Advertisement
- Corrigendum (January 2021)
Brief Report Free access | 10.1172/JCI61745
Transient telomere dysfunction induces chromosomal instability and promotes carcinogenesis
Yvonne Begus-Nahrmann,1 Daniel Hartmann,2,3,4 Johann Kraus,5 Parisa Eshraghi,2 Annika Scheffold,2 Melanie Grieb,4,5 Volker Rasche,6 Peter Schirmacher,7 Han-Wong Lee,8 Hans A. Kestler,5 André Lechel,2 and K. Lenhard Rudolph2
1Institute for Molecular Oncology, University Medical Center, GÖttingen, Germany. 2Institute of Molecular Medicine and Max-Planck-Research Department on Stem Cell Aging, Ulm University, Ulm, Germany. 3Department of Surgery, Technische Universität München, Munich, Germany. 4International Graduate School in Molecular Medicine, 5Research Group Bioinformatics and Systems Biology, Institute of Neural Information Processing, and 6Experimental Cardiovascular Imaging, Core Facility Small Animal MRI, Ulm University, Ulm, Germany. 7Institute of Pathology, University Hospital Heidelberg, Heidelberg, Germany. 8Department of Biochemistry, Yonsei University, Seoul, Republic of Korea.
Address correspondence to: K. Lenhard Rudolph or André Lechel, Institute of Molecular Medicine and Max-Planck-Research Group on Stem Cell Aging, Ulm University, 89081 Ulm, Germany. Phone: 49.731.503.6100; Fax: 49.731.503.6102; E-mail: KLRudolph@fli-leibniz.de (K.L. Rudolph). Phone: 49.731.503.6110; Fax: 49.731.503.6102; E-mail: Andre.Lechel@uni-ulm.de (A. Lechel).
Authorship note: Yvonne Begus-Nahrmann and Daniel Hartmann contributed equally to this work.
Find articles by Begus-Nahrmann, Y. in: JCI | PubMed | Google Scholar
1Institute for Molecular Oncology, University Medical Center, GÖttingen, Germany. 2Institute of Molecular Medicine and Max-Planck-Research Department on Stem Cell Aging, Ulm University, Ulm, Germany. 3Department of Surgery, Technische Universität München, Munich, Germany. 4International Graduate School in Molecular Medicine, 5Research Group Bioinformatics and Systems Biology, Institute of Neural Information Processing, and 6Experimental Cardiovascular Imaging, Core Facility Small Animal MRI, Ulm University, Ulm, Germany. 7Institute of Pathology, University Hospital Heidelberg, Heidelberg, Germany. 8Department of Biochemistry, Yonsei University, Seoul, Republic of Korea.
Address correspondence to: K. Lenhard Rudolph or André Lechel, Institute of Molecular Medicine and Max-Planck-Research Group on Stem Cell Aging, Ulm University, 89081 Ulm, Germany. Phone: 49.731.503.6100; Fax: 49.731.503.6102; E-mail: KLRudolph@fli-leibniz.de (K.L. Rudolph). Phone: 49.731.503.6110; Fax: 49.731.503.6102; E-mail: Andre.Lechel@uni-ulm.de (A. Lechel).
Authorship note: Yvonne Begus-Nahrmann and Daniel Hartmann contributed equally to this work.
Find articles by Hartmann, D. in: JCI | PubMed | Google Scholar
1Institute for Molecular Oncology, University Medical Center, GÖttingen, Germany. 2Institute of Molecular Medicine and Max-Planck-Research Department on Stem Cell Aging, Ulm University, Ulm, Germany. 3Department of Surgery, Technische Universität München, Munich, Germany. 4International Graduate School in Molecular Medicine, 5Research Group Bioinformatics and Systems Biology, Institute of Neural Information Processing, and 6Experimental Cardiovascular Imaging, Core Facility Small Animal MRI, Ulm University, Ulm, Germany. 7Institute of Pathology, University Hospital Heidelberg, Heidelberg, Germany. 8Department of Biochemistry, Yonsei University, Seoul, Republic of Korea.
Address correspondence to: K. Lenhard Rudolph or André Lechel, Institute of Molecular Medicine and Max-Planck-Research Group on Stem Cell Aging, Ulm University, 89081 Ulm, Germany. Phone: 49.731.503.6100; Fax: 49.731.503.6102; E-mail: KLRudolph@fli-leibniz.de (K.L. Rudolph). Phone: 49.731.503.6110; Fax: 49.731.503.6102; E-mail: Andre.Lechel@uni-ulm.de (A. Lechel).
Authorship note: Yvonne Begus-Nahrmann and Daniel Hartmann contributed equally to this work.
Find articles by Kraus, J. in: JCI | PubMed | Google Scholar
1Institute for Molecular Oncology, University Medical Center, GÖttingen, Germany. 2Institute of Molecular Medicine and Max-Planck-Research Department on Stem Cell Aging, Ulm University, Ulm, Germany. 3Department of Surgery, Technische Universität München, Munich, Germany. 4International Graduate School in Molecular Medicine, 5Research Group Bioinformatics and Systems Biology, Institute of Neural Information Processing, and 6Experimental Cardiovascular Imaging, Core Facility Small Animal MRI, Ulm University, Ulm, Germany. 7Institute of Pathology, University Hospital Heidelberg, Heidelberg, Germany. 8Department of Biochemistry, Yonsei University, Seoul, Republic of Korea.
Address correspondence to: K. Lenhard Rudolph or André Lechel, Institute of Molecular Medicine and Max-Planck-Research Group on Stem Cell Aging, Ulm University, 89081 Ulm, Germany. Phone: 49.731.503.6100; Fax: 49.731.503.6102; E-mail: KLRudolph@fli-leibniz.de (K.L. Rudolph). Phone: 49.731.503.6110; Fax: 49.731.503.6102; E-mail: Andre.Lechel@uni-ulm.de (A. Lechel).
Authorship note: Yvonne Begus-Nahrmann and Daniel Hartmann contributed equally to this work.
Find articles by Eshraghi, P. in: JCI | PubMed | Google Scholar
1Institute for Molecular Oncology, University Medical Center, GÖttingen, Germany. 2Institute of Molecular Medicine and Max-Planck-Research Department on Stem Cell Aging, Ulm University, Ulm, Germany. 3Department of Surgery, Technische Universität München, Munich, Germany. 4International Graduate School in Molecular Medicine, 5Research Group Bioinformatics and Systems Biology, Institute of Neural Information Processing, and 6Experimental Cardiovascular Imaging, Core Facility Small Animal MRI, Ulm University, Ulm, Germany. 7Institute of Pathology, University Hospital Heidelberg, Heidelberg, Germany. 8Department of Biochemistry, Yonsei University, Seoul, Republic of Korea.
Address correspondence to: K. Lenhard Rudolph or André Lechel, Institute of Molecular Medicine and Max-Planck-Research Group on Stem Cell Aging, Ulm University, 89081 Ulm, Germany. Phone: 49.731.503.6100; Fax: 49.731.503.6102; E-mail: KLRudolph@fli-leibniz.de (K.L. Rudolph). Phone: 49.731.503.6110; Fax: 49.731.503.6102; E-mail: Andre.Lechel@uni-ulm.de (A. Lechel).
Authorship note: Yvonne Begus-Nahrmann and Daniel Hartmann contributed equally to this work.
Find articles by Scheffold, A. in: JCI | PubMed | Google Scholar
1Institute for Molecular Oncology, University Medical Center, GÖttingen, Germany. 2Institute of Molecular Medicine and Max-Planck-Research Department on Stem Cell Aging, Ulm University, Ulm, Germany. 3Department of Surgery, Technische Universität München, Munich, Germany. 4International Graduate School in Molecular Medicine, 5Research Group Bioinformatics and Systems Biology, Institute of Neural Information Processing, and 6Experimental Cardiovascular Imaging, Core Facility Small Animal MRI, Ulm University, Ulm, Germany. 7Institute of Pathology, University Hospital Heidelberg, Heidelberg, Germany. 8Department of Biochemistry, Yonsei University, Seoul, Republic of Korea.
Address correspondence to: K. Lenhard Rudolph or André Lechel, Institute of Molecular Medicine and Max-Planck-Research Group on Stem Cell Aging, Ulm University, 89081 Ulm, Germany. Phone: 49.731.503.6100; Fax: 49.731.503.6102; E-mail: KLRudolph@fli-leibniz.de (K.L. Rudolph). Phone: 49.731.503.6110; Fax: 49.731.503.6102; E-mail: Andre.Lechel@uni-ulm.de (A. Lechel).
Authorship note: Yvonne Begus-Nahrmann and Daniel Hartmann contributed equally to this work.
Find articles by Grieb, M. in: JCI | PubMed | Google Scholar
1Institute for Molecular Oncology, University Medical Center, GÖttingen, Germany. 2Institute of Molecular Medicine and Max-Planck-Research Department on Stem Cell Aging, Ulm University, Ulm, Germany. 3Department of Surgery, Technische Universität München, Munich, Germany. 4International Graduate School in Molecular Medicine, 5Research Group Bioinformatics and Systems Biology, Institute of Neural Information Processing, and 6Experimental Cardiovascular Imaging, Core Facility Small Animal MRI, Ulm University, Ulm, Germany. 7Institute of Pathology, University Hospital Heidelberg, Heidelberg, Germany. 8Department of Biochemistry, Yonsei University, Seoul, Republic of Korea.
Address correspondence to: K. Lenhard Rudolph or André Lechel, Institute of Molecular Medicine and Max-Planck-Research Group on Stem Cell Aging, Ulm University, 89081 Ulm, Germany. Phone: 49.731.503.6100; Fax: 49.731.503.6102; E-mail: KLRudolph@fli-leibniz.de (K.L. Rudolph). Phone: 49.731.503.6110; Fax: 49.731.503.6102; E-mail: Andre.Lechel@uni-ulm.de (A. Lechel).
Authorship note: Yvonne Begus-Nahrmann and Daniel Hartmann contributed equally to this work.
Find articles by Rasche, V. in: JCI | PubMed | Google Scholar
1Institute for Molecular Oncology, University Medical Center, GÖttingen, Germany. 2Institute of Molecular Medicine and Max-Planck-Research Department on Stem Cell Aging, Ulm University, Ulm, Germany. 3Department of Surgery, Technische Universität München, Munich, Germany. 4International Graduate School in Molecular Medicine, 5Research Group Bioinformatics and Systems Biology, Institute of Neural Information Processing, and 6Experimental Cardiovascular Imaging, Core Facility Small Animal MRI, Ulm University, Ulm, Germany. 7Institute of Pathology, University Hospital Heidelberg, Heidelberg, Germany. 8Department of Biochemistry, Yonsei University, Seoul, Republic of Korea.
Address correspondence to: K. Lenhard Rudolph or André Lechel, Institute of Molecular Medicine and Max-Planck-Research Group on Stem Cell Aging, Ulm University, 89081 Ulm, Germany. Phone: 49.731.503.6100; Fax: 49.731.503.6102; E-mail: KLRudolph@fli-leibniz.de (K.L. Rudolph). Phone: 49.731.503.6110; Fax: 49.731.503.6102; E-mail: Andre.Lechel@uni-ulm.de (A. Lechel).
Authorship note: Yvonne Begus-Nahrmann and Daniel Hartmann contributed equally to this work.
Find articles by Schirmacher, P. in: JCI | PubMed | Google Scholar
1Institute for Molecular Oncology, University Medical Center, GÖttingen, Germany. 2Institute of Molecular Medicine and Max-Planck-Research Department on Stem Cell Aging, Ulm University, Ulm, Germany. 3Department of Surgery, Technische Universität München, Munich, Germany. 4International Graduate School in Molecular Medicine, 5Research Group Bioinformatics and Systems Biology, Institute of Neural Information Processing, and 6Experimental Cardiovascular Imaging, Core Facility Small Animal MRI, Ulm University, Ulm, Germany. 7Institute of Pathology, University Hospital Heidelberg, Heidelberg, Germany. 8Department of Biochemistry, Yonsei University, Seoul, Republic of Korea.
Address correspondence to: K. Lenhard Rudolph or André Lechel, Institute of Molecular Medicine and Max-Planck-Research Group on Stem Cell Aging, Ulm University, 89081 Ulm, Germany. Phone: 49.731.503.6100; Fax: 49.731.503.6102; E-mail: KLRudolph@fli-leibniz.de (K.L. Rudolph). Phone: 49.731.503.6110; Fax: 49.731.503.6102; E-mail: Andre.Lechel@uni-ulm.de (A. Lechel).
Authorship note: Yvonne Begus-Nahrmann and Daniel Hartmann contributed equally to this work.
Find articles by Lee, H. in: JCI | PubMed | Google Scholar
1Institute for Molecular Oncology, University Medical Center, GÖttingen, Germany. 2Institute of Molecular Medicine and Max-Planck-Research Department on Stem Cell Aging, Ulm University, Ulm, Germany. 3Department of Surgery, Technische Universität München, Munich, Germany. 4International Graduate School in Molecular Medicine, 5Research Group Bioinformatics and Systems Biology, Institute of Neural Information Processing, and 6Experimental Cardiovascular Imaging, Core Facility Small Animal MRI, Ulm University, Ulm, Germany. 7Institute of Pathology, University Hospital Heidelberg, Heidelberg, Germany. 8Department of Biochemistry, Yonsei University, Seoul, Republic of Korea.
Address correspondence to: K. Lenhard Rudolph or André Lechel, Institute of Molecular Medicine and Max-Planck-Research Group on Stem Cell Aging, Ulm University, 89081 Ulm, Germany. Phone: 49.731.503.6100; Fax: 49.731.503.6102; E-mail: KLRudolph@fli-leibniz.de (K.L. Rudolph). Phone: 49.731.503.6110; Fax: 49.731.503.6102; E-mail: Andre.Lechel@uni-ulm.de (A. Lechel).
Authorship note: Yvonne Begus-Nahrmann and Daniel Hartmann contributed equally to this work.
Find articles by Kestler, H. in: JCI | PubMed | Google Scholar
1Institute for Molecular Oncology, University Medical Center, GÖttingen, Germany. 2Institute of Molecular Medicine and Max-Planck-Research Department on Stem Cell Aging, Ulm University, Ulm, Germany. 3Department of Surgery, Technische Universität München, Munich, Germany. 4International Graduate School in Molecular Medicine, 5Research Group Bioinformatics and Systems Biology, Institute of Neural Information Processing, and 6Experimental Cardiovascular Imaging, Core Facility Small Animal MRI, Ulm University, Ulm, Germany. 7Institute of Pathology, University Hospital Heidelberg, Heidelberg, Germany. 8Department of Biochemistry, Yonsei University, Seoul, Republic of Korea.
Address correspondence to: K. Lenhard Rudolph or André Lechel, Institute of Molecular Medicine and Max-Planck-Research Group on Stem Cell Aging, Ulm University, 89081 Ulm, Germany. Phone: 49.731.503.6100; Fax: 49.731.503.6102; E-mail: KLRudolph@fli-leibniz.de (K.L. Rudolph). Phone: 49.731.503.6110; Fax: 49.731.503.6102; E-mail: Andre.Lechel@uni-ulm.de (A. Lechel).
Authorship note: Yvonne Begus-Nahrmann and Daniel Hartmann contributed equally to this work.
Find articles by Lechel, A. in: JCI | PubMed | Google Scholar
1Institute for Molecular Oncology, University Medical Center, GÖttingen, Germany. 2Institute of Molecular Medicine and Max-Planck-Research Department on Stem Cell Aging, Ulm University, Ulm, Germany. 3Department of Surgery, Technische Universität München, Munich, Germany. 4International Graduate School in Molecular Medicine, 5Research Group Bioinformatics and Systems Biology, Institute of Neural Information Processing, and 6Experimental Cardiovascular Imaging, Core Facility Small Animal MRI, Ulm University, Ulm, Germany. 7Institute of Pathology, University Hospital Heidelberg, Heidelberg, Germany. 8Department of Biochemistry, Yonsei University, Seoul, Republic of Korea.
Address correspondence to: K. Lenhard Rudolph or André Lechel, Institute of Molecular Medicine and Max-Planck-Research Group on Stem Cell Aging, Ulm University, 89081 Ulm, Germany. Phone: 49.731.503.6100; Fax: 49.731.503.6102; E-mail: KLRudolph@fli-leibniz.de (K.L. Rudolph). Phone: 49.731.503.6110; Fax: 49.731.503.6102; E-mail: Andre.Lechel@uni-ulm.de (A. Lechel).
Authorship note: Yvonne Begus-Nahrmann and Daniel Hartmann contributed equally to this work.
Find articles by Rudolph, K. in: JCI | PubMed | Google Scholar
Authorship note: Yvonne Begus-Nahrmann and Daniel Hartmann contributed equally to this work.
Published May 24, 2012 - More info
J Clin Invest. 2021;122(6):2283–2288. https://doi.org/10.1172/JCI61745.
© 2012 The American Society for Clinical Investigation
Received: November 3, 2011; Accepted: April 6, 2012
Related article:
Abstract
Previous studies in mice have demonstrated antagonistic effects of telomerase loss on carcinogenesis. Telomere attrition can promote genome instability, thereby stimulating initiation of early-stage cancers, but can also inhibit tumorigenesis by promoting permanent cell growth arrest or death. Human cancers likely develop in cell lineages with low levels of telomerase, leading to telomere losses in early lesions, followed by subsequent activation of telomerase. Mouse models constitutively lacking telomerase have thus not addressed how telomere losses within telomerase-proficient cells have an impact on carcinogenesis. Using a novel transgenic mouse model, Begus-Nahrmann et al. demonstrate in this issue of the JCI that transient telomere dysfunction in telomerase-proficient animals is a potent stimulus of tumor formation.
Authors
Jennifer J. Wanat, F. Brad Johnson
-
Abstract
Telomere shortening limits the proliferative capacity of a cell, but perhaps surprisingly, shortening is also known to be associated with increased rates of tumor initiation. A current hypothesis suggests that telomere dysfunction increases tumor initiation by induction of chromosomal instability, but that initiated tumors need to reactivate telomerase for genome stabilization and tumor progression. This concept has not been tested in vivo, since appropriate mouse models were lacking. Here, we analyzed hepatocarcinogenesis in a mouse model of inducible telomere dysfunction on a telomerase-proficient background, in telomerase knockout mice with chronic telomere dysfunction (G3 mTerc–/–), and in WT mice with functional telomeres and telomerase. Transient or chronic telomere dysfunction enhanced the rates of chromosomal aberrations during hepatocarcinogenesis, but only telomerase-proficient mice exhibited significantly increased rates of macroscopic tumor formation in response to telomere dysfunction. In contrast, telomere dysfunction resulted in pronounced accumulation of DNA damage, cell-cycle arrest, and apoptosis in telomerase-deficient liver tumors. Together, these data provide in vivo evidence that transient telomere dysfunction during early or late stages of tumorigenesis promotes chromosomal instability and carcinogenesis in telomerase-proficient mice.
-
Introduction
Telomere shortening limits the proliferative life span of cells by induction of senescence or crisis (1). Both checkpoints represent potent tumor-suppressor mechanisms, and tumor cells need to activate telomere maintenance mechanisms — telomerase activation or alternative mechanisms of telomere lengthening (ALT) — in order to gain immortal growth capacity (2). Genetic deletion or pharmacological inhibition of telomerase significantly suppressed the tumor-forming capacity of human cancer cells as well as the progression of tumors in mouse models (3–7).
In contrast to the role of telomere shortening in tumor suppression, telomere shortening is also associated with increased rates of tumor initiation (3, 7, 8). Studies on late-generation telomerase knockout mice with short telomeres (G3 mTerc–/–) revealed that telomere shortening increases the rate of tumor initiation by induction of chromosomal instability (3, 7), especially when the p53 tumor-suppressor checkpoint is abrogated (9).
The dual role of telomere shortening and telomerase reactivation in tumor initiation and progression led to the model that both processes may occur sequentially during carcinogenesis (3, 6, 7). Specifically, telomere shortening (as a consequence of aging or chronic diseases) leads to the induction of chromosomal instability and cancer initiation followed by activation of telomere maintenance mechanisms (telomerase or ALT) required for tumor progression. This model is supported by correlative data on human carcinogenesis (7, 10). However, the model was not tested directly in vivo due to the lack of appropriate mouse models allowing a transient induction of telomere dysfunction at an early or late stage of tumorigenesis in a telomerase-proficient background.
The initiation of human hepatocellular carcinoma (HCC) is associated with telomere shortening at precancerous disease stages and in early tumors (10–12). However, telomerase reactivation occurs in the vast majority of human HCCs during tumor progression (10, 11). These data suggest that hepatocarcinogenesis in humans proceeds in accordance with the model of tumor initiation by telomere dysfunction followed by telomerase reactivation during tumor progression. To experimentally test the role of this sequence during in vivo tumorigenesis, we analyzed the consequences of transient telomere dysfunction during cancer initiation and progression of carcinogen-induced HCCs in telomerase-proficient mice.
-
Results and Discussion
Transient telomere dysfunction increases hepatocarcinogenesis in telomerase-proficient mice. Previous studies showed that expression of TRF2ΔBΔM (a dominant-negative truncation of TRF2) induces telomere uncapping and chromosomal fusion (13, 14). Here, we generated a liver-specific, doxycycline-inducible TRF2ΔBΔM-transgenic mouse model (Supplemental Methods and Supplemental Figure 1A; supplemental material available online with this article; doi: 10.1172/JCI61745DS1). Control experiments verified that doxycycline administration to Tg(tetopTRF2ΔBΔM)3, rTALAP-1–double-transgenic mice (TTD+) mice on a C57BL/6 background led to a transient expression of TRF2ΔBΔM and the induction of chromosomal instability in the liver (Supplemental Figure 1, B–E). In contrast, single-transgenic (expressing either TALAP-1 or C57BL/6J;Tg[tetopTRF2ΔBΔM]) and WT mice (grouped as TTD–) did not show TRF2ΔBΔM expression or chromosomal fusions when injected with doxycycline (Supplemental Figure 1, B–E).
To evaluate the influence of transient telomere dysfunction on hepatocarcinogenesis, 15-day-old mice were treated with diethylnitrosamine (DEN), which induces HCCs in 10- to 12-month-old mice (3). Transient telomere dysfunction by expression of TRF2ΔBΔM was induced before DEN-treated mice developed tumor lesions at 2 to 3 months (Supplemental Figure 1F). Gene array analysis and hierarchical clustering revealed no global changes in gene expression in mouse livers in acute response to TRF2ΔBΔM expression, suggesting that the inducible transgene did not regulate other biological processes aside from the induction of telomere dysfunction (Supplemental Figure 2).
To investigate the influence of transient telomere dysfunction on tumor initiation, a cohort of male mice was analyzed at 6 months of age. TTD+ mice exhibited a significant increase in the numbers of dysplastic foci (2.857 foci/liver) and macroscopic liver tumors (2.143 tumors/liver) compared with TTD– mice (1.389 foci/liver, P = 0.0339; 0.556 tumors/liver, P = 0.0183; Figure 1, A and B). In agreement with previous studies, hepatocarcinogenesis was delayed in female compared with male mice (3, 15). Transient telomere dysfunction at early stages of tumorigenesis also led to a significant increase in the number of hepatic tumors in 13-month-old female mice (Figure 1, C and D). In addition, transient telomere dysfunction promoted the growth of initiated liver tumors (Figure 1E and Supplemental Figure 3, A and B).
 Figure 1
Figure 1Transient telomere dysfunction promotes hepatocarcinogenesis. Mice were treated with the liver carcinogen DEN at P15. Transient telomere dysfunction was induced by doxycycline-inducible TRF2ΔBΔM expression in TTD+ but not in TTD– mice. (A and B) The incidence of dysplastic foci (A) and macroscopic liver tumors (B) in 6-month-old TTD+ male mice (n = 7) was significantly increased compared with that in age-matched TTD– male mice (n = 18). (C and D) Analysis of foci (C) and tumors (D) in 13-month-old TTD+, TTD–, and G3 mTerc–/– female mice (n = 19, n = 20, and n = 17, respectively). (E) The scatter plot shows the distribution of tumor size from all tumors analyzed in 13-month-old male mice on a logarithmic scale. Tumor volume was significantly increased in TTD+ (n = 136) compared with TDD– mice (n = 233) and G3 mTerc–/– mice (n = 302). (F–H) Transient telomere dysfunction was induced after establishment of macroscopic HCCs in tumors of 12- to 14-month-old female mice. MRI imaging determined the tumor volume before and 1 month after induction of telomere dysfunction. (F) Transient telomere dysfunction in TTD+ mice led to a significant increase in tumor size (n = 31) compared with the control groups (n = 60; P = 0.0002). Dox, doxycycline. (G) Representative MRI images of TTD+ and TTD– mice before and after doxycycline treatment (circles highlight detected tumors). (H) Tumor volumes of both groups prior to doxycycline treatment. Data represent mean ± SEM.
To determine whether transient telomere dysfunction would also promote the progression of established, macroscopic liver tumors, DEN-treated female mice were followed for 11 to 13 months without induction of telomere dysfunction. At this age, the size of liver tumors was determined by MRI followed by doxycycline administration to transiently induce telomere dysfunction in established tumors. Tumor progression was monitored by a second MRI scan 4 weeks after the last injection. This analysis revealed a significant increase in tumor size in TTD+ mice (n = 31) compared with controls (n = 60, P = 0.0002; Figure 1, F–H).
In contrast to the effects of transient telomere dysfunction in telomerase-proficient mice, the formation of macroscopic liver tumors was significantly impaired in third-generation telomerase-deficient mice (G3 mTerc–/–) with dysfunctional telomeres (Figure 1, D and E, and Supplemental Figure 3, A and B). These data indicate that transient telomere dysfunction leads to an acceleration of tumorigenesis in telomerase-proficient mice, but impairment of macroscopic liver tumor formation in telomerase-deficient mice. Previous studies showed that telomerase deficiency per se does not influence tumor formation in Terc–/– G1 mice compared with mTerc+/+ mice (7, 16). Histological analysis of the tumors in TTD+ compared with TTD– mice revealed no influence of telomere dysfunction on the grading/differentiation of liver tumors (Supplemental Figure 4, A and B).
Transient telomere dysfunction induces telomere shortening and chromosomal instability in tumor cells. Quantitative FISH (qFISH) (8, 17) revealed no significant difference in telomere length in nontransformed liver of TTD+ mice compared with TTD– mice (Supplemental Figure 5, A–C). Telomere length was reduced in tumors of TTD+ mice compared with nontransformed liver (Supplemental Figure 5C), suggesting that transient telomere dysfunction enhanced tumor initiation originating from cells with short telomeres. qFISH analysis showed shorter telomeres in HCCs from G3 mTerc–/– and TTD+ mice compared with TTD– HCCs (Figure 2, A–C). However, the percentage of cells with critically short telomeres was significantly higher in HCCs of G3 mTerc–/– mice compared with TTD+ HCCs (Figure 2, B and C). Specifically, HCCs from TTD+ mice contained higher numbers of cells with short telomeres (53/390) compared with HCCs from TTD– mice (16/432, P < 0.0001). However, HCCs from G3 mTerc–/– mice displayed a further significant increase of cells with short telomeres compared with HCCs from TTD+ mice (52/1202, P = 0.004). These results correlated with a higher number of telomere-associated DNA damage foci (TIFs) in HCCs of G3 mTerc–/– mice (Supplemental Figure 5D).
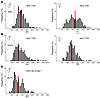 Figure 2
Figure 2Transient telomere dysfunction induces HCCs with shortened telomeres. Telomere length was measured by qFISH. (A–C) Distribution of mean telomere fluorescence intensities (TFI) in HCCs of the indicated genotypes and sex (n = 5–7). Red lines indicate the mean fluorescence intensities. The numbers on the left of the dotted line indicate the percentage of tumor cells with critically short telomeres (TFI < 800).
Array comparative genome hybridization (aCGH) analysis revealed that transient telomere dysfunction significantly increased the frequency of chromosomal aberrations in telomerase-proficient HCCs (3.6 aberrations/TTD+ tumor vs. 1.2 aberrations/TTD– tumor, P = 0.0002; Figure 3, A and B, and Table 1). A comparison of known chromosomal aberrations in human HCCs (Progenetix database, http://www.progenetix.net; ref. 18) with the mouse data showed a significantly higher overlap of TTD+ HCCs versus TTD– HCCs (Supplemental Figure 6, P = 2.2 × 10–16). One of the most prevalent lesions in TTD+ HCCs was the gain in chromosome 15 carrying the c-Myc locus, which is implicated in human liver carcinogenesis (12). Gene set enrichment analysis of differentially regulated genes in TTD+ versus TTD– tumors revealed a significant deregulation of ribosomal genes — a known molecular feature of tumor progression (19), which is regulated by c-MYC (19). Most of the enriched pathways in TTD+ tumors were also associated with HCCs in humans (Supplemental Figure 7 and Supplemental Table 1).
 Figure 3
Figure 3Transient telomere dysfunction induces chromosomal instability in telomerase-proficient liver without strong accumulation of DNA damage. (A) Ideogram displaying chromosomal gains and losses in tumors of the indicated cohorts (n = 6–10). The bars on the right side of each chromosome indicate the frequency (%) of gains (red) and losses (green). (B) Average number of chromosomal alterations in HCCs of the indicated genotypes (n = 6–10). (C) The histogram shows the number of γH2AX-positive cells in HCCs of the indicated genotypes (n = 5–9). (D) Representative immunofluorescence staining of γH2AX foci as an indicator of DNA breaks. Original magnification, ×200. (E) Number of PCNA-positive cells in nontumorous livers and HCCs of mice of the indicated genotypes (n = 5–9). Data represent mean ± SEM.
 Table 1
Table 1The contingency table represents the number of chromosomal aberrations in HCCs of 13-month-old mice of the indicated genotype
Telomerase promotes progression of chromosomal-unstable tumors by limiting DNA damage, cell-cycle arrest, and apoptosis. In contrast to the tumor-promoting effects of transient telomere dysfunction in telomerase-proficient mice, telomere dysfunction impaired tumor progression in telomerase-knockout mice (4, 6). aCGH analysis of HCCs from G3 mTerc–/– revealed similarly high levels of chromosomal aberrations compared with tumors of TTD+ mice (Figure 3, A and B, and Table 1). These data indicated that differences in tumor progression in G3 mTerc–/– compared with TTD+ mice (Figure 1, D and E, and Supplemental Figure 3, A and B) were not associated with the levels of chromosomal aberrations per se. Instead, the data suggested that telomerase activity was required for the survival of chromosomal-unstable tumor cells by limiting levels of telomere dysfunction (Figure 2, A–C, and Supplemental Figure 5D). An analysis of telomerase activity revealed an activation of telomerase in HCCs of TTD+ and TTD– mice (n = 3–7; Supplemental Figure 8, A and B). DNA breaks monitored as γH2AX foci pointed to an increase in DNA damage in HCCs of TTD+ versus TTD– mice, but HCCs of G3 mTerc–/– mice exhibited a strong further increase compared with the other 2 groups (P = 0.0001) (Figure 3, C and D, and Supplemental Figure 8, C and D). This increase was higher than the increase in TIFs (Supplemental Figure 5D), suggesting that it involved an accumulation of intrachromosomal breaks that could be induced by fusion-bridge-breakage cycles in response to telomere dysfunction. Increases in DNA damage accumulation correlated with increased rates of apoptosis and impaired cell proliferation in tumors of G3 mTerc–/– compared with TTD– mice (Figure 3E and Supplemental Figure 8E). In contrast, cell proliferation rates were significantly increased in HCCs of TTD+ compared with TTD– mice (P = 0.0004, n = 7–9; Figure 3E), and TUNEL staining did not reveal a difference in apoptosis of tumors of TTD+ compared with TTD– mice (Supplemental Figure 8E).
This study demonstrates that transient telomere dysfunction induced by a dominant-negative version of TRF2 enhances tumor formation in carcinogen-treated mice. TRF2 is essential for telomere capping, and inhibition of TRF2 induces hallmark features of telomere dysfunction that also occur in response to telomere shortening (14, 20). The molecular pathways of chromosomal fusion formation show differences when telomere dysfunction is induced by telomere shortening or TRF2 inhibition (21). Thus, the data of the current study may also be relevant for human tumors exhibiting an aberrant expression of telomere-binding proteins — a frequent event in human tumors (22, 23).
During the revision of this study, two recent publications showed that telomerase reactivation in late generation of telomerase-deficient mice promotes tumor metastasis and malignancy in the background of genetic lesions inducing prostate cancer (24) or T cell lymphoma (5). In both studies, telomerase was reactivated in tissues exposed to long-term telomere dysfunction (from germ line to adulthood). The current study provides what we believe is the first experimental evidence that transient telomere dysfunction during early or late stages of tumorigenesis promotes cancer formation in telomerase-proficient mice that did not experience telomere dysfunction prior to the oncogenic process. Together, these studies indicate that telomere dysfunction can promote tumorigenesis when occurring in precancerous tissues or at early or late stages of carcinogenesis. In each of these scenarios, telomere dysfunction promotes carcinogenesis only when followed by telomerase reactivation. In contrast, telomere dysfunction impairs tumor progression in telomerase-deficient mice. The average number of chromosomal aberrations in tumor cells was not different in both scenarios, but telomerase-deficient mice accumulated significantly higher numbers of DNA breaks leading to the induction of cell-cycle arrest and apoptosis. These results stand in agreement with the hypothesis that telomerase is required to promote survival of chromosomal-unstable tumors by preventing the accumulation of DNA damage. Telomere dysfunction leads to an accumulation of DNA breaks by the induction of chromosomal fusions and breakage of chromosomes during the cell cycle. In this context, telomerase activation stabilizes telomeres, thereby preventing accumulating DNA breakage and checkpoint induction.
Together, these data provide the first in vivo proof-of-concept that transient telomere dysfunction during early or late stages of tumorigenesis can increase cancer initiation and progression in telomerase-proficient mice and support the stepwise role of crisis and telomerase reactivation during tumor initiation and progression.
-
Methods
Generation of transgenic mice, histological analysis, and MRI. C57BL/6J;Tg(tetopTRF2ΔBΔM)3 mice were obtained by pronuclear injection of a transgenic construct containing a purified 2.7-kb enzyme fragment containing the human TRF2ΔBΔM transgene under the control of a tetracycline-regulated promoter (TRE; tetO) and an SV40 poly A signal sequence into oocytes from C57BL/6J mice. rTALAP-1 (25) mice were backcrossed 7 times to C57Bl/6J mice and then bred to C57BL/6J;Tg(tetopTRF2ΔBΔM) 3 mice expressing a dominant-negative form of TRF2 (TRF2ΔBΔM) under the tetO promoter. DEN was administered to the progeny of this intercross at day 15 after birth (8 mg/kg body weight) by intraperitoneal injection. For transient hepatic expression of TRF2ΔBΔM, mice were treated with intrasplenic injections of doxycycline (50 μg/g body weight) at either 2, 2.5, and 3 months of age or at 12 to 14 months of age. See Supplemental Methods for further details.
Protein isolation and Western blot, qFISH, immunofluorescence stainings, array CGH profiling, cross species analysis, and gene expression analysis. The GEO accession number is GSE36813. See Supplemental Methods for details.
Statistics. Statistical analysis was performed using Microsoft Excel and GraphPad Prism software. The 2-tailed Student’s t test was used to calculate P values, except for those in Table 1, for which the χ2 test was used. Data represent mean ± SEM. P < 0.05 was considered significant.
Study approval. The Institutional Review Board of the University of Ulm approved the study. Animal experiments were approved by the state government of Baden-Württemberg, Tübingen, Germany.
-
Acknowledgments
We thank Hermann Bujard and Kai SchÖnig for providing the rTALAP-1 transgenic mice (DKFZ, Heidelberg). K.L. Rudolph is supported by the Deutsche Forschungsgemeinschaft (KFO 167, Ru745-10/1), Deutsche Krebshilfe e.V. (Tumorstammzellverbund D.2787), and the European Union (GENINCA, contract number 202230). D. Hartmann is funded by the Else KrÖner-Fresenius-Stiftung (EKFS Memorial Scholarship). H.A. Kestler is supported by the BMBF (Federal Ministry of Education and Research) (Forschungskern SyStaR and NGFN+ PaCa-Net).
Address correspondence to: K. Lenhard Rudolph or André Lechel, Institute of Molecular Medicine and Max-Planck-Research Group on Stem Cell Aging, Ulm University, 89081 Ulm, Germany. Phone: 49.731.503.6100; Fax: 49.731.503.6102; E-mail: KLRudolph@fli-leibniz.de (K.L. Rudolph). Phone: 49.731.503.6110; Fax: 49.731.503.6102; E-mail: Andre.Lechel@uni-ulm.de (A. Lechel).
-
Footnotes
Conflict of interest: The authors have declared that no conflict of interest exists.
Reference information: J Clin Invest. 2012;122(6):2283–2288. doi:10.1172/JCI61745.
See the related article at Telomere stability and carcinogenesis: an off-again, on-again relationship.
-
References
- Maser RS, DePinho RA. Connecting chromosomes, crisis, and cancer. Science. 2002;297(5581):565–569.
- Kim NW, et al. Specific association of human telomerase activity with immortal cells and cancer. Science. 1994;266(5193):2011–2015.
- Farazi PA, Glickman J, Jiang S, Yu A, Rudolph KL, DePinho RA. Differential impact of telomere dysfunction on initiation and progression of hepatocellular carcinoma. Cancer Res. 2003;63(16):5021–5027.View this article via: PubMed Google Scholar
- Greenberg RA, Allsopp RC, Chin L, Morin GB, DePinho RA. Expression of mouse telomerase reverse transcriptase during development, differentiation and proliferation. Oncogene. 1998;16(13):1723–1730.
- Hu J, et al. Antitelomerase therapy provokes ALT and mitochondrial adaptive mechanisms in cancer. Cell. 2012;148(4):651–663.
- Lechel A, et al. Telomerase deletion limits progression of p53-mutant hepatocellular carcinoma with short telomeres in chronic liver disease. Gastroenterology. 2007;132(4):1465–1475.
- Rudolph KL, Millard M, Bosenberg MW, DePinho RA. Telomere dysfunction and evolution of intestinal carcinoma in mice and humans. Nat Genet. 2001;28(2):155–159.
- Rudolph KL, et al. Longevity, stress response, and cancer in aging telomerase-deficient mice. Cell. 1999;96(5):701–712.
- Artandi SE, et al. Telomere dysfunction promotes non-reciprocal translocations and epithelial cancers in mice. Nature. 2000;406(6796):641–645.
- Satyanarayana A, Manns MP, Rudolph KL. Telomeres and telomerase: a dual role in hepatocarcinogenesis. Hepatology. 2004;40(2):276–283.
- Miura N, et al. Progressive telomere shortening and telomerase reactivation during hepatocellular carcinogenesis. Cancer Genet Cytogenet. 1997;93(1):56–62.
- Plentz RR, et al. Telomere shortening correlates with increasing aneuploidy of chromosome 8 in human hepatocellular carcinoma. Hepatology. 2005;42(3):522–526.
- van Steensel B, Smogorzewska A, de Lange T. TRF2 protects human telomeres from end-to-end fusions. Cell. 1998;92(3):401–413.
- Karlseder J, Broccoli D, Dai Y, Hardy S, de Lange T. p53- and ATM-dependent apoptosis induced by telomeres lacking TRF2. Science. 1999;283(5406):1321–1325.
- Naugler WE, et al. Gender disparity in liver cancer due to sex differences in MyD88-dependent IL-6 production. Science. 2007;317(5834):121–124.
- Greenberg RA, et al. Short dysfunctional telomeres impair tumorigenesis in the INK4a(delta2/3) cancer-prone mouse. Cell. 1999;97(4):515–525.
- Blasco MA, et al. Telomere shortening and tumor formation by mouse cells lacking telomerase RNA. Cell. 1997;91(1):25–34.
- Baudis M, Cleary ML. Progenetix.net: an online repository for molecular cytogenetic aberration data. Bioinformatics. 2001;17(12):1228–1229.
- Ruggero D, Pandolfi PP. Does the ribosome translate cancer? Nat Rev Cancer. 2003;3(3):179–192.
- Lechel A, et al. The cellular level of telomere dysfunction determines induction of senescence or apoptosis in vivo. EMBO Rep. 2005;6(3):275–281.
- Smogorzewska A, de Lange T. Different telomere damage signaling pathways in human and mouse cells. EMBO J. 2002;21(16):4338–4348.
- Poncet D, et al. Changes in the expression of telomere maintenance genes suggest global telomere dysfunction in B-chronic lymphocytic leukemia. Blood. 2008;111(4):2388–2391.
- Raynaud CM, et al. Telomere shortening is correlated with the DNA damage response and telomeric protein down-regulation in colorectal preneoplastic lesions. Ann Oncol. 2008;19(11):1875–1881.
- Ding Z, et al. Telomerase reactivation following telomere dysfunction yields murine prostate tumors with bone metastases. Cell. 2012;148(5):896–907.
- Schonig K, Schwenk F, Rajewsky K, Bujard H. Stringent doxycycline dependent control of CRE recombinase in vivo. Nucleic Acids Res. 2002;30(23):e134.
-
Version history
- Version 1 (May 24, 2012): No description
- Version 2 (June 1, 2012): No description
- Version 3 (January 7, 2021): Updated Supplemental Figure 5D



Copyright © 2024 American Society for Clinical Investigation
ISSN: 0021-9738 (print), 1558-8238 (online)
