CN102409045B - Tag library constructing method based on DNA (deoxyribonucleic acid) adapter connection as well as used tag and tag adapter - Google Patents
Tag library constructing method based on DNA (deoxyribonucleic acid) adapter connection as well as used tag and tag adapter Download PDFInfo
- Publication number
- CN102409045B CN102409045B CN 201010299257 CN201010299257A CN102409045B CN 102409045 B CN102409045 B CN 102409045B CN 201010299257 CN201010299257 CN 201010299257 CN 201010299257 A CN201010299257 A CN 201010299257A CN 102409045 B CN102409045 B CN 102409045B
- Authority
- CN
- China
- Prior art keywords
- dna
- index
- tag
- adapter
- tags
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Active
Links
Images
Classifications
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q1/00—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions
- C12Q1/68—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions involving nucleic acids
- C12Q1/6869—Methods for sequencing
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/10—Processes for the isolation, preparation or purification of DNA or RNA
- C12N15/1034—Isolating an individual clone by screening libraries
- C12N15/1065—Preparation or screening of tagged libraries, e.g. tagged microorganisms by STM-mutagenesis, tagged polynucleotides, gene tags
Landscapes
- Life Sciences & Earth Sciences (AREA)
- Chemical & Material Sciences (AREA)
- Health & Medical Sciences (AREA)
- Organic Chemistry (AREA)
- Engineering & Computer Science (AREA)
- Genetics & Genomics (AREA)
- Zoology (AREA)
- Wood Science & Technology (AREA)
- Proteomics, Peptides & Aminoacids (AREA)
- Biotechnology (AREA)
- General Engineering & Computer Science (AREA)
- Bioinformatics & Cheminformatics (AREA)
- Biochemistry (AREA)
- Molecular Biology (AREA)
- Physics & Mathematics (AREA)
- Microbiology (AREA)
- General Health & Medical Sciences (AREA)
- Biophysics (AREA)
- Biomedical Technology (AREA)
- Analytical Chemistry (AREA)
- Immunology (AREA)
- Bioinformatics & Computational Biology (AREA)
- Crystallography & Structural Chemistry (AREA)
- Plant Pathology (AREA)
- Measuring Or Testing Involving Enzymes Or Micro-Organisms (AREA)
Abstract
本发明基于illumina公司的solexa测序平台提供的DNA标签文库制备方法,设计了长度为6bp独特的标签序列,将标签嵌入DNA接头中,通过连接DNA接头连接来导入标签序列,成功的建立了DNA标签文库的建库方法,并应用于solexa DNA测序。The present invention is based on the DNA tag library preparation method provided by Illumina's solexa sequencing platform, designed a unique tag sequence with a length of 6 bp, embedded the tag into the DNA adapter, and introduced the tag sequence by connecting the DNA adapter to successfully establish the DNA tag Library construction method, and applied to solexa DNA sequencing.
Description
技术领域 technical field
本发明涉及核酸测序技术领域,特别是DNA标签文库制备方法。另外,本发明还涉及标签技术,以及实现多个样品在同一反应体系中进行构建标签文库的方法。本发明的方法特别适用于第二代测序技术,尤其是solexa测序技术。The invention relates to the technical field of nucleic acid sequencing, in particular to a method for preparing a DNA tag library. In addition, the present invention also relates to labeling technology and a method for constructing a label library in the same reaction system for multiple samples. The method of the present invention is particularly suitable for the second generation sequencing technology, especially the solexa sequencing technology.
背景技术 Background technique
Illumina公司提供的Solexa DNA测序平台,可在一个反应中同时加入四种带荧光标记的核苷酸,采用边合成边测序(Sequencing BySynthesis,SBS),具有所需样品量少,高通量,高精确性,拥有简单易操作的自动化平台和功能强大等特点[1-4]。文库构建首先需要将目的片段进行末端修复,在目的片段的3’末端连接“A”碱基,将3’末端带有A碱基的目的片段与DNA接头(也称为adapter)连接,通过PCR反应将目的片段进行扩增,最后回收含有DNA接头的目的片段文库,见图1。目的片段文库与测序芯片上面的DNA接头进行杂交,通过桥式PCR进行扩增后,最后边合成边测序。在每个循环过程里,荧光标记的核苷和聚合酶被加入到单分子阵列中。互补的核苷和核苷酸片断的第一个碱基配对,通过酶加入到引物上。多余的核苷被移走。这样每个单链DNA分子通过互补碱基的配对被延伸,针对每种碱基的特定波长的激光激发结合上的核苷的标记,这个标记会释放出荧光,最后收集到的荧光信号来翻译成碱基序列。目前这种DNA建库方法可以根据需求运用于各种研究领域,如基因组的De Novo测序,基因组重测序、转录组测序和表观基因组测序等。The Solexa DNA sequencing platform provided by Illumina can simultaneously add four kinds of fluorescently labeled nucleotides in one reaction, and adopts Sequencing By Synthesis (SBS), which requires less sample volume, high throughput, high Accuracy, has the characteristics of simple and easy-to-operate automation platform and powerful functions [1-4]. Library construction first needs to carry out end repair on the target fragment, connect the "A" base at the 3' end of the target fragment, connect the target fragment with the A base at the 3' end to a DNA adapter (also called an adapter), and pass PCR The reaction amplifies the target fragment, and finally recovers the target fragment library containing the DNA linker, as shown in Figure 1. The target fragment library is hybridized with the DNA adapter on the sequencing chip, amplified by bridge PCR, and finally sequenced while being synthesized. During each cycle, fluorescently labeled nucleosides and polymerase are added to single-molecule arrays. Complementary nucleosides and the first base pairing of nucleotide fragments are enzymatically added to the primer. Excess nucleosides are removed. In this way, each single-stranded DNA molecule is extended through the pairing of complementary bases, and the laser of a specific wavelength for each base excites the label of the bound nucleoside, which will release fluorescence, and finally collect the fluorescent signal for translation into a base sequence. At present, this DNA library construction method can be used in various research fields according to the needs, such as genome De Novo sequencing, genome resequencing, transcriptome sequencing and epigenome sequencing, etc.
基于上述建库方法illumina公司也推出了DNA标签(也称为index)建库方法,如图2所示。在DNA标签建库流程中,PCR过程使用了3条PCR标签引物,通过PCR导入标签来构建DNA标签文库[5]。专利申请WO2005068656A1和WO2008093098A2中公开了一种使用标签序列标记核酸样品的来源从而可以将样品进行混合测序的方法,可以通过PCR的过程将特定的核苷酸序列(标签序列)通过PCR导入到文库中,PCR标签引物序列见表1。这些带有标签的文库可以根据需求进行任意混合,然后通过solexa测序仪器进行测序,最后将数据按标签序列进行分类。Based on the above-mentioned library construction method, Illumina Company also introduced a DNA tag (also called index) library construction method, as shown in FIG. 2 . In the DNA label library construction process, the PCR process uses 3 PCR label primers, and the DNA label library is constructed by introducing labels through PCR [5]. Patent applications WO2005068656A1 and WO2008093098A2 disclose a method of using tag sequences to mark the source of nucleic acid samples so that samples can be mixed and sequenced, and a specific nucleotide sequence (tag sequence) can be introduced into the library by PCR , See Table 1 for the sequences of PCR index primers. These tagged libraries can be arbitrarily mixed according to requirements, and then sequenced by a solexa sequencing instrument, and finally the data are classified according to the tag sequence.
表1 illumina公司提供的标签(index)序列及相对应的PCR标签引物(indexN PCR引物)序列Table 1 The index (index) sequence provided by illumina company and the corresponding PCR index primer (indexN PCR primer) sequence
但是illumina公司提供的标签文库制备的方法存在着一些缺陷:第一、目前illumina公司只提供12个长度为6bp的标签序列,标签的数量较少,随着solexa测序通量的增加,不能对大量样本进行混合测序将是一个巨大的缺陷;第二、目前illumina公司提供的标签建库方法是通过PCR反应将标签序列导入到目的片段文库中,需要3条PCR引物对目的片段进行扩增(两条公用引物和一条标签PCR引物,如表1),而且PCR扩增效率不高。第三、illumina公司提供的标签建库方法中接头不包含标签序列,每一个标签文库需要通过一个PCR反应来导入标签序列,然后针对每一个标签文库都需要切胶回收,然后将切胶回收后的标签目的片段文库进行混合,这样不仅费时费力,而且费用也较高。However, there are some defects in the method for preparing the tag library provided by Illumina: first, at present, Illumina only provides 12 tag sequences with a length of 6 bp, and the number of tags is small. Mixed sequencing of samples will be a huge defect; second, the current label library construction method provided by Illumina is to import the label sequence into the target fragment library through PCR reaction, which requires 3 PCR primers to amplify the target fragment (two common primer and a label PCR primer, as shown in Table 1), and the PCR amplification efficiency is not high. Third, the linker in the label library construction method provided by illumina does not contain the label sequence. Each label library needs to pass a PCR reaction to import the label sequence, and then for each label library, it needs to be cut and recovered, and then the glue is recovered. It is not only time-consuming and labor-intensive, but also expensive.
因此,对标签的序列及标签引入方法进行优化和改进,使标签引入的效率提高,扩大标签序列的数量,才能满足高通量的建库的要求,以适应测序通量不断提高的现状,使测序仪器的产能充分利用,降低测序的成本。Therefore, only by optimizing and improving the tag sequence and tag introduction method to improve the efficiency of tag introduction and expand the number of tag sequences can we meet the requirements of high-throughput library construction and adapt to the current situation of increasing sequencing throughput. The capacity of the sequencing instrument is fully utilized to reduce the cost of sequencing.
发明内容 Contents of the invention
本发明将标签嵌入DNA接头中,通过连接DNA标签接头来构建DNA标签文库,将连接上DNA标签接头的产物混合在一起,一次PCR反应就可完成对需要混合的所有标签文库的构建。不仅能提高目前DNA样品的测序通量,也能提高文库制备的效率和标签的识别率,极大的降低了单个文库的测序费用。The present invention embeds tags into DNA adapters, constructs DNA tag libraries by connecting DNA tag adapters, mixes products connected with DNA tag adapters, and completes the construction of all tag libraries that need to be mixed in one PCR reaction. It can not only improve the sequencing throughput of current DNA samples, but also improve the efficiency of library preparation and the recognition rate of tags, greatly reducing the sequencing cost of a single library.
鉴于目前illumina公司的solexa测序平台提供的DNA标签文库制备方法的不足,本发明改良了DNA标签文库的制备方法,设计了长度为6bp独特的标签序列以及含有所述标签序列的接头,通过包含标签序列的DNA接头的连接来导入标签序列,成功的建立了DNA标签文库的建库方法,并应用于solexa DNA测序,提高了DNA标签文库的制备的效率,增大了DNA样品的测序通量,降低了单个样品的solexa测序费用。In view of the shortcomings of the DNA tag library preparation method provided by the solexa sequencing platform of Illumina, the present invention improves the preparation method of the DNA tag library, and designs a unique tag sequence with a length of 6 bp and a linker containing the tag sequence. By including the tag The connection of the DNA adapter of the sequence to introduce the tag sequence, successfully established the library construction method of the DNA tag library, and applied it to solexa DNA sequencing, which improved the efficiency of DNA tag library preparation and increased the sequencing throughput of DNA samples. Reduced the cost of Solexa sequencing for a single sample.
在本发明中,标签设计首先需要考虑标签序列之间的可识别性和识别率的问题,然后需要考虑标签序列混合之后的每个位点的GT与AC碱基含量的平衡问题,最后考虑数据产出的可重复性和准确性。在设计标签的过程中,本发明充分考虑到以上几个因素,同时避免了标签的核酸序列出现3或3个以上连续相同的碱基,这样可以降低序列在合成过程中或测序过程中的错误率。同时尽量避免DNA标签接头自身形成发夹结构,提高DNA标签接头的连接效率。In the present invention, the label design first needs to consider the identifiability and recognition rate between the label sequences, and then needs to consider the balance of the GT and AC base content of each site after the label sequence is mixed, and finally consider the data Repeatability and accuracy of output. In the process of designing tags, the present invention fully considers the above factors, and at the same time avoids 3 or more consecutive identical bases in the nucleic acid sequence of the tag, which can reduce sequence errors during synthesis or sequencing Rate. At the same time, try to avoid the formation of hairpin structure by the DNA tag adapter itself, so as to improve the connection efficiency of the DNA tag adapter.
本发明对illumina提供的DNA接头序列进行优化,在接头中引入标签序列,通过DNA标签接头的连接将标签序列导入目的文库中。在接头连接后的PCR反应中,也就无需使用额外的标签PCR引物,从而节省了引物合成的步骤,降低了PCR反应的难度,提高了PCR反应的特异性。目前为止,通过这些DNA标签接头导入标签的建库方法及标签序列,并没有相关的报道。优化后的DNA标签接头,与illumina公司的DNA接头相比,提高了接头连接的效率,并提高了标签序列的识别效率及标签的数量。如图3所示为illumina公司的DNA标签建库流程图,图4为优化后基于接头连接的DNA标签文库构建方法实验流程图。The present invention optimizes the DNA linker sequence provided by illumina, introduces a tag sequence into the linker, and introduces the tag sequence into the target library through the connection of the DNA tag linker. In the PCR reaction after adapter ligation, there is no need to use additional label PCR primers, thereby saving the step of primer synthesis, reducing the difficulty of PCR reaction, and improving the specificity of PCR reaction. So far, there are no related reports on the library construction method and tag sequence introduced by these DNA tag adapters. Compared with Illumina's DNA adapters, the optimized DNA tag adapters have improved the efficiency of adapter ligation, and improved the recognition efficiency of tag sequences and the number of tags. Figure 3 shows the flow chart of Illumina's DNA tag library construction, and Figure 4 shows the experimental flow chart of the optimized DNA tag library construction method based on adapter ligation.
本发明基于目前illumina公司提供的Solexa Paired End测序平台,设计一段长度为6bp的特定标签核酸序列。通过测试DNA标签PCR引物的扩增效率和标签核酸序列的识别率,最后优化并筛选出67条长度为6bp的DNA标签序列(如表2,6bp的DNA标签序列DNAIndex-N)及DNA标签接头序列(DNA Index-NF/R_adapter)。这些长度为6bp的标签之间的差异至少有3个碱基,即至少3个碱基序列不同。当标签的6个碱基中的任意一个碱基出现测序错误或合成错误,都不影响到标签的最终识别。The present invention is based on the Solexa Paired End sequencing platform currently provided by Illumina, and designs a specific tag nucleic acid sequence with a length of 6 bp. By testing the amplification efficiency of DNA tag PCR primers and the recognition rate of tag nucleic acid sequences, 67 DNA tag sequences with a length of 6 bp (such as Table 2, 6 bp DNA tag sequence DNAIndex-N) and DNA tag adapters were optimized and screened Sequence (DNA Index-NF/R_adapter). The difference between these tags with a length of 6bp is at least 3 bases, that is, at least 3 bases are different in sequence. When any one of the 6 bases of the tag has a sequencing error or a synthesis error, it will not affect the final identification of the tag.
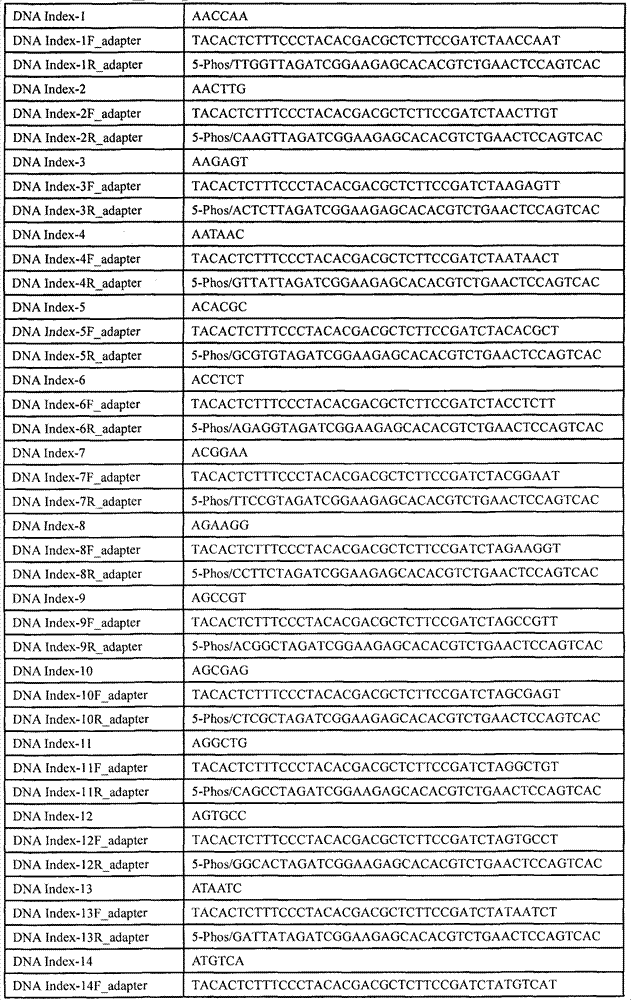
表2长度为6bp的DNA标签序列(DNA Index-N)及其相应的DNA标签接头的正向和反向序列(DNA Index-NF_adapter和DNA Index-NR_adapter),表中所示序列方向均是5’-3’方向。Table 2 is the forward and reverse sequences (DNA Index-NF_adapter and DNA Index-NR_adapter) of the DNA tag sequence (DNA Index-N) and its corresponding DNA tag adapter (DNA Index-NF_adapter and DNA Index-NR_adapter) whose length is 6bp, and the sequence directions shown in the table are all 5 '-3' direction.
使用Lasergene的PrimerSelect软件,通过分析两条序列之间形成的能量值来判断双链体之间的亲和力参数,能量值(kcal/mol)的绝对值越大表示双链体的结果越稳定,以下分别为分析了上述DNA标签接头的亲和力的能量值,说明这些DNA标签接头形成的结构非常稳定。Using Lasergene's PrimerSelect software, the affinity parameter between the duplexes is judged by analyzing the energy value formed between the two sequences. The larger the absolute value of the energy value (kcal/mol), the more stable the duplex result is, as follows are the energy values of the affinities of the above-mentioned DNA tag adapters, respectively, indicating that the structures formed by these DNA tag adapters are very stable.
预测的DNA标签接头形成的二级结构及能量值。最稳定接头序列(the most stable dimer overall)。Predicted secondary structure and energy values for DNA-tagged adapter formation. The most stable linker sequence (the most stable dimer overall).
DNA Index-1接头DNA Index-1 Adapter
The most stable dimer overall:19bp,-35.5kcal/molThe most stable dimer overall: 19bp, -35.5kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTAACCAAT 3′5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTAACCAAT 3′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGATTGGTT 5′3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGATTGGTT 5′
DNA Index-2接头DNA Index-2 Adapter
The most stable dimer overall:19bp,-34.0kcal/molThe most stable dimer overall: 19bp, -34.0kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTAACTTGT 3′5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTAACTTGT 3′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGATTGAAC 5′3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGATTGAAC 5′
DNA Index-3接头DNA Index-3 Adapter
The most stable dimer overall:19bp,-33.4kcal/molThe most stable dimer overall: 19bp, -33.4kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTAAGAGTT 3′5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTAAGAGTT 3′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGATTCTCA 5′3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGATTCTCA 5′
DNA Index-4接头DNA Index-4 Adapter
The most stable dimer overall:19bp,-33.0kcal/molThe most stable dimer overall: 19bp, -33.0kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTAATAACT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGATTATTG 5′3′
DNA Index-5接头DNA Index-5 Adapter
The most stable dimer overall:19bp,-36.6kcal/molThe most stable dimer overall: 19bp, -36.6kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTACACGCT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGATGTGCG 5′3′
DNA Index-6接头DNA Index-6 Adapter
The most stable dimer overall:19bp,-34.6kcal/molThe most stable dimer overall: 19bp, -34.6kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTACCTCTT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGATGGAGA 5′3′
DNA Index-7接头DNA Index-7 Adapter
The most stable dimer ove rall:19bp,-36.9kcal/molThe most stable dimer over rall: 19bp, -36.9kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTACGGAAT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGATGCCTT 5′3′
DNA Index-8接头DNA Index-8 Adapter
The most stable dimer overall:19bp,-35.2kcal/molThe most stable dimer overall: 19bp, -35.2kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTAGAAGGT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGATCTTCC 5′3′
DNA Index-9接头DNA Index-9 Adapter
The most stable dimer overall:19bp,-38.1kcal/molThe most stable dimer overall: 19bp, -38.1kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTAGCCGTT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGATCGGCA 5′3′
DNA Index-10接头DNA Index-10 Adapter
The most stable dimer overall:19bp,-36.9kcal/molThe most stable dimer overall: 19bp, -36.9kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTAGCGAGT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGATCGCTC 5′3′
DNA Index-11接头DNA Index-11 Adapter
The most stable dimer overall:19bp,-36.7kcal/molThe most stable dimer overall: 19bp, -36.7kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTAGGCTGT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGATCCGAC 5′3′
DNA Index-12接头DNA Index-12 Adapter
The most stable dimer overall:19bp,-36.4kcal/molThe most stable dimer overall: 19bp, -36.4kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTAGTGCCT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGATCACGG 5′3′
DNA Index-13接头DNA Index-13 linker
The most stable dimer overall:19bp,-32.9kcal/molThe most stable dimer overall: 19bp, -32.9kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTATAATCT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGATATTAG 5′3′
DNA Index-14接头DNA Index-14 Adapter
The most stable dimer overall:19bp,-33.6kcal/molThe most stable dimer overall: 19bp, -33.6kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTATGTCAT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGATACAGT 5′3′
DNA Index-15接头DNA Index-15 Adapter
The most stable dimer overall:19bp,-35.1kcal/molThe most stable dimer overall: 19bp, -35.1kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTATTCCTT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGATAAGGA 5′3′
DNA Index-16接头DNA Index-16 Adapter
The most stable dimer overall:19bp,-34.3kcal/molThe most stable dimer overall: 19bp, -34.3kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTCAACACT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGAGTTGTG 5′3′
DNA Index-17接头DNA Index-17 Adapter
The most stable dimer overall:19bp,-34.6kcal/molThe most stable dimer overall: 19bp, -34.6kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTCACAAGT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGAGTGTTC 5′3′
DNA Index-18接头DNA Index-18 Adapter
The most stable dimer overall:19bp,-37.2kcal/molThe most stable dimer overall: 19bp, -37.2kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTCACGGTT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGAGTGCCA 5′3′
DNA Index-19接头DNA Index-19 Adapter
The most stable dimer overall:19bp,-34.3kcal/molThe most stable dimer overall: 19bp, -34.3kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTCACTCAT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGAGTGAGT 5′3′
DNA Index-20接头DNA Index-20 Adapter
The most stable dimer overall:19bp,-37.8kcal/molThe most stable dimer overall: 19bp, -37.8kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTCCAACGT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGAGGTTGC 5′3′
DNA Index-21接头DNA Index-21 Adapter
The most stable dimer overall:19bp,-37.3kcal/molThe most stable dimer overall: 19bp, -37.3kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTCCAGGAT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGAGGTCCT 5′3′
DNA Index-22接头DNA Index-22 Adapter
The most stable dimer overall:19bp,-40.5kcal/molThe most stable dimer overall: 19bp, -40.5kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTCCGCCTT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGAGGCGGA 5′3′
DNA Index-23接头DNA Index-23 Adapter
The most stable dimer overall:19bp,-34.9kcal/molThe most stable dimer overall: 19bp, -34.9kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTCCTTACT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGAGGAATG 5′3′
DNA Index-24接头DNA Index-24 Adapter
The most stable dimer overall:19bp,-35.6kcal/molThe most stable dimer overall: 19bp, -35.6kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTCGAATAT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGAGCTTAT 5′3′
DNA Index-25接头DNA Index-25 Adapter
The most stable dimer overall:19bp,-39.2kcal/molThe most stable dimer overall: 19bp, -39.2kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTCGCGTCT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGAGCGCAG 5′3′
DNA Index-26接头DNA Index-26 Adapter
The most stable dimer overall:19bp,-36.9kcal/molThe most stable dimer overall: 19bp, -36.9kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTCGGTAAT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGAGCCATT 5′3′
DNA Index-27接头DNA Index-27 Adapter
The most stable dimer overall:19bp,-39.2kcal/molThe most stable dimer overall: 19bp, -39.2kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTCGTCGGT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGAGCAGCC 5′3′
DNA Index-28接头DNA Index-28 Adapter
The most stable dimer overall:19bp,-34.9kcal/molThe most stable dimer overall: 19bp, -34.9kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTCTAGCTT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGAGATCGA 5′3′
DNA Index-29接头DNA Index-29 Adapter
The most stable dimer overall:19bp,-37.5kcal/molThe most stable dimer overall: 19bp, -37.5kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTCTCCGAT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGAGAGGCT 5′3′
DNA Index-30接头DNA Index-30 Adapter
The most stable dimer overall:19bp,-34.6kcal/molThe most stable dimer overall: 19bp, -34.6kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTCTGTTGT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGAGACAAC 5′3′
DNA Index-31接头DNA Index-31 Adapter
The most stable dimer overall:19bp,-33.4kcal/molThe most stable dimer overall: 19bp, -33.4kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTCTTAGTT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGAGAATCA 5′3′
DNA Index-32接头DNA Index-32 Adapter
The most stable dimer overall:19bp,-35.5kcal/molThe most stable dimer overall: 19bp, -35.5kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTGACCTCT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGACTGGAG 5′3′
DNA Index-33接头DNA Index-33 Adapter
The most stable dimer overall:19bp,-37.6kcal/molThe most stable dimer overall: 19bp, -37.6kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTGAGCCAT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGACTCGGT 5′3′
DNA Index-34接头DNA Index-34 Adapter
The most stable dimer overall:19bp,-33.3kcal/molThe most stable dimer overall: 19bp, -33.3kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTGAGTATT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGACTCATA 5′3′
DNA Index-35接头DNA Index-35 Adapter
The most stable dimer overall:19bp,-36.0kcal/molThe most stable dimer overall: 19bp, -36.0kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTGATGGAT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGACTACCT 5′3′
DNA Index-36接头DNA Index-36 Adapter
The most stable dimer overall:19bp,-35.4kcal/molThe most stable dimer overall: 19bp, -35.4kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTGCATAGT 3′5′
::: :|||||||||||||||||||::: :||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGACGTATC 5′3′
DNA Index-37接头DNA Index-37 Adapter
The moststable dimer overall:19bp,-42.3kcal/molThe most stable dimer overall: 19bp, -42.3kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTGCCGGCT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGACGGCCG 5′3′
DNA Index-38接头DNA Index-38 Adapter
The most stable dimer overa1l:19bp,-37.2kcal/molThe most stable dimer overa1l: 19bp, -37.2kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTGCGTTAT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGACGCAAT 5′3′
DNA Index-39接头DNA Index-39 Adapter
The most stable dimer overall:19bp,-36.1kcal/molThe most stable dimer overall: 19bp, -36.1kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTGGAGAAT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGACCTCTT 5′3′
DNA Index-40接头DNA Index-40 Adapter
The most stable dimer overall:19bp,-37.8kcal/molThe most stable dimer overall: 19bp, -37.8kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTGGCATGT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGACCGTAC 5′3′
DNA Index-41接头DNA Index-41 Adapter
The most stable dimer overall:19bp,-40.8kcal/molThe most stable dimer overall: 19bp, -40.8kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTGGCGCTT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGACCGCGA 5′3′
DNA Index-42接头DNA Index-42 Adapter
The most stable dimer overall:19bp,-36.4kcal/molThe most stable dimer overall: 19bp, -36.4kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTGGTAGCT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGACCATCG 5′3′
DNA Index-43接头DNA Index-43 Adapter
The most stable dimer overall:19bp,-37.8kcal/molThe most stable dimer overall: 19bp, -37.8kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTGGTTCGT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGACCAAGC 5′3′
DNA Index-44接头DNA Index-44 Adapter
The most stable dimer overa1l:19bp,-33.3kcal/molThe most stable dimer overa1l: 19bp, -33.3kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTGTACATT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGACATGTA 5′3′
DNA Index-45接头DNA Index-45 Adapter
The most stable dimer overall:19bp,-34.8kcal/molThe most stable dimer overall: 19bp, -34.8kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTGTATCCT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGACATAGG 5′3′
DNA Index-46接头DNA Index-46 Adapter
The most stable dimer overall:19bp,-34.0kcal/molThe most stable dimer overall: 19bp, -34.0kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTGTCTGTT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGACAGACA 5′3′
DNA Index-47接头DNA Index-47 Adapter
The most stable dimer overall:19bp,-39.3kcal/molThe most stable dimer overall: 19bp, -39.3kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTGTGCGCT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGACACGCG 5′3′
DNA Index-48接头DNA Index-48 Adapter
The most stable dimer overall:19bp,-39.3kcal/molThe most stable dimer overall: 19bp, -39.3kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTGTGGCGT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGACACCGC 5′3′
DNA Index-49接头DNA Index-49 Adapter
The most stable dimer overall:19bp,-33.7kcal/molThe most stable dimer overall: 19bp, -33.7kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTGTTACAT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGACAATGT 5′3′
DNA Index-50接头DNA Index-50 Adapter
The most stable dimer overall:19bp,-34.3kcal/molThe most stable dimer overall: 19bp, -34.3kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTGTTGACT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGACAACTG 5′3′
DNA Index-51接头DNA Index-51 Adapter
The most stable dimer overall:19bp,-35.9kcal/molThe most stable dimer overall: 19bp, -35.9kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTTAATCGT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGAATTAGC 5′3′
DNA Index-52接头DNA Index-52 Adapter
The most stable dimer overall:19bp,-35.2kcal/molThe most stable dimer overall: 19bp, -35.2kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTTAGGAGT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGAATCCTC 5′3′
DNA Index-53接头DNA Index-53 Adapter
The most stable dimer overall:19bp,-35.2kcal/molThe most stable dimer overall: 19bp, -35.2kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTTAGTGCT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGAATCACG 5′3′
DNA Index-54接头DNA Index-54 Adapter
The most stable dimer overall:19bp,-35.4kcal/molThe most stable dimer overall: 19bp, -35.4kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTTATGCTT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGAATACGA 5′3′
DNA Index-55接头DNA Index-55 Adapter
The most stable dimer overall:19bp,-34.5kcal/molThe most stable dimer overall: 19bp, -34.5kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTTCAGATT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGAAGTCTA 5′3′
DNA Index-56接头DNA Index-56 Adapter
The most stable dimer overall:19bp,-34.8kcal/molThe most stable dimer overall: 19bp, -34.8kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTTCATTCT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGAAGTAAG 5′3′
DNA Index-57接头DNA Index-57 Adapter
The most stable dimer overall:19bp,-37.6kcal/molThe most stable dimer overall: 19bp, -37.6kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTTCCAGGT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGAAGGTCC 5′3′
DNA Index-58接头DNA Index-58 Adapter
The most stable dimer overall:19bp,-39.6kcal/molThe most stable dimer overall: 19bp, -39.6kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTTCCGCAT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGAAGGCGT 5′3′
DNA Index-59接头DNA Index-59 Adapter
The most stable dimer overall:19bp,-34.9kcal/molThe most stable dimer overall: 19bp, -34.9kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTTCTACCT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGAAGATGG 5′3′
DNA Index-60接头DNA Index-60 Adapter
The most stable dimer overall:19bp,-36.0kcal/molThe most stable dimer overall: 19bp, -36.0kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTTCTCGTT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGAAGAGCA 5′3′
DNA Index-61接头DNA Index-61 Adapter
The most stable dimer overall:19bp,-35.8kcal/molThe most stable dimer overall: 19bp, -35.8kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTTGCTTAT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGAACGAAT 5′3′
DNA Index-62接头DNA Index-62 Adapter
The most stable dimer overall:19bp,-36.1kcal/molThe most stable dimer overall: 19bp, -36.1kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTTGGAGAT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGAACCTCT 5′3′
DNA Index-63接头DNA Index-63 Adapter
The most stable dimer overall:19bp,-35.8kcal/molThe most stable dimer overall: 19bp, -35.8kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTTGGTCTT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGAACCAGA 5′3′
DNA Index-64接头DNA Index-64 Adapter
The most stable dimer ove rall:19bp,-33.9kcal/molThe most stable dimer over rall: 19bp, -33.9kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTTGTAATT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGAACATTA 5′3′
DNA Index-65接头DNA Index-65 Adapter
The most stable dimer overall:19bp,-34.0kcal/molThe most stable dimer overall: 19bp, -34.0kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTTTACTGT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGAAATGAC 5′3′
DNA Index-66接头DNA Index-66 linker
The most stable dimer overall:19bp,-33.6kcal/molThe most stable dimer overall: 19bp, -33.6kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTTTATAAT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGAAATATT 5′3′
DNA Index-67接头DNA Index-67 Adapter
The most stable dimer overall:19bp,-36.1kcal/molThe most stable dimer overall: 19bp, -36.1kcal/mol
5′TACACTCTTTCCCTACACGACGCTCTTCCGATCTTTCCACT 3′5′
::: : |||||||||||||||||||::: : : ||||||||||||||||||||
3′CACTGACCTCAAGTCTGCACACGAGAAGGCTAGAAAGGTG 5′3′
附图说明 Description of drawings
图1:illumina公司提供的常规DNA建库流程示意图。Figure 1: Schematic diagram of the routine DNA library construction process provided by Illumina.
图2:illumina公司提供的常规DNA标签建库流程示意图。Figure 2: Schematic diagram of the conventional DNA label library construction process provided by Illumina.
图3:illumina公司的DNA标签建库流程示意图。Figure 3: Schematic diagram of Illumina's DNA label library construction process.
图4:本发明的优化后基于接头连接的DNA标签文库构建方法的流程示意图。Figure 4: Schematic flow chart of the optimized DNA tag library construction method based on adapter ligation of the present invention.
图5:构建67个DNA标签文库电泳结果。(a)DNA标签接头文库测试(index1~index23)电泳检测结果(泳道1和泳道25分别是D2000makrer和50bp marker、泳道2到泳道24分别是使用DNA标签接头index1~index23构建的文库);(b)DNA标签接头文库测试(index23~index44)电泳检测结果(泳道1和泳道25分别是D2000makrer和50bp marker、泳道2到泳道24分别是使用DNA标签接头index23~index44构建的文库,其中泳道14为试验的阴性对照,即没有样品);(c)DNA标签接头文库测试(index45~index67)电泳检测结果(泳道1和泳道25分别是D2000 makrer和50bp marker、泳道2到泳道24分别使用DNA标签接头index45~index67构建的文库)。Figure 5: Electrophoresis results of the construction of 67 DNA tag libraries. (a) Electrophoresis detection results of DNA tag adapter library test (index1~index23) (
图6:构建的DNA标签文库使用Agilent2100检测的结果。样品名为Agilent3,图中峰从左到右分别代表Marker,样品片段大小,Marker,所测文库片段大小为284bp,浓度为32.64ng/ul。文库大小和浓度均合格。Figure 6: The detection results of the constructed DNA tag library using Agilent2100. The sample name is Agilent3. The peaks in the figure represent Marker, sample fragment size, and Marker from left to right. The measured library fragment size is 284bp, and the concentration is 32.64ng/ul. The library size and concentration are acceptable.
具体实施方式 Detailed ways
下面将结合实施例对本发明的实施方案进行详细描述,但是本领域技术人员将会理解,下列实施例仅用于说明本发明,而不应视为限定本发明的范围。Embodiments of the present invention will be described in detail below in conjunction with examples, but those skilled in the art will understand that the following examples are only used to illustrate the present invention, and should not be considered as limiting the scope of the present invention.
本发明一方面提供了一组DNA标签,所述DNA标签包括如下或由如下组成:表2所示DNA标签与之相差1个碱基的DNA标签中的至少5个,或至少10个,或至少15个,或至少20个,至少25个,或至少30个,或至少35个,或至少40个,或45个,或至少50个,或至少55个,或至少60个,或全部67个,One aspect of the present invention provides a set of DNA tags, the DNA tags include or consist of the following: at least 5, or at least 10, of the DNA tags shown in Table 2 and which differ by 1 base, or At least 15, or at least 20, or at least 25, or at least 30, or at least 35, or at least 40, or 45, or at least 50, or at least 55, or at least 60, or all 67 indivual,
所述DNA标签优选地至少包括表2所示的67个DNA标签的DNAIndex-1~DNA Index-5,或DNA Index-6~DNA Index-10,或DNA Index-11~DNA Index-15,或DNA Index-16~DNA Index-20,或DNA Index-21~DNA Index-25,或DNA Index-26~DNAIndex-30,或DNA Index-31~DNA Index-35,或DNA Index-36~DNA Index-40,或DNA Index-41~DNA Index-45,或DNA Index-46~DNA Index-50,或DNA Index-51~DNA Index-55,或DNAIndex-56~DNA Index-60,或DNA Index-61~DNA Index-65,或DNA Index-63~DNA Index-67,或者他们任何两个或多个的组合。The DNA tags preferably include at least DNA Index-1 to DNA Index-5 of the 67 DNA tags shown in Table 2, or DNA Index-6 to DNA Index-10, or DNA Index-11 to DNA Index-15, or DNA Index-16~DNA Index-20, or DNA Index-21~DNA Index-25, or DNA Index-26~DNA Index-30, or DNA Index-31~DNA Index-35, or DNA Index-36~DNA Index -40, or DNA Index-41~DNA Index-45, or DNA Index-46~DNA Index-50, or DNA Index-51~DNA Index-55, or DNA Index-56~DNA Index-60, or DNA Index- 61~DNA Index-65, or DNA Index-63~DNA Index-67, or any combination of two or more of them.
在本发明的一个具体实施方式中,本发明提供的DNA标签中,其中相差1个碱基包括所述标签中1个碱基的取代、添加或删除。In a specific embodiment of the present invention, in the DNA tag provided by the present invention, the difference of 1 base includes the substitution, addition or deletion of 1 base in the tag.
本发明另一方面,提供了所述DNA标签用于构建DNA标签文库的用途,其中DNA标签文库的DNA标签接头在3’末端包含所述DNA标签,从而构成各自相对应的DNA标签接头,所述DNA标签接头同时作为DNA标签文库的5’接头和3’接头使用。Another aspect of the present invention provides the use of the DNA tag for constructing a DNA tag library, wherein the DNA tag adapter of the DNA tag library contains the DNA tag at the 3' end, thereby constituting each corresponding DNA tag adapter, so The above-mentioned DNA tag adapter is used as the 5' adapter and the 3' adapter of the DNA tag library at the same time.
在本发明的一个具体实施方式中,提供了所述DNA标签用于构建DNA标签文库的用途,其中所述DNA标签插入DNA标签接头中的3’末端中,或通过或不通过连接子连接在DNA接头的3’末端,优选地不通过连接子连接在DNA接头的3’末端。In a specific embodiment of the present invention, the use of the DNA tag for constructing a DNA tag library is provided, wherein the DNA tag is inserted into the 3' end of the DNA tag adapter, or is connected to the DNA tag adapter with or without a linker. The 3' end of the DNA adapter is preferably ligated to the 3' end of the DNA adapter without a linker.
本发明另一方面提供了含有权利要求1或2所述的DNA标签的一组DNA标签接头,其中DNA标签文库的DNA标签接头在3’末端包含所述的标签,并且同时用作5’接头和3’接头,所述DNA标签接头包括如下或由如下组成:表2所示67个DNA标签接头与其所包含的DNA标签序列相差1个碱基的DNA标签接头中的至少5个,或至少10个,或至少15个,或至少20个,至少25个,或至少30个,或至少35个,或至少40个,或45个,或至少50个,或至少55个,或至少60个,或全部67个,Another aspect of the present invention provides a set of DNA tag adapters containing the DNA tags described in
所述DNA标签接头优选地至少包括表2所示的67个DNA标签接头中的DNA Index-1F/R_adapte~DNA Index-5F/R_adapter,或DNAIndex-6F/R_adapte~DNA Index-10F/R_adapter,或DNA Index-11F/R_adapte~DNA Index-15F/R_adapter,或DNA Index-16F/R_adapte~DNA Index-20F/R_adapter,或DNA Index-21F/R_adapte~DNA Index-25F/R_adapter,或DNA Index-26F/R_adapte~DNA Index-30F/R_adapter,或DNA Index-31F/R_adapte~DNA Index-35F/R_adapter,或DNA Index-36F/R_adapte~DNA Index-40F/R_adapter,或DNA Index-41F/R_adapte~DNA Index-45F/R_adapter,或DNA Index-46F/R_adapte~DNA Index-50F/R_adapter,或DNA Index-51F/R_adapte~DNA Index-55F/R_adapter,或DNA Index-56F/R_adapte~DNA Index-60F/R_adapter,或DNA Index-61F/R_adapte~DNA Index-65F/R_adapter,或DNA Index-66F/R_adapte~DNA Index-67F/R_adapter,或者他们任何两个或多个的组合。The DNA tag adapter preferably includes at least DNA Index-1F/R_adapte~DNA Index-5F/R_adapter, or DNAIndex-6F/R_adapte~DNA Index-10F/R_adapter among the 67 DNA tag adapters shown in Table 2, or DNA Index-11F/R_adapte~DNA Index-15F/R_adapter, or DNA Index-16F/R_adapte~DNA Index-20F/R_adapter, or DNA Index-21F/R_adapte~DNA Index-25F/R_adapter, or DNA Index-26F/ R_adapte~DNA Index-30F/R_adapter, or DNA Index-31F/R_adapte~DNA Index-35F/R_adapter, or DNA Index-36F/R_adapte~DNA Index-40F/R_adapter, or DNA Index-41F/R_adapte~DNA Index- 45F/R_adapter, or DNA Index-46F/R_adapte~DNA Index-50F/R_adapter, or DNA Index-51F/R_adapte~DNA Index-55F/R_adapter, or DNA Index-56F/R_adapte~DNA Index-60F/R_adapter, or DNA Index-61F/R_adapte~DNA Index-65F/R_adapter, or DNA Index-66F/R_adapte~DNA Index-67F/R_adapter, or any combination of two or more of them.
在本发明的一个具体实施方式中,所述的DNA标签接头中,所述的相差1个碱基包括标签中1个碱基的取代、添加或删除。In a specific embodiment of the present invention, in the DNA tag adapter, the difference of 1 base includes the substitution, addition or deletion of 1 base in the tag.
在本发明的一个进一步的方面中,提供了所述的DNA标签接头用于构建DNA标签文库的用途,优选地所述DNA标签接头同时用作DNA标签文库的5’接头和3’接头。In a further aspect of the present invention, the use of the DNA tag adapter for constructing a DNA tag library is provided, preferably the DNA tag adapter is used as both the 5' adapter and the 3' adapter of the DNA tag library.
在本发明的另一方面中,提供了通过本发明所提供的DNA标签接头构建的DNA标签文库。In another aspect of the present invention, a DNA tag library constructed by the DNA tag adapter provided in the present invention is provided.
本发明另一方面提供了一种标签文库的构建方法,所述方法的特征在于使用包含标签的DNA接头来构建标签文库。Another aspect of the present invention provides a method for constructing a tag library, which is characterized in that a tag-containing DNA linker is used to construct the tag library.
本发明进一步的方面中,提供了一种标签文库的构建方法,其包括:In a further aspect of the present invention, a method for constructing a tag library is provided, which includes:
1)提供n个DNA样品,n为整数且1≤n≤67的整数,优选地n为整数且2≤n≤67,所述DNA样品来自所有真核和原核DNA样品,包括但不限于人DNA样品;1) Provide n DNA samples, n is an integer and an integer of 1≤n≤67, preferably n is an integer and 2≤n≤67, the DNA samples are from all eukaryotic and prokaryotic DNA samples, including but not limited to human DNA samples;
2)将人基因组DNA打断,其中打断方法包括但不限于超声波打断方法,优选地使打断后的DNA条带集中在180bp左右;2) Fragmenting human genomic DNA, wherein the fragmentation method includes but not limited to the ultrasonic fragmentation method, preferably the fragmented DNA bands are concentrated at about 180bp;
3)末端修复;3) End repair;
4)DNA片段3’末端加“A”碱基;4) Add "A" base to the 3' end of the DNA fragment;
5)连接DNA标签接头,其中优选地每一个标签接头连接到DNA片段的两端,;5) connecting DNA tag adapters, wherein preferably each tag adapter is connected to both ends of the DNA fragment;
6)将步骤5)得到的连接产物进行凝胶回收纯化,优选地通过2%的琼脂糖胶进行电泳并回收,并将各个DNA样品的回收产物混合在一起;6) Gel recovery and purification of the ligated product obtained in step 5), preferably through 2% agarose gel electrophoresis and recovery, and mixing the recovered products of each DNA sample together;
7)PCR反应,使用步骤6)的回收产物的混合物作为模板,在适于扩增目的核酸的条件下进行PCR扩增,将PCR产物进行胶回收纯化,优选地回收280~300bp的目的片段。7) PCR reaction, using the mixture of recovered products in step 6) as a template, performing PCR amplification under conditions suitable for amplifying the target nucleic acid, gel recovery and purification of the PCR product, preferably recovering a target fragment of 280-300 bp.
在本发明的一个具体实施方式中,在本发明提供了一种标签文库的构建方法中,所述DNA标签接头包括如下或由如下组成:表2所示67个DNA标签接头与其所包含的DNA标签序列相差1个碱基的DNA标签接头中的至少5个,或至少10个,或至少15个,或至少20个,至少25个,或至少30个,或至少35个,或至少40个,或45个,或至少50个,或至少55个,或至少60个,或全部67个,In a specific embodiment of the present invention, in the method for constructing a tag library provided by the present invention, the DNA tag adapters include or consist of the following: the 67 DNA tag adapters shown in Table 2 and the DNA contained therein At least 5, or at least 10, or at least 15, or at least 20, at least 25, or at least 30, or at least 35, or at least 40 of the DNA tag adapters whose tag sequences differ by 1 base , or 45, or at least 50, or at least 55, or at least 60, or all 67,
所述DNA标签接头优选地至少包括表2所示的67个DNA标签接头中的DNA Index-1F/R_adapte~DNA Index-5F/R_adapter,或DNAIndex-6F/R_adapte~DNA Index-10F/R_adapter,或DNA Index-11F/R_adapte~DNA Index-15F/R_adapter,或DNA Index-16F/R_adapte~DNA Index-20F/R_adapter,或DNA Index-21F/R_adapte~DNA Index-25F/R_adapter,或DNA Index-26F/R_adapte~DNA Index-30F/R_adapter,或DNA Index-31F/R_adapte~DNA Index-35F/R_adapter,或DNA Index-36F/R_adapte~DNA Index-40F/R_adapter,或DNA Index-41F/R_adapte~DNA Index-45F/R_adapter,或DNA Index-46F/R_adapte~DNA Index-50F/R_adapter,或DNA Index-51F/R_adapte~DNA Index-55F/R_adapter,或DNA Index-56F/R_adapte~DNA Index-60F/R_adapter,或DNA Index-61F/R_adapte~DNA Index-65F/R_adapter,或DNA Index-66F/R_adapte~DNA Index-67F/R_adapter,或者他们任何两个或多个的组合。The DNA tag adapter preferably includes at least DNA Index-1F/R_adapte~DNA Index-5F/R_adapter, or DNAIndex-6F/R_adapte~DNA Index-10F/R_adapter among the 67 DNA tag adapters shown in Table 2, or DNA Index-11F/R_adapte~DNA Index-15F/R_adapter, or DNA Index-16F/R_adapte~DNA Index-20F/R_adapter, or DNA Index-21F/R_adapte~DNA Index-25F/R_adapter, or DNA Index-26F/ R_adapte~DNA Index-30F/R_adapter, or DNA Index-31F/R_adapte~DNA Index-35F/R_adapter, or DNA Index-36F/R_adapte~DNA Index-40F/R_adapter, or DNA Index-41F/R_adapte~DNA Index- 45F/R_adapter, or DNA Index-46F/R_adapte~DNA Index-50F/R_adapter, or DNA Index-51F/R_adapte~DNA Index-55F/R_adapter, or DNA Index-56F/R_adapte~DNA Index-60F/R_adapter, or DNA Index-61F/R_adapte~DNA Index-65F/R_adapter, or DNA Index-66F/R_adapte~DNA Index-67F/R_adapter, or any combination of two or more of them.
在本发明的一个具体实施方式中,在本发明提供了一种标签文库的构建方法中,所述的相差1个碱基包括标签中1个碱基的取代、添加或删除。In a specific embodiment of the present invention, in the method for constructing a tag library provided by the present invention, the difference of 1 base includes the substitution, addition or deletion of 1 base in the tag.
在本发明的一个具体实施方式中,在本发明提供了一种标签文库的构建方法中,其中步骤7)PCR反应中使用的引物包括In a specific embodiment of the present invention, the present invention provides a method for constructing a tag library, wherein the primers used in step 7) PCR reaction include
PE PCR Primers 1.0:PE PCR Primers 1.0:
AATGATACGGCGACCACCGAGATCTACACTCTTTCCCTACACGACGCTCTTCCGATCT;和AATGATACGGCGACCACCGAGATCTACACTCTTTCCCTACACGACGCTCTTCCGATCT; and
PE PCR Primers 2.0:PE PCR Primers 2.0:
CAAGCAGAAGACGGCATACGAGATGTGACTGGAGTTCAGACGTGTGCTCTTCCGATCT。CAAGCAGAAGACGGCATACGAGATGTGACTGGAGTTCAGACGTGTGCTCTTCCGATCT.
本发明进一步的另一方面中,提供了通过本发明提供的标签文库的构建方法构建的标签文库。In a further aspect of the present invention, a tag library constructed by the method for constructing a tag library provided in the present invention is provided.
实施例1Example 1
Paired End DNA寡核苷酸序列:Paired End DNA Oligonucleotide Sequence:
PE PCR Primers 1.0PE PCR Primers 1.0
AATGATACGGCGACCACCGAGATCTACACTCTTTCCCTACACGACGCTCTTCCGATCTAATGATACGGCGACCACCGAGATCTACACTCTTTCCCTACACGACGCTCTTCCGATCT
PE PCR Primers 2.0PE PCR Primers 2.0
CAAGCAGAAGACGGCATACGAGATGTGACTGGAGTTCAGACGTGTGCTCTTCCGATCTCAAGCAGAAGACGGCATACGAGATGTGACTGGAGTTCAGACGTGTGCTCTTCCGATCT
主要实验仪器及试剂Main experimental instruments and reagents
实施例1DNA标签文库的构建Construction of Example 1DNA Tag Library
1.方法步骤1. Method steps
1.1DNA打断1.1 DNA interruption
将人全血基因组DNA 5ug使用Covaris打碎仪打断6分钟(参数设置:Duty cycle(负载比)-20%;Intensity(强度)-5.0;Bursts persecond(每秒钟脉冲)-200;Duration(持续时间)-40seconds;Mode(模式)-Frequency sweeping(频率扫描);Power(功率)-33-34W;Temperature(温度)-5.5to 6℃),使其在琼脂糖电泳中显示的主要条带集中在180bp左右[5]。5ug of human whole blood genomic DNA was interrupted for 6 minutes using a Covaris breaker (parameter settings: Duty cycle (load ratio)-20%; Intensity (strength)-5.0; Bursts persecond (pulse per second)-200; Duration ( Duration)-40seconds; Mode (mode)-Frequency sweeping (frequency sweep); Power (power)-33-34W; Temperature (temperature)-5.5to 6°C), making it the main band displayed in agarose electrophoresis Concentrate on around 180bp [5].
1.2末端修复1.2 End Repair
按照下列的配比准备反应混合:Prepare the reaction mix in the following proportions:
打断后的DNA片段 35μLFragmented DNA 35μL
T4 DNA连接酶缓冲液 50μLT4 DNA Ligase Buffer 50μL
dNTPs混合液 4μLdNTPs mixture 4μL
T4 DNA聚合酶 5μL
Klenow DNA聚合酶 1μL
T4多聚核苷酸激酶 5μL
总体积 100μLTotal volume 100μL
将舒适型恒温混匀器调至20℃,反应30min,然后用QIAquick PCR纯化试剂盒进行纯化,最后将样品溶于32μL溶解缓冲液。Adjust the comfortable constant temperature mixer to 20°C, react for 30min, then purify with the QIAquick PCR purification kit, and finally dissolve the sample in 32μL dissolution buffer.
1.3DNA片段3′末端加“A”碱基1.3 Add "A" base to the 3' end of the DNA fragment
按照下列的配比准备反应混合物:Prepare the reaction mixture according to the following proportions:
末端修复后的DNA 32μLDNA after end repair 32μL
Klenow酶缓冲液 5μL
dATP(1mM) 10μLdATP(1mM) 10μL
Klenow酶(3′到5′外切酶活性) 3μLKlenow enzyme (3′ to 5′ exonuclease activity) 3 μL
总体积 50μLTotal volume 50μL
将舒适型恒温混匀器调至37℃,反应30min,然后用MiniElute PCR纯化试剂盒进行纯化,最后将样品溶于10μL Elution Buffer。Adjust the comfortable thermostatic mixer to 37°C, react for 30 minutes, then purify with MiniElute PCR purification kit, and finally dissolve the sample in 10 μL Elution Buffer.
1.4连接DNA标签接头1.4 Ligation of DNA tag adapters
按照下列的配比准备反应混合物,:Prepare the reaction mixture according to the following proportions:
上述步骤得到的DNA 10μLDNA obtained from the above steps 10μL
T4DNA连接酶缓冲液 25μLT4
DNA Index-N接头(takara公司合成) 10μLDNA Index-N adapter (synthesized by Takara Company) 10 μL
T4DNA连接酶 5μL
总体积 50μLTotal volume 50μL
注:对于不同的样品,使用不同的DNA Index-N接头(N=1~67)。所使用的DNA Index-N接头可为表2中所示任一对DNA Index-NF_adapter和DNA Index-NR_adapter退火后形成的DNA标签接头。Note: For different samples, use different DNA Index-N adapters (N=1~67). The DNA Index-N linker used can be the DNA index linker formed after the annealing of any pair of DNA Index-NF_adapter and DNA Index-NR_adapter shown in Table 2.
将舒适型恒温混匀器(调至20℃,反应15min,然后用QIAquickPCR纯化试剂盒进行纯化,最后将样品溶于30μL溶解缓冲液中。Adjust the comfortable constant temperature mixer (adjusted to 20°C, react for 15min, then use the QIAquickPCR purification kit to purify, and finally dissolve the sample in 30μL of dissolution buffer.
1.5连接产物的胶回收纯化1.5 Gel recovery and purification of ligation products
将连接产物于2%的琼脂糖胶中进行电泳分离;随后将280~300bp的目的片段放入Eppendorf管中。用QIAquick胶纯化试剂盒进行胶纯化回收,回收产物溶于30μL Elution Buffer。The ligation product was separated by electrophoresis in 2% agarose gel; then the target fragment of 280-300 bp was put into an Eppendorf tube. Use the QIAquick Gel Purification Kit for gel purification and recovery, and the recovered product is dissolved in 30 μL Elution Buffer.
1.6PCR反应1.6 PCR reaction
PCR反应:按照下列的反应体系准备反应混合物,将试剂放置于冰上。PCR reaction: Prepare the reaction mixture according to the following reaction system, and place the reagents on ice.
胶回收纯化后的DNA 10μLGel recovery purified DNA 10μL
PE PCR Primers 1.0 1μLPE PCR Primers 1.0 1μL
PE PCR Primers 2.0 1μLPE PCR Primers 2.0 1μL
Phusion DNA聚合酶 25μLPhusion DNA Polymerase 25μL
ddH2O 13μLddH 2 O 13 μL
总体积 50μLTotal volume 50μL
PCR反应条件PCR reaction conditions
9g℃30s9g℃30s
72℃5min72℃5min
4℃保存Store at 4°C
1.7PCR产物的胶回收纯化1.7 Gel recovery and purification of PCR products
将PCR产物于2%琼脂糖胶中电泳分离,切割回收280~300bp的目的片段,用QIAquick胶纯化试剂盒进行胶纯化回收,回收产物溶于30μL Elution Buffer。The PCR product was separated by electrophoresis in 2% agarose gel, and the target fragment of 280-300 bp was cut and recovered, and the gel purification was performed with the QIAquick gel purification kit, and the recovered product was dissolved in 30 μL Elution Buffer.
1.8DNA制备产物检测1.8 DNA preparation product detection
使用Agilent 2100 Bioanalyzer根据生产商说明书的操作方法,检测文库产量(见图6)。所测文库片段大小为284bp,浓度为32.64ng/ul。文库大小和浓度均合格。Use the Agilent 2100 Bioanalyzer to detect the library yield according to the manufacturer's instructions (see Figure 6). The size of the measured library fragment was 284bp, and the concentration was 32.64ng/ul. The library size and concentration are acceptable.
2.结果分析2. Result analysis
构建的67个DNA标签文库电泳结果如下图5所示,图5中分别使用了D2000和50bp的marker,分别产自天根公司和NEB公司;箭头所标记的为目的文库片段大小。The electrophoresis results of the 67 constructed DNA tag libraries are shown in Figure 5 below. In Figure 5, D2000 and 50bp markers were used, respectively, produced by Tiangen Company and NEB Company; the arrows marked the target library fragment size.
Solexa测序结果统计:标签完全识别即0错误匹配(mismatch)占98.43%,标签测出错误1个碱基即1错误匹配的占0.09%,其他读取结果(other reads)占1.48%,所以测序结果标签的识别率为98.5%,可以满足solexa DNA index的测序要求。Statistics of Solexa sequencing results: 98.43% of tags are fully identified,
尽管本发明的具体实施方式已经得到详细的描述,本领域技术人员将会理解。根据已经公开的所有教导,可以对那些细节进行各种修改和替换,这些改变均在本发明的保护范围之内。本发明的全部范围由所附权利要求及其任何等同物给出。Although specific embodiments of the present invention have been described in detail, those skilled in the art will understand. Based on all the teachings that have been disclosed, various modifications and substitutions can be made to those details, and these changes are all within the scope of the invention. The full scope of the invention is given by the appended claims and any equivalents thereof.
参考文献references
1、Paired-End sequencing User Guide;illumina part#10038801. Paired-End sequencing User Guide; illumina part#1003880
2、Preparing samples for ChIP sequencing for DNA;illuminapart#11257047 Rev.A;2. Preparing samples for ChIP sequencing for DNA; illuminapart#11257047 Rev.A;
3、mRNA sequencing sample preparation Guide;illuminapart#1004898 Rev.D3. mRNA sequencing sample preparation Guide; illuminapart#1004898 Rev.D
4、Preparing 2-5kb samples for mate pair library sequencing;illumina part#1005363 Rev.B;4. Preparing 2-5kb samples for mate pair library sequencing; illumina part#1005363 Rev.B;
5、Preparing samples for multiplexed Paired-End sequencing;illumina part#1005361 Rev.B;5. Preparing samples for multiplexed Paired-End sequencing; illumina part#1005361 Rev.B;
Claims (44)
Priority Applications (3)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| CN 201010299257 CN102409045B (en) | 2010-09-21 | 2010-09-21 | Tag library constructing method based on DNA (deoxyribonucleic acid) adapter connection as well as used tag and tag adapter |
| PCT/CN2011/079898 WO2012037876A1 (en) | 2010-09-21 | 2011-09-20 | Dna tag and application thereof |
| HK12109359.2A HK1168626B (en) | 2012-09-24 | Tag library constructing method based on dna (deoxyribonucleic acid) adapter connection as well as used tag and tag adapter |
Applications Claiming Priority (1)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| CN 201010299257 CN102409045B (en) | 2010-09-21 | 2010-09-21 | Tag library constructing method based on DNA (deoxyribonucleic acid) adapter connection as well as used tag and tag adapter |
Publications (2)
| Publication Number | Publication Date |
|---|---|
| CN102409045A CN102409045A (en) | 2012-04-11 |
| CN102409045B true CN102409045B (en) | 2013-09-18 |
Family
ID=45873441
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| CN 201010299257 Active CN102409045B (en) | 2010-09-21 | 2010-09-21 | Tag library constructing method based on DNA (deoxyribonucleic acid) adapter connection as well as used tag and tag adapter |
Country Status (2)
| Country | Link |
|---|---|
| CN (1) | CN102409045B (en) |
| WO (1) | WO2012037876A1 (en) |
Families Citing this family (20)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN103374759B (en) * | 2012-04-26 | 2015-09-30 | 中国科学院上海生命科学研究院 | A kind of detection of lung cancer shifts method and the application thereof of significant SNP |
| CN103571822B (en) * | 2012-07-20 | 2016-03-30 | 中国科学院植物研究所 | A kind of multipurpose DNA fragmentation enriching method analyzed for new-generation sequencing |
| CN102952877B (en) * | 2012-08-06 | 2014-09-24 | 深圳华大基因研究院 | Method and system for detecting copy number of alpha globin gene |
| CN103290104B (en) * | 2013-01-23 | 2016-03-02 | 北京诺禾致源生物信息科技有限公司 | A kind of genomic samples breaking method being applied to the simple and direct cheapness of s-generation order-checking |
| US10302614B2 (en) | 2014-05-06 | 2019-05-28 | Safetraces, Inc. | DNA based bar code for improved food traceability |
| JP6483518B2 (en) * | 2014-05-20 | 2019-03-13 | 信越化学工業株式会社 | Conductive polymer composite and substrate |
| CN107075513B (en) | 2014-09-12 | 2020-11-03 | 深圳华大智造科技有限公司 | Isolated oligonucleotides and their use in nucleic acid sequencing |
| WO2016049929A1 (en) * | 2014-09-30 | 2016-04-07 | 天津华大基因科技有限公司 | Method for constructing sequencing library and application thereof |
| US10962512B2 (en) * | 2015-08-03 | 2021-03-30 | Safetraces, Inc. | Pathogen surrogates based on encapsulated tagged DNA for verification of sanitation and wash water systems for fresh produce |
| CN105506125B (en) * | 2016-01-12 | 2019-01-22 | 上海美吉生物医药科技有限公司 | A kind of sequencing approach and a kind of two generation sequencing libraries of DNA |
| CN105734048A (en) * | 2016-02-26 | 2016-07-06 | 武汉冰港生物科技有限公司 | PCR-free sequencing library preparation method for genome DNA |
| RU2760737C2 (en) * | 2016-12-27 | 2021-11-30 | Еги Тек (Шэнь Чжэнь) Ко., Лимитед | Method for sequencing based on one fluorescent dye |
| CN108728903A (en) * | 2017-04-21 | 2018-11-02 | 深圳市乐土精准医疗科技有限公司 | The banking process of thalassemia large sample screening is used for based on high-flux sequence |
| CN108949905B (en) * | 2017-05-23 | 2022-05-17 | 深圳华大基因股份有限公司 | Control library and its construction method |
| US10926264B2 (en) | 2018-01-10 | 2021-02-23 | Safetraces, Inc. | Dispensing system for applying DNA taggants used in combinations to tag articles |
| US10556032B2 (en) | 2018-04-25 | 2020-02-11 | Safetraces, Inc. | Sanitation monitoring system using pathogen surrogates and surrogate tracking |
| US11853832B2 (en) | 2018-08-28 | 2023-12-26 | Safetraces, Inc. | Product tracking and rating system using DNA tags |
| US11200383B2 (en) | 2018-08-28 | 2021-12-14 | Safetraces, Inc. | Product tracking and rating system using DNA tags |
| CN110468188B (en) * | 2019-08-22 | 2023-08-22 | 广州微远医疗器械有限公司 | Tag sequence set for next generation sequencing and its design method and application |
| US12258638B2 (en) | 2020-04-16 | 2025-03-25 | Safetraces, Inc. | Airborne pathogen simulants and mobility testing |
Citations (3)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO2005068656A1 (en) * | 2004-01-12 | 2005-07-28 | Solexa Limited | Nucleic acid characterisation |
| CN101100764A (en) * | 2007-06-13 | 2008-01-09 | 北京万达因生物医学技术有限责任公司 | Molecule substitution label sequencing parallel detection method-oligomictic nucleic acid coding label molecule library micro-sphere array analysis |
| WO2008093098A2 (en) * | 2007-02-02 | 2008-08-07 | Illumina Cambridge Limited | Methods for indexing samples and sequencing multiple nucleotide templates |
Family Cites Families (4)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| ES2446927T3 (en) * | 2006-03-01 | 2014-03-10 | Keygene N.V. | Rapid sequence-based SNP detection using ligation assays |
| CN101434988B (en) * | 2007-11-16 | 2013-05-01 | 深圳华因康基因科技有限公司 | High throughput oligonucleotide sequencing method |
| EP4335932A3 (en) * | 2008-11-07 | 2024-06-26 | Adaptive Biotechnologies Corporation | Methods of monitoring conditions by sequence analysis |
| CN101748213B (en) * | 2008-12-12 | 2013-05-08 | 深圳华大基因研究院 | Environmental microorganism detection method and system |
-
2010
- 2010-09-21 CN CN 201010299257 patent/CN102409045B/en active Active
-
2011
- 2011-09-20 WO PCT/CN2011/079898 patent/WO2012037876A1/en not_active Ceased
Patent Citations (3)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO2005068656A1 (en) * | 2004-01-12 | 2005-07-28 | Solexa Limited | Nucleic acid characterisation |
| WO2008093098A2 (en) * | 2007-02-02 | 2008-08-07 | Illumina Cambridge Limited | Methods for indexing samples and sequencing multiple nucleotide templates |
| CN101100764A (en) * | 2007-06-13 | 2008-01-09 | 北京万达因生物医学技术有限责任公司 | Molecule substitution label sequencing parallel detection method-oligomictic nucleic acid coding label molecule library micro-sphere array analysis |
Also Published As
| Publication number | Publication date |
|---|---|
| HK1168626A1 (en) | 2013-01-04 |
| CN102409045A (en) | 2012-04-11 |
| WO2012037876A1 (en) | 2012-03-29 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| CN102409045B (en) | Tag library constructing method based on DNA (deoxyribonucleic acid) adapter connection as well as used tag and tag adapter | |
| CN101967476B (en) | Joint connection-based deoxyribonucleic acid (DNA) polymerase chain reaction (PCR)-free tag library construction method | |
| CN102409049B (en) | A method for constructing DNA tag library based on PCR | |
| CN102409048B (en) | A DNA tagging library construction method based on high-throughput sequencing | |
| CN102653784B (en) | Tag used for multiple nucleic acid sequencing and application method thereof | |
| CN102409408B (en) | Method for accurate detection of whole genome methylation sites by utilizing trace genome DNA (deoxyribonucleic acid) | |
| CN103088433B (en) | Construction method and application of genome-wide methylation high-throughput sequencing library | |
| CN106192021B (en) | Method for constructing series connection RAD [restriction-site-associated DNA (deoxyribonucleic acid)] tag sequencing libraries | |
| CN102409046B (en) | Small RNA (ribonucleic acid) tags | |
| CN103602735B (en) | Utilize the method that high-flux sequence Accurate Measurement Mitochondrial DNA high and low frequency suddenlys change | |
| WO2012037875A1 (en) | Dna tags and use thereof | |
| CN104313172A (en) | Method for simultaneous genotyping of large number of samples | |
| WO2022199242A1 (en) | Set of barcode linkers and medium-flux multi-single-cell representative dna methylation library construction and sequencing method | |
| CN115873922B (en) | A single-cell full-length transcript library construction and sequencing method | |
| Albrecht et al. | Amplicon-based bisulfite conversion-NGS DNA methylation analysis protocol | |
| CN102409044B (en) | Indexes for digital gene expression profiling (DGE) and use method thereof | |
| CN111005075B (en) | Y-shaped adapter for double-sample co-construction sequencing library and method for co-construction sequencing library of two samples | |
| CN108342385A (en) | A kind of connector and the method that sequencing library is built by way of high efficiency cyclisation | |
| HK1168626B (en) | Tag library constructing method based on dna (deoxyribonucleic acid) adapter connection as well as used tag and tag adapter | |
| CN113736777A (en) | Design and synthesis method of nucleic acid coding probe for high-throughput sequencing | |
| CN116042770B (en) | Method and kit for preparing miRNA library in urine and quantifying expression | |
| HK1153226B (en) | An adapter ligation-based method for pcr-free indexed dna library construction | |
| HK1168627B (en) | Dna (deoxyribonucleic acid) index library building method based on pcr (polymerase chain reaction) | |
| HK1168630B (en) | Dna index library building method based on high throughput sequencing | |
| CN114622286A (en) | A high-throughput transcriptome sequencing library construction method and its application |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| C06 | Publication | ||
| PB01 | Publication | ||
| C10 | Entry into substantive examination | ||
| SE01 | Entry into force of request for substantive examination | ||
| REG | Reference to a national code |
Ref country code: HK Ref legal event code: DE Ref document number: 1168626 Country of ref document: HK |
|
| ASS | Succession or assignment of patent right |
Owner name: BGI TECHNOLOGY SOLUTIONS CO., LTD. Free format text: FORMER OWNER: BGI-SHENZHEN CO., LTD. Effective date: 20130715 Free format text: FORMER OWNER: BGI-SHENZHEN Effective date: 20130715 |
|
| C41 | Transfer of patent application or patent right or utility model | ||
| TA01 | Transfer of patent application right |
Effective date of registration: 20130715 Address after: 518083 science and Technology Pioneer Park, comprehensive building, Beishan Industrial Zone, Yantian District, Guangdong, Shenzhen 201 Applicant after: BGI Technology Solutions Co., Ltd. Address before: Beishan Industrial Zone Building in Yantian District of Shenzhen city of Guangdong Province in 518083 Applicant before: BGI-Shenzhen Co., Ltd. Applicant before: BGI-Shenzhen |
|
| C14 | Grant of patent or utility model | ||
| GR01 | Patent grant | ||
| REG | Reference to a national code |
Ref country code: HK Ref legal event code: GR Ref document number: 1168626 Country of ref document: HK |
|
| TR01 | Transfer of patent right |
Effective date of registration: 20190726 Address after: 214200 Xingye Road, Xinjie Street, Yixing City, Wuxi City, Jiangsu Province, 298 Patentee after: Huada Qinglan Biotechnology (Wuxi) Co., Ltd. Address before: 518083 science and Technology Pioneer Park, comprehensive building, Beishan Industrial Zone, Yantian District, Guangdong, Shenzhen 201 Patentee before: BGI Technology Solutions Co., Ltd. |
|
| TR01 | Transfer of patent right |