CN102435665A - Serum tumor marker for early diagnosis of pancreatic cancer, detection method and diagnosis model - Google Patents
Serum tumor marker for early diagnosis of pancreatic cancer, detection method and diagnosis model Download PDFInfo
- Publication number
- CN102435665A CN102435665A CN2011102863859A CN201110286385A CN102435665A CN 102435665 A CN102435665 A CN 102435665A CN 2011102863859 A CN2011102863859 A CN 2011102863859A CN 201110286385 A CN201110286385 A CN 201110286385A CN 102435665 A CN102435665 A CN 102435665A
- Authority
- CN
- China
- Prior art keywords
- protein
- early diagnosis
- cancer
- tumor markers
- blood serum
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Pending
Links
- 210000002966 serum Anatomy 0.000 title claims abstract description 50
- 206010061902 Pancreatic neoplasm Diseases 0.000 title claims abstract description 43
- 208000015486 malignant pancreatic neoplasm Diseases 0.000 title claims abstract description 43
- 238000013399 early diagnosis Methods 0.000 title claims abstract description 25
- 238000001514 detection method Methods 0.000 title claims abstract description 18
- 208000008443 pancreatic carcinoma Diseases 0.000 title abstract description 15
- 238000003745 diagnosis Methods 0.000 title abstract description 10
- 201000002528 pancreatic cancer Diseases 0.000 title abstract description 8
- 239000000439 tumor marker Substances 0.000 title abstract description 5
- 102000004169 proteins and genes Human genes 0.000 claims abstract description 53
- 108090000623 proteins and genes Proteins 0.000 claims abstract description 53
- 238000000034 method Methods 0.000 claims abstract description 16
- 206010028980 Neoplasm Diseases 0.000 claims description 35
- 230000035945 sensitivity Effects 0.000 claims description 4
- 230000002068 genetic effect Effects 0.000 claims description 3
- 238000003795 desorption Methods 0.000 claims description 2
- 238000000756 surface-enhanced laser desorption--ionisation time-of-flight mass spectrometry Methods 0.000 abstract description 5
- 238000004393 prognosis Methods 0.000 abstract description 4
- 230000007812 deficiency Effects 0.000 abstract description 2
- 238000003384 imaging method Methods 0.000 abstract description 2
- 238000012706 support-vector machine Methods 0.000 description 10
- WEVYAHXRMPXWCK-UHFFFAOYSA-N Acetonitrile Chemical compound CC#N WEVYAHXRMPXWCK-UHFFFAOYSA-N 0.000 description 6
- 230000000875 corresponding effect Effects 0.000 description 6
- 238000001819 mass spectrum Methods 0.000 description 6
- 102000001708 Protein Isoforms Human genes 0.000 description 4
- 108010029485 Protein Isoforms Proteins 0.000 description 4
- 239000012148 binding buffer Substances 0.000 description 4
- 238000002474 experimental method Methods 0.000 description 4
- 238000000926 separation method Methods 0.000 description 4
- 238000000672 surface-enhanced laser desorption--ionisation Methods 0.000 description 4
- XLYOFNOQVPJJNP-UHFFFAOYSA-N water Chemical compound O XLYOFNOQVPJJNP-UHFFFAOYSA-N 0.000 description 4
- 125000003275 alpha amino acid group Chemical group 0.000 description 3
- 201000011510 cancer Diseases 0.000 description 3
- 238000006243 chemical reaction Methods 0.000 description 3
- 238000010828 elution Methods 0.000 description 3
- 239000012634 fragment Substances 0.000 description 3
- 239000007788 liquid Substances 0.000 description 3
- 230000008569 process Effects 0.000 description 3
- 238000011160 research Methods 0.000 description 3
- 239000002904 solvent Substances 0.000 description 3
- 102000004506 Blood Proteins Human genes 0.000 description 2
- 108010017384 Blood Proteins Proteins 0.000 description 2
- 241000283707 Capra Species 0.000 description 2
- 102100025942 Chemokine-like protein TAFA-5 Human genes 0.000 description 2
- 101000788164 Homo sapiens Chemokine-like protein TAFA-5 Proteins 0.000 description 2
- VMHLLURERBWHNL-UHFFFAOYSA-M Sodium acetate Chemical compound [Na+].CC([O-])=O VMHLLURERBWHNL-UHFFFAOYSA-M 0.000 description 2
- 238000004458 analytical method Methods 0.000 description 2
- 239000003153 chemical reaction reagent Substances 0.000 description 2
- 238000004140 cleaning Methods 0.000 description 2
- 238000011161 development Methods 0.000 description 2
- 230000000694 effects Effects 0.000 description 2
- 238000002156 mixing Methods 0.000 description 2
- 238000012544 monitoring process Methods 0.000 description 2
- 210000000496 pancreas Anatomy 0.000 description 2
- 238000002360 preparation method Methods 0.000 description 2
- 238000012216 screening Methods 0.000 description 2
- 235000017281 sodium acetate Nutrition 0.000 description 2
- 239000001632 sodium acetate Substances 0.000 description 2
- 238000012360 testing method Methods 0.000 description 2
- UMCMPZBLKLEWAF-BCTGSCMUSA-N 3-[(3-cholamidopropyl)dimethylammonio]propane-1-sulfonate Chemical compound C([C@H]1C[C@H]2O)[C@H](O)CC[C@]1(C)[C@@H]1[C@@H]2[C@@H]2CC[C@H]([C@@H](CCC(=O)NCCC[N+](C)(C)CCCS([O-])(=O)=O)C)[C@@]2(C)[C@@H](O)C1 UMCMPZBLKLEWAF-BCTGSCMUSA-N 0.000 description 1
- 102000002659 Amyloid Precursor Protein Secretases Human genes 0.000 description 1
- 108010043324 Amyloid Precursor Protein Secretases Proteins 0.000 description 1
- 102400000269 Amyloid protein A Human genes 0.000 description 1
- 101710144835 Amyloid protein A Proteins 0.000 description 1
- 206010006187 Breast cancer Diseases 0.000 description 1
- 208000026310 Breast neoplasm Diseases 0.000 description 1
- 102100040530 CKLF-like MARVEL transmembrane domain-containing protein 1 Human genes 0.000 description 1
- 101710165701 CKLF-like MARVEL transmembrane domain-containing protein 1 Proteins 0.000 description 1
- 102000018832 Cytochromes Human genes 0.000 description 1
- 108010052832 Cytochromes Proteins 0.000 description 1
- 241001269238 Data Species 0.000 description 1
- ROHFNLRQFUQHCH-YFKPBYRVSA-N L-leucine Chemical compound CC(C)C[C@H](N)C(O)=O ROHFNLRQFUQHCH-YFKPBYRVSA-N 0.000 description 1
- ROHFNLRQFUQHCH-UHFFFAOYSA-N Leucine Natural products CC(C)CC(N)C(O)=O ROHFNLRQFUQHCH-UHFFFAOYSA-N 0.000 description 1
- 206010058467 Lung neoplasm malignant Diseases 0.000 description 1
- 206010027336 Menstruation delayed Diseases 0.000 description 1
- 102000013566 Plasminogen Human genes 0.000 description 1
- 108010051456 Plasminogen Proteins 0.000 description 1
- 208000005718 Stomach Neoplasms Diseases 0.000 description 1
- XSQUKJJJFZCRTK-UHFFFAOYSA-N Urea Chemical compound NC(N)=O XSQUKJJJFZCRTK-UHFFFAOYSA-N 0.000 description 1
- 238000010521 absorption reaction Methods 0.000 description 1
- 230000004308 accommodation Effects 0.000 description 1
- 238000013459 approach Methods 0.000 description 1
- 238000003766 bioinformatics method Methods 0.000 description 1
- 210000004369 blood Anatomy 0.000 description 1
- 239000008280 blood Substances 0.000 description 1
- 210000001124 body fluid Anatomy 0.000 description 1
- 239000010839 body fluid Substances 0.000 description 1
- 210000004899 c-terminal region Anatomy 0.000 description 1
- 239000004202 carbamide Substances 0.000 description 1
- 210000004027 cell Anatomy 0.000 description 1
- 230000008859 change Effects 0.000 description 1
- 230000035572 chemosensitivity Effects 0.000 description 1
- 230000000295 complement effect Effects 0.000 description 1
- 238000007796 conventional method Methods 0.000 description 1
- 238000012937 correction Methods 0.000 description 1
- 230000002596 correlated effect Effects 0.000 description 1
- 238000002790 cross-validation Methods 0.000 description 1
- 239000008367 deionised water Substances 0.000 description 1
- 229910021641 deionized water Inorganic materials 0.000 description 1
- 238000010586 diagram Methods 0.000 description 1
- 201000010099 disease Diseases 0.000 description 1
- 208000037265 diseases, disorders, signs and symptoms Diseases 0.000 description 1
- 229940079593 drug Drugs 0.000 description 1
- 238000001914 filtration Methods 0.000 description 1
- 239000012467 final product Substances 0.000 description 1
- 238000004108 freeze drying Methods 0.000 description 1
- 238000002290 gas chromatography-mass spectrometry Methods 0.000 description 1
- 206010017758 gastric cancer Diseases 0.000 description 1
- 230000036541 health Effects 0.000 description 1
- 238000004128 high performance liquid chromatography Methods 0.000 description 1
- 238000000265 homogenisation Methods 0.000 description 1
- 210000003734 kidney Anatomy 0.000 description 1
- 210000002429 large intestine Anatomy 0.000 description 1
- 238000004895 liquid chromatography mass spectrometry Methods 0.000 description 1
- 238000001294 liquid chromatography-tandem mass spectrometry Methods 0.000 description 1
- 201000005202 lung cancer Diseases 0.000 description 1
- 208000020816 lung neoplasm Diseases 0.000 description 1
- 230000036210 malignancy Effects 0.000 description 1
- 239000003550 marker Substances 0.000 description 1
- 238000001840 matrix-assisted laser desorption--ionisation time-of-flight mass spectrometry Methods 0.000 description 1
- 230000002438 mitochondrial effect Effects 0.000 description 1
- 238000012986 modification Methods 0.000 description 1
- 230000004048 modification Effects 0.000 description 1
- 201000010879 mucinous adenocarcinoma Diseases 0.000 description 1
- 238000012856 packing Methods 0.000 description 1
- 230000007170 pathology Effects 0.000 description 1
- 230000007310 pathophysiology Effects 0.000 description 1
- 230000002093 peripheral effect Effects 0.000 description 1
- 230000002980 postoperative effect Effects 0.000 description 1
- 238000012545 processing Methods 0.000 description 1
- 238000000746 purification Methods 0.000 description 1
- 239000002994 raw material Substances 0.000 description 1
- 238000010183 spectrum analysis Methods 0.000 description 1
- 238000012109 statistical procedure Methods 0.000 description 1
- 210000002784 stomach Anatomy 0.000 description 1
- 201000011549 stomach cancer Diseases 0.000 description 1
- 239000006228 supernatant Substances 0.000 description 1
- 230000004083 survival effect Effects 0.000 description 1
- 238000004885 tandem mass spectrometry Methods 0.000 description 1
- 230000009466 transformation Effects 0.000 description 1
- 210000003462 vein Anatomy 0.000 description 1
Images
Landscapes
- Investigating Or Analysing Biological Materials (AREA)
Abstract
The invention belongs to the field of medical diagnosis, and particularly relates to a serum tumor marker for early diagnosis of pancreatic cancer, which consists of 11 protein mass-to-charge ratio peaks: 6684Da, 6668Da, 8591Da, 6471Da, 4121Da, 8775Da, 4290Da, 6655Da, 2959Da, 5913Da and 5346 Da. The invention also provides a detection method and a diagnosis model of the serum tumor marker for early diagnosis of pancreatic cancer. The invention utilizes SELDI-TOF-MS technology to detect the combined tumor marker from a low-abundance serum sample, establishes a protein fingerprint diagnosis model for early diagnosis of pancreatic cancer, provides a new way and a new method for early diagnosis of pancreatic cancer patients, makes up for the deficiency of imaging, is expected to improve the early diagnosis level of pancreatic cancer, and is helpful for improving the prognosis of pancreatic cancer patients.
Description
Technical field
The invention belongs to area of medical diagnostics, particularly to cancer of pancreas early diagnosis sensitivity, blood serum tumor markers, detection method and protein fingerprint figure diagnostic model that specificity is high.
Background technology
Cancer of pancreas is one of tumour that grade malignancy is the highest, prognosis is the poorest, and cancer of pancreas morbidity rate and case fatality rate rise rapidly, and age of onset has caused common concern gradually in advance.Because lack good tumor-marker object detecting method clinically, most Pancreas cancer patients has been a middle and advanced stage when making a definite diagnosis.Early diagnosis, early treatment are the keys that improves the cancer of pancreas survival rate and reduce mortality ratio.But at present, the cancer of pancreas early diagnosis lacks effective means, and at present main means are leaned on imaging examination, but the result is unsatisfactory.Stage in early days, especially subclinical stage CT, MR are then powerless.The diagnosis of tumor markers is the focus of Recent study.Yet single tumor markers often is difficult to reflect the overall picture of tumour, and theoretically, what most possibly become desirable tumor markers should be several or a plurality of tumor markerses of associating.
Summary of the invention
Problem to above-mentioned prior art existence; In order to improve cancer of pancreas early diagnosis level; Thereby help to improve the prognosis of Pancreas cancer patients; The object of the invention is intended to set up the protein fingerprint figure diagnostic model of cancer of pancreas early diagnosis, further can filter out the new body fluid protein tumor markers of cancer of pancreas early diagnosis.
The objective of the invention is to be able to implement through following technical scheme:
The present invention at first provides the blood serum tumor markers of cancer of pancreas early diagnosis, and described blood serum tumor markers is made up of 11 protein mass-to-charge ratio peaks: 6684Da, 6668Da, 8591Da, 6471Da, 4121Da, 8775Da, 4290Da, 6655Da, 2959Da, 5913Da and 5346Da.
The present invention also provides a kind of detection method of blood serum tumor markers of cancer of pancreas early diagnosis, comprising:
(1) serum is prepared,
(2) mass spectrometric data is collected, and filters out the differential protein peak, sets up the protein fingerprint figure diagnostic model of cancer of pancreas early diagnosis,
(3) further filter out blood serum tumor markers with the method for genetic algorithm combination supporting vector machine model.
As preferred version; According to the detection method of the blood serum tumor markers of a kind of cancer of pancreas early diagnosis of the present invention, wherein, in the described step (2): mass spectrometric data is collected laser intensity is set is 180; Detection sensitivity is 6; Upper limit of detection is 100000m/z, and collecting data area is 2000-20000 m/z, and the signal collection position is from 20-80.
The present invention also provides a kind of protein fingerprint figure diagnostic model of blood serum tumor markers of cancer of pancreas early diagnosis; Wherein, Described blood serum tumor markers is made up of 11 protein mass-to-charge ratio peaks: 6684Da, 6668Da, 8591Da, 6471Da, 4121Da, 8775Da, 4290Da, 6655Da, 2959Da, 5913Da and 5346Da
Described protein fingerprint figure diagnostic model is set up as follows:
(1) utilize the surface to strengthen the proteomic map that laser desorption ionisation-time of-flight mass spectrometer is measured tumor patient and healthy subjects serum specimen,
(2) combine the method for bioinformatics to filter out corresponding blood serum tumor markers and set up diagnostic model.
The present invention compared with prior art has the following advantages:
(1) utilize the SELDI-TOF-MS technology from low-abundance serum sample, to detect blood serum tumor markers.
(2) unite the candidate's who in serum, screens tumor markers, new approaches and methods is provided, remedied the deficiency of iconography the early diagnosis of Pancreas cancer patients.The iconography development of technology, Pancreas cancer patients diagnosis aspect has obtained certain breakthrough in early days.
The bioinformatics Development of Software and the application such as SVMs of (3) being correlated with improve accurate rate of diagnosis.
(4) the present invention is expected to improve cancer of pancreas early diagnosis level, thereby helps to improve the prognosis of Pancreas cancer patients from the associating protein tumor markers of a brand-new technology platform and theory detection cancer of pancreas.
Figure of description
Fig. 1 is the schematic flow sheet of experimental technique of the present invention.
Fig. 2 is that the mass spectrum protein peak that progressively increases of all samples of the present invention shows the intention that distributes.
Fig. 3 is an early stage cancer of pancreas group and the diffusing point of two groups of sample SVMs of a healthy subjects control group distribution plan; Among the figure; Ordinate principal component representes major component; Horizontal ordinate SVM predict labels expresses support for the vector machine predicted value, and SVM Result Scatter Plot expresses support for the diffusing point of a vector machine distribution results.
Fig. 4 is the mass spectrum expression figure of all samples.
Fig. 5 is that human pancreas cancer serum sample and corresponding goat anti-human antibody reaction are spent the night after SELDI testing result figure.
Embodiment
Below in conjunction with embodiment, content of the present invention is described more specifically.Should be appreciated that enforcement of the present invention is not limited to following embodiment, all will fall into protection domain of the present invention any pro forma accommodation and/or the change that the present invention made.
In the present invention, if not refer in particular to, all part, number percents are unit of weight, and all equipment and raw material etc. all can be buied from market or the industry is commonly used.Method among the following embodiment if no special instructions, is the conventional method of this area.
1, sample and data
Pancreas cancer patients: totally 54 examples (early stage 22, late period 32), postoperative pathology is made a definite diagnosis, 65 years old mean age.
The normal healthy controls group: totally 20 examples, derive from health check-up, the mean age is 63 years old.
Be divided into two groups of protein spectrum analysis contrasts of carrying out serum:
1. early stage cancer of pancreas group vs healthy subjects control group,
2. early stage cancer of pancreas vs advanced pancreatic cancer group.
Each organizes the age and sex is all matched; Similar exposure history is arranged, does not have other relevant diseases influence protein content in the serum, gather healthy subjects and patient treatment on an empty stomach early morning before peripheral vein whole blood 5ml; 4 ℃ leave standstill 1-2h; Centrifugal 10min under the 3000 r/min conditions, separation of serum ,-80 ℃ of refrigerators are preserved subsequent use.
2, experimental technique
Key instrument equipment and reagent:
L Ciphergen PBS-II-PLUS type SELDI-TOF-MS system
L protein-chip: CM10 chip
The preparation of experiment reagent:
1.U9 serum denaturing soln: 9 mol/l urea, 2 % CHAPS, 1 % DTT.Packing after preparation is accomplished ,-80 ℃ of low temperature refrigerators are preserved.
2.50% saturated SPA solution: the solvent of SPA is 50% H
2O+50% acetonitrile+0.5% TFA.SPA is dissolved in its solvent until supersaturation, 13,000rpm gets supernatant after centrifugal 1 minute, mixes getting final product with solvent 1:1.Room temperature keeps in Dark Place.
3.CM10 chip Binding Buffer:50 mmol/l sodium acetate (pH=4.0).Sodium acetate 2.051g is dissolved in the deionized water, and HCl regulates pH value to 4.0, and constant volume is 500ml.
Experimental technique(process flow diagram is seen accompanying drawing 1)
The pre-service of chip
Add 200 μ l Binding Buffer in the every hole of l
Remove liquid after vibration (600rpm, room temperature) made abundant balance in 5 minutes on the l shaking table, repeat balance once.
The pre-service of serum
Slowly melt serum (30~60 minutes) on the l ice bath, centrifugal 5 minutes of 10000rpm (4 ℃).
Add serum 5 μ L in the l 10 μ l U9 serum denaturing solns, vibration makes abundant mixing (600rpm, 4 ℃, 30 minutes) on the shaking table.
Serum after l handles U9 with Binding Buffer is diluted to 200 μ l and is used for appearance, and vibration makes abundant mixing (600rpm, 4 ℃, 2 minutes) on the shaking table.So far serum is diluted about 40 times.
Protein and chips incorporate
L gets and above-mentionedly handles and the serum 100 μ l that diluted are added on the chip through U9, and scrutiny is also removed the bubble in each hole in order to avoid it influences combining of protein and chip.
L chip and serum fully react 1 hour (vibrating 4 ℃ on the 600rpm shaking table) back and remove sample.
In conjunction with post-flush
The every hole of l adds the corresponding Binding Buffer of 200 μ l, and vibration was removed liquid after (600rpm, room temperature) in 5 minutes on the shaking table.Repeat this process 2 times.
L water (purity HPLC level) 200 each hole of μ l Rapid Cleaning 1 time firmly dry and Bio-processor are tipped upside down on the thieving paper of cleaning and pat to remove unnecessary water.
L takes out chip from Bio-processor, air dry.
Add energy absorption molecule (EAM)
Every hole point adds 50% saturated SPA solution 1 μ l, treats to repeat a little to add 1 time after its air dry.Air dry can be gone up machine testing.
Data acquisition and statistical procedures
The parameter setting of SELDI mass spectrometer and raw data acquisition and output
At first with the standard protein chip All-in-One of known molecular amount with the molecular weight error correction of SELDI mass spectrometer system to < 0.1%, the chip of the protein that will be combined is again put into mass spectrometer and detected.Raw data is by Proteinchip Software 3.2 software collections, and it is 180 that laser intensity is set, detection sensitivity 6, upper limit of detection 100,000
M/z, optimize and collect data area 2000~20000
M/z, the signal collection position is from 20~80, and each sample is got the mean value of 168 collected signals of point.With Proteinchip Software 3.2 softwares raw data is exported with the form of xlm.
Data processing
Adopt ZJU-PDAS (ProteinChip Data Analyze System) software analysis, whole flow process is following:
1. raw mass spectrum figure uploads onto the server,
2. remove the noise that mass spectrometer itself causes with wavelet transformation (UDWT undecimated discrete wavelet transform) earlier,
3. revise to remove the baseline of the mass spectrogram behind the noise,
4. proofread and correct the molecular weight values of whole collection of illustrative plates,
5. find out albumen value peak with the local extremum method, the ratio filtration protein peak that in each sample, occurs with signal to noise ratio (S/N ratio) and this peak,
6. all sample datas of homogenization,
7. the intact protein peak that screens of pre-service is done further check analysis, filter out P 0.05 differential protein peak,
8. the differential protein peak to screening further screens best model with the method for genetic algorithm combination supporting vector machine model; Prediction effect with the leaving-one method assessment models; Select the highest combination of the youden index of setting up supporting vector machine model prediction as final candidate markers, the result of the model of foundation and leaving-one method cross validation is as final result.
9. export various statisticses and picture.
The result:
1, early stage cancer of pancreas group (positive group) vs healthy subjects control group
Early stage cancer of pancreas groups of 22 examples and 20 routine healthy subjects control groups carry out the t check, and examination p value is less than 0.05 differential protein.Detect 153 altogether through signal to noise ratio (S/N ratio)s and the high-quality mass spectrum protein peaks that stress to filter, find to have 14 protein peak content have significant difference (
P<0.05), therefrom filter out 11 protein peaks and be used to set up the SVM discrimination model.The mass spectrum protein peak that progressively increases of all samples shows that distribution is intended to see Fig. 2.The two groups of diffusing point of sample SVMs distribution plans are seen Fig. 3, and every bit is represented a sample, and ordinate is a major component, and horizontal ordinate is the SVM predicted value, can distinguish clearly for two groups.
Explain: among the present invention in all charts Group0 or Class0 be control group, positive group of Group1 or Class1.
11 peaks of screening are used to set up the mark of SVM (SVMs) model of distinguishing two groups from above-mentioned table 1: 6684.598Da, 6668.8192Da, 8591.8289Da, 6471.9468Da, 4121.3689Da, 8775.5261Da, 4290.4956Da, 6655.6858Da, 2959.1113Da
,5913.514Da with 5346.1731 Da.
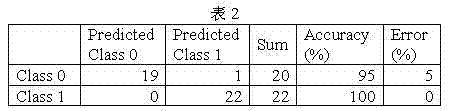
It is following that the result is differentiated in the checking of the best model of setting up:
Can find out that from table 2 specificity of model is 95%, susceptibility is 100%.
Fig. 4 is the mass spectrum expression figure of all samples, and for example: peak 6685.emf is MZ6684.598, and among the figure, horizontal ordinate is a mass-to-charge ratio, and ordinate is an expression intensity, and every line indicates a sample, arrow → and be the position at this peak of MZ6684.598.As can be seen from Figure 4 the MZ6684.598 mark is expressed higher in the healthy subjects control group.
Embodiment 2The responsive special blood serum tumor markers that filters out is separated and identifies, formulate monoclonal antibody
1,5.9kDa (5913.3631) protein equal high expressed in two models, the inventor identifies it.
(1) 5.9kDa (5913.3631) protein peak is identified
With SPE post enrichment protein; The LC-MS separation is also confirmed albumen; Components separated is used trypsinization; It is checked order with MS-MS, this research is expressed the protein (5.9 kDa) that significantly raises in early days in the cancer of pancreas to one and is identified that result is an a kind of fibrinous segment again.Be specially: separatory sample is a 1ml cancer of pancreas early stage patient serum.Serum is earlier with SPE post (anti-phase, acetonitrile gradient) initial gross separation (with the elution component at SELDI-TOF-MS monitoring objective albumen place), with the enrichment of lyophilization appearance.Again the dissolving back is with high performance liquid chromatography-GC-MS (HPLC-MS) separation and purification (with the elution component at MALDI-TOF-MS monitoring objective albumen place); Target elution component is further used trypsinization; It is that 3 peaks of 2040.8,1189.5 and 1139.5 get into the LC/MS/MS order-checking that the component of purifying is chosen molecular weight through trypsinization, and the result of order-checking is following:
Molecular weight 2040.8: qftsstsyn rgdstfesk
Molecular weight 1189.5: qftsstsyn r
Molecular weight 1139.5: gse sgiftntk
The amino acid sequence of these 3 molecular weight segments to MASCOT (MatrixScience, www.matrixscience.com) go up comparison with, all be complementary with a fibrinous segment, its complete sequence is following:
mfsmrivclv?lsvvgtawta?dsgegdflae?gggvrgprvv?erhqsackds?dwpfcsdedw
nykcpsgcrm?kglidevnqd?ftnrinklkn?slfeyqknnk?dshslttnim?eilrgdfssa
nnrdntynrv?sedlrsriev?lkrkviekvq?hiqllqknvr?aqlvdmkrle?vdidikirsc
rgscsralar?evdlkdyedq?qkqleqviak?dllpsrdrqh?lplikmkpvp?dlvpgnfksq
lqkvppewka?ltdmpqmrme?lerpggneit?rggstsygtg?setesprnps?sagswnsgss
gpgstgnrnp?gssgtggtat?wkpgssgpgs?tgswnsgssg?tgstgnqnpg?sprpgstgtw
npgssergsa?ghwtsessvs?gstgqwhses?gsfrpdspgs?gnarpnnpdw?gtfeevsgnv
spgtrreyht?eklvtskgdk?elrtgkekvt?sgsttttrrs?csktvtktvi?gpdghkevtk
evvtsedgsd?cpeamdlgtl?sgigtldgfr?hrhpdeaaff?dtastgktfp?gffspmlgef
vsetesrgse?sgiftntkes?sshhpgiaef?psrgksssys?kqftsstsyn?rgdstfesks
ykmadeagse?adhegthstk?rghaksrpvr?dcddvlqthp?sgtqsgifni?klpgsskifs
vycdqetslg?gwlliqqrmd?gslnfnrtwq?dykrgfgsln?degegefwlg?ndylhlltqr
gsvlrveled?wagneayaey?hfrvgseaeg?yalqvssyeg?tagdaliegs?veegaeytsh
nnmqfstfdr?dadqweenca?evygggwwyn?ncqaanlngi?yypggsydpr?nnspyeieng
vvwvsfrgad?yslravrmki?rplvtq
(2) spend the night with the reaction of corresponding goat anti-human antibody and human pancreas's serum sample after SELDI detects this peak of 5.9kDa obviously disappears, as shown in Figure 5, among the figure 1. be not with the serum of corresponding antibodies reaction, 2. be and the serum of corresponding antibodies after reacting.
2, other blood serum tumor markers are further searched for the UniProtKB/Swiss-Prot database with TagIdent tool instrument, identify other protein respectively, the result is following:
(1). molecular weight: 6667.6807
Protein is called for short CKLF1_HUMAN (Q8IZ96-10)
The amino acid sequence Chain:1-65 of detected protein fragments,
The pI value: 8.19,
Theoretical molecular Mw:6669
(2). molecular weight: 8572.3799
Protein is called for short SAA_HUMAN (P02735)
Protein title Amyloid protein A.
Chain:?19-94,pI:?5.56,Mw:?8575
(3). molecular weight: 2958.761
Protein is called for short CKLF1_HUMAN (Q8IZ96-15)
Protein title CKLF-like MARVEL transmembrane domain-containing protein 1
Chain:?1-28,pI:?4.95,Mw:?2962
(4). molecular weight: 6441.593
LUZP6_HUMAN?(Q538Z0)
Leucine?zipper?protein?6.
Chain:?1-58,?pI:?9.69,?Mw:?6437
(5). molecular weight: 6668.8192
Protein is called for short CKLF1_HUMAN (Q8IZ96-10)
The amino acid sequence Chain:1-65 of detected protein fragments,
The pI value: 8.19,
Theoretical molecular Mw:6669
(6). molecular weight: 8775.5261
PLGA_HUMAN?(Q15195)
Plasminogen-related?protein?A.
Chain:?20-96,?pI:?5.83,?Mw:?8778
(7). molecular weight: 6655.6858
A4_HUMAN?(P05067)
Gamma-secretase?C-terminal?fragment?57.
Chain:?714-770,?pI:?6.74,?Mw:?6651
(8). molecular weight: 2959.1113
Recommended?name:
CKLF-like?MARVEL?transmembrane?domain-containing?protein?1
Alternative?name(s):
Chemokine-like?factor?superfamily?member?1
CKLF1_HUMAN?(Q8IZ96-15)
Isoform?16?of?CKLF-like?MARVEL?transmembrane?domain-conta...
Chain:?1-28,?pI:?4.95,?Mw:?2962
(9). molecular weight: 6119.674
F19A5_HUMAN?(Q7Z5A7-3)
Isoform?3?of?Protein?FAM19A5?OS=Homo?sapiens?GN=FAM19A5
Chain:?1-53,?pI:?8.48,?Mw:?6119
(10). molecular weight: 5346.1731
COX7C_HUMAN?(P15954)
Cytochrome?coxidase?subunit?7C,?mitochondrial.
Chain:?17-63,?pI:?8.97,?Mw:?5356
Above-mentioned preferred embodiment just is used for explanation and explains content of the present invention not constituting the restriction to content of the present invention.Although the inventor has done in more detail the present invention and has enumerated; But; The content that those skilled in the art is disclosed according to summary of the invention part and embodiment can be made various modifications or/and additional or to adopt similar mode to substitute be obvious to described specific embodiment, and can realize technique effect of the present invention; Therefore, give unnecessary details no longer one by one here.The term that occurs among the present invention is used for the elaboration of technical scheme of the present invention and understanding are not construed as limiting the invention.
List of references:
1. thank to gorgeous eating, Huang Jianjin, Yu Jiekai, Li Xufen, fourth is insulted the research of SELDI-TOF-MS technology in detection of lung cancer chemosensitivity related protein, oncology magazine, 2010,16 (8): 610-613.
2. recklessly jump, Yu Jiekai, Zhang Suzhan, Liu builds, the Zheng Shu protein fingerprint pattern breast cancer diagnosis with follow up a case by regular visits in applied research, Chinese Journal of Pathophysiology, 2010,26 (4): 700-704.
3. Korea Spro is little grand, Mao Qiaoxia, and Li Xiaochun, Shen Jianfa, Lu Guofeng, Yu Jiekai is based on the application of serum mark object model in diagnosing gastric cancer of bioinformatics method, Chinese cancer magazine, 2010,20 (5): 364-368.
4. Xu Wen letter, Chen Yiding jumps recklessly, Yu Jiekai, Wang Liancong, Zheng Shu, Zhang Suzhan based on the large intestine mucinous adenocarcinoma art of SVMs before serum proteins marker detection and analysis, cell biology magazine, 2008,30:819 – 822.
5. old Xinzhou, Wang Yujie, Zhang Qiong, Wang Qingrong, the application of the triumphant serum protein fingerprint diagnostic model of Yu Jie in kidney, Xinjiang Medicine University's journal, 2010,33 (2): 141-148.
6. Liu Ying is refined, and the scope of the wide body of pancreas tail of Wu Wen cancer radical excision is inquired into, general outer basis of China and clinical magazine, 20 11,18 (1): 11-13.
Claims (4)
1. the blood serum tumor markers of cancer of pancreas early diagnosis; It is characterized in that described blood serum tumor markers is made up of 11 protein mass-to-charge ratio peaks: 6684Da, 6668Da, 8591Da, 6471Da, 4121Da, 8775Da, 4290Da, 6655Da, 2959Da, 5913Da and 5346Da.
2. the detection method of the blood serum tumor markers of a cancer of pancreas early diagnosis is characterized in that, described detection method comprises:
(1) serum is prepared,
(2) mass spectrometric data is collected, and filters out the differential protein peak, sets up the protein fingerprint figure diagnostic model of cancer of pancreas early diagnosis,
(3) further filter out blood serum tumor markers with the method for genetic algorithm combination supporting vector machine model.
3. the detection method of the blood serum tumor markers of a kind of cancer of pancreas early diagnosis according to claim 2; It is characterized in that; In the described step (2): mass spectrometric data is collected laser intensity is set is 180, and detection sensitivity is 6, and upper limit of detection is 100000m/z; Collecting data area is 2000-20000 m/z, and the signal collection position is from 20-80.
4. the protein fingerprint figure diagnostic model of the blood serum tumor markers of a cancer of pancreas early diagnosis; It is characterized in that; Described blood serum tumor markers is made up of 11 protein mass-to-charge ratio peaks: 6684Da, 6668Da, 8591Da, 6471Da, 4121Da, 8775Da, 4290Da, 6655Da, 2959Da, 5913Da and 5346Da
Described protein fingerprint figure diagnostic model is set up as follows:
(1) utilize the surface to strengthen the proteomic map that laser desorption ionisation-time of-flight mass spectrometer is measured tumor patient and healthy subjects serum specimen,
(2) combine the method for bioinformatics to filter out corresponding blood serum tumor markers and set up diagnostic model.
Priority Applications (1)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| CN2011102863859A CN102435665A (en) | 2011-09-23 | 2011-09-23 | Serum tumor marker for early diagnosis of pancreatic cancer, detection method and diagnosis model |
Applications Claiming Priority (1)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| CN2011102863859A CN102435665A (en) | 2011-09-23 | 2011-09-23 | Serum tumor marker for early diagnosis of pancreatic cancer, detection method and diagnosis model |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| CN102435665A true CN102435665A (en) | 2012-05-02 |
Family
ID=45983830
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| CN2011102863859A Pending CN102435665A (en) | 2011-09-23 | 2011-09-23 | Serum tumor marker for early diagnosis of pancreatic cancer, detection method and diagnosis model |
Country Status (1)
| Country | Link |
|---|---|
| CN (1) | CN102435665A (en) |
Cited By (5)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN105102637A (en) * | 2013-04-17 | 2015-11-25 | Lg电子株式会社 | Method for extracting pancreatic cancer diagnostic biomarkers, computing device used for the method, pancreatic cancer diagnostic biomarkers, and pancreatic cancer diagnostic device comprising the same |
| CN106909781A (en) * | 2017-02-22 | 2017-06-30 | 北京航空航天大学 | A kind of method for optimum option cerebral apoplexy associated biomarkers |
| CN108445097A (en) * | 2017-03-31 | 2018-08-24 | 北京谷海天目生物医学科技有限公司 | Molecular typing of diffuse type gastric cancer, protein marker for typing, screening method and application thereof |
| CN115925864A (en) * | 2015-12-11 | 2023-04-07 | 国立大学法人高知大学 | Markers of pancreatic cancer and intraductal papillary mucinous neoplasms |
| US12435130B2 (en) | 2012-02-15 | 2025-10-07 | Neuracle Science Co., Ltd. | Pharmaceutical use of FAM19A5 involved in regulating gliogenesis |
-
2011
- 2011-09-23 CN CN2011102863859A patent/CN102435665A/en active Pending
Cited By (7)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US12435130B2 (en) | 2012-02-15 | 2025-10-07 | Neuracle Science Co., Ltd. | Pharmaceutical use of FAM19A5 involved in regulating gliogenesis |
| CN105102637A (en) * | 2013-04-17 | 2015-11-25 | Lg电子株式会社 | Method for extracting pancreatic cancer diagnostic biomarkers, computing device used for the method, pancreatic cancer diagnostic biomarkers, and pancreatic cancer diagnostic device comprising the same |
| CN105102637B (en) * | 2013-04-17 | 2018-05-22 | Lg电子株式会社 | Method for extracting pancreatic cancer diagnostic biomarkers, computing device used for the method, pancreatic cancer diagnostic biomarkers, and pancreatic cancer diagnostic device comprising the same |
| CN115925864A (en) * | 2015-12-11 | 2023-04-07 | 国立大学法人高知大学 | Markers of pancreatic cancer and intraductal papillary mucinous neoplasms |
| CN106909781A (en) * | 2017-02-22 | 2017-06-30 | 北京航空航天大学 | A kind of method for optimum option cerebral apoplexy associated biomarkers |
| CN106909781B (en) * | 2017-02-22 | 2019-12-31 | 北京航空航天大学 | A method for optimal selection of stroke-related biomarkers |
| CN108445097A (en) * | 2017-03-31 | 2018-08-24 | 北京谷海天目生物医学科技有限公司 | Molecular typing of diffuse type gastric cancer, protein marker for typing, screening method and application thereof |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| Sigdel et al. | Shotgun proteomics identifies proteins specific for acute renal transplant rejection | |
| Chung et al. | Novel serum protein biomarker panel revealed by mass spectrometry and its prognostic value in breast cancer | |
| EP2310861B1 (en) | Healthy kidney biomarkers | |
| Yang et al. | Identification of candidate biomarkers for the early detection of nasopharyngeal carcinoma by quantitative proteomic analysis | |
| Guipaud | Serum and plasma proteomics and its possible use as detector and predictor of radiation diseases | |
| KR20160045547A (en) | Composition for diagnosing pancreatic cancer and method for diagnosing pancreatic cancer using the same | |
| CN102323351A (en) | Bladder cancer patients urine specific metabolic thing spectrum, method for building up and purposes | |
| CN115678994B (en) | A biomarker combination, a reagent containing the same and its application | |
| CN102435665A (en) | Serum tumor marker for early diagnosis of pancreatic cancer, detection method and diagnosis model | |
| JP2015105951A (en) | Blood-derived peptide marker for cancer diagnosis and method for cancer diagnosis using the same | |
| WO2015117133A1 (en) | Integrated quantification method for protein measurements in clinical proteomics | |
| Gonzalez | Novel biomarkers for hepatocellular carcinoma surveillance: has the future arrived? | |
| EP3092495B1 (en) | Srm assay for pd-l1 | |
| Acharya et al. | Quantitative plasma proteomics identifies metallothioneins as a marker of acute-on-chronic liver failure associated acute kidney injury | |
| EP2850229A1 (en) | Srm/mrm assay for subtyping lung histology | |
| Huang et al. | Liquid chromatography–mass spectrometry based serum peptidomic approach for renal clear cell carcinoma diagnosis | |
| KR20150062915A (en) | Serological markers for cancer diagnosis using blood sample | |
| CN107255711B (en) | Use of osteopontin for preparing or screening diagnostic reagents for acute-on-chronic liver failure | |
| CN110554189A (en) | Pancreatic cancer diagnostic markers and their application | |
| CN108152508A (en) | A kind of screening technique of hurtless measure gastric cancer saliva biomarker | |
| CN113358881B (en) | Biomarker for NMOSD prediction or recurrence monitoring and application thereof | |
| US20050158745A1 (en) | Methods and compositions for detection of nasopharyngeal carcinoma | |
| Li et al. | An integrated platform for decoding hydrophilic peptide fingerprints of hepatocellular carcinoma using artificial intelligence and two-dimensional nanosheets | |
| CN109342738B (en) | Application of group of serum differential protein combinations in preparation of reagent for detecting Alzheimer's disease | |
| CN108334747A (en) | It obtains the method for tumour Urine proteins marker and obtained tumour correlation peels off Urine proteins library |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| C06 | Publication | ||
| PB01 | Publication | ||
| C10 | Entry into substantive examination | ||
| SE01 | Entry into force of request for substantive examination | ||
| C12 | Rejection of a patent application after its publication | ||
| RJ01 | Rejection of invention patent application after publication |
Application publication date: 20120502 |