Ribonucleotide
In biochemistry, a ribonucleotide is a nucleotide containing ribose as its pentose component. It is considered a molecular precursor of nucleic acids. Nucleotides are the basic building blocks of DNA and RNA. Ribonucleotides themselves are basic monomeric building blocks for RNA. Deoxyribonucleotides, formed by reducing ribonucleotides with the enzyme ribonucleotide reductase (RNR), are essential building blocks for DNA.[1] There are several differences between DNA deoxyribonucleotides and RNA ribonucleotides. Successive nucleotides are linked together via phosphodiester bonds.
Ribonucleotides are also utilized in other cellular functions. These special monomers are utilized in both cell regulation and cell signaling as seen in adenosine-monophosphate (AMP). Furthermore, ribonucleotides can be converted to adenosine triphosphate (ATP), the energy currency in organisms. Ribonucleotides can be converted to cyclic adenosine monophosphate (cyclic AMP) to regulate hormones in organisms as well.[1] In living organisms, the most common bases for ribonucleotides are adenine (A), guanine (G), cytosine (C), or uracil (U). The nitrogenous bases are classified into two parent compounds, purine and pyrimidine.

Structure
[edit]General structure
[edit]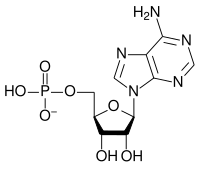



The general structure of a ribonucleotide consists of a phosphate group, a ribose sugar group, and a nucleobase, in which the nucleobase can either be adenine, guanine, cytosine, or uracil. Without the phosphate group, the composition of the nucleobase and sugar is known as a nucleoside. The interchangeable nitrogenous nucleobases are derived from two parent compounds, purine and pyrimidine. Nucleotides are heterocyclic compounds, that is, they contain at least two different chemical elements as members of its rings.[citation needed]
Both RNA and DNA contain two major purine bases, adenine (A) and guanine (G), and two major pyrimidines. In both DNA and RNA, one of the pyrimidines is cytosine (C). However, DNA and RNA differ in the second major pyrimidine. DNA contains thymine (T) while RNA contains uracil (U). There are some rare cases where thymine does occur in RNA and uracil in DNA.[1]
Here are the 4 major ribonucleotides (ribonucleoside 5'-monophosphate) which are the structural units of RNAs.
| Nucleotide | Symbols | Nucleoside |
|---|---|---|
| Adenylate (adenosine 5'-monophosphate) | A, AMP | Adenosine |
| Guanylate (guanosine 5'-monophosphate) | G, GMP | Guanosine |
| Uridylate (uridine 5'-monophosphate) | U, UMP | Uridine |
| Cytidylate (cytidine 5'-monophosphate) | C, CMP | Cytidine |
DNA deoxyribonucleotides versus RNA ribonucleotides
[edit]In ribonucleotides, the sugar component is ribose while in deoxyribonucleotides, the sugar component is deoxyribose. Instead of a hydroxyl group at the second carbon in the ribose ring, it is replaced by a hydrogen atom.[2]
Both types of pentoses in DNA and RNA are in their β-furanose (closed five-membered ring) form and they define the identity of a nucleic acid. DNA is defined by containing 2'-deoxy-ribose nucleic acid while RNA is defined by containing ribose nucleic acid.[1]
In some occasions, DNA and RNA may contain some minor bases. Methylated forms of the major bases are most common in DNA. In viral DNA, some bases may be hydroxymethylated or glucosylated. In RNA, minor or modified bases occur more frequently. Some examples include hypoxanthine, dihydrouracil, methylated forms of uracil, cytosine, and guanine, as well as modified nucleoside pseudouridine.[3] Nucleotides with phosphate groups in positions other than on the 5' carbon have also been observed. Examples include ribonucleoside 2',3'-cyclic monophosphates which are isolatable intermediates, and ribonucleoside 3'-monophosphates which are end products of the hydrolysis of RNA by certain ribonucleases. Other variations include adenosine 3',5'-cyclic monophosphate (cAMP) and guanosine 3',5'-cyclic monophosphate (cGMP).[4]
Linking successive nucleotides
[edit]Ribonucleotides are linked together to form RNA strands via phosphodiester bonds. The 5'-phosphate group of one nucleotide is linked to the 3'-hydroxyl group of the next nucleotide, creating a backbone of alternating phosphate and pentose residues. There is no phosphodiester bond at each end of the polynucleotide.[5] Phosphodiester bonds are formed between ribonucleotides by the enzyme RNA polymerase. The RNA chain is synthesized from the 5' end to the 3' end as the 3'-hydroxyl group of the last ribonucleotide in the chain acts as a nucleophile and launches a hydrophilic attack on the 5'-triphosphate of the incoming ribonucleotide, releasing pyrophosphate as a by-[6] product. Due to the physical properties of the nucleotides, the backbone of RNA is very hydrophilic and polar. At neutral pH, nucleic acids are highly charged as each phosphate group carries a negative charge.[7]
Both DNA and RNA are built from nucleoside phosphates, also known as mononucleotide monomers, which are thermodynamically less likely to combine than amino acids. Phosphodiester bonds, when hydrolyzed, release a considerable amount of free energy. Therefore, nucleic acids tend to spontaneously hydrolyze into mononucleotides. The precursors for RNA are GTP, CTP, UTP and ATP, which is a major source of energy in group-transfer reactions.[8]
Function
[edit]Precursors of deoxyribonucleotides
[edit]Scientists believe that RNA was developed before DNA.[9]
The reduction of ribonucleotides to deoxyribonucleotides is catalyzed by ribonucleotide reductase. Ribonucleotide reductase (RNR) is an essential enzyme for all living organisms since it is responsible for the last step in the synthesis of the four deoxyribonucleotides (dNTPs) necessary for DNA replication and repair.[10] The reaction also requires two other proteins: thioredoxin and thioredoxin reductase. Ribonucleoside diphosphate (NDP) is reduced by thioredoxin to a deoxyribonucleoside diphosphate (dNTP).
The general reaction is:
Ribonucleoside diphosphate + NADPH + H+ → Deoxyribonucleoside diphosphate + NADP+ + H2O
[11]
To illustrate this equation, dATP and dGTP are synthesized from ADP and GDP, respectively. They are first reduced by RNR and then phosphorylated by nucleoside diphosphate kinases to dATP and dGTP. Ribonucleotide reductase is controlled by allosteric interactions. Once dATP binds to ribonucleotide reductase, the overall catalytic activity of the enzyme decreases, as it signifies an abundance of deoxyribonucleotides. This feedback inhibition is reversed once ATP binds.[12]
Ribonucleotide discrimination
[edit]During DNA synthesis, DNA polymerases must select against ribonucleotides, present at much higher levels compared with deoxyribonucleotides. It is crucial that there is selectivity as DNA replication has to be accurate to maintain the organism's genome. It has been shown that the active sites of Y-family DNA polymerases are responsible for maintaining a high selectivity against ribonucleotides.[13] Most DNA polymerases are also equipped to exclude ribonucleotides from their active site through a bulky side chain residue that can sterically block the 2'-hydroxyl group of the ribose ring. However, many nuclear replicative and repair DNA polymerases incorporate ribonucleotides into DNA,[14][15] suggesting that the exclusion mechanism is not perfect.[16]
Synthesis
[edit]Ribonucleotide synthesis
[edit]Ribonucleotides can be synthesized in organisms from smaller molecules through the de novo pathway or recycled through the salvage pathway. In the case of the de novo pathway, both purines and pyrimidines are synthesized from components derived from precursors of amino acids, ribose-5-phosphates, CO2, and NH3.[17][12]

 |
The biosynthetic origins of purine ring atoms N1 arises from the amine group of Asp C2 and C8 originate from formate N3 and N9 are contributed by the amide group of Gln C4, C5 and N7 are derived from Gly C6 comes from HCO−3(CO2) |
De novo biosynthesis of purine nucleotides is fairly complex, consisting of several enzymatic reactions. Utilizing the five-ring sugar structure as a base, the purine ring is built a few atoms at a time in an eleven-step process that leads to the formation of inosinate (IMP). Essentially, IMP is converted into the purine nucleotides required for nucleic acid synthesis.[1]
The pathway begins with the conversion of Ribose-5-Phosphate(R5P) to phosphoribosyl pyrophosphate (PRPP) by enzyme ribose-phosphate diphosphokinase (PRPS1). PRPP is then converted to 5-phosphoribosylamine (5-PRA) as glutamine donates an amino group to the C-1 of PRPP. In a condensation reaction, enzyme GAR synthetase, along with glycine and ATP, activates the glycine carboxylase group of 5-PRA to form Glycinamide ribonucleotide (GAR). Co-enzyme N10-formyl-THF, along with enzyme GAR transformylase, then donates a one-carbon unit to the amino group onto the glycine of GAR, followed by glutamine addition by enzyme FGAR amidotransferase, leading to the formation of formylglycinamidine ribonucleotide (FGAM). Dehydration of FGAM by enzyme FGAM cyclase results in the closure of the imidazole ring, as 5-aminoimidazole ribonucleotide (AIR). A carboxyl group is attached to AIR by N5-CAIR synthetase to form N5-Carboxyaminoimidazole ribonucleotide (N5-CAIR), which is then converted to Carboxyamino-imidazole ribonucleotide (CAIR) with enzyme N5-CAIR mutase. Enzyme SAICAR synthetase, along with amino group from aspartate forms an amide bond to create N-succinyl-5-aminoimidazole-4-carboxamide ribonucleotide (SAICAR). Continuing down the pathway, the removal of the carbon skeleton of aspartate by SAICAR lyase results in 5-aminoimidazole-4-carboxamide ribonucleotide (AICAR). Enzyme AICAR transformylase assists in the final carbon transfer from N10-formyltetrahydrofolate, forming N-formylaminoimidazole-4-carboxamide ribonucleotide (FAICAR). Lastly, closure of the second ring structure is carried out by IMP synthase to form IMP, where IMP fate would lead to the formation of a purine nucleotide.[1]

Synthesis of pyrimidine nucleotides is a much simpler process. The formation of the pyrimidine ring begins with the conversion of Aspartate to N-Carbamoylaspartate by undergoing a condensation reaction with carbamoyl phosphate. Dihydroorotase and dihydroorotase dehydrogenase then converts N-Carbamoylaspartate to orotate. Orotate is covalently linked with phosphoribosyl pyrophosphate (PRPP) by orotate phosphoribysol-transferase yielding orotidine monophosphate (OMP). OMP follows with the decarboxylation by orotidylate decarboxylase to form the Uridylate (UMP) ribonucleotide structure. UMP can then be converted to Uridine-5'-trisphosphate (UTP) by two kinases reaction. Formation of Cytidine-5'-trisphosphate (CTP) from UTP can be achieved by cytidylate synthetase by an acyl phosphate intermediate.[1]
Prebiotic synthesis of ribonucleotides
[edit]In order to understand how life arose, knowledge is required of the chemical pathways that permit formation of the key building blocks of life under plausible prebiotic conditions. According to the RNA world hypothesis free-floating ribonucleotides were present in the primitive soup. These were the fundamental molecules that combined in series to form RNA. Molecules as complex as RNA must have arisen from small molecules whose reactivity was governed by physico-chemical processes. RNA is composed of purine and pyrimidine nucleotides, both of which are necessary for reliable information transfer, and thus Darwinian natural selection and evolution. The synthesis of activated pyrimidine ribonucleotides was demonstrated under plausible prebiotic conditions.[18] The starting materials for the synthesis (cyanamide, cyanoacetylene, glycolaldehyde, glyceraldehyde and inorganic phosphate) were considered to be plausible prebiotic feedstock molecules.[18] Nam et al. demonstrated the direct condensation of nucleobases with ribose to give ribonucleosides in aqueous microdroplets, a key step leading to RNA formation.[19] Also, a plausible prebiotic process for synthesizing pyrimidine and purine ribonucleotides using wet-dry cycles was presented by Becker et al.[20]
History
[edit]
Prior to James Watson and Francis Crick's landmark paper that detailed the structure of DNA from Rosalind Franklin's X-ray crystallography image, there were several historical scientists that also contributed to its discovery.[21] Friedrich Miescher, a Swiss physician, who, in 1869, was first to isolate and identify nucleic substance from the nuclei of white blood cells he later called "nuclein", paving the way for the discovery of DNA.[22] Following Mieschers work, was the German biochemist, Albrecht Kossel, who, in 1878, isolated the non-protein components of "nuclein", and discovered the five nucleobases present in nucleic acids: adenine, cytosine, guanine, thymine and uracil.[23] Although some fundamental facts were known about nucleic acids due to these early discoveries, its structure and function remained a mystery.
It wasn't until the discovery of nucleotides in 1919 by Phoebus Levene, a Russian-Lithuanian biochemist that re-opened the gates of the DNA discovery. Levene first identified the carbohydrate component present in yeast RNA was in fact ribose. However, it was not until his discovery that the carbohydrate component in thymus nucleic acid was also a sugar but lacked one oxygen atom, termed deoxyribose, that his discovery was widely appreciated by the scientific community. Eventually, Levene was able to identify the correct order of which the components of RNA and DNA are put together, a phosphate-sugar-base unit, in which he later called a nucleotide. Although the order of nucleotide components were well understood by Levene, the structure of nucleotide arrangement in space and its genetic code still remained a mystery during the early years of his career.[24]
See also
[edit]- Ribonucleosides or ribosides
References
[edit]- ^ a b c d e f g Nelson, David L.; Lehninger, Albert L.; Cox, Michael M. (2008). Lehninger Principles of Biochemistry (5th ed.). W H Freeman. pp. 272–3. ISBN 978-0-7167-7108-1.
- ^ Newsholme, Eric A.; Leech, Anthony R.; Board, Mary (2008). Functional biochemistry in health & disease: metabolic regulation in health and disease (2nd ed.). Hoboken, N.J.: Wiley. ISBN 978-0-471-98820-5.
- ^ Das, Debajyoti (2010). Biochemistry. Bimal Kumar Dhur of Academic Publishers.
- ^ Cox, Michael M.; Nelson, David L. (2008). Principles of Biochemistry. W H Freeman. ISBN 978-1-4292-2263-1. OCLC 216940502.
- ^ Raymond, Kenneth W. (2010). General, organic, and biological chemistry: an integrated approach (3rd ed.). Wiley. ISBN 978-0-470-55124-0. OCLC 699600111.
- ^ Schaechter, Moselio; Lederberg, Joshua, eds. (2004). The Desk Encyclopedia of Microbiology (1st ed.). Amsterdam: Elsevier. ISBN 0-12-621361-5. OCLC 1055874153.
- ^ Turner, Phil; et al. (2005). Molecular Biology. Instant Notes (3rd ed.). CRC, Taylor & Francis. ISBN 0-415-35167-7. OCLC 61745541.
- ^ Nelson, David (2008). Lehninger Principles of Biochemistry. W H Freeman and Co. pp. 274–275.
- ^ Chauhan, Ashok K.; Varma, Ajit, eds. (2009). A textbook of molecular biotechnology. New Delhi: I.K. International Pub. House. ISBN 978-93-80026-37-4.
- ^ Cendra Mdel, M; Juárez, A; Torrents, E (2012). "Biofilm modifies expression of ribonucleotide reductase genes in Escherichia coli". PLOS ONE. 7 (9): e46350. Bibcode:2012PLoSO...746350C. doi:10.1371/journal.pone.0046350. PMC 3458845. PMID 23050019.
- ^ Campbell, Mary K.; Farrell, Shawn O. (2009). Biochemistry (7th ed.). Belmont, CA: Brooks/Cole Cengage Learning. ISBN 978-0-8400-6858-3.
- ^ a b Berg, J.M.; Tymoczko, J.L.; Stryer, L. (2002). "§25.2 Purine Bases can be Synthesized by de Novo or Recycled by Salvage Pathways". Biochemistry. W.H. Freeman. ISBN 9780716730514. OCLC 48055706.
- ^ Kirouac, Kevin N.; Suo, Zucai; Ling, Hong (1 April 2011). "Structural Mechanism of Ribonucleotide Discrimination by a Y-Family DNA Polymerase". Journal of Molecular Biology. 407 (3): 382–390. doi:10.1016/j.jmb.2011.01.037. PMID 21295588.
- ^ Nick McElhinny, SA; Kumar, D; Clark, AB; Watt, DL; Watts, BE; Lundström, EB; Johansson, E; Chabes, A; Kunkel, TA (October 2010). "Genome instability due to ribonucleotide incorporation into DNA". Nature Chemical Biology. 6 (10): 774–81. doi:10.1038/nchembio.424. PMC 2942972. PMID 20729855.
- ^ Nick McElhinny, SA; Watts, BE; Kumar, D; Watt, DL; Lundström, EB; Burgers, PM; Johansson, E; Chabes, A; Kunkel, TA (16 March 2010). "Abundant ribonucleotide incorporation into DNA by yeast replicative polymerases". Proceedings of the National Academy of Sciences of the United States of America. 107 (11): 4949–54. Bibcode:2010PNAS..107.4949N. doi:10.1073/pnas.0914857107. PMC 2841928. PMID 20194773.
- ^ Kasiviswanathan, R; Copeland, WC (Sep 9, 2011). "Ribonucleotide discrimination and reverse transcription by the human mitochondrial DNA polymerase". The Journal of Biological Chemistry. 286 (36): 31490–500. doi:10.1074/jbc.M111.252460. PMC 3173122. PMID 21778232.
- ^ Nelson, Lehninger & Cox 2008, pp. 881–894
- ^ a b Powner MW, Gerland B, Sutherland JD (May 2009). "Synthesis of activated pyrimidine ribonucleotides in prebiotically plausible conditions". Nature. 459 (7244): 239–42. Bibcode:2009Natur.459..239P. doi:10.1038/nature08013. PMID 19444213.
- ^ Nam I, Nam HG, Zare RN (January 2018). "Abiotic synthesis of purine and pyrimidine ribonucleosides in aqueous microdroplets". Proc Natl Acad Sci U S A. 115 (1): 36–40. Bibcode:2018PNAS..115...36N. doi:10.1073/pnas.1718559115. PMC 5776833. PMID 29255025.
- ^ Becker S, Feldmann J, Wiedemann S, Okamura H, Schneider C, Iwan K, Crisp A, Rossa M, Amatov T, Carell T (October 2019). "Unified prebiotically plausible synthesis of pyrimidine and purine RNA ribonucleotides". Science. 366 (6461): 76–82. Bibcode:2019Sci...366...76B. doi:10.1126/science.aax2747. PMID 31604305.
- ^ Watson, JD; Crick, FH (Apr 25, 1953). "Molecular structure of nucleic acids; a structure for deoxyribose nucleic acid". Nature. 171 (4356): 737–8. Bibcode:1953Natur.171..737W. doi:10.1038/171737a0. PMID 13054692. S2CID 4253007.
- ^ Dahm, R (January 2008). "Discovering DNA: Friedrich Miescher and the early years of nucleic acid research". Human Genetics. 122 (6): 565–81. doi:10.1007/s00439-007-0433-0. PMID 17901982. S2CID 915930.
- ^ Jones, ME (September 1953). "Albrecht Kossel, a biographical sketch". The Yale Journal of Biology and Medicine. 26 (1): 80–97. PMC 2599350. PMID 13103145.
- ^ Levene, Phoebus (1919). "The structure of yeast nucleic acid IV. Ammonia Hydrolysis". Journal of Biological Chemistry. 40 (2): 415–424. doi:10.1016/S0021-9258(18)87254-4.