001413 amino acids Chemical group 0.000 claims 2
- 239000006144 Dulbecco’s modified Eagle's medium Substances 0.000 claims 1
- 102000016854 Interferon Regulatory Factors Human genes 0.000 claims 1
- WCUXLLCKKVVCTQ-UHFFFAOYSA-M Potassium chloride Chemical compound [Cl-].[K+] WCUXLLCKKVVCTQ-UHFFFAOYSA-M 0.000 claims 1
- 239000001103 potassium chloride Substances 0.000 claims 1
- 102000004190 Enzymes Human genes 0.000 abstract description 21
- 108090000790 Enzymes Proteins 0.000 abstract description 21
- 230000003197 catalytic effect Effects 0.000 abstract description 8
- 230000004071 biological effect Effects 0.000 abstract description 7
- 230000014509 gene expression Effects 0.000 abstract description 6
- 230000028993 immune response Effects 0.000 abstract description 5
- 239000002671 adjuvant Substances 0.000 abstract description 4
- 230000019491 signal transduction Effects 0.000 abstract description 4
- 238000011031 large-scale manufacturing process Methods 0.000 abstract description 3
- 239000012190 activator Substances 0.000 abstract description 2
- 230000000259 anti-tumor effect Effects 0.000 abstract description 2
- 230000002155 anti-virotic effect Effects 0.000 abstract description 2
- 238000011161 development Methods 0.000 abstract description 2
- 229940079593 drug Drugs 0.000 abstract description 2
- 239000003814 drug Substances 0.000 abstract description 2
- 239000003623 enhancer Substances 0.000 abstract description 2
- 241000282898 Sus scrofa Species 0.000 description 27
- 229940088598 enzyme Drugs 0.000 description 20
- 108020004414 DNA Proteins 0.000 description 15
- 238000006243 chemical reaction Methods 0.000 description 15
- 108090000623 proteins and genes Proteins 0.000 description 13
- 102000053602 DNA Human genes 0.000 description 9
- 241000894006 Bacteria Species 0.000 description 8
- 239000007788 liquid Substances 0.000 description 7
- 239000013604 expression vector Substances 0.000 description 6
- 108091028043 Nucleic acid sequence Proteins 0.000 description 5
- 125000003275 alpha amino acid group Chemical group 0.000 description 5
- 238000009472 formulation Methods 0.000 description 5
- 101710163270 Nuclease Proteins 0.000 description 4
- 210000001744 T-lymphocyte Anatomy 0.000 description 4
- 239000011543 agarose gel Substances 0.000 description 4
- 238000004140 cleaning Methods 0.000 description 4
- 239000000287 crude extract Substances 0.000 description 4
- 238000009396 hybridization Methods 0.000 description 4
- 239000012528 membrane Substances 0.000 description 4
- 239000000137 peptide hydrolase inhibitor Substances 0.000 description 4
- 102000004169 proteins and genes Human genes 0.000 description 4
- 238000002415 sodium dodecyl sulfate polyacrylamide gel electrophoresis Methods 0.000 description 4
- 101000708016 Caenorhabditis elegans Sentrin-specific protease Proteins 0.000 description 3
- 241000588724 Escherichia coli Species 0.000 description 3
- 229940124158 Protease/peptidase inhibitor Drugs 0.000 description 3
- 230000000840 anti-viral effect Effects 0.000 description 3
- 239000002246 antineoplastic agent Substances 0.000 description 3
- 229940041181 antineoplastic drug Drugs 0.000 description 3
- 239000003443 antiviral agent Substances 0.000 description 3
- 239000006143 cell culture medium Substances 0.000 description 3
- 238000005516 engineering process Methods 0.000 description 3
- 238000001976 enzyme digestion Methods 0.000 description 3
- 238000002474 experimental method Methods 0.000 description 3
- 230000004927 fusion Effects 0.000 description 3
- 230000000091 immunopotentiator Effects 0.000 description 3
- 238000011534 incubation Methods 0.000 description 3
- 230000002757 inflammatory effect Effects 0.000 description 3
- 239000008191 permeabilizing agent Substances 0.000 description 3
- 238000003786 synthesis reaction Methods 0.000 description 3
- 210000004881 tumor cell Anatomy 0.000 description 3
- 238000000108 ultra-filtration Methods 0.000 description 3
- 101001011382 Homo sapiens Interferon regulatory factor 3 Proteins 0.000 description 2
- 102000043138 IRF family Human genes 0.000 description 2
- 108010050904 Interferons Proteins 0.000 description 2
- DSFYPIUSAMSERP-IHRRRGAJSA-N Leu-Leu-Arg Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](CC(C)C)C(=O)N[C@H](C(O)=O)CCCN=C(N)N DSFYPIUSAMSERP-IHRRRGAJSA-N 0.000 description 2
- 102000016943 Muramidase Human genes 0.000 description 2
- 108010014251 Muramidase Proteins 0.000 description 2
- 108010062010 N-Acetylmuramoyl-L-alanine Amidase Proteins 0.000 description 2
- 206010028980 Neoplasm Diseases 0.000 description 2
- 108010034546 Serratia marcescens nuclease Proteins 0.000 description 2
- 241000700605 Viruses Species 0.000 description 2
- 230000003213 activating effect Effects 0.000 description 2
- 238000004458 analytical method Methods 0.000 description 2
- 230000015572 biosynthetic process Effects 0.000 description 2
- 239000000872 buffer Substances 0.000 description 2
- 230000002255 enzymatic effect Effects 0.000 description 2
- 239000012634 fragment Substances 0.000 description 2
- 210000003976 gap junction Anatomy 0.000 description 2
- 238000010353 genetic engineering Methods 0.000 description 2
- 238000000338 in vitro Methods 0.000 description 2
- BPHPUYQFMNQIOC-NXRLNHOXSA-N isopropyl beta-D-thiogalactopyranoside Chemical compound CC(C)S[C@@H]1O[C@H](CO)[C@H](O)[C@H](O)[C@H]1O BPHPUYQFMNQIOC-NXRLNHOXSA-N 0.000 description 2
- 229960000274 lysozyme Drugs 0.000 description 2
- 239000004325 lysozyme Substances 0.000 description 2
- 235000010335 lysozyme Nutrition 0.000 description 2
- 108010009298 lysylglutamic acid Proteins 0.000 description 2
- 230000035800 maturation Effects 0.000 description 2
- 239000002773 nucleotide Substances 0.000 description 2
- 125000003729 nucleotide group Chemical group 0.000 description 2
- 230000009465 prokaryotic expression Effects 0.000 description 2
- 238000000746 purification Methods 0.000 description 2
- 238000011002 quantification Methods 0.000 description 2
- 238000003259 recombinant expression Methods 0.000 description 2
- 238000011160 research Methods 0.000 description 2
- 230000028327 secretion Effects 0.000 description 2
- 238000000926 separation method Methods 0.000 description 2
- 230000004936 stimulating effect Effects 0.000 description 2
- 210000002845 virion Anatomy 0.000 description 2
- HHGYNJRJIINWAK-FXQIFTODSA-N Ala-Ala-Arg Chemical compound C[C@H](N)C(=O)N[C@@H](C)C(=O)N[C@H](C(O)=O)CCCN=C(N)N HHGYNJRJIINWAK-FXQIFTODSA-N 0.000 description 1
- RLMISHABBKUNFO-WHFBIAKZSA-N Ala-Ala-Gly Chemical compound C[C@H](N)C(=O)N[C@@H](C)C(=O)NCC(O)=O RLMISHABBKUNFO-WHFBIAKZSA-N 0.000 description 1
- CXRCVCURMBFFOL-FXQIFTODSA-N Ala-Ala-Pro Chemical compound C[C@H](N)C(=O)N[C@@H](C)C(=O)N1CCC[C@H]1C(O)=O CXRCVCURMBFFOL-FXQIFTODSA-N 0.000 description 1
- SVBXIUDNTRTKHE-CIUDSAMLSA-N Ala-Arg-Glu Chemical compound [H]N[C@@H](C)C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CCC(O)=O)C(O)=O SVBXIUDNTRTKHE-CIUDSAMLSA-N 0.000 description 1
- TTXMOJWKNRJWQJ-FXQIFTODSA-N Ala-Arg-Ser Chemical compound OC[C@@H](C(O)=O)NC(=O)[C@@H](NC(=O)[C@@H](N)C)CCCN=C(N)N TTXMOJWKNRJWQJ-FXQIFTODSA-N 0.000 description 1
- IFTVANMRTIHKML-WDSKDSINSA-N Ala-Gln-Gly Chemical compound C[C@H](N)C(=O)N[C@@H](CCC(N)=O)C(=O)NCC(O)=O IFTVANMRTIHKML-WDSKDSINSA-N 0.000 description 1
- KXEVYGKATAMXJJ-ACZMJKKPSA-N Ala-Glu-Asp Chemical compound C[C@H](N)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CC(O)=O)C(O)=O KXEVYGKATAMXJJ-ACZMJKKPSA-N 0.000 description 1
- OMMDTNGURYRDAC-NRPADANISA-N Ala-Glu-Val Chemical compound [H]N[C@@H](C)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](C(C)C)C(O)=O OMMDTNGURYRDAC-NRPADANISA-N 0.000 description 1
- ZVFVBBGVOILKPO-WHFBIAKZSA-N Ala-Gly-Ala Chemical compound C[C@H](N)C(=O)NCC(=O)N[C@@H](C)C(O)=O ZVFVBBGVOILKPO-WHFBIAKZSA-N 0.000 description 1
- CCDFBRZVTDDJNM-GUBZILKMSA-N Ala-Leu-Glu Chemical compound [H]N[C@@H](C)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CCC(O)=O)C(O)=O CCDFBRZVTDDJNM-GUBZILKMSA-N 0.000 description 1
- MDNAVFBZPROEHO-DCAQKATOSA-N Ala-Lys-Val Chemical compound [H]N[C@@H](C)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](C(C)C)C(O)=O MDNAVFBZPROEHO-DCAQKATOSA-N 0.000 description 1
- MDNAVFBZPROEHO-UHFFFAOYSA-N Ala-Lys-Val Natural products CC(C)C(C(O)=O)NC(=O)C(NC(=O)C(C)N)CCCCN MDNAVFBZPROEHO-UHFFFAOYSA-N 0.000 description 1
- OLVCTPPSXNRGKV-GUBZILKMSA-N Ala-Pro-Pro Chemical compound C[C@H](N)C(=O)N1CCC[C@H]1C(=O)N1[C@H](C(O)=O)CCC1 OLVCTPPSXNRGKV-GUBZILKMSA-N 0.000 description 1
- RTZCUEHYUQZIDE-WHFBIAKZSA-N Ala-Ser-Gly Chemical compound C[C@H](N)C(=O)N[C@@H](CO)C(=O)NCC(O)=O RTZCUEHYUQZIDE-WHFBIAKZSA-N 0.000 description 1
- KWKQGHSSNHPGOW-BQBZGAKWSA-N Arg-Ala-Gly Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](C)C(=O)NCC(O)=O KWKQGHSSNHPGOW-BQBZGAKWSA-N 0.000 description 1
- VWVPYNGMOCSSGK-GUBZILKMSA-N Arg-Arg-Asn Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CC(N)=O)C(O)=O VWVPYNGMOCSSGK-GUBZILKMSA-N 0.000 description 1
- IASNWHAGGYTEKX-IUCAKERBSA-N Arg-Arg-Gly Chemical compound NC(N)=NCCC[C@H](N)C(=O)N[C@@H](CCCN=C(N)N)C(=O)NCC(O)=O IASNWHAGGYTEKX-IUCAKERBSA-N 0.000 description 1
- USNSOPDIZILSJP-FXQIFTODSA-N Arg-Asn-Asn Chemical compound NC(N)=NCCC[C@H](N)C(=O)N[C@@H](CC(N)=O)C(=O)N[C@@H](CC(N)=O)C(O)=O USNSOPDIZILSJP-FXQIFTODSA-N 0.000 description 1
- OTCJMMRQBVDQRK-DCAQKATOSA-N Arg-Asp-Leu Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CC(O)=O)C(=O)N[C@@H](CC(C)C)C(O)=O OTCJMMRQBVDQRK-DCAQKATOSA-N 0.000 description 1
- NKBQZKVMKJJDLX-SRVKXCTJSA-N Arg-Glu-Leu Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CC(C)C)C(O)=O NKBQZKVMKJJDLX-SRVKXCTJSA-N 0.000 description 1
- WVNFNPGXYADPPO-BQBZGAKWSA-N Arg-Gly-Ser Chemical compound NC(N)=NCCC[C@H](N)C(=O)NCC(=O)N[C@@H](CO)C(O)=O WVNFNPGXYADPPO-BQBZGAKWSA-N 0.000 description 1
- GMFAGHNRXPSSJS-SRVKXCTJSA-N Arg-Leu-Gln Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CCC(N)=O)C(O)=O GMFAGHNRXPSSJS-SRVKXCTJSA-N 0.000 description 1
- FSNVAJOPUDVQAR-AVGNSLFASA-N Arg-Lys-Arg Chemical compound NC(=N)NCCC[C@H](N)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O FSNVAJOPUDVQAR-AVGNSLFASA-N 0.000 description 1
- CVXXSWQORBZAAA-SRVKXCTJSA-N Arg-Lys-Glu Chemical compound OC(=O)CC[C@@H](C(O)=O)NC(=O)[C@H](CCCCN)NC(=O)[C@@H](N)CCCN=C(N)N CVXXSWQORBZAAA-SRVKXCTJSA-N 0.000 description 1
- VIINVRPKMUZYOI-DCAQKATOSA-N Arg-Met-Glu Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CCSC)C(=O)N[C@@H](CCC(O)=O)C(O)=O VIINVRPKMUZYOI-DCAQKATOSA-N 0.000 description 1
- DNBMCNQKNOKOSD-DCAQKATOSA-N Arg-Pro-Gln Chemical compound NC(N)=NCCC[C@H](N)C(=O)N1CCC[C@H]1C(=O)N[C@@H](CCC(N)=O)C(O)=O DNBMCNQKNOKOSD-DCAQKATOSA-N 0.000 description 1
- ASQKVGRCKOFKIU-KZVJFYERSA-N Arg-Thr-Ala Chemical compound C[C@H]([C@@H](C(=O)N[C@@H](C)C(=O)O)NC(=O)[C@H](CCCN=C(N)N)N)O ASQKVGRCKOFKIU-KZVJFYERSA-N 0.000 description 1
- AIFHRTPABBBHKU-RCWTZXSCSA-N Arg-Thr-Arg Chemical compound NC(N)=NCCC[C@H](N)C(=O)N[C@@H]([C@H](O)C)C(=O)N[C@@H](CCCN=C(N)N)C(O)=O AIFHRTPABBBHKU-RCWTZXSCSA-N 0.000 description 1
- ULBHWNVWSCJLCO-NHCYSSNCSA-N Arg-Val-Glu Chemical compound OC(=O)CC[C@@H](C(O)=O)NC(=O)[C@H](C(C)C)NC(=O)[C@@H](N)CCCN=C(N)N ULBHWNVWSCJLCO-NHCYSSNCSA-N 0.000 description 1
- GMRGSBAMMMVDGG-GUBZILKMSA-N Asn-Arg-Arg Chemical compound C(C[C@@H](C(=O)N[C@@H](CCCN=C(N)N)C(=O)O)NC(=O)[C@H](CC(=O)N)N)CN=C(N)N GMRGSBAMMMVDGG-GUBZILKMSA-N 0.000 description 1
- COUZKSSMBFADSB-AVGNSLFASA-N Asn-Glu-Phe Chemical compound C1=CC=C(C=C1)C[C@@H](C(=O)O)NC(=O)[C@H](CCC(=O)O)NC(=O)[C@H](CC(=O)N)N COUZKSSMBFADSB-AVGNSLFASA-N 0.000 description 1
- GKKUBLFXKRDMFC-BQBZGAKWSA-N Asn-Pro-Gly Chemical compound [H]N[C@@H](CC(N)=O)C(=O)N1CCC[C@H]1C(=O)NCC(O)=O GKKUBLFXKRDMFC-BQBZGAKWSA-N 0.000 description 1
- YQPSDMUGFKJZHR-QRTARXTBSA-N Asn-Trp-Val Chemical compound CC(C)[C@@H](C(=O)O)NC(=O)[C@H](CC1=CNC2=CC=CC=C21)NC(=O)[C@H](CC(=O)N)N YQPSDMUGFKJZHR-QRTARXTBSA-N 0.000 description 1
- WSGVTKZFVJSJOG-RCOVLWMOSA-N Asp-Gly-Val Chemical compound [H]N[C@@H](CC(O)=O)C(=O)NCC(=O)N[C@@H](C(C)C)C(O)=O WSGVTKZFVJSJOG-RCOVLWMOSA-N 0.000 description 1
- GXIUDSXIUSTSLO-QXEWZRGKSA-N Asp-Val-Met Chemical compound CC(C)[C@@H](C(=O)N[C@@H](CCSC)C(=O)O)NC(=O)[C@H](CC(=O)O)N GXIUDSXIUSTSLO-QXEWZRGKSA-N 0.000 description 1
- 102000008096 B7-H1 Antigen Human genes 0.000 description 1
- 108010074708 B7-H1 Antigen Proteins 0.000 description 1
- UKVGHFORADMBEN-GUBZILKMSA-N Cys-Arg-Arg Chemical compound [H]N[C@@H](CS)C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O UKVGHFORADMBEN-GUBZILKMSA-N 0.000 description 1
- CPTUXCUWQIBZIF-ZLUOBGJFSA-N Cys-Asn-Ser Chemical compound SC[C@H](N)C(=O)N[C@@H](CC(N)=O)C(=O)N[C@@H](CO)C(O)=O CPTUXCUWQIBZIF-ZLUOBGJFSA-N 0.000 description 1
- CFQVGYWKSLKWFX-KBIXCLLPSA-N Cys-Glu-Ile Chemical compound [H]N[C@@H](CS)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H]([C@@H](C)CC)C(O)=O CFQVGYWKSLKWFX-KBIXCLLPSA-N 0.000 description 1
- UCSXXFRXHGUXCQ-SRVKXCTJSA-N Cys-Leu-Lys Chemical compound CC(C)C[C@@H](C(=O)N[C@@H](CCCCN)C(=O)O)NC(=O)[C@H](CS)N UCSXXFRXHGUXCQ-SRVKXCTJSA-N 0.000 description 1
- FCXJJTRGVAZDER-FXQIFTODSA-N Cys-Val-Ala Chemical compound [H]N[C@@H](CS)C(=O)N[C@@H](C(C)C)C(=O)N[C@@H](C)C(O)=O FCXJJTRGVAZDER-FXQIFTODSA-N 0.000 description 1
- 102100039498 Cytotoxic T-lymphocyte protein 4 Human genes 0.000 description 1
- RZSLYUUFFVHFRQ-FXQIFTODSA-N Gln-Ala-Glu Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H](C)C(=O)N[C@@H](CCC(O)=O)C(O)=O RZSLYUUFFVHFRQ-FXQIFTODSA-N 0.000 description 1
- JFSNBQJNDMXMQF-XHNCKOQMSA-N Gln-Asp-Pro Chemical compound C1C[C@@H](N(C1)C(=O)[C@H](CC(=O)O)NC(=O)[C@H](CCC(=O)N)N)C(=O)O JFSNBQJNDMXMQF-XHNCKOQMSA-N 0.000 description 1
- SNLOOPZHAQDMJG-CIUDSAMLSA-N Gln-Glu-Glu Chemical compound NC(=O)CC[C@H](N)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCC(O)=O)C(O)=O SNLOOPZHAQDMJG-CIUDSAMLSA-N 0.000 description 1
- IULKWYSYZSURJK-AVGNSLFASA-N Gln-Leu-Lys Chemical compound NC(=O)CC[C@H](N)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CCCCN)C(O)=O IULKWYSYZSURJK-AVGNSLFASA-N 0.000 description 1
- MQJDLNRXBOELJW-KKUMJFAQSA-N Gln-Pro-Phe Chemical compound N[C@@H](CCC(N)=O)C(=O)N1CCC[C@H]1C(=O)N[C@@H](Cc1ccccc1)C(O)=O MQJDLNRXBOELJW-KKUMJFAQSA-N 0.000 description 1
- FVEMBYKESRUFBG-SZMVWBNQSA-N Gln-Trp-His Chemical compound C1=CC=C2C(=C1)C(=CN2)C[C@@H](C(=O)N[C@@H](CC3=CN=CN3)C(=O)O)NC(=O)[C@H](CCC(=O)N)N FVEMBYKESRUFBG-SZMVWBNQSA-N 0.000 description 1
- IIMZHVKZBGSEKZ-SZMVWBNQSA-N Gln-Trp-Leu Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H](CC1=CNC2=C1C=CC=C2)C(=O)N[C@@H](CC(C)C)C(O)=O IIMZHVKZBGSEKZ-SZMVWBNQSA-N 0.000 description 1
- MXPBQDFWIMBACQ-ACZMJKKPSA-N Glu-Cys-Cys Chemical compound OC(=O)CC[C@H](N)C(=O)N[C@@H](CS)C(=O)N[C@@H](CS)C(O)=O MXPBQDFWIMBACQ-ACZMJKKPSA-N 0.000 description 1
- BUAKRRKDHSSIKK-IHRRRGAJSA-N Glu-Glu-Tyr Chemical compound OC(=O)CC[C@H](N)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@H](C(O)=O)CC1=CC=C(O)C=C1 BUAKRRKDHSSIKK-IHRRRGAJSA-N 0.000 description 1
- QXDXIXFSFHUYAX-MNXVOIDGSA-N Glu-Ile-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)[C@H]([C@@H](C)CC)NC(=O)[C@@H](N)CCC(O)=O QXDXIXFSFHUYAX-MNXVOIDGSA-N 0.000 description 1
- HVYWQYLBVXMXSV-GUBZILKMSA-N Glu-Leu-Ala Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](C)C(O)=O HVYWQYLBVXMXSV-GUBZILKMSA-N 0.000 description 1
- RBXSZQRSEGYDFG-GUBZILKMSA-N Glu-Lys-Ser Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CO)C(O)=O RBXSZQRSEGYDFG-GUBZILKMSA-N 0.000 description 1
- QNJNPKSWAHPYGI-JYJNAYRXSA-N Glu-Phe-Leu Chemical compound OC(=O)CC[C@H](N)C(=O)N[C@H](C(=O)N[C@@H](CC(C)C)C(O)=O)CC1=CC=CC=C1 QNJNPKSWAHPYGI-JYJNAYRXSA-N 0.000 description 1
- ZIYGTCDTJJCDDP-JYJNAYRXSA-N Glu-Phe-Lys Chemical compound C1=CC=C(C=C1)C[C@@H](C(=O)N[C@@H](CCCCN)C(=O)O)NC(=O)[C@H](CCC(=O)O)N ZIYGTCDTJJCDDP-JYJNAYRXSA-N 0.000 description 1
- RXJFSLQVMGYQEL-IHRRRGAJSA-N Glu-Tyr-Glu Chemical compound OC(=O)CC[C@H](N)C(=O)N[C@H](C(=O)N[C@@H](CCC(O)=O)C(O)=O)CC1=CC=C(O)C=C1 RXJFSLQVMGYQEL-IHRRRGAJSA-N 0.000 description 1
- FKJQNJCQTKUBCD-XPUUQOCRSA-N Gly-Ala-His Chemical compound NCC(=O)N[C@@H](C)C(=O)N[C@@H](CC1=CNC=N1)C(=O)O FKJQNJCQTKUBCD-XPUUQOCRSA-N 0.000 description 1
- QXPRJQPCFXMCIY-NKWVEPMBSA-N Gly-Ala-Pro Chemical compound C[C@@H](C(=O)N1CCC[C@@H]1C(=O)O)NC(=O)CN QXPRJQPCFXMCIY-NKWVEPMBSA-N 0.000 description 1
- LERGJIVJIIODPZ-ZANVPECISA-N Gly-Ala-Trp Chemical compound C1=CC=C2C(C[C@H](NC(=O)[C@@H](NC(=O)CN)C)C(O)=O)=CNC2=C1 LERGJIVJIIODPZ-ZANVPECISA-N 0.000 description 1
- XRTDOIOIBMAXCT-NKWVEPMBSA-N Gly-Asn-Pro Chemical compound C1C[C@@H](N(C1)C(=O)[C@H](CC(=O)N)NC(=O)CN)C(=O)O XRTDOIOIBMAXCT-NKWVEPMBSA-N 0.000 description 1
- GDOZQTNZPCUARW-YFKPBYRVSA-N Gly-Gly-Glu Chemical compound NCC(=O)NCC(=O)N[C@H](C(O)=O)CCC(O)=O GDOZQTNZPCUARW-YFKPBYRVSA-N 0.000 description 1
- IEGFSKKANYKBDU-QWHCGFSZSA-N Gly-Phe-Pro Chemical compound C1C[C@@H](N(C1)C(=O)[C@H](CC2=CC=CC=C2)NC(=O)CN)C(=O)O IEGFSKKANYKBDU-QWHCGFSZSA-N 0.000 description 1
- WDXLKVQATNEAJQ-BQBZGAKWSA-N Gly-Pro-Asp Chemical compound NCC(=O)N1CCC[C@H]1C(=O)N[C@@H](CC(O)=O)C(O)=O WDXLKVQATNEAJQ-BQBZGAKWSA-N 0.000 description 1
- HAOUOFNNJJLVNS-BQBZGAKWSA-N Gly-Pro-Ser Chemical compound NCC(=O)N1CCC[C@H]1C(=O)N[C@@H](CO)C(O)=O HAOUOFNNJJLVNS-BQBZGAKWSA-N 0.000 description 1
- GWCJMBNBFYBQCV-XPUUQOCRSA-N Gly-Val-Ala Chemical compound NCC(=O)N[C@@H](C(C)C)C(=O)N[C@@H](C)C(O)=O GWCJMBNBFYBQCV-XPUUQOCRSA-N 0.000 description 1
- WZOGEMJIZBNFBK-CIUDSAMLSA-N His-Asp-Asn Chemical compound [H]N[C@@H](CC1=CNC=N1)C(=O)N[C@@H](CC(O)=O)C(=O)N[C@@H](CC(N)=O)C(O)=O WZOGEMJIZBNFBK-CIUDSAMLSA-N 0.000 description 1
- HQKADFMLECZIQJ-HVTMNAMFSA-N His-Glu-Ile Chemical compound CC[C@H](C)[C@@H](C(=O)O)NC(=O)[C@H](CCC(=O)O)NC(=O)[C@H](CC1=CN=CN1)N HQKADFMLECZIQJ-HVTMNAMFSA-N 0.000 description 1
- CHZRWFUGWRTUOD-IUCAKERBSA-N His-Gly-Gln Chemical compound C1=C(NC=N1)C[C@@H](C(=O)NCC(=O)N[C@@H](CCC(=O)N)C(=O)O)N CHZRWFUGWRTUOD-IUCAKERBSA-N 0.000 description 1
- LBQAHBIVXQSBIR-HVTMNAMFSA-N His-Ile-Glu Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CCC(=O)O)C(=O)O)NC(=O)[C@H](CC1=CN=CN1)N LBQAHBIVXQSBIR-HVTMNAMFSA-N 0.000 description 1
- 101000889276 Homo sapiens Cytotoxic T-lymphocyte protein 4 Proteins 0.000 description 1
- 101001032342 Homo sapiens Interferon regulatory factor 7 Proteins 0.000 description 1
- 101000665442 Homo sapiens Serine/threonine-protein kinase TBK1 Proteins 0.000 description 1
- 101150074358 IFIT2 gene Proteins 0.000 description 1
- QSPLUJGYOPZINY-ZPFDUUQYSA-N Ile-Asp-Lys Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CC(=O)O)C(=O)N[C@@H](CCCCN)C(=O)O)N QSPLUJGYOPZINY-ZPFDUUQYSA-N 0.000 description 1
- KUHFPGIVBOCRMV-MNXVOIDGSA-N Ile-Gln-Leu Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CCC(=O)N)C(=O)N[C@@H](CC(C)C)C(=O)O)N KUHFPGIVBOCRMV-MNXVOIDGSA-N 0.000 description 1
- MTFVYKQRLXYAQN-LAEOZQHASA-N Ile-Glu-Gly Chemical compound [H]N[C@@H]([C@@H](C)CC)C(=O)N[C@@H](CCC(O)=O)C(=O)NCC(O)=O MTFVYKQRLXYAQN-LAEOZQHASA-N 0.000 description 1
- TWPSALMCEHCIOY-YTFOTSKYSA-N Ile-Ile-Leu Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H]([C@@H](C)CC)C(=O)N[C@@H](CC(C)C)C(=O)O)N TWPSALMCEHCIOY-YTFOTSKYSA-N 0.000 description 1
- OVDKXUDMKXAZIV-ZPFDUUQYSA-N Ile-Lys-Asn Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CC(=O)N)C(=O)O)N OVDKXUDMKXAZIV-ZPFDUUQYSA-N 0.000 description 1
- VGSPNSSCMOHRRR-BJDJZHNGSA-N Ile-Ser-Lys Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CO)C(=O)N[C@@H](CCCCN)C(=O)O)N VGSPNSSCMOHRRR-BJDJZHNGSA-N 0.000 description 1
- 108010065920 Insulin Lispro Proteins 0.000 description 1
- 102100029843 Interferon regulatory factor 3 Human genes 0.000 description 1
- 102100038070 Interferon regulatory factor 7 Human genes 0.000 description 1
- 108010074328 Interferon-gamma Proteins 0.000 description 1
- 102100027303 Interferon-induced protein with tetratricopeptide repeats 2 Human genes 0.000 description 1
- 102000014150 Interferons Human genes 0.000 description 1
- 108010002350 Interleukin-2 Proteins 0.000 description 1
- PMGDADKJMCOXHX-UHFFFAOYSA-N L-Arginyl-L-glutamin-acetat Natural products NC(=N)NCCCC(N)C(=O)NC(CCC(N)=O)C(O)=O PMGDADKJMCOXHX-UHFFFAOYSA-N 0.000 description 1
- HGCNKOLVKRAVHD-UHFFFAOYSA-N L-Met-L-Phe Natural products CSCCC(N)C(=O)NC(C(O)=O)CC1=CC=CC=C1 HGCNKOLVKRAVHD-UHFFFAOYSA-N 0.000 description 1
- SITWEMZOJNKJCH-UHFFFAOYSA-N L-alanine-L-arginine Natural products CC(N)C(=O)NC(C(O)=O)CCCNC(N)=N SITWEMZOJNKJCH-UHFFFAOYSA-N 0.000 description 1
- SENJXOPIZNYLHU-UHFFFAOYSA-N L-leucyl-L-arginine Natural products CC(C)CC(N)C(=O)NC(C(O)=O)CCCN=C(N)N SENJXOPIZNYLHU-UHFFFAOYSA-N 0.000 description 1
- 239000012880 LB liquid culture medium Substances 0.000 description 1
- CZCSUZMIRKFFFA-CIUDSAMLSA-N Leu-Ala-Asn Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](C)C(=O)N[C@@H](CC(N)=O)C(O)=O CZCSUZMIRKFFFA-CIUDSAMLSA-N 0.000 description 1
- KVMULWOHPPMHHE-DCAQKATOSA-N Leu-Glu-Gln Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCC(N)=O)C(O)=O KVMULWOHPPMHHE-DCAQKATOSA-N 0.000 description 1
- HQUXQAMSWFIRET-AVGNSLFASA-N Leu-Glu-Lys Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@H](C(O)=O)CCCCN HQUXQAMSWFIRET-AVGNSLFASA-N 0.000 description 1
- QNBVTHNJGCOVFA-AVGNSLFASA-N Leu-Leu-Glu Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](CC(C)C)C(=O)N[C@H](C(O)=O)CCC(O)=O QNBVTHNJGCOVFA-AVGNSLFASA-N 0.000 description 1
- JLWZLIQRYCTYBD-IHRRRGAJSA-N Leu-Lys-Arg Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O JLWZLIQRYCTYBD-IHRRRGAJSA-N 0.000 description 1
- WXUOJXIGOPMDJM-SRVKXCTJSA-N Leu-Lys-Asn Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CC(N)=O)C(O)=O WXUOJXIGOPMDJM-SRVKXCTJSA-N 0.000 description 1
- BGZCJDGBBUUBHA-KKUMJFAQSA-N Leu-Lys-Leu Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CC(C)C)C(O)=O BGZCJDGBBUUBHA-KKUMJFAQSA-N 0.000 description 1
- PTRKPHUGYULXPU-KKUMJFAQSA-N Leu-Phe-Ser Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](CC1=CC=CC=C1)C(=O)N[C@@H](CO)C(O)=O PTRKPHUGYULXPU-KKUMJFAQSA-N 0.000 description 1
- IDGZVZJLYFTXSL-DCAQKATOSA-N Leu-Ser-Arg Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](CO)C(=O)N[C@H](C(O)=O)CCCN=C(N)N IDGZVZJLYFTXSL-DCAQKATOSA-N 0.000 description 1
- 241001424413 Lucia Species 0.000 description 1
- ABHIXYDMILIUKV-CIUDSAMLSA-N Lys-Asn-Asn Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CC(N)=O)C(=O)N[C@@H](CC(N)=O)C(O)=O ABHIXYDMILIUKV-CIUDSAMLSA-N 0.000 description 1
- FACUGMGEFUEBTI-SRVKXCTJSA-N Lys-Asn-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)[C@H](CC(N)=O)NC(=O)[C@@H](N)CCCCN FACUGMGEFUEBTI-SRVKXCTJSA-N 0.000 description 1
- WGCKDDHUFPQSMZ-ZPFDUUQYSA-N Lys-Asp-Ile Chemical compound CC[C@H](C)[C@@H](C(O)=O)NC(=O)[C@H](CC(O)=O)NC(=O)[C@@H](N)CCCCN WGCKDDHUFPQSMZ-ZPFDUUQYSA-N 0.000 description 1
- CFVQPNSCQMKDPB-CIUDSAMLSA-N Lys-Cys-Cys Chemical compound C(CCN)C[C@@H](C(=O)N[C@@H](CS)C(=O)N[C@@H](CS)C(=O)O)N CFVQPNSCQMKDPB-CIUDSAMLSA-N 0.000 description 1
- YVMQJGWLHRWMDF-MNXVOIDGSA-N Lys-Gln-Ile Chemical compound CC[C@H](C)[C@@H](C(=O)O)NC(=O)[C@H](CCC(=O)N)NC(=O)[C@H](CCCCN)N YVMQJGWLHRWMDF-MNXVOIDGSA-N 0.000 description 1
- PBIPLDMFHAICIP-DCAQKATOSA-N Lys-Glu-Glu Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCC(O)=O)C(O)=O PBIPLDMFHAICIP-DCAQKATOSA-N 0.000 description 1
- GCMWRRQAKQXDED-IUCAKERBSA-N Lys-Glu-Gly Chemical compound [NH3+]CCCC[C@H]([NH3+])C(=O)N[C@@H](CCC([O-])=O)C(=O)NCC([O-])=O GCMWRRQAKQXDED-IUCAKERBSA-N 0.000 description 1
- LPAJOCKCPRZEAG-MNXVOIDGSA-N Lys-Glu-Ile Chemical compound CC[C@H](C)[C@@H](C(O)=O)NC(=O)[C@H](CCC(O)=O)NC(=O)[C@@H](N)CCCCN LPAJOCKCPRZEAG-MNXVOIDGSA-N 0.000 description 1
- GQFDWEDHOQRNLC-QWRGUYRKSA-N Lys-Gly-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)CNC(=O)[C@@H](N)CCCCN GQFDWEDHOQRNLC-QWRGUYRKSA-N 0.000 description 1
- KZJQUYFDSCFSCO-DLOVCJGASA-N Lys-His-Ala Chemical compound C[C@@H](C(=O)O)NC(=O)[C@H](CC1=CN=CN1)NC(=O)[C@H](CCCCN)N KZJQUYFDSCFSCO-DLOVCJGASA-N 0.000 description 1
- JYXBNQOKPRQNQS-YTFOTSKYSA-N Lys-Ile-Ile Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H]([C@@H](C)CC)C(=O)N[C@@H]([C@@H](C)CC)C(O)=O JYXBNQOKPRQNQS-YTFOTSKYSA-N 0.000 description 1
- ONPDTSFZAIWMDI-AVGNSLFASA-N Lys-Leu-Gln Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CCC(N)=O)C(O)=O ONPDTSFZAIWMDI-AVGNSLFASA-N 0.000 description 1
- ATNKHRAIZCMCCN-BZSNNMDCSA-N Lys-Lys-Phe Chemical compound C1=CC=C(C=C1)C[C@@H](C(=O)O)NC(=O)[C@H](CCCCN)NC(=O)[C@H](CCCCN)N ATNKHRAIZCMCCN-BZSNNMDCSA-N 0.000 description 1
- VSTNAUBHKQPVJX-IHRRRGAJSA-N Lys-Met-Leu Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CCSC)C(=O)N[C@@H](CC(C)C)C(O)=O VSTNAUBHKQPVJX-IHRRRGAJSA-N 0.000 description 1
- JPYPRVHMKRFTAT-KKUMJFAQSA-N Lys-Phe-Cys Chemical compound C1=CC=C(C=C1)C[C@@H](C(=O)N[C@@H](CS)C(=O)O)NC(=O)[C@H](CCCCN)N JPYPRVHMKRFTAT-KKUMJFAQSA-N 0.000 description 1
- MIFFFXHMAHFACR-KATARQTJSA-N Lys-Ser-Thr Chemical compound C[C@@H](O)[C@@H](C(O)=O)NC(=O)[C@H](CO)NC(=O)[C@@H](N)CCCCN MIFFFXHMAHFACR-KATARQTJSA-N 0.000 description 1
- TVOOGUNBIWAURO-KATARQTJSA-N Lys-Thr-Cys Chemical compound C[C@H]([C@@H](C(=O)N[C@@H](CS)C(=O)O)NC(=O)[C@H](CCCCN)N)O TVOOGUNBIWAURO-KATARQTJSA-N 0.000 description 1
- OHXUUQDOBQKSNB-AVGNSLFASA-N Lys-Val-Arg Chemical compound NCCCC[C@H](N)C(=O)N[C@@H](C(C)C)C(=O)N[C@@H](CCCN=C(N)N)C(O)=O OHXUUQDOBQKSNB-AVGNSLFASA-N 0.000 description 1
- YRAWWKUTNBILNT-FXQIFTODSA-N Met-Ala-Ala Chemical compound CSCC[C@H](N)C(=O)N[C@@H](C)C(=O)N[C@@H](C)C(O)=O YRAWWKUTNBILNT-FXQIFTODSA-N 0.000 description 1
- AOFZWWDTTJLHOU-ULQDDVLXSA-N Met-Lys-Tyr Chemical compound CSCC[C@H](N)C(=O)N[C@@H](CCCCN)C(=O)N[C@H](C(O)=O)CC1=CC=C(O)C=C1 AOFZWWDTTJLHOU-ULQDDVLXSA-N 0.000 description 1
- XZFYRXDAULDNFX-UHFFFAOYSA-N N-L-cysteinyl-L-phenylalanine Natural products SCC(N)C(=O)NC(C(O)=O)CC1=CC=CC=C1 XZFYRXDAULDNFX-UHFFFAOYSA-N 0.000 description 1
- AUEJLPRZGVVDNU-UHFFFAOYSA-N N-L-tyrosyl-L-leucine Natural products CC(C)CC(C(O)=O)NC(=O)C(N)CC1=CC=C(O)C=C1 AUEJLPRZGVVDNU-UHFFFAOYSA-N 0.000 description 1
- 108010079364 N-glycylalanine Proteins 0.000 description 1
- 102000003945 NF-kappa B Human genes 0.000 description 1
- 108010057466 NF-kappa B Proteins 0.000 description 1
- WMGVYPPIMZPWPN-SRVKXCTJSA-N Phe-Asp-Asn Chemical compound C1=CC=C(C=C1)C[C@@H](C(=O)N[C@@H](CC(=O)O)C(=O)N[C@@H](CC(=O)N)C(=O)O)N WMGVYPPIMZPWPN-SRVKXCTJSA-N 0.000 description 1
- DMEYUTSDVRCWRS-ULQDDVLXSA-N Phe-Lys-Arg Chemical compound NC(=N)NCCC[C@@H](C(O)=O)NC(=O)[C@H](CCCCN)NC(=O)[C@@H](N)CC1=CC=CC=C1 DMEYUTSDVRCWRS-ULQDDVLXSA-N 0.000 description 1
- DOXQMJCSSYZSNM-BZSNNMDCSA-N Phe-Lys-Leu Chemical compound [H]N[C@@H](CC1=CC=CC=C1)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CC(C)C)C(O)=O DOXQMJCSSYZSNM-BZSNNMDCSA-N 0.000 description 1
- TXJJXEXCZBHDNA-ACRUOGEOSA-N Phe-Phe-His Chemical compound C1=CC=C(C=C1)C[C@@H](C(=O)N[C@@H](CC2=CC=CC=C2)C(=O)N[C@@H](CC3=CN=CN3)C(=O)O)N TXJJXEXCZBHDNA-ACRUOGEOSA-N 0.000 description 1
- RGMLUHANLDVMPB-ULQDDVLXSA-N Phe-Val-Lys Chemical compound CC(C)[C@@H](C(=O)N[C@@H](CCCCN)C(=O)O)NC(=O)[C@H](CC1=CC=CC=C1)N RGMLUHANLDVMPB-ULQDDVLXSA-N 0.000 description 1
- OOLOTUZJUBOMAX-GUBZILKMSA-N Pro-Ala-Val Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](C)C(=O)N[C@@H](C(C)C)C(O)=O OOLOTUZJUBOMAX-GUBZILKMSA-N 0.000 description 1
- NHDVNAKDACFHPX-GUBZILKMSA-N Pro-Arg-Ala Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](C)C(O)=O NHDVNAKDACFHPX-GUBZILKMSA-N 0.000 description 1
- IHCXPSYCHXFXKT-DCAQKATOSA-N Pro-Arg-Glu Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CCC(O)=O)C(O)=O IHCXPSYCHXFXKT-DCAQKATOSA-N 0.000 description 1
- UAYHMOIGIQZLFR-NHCYSSNCSA-N Pro-Gln-Val Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H](C(C)C)C(O)=O UAYHMOIGIQZLFR-NHCYSSNCSA-N 0.000 description 1
- CLNJSLSHKJECME-BQBZGAKWSA-N Pro-Gly-Ala Chemical compound OC(=O)[C@H](C)NC(=O)CNC(=O)[C@@H]1CCCN1 CLNJSLSHKJECME-BQBZGAKWSA-N 0.000 description 1
- HAEGAELAYWSUNC-WPRPVWTQSA-N Pro-Gly-Val Chemical compound [H]N1CCC[C@H]1C(=O)NCC(=O)N[C@@H](C(C)C)C(O)=O HAEGAELAYWSUNC-WPRPVWTQSA-N 0.000 description 1
- UREQLMJCKFLLHM-NAKRPEOUSA-N Pro-Ile-Ser Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H]([C@@H](C)CC)C(=O)N[C@@H](CO)C(O)=O UREQLMJCKFLLHM-NAKRPEOUSA-N 0.000 description 1
- FDMKYQQYJKYCLV-GUBZILKMSA-N Pro-Pro-Ser Chemical compound OC[C@@H](C(O)=O)NC(=O)[C@@H]1CCCN1C(=O)[C@H]1NCCC1 FDMKYQQYJKYCLV-GUBZILKMSA-N 0.000 description 1
- POQFNPILEQEODH-FXQIFTODSA-N Pro-Ser-Ala Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](CO)C(=O)N[C@@H](C)C(O)=O POQFNPILEQEODH-FXQIFTODSA-N 0.000 description 1
- FNGOXVQBBCMFKV-CIUDSAMLSA-N Pro-Ser-Glu Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](CO)C(=O)N[C@@H](CCC(O)=O)C(O)=O FNGOXVQBBCMFKV-CIUDSAMLSA-N 0.000 description 1
- SXJOPONICMGFCR-DCAQKATOSA-N Pro-Ser-Lys Chemical compound C1C[C@H](NC1)C(=O)N[C@@H](CO)C(=O)N[C@@H](CCCCN)C(=O)O SXJOPONICMGFCR-DCAQKATOSA-N 0.000 description 1
- 102100040678 Programmed cell death protein 1 Human genes 0.000 description 1
- 101710089372 Programmed cell death protein 1 Proteins 0.000 description 1
- 108700008625 Reporter Genes Proteins 0.000 description 1
- 108091027981 Response element Proteins 0.000 description 1
- FIXILCYTSAUERA-FXQIFTODSA-N Ser-Ala-Arg Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](C)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O FIXILCYTSAUERA-FXQIFTODSA-N 0.000 description 1
- BRKHVZNDAOMAHX-BIIVOSGPSA-N Ser-Ala-Pro Chemical compound C[C@@H](C(=O)N1CCC[C@@H]1C(=O)O)NC(=O)[C@H](CO)N BRKHVZNDAOMAHX-BIIVOSGPSA-N 0.000 description 1
- YQHZVYJAGWMHES-ZLUOBGJFSA-N Ser-Ala-Ser Chemical compound OC[C@H](N)C(=O)N[C@@H](C)C(=O)N[C@@H](CO)C(O)=O YQHZVYJAGWMHES-ZLUOBGJFSA-N 0.000 description 1
- BLPYXIXXCFVIIF-FXQIFTODSA-N Ser-Cys-Arg Chemical compound C(C[C@@H](C(=O)O)NC(=O)[C@H](CS)NC(=O)[C@H](CO)N)CN=C(N)N BLPYXIXXCFVIIF-FXQIFTODSA-N 0.000 description 1
- SMIDBHKWSYUBRZ-ACZMJKKPSA-N Ser-Glu-Ala Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](C)C(O)=O SMIDBHKWSYUBRZ-ACZMJKKPSA-N 0.000 description 1
- OHKFXGKHSJKKAL-NRPADANISA-N Ser-Glu-Val Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](C(C)C)C(O)=O OHKFXGKHSJKKAL-NRPADANISA-N 0.000 description 1
- JFWDJFULOLKQFY-QWRGUYRKSA-N Ser-Gly-Phe Chemical compound [H]N[C@@H](CO)C(=O)NCC(=O)N[C@@H](CC1=CC=CC=C1)C(O)=O JFWDJFULOLKQFY-QWRGUYRKSA-N 0.000 description 1
- VMLONWHIORGALA-SRVKXCTJSA-N Ser-Leu-Leu Chemical compound CC(C)C[C@@H](C([O-])=O)NC(=O)[C@H](CC(C)C)NC(=O)[C@@H]([NH3+])CO VMLONWHIORGALA-SRVKXCTJSA-N 0.000 description 1
- WGDYNRCOQRERLZ-KKUMJFAQSA-N Ser-Lys-Phe Chemical compound C1=CC=C(C=C1)C[C@@H](C(=O)O)NC(=O)[C@H](CCCCN)NC(=O)[C@H](CO)N WGDYNRCOQRERLZ-KKUMJFAQSA-N 0.000 description 1
- FPCGZYMRFFIYIH-CIUDSAMLSA-N Ser-Lys-Ser Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CO)C(O)=O FPCGZYMRFFIYIH-CIUDSAMLSA-N 0.000 description 1
- MQUZANJDFOQOBX-SRVKXCTJSA-N Ser-Phe-Ser Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CC1=CC=CC=C1)C(=O)N[C@@H](CO)C(O)=O MQUZANJDFOQOBX-SRVKXCTJSA-N 0.000 description 1
- SZRNDHWMVSFPSP-XKBZYTNZSA-N Ser-Thr-Gln Chemical compound C[C@H]([C@@H](C(=O)N[C@@H](CCC(=O)N)C(=O)O)NC(=O)[C@H](CO)N)O SZRNDHWMVSFPSP-XKBZYTNZSA-N 0.000 description 1
- HXPNJVLVHKABMJ-KKUMJFAQSA-N Ser-Tyr-His Chemical compound C1=CC(=CC=C1C[C@@H](C(=O)N[C@@H](CC2=CN=CN2)C(=O)O)NC(=O)[C@H](CO)N)O HXPNJVLVHKABMJ-KKUMJFAQSA-N 0.000 description 1
- PCMZJFMUYWIERL-ZKWXMUAHSA-N Ser-Val-Asn Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](C(C)C)C(=O)N[C@@H](CC(N)=O)C(O)=O PCMZJFMUYWIERL-ZKWXMUAHSA-N 0.000 description 1
- UKKROEYWYIHWBD-ZKWXMUAHSA-N Ser-Val-Asp Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](C(C)C)C(=O)N[C@@H](CC(O)=O)C(O)=O UKKROEYWYIHWBD-ZKWXMUAHSA-N 0.000 description 1
- 102100038192 Serine/threonine-protein kinase TBK1 Human genes 0.000 description 1
- MQCPGOZXFSYJPS-KZVJFYERSA-N Thr-Ala-Arg Chemical compound [H]N[C@@H]([C@@H](C)O)C(=O)N[C@@H](C)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O MQCPGOZXFSYJPS-KZVJFYERSA-N 0.000 description 1
- GKWNLDNXMMLRMC-GLLZPBPUSA-N Thr-Glu-Gln Chemical compound C[C@H]([C@@H](C(=O)N[C@@H](CCC(=O)O)C(=O)N[C@@H](CCC(=O)N)C(=O)O)N)O GKWNLDNXMMLRMC-GLLZPBPUSA-N 0.000 description 1
- LHEZGZQRLDBSRR-WDCWCFNPSA-N Thr-Glu-Leu Chemical compound [H]N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CC(C)C)C(O)=O LHEZGZQRLDBSRR-WDCWCFNPSA-N 0.000 description 1
- DJDSEDOKJTZBAR-ZDLURKLDSA-N Thr-Gly-Ser Chemical compound C[C@@H](O)[C@H](N)C(=O)NCC(=O)N[C@@H](CO)C(O)=O DJDSEDOKJTZBAR-ZDLURKLDSA-N 0.000 description 1
- MEJHFIOYJHTWMK-VOAKCMCISA-N Thr-Leu-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)[C@H](CC(C)C)NC(=O)[C@@H](N)[C@@H](C)O MEJHFIOYJHTWMK-VOAKCMCISA-N 0.000 description 1
- ZOCJFNXUVSGBQI-HSHDSVGOSA-N Thr-Trp-Arg Chemical compound C[C@H]([C@@H](C(=O)N[C@@H](CC1=CNC2=CC=CC=C21)C(=O)N[C@@H](CCCN=C(N)N)C(=O)O)N)O ZOCJFNXUVSGBQI-HSHDSVGOSA-N 0.000 description 1
- BKVICMPZWRNWOC-RHYQMDGZSA-N Thr-Val-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)[C@H](C(C)C)NC(=O)[C@@H](N)[C@@H](C)O BKVICMPZWRNWOC-RHYQMDGZSA-N 0.000 description 1
- XGFOXYJQBRTJPO-PJODQICGSA-N Trp-Pro-Ala Chemical compound [H]N[C@@H](CC1=CNC2=C1C=CC=C2)C(=O)N1CCC[C@H]1C(=O)N[C@@H](C)C(O)=O XGFOXYJQBRTJPO-PJODQICGSA-N 0.000 description 1
- CDKZJGMPZHPAJC-ULQDDVLXSA-N Tyr-Leu-Val Chemical compound CC(C)[C@@H](C(O)=O)NC(=O)[C@H](CC(C)C)NC(=O)[C@@H](N)CC1=CC=C(O)C=C1 CDKZJGMPZHPAJC-ULQDDVLXSA-N 0.000 description 1
- FASACHWGQBNSRO-ZEWNOJEFSA-N Tyr-Phe-Ile Chemical compound CC[C@H](C)[C@@H](C(=O)O)NC(=O)[C@H](CC1=CC=CC=C1)NC(=O)[C@H](CC2=CC=C(C=C2)O)N FASACHWGQBNSRO-ZEWNOJEFSA-N 0.000 description 1
- SCZJKZLFSSPJDP-ACRUOGEOSA-N Tyr-Phe-Leu Chemical compound [H]N[C@@H](CC1=CC=C(O)C=C1)C(=O)N[C@@H](CC1=CC=CC=C1)C(=O)N[C@@H](CC(C)C)C(O)=O SCZJKZLFSSPJDP-ACRUOGEOSA-N 0.000 description 1
- JQOMHZMWQHXALX-FHWLQOOXSA-N Tyr-Tyr-Glu Chemical compound [H]N[C@@H](CC1=CC=C(O)C=C1)C(=O)N[C@@H](CC1=CC=C(O)C=C1)C(=O)N[C@@H](CCC(O)=O)C(O)=O JQOMHZMWQHXALX-FHWLQOOXSA-N 0.000 description 1
- LABUITCFCAABSV-UHFFFAOYSA-N Val-Ala-Tyr Natural products CC(C)C(N)C(=O)NC(C)C(=O)NC(C(O)=O)CC1=CC=C(O)C=C1 LABUITCFCAABSV-UHFFFAOYSA-N 0.000 description 1
- IRLYZKKNBFPQBW-XGEHTFHBSA-N Val-Cys-Thr Chemical compound C[C@H]([C@@H](C(=O)O)NC(=O)[C@H](CS)NC(=O)[C@H](C(C)C)N)O IRLYZKKNBFPQBW-XGEHTFHBSA-N 0.000 description 1
- YDPFWRVQHFWBKI-GVXVVHGQSA-N Val-Glu-His Chemical compound CC(C)[C@@H](C(=O)N[C@@H](CCC(=O)O)C(=O)N[C@@H](CC1=CN=CN1)C(=O)O)N YDPFWRVQHFWBKI-GVXVVHGQSA-N 0.000 description 1
- QRVPEKJBBRYISE-XUXIUFHCSA-N Val-Lys-Ile Chemical compound CC[C@H](C)[C@@H](C(=O)O)NC(=O)[C@H](CCCCN)NC(=O)[C@H](C(C)C)N QRVPEKJBBRYISE-XUXIUFHCSA-N 0.000 description 1
- XPKCFQZDQGVJCX-RHYQMDGZSA-N Val-Lys-Thr Chemical compound C[C@H]([C@@H](C(=O)O)NC(=O)[C@H](CCCCN)NC(=O)[C@H](C(C)C)N)O XPKCFQZDQGVJCX-RHYQMDGZSA-N 0.000 description 1
- JMCOXFSCTGKLLB-FKBYEOEOSA-N Val-Phe-Trp Chemical compound CC(C)[C@@H](C(=O)N[C@@H](CC1=CC=CC=C1)C(=O)N[C@@H](CC2=CNC3=CC=CC=C32)C(=O)O)N JMCOXFSCTGKLLB-FKBYEOEOSA-N 0.000 description 1
- YTNGABPUXFEOGU-SRVKXCTJSA-N Val-Pro-Arg Chemical compound CC(C)[C@H](N)C(=O)N1CCC[C@H]1C(=O)N[C@@H](CCCN=C(N)N)C(O)=O YTNGABPUXFEOGU-SRVKXCTJSA-N 0.000 description 1
- RYQUMYBMOJYYDK-NHCYSSNCSA-N Val-Pro-Glu Chemical compound CC(C)[C@@H](C(=O)N1CCC[C@H]1C(=O)N[C@@H](CCC(=O)O)C(=O)O)N RYQUMYBMOJYYDK-NHCYSSNCSA-N 0.000 description 1
- HTONZBWRYUKUKC-RCWTZXSCSA-N Val-Thr-Val Chemical compound CC(C)[C@H](N)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](C(C)C)C(O)=O HTONZBWRYUKUKC-RCWTZXSCSA-N 0.000 description 1
- 208000036142 Viral infection Diseases 0.000 description 1
- 238000002835 absorbance Methods 0.000 description 1
- 238000001042 affinity chromatography Methods 0.000 description 1
- 108010008685 alanyl-glutamyl-aspartic acid Proteins 0.000 description 1
- 108010024078 alanyl-glycyl-serine Proteins 0.000 description 1
- 108010086434 alanyl-seryl-glycine Proteins 0.000 description 1
- 108010087924 alanylproline Proteins 0.000 description 1
- 125000000539 amino acid group Chemical group 0.000 description 1
- 230000002924 anti-infective effect Effects 0.000 description 1
- 230000006907 apoptotic process Effects 0.000 description 1
- 108010008355 arginyl-glutamine Proteins 0.000 description 1
- 108010018691 arginyl-threonyl-arginine Proteins 0.000 description 1
- 108010068380 arginylarginine Proteins 0.000 description 1
- 108010060035 arginylproline Proteins 0.000 description 1
- 108010093581 aspartyl-proline Proteins 0.000 description 1
- 238000003556 assay Methods 0.000 description 1
- 230000001580 bacterial effect Effects 0.000 description 1
- 238000002306 biochemical method Methods 0.000 description 1
- 238000007036 catalytic synthesis reaction Methods 0.000 description 1
- 239000012930 cell culture fluid Substances 0.000 description 1
- 238000012512 characterization method Methods 0.000 description 1
- 238000002512 chemotherapy Methods 0.000 description 1
- 238000004587 chromatography analysis Methods 0.000 description 1
- 238000003501 co-culture Methods 0.000 description 1
- 108010004073 cysteinylcysteine Proteins 0.000 description 1
- 230000009089 cytolysis Effects 0.000 description 1
- 230000029087 digestion Effects 0.000 description 1
- FSXRLASFHBWESK-UHFFFAOYSA-N dipeptide phenylalanyl-tyrosine Natural products C=1C=C(O)C=CC=1CC(C(O)=O)NC(=O)C(N)CC1=CC=CC=C1 FSXRLASFHBWESK-UHFFFAOYSA-N 0.000 description 1
- 229940042399 direct acting antivirals protease inhibitors Drugs 0.000 description 1
- 201000010099 disease Diseases 0.000 description 1
- 208000037265 diseases, disorders, signs and symptoms Diseases 0.000 description 1
- 239000012154 double-distilled water Substances 0.000 description 1
- 238000001962 electrophoresis Methods 0.000 description 1
- 108010078144 glutaminyl-glycine Proteins 0.000 description 1
- 108010008237 glutamyl-valyl-glycine Proteins 0.000 description 1
- VPZXBVLAVMBEQI-UHFFFAOYSA-N glycyl-DL-alpha-alanine Natural products OC(=O)C(C)NC(=O)CN VPZXBVLAVMBEQI-UHFFFAOYSA-N 0.000 description 1
- JYPCXBJRLBHWME-UHFFFAOYSA-N glycyl-L-prolyl-L-arginine Natural products NCC(=O)N1CCCC1C(=O)NC(CCCN=C(N)N)C(O)=O JYPCXBJRLBHWME-UHFFFAOYSA-N 0.000 description 1
- 108010015792 glycyllysine Proteins 0.000 description 1
- 108010037850 glycylvaline Proteins 0.000 description 1
- 239000001963 growth medium Substances 0.000 description 1
- 108010025306 histidylleucine Proteins 0.000 description 1
- 210000000987 immune system Anatomy 0.000 description 1
- 239000000568 immunological adjuvant Substances 0.000 description 1
- 230000006872 improvement Effects 0.000 description 1
- 230000006698 induction Effects 0.000 description 1
- 230000001939 inductive effect Effects 0.000 description 1
- 208000015181 infectious disease Diseases 0.000 description 1
- 230000015788 innate immune response Effects 0.000 description 1
- 229940079322 interferon Drugs 0.000 description 1
- 231100000225 lethality Toxicity 0.000 description 1
- 108010000761 leucylarginine Proteins 0.000 description 1
- 108010057821 leucylproline Proteins 0.000 description 1
- 108010064235 lysylglycine Proteins 0.000 description 1
- 239000000463 material Substances 0.000 description 1
- 230000001404 mediated effect Effects 0.000 description 1
- 108010068488 methionylphenylalanine Proteins 0.000 description 1
- 238000001471 micro-filtration Methods 0.000 description 1
- 239000008188 pellet Substances 0.000 description 1
- 230000026731 phosphorylation Effects 0.000 description 1
- 238000006366 phosphorylation reaction Methods 0.000 description 1
- 239000002244 precipitate Substances 0.000 description 1
- 230000008569 process Effects 0.000 description 1
- 108010029020 prolylglycine Proteins 0.000 description 1
- 108010053725 prolylvaline Proteins 0.000 description 1
- 238000012113 quantitative test Methods 0.000 description 1
- 230000035945 sensitivity Effects 0.000 description 1
- 238000012163 sequencing technique Methods 0.000 description 1
- 108010026333 seryl-proline Proteins 0.000 description 1
- 108010005652 splenotritin Proteins 0.000 description 1
- 230000002194 synthesizing effect Effects 0.000 description 1
- 230000009885 systemic effect Effects 0.000 description 1
- 238000012360 testing method Methods 0.000 description 1
- 230000000451 tissue damage Effects 0.000 description 1
- 231100000827 tissue damage Toxicity 0.000 description 1
- 238000012546 transfer Methods 0.000 description 1
- 108010078580 tyrosylleucine Proteins 0.000 description 1
- 108010073969 valyllysine Proteins 0.000 description 1
- 238000012795 verification Methods 0.000 description 1
- 230000009385 viral infection Effects 0.000 description 1
Images
Landscapes
- Enzymes And Modification Thereof (AREA)
Abstract
The invention discloses a preparation method and an activity identification method of a porcine second messenger molecule 2 '3' -cGAMP. The invention prepares the soluble expression recombinant porcine 2 '3' -cGAMP synthetase pcGAS, and then provides a simple and rapid preparation method of 2 '3' -cGAMP with low cost, mild condition, high yield and large-scale production; also provided is a method for identifying the biological activity of the recombinant pcGAS enzyme and the catalytic product 2 '3' -cGAMP thereof. The 2 '3' -cGAMP prepared by the invention is an effective activator of the central knob molecule STING of a natural immune signal pathway, and can quickly activate and enhance the natural immune response of a host; the 2 '3' -cGAMP has wide biological activities of antivirus, antitumor, immune enhancement, immune adjuvant and the like, and has huge development prospects of medicines, immune enhancers and immune adjuvants.
Description
Technical Field
The invention relates to the technical field of biochemistry and genetic engineering, in particular to a preparation method and an activity identification method of a porcine second messenger molecule 2 '3' -cGAMP.
Background
2 '3' -cGAMP is used as a second messenger molecule, can combine and activate interferon gene stimulating molecules (STING), promotes the phosphorylation of IRF3/7 and NF-kB and enters the nucleus by activating TBK1 and IKKs, induces the expression of I type IFNs and inflammatory factors, and finally causes the natural immune response of a host. It was found that 2 '3' -cGAMP induces the expression of type I IFNs and inflammatory factors not only through the recognition of STING but also through the rapid transfer of gap junctions (gap junctions) from producer cells to neighboring cells, rapidly activating the STING of neighboring cells to induce the secretion of type I IFN. In addition, 2 '3' -cGAMP can be packaged by enveloped viruses into new virions, and can rapidly elicit STING-mediated antiviral innate immune responses when new virions infect new cells. Therefore, the 2 '3' -cGAMP can be applied and developed as a potential antiviral drug, an antitumor drug, an immunopotentiator and an immunologic adjuvant.
2 '3' -cGAMP is used as an antiviral drug, induces the expression of I type IFNs through a STING-TBK1-IRF3 signal channel, activates the antiviral natural immune response of the organism, and then resists the local and systemic infection of viruses. In addition, 2 '3' -cGAMP not only induces the apoptosis, nuclear lysis and tumor tissue damage of tumor cells through STING, but also induces the release of I-type IFNs and inflammatory factors through STING-TBK1-IRF3 signaling pathway, thereby mediating the maturation of DCs and CD8+Activation of T cells, and improvement of the activity of the anti-tumor drug 5-FU; when 2 '3' -cGAMP is used in combination with PD-1 antibody, PD-L1 antibody or CTLA4 antibody, DCs maturation and CD8 can be induced more effectively+Activation of T cells, increased lethality of the immune system to tumor cells, and increased sensitivity of tumor cells to chemotherapy, and treatment of a wider range of tumor-type diseases. Further research shows that the 2 '3' -cGAMP can also be used as an immunopotentiator to stimulate the body to produce high-level IgG1, IgG2a, IgG2c, IgA and cytokines IFN-gamma and IL-2, and activate DCs, CD4+T cell, CD8+T cells, thereby effectively preventing and treating viral infection. Because of the huge potential application value of the 2 '3' -cGAMP in the aspect of host anti-infection natural immune response and the huge market application prospect of the 2 '3' -cGAMP as a novel antiviral and antitumor drug and an immunopotentiator,there is great interest in the synthesis of 2 '3' -cGAMP. In terms of the prior art, the currently obtained 2 '3' -cGAMP is mainly prepared by tissue purification and chemical synthesis methods, but the two methods have low efficiency, difficult separation and purification, complex process and high cost, and limit the wide application of the two methods. Therefore, it is important for practical applications to find a method for preparing 2 '3' -cGAMP in a simple and rapid manner, at low cost, under mild conditions, in high yield, and in a large scale.
Disclosure of Invention
The invention aims to provide a preparation method and an activity identification method of a porcine second messenger molecule 2 '3' -cGAMP.
The invention provides a method for preparing a porcine second messenger molecule 2 '3' -cGAMP, which comprises the following steps: ATP and GTP are taken as substrates, and pig-derived cyclic guanosine monophosphate-adenylate synthetase is taken as a catalyst to catalyze and synthesize pig-derived second messenger molecule 2 '3' -cGAMP through in vitro enzymatic reaction under the stimulation of DNA.
The preparation method of the pig-derived cyclic guanosine monophosphate-adenylate synthetase comprises the following steps: introducing a gene for coding the pig-derived cyclic guanosine monophosphate-adenylate synthetase into a host bacterium to obtain a recombinant bacterium; culturing the recombinant strain to obtain the porcine cyclic guanosine monophosphate-adenylate synthetase from the recombinant strain.
The gene for encoding the pig-derived cyclic guanosine monophosphate-adenylate synthetase is any one of the following genes (b1) - (b 4):
(b1) DNA molecule shown by 406-position 1488 nucleotide from 5' end of sequence 2 in the sequence table;
(b2) DNA molecule shown in sequence 2 in the sequence table;
(b3) a DNA molecule which hybridizes under stringent conditions to the DNA sequence defined in (b1) or (b2) and which encodes a cyclic guanosine-adenylate synthetase of porcine origin;
(b4) a DNA molecule which has more than 90% homology with the DNA sequence defined in (b1) or (b2) or (b3) and encodes a cyclic guanosine-adenosine synthetase derived from swine.
The stringent conditions can be hybridization and membrane washing with a solution of 0.1 XSSPE (or 0.1 XSSC), 0.1% SDS at 65 ℃ in DNA or RNA hybridization experiments.
The host bacterium may specifically be E.coli, more specifically E.coli Rosetta (DE 3).
The gene for coding the pig-derived cyclic guanosine monophosphate-adenylic acid synthetase can be specifically introduced into a host bacterium through an expression vector containing the gene. The expression vector can be obtained by replacing the fragment between the BamH I site and the Hind III site of the prokaryotic expression vector pET-28a-SUMO with a DNA sequence shown in the 406-1488 site from the 5' end of the sequence 2 in the sequence table.
The method for obtaining the porcine cyclic guanylic acid-adenylic acid synthetase from the recombinant bacteria specifically comprises the following steps (1) and (2):
(1) crushing the recombinant bacteria to obtain a crude extract containing the porcine cyclic guanylic acid-adenylic acid synthetase;
(2) separating the pig-derived cyclic guanosine monophosphate-adenylate synthetase in the crude extract and removing the HIS6-SUMO fusion tag to obtain the pig-derived cyclic guanosine monophosphate-adenylate synthetase.
The separation of the cyclic guanosine monophosphate-adenylate synthetase in the crude extract can be carried out by purifying the cyclic guanosine monophosphate-adenylate synthetase in the crude extract by adopting an affinity chromatography method, and specifically can be carried out by adopting a Ni-NTA agarose gel affinity column.
The reaction system of the enzymatic reaction comprises the following components: pig-derived cyclic guanosine monophosphate-adenylate synthetase, HEPES and MgCl2ATP, GTP and HT-DNA.
The concentration of each component in the reaction system is as follows: pig-derived cyclic guanosine monophosphate-adenylate synthetase 5 mu M, HEPES 20mM and MgCl25mM, ATP 2mM, GTP 2mM and HT-DNA0.1mg/mL.
The reaction conditions of the enzymatic reaction are as follows: incubate at 37 ℃ for 2 h.
After the enzymatic reaction is finished, adding nuclease into the reaction system, and incubating for 30min at 37 ℃; then incubating the reaction product at 95 ℃ for 10min, centrifuging and taking supernatant; the supernatant was filtered through 0.5ml/10kDa ultrafiltration centrifuge tubes to give 2 '3' -cGAMP.
The invention also protects the application of the ring-derived guanylic acid-adenylate synthetase or the coding gene thereof in the preparation of 2 '3' -cGAMP; the pig-derived cyclic guanosine monophosphate-adenylate synthetase is any one of the following (a1) - (a 4):
(a1) the amino acid sequence shown in the No. 136-495 bit of the sequence 1 in the sequence table from the N terminal;
(a2) an amino acid sequence shown in a sequence 1 in a sequence table;
(a3) amino acid sequences with the same function obtained by substituting and/or deleting and/or adding one or more amino acid residues in (a1) or (a 2);
(a4) and (b) an amino acid sequence which has 75% or more homology with (a1) or (a2) and has the same function.
The encoding gene of the pig-derived cyclic guanosine monophosphate-adenylate synthetase is any one of the following genes (b1) - (b 4):
(b1) DNA molecule shown by 406-position 1488 nucleotide from 5' end of sequence 2 in the sequence table;
(b2) DNA molecule shown in sequence 2 in the sequence table;
(b3) a DNA molecule which hybridizes under stringent conditions to the DNA sequence defined in (b1) or (b2) and which encodes a cyclic guanosine-adenylate synthetase of porcine origin;
(b4) a DNA molecule which has more than 90% homology with the DNA sequence defined in (b1) or (b2) or (b3) and encodes a cyclic guanosine-adenosine synthetase derived from swine.
The stringent conditions can be hybridization and membrane washing with a solution of 0.1 XSSPE (or 0.1 XSSC), 0.1% SDS at 65 ℃ in DNA or RNA hybridization experiments.
The invention also provides a kit for preparing the porcine second messenger molecule 2 '3' -cGAMP, which comprises porcine cyclic guanylic acid-adenylic acid synthetase, HEPES, MgCl2ATP, GTP and HT-DNA.
The invention also provides a method for detecting the activity of the pig-derived cyclic guanosine monophosphate-adenylate synthetase and/or the activity of the product 2 '3' -cGAMP thereof, which comprises the following steps:
(1) using ATP and GTP as substrates, and adopting pig-derived cyclic guanosine monophosphate-adenylate synthetase as a catalyst to carry out enzymatic reaction under the stimulation of DNA to obtain a reaction product 2 '3' -cGAMP;
(2) co-culturing the reaction product 2 '3' -cGAMP in the step (1) and a report cell, and determining the activity of the pig-derived cyclic guanosine monophosphate-adenylate synthetase and the activity of the product 2 '3' -cGAMP by detecting the fluorescence intensity of a culture system;
the reporter cell reflects the activation condition of the interferon regulatory factor after 2 '3' -cGAMP stimulation through luciferase activity in the cell, thereby achieving the purpose of detecting the activity of the porcine cyclic guanosine monophosphate-adenosine synthetase and/or the product 2 '3' -cGAMP thereof.
In step (1) of the method, the reaction system of the enzymatic reaction comprises the following components: pig-derived cyclic guanosine monophosphate-adenylate synthetase, HEPES and MgCl2ATP, GTP and HT-DNA. The concentration of each component in the reaction system is as follows: pig-derived cyclic guanosine monophosphate-adenylate synthetase 5 mu M, HEPES 20mM and MgCl25mM, ATP 2mM, GTP 2mM and HT-DNA0.1mg/mL. The reaction conditions of the enzymatic reaction are as follows: incubate at 37 ℃ for 2 h. After the enzymatic reaction is finished, adding nuclease into the reaction system, and incubating for 30min at 37 ℃; then incubating the reaction product at 95 ℃ for 10min, centrifuging and taking supernatant; the supernatant was filtered through a 0.5ml/10kDa ultrafiltration tube to give the reaction product 2 '3' -cGAMP.
In step (2) of the method, the reporter cells were first washed once with permeabilized cell culture fluid before co-culturing with the reaction product 2 '3' -cGAMP. The formulation of the permeabilized cell culture medium is shown in table 2.
The step (2) of the method is specifically as follows: after the reporter cell is washed once by using a permeabilized cell culture solution, 1 volume part of the reaction product 2 '3' -cGAMP and 1 volume part of 2 multiplied by cell membrane penetrating agent are mixed and then co-cultured with the reporter cell (co-culture at 37 ℃ for 30min), and after the reporter cell is washed once by using the permeabilized cell culture solution, the permeabilized cell culture solution is added and cultured for 20h at 37 ℃. Then, the culture system supernatant was taken, added with a luciferase substrate, and the fluorescence value was detected at a wavelength of 700nm, and the relative fluorescence intensity (fold) was calculated, thereby determining the activity of the porcine-derived cyclic guanosine-adenylate synthetase and/or the product thereof, 2 '3' -cGAMP. The formulation of the 2 × cell membrane permeabilizing agent is shown in table 3.
The invention also provides a kit for detecting the activity of the pig-derived cyclic guanosine monophosphate-adenylate synthetase and/or the activity of the product 2 '3' -cGAMP, which comprises ATP, GTP, HT-DNA and a report cell; the reporter cell reflects the activation condition of the interferon regulatory factor after 2 '3' -cGAMP stimulation through luciferase activity in the cell, thereby achieving the purpose of detecting the activity of the cyclic pig guanylate-adenylate synthetase and/or the product 2 '3' -cGAMP thereof.
The kit also comprises the permeabilized cell culture solution, a2 x cell permeabilized agent and a luciferase substrate (specifically, a product with the product number of rep-qlc1 from French invivogen).
Any one of the above reporter cells may specifically be RAW264.7-LuciaTMISG cells (product of Rawl-ISG, Invivogen, France).
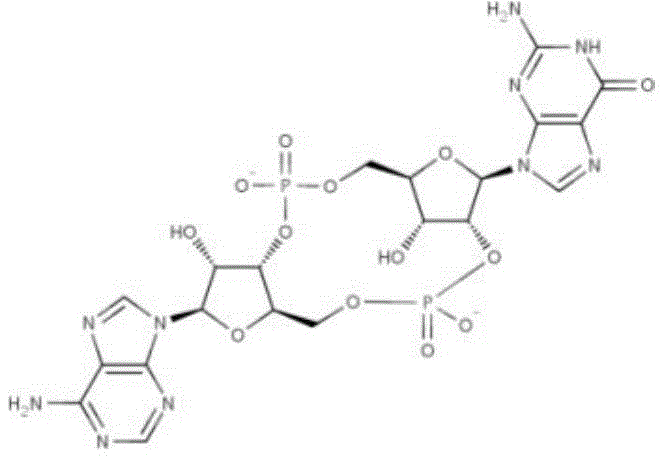
The molecular formula of any one of the porcine second messenger molecules 2 '3' -cGAMP is C20H22N10O13P2CAS number 1441190-66-4, structural formula as follows:
compared with the existing preparation method of 2 '3' -cGAMP, the main technical advantages of the invention are as follows: (1) the method has the characteristics of simple and quick operation, low cost, mild condition, high yield, large-scale production and the like; (2) the recombinant pcGAS enzyme and the catalytic product 2 '3' -cGAMP have the characteristics of high activity and specificity; (3) the method can also identify the biological activity of the recombinant pcGAS enzyme and the 2 '3' -cGAMP catalytic synthesis product thereof, and is convenient for subsequent application.
The invention utilizes the genetic engineering technology to firstly prepare the soluble expressed recombinant porcine 2 '3' -cGAMP synthetase pcGAS, then utilizes the biochemical method to take ATP and GTP as substrates, and catalyzes and synthesizes the 2 '3' -cGAMP with high activity and specificity under the stimulation of DNA by the biological enzyme, thereby providing the simple and rapid preparation method of the 2 '3' -cGAMP with low cost, mild condition, high yield and large-scale production; also provided is a method for identifying the biological activity of the recombinant pcGAS enzyme and the catalytic product 2 '3' -cGAMP thereof. The 2 '3' -cGAMP prepared by the invention is an effective activator of the central knob molecule STING of a natural immune signal pathway, and can quickly activate and enhance the natural immune response of a host; the 2 '3' -cGAMP has wide biological activities of antivirus, antitumor, immune enhancement, immune adjuvant and the like, and has huge development prospects of medicines, immune enhancers and immune adjuvants.
Drawings
FIG. 1 is an SDS-PAGE electrophoresis of purified recombinant pcGAS enzyme.
FIG. 2 is an SDS-PAGE of the recombinant pcGAS enzyme purified after removal of the His6-SUMO fusion tag protein.
FIG. 3 is an assay of the activity of recombinant pcGAS enzyme and the enzymatic reaction product 2 '3' -cGAMP.
Detailed Description
The following examples are given to facilitate a better understanding of the invention, but do not limit the invention. The experimental procedures in the following examples are conventional unless otherwise specified. The test materials used in the following examples were purchased from conventional biochemicals, unless otherwise specified. The quantitative tests in the following examples, all set up three replicates and the results averaged.
2 '3' -cGAMP: the molecular formula is as follows: c20H22N10O13P2CAS number: 1441190-66-4, the structural formula is as follows:
coli Rosetta (DE3) competent cells: beijing Huayuyo Biotech Co., Ltd., Cat No.: SJ 00010.
Protease inhibitors: roche, cat number: 04693132001, respectively; 1 tablet of protease inhibitor was dissolved in 50mL of the solution.
Lysozyme: amresco, cat No.: 0663-5G, working concentration 10 mg/ml.
Ni-NTA agarose gel affinity column: wechbo hui chromatography technologies ltd, cat #: CS-A01-02.
PierceTMCoomassie (bradford) protein quantification kit: sammer Feishale science and technology, China Co., Ltd., Cat No.: 23200.
HT-DNA: sigma aldrich shanghai trade ltd, cat #: D6898.
nuclease Benzonase: sigma aldrich shanghai trade ltd, cat #: e1014-5 KU.
RAW264.7-LuciaTMISG cells: french invivogen, cat No.: rawl-isg.
Luciferase substrate: french invivogen, cat No.: rep-qlc 1.
Example 1 preparation of recombinant porcine 2 '3' -cGAMP synthetase pcGAS
The amino acid sequence of pcGAS enzyme (pig-derived cyclic guanosine monophosphate-adenylate synthetase) is shown as sequence 1 in the sequence table, wherein the 495 th site from the N end 136-495 th site is the catalytic domain of the enzyme activity. The coding gene of the pcGAS enzyme is shown as a sequence 2 in the sequence table, wherein the 406-th 1488 site of the 5' end codes the catalytic domain of the enzyme activity.
1. The fragment between the BamH I and Hind III sites of the prokaryotic expression vector pET-28a-SUMO (Wuhan vast Ling Biotech Co., Ltd., product number: P0028) is replaced by a DNA molecule shown in the 406 th 1488 th site from the 5' end of the sequence 2 of the sequence table to obtain the recombinant expression vector pET-28a-SUMO-pcGAS (sequencing verification has been carried out).
2. And (2) introducing the recombinant expression vector pET-28a-SUMO-pcGAS obtained in the step (1) into escherichia coli Rosetta (DE3) competent cells to obtain recombinant bacteria.
3. Inoculating the recombinant bacteria obtained in the step 2 into an LB liquid culture medium, and culturing at 37 ℃ and 180rpm until the OD of the bacterial liquid600nmWhen the concentration is 0.6-0.8, adding IPTG (concentration of IPTG in the culture system is 0.5mmol/L), inducing at 18 deg.C and 120rpm for 16h, centrifuging at 4 deg.C and 6000r/min for 5min after induction, and collecting thallus precipitate.
4. The cell pellet obtained in step 3 was washed 3 times with 50mL of 20mM Tris-Cl buffer (pH 7.5) containing protease inhibitor, then resuspended with 4mL of Tris-Cl containing lysozyme and protease inhibitor, freeze-thaw repeated 3 times, and disrupted by ultrasonic cell disruption apparatus (ultrasonic setting: 5s, 5s pause, 20min total).
5. After completion of step 4, the mixture was centrifuged at 12000rpm/min at 4 ℃ for 30min, and the supernatant was collected.
6. The supernatant obtained in step 5 was filtered through a 0.45 μm microfiltration membrane and then preliminarily separated and purified by a Ni-NTA agarose gel affinity column (according to the instructions) to obtain pcGAS, which was identified by SDS-PAGE analysis as purified pcGAS (purity: 90%), and the results are shown in FIG. 1.
7. Taking the pcGAS obtained in the step 6, carrying out enzyme digestion by SUMO protease ULP (Beijing Sorley technologies, Ltd.) to remove HIS6-SUMO fusion tag protein, and then separating and purifying by a Ni-NTA agarose gel affinity column (operating according to the instruction) to obtain the recombinant pcGAS enzyme. The recombinant pcGAS enzyme (95% pure) was identified by SDS-PAGE analysis and the results are shown in FIG. 2.
An enzyme digestion reaction system: recombinant pcGAS enzyme 1000. mu.g, SUMO Protease Buffer 20. mu.L, SUMO Protease 2. mu.L, ddH2O is metered to 1000. mu.L. And (3) enzyme digestion reaction program: the digestion was carried out at 4 ℃ for 12h or overnight.
8. Use of PierceTMThe Coomassie (Bradford) protein quantification kit (according to the instructions) determined a concentration of 10.75mg/ml of recombinant pcGAS enzyme.
Example 2 enzymatic Synthesis of porcine 2 '3' -cGAMP Using recombinant pcGAS
The porcine recombinant pcGAS enzyme prepared in the example 1 is used for catalyzing and synthesizing the porcine 2 '3' -cGAMP in vitro efficiently and specifically as follows:
1. an enzymatic reaction system was configured (as shown in table 1). The enzymatic reaction system was set up on ice.
TABLE 1 enzymatic reaction System
2. After completion of step 1, incubation was carried out at 37 ℃ for 2h, followed by addition of 50. mu.L nuclease Benzonase and incubation at 37 ℃ for 30 min.
3. After step 2 was completed, the reaction product was incubated at 95 ℃ for 10min, and then centrifuged at 16000g/min for 10min to obtain a supernatant. The supernatant was filtered through a 0.5ml/10kDa ultrafiltration centrifuge tube (Merck-Millipore, cat # UFC501096) to obtain a reaction product.
The absorbance values of the reaction product obtained in step 3 and the enzymatic reaction system of step 1 were measured at 260nm using a spectrophotometer, and the reaction yield was 97.83%.
Reaction yield-reaction product OD260nmEnzymatic reaction System OD260nm。
Example 3 Activity characterization of porcine pcGAS and catalytic product 2 '3' -cGAMP
RAW264.7-LuciaTMISG cells are reporter cells in which the ISG54 promoter and ISRE, an interferon-stimulated response element, co-drive the expression of a luciferase gene (Lucia luciferase gene). When 2 '3' -cGAMP is stimulated, RAW264.7-LuciaTMThe transcription factors IRF3 and IRF7 are activated after recognition by STING molecule expressed by ISG cells, then combined with ISRE to induce the secretion of luciferase gene, and further processed by QUANTI-LucTMLuciferase fluorescence was measured to reflect pcGAS enzymatic activity and 2 '3' -cGAMP stimulatory activity.
1. Mixing RAW264.7-LuciaTMISG cells were adjusted to a density of 5X 105Perml, seeded in 96-well plates (100. mu.L per well, cell number 5X 104/mL) at 37 ℃ until the cell density reaches 70-80%.
2. After completion of step 1, the 96-well plate was removed, the culture medium was discarded, and cells were washed once with 100 μ L of permeabilized cell culture medium. The formulation of the permeabilized cell culture medium is shown in Table 2.
TABLE 2 formula of permeabilized cell culture solution
3. After completion of step 2, the following six groups were set up for operation (6 replicates per group):
pcGAS (10. mu.M) group: mixing the enzymatic reaction product with 2 Xcell cell membrane permeable agent according to the volume ratio of 1:1, adding into the hole (50. mu.L per hole), incubating at 37 ℃ for 30min, removing the liquid in the hole, adding 100. mu.L of the permeable cell culture solution shown in Table 2, carefully cleaning the cells once, adding 50. mu.L of the permeable cell culture solution, and culturing at 37 ℃ for 20 h. The enzymatic reaction product was the reaction product obtained in example 2 (wherein the concentration of the recombinant pcGAS enzyme in the reaction system was changed to 10. mu.M).
pcGAS (5. mu.M) group: mixing the enzymatic reaction product with 2 Xcell cell membrane permeable agent according to the volume ratio of 1:1, adding into the hole (50. mu.L per hole), incubating at 37 ℃ for 30min, removing the liquid in the hole, adding 100. mu.L of the permeable cell culture solution shown in Table 2, carefully cleaning the cells once, adding 50. mu.L of the permeable cell culture solution, and culturing at 37 ℃ for 20 h. The enzymatic reaction product is the reaction product obtained in example 2.
Group 2 '3' -cGAMP (50 ng/ml): 100ng/ml of 2 '3' -cGAMP (French invivogen, cat # tlrl-nacga23) and 2 Xcell permeant were mixed at a volume ratio of 1:1 and added to wells (50. mu.L/well, working concentration 50ng/ml), and after incubation at 37 ℃ for 30min, the liquid in the wells was discarded, 100. mu.L of the permeabilized cell culture shown in Table 2 was added and the cells were washed once carefully, and 50. mu.L of the permeabilized cell culture was added and cultured at 37 ℃ for 20 hours.
ATP and GTP free group: mixing the enzymatic reaction product with 2 Xcell cell membrane permeable agent according to the volume ratio of 1:1, adding into the hole (50. mu.L per hole), incubating at 37 ℃ for 30min, removing the liquid in the hole, adding 100. mu.L of the permeable cell culture solution shown in Table 2, carefully cleaning the cells once, adding 50. mu.L of the permeable cell culture solution, and culturing at 37 ℃ for 20 h. The enzymatic reaction product was the reaction product obtained in example 2 (wherein ATP and GTP were not added to the reaction system).
pcGAS-free group: mixing the enzymatic reaction product with 2 Xcell cell membrane permeable agent according to the volume ratio of 1:1, adding into the hole (50. mu.L per hole), incubating at 37 ℃ for 30min, removing the liquid in the hole, adding 100. mu.L of the permeable cell culture solution shown in Table 2, carefully cleaning the cells once, adding 50. mu.L of the permeable cell culture solution, and culturing at 37 ℃ for 20 h. The enzymatic reaction product was the reaction product obtained in example 2 (wherein no recombinant pcGAS enzyme was added to the reaction system).
Control group: 2 Xcell membrane permeabilizing agent is added into the holes (50. mu.L of each hole), the cells are incubated at 37 ℃ for 30min, then the liquid in the holes is discarded, 100. mu.L of permeabilized cell culture solution shown in Table 2 is added, the cells are carefully cleaned once, 50. mu.L of permeabilized cell culture solution is added, and the cells are cultured at 37 ℃ for 20 h.
The formulation of the 2 × cell membrane permeabilizing agent is shown in table 3.
TABLE 32X cell permeant formulations
| Components | Concentration in the reaction System | Volume of |
| HEPES(pH7.5) | 100mM | 500μL |
| MgCl2 | 6mM | 30μL |
| KCl | 200mM | 1ml |
| DTT | 0.2mM | 10μL |
| Sucrose | 170mM | 850μL |
| BSA | 7.5% | 267μL |
| ATP | 2mM | 100μL |
| GTP | 0.2mM | 10μL |
| Digitonin (Digitonin) | 20μg/mL | 20μL |
| DEPCH2O | Make up to 5ml |
4. After completion of step 3, 25. mu.L of the supernatant was pipetted into a 96-well microwell plate and 50. mu.L of luciferase substrate was added, and the fluorescence value was immediately detected at a wavelength of 700nm using a VICTORNivo multimode plate reader (Perkin Elmer Co., Ltd.) and the relative fluorescence intensity (fold) was calculated, thereby determining the biological activity of the recombinant pcGAS enzyme and its catalytic product, 2 '3' -cGAMP.
The results are shown in FIG. 3. The results show that the recombinant pcGAS enzyme and the catalytic product 2 '3' -cGAMP thereof have high biological activity, the expression of luciferase reporter gene can be strongly induced through the STING-TBK1-IRF3/7 signal pathway, and the stimulation activity of the 2 '3' -cGAMP is related to the concentration of pcGAS.
Sequence listing
<110> Lanzhou veterinary research institute of Chinese academy of agricultural sciences
<120> preparation and activity identification method of pig-derived second messenger molecule 2 '3' -cGAMP
<160> 2
<170> SIPOSequenceListing 1.0
<210> 1
<211> 495
<212> PRT
<213> pig (Sus scrofa)
<400> 1
Met Ala Ala Arg Arg Gly Lys Ser Thr Arg Thr Ala Ser Glu Val Gly
1 5 10 15
Ala Ala Gly Pro Arg Ala Ser Ala Arg Ser Val Asn Gly Ala Pro Thr
20 25 30
Val Pro Glu Ala Ala Arg Pro Gly Ala Arg Arg Asn Gly Pro Ser Arg
35 40 45
Ala Ser Gly Cys Arg Arg Glu Lys Ser Gly Pro Asp Pro Arg Glu Lys
50 55 60
Pro Gln Val Arg Thr Arg Thr Ala Arg Ala Glu Asp Gln Ala Glu Gly
65 70 75 80
Pro Ser Ala Pro Ser Glu Arg Val Glu Pro Pro Ser Ala Gln Gly Ala
85 90 95
Ser Leu Leu Arg Ala Gly Ser Cys Arg Ala Arg Glu Ala Arg Ser Ala
100 105 110
Arg Glu Leu Arg Pro Gln Ala Gly Ala Thr Glu Leu Ala Ala Pro Ala
115 120 125
Arg Met Glu Ala Pro Pro Gly Ala Trp Lys Leu Gln Thr Val Leu Glu
130 135 140
Lys Val Arg Leu Ser Arg His Glu Ile Ser Glu Ala Ala Glu Val Val
145 150 155 160
Asn Trp Val Val Glu His Leu Leu Arg Arg Leu Gln Gly Gly Glu Ser
165 170 175
Glu Phe Lys Gly Val Ala Leu Leu Arg Thr Gly Ser Tyr Tyr Glu Arg
180 185 190
Val Lys Ile Ser Ala Pro Asn Glu Phe Asp Val Met Phe Lys Leu Glu
195 200 205
Val Pro Arg Ile Gln Leu Glu Glu Tyr Cys Asn Ser Gly Ala His Tyr
210 215 220
Phe Val Lys Phe Lys Arg Asn Pro Gly Gly Asn Pro Leu Glu Gln Phe
225 230 235 240
Leu Glu Lys Glu Ile Leu Ser Ala Ser Lys Met Leu Ser Lys Phe Arg
245 250 255
Lys Ile Ile Lys Glu Glu Ile Lys Asn Ile Glu Gly Val Thr Val Glu
260 265 270
Arg Lys Arg Arg Gly Ser Pro Ala Val Thr Leu Leu Ile Ser Lys Pro
275 280 285
Lys Glu Ile Ser Val Asp Ile Ile Leu Ala Leu Glu Ser Lys Ser Ser
290 295 300
Trp Pro Ala Ser Thr Gln Lys Gly Leu Pro Ile Ser Gln Trp Leu Gly
305 310 315 320
Ala Lys Val Lys Asn Asn Leu Lys Arg Gln Pro Phe Tyr Leu Val Pro
325 330 335
Lys His Ala Lys Glu Gly Ser Gly Phe Gln Glu Glu Thr Trp Arg Leu
340 345 350
Ser Phe Ser His Ile Glu Lys Asp Ile Leu Lys Asn His Gly Gln Ser
355 360 365
Lys Thr Cys Cys Glu Ile Asp Gly Val Lys Cys Cys Arg Lys Glu Cys
370 375 380
Leu Lys Leu Met Lys Tyr Leu Leu Glu Gln Leu Lys Lys Lys Phe Gly
385 390 395 400
Asn Arg Arg Glu Leu Ala Lys Phe Cys Ser Tyr His Val Lys Thr Ala
405 410 415
Phe Phe His Val Cys Thr Gln Asp Pro His Asp Asn Gln Trp His Leu
420 425 430
Lys Asn Leu Glu Cys Cys Phe Asp Asn Cys Val Ala Tyr Phe Leu Gln
435 440 445
Cys Leu Lys Thr Glu Gln Leu Ala Asn Tyr Phe Ile Pro Gly Val Asn
450 455 460
Leu Phe Ser Arg Asp Leu Ile Asp Lys Pro Ser Lys Glu Phe Leu Ser
465 470 475 480
Lys Gln Ile Glu Tyr Glu Arg Asn Asn Gly Phe Pro Val Phe Trp
485 490 495
<210> 2
<211> 1488
<212> DNA
<213> pig (Sus scrofa)
<400> 2
atggcggccc ggcggggaaa gtccacgcgg acagcttcag aggtgggagc cgctggcccc 60
agggcctcag cgcggagtgt aaatggcgct ccaacggtgc ccgaggccgc ccggcctggc 120
gcacggagga acggcccctc cagggcgtcg ggatgccgac gggagaagag cggcccggac 180
ccccgggaga agccgcaggt acgcacgagg acggcccgcg ccgaggacca ggcggagggc 240
ccatcggctc cctccgaaag ggtagagcct ccttcggccc agggggcttc ccttctcagg 300
gctggatctt gccgcgcgag agaggcgcgc tcggcgcggg aactgagacc ccaggccggg 360
gccacagagc tcgcagcccc cgcgcggatg gaggcacccc ccggcgcctg gaagctccag 420
acggtgctgg agaaggtgag gctgagccgc cacgaaatct cagaagcggc ggaggtggtg 480
aactgggtcg tggagcacct gctgcggagg ctgcagggcg gagagtccga gttcaaaggc 540
gttgccctgc tgcgcaccgg gagctactat gagcgagtga agatttctgc tcccaatgaa 600
tttgatgtta tgttcaaact ggaagttccc cgaattcagc tagaggaata ttgcaacagt 660
ggtgctcatt attttgtcaa gttcaaaaga aatcctggag gaaatcctct ggaacagttc 720
ttagaaaagg aaatattatc agcttctaag atgctgtcca agtttaggaa aattattaag 780
gaagaaatta aaaatataga aggtgtcact gtggagagga agaggagagg gagccctgca 840
gtaacacttc tgattagcaa acctaaagaa atatctgtgg atataatcct ggctttggaa 900
tctaaaagca gctggcctgc tagcacccag aaaggcctgc ccatcagtca gtggcttgga 960
gcaaaagtta aaaacaatct gaaacgacag ccattttacc tggtacccaa gcatgccaag 1020
gaaggaagtg gttttcaaga agaaacatgg aggctttcct tctctcacat tgaaaaggac 1080
attttgaaaa atcatggaca gtctaaaaca tgctgtgaaa ttgatggagt gaaatgttgc 1140
aggaaagagt gtttaaaact aatgaaatac cttttagaac aactgaaaaa aaagtttgga 1200
aaccgaaggg aactggctaa gttctgttct tatcacgtga aaactgcctt ctttcacgtc 1260
tgtacccagg accctcatga caatcagtgg cacctcaaga accttgagtg ctgctttgat 1320
aactgtgtgg catattttct gcagtgcctc aagacagaac aactggccaa ttatttcatt 1380
cctggagtca atctgttctc tcgagaccta attgacaaac caagtaaaga atttctgtca 1440
aagcaaattg aatatgaaag aaacaatgga tttccagttt tttggtga 1488
Claims (1)
1. A method for detecting the activity of pig cyclic guanosine monophosphate synthetase and/or the activity of 2 '3' -cGAMP of a product thereof, which comprises the following steps:
(1) using ATP and GTP as substrates, and adopting pig-derived cyclic guanosine monophosphate-adenylate synthetase as a catalyst to carry out enzymatic reaction under the stimulation of DNA to obtain a reaction product 2 '3' -cGAMP;
(2) co-culturing the reaction product 2 '3' -cGAMP in the step (1) and a report cell, and determining the activity of the pig-derived cyclic guanosine monophosphate-adenylate synthetase and the activity of the product 2 '3' -cGAMP by detecting the fluorescence intensity of a culture system;
the reporter cell reflects the activation condition of the interferon regulatory factor after 2 '3' -cGAMP stimulation through luciferase activity in the cell, thereby achieving the purpose of detecting the activity of the porcine cyclic guanosine monophosphate-adenosine synthetase and/or the product 2 '3' -cGAMP thereof;
the step (2) is as follows: before the report cell is co-cultured with the reaction product 2 '3' -cGAMP, firstly, washing the report cell once by using a permeabilized cell culture solution, mixing 1 volume part of the reaction product 2 '3' -cGAMP with 1 volume part of 2 multiplied cell membrane penetrating agent, then, co-culturing the mixture with the report cell at 37 ℃ for 30min, washing the report cell once by using the permeabilized cell culture solution, then, adding the permeabilized cell culture solution, and culturing the mixture for 20h at 37 ℃; then taking culture system supernatant, adding luciferase substrate, detecting fluorescence value at 700nm wavelength, and calculating relative fluorescence intensity, thereby determining activity of pig-derived cyclic guanosine-adenylate synthetase and/or product 2 '3' -cGAMP thereof;
the permeabilized cell culture solution consists of FBS, HEPES, L-glutamine, Zeolin and DMEM, wherein the concentration of FBS is 10%, the pH value of HEPES is 7.5, the concentration is 25 mM, the concentration of L-glutamine is 2mM, and the concentration of Zeolin is 200 mu g/mL;
the 2 × cell membrane-penetrating agent is prepared from HEPES, MgCl2, KCl, DTT, sucrose, BSA, ATP, GTP, Digitonin and DEPC H2O, wherein HEPES has a pH of 7.5 and a concentration of 100 mM, MgCl2Has a concentration of 6 mM, KCl concentration of 200 mM, DTT concentration of 0.2 mM, sucrose concentration of 170 mM, BSA content of 7.5%, ATP concentration of 2mM, GTP concentration of 0.2 mM; the porcine cyclic guanylic acid-adenylic acid synthetase is (a1) or (a 2):
(a1) the amino acid sequence shown in the No. 136-495 bit of the sequence 1 in the sequence table from the N terminal;
(a2) an amino acid sequence shown in a sequence 1 in a sequence table.
Priority Applications (1)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| CN201910307619.XA CN109929894B (en) | 2019-04-17 | 2019-04-17 | A kind of preparation and activity identification method of porcine second messenger molecule 2'3'-cGAMP |
Applications Claiming Priority (1)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| CN201910307619.XA CN109929894B (en) | 2019-04-17 | 2019-04-17 | A kind of preparation and activity identification method of porcine second messenger molecule 2'3'-cGAMP |
Publications (2)
| Publication Number | Publication Date |
|---|---|
| CN109929894A CN109929894A (en) | 2019-06-25 |
| CN109929894B true CN109929894B (en) | 2021-06-01 |
Family
ID=66990164
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| CN201910307619.XA Active CN109929894B (en) | 2019-04-17 | 2019-04-17 | A kind of preparation and activity identification method of porcine second messenger molecule 2'3'-cGAMP |
Country Status (1)
| Country | Link |
|---|---|
| CN (1) | CN109929894B (en) |
Families Citing this family (1)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN114790470B (en) * | 2022-04-14 | 2023-09-29 | 山东大学 | A method for preparing the second messenger molecule 2’3’-cGAMP through solid-phase multi-enzyme coupling |
Family Cites Families (4)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO2016100261A2 (en) * | 2014-12-17 | 2016-06-23 | Lipogen Llc | Method of treating cancer with cgamp or cgasmp |
| CN106190999A (en) * | 2015-05-10 | 2016-12-07 | 聊城市奥润生物医药科技有限公司 | The high efficiency preparation method of ring dinucleotide cGAMP |
| EP3429596B1 (en) * | 2016-03-18 | 2022-08-31 | Immune Sensor, LLC | Cyclic di-nucleotide compounds and methods of use |
| JOP20170188A1 (en) * | 2016-11-25 | 2019-01-30 | Janssen Biotech Inc | Cyclic dinucleotides as sting agonists |
-
2019
- 2019-04-17 CN CN201910307619.XA patent/CN109929894B/en active Active
Non-Patent Citations (4)
| Title |
|---|
| Cyclic GMP-AMP Containing Mixed Phosphodiester Linkages Is An Endogenous High-Affinity Ligand for STING;Xu Zhang等;《Molecular Cell》;20130725;第51卷(第2期);第226-235页 * |
| cyclic GMP-AMP synthase [Sus scrofa];NCBI Reference Sequence: XP_013840602.1;《NCBI》;20170513;第1页 * |
| 猪 cGAS基因的克隆与原核表达;杜丽丽等;《中国农业科学》;20160930;第49卷(第9期);第1803-1809页 * |
| 猪环磷酸鸟苷-腺苷酸合成酶cGAS基因的克隆及其功能鉴定;杜丽丽;《中国优秀硕士学位论文全文数据库 农业科技辑》;中国学术期刊(光盘版)电子杂志社;20160615(第06期);D050-124 * |
Also Published As
| Publication number | Publication date |
|---|---|
| CN109929894A (en) | 2019-06-25 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| CN108330119B (en) | A kind of chitosanase and its application in the preparation of chitosan oligosaccharide | |
| CN107400666B (en) | A kind of aminopeptidase and its coding gene and application | |
| CN113736763B (en) | Myrosinase Rmmr and application thereof in preparation of sulforaphane and sulforaphane | |
| CN109929894B (en) | A kind of preparation and activity identification method of porcine second messenger molecule 2'3'-cGAMP | |
| CN110592049A (en) | An Aspergillus niger ester hydrolase AnCu3, its coding gene and its application in hydrolyzing DEHP | |
| CN120641560A (en) | RNA polymerase variants and their applications | |
| CN104232606B (en) | The β glucuroides and its expressing gene of a kind of transformation and application | |
| CN114107341B (en) | Application of fungal source alpha-L-rhamnosidase in icariin production | |
| WO2014166326A1 (en) | β-GLUCOSIDASE AND EXPRESSION GENE AND USAGE THEREOF | |
| CN109402092B (en) | A marine environment-derived chitinase and its gene | |
| CN112322599B (en) | Transaminase UPTA, preparation method and application | |
| CN110499323A (en) | Method for synthesizing cyclic guanosine diphosphate catalyzed by guanylate cyclase | |
| CN112301010B (en) | A kind of amine oxidase ACAO, preparation method and application | |
| CN113265435A (en) | Preparation method of bacterial second messenger molecule cyclic dinucleotide | |
| CN109022405A (en) | A kind of Cold tolerance algin catenase AlgA5 and its application | |
| CN111471667B (en) | Chitosanase Csn-PT and Its Application | |
| CN109182439A (en) | The bioconversion method of the rare saponin(e Rg3 of ginseng | |
| CN108588049B (en) | A kind of glucosamine synthase, engineering bacteria and application thereof | |
| CN106834328A (en) | A kind of S ribosylhomocysteines lyase gene recombinant expression carrier and its expression and application | |
| CN118147114A (en) | Beta-glucosidase mutant for catalyzing phlorizin to prepare phloretin, coding gene thereof and application of beta-glucosidase mutant | |
| CN111607575B (en) | A kind of transaminase PHTA, preparation method and application | |
| CN111549007B (en) | Transaminase TSTA, preparation method and application | |
| RU2619170C2 (en) | RECOMBINANT PLASMID DNA pER-ARNS3, ENCODING FUSION PROTEIN CAPABLE OF AUTOCATALYTIC SPLITTING WITH FORMATION OF ARNS3, ESCHERICHIA COLI STRAIN C3030/pER-ARNS3 PRODUCER OF SAID PROTEINS AND METHODS FOR RECOMBINANT ARNS3 PRODUCTION | |
| CN109022404A (en) | A kind of novel Cold tolerance algin catenase AlgA7 and its application | |
| CN114380896A (en) | Expression method of novel coronavirus S protein |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| PB01 | Publication | ||
| PB01 | Publication | ||
| SE01 | Entry into force of request for substantive examination | ||
| SE01 | Entry into force of request for substantive examination | ||
| GR01 | Patent grant | ||
| GR01 | Patent grant |