CN102373288A - Method and kit for sequencing target areas - Google Patents
Method and kit for sequencing target areas Download PDFInfo
- Publication number
- CN102373288A CN102373288A CN2011103919902A CN201110391990A CN102373288A CN 102373288 A CN102373288 A CN 102373288A CN 2011103919902 A CN2011103919902 A CN 2011103919902A CN 201110391990 A CN201110391990 A CN 201110391990A CN 102373288 A CN102373288 A CN 102373288A
- Authority
- CN
- China
- Prior art keywords
- sequencing
- target region
- linker
- product
- adapter
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Granted
Links
Images
Landscapes
- Measuring Or Testing Involving Enzymes Or Micro-Organisms (AREA)
Abstract
本发明涉及基因工程领域,提供了一种对目标区域进行测序的方法及试剂盒,该方法包括以下步骤:A.利用特异性引物,对待测样品中的多个目标区域进行扩增,并基于扩增产物构建测序文库;B.对测序文库进行单分子扩增,得到与所述多个目标区域对应的多个单分子扩增产物;C.同时对所述多个单分子扩增产物进行高通量基因测序,得到所述多个目标区域的序列信息。该方法及其相应的试剂盒利用高通量测序技术对多个目标区域同时进行区域测序,能准确得出这些区域的序列信息,包括已知突变和未知突变位点的变异情况,检测灵敏度高,并能进一步同时对大量样品进行检测。
The present invention relates to the field of genetic engineering, and provides a method and a kit for sequencing a target region. The method includes the following steps: A. using specific primers to amplify multiple target regions in a sample to be tested, and based on Amplifying the product to construct a sequencing library; B. performing single-molecule amplification on the sequencing library to obtain a plurality of single-molecule amplification products corresponding to the multiple target regions; C. simultaneously performing a single-molecule amplification on the multiple single-molecule amplification products High-throughput gene sequencing to obtain sequence information of the multiple target regions. This method and its corresponding kit use high-throughput sequencing technology to simultaneously perform regional sequencing on multiple target regions, and can accurately obtain the sequence information of these regions, including the variation of known mutations and unknown mutation sites, with high detection sensitivity , and can further detect a large number of samples at the same time.
Description
技术领域 technical field
本发明涉及基因工程领域,更具体地说,涉及一种对目标区域进行测序的方法及试剂盒。The invention relates to the field of genetic engineering, more specifically, to a method and a kit for sequencing a target region.
背景技术 Background technique
自从人类基因组核苷酸序列草图和许多其他基因组草图问世,以及人类和许多其他生物体基因组草图中大量单核苷酸多态性(SNP)和小的插入/缺失多态性的发现,人们开始对核酸测定产生兴趣。研究表明,某些特定位置的核酸序列变异包含了大量的生物学信息,可能与人类或动植物的某些性状相关,这些信息将对医药和农业等的方方面面产生广泛的影响。举例来说,可以预期的是,这个领域的发展有可能实现个性化医疗。Since the advent of the draft nucleotide sequence of the human genome and many other draft genomes, as well as the discovery of a large number of single nucleotide polymorphisms (SNPs) and small insertion/deletion polymorphisms in the draft genomes of humans and many other organisms, people began to Interested in nucleic acid determination. Studies have shown that nucleic acid sequence variations at specific positions contain a large amount of biological information, which may be related to certain traits of humans or animals and plants. These information will have a wide impact on all aspects of medicine and agriculture. For example, it is anticipated that developments in this field may enable personalized medicine.
目前,对目标区域(特定位置)进行测序的常见方法为Sanger测序法,通过Sanger测序法能够对目标区域进行区域检测,但是Sanger测序法每次只能对一个样品的某一段区域进行测序,测序成本高,灵敏度低(20%),且不能得出各突变位点发生变异的精确数据。At present, the common method for sequencing the target region (specific position) is the Sanger sequencing method. The Sanger sequencing method can detect the target region, but the Sanger sequencing method can only sequence a certain region of a sample at a time. The cost is high, the sensitivity is low (20%), and accurate data on the variation of each mutation site cannot be obtained.
因此,需要一种能同时对待测样品的多个目标区域进行区域检测的新方法,该方法能够准确得到这些区域的序列信息,包括已知突变和未知突变的各突变位点的变异情况,检测灵敏度高,还能进一步同时对大量样品进行检测。Therefore, there is a need for a new method for regional detection of multiple target regions of the sample to be tested at the same time. This method can accurately obtain the sequence information of these regions, including the variation of each mutation site of known mutations and unknown mutations, and detect The sensitivity is high, and it can further detect a large number of samples at the same time.
发明内容 Contents of the invention
本发明的目的在于提供一种对目标区域进行测序的方法及试剂盒,能同时对待测样品的多个目标区域进行区域检测,准确得到这些区域的序列信息,包括已知突变和未知突变的各突变位点的变异情况,检测灵敏度高,还能进一步同时对大量样品进行检测。The object of the present invention is to provide a method and a kit for sequencing target regions, which can detect multiple target regions of a sample to be tested at the same time, and accurately obtain the sequence information of these regions, including various mutations of known mutations and unknown mutations. The variation of the mutation site has high detection sensitivity, and it can further detect a large number of samples at the same time.
本发明是这样实现的,一种对目标区域进行测序的方法,包括以下步骤:The present invention is achieved in this way, a method for sequencing the target region, comprising the following steps:
A.利用特异性引物,对待测样品中的多个目标区域进行扩增,并基于扩增产物构建测序文库;A. Use specific primers to amplify multiple target regions in the sample to be tested, and construct a sequencing library based on the amplified products;
B.对测序文库进行单分子扩增,得到与所述多个目标区域对应的多个单分子扩增产物;B. performing single-molecule amplification on the sequencing library to obtain multiple single-molecule amplification products corresponding to the multiple target regions;
C.同时对所述多个单分子扩增产物进行高通量基因测序,得到所述多个目标区域的序列信息。C. Simultaneously performing high-throughput gene sequencing on the multiple single-molecule amplification products to obtain sequence information of the multiple target regions.
其中,所述步骤A包括以下步骤:Wherein, described step A comprises the following steps:
A1.利用特异性引物,对待测样品中的多个目标区域进行扩增,得到与所述多个目标区域对应的扩增产物;A1. using specific primers to amplify multiple target regions in the sample to be tested, and obtain amplification products corresponding to the multiple target regions;
A2.利用接头元件,与所述多个目标区域对应的扩增产物进行连接,得到测序文库;所述接头元件采用平末端接头、突出末端接头、带茎环结构的接头和分叉接头中的至少一种。A2. Use adapter elements to connect the amplified products corresponding to the multiple target regions to obtain a sequencing library; the adapter elements use blunt end adapters, protruding end adapters, adapters with stem-loop structures, and bifurcated adapters. at least one.
其中,步骤A2包括以下步骤:Wherein, step A2 includes the following steps:
A21.对与所述多个目标区域对应的扩增产物进行片段化,得到片段化产物;A21. Fragmenting the amplified products corresponding to the multiple target regions to obtain fragmented products;
A22.利用接头元件,与片段化产物进行连接,构建测序文库。A22. Use the adapter element to connect the fragmented product to construct a sequencing library.
步骤A所述的目标区域测序文库的长度无特殊限制,优选为25bp~500bp。更优选为50bp~200bp,更优选为70bp~130bp。The length of the target region sequencing library described in step A is not particularly limited, and is preferably 25bp-500bp. More preferably, it is 50bp to 200bp, and more preferably, it is 70bp to 130bp.
其中,所述步骤A22包括以下步骤:Wherein, said step A22 includes the following steps:
A221.利用第一接头与片段化产物的两端连接,得第一连接产物;A221. Using the first linker to connect the two ends of the fragmentation product to obtain the first ligation product;
A222.环化第一连接产物,得环化产物;A222. cyclization of the first connection product to obtain a cyclization product;
A223.II s型限制性内切酶酶切环化产物,得酶切产物;A223. II s type restriction endonuclease digests the cyclization product to obtain the digested product;
A224.在酶切产物两端接上第二接头和第三接头,得测序文库。A224. Connect the second linker and the third linker to both ends of the restriction product to obtain a sequencing library.
其中,所述步骤A22包括以下步骤:Wherein, said step A22 includes the following steps:
A221’.利用第四接头与片段化产物连接,得第二连接产物;A221'. Use the fourth linker to connect with the fragmentation product to obtain the second connection product;
A222’.II s型限制性内切酶酶切第二连接产物,得带有第四接头的酶切片段;A222'. II s type restriction endonuclease digests the second connection product to obtain an enzyme-cut fragment with the fourth linker;
A223’.带有第四接头的酶切片段与第五接头连接,形成测序文库。A223'. The digested fragment with the fourth linker is connected with the fifth linker to form a sequencing library.
其中,步骤A2所述接头元件中的至少一个接头包含有第一标签序列,用于在文库构建过程中,对不同待测样品的测序文库进行标记。所述第一标签序列,优选为带有特定碱基序列的核酸分子,其碱基数不限。Wherein, at least one of the linker elements in the step A2 includes a first tag sequence, which is used to tag the sequencing libraries of different samples to be tested during the library construction process. The first tag sequence is preferably a nucleic acid molecule with a specific base sequence, and the number of bases is not limited.
进一步的,所述第一标签序列碱基数为3~20,更优选为4~10。Further, the number of bases in the first tag sequence is 3-20, more preferably 4-10.
其中,步骤A所述的特异性引物与目标区域完全互补或部分互补。Wherein, the specific primer described in step A is fully or partially complementary to the target region.
进一步的,每个目标区域对应的特异性引物中,至少一条引物与该目标区域部分互补,该部分互补的引物的5’端包含有第二标签序列,用于在扩增目标区域过程中,对不同待测样品的目标区域扩增产物进行标记。所述第二标签序列,优选为带有特定碱基序列的核酸分子,其碱基数不限。Further, among the specific primers corresponding to each target region, at least one primer is partially complementary to the target region, and the 5' end of the partially complementary primer contains a second tag sequence, which is used in the process of amplifying the target region, The amplification products of target regions of different samples to be tested are labeled. The second tag sequence is preferably a nucleic acid molecule with a specific base sequence, and the number of bases is not limited.
进一步的,所述第二标签序列碱基数为3~20,更优选为4~10。Further, the number of bases in the second tag sequence is 3-20, more preferably 4-10.
其中,所述步骤A中同一样品的各目标区域的扩增,同时进行或部分同时进行或分别独立进行。Wherein, the amplification of each target region of the same sample in the step A is carried out simultaneously or partly simultaneously or independently.
其中,步骤B所述的单分子扩增的方法为乳液PCR、桥式PCR中的至少一种。Wherein, the single-molecule amplification method described in step B is at least one of emulsion PCR and bridge PCR.
其中,步骤C所述的高通量测序技术为基于聚合酶的合成测序法或基于连接酶的连接测序法。Wherein, the high-throughput sequencing technology described in step C is a polymerase-based sequencing-by-synthesis method or a ligase-based sequencing-by-ligation method.
本发明的还提供了一种能够用于本发明的任一种测序方法的试剂盒,本发明是这样实现的,一种对目标区域进行测序的试剂盒,包括:The present invention also provides a kit that can be used in any of the sequencing methods of the present invention, the present invention is achieved in this way, a kit for sequencing the target region, comprising:
特异性引物,用于对待测样品中的多个目标区域进行扩增;Specific primers for amplifying multiple target regions in the sample to be tested;
接头元件,用于与扩增产物结合构建测序文库。Linker elements are used to combine with amplification products to construct sequencing libraries.
其中,所述接头元件采用平末端接头、突出末端接头、带茎环结构的接头和分叉接头中的至少一种。Wherein, the joint element adopts at least one of a blunt end joint, a protruding end joint, a joint with a stem-loop structure and a bifurcated joint.
其中,所述接头元件中的至少一个接头包含有第一标签序列,用于在文库构建过程中,对不同待测样品的测序文库进行标记。所述第一标签序列优选为带有特定碱基序列的核酸分子,其碱基数不限,优选为3~20。Wherein, at least one adapter in the adapter element includes a first tag sequence, which is used to mark the sequencing libraries of different samples to be tested during the library construction process. The first tag sequence is preferably a nucleic acid molecule with a specific base sequence, and the number of bases is not limited, preferably 3-20.
其中,所述的特异性引物与目标区域完全互补或部分互补。Wherein, the specific primer is fully or partially complementary to the target region.
进一步的,每个目标区域对应的特异性引物中,至少一条引物与该目标区域部分互补,该部分互补的引物的5’端包含有第二标签序列,用于在扩增目标区域过程中,对不同待测样品的目标区域扩增产物进行标记。所述第二标签序列优选为带有特定碱基序列的核酸分子,其碱基数不限,优选为3~20,更优选为4~10。Further, among the specific primers corresponding to each target region, at least one primer is partially complementary to the target region, and the 5' end of the partially complementary primer contains a second tag sequence, which is used in the process of amplifying the target region, The amplification products of target regions of different samples to be tested are labeled. The second tag sequence is preferably a nucleic acid molecule with a specific base sequence, and the number of bases is not limited, preferably 3-20, more preferably 4-10.
与现有技术相比,本发明的方法及试剂盒通过高通量测序技术对待测样品的多个目标区域同时进行深度测序,准确得出这些目标区域的序列信息,包括已知突变和未知突变的各突变位点的变异情况,精确得出各样品的各突变位点发生变异的频率,检测灵敏度高,并能进一步对大量样品同时进行多区域测序。Compared with the prior art, the method and kit of the present invention perform deep sequencing on multiple target regions of the sample to be tested simultaneously through high-throughput sequencing technology, and accurately obtain the sequence information of these target regions, including known mutations and unknown mutations The variation of each mutation site in each sample can be accurately obtained, and the frequency of each mutation site of each sample can be accurately obtained. The detection sensitivity is high, and it can further perform multi-region sequencing on a large number of samples at the same time.
附图说明 Description of drawings
图1是本发明一个实施例中的对目标区域进行测序的的方法流程图;FIG. 1 is a flowchart of a method for sequencing a target region in one embodiment of the present invention;
图2是本发明一个实施例中的单突出末端接头的接头示意图;Figure 2 is a joint schematic diagram of a single protruding end joint in one embodiment of the present invention;
图3是本发明一个实施例中的双突出末端接头的结构示意图;Fig. 3 is a structural schematic diagram of a double protruding end joint in an embodiment of the present invention;
图4是本发明一个实施例中的带茎环结构的接头的结构示意图;Fig. 4 is a structural schematic diagram of a joint with a stem-loop structure in one embodiment of the present invention;
图5是本发明一个实施例中的分叉接头的结构示意图;Fig. 5 is a schematic structural view of a bifurcated joint in an embodiment of the present invention;
图6是本发明一个实施例中的T末端分叉接头的结构示意图;FIG. 6 is a schematic structural view of a T-terminal bifurcation adapter in an embodiment of the present invention;
图7是本发明另一个实施例中的分叉接头的结构示意图;Fig. 7 is a schematic structural view of a bifurcated joint in another embodiment of the present invention;
图8是本发明一个实施例中的双脱氧分叉接头的结构示意图;Figure 8 is a schematic structural view of a dideoxy bifurcated linker in one embodiment of the present invention;
图9是本发明一个实施例中利用片段化产物和接头元件构建测序文库的方法流程图;Fig. 9 is a flowchart of a method for constructing a sequencing library using fragmentation products and linker elements in an embodiment of the present invention;
图10是本发明另一个实施例中利用片段化产物和接头元件构建测序文库的方法流程图;Fig. 10 is a flowchart of a method for constructing a sequencing library using fragmentation products and linker elements in another embodiment of the present invention;
图11是本发明另一个实施例中利用片段化产物和接头元件构建测序文库的方法流程图。Fig. 11 is a flowchart of a method for constructing a sequencing library using fragmentation products and linker elements in another embodiment of the present invention.
具体实施方式 Detailed ways
为了使本发明的目的、技术方案及优点更加清楚明白,以下结合附图及实施例,对本发明进行进一步详细说明。In order to make the object, technical solution and advantages of the present invention clearer, the present invention will be further described in detail below in conjunction with the accompanying drawings and embodiments.
本发明所述的目标区域,为任意基因组上的任意序列,可根据需要进行选择,包括但不限于基因的内部序列、基因的外部调控区域,所述基因的内部序列,包括但不限于基因的内含子区域、外显子区域、同时含有内含子与外显子的区域。The target region of the present invention is any sequence on any genome, which can be selected according to needs, including but not limited to the internal sequence of the gene, the external regulatory region of the gene, the internal sequence of the gene, including but not limited to the internal sequence of the gene Intron regions, exon regions, regions containing both introns and exons.
图1示出了本发明的一种对目标区域进行测序的方法流程,该方法包括以下步骤:Fig. 1 shows a kind of method flow process of sequencing target region of the present invention, and this method comprises the following steps:
S1.利用特异性引物,对待测样品中的多个目标区域进行扩增,并基于扩增产物构建测序文库;S1. Use specific primers to amplify multiple target regions in the sample to be tested, and construct a sequencing library based on the amplified products;
S2.对测序文库进行单分子扩增,得到与所述多个目标区域对应的多个单分子扩增产物;S2. performing single-molecule amplification on the sequencing library to obtain multiple single-molecule amplification products corresponding to the multiple target regions;
S3.同时对所述多个单分子扩增产物进行高通量基因测序,得到所述多个目标区域的序列信息。S3. Simultaneously performing high-throughput gene sequencing on the multiple single-molecule amplification products to obtain sequence information of the multiple target regions.
本方法通过高通量测序技术对待测样品的目标区域进行深度测序,该方法能同时对待测样品的多个目标区域进行检测,准确得到这些区域的序列信息,包括已知突变和未知突变的各突变位点的变异情况,检测灵敏度高,能精确得出各样品的各突变位点发生变异的频率。此外,通过控制各目标区域扩增产物的大小,可免去了片段化步骤,减少了实验步骤,提高了实验效率,降低了成本。This method uses high-throughput sequencing technology to perform deep sequencing on the target region of the sample to be tested. This method can detect multiple target regions of the sample to be tested at the same time, and accurately obtain the sequence information of these regions, including various mutations of known mutations and unknown mutations. The variation of the mutation site, the detection sensitivity is high, and the frequency of the mutation of each mutation site in each sample can be accurately obtained. In addition, by controlling the size of the amplified product of each target region, the fragmentation step can be eliminated, the experimental steps can be reduced, the experimental efficiency can be improved, and the cost can be reduced.
通过本方法所得的序列信息,可用于各种科学研究,包括但不限于人群序列分析、基因功能研究、蛋白功能研究。The sequence information obtained by this method can be used in various scientific researches, including but not limited to population sequence analysis, gene function research, and protein function research.
需要说明的是:It should be noted:
步骤S1中所述待测样品为能提取核酸的任意形式的样品,包括但不限于:全血、血清、血浆和组织样品;所述组织样品包括但不限于:石蜡包埋组织、新鲜组织和冰冻切片。The sample to be tested in step S1 is any form of sample that can extract nucleic acid, including but not limited to: whole blood, serum, plasma and tissue samples; the tissue samples include but not limited to: paraffin-embedded tissue, fresh tissue and frozen slice.
步骤S1所得测序文库中,存在多种测序文库分子,对测序文库进行单分子扩增,即是指,将测序文库中的多种文库分子,以极微量(甚至单分子)的形式在空间上隔离(但这些文库分子整体上还是属于同一个反应体系),并且在各自的空间内实现扩增,以提升各种分子在后续测序反应中的信号。In the sequencing library obtained in step S1, there are multiple sequencing library molecules, and the single-molecule amplification of the sequencing library means that the various library molecules in the sequencing library are spaced in the form of a very small amount (or even a single molecule). Isolate (but these library molecules still belong to the same reaction system as a whole), and achieve amplification in their respective spaces to improve the signal of various molecules in subsequent sequencing reactions.
现有技术中,Sanger测序技术每次只能对一个样品的某一段区域进行测序,要实现对多个目标区域的测序,只能是通过多次反应来实现。而在本发明中,测序文库中的各个分子经过单分子扩增后,每个测序文库分子均形成单分子拷贝阵列,各单分子拷贝阵列在进行高通量基因测序时处于不同的位置,使得测序引物与单分子拷贝阵列之间的杂交,以及在酶作用下的延伸反应可同时进行,相互之间互不干扰。因此,可以同时对大量的(成百万上千万,甚至更多的)单分子拷贝阵列同时进行测序反应,然后通过采集相应的信号,进而获得所需的序列信息,且测序的灵敏度较Sanger更高。In the prior art, the Sanger sequencing technology can only sequence a certain region of a sample at a time, and the sequencing of multiple target regions can only be achieved through multiple reactions. However, in the present invention, after each molecule in the sequencing library undergoes single-molecule amplification, each molecule of the sequencing library forms a single-molecule copy array, and each single-molecule copy array is in a different position during high-throughput gene sequencing, so that The hybridization between the sequencing primer and the single-molecule copy array, and the extension reaction under the action of the enzyme can be carried out simultaneously without mutual interference. Therefore, a large number (millions, tens of millions, or even more) of single-molecule copy arrays can be sequenced at the same time, and then the required sequence information can be obtained by collecting the corresponding signals, and the sensitivity of sequencing is higher than that of Sanger higher.
其中,步骤S1所述的特异性引物与目标区域完全互补或部分互补。Wherein, the specific primer described in step S1 is fully or partially complementary to the target region.
进一步的,每对针对目标区域的引物中,至少一条引物与目标区域部分互补,且该引物的5’端带有第二标签序列。该第二标签序列,用于在扩增目标区域过程中,对不同待测样品的目标区域扩增产物进行标记。Further, in each pair of primers for the target region, at least one primer is partially complementary to the target region, and the 5' end of the primer has a second tag sequence. The second tag sequence is used to mark the amplified products of the target region of different samples to be tested during the process of amplifying the target region.
该第二标签序列,优选为带有特定序列的核酸分子,其碱基数不限。该第二标签序列的碱基数优选为3~20,更优选为4~10。The second tag sequence is preferably a nucleic acid molecule with a specific sequence, and its number of bases is not limited. The base number of the second tag sequence is preferably 3-20, more preferably 4-10.
此外,所述特异性引物还可带有其它标记物,包括但不限于:生物素标记、多聚组氨酸标记、抗原、抗体,从而使得目标区域扩增产物的纯化极为方便。In addition, the specific primers can also carry other labels, including but not limited to: biotin label, polyhistidine label, antigen, antibody, so that the purification of the amplification product of the target region is very convenient.
另外,同一待测样品的不同目标区域的扩增可同时进行或分别独立进行或部分同时进行。在具体的实验过程中,可根据需要选用上述任一种方案进行。In addition, the amplification of different target regions of the same sample to be tested can be performed simultaneously or independently or partly simultaneously. In the specific experimental process, any of the above-mentioned schemes can be selected according to the needs.
如果分别对各目标区域进行分别扩增的话,能够通过测定扩增产物的量来确保步骤S1中用于构建目标区域测序文库的目标区域扩增产物的分子数保持一致,不会因为扩增步骤而导致同一待测样品的不同目标区域拷贝数不同,进而影响后续的测序反应结果。If each target region is amplified separately, the amount of the amplified product can be determined to ensure that the number of molecules of the amplified product of the target region used to construct the target region sequencing library in step S1 remains consistent, and there will be no possibility of amplification due to the amplification step. As a result, the copy numbers of different target regions of the same sample to be tested are different, which in turn affects the subsequent sequencing reaction results.
当然,当各目标区域的大小相近,GC含量也相近的时候,通过合理的设计引物,利用多重PCR技术,步骤S1可对多个目标区域进行同时扩增,且保证各目标区域之间的扩增效率保持基本一致,这样就能够有效的提高实验效率,降低反应的成本。Of course, when the size of each target region is similar and the GC content is also similar, by rationally designing primers and using multiplex PCR technology, step S1 can simultaneously amplify multiple target regions, and ensure the amplification between target regions. The increasing efficiency remains basically the same, so that the experimental efficiency can be effectively improved and the cost of the reaction can be reduced.
当待测样品有多个时,不同样品的目标区域的扩增必须分别进行。When there are multiple samples to be tested, the amplification of target regions of different samples must be performed separately.
在一个实施例中,步骤S1的具体实现过程是:In one embodiment, the specific implementation process of step S1 is:
S11.利用特异性引物,对待测样品中的多个目标区域进行扩增,得到与所述多个目标区域对应的扩增产物;S11. Using specific primers, amplify multiple target regions in the sample to be tested to obtain amplification products corresponding to the multiple target regions;
S12.利用接头元件,与所述多个目标区域对应的扩增产物进行连接,得到测序文库;所述接头元件采用平末端接头、突出末端接头、带茎环结构的接头和分叉接头中的至少一种。S12. Using adapter elements to connect the amplified products corresponding to the multiple target regions to obtain a sequencing library; the adapter elements use blunt end adapters, protruding end adapters, adapters with stem-loop structures, and bifurcated adapters. at least one.
需要说明的是:It should be noted:
步骤S11中,对同一待测样品中的多个目标区域的扩增,可同时进行或分别独立进行或部分同时进行。可根据实际情况,如:扩增特异性引物的退火温度,扩增的目标区域的大小、GC含量,扩增的目标区域的数量等,进行相应的调整。不同待测样品的目标区域的扩增必须分别进行。In step S11, the amplification of multiple target regions in the same sample to be tested can be performed simultaneously or independently or partly simultaneously. Corresponding adjustments can be made according to the actual situation, such as: the annealing temperature of the amplification-specific primers, the size of the amplified target region, the GC content, the number of amplified target regions, etc. Amplification of target regions for different samples to be tested must be performed separately.
步骤S12中,所述接头元件与扩增产物的连接方式,可以采用多种方式实现,包括接头元件与扩增产物直接连接,或对扩增产物进行处理之后再进行连接。In step S12, the connection of the adapter element and the amplification product can be realized in various ways, including direct connection of the adapter element and the amplification product, or connection after the amplification product is processed.
步骤S12中的接头元件,用于构建测序文库,可包括一种或多种接头。The linker element in step S12 is used to construct the sequencing library, and may include one or more linkers.
其中,步骤S12所述接头元件中的至少一个接头包含有第一标签序列,该第一标签序列,用于在文库构建过程中,对不同待测样品的测序文库进行标记。这样,在分别获得目标区域测序文库后,不同的待测样品的目标区域测序文库可以混合在同一个反应体系中,进行单分子扩增反应,进而同时进行高通量测序。提高测序反应的效率,降低了样品检测的成本。Wherein, at least one adapter among the adapter elements described in step S12 includes a first tag sequence, and the first tag sequence is used to tag the sequencing libraries of different samples to be tested during the library construction process. In this way, after the target region sequencing libraries are respectively obtained, the target region sequencing libraries of different samples to be tested can be mixed in the same reaction system for single-molecule amplification reaction, and then high-throughput sequencing can be performed at the same time. Improve the efficiency of the sequencing reaction and reduce the cost of sample detection.
该第一标签序列优选为带有特定碱基序列的核酸分子,其碱基数不限。进一步的,所述第一标签序列的碱基数为3~20,这样,每次至少能够对43个样品进行同时检测。在综合考虑各种情况后,如:标签的特异性、接头的成本、接头的长度等,第一标签序列的碱基数优选为4~10。通过第二标签和第一标签的结合,本发明的发明每次能够检测至少43×43个样品,即4096个样品。The first tag sequence is preferably a nucleic acid molecule with a specific base sequence, and the number of bases is not limited. Further, the number of bases in the first tag sequence is 3-20, so that at least 43 samples can be detected simultaneously at a time. After comprehensive consideration of various conditions, such as: specificity of the tag, cost of the linker, length of the linker, etc., the base number of the first tag sequence is preferably 4-10. Through the combination of the second label and the first label, the invention of the present invention can detect at least 4 3 × 4 3 samples each time, that is, 4096 samples.
接头的修饰方式有多种,包括但不限于:被生物素化或甲基化,或同时被生物素化和甲基化。在一个实施例中,该接头被生物素化,并与未生物素化的片段化产物连接,生物素的存在有利于构建的测序文库的分离纯化。在另一个实施例中,该接头被甲基化,并与未甲基化的片段化产物连接,然后用仅切割甲基化DNA的限制性内切酶消化连接产物,因为只有成功连接的连接产物才能被切割,所以只有成功连接的连接物被切割,从而确保酶切产物的单一性。Linkers can be modified in various ways, including but not limited to: being biotinylated or methylated, or both biotinylated and methylated. In one embodiment, the linker is biotinylated and connected to non-biotinylated fragmentation products, and the presence of biotin facilitates the separation and purification of the constructed sequencing library. In another embodiment, the adapter is methylated and ligated to unmethylated fragments, and the ligation products are then digested with a restriction enzyme that cleaves only methylated DNA, since only successfully ligated ligation Only the product can be cleaved, so only the successfully ligated linker is cleaved, thus ensuring the uniqueness of the digested product.
接头的结构形式也有多种,包括但不限于:平末端接头、突出末端接头、带茎环结构的接头和分叉接头。构建测序文库过程中可以使用一种或多种接头。其中,突出末端接头、带茎环结构的接头和分叉接头均能够有效防止在连接过程中多个接头自连现象的发生。针对接头的上述接头形式,以下将提供多个实施例。There are also various structural forms of linkers, including but not limited to: blunt-ended linkers, protruding-end linkers, linkers with stem-loop structures, and bifurcated linkers. One or more adapters can be used in the construction of a sequencing library. Among them, protruding end joints, joints with stem-loop structures and bifurcated joints can effectively prevent the self-connection of multiple joints during the connection process. Regarding the above joint forms of the joint, several embodiments will be provided below.
在第一实施例中,接头元件采用平末端接头,该接头是双链完全互补的核酸分子。In a first embodiment, the linker element employs a blunt-ended linker, which is a double-stranded fully complementary nucleic acid molecule.
在第二实施例中,接头元件采用突出末端接头,该接头是双链核酸分子,该双链核酸分子至少包括一突出末端。该突出末端的碱基数无具体限制,优选为1~10个碱基。根据该双链核酸分子的结构,突出末端接头可分成两类,分别是单突出末端接头、双突出末端接头。In a second embodiment, the adapter element is an overhanging end adapter, which is a double-stranded nucleic acid molecule including at least one overhanging end. The number of bases at the protruding end is not particularly limited, and is preferably 1 to 10 bases. According to the structure of the double-stranded nucleic acid molecule, the overhanging end adapters can be divided into two types, namely single overhanging end adapters and double overhanging end adapters.
如图2所示的单突出末端接头,其一端为平末端,另一端为突出末端。其中带有分叉接头的单突出末端接头能够防止接头自连。为了防止一端为平末端的单突出末端接头自连,可对平末端上的3’OH进行修饰(包括但不限于用氨基封闭羟基),或将平末端上的5’磷酸基团去除。A single protruding end fitting as shown in Figure 2 has a blunt end at one end and a protruding end at the other end. Single protruding end fittings with bifurcated fittings prevent self-ligation of the fittings. In order to prevent self-ligation of the single protruding end adapter with a blunt end, the 3'OH on the blunt end can be modified (including but not limited to blocking the hydroxyl with an amino group), or the 5' phosphate group on the blunt end can be removed.
如图3所示的双突出末端接头,其含有两个突出末端,这两个突出末端可在一条核苷酸链上(图3a)或在不同的核苷酸链上(图3b)。当这两个突出末端在不同的核酸链上时,他们相互之间不互补,以防在连接时出现接头自连。A double overhang adapter as shown in Figure 3, which contains two overhangs, which can be on one nucleotide strand (Figure 3a) or on different nucleotide strands (Figure 3b). When the two overhanging ends are on different nucleic acid strands, they are not complementary to each other to prevent adapter self-ligation during ligation.
在第三实施例中,接头元件采用带茎环结构的接头,如图4所示。该接头为单链核酸分子,该单链核酸分子包括第一互补配对区1、茎环区2和第二互补配对区3(图4a),第一互补配对区1能够与第二互补配对区3互补配对,且它们形成的互补配对区包括至少一个限制性内切酶识别位点,而通过该酶切识别位点,特定的酶能够将茎环区切开或切除,从而将单链核酸分子变成双链核酸分子,以便于后续的操作。如图4b所示,带茎环结构的接头还可带有突出末端4,该突出末端可位于单链核酸分子的5’端或3’端。突出末端4的存在能够进一步的防止接头自连现象的发生。该突出末端优选为T。In the third embodiment, the joint element adopts a joint with a stem-loop structure, as shown in FIG. 4 . The linker is a single-stranded nucleic acid molecule, which comprises a first complementary pairing region 1, a stem-loop region 2 and a second complementary pairing region 3 (Fig. 4a), and the first complementary pairing region 1 can be combined with the second
在第四实施例中,接头元件采用分叉接头,如图5所示,该接头是双链核酸分子,包括互补区和分叉区,所述分叉区的两条单链各包含至少一个扩增引物结合位点。优选的,所述分叉接头的互补区包括至少一个限制性内切酶识别位点,该酶切识别位点可以在建库过程中酶切形成末端,以便于进行后续的操作。In the fourth embodiment, the joint element adopts a bifurcated joint, as shown in Figure 5, the joint is a double-stranded nucleic acid molecule, including a complementary region and a bifurcated region, and each of the two single strands of the bifurcated region contains at least one Amplify primer binding sites. Preferably, the complementary region of the bifurcation adapter includes at least one restriction endonuclease recognition site, which can be digested to form a terminal during the library construction process, so as to facilitate subsequent operations.
该分叉接头的分叉设计能够在建库过程避免多个接头自连现象的出现;所述分叉区上包含的扩增引物结合位点,可以直接用于结合扩增引物,进行扩增反应The bifurcated design of the bifurcated adapter can avoid the phenomenon of self-connection of multiple adapters during the library construction process; the amplification primer binding site contained on the bifurcated region can be directly used to bind the amplification primer for amplification reaction
其中,所述分叉接头的分叉区的每条链各含有N个核苷酸;优选的,9≤N≤30。其中,所述分叉接头的配对区互补配对的核苷酸对数不限;优选的,互补配对的核苷酸对数为7~15,更优选的,互补配对的核苷酸对数为9~13。Wherein, each strand of the bifurcation region of the bifurcation adapter contains N nucleotides; preferably, 9≤N≤30. Wherein, there is no limit to the number of nucleotide pairs in the pairing region of the bifurcated junction; preferably, the number of nucleotide pairs in complementary pairing is 7 to 15, and more preferably, the number of nucleotide pairs in complementary pairing is 9~13.
其中,所述分叉接头的配对区的3’末端为突出末端或平末端。优选的,所述分叉接头的配对区的3’末端为突出末端,该突出末端可与所述片段化产物的粘性末端互补配对,提高了连接效率,以利于构建测序文库反应的顺利进行。Wherein, the 3' end of the pairing region of the bifurcated linker is a protruding end or a blunt end. Preferably, the 3' end of the pairing region of the bifurcated adapter is a protruding end, and the protruding end can be complementary paired with the sticky end of the fragmented product, which improves the ligation efficiency and facilitates the smooth progress of the sequencing library construction reaction.
优选的,所述分叉接头为T末端分叉接头,该接头的配对区的3’末端为突出末端,且突出末端最后一个碱基为T;例如图6所示的T末端分叉接头,图中N为A、T、C、G碱基中的任一种。Preferably, the bifurcated joint is a T-terminal bifurcated joint, the 3' end of the pairing region of the joint is a protruding end, and the last base of the protruding end is T; for example, the T-terminal bifurcated joint shown in Figure 6, N in the figure is any one of A, T, C, and G bases.
优选的,所述分叉接头的配对区的3’末端为突出末端,且突出末端的核苷酸包括通用碱基。其中,突出末端的碱基数无特殊限制,优选为1~4。例如图7所示的分叉接头,图中N为A、T、C、G碱基中的任一种,X为通用碱基。Preferably, the 3' end of the pairing region of the bifurcation linker is an overhang end, and the nucleotides at the overhang end include universal bases. Among them, the number of bases at the protruding end is not particularly limited, but is preferably 1-4. For example, in the bifurcated linker shown in Figure 7, N in the figure is any one of A, T, C, and G bases, and X is a universal base.
优选的,所述分叉接头为双脱氧接头,该接头的配对区的3’末端为平末端,且3’末端最后一个核苷酸为带有双脱氧碱基的核苷酸;例如图8所示的双脱氧分叉接头,图中N为A、T、C、G碱基中的任一种,dd表示该3’末端最后一个核苷酸为带有双脱氧碱基的胞嘧啶核苷酸。Preferably, the bifurcated linker is a dideoxy linker, the 3' end of the pairing region of the linker is a blunt end, and the last nucleotide at the 3' end is a nucleotide with a dideoxy base; for example, Figure 8 In the dideoxy bifurcated linker shown, N in the figure is any one of A, T, C, and G bases, and dd indicates that the last nucleotide at the 3' end is a cytosine nucleus with a dideoxy base glycosides.
应当说明的是,上述接头元件只是部分实施例,并不用以限制本发明的保护范围。It should be noted that the above joint components are only some examples, and are not intended to limit the protection scope of the present invention.
关于步骤S12的实现方式:About the implementation of step S12:
在一个实施例中,步骤S12采用接头元件与扩增产物直接连接的方式,构建出测序文库。In one embodiment, step S12 constructs a sequencing library by directly connecting adapter elements with amplification products.
在另一个实施例中,当目标区域扩增产物较大时或者需要构建的目标区域测序文库较小时,则对扩增产物进行片段化处理之后,片段化产物再与接头元件进行连接,构建测序文库,如图9所示,步骤S12包括以下步骤:In another embodiment, when the amplified product of the target region is large or the sequencing library of the target region to be constructed is small, the amplified product is fragmented, and then the fragmented product is ligated with adapter elements to construct a sequencing library. Library, as shown in Figure 9, step S12 comprises the following steps:
S121.对与所述多个目标区域对应的扩增产物进行片段化,得到片段化产物;S121. Fragment the amplified products corresponding to the multiple target regions to obtain fragmented products;
S122.利用接头元件,与片段化产物进行连接,构建测序文库。S122. Using the adapter element to connect the fragmented product to construct a sequencing library.
通过片段化步骤S121将目标区域扩增产物变成较小的片段化产物,从而有助于对目标区域的进一步深度测序。此外,还可以通过片段化处理,将不同的扩增产物变成长短相似的片段化产物,能够有助于后续的统一测序。Through the fragmentation step S121, the amplified product of the target region is converted into smaller fragmented products, thereby facilitating further deep sequencing of the target region. In addition, different amplification products can be transformed into fragmented products of similar length through fragmentation processing, which can facilitate subsequent unified sequencing.
需要说明的是:It should be noted:
步骤S121中,所述片段化目标区域扩增产物的方法有多种,包括但不限于:超声法、喷雾法、化学剪切法和酶切法。可根据实际情况,采用相适应的方法进行实验。In step S121, there are various methods for fragmenting the amplified product of the target region, including but not limited to: ultrasonic method, spray method, chemical shearing method and enzyme cleavage method. According to the actual situation, the experiment can be carried out with an appropriate method.
所述片段化处理之后还可包括片段化产物的分离纯化以及末端修饰的步骤。根据测序的片段长度需要,对于片段化得到的核酸片段,进行目的核酸片段的分离纯化,分离方法可以采用常用方法,如凝胶电泳、蔗糖梯度或氯化铯梯度沉降、柱层析分离等。根据所使用的片段化方法,对所得的目的核酸片段进一步的末端修饰,包括但不限于:磷酸化或去磷酸化、末端补平和末端加A,以便于后续的接头元件连接。上述目的核酸片段长短不限,优选为25bp~500bp,更优选为30bp~200bp,更优选为40~100bp。After the fragmentation treatment, the steps of separation and purification of fragmentation products and terminal modification may also be included. According to the length of the sequenced fragments, for the fragmented nucleic acid fragments, separate and purify the target nucleic acid fragments. The separation methods can be commonly used methods, such as gel electrophoresis, sucrose gradient or cesium chloride gradient sedimentation, column chromatography separation, etc. According to the fragmentation method used, the further terminal modification of the obtained target nucleic acid fragments includes but not limited to: phosphorylation or dephosphorylation, end filling and A at the end, so as to facilitate subsequent linker element ligation. The above-mentioned target nucleic acid fragment is not limited in length, preferably 25bp-500bp, more preferably 30bp-200bp, more preferably 40-100bp.
步骤S121所述片段化产物的长度不限,优选为25bp~500bp,更优选为30bp~200bp,更优选为40bp~100bp。在实现对目标区域的测序的前提下,随着目标区域测序文库分子中含有的目标区域片段长度的减短,高通量测序技术的对目标区域的测序深度加深;而测序深度越深,即对目标区域的每一个碱基位置的测序次数越多,测序结果越准确,对样品中的少量突变的检测就越灵敏;这样就能够有效防止因为样品中带有突变的目标区域的比例偏低,而导致该突变的测序信号的绝对值偏低,发生测序结果不准确的现象。The length of the fragmented product in step S121 is not limited, preferably 25bp-500bp, more preferably 30bp-200bp, more preferably 40bp-100bp. On the premise of realizing the sequencing of the target region, as the length of the target region fragments contained in the target region sequencing library molecule shortens, the sequencing depth of the target region of the high-throughput sequencing technology will be deepened; and the deeper the sequencing depth, that is The more times of sequencing for each base position of the target region, the more accurate the sequencing result, and the more sensitive the detection of a small number of mutations in the sample; this can effectively prevent the low proportion of the target region with mutations in the sample , and the absolute value of the sequencing signal leading to the mutation is low, resulting in inaccurate sequencing results.
为了实现对目标区域测序文库大小的限制,可在步骤S121或S122之后,对片段化产物或目标区域测序文库进行分离纯化。分离纯化的方法有多种,包括但不限于:凝胶方法、蔗糖梯度或氯化铯梯度沉降和柱层析分离。可根据实际情况,采用相适应的方法进行实验。In order to limit the size of the sequencing library of the target region, after step S121 or S122, the fragmented products or the sequencing library of the target region can be separated and purified. There are many separation and purification methods, including but not limited to: gel method, sucrose gradient or cesium chloride gradient sedimentation and column chromatography separation. According to the actual situation, the experiment can be carried out with an appropriate method.
根据上述接头元件,针对步骤S122,下面将通过多个实施例和附图对本步骤进行进一步的说明。According to the above joint components, with regard to step S122, this step will be further described below through multiple embodiments and drawings.
在本发明的一个实施例中,直接在片段化产物的两端接上接头形成测序文库。In one embodiment of the present invention, adapters are directly connected to both ends of the fragmented products to form a sequencing library.
所述接头可采用上述的平末端接头、突出末端接头、带茎环结构的接头和分叉接头中的至少一种。The joint can be at least one of the above-mentioned blunt end joint, protruding end joint, joint with stem-loop structure and bifurcated joint.
在本发明的另一个实施例中,如图10所示,步骤S122具体可由以下步骤实现:In another embodiment of the present invention, as shown in FIG. 10, step S122 can be specifically implemented by the following steps:
S1221.利用第一接头与片段化产物的两端连接,得第一连接产物;S1221. Using the first linker to connect the two ends of the fragmented product to obtain a first ligated product;
S1222.环化第一连接产物,得环化产物;S1222. Cyclizing the first ligated product to obtain a cyclized product;
S1223.II s型限制性内切酶酶切环化产物,得酶切产物;S1223. Type II s restriction endonuclease digests the cyclization product to obtain the digested product;
S1224.在酶切产物两端接上第二接头和第三接头,得测序文库。S1224. Connecting a second linker and a third linker to both ends of the digested product to obtain a sequencing library.
在步骤S1221中,所述第一接头可采用平末端接头、突出末端接头、带茎环结构的接头和分叉接头中的一种,所述第一接头包含有II s型限制性内切酶酶切识别位点;所述的II s型限制性内切酶为切割位点在识别序列之外的限制性内切酶,包括但不限于:Acu I、Alw I、Bbs I、BbV I、Bcc I、BceA I、BciV I、BfuA I、Bmr I、Bpm I、BpuE I、Bsa I、BseM II、BseR I、Bsg I、BsmA I、BsmB I、BsmF I、BspCN I、BspM I、BspQ I、BtgZ I、Ear I、Eci I、EcoP15 I、Fau I、Fok I、Hga I、Hph I、HpyAV、MboⅡ、Mly I、Mme I、Mnl I、NmeAIII、Ple I、Sap I、SfaN I和TspDT I,优选为Acu I、Bsg I、EcoP15 I或Mme I。In step S1221, the first linker can be one of a blunt end linker, a protruding end linker, a linker with a stem-loop structure, and a bifurcated linker, and the first linker contains a type II s restriction endonuclease Enzyme cutting recognition site; the type II s restriction endonuclease is a restriction endonuclease whose cutting site is outside the recognition sequence, including but not limited to: Acu I, Alw I, Bbs I, BbV I, Bcc I, BceA I, BciV I, BfuA I, Bmr I, Bpm I, BpuE I, Bsa I, BseM II, BseR I, Bsg I, BsmA I, BsmB I, BsmF I, BspCN I, BspM I, BspQ I , BtgZ I, Ear I, Eci I, EcoP15 I, Fau I, Fok I, Hga I, Hph I, HpyAV, MboⅡ, Mly I, Mme I, Mnl I, NmeAIII, Ple I, Sap I, SfaN I, and TspDT I, preferably Acu I, Bsg I, EcoP15 I or Mme I.
当所述第一接头为分叉接头时,该II s型限制性内切酶酶切识别位点位于互补区;当所述第一接头为带茎环结构的接头时,限制性内切酶识别位点与茎环结构之间的距离,较II s型限制性内切酶酶切识别位点与茎环结构之间的距离近。When the first linker is a bifurcated linker, the restriction endonuclease recognition site of type II s is located in the complementary region; when the first linker is a linker with a stem-loop structure, the restriction endonuclease The distance between the recognition site and the stem-loop structure is shorter than the distance between the recognition site for restriction endonuclease digestion of type II s and the stem-loop structure.
若片段化产物经过末端修复酶修复,以及末端加A反应,所述第一接头优选为带T末端的分叉接头。若片段化产物只是经过末端修复酶修复,将片段化产物的末端补平,则所述第一接头优选双脱氧接头。If the fragmented product is repaired by an end repair enzyme and A is added to the end, the first linker is preferably a bifurcated linker with a T-terminal. If the fragmented product is only repaired by an end repair enzyme to fill in the ends of the fragmented product, the first linker is preferably a dideoxy linker.
在步骤S1222中,环化第一连接产物有多种实现方式。In step S1222, there are many ways to realize the cyclization of the first ligation product.
在本发明的一实施例中,步骤S1222包括以下步骤:In an embodiment of the present invention, step S1222 includes the following steps:
S12221.利用酶切引物对第一连接产物进行扩增,得扩增产物;S12221. Amplifying the first ligation product with enzyme-cutting primers to obtain an amplification product;
S12222.对扩增产物进行酶切,使得扩增产物形成粘性末端,并自身环化成环化产物。S12222. Enzymatic digestion is performed on the amplified product, so that the amplified product forms a cohesive end and circularizes itself into a circularized product.
所述酶切引物的3’端分别与第一连接产物的两个末端部分互补,5’端均含有限制性内切酶识别位点。经过步骤S12221的扩增形成的扩增产物,其两个末端均含有限制性内切酶识别位点,然后在相应的酶的作用下,使扩增产物的两端形成粘性末端,且这两个粘性末端互补,能够进行自身环化。The 3' ends of the restriction primers are respectively complementary to the two ends of the first ligation product, and the 5' ends both contain restriction endonuclease recognition sites. Both ends of the amplified product formed by amplification in step S12221 contain restriction endonuclease recognition sites, and then under the action of the corresponding enzyme, the two ends of the amplified product form sticky ends, and the two The cohesive ends are complementary and capable of self-circularization.
在本发明的另一个实施例中,所述第一接头包含有2个酶切识别位点,其中一个为限制性内切酶识别位点,用于使步骤S1222形成的第一连接产物的两端在相应的酶的作用下,形成粘性末端,且它们之间互补,能够进行自身环化。另一个为II s型限制性内切酶酶切识别位点,用于在步骤S1223中,利用识别该酶切位点的酶识别环化产物,进行酶切,进而得到酶切产物。In another embodiment of the present invention, the first linker contains two enzyme recognition sites, one of which is a restriction endonuclease recognition site, which is used to make the two ends of the first ligation product formed in step S1222 Under the action of the corresponding enzymes, the cohesive ends are formed, and they are complementary to each other and can undergo self-circularization. The other is a type II s restriction endonuclease recognition site, which is used in step S1223 to recognize the cyclization product with an enzyme that recognizes the restriction endonuclease, perform digestion, and then obtain a digestion product.
应当说明,以上两个实施例仅为本发明中的两种实现环化第一连接产物的实施方案,对于本发明的保护范围不做任何具体的限制。It should be noted that the above two examples are only two implementations of the present invention for realizing the cyclization of the first ligation product, and no specific limitation is imposed on the protection scope of the present invention.
在步骤S1223中,利用能够识别第一接头上的酶切识别位点,并切割环化产物(DNA)但不切割第一接头的酶进行酶切。所述第一接头上的酶切识别位点包括但不限于:Mme I酶切识别位点、Acu I酶切识别位点、Bsg I酶切识别位点。In step S1223, an enzyme that can recognize the restriction recognition site on the first linker and cut the circularization product (DNA) but not the first linker is used for digestion. The restriction recognition sites on the first adapter include but are not limited to: Mme I restriction recognition sites, Acu I restriction recognition sites, and Bsg I restriction recognition sites.
步骤S1224中,所述第二接头可为平末端接头、突出末端接头、带茎环结构的接头和分叉接头中的一种,可带有生物素标记。所述第三接头可为平末端接头、突出末端接头、带茎环结构的接头和分叉接头中的一种。第二接头与第三接头可相同或不同。优选的,所述第二接头和第三接头相同,均为分叉接头。更优选的,所述分叉接头为T末端分叉接头或双脱氧分叉接头。In step S1224, the second linker can be one of a blunt end linker, a protruding end linker, a linker with a stem-loop structure and a bifurcated linker, and can be labeled with biotin. The third joint may be one of a blunt end joint, a protruding end joint, a joint with a stem-loop structure and a bifurcated joint. The second linker and the third linker can be the same or different. Preferably, the second joint and the third joint are the same and are bifurcated joints. More preferably, the bifurcated junction is a T-terminal bifurcated junction or a dideoxy bifurcated junction.
需要特别说明的是,步骤S1222与S1223之间还可包括步骤S1222A:滚环扩增环化产物,得滚环扩增产物。通过步骤S1222A,能够保证后续的酶切步骤S1223有足够的原材料。It should be noted that between steps S1222 and S1223, step S1222A: rolling circle amplification of the circularized product may also be included to obtain a rolling circle amplification product. Through step S1222A, it is possible to ensure sufficient raw materials for the subsequent enzyme digestion step S1223.
又或者,在步骤S1221与S1222之间还可包括步骤S1221A:利用扩增引物对第一连接产物进行扩增,得扩增产物。所述扩增引物分别与第一连接产物两端的接头序列互补。通过步骤S1221A,能够保证后续的环化步骤有足够的原材料。Alternatively, between steps S1221 and S1222, step S1221A may also be included: using amplification primers to amplify the first ligation product to obtain an amplification product. The amplification primers are respectively complementary to the linker sequences at both ends of the first ligation product. Through step S1221A, sufficient raw materials can be ensured for the subsequent cyclization step.
本方案可避免在步骤S1222之后进行滚环扩增,而用步骤S1222之前的步骤S1221A进行普通的PCR扩增代替,可有效减少后续酶切步骤中,II s型限制性内切酶的用量。This protocol can avoid rolling circle amplification after step S1222, and replace it with ordinary PCR amplification in step S1221A before step S1222, which can effectively reduce the amount of restriction endonuclease II s in the subsequent enzyme digestion step.
其中,所述第一接头优选为分叉接头。Wherein, the first joint is preferably a bifurcated joint.
其中,所述分叉接头的互补区可包含至少一个酶切识别位点。所述酶切识别位点可为普通的限制性内切酶识别位点,也可为II s型限制性内切酶酶切识别位点。Wherein, the complementary region of the bifurcated linker may contain at least one enzyme recognition site. The restriction endonuclease recognition site can be a common restriction endonuclease recognition site, and can also be an IIs type restriction endonuclease recognition site.
其中,步骤S1221A所述扩增引物优选为生物素化引物,有利于扩增产物的回收纯化。Wherein, the amplification primer described in step S1221A is preferably a biotinylated primer, which is beneficial to the recovery and purification of the amplification product.
其中,步骤S1221A所述扩增引物带有至少一个特异性酶切识别位点。Wherein, the amplification primer in step S1221A has at least one specific restriction enzyme recognition site.
若扩增引物上所带的特异性酶切识别位点是尿嘧啶碱基,则步骤S1222中利用尿嘧啶特异性切除试剂进行酶切,然后再进行连接环化。If the specific enzyme recognition site carried on the amplification primer is a uracil base, then in step S1222, the uracil-specific excision reagent is used for enzyme digestion, and then ligation and cyclization are performed.
若扩增引物所带特异性酶切识别位点为限制性内切酶识别位点,则步骤S1222中利用相应的限制性内切酶进行酶切,然后再进行连接环化。If the specific restriction endonuclease recognition site carried by the amplification primer is a restriction endonuclease recognition site, then in step S1222, the corresponding restriction endonuclease is used for digestion, and then ligation and circularization are performed.
在本发明的另一个实施例中,如图11所示,步骤S122具体可由以下步骤实现:In another embodiment of the present invention, as shown in FIG. 11, step S122 can be specifically implemented by the following steps:
S1221’.利用第四接头与片段化产物连接,得第二连接产物;S1221'. Use the fourth linker to connect with the fragmented product to obtain a second ligated product;
S1222’.II s型限制性内切酶酶切第二连接产物,得带有第四接头的酶切片段;S1222'. II s type restriction endonuclease digests the second connection product to obtain a fragment with the fourth linker;
S1223’.带有第四接头的酶切片段与第五接头连接,形成测序文库。S1223'. The enzyme-cut fragment with the fourth linker is connected to the fifth linker to form a sequencing library.
需要说明的是:It should be noted:
在步骤S1221’中,所述第四接头可采用平末端接头、突出末端接头、带茎环结构的接头和分叉接头中的一种,所述第四接头包含有II s型限制性内切酶酶切识别位点;当所述第四接头为分叉接头时,该II s型限制性内切酶酶切识别位点位于互补区;当所述第四接头为带茎环结构的接头时,限制性内切酶识别位点与茎环结构之间的距离,较II s型限制性内切酶酶切识别位点与茎环结构之间的距离近。In step S1221', the fourth linker can be one of a blunt end linker, a protruding end linker, a linker with a stem-loop structure, and a bifurcated linker, and the fourth linker contains a type II s restriction Enzyme digestion recognition site; when the fourth joint is a bifurcated joint, the II s type restriction endonuclease recognition site is located in the complementary region; when the fourth joint is a joint with a stem-loop structure When , the distance between the restriction endonuclease recognition site and the stem-loop structure is shorter than the distance between the II s type restriction endonuclease recognition site and the stem-loop structure.
在步骤S1222’中,利用能够识别第四接头上的酶切识别位点,并切割环化产物(DNA)但不切割第四接头的酶进行酶切。所述第四接头上的酶切识别位点包括但不限于:Mme I酶切识别位点、Acu I酶切识别位点、Bsg I酶切识别位点。若片段化产物经过末端修复酶修复,以及末端加A反应,所述第四接头优选为带T末端的分叉接头。若片段化产物只是经过末端修复酶修复,则所述第四接头优选双脱氧接头。In step S1222', an enzyme that can recognize the enzyme recognition site on the fourth linker and cut the circularization product (DNA) but not the fourth linker is used for enzyme digestion. The restriction recognition sites on the fourth adapter include but are not limited to: Mme I restriction recognition sites, Acu I restriction recognition sites, and Bsg I restriction recognition sites. If the fragmented product is repaired by an end-repair enzyme and A is added to the end, the fourth linker is preferably a bifurcated linker with a T-terminal. If the fragmented product is only repaired by an end repair enzyme, the fourth linker is preferably a dideoxy linker.
在步骤S1223’中,所述第五接头可为平末端接头、突出末端接头、带茎环结构的接头和分叉接头中的一种,可带有生物素标记。In step S1223', the fifth linker can be one of a blunt end linker, a protruding end linker, a linker with a stem-loop structure and a bifurcated linker, and can be labeled with biotin.
在本发明的另一个实施例中,步骤S122具体包括以下步骤:In another embodiment of the present invention, step S122 specifically includes the following steps:
S1221”.直接在片段化产物的两端接上带茎环结构的接头,形成带茎环结构的片段化产物;S1221". Directly connect adapters with a stem-loop structure to both ends of the fragmented product to form a fragmented product with a stem-loop structure;
S1222”.利用限制性内切酶,将带茎环结构的片段化产物的茎环区切开或切除,从而形成测序文库。S1222". Use restriction endonuclease to cut or excise the stem-loop region of the fragmented product with stem-loop structure, so as to form a sequencing library.
本技术方案利用带茎环结构的接头,防止了多个接头自连现象的发生。The technical scheme utilizes the joint with the stem-loop structure, which prevents the self-connection phenomenon of multiple joints.
当通过上述几个方案形成的测序文库的数量过少,不利于后续的单分子扩增步骤时,还可以进行以下步骤:When the number of sequencing libraries formed by the above-mentioned schemes is too small, which is not conducive to the subsequent single-molecule amplification steps, the following steps can also be performed:
利用与目标区域测序文库两端的接头部分互补的引物,以所得的测序文库为模版进行扩增,得扩增后的测序文库。该步骤可满足后续实验对目标测序文库的量的要求。Using primers complementary to the adapters at both ends of the sequencing library in the target region, amplifying the obtained sequencing library as a template to obtain an amplified sequencing library. This step can meet the requirements for the amount of the target sequencing library in subsequent experiments.
其中,步骤S2所述的单分子扩增是指对目标区域测序文库中的分子,以极微量(甚至单分子)的形式在空间上隔离(但这些文库分子整体上还是属于同一个反应体系),并且在各自的空间内实现扩增,以提升各种分子在后续测序反应中的信号。所述单分子扩增的方法包括但不限于:乳液PCR(Emulsion PCR,EPCR)、桥式PCR。Wherein, the single-molecule amplification described in step S2 refers to the sequencing of the molecules in the target region in the library, and spatial isolation in the form of a very small amount (even a single molecule) (but these library molecules still belong to the same reaction system as a whole) , and achieve amplification in their respective spaces to enhance the signals of various molecules in subsequent sequencing reactions. The method of single molecule amplification includes but not limited to: emulsion PCR (Emulsion PCR, EPCR), bridge PCR.
所述EPCR最大的特点是可以形成数目庞大的独立反应空间以进行DNA扩增。基本过程是在PCR反应前,将包含PCR所有反应成分的水溶液注入到高速旋转的矿物油表面,水溶液瞬间形成无数个被矿物油包裹的小水滴。这些小水滴就构成了独立的PCR反应空间。理想状态下,每个小水滴只含一个DNA模板(目标区域测序文库分子)和一个磁珠,磁珠上含有与目标区域测序文库分子的共有序列(由接头元件引入)互补的引物,在PCR反应后,磁珠表面就固定有拷贝数目巨大的同来源的DNA模板扩增产物。EPCR的具体步骤可参考PCR amplification from single DNA molecules on magnetic beads in emulsion:application for high-throughput screening of transcription factor targets,TakaakiKojima,Yoshiaki Takei,Miharu Ohtsuka et al,Nucleic Acids Research,2005,Vol.33,No.17;Dual primer emulsion PCR for nextgeneration DNA sequencing,MingYan Xu,Anthony D.Aragon,Monica R.Mascarenas et al,BioTechniques48:409-412(May 2010);BEAMing:single-molecule PCR on microparticles inwater-in-oil emulsions,Frank Diehl,Meng Li,Yiping He,nature methods,Vol.3,No.7,July 2006等。The biggest feature of the EPCR is that it can form a large number of independent reaction spaces for DNA amplification. The basic process is that before the PCR reaction, the aqueous solution containing all the reaction components of the PCR is injected onto the surface of the mineral oil rotating at high speed, and the aqueous solution instantly forms countless small water droplets wrapped by the mineral oil. These small water droplets constitute an independent PCR reaction space. Ideally, each droplet contains only one DNA template (target region sequencing library molecule) and one magnetic bead, which contains a primer complementary to the consensus sequence (introduced by the adapter element) of the target region sequencing library molecule. After the reaction, the amplified product of the same source DNA template with a huge copy number is immobilized on the surface of the magnetic beads. For the specific steps of EPCR, please refer to PCR amplification from single DNA molecules on magnetic beads in emulsion: application for high-throughput screening of transcription factor targets, Takaaki Kojima, Yoshiaki Takei, Miharu Ohtsuka et al,
所述桥式PCR的基本原理是,桥式PCR的引物被固定在固相载体上,PCR过程中PCR扩增产物会被固定在固相载体上,且PCR扩增产物能够与固相载体上的引物互补配对,成桥状,然后互补配对的引物以与其成桥的扩增产物为模板进行扩增。通过控制初始模板加入的量,桥式PCR反应完成后,扩增产物在固相载体上以一簇簇的形式存在,且每一簇的扩增产物为同来源的DNA模板扩增产物。其具体的原理和实施方案可参考以下文献:CN20061009879.X、US6227604。The basic principle of the bridge-type PCR is that the primers of the bridge-type PCR are fixed on the solid-phase carrier, and the PCR amplification product will be fixed on the solid-phase carrier during the PCR process, and the PCR amplification product can be combined with the solid-phase carrier. The primers of the complementary pairs are complementary to form a bridge, and then the complementary paired primers are amplified with the bridged amplification product as a template. By controlling the amount of initial template added, after the bridge PCR reaction is completed, the amplified products exist in clusters on the solid-phase support, and the amplified products of each cluster are amplified products of DNA templates from the same source. For its specific principles and implementations, please refer to the following documents: CN20061009879.X, US6227604.
如前所述,现有技术中,Sanger测序技术由于自身的技术限制,每次只能对一个样品的某一段区域进行测序。为了一次性实现对样品中多个区域的同时检测,本发明在测序方法上采取高通量基因测序方法。高通量基因测序相对Sanger测序法检测序列信息更为方便灵敏,测序文库中的各个分子经过单分子扩增后,每个测序文库分子均形成单分子拷贝阵列,各单分子拷贝阵列在进行高通量基因测序时处于不同的位置,使得测序引物与单分子拷贝阵列之间的杂交,以及在酶作用下的延伸反应可同时进行,相互之间互不干扰。因此,可以同时对大量的(成百万上千万,甚至更多的)单分子拷贝阵列同时进行测序反应,然后通过采集相应的信号,进而准确的获得所需的序列信息,且测序的灵敏度较Sanger更高。尤其是对多个目标区域的扩增产物进行了片段化处理,相当于对相同序列的目标区域分子的每个碱基的测序次数增加了,能够进一步提高测序的灵敏度。As mentioned above, in the prior art, Sanger sequencing technology can only sequence a certain region of a sample at a time due to its own technical limitations. In order to realize the simultaneous detection of multiple regions in the sample at one time, the present invention adopts a high-throughput gene sequencing method in the sequencing method. Compared with Sanger sequencing, high-throughput gene sequencing is more convenient and sensitive to detect sequence information. After each molecule in the sequencing library is amplified by single molecule, each molecule of the sequencing library forms a single-molecule copy array. Throughput gene sequencing is in different positions, so that the hybridization between the sequencing primer and the single-molecule copy array, and the extension reaction under the action of the enzyme can be carried out simultaneously without mutual interference. Therefore, a large number (millions, tens of millions, or even more) of single-molecule copy arrays can be sequenced at the same time, and then the required sequence information can be accurately obtained by collecting the corresponding signals, and the sensitivity of sequencing Higher than Sanger. In particular, the fragmentation of the amplification products of multiple target regions is equivalent to an increase in the number of sequencing of each base of the target region molecules of the same sequence, which can further improve the sensitivity of sequencing.
其中,步骤S3所述的高通量测序技术包括但不限于:基于聚合酶的合成测序法、基于连接酶的连接测序法。Wherein, the high-throughput sequencing technology described in step S3 includes, but is not limited to: polymerase-based sequencing-by-synthesis and ligase-based sequencing-by-ligation.
基于聚合酶的合成测序法是基于带可去除标记的核苷酸进行的。在每次合成反应中,每个模板链至多只能延伸一次,基于聚合酶的合成测序法的大致流程如下:Polymerase-based sequencing-by-synthesis methods are based on removably labeled nucleotides. In each synthesis reaction, each template strand can only be extended once at most. The general flow of the polymerase-based sequencing-by-synthesis method is as follows:
a.测序引物通过互补配对结合在单分子扩增产物共有的已知序列上(该单分子扩增产物固定在引物-固相载体复合物上),在DNA聚合酶的作用下,以带可去除标记的核苷酸进行单碱基延伸合成反应,收集该次加入核苷酸的标记信号,即可得到与测序引物3’最末端碱基互补的单分子扩增产物(固定在引物-固相载体复合物上)的下一位的碱基序列信息。a. Sequencing primers bind to the known sequence shared by the single-molecule amplification product through complementary pairing (the single-molecule amplification product is fixed on the primer-solid phase carrier complex), and under the action of DNA polymerase, the Remove the labeled nucleotides to perform a single base extension synthesis reaction, collect the labeling signal of the added nucleotides, and then obtain a single-molecule amplification product complementary to the 3'most base of the sequencing primer (fixed on the primer-solid The base sequence information of the next bit on the phase-carrier complex).
b.切除可去除标记,然后在DNA聚合酶的作用下,以带可去除标记的核苷酸继续进行单碱基延伸合成反应,收集加入核苷酸的标记信号,即可得到与测序引物3’末端碱基互补的单分子扩增产物的下两位的碱基序列信息。b. Excise the removable marker, and then under the action of DNA polymerase, continue the single-base extension synthesis reaction with the nucleotide with the removable marker, collect the marker signal of the added nucleotide, and then obtain the sequence primer 3 'The base sequence information of the next two bits of the single-molecule amplification product with complementary terminal bases.
重复b步骤,直至不能继续进行合成反应为止,从而获得单分子扩增产物的全部序列信息。Step b is repeated until the synthesis reaction cannot be continued, so as to obtain all sequence information of the single-molecule amplification product.
基于连接酶的连接测序法是均是基于带荧光标记的寡核苷酸探针进行的。其中一种连接酶的连接测序法是基于在特定位置带有荧光标记的寡核苷酸探针进行的,该寡核苷酸探针带有n个碱基,从其5’端数起的第a位带有荧光标记,其中不同的碱基对应特定的荧光标记,因为该寡核苷酸探针的3’端进行了特定的修饰,寡核苷酸探针之间不能直接相互连接,每次连接反应,每个单分子扩增产物只能连接一个寡核苷酸探针。该连接测序法的大致流程如下:Ligase-based ligation sequencing methods are all based on fluorescently labeled oligonucleotide probes. One of the ligase ligation sequencing methods is based on a fluorescently labeled oligonucleotide probe with n bases at a specific position, counting from its 5' end. The a position has a fluorescent label, where different bases correspond to specific fluorescent labels, because the 3' end of the oligonucleotide probe has been specifically modified, and the oligonucleotide probes cannot be directly connected to each other. For each ligation reaction, only one oligonucleotide probe can be ligated to each single-molecule amplification product. The general flow of the ligation sequencing method is as follows:
A.测序引物通过互补配对结合在单分子扩增产物共有的已知序列上(该单分子扩增产物固定在引物-固相载体复合物上)上,利用上述的寡核苷酸探针,在连接酶的作用下,将核酸探针与上述寡核苷酸链连接,然后采集荧光信号,即可得到与单分子扩增产物共有的已知序列的3’末端后第a位碱基序列信息。A. Sequencing primers are bound by complementary pairing on the known sequence shared by the unimolecular amplification product (the unimolecular amplification product is immobilized on the primer-solid phase carrier complex), using the above-mentioned oligonucleotide probes, Under the action of ligase, connect the nucleic acid probe to the above-mentioned oligonucleotide chain, and then collect the fluorescent signal to obtain the a-th base sequence after the 3' end of the known sequence shared by the single-molecule amplification product information.
B.切除寡核苷酸上的荧光标记,在连接酶的作用下,以上述的寡核苷酸探针为原料,继续进行连接反应,然后采集荧光信号,从而得到单分子扩增产物共有的已知序列的3’末端后第2a位的碱基序列信息。B. Cut off the fluorescent label on the oligonucleotide, under the action of ligase, use the above-mentioned oligonucleotide probe as the raw material, continue the ligation reaction, and then collect the fluorescent signal, so as to obtain the common value of the single-molecule amplification product The base sequence information at the 2a position after the 3' end of the known sequence.
重复B步骤,直至不能继续进行连接反应为止,从而获得单分子扩增产物共有的已知序列的3’末端后第a、2a、3a、4a......位的碱基序列信息。Repeat step B until the ligation reaction cannot be continued, so as to obtain the base sequence information of positions a, 2a, 3a, 4a... after the 3' end of the known sequence shared by the unimolecular amplification products.
然后将测序引物及其所连接的寡核苷酸探针从单分子扩增产物上变性洗脱下来,换用与之前的测序引物相比3’末端少一个碱基的引物重复上述反应,从而获得单分子扩增产物共有的已知序列的3’末端后第a-1、2a-1、3a-1、4a-1......位的碱基序列信息。重复此步骤,最后获得单分子扩增产物共有的已知序列的3’末端后第a-(a-1)、2a-(a-1)、3a-(a-1)、4a-(a-1)......位的碱基序列信息,从而获得单链扩增产物的全部序列信息。Then, the sequencing primer and the oligonucleotide probe connected thereto are denatured and eluted from the single-molecule amplification product, and the above reaction is repeated with a primer that has one base less at the 3' end than the previous sequencing primer, thereby Obtain the base sequence information of positions a-1, 2a-1, 3a-1, 4a-1... after the 3' end of the known sequence shared by the unimolecular amplification products. Repeat this step to finally obtain the a-(a-1), 2a-(a-1), 3a-(a-1), 4a-(a -1) The base sequence information of ..., so as to obtain the entire sequence information of the single-stranded amplification product.
另一种连接酶的连接测序法同样也是基于带有荧光标记的寡核苷酸探针进行的,该寡核苷酸探针带有n个碱基,分为h(h≤n)组,同一组寡核苷酸探针的不同荧光标记对应同一特定位置的不同碱基序列,不同组之间的区别在于:不同荧光标记对应的特定位置不同,因为该寡核苷酸探针的3’端进行了特定的修饰,寡核苷酸探针之间不能直接相互连接,每次连接反应,每个单分子扩增产物只能连接一个寡核苷酸探针。该连接测序法的大致流程如下:Another ligase ligation sequencing method is also based on fluorescently labeled oligonucleotide probes, the oligonucleotide probes have n bases and are divided into h (h≤n) groups, Different fluorescent labels of the same group of oligonucleotide probes correspond to different base sequences at the same specific position, and the difference between different groups is that the specific positions corresponding to different fluorescent labels are different, because the 3' of the oligonucleotide probe Specific modification is carried out at the end, and the oligonucleotide probes cannot be directly connected to each other. In each ligation reaction, only one oligonucleotide probe can be connected to each single-molecule amplification product. The general flow of the ligation sequencing method is as follows:
a.测序引物通过互补配对结合在单分子扩增产物共有的已知序列上(该单分子扩增产物固定在引物-固相载体复合物上)上,利用上述寡核苷酸探针中的一组(荧光标记对应的碱基位置为x,x≤h),在连接酶的作用下,将核酸探针与上述寡核苷酸链连接,然后采集荧光信号,即可得到与单链扩增产物共有的已知序列的3’末端后第x位碱基序列信息,将测序引物及其所连接的寡核苷酸探针从单分子扩增产物上变性洗脱下来。a. The sequencing primer is bound to the known sequence shared by the single molecule amplification product (the single molecule amplification product is immobilized on the primer-solid phase carrier complex) by complementary pairing, using the above-mentioned oligonucleotide probe One group (the base position corresponding to the fluorescent label is x, x≤h), under the action of ligase, connect the nucleic acid probe to the above-mentioned oligonucleotide chain, and then collect the fluorescent signal to obtain the The base sequence information at position x after the 3' end of the known sequence common to the amplified products is used to denature and elute the sequencing primers and their connected oligonucleotide probes from the single-molecule amplified products.
b.然后重新将测序引物结合在单分子扩增产物上,换用与a步骤不同的寡核苷酸探针组(荧光标记对应的碱基位置为y,y≤h),在连接酶的作用下,将核酸探针与上述寡核苷酸链连接,然后采集荧光信号,即可得到与单链扩增产物共有的已知序列的3’末端后第y位碱基序列信息,将测序引物及其所连接的寡核苷酸探针从单分子扩增产物上变性洗脱下来。b. Then re-combine the sequencing primers on the single-molecule amplification product, and use an oligonucleotide probe set different from that in step a (the base position corresponding to the fluorescent label is y, y≤h), in the ligase Under the action, the nucleic acid probe is connected to the above-mentioned oligonucleotide chain, and then the fluorescent signal is collected to obtain the y-th base sequence information after the 3' end of the known sequence shared by the single-stranded amplification product, and the sequencing Primers and their attached oligonucleotide probes are denatured and eluted from single-molecule amplification products.
c.重复步骤b,直至h组寡核苷酸探针均分别进行过一次连接反应,从而获得单分子扩增产物共有的已知序列的3’末端后第1、2......、h位的碱基序列信息。c. Repeat step b until the oligonucleotide probes of group h have undergone a ligation reaction respectively, so as to obtain the 1st, 2nd... , The base sequence information of the h position.
换用与之前的测序引物相比3’末端多一个或各通用碱基的引物按上述原理进行反应,能够延长获得的单分子扩增产物共有的已知序列的3’末端碱基序列的读长。Replace the primers with one or more universal bases at the 3' end compared with the previous sequencing primers to carry out the reaction according to the above principle, which can extend the reading of the 3' end base sequence of the known sequence shared by the obtained single molecule amplification products long.
这种基于连接酶的连接测序法的原理和具体实施方案可参考CN200710170507.1。For the principle and specific implementation of this ligase-based ligation sequencing method, please refer to CN200710170507.1.
以检测EGFR基因的外显子19、21为例,本发明提出一实施例,设计用于扩增EGFR基因外显子19、21的特异性引物:19F(SEQ ID NO:1)、19R(SEQID NO:2)、21F(SEQ ID NO:3)、21R(SEQ ID NO:4)。Taking the detection of exons 19 and 21 of the EGFR gene as an example, the present invention proposes an embodiment, designing specific primers for amplifying exons 19 and 21 of the EGFR gene: 19F (SEQ ID NO: 1), 19R ( SEQ ID NO: 2), 21F (SEQ ID NO: 3), 21R (SEQ ID NO: 4).
一、待测样品DNA的提取1. Extraction of DNA from samples to be tested
利用市场上常见的核酸提取试剂盒分别提取全血样品(1至10)、血清样品(11至20)、石蜡组织样品(21至30)的DNA,并分别做上相应的标记。DNA was extracted from whole blood samples (1 to 10), serum samples (11 to 20), and paraffin tissue samples (21 to 30) using common nucleic acid extraction kits on the market, and marked accordingly.
二、目标区域的扩增2. Amplification of the target area
同一样品的EGFR基因外显子19、21的扩增在同一反应体系中进行,反应体系如下:The amplification of exons 19 and 21 of the EGFR gene of the same sample was carried out in the same reaction system, and the reaction system was as follows:
19F(10μM) 2μL;19F (10μM) 2μL;
19R(10μM) 2μL;19R (10μM) 2μL;
21F(10μM) 2μL;21F (10μM) 2μL;
21R(10μM) 2μL;21R (10μM) 2μL;
dNTP(各2.5mM) 4μL;dNTP (2.5mM each) 4μL;
待测样品DNA 20ng;Sample DNA to be tested 20ng;
Ex Taq(5U/μL) 0.25μL;Ex Taq(5U/μL) 0.25μL;
10×Ex Taq Buffer 5μL;10×Ex Taq Buffer 5μL;
ddH2O up to 50μL。ddH 2 O up to 50 μL.
PCR反应条件如下:The PCR reaction conditions are as follows:
95℃3min;95°C for 3 minutes;
94℃30s,58℃30s,72℃30s;重复25个循环;94°C for 30s, 58°C for 30s, 72°C for 30s; repeat 25 cycles;
72℃7min。72°C for 7min.
利用PCR清洁试剂盒,分别对各样品的扩增产物进行清洁,除去未扩增的引物和dNTP,回收扩增产物。The PCR cleaning kit was used to clean the amplified products of each sample, remove unamplified primers and dNTPs, and recover the amplified products.
三、构建目标区域测序文库3. Construction of target region sequencing library
此步骤可有多种实施方案,本发明的一个实施例中,包括以下两个步骤:This step can have multiple implementations, and in one embodiment of the present invention, comprises the following two steps:
1.扩增产物的片段化1. Fragmentation of amplified products
利用超声法进行片段化处理。具体操作为:将每个样品的PCR清洁回收后的扩增产物(约50μL)加入至400μL的TE buffer中,在430W功率条件下超声4s,间隔20s,反复5次,得片段化产物。利用1%琼脂糖凝胶对各样品的片段化产物进行分离纯化,切胶回收大小在40bp至100bp之间的片段化产物。Fragmentation was performed using sonication. The specific operation is as follows: add the amplified product (about 50 μL) recovered by PCR cleaning of each sample into 400 μL TE buffer, sonicate for 4 s under 430 W power, with an interval of 20 s, and
2.构建测序文库2. Construction of sequencing library
在构建目标区域测序文库之前,需对切胶回收的片段化产物分别进行末端修饰,以便于接头元件的连接。在本实施例中,对切胶回收的片段化产物的末端修饰包括磷酸化、末端补平和末端加A反应。Before constructing the sequencing library of the target region, it is necessary to modify the ends of the fragmented products recovered from gel cutting to facilitate the connection of adapter elements. In this example, the end modification of the fragmented products recovered from gel cutting includes phosphorylation, end filling and A reaction at the end.
具体实现如下:The specific implementation is as follows:
1)磷酸化以及末端补平反应1) Phosphorylation and end-filling reactions
体系为:The system is:
切胶回收的片段化产物 20μL(约2000ng);20μL (about 2000ng) of the fragmented product recovered by gel cutting;
10mM dNTP 1.5μL;10mM dNTP 1.5μL;
T4DNA聚合酶(5U/μL) 1μL;T4 DNA polymerase (5U/μL) 1 μL;
Klenow DNA聚合酶 0.1μL;Klenow DNA polymerase 0.1 μL;
T4多核苷酸激酶(10U/μL) 0.5μL;T4 polynucleotide kinase (10U/μL) 0.5μL;
10m MATP 1.5μL;10m MATP 1.5μL;
10×T4连接酶缓冲液 10μL;10×T4 ligase buffer 10μL;
加ddH2O至100μL。Add ddH 2 O to 100 μL.
反应条件为:20℃孵育20min。反应结束后利用回收试剂盒进行纯化回收。The reaction conditions are: incubate at 20°C for 20 min. After the reaction, the recovery kit was used for purification and recovery.
2)末端加A尾2) Add A tail at the end
反应体系为:The reaction system is:
磷酸化以及末端补平后的回收产物 60μL(约1000ng);60μL (about 1000ng) of the recovered product after phosphorylation and end-filling;
Klenow缓冲液(NEB Buffer2) 10μL;Klenow buffer (NEB Buffer2) 10μL;
10mM dATP 2μL;10mM dATP 2μL;
Klenow酶(3’to 5’exo minus,10U/μL) 1μL;Klenow enzyme (3'to 5'exo minus, 10U/μL) 1μL;
加ddH2O至100μL。Add ddH 2 O to 100 μL.
反应条件为:37℃孵育30min。反应结束后利用纯化试剂盒纯化回收。The reaction conditions are: incubate at 37°C for 30 min. After the reaction was completed, it was purified and recovered using a purification kit.
3)连接接头13) Connection connector 1
本实施例中,采用如图6所示的T末端分叉接头作为接头1,同一样品的所使用的T末端分叉接头相同,不同样品的T末端分叉接头不同,区别在于标签序列不同,不同样品对应的标签序列如下表所示。In this example, the T-terminal bifurcated adapter as shown in FIG. 6 is used as the adapter 1. The T-terminal bifurcated adapters used in the same sample are the same, and the T-terminal bifurcated adapters of different samples are different. The difference lies in the different tag sequences. The tag sequences corresponding to different samples are shown in the table below.
在T4连接酶的作用下,上述T末端分叉接头分别与加A尾后纯化回收的产物进行连接,形成带接头1的片段。连接体系为:Under the action of T4 ligase, the above-mentioned T-terminal bifurcated adapters were respectively ligated with the products purified and recovered after adding A tails to form fragments with adapter 1. The connection system is:
加A尾后纯化回收的产物 50μL(约500ng);Add 50μL (about 500ng) of the recovered product after adding A tail;
接头1 2μL(约3000ng);Connector 1 2μL (about 3000ng);
10mM ATP 5μL;10mM ATP 5μL;
T4DNA连接酶(10U/μL) 1μL;T4 DNA ligase (10U/μL) 1 μL;
10×T4连接酶缓冲液 10μL;10×T4 ligase buffer 10μL;
加ddH2O至100μL。Add ddH 2 O to 100 μL.
反应条件为:16℃孵育4h以上。反应结束后利用纯化试剂盒纯化回收。The reaction conditions are: incubate at 16°C for more than 4h. After the reaction was completed, it was purified and recovered using a purification kit.
4)PCR扩增带接头1的片段4) PCR amplification of the fragment with adapter 1
扩增体系为:The amplification system is:
带接头1的片段 约300ng;The fragment with linker 1 is about 300ng;
10×Ex Taq Buffer 100μL;10×Ex Taq Buffer 100μL;
Primer F(100μM,SEQ ID NO:11) 10μL;Primer F (100 μM, SEQ ID NO: 11) 10 μL;
Primer R(100μM,SEQ ID NO:12) 10μL;Primer R (100 μM, SEQ ID NO: 12) 10 μL;
Ex Taq(5U/μL) 7.5μL;Ex Taq(5U/μL) 7.5μL;
ddH2O up to 1000μL。ddH 2 O up to 1000 μL.
PCR反应条件如下:The PCR reaction conditions are as follows:
95℃3min;95°C for 3 minutes;
94℃30s,58℃30s,72℃30s;重复25个循环;94°C for 30s, 58°C for 30s, 72°C for 30s; repeat 25 cycles;
72℃7min。72°C for 7min.
利用PCR清洁试剂盒,分别对各样品的扩增产物进行清洁,除去未扩增的引物和dNTP,回收带接头1的片段的扩增产物。The PCR cleaning kit was used to clean the amplified products of each sample, remove unamplified primers and dNTPs, and recover the amplified products of the fragments with adapter 1.
5)II s型限制性内切酶酶切5) Type II s restriction endonuclease digestion
利用Acu I酶切回收后的带接头1的片段的扩增产物,反应体系如下:The amplified product of the recovered fragment with adapter 1 was digested with Acu I, and the reaction system was as follows:
回收后的带接头1的片段的扩增产物 60μL(3~5μg);60 μL (3-5 μg) of the amplified product of the recovered fragment with adapter 1;
10×NEB Buffer 4 8μL;10×NEB Buffer 4 8μL;
Acu I(NEB) 2μL(10U);Acu I(NEB) 2μL(10U);
SAM(3.2mM) 1μL(终浓度50μM);SAM (3.2mM) 1μL (final concentration 50μM);
ddH2O up to 80μL。ddH 2 O up to 80 μL.
反应条件:37℃孵育1h。反应结束后利用纯化试剂盒纯化回收酶切产物。Reaction conditions: incubate at 37°C for 1 hour. After the reaction, the digested product was purified and recovered using a purification kit.
6)连接接头26) Connect connector 2
利用如图7所示的突出末端分叉接头作为接头2,与回收的酶切产物进行连接,得测序文库,连接体系如下:Use the protruding end bifurcated adapter as shown in Figure 7 as the adapter 2, and connect with the recovered enzyme digestion product to obtain a sequencing library. The connection system is as follows:
回收的酶切产物 2μg;Recovered digested product 2 μg;
接头2 1μL(10pM);Connector 2 1μL (10pM);
10T4DNA连接酶(3U/μL) 1μL;10T4 DNA ligase (3U/μL) 1 μL;
10×T4连接酶缓冲液 2μL;10×T4 ligase buffer 2μL;
加ddH2O至100μL。Add ddH 2 O to 100 μL.
连接条件,14℃孵育2h。反应结束后利用纯化试剂盒纯化回收,然后变性形成单链,得测序文库。In connection conditions, incubate at 14°C for 2h. After the reaction, use a purification kit to purify and recover, and then denature to form a single strand to obtain a sequencing library.
四、对目标区域测序文库进行单分子扩增4. Single-molecule amplification of the target region sequencing library
测定(三)步骤所获得30个目标区域测序文库各自的浓度,然后按同等浓度混合,然后进行单分子扩增,得单分子扩增产物,所述单分子扩增的方法可采用EPCR或桥式PCR。Measure the respective concentrations of the 30 target region sequencing libraries obtained in step (3), then mix them at the same concentration, and then perform single-molecule amplification to obtain single-molecule amplification products. The single-molecule amplification method can be EPCR or bridge PCR.
优选为EPCR扩增,单分子扩增引物优选为:SEQ ID NO:5和SEQ ID NO:6。It is preferably EPCR amplification, and the unimolecular amplification primers are preferably: SEQ ID NO: 5 and SEQ ID NO: 6.
五、对单分子扩增产物进行高通量测序5. High-throughput sequencing of single-molecule amplification products
所述测序方法可采用基于合成酶的合成测序法,也可采用基于连接酶的连接测序法。对测序结果进行生物信息学分析,即可得到上述30个样品的EGFR基因的外显子19和21的序列信息。分析序列结果可知,除样品23的EGFR基因的外显子19发生突变外(序列信息见SEQ ID NO:8,突变率为6.2%),其它样品的各外显子均为野生型。The sequencing method may adopt a synthetase-based sequencing-by-synthesis method or a ligase-based sequencing-by-ligation method. By performing bioinformatics analysis on the sequencing results, the sequence information of exons 19 and 21 of the EGFR gene of the above-mentioned 30 samples can be obtained. Analysis of the sequence results shows that, except for the mutation in exon 19 of the EGFR gene in sample 23 (see SEQ ID NO: 8 for sequence information, the mutation rate is 6.2%), all exons in other samples are wild type.
应当说明的是,本实施例只是本发明的一个具体实施例,对本发明无任何限定作用,例如EGFR基因的特异性引物可用其他的经过合理设计的引物进行替换;除此之外,本实施例中的多个步骤均可参照前述的方法进行替换,在此不再赘述。It should be noted that this embodiment is only a specific embodiment of the present invention, and does not have any limiting effect on the present invention. For example, the specific primer of EGFR gene can be replaced by other rationally designed primers; in addition, this embodiment Many of the steps in can be replaced with reference to the aforementioned method, and will not be repeated here.
针对本发明的对目标区域进行测序的方法的灵敏度,本发明采用下述实施例进行验证。Regarding the sensitivity of the method for sequencing the target region of the present invention, the present invention uses the following examples to verify.
通过常规设计,构建分别含有EGFR基因外显子19野生型序列(SEQ IDNO:7)和突变型序列(SEQ ID NO:8)的质粒;含有EGFR基因外显子21野生型序列(SEQ ID NO:9)和突变型序列(SEQ ID NO:10)的质粒。By conventional design, construct plasmids containing respectively EGFR gene exon 19 wild-type sequence (SEQ ID NO: 7) and mutant sequence (SEQ ID NO: 8); containing EGFR gene exon 21 wild-type sequence (SEQ ID NO : 9) and the plasmid of the mutant sequence (SEQ ID NO: 10).
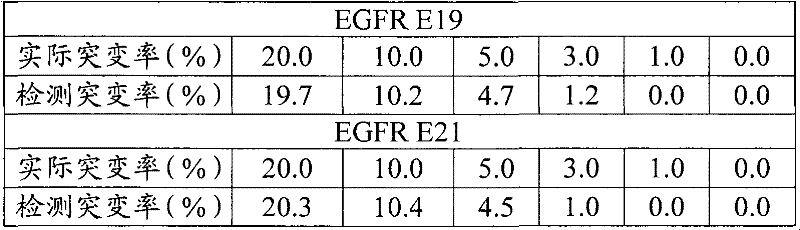
然后配制突变率分别为20%、10%、5%、3%、1%、0%的质粒混合液。以含有1000个拷贝质粒的上述质粒混合液为模板,然后按参照上述实施例的方法进行检测。检测结果如下表所示:Then prepare plasmid mixtures with mutation rates of 20%, 10%, 5%, 3%, 1%, and 0%, respectively. The above-mentioned plasmid mixture containing 1000 copies of the plasmid was used as a template, and then the detection was carried out according to the method referring to the above-mentioned examples. The test results are shown in the table below:
结果显示,本发明的对目标区域进行测序的方法的最低检测限为3%左右,在5%及以上的检测结果与实际情况基本一致。The results show that the minimum detection limit of the method for sequencing the target region of the present invention is about 3%, and the detection results of 5% and above are basically consistent with the actual situation.
一种对目标区域进行测序的试剂盒,包括:A kit for sequencing a region of interest, comprising:
特异性引物,用于对待测样品中的多个目标区域进行扩增;Specific primers for amplifying multiple target regions in the sample to be tested;
接头元件,用于与扩增产物结合构建测序文库。Linker elements are used to combine with amplification products to construct sequencing libraries.
该试剂盒用于目标区域测序,能同时对样本的多个目标区域进行深度测序,准确得出这些目标区域的序列信息,包括已知突变和位置突变的各突变位点的变异情况,精确得出各样品的各突变位点发生变异的频率,检测灵敏度高,能用于目标区域的大规模人群筛查,确定该目标区域的各种基因型的比例。The kit is used for the sequencing of target regions, and can perform deep sequencing on multiple target regions of the sample at the same time, and accurately obtain the sequence information of these target regions, including the variation of each mutation site with known mutations and positional mutations. The mutation frequency of each mutation site in each sample can be obtained, and the detection sensitivity is high, which can be used for large-scale population screening in the target area to determine the proportion of various genotypes in the target area.
其中,所述接头元件用于构建目标区域测序文库,包括至少一种或多种接头。Wherein, the linker element is used to construct a target region sequencing library, including at least one or more linkers.
其中,所述接头元件可采用多种形式,如:平末端接头、突出末端接头、带茎环结构的接头、分叉接头中的至少一种。Wherein, the joint element may adopt various forms, such as at least one of a blunt end joint, a protruding end joint, a joint with a stem-loop structure, and a bifurcated joint.
其中,所述接头可被甲基化或生物素化,或同时被生物素化和甲基化。Wherein, the linker can be methylated or biotinylated, or biotinylated and methylated simultaneously.
其中,所述接头元件中的至少一个接头包含有第一标签序列,用于在文库构建过程中,对不同待测样品的测序文库进行标记。所述第一标签序列优选为带有特定碱基序列的核酸分子,其碱基数不限,优选为3~20,更优选为4~20。Wherein, at least one adapter in the adapter element includes a first tag sequence, which is used to mark the sequencing libraries of different samples to be tested during the library construction process. The first tag sequence is preferably a nucleic acid molecule with a specific base sequence, and the number of bases is not limited, preferably 3-20, more preferably 4-20.
其中,所述的特异性引物与目标区域互补或部分互补。Wherein, the specific primer is complementary or partially complementary to the target region.
进一步的,每个目标区域对应的特异性引物中,至少一条引物与该目标区域部分互补,该引物的5’端包含有第二标签序列,用于在扩增目标区域过程中,对不同待测样品的目标区域扩增产物进行标记。所述第二标签序列优选为带有特定碱基序列的核酸分子,其碱基长度不限。Further, among the specific primers corresponding to each target region, at least one primer is partially complementary to the target region, and the 5' end of the primer contains a second tag sequence, which is used to treat different target regions in the process of amplifying the target region. The amplified product of the target region of the test sample is labeled. The second tag sequence is preferably a nucleic acid molecule with a specific base sequence, and its base length is not limited.
更进一步的,所述第二标签序列的碱基数为3~20,更优选为4~10。Furthermore, the number of bases in the second tag sequence is 3-20, more preferably 4-10.
其中,所述试剂盒还可以包括PCR扩增试剂,所述PCR扩增试剂包括PCR酶、dNTP、PCR缓冲液、Mg2+溶液。Wherein, the kit may also include PCR amplification reagents, and the PCR amplification reagents include PCR enzyme, dNTP, PCR buffer, and Mg 2+ solution.
其中,所述试剂盒还可包括用于识别接头元件上的酶切识别位点的酶。Wherein, the kit may further include an enzyme for recognizing the restriction enzyme recognition site on the linker element.
应当说明的是,通过本发明的方法或试剂盒检测得到的基因序列信息,可用于各种科学研究,包括但不限于人群序列分析、基因功能研究、蛋白功能研究。例如,结合后续进一步的分子生物学试验、临床试验、临床观察及综合数据的统计分析等,从而实现多种科研目的,包括但不限于:某特定人群的某些特定基因的基因型分布、各基因的突变类型与相关生理活动的关系等。It should be noted that the gene sequence information detected by the method or kit of the present invention can be used in various scientific research, including but not limited to population sequence analysis, gene function research, and protein function research. For example, combined with subsequent further molecular biology tests, clinical trials, clinical observations and statistical analysis of comprehensive data, etc., to achieve a variety of scientific research purposes, including but not limited to: genotype distribution of certain specific genes in a specific population, various The relationship between gene mutation types and related physiological activities, etc.
以上所述仅为本发明的较佳实施例而已,并不用以限制本发明,凡在本发明的精神和原则之内所作的任何修改、等同替换和改进等,均应包含在本发明的保护范围之内。The above descriptions are only preferred embodiments of the present invention, and are not intended to limit the present invention. Any modifications, equivalent replacements and improvements made within the spirit and principles of the present invention should be included in the protection of the present invention. within range.
Claims (11)
Priority Applications (1)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| CN2011103919902A CN102373288B (en) | 2011-11-30 | 2011-11-30 | Method and kit for sequencing target areas |
Applications Claiming Priority (1)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| CN2011103919902A CN102373288B (en) | 2011-11-30 | 2011-11-30 | Method and kit for sequencing target areas |
Publications (2)
| Publication Number | Publication Date |
|---|---|
| CN102373288A true CN102373288A (en) | 2012-03-14 |
| CN102373288B CN102373288B (en) | 2013-12-11 |
Family
ID=45792515
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| CN2011103919902A Expired - Fee Related CN102373288B (en) | 2011-11-30 | 2011-11-30 | Method and kit for sequencing target areas |
Country Status (1)
| Country | Link |
|---|---|
| CN (1) | CN102373288B (en) |
Cited By (21)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN102373287A (en) * | 2011-11-30 | 2012-03-14 | 盛司潼 | Method and kit for detecting lung cancer susceptibility gene |
| CN102628079A (en) * | 2012-03-31 | 2012-08-08 | 盛司潼 | Method for constructing sequencing library by cyclizing method |
| CN104450869A (en) * | 2013-09-12 | 2015-03-25 | 中国科学院上海巴斯德研究所 | Dideoxyribonucleoside modified primer, method, reaction system and application thereof to mutation detection |
| CN104480534A (en) * | 2014-12-29 | 2015-04-01 | 深圳华因康基因科技有限公司 | Rapid library building method |
| CN104695027A (en) * | 2013-12-06 | 2015-06-10 | 中国科学院北京基因组研究所 | Sequencing library, preparation and application thereof |
| CN104745679A (en) * | 2013-12-31 | 2015-07-01 | 北京贝瑞和康生物技术有限公司 | Method and kit for non-invasive detection of EGFR (epidermal growth factor receptor) gene mutation |
| CN104790042A (en) * | 2015-05-05 | 2015-07-22 | 天津诺禾致源生物信息科技有限公司 | High-throughput sequencing library and building method thereof |
| WO2016058121A1 (en) * | 2014-10-13 | 2016-04-21 | 深圳华大基因科技有限公司 | Nucleic acid fragmentation method and sequence combination |
| CN106011230A (en) * | 2016-05-10 | 2016-10-12 | 人和未来生物科技(长沙)有限公司 | Primer composition for detecting fragmentized DNA target area and application thereof |
| CN106192018A (en) * | 2015-05-07 | 2016-12-07 | 深圳华大基因研究院 | A kind of method of grappling Nest multiplex PCR enrichment DNA target area and test kit |
| WO2018040961A1 (en) * | 2016-08-30 | 2018-03-08 | 广州康昕瑞基因健康科技有限公司 | Method for building library and method for snp typing |
| WO2018040962A1 (en) * | 2016-08-30 | 2018-03-08 | 广州康昕瑞基因健康科技有限公司 | Method for building library and method for snp typing |
| CN107881242A (en) * | 2016-09-29 | 2018-04-06 | 广州康昕瑞基因健康科技有限公司 | EGFR gene detection kit and detection method |
| CN107988347A (en) * | 2017-12-19 | 2018-05-04 | 光瀚健康咨询管理(上海)有限公司 | A kind of detection method of alzheimer's disease tumor susceptibility gene and its application |
| CN108350453A (en) * | 2015-09-11 | 2018-07-31 | 通用医疗公司 | The rough complete querying of nuclease DSB and sequencing (FIND-SEQ) |
| CN108467886A (en) * | 2017-02-23 | 2018-08-31 | 广州康昕瑞基因健康科技有限公司 | Detection method of gene mutation |
| CN110225979A (en) * | 2017-05-23 | 2019-09-10 | 深圳华大基因股份有限公司 | Genome target region enrichment method based on rolling circle amplification and its application |
| CN111662969A (en) * | 2020-05-18 | 2020-09-15 | 北京优吉科技有限公司 | Gene transcription region multi-variable region sequencing method |
| US11725228B2 (en) | 2017-10-11 | 2023-08-15 | The General Hospital Corporation | Methods for detecting site-specific and spurious genomic deamination induced by base editing technologies |
| US11898203B2 (en) | 2018-04-17 | 2024-02-13 | The General Hospital Corporation | Highly sensitive in vitro assays to define substrate preferences and sites of nucleic-acid binding, modifying, and cleaving agents |
| US12104207B2 (en) | 2014-06-23 | 2024-10-01 | The General Hospital Corporation | Genomewide unbiased identification of DSBs evaluated by sequencing (GUIDE-Seq) |
Families Citing this family (1)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN110699425B (en) * | 2019-09-20 | 2024-01-26 | 上海臻迪基因科技有限公司 | Enrichment methods and systems for gene target regions |
Citations (10)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US5710000A (en) * | 1994-09-16 | 1998-01-20 | Affymetrix, Inc. | Capturing sequences adjacent to Type-IIs restriction sites for genomic library mapping |
| WO2001083696A2 (en) * | 2000-04-28 | 2001-11-08 | Digital Gene Tech Inc | Methods for rapid isolation and sequence determination of gene-specific sequences |
| CN101278058A (en) * | 2005-06-23 | 2008-10-01 | 科因股份有限公司 | Improved strategies for sequencing complex genomes using high-throughput sequencing technologies |
| CN101432438A (en) * | 2006-04-04 | 2009-05-13 | 凯津公司 | High throughput detection of molecular markers based on AFLP and high throughput sequencing |
| CN101434988A (en) * | 2007-11-16 | 2009-05-20 | 深圳华因康基因科技有限公司 | High throughput oligonucleotide sequencing method |
| WO2010039991A2 (en) * | 2008-10-02 | 2010-04-08 | The Texas A&M University System | Method of generating informative dna templates for high-throughput sequencing applications |
| CN101965410A (en) * | 2008-02-27 | 2011-02-02 | 霍夫曼-拉罗奇有限公司 | Systems and methods for improved nucleic acid processing for generating sequenceable libraries |
| WO2011074960A1 (en) * | 2009-12-17 | 2011-06-23 | Keygene N.V. | Restriction enzyme based whole genome sequencing |
| US20110189677A1 (en) * | 2010-02-03 | 2011-08-04 | Massachusetts Institute Of Technology | Methods For Preparing Sequencing Libraries |
| CN102212612A (en) * | 2011-03-23 | 2011-10-12 | 上海美吉生物医药科技有限公司 | Constructing method of double-end library for high throughput 454 sequencing |
-
2011
- 2011-11-30 CN CN2011103919902A patent/CN102373288B/en not_active Expired - Fee Related
Patent Citations (10)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US5710000A (en) * | 1994-09-16 | 1998-01-20 | Affymetrix, Inc. | Capturing sequences adjacent to Type-IIs restriction sites for genomic library mapping |
| WO2001083696A2 (en) * | 2000-04-28 | 2001-11-08 | Digital Gene Tech Inc | Methods for rapid isolation and sequence determination of gene-specific sequences |
| CN101278058A (en) * | 2005-06-23 | 2008-10-01 | 科因股份有限公司 | Improved strategies for sequencing complex genomes using high-throughput sequencing technologies |
| CN101432438A (en) * | 2006-04-04 | 2009-05-13 | 凯津公司 | High throughput detection of molecular markers based on AFLP and high throughput sequencing |
| CN101434988A (en) * | 2007-11-16 | 2009-05-20 | 深圳华因康基因科技有限公司 | High throughput oligonucleotide sequencing method |
| CN101965410A (en) * | 2008-02-27 | 2011-02-02 | 霍夫曼-拉罗奇有限公司 | Systems and methods for improved nucleic acid processing for generating sequenceable libraries |
| WO2010039991A2 (en) * | 2008-10-02 | 2010-04-08 | The Texas A&M University System | Method of generating informative dna templates for high-throughput sequencing applications |
| WO2011074960A1 (en) * | 2009-12-17 | 2011-06-23 | Keygene N.V. | Restriction enzyme based whole genome sequencing |
| US20110189677A1 (en) * | 2010-02-03 | 2011-08-04 | Massachusetts Institute Of Technology | Methods For Preparing Sequencing Libraries |
| CN102212612A (en) * | 2011-03-23 | 2011-10-12 | 上海美吉生物医药科技有限公司 | Constructing method of double-end library for high throughput 454 sequencing |
Cited By (31)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN102373287A (en) * | 2011-11-30 | 2012-03-14 | 盛司潼 | Method and kit for detecting lung cancer susceptibility gene |
| CN102628079A (en) * | 2012-03-31 | 2012-08-08 | 盛司潼 | Method for constructing sequencing library by cyclizing method |
| CN102628079B (en) * | 2012-03-31 | 2014-02-19 | 盛司潼 | Method for constructing sequencing library by cyclizing method |
| CN104450869A (en) * | 2013-09-12 | 2015-03-25 | 中国科学院上海巴斯德研究所 | Dideoxyribonucleoside modified primer, method, reaction system and application thereof to mutation detection |
| CN104450869B (en) * | 2013-09-12 | 2020-07-03 | 中国科学院上海巴斯德研究所 | Dideoxynucleoside modified primer method, reaction system and application thereof in mutation detection |
| CN104695027A (en) * | 2013-12-06 | 2015-06-10 | 中国科学院北京基因组研究所 | Sequencing library, preparation and application thereof |
| CN104745679A (en) * | 2013-12-31 | 2015-07-01 | 北京贝瑞和康生物技术有限公司 | Method and kit for non-invasive detection of EGFR (epidermal growth factor receptor) gene mutation |
| US12104207B2 (en) | 2014-06-23 | 2024-10-01 | The General Hospital Corporation | Genomewide unbiased identification of DSBs evaluated by sequencing (GUIDE-Seq) |
| US10344317B2 (en) | 2014-10-13 | 2019-07-09 | Mgi Tech Co., Ltd | Method and a sequence combination for producing nucleic acid fragments |
| WO2016058121A1 (en) * | 2014-10-13 | 2016-04-21 | 深圳华大基因科技有限公司 | Nucleic acid fragmentation method and sequence combination |
| CN104480534A (en) * | 2014-12-29 | 2015-04-01 | 深圳华因康基因科技有限公司 | Rapid library building method |
| CN104790042A (en) * | 2015-05-05 | 2015-07-22 | 天津诺禾致源生物信息科技有限公司 | High-throughput sequencing library and building method thereof |
| CN106192018B (en) * | 2015-05-07 | 2020-03-24 | 深圳华大智造科技有限公司 | Method and kit for enriching DNA target region by anchoring nested multiplex PCR |
| CN106192018A (en) * | 2015-05-07 | 2016-12-07 | 深圳华大基因研究院 | A kind of method of grappling Nest multiplex PCR enrichment DNA target area and test kit |
| CN108350453A (en) * | 2015-09-11 | 2018-07-31 | 通用医疗公司 | The rough complete querying of nuclease DSB and sequencing (FIND-SEQ) |
| US11028429B2 (en) | 2015-09-11 | 2021-06-08 | The General Hospital Corporation | Full interrogation of nuclease DSBs and sequencing (FIND-seq) |
| CN106011230A (en) * | 2016-05-10 | 2016-10-12 | 人和未来生物科技(长沙)有限公司 | Primer composition for detecting fragmentized DNA target area and application thereof |
| CN108300764B (en) * | 2016-08-30 | 2021-11-09 | 武汉康昕瑞基因健康科技有限公司 | Library building method and SNP typing method |
| WO2018040962A1 (en) * | 2016-08-30 | 2018-03-08 | 广州康昕瑞基因健康科技有限公司 | Method for building library and method for snp typing |
| WO2018040961A1 (en) * | 2016-08-30 | 2018-03-08 | 广州康昕瑞基因健康科技有限公司 | Method for building library and method for snp typing |
| CN108300764A (en) * | 2016-08-30 | 2018-07-20 | 广州康昕瑞基因健康科技有限公司 | A kind of banking process and SNP classifying methods |
| CN107881242A (en) * | 2016-09-29 | 2018-04-06 | 广州康昕瑞基因健康科技有限公司 | EGFR gene detection kit and detection method |
| CN108467886A (en) * | 2017-02-23 | 2018-08-31 | 广州康昕瑞基因健康科技有限公司 | Detection method of gene mutation |
| CN108467886B (en) * | 2017-02-23 | 2022-01-21 | 武汉康昕瑞基因健康科技有限公司 | Gene mutation detection method |
| CN110225979A (en) * | 2017-05-23 | 2019-09-10 | 深圳华大基因股份有限公司 | Genome target region enrichment method based on rolling circle amplification and its application |
| CN110225979B (en) * | 2017-05-23 | 2024-05-31 | 深圳华大基因股份有限公司 | Method for enriching genomic target regions based on rolling circle amplification and its application |
| US11725228B2 (en) | 2017-10-11 | 2023-08-15 | The General Hospital Corporation | Methods for detecting site-specific and spurious genomic deamination induced by base editing technologies |
| CN107988347A (en) * | 2017-12-19 | 2018-05-04 | 光瀚健康咨询管理(上海)有限公司 | A kind of detection method of alzheimer's disease tumor susceptibility gene and its application |
| US11898203B2 (en) | 2018-04-17 | 2024-02-13 | The General Hospital Corporation | Highly sensitive in vitro assays to define substrate preferences and sites of nucleic-acid binding, modifying, and cleaving agents |
| US11976324B2 (en) | 2018-04-17 | 2024-05-07 | The General Hospital Corporation | Highly sensitive in vitro assays to define substrate preferences and sites of nucleic-acid binding, modifying, and cleaving agents |
| CN111662969A (en) * | 2020-05-18 | 2020-09-15 | 北京优吉科技有限公司 | Gene transcription region multi-variable region sequencing method |
Also Published As
| Publication number | Publication date |
|---|---|
| CN102373288B (en) | 2013-12-11 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| CN102373288B (en) | Method and kit for sequencing target areas | |
| CN102586423B (en) | Method and kit for detecting susceptibility gene of colorectal cancer | |
| JP7500549B2 (en) | Methods for detecting nucleic acids | |
| JP6766236B2 (en) | Nucleic acid probe and genome fragment detection method | |
| CN102373287A (en) | Method and kit for detecting lung cancer susceptibility gene | |
| KR102307230B1 (en) | Multiplex detection of nucleic acids | |
| EP3068883B1 (en) | Compositions and methods for identification of a duplicate sequencing read | |
| CN105714383B (en) | A kind of sequencing library construction method and reagent based on the reverse probe of molecule | |
| KR102592367B1 (en) | Systems and methods for clonal replication and amplification of nucleic acid molecules for genomic and therapeutic applications | |
| CN102586420A (en) | Method and kit for assaying breast cancer susceptibility genes | |
| CN108611398A (en) | Genotyping is carried out by new-generation sequencing | |
| CN110079592B (en) | High throughput sequencing-targeted capture of target regions for detection of genetic mutations and known, unknown gene fusion types | |
| TWI868882B (en) | Methods for accurate parallel detection and quantification of nucleic acids | |
| CN106554955B (en) | Method and kit for constructing sequencing library of PKHD1 gene mutation and application thereof | |
| KR20220130592A (en) | A high-sensitivity method for accurate parallel quantitation of nucleic acids | |
| KR20220130591A (en) | Methods for accurate parallel quantification of nucleic acids in dilute or non-purified samples | |
| US20250145988A1 (en) | Methods of enriching nucleic acids | |
| CN115873922B (en) | A single-cell full-length transcript library construction and sequencing method | |
| CN103374759A (en) | Method for detecting symbolic SNP (Single Nucleotide Polymorphism) of lung cancer metastasis and application thereof | |
| CN112080555A (en) | DNA methylation detection kit and detection method | |
| TWI899628B (en) | Highly sensitive methods for accurate parallel quantification of variant nucleic acids | |
| CN102586472A (en) | Method and kit for assaying hepatitis B virus DNA (deoxyribonucleic acid) sequence | |
| CN111718981A (en) | ACE gene detection primer set, detection kit and detection method | |
| CN102533991A (en) | Method and kit for detecting drinking gene | |
| HK40072445A (en) | Highly sensitive methods for accurate parallel quantification of nucleic acids |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| C06 | Publication | ||
| PB01 | Publication | ||
| C10 | Entry into substantive examination | ||
| SE01 | Entry into force of request for substantive examination | ||
| C14 | Grant of patent or utility model | ||
| GR01 | Patent grant | ||
| C41 | Transfer of patent application or patent right or utility model | ||
| TR01 | Transfer of patent right |
Effective date of registration: 20160927 Address after: 430040 No. 388, No. 21, No. three, biological medicine accelerator, two hi tech Road, East Lake New Technology Development Zone, Hubei, Wuhan Patentee after: WUHAN CONSIDERIN GENE & HEALTH TECHNOLOGY Co.,Ltd. Address before: 518057, Guangdong, Nanshan District hi tech Zone, two science and Technology Park, two software park, 11, 4, building 402, north room, Shenzhen Patentee before: Sheng Sichong |
|
| CF01 | Termination of patent right due to non-payment of annual fee | ||
| CF01 | Termination of patent right due to non-payment of annual fee |
Granted publication date: 20131211 |