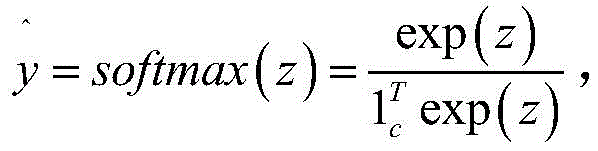CN109722466B - Method for rapidly detecting strain producing carbapenemase by using AI model - Google Patents
Method for rapidly detecting strain producing carbapenemase by using AI model Download PDFInfo
- Publication number
- CN109722466B CN109722466B CN201910090138.8A CN201910090138A CN109722466B CN 109722466 B CN109722466 B CN 109722466B CN 201910090138 A CN201910090138 A CN 201910090138A CN 109722466 B CN109722466 B CN 109722466B
- Authority
- CN
- China
- Prior art keywords
- model
- value
- solution
- training
- lstm
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Active
Links
Images
Landscapes
- Measuring Or Testing Involving Enzymes Or Micro-Organisms (AREA)
Abstract
本发明公开了一种运用AI模型快速检测产生碳青霉烯酶菌株的方法,通过使用溴百里酚蓝溶液,测定混合溶液的OD值,最后运用AI模型通过分析混合溶液OD值的变化趋势判断菌株是否能够产生碳青霉烯酶;本发明通过检测菌株的OD值随时间变化的趋势降低了误差;通过检测菌株溶液的OD值,能够在极短时间内快速判断菌株是否产生碳青霉烯酶,极大地节省了时间;对于弱产酶的菌株,运用AI模型分析菌株溶液OD值的变化,快速判断出来;将OD值导入到LSTM模型中,即可立刻得到菌株是否能够产酶以及产酶能力的强弱,且能直观的看到一批次中菌株产酶阳性数量与产酶阴性数量的多少,并直接导出产酶结果,不必人工记录实验结果,高效便捷。
The invention discloses a method for rapidly detecting carbapenemase-producing strains by using an AI model. By using a bromothymol blue solution, the OD value of a mixed solution is measured, and finally the AI model is used to analyze the changing trend of the OD value of the mixed solution. Judging whether the strain can produce carbapenemase; the invention reduces errors by detecting the trend of the OD value of the strain changing with time; by detecting the OD value of the strain solution, it can quickly judge whether the strain produces carbapenemase in a very short time Enzyme, which greatly saves time; for weak enzyme-producing strains, the AI model is used to analyze the change of the OD value of the strain solution and quickly judge; import the OD value into the LSTM model, and you can immediately get whether the strain can produce enzymes and The enzyme-producing ability is strong and weak, and the number of positive and negative enzyme-producing strains in a batch can be intuitively seen, and the enzyme-producing results can be directly derived, without the need to manually record the experimental results, which is efficient and convenient.
Description
技术领域technical field
本发明涉及人工智能和生物技术的研究领域,特别涉及一种运用AI模型快速检测产生碳青霉烯酶菌株的方法。The invention relates to the research fields of artificial intelligence and biotechnology, in particular to a method for rapidly detecting carbapenemase-producing strains by using an AI model.
背景技术Background technique
碳青霉烯类药物是目前临床上治疗革兰氏阴性菌感染的最后防线之一,尤其是对产碳青霉烯酶肠杆菌(Carbapenemase-producing Enterobacteriaceae,CRE),而且近年来碳青霉烯类药物的耐药率逐年增高。因此,开发一种能够准确快速检测碳青霉烯酶的方法对临床治疗来说至关重要。Carbapenems are currently one of the last lines of defense in the clinical treatment of Gram-negative bacterial infections, especially for Carbapenemase-producing Enterobacteriaceae (CRE). The drug resistance rate is increasing year by year. Therefore, developing a method for accurate and rapid detection of carbapenemase is crucial for clinical treatment.
2012年Nordmann等人报道Carba-NP方法通过使用酚红指示剂在不同下的颜色变化来检测菌株是否产碳青霉烯酶,产碳青霉烯酶的菌株,其β-内酰胺环发生水解时,会产生羧基,最终导致酚红指示剂颜色随pH的降低有红色变为黄色,从而判断出菌株是否产碳青霉烯酶,缺点是:由于酚红从红色变为黄色的颜色变化幅度不够大,对于一些弱产酶的菌株并不能很好的判断期是否产碳青霉烯酶。2013年J.Pires等人报道了Blue-Carba方法,即利用溴百里酚蓝的宽变色范围(pH=6.0-7.6),由蓝色变为黄色,使其更容易判断菌株是否产碳青霉烯,缺点是:对于弱产酶的菌株也不能很好的判断,且也需要2h的菌株生长时间;2017年Jeremy Surre等通过运用紫外分光光度计的方法检测相应溶液在558nm的OD值,通过OD值的变化来判断菌株是否产酶,缺点是:通过运用紫外分光光度计仍需2h才能完全检测菌株是否产碳青霉烯酶;Lisong Shen等人通过MALDI-TOF检测血液培养的产碳青霉烯酶的肠杆菌科菌株,该方法是以麦氏杆菌为底物的亚胺培南水解实验方法,根据亚胺培南及其水解产物的峰值强度计算出logQ值,以区分菌株是否产碳青霉烯酶,该方法能够用于血液样本的快速鉴定和药敏实验报告,但是NDM-1亚型的肠杆菌科菌株有41.7%被判断为阴性,因此其结果敏感性不是很高。In 2012, Nordmann et al. reported that the Carba-NP method detects whether the strain produces carbapenemase by using the color change of the phenol red indicator under different conditions. The carbapenemase-producing strain has hydrolysis of its β-lactam ring. When the pH value decreases, carboxyl groups will be generated, and the color of the phenol red indicator will change from red to yellow with the decrease of pH, so as to determine whether the strain produces carbapenemase. It is not large enough, and for some strains with weak enzyme production, it is not very good to judge whether it produces carbapenemase. In 2013, J.Pires et al. reported the Blue-Carba method, which uses the wide discoloration range of bromothymol blue (pH=6.0-7.6) to change from blue to yellow, making it easier to judge whether the strain produces carbon blue. The disadvantage is that: the strains with weak enzyme production cannot be well judged, and the strain growth time of 2h is also required; in 2017, Jeremy Surre et al. detected the OD value of the corresponding solution at 558nm by using a UV spectrophotometer. The change of OD value is used to judge whether the strain produces enzyme. The disadvantage is that it still takes 2 hours to completely detect whether the strain produces carbapenemase by using UV spectrophotometer; Lisong Shen et al. detected the carbon production of blood culture by MALDI-TOF. Enterobacteriaceae strains of penemase, the method is a hydrolysis test method of imipenem with Mysterella as the substrate, and the logQ value is calculated according to the peak intensity of imipenem and its hydrolyzed products to distinguish whether the strain is not Carbapenemase production, this method can be used for rapid identification and drug susceptibility test report of blood samples, but 41.7% of Enterobacteriaceae strains of NDM-1 subtype were judged negative, so the sensitivity of the results is not very high .
发明内容SUMMARY OF THE INVENTION
本发明的主要目的在于克服现有技术的缺点与不足,提供一种运用AI模型快速检测产生碳青霉烯酶菌株的方法,能自动分析菌株是否产碳青霉烯酶以及产酶阳性菌株的数量,进而快速判断菌株是否产碳青霉烯酶以及菌株产酶能力的强弱,且对一些弱产酶菌株也能快速判断其是否产碳青霉烯酶,并将实验结果直接导出到电脑中,减少人工记录实验结果的时间。解决了人为肉眼观察颜色变化的不敏感性以及产酶时间长的问题。The main purpose of the present invention is to overcome the shortcomings and deficiencies of the prior art, and to provide a method for rapidly detecting carbapenemase-producing strains by using an AI model, which can automatically analyze whether the strains produce carbapenemase and whether the strains produce carbapenemase-positive strains. Quantity, and then quickly determine whether the strain produces carbapenemase and the strength of the strain's enzyme-producing ability, and can also quickly determine whether the strain produces carbapenemase for some weak enzyme-producing strains, and export the experimental results directly to the computer. , reducing the time for manual recording of experimental results. It solves the problems of the insensitivity of human visual observation of color changes and the long time for enzyme production.
本发明的目的通过以下的技术方案实现:The object of the present invention is achieved through the following technical solutions:
一种运用AI模型快速检测产生碳青霉烯酶菌株的方法,包括以下步骤:A method for rapidly detecting carbapenemase-producing strains using an AI model, comprising the following steps:
使用Blue-Carba方法,配制solution溶液;即在溴百里酚蓝溶液中加入亚胺培南,再加入ZnSO4溶液,最后调节溶液PH,得到solution溶液;Use the Blue-Carba method to prepare the solution solution; namely, add imipenem to the bromothymol blue solution, then add ZnSO4 solution, and finally adjust the pH of the solution to obtain the solution solution;
选择野生菌株,用PBS液配制细菌悬浮液:将样品在麦康凯琼脂培养基上分离纯化,之后在LB琼脂培养基上划菌苔,培养一定时间,通过MALDI-TOF鉴定菌株种属;用接种环刮一环菌苔到PBS液中,涡旋混匀,调节溶液OD值,得到细菌悬浮液;Select wild strains and prepare bacterial suspension with PBS solution: isolate and purify the samples on MacConkey agar medium, then streaked on LB agar medium, cultivate for a certain period of time, and identify the strain species by MALDI-TOF; Scrape a circle of bacterial moss into the PBS solution, vortex to mix, and adjust the OD value of the solution to obtain a bacterial suspension;
将solution溶液和细菌悬浮液混匀;用排枪吸取solution溶液加入到96孔板中,再吸取细菌悬浮液与solution溶液混匀;Mix the solution solution and the bacterial suspension; use a discharge gun to draw the solution solution and add it to the 96-well plate, and then draw the bacterial suspension and mix it with the solution solution;
使用全自动酶标仪检测混合溶液的OD值;将混合溶液的96孔板放入全自动酶标仪中,设置温度,检测出混合溶液的OD值,并导出原始数据;Use an automatic microplate reader to detect the OD value of the mixed solution; put the 96-well plate of the mixed solution into the automatic microplate reader, set the temperature, detect the OD value of the mixed solution, and export the original data;
选择AI模型,并进行模型训练和验证,得到最优AI模型,将混合溶液的OD值原始数据导入最优AI模型中,经由LSTM循环神经元循环计算,输出特征向量,再经过一个全连接网络,分析得到菌株产碳青霉烯酶的结果;Select the AI model, and perform model training and verification to obtain the optimal AI model, import the raw data of the OD value of the mixed solution into the optimal AI model, and cyclically calculate through the LSTM cyclic neurons, output the feature vector, and then go through a fully connected network. , and analyze the results of carbapenemase production by the strain;
进一步地,所述进行模型训练和验证,具体过程为:Further, the described model training and verification, the specific process is:
用80%数据作为学习样本,进行模型训练,用20%数据作为验证数据;Use 80% of the data as the learning sample to train the model, and use 20% of the data as the validation data;
所述模型训练采用cross entropy作为损失函数,采用Adam梯度下降方式,利用训练数据,进行多次迭代优化模型参数,使损失函数最小化;The model training adopts cross entropy as the loss function, adopts the Adam gradient descent method, and uses the training data to perform multiple iterations to optimize the model parameters to minimize the loss function;
S1,LSTM的计算过程:S1, the calculation process of LSTM:
LSTM以每个时间点OD值作为输入,时间点个数记为n,LSTM计算时遍历每个时间点,单独作为输入,进行一次循环计算;设定LSTM的神经元个数为k;设t为当前时间点;LSTM takes the OD value of each time point as input, and the number of time points is denoted as n. When LSTM calculates, it traverses each time point and uses it as input to perform a loop calculation; set the number of neurons of LSTM to k; set t is the current point in time;
LSTM有两个传输状态,一个Ct、一个ht;LSTM的当前输入xt和上一个状态传递下来的ht-1拼接计算得到四个状态:LSTM has two transmission states, one C t and one h t ; the current input x t of LSTM and the h t-1 passed from the previous state are concatenated to obtain four states:
ft=σ(Wf*[ht-1,xt]+bf),f t =σ(W f *[h t-1 ,x t ]+b f ),
it=σ(Wi*[ht-1,xt]+bi),i t =σ(W i *[h t-1 ,x t ]+ bi ),
ot=σ(Wo*[ht-1,xt]+bo),o t =σ(W o *[h t-1 ,x t ]+b o ),
其中,ft为忘记门控,it为记忆门控,为输入数据,ot为控制当前输出;Wf为忘记门控的权重矩阵,bf为忘记门控的偏置;Wi为记忆门控的权重矩阵,bi为记忆门控的偏置;Wc为输入数据的权重矩阵,bc为输入数据的偏置;Wo为控制当前输出的权重矩阵,bo为控制当前输出的偏置;σ为激活函数;Among them, f t is the forget gate, it is the memory gate, is the input data, o t is the control current output; W f is the weight matrix of forget gate, b f is the bias of forget gate; Wi is the weight matrix of memory gate , b i is the bias of memory gate ; W c is the weight matrix of the input data, b c is the bias of the input data; W o is the weight matrix that controls the current output, bo is the bias that controls the current output; σ is the activation function;
不同状态具有不同权重矩阵W和偏置b,需要训练优化参数,得到训练参数;权重矩阵为一个(k+1)*k的矩阵,与[ht-1,xt]相乘得到长度为k的向量,即k个神经元;其中ht-1是长度为k的向量;xt为t时刻的OD值;所有的权重矩阵与偏置的初始值基于均值为0,方差为1的正态分布随机生成;Different states have different weight matrices W and bias b, and the optimization parameters need to be trained to obtain the training parameters; the weight matrix is a (k+1)*k matrix, which is multiplied by [h t-1 ,x t ] to obtain a length of A vector of k, that is, k neurons; where h t-1 is a vector of length k; x t is the OD value at time t; the initial values of all weight matrices and biases are based on a mean of 0 and a variance of 1. The normal distribution is randomly generated;
ft为忘记门控,针对对象是之前的状态Ct-1;it为记忆门控,针对对象是现在的输入则Ct由忘记门控和记忆门控为依据进行更新:f t is the forget gate, which is the previous state C t-1 for the object; i t is the memory gate, which is the current input for the object Then C t is updated according to forget gate and memory gate:
ot控制当前输出,同时ht依据ot进行更新:o t controls the current output, while h t is updated according to o t :
ht=ot*tanh(Ct),h t =o t *tanh(C t ),
LSTM计算完成后,经过一个全连接网络,使最终输出节点为z,表示产酶与否:After the LSTM calculation is completed, through a fully connected network, the final output node is z, indicating whether the enzyme is produced or not:
z=Wz*hn+bz;z=W z *h n +b z ;
其中,Wz为全连接网络的权重矩阵,bz为全连接网络的偏置,hn即上述LSTM的最终输出;Among them, W z is the weight matrix of the fully connected network, b z is the bias of the fully connected network, and h n is the final output of the above LSTM;
S2、损失函数的计算过程:S2, the calculation process of the loss function:
使用cross entropy作为损失函数,记cross entropy为LOSS,计算过程如下:Use cross entropy as the loss function, record cross entropy as LOSS, and the calculation process is as follows:
其中,c为样本量;z为所有样本经过上一步的计算后所组成的矩阵,每个样本计算得到一个向量,向量长度为2,含义为产酶或不产酶,c个样本则得到大小为c*2的矩阵;exp为自然常数e为底的指数函数;softmax函数用于将输入值的值域转换为0到1之间,以模拟概率;为预测值,经softmax计算得到,为c*2的矩阵;y为真实值,其同样为c*2的矩阵,对于每个样本,若产酶,则有向量[1,0],若不产酶,则有向量[0,1];yT为矩阵y的转置;为单位矩阵;Among them, c is the sample size; z is the matrix formed by all samples after the calculation in the previous step, each sample is calculated to obtain a vector, the vector length is 2, which means enzyme production or non-enzyme production, and c samples get the size is a matrix of c*2; exp is an exponential function with the base of natural constant e; the softmax function is used to convert the range of the input value to between 0 and 1 to simulate probability; is the predicted value, which is calculated by softmax and is a matrix of c*2; y is the real value, which is also a matrix of c*2. For each sample, if enzyme is produced, there will be a vector [1,0], if not If the enzyme is produced, there is a vector [0,1]; y T is the transpose of the matrix y; is the unit matrix;
S3、训练算法的计算过程:S3. The calculation process of the training algorithm:
使用参数优选算法Adam为训练算法,参数设置有:学习率α,α=0.001;一阶矩估计的指数衰减率β1,β1=0.9;一阶矩估计的指数衰减率β2,β2=0.999;参数ε,ε=10e-8;训练次数阈值THRESHOLD,THRESHOLD=2000;The parameter optimization algorithm Adam is used as the training algorithm, and the parameters are set as follows: learning rate α, α=0.001; exponential decay rate β 1 , β 1 =0.9 for first-order moment estimation; exponential decay rate β 2 , β 2 for first-order moment estimation =0.999; parameters ε, ε=10e-8; training times threshold THRESHOLD, THRESHOLD=2000;
参数优选算法Adam针对损失函数,损失函数以全体训练参数为自变量,即所有权重矩阵和偏置;对损失函数求梯度▽θFt(θt-1);The parameter optimization algorithm Adam is aimed at the loss function. The loss function takes all the training parameters as independent variables, that is, all weight matrices and biases; the gradient of the loss function is calculated ▽ θ F t (θ t-1 );
设当前训练次数为t,则参数优选算法Adam一轮计算过程如下:Assuming that the current training times is t, the calculation process of the parameter optimization algorithm Adam is as follows:
gt=▽θFt(θt-1),g t =▽ θ F t (θ t-1 ),
mt=β1*mt-1+(1-β1)*gt,m t =β 1 *m t-1 +(1-β 1 )*g t ,
其中,gt为一阶矩估计,为二阶矩估计;α为学习率;β1为一阶矩估计的指数衰减率;β2为二阶矩估计的指数衰减率;ε为非常小的数,其为了防止在实现中除以零;mt,vt为指数加权移动平均;为偏差修正;θt-1为上一轮训练完成的参数集合;θt为此轮训练完成的参数集合;where g t is the first-order moment estimate, is the second-order moment estimation; α is the learning rate; β1 is the exponential decay rate of the first -order moment estimation; β2 is the exponential decay rate of the second -order moment estimation; zero; m t , v t are exponentially weighted moving averages; is the bias correction; θ t-1 is the parameter set completed in the previous round of training; θ t is the parameter set completed in this round of training;
训练次数达到THRESHOLD,则训练完成;When the number of training reaches THRESHOLD, the training is completed;
所述验证数据采用交叉验证的方法,分多次重复训练验证,取验证指标的平均值作为模型效能的量化展现,并得到模型准确率的标准差;The verification data adopts the method of cross-validation, repeats the training and verification in multiple times, takes the average value of the verification indicators as the quantitative display of the model performance, and obtains the standard deviation of the accuracy rate of the model;
进一步地,所述AI模型包含:决策树模型、AdaBoost模型、SVM模型、LSTM模型Further, the AI model includes: decision tree model, AdaBoost model, SVM model, LSTM model
进一步地,所述最优AI模型为LSTM模型;Further, the optimal AI model is an LSTM model;
进一步地,所述溴百里酚蓝溶液为0.4%,PH=6.0;Further, the bromothymol blue solution is 0.4%, PH=6.0;
进一步地,所述solution溶液中,亚胺培南含量为3mg/mL;所述solution溶液中,Zn2+浓度为0.1mmol/mL;所述最后调节溶液PH,PH为7.0;Further, in the solution solution, the imipenem content is 3mg/mL; in the solution solution, the Zn concentration is 0.1mmol/mL; The pH of the final adjustment solution is 7.0;
进一步地,所述野生菌株为大肠埃希菌、恶臭假单胞菌、肺炎克雷伯、弗氏柠檬酸菌、阴沟肠杆菌、粘沙肠杆菌其中之一;Further, the wild strain is one of Escherichia coli, Pseudomonas putida, Klebsiella pneumoniae, Citrobacter freundii, Enterobacter cloacae, and Enterobacter myxa;
进一步地,所述培养一定时间,具体为在37℃培养16h-24h;Further, the culturing for a certain period of time, specifically, culturing at 37°C for 16h-24h;
进一步地,所述细菌悬浮液,具体配置过程为:用10uL的接种环刮一环菌苔到500uLPBS液中,PBS液的PH为7.4,涡旋混匀,在600nm波长调节细菌悬浮液OD值在1.3OD-1.7OD之间;Further, the specific configuration process of the bacterial suspension is as follows: scrape a bacterial lawn with a 10uL inoculation ring into a 500uL PBS solution, the pH of the PBS solution is 7.4, vortex and mix, and adjust the OD value of the bacterial suspension at a wavelength of 600nm. Between 1.3OD-1.7OD;
进一步地,所述检测混合溶液的OD值,具体为:将加入混合溶液的96孔板放入全自动酶标仪中,设置酶标仪温度为37°,每隔5分钟分别在615nm与720nm波长出检测混合溶液的OD值,检测时间为1h,导出混合溶液OD值原始数据。Further, the OD value of the detection mixed solution is specifically: put the 96-well plate into the automatic microplate reader, set the temperature of the microplate reader to 37°, and set the temperature of the microplate reader at 615nm and 720nm every 5 minutes. The wavelength is out to detect the OD value of the mixed solution, the detection time is 1h, and the original data of the OD value of the mixed solution is derived.
本发明与现有技术相比,具有如下优点和有益效果:Compared with the prior art, the present invention has the following advantages and beneficial effects:
1、本发明通过检测菌株的OD值随时间变化的趋势大大降低了肉眼观察溶液颜色变化造成的误差,具有更高的客观性;1, the present invention greatly reduces the error caused by the change of the color of the solution observed by the naked eye by detecting the trend of the OD value of the strain changing with time, and has higher objectivity;
2、本发明只需将OD值导入到LSTM模型中,即可立刻得到菌株是否能够产酶以及产酶能力的强弱,且可以直观的看到一批次中菌株产酶阳性数量与产酶阴性数量的多少,高效便捷;2. In the present invention, it is only necessary to import the OD value into the LSTM model, and it is possible to immediately obtain whether the strain can produce enzymes and the strength of the enzyme production ability, and can intuitively see the number of positive strains producing enzymes in a batch and the enzyme production capacity. The number of negatives is efficient and convenient;
3、本发明只检测两个时间点的OD值变化,就能得到菌株是否能够产生碳青霉烯酶的结果,能够节省时间;3. The present invention only detects the change of the OD value at two time points to obtain the result of whether the strain can produce carbapenemase, which can save time;
4、本发明直观的看到一批次中菌株产酶阳性数量与产酶阴性数量的多少,并且可以直接导出一批次菌株产酶结果,不必人工记录实验结果,高效便捷;4. The present invention intuitively sees the number of positive and negative enzyme-producing strains in a batch, and can directly derive the enzyme-producing results of a batch of strains without manually recording the experimental results, which is efficient and convenient;
5、本发明通过检测两个时间点OD值的变化,能快速得到弱产酶菌株是否产碳青霉烯酶的结果。5. The present invention can quickly obtain the result of whether the weak enzyme-producing strain produces carbapenemase by detecting the change of the OD value at two time points.
附图说明Description of drawings
图1是本发明所述一种运用AI模型快速检测产生碳青霉烯酶菌株的方法的方法流程图;Fig. 1 is a kind of method flow chart of the present invention that utilizes AI model to rapidly detect the method for producing carbapenemase strain;
图2是本发明所述实施例中决策树模型的测试结果图;Fig. 2 is the test result diagram of decision tree model in the described embodiment of the present invention;
图3是本发明所述实施例中AdaBoost模型的测试结果图;Fig. 3 is the test result diagram of AdaBoost model in the described embodiment of the present invention;
图4是本发明所述实施例中SVM模型的测试结果图;Fig. 4 is the test result diagram of SVM model in the described embodiment of the present invention;
图5是本发明所述实施例中LSTM模型的测试结果图;Fig. 5 is the test result diagram of LSTM model in the described embodiment of the present invention;
图6是本发明所述实施例中四个mix的准确度和标准差对比图;Fig. 6 is the accuracy and standard deviation contrast figure of four mixes in the described embodiment of the present invention;
图7是本发明所述实施例中LSTM模型结构图;Fig. 7 is the LSTM model structure diagram in the described embodiment of the present invention;
图8是本发明所述实施例中LSTM训练曲线图;Fig. 8 is the LSTM training curve diagram in the described embodiment of the present invention;
图9是本发明所述实施例中10折交叉验证示意图;9 is a schematic diagram of 10-fold cross-validation in the embodiment of the present invention;
图10是本发明所述实施例中8组时间点训练过程图;Fig. 10 is a training process diagram of 8 groups of time points in the embodiment of the present invention;
图11是本发明所述实施例中8组时间点训练结果准确率比较图;FIG. 11 is a comparison chart of the accuracy rate of training results of 8 groups of time points in the embodiment of the present invention;
图12是本发明所述实施例中LSTM模型分析菌株OD值随时间的变化趋势图;Fig. 12 is the variation trend diagram of OD value of LSTM model analysis strain over time in the embodiment of the present invention;
图13是本发明所述实施例中LSTM模型分析菌株OD值得到的产酶结果示意图。FIG. 13 is a schematic diagram of the enzyme production results obtained by analyzing the OD value of the strain by the LSTM model in the embodiment of the present invention.
具体实施方式Detailed ways
下面结合实施例及附图对本发明作进一步详细的描述,但本发明的实施方式不限于此。The present invention will be described in further detail below with reference to the embodiments and the accompanying drawings, but the embodiments of the present invention are not limited thereto.
实施例:Example:
一种运用AI模型快速检测产生碳青霉烯酶菌株的方法,如图1所示,包含以下步骤:A method for rapidly detecting carbapenemase-producing strains using an AI model, as shown in Figure 1, includes the following steps:
第一步:配制solutionA溶液:Step 1: Prepare solutionA solution:
使用Blue-Carba方法:在配置好的0.4%溴百里酚蓝溶液(pH=6.0)中加入亚胺培南,使溶液中亚胺培南的药物终含量为3mg/mL,再加入ZnSO4溶液,使溶液中Zn2+的终浓度为0.1mmol/mL,最后调节溶液pH为7.0。Using the Blue-Carba method: add imipenem to the prepared 0.4% bromothymol blue solution (pH=6.0), so that the final drug content of imipenem in the solution is 3 mg/mL, and then add ZnSO4 solution , the final concentration of Zn 2+ in the solution was 0.1 mmol/mL, and the pH of the solution was finally adjusted to 7.0.
第二步:用PBS液配制细菌悬浮液:Step 2: Prepare bacterial suspension with PBS solution:
菌株:受试菌株来自于本实验分离得到的野生菌株,包括大肠埃希菌,恶臭假单胞菌,肺炎克雷伯,弗氏柠檬酸菌,阴沟肠杆菌,粘沙肠杆菌。首先将样品在麦康凯琼脂培养基上分离纯化,之后在LB琼脂培养基上划菌苔,37℃培养16-24h,通过MALDI-TOF鉴定菌株种属。阴性对照菌株ATCC25922为实验室保存。Strains: The tested strains are from wild strains isolated in this experiment, including Escherichia coli, Pseudomonas putida, Klebsiella pneumoniae, Citrobacter freundii, Enterobacter cloacae, and Enterobacter myxa. The samples were first isolated and purified on MacConkey agar medium, then streaked on LB agar medium, cultured at 37°C for 16-24h, and the strains were identified by MALDI-TOF. The negative control strain ATCC25922 was kept in the laboratory.
配制细菌悬浮液:用10uL的接种环刮一环菌苔到500uLPBS(pH=7.4)液中,涡旋混匀,在600nm波长调节细菌悬浮液OD值在1.3OD-1.7OD之间。Preparation of bacterial suspension: scrape a loop of bacterial lawn with a 10uL inoculation ring into 500uL PBS (pH=7.4) solution, vortex and mix, and adjust the OD value of the bacterial suspension at 600nm wavelength between 1.3OD-1.7OD.
第三步:将细菌悬浮液与solution溶液混匀Step 3: Mix the bacterial suspension with the solution
用排枪吸取100ul的solutionA溶液加入到96孔板中,在吸取100ul的细菌悬浮液与solutionA溶液混匀。Pipette 100ul of solutionA solution into the 96-well plate with a drain gun, and then pipette 100ul of bacterial suspension and mix with solutionA solution.
第四步:将96孔板放入全自动酶标仪中检测混合溶液的OD值:Step 4: Put the 96-well plate into the automatic microplate reader to detect the OD value of the mixed solution:
将上一步加入混合溶液的96孔板放入全自动酶标仪中,设置酶标仪温度为37°,每隔5min分别在615nm与720nm波长处检测混合溶液的OD值,检测1h,导出混合溶液OD值原始数据。Put the 96-well plate into which the mixed solution was added in the previous step into the automatic microplate reader, set the temperature of the microplate reader to 37°, and detect the OD value of the mixed solution at the wavelengths of 615nm and 720nm every 5min, detect for 1h, and derive the mixed solution. Raw data of solution OD value.
第五步:选择AI模型,并进行模型训练和验证,得到最优AI模型,将混合溶液的OD值原始数据导入最优AI模型中,选出进行循环计算,输出特征向量,再经过一个全连接网络,分析得到菌株产碳青霉烯酶的结果Step 5: Select the AI model, and perform model training and verification to obtain the optimal AI model, import the raw data of the OD value of the mixed solution into the optimal AI model, select it for cyclic calculation, output the feature vector, and then go through a full Connect the network and analyze the results of carbapenemase production by strains
LSTM模型的设计:Design of LSTM model:
为了更加全面地验证分析AI模型在产酶分辨上的效能,本实验选取4种常见的智能模型:决策树模型、AdaBoost模型、SVM模型、LSTM模型。选用决策树的原因在于其解释性强,能训练出一个易于理解的模型。而SVM则是小样本分类算法中常用的算法,适用于本实验的样本情况。AdaBoost则是集成学习的代表之一,可以理解为集合多棵决策树的综合决策模型。LSTM则为深度学习时序模型的代表,在挖掘样本时序特征上效果突出。更具体地,决策树模型使用了CART分类回归树算法。AdaBoost模型使用了50棵决策树来实现综合决策。SVM模型则使用了高斯核函数,以处理非线性数据。LSTM模型使用了128个神经元的单层LSTM。以上模型设置是在训练集上反复验证调整,以得到的最佳选择。In order to more comprehensively verify the effectiveness of the analysis AI model in enzyme production discrimination, four common intelligent models were selected in this experiment: decision tree model, AdaBoost model, SVM model, and LSTM model. The reason for choosing a decision tree is that it is highly explanatory and can train an easy-to-understand model. The SVM is a commonly used algorithm in the small sample classification algorithm, which is suitable for the sample situation of this experiment. AdaBoost is one of the representatives of ensemble learning, which can be understood as a comprehensive decision-making model that integrates multiple decision trees. LSTM is a representative of deep learning time series models, and it has outstanding effects in mining sample time series features. More specifically, the decision tree model uses the CART classification regression tree algorithm. The AdaBoost model used 50 decision trees for comprehensive decision making. The SVM model uses a Gaussian kernel function to deal with nonlinear data. The LSTM model used a single-layer LSTM with 128 neurons. The above model settings are the best choices obtained by repeatedly validating and adjusting on the training set.
实验收集了517株菌的1个小时内的13个时间点的产酶实验OD值数据,以其中80%的数据作为AI模型的学习样本,进行模型训练,使其达到较高的精度,以构建最终模型。剩余的20%数据则作为最后验证模型精度的数据。为进一步精确地评估模型的效能,实验使用了交叉验证的方法,分多次重复训练验证,取其验证指标的平均值作为模型效能的量化展现,同时还能得到多次重复训练验证中模型准确率的标准差,以评估模型的稳定性。The experiment collected the OD value data of the enzyme production experiment of 517 strains at 13 time points within 1 hour, and used 80% of the data as the learning sample of the AI model to train the model to achieve a high accuracy. Build the final model. The remaining 20% of the data is used as the final data to verify the accuracy of the model. In order to further accurately evaluate the performance of the model, the experiment uses the method of cross-validation, repeats the training and verification for many times, and takes the average value of the verification indicators as the quantitative display of the model performance. The standard deviation of the rate to assess the stability of the model.
其中,图2为决策树模型测试结果,准确率和标准差分别为0.9766与0.0218,验证过程中,决策错误的样本数为12个,从辨别错误的样本中可以看出决策树没有能抓住时序关系,导致其错误率较高;图3为AdaBoost模型测试结果,准确率和标准差分别为0.9883与0.0157,验证过程中,错误的样本数为6个,比起决策树减少了一倍;图4为SVM模型测试结果,准确率和标准差分别为0.9902与0.01312,SVM得到了与AdaBoost同个级别的效果,只比AdaBoost多检测正确一个样本,错误的样本数为5个;图5为LSTM模型测试结果,准确率和标准差分别为0.9941与0.01251,可见准确率又提高了一个数量级,验证过程中,错误的样本数仅为3个;图6为4个模型的准确率与标准差对比。Among them, Figure 2 shows the test results of the decision tree model. The accuracy and standard deviation are 0.9766 and 0.0218 respectively. During the verification process, the number of samples with wrong decision-making is 12. It can be seen from the samples that identify the wrong decision tree. The time series relationship leads to a high error rate; Figure 3 shows the test results of the AdaBoost model. The accuracy and standard deviation are 0.9883 and 0.0157 respectively. During the verification process, the number of wrong samples is 6, which is twice as much as the decision tree; Figure 4 shows the test results of the SVM model. The accuracy and standard deviation are 0.9902 and 0.01312, respectively. SVM obtains the same level of effect as AdaBoost, and only detects one more correct sample than AdaBoost, and the number of wrong samples is 5; Figure 5 shows In the test results of the LSTM model, the accuracy and standard deviation are 0.9941 and 0.01251, respectively. It can be seen that the accuracy has increased by an order of magnitude. During the verification process, the number of wrong samples is only 3; Figure 6 shows the accuracy and standard deviation of the 4 models. Compared.
由于数据为时序数据,故使用LSTM作为实验模型。LSTM是一种循环神经网络,其通过门限机制保留有用的长期信息,去除无用的短期信息,以实现时序信息的挖掘。Since the data is time series data, LSTM is used as the experimental model. LSTM is a recurrent neural network that retains useful long-term information and removes useless short-term information through a threshold mechanism to realize the mining of time series information.
将各个时刻的OD值依次输入LSTM的循环神经模块中,LSTM经过循环计算后,输出特征向量,再经由一个全连接网络,最终得到产酶阳性结果。The OD value at each moment is input into the cyclic neural module of LSTM in turn. After cyclic calculation, LSTM outputs the feature vector, and then passes through a fully connected network to finally obtain the positive result of enzyme production.
LSTM模型详细结构图如7所示。The detailed structure diagram of the LSTM model is shown in Figure 7.
模型训练与验证Model training and validation
实验收集了517株菌的1个小时内的13个时间点的产酶实验原始OD值数据,以其中80%的数据作为AI模型的学习样本,进行模型训练,使其达到较高的精度,以构建最终模型。剩余的20%数据则作为最后验证模型精度的数据。The experiment collected the original OD value data of the enzyme production experiment of 517 strains at 13 time points within 1 hour, and used 80% of the data as the learning sample of the AI model to train the model to achieve high accuracy. to build the final model. The remaining 20% of the data is used as the final data to verify the accuracy of the model.
训练计划中,采用Adam梯度下降函数作为模型参数优化器,使用crossentropy作为损失函数,梯度截断阈值为1.0,学习率为0.001,迭代次数为2000。In the training plan, the Adam gradient descent function is used as the model parameter optimizer, the crossentropy is used as the loss function, the gradient truncation threshold is 1.0, the learning rate is 0.001, and the number of iterations is 2000.
设x为输入的OD值,即训练数据,W为LSTM模型的参数,y为输出的结果,y’为标签,即正确的结果,则训练过程如下:Let x be the input OD value, that is, the training data, W is the parameter of the LSTM model, y is the output result, y' is the label, that is, the correct result, the training process is as follows:
所述进行模型训练和验证,具体过程为:The specific process of model training and verification is as follows:
用80%数据作为学习样本,进行模型训练,用20%数据作为验证数据;Use 80% of the data as the learning sample to train the model, and use 20% of the data as the validation data;
所述模型训练采用cross entropy作为损失函数,采用Adam梯度下降方式,利用训练数据,进行多次迭代优化模型参数,使损失函数最小化;The model training adopts cross entropy as the loss function, adopts the Adam gradient descent method, and uses the training data to perform multiple iterations to optimize the model parameters to minimize the loss function;
S1,LSTM的计算过程:S1, the calculation process of LSTM:
LSTM以每个时间点OD值作为输入,时间点个数记为n,LSTM计算时遍历每个时间点,单独作为输入,进行一次循环计算;设定LSTM的神经元个数为8;设t为当前时间点;LSTM takes the OD value of each time point as input, and the number of time points is denoted as n. When LSTM calculates, it traverses each time point and uses it as input to perform a loop calculation; set the number of neurons of LSTM to 8; set t is the current point in time;
LSTM有两个传输状态,一个Ct、一个ht;LSTM的当前输入xt和上一个状态传递下来的ht-1拼接计算得到四个状态:LSTM has two transmission states, one C t and one h t ; the current input x t of LSTM and the h t-1 passed from the previous state are concatenated to obtain four states:
ft=σ(Wf*[ht-1,xt]+bf),f t =σ(W f *[h t-1 ,x t ]+b f ),
it=σ(Wi*[ht-1,xt]+bi),i t =σ(W i *[h t-1 ,x t ]+ bi ),
ot=σ(Wo*[ht-1,xt]+bo),o t =σ(W o *[h t-1 ,x t ]+b o ),
其中,ft为忘记门控,it为记忆门控,为输入数据,ot为控制当前输出;Wf为忘记门控的权重矩阵,bf为忘记门控的偏置;Wi为记忆门控的权重矩阵,bi为记忆门控的偏置;Wc为输入数据的权重矩阵,bc为输入数据的偏置;Wo为控制当前输出的权重矩阵,bo为控制当前输出的偏置;σ为激活函数;Among them, f t is the forget gate, it is the memory gate, is the input data, o t is the control current output; W f is the weight matrix of forget gate, b f is the bias of forget gate; Wi is the weight matrix of memory gate , b i is the bias of memory gate ; W c is the weight matrix of the input data, b c is the bias of the input data; W o is the weight matrix that controls the current output, bo is the bias that controls the current output; σ is the activation function;
不同状态具有不同权重矩阵W和偏置b,需要训练优化参数,得到训练参数;权重矩阵为一个(k+1)*k的矩阵,与[ht-1,xt]相乘得到长度为k的向量,即k个神经元;其中ht-1是长度为k的向量;xt为t时刻的OD值;所有的权重矩阵与偏置的初始值基于均值为0,方差为1的正态分布随机生成;Different states have different weight matrices W and bias b, and the optimization parameters need to be trained to obtain the training parameters; the weight matrix is a (k+1)*k matrix, which is multiplied by [h t-1 ,x t ] to obtain a length of A vector of k, that is, k neurons; where h t-1 is a vector of length k; x t is the OD value at time t; the initial values of all weight matrices and biases are based on a mean of 0 and a variance of 1. The normal distribution is randomly generated;
ft为忘记门控,针对对象是之前的状态Ct-1;it为记忆门控,针对对象是现在的输入则Ct由忘记门控和记忆门控为依据进行更新:f t is the forget gate, which is the previous state C t-1 for the object; i t is the memory gate, which is the current input for the object Then C t is updated according to forget gate and memory gate:
ot控制当前输出,同时ht依据ot进行更新:o t controls the current output, while h t is updated according to o t :
ht=ot*tanh(Ct),h t =o t *tanh(C t ),
LSTM计算完成后,经过一个全连接网络,使最终输出节点为z,表示产酶与否:After the LSTM calculation is completed, through a fully connected network, the final output node is z, indicating whether the enzyme is produced or not:
z=Wz*hn+bz;z=W z *h n +b z ;
其中,Wz为全连接网络的权重矩阵,bz为全连接网络的偏置,hn即上述LSTM的最终输出;Among them, W z is the weight matrix of the fully connected network, b z is the bias of the fully connected network, and h n is the final output of the above LSTM;
S2、损失函数的计算过程:S2, the calculation process of the loss function:
使用cross entropy作为损失函数,记cross entropy为LOSS,计算过程如下:Use cross entropy as the loss function, record cross entropy as LOSS, and the calculation process is as follows:
其中,c为样本量;z为所有样本经过上一步的计算后所组成的矩阵,每个样本计算得到一个向量,向量长度为2,含义为产酶或不产酶,c个样本则得到大小为c*2的矩阵;exp为自然常数e为底的指数函数;softmax函数用于将输入值的值域转换为0到1之间,以模拟概率;为预测值,经softmax计算得到,为c*2的矩阵;y为真实值,其同样为c*2的矩阵,对于每个样本,若产酶,则有向量[1,0],若不产酶,则有向量[0,1];yT为矩阵y的转置;为单位矩阵;Among them, c is the sample size; z is the matrix formed by all samples after the calculation in the previous step, each sample is calculated to obtain a vector, the vector length is 2, which means enzyme production or non-enzyme production, and c samples get the size is a matrix of c*2; exp is an exponential function with the base of natural constant e; the softmax function is used to convert the range of the input value to between 0 and 1 to simulate probability; is the predicted value, which is calculated by softmax and is a matrix of c*2; y is the real value, which is also a matrix of c*2. For each sample, if enzyme is produced, there will be a vector [1,0], if not If the enzyme is produced, there is a vector [0,1]; y T is the transpose of the matrix y; is the unit matrix;
S3、训练算法的计算过程S3, the calculation process of the training algorithm
使用参数优选算法Adam为训练算法,参数设置有:学习率α,α=0.001;一阶矩估计的指数衰减率β1,β1=0.9;一阶矩估计的指数衰减率β2,β2=0.999;参数ε,ε=10e-8;训练次数阈值THRESHOLD,THRESHOLD=2000;The parameter optimization algorithm Adam is used as the training algorithm, and the parameters are set as follows: learning rate α, α=0.001; exponential decay rate β 1 , β 1 =0.9 for first-order moment estimation; exponential decay rate β 2 , β 2 for first-order moment estimation =0.999; parameters ε, ε=10e-8; training times threshold THRESHOLD, THRESHOLD=2000;
参数优选算法Adam针对损失函数,损失函数以全体训练参数为自变量,即所有权重矩阵和偏置;对损失函数求梯度▽θFt(θt-1);The parameter optimization algorithm Adam is aimed at the loss function. The loss function takes all the training parameters as independent variables, that is, all weight matrices and biases; the gradient of the loss function is calculated ▽ θ F t (θ t-1 );
设当前训练次数为t,则参数优选算法Adam一轮计算过程如下:Assuming that the current training times is t, the calculation process of the parameter optimization algorithm Adam is as follows:
gt=▽θFt(θt-1),g t =▽ θ F t (θ t-1 ),
mt=β1*mt-1+(1-β1)*gt,m t =β 1 *m t-1 +(1-β 1 )*g t ,
其中,gt为一阶矩估计,为二阶矩估计;α为学习率;β1为一阶矩估计的指数衰减率;β2为二阶矩估计的指数衰减率;ε为非常小的数,其为了防止在实现中除以零;mt,vt为指数加权移动平均;为偏差修正;θt-1为上一轮训练完成的参数集合;θt为此轮训练完成的参数集合;where g t is the first-order moment estimate, is the second-order moment estimation; α is the learning rate; β1 is the exponential decay rate of the first -order moment estimation; β2 is the exponential decay rate of the second -order moment estimation; zero; m t , v t are exponentially weighted moving averages; is the bias correction; θ t-1 is the parameter set completed in the previous round of training; θ t is the parameter set completed in this round of training;
训练次数达到THRESHOLD,则训练完成;训练曲线如图8所示。When the number of training reaches THRESHOLD, the training is completed; the training curve is shown in Figure 8.
模型验证使用交叉验证的方式,分多次重复训练验证,取其验证指标的平均值作为模型效能的量化展现,同时还能得到多次重复训练验证中模型准确率的标准差,以评估模型的稳定性。图9为10折交叉验证的示意图,即每次验证将训练集中不同的十分之一的数据作为验证,其余的作为训练。Model validation uses the method of cross-validation to repeat the training and validation in multiple times, and takes the average value of the validation indicators as the quantitative display of the model performance. stability. Figure 9 is a schematic diagram of 10-fold cross-validation, that is, each validation uses one tenth of the different data in the training set as validation, and the rest as training.
交叉验证最终结果为:The final result of cross-validation is:
精确性:0.994174757282Accuracy: 0.994174757282
标准差:0.0125143666203Standard deviation: 0.0125143666203
精确度:0.997347480106Accuracy: 0.997347480106
回归性:0.994708994709Regression: 0.994708994709
模型验证结果分析Model Validation Results Analysis
LSTM模型最终的测试结果如图6所示,以准确率、标准差、以及检测错误的样本来展现。The final test results of the LSTM model are shown in Figure 6, which are presented in terms of accuracy, standard deviation, and samples with detected errors.
LSTM补充实验LSTM supplementary experiment
上述实验将13个时间点的OD值数据全部用上时,LSTM模型正确率以达很高,故实验进一步尝试了使用0min分别与5min,10min,15min,20min,25min,30min,35min,40min,45min这8批两个时间点的OD值,以二元组的形式作为LSTM神经网络的输入。最终得到8个对应的LSTM模型,其训练过程如图10所示。In the above experiment, when all the OD value data of 13 time points were used, the correct rate of the LSTM model was very high, so the experiment further tried to use 0min and 5min, 10min, 15min, 20min, 25min, 30min, 35min, 40min, respectively. The OD values of the 8 batches of two time points at 45min are used as the input of the LSTM neural network in the form of two-tuples. Finally, 8 corresponding LSTM models are obtained, and the training process is shown in Figure 10.
使用验证数据对LSTM的识别效果进行分析,其结果如图11所示。由图10可知,不同输入所对应的准确率都在92%以上,其中以0min和5min的OD值为输入的LSTM模型的准确率为92%,为所有模型最低,其余模型的准确率都在95%以上,其中以0min和35min的OD值为输入的LSTM模型的准确率为98.6%,为各个模型最高。Using the verification data to analyze the recognition effect of LSTM, the results are shown in Figure 11. It can be seen from Figure 10 that the accuracy rates corresponding to different inputs are all above 92%. Among them, the accuracy rate of the LSTM model with the OD value of 0min and 5min as input is 92%, which is the lowest among all models, and the accuracy rates of the other models are all The accuracy rate of the LSTM model with OD values of 0min and 35min as input is 98.6%, which is the highest among all models.
时间大于10min以后,任意两个时间点的准确率就都在95%以上,0min-20min的准确率为97.5%,0min-35min两个时间点的准确率最高达到了99%,因此利用计算机模型可以通过分析0min与10min以后任意一个时间点的OD值来迅速判断菌株是否产碳青霉烯酶,这一方法极大的减少了临床治疗上判断碳青霉烯耐药菌株的人力物力以及时间成本。图中横轴表示训练迭代的次数,纵轴表示模型的误差,以cross entropy损失函数值来表示。After the time is greater than 10min, the accuracy of any two time points is above 95%, the accuracy of 0min-20min is 97.5%, and the accuracy of the two time points of 0min-35min is up to 99%. Therefore, the computer model is used. It can be quickly judged whether the strain produces carbapenemase by analyzing the OD value at any time point after 0min and 10min. This method greatly reduces the manpower, material resources and time for judging carbapenem-resistant strains in clinical treatment. cost. The horizontal axis in the figure represents the number of training iterations, and the vertical axis represents the error of the model, which is represented by the value of the cross entropy loss function.
实验最开始分析了菌株在1个小时内的13个时间点的OD值,最终与Blue-Carba相对比的准确率为99.41%,说明利用通过AI模型分析菌株在特定波长下的OD值数据进而判断菌株是否为产碳青霉烯酶阳性菌株的方法是可行的,且具有极高的准确率与高效性。At the beginning of the experiment, the OD value of the strain at 13 time points within 1 hour was analyzed, and the final accuracy compared with Blue-Carba was 99.41%, indicating that the AI model was used to analyze the OD value of the strain at a specific wavelength. The method of judging whether the strain is a carbapenemase-positive strain is feasible, and has extremely high accuracy and high efficiency.
LSTM结果分析LSTM result analysis
将数据导入LSTM模型中后,通过LSTM模型的深度分析可以立即直观的看到菌株OD间变化的趋势,菌株溶液OD值随着时间的延长逐渐降低,阳性对照(NDM-5)OD值基本上从一开始就在0.2OD-.05OD之间,且曲线为一条稳定的直线,阴性对照菌株OD值则在1.5OD左右,其余产酶菌株的OD值则从1.5OD逐渐降到0.2OD左右,在20min基本上就可以大概判断菌株是否产碳青霉烯酶。且不同产酶强度菌株OD值变化曲线明显不同,VIM-2耐药基因的OD值变化曲线明显比NDM-1及NDM-5的缓慢的多,且同时携带NDM-1与VIM-4的菌株的OD值变化明显比只携带单一耐药基因的其他菌株变化慢,且之后一直呈缓慢上升趋势,这一现象值得仔细研究。因此亦可以通过菌株OD值的变化判断菌株产酶能力的强弱。如图12所示,且不同的OD值变化趋势也直观的表示了菌株产碳青霉烯酶能力的强弱。After importing the data into the LSTM model, through the in-depth analysis of the LSTM model, the trend of changes in OD among strains can be immediately and intuitively seen. The OD value of the strain solution gradually decreases with time, and the OD value of the positive control (NDM-5) is basically From the beginning, it was between 0.2OD-.05OD, and the curve was a stable straight line. The OD value of the negative control strain was around 1.5OD, and the OD value of the other enzyme-producing strains gradually decreased from 1.5OD to about 0.2OD. Basically in 20min, it is possible to roughly judge whether the strain produces carbapenemase. And the OD value change curve of strains with different enzyme production intensity is obviously different, the OD value change curve of VIM-2 drug resistance gene is significantly slower than that of NDM-1 and NDM-5, and the strains carrying NDM-1 and VIM-4 at the same time The change of the OD value of the strain was significantly slower than that of other strains carrying only a single drug resistance gene, and it has been showing a slow upward trend since then, which is worthy of careful study. Therefore, the enzyme-producing ability of the strain can also be judged by the change of the OD value of the strain. As shown in Figure 12, and the change trend of different OD values also intuitively indicates the strength of the strain's ability to produce carbapenemase.
如图13所示为LSTM模型可以检测多个二元组时间点的组合,这样在实验中就可以通过检测任意两个时间点菌株的OD值变化来判断菌株是否产碳青霉烯酶,且可以直接得出一批菌中有多少株产酶阳性菌株。As shown in Figure 13, the LSTM model can detect the combination of multiple binary time points, so in the experiment, it can be determined whether the strain produces carbapenemase by detecting the change of the OD value of the strain at any two time points, and The number of enzyme-producing strains in a batch of bacteria can be directly obtained.
上述实施例为本发明较佳的实施方式,但本发明的实施方式并不受上述实施例的限制,其他的任何未背离本发明的精神实质与原理下所作的改变、修饰、替代、组合、简化,均应为等效的置换方式,都包含在本发明的保护范围之内。The above-mentioned embodiments are preferred embodiments of the present invention, but the embodiments of the present invention are not limited by the above-mentioned embodiments, and any other changes, modifications, substitutions, combinations, The simplification should be equivalent replacement manners, which are all included in the protection scope of the present invention.
Claims (6)
Priority Applications (1)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| CN201910090138.8A CN109722466B (en) | 2019-01-30 | 2019-01-30 | Method for rapidly detecting strain producing carbapenemase by using AI model |
Applications Claiming Priority (1)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| CN201910090138.8A CN109722466B (en) | 2019-01-30 | 2019-01-30 | Method for rapidly detecting strain producing carbapenemase by using AI model |
Publications (2)
| Publication Number | Publication Date |
|---|---|
| CN109722466A CN109722466A (en) | 2019-05-07 |
| CN109722466B true CN109722466B (en) | 2022-03-25 |
Family
ID=66300400
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| CN201910090138.8A Active CN109722466B (en) | 2019-01-30 | 2019-01-30 | Method for rapidly detecting strain producing carbapenemase by using AI model |
Country Status (1)
| Country | Link |
|---|---|
| CN (1) | CN109722466B (en) |
Families Citing this family (1)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN116705170A (en) * | 2023-05-08 | 2023-09-05 | 宁波市医疗中心李惠利医院 | Escherichia coli strain-level identification method based on MALDI-TOF MS combined with LSTM |
Citations (4)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO2015193501A1 (en) * | 2014-06-19 | 2015-12-23 | Université Catholique de Louvain | Method for detecting enzyme activity hydrolyzing beta-lactam ring antimicrobial agents |
| CN105861630A (en) * | 2011-06-22 | 2016-08-17 | 国家医疗保健研究所 | Method for detecting the presence of carbapenemase-producing bacteria in a sample |
| WO2018203084A1 (en) * | 2017-05-04 | 2018-11-08 | Oxford Nanopore Technologies Limited | Machine learning analysis of nanopore measurements |
| CN108764050B (en) * | 2018-04-28 | 2021-02-26 | 中国科学院自动化研究所 | Method, system and equipment for recognizing skeleton behavior based on angle independence |
-
2019
- 2019-01-30 CN CN201910090138.8A patent/CN109722466B/en active Active
Patent Citations (4)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN105861630A (en) * | 2011-06-22 | 2016-08-17 | 国家医疗保健研究所 | Method for detecting the presence of carbapenemase-producing bacteria in a sample |
| WO2015193501A1 (en) * | 2014-06-19 | 2015-12-23 | Université Catholique de Louvain | Method for detecting enzyme activity hydrolyzing beta-lactam ring antimicrobial agents |
| WO2018203084A1 (en) * | 2017-05-04 | 2018-11-08 | Oxford Nanopore Technologies Limited | Machine learning analysis of nanopore measurements |
| CN108764050B (en) * | 2018-04-28 | 2021-02-26 | 中国科学院自动化研究所 | Method, system and equipment for recognizing skeleton behavior based on angle independence |
Non-Patent Citations (3)
| Title |
|---|
| Blue-carba, an easy biochemical test for detection of diverse carbapenemase producers directly from bacterial cultures;J Pires等;《J Clin Microbiol》;20131009;第51卷(第12期);第38-71、81页 * |
| 耐药性细菌碳青霉烯酶检测方法研究进展;徐建民等;《标记免疫分析与临床》;20170330;第24卷(第3期);第349-355页 * |
| 色差法检测乳品细菌数量的数学模型;王树忠等;《哈尔滨理工大学学报》;20110630;第16卷(第3期);第4281-4283页 * |
Also Published As
| Publication number | Publication date |
|---|---|
| CN109722466A (en) | 2019-05-07 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| CN102231057B (en) | Method for carrying out soft-sensing on lysine fermenting process on basis of chaos particle swarm optimization (CPSO) | |
| CN103218669B (en) | A kind of live fish cultivation water quality comprehensive forecasting method of intelligence | |
| CN101556242B (en) | Method for discriminating microorganism by utilizing Fourier infrared spectrum | |
| CN107066823A (en) | Based on plant, soil, microorganism heavy-metal contaminated soil repairing effect integrated evaluating method | |
| Chen et al. | Diagnosing of rice nitrogen stress based on static scanning technology and image information extraction | |
| CN102876816A (en) | Fermentation process statue monitoring and controlling method based on multi-sensor information fusion | |
| CN117807501B (en) | Crop growth environment monitoring method and system based on big data | |
| CN105158175A (en) | Method for identifying bacteria in water by using transmitted spectrum | |
| CN109722466B (en) | Method for rapidly detecting strain producing carbapenemase by using AI model | |
| CN112735511B (en) | A method for predicting PSII potential activity of cold-damaged cucumber based on QGA-SVR | |
| CN117034730A (en) | Dissolved oxygen prediction method for lysozyme fermentation based on DA-TCN-LSTM | |
| CN113049499A (en) | Indirect remote sensing inversion method for water total nitrogen concentration, storage medium and terminal equipment | |
| CN119884964A (en) | Reservoir water source raw water quality abnormality simulation early warning and auxiliary decision making method and system | |
| CN102175661A (en) | Online analyzer of Escherichia coli | |
| Soon et al. | Smart sensing and anomaly detection for microalgae culture based on LoRaWAN sensors and LSTM autoencoder | |
| CN114965299B (en) | Remote sensing monitoring method for aquatic plants in different life forms | |
| CN118762776A (en) | A tap water quality detection and analysis instrument system | |
| CN119375188B (en) | Water quality monitoring method and monitoring system of blue crab culture system | |
| CN118551922A (en) | A breeding management method based on photovoltaics and fisheries | |
| CN118425285A (en) | A method for detecting the concentration of potentially toxic elements in aquatic products based on artificial intelligence and environmental magnetism | |
| CN113221436B (en) | Sewage suspended matter concentration soft measurement method based on improved RBF neural network | |
| CN121090447A (en) | Methods and systems for detecting microalgal pigments and cell concentrations based on spectral inversion | |
| CN112082961A (en) | Method for testing toxicity of microbial plastics by using scenedesmus obliquus | |
| CN120142595B (en) | Method and system for identifying water body odor characteristics | |
| CN121435196A (en) | A method for predicting 2-MIB concentration in raw water based on multi-source sensing data |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| PB01 | Publication | ||
| PB01 | Publication | ||
| SE01 | Entry into force of request for substantive examination | ||
| SE01 | Entry into force of request for substantive examination | ||
| GR01 | Patent grant | ||
| GR01 | Patent grant |