CN104962567B - 6 type recombinant human papilloma virus-like particle and preparation method thereof - Google Patents
6 type recombinant human papilloma virus-like particle and preparation method thereof Download PDFInfo
- Publication number
- CN104962567B CN104962567B CN201410685769.1A CN201410685769A CN104962567B CN 104962567 B CN104962567 B CN 104962567B CN 201410685769 A CN201410685769 A CN 201410685769A CN 104962567 B CN104962567 B CN 104962567B
- Authority
- CN
- China
- Prior art keywords
- protein
- hpv
- leu
- gly
- lys
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Active
Links
Images
Landscapes
- Peptides Or Proteins (AREA)
- Medicines Containing Antibodies Or Antigens For Use As Internal Diagnostic Agents (AREA)
- Preparation Of Compounds By Using Micro-Organisms (AREA)
Abstract
Description
技术领域technical field
本发明涉及人乳头状瘤病毒的病毒样颗粒及其制备方法。更具体而言,本发明涉及一种重组的人乳头瘤病毒L1蛋白的五聚体及病毒样颗粒(Virus-like Particle,VLP)及其制备方法,及含该病毒样颗粒的疫苗组合物在预防人乳头瘤病毒感染中的应用。The invention relates to a virus-like particle of human papilloma virus and a preparation method thereof. More specifically, the present invention relates to a pentamer of recombinant human papillomavirus L1 protein and a virus-like particle (Virus-like Particle, VLP) and a preparation method thereof, and a vaccine composition containing the virus-like particle in Use in the prevention of human papillomavirus infection.
背景技术Background technique
人乳头瘤病毒(Human Papillomavirus,简称HPV)主要通过人体密切接触,如性传播的病毒,可引起人类多种增殖性上皮病变,包括乳头状瘤(疣)和瘤样病变。具体来讲,HPV诱发的疾病主要包括3大类,第1类:宫颈、阴道、女性外阴、阴茎和肛门的癌症及某些类型的头颈部肿瘤等恶性病变。100%的宫颈癌患者都是HPV感染所导致的,90%的肛门癌,40%的外阴、阴道及阴茎,12%的口咽及3%的口腔癌症归因于HPV感染。第2类:良性病变如扁平疣、尖锐湿疣等生殖器疣,是一种性传播疾病,在性行为活跃的人群中很常见。虽然生殖器疣不会像癌症一样造成严重的后果,但是病变通常会引起病人较为痛苦的临床症状如灼痛、出血和疼痛,同时产生尴尬、焦虑和自卑等负而的心理反应,而且反复治疗的过程浪费了大量的医疗资源。在世界范围内估计由非致癌性HPV(主要是6和11型)引起的生殖器疣有3000万,其中20~50%的病变中还包含有高危型HPV的混合感染。第3类:HPV感染还能引起复发性呼吸道乳头瘤( RRP),这是一种罕见的、具有潜在致命性的疾病,主要发生在青少年时期,有时,大量的乳头瘤可以引起呼吸困难并导致较小年龄儿童死亡。所以预防或治疗HPV感染对人类健康意义十分重大。Human papillomavirus (HPV) mainly through close contact with human body, such as sexually transmitted virus, can cause a variety of human proliferative epithelial lesions, including papilloma (wart) and tumor-like lesions. Specifically, HPV-induced diseases mainly include three categories, the first category: cancers of the cervix, vagina, female vulva, penis and anus, and certain types of malignant lesions such as head and neck tumors. 100% of cervical cancer patients are caused by HPV infection, 90% of anal cancer, 40% of vulva, vagina and penis, 12% of oropharyngeal and 3% of oral cancer are attributed to HPV infection. Type 2: Benign lesions such as flat warts, condyloma acuminatum and other genital warts, which are sexually transmitted diseases and are very common among sexually active people. Although genital warts will not cause serious consequences like cancer, the lesions usually cause patients with more painful clinical symptoms such as burning pain, bleeding and pain, and at the same time produce negative psychological reactions such as embarrassment, anxiety and low self-esteem, and repeated treatment The process wastes a lot of medical resources. It is estimated that there are 30 million genital warts caused by non-carcinogenic HPV (mainly
HPV是无囊膜的双链DNA病毒,主要由病毒外壳和基因组DNA组成(Bernard, Burket al. 2011)。HPV病毒外壳是由360个L1蛋白质(形成72个五聚体)和至多72个L2蛋白质构成的二十面体结构,直径55~60 nm(Howley and Lowy 2007)。病毒外壳蛋白质具有自组装特性,在体外L1蛋白质单独或与L2蛋白质共同自组装形成病毒样样颗粒(Virus-likeParticle,VLP)(Chen, Garcea et al. 2000, Finnen, Erickson et al. 2003, Buck,Cheng et al. 2008, Wang and Roden 2013)。HPV is a non-enveloped double-stranded DNA virus, mainly composed of viral coat and genomic DNA (Bernard, Burket al. 2011). The HPV viral coat is an icosahedral structure composed of 360 L1 proteins (forming 72 pentamers) and up to 72 L2 proteins, with a diameter of 55–60 nm (Howley and Lowy 2007). Viral coat proteins have self-assembly properties. In vitro, L1 proteins can self-assemble alone or together with L2 proteins to form virus-like particles (Virus-likeParticle, VLP) (Chen, Garcea et al. 2000, Finnen, Erickson et al. 2003, Buck , Cheng et al. 2008, Wang and Roden 2013).
由于HPV不能在体外细胞培养, 要获得该病毒的特异性抗原, 只能用重组DNA 技术的方法制备基因工程疫苗。重组Ll或L1/L2组装形成的病毒样颗粒VLPs,无病毒DNA,安全性好,具有和天然病毒颗粒相似的抗原表位,刺激机体后可产生中和抗体IgG和IgA,因此HPV VLPs可作为预防性疫苗,从而大大降低因感染HPV导致产生相关肿瘤的可能性(Howley and Lowy 2007) 。Since HPV cannot be cultured in vitro, the only way to obtain the specific antigen of the virus is to use recombinant DNA technology to prepare genetically engineered vaccines. Virus-like particles (VLPs) formed by recombinant L1 or L1/L2 assembly have no viral DNA, are safe, have antigenic epitopes similar to natural virus particles, and can produce neutralizing antibodies IgG and IgA after stimulating the body. Therefore, HPV VLPs can be used as Preventive vaccines, thereby greatly reducing the possibility of HPV-related tumors (Howley and Lowy 2007).
HPV疫苗研制的关键是能够大量制备高纯度、稳定的HPV抗原。在HPV疫苗抗原制备技术方面,目前较为常用的生产HPV抗原的表达系统可以分为真核表达系统及原核表达系统。常用的真核表达系统有痘病毒表达系统、昆虫杆状病毒表达系统、酵母表达系统。在真核表达系统中所表达的HPV L1能自发的形成VLP,往往只需进行简单的纯化即可获得VLP。但是由于真核表达系统的表达量低,培养成本高,给大规模工业化生产带来了极大困难。原核表达系统中利用大肠杆菌表达系统表达HPV L1蛋白质已有报道。但是由于大肠杆菌所表达的HPV L1蛋白质可溶性低,目前已知的纯化方法大多通过无盐沉淀或变性复性等步骤从蛋白质种类繁杂的细胞液中最终纯化得到HPV VLP。例如:在专利CN02129070.9中公开通过原核细胞表达和制备HPV L1多聚体的方法,其中纯化工艺包括通过3.3M尿素处理和透析复性过程;在WO-0204007专利中对L1-GST融合蛋白质的纯化方法也是通过尿素变性处理并进行透析复性;在现有技术中也有公开L1蛋白质的纯化方法是包括磷酸缓冲液超滤透析和离心,使目的蛋白沉淀再复溶的步骤。但是在这些纯化过程中蛋白质损失量大,得率低,难以在大规模生产上应用。The key to the development of HPV vaccines is the ability to produce high-purity, stable HPV antigens in large quantities. In terms of preparation technology of HPV vaccine antigens, the commonly used expression systems for producing HPV antigens can be divided into eukaryotic expression systems and prokaryotic expression systems. Commonly used eukaryotic expression systems include poxvirus expression systems, insect baculovirus expression systems, and yeast expression systems. The HPV L1 expressed in the eukaryotic expression system can spontaneously form VLP, and often only need simple purification to obtain VLP. However, due to the low expression level of the eukaryotic expression system and the high cost of cultivation, it has brought great difficulties to large-scale industrial production. In the prokaryotic expression system, it has been reported that the expression system of Escherichia coli expresses HPV L1 protein. However, due to the low solubility of the HPV L1 protein expressed by Escherichia coli, most of the currently known purification methods obtain HPV VLP from the cell fluid with various protein types through steps such as salt-free precipitation or denaturation and renaturation. For example: in the patent CN02129070.9, the method for expressing and preparing HPV L1 multimers by prokaryotic cells is disclosed, wherein the purification process includes 3.3M urea treatment and dialysis refolding process; in the WO-0204007 patent, the L1-GST fusion protein The purification method is also through urea denaturation treatment and dialysis refolding; in the prior art, the purification method of L1 protein is also disclosed, which includes the steps of phosphate buffer ultrafiltration, dialysis and centrifugation, so that the target protein is precipitated and then redissolved. However, in these purification processes, the loss of protein is large and the yield is low, which makes it difficult to apply in large-scale production.
在HPV疫苗抗原蛋白质VLP的均一性方面,现有技术中所组装的HPV L1 VLP的粒径分散度有使用polyd值表示,polyd值<15%说明颗粒有很好的均一性,15%到30%之间说明颗粒有较大的不均一性,大于30%说明颗粒完全不够均一。现有技术中制备的HPV L1 VLP多大于15%。另一个说明粒径均一的指标是PdI值,PdI值为粒径分布系数,小于0.05为高度均一的样品;0.05~0.1为准均一的样品,0.1~0.3为均一性较差的样品,大于0.3为不均一的样品。在US7205125B2专利中公开两个型别HPV L1 VLP的混合蛋白液的PdI为0.07。In terms of the homogeneity of the HPV vaccine antigen protein VLP, the particle size dispersion of the HPV L1 VLP assembled in the prior art is represented by the polyd value, and the polyd value <15% indicates that the particles have good uniformity, 15% to 30 % means that the particles have greater heterogeneity, and greater than 30% means that the particles are not uniform at all. Most of the HPV L1 VLPs prepared in the prior art are greater than 15%. Another indicator of uniform particle size is the PdI value, the PdI value is the particle size distribution coefficient, less than 0.05 is a highly uniform sample; for non-uniform samples. In the US7205125B2 patent, it is disclosed that the PdI of the mixed protein solution of two types of HPV L1 VLP is 0.07.
因此,本领域仍然需要成本低、纯度高、产量高、质量稳定的HPV L1蛋白质生产技术和大规模工业化生产重组HPV L1 VLP的新方法。Therefore, there is still a need in the art for a low-cost, high-purity, high-yield, stable-quality HPV L1 protein production technology and a new method for large-scale industrial production of recombinant HPV L1 VLP.
发明内容Contents of the invention
本发明的目的是公开一种优化的编码HPV6 L1 蛋白质的核苷酸序列,包含该核苷酸序列的载体、包括载体的宿主细胞,以及由该多核苷酸序列翻译表达的HPV L1蛋白质,Tag-HPV-L1重组蛋白,由该L1蛋白质形成的五聚体和VLP,以及由该五聚体和VLP作为抗原组成的预防HPV感染的疫苗。 The object of the present invention is to disclose an optimized nucleotide sequence encoding HPV6 L1 protein, a vector comprising the nucleotide sequence, a host cell comprising the vector, and an HPV L1 protein expressed by translation of the polynucleotide sequence, Tag - HPV-L1 recombinant protein, pentamers and VLPs formed by the L1 protein, and a vaccine for preventing HPV infection composed of the pentamers and VLPs as antigens.
本发明第一方面提供一种经密码子优化的HPV6 L1的基因,其核苷酸序列为SEQNO:2。The first aspect of the present invention provides a codon-optimized HPV6 L1 gene, the nucleotide sequence of which is SEQ NO:2.
本发明第二方面提供一种构建的表达载体,其包含本发明第一方面的经密码子优化的HPV6 L1的基因。所述载体适合驱动异源DNA在细菌中翻译表达HPV L1蛋白质。在一个实施方案中,所述表达载体优选pGEX-6p-1、pGEX-4T-2、 pMAL或pET28a。The second aspect of the present invention provides a constructed expression vector comprising the codon-optimized HPV6 L1 gene of the first aspect of the present invention. The vector is suitable for driving heterologous DNA to translate and express HPV L1 protein in bacteria. In one embodiment, the expression vector is preferably pGEX-6p-1, pGEX-4T-2, pMAL or pET28a.
本发明的第三方面提供一种构建的工程菌细胞,该细胞包含本发明第一方面的基因,或第二方面的表达载体。所述的工程菌宿主细胞是大肠杆菌,在一个实施方案中,所述宿主细胞优选BL21细胞株。The third aspect of the present invention provides a constructed engineered bacterial cell comprising the gene of the first aspect of the present invention, or the expression vector of the second aspect. The host cell of the engineering bacteria is Escherichia coli, and in one embodiment, the host cell is preferably BL21 cell strain.
本发明第四方面提供一种Tag-HPV6 L1融合蛋白,其中标签Tag为6*His.Tag,GST.Tag,SUMO.Tag,MBP.Tag,6*His - SUMO.Tag或GST- SUMO.Tag;L1为HPV6 L1全长蛋白质和/或C端截短5个、10个、15个或不多于30个氨基酸和/或N端截短2个、4个、6个或不多于10个氨基酸的L1蛋白质。The fourth aspect of the present invention provides a Tag-HPV6 L1 fusion protein, wherein the tag Tag is 6*His.Tag, GST.Tag, SUMO.Tag, MBP.Tag, 6*His-SUMO.Tag or GST-SUMO.Tag ; L1 is HPV6 L1 full-length protein and/or C-terminal truncated by 5, 10, 15 or no more than 30 amino acids and/or N-terminal truncated by 2, 4, 6 or no more than 10 amino acid L1 protein.
编码Tag-HPVL1融合蛋白GST-HPV6 L1的核苷酸序列为SEQ NO:3、SEQ NO:11,,GST-SUMO-HPV6 L1的核苷酸序列为SEQ NO:4、SEQ NO:12,MBP的核苷酸序列SEQ NO:5、 SEQNO:13,6*His-HPV6 L1的核苷酸序列为SEQ NO:6, 6*His-SUMO-HPV6 L1的核苷酸序列为SEQ NO:7。The nucleotide sequence encoding Tag-HPVL1 fusion protein GST-HPV6 L1 is SEQ NO: 3, SEQ NO: 11, the nucleotide sequence of GST-SUMO-HPV6 L1 is SEQ NO: 4, SEQ NO: 12, MBP The nucleotide sequence of SEQ NO: 5, SEQ NO: 13, the nucleotide sequence of 6*His-HPV6 L1 is SEQ NO: 6, the nucleotide sequence of 6*His-SUMO-HPV6 L1 is SEQ NO: 7.
编码Tag-HPVL1融合蛋白GST-HPV6 L1的氨基酸序列为SEQ NO:8,GST-SUMO-HPV6L1的氨基酸序列为SEQ NO:9,MBP的氨基酸序列SEQ NO:10。The amino acid sequence encoding Tag-HPVL1 fusion protein GST-HPV6 L1 is SEQ NO: 8, the amino acid sequence of GST-SUMO-HPV6L1 is SEQ NO: 9, and the amino acid sequence of MBP is SEQ NO: 10.
本发明第五方面提供Tag-HPVL1融合蛋白质经过纯化后获得的HPV L1的五聚体,及由五聚体组装的VLP。在一个优选实施例中HPV6 L1五聚体蛋白平均粒径10~15nm PdI<0.1。在一个优选实施例中HPV6 L1VLP的平均粒径45~65nm PdI<0.1。The fifth aspect of the present invention provides the HPV L1 pentamer obtained after the Tag-HPVL1 fusion protein is purified, and the VLP assembled from the pentamer. In a preferred embodiment, the average particle size of the HPV6 L1 pentameric protein is 10-15 nm PdI<0.1. In a preferred embodiment, the average particle size of HPV6 L1 VLP is 45-65nm PdI<0.1.
本发明第六方面提供了一种疫苗组合物,其包含本发明HPV L1的五聚体或HPV L1的VLP,所述组合物中进一步包含可药用的赋形剂和药用佐剂。The sixth aspect of the present invention provides a vaccine composition, which comprises the HPV L1 pentamer or HPV L1 VLP of the present invention, and the composition further comprises pharmaceutically acceptable excipients and pharmaceutically acceptable adjuvants.
在一个实施方案中将含有HPV6 L1五聚体或VLP蛋白原液(根据上述方法制备所得)分别与氢氧化铝佐剂生理盐水溶液按照蛋白与铝含量1:10比例进行吸附配制即可制得重组HPV L1蛋白质五聚体或VLP疫苗,在4℃保存待用。In one embodiment, the stock solution containing HPV6 L1 pentamer or VLP protein (prepared according to the above method) and aluminum hydroxide adjuvant physiological saline solution are prepared by adsorption according to the ratio of protein to aluminum content of 1:10 to obtain recombinant HPV L1 protein pentamer or VLP vaccine, stored at 4°C until use.
另一方面,本发明还提供一种获得Tag-HPVL1融合蛋白的方法,包括如下步骤:On the other hand, the present invention also provides a method for obtaining Tag-HPVL1 fusion protein, comprising the steps of:
A.通过用大肠杆菌偏爱的密码子取代HPV6 L1基因序列的翻译同种蛋白的密码子,得到大肠杆菌表达系统偏爱的密码子优化的HPV6 L1的基因;A. by substituting the codons of the translation homologous protein of the HPV6 L1 gene sequence with the codons favored by Escherichia coli, the gene of the codon-optimized HPV6 L1 favored by the Escherichia coli expression system is obtained;
B.构建HPV6 L1基因的大肠杆菌表达载体;B. construct the Escherichia coli expression vector of HPV6 L1 gene;
C.构建Tag-HPV6 L1的大肠杆菌表达工程菌株;C. construct the Escherichia coli expression engineering strain of Tag-HPV6 L1;
D.诱导表达并纯化得融合蛋白Tag-HPV6 L1。D. Induced expression and purification of the fusion protein Tag-HPV6 L1.
上述制备融合蛋白Tag-HPV6 L1方法中原核宿主细胞选自但不限于GI698,ER2566,BL21 (DE3),XA90,B834 (DE3),BLR (DE3)。The prokaryotic host cell in the above method for preparing fusion protein Tag-HPV6 L1 is selected from but not limited to GI698, ER2566, BL21 (DE3), XA90, B834 (DE3), BLR (DE3).
上述制备融合蛋白Tag-HPV6 L1方法中表达条件是:20~37℃温度条件下,诱导表达3~20小时。在一个具体实施例中优选在28℃温度条件下,诱导表达 16小时。The expression condition in the above method for preparing the fusion protein Tag-HPV6 L1 is: under the temperature condition of 20-37° C., the expression is induced for 3-20 hours. In a specific embodiment, expression is preferably induced for 16 hours at a temperature of 28°C.
本发明还提供一种获得HPV6 L1五聚体的方法,包括如下步骤:The present invention also provides a method for obtaining HPV6 L1 pentamers, comprising the steps of:
a)用亲和层析方法吸附融合蛋白Tag-HPV6 L1;a) Adsorb the fusion protein Tag-HPV6 L1 by affinity chromatography;
b)加入蛋白质水解酶切除Tag标签,得到HPV6 L1五聚体蛋白质;b) adding proteolytic enzymes to remove the Tag tag to obtain HPV6 L1 pentameric protein;
c)纯化HPVL1五聚体蛋白质、得到纯度>98%,平均粒径10~15nm PdI<0.1的L1五聚体蛋白质。c) Purifying the HPV L1 pentameric protein to obtain L1 pentameric protein with a purity >98% and an average particle size of 10-15 nm PdI<0.1.
上述制备HPV6 L1五聚体方法中所述用于蛋白酶为切除Tag标签的位点专一的蛋白质水解酶:重组3C蛋白酶,凝血酶,SUMO蛋白酶,SENP1或TEV蛋白酶。The protease used in the method for preparing the HPV6 L1 pentamer described above is a site-specific proteolytic enzyme that cuts off the Tag tag: recombinant 3C protease, thrombin, SUMO protease, SENP1 or TEV protease.
上述制备HPV6 L1五聚体方法中纯化方法选自但不限于离子交换色谱法,疏水性色谱法,分子筛(或称凝胶过滤或分子排阻)色谱法;优选地纯化包括离子交换色谱法和分子筛色谱法。The purification method in the above method for preparing HPV6 L1 pentamer is selected from but not limited to ion exchange chromatography, hydrophobic chromatography, molecular sieve (or gel filtration or molecular exclusion) chromatography; preferably purification includes ion exchange chromatography and Molecular sieve chromatography.
上述制备HPV6 L1五聚体方法中纯化方法还包括使用还原剂,例如加入DTT。The purification method in the above method for preparing the HPV6 L1 pentamer also includes the use of a reducing agent, such as adding DTT.
上述制备HPV6 L1五聚体方法中最终纯化后所得到HPV6 L1五聚体蛋白平均粒径10~15nm PdI<0.1。The average particle size of the HPV6 L1 pentamer protein obtained after the final purification in the above method for preparing the HPV6 L1 pentamer is 10-15 nm PdI<0.1.
本发明还提供了一种HPV6 L1五聚体组装成VLP的方法,包括如下步骤:The present invention also provides a method for assembling HPV6 L1 pentamers into VLPs, comprising the steps of:
将平均粒径10~15nm PdI<0.1的L1五聚体蛋白质液与组装缓冲液混合,最终获得pH值为5.0~5.9,盐浓度为500~2000 mM,平均粒径45~65nm PdI<0.1的HPV6 L1 VLP蛋白质液,优选获得pH值为5.7,盐浓度为1300 mM的HPV6 L1 VLP蛋白质液。Mix the L1 pentamer protein solution with an average particle size of 10-15nm PdI<0.1 with the assembly buffer, and finally obtain a pH value of 5.0-5.9, a salt concentration of 500-2000 mM, and an average particle size of 45-65nm PdI<0.1 For the HPV6 L1 VLP protein liquid, it is preferable to obtain the HPV6 L1 VLP protein liquid with a pH value of 5.7 and a salt concentration of 1300 mM.
组装缓冲液包括但不限于Tris缓冲液,磷酸盐缓冲液,醋酸缓冲液,HEPES缓冲液,MOPS缓冲液,枸橼酸缓冲液、组氨酸缓冲液,硼酸缓冲液等。Assembly buffers include but are not limited to Tris buffer, phosphate buffer, acetate buffer, HEPES buffer, MOPS buffer, citrate buffer, histidine buffer, borate buffer, etc.
上述HPV6 L1五聚体组装成VLP的方法中HPV6 L1-VLP蛋白质液中还可以加入保护剂,例如:0.01~0.1聚山梨酯80。In the above method of assembling VLPs from HPV6 L1 pentamers, a protective agent, such as 0.01-0.1 polysorbate 80, may also be added to the HPV6 L1-VLP protein solution.
本发明还提供了另一种组装VLP的方法—低温冷冻处理组装法,包括如下步骤:The present invention also provides another method for assembling VLPs—low temperature freezing treatment assembly method, comprising the following steps:
将HPV L1五聚体蛋白质液置于pH值为5.5~8.0 盐浓度为150~1000 mM条件下缓冲液,在-20~-80℃条件下完全冷冻,优选冷冻24小时后,再放置室温至蛋白质原液融解,获得平均粒径45~65nm PdI<0.1的HPV6 L1VLP蛋白质液。Put the HPV L1 pentamer protein solution in buffer solution with a pH value of 5.5 to 8.0 and a salt concentration of 150 to 1000 mM, and completely freeze it at -20 to -80°C, preferably for 24 hours, and then place it at room temperature to The protein stock solution was melted to obtain the HPV6 L1VLP protein solution with an average particle size of 45-65nm PdI<0.1.
另一方面,本发明还提供了HPV L1的五聚体、VLP和包括五聚体或VLP的疫苗组合物在制备预防HPV感染的药物中的应用。On the other hand, the present invention also provides the application of the HPV L1 pentamer, VLP and the vaccine composition comprising the pentamer or VLP in the preparation of a medicament for preventing HPV infection.
根据本发明,本发明的疫苗可采用患者可接受的形式,包括但不限于注射或鼻腔或口腔吸入或者阴道给药,优选注射剂和肌内注射。According to the present invention, the vaccine of the present invention may be in a form acceptable to the patient, including but not limited to injection or nasal or oral inhalation or vaginal administration, preferably injection and intramuscular injection.
本发明中相关术语的说明及解释Explanation and interpretation of relevant terms in the present invention
根据本发明,术语“大肠杆菌表达系统”是指由大肠杆菌(菌株)与载体组成,其中大肠杆菌(菌株)来源于市场上可得到的,在此举例但不限于: GI698,ER2566,BL21 (DE3),XA90,DH(5a)、B834 (DE3),BLR (DE3)。 According to the present invention, the term "Escherichia coli expression system" refers to the composition of Escherichia coli (bacterial strains) and vectors, wherein Escherichia coli (bacterial strains) are derived from commercially available, examples but not limited to: GI698, ER2566, BL21 ( DE3), XA90, DH(5a), B834 (DE3), BLR (DE3).
根据本发明,术语“载体”一词指的是,可将某编码蛋白质的多聚核苷酸插入其中并使蛋白质获得表达的一种核酸运载工具。载体可以通过转化,转导或者转染宿主细胞,使其携带的遗传物质元件在宿主细胞中获得表达。举例来说,载体包括:质粒;噬菌体;柯斯质粒等等。According to the present invention, the term "vector" refers to a nucleic acid delivery tool into which a polynucleotide encoding a protein can be inserted and the protein can be expressed. The vector can transform, transduce or transfect the host cell, so that the genetic material elements carried by it can be expressed in the host cell. For example, vectors include: plasmids; bacteriophages; cosmids, and the like.
根据本发明,术语“疫苗用赋形剂或载体“是指选自一种或多种,包括但不限于:pH调节剂,表面活性剂,佐剂,离子强度增强剂。例如,pH调节剂举例但不限于磷酸盐缓冲液,表面活性剂包括阳离子,阴离子或者非离子型表面活性剂。举例但不限于:聚山梨酯80。佐剂举例但不限于氢氧化铝,磷酸铝、氟氏完全佐剂、氟氏不完全佐剂等。离子强度增强剂举例但不限于氯化钠。According to the present invention, the term "vaccine excipient or carrier" refers to one or more of them, including but not limited to: pH regulators, surfactants, adjuvants, and ionic strength enhancers. For example, pH regulators are exemplified but not limited to phosphate buffer, and surfactants include cationic, anionic or nonionic surfactants. For example but not limited to: Polysorbate 80. Adjuvants include but are not limited to aluminum hydroxide, aluminum phosphate, complete Freund's adjuvant, incomplete Freund's adjuvant and the like. Ionic strength enhancers are exemplified but not limited to sodium chloride.
根据本发明,术语“色谱”包括但不限于:离子交换色谱(例如阳离子交换色谱、阴离子交换色谱)、疏水相互作用色谱、吸附色谱层析法(例如羟基磷灰石色谱)、分子筛色谱层析(凝胶过滤或分子排阻层析)、亲和色谱层析法。According to the present invention, the term "chromatography" includes, but is not limited to: ion exchange chromatography (e.g. cation exchange chromatography, anion exchange chromatography), hydrophobic interaction chromatography, adsorption chromatography (e.g. hydroxyapatite chromatography), molecular sieve chromatography (gel filtration or size exclusion chromatography), affinity chromatography.
根据本发明,在本发明获得的重组HPV L1蛋白质的方法中,缓冲液是指一种能在加入少量酸或碱和水时大大降低pH变动幅度的溶液,包括但不限于Tris缓冲液,磷酸盐缓冲液,醋酸缓冲液,HEPES缓冲液,MOPS缓冲液,枸橼酸缓冲液、组氨酸缓冲液,硼酸缓冲液等。According to the present invention, in the method for recombinant HPV L1 protein obtained in the present invention, the buffer refers to a solution that can greatly reduce the range of pH fluctuations when adding a small amount of acid or alkali and water, including but not limited to Tris buffer, phosphoric acid Salt buffer, acetate buffer, HEPES buffer, MOPS buffer, citrate buffer, histidine buffer, boric acid buffer, etc.
根据本发明,所述细胞破碎包括但不限于通过匀浆器破碎、均质机破碎、超声波处理、研磨、高压挤压、溶菌酶处理中的一项或者多项方法来实现;According to the present invention, the cell disruption includes, but is not limited to, one or more methods of homogenizer disruption, homogenizer disruption, ultrasonic treatment, grinding, high-pressure extrusion, and lysozyme treatment;
根据本发明,在本发明获得的重组HPV L1蛋白质的方法中,所用的盐包括但不限于是中性盐,特别是碱金属盐、铵盐、盐酸盐、硫酸盐,碳酸氢盐,磷酸盐或磷酸氢盐,特别是NaCI、KCl、CaCl2、NH4Cl、KCI、NH4CI、MgSO4 、(NH4)2SO4中的一种或几种。优选NaCI。所用的还原剂包括但不限于DTT,2-巯基乙醇。所用量包括但不限于2mM~lO0mM,优选10~15mM。According to the present invention, in the method for the recombinant HPV L1 protein that the present invention obtains, used salt includes but not limited to neutral salt, particularly alkali metal salt, ammonium salt, hydrochloride, sulfate, bicarbonate, phosphoric acid Salt or hydrogen phosphate, especially one or more of NaCl, KCl, CaCl2, NH4Cl, KCl, NH4Cl, MgSO4, (NH4)2SO4. NaCI is preferred. The reducing agents used include, but are not limited to, DTT, 2-mercaptoethanol. The amount used includes, but is not limited to, 2 mM to 100 mM, preferably 10 to 15 mM.
有益效果Beneficial effect
本发明提供了一种合成基因,该基因序列是根据大肠杆菌的密码子偏好进行过密码子优化的核苷酸序列,该序列编码了HPV L1蛋白氨基酸序列。研究发现经过密码子优化的核酸序列相对于未经密码子优化的核酸序列的L1蛋白的表达量有显著提高。The invention provides a synthetic gene, the gene sequence is a codon-optimized nucleotide sequence according to the codon preference of Escherichia coli, and the sequence encodes the amino acid sequence of HPV L1 protein. The study found that the expression level of L1 protein in the codon-optimized nucleic acid sequence was significantly increased compared with that in the non-codon-optimized nucleic acid sequence.
本发明公开的大肠杆菌表达系统具有表达量高、 易于培养和操作以及生产成本低等优点。但是, 仅仅使用该表达系统仍难以直接获得大量可溶性的HPV L1蛋白, 其原因在于L1蛋白极容易形成包涵体,即无生物学活性的不溶性聚合体。此外, 即使获得大量的包涵体, 为了得到有生物学活性的蛋白, 还必须对包涵体进行变性、 复性处理,这个过程往往损失大量的蛋白。为了解决这一难题,本发明采用融合技术,将L1基因与具有协助蛋白质正确折叠的蛋白如谷胱甘肽硫转移酶(GST)、SUMO、MBP、6*His- SUMO或GST- SUMO等进行融合表达,不仅蛋白的可溶性及收率有所提高,而且GST-SUMO-HPVL1,6*His-SUMO-HPVL1使得在HPV L1蛋白质N端没有外源氨基酸的残留,同时发现其中的GST-SUMO作为重组蛋白HPVL1表达的融合标签和分子伴侣, 具有抗蛋白酶水解、显著增加重组蛋白表达量以及促进靶蛋白正确折叠, 提高可溶性等功能。因此本发明采用的技术路线是在构建HPV L1蛋白表达载体时采用了标签蛋白融合技术,一方面通过标签蛋白与L1蛋白形成的融合蛋白来提高目的蛋白的可溶性、提高产量,另一方面通过GST融合标签可以利用亲和层析和蛋白水解酶切除融合质标签方法进行目的蛋白的纯化特点,从而实现了从种类繁杂的细胞裂解液中一步纯化即可获得纯度达到70%以上的HPV L1蛋白,大大提高了纯化效率,从而提高了终产物HPV L1蛋白的产量。The Escherichia coli expression system disclosed in the present invention has the advantages of high expression level, easy cultivation and operation, and low production cost. However, it is still difficult to directly obtain a large amount of soluble HPV L1 protein only by using this expression system, because the L1 protein is very easy to form inclusion bodies, that is, insoluble polymers without biological activity. In addition, even if a large number of inclusion bodies are obtained, in order to obtain biologically active proteins, the inclusion bodies must be denatured and refolded, and a large amount of protein is often lost during this process. In order to solve this problem, the present invention adopts fusion technology to combine L1 gene with proteins that assist in correct protein folding, such as glutathione sulfur transferase (GST), SUMO, MBP, 6*His-SUMO or GST-SUMO, etc. Fusion expression not only improves the solubility and yield of the protein, but also GST-SUMO-HPVL1, 6*His-SUMO-HPVL1 makes there is no residue of exogenous amino acids at the N-terminus of the HPV L1 protein, and GST-SUMO is found as The fusion tag and molecular chaperone expressed by the recombinant protein HPVL1 have the functions of resisting protease hydrolysis, significantly increasing the expression of recombinant protein, promoting the correct folding of the target protein, and improving solubility. Therefore, the technical route adopted in the present invention is to use the tag protein fusion technology when constructing the HPV L1 protein expression vector. On the one hand, the fusion protein formed by the tag protein and the L1 protein is used to improve the solubility and yield of the target protein; The fusion tag can use affinity chromatography and proteolytic enzymes to remove the fusion tag method to purify the target protein, so that the HPV L1 protein with a purity of more than 70% can be obtained in one step from a variety of cell lysates , greatly improving the purification efficiency, thereby increasing the yield of the final product HPV L1 protein.
本发明提供的先表达、分离纯化获得高纯度的HPV L1五聚体蛋白后再人工控制组装形成VLP的技术路线,可以解决当前公知技术存在的从杂蛋白种类繁多的细胞破碎液中直接纯化VLP纯度低、降解比例高,收率低的问题,得到高纯度五聚体体外组装VLP及VLP保存条件。The technical route provided by the present invention to obtain high-purity HPV L1 pentameric protein by first expressing, separating and purifying, and then manually controlling the assembly to form VLP can solve the problem of directly purifying VLP from cell disruption liquid with various miscellaneous proteins in the current known technology The problems of low purity, high degradation ratio, and low yield were obtained by in vitro assembly of high-purity pentamer VLP and storage conditions of VLP.
另外,本发明人出人意料地发现了一种新的组装条件和方法:即低温冷冻处理组装法。通过该方法得到的VLP可将冻融前组装的粒径不均一的蛋白质(PdI大于0.1)变成粒径大小符合理论预期而且均一的,PdI小于0.1的VLP,对比现有技术得到的VLP更加稳定,并且可保存于不同盐浓度、pH值范围更广泛的缓冲液中,更便于最终疫苗制剂的稀释和配制。In addition, the present inventors unexpectedly discovered a new assembly condition and method: that is, low-temperature freezing treatment assembly method. The VLP obtained by this method can transform the protein with uneven particle size (PdI greater than 0.1) assembled before freezing and thawing into a VLP with a uniform particle size in line with theoretical expectations and a PdI less than 0.1, which is more efficient than the VLP obtained by the prior art. It is stable and can be stored in buffers with different salt concentrations and a wider pH range, which is more convenient for dilution and preparation of the final vaccine preparation.
本发明经重组所得的HPV L1 VLP蛋白质,具有良好的免疫原性,可以诱导高滴度的针对同型HPV的中和抗体,预防HPV对人体的感染,是一种良好的疫苗形式。The recombined HPV L1 VLP protein of the invention has good immunogenicity, can induce high-titer neutralizing antibodies against the same type of HPV, prevents HPV from infecting human body, and is a good vaccine form.
在参考下列详述和附图后,本发明的这些和其它方面将是显然的。此处公开的所有参考文献在此均完整引用作为参考。These and other aspects of the invention will be apparent upon reference to the following detailed description and accompanying drawings. All references disclosed herein are hereby incorporated by reference in their entirety.
附图说明Description of drawings
图l:GST-HPV6 L1 蛋白质亲和与酶解后的SDS-PAGE凝胶电泳图。M泳道为蛋白质量标准泳道从上至下为:94kDa,66kDa,45kDa,33kDa,26kDa,20kdat,左泳道为亲和吸附GST-L1的树脂,右泳道为酶解后GST与L1的树脂。Figure 1: GST-HPV6 L1 protein affinity and SDS-PAGE gel electrophoresis after enzymatic hydrolysis. Lane M is the protein quality standard lane from top to bottom: 94kDa, 66kDa, 45kDa, 33kDa, 26kDa, 20kdat, the left lane is the resin for affinity adsorption of GST-L1, and the right lane is the resin for GST and L1 after enzymatic hydrolysis.
图2:GST-SUMO-HPV6 L1 蛋白质经亲和与酶解后的SDS-PAGE凝胶电泳图。M泳道为为蛋白质量标准(从上至下为:94kDa,66kDa,45kDa,33kDa,26kDa,20kda),左泳道为亲和吸附GST-SUMO -L1的树脂,右泳道为酶解后GST-SUMO与L1的树脂。Figure 2: SDS-PAGE gel electrophoresis of GST-SUMO-HPV6 L1 protein after affinity and enzymatic digestion. Lane M is the protein standard (from top to bottom: 94kDa, 66kDa, 45kDa, 33kDa, 26kDa, 20kda), the left lane is the resin for affinity adsorption of GST-SUMO-L1, and the right lane is GST-SUMO after enzymatic digestion Resin with L1.
图3:MBP-HPV6 L1 蛋白质经亲和与酶解后的SDS-PAGE凝胶电泳图。M泳道为为蛋白质量标准(从上至下为:94kDa,66kDa,45kDa,33kDa,26kDa,20kda),左泳道为亲和吸附MBP -L1的树脂,右泳道为酶解后MBP与L1的树脂。Figure 3: SDS-PAGE gel electrophoresis of MBP-HPV6 L1 protein after affinity and enzymatic digestion. Lane M is the protein standard (from top to bottom: 94kDa, 66kDa, 45kDa, 33kDa, 26kDa, 20kda), the left lane is the resin for affinity adsorption of MBP-L1, and the right lane is the resin for MBP and L1 after enzymatic hydrolysis .
图4: 6*HIS-SUMO-HPV6 L1 蛋白质经亲和与酶解后的SDS-PAGE凝胶电泳图。M泳道为为蛋白质量标准(从上至下为:94kDa,66kDa,45kDa,33kDa,26kDa,20kda),左泳道为亲和吸附6*HIS-SUMO - L1的树脂,右泳道为酶解后6*HIS-SUMO与L1的树脂。通过凝胶电泳图显示蛋白酶未切开带有6*HIS-SUMO标签的溶合蛋白。Figure 4: SDS-PAGE gel electrophoresis of 6*HIS-SUMO-HPV6 L1 protein after affinity and enzymatic digestion. Lane M is the protein standard (from top to bottom: 94kDa, 66kDa, 45kDa, 33kDa, 26kDa, 20kda), the left lane is the resin for affinity adsorption of 6*HIS-SUMO-L1, and the right lane is the 6*HIS-SUMO-L1 after enzymatic digestion. *HIS-SUMO and L1 resins. Gel electrophoresis showed that the protease did not cut the fusion protein with 6*HIS-SUMO tag.
图5:本发明经过分子筛色谱纯化后的重组HPV6 L1五聚体蛋白质SDS-PAGE凝胶电泳图。M泳道为蛋白质量标准(从上至下为:94kDa,66kDa,45kDa,33kDa,26kDa,20kda),另一泳道为HPV L1蛋白。Fig. 5: SDS-PAGE gel electrophoresis picture of recombinant HPV6 L1 pentameric protein purified by molecular sieve chromatography of the present invention. Lane M is the protein standard (from top to bottom: 94kDa, 66kDa, 45kDa, 33kDa, 26kDa, 20kda), and the other lane is HPV L1 protein.
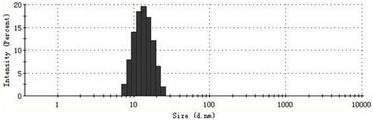
图6:HPV6 L1五聚体的动态光散射观测结果。结果显示五聚体的粒径直径为12.89nM,粒度分布PdI为 0.058。Figure 6: Dynamic light scattering observations of the HPV6 L1 pentamer. The results show that the particle diameter of the pentamer is 12.89nM, and the particle size distribution PdI is 0.058.
图7: HPV6 L1 VLP的动态光散射观测结果。结果显示VLP的粒径直径为 50.29 nM,粒度分布PdI为0.040。Figure 7: Dynamic light scattering observations of HPV6 L1 VLPs. The results showed that the particle size diameter of VLP was 50.29 nM, and the particle size distribution PdI was 0.040.
图8:HPV6 L1五聚体蛋白的透射电镜照片。Figure 8: Transmission electron micrographs of HPV6 L1 pentameric protein.
图9:HPV6 L1 VLP蛋白的透射电镜照片。Figure 9: Transmission electron micrographs of HPV6 L1 VLP protein.
图10:HPV6 L1五聚体蛋白质的高压液相分子筛色谱图,图中显示经高度纯化的L1五聚体蛋白质纯度大于98%。Figure 10: High pressure liquid phase molecular sieve chromatogram of HPV6 L1 pentamer protein, which shows that the purity of the highly purified L1 pentamer protein is greater than 98%.
图11:HPV6 L1 VLP蛋白质的高压液相分子筛色谱图,图中显示经高度纯化的VLP蛋白质纯度大于98%。Figure 11: High pressure liquid phase molecular sieve chromatogram of HPV6 L1 VLP protein, which shows that the purity of the highly purified VLP protein is greater than 98%.
图12:HPV6 L1五聚体各实验组疫苗接种小鼠后,在第二次加强免疫4周后,检测中和抗体的平均滴度水平。Figure 12: After the mice were vaccinated with the HPV6 L1 pentamer vaccine in each experimental group, the average titer levels of neutralizing antibodies were detected 4 weeks after the second booster immunization.
图13:HPV6 L1 VLP各实验组疫苗接种小鼠后,在第二次加强免疫4周后,检测中和抗体的平均滴度水平。Figure 13: The average titer levels of neutralizing antibodies were detected 4 weeks after the second booster immunization after mice were inoculated with HPV6 L1 VLP vaccine in each experimental group.
下面结合实施例对本发明进一步举例描述。这些实施例是非限制性的。Below in conjunction with embodiment the present invention is further described by way of example. These examples are non-limiting.
实施例l:密码子优化的HPV L1基因的设计与合成Embodiment 1: the design and synthesis of the HPV L1 gene of codon optimization
基因序列来源于PUBMED上已公开的各型HPV序列。参照大肠杆菌对基因转录密码子的偏好对选定的各型HPV DNA序列进行密码子优化后合成所有HPV DNA序列。根据合成DNA序列设计引物,利用合成基因为模板进行PCR扩增。所得的密码子优化序列通过DNA序列测定验证。The gene sequences were derived from the various types of HPV sequences published on PUBMED. All HPV DNA sequences were synthesized after codon optimization of the selected HPV DNA sequences according to the codon preference of Escherichia coli for gene transcription. Primers were designed according to the synthetic DNA sequence, and the synthetic gene was used as a template for PCR amplification. The resulting codon-optimized sequences were verified by DNA sequencing.
优化前与优化后的HPV各型DNA序列:DNA sequences of various types of HPV before and after optimization:
SEQ NO.1:优化前的HPV6型L1的DNA序列SEQ NO.1: DNA sequence of HPV6 type L1 before optimization
SEQ NO.2:优化后的HPV6型L1的DNA序列SEQ NO.2: DNA sequence of optimized HPV6 type L1
实施例2:重组载体pGEX-6P-1-GST- HPV6 L1的构建及鉴定:Example 2: Construction and identification of recombinant vector pGEX-6P-1-GST-HPV6 L1:
扩增HPV6 L1 的DNA片段引物:(酶切位点分别是BamHI和XhoI)Primers for amplifying the DNA fragment of HPV6 L1: (the restriction sites are BamHI and XhoI, respectively)
Forward-HPV6 L1-ApaI:5’ACTTCAGGATCC ATGTGGCGTCCGTCTGACTCTAForward-HPV6 L1-ApaI:5'ACTTCAGGATCC ATGTGGCGTCCGTCTGACTCTA
Reverse-HPV6 L1-XhoI:5’ATCTCACTCGAGCTA ACGTTTGGTT TTAGCACGTTTReverse-HPV6 L1-XhoI: 5'ATCTCACTCGAGCTA ACGTTTGGTT TTAGCACGTTT
PCR扩增反应体系:10 x Pfu buffer 20 μL,Pfu酶 4 μL,10 mM dNTP 2.5 μL,5’Primer (5μM) 10μL,3’ Primer (5μM) 10μL,模 板DNA 50 ng,加d2H2O至200μL。PCR amplification reaction system: 10 x
基因PCR扩增条件:95℃ 3 min; 95℃ 30 sec,58℃ 30 sec,72℃ 4 min;循环32次;72℃ 10 min。Gene PCR amplification conditions: 95°C for 3 min; 95°C for 30 sec, 58°C for 30 sec, 72°C for 4 min; cycle 32 times; 72°C for 10 min.
将含有BamH I和XhoI酶切位点的L1基因片段以及载体pGEX-6P-1进行BamH I/XhoI双酶切处理,之后利用T4 DNA连接酶将回收的基因片段与含有对应粘性末端的pGEX-6P-1进行连接反应,16 ℃ 10~15 h。The L1 gene fragment containing BamH I and XhoI restriction sites and the vector pGEX-6P-1 were subjected to BamH I/XhoI double digestion treatment, and then the recovered gene fragment was combined with pGEX-6P-1 containing the corresponding cohesive end using T4 DNA ligase. 6P-1 was subjected to ligation reaction at 16°C for 10-15 h.
连接反应后转化连接产物到宿主菌DH5α中进行重组子的筛选。将筛选的单克隆菌落进行扩大培养并进行质粒的提取,之后由上海生工生物公司进行测序,得到融合重组GST-HPV6-L1蛋白质的核苷酸序列为SEQ NO.3,氨基酸序列为SEQ NO.8。After the ligation reaction, the ligation product was transformed into the host strain DH5α for recombinant screening. The screened monoclonal colony was expanded and cultured and the plasmid was extracted, and then sequenced by Shanghai Sangon Biotechnology Co., Ltd., the nucleotide sequence of the fusion recombinant GST-HPV6-L1 protein was obtained as SEQ NO.3, and the amino acid sequence was SEQ NO. .8.
参照该实施例方法制备带有GST标签的融合重组载体GST-HPV-L1,其基因序列SEQNO.11。The fusion recombinant vector GST-HPV-L1 with GST tag was prepared according to the method in this example, and its gene sequence was SEQ NO.11.
实施例3:重组载体pGEX-6P-1m-GST-SUMO- HPV6 L1 载体构建Example 3: Construction of recombinant vector pGEX-6P-1m-GST-SUMO-HPV6 L1 vector
pGEX-6p-1m载体构建:为使得多酶切位点附近的ApaI酶切位点 (GGGCCC)为载体的唯一ApaI酶切位点, 在不改变lacI基因蛋白质表达序列的前提下,通过点突变技术将市售的pGEX-6p-1载体的另一ApaI识别序列GGGCCC中的Gly密码子GGC改变为它的同义密码子GGT,即可消除ApaI(3890)。通过这样的改造使得ApaI 成为可用来插入表达基因的位点。Construction of pGEX-6p-1m vector: In order to make the ApaI restriction site (GGGCCC) near the multi-restriction site the only ApaI restriction site of the vector, point mutation was carried out without changing the protein expression sequence of the lacI gene ApaI (3890) can be eliminated by changing the Gly codon GGC in another ApaI recognition sequence GGGCCC of the commercially available pGEX-6p-1 vector to its synonymous codon GGT. Through such modification, ApaI can be used to insert and express genes.
扩增SUMO 的DNA片段引物:(酶切位点分别是ApaI和BamHI)Primers for amplifying the DNA fragment of SUMO: (restriction sites are ApaI and BamHI respectively)
Forward -SUMO-ApaI: ACTTCAGGGCCCTCTGACCAGGAAGCTAAACCGTCForward -SUMO-ApaI: ACTTCAGGGCCCTCTGACCAGGAAGCTAAACCGTC
Reverse-SUMO-BamHI: CGCGGATCCACCGGTCTGTTCCTGGTAAACReverse-SUMO-BamHI: CGCGGATCCACCGGTCTGTTCCTGGTAAAC
扩增HPV6 L1 的DNA片段引物:(酶切位点分别是BamHI和XhoI)Primers for amplifying the DNA fragment of HPV6 L1: (the restriction sites are BamHI and XhoI, respectively)
Forward-HPV6 L1-ApaI:5’ACTTCAGGATCC ATGTGGCGTCCGTCTGACTCTAForward-HPV6 L1-ApaI:5'ACTTCAGGATCC ATGTGGCGTCCGTCTGACTCTA
Reverse-HPV6 L1-XhoI:5’ATCTCACTCGAGCTA ACGTTTGGTT TTAGCACGTTTReverse-HPV6 L1-XhoI: 5'ATCTCACTCGAGCTA ACGTTTGGTT TTAGCACGTTT
PCR扩增反应体系:10 x Pfu buffer 20 μL,Pfu酶 4 μL,10 mM dNTP 2.5 μL,5’Primer (5μM) 10μL,3’ Primer (5μM) 10μL,模 板DNA 50 ng,加d2H2O至200μL。PCR amplification reaction system: 10 x
基因PCR扩增条件:95℃ 1.5 min; 95℃ 30 sec,58℃ 30 sec,72℃ 1 min;循环32次;72℃ 10 min。Gene PCR amplification conditions: 1.5 min at 95°C; 30 sec at 95°C, 30 sec at 58°C, 1 min at 72°C; 32 cycles; 10 min at 72°C.
基因PCR扩增条件同上述实施例。The conditions for gene PCR amplification are the same as those in the above-mentioned examples.
酶切连接:将含有ApaI和BamHI酶切位点的SUMO基因片段以及载体pGEX-6P-1m进行Apa I/ BamHI双酶切处理,之后利用T4 DNA连接酶将回收的基因片段与含有对应粘性末端的pGEX-6P-1m进行连接反应,16 ℃ 10~15h。Enzyme digestion ligation: The SUMO gene fragment containing ApaI and BamHI restriction sites and the vector pGEX-6P-1m were subjected to ApaI/BamHI double digestion treatment, and then the recovered gene fragment was combined with the corresponding cohesive end using T4 DNA ligase The pGEX-6P-1m was used for ligation reaction at 16°C for 10-15h.
转化鉴定:连接反应后转化连接产物到宿主菌DH5α中进行重组子的筛选。将筛选的单克隆菌落进行扩大培养并进行质粒的提取,之后由苏州金唯智生物科技有限公司进行测序,得到融合重组载体pGSTSUMO-6p-1m。Transformation identification: After the ligation reaction, transform the ligated product into the host strain DH5α for recombinant screening. The screened monoclonal colony was expanded and cultured and the plasmid was extracted, and then sequenced by Suzhou Jinweizhi Biotechnology Co., Ltd. to obtain the fusion recombinant vector pGSTSUMO-6p-1m.
再次酶切连接:将含有BamHI和Xho1酶切位点的L1基因片段以及重组载体pGSTSUMO-6p-1m进行BamHI/Xho1双酶切处理,之后利用T4 DNA连接酶将回收的基因片段与含有对应粘性末端的pGSTSUMO-6p1m进行连接反应,16 ℃ 10~15 h。Reenzyme digestion and ligation: The L1 gene fragment containing BamHI and Xho1 restriction sites and the recombinant vector pGSTSUMO-6p-1m were subjected to BamHI/Xho1 double digestion treatment, and then the recovered gene fragment was combined with the corresponding cohesive DNA using T4 DNA ligase The terminal pGSTSUMO-6p1m was ligated at 16°C for 10-15 h.
再次转化鉴定:连接反应后转化连接产物到宿主菌DH5α中进行重组子的筛选。将筛选的单克隆菌落进行扩大培养并进行质粒的提取,之后由苏州金唯智生物科技有限公司进行测序,得到带有GST-SUMO标签的融合重组载体GST-SUMO-L1,其基因序列SEQ NO.4,氨基酸序列为SEQ NO.9。Re-transformation identification: after the ligation reaction, transform the ligation product into the host strain DH5α for recombinant screening. The screened monoclonal colony was expanded and cultured and the plasmid was extracted, and then sequenced by Suzhou Jinweizhi Biotechnology Co., Ltd. to obtain the fusion recombinant vector GST-SUMO-L1 with the GST-SUMO tag. Its gene sequence is SEQ NO. 4. The amino acid sequence is SEQ NO.9.
参照该实施例方法制备带有GST-SUMO标签的融合重组载体GST-SUMO-L1,其基因序列SEQ NO.12。The fusion recombinant vector GST-SUMO-L1 with the GST-SUMO tag was prepared according to the method in this example, and its gene sequence is SEQ NO.12.
实施例4:重组载体pMAL—MBP-HPV6 L1的构建Embodiment 4: Construction of recombinant vector pMAL-MBP-HPV6 L1
扩增HPV6 L1 的DNA片段引物:(酶切位点分别是EcoRI和HindIII)Primers for amplifying the DNA fragment of HPV6 L1: (the enzyme cutting sites are EcoRI and HindIII respectively)
Forward-HPV6 L1-EcoRI:5’ ACTTCA GAATTC ATGTGGCGTCCGTCTGACTCTAForward-HPV6 L1-EcoRI:5’ ACTTCA GAATTC ATGTGGCGTCCGTCTGACTCTA
Reverse-HPV6 L1-HindIII:5’ ATCTCA AAGCTTCTA ACGTTTGGTT TTAGCACGTTTReverse-HPV6 L1-HindIII:5’ ATCTCA AAGCTTCTA ACGTTTGGTT TTAGCACGTTT
将含有EcoRI和HindIII酶切位点的L1基因片段以及载体pMAL进行EcoRI/HindIII双酶切处理,之后利用T4 DNA连接酶将回收的基因片段与含有对应粘性末端的pMAL进行连接反应,16 ℃ 10~15 h。The L1 gene fragment containing EcoRI and HindIII restriction sites and the carrier pMAL were subjected to EcoRI/HindIII double digestion treatment, and then the recovered gene fragment was ligated with the pMAL containing the corresponding cohesive end using T4 DNA ligase, 16 ℃ 10 ~15 hours.
连接反应后转化连接产物到宿主菌DH5α中进行重组子的筛选。将筛选的单克隆菌落进行扩大培养并进行质粒的提取,之后由上海生工生物公司进行测序,得到融合重组MBP-HPV6-L1蛋白质的基因序列SEQ NO.5,氨基酸序列为SEQ NO.10。After the ligation reaction, the ligation product was transformed into the host strain DH5α for recombinant screening. The screened monoclonal colony was expanded and cultured and the plasmid was extracted, and then sequenced by Shanghai Sangon Biotechnology Co., Ltd. to obtain the gene sequence SEQ NO.5 of the fusion recombinant MBP-HPV6-L1 protein, and the amino acid sequence was SEQ NO.10.
参照该实施例方法制备带有MBP标签的融合重组载体MBP-HPV-L1,其基因序列SEQNO.13。The fusion recombinant vector MBP-HPV-L1 with the MBP tag was prepared according to the method in this example, and its gene sequence was SEQ NO.13.
实施例5:重组载体pET28a-6*His-HPV6 L1的构建Example 5: Construction of recombinant vector pET28a-6*His-HPV6 L1
扩增HPV6 L1 的DNA片段引物:(酶切位点分别是NdeI和XhoI, pET28a)Primers for amplifying the DNA fragment of HPV6 L1: (The enzyme cutting sites are NdeI and XhoI, respectively, pET28a)
Forward-HPV6 L1-NdeI:5’ GACTTCA CATATGATGTGGCGTCCGTCTGACTCTAForward-HPV6 L1-NdeI:5’ GACTTCA CATATGATGTGGCGTCCGTCTGACTCTA
Reverse-HPV6 L1-XhoI:5’ CATCTCACTCGAGCTA ACGTTTGGTT TTAGCACGTTTReverse-HPV6 L1-XhoI:5' CATCTCACTCGAGCTA ACGTTTGGTT TTAGCACGTTT
将含有NdeI和XhoI酶切位点的L1基因片段以及载体pMAL进行NdeI/XhoI双酶切处理,之后利用T4 DNA连接酶将回收的基因片段与含有对应粘性末端的pET28a进行连接反应,16℃ 10~15 h。The L1 gene fragment containing NdeI and XhoI restriction sites and the carrier pMAL were subjected to NdeI/XhoI double digestion treatment, and then the recovered gene fragment was ligated with pET28a containing the corresponding cohesive end using T4 DNA ligase, at 16°C for 10 ~15 hours.
连接反应后转化连接产物到宿主菌DH5α中进行重组子的筛选。将筛选的单克隆菌落进行扩大培养并进行质粒的提取,之后由上海生工生物公司进行测序,得到融合重组MBP-HPV6-L1蛋白质的基因序列SEQ NO.6。After the ligation reaction, the ligation product was transformed into the host strain DH5α for recombinant screening. The screened monoclonal colony was expanded and cultured and the plasmid was extracted, and then sequenced by Shanghai Sangon Biotechnology Co., Ltd. to obtain the gene sequence SEQ NO.6 of the fusion recombinant MBP-HPV6-L1 protein.
实施例6:重组载体6*His-SUMO-HPV6 L1 载体构建Example 6: Construction of
扩增SUMO 的DNA片段引物:(酶切位点分别是NdeI和BamHI)Primers for amplifying the DNA fragment of SUMO: (the enzyme cutting sites are NdeI and BamHI respectively)
Forward -SUMO-NdeI: GGAATTCCATATGTCTGACCAGGAAGCTAAACCGTCForward -SUMO-NdeI: GGAATTCCATATGTCTGACCAGGAAGCTAAACCGTC
Reverse-SUMO-BamHI: CGC GGATCCACCGGTCTGTTCCTGGTAAACReverse-SUMO-BamHI: CGC GGATCCACCGGTCTGTTCCTGGTAAAC
扩增HPV6 L1 的DNA片段引物:(酶切位点分别是BamHI和XhoI)Primers for amplifying the DNA fragment of HPV6 L1: (the restriction sites are BamHI and XhoI, respectively)
Forward-HPV6 L1-ApaI:5’ACTTCAGGATCC ATGTGGCGTCCGTCTGACTCTAForward-HPV6 L1-ApaI:5'ACTTCAGGATCC ATGTGGCGTCCGTCTGACTCTA
Reverse-HPV6 L1-XhoI:5’ATCTCACTCGAGCTA ACGTTTGGTT TTAGCACGTTTReverse-HPV6 L1-XhoI: 5'ATCTCACTCGAGCTA ACGTTTGGTT TTAGCACGTTT
SUMO基因、L1基因PCR扩增条件、反应体系同上述实施例所述。The PCR amplification conditions and reaction system of SUMO gene and L1 gene are the same as those described in the above examples.
酶切连接:将含有NdeI和BamHI酶切位点的SUMO基因片段以及载体pET-28a进行NdeI/ BamHI双酶切处理,之后利用T4 DNA连接酶将回收的基因片段与含有对应粘性末端的pET28a进行连接反应,16 ℃ 10~15 h。Enzyme digestion ligation: The SUMO gene fragment containing NdeI and BamHI restriction sites and the vector pET-28a were subjected to NdeI/BamHI double digestion treatment, and then the recovered gene fragment was combined with pET28a containing the corresponding sticky end using T4 DNA ligase Ligation reaction, 10-15 h at 16 °C.
转化鉴定:连接反应后转化连接产物到宿主菌DH5α中进行重组子的筛选。将筛选的单克隆菌落进行扩大培养并进行质粒的提取,之后由苏州金唯智生物科技有限公司进行测序,得到融合重组载体pETSUMO-28a。Transformation identification: After the ligation reaction, transform the ligated product into the host strain DH5α for recombinant screening. The screened monoclonal colony was expanded and cultured and the plasmid was extracted, and then sequenced by Suzhou Jinweizhi Biotechnology Co., Ltd. to obtain the fusion recombinant vector pETSUMO-28a.
再次酶切连接:将含有BamHI和Xho1酶切位点的L1基因片段以及重组载体pETSUMO-28a进行BamHI/Xho1双酶切处理,之后利用T4 DNA连接酶将回收的基因片段与含有对应粘性末端的pETSUMO-28a进行连接反应,16 ℃ 10~15 h。Reenzyme digestion and ligation: The L1 gene fragment containing BamHI and Xho1 restriction sites and the recombinant vector pETSUMO-28a were subjected to BamHI/Xho1 double digestion treatment, and then the recovered gene fragment was combined with the corresponding cohesive end using T4 DNA ligase. pETSUMO-28a was subjected to ligation reaction at 16°C for 10-15 h.
再次转化鉴定:连接反应后转化连接产物到宿主菌DH5α中进行重组子的筛选。将筛选的单克隆菌落进行扩大培养并进行质粒的提取,之后由上海生工生物公司进行测序,得到融合重组MBP-HPV6-L1蛋白质的基因序列SEQ NO.7。Re-transformation identification: after the ligation reaction, transform the ligation product into the host strain DH5α for recombinant screening. The screened monoclonal colony was expanded and cultured and the plasmid was extracted, and then sequenced by Shanghai Sangon Biotechnology Co., Ltd. to obtain the gene sequence SEQ NO.7 of the fusion recombinant MBP-HPV6-L1 protein.
实施例7:重组HPV L1五聚体蛋白质的表达Embodiment 7: Expression of recombinant HPV L1 pentameric protein
将测序结果正确实施例2、3、4、5和6的重组载体转化大肠杆菌BL21宿主细胞,并作为表达重组蛋白质的工程菌进行HPV L1蛋白的表达。工程菌培养基为2YT培养基(10 g/L胰化蛋白胨;5 g/L酵母粉;10 g/L NaCl)。挑取含重组质粒的菌体单斑于10ml 2YT培养基(含100μg/ml氨苄青霉素)中,230转/分钟(rpm),37℃振荡培养过夜。转接5 ml过夜菌于500 ml(含100 μg/ml 氨苄青霉素)2YT液体培养基中,37℃震荡培养至重组工程菌生长至OD600nm≈0.4~1时,加入终浓度0.2mM的IPTG诱导,在28℃的条件下进行6h以上重组蛋白质的诱导表达。The recombinant vectors of Examples 2, 3, 4, 5 and 6 with correct sequencing results were transformed into Escherichia coli BL21 host cells, and used as engineering bacteria expressing recombinant proteins to express HPV L1 protein. The engineering culture medium was 2YT medium (10 g/L tryptone; 5 g/L yeast powder; 10 g/L NaCl). Pick a single spot of bacteria containing the recombinant plasmid and place it in 10ml 2YT medium (containing 100μg/ml ampicillin) at 230 revolutions per minute (rpm), shaking at 37°C overnight.
细胞收集及破碎:对发酵培养物进行离心,弃上清,收获菌体沉淀,称重;使用buffer L(pH 8.0,50 mM Tris,200 mM NaCl,5mM DTT)洗涤沉淀,然后将其重悬于bufferL中进行超声波破碎,随后通过高速离心机对破菌液进行离心(16000 rpm,30 min,4℃),收集上清液。Cell collection and crushing: Centrifuge the fermentation culture, discard the supernatant, harvest the cell pellet, and weigh it; wash the pellet with buffer L (pH 8.0, 50 mM Tris, 200 mM NaCl, 5 mM DTT), and then resuspend it Ultrasonic disruption was carried out in buffer L, and then the broken bacteria liquid was centrifuged by a high-speed centrifuge (16000 rpm, 30 min, 4°C), and the supernatant was collected.
实施例8:重组HPV L1五聚体蛋白在大肠杆菌中表达量的检测Example 8: Detection of recombinant HPV L1 pentamer protein expression in Escherichia coli
采用ELISA夹心法检测亲和层析上样前Tag-HPV L1五聚体蛋白在大肠杆菌中表达量,样品及供试品:ELISA sandwich method was used to detect the expression level of Tag-HPV L1 pentamer protein in Escherichia coli before loading on affinity chromatography, samples and test items:
包被抗体:自制抗HPV6 L1小鼠单抗。Coating antibody: self-made anti-HPV6 L1 mouse monoclonal antibody.
对照品:自制高纯度的HPV6 L1蛋白。Control substance: self-made high-purity HPV6 L1 protein.
供试品:用样品稀释液将供试品Tag-HPV6 L1稀释至浓度在对照品梯度稀释浓度范围内。Test product: Dilute the test product Tag-HPV6 L1 with the sample diluent until the concentration is within the range of the gradient dilution concentration of the reference product.
酶标抗体:自制的辣根过氧化物酶标记的兔抗HPV6 L1蛋白多抗。Enzyme-labeled antibody: homemade horseradish peroxidase-labeled rabbit anti-HPV6 L1 protein polyclonal antibody.
结果计算:计算平行孔的平均值,以对照品系列浓度OD450吸收值对其相应的L1蛋白抗原作直线方程,平行样品孔间变异系数不得大于10%,直线回归方程R2不得小于0.980,将供试品的OD450吸收值代入方程计算出稀释后供试品L1蛋白抗原含量,再乘以相应的稀释倍数即为供试品中L1蛋白抗原含量,见表1。Calculation of results: calculate the average value of parallel wells, make a linear equation for its corresponding L1 protein antigen with the concentration OD of the reference substance series concentration, the coefficient of variation between parallel sample wells must not be greater than 10%, and the linear regression equation R must not be less than 0.980, Substituting the OD 450 absorption value of the test product into the equation to calculate the L1 protein antigen content of the test product after dilution, and then multiplying by the corresponding dilution factor is the L1 protein antigen content in the test product, see Table 1.
表1 检测表达后Tag-HPV L1蛋白抗原含量Table 1 Antigen content of Tag-HPV L1 protein after detection of expression
实施例9:重组HPV L1五聚体蛋白质亲和层析Embodiment 9: Recombinant HPV L1 pentamer protein affinity chromatography
带GST标签重组蛋白的亲和层析:亲和柱中装入GST琼脂糖亲和层析介质5ml,以buffer L(pH 8.0,50 mM Tris,200 mM NaCl,5mM DTT)平衡层析柱,然后上样实施例8中带有GST或GST-SUMO标签的蛋白液,完毕后以Buffer L洗至无蛋白质流出,亲和完毕。以5mLBuffer L悬浮亲和介质,取样检测并计算介质中结合L1蛋白质的总量。Affinity chromatography of recombinant protein with GST tag: 5 ml of GST agarose affinity chromatography medium is loaded into the affinity column, and the column is equilibrated with buffer L (pH 8.0, 50 mM Tris, 200 mM NaCl, 5 mM DTT), Then load the protein solution with the GST or GST-SUMO label in Example 8, and wash with Buffer L until no protein flows out, and the affinity is completed. Suspend the affinity medium with 5mL Buffer L, take a sample to detect and calculate the total amount of bound L1 protein in the medium.
带MBP标签重组蛋白的亲和层析:亲和柱中装入Amylose-Resin亲和层析介质5ml,以buffer L(pH 8.0,50 mM Tris,200 mM NaCl,5mM DTT)平衡层析柱,然后上样实施例8中带有GST或GST-SUMO标签的蛋白液,完毕后以Buffer L洗至无蛋白质流出,亲和完毕。以5mLBuffer L悬浮亲和介质,取样检测并计算介质中结合L1蛋白质的总量。Affinity chromatography of recombinant protein with MBP tag: put 5ml of Amylose-Resin affinity chromatography medium into the affinity column, equilibrate the column with buffer L (pH 8.0, 50 mM Tris, 200 mM NaCl, 5mM DTT), Then load the protein solution with the GST or GST-SUMO label in Example 8, and wash with Buffer L until no protein flows out, and the affinity is completed. Suspend the affinity medium with 5mL Buffer L, take a sample to detect and calculate the total amount of bound L1 protein in the medium.
带6*HIS标签重组蛋白的亲和层析:取5ml Ni-NTA凝胶装柱,在柱上缓慢加入10倍柱体积的平衡液(50mmol/L NaH2PO4,300mmol/L Nacl,20mmol/L imidazole,用NaOH调整PH值至8),以充分平衡Ni-NTA凝胶,流速为1ml/min。取实施例8中过滤后的带有6*His标签的上清液,完全进入凝胶后,用10倍柱体积的平衡液继续洗涤凝胶,保存流速为1ml/min。用平衡液洗脱至无蛋白质流出,亲和完毕。取样检测并计算介质中结合L1蛋白质的总量。Affinity chromatography of recombinant protein with 6*HIS tag: take 5ml Ni-NTA gel column, slowly add 10 times column volume of equilibrium solution (50mmol/L NaH2PO4, 300mmol/L Nacl, 20mmol/L imidazole , adjust the pH value to 8 with NaOH) to fully balance the Ni-NTA gel, the flow rate is 1ml/min. Take the filtered supernatant with 6*His tag in Example 8, and after it completely enters the gel, continue to wash the gel with 10 times the column volume of the equilibrium solution, and keep the flow rate at 1ml/min. Elute with equilibrium solution until no protein flows out, and the affinity is completed. Take a sample to detect and calculate the total amount of bound L1 protein in the medium.
实施例10:重组Tag-HPV L1蛋白质的酶切纯化Example 10: Enzymatic digestion and purification of recombinant Tag-HPV L1 protein
按照目的蛋白质与蛋白酶质量比100:1加入酶量,其中带有GST-HPV-L1的蛋白质用3C蛋白酶切,带有GST-SUMO-HPV-L1和6*His-SUMO-HPV-L1 的蛋白质用SENP1蛋白酶切,带有Mbp-HPV-L1的蛋白质用Factor Xa蛋白酶切,带有6*His-HPV-L1的蛋白质用Thrombin蛋白酶切,分别混合酶切2h后,分别洗脱收集各个蛋白酶切后所得的HPV6 L1五聚体蛋白质溶液。Add the amount of enzyme according to the mass ratio of the target protein to protease 100:1, the protein with GST-HPV-L1 is digested with 3C protease, and the protein with GST-SUMO-HPV-L1 and 6*His-SUMO-HPV-L1 Digest with SENP1 protease, digest protein with Mbp-HPV-L1 with Factor Xa protease, digest protein with 6*His-HPV-L1 with Thrombin protease, mix and digest for 2 hours, then elute and collect each protease The resulting HPV6 L1 pentamer protein solution.
将3C酶酶切GST标签后的L1蛋白质溶液用SDS- PAGE凝胶电泳检测,结果见图1亲和层析电泳结果,实验表明,可将90%的目的蛋白切下。图2为SENP1蛋白酶切带有GST-SUMO-HPV-L1的蛋白质,用SDS- PAGE凝胶电泳检测。图3为Factor Xa蛋白酶切带有Mbp-HPV-L1的蛋白质,用SDS- PAGE凝胶电泳检测。图1-图3说明说明得到了55kDa 的HPV6 L1蛋白。The L1 protein solution after the 3C enzyme digested the GST tag was detected by SDS-PAGE gel electrophoresis, and the results are shown in Figure 1. The results of affinity chromatography electrophoresis showed that 90% of the target protein could be cut off. Fig. 2 shows that SENP1 protease cleaves the protein with GST-SUMO-HPV-L1, which is detected by SDS-PAGE gel electrophoresis. Figure 3 shows Factor Xa protease digestion of protein with Mbp-HPV-L1, detected by SDS-PAGE gel electrophoresis. Figures 1 to 3 illustrate that HPV6 L1 protein of 55 kDa was obtained.
Thrombin蛋白酶没有切开6*His-HPV-L1的蛋白质;用SENP1酶切6*His-SUMO-L1的蛋白质溶液用SDS- PAGE凝胶电泳检测,结果见图4,显示SENP1蛋白酶未能切开带有6*His-SUMO标签的溶合蛋白。Thrombin protease did not cut 6*His-HPV-L1 protein; the protein solution of 6*His-SUMO-L1 digested with SENP1 was detected by SDS-PAGE gel electrophoresis, the results are shown in Figure 4, showing that SENP1 protease failed to cut Fusion protein with 6*His-SUMO tag.
实施例11:重组HPV L1五聚体蛋白质的纯化Embodiment 11: Purification of recombinant HPV L1 pentameric protein
分子筛色谱纯化:将上一个实施例收集的酶切纯化后的HPV6 L1五聚体蛋白质分别进行纯化,可先经过离子交换色谱收集的HPV6 L1五聚体蛋白质,或不经过离子交换步骤直接用Superdex200(GE公司生产)的凝胶过滤介质进行进一步分子筛层析,分子筛流动相为pH8.0,10 mM Tris,100 mM NaCl,收集HPV6 L1五聚体蛋白质紫外吸收峰的馏分。Molecular sieve chromatographic purification: Purify the HPV6 L1 pentamer protein collected by enzyme digestion and purification in the previous example, first through the HPV6 L1 pentamer protein collected by ion exchange chromatography, or directly use Superdex200 without ion exchange step The gel filtration medium (produced by GE Company) was further subjected to molecular sieve chromatography, and the molecular sieve mobile phase was pH8.0, 10 mM Tris, 100 mM NaCl, and the fractions of the UV absorption peak of the HPV6 L1 pentameric protein were collected.
纯化后测定样品纯度:将收集的蛋白质溶液取样用SDS- PAGE凝胶电泳检测,目的蛋白质HPV6 L1五聚体经过分子筛层析纯化后最终纯度均大于98%,详见图5,经过分子筛色谱纯化后的重组HPV6 L1五聚体蛋白质SDS-PAGE凝胶电泳图。Determination of sample purity after purification: The collected protein solution was sampled and detected by SDS-PAGE gel electrophoresis. The final purity of the target protein HPV6 L1 pentamer was greater than 98% after purification by molecular sieve chromatography. See Figure 5 for details. After purification by molecular sieve chromatography SDS-PAGE gel electrophoresis picture of recombinant HPV6 L1 pentameric protein.
测定样品蛋白浓度:用Bradford法进行蛋白浓度检测,使用标样2mg/ml BAS配制从100ug/ul稀释到500ug/ul,样品反应体系取10ul稀释的BSA+200ulBradford工作液:标准曲线为y = 0.0013 x - 0.0294 ,R² = 0.9986 ,测定样品的 OD595,代入标准曲线,计算样品的蛋白浓度,结果见表2。Determination of sample protein concentration: use the Bradford method for protein concentration detection, use the standard sample 2mg/ml BAS to prepare and dilute from 100ug/ul to 500ug/ul, take 10ul diluted BSA+200ul Bradford working solution for the sample reaction system: the standard curve is y = 0.0013 x - 0.0294, R² = 0.9986, measure the OD 595 of the sample, substitute it into the standard curve, and calculate the protein concentration of the sample. The results are shown in Table 2.
表2 Bradford法检测重组HPV6 L1五聚体蛋白浓度Table 2 Bradford method to detect recombinant HPV6 L1 pentamer protein concentration
注:样品组1为GST-HPV L1经分子筛纯化后得HPV L1五聚体蛋白溶液;样品组2为GST-SUMO-HPV L1经分子筛纯化后得HPV L1五聚体蛋白溶液;样品组3为Mbp-HPV L1经分子筛纯化后得HPV L1五聚体蛋白溶液。Note:
实施例12:重组HPV6 L1五聚体蛋白质组装成VLPEmbodiment 12: Recombinant HPV6 L1 pentamer protein is assembled into VLP
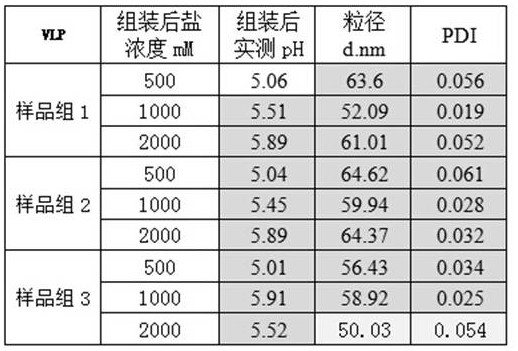
在置于如下盐浓度(NaCl)和PH值条件下,HPV L1五聚体溶液样品组1、2和3,放置稳定后,使用马尔文 Zetasizer NanoZS的动态光散射粒径仪,进行粒径及粒径分布测定(粒径分布系数PdI值为粒径分散度指标,小于0.05为高度均一的样品;0.05~0.1为准均一的样品,0.1~0.3为均一性较差的样品,大于0.3为不均一的样品),HPV6 L1五聚体蛋白组装得到粒径均一的VLP(PdI<0.05)。Under the conditions of the following salt concentration (NaCl) and pH value, HPV L1 pentamer
表3 不同pH和盐浓度条件下组装HPV6 L1 VLP的粒径检测Table 3 The particle size detection of assembled HPV6 L1 VLP under different pH and salt concentration conditions
注:样品组1为GST-HPV L1经分子筛纯化后得HPV L1 VLP蛋白溶液;样品组2为GST-SUMO-HPV L1经分子筛纯化后得HPV L1VLP蛋白溶液;样品组3为Mbp-HPV L1经分子筛纯化后得HPV L1VLP蛋白溶液。Note:
实施例13:动态光散射(DLS)对L1五聚体和VLP蛋白质粒径测定Example 13: Particle Size Determination of L1 Pentamer and VLP Proteins by Dynamic Light Scattering (DLS)
仪器为马尔文 Zetasizer NanoZS的动态光散射粒径仪,取各样品组最终制得的HPV6 L1五聚体和HPV6 L1 VLP蛋白质进行检测,测平均粒径和分散性指数PdI(表明蛋白质的均一性),说明各组样品最终制备的L1五聚体和VLP蛋白均一。其中样品组2最终制得的五聚体蛋白质和其组装获得的HPV6 L1 VLP蛋白质粒径分布详见附图6和7。The instrument is the dynamic light scattering particle size analyzer of Malvern Zetasizer NanoZS, and the HPV6 L1 pentamer and HPV6 L1 VLP protein finally prepared in each sample group are taken for detection, and the average particle size and dispersibility index PdI (indicating the homogeneity of the protein ), indicating that the final preparation of L1 pentamer and VLP protein in each group of samples was uniform. The particle size distribution of the pentameric protein finally produced in
实施例14:HPV6 L1五聚体和VLP的制备Example 14: Preparation of HPV6 L1 Pentamer and VLP
依据本发明上述实施例1-13所采用的技术,制备具有序列11,12,13的HPV6 L1蛋白,以上蛋白均可纯化得到纯度达到98%以上的蛋白,得到平均粒径10~15nm PdI<0.1的HPV6 L1五聚体蛋白。进一步组装得到平均粒径45~65nm PdI<0.1的HPV6 L1VLP蛋白。According to the technology adopted in the above-mentioned Examples 1-13 of the present invention, HPV6 L1 proteins with
实施例15:HPV6 L1 五聚体和VLP的形态学检测Example 15: Morphological detection of HPV6 L1 pentamers and VLPs
透射电镜观察:将实施例中各个纯化获得的HPV6 L1五聚体蛋白质、组装获得的HPV6 L1-VLP蛋白质,通过中国科学院生物物理所利用透射电镜平台观察。冷冻样品制备及拍照流程:Transmission electron microscope observation: The purified HPV6 L1 pentamer protein and the assembled HPV6 L1-VLP protein obtained in the examples were observed by the Institute of Biophysics, Chinese Academy of Sciences using a transmission electron microscope platform. Frozen sample preparation and photographing process:
1)将液氮盒装满液氮,待液面不沸腾时,将乙烷缓慢注入冷却的铜碗中,使之冷却为液态。1) Fill the liquid nitrogen box with liquid nitrogen, and when the liquid level is not boiling, slowly inject ethane into the cooling copper bowl to cool it into a liquid state.
2)将铜网在PDC-32型等离子清洗器做亲水性处理。2) Make the copper mesh hydrophilic in the PDC-32 plasma cleaner.
3)在Vitrobot TM Mark IV冷冻样品制备设备中,将 3.5 μL 的五聚体及VLP样品吸附在 300目的 QUANTIFOIL 铜网中,吸水4s后,通过液态乙烷冷冻样品。3) In the Vitrobot TM Mark IV frozen sample preparation equipment, 3.5 μL of pentamer and VLP samples were adsorbed on a 300-mesh QUANTIFOIL copper grid, and after absorbing water for 4 seconds, the samples were frozen by liquid ethane.
4)迅速将样品转移到液氮中保存。4) Quickly transfer the sample to liquid nitrogen for storage.
5)收集冷冻照片时,电子剂量为 20 e-/Å2。数据通过300 KV 的 300 kV TitanKrios 透射电子显微镜的 Gatan UltraScan 4000 CCD记录。加速电压为300 kV。5) When collecting cryographs, the electron dose is 20 e-/Å2. Data were recorded with a Gatan UltraScan 4000 CCD on a 300 kV TitanKrios transmission electron microscope at 300 kV. The accelerating voltage is 300 kV.
结果显示,在HPV6 L1五聚体蛋白质样品组中,视野中可见大量直径与理论大小相符的10nm左右的五聚体蛋白;在HPV6 L1-VLP蛋白质样品组中,可见颗粒大小与理论相符的大量直径为50nm左右的病毒样颗粒(VLP),均匀一致。其中GST-SUMO标签组(样品组2)酶切纯化后HPV6 L1五聚体所得样品的透射电镜照片见附图8, Mbp标签组(样品组3)酶切纯化后再组装的VLP蛋白的透射电镜照片见附图9。The results show that in the HPV6 L1 pentamer protein sample group, a large number of pentameric proteins with a diameter consistent with the theoretical size can be seen in the field of vision; in the HPV6 L1-VLP protein sample group, a large number of pentameric proteins with a particle size consistent with the theoretical size can be seen Virus-like particles (VLP) with a diameter of about 50nm, uniform and consistent. Among them, the transmission electron micrograph of the sample obtained by digesting and purifying the HPV6 L1 pentamer of the GST-SUMO tag group (sample group 2) is shown in Figure 8. The electron microscope photo is shown in attached drawing 9.
实施例16:HPV6 L1蛋白质原液纯度检测Example 16: Purity detection of HPV6 L1 protein stock solution
分子排阻高效液相色谱测定:色谱柱Agilent Bio SEC-5um,2000Å,7.8×300mm,柱体积约15 m1,分子量范围≥lO,OOOkDa;以pH6.8 的0.1mol/L磷酸盐缓冲液(称取磷酸氢二钠25.8g,磷酸二氢钠4.37g,加超纯水使溶解,用磷酸调pH至6.8,超纯水定容成1000ml)为流动相;流速为1ml/min;检测波长280nm;柱温25℃,上样量不得小于20ug,样品主峰理论塔板数不低于1000,拖尾因子小于2.0,连续进样5针,峰面积的相对标准偏差不得大于3%。Molecular exclusion high-performance liquid chromatography determination: chromatographic column Agilent Bio SEC-5um, 2000Å, 7.8 × 300mm, column volume about 15 ml, molecular weight range ≥ 10,000kDa; 0.1mol/L phosphate buffer with pH 6.8 ( Weigh 25.8g of disodium hydrogen phosphate and 4.37g of sodium dihydrogen phosphate, add ultrapure water to dissolve, adjust the pH to 6.8 with phosphoric acid, and adjust the volume to 1000ml with ultrapure water) as the mobile phase; the flow rate is 1ml/min; the detection wavelength 280nm;
取纯化后的样品2组最终制得的HPV6 L1五聚体和组装后的VLP的蛋白质原液,分别稀释浓度为1mg/ml,上样量20ul注入高压液相色谱仪,按照上述方法检测,按面积归一法计算纯度,所有处理蛋白质纯度均大于98%,结果见附图10和表4、附图11和表5。Take 2 groups of purified samples and finally prepare the HPV6 L1 pentamer and the protein stock solution of the assembled VLP, respectively dilute the concentration to 1mg/ml, inject 20ul of the sample into the high-pressure liquid chromatograph, detect according to the above method, press The purity was calculated by the area normalization method, and the purity of all treated proteins was greater than 98%. The results are shown in Figure 10 and Table 4, Figure 11 and Table 5.
表4 HPV6 L1五聚体的HPLC蛋白质纯度检测Table 4 HPLC protein purity detection of HPV6 L1 pentamer
表5 HPV6 L1组装后VLP的HPLC蛋白质纯度检测
实施例17:HPV VLP稳定性实验Embodiment 17: HPV VLP stability experiment
将各个样品组最终制得的HPV6 VLP蛋白质在下表的缓冲液条件下,在25℃放置14天至28天,进行粒径检测,结果见下表,证明HPV6 VLP在pH 5.0至5.9,盐浓度500~2000mM下存放稳定。样品组3所得HPV6 VLP在pH 5.0至5.9,盐浓度500~2000mM下放置14-28天后检测结果详见如下表。The final HPV6 VLP protein prepared by each sample group was placed at 25°C for 14 to 28 days under the buffer conditions in the following table, and the particle size was detected. Stable storage at 500-2000mM. The test results of the HPV6 VLP obtained in
表6 HPV6 L1 VLP 在25℃下放置14-28天粒径检测结果Table 6 The particle size detection results of HPV6 L1 VLP stored at 25°C for 14-28 days
实施例18:制备含有HPV L1 五聚体或VLP的单价疫苗 Example 18: Preparation of monovalent vaccine containing HPV L1 pentamer or VLP
将含有各个样品组的HPV6 L1五聚体或VLP蛋白原液分别与氢氧化铝佐剂生理盐水溶液按照蛋白与铝含量1:10比例进行吸附配制即可制得重组HPV L1蛋白质五聚体或VLP疫苗,在4℃保存待用。Recombinant HPV L1 protein pentamer or VLP can be prepared by adsorbing the HPV6 L1 pentamer or VLP protein stock solution containing each sample group with aluminum hydroxide adjuvant saline solution according to the ratio of protein to aluminum content of 1:10 Vaccines are stored at 4°C until use.
实施例19:HPV L1 五聚体和VLP的免疫原性测定Example 19: Immunogenicity Assay of HPV L1 Pentamers and VLPs
分别取上述L1五聚体或VLP疫苗,加入灭菌过的生理盐水分别稀释成20μg/ml浓度的五聚体或VLP蛋白疫苗,以每只0.1ml肌肉注射BALB/c小鼠,每组10只。小鼠每4周加强免疫一次,共免疫2次。加强免疫4周后,采用假病毒细胞中和实验法分别测定每次免疫后的小鼠血清中针对同型HPV的中和抗体滴度,结果如附图12、13所示。Take the above-mentioned L1 pentamer or VLP vaccine respectively, add sterilized normal saline and dilute to 20 μg/ml concentration of the pentamer or VLP protein vaccine, inject BALB/c mice intramuscularly with 0.1ml each, 10 in each group Only. Mice were boosted every 4 weeks for a total of 2 immunizations. Four weeks after the booster immunization, the neutralizing antibody titer against the same type of HPV in the mouse serum after each immunization was measured by the pseudovirus cell neutralization assay, and the results are shown in Figures 12 and 13 .
结果表明,HPV L1五聚体和VLP蛋白疫苗接种小鼠,二次免疫后4周中和抗体即可达到很高的水平。实验结果证明,HPV L1五聚体和组装的VLP疫苗均可以在动物体内产生中和抗体,说明HPV L1五聚体和VLP蛋白质疫苗在人体临床试验中都具有免疫原性,即能预防HPV同型病毒引起的疾病。The results showed that HPV L1 pentamer and VLP protein vaccinated mice, and neutralizing antibodies could reach a very high level 4 weeks after the second immunization. The experimental results prove that both the HPV L1 pentamer and the assembled VLP vaccine can produce neutralizing antibodies in animals, indicating that both the HPV L1 pentamer and the VLP protein vaccine have immunogenicity in human clinical trials, that is, they can prevent HPV isotypes Diseases caused by viruses.
SEQUENCE LISTING SEQUENCE LISTING
<110> 北京康乐卫士生物技术股份有限公司<110> Beijing Kangleguard Biotechnology Co., Ltd.
<120> 6型重组人乳头瘤病毒病毒样颗粒及其制备方法<120>
<130> 2014<130> 2014
<160> 13<160> 13
<170> PatentIn version 3.3<170> PatentIn version 3.3
<210> 1<210> 1
<211> 1503<211> 1503
<212> DNA<212>DNA
<213> 人工序列<213> Artificial sequence
<400> 1<400> 1
atgtggcggc ctagcgacag cacagtatat gtgcctcctc ctaaccctgt atccaaagtt 60atgtggcggc ctagcgacag cacagtatat gtgcctcctc ctaaccctgt atccaaagtt 60
gttgccacgg atgcttatgt tactcgcacc aacatatttt atcatgccag cagttctaga 120gttgccacgg atgcttatgt tactcgcacc aacatatttt atcatgccag cagttctaga 120
cttcttgcag tgggacatcc ttatttttcc ataaaacggg ctaacaaaac tgttgtgcca 180cttcttgcag tgggacatcc ttattttcc ataaaacggg ctaacaaaac tgttgtgcca 180
aaggtgtcag gatatcaata cagggtattt aaggtggtgt taccagatcc taacaaattt 240aaggtgtcag gatatcaata cagggtattt aaggtggtgt taccagatcc taacaaattt 240
gcattgcctg actcgtctct tttcgatccc acaacacaac gtttagtatg ggcatgcaca 300gcattgcctg actcgtctct tttcgatccc acaacacaac gtttagtatg ggcatgcaca 300
ggcctagagg tgggcagggg acagccatta ggtgtgggtg taagtggaca tcctttccta 360ggcctagagg tgggcagggg acagccatta ggtgtgggtg taagtggaca tcctttccta 360
aataaatatg atgatgttga aaattcaggg agtggtggta accctggaca ggataacagg 420aataaatatg atgatgttga aaattcaggg agtggtggta accctggaca ggataacagg 420
gttaatgtag gtatggatta taaacaaaca caattatgca tggttggatg tgccccccct 480gttaatgtag gtatggatta taaacaaaca caattatgca tggttggatg tgccccccct 480
ttgggcgagc attggggtaa aggtaaacag tgtactaata cacctgtaca ggctggtgac 540ttgggcgagc attggggtaa aggtaaacag tgtactaata cacctgtaca ggctggtgac 540
tgcccgccct tagaacttat taccagtgtt atacaggatg gcgatatggt tgacacaggc 600tgcccgccct tagaacttat taccagtgtt atacagatg gcgatatggt tgacacaggc 600
tttggtgcta tgaattttgc tgatttgcag accaataaat cagatgttcc tattgacata 660tttggtgcta tgaattttgc tgatttgcag accaataaat cagatgttcc tattgacata 660
tgtggcacta catgtaaata tccagattat ttacaaatgg ctgcagaccc atatggtgat 720tgtggcacta catgtaaata tccagattat ttacaaatgg ctgcagaccc atatggtgat 720
agattatttt tttttctacg gaaggaacaa atgtttgcca gacatttttt taacagggct 780agattatttt tttttctacg gaaggaacaa atgtttgcca gacatttttt taacagggct 780
ggcgaggtgg gggaacctgt gcctgataca cttataatta agggtagtgg aaatcgcacg 840ggcgaggtgg gggaacctgt gcctgataca cttataatta agggtagtgg aaatcgcacg 840
tctgtaggga gtagtatata tgttaacacc ccgagcggct ctttggtgtc ctctgaggca 900tctgtaggga gtagtatata tgttaacacc ccgagcggct ctttggtgtc ctctgaggca 900
caattgttta ataagccata ttggctacaa aaagcccagg gacataacaa tggtatttgt 960caattgttta ataagccata ttggctacaa aaagcccagg gacataacaa tggtatttgt 960
tggggtaatc aactgtttgt tactgtggta gataccacac gcagtaccaa catgacatta 1020tggggtaatc aactgtttgt tactgtggta gataccacac gcagtaccaa catgacatta 1020
tgtgcatccg taactacatc ttccacatac accaattctg attataaaga gtacatgcgt 1080tgtgcatccg taactacatc ttccacatac accaattctg attataaaga gtacatgcgt 1080
catgtggaag agtatgattt acaatttatt tttcaattat gtagcattac attgtctgct 1140catgtggaag agtatgattt acaatttatt tttcaattat gtagcattac attgtctgct 1140
gaagtaatgg cctatattca cacaatgaat ccctctgttt tggaagactg gaactttggg 1200gaagtaatgg cctatattca cacaatgaat ccctctgttt tggaagactg gaactttggg 1200
ttatcgcctc ccccaaatgg tacattagaa gatacctata ggtatgtgca gtcacaggcc 1260ttatcgcctc ccccaaatgg tacattagaa gatacctata ggtatgtgca gtcacaggcc 1260
attacctgtc aaaagcccac tcctgaaaag gaaaagccag atccctataa gaaccttagt 1320attacctgtc aaaagcccac tcctgaaaag gaaaagccag atccctataa gaaccttagt 1320
ttttgggagg ttaatttaaa agaaaagttt tctagtgaat tggatcagta tcctttggga 1380ttttgggagg ttaatttaaa agaaaagttt tctagtgaat tggatcagta tcctttggga 1380
cgcaagtttt tgttacaaag tggatatagg ggacggtcct ctattcgtac aggtgttaag 1440cgcaagtttt tgttacaaag tggatatagg ggacggtcct ctattcgtac aggtgttaag 1440
cgccctgctg tttccaaagc ctctgctgcc cctaaacgta agcgcgccaa aactaaaagg 1500cgccctgctg tttccaaagc ctctgctgcc cctaaacgta agcgcgccaa aactaaaagg 1500
taa 1503taa 1503
<210> 2<210> 2
<211> 1500<211> 1500
<212> DNA<212>DNA
<213> 人工序列<213> Artificial sequence
<400> 2<400> 2
atgtggcgtc cgtctgactc taccgtttac gttccgccgc cgaacccggt ttctaaagtt 60atgtggcgtc cgtctgactc taccgtttac gttccgccgc cgaacccggt ttctaaagtt 60
gttgctaccg acgcttacgt tacccgtacc aacatcttct accacgcttc ttcttctcgt 120gttgctaccg acgcttacgt tacccgtacc aacatcttct accacgcttc ttcttctcgt 120
ctgctggctg ttggtcaccc gtacttctct atcaaacgtg ctaacaaaac cgttgttccg 180ctgctggctg ttggtcaccc gtacttctct atcaaacgtg ctaacaaaac cgttgttccg 180
aaagtttctg gttaccagta ccgtgttttc aaagttgttc tgccggaccc gaacaaattc 240aaagtttctg gttaccagta ccgtgttttc aaagttgttc tgccggaccc gaacaaattc 240
gctctgccgg actcttctct gttcgacccg accacccagc gtctggtttg ggcttgcacc 300gctctgccgg actcttctct gttcgacccg accacccagc gtctggtttg ggcttgcacc 300
ggtctggaag ttggtcgtgg tcagccgctg ggtgttggtg tttctggtca cccgttcctg 360ggtctggaag ttggtcgtgg tcagccgctg ggtgttggtg tttctggtca cccgttcctg 360
aacaaatacg acgacgttga aaactctggt tctggtggta acccgggtca ggacaaccgt 420aacaaatacg acgacgttga aaactctggt tctggtggta acccgggtca ggacaaccgt 420
gttaacgttg gtatggacta caaacagacc cagctgtgca tggttggttg cgctccgccg 480gttaacgttg gtatggacta caaacagacc cagctgtgca tggttggttg cgctccgccg 480
ctgggtgaac actggggtaa aggtaaacag tgcaccaaca ccccggttca ggctggtgac 540ctgggtgaac actggggtaa aggtaaacag tgcaccaaca ccccggttca ggctggtgac 540
tgcccgccgc tggaactgat cacctctgtt atccaggacg gtgacatggt tgacaccggt 600tgcccgccgc tggaactgat cacctctgtt atccaggacg gtgacatggt tgacaccggt 600
ttcggtgcta tgaacttcgc tgacctgcag accaacaaat ctgacgttcc gatcgacatc 660ttcggtgcta tgaacttcgc tgacctgcag accaacaaat ctgacgttcc gatcgacatc 660
tgcggtacca cctgcaaata cccggactac ctgcagatgg ctgctgaccc gtacggtgac 720tgcggtacca cctgcaaata cccggactac ctgcagatgg ctgctgaccc gtacggtgac 720
cgtctgttct tcttcctgcg taaagaacag atgttcgctc gtcacttctt caaccgtgct 780cgtctgttct tcttcctgcg taaagaacag atgttcgctc gtcacttctt caaccgtgct 780
ggtgaagttg gtgaaccggt tccggacacc ctgatcatca aaggttctgg taaccgtacc 840ggtgaagttg gtgaaccggt tccggacacc ctgatcatca aaggttctgg taaccgtacc 840
tctgttggtt cttctatcta cgttaacacc ccgtctggtt ctctggtttc ttctgaagct 900tctgttggtt cttctatcta cgttaacacc ccgtctggtt ctctggtttc ttctgaagct 900
cagctgttca acaaaccgta ctggctgcag aaagctcagg gtcacaacaa cggtatctgc 960cagctgttca acaaaccgta ctggctgcag aaagctcagg gtcacaacaa cggtatctgc 960
tggggtaacc agctgttcgt taccgttgtt gacaccaccc gttctaccaa catgaccctg 1020tggggtaacc agctgttcgt taccgttgtt gacaccaccc gttctaccaa catgaccctg 1020
tgcgcttctg ttaccacctc ttctacctac accaactctg actacaaaga atacatgcgt 1080tgcgcttctg ttaccacctc ttctacctac accaactctg actacaaaga atacatgcgt 1080
cacgttgaag aatacgacct gcagttcatc ttccagctgt gctctatcac cctgtctgct 1140cacgttgaag aatacgacct gcagttcatc ttccagctgt gctctatcac cctgtctgct 1140
gaagttatgg cttacatcca caccatgaac ccgtctgttc tggaagactg gaacttcggt 1200gaagttatgg cttacatcca caccatgaac ccgtctgttc tggaagactg gaacttcggt 1200
ctgtctccgc cgccgaacgg taccctggaa gacacctacc gttacgttca gtctcaggct 1260ctgtctccgc cgccgaacgg taccctggaa gacacctacc gttacgttca gtctcaggct 1260
atcacctgcc agaaaccgac cccggaaaaa gaaaaaccgg acccgtacaa aaacctgtct 1320atcacctgcc agaaaccgac cccggaaaaa gaaaaaccgg acccgtacaa aaacctgtct 1320
ttctgggaag ttaacctgaa agaaaaattc tcttctgaac tggaccagta cccgctgggt 1380ttctgggaag ttaacctgaa agaaaaattc tcttctgaac tggaccagta cccgctgggt 1380
cgtaaattcc tgctgcagtc tggttaccgt ggtcgttctt ctatccgtac cggtgttaaa 1440cgtaaattcc tgctgcagtc tggttaccgt ggtcgttctt ctatccgtac cggtgttaaa 1440
cgtccggctg tttctaaagc ttctgctgct ccgaaacgta aacgtgctaa aaccaaacgt 1500cgtccggctg tttctaaagc ttctgctgct ccgaaacgta aacgtgctaa aaccaaacgt 1500
<210> 3<210> 3
<211> 2193<211> 2193
<212> DNA<212>DNA
<213> 人工序列<213> Artificial sequence
<400> 3<400> 3
atgtccccta tactaggtta ttggaaaatt aagggccttg tgcaacccac tcgacttctt 60atgtccccta tactaggtta ttggaaaatt aagggccttg tgcaacccac tcgacttctt 60
ttggaatatc ttgaagaaaa atatgaagag catttgtatg agcgcgatga aggtgataaa 120ttggaatatc ttgaagaaaa atatgaagag catttgtatg agcgcgatga aggtgataaa 120
tggcgaaaca aaaagtttga attgggtttg gagtttccca atcttcctta ttatattgat 180tggcgaaaca aaaagtttga attgggtttg gagtttccca atcttcctta ttatattgat 180
ggtgatgtta aattaacaca gtctatggcc atcatacgtt atatagctga caagcacaac 240ggtgatgtta aattaacaca gtctatggcc atcatacgtt atatagctga caagcacaac 240
atgttgggtg gttgtccaaa agagcgtgca gagatttcaa tgcttgaagg agcggttttg 300atgttgggtg gttgtccaaa agagcgtgca gagatttcaa tgcttgaagg agcggttttg 300
gatattagat acggtgtttc gagaattgca tatagtaaag actttgaaac tctcaaagtt 360gatattagat acggtgtttc gagaattgca tatagtaaag actttgaaac tctcaaagtt 360
gattttctta gcaagctacc tgaaatgctg aaaatgttcg aagatcgttt atgtcataaa 420gattttctta gcaagctacc tgaaatgctg aaaatgttcg aagatcgttt atgtcataaa 420
acatatttaa atggtgatca tgtaacccat cctgacttca tgttgtatga cgctcttgat 480acatatttaa atggtgatca tgtaacccat cctgacttca tgttgtatga cgctcttgat 480
gttgttttat acatggaccc aatgtgcctg gatgcgttcc caaaattagt ttgttttaaa 540gttgttttat acatggaccc aatgtgcctg gatgcgttcc caaaattagt ttgttttaaa 540
aaacgtattg aagctatccc acaaattgat aagtacttga aatccagcaa gtatatagca 600aaacgtattg aagctatccc acaaattgat aagtacttga aatccagcaa gtatatagca 600
tggcctttgc agggctggca agccacgttt ggtggtggcg accatcctcc aaaatcggat 660tggcctttgc agggctggca agccacgttt ggtggtggcg accatcctcc aaaatcggat 660
ctggaagttc tgttccaggg gcccctggga tccatgtggc gtccgtctga ctctaccgtt 720ctggaagttc tgttccaggg gcccctggga tccatgtggc gtccgtctga ctctaccgtt 720
tacgttccgc cgccgaaccc ggtttctaaa gttgttgcta ccgacgctta cgttacccgt 780tacgttccgc cgccgaaccc ggtttctaaa gttgttgcta ccgacgctta cgttacccgt 780
accaacatct tctaccacgc ttcttcttct cgtctgctgg ctgttggtca cccgtacttc 840accaacatct tctaccacgc ttcttcttct cgtctgctgg ctgttggtca cccgtacttc 840
tctatcaaac gtgctaacaa aaccgttgtt ccgaaagttt ctggttacca gtaccgtgtt 900tctatcaaac gtgctaacaa aaccgttgtt ccgaaagttt ctggttacca gtaccgtgtt 900
ttcaaagttg ttctgccgga cccgaacaaa ttcgctctgc cggactcttc tctgttcgac 960ttcaaagttg ttctgccgga cccgaacaaa ttcgctctgc cggactcttc tctgttcgac 960
ccgaccaccc agcgtctggt ttgggcttgc accggtctgg aagttggtcg tggtcagccg 1020ccgaccaccc agcgtctggt ttgggcttgc accggtctgg aagttggtcg tggtcagccg 1020
ctgggtgttg gtgtttctgg tcacccgttc ctgaacaaat acgacgacgt tgaaaactct 1080ctgggtgttg gtgtttctgg tcacccgttc ctgaacaaat acgacgacgt tgaaaactct 1080
ggttctggtg gtaacccggg tcaggacaac cgtgttaacg ttggtatgga ctacaaacag 1140ggttctggtg gtaacccggg tcaggacaac cgtgttaacg ttggtatgga ctacaaacag 1140
acccagctgt gcatggttgg ttgcgctccg ccgctgggtg aacactgggg taaaggtaaa 1200accccagctgt gcatggttgg ttgcgctccg ccgctgggtg aacactgggg taaaggtaaa 1200
cagtgcacca acaccccggt tcaggctggt gactgcccgc cgctggaact gatcacctct 1260cagtgcacca acaccccggt tcaggctggt gactgcccgc cgctggaact gatcacctct 1260
gttatccagg acggtgacat ggttgacacc ggtttcggtg ctatgaactt cgctgacctg 1320gttatccagg acggtgacat ggttgacacc ggtttcggtg ctatgaactt cgctgacctg 1320
cagaccaaca aatctgacgt tccgatcgac atctgcggta ccacctgcaa atacccggac 1380cagaccaaca aatctgacgt tccgatcgac atctgcggta ccacctgcaa atacccggac 1380
tacctgcaga tggctgctga cccgtacggt gaccgtctgt tcttcttcct gcgtaaagaa 1440tacctgcaga tggctgctga cccgtacggt gaccgtctgt tcttcttcct gcgtaaagaa 1440
cagatgttcg ctcgtcactt cttcaaccgt gctggtgaag ttggtgaacc ggttccggac 1500cagatgttcg ctcgtcactt cttcaaccgt gctggtgaag ttggtgaacc ggttccggac 1500
accctgatca tcaaaggttc tggtaaccgt acctctgttg gttcttctat ctacgttaac 1560accctgatca tcaaaggttc tggtaaccgt acctctgttg gttcttctat ctacgttaac 1560
accccgtctg gttctctggt ttcttctgaa gctcagctgt tcaacaaacc gtactggctg 1620accccgtctg gttctctggt ttcttctgaa gctcagctgt tcaacaaacc gtactggctg 1620
cagaaagctc agggtcacaa caacggtatc tgctggggta accagctgtt cgttaccgtt 1680cagaaagctc agggtcacaa caacggtatc tgctggggta accagctgtt cgttaccgtt 1680
gttgacacca cccgttctac caacatgacc ctgtgcgctt ctgttaccac ctcttctacc 1740gttgacacca cccgttctac caacatgacc ctgtgcgctt ctgttaccac ctcttctacc 1740
tacaccaact ctgactacaa agaatacatg cgtcacgttg aagaatacga cctgcagttc 1800tacaccaact ctgactacaa agaatacatg cgtcacgttg aagaatacga cctgcagttc 1800
atcttccagc tgtgctctat caccctgtct gctgaagtta tggcttacat ccacaccatg 1860atcttccagc tgtgctctat caccctgtct gctgaagtta tggcttacat ccacaccatg 1860
aacccgtctg ttctggaaga ctggaacttc ggtctgtctc cgccgccgaa cggtaccctg 1920aacccgtctg ttctggaaga ctggaacttc ggtctgtctc cgccgccgaa cggtaccctg 1920
gaagacacct accgttacgt tcagtctcag gctatcacct gccagaaacc gaccccggaa 1980gaagacacct accgttacgt tcagtctcag gctatcacct gccagaaacc gaccccggaa 1980
aaagaaaaac cggacccgta caaaaacctg tctttctggg aagttaacct gaaagaaaaa 2040aaagaaaaac cggacccgta caaaaacctg tctttctggg aagttaacct gaaagaaaaa 2040
ttctcttctg aactggacca gtacccgctg ggtcgtaaat tcctgctgca gtctggttac 2100ttctcttctg aactggacca gtacccgctg ggtcgtaaat tcctgctgca gtctggttac 2100
cgtggtcgtt cttctatccg taccggtgtt aaacgtccgg ctgtttctaa agcttctgct 2160cgtggtcgtt cttctatccg taccggtgtt aaacgtccgg ctgtttctaa agcttctgct 2160
gctccgaaac gtaaacgtgc taaaaccaaa cgt 2193gctccgaaac gtaaacgtgc taaaaccaaa cgt 2193
<210> 4<210> 4
<211> 2475<211> 2475
<212> DNA<212>DNA
<213> 人工序列<213> Artificial sequence
<400> 4<400> 4
atgtccccta tactaggtta ttggaaaatt aagggccttg tgcaacccac tcgacttctt 60atgtccccta tactaggtta ttggaaaatt aagggccttg tgcaacccac tcgacttctt 60
ttggaatatc ttgaagaaaa atatgaagag catttgtatg agcgcgatga aggtgataaa 120ttggaatatc ttgaagaaaa atatgaagag catttgtatg agcgcgatga aggtgataaa 120
tggcgaaaca aaaagtttga attgggtttg gagtttccca atcttcctta ttatattgat 180tggcgaaaca aaaagtttga attgggtttg gagtttccca atcttcctta ttatattgat 180
ggtgatgtta aattaacaca gtctatggcc atcatacgtt atatagctga caagcacaac 240ggtgatgtta aattaacaca gtctatggcc atcatacgtt atatagctga caagcacaac 240
atgttgggtg gttgtccaaa agagcgtgca gagatttcaa tgcttgaagg agcggttttg 300atgttgggtg gttgtccaaa agagcgtgca gagatttcaa tgcttgaagg agcggttttg 300
gatattagat acggtgtttc gagaattgca tatagtaaag actttgaaac tctcaaagtt 360gatattagat acggtgtttc gagaattgca tatagtaaag actttgaaac tctcaaagtt 360
gattttctta gcaagctacc tgaaatgctg aaaatgttcg aagatcgttt atgtcataaa 420gattttctta gcaagctacc tgaaatgctg aaaatgttcg aagatcgttt atgtcataaa 420
acatatttaa atggtgatca tgtaacccat cctgacttca tgttgtatga cgctcttgat 480acatatttaa atggtgatca tgtaacccat cctgacttca tgttgtatga cgctcttgat 480
gttgttttat acatggaccc aatgtgcctg gatgcgttcc caaaattagt ttgttttaaa 540gttgttttat acatggaccc aatgtgcctg gatgcgttcc caaaattagt ttgttttaaa 540
aaacgtattg aagctatccc acaaattgat aagtacttga aatccagcaa gtatatagca 600aaacgtattg aagctatccc acaaattgat aagtacttga aatccagcaa gtatatagca 600
tggcctttgc agggctggca agccacgttt ggtggtggcg accatcctcc aaaatcggat 660tggcctttgc agggctggca agccacgttt ggtggtggcg accatcctcc aaaatcggat 660
ctggaagttc tgttccaggg gccctctgac caggaagcta aaccgtctac cgaagacctg 720ctggaagttc tgttccaggg gccctctgac caggaagcta aaccgtctac cgaagacctg 720
ggtgacaaaa aagaaggtga atacatcaaa ctgaaagtta tcggtcagga ctcttctgaa 780ggtgacaaaa aagaaggtga atacatcaaa ctgaaagtta tcggtcagga ctcttctgaa 780
atccacttca aagttaaaat gaccacccac ctgaaaaaac tgaaagaatc ttactgccag 840atccacttca aagttaaaat gaccaccac ctgaaaaaac tgaaagaatc ttactgccag 840
cgtcagggtg ttccgatgaa ctctctgcgt ttcctgttcg aaggtcagcg tatcgctgac 900cgtcagggtg ttccgatgaa ctctctgcgt ttcctgttcg aaggtcagcg tatcgctgac 900
aaccacaccc cgaaagaact gggtatggaa gaagaagacg ttatcgaagt ttaccaggaa 960aaccaacaccc cgaaagaact gggtatggaa gaagaagacg ttatcgaagt ttaccaggaa 960
cagaccggtg gatccatgtg gcgtccgtct gactctaccg tttacgttcc gccgccgaac 1020cagaccggtg gatccatgtg gcgtccgtct gactctaccg tttacgttcc gccgccgaac 1020
ccggtttcta aagttgttgc taccgacgct tacgttaccc gtaccaacat cttctaccac 1080ccggtttcta aagttgttgc taccgacgct tacgttaccc gtaccaacat cttctaccac 1080
gcttcttctt ctcgtctgct ggctgttggt cacccgtact tctctatcaa acgtgctaac 1140gcttcttctt ctcgtctgct ggctgttggt cacccgtact tctctatcaa acgtgctaac 1140
aaaaccgttg ttccgaaagt ttctggttac cagtaccgtg ttttcaaagt tgttctgccg 1200aaaaccgttg ttccgaaagt ttctggttac cagtaccgtg ttttcaaagt tgttctgccg 1200
gacccgaaca aattcgctct gccggactct tctctgttcg acccgaccac ccagcgtctg 1260gacccgaaca aattcgctct gccggactct tctctgttcg acccgaccac ccagcgtctg 1260
gtttgggctt gcaccggtct ggaagttggt cgtggtcagc cgctgggtgt tggtgtttct 1320gtttgggctt gcaccggtct ggaagttggt cgtggtcagc cgctgggtgt tggtgtttct 1320
ggtcacccgt tcctgaacaa atacgacgac gttgaaaact ctggttctgg tggtaacccg 1380ggtcacccgt tcctgaacaa atacgacgac gttgaaaact ctggttctgg tggtaacccg 1380
ggtcaggaca accgtgttaa cgttggtatg gactacaaac agacccagct gtgcatggtt 1440ggtcaggaca accgtgttaa cgttggtatg gactacaaac agacccagct gtgcatggtt 1440
ggttgcgctc cgccgctggg tgaacactgg ggtaaaggta aacagtgcac caacaccccg 1500ggttgcgctc cgccgctggg tgaacactgg ggtaaaggta aacagtgcac caacacccccg 1500
gttcaggctg gtgactgccc gccgctggaa ctgatcacct ctgttatcca ggacggtgac 1560gttcaggctg gtgactgccc gccgctggaa ctgatcacct ctgttatcca ggacggtgac 1560
atggttgaca ccggtttcgg tgctatgaac ttcgctgacc tgcagaccaa caaatctgac 1620atggttgaca ccggtttcgg tgctatgaac ttcgctgacc tgcagaccaa caaatctgac 1620
gttccgatcg acatctgcgg taccacctgc aaatacccgg actacctgca gatggctgct 1680gttccgatcg acatctgcgg taccacctgc aaatacccgg actacctgca gatggctgct 1680
gacccgtacg gtgaccgtct gttcttcttc ctgcgtaaag aacagatgtt cgctcgtcac 1740gacccgtacg gtgaccgtct gttcttcttc ctgcgtaaag aacagatgtt cgctcgtcac 1740
ttcttcaacc gtgctggtga agttggtgaa ccggttccgg acaccctgat catcaaaggt 1800ttcttcaacc gtgctggtga agttggtgaa ccggttccgg acaccctgat catcaaaggt 1800
tctggtaacc gtacctctgt tggttcttct atctacgtta acaccccgtc tggttctctg 1860tctggtaacc gtacctctgt tggttcttct atctacgtta acaccccgtc tggttctctg 1860
gtttcttctg aagctcagct gttcaacaaa ccgtactggc tgcagaaagc tcagggtcac 1920gtttcttctg aagctcagct gttcaacaaa ccgtactggc tgcagaaagc tcagggtcac 1920
aacaacggta tctgctgggg taaccagctg ttcgttaccg ttgttgacac cacccgttct 1980aacaacggta tctgctgggg taaccagctg ttcgttaccg ttgttgacac cacccgttct 1980
accaacatga ccctgtgcgc ttctgttacc acctcttcta cctacaccaa ctctgactac 2040accaacatga ccctgtgcgc ttctgttacc acctcttcta cctacaccaa ctctgactac 2040
aaagaataca tgcgtcacgt tgaagaatac gacctgcagt tcatcttcca gctgtgctct 2100aaagaataca tgcgtcacgt tgaagaatac gacctgcagt tcatcttcca gctgtgctct 2100
atcaccctgt ctgctgaagt tatggcttac atccacacca tgaacccgtc tgttctggaa 2160atcaccctgt ctgctgaagt tatggcttac atccacacca tgaacccgtc tgttctggaa 2160
gactggaact tcggtctgtc tccgccgccg aacggtaccc tggaagacac ctaccgttac 2220gactggaact tcggtctgtc tccgccgccg aacggtaccc tggaagacac ctaccgttac 2220
gttcagtctc aggctatcac ctgccagaaa ccgaccccgg aaaaagaaaa accggacccg 2280gttcagtctc aggctatcac ctgccagaaa ccgaccccgg aaaaagaaaa accggacccg 2280
tacaaaaacc tgtctttctg ggaagttaac ctgaaagaaa aattctcttc tgaactggac 2340tacaaaaacc tgtctttctg ggaagttaac ctgaaagaaa aattctcttc tgaactggac 2340
cagtacccgc tgggtcgtaa attcctgctg cagtctggtt accgtggtcg ttcttctatc 2400cagtacccgc tgggtcgtaa attcctgctg cagtctggtt accgtggtcg ttcttctatc 2400
cgtaccggtg ttaaacgtcc ggctgtttct aaagcttctg ctgctccgaa acgtaaacgt 2460cgtaccggtg ttaaacgtcc ggctgtttct aaagcttctg ctgctccgaa acgtaaacgt 2460
gctaaaacca aacgt 2475gctaaaacca aacgt 2475
<210> 5<210> 5
<211> 2673<211> 2673
<212> DNA<212>DNA
<213> 人工序列<213> Artificial sequence
<400> 5<400> 5
atgaaaatcg aagaaggtaa actggtaatc tggattaacg gcgataaagg ctataacggt 60atgaaaatcg aagaaggtaa actggtaatc tggattaacg gcgataaagg ctataacggt 60
ctcgctgaag tcggtaagaa attcgagaaa gataccggaa ttaaagtcac cgttgagcat 120ctcgctgaag tcggtaagaa attcgagaaa gataccggaa ttaaagtcac cgttgagcat 120
ccggataaac tggaagagaa attcccacag gttgcggcaa ctggcgatgg ccctgacatt 180ccggataaac tggaagagaa attcccacag gttgcggcaa ctggcgatgg ccctgacatt 180
atcttctggg cacacgaccg ctttggtggc tacgctcaat ctggcctgtt ggctgaaatc 240atcttctggg cacacgaccg ctttggtggc tacgctcaat ctggcctgtt ggctgaaatc 240
accccggaca aagcgttcca ggacaagctg tatccgttta cctgggatgc cgtacgttac 300accccggaca aagcgttcca ggacaagctg tatccgttta cctgggatgc cgtacgttac 300
aacggcaagc tgattgctta cccgatcgct gttgaagcgt tatcgctgat ttataacaaa 360aacggcaagc tgattgctta cccgatcgct gttgaagcgt tatcgctgat ttataacaaa 360
gatctgctgc cgaacccgcc aaaaacctgg gaagagatcc cggcgctgga taaagaactg 420gatctgctgc cgaacccgcc aaaaacctgg gaagagatcc cggcgctgga taaagaactg 420
aaagcgaaag gtaagagcgc gctgatgttc aacctgcaag aaccgtactt cacctggccg 480aaagcgaaag gtaagagcgc gctgatgttc aacctgcaag aaccgtactt cacctggccg 480
ctgattgctg ctgacggggg ttatgcgttc aagtatgaaa acggcaagta cgacattaaa 540ctgattgctg ctgacgggggg ttatgcgttc aagtatgaaa acggcaagta cgacattaaa 540
gacgtgggcg tggataacgc tggcgcgaaa gcgggtctga ccttcctggt tgacctgatt 600gacgtgggcg tggataacgc tggcgcgaaa gcgggtctga ccttcctggt tgacctgatt 600
aaaaacaaac acatgaatgc agacaccgat tactccatcg cagaagctgc ctttaataaa 660aaaaacaaac acatgaatgc agacaccgat tactccatcg cagaagctgc ctttaataaa 660
ggcgaaacag cgatgaccat caacggcccg tgggcatggt ccaacatcga caccagcaaa 720ggcgaaacag cgatgaccat caacggcccg tgggcatggt ccaacatcga caccagcaaa 720
gtgaattatg gtgtaacggt actgccgacc ttcaagggtc aaccatccaa accgttcgtt 780gtgaattatg gtgtaacggt actgccgacc ttcaagggtc aaccatccaa accgttcgtt 780
ggcgtgctga gcgcaggtat taacgccgcc agtccgaaca aagagctggc aaaagagttc 840ggcgtgctga gcgcaggtat taacgccgcc agtccgaaca aagagctggc aaaagagttc 840
ctcgaaaact atctgctgac tgatgaaggt ctggaagcgg ttaataaaga caaaccgctg 900ctcgaaaact atctgctgac tgatgaaggt ctggaagcgg ttaataaaga caaaccgctg 900
ggtgccgtag cgctgaagtc ttacgaggaa gagttggcga aagatccacg tattgccgcc 960ggtgccgtag cgctgaagtc ttacgaggaa gagttggcga aagatccacg tattgccgcc 960
actatggaaa acgcccagaa aggtgaaatc atgccgaaca tcccgcagat gtccgctttc 1020actatggaaa acgcccagaa aggtgaaatc atgccgaaca tcccgcagat gtccgctttc 1020
tggtatgccg tgcgtactgc ggtgatcaac gccgccagcg gtcgtcagac tgtcgatgaa 1080tggtatgccg tgcgtactgc ggtgatcaac gccgccagcg gtcgtcagac tgtcgatgaa 1080
gccctgaaag acgcgcagac taattcgagc tcgaacaaca acaacaataa caataacaac 1140gccctgaaag acgcgcagac taattcgagc tcgaacaaca acaacaataa caataacaac 1140
aacctcggga tcgagggaag gatttcagaa ttcatgtggc gtccgtctga ctctaccgtt 1200aacctcggga tcgagggaag gatttcagaa ttcatgtggc gtccgtctga ctctaccgtt 1200
tacgttccgc cgccgaaccc ggtttctaaa gttgttgcta ccgacgctta cgttacccgt 1260tacgttccgc cgccgaaccc ggtttctaaa gttgttgcta ccgacgctta cgttacccgt 1260
accaacatct tctaccacgc ttcttcttct cgtctgctgg ctgttggtca cccgtacttc 1320accaacatct tctaccacgc ttcttcttct cgtctgctgg ctgttggtca cccgtacttc 1320
tctatcaaac gtgctaacaa aaccgttgtt ccgaaagttt ctggttacca gtaccgtgtt 1380tctatcaaac gtgctaacaa aaccgttgtt ccgaaagttt ctggttacca gtaccgtgtt 1380
ttcaaagttg ttctgccgga cccgaacaaa ttcgctctgc cggactcttc tctgttcgac 1440ttcaaagttg ttctgccgga cccgaacaaa ttcgctctgc cggactcttc tctgttcgac 1440
ccgaccaccc agcgtctggt ttgggcttgc accggtctgg aagttggtcg tggtcagccg 1500ccgaccacccc agcgtctggt ttgggcttgc accggtctgg aagttggtcg tggtcagccg 1500
ctgggtgttg gtgtttctgg tcacccgttc ctgaacaaat acgacgacgt tgaaaactct 1560ctgggtgttg gtgtttctgg tcacccgttc ctgaacaaat acgacgacgt tgaaaactct 1560
ggttctggtg gtaacccggg tcaggacaac cgtgttaacg ttggtatgga ctacaaacag 1620ggttctggtg gtaacccggg tcaggacaac cgtgttaacg ttggtatgga ctacaaacag 1620
acccagctgt gcatggttgg ttgcgctccg ccgctgggtg aacactgggg taaaggtaaa 1680accccagctgt gcatggttgg ttgcgctccg ccgctgggtg aacactgggg taaaggtaaa 1680
cagtgcacca acaccccggt tcaggctggt gactgcccgc cgctggaact gatcacctct 1740cagtgcacca acaccccggt tcaggctggt gactgcccgc cgctggaact gatcacctct 1740
gttatccagg acggtgacat ggttgacacc ggtttcggtg ctatgaactt cgctgacctg 1800gttatccagg acggtgacat ggttgacacc ggtttcggtg ctatgaactt cgctgacctg 1800
cagaccaaca aatctgacgt tccgatcgac atctgcggta ccacctgcaa atacccggac 1860cagaccaaca aatctgacgt tccgatcgac atctgcggta ccacctgcaa atacccggac 1860
tacctgcaga tggctgctga cccgtacggt gaccgtctgt tcttcttcct gcgtaaagaa 1920tacctgcaga tggctgctga cccgtacggt gaccgtctgt tcttcttcct gcgtaaagaa 1920
cagatgttcg ctcgtcactt cttcaaccgt gctggtgaag ttggtgaacc ggttccggac 1980cagatgttcg ctcgtcactt cttcaaccgt gctggtgaag ttggtgaacc ggttccggac 1980
accctgatca tcaaaggttc tggtaaccgt acctctgttg gttcttctat ctacgttaac 2040accctgatca tcaaaggttc tggtaaccgt acctctgttg gttcttctat ctacgttaac 2040
accccgtctg gttctctggt ttcttctgaa gctcagctgt tcaacaaacc gtactggctg 2100accccgtctg gttctctggt ttcttctgaa gctcagctgt tcaacaaacc gtactggctg 2100
cagaaagctc agggtcacaa caacggtatc tgctggggta accagctgtt cgttaccgtt 2160cagaaagctc agggtcacaa caacggtatc tgctggggta accagctgtt cgttaccgtt 2160
gttgacacca cccgttctac caacatgacc ctgtgcgctt ctgttaccac ctcttctacc 2220gttgacacca cccgttctac caacatgacc ctgtgcgctt ctgttaccac ctcttctacc 2220
tacaccaact ctgactacaa agaatacatg cgtcacgttg aagaatacga cctgcagttc 2280tacaccaact ctgactacaa agaatacatg cgtcacgttg aagaatacga cctgcagttc 2280
atcttccagc tgtgctctat caccctgtct gctgaagtta tggcttacat ccacaccatg 2340atcttccagc tgtgctctat caccctgtct gctgaagtta tggcttacat ccacaccatg 2340
aacccgtctg ttctggaaga ctggaacttc ggtctgtctc cgccgccgaa cggtaccctg 2400aacccgtctg ttctggaaga ctggaacttc ggtctgtctc cgccgccgaa cggtaccctg 2400
gaagacacct accgttacgt tcagtctcag gctatcacct gccagaaacc gaccccggaa 2460gaagacacct accgttacgt tcagtctcag gctatcacct gccagaaacc gaccccggaa 2460
aaagaaaaac cggacccgta caaaaacctg tctttctggg aagttaacct gaaagaaaaa 2520aaagaaaaac cggacccgta caaaaacctg tctttctggg aagttaacct gaaagaaaaa 2520
ttctcttctg aactggacca gtacccgctg ggtcgtaaat tcctgctgca gtctggttac 2580ttctcttctg aactggacca gtacccgctg ggtcgtaaat tcctgctgca gtctggttac 2580
cgtggtcgtt cttctatccg taccggtgtt aaacgtccgg ctgtttctaa agcttctgct 2640cgtggtcgtt cttctatccg taccggtgtt aaacgtccgg ctgtttctaa agcttctgct 2640
gctccgaaac gtaaacgtgc taaaaccaaa cgt 2673gctccgaaac gtaaacgtgc taaaaccaaa cgt 2673
<210> 6<210> 6
<211> 1563<211> 1563
<212> DNA<212>DNA
<213> 人工序列<213> Artificial sequence
<400> 6<400> 6
atgggcagca gccatcatca tcatcatcac agcagcggcc tggtgccgcg cggcagccat 60atgggcagca gccatcatca tcatcatcac agcagcggcc tggtgccgcg cggcagccat 60
atgatgtggc gtccgtctga ctctaccgtt tacgttccgc cgccgaaccc ggtttctaaa 120atgatgtggc gtccgtctga ctctaccgtt tacgttccgc cgccgaaccc ggtttctaaa 120
gttgttgcta ccgacgctta cgttacccgt accaacatct tctaccacgc ttcttcttct 180gttgttgcta ccgacgctta cgttacccgt accaacatct tctaccacgc ttcttcttct 180
cgtctgctgg ctgttggtca cccgtacttc tctatcaaac gtgctaacaa aaccgttgtt 240cgtctgctgg ctgttggtca cccgtacttc tctatcaaac gtgctaacaa aaccgttgtt 240
ccgaaagttt ctggttacca gtaccgtgtt ttcaaagttg ttctgccgga cccgaacaaa 300ccgaaagttt ctggttacca gtaccgtgtt ttcaaagttg ttctgccgga cccgaacaaa 300
ttcgctctgc cggactcttc tctgttcgac ccgaccaccc agcgtctggt ttgggcttgc 360ttcgctctgc cggactcttc tctgttcgac ccgaccaccc agcgtctggt ttgggcttgc 360
accggtctgg aagttggtcg tggtcagccg ctgggtgttg gtgtttctgg tcacccgttc 420accggtctgg aagttggtcg tggtcagccg ctgggtgttg gtgtttctgg tcacccgttc 420
ctgaacaaat acgacgacgt tgaaaactct ggttctggtg gtaacccggg tcaggacaac 480ctgaacaaat acgacgacgt tgaaaactct ggttctggtg gtaacccggg tcaggacaac 480
cgtgttaacg ttggtatgga ctacaaacag acccagctgt gcatggttgg ttgcgctccg 540cgtgttaacg ttggtatgga ctacaaacag acccagctgt gcatggttgg ttgcgctccg 540
ccgctgggtg aacactgggg taaaggtaaa cagtgcacca acaccccggt tcaggctggt 600ccgctgggtg aacactgggg taaaggtaaa cagtgcacca acaccccggt tcaggctggt 600
gactgcccgc cgctggaact gatcacctct gttatccagg acggtgacat ggttgacacc 660gactgcccgc cgctggaact gatcacctct gttatccagg acggtgacat ggttgacacc 660
ggtttcggtg ctatgaactt cgctgacctg cagaccaaca aatctgacgt tccgatcgac 720ggtttcggtg ctatgaactt cgctgacctg cagaccaaca aatctgacgt tccgatcgac 720
atctgcggta ccacctgcaa atacccggac tacctgcaga tggctgctga cccgtacggt 780atctgcggta ccacctgcaa atacccggac tacctgcaga tggctgctga cccgtacggt 780
gaccgtctgt tcttcttcct gcgtaaagaa cagatgttcg ctcgtcactt cttcaaccgt 840gaccgtctgt tcttcttcct gcgtaaagaa cagatgttcg ctcgtcactt cttcaaccgt 840
gctggtgaag ttggtgaacc ggttccggac accctgatca tcaaaggttc tggtaaccgt 900gctggtgaag ttggtgaacc ggttccggac accctgatca tcaaaggttc tggtaaccgt 900
acctctgttg gttcttctat ctacgttaac accccgtctg gttctctggt ttcttctgaa 960acctctgttg gttcttctat ctacgttaac accccgtctg gttctctggt ttcttctgaa 960
gctcagctgt tcaacaaacc gtactggctg cagaaagctc agggtcacaa caacggtatc 1020gctcagctgt tcaacaaacc gtactggctg cagaaagctc agggtcacaa caacggtatc 1020
tgctggggta accagctgtt cgttaccgtt gttgacacca cccgttctac caacatgacc 1080tgctggggta accagctgtt cgttaccgtt gttgacacca cccgttctac caacatgacc 1080
ctgtgcgctt ctgttaccac ctcttctacc tacaccaact ctgactacaa agaatacatg 1140ctgtgcgctt ctgttaccac ctcttctacc tacaccaact ctgactacaa agaatacatg 1140
cgtcacgttg aagaatacga cctgcagttc atcttccagc tgtgctctat caccctgtct 1200cgtcacgttg aagaatacga cctgcagttc atcttccagc tgtgctctat caccctgtct 1200
gctgaagtta tggcttacat ccacaccatg aacccgtctg ttctggaaga ctggaacttc 1260gctgaagtta tggcttacat ccacaccatg aacccgtctg ttctggaaga ctggaacttc 1260
ggtctgtctc cgccgccgaa cggtaccctg gaagacacct accgttacgt tcagtctcag 1320ggtctgtctc cgccgccgaa cggtaccctg gaagaacacct accgttacgt tcagtctcag 1320
gctatcacct gccagaaacc gaccccggaa aaagaaaaac cggacccgta caaaaacctg 1380gctatcacct gccagaaacc gaccccggaa aaagaaaaac cggacccgta caaaaacctg 1380
tctttctggg aagttaacct gaaagaaaaa ttctcttctg aactggacca gtacccgctg 1440tctttctggg aagttaacct gaaagaaaaa ttctcttctg aactggacca gtacccgctg 1440
ggtcgtaaat tcctgctgca gtctggttac cgtggtcgtt cttctatccg taccggtgtt 1500ggtcgtaaat tcctgctgca gtctggttac cgtggtcgtt cttctatccg taccggtgtt 1500
aaacgtccgg ctgtttctaa agcttctgct gctccgaaac gtaaacgtgc taaaaccaaa 1560aaacgtccgg ctgtttctaa agcttctgct gctccgaaac gtaaacgtgc taaaaccaaa 1560
cgt 1563cgt 1563
<210> 7<210> 7
<211> 1854<211> 1854
<212> DNA<212>DNA
<213> 人工序列<213> Artificial sequence
<400> 7<400> 7
atgggcagca gccatcatca tcatcatcac agcagcggcc tggtgccgcg cggcagccat 60atgggcagca gccatcatca tcatcatcac agcagcggcc tggtgccgcg cggcagccat 60
atgtctgacc aggaagctaa accgtctacc gaagacctgg gtgacaaaaa agaaggtgaa 120atgtctgacc aggaagctaa accgtctacc gaagacctgg gtgacaaaaa agaaggtgaa 120
tacatcaaac tgaaagttat cggtcaggac tcttctgaaa tccacttcaa agttaaaatg 180tacatcaaac tgaaagttat cggtcaggac tcttctgaaa tccacttcaa agttaaaatg 180
accacccacc tgaaaaaact gaaagaatct tactgccagc gtcagggtgt tccgatgaac 240accacccacc tgaaaaaact gaaagaatct tactgccagc gtcagggtgttccgatgaac 240
tctctgcgtt tcctgttcga aggtcagcgt atcgctgaca accacacccc gaaagaactg 300tctctgcgtt tcctgttcga aggtcagcgt atcgctgaca accacaccccc gaaagaactg 300
ggtatggaag aagaagacgt tatcgaagtt taccaggaac agaccggtgg atccatgtgg 360ggtatggaag aagaagacgt tatcgaagtt taccaggaac agaccggtgg atccatgtgg 360
cgtccgtctg actctaccgt ttacgttccg ccgccgaacc cggtttctaa agttgttgct 420cgtccgtctg actctaccgt ttacgttccg ccgccgaacc cggtttctaa agttgttgct 420
accgacgctt acgttacccg taccaacatc ttctaccacg cttcttcttc tcgtctgctg 480accgacgctt acgttacccg taccaacatc ttctaccacg cttcttcttc tcgtctgctg 480
gctgttggtc acccgtactt ctctatcaaa cgtgctaaca aaaccgttgt tccgaaagtt 540gctgttggtc acccgtactt ctctatcaaa cgtgctaaca aaaccgttgt tccgaaagtt 540
tctggttacc agtaccgtgt tttcaaagtt gttctgccgg acccgaacaa attcgctctg 600tctggttacc agtaccgtgttttcaaagtt gttctgccgg acccgaacaa attcgctctg 600
ccggactctt ctctgttcga cccgaccacc cagcgtctgg tttgggcttg caccggtctg 660ccggactctt ctctgttcga cccgaccacc cagcgtctgg tttgggcttg caccggtctg 660
gaagttggtc gtggtcagcc gctgggtgtt ggtgtttctg gtcacccgtt cctgaacaaa 720gaagttggtc gtggtcagcc gctgggtgtt ggtgtttctg gtcacccgtt cctgaacaaa 720
tacgacgacg ttgaaaactc tggttctggt ggtaacccgg gtcaggacaa ccgtgttaac 780tacgacgacg ttgaaaactc tggttctggt ggtaacccgg gtcaggaca ccgtgttaac 780
gttggtatgg actacaaaca gacccagctg tgcatggttg gttgcgctcc gccgctgggt 840gttggtatgg actacaaaca gacccagctg tgcatggttg gttgcgctcc gccgctgggt 840
gaacactggg gtaaaggtaa acagtgcacc aacaccccgg ttcaggctgg tgactgcccg 900gaacactggg gtaaaggtaa acagtgcacc aacacccccgg ttcaggctgg tgactgcccg 900
ccgctggaac tgatcacctc tgttatccag gacggtgaca tggttgacac cggtttcggt 960ccgctggaac tgatcacctc tgttatccag gacggtgaca tggttgacac cggtttcggt 960
gctatgaact tcgctgacct gcagaccaac aaatctgacg ttccgatcga catctgcggt 1020gctatgaact tcgctgacct gcagaccaac aaatctgacg ttccgatcga catctgcggt 1020
accacctgca aatacccgga ctacctgcag atggctgctg acccgtacgg tgaccgtctg 1080accacctgca aatacccgga ctacctgcag atggctgctg accccgtacgg tgaccgtctg 1080
ttcttcttcc tgcgtaaaga acagatgttc gctcgtcact tcttcaaccg tgctggtgaa 1140ttcttcttcc tgcgtaaaga acagatgttc gctcgtcact tcttcaaccg tgctggtgaa 1140
gttggtgaac cggttccgga caccctgatc atcaaaggtt ctggtaaccg tacctctgtt 1200gttggtgaac cggttccgga caccctgatc atcaaaggtt ctggtaaccg taccctctgtt 1200
ggttcttcta tctacgttaa caccccgtct ggttctctgg tttcttctga agctcagctg 1260ggttcttcta tctacgttaa caccccgtct ggttctctgg tttcttctga agctcagctg 1260
ttcaacaaac cgtactggct gcagaaagct cagggtcaca acaacggtat ctgctggggt 1320ttcaacaaac cgtactggct gcagaaagct cagggtcaca acaacggtat ctgctggggt 1320
aaccagctgt tcgttaccgt tgttgacacc acccgttcta ccaacatgac cctgtgcgct 1380aaccagctgt tcgttaccgt tgttgacacc acccgttcta ccaacatgac cctgtgcgct 1380
tctgttacca cctcttctac ctacaccaac tctgactaca aagaatacat gcgtcacgtt 1440tctgttacca cctcttctac cctaccaac tctgactaca aagaatacat gcgtcacgtt 1440
gaagaatacg acctgcagtt catcttccag ctgtgctcta tcaccctgtc tgctgaagtt 1500gaagaatacg acctgcagtt catcttccag ctgtgctcta tcaccctgtc tgctgaagtt 1500
atggcttaca tccacaccat gaacccgtct gttctggaag actggaactt cggtctgtct 1560atggcttaca tccacaccat gaacccgtct gttctggaag actggaactt cggtctgtct 1560
ccgccgccga acggtaccct ggaagacacc taccgttacg ttcagtctca ggctatcacc 1620ccgccgccga acggtaccct ggaagacacc taccgttacg ttcagtctca ggctatcacc 1620
tgccagaaac cgaccccgga aaaagaaaaa ccggacccgt acaaaaacct gtctttctgg 1680tgccagaaac cgaccccgga aaaagaaaaa ccggacccgt acaaaaacct gtctttctgg 1680
gaagttaacc tgaaagaaaa attctcttct gaactggacc agtacccgct gggtcgtaaa 1740gaagttaacc tgaaagaaaa attctcttct gaactggacc agtacccgct gggtcgtaaa 1740
ttcctgctgc agtctggtta ccgtggtcgt tcttctatcc gtaccggtgt taaacgtccg 1800ttcctgctgc agtctggtta ccgtggtcgt tcttctatcc gtaccggtgt taaacgtccg 1800
gctgtttcta aagcttctgc tgctccgaaa cgtaaacgtg ctaaaaccaa acgt 1854gctgtttcta aagcttctgc tgctccgaaa cgtaaacgtg ctaaaaccaa acgt 1854
<210> 8<210> 8
<211> 731<211> 731
<212> PRT<212> PRT
<213> 人工序列<213> Artificial sequence
<400> 8<400> 8
Met Ser Pro Ile Leu Gly Tyr Trp Lys Ile Lys Gly Leu Val Gln ProMet Ser Pro Ile Leu Gly Tyr Trp Lys Ile Lys Gly Leu Val Gln Pro
1 5 10 151 5 10 15
Thr Arg Leu Leu Leu Glu Tyr Leu Glu Glu Lys Tyr Glu Glu His LeuThr Arg Leu Leu Leu Glu Tyr Leu Glu Glu Lys Tyr Glu Glu His Leu
20 25 30 20 25 30
Tyr Glu Arg Asp Glu Gly Asp Lys Trp Arg Asn Lys Lys Phe Glu LeuTyr Glu Arg Asp Glu Gly Asp Lys Trp Arg Asn Lys Lys Phe Glu Leu
35 40 45 35 40 45
Gly Leu Glu Phe Pro Asn Leu Pro Tyr Tyr Ile Asp Gly Asp Val LysGly Leu Glu Phe Pro Asn Leu Pro Tyr Tyr Ile Asp Gly Asp Val Lys
50 55 60 50 55 60
Leu Thr Gln Ser Met Ala Ile Ile Arg Tyr Ile Ala Asp Lys His AsnLeu Thr Gln Ser Met Ala Ile Ile Arg Tyr Ile Ala Asp Lys His Asn
65 70 75 8065 70 75 80
Met Leu Gly Gly Cys Pro Lys Glu Arg Ala Glu Ile Ser Met Leu GluMet Leu Gly Gly Cys Pro Lys Glu Arg Ala Glu Ile Ser Met Leu Glu
85 90 95 85 90 95
Gly Ala Val Leu Asp Ile Arg Tyr Gly Val Ser Arg Ile Ala Tyr SerGly Ala Val Leu Asp Ile Arg Tyr Gly Val Ser Arg Ile Ala Tyr Ser
100 105 110 100 105 110
Lys Asp Phe Glu Thr Leu Lys Val Asp Phe Leu Ser Lys Leu Pro GluLys Asp Phe Glu Thr Leu Lys Val Asp Phe Leu Ser Lys Leu Pro Glu
115 120 125 115 120 125
Met Leu Lys Met Phe Glu Asp Arg Leu Cys His Lys Thr Tyr Leu AsnMet Leu Lys Met Phe Glu Asp Arg Leu Cys His Lys Thr Tyr Leu Asn
130 135 140 130 135 140
Gly Asp His Val Thr His Pro Asp Phe Met Leu Tyr Asp Ala Leu AspGly Asp His Val Thr His Pro Asp Phe Met Leu Tyr Asp Ala Leu Asp
145 150 155 160145 150 155 160
Val Val Leu Tyr Met Asp Pro Met Cys Leu Asp Ala Phe Pro Lys LeuVal Val Leu Tyr Met Asp Pro Met Cys Leu Asp Ala Phe Pro Lys Leu
165 170 175 165 170 175
Val Cys Phe Lys Lys Arg Ile Glu Ala Ile Pro Gln Ile Asp Lys TyrVal Cys Phe Lys Lys Arg Ile Glu Ala Ile Pro Gln Ile Asp Lys Tyr
180 185 190 180 185 190
Leu Lys Ser Ser Lys Tyr Ile Ala Trp Pro Leu Gln Gly Trp Gln AlaLeu Lys Ser Ser Lys Tyr Ile Ala Trp Pro Leu Gln Gly Trp Gln Ala
195 200 205 195 200 205
Thr Phe Gly Gly Gly Asp His Pro Pro Lys Ser Asp Leu Glu Val LeuThr Phe Gly Gly Gly Asp His Pro Pro Lys Ser Asp Leu Glu Val Leu
210 215 220 210 215 220
Phe Gln Gly Pro Leu Gly Ser Met Trp Arg Pro Ser Asp Ser Thr ValPhe Gln Gly Pro Leu Gly Ser Met Trp Arg Pro Ser Asp Ser Thr Val
225 230 235 240225 230 235 240
Tyr Val Pro Pro Pro Asn Pro Val Ser Lys Val Val Ala Thr Asp AlaTyr Val Pro Pro Pro Asn Pro Val Ser Lys Val Val Ala Thr Asp Ala
245 250 255 245 250 255
Tyr Val Thr Arg Thr Asn Ile Phe Tyr His Ala Ser Ser Ser Arg LeuTyr Val Thr Arg Thr Asn Ile Phe Tyr His Ala Ser Ser Ser Arg Leu
260 265 270 260 265 270
Leu Ala Val Gly His Pro Tyr Phe Ser Ile Lys Arg Ala Asn Lys ThrLeu Ala Val Gly His Pro Tyr Phe Ser Ile Lys Arg Ala Asn Lys Thr
275 280 285 275 280 285
Val Val Pro Lys Val Ser Gly Tyr Gln Tyr Arg Val Phe Lys Val ValVal Val Pro Lys Val Ser Gly Tyr Gln Tyr Arg Val Phe Lys Val Val
290 295 300 290 295 300
Leu Pro Asp Pro Asn Lys Phe Ala Leu Pro Asp Ser Ser Leu Phe AspLeu Pro Asp Pro Asn Lys Phe Ala Leu Pro Asp Ser Ser Leu Phe Asp
305 310 315 320305 310 315 320
Pro Thr Thr Gln Arg Leu Val Trp Ala Cys Thr Gly Leu Glu Val GlyPro Thr Thr Gln Arg Leu Val Trp Ala Cys Thr Gly Leu Glu Val Gly
325 330 335 325 330 335
Arg Gly Gln Pro Leu Gly Val Gly Val Ser Gly His Pro Phe Leu AsnArg Gly Gln Pro Leu Gly Val Gly Val Ser Gly His Pro Phe Leu Asn
340 345 350 340 345 350
Lys Tyr Asp Asp Val Glu Asn Ser Gly Ser Gly Gly Asn Pro Gly GlnLys Tyr Asp Asp Val Glu Asn Ser Gly Ser Gly Gly Asn Pro Gly Gln
355 360 365 355 360 365
Asp Asn Arg Val Asn Val Gly Met Asp Tyr Lys Gln Thr Gln Leu CysAsp Asn Arg Val Asn Val Gly Met Asp Tyr Lys Gln Thr Gln Leu Cys
370 375 380 370 375 380
Met Val Gly Cys Ala Pro Pro Leu Gly Glu His Trp Gly Lys Gly LysMet Val Gly Cys Ala Pro Pro Leu Gly Glu His Trp Gly Lys Gly Lys
385 390 395 400385 390 395 400
Gln Cys Thr Asn Thr Pro Val Gln Ala Gly Asp Cys Pro Pro Leu GluGln Cys Thr Asn Thr Pro Val Gln Ala Gly Asp Cys Pro Pro Leu Glu
405 410 415 405 410 415
Leu Ile Thr Ser Val Ile Gln Asp Gly Asp Met Val Asp Thr Gly PheLeu Ile Thr Ser Val Ile Gln Asp Gly Asp Met Val Asp Thr Gly Phe
420 425 430 420 425 430
Gly Ala Met Asn Phe Ala Asp Leu Gln Thr Asn Lys Ser Asp Val ProGly Ala Met Asn Phe Ala Asp Leu Gln Thr Asn Lys Ser Asp Val Pro
435 440 445 435 440 445
Ile Asp Ile Cys Gly Thr Thr Cys Lys Tyr Pro Asp Tyr Leu Gln MetIle Asp Ile Cys Gly Thr Thr Cys Lys Tyr Pro Asp Tyr Leu Gln Met
450 455 460 450 455 460
Ala Ala Asp Pro Tyr Gly Asp Arg Leu Phe Phe Phe Leu Arg Lys GluAla Ala Asp Pro Tyr Gly Asp Arg Leu Phe Phe Phe Leu Arg Lys Glu
465 470 475 480465 470 475 480
Gln Met Phe Ala Arg His Phe Phe Asn Arg Ala Gly Glu Val Gly GluGln Met Phe Ala Arg His Phe Phe Asn Arg Ala Gly Glu Val Gly Glu
485 490 495 485 490 495
Pro Val Pro Asp Thr Leu Ile Ile Lys Gly Ser Gly Asn Arg Thr SerPro Val Pro Asp Thr Leu Ile Ile Lys Gly Ser Gly Asn Arg Thr Ser
500 505 510 500 505 510
Val Gly Ser Ser Ile Tyr Val Asn Thr Pro Ser Gly Ser Leu Val SerVal Gly Ser Ser Ile Tyr Val Asn Thr Pro Ser Gly Ser Leu Val Ser
515 520 525 515 520 525
Ser Glu Ala Gln Leu Phe Asn Lys Pro Tyr Trp Leu Gln Lys Ala GlnSer Glu Ala Gln Leu Phe Asn Lys Pro Tyr Trp Leu Gln Lys Ala Gln
530 535 540 530 535 540
Gly His Asn Asn Gly Ile Cys Trp Gly Asn Gln Leu Phe Val Thr ValGly His Asn Asn Gly Ile Cys Trp Gly Asn Gln Leu Phe Val Thr Val
545 550 555 560545 550 555 560
Val Asp Thr Thr Arg Ser Thr Asn Met Thr Leu Cys Ala Ser Val ThrVal Asp Thr Thr Arg Ser Thr Asn Met Thr Leu Cys Ala Ser Val Thr
565 570 575 565 570 575
Thr Ser Ser Thr Tyr Thr Asn Ser Asp Tyr Lys Glu Tyr Met Arg HisThr Ser Ser Thr Tyr Thr Asn Ser Asp Tyr Lys Glu Tyr Met Arg His
580 585 590 580 585 590
Val Glu Glu Tyr Asp Leu Gln Phe Ile Phe Gln Leu Cys Ser Ile ThrVal Glu Glu Tyr Asp Leu Gln Phe Ile Phe Gln Leu Cys Ser Ile Thr
595 600 605 595 600 605
Leu Ser Ala Glu Val Met Ala Tyr Ile His Thr Met Asn Pro Ser ValLeu Ser Ala Glu Val Met Ala Tyr Ile His Thr Met Asn Pro Ser Val
610 615 620 610 615 620
Leu Glu Asp Trp Asn Phe Gly Leu Ser Pro Pro Pro Asn Gly Thr LeuLeu Glu Asp Trp Asn Phe Gly Leu Ser Pro Pro Pro Asn Gly Thr Leu
625 630 635 640625 630 635 640
Glu Asp Thr Tyr Arg Tyr Val Gln Ser Gln Ala Ile Thr Cys Gln LysGlu Asp Thr Tyr Arg Tyr Val Gln Ser Gln Ala Ile Thr Cys Gln Lys
645 650 655 645 650 655
Pro Thr Pro Glu Lys Glu Lys Pro Asp Pro Tyr Lys Asn Leu Ser PhePro Thr Pro Glu Lys Glu Lys Pro Asp Pro Tyr Lys Asn Leu Ser Phe
660 665 670 660 665 670
Trp Glu Val Asn Leu Lys Glu Lys Phe Ser Ser Glu Leu Asp Gln TyrTrp Glu Val Asn Leu Lys Glu Lys Phe Ser Ser Glu Leu Asp Gln Tyr
675 680 685 675 680 685
Pro Leu Gly Arg Lys Phe Leu Leu Gln Ser Gly Tyr Arg Gly Arg SerPro Leu Gly Arg Lys Phe Leu Leu Gln Ser Gly Tyr Arg Gly Arg Ser
690 695 700 690 695 700
Ser Ile Arg Thr Gly Val Lys Arg Pro Ala Val Ser Lys Ala Ser AlaSer Ile Arg Thr Gly Val Lys Arg Pro Ala Val Ser Lys Ala Ser Ala
705 710 715 720705 710 715 720
Ala Pro Lys Arg Lys Arg Ala Lys Thr Lys ArgAla Pro Lys Arg Lys Arg Ala Lys Thr Lys Arg
725 730 725 730
<210> 9<210> 9
<211> 825<211> 825
<212> PRT<212> PRT
<213> 人工序列<213> Artificial sequence
<400> 9<400> 9
Met Ser Pro Ile Leu Gly Tyr Trp Lys Ile Lys Gly Leu Val Gln ProMet Ser Pro Ile Leu Gly Tyr Trp Lys Ile Lys Gly Leu Val Gln Pro
1 5 10 151 5 10 15
Thr Arg Leu Leu Leu Glu Tyr Leu Glu Glu Lys Tyr Glu Glu His LeuThr Arg Leu Leu Leu Glu Tyr Leu Glu Glu Lys Tyr Glu Glu His Leu
20 25 30 20 25 30
Tyr Glu Arg Asp Glu Gly Asp Lys Trp Arg Asn Lys Lys Phe Glu LeuTyr Glu Arg Asp Glu Gly Asp Lys Trp Arg Asn Lys Lys Phe Glu Leu
35 40 45 35 40 45
Gly Leu Glu Phe Pro Asn Leu Pro Tyr Tyr Ile Asp Gly Asp Val LysGly Leu Glu Phe Pro Asn Leu Pro Tyr Tyr Ile Asp Gly Asp Val Lys
50 55 60 50 55 60
Leu Thr Gln Ser Met Ala Ile Ile Arg Tyr Ile Ala Asp Lys His AsnLeu Thr Gln Ser Met Ala Ile Ile Arg Tyr Ile Ala Asp Lys His Asn
65 70 75 8065 70 75 80
Met Leu Gly Gly Cys Pro Lys Glu Arg Ala Glu Ile Ser Met Leu GluMet Leu Gly Gly Cys Pro Lys Glu Arg Ala Glu Ile Ser Met Leu Glu
85 90 95 85 90 95
Gly Ala Val Leu Asp Ile Arg Tyr Gly Val Ser Arg Ile Ala Tyr SerGly Ala Val Leu Asp Ile Arg Tyr Gly Val Ser Arg Ile Ala Tyr Ser
100 105 110 100 105 110
Lys Asp Phe Glu Thr Leu Lys Val Asp Phe Leu Ser Lys Leu Pro GluLys Asp Phe Glu Thr Leu Lys Val Asp Phe Leu Ser Lys Leu Pro Glu
115 120 125 115 120 125
Met Leu Lys Met Phe Glu Asp Arg Leu Cys His Lys Thr Tyr Leu AsnMet Leu Lys Met Phe Glu Asp Arg Leu Cys His Lys Thr Tyr Leu Asn
130 135 140 130 135 140
Gly Asp His Val Thr His Pro Asp Phe Met Leu Tyr Asp Ala Leu AspGly Asp His Val Thr His Pro Asp Phe Met Leu Tyr Asp Ala Leu Asp
145 150 155 160145 150 155 160
Val Val Leu Tyr Met Asp Pro Met Cys Leu Asp Ala Phe Pro Lys LeuVal Val Leu Tyr Met Asp Pro Met Cys Leu Asp Ala Phe Pro Lys Leu
165 170 175 165 170 175
Val Cys Phe Lys Lys Arg Ile Glu Ala Ile Pro Gln Ile Asp Lys TyrVal Cys Phe Lys Lys Arg Ile Glu Ala Ile Pro Gln Ile Asp Lys Tyr
180 185 190 180 185 190
Leu Lys Ser Ser Lys Tyr Ile Ala Trp Pro Leu Gln Gly Trp Gln AlaLeu Lys Ser Ser Lys Tyr Ile Ala Trp Pro Leu Gln Gly Trp Gln Ala
195 200 205 195 200 205
Thr Phe Gly Gly Gly Asp His Pro Pro Lys Ser Asp Leu Glu Val LeuThr Phe Gly Gly Gly Asp His Pro Pro Lys Ser Asp Leu Glu Val Leu
210 215 220 210 215 220
Phe Gln Gly Pro Ser Asp Gln Glu Ala Lys Pro Ser Thr Glu Asp LeuPhe Gln Gly Pro Ser Asp Gln Glu Ala Lys Pro Ser Thr Glu Asp Leu
225 230 235 240225 230 235 240
Gly Asp Lys Lys Glu Gly Glu Tyr Ile Lys Leu Lys Val Ile Gly GlnGly Asp Lys Lys Glu Gly Glu Tyr Ile Lys Leu Lys Val Ile Gly Gln
245 250 255 245 250 255
Asp Ser Ser Glu Ile His Phe Lys Val Lys Met Thr Thr His Leu LysAsp Ser Ser Glu Ile His Phe Lys Val Lys Met Thr Thr His Leu Lys
260 265 270 260 265 270
Lys Leu Lys Glu Ser Tyr Cys Gln Arg Gln Gly Val Pro Met Asn SerLys Leu Lys Glu Ser Tyr Cys Gln Arg Gln Gly Val Pro Met Asn Ser
275 280 285 275 280 285
Leu Arg Phe Leu Phe Glu Gly Gln Arg Ile Ala Asp Asn His Thr ProLeu Arg Phe Leu Phe Glu Gly Gln Arg Ile Ala Asp Asn His Thr Pro
290 295 300 290 295 300
Lys Glu Leu Gly Met Glu Glu Glu Asp Val Ile Glu Val Tyr Gln GluLys Glu Leu Gly Met Glu Glu Glu Asp Val Ile Glu Val Tyr Gln Glu
305 310 315 320305 310 315 320
Gln Thr Gly Gly Ser Met Trp Arg Pro Ser Asp Ser Thr Val Tyr ValGln Thr Gly Gly Ser Met Trp Arg Pro Ser Asp Ser Thr Val Tyr Val
325 330 335 325 330 335
Pro Pro Pro Asn Pro Val Ser Lys Val Val Ala Thr Asp Ala Tyr ValPro Pro Pro Asn Pro Val Ser Lys Val Val Ala Thr Asp Ala Tyr Val
340 345 350 340 345 350
Thr Arg Thr Asn Ile Phe Tyr His Ala Ser Ser Ser Arg Leu Leu AlaThr Arg Thr Asn Ile Phe Tyr His Ala Ser Ser Ser Arg Leu Leu Ala
355 360 365 355 360 365
Val Gly His Pro Tyr Phe Ser Ile Lys Arg Ala Asn Lys Thr Val ValVal Gly His Pro Tyr Phe Ser Ile Lys Arg Ala Asn Lys Thr Val Val
370 375 380 370 375 380
Pro Lys Val Ser Gly Tyr Gln Tyr Arg Val Phe Lys Val Val Leu ProPro Lys Val Ser Gly Tyr Gln Tyr Arg Val Phe Lys Val Val Leu Pro
385 390 395 400385 390 395 400
Asp Pro Asn Lys Phe Ala Leu Pro Asp Ser Ser Leu Phe Asp Pro ThrAsp Pro Asn Lys Phe Ala Leu Pro Asp Ser Ser Leu Phe Asp Pro Thr
405 410 415 405 410 415
Thr Gln Arg Leu Val Trp Ala Cys Thr Gly Leu Glu Val Gly Arg GlyThr Gln Arg Leu Val Trp Ala Cys Thr Gly Leu Glu Val Gly Arg Gly
420 425 430 420 425 430
Gln Pro Leu Gly Val Gly Val Ser Gly His Pro Phe Leu Asn Lys TyrGln Pro Leu Gly Val Gly Val Ser Gly His Pro Phe Leu Asn Lys Tyr
435 440 445 435 440 445
Asp Asp Val Glu Asn Ser Gly Ser Gly Gly Asn Pro Gly Gln Asp AsnAsp Asp Val Glu Asn Ser Gly Ser Gly Gly Asn Pro Gly Gln Asp Asn
450 455 460 450 455 460
Arg Val Asn Val Gly Met Asp Tyr Lys Gln Thr Gln Leu Cys Met ValArg Val Asn Val Gly Met Asp Tyr Lys Gln Thr Gln Leu Cys Met Val
465 470 475 480465 470 475 480
Gly Cys Ala Pro Pro Leu Gly Glu His Trp Gly Lys Gly Lys Gln CysGly Cys Ala Pro Pro Leu Gly Glu His Trp Gly Lys Gly Lys Gln Cys
485 490 495 485 490 495
Thr Asn Thr Pro Val Gln Ala Gly Asp Cys Pro Pro Leu Glu Leu IleThr Asn Thr Pro Val Gln Ala Gly Asp Cys Pro Pro Leu Glu Leu Ile
500 505 510 500 505 510
Thr Ser Val Ile Gln Asp Gly Asp Met Val Asp Thr Gly Phe Gly AlaThr Ser Val Ile Gln Asp Gly Asp Met Val Asp Thr Gly Phe Gly Ala
515 520 525 515 520 525
Met Asn Phe Ala Asp Leu Gln Thr Asn Lys Ser Asp Val Pro Ile AspMet Asn Phe Ala Asp Leu Gln Thr Asn Lys Ser Asp Val Pro Ile Asp
530 535 540 530 535 540
Ile Cys Gly Thr Thr Cys Lys Tyr Pro Asp Tyr Leu Gln Met Ala AlaIle Cys Gly Thr Thr Thr Cys Lys Tyr Pro Asp Tyr Leu Gln Met Ala Ala
545 550 555 560545 550 555 560
Asp Pro Tyr Gly Asp Arg Leu Phe Phe Phe Leu Arg Lys Glu Gln MetAsp Pro Tyr Gly Asp Arg Leu Phe Phe Phe Leu Arg Lys Glu Gln Met
565 570 575 565 570 575
Phe Ala Arg His Phe Phe Asn Arg Ala Gly Glu Val Gly Glu Pro ValPhe Ala Arg His Phe Phe Asn Arg Ala Gly Glu Val Gly Glu Pro Val
580 585 590 580 585 590
Pro Asp Thr Leu Ile Ile Lys Gly Ser Gly Asn Arg Thr Ser Val GlyPro Asp Thr Leu Ile Ile Lys Gly Ser Gly Asn Arg Thr Ser Val Gly
595 600 605 595 600 605
Ser Ser Ile Tyr Val Asn Thr Pro Ser Gly Ser Leu Val Ser Ser GluSer Ser Ile Tyr Val Asn Thr Pro Ser Gly Ser Leu Val Ser Ser Glu
610 615 620 610 615 620
Ala Gln Leu Phe Asn Lys Pro Tyr Trp Leu Gln Lys Ala Gln Gly HisAla Gln Leu Phe Asn Lys Pro Tyr Trp Leu Gln Lys Ala Gln Gly His
625 630 635 640625 630 635 640
Asn Asn Gly Ile Cys Trp Gly Asn Gln Leu Phe Val Thr Val Val AspAsn Asn Gly Ile Cys Trp Gly Asn Gln Leu Phe Val Thr Val Val Asp
645 650 655 645 650 655
Thr Thr Arg Ser Thr Asn Met Thr Leu Cys Ala Ser Val Thr Thr SerThr Thr Arg Ser Thr Asn Met Thr Leu Cys Ala Ser Val Thr Thr Ser
660 665 670 660 665 670
Ser Thr Tyr Thr Asn Ser Asp Tyr Lys Glu Tyr Met Arg His Val GluSer Thr Tyr Thr Asn Ser Asp Tyr Lys Glu Tyr Met Arg His Val Glu
675 680 685 675 680 685
Glu Tyr Asp Leu Gln Phe Ile Phe Gln Leu Cys Ser Ile Thr Leu SerGlu Tyr Asp Leu Gln Phe Ile Phe Gln Leu Cys Ser Ile Thr Leu Ser
690 695 700 690 695 700
Ala Glu Val Met Ala Tyr Ile His Thr Met Asn Pro Ser Val Leu GluAla Glu Val Met Ala Tyr Ile His Thr Met Asn Pro Ser Val Leu Glu
705 710 715 720705 710 715 720
Asp Trp Asn Phe Gly Leu Ser Pro Pro Pro Asn Gly Thr Leu Glu AspAsp Trp Asn Phe Gly Leu Ser Pro Pro Pro Asn Gly Thr Leu Glu Asp
725 730 735 725 730 735
Thr Tyr Arg Tyr Val Gln Ser Gln Ala Ile Thr Cys Gln Lys Pro ThrThr Tyr Arg Tyr Val Gln Ser Gln Ala Ile Thr Cys Gln Lys Pro Thr
740 745 750 740 745 750
Pro Glu Lys Glu Lys Pro Asp Pro Tyr Lys Asn Leu Ser Phe Trp GluPro Glu Lys Glu Lys Pro Asp Pro Tyr Lys Asn Leu Ser Phe Trp Glu
755 760 765 755 760 765
Val Asn Leu Lys Glu Lys Phe Ser Ser Glu Leu Asp Gln Tyr Pro LeuVal Asn Leu Lys Glu Lys Phe Ser Ser Glu Leu Asp Gln Tyr Pro Leu
770 775 780 770 775 780
Gly Arg Lys Phe Leu Leu Gln Ser Gly Tyr Arg Gly Arg Ser Ser IleGly Arg Lys Phe Leu Leu Gln Ser Gly Tyr Arg Gly Arg Ser Ser Ile
785 790 795 800785 790 795 800
Arg Thr Gly Val Lys Arg Pro Ala Val Ser Lys Ala Ser Ala Ala ProArg Thr Gly Val Lys Arg Pro Ala Val Ser Lys Ala Ser Ala Ala Pro
805 810 815 805 810 815
Lys Arg Lys Arg Ala Lys Thr Lys ArgLys Arg Lys Arg Ala Lys Thr Lys Arg
820 825 820 825
<210> 10<210> 10
<211> 891<211> 891
<212> PRT<212> PRT
<213> 人工序列<213> Artificial sequence
<400> 10<400> 10
Met Lys Ile Glu Glu Gly Lys Leu Val Ile Trp Ile Asn Gly Asp LysMet Lys Ile Glu Glu Gly Lys Leu Val Ile Trp Ile Asn Gly Asp Lys
1 5 10 151 5 10 15
Gly Tyr Asn Gly Leu Ala Glu Val Gly Lys Lys Phe Glu Lys Asp ThrGly Tyr Asn Gly Leu Ala Glu Val Gly Lys Lys Phe Glu Lys Asp Thr
20 25 30 20 25 30
Gly Ile Lys Val Thr Val Glu His Pro Asp Lys Leu Glu Glu Lys PheGly Ile Lys Val Thr Val Glu His Pro Asp Lys Leu Glu Glu Lys Phe
35 40 45 35 40 45
Pro Gln Val Ala Ala Thr Gly Asp Gly Pro Asp Ile Ile Phe Trp AlaPro Gln Val Ala Ala Thr Gly Asp Gly Pro Asp Ile Ile Phe Trp Ala
50 55 60 50 55 60
His Asp Arg Phe Gly Gly Tyr Ala Gln Ser Gly Leu Leu Ala Glu IleHis Asp Arg Phe Gly Gly Tyr Ala Gln Ser Gly Leu Leu Ala Glu Ile
65 70 75 8065 70 75 80
Thr Pro Asp Lys Ala Phe Gln Asp Lys Leu Tyr Pro Phe Thr Trp AspThr Pro Asp Lys Ala Phe Gln Asp Lys Leu Tyr Pro Phe Thr Trp Asp
85 90 95 85 90 95
Ala Val Arg Tyr Asn Gly Lys Leu Ile Ala Tyr Pro Ile Ala Val GluAla Val Arg Tyr Asn Gly Lys Leu Ile Ala Tyr Pro Ile Ala Val Glu
100 105 110 100 105 110
Ala Leu Ser Leu Ile Tyr Asn Lys Asp Leu Leu Pro Asn Pro Pro LysAla Leu Ser Leu Ile Tyr Asn Lys Asp Leu Leu Pro Asn Pro Pro Lys
115 120 125 115 120 125
Thr Trp Glu Glu Ile Pro Ala Leu Asp Lys Glu Leu Lys Ala Lys GlyThr Trp Glu Glu Ile Pro Ala Leu Asp Lys Glu Leu Lys Ala Lys Gly
130 135 140 130 135 140
Lys Ser Ala Leu Met Phe Asn Leu Gln Glu Pro Tyr Phe Thr Trp ProLys Ser Ala Leu Met Phe Asn Leu Gln Glu Pro Tyr Phe Thr Trp Pro
145 150 155 160145 150 155 160
Leu Ile Ala Ala Asp Gly Gly Tyr Ala Phe Lys Tyr Glu Asn Gly LysLeu Ile Ala Ala Asp Gly Gly Tyr Ala Phe Lys Tyr Glu Asn Gly Lys
165 170 175 165 170 175
Tyr Asp Ile Lys Asp Val Gly Val Asp Asn Ala Gly Ala Lys Ala GlyTyr Asp Ile Lys Asp Val Gly Val Asp Asn Ala Gly Ala Lys Ala Gly
180 185 190 180 185 190
Leu Thr Phe Leu Val Asp Leu Ile Lys Asn Lys His Met Asn Ala AspLeu Thr Phe Leu Val Asp Leu Ile Lys Asn Lys His Met Asn Ala Asp
195 200 205 195 200 205
Thr Asp Tyr Ser Ile Ala Glu Ala Ala Phe Asn Lys Gly Glu Thr AlaThr Asp Tyr Ser Ile Ala Glu Ala Ala Phe Asn Lys Gly Glu Thr Ala
210 215 220 210 215 220
Met Thr Ile Asn Gly Pro Trp Ala Trp Ser Asn Ile Asp Thr Ser LysMet Thr Ile Asn Gly Pro Trp Ala Trp Ser Asn Ile Asp Thr Ser Lys
225 230 235 240225 230 235 240
Val Asn Tyr Gly Val Thr Val Leu Pro Thr Phe Lys Gly Gln Pro SerVal Asn Tyr Gly Val Thr Val Leu Pro Thr Phe Lys Gly Gly Gln Pro Ser
245 250 255 245 250 255
Lys Pro Phe Val Gly Val Leu Ser Ala Gly Ile Asn Ala Ala Ser ProLys Pro Phe Val Gly Val Leu Ser Ala Gly Ile Asn Ala Ala Ser Pro
260 265 270 260 265 270
Asn Lys Glu Leu Ala Lys Glu Phe Leu Glu Asn Tyr Leu Leu Thr AspAsn Lys Glu Leu Ala Lys Glu Phe Leu Glu Asn Tyr Leu Leu Thr Asp
275 280 285 275 280 285
Glu Gly Leu Glu Ala Val Asn Lys Asp Lys Pro Leu Gly Ala Val AlaGlu Gly Leu Glu Ala Val Asn Lys Asp Lys Pro Leu Gly Ala Val Ala
290 295 300 290 295 300
Leu Lys Ser Tyr Glu Glu Glu Leu Ala Lys Asp Pro Arg Ile Ala AlaLeu Lys Ser Tyr Glu Glu Glu Leu Ala Lys Asp Pro Arg Ile Ala Ala
305 310 315 320305 310 315 320
Thr Met Glu Asn Ala Gln Lys Gly Glu Ile Met Pro Asn Ile Pro GlnThr Met Glu Asn Ala Gln Lys Gly Glu Ile Met Pro Asn Ile Pro Gln
325 330 335 325 330 335
Met Ser Ala Phe Trp Tyr Ala Val Arg Thr Ala Val Ile Asn Ala AlaMet Ser Ala Phe Trp Tyr Ala Val Arg Thr Ala Val Ile Asn Ala Ala
340 345 350 340 345 350
Ser Gly Arg Gln Thr Val Asp Glu Ala Leu Lys Asp Ala Gln Thr AsnSer Gly Arg Gln Thr Val Asp Glu Ala Leu Lys Asp Ala Gln Thr Asn
355 360 365 355 360 365
Ser Ser Ser Asn Asn Asn Asn Asn Asn Asn Asn Asn Asn Leu Gly IleSer Ser Ser Asn Asn Asn Asn Asn Asn Asn Asn Asn Asn Asn Asn Leu Gly Ile
370 375 380 370 375 380
Glu Gly Arg Ile Ser Glu Phe Met Trp Arg Pro Ser Asp Ser Thr ValGlu Gly Arg Ile Ser Glu Phe Met Trp Arg Pro Ser Asp Ser Thr Val
385 390 395 400385 390 395 400
Tyr Val Pro Pro Pro Asn Pro Val Ser Lys Val Val Ala Thr Asp AlaTyr Val Pro Pro Pro Asn Pro Val Ser Lys Val Val Ala Thr Asp Ala
405 410 415 405 410 415
Tyr Val Thr Arg Thr Asn Ile Phe Tyr His Ala Ser Ser Ser Arg LeuTyr Val Thr Arg Thr Asn Ile Phe Tyr His Ala Ser Ser Ser Arg Leu
420 425 430 420 425 430
Leu Ala Val Gly His Pro Tyr Phe Ser Ile Lys Arg Ala Asn Lys ThrLeu Ala Val Gly His Pro Tyr Phe Ser Ile Lys Arg Ala Asn Lys Thr
435 440 445 435 440 445
Val Val Pro Lys Val Ser Gly Tyr Gln Tyr Arg Val Phe Lys Val ValVal Val Pro Lys Val Ser Gly Tyr Gln Tyr Arg Val Phe Lys Val Val
450 455 460 450 455 460
Leu Pro Asp Pro Asn Lys Phe Ala Leu Pro Asp Ser Ser Leu Phe AspLeu Pro Asp Pro Asn Lys Phe Ala Leu Pro Asp Ser Ser Leu Phe Asp
465 470 475 480465 470 475 480
Pro Thr Thr Gln Arg Leu Val Trp Ala Cys Thr Gly Leu Glu Val GlyPro Thr Thr Gln Arg Leu Val Trp Ala Cys Thr Gly Leu Glu Val Gly
485 490 495 485 490 495
Arg Gly Gln Pro Leu Gly Val Gly Val Ser Gly His Pro Phe Leu AsnArg Gly Gln Pro Leu Gly Val Gly Val Ser Gly His Pro Phe Leu Asn
500 505 510 500 505 510
Lys Tyr Asp Asp Val Glu Asn Ser Gly Ser Gly Gly Asn Pro Gly GlnLys Tyr Asp Asp Val Glu Asn Ser Gly Ser Gly Gly Asn Pro Gly Gln
515 520 525 515 520 525
Asp Asn Arg Val Asn Val Gly Met Asp Tyr Lys Gln Thr Gln Leu CysAsp Asn Arg Val Asn Val Gly Met Asp Tyr Lys Gln Thr Gln Leu Cys
530 535 540 530 535 540
Met Val Gly Cys Ala Pro Pro Leu Gly Glu His Trp Gly Lys Gly LysMet Val Gly Cys Ala Pro Pro Leu Gly Glu His Trp Gly Lys Gly Lys
545 550 555 560545 550 555 560
Gln Cys Thr Asn Thr Pro Val Gln Ala Gly Asp Cys Pro Pro Leu GluGln Cys Thr Asn Thr Pro Val Gln Ala Gly Asp Cys Pro Pro Leu Glu
565 570 575 565 570 575
Leu Ile Thr Ser Val Ile Gln Asp Gly Asp Met Val Asp Thr Gly PheLeu Ile Thr Ser Val Ile Gln Asp Gly Asp Met Val Asp Thr Gly Phe
580 585 590 580 585 590
Gly Ala Met Asn Phe Ala Asp Leu Gln Thr Asn Lys Ser Asp Val ProGly Ala Met Asn Phe Ala Asp Leu Gln Thr Asn Lys Ser Asp Val Pro
595 600 605 595 600 605
Ile Asp Ile Cys Gly Thr Thr Cys Lys Tyr Pro Asp Tyr Leu Gln MetIle Asp Ile Cys Gly Thr Thr Cys Lys Tyr Pro Asp Tyr Leu Gln Met
610 615 620 610 615 620
Ala Ala Asp Pro Tyr Gly Asp Arg Leu Phe Phe Phe Leu Arg Lys GluAla Ala Asp Pro Tyr Gly Asp Arg Leu Phe Phe Phe Leu Arg Lys Glu
625 630 635 640625 630 635 640
Gln Met Phe Ala Arg His Phe Phe Asn Arg Ala Gly Glu Val Gly GluGln Met Phe Ala Arg His Phe Phe Asn Arg Ala Gly Glu Val Gly Glu
645 650 655 645 650 655
Pro Val Pro Asp Thr Leu Ile Ile Lys Gly Ser Gly Asn Arg Thr SerPro Val Pro Asp Thr Leu Ile Ile Lys Gly Ser Gly Asn Arg Thr Ser
660 665 670 660 665 670
Val Gly Ser Ser Ile Tyr Val Asn Thr Pro Ser Gly Ser Leu Val SerVal Gly Ser Ser Ile Tyr Val Asn Thr Pro Ser Gly Ser Leu Val Ser
675 680 685 675 680 685
Ser Glu Ala Gln Leu Phe Asn Lys Pro Tyr Trp Leu Gln Lys Ala GlnSer Glu Ala Gln Leu Phe Asn Lys Pro Tyr Trp Leu Gln Lys Ala Gln
690 695 700 690 695 700
Gly His Asn Asn Gly Ile Cys Trp Gly Asn Gln Leu Phe Val Thr ValGly His Asn Asn Gly Ile Cys Trp Gly Asn Gln Leu Phe Val Thr Val
705 710 715 720705 710 715 720
Val Asp Thr Thr Arg Ser Thr Asn Met Thr Leu Cys Ala Ser Val ThrVal Asp Thr Thr Arg Ser Thr Asn Met Thr Leu Cys Ala Ser Val Thr
725 730 735 725 730 735
Thr Ser Ser Thr Tyr Thr Asn Ser Asp Tyr Lys Glu Tyr Met Arg HisThr Ser Ser Thr Tyr Thr Asn Ser Asp Tyr Lys Glu Tyr Met Arg His
740 745 750 740 745 750
Val Glu Glu Tyr Asp Leu Gln Phe Ile Phe Gln Leu Cys Ser Ile ThrVal Glu Glu Tyr Asp Leu Gln Phe Ile Phe Gln Leu Cys Ser Ile Thr
755 760 765 755 760 765
Leu Ser Ala Glu Val Met Ala Tyr Ile His Thr Met Asn Pro Ser ValLeu Ser Ala Glu Val Met Ala Tyr Ile His Thr Met Asn Pro Ser Val
770 775 780 770 775 780
Leu Glu Asp Trp Asn Phe Gly Leu Ser Pro Pro Pro Asn Gly Thr LeuLeu Glu Asp Trp Asn Phe Gly Leu Ser Pro Pro Pro Asn Gly Thr Leu
785 790 795 800785 790 795 800
Glu Asp Thr Tyr Arg Tyr Val Gln Ser Gln Ala Ile Thr Cys Gln LysGlu Asp Thr Tyr Arg Tyr Val Gln Ser Gln Ala Ile Thr Cys Gln Lys
805 810 815 805 810 815
Pro Thr Pro Glu Lys Glu Lys Pro Asp Pro Tyr Lys Asn Leu Ser PhePro Thr Pro Glu Lys Glu Lys Pro Asp Pro Tyr Lys Asn Leu Ser Phe
820 825 830 820 825 830
Trp Glu Val Asn Leu Lys Glu Lys Phe Ser Ser Glu Leu Asp Gln TyrTrp Glu Val Asn Leu Lys Glu Lys Phe Ser Ser Glu Leu Asp Gln Tyr
835 840 845 835 840 845
Pro Leu Gly Arg Lys Phe Leu Leu Gln Ser Gly Tyr Arg Gly Arg SerPro Leu Gly Arg Lys Phe Leu Leu Gln Ser Gly Tyr Arg Gly Arg Ser
850 855 860 850 855 860
Ser Ile Arg Thr Gly Val Lys Arg Pro Ala Val Ser Lys Ala Ser AlaSer Ile Arg Thr Gly Val Lys Arg Pro Ala Val Ser Lys Ala Ser Ala
865 870 875 880865 870 875 880
Ala Pro Lys Arg Lys Arg Ala Lys Thr Lys ArgAla Pro Lys Arg Lys Arg Ala Lys Thr Lys Arg
885 890 885 890
<210> 11<210> 11
<211> 2088<211> 2088
<212> DNA<212>DNA
<213> 人工序列<213> Artificial sequence
<400> 11<400> 11
atgtccccta tactaggtta ttggaaaatt aagggccttg tgcaacccac tcgacttctt 60atgtccccta tactaggtta ttggaaaatt aagggccttg tgcaacccac tcgacttctt 60
ttggaatatc ttgaagaaaa atatgaagag catttgtatg agcgcgatga aggtgataaa 120ttggaatatc ttgaagaaaa atatgaagag catttgtatg agcgcgatga aggtgataaa 120
tggcgaaaca aaaagtttga attgggtttg gagtttccca atcttcctta ttatattgat 180tggcgaaaca aaaagtttga attgggtttg gagtttccca atcttcctta ttatattgat 180
ggtgatgtta aattaacaca gtctatggcc atcatacgtt atatagctga caagcacaac 240ggtgatgtta aattaacaca gtctatggcc atcatacgtt atatagctga caagcacaac 240
atgttgggtg gttgtccaaa agagcgtgca gagatttcaa tgcttgaagg agcggttttg 300atgttgggtg gttgtccaaa agagcgtgca gagatttcaa tgcttgaagg agcggttttg 300
gatattagat acggtgtttc gagaattgca tatagtaaag actttgaaac tctcaaagtt 360gatattagat acggtgtttc gagaattgca tatagtaaag actttgaaac tctcaaagtt 360
gattttctta gcaagctacc tgaaatgctg aaaatgttcg aagatcgttt atgtcataaa 420gattttctta gcaagctacc tgaaatgctg aaaatgttcg aagatcgttt atgtcataaa 420
acatatttaa atggtgatca tgtaacccat cctgacttca tgttgtatga cgctcttgat 480acatatttaa atggtgatca tgtaacccat cctgacttca tgttgtatga cgctcttgat 480
gttgttttat acatggaccc aatgtgcctg gatgcgttcc caaaattagt ttgttttaaa 540gttgttttat acatggaccc aatgtgcctg gatgcgttcc caaaattagt ttgttttaaa 540
aaacgtattg aagctatccc acaaattgat aagtacttga aatccagcaa gtatatagca 600aaacgtattg aagctatccc acaaattgat aagtacttga aatccagcaa gtatatagca 600
tggcctttgc agggctggca agccacgttt ggtggtggcg accatcctcc aaaatcggat 660tggcctttgc agggctggca agccacgttt ggtggtggcg accatcctcc aaaatcggat 660
ctggaagttc tgttccaggg gccctctgac tctaccgttt acgttccgcc gccgaacccg 720ctggaagttc tgttccaggg gccctctgac tctaccgttt acgttccgcc gccgaacccg 720
gtttctaaag ttgttgctac cgacgcttac gttacccgta ccaacatctt ctaccacgct 780gtttctaaag ttgttgctac cgacgcttac gttacccgta ccaacatctt ctaccacgct 780
tcttcttctc gtctgctggc tgttggtcac ccgtacttct ctatcaaacg tgctaacaaa 840tcttcttctc gtctgctggc tgttggtcac ccgtacttct ctatcaaacg tgctaacaaa 840
accgttgttc cgaaagtttc tggttaccag taccgtgttt tcaaagttgt tctgccggac 900accgttgttc cgaaagtttc tggttaccag taccgtgttt tcaaagttgt tctgccggac 900
ccgaacaaat tcgctctgcc ggactcttct ctgttcgacc cgaccaccca gcgtctggtt 960ccgaacaaat tcgctctgcc ggactcttct ctgttcgacc cgaccaccca gcgtctggtt 960
tgggcttgca ccggtctgga agttggtcgt ggtcagccgc tgggtgttgg tgtttctggt 1020tgggcttgca ccggtctgga agttggtcgt ggtcagccgc tgggtgttgg tgtttctggt 1020
cacccgttcc tgaacaaata cgacgacgtt gaaaactctg gttctggtgg taacccgggt 1080cacccgttcc tgaacaaata cgacgacgtt gaaaactctg gttctggtgg taacccgggt 1080
caggacaacc gtgttaacgt tggtatggac tacaaacaga cccagctgtg catggttggt 1140caggacaacc gtgttaacgt tggtatggac tacaaacaga cccagctgtg catggttggt 1140
tgcgctccgc cgctgggtga acactggggt aaaggtaaac agtgcaccaa caccccggtt 1200tgcgctccgc cgctgggtga aacactggggt aaaggtaaac agtgcaccaa caccccggtt 1200
caggctggtg actgcccgcc gctggaactg atcacctctg ttatccagga cggtgacatg 1260caggctggtg actgcccgcc gctggaactg atcacctctg ttatccagga cggtgacatg 1260
gttgacaccg gtttcggtgc tatgaacttc gctgacctgc agaccaacaa atctgacgtt 1320gttgacaccg gtttcggtgc tatgaacttc gctgacctgc agaccaacaa atctgacgtt 1320
ccgatcgaca tctgcggtac cacctgcaaa tacccggact acctgcagat ggctgctgac 1380ccgatcgaca tctgcggtac cacctgcaaa tacccggact acctgcagat ggctgctgac 1380
ccgtacggtg accgtctgtt cttcttcctg cgtaaagaac agatgttcgc tcgtcacttc 1440ccgtacggtg accgtctgtt cttcttcctg cgtaaagaac agatgttcgc tcgtcacttc 1440
ttcaaccgtg ctggtgaagt tggtgaaccg gttccggaca ccctgatcat caaaggttct 1500ttcaaccgtg ctggtgaagt tggtgaaccg gttccggaca ccctgatcat caaaggttct 1500
ggtaaccgta cctctgttgg ttcttctatc tacgttaaca ccccgtctgg ttctctggtt 1560ggtaaccgta cctctgttgg ttcttctatc tacgttaaca ccccgtctgg ttctctggtt 1560
tcttctgaag ctcagctgtt caacaaaccg tactggctgc agaaagctca gggtcacaac 1620tcttctgaag ctcagctgtt caacaaaccg tactggctgc agaaagctca gggtcacaac 1620
aacggtatct gctggggtaa ccagctgttc gttaccgttg ttgacaccac ccgttctacc 1680aacggtatct gctggggtaa ccagctgttc gttaccgttg ttgacaccac ccgttctacc 1680
aacatgaccc tgtgcgcttc tgttaccacc tcttctacct acaccaactc tgactacaaa 1740aacatgaccc tgtgcgcttc tgttaccacc tcttctacct acaccaactc tgactacaaa 1740
gaatacatgc gtcacgttga agaatacgac ctgcagttca tcttccagct gtgctctatc 1800gaatacatgc gtcacgttga agaatacgac ctgcagttca tcttccagct gtgctctatc 1800
accctgtctg ctgaagttat ggcttacatc cacaccatga acccgtctgt tctggaagac 1860accctgtctg ctgaagttat ggcttacatc cacaccatga accccgtctgt tctggaagac 1860
tggaacttcg gtctgtctcc gccgccgaac ggtaccctgg aagacaccta ccgttacgtt 1920tggaacttcg gtctgtctcc gccgccgaac ggtaccctgg aagacaccta ccgttacgtt 1920
cagtctcagg ctatcacctg ccagaaaccg accccggaaa aagaaaaacc ggacccgtac 1980cagtctcagg ctatcacctg ccagaaaccg accccggaaa aagaaaaacc ggacccgtac 1980
aaaaacctgt ctttctggga agttaacctg aaagaaaaat tctcttctga actggaccag 2040aaaaacctgt ctttctggga agttaacctg aaagaaaaat tctcttctga actggaccag 2040
tacccgctgg gtcgtaaatt cctgctgcag tctggttacc gtggttag 2088tacccgctgg gtcgtaaatt cctgctgcag tctggttacc gtggttag 2088
<210> 12<210> 12
<211> 2457<211> 2457
<212> DNA<212>DNA
<213> 人工序列<213> Artificial sequence
<400> 12<400> 12
atgtccccta tactaggtta ttggaaaatt aagggccttg tgcaacccac tcgacttctt 60atgtccccta tactaggtta ttggaaaatt aagggccttg tgcaacccac tcgacttctt 60
ttggaatatc ttgaagaaaa atatgaagag catttgtatg agcgcgatga aggtgataaa 120ttggaatatc ttgaagaaaa atatgaagag catttgtatg agcgcgatga aggtgataaa 120
tggcgaaaca aaaagtttga attgggtttg gagtttccca atcttcctta ttatattgat 180tggcgaaaca aaaagtttga attgggtttg gagtttccca atcttcctta ttatattgat 180
ggtgatgtta aattaacaca gtctatggcc atcatacgtt atatagctga caagcacaac 240ggtgatgtta aattaacaca gtctatggcc atcatacgtt atatagctga caagcacaac 240
atgttgggtg gttgtccaaa agagcgtgca gagatttcaa tgcttgaagg agcggttttg 300atgttgggtg gttgtccaaa agagcgtgca gagatttcaa tgcttgaagg agcggttttg 300
gatattagat acggtgtttc gagaattgca tatagtaaag actttgaaac tctcaaagtt 360gatattagat acggtgtttc gagaattgca tatagtaaag actttgaaac tctcaaagtt 360
gattttctta gcaagctacc tgaaatgctg aaaatgttcg aagatcgttt atgtcataaa 420gattttctta gcaagctacc tgaaatgctg aaaatgttcg aagatcgttt atgtcataaa 420
acatatttaa atggtgatca tgtaacccat cctgacttca tgttgtatga cgctcttgat 480acatatttaa atggtgatca tgtaacccat cctgacttca tgttgtatga cgctcttgat 480
gttgttttat acatggaccc aatgtgcctg gatgcgttcc caaaattagt ttgttttaaa 540gttgttttat acatggaccc aatgtgcctg gatgcgttcc caaaattagt ttgttttaaa 540
aaacgtattg aagctatccc acaaattgat aagtacttga aatccagcaa gtatatagca 600aaacgtattg aagctatccc acaaattgat aagtacttga aatccagcaa gtatatagca 600
tggcctttgc agggctggca agccacgttt ggtggtggcg accatcctcc aaaatcggat 660tggcctttgc agggctggca agccacgttt ggtggtggcg accatcctcc aaaatcggat 660
ctggaagttc tgttccaggg gccctctgac caggaagcta aaccgtctac cgaagacctg 720ctggaagttc tgttccaggg gccctctgac caggaagcta aaccgtctac cgaagacctg 720
ggtgacaaaa aagaaggtga atacatcaaa ctgaaagtta tcggtcagga ctcttctgaa 780ggtgacaaaa aagaaggtga atacatcaaa ctgaaagtta tcggtcagga ctcttctgaa 780
atccacttca aagttaaaat gaccacccac ctgaaaaaac tgaaagaatc ttactgccag 840atccacttca aagttaaaat gaccaccac ctgaaaaaac tgaaagaatc ttactgccag 840
cgtcagggtg ttccgatgaa ctctctgcgt ttcctgttcg aaggtcagcg tatcgctgac 900cgtcagggtg ttccgatgaa ctctctgcgt ttcctgttcg aaggtcagcg tatcgctgac 900
aaccacaccc cgaaagaact gggtatggaa gaagaagacg ttatcgaagt ttaccaggaa 960aaccaacaccc cgaaagaact gggtatggaa gaagaagacg ttatcgaagt ttaccaggaa 960
cagaccggtg gatcctctac cgtttacgtt ccgccgccga acccggtttc taaagttgtt 1020cagaccggtg gatcctctac cgtttacgtt ccgccgccga acccggtttc taaagttgtt 1020
gctaccgacg cttacgttac ccgtaccaac atcttctacc acgcttcttc ttctcgtctg 1080gctaccgacg cttacgttac ccgtaccaac atcttctacc acgcttcttc ttctcgtctg 1080
ctggctgttg gtcacccgta cttctctatc aaacgtgcta acaaaaccgt tgttccgaaa 1140ctggctgttg gtcacccgta cttctctatc aaacgtgcta acaaaaccgt tgttccgaaa 1140
gtttctggtt accagtaccg tgttttcaaa gttgttctgc cggacccgaa caaattcgct 1200gtttctggtt accagtaccg tgttttcaaa gttgttctgc cggacccgaa caaattcgct 1200
ctgccggact cttctctgtt cgacccgacc acccagcgtc tggtttgggc ttgcaccggt 1260ctgccggact cttctctgtt cgacccgacc acccagcgtc tggtttgggc ttgcaccggt 1260
ctggaagttg gtcgtggtca gccgctgggt gttggtgttt ctggtcaccc gttcctgaac 1320ctggaagttg gtcgtggtca gccgctgggt gttggtgttt ctggtcaccc gttcctgaac 1320
aaatacgacg acgttgaaaa ctctggttct ggtggtaacc cgggtcagga caaccgtgtt 1380aaatacgacg acgttgaaaa ctctggttct ggtggtaacc cgggtcagga caaccgtgtt 1380
aacgttggta tggactacaa acagacccag ctgtgcatgg ttggttgcgc tccgccgctg 1440aacgttggta tggactacaa acagacccag ctgtgcatgg ttggttgcgc tccgccgctg 1440
ggtgaacact ggggtaaagg taaacagtgc accaacaccc cggttcaggc tggtgactgc 1500ggtgaacact ggggtaaagg taaacagtgc accaacaccc cggttcaggc tggtgactgc 1500
ccgccgctgg aactgatcac ctctgttatc caggacggtg acatggttga caccggtttc 1560ccgccgctgg aactgatcac ctctgttatc caggacggtg acatggttga caccggtttc 1560
ggtgctatga acttcgctga cctgcagacc aacaaatctg acgttccgat cgacatctgc 1620ggtgctatga acttcgctga cctgcagacc aacaaatctg acgttccgat cgacatctgc 1620
ggtaccacct gcaaataccc ggactacctg cagatggctg ctgacccgta cggtgaccgt 1680ggtaccacct gcaaataccc ggactacctg cagatggctg ctgacccgta cggtgaccgt 1680
ctgttcttct tcctgcgtaa agaacagatg ttcgctcgtc acttcttcaa ccgtgctggt 1740ctgttcttct tcctgcgtaa agaacagatg ttcgctcgtc acttcttcaa ccgtgctggt 1740
gaagttggtg aaccggttcc ggacaccctg atcatcaaag gttctggtaa ccgtacctct 1800gaagttggtg aaccggttcc ggacaccctg atcatcaaag gttctggtaa ccgtacctct 1800
gttggttctt ctatctacgt taacaccccg tctggttctc tggtttcttc tgaagctcag 1860gttggttctt ctatctacgt taacaccccg tctggttctc tggtttcttc tgaagctcag 1860
ctgttcaaca aaccgtactg gctgcagaaa gctcagggtc acaacaacgg tatctgctgg 1920ctgttcaaca aaccgtactg gctgcagaaa gctcagggtc acaacaacgg tatctgctgg 1920
ggtaaccagc tgttcgttac cgttgttgac accacccgtt ctaccaacat gaccctgtgc 1980ggtaaccagc tgttcgttac cgttgttgac accacccgtt ctaccaacat gaccctgtgc 1980
gcttctgtta ccacctcttc tacctacacc aactctgact acaaagaata catgcgtcac 2040gcttctgtta ccaccctcttc tacctacacc aactctgact acaaagaata catgcgtcac 2040
gttgaagaat acgacctgca gttcatcttc cagctgtgct ctatcaccct gtctgctgaa 2100gttgaagaat acgacctgca gttcatcttc cagctgtgct ctatcaccct gtctgctgaa 2100
gttatggctt acatccacac catgaacccg tctgttctgg aagactggaa cttcggtctg 2160gttatggctt acatccacac catgaacccg tctgttctgg aagactggaa cttcggtctg 2160
tctccgccgc cgaacggtac cctggaagac acctaccgtt acgttcagtc tcaggctatc 2220tctccgccgc cgaacggtac cctggaagac acctaccgtt acgttcagtc tcaggctatc 2220
acctgccaga aaccgacccc ggaaaaagaa aaaccggacc cgtacaaaaa cctgtctttc 2280acctgccaga aaccgacccc ggaaaaagaa aaaccggacc cgtacaaaaa cctgtctttc 2280
tgggaagtta acctgaaaga aaaattctct tctgaactgg accagtaccc gctgggtcgt 2340tgggaagtta acctgaaaga aaaattctct tctgaactgg accagtaccc gctgggtcgt 2340
aaattcctgc tgcagtctgg ttaccgtggt cgttcttcta tccgtaccgg tgttaaacgt 2400aaattcctgc tgcagtctgg ttaccgtggt cgttcttcta tccgtaccgg tgttaaacgt 2400
ccggctgttt ctaaagcttc tgctgctccg aaacgtaaac gtgctaaaac caaacgt 2457ccggctgttt ctaaagcttc tgctgctccg aaacgtaaac gtgctaaaac caaacgt 2457
<210> 13<210> 13
<211> 2628<211> 2628
<212> DNA<212>DNA
<213> 人工序列<213> Artificial sequence
<400> 13<400> 13
atgaaaatcg aagaaggtaa actggtaatc tggattaacg gcgataaagg ctataacggt 60atgaaaatcg aagaaggtaa actggtaatc tggattaacg gcgataaagg ctataacggt 60
ctcgctgaag tcggtaagaa attcgagaaa gataccggaa ttaaagtcac cgttgagcat 120ctcgctgaag tcggtaagaa attcgagaaa gataccggaa ttaaagtcac cgttgagcat 120
ccggataaac tggaagagaa attcccacag gttgcggcaa ctggcgatgg ccctgacatt 180ccggataaac tggaagagaa attcccacag gttgcggcaa ctggcgatgg ccctgacatt 180
atcttctggg cacacgaccg ctttggtggc tacgctcaat ctggcctgtt ggctgaaatc 240atcttctggg cacacgaccg ctttggtggc tacgctcaat ctggcctgtt ggctgaaatc 240
accccggaca aagcgttcca ggacaagctg tatccgttta cctgggatgc cgtacgttac 300accccggaca aagcgttcca ggacaagctg tatccgttta cctgggatgc cgtacgttac 300
aacggcaagc tgattgctta cccgatcgct gttgaagcgt tatcgctgat ttataacaaa 360aacggcaagc tgattgctta cccgatcgct gttgaagcgt tatcgctgat ttataacaaa 360
gatctgctgc cgaacccgcc aaaaacctgg gaagagatcc cggcgctgga taaagaactg 420gatctgctgc cgaacccgcc aaaaacctgg gaagagatcc cggcgctgga taaagaactg 420
aaagcgaaag gtaagagcgc gctgatgttc aacctgcaag aaccgtactt cacctggccg 480aaagcgaaag gtaagagcgc gctgatgttc aacctgcaag aaccgtactt cacctggccg 480
ctgattgctg ctgacggggg ttatgcgttc aagtatgaaa acggcaagta cgacattaaa 540ctgattgctg ctgacgggggg ttatgcgttc aagtatgaaa acggcaagta cgacattaaa 540
gacgtgggcg tggataacgc tggcgcgaaa gcgggtctga ccttcctggt tgacctgatt 600gacgtgggcg tggataacgc tggcgcgaaa gcgggtctga ccttcctggt tgacctgatt 600
aaaaacaaac acatgaatgc agacaccgat tactccatcg cagaagctgc ctttaataaa 660aaaaacaaac acatgaatgc agacaccgat tactccatcg cagaagctgc ctttaataaa 660
ggcgaaacag cgatgaccat caacggcccg tgggcatggt ccaacatcga caccagcaaa 720ggcgaaacag cgatgaccat caacggcccg tgggcatggt ccaacatcga caccagcaaa 720
gtgaattatg gtgtaacggt actgccgacc ttcaagggtc aaccatccaa accgttcgtt 780gtgaattatg gtgtaacggt actgccgacc ttcaagggtc aaccatccaa accgttcgtt 780
ggcgtgctga gcgcaggtat taacgccgcc agtccgaaca aagagctggc aaaagagttc 840ggcgtgctga gcgcaggtat taacgccgcc agtccgaaca aagagctggc aaaagagttc 840
ctcgaaaact atctgctgac tgatgaaggt ctggaagcgg ttaataaaga caaaccgctg 900ctcgaaaact atctgctgac tgatgaaggt ctggaagcgg ttaataaaga caaaccgctg 900
ggtgccgtag cgctgaagtc ttacgaggaa gagttggcga aagatccacg tattgccgcc 960ggtgccgtag cgctgaagtc ttacgaggaa gagttggcga aagatccacg tattgccgcc 960
actatggaaa acgcccagaa aggtgaaatc atgccgaaca tcccgcagat gtccgctttc 1020actatggaaa acgcccagaa aggtgaaatc atgccgaaca tcccgcagat gtccgctttc 1020
tggtatgccg tgcgtactgc ggtgatcaac gccgccagcg gtcgtcagac tgtcgatgaa 1080tggtatgccg tgcgtactgc ggtgatcaac gccgccagcg gtcgtcagac tgtcgatgaa 1080
gccctgaaag acgcgcagac taattcgagc tcgaacaaca acaacaataa caataacaac 1140gccctgaaag acgcgcagac taattcgagc tcgaacaaca acaacaataa caataacaac 1140
aacctcggga tcgagggaag gatttcagaa ttcatgtggc gtccgtctga ctctaccgtt 1200aacctcggga tcgagggaag gatttcagaa ttcatgtggc gtccgtctga ctctaccgtt 1200
tacgttccgc cgccgaaccc ggtttctaaa gttgttgcta ccgacgctta cgttacccgt 1260tacgttccgc cgccgaaccc ggtttctaaa gttgttgcta ccgacgctta cgttacccgt 1260
accaacatct tctaccacgc ttcttcttct cgtctgctgg ctgttggtca cccgtacttc 1320accaacatct tctaccacgc ttcttcttct cgtctgctgg ctgttggtca cccgtacttc 1320
tctatcaaac gtgctaacaa aaccgttgtt ccgaaagttt ctggttacca gtaccgtgtt 1380tctatcaaac gtgctaacaa aaccgttgtt ccgaaagttt ctggttacca gtaccgtgtt 1380
ttcaaagttg ttctgccgga cccgaacaaa ttcgctctgc cggactcttc tctgttcgac 1440ttcaaagttg ttctgccgga cccgaacaaa ttcgctctgc cggactcttc tctgttcgac 1440
ccgaccaccc agcgtctggt ttgggcttgc accggtctgg aagttggtcg tggtcagccg 1500ccgaccacccc agcgtctggt ttgggcttgc accggtctgg aagttggtcg tggtcagccg 1500
ctgggtgttg gtgtttctgg tcacccgttc ctgaacaaat acgacgacgt tgaaaactct 1560ctgggtgttg gtgtttctgg tcacccgttc ctgaacaaat acgacgacgt tgaaaactct 1560
ggttctggtg gtaacccggg tcaggacaac cgtgttaacg ttggtatgga ctacaaacag 1620ggttctggtg gtaacccggg tcaggacaac cgtgttaacg ttggtatgga ctacaaacag 1620
acccagctgt gcatggttgg ttgcgctccg ccgctgggtg aacactgggg taaaggtaaa 1680accccagctgt gcatggttgg ttgcgctccg ccgctgggtg aacactgggg taaaggtaaa 1680
cagtgcacca acaccccggt tcaggctggt gactgcccgc cgctggaact gatcacctct 1740cagtgcacca acaccccggt tcaggctggt gactgcccgc cgctggaact gatcacctct 1740
gttatccagg acggtgacat ggttgacacc ggtttcggtg ctatgaactt cgctgacctg 1800gttatccagg acggtgacat ggttgacacc ggtttcggtg ctatgaactt cgctgacctg 1800
cagaccaaca aatctgacgt tccgatcgac atctgcggta ccacctgcaa atacccggac 1860cagaccaaca aatctgacgt tccgatcgac atctgcggta ccacctgcaa atacccggac 1860
tacctgcaga tggctgctga cccgtacggt gaccgtctgt tcttcttcct gcgtaaagaa 1920tacctgcaga tggctgctga cccgtacggt gaccgtctgt tcttcttcct gcgtaaagaa 1920
cagatgttcg ctcgtcactt cttcaaccgt gctggtgaag ttggtgaacc ggttccggac 1980cagatgttcg ctcgtcactt cttcaaccgt gctggtgaag ttggtgaacc ggttccggac 1980
accctgatca tcaaaggttc tggtaaccgt acctctgttg gttcttctat ctacgttaac 2040accctgatca tcaaaggttc tggtaaccgt acctctgttg gttcttctat ctacgttaac 2040
accccgtctg gttctctggt ttcttctgaa gctcagctgt tcaacaaacc gtactggctg 2100accccgtctg gttctctggt ttcttctgaa gctcagctgt tcaacaaacc gtactggctg 2100
cagaaagctc agggtcacaa caacggtatc tgctggggta accagctgtt cgttaccgtt 2160cagaaagctc agggtcacaa caacggtatc tgctggggta accagctgtt cgttaccgtt 2160
gttgacacca cccgttctac caacatgacc ctgtgcgctt ctgttaccac ctcttctacc 2220gttgacacca cccgttctac caacatgacc ctgtgcgctt ctgttaccac ctcttctacc 2220
tacaccaact ctgactacaa agaatacatg cgtcacgttg aagaatacga cctgcagttc 2280tacaccaact ctgactacaa agaatacatg cgtcacgttg aagaatacga cctgcagttc 2280
atcttccagc tgtgctctat caccctgtct gctgaagtta tggcttacat ccacaccatg 2340atcttccagc tgtgctctat caccctgtct gctgaagtta tggcttacat ccacaccatg 2340
aacccgtctg ttctggaaga ctggaacttc ggtctgtctc cgccgccgaa cggtaccctg 2400aacccgtctg ttctggaaga ctggaacttc ggtctgtctc cgccgccgaa cggtaccctg 2400
gaagacacct accgttacgt tcagtctcag gctatcacct gccagaaacc gaccccggaa 2460gaagacacct accgttacgt tcagtctcag gctatcacct gccagaaacc gaccccggaa 2460
aaagaaaaac cggacccgta caaaaacctg tctttctggg aagttaacct gaaagaaaaa 2520aaagaaaaac cggacccgta caaaaacctg tctttctggg aagttaacct gaaagaaaaa 2520
ttctcttctg aactggacca gtacccgctg ggtcgtaaat tcctgctgca gtctggttac 2580ttctcttctg aactggacca gtacccgctg ggtcgtaaat tcctgctgca gtctggttac 2580
cgtggtcgtt cttctatccg taccggtgtt aaacgtccgg ctgtttct 2628cgtggtcgtt cttctatccg taccggtgtt aaacgtccgg ctgtttct 2628
Claims (3)
Priority Applications (1)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| CN201410685769.1A CN104962567B (en) | 2013-12-03 | 2014-11-25 | 6 type recombinant human papilloma virus-like particle and preparation method thereof |
Applications Claiming Priority (3)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| CN201310635249 | 2013-12-03 | ||
| CN2013106352495 | 2013-12-03 | ||
| CN201410685769.1A CN104962567B (en) | 2013-12-03 | 2014-11-25 | 6 type recombinant human papilloma virus-like particle and preparation method thereof |
Publications (2)
| Publication Number | Publication Date |
|---|---|
| CN104962567A CN104962567A (en) | 2015-10-07 |
| CN104962567B true CN104962567B (en) | 2022-11-08 |
Family
ID=54216657
Family Applications (5)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| CN201410672158.3A Active CN105177025B (en) | 2013-12-03 | 2014-11-22 | 18 type recombinant human papilloma virus-like particle and preparation method thereof |
| CN201410672159.8A Active CN105002190B (en) | 2013-12-03 | 2014-11-22 | 11 type recombinant human papilloma virus-like particle and preparation method thereof |
| CN201410672161.5A Active CN105039358B (en) | 2013-12-03 | 2014-11-22 | 58 type recombinant human papilloma virus-like particle and preparation method thereof |
| CN201410685769.1A Active CN104962567B (en) | 2013-12-03 | 2014-11-25 | 6 type recombinant human papilloma virus-like particle and preparation method thereof |
| CN201410683185.0A Active CN105039359B (en) | 2013-12-03 | 2014-11-25 | 16 type recombinant human papilloma virus-like particle and preparation method thereof |
Family Applications Before (3)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| CN201410672158.3A Active CN105177025B (en) | 2013-12-03 | 2014-11-22 | 18 type recombinant human papilloma virus-like particle and preparation method thereof |
| CN201410672159.8A Active CN105002190B (en) | 2013-12-03 | 2014-11-22 | 11 type recombinant human papilloma virus-like particle and preparation method thereof |
| CN201410672161.5A Active CN105039358B (en) | 2013-12-03 | 2014-11-22 | 58 type recombinant human papilloma virus-like particle and preparation method thereof |
Family Applications After (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| CN201410683185.0A Active CN105039359B (en) | 2013-12-03 | 2014-11-25 | 16 type recombinant human papilloma virus-like particle and preparation method thereof |
Country Status (1)
| Country | Link |
|---|---|
| CN (5) | CN105177025B (en) |
Families Citing this family (13)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN105296521B (en) * | 2015-12-07 | 2020-08-28 | 郑州大学 | Recombinant plasmid for expressing soluble human papilloma virus 16 subtype L1 protein and expression method thereof |
| CN105548420B (en) * | 2015-12-30 | 2019-08-02 | 上海泽润生物科技有限公司 | The detection method of MOPS residual quantity |
| CN107188932B (en) * | 2016-03-15 | 2020-02-11 | 中国医学科学院基础医学研究所 | Truncated human papilloma virus 16 type L1 protein and application thereof |
| CN107188931B (en) * | 2016-03-15 | 2020-02-11 | 中国医学科学院基础医学研究所 | Truncated human papilloma virus 58 type L1 protein and application thereof |
| CN109952372A (en) * | 2016-09-16 | 2019-06-28 | 白血球保健股份有限公司 | Obtain the new method of the composition based on viral vectors effective for vaccine inoculation or gene therapy |
| CN108239652B (en) * | 2016-12-27 | 2020-03-17 | 天津大学 | Zika virus capsid protein rigid fusion expression vector and construction and application thereof |
| CN107488747A (en) * | 2017-09-14 | 2017-12-19 | 温州美众医学检验所 | Quantitative fluorescent PCR primer and probe combinations and its reaction system |
| WO2021013063A1 (en) * | 2019-07-19 | 2021-01-28 | 神州细胞工程有限公司 | Chimeric human papillomavirus type 16 l1 protein |
| CN114539363B (en) * | 2020-11-26 | 2023-12-01 | 中国医学科学院基础医学研究所 | C-terminal modified human papilloma virus 11 type L1 protein and application thereof |
| CN114681602B (en) * | 2020-12-25 | 2023-12-01 | 中国食品药品检定研究院 | Bivalent human papillomavirus vaccine |
| CN113549634B (en) * | 2021-06-07 | 2023-03-31 | 郑州大学 | Gene for coding soluble HPV58L1 protein and construction and application of recombinant plasmid thereof |
| CN116023446A (en) * | 2022-10-28 | 2023-04-28 | 北京康乐卫士生物技术股份有限公司 | Expression of human papillomavirus HPV68 L1 protein, virus-like particles and preparation method thereof |
| CN116200416B (en) * | 2023-02-15 | 2024-03-12 | 北京康乐卫士生物技术股份有限公司 | Construction and use of a plasmid expression vector based on Tac promoter |
Citations (3)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN1259051A (en) * | 1997-04-08 | 2000-07-05 | 麦克公司 | Stabilized human papillomavirus formulations |
| CN101016542A (en) * | 2007-02-14 | 2007-08-15 | 马润林 | Method of increasing yield of human papilloma virus L1 albumen pronucleus expression |
| CN101343315A (en) * | 2007-05-29 | 2009-01-14 | 厦门大学 | truncated human papillomavirus type 6 L1 protein |
Family Cites Families (15)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| EP1386611A4 (en) * | 2001-04-19 | 2009-05-13 | Teika Pharmaceutical Co Ltd | MEDICINES AND MEDICAL KITS |
| CN101153280B (en) * | 2006-09-29 | 2015-08-19 | 厦门大学 | The method of purifying human papilloma virus advanced protein L1 from prokaryotic organism |
| CN101245099A (en) * | 2007-02-14 | 2008-08-20 | 马润林 | Amino acid sequence of recombined human papilloma virus L1 capsid protein and uses thereof |
| DK2147926T3 (en) * | 2007-04-29 | 2016-11-28 | Beijing Wantai Biological Pharmacy Entpr Co Ltd | Truncated human papillomavirus type 18 L1 protein |
| BRPI0811016B1 (en) * | 2007-04-29 | 2021-09-21 | Xiamen Innovax Biotech Co., Ltd. | LI TRUNCATE PROTEIN FROM HUMAN PAPILOMA VIRUS TYPE 16 |
| DK2907821T3 (en) * | 2007-05-29 | 2017-04-10 | Univ Xiamen | Process for Preparation of an N-Terminal Truncated L1 Protein of Human Papillomavirus (HPV) |
| CN101481407A (en) * | 2008-01-07 | 2009-07-15 | 马润林 | Modification sequence of recombinant human mammilla tumor virus L1 capsid protein |
| CN101735310B (en) * | 2008-11-20 | 2012-07-18 | 中国疾病预防控制中心病毒病预防控制所 | Human papilloma virus (HPV) fusion protein, gene, carrier, strain, preparation method and application |
| CN106191270B (en) * | 2010-05-05 | 2020-03-03 | 德国癌症研究中心 | Diagnostic transcript and splicing patterns of HR-HPV in different cervical lesions |
| CN101857870B (en) * | 2010-06-12 | 2012-09-05 | 上海泽润生物科技有限公司 | HPV58 L1 gene, vector, strain and expression method |
| US9382296B2 (en) * | 2010-07-16 | 2016-07-05 | Xiamen University | Truncated L1 protein of human papillomavirus type 58 |
| WO2012158639A2 (en) * | 2011-05-14 | 2012-11-22 | Sanofi Pasteur Biologics, Llc | Recombinant fusion proteins and methods for use thereof for treatment or prevention of papillomavirus infection |
| CN103864936B (en) * | 2012-12-11 | 2018-01-23 | 中国疾病预防控制中心病毒病预防控制所 | HPV18 type L2NE7E6 antigen-4 fusion protein genes, expression vector, method, bacterial strain and purposes |
| CN104045696B (en) * | 2012-12-18 | 2018-10-19 | 北京康乐卫士生物技术股份有限公司 | L 1 Protein of Human Papillomavirus Type 16 of recombination and application thereof |
| CN104045695B (en) * | 2012-12-19 | 2019-01-25 | 北京康乐卫士生物技术股份有限公司 | Recombinant human papillomavirus type 18 L1 protein and use thereof |
-
2014
- 2014-11-22 CN CN201410672158.3A patent/CN105177025B/en active Active
- 2014-11-22 CN CN201410672159.8A patent/CN105002190B/en active Active
- 2014-11-22 CN CN201410672161.5A patent/CN105039358B/en active Active
- 2014-11-25 CN CN201410685769.1A patent/CN104962567B/en active Active
- 2014-11-25 CN CN201410683185.0A patent/CN105039359B/en active Active
Patent Citations (3)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN1259051A (en) * | 1997-04-08 | 2000-07-05 | 麦克公司 | Stabilized human papillomavirus formulations |
| CN101016542A (en) * | 2007-02-14 | 2007-08-15 | 马润林 | Method of increasing yield of human papilloma virus L1 albumen pronucleus expression |
| CN101343315A (en) * | 2007-05-29 | 2009-01-14 | 厦门大学 | truncated human papillomavirus type 6 L1 protein |
Also Published As
| Publication number | Publication date |
|---|---|
| CN105002190B (en) | 2022-11-08 |
| CN105177025A (en) | 2015-12-23 |
| CN105039358A (en) | 2015-11-11 |
| CN105177025B (en) | 2022-11-01 |
| CN104962567A (en) | 2015-10-07 |
| CN105039359A (en) | 2015-11-11 |
| CN105039358B (en) | 2020-02-28 |
| CN105002190A (en) | 2015-10-28 |
| CN105039359B (en) | 2020-02-28 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| CN104962567B (en) | 6 type recombinant human papilloma virus-like particle and preparation method thereof | |
| CN101487010A (en) | Method for preparing vaccine for anti-HPV 18 infection by pichia yeast expression system | |
| EP1305039B1 (en) | Stable (fixed) forms of viral l1 capsid proteins, fusion proteins and uses thereof | |
| CN106831958B (en) | A mutant of human papillomavirus type 11 L1 protein | |
| WO2008145021A1 (en) | A truncated l1 protein of human papillomavirus 6 | |
| CN106399329B (en) | 33 type recombinant human papilloma virus-like particle and preparation method thereof | |
| CN103936840B (en) | Recombinant human papillomavirus type 33 L1 protein and use thereof | |
| CN113121704B (en) | Nanoparticle-based coronavirus vaccines | |
| CN110551183A (en) | Mutant of human papilloma virus 39 type L1 protein | |
| CN106701797B (en) | Type 31 recombinant human papillomavirus virus-like particle and preparation method thereof | |
| CN106701796B (en) | 52 type recombinant human papilloma virus-like particle and preparation method thereof | |
| CN103992395B (en) | Recombinate the VLP vaccines and preparation method thereof of HPV-58 types L1 | |
| CN104045696B (en) | L 1 Protein of Human Papillomavirus Type 16 of recombination and application thereof | |
| CN103819543B (en) | 6 type L1 albumen of human papilloma virus of recombination and application thereof | |
| CN106701798B (en) | Type 45 recombinant human papillomavirus virus-like particle and preparation method thereof | |
| SG183776A1 (en) | Improved methods for protein production | |
| CN104045695B (en) | Recombinant human papillomavirus type 18 L1 protein and use thereof | |
| CN115073563B (en) | A GP234 recombinant protein, a gene encoding the same, a production vector, a preparation method and applications thereof | |
| CN110551186A (en) | Mutant of human papilloma virus 45 type L1 protein | |
| CN110551182A (en) | Mutant of human papilloma virus 18 type L1 protein | |
| CN110551181A (en) | mutant of human papilloma virus 66 type L1 protein | |
| CN103214561B (en) | Human hepatitis c virus core antigen and preparation method and application thereof | |
| JP2023530095A (en) | Methods of making virus-like particles | |
| LV15007B (en) | Production of potato pvm virus-like particles |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| C06 | Publication | ||
| PB01 | Publication | ||
| C10 | Entry into substantive examination | ||
| SE01 | Entry into force of request for substantive examination | ||
| GR01 | Patent grant | ||
| GR01 | Patent grant | ||
| PP01 | Preservation of patent right |
Effective date of registration: 20251212 Granted publication date: 20221108 |
|
| PP01 | Preservation of patent right |