Comparative analysis and implications of the chloroplast genomes of three thistles (Carduus L., Asteraceae)
- Published
- Accepted
- Received
- Academic Editor
- Jonathan Thomas
- Subject Areas
- Genomics, Plant Science
- Keywords
- Carduus crispus, Chloroplast genome, Invasive species, Molecular markers, Plumeless thistles
- Copyright
- © 2021 Jung et al.
- Licence
- This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, reproduction and adaptation in any medium and for any purpose provided that it is properly attributed. For attribution, the original author(s), title, publication source (PeerJ) and either DOI or URL of the article must be cited.
- Cite this article
- 2021. Comparative analysis and implications of the chloroplast genomes of three thistles (Carduus L., Asteraceae) PeerJ 9:e10687 https://doi.org/10.7717/peerj.10687
Abstract
Background
Carduus, commonly known as plumeless thistles, is a genus in the Asteraceae family that exhibits both medicinal value and invasive tendencies. However, the genomic data of Carduus (i.e., complete chloroplast genomes) have not been sequenced.
Methods
We sequenced and assembled the chloroplast genome (cpDNA) sequences of three Carduus species using the Illumina Miseq sequencing system and Geneious Prime. Phylogenetic relationships between Carduus and related taxa were reconstructed using Maximum Likelihood and Bayesian Inference analyses. In addition, we used a single nucleotide polymorphism (SNP) in the protein coding region of the matK gene to develop molecular markers to distinguish C. crispus from C. acanthoides and C. tenuiflorus.
Results
The cpDNA sequences of C. crispus, C. acanthoides, and C. tenuiflorus ranged from 152,342 bp to 152,617 bp in length. Comparative genomic analysis revealed high conservation in terms of gene content (including 80 protein-coding, 30 tRNA, and four rRNA genes) and gene order within the three focal species and members of subfamily Carduoideae. Despite their high similarity, the three species differed with respect to the number and content of repeats in the chloroplast genome. Additionally, eight hotspot regions, including psbI-trnS_GCU, trnE_UUC-rpoB, trnR_UCU-trnG_UCC, psbC-trnS_UGA, trnT_UGU-trnL_UAA, psbT-psbN, petD-rpoA, and rpl16-rps3, were identified in the study species. Phylogenetic analyses inferred from 78 protein-coding and non-coding regions indicated that Carduus is polyphyletic, suggesting the need for additional studies to reconstruct relationships between thistles and related taxa. Based on a SNP in matK, we successfully developed a molecular marker and protocol for distinguishing C. crispus from the other two focal species. Our study provides preliminary chloroplast genome data for further studies on plastid genome evolution, phylogeny, and development of species-level markers in Carduus.
Introduction
Carduus L. (subfamily Carduoideae; Asteraceae), commonly known as plumeless thistles, comprises 90 species native to Eurasia and Africa (Angiosperm Phylogeny Group, 2016). Several Carduus species are invasive, noxious weeds on other continents (Doing, Biddiscombe & Knedlhans, 1969). Four species, including C. acanthoides L. (spiny plumeless thistle), C. tenuiflorus Curtis (sheep thistle), C. pycnocephalus L. (Italian thistle), and C. crispus Guirão ex Nyman (welted thistle), all of which originate in Eurasia and Africa, are considered invasive in North America (Dunn, 1976; Verloove, 2014). Carduus crispus, also called curly plumeless thistle, is also considered an invasive species in Korea (Jung et al., 2017). This species differs from other Carduus in having soft, sparsely arachnoid-hairy leaves with short marginal bristles, and apically recurved involucral bracts (Todorov et al., 2018). Among Carduus species, C. crispus contains chemicals with the potential to treat various diseases (Xie, Li & Jia, 2005; Davaakhuu, Sukhdolgor & Gereltu, 2010; Lee et al., 2011; Tunsag, Davaakhuu & Batsuren, 2011). Specifically, certain compounds extracted from C. crispus have potential value in the treatment of obesity and cancer (Davaakhuu, Sukhdolgor & Gereltu, 2010; Lee et al., 2011). While Carduus has been studied from various perspectives (i.e., invasion, phylogeny, and medicinal effects), its chloroplast genome has not been sequenced. It is therefore worthwhile to study the genome of Carduus, and particularly that of C. crispus, which has potential medicinal benefits.
In most angiosperms, the chloroplast genome (cpDNA) contains genes essential to photosynthesis (Sugiura, 1992). Genomic events (i.e., gene deletion, inversion, or duplication) in cpDNA may provide information about species’ evolutionary history (Cosner, Raubeson & Jansen, 2004; Do & Kim, 2017; Haberle et al., 2008). For example, the Fabaceae includes clades that are characterised by large inversions and the loss of inverted repeat regions (Choi & Choi, 2017). Inversions have also been recorded in the cpDNA of Asteraceae (Kim, Choi & Jansen, 2005). Specifically, a large inversion comprising a 22.8 kb sequence occurred simultaneously with a small inversion of a 3.3 kb fragment; this event coincided with the split between major clades (excluding Barnadesioideae) in the evolution of Asteraceae. cpDNA data can also be used to develop molecular markers based on nucleotide polymorphisms (i.e., single nucleotide polymorphism (SNP) markers and microsatellite markers). Molecular authentication has been reported for various plant species, with a focus on invasive plants, endangered species, and taxa with potential medicinal value (Kim et al., 2012; Ishikawa, Sakaguchi & Ito, 2016; Luo et al., 2016; Park et al., 2016; Marochio et al., 2017; Han et al., 2018; Do et al., 2019). Specific regions of cpDNA have been identified for developing molecular markers in plants, including the commonly-used matK region (Poovitha et al., 2016; Vu et al., 2017). Among the Asteraceae, studies on molecular markers have been conducted for rubber dandelion (Taraxacum kok-saghyz LE Rodin), horseweed (Conyza sp.), Indian Chrysanthemum (Chrysanthemum indicum L.), the endemic herb Aster savatieri Makino, and the invasive plant Tithonia diversifolia (Hemsl.) A Gray (Ishikawa, Sakaguchi & Ito, 2016; Luo et al., 2016; Zhang et al., 2017; Marochio et al., 2017; Han et al., 2018). In addition to the development of these molecular markers, complete cpDNA sequences have been reported for various Asteraceae species (Kim, Choi & Jansen, 2005; Choi & Park, 2015; Wang et al., 2015a; Wang et al., 2015b; Yun, Gil & Kim, 2017; Liu et al., 2018; Ma, Sun & Zhao, 2018; Su et al., 2018). cpDNA sequences may be used to elucidate the phylogeny of angiosperms from the clade that is basal to monocots and eudicots (Angiosperm Phylogeny Group, 2016). Previous investigations into phylogenetic relationships among members of the Asteraceae have been conducted using a range of molecular data types, including rbcL, ndhF, matK, chloroplast DNA restriction sites, ITS sequence data, and nuclear loci (Jansen, Michaels & Palmer, 1991; Häffner & Hellwig, 1999; Fu et al., 2016; Mandel et al., 2019). However, the paucity of available sequence data may have resulted in ambiguous relationships between Carduus and related taxa (Häffner & Hellwig, 1999; Fu et al., 2016). In particular, ITS sequence data suggest that C. leptacanthus is sister to Cirsium and Notobasis, whereas another Carduus species is sister to Cirsium and Tyrimmus (Häffner & Hellwig, 1999). As such, clarification of relationships between Carduus and related species will require studies that include a larger number of Carduus species and different data types (i.e., chloroplast and mitochondrial genomes).
We used next-generation sequencing (NGS) to sequence and characterise the chloroplast genomes of Carduus crispus, C. acanthoides, and C. tenuiflorus, which exhibit both invasive tendencies and potential medical utility (particularly C. crispus). We then conducted comparative genomic analyses to explore genomic diversity among the three species with respect to highly variable regions, and the types and numbers of repeats. In addition, we reconstructed the formerly ambiguous relationship between Carduus and related taxa based on 78 protein-coding regions and non-coding sequences. Finally, we developed a specific molecular marker for C. crispus based on a SNP in the matK gene. This molecular marker provides useful information for managing C. crispus invasions, particularly with respect to the identification of immature (vegetative) individuals, which tend to be morphologically similar to other Carduus species (i.e., having winged stems with apical spines and spiny leaves). This molecular marker may also support positive identification of C. crispus for medical usage.
Materials & Methods
Taxon sampling, total DNA extraction, chloroplast genome assembly, and comparative analysis
Notes: Asterisks indicate samples for next generation sequencing (NGS) analysis.
No.
Species
Voucher
Location
1
Carduus crispus Guirão ex Nyman
Korea National Arboretum (LK0908)
Mt. Seokbyeong, Imgye-myeon, Jeongseon-gun, Gangwon-do, Republic of Korea
2
Carduus crispus Guirão ex Nyman*
Korea National Arboretum (LK0943)
Mt. cheongog, Hajang-myeon, Samcheok-si, Gangwon-do, Republic of Korea
3
Carduus crispus Guirão ex Nyman
Korea National Arboretum (LK1497)
Mt. Nochu, Yeoryang-myeon, Jeongseon-gun, Gangwon-do, Republic of Korea
4
Carduus crispus Guirão ex Nyman
Korea National Arboretum (CNUFR0470)
295, Sinseong-ri, Bukha-myeon, Jangseong-gun, Jeollanam-do, Republic of Korea
5
Carduus crispus Guirão ex Nyman
Korea National Arboretum (LK0430)
Mt. cheongog, Imgye-myeon, Jeongseon-gun, Gangwon-do, Republic of Korea
6
Carduus crispus Guirão ex Nyman
National Institute of Biological Resources (NIBRVP0000619716)
Mt. Jaam, Namhu-myeon, Andong-si, Gyeongsangbuk-do, Republic of Korea
7
Carduus crispus Guirão ex Nyman
National Institute of Biological Resources (NIBRVP0000601328)
Sanghwa-ri, Danchon-myeon, Uiseong-gun, Gyeongsangbuk-do, Republic of Korea
8
Carduus crispus Guirão ex Nyman
National Institute of Biological Resources (NIBRVP0000524207)
Mt. Beophwa, Yugu-eup, Gongju-si, Chungcheongnam-do, Republic of Korea
9
Carduus crispus Guirão ex Nyman
National Institute of Biological Resources (NIBRVP0000613325)
Mt. Mani, Hwado-myeon, Ganghwa-gun, Incheon, Republic of Korea
10
Carduus crispus Guirão ex Nyman
National Institute of Biological Resources (NIBRVP0000580633)
Mt. Jeonggwang, Mohyeon-eup, Cheoin-gu, Yongin-si, Gyeonggi-do, Republic of Korea
11
Carduus crispus Guirão ex Nyman
National Institute of Biological Resources (NIBRVP0000578538)
Mt. Jangam, Pyeongchang-eup, Pyeongchang-gun, Gangwon-do, Republic of Korea
12
Carduus crispus Guirão ex Nyman
National Institute of Biological Resources (NIBRVP0000580501)
Mt. Gwangdeok, Dongnam-gu, Cheonan-si, Chungcheongnam-do, Republic of Korea
13
Carduus acanthoides L.*
Kim 2018-001, Nevada city, Ca, USA
USA, California, Nevada city
14
Carduus acanthoides L.
NewYork Botanical Garden (00532263)
USA, Colorado, Pitkin Co., South side of State Highway 82, 5 miles W of Aspen, Airport., 2256 - 2256m
15
Carduus acanthoides L.
NewYork Botanical Garden (00532244)
USA, Wisconsin, Richland Co., 3 miles SE of Richland Center., 43.300443 -90.321019
16
Carduus acanthoides L.
NewYork Botanical Garden (00532262)
USA, Wyoming, Platte Co., 1402 - 1402m
17
Carduus acanthoides L.
Carnegie Museum Herbarium (267884)
Romania, Oltenia, distr. Dolj, inter vicos Lascar Catargiu et Popoveni ad Canalul colector, 75m
18
Carduus acanthoides L.
Carnegie Museum Herbarium (528144)
USA, Pennsylvania, Mifflin, Maple & Walnut Sts, Belleville
19
Carduus acanthoides L.
United States National Herbarium (1944312)
Czech Republic, Bohemia centralis: Paraha-Troja. In ruderatis
20
Carduus acanthoides L.
United States National Herbarium (3419733)
Ukraine, Prov. Czerkassy, prope opp. Umanj, in ruderatis
21
Carduus tenuiflorus Curtis*
Kim 2018-002, Nevada city, Ca, USA
USA, California, Nevada city
22
Carduus tenuiflorus Curtis
NewYork Botanical Garden (00366662)
Mexico, El Cercado. Santiago, N. L., 495–495 m
23
Carduus tenuiflorus Curtis
Carnegie Museum Herbarium (519240)
USA, California, Humboldt, Angels Ranch, toward Hungry Hollow, Bald Mountain
24
Carduus tenuiflorus Curtis
Carnegie Museum Herbarium (282243)
USA, California, Alameda, Berkeley
Phylogenetic analysis of Carduus and related taxa
A total of 78 protein-coding regions were extracted from the complete cpDNA of the focal species and other related taxa (Table S1). Sequences were aligned using MUSCLE embedded in Geneious Prime (Edgar, 2004). We used jModeltest (Posada, 2008) to find the best model for the aligned DNA sequences; GTR + I + R was selected as the most suitable model and was used in Maximum Likelihood (ML) and Bayesian Inference (BI) analyses. The ML analysis was conducted with the IQ-tree web server (http://iqtree.cibiv.univie.ac.at), using 1,000 bootstrap replications to calculate branch support values (Trifinopoulos et al., 2016). We used MrBayes v3.2 (Ronquist et al., 2012) for BI analyses. The Markov chain Monte Carlo (MCMC) analysis was run for 1,000,000 generations, and a tree was assembled every 1000 generations. A 25% burn-in setting was used for summarising trees. Figtree v4.0 (http://tree.bio.ed.ac.uk/software/figtree/) was used to visualise phylogenetic trees. Other datasets, including whole chloroplast genomes (excluding one IR region), non-coding regions of cpDNA, and hotspot regions derived from the cpDNA of Carduus species, were used in phylogenetic analysis in addition to protein-coding regions (Table S1). Analytical procedures for these additional datasets were identical to those used for the protein coding regions; however, we used the TVM+I+G model for the whole chloroplast genome (excluding one inverted repeat [IR] region) and all non-coding regions, and the TVM+G model for the hotspot regions dataset.
SNP identification, primer design, and multiplex PCR
The complete matK gene, extracted from the cpDNA of the three focal species, was aligned using MUSCLE to identify SNPs (Edgar, 2004; Fig. S1). The selected SNP for C. crispus was then confirmed by aligning the available matK sequences of other Carduus species on NCBI to those of the focal species (Fig. S2). Based on SNP data, primer pairs were designed using Primer3 to distinguish C. crispus from other Carduus species (Untergasser et al., 2012). Primer sequences included matK_463F (5′-CATCTGGAAATCTTGGTTCAG-3′), matK_1162R (5′-GATGCCCCAATGCGTTACAA-3′), CD_SNP_F1 (5′-AATTCTTGCTTCAAAAGG GTCC- 3′), CD_SNP_R1 (5′-TTCCATTTATTCATCAA AAGATAC-3′), CD_SNP_F2 (5′-AATTCTTGCTTCAAAAGGGTCG-3′), and CD_SNP_R2 (5′-TTCCATTTATTCATCAAAAGATAG- 3′). The multiplex PCR of matK_463F, matK_1162R, CD_SNP_F1, and CD_SNP_R1 was designed to yield the 323 bp band for C. crispus, the 421 bp band for other Carduus, and the 700 bp band for all examined samples (Fig. S3). By contrast, the combination of matK_463F, matK_1162R, CD_SNP_F2, and CD_SNP_R2 yielded a 421 bp PCR product for C. crispus and a 323 bp band for other Carduus species (Fig. S3). Reactions were conducted in 25 µl solution consisting of 50 ng of template DNA, 2.5 µl of 10× reaction buffer, 0.5 U of E-taq DNA polymerase, 50 mM MgCl2, and 5 mM dNTPs. Concentrations of outer primer pairs (matK_463F and matK_1162R) and inner primer pairs (CD_SNP_F1 and CD_SNP_R1, CD_SNP_F2 and CD_SNP_R2) were 0.75 pM and 0.5 pM, respectively. The PCR procedure consisted of 1 min at 94 °C, followed by denaturing for 1 min at 94 °C, annealing for 40 s at 55 °C, an extension stage of 50 s at 72 °C, and an additional extension of 7 min at 72 °C.
| Species | Carduus crispus (MK652229) | C. tenuiflorus (MK652230) | C. acanthoides (MK652228) | Cynara humilis (KP299292) | C. baetica (KP842706) | C. cornigera (KP842707) | C. cardunculus var. scolymus (KP842708) | C. cardunculus var sylvestris (KP842721) | Cirsium arvense (KY562583) | Helianthus annus (NC007977) |
|---|---|---|---|---|---|---|---|---|---|---|
| Total reads | 21,118,624 | 4,621,758 | 4,300,959 | – | – | – | – | – | – | – |
| Assemble read | 805,076 (3.8%) | 189,215 (4.1%) | 180,245 (4.2%) | – | – | – | – | – | – | – |
| Coverage | 1,585 | 372 | 354 | – | – | – | – | – | – | – |
| Number of contigs | 126 | 119 | 14 | – | – | – | – | – | – | – |
| N50 value (bp) | 95,199 | 50,873 | 128,044 | – | – | – | – | – | – | – |
| Total length | 152,342 | 152,426 | 152,617 | 152,585 | 152,548 | 152,550 | 152,529 | 152,528 | 152,855 | 151,104 |
| LSC | 83,254 | 83,360 | 83,532 | 83,622 | 83,599 | 83,580 | 83,578 | 83,577 | 83,859 | 83,530 |
| SSC | 18,706 | 18,674 | 18,693 | 18,651 | 18,639 | 18,660 | 18,641 | 18,641 | 18,633 | 18,308 |
| IR | 25,191 | 25,196 | 25,196 | 25,156 | 25,155 | 25,155 | 25,155 | 25,155 | 25,182 | 24,633 |
| Protein-coding genes | 80 | 80 | 80 | 80 | 80 | 80 | 80 | 80 | 80 | 80 |
| tRNA | 30 | 30 | 30 | 30 | 30 | 30 | 30 | 30 | 30 | 30 |
| rRNA | 4 | 4 | 4 | 4 | 4 | 4 | 4 | 4 | 4 | 4 |
| LSC-IR junction | rps19 (60 bp) | rps19 (60 bp) | rps19 (61 bp) | rps19 (60 bp) | rps19 (60 bp) | rps19 (60 bp) | rps19 (60 bp) | rps19 (60 bp) | rps19 (60 bp) | rps19 (101 bp) |
| SSC-IR junction | ycf1 (565 bp) | ycf1 (565 bp) | ycf1 (568 bp) | ycf1 (567 bp) (ycf1-ndhF overlap 17 bp) | ycf1 (567 bp) (ycf1-ndhF overlap 17 bp) | ycf1 (567 bp) (ycf1-ndhF overlap 17 bp) | ycf1 (567 bp) (ycf1-ndhF overlap 17 bp) | ycf1 (567 bp) (ycf1-ndhF overlap 17 bp) | ycf1 (565 bp) | ycf1 (576 bp) |
| Pairwise identity (%) | ||||||||||
| C. crispus | 100 | 98.5 | 98.5 | 98.5 | 98.5 | 98.5 | 98.8 | 92.9 | ||
| C. tenuiflorus | 99.6 | 100 | 98.5 | 98.5 | 98.5 | 98.5 | 98.5 | 98.9 | 92.3 | |
| C. acanthoides | 99.2 | 99.3 | 100 | 98.7 | 98.6 | 98.7 | 98.7 | 98.7 | 98.7 | 92.3 |
Notes:
The dashes (–) mean no data in this study.
Results
Comparative chloroplast genome analysis of the focal species
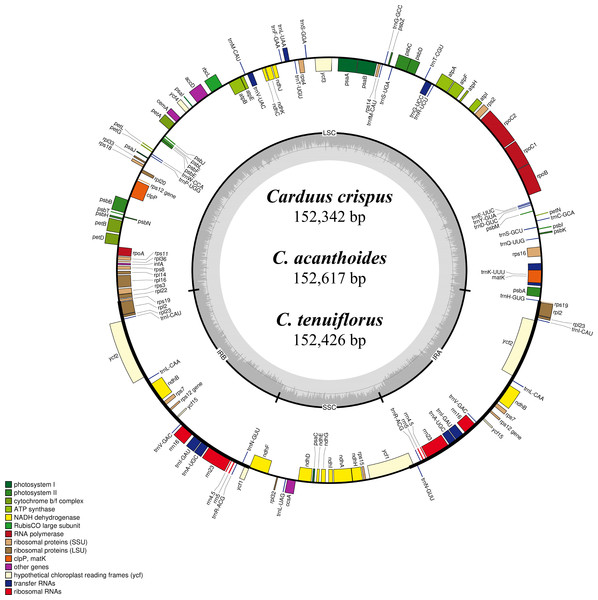
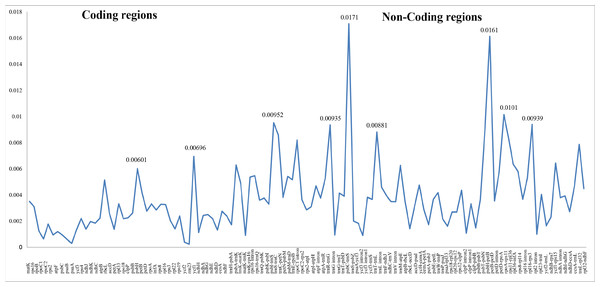
Differing numbers of reads were obtained from NGS data, resulting in varying cpDNA coverage rates among the three focal species (Table 2). Total cpDNA length differed among species, ranging from 152,342 bp to 152,617 bp, and included a large single copy (LSC), a small single copy (SSC), and two IR regions (Fig. 1). By contrast, all three species had identical numbers of protein coding (80), tRNA (30), and rRNA (4) genes (Table 2, Table S3). The IR-LSC and IR-SSC junctions were located in the rps19 and ycf1 coding regions, respectively, but were longer in C. acanthoides (rps19 = 61 bp and ycf1 = 568 bp) than in the other two species (rps19 = 60 bp and ycf1 = 565 bp). In addition, pairwise identity indicated that C. crispus is more similar to C. tenuiflorus (99.6%) than C. acanthoides (99.2%). Observations of nucleotide diversity indicated that 119 of 131 surveyed regions differ among the three focal species (Table S2, Fig. 2). Compared to coding regions, non-coding sequences had higher Pi values (Fig. 2). The highest Pi values were found in the psbC-trnS (0.0171) and psbH-petB (0.0161) regions. The highest value in coding regions was 0.00696 for ycf1 (Fig. 2). High nucleotide diversity regions (Pi values >0.008) included psbI-trnS_GCU, trnE_UUC-rpoB, trnR_UCU-trnG_UCC, psbC-trnS_UGA, trnT_UGU-trnL_UAA, psbT-psbN, petD-rpoA, and rpl16-rps3.
Figure 1: Map of the chloroplast genomes of Carduus.
Genes inside the circle are transcribed clockwise whereas genes outside the circle are transcribed counterclockwise. LSC, large single copy; SSC, small single copy; IRA–IRB, inverted repeat regions.Figure 2: Nucleotide diversity (Pi values) among the three Carduus species.
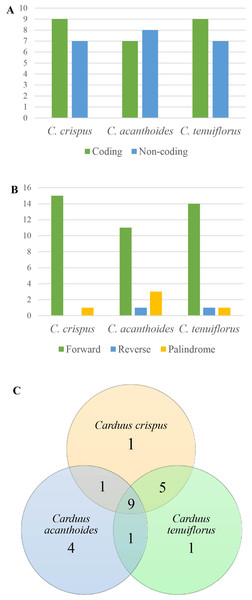
Figure 3: Repeats in the three Carduus species.
(A) Distribution of repeats in the coding and non-coding regions, (B) composition of three types of repeat, and (C) unique and shared repeats in the three Carduus species.Features of cpDNA repeats
Analysis of SSRs yielded 43 SSRs in C. crispus, 40 in C. tenuiflorus, and 31 in C. acanthoides (Table S4). SSRs occupied the same position in all three species and were mostly located in non-coding regions. Although four types of SSR (i.e., mono-, di-, tri-, and tetra-nucleotides) were identified, most SSRs were mononucleotides composed of A and T nucleotides (Table S4). All 31 SSRs found in C. acanthoides were also present in the other two species (Table S4). By contrast, C. crispus had three unique SSRs and shared nine SSRs with C. tenuiflorus. There were no unique SSRs in C. acanthoides or specific shared SSRs between C. acanthoides and either C. crispus or C. tenuiflorus.
Among the focal species, 16 repeats were identified for both C. crispus and C. tenuiflorus, compared to 15 for C. acanthoides (Fig. 3A, Table S5). There were more repeats in coding regions than in non-coding areas, with the exception of C. acanthoides. Three types of repeats (i.e., forward, reverse, and palindrome) were identified in C. tenuiflorus and C. acanthoides; by contrast, only forward and palindrome repeats were found in C. crispus. Forward repeats were more abundant than reverse and palindrome repeats (Fig. 3B). Among recorded repeats, nine were common among all three species (Fig. 3C). Carduus acanthoides had four unique repeats, whereas C. tenuiflorus and C. crispus each had a single unique repeat. Carduus acanthoides shared one specific repeat with both C. crispus and C. tenuiflorus, whereas C. crispus and C. tenuiflorus shared five specific repeat regions (Fig. 3C).
Phylogenetic relationships between Carduus and related taxa
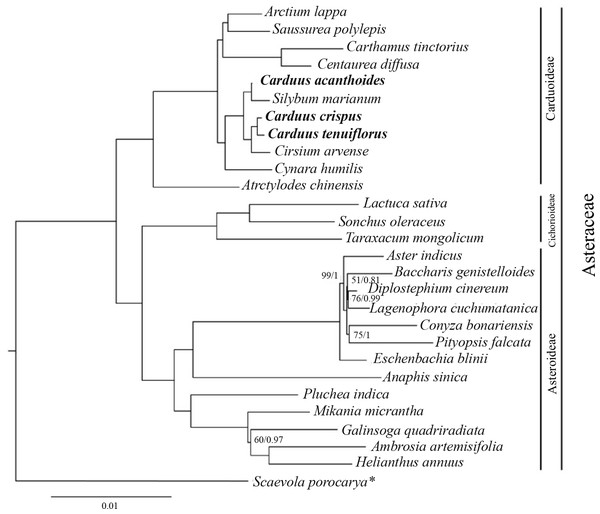
The ML and BI analyses, based on 78 protein-coding genes from Carduus and related taxa, yielded identical topologies (Fig. 4). In particular, both analyses confirmed the monophyly of Asteraceae subfamilies (i.e., Carduoideae, Chichorioideae, and Asteroideae). In contrast to the high support for Carduoideae and Cichorioideae (Bootstrap = 100/Posterior Probability = 1), low support was found for Asteroideae clades (Fig. 4). Notably, monophyly of the three Carduus species was not supported by either analysis. For example, C. acanthoides was sister to Silybum marianum, whereas C. crispus and C. tenuiflorus were sister to Cirsium arvense. Additional ML and BI analyses of full cpDNA sequences and non-coding regions suggested similar relationships (Fig. S4).
Figure 4: BI tree of Carduus and related taxa inferred from 78 protein coding cpDNA regions.
Numbers indicate supporting values (BP, Bootstrap; PP, Posterior Probability). The asterisk indicates the Scaevola porocarya branch, which is compared with other species in the tree; this branch was reduced five times. Only supporting values under 100 (BP) or 1 (PP) are shown.Multiplex PCR and specific markers for C. crispus
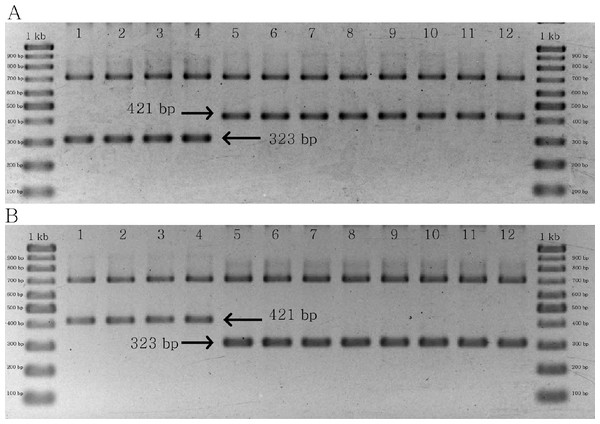
The results of multiplex PCR for the two groups of primer pairs yielded similar products, both of which were designed to identify C. crispus. In the first group, a 323 bp band was found in C. crispus, whereas a 421 bp band was identified in C. acanthoides and C. tenuiflorus (Fig. 5A). By contrast, the combination of matK_463F, matK_1162R, CD_SNP_F2, and CD_SNP_R2 yielded a longer PCR product for C. crispus in comparison with the other species (Fig. 5B). The designed primer pairs were specific to all C. crispus samples examined in this study (Figs. S5 and S6).
Figure 5: PCR results of specific primer pairs for C. crispus.
(A) Combination of matK_463F, matK_1162R, CD_SNP_F1, and CD_SNP_R1, and (B) combination of matK_463F, matK_1162R, CD_SNP_F2, and CD_SNP_R2. Numbers 1–4 = C. crispus; 5–8 = C. acanthoides, and 9–12 = C. tenuiflorus.Discussion
Conservatism of Carduus cpDNA
Chloroplast genomes are highly conserved in angiosperms with respect to gene content and order (Sugiura, 1992). This conservative tendency was observed in the three newly sequenced Carduus cpDNA genomes, compared to other Asteraceae members (Table 2). Other cpDNA sequences have revealed unique genomic events in Asteraceae. For example, the atpB gene, which encodes the CF1 ATPase beta subunit, is annotated as a pseudogene in Aster spathulifolius due to a deletion within the coding region (Choi and Park, 2015). Similarly, trnT_GGU was completely deleted or pseudogenised in the tribe Gnaphalieae (Lee et al., 2017). Duplication of trnF_GAA has been identified in Taraxacum (Salih et al., 2017). No comparable genomic events are present in Carduus or other members of subfamily Carduoideae (Table 2, Fig. S7). However, nucleotide diversity data pointed to potential regions for further study of phylogeny and population genetics, and the development of Carduus-specific molecular markers (Table S2, Fig. 2). The number of species we examined for this study was low relative to the approximately 32,000 known Asteraceae species. Therefore, additional studies that include the majority of Asteraceae species should be conducted to explore the overall evolutionary trends in the chloroplast genomes of this globally-distributed family.
Chloroplast genomes provide useful molecular data for reconstructing phylogeny, exploring biogeography, and estimating divergence time in angiosperm lineages (Do, Kim & Kim, 2014; Nguyen, Kim & Kim, 2015; Kim, Kim & Kim, 2016; Kim & Kim, 2018). Repetitive sequences in the chloroplast genome provide useful information for studying genomic rearrangement and phylogeny (Cavalier-Smith, 2002; Nie et al., 2012; Yi et al., 2013; Kim & Kim, 2018). In addition, existing repeats might result in the accumulation of new repeats in cpDNA (Asano et al., 2004). One of the crucial molecular data types in cpDNA is SSR sequences. Other studies have used SSRs to develop specific markers for different species and to study the genetic diversity of angiosperms (Ishikawa, Sakaguchi & Ito, 2016; Luo et al., 2016; Marochio et al., 2017; Han et al., 2018). In this study, although cpDNA sequences were highly conserved, the three Carduus species were found to have different numbers of SSRs (Table S4). While we did not develop SSR markers or conduct population studies of Carduus, the SSR information we obtained may be useful in future studies on Carduus species. In addition to the repeats shared among the three species, C. crispus had three unique repetitive sequences (Table S4), which may be useful in population studies, phylogenetic analyses, and the development of additional molecular markers.
Uncertain relationships among Carduus
Phylogenetic analyses of the Asteraceae have identified ambiguous relationships between Carduus, Cirsium, and Silybum (Fu et al., 2016; Panero, 2016; Arnelas et al., 2018); for example, ITS data suggests that Carduus is polyphyletic (Häffner & Hellwig, 1999). Although three coding regions (matK, rbcL, and ndhF) were used to reconstruct phylogenetic relationships, the position of Carduus remained unresolved (Fu et al., 2016). We used 78 protein-coding regions to clarify these relationships; however, the phylogeny of Carduus and related taxa remains unclear (Fig. 4). Specifically, C. acanthoides was found to be close to Silybum marianum whereas C. crispus and C. tenuiflorus form a clade with Cirsium arvense. While non-coding regions can be useful in reconstructing the phylogeny of lower taxa, we were unable to recover the monophyly of Carduus using data from non-coding regions, including the eight hotspot areas as well as the combined data from coding and non-coding regions (Fig. S4). These issues suggest the need for additional studies on the phylogeny of Carduus and other members of the subfamily Carduoideae using supplementary molecular data and morphology.
Implications of SNP data for developing molecular markers for Carduus
SNPs are useful in population studies due to its extremely abundant presence in the angiosperms genomes (Cui et al., 2017; Fischer et al., 2017; Pantoja et al., 2017), and are effective in phylogenetic analysis (Leaché & Oaks, 2017). In addition, various molecular markers have been developed for different angiosperm species based on SNP data from chloroplast genomes (Khlestkina & Salina, 2006; Wang et al., 2015a; Wang et al., 2015b; Hyun et al., 2019; Xia et al., 2019). We successfully developed a molecular marker, inferred from SNP data, to distinguish C. crispus from C. acanthoides and C. tenuiflorus (Fig. S3). Our marker demonstrates that nucleotide sequence variations can provide rapid molecular identification of C. crispus. We focused on C. crispus because it exhibits the characteristics of an invasive species (Dunn, 1976; Verloove, 2014; Jung et al., 2017), and may also have value for the treatment of obesity and cancer (Xie, Li & Jia, 2005; Davaakhuu, Sukhdolgor & Gereltu, 2010; Lee et al., 2011; Tunsag, Davaakhuu & Batsuren, 2011). Various DNA-based markers (i.e., inter-simple sequence repeats [ISSRs], sequence characterisation of amplified regions [SCARs], and SSRs) have been developed to authenticate medicinal plants to ensure safety and efficacy (Hao et al., 2010; Sarwat et al., 2012; Ganie et al., 2015; Ward, Gaskin & Wilson, 2008). Additionally, molecular data are useful for understanding invasion processes of alien plants (Ward et al., 2008). We developed a SNP-based molecular marker for C. crispus (Fig. 5, Figs. S5 and S6) that may be used to detect C. crispus invasions in their early stages, and to develop suitable management strategies. Our alignment results also identified specific SNPs for C. acanthoides and C. tenuiflorus, which may be used to create molecular markers for these species (Fig. S1). Although we only used the SNP in matK, use of the complete cpDNA sequences of Carduus will enable the mining of SNPs from other regions for developing molecular markers for C. crispus and related species.
Conclusions
In this study, we provided the first complete cpDNA sequences for Carduus species. Despite the absence of significant differences (i.e., inversions, deletions, and duplications) between the chloroplast genomes of Carduus and those of related taxa, the newly acquired cpDNA sequences have value as a resource in future studies of the evolution of the chloroplast genome in Carduoideae and Asteraceae. Additionally, the 78 protein-coding regions of the chloroplast genome revealed uncertainty regarding the position of Carduus within the subfamily Carduoideae, and suggested the need for additional studies to reconstruct relationships not only among thistles, but among other members of the Asteraceae as well. The methods and protocols used in developing molecular markers for C. crispus are easy to apply and may be useful as a standard method in other studies of Asteraceae species.
Supplemental Information
Supplemental tables and figures
Table S1: List of species for phylogenomic analysis and whole cpDNA alignment. Table S2: List of regions for calculating Pi values. Table S3. Genes composition of the Carduus chloroplast genomes. Table S4: Feature of SSR in three Carduus species. Table S5: Feature of repeats in three Carduus species. Figure S1: The alignment of matK among three Carduus. The asterisk indicates the positions of single nucleotide polymorphism (SNP). The red square indicates SNP site for design primer pairs. Figure S2: The design of the primer pairs based on SNP site that is specific for Carduus crispus. Figure S3: The Bayesian Inference tree of Carduus and related taxa inferred from whole cpDNA sequences (A), non-coding regions of cpDNA (B), and eight hotspot regions (C).Figure S4: The PCR results of specific primer pairs for Carduus crispus. The combination of matK_463F, matK_1162R, CD_SNP_F1, and CD_SNP_R1. The number from 1 to 4: Carduus crispus; from 5 to 8: Carduus acanthoides; from 9 to 12: Carduus tenuiflorus. Figure S5: The PCR results of specific primer pairs for Carduus crispus. The combination of matK_463F, matK_1162R, CD_SNP_F2, and CD_SNP_R2. The number from 1 to 4: Carduus crispus; from 5 to 8: Carduus acanthoides; from 9 to 12: Carduus tenuiflorus. Figure S6: The MAUVE alignment of chloroplast genomes among Carduus and related species.